code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: stdl_env
# language: python
# name: stdl_env
# ---
import gc
import scanit
import torch
import random
import scanpy as sc
import pandas as pd
import anndata
import numpy as np
from scipy import sparse
from sklearn.metrics import normalized_mutual_info_score, adjusted_rand_score
from sklearn.cluster import SpectralClustering, KMeans
import matplotlib.pyplot as plt
import stlearn as st
from pathlib import Path
sp_datadir = "./data"
pts = np.loadtxt(sp_datadir+'/positions.csv')
X_sp = sparse.load_npz(sp_datadir+'/expression.npz')
X_sp = X_sp.toarray()
genes_sp = np.loadtxt(sp_datadir+'/genes.txt', dtype=str)
df_sp = pd.DataFrame(data=X_sp, columns=genes_sp)
adata = anndata.AnnData(X = X_sp, var=pd.DataFrame(index=genes_sp))
adata.obsm['spatial'] = pts
adata.obsm['spatial'][:,1] = -adata.obsm['spatial'][:,1]
adata.shape
n_sv_genes = 3000
adata_sp = adata.copy()
sc.pp.normalize_total(adata_sp)
df_somde = pd.read_csv('./data/somde_result.csv')
sv_genes = list( df_somde['g'].values[:n_sv_genes] )
adata_sp = adata_sp[:, sv_genes]
sc.pp.log1p(adata_sp)
sc.pp.scale(adata_sp)
scanit.tl.spatial_graph(adata_sp, method='alpha shape', alpha_n_layer=1, knn_n_neighbors=5)
scanit.tl.spatial_representation(adata_sp, n_h=10, n_epoch=2000, lr=0.001, device='cuda', n_consensus=5, projection='mds',
python_seed=0, torch_seed=0, numpy_seed=0)
sc.pp.neighbors(adata_sp, use_rep='X_scanit', n_neighbors=10)
sc.tl.leiden(adata_sp, resolution=0.3)
import plotly
pts = adata_sp.obsm['spatial']
color_list = plotly.colors.qualitative.Light24
labels_pred = np.array( adata_sp.obs['leiden'], int )
labels = list(set(labels_pred))
labels = list(np.sort(labels))
for j in range(len(labels)):
label = labels[j]
idx = np.where(labels_pred == label)[0]
plt.scatter(pts[idx,0], -pts[idx,1], c=color_list[j], label = label, s=15, linewidth=0)
plt.axis('equal')
plt.axis('off')
plt.legend(markerscale=3, loc='center left', bbox_to_anchor=(1,0.5), fontsize=12)
plt.tight_layout()
plt.savefig('./figures/scanit_segmentation.pdf', bbox_inches='tight')
adata_sp_marker = adata.copy()
adata_sp_marker.obs['leiden'] = adata_sp.obs['leiden']
sc.pp.normalize_total(adata_sp_marker)
sc.pp.log1p(adata_sp_marker)
adata_sp_marker.raw = adata_sp_marker
sc.pp.scale(adata_sp_marker)
sc.tl.rank_genes_groups(adata_sp_marker, 'leiden', method='wilcoxon')
sc.pl.rank_genes_groups(adata_sp_marker, n_genes=25, sharey=False)
result = adata_sp_marker.uns['rank_genes_groups']
groups = result['names'].dtype.names
marker_genes = []
for group in groups:
marker_genes.extend(list(result['names'][group][:3]))
import matplotlib as mpl
mpl.rcParams['font.size'] = 17
gs = sc.pl.matrixplot(adata_sp_marker, marker_genes, groupby='leiden', dendrogram=False,
use_raw=True, cmap='Blues', swap_axes=True, figsize=(5,6), save="scanit_segmentation.pdf")
result = adata_sp_marker.uns['rank_genes_groups']
groups = result['names'].dtype.names
marker_genes = []
for group in groups:
marker_genes.extend(list(result['names'][group][:1]))
adata_sp_plot = adata.copy()
sc.pp.normalize_total(adata_sp_plot)
sc.pp.log1p(adata_sp_plot)
pts = adata_sp_plot.obsm['spatial']
for i in range(len(marker_genes)):
expr = np.array(adata_sp_plot[:,marker_genes[i]].X).reshape(-1)
idx = np.argsort(expr)
plt.scatter(pts[idx,0], -pts[idx,1], c=expr[idx], linewidth=0, s=10, cmap='coolwarm')
plt.axis('equal'); plt.axis('off')
plt.colorbar()
plt.savefig("./figures/feature_plot_domain%d_%s.pdf" %(i, marker_genes[i]))
plt.clf()
# ### **Annotated cell types (by Giotto)**
X = pd.read_csv("./giotto/data/raw_exprs.csv", index_col=0)
adata_sc = anndata.AnnData(X=X)
celltypes = pd.read_csv("./giotto/data/celltype_annotation.csv", index_col=0)
adata_sc.obs['celltype_annotated'] = celltypes['x'].astype('category').values
df_pts = pd.read_csv("./giotto/data/spatial_locs.csv", index_col=0)
pts = df_pts[['sdimx', 'sdimy']].values
pts[:,1] = -pts[:,1]
adata_sc.obsm['spatial'] = pts
sc.pl.spatial(adata_sc, spot_size=100, color='celltype_annotated', frameon=False, palette='tab20', legend_fontsize=12, save="_giotto_celltype.pdf")
adata_sc_marker = adata_sc.copy()
sc.pp.normalize_total(adata_sc_marker)
sc.pp.log1p(adata_sc_marker)
adata_sc_marker.raw = adata_sc_marker
sc.pp.scale(adata_sc_marker)
sc.tl.rank_genes_groups(adata_sc_marker, 'celltype_annotated', method='wilcoxon')
sc.pl.rank_genes_groups(adata_sc_marker, n_genes=25, sharey=False)
result = adata_sc_marker.uns['rank_genes_groups']
groups = result['names'].dtype.names
marker_genes = []
for group in groups:
marker_genes.extend(list(result['names'][group][:3]))
import matplotlib as mpl
mpl.rcParams['font.size'] = 13
gs = sc.pl.matrixplot(adata_sc_marker, marker_genes, groupby='celltype_annotated', dendrogram=False,
use_raw=True, cmap='Blues', swap_axes=True, figsize=(5,6), save="celltype_markers.pdf")
| examples/seqFISH-mouse-SScortex/scanit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
from functools import wraps
type('a').__mro__
def extract_var(func, var, i=1):
return {k: v for k, v in sys._getframe(i).f_locals.items()
if not k.startswith('__')}
def foo(a, b, c=3, **kwargs):
d = a + b
return a*b*c
foo(5, 4)
res = extract_var(foo, 'd', 2)
res
from contextlib import redirect_stdout
from functools import partial
import io
import os
from pathlib import Path
def log_stdout(func=None, fname=''):
if not func:
return partial(log_stdout, fname=Path(fname))
if not fname:
fname = Path(f'./logs/{func.__name__}.log')
@wraps(func)
def wrapper(*args, **kwargs):
os.makedirs(fname.parent, exist_ok=True)
with open(fname, 'w') as f:
with redirect_stdout(f):
out = func(*args, **kwargs)
return out
return wrapper
# @log_stdout
def foo(a, b):
for i in range(a):
print(i*b, i/a, i*'-')
foo(3, 4)
# !ls logs
# !cat logs/foo.log
def log_stdout_new(func=None, fname=''):
if not func:
return partial(log_stdout, fname=Path(fname))
if not fname:
fname = Path(f'./logs/{func.__name__}.log')
@wraps(func)
def wrapper(*args, **kwargs):
os.makedirs(fname.parent, exist_ok=True)
with open(fname, 'w') as f:
with redirect_stdout(f):
out = func(*args, **kwargs)
return out
return wrapper
class MultiOutput:
def __init__(self):
self.stored = ''
def write(self, data):
print(type(data))
# self.stored += data
# print(data)
with redirect_stdout(MultiOutput()):
foo(3, 4)
| notebooks/scratch_extract_var.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import time
import numpy as np
x1 = [9, 2, 5, 0, 0, 7, 5, 0, 0, 0, 9, 2, 5, 0, 0]
x2 = [9, 2, 2, 9, 0, 9, 2, 5, 0, 0, 9, 2, 5, 0, 0]
### VECTORIZED DOT PRODUCT OF VECTORS ###
tic = time.process_time()
dot = np.dot(x1,x2)
toc = time.process_time()
print ("dot = " + str(dot) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
### VECTORIZED OUTER PRODUCT ###
tic = time.process_time()
outer = np.outer(x1,x2)
toc = time.process_time()
print ("outer = " + str(outer) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
### VECTORIZED ELEMENTWISE MULTIPLICATION ###
tic = time.process_time()
mul = np.multiply(x1,x2)
toc = time.process_time()
print ("elementwise multiplication = " + str(mul) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
### VECTORIZED GENERAL DOT PRODUCT ###
tic = time.process_time()
dot = np.dot(W,x1)
toc = time.process_time()
print ("gdot = " + str(dot) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
| coursera/deeplearning/week2/week2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import nltk
data = pd.read_json('https://raw.githubusercontent.com/risan/quran-json/master/json/translations/en.pretty.json')
data
data.drop(['surah_number'],axis=1,inplace=True)
data
data.describe()
data.head(27)
data.info()
| notebook/eda/Suraiya-Project/old-scrap-nltk/eng-json-csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Plot tractometry results
#
# You have to install seaborn 0.9 and run the following script.
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from tractseg.libs import exp_utils
values = {}
#Load tractometry results subject 1
values["s0"] = np.loadtxt("Tractometry_subject1.csv", delimiter=";", skiprows=1).transpose()
#Load tractometry results subject 2
values["s1"] = np.loadtxt("Tractometry_subject2.csv", delimiter=";", skiprows=1).transpose()
#todo: load more subjects
subjects_controls = ["s0"]
subjects_patients = ["s1"]
#Define bundles you want to plot
selected_bundles = ["CST_right", "SCP_left", "UF_right", "CG_right", "CC_4", "AF_right"]
NR_POINTS = values[subjects_controls[0]].shape[1]
bundles = dataset_specific_utils.get_bundle_names("All")[1:]
selected_bun_indices = [bundles.index(b) for b in selected_bundles]
a4_dims = (15, 10)
#You have to adapt nr of subplots if you want to show more bundles
f, axes = plt.subplots(2, 3, figsize=a4_dims)
axes = axes.flatten()
sns.set(font_scale=1.2)
sns.set_style("whitegrid")
for i, b_idx in enumerate(selected_bun_indices):
#Bring data into right format for seaborn
data = { "position": [],
"fa": [],
"group": [],
"subject": []}
for j, subject in enumerate(subjects_controls + subjects_patients):
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
if subject in subjects_controls:
data["group"].append("Group A")
else:
data["group"].append("Group B")
#Plot
ax = sns.lineplot(x="position", y="fa", data=data, ax=axes[i], hue="group")
ax.set(xlabel='position', ylabel='FA')
ax.set_title(bundles[b_idx])
if i > 0:
ax.legend_.remove() #only show legend on first subplot
plt.tight_layout() #If this command creates errors you can remove it
plt.show()
#save as image
#plt.savefig("tractometry_plot.png", dpi=200)
| examples/.ipynb_checkpoints/plot_tractometry_results-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Upload data to CARTO
#
# This example illustrates how to upload local data to a CARTO account.
#
# _Note: You'll need [CARTO Account](https://carto.com/signup) credentials to reproduce this example._
# +
from cartoframes.auth import set_default_credentials
set_default_credentials('creds.json')
# +
from cartoframes import read_carto
gdf = read_carto("SELECT * FROM starbucks_brooklyn WHERE revenue > 1200000")
gdf.head()
# +
from cartoframes import to_carto
to_carto(gdf, 'starbucks_brooklyn_filtered', if_exists='replace')
| docs/examples/data_management/upload_to_carto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SarahRebulado/OOP1_2/blob/main/OOP_CONCEPTS_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="2kKaWebOVITZ"
# Classes with Multiple Objects
# + colab={"base_uri": "https://localhost:8080/"} id="v3wkkloXT1WW" outputId="b15613fa-edf7-42be-d013-fedb052bcfa1"
class Birds:
def __init__(self, bird_name):
self.bird_name= bird_name
def flying_birds(self):
print(f"{self.bird_name}flies above the sky")
def non_flying_birds(self):
print(f"{self.bird_name}is the national bird of the Philippines")
vulture= Birds("Griffon Vulture")
crane= Birds("Common Crane")
emu= Birds("Emu")
vulture.flying_birds()
crane.flying_birds()
emu.non_flying_birds()
# + [markdown] id="hTkcWLmtXVEh"
# Encapsulation using mangling with double underscores
# + colab={"base_uri": "https://localhost:8080/"} id="GNxa7vSfXUqu" outputId="377717ff-12b4-4154-f654-e109fadf5eaa"
class foo:
def __init__(self, a, b):
self.__a= a
self.__b= b
def add(self):
return self.__a + self.__b #Private attributes
number= foo(3,4)
number.add()
number.a= 7
number.add()
# + [markdown] id="7jTRaEhNY-rq"
# Encapsulation with Private Attributes
# + colab={"base_uri": "https://localhost:8080/"} id="v2O2ZoJWY-_Y" outputId="de8cbfe2-42c3-4165-dd8c-53ad8ab02316"
class Counter:
def __init__(self):
self.__current=0
def increment(self):
self.__current+=1
def value(self):
return self.__current
def reset(self):
self.__current=0
num=Counter()
num.counter= 1
num.increment() #counter=counter+1
num.increment()
num.increment()
num.value()
# + [markdown] id="qfloJfU1dJ16"
# Inheritance
# + colab={"base_uri": "https://localhost:8080/"} id="bD-yttdDchAC" outputId="e695a7f0-4dc5-48bd-8073-7c3f3ede330c"
class Person:
def __init__(self, firstname, surname):
self.firstname= firstname
self.surname= surname
def printname(self):
print(self.firstname,self.surname)
person=Person("Ana","Santos")
person.printname()
class Teacher(Person):
pass
person2=Teacher("Maria","Sayo")
person2.printname()
class Student(Person):
pass
person3= Student("Jhoriz", "Aquino")
person3.printname()
# + [markdown] id="1izO_HW1fK1-"
# Polymorphism
# + colab={"base_uri": "https://localhost:8080/"} id="RzzjPyxkfMfx" outputId="afebdab6-1106-4aca-806c-2ce34d41e645"
class RegularPolygon:
def __init__(self,side):
self.side = side
class Square(RegularPolygon):
def area(self):
return self.side * self.side
class EquilateralTriangle(RegularPolygon):
def area(self):
return self.side * self.side * 0.433
object= Square(4)
print(object.area())
object2=EquilateralTriangle(3)
print(object2.area())
# + [markdown] id="omAKbmzZhknl"
# Application 1
#
# 1. Create a Python program that the name of three students (Student 1, Student 2, and Student 3) and their term grades
# 2. Create a class name Person and attributes -std1, std2,, pre, mid, fin
# 3. Compute the average of each term grade using Grade() method
# 4. Information about student's grades must be hidden from others
# + colab={"base_uri": "https://localhost:8080/"} id="mLPFMcE73aVS" outputId="25ced18d-870a-466a-d08f-f2dced2a4949"
class Person():
def __init__(self, std, pre, mid, fin):
self.std= std
self.pre= pre
self.mid= mid
self.fin= fin
def Grade(self):
return round((self.pre + self.mid + self.fin)/3,2)
class Anna_Smith(Person):
pass
print("Student1 Grade")
std1= str(input("Name:"))
pre1= float(input("Enter Prelim Grade: "))
mid1= float(input("Enter Midterm Grade: "))
fin1= float(input("Enter Final Grade: "))
Anna_Smith= Person(std1, pre1, mid1, fin1)
print()
class Jona_Reyes(Person):
pass
print("Student2 Grade")
std2= str(input("Name: "))
pre2= float(input("Enter Prelim Grade: "))
mid2= float(input("Enter Midterm Grade: "))
fin2= float(input("Enter Final Grade: "))
Jona_Reyes= Person(std2, pre2, mid2, fin2)
print()
class Michele_Cruz(Person):
pass
print("Student3 Grade")
std3= str(input("Name: "))
pre3= float(input("Enter Prelim Grade: "))
mid3= float(input("Enter Midterm Grade: "))
fin3= float(input("Enter Final Grade: "))
Michele_Cruz= Person(std3, pre3, mid3, fin3)
print()
print("View Student's Grade")
Student_Name= str(input("Enter Name: "))
if Student_Name==std1:
print("Average Grade: ", Anna_Smith.Grade())
else:
if Student_Name==std2:
print("Average Grade: ", Jona_Reyes.Grade())
else:
if Student_Name==std3:
print("Average Grade: ", Michele_Cruz.Grade())
else:
print("No student record.")
| OOP_CONCEPTS_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
data1 = np.random.normal(0, 1, 1000)
data2 = np.random.normal(0, 1, size=(2, 1000))
data3 = np.random.normal(0, 1, size=(4, 1000))
# -
# # Shannon Entropy
# Shannon entropy H is given by the formula $H=-\sum_{i}p_{i}\log_{b}(p_{i})$
# where $p_{i}$ is the probability of character number $i$ appearing in the
# stream of characters of the message.
#
# Consider a simple digital circuit which has a two-bit input ($X$, $Y$) and a two-bit output ($X$ and $Y$, $X$ or $Y$). Assuming that the two input bits $X$ and $Y$ have mutually independent chances of $50%$ of being *HIGH*, then the input combinations $(0,0)$, $(0,1)$, $(1,0)$, and ($1,1)$ each have a 1/4 chance of occurring, so the circuit's Shannon entropy on the input side is $H(X,Y)=4{\Big (}-{1 \over 4}\log _{2}{1 \over 4}{\Big )}=2$ Then the possible output combinations are (0,0), (0,1) and (1,1) with respective chances of 1/4, 1/2, and 1/4 of occurring, so the circuit's Shannon entropy on the output side is $H(X{\text{ and }}Y,X{\text{ or }}Y)=2{\Big (}-{1 \over 4}\log _{2}{1 \over 4}{\Big )}-{1 \over 2}\log _{2}{1 \over 2}=1+{1 \over 2}=1{1 \over 2}$, so the circuit reduces (or "orders") the information going through it by half a
# bit of Shannon entropy due to its logical irreversibility.
# +
from gcpds.entropies import Shannon
ent = Shannon(data1)
print(f"Input data shape: {data1.shape}")
print(f"Entropy: {ent}", end='\n\n')
# -
Shannon(data1, base=10) # Default base is 2
Shannon(data1, bins=12) # Default bins value used to calculate the distribution is 16
Shannon(data2, conditional=1)
# ## Joint entropy
#
# For 2 variables:
# ${\displaystyle \mathrm {H} (X,Y)=-\sum _{x\in {\mathcal {X}}}\sum _{y\in {\mathcal {Y}}}P(x,y)\log _{2}[P(x,y)]}$
# +
ent = Shannon(data2)
print(f"Input data shape: {data2.shape}")
print(f"Entropy: {ent}")
# -
# For more than two random variables ${\displaystyle X_{1},...,X_{n}} X_{1},...,X_{n}$ this expands to
# ${\displaystyle \mathrm {H} (X_{1},...,X_{n})=-\sum _{x_{1}\in {\mathcal {X}}_{1}}...\sum _{x_{n}\in {\mathcal {X}}_{n}}P(x_{1},...,x_{n})\log _{2}[P(x_{1},...,x_{n})]}$
# +
ent = Shannon(data3)
print(f"Input data shape: {data3.shape}")
print(f"Entropy: {ent}")
# -
# ## Conditional entropy
#
# Joint entropy is used in the definition of conditional entropy
# ${\displaystyle \mathrm {H} (X|Y)=\mathrm {H} (X,Y)-\mathrm {H} (Y)}$
# +
ent = Shannon(data3, conditional=0) # `conditional` is an index of the input array
print(f"Input data shape: {data3.shape}")
print(f"Entropy: {ent}")
# -
# ----
# ### References
#
# * <NAME>; <NAME>. Elements of Information Theory. Hoboken, New Jersey: Wiley. ISBN 0-471-24195-4.
| notebooks/01-shannon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from keras.datasets import mnist
from keras.utils import np_utils
from keras.models import Sequential,load_model
from keras.layers import Dense
from keras.optimizers import SGD,Adam
# +
(x_train,y_train),(x_test,y_test) = mnist.load_data()
print('x_shpae:',x_train.shape)
print('y_shapr:',y_train.shape)
x_train = x_train.reshape(x_train.shape[0],-1)/255.0
x_test = x_test.reshape(x_test.shape[0],-1)/255.0
y_train = np_utils.to_categorical(y_train,num_classes=10)
y_test = np_utils.to_categorical(y_test,num_classes=10)
model = load_model('model.h5')
loss,accuracy = model.evaluate(x_test,y_test)
print('\n terst loss ',loss)
print('accuracy:',accuracy)
# -
| test/loadmodel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''pyenv'': venv)'
# language: python
# name: python38564bitpyenvvenvbe3b140a556347e4bb4bb25d3b42328a
# ---
# +
from IPython.display import Audio
from IPython.display import Video
from IPython.display import Image
from moviepy import *
from moviepy.editor import *
# -
w,h = moviesize = (720, 380)
# +
# >>> myClip.resize(lambda t : 1+0.02*t) # slow swelling of the clip
# -
clip = VideoFileClip("data/footage/gradient.mp4").subclip(10,15).resize((640, 480))
music = AudioFileClip("data/sound/sunday.mp3")
# +
txt_clip = TextClip("Replace value\nin each cell", font='Bungee-Shade', fontsize=60,color='white').set_pos('center').set_duration(4)
video = CompositeVideoClip([clip, txt_clip]).set_audio(music.subclip(0, clip.duration))
# -
myClip = video.resize(0.60)
# +
# video.write_videofile("vid.mp4", preset="ultrafast", threads=4)
# video.write_videofile("vid.webm")
# + active=""
# # To demo stuff : stackoverlfow.com/18019477
# display(audio)
# Video("somevideofile.mp4")
# display(Image("somefile.png"))
# +
# Ways to preview video
# myClip.show()
# video.preview()
| pysc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Luther - Data Cleaning and Merging Dataframes
# This is part 2 of the Luther Project. In part 1, I've created 6 different dataframes that will be merged and cleaned in this notebook. The final merged dataframe "merged2.pkl" will be used in the last notebook "03 - Luther - Linear Regression" to develop a final linear regression model for predicting movie ticket sales on opening week (as well as opening gross adjusted to ticket price).
import pickle
import requests
from bs4 import BeautifulSoup
import re
import pandas as pd
import time
import seaborn as sns
import datetime
# Import all the dataframes
mojog_df = pd.read_pickle('data/mojog_2018.pkl')
#unemp_df = pd.read_pickle("data/unemp_df.pkl")
youtube_df = pd.read_pickle('data/youtube_2018.pkl')
omdb_df = pd.read_pickle('data/omdb_2018.pkl')
director_df = pd.read_pickle('data/director_df.pkl')
actor_df = pd.read_pickle('data/actor_df.pkl')
#clean mojo df
mojog_df.opening = mojog_df.opening.replace("\$","", regex = True).replace(",","", regex = True).str.strip()
mojog_df.opening = pd.to_numeric(mojog_df.opening)
mojog_df.tot_gross = mojog_df.tot_gross.replace("\$","", regex = True).replace(",","", regex = True).str.strip()
mojog_df.tot_gross = pd.to_numeric(mojog_df.tot_gross)
mojog_df.theaters = mojog_df.theaters.replace(",","", regex = True).str.strip()
mojog_df.theaters = pd.to_numeric(mojog_df.theaters)
mojog_df.info()
# +
# Clean youtube df
# select youtube df relevant data
youtube_df = youtube_df[["movie_name", "viewCount", "commentCount", "dislikeCount", "likeCount"]]
#replace nulls with 0
youtube_df = youtube_df.fillna(0)
# convert numeric str values to int
youtube_df.viewCount = pd.to_numeric(youtube_df.viewCount)
youtube_df.dislikeCount = pd.to_numeric(youtube_df.dislikeCount)
youtube_df.commentCount = pd.to_numeric(youtube_df.commentCount)
youtube_df.likeCount = pd.to_numeric(youtube_df.likeCount)
youtube_df = youtube_df.rename(columns ={"viewCount":"Yviews", "commentCount": "Ycomments",
"dislikeCount": "Ydislikes", "likeCount":"Ylikes"})
youtube_df.info()
# -
omdb_df
# Clean omdb data
# omdb_df.imdb = pd.to_numeric(omdb_df.imdb.replace("/10","", regex = True).str.strip())
# omdb_df.metacritic = pd.to_numeric(omdb_df.metacritic.replace("/100","", regex = True).str.strip())
# omdb_df.rotten_tomatoes = pd.to_numeric(omdb_df.rotten_tomatoes.replace("%","", regex = True).str.strip())
# omdb_df.runtime = pd.to_numeric(omdb_df.runtime.replace("min","", regex = True).str.strip())
# omdb_df.year = pd.to_numeric(omdb_df.year)
omdb_df.info()
# Clean directors data
# Convert str to numeric
director_df.dir_agross = pd.to_numeric(director_df.dir_agross.replace("\$","", regex = True).str.strip())
director_df.dir_gross = pd.to_numeric(director_df.dir_gross.replace("\$","", regex = True)
.replace(",","", regex = True).str.strip())
director_df.dir_nmovies = pd.to_numeric(director_df.dir_nmovies)
director_df.info()
# Clean directors data
# Convert str to numeric
actor_df.act_agross = pd.to_numeric(actor_df.act_agross.replace("\$", "", regex = True)
.replace(",", "", regex = True).str.strip())
actor_df.act_gross = pd.to_numeric(actor_df.act_gross.replace("\$", "", regex = True)
.replace(",", "", regex = True).str.strip())
actor_df.act_nmovies = pd.to_numeric(actor_df.act_nmovies)
actor_df.info()
actor_df.head()
# +
# # Split actor & director strings into lists
# # for index in range(len(omdb_df)):
# # omdb_df.actors[index] = [x.strip() for x in omdb_df.actors[index].split(',')]
# omdb_df.actors = omdb_df.actors.apply(lambda x :[x.strip() for x in x.split(',')])
# omdb_df.director = omdb_df.director.apply(lambda x :[x.strip() for x in x.split(',')])
# omdb_df.head()
# -
mojog_df
# +
# unemp rate for this april 2018 was 4.1
mojog_df["unemp_rate"] = 4.1
mojog_df.head()
# -
# # Merge the dataframes
# Merge mojo and youtube
merged = pd.DataFrame.merge(mojog_df, youtube_df,on='movie_name', how = 'inner')
merged.head()
# Merge omdb
merged2 = pd.DataFrame.merge(merged,omdb_df,on="movie",how="inner")
merged2.head()
# Adjust the youtube views based on google trends data
merged2["Yviews_adj"] = round(merged2.Yviews * merged2.gtrend)
merged2["Ylikes_adj"] = round(merged2.Ylikes * merged2.gtrend)
merged2["Ydis_adj"] = round(merged2.Ydislikes * merged2.gtrend)
merged2["Ycom_adj"] = round(merged2.Ycomments * merged2.gtrend)
#merged2["Yviews_adj2"] = round(merged2.Yviews * merged2.gtrend)
merged2 = merged2.drop(["gtrend","Yviews","Ycomments","Ylikes","Ydislikes"],1)
#merged2 = merged2.drop(["gtrend","gtrend2"],1)
merged2
# First create a dataframe for year and corresponding movie ticket
merged2["tick"] = 9.16
# Create new column for merged
est_tick = (merged2.opening // merged2.tick).astype("int64")
merged2.insert(loc=3, column='est_tick', value=est_tick)
merged2 = merged2.rename(columns={"opening":"op_gross"})
merged2 = merged2.fillna(0)
merged2.columns
# # Generate a metric for actor and directors
# #### actors
# I took the max total gross or total # movies featured for the actor/director for each given movie.
# +
import numpy as np
act_list = []
for i in range(len(merged2)):
#for each of the actors in a given movie, sum up their total gross & movies
gross_list = []
nmovies_list = []
if (merged2.actors[i] == 0) or (merged2.actors[i] == []):
act_gross, act_nmovies = 0, 0
else:
actor_count = len(merged2.actors[i])
for actoriter in merged2.actors[i]:
if actor_df[actor_df.actor == actoriter].empty:
actor_count -= 1
#Subtract the movie total gross (movie i want to predict) from actor total gross
else:
gross_list.append(actor_df[actor_df.actor == actoriter].act_gross.iloc[0] - \
merged2.tot_gross[i]/(1e6))
nmovies_list.append(actor_df[actor_df.actor == actoriter].act_nmovies.iloc[0])
#take the sum and average over number of actors featured
if (actor_count <= 0) or (merged2.actors[i] == []) or (merged2.actors[i] == 0):
act_gross, act_nmovies = 0, 0
else:
act_gross = round(max(gross_list),1)
act_nmovies = round(max(nmovies_list),1)
act_dict = {'movie': merged2.movie[i], 'act_gross':act_gross,
'act_nmovies': act_nmovies}
act_list.append(act_dict)
actor_metric = pd.DataFrame(act_list)
# -
# #### directors
# +
import numpy as np
dir_list = []
for i in range(len(merged2)):
#for each of the actors in a given movie, sum up their total gross & movies
gross_list = []
nmovies_list = []
if (merged2.directors[i] == 0) or (merged2.directors[i] == []):
dir_gross, dir_nmovies = 0, 0
else:
director_count = len(merged2.directors[i])
for directoriter in merged2.directors[i]:
if director_df[director_df.director == directoriter].empty:
director_count -= 1
#Subtract the movie total gross (movie i want to predict) from actor total gross
else:
gross_list.append(director_df[director_df.director == directoriter].
dir_gross.iloc[0] - \
merged2.tot_gross[i]/(1e6))
nmovies_list.append(director_df[director_df.director == directoriter]
.dir_nmovies.iloc[0])
#take the sum and average over number of actors featured
if (director_count <= 0) or (merged2.directors[i] == []) or (merged2.directors[i] == 0):
dir_gross, dir_nmovies = 0, 0
else:
dir_gross = round(max(gross_list),1)
dir_nmovies = round(max(nmovies_list),1)
dir_dict = {'movie': merged2.movie[i], 'dir_gross':dir_gross,
'dir_nmovies': dir_nmovies}
dir_list.append(dir_dict)
director_metric = pd.DataFrame(dir_list)
# -
# # Merge the director & actor metrics to df
merged2 = pd.merge(merged2, actor_metric, on="movie", how="inner")
merged2 = pd.merge(merged2, director_metric, on="movie", how="inner")
merged2 = merged2.drop(["actors","directors"],1)
merged2.head()
# ### Save the merged dataframe
merged2.to_pickle("data/merged_2018.pkl")
len(merged2[(merged2.dir_nmovies == 0)])# | (merged2.act_gross == 0)])
| 02-Movie_Opening_Gross_Prediction/Luther - test on 2018 avengers movie - part 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center> NumPy <center>
# <img src = "https://github.com/saeed-saffari/alzahra-workshop-spr2021/blob/main/lecture/PIC/Numpy.png?raw=true">
# ## Installation
#
# - Conda install numpy
# - pip install numpy
# - pip install --upgrade numpy
pip install --upgrade numpy
# ## Import
# ## Specification
np.array()
np.shape
np.size
reshape
np.linspace
round
np.zeros
np.ones
np.eye
np.diag
np.full
np.matmul
@
np.transpose
np.linalg.inv
np.linalg.det
np.random.randint
np.random.normal
np.mean
np.var
np.std
print ('Overall mean of matrix is \n%s\n'%np.mean(mat))
print ('Column mean of matrix is \n%s\n'%np.mean(mat, axis=1))
print ('Overall varience of matrix is %s\n'%np.var(mat))
print ('Overall standard deviation of matrix is %s\n'%np.std(mat))
print ('Overall sum of matrix is %s\n'%np.sum(mat))
print ('Overall min of matrix is %s\n'%np.min(mat))
print ('Overall max of matrix is %s\n'%np.max(mat))
v1 = np.arange(5)
print ('Creating a numpy array via arange (stop=5) : %s\n'%v1)
v2 = np.arange(2,5)
print ('Creating a numpy array via arange (start=2,stop=5) : %s\n'%v2)
v3 = np.arange(0,-10,-2)
print ('Creating a numpy array via arange (start=0,stop=-10,step=-2) : %s\n'%v3)
| Planning Economics/.ipynb_checkpoints/03 - NumPy-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 01 Procesamiento de Datos: [NOMBRE_DEL_PROYECTO]
#
# ##### Metas:
# 1. Limpiar las columnas XXX
# 2. Reordenar el **dataset** a un formato útil
# 3. Identificar variables útiles/de interés
#
# ##### Cambios:
# * fecha_del_ultimo_cambio: Descripción del cambio.
# * otro cambio: otra descripción.
#
# ***
# __Preparación__
import pandas as pd
from pathlib import Path
from datetime import datetime
# Inicializa rutas hacia los archivos.
fecha_hoy = datetime.today()
archivos_en_bruto = Path("../datos/en_bruto/")
archivos_procesados = Path("../datos/procesados/")
archivo_final = Path("../datos/procesados/") / f"datos_procesados_{fecha_hoy:%b-%d-%Y}.csv"
archivo = archivos_en_bruto / 'REEMPLAZA_ESTO_CON_EL_NOMBRE_DE_TU_ARCHIVO'
# +
# Leer y describir el conjunto de datos.
datos = pd.read_csv(archivo)
print("-"*25)
print(f"{datos.shape[0]} filas, {datos.shape[1]} columnas.")
print("-"*25)
datos.describe()
print("-"*25)
datos.head()
| planillas/01_procesar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# tgb - 11/26/2019 - Mimic notebook 029 but train on +4K to see what is missed on 0K
# tgb - 11/13/2019 - Continuity of 028 but for simultaneous training while files are pre-processing
# # 0) Imports
# +
from cbrain.imports import *
from cbrain.data_generator import *
from cbrain.cam_constants import *
from cbrain.losses import *
from cbrain.utils import limit_mem
from cbrain.layers import *
from cbrain.data_generator import DataGenerator
import tensorflow as tf
import tensorflow.math as tfm
from tensorflow.keras.layers import *
from tensorflow.keras.models import *
import xarray as xr
import numpy as np
from cbrain.model_diagnostics import ModelDiagnostics
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.image as imag
import scipy.integrate as sin
TRAINDIR = '/local/Tom.Beucler/SPCAM_PHYS/'
DATADIR = '/project/meteo/w2w/A6/S.Rasp/SP-CAM/fluxbypass_aqua/'
PREFIX = '8col009_01_'
# %cd /filer/z-sv-pool12c/t/Tom.Beucler/SPCAM/CBRAIN-CAM
# Otherwise tensorflow will use ALL your GPU RAM for no reason
limit_mem()
# -
# # 1) NN with only q and T as inputs
# ## 1.1) Rescaling
scale_dict = {
'PHQ': L_V/G,
'TPHYSTND': C_P/G,
'FSNT': 1,
'FSNS': 1,
'FLNT': 1,
'FLNS': 1,
}
# Takes representative value for PS since purpose is normalization
PS = 1e5; P0 = 1e5;
P = P0*hyai+PS*hybi; # Total pressure [Pa]
dP = P[1:]-P[:-1]; # Differential pressure [Pa]
for v in ['PHQ','TPHYSTND']:
scale_dict[v] *= dP
save_pickle('./nn_config/scale_dicts/100_POG_scaling.pkl', scale_dict)
in_vars = ['QBP','TBP','PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_vars = ['PHQ','TPHYSTND','FSNT','FSNS','FLNT','FLNS']
train_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/102_train_shuffle.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/100_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
valid_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/102_valid.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/100_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
X, Y = valid_gen[0]; X.shape, Y.shape
# ## 1.2) Model and training
inp = Input(shape=(64,))
densout = Dense(128, activation='linear')(inp)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (5):
densout = Dense(128, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(64, activation='linear')(densout)
out = LeakyReLU(alpha=0.3)(densout)
NNmodel = tf.keras.models.Model(inp,out)
NNmodel.summary()
earlyStopping = EarlyStopping(monitor='val_loss', patience=10, verbose=0, mode='min')
mcp_save = ModelCheckpoint('/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/POG102.hdf5',save_best_only=True, monitor='val_loss', mode='min')
NNmodel.compile(tf.keras.optimizers.Adam(),loss=mse)
# Trained for 15 epochs in total
Nep = 15
NNmodel.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen,callbacks=[earlyStopping,mcp_save])
# # 2) NN with only RH and T as inputs
# ## 2.1) Rescaling
in_vars = ['RH','TBP','PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_vars = ['PHQ','TPHYSTND','FSNT','FSNS','FLNT','FLNS']
train_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/105_train_shuffle.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/103_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
valid_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/105_valid.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/103_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
X, Y = valid_gen[0]; X.shape, Y.shape
# ## 2.2) Model and training
inp2 = Input(shape=(64,))
densout = Dense(128, activation='linear')(inp2)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (5):
densout = Dense(128, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(64, activation='linear')(densout)
out2 = LeakyReLU(alpha=0.3)(densout)
NNmodel2 = tf.keras.models.Model(inp2,out2)
NNmodel2.summary()
earlyStopping = EarlyStopping(monitor='val_loss', patience=10, verbose=0, mode='min')
mcp_save = ModelCheckpoint('/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/POG105.hdf5',save_best_only=True, monitor='val_loss', mode='min')
NNmodel2.compile(tf.keras.optimizers.Adam(),loss=mse)
Nep = 15
NNmodel2.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen,callbacks=[earlyStopping,mcp_save])
# # 3) NN with only QBP and TfromMA as inputs
# ## 3.1) Rescaling
in_vars = ['QBP','TfromMA','PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_vars = ['PHQ','TPHYSTND','FSNT','FSNS','FLNT','FLNS']
train_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/108_train_shuffle.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/106_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
valid_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/108_valid.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/106_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
X, Y = train_gen[0]; X.shape, Y.shape
# ## 3.2) Model and training
inp3 = Input(shape=(64,))
densout = Dense(128, activation='linear')(inp3)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (5):
densout = Dense(128, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(64, activation='linear')(densout)
out3 = LeakyReLU(alpha=0.3)(densout)
NNmodel3 = tf.keras.models.Model(inp3,out3)
NNmodel3.summary()
earlyStopping = EarlyStopping(monitor='val_loss', patience=10, verbose=0, mode='min')
mcp_save = ModelCheckpoint('/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/POG108.hdf5',save_best_only=True, monitor='val_loss', mode='min')
NNmodel3.compile(tf.keras.optimizers.Adam(),loss=mse)
Nep = 15
NNmodel3.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen,callbacks=[earlyStopping,mcp_save])
# # 4) NN using only RH and TfromMA as inputs
# ## 4.2) Rescaling
in_vars = ['RH','TfromMA','PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_vars = ['PHQ','TPHYSTND','FSNT','FSNS','FLNT','FLNS']
train_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/111_train_shuffle.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/109_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
valid_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/111_valid.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/109_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
X, Y = train_gen[0]; X.shape, Y.shape
# ## 4.3) Model and training
inp4 = Input(shape=(64,))
densout = Dense(128, activation='linear')(inp4)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (5):
densout = Dense(128, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(64, activation='linear')(densout)
out4 = LeakyReLU(alpha=0.3)(densout)
NNmodel4 = tf.keras.models.Model(inp4,out4)
NNmodel4.summary()
earlyStopping = EarlyStopping(monitor='val_loss', patience=10, verbose=0, mode='min')
mcp_save = ModelCheckpoint('/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/POG111.hdf5',save_best_only=True, monitor='val_loss', mode='min')
NNmodel4.compile(tf.keras.optimizers.Adam(),loss=mse)
Nep = 15
NNmodel4.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen,callbacks=[earlyStopping,mcp_save])
# ## 4.4) Preprocess +4K
# # 5) NN using QBP and Carnotmax as inputs
#
# ## 5.1) Rescaling
# +
in_vars = ['QBP','Carnotmax','PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_vars = ['PHQ','TPHYSTND','FSNT','FSNS','FLNT','FLNS']
train_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/114_train_shuffle.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/112_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
valid_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/114_valid.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/112_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
X, Y = train_gen[12]; X.shape, Y.shape
# -
# ## 5.2) Model and training
inp5 = Input(shape=(64,))
densout = Dense(128, activation='linear')(inp5)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (5):
densout = Dense(128, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(64, activation='linear')(densout)
out5 = LeakyReLU(alpha=0.3)(densout)
NNmodel5 = tf.keras.models.Model(inp5,out5)
NNmodel5.summary()
earlyStopping = EarlyStopping(monitor='val_loss', patience=10, verbose=0, mode='min')
mcp_save = ModelCheckpoint('/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/POG114.hdf5',save_best_only=True, monitor='val_loss', mode='min')
NNmodel5.compile(tf.keras.optimizers.Adam(),loss=mse)
Nep = 15
NNmodel5.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen,callbacks=[earlyStopping,mcp_save])
# # 6) NN using only Q and (T-TS) as inputs
# # 6.1) Rescaling
in_vars = ['QBP','TfromTS','PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_vars = ['PHQ','TPHYSTND','FSNT','FSNS','FLNT','FLNS']
train_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/120_train_shuffle.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/118_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
valid_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/120_valid.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/118_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
X, Y = train_gen[25]; X.shape, Y.shape
# ## 6.2) Model and training
inp6 = Input(shape=(64,))
densout = Dense(128, activation='linear')(inp6)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (5):
densout = Dense(128, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(64, activation='linear')(densout)
out6 = LeakyReLU(alpha=0.3)(densout)
NNmodel6 = tf.keras.models.Model(inp6,out6)
NNmodel6.summary()
# +
earlyStopping = EarlyStopping(monitor='val_loss', patience=10, verbose=0, mode='min')
mcp_save = ModelCheckpoint('/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/POG120.hdf5',save_best_only=True, monitor='val_loss', mode='min')
NNmodel6.compile(tf.keras.optimizers.Adam(),loss=mse)
Nep = 15
NNmodel6.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen,callbacks=[earlyStopping,mcp_save])
# -
# # 7) NN using RH and (T-Ts) as inputs
# +
in_vars = ['RH','TfromTS','PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_vars = ['PHQ','TPHYSTND','FSNT','FSNS','FLNT','FLNS']
train_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/123_train_shuffle.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/121_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
valid_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/123_valid.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/121_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
# -
X, Y = train_gen[42]; X.shape, Y.shape
inp7 = Input(shape=(64,))
densout = Dense(128, activation='linear')(inp7)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (5):
densout = Dense(128, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(64, activation='linear')(densout)
out7 = LeakyReLU(alpha=0.3)(densout)
NNmodel7 = tf.keras.models.Model(inp7,out7)
NNmodel7.summary()
# +
earlyStopping = EarlyStopping(monitor='val_loss', patience=10, verbose=0, mode='min')
mcp_save = ModelCheckpoint('/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/POG123.hdf5',save_best_only=True, monitor='val_loss', mode='min')
NNmodel7.compile(tf.keras.optimizers.Adam(),loss=mse)
Nep = 15
NNmodel7.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen,callbacks=[earlyStopping,mcp_save])
# -
# # 8) NN using RH and Carnotmax as inputs
# +
in_vars = ['RH','Carnotmax','PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_vars = ['PHQ','TPHYSTND','FSNT','FSNS','FLNT','FLNS']
train_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/126_train_shuffle.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/124_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
valid_gen = DataGenerator(
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/126_valid.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = '/local/Tom.Beucler/SPCAM_PHYS/124_norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
# -
X, Y = train_gen[12]; X.shape, Y.shape
inp8 = Input(shape=(64,))
densout = Dense(128, activation='linear')(inp8)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (5):
densout = Dense(128, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(64, activation='linear')(densout)
out8 = LeakyReLU(alpha=0.3)(densout)
NNmodel8 = tf.keras.models.Model(inp8,out8)
NNmodel8.summary()
# +
earlyStopping = EarlyStopping(monitor='val_loss', patience=10, verbose=0, mode='min')
mcp_save = ModelCheckpoint('/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/POG126.hdf5',save_best_only=True, monitor='val_loss', mode='min')
NNmodel8.compile(tf.keras.optimizers.Adam(),loss=mse)
Nep = 15
NNmodel8.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen,callbacks=[earlyStopping,mcp_save])
# -
| notebooks/tbeucler_devlog/031_NN_training_on_p4K_only.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Similarity & Clustering
# ## What is Clustering?
#
# A cluster is a collection of data objects or a natural grouping of any sort.
#
# * Data objects are **similar** to one another within the same cluster
# * Data objects are **dissimilar** to the objects in other clusters
#
# Clustering is a part of unsupervised learning and often part of EDA (exploratory data analysis) as an approach to analyze data sets to summarize their main characteristics. This is more specifically known as **unsupervised segmentation** as they are creating segments without predefined targets.
#
# **What are some applications of clustering?**
#
# Clustering has two broad categories of application. It can be a **stand-alone tool** to get insight into underlying data by grouping related data together, or it can be a **data pre-processing step** before other algorithms are run.
#
# A good clustering method produces high-quality clusters. A high-quality cluster would contain data objects showing high similarity within a cluster and low similarity across clusters. The quality of clusters depends on the metric at which we define similarity, how clustering was implemented, and the ability of the clusters to discover hidden patterns.
#
# A simplified scientific approach to clustering would first require you to quantify object characteristics with numerical values. Next, calculate the distance between objects based on characteristics, utilizing a “similarity” metric that can be used to compare variables.
# | |
# |:--:|
# |<b>Fig. 1 - Object Characteristics</b>|
# **Euclidean distance** is a commonly used metric to measure distance between clusters. It can be calculated by utilizing the following equation: $$ d(x,y) = \sqrt \Sigma(x_{i} - y_{i})^{2} $$
#
# Euclidean distance is a special case of **Minkowski Distance.**
#
# Another case of Minkowski Distance is **Manhattan Distance.**
# | |
# |:--:|
# |<b>Fig. 2</b>|
# Assume that each side of the square is **one unit**.
#
# The distance between A and B is as follows:
#
# (Assume A at origin, and B at (6,6))
#
# **Euclidean Distance**: $$ \sqrt (6^{2} + 6^{2}) \approx 8.485 $$
#
# **Manhattan Distance**: $$ | 0 - 6 | + | 0 - 6 | = 12 $$
# ## Rescaling Data
# **Standardize**: subtract mean and divide by the standard deviation.
#
# $$ x_{new} = \frac{x - \mu}{\sigma} $$
#
# One way we can standardize data in Python is by using the StandardScaler() function.
#
# After partitioning your data set into a train and test set as discussed in Chapter X, the StandardScaler () function can be utilized by:
# ```
# from sklearn.preprocessing import StandardScaler
#
# scaler = StandardScaler()
# train_X=scaler.fit_transform(train_X)
# valid_X=scaler.transform(valid_X)
# ```
# **Normalize**: scale to [0,1] by subtracting minimum and dividing by range
#
# $$ x_{new} = \frac{x - x_{min}}{x_{max} - x_{min}} $$
#
# We can normalize our data and use imputation for any missing values in Python by using the MinMaxScaler():
# ```
# # Scale the data to be between 0 and 1 (default range)
# mms = MinMaxScaler()
# data_df_array = mms.fit_transform(data_df_sub) # results stored in a numpy array, not a dataframe.
#
# # IMPUTATION
# # initialize imputer, which uses mean substitution
# # does this by taking the mean of the values of the two nearest neighbors
# # n_neighbors: number of neighbors
# # weights: whether and how to weight values; we don't weight
# imputer = KNNImputer(n_neighbors=2, weights="uniform")
#
# # apply the imputer function to the array
# data_df_array=imputer.fit_transform(data_df_array)
#
# # convert the array back into a dataframe
# data_df_norm = pd.DataFrame(data_df_array, columns=data_df_sub.columns)
# ```
# What makes a good similarity metric?
#
# * Symmetry: d(x,y) = d(y,x)
# * Satisfy triangle inequality: d(x,y) <= d(x,z) + d(z,y)
# * **Can** distinguish between different objects: if d(x,y) != 0 then x != y
# * **Can’t** distinguish between identical objects: if d(x,y) = 0 then x = y
# ## Clustering Algorithm
#
# There are two popular algorithms for partitional clustering: k-means and k-medoids. Both aim to partition a dataset with n data objects into k clusters. This is accomplished by finding a partition of k clusters that optimizes the chosen partitioning criterion. The ideal solution to this would be finding the Global Optimum by exhaustively enumerating all partitions. For the purposes of this course, we will focus on k-means clustering.
# | |
# |:--:|
# |<b>Fig. 3 - Distances between data points and a pre-determined centroid.</b>|
# In k-means clustering, we arbitrarily choose k initial cluster centroids and cluster data objects around them. Then, for the current partition, we compute new cluster centroids. Those new centroids are used to create new clusters and data objects are assigned to clusters based on the nearest new centroid. This process is repeated until there is no change to the centroids.
#
# K-Means Clustering Steps:
#
# 1. **Randomly** cluster objects into k number of clusters around k number of centroids
# 2. For the current partition, compute new cluster centroids
# 1. Centroids should be at the center (the mean value) of the cluster
# 3. Create new clusters, assigning each object to cluster with the nearest new centroid
# 4. Repeat steps 2 and 3 until there is no change to your centroids
# | |
# |:--:|
# |<b>Fig. 4 - K-Means Clustering Steps</b>|
# How do we choose a value for k?
#
# We can create an elbow plot. An elbow plot showcases the distortion score for each k value. The graph will show an inflection point - an “elbow” point - at the most ideal k-value.
# | |
# |:--:|
# |<b>Fig. 5 - Elbow Plot</b>|
# Note that we calculate average distortion scores utilizing the normalized data. In this case, distortion is the sum of mean Euclidean distances between data points and the centroids of their assigned clusters.
#
# Here is how we would go about creating an elbow plot for n = 15. This means we are looking at average distances for k = 1, 2, 3… 14, 15.
#
# ```
# # from scipy.cluster.vq import kmeans, vq
#
# # Declare variables for use
# distortions = []
# #How many clusters are you going to try? Specify the range
# num_clusters = range(1,16)
#
# # Create a list of distortions from the kmeans function
# for i in num_clusters:
# cluster_centers, distortion = kmeans(random_df_norm[['variable_of_interest_one', 'variable_of_interest_two']], i)
# distortions.append(distortion)
#
# # Create a data frame with two lists - number of clusters and distortions
# elbow_plot = pd.DataFrame({'num_clusters': num_clusters, 'distortions': distortions})
#
# # Create a line plot of num_clusters and distortions
# sns.lineplot(x='num_clusters', y='distortions', data=elbow_plot)
# plt.xticks(num_clusters)
# plt.show()
# ```
# ## Strengths + Weaknesses of k-means:
#
# The k-means method is efficient and much faster than hierarchical clustering. It is also straightforward and intuitively implementable.
#
# Some weaknesses of the k-means method are that we need to specify the value of k before running the algorithm which directly affects the final outcome. Also, this method is very sensitive to outliers. K-medoids clustering can address this (where the centroid could be a data point itself). Lastly, the k-means method is not very helpful for categorical data and only applies when a mean and centroid values are defined.
# ## References:
# ## Glossary:
#
# **Unsupervised Segmentation:**
#
# **Data pre-processing:**
#
# **Minkowski Distance:**
#
# **Euclidean Distance:**
#
# **Manhattan Distance:**
#
# **Standardize:**
#
# **Normalize:**
#
# **Global Optimum:**
#
# **K-Means Clustering:**
#
# **Elbow Plot:**
#
# **Hierarchical Clustering:**
| _build/jupyter_execute/Similarity & Clustering Chapter 4 Draft v2.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.2
# language: julia
# name: julia-1.4
# ---
# ### Cómo podemos hacer para distribuir la carga?
# La topología que nosotros elegimos es de un nodo coordinador y 4 workers.
# 
# Julia se comunica con los workers mediante una comunicación por SSH.
using Distributed
workervec = [("montecarlo@worker_1:22",1),
("montecarlo@worker_2:22",1),
("montecarlo@worker_3:22",1),
("montecarlo@worker_4:22",1)]
addprocs(workervec; tunnel=true)
println("Number procs: $(nprocs())")
println("Number of workers: $(nworkers())")
addprocs(2)
println("Number procs: $(nprocs())")
println("Number of workers: $(nworkers())")
| workdir/procesamiento_distribuido_en_julia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Serving
from __future__ import print_function
from PIL import Image
from grpc.beta import implementations
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow_serving.apis import predict_pb2
from tensorflow_serving.apis import prediction_service_pb2
import requests
import numpy as np
from StringIO import StringIO
server = 'localhost:8500'
host, port = server.split(':')
# define the image url to be sent to the model for prediction
image_url = "https://www.publicdomainpictures.net/pictures/60000/nahled/bird-1382034603Euc.jpg"
response = requests.get(image_url)
image = np.array(Image.open(StringIO(response.content)))
height = image.shape[0]
width = image.shape[1]
print("Image shape:", image.shape)
plt.imshow(image)
plt.show()
# create the request object and set the name and signature_name params
request = predict_pb2.PredictRequest()
request.model_spec.name = 'deeplab'
request.model_spec.signature_name = 'predict_images'
# +
# fill in the request object with the necessary data
request.inputs['images'].CopyFrom(
tf.contrib.util.make_tensor_proto(image.astype(dtype=np.float32), shape=[1, height, width, 3]))
request.inputs['height'].CopyFrom(tf.contrib.util.make_tensor_proto(height, shape=[1]))
request.inputs['width'].CopyFrom(tf.contrib.util.make_tensor_proto(width, shape=[1]))
# -
# create the RPC stub
channel = implementations.insecure_channel(host, int(port))
stub = prediction_service_pb2.beta_create_PredictionService_stub(channel)
# +
# sync requests
result_future = stub.Predict(request, 30.)
# For async requests
# result_future = stub.Predict.future(request, 10.)
# result_future = result_future.result()
# +
# get the results
output = np.array(result_future.outputs['segmentation_map'].int64_val)
height = result_future.outputs['segmentation_map'].tensor_shape.dim[1].size
width = result_future.outputs['segmentation_map'].tensor_shape.dim[2].size
image_mask = np.reshape(output, (height, width))
plt.imshow(image_mask)
plt.show()
# -
plt.figure(figsize=(14,10))
plt.subplot(1,2,1)
plt.imshow(image, 'gray', interpolation='none')
plt.subplot(1,2,2)
plt.imshow(image, 'gray', interpolation='none')
plt.imshow(image_mask, 'jet', interpolation='none', alpha=0.7)
plt.show()
| serving/deeplab_client.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="aPIA-10zdv4P"
# ## ART Randomized Smoothing
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="CGDOyI0HgDfx" outputId="2d61711f-6f8a-41b5-f05c-1085fd00fa13"
import keras.backend as k
from keras.models import load_model
from keras.models import Sequential
from keras.layers import Dense, Flatten, Conv2D, MaxPooling2D, Dropout
from art import DATA_PATH
from art.defences import GaussianAugmentation
from art.attacks import FastGradientMethod
from art.classifiers import KerasClassifier
from art.utils import load_dataset, get_file, compute_accuracy
from art.wrappers import RandomizedSmoothing
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# + [markdown] colab_type="text" id="FqXvuMM9dv4U"
# ### Load data
# + colab={} colab_type="code" id="z9OztmSidv4V"
# Read MNIST dataset
(x_train, y_train), (x_test, y_test), min_, max_ = load_dataset(str('mnist'))
num_samples_test = 250
x_test = x_test[0:num_samples_test]
y_test = y_test[0:num_samples_test]
# + [markdown] colab_type="text" id="xDCzquK1dv4X"
# ### Train classifiers
# + colab={} colab_type="code" id="G-mh9wSAHm-Z"
# create Keras convolutional neural network - basic architecture
def cnn_mnist(input_shape, min_val, max_val):
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3), activation='relu', input_shape=input_shape))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dense(10, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
classifier = KerasClassifier(clip_values=(min_val, max_val),
model=model, use_logits=False)
return classifier
# + colab={} colab_type="code" id="tGbe8Cjmdv4a"
num_epochs = 3
# # Construct and train a convolutional neural network
# classifier = cnn_mnist(x_train.shape[1:], min_, max_)
# classifier.fit(x_train, y_train, nb_epochs=num_epochs, batch_size=128)
# import trained model to save time :)
path = get_file('mnist_cnn_original.h5', extract=False, path=DATA_PATH,
url='https://www.dropbox.com/s/p2nyzne9chcerid/mnist_cnn_original.h5?dl=1')
classifier_model = load_model(path)
classifier = KerasClassifier(clip_values=(min_, max_), model=classifier_model,
use_logits=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="qH0lH14Ddv4d" outputId="058fe69b-70af-46ad-8d71-bc7f923f2202"
# add Gaussian noise and train two classifiers
sigma1 = 0.25
sigma2 = 0.5
# sigma = 0.25
ga = GaussianAugmentation(sigma=sigma1, augmentation=False)
x_new1, _ = ga(x_train)
classifier_ga1 = cnn_mnist(x_train.shape[1:], min_, max_)
classifier_ga1.fit(x_new1, y_train, nb_epochs=num_epochs, batch_size=128)
# sigma = 0.5
ga = GaussianAugmentation(sigma=sigma2, augmentation=False)
x_new2, _ = ga(x_train)
classifier_ga2 = cnn_mnist(x_train.shape[1:], min_, max_)
classifier_ga2.fit(x_new2, y_train, nb_epochs=num_epochs, batch_size=128)
# + colab={} colab_type="code" id="XYJN3rCpdv4h"
# create smoothed classifiers
classifier_rs = RandomizedSmoothing(classifier, sample_size=100, scale=0.25, alpha=0.001)
classifier_rs1 = RandomizedSmoothing(classifier_ga1, sample_size=100, scale=sigma1, alpha=0.001)
classifier_rs2 = RandomizedSmoothing(classifier_ga2, sample_size=100, scale=sigma2, alpha=0.001)
# + [markdown] colab_type="text" id="kukXRDcedv4j"
# ### Prediction
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="jcPkXptcdv4k" outputId="ee65b562-0839-483b-9b1f-b7c911e3131a"
# compare prediction of randomized smoothed models to original model f
x_preds = classifier.predict(x_test)
x_preds_rs1 = classifier_rs1.predict(x_test)
x_preds_rs2 = classifier_rs2.predict(x_test)
acc, cov = compute_accuracy(x_preds, y_test)
acc_rs1, cov_rs1 = compute_accuracy(x_preds_rs1, y_test)
acc_rs2, cov_rs2 = compute_accuracy(x_preds_rs2, y_test)
print("Original test data (first 250 images):")
print("Original Classifier")
print("Accuracy: {}".format(acc))
print("Coverage: {}".format(cov))
print("Smoothed Classifier, sigma=" + str(sigma1))
print("Accuracy: {}".format(acc_rs1))
print("Coverage: {}".format(cov_rs1))
print("Smoothed Classifier, sigma=" + str(sigma2))
print("Accuracy: {}".format(acc_rs2))
print("Coverage: {}".format(cov_rs2))
# + [markdown] colab_type="text" id="hqea3xvMdv4n"
# ### Certification accuracy and radius
# + colab={} colab_type="code" id="D6Va8ST8dv4n"
# calculate certification accuracy for a given radius
def getCertAcc(radius, pred, y_test):
rad_list = np.linspace(0,2.25,201)
cert_acc = []
num_cert = len(np.where(radius > 0)[0])
for r in rad_list:
rad_idx = np.where(radius > r)[0]
y_test_subset = y_test[rad_idx]
cert_acc.append(np.sum(pred[rad_idx] == np.argmax(y_test_subset, axis=1))/num_cert)
return cert_acc
# + colab={} colab_type="code" id="iPWY6KFMdv4p"
# compute certification
pred0, radius0 = classifier_rs.certify(x_test, n=500)
pred1, radius1 = classifier_rs1.certify(x_test, n=500)
pred2, radius2 = classifier_rs2.certify(x_test, n=500)
# + colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" id="ZZv5wDHSdv4s" outputId="a6fbe7ba-dfbb-47bd-8e56-794fb689cb14"
# plot certification accuracy wrt to radius
rad_list = np.linspace(0,2.25,201)
plt.plot(rad_list, getCertAcc(radius0, pred0, y_test), 'r-', label='original')
plt.plot(rad_list, getCertAcc(radius1, pred1, y_test), '-', color='cornflowerblue', label='smoothed, $\sigma=$' + str(sigma1))
plt.plot(rad_list, getCertAcc(radius2, pred2, y_test), '-', color='royalblue', label='smoothed, $\sigma=$' + str(sigma2))
plt.xlabel('radius')
plt.ylabel('certified accuracy')
plt.legend()
plt.show()
| notebooks/output_randomized_smoothing_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Note:** I tried several times to use PySpark to use `Logistic Regression` procedure, but most of times I got stuck on its processing. Therefore, I did a research and learn how to do that using SKLearn instead. Fortunately, I got better results using it rather than PySpark Framework for this purpose.
# +
import pandas as pd
from pandas.core.frame import DataFrame
from pandas.core.series import Series
import string
import nltk
from nltk.corpus import stopwords
from numpy import ndarray
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sklearn.utils import shuffle
# -
nltk.download('stopwords', quiet=True)
# Number of entries per dataframe
NUMBER_ENTRIES_PER_DF = 100
# +
def get_subsets(df:DataFrame, subset:str)->type(list):
return df.drop_duplicates(subset=[subset])[subset].to_list()
def get_accuracy(s:Series, arr:ndarray)->type(int):
return round(accuracy_score(s, arr) * 100, 2)
def get_random_df(df:DataFrame)->type(DataFrame):
n_df = shuffle(df.reset_index(drop=True))\
.head(NUMBER_ENTRIES_PER_DF)\
.reset_index(drop=True)
n_df.info()
return n_df
# -
t_df = pd.read_csv('../Datasets/True.csv')
f_df = pd.read_csv('../Datasets/Fake.csv')
t_df['label'] = 'Real News'
f_df['label'] = 'Fake News'
print("Real News - DF info:")
t_df = get_random_df(t_df)
print("Fake News - DF info:")
f_df = get_random_df(f_df)
df = shuffle(pd\
.concat([t_df, f_df])\
.reset_index(drop=True))
df = df.reset_index(drop=True)
df.drop(['date'], axis=1, inplace=True)
df.info()
df['text'] = df['text']\
.map(lambda x : x.lower()\
.translate(str\
.maketrans('', '', string.punctuation))
.join([word for word in x.split() if word not in stopwords.words('english')]))
print("Subjects: {}".format(get_subsets(df, 'subject')))
df.groupby(['subject'])['label'].count()
X_training, X_testing, y_training, y_testing = train_test_split(
df['text'],
df['label'],
test_size=0.3
)
ml_pipeline = Pipeline([
('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('model', LogisticRegression())
])
ml_model = ml_pipeline.fit(X_training, y_training)
ml_preds = ml_model.predict(X_testing)
print("Prediction accuracy: {}%".format(get_accuracy(y_testing, ml_preds)))
| PyDM.Module4/project2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import matplotlib
import seaborn as sns
import matplotlib.pyplot as plt
import tensorflow as tf
import numpy as np
from tensorflow.contrib import signal
import os
import sys
import re
import config as cfg
from dataload import load_test_batch, load_data, load_batch, load_config
from preprocessing import signalProcessBatch, tf_diff_axis
from models import buildModel
# +
# Tensorflow setup
sess = None
tf.logging.set_verbosity(tf.logging.INFO)
def reset_vars():
"""Initializes all tf variables"""
sess.run(tf.global_variables_initializer())
def reset_tf():
"""Closes the current tf session and opens new session"""
global sess
if sess:
sess.close()
tf.reset_default_graph()
sess = tf.Session()
# -
# ## Load some test files
reset_tf()
X_data_values, X_filenames = load_test_batch(cfg.DATA_DIR, idx=10, batch_size=2, samples=cfg.SAMRATE)
X_data = tf.placeholder(tf.float32, [None, cfg.SAMRATE], name='X_data')
x_mfcc, x_mel, x_zcr, x_rmse = signalProcessBatch(X_data,
noise_factor=0.0,
noise_frac=0.0,
window=512,
maxamps=cfg.MAXAMPS, sr=cfg.SAMRATE,
num_mel_bins=64,
num_mfccs=20)
x_mfcc_val, x_mel_val, x_zcr_val, x_rmse_val = sess.run(
[x_mfcc, x_mel, x_zcr, x_rmse],
feed_dict={X_data: X_data_values})
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.pcolormesh(x_mfcc_val[0].T)
plt.subplot(122)
plt.pcolormesh(x_mfcc_val[1].T);
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.pcolormesh(x_mel_val[0].T)
plt.subplot(122)
plt.pcolormesh(x_mel_val[1].T);
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.plot(x_zcr_val[0])
plt.subplot(122)
plt.plot(x_zcr_val[1]);
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.plot(x_rmse_val[0])
plt.subplot(122)
plt.plot(x_rmse_val[1]);
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.plot(x_mfcc_val[0].T[0])
plt.subplot(122)
plt.plot(x_mfcc_val[1].T[0]);
# Deltas
paddings = tf.constant([[0, 0], [0, 1]])
zcr_delta = sess.run(tf.pad(tf_diff_axis(x_zcr_val), paddings, 'CONSTANT'))
rmse_delta = sess.run(tf.pad(tf_diff_axis(x_rmse_val), paddings, 'CONSTANT'))
print zcr_delta.shape
print x_zcr_val.shape
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.plot(zcr_delta[0])
plt.subplot(122)
plt.plot(zcr_delta[1]);
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.plot(rmse_delta[0])
plt.subplot(122)
plt.plot(rmse_delta[1]);
# +
# Reshape to [audio file number, time size, 1]
x_zcr_val_rs = tf.reshape(x_zcr_val, [-1, 122, 1])
zcr_delta_rs = tf.reshape(zcr_delta, [-1, 122, 1])
x_rmse_val_rs = tf.reshape(x_rmse_val, [-1, 122, 1])
rmse_delta_rs = tf.reshape(rmse_delta, [-1, 122, 1])
# Stack together zcr and rmse features using tf.concat
zr_stack = tf.concat([x_zcr_val_rs, zcr_delta_rs, x_rmse_val_rs, rmse_delta_rs], 2)
# Stack with the mfccs using tf.concat
fingerprint = tf.concat([zr_stack, x_mfcc_val], 2)
# -
# Normalize
fingerprint_norm = sess.run(tf.nn.l2_normalize(fingerprint, 1, epsilon=1e-8))
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.pcolormesh(fingerprint_norm[0].T)
plt.subplot(122)
plt.pcolormesh(fingerprint_norm[1].T);
| exploratory_notebooks/tf_full_test_on_audio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Elasticity in 2D
#
# ## Introduction
#
# This example provides a demonstration of using PyMKS to compute the linear strain field for a two-phase composite material. The example introduces the governing equations of linear elasticity, along with the boundary conditions required for the MKS. It subsequently demonstrates how to generate data for delta microstructures and then use this data to calibrate the first order MKS influence coefficients for all strain fields. The calibrated influence coefficients are used to predict the strain response for a random microstructure and the results are compared with those from finite element. Finally, the influence coefficients are scaled up and the MKS results are again compared with the finite element data for a large problem.
#
# PyMKS uses the finite element tool [SfePy](http://sfepy.org) to generate both the strain fields to fit the MKS model and the verification data to evaluate the MKS model's accuracy.
# ### Elastostatics Equations
#
# For the sake of completeness, a description of the equations of linear elasticity is included. The constitutive equation that describes the linear elastic phenomena is Hook's law.
#
# $$ \sigma_{ij} = C_{ijkl}\varepsilon_{kl} $$
#
# $\sigma$ is the stress, $\varepsilon$ is the strain, and $C$ is the stiffness tensor that relates the stress to the strain fields. For an isotropic material the stiffness tensor can be represented by lower dimension terms which can relate the stress and the strain as follows.
#
# $$ \sigma_{ij} = \lambda \delta_{ij} \varepsilon_{kk} + 2\mu \varepsilon_{ij} $$
#
# $\lambda$ and $\mu$ are the first and second Lame parameters and can be defined in terms of the Young's modulus $E$ and Poisson's ratio $\nu$ in 2D.
#
# $$ \lambda = \frac{E\nu}{(1-\nu)(1-2\nu)} $$
#
# $$ \mu = \frac{E}{3(1+\nu)} $$
#
#
# Linear strain is related to displacement using the following equation.
#
# $$ \varepsilon_{ij} = \frac{u_{i,j}+u_{j,i}}{2} $$
#
# We can get an equation that relates displacement and stress by plugging the equation above back into our expression for stress.
#
# $$ \sigma_{ij} = \lambda u_{k,k} + \mu( u_{i,j}+u_{j,i}) $$
#
# The equilibrium equation for elastostatics is defined as
#
# $$ \sigma_{ij,j} = 0 $$
#
# and can be cast in terms of displacement.
#
# $$ \mu u_{i,jj}+(\mu + \lambda)u_{j,ij}=0 $$
#
# In this example, a displacement controlled simulation is used to calculate the strain. The domain is a square box of side $L$ which has an macroscopic strain $\bar{\varepsilon}_{xx}$ imposed.
#
# In general, generating the calibration data for the MKS requires boundary conditions that are both periodic and displaced, which are quite unusual boundary conditions and are given by:
#
# $$ u(L, y) = u(0, y) + L\bar{\varepsilon}_{xx}$$
# $$ u(0, L) = u(0, 0) = 0 $$
# $$ u(x, 0) = u(x, L) $$
# ## Modeling with MKS
#
# ### Calibration Data and Delta Microstructures
#
# The first order MKS influence coefficients are all that is needed to compute a strain field of a random microstructure, as long as the ratio between the elastic moduli (also known as the contrast) is less than 1.5. If this condition is met, we can expect a mean absolute error of 2% or less, when comparing the MKS results with those computed using finite element methods [[1]](#References).
#
# Because we are using distinct phases and the contrast is low enough to only need the first order coefficients, delta microstructures and their strain fields are all that we need to calibrate the first order influence coefficients [[2]](#References).
# +
from sklearn.pipeline import Pipeline
import dask.array as da
import numpy as np
from pymks import (
generate_delta,
plot_microstructures,
solve_fe,
PrimitiveTransformer,
LocalizationRegressor,
coeff_to_real
)
# +
#PYTEST_VALIDATE_IGNORE_OUTPUT
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
# Here we use the `generate_delta` function to create the two delta microstructures needed to calibrate the first order influence coefficients.
x_delta = generate_delta(n_phases=2, shape=(21, 21)).persist()
plot_microstructures(x_delta[0], x_delta[1], titles=("X[0]", "X[1]"), cmap='gray')
# Using delta microstructures for the calibration of the first order influence coefficients is essentially the same as using a unit [impulse response](http://en.wikipedia.org/wiki/Impulse_response) to find the kernel of a system in signal processing. Any given delta microstructure is composed of only two phases with the center cell having an alternative phase from the remainder of the domain.
# ### Generating Calibration Data
#
# The `solve_fe` function provides an interface to generate strain fields, which can then be used for calibration of the influence coefficients.
#
# This example uses a microstructure with elastic moduli values of 100 and 120 and Poisson's ratio values of 0.3 and 0.3, respectively. The macroscopic imposed strain equal to 0.02. These parameters must be passed into the `solve_fe` function.
# +
strain_xx = lambda x: solve_fe(
x,
elastic_modulus=(100, 120),
poissons_ratio=(0.3, 0.3),
macro_strain=0.02
)['strain'][...,0]
y_delta = strain_xx(x_delta).persist()
# -
# Observe the strain fields.
plot_microstructures(
y_delta[0],
y_delta[1],
titles=(r'$\mathbf{\varepsilon_{xx}}$ [0]', r'$\mathbf{\varepsilon_{xx}}$ [1]')
)
# ### Calibrating First Order Influence Coefficients
#
# The following creates a model using an Scikit-learn pipeline using the `PrimitiveTransformer` to discretize and the `LocalizationRegressor` to perform regression in Fourier space. `n_state` is set to 2 as there are 2 states.
model = Pipeline(steps=[
('discretize', PrimitiveTransformer(n_state=2, min_=0.0, max_=1.0)),
('regressor', LocalizationRegressor())
])
# The delta microstructures are used to calibrate the influence coefficients.
model.fit(x_delta, y_delta);
# The influence coefficient have been calibrated. The influence coefficients need to be converted into real space to view. A helper function, `to_real`, is used to get the real space coefficients from the model.
# +
to_real = lambda x: coeff_to_real(x.steps[1][1].coeff).real
coeff = to_real(model)
plot_microstructures(coeff[...,0], coeff[...,1], titles=['Influence coeff [0]', 'Influence coeff [1]'])
# -
# The influence coefficients have a Gaussian-like shape.
# ### Predict the Strain Field for a Random Microstructure
#
# Let's use the calibrated `model` to compute the strain field for a random two phase microstructure and compare it with the results from a finite element simulation. The `strain_xx` helper function is used to generate the strain field.
# +
da.random.seed(99)
x_data = da.random.randint(2, size=(1,) + x_delta.shape[1:]).persist()
y_data = strain_xx(x_data).persist()
# -
plot_microstructures(y_data[0], titles=[r'$\mathbf{\varepsilon_{xx}}$'])
# **Note that the calibrated influence coefficients can only be used to reproduce the simulation with the same boundary conditions that they were calibrated with.**
#
# Get the predicted strain field using `model.predict`.
y_predict = model.predict(x_data)
# Finally, compare the results from finite element simulation and the MKS model.
plot_microstructures(y_data[0], y_predict[0], titles=['Actual', 'Predicted'])
# Lastly, observe the difference between the two strain fields.
plot_microstructures(y_data[0] - y_predict[0], titles=['Finite Element - MKS'])
# The MKS model is able to capture the strain field for the random microstructure after being calibrated with delta microstructures.
# ## Resizing the Coefficients to use on Larger Microstructures
#
# The influence coefficients that were calibrated on a smaller microstructure can be used to predict the strain field on a larger microstructure though spectral interpolation [[3]](#References), but accuracy of the MKS model drops slightly. To demonstrate how this is done, generate a new random microstructure that is 3x larger
new_shape = tuple(np.array(x_delta.shape[1:]) * 3)
x_large = da.random.randint(2, size=(1,) + new_shape).persist()
y_large = strain_xx(x_large).persist()
plot_microstructures(y_large[0], titles=[r'$\mathbf{\varepsilon_{xx}}$'])
# The influence coefficients that have already been calibrated need to be resized to match the shape of the new larger microstructure that we want to compute the strain field for. This can be done by passing the shape of the new larger microstructure into the `resize_coeff` method.
model.steps[1][1].coeff_resize(x_large[0].shape);
# Observe the resized influence coefficients.
coeff = to_real(model)
plot_microstructures(coeff[...,0], coeff[...,1], titles=['Influence coeff [0]', 'Influence coeff [1]'])
# The coefficients have been resized so will only work on the 63x63 microstructures. As before, pass the microstructure as the argument to the `predict` method to get the strain field.
y_predict_large = model.predict(x_large).persist()
plot_microstructures(y_large[0], y_predict_large[0], titles=['Actual', 'Predicted'])
# Observe the difference between the two strain fields.
plot_microstructures(y_large[0] - y_predict_large[0], titles=['Finite Element - MKS'])
# The results from the strain field computed with the resized influence coefficients are not as accurate, but still acceptable for engineering purposes. This decrease in accuracy is expected when using spectral interpolation [[4]](#References).
#
# ## References
#
# <a id="ref1"></a>
# [1] <NAME>., <NAME>., <NAME>., A new spectral framework for establishing localization relationships for elastic behavior of composites and their calibration to finite-element models. Acta Materialia, 2008. 56 (10) p. 2272-2282 [doi:10.1016/j.actamat.2008.01.017](http://dx.doi.org/10.1016/j.actamat.2008.01.017).
#
# <a id="ref2"></a>
# [2] <NAME>., S.R. Niezgoda, S.R. Kalidindi, Multi-scale modeling of elastic response of three-dimensional voxel-based microstructure datasets using novel DFT-based knowledge systems. Acta Materialia, 2009. 58 (7): p. 2716-2725 [doi:10.1016/j.actamat.2010.01.007](http://dx.doi.org/10.1016/j.actamat.2010.01.007).
#
# <a id="ref3"></a>
# [3] <NAME>., <NAME>., <NAME>., Computationally efficient database and spectral interpolation for fully plastic Taylor-type crystal plasticity calculations of face-centered cubic polycrystals. International Journal of Plasticity 24 (2008) 1264–1276 [doi:10.1016/j.ijplas.2007.12.002](http://dx.doi.org/10.1016/j.ijplas.2007.12.002).
#
# <a id="ref4"></a>
# [4] <NAME>-<NAME>. , <NAME>., Crystal plasticity simulations using discrete Fourier transforms. Acta Materialia 57 (2009) 1777–1784 [doi:10.1016/j.actamat.2008.12.017](http://dx.doi.org/10.1016/j.actamat.2008.12.017).
| notebooks/elasticity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/danilo-uea/RNA-2021.1-AA1.2/blob/main/RNA_AA_1_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="1Ok_1jxEk3L6"
# # Redes Neurais Artificiais - Atividade Avaliativa 1.2
# > Criado por: <NAME> & <NAME>
#
#
#
# + id="a03rnx05qRD7"
import pandas as pd
import numpy as np
# + [markdown] id="6rojGRaUorE3"
# ---
# ## Ocorrências:
# O DataSet tem no total 243.984 casos.
#
# ## Atributos
# Os atributos deste DataSet são:
#
# * Data em que houve alguma evolução;
# * Data em que ocorreu a notificação;
# * Data em que foram notados os sintomas;
# * Classificação (resultado dos exames);
# * Conclusão (estado final do paciente);
# * Origem dos exames;
# * Existencia de comorbidade cardiaca;
# * Existencia de comorbidade cromossomica;
# * Existencia de comorbidade diabetes;
# * Existencia de comorbidade hemato;
# * Existencia de comorbidade imunologica;
# * Existencia de comorbidade neurologica;
# * Existencia de comorbidade obessidade;
# * Existencia de comorbidade renal;
# * Existencia de comorbidade respiratoria;
# * Criterio de avaliação dos exames;
# * Etnia do indivíduo;
# * Evoluçao do seu quadro clínico;
# * Idade do indivíduo;
# * Faixa etária do indivíduo;
# * Se o indivíduo é uma gestante;
# * Se o indivíduo é um profissional da saúde;
# * Raça do indivíduo;
# * se notificação;
# * Sexo do indivíduo;
# * Existencia do sintoma de dispneia;
# * Existencia do sintoma de febre;
# * Existencia de sintomas na garganta;
# * Existencia do sintoma de tosse;
# * Existencia de outros sintomas;
# * Tipo do SRAG;
# * Número de testes de anticorpos realizados;
# * Número de testes de antigeno realizados;
# * Número de testes PCR realizados;
# * Bairro em que o indivíduo reside;
# * Bairro em que o indivíduo vive baseando-se no mapa;
# * Distrito em que a pessoa reside;
# * Taxa de transmição;
# + id="tqVvc9x1g5yD"
# from google.colab import drive
# drive.mount('/content/drive')
# + id="fnzdyNd6n7TM"
# covid = pd.read_csv("/content/drive/My Drive/Colab Notebooks/RNA/Manaus.csv", sep = ";") # Yasser
covid = pd.read_csv("/content/drive/MyDrive/dados_colab/RNA/Manaus.csv", sep = ";", low_memory=False) # Danilo
# + id="H31UjfDhs7oT" colab={"base_uri": "https://localhost:8080/"} outputId="1403d323-7b6b-4123-e266-68df2cdae2fb"
print('O arquivo possui {} atributos no total:'.format(len(covid.columns)))
atributos = covid.columns.values
for i in range(len(atributos)):
print(atributos[i])
# + [markdown] id="J4edGNC8zCJ4"
# ---
# ## Limpeza dos dados
# Para esta atividade, iremos apenas usar os casos cuja a classificação sejam de casos confirmados, o DataSet apresenta no total 80420 casos confirmados. O primeiro caso confirmado neste DataSet foi em 16 de Fevereiro de 2020, e o ultimo caso confirmado foi em 29 de Dezembro de 2020.
#
# Para este trabalho, foram removidas colunas que não serão uteis para o escopo das atividades e os casos que apresentam algum dado faltante.
# Restaram apenas 9 colunas (_dt_notificacao, _classificacao, _conclusao, _idade, _sexo, _teste_anticorpo, _teste_antigeno, _teste_pcr e _bairro), e 246 casos.
#
#
#
# + id="rl6hzOwTol51"
covid = covid[covid["_classificacao"] == "Confirmado"]
# + id="gnn7bH1xtcQ7" colab={"base_uri": "https://localhost:8080/"} outputId="8130f3e2-f6f1-40c2-eb20-6ae3f7173d23"
print('Um total de {} casos confirmados'.format(len(covid)))
# + id="vjOrE9eTm7eD" colab={"base_uri": "https://localhost:8080/"} outputId="4998046d-85ae-4310-db82-9250015c27d0"
dates = covid['_dt_notificacao'].dropna() # Exclui linhas com elementos vazios ou nulos
dates = pd.to_datetime(dates, format='%d/%m/%Y %H:%M')
dates = dates.sort_values()
primeiro = dates.iloc[0]
ultimo = dates.iloc[-1]
print("Data do primeiro caso notificado: ")
print(primeiro)
print("Data do primeiro caso notificado: ")
print(ultimo)
# + id="yzTYppt4LNZq" colab={"base_uri": "https://localhost:8080/"} outputId="7aac9a1d-a4ef-4662-a634-0c67145c5fe9"
#removendo as colunas e as linhas com elementos nulos, sobraram apenas 240 linhas
if (len(covid.columns.values) == 39):
covid.drop(["_bairro_mapa","_taxa","_origem","_distrito","_faixa etária","_sintoma_outros","_sintoma_tosse","_sintoma_garganta","_sintoma_febre","_sintoma_dispneia","_srag","_se_notificacao","_evolução","_criterio","_gestante","_raca","_dt_sintomas","_dt_evolucao","_profiss_saude","_etnia","_dt_sintomas","_comorb_cardio","_comorb_cromossomica","_comorb_diabetes","_comorb_hemato","_comorb_hepatica","_comorb_imuno","_comorb_neurologica","_comorb_obessidade","_comorb_renal","_comorb_respiratoria"],axis=1,inplace=True)
#covid = covid.dropna(subset=['_idade'])
covid = covid.dropna() # remove todas as linhas com nop minimo um atributo nulo
atributos = covid.columns.values
for i in range(len(atributos)):
print(atributos[i])
# + [markdown] id="co-AV1LR1Tgs"
# ---
# ## Análise exploratória dos dados:
#
#
# 1) Quantos exemplos e atributos há na base de dados após a limpeza e organização?
#
#
# > A limpeza e a organização da base de dados resultou em um total de 246 exemplos, cada um com 9 atributos (_dt_notificacao, _classificacao, _conclusao, _idade, _sexo, _teste_anticorpo, _teste_antigeno, _teste_pcr e _bairro), nenhum atributo de nenhum exemplo na base de dados tem valor nulo.
#
#
# + id="1W7--gtK6RQe" colab={"base_uri": "https://localhost:8080/"} outputId="4bb149eb-1fc7-4afe-b4af-ac504b54e51b"
# 1.1
print('Qtd elementos: {}'.format(len(covid)))
print('Qtd atributos: {}'.format(len(covid.columns.values)))
# + [markdown] id="mY4ihmb29V5n"
# 2) Qual a porcentagem de indivíduos recuperados em relação ao todo?
# + id="ilksjf9ukfKi" colab={"base_uri": "https://localhost:8080/"} outputId="e1233182-1a03-41ea-f8fa-5528376bceea"
# 1.2
total_recuperados = len(covid[covid['_conclusao'] == 'Recuperado'])
print("Total de recuperados: " + str(total_recuperados))
print("Porcentagem: " + str(total_recuperados/len(covid)))
# + [markdown] id="ri9gfQuc8rVF"
# 3) Os casos acometeram mais indivíduos do sexo masculino ou feminino?
# + id="WcGjIDRbmuMw" colab={"base_uri": "https://localhost:8080/"} outputId="790bb753-6ea2-46f1-f9ac-0afac8eccb1e"
# 1.3
total_masculino = len(covid[covid['_sexo'] == 'M'])
total_feminino = len(covid[covid['_sexo'] == 'F'])
print("total de individuos masculinos que foram acometidos: " + str(total_masculino))
print("total de individuos femininos que foram acometidos: " + str(total_feminino))
# + [markdown] id="l9PVR9_f9hPF"
# 4) Qual a média e desvio padrão de idade dos indivíduos que contraíram COVID-19? Qual o
# indivíduo mais jovem e o mais idoso a contraírem tal enfermidade?
# + id="phvdA6oEnkE2" colab={"base_uri": "https://localhost:8080/"} outputId="56f76d1b-9ec4-49f0-e0da-b3ec81f7172a"
# 1.4
covid = covid.dropna(subset=['_idade']) # remove todas as linhas com nop minimo um atributo nulo
idades = covid['_idade'].sort_values()
mais_novo = idades.iloc[0]
mais_velho = idades.iloc[-1]
media = idades.mean()
desvio_padrao = idades.std()
if (mais_velho > 80):
mais_velho = 'Acima de 80 anos'
print("Idade do individuo mais novo: " + str(mais_novo))
print("Idade do individuo mais velho: " + str(mais_velho))
print("Media das idades: " + str(media))
print("Desvio padrao das idades: " + str(desvio_padrao))
# + [markdown] id="yU-Pw-3S9uPa"
# Devido a inconsistencias em relação aos nomes dos bairros devido a forma como foram inseridos, para padronizar os nomes dos bairros, iremos considerar que dois nomes se referem ao mesmo bairro se suas 5 primeiras letras forem iguais, e iremos remover as acentuações e considera-los em caixa alta, foi adicionado uma nova coluna na base de dados (_bairro_id) contendo o identificador do bairro de forma padronizada.
#
# 5) Qual o bairro com maior incidência de casos?
#
# 6) Quais os três bairros com maior incidência de casos recuperados?
# + id="ojFSvUO7pNKl" colab={"base_uri": "https://localhost:8080/"} outputId="523a11e9-e3a7-4b3a-9ae9-281fc599806b"
covid['_bairro_id'] = covid['_bairro'].str.extract(r'(^.{0,5})') #extrai os primeiros 5 caracteres e os adiciona em uma nova coluna em covid
covid['_bairro_id'] = covid['_bairro_id'].str.normalize('NFKD').str.encode('ascii', errors='ignore').str.decode('utf-8') # remove as acentuações
covid['_bairro_id'] = covid['_bairro_id'].str.upper() # torna todos os itens em caixa alta
covid['_bairro_id'].unique()
# + id="RyBJ_NkiCjba" colab={"base_uri": "https://localhost:8080/"} outputId="57ab31dc-b56c-4c0e-bda5-1f5a9f2933e6"
# 1.5 e 1.6
maior_freq = covid['_bairro_id'].mode()
recuperados = covid[covid['_conclusao'] == 'Recuperado']
freqs_recuperados = recuperados['_bairro_id'].value_counts()
print("Bairro com a maior frequencia: " + maior_freq[0])
print("\nOs 3 bairros com a maior incidencia de casos recuperados")
print(freqs_recuperados.index[0])
print(freqs_recuperados.index[1])
print(freqs_recuperados.index[2])
# + [markdown] id="Hll0bA9J_ktB"
# 7) Quais os tipos de testes efetuados, segundo os dados? Indique os dados de maneira quantitativa e percentual.
# + id="CpN6RxXIqSlc" colab={"base_uri": "https://localhost:8080/"} outputId="d296bb9c-91bc-4a31-de5f-4af3968387b8"
# 1.7
num_anticorpo = covid['_teste_anticorpo'].sum()
num_antigeno = covid['_teste_antigeno'].sum()
num_pcr = covid['_teste_pcr'].sum()
total = num_anticorpo + num_antigeno + num_pcr
print("Testes anticorpo\nTotal de aplicacoes: " + str(num_anticorpo) + "\nProporcao: " + str((num_anticorpo/total) * 100) + " %\n")
print("Testes antigeno\nTotal de aplicacoes: " + str(num_antigeno) + "\nProporcao: " + str((num_antigeno/total) * 100) + " %\n")
print("Testes pcr\nTotal de aplicacoes: " + str(num_pcr) + "\nProporcao: " + str((num_pcr/total) * 100) + " %\n")
# + [markdown] id="RBNNtny8_s0a"
# 8) Qual taxa de letalidade pode ser calculada a partir do conjunto de dados? Para calcular esta taxa, considere a fração do total de óbitos pelo total de casos.
# + id="dJOqszFwdK5x" colab={"base_uri": "https://localhost:8080/"} outputId="3d0db9c0-b58d-4fe0-e0e8-a7a0df3085a2"
#1.8
total_obitos = len(covid[covid['_conclusao'] != 'Recuperado'])
print("Total de obitos: " + str(total_obitos))
print("Porcentagem: " + str(total_obitos/len(covid)))
# + [markdown] id="i2VvvQUL_yrn"
# 9) Qual o tipo de correlação, mediante coeficiente de correlação de Pearson, entre a idade e o
# número de casos? Para responder a esta pergunte, agrupe o número de casos por idade e efetue
# o cálculo de tal coeficiente. Indique, a partir do resultado, a natureza desta correlação, se é
# positiva ou negativa, e qual sua intensidade.
# + id="btG8phiJrFwf" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="4a3571c3-4a29-4200-dbb8-0532e70d400a"
#1.9
idade_sort = covid.sort_values(by=['_idade'])
idade_sort.corr(method='pearson')
# + [markdown] id="tKLGQxJ__6-D"
# ---
# ## Visualização de dados:
#
#
# 1) Construa um histograma denotando a quantidade de casos nos 10 bairros em que houve mais casos registrados. Inclua todos os bairros remanescentes em uma categoria denominada “Outros.” Denote as informações de maneira percentual.
#
# + id="dOt0L7yoda_2" colab={"base_uri": "https://localhost:8080/", "height": 338} outputId="7de0c049-d32d-4983-e421-b292dbbe1921"
# 2.1
import matplotlib.pyplot as plt
nomes = []
valores = []
total_casos = len(covid)
soma = 0
ordem_freq = covid['_bairro_id'].value_counts()
for i in range(10):
valor = 100 * (ordem_freq[i]/total_casos)
soma = soma + valor
valores.append(valor)
nomes.append(ordem_freq.index[i])
valores.append(100 - soma)
nomes.append('OUTROS')
figura1, ax_1 = plt.subplots(figsize=(10, 5))
ax_1.bar(nomes, valores)
ax_1.set_ylabel('Quantidade de casos (%)', size=14)
ax_1.set_xlabel('Bairros', size=14)
figura1.show()
# + [markdown] id="OsHQx-szvOpm"
# 2) Denote, por sexo, o boxplot da idade dos casos confirmados. Há outliers?
#
# > Existe um outlier no sexo feminino
# + id="JnbkKUKMdeYu" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="0c4cade1-c087-41ca-f93f-79adad41ef88"
# 2.2
masculino = covid[covid["_sexo"] == "M"]
masculino = np.array(masculino['_idade'], dtype=object)
feminino = covid[covid["_sexo"] == "F"]
feminino = np.array(feminino['_idade'], dtype=object)
labels = ['Masculino', 'Feminino']
data = np.array([masculino, feminino], dtype=object)
figura_2, ax_2 = plt.subplots()
ax_2.set_title('Boxplot da idade por sexo')
ax_2.boxplot(data, labels=labels)
figura_2.show()
# + [markdown] id="5CGT_e0avS3D"
# 3) Denote em um gráfico de barras o número de novos casos por dia, considerando os 10 últimos dias existentes na base de dados.
# + id="kFui8L2adfX-" colab={"base_uri": "https://localhost:8080/", "height": 395} outputId="f668116a-9254-41f8-ca9d-3c9e0fe66ca3"
# 2.3
nomes = []
valores = []
dates = covid['_dt_notificacao'].dropna() # Exclui linhas com elementos vazios ou nulos
dates = pd.to_datetime(dates, format='%d/%m/%Y %H:%M')
data_qtd = dates.value_counts()
data_ordem = dates.sort_values(ascending = False)
data_ordem = data_ordem.unique()
for i in range(9, -1, -1):
valores.append(data_qtd[data_ordem[i]])
nomes.append(data_ordem[i])
figura3, ax_3 = plt.subplots(figsize=(8, 5))
ax_3.bar(nomes, valores)
ax_3.set(xlabel='Dias', ylabel='Quantidade', title='Novos casos nos últimos 10 dias')
plt.xticks(rotation=60)
figura3.show()
# + [markdown] id="8YuQV1thvY-d"
# 4) Repita o gráfico anterior considerando o número de casos recuperados.
# + id="XhEtfElNdhU9" colab={"base_uri": "https://localhost:8080/", "height": 395} outputId="d09da261-85c4-4695-98d0-2cdf2b96a6b6"
# 2.4
nomes = []
valores = []
recuperados = covid[covid['_conclusao'] == 'Recuperado']
dates = recuperados['_dt_notificacao'].dropna() # Exclui linhas com elementos vazios ou nulos
dates = pd.to_datetime(dates, format='%d/%m/%Y %H:%M')
data_qtd = dates.value_counts()
data_ordem = dates.sort_values(ascending = False)
data_ordem = data_ordem.unique()
for i in range(9, -1, -1):
valores.append(data_qtd[data_ordem[i]])
nomes.append(data_ordem[i])
figura3, ax_3 = plt.subplots(figsize=(8, 5))
ax_3.bar(nomes, valores)
ax_3.set(xlabel='Dias', ylabel='Quantidade', title='Casos recuperados últimos 10 dias')
plt.xticks(rotation=60)
figura3.show()
# + [markdown] id="62rjP1M1vdy8"
# 5) Construa um histograma que denote a quantidade percentual de casos por grupo etário, considerando que cada grupo contempla uma década (0 a 10 anos, 11 a 20 anos, etc.).
# + id="Ny2PrzlaeJQB" colab={"base_uri": "https://localhost:8080/", "height": 380} outputId="ff9d3500-88f2-4f26-91be-0d4879a5240b"
# 2.5
idades = covid["_idade"]
idades = idades.value_counts()
nomes = []
nomes.append('0 a 10')
nomes.append('11 a 20')
nomes.append('21 a 30')
nomes.append('31 a 40')
nomes.append('41 a 50')
nomes.append('51 a 60')
nomes.append('61 a 70')
nomes.append('71 a 80')
nomes.append('81 ou +')
valores = []
valores.append(0)
valores.append(0)
valores.append(0)
valores.append(0)
valores.append(0)
valores.append(0)
valores.append(0)
valores.append(0)
valores.append(0)
for i in range(len(idades)):
if (idades.index[i] >= 0 and idades.index[i] <= 10):
valores[0] = valores[0] + idades[idades.index[i]]
if (idades.index[i] >= 11 and idades.index[i] <= 20):
valores[1] = valores[1] + idades[idades.index[i]]
if (idades.index[i] >= 21 and idades.index[i] <= 30):
valores[2] = valores[2] + idades[idades.index[i]]
if (idades.index[i] >= 31 and idades.index[i] <= 40):
valores[3] = valores[3] + idades[idades.index[i]]
if (idades.index[i] >= 41 and idades.index[i] <= 50):
valores[4] = valores[4] + idades[idades.index[i]]
if (idades.index[i] >= 51 and idades.index[i] <= 60):
valores[5] = valores[5] + idades[idades.index[i]]
if (idades.index[i] >= 61 and idades.index[i] <= 70):
valores[6] = valores[6] + idades[idades.index[i]]
if (idades.index[i] >= 71 and idades.index[i] <= 80):
valores[7] = valores[7] + idades[idades.index[i]]
if (idades.index[i] >= 81):
valores[8] = valores[8] + idades[idades.index[i]]
total = len(covid)
for i in range(len(valores)):
valores[i] = (valores[i] * 100) / total
figura5, ax_5 = plt.subplots(figsize=(8, 5))
ax_5.bar(nomes, valores)
ax_5.set(xlabel='Grupo etário', ylabel='Quantidade percentual', title='Casos por grupo etário')
plt.xticks(rotation=60)
figura5.show()
# + [markdown] id="YzVFNcOMvlqY"
# 6) Elabore um gráfico que mostra o cumulativo de casos notificados ao longo do tempo.
# + id="T41P5tCjeTNv" colab={"base_uri": "https://localhost:8080/", "height": 381} outputId="4e8f8184-a1b7-49ab-8f89-188d388b29c0"
# 2.6
nomes = []
valores = []
dates = covid['_dt_notificacao'].dropna() # Exclui linhas com elementos vazios ou nulos
dates = pd.to_datetime(dates, format='%d/%m/%Y %H:%M')
data_qtd = dates.value_counts() # Extrai a quantidade de casos notificados por data
data_ordem = dates.sort_values(ascending = False) # Ordena as datas
figura6, ax_6 = plt.subplots(figsize=(8, 5))
ax_6.hist(data_ordem, 40, rwidth=0.9)
ax_6.set(xlabel='Dias', ylabel='Quantidade', title='Cumulativo de casos notificados ao longo do tempo')
plt.xticks(rotation=60)
figura6.show()
# + [markdown] id="dacEqp0Jv5ya"
# 7) Faça um gráfico do tipo scatterplot que denote a idade versus o número total de casos registrado para aquela idade. Aproveite o processamento efetuado para o cálculo da correlação. É possível observar alguma tendência?
#
#
# > Existe uma tendência de uma maior quantidade de casos nas idades entre 32 e 45 anos.
#
#
# + id="3ToMQPJW-r2i" colab={"base_uri": "https://localhost:8080/", "height": 350} outputId="f0de48a2-aefd-4463-cd06-519a55c02e6f"
# 2.7
dados = covid["_idade"]
dados = dados.value_counts() # A frequência de casos por idade
idade = []
frequencia = []
for i in range(len(dados)):
idade.append(dados.index[i])
frequencia.append(dados[dados.index[i]])
figura6, ax_6 = plt.subplots(figsize=(8, 5))
ax_6.scatter(idade, frequencia)
ax_6.set(xlabel='Idade', ylabel='Número de casos', title='Idade vs número de casos para esta idade')
figura6.show()
# + [markdown] id="OVQmjUBS6mr3"
# ---
# ## Tarefa de Classificação:
#
#
# > A tarefa de classificação proposta seria determinar com base nos sintomas, comorbidades e idade de um paciente se ele virá a necessitar de uma internação no hospital, ou em uma UTI, ou se ele pode realizar o tratamento em casa.
#
#
#
# * **Atributos Preditores:**
# 1. _idade;
# 2. _gestante;
# 3. _comorb_cardio;
# 4. _comorb_cromossomica;
# 5. _comorb_diabetes;
# 6. _comorb_hemato;
# 7. _comorb_hepatica;
# 8. _comorb_imuno;
# 9. _comorb_neurologica;
# 10. _comorb_obessidade;
# 11. _comorb_renal;
# 12. _comorb_respiratoria;
# 13. _sintoma_dispneia;
# 14. _sintoma_febre;
# 15. _sintoma_garganta;
# 16. _sintoma_tosse;
# 17. _sintoma_outros;
# 18. _srag;
#
#
#
# * **Atributo-Alvo:** _evolução (tratamento em casa, internação ou internação em UTI)
#
#
#
# * **Metricas de desempenho:** Como se trata de uma tarefa de classificar multi-classes, e como a quantidade de classes é desbalanceada, será necessario usar a tecnica de micro-averaging
#
# 1. Precisão, para determinar a taxa de acertos na classe positiva;
# 2. Revocação, para determinar a taxa de acertos entre os elementos considerados da classe positiva.
# 3. Especificidade, para determinar a taxa de acertos entre os elementos considerados na classe negativa
# 5. F-Score, a média harmonica
# 6. G-Score, pois a quantidade dos exemplos das classes é muito desbalanceadas
#
#
#
# + [markdown] id="fs8jr-ON6cMo"
# ---
# ## Tarefa de Regressão:
# A tarefa de regressão proposta seria prever a taxa de transmissão em um bairro com base nas datas de notificação, evolução e sintoma, Srag, classificação e no numero total de individuos por bairro.
#
#
# * **Atributos Preditores:**
# 1. _dt_evolucao;
# 2. _dt_notificacao;
# 3. _dt_sintomas;
# 4. _classificacao;
# 5. _srag;
# 6. _bairro;
# 7. _bairro_mapa;
# 8. _distrito;
#
#
#
# * **Atributo-Alvo:** _taxa
| RNA_AA_1_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Generating a model
#
# Activity: ADP-ribose-1″-phosphate phosphatase
#
# https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1563857/
#
# 2FAV. N41 catalytic. Mystery. No covalents?
#
# 1"P would clash. Alternate unknown form?
# +
pdbcode = '6WOJ' # ADP bound.
from fragmenstein import Victor, Igor
Igor.download_map(pdbcode, pdbcode+'.ccp4')
# structuure manually inspected. Chain A
# +
from rdkit_to_params import Params
p = Params.from_smiles_w_pdbfile(pdb_file='mono.pdb',
smiles='Nc1ncnc2n(cnc12)[C@@H]3O[C@H](CO[P@@]([O-])(=O)O[P@@]([O-])(=O)OC[C@H]4O[C@@H](O)[C@H](O)[C@@H]4O)[C@@H](O)[C@H]3O',
name='APR',
proximityBonding=False)
p.dump('APR.params')
# +
import pyrosetta
pyrosetta.init(extra_options='-no_optH false -mute all -ex1 -ex2 -ignore_unrecognized_res false -load_PDB_components false -ignore_waters false')
import nglview
nglview.show_rosetta(p.test())
# +
params_file = 'APR.params'
pdbfile = 'mono.pdb'
pose = pyrosetta.Pose()
params_paths = pyrosetta.rosetta.utility.vector1_string()
params_paths.extend([params_file])
pyrosetta.generate_nonstandard_residue_set(pose, params_paths)
pyrosetta.rosetta.core.import_pose.pose_from_file(pose, pdbfile)
Igor.relax_with_ED(pose=pose, ccp4_file=pdbcode+'.ccp4')
pose.dump_pdb('mono.r.pdb')
# -
# inspected and aligned to XChem in PyMOL...
| scripts/template_making.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %pylab
# %matplotlib inline
# +
import os
import glob
import multiprocessing
import itertools
import argparse
import numpy as np
import pandas as pd
import matplotlib.image as mpimg
from sklearn.cluster import DBSCAN
from subprocess import PIPE, Popen
import scipy.spatial
from scipy.optimize import curve_fit
import warnings
warnings.simplefilter(action='ignore', category=Warning)
import qrdar
import pcd_io
import ply_io
# +
# def apply_rotation(M, df):
# if 'a' not in df.columns:
# df.loc[:, 'a'] = 1
# r_ = np.dot(M, df[['x', 'y', 'z', 'a']].T).T
# df.loc[:, ['x', 'y', 'z']] = r_[:, :3]
# return df[['x', 'y', 'z']]
# def apply_rotation_2D(M, df):
# if 'a' not in df.columns:
# df.loc[:, 'a'] = 1
# r_ = np.dot(M, df[['x', 'y', 'a']].T).T
# df.loc[:, ['x', 'y']] = r_[:, :2]
# return df[['x', 'y']]
def rigid_transform_3D(A, B, d=3):
"""
http://nghiaho.com/uploads/code/rigid_transform_3D.py_
"""
assert len(A) == len(B)
A = np.matrixlib.defmatrix.matrix(A)
B = np.matrixlib.defmatrix.matrix(B)
N = A.shape[0]; # total points
centroid_A = mean(A, axis=0).reshape(1, d)
centroid_B = mean(B, axis=0).reshape(1, d)
# centre the points
AA = A - np.tile(centroid_A, (N, 1))
BB = B - np.tile(centroid_B, (N, 1))
# dot is matrix multiplication for array
H = transpose(AA) * BB
U, S, Vt = linalg.svd(H)
R = np.dot(Vt.T, U.T)
t = -R*centroid_A.T + centroid_B.T
M, N = np.identity(d+1), np.identity(d+1)
M[:d, :d] = R
N[:d, d] = t.reshape(-1, d)
return np.dot(N, M)
def read_aruco2(pc,
expected,
figs=False,
marker_template=None,
codes_dict='aruco_mip_16h3',
verbose=False):
if verbose: print ("extracting aruco")
pc.loc[:, 'intensity'] = pc.refl
targets = qrdar.identify_codes(pc,
expected=expected,
print_figure=True,
marker_template=marker_template,
codes_dict=codes_dict,
verbose=verbose)
targets.rename(columns={'code':'aruco'}, inplace=True)
targets = targets[targets.confidence == 1]
targets.reset_index(inplace=True)
sticker_centres = pd.DataFrame(columns=['x', 'y', 'z', 'aruco'])
i = 0
for ix, row in targets.iterrows():
for col in ['c0', 'c1', 'c2', 'c3']:
if isinstance(row[col], float): continue
sticker_centres.loc[i, :] = list(row[col]) + [row.aruco]
i += 1
return sticker_centres#[['aruco', 'x', 'y']]
def identify_ground2(pc, target_centres):
nominal_plane = target_centres[['x', 'y', 'z']].copy()
nominal_plane.z = 0
M = qrdar.common.rigid_transform_3D(target_centres[['x', 'y', 'z']].astype(float).values,
nominal_plane.astype(float).values)
pc.loc[:, ['x', 'y', 'z']] = qrdar.common.apply_rotation(M, pc)
pc.loc[pc.z < .05, 'is_branch'] = False
return pc, M
def find_buckets(pc, target_centres, N, bucket_height=.38, bucket_radius=.15):
"""
Returns: pc, bucket_centres
"""
### find buckets and remove ###
print ('finding buckets')
buckets = pc[pc.z.between(.1, .4)]
# voxelise to speed-up dbscan
buckets.loc[:, 'xx'] = (buckets.x // .005) * .005
buckets.loc[:, 'yy'] = (buckets.y // .005) * .005
buckets.loc[:, 'zz'] = (buckets.z // .005) * .005
buckets.sort_values(['xx', 'yy', 'zz', 'refl'], inplace=True)
bucket_voxels = buckets[~buckets[['xx', 'yy', 'zz']].duplicated()]
print(buckets)
dbscan = DBSCAN(min_samples=20, eps=.05).fit(bucket_voxels[['xx', 'yy', 'zz']])
bucket_voxels.loc[:, 'labels_'] = dbscan.labels_
# merge results back
buckets = pd.merge(buckets, bucket_voxels[['xx', 'yy', 'zz', 'labels_']], on=['xx', 'yy', 'zz'])
# find three largest targets (assumed buckets)
labels = buckets.labels_.value_counts().index[:N]
buckets = buckets[buckets.labels_.isin(labels)]
bucket_centres = buckets.groupby('labels_')[['x', 'y']].mean().reset_index()
bucket_centres.loc[:, 'aruco'] = -1
try:
# pair up aruco and buckets , identify and label bucket points
for i, lbl in enumerate(buckets.labels_.unique()):
bucket = buckets[buckets.labels_ == lbl]
X, Y = bucket[['x', 'y']].mean(), target_centres[['x', 'y']].astype(float)
dist2bucket = np.linalg.norm(X - Y, axis=1)
aruco = target_centres.loc[np.where(dist2bucket == dist2bucket.min())].aruco.values[0]
print ('bucket {} associated with aruco {}'.format(lbl, aruco))
bucket_centres.loc[bucket_centres.labels_ == lbl, 'aruco'] = aruco
# identify buckets points
x_shift = bucket_centres[bucket_centres.aruco == aruco].x.values
y_shift = bucket_centres[bucket_centres.aruco == aruco].y.values
pc.dist = np.sqrt((pc.x - x_shift)**2 + (pc.y - y_shift)**2)
idx = pc[(pc.z < bucket_height) & (pc.dist < bucket_radius) & (pc.is_branch)].index
pc.loc[idx, 'is_branch'] = False
# label branch base with aruco
idx = pc[(pc.z < bucket_height + .5) & (pc.dist < bucket_radius)].index
pc.loc[idx, 'aruco'] = aruco
except Exception as err:
plt.scatter(buckets.x.loc[::100], buckets.y.loc[::100], c=buckets.labels_.loc[::100])
plt.scatter(target_centres.x, target_centres.y)
[plt.text(r.x, r.y, r.aruco) for ix, r in target_centres.iterrows()]
raise Exception
return pc, bucket_centres
def isolate_branches(pc, N, translation, odir):
print ('\tsegmenting branches')
min_sample, iterate = 10, True
while iterate:
branches = pc[pc.is_branch]
branches.loc[:, 'xx'] = (branches.x // .005) * .005
branches.loc[:, 'yy'] = (branches.y // .005) * .005
branches.loc[:, 'zz'] = (branches.z // .005) * .005
branch_voxels = branches[~branches[['xx', 'yy', 'zz']].duplicated()]
dbscan = DBSCAN(min_samples=min_sample, eps=.02).fit(branch_voxels[['xx', 'yy', 'zz']])
branch_voxels.loc[:, 'labels_'] = dbscan.labels_
branches = pd.merge(branches, branch_voxels[['xx', 'yy', 'zz', 'labels_']], on=['xx', 'yy', 'zz'])
labels = branches.labels_.value_counts().index[:N]
branches = branches[branches.labels_.isin(labels)]
width = branches.groupby('labels_').agg({'x':np.ptp, 'y':np.ptp})
if np.any(width < .1):
min_sample += 10
else: iterate = False
cols = [u'pid', u'tot_rtn', u'x', u'y', u'z', u'dev', u'refl', u'rtn_N', u'sel', u'sp', u'rng', u'spot_size']
for i, label in enumerate(branches.labels_.unique()):
b = branches[branches.labels_ == label]
aruco = b[(b.z < .5) & (~np.isnan(b.aruco))].aruco.value_counts().index[0]
tag = translation[(translation.aruco == aruco)].tag.values[0]
b.loc[:, ['x', 'y', 'z']] = qrdar.common.apply_rotation(np.linalg.inv(M), b)
ply_io.write_ply(os.path.join(odir, '{}.ply'.format(tag)), b[cols])
print ('\tsaved branch to:', os.path.join(odir, '{}.ply'.format(tag)))
def read_pc(args):
pc = qrdar.io.read_ply(args.pc)
pc = pc[pc.dev <= 10]
pc.loc[:, 'is_branch'] = True
pc.loc[:, 'aruco'] = np.nan
if args.verbose: print ("number of points:", len(pc))
return pc
# +
class ARGS:
def __init__(self):
self.pc = '/data/TLS/uk/mk-street/branches_indoors/2021-07-11.003.riproject/2021-07-11-003.ply'
self.translation = '/data/TLS/uk/mk-street/branches_indoors/qrdar.csv'
self.odir = '/data/TLS/uk/mk-street/branches_indoors/branches/'
self.verbose = True
self.step = 2
self.bucket_height = 40
self.bucket_radius = 15
args = ARGS()
# +
# if __name__ == "__main__":
# parser = argparse.ArgumentParser()
# parser.add_argument('-p', '--pc', type=str, help='path to point cloud')
# parser.add_argument('-t', '--translation', type=str, help='path to .csv with tag translation,\
# this should have the form "name, project, code" \
# where name is the branch name, project is the name\
# of the file and code is the qrDAR number')
# parser.add_argument('-o', '--odir', type=str, help='output directory for branches')
# parser.add_argument('--bucket-height', type=float, default=.4, help='height of the bucket')
# parser.add_argument('--bucket-radius', type=float, default=.15, help='radius of the bucket')
# parser.add_argument('--verbose', action='store_true', help='print something')
# args = parser.parse_args()
# path = '2019-07-26.012.riproject/ascii/2019-07-26.012.ply'
project = os.path.split(args.pc)[1].split('.')[0]
if args.verbose: print ('processing project:', project)
# reading in translation will need to be edited
# dependent on formatting etc.
ctag = lambda row: '{}-{}-{}'.format(*row[['plot', 'treetag', 'light']])
translation = pd.read_csv(args.translation)
translation.rename(columns={c:c.lower() for c in translation.columns}, inplace=True)
#translation.loc[:, 'tag'] = translation.apply(ctag, axis=1)
#translation.tag = [t.replace('-nan', '') for t in translation.tag]
translation = translation[translation.project == project]
n_targets = len(translation[translation.project == project])
expected = translation[translation.project == project].code.astype(int).values
if args.verbose: print('expecting targets:', n_targets)
# read in branch scan
pc = read_pc(args)
### read aruco targets ###
sticker_centres = read_aruco2(pc, expected, verbose=args.verbose)
if args.verbose: print('targets identified')
### identify ground ###
pc, M = identify_ground2(pc, sticker_centres)
if args.verbose: print('ground identified')
# -
sticker_centres
# +
### find buckets ###
pc, buket_centres = find_buckets(pc, sticker_centres, n_targets,
bucket_height=args.bucket_height,
bucket_radius=args.bucket_radius)
if args.verbose: print('buckets found')
### isolate branches ###
isolate_branches(pc, n_targets, translation, args.odir)
# -
| python/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Magnitude of the Gradient
# 
# With the result of the last quiz, you can now take the gradient in x or y and set thresholds to identify pixels within a certain gradient range. If you play around with the thresholds a bit, you'll find the x-gradient does a cleaner job of picking up the lane lines, but you can see the lines in the y-gradient as well.
#
# In this next exercise, your goal is to apply a threshold to the overall magnitude of the gradient, in both x and y.
#
# The magnitude, or absolute value, of the gradient is just the square root of the squares of the individual x and y gradients. For a gradient in both the xx and yy directions, the magnitude is the square root of the sum of the squares.
#
# **abs_sobelx** = \sqrt{(sobel_x)^2}
#
# **abs_sobely** = \sqrt{(sobel_y)^2}
#
# **abs_sobelxy** = \sqrt{(sobel_x)^2+(sobel_y)^2}
# It's also worth considering the size of the region in the image over which you'll be taking the gradient. You can modify the kernel size for the Sobel operator to change the size of this region. Taking the gradient over larger regions can smooth over noisy intensity fluctuations on small scales. The default Sobel kernel size is 3, but here you'll define a new function that takes kernel size as a parameter.
#
# It's important to note here that the kernel size should be an **odd** number. Since we are searching for the gradient around a given pixel, we want to have an equal number of pixels in each direction of the region from this central pixel, leading to an odd-numbered filter size - a filter of size three has the central pixel with one additional pixel in each direction, while a filter of size five has an additional two pixels outward from the central pixel in each direction.
#
# The function you'll define for the exercise below should take in an image and optional Sobel kernel size, as well as thresholds for gradient magnitude. Next, you'll compute the gradient magnitude, apply a threshold, and create a binary output image showing where thresholds were met.
# **Steps** to take in this exercise:
#
# 1. Fill out the function in the editor below to return a thresholded gradient magnitude.
# 2. Again, you can apply exclusive (**<, >**) or inclusive (**<=, >=**) thresholds.
# 3. Test that your function returns output similar to the example below for **sobel_kernel=9, mag_thresh=(30, 100)**.
# +
import numpy as np
import cv2
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import pickle
# Read in an image
image = mpimg.imread('img/signs_vehicles_xygrad.png')
# Define a function that applies Sobel x and y,
# then computes the magnitude of the gradient
# and applies a threshold
def mag_thresh(img, sobel_kernel=3, mag_thresh=(0, 255)):
# Apply the following steps to img
# 1) Convert to grayscale
# 2) Take the gradient in x and y separately
# 3) Calculate the magnitude
# 4) Scale to 8-bit (0 - 255) and convert to type = np.uint8
# 5) Create a binary mask where mag thresholds are met
# 6) Return this mask as your binary_output image
# Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Take both Sobel x and y gradients
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# Calculate the gradient magnitude
gradmag = np.sqrt(sobelx**2 + sobely**2)
# Rescale to 8 bit
scale_factor = np.max(gradmag)/255
gradmag = (gradmag/scale_factor).astype(np.uint8)
# Create a binary image of ones where threshold is met, zeros otherwise
binary_output = np.zeros_like(gradmag)
binary_output[(gradmag >= mag_thresh[0]) & (gradmag <= mag_thresh[1])] = 1
#binary_output = np.copy(img) # Remove this line
return binary_output
# Run the function
mag_binary = mag_thresh(image, sobel_kernel=3, mag_thresh=(30, 100))
# Plot the result
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
f.tight_layout()
ax1.imshow(image)
ax1.set_title('Original Image', fontsize=50)
ax2.imshow(mag_binary, cmap='gray')
ax2.set_title('Thresholded Magnitude', fontsize=50)
plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
plt.show()
# -
| 04_Gradients_Color_Spaces/4_4_Magnitude_of_Gradient.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import database
from snowballing.operations import load_work, reload, work_by_varname
from snowballing.snowballing import Converter
from snowballing.snowballing import BackwardSnowballing
from snowballing.dbmanager import insert, set_attribute
Converter().browser()
len(article_list)
BackwardSnowballing("murta2014a", articles=article_list)
| snowballing/bibtex/Backward.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import adafdr.method as md
import adafdr.data_loader as dl
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# # RNA-seq: pasilla data
# ## Load the pasilla data
p,x = dl.data_pasilla()
print('p', p.shape)
print('x:', x.shape)
# ## covariate visualization
md.adafdr_explore(p, x, output_folder=None)
# ## hypothesis testing
res = md.adafdr_test(p, x, fast_mode=False, single_core=False)
n_rej = res['n_rej']
t_rej = res['threshold']
print('number of discoveris: %d'%np.sum(p<=t_rej))
plt.figure()
plt.scatter(x, p, alpha=0.2, s=4)
plt.scatter(x, t_rej, s=16, label='threshold')
plt.xlabel('covariate x', fontsize=16)
plt.ylabel('p-value', fontsize=16)
plt.legend(fontsize=16)
plt.show()
| vignettes/.ipynb_checkpoints/passila-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import scipy.optimize as sopt
from pysimu import ode2numba, ssa
from ipywidgets import *
# -
# %matplotlib notebook
# ## System definition
# +
S_base = 100.0e6
U_base = 20e3
Z_base = U_base**2/S_base
r_km = 0.127 # ohm/km
x_km = 0.113 # ohm/km
lenght = 1.0
R = r_km*lenght/Z_base
X = x_km*lenght/Z_base
Z = R +1j*X
Y = 1.0/Z
G_s_inf, B_s_inf = Y.real, Y.imag
sys = { 't_end':20.0,'Dt':0.01,'solver':'forward-euler', 'decimation':10, 'name':'vsg_pi_z2',
'models':[{'params':
{'K_p':0.01,
'K_i':0.1,
'K_q':0.1,
'T_q':0.1,
'Omega_b' : 2*np.pi*50,
'R_g':0.01,
'X_g':0.05,
'V_g': 1.0,
'K_f': 0.0,
'K_s': 1.0,
'H': 5.0,
'R_s':0.01,
'X_s':0.01,
},
'f':[
'dphi_s = Omega_b*(omega_s - 1)',
'dxi_p = epsilon_p',
'dxi_q = epsilon_q',
'dphi_g = Omega_b*(omega_g - 1) -1e-5*phi_g',
'domega_g = RoCoFpu - 1e-5*(omega_g - 1)'
],
'g':[
'omega_s @ -omega_s + K_p*epsilon_p + K_i*xi_p + 1',
'p_s_ref @ -p_s_ref + p_m',
'p_h @ p_h + 2*H*RoCoFpu',
'p_m0 @ p_m0 - p_m',
'v_gr @-v_gr + V_g*cos(phi_g)',
'v_gi @-v_gi + V_g*sin(phi_g)',
'i_sr @ -i_sr -(R_g*v_gr - R_g*v_sr + X_g*v_gi - X_g*v_si)/(R_g**2 + X_g**2)', # kron reference frame
'i_si @ -i_si -(R_g*v_gi - R_g*v_si - X_g*v_gr + X_g*v_sr)/(R_g**2 + X_g**2)', # kron reference frame
'i_sd_s@-i_sd_s + i_si', # phasor to dq in POI reference frame (v_sd_s = 0)
'i_sq_s@-i_sq_s - i_sr', # phasor to dq
'i_sd @-i_sd + cos(phi_s)*i_sd_s + cos(phi_s-pi/2)*i_sq_s',# virtual rotor reference frame
'i_sq @-i_sq - sin(phi_s)*i_sd_s - sin(phi_s-pi/2)*i_sq_s',# virtual rotor reference frame
'v_sd @ -v_sd + 0.0 + R_s*i_sd + X_s*i_sq',# virtual rotor reference frame
'v_sq @ -v_sq -1 + K_q*(epsilon_q + xi_q/T_q) + R_s*i_sq - X_s*i_sd',# virtual rotor reference frame
'v_sd_s @ -v_sd_s + cos(phi_s)*v_sd - sin(phi_s)*v_sq', # kron reference frame
'v_sq_s @ -v_sq_s + cos(phi_s-pi/2)*v_sd - sin(phi_s-pi/2)*v_sq', # kron reference frame
'v_si@-v_si + v_sd_s', # dq to phasor
'v_sr@-v_sr - v_sq_s', # dq to phasor
'epsilon_p@-epsilon_p + p_s_ref - p_s',
'epsilon_q@-epsilon_q + q_s_ref - q_s',
'p_s@-p_s+ i_sd*v_sd + i_sq*v_sq', # active power equation
'q_s@-q_s+ i_sd*v_sq - i_sq*v_sd', # reactive power equation
],
'u':{'p_m':0.4,'q_s_ref':0.1, 'RoCoFpu':0.0},
'y':['omega_s','p_s_ref','p_h','p_m0','v_gr','v_gi','i_sr', 'i_si', 'i_sd_s', 'i_sq_s', 'i_sd','i_sq','v_sd','v_sq','v_sd_s','v_sq_s',
'v_sr','v_si','epsilon_p','epsilon_q','p_s','q_s'],
'y_ini':['omega_s','p_s_ref','p_h','p_m0','v_gr','v_gi','i_sr', 'i_si', 'i_sd_s', 'i_sq_s', 'i_sd','i_sq','v_sd','v_sq','v_sd_s','v_sq_s',
'v_sr','v_si','epsilon_p','epsilon_q','p_s','q_s'],
'h':[
'p_m'
]}
],
'perturbations':[{'type':'step','time':100.0,'var':'p_m','final':1.01} ]
}
x,f = ode2numba.system(sys) ;
# -
25000*25/40/12
| examples/notebooks/uvsg/vsg_pi_z_builder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Real-Time Twitter Sentiment Plot
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.animation import FuncAnimation
# %matplotlib notebook
# +
plt.style.use('fivethirtyeight')
frame_len = 10000
fig = plt.figure(figsize=(9,6))
def animate(i):
data = pd.read_csv('sentiment.csv')
y1 = data['usa']
y2 = data['china']
if len(y1) <= frame_len:
plt.cla()
plt.plot(y1, label = 'USA')
plt.plot(y2, label = 'China')
else:
plt.cla()
plt.plot(y1[-frame_len:], label = 'USA')
plt.plot(y2[-frame_len:], label = 'China')
plt.legend(loc = 'upper-left')
plt.title('Twitter Sentiment Analysis for USA vs China')
plt.tight_layout()
ani = FuncAnimation(plt.gcf(), animate, interval = 100)
# -
| Real_time_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="MhoQ0WE77laV" colab_type="text"
# ##### Copyright 2018 The TensorFlow Authors.
# + id="_ckMIh7O7s6D" colab_type="code" cellView="form" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + id="vasWnqRgy1H4" colab_type="code" cellView="form" colab={}
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a
# # copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
# + [markdown] id="jYysdyb-CaWM" colab_type="text"
# # İlk sinir ağınızı eğitin: temel sınıflandırma
# + [markdown] id="S5Uhzt6vVIB2" colab_type="text"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/tr/r1/tutorials/keras/basic_classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Google Colab’da Çalıştır</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/tr/r1/tutorials/keras/basic_classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />GitHub'da Kaynağı Görüntüle</a>
# </td>
# </table>
# + [markdown] id="EVlm-3_0Q4zZ" colab_type="text"
# Note: Bu dökümanlar TensorFlow gönüllü kullanıcıları tarafından çevirilmiştir.
# Topluluk tarafından sağlananan çeviriler gönüllülerin ellerinden geldiğince
# güncellendiği için [Resmi İngilizce dökümanlar](https://www.tensorflow.org/?hl=en)
# ile bire bir aynı olmasını garantileyemeyiz. Eğer bu tercümeleri iyileştirmek
# için önerileriniz var ise lütfen [tensorflow/docs](https://github.com/tensorflow/docs)
# havuzuna pull request gönderin. Gönüllü olarak çevirilere katkıda bulunmak için
# [<EMAIL>](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-tr)
# listesi ile iletişime geçebilirsiniz.
# + [markdown] id="FbVhjPpzn6BM" colab_type="text"
# Bu yardımcı döküman, spor ayakkabısı ve gömlek gibi çeşitli giysi görüntülerini sınıflandırmak için bir sinir ağı modelini eğitir. Örnekte yer alan tüm detayları anlayamadıysanız bu sorun değil, aşağıda ilerledikçe ayrıntıları açıklanacak olan Tensorflow'a hızlı bir genel bakış yapılmaktadır.
#
# Bu örnekte, Tensorflow'da yapay zeka modellerinin oluşturulması ve eğitilmesinde kullanılan yüksek-seviye API olan, [tf.keras](https://www.tensorflow.org/r1/guide/keras) kullanmaktadır.
# + id="dzLKpmZICaWN" colab_type="code" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
# TensorFlow ve tf.keras
import tensorflow as tf
from tensorflow import keras
# Yardimci kutuphaneler
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
# + [markdown] id="yR0EdgrLCaWR" colab_type="text"
# ## Fashion MNIST veri kümesini yükleyelim
# + [markdown] id="DLdCchMdCaWQ" colab_type="text"
# Bu örnek uygulama, 10 kategoride 70,000 siyah-beyaz görüntü içeren [Fashion MNIST](https://github.com/zalandoresearch/fashion-mnist) veri kümesini kullanmaktadır. Aşağıda görüldüğü gibi bu veri kümesi, çeşitli giyim eşyalarının düşük çüzünürlükteki (28 x 28 piksel) görüntülerini içermektedir :
#
# <table>
# <tr><td>
# <img src="https://tensorflow.org/images/fashion-mnist-sprite.png"
# alt="Fashion MNIST sprite" width="600">
# </td></tr>
# <tr><td align="center">
# <b>Figür 1.</b> <a href="https://github.com/zalandoresearch/fashion-mnist">Fashion-MNIST örnekleri</a> (Zalando tarafından, MIT lisansı ile).<br/>
# </td></tr>
# </table>
#
# Fashion MNIST, klasik [MNIST](http://yann.lecun.com/exdb/mnist/) veri kümesinin yerine kolayca kullanılabilecek şekilde geliştirilmiştir. Klasik MNIST veri kümesi, yukarıda yer alan giysi görüntüleri ile aynı formatta, el yazısı rakam (0, 1, 2, vb) görüntülerini içermektedir.
#
# Fashion MNIST, klasik MNIST'e göre biraz daha zorlayıcı olduğu için ve çeşitliliğin arttırılması amacıyla kullanılmıştır. İki veri kümesi de nispeten küçüktür ve algoritmaların beklendiği gibi çalışıp çalışmadığının doğrulanmasında kullanılırlar. Ayrıca, yazdığımız kodun test edilmesi ve hataların tespit edilmesinde oldukça iyi bir başlangıç noktası oluştururlar.
#
# Oluşturacağımız sinir ağının eğitilmesinde 60,000 görüntü, eğitilmiş sinir ağının görüntü sınıflandırma doğruluğunu değerlendirmek içinse 10,000 görüntü kullanacağız. Fashion MNIST'e TensorFlow içerisinde doğrudan ulaşabilirsiniz, bunun için yapmanız gereken sadece veriyi içeri almak ve yüklemek:
# + id="7MqDQO0KCaWS" colab_type="code" colab={}
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# + [markdown] id="t9FDsUlxCaWW" colab_type="text"
# Veri kümesinin yüklenmesi sonucunda 4 NumPy dizisi oluşur:
#
# * `train_images` ve `train_labels` dizileri *eğitim veri setidir* - modelin eğitilmesinde kullanılır.
# * `test_images` ve `test_labels` dizileri *test veri setidir* - modelin test edilmesinde kullanılır.
#
# *train_images, test_images* 28x28 boyutunda ve piksel değerleri 0 ile 255 arasında değişen NumPy dizileridir. *train_labels, test_labels* ise 0 ile 9 arasında değişen ve her biri bir giyim eşyası sınıfı ile eşleşen tam sayı dizisidir:
#
# <table>
# <tr>
# <th>Etiket</th>
# <th>Sınıf</th>
# </tr>
# <tr>
# <td>0</td>
# <td>Tişört/Üst</td>
# </tr>
# <tr>
# <td>1</td>
# <td>Pantolon</td>
# </tr>
# <tr>
# <td>2</td>
# <td>Kazak</td>
# </tr>
# <tr>
# <td>3</td>
# <td>Elbise</td>
# </tr>
# <tr>
# <td>4</td>
# <td>Mont</td>
# </tr>
# <tr>
# <td>5</td>
# <td>Sandal</td>
# </tr>
# <tr>
# <td>6</td>
# <td>Gömlek</td>
# </tr>
# <tr>
# <td>7</td>
# <td>Spor Ayakkabı</td>
# </tr>
# <tr>
# <td>8</td>
# <td>Çanta</td>
# </tr>
# <tr>
# <td>9</td>
# <td>Yarım Bot</td>
# </tr>
# </table>
#
# Veri kümesi içerisindeki her bir görüntü tek bir etiket ile eşleştirilmiştir. *Sınıf isimleri* veri kümesi içerisinde yer almadığı için, daha sonra görüntüleri ekrana yazdırmak için bunları aşağıdaki gibi bir dizi içerisinde saklayalım:
# + id="IjnLH5S2CaWx" colab_type="code" colab={}
class_names = ['Tişört/Üst', 'Pantolon', 'Kazak', 'Elbise', 'Mont',
'Sandal', 'Gömlek', 'Spor Ayakkabı', 'Çanta', 'Yarım Bot']
# + [markdown] id="Brm0b_KACaWX" colab_type="text"
# ## Veriyi inceleyelim
#
# Modeli eğitmeye başlamadan önce, veri kümesi yapısını birlikte inceleyelim. Aşağıda, modelin eğitilmesinde kullanılan veri setinin 60,000 görüntüden oluştuğu ve her birinin 28 x 28 piksel olduğunu görmektesiniz:
# + id="zW5k_xz1CaWX" colab_type="code" colab={}
train_images.shape
# + [markdown] id="cIAcvQqMCaWf" colab_type="text"
# Benzer şekilde, eğitim veri setinde 60,000 adet etiket bilgisi yer almaktadır:
# + id="TRFYHB2mCaWb" colab_type="code" colab={}
len(train_labels)
# + [markdown] id="YSlYxFuRCaWk" colab_type="text"
# Her bir etiket 0 ile 9 arasında bir tam sayıdır:
# + id="XKnCTHz4CaWg" colab_type="code" colab={}
train_labels
# + [markdown] id="TMPI88iZpO2T" colab_type="text"
# Test veri kümesinde 10,000 görüntü mevcuttur. Her bir görüntü, benzer şekilde 28 x 28 piksel den oluşmaktadır:
# + id="2KFnYlcwCaWl" colab_type="code" colab={}
test_images.shape
# + [markdown] id="rd0A0Iu0CaWq" colab_type="text"
# Ve test veri seti 10,000 etiket bilgisini kapsamaktadır:
# + id="iJmPr5-ACaWn" colab_type="code" colab={}
len(test_labels)
# + [markdown] id="ES6uQoLKCaWr" colab_type="text"
# ## Verileri Ön İşleme
#
# Sinir ağının eğitilmesinden önce verinin bir ön işleme tabi tutulması gerekmektedir. Eğitim veri setindeki ilk görüntüyü inceleyecek olursanız, piksel değerlerinin 0 ile 255 arasında olduğunu göreceksiniz:
# + id="m4VEw8Ud9Quh" colab_type="code" colab={}
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
plt.show()
# + [markdown] id="Wz7l27Lz9S1P" colab_type="text"
# Bu görüntüler ile sinir ağını beslemeden önce, görüntülerin piksel değerlerini 0 ile 1 aralığına ölçekleyeceğiz. Bunun için, piksel değerlerini 255'e böleceğiz. Bu noktada *eğitim veri seti* ile *test veri seti*'nin aynı şekilde ön işlemden geçirilmesi, modelimizin doğru sonuç vermesi açısından önem taşımaktadır:
# + id="bW5WzIPlCaWv" colab_type="code" colab={}
train_images = train_images / 255.0
test_images = test_images / 255.0
# + [markdown] id="Ee638AlnCaWz" colab_type="text"
# *eğitim veri seti*'nin ilk 25 görüntüsünü, her bir görüntünün altında sınıf etiketi yazacak şekilde ekranda gösterelim. Verinin doğru formatta olduğunu doğruladıktan sonra artık modeli oluşturup eğitmeye hazırız.
# + id="oZTImqg_CaW1" colab_type="code" colab={}
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
plt.show()
# + [markdown] id="59veuiEZCaW4" colab_type="text"
# ## Modelin oluşturulması
#
# Sinir ağının oluşturulması için öncelikle model katmanlarının yapılandırılması ve sonrasında modelin derlenmesi gerekmektedir.
# + [markdown] id="Gxg1XGm0eOBy" colab_type="text"
# ### Katmanların hazırlanması
#
# Sinir ağını oluşturan temel yapı taşları *katman*'lardır. Katmanlar, kendilerine beslenen verileri kullanarak bu verilere ait çıkarımlar oluştururlar. Bu çıkarımların, bu örnekte görüntülerin sınıflandırılması olarak karşımıza çıkan problemin çözümüne yardımcı olması beklenir.
#
# Çoğu derin öğrenme modeli, birbirlerine bağlanmış birçok basit katman içermektedir. Çoğu katman, `tf.keras.layers.Dense` gibi, eğitme sürecinde öğrenilen parametrelere sahiptir.
# + id="9ODch-OFCaW4" colab_type="code" colab={}
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation=tf.nn.relu),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
# + [markdown] id="gut8A_7rCaW6" colab_type="text"
# Ağımızın ilk katmanı olan `tf.keras.layers.Flatten`, görüntülerin formatını 2 boyutlu sayı dizisinden (28 x 28 piksel), 28 * 28 = 784 piksel değerinden oluşan tek boyutlu bir sayı dizisine çevirir. Bu katmanın, görüntüleri oluşturan piksel satırlarının çıkarılarak, art arda birleştirilmesi ile oluştuğunu düşünebilirsiniz. Bu katmanda öğrenilecek parametre olmayıp, sadece görüntünün formatını düzenler.
#
# Görüntüyü oluşturan pikselleri tek boyutlu sayı dizisine düzleştirdikten sonra, ağımız ardaşık iki `tf.keras.layers.Dense` katmanını içerir. Bunlara, yoğun-bağlı veya tam-bağlı ağ katmanları denir. İlk 'Yoğun' katman 128 neron'a (düğüm) sahiptir. İkinci katman is 10 neronlu 'softmax' katmanıdır. Bu son katmanın çıktısı, toplam değerleri 1' eşit olan ve 10 farklı olasılık sonucunu içeren sayı dizisidir. Her bir düğüm, mevcut görüntünün hangi sınıfa ait olduğu belirten olasılık değerini içerir.
#
# ### Modelin derlenmesi
#
# Modelin eğitilmeye tamamıyla hazır olması öncesinde bir kaç düzenleme daha yapılması gerekmektedir. Bu düzenlemeler modelin 'derlenme' adımında eklenmektedir:
#
# * *Kayıp Fonksiyonu - Loss Function* — Bu fonksiyon modelin eğitim sürecinde ne kadar doğru sonuç verdiğini ölçer. Bu fonksiyonun değerini en aza indirgeyerek, modelin doğru istikamete "yönlendirmek" isteriz.
# * *Eniyileme - Optimizer* — Beslenen veriler ve kayıp fonksiyonu ile modelin nasıl güncellediğini belirler
# * *Metrikler - Metrics* — Eğitim ve test adımlarını gözlemlemek için kullanılır. Aşağıdaki örnekte, *doğruluk-accuracy*, modelin doğru sınıfladığı görüntü oranı, kullanılmaktadır.
# + id="Lhan11blCaW7" colab_type="code" colab={}
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# + [markdown] id="qKF6uW-BCaW-" colab_type="text"
# ## Modelin eğitilmesi
#
# Sinir ağının eğitilmesi aşağıdaki adımları gerektirir:
#
# 1. Eğitim veri setinin modele beslenmesi - bu örnekte veri setimiz yukarıda açıklanan 'train_images' ve 'train_labels' dizileridir.
# 2. Model etiketleri ve görüntüleri kullanarak çıkarımlar yapar ve öğrenir.
# 3. Modelden test veri setini - bu örnekte 'test_images' dizisidir, kullanarak tahminleme yapmasını isteriz. Sonucu 'test_labels' dizisindeki etiket ile eşleştirerek, bu kestirimlerin doğruluğunu teyid edebiliriz.
#
# Eğitimi başlatmak için 'model.fit' methodu çalıştırılır:
# + id="xvwvpA64CaW_" colab_type="code" colab={}
model.fit(train_images, train_labels, epochs=5)
# + [markdown] id="W3ZVOhugCaXA" colab_type="text"
# Model eğitimi süresince, kayıp ve doğruluk metrikleri ekranda gösterilir. Örneğimizdeki model, eğitim veri setiyle 0.88 (or 88%) doğruluk eğerine ulaşır.
# + [markdown] id="oEw4bZgGCaXB" colab_type="text"
# ## Model doğruluğunun değerlendirlmesi
#
# Sonrasında, modelin test veri seti ile nasıl bir performans gösterdiğini karşılaştıralım:
# + id="VflXLEeECaXC" colab_type="code" colab={}
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
print('Test accuracy:', test_acc)
# + [markdown] id="yWfgsmVXCaXG" colab_type="text"
# Test veri seti ile aldığımız model doğruluk değerinin, eğitim veri seti ile aldığımız model doğruluk değerinden biraz daha düşük olduğunu görmekteyiz. Eğitim doğruluk değeri ile test doğruluk değeri arasındaki bu farka *aşırı uyum-overfitting* denir. Aşırı uyum, modelin yeni veriler ile tahminleme yaparken, eğitim veri setine göre daha kötü performans göstermesi durumudur.
# + [markdown] id="xsoS7CPDCaXH" colab_type="text"
# ## Modeli kullanarak tahminleme yapalım
#
# Eğitilmiş modelimizi kullanarak, bir kaç görüntü için tahminleme yapabiliriz:
# + id="Gl91RPhdCaXI" colab_type="code" colab={}
predictions = model.predict(test_images)
# + [markdown] id="x9Kk1voUCaXJ" colab_type="text"
# Aşağıda, test veri setinde yer alan her bir görüntü için, modelimiz etiket sınıflandırması yapmaktadır. İlk tahminlemeye birlikte bakalım:
# + id="3DmJEUinCaXK" colab_type="code" colab={}
predictions[0]
# + [markdown] id="-hw1hgeSCaXN" colab_type="text"
# Tahminleme sonucu, 10 sayıdan oluşan bir dizi elde ederiz. Bu sayı dizisi bize, görüntünün 10 farklı sınıftan hangi giysi türüne ait olduğuna dair modelin "güvenini" tanımlamaktadır. Bu değerlere bakarak, hangi etiket sınıfının en yüksek güven değerine sahip olduğunu görebiliriz:
# + id="qsqenuPnCaXO" colab_type="code" colab={}
np.argmax(predictions[0])
# + [markdown] id="E51yS7iCCaXO" colab_type="text"
# Modelimiz yarım bot etiketi, (`veya class_names[9]`) için en yüksek kestirim güven değeri vermektedir. Ve test veri setindeki etikete bakarak sonucun doğru olduğunu görebiliriz:
# + id="Sd7Pgsu6CaXP" colab_type="code" colab={}
test_labels[0]
# + [markdown] id="ygh2yYC972ne" colab_type="text"
# 10 farklı sınıfın tümüne bakabilmek için sonucun grafiğini oluşturabiliriz:
# + id="DvYmmrpIy6Y1" colab_type="code" colab={}
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# + [markdown] id="d4Ov9OFDMmOD" colab_type="text"
# 0'ıncı görüntüye, tahminlere ve tahmin dizisine bakalım:
# + id="HV5jw-5HwSmO" colab_type="code" colab={}
i = 0
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions, test_labels)
plt.show()
# + id="Ko-uzOufSCSe" colab_type="code" colab={}
i = 12
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions, test_labels)
plt.show()
# + [markdown] id="kgdvGD52CaXR" colab_type="text"
# Çeşitli görüntüleri, tahminlemeleri ile ekranda gösterelim. Doğru tahminleme sonuçları mavi, yanlış olanlar ise kırmızı ile ekranda gösterilecektir. Rakamlar ise, 100 üzerinden, yapılan tahminlemenin güven değerini vermektedir. Güven değeri yüksek olsa bile, sonucun yanlış olabileceğini görebilirsiniz.
# + id="hQlnbqaw2Qu_" colab_type="code" colab={}
# Ilk X resmi, tahmin edilen etiketini ve asil etiketlerini cizelim.
# Dogru tahminler mavi, yanlis tahminler ise kirmizi renktedir.
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions, test_labels)
plt.show()
# + [markdown] id="R32zteKHCaXT" colab_type="text"
# Son olarak, eğitilmiş modeli kullanarak tek bir görüntü üzerinden tahminleme yapalım:
# + id="yRJ7JU7JCaXT" colab_type="code" colab={}
# Test veri setinden bir resim secelim.
img = test_images[0]
print(img.shape)
# + [markdown] id="vz3bVp21CaXV" colab_type="text"
# `tf.keras` modelleri, *veri yığınları* içerisindeki örnekler üzerinden tahminleme yapmak üzere optimize edilmiştirler. Tek bir görüntü kullanmamıza rağmen, bu nedenle görüntüyü bir listeye aktarmamız gerekmektedir:
# + id="lDFh5yF_CaXW" colab_type="code" colab={}
# Resmi tek ogesi kendisi olacagi bir listeye aktaralim.
img = (np.expand_dims(img,0))
print(img.shape)
# + [markdown] id="EQ5wLTkcCaXY" colab_type="text"
# Şimdi görüntüyü tahminleyelim:
# + id="o_rzNSdrCaXY" colab_type="code" colab={}
predictions_single = model.predict(img)
print(predictions_single)
# + id="6Ai-cpLjO-3A" colab_type="code" colab={}
plot_value_array(0, predictions_single, test_labels)
plt.xticks(range(10), class_names, rotation=45)
plt.show()
# + [markdown] id="cU1Y2OAMCaXb" colab_type="text"
# `model.predict` çalıştırıldığında, veri yığını içerisindeki her bir görüntüye ait bir liste verir. Yığın içerisinden görüntümüze (örneğimizdeki tek görüntü) ait tahminleme sonuçlarını alalım:
# + id="2tRmdq_8CaXb" colab_type="code" colab={}
prediction_result = np.argmax(predictions_single[0])
print(prediction_result)
# + [markdown] id="YFc2HbEVCaXd" colab_type="text"
# Daha önceden olduğu gibi, modelimiz etiket değeri olarak 9'u vermektedir.
| site/tr/r1/tutorials/keras/basic_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # %load /Users/facai/Study/book_notes/preconfig.py
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(color_codes=True)
#sns.set(font='SimHei')
plt.rcParams['axes.grid'] = False
#from IPython.display import SVG
def show_image(filename, figsize=None, res_dir=True):
if figsize:
plt.figure(figsize=figsize)
if res_dir:
filename = './res/{}'.format(filename)
plt.imshow(plt.imread(filename))
# -
# Chapter 8 Optimization for Training Deep Models
# =========
#
# optimization: finding the parameters $\theta$ of a neural network that significantly reduce a cost function $J(\theta)$.
# ### 8.1 How Learning Differs from Pure Optimization
#
# expectation is taken across **the data generating distribution** $p_{data}$ rather than just over the finite training set:
#
# \begin{equation}
# J^*(\theta) = \mathcal{E}_{(x, y) \sim p_{data}} L(f(x; \theta), y)
# \end{equation}
#
#
# #### 8.1.1 Empirical Risk Minimization
#
# Rather than optimizing the risk direcly, we optimize the empirical risk, and hope that the risk decreases significantly as well.
#
#
# #### 8.1.2 Surrogate Loss Functions and Early Stopping
#
#
# #### 8.1.3 Batch and Minibatch Algorithms
#
# Small batches can offer a regularing effect.
#
# gradient can handle smaller batch size like 100, while second-order methods typically require much large batch sizes like 10,000.
#
# minibatches must be selected randomly. For very large datasets, it is usually sufficient to shuffle the order of the dataset once and then store it in shuffled fashion.
#
# On the second pass, the estimator becomes biased because it is formed by re-sampling values used before.
# ### 8.2 Challenges in Neural Network Optimization
#
#
# #### 8.2.1 Ill-Conditioning
#
# To determin whether ill-conditioning, one can monitor the squared gradient norm $g^T g$ and the $g^T H g$ term. In many cases, the gradient norm does not shrink significantly throughout learning, but the $g^T H g$ term grows by more than order of magnitude.
#
#
# #### 8.2.2 Local Minima
#
# model identifiability problem: models with latent variables are often not identifiable <= weight space symmetry.
#
#
# #### 8.2.3 Plateaus, Saddle Points and Other Flat Regions
#
# + saddle point: local minimum along one cross-section, and local maximum along another another cross-section.
# - in higher dimensional spaces, local minima are rare and saddle points are more common.
# - difficult for newton's method, while easy for gradient descent.
#
# + maxima
# + wide, flat regions of constant value
#
#
# #### 8.2.4 Cliffs and Exploding Gradients
show_image("fig8_3.png")
# can be avoided using the *gradient clipping* heuristic
#
#
# #### 8.2.5 Long-Term Dependencies
#
# when graph becomes extremely deep => vanishing and exploding gradient problem
#
# Vanishing gradients make it difficult to know which direction the parameters should move to improve to the cost function, while exploding gradients can make learning unstable.
#
#
# #### 8.2.6 Inexact Gradients
#
#
# #### 8.2.7 Poor Correspondence between Local and Global Structure
#
# Many existing research directions are aimed at finding good initial points, rather than developing algorithms that use non-local moves.
#
#
# #### 8.2.8 Theoretical Limits of Optimization
# ### 8.3 Basic Algorithms
#
#
# #### 8.3.1 Stochastic Gradient Descent
#
# In practice, it is necessary to gradually decrease the learning rate over time.
#
# In practice, it is common to decay the learning rate linearly until iteration $\tau$:
#
# \begin{equation}
# \epsilon_k = (1 - \alpha) \epsilon_0 + \alpha \epsilon_{\tau}
# \end{equation}
#
# + $\tau$: a few hundred passes through the training set
# + $\epsilon_\tau \approx 1\% \, \epsilon_0$
# + $\epsilon_0$: monitor the first several iterations and use a learning rate that is higher than the best-performing learning rate at this time, but not so high that it causes severe instability.
#
# To study the convergence rate of an optimization algorithm, it is common to measure the *excess error* $J(\theta) - \min_\theta J(\theta)$.
# + SGD is applied to a convex problem: $O(\frac{1}{\sqrt{k}}$
# + in the stronly convex case it is $O(\frac{1}{k})$.
#
#
# #### 8.3.2 Momentum
#
# The momentum algorithm accumulates an exponentially decaying moving average of past gradients and continues to move in their direction.
#
# \begin{align}
# v &\gets \alpha v - \epsilon \Delta_\theta \left ( \frac1{m} \displaystyle \sum^m_{i = 1} L \left ( f(x^{(i)}; \theta), y^{(i)} \right ) \right ) \\
# \theta &\gets \theta + v
# \end{align}
#
# The larger $\alpha$ is relative to $\epsilon$, the more previous gradients affect the current direction.
show_image("fig8_5.png")
# the size of each step is $\frac{\epsilon \| g \|}{1 - \alpha} \implies$ it is thus helpful to think of the momentum hyperparameter in terms of $\frac1{1 - \alpha}$.
#
# + Common values of $\alpha$ used in practice include 0.5, 0.9 and 0.99.
# + Typically it begins with a small value and is later raised.
# + It is less important to adapt $\alpha$ over time than to shrink $\epsilon$ over time.
#
#
# #### 8.3.3 Nesterov Momentum
#
# Nesterov momentum: the gradient is evaluated after the current velocity is applied.
#
# \begin{align}
# v &\gets \alpha v - \epsilon \Delta_\theta \left ( \frac1{m} \displaystyle \sum^m_{i = 1} L \left ( f(x^{(i)}; \theta + \color{blue}{\alpha v}), y^{(i)} \right ) \right ) \\
# \theta &\gets \theta + v
# \end{align}
#
# 考虑了提前量
# ### 8.4 Parameter Initialization Strategies
#
# Designing improved initialization strategies is a difficult task because neural network optimization is not yet well understood.
#
# A further difficulty is that some initial points may be benefical from the viewpoint of optimization but detrimental from the viewpoint of generalization.
#
# complete certainty: break symmetry between different units
#
# + initialize each unit to compute a different function from all of the other units.
# + random initialization of the parameters.
# - Typically, set biases for each unit to heuristically chosen constants, and initilize only the weights randomly.
#
# ##### weight
#
# We can think of initializing the parameters $\theta$ to $\theta_0$ as being similar to imposing a Gaussian prior $p(\theta)$ with mean $\theta_0$.
# $\implies$ choose $\theta_0$ to be near 0 = more likely that units do not interact with each other than that they do interact.
#
# 1. normalized initialization: $W_{i, j} \sim U \left ( - \frac{6}{\sqrt{m + n}}, \frac{6}{\sqrt{m + n}} \right )$
# 2. initializing to random orthogonal matrices
# 3. perserve norms
# 4. sparse initialization
#
# A good rule of thumb for choosing the initial scales is to look at the range or standard deviation of activations or gradients on a single minibatch of data.
#
# ##### biase
#
# 1. Setting the biases to zero is compatible with most weight initialization schemes.
# 2. a few situations where we may set some biases to non-zero values:
# + for an output unit, often feneficial to initialize the bias to obtain the right marginal statistics of the output.
# + choose the bias to avoid causing too much saturation at initialization.
# eg: set the bias of ReLU hidden unit to 0.1 rather than 0
# + Sometimes a unit controls whether other units are able to participate in a function => all units have a chance to learn.
#
# ##### initialize model parameters using machine learning
#
# eg: to initialize a supervised model with the parameters learned by an unsupervised model trained on the same inputs.
# ### 8.5 Algorithms with Adaptive Learning Rates
#
# the cost is often highly sensitive to some directions in parameter space and insensitive to others. => adapt the learning rates of model parameters.
#
#
# #### 8.5.1 AdaGrad
#
# gradient accumulation
#
# $\text{rate} = \frac1{\sum \text{squared gradients}}$
#
#
# #### 8.5.2 RMSProp
#
# changing the gradient accumulation into an exponentially weighted moving average.
#
#
# #### 8.5.3 Adam
#
# adaptive moments: combination of RMSProp and momentum
#
# Currently, the most popular optimization algorithms actively in use include
# + SGD,
# + SGD with momentum,
# + RMSProp,
# + RMSProp with momentum,
# + AdaDelta and Adam.
#
# The choice of which algorithm to use, at this point, seems to depend largely on the user’s familiarity with the algorithm (for ease of hyperparameter tuning).
# ### 8.6 Approximate Second-Order Methods
#
# #### 8.6.1 Newton's Method
#
# a two-step iterative procedure:
#
# + update or compute the inverse Hessian
# + update the parameters: $\theta^* = \theta_0 - \mathbf{H}^{-1} \Delta_\theta J(\theta_0)$
#
#
# #### 8.6.2 Conjugate Gradients
#
#
# #### 8.6.3 BFGS
#
# L-BFGS
# ### 8.7 Optimization Strategies and Meta-Algorithms
#
# #### 8.7.1 Batch Normalization
#
# adaptive reparametrization => training very deep models
#
#
# #### 8.7.2 Coordinate Descent
#
# bad: variables are dependent.
#
#
# #### 8.7.3 Polyak Averaging
#
# averaging points.
#
#
# #### 8.7.4 Supervised Pretraining
#
# training sample models on simple tasks => then make the model more complex.
#
#
# #### 8.7.5 Designing Models to Aid Optimization
#
# In practice, it is more important to choose a model family that is easy to optimize than to use a powerful optimization algorithm.
#
#
# #### 8.7.6 Continuous Methods and Curriculumn Learning
| deep_learning/Optimization_for_Training_Deep_Models/note.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Brief Honor Code**. Do the homework on your own. You may discuss ideas with your classmates, but DO NOT copy the solutions from someone else or the Internet. If stuck, discuss with TA.
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# **1**. (20 points)
#
# Find the gradient and Hessian for the following equation
#
# $$
# f(x, y) = 1 + 2x + 3y + 4x^2 + 2xy + y^2
# $$
#
# - Plot the contours of this function using `matplotlib` in the box $-5 \le x \le 5$ and $-5 \le y \le 5$ using a $100 \times 100$ grid.
# - Then plot the gradient vectors using the `quiver` function on top of the contour plot using a $10 \times 10$ grid. Are the gradients orthogonal to the contours?
#
# Hint: Use `numpy.meshgrid`, `matplotlib.contour` and `matplotllib.quiver`.
# Gradient is
#
# $$
# \begin{bmatrix}
# 2 + 8x + 2y \\
# 3 + 2x + 2y
# \end{bmatrix}
# $$
#
# Hessian is
# $$
# \begin{bmatrix}
# 8 & 2 \\
# 2 & 2
# \end{bmatrix}
# $$
#
#
# +
x = np.linspace(-10, 10, 100)
y = np.linspace(-10, 10, 100)
X, Y = np.meshgrid(x, y)
Z = 1 + 2*X + 3*Y + 4*X**2 + 2*X*Y + Y**2
plt.contour(X, Y, Z, 15)
x = np.linspace(-10, 10, 10)
y = np.linspace(-10, 10, 10)
X, Y = np.meshgrid(x, y)
U = 2 + 8*X + 2*Y
V = 3 + 2*X + 2*Y
plt.quiver(X, Y, U, V, edgecolor='k', facecolor='r', linewidth=.5, minlength=5)
plt.axis('square')
pass
# -
# **2**. (30 points)
#
# This exercise is about using Newton's method to find the cube roots of unity - find $z$ such that $z^3 = 1$. From the fundamental theorem of algebra, we know there must be exactly 3 complex roots since this is a degree 3 polynomial.
#
# We start with Euler's equation
# $$
# e^{ix} = \cos x + i \sin x
# $$
#
# Raising $e^{ix}$ to the $n$th power where $n$ is an integer, we get from Euler's formula with $nx$ substituting for $x$
# $$
# (e^{ix})^n = e^{i(nx)} = \cos nx + i \sin nx
# $$
#
# Whenever $nx$ is an integer multiple of $2\pi$, we have
# $$
# \cos nx + i \sin nx = 1
# $$
#
# So
# $$
# e^{2\pi i \frac{k}{n}}
# $$
# is a root of 1 whenever $k/n = 0, 1, 2, \ldots$.
#
# So the cube roots of unity are $1, e^{2\pi i/3}, e^{4\pi i/3}$.
#
# While we can do this analytically, the idea is to use Newton's method to find these roots, and in the process, discover some rather perplexing behavior of Newton's method.
from sympy import Symbol, exp, I, pi, N, expand
expand(exp(2*pi*I/3), complex=True)
expand(exp(4*pi*I/3), complex=True)
plt.figure(figsize=(4,4))
roots = np.array([[1,0], [-0.5, np.sqrt(3)/2], [-0.5, -np.sqrt(3)/2]])
plt.scatter(roots[:,0], roots[:,1], s=50, c='red')
xp = np.linspace(0, 2*np.pi, 100)
plt.plot(np.cos(xp), np.sin(xp), c='blue');
# Newton's method for functions of complex variables - stability and basins of attraction. (30 points)
#
# 1. Write a function with the following function signature `newton(z, f, fprime, max_iter=100, tol=1e-6)` where
# - `z` is a starting value (a complex number e.g. ` 3 + 4j`)
# - `f` is a function of `z`
# - `fprime` is the derivative of `f`
# The function will run until either max_iter is reached or the absolute value of the Newton step is less than tol. In either case, the function should return the number of iterations taken and the final value of `z` as a tuple (`i`, `z`).
#
# 2. Define the function `f` and `fprime` that will result in Newton's method finding the cube roots of 1. Find 3 starting points that will give different roots, and print both the start and end points.
#
# Write the following two plotting functions to see some (pretty) aspects of Newton's algorithm in the complex plane.
#
# 3. The first function `plot_newton_iters(f, fprime, n=200, extent=[-1,1,-1,1], cmap='hsv')` calculates and stores the number of iterations taken for convergence (or max_iter) for each point in a 2D array. The 2D array limits are given by `extent` - for example, when `extent = [-1,1,-1,1]` the corners of the plot are `(-i, -i), (1, -i), (1, i), (-1, i)`. There are `n` grid points in both the real and imaginary axes. The argument `cmap` specifies the color map to use - the suggested defaults are fine. Finally plot the image using `plt.imshow` - make sure the axis ticks are correctly scaled. Make a plot for the cube roots of 1.
#
# 4. The second function `plot_newton_basins(f, fprime, n=200, extent=[-1,1,-1,1], cmap='jet')` has the same arguments, but this time the grid stores the identity of the root that the starting point converged to. Make a plot for the cube roots of 1 - since there are 3 roots, there should be only 3 colors in the plot.
def newton(z, f, fprime, max_iter=100, tol=1e-6):
"""The Newton-Raphson method."""
for i in range(max_iter):
step = f(z)/fprime(z)
if abs(step) < tol:
return i, z
z -= step
return i, z
def plot_newton_iters(p, pprime, n=200, extent=[-1,1,-1,1], cmap='hsv'):
"""Shows how long it takes to converge to a root using the Newton-Rahphson method."""
m = np.zeros((n,n))
xmin, xmax, ymin, ymax = extent
for r, x in enumerate(np.linspace(xmin, xmax, n)):
for s, y in enumerate(np.linspace(ymin, ymax, n)):
z = x + y*1j
m[s, r] = newton(z, p, pprime)[0]
plt.imshow(m, cmap=cmap, extent=extent)
def plot_newton_basins(p, pprime, n=200, extent=[-1,1,-1,1], cmap='jet'):
"""Shows basin of attraction for convergence to each root using the Newton-Raphson method."""
root_count = 0
roots = {}
m = np.zeros((n,n))
xmin, xmax, ymin, ymax = extent
for r, x in enumerate(np.linspace(xmin, xmax, n)):
for s, y in enumerate(np.linspace(ymin, ymax, n)):
z = x + y*1j
root = np.round(newton(z, p, pprime)[1], 1)
if not root in roots:
roots[root] = root_count
root_count += 1
m[s, r] = roots[root]
plt.imshow(m, cmap=cmap, extent=extent)
plt.grid('off')
plot_newton_iters(lambda x: x**3 - 1, lambda x: 3*x**2)
plt.grid('off')
m = plot_newton_basins(lambda x: x**3 - 1, lambda x: 3*x**2)
# **3**. (20 points)
#
# Consider the following function on $\mathbb{R}^2$:
#
# $$
# f(x_1,x_2) = -x_1x_2e^{-\frac{(x_1^2+x_2^2)}{2}}
# $$
#
# - Find the minimum under the constraint
# $$g(x) = x_1^2+x_2^2 \leq 10$$
# and
# $$h(x) = 2x_1 + 3x_2 = 5$$ using `scipy.optimize.minimize`.
# - Plot the function contours using `matplotlib`, showing the constraints $g$ and $h$ and indicate the constrained minimum with an `X`.
import scipy.optimize as opt
# +
def f(x):
return -x[0] * x[1] * np.exp(-(x[0]**2+x[1]**2)/2)
cons = ({'type': 'eq',
'fun' : lambda x: np.array([2.0*x[0] + 3.0*x[1] - 5.0]),
'jac' : lambda x: np.array([2.0,3.0])},
{'type': 'ineq',
'fun' : lambda x: np.array([-x[0]**2.0 - x[1]**2.0 + 10.0])})
x0 = [1.5,1.5]
cx = opt.minimize(f, x0, constraints=cons)
# +
x = np.linspace(-5, 5, 200)
y = np.linspace(-5, 5, 200)
X, Y = np.meshgrid(x, y)
Z = f(np.vstack([X.ravel(), Y.ravel()])).reshape((200,200))
plt.contour(X, Y, Z)
# g constraint
plt.plot(x, (5-2*x)/3, 'k:', linewidth=1)
# h constraint
theta = np.linspace(0, 2*np.pi, 100)
x = np.sqrt(10) * np.cos(theta)
y = np.sqrt(10) * np.sin(theta)
plt.plot(x, y, 'k:', linewidth=1)
plt.fill_between(x,y,alpha=0.15)
plt.text(cx['x'][0], cx['x'][1], 'x', va='center', ha='center', size=20, color='red')
plt.axis([-5,5,-5,5])
plt.title('Contour plot of f(x) subject to constraints g(x) and h(x)')
plt.xlabel('x1')
plt.ylabel('x2')
pass
# -
# **4** (30 points)
#
# Find solutions to $x^3 + 4x^2 -3 = x$.
#
# - Write a function to find brackets, assuming roots are always at least 1 unit apart and that the roots lie between -10 and 10
# - For each bracket, find the enclosed root using
# - a bisection method
# - Newton-Raphson (no guarantee to stay within brackets)
# - Use the end points of the bracket as starting points for the bisection methods and the midpoint for Newton-Raphson.
# - Use the companion matrix and characteristic polynomial to find the solutions
# - Plot the function and its roots (marked with a circle) in a window just large enough to contain all roots.
#
# Use a tolerance of 1e-6.
def f(x):
"""Fucntion to find zeros for."""
return x**3 + 4*x**2 -x - 3
def fp(x):
"""Derivative of f."""
return 3*x**2 + 8*x -1
# ### Bracketing function
def bracket(f, start, stop, step):
"""Find brackets where end points have different signs."""
brackets = []
for a in np.arange(start, stop, step):
b = a + step
if f(a) * f(b) < 0:
brackets.append([a, b])
return brackets
brackets = bracket(f, -10, 10, 1)
brackets
# ### Bisection
def bisect(f, a, b, tol=1e-6):
"""Bisection method."""
while np.abs(b - a) >= tol:
c = (a + b)/2
if (f(a) * f(c )) < 0:
b = c
else:
a = c
return c
for bracket in brackets:
a, b = bracket
x = bisect(f, a, b)
print(x)
# ### Newton-Raphson
def newton(f, fp, x, tol=1e-6):
"""Newton Raphson method."""
while np.abs(f(x)) >= tol:
x = x - f(x)/fp(x)
return x
for bracket in brackets:
a, b = bracket
x = (a + b)/2
x = newton(f, fp, x)
print(x)
# ### Companion matrix
A = np.array([
[-4, 1, 3],
[1,0,0],
[0,1,0]
])
roots = np.linalg.eigvals(A)
roots
x = np.linspace(-5, 2, 100)
plt.plot(x, f(x))
plt.axhline(0, c='black')
plt.scatter(roots, np.zeros_like(roots), s=50, c='red')
pass
| labs/Lab07_Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# This file describes software that supports the data and analysis in Starn and Belitz (in review) submitted to Water Resources Research (2017WR021531). This readme file describes the order of processing, required input, dependencies, and output from Jupyter Notebooks (JN) for the creation and processing of groundwater residence time distributions (GRTD) from groundwater flow models created using MODFLOW. Software and data supporting this work is available through the Science Base catalog of the U.S. Geological Survey at https://doi.org/10.5066/F7FQ9VTD.
# ## Disclaimer
# This software is preliminary or provisional and is subject to revision. It is being provided to meet the need for timely best science. The software has not received final approval by the U.S. Geological Survey (USGS). No warranty, expressed or implied, is made by the USGS or the U.S. Government as to the functionality of the software and related material nor shall the fact of release constitute any such warranty. The software is provided on the condition that neither the USGS nor the U.S. Government shall be held liable for any damages resulting from the authorized or unauthorized use of the software.
# ## Dependencies
# ### Provided in /Data
#
# * **P** -- shapefile of head calibration points
# * **W** -- shapefile of watershed boundaries used to create the General Models
# * **S** -- shapefile of sample data
# * **T** -- Excel workbook containing tracer curves (TracerLPM)
# ### Provided in /Models
#
# * **R** -- raster images of land surface (ned), model top (top), and model active area (ibound)
# * **F** -- a MODFLOW model readable by FloPy; may include head and budget output files for some JN
# * **K** -- `par.csv` file containing MODFLOW parameter information (created by General Model JN4)
# ### Created in these JNs
#
# * **M** -- MODPATH endpoint file
# * **A** -- `tau.txt` file containing mean age and recharge information
# # GRTD JNs
# Some JN are meant to be run on individual models and others are meant to process a number of models at once in batch mode. In either case, the model(s) to be processed are those that have `.nam` files in a directory in the `homes` directory list. Results from JN are in model directories (Individual) or `fig_dir` (Batch). The assignment of directory names in `homes` and `fig_dir` is made in each JN. Curly braces `{}` in file names below mean that the file is created in individual model directories and the braces are to be replaced with the model name.
#
# * MODFLOW model output is required and can be created by executing the `runMF.ipynb`.
#
# * The number and letter at the beginning of each JN name indicates the suggested order of processing; however, JN can be run in any order if the required input is present.
#
# * These JN were used to generate figures in Starn and Belitz (in review). The figure number is listed in the description section.
# ---
# ## 01a Calculate flux-weighted whole aquifer--Particles.ipynb
# *Individual *
# ### Input
# * **F**
# ### Output
# * **M, A**
# ---
# ## 01b Calculate flux-weighted whole aquifer--RTDs.ipynb
# *Individual *
# ### Input
# * **F, M, A**
# ### Output
# * `fit_dict_res_all_layers.pickle`
# * plots
# ### Description
# * `fit_dict_res_all_layers.pickle` contains all the information about particle travel times and the RTDs that fit them.
# * Plots show fits to individual models for all distributions including mixtures
# * Figure 6 shows plot for model number 4.
#
# ---
# ## 02 General Model model summary.ipynb
# *Batch *
# ### Input
# * **F, A, K**
# ### Output
# * `master_modflow_table.csv `
# * `tau_summary.csv`
# ### Description
# * `master_modflow_table.csv` contains a summary of explanatory features in multiple models
# * `tau_summary.csv` contains a concatenation of individual tau.txt files
#
# ---
# ## 03 Age grids and sections.ipynb
# *Individual *
# ### Input
# * **F, M**
# ### Output
# * plots
# ### Description
# * Plots show maps and cross sections of age and head
# * Figure 3 shows plots of age for model number 4
#
# ---
# ## 04 Compile multi-model head residuals.ipynb
# *Batch *
# ### Input
# * **P, W, F**
# ### Output
# * `head_resid_df.csv`
# * plots
# ### Description
# * `head_resid_df.csv` contains head residual information for multiple models
# * Figure 2 shows boxplots of residuals by model
#
# ---
# ## 05 Plot boxplots of multi-model distribution fits.ipynb
# *Batch *
# ### Input
# * `fit_dict_res_all_layers.pickle`
# ### Output
# * `rtd_error_all_layers_summary.csv`
# * plots
# ### Description
# * `rtd_error_all_layers_summary.csv` contains the RMSE errors for all fitted disitrbutions for multiple models
# * Figure 4 shows boxplots of RMSE errors for multiple models
# ---
# ## 06 Plot multi-model RTDs.ipynb
# *Batch *
# ### Input
# * **A, W**
# * `fit_dict_res_all_layers.pickle`
# ### Output
# * `weib_and_exp_fit.csv`
# * plots
# ### Description
# * `weib_and_exp_fit.csv` contains parameters and errors on Weibull and exponential fits
# * Figure 5 shows XY plots of RTD from multiple models.
# ---
# ## 07a Calculate flux-weighted whole aquifer--Process Wells.ipynb
# *Batch *
# ### Input
# * **F, S, R, W**
# ### Output
# * `node_df.csv`
# * `well_gdf.shp`
# * `sample_gdf.shp`
# * plots
# ### Description
# * `sample_gdf.shp` contains point locations and sampling data
# * `well_gdf.shp` contains point locations and well construction details
# * `node_df.csv` contains well information for each model layer the well penetrates
# * `well diagrams` is a directory that contains plots of well construction
# ---
# ## 07b Calculate flux-weighted whole aquifer--Well RTDS.ipynb
# *Batch *
# ### Input
# * **F, M**
# * `node_df.csv`
# * `well_gdf.shp`
# * `sample_gdf.shp`
# ### Output
# * `fit_dict_wells_{}.pickle`
# ### Description
#
# * `fit_dict_wells_{}.pickle` contains the distribution parameters at well cells
# ---
# ## 08 Plot BTCs and sample.ipynb
# *Batch or individual *
# ### Input
# * **T**
# * `fit_dict_wells_{}.pickle`
# ### Output
# * `sample_dict_wells.csv` (batch)
# * plots (indiviudal)
# ### Description
# * `sample_dict_wells.csv` is created in individual model directories and contains sample information, simulated equivalent concentrations, and the optimal porosity that produced them
# * Figure 8 shows an XY plot of ${^3}H$ breakthrough in two wells in model number 4
#
# ---
# ## 09 Plot Observed and Simulated Tracers with porosity adjusted.ipynb
# *Batch *
# ### Input
# * `sample_dict_wells.csv`
# ### Output
# * `master_sample_fit.csv`
# * `trit_fit_df.csv`
# * plot
# ### Description
# * `master_sample_fit.csv` contains a concatenation of `sample_dict_wells.csv` from all individual models
# * `trit_fit_df.csv` contains the results of the Kendall Tau statistical test
# * Figure 7 shows XY plot of ${^3}H$ residuals
# ---
# ## 10 Create and plot RTD metamodel.ipynb
# *Batch *
# ### Input
# * **W**
# * `master_modflow_table.csv`
# * `weib_and_exp_fit.csv`
# ### Output
# * plots
# ### Description
# * Figure 9a shows XY plot showing results of LASSO tuning
# * Figure 9b shows XY plot showing fitted parameters versus independent variables (labels)
# * Figure 9c shows bar chart showing importance of variables
# ---
# ## 11 Create and plot RTD poly metamodel.ipynb
# same as above but with quadratic expansion of the explanatory features
#
# Figure 10a-c shows results
| Notebooks/GRTD README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Europe Health
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import plotly.express as px
import plotly.graph_objects as go
from plotly.subplots import make_subplots
import plotly.figure_factory as ff
import os, shutil
# ## Open Questions
# * Data Source
# * Data current date
# ### Importing the data
#description of the respective features
pd.set_option('display.max_colwidth', None)
desc_df = pd.read_csv('Supply_Food_Data_Descriptions.csv', index_col = 'Categories')
desc_df
# +
# change the directory to the one of your "food" folder here
files = os.listdir('/Users/franz/Desktop/DV Project/Dashboard/diet_corona/food/')
if '.DS_Store' in files:
files.remove('.DS_Store')
directory = '/Users/franz/Desktop/DV Project/Dashboard/diet_corona/food/' + files[0]
this_df = pd.read_csv(directory)[["Area", "Item", "Value"]].set_index("Area", drop=True)
name = this_df.iloc[0,0]
this_df = this_df.rename(columns={"Value":name}).drop(columns="Item")
food_supply = this_df
for i in range(1,len(files)):
directory = '/Users/franz/Desktop/DV Project/Dashboard/diet_corona/food/' + files[i]
this_df = pd.read_csv(directory)[["Area", "Item", "Value"]].set_index("Area", drop=True)
name = this_df.iloc[0,0]
this_df = this_df.rename(columns={"Value":name}).drop(columns="Item")
food_supply = pd.concat([food_supply, this_df], axis=1)
# -
smo = pd.read_csv("share-of-adults-who-smoke.csv")
#retrieved from https://ourworldindata.org/smoking
smo = smo[smo["Year"] == 2016].rename(columns={"Code":"iso_a3", "Smoking prevalence, total (ages 15+)":"Smoking"})
smo = smo[["Entity", "Smoking"]].set_index("Entity", drop = True)
food_supply = pd.concat([food_supply, smo], axis = 1)
smo["Smoking"].std()
obes = pd.read_csv('Food_Supply_Quantity_kg_Data.csv')[["Country", "Obesity"]].set_index("Country")
food_supply = pd.concat([food_supply, obes], axis = 1)
food_supply = food_supply.reset_index().rename(columns={"index":"Country"})
# +
exp = pd.read_excel("h_exp.xlsx")
#https://ec.europa.eu/eurostat/databrowser/view/HLTH_SHA11_HF__custom_227597/bookmark/table?lang=en&bookmarkId=1530a1e6-767e-4661-9e15-0ed2f7fae0d5
exp.head()
# -
# ### Data Preprocessing
# +
#keep only countries from the EU
eu_countries_list = ["Austria","Belgium","Bulgaria","Croatia","Cyprus","Czechia","Denmark","Estonia","Finland","France",
"Germany","Greece","Hungary","Ireland","Italy","Latvia","Lithuania","Luxembourg","Malta","Netherlands","Poland",
"Portugal","Romania","Slovakia","Slovenia","Spain","Sweden"]
food_supply = food_supply[food_supply["Country"].isin(eu_countries_list)]
# +
def cz(s):
if s == "Czech Republic":
return "Czechia"
elif s == "Slovak Republic":
return "Slovakia"
else:
return s
exp["Country Name"] = exp["Country Name"].apply(cz)
exp = exp[exp["Country Name"].isin(eu_countries_list)]
len(exp)
# -
#check for nan values
if food_supply.dropna().shape == food_supply.shape:
print("There are no nan values!")
else:
print(food_supply.isna().sum())
#drop columns with nans
food_supply = food_supply.drop(columns=["Sugar Crops", "Miscellaneous"])
# ### Assessing the variables
print("There are", food_supply.shape[1], "different columns in the food_supply DataFrame!")
food_supply.columns
food_supply.info()
# ### Adding Covid-19 realted Data
# +
# cases, deaths, tests, vaccinations
# -
vacc = pd.read_csv("https://covid.ourworldindata.org/data/owid-covid-data.csv?v=2021-03-23")
vacc.columns
#retrieved from https://covid.ourworldindata.org/data/owid-covid-data.csv?v=2021-03-23
vacc = vacc[vacc["date"] == "2021-03-20"]
vacc = vacc[vacc["location"].isin(eu_countries_list)]
vacc = vacc[["location", "total_cases_per_million", "total_deaths_per_million", "life_expectancy", "human_development_index", "population_density", "median_age", "gdp_per_capita", "cardiovasc_death_rate", "diabetes_prevalence"]] # , "total_tests_per_thousand", "total_vaccinations_per_hundred
food_supply = food_supply.set_index("Country", drop=True)
exp = exp.set_index("Country Name", drop=True)
vacc = vacc.set_index("location", drop = True)
food_supply = pd.concat([food_supply, vacc, exp], axis = 1)
food_supply.head()
#check for nan values
if food_supply.dropna().shape == food_supply.shape:
print("There are no nan values!")
else:
print(food_supply.isna().sum())
# ### Checking correlations
def corr(df, color):
fig = plt.figure(figsize=(20,16))
mask = np.zeros_like(df.corr())
mask[np.triu_indices_from(mask)] = True
sns.heatmap(df.corr().round(2), annot=True, cmap=color, linewidths=0.2, mask=mask, vmin=-1, vmax=1)
plt.xticks(fontsize=11)
plt.yticks(fontsize=11)
plt.savefig("correlation.pdf");
food_supply = food_supply.rename(columns = {"life_expectancy":"Life Expectancy", "human_development_index":"Human Development Index",
"population_density":"Population Density", "median_age":"Median Age",
"gdp_per_capita":"GDP per Capita", "cardiovasc_death_rate":"Cardiovascular Death Rate",
"diabetes_prevalence":"Diabetes Prevalence"})
corr(food_supply, "plasma")
# ### Feature engineering
food_supply["Country"] = food_supply.index
food_supply.to_csv("scatter_data.csv")
# # Part 1: Eating habits of the 27 EU countries - Map(s)
# This part contains the per country per capita per year food consumption of different kinds of foods in kilogram. This includes the following 19 food categories:
foods = food_supply.iloc[:,:20].columns.tolist()
foods
# # Part 2: Correlation of the food habits and general information about the country
# ## Watch out for outliers!
# This part investigates if there are any correlations between a countries eating habits and the general data about the country. This general data includes the following variables:
# * GDP per capita
# * Human Development Index
# * Population Density
# * Median Age
def scatter_plot(df, x, y, size):
'''insert x and y as string while x entails color coding (abv avg...)'''
# feature engineering
col_name = str(y) + " above avg"
food_supply[col_name] = (df[y] > df[y].mean()).astype(int)
size = df[size] * 1/4
# plotting
fig = px.scatter(df, x=x, y =y, size=size, color_continuous_scale=px.colors.sequential.Plasma, color=col_name,hover_name=df.index, log_x=False, trendline = "ols", trendline_color_override="#bd3786",marginal_x = "box",marginal_y = "box", template="simple_white")
fig.show()
general_food_cor = food_supply.corr()[["GDP per Capita", "Human Development Index", "Median Age", "Population Density"]].T[foods]
fig = plt.figure(figsize=(20,5))
sns.heatmap(general_food_cor.round(2), annot=True, cmap="plasma", linewidths=0.2, vmin=-1, vmax=1)
general_cor = food_supply[["GDP per Capita", "Human Development Index", "Median Age", "Population Density"]].corr()
general_cor
scatter_plot(food_supply , "GDP per Capita", "Stimulants", "Human Development Index")
# # Part 3: Correlation of the food habits and health information about the country
# This part investigates if there are any correlations between a countries eating habits and the health data about the country. This data includes the following variables:
# * Obesity
# * Life Expectancy
# * Cardiovasc Death Rate
health_food_cor = food_supply.corr()[["Obesity", "Diabetes Prevalence", "Cardiovascular Death Rate", "Life Expectancy", "Health Expenditure" ]].T[foods].T
fig = plt.figure(figsize=(10,10))
sns.heatmap(health_food_cor.round(2), annot=True, cmap="plasma", linewidths=0.2, vmin=-1, vmax=1)#.set_yticklabels(["Obesity", "Diabetes Prevalence", "Cardiovascular Death Rate", "Life Expectancy", "Health Expenditure (% of GDP)"], rotation=0)
# +
df_corr_round = food_supply.corr()[["Obesity", "Diabetes Prevalence", "Cardiovascular Death Rate", "Life Expectancy", "Health Expenditure" ]].T[foods].T.round(2)
fig = ff.create_annotated_heatmap(
z=df_corr_round.to_numpy(),
x=df_corr_round.columns.tolist(),
y=df_corr_round.index.tolist(),
zmax=1, zmin=-1,
showscale=True,
hoverongaps=True,
ygap=3
)
fig.update_layout(yaxis_tickangle=-45, yaxis=dict(showgrid=False), xaxis=dict(showgrid=False))
# add title
fig.update_layout(title_text='<i><b>Correlation of Food Consumption and Health</b></i>',width=600, height=800)
# -
food_supply.columns
#health_cor = food_supply[["Obesity", "life_expectancy", "cardiovasc_death_rate", "gdp_per_capita", "human_development_index", "median_age", "population_density"]].corr()[["Obesity", "life_expectancy", "cardiovasc_death_rate"]].T
#health_cor
# +
import plotly.graph_objs as go
import statsmodels.api as sm
df = food_supply.copy()
col_name = str("Fruits - Excluding Wine") + "(above Average)"
df[col_name] = (df["Fruits - Excluding Wine"] > df["Fruits - Excluding Wine"].mean())#.astype(int)
fig = px.scatter(df, x="Life Expectancy", y ="Fruits - Excluding Wine", size=df["GDP per Capita"], color=col_name,hover_name=df.index, log_x=False,marginal_x = "box",marginal_y = "box", template="simple_white", color_discrete_sequence=["#0d0887", "#9c179e"])
# linear regression
regline = sm.OLS(df["Fruits - Excluding Wine"],sm.add_constant(df["Life Expectancy"])).fit().fittedvalues
# add linear regression line for whole sample
fig.add_traces(go.Scatter(x=df["Life Expectancy"], y=regline,
mode = 'lines',
marker_color='#fb9f3a',
name='OLS Trendline')
)
fig.show()
# +
df = food_supply.copy()
col_name = str("Fruits - Excluding Wine") + ": above Average"
df[col_name] = (df["Fruits - Excluding Wine"] > df["Fruits - Excluding Wine"].mean())#.astype(int)
size = df["Cardiovascular Death Rate"]
fig = px.scatter(df, x="Life Expectancy", y ="Fruits - Excluding Wine", size=size, color=col_name,hover_name=df.index, log_x=False, trendline = "ols", trendline_color_override="#bd3786",marginal_x = "box",marginal_y = "box", template="simple_white", color_discrete_sequence=px.colors.qualitative.G10)
fig.show()
# -
scatter_plot(food_supply , "Life Expectancy", "Fruits - Excluding Wine", "Cardiovascular Death Rate")
# # Part 4: Correlation of the food habits and Covid-19 data about the country
# This part investigates if there are any correlations between a countries eating habits and the Covid-19 data about the country. This data includes the following variables:
# * relative Cases
# * relative Deaths
health_food_cor = food_supply.corr()[["total_cases_per_million", "total_deaths_per_million"]].T[foods]
fig = plt.figure(figsize=(17,5))
sns.heatmap(health_food_cor.round(2), annot=True, cmap="plasma", linewidths=0.2, vmin=-1, vmax=1).set_yticklabels(["total_cases_per_million", "total_deaths_per_million"], rotation=0)
# +
#health_cor = food_supply[["total_cases_per_million", "total_deaths_per_million", "Obesity", "life_expectancy", "cardiovasc_death_rate", "gdp_per_capita", "human_development_index", "median_age", "population_density"]].corr()[["total_cases_per_million", "total_deaths_per_million"]].T
#health_cor
# -
scatter_plot(food_supply , "total_cases_per_million", "Alcoholic Beverages", "total_deaths_per_million")
# ## Evtl.: Part 4.0: Covid - general timelines per chosen country
# # Introducing health variables
# +
health = food_supply[["Country","Obesity", "Diabetes Prevalence", "Cardiovascular Death Rate", "Life Expectancy", "Health Expenditure" ]]
# +
health = food_supply[["Country","Obesity", "Diabetes Prevalence", "Cardiovascular Death Rate", "Life Expectancy", "Health Expenditure" ]]
fig = go.Figure()
fig.add_trace(go.Bar(
x=health["Country"],
y=health["Obesity"],
name="Obesity",
marker_color='#0d0887'
))
fig.add_trace(go.Bar(
x=health["Country"],
y=health["Diabetes Prevalence"],
name="Diabetes Prevalence",
marker_color='#7201a8'
))
fig.add_trace(go.Bar(
x=health["Country"],
y=health["Cardiovascular Death Rate"],
name="Cardiovascular Death Rate",
marker_color='#bd3786'
))
fig.add_trace(go.Bar(
x=health["Country"],
y=health["Life Expectancy"],
name="Life Expectancy",
marker_color='#ed7953'
))
fig.add_trace(go.Bar(
x=health["Country"],
y=health["Health Expenditure"],
name="Health Expenditure (% of GDP)",
marker_color='#fdca26'
))
# Here we modify the tickangle of the xaxis, resulting in rotated labels.
fig.update_layout(barmode='group', xaxis_tickangle=-45)
fig.update_layout(plot_bgcolor='white')
fig.update_yaxes(showline=True, linewidth=2, linecolor='black', gridcolor='grey')
fig.update_xaxes(showline=True, linewidth=2, linecolor='black')
# +
health = food_supply[["Country","Obesity", "Diabetes Prevalence", "Cardiovascular Death Rate", "Life Expectancy", "Health Expenditure" ]]
fig = go.Figure()
fig.add_trace(go.Bar(
x=health["Country"],
y=np.log(health["Obesity"]),
name="Obesity",
marker_color='#0d0887'
))
fig.add_trace(go.Bar(
x=health["Country"],
y=np.log(health["Diabetes Prevalence"]),
name="Diabetes Prevalence",
marker_color='#7201a8'
))
fig.add_trace(go.Bar(
x=health["Country"],
y=np.log(health["Cardiovascular Death Rate"]),
name="Cardiovascular Death Rate",
marker_color='#bd3786'
))
fig.add_trace(go.Bar(
x=health["Country"],
y=np.log(health["Life Expectancy"]),
name="Life Expectancy",
marker_color='#ed7953'
))
fig.add_trace(go.Bar(
x=health["Country"],
y=np.log(health["Health Expenditure"]),
name="Health Expenditure (% of GDP)",
marker_color='#fdca26'
))
# Here we modify the tickangle of the xaxis, resulting in rotated labels.
fig.update_layout(barmode='group', xaxis_tickangle=-45)
fig.update_layout(plot_bgcolor='white')
fig.update_yaxes(title_text="log")
fig.update_yaxes(showline=True, linewidth=2, linecolor='black', gridcolor='grey')
fig.update_xaxes(showline=True, linewidth=2, linecolor='black')
# -
print("Hey", "\<b> Hey \</b>" )
print('\033[1m{:10s}\033[0m'.format('Hey'))
BOLD = '\033[1m'
END = '\033[0m'
print('{}{}{}'.format(BOLD, "Hey", END))
food = ["Alcoholic Beverages", "Animal fats", "Cereals - Excluding Beer", "Eggs",
"Fish, Seafood", "Fruits - Excluding Wine", "Meat", "Milk - Excluding Butter",
"Offals", "Oilcrops", "Pulses", "Spices", "Starchy Roots","Stimulants",
"Sugar & Sweeteners", "Treenuts", "Vegetable Oils",
"Vegetables" ]
food.reverse()
print(food)
| apps/dash-food-consumption/notebooks/europe_health.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Beam bending calculator
#
# * Leave *lines 1-6 and 21-23 as they are*; they ensure that all necessary modules are imported and active.
#
# * Whatever code is written to the right of the numeral sign (**#**) is a comment. These comments are not executed, so you can safely ignore or delete them.
#
# The current code reproduces [this example](https://alfredocarella.github.io/simplebendingpractice/examples/example_1.html).
# +
# Initialization (leave this section as it is, unless you know what you are doing)
# %matplotlib inline
from beambending import Beam, DistributedLoadH, DistributedLoadV, PointLoadH, PointLoadV, x
# Go on and play around with the values in this section. See what happens!
### ------------------------------------------------------------------------ ###
beam = Beam(9)
beam.pinned_support = 2 # x-coordinate of the pinned support
beam.rolling_support = 7 # x-coordinate of the rolling support
beam.add_loads((
PointLoadH(10, 3), # 10kN pointing right, at x=3m
PointLoadV(-20, 3), # 20kN downwards, at x=3m
DistributedLoadV(-10, (3, 9)), # 10 kN/m, downwards, for 3m <= x <= 9m
DistributedLoadV(-20 + x**2, (0, 2)), # variable load, for 0m <= x <= 2m
))
### ------------------------------------------------------------------------ ###
# Output generation (leave this section as it is, unless you know what you are doing)
fig = beam.plot()
fig.savefig("./results.pdf")
# -
# Click the link to download [a pdf file with your case results](./results.pdf "Come on and click! You know you want to.").
# You can now go back up and continue to play around choosing different loads, beam lengths and placement of the beam supports.
# Add as many loads as you want to the list (but don't forget to put a comma between two loads).
#
# After you have made some changes, run the code cell again to calculate your new results.
#
# When you are comfortable with that, check what happens if you try more interesting expressions for the distributed loads, for example:
#
# ```python
# DistributedLoadV("2 * x**2 + cos(5)", (0, 3))
# ```
# By the way, a double asterisk is how you write a power in Python: ```"2 * x**2 + cos(5)"``` means $2x^2 + \cos(5)$
#
# **Note:** you can also try to include trigonometric functions, but in this case the whole mathematical expression __must be surrounded by quotation marks__, just as in the example above.
| simple_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import imagej
ij = imagej.init('/home/djproc/Fiji.app')
ij.getVersion()
# +
args = {
'name': 'Chuckles',
'age': 13,
'city': 'Nowhere'
}
result = ij.py.run_macro(macro_test.ijm, args)
print(result.getOutput('greeting'))
# -
| Run a macro from imagej.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.13 64-bit (''venv'': conda)'
# name: python3613jvsc74a57bd09fcd11040269ff8a468a26bab548888089e1e15f5655977e7680097ba56865af
# ---
# +
# # mount on google drive
# from google.colab import drive
# drive.mount('/content/drive/')
# -
import config
import sys
sys.path.append(config.root)
import matplotlib.pyplot as plt
import numpy as np
from utils.tiff_io import readTiff, writeTiff
from utils.imgShow import imgShow
# +
# sen2 images
scene_num = 'l5_scene_01'
path_img = config.root + '/data/dataset-l578/' + scene_num+ '.tif'
path_truth = config.root + '/data/dataset-l578/' + scene_num+ '_truth.tif'
image_src, image = readTiff(path_in=path_img)
truth_src, truth = readTiff(path_in=path_truth)
image = np.float32(np.clip(image/10000, a_min=0, a_max=1))
print(image.shape)
print(truth.shape)
# -
plt.figure(figsize=(9,5))
plt.subplot(1,2,1)
imgShow(image, col_bands=(2,1,0), clip_percent=2)
plt.subplot(1,2,2)
imgShow(truth, col_bands=(0,0,0), clip_percent=2)
| ipynb/img_show.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
import scipy
from matplotlib.colors import LogNorm
import pandas as pd
import seaborn as sns
# %matplotlib inline
# sys.path.insert(1, "/users/PAS0654/osu8354/ARA_cvmfs/source/AraRoot/analysis/ARA_analysis/SourceSearch")
# import deDisperse_util as util
import matplotlib.colors as colors
from matplotlib.pyplot import cm
# my_path_plots = os.path.abspath("./plots/")
# +
# mpl.use('agg')
mpl.rcParams['text.usetex'] = True
mpl.rcParams['mathtext.rm'] = 'Times New Roman'
mpl.rcParams['mathtext.it'] = 'Times New Roman:italic'
mpl.rcParams['mathtext.bf'] = 'Times New Roman:bold'
mpl.rcParams['text.latex.preamble'] = [r'\usepackage{amsmath}'] #for \text command
mpl.rc('font', family='serif', size=12)
mpl.rcParams['xtick.labelsize'] = 14
mpl.rcParams['ytick.labelsize'] = 14
mpl.rcParams['xtick.major.size'] = 5
mpl.rcParams['ytick.major.size'] = 5
mpl.rcParams['axes.titlesize'] = 18
mpl.rcParams['axes.labelsize'] = 18
# mpl.rc('font', size=16)
mpl.rc('axes', titlesize=20)
current_palette = sns.color_palette('colorblind', 10)
import warnings
warnings.filterwarnings("ignore")
# -
data_2NuBB = pd.read_csv("./data/2NuBB.csv")
data_2NuBB.head()
from mpl_toolkits.axes_grid.inset_locator import (inset_axes, InsetPosition,
mark_inset)
from scipy.interpolate import make_interp_spline
# +
x = np.linspace(0,6,6000)
def f(K,T0):
return (K*(T0-K)**5)*(1+2*K+(4/3)*K**2+(1/3)*K**3+(1/30)*K**4)#Primakoffand Rosen: <NAME> and <NAME>, Phys. Rev. 184, 1925 (1969)
T0=(2600)/511
I = quad(f, 0, 3, args=(T0))
y = f(x,T0)/I[0]
x2 = np.linspace(0.95,1.05,600)
# -
def gaussian(x, mu, sig):
return np.exp(-np.power(x - mu, 2.) / (2 * np.power(sig, 2.)))
# +
fig, axes = plt.subplots(nrows=1, ncols=1, figsize=(8, 6))
axes.plot(x/T0,y, ls = "--", label = r"$2\nu\beta\beta$")
axes.plot(x2, gaussian(x2,1,0.01)/50, label = r"$0\nu\beta\beta$")
axes.set_ylim(0,0.6)
axes.set_xlim(0,1.2)
axes.set_xlabel(r"$\frac{E_{e_1}+E_{e_2}}{Q_{\text{value}}}$")
axes.set_ylabel("Differential decay rate [arb.]")
plt.legend(loc = "upper left", fontsize = 16, title="Decay mode", fancybox=True)
left, bottom, width, height = [0.7, 0.63, 0.25, 0.2]
ins1 = fig.add_axes([left, bottom, width, height])
# ins2 = fig.add_axes([left, bottom, width, height])
# ins2.set_xticks([])
# ins2.set_yticks([])
ip = InsetPosition(axes, [left, bottom, width, height])
ins1.plot(x2, gaussian(x2,1,0.01)/1E5, c="C1");
# ins2.plot(x2,f(x2,T0)/(10000*I[0]))
# ins2.plot(x2, gaussian(x2,1,0.01)/1E5, c="C1");
# ax3 = fig.add_axes([left, bottom, width, height])
# ax3.plot(x/T0,y, ls = "--")
# ax3.set_xlim([-1,5])
# ax3.set_ylim([-1,5])
# ax3.set_axes_locator(ip)
mark_inset(axes, ins1, loc1=3, loc2=4, fc="none", ec='0.7')
plt.tight_layout()
plt.savefig("0NuBBdecay_spectrum.pdf")
# -
plt.plot(x2,f(x2,T0)/I[0])
| EnergySpectra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="dM_VpZ33OQ2u"
# 
# + [markdown] id="LR3OIVkDOQ2w"
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Public/7.Context_Spell_Checker.ipynb)
# + [markdown] id="V52RWZgkKskS"
# # 7 Context Spell Checker
# + id="OXiLnK8kOQ2x" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="f31b338c-a78a-434c-c96c-5b7f1f9b724c"
import os
# Install java
# ! apt-get update -qq
# ! apt-get install -y openjdk-8-jdk-headless -qq > /dev/null
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["PATH"] = os.environ["JAVA_HOME"] + "/bin:" + os.environ["PATH"]
# ! java -version
# Install pyspark
# ! pip install --ignore-installed -q pyspark==2.4.4
# ! pip install --ignore-installed -q spark-nlp==2.7.1
import sparknlp
spark = sparknlp.start() # for GPU training >> sparknlp.start(gpu = True) # for Spark 2.3 =>> sparknlp.start(spark23 = True)
from sparknlp.base import *
from sparknlp.annotator import *
from pyspark.ml import Pipeline
import pandas as pd
print("Spark NLP version", sparknlp.version())
print("Apache Spark version:", spark.version)
spark
# + [markdown] id="vzFwQ6KtOQ22"
#
# <b> if you want to work with Spark 2.3 </b>
# ```
# import os
#
# # Install java
# # # ! apt-get update -qq
# # # ! apt-get install -y openjdk-8-jdk-headless -qq > /dev/null
#
# # # !wget -q https://archive.apache.org/dist/spark/spark-2.3.0/spark-2.3.0-bin-hadoop2.7.tgz
#
# # # !tar xf spark-2.3.0-bin-hadoop2.7.tgz
# # # !pip install -q findspark
#
# os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
# os.environ["PATH"] = os.environ["JAVA_HOME"] + "/bin:" + os.environ["PATH"]
# os.environ["SPARK_HOME"] = "/content/spark-2.3.0-bin-hadoop2.7"
# # # ! java -version
#
# import findspark
# findspark.init()
# from pyspark.sql import SparkSession
#
# # # ! pip install --ignore-installed -q spark-nlp==2.5.5
#
# import sparknlp
#
# spark = sparknlp.start(spark23=True)
# ```
# + [markdown] id="TOjVN8NKOQ22"
# <H1> Noisy Channel Model Spell Checker - Introduction </H1>
#
# blogpost : https://medium.com/spark-nlp/applying-context-aware-spell-checking-in-spark-nlp-3c29c46963bc
#
# <div>
# <p><br/>
# The idea for this annotator is to have a flexible, configurable and "re-usable by parts" model.<br/>
# Flexibility is the ability to accommodate different use cases for spell checking like OCR text, keyboard-input text, ASR text, and general spelling problems due to orthographic errors.<br/>
# We say this is a configurable annotator, as you can adapt it yourself to different use cases avoiding re-training as much as possible.<br/>
# </p>
# </div>
#
#
# <b> Spell Checking at three levels: </b>
# The final ranking of a correction sequence is affected by three things,
#
#
# 1. Different correction candidates for each word - __word level__.
# 2. The surrounding text of each word, i.e. it's context - __sentence level__.
# 3. The relative cost of different correction candidates according to the edit operations at the character level it requires - __subword level__.
#
#
#
# + [markdown] id="jUCfqQbLOQ23"
# ### Initial Setup
# As it's usual in Spark-NLP let's start with building a pipeline; a _spell correction pipeline_. We will use a pretrained model from our library.
# + id="9wK6EnGvOQ24"
from sparknlp.annotator import *
from sparknlp.common import *
from sparknlp.base import *
from IPython.utils.text import columnize
# + id="-cBsZyHaOQ27" colab={"base_uri": "https://localhost:8080/"} outputId="4ddbec07-c3c6-4fdc-f32a-fd06d8bafa21"
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
tokenizer = RecursiveTokenizer()\
.setInputCols(["document"])\
.setOutputCol("token")\
.setPrefixes(["\"", "(", "[", "\n"])\
.setSuffixes([".", ",", "?", ")","!", "'s"])
spellModel = ContextSpellCheckerModel\
.pretrained('spellcheck_dl')\
.setInputCols("token")\
.setOutputCol("checked")\
.setErrorThreshold(4.0)\
.setTradeoff(6.0)
# + id="gY5j13B6OQ3A"
finisher = Finisher()\
.setInputCols("checked")
pipeline = Pipeline(
stages = [
documentAssembler,
tokenizer,
spellModel,
finisher
])
empty_ds = spark.createDataFrame([[""]]).toDF("text")
lp = LightPipeline(pipeline.fit(empty_ds))
# + [markdown] id="5Hcev_C7OQ3D"
# Ok!, at this point we have our spell checking pipeline as expected. Let's see what we can do with it,
# + id="IytF5v0_OQ3E" colab={"base_uri": "https://localhost:8080/"} outputId="615aab5c-07af-4485-aaf6-cf0a50f8162e"
lp.annotate("Plaese alliow me tao introdduce myhelf, I am a man of waelth und tiaste")
# + [markdown] id="EfgK96HuOQ3K"
# ### Word Level Corrections
# Continuing with our pretrained model, let's try to see how corrections work at the word level. Each Context Spell Checker model that you can find in Spark-NLP library comes with two sources for word candidates:
# # + a general vocabulary that is built during training(and remains unmutable during the life of the model), and
# # + special classes for dealing with special types of words like numbers or dates. These are dynamic, and you can modify them so they adjust better to your data.
#
# The general vocabulary is learned during training, and cannot be modified, however, the special classes can be updated after training has happened on a pre-trained model.
# This means you can modify how existing classes produce corrections, but not the number or type of the classes.
# Let's see how we can accomplish this.
# + id="xrECOVImOQ3L" colab={"base_uri": "https://localhost:8080/"} outputId="599ad399-61b1-449a-f53d-e2a76b99c6f3"
# First let's start with a loaded model, and check which classes it has been trained with
spellModel.getWordClasses()
# + [markdown] id="bnXKtN9JOQ3P"
# We have five classes, of two different types: some are vocabulary based and others are regex based,
# # + __Vocabulary based classes__ can propose correction candidates from the provided vocabulary, for example a dictionary of names.
# # + __Regex classes__ are defined by a regular expression, and they can be used to generate correction candidates for things like numbers. Internally, the Spell Checker will enumerate your regular expression and build a fast automaton, not only for recognizing the word(number in this example) as valid and preserve it, but also for generating a correction candidate.
# Thus the regex should be a finite regex(it must define a finite regular language).
#
# Now suppose that you have a new friend from Poland whose name is 'Jowita', let's see how the pretrained Spell Checker does with this name.
# + id="LyNv27gBO4y4"
beautify = lambda annotations: [columnize(sent['checked']) for sent in annotations]
# + id="j5rqzNm1OQ3P" colab={"base_uri": "https://localhost:8080/"} outputId="a6ad8826-380f-4694-ad02-8bcb463c6fe2"
# Foreign name without errors
sample = 'We are going to meet Jowita in the city hall.'
beautify([lp.annotate(sample)])
# + [markdown] id="qKyqcdy9OQ3S"
# Well, the result is not very good, that's because the Spell Checker has been trained mainly with American English texts. At least, the surrounding words are helping to obtain a correction that is a name. We can do better, let's see how.
#
# ## Updating a predefined word class
#
# ### Vocabulary Classes
#
# In order for the Spell Checker to be able to preserve words, like a foreign name, we have the option to update existing classes so they can cover new words.
# + id="hpKgt58OOQ3T" colab={"base_uri": "https://localhost:8080/"} outputId="db1883c2-4db0-4f3a-9788-b41ea4f194c1"
# add some more, in case we need them
spellModel.updateVocabClass('_NAME_', ['Monika', 'Agnieszka', 'Inga', 'Jowita', 'Melania'], True)
# Let's see what we get now
sample = 'We are going to meet Jowita at the city hall.'
beautify([lp.annotate(sample)])
# + [markdown] id="qoWZ657hOQ3W"
# Much better, right? Now suppose that we want to be able to not only preserve the word, but also to propose meaningful corrections to the name of our foreign friend.
# + id="azIAKc8UOQ3X" colab={"base_uri": "https://localhost:8080/"} outputId="3d665c94-a6b3-4f99-b622-3154cf8042d6"
# Foreign name with an error
sample = 'We are going to meet Jovita in the city hall.'
beautify([lp.annotate(sample)])
# + [markdown] id="z4fvnwl7OQ3b"
# Here we were able to add the new word to the class and propose corrections for it, but also, the new word has been treated as a name, that meaning that the model used information about the typical context for names in order to produce the best correction.
# + [markdown] id="JrpDPJ4YOQ3c"
# ### Regex Classes
# We can do something similar for classes defined by regex. We can add a regex, to for example deal with a special format for dates, that will not only preserve the date with the special format, but also be able to correct it.
# + id="fg5-ajMDOQ3d" colab={"base_uri": "https://localhost:8080/"} outputId="f5f0983c-1a95-4bbb-a760-c659c2bab67e"
# Date with custom format
sample = 'We are going to meet her in the city hall on february-3.'
beautify([lp.annotate(sample)])
# + id="YP2iOBsGOQ3g" colab={"base_uri": "https://localhost:8080/"} outputId="cf226ca9-5402-4fb4-d0c7-cee8e9fbabcd"
# this is a sample regex, for simplicity not covering all months
spellModel.updateRegexClass('_DATE_', '(january|february|march)-[0-31]')
beautify([lp.annotate(sample)])
# + [markdown] id="74XHt67TOQ3j"
# Now our date wasn't destroyed!
# + id="v4BxqhHNOQ3j" colab={"base_uri": "https://localhost:8080/"} outputId="4d21f0ba-3882-4f00-9c40-8c339238e682"
# now check that it produces good corrections to the date
sample = 'We are going to meet her in the city hall on febbruary-3.'
beautify([lp.annotate(sample)])
# + [markdown] id="0_qhmeH7OQ3n"
# And the model produces good corrections for the special regex class. Remember that each regex that you enter to the model must be finite. In all these examples the new definitions for our classes didn't prevent the model to continue using the context to produce corrections. Let's see why being able to use the context is important.
# ### Sentence Level Corrections
# The Spell Checker can leverage the context of words for ranking different correction sequences. Let's take a look at some examples,
# + id="FDuBuz29OQ3o" colab={"base_uri": "https://localhost:8080/"} outputId="17f89acd-a223-44dd-e19d-c8166ff2169f"
# check for the different occurrences of the word "siter"
example1 = ["I will call my siter.",\
"Due to bad weather, we had to move to a different siter.",\
"We travelled to three siter in the summer."]
beautify(lp.annotate(example1))
# + id="D4wn2v2XOQ3s" colab={"base_uri": "https://localhost:8080/"} outputId="b7052541-8e61-42ab-916e-1a50fd1a7c5b"
# check for the different occurrences of the word "ueather"
example2 = ["During the summer we have the best ueather.",\
"I have a black ueather jacket, so nice.",\
"I introduce you to my sister, she is called ueather."]
beautify(lp.annotate(example2))
# + [markdown] id="xd1gXbwyOQ3u"
# Notice that in the first example, 'siter' is indeed a valid English word, <br/> https://www.merriam-webster.com/dictionary/siter <br/>
# The only way to customize how the use of context is performed is to train the language model by training a Spell Checker from scratch. If you want to be able to train your custom language model, please refer to the Training notebook.
# Now we've learned how the context can help to pick the best possible correction, and why it is important to be able to leverage the context even when the other parts of the Spell Checker were updated.
# + [markdown] id="tAXDhU-LOQ3v"
# ### Subword level corrections
# Another fine tunning that our Spell Checker accepts is to assign different costs to different edit operations that are necessary to transform a word into a correction candidate.
# So, why is this important? Errors can come from different sources,
# # + Homophones are words that sound similar, but are written differently and have different meaning. Some examples, {there, their, they're}, {see, sea}, {to, too, two}. You will typically see these errors in text obtained by Automatic Speech Recognition(ASR).
# # + Characters can also be confused because of looking similar. So a 0(zero) can be confused with a O(capital o), or a 1(number one) with an l(lowercase l). These errors typically come from OCR.
# # + Input device related, sometimes keyboards cause certain patterns to be more likely than others due to letter locations, for example in a QWERTY keyboard.
# # + Last but not least, ortographic errors, related to the writter making mistakes. Forgetting a double consonant, or using it in the wrong place, interchanging letters(i.e., 'becuase' for 'because'), and many others.
#
# The goal is to continue using all the other features of the model and still be able to adapt the model to handle each of these cases in the best possible way. Let's see how to accomplish this.
# + id="nsvSTA5TOQ3v" colab={"base_uri": "https://localhost:8080/"} outputId="2570ee80-d1db-481d-827e-ab0e600f5c28"
# sending or lending ?
sample = 'I will be 1ending him my car'
lp.annotate(sample)
# + id="V31KhduLOQ35" colab={"base_uri": "https://localhost:8080/"} outputId="bd7e3e21-fdb4-4d62-873c-a8c70343751c"
# let's make the replacement of an '1' for an 'l' cheaper
weights = {'1': {'l': .1}}
spellModel.setWeights(weights)
lp.annotate(sample)
# + [markdown] id="oYMAj2YjOQ37"
# Assembling this matrix by hand could be a daunting challenge. There is one script in Python that can do this for you.
# This is something to be soon included like an option during training for the Context Spell Checker. Stay tuned on new releases!
# + [markdown] id="CF6roiiQOQ38"
# ## Advanced - the mysterious tradeoff parameter
# There's a clear tension between two forces here,
# # + The context information: by which the model wants to change words based on the surrounding words.
# # + The word information: by which the model wants to preserve as much an input word as possible to avoid destroying the input.
#
# Changing words that are in the vocabulary for others that seem more suitable according to the context is one of the most challenging tasks in spell correction. This is because you run into the risk of destroying existing 'good' words.
# The models that you will find in the Spark-NLP library have already been configured in a way that balances these two forces and produces good results in most of the situations. But your dataset can be different from the one used to train the model.
# So we encourage the user to play a bit with the hyperparameters, and for you to have an idea on how it can be modified, we're going to see the following example,
# + id="trwTZ0YROQ38" colab={"base_uri": "https://localhost:8080/"} outputId="53773215-a50a-4dd1-d4ab-b9ae184fef1a"
sample = 'have you been two the falls?'
beautify([lp.annotate(sample)])
# + [markdown] id="nvp4QocxOQ3-"
# Here 'two' is clearly wrong, probably a typo, and the model should be able to choose the right correction candidate according to the context. <br/>
# Every path is scored with a cost, and the higher the cost the less chances for the path being chosen as the final answer.<br/>
# In order for the model to rely more on the context and less on word information, we have the setTradeoff() method. You can think of the tradeoff as how much a single edition(insert, delete, etc) operation affects the influence of a word when competing inside a path in the graph.<br/>
# So the lower the tradeoff, the less we care about the edit operations in the word, and the more we care about the word fitting properly into its context. The tradeoff parameter typically ranges between 5 and 25. <br/>
# Let's see what happens when we relax how much the model cares about individual words in our example,
# + id="WUpyUecvOQ3_" colab={"base_uri": "https://localhost:8080/"} outputId="7bba5ef2-3998-41d0-f8d9-8cfbbac878b5"
spellModel.getTradeoff()
# + id="5zmQLB_UOQ4C" colab={"base_uri": "https://localhost:8080/"} outputId="020bf6dd-20f9-4622-905e-8b73b51424df"
# let's decrease the influence of word-level errors
# TODO a nicer way of doing this other than re-creating the pipeline?
spellModel.setTradeoff(5.0)
pipeline = Pipeline(
stages = [
documentAssembler,
tokenizer,
spellModel,
finisher
])
empty_ds = spark.createDataFrame([[""]]).toDF("text")
lp = LightPipeline(pipeline.fit(empty_ds))
beautify([lp.annotate(sample)])
# + [markdown] id="LD1RZYWCOQ4F"
# ## Advanced - performance
# + [markdown] id="H-EyNb0HOQ4G"
# The discussion about performance revolves around _error detection_. The more errors the model detects the more populated is the candidate diagram we showed above[TODO add diagram or convert this into blogpost], and the more alternative paths need to be evaluated. </br>
# Basically the error detection stage of the model can decide whether a word needs a correction or not; with two reasons for a word to be considered as incorrect,
# # + The word is OOV: the word is out of the vocabulary.
# # + The context: the word doesn't fit well within its neighbouring words.
# The only parameter that we can control at this point is the second one, and we do so with the setErrorThreshold() method that contains a max perplexity above which the word will be considered suspicious and a good candidate for being corrected.</br>
# The parameter that comes with the pretrained model has been set so you can get both a decent performance and accuracy. For reference, this is how the F-score, and time varies in a sample dataset for different values of the errorThreshold,
#
#
# |fscore |totaltime|threshold|
# |-------|---------|---------|
# |52.69 |405s | 8f|
# |52.43 |357s |10f|
# |52.25 |279s |12f|
# |52.14 |234s |14f|
#
# You can trade some minor points in accuracy for a nice speedup.
#
# + id="Pt7ca87zQaCP"
def sparknlp_spell_check(text):
return beautify([lp.annotate(text)])[0].rstrip()
# + id="vIFJmn6pPobo" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5c098e79-6fc2-4a6a-95a4-420f4c69bfad"
sparknlp_spell_check('I will go to Philadelhia tomorrow')
# + id="nO2315WwPtG4" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0e8c75a4-53e2-4e85-e1b7-d5fa64b3bebb"
sparknlp_spell_check('I will go to Philadhelpia tomorrow')
# + id="exFUMUU2P10V" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="300c9b00-5088-4257-9916-c8e35fc9fbbb"
sparknlp_spell_check('I will go to Piladelphia tomorrow')
# + id="cskimhCBP7jm" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="edde97f9-579a-4400-f024-916c0690d67b"
sparknlp_spell_check('I will go to Philadedlphia tomorrow')
# + id="FPzpMT5-QUg9" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="c727bb3d-7eb2-4ec3-aa6f-950aa953b8d4"
sparknlp_spell_check('I will go to Phieladelphia tomorrow')
# + id="eHESDrGyQ5aT"
| tutorials/Certification_Trainings/Public/7.Context_Spell_Checker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import keras
import os, time, pickle
from tqdm.notebook import tqdm
from sl_model import SL512, DSODSL512
from ssd_data import InputGenerator
from sl_utils import PriorUtil
from sl_training import SegLinkLoss, SegLinkFocalLoss
from utils.model import load_weights, calc_memory_usage
from utils.training import MetricUtility
# -
# ### Data
# +
from data_synthtext import GTUtility
file_name = 'gt_util_synthtext_seglink.pkl'
with open(file_name, 'rb') as f:
gt_util = pickle.load(f)
gt_util_train, gt_util_val = gt_util.split(0.9)
print(gt_util_train)
# -
# ### Model
# SegLink
model = SL512()
weights_path = './models/ssd512_voc_weights_fixed.hdf5'
#weights_path = '~/.keras/models/vgg16_weights_tf_dim_ordering_tf_kernels_notop.h5'
batch_size = 24
experiment = 'sl512_synthtext'
#experiment = 'sl512fl_synthtext'
# SegLink + DenseNet
model = DSODSL512()
#model = DSODSL512(activation='leaky_relu')
weights_path = None
batch_size = 6
experiment = 'dsodsl512_synthtext'
# SegLink + ResNet
from sl_model import SL512_resnet
model = SL512_resnet()
weights_path = None
batch_size = 10
experiment = 'sl512_resnet_synthtext'
# NOTE: did not converge and I had no time for further investigations
# +
freeze = []
if weights_path is not None:
if weights_path.find('ssd512') > -1:
layer_list = [
'conv1_1', 'conv1_2',
'conv2_1', 'conv2_2',
'conv3_1', 'conv3_2', 'conv3_3',
'conv4_1', 'conv4_2', 'conv4_3',
'conv5_1', 'conv5_2', 'conv5_3',
'fc6', 'fc7',
'conv6_1', 'conv6_2',
'conv7_1', 'conv7_2',
'conv8_1', 'conv8_2',
'conv9_1', 'conv9_2',
]
freeze = [
'conv1_1', 'conv1_2',
'conv2_1', 'conv2_2',
'conv3_1', 'conv3_2', 'conv3_3',
#'conv4_1', 'conv4_2', 'conv4_3',
#'conv5_1', 'conv5_2', 'conv5_3',
]
load_weights(model, weights_path, layer_list)
for layer in model.layers:
layer.trainable = not layer.name in freeze
else:
load_weights(model, weights_path)
prior_util = PriorUtil(model)
# -
# ### Training
# +
epochs = 100
initial_epoch = 0
#optimizer = tf.optimizers.SGD(learning_rate=1e-3, momentum=0.9, decay=0, nesterov=True)
optimizer = tf.optimizers.Adam(learning_rate=1e-3, beta_1=0.9, beta_2=0.999, epsilon=0.001, decay=0.0)
#loss = SegLinkLoss(lambda_offsets=1.0, lambda_links=1.0, neg_pos_ratio=3.0)
loss = SegLinkFocalLoss(lambda_segments=100.0, lambda_offsets=1.0, lambda_links=100.0, gamma_segments=2, gamma_links=2)
#regularizer = None
regularizer = keras.regularizers.l2(5e-4) # None if disabled
gen_train = InputGenerator(gt_util_train, prior_util, batch_size, model.image_size, augmentation=False)
gen_val = InputGenerator(gt_util_val, prior_util, batch_size, model.image_size, augmentation=False)
dataset_train, dataset_val = gen_train.get_dataset(), gen_val.get_dataset()
iterator_train, iterator_val = iter(dataset_train), iter(dataset_val)
checkdir = './checkpoints/' + time.strftime('%Y%m%d%H%M') + '_' + experiment
if not os.path.exists(checkdir):
os.makedirs(checkdir)
with open(checkdir+'/source.py','wb') as f:
source = ''.join(['# In[%i]\n%s\n\n' % (i, In[i]) for i in range(len(In))])
f.write(source.encode())
print(checkdir)
for l in model.layers:
l.trainable = not l.name in freeze
if regularizer and l.__class__.__name__.startswith('Conv'):
model.add_loss(lambda l=l: regularizer(l.kernel))
metric_util = MetricUtility(loss.metric_names, logdir=checkdir)
@tf.function
def step(x, y_true, training=False):
if training:
with tf.GradientTape() as tape:
y_pred = model(x, training=True)
metric_values = loss.compute(y_true, y_pred)
total_loss = metric_values['loss']
if len(model.losses):
total_loss += tf.add_n(model.losses)
gradients = tape.gradient(total_loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
else:
y_pred = model(x, training=True)
metric_values = loss.compute(y_true, y_pred)
return metric_values
#tf.profiler.experimental.start('./tblog')
for k in tqdm(range(initial_epoch, epochs), 'total', leave=False):
print('\nepoch %i/%i' % (k+1, epochs))
metric_util.on_epoch_begin()
for i in tqdm(range(gen_train.num_batches//4), 'training', leave=False):
x, y_true = next(iterator_train)
metric_values = step(x, y_true, training=True)
metric_util.update(metric_values, training=True)
#if i == 100: break
model.save_weights(checkdir+'/weights.%03i.h5' % (k+1,))
for i in tqdm(range(gen_val.num_batches), 'validation', leave=False):
x, y_true = next(iterator_val)
metric_values = step(x, y_true, training=False)
metric_util.update(metric_values, training=False)
#if i == 10: break
metric_util.on_epoch_end(verbose=1)
#if k == 1: break
#tf.profiler.experimental.stop()
# -
| SL_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
from pickle import Unpickler
infile = open('tests.pickle','rb')
#new_dict = pickle.loads(infile)
#infile.close()
with open('tests.pickle', 'rb') as f:
try:
test_data = pickle.load(f, encoding='latin1')
except TypeError:
test_data = pickle.load(f)
test_data['linear_grad_W']['outputs']
| week_2/local_tests/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import joblib
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
student = { 0: 0.,
1: 12.7062, 2: 4.3027, 3: 3.1824, 4: 2.7764, 5: 2.5706, 6: 2.4469, 7: 2.3646, 8: 2.3060,
9: 2.2622, 10: 2.2281, 11: 2.2010, 12: 2.1788, 13: 2.1604, 14: 2.1448, 15: 2.1314, 16: 2.1199,
17: 2.1098, 18: 2.1009, 19: 2.0930, 20: 2.0860}
# **1. Построим график средней точности при конкретном значении $\lambda$.**
data1 = joblib.load('convolutional-ewc-mas-stabilized.dmp')
data2 = joblib.load('convolutional-ewc-si-stabilized.dmp')
# +
lmbda1, lmbda2 = 40., 4.
means1 = np.mean(data1[lmbda1], axis=0)
intervals1 = np.std(data1[lmbda1], axis=0) * student[len(data1[lmbda1])-1] / np.sqrt(len(data1[lmbda1]))
means2 = np.mean(data2[lmbda2], axis=0)
intervals2 = np.std(data2[lmbda2], axis=0) * student[len(data2[lmbda2])-1] / np.sqrt(len(data2[lmbda2]))
x = range(1, len(means1) + 1)
fig, (ax1, ax2) = plt.subplots(
nrows=1, ncols=2,
figsize=(17, 4)
)
for ax, means, intervals in zip((ax1, ax2), (means1, means2), (intervals1, intervals2)):
ax.set_ylim(np.min(means-intervals), np.max(means+intervals))
ax.set_xlim(min(x), max(x))
ax.set_ylabel('Total accuracy')
ax.set_xlabel('Number of tasks')
ax.fill_between(x, means-intervals, means+intervals, color='grey', alpha=.2, label='confidence interval')
ax.plot(x, means, marker=".")
ax1.set_title(f'Mean accuracy degradation while continual learning\non {len(means1)} datasets'
f'with lambda {lmbda1} (log1)')
ax2.set_title(f'Mean accuracy degradation while continual learning\non {len(means2)} datasets '
f'with lambda {lmbda2} (log2)')
plt.legend()
plt.show()
# +
def func1(X, Y):
return [np.mean(data1[l], axis=0)[n-1] for l, n in zip(X, Y)]
def func2(X, Y):
return [np.mean(data2[l], axis=0)[n-1] for l, n in zip(X, Y)]
# +
fig = plt.figure(figsize=(29, 7))
ax1 = fig.add_subplot(111, projection='3d')
ax2 = fig.add_subplot(122, projection='3d')
x1 = sorted(list(data1.keys()))
y1 = list(range(1, 5))
X1, Y1 = np.meshgrid(x1, y1)
zs1 = np.array(func1(np.ravel(X1), np.ravel(Y1)))
Z1 = zs1.reshape(X1.shape)
ax1.plot_surface(X1, Y1, Z1, cmap=cm.coolwarm)
ax1.set_xlabel('Lambda')
ax1.set_ylabel('Number of tasks')
ax1.set_zlabel('Accuracy')
ax1.set_zlim(min(zs1), max(zs1))
ax1.view_init(5, 120)
x2 = sorted(list(data2.keys()))
y2 = list(range(1, 5))
X2, Y2 = np.meshgrid(x2, y2)
zs2 = np.array(func2(np.ravel(X2), np.ravel(Y2)))
Z2 = zs2.reshape(X2.shape)
ax2.plot_surface(X2, Y2, Z2, cmap=cm.coolwarm)
ax2.set_xlabel('Lambda')
ax2.set_ylabel('Number of tasks')
ax2.set_zlabel('Accuracy')
ax2.set_zlim(min(zs2), max(zs2))
ax2.view_init(5, 120)
plt.show()
# -
# **2. Посмотрим график точности после обучения 10-му датасету.**
data1 = joblib.load('convolutional-ewc-fisher-stabilized.dmp')
data2 = joblib.load('convolutional-ewc-mas-stabilized.dmp')
data3 = joblib.load('convolutional-ewc-si-stabilized.dmp')
lmbda1, lmbda2, lmbda3 = 2100, 40, 4
means1 = np.mean(data1[lmbda1], axis=0)
intervals1 = np.std(data1[lmbda1], axis=0) * student[len(data1[lmbda1])-1] / np.sqrt(len(data1[lmbda1]))
means2 = np.mean(data2[lmbda2], axis=0)
intervals2 = np.std(data2[lmbda2], axis=0) * student[len(data2[lmbda2])-1] / np.sqrt(len(data2[lmbda2]))
means3 = np.mean(data3[lmbda3], axis=0)
intervals3 = np.std(data3[lmbda3], axis=0) * student[len(data3[lmbda3])-1] / np.sqrt(len(data3[lmbda3]))
# +
fig, (ax1, ax2, ax3) = plt.subplots(nrows=1, ncols=3, figsize=(17, 4))
x = np.arange(1, 5)
for ax, means, intervals in zip((ax1, ax2, ax3), (means1, means2, means3), (intervals1, intervals2, intervals3)):
ax.set_ylim(np.min(means-intervals), np.max(means+intervals))
ax.set_xlim(min(x), max(x))
ax.set_ylabel('Total accuracy')
ax.set_xlabel('Number of tasks')
ax.fill_between(x, means-intervals, means+intervals, color='grey', alpha=.2, label='confidence interval')
ax.plot(x, means, marker=".")
ax1.set_title(f'Accuracy degradation while learning on {len(means1)} tasks\n'
f'(log1)')
ax2.set_title(f'Accuracy degradation while learning on {len(means2)} tasks\n'
f'(log2)')
ax3.set_title(f'Accuracy degradation while learning on {len(means3)} tasks\n'
f'(log3)')
plt.legend()
plt.show()
# -
print(sorted(data1.keys()))
print(sorted(data2.keys()))
print(sorted(data3.keys()))
# +
l1 = [k for k in sorted(list(data1.keys())) if k <6000] # sorted(list(data1.keys()))
means1 = np.asarray([np.mean(data1[l], axis=0)[3] for l in l1])
intervals1 = np.asarray([np.std(data1[l], axis=0)[3] * student[len(data1[l])-1] / np.sqrt(len(data1[l])) for l in l1])
l2 = [k for k in sorted(list(data2.keys())) if k <6000] # sorted(list(data2.keys())) #
means2 = np.asarray([np.mean(data2[l], axis=0)[3] for l in l2])
intervals2 = np.asarray([np.std(data2[l], axis=0)[3] * student[len(data2[l])-1] / np.sqrt(len(data2[l])) for l in l2])
l3 = [k for k in sorted(list(data3.keys())) if k <6000] # sorted(list(data3.keys()))
means3 = np.asarray([np.mean(data3[l], axis=0)[3] for l in l3])
intervals3 = np.asarray([np.std(data3[l], axis=0)[3] * student[len(data3[l])-1] / np.sqrt(len(data3[l])) for l in l3])
# -
for l, means, intervals in zip([l1, l2, l3], [means1, means2, means3], [intervals1, intervals2, intervals3]):
maxmean = max(means)
for idx in range(len(l)):
if means[idx]==maxmean: break
print(f'lambda: {l[idx]}, max mean accuracy {means[idx]}, interval {intervals[idx]}')
# +
fig, (ax1, ax2, ax3) = plt.subplots(
nrows=3, ncols=1,
figsize=(6, 18)
)
for ax, means, intervals, l in zip((ax1, ax2, ax3), (means1, means2, means3), (intervals1, intervals2, intervals3), (l1,l2,l3)):
ax.set_ylim(0.45, 0.63) # np.min(means-intervals), np.max(means+intervals)
ax.set_xlim(min(l), max(l)) #max(l)
ax.set_ylabel('Средняя точность после 4 задач')
ax.set_xlabel(r'$\lambda$')
ax.fill_between(l, means-intervals, means+intervals, color='grey', alpha=.2, label='confidence interval')
ax.plot(l, means, marker=".")
ax.plot([min(l), max(l)], [0.532, 0.532])
ax.plot([min(l), max(l)], [0.532, 0.532])
ax1.set_title('Важность весов по матрице Фишера')
ax2.set_title('Важность весов методом MAS')
ax3.set_title('Важность весов методом SI')
plt.legend()
plt.show()
# -
| section3/result-plots-conv-stabilized.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import glob
import os
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image, ImageOps, ImageFilter
from scipy.ndimage.filters import laplace, generic_filter
from scipy.ndimage import (gaussian_gradient_magnitude, sobel,
maximum_filter, gaussian_laplace)
from dataset_utils import load_img, img_to_dataset, grid_plot, get_training_windows
from tensorflow import keras as K
import tensorflow as tf
# -
tf.config.list_physical_devices('GPU')
# ## loading the data
data_source = '../../data/antibodies/DL-dataset/masked/'
glob.glob(data_source + '*')
class_files = {}
for name in ['NUC', 'ACA', 'ZIA', 'HOM']:
files = list(sorted(glob.glob(f'{data_source}/{name}/*')))
class_files[name] = files
# +
class_indices = {
'NUC': 0,
'ACA': 1,
'ZIA': 2,
'HOM': 3
}
masked_data = {}
for name, files in class_files.items():
class_index = class_indices[name]
imgs = [load_img(f, grayscale=True) for i, f in enumerate(files) if i % 2 == 1]
masks = [load_img(f, grayscale=True) for i, f in enumerate(files) if i % 2 == 0]
masked_data[name] = []
for img, mask in zip(imgs, masks):
masked_data[name].append((img, mask, class_index))
# +
sample_imgs = [masked_data['NUC'][i][0] for i in range(5)]
sample_masks = [masked_data['NUC'][i][1] for i in range(5)]
plt.figure(figsize=(20, 8))
grid_plot(sample_imgs + sample_masks, cmap='gray', rows=2, cols=5)
# -
# ## defining datasets
# +
def extract_meaningful_crops(img, mask, crop_size, crop_stride, mask_fill_threshold):
bin_mask = mask < 100
_, pos = get_training_windows(img, bin_mask,
pos_threshold=mask_fill_threshold, neg_threshold=1,
window_size=crop_size, stride=crop_stride)
return pos
def get_dataset_from_records(masked_records, crop_size, crop_stride, mask_fill_threshold):
X = []
y = []
for record in masked_records:
img, mask, class_index = record
crops = extract_meaningful_crops(img, mask, crop_size, crop_stride, mask_fill_threshold)
X.append(crops)
y.append(np.full(crops.shape[0], fill_value=class_index))
X = np.concatenate(X)
y = np.concatenate(y)
return X, y
def get_dataset(masked_data, train_samples, test_samples, crop_size, crop_stride, mask_fill_threshold):
X_train = []
y_train = []
X_test = []
y_test = []
for name, data in masked_data.items():
train_count = 0
test_count = 0
tmp_X_train = []
tmp_y_train = []
tmp_X_test = []
tmp_y_test = []
for record in data:
X, y = get_dataset_from_records([record], crop_size, crop_stride, mask_fill_threshold)
if train_count < train_samples:
train_count += X.shape[0]
tmp_X_train.append(X)
tmp_y_train.append(y)
elif test_count < test_samples:
test_count += X.shape[0]
tmp_X_test.append(X),
tmp_y_test.append(y)
tmp_X_train = np.concatenate(tmp_X_train)
tmp_y_train = np.concatenate(tmp_y_train)
tmp_X_test = np.concatenate(tmp_X_test)
tmp_y_test = np.concatenate(tmp_y_test)
X_train.append(tmp_X_train[:train_samples])
y_train.append(tmp_y_train[:train_samples])
X_test.append(tmp_X_test[:test_samples])
y_test.append(tmp_y_test[:test_samples])
print(f'finished {name}, train samples: {train_count}, test_samples: {test_count}')
X_train = np.concatenate(X_train)
y_train = np.concatenate(y_train)
X_test = np.concatenate(X_test)
y_test = np.concatenate(y_test)
return X_train, y_train, X_test, y_test
# -
# %%time
dataset = get_dataset(masked_data, train_samples=2000, test_samples=500,
crop_size=128, crop_stride=16, mask_fill_threshold=0.6)
X_train, y_train, X_test, y_test = dataset
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
# ## network training
from tensorflow.keras.applications import resnet_v2, inception_v3
# input_tensor = K.layers.Input(shape=(128, 128, 3))
base_model = inception_v3.InceptionV3(weights='imagenet', include_top=False)
base_model.trainable = False
base_model.summary()
model_top = K.layers.Conv2D(4, 2, activation='softmax')(base_model.output)
# model_top = K.layers.Dense(4, activation='softmax')(model_top)
full_model = K.Model(base_model.input, model_top, name='cells-transfer')
full_model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
full_model.summary()
# +
def prepare_X(X, expand_dims=False):
X_prep = X.copy()
X_prep = X_prep.astype(np.float32)
if expand_dims:
X_prep = tf.expand_dims(X_prep, axis=0)
X_prep = tf.expand_dims(X_prep, axis=-1)
X_prep = tf.repeat(X_prep, 3, axis=-1)
X_prep = tf.map_fn(inception_v3.preprocess_input, X_prep)
return X_prep
def prepare_y(y, depth=None):
if depth is not None:
y = tf.one_hot(y, depth=depth)
return tf.reshape(y, (-1, 1, 1, y.shape[-1]))
# +
# %%time
X_train_prep = prepare_X(X_train)
y_train_prep = prepare_y(y_train, depth=4)
X_test_prep = prepare_X(X_test)
y_test_prep = prepare_y(y_test, depth=4)
# -
X_train_prep.shape
# +
# run after pretraining
base_model.trainable = True
full_model.compile(optimizer=K.optimizers.Adam(learning_rate=0.0003),
loss='sparse_categorical_crossentropy', metrics=['accuracy'])
full_model.summary()
# -
history = full_model.fit(
X_train_prep, y_train_prep,
epochs=10,
batch_size=256,
validation_data=(X_test_prep, y_test_prep)
)
# +
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
# acc = np.concatenate([acc, history.history['accuracy']])
# val_acc = np.concatenate([val_acc, history.history['val_accuracy']])
plt.plot(acc, c='red', label='training accuracy')
plt.plot(val_acc, c='blue', label='validation accuracy')
plt.legend()
# -
# ### heatmap visualization
test_img = masked_data['NUC'][-1][0]
test_img_mask = masked_data['NUC'][-1][1]
test_img_prep = np.expand_dims(test_img, axis=0)
test_img_prep = prepare_X(test_img_prep)
plt.imshow(test_img)
# %%time
test_pred = full_model.predict(test_img_prep)[0]
test_pred_classes = np.argmax(test_pred, axis=-1)
plt.imshow(test_pred_classes)
plt.colorbar()
class_indices
# +
test_img_crops = extract_meaningful_crops(test_img, test_img_mask,
crop_size=128, crop_stride=16, mask_fill_threshold=0.6)
test_img_crops.shape
# -
test_crops_prep = prepare_X(test_img_crops)
test_crops_prep.shape
# %%time
crops_pred = full_model.predict(test_crops_prep)
crops_pred_classes = np.argmax(crops_pred, axis=-1)
crops_pred_classes = crops_pred_classes.reshape(-1)
crops_pred_classes.shape
plt.hist(crops_pred_classes)
print(np.unique(crops_pred_classes, return_counts=True))
print(np.unique(test_pred_classes, return_counts=True))
| new-development/transfer-learning-conv-output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p>
# -
import numpy as np
import pandas as pd
vgg = pd.read_csv('submission_10.csv')
inception = pd.read_csv('submission_08.csv')
resnet = pd.read_csv('submission_09.csv')
np.all(resnet.name == inception.name)
np.all(vgg.name == inception.name)
df = pd.DataFrame({'name': vgg.name, 'invasive': (vgg.invasive + inception.invasive)/2})
df = df[['name', 'invasive']]
df.to_csv("submission_ensem_4.csv", encoding="utf8", index=False)
from IPython.display import FileLink
FileLink('submission_ensem_4.csv')
# +
# Got 0.99527 on LB
# -
df = pd.DataFrame({'name': vgg.name, 'invasive': (vgg.invasive + inception.invasive + resnet.invasive)/3})
df = df[['name', 'invasive']]
df.to_csv("submission_ensem_5.csv", encoding="utf8", index=False)
from IPython.display import FileLink
FileLink('submission_ensem_5.csv')
# +
# Got 0.99397 on LB
| 10-ensemble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: python
# name: pysparkkernel
# ---
# <center>
# <a href="https://github.com/kamu-data/kamu-cli">
# <img alt="kamu" src="https://raw.githubusercontent.com/kamu-data/kamu-cli/master/docs/readme_files/kamu_logo.png" width=270/>
# </a>
# </center>
#
# <br/>
#
# <center><i>World's first decentralized real-time data warehouse, on your laptop</i></center>
#
# <br/>
#
# <div align="center">
# <a href="https://docs.kamu.dev/cli/">Docs</a> |
# <a href="https://docs.kamu.dev/cli/learn/learning-materials/">Tutorials</a> |
# <a href="https://docs.kamu.dev/cli/learn/examples/">Examples</a> |
# <a href="https://docs.kamu.dev/cli/get-started/faq/">FAQ</a> |
# <a href="https://discord.gg/nU6TXRQNXC">Discord</a> |
# <a href="https://kamu.dev">Website</a>
# </div>
#
#
# <center>
#
# <br/>
# <br/>
#
# # 1. Introduction
#
# </center>
# ## Welcome
#
# Hi, and thank you for checking out [kamu](https://github.com/kamu-data/kamu-cli) - the **new generation data management tool**!
#
# This environment comes with `kamu` command-line tool pre-installed, so give it a try now.
#
# <div class="alert alert-block alert-success">
# <b>Your turn:</b> Open the <b>Terminal</b> tab in Jupyter and run:
#
# <p style="background:black">
# <code style="background:black;color:white">$ kamu
# </code>
# </p>
# </div>
#
# <div class="alert alert-block alert-warning">
# <details>
# <summary style="display:list-item"><b>New to Jupyter?</b></summary>
#
# * Go back to the Jupyter's <b>main tab</b> that shows the list of files
# * In the top right corner click <b>New -> Terminal</b>
# * Now you can switch between the terminal tab and this lesson as you continue
#
# </details>
# </div>
#
# ## What is Kamu for?
#
# [Kamu](https://github.com/kamu-data/kamu-cli) is a tool based on [Open Data Fabric](http://opendatafabric.org/) protocol that connects publishers and consumers of data into a <mark>decentralized data supply chain</mark>. It allows you to get data fast, in a ready-to-use form for analysis and ML tasks, ensure it is trustworthy and easy to keep up to date.
#
# In this demo, we are going to explore some of the key features of `kamu` through some <mark>real world examples</mark>.
#
# <div class="alert alert-block alert-warning">
# <b>Short on time?</b> See <a href="https://www.youtube.com/watch?v=oUTiWW6W78A&list=PLV91cS45lwVG20Hicztbv7hsjN6x69MJk">this video</a> for a quick tour of key features.
# </div>
#
# If you have any questions throughout this demo - you can chat to us on [Discord](https://discord.gg/nU6TXRQNXC) or create an issue in [kamu-cli](https://github.com/kamu-data/kamu-cli) GitHub repository.
# ## Workspaces
#
# You start working with `kamu` by creating a workspace. A workspace is just a directory where `kamu` stores data and metadata of the datasets.
#
# <div class="alert alert-block alert-success">
# Go ahead and create your first workspace (we'll do it right in the home directory):<br/>
#
# <p style="background:black">
# <code style="background:black;color:white">$ kamu init
# </code>
# </p>
# </div>
#
# <div class="alert alert-block alert-info">
# <b>Note:</b> Similarly to `git` it will create a `.kamu` directory in the folder you ran the command in.
# </div>
#
# Your new workspace is currently empty. Confirm that by running:
#
# <p style="background:black">
# <code style="background:black;color:white">$ kamu list
# </code>
# </p>
# ## Adding the first dataset
# Let's add some new datasets to our workspace!
#
# In `kamu`, datasets are defined using `.yaml` files. You can import them with `kamu add` command.
#
# In this demo, we are going to work with some disagregated COVID-19 datasets published by different provinces of Canada.
#
# <div class="alert alert-block alert-success">
# To add a dataset to the workspace run:<br/>
#
# <p style="background:black">
# <code style="background:black;color:white">$ kamu add demo/datasets/ca.bccdc.covid19.case-details.yaml
# </code>
# </p>
# </div>
#
# <div class="alert alert-block alert-info">
# <b>Note:</b> Every command in <code>kamu</code> is well documented. Try running <code>kamu add -h</code> or <code>kamu add --help</code> to see all the parameters and useful examples.
# </div>
#
# This particular dataset includes case data form British Columbia, including day, age group, gender and area where the case was registered. Such datasets that ingest or receive external data are called `root` datasets and contain valuable data that cannot be reconstructed if lost (also known as **source data**).
#
# The dataset definition file looks like this:
#
# ```yaml
# version: 1
# kind: DatasetSnapshot
# content:
# id: ca.bccdc.covid19.case-details
# source:
# kind: root
# fetch:
# kind: url
# url: http://www.bccdc.ca/Health-Info-Site/Documents/BCCDC_COVID19_Dashboard_Case_Details.csv
# read:
# kind: csv
# separator: ','
# header: true
# nullValue: ''
# preprocess:
# kind: sql
# engine: spark
# query: >
# SELECT
# CAST(UNIX_TIMESTAMP(Reported_Date, "yyyy-MM-dd") as TIMESTAMP) as reported_date,
# Classification_Reported as classification,
# id,
# ha,
# sex,
# age_group
# FROM input
# merge:
# kind: ledger
# primaryKey:
# - id
# vocab:
# eventTimeColumn: reported_date
# ```
#
# As you can see, it tells `kamu` where to fetch the data from, what type of data to expect, and all the pre-processing steps needed to shape the data into a nice typed schema.
#
# <div class="alert alert-block alert-info">
# <b>Note:</b> Kamu strictly follows <b>"data as code"</b> philosophy in which you never alter the data directly. Instead, you express all transformations with queries (SQL in this case).
# </div>
# ### Pulling the data in
#
# We can now see the dataset in our workspace, but it is still empty:
#
# <p style="background:black">
# <code style="background:black;color:white">$ kamu list
# </code>
# </p>
#
# We told `kamu` where to get data from, but did not fetch it yet.
#
# <div class="alert alert-block alert-success">
# So let's run the following command to fetch data:<br/>
#
# <p style="background:black">
# <code style="background:black;color:white">$ kamu pull ca.bccdc.covid19.case-details
# </code>
# </p>
# </div>
#
# <div class="alert alert-block alert-info">
# Make sure to use <b>shell completions</b>, they will save you a lot of typing!
# <p style="background:black">
# <code style="background:black;color:white">$ kamu pull c<TAB>
# </code>
# </p>
# </div>
#
# During this time `kamu` will fetch the data from its source, read and preprocess it as specified.
#
# <div class="alert alert-block alert-success">
# Once completed, we can use <code>tail</code> command to see a sample of the new data:
#
# <p style="background:black">
# <code style="background:black;color:white">$ kamu tail ca.bccdc.covid19.case-details
# </code>
# </p>
# </div>
# ## Ledger nature of Data and Metadata
#
# A very important aspect of `kamu` is that it stores the history of data, not just snapshots. If we run the `pull` command on these datasets tomorrow, it will only add the records that were not previously observed.
#
# <div class="alert alert-block alert-success">
# Run <code>pull</code> command again to verify that our data is still up-to-date with the source:
#
# <p style="background:black">
# <code style="background:black;color:white">$ kamu pull ca.bccdc.covid19.case-details
# </code>
# </p>
# </div>
#
# In other words, in `kamu` <mark>data is a ledger</mark> - an append-only record of events where past events never change (are immutable). The exact way how external data is transformed into a ledger is determined by the merge strategies documented [here](https://github.com/kamu-data/kamu-cli/blob/master/docs/merge_strategies.md).
#
# Additionally, every event that affects the dataset is stored in so-called **metadata chain**.
#
# <div class="alert alert-block alert-success">
# Inspect the metadata chain using the <code>log</code> command ("Q" to close):
#
# <p style="background:black">
# <code style="background:black;color:white">$ kamu log ca.bccdc.covid19.case-details
# </code>
# </p>
# </div>
#
# As you can see, **metadata is also a ledger**! There are two metadata **blocks**:
# - First, corresponding to the dataset creation
# - Second, corresponding to N new records added by the last `pull`
#
# You can think of the metadata chain as `git` commit log, except instead of data - it stores an accurate **history of events** that affected how dataset looks like throughought its entire lifetime.
# ## Analyzing Data
#
# Getting raw data in is just a small first step on our journey towards collaboration on data, but before we continue, let's take a quick break and see how you can analyze the data that we already have.
#
# ### SQL Shell
#
# Kamu has a built-in SQL shell which you can start by running:
#
# <p style="background:black">
# <code style="background:black;color:white">$ kamu sql
# </code>
# </p>
#
# <div class="alert alert-block alert-info">
# The default SQL shell is based on the <a href="https://spark.apache.org/">Apache Spark</a>.
# </div>
#
# <div class="alert alert-block alert-success">
# Once the shell starts, try the following queries:
#
# <p style="background:black">
# <code style="background:black;color:white">> show tables;</code>
# </p>
# <p style="background:black">
# <code style="background:black;color:white">> describe `ca.bccdc.covid19.case-details`;</code>
# </p>
# <p style="background:black">
# <code style="background:black;color:white">> select * from `ca.bccdc.covid19.case-details` limit 10;</code>
# </p>
# </div>
#
# Press **Ctrl + D** to exit.
#
# ### Notebooks
#
# When you install `kamu` on your computer you can use `kamu notebook` command to start an integrated Jupyter
# Notebook environment, identical to the one you are currently using.
#
# Since we're already in the notebook environment - let's give this integration a try!
#
# <div class="alert alert-block alert-success">
# Start by loading <code>kamu</code> Jupyter extension:
# </div>
# %load_ext kamu
# <div class="alert alert-block alert-warning">
# <details>
# <summary style="display:list-item"><b>New to Jupyter?</b></summary>
#
# Jupyter notebooks contain cells that are **executable**, so static text can me mixed with computations and data visualization.
#
# **You** are in control of what runs when, so you'll need to **select the code cell above** and then click the **"Run"** button on the top panel, or press `Shift + Enter`.
#
# </details>
# </div>
#
# We can now import the dataset we have in our workspace into this notebook environment. We can also give it a less verbose alias.
#
# <div class="alert alert-block alert-success">
# Run the below to import the dataset (may take 15 or so seconds first time):
# </div>
# %import_dataset ca.bccdc.covid19.case-details --alias bc_covid19
# <div class="alert alert-block alert-success">
# To see the schema and number of records in the dataset run:
# </div>
bc_covid19.printSchema()
bc_covid19.count()
# <div class="alert alert-block alert-info">
# <details>
# <summary style="display:list-item"><b>What did we just run?</b></summary>
#
# The code you type into a regular cell is executed by [PySpark](https://spark.apache.org/docs/latest/api/python/) server that `kamu` runs when you are working with notebooks.
#
# So it's a Python code, but it is **executed remotely**, not in the notebook kernel. We will discuss benefits of this later.
#
# </details>
# </div>
#
# You can use the `%%sql` cell command to run SQL queries on the imported datasets.
#
# <div class="alert alert-block alert-success">
# To see a sample of data run:
# </div>
# + language="sql"
# select * from bc_covid19
# order by reported_date desc
# limit 5
# -
# <div class="alert alert-block alert-info">
# <details>
# <summary style="display:list-item"><b>What did we just run?</b></summary>
#
# Similarly to the PySpark code, the queries in `%%sql` cells are sent to and executed by the Spark SQL engine. The results are then returned back to the notebook kernel.
#
# </details>
# </div>
#
# <div class="alert alert-block alert-success">
# Let's run this simple SQL query to build a histogram of cases by the age group:
# </div>
# + language="sql"
# select
# age_group,
# count(*) as case_count
# from bc_covid19
# group by age_group
# -
# <div class="alert alert-block alert-success">
#
# Once you get the results, try using the built-in data visualizer to plot the data as a **bar chart**
#
# </div>
#
# SQL is great for shaping and aggregating data, but for more advanced processing or visualizations you might need more tools. Using `-o <variable_name>` parameter of the `%%sql` command we can ask for the result of a query to be returned into the notebook as **Pandas dataframe**.
#
# <div class="alert alert-block alert-success">
#
# Let's count the number of cases per day and pull the result from Spark into our notebook:
#
# </div>
# + magic_args="-o df" language="sql"
# select
# reported_date as date,
# count(*) as case_count
# from bc_covid19
# group by Date
# order by Date
# -
# We now have a variable `df` containing the data as Pandas dataframe, and you are free to do with it anything you'd normally do in Jupyter.
#
# <div class="alert alert-block alert-warning">
#
# Note that if you just type `df` in a cell - you will get an error. That's because by default this kernel executes operations in the remore PySpark environment. To access `df` you need to use `%%local` cell command which will execute code in this local Python kernel.
#
# </div>
#
# This environment already comes with some popular plotting libraries pre-installed (like `plotly`, `bokeh`, `mapbox`, etc.), but if your favorite library is missing - you can always `pip install` it from the terminal.
#
# <div class="alert alert-block alert-success">
#
# Let's do some basic plotting:
#
# </div>
# +
# %%local
import plotly.express as px
fig = px.scatter(
df, x="date", y="case_count",
trendline="rolling", trendline_options=dict(window=7),
trendline_color_override="red")
fig.show()
# -
# ---
#
# ## Up Next
# 🎉 Well done so far! 🎉
#
# Now that we covered the basics of root datasets and data exploration - you are ready to move on to the next chapter where we will take a look at <mark>the key feature</mark> of `kamu` - **data collaboration**!
| images/demo/user-home/demo/01 - Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/elliotgunn/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/LS_DS_131_Statistics_Probability_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="IZHmg86oxREO" colab_type="text"
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
# <br></br>
#
# ## *Data Science Unit 1 Sprint 3 Assignment 1*
#
# # Apply the t-test to real data
#
# Your assignment is to determine which issues have "statistically significant" differences between political parties in this [1980s congressional voting data](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records). The data consists of 435 instances (one for each congressperson), a class (democrat or republican), and 16 binary attributes (yes or no for voting for or against certain issues). Be aware - there are missing values!
#
# Your goals:
#
# 1. Load and clean the data (or determine the best method to drop observations when running tests)
# 2. Using hypothesis testing, find an issue that democrats support more than republicans with p < 0.01
# 3. Using hypothesis testing, find an issue that republicans support more than democrats with p < 0.01
# 4. Using hypothesis testing, find an issue where the difference between republicans and democrats has p > 0.1 (i.e. there may not be much of a difference)
#
# Note that this data will involve *2 sample* t-tests, because you're comparing averages across two groups (republicans and democrats) rather than a single group against a null hypothesis.
#
# Stretch goals:
#
# 1. Refactor your code into functions so it's easy to rerun with arbitrary variables
# 2. Apply hypothesis testing to your personal project data (for the purposes of this notebook you can type a summary of the hypothesis you formed and tested)
# + id="93nZqTk3xREn" colab_type="code" colab={}
### YOUR CODE STARTS HERE
voting_url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/voting-records/house-votes-84.data'
# + id="O4oObpOatEVe" colab_type="code" colab={}
import pandas as pd
# + id="l3rnuTEiubnY" colab_type="code" colab={}
# headers
column_headers = ['party','handicapped-infants','water-project-cost-sharing',
'adoption-of-the-budget-resolution','physician-fee-freeze',
'el-salvador-aid','religious-groups-in-schools',
'anti-satellite-test-ban','aid-to-nicaraguan-contras',
'mx-missile','immigration','synfuels-corporation-cutback',
'education-spending','superfund-right-to-sue','crime',
'duty-free-exports','export-administration-act-south-africa']
# + id="8RXrsWt2tFww" colab_type="code" colab={}
voting_data = pd.read_csv(voting_url, names=column_headers)
# + id="cXp94OYOtPVA" colab_type="code" outputId="ae991f33-447f-4085-b8b2-0ee1c44bce97" colab={"base_uri": "https://localhost:8080/", "height": 275}
voting_data.head()
# + [markdown] id="E7ZcywGRy85k" colab_type="text"
# NaNs could mean absent or abstaining
# + id="0sdFNSA4tnwz" colab_type="code" colab={}
# fill ?s with NaN
import numpy as np
voting_data = voting_data.replace('?', np.NaN)
# + id="i8DbzAD9yEGU" colab_type="code" colab={}
# change y = 1.0
# change n = 0.0
voting_data = voting_data.replace({'y':1, 'n':0})
# + id="Q5pM2lXLytJi" colab_type="code" outputId="702ad785-3199-4891-f6c9-b7fb14770e82" colab={"base_uri": "https://localhost:8080/", "height": 275}
voting_data.head()
# + id="n4E50dney4Z-" colab_type="code" colab={}
# split into two datasets
rep = voting_data[voting_data['party'] == 'republican']
dem = voting_data[voting_data['party'] == 'democrat']
# + [markdown] id="z1Dl8ycBzcE3" colab_type="text"
# # 1-sample t-test
# + id="3AZujz4rzyRE" colab_type="code" outputId="bc918128-ded3-4e83-e5a4-f81eb572c8a8" colab={"base_uri": "https://localhost:8080/", "height": 68}
dem['handicapped-infants'].value_counts()
# + id="vrPMvkhr0JPY" colab_type="code" outputId="ecca15b1-60bf-40b0-8c92-4a2509f1c00b" colab={"base_uri": "https://localhost:8080/", "height": 34}
dem['handicapped-infants'].isnull().sum()
# + id="98-TiuFozeUJ" colab_type="code" outputId="6d109671-db21-4aa5-ab3e-74c46f8537c1" colab={"base_uri": "https://localhost:8080/", "height": 34}
from scipy.stats import ttest_1samp
ttest_1samp(dem['handicapped-infants'], 0, nan_policy='omit')
# this is questioning proportion of dems voting yes compared to null of no ('0')
# can i say there is no dem support?
# p value says nope, you can't say that
# + [markdown] id="S5VZyQFQ0nEe" colab_type="text"
# Null: there is 0 dem support for this bill
# alt hypothesis: that there is non-0 dem support
# p-value threshold (confidence level): 95% confidence level (0.05 p value threshold)
#
# Given the result of the above test I would reject the null that dem support is 0 at the 95% significance level.
# + [markdown] id="ZkCUgz97zet_" colab_type="text"
# # 2-sample t-test (for means) example
# + id="ugk4LXYL2GdH" colab_type="code" colab={}
from scipy.stats import ttest_ind, ttest_ind_from_stats, ttest_rel
# + id="H9iehwyF1VAs" colab_type="code" outputId="5a11bf4e-8f86-4c45-bfce-edcd707f84f9" colab={"base_uri": "https://localhost:8080/", "height": 51}
print("Democrat Support: ", dem['export-administration-act-south-africa'].mean())
print("Republican Support: ", rep['export-administration-act-south-africa'].mean())
# + id="UqTN5nya1ze3" colab_type="code" outputId="25fa0c77-2b44-4f64-ed6c-b702ef87deed" colab={"base_uri": "https://localhost:8080/", "height": 34}
ttest_ind(dem['export-administration-act-south-africa'],
rep['export-administration-act-south-africa'],
nan_policy='omit')
# + [markdown] id="9feOu5JR2N6K" colab_type="text"
# is rate of support in dems same as rep?
#
# Null: the mean of dem support == mean of repub support (two parties support the bill at the same rate)
#
# alt: means are different (not the same level of support)
#
# T stat goes up, p value goes down (inversely correlated)
#
# + [markdown] id="wyvrx10928UU" colab_type="text"
# # Assignment
# 2. Using hypothesis testing, find an issue that democrats support more than republicans with p < 0.01
# 3. Using hypothesis testing, find an issue that republicans support more than democrats with p < 0.01
# 4. Using hypothesis testing, find an issue where the difference between republicans and democrats has p > 0.1 (i.e. there may not be much of a difference)
# + [markdown] id="mMCjDK742-Fx" colab_type="text"
# write out hypothesis, write out conclusions
# + [markdown] id="-a9pQAZX4daC" colab_type="text"
# ## Dems support more than repubs
# + id="cMhFg4KJ4fph" colab_type="code" outputId="26aeb7e0-6c5d-422f-cdac-42a9bbcbf5d0" colab={"base_uri": "https://localhost:8080/", "height": 51}
print("Democrat Support: ", dem['aid-to-nicaraguan-contras'].mean())
print("Republican Support: ", rep['aid-to-nicaraguan-contras'].mean())
# + id="ibY79yPs4lrk" colab_type="code" outputId="9d3a3a8a-3d85-4881-c507-ae35eb924d0d" colab={"base_uri": "https://localhost:8080/", "height": 34}
ttest_ind(dem['aid-to-nicaraguan-contras'],
rep['aid-to-nicaraguan-contras'],
nan_policy='omit')
# + [markdown] id="FoLhNYBd4r1u" colab_type="text"
# Null: the mean of dem support == mean of repub support (two parties support the bill at the same rate)
#
# alt: means are different (not the same level of support)
#
# Given the result of the above test I would reject the null that both parties support the same issue at the same rate.
# + [markdown] id="QnajuL7b4ZvM" colab_type="text"
# ## Repubs support more than dems
# + id="mA7Ko6333bQ9" colab_type="code" outputId="57cb074f-f2e4-427a-f480-23b0d73f6a33" colab={"base_uri": "https://localhost:8080/", "height": 275}
voting_data.head()
# + id="TAoJswsp3Lho" colab_type="code" outputId="121e6755-7386-47ac-8eb3-49d29e6d3f8a" colab={"base_uri": "https://localhost:8080/", "height": 51}
print("Democrat Support: ", dem['religious-groups-in-schools'].mean())
print("Republican Support: ", rep['religious-groups-in-schools'].mean())
# + id="R9OiDvC_3nCu" colab_type="code" outputId="4ae62a41-1aae-4c5b-c00d-db91f52a8b63" colab={"base_uri": "https://localhost:8080/", "height": 34}
ttest_ind(dem['religious-groups-in-schools'],
rep['religious-groups-in-schools'],
nan_policy='omit')
# + [markdown] id="_Ynq8xyd32bO" colab_type="text"
# Null: the mean of dem support == mean of repub support (two parties support the bill at the same rate)
#
# alt: means are different (not the same level of support)
#
# Given the result of the above test I would reject the null that both parties support the same issue at the same rate.
#
# + [markdown] id="evRqe1tX4t3I" colab_type="text"
# ## difference between republicans and democrats has p > 0.1
# + id="iotesGaO5T12" colab_type="code" outputId="8894d176-122f-433d-aba1-bc5b4da8f49f" colab={"base_uri": "https://localhost:8080/", "height": 51}
print("Democrat Support: ", dem['water-project-cost-sharing'].mean())
print("Republican Support: ", rep['water-project-cost-sharing'].mean())
# + id="-0LEwcvD4NQy" colab_type="code" outputId="f4a6e478-4d2e-445c-b557-a286c571d53f" colab={"base_uri": "https://localhost:8080/", "height": 34}
ttest_ind(dem['water-project-cost-sharing'],
rep['water-project-cost-sharing'],
nan_policy='omit')
# + [markdown] id="QzstqgWz4-bb" colab_type="text"
# Null: the mean of dem support == mean of repub support (two parties support the bill at the same rate)
#
# alt: means are different (not the same level of support)
#
# Given the result of the above test I would fail to reject the null that both parties support the same issue at the same rate.
| LS_DS_131_Statistics_Probability_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kerim371/xeus-cling-on-colab/blob/main/xeus_cling_on_colab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="C85IKatP-piA" outputId="3d71a7e8-abef-439c-ce83-a9aa27d1c7dc"
# !wget https://repo.anaconda.com/miniconda/Miniconda3-latest-Linux-x86_64.sh
# + id="jBr9_MV--1np"
# !chmod +x Miniconda3-latest-Linux-x86_64.sh
# + colab={"base_uri": "https://localhost:8080/"} id="x_QGrfUE_KDb" outputId="b0ba7120-dd22-4291-b04a-35ec77310cc2"
# !bash ./Miniconda3-latest-Linux-x86_64.sh -b -f -p /usr/local
# + id="um4yEclG_dru"
# !conda install xeus-cling -c conda-forge
| xeus_cling_on_colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import numpy as np
import pandas as pd
import json
# # Experiment results
#
# The experiment was based on the classical POMDP problem - Tiger problem.
#
# ### Tiger Problem
#
# In the problem setting, there are two doors and a tiger. In each run of the problem, the tiger is initialized randomly behind one door and is unknown to the player. The player could take three possible actions for each step - listen, open door 1 and open door 2.
#
# - Listen: by listening, player will heard a sound from either door 1 or 2. However the origin of sound is not guaranteed correct. Listening action results in a tiny cost to player (-1)
# - Open door 1/2: by opening one door, player enters the exit state with either a large reward (+10) if the tiger doesn't stand behind the door, or a even larger penalty (-20) otherwise.
#
#
# ### Experiment method
#
# The experiment programs the Tiger problem setting and the agent using "Linear Value Function Approximation" solving the POMDP. We also added choices of applying three kinds of deceptive mask onto player's observations - 'opposite', 'random', 'probabilistic', together with the original observation model.
#
# - Original observation model(baseline): player has a higher probability (0.85) to hear the true origin of tiger's sound and small chance to hear it wrongly. i.e. [True, False] = [0.85, 0.15]
#
# - 'opposite' mask gives opposite observation from true origin of the sound. [True, False] = [0, 1]
#
# - 'random' mask gives randomly generated observation and is impartial to two doors. [True, False] = [0.5, 0.5]
#
# - 'probabilistic' mask provides a relatively "less correct" observation compared to original. [True, False] = [0.6, 0.6]
#
# Baseline experiment and experiment with 'random' mask was run twice; experiment with 'opposite' and 'probabilistic' mask was run 4 times respectively.
#
# During each 'run', the Tiger problem was played 5000 times; the agent was trained every 9 from 10 epochs and the rest was test epoch. During the test epoch, we records the player's action taken, observation and reward received for every step.
#
# ## Direct results from experiment
#
# For the below experiments, the average undiscounted/discounted return/step results are:
#
# - Baseline:
# - run 1 - undiscounted: 4.90 +- 0.30, discounted: 4.42 +- 0.28
# - run 2 - undiscounted: 4.69 +- 0.33, discounted: 4.41 +- 0.31
#
# - "Opposite":
# - run 1 - undiscounted: -20.47 +- 0.10, discounted: -19.50 +- 0.00
# - run 2 - undiscounted: -20.47 +- 0.10, discounted: -19.50 +- 0.10
# - run 3 - undiscounted: -20.91 +- 0.12, discounted: -19.49 +- 0.10
# - run 4 - undiscounted: -20.47 +- 0.10, discounted: -19.49 +- 0.10
#
# - "Random":
# - run 1 - undiscounted: -9.96 +- 0.54, discounted: -7.61 +- 0.42
# - run 2 - undiscounted: -9.57 +- 0.55, discounted: -7.14 +- 0.42
#
# - "Probabilistic" ($p_{correct} = 0.6$):
# - run 1 - undiscounted: -2.51 +- 0.32, discounted: -2.39 +- 0.30
# - run 2 - undiscounted: -3.69 +- 0.33, discounted: -3.53 +- 0.30
# - run 3 - undiscounted: -2.58 +- 0.32, discounted: -2.47 +- 0.30
# - run 4 - undiscounted: -3.01 +- 0.33, discounted: -2.91 +- 0.31
# ### Look into the stepwise results
#
# We further look into the stepwise outcome for each epoch in each run of experiments.
#
# We parse the verbal records into numerical statistics. Below are related codes, jump to:
#
# - [link to data processing](#processing)
# - [link to results](#results)
# for analysis.
#
#
def parse(path):
results = dict()
with open(path, 'r', encoding='utf-8') as f:
content = f.readlines()
if not content:
raise ValueError("path not valid")
temp_epoch = -1
temp_block = dict()
temp_step = -1
temp_reward = 0
for line in content:
line = line.strip()
if len(line)==0:
temp_block['r'] = temp_reward
if 'fr' not in temp_block:
temp_block['fr'] = temp_reward
if temp_epoch==5000 and len(temp_block)==1:
break
results[temp_epoch] = temp_block
# renew the temps
temp_block = dict()
temp_reward = 0
elif line[:3]=='eva':
temp_epoch = int(line.strip('.').split(' ')[-1])
#print(temp_epoch)
elif line[:2]=='b ':
line = line.split(' ')
temp_step = int(line[1])
b1 = float(line[2].strip('['))
b2 = float(line[3].strip(']'))
temp_block[temp_step] = (b1, b2)
#print(temp_step, b1, b2)
elif line[20:23]=='Rew':
temp_v = float(line.split(' ')[-1])
if temp_v != -1.0:
temp_block['fr'] = temp_v
temp_reward += temp_v
results.pop(-1, None)
return results
# +
path = './temp/tiger_res/'
oppo_fn = ['oppo1', 'oppo2', 'oppo3', 'oppo4']
prob_fn = ['prob1_60','prob2_60','prob3_60','prob1_60']
rand_fn = ['rand1', 'rand2']
valid_fn = ['valid', 'valid2']
# -
def save_json(file_list):
for fn in file_list:
res = parse(path+fn+'.txt')
with open(path+fn+'.json', 'w') as fp:
json.dump(res, fp)
print("Json saved.")
save_json(oppo_fn)
save_json(prob_fn)
save_json(rand_fn)
save_json(valid_fn)
# +
def get_stat(res_dict):
belief_stat = dict()
reward_stat = dict()
for _, epoch in res_dict.items():
for k,v in epoch.items():
if k == 'r':
if v in reward_stat:
reward_stat[v] += 1
else:
reward_stat[v] = 1
else:
if v[0] in belief_stat:
belief_stat[v[0]] += 1
else:
belief_stat[v[0]] = 1
return belief_stat, reward_stat
def get_stat_count(res_dict):
b1_all = []
for _, epoch in res_dict.items():
for k,v in epoch.items():
if k != 'r' and k != 'fr':
b1_all.append(v)
return b1_all
def get_belief_reward(res_dict):
b1_rwds = []
fr_stat = dict()
for k, epoch in res_dict.items():
temp_rew = epoch.pop('r')
temp_fr = epoch.pop('fr')
if temp_fr in fr_stat:
fr_stat[temp_fr] += 1
else:
fr_stat[temp_fr] = 1
for bs in epoch.values():
b1_rwds.append((bs[0], temp_rew))
return b1_rwds, fr_stat
# -
import matplotlib.pyplot as plt
from functools import reduce
# <a id='processing'></a>
#
# ### Explanation
#
# I collect the believes of the player from every step of every testing, together with the resulting rewards at the final step - the exit state which marks whether the player successfully avoid the hazard or not. By combining the belief in the process with the final rewards, we aim to analyze how the rewards distribute over belief.
# ### baseline with no deceptive mask
_, axs = plt.subplots(1, 2, sharey=True, tight_layout=True, figsize=(8,4))
valid_b1_rew = []
for i in range(len(valid_fn)):
temp_res = parse(path+valid_fn[i]+'.txt')
b1_rew, fr_stat = get_belief_reward(temp_res)
msg = "Run {} have "+'{} in {} results; '*len(fr_stat)
print(msg.format(i+1, *list(reduce(lambda x, y: x + y, fr_stat.items()))))
valid_b1_rew += b1_rew
b1_rew = np.asarray(b1_rew)
axs[i].hist2d(b1_rew[:, 0], b1_rew[:, 1], bins=10)
# ### baseline with opposite mask
_, axs = plt.subplots(1, len(oppo_fn), sharey=True, tight_layout=True, figsize=(12,3))
oppo_b1_rew = []
for i in range(len(oppo_fn)):
temp_res = parse(path+oppo_fn[i]+'.txt')
b1_rew, fr_stat = get_belief_reward(temp_res)
msg = "Run {} have "+'{} in {} results; '*len(fr_stat)
print(msg.format(i+1, *list(reduce(lambda x, y: x + y, fr_stat.items()))))
oppo_b1_rew += b1_rew
b1_rew = np.asarray(b1_rew)
axs[i].hist2d(b1_rew[:, 0], b1_rew[:, 1], bins=10)
# ### baseline with probabilistic deceptive mask
_, axs = plt.subplots(1, 4, sharey=True, tight_layout=True, figsize=(12,4))
prob_b1_rew = []
for i in range(len(prob_fn)):
temp_res = parse(path+prob_fn[i]+'.txt')
b1_rew, fr_stat = get_belief_reward(temp_res)
msg = "Run {} have "+'{} in {} results; '*len(fr_stat)
print(msg.format(i+1, *list(reduce(lambda x, y: x + y, fr_stat.items()))))
prob_b1_rew += b1_rew
b1_rew = np.asarray(b1_rew)
axs[i].hist2d(b1_rew[:, 0], b1_rew[:, 1], bins=10)
# ### baseline with random mask
_, axs = plt.subplots(1, len(rand_fn), sharey=True, tight_layout=True, figsize=(8,4))
rand_b1_rew= []
for i in range(len(rand_fn)):
temp_res = parse(path+rand_fn[i]+'.txt')
b1_rew, fr_stat = get_belief_reward(temp_res)
msg = "Run {} have "+'{} in {} results; '*len(fr_stat)
print(msg.format(i+1, *list(reduce(lambda x, y: x + y, fr_stat.items()))))
rand_b1_rew += b1_rew
b1_rew = np.asarray(b1_rew)
axs[i].hist2d(b1_rew[:, 0], b1_rew[:, 1], bins=10)
# ### Aggregate the results from 4 settings and cross comparing
# +
#_, axs = plt.subplots(1, 4, sharey=True, tight_layout=True, figsize=(12,3))
valid_b1_rew = np.asarray(valid_b1_rew)
oppo_b1_rew = np.asarray(oppo_b1_rew)
prob_b1_rew = np.asarray(prob_b1_rew)
rand_b1_rew = np.asarray(rand_b1_rew)
fig = plt.figure(figsize=(15,8))
ax1 = fig.add_subplot(221)
ax2 = fig.add_subplot(222)
ax3 = fig.add_subplot(223)
ax4 = fig.add_subplot(224)
ax1.title.set_text('Baseline')
ax2.title.set_text('Opposite Mask')
ax3.title.set_text('Prob Mask')
ax4.title.set_text('Random Mask')
_ = ax1.hist2d(valid_b1_rew[:, 0], valid_b1_rew[:, 1], bins=10)
_ = ax2.hist2d(oppo_b1_rew[:, 0], oppo_b1_rew[:, 1], bins=10)
_ = ax3.hist2d(prob_b1_rew[:, 0], prob_b1_rew[:, 1], bins=10)
_ = ax4.hist2d(rand_b1_rew[:, 0], rand_b1_rew[:, 1], bins=10)
# -
# <a id='results'></a>
# ## Results
#
# As shown in statistical results, for baseline model with no deception on observation, the agent achieves success (away from tiger) with approximate 90% probability.
#
# When applying probabilistic deceptive mask with 0.6 correct rate, the success rate of the player drops to 62.3%.
#
# When applying randomly deceptive mask, the success rate drops to 50% approximately. This makes sense because the tiger is initialized randomly and impartially behind two doors. Applying random mask is mathematically equivalent to making random guess from two doors.
#
# When applying opposite mask, the player won't survive a single attempt.
#
# ### Regarding the distribution of resulting rewards over belief,
# we see that "opposite" mask serves as a quick and direct deception - the player will simply listen once, get a falsified observation and make wrong selection of the door. It's a short "pain" for the player with no success chance.
#
# The "random" mask deceives the player in another approach. The player still wins at almost 50% chance, which equals to random guess. However, in many runs of the model, the player suffered from "long pain" when they repeatedly tried to listen for the correct sound and yet get puzzled by the falsified feedback generated by random mask. Some runs resulted in an even worse reward/penalty than an direct fail as small costs of listening aggregate large. Some player eventually made max number of steps allowed (50) and was forced to exit.
#
# The "probabilistic" mask works as a "subtle" deception. The distribution of final rewards resembles the one from baseline, yet the player has less chance to success in the end.
# ### histogram 1D - belief for door 1
# +
_, axs = plt.subplots(1, 2, sharey=True, tight_layout=True, figsize=(8,4))
for i in range(len(valid_fn)):
temp_res = parse(path+valid_fn[i]+'.txt')
temp_b1 = np.asarray(get_stat_count(temp_res))
axs[i].hist(temp_b1[:, 0], bins=20)
# -
_, oppo_ax = plt.subplots(1, len(oppo_fn), sharey=True, tight_layout=True, figsize=(12,3))
for i in range(len(oppo_fn)):
temp_res = parse(path+oppo_fn[i]+'.txt')
temp_belief_1 = np.asarray(get_stat_count(temp_res))
oppo_ax[i].hist(temp_belief_1[:,0], bins=20)
_, prob_ax = plt.subplots(1, 4, sharey=True, tight_layout=True, figsize=(14,3))
for i in range(len(prob_fn)):
temp_res = parse(path+prob_fn[i]+'.txt')
temp_belief_1 = np.asarray(get_stat_count(temp_res))
prob_ax[i].hist(temp_belief_1[:,0], bins=20)
_, rand_ax = plt.subplots(1, len(rand_fn), sharey=True, tight_layout=True, figsize=(12,3))
for i in range(len(rand_fn)):
temp_res = parse(path+rand_fn[i]+'.txt')
temp_belief_1 = np.asarray(get_stat_count(temp_res))
rand_ax[i].hist(temp_belief_1[:,0], bins=20)
# Plot alpha vectors
from experiments.scripts import plot_alpha_vectors
from experiments.scripts import pickle_wrapper
import os
# +
n_actions = 3
weight_dir = os.path.join('./experiments', 'pickle_jar')
valid_pkl = ['console.pkl', 'console2.pkl']
for pkl in valid_pkl:
gamma = pickle_wrapper.load_pkl(os.path.join(weight_dir, pkl))
plot_alpha_vectors.plot_alpha_vectors('Alpha vectors computed with linear function approximation', gamma, n_actions)
# -
for oppo in oppo_fn:
gamma = pickle_wrapper.load_pkl(os.path.join(weight_dir, oppo+'.pkl'))
plot_alpha_vectors.plot_alpha_vectors('Alpha vectors computed with linear function approximation', gamma, n_actions)
for rand in rand_fn:
gamma = pickle_wrapper.load_pkl(os.path.join(weight_dir, rand+'.pkl'))
plot_alpha_vectors.plot_alpha_vectors('Alpha vectors computed with linear function approximation', gamma, n_actions)
for prob in prob_fn:
gamma = pickle_wrapper.load_pkl(os.path.join(weight_dir, prob+'.pkl'))
plot_alpha_vectors.plot_alpha_vectors('Alpha vectors computed with linear function approximation', gamma, n_actions)
| experiment_result.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from thesis_initialise import *
import numpy as _np
# -
from dataclasses import dataclass, fields, Field, InitVar
# +
################################################################################
from functools import lru_cache as _lru_cache, reduce as _reduce
import numpy as _np
@dataclass
class ChesterWorld:
nTimes: int = 24
levels: int = 3
popDensity: int = 1500 # persons per km 2
res: float = 0.016 # innermost edge km
init: InitVar(bool) = True
def __post_init__(self, init):
self.maxLevel = (levels := self.levels) - 1
self.nTiles = nTiles = 2 ** (8 * levels)
self.length = length = (res := self.res) * 16 ** levels
self.area = area = length ** 2
self.nAgents = nAgents = round(self.popDensity * area)
self.agentsPerTile = nAgents / nTiles
self.agentIDs = _np.arange(nAgents)
self._hashVal = np.random.randint(10 ** 12, 10 ** 13 - 1)
if init:
self.initialise()
def initialise(self):
shape = (self.nAgents, self.nTimes, self.levels)
self.schedule = _np.random.randint(0, 2**8, shape, 'uint8')
# self.places = _np.zeros(self.nAgents,)
@_lru_cache
def get_potential_contacts(self, agentID: int, level: int = None, /):
level = self.maxLevel if level is None else level
if level:
shortlist = self.get_potential_contacts(agentID, level - 1)
else:
shortlist = self.agentIDs
schedule = self.schedule[:, :, level][shortlist]
matches = schedule == schedule[agentID]
return shortlist[_np.unique(_np.nonzero(matches)[0])]
def __hash__(self):
return self._hashVal
################################################################################
# if level = 0:
# schedule = self.schedule
# else:
# schedule = self.get_potential_contacts(agentID, level - 1)
# schedule = self.schedule[:, :, :level+1]
# contacts = _np.nonzero(schedule == schedule[agentID])[0]
# return sorted(set(contacts))
# def get_contacts(self, agentID: int):
# schedule = self.schedule
# f
# return _reduce(
# self.get_potential_contacts,
# (agentID, )
# )
# -
# %time model = ChesterWorld(nTimes = 1)
# %time contacts = model.get_potential_contacts(0, 0)
# %time contacts = model.get_potential_contacts(0, 1)
# %time contacts = model.get_potential_contacts(0, 2)
print(f"nContacts = {len(contacts)}")
assert all(_np.any(model.schedule[0] == model.schedule[contact]) for contact in contacts)
assert all(_np.any(model.schedule[0, :, 0] == model.schedule[contact, :, 0]) for contact in contacts)
# %time model = ChesterWorld(nTimes = 24)
# %time contacts = model.get_potential_contacts(0, 0)
# %time contacts = model.get_potential_contacts(0, 1)
# %time contacts = model.get_potential_contacts(0, 2)
print(f"nContacts = {len(contacts)}")
assert all(_np.any(model.schedule[0] == model.schedule[contact]) for contact in contacts)
assert all(_np.any(model.schedule[0, :, 0] == model.schedule[contact, :, 0]) for contact in contacts)
self = ChesterWorld(nTimes = 4)
contacts = self.get_potential_contacts(0)
subSchedule = (schedule := self.schedule)[:, :, -1]
contactMatrix = schedule[_np.nonzero(subSchedule[contacts] == subSchedule[0])]
print(self.schedule[0])
print(contactMatrix)
contacts
tracerSchedule = schedule[0]
tracerSchedule
subSchedule[contacts] == subSchedule[0]
_np.argwhere(contactMatrix)
| working/chesterworld_002.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# +
# SANITY CHECK 1
# Are simulated choice probabilities the same that we get from Navarro & Fuss ?
# -
# Import relevant files
import make_data_wfpt as mdw
import ddm_data_simulation as ddm_sim
# Simulate ddm
(_ , sim_choices) = ddm_sim.ddm_simulate_fast(v = 1, a = 1, w = 0.5, n_samples = 20000)
# Calculate choice probabilities
# 1. From simulated data
sum(sim_choices > 0) / len(sim_choices)
# 2. From navarro fuss
mdw.choice_probabilities(v = 1, a = 1, w = 0.5)
| deprecated/wfpt_sanity_checks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Summary
# This notebook is about simple PoC of using basic statistics to detect anomaly. After this, I would incorporate Time Series / Machine learning / Deep learning on anomaly detection
#
#
# Overall Anomaly Detection
#
# - Goal: The AI/ML will detect the anomaly based on deviation in pattern of data from last 30 days
# - How it Works: There will be score given to the anomaly based on below attributes:
# - The criticality of data
# - Minor
# - Major
# - Critical
# - Duration of anomaly
# - Frequency of anomaly in last 30 days
#
# The Anomaly will be stored in MySQL/NoSQL database, with below details:
# - Anomaly id
# - Anomaly first occurrence
# - Anomaly Duration
# - Anomaly Status
# - Involved network parameters
# - Impacted Customers
# - Impacted machines
#
# ## Preparation
# ### Connect Data
# +
import numpy as np
import mysql.connector
import pandas as pd
from pandas_profiling import ProfileReport
# Initiate with Parameters
db_name= "core_stats"
col = "peak_upload_speed"
# Start Database Connection
db_connection = mysql.connector.connect(
host="192.168.3.11",
user="gyan",
password="<PASSWORD>",
database="gyan_db"
)
# Create Database Cursor for SQL Queries
mycursor = db_connection.cursor()
mycursor.execute("SELECT * FROM {} LIMIT 5".format(db_name))
myresult = mycursor.fetchall()
for x in myresult:
print(x)
# Load data from database and store as pandas Dataframe
df = pd.read_sql('SELECT * FROM {}'.format(db_name), con=db_connection)
df.head()
# -
#
# ### Data Cleaning
# Load functions from Jupyter notebook, Could change to .py as Well
# %run helper_functions.ipynb
getSummaryTable(df)
Summary_table= getSummaryTable(df,True)
getDuplicateColumns(df)
temp=df
# # Phrase 1 - Starter Anomaly Detector
# **How it Works?**
# 1. Summarize first, forecast later.
# - Exaplin how a metric is identified as anomaly data point comparing to last 30 days.
#
# 2. Anomaly Detector Category
# - Statistical (Outlier, Z-score)
# - Time Series (Prophet, Arima | Kalman Filter)
# - Machine Learning (Clustering, Classification: Isolation Forest |)
# - Deep Learning (LSTM, Autoencoder | Clockwork RNN, Depth Gated RNN)
#
#
# **How it's different from the Final Product**
# 1. No enough data.
# - As for adjustments, I'll use moving window 7 days instead of 30 days for PoC.
# - e.g. What data are anomlies comparing to historical 7 (30) days
#
#
# 2. No enough column.
# - We simply want to focus on the most important column first.
# - However, the function is designed to apply to more columns in type aspect.
#
#
# 3. Not Focusing on Quality, but automated process.
#
# 4. Individual labels or Scores instead of an Overall Score.
#
# 5. Keep comparison with Exiting 3rd Party Anomaly Detection tools like Anodot.
#
# ## Stats Calculations
#
# - Mean
# - Median
# - IQR
# - Outlier: x> Q3+1.5IQR or x < Q1-1.5IQR
#
#
# **Business Explanation**
# Big jump are going on going down on upload or download speed.
# We can identify there's a big jump in the
#
# WHen the jump happends, we send an alert.
#
#
# **Column of interest:**
# - Peak download speed
# ### Basic Summary Stats
# +
col= "peak_upload_speed"
df[col].describe()
# -
# ### Z - Score
#
# - Function Details
# - Input: dataframe, one column
# - Output:
# - One Score Column
# - e.g. Z-score (negative 3 to 3 )-> Adjusted Z-score ( 0~1)
# - One Label Column
#
#
add_Z_score_column(df,col)
# ### Outlier
add_outlier_column(df,col)
# ## Automation
# ### Apply Stats Functions for All column in one Database
keeped_column_name = list(Summary_table["Columns_Name"])
print("We keep columns: ", keeped_column_name,"\n")
keeped_column_name.remove("client_id")
# +
# Create a new table to store all the computed metrics
Stats_summary_core = df
client_id_list= list(df.client_id.unique())
Stats_summary_core_new= pd.DataFrame()
for clientID in client_id_list:
temp_df = Stats_summary_core[Stats_summary_core["client_id"]==clientID]
for col in keeped_column_name:
temp_df=add_Z_score_column(temp_df,col)
temp_df=add_outlier_column(temp_df,col)
Stats_summary_core_new=Stats_summary_core_new.append(temp_df, ignore_index = True)
filter_col = [col for col in Stats_summary_core if col.startswith('label')]
# -
Stats_summary_core= Stats_summary_core_new
# Find percentage of labels that's been marked as anomalies, 0.01 means 1%
np.round(Stats_summary_core[filter_col].sum()/Stats_summary_core.shape[0],2)
# ### Export to CSVs
export_full_option=False
if export_full_option == True:
Anomaly_summary = Stats_summary_core[Stats_summary_core[filter_col].sum(axis=1)>=1].reset_index(drop=True)
Anomaly_summary.to_csv("Anomaly_summary.csv")
# +
import os.path
# folder_path = "C:\Users\<NAME>\Desktop\Main_Anomaly_Detection\Generated_csv\"
for col in keeped_column_name:
# Filter columns with at least one record that have anomaly label
condition= (Stats_summary_core["label_Z-score_"+ col]!=0) | (Stats_summary_core["label_outlier_"+col]!=0)
subset_columns=["client_id","stats_timestamp",col,"label_Z-score_"+col,"label_outlier_"+col]
print("{} rows of anomaly detected for column {}".format(sum(condition),col))
subset_Summary= Stats_summary_core[condition][subset_columns]
csv_filename = "Anomaly_{}_summary.csv".format(col)
if os.path.exists(csv_filename):
print("File {} already generated".format(csv_filename))
pass
else:
subset_Summary.to_csv(csv_filename,index=False)
print("Done for:", col,"\n")
# -
export_anomaly_df= pd.DataFrame()
# +
# folder_path = "C:\Users\<NAME>\Desktop\Main_Anomaly_Detection\Generated_csv\"
export_anomaly_df= pd.DataFrame()
for col in keeped_column_name:
# Filter columns with at least one record that have anomaly label
condition= (Stats_summary_core["label_Z-score_"+ col]!=0) | (Stats_summary_core["label_outlier_"+col]!=0)
subset_columns=["client_id","stats_timestamp",col,"label_Z-score_"+col,"label_outlier_"+col]
print("{} rows of anomaly detected for column {}".format(sum(condition),col))
subset_Summary= Stats_summary_core[condition][subset_columns]
export_anomaly_df
print("Done for:", col,"\n")
# -
def reorder_columns(dataframe, col_name, position):
"""Reorder a dataframe's column.
Args:
dataframe (pd.DataFrame): dataframe to use
col_name (string): column name to move
position (0-indexed position): where to relocate column to
Returns:
pd.DataFrame: re-assigned dataframe
"""
temp_col = dataframe[col_name]
dataframe = dataframe.drop(columns=[col_name])
dataframe.insert(loc=position, column=col_name, value=temp_col)
return dataframe
# +
# Filter columns with at least one record that have anomaly label
for col in keeped_column_name:
condition= (Stats_summary_core["label_Z-score_"+ col]!=0) | (Stats_summary_core["label_outlier_"+col]!=0)
subset_columns=["client_id","stats_timestamp",col,"label_Z-score_"+col,"label_outlier_"+col]
print("{} rows of anomaly detected for column {}".format(sum(condition),col))
subset_Summary= Stats_summary_core[condition][subset_columns]
subset_Summary["Attribute_Name"] = col
subset_Summary = reorder_columns(subset_Summary,"Attribute_Name",2)
subset_Summary=subset_Summary.rename(columns={str(col):"Attribute_Value","label_Z-score_"+col: "Attribute_Label_Z_Score", "label_outlier_"+col: "Attribute_Label_Outlier"})
export_anomaly_df=export_anomaly_df.append(subset_Summary, ignore_index = True)
# -
export_anomaly_df.to_csv("export_anomaly_df.csv")
# ### Anomaly Per Day
Anomaly_summary=Stats_summary_core
# +
ans_time_delta= max(df.stats_timestamp)-min(df.stats_timestamp)
print("Q: What's the number of days for data we have?","\nA:",ans_time_delta)
num_days = ans_time_delta.days
ans_avg_anomaly_by_day = np.round(Anomaly_summary.shape[0]/num_days,1)
print("Q: How many records per day is classified as anomaly by at least one label?","\nA:",ans_avg_anomaly_by_day)
for col in keeped_column_name:
condition= (Stats_summary_core["label_Z-score_"+ col]!=0) | (Stats_summary_core["label_outlier_"+col]!=0)
subset_columns=["client_id","stats_timestamp",col,"label_Z-score_"+col,"label_outlier_"+col]
subset_Summary= Stats_summary_core[condition][subset_columns]
ans_col_avg_anomaly_by_day= np.round(subset_Summary.shape[0] / num_days,2)
#print("\n")
print(col)
print("Q: How many Total Anomaly Occurs per day?","\nA:",ans_col_avg_anomaly_by_day)
# -
# ### Addtional Summary
# - Anoamly for which Column
# - Client
# - Anomaly Time
#
# **Data Structure**
# ### Automate Process for Anomly Experimentation
def experiment_Stats(z_score=3,iqr=1.5, num_days=14):
Stats_summary_core = df
for col in keeped_column_name:
Stats_summary_core=add_Z_score_column(Stats_summary_core,col,z_score_threshold=z_score)
Stats_summary_core=add_outlier_column(Stats_summary_core,col, iqr_factor=iqr)
print("Anomaly Detected Rocord per Day")
print(np.round(Stats_summary_core[filter_col].sum()/num_days,2))
print("\n")
print("Anomaly Percentage")
print(np.round(Stats_summary_core[filter_col].sum()/Stats_summary_core.shape[0],2))
experiment_Stats(4,2)
filter_col_Z = [col for col in Stats_summary_core if col.startswith('label_Z')]
Stats_summary_core[filter_col_Z].head(3)
Stats_summary_core.head(3)
matrix_names= filter_col_Z
Stats_summary_matrix = Stats_summary_core[filter_col_Z].to_numpy()
filter_col
# ### Scoring for Different Types of Anomalies
Stats_summary_df= pd.DataFrame(Stats_summary_matrix*[12,13,14,14,13,11,14,11])
score_list = list(range(0,110,10))
value_list= []
# +
for score in score_list:
v= (Stats_summary_df.sum(axis=1)>score).sum()
value_list.append(v)
value_list
# -
Stats_summary_df.loc[Stats_summary_df.sum(axis=1)>10,]
Stats_summary_df.loc[Stats_summary_df.sum(axis=1)>40,].head(3)
Stats_summary_df.loc[Stats_summary_df.sum(axis=1)>30,].head(3)
theory_data_point = round(24*60/5)
df.shape[0]/14*0.01
24*60/5*0.01
# ### Create Database and Insert Core stats
new_db= db_name+"_Anomaly_Summary"
# mycursor.execute("CREATE DATABASE {}".format(new_db))
# # Phrase 2: Business & Analytics and Automation
# # Phrase 3: Advanced Anomaly Detection Methods
# ## Time Series
# ## Functions need exploring
# +
#https://github.com/Vicam/Unsupervised_Anomaly_Detection/blob/master/custom_function.py
#from pyemma import msm
import pandas as pd
import numpy as np
# return Series of distance between each point and his distance with the closest centroid
def getDistanceByPoint(data, model):
distance = pd.Series()
for i in range(0,len(data)):
Xa = np.array(data.loc[i])
Xb = model.cluster_centers_[model.labels_[i]-1]
distance.set_value(i, np.linalg.norm(Xa-Xb))
return distance
# train markov model to get transition matrix
# def getTransitionMatrix (df):
# df = np.array(df)
# model = msm.estimate_markov_model(df, 1)
# return model.transition_matrix
# return the success probability of the state change
def successProbabilityMetric(state1, state2, transition_matrix):
proba = 0
for k in range(0,len(transition_matrix)):
if (k != (state2-1)):
proba += transition_matrix[state1-1][k]
return 1-proba
# return the success probability of the whole sequence
def sucessScore(sequence, transition_matrix):
proba = 0
for i in range(1,len(sequence)):
if(i == 1):
proba = successProbabilityMetric(sequence[i-1], sequence[i], transition_matrix)
else:
proba = proba*successProbabilityMetric(sequence[i-1], sequence[i], transition_matrix)
return proba
# return if the sequence is an anomaly considering a threshold
def anomalyElement(sequence, threshold, transition_matrix):
if (sucessScore(sequence, transition_matrix) > threshold):
return 0
else:
return 1
# return a dataframe containing anomaly result for the whole dataset
# choosing a sliding windows size (size of sequence to evaluate) and a threshold
def markovAnomaly(df, windows_size, threshold):
transition_matrix = getTransitionMatrix(df)
real_threshold = threshold**windows_size
df_anomaly = []
for j in range(0, len(df)):
if (j < windows_size):
df_anomaly.append(0)
else:
sequence = df[j-windows_size:j]
sequence = sequence.reset_index(drop=True)
df_anomaly.append(anomalyElement(sequence, real_threshold, transition_matrix))
return df_anomaly
# +
import numpy as np
import matplotlib.pyplot as plt
# multiply and add by random numbers to get some real values
data = np.random.randn(50000) * 20 + 20
# Function to Detection Outlier on one-dimentional datasets.
def find_anomalies(data):
#define a list to accumlate anomalies
anomalies = []
# Set upper and lower limit to 3 standard deviation
data_std = np.std(data)
data_mean = np.mean(data)
anomaly_cut_off = data_std * 3
lower_limit = data_mean - anomaly_cut_off
upper_limit = data_mean + anomaly_cut_off
print(lower_limit)
# Generate outliers
for outlier in data:
if outlier > upper_limit or outlier < lower_limit:
anomalies.append(outlier)
return anomalies
find_anomalies(data)
| Stage1_Summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting started with K8
#
# After years of talking about it, I've started taking the time to really dig into a k8 environment. Having spent most of my career working with AWS. I've opt'd to learn more about GCP this time around. With it, I'm extending my vocabulary and looking into certifications. In the meantime, I'll make regular updates for moving from a Cloud Deployment strategy to Kubernetes.
# To begin, lets become oriented with how to access k8 pods. K8 pods are one of the more common K8 datatypes. They provide a compute resource that'll execute a docker image. Either as a Job, Deployment, CronJob, etc. Pods pretty much do everything inside a K8 cluster.
#
# ```
# $ kubectl get pods --namespace=default
# NAME READY STATUS RESTARTS AGE
# cassandra-cassandra-0 1/1 Running 0 45m
# redis-master-0 1/1 Running 0 10h
# ```
#
# Already installed in the cluster, we have [Apache Cassandra](https://cassandra.apache.org) and [Redis](https://redis.io). To deploy the instances, we're using [helm](https://helm.sh/).
#
# Cassandra and Redis are running as a single instance. We're not to worried about availability with this cluster mainly do to it being a utility for scheduled [CronJobs](https://kubernetes.io/docs/concepts/workloads/controllers/cron-jobs/). I'm still interested in monitoring it, lets go ahead and deploy DataDog using helm.
#
# ```
# $ helm install data-dog -f values/data-dog.yaml datadog/datadog
# NAME: data-dog
# LAST DEPLOYED: Wed Nov 25 23:34:04 2020
# NAMESPACE: default
# STATUS: deployed
# REVISION: 1
# TEST SUITE: None
# NOTES:
# Datadog agents are spinning up on each node in your cluster. After a few
# minutes, you should see your agents starting in your event stream:
# https://app.datadoghq.com/event/stream
#
# ```
#
# Lets check to see how many new pods we have running
#
# ```
# $ $ kubectl get pods --namespace=default
# NAME READY STATUS RESTARTS AGE
# data-dog-datadog-rgckw 2/2 Running 0 40m
# data-dog-kube-state-metrics-8546d8989-rgm4w 1/1 Running 0 40m
# ```
#
# Looking good. Lets clean up our cluster a little bit before we finish.
#
# ```
# $ kubectl delete pod --field-selector=status.phase==Succeeded
# ```
#
| notebooks/k8-getting-started/k8-getting-started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# In this tutorial, you'll learn how to investigate data types within a DataFrame or Series. You'll also learn how to find and replace entries.
#
# **To start the exercise for this topic, please click [here](#$NEXT_NOTEBOOK_URL$).**
#
# # Dtypes
#
# The data type for a column in a DataFrame or a Series is known as the **dtype**.
#
# You can use the `dtype` property to grab the type of a specific column. For instance, we can get the dtype of the `price` column in the `reviews` DataFrame:
#$HIDE_INPUT$
import pandas as pd
reviews = pd.read_csv("../input/wine-reviews/winemag-data-130k-v2.csv", index_col=0)
pd.set_option('max_rows', 5)
reviews.price.dtype
# Alternatively, the `dtypes` property returns the `dtype` of _every_ column in the DataFrame:
reviews.dtypes
# Data types tell us something about how pandas is storing the data internally. `float64` means that it's using a 64-bit floating point number; `int64` means a similarly sized integer instead, and so on.
#
# One peculiarity to keep in mind (and on display very clearly here) is that columns consisting entirely of strings do not get their own type; they are instead given the `object` type.
#
# It's possible to convert a column of one type into another wherever such a conversion makes sense by using the `astype()` function. For example, we may transform the `points` column from its existing `int64` data type into a `float64` data type:
reviews.points.astype('float64')
# A DataFrame or Series index has its own `dtype`, too:
reviews.index.dtype
# Pandas also supports more exotic data types, such as categorical data and timeseries data. Because these data types are more rarely used, we will omit them until a much later section of this tutorial.
# # Missing data
#
# Entries missing values are given the value `NaN`, short for "Not a Number". For technical reasons these `NaN` values are always of the `float64` dtype.
#
# Pandas provides some methods specific to missing data. To select `NaN` entries you can use `pd.isnull()` (or its companion `pd.notnull()`). This is meant to be used thusly:
reviews[pd.isnull(reviews.country)]
# Replacing missing values is a common operation. Pandas provides a really handy method for this problem: `fillna()`. `fillna()` provides a few different strategies for mitigating such data. For example, we can simply replace each `NaN` with an `"Unknown"`:
reviews.region_2.fillna("Unknown")
# Or we could fill each missing value with the first non-null value that appears sometime after the given record in the database. This is known as the backfill strategy.
# Alternatively, we may have a non-null value that we would like to replace. For example, suppose that since this dataset was published, reviewer <NAME> has changed her Twitter handle from `@kerinokeefe` to `@kerino`. One way to reflect this in the dataset is using the `replace()` method:
reviews.taster_twitter_handle.replace("@kerinokeefe", "@kerino")
# The `replace()` method is worth mentioning here because it's handy for replacing missing data which is given some kind of sentinel value in the dataset: things like `"Unknown"`, `"Undisclosed"`, `"Invalid"`, and so on.
#
# # Your turn
#
# If you haven't started the exercise, you can **[get started here](#$NEXT_NOTEBOOK_URL$)**.
| notebooks/pandas/raw/tut_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Let's make it more idiomatic
# Your task is to refactor the following report generation code to more idiomatic. The existing implementation was written by an unknown developer who did not know anything about the idioms of Python. Luckily the unkown developer documented the implementation decently and wrote some tests for it.
#
# ### The specification of the report generation
#
# This file content:
# ```
# something
# 1
# 7
# somEThing
#
# 2
# wassup
# woop
# woop
# something
# WoOP
# ```
#
# Should yield this report:
# ```
# missing values: 1
# highest number: 7.0
# most common words: something, woop
# occurrences of most common: 3
# #####
# numbers: [1.0, 7.0, 2.0]
# words: ['something', 'something', 'wassup', 'woop', 'woop', 'something', 'woop']
# ```
#
# Note:
# * all numbers of the input file should be presented as floats in the report
# * all words are presented as lowercased in the report
# * while calculating the most common words, the count should be done as case insensitive (in other words, `'WoOp'` should be considered the same word as `'woop'`)
# * if there are multiple different most common words, they should be presented in the format presented above
# * there are more examples in the tests
#
# Run the cell of the existing implementation and then run the tests to verify that it works correctly. Then make sure that you understand how the legacy implementation works. After that, start refactoring, a function by function. Good luck!
# + editable=false
def get_report(path):
"""
Creates a report of the file specified as argument.
:param path: path to file from which the report should be created (string)
:return: the report (string)
"""
data = _read_file(path)
missing_count = data[0]
numbers = data[1]
words = data[2]
report = _make_report(missing_count, numbers, words)
return report
def _read_file(path):
"""
Reads and returns the data from the file specified as argument.
:param path: path to the file to be read.
:return: a tuple containing
1. the number of empty lines (int)
2. numeric values (list of floats)
3. non-numeric values (list of strings)
"""
data_file = open(path, 'r')
lines = data_file.readlines()
line_count = len(lines)
idx = 0
empty_lines = 0
words = []
numbers = []
while idx < line_count:
line = lines[idx]
line = line.strip()
if line == '':
empty_lines = empty_lines + 1
else:
is_number = False
try:
number = float(line)
is_number = True
except Exception:
pass
if is_number:
numbers.append(number)
else:
words.append(line)
idx = idx + 1
data_file.close()
return empty_lines, numbers, words
def _make_report(missing_values, numbers, words):
"""
Creates and a report based on data given as arguments.
:param missing_values: number of empty lines (int)
:param numbers: numeric values (list of floats)
:param words: non numeric values (list of strings)
:return: the generated report (string)
"""
max_value = _get_max_value(numbers)
lower_case_words = _words_to_lowercase(words)
most_common_info = _get_most_common_words(lower_case_words)
most_common_words = most_common_info[0]
most_common_count = most_common_info[1]
most_common_str = ''
for idx in range(len(most_common_words)):
most_common_str += most_common_words[idx] + ', '
# remove the last comma and space
most_common_str = most_common_str[0:len(most_common_str) - 2]
report = ('missing values: {}\n'
'highest number: {}\n'
'most common words: {}\n'
'occurrences of most common: {}\n'
'#####\n'
'numbers: {}\n'
'words: {}').format(missing_values, max_value, most_common_str,
most_common_count, numbers, lower_case_words)
return report
def _get_max_value(numbers):
"""
Returns the greatest value of the list given as argument.
:param numbers: numbers (list of numeric values)
:return: greatest value of numbers, None if numbers is an empty list
"""
max_value = None
if len(numbers) > 0:
max_value = numbers[0]
for idx in range(len(numbers)):
if numbers[idx] > max_value:
max_value = numbers[idx]
return max_value
def _words_to_lowercase(words):
"""
:param words: words to be converted (list of strings)
:return: lowercased words (list of strings)
"""
lowercased = []
for idx in range(len(words)):
value = words[idx].lower()
lowercased.append(value)
return lowercased
def _get_most_common_words(words):
"""
Finds the most common words in a list of words.
If there are multiple different words with the same amount of occurrences,
they are all included in the return value sorted alphabetically.
In addition to returning the most common words, the return value
includes also the count of occurrences of the most common words.
:param words: list of words (list of strings)
:return: a tuple containing:
1. most common words (list of strings)
2. the count of occurrences of the most common words (int)
"""
word_counts = {}
idx = 0
while idx < len(words):
value = words[idx]
if value not in word_counts.keys():
word_counts[value] = 1
else:
word_counts[value] += 1
idx = idx + 1
max_count = 0
for value in word_counts.values():
if value > max_count:
max_count = value
most_common_words = []
for word in word_counts.keys():
count = word_counts[word]
if count == max_count:
most_common_words.append(word)
most_common_words = sorted(most_common_words)
return most_common_words, max_count
# -
# Now it's time refactor the existing code to make it more idiomatic.
#
# It's desirable that you do the refactoring in small chunks. Consider using the following workflow:
# 1. Copy-paste a single function from the above ugly implementation to the cell below
# 2. Refactor the function
# 3. Run the tests to verify that you did not break anything
#
# This way you can consider each function as a separate sub task.
# +
# Your beautiful refactored, idiomatic, pythonic solution here
# -
# The tests are here. Run these often while refactoring!
# + editable=false
import os
CURRENT_DIR = os.getcwd()
DATA_DIR = os.path.join(os.path.dirname(CURRENT_DIR), 'data')
DATA_FILE1 = os.path.join(DATA_DIR, 'misc_data1.txt')
DATA_FILE2 = os.path.join(DATA_DIR, 'misc_data2.txt')
DATA_FILE3 = os.path.join(DATA_DIR, 'empty.txt')
expected1 = '''missing values: 2
highest number: 99.0
most common words: john
occurrences of most common: 4
#####
numbers: [1.0, 2.0, 99.0, 6.72, 2.0, 2.0, 2.0]
words: ['john', 'doe', 'john', 'john', 'was', 'here', 'this', 'is', 'totally', 'random', 'john']'''
expected2 = '''missing values: 3
highest number: 101.0
most common words: doe, john
occurrences of most common: 4
#####
numbers: [1.0, 2.0, 101.0, 6.72, 2.0, 2.0, 67.0, 2.0]
words: ['john', 'doe', 'john', 'john', 'doe', 'was', 'doe', 'here', 'this', 'is', 'totally', 'random', 'john', 'doe']'''
expected3 = '''missing values: 0
highest number: None
most common words:
occurrences of most common: 0
#####
numbers: []
words: []'''
assert get_report(DATA_FILE1) == expected1
print('First one OK!')
assert get_report(DATA_FILE2) == expected2
print('Second one OK!')
assert get_report(DATA_FILE3) == expected3
print('All OK, woop woop!')
# +
# If the tests are failing, you can debug here.
report = get_report(DATA_FILE1)
print(report)
| notebooks/intermediate/exercises/idiomatic_python_exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .venv_calfews2
# language: python
# name: .venv_calfews2
# ---
import numpy as np
import pandas as pd
import h5py
import json
from itertools import compress
from datetime import datetime
from calfews_src import *
from calfews_src.util import *
import ipywidgets as widgets
from IPython.display import display, clear_output, FileLink
from bqplot import pyplot as plt
from bqplot import LinearScale, DateScale, Lines, Axis, Figure
from bqplot.interacts import BrushSelector, BrushIntervalSelector, panzoom
import threading
import time
import tornado
import warnings
import os
from datetime import datetime
warnings.filterwarnings('ignore')
# +
### flags for command line, diff for windows vs unix
if os.name == 'nt':
mkdir_os = 'mkdir '
sep_os = '\\'
else:
mkdir_os = 'mkdir -p '
sep_os = '/'
### flip slash
def slash_os(s):
return s.replace('/', sep_os)
#######################################################################
### set up functions
#######################################################################
### function to run simulation using chosen options
def run_sim(progress_w, ioloop):
global sim_finished, main_cy_obj
start_time = datetime.now()
print('#######################################################')
print('Begin initialization...')
### delete old results, then recreate folder to store new results
# !rm -rf {slash_os(temp_results_sim + filname)}
# !{mkdir_os} {slash_os(temp_results_sim + filname)}
### initialize model
import main_cy
main_cy_obj = main_cy.main_cy(temp_results_sim + filname, model_mode, flow_input_type, flow_input_source)
print('Initialization complete, ', datetime.now() - start_time)
### setup thread to continuously check model progress, end when model finishes
def update_progress(progress, progress_w=progress_w):
progress_w.value = progress
def check_progress_thread():
### update progress bar asynchronously
while not sim_finished.isSet():
sim_finished.wait(5)
ioloop.add_callback(update_progress, main_cy_obj.progress)
t_check_progress = threading.Thread(target = check_progress_thread)
t_check_progress.start()
### run model
main_cy_obj.run_sim_py(start_time)
print ('Simulation complete,', datetime.now() - start_time)
main_cy_obj.calc_objectives()
main_cy_obj.output_results()
print ('Data output complete,', datetime.now() - start_time)
### move results to permanent storage (if different)
if temp_results_sim != permanent_results:
# !cp -r {temp_results_sim + filname} {permanent_results}
print('Copying data to ' + permanent_results + filname)
print('#######################################################')
print('Finished! Proceed to "Plot data" tab.')
### function to deal with different label in gui label vs dataset
def get_alternative_label(output_type, bin_is_attribute, new_column, old_column, old_label):
if output_type in ['District'] and bin_is_attribute == 1 and old_column == 'Object Attributes':
### check if label contains Contract/District/Waterbank
try:
name, old_att = old_label.split('_', 1)
if name in results_keys['Contract'][0]['Object List Model'].values.tolist():
new_name = get_alternative_label('Contract', 0, 'Object Name', 'Object List Model', name)
new_att = get_alternative_label('District', 1, 'Label', 'Object Attributes', '*contract_' + old_att)
return new_name + ': ' + new_att
elif name in results_keys['District'][0]['Object Key'].values.tolist():
new_name = get_alternative_label('District', 0, 'Object Name', 'Object Key', name)
new_att = get_alternative_label('District', 1, 'Label', 'Object Attributes', '*district_' + old_att)
return new_name + ': ' + new_att
elif name in results_keys['Waterbank'][0]['Object Key'].values.tolist():
new_name = get_alternative_label('Waterbank', 0, 'Object Name', 'Object Key', name)
new_att = get_alternative_label('District', 1, 'Label', 'Object Attributes', '*waterbank_' + old_att)
return new_name + ': ' + new_att
else:
pass
except:
# ### check if key is Private/District with bank account at this District
if old_label in results_keys['Private'][0]['Object Key'].values.tolist():
new_name = get_alternative_label('Private', 0, 'Object Name', 'Object Key', old_label)
new_att = get_alternative_label('District', 1, 'Label', 'Object Attributes', '*bank_member_key')
return new_name + ': ' + new_att
if old_label in results_keys['District'][0]['Object Key'].values.tolist():
new_name = get_alternative_label('District', 0, 'Object Name', 'Object Key', old_label)
new_att = get_alternative_label('District', 1, 'Label', 'Object Attributes', '*bank_member_key')
return new_name + ': ' + new_att
else:
pass
elif output_type in ['District'] and bin_is_attribute == 1 and old_column == 'Label':
### check if label containts Contract/District/Waterbank
try:
name, old_att = old_label.split(': ', 1)
if name in results_keys['Contract'][0]['Object Name'].values.tolist():
new_name = get_alternative_label('Contract', 0, 'Object List Model', 'Object Name', name)
new_att = get_alternative_label('District', 1, 'Object Attributes', 'Label', old_att)
if new_att == '*bank_member_key':
return get_alternative_label('Contract', 0, 'Object Key', 'Object List Model', new_name)
else:
return new_att.replace('*contract', new_name)
elif name in results_keys['District'][0]['Object Name'].values.tolist():
new_name = get_alternative_label('District', 0, 'Object Key', 'Object Name', name)
new_att = get_alternative_label('District', 1, 'Object Attributes', 'Label', old_att)
if new_att == '*bank_member_key':
return new_name
else:
return new_att.replace('*district', new_name)
elif name in results_keys['Waterbank'][0]['Object Name'].values.tolist():
new_name = get_alternative_label('Waterbank', 0, 'Object Key', 'Object Name', name)
new_att = get_alternative_label('District', 1, 'Object Attributes', 'Label', old_att)
if new_att == '*bank_member_key':
return new_name
else:
return new_att.replace('*waterbank', new_name)
elif name in results_keys['Private'][0]['Object Name'].values.tolist():
new_name = get_alternative_label('Private', 0, 'Object Key', 'Object Name', name)
new_att = get_alternative_label('District', 1, 'Object Attributes', 'Label', old_att)
if new_att == '*bank_member_key':
return new_name
else:
return new_att.replace('*private', new_name)
else:
pass
except:
pass
elif output_type in ['Private'] and bin_is_attribute == 1 and old_column == 'Object Attributes':
### check if label contains Contract/District/Waterbank
try:
name1, old_att1 = old_label.split('_', 1)
if name1 in results_keys['District'][0]['Object Key'].values.tolist():
new_name1 = get_alternative_label('District', 0, 'Object Name', 'Object Key', name1)
### check if attribute contains Contract/District/Waterbank
try:
name2, old_att2 = old_att1.split('_', 1)
if name2 in results_keys['Contract'][0]['Object List Model'].values.tolist():
new_name2 = get_alternative_label('Contract', 0, 'Object Name', 'Object List Model', name2)
new_att2 = get_alternative_label('Private', 1, 'Label', 'Object Attributes', '*location_*contract_' + old_att2)
return new_name1 + ': ' + new_name2 + ': ' + new_att2
elif name2 in results_keys['District'][0]['Object Key'].values.tolist():
new_name2 = get_alternative_label('District', 0, 'Object Name', 'Object Key', name2)
new_att2 = get_alternative_label('Private', 1, 'Label', 'Object Attributes', '*location_*district_' + old_att2)
return new_name1 + ': ' + new_name2 + ': ' + new_att2
elif name2 in results_keys['Waterbank'][0]['Object Key'].values.tolist():
new_name2 = get_alternative_label('Waterbank', 0, 'Object Name', 'Object Key', name2)
new_att2 = get_alternative_label('Private', 1, 'Label', 'Object Attributes', '*location_*waterbank_' + old_att2)
return new_name1 + ': ' + new_name2 + ': ' + new_att2
else:
pass
except:
pass
except:
pass
elif output_type in ['Private'] and bin_is_attribute == 1 and old_column == 'Label':
### first should be key for District associated with Private water account
try:
name1, old_att1 = old_label.split(': ', 1)
if name1 in results_keys['District'][0]['Object Name'].values.tolist():
new_name1 = get_alternative_label('District', 0, 'Object Key', 'Object Name', name1)
## check if attribute contains Contract/District/Waterbank
try:
name2, old_att2 = old_att1.split(': ', 1)
if name2 in results_keys['Contract'][0]['Object Name'].values.tolist():
new_name2 = get_alternative_label('Contract', 0, 'Object List Model', 'Object Name', name2)
new_att2 = get_alternative_label('Private', 1, 'Object Attributes', 'Label', old_att2)
return new_att2.replace('*location_*contract', new_name1 + '_' + new_name2)
elif name2 in results_keys['District'][0]['Object Name'].values.tolist():
new_name2 = get_alternative_label('District', 0, 'Object Key', 'Object Name', name2)
new_att2 = get_alternative_label('Private', 1, 'Object Attributes', 'Label', old_att2)
return new_att2.replace('*location_*district', new_name1 + '_' + new_name2)
elif name2 in results_keys['Waterbank'][0]['Object Name'].values.tolist():
new_name2 = get_alternative_label('Waterbank', 0, 'Object Key', 'Object Name', name2)
new_att2 = get_alternative_label('Private', 1, 'Object Attributes', 'Label', old_att2)
return new_att2.replace('*location_*waterbank', new_name1 + '_' + new_name2)
else:
pass
except:
pass
except:
pass
elif output_type in ['Waterbank'] and bin_is_attribute == 1 and old_column == 'Object Attributes':
### check if label contains Contract/District/Waterbank
try:
# ### check if key is Private/District with bank account at this District
if old_label in results_keys['District'][0]['Object Key'].values.tolist():
new_name = get_alternative_label('District', 0, 'Object Name', 'Object Key', old_label)
new_att = get_alternative_label('Waterbank', 1, 'Label', 'Object Attributes', '*bank_member_key')
return new_name + ': ' + new_att
elif old_label in results_keys['Private'][0]['Object Key'].values.tolist():
new_name = get_alternative_label('Private', 0, 'Object Name', 'Object Key', old_label)
new_att = get_alternative_label('Waterbank', 1, 'Label', 'Object Attributes', '*bank_member_key')
return new_name + ': ' + new_att
else:
pass
except:
pass
elif output_type in ['Waterbank'] and bin_is_attribute == 1 and old_column == 'Label':
### check if label containts Contract/District/Waterbank
try:
name, old_att = old_label.split(': ', 1)
if name in results_keys['District'][0]['Object Name'].values.tolist():
new_name = get_alternative_label('District', 0, 'Object Key', 'Object Name', name)
new_att = get_alternative_label('Waterbank', 1, 'Object Attributes', 'Label', old_att)
if new_att == '*bank_member_key':
return new_name
else:
pass
elif name in results_keys['Private'][0]['Object Name'].values.tolist():
new_name = get_alternative_label('Private', 0, 'Object Key', 'Object Name', name)
new_att = get_alternative_label('Waterbank', 1, 'Object Attributes', 'Label', old_att)
if new_att == '*bank_member_key':
return new_name
else:
pass
else:
pass
except:
pass
labels_df = results_keys[output_type][bin_is_attribute]
try:
new_label = labels_df[new_column].loc[labels_df[old_column] == old_label].values[0]
return new_label
except:
return None
def plot_data_initial():
global fig, fig_zoom
xdata = []
ydata = []
labels = []
xsc = DateScale()
ysc = LinearScale()
k1 = output_type_w.value
for output in output_split2_w.value:
if output_order_w.value == output_orders[k1][0]:
obj_key = get_alternative_label(k1, 0, 'Object List', 'Object Name', output_split1_w.value)
att_key = get_alternative_label(k1, 1, 'Object Attributes', 'Label', output)
else:
obj_key = get_alternative_label(k1, 0, 'Object List', 'Object Name', output)
att_key = get_alternative_label(k1, 1, 'Object Attributes', 'Label', output_split1_w.value)
key = obj_key + '_' + att_key
xdata.append(datDaily.index)
ydata.append(datDaily[key])
labels.append(output)
lines = Lines(x=xdata, y=ydata, scales={'x':xsc, 'y':ysc}, labels=labels, display_legend=True)
axx = Axis(scale=xsc, label='Date')
title = output_split1_w.value
axy = Axis(scale=ysc, orientation='vertical')
### set up first plot with brushing capability
brush_selector_w = BrushIntervalSelector(scale=xsc, marks=[lines])
fig = Figure(marks=[lines], axes=[axx, axy], interaction=brush_selector_w, title=title, legend_location=leg_loc)
fig.legend_style = {'stroke-width': 0}
# ### set up second plot, which will be zoomed to box from first plot
xsc_zoom = DateScale()
ysc_zoom = LinearScale()
lines_zoom = Lines(x=xdata, y=ydata, scales={'x':xsc_zoom, 'y':ysc_zoom}, labels=labels, display_legend=True)
axx_zoom = Axis(scale=xsc_zoom, label='Date')
axy_zoom = Axis(scale=ysc_zoom, orientation='vertical')
fig_zoom = Figure(marks=[lines_zoom], axes=[axx_zoom, axy_zoom], title=title, legend_location=leg_loc)
fig_zoom.legend_style = {'stroke-width': 0}
def brush_bounds_update(b):
if brush_selector_w.selected is not None:
xsc_zoom.min, xsc_zoom.max = brush_selector_w.selected
else:
xsc_zoom.min, xsc_zoom.max = np.min(fig_zoom.marks[0].x), np.max(fig_zoom.marks[0].x)
fig_zoom.axes[0].scale = xsc_zoom
brush_selector_w.observe(brush_bounds_update, names=['brushing'])
plot_html3_w = widgets.HTML(value = '<b>Figure 1: Full time series (click and drag to select zoom region for Figure 2)</b><br />\n')
plot_html4_w = widgets.HTML(value = '<b>Figure 2: Zoomed in time series</b><br />\n')
display(widgets.VBox([plot_html3_w, fig, plot_html4_w, fig_zoom]))
def plot_data_output_type(output_type):
global output_split2_w
output_split2_w.value = [output_split2_w.options[0]]
def plot_data_output_split1(output_split1):
global fig, fig_zoom
xdata = []
ydata = []
labels = []
k1 = output_type_w.value
for output in output_split2_w.value:
if output_order_w.value == output_orders[k1][0]:
obj_key = get_alternative_label(k1, 0, 'Object List', 'Object Name', output_split1['new'])
att_key = get_alternative_label(k1, 1, 'Object Attributes', 'Label', output)
else:
obj_key = get_alternative_label(k1, 0, 'Object List', 'Object Name', output)
att_key = get_alternative_label(k1, 1, 'Object Attributes', 'Label', output_split1['new'])
key = obj_key + '_' + att_key
xdata.append(datDaily.index)
ydata.append(datDaily[key])
labels.append(output)
fig.marks[0].x = xdata
fig.marks[0].y = ydata
fig.marks[0].labels = labels
fig_zoom.marks[0].x = xdata
fig_zoom.marks[0].y = ydata
fig_zoom.marks[0].labels = labels
title = output_split1['new']
fig.title = title
fig_zoom.title = title
def plot_data_output_split2(output_split2):
global fig, fig_zoom
xdata = []
ydata = []
labels = []
k1 = output_type_w.value
for output in output_split2['new']:
if output_order_w.value == output_orders[k1][0]:
obj_key = get_alternative_label(k1, 0, 'Object List', 'Object Name', output_split1_w.value)
att_key = get_alternative_label(k1, 1, 'Object Attributes', 'Label', output)
else:
obj_key = get_alternative_label(k1, 0, 'Object List', 'Object Name', output)
att_key = get_alternative_label(k1, 1, 'Object Attributes', 'Label', output_split1_w.value)
key = obj_key + '_' + att_key
xdata.append(datDaily.index)
ydata.append(datDaily[key])
labels.append(output)
fig.marks[0].x = xdata
fig.marks[0].y = ydata
fig.marks[0].labels = labels
fig_zoom.marks[0].x = xdata
fig_zoom.marks[0].y = ydata
fig_zoom.marks[0].labels = labels
def update_tab_display(widget):
# get the correct Output widget based on the index of the chosen tab
tab_idx = widget['new']
output_widget = dynamic_outputs[tab_idx]
with output_widget:
clear_output()
tab_dispatcher(tab_idx)
def tab_intro():
clear_output()
intro_html = '<b>Graphical User Interface for the California Food-Energy-Water System (CALFEWS) simulation model</b><br />\n'
intro_html += '<p>This tool provides a user interface for the <a href="https://github.com/hbz5000/CALFEWS">California Food-Energy-Water System (CALFEWS)</a>, an open-sourced, Python-based model for simulating the integrated, multi-sector dynamics of water supply in the Central Valley of California. CALFEWS captures system dynamics across multiple scales, from coordinated management of inter-basin water supply projects at the state and regional scale, to agent-based representation of conjunctive surface water and groundwater supplies at the scale of irrigation and water storage districts. This user interface can be used to simulate operations under different hydrologic conditions and visualize the results using the interactive plotting features.</p><br />\n'
intro_html += '<p>More information on the CALFEWS model, and comparison of model output to historical data, can be found in the following manuscript:</p><br />\n'
intro_html += '<p><NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., and <NAME>. (2020). "California\'s Food-Energy-Water System: An Open Source Simulation Model of Adaptive Surface and Groundwater Management in the Central Valley". (In review, <a href="https://engrxiv.org/sqr7e/")>preprint available here</a>.)</p><br />\n'
intro_html_w = widgets.HTML(value = intro_html)
display(intro_html_w)
def tab_simulation():
global model_mode, flow_input_type, flow_input_source, filname_w, filname, sim_finished
### This tab will allow user to run a new dataset
clear_output()
### add instructions as html
sim_html1_w = widgets.HTML(value = '<b>Select inflow scenario</b><br />\n')
# display(sim_html1_w)
### get input file scenarios from key file
input_key = pd.read_excel('gui_key.xlsx', sheet_name='input_data', engine="openpyxl")
input_file_options = {input_key['Inflow scenario'][i]: input_key['model_mode'][i] + ':' + input_key['flow_input_type'][i] + ':' + input_key['flow_input_source'][i] for i in range(input_key.shape[0])}
### select widget for input scenario
input_select_w = widgets.Select(options = list(input_file_options.keys()))
input_select_w.value = input_select_w.options[0]
### table explaining where data came from
def make_sim_html_table(headers, values):
yield '<head><style>table, th, td {border: 1px solid black;}table.center {margin-left: auto;margin-right: auto;}th, td {padding: 6px;}</style></head><body><table>'
yield '<tr><th>'
yield '</th><th>'.join(headers)
yield '</th></tr>'
for sublist in values:
yield '<tr><td>'
yield '</td><td>'.join(sublist)
yield '</td></tr>'
yield '</table></body>'
sim_table = [input_key.iloc[i, :3].values.tolist() for i in range(input_key.shape[0])]
sim_table = [[str(item) for item in l] for l in sim_table]
sim_html_table = ''.join(make_sim_html_table(input_key.columns[:3], sim_table))
table_layout = widgets.Layout(max_height='308px', overflow_y='auto')#width='70%',
sim_html_table_w = widgets.HTML(value = sim_html_table, layout=table_layout)
### choose filename for saving data
sim_html2_w = widgets.HTML(value = '<b>Folder name to save results (no spaces)</b><br />\n')
filname = input_select_w.value.replace(' ', '_')
model_mode, flow_input_type, flow_input_source = input_file_options[input_select_w.value].split(':')
filname_w = widgets.Text(value = filname)
### monitor folder name and replace spaces with '_'
def update_filname(fil):
global filname
filname = fil['new'].replace(' ', '_')
filname_w.observe(update_filname, 'value')
### update default folder name when new scenario chosen
def input_update(input_select):
global model_mode, flow_input_type, flow_input_source, filname_w
filname_w.value = input_select['new'].replace(' ', '_')
model_mode, flow_input_type, flow_input_source = input_file_options[input_select['new']].split(':')
input_select_w.observe(input_update, 'value')
### setup progress bar for simulation
progress_w = widgets.FloatProgress(value=0, min=0, max=1)
sim_finished = threading.Event()
def run_sim_thread(*args, **kwargs):
t_run_sim = threading.Thread(target=run_sim, args=(progress_w, tornado.ioloop.IOLoop.instance()))
t_run_sim.start()
### This button will start simulation and make plotting button available afterwards
sim_html3_w = widgets.HTML(value = '<b>Click to begin</b><br />\n')
button = widgets.Button(description='Start')
button.on_click(run_sim_thread)
# display(button)
### print progress bar
sim_html4_w = widgets.HTML(value = '<b>Progress</b><br />\n')
# print('Progress:')
# display(progress_w)
### stack widgets for display
sim_left_display_w = widgets.VBox([sim_html1_w, input_select_w, sim_html2_w, filname_w, sim_html3_w, button, sim_html4_w, progress_w])
sim_left_display_w.layout.min_width = '310px'
sim_display_w = widgets.HBox([sim_left_display_w, sim_html_table_w])
sim_display_w.layout.grid_gap = '12px'
display(sim_display_w)
def tab_plotting_dynamic():
clear_output()
with static_outputs[2]:
global data_w, output_plot_w
### This tab will allow user to visualize their data
clear_output()
### add instructions as html
plot_html = '<b>Select dataset</b><br />\n'
plot_html_w = widgets.HTML(value = plot_html)
display(plot_html_w)
# folders = !ls {permanent_results}
try:
folders.remove('.ipynb_checkpoints')
except:
pass
data_w = widgets.Select(options=folders, value=None)
display(data_w)
# time.sleep(0.1)
# data_w.value = None
data_w.observe(get_data, 'value')
def tab_dispatcher(tab_idx):
if tab_idx == 0:
pass
elif tab_idx == 1:
tab_simulation()
elif tab_idx == 2:
tab_plotting_dynamic()
### function to get all possible output splits for plotting
def get_output_splits():
global results_keys, output_objects, output_split1s, output_split2s, colsplit1, colsplit2
### get results attribute keys
results_keys = {}
for k1 in ['Reservoir','Delta','Contract','District','Private','Waterbank']:
results_keys[k1] = {}
### get objects within class (e.g. shasta for Reservoir)
objects = pd.read_excel('gui_key.xlsx', sheet_name=k1, engine="openpyxl")
### strip whitespace
for c in objects.columns:
try:
objects[c] = [s.strip() for s in objects[c]]
except:
pass
results_keys[k1][0] = objects
### get attributes within class (e.g. storage for Reservoir)
attributes = pd.read_excel('gui_key.xlsx', sheet_name= k1 + '_attributes', engine="openpyxl")
### only include attributes of interest
attributes = attributes.loc[attributes['Include'] == 1, :]
### strip whitespace
for c in objects.columns:
try:
objects[c] = [s.strip() for s in objects[c]]
except:
pass
### add units to label
attributes['Label'] = attributes['Label'] + ' (' + attributes['Attribute Unit'] + ')'
results_keys[k1][1] = attributes
output_objects = {}
for k1 in output_types:
output_objects[k1] = results_keys[k1][0]['Object Name'].values.tolist()
### get options for dropdown 2 based on choice of dropdown 1
cols = datDaily.columns
colsplit = [c.split('_', 1) for c in cols]
colsplit1 = [c[0] for c in colsplit]
colsplit2 = [c[1] for c in colsplit]
output_split1s = {}
for k1 in output_types:
output_split1s[k1] = {}
for output_order in output_orders[k1]:
output_split1s[k1][output_order] = {}
if output_order == output_orders[k1][0]:
l = []
for k2 in output_objects[k1]:
obj_short = get_alternative_label(k1, 0, 'Object List', 'Object Name', k2)
ind = [c == obj_short for c in colsplit1]
if sum(ind) > 0:
l.append(k2)
output_split1s[k1][output_order] = l
else:
l = []
for k2 in output_objects[k1]:
obj_short = get_alternative_label(k1, 0, 'Object List', 'Object Name', k2)
ind = [c == obj_short for c in colsplit1]
if sum(ind) > 0:
ltemp = [get_alternative_label(k1, 1, 'Label', 'Object Attributes', ival) for i,ival in enumerate(colsplit2) if ind[i]]
l.append([item for item in ltemp if item])
l = [item for sublist in l for item in sublist]
lunique = []
for item in l:
if item not in lunique:
lunique.append(item)
output_split1s[k1][output_order] = lunique
### get options for dropdown 3 based on choice of dropdown 1&2
output_split2s = {}
for k1 in output_types:
output_split2s[k1] = {}
for output_order in output_orders[k1]:
output_split2s[k1][output_order] = {}
for k2 in output_split1s[k1][output_order]:
if output_order == output_orders[k1][0]:
obj_short = get_alternative_label(k1, 0, 'Object List', 'Object Name', k2)
ind = [c == obj_short for c in colsplit1]
if sum(ind) > 0:
ltemp = [get_alternative_label(k1, 1, 'Label', 'Object Attributes', ival) for i,ival in enumerate(colsplit2) if ind[i]]
output_split2s[k1][output_order][k2] = [item for item in ltemp if item]
else:
att_short = get_alternative_label(k1, 1, 'Object Attributes', 'Label', k2)
ind = [c == att_short for c in colsplit2]
if sum(ind) > 0:
ltemp = [get_alternative_label(k1, 0, 'Object Name', 'Object List', ival) for i,ival in enumerate(colsplit1) if ind[i]]
output_split2s[k1][output_order][k2] = [item for item in ltemp if item]
### function to retrieve dataset
def get_data(data):
global datDaily, output_type_w, output_split2_w, output_split1_w, output_order_w, output_orders, results_keys, output_types
with dynamic_outputs[2]:
clear_output()
### load data once sim finished
# results hdf5 file location from CALFEWS simulations
output_folder_sim = permanent_results + data['new'] + '/'
got_data = False
# try:
### copy results from permanent storage to app file tree (if different)
if permanent_results != temp_results_plt:
# !cp -r {output_folder_sim} {temp_results_plt}
print('Loading data from ' + output_folder_sim)
datDaily = get_results_sensitivity_number_outside_model(temp_results_plt + data['new'] + '/results.hdf5', '')
got_data = True
# except:
# print('Enter folder name with valid results')
if got_data:
### create interactive dropdown selectors for which data to plot
output_types = ['Reservoir', 'Delta', 'Contract', 'District', 'Private', 'Waterbank']
output_orders = {'Reservoir': ['Choose reservoir, compare attributes', 'Choose attribute, compare reservoirs'],
'Delta': ['Compare delta attributes'],
'Contract': ['Choose contract, compare attributes', 'Choose attribute, compare contracts'],
'District': ['Choose district, compare attributes', 'Choose attribute, compare districts'],
'Private': ['Choose private, compare attributes', 'Choose attribute, compare privates'],
'Waterbank': ['Choose waterbank, compare attributes', 'Choose attribute, compare waterbank']}
### get possible plotting combinations for dropdown menu
get_output_splits()
### interactive dropdown menu for data selection
plot_html1_w = widgets.HTML(value = '<b>Select subset of data for figures</b><br />\n')
output_type_w = widgets.Dropdown(options = output_types)
output_order_w = widgets.Dropdown(options = output_orders[output_type_w.value])
output_split1_w = widgets.Dropdown(options = output_split1s[output_type_w.value][output_order_w.value])
output_split2_w = widgets.SelectMultiple(options = output_split2s[output_type_w.value][output_order_w.value][output_split1_w.value], value=[output_split2s[output_type_w.value][output_order_w.value][output_split1_w.value][0]])
### create buttons for data download
plot_html2_w = widgets.HTML(value = '<b>Download full dataset or subset of data</b><br />\n')
### button to save full data as csv and trigger download link
def create_download_full(b):
### get alternative label for each column in datDaily
labels = []
include = []
for c in datDaily.columns:
name, att = c.split('_', 1)
for obj_type in ['Reservoir','Delta','Contract','District','Private','Waterbank']:
if name in results_keys[obj_type][0]['Object List'].values:
try:
new_name = get_alternative_label(obj_type, 0, 'Object Name', 'Object List', name)
new_att = get_alternative_label(obj_type, 1, 'Label', 'Object Attributes', att)
labels.append(new_name + ': ' + new_att)
include.append(True)
break
except:
include.append(False)
datTemp = datDaily.loc[:, include]
datTemp.columns = labels
### make temp folder to hold results (has to be inside temporary instance of tool, FileLink can't access home directory since it is "upstream" of jupyter home dir)
# !{mkdir_os} temp
datTemp.to_csv('temp/results_full.csv')
full_download = FileLink('temp/results_full.csv', result_html_prefix="Click for full dataset: ")
display(full_download)
### This button will start simulation and make plotting button available afterwards
button_download_full = widgets.Button(description='Full dataset')
button_download_full.on_click(create_download_full)
### button to save full data as csv and trigger download link
def create_download_subset(b):
keys = []
labels = []
k1 = output_type_w.value
for output in output_split2_w.value:
if output_order_w.value == output_orders[k1][0]:
obj_key = get_alternative_label(k1, 0, 'Object List', 'Object Name', output_split1_w.value)
att_key = get_alternative_label(k1, 1, 'Object Attributes', 'Label', output)
else:
obj_key = get_alternative_label(k1, 0, 'Object List', 'Object Name', output)
att_key = get_alternative_label(k1, 1, 'Object Attributes', 'Label', output_split1_w.value)
keys.append(obj_key + '_' + att_key)
labels.append(output_split1_w.value + ': ' + output)
datDaily_subset = datDaily.loc[:, keys]
datDaily_subset.columns = labels
### make temp folder to hold results (has to be inside temporary instance of tool, FileLink can't access home directory since it is "upstream" of jupyter home dir)
# !{mkdir_os} temp
datDaily_subset.to_csv('temp/results_subset.csv')
subset_download = FileLink('temp/results_subset.csv', result_html_prefix="Click for subsetted dataset: ")
display(subset_download)
### This button will start simulation and make plotting button available afterwards
button_download_subset = widgets.Button(description='Subsetted dataset')
button_download_subset.on_click(create_download_subset)
buttons_download_w = widgets.HBox([button_download_full, button_download_subset])
### box for displaying data selection
plot_top_display_w = widgets.VBox([plot_html1_w, output_type_w, output_order_w, output_split1_w, output_split2_w])
display(plot_top_display_w)
def update_output_order_w(output_type):
global output_order_w
output_order_w.options = output_orders[output_type['new']]
# output_order_w.value = [output_order_w.options[0]]
output_type_w.observe(update_output_order_w, 'value')
def update_output_split1_w(output_order):
global output_split1_w
output_split1_w.options = output_split1s[output_type_w.value][output_order['new']]
# output_split1_w.value = [output_split1s_w.options[0]]
output_order_w.observe(update_output_split1_w, 'value')
def update_output_split2_w(output_split1):
global output_split2_w
output_split2_w.options = output_split2s[output_type_w.value][output_order_w.value][output_split1['new']]
output_split2_w.value = [output_split2_w.options[0]]
output_split1_w.observe(update_output_split2_w, 'value')
### plot data using initial selection
plot_data_initial()
### observe selection and replot when anything changes
output_type_w.observe(plot_data_output_type, 'value')
output_order_w.observe(plot_data_output_type, 'value')
output_split1_w.observe(plot_data_output_split1, 'value')
output_split2_w.observe(plot_data_output_split2, 'value')
### box for displaying plot options
plot_bottom_display_w = widgets.VBox([plot_html2_w, buttons_download_w])
display(plot_bottom_display_w)
### create folder for results in user's storage space
temp_results_sim = 'results'
temp_results_plt = 'results'
permanent_results = "results"
if not os.path.isdir(permanent_results):
# !{mkdir_os} {slash_os(permanent_results)}
if temp_results_sim != permanent_results:
# !{mkdir_os} {slash_os(temp_results_sim)}
if temp_results_plt != permanent_results:
# !{mkdir_os} {slash_os(temp_results_plt)}
temp_results_sim += '/'
temp_results_plt += '/'
permanent_results += '/'
# legend location
leg_loc = 'top-left'
# set up a dictionary of Output widgets
static_outputs = {i: widgets.Output() for i in range(0,3)}
dynamic_outputs = {i: widgets.Output() for i in range(0,3)}
output_vbox = static_outputs.copy()
output_vbox[0] = widgets.VBox([static_outputs[0], dynamic_outputs[0]])
output_vbox[1] = widgets.VBox([static_outputs[1], dynamic_outputs[1]])
output_vbox[2] = widgets.VBox([static_outputs[2], dynamic_outputs[2]])
# add the Output widgets as tab childen
tab = widgets.Tab()
tab.children = list(output_vbox.values())
tab_labels = ['About', 'Run simulation', 'Plot data']
for i, title in static_outputs.items():
tab.set_title(i, tab_labels[i])
with static_outputs[0]: tab_intro()
tab.observe(update_tab_display, names='selected_index')
display(tab)
# -
| CALFEWS_GUI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import streamlit as st
from streamlit_multipage import MultiPage
def input_page(st, **state):
st.title("Body Mass Index")
weight_ = state["weight"] if "weight" in state else 0.0
weight = st.number_input("Your weight (Kg): ", value=weight_)
height_ = state["height"] if "height" in state else 0.0
height = st.number_input("Your height (m): ", value=height_)
if height and weight:
MultiPage.save({"weight": weight, "height": height})
def compute_page(st, **state):
st.title("Body Mass Index")
if "weight" not in state or "height" not in state:
st.warning("Enter your data before computing. Go to the Input Page")
return
weight = state["weight"]
height = state["height"]
st.metric("BMI", round(weight / height ** 2, 2))
app = MultiPage()
app.st = st
app.add_app("Input Page", input_page)
app.add_app("BMI Result", compute_page)
app.run()
# -
| Script/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using and analyzing foldlabel files
# # Imports and definitions
import numpy as np
from soma import aims
import anatomist.notebook as ana
a = ana.Anatomist()
print(a.headless_info.__dict__)
src_dir="/neurospin/dico/data/deep_folding/current/datasets/hcp/crops/2mm/CINGULATE/mask/Rlabels"
skel_dir="/neurospin/dico/data/deep_folding/current/datasets/hcp/crops/2mm/CINGULATE/mask/Rcrops"
subject="100206"
# Defines the function to plot the volume in anatomist
def plot_sagittal(vol):
global a
print(vol.header())
a_vol = a.toAObject(vol)
sagittal = a.createWindow('Sagittal')
sagittal.addObjects(a_vol)
return sagittal
foldlabel_file = f"{src_dir}/{subject}_cropped_foldlabel.nii.gz"
skeleton_file = f"{skel_dir}/{subject}_cropped_skeleton.nii.gz"
# # Checks and plots
vol = aims.read(foldlabel_file)
vol_skel = aims.read(skeleton_file)
win1 = plot_sagittal(vol)
# # General computation
arr_foldlabel = np.asarray(vol)
arr_skel = np.asarray(vol_skel)
arr_foldlabel.shape
branches, nb_per_branch = np.unique(arr_foldlabel, return_counts=True)
print(branches)
print(nb_per_branch)
print(f"nb of branches = {branches.shape[0]}")
print(type(nb_per_branch))
nb_per_branch[nb_per_branch>6].sum()-nb_per_branch[0]
np.unique(arr_skel, return_counts=True)
arr_skel.shape
# Counts non-zero values
(arr_foldlabel != 0).sum()
(arr_skel != 0).sum()
diff = arr_skel[arr == 0]
np.count_nonzero(diff)
diff_inverse = arr_foldlabel[arr_skel == 0]
np.count_nonzero(diff_inverse)
nb_per_branch
# # Algorithm to remove branches
branches.size
np.random.randint(0,52)+1
def select_one_random_branch(arr_foldlabel):
"""It selects randomly one of the branch
The branch is characterized by a number.
This number is present on several pixels in the array"""
branches = np.unique(arr_foldlabel)
nb_branches = branches.size
# 0 is not a branch
selected_branch = np.random.randint(0,nb_branches-1)+1
return branches[selected_branch]
def remove_branch(arr_foldlabel, arr_skel, selected_branch):
"""It masks the selected branch in arr_skel
"""
# print((arr_foldlabel > 0).sum())
mask = ( (arr_foldlabel != 0) & (arr_foldlabel != selected_branch))
mask = mask.astype(int)
# print(mask.sum())
return arr_skel * mask
select_one_random_branch(arr_foldlabel)
selected_branch=1005
arr_skel_without_branch = remove_branch(arr_foldlabel, arr_skel, selected_branch)
# Expected 23
np.count_nonzero(arr_skel)-np.count_nonzero(arr_skel_without_branch)
selected_branch=1311
arr_skel_without_branch = mask_branch(arr_foldlabel, arr_skel, selected_branch)
# Expected 1
np.count_nonzero(arr_skel)-np.count_nonzero(arr_skel_without_branch)
# We want to check which branches are not represented at all in skeleton.
# Indeed, there are 111 pixels in foldlabel that have a 0 value correspondence
# in skeleton
branches = np.unique(arr_foldlabel)
print(branches)
indexes = np.arange(branches.size-1)+1
np.random.shuffle(indexes)
indexes
histo_foldlabel = np.unique(arr_foldlabel, return_counts=True)
histo_foldlabel = np.stack((histo_foldlabel),axis=-1)
for branch in branches:
if branch != 0:
arr_skel_without_branch = mask_branch(arr_foldlabel, arr_skel, branch)
diff = np.count_nonzero(arr_skel)-np.count_nonzero(arr_skel_without_branch)
if diff == 0:
index = np.argwhere(histo_foldlabel[:,0]==branch)
print(histo_foldlabel[index])
# We note and conclude that very few branches have no correspondence with skeleton
# # Full program to remove branches up to the point for which a certain percentage of pixels have been removed
def remove_branches_up_to_percent(arr_foldlabel, arr_skel, percentage):
"""Removes from arr_skel random branches up percentage of pixels
"""
branches = np.unique(arr_foldlabel)
# We take as index branches indexes that are not 0
indexes = np.arange(branches.size-1) + 1
# We take random branches
np.random.shuffle(indexes)
arr_skel_without_branches = arr_skel
total_pixels = (arr_skel !=0 ).sum()
total_pixels_after=total_pixels
for index in indexes:
if total_pixels_after <= total_pixels*(100-percentage)/100:
break
arr_skel_without_branches = \
remove_branch(arr_foldlabel,
arr_skel_without_branches,
branches[index])
total_pixels_after = (arr_skel_without_branches != 0).sum()
print(f"total_pixels_after = {total_pixels_after}")
print(f"% removed pixels = {(total_pixels-total_pixels_after)/total_pixels*100}")
return arr_skel_without_branches
arr_skel_without_branches = remove_branches_up_to_percent(arr_foldlabel, arr_skel, 10)
| contrastive/notebooks/18_use_foldlabel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import numpy
from tensorflow.keras.preprocessing.image import ImageDataGenerator,array_to_img,img_to_array, load_img
import tensorflow as tf
from tensorflow import keras
from keras.models import Sequential
from keras.models import Model
from keras.layers import Conv2D,MaxPooling2D,Activation,Dropout,Flatten,Dense,BatchNormalization
from keras.applications.vgg19 import VGG19, preprocess_input
import keras.backend as K
from sklearn.model_selection import train_test_split
from tensorflow.keras import Input, Model
import PIL
from PIL import Image,ImageOps
from PIL import ImageDraw
import shutil
import os
from os import listdir
from PIL import Image
import xml.etree.ElementTree as ET
from tqdm import tqdm
import seaborn as sns
import string
import nltk
from nltk.tokenize import word_tokenize
import pickle
import re
from numpy import savez_compressed
# Tag Prediction
train=pd.read_csv('traindata.csv',nrows=20)
test=pd.read_csv('testdata.csv',nrows=10)
train.head()
train.drop('Unnamed: 0',axis='columns',inplace=True)
test.drop('Unnamed: 0',axis='columns',inplace=True)
train.head()
test.head()
with open('train_features_list_new.pkl', 'rb') as f:
train_feat_list = pickle.load(f)
with open('test_features_list_new.pkl', 'rb') as f:
test_feat_list = pickle.load(f)
train_feat_list_new = []
for i in range(0,20):
train_feat_list_new.append(train_feat_list[i])
test_feat_list_new = []
for i in range(0,10):
test_feat_list_new.append(test_feat_list[i])
len(train_feat_list_new), len(test_feat_list_new)
df_tags = pd.read_csv("tags.csv")
df_tags.head()
df_tags.reset_index(drop=True, inplace=True)
df_new_2 = pd.DataFrame(train.merge(df_tags, on='patient_id', how='left'))
df_new_2.describe()
df_new_2.head()
df_new_2.isnull().sum()
df_new_2.head()
df_new_2['tags'].replace('\d+', '', regex=True, inplace=True)
df_new_2.head()
df_new_2.to_csv('df_coatt.csv')
# +
def create_tag_mapping(mapping_csv):
labels = set()
for i in range(len(mapping_csv)):
tags = mapping_csv['tags'][i].split(' ')
labels.update(tags)
labels = list(labels)
labels.sort()
labels_map = {labels[i]:i for i in range(len(labels))}
inv_labels_map = {i:labels[i] for i in range(len(labels))}
return labels_map, inv_labels_map
def create_file_mapping(mapping_csv):
mapping = dict()
for i in range(len(mapping_csv)):
name, tags = mapping_csv['image1_id'][i], mapping_csv['tags'][i]
mapping[name] = tags.split(' ')
return mapping
def one_hot_encode(tags, mapping):
encoding = np.zeros(len(mapping), dtype='uint8')
for tag in tags:
encoding[mapping[tag]] = 1
return encoding
def load_dataset(path, file_mapping, tag_mapping):
photos, targets = list(), list()
for i in file_mapping:
for filename in os.listdir(folder):
image_name = os.path.splitext(filename)[0]
if(i==image_name):
photo = Image.open(path + image_name + '.png')
photo=img_to_array(photo)
photo = preprocess_input(photo)
photo=cv2.resize(photo,(128,128))
photo = img_to_array(photo, dtype='uint8')
tags = file_mapping[image_name]
target = one_hot_encode(tags, tag_mapping)
photos.append(photo)
targets.append(target)
X = np.asarray(photos, dtype='uint8')
y = np.asarray(targets, dtype='uint8')
return X, y
filename = 'df_coatt.csv'
mapping_csv = pd.read_csv(filename)
tag_mapping, _ = create_tag_mapping(mapping_csv)
file_mapping = create_file_mapping(mapping_csv)
folder = 'C:/Users/INDIRA/Downloads/data rescon/images/'
X, y = load_dataset(folder, file_mapping, tag_mapping)
print(X.shape, y.shape)
savez_compressed('data_coatt.npz', X, y)
# -
def load_dataset():
data = np.load('data_coatt.npz')
X, y = data['arr_0'], data['arr_1']
trainX, trainY = X, y
return trainX, trainY
trainX.shape , trainY.shape
def define_model(in_shape=(128, 128, 3), out_shape=30):
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same', input_shape=in_shape))
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu', kernel_initializer='he_uniform'))
model.add(Dense(out_shape, activation='softmax'))
# compile model
opt = tf.keras.optimizers.SGD(lr=0.001, momentum=0.9)
model.compile(optimizer=opt, loss='binary_crossentropy', metrics='accuracy')
return model
datagen = ImageDataGenerator(rescale=1.0/255.0)
trainX, trainY = load_dataset()
train_it = datagen.flow(trainX, trainY, batch_size=5)
model = define_model()
history = model.fit_generator(train_it, steps_per_epoch=len(train_it), epochs=10, verbose=0)
loss, fbeta = model.evaluate_generator(train_it, steps=len(train_it), verbose=0)
print('> loss=%.3f, fbeta=%.3f' % (loss, fbeta))
yhat = model.predict(trainX)
yhat.shape
yhat_list = []
yhat_list = yhat.max(axis=1)
len(yhat_list)
yhat_list
labels = np.argmax(yhat, axis=-1)
labels
_, inv_tag_mapping = create_tag_mapping(mapping_csv)
inv_tag_mapping
semantic_feat = []
for keys in inv_tag_mapping.keys():
for item in labels:
if item == keys:
semantic_feat.append(inv_tag_mapping[keys])
semantic_feat
tag_mapping, _ = create_tag_mapping(mapping_csv)
semantic_encoding = one_hot_encode(semantic_feat,tag_mapping)
semantic_feat_encoding = list(semantic_encoding)
# Semantic Features
len(semantic_feat_encoding)
co_att_feat = []
# Visual Features
len(train_feat_list_new)
# Concatenating two features
co_att_feat = train_feat_list_new + semantic_feat_encoding
len(co_att_feat)
co_att_feat_arr = x = np.asarray(co_att_feat)
co_att_feat_arr.shape
type(co_att_feat_arr)
# Co-attention layer to generate context vector
inp = Input(shape=co_att_feat_arr.shape[0])
hidden = Dense(8, activation='relu')(inp)
flat = Flatten()(hidden)
out = Dense(2)(flat)
model = Model(inputs=inp, outputs=out)
| co-attention.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# +
import random
import matplotlib.pyplot as plt
random.seed()
result_ratio_arrays = []
# +
def get_roll(die_size):
if die_size<1:
do_negative=True
die_size=2
roll1 = random.randint(1, 6)
roll2 = random.randint(1,6)
return min(roll1, roll2)
maxRoll=0
sixCount=0
for i in range(die_size):
roll = random.randint(1, 6)
if roll > maxRoll:
maxRoll = roll
if roll == 6:
sixCount=sixCount+1
if sixCount <2:
return maxRoll
else:
return 7
# +
def get_result(roll):
if roll < 4:
return 0
if roll < 6:
return 1
if roll < 7:
return 2
return 3
# +
def get_results(dice_size, run_count):
results = []
result_counts = [0,0,0,0]
for i in range(run_count):
result = get_result(get_roll(dice_size))
results.append(result)
result_counts[result] = result_counts[result]+1
return (results, result_counts)
# -
def do_plot(size):
results, result_counts = get_results(size, 100000)
total_results = sum(result_counts)
result_ratios = [0,0,0,0]
for i in range(4):
result_ratios[i] = (result_counts[i]*100) / total_results
register = 0,1,2,3
result_types = "bad_outcome", "partial success", "full success", "critical success"
plt.figure(figsize = (8,4))
plt.bar(register, result_ratios, width = 0.8)
plt.title("likelyhood of outcomes with {0} dice".format(size), fontsize=16)
plt.xticks(register, result_types)
plt.xlabel("outcomes")
plt.ylabel("percent chance")
plt.ylim(0,119)
for x, y in zip(range(4), result_ratios):
label = "{:.0f}%".format(y)
plt.annotate(label,
(x,y), # this is the point to label
textcoords="offset points", # how to position the text
xytext=(0,10), # distance from text to points (x,y)
ha='center') # horizontal alignment can be left, right or center
plt.show()
return result_ratios;
def plot_scenario_chance(result_ratio_arrays, max_dice_size, scenario_text, result_array):
full_success_data = []
dice_sizes = []
for i in range(max_dice_size):
full_success = 0
for j in result_array:
full_success = full_success + result_ratio_arrays[i][j]
full_success_data.append(full_success)
dice_sizes.append(str(i))
register = range(max_dice_size)
plt.figure(figsize = (8,4))
plt.bar(register, full_success_data, width = 0.8)
plt.title("likelyhood of {0} by dice count".format(scenario_text), fontsize=16)
plt.xticks(register, dice_sizes)
plt.xlabel("number of dice")
plt.ylabel("percent chance of {0}".format(scenario_text))
plt.ylim(0,119)
for x, y in zip(range(max_dice_size), full_success_data):
label = "{:.0f}%".format(y)
plt.annotate(label,
(x,y), # this is the point to label
textcoords="offset points", # how to position the text
xytext=(0,10), # distance from text to points (x,y)
ha='center') # horizontal alignment can be left, right or center
plt.show()
result_ratio_arrays.append(do_plot(0))
result_ratio_arrays.append(do_plot(1))
result_ratio_arrays.append(do_plot(2))
result_ratio_arrays.append(do_plot(3))
result_ratio_arrays.append(do_plot(4))
result_ratio_arrays.append(do_plot(5))
result_ratio_arrays.append(do_plot(6))
plot_scenario_chance(result_ratio_arrays, 7, "at least full success", [2,3])
plot_scenario_chance(result_ratio_arrays, 7, "partial sucess or worse", [0,1])
plot_scenario_chance(result_ratio_arrays, 7, "at least partial sucess", [1,2,3])
plot_scenario_chance(result_ratio_arrays, 7, "critical sucess", [3])
plot_scenario_chance(result_ratio_arrays, 7, "bad outcome", [0])
| Blades Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:expression-classification]
# language: python
# name: conda-env-expression-classification-py
# ---
# # Evaluate Classifier Predictions
#
# **<NAME>, 2018**
#
# In the following notebook I evaluate the predictions made by the Ras, _NF1_, and _TP53_ classifiers in the input PDX RNAseq data.
#
# ## Procedure
#
# 1. Load status matrices
# * These files store the mutation status for _TP53_ and Ras pathway genes for the input samples
# 2. Align barcode identifiers
# * The identifiers matching the RNAseq data to the status matrix are not aligned.
# * I use an intermediate dictionary to map common identifiers
# 3. Load predictions (see `1.apply-classifier.ipynb` for more details)
# 4. Evaluate predictions
# * I visualize the distribution of predictions between wild-type and mutant samples for both classifiers
#
# ## Output
#
# The output of this notebook are several evaluation figures demonstrating the predictive performance on the input data for the three classifiers. Included in this output are predictions stratified by histology.
# +
import os
import random
from decimal import Decimal
from scipy.stats import ttest_ind
import numpy as np
import pandas as pd
from sklearn.metrics import average_precision_score, roc_auc_score
from sklearn.metrics import roc_curve, precision_recall_curve
import seaborn as sns
import matplotlib.pyplot as plt
from utils import get_mutant_boxplot, perform_ttest
# -
# %matplotlib inline
np.random.seed(123)
# ## Load Status Matrix
# +
file = os.path.join('data', 'raw', '2019-02-14-ras-tp53-nf1-alterations.txt')
status_df = pd.read_table(file)
print(status_df.shape)
status_df.head(3)
# -
status_df.Hugo_Symbol.value_counts()
status_df.Variant_Classification.value_counts()
pd.crosstab(status_df['Histology.Detailed'], status_df.Hugo_Symbol)
# Obtain a binary status matrix
full_status_df = pd.crosstab(status_df['Model'], status_df.Hugo_Symbol)
full_status_df[full_status_df > 1] = 1
full_status_df = full_status_df.reset_index()
# +
histology_df = status_df.loc[:, ['Model', 'Histology.Detailed']]
histology_df.columns = ['Model', 'Histology_Full']
full_status_df = (
full_status_df
.merge(histology_df, how='left', on="Model")
.drop_duplicates()
.reset_index(drop=True)
)
print(full_status_df.shape)
full_status_df.head()
# -
# ## Extract Gene Status
# +
# Ras Pathway Alterations
ras_genes = ['KRAS', 'HRAS', 'NRAS']
tp53_genes = ["TP53"]
nf1_genes = ["NF1"]
full_status_df = (
full_status_df
.assign(ras_status = full_status_df.loc[:, ras_genes].sum(axis=1),
tp53_status = full_status_df.loc[:, tp53_genes].sum(axis=1),
nf1_status = full_status_df.loc[:, nf1_genes].sum(axis=1))
)
full_status_df.head()
# -
# ## Load Clinical Data Information
#
# This stores histology information
# +
file = os.path.join('data', 'raw', 'pptc-pdx-clinical-web.txt')
clinical_df = pd.read_table(file)
print(clinical_df.shape)
clinical_df.head(3)
# -
# ## Load Predictions and Merge with Clinical and Alteration Data
# +
file = os.path.join('results', 'classifier_scores.tsv')
scores_df = pd.read_table(file)
scores_df = (
scores_df.merge(
clinical_df,
how='left', left_on='sample_id', right_on='Model'
)
.merge(
full_status_df,
how='left', left_on='sample_id', right_on='Model'
)
)
print(scores_df.shape)
scores_df.head()
# +
gene_status = ['tp53_status', 'ras_status', 'nf1_status']
scores_df.loc[:, gene_status] = (
scores_df.loc[:, gene_status].fillna(0)
)
scores_df.loc[scores_df['tp53_status'] != 0, 'tp53_status'] = 1
scores_df.loc[scores_df['ras_status'] != 0, 'ras_status'] = 1
scores_df.loc[scores_df['nf1_status'] != 0, 'nf1_status'] = 1
scores_df['tp53_status'] = scores_df['tp53_status'].astype(int)
scores_df['ras_status'] = scores_df['ras_status'].astype(int)
scores_df['nf1_status'] = scores_df['nf1_status'].astype(int)
scores_df.head(2)
# -
# ## Load Histology Color Codes
file = os.path.join('data', '2019-07-09-all-hist-colors.txt')
color_code_df = pd.read_csv(file)
color_code_df.head(2)
color_dict = dict(zip(color_code_df.Histology, color_code_df.Color))
color_dict
# ## Determine Status Counts
scores_df.tp53_status.value_counts()
scores_df.ras_status.value_counts()
scores_df.nf1_status.value_counts()
# ## Perform ROC and Precision-Recall Analysis using all Alteration Information
# +
n_classes = 3
labels = ['Ras', 'NF1', 'TP53']
colors = ['#1b9e77', '#d95f02', '#7570b3']
fpr_pdx = {}
tpr_pdx = {}
thresh_pdx = {}
precision_pdx = {}
recall_pdx = {}
auroc_pdx = {}
aupr_pdx = {}
fpr_shuff = {}
tpr_shuff = {}
thresh_shuff = {}
precision_shuff = {}
recall_shuff = {}
auroc_shuff = {}
aupr_shuff = {}
all_roc_list = []
idx = 0
for status, score, shuff in zip(('ras_status', 'nf1_status', 'tp53_status'),
('ras_score', 'nf1_score', 'tp53_score'),
('ras_shuffle', 'nf1_shuffle', 'tp53_shuffle')):
# Obtain Metrics
sample_status = scores_df.loc[:, status]
sample_score = scores_df.loc[:, score]
shuffle_score = scores_df.loc[:, shuff]
# Get Metrics
fpr_pdx[idx], tpr_pdx[idx], thresh_pdx[idx] = roc_curve(sample_status, sample_score, drop_intermediate=False)
precision_pdx[idx], recall_pdx[idx], _ = precision_recall_curve(sample_status, sample_score)
auroc_pdx[idx] = roc_auc_score(sample_status, sample_score)
aupr_pdx[idx] = average_precision_score(sample_status, sample_score)
# Obtain Shuffled Metrics
fpr_shuff[idx], tpr_shuff[idx], thresh_shuff[idx] = roc_curve(sample_status, shuffle_score, drop_intermediate=False)
precision_shuff[idx], recall_shuff[idx], _ = precision_recall_curve(sample_status, shuffle_score)
auroc_shuff[idx] = roc_auc_score(sample_status, shuffle_score)
aupr_shuff[idx] = average_precision_score(sample_status, shuffle_score)
roc_df = (
pd.DataFrame([fpr_pdx[idx], tpr_pdx[idx], thresh_pdx[idx]], index=['fpr', 'tpr', 'threshold'])
.transpose()
.assign(gene=labels[idx],
shuffled=False)
)
roc_shuffled_df = (
pd.DataFrame([fpr_shuff[idx], tpr_shuff[idx], thresh_shuff[idx]], index=['fpr', 'tpr', 'threshold'])
.transpose()
.assign(gene=labels[idx],
shuffled=True)
)
all_roc_list.append(roc_df)
all_roc_list.append(roc_shuffled_df)
idx += 1
# -
os.makedirs('figures', exist_ok=True)
# +
# Visualize ROC curves
plt.subplots(figsize=(4, 4))
for i in range(n_classes):
plt.plot(fpr_pdx[i], tpr_pdx[i],
label='{} (AUROC = {})'.format(labels[i], round(auroc_pdx[i], 2)),
linestyle='solid',
color=colors[i])
# Shuffled Data
plt.plot(fpr_shuff[i], tpr_shuff[i],
label='{} Shuffle (AUROC = {})'.format(labels[i], round(auroc_shuff[i], 2)),
linestyle='dotted',
color=colors[i])
plt.axis('equal')
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.xlabel('False Positive Rate', fontsize=12)
plt.ylabel('True Positive Rate', fontsize=12)
plt.tick_params(labelsize=10)
lgd = plt.legend(bbox_to_anchor=(1.03, 0.85),
loc=2,
borderaxespad=0.,
fontsize=10)
file = os.path.join('figures', 'classifier_roc_curve.pdf')
plt.savefig(file, bbox_extra_artists=(lgd,), bbox_inches='tight')
# +
# Visualize PR curves
plt.subplots(figsize=(4, 4))
for i in range(n_classes):
plt.plot(recall_pdx[i], precision_pdx[i],
label='{} (AUPR = {})'.format(labels[i], round(aupr_pdx[i], 2)),
linestyle='solid',
color=colors[i])
# Shuffled Data
plt.plot(recall_shuff[i], precision_shuff[i],
label='{} Shuffle (AUPR = {})'.format(labels[i], round(aupr_shuff[i], 2)),
linestyle='dotted',
color=colors[i])
plt.axis('equal')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.xlabel('Recall', fontsize=12)
plt.ylabel('Precision', fontsize=12)
plt.tick_params(labelsize=10)
lgd = plt.legend(bbox_to_anchor=(1.03, 0.85),
loc=2,
borderaxespad=0.,
fontsize=10)
file = os.path.join('figures', 'classifier_precision_recall_curve.pdf')
plt.savefig(file, bbox_extra_artists=(lgd,), bbox_inches='tight')
# -
# ## Perform t-test against status classification
t_results_ras = perform_ttest(scores_df, gene='ras')
t_results_ras
t_results_nf1 = perform_ttest(scores_df, gene='nf1')
t_results_nf1
t_results_tp53 = perform_ttest(scores_df, gene='tp53')
t_results_tp53
# ## Observe broad differences across sample categories
# Ras
get_mutant_boxplot(df=scores_df,
gene="Ras",
t_test_results=t_results_ras)
# NF1
get_mutant_boxplot(df=scores_df,
gene="NF1",
t_test_results=t_results_nf1)
# TP53
get_mutant_boxplot(df=scores_df,
gene="TP53",
t_test_results=t_results_tp53)
# ## Write output files for downstream analysis
# +
# Classifier scores with clinical data and alteration status
scores_file = os.path.join("results", "classifier_scores_with_clinical_and_alterations.tsv")
genes = ras_genes + ['TP53']
scores_df = scores_df.drop(['Model_x', 'Model_y', 'Histology_Full'], axis='columns')
scores_df[genes] = scores_df[genes].fillna(value=0)
scores_df.sort_values(by='sample_id').to_csv(scores_file, sep='\t', index=False)
# +
# Output classifier scores for the specific variants observed
status_scores_file = os.path.join("results", "classifier_scores_with_variants.tsv")
classifier_scores_df = scores_df[['sample_id', 'ras_score' ,'tp53_score', 'nf1_score', 'Histology.Detailed']]
classifier_scores_df = (
status_df
.drop(['Histology.Detailed'], axis='columns')
.merge(classifier_scores_df, how='left', left_on='Model', right_on='sample_id')
)
classifier_scores_df.sort_values(by='Model').to_csv(status_scores_file, sep='\t', index=False)
# +
# ROC Curve Estimates
file = os.path.join("results", "full_roc_threshold_results.tsv")
full_roc_df = pd.concat(all_roc_list, axis='rows')
full_roc_df.to_csv(file, sep='\t', index=False)
| 2.evaluate-classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="MhoQ0WE77laV"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="_ckMIh7O7s6D"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="jYysdyb-CaWM"
# # Custom training with tf.distribute.Strategy
# + [markdown] colab_type="text" id="S5Uhzt6vVIB2"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/distribute/custom_training"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/distribute/custom_training.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/distribute/custom_training.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/distribute/custom_training.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="FbVhjPpzn6BM"
# This tutorial demonstrates how to use [`tf.distribute.Strategy`](https://www.tensorflow.org/guide/distribute_strategy) with custom training loops. We will train a simple CNN model on the fashion MNIST dataset. The fashion MNIST dataset contains 60000 train images of size 28 x 28 and 10000 test images of size 28 x 28.
#
# We are using custom training loops to train our model because they give us flexibility and a greater control on training. Moreover, it is easier to debug the model and the training loop.
# + colab={} colab_type="code" id="dzLKpmZICaWN"
from __future__ import absolute_import, division, print_function, unicode_literals
# Import TensorFlow
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
# Helper libraries
import numpy as np
import os
print(tf.__version__)
# + [markdown] colab_type="text" id="MM6W__qraV55"
# ## Download the fashion MNIST dataset
# + colab={} colab_type="code" id="7MqDQO0KCaWS"
fashion_mnist = tf.keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# Adding a dimension to the array -> new shape == (28, 28, 1)
# We are doing this because the first layer in our model is a convolutional
# layer and it requires a 4D input (batch_size, height, width, channels).
# batch_size dimension will be added later on.
train_images = train_images[..., None]
test_images = test_images[..., None]
# Getting the images in [0, 1] range.
train_images = train_images / np.float32(255)
test_images = test_images / np.float32(255)
# + [markdown] colab_type="text" id="4AXoHhrsbdF3"
# ## Create a strategy to distribute the variables and the graph
# + [markdown] colab_type="text" id="5mVuLZhbem8d"
# How does `tf.distribute.MirroredStrategy` strategy work?
#
# * All the variables and the model graph is replicated on the replicas.
# * Input is evenly distributed across the replicas.
# * Each replica calculates the loss and gradients for the input it received.
# * The gradients are synced across all the replicas by summing them.
# * After the sync, the same update is made to the copies of the variables on each replica.
#
# Note: You can put all the code below inside a single scope. We are dividing it into several code cells for illustration purposes.
#
# + colab={} colab_type="code" id="F2VeZUWUj5S4"
# If the list of devices is not specified in the
# `tf.distribute.MirroredStrategy` constructor, it will be auto-detected.
strategy = tf.distribute.MirroredStrategy()
# + colab={} colab_type="code" id="ZngeM_2o0_JO"
print ('Number of devices: {}'.format(strategy.num_replicas_in_sync))
# + [markdown] colab_type="text" id="k53F5I_IiGyI"
# ## Setup input pipeline
# + [markdown] colab_type="text" id="0Qb6nDgxiN_n"
# Export the graph and the variables to the platform-agnostic SavedModel format. After your model is saved, you can load it with or without the scope.
# + colab={} colab_type="code" id="jwJtsCQhHK-E"
BUFFER_SIZE = len(train_images)
BATCH_SIZE_PER_REPLICA = 64
GLOBAL_BATCH_SIZE = BATCH_SIZE_PER_REPLICA * strategy.num_replicas_in_sync
EPOCHS = 10
# + [markdown] colab_type="text" id="J7fj3GskHC8g"
# Create the datasets and distribute them:
# + colab={} colab_type="code" id="WYrMNNDhAvVl"
train_dataset = tf.data.Dataset.from_tensor_slices((train_images, train_labels)).shuffle(BUFFER_SIZE).batch(GLOBAL_BATCH_SIZE)
test_dataset = tf.data.Dataset.from_tensor_slices((test_images, test_labels)).batch(GLOBAL_BATCH_SIZE)
train_dist_dataset = strategy.experimental_distribute_dataset(train_dataset)
test_dist_dataset = strategy.experimental_distribute_dataset(test_dataset)
# + [markdown] colab_type="text" id="bAXAo_wWbWSb"
# ## Create the model
#
# Create a model using `tf.keras.Sequential`. You can also use the Model Subclassing API to do this.
# + colab={} colab_type="code" id="9ODch-OFCaW4"
def create_model():
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu'),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Conv2D(64, 3, activation='relu'),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
return model
# + colab={} colab_type="code" id="9iagoTBfijUz"
# Create a checkpoint directory to store the checkpoints.
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
# + [markdown] colab_type="text" id="e-wlFFZbP33n"
# ## Define the loss function
#
# Normally, on a single machine with 1 GPU/CPU, loss is divided by the number of examples in the batch of input.
#
# *So, how should the loss be calculated when using a `tf.distribute.Strategy`?*
#
# * For an example, let's say you have 4 GPU's and a batch size of 64. One batch of input is distributed
# across the replicas (4 GPUs), each replica getting an input of size 16.
#
# * The model on each replica does a forward pass with its respective input and calculates the loss. Now, instead of dividing the loss by the number of examples in its respective input (BATCH_SIZE_PER_REPLICA = 16), the loss should be divided by the GLOBAL_BATCH_SIZE (64).
#
# *Why do this?*
#
# * This needs to be done because after the gradients are calculated on each replica, they are synced across the replicas by **summing** them.
#
# *How to do this in TensorFlow?*
# * If you're writing a custom training loop, as in this tutorial, you should sum the per example losses and divide the sum by the GLOBAL_BATCH_SIZE:
# `scale_loss = tf.reduce_sum(loss) * (1. / GLOBAL_BATCH_SIZE)`
# or you can use `tf.nn.compute_average_loss` which takes the per example loss,
# optional sample weights, and GLOBAL_BATCH_SIZE as arguments and returns the scaled loss.
#
# * If you are using regularization losses in your model then you need to scale
# the loss value by number of replicas. You can do this by using the `tf.nn.scale_regularization_loss` function.
#
# * Using `tf.reduce_mean` is not recommended. Doing so divides the loss by actual per replica batch size which may vary step to step.
#
# * This reduction and scaling is done automatically in keras `model.compile` and `model.fit`
#
# * If using `tf.keras.losses` classes (as in the example below), the loss reduction needs to be explicitly specified to be one of `NONE` or `SUM`. `AUTO` and `SUM_OVER_BATCH_SIZE` are disallowed when used with `tf.distribute.Strategy`. `AUTO` is disallowed because the user should explicitly think about what reduction they want to make sure it is correct in the distributed case. `SUM_OVER_BATCH_SIZE` is disallowed because currently it would only divide by per replica batch size, and leave the dividing by number of replicas to the user, which might be easy to miss. So instead we ask the user do the reduction themselves explicitly.
# + colab={} colab_type="code" id="R144Wci782ix"
with strategy.scope():
# Set reduction to `none` so we can do the reduction afterwards and divide by
# global batch size.
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(
reduction=tf.keras.losses.Reduction.NONE)
# or loss_fn = tf.keras.losses.sparse_categorical_crossentropy
def compute_loss(labels, predictions):
per_example_loss = loss_object(labels, predictions)
return tf.nn.compute_average_loss(per_example_loss, global_batch_size=GLOBAL_BATCH_SIZE)
# + [markdown] colab_type="text" id="w8y54-o9T2Ni"
# ## Define the metrics to track loss and accuracy
#
# These metrics track the test loss and training and test accuracy. You can use `.result()` to get the accumulated statistics at any time.
# + colab={} colab_type="code" id="zt3AHb46Tr3w"
with strategy.scope():
test_loss = tf.keras.metrics.Mean(name='test_loss')
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(
name='train_accuracy')
test_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(
name='test_accuracy')
# + [markdown] colab_type="text" id="iuKuNXPORfqJ"
# ## Training loop
# + colab={} colab_type="code" id="OrMmakq5EqeQ"
# model and optimizer must be created under `strategy.scope`.
with strategy.scope():
model = create_model()
optimizer = tf.keras.optimizers.Adam()
checkpoint = tf.train.Checkpoint(optimizer=optimizer, model=model)
# + colab={} colab_type="code" id="3UX43wUu04EL"
with strategy.scope():
def train_step(inputs):
images, labels = inputs
with tf.GradientTape() as tape:
predictions = model(images, training=True)
loss = compute_loss(labels, predictions)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_accuracy.update_state(labels, predictions)
return loss
def test_step(inputs):
images, labels = inputs
predictions = model(images, training=False)
t_loss = loss_object(labels, predictions)
test_loss.update_state(t_loss)
test_accuracy.update_state(labels, predictions)
# + colab={} colab_type="code" id="gX975dMSNw0e"
with strategy.scope():
# `experimental_run_v2` replicates the provided computation and runs it
# with the distributed input.
@tf.function
def distributed_train_step(dataset_inputs):
per_replica_losses = strategy.experimental_run_v2(train_step,
args=(dataset_inputs,))
return strategy.reduce(tf.distribute.ReduceOp.SUM, per_replica_losses,
axis=None)
@tf.function
def distributed_test_step(dataset_inputs):
return strategy.experimental_run_v2(test_step, args=(dataset_inputs,))
for epoch in range(EPOCHS):
# TRAIN LOOP
total_loss = 0.0
num_batches = 0
for x in train_dist_dataset:
total_loss += distributed_train_step(x)
num_batches += 1
train_loss = total_loss / num_batches
# TEST LOOP
for x in test_dist_dataset:
distributed_test_step(x)
if epoch % 2 == 0:
checkpoint.save(checkpoint_prefix)
template = ("Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, "
"Test Accuracy: {}")
print (template.format(epoch+1, train_loss,
train_accuracy.result()*100, test_loss.result(),
test_accuracy.result()*100))
test_loss.reset_states()
train_accuracy.reset_states()
test_accuracy.reset_states()
# + [markdown] colab_type="text" id="Z1YvXqOpwy08"
# Things to note in the example above:
#
# * We are iterating over the `train_dist_dataset` and `test_dist_dataset` using a `for x in ...` construct.
# * The scaled loss is the return value of the `distributed_train_step`. This value is aggregated across replicas using the `tf.distribute.Strategy.reduce` call and then across batches by summing the return value of the `tf.distribute.Strategy.reduce` calls.
# * `tf.keras.Metrics` should be updated inside `train_step` and `test_step` that gets executed by `tf.distribute.Strategy.experimental_run_v2`.
# *`tf.distribute.Strategy.experimental_run_v2` returns results from each local replica in the strategy, and there are multiple ways to consume this result. You can do `tf.distribute.Strategy.reduce` to get an aggregated value. You can also do `tf.distribute.Strategy.experimental_local_results` to get the list of values contained in the result, one per local replica.
#
# + [markdown] colab_type="text" id="-q5qp31IQD8t"
# ## Restore the latest checkpoint and test
# + [markdown] colab_type="text" id="WNW2P00bkMGJ"
# A model checkpointed with a `tf.distribute.Strategy` can be restored with or without a strategy.
# + colab={} colab_type="code" id="pg3B-Cw_cn3a"
eval_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(
name='eval_accuracy')
new_model = create_model()
new_optimizer = tf.keras.optimizers.Adam()
test_dataset = tf.data.Dataset.from_tensor_slices((test_images, test_labels)).batch(GLOBAL_BATCH_SIZE)
# + colab={} colab_type="code" id="7qYii7KUYiSM"
@tf.function
def eval_step(images, labels):
predictions = new_model(images, training=False)
eval_accuracy(labels, predictions)
# + colab={} colab_type="code" id="LeZ6eeWRoUNq"
checkpoint = tf.train.Checkpoint(optimizer=new_optimizer, model=new_model)
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
for images, labels in test_dataset:
eval_step(images, labels)
print ('Accuracy after restoring the saved model without strategy: {}'.format(
eval_accuracy.result()*100))
# + [markdown] colab_type="text" id="EbcI87EEzhzg"
# ## Alternate ways of iterating over a dataset
#
# ### Using iterators
#
# If you want to iterate over a given number of steps and not through the entire dataset you can create an iterator using the `iter` call and explicity call `next` on the iterator. You can choose to iterate over the dataset both inside and outside the tf.function. Here is a small snippet demonstrating iteration of the dataset outside the tf.function using an iterator.
#
# + colab={} colab_type="code" id="7c73wGC00CzN"
with strategy.scope():
for _ in range(EPOCHS):
total_loss = 0.0
num_batches = 0
train_iter = iter(train_dist_dataset)
for _ in range(10):
total_loss += distributed_train_step(next(train_iter))
num_batches += 1
average_train_loss = total_loss / num_batches
template = ("Epoch {}, Loss: {}, Accuracy: {}")
print (template.format(epoch+1, average_train_loss, train_accuracy.result()*100))
train_accuracy.reset_states()
# + [markdown] colab_type="text" id="GxVp48Oy0m6y"
# ### Iterating inside a tf.function
# You can also iterate over the entire input `train_dist_dataset` inside a tf.function using the `for x in ...` construct or by creating iterators like we did above. The example below demonstrates wrapping one epoch of training in a tf.function and iterating over `train_dist_dataset` inside the function.
# + colab={} colab_type="code" id="-REzmcXv00qm"
with strategy.scope():
@tf.function
def distributed_train_epoch(dataset):
total_loss = 0.0
num_batches = 0
for x in dataset:
per_replica_losses = strategy.experimental_run_v2(train_step,
args=(x,))
total_loss += strategy.reduce(
tf.distribute.ReduceOp.SUM, per_replica_losses, axis=None)
num_batches += 1
return total_loss / tf.cast(num_batches, dtype=tf.float32)
for epoch in range(EPOCHS):
train_loss = distributed_train_epoch(train_dist_dataset)
template = ("Epoch {}, Loss: {}, Accuracy: {}")
print (template.format(epoch+1, train_loss, train_accuracy.result()*100))
train_accuracy.reset_states()
# + [markdown] colab_type="text" id="MuZGXiyC7ABR"
# ### Tracking training loss across replicas
#
# Note: As a general rule, you should use `tf.keras.Metrics` to track per-sample values and avoid values that have been aggregated within a replica.
#
# We do *not* recommend using `tf.metrics.Mean` to track the training loss across different replicas, because of the loss scaling computation that is carried out.
#
# For example, if you run a training job with the following characteristics:
# * Two replicas
# * Two samples are processed on each replica
# * Resulting loss values: [2, 3] and [4, 5] on each replica
# * Global batch size = 4
#
# With loss scaling, you calculate the per-sample value of loss on each replica by adding the loss values, and then dividing by the global batch size. In this case: `(2 + 3) / 4 = 1.25` and `(4 + 5) / 4 = 2.25`.
#
# If you use `tf.metrics.Mean` to track loss across the two replicas, the result is different. In this example, you end up with a `total` of 3.50 and `count` of 2, which results in `total`/`count` = 1.75 when `result()` is called on the metric. Loss calculated with `tf.keras.Metrics` is scaled by an additional factor that is equal to the number of replicas in sync.
# + [markdown] colab_type="text" id="xisYJaV9KZTN"
# ### Examples and Tutorials
# Here are some examples for using distribution strategy with custom training loops:
#
# 1. [Tutorial](../tutorials/distribute/training_loops.ipynb) to train MNIST using `MirroredStrategy`.
# 2. [DenseNet](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/densenet/distributed_train.py) example using `MirroredStrategy`.
# 1. [BERT](https://github.com/tensorflow/models/blob/master/official/bert/run_classifier.py) example trained using `MirroredStrategy` and `TPUStrategy`.
# This example is particularly helpful for understanding how to load from a checkpoint and generate periodic checkpoints during distributed training etc.
# 2. [NCF](https://github.com/tensorflow/models/blob/master/official/recommendation/ncf_keras_main.py) example trained using `MirroredStrategy` that can be enabled using the `keras_use_ctl` flag.
# 3. [NMT](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/nmt_with_attention/distributed_train.py) example trained using `MirroredStrategy`.
#
# More examples listed in the [Distribution strategy guide](../../guide/distribute_strategy.ipynb#examples_and_tutorials)
# + [markdown] colab_type="text" id="6hEJNsokjOKs"
# ## Next steps
#
# Try out the new `tf.distribute.Strategy` API on your models.
| site/en/tutorials/distribute/custom_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Master Drifter Evaluation Notebook
#
# ***
# +
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
from datetime import datetime, timedelta
from scipy.io import loadmat
from sys import float_info as floats
from salishsea_tools import viz_tools
# %matplotlib inline
# -
plt.rcParams['font.size'] = 14
# ***
#
# ### Skill score definition
#
# [<NAME> 2011, JGR Oceans](https://agupubs.onlinelibrary.wiley.com/doi/full/10.1029/2010JC006837)
#
# $$\left. s = \sum_{i=1}^Nd_i \middle/ \sum_{i=1}^NI_{oi} \right.$$
def mtime2datetime(mtime):
"""Convert matlab time to python datetime object
"""
pytime = datetime.fromordinal(int(mtime) - 366) + timedelta(days=mtime%1)
return pytime
# Paths, files and definitions
paths = {
'coords': '/data/bmoorema/MEOPAR/grid/coordinates_seagrid_SalishSea201702.nc',
'mask': '/data/bmoorema/MEOPAR/grid/mesh_mask201702.nc',
'drifters': '/ocean/shared/drifters/data/L3/Salish_L3_20190728T103529.mat',
'out': '/data/bmoorema/results/parcels/drifters_base/',
}
grid = xr.open_dataset(paths['coords'], decode_times=False)
mask = xr.open_dataset(paths['mask'])
drifters = loadmat(paths['drifters'])['drift'][0]
IDs = [4, 32, 73, 82, 94, 106, 132, 142]
deg2m = 111000 * np.cos(50 * np.pi / 180)
duration = timedelta(days=5)
# +
# Make figure
fig = plt.figure(figsize=(15, 15))
gs = plt.GridSpec(5, 4, height_ratios=[3, 3, 2, 2, 2])
axs_maps = [fig.add_subplot(gs[row, col]) for row in range(2) for col in range(4)]
axs_metrics = [fig.add_subplot(gs[row+2, :]) for row in range(3)]
ylims, ylabels = [(0, 100), (0, 100), (0, 2)], ['$d$ [km]', '$I_0$ [km]', '$ss$']
axs_metrics[2].set_xlabel('Time elapsed [h]')
for ax, ylim, ylabel in zip(axs_metrics, ylims, ylabels):
ax.set_xlim([0, 71])
ax.set_ylim(ylim)
ax.set_ylabel(ylabel)
if ylabel != '$ss$': ax.xaxis.set_ticklabels('')
# Loop through simulations
for ax, ID, color in zip(axs_maps, IDs, ['cornflowerblue', 'orange', 'g', 'm', 'r', 'gray', 'brown', 'c']):
# Parse ID
dindex = np.where(drifters['id'].astype(int) == ID)[0][0]
t_obs = np.array([mtime2datetime(float(t)) for t in drifters['mtime'][dindex]])
fn = f'drifter{ID:03d}_' + '_'.join(d.strftime('%Y%m%d') for d in [t_obs[0], t_obs[0]+duration]) + '.nc'
metrics = {'dl': np.empty(0), 'd': np.empty((0, 50))}
label = t_obs[0].strftime('%Y-%b-%d')
# Make map area
ax.contourf(grid.nav_lon, grid.nav_lat, mask.tmask[0, 0, ...], levels=[-0.01, 0.01], colors='gray')
ax.contour(grid.nav_lon, grid.nav_lat, mask.tmask[0, 0, ...], levels=[-0.01, 0.01], colors='k')
ax.set_xlim([-124, -122.9])
ax.set_ylim([48.6, 49.5])
ax.set_title(label)
ax.xaxis.set_ticks([-124, -123.5, -123])
ax.yaxis.set_ticks([49, 49.5])
ax.xaxis.set_ticklabels(['124$^{\circ}$W', '123.5$^{\circ}$W', '123$^{\circ}$W'])
ax.yaxis.set_ticklabels(['49$^{\circ}$N', '49.5$^{\circ}$N'])
viz_tools.set_aspect(ax)
if (ID != 4) and (ID != 94): ax.yaxis.set_ticklabels('')
if ID < 94: ax.xaxis.set_ticklabels('')
# Load simulation
with xr.open_dataset(paths['out'] + fn) as data:
# Load and plot trajectories
times = data.time[0, :].values.astype('datetime64[s]').astype(datetime)
lons, lats = data.lon.values, data.lat.values
# Loop through simulation time points
index = 0
for time, lon, lat in zip(times[:72], lons.T, lats.T):
# Extract observations
ilast = index
index = abs(t_obs - time).argmin()
lon_obs, lat_obs = drifters['lon'][dindex], drifters['lat'][dindex]
ax.plot(lon, lat, 'ko', zorder=1)
ax.plot(lon_obs[index], lat_obs[index], 'ko', markerfacecolor=color, zorder=2)
# Calculate metrics
d = np.sqrt((lon - lon_obs[index])**2 + (lat - lat_obs[index])**2) * deg2m * 1e-3
dl = np.sqrt((lon_obs[index] - lon_obs[ilast])**2 + (lat_obs[index] - lat_obs[ilast])**2) * deg2m * 1e-3
metrics['dl'] = np.append(metrics['dl'], dl)
metrics['d'] = np.concatenate((metrics['d'], d[np.newaxis, :]))
# Calc skill score
I_0 = np.cumsum(metrics['dl'])[:, np.newaxis]
ss = np.cumsum(metrics['d'], axis=0) / np.cumsum(I_0, axis=0)
for ax, var in zip(axs_metrics, [metrics['d'], I_0, ss]):
ax.plot(var.mean(axis=1), 'o-', color=color, markeredgecolor='k')
fig.savefig('/home/bmoorema/Desktop/skill_scores.png', bbox_inches='tight')
# -
| notebooks/OpenDrift/drifter_evaluation_master.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # 41. Export Run History as Tensorboard logs
#
# 1. Run some training and log some metrics into Run History
# 2. Export the run history to some directory as Tensorboard logs
# 3. Launch a local Tensorboard to view the run history
# ## Prerequisites
# Make sure you go through the [00. Installation and Configuration](00.configuration.ipynb) Notebook first if you haven't.
# +
# Check core SDK version number
import azureml.core
print("SDK version:", azureml.core.VERSION)
# -
# ## Initialize Workspace
#
# Initialize a workspace object from persisted configuration.
# +
from azureml.core import Workspace, Run, Experiment
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
# -
# ## Set experiment name and start the run
experiment_name = 'export-to-tensorboard'
exp = Experiment(ws, experiment_name)
root_run = exp.start_logging()
# +
# load diabetes dataset, a well-known built-in small dataset that comes with scikit-learn
from sklearn.datasets import load_diabetes
from sklearn.linear_model import Ridge
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
X, y = load_diabetes(return_X_y=True)
columns = ['age', 'gender', 'bmi', 'bp', 's1', 's2', 's3', 's4', 's5', 's6']
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
data = {
"train":{"x":x_train, "y":y_train},
"test":{"x":x_test, "y":y_test}
}
# +
# Example experiment
from tqdm import tqdm
alphas = [.1, .2, .3, .4, .5, .6 , .7]
# try a bunch of alpha values in a Linear Regression (Ridge) model
for alpha in tqdm(alphas):
# create a bunch of child runs
with root_run.child_run("alpha" + str(alpha)) as run:
# More data science stuff
reg = Ridge(alpha=alpha)
reg.fit(data["train"]["x"], data["train"]["y"])
# TODO save model
preds = reg.predict(data["test"]["x"])
mse = mean_squared_error(preds, data["test"]["y"])
# End train and eval
# log alpha, mean_squared_error and feature names in run history
root_run.log("alpha", alpha)
root_run.log("mse", mse)
# -
# ## Export Run History to Tensorboard logs
# +
# Export Run History to Tensorboard logs
from azureml.contrib.tensorboard.export import export_to_tensorboard
import os
import tensorflow as tf
logdir = 'exportedTBlogs'
log_path = os.path.join(os.getcwd(), logdir)
try:
os.stat(log_path)
except os.error:
os.mkdir(log_path)
print(logdir)
# export run history for the project
export_to_tensorboard(root_run, logdir)
# or export a particular run
# export_to_tensorboard(run, logdir)
# -
root_run.complete()
# ## Start Tensorboard
#
# Or you can start the Tensorboard outside this notebook to view the result
# +
from azureml.contrib.tensorboard import Tensorboard
# The Tensorboard constructor takes an array of runs, so be sure and pass it in as a single-element array here
tb = Tensorboard([], local_root=logdir, port=6006)
# If successful, start() returns a string with the URI of the instance.
tb.start()
# -
# ## Stop Tensorboard
#
# When you're done, make sure to call the `stop()` method of the Tensorboard object.
tb.stop()
| training/08.export-run-history-to-tensorboard/08.export-run-history-to-tensorboard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="AVEeBulfJkzl" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 1*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Regression 1
#
# ## Assignment
#
# You'll use another **New York City** real estate dataset.
#
# But now you'll **predict how much it costs to rent an apartment**, instead of how much it costs to buy a condo.
#
# The data comes from renthop.com, an apartment listing website.
#
# - [ ] Look at the data. Choose a feature, and plot its relationship with the target.
# - [ ] Use scikit-learn for linear regression with one feature. You can follow the [5-step process from Jake VanderPlas](https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html#Basics-of-the-API).
# - [ ] Define a function to make new predictions and explain the model coefficient.
# - [ ] Organize and comment your code.
#
# > [Do Not Copy-Paste.](https://docs.google.com/document/d/1ubOw9B3Hfip27hF2ZFnW3a3z9xAgrUDRReOEo-FHCVs/edit) You must type each of these exercises in, manually. If you copy and paste, you might as well not even do them. The point of these exercises is to train your hands, your brain, and your mind in how to read, write, and see code. If you copy-paste, you are cheating yourself out of the effectiveness of the lessons.
#
# If your **Plotly** visualizations aren't working:
# - You must have JavaScript enabled in your browser
# - You probably want to use Chrome or Firefox
# - You may need to turn off ad blockers
# - [If you're using Jupyter Lab locally, you need to install some "extensions"](https://plot.ly/python/getting-started/#jupyterlab-support-python-35)
#
# ## Stretch Goals
# - [ ] Do linear regression with two or more features.
# - [ ] Read [The Discovery of Statistical Regression](https://priceonomics.com/the-discovery-of-statistical-regression/)
# - [ ] Read [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/ISLR%20Seventh%20Printing.pdf), Chapter 2.1: What Is Statistical Learning?
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + colab_type="code" id="4S2wXSrFV_g4" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="e01673c0-0c9c-4a6a-dd26-bf8f68a70f5a"
# Read New York City apartment rental listing data
import pandas as pd
df = pd.read_csv(DATA_PATH+'apartments/renthop-nyc.csv',
parse_dates=['created'])
assert df.shape == (49352, 34)
df.head()
# + id="gv9_GPj1Nr5x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 728} outputId="28216d4d-29ee-4132-e19d-5d286e4fb134"
df.info()
# + id="RaU-KH-XJkzv" colab_type="code" colab={}
# Remove outliers:
# the most extreme 1% prices,
# the most extreme .1% latitudes, &
# the most extreme .1% longitudes
df = df[(df['price'] >= 1375) & (df['price'] <= 15500) &
(df['latitude'] >=40.57) & (df['latitude'] < 40.99) &
(df['longitude'] >= -74.1) & (df['longitude'] <= -73.38)]
# + [markdown] id="gXqnnDsdOk09" colab_type="text"
# ## EDA
# + id="OAq8WiBmOUxs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="c1d53e1c-fffb-40f0-deed-faba26e758bc"
#Choose a feature, and plot its relationship with the target.
df['price'].plot(kind='hist')
# + id="GbCD12vuP52i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="37266470-0785-4981-9bb7-ff168f1a8262"
df.describe()['price']
# + id="Q_hmn1sUP8kI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="7da1b68d-b27b-4458-b682-b3f4eb0df13e"
df['bathrooms'].plot(kind='hist')
# + id="9R8nLNvKi4P7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="4c6b1a3e-6f2b-40c1-8f92-382b38d79214"
df.describe()['bathrooms']
# + id="mMh9L81KQuxs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 446} outputId="571462eb-3f2e-49c6-aade-f9563f7539da"
pd.crosstab(df['price'],df['bathrooms'])
# + [markdown] id="YSyXEiK5RKi-" colab_type="text"
# ## split data
# + [markdown] id="JuX2KKGMRIym" colab_type="text"
#
# + id="a0wu6N39RQhv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="429c1513-c619-4b9f-f7d2-7191a65e2eb8"
y = df['price']
X = df[['bathrooms']]
import matplotlib.pyplot as plt
plt.scatter(X, y)
plt.xlabel('Bathrooms')
plt.ylabel('Price')
#line of best fit
# + [markdown] id="veS2tId7TYLw" colab_type="text"
# ## train validation split
# + id="WwxGN1M-Tax1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 156} outputId="abf1f133-c477-47d9-ff8c-85ebd8920bf4"
X.info()
# + id="vPNMi9uhTi0B" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="08d9bd54-155b-42b5-dd14-7c588bfcd40c"
X.shape
# + id="72X7lUiAx6mf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d3f42d51-1c3f-4730-eb9e-e112b76f3d4f"
#create a mask to split data into 20% and 80% data
mask = X.index < (48818*0.2)
mask
# + id="zAiugfelyKi1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="564c94df-496a-41cb-d8ba-cc37bb82ad29"
X_train, y_train = X.loc[mask], y.loc[mask]
X_val, y_val = X.loc[~mask], y.loc[~mask]
print(X_train.shape)
print(X_val.shape)
print(y_train.shape)
print(y_val.shape)
# + id="BDLBQC5ZytNA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="16932fb8-cbf8-4273-9685-7bbf5aa96ffa"
#creating a baseline
baseline_guess = y_train.mean()
MAE = abs(y_train - baseline_guess).mean()
print(f'''if baseline model always predicts {baseline_guess},
on average, the prediction will be off by {MAE}.''')
# + id="jD_Co8YDzoH1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d0c4873b-33a8-4432-a3e3-462403db8a79"
#Use scikit-learn for linear regression with one feature.
from sklearn.linear_model import LinearRegression
#instantiate predictor
lr = LinearRegression()
#train predictor using training data
lr.fit(X_train, y_train)
# + id="_jYhE7Bu0XK0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a87bb588-ec3f-407f-fdd5-ca54b283b721"
#always need the zeroth index of lr.coef bc trained on one feature
lr.coef_[0]
# + id="raiNx_V00b-r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="445d699c-ad6a-4c28-d897-3ea6f742cc05"
lr.intercept_
# + id="SsEk6gtx0fum" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="62a3a9c5-b106-4e3e-a98d-ebd3c35b6515"
#Define a function to make new predictions and explain the model coefficient.
#rent price = # of bathrooms * 2454.36 + 612.41
def rent_price(bath):
rent = bath * lr.coef_[0] + lr.intercept_
return rent
rent_price(1)
| module1-regression-1/LS_DS_211_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import panel
from panel.widgets import *
panel.extension()
# -
# ``Panel`` provides a number of widgets which sync their state with the parameter values in a notebook and in a deployed bokeh dashboard.
widget = TextInput(name='A widget', value='A string')
widget
# Changing the text value will automatically update the corresponding parameter:
widget.value
# Updating the parameter value will also update the widget:
# +
widget.width = 300
widget.value = 'ABCDEFG'
# -
# To listen to a parameter change we can call ``widget.param.watch`` with the parameter to watch and a function:
widget.param.watch(print, 'value')
# If we change the ``widget.value`` now, the resulting change event will be printed:
widget.value = 'A'
# In combination with ``Panel`` objects, widgets make it possible to build interactive dashboards and visualizations very easily.
# ## Laying out widgets
# To compose multiple widgets they can be added to a WidgetBox, which also allows defining a fixed width. To learn more about laying out widgets and panels see the [layout user guide](Layouts.ipynb).
slider = FloatSlider(name='Another widget', width=200)
WidgetBox(widget, slider, width=200)
# ## Supported widgets
# ### TextInput
#
# Allows entering arbitrary text:
text = TextInput(name='A text widget', value='Some text', width=300)
text
# ### LiteralInput
#
# ``LiteralInput`` allows entering any Python literal using a text entry box. Optionally a ``type`` may be declared to validate the literal before updating the parameter.
literal = LiteralInput(name='Type Python literal here:', value={'key': [1, 2, 3]}, type=dict)
literal
# Returns a Python literal:
literal.value
# ### DatetimeInput
#
# ``DatetimeInput`` allows entering a ``datetime`` object as a string. Optionally a ``format`` to change the formatting and input parsing may be declared and ``start`` and ``end`` values may be defined to validate the bounds.
# +
import datetime as dt
dt_input = DatetimeInput(name='Type a datetime string here:', value=dt.datetime(2018, 9, 23), end=dt.datetime(2018, 9, 30), format='%Y-%m-%d %H:%M:%S')
dt_input
# -
# The widget will parse the string that is entered as long as it conforms to the specified ``format`` and return the value as a datetime object:
dt_input.value
# ### Select
# ``Select`` allows choosing between the specified options which may be of any type:
select = Select(name='Select:', options=['A', 1], value=1)
select
select.value
select.value = 'A'
# ### MultiSelect
# MultiSelect allows selecting multiple values which may be of any type:
multi_select = MultiSelect(options=['A', 1.2, 1, 0j], value=[1.2, 1], name='MultiSelect')
multi_select
multi_select.value = ['A', 1, 0j]
# ### DatePicker
# The DatePicker allows selecting a date using arrow keys and a date selection widget that pops up:
date_picker = DatePicker(name='Date Picker', value=dt.datetime(2017, 1, 2), start=dt.datetime(2017, 1, 1))
date_picker
# Just like the ``DatetimeInput`` widget it returns a datetime object:
date_picker.value
# ### DateRangeSlider
# DateRangeSlider allows selecting a range of dates:
date_range = DateRangeSlider(start=dt.datetime(2017, 1, 1), end=dt.datetime(2017, 1, 5),
value=(dt.datetime(2017, 1, 2), dt.datetime(2017, 1, 3)), name='DateRange')
date_range
# The value is returned as a tuple of datetimes:
date_range.value
# ### Checkbox
# A ``Checkbox`` allows toggling between true and false states:
checkbox = Checkbox(name='Checkbox', height=15, value=False)
checkbox
checkbox.value
# ### RangeSlider
# A RangeSlider allows selecting a range of numeric values:
range_slider = RangeSlider(name='Range Slider', start=0, end=10, value=(2, 10))
range_slider
# The current range is returned as a tuple of the lower and upper bound:
range_slider.value
# ### FloatSlider
# A FloatSlider allows defining a single float value within a certain range and allows customizing the step:
float_slider = FloatSlider(name='Slider', start=0, end=10, value=3.14, step=0.1)
float_slider
float_slider.value
# ### IntSlider
# A FloatSlider allows selecting a single integer value within a certain range and allows customizing the (integer) step:
int_slider = IntSlider(name='Slider', start=0, end=15, value=5, step=5)
int_slider
int_slider.value
# ### DiscreteSlider
# A DiscreteSlider allows selecting between any number of discrete numeric values defined via the ``options``:
discrete_slider = DiscreteSlider(options=[0.1, 1, 3.14, 10, 100], value=1, name='Discrete Slider')
discrete_slider
discrete_slider.value
# ### Button
button = Button(name='Click me', button_type='danger')
button
button.clicks
# ### Toggle
# A ``Toggle`` button allows toggling between boolean True and False states:
toggle = Toggle(name='Toggle me', active=True, button_type='primary')
toggle
# The current state is available as the ``active`` parameter value:
toggle.active
# ### Toggle Groups
# A `ToggleGroup` is a group of `widgets` which can be switched 'on' or 'off' (True,False)
#
# Two types of widgets are available. `ToggleGroup` accept a `widget_type` argument which can be set to :
# - **button** (default)
# - **box**
#
# Two different behaviors are available through `behavior` argument:
# - **check** (default) : Any number of widgets can be selected. In this case `value` is a list of objects
# - **radio** : One and only one widget is switched on. In this case `value` is an object
options_list = [1,2,3]
options_dict = {'a':1,'b':2,'c':3}
toggle_group_check = ToggleGroup(options = options_list)
toggle_group_check.value = [2,3]
panel.Column(
toggle_group_check,
panel.Row(
ToggleGroup(options=options_dict, value=2, widget_type='button', behavior='radio'),
ToggleGroup(options=options_list, value=1, widget_type='box', behavior='radio'),
),
panel.Row(
ToggleGroup(options=options_dict, value=[2,3], widget_type='button', behavior='check'),
ToggleGroup(options=options_dict, value=[], widget_type='box', behavior='check')
),
toggle_group_check.value
)
# ## Player
#
# The Player widget allows playing and skipping through a number of frames defined by the ``length``.
player = Player(length=100)
player
# The current value of the player widget can be read out:
player.value
# All the Player state including the ``length``, ``value``, ``interval`` and ``loop_policy`` can be set dynamically:
player.loop_policy = 'loop'
player.length = 20
# ## CrossSelector
# The CrossSelector is a more powerful alternative to the ``MultiSelect`` widget. It allows selecting items by moving them between two lists with a set of buttons. It also provides query fields, which support regex expressions, that allow filtering the selected and unselected items to move them in bulk.
cross_select = CrossSelector(options=[1, 2, 3, 4, 'A', 'B', 'C', 'D'], value=[3, 'A'])
cross_select
# Like most other widgets the selected values can be accessed on the value parameter:
cross_select.value
| examples/user_guide/Widgets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
def fit(X_train,y_train):
result = {}
class_values = set(y_train) #Gives distinct Class Values
for current_class in class_values:
result[current_class] = {}
result['total_data'] = len(y_train)
#Getting Rows of a particular Class
current_class_rows = (y_train == current_class) #Boolean Array
X_train_current = X_train[current_class_rows]
y_train_current = y_train[current_class_rows]
result[current_class]["total_count"] = len(y_train_current)
num_features = X_train.shape[1]
for j in range(1,num_features+1): #instead of 0,1,2,3,4 we want 1,2,3,4,5
result[current_class][j] = {}
all_possible_values = set(X_train[:,j-1]) #x^j #j-1 because starting from 1
for current_value in all_possible_values:
result[current_class][j][current_value] = (X_train[:,j-1] == current_value).sum() #Boolean,sum=count of current value
return result
def probability(dictionary,x,current_class):
#output = 1
#class_prob = dictionary[current_class][total_count] / dictionary['total_data']
#output = dictionary[current_class][total_count] / dictionary['total_data']
output = np.log(dictionary[current_class]['total_count']) - np.log(dictionary['total_data'])
num_features = len(dictionary[current_class].keys()) - 1 #One is total_count
for j in range(1,num_features+1):
xj = x[j-1]
count_current_class_with_value_xj = dictionary[current_class][j][xj] + 1
count_current_class = dictionary[current_class]['total_count'] + len(dictionary[current_class][j].keys())
#current_xj_prob = count_current_class_with_value_xj / count_current_class
current_xj_prob = np.log(count_current_class_with_value_xj) - np.log(count_current_class)
#output *= current_xj_prob
output += current_xj_prob
return output
def predictSinglePoint(dictionary,x):
classes = dictionary.keys()
#best_p = -1000
#best_class = -1
firstRun = True
for current_class in classes:
if(current_class=='total_data'):
continue
p_current_class = probability(dictionary,x,current_class)
if(firstRun or p_current_class>best_p):
best_p = p_current_class
best_class = current_class
firstRun=False
return best_class
def predict(dictionary ,X_test): #result = dictionary
y_pred = []
for x in X_test:
x_class = predictSinglePoint(dictionary,x)
y_pred.append(x_class)
return y_pred
# # Applying to iris
def makeLabelled(column):
second_limit = column.mean()
first_limit = 0.5 * second_limit
third_limit = 1.5 * second_limit
for i in range(len(column)):
if(column[i]<first_limit):
column[i] = 0
elif(column[i]<second_limit):
column[i] = 1
elif(column[i]<third_limit):
column[i] = 2
else:
column[i] = 3
from sklearn import datasets
iris = datasets.load_iris()
X = iris.data
y = iris.target
for i in range(0,X.shape[-1]):
X[:,i] = makeLabelled(X[:,i])
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y)
dictionary = fit(X_train,y_train)
y_pred = predict(dictionary,X_test)
from sklearn.metrics import classification_report,confusion_matrix
print(classification_report(y_test,y_pred))
print(confusion_matrix(y_test,y_pred))
| 12. Implementing Naive Bayes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (nilmtk-env)
# language: python
# name: nilmtk-env
# ---
# + [markdown] toc="true"
# # Tabla de Contenidos
# 1 - Preparación de datos
#
# 2 - Canalización de datos
#
# 2.1 - Selección de aparatos
#
# 2.2 - Selección de secuencias de tiempo
#
# 2.3 - Selección de casas
#
# 2.4 - Tratamiento de datos desequilibrados: selección de ventanas de datos agregados
#
# 2.5 - Datos agregados sintéticos
#
# 3 - Estandarización de los datos de entrada (datos agregados)
#
# 4 - Datos de salida (hora de inicio, hora de finalización y potencia media)
#
# 5 - Puntuaciones - evaluación del rendimiento de la red neuronal
#
# 6 - Estrategia de implementación para el aumento de datos en tiempo real
#
# 7 - Arquitectura de red
#
# 8 - Función de pérdida y optimizador
#
# 8.1 - Función de pérdida
#
# 8.2 - Optimizador
#
# 9 - Experimentando con arquitecturas ConvNet
#
# 9.1 - Una capa convolucional
#
# 10 - Dos capas convolucionales
#
# 10.1 - Resultados para el microondas
# -
# Este cuaderno es el informe sobre la implementación de redes neuronales convolucionales aplicadas a la desagregación de energía. Todos los códigos están en el repositorio: https://github.com/tperol/neuralnilm. No transferimos todos los códigos a este depósito final para mantenerlo limpio. Los datos también se pueden encontrar en https://github.com/tperol/neuralnilm. Algunos de los códigos (principalmente preprocesamiento de los datos, aunque se hizo para otro conjunto de datos) se han bifurcado de <NAME> github Repository Neuralnilm (https://github.com/JackKelly/neuralnilm). Hemos implementado aquí todas las arquitecturas de redes neuronales probadas, así como el preprocesamiento y las métricas de puntuación. El preprocesamiento de datos se ha editado ligeramente del código de <NAME> para alimentar la red neuronal a través de un generador de python que aumenta los datos en la CPU mientras la GPU entrena la red (vea los detalles más adelante).
# +
# for plotting purposes
# %matplotlib inline
from matplotlib.pylab import plt
from matplotlib import rcParams
dark_colors = ["#A51C30", "#808080",
(0.8509803921568627, 0.37254901960784315, 0.00784313725490196),
(0.4588235294117647, 0.4392156862745098, 0.7019607843137254),
(0.9058823529411765, 0.1607843137254902, 0.5411764705882353),
(0.4, 0.6509803921568628, 0.11764705882352941),
(0.9019607843137255, 0.6705882352941176, 0.00784313725490196),
(0.6509803921568628, 0.4627450980392157, 0.11372549019607843),
(0.4, 0.4, 0.4)]
# dark_colors = ['#A51C30', '#808080']
rcParams['figure.figsize'] = (13, 6)
rcParams['figure.dpi'] = 150
rcParams['axes.color_cycle'] = dark_colors
rcParams['lines.linewidth'] = 2
rcParams['axes.facecolor'] = "white"
rcParams['axes.titlesize'] = 20
rcParams['axes.labelsize'] = 17.5
rcParams['xtick.labelsize'] = 15
rcParams['ytick.labelsize'] = 15
rcParams['legend.fontsize'] = 17.5
rcParams['patch.edgecolor'] = 'none'
rcParams['grid.color']="gray"
rcParams['grid.linestyle']="-"
rcParams['grid.linewidth'] = 0.3
rcParams['grid.alpha']=1
rcParams['text.color'] = "444444"
rcParams['axes.labelcolor'] = "444444"
rcParams['ytick.color'] = "444444"
rcParams['xtick.color'] = "444444"
# -
# # Preparación de datos
# Tenemos datos de energía para 6 casas de medidores principales que miden el consumo de energía agregado de la casa y de cada electrodoméstico por separado. Por ejemplo, aquí mostramos los datos agregados (suma de todos los electrodomésticos de la casa) para la casa 2.
# +
from __future__ import print_function
import nilmtk
from nilmtk.utils import print_dict
from nilmtk import DataSet
# import dataset
dataset = DataSet('./redd_data/redd.h5')
dataset.set_window("2011-04-01", "2011-05-01")
# look at house 2
BUILDING = 2
elec = dataset.buildings[BUILDING].elec
# plot mains
elec.mains().plot(kind = 'sum');
# -
# Una contribución a estos datos agregados son los ciclos del refrigerador que se muestran a continuación.
elec['fridge'].plot();
# Por supuesto, hay muchos otros aparatos que contribuyen a los datos agregados observados anteriormente. La lista de electrodomésticos enumerados en esta casa se muestra aquí:
elec.submeters().appliances
# La suma de todos estos dispositivos está altamente correlacionada con los datos agregados.
corr = elec.correlation_of_sum_of_submeters_with_mains()
print
print('------------------------------------------------------------------------------')
print('the correlation between the sum of the submeters and the main meter is:', corr)
print('------------------------------------------------------------------------------')
# Esto lo podemos visualizar mirando los contadores principales de un día y la suma de todos los electrodomésticos de un día.
dataset = DataSet('./redd_data/redd.h5')
dataset.set_window("2011-04-19", "2011-04-20")
dataset.buildings[2].elec.mains().plot(kind= 'sum');
plt.figure(figsize = (16,8))
dataset.buildings[2].elec.submeters().plot(kind = 'sum');
# La correlación es alta porque los 9 electrodomésticos enumerados anteriormente consumen una proporción significativa de la energía y representan un rango de "firmas" de energía diferentes. Aquí hay una firma del refrigerador y una firma del microondas.
elec['fridge'].get_activations()[0].plot();
elec['microwave'].get_activations()[0].plot();
# Obviamente, la firma del microondas es una imagen de corta duración, mientras que el ciclo del refrigerador es un poco más largo. En este estudio tratamos de recuperar de los datos agregados las veces que se enciende un electrodoméstico, las veces que se apaga y el consumo medio de energía en cada ciclo. A esto lo llamamos el método del rectángulo. Esencialmente, queremos recuperar de los datos agregados, este tipo de representación:
dataset.buildings[2].elec.plot_when_on()
# # Canalización de datos
# ## Selección de electrodomésticos
# Entrenamos cada red neuronal por dispositivo. Esto es diferente de los métodos CO y FHMM.
# ## Selección de secuencias de tiempo
#
# Reducimos el muestreo de los medidores principales y los submedidores a 6 muestras por segundo para que las secuencias agregadas y submétricas estén correctamente alineadas. Descartamos cualquier activación que tenga una duración inferior a un umbral para evitar picos falsos.
# ## Seleccionando casas
#
# Elegimos entrenar el algoritmo en la casa 1, 2, 3 y 6 y probar los datos en la casa 6.
# ## Manejo de datos desequilibrados: selección de ventanas de datos agregados
# Primero extraemos usando las bibliotecas NILMTK las activaciones del dispositivo (aquí el refrigerador) en la serie de tiempo. Concatenamos las series de tiempo de la casa 1, 2, 3 y 6 para el conjunto de entrenamiento y probaremos en la casa 5. Alimentamos nuestro algoritmo de red neuronal (detallado más adelante) mini lotes balanceados de secuencias de datos agregados en los que el frigorífico está activado y secuencias en las que no está activado. Esta es una forma de lidiar con datos desequilibrados: hay más secuencias en las que el refrigerador no está activado que secuencias con el refrigerador activado. La mayor parte de la canalización de datos utilizada se toma prestada de https://github.com/JackKelly/neuralnilm.
# ## Datos agregados sintéticos
# Usamos el método de <NAME> para crear datos sintéticos. Para crear una sola secuencia de datos sintéticos, comenzamos con dos vectores de ceros: un vector se convertirá en la entrada a la red; el otro se convertirá en el objetivo. La longitud de cada vector define el "ancho de la ventana" de datos que ve la red. Pasamos por cinco clases de dispositivos y decidimos si agregar o no una activación de esa clase a la secuencia de entrenamiento. Hay un 50 % de posibilidades de que el dispositivo de destino aparezca en la secuencia y un 25 % de posibilidades de que cada uno de los demás dispositivos "distractores". Para cada clase de dispositivo seleccionada, seleccionamos aleatoriamente una activación de dispositivo y luego elegimos aleatoriamente dónde agregar esa activación en el vector de entrada. Los dispositivos distractores pueden aparecer en cualquier parte de la secuencia (incluso si esto significa que solo una parte de la activación se incluirá en la secuencia). La activación del dispositivo de destino debe estar completamente contenida dentro de la secuencia (a menos que sea demasiado grande para caber).
#
# Ejecutamos redes neuronales con y sin datos agregados sintéticos. Descubrimos que los datos sintéticos actúan como reguladores, mejoran los puntajes en la casa de uso.
#
# Todos los códigos que realizan la operación descrita en este apartado 2 se encuentran en https://github.com/tperol/neuralnilm.
# +
# here are the code that perform the preprocessing of the mini batches
import nilmtk
from nilmtk.utils import print_dict
from nilmtk import DataSet
from neuralnilm.data.loadactivations import load_nilmtk_activations
from neuralnilm.data.syntheticaggregatesource import SyntheticAggregateSource
from neuralnilm.data.realaggregatesource import RealAggregateSource
from neuralnilm.data.stridesource import StrideSource
from neuralnilm.data.datapipeline import DataPipeline
from neuralnilm.data.processing import DivideBy, IndependentlyCenter
# ------------
# create dictionary with train, unseen_house, unseen_appliance
# ------------
def select_windows(train_buildings, unseen_buildings):
windows = {fold: {} for fold in DATA_FOLD_NAMES}
def copy_window(fold, i):
windows[fold][i] = WINDOWS[fold][i]
for i in train_buildings:
copy_window('train', i)
copy_window('unseen_activations_of_seen_appliances', i)
for i in unseen_buildings:
copy_window('unseen_appliances', i)
return windows
def filter_activations(windows, activations):
new_activations = {
fold: {appliance: {} for appliance in APPLIANCES}
for fold in DATA_FOLD_NAMES}
for fold, appliances in activations.iteritems():
for appliance, buildings in appliances.iteritems():
required_building_ids = windows[fold].keys()
required_building_names = [
'UK-DALE_building_{}'.format(i) for i in required_building_ids]
for building_name in required_building_names:
try:
new_activations[fold][appliance][building_name] = (
activations[fold][appliance][building_name])
except KeyError:
pass
return activations
NILMTK_FILENAME = './redd_data/redd.h5'
SAMPLE_PERIOD = 6
STRIDE = None
APPLIANCES = ['fridge']
WINDOWS = {
'train': {
1: ("2011-04-19", "2011-05-21"),
2: ("2011-04-19", "2013-05-01"),
3: ("2011-04-19", "2013-05-26"),
6: ("2011-05-22", "2011-06-14"),
},
'unseen_activations_of_seen_appliances': {
1: ("2011-04-19", None),
2: ("2011-04-19", None),
3: ("2011-04-19", None),
6: ("2011-05-22", None),
},
'unseen_appliances': {
5: ("2011-04-19", None)
}
}
# get the dictionary of activations for each appliance
activations = load_nilmtk_activations(
appliances=APPLIANCES,
filename=NILMTK_FILENAME,
sample_period=SAMPLE_PERIOD,
windows=WINDOWS
)
# ------------
# get pipeline for the fridge example
# ------------
num_seq_per_batch = 16
target_appliance = 'fridge'
seq_length = 512
train_buildings = [1, 2, 3, 6]
unseen_buildings = [5]
DATA_FOLD_NAMES = (
'train', 'unseen_appliances', 'unseen_activations_of_seen_appliances')
filtered_windows = select_windows(train_buildings, unseen_buildings)
filtered_activations = filter_activations(filtered_windows, activations)
synthetic_agg_source = SyntheticAggregateSource(
activations=filtered_activations,
target_appliance=target_appliance,
seq_length=seq_length,
sample_period=SAMPLE_PERIOD
)
real_agg_source = RealAggregateSource(
activations=filtered_activations,
target_appliance=target_appliance,
seq_length=seq_length,
filename=NILMTK_FILENAME,
windows=filtered_windows,
sample_period=SAMPLE_PERIOD
)
# ------------
# needed to rescale the input aggregated data
# rescaling is done using the a first batch of num_seq_per_batch sequences
sample = real_agg_source.get_batch(num_seq_per_batch=1024).next()
sample = sample.before_processing
input_std = sample.input.flatten().std()
target_std = sample.target.flatten().std()
# ------------
pipeline = DataPipeline(
[synthetic_agg_source, real_agg_source],
num_seq_per_batch=num_seq_per_batch,
input_processing=[DivideBy(input_std), IndependentlyCenter()],
target_processing=[DivideBy(target_std)]
)
# ------------
# create the validation set
# ------------
num_test_seq = 101
X_valid = np.empty((num_test_seq*num_seq_per_batch, seq_length))
Y_valid = np.empty((num_test_seq*num_seq_per_batch, 3))
for i in range(num_test_seq):
(x_valid,y_valid) = pipeline.train_generator(fold = 'unseen_appliances', source_id = 1).next()
X_valid[i*num_seq_per_batch: (i+1)*num_seq_per_batch,:] = x_valid[:,:,0]
Y_valid[i*num_seq_per_batch: (i+1)*num_seq_per_batch,:] = y_valid
X_valid = np.reshape(X_valid, [X_valid.shape[0],X_valid.shape[1],1])
# -
# # Estandarización de los datos de entrada (datos agregados)
# Un paso típico en la tubería de datos de la red neuronal es la estandarización de los datos. Para cada secuencia de 512 muestras (= 85 segundos) restamos la media para centrar la secuencia. Además, cada secuencia de entrada se divide por la desviación estándar de una muestra aleatoria en el conjunto de entrenamiento. En este caso no podemos dividir cada secuencia por su propia desviación estándar porque borraría información sobre la escala de la señal.
#
# Esto se hace a través de:
# needed to rescale the input aggregated data
# rescaling is done using the a first batch of num_seq_per_batch sequences
sample = real_agg_source.get_batch(num_seq_per_batch=1024).next()
sample = sample.before_processing
input_std = sample.input.flatten().std()
target_std = sample.target.flatten().std()
# ------------
# # Datos de salida (hora de inicio, hora de finalización y potencia media)
# La salida de la red neuronal es de 3 neuronas: hora de inicio, hora de finalización y potencia media. Reescalamos el tiempo al intervalo [0,1]. Por lo tanto, si la nevera comienza en medio de las secuencias de entrada, la salida de la primera neurona es 0,5. Si se detiene después del final de la ventana de entrada, la salida de la segunda neurona se establece en 1. La tercera neurona es la potencia promedio durante el período de activación. Por supuesto, esto se establece en 0 cuando no se activa durante la secuencia de entrada. También procesamos los datos mediante la configuración de una hora de inicio inferior a 0 a 0 y una hora de finalización superior a 1 a 1. Creamos un umbral de potencia promedio establecido en 0,1 que indica si el dispositivo estaba activo o no (por debajo del umbral en el que el dispositivo está activo). se considera apagado, encima se considera encendido).
#
# Aquí mostramos como ejemplo los datos de entrada y la salida calculada por una red entrenada. Comparamos esto con la activación real del dispositivo. <img src='./figures/output_example.png'>
#
# Como podemos ver aquí, la red hace un muy buen trabajo al detectar la activación del refrigerador. La línea roja son los datos agregados. En la región plana sería imposible detectar la activación de la nevera con el ojo humano. Tendemos a poner una activación en la región del paso. Sin embargo, la red hace una predicción muy precisa de la activación de la nevera.
# # Puntuaciones - evaluación del rendimiento de la red neuronal
# Debido a la dimensión de la salida, elegimos métricas de puntuación de clasificación. Cuando la hora de inicio y la hora de finalización son 0, lo llamamos negativo. También llamamos negativo si el promedio de potencia es inferior al umbral. En caso contrario es positivo (el aparato está activado). Llamamos TP verdadero positivo, TN verdadero negativo, FP falso positivo y FN falso negativo. Las diversas métricas/puntuaciones utilizadas en este estudio son
#
# $$ recall = \frac{TP}{TP + FN} $$
#
# $$ precision = \frac{TP}{TP + FP} $$
#
# $$ F1 = 2 * \frac{precision* recall}{precision + recall} $$
#
# $$ accuracy = \frac{TP + TN}{P + N} $$
#
# donde P es el número de positivos y N el número de negativos.
# El código para calcular se puede encontrar en neuralnilm/scores y se reproduce parcialmente aquí.
def scores(Y_pred, Y_test, activation_threshold = 0.1 ,plot_results= True, print_results = False):
"""
a function that computes the classification scores with various metrics
return: dictionary with the various scores
"""
# post process the data
np.putmask(Y_pred[:,0], Y_pred[:,0] <=0, 0)
np.putmask(Y_pred[:,1], Y_pred[:,1] >=1, 1)
np.putmask(Y_pred[:,0],Y_pred[:,1] < Y_pred[:,0],0)
np.putmask(Y_pred[:,1],Y_pred[:,1] < Y_pred[:,0],0)
np.putmask(Y_pred[:,1],Y_pred[:,2] < activation_threshold,0)
np.putmask(Y_pred[:,0],Y_pred[:,2] < activation_threshold,0)
# find negative in prediction
pred_negatives = (Y_pred[:,0] ==0) &(Y_pred[:,1] ==0)
pred_positives = ~pred_negatives
obs_negatives = (Y_test[:,0] ==0) &(Y_test[:,1] ==0)
obs_positives = ~obs_negatives
TP = obs_positives[pred_positives].sum()
FN = obs_positives[pred_negatives].sum()
TN = obs_negatives[pred_negatives].sum()
FP = obs_negatives[pred_positives].sum()
recall = TP / float(TP + FN)
precision = TP / float(TP+ FP)
f1 = 2* precision*recall / (precision + recall)
accuracy = (TP + TN)/ float(obs_negatives.sum() +obs_positives.sum() )
if print_results:
print('number of Predicted negatives:',pred_negatives.sum() )
print('number of Predicted positives:',pred_positives.sum() )
print('number of Observed negatives:', obs_negatives.sum() )
print('number of Observed positives:', obs_positives.sum() )
print('f1:', f1)
print('precision :' ,precision)
print('recall : ', recall)
print('accuracy:', accuracy)
results = {
'accuracy': accuracy,
'f1_score': f1,
'precision': precision,
'recall_score': recall}
if plot_results:
pd_results = pd.DataFrame.from_dict(results, orient = 'index')
pd_results = pd_results.transpose()
sns.barplot(data = pd_results)
return results
# # Estrategia de implementación para el aumento de datos en tiempo real
# Mientras que la red neuronal ejecuta una **NVIDIA GeForce GT 750M (GPU)**, mantenemos la **CPU** ocupada haciendo el aumento de datos en tiempo real (cargar datos agregados, crear datos sintéticos, preprocesar el mini lote para ser alimentado a la red neuronal). Para esto creamos un **generador de python** que crea una cola de 50 mini-lotes y los alimenta sucesivamente a la GPU para su entrenamiento.
#
# La clase de canalización se puede encontrar en neuralnilm.data.datapiline en https://github.com/tperol/neuralnilm y se reproduce parcialmente aquí. Hacemos lo mismo para generar el conjunto de validación y prueba.
def train_generator(self, fold='train', enable_all_appliances=False,
source_id=None, reset_iterator=False,
validation=False ):
"""
class method that yield generator for training the neural network
parameters:
----------
fold: 'train' for training or 'unseen_appliance' for testing/validation
source_id: if None we use both real and synthetic data with 50:50 ratio
if 1 , only use the real data
if 0, only use the synthetic data
output:
---------
generator (X_train,Y_train): a tuple with X_train being the mini-batch for training
and Y_train being the mini-batch of labels
"""
while 1:
batch_iter = self.get_batch(fold, enable_all_appliances, source_id, reset_iterator,validation)
X_train = batch_iter.input
input_dim = X_train.shape[1]
Y_train = self._get_output_neurons(batch_iter)
yield (np.reshape(X_train, [self.num_seq_per_batch, input_dim, 1]), Y_train.astype(np.float32))
def _get_output_neurons(self, new_batch):
"""
class method that generates the indices needed for the generator
Once the network has seen all the sample, the algorithm feed again the mini-batch
by reinitialization of the generator (there is no StopIteration error raised)
Suffling of the mini-batch is the default strategy implemented
"""
batch_size = new_batch.target.shape[0]
neural_net_output = np.empty((batch_size, 3))
for b in range(batch_size):
seq = new_batch.target[b]
# case 1 and 2: if the signal start at 0
if seq[0] > 0:
start = 0
stop_array = np.where(seq > 0)[0]
# case 2: signal stops after 1
# set stop to the last element
if len(stop_array) == 0:
stop = seq[-1]
# case 1: signal stops before 1
else:
stop = stop_array[-1]
# calculate avg power
avg_power = np.mean(seq[start:stop + 1])
# case 3: signal starts after 0 and before 1
else:
start_array = np.where(seq > 0)[0]
if len(start_array) == 0:
# case 5: there is no signal in the window
start = 0
stop = 0
avg_power = 0
else:
start = start_array[0]
# find stop
stop_array = np.where(seq > 0)[0]
# case 4: signal stops after 1
# set to the last element
if len(stop_array) == 0:
stop = seq[-1]
else:
stop = stop_array[-1]
avg_power = np.mean(seq[start:stop + 1])
start = start / float(new_batch.target.shape[1] - 1)
stop = stop / float(new_batch.target.shape[1] - 1)
if stop < start:
raise ValueError("start must be before stop in sequence {}".format(b))
neural_net_output[b, :] = np.array([start, stop, avg_power])
return neural_net_output
# # Red de arquitectura
# Usamos una red neuronal convolucional (ConvNet) para aprovechar la invariancia de traducción. Queremos que ConvNet reconozca la activación del dispositivo de destino en cualquier lugar de la secuencia. Para este proyecto hemos probado múltiples arquitecturas que se reportan más adelante. Todas estas arquitecturas tienen una primera capa convolucional de tamaño de filtro 3 y paso 1. Hemos jugado tanto con el tamaño del filtro como con la cantidad de filtros de salida en la primera capa. Descubrimos que 16 filtros es un número razonable: aumentar el número de filtros en la primera capa no mejoró significativamente las puntuaciones.
#
# La mejor red neuronal que encontramos consiste en
#
# * Capa de entrada: un canal y duración de 512 muestras
# * Capa convolucional 1D (tamaño de filtro = 3, zancada = 1, número de filtros = 16, función de activación = relu, modo de borde = válido, inicialización de peso = distribución normal)
# * Capa completamente conectada (N = 1024, función de activación = relu, inicialización de peso = distribución normal)
# * Capa completamente conectada (N = 512, función de activación = relu, inicialización de peso = distribución normal)
# * Capa totalmente conectada (N= 3, función de activación = relu)
#
# La salida tiene 3 neuronas activadas por una función de activación relu ya que la salida no puede ser negativa. Hemos probado otras redes que se informan más adelante en este cuaderno. Sin embargo, este es el diseño del mejor que encontramos.
# # Función de pérdida y optimizador
# ## Función de pérdida
# Dado que las neuronas de salida abarcan el eje real, no hay otra opción que usar una norma L2 para la función de pérdida. Esto es (hora de inicio pronosticada - hora de inicio real) 2 + (hora de finalización pronosticada - hora de finalización real) 2 + (potencia promedio pronosticada - potencia promedio real) 2 . La función de pérdida total es la suma de la función de pérdida de toda la muestra en un mini lote.
# ## Optimizador
# Hemos probado varios optimizadores para encontrar el mejor. Usamos un clásico **Descenso de gradiente estocástico** para actualizar los pesos donde alimentamos un mini lote elegido al azar a la red neuronal y luego actualizamos cada peso
#
# $$w_j = w_j - \eta \frac{\parcial L}{\parcial w_j} $$
#
# donde $L$ es la función de pérdida evaluada para el mini lote dado. El gradiente de la función de pérdida se calcula usando el algoritmo de retropropagación (no detallado aquí por simplicidad). En cada época, disminuimos la tasa de aprendizaje $eta$ para permitir que el algoritmo converja hacia un mínimo local.
#
# Probamos una variación de SGD usando el **método de impulso**. Este método tiene alguna interpretación física donde $\mu$ es el coeficiente de fricción. En este caso los pesos se actualizan usando
#
# $$w_j = w_j + \mu v - \eta \frac{\parcial L}{\parcial w_j} $$
#
# donde $v$ es la velocidad. Otra implementación probada es el **momento de Nesterov** en el que, en una posición dada en el panorama del peso, miramos un paso adelante con el momento y luego evaluamos el gradiente allí para calcular el nuevo valor del peso. Un pseudocódigo para esto sería
w_ahead = w + mu * v
# evaluate dw_ahead
v = mu * v - learning_rate * dw_ahead
x += v
#
# Experimentando encontramos que el mejor optimizador es Adam (http://arxiv.org/pdf/1412.6980v8.pdf). Un pseudocódigo para Adam es
m = beta1*m + (1-beta1)*dw
v = beta2*v + (1-beta2)*(dw**2)
w += - learning_rate * m / (np.sqrt(v) + eps)
# donde *dw* es el gradiente de la función perdida con respecto al peso considerado y *w* el peso considerado. Los hiperparámetros de este optimizador son beta1, beta2 y eps. Configuramos beta_1=0.9, beta_2=0.999, epsilon=1e-08 y la tasa de aprendizaje = 1e-3.
# # Experimentando con arquitecturas ConvNet
# Para la implementación de ConvNet usamos Keras (http://keras.io). Esta es una biblioteca implementada sobre Theano y Tensorflow (en este caso, usamos Theano para aprovechar la GPU, el entrenamiento de GPU aún no está disponible en Mac OS con TensorFlow). Aquí mostramos el código ejecutado usando GPU y Keras para entrenar nuestra mejor ConvNet.
# +
# import Keras related libraries
from keras.layers import Input, Dense, Flatten, MaxPooling1D, AveragePooling1D, Convolution1D
from keras.models import Model
import keras.callbacks
from keras.callbacks import ModelCheckpoint
import time
from keras.models import model_from_json
import pickle
# ------------
exp_number = 13
output_architecture = './tmpdata/convnet_architecture_exp' + str(exp_number) + '.json'
best_weights_during_run = './tmpdata/weights_exp' + str(exp_number) + '.h5'
final_weights = './tmpdata/weights_exp' + str(exp_number) + '_final.h5'
loss_history = './tmpdata/history_exp' + str(exp_number) + '.pickle'
# ------------
# ------------
# a class used to record the training and validation loss
# at the end of each epoch
class LossHistory(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.train_losses = []
self.valid_losses = []
def on_epoch_end(self, epoch, logs = {}):
self.train_losses.append(logs.get('loss'))
self.valid_losses.append(logs.get('val_loss'))
# ------------
# input sequence
input_seq = Input(shape = (seq_length, 1))
# first convolutional layer
conv1_layer = Convolution1D(nb_filter = 16, filter_length = 3, border_mode='valid',
init = 'normal', activation = 'relu')
conv1 = conv1_layer(input_seq)
# flatten the weights
flat = Flatten()(conv1)
# first dense layer
dense1 = Dense(1024, activation = 'relu')(flat)
# second dense layer
dense2 = Dense(512, activation = 'relu', init= 'normal')(dense1)
# output layer
predictions = Dense(3, activation = 'linear')(dense2)
# create the model
model = Model(input=input_seq, output=predictions)
# compile the model -- define the loss and the optimizer
model.compile(loss='mean_squared_error', optimizer='Adam')
# record the loss history
history = LossHistory()
# save the weigths when the vlaidation lost decreases only
checkpointer = ModelCheckpoint(filepath=best_weights_during_run, save_best_only=True, verbose =1 )
# fit the network using the generator of mini-batches.
model.fit_generator(pipeline.train_generator(fold = 'train'), \
samples_per_epoch = 30000, \
nb_epoch = 20, verbose = 1, callbacks=[history, checkpointer],
validation_data = (x_valid,y_valid), max_q_size = 50)
losses_dic = {'train_loss': history.train_losses, 'valid_loss':history.valid_losses}
# save history
losses_dic = {'train_loss': history.train_losses, 'valid_loss':history.valid_losses}
with open(loss_history, 'wb') as handle:
pickle.dump(losses_dic, handle)
print('\n saving the architecture of the model \n')
json_string = model.to_json()
open(output_architecture, 'w').write(json_string)
print('\n saving the final weights ... \n')
model.save_weights(final_weights, overwrite = True)
print('done saving the weights')
print('\n saving the training and validation losses')
print('This was the model trained')
print(model.summary())
# -
# ## Una capa convolucional
# Aquí presentamos los resultados de ConvNet detallados en la celda anterior. Primero cargamos la arquitectura ConvNet, los pesos y el historial de pérdidas de entrenamiento y validación (la red ha sido entrenada previamente mediante GPU).
from keras.models import model_from_json
# load experiments number
exp_number = 13
# load the model architecture
output_architecture = './tmpdata/convnet_architecture_exp' + str(exp_number) + '.json'
# load the weights for the lowest validation loss during training
best_weights_during_run = './tmpdata/weights_exp' + str(exp_number) + '.h5'
# load the final weights at the end of the 20 epochs
final_weights = './tmpdata/weights_exp' + str(exp_number) + '_final.h5'
model = model_from_json(open(output_architecture).read())
# load intermediate or final weights
model.load_weights(best_weights_during_run)
model.compile(loss='mean_squared_error', optimizer='sgd')
# print the summary of the architecture
model.summary()
# load the loss summary (training and validation losses)
import pickle
losses = pickle.load( open('./tmpdata/history_exp' + str(exp_number) + '.pickle' , 'rb'))
# load the test set
test_set = pickle.load( open('./tmpdata/TestSet.pickle', 'rb'))
X_test = test_set['X_test']
Y_test = test_set["Y_test"]
# Here we predict the output from the neural network and show the scores
import neuralnilm.scores
Y_pred = model.predict(X_test)
scores(Y_pred, Y_test)
# Esta es una muy buena puntuación de F1, mucho mejor que con cualquier otro método probado antes. Aquí mostramos la evolución de la pérdida de entrenamiento y la pérdida de validación.
from neuralnilm.scores import plot_loss
plot_loss(losses)
#
# La pérdida de entrenamiento y validación durante el entrenamiento disminuye simuladamente. No hay sobreajuste. Detenemos la simulación después de 20 épocas pero el modelo seguía mejorando. 20 épocas en GPU tomaron alrededor de 1,5 horas.
# # Dos capas convolucionales
# Dado que no estamos sobreajustando, no agregamos ninguna capa de agrupación o abandono a la red anterior. Sin embargo, experimentamos con otra red con una capa convolucional adicional y entrenamos el modelo durante 30 épocas. El modelo es el siguiente:
exp_number = 14
output_architecture = './tmpdata/convnet_architecture_exp' + str(exp_number) + '.json'
best_weights_during_run = './tmpdata/weights_exp' + str(exp_number) + '.h5'
final_weights = './tmpdata/weights_exp' + str(exp_number) + '_final.h5'
model = model_from_json(open(output_architecture).read())
# load intermediate or final weights
model.load_weights(best_weights_during_run)
model.compile(loss='mean_squared_error', optimizer='sgd')
model.summary()
# load the loss summary
import pickle
losses = pickle.load( open('./tmpdata/history_exp' + str(exp_number) + '.pickle' , 'rb'))
# load the test set
test_set = pickle.load( open('./tmpdata/TestSet.pickle', 'rb'))
X_test = test_set['X_test']
Y_test = test_set["Y_test"]
#
# Hay 2 capas convolucionales con 16 filtros y 2 capas densas y la capa de salida. En este caso, los resultados son bastante similares a la red presentada anteriormente.
import neuralnilm.scores
Y_pred = model.predict(X_test)
scores(Y_pred, Y_test)
from neuralnilm.scores import plot_loss
plot_loss(losses)
# ## Resultados para el microondas
# Aquí ejecutamos ConvNet con una capa en el microondas. Entrenamos usando la casa 1 y 2 y probamos en la casa 5.
from keras.models import model_from_json
exp_number = 13
output_architecture = './tmpdata/convnet_architecture_exp' + str(exp_number) + '.json'
best_weights_during_run = './tmpdata/weights_exp' + str(exp_number) + '.h5'
final_weights = './tmpdata/weights_exp' + str(exp_number) + '_final.h5'
model = model_from_json(open(output_architecture).read())
# load intermediate or final weights
model.load_weights(best_weights_during_run)
model.compile(loss='mean_squared_error', optimizer='sgd')
model.summary()
# load the loss summary
import pickle
losses = pickle.load( open('./tmpdata/history_exp' + str(exp_number) + '.pickle' , 'rb'))
# load the test set
test_set = pickle.load( open('./tmpdata/TestSet_microwave.pickle', 'rb'))
X_test_microwave = test_set['X_test']
Y_test_microwave = test_set["Y_test"]
from neuralnilm.scores import scores
Y_pred_microwave = model.predict(X_test_microwave)
scores(Y_pred_microwave, Y_test_microwave)
| Report_convnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (ICML2020)
# language: python
# name: icml2020
# ---
# +
from IPython.core.debugger import set_trace
import numpy as np
import import_ipynb
from environment import *
# -
def heuristicFCFS(num_jobs, num_mc, machines, durations):
machines_ = np.array(machines)
tmp = np.zeros((num_jobs,num_mc+1), dtype=int)
tmp[:,:-1] = machines_
machines_ = tmp
durations_ = np.array(durations)
tmp = np.zeros((num_jobs,num_mc+1), dtype=int)
tmp[:,:-1] = durations_
durations_ = tmp
indices = np.zeros([num_jobs], dtype=int)
# Internal variables
previousTaskReadyTime = np.zeros([num_jobs], dtype=int)
machineReadyTime = np.zeros([num_mc], dtype=int)
placements = [[] for _ in range(num_mc)]
# While...
while(not np.array_equal(indices, np.ones([num_jobs], dtype=int)*num_mc)):
machines_Idx = machines_[range(num_jobs),indices]
durations_Idx = durations_[range(num_jobs),indices]
# 1: Check previous Task and machine availability
mask = np.zeros([num_jobs], dtype=bool)
for j in range(num_jobs):
if previousTaskReadyTime[j] == 0 and machineReadyTime[machines_Idx[j]] == 0 and indices[j]<num_mc:
mask[j] = True
# 2: Competition SPT
for m in range(num_mc):
job = None
duration = 0
for j in range(num_jobs):
if machines_Idx[j] == m and durations_Idx[j] > duration and mask[j]:
job = j
duration = durations_Idx[j]
placements[m].append([job, indices[job]])
previousTaskReadyTime[job] += durations_Idx[job]
machineReadyTime[m] += durations_Idx[job]
indices[job] += 1
break
# time+1
previousTaskReadyTime = np.maximum(previousTaskReadyTime - 1 , np.zeros([num_jobs], dtype=int))
machineReadyTime = np.maximum(machineReadyTime - 1 , np.zeros([num_mc], dtype=int))
return placements
if __name__ == "__main__":
# Import environment
config = Config()
config.machine_profile = "xsmall_default"
config.job_profile = "xsmall_default"
config.reconfigure()
# Configure environment
env = Environment(config)
env.clear()
# Read problem instance
filename = "datasets/inference/dataset_xsmall.data"
with open(filename, "r") as file:
NB_JOBS, NB_MACHINES = [int(v) for v in file.readline().split()]
JOBS = [[int(v) for v in file.readline().split()] for i in range(NB_JOBS)]
#-----------------------------------------------------------------------------
# Prepare the data for modeling
#-----------------------------------------------------------------------------
# Build list of machines. MACHINES[j][s] = id of the machine for the operation s of the job j
machines = [[JOBS[j][2 * s] for s in range(NB_MACHINES)] for j in range(NB_JOBS)]
# Build list of durations. DURATION[j][s] = duration of the operation s of the job j
durations = [[JOBS[j][2 * s + 1] for s in range(NB_MACHINES)] for j in range(NB_JOBS)]
placements = heuristicFCFS(NB_JOBS, NB_MACHINES, machines, durations)
env.step(machines, durations, placements)
print("Makespan: ", env.makespan)
env.plot(save=False)
| heuristic_FCFS.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# # Calculating the *steady* solution for $\eta$ and $\theta$ domain using BVP solvers and upwinding
# \begin{equation}\label{p1introhpa} \Psi _{\eta \eta \tau } = \frac{1}{{R_s }}\Psi _{\eta \eta \eta \eta } + \Psi _{\eta \eta \eta } \Psi _\theta - \Psi _{\eta \eta \theta } \Psi _\eta \\
# \\
# \quad \Psi(\eta=\mp 1) = 0, \qquad \Psi_\eta(\eta=\mp 1) = - sin\theta, \qquad
# -1 \le \eta \le1, \quad 0 \le \theta \le \pi \end{equation}
#
# This is the notebook uses upwinding, **this numerical method is unstable with respect to shooting** and numerically too intensive for the BVP solver to even return an answer. **This code should be avoided.**
using DifferentialEquations
using Plots
using Sundials
using BenchmarkTools
using Calculus
using DelimitedFiles
using Interpolations
using LaTeXStrings
using SparseArrays
using Debugger
#using LSODA
using LinearAlgebra
# In this code the vector **u** is used to store **u** = $[\Psi ... \Psi_\eta...\Psi_{\eta\eta} ... \Psi_{\eta\eta\eta}]$ in a single 1-dimensional vector. The BVPSolver runs perfectly with the latter structure. However the BVPSolver has a few issues if a matrix M is used to store the derivatives of $\Psi$ in columns e.g. $M = [\Psi, \Psi_\eta,\Psi_{\eta\eta}, \Psi_{\eta\eta\eta}]$ this is why the matrix form is not used in this code, interesting the ODE solver works perfectly with the Matrix form $M$.
# +
pyplot()
function Ψ_smallr(θ::Float64, η::Float64 ,r::Float64)
return(-η*(η^2-1)*sin(θ)/2.0 + r*η*(η^2-1)^2.0*(2.0+η^2)*sin(2*θ)/560.0 +
r^2.0*η*(η^2-1)^2*((-591 -2294*η^2 + 161*η^4 + 1428*η^6)*sin(θ) + 3.0*(423 + 166*η^2 - 553*η^4 - 84*η^6)sin(3*θ))/62092800)
end
function Ψ_smallr_d1(θ::Float64, η::Float64 ,r::Float64)
return(-(3.0*η^2-1)*sin(θ)/2.0 + r*(7*η^6-9*η^2+2)*sin(2*θ)/560.0 +
r^2.0*(3*(-197 - 1112*η^2 + 6930*η^4 - 2772*η^6 - 8085*η^8 + 5236*η^10)*sin(θ) + (1269 - 6120*η^2 - 6930*η^4 + 24948*η^6 - 10395*η^8 - 2772*η^10)*sin(3*θ))/62092800.0)
end
function Ψ_smallr_d2(θ::Float64, η::Float64 ,r::Float64)
return(-3.0*η*sin(θ) + r*(42*η^5-18*η)*sin(2*θ)/560.0 +
r^2.0*(3*( - 2224*η + 27720*η^3 - 16632*η^5 - 64680*η^7 + 52360*η^9)*sin(θ) + (- 12240*η - 27720*η^3 + 149688*η^5 - 83160*η^7 - 27720*η^9)*sin(3*θ))/62092800.0)
end
function Ψ_smallr_d3(θ::Float64, η::Float64 ,r::Float64)
return(-3.0*sin(θ) + r*(210*η^4-18)*sin(2*θ)/560.0 +
r^2.0*(3*( - 2224 + 83160*η^2 - 83160*η^4 - 452760*η^6 + 471240*η^8)*sin(θ) + (- 12240 - 83160*η^2 + 748440*η^4 - 582120*η^6 - 249480*η^8)*sin(3*θ))/62092800.0)
end
function Ψ_bigr(θ::Float64, η::Float64)
return(sin(pi*η)*sin(θ)/pi)
end
function Ψ_bigr_d1(θ::Float64, η::Float64)
return(cos(pi*η)*sin(θ))
end
function Ψ_bigr_d2(θ::Float64, η::Float64)
return(-sin(pi*η)*sin(θ)*pi)
end
function Ψ_bigr_d3(θ::Float64, η::Float64)
return(-cos(pi*η)*sin(θ)*pi^2)
end
"""This is the differential equation for the ``\\psi(\\eta)^{(0)}`` term in the series.
``\\Psi_{\\eta \\eta \\eta \\eta
}^{(n)} = R_s( \\Psi_{\\eta}^{(0)} \\Psi _{\\eta \\eta}^{(0)} - \\Psi ^{(0)} \\Psi _{\\eta \\eta
\\eta }^{(0)})``
"""
function steady_diffeq_psi_bvp!(dψ, ψ, p, η) # This works for the BVPsolver psi a vector of vectors
#p[4] psi0 initial condition
#p[3] Δθ
#p[2] is the J value for the number of steps in the J direction
#p[1] is the Reynolds number
#dψ[1:j+1] .= ψ[j+2:2*j+2]
#dψ[j+2:2*j+2] .= ψ[2*j+3:3*j+3]
#dψ[2*j+3:3*j+3] .= ψ[3*j+4:4*j+4]
#dψ[3*j+4:4*j+4] = (ψ[j+2:2*j+2].*(p[3]*ψ[2*j+3:3*j+3])-ψ[3*j+4:4*j+4].*(p[3]*ψ[1:j+1])).*p[1]
r, J, Δθ, psi0 = p
Psietaetaeta = @view ψ[3*J+4:4*J+4] #View are non allocating and are very fast to work with.
Psietaeta = @view ψ[2*J+3:3*J+3]
Psieta = @view ψ[J+2:2*J+2]
Psi = @view ψ[1:J+1]
DPsi4eta = @view dψ[3*J+4:4*J+4]
DPsi3eta = @view dψ[2*J+3:3*J+3]
DPsi2eta = @view dψ[J+2:2*J+2]
DPsi1eta = @view dψ[1:J+1]
Psi[1] = Psi[J+1] = 0.0
#The first 3 derivatives at the next η step are assigned, the last ψ_ηηηη derivative is assigned manually
@. DPsi1eta = Psieta
@. DPsi2eta = Psietaeta
@. DPsi3eta = Psietaetaeta
#Near θ=0 boundary we have to use first order one sided θ derivative for ψ_ηηθ and ψ_θ
DPsietaetaDtheta=(Psietaeta[2]-Psietaeta[1])/Δθ
DPsiDtheta =(Psi[2]-Psi[1])/Δθ
DPsi4eta[1] = (Psieta[1]*DPsietaetaDtheta - Psietaetaeta[1]*DPsiDtheta)*r
#Near θ=π boundary we have to use first order one sided θ derivative for ψ_ηηθ and ψ_θ
DPsietaetaDtheta=(Psietaeta[J+1]-Psietaeta[J])/Δθ
DPsiDtheta=(Psi[J+1]-Psi[J])/Δθ
DPsi4eta[J+1] = (Psieta[J+1]*DPsietaetaDtheta - Psietaetaeta[J+1]*DPsiDtheta)*r
#Upwinding
for j = 2:J #If \psi_eta < 0 then upwind with forward difference
if Psieta[j] < 0.0
DPsietaetaDtheta=(Psietaeta[j+1]-Psietaeta[j])/Δθ
if( j < J )
DPsietaetaDtheta=(-0.5*Psietaeta[j+2]+2*Psietaeta[j+1]-1.5*Psietaeta[j])/Δθ
end
else
DPsietaetaDtheta=(Psietaeta[j] -Psietaeta[j-1])/Δθ
if (j > 2)
DPsietaetaDtheta=( 0.5*Psietaeta[j-2]-2*Psietaeta[j-1]+1.5*Psietaeta[j])/Δθ
end
end
if Psietaetaeta[j] > 0.0
DPsiDtheta=(Psi[j+1]-Psi[j])/Δθ
if j < J
DPsiDtheta=(-0.5*Psi[j+2]+2*Psi[j+1]-1.5*Psi[j])/Δθ
end
else
DPsiDtheta=(Psi[j] -Psi[j-1])/Δθ
if j > 2
DPsiDtheta=( 0.5*Psi[j-2]-2*Psi[j-1]+1.5*Psi[j])/Δθ
end
end
DPsi4eta[j] = (Psieta[j]*DPsietaetaDtheta - Psietaetaeta[j]*DPsiDtheta)*r
end
#Fully expanded loops for the copying this makes the
#@inbounds for i =1:j+1
# dψ[i] = ψ[j+1+i]
# dψ[j+1+i] = ψ[2*j+2+i]
# dψ[2*j+2+i] = ψ[3*j+3+i]
#end
end
function bc_bvp!(residual, u, p, η) # psi[1] is the beginning of the etaspan, and psi[end] is the ending
#p[4] is the initial condition psi0
#p[3] is the matrix A
#p[2] is the J value for the number of steps in the J direction
j=p[2]
@inbounds for i = 1:j+1
residual[i] = u[1][i] # The psi[1] (i.e,. psi^{0}) solution at the beginning of the time span should be 0
residual[j+1+i] = u[1][j+1+i] - p[4][j+1+i] #First derivative of should be -1 at first time step
residual[2*j+2+i] = u[end][i] # the solution at the end of the time span should be 0
residual[3*j+3+i] = u[end][j+1+i] - p[4][j+1+i] #First derivative should be -sinθ at end time step
end
end
# +
const Reynolds_number = 5.0
const J = 64 #Number of steps take in the theta direction from 0 to pi, restart kernel if this value is changed
const Δθ = pi/J
const etaspan=(-1.0,1.0)
Dpsi_eta_eta_Dtheta = ones(J+1); #For the Boundary Value solver Dpsi_eta_eta_Dtheta can be declared as Float and not Real, mult! function can be unstable
Dpsi_Dtheta = ones(J+1);
if (Reynolds_number < 40.0 )
psi_eta = [-sin(j*Δθ) for j =0:J]
psi_eta_eta= [ Ψ_smallr_d2(j*Δθ, -1.0, Reynolds_number) for j =0:J]
psi_eta_eta_eta= [ Ψ_smallr_d3(j*Δθ, -1.0, Reynolds_number) for j =0:J]
else
psi_eta = [-sin(j*Δθ) for j =0:J]
psi_eta_eta= [ Ψ_bigr_d2(j*Δθ, -1.0) for j =0:J]
psi_eta_eta_eta= [ Ψ_bigr_d3(j*Δθ, -1.0) for j =0:J]
end
# +
#The initial conditions psi0 (note here psi0 means psi@t=0) must be sent as a matrix
#This is the ODESolver variable psi0
#Uncomment the code below to solve the ODE IVP problem
#ψ0= hcat(zeros(J+1), psi_eta, psi_eta_eta , psi_eta_eta_eta) #This works for the ODESolver AND BVPSolver
#ψ0 is declared as a Matrix
#p= (Reynolds_number, J, A, ψ0, Dpsi_Dtheta, Dpsi_eta_eta_Dtheta)
#prob = ODEProblem(steady_diffeq_psi!, ψ0, etaspan, p)
#sol = @time solve(prob, reltol=1e-5, alg_hints = [:stiff], Rosenbrock23())
#Dpsi_eta_eta_Dtheta = Array{Real,1}(undef,J+1) #This has to be declared as real type because stiff solvers output reals
#Dpsi_Dtheta = Array{Real,1}(undef,J+1)
#This works fine for R < 10 with Vern 7 but there is a problem with MethodError: no method matching Float64
#ψ0 is declared as a vector, Rosenbrock cannot handle VECTOR when called
#ψ0= vcat(zeros(J+1), psi_eta, psi_eta_eta , psi_eta_eta_eta) #This works for the ODESolver AND BVPSolver
#Dpsi_eta_eta_Dtheta = Array{Real,1}(undef,J+1) #This has to be declared as real type because stiff solvers output reals
#Dpsi_Dtheta = Array{Real,1}(undef,J+1)
#p= (Reynolds_number, J, A, ψ0, Dpsi_Dtheta, Dpsi_eta_eta_Dtheta)
#prob = ODEProblem(steady_diffeq_psi_bvp!, ψ0, etaspan, p)
#sol = @time solve(prob, reltol=1e-5, alg_hints = [:stiff], Vern7())
#This is the BVP solver, works with J =20 upto R=30 with small R bc, and upto R=60, 88 with bigR boundary conditions.
#ψ0 is declared as a VECTOR bvp only knows how to handle vectors
ψ0= vcat(zeros(J+1), psi_eta, psi_eta_eta , psi_eta_eta_eta) #This works for the ODESolver AND BVPSolver
p= (Reynolds_number, J, Δθ, ψ0)
bvp_psi_2point = TwoPointBVProblem(steady_diffeq_psi_bvp!, bc_bvp!, ψ0, etaspan, p)
sol = @time solve(bvp_psi_2point, alg_hints = [:stiff], GeneralMIRK4(),dt=0.001) #Very accurate solver
#sol_shooting = @time solve(bvp_psi_2point, alg_hints=[:stiff], reltol=1e-6, abstol=1e-6 , Shooting(Rosenbrock23()))
# -
plot(sol_shooting)
# +
MatrixPsis=readdlm("psifin.m",Float64,);
size(MatrixPsis)
# -
xinterval=range(-1,stop=1,length=101)
plot(xinterval,MatrixPsis[2:102,77])
theta_step=30
mypsi = [i[theta_step] for i in sol.u]
etarange = [i for i in sol.t]
plot!(etarange, mypsi, linecolor=:orange)
# +
theta_step=20
mypsi = [i[theta_step] for i in sol.u]
etarange = [i for i in sol.t]
plot(etarange, mypsi, linecolor=:orange)
plot!(x->Ψ_bigr((theta_step-1)*Δθ,x),-1,1, label="Large R analytical", linestyle=:dash, linecolor=:red,xlabel = L"\eta", ylabel = L"\Psi",size=(800,400),yguidefontrotation=-90, legend=:outertopright)
plot!(x->Ψ_smallr((theta_step-1)*Δθ,x,Reynolds_number),-1,1, label="Small R analytical", linestyle=:dot, linecolor=:blue,xlabel = L"\eta", ylabel = L"\Psi",size=(800,400),yguidefontrotation=-90, legend=:outertopright)
title!(L"\Psi\ at\ \theta = "*string((theta_step-1))*L"\pi/"*string(J))
# -
#First derivative of psi wrt to eta at the theta=0 boundary
print(sol.u[1][2*J+3:3*J+3])
#First derivative of intitial psi wrt to eta at the theta=0 boundary
#print(ψ0[2*J+3:3*J+3])
#for i=2*J+3:3*J+3
for i=3*J+4:4*J+4
println(sol.u[1][i]," ", Ψ_bigr_d3(Δθ*(i-3*J-4), -1.0)," ", Ψ_smallr_d3(Δθ*(i-3*J-4), -1.0, Reynolds_number),)
end
# +
eta_vector = [i for i in sol.t]
eta_totalsteps = length(sol.u)
psi_sol = zeros((J+1)*eta_totalsteps)
counter = 1
for j = 1:J+1
for matrix in sol.u
psi_sol[counter] = matrix[j,1]
counter += 1
end
end
# -
#sol.u[end][1:J+1]
mypsi
for plot_counter =1:5
theta_step = plot_counter
if (theta_step > J+1)
println("The value of theta_step should be lower than "*string(J+2))
elseif (theta_step < 1)
println("The value of theta_step should be greater than 0")
end
psi_attheta = @view psi_sol[ (eta_totalsteps*(theta_step-1) +1) : eta_totalsteps*theta_step ]
plot(eta_vector,psi_attheta,label="Julia numerical solution", size=(800,600), legend=:outertopright)
temp_plot =plot!(x->Ψ_smallr((theta_step-1)*Δθ,x,Reynolds_number),-1,1, label="Small R analytical", xlabel = L"\eta", ylabel = L"\Psi",size=(800,400),yguidefontrotation=-90, legend=:outertopright)
title!(L"\Psi\ at\ \theta = "*string((theta_step-1))*L"\pi/"*string(J))
display(temp_plot)
sleep(1)
end
ψ0[:,1]
length(sol.t)
eta_n = length(sol.t)
eta_values= [i for i in sol.t]
theta_values=
X = x'.*ones(n)
y=10:15
Y = y'.*ones(n)
z=[i for i =1:n^2]
z= reshape(z,(n,n))
x = [j for j=1:n for i=1:n]
y = [i for j=1:n for i=1:n]
eta_values
pyplot()
plot(x,y,z,seriestype=:scatter, markersize = 7,camera=(-30,30))
?TwoPointBVProblem
| SteadyPsiEtaTheta_BVP_Upwind.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 8. Calibration of double ended measurement with WLS and confidence intervals
# ## Calibration procedure
# A double ended calibration is performed with weighted least squares. Over all timesteps simultaneous. $\gamma$ and $\alpha$ remain constant, while $C$ varies over time. The weights are not considered equal here. The weights kwadratically decrease with the signal strength of the measured Stokes and anti-Stokes signals.
#
# The confidence intervals can be calculated as the weights are correctly defined.
# The confidence intervals consist of two sources of uncertainty.
#
# 1. Measurement noise in the measured Stokes and anti-Stokes signals. Expressed in a single variance value.
# 2. Inherent to least squares procedures / overdetermined systems, the parameters are estimated with limited certainty and all parameters are correlated. Which is expressen in the covariance matrix.
#
# Both sources of uncertainty are propagated to an uncertainty in the estimated temperature via Monte Carlo.
# +
import os
from dtscalibration import read_silixa_files
import matplotlib.pyplot as plt
# %matplotlib inline
# +
filepath = os.path.join('..', '..', 'tests', 'data', 'double_ended2')
ds_ = read_silixa_files(
directory=filepath,
timezone_netcdf='UTC',
file_ext='*.xml')
ds = ds_.sel(x=slice(0, 100)) # only calibrate parts of the fiber
sections = {
'probe1Temperature': [slice(7.5, 17.), slice(70., 80.)], # cold bath
'probe2Temperature': [slice(24., 34.), slice(85., 95.)], # warm bath
}
ds.sections = sections
# -
st_label = 'ST'
ast_label = 'AST'
rst_label = 'REV-ST'
rast_label = 'REV-AST'
# First calculate the variance in the measured Stokes and anti-Stokes signals, in the forward and backward direction.
#
# The Stokes and anti-Stokes signals should follow a smooth decaying exponential. This function fits a decaying exponential to each reference section for each time step. The variance of the residuals between the measured Stokes and anti-Stokes signals and the fitted signals is used as an estimate of the variance in measured signals.
st_var, resid = ds.variance_stokes(st_label=st_label)
ast_var, _ = ds.variance_stokes(st_label=ast_label)
rst_var, _ = ds.variance_stokes(st_label=rst_label)
rast_var, _ = ds.variance_stokes(st_label=rast_label)
resid.plot(figsize=(12, 8));
# We calibrate the measurement with a single method call. The labels refer to the keys in the DataStore object containing the Stokes, anti-Stokes, reverse Stokes and reverse anti-Stokes. The variance in those measurements were calculated in the previous step. We use a sparse solver because it saves us memory.
ds.calibration_double_ended(
st_label=st_label,
ast_label=ast_label,
rst_label=rst_label,
rast_label=rast_label,
st_var=st_var,
ast_var=ast_var,
rst_var=rst_var,
rast_var=rast_var,
store_tmpw='TMPW',
method='wls',
solver='sparse')
ds.TMPW.plot()
# ## Confidence intervals
# With another method call we estimate the confidence intervals. If the method is `wls` and confidence intervals are passed to `conf_ints`, confidence intervals calculated. As weigths are correctly passed to the least squares procedure, the covariance matrix can be used as an estimator for the uncertainty in the parameters. This matrix holds the covariances between all the parameters. A large parameter set is generated from this matrix as part of the Monte Carlo routine, assuming the parameter space is normally distributed with their mean at the best estimate of the least squares procedure.
#
# The large parameter set is used to calculate a large set of temperatures. By using `percentiles` or `quantile` the 95% confidence interval of the calibrated temperature between 2.5% and 97.5% are calculated.
#
# The confidence intervals differ per time step. If you would like to calculate confidence intervals of all time steps together you have the option `ci_avg_time_flag=True`. 'We can say with 95% confidence that the temperature remained between this line and this line during the entire measurement period'. This is ideal if you'd like to calculate the background temperature with a confidence interval.
ds.conf_int_double_ended(
p_val='p_val',
p_cov='p_cov',
st_label=st_label,
ast_label=ast_label,
rst_label=rst_label,
rast_label=rast_label,
st_var=st_var,
ast_var=ast_var,
rst_var=rst_var,
rast_var=rast_var,
store_tmpf='TMPF',
store_tmpb='TMPB',
store_tmpw='TMPW',
store_tempvar='_var',
conf_ints=[2.5, 50., 97.5],
mc_sample_size=500, # <- choose a much larger sample size
ci_avg_time_flag=False)
ds1 = ds.isel(time=-1) # take only the first timestep
ds1.TMPW.plot(linewidth=0.7, figsize=(12, 8))
ds1.TMPW_MC.isel(CI=0).plot(linewidth=0.7, label='CI: 2.5%')
ds1.TMPW_MC.isel(CI=2).plot(linewidth=0.7, label='CI: 97.5%')
plt.legend();
# The DataArrays `TMPF_MC` and `TMPB_MC` and the dimension `CI` are added. `MC` stands for monte carlo and the `CI` dimension holds the confidence interval 'coordinates'.
(ds1.TMPW_MC_var**0.5).plot(figsize=(12, 4));
plt.ylabel('$\sigma$ ($^\circ$C)');
ds.data_vars
| examples/notebooks/08Calibrate_double_wls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression with news (all)
# +
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import sys
sys.version
import pandas as pd
import os
import copy
import numpy as np
import xgboost
import pickle
from pythainlp.corpus import stopwords
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, mean_absolute_error, accuracy_score
from sklearn import preprocessing
from sklearn.linear_model import Ridge
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
from plotly import tools
import plotly.graph_objs as go
from datetime import datetime, timedelta
from tqdm import tqdm_notebook
import matplotlib.pyplot as plt
% matplotlib inline
# +
target_stocks = ['BANPU','IRPC','PTT','BBL','KBANK','SCB','AOT','THAI','CPF','MINT',
'TU','SCC','CPN','CK','CPALL','HMPRO','BDMS','BH','ADVANC','JAS','TRUE']
df_price = pd.read_csv('merged_2013_2018.csv')
df_price['Date'] = pd.to_datetime(df_price['Date'], format='%Y-%m-%d')
df_price = df_price.loc[df_price['Ticker'].isin(target_stocks)]
df_price['Date'] = df_price['Date'].dt.date
df_price = df_price.set_index('Date')
df_price.tail(3)
len(df_price)
df_kaohoon = pd.read_csv('data/kaohoon_all.csv')
df_kaohoon['Date'] = pd.to_datetime(df_kaohoon['Date'], format='%Y-%m-%d')
df_kaohoon = df_kaohoon.set_index('Date')
df_kaohoon = df_kaohoon[:'2018-2-8']
df_kaohoon.index = df_kaohoon.index.date
df_kaohoon.tail(3)
len(df_kaohoon)
df_moneych = pd.read_csv('data/moneychanel_all.csv')
df_moneych['Date'] = pd.to_datetime(df_moneych['Date'], format='%Y-%m-%d')
df_moneych = df_moneych.set_index('Date')
df_moneych = df_moneych[:'2018-2-8']
df_moneych.index = df_moneych.index.date
df_moneych.tail(3)
len(df_moneych)
df_news = pd.concat([df_moneych, df_kaohoon])
df_news = df_news.sort_index()
# -
# # Lag & Horizon Construction
# +
N_lags = 3
N_horizon = 1
df_train = []
df_test = []
for stock in tqdm_notebook(target_stocks):
news_stocks = []
df_stock = df_news.loc[df_news['Ticker'] == stock]
prev_date = None
prev_text = None
# pbar = tqdm_notebook(total=len(df_stock))
for date, row in df_stock.iterrows():
if prev_date == None:
prev_date = date
prev_text = row['Text']
elif prev_date != date:
# horizon
tmp_date = copy.deepcopy(prev_date)
tmp_date += timedelta(days=1)
prices = []
count_lags = 0
while count_lags < N_horizon:
price = df_price.loc[(df_price.index == tmp_date) & (df_price['Ticker'] == stock)].values
tmp_date += timedelta(days=1)
if len(price) == 0: continue
prices.append(price[0][4]) # Close price next day(s)
count_lags+=1
# lag
tmp_date = copy.deepcopy(prev_date)
count_lags = 0
while count_lags <= N_lags:
price = df_price.loc[(df_price.index == tmp_date) & (df_price['Ticker'] == stock)].values
tmp_date -= timedelta(days=1)
if len(price) == 0: continue
for val in price[0][:-1]:
if type(val) != str: prices.append(val)
count_lags+=1
news_stocks.append([prev_date, stock, prev_text] + prices)
prev_date = date
prev_text = row['Text']
elif prev_date == date:
prev_text += ' '+row['Text']
# pbar.update(1)
# pbar.close()
news_stocks = pd.DataFrame.from_records(news_stocks)
news_stocks.columns = ['Date', 'Ticker', 'Text',
'Close(t+1)', # 'Close(t+2)','Close(t+3)','Close(t+4)','Close(t+5)',
'Open(t)', 'High(t)', 'Low(t)', 'Close(t)',
'Open(t-1)', 'High(t-1)', 'Low(t-1)', 'Close(t-1)',
'Open(t-2)', 'High(t-2)', 'Low(t-2)', 'Close(t-2)',
'Open(t-3)', 'High(t-3)', 'Low(t-3)', 'Close(t-3)',
# 'Open(t-4)', 'High(t-4)', 'Low(t-4)', 'Close(t-4)',
# 'Open(t-5)', 'High(t-5)', 'Low(t-5)', 'Close(t-5)'
]
news_stocks = news_stocks.set_index('Date')
train_size = int(len(news_stocks) * 0.95)
test_size = len(news_stocks) - train_size
train, test = news_stocks.iloc[:train_size], news_stocks.iloc[train_size:]
print(stock, ':\t',len(train), len(test))
df_train.append(train)
df_test.append(test)
df_train = pd.concat(df_train, axis=0)
df_test = pd.concat(df_test, axis=0)
# +
df_train.to_csv('data/df_train_news_all.csv')
df_test.to_csv('data/df_test_news_all.csv')
df_train = pd.read_csv('data/df_train_news_all.csv')
df_test = pd.read_csv('data/df_test_news_all.csv')
df_train = df_train.set_index('Date')
df_test = df_test.set_index('Date')
len(df_train), len(df_test)
df_train.head(2)
df_test.head(2)
# -
# # TF-IDF Vetorization
# +
stop_words = stopwords.words('thai')
stop_words.remove('ขึ้น')
stop_words.remove('ลง')
vertorizer = TfidfVectorizer(stop_words=stop_words,
max_df=0.9,
min_df=2,
max_features=1000)
tfidf_train = vertorizer.fit_transform(df_train['Text'])
tfidf_test = vertorizer.transform(df_test['Text'])
df_tfidf_train = pd.DataFrame.from_records(tfidf_train.toarray())
df_tfidf_test = pd.DataFrame.from_records(tfidf_test.toarray())
df_tfidf_train = df_tfidf_train.set_index(df_train.index)
df_tfidf_test = df_tfidf_test.set_index(df_test.index)
len(df_tfidf_train), len(df_tfidf_test)
# replace Text with TF-IDF vector
x_train = df_train.drop(['Text'], axis=1)
x_train = pd.concat([x_train, df_tfidf_train], axis=1)
x_test = df_test.drop(['Text'], axis=1)
x_test = pd.concat([x_test, df_tfidf_test], axis=1)
# Label Encoding
le = preprocessing.LabelEncoder()
x_train['Ticker'] = le.fit_transform(x_train['Ticker'])
x_test['Ticker'] = le.transform(x_test['Ticker'])
x_train.head(2)
x_test.head(2)
le.classes_
# -
# # Create x_train and y_train
Horizon = 'Close(t+1)'
y_train = x_train[[Horizon]]
x_train = x_train.drop(['Close(t+1)'], axis=1).copy()
x_train.shape, y_train.shape
# # Evaluate Each Stcok
# +
def mean_absolute_percentage_error(y_true, y_pred):
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
def evaluator(clf, df_test, le, isXGB=False, isLSTM=False):
RMSEs, MAEs, MAPEs, DAs = [], [], [], []
results = []
for stock in target_stocks:
x_tmp = df_test.loc[df_test['Ticker'] == le.transform([stock])[0]].copy()
y_tmp = x_tmp[Horizon].values
# Directional Accuracy
changes = x_tmp[Horizon] - x_tmp['Close(t)']
y_true_da = []
for change in changes:
y_true_da.append(1 if change >= 0 else 0)
x_tmp = x_tmp.drop(['Close(t+1)'], axis=1)
if isXGB:
y_pred = clf.predict(xgboost.DMatrix(x_tmp))
elif isLSTM:
x = x_tmp.values
x = x.reshape((x.shape[0], x.shape[1], 1))
y_pred = clf.predict(x)
else:
y_pred = clf.predict(x_tmp.as_matrix())
# Directional Accuracy Pred
changes = np.reshape(y_pred, (-1,1)) - np.reshape(x_tmp['Close(t)'].values,(-1,1))
y_pred_da = []
for change in changes:
y_pred_da.append(1 if change >= 0 else 0)
RMSE = np.sqrt(mean_squared_error(y_tmp, y_pred))
MAE = mean_absolute_error(y_tmp, y_pred)
MAPE = mean_absolute_percentage_error(y_tmp, y_pred)
DA = accuracy_score(y_true_da, y_pred_da)
print(stock, "\tRMSE: %.2f\t MAE: %.2f \tMAPE: %.2f \tDA: %.2f" % (RMSE, MAE, MAPE, DA))
RMSEs.append(RMSE)
MAEs.append(MAE)
MAPEs.append(MAPE)
DAs.append(DA)
print('\nmean RMSE:', round(np.mean(RMSEs),2))
print('mean MAE:', round(np.mean(MAEs),2))
print('mean MAPE:', round(np.mean(MAPEs),2))
print('mean DA:', round(np.mean(DAs),4))
# -
def ensemble_evaluator(bagging, ada_dt, ada_rf, xgb, stack, stack_da, df_test, le, feature_importances, feature_importances_da):
RMSEs, MAEs, MAPEs, DAs = [], [], [], []
results = []
for stock in target_stocks:
x_tmp = df_test.loc[df_test['Ticker'] == le.transform([stock])[0]].copy()
# Directional Accuracy
changes = x_tmp[Horizon] - x_tmp['Close(t)']
y_true_da = []
for change in changes:
y_true_da.append(1 if change >= 0 else 0)
y_tmp = x_tmp[Horizon].values.reshape(-1,1)
x_tmp = x_tmp.drop([Horizon], axis=1)
# Prediction
y_pred = np.concatenate((
bagging.predict(x_tmp).reshape(-1,1),
ada_dt.predict(x_tmp).reshape(-1,1),
ada_rf.predict(x_tmp).reshape(-1,1),
xgb.predict(xgboost.DMatrix(x_tmp)).reshape(-1,1)),
axis=1)
df_pred = pd.DataFrame.from_records(y_pred).round(2)
df_pred.columns = ['Bagging_DT', 'Ada_DT', 'Ada_RF', 'XGB']
df_pred.head()
# Directional Accuracy Pred
close_t = np.reshape(x_tmp['Close(t)'].values, (-1, 1))
y_changes = np.concatenate((
np.array(y_pred[:,0]).reshape(-1,1)-close_t,
np.array(y_pred[:,1]).reshape(-1,1)-close_t,
np.array(y_pred[:,2]).reshape(-1,1)-close_t,
np.array(y_pred[:,3]).reshape(-1,1)-close_t,
), axis=1)
y_pred_da = []
for row in y_changes:
tmp_row = []
for change in row:
tmp_row.append(1 if change>=0 else 0)
y_pred_da.append(tmp_row)
df_pred_da = pd.DataFrame.from_records(y_pred_da)
df_pred_da.columns = ['Bagging_DT', 'Ada_DT', 'Ada_RF', 'XGB']
df_pred_da.head()
df_pred['weight'] = (
df_pred['Bagging_DT']*feature_importances[0] +
df_pred['Ada_DT']*feature_importances[1] +
df_pred['Ada_RF']*feature_importances[2] +
df_pred['XGB']*feature_importances[3]
)
df_pred_da['vote'] = (
df_pred_da['Bagging_DT']*feature_importances_da[0] +
df_pred_da['Ada_DT']*feature_importances_da[1] +
df_pred_da['Ada_RF']*feature_importances_da[2] +
df_pred_da['XGB']*feature_importances_da[3]
).round(0).astype(int)
y_pred = stack.predict(y_pred).reshape(-1,1)
y_pred_da = stack_da.predict(y_pred_da).reshape(-1,1).round(0).astype(int)
# y_pred = df_pred['weight'].values.reshape(-1,1)
# y_pred_da = df_pred_da['vote'].values.reshape(-1,1)
RMSE = np.sqrt(mean_squared_error(y_tmp, y_pred))
MAE = mean_absolute_error(y_tmp, y_pred)
MAPE = mean_absolute_percentage_error(y_tmp, y_pred)
DA = accuracy_score(y_true_da, y_pred_da)
print(stock, "\tRMSE: %.2f\t MAE: %.2f \tMAPE: %.2f \tDA: %.2f" % (RMSE, MAE, MAPE, DA))
RMSEs.append(RMSE)
MAEs.append(MAE)
MAPEs.append(MAPE)
DAs.append(DA)
print('\nmean RMSE:', round(np.mean(RMSEs),2))
print('mean MAE:', round(np.mean(MAEs),2))
print('mean MAPE:', round(np.mean(MAPEs),2))
print('mean DA:', round(np.mean(DAs),4))
# # Ensemble
ensemble_evaluator(
bagging,
adaboost_dt_regr,
adaboost_rf_regr,
xgb,
stack, stack_da,
x_test, le, feature_importances, feature_importances_da)
# # Linear Regression
# +
from sklearn import linear_model
lineregr = linear_model.LinearRegression()
lineregr.fit(x_train, y_train)
evaluator(lineregr, x_test, le)
# -
# # Support Vector Regressor
# +
# from sklearn.svm import SVR
# svr = SVR()
# svr.fit(x_train, y_train)
# evaluator(svr, x_test, le)
# -
# # Decistion Tree Regressor
# +
from sklearn import tree
decis_tree_regr = tree.DecisionTreeRegressor(max_depth=10)
decis_tree_regr.fit(x_train, y_train.values.ravel())
evaluator(decis_tree_regr, x_test, le)
# -
# # Random Forest Regrssor
# +
from sklearn import ensemble
rnd_forest_regr = ensemble.RandomForestRegressor(n_jobs=-1)
rnd_forest_regr.fit(x_train, y_train.values.ravel())
evaluator(rnd_forest_regr, x_test, le)
# -
# # Bagging Regressor
bagging = ensemble.BaggingRegressor(base_estimator=None,
n_estimators=30,n_jobs=-1)
bagging.fit(x_train, y_train.values.ravel())
evaluator(bagging, x_test, le)
# # AdaBoost Regressor
# +
adaboost_dt_regr = ensemble.AdaBoostRegressor(base_estimator=tree.DecisionTreeRegressor(),
learning_rate=1,
n_estimators=10,
loss='linear')
adaboost_dt_regr.fit(x_train, y_train.values.ravel())
evaluator(adaboost_dt_regr, x_test, le)
# +
adaboost_rf_regr = ensemble.AdaBoostRegressor(base_estimator=ensemble.RandomForestRegressor(n_jobs=-1),
learning_rate=1,
n_estimators=10,
loss='linear')
adaboost_rf_regr.fit(x_train, y_train.values.ravel())
evaluator(adaboost_rf_regr, x_test, le)
# -
# # Gradient Boosting Regressor
# +
gbr = ensemble.GradientBoostingRegressor(n_estimators=100,
learning_rate=0.1,
max_depth=6,
min_samples_split=2,
loss='ls',
)
gbr.fit(x_train, y_train.values.ravel())
evaluator(gbr, x_test, le)
# -
# # XGBoost Regressor
# +
import xgboost
from sklearn.model_selection import train_test_split
d_train, d_valid, y_d_train, y_d_valid = train_test_split(x_train, y_train, test_size=0.1, random_state=10)
len(d_train), len(d_valid)
d_train = xgboost.DMatrix(d_train, label=y_d_train)
d_valid = xgboost.DMatrix(d_valid, label=y_d_valid)
# -
# Parameters
# http://xgboost.readthedocs.io/en/latest//parameter.html
# +
params = {
'booster':'dart',
'max_depth': 4,
'learning_rate': 0.05,
'subsample': 1,
'objective': 'reg:tweedie',
'eval_metric': 'mae',
'reg_lambda': 0.8,
'reg_alpha': 0.2,
'silent': 1,
'sample_type':"weighted"
}
xgb = xgboost.train(params, d_train,
num_boost_round=5000,
evals=[(d_train, 'train'), (d_valid, 'valid')],
early_stopping_rounds=50,
verbose_eval=100
)
# -
evaluator(xgb, x_test, le, isXGB=True)
# # Save ML models
pickle.dump(decis_tree_regr, open('models/decis_tree_regr_news_all.pkl', 'wb'))
pickle.dump(rnd_forest_regr, open('models/rnd_forest_regr_news_all.pkl', 'wb'))
pickle.dump(bagging, open('models/bagging_regr_news_all.pkl', 'wb'))
pickle.dump(adaboost_dt_regr, open('models/adaboost_dt_regr_news_all.pkl', 'wb'))
pickle.dump(adaboost_rf_regr, open('models/adaboost_rf_regr_news_all.pkl', 'wb'))
pickle.dump(xgb, open('models/xgb_news_all.pkl', 'wb'))
# # Ensemble Stacking
dt = pickle.load(open('models/decis_tree_regr_news_all.pkl', 'rb'))
rf = pickle.load(open('models/rnd_forest_regr_news_all.pkl', 'rb'))
bagging = pickle.load(open('models/bagging_regr_news_all.pkl', 'rb'))
ada_dt = pickle.load(open('models/adaboost_dt_regr_news_all.pkl', 'rb'))
ada_rf = pickle.load(open('models/adaboost_rf_regr_news_all.pkl', 'rb'))
xgb = pickle.load(open('models/xgb_news_all.pkl', 'rb'))
x_train_stack = np.concatenate((
bagging.predict(x_train).reshape(-1,1),
ada_dt.predict(x_train).reshape(-1,1),
ada_rf.predict(x_train).reshape(-1,1),
xgb.predict(xgboost.DMatrix(x_train)).reshape(-1,1)), axis=1)
# +
y_test = x_test[Horizon].values.reshape(-1,1)
x_test_stack = np.concatenate((
bagging.predict(x_test.drop(['Close(t+1)'], axis=1)).reshape(-1,1),
ada_dt.predict(x_test.drop(['Close(t+1)'], axis=1)).reshape(-1,1),
ada_rf.predict(x_test.drop(['Close(t+1)'], axis=1)).reshape(-1,1),
xgb.predict(xgboost.DMatrix((x_test.drop(['Close(t+1)'], axis=1)))).reshape(-1,1)), axis=1)
# +
stack = ensemble.RandomForestRegressor(n_jobs=-1)
stack.fit(x_train_stack, y_train.values.ravel())
y_pred_stack = stack.predict(x_test_stack).reshape(-1,1)
for i in range(x_test_stack.shape[1]):
RMSE = np.sqrt(mean_squared_error(y_test, x_test_stack[:,i]))
MAE = mean_absolute_error(y_test, x_test_stack[:,i])
MAPE = mean_absolute_percentage_error(y_test, x_test_stack[:,i].reshape(-1,1))
print("RMSE: %.2f \tMAE: %.2f \tMAPE: %.2f" % (RMSE, MAE, MAPE))
RMSE = np.sqrt(mean_squared_error(y_test, y_pred_stack))
MAE = mean_absolute_error(y_test, y_pred_stack)
MAPE = mean_absolute_percentage_error(y_test, y_pred_stack)
print("\nRMSE: %.2f \tMAE: %.2f \tMAPE: %.2f" % (RMSE, MAE, MAPE))
# -
feature_importances = stack.feature_importances_
feature_importances
# # Stack DA
# +
close_t = np.reshape(x_train['Close(t)'].values, (-1, 1))
changes = y_train.values.reshape(-1,1) - close_t
y_train_da = np.array([1 if change >= 0 else 0 for change in changes]).reshape(-1,1)
x_changes = np.concatenate((
np.array(x_train_stack[:,0]).reshape(-1,1)-close_t,
np.array(x_train_stack[:,1]).reshape(-1,1)-close_t,
np.array(x_train_stack[:,2]).reshape(-1,1)-close_t,
np.array(x_train_stack[:,3]).reshape(-1,1)-close_t,
), axis=1)
x_train_stack_da = []
for row in x_changes:
tmp_row = []
for change in row:
tmp_row.append(1 if change>=0 else 0)
x_train_stack_da.append(tmp_row)
# +
close_t = np.reshape(x_test['Close(t)'].values, (-1, 1))
changes = y_test - close_t
y_test_da = np.array([1 if change >= 0 else 0 for change in changes]).reshape(-1,1)
x_changes = np.concatenate((
np.array(x_test_stack[:,0]).reshape(-1,1)-close_t,
np.array(x_test_stack[:,1]).reshape(-1,1)-close_t,
np.array(x_test_stack[:,2]).reshape(-1,1)-close_t,
np.array(x_test_stack[:,3]).reshape(-1,1)-close_t,
), axis=1)
x_test_stack_da = []
for row in x_changes:
tmp_row = []
for change in row:
tmp_row.append(1 if change>=0 else 0)
x_test_stack_da.append(tmp_row)
x_test_stack_da = np.array(x_test_stack_da)
# +
stack_da = ensemble.GradientBoostingRegressor()
stack_da.fit(x_train_stack_da, y_train_da.ravel())
y_stack_da = stack_da.predict(x_test_stack_da).reshape(-1,1).round(0).astype(int)
for i in range(x_test_stack_da.shape[1]):
acc = accuracy_score(y_test_da, x_test_stack_da[:,i])
print("Accuracy: %.4f" % (acc))
acc = accuracy_score(y_test_da, y_stack_da)
print("\nAccuracy: %.4f" % (acc))
# -
feature_importances_da = stack_da.feature_importances_
feature_importances_da
# # Visualize Error
def visualize(ticker):
x_tmp = x_test.loc[x_test['Ticker'] == le.transform([ticker])[0]].copy()
y_tmp = x_tmp[Horizon].values.reshape(-1,1)
x_tmp = x_tmp.drop([Horizon], axis=1)
y_pred = xgb.predict(xgboost.DMatrix((x_tmp))).reshape(-1,1)
df = np.concatenate((y_tmp,
y_pred), axis=1)
df = pd.DataFrame.from_records(df)
df.columns = ['actual', 'predicted']
df['AE'] = 100*(df['actual']-df['predicted']).abs()/df['actual']
trace_1 = go.Scatter(
x = x_tmp.index,
y = df.actual,
mode = 'lines',
name = 'Actual',
line = dict(width = 4)
)
trace_2 = go.Scatter(
x = x_tmp.index,
y = df.predicted,
mode = 'lines',
name = 'Predicted',
line = dict(width = 4)
)
ln = make_pipeline(PolynomialFeatures(4), Ridge())
ln = LinearRegression()
ln.fit(df.index.values.reshape(-1,1), df.AE)
trace_3 = go.Scatter(
x = x_tmp.index,
y = ln.predict(df.index.values.reshape(-1,1)),
mode = 'lines',
name = 'Trend Line',
line = dict(width = 4)
)
trace_4 = go.Scatter(
x = x_tmp.index,
y = df.AE,
mode = 'lines',
name = 'Errors',
line = dict(width = 4)
)
fig = tools.make_subplots(rows=2, cols=1, subplot_titles=(ticker+': Actual vs. Predicted', 'Absolute Error'))
fig.append_trace(trace_1, 1, 1)
fig.append_trace(trace_2, 1, 1)
fig.append_trace(trace_3, 2, 1)
fig.append_trace(trace_4, 2, 1)
fig['layout'].update(height=1000, width=1200,
paper_bgcolor='rgba(0,0,0,0)',
plot_bgcolor='rgba(0,0,0,0)',
font=dict(size=16, color='#bfbfbf'))
fig['layout']['yaxis1'].update(title='Price')
fig['layout']['yaxis2'].update(title='Absolute Error')
iplot(fig, filename='line-mode')
visualize('SCB')
# # LSTM
from keras.models import Sequential
from keras.layers import Dense, Dropout, LSTM
# +
train_X = x_train.values
val_X = x_valid.values
# test_X = x_test.values
train_y = y_train.values
val_y = y_valid.values
# test_y = y_test.values
train_X = train_X.reshape(train_X.shape[0], train_X.shape[1], 1)
val_X = val_X.reshape(val_X.shape[0], val_X.shape[1], 1)
# test_X = test_X.reshape((test_X.shape[0], 1, test_X.shape[1]))
print(train_X.shape, train_y.shape)
print(val_X.shape, val_y.shape)
# print(test_X.shape, test_y.shape)
# -
model = Sequential()
model.add(LSTM(64, return_sequences=True, input_shape=(train_X.shape[1], 1)))
model.add(Dropout(0.3))
model.add(LSTM(128, return_sequences=True))
model.add(Dropout(0.3))
# model.add(LSTM(256, return_sequences=True))
# model.add(Dropout(0.3))
# model.add(LSTM(256, return_sequences=True))
# model.add(Dropout(0.4))
model.add(LSTM(128))
model.add(Dense(1))
model.compile(loss='mean_absolute_error', optimizer='adam')
# +
from keras.callbacks import ModelCheckpoint, TensorBoard, EarlyStopping
checkpoint = ModelCheckpoint(filepath="model/LSTM.h5",
monitor='val_loss',
verbose=1,
save_best_only=True,
save_weights_only=False,
mode='auto',
period=1
)
earlystopping = EarlyStopping(monitor='val_loss',
min_delta=0,
patience=100,
verbose=1,
mode='auto')
# access via $ tensorboard --logdir=./logs
tensorboard = TensorBoard(log_dir='./logs')
# -
model.fit(x=train_X,
y=train_y,
epochs=1000,
batch_size=32,
validation_data=(val_X, val_y),
verbose=1,
shuffle=False,
callbacks=[checkpoint, earlystopping, tensorboard]
)
evaluator(model, x_test, le, isLSTM=True)
from keras.layers import Bidirectional
model = Sequential()
model.add(Bidirectional(LSTM(128, return_sequences=True), input_shape=(2, train_X.shape[2])))
model.add(Dropout(0.2))
model.add(Bidirectional(LSTM(128, return_sequences=True)))
model.add(Dropout(0.3))
model.add(Bidirectional(LSTM(128, return_sequences=True)))
model.add(Dropout(0.4))
model.add(Bidirectional(LSTM(128)))
model.add(Dense(1))
model.compile(loss='mean_absolute_error', optimizer='adam')
model.fit(x=train_X,
y=train_y,
epochs=10000,
batch_size=1024,
validation_data=(val_X, val_y),
verbose=1,
shuffle=False,
callbacks=[checkpoint, earlystopping, tensorboard]
)
'hey'
evaluator(model, x_test, le, isLSTM=True)
data_dim = 16
timesteps = 8
num_classes = 10
np.random.random((10, timesteps, data_dim)).shape
# +
train_X = x_train.values
val_X = x_valid.values
train_X.shape
train_X = train_X.reshape(train_X.shape[0], train_X.shape[1], 1)
train_X.shape
# -
| 1_regression_news_all.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting Started - Further Details
#
# This notebook will introduce all of the other parameters currently in the iDEA code. Note that the explanations are only very brief, so if a proper and full understanding is to be gained, study the relevent theory notebooks. There are elementary exercises at the end of each section to test understanding, with model answers at the end of the notebook. *They are considered elementary as they should only involve changing the values of relevent parameters and plotting the result*
#
# +
# Setting up - same as previous notebook
from iDEA.input import Input
pm = Input()
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import animation, rc
from IPython.display import HTML
from iDEA.input import Input
from iDEA.results import Results as rs
# -
#
# ## Further Parameters
#
# ### Exact Many-body Calculation
#
# The exact many-body calculation is done by propagating the wavefunction in imaginary time, which eventually converges on the ground-state energy. The exact (ext) parameters are as follows:
#
# * **ext.itol** - Tolerance of imaginary time propagation (~ 1e-12)
# * **ext.itol_solver** - Tolerance of linear solver in imaginary time propagation (~ 1e-14)
# * **ext.rtol_solver** - Tolerance of linear solver in real time propagation (~ 1e-12)
# * **ext.itmax** - Total imaginary time
# * **ext.iimax** - Imaginary time iterations
# * **ext.ideltat** - Imaginary time step (DERIVED)
# * **ext.RE** - Reverse engineer many-body density
# * **ext.OPT** - Calculate the external potential for the exact density
# * **ext.excited_states** - Number of excited states to calculate (0 for only ground-state)
# * **ext.psi_gs** - Save the reduced ground-state wavefunction to file
# * **ext.psi_es** - Save the reduced excited-state wavefunctions to file
# * **ext.initial_psi** - initial wavefunction: qho, non, hf, lda1/2/3/heg, ext or wavefunction from previous run (e.g run_name)
#
# Tolerance describes the difference between, say for example, densities between iterations before a successful convergence has occured.
#
# Note that in the exercises, if a variable is not specified, such as electron number, take it to be the default value set in the parameters file.
# #### Exercise 1
#
# Perfom exact many-body calculations for the ground state first excited state and calculate the relevent ELF. Display these results in a graph.
# *HINT: start by printing the parameters to find out what needs to be changed to get the desired result.*
# ### Non-interaction approximation
#
# This is the simplest form of DFT, which acts as a baseline comparison to see whether an approximation, like the LDA, gives a good result compared to the non interacting result. The parameters are as follows:
#
# * **non.rtol_solver** - Tolerance of linear solver in real time propagation (~e-13)
# * **non.save_eig** - Save eigenfunctions and eigenvalues of Hamiltonian
# * **non.RE** - Reverse-engineer non-interacting density
# * **non.OPT** - Calculate the external potential for the non-interacting density
#
#
# ### Local Density Approximation
#
# This is the most common approximation used in DFT. The parameters are as follows:
#
# * **lda.NE** - number of electrons used for LDA construction (1, 2, 3, 'heg')
# * **lda.scf_type** - scf type (linear, pulay, cg)
# * **lda.mix** - mixing parameter for linear and Pulay mixing (between 0 and 1)
# * **lda.pulay_order** - history length for Pulay mixing (max: lda.max_iter)
# * **lda.pulay_preconditioner** - preconditioner for Pulay mixing (None, kerker, rpa)
# * **lda.kerker_length** - length over which density flunctuations are screened (Kerker)
# * **lda.tol** - convergence tolerance in the density
# * **lda.etol** - convergence tolerance in the energy
# * **lda.max_iter** - maximum number of self-consistency iterations
# * **lda.save_eig** - save eigenfunctions and eigenvalues of Hamiltonian
# * **lda.OPT** - calculate the external potential for the LDA density
#
# For the number of electrons, 'heg' is an acronym for 'homogeneous electron gas'. There are also the types of self-consistency available: 'linear', 'pulay' and 'cg'. 'Linear' is the least complicated and used in most situations. Density fluctuations can occur, which prevents LDA from reaching self-consistency. These different methods and mixing of methods will help reach self-consistency.
#
# #### Exercise 2
#
# Perform DFT calculations for 2 electrons with the LDA approximations using 'heg' and compare with the non-interaction approximation and exact calculation by plotting the electron densities. *HINT: take a look at the parameters file directly if one cannot determine which parameter needs to be changed from the notebook's description of the parameters.*
# ### Hartree-Fock calculation
#
# The Hartree-Fock method is an alternative to DFT and is essentially a simplified version of many-body perturbation theory. The parameters are as follows:
#
# * **hf.fock** - include Fock term ( 0 = Hartree approximation, 1 = Hartree-Fock approximation)
# * **hf.con** - tolerance
# * **hf.nu** - mixing term
# * **hf.save_eig** - save eigenfunctions and eigenvalues of the Hamiltonian
# * **hf.RE** - reverse engineering HF density
# * **hf.OPT** - calculate the external potential from the HF density
#
#
# #### Exercise 3
#
# Compare the electron densities for a two-electron system with the exact, non-interacting and HF. *HINT: each cycle will produce an output file. If ever in doubt of what your outputs are, take a direct look in **outputs/run_name/raw**. (run_name being the default name)*
# ### Reverse Engineering
#
# The Reverse Engineering algorithm (for both time-independent and time-dependent systems) take the exact electron density and 'reverse engineers' the exact Kohn-Sham potential for the system. The parameters are as follows:
#
# * **re.save_eig** - save Kohn-Sham eigenfunctions and eigen values of reverse-engineered potential
# * **re.stencil** - discretisation of 1st derivative (5 or 7)
# * **re.mu** - 1st convergence parameter in the ground-state reverse-engineering algorithm
# * **re.p** - 2nd convergence parameter in the GS RE algorithm
# * **re.nu** - convergence parameter in the time-dependent RE algorithm
# * **re.rtol_solver** - tolerance of the linear solver in real-time propagation (~1e-12)
# * **re.density_tolerance** - tolerance of the error in the time_dependent density
# * **re.cdensity_tolerance** - tolerance of the error in the current density
# * **re.max_iterations** - maximum number of iterations per time step to find the Kohn-Sham potential
# * **re.damping** - damping factor used when filtering out noise in the Kohn-Sham vector potential (0: none)
#
# #### Exercise 4
#
# Reverse engineer the Kohn-Sham potential from the exact density from one of the previous exercises. The exact densities are, in fact, the same for all the exercises so the result should be the same regardless. *HINT: make sure to set the reverse-engineering parameter to True.*
# ## Model Answers
# +
#Exercise 1
# As always set a run_name
pm.run.name = 'exercise_1'
# Turn on the correct parameters
pm.run.NON = False
pm.run.EXT = True
pm.ext.elf_gs = True
pm.ext.elf_es = True
pm.ext.excited_states = 1
# Start the simulation
results = pm.execute()
# Plotting the results
ex1_results = rs.read("es_ext_elf1",pm)
plt.plot(ex1_results, label="ELF - 1st excited state")
plt.legend()
plt.show()
# +
#Exercise 2
# Setting up the parameters
# run
pm.run.name = 'exercise_2'
pm.run.NON = True
pm.run.LDA = True
pm.run.EXT = True
# sys
pm.sys.NE = 2
# lda
pm.lda.NE = 'heg'
# Starting the simulation
results = pm.execute()
# Plotting the results
ex2_lda = rs.read("gs_ldaheg_den",pm)
ex2_ext = rs.read("gs_ext_den",pm)
ex2_non = rs.read("gs_non_den",pm)
plt.plot(ex2_lda, label="lda - heg")
plt.plot(ex2_ext, label="ext")
plt.plot(ex2_non, label="non")
plt.legend()
plt.show()
# +
# Exercise 3
# Setting up the parameters
# run
pm.run.name = 'exercise_3'
pm.run.NON = True
pm.run.EXT = True
pm.run.HF = True
#results = pm.execute()
# Plotting the results
ex3_ext = rs.read("gs_ext_den",pm)
ex3_non = rs.read("gs_non_den",pm)
ex3_hf = rs.read("gs_hf_den",pm)
plt.plot(ex3_ext, label="ext")
plt.plot(ex3_non, label="non")
plt.plot(ex3_hf, label="hf")
plt.legend()
plt.show()
# +
# Exercise 4
# Setting up the parameters
# Reverse-engineering the exact density from exercise 2, so need to be
# working out of the subdirectory containing that.
pm.run.name = 'exercise_2'
# turn off exact so that it won't rerun and overwite the previous data
pm.run.EXT = False
# turn off any unrelated parameters
pm.run.NON = False
# Turning on reverse-engineering
pm.ext.RE = True
#results = pm.execute()
# Plotting the results
ex4_RE = rs.read("gs_extre_vks",pm)
plt.plot(ex4_RE, label="reverse-engineered ext")
plt.legend()
plt.show()
| 02_get_started_further/get_started_further.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false
# Initialize Otter
import otter
grader = otter.Notebook("hw07.ipynb")
# -
# # Homework 7: Testing Hypotheses
# **Reading**:
# * [Testing Hypotheses](https://www.inferentialthinking.com/chapters/11/testing-hypotheses.html)
# Please complete this notebook by filling in the cells provided.
#
# Directly sharing answers is not okay, but discussing problems with the course staff or with other students is encouraged. Refer to the policies page to learn more about how to learn cooperatively.
#
# For all problems that you must write our explanations and sentences for, you **must** provide your answer in the designated space. Moreover, throughout this homework and all future ones, please be sure to not re-assign variables throughout the notebook! For example, if you use `max_temperature` in your answer to one question, do not reassign it later on.
# +
# Don't change this cell; just run it.
import numpy as np
from datascience import *
# These lines do some fancy plotting magic.
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
import warnings
warnings.simplefilter('ignore', FutureWarning)
# -
# ## 1. Spam Calls
#
# ## Part 1: 781 Fun
# Yanay gets a lot of spam calls. An area code is defined to be a three digit number from 200-999 inclusive. In reality, many of these area codes are not in use, but for this question we'll simplify things and assume they all are. **Throughout these questions, you should assume that Yanay's area code is 781.**
# + [markdown] deletable=false editable=false
# **Question 1.** Assuming each area code is just as likely as any other, what's the probability that the area code of two back to back spam calls are 781?
#
#
# <!--
# BEGIN QUESTION
# name: q1_1
# manual: false
# -->
# + manual_grade=true manual_problem_id="catching_cheaters_1"
prob_781 = ...
prob_781
# + deletable=false editable=false
grader.check("q1_1")
# + [markdown] deletable=false editable=false
# **Question 2.** Rohan already knows that Yanay's area code is 781. Rohan randomly guesses the last 7 digits (0-9 inclusive) of his phone number. What's the probability that Rohan correctly guesses Yanay's number, assuming he’s equally likely to choose any digit?
#
# *Note: A phone number contains an area code and 7 additional digits, i.e. xxx-xxx-xxxx*
#
# <!--
# BEGIN QUESTION
# name: q1_2
# manual: false
# -->
# -
prob_yanay_num = ...
prob_yanay_num
# + deletable=false editable=false
grader.check("q1_2")
# -
# Yanay suspects that there's a higher chance that the spammers are using his area code (781) to trick him into thinking it's someone from his area calling him. Ashley thinks that this is not the case, and that spammers are just choosing area codes of the spam calls at random from all possible area codes (*Remember, for this question we’re assuming the possible area codes are 200-999, inclusive*). Yanay wants to test his claim using the 50 spam calls he received in the past month.
#
# Here's a dataset of the area codes of the 50 spam calls he received in the past month.
# Just run this cell
spam = Table().read_table('spam.csv')
spam
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 3.** Define the null hypothesis and alternative hypothesis for this investigation.
#
# *Hint: Don’t forget that your null hypothesis should fully describe a probability model that we can use for simulation later.*
#
#
# <!--
# BEGIN QUESTION
# name: q1_3
# manual: true
# -->
# -
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# **Question 4.** Which of the following test statistics would be a reasonable choice to help differentiate between the two hypotheses?
#
# *Hint*: For a refresher on choosing test statistics, check out the textbook section on [Test Statistics](https://www.inferentialthinking.com/chapters/11/3/decisions-and-uncertainty.html#Step-2:-The-Test-Statistic).
#
# 1. The proportion of area codes that are 781 in 50 random spam calls
# 2. The total variation distance (TVD) between probability distribution of randomly chosen area codes, and the observed distribution of area codes. (*Remember the possible area codes are 200-999 inclusive*)
# 3. The probability of getting an area code of 781 out of all the possible area codes.
# 4. The proportion of area codes that are 781 in 50 random spam calls divided by 2
# 5. The number of times you see the area code 781 in 50 random spam calls
#
# Assign `reasonable_test_statistics` to an array of numbers corresponding to these test statistics.
#
# <!--
# BEGIN QUESTION
# name: q1_4
# manual: false
# -->
# -
reasonable_test_statistics = ...
# + deletable=false editable=false
grader.check("q1_4")
# + [markdown] deletable=false editable=false
# <div class="hide">\pagebreak</div>
#
# **For the rest of this question, suppose you decide to use the number of times you see the area code 781 in 50 spam calls as your test statistic.**
#
# **Question 5.**
# Write a function called `simulate` that generates exactly one simulated value of your test statistic under the null hypothesis. It should take no arguments and simulate 50 area codes under the assumption that the result of each area is sampled from the range 200-999 inclusive with equal probability. Your function should return the number of times you saw the 781 area code in those 50 random spam calls.
#
#
# <!--
# BEGIN QUESTION
# name: q1_5
# manual: false
# -->
# +
possible_area_codes = ...
def simulate():
...
# Call your function to make sure it works
simulate()
# + deletable=false editable=false
grader.check("q1_5")
# + [markdown] deletable=false editable=false
# **Question 6.** Generate 20,000 simulated values of the number of times you see the area code 781 in 50 random spam calls. Assign `test_statistics_under_null` to an array that stores the result of each of these trials.
#
# *Hint*: Use the function you defined in Question 5.
#
# <!--
# BEGIN QUESTION
# name: q1_6
# manual: false
# -->
# + for_assignment_type="solution"
test_statistics_under_null = ...
repetitions = ...
...
test_statistics_under_null
# + deletable=false editable=false
grader.check("q1_6")
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 7.** Using the results from Question 6, generate a histogram of the empirical distribution of the number of times you saw the area code 781 in your simulation. **NOTE: Use the provided bins when making the histogram**
#
# <!--
# BEGIN QUESTION
# name: q1_7
# manual: true
# -->
# + export_pdf=true for_assignment_type="solution"
bins = np.arange(0,5,1) # Use these provided bins
...
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <div class="hide">\pagebreak</div>
#
# **Question 8.** Compute an empirical P-value for this test.
#
#
# <!--
# BEGIN QUESTION
# name: q1_8
# manual: false
# -->
# + for_assignment_type="solution"
# First calculate the observed value of the test statistic from the `spam` table.
observed_val = ...
p_value = ...
p_value
# + deletable=false editable=false
grader.check("q1_8")
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 9.** Suppose you use a P-value cutoff of 1%. What do you conclude from the hypothesis test? Why?
#
# <!--
# BEGIN QUESTION
# name: q1_9
# manual: true
# -->
# -
# _Type your answer here, replacing this text._
# <!-- END QUESTION -->
#
#
#
# ## Part 2: Multiple Spammers
# Instead of checking if the area code is equal to his own, Yanay decides to check if the area code matches the area code of one of the 8 places he's been to recently, and wants to test if it's more likely to receive a spam call with an area code from any of those 8 places. These are the area codes of the places he's been to recently: 781, 617, 509, 510, 212, 858, 339, 626.
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 10.** Define the null hypothesis and alternative hypothesis for this investigation.
#
# *Reminder: Don’t forget that your null hypothesis should fully describe a probability model that we can use for simulation later.*
#
#
# <!--
# BEGIN QUESTION
# name: q1_10
# manual: true
# -->
# -
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <div class="hide">\pagebreak</div>
#
# **Suppose you decide to use the number of times you see any of the area codes of the places Yanay has been to in 50 spam calls as your test statistic.**
#
# **Question 11.**
# Write a function called `simulate_visited_area_codes` that generates exactly one simulated value of your test statistic under the null hypothesis. It should take no arguments and simulate 50 area codes under the assumption that the result of each area is sampled from the range 200-999 inclusive with equal probability. Your function should return the number of times you saw any of the area codes of the places Yanay has been to in those 50 spam calls.
#
# *Hint*: You may find the textbook [section](https://www.inferentialthinking.com/chapters/11/1/Assessing_Models#Predicting-the-Statistic-Under-the-Model) on the `sample_proportions` function to be useful.
#
# <!--
# BEGIN QUESTION
# name: q1_11
# manual: false
# -->
# +
model_proportions = make_array(8/800, 792/800)
def simulate_visited_area_codes():
...
# Call your function to make sure it works
simulate_visited_area_codes()
# + deletable=false editable=false
grader.check("q1_11")
# + [markdown] deletable=false editable=false
# **Question 12.** Generate 20,000 simulated values of the number of times you see any of the area codes of the places Yanay has been to in 50 random spam calls. Assign `test_statistics_under_null` to an array that stores the result of each of these trials.
#
# *Hint*: Use the function you defined in Question 11.
#
# <!--
# BEGIN QUESTION
# name: q1_12
# manual: false
# -->
# + for_assignment_type="solution"
visited_test_statistics_under_null = ...
repetitions = ...
...
visited_test_statistics_under_null
# + deletable=false editable=false
grader.check("q1_12")
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 13.** Using the results from Question 12, generate a histogram of the empirical distribution of the number of times you saw any of the area codes of the places Yanay has been to in your simulation. **NOTE: Use the provided bins when making the histogram**
#
# <!--
# BEGIN QUESTION
# name: q1_13
# manual: true
# -->
# + export_pdf=true for_assignment_type="solution"
bins_visited = np.arange(0,6,1) # Use these provided bins
...
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <div class="hide">\pagebreak</div>
#
# **Question 14.** Compute an empirical P-value for this test.
#
#
# <!--
# BEGIN QUESTION
# name: q1_14
# manual: false
# -->
# + for_assignment_type="solution"
visited_area_codes = make_array(781, 617, 509, 510, 212, 858, 339, 626)
# First calculate the observed value of the test statistic from the `spam` table.
visited_observed_value = ...
p_value = ...
p_value
# + deletable=false editable=false
grader.check("q1_14")
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 15.** Suppose you use a P-value cutoff of 0.05% (**Note: that’s 0.05%, not our usual cutoff of 5%**). What do you conclude from the hypothesis test? Why?
#
# <!--
# BEGIN QUESTION
# name: q1_15
# manual: true
# -->
# -
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# **Question 16.** Is `p_value`:
#
# * (a) the probability that the spam calls favored the visited area codes,
# * (b) the probability that they didn't favor, or
# * (c) neither
#
# If you chose (c), explain what it is instead.
#
#
# <!--
# BEGIN QUESTION
# name: q1_16
# manual: true
# -->
# -
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# **Question 17.** Is 0.05% (the P-value cutoff):
#
# * (a) the probability that the spam calls favored the visited area codes,
# * (b) the probability that they didn't favor, or
# * (c) neither
#
# If you chose (c), explain what it is instead.
#
# <!--
# BEGIN QUESTION
# name: q1_17
# manual: true
# -->
# -
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# **Question 18.** Suppose you run this test for 4000 different people after observing each person's last 50 spam calls. When you reject the null hypothesis for a person, you accuse the spam callers of favoring the area codes that person has visited. If the spam callers were not actually favoring area codes that people have visited, can we compute how many times we will incorrectly accuse the spam callers of favoring area codes that people have visited? If so, what is the number? Explain your answer. Assume a 0.05% P-value cutoff.
#
# <!--
# BEGIN QUESTION
# name: q1_18
# manual: true
# -->
# -
# _Type your answer here, replacing this text._
# <!-- END QUESTION -->
#
#
#
# ## Part 3: Practice with A/B Tests
# Yanay collects information about this month's spam calls. The table `with_labels` is a sampled table, where the `Area Code Visited` column contains either `"Yes"` or `"No"` which represents whether or not Yanay has visited the location of the area code. The `Picked Up` column is `1` if Yanay picked up and `0` if he did not pick up.
# + deletable=false editable=false
# Just run this cell
with_labels = Table().read_table("spam_picked_up.csv")
with_labels
# -
# Yanay is going to perform an A/B Test to see whether or not he is more likely to pick up a call from an area code he has visited. Specifically, his null hypothesis is that there is no difference in the distribution of calls he picked up between visited and not visited area codes, with any difference due to chance. His alternative hypothesis is that there is a difference between the two categories, specifically that he thinks that he is more likely to pick up if he has visited the area code. We are going to perform a [permutation test](https://www.inferentialthinking.com/chapters/12/1/AB_Testing.html#Permutation-Test) to test this. Our test statistic will be the difference in proportion of calls picked up between the area codes Yanay visited and the area codes he did not visit.
# + [markdown] deletable=false editable=false
# **Question 19.** Complete the `difference_in_proportion` function to have it calculate this test statistic, and use it to find the observed value. The function takes in a sampled table which can be any table that has the same columns as `with_labels`. We'll call `difference_in_proportion` with the sampled table `with_labels` in order to find the observed difference in proportion.
#
# <!--
# BEGIN QUESTION
# name: q1_19
# manual: false
# -->
# +
def difference_in_proportion(sample):
# Take a look at the code for `proportion_visited` and use that as a
# hint of what `proportions` should be assigned to
proportions = ...
proportion_visited = proportions.where("Area Code Visited", "Yes").column("Picked Up mean").item(0)
proportion_not_visited = proportions.where("Area Code Visited", "No").column("Picked Up mean").item(0)
...
observed_diff_proportion = difference_in_proportion(with_labels)
observed_diff_proportion
# + deletable=false editable=false
grader.check("q1_19")
# + [markdown] deletable=false editable=false
# **Question 20.** To perform a permutation test we shuffle the labels, because our null hypothesis is that the labels don't matter because the distribution of calls he picked up between visited and not visited area codes come from same underlying distribution. The labels in this case is the `"Area Code Visited"` column containing `"Yes"` and `"No"`.
#
# Write a function to shuffle the table and return a test statistic using the function you defined in question 19.
#
# *Hint: To shuffle labels, we sample without replacement and then replace the appropriate column with the new shuffled column.*
#
# <!--
# BEGIN QUESTION
# name: q1_20
# manual: false
# -->
# +
def simulate_one_stat():
shuffled = ...
original_with_shuffled_labels = ...
return difference_in_proportion(original_with_shuffled_labels)
one_simulated_test_stat = simulate_one_stat()
one_simulated_test_stat
# + deletable=false editable=false
grader.check("q1_20")
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 21.** Generate 1,000 simulated test statistic values. Assign `test_stats` to an array that stores the result of each of these trials.
#
# *Hint*: Use the function you defined in Question 20.
#
# We also provided code that'll generate a histogram for you after generating a 1000 simulated test statistic values.
#
# <!--
# BEGIN QUESTION
# name: q1_21
# manual: true
# -->
# + export_pdf=true
trials = ...
test_stats = ...
...
# here's code to generate a histogram of values and the red dot is the observed value
Table().with_column("Simulated Proportion Difference", test_stats).hist("Simulated Proportion Difference");
plt.plot(observed_diff_proportion, 0, 'ro', markersize=15);
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# **Question 22.** Compute the empirical p-value for this test, and assign it to `p_value_ab`.
#
# <!--
# BEGIN QUESTION
# name: q1_22
# manual: false
# -->
# -
p_value_ab = ...
p_value_ab
# + deletable=false editable=false
grader.check("q1_22")
# -
# For `p_value_ab`, you should be getting a value around 10-15%. If our p-value cutoff is 5%, the data is more consistent with the null hypothesis - that there is no difference in the distribution of calls Yanay picked up between visited and not visited area codes.
# + [markdown] deletable=false editable=false
# ---
#
# To double-check your work, the cell below will rerun all of the autograder tests.
# + deletable=false editable=false
grader.check_all()
# + [markdown] deletable=false editable=false
# ## Submission
#
# Make sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output. The cell below will generate a zip file for you to submit. **Please save before exporting!**
# + deletable=false editable=false
# Save your notebook first, then run this cell to export your submission.
grader.export()
# -
#
| hw/hw07/.ipynb_checkpoints/hw07-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Introducción al Cálculo Científico
# + [markdown] slideshow={"slide_type": "slide"}
# En esta clase introduciremos algunos conceptos de computación cientifica en Python, principalmente utilizando la biblioteca `NumPy`, piedra angular de otras librerías científicas.
# + [markdown] slideshow={"slide_type": "slide"}
# ## SciPy.org
# -
# **SciPy** es un ecosistema de software _open-source_ para matemática, ciencia y engeniería. Las principales bibliotecas son:
#
# * NumPy: Arrays N-dimensionales. Librería base, integración con C/C++ y Fortran.
# * SciPy library: Computación científica (integración, optimización, estadística, etc.)
# * Matplotlib: Visualización 2D:
# * IPython: Interactividad (Project Jupyter).
# * SimPy: Matemática Simbólica.
# * Pandas: Estructura y análisis de datos.
# ## Numpy
# NumPy es el paquete fundamental para la computación científica en Python. Proporciona un objeto de matriz multidimensional, varios objetos derivados (como matrices y arreglos) y una variedad de rutinas para operaciones rápidas en matrices, incluida la manipulación matemática, lógica, de formas, clasificación, selección, I/O, transformadas discretas de Fourier, álgebra lineal básica, operaciones estadísticas básicas, simulación y mucho más. [Fuente.](https://numpy.org/devdocs/user/whatisnumpy.html)
# Para comenzar, la forma usual de importar `NumPy` es utilizando el alias `np`. Lo verás así en una infinidad de ejmplos, libros, blogs, etc.
import numpy as np
# ### Lo básico
# Los objetos principales de Numpy son los comúnmente conocidos como NumPy Arrays (la clase se llama `ndarray`), corresponden a una tabla de elementos, todos del mismo tipo, indexados por una tupla de enternos no-negativos. En NumPy, las dimensiones son llamadas `axes` (ejes) y su singular `axis` (eje), similar a un plano cartesiano generalizado. Esta parte de la clase está basada en el _Quickstart tutorial_ en la página oficial ([link](https://numpy.org/devdocs/user/quickstart.html)).
#
# Instanciar un NumPy Array es simple es utilizando el constructor propio de la biblioteca.
a = np.array(
[
[ 0, 1, 2, 3, 4],
[ 5, 6, 7, 8, 9],
[10, 11, 12, 13, 14]
]
)
type(a)
# Los atributos más importantes de un `ndarray` son:
a.shape # the dimensions of the array.
a.ndim # the number of axes (dimensions) of the array.
a.size # the total number of elements of the array.
a.dtype # an object describing the type of the elements in the array.
a.itemsize # the size in bytes of each element of the array.
# ### Crear Numpy Arrays
# Hay varias formas de crear arrays, el constructor básico es el que se utilizó hace unos momentos, `np.array`. El _type_ del array resultante es inferido de los datos proporcionados.
# +
a_int = np.array([2, 6, 10])
a_float = np.array([2.1, 6.1, 10.1])
print(f"a_int: {a_int.dtype.name}")
print(f"a_float: {a_float.dtype.name}")
# -
# También es posible utilizar otras estructuras de Python, como listas o tuplas.
a_list = [1, 1, 2, 3, 5]
np.array(a_list)
a_tuple = (1, 1, 1, 3, 5, 9)
np.array(a_tuple)
# __¡Cuidado!__ Es fácil confundirse con las dimensiones o el tipo de argumento en los contructores de NumPy, por ejemplo, utilizando una lista podríamos crear un arreglo de una o dos dimensiones si no tenemos cuidado.
one_dim_array = np.array(a_list)
two_dim_array = np.array([a_list])
print(f"np.array(a_list) = {one_dim_array} tiene shape: {one_dim_array.shape}, es decir, {one_dim_array.ndim} dimensión(es).")
print(f"np.array([a_list]) = {two_dim_array} tiene shape: {two_dim_array.shape}, es decir, {two_dim_array.ndim} dimensión(es).")
# Una funcionalidad útil son los constructores especiales a partir de constantes.
np.zeros((3, 4))
np.ones((2, 3, 4), dtype=np.int) # dtype can also be specified
np.identity(4) # Identity matrix
# Por otro lado, NumPy proporciona una función análoga a `range`.
range(10)
type(range(10))
np.arange(10)
type(np.arange(10))
np.arange(3, 10)
np.arange(2, 20, 3, dtype=np.float)
np.arange(9).reshape(3, 3)
# __Bonus:__ Utilizar `np.arange` tiene como _"ingredientes"_ el inicio (_start_), fin (_stop_) y el tamaño del espacio entre valores (_step_) y el largo (`len`) depende estos argumentos. Sin embargo, existe la función `np.linspace` que construye un `np.array` con un inicio y un fin, pero indicando la cantidad de elementos (y por lo tanto, el espaciado depende es este).
np.linspace(0, 100, 5)
# Esto puede causar confusiones en ocasiones, pues recuerda que la indexación de Python (y por lo tanto NumPy) comienza en cero, por lo que si quieres replicar el `np.array` anterior con `np.arange` debes tener esto en consideración. Es decir:
np.arange(start=0, stop=100, step=25) # stop = 100
np.arange(start=0, stop=101, step=25) # stop = 101
# No podía faltar la instanciación a través de elementos aleatorios
np.random.random(size=3) # Elementos entre 0 y 1
np.random.uniform(low=3, high=7, size=5) # Desde una distribución uniforme
np.random.normal(loc=100, scale=10, size=(2, 3)) # Desde una distribución normal indicando media y desviación estándar
# ### Acceder a los elementos de un array
# Es muy probable que necesites acceder a elementos o porciones de un array, para ello NumPy tiene una sintáxis consistente con Python.
x1 = np.arange(0, 30, 4)
x2 = np.arange(0, 60, 3).reshape(4, 5)
print("x1:")
print(x1)
print("\nx2:")
print(x2)
x1[1] # Un elemento de un array 1D
x1[:3] # Los tres primeros elementos
x2[0, 2] # Un elemento de un array 2D
x2[0] # La primera fila
x2[:, 1] # Todas las filas y la segunda columna
x2[:, 1:3] # Todas las filas y de la segunda a la tercera columna
# Nuevamente, recordar que Python tiene indexación partiendo desde cero. Además, la dimensión del arreglo también depende de la forma en que se haga la selección.
x2[:, 2]
x2[:, 2:3] # What?!
# En el ejemplo anterior los valores son los mismos, pero las dimensiones no. En el primero se utiliza `indexing` para acceder a la tercera columna, mientras que en el segundo `slicing` para acceder desde la tercera columna a la tercera columna.
print(x2[:, 2].shape)
print(x2[:, 2:3].shape)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Operaciones Básias
# -
# Numpy provee operaciones vectorizadas, con tal de mejorar el rendimiento de la ejecución.
# Por ejemplo, pensemos en la suma de dos arreglos 2D.
A = np.random.random((5,5))
B = np.random.random((5,5))
# Con los conocimientos de la clase pasada, podríamos pensar en iterar a través de dos `for`, con tal de llenar el arreglo resultando. algo así:
def my_sum(A, B):
n, m = A.shape
C = np.empty(shape=(n, m))
for i in range(n):
for j in range(m):
C[i, j] = A[i, j] + B[i, j]
return C
# %timeit my_sum(A, B)
# Pero la suma de `ndarray`s es simplemente con el signo de suma (`+`):
# %timeit A + B
# Para dos arrays tan pequeños la diferencia de tiempo es considerable, ¡Imagina con millones de datos!
# Los clásicos de clásicos:
x = np.arange(5)
print(f"x = {x}")
print(f"x + 5 = {x + 5}")
print(f"x - 5 = {x - 5}")
print(f"x * 2 = {x * 2}")
print(f"x / 2 = {x / 2}")
print(f"x // 2 = {x // 2}")
print(f"x ** 2 = {x ** 2}")
print(f"x % 2 = {x % 2}")
# ¡Júntalos como quieras!
-(0.5 + x + 3) ** 2
# Al final del día, estos son alias para funciones de Numpy, por ejemplo, la operación suma (`+`) es un _wrapper_ de la función `np.add`
np.add(x, 5)
# Podríamos estar todo el día hablando de operaciones, pero básicamente, si piensas en alguna operación lo suficientemente común, es que la puedes encontrar implementada en Numpy. Por ejemplo:
np.abs(-(0.5 + x + 3) ** 2)
np.log(x + 5)
np.exp(x)
np.sin(x)
# Para dimensiones mayores la idea es la misma, pero siempre hay que tener cuidado con las dimensiones y `shape` de los arrays.
print("A + B: \n")
print(A + B)
print("\n" + "-" * 80 + "\n")
print("A - B: \n")
print(A - B)
print("\n" + "-" * 80 + "\n")
print("A * B: \n")
print(A * B) # Producto elemento a elemento
print("\n" + "-" * 80 + "\n")
print("A / B: \n")
print(A / B) # División elemento a elemento
print("\n" + "-" * 80 + "\n")
print("A @ B: \n")
print(A @ B) # Producto matricial
# ### Operaciones Booleanas
print(f"x = {x}")
print(f"x > 2 = {x > 2}")
print(f"x == 2 = {x == 2}")
print(f"x == 2 = {x == 2}")
# +
aux1 = np.array([[1, 2, 3], [2, 3, 5], [1, 9, 6]])
aux2 = np.array([[1, 2, 3], [3, 5, 5], [0, 8, 5]])
B1 = aux1 == aux2
B2 = aux1 > aux2
print("B1: \n")
print(B1)
print("\n" + "-" * 80 + "\n")
print("B2: \n")
print(B2)
print("\n" + "-" * 80 + "\n")
print("~B1: \n")
print(~B1) # También puede ser np.logical_not(B1)
print("\n" + "-" * 80 + "\n")
print("B1 | B2 : \n")
print(B1 | B2)
print("\n" + "-" * 80 + "\n")
print("B1 & B2 : \n")
print(B1 & B2)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Broadcasting
# -
# ¿Qué pasa si las dimensiones no coinciden? Observemos lo siguiente:
a = np.array([0, 1, 2])
b = np.array([5, 5, 5])
a + b
# Todo bien, dos arrays 1D de 3 elementos, la suma retorna un array de 3 elementos.
a + 3
# Sigue pareciendo normal, un array 1D de 3 elementos, se suma con un `int`, lo que retorna un array 1D de tres elementos.
M = np.ones((3, 3))
M
M + a
# Magia! Esto es _broadcasting_. Una pequeña infografía para digerirlo:
# 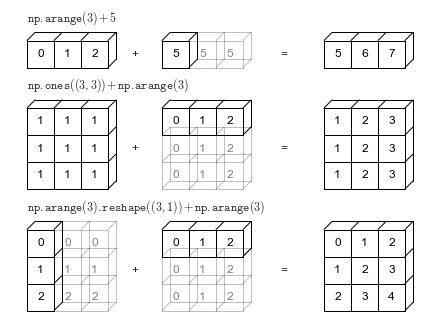
# Resumen: A lo menos los dos arrays deben coincidir en una dimensión. Luego, el array de dimensión menor se extiende con tal de ajustarse a las dimensiones del otro.
#
# La documentación oficial de estas reglas la puedes encontrar [aquí](https://numpy.org/devdocs/user/basics.broadcasting.html).
| lessons/M2L01_scientific_computing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="nCc3XZEyG3XV"
# Lambda School Data Science, Unit 2: Predictive Modeling
#
# # Applied Modeling, Module 1
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your decisions.
#
# - [X] Choose your target. Which column in your tabular dataset will you predict?
# - [X] Choose which observations you will use to train, validate, and test your model. And which observations, if any, to exclude.
# - [X] Determine whether your problem is regression or classification.
# - [X] Choose your evaluation metric.
# - [X] Begin with baselines: majority class baseline for classification, or mean baseline for regression, with your metric of choice.
# - [X] Begin to clean and explore your data.
# - [X] Choose which features, if any, to exclude. Would some features "leak" information from the future?
#
# ## Reading
# - [Attacking discrimination with smarter machine learning](https://research.google.com/bigpicture/attacking-discrimination-in-ml/), by Google Research, with interactive visualizations. _"A threshold classifier essentially makes a yes/no decision, putting things in one category or another. We look at how these classifiers work, ways they can potentially be unfair, and how you might turn an unfair classifier into a fairer one. As an illustrative example, we focus on loan granting scenarios where a bank may grant or deny a loan based on a single, automatically computed number such as a credit score."_
# - [How Shopify Capital Uses Quantile Regression To Help Merchants Succeed](https://engineering.shopify.com/blogs/engineering/how-shopify-uses-machine-learning-to-help-our-merchants-grow-their-business)
# - [Maximizing Scarce Maintenance Resources with Data: Applying predictive modeling, precision at k, and clustering to optimize impact](https://towardsdatascience.com/maximizing-scarce-maintenance-resources-with-data-8f3491133050), **by Lambda DS3 student** <NAME>. His blog post extends the Tanzania Waterpumps scenario, far beyond what's in the lecture notebook.
# - [Notebook about how to calculate expected value from a confusion matrix by treating it as a cost-benefit matrix](https://github.com/podopie/DAT18NYC/blob/master/classes/13-expected_value_cost_benefit_analysis.ipynb)
# - [Simple guide to confusion matrix terminology](https://www.dataschool.io/simple-guide-to-confusion-matrix-terminology/) by <NAME>, with video
# - [Visualizing Machine Learning Thresholds to Make Better Business Decisions](https://blog.insightdatascience.com/visualizing-machine-learning-thresholds-to-make-better-business-decisions-4ab07f823415)
# -
# Load the data so we can use it
import pandas as pd
bitcoin = pd.read_csv('Historical data for Bitcoin.csv')
bitcoin.head(10)
bitcoin.dtypes
# +
from tqdm import tnrange
# A little bit of feature engineering
bitcoin['numeric_date'] = bitcoin['Date'].copy()
bitcoin['avg_daily_price'] = (bitcoin['Open*']+bitcoin['High']+bitcoin['Low']+bitcoin['Close**'])/4
bitcoin['Date'] = pd.to_datetime(bitcoin['Date'])
bitcoin['Year'] = bitcoin['Date'].dt.year
bitcoin['Previous higher?'] = ""
for i in tnrange(len(bitcoin)-1,0,-1):
if(i>0):
bitcoin['Previous higher?'][i-1] = bitcoin['Low'][i].copy() > bitcoin['Low'][i-1].copy()
bitcoin['Previous higher?'][2297] = False
# -
new_columns = {True: 1, False:0}
bitcoin['Previous higher?'] = [new_columns[item] for item in bitcoin['Previous higher?']]
train = bitcoin[bitcoin.Year < 2016]
val = bitcoin[(bitcoin.Year >2015) & (bitcoin.Year < 2018)]
test = bitcoin[bitcoin.Year > 2017]
# Get the X and y parts for train, val, and test
drop_columns = ['Previous higher?','Date','numeric_date']
target = 'Previous higher?'
X_train = train.drop(columns=drop_columns)
y_train = train[target]
X_val = val.drop(columns=drop_columns)
y_val = val[target]
X_test = test.drop(columns=drop_columns)
# This is a classification problem.
accuracy = y_train.value_counts(normalize=True)[0]
print('Accuracy is', accuracy)
bitcoin.head()
# Lambda School Data Science, Unit 2: Predictive Modeling
#
# # Applied Modeling, Module 2
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your work.
#
# - [X] Plot the distribution of your target.
# - Regression problem: Is your target skewed? Then, log-transform it.
# - Classification: Are your classes imbalanced? Then, don't use just accuracy. And try `class_balance` parameter in scikit-learn.
# - [X] Continue to clean and explore your data. Make exploratory visualizations.
# - [X] Fit a model. Does it beat your baseline?
# - [ ] Share at least 1 visualization on Slack.
#
# You need to complete an initial model today, because the rest of the week, we're making model interpretation visualizations.
#
#
# ## Reading
#
# ### Today
# - [imbalance-learn](https://github.com/scikit-learn-contrib/imbalanced-learn)
# - [Learning from Imbalanced Classes](https://www.svds.com/tbt-learning-imbalanced-classes/)
# - [Machine Learning Meets Economics](http://blog.mldb.ai/blog/posts/2016/01/ml-meets-economics/)
# - [ROC curves and Area Under the Curve explained](https://www.dataschool.io/roc-curves-and-auc-explained/)
# - [The philosophical argument for using ROC curves](https://lukeoakdenrayner.wordpress.com/2018/01/07/the-philosophical-argument-for-using-roc-curves/)
#
#
# ### Yesterday
# - [Attacking discrimination with smarter machine learning](https://research.google.com/bigpicture/attacking-discrimination-in-ml/), by Google Research, with interactive visualizations. _"A threshold classifier essentially makes a yes/no decision, putting things in one category or another. We look at how these classifiers work, ways they can potentially be unfair, and how you might turn an unfair classifier into a fairer one. As an illustrative example, we focus on loan granting scenarios where a bank may grant or deny a loan based on a single, automatically computed number such as a credit score."_
# - [How Shopify Capital Uses Quantile Regression To Help Merchants Succeed](https://engineering.shopify.com/blogs/engineering/how-shopify-uses-machine-learning-to-help-our-merchants-grow-their-business)
# - [Maximizing Scarce Maintenance Resources with Data: Applying predictive modeling, precision at k, and clustering to optimize impact](https://towardsdatascience.com/maximizing-scarce-maintenance-resources-with-data-8f3491133050), **by Lambda DS3 student** <NAME>. His blog post extends the Tanzania Waterpumps scenario, far beyond what's in the lecture notebook.
# - [Notebook about how to calculate expected value from a confusion matrix by treating it as a cost-benefit matrix](https://github.com/podopie/DAT18NYC/blob/master/classes/13-expected_value_cost_benefit_analysis.ipynb)
# - [Simple guide to confusion matrix terminology](https://www.dataschool.io/simple-guide-to-confusion-matrix-terminology/) by <NAME>, with video
# - [Visualizing Machine Learning Thresholds to Make Better Business Decisions](https://blog.insightdatascience.com/visualizing-machine-learning-thresholds-to-make-better-business-decisions-4ab07f823415)
import seaborn as sns
y = bitcoin['Close**']
sns.distplot(y)
import numpy as np
y_log = np.log1p(y)
bitcoin['log(Bitcoin Close)'] = y_log
sns.distplot(bitcoin['log(Bitcoin Close)']);
# +
import matplotlib.pyplot as plt
import matplotlib
dates = matplotlib.dates.date2num(bitcoin['Date'])
plt.plot_date(dates, bitcoin['Close**']);
# +
from sklearn.pipeline import make_pipeline
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(),
RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
# -
pipeline.fit(X_train, y_train)
print('Validation Accuracy', pipeline.score(X_val, y_val))
bitcoin.dtypes
bitcoin.head(10)
| assignment_applied_modeling_1&2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Optimally Creating and Assigning Work Orders Based on Routes
#
# A pretty common task for organizations is optimally distributing work orders. Suppose our organization needs to perform restaurant/brewery inspections in the Greater Portland, Maine area. Let's assume that there are around 25 breweries that need to be inspected and that there are 5 workers that are available to do the inspections. As the supervisor of these workers I'm going to develop a Python Script (well, Jupyter Notebook in this case) that will optimally create distinct routes for my workers, create assignments at the hydrant locations, and then assign the assignment to the correct worker.
# ## Scenario 1: Creating and Assigning Assignments From Planned Routes
# In this scenario we are going to generate one route per worker. Each route will have up to 5 breweries (stored in an existing Feature Layer) that must be visisted and inspected. For each of the genenerated routes, we'll see which breweries need to be inspected and create assignments for them. We'll also assign the assignments to the worker that will be driving that route.
# ### Connecting to the Organization and Workforce project
# First let's connect to our GIS and fetch the Brewery Inspection Workforce Project.
# +
import pandas as pd
import arcgis
from arcgis.gis import GIS
from arcgis.apps import workforce
pd.options.display.max_columns = None
gis = GIS("https://arcgis.com", "workforce_scripts")
project = workforce.Project(gis.content.search("type:'Workforce Project' Brewery Inspections")[0])
# -
# ### Viewing the breweries that need to be inspected
# Now let's fetch the Breweries Feature Layer that our organization maintains.
breweries_item = gis.content.search("type:'Feature Service' owner:workforce_scripts Maine Breweries")[0]
breweries_item
# Let's query to find all of the breweries in the layer. You can see some of the detailed information in the dataframe below.
breweries_layer = breweries_item.layers[0]
breweries_filter = "location in ('Portland','South Portland','Gorham','Biddeford','Scarborough', 'Topsham','Freeport')"
breweries_df = breweries_layer.query(where=breweries_filter,out_fields="objectid,name,location,url", as_df=True)
breweries_df
# ### Creating optimal routes for each worker
# Now that we know what we're working with, let's use the [Plan Routes](https://doc.arcgis.com/en/arcgis-online/analyze/plan-routes.htm) tool to generate the most optimal routes for each of the workers. First we need to define where the workers will start their routes. Each worker will begin from the main office located at 100 Commercial Street, Portland Maine. We'll use the [geocoding module](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.geocoding.html#geocode) to get an exact location for this address.
from arcgis.geocoding import geocode
start_location = geocode("100 Commercial Street, Portland, ME", out_sr={"wkid": 102100})[0]["location"]
start_location
# Next, we need to convert this location into an in-memory feature layer that we can submit to the Plan Routes tools. First, we'll add the spatial reference to the location; this will help us later on when we need to create a feature collection.
start_location["spatialReference"] = {"wkid": 102100}
# Then we'll create a [Feature](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.features.toc.html#feature) from this location and supply a "Name" field.
feature = arcgis.features.Feature(
attributes={
"ObjectID": 1,
"Name": "Office"
},
geometry=start_location
)
# Next, we'll create a [Feature Set](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.features.toc.html#featureset) from the feature. Then we'll create a [Feature Collection](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.features.toc.html#featurecollection) from the Feature Set. Finally, we'll format the layer so that it conforms to the expected input format defined [here](https://doc.arcgis.com/en/arcgis-online/analyze/plan-routes.htm).
feature_set = arcgis.features.FeatureSet([feature])
feature_collection = arcgis.features.FeatureCollection.from_featureset(feature_set)
start_layer = {"layerDefinition": feature_collection.properties["layers"][0]["layerDefinition"], "featureSet": feature_set.value}
# Then we'll run the [Plan Routes](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.features.analysis.html#plan-routes) tool using the breweries layer as list of stops to route to. We'll set the number of routes equal to the number of workers. We'll also set the start time and start location as well as few other parameters.
from datetime import datetime
workers = project.workers.search()
breweries_layer.filter = breweries_filter
results = arcgis.features.analysis.plan_routes(breweries_layer, # Feature Layer of Stops
len(workers), # Number of routes to generate
5, # Maximum stops per route
datetime.now(), # Start time of route
start_layer, # The dictionary we created to represent the start location
stop_service_time=60, # How much time in minutes to spend at each stop
max_route_time=480, # The maximum time for the worker to complete the route
)
results
# As shown above, the output of the Plan Routes tool is a dictionary of 3 Feature Collections. One for the generated routes, one for the stops that were assigned to a route, and one for the stops that were not assigned a route. Let's see what information is provided in a route.
routes = results['routes_layer'].query().sdf
routes
# You can see that each route has a name, total time, and total distance among other things. Let's see what information is provided in an assigned stop.
stops = results['assigned_stops_layer'].query().sdf
stops
# You can see each row in the above table contains the attributes of each Brewery along with information about which route it is on. You'll also notice that there are several additional stops not related to a brewery. These are the starting and ending locations of each route.
# ### Create Assignment and Assign To Worker
# For each route that was generated we will select a random worker to complete that route. Then we'll find the breweries that were assigned to that route and create an Inspection Assignment for each one. Notice that when the assignment is created we are also assigning it to a worker.
#
# An important thing to note is that we are setting the due date of the assignment to the departure date of the stop. This means that a mobile worker will be able to sort their "To Do" list by due date and see the assignments in the correct order (according to the route).
# +
import random
assignments_to_add = []
for _, row in routes.iterrows():
worker = random.choice(workers)
workers.remove(worker)
route_stops = stops.loc[(stops['RouteName'] == row["RouteName"]) & stops['globalid'].notnull()]
for _, stop in route_stops.iterrows():
assignments_to_add.append(workforce.Assignment(
project,
assignment_type="Inspection",
location=stop["name"],
status="assigned",
worker=worker,
assigned_date=datetime.now(),
due_date=stop["DepartTime"],
geometry=stop["SHAPE"]
))
assignments = project.assignments.batch_add(assignments_to_add)
# -
# Let's see what this looks like in a map where the color of the assignment and route corresponds to the assigned worker. You can see that, in general, the colors are grouped together which is what we would expect. For example, the purple assignments are all placed on route that travels from Portland to Brunswick.
# 
# ## Scenario 2: Assigning Existing Assignments Based on Planned Routes
# In this scenario we already have assignments added to a Workforce Project. We are going to do something very similar to what we previously did, but this time instead of creating new assignments, we're going to assign workers to existing assignments. To accomplish this we'll generate one route per worker. Each route will have up to 5 assignments that must completed. For all of the assignments along each generated route, we'll assign a worker.
# ### Connecting to the Organization and Workforce project
# First let's connect to our GIS and fetch the Restaurant Inspections Workforce Project.
# +
import arcgis
from arcgis.gis import GIS
from arcgis.apps import workforce
gis = GIS("https://arcgis.com", "workforce_scripts")
project = workforce.Project(gis.content.search("type:'Workforce Project' Restaurant Inspections")[0])
# -
# ### Run Plan Routes tool using the assignments layer as the input
# Each worker will begin from the main office located at 100 Commercial Street, Portland Maine. We'll use the [geocoding module](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.geocoding.html#geocode) to get an exact location for this address.
from arcgis.geocoding import geocode
start_location = geocode("100 Commercial Street, Portland, ME", out_sr={"wkid": 102100})[0]["location"]
start_location
# Then we'll create a new [Feature](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.features.toc.html#feature) for this location.
# +
start_location["spatialReference"] = {"wkid": 102100}
start_location
feature = arcgis.features.Feature(
attributes={
"ObjectID": 1,
"Name": "Office"
},
geometry=start_location
)
# -
# Next, we'll create the layer that represents where the route should start.
feature_set = arcgis.features.FeatureSet([feature])
feature_collection = arcgis.features.FeatureCollection.from_featureset(feature_set)
start_layer = {"layerDefinition": feature_collection.properties["layers"][0]["layerDefinition"], "featureSet": feature_set.value}
# Next, we'll execute the [Plan Routes](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.features.analysis.html#plan-routes) tool using all unassigned assignments as the input layer. We'll set the number of routes equal to the number of workers. We'll allow up to 7 assignments per route.
from datetime import datetime
workers = project.workers.search()
assignments_filter = "status = 0"
project.assignments_layer.filter = assignments_filter
results = arcgis.features.analysis.plan_routes(project.assignments_layer, # Feature Layer of Stops
len(workers), # Number of routes to generate
7, # Maximum stops per route
datetime.now(), # Start time of route
start_layer, # The dictionary we created to represent the start location
stop_service_time=60, # How much time in minutes to spend at each stop
max_route_time=480, # The maximum time for the worker to complete the route
)
results
# Let's inspect the routes. There are 4 generated routes each with a unique name.
routes = results["routes_layer"].query().sdf
routes
# Let's inspect the assigned stops. Notice how all of the assignment fields are persisted.
stops = results["assigned_stops_layer"].query().sdf
stops
# ### Randomly assign a worker to each stop in a route
# To assign a worker to the assignments on a route, we will first fetch all of the unassigned assignments as well as all of the workers. Then we'll iterate over each route while selecting a random worker for that route. Then we'll iterate over all of the stops along that route. Next, we'll look up the specific assignment, using the GlobalID field in the stop, and assign it to that worker. Finally, we'll push our changes to the server using the batch update method.
import random
workers = project.workers.search()
assignments = project.assignments.search(assignments_filter)
assignments_dict = {assignment.global_id: assignment for assignment in assignments}
for _, row in routes.iterrows():
worker = random.choice(workers)
workers.remove(worker)
route_stops = stops.loc[(stops['RouteName'] == row["RouteName"]) & stops['GlobalID'].notnull()]
for _, stop in route_stops.iterrows():
# remove the brackets around the GlobalID field and lowercase it
g_id = stop["GlobalID"].replace("}","").replace("{","").lower()
assignments_dict[g_id].worker = worker
assignments_dict[g_id].assigned_date = datetime.now()
assignments_dict[g_id].status = "assigned"
assignments_dict[g_id].due_date = stop["DepartTime"]
assignments = project.assignments.batch_update(assignments)
# ## Summary
# We've demonstrated how work orders can be created and assigned on a per-route basis by using the Plan Routes tool. We've also shown how existing assignments can be assigned on a per-route basis. Workflows such as these can significantly improve the overall output by your workers by optimally assigning the work across time, space, and resources.
| notebooks/examples/4 - Optimally Creating and Assigning Work Orders Based on Routes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Set up
#
#
# ```{tip}
# Please do all of these steps before the first class!
#
# There is a youtube video at the bottom of this page, if you prefer that.
# ```
#
# Let's install our programming "stack".
#
# 1. [Create a GitHub account](https://github.com/join). Please use your name in your username to make discussions easier. Then,
# - [Fill out this survey so we know your username and get to know you](https://forms.gle/425BySmg2TxFMoid6).
# - Join the class's GitHub organization: Go to [coursesite](https://coursesite.lehigh.edu/course/view.php?id=236371), and click the first link under "Assignment links".
# - Please add a profile photo on GitHub, to make our exchanges and community more personal (don't be a "Twitter egg"!)
# - Optional: Sign up for a [student discount](https://education.github.com/discount_requests/new).
#
# 1. Install [Anaconda](https://www.anaconda.com/products/individual#Downloads) (not [the Nicki Minaj song](https://www.youtube.com/watch?v=LDZX4ooRsWs)).
# - _Install the latest version, make Anaconda your default Python installation, and accept all the defaults otherwise._
# - **Mac users**: The TA used the "64-Bit Graphical Installer (435 MB)"
# - Update Anaconda: Open up Anaconda Powershell Prompt (on windows, or terminal on Mac) and type
#
# `conda update --all`
#
# then click enter
#
# 4. Test start Jupyter Lab (simply see if you can open it, then close it)
# - Open Anaconda Prompt (or terminal on a Mac) and type `jupyter lab`, click enter, and an internet tab should open with the url `http://localhost:8888/lab`. This is Jupyter Lab, and where we will code this semester!
# - _The "point and click: option: Open `Anaconda Navigator` and launch Jupyter Lab from there_
# - _I recommend using the Powershell Prompt, because getting comfortable with that will be helpful_
# - Bonus: Play around and explore Jupyter Lab. [For a walkthough of what you're seeing, this page should help](https://jupyterlab.readthedocs.io/en/latest/user/interface.html).
#
# ```{tip}
# Once Jupyter Lab is open, it often helps to open a **second** powershell window. This will let you run terminal commands, which you can't do in the first powershell window (because that one is running jupyter lab).
# ```
#
# 2. Install [Git](https://git-scm.com/download/).
# - **Mac users**: Your TA installed Git via Homebrew. Go to [https://brew.sh](https://brew.sh), copy the home-brew address (they have a little copy paste icon which makes it easier for students), then type `brew install git` into the terminal and it'll install git. (There is no config necessary, type "y " when prompted with `y/n?`)
#
# ```{warning}
# If Mac users install Git via Xcode, you'll have to install Xcode and essentially waste 4gb of space you probably won't be using.
# ```
#
#
# 3. Install [GitHub Desktop](https://desktop.github.com/). _I accepted the default settings._
#
# ````{margin}
# ```{admonition} **Is this a "fun" fact?**
# :class: tip
# Git $\neq$ GitHub $\neq$ GitHub Desktop!
# 
#
# It is annoying that the names are so similar, but this will make more sense soon, I promise!
# ```
# ````
#
# 6. Optional, but recommended: Install a good text editor. [Atom](https://atom.io/), [Sublime](https://www.sublimetext.com/), or [Visual Studio (this link includes suggestions for extensions)](https://www.youtube.com/watch?v=jY0o1nkW0ow) are the most popular according to prior students of this class.
#
# 7. **Strongly** recommended: [Add some superpowers to JupyterLab to make coding easier](#step-7-setting-up-jupyterlab)
# ## Steps 1-6 on video
#
# I put together a video of me installing everything due to popular demand:
# <iframe width="560" height="315" src="https://www.youtube.com/embed/zKqkLEMmwBw" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen>Loading...</iframe>
#
# If you have completed all the steps above, you are ready to go! If you got stuck on a step, message the RA or post on the classmates group. We will get you going quickly!
# ## Step 7: Setting up JupyterLab
#
# These extensions will give you spell checking, code hints, and automatic formatting for your code 😍.
#
# 1. Run these in terminal (Mac) or Anaconda Powershell Prompt (Windows) one at a time:
# ```text
# pip install jupyterlab-spellchecker
# pip install jupyterlab_code_formatter
# pip install black
# pip install jupyterlab-lsp
# pip install python-lsp-server[all]
# pip install --upgrade lckr-jupyterlab-variableinspector
# ```
#
# - Mac users: To install the spellchecker, your TA had to install node.js in order to install the spell checker extension. MacOS users who also installed homebrew can install this in terminal via `brew install node`.
# - Windows users: The last `pip install` command often fails. [Use this fix.](https://github.com/lckr/jupyterlab-variableInspector/issues/183#issuecomment-806579896) As of 2022, this fix isn't working for everyone. If it doesn't work on your computer, don't worry - just skip ahead to the next step, and let me know.
#
# 2. Now run `jupyter lab` in the terminal window to open Jupterlab.
#
# 3. Click on settings, then Advanced Settings Editor. Click on the Keyboard Shortcuts menu. Then look to the right side of the screen: In the user preference box, add the text below. After you copy it in, save the user settings. (Either CTRL+S or using the the icon to the upper right of the user preference box.)
#
# ```text
# {
# "shortcuts": [
# {
# "command": "jupyterlab_code_formatter:format",
# "keys": [
# "Ctrl Shift F"
# ],
# "selector": ".jp-Notebook.jp-mod-editMode"
# }
# ]
# }
# ```
#
# - Mac users: You might want to use "Cmd" instead of "Ctrl"
#
# 4. Still in the Advanced Settings Editor, Click on the Code Completion menu. Then look to the right side of the screen: In the user preference box, add the text below. After you copy it in, save the user settings. (Either CTRL+S or using the the icon to the upper right of the user preference box.)
# ```text
# {
# "continuousHinting": true,
# }
# ```
#
| content/01/02_Setup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Test a Hilbert transform filter
import numpy as np
from scipy.fftpack import fft, fft2, ifft, ifft2, fftfreq, fftshift
from scipy import signal
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# Create a test signal with positive and negative frequencies
x = np.arange(0,1e5,500)
#lambda_x = [9923.0, -19923.] # Noninteger ratios of dx
lambda_x = [10000.0, -70000.0] # Integer ratios of dx
k1 = 2*np.pi/lambda_x[0]
k2 = 2*np.pi/lambda_x[1]
# Complex amplitude of each signal
A1 = 1.0 + 0*1j
A2 = 1.3 + 0*1j
y = A1*np.exp(1j*k1*x) + A2*np.exp(1j*k2*x)
plt.plot(x,np.real(y))
# +
# Compute the fft of the signal
M = x.size
# Window function
#W = np.hanning(M)
W = 1
Y = fft(y*W)
# Compute the frequencies (wavenumbers)
dx = x[1]-x[0]
k = fftfreq(M, dx/(2*np.pi))
dk = 1/(M*dx)
#
plt.plot(2*np.pi/k,Y,'k')
plt.plot(2*np.pi/k[:M//2],Y[:M//2]) # Positve frequencies
plt.plot(2*np.pi/k[M//2::],Y[M//2::],'r') # Negative frequencies
# +
# Design a butterworth filter
cutoff_x = 90000. # Close to the grid size
Wn = dx/cutoff_x
print Wn
b, a = signal.butter(3, Wn, btype='high', analog=False, output='ba')
w, h = signal.freqz(b, a, worN=k*dx)
plt.plot(2*np.pi*dx/w, h)
# -
plt.plot(abs(h[0:100]))
# +
# Filter using the butter worth weights
hpos = fftshift(h)
hpos[k<0] = 0
#hpos[:M//2] = h[:M//2]
ypos = ifft(Y*hpos)
hneg = fftshift(h)
hneg[k>0] = 0
yneg = ifft(Y*hneg)
plt.plot(x, ypos,'k')
plt.plot(x, yneg,'r')
plt.legend(('k > 0','k < 0'))
# -
# Calculate the maximum amplitudes of the positve and negative frequencies
print 'Amplitude of the maximum positive wavenumber', np.max( np.abs(Y*hpos)/M ), np.abs(A1)
print 'Amplitude of the maximum negative wavenumber', np.max( np.abs(Y*hneg)/M ), np.abs(A2)
# +
# Filter the positive wavenumbers
Yfiltpos = Y.copy()
Yfiltpos[k<0] = 0
Yfiltneg = Y.copy()
Yfiltneg[k>0] = 0
ypos = ifft(Yfiltpos)
yneg = ifft(Yfiltneg)
plt.plot(x, ypos,'k')
plt.plot(x, yneg,'r')
plt.legend(('k > 0','k < 0'))
yneg[10:-10].max(), ypos[10:-10].max()
# -
plt.plot(x, ifft(Y),'r',lw=3)
plt.plot(x,y)
plt.plot(x,ypos+yneg,'y')
# Calculate the maximum amplitudes of the positve and negative frequencies
print 'Amplitude of the maximum positive wavenumber', np.max( np.abs(Yfiltpos)/M ), np.abs(A1)
print 'Amplitude of the maximum negative wavenumber', np.max( np.abs(Yfiltneg)/M ), np.abs(A2)
# +
# Play around with Welch's method
print x.size, M
# #signal.welch?
next2 = int(2**np.ceil(np.log2(M)))
fs = 1/dx
nperseg = 2*next2
#f, Pxx = signal.welch(y, fs=fs)
f, Pxx = signal.welch(y, fs=fs, nperseg=nperseg, noverlap=next2//4, \
return_onesided=False)
w = np.hanning(M)
w = w / sum(w) # normalize analysis window
plt.figure()
plt.plot(f, Pxx)
# Filter the positive wavenumbers
Yfiltpos = Pxx.copy()
Yfiltpos[f<0] = 0
Yfiltneg = Pxx.copy()
Yfiltneg[f>0] = 0
ypos = ifft(Yfiltpos)
yneg = ifft(Yfiltneg)
# Calculate the maximum amplitudes of the positve and negative frequencies
print 'Amplitude of the maximum positive wavenumber', np.max( np.abs(Pxx[f<0])/M*fs ), np.abs(A1)
print 'Amplitude of the maximum negative wavenumber', np.max( np.abs(Pxx[f>0])/M*fs ), np.abs(A2)
plt.figure()
plt.plot(x, ypos*w,'k')
plt.plot(x, yneg*w,'r')
plt.legend(('k > 0','k < 0'))
# +
# Test the hilbert transform filter
Hx = signal.hilbert?
# -
Hx = signal.hilbert
# +
Hx = signal.hilbert(np.real(y))
z = y+1j*Hx
plt.plot(np.imag(Hx))
plt.plot(np.real(Hx))
# +
# Zero pad the edges
N = 2*int(2**np.ceil(np.log2(M)))
w = np.hamming(M)
w = w / sum(w) # normalize analysis window
#w = 1
#hN = (N/2)+1 # size of positive spectrum, it includes sample 0
#hM1 = int(np.floor((M+1)/2)) # half analysis window size by rounding
#hM2 = int(np.floor(M/2)) # half analysis window size by floor
hM1 = N//4
hM2 = hM1+M
fftbuffer = np.zeros(N) # initialize buffer for FFT
yw = y*w # window the input sound
fftbuffer[hM1:hM2] = yw # zero-pad
Y = fft(fftbuffer)
Ypos = Y.copy()
Ypos[N//2::] = Y[N//2::]
plt.figure()
plt.plot(fftbuffer)
plt.figure()
plt.plot(Y)
plt.figure()
plt.plot(ifft(Ypos))
# +
# Try zero buffering
# M = size of time series
# N = size of fft window
N = 4*int(2**np.ceil(np.log2(M)))
w = np.hamming(M)
w = w / sum(w) # normalize analysis window
hN = (N/2)+1 # size of positive spectrum, it includes sample 0
hM1 = int(np.floor((M+1)/2)) # half analysis window size by rounding
hM2 = int(np.floor(M/2)) # half analysis window size by floor
fftbuffer = np.zeros(N) # initialize buffer for FFT
yw = y*w # window the input sound
fftbuffer[:hM1] = yw[hM2:] # zero-phase window in fftbuffer
fftbuffer[-hM2:] = yw[:hM2]
Y = fft(fftbuffer)
absX = abs(Y[:hN]) # compute ansolute value of positive side
absX[absX<np.finfo(float).eps] = np.finfo(float).eps # if zeros add epsilon to handle log
mX = 20 * np.log10(absX) # magnitude spectrum of positive frequencies in dB
tol = 1e-14
Y[:hN].real[np.abs(Y[:hN].real) < tol] = 0.0 # for phase calculation set to 0 the small values
Y[:hN].imag[np.abs(Y[:hN].imag) < tol] = 0.0 # for phase calculation set to 0 the small values
pX = np.unwrap(np.angle(Y[:hN])) # unwrapped phase spectrum of positive frequencies
# Compute the frequencies (wavenumbers)
dx = x[1]-x[0]
k = fftfreq(N, dx/(2*np.pi))
dk = 1/(N*dx)
#
plt.figure(figsize=(12,6))
plt.plot(mX)
# Compute the inverse fft and undo the phase buffer
Ypos = np.zeros(N, dtype=np.complex)
Yneg = np.zeros(N, dtype=np.complex)
Ypos[:hN] = 10**(mX/20) * np.exp(1j*pX) # generate positive frequencies
Yneg[hN:] = 10**(mX[-2:0:-1]/20) * np.exp(-1j*pX[-2:0:-1]) # generate negative frequencies
fftbuffer = np.real(ifft(Ypos)) # compute inverse FFT
ypos = np.zeros(M)
ypos[:hM2] = fftbuffer[-hM2:] # undo zero-phase window
ypos[hM2:] = fftbuffer[:hM1]
fftbuffer = np.real(ifft(Yneg))
yneg = np.zeros(M)
yneg[:hM2] = fftbuffer[-hM2:] # undo zero-phase window
yneg[hM2:] = fftbuffer[:hM1]
plt.figure()
plt.plot(ypos/w)
plt.plot(yneg/w,'r')
#plt.plot(2*np.pi/k,Y,'k')
#plt.plot(2*np.pi/k[:M//2],Y[:M//2]) # Positve frequencies
#plt.plot(2*np.pi/k[M//2::],Y[M//2::],'r') # Negative frequencies
# -
# # Do the same in 2D
# +
# Create a test signal with positive and negative frequencies
x = np.arange(0,1e5,5e2)
y = np.arange(0,1e5,5e2)
xx, yy = np.meshgrid(x,y)
#lambda_x = [9923.0, -20017.0] # Noninteger ratios of dx
lambda_x = [10000.0, -20000.0] # Integer ratios of dx
lambda_y = [5000.0, -21002.] # Integer ratios of dy
k1 = 2*np.pi/lambda_x[0]
k2 = 2*np.pi/lambda_x[1]
l1 = 2*np.pi/lambda_y[0]
l2 = 2*np.pi/lambda_y[1]
# Complex amplitude of each signal
A1 = 1.0 + 0*1j
A2 = 1.3 + 0*1j
z = A1*np.exp(1j*(k1*xx+l1*yy)) + A2*np.exp(1j*(k2*xx+l2*yy))
plt.pcolormesh(x,y,np.real(z), cmap='PuOr')
plt.colorbar()
# +
# Compute the fft
Mx = x.size
My = y.size
Z = fft2(z, axes=(1,0))
# Compute zonal frequencies
dx = x[1]-x[0]
k = fftfreq(Mx, dx/(2*np.pi))
dk = 1/(Mx*dx)
# Compute meridional frequencies
dy = y[1]-y [0]
l = fftfreq(Mx, dx/(2*np.pi))
dl = 1/(My*dy)
# +
# Create filter matrices for each of the four quadrant
Z_posk_posl = np.zeros_like(Z)
Z_posk_posl[:My//2, :Mx//2] = Z[:My//2, :Mx//2]
z_posk_posl = ifft2(Z_posk_posl)
Z_posk_negl = np.zeros_like(Z)
Z_posk_negl[My//2::, :Mx//2] = Z[My//2::, :Mx//2]
z_posk_negl = ifft2(Z_posk_negl)
Z_negk_negl = np.zeros_like(Z)
Z_negk_negl[My//2::, Mx//2::] = Z[My//2::, Mx//2::]
z_negk_negl = ifft2(Z_negk_negl)
Z_negk_posl = np.zeros_like(Z)
Z_negk_posl[:My//2, Mx//2::] = Z[:My//2, Mx//2::]
z_negk_posl = ifft2(Z_negk_posl)
plt.figure(figsize=(12,12))
plt.subplot(221)
plt.pcolormesh(x,y,np.real(z_posk_negl), cmap='PuOr')
plt.colorbar()
plt.subplot(222)
plt.pcolormesh(x,y,np.real(z_posk_posl), cmap='PuOr')
plt.colorbar()
plt.subplot(223)
plt.pcolormesh(x,y,np.real(z_negk_negl), cmap='PuOr')
plt.colorbar()
plt.subplot(224)
plt.pcolormesh(x,y,np.real(z_negk_posl), cmap='PuOr')
plt.colorbar()
# -
print 'Amplitude of the maximum positive x and y wavenumber', np.max( np.abs(Z_posk_posl)/(Mx*My))
print 'Amplitude of the maximum negative x and y wavenumber', np.max( np.abs(Z_negk_negl)/(Mx*My) )
print 'Amplitude of the maximum -ve x and +ve y wavenumber', np.max( np.abs(Z_negk_posl)/(Mx*My) )
print 'Amplitude of the maximum +ve x and -ve y wavenumber', np.max( np.abs(Z_posk_negl)/(Mx*My) )
| hilbert_filter_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Maximum Entropy
#
# Information entropy is a function of probability distributions measure the quantity of information contained in a random variable (typically in bits). Consider a binary random variable that takes on two values, $H$ and $L$. The probability it takes the value $H$ is $p$ and the probability it takes value $L$ is $1-p$. Then the average information content of this random variable is measured as,
#
# \begin{align}
# H(p) &= -\left( p \log(p) + (1-p)\log(1-p) \right)
# \end{align}
#
# Choose the value of $p$ that maximizes the entropy function (i.e., maximizes the uncertainty, or information content of the binary random variable.)
from sympy import *
from sympy.plotting import plot
init_printing()
# +
# Declare our symbols and setup the
# entropy function.
p = Symbol('p', real=True, positive=True)
H = -1*(p * log(p) + (1 - p) * log(1-p))
Eq(Symbol('H(p)'), H)
# -
# Take the derivative with respect to
# the probability of H: p.
DH = H.diff(p)
Eq(Symbol('\\frac{\partial H(p)}{\partial p}'), DH)
# Set the derivative equal to zero
# and solve for p to find the value that
# satisfies the first order condition for
# a maximum.
p_star = solve(DH, p)[0]
Eq(Symbol('p^*'), p_star)
# Find the second derivative so we can
# test the second order condition for
# a maximum.
D2H = H.diff(p).diff(p)
Eq(Symbol('\\frac{\partial^2 H(p)}{\partial p^2}'), D2H)
# Evaluate the sign of the second derivative
# at the critical point we found, p*.
Eq(Symbol('\\frac{\partial^2 H(p)}{\partial p^2}|_{p^*}'), D2H.subs(p, p_star))
# As we can see, the second derivative is negative and the critical point we found $p^* = 1/2$ is a maximum. Thus, maximum entropy occurs at the uniform distribution $(p, 1-p) = (1/2, 1/2)$.
plot(H, (p, 0.01, 0.99))
| assets/pdfs/math_bootcamp/notebooks/maximum_entropy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from pyvesc.VESC import MultiVESC
class MobileBase:
def __init__(
self,
serial_port='/dev/ttyACM0',
left_wheel_id=24,
right_wheel_id=72,
back_wheel_id=None,
) -> None:
params = [
{'can_id': left_wheel_id, 'has_sensor': True, 'start_heartbeat': True},
{'can_id': right_wheel_id, 'has_sensor': True, 'start_heartbeat': True},
{'can_id': back_wheel_id, 'has_sensor': True, 'start_heartbeat': True},
]
self._multi_vesc = MultiVESC(serial_port=serial_port, vescs_params=params)
self.left_wheel, self.right_wheel, self.back_wheel = self._multi_vesc.controllers
omnibase = MobileBase()
# _____________
# RPM and Duty Cycle testing
# +
duty = 0
omnibase.back_wheel.set_duty_cycle(duty)
#omnibase.left_wheel.set_duty_cycle(duty)
#omnibase.right_wheel.set_duty_cycle(duty)
# -
omnibase.back_wheel.set_rpm(0)
# +
rpm = 0
omnibase.back_wheel.set_rpm(rpm)
omnibase.left_wheel.set_rpm(rpm)
omnibase.right_wheel.set_rpm(rpm)
# -
omnibase.back_wheel.set_rpm(100)
omnibase.back_wheel.set_rpm(0)
# With Joysticks
import pygame
import time
import numpy as np
# +
pygame.init()
pygame.joystick.init()
J = pygame.joystick.Joystick(0)
# -
# Close Loop
def speeds_from_joystick(joystick):
speed_max_t = 500
speed_max_r = 300
y = -joystick.get_axis(1) * speed_max_t
x = joystick.get_axis(0) * speed_max_t
rot = -joystick.get_axis(3) * speed_max_r
speed_back = x + rot
speed_right = (x*np.cos(120*np.pi/180)) + (y*np.sin(120*np.pi/180)) + rot
speed_left = (x*np.cos(240*np.pi/180)) + (y*np.sin(240*np.pi/180)) + rot
return int(speed_back), int(speed_right), int(speed_left)
while True:
for event in pygame.event.get():
pass
time.sleep(0.1)
omnibase.back_wheel.set_rpm(speeds_from_joystick(J)[0])
omnibase.left_wheel.set_rpm(speeds_from_joystick(J)[2])
omnibase.right_wheel.set_rpm(speeds_from_joystick(J)[1])
# Open Loop
# Duty Cycle
def cycle_from_joystick(joystick):
cycle_max_t = 0.4
cycle_max_r = 0.2
y = -joystick.get_axis(1) * cycle_max_t
x = joystick.get_axis(0) * cycle_max_t
rot = -joystick.get_axis(3) * cycle_max_r
cycle_back = x + rot
cycle_right = (x*np.cos(120*np.pi/180)) + (y*np.sin(120*np.pi/180)) + rot
cycle_left = (x*np.cos(240*np.pi/180)) + (y*np.sin(240*np.pi/180)) + rot
#print(cycle_back, cycle_left, cycle_right)
return (cycle_back), (cycle_right), (cycle_left)
while True:
for event in pygame.event.get():
pass
time.sleep(0.1)
omnibase.back_wheel.set_duty_cycle(cycle_from_joystick(J)[0])
omnibase.left_wheel.set_duty_cycle(cycle_from_joystick(J)[2])
omnibase.right_wheel.set_duty_cycle(cycle_from_joystick(J)[1])
# Current
def current_from_joystick(joystick):
current_max_t = 1
current_max_r = 0.5
y = -joystick.get_axis(1) * current_max_t
x = joystick.get_axis(0) * current_max_t
rot = -joystick.get_axis(3) * current_max_r
current_back = x + rot
current_right = (x*np.cos(120*np.pi/180)) + (y*np.sin(120*np.pi/180)) + rot
current_left = (x*np.cos(240*np.pi/180)) + (y*np.sin(240*np.pi/180)) + rot
#print(cycle_back, cycle_left, cycle_right)
return (current_back), (current_right), (current_left)
while True:
for event in pygame.event.get():
pass
time.sleep(0.1)
omnibase.back_wheel.set_current(current_from_joystick(J)[0])
omnibase.left_wheel.set_current(current_from_joystick(J)[2])
omnibase.right_wheel.set_current(current_from_joystick(J)[1])
| pyvesc/examples/test_omnibase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/TerraFusion/pytaf/blob/master/examples/notebook/pytaf_misr2ceres_pydap.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="FFzcpkU99wPO" colab_type="text"
# Copyright (C) 2019 The HDF Group
#
# This example code illustrates how to access Basic Fusion data product in Python via OPeNDAP.
#
# If you have any questions, suggestions, or comments on this example, please use the HDF-EOS Forum (http://hdfeos.org/forums).
#
# If you would like to see an example of any other NASA HDF/HDF-EOS data
# product that is not listed in the HDF-EOS Comprehensive Examples page
# (http://hdfeos.org/zoo), feel free to contact us at <EMAIL> or post it at the HDF-EOS Forum (http://hdfeos.org/forums).
#
# Tested under: Google CoLab with Python 3.0 runtime
#
# Last updated: 2019-05-28
#
#
# # Subset TerraFusion data via OPeNDAP using Pydap
# + id="YWcrh4S5G9Ef" colab_type="code" outputId="e275fc1f-043d-4cb8-d8df-4dafef3a93ce" colab={"base_uri": "https://localhost:8080/", "height": 957}
# !pip install pydap
# !pip install matplotlib
# !pip install pyproj==1.9.6
# !apt install proj-bin libproj-dev libgeos-dev
# !pip install https://github.com/matplotlib/basemap/archive/v1.2.0rel.tar.gz
# !pip install git+https://github.com/TerraFusion/pytaf.git
# + id="QiA8N9Zt9wPT" colab_type="code" outputId="eea98d6f-85ba-4f68-f0b8-a55923e7d0c4" colab={"base_uri": "https://localhost:8080/", "height": 295}
from pydap.client import open_url, open_dods
import numpy as np
import time
start_time = time.time()
server = 'https://eosdap.hdfgroup.org:8080/opendap/data/hdf5/'
file_name = 'TERRA_BF_L1B_O53557_20100112014327_F000_V001.h5'
url = server + file_name
lat_s = np.array([])
lon_s = np.array([])
data_s = np.array([])
data_v = np.array([])
# There are 3 CERES granules.
for i in range(1,4):
print(i)
varname = 'CERES_granule_201001120'+str(i)+'_FM1_Viewing_Angles_Solar_Zenith'
vzaname = 'CERES_granule_201001120'+str(i)+'_FM1_Viewing_Angles_Viewing_Zenith'
latname = 'CERES_granule_201001120'+str(i)+'_FM1_Time_and_Position_Latitude'
lonname = 'CERES_granule_201001120'+str(i)+'_FM1_Time_and_Position_Longitude'
dataset = open_dods(url+'.dods?'+varname)
data = dataset[varname].data
data_s = np.append(data_s, data)
print(data[0])
dataset_vza = open_dods(url+'.dods?'+vzaname)
datav = dataset_vza[vzaname].data
data_v = np.append(data_v, datav)
print(datav[0])
latset = open_dods(url+'.dods?'+latname)
lat = latset[latname].data
print(lat[0])
lat_s = np.append(lat_s, lat)
lonset = open_dods(url+'.dods?'+lonname)
lon = lonset[lonname].data
print(lon[0])
lon_s = np.append(lon_s, lon)
print("--- %s seconds ---" % (time.time() - start_time))
# + id="StrpDWBDuN25" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="324290bf-e4fc-44b0-8024-6192c5c4d7ec"
# Apply criteria.
idx = np.where((data_s>=0)&(data_s<=89)&(data_v>=0)&(data_v<=25))[0]
print(idx.shape)
target_lat = lat_s[idx]
target_lon = lon_s[idx]
# + colab_type="code" id="cEBJkbo9qUwU" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="8fca5d37-f16d-45bf-ef15-88ac9c52060b"
# ---- MISR part ----
varname = 'MISR_AN_Data_Fields_Red_Radiance'
latname = 'MISR_HRGeolocation_GeoLatitude'
lonname = 'MISR_HRGeolocation_GeoLongitude'
dataset = open_dods(url+'.dods?'+varname)
var = dataset[varname].data
# print(var[0])
latset = open_dods(url+'.dods?'+latname)
lat = latset[latname].data
# print(lat[0])
lonset = open_dods(url+'.dods?'+lonname)
lon = lonset[lonname].data
# print(lon[0])
# Convert 3-D MISR grids to 2-D.
src_var = np.vstack(var).astype(np.float64)
print(src_var.shape)
src_lat = np.vstack(lat).astype(np.float64)
src_lon = np.vstack(lon).astype(np.float64)
# Call resample using summary interpolation.
sd = np.zeros(src_var.shape, dtype=src_var.dtype)
npix = np.zeros(src_var.shape, dtype=np.int32)
max_radius = 20000
# Make copies of target lat/lon because resample will modify them.
tlat = target_lat.copy()
tlon = target_lon.copy()
print("--- %s seconds ---" % (time.time() - start_time))
# + id="iNYQWuEx5Ye-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 86} outputId="b7b00ea8-c5d2-418a-e263-2dcca7f9e7ca"
import pytaf
trg_data = pytaf.resample_s(src_lat, src_lon, target_lat, target_lon,
src_var, max_radius, sd, npix)
print(trg_data.shape)
print(trg_data.size)
print(trg_data)
print("--- %s seconds ---" % (time.time() - start_time))
# + id="kSrVeFoV4hZH" colab_type="code" colab={}
import h5py
# Write data for plotting.
f3 = h5py.File('misr2ceres.h5', 'w')
dset = f3.create_dataset('/Target/Data_Fields/MISR_AN_Red_Radiance', data=trg_data)
dset3 = f3.create_dataset('/Geolocation/Latitude', data=tlat)
dset4 = f3.create_dataset('/Geolocation/Longitude', data=tlon)
f3.close()
# + id="IvQGxxfhzGV7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 307} outputId="8f8217e9-a334-4d00-86b7-870bb3622918"
# Open the file.
file_name = 'misr2ceres.h5'
with h5py.File(file_name, 'r') as f:
# Read MISR Radiance dataset.
misr_dset = f['/Target/Data_Fields/MISR_AN_Red_Radiance']
misr_data = misr_dset[:].astype(np.float64)
ds_lat = f['/Geolocation/Latitude']
lat = ds_lat[:].astype(np.float64)
ds_lon = f['/Geolocation/Longitude']
lon = ds_lon[:].astype(np.float64)
import matplotlib as mpl
import matplotlib.pyplot as plt
from mpl_toolkits.basemap import Basemap
# Plot data.
data = misr_data
data[data == -999] = np.nan
datam = np.ma.masked_array(data, np.isnan(data))
m = Basemap(projection='cyl', resolution='l',
llcrnrlat=np.min(lat), urcrnrlat=np.max(lat),
llcrnrlon=np.min(lon), urcrnrlon=np.max(lon))
slat = (np.ceil(np.max(lat)) - np.floor(np.min(lat))) / 6.0
slon = (np.ceil(np.max(lon)) - np.floor(np.min(lon))) / 6.0
m.drawcoastlines(linewidth=0.5)
m.drawparallels(np.arange(np.floor(np.min(lat)),
np.ceil(np.max(lat)), slat),
labels=[1, 0, 0, 0])
m.drawmeridians(np.arange(np.floor(np.min(lon)),
np.ceil(np.max(lon)), slon),
labels=[0, 0, 0, 1])
m.scatter(lon, lat, c=datam, s=1, edgecolors=None, linewidth=0)
fig = plt.gcf()
fig.suptitle('{0}'.format(file_name))
pngfile = file_name+'.py.png'
fig.savefig(pngfile)
print("--- %s seconds ---" % (time.time() - start_time))
| examples/notebook/pytaf_misr2ceres_pydap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import pandas as pd
from pandas_profiling import ProfileReport
import matplotlib.pyplot as plt
import mplcyberpunk
# %matplotlib inline
# plt.style.use('ggplot')
plt.style.use("cyberpunk")
import os
os.getcwd()
# -
# Import Custom Modules
from src.helpers import *
from src.final_rec import *
# Import Data
items = pd.read_csv('data/items_final.csv')
users = pd.read_csv('data/user_final.csv')
chemicals = pd.read_csv('data/chemical_effects_final.csv')
# # LDA for Ingredient Features
import pyLDAvis
import pyLDAvis.sklearn
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.decomposition import LatentDirichletAllocation
pyLDAvis.enable_notebook()
# +
# Get Ingredients
ingredients = items['ingredients']
# Vectorize Ingredients
# Count Vec
cnt_vec = CountVectorizer(strip_accents='unicode', lowercase=True, token_pattern=r'\b[a-zA-Z]{3,}\b')
doc_term_matrix = cnt_vec.fit_transform(ingredients)
# TFIDF Vec
tfidf_vec = TfidfVectorizer(**cnt_vec.get_params())
doc_term_matrix_tfidf = tfidf_vec.fit_transform(ingredients)
# -
n_comp = 5
num_words = 10
# Perform LDA
# TF ONLY
lda_term_freq = LatentDirichletAllocation(n_components=n_comp, random_state=0)
lda_term_freq.fit(doc_term_matrix)
# TFIDF
lda_term_freq_idf = LatentDirichletAllocation(n_components=n_comp, random_state=0)
lda_term_freq_idf.fit(doc_term_matrix_tfidf)
# Print the topics found by the LDA model
print("Topics found via LDA:")
print_topics(lda_term_freq, cnt_vec, num_words)
# Visualize LDA
LDA_prepared = pyLDAvis.sklearn.prepare(lda_term_freq, doc_term_matrix_tfidf, cnt_vec)
# +
# Write to File
# pyLDAvis.save_html(LDA_prepared, 'media/previews/lda.html')
# -
# # Modeling with Surprise
# +
import surprise
from surprise import Dataset, Reader
from surprise import SVD
from surprise import accuracy
from surprise.model_selection import cross_validate
from surprise.model_selection import train_test_split
from surprise.model_selection import RandomizedSearchCV
from surprise.model_selection import train_test_split
from surprise.model_selection import LeaveOneOut
from surprise.model_selection import cross_validate
from surprise import KNNBaseline
from surprise import SVD, SVDpp, SlopeOne, NMF, NormalPredictor, KNNBaseline, KNNBasic, KNNWithMeans, KNNWithZScore, BaselineOnly, CoClustering
# -
# ## SVD
# +
# Start with SVD
data = users[['user_id', 'item_id', 'star_rating']]
reader = Reader(line_format='user item rating', sep=',')
data = Dataset.load_from_df(data, reader=reader)
trainset, testset = train_test_split(data, test_size=.5)
# -
# Train SVD Model
svd = SVD(n_epochs=100)
svd.fit(trainset)
svd_predictions = svd.test(testset)
accuracy.rmse(svd_predictions)
accuracy.mae(svd_predictions)
# Predictions look decent, with testing on different test sizes, it was found nearing 0.2 test_split resulted in best RMSE
# ## SVDpp
svd_pp = SVDpp(n_epochs=100)
svd_pp.fit(trainset)
svdpp_predictions = svd_pp.test(testset)
accuracy.rmse(svdpp_predictions)
accuracy.mae(svdpp_predictions)
# # Modeling with LightFM
from scipy import sparse
from lightfm import LightFM
from sklearn.metrics.pairwise import cosine_similarity
from lightfm.evaluation import auc_score
from lightfm.evaluation import precision_at_k, recall_at_k
# ### Collaborative Filtering
# Create Utility Matrix
utility_matrix = create_utility_mat(df=users, user_col='user_id', item_col = 'item_id', rating_col='star_rating')
utility_matrix
# +
# Run Matrix Factorization
n_comp = 30
loss_metric = 'warp'
k = 15
epoch = 30
n_jobs = 4
# Create Matrix Factorization Model
x = sparse.csr_matrix(utility_matrix.values)
mf_model = LightFM(no_components=n_comp,
loss=loss_metric,
k=k
)
mf_model.fit(x,
epochs=epoch,
num_threads=n_jobs
)
train_auc = auc_score(mf_model,
x,
num_threads=n_jobs).mean()
print('Collaborative filtering AUC: %s' % train_auc)
print("Train precision: %.4f" % precision_at_k(mf_model,
x,
k=k,
num_threads=n_jobs).mean())
# -
# Create Dictionaries for user and item for recommendations
user_dictionary = create_user_dict(utility_matrix)
product_dictionary = create_item_dict(users,'item_id', 'item_name')
user_id = 1
# Final Recomendations (sample)
recommendations = recommendation_user(mf_model, utility_matrix,
user_id, user_dictionary,
product_dictionary, 4,
5, True)
# # K Means Clustering of Users
from scipy.sparse import csr_matrix
from sklearn.cluster import KMeans
# Use CSR Matrix for Sparcity
sparse_ratings = csr_matrix(utility_matrix)
# sparse_ratings = utility_matrix
# Predict 20 clusters
predictions = KMeans(n_clusters=10, algorithm='full').fit_predict(sparse_ratings)
# +
# clustered.group.unique
# utility_matrix
# +
# Plot Heatmap
max_users = 2000
max_items = 10
clustered = pd.concat([utility_matrix.reset_index(), pd.DataFrame({'group':predictions})], axis=1)
draw_clusters(clustered, max_users, max_items)
| modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Combien de numéros de mobiles français sont des nombres premiers ?
#
# A question simple, réponse simple :
# ## Dépendances
from sympy import isprime
print(isprime.__doc__[:180])
# ## Réponse
# +
first_number = 6_00_00_00_00
last_number = 7_99_99_99_99
# test rapide
#last_number = first_number + 20
all_numbers = range(first_number, last_number + 1)
# -
def count_prime_numbers_in_range(some_range):
count = 0
for number in some_range:
if isprime(number):
count += 1
return count
# ## Conclusion
# +
count = count_prime_numbers_in_range(all_numbers)
print(f"Pour des numéros de téléphones, nombres entre {first_number} et {last_number} (inclus), il y a {count} nombres premiers.")
# -
# Et donc, on peut calculer la part de nombres premiers parmi les numéros de téléphones mobiles français.
# De souvenir, c'était environ 5.1%, vérifions :
# +
total_number = len(all_numbers)
print(f"Pour des numéros de téléphones, nombres entre {first_number} et {last_number} (inclus), il y a {count/total_number:%} nombres premiers.")
# -
# Et voilà, c'était simple !
| Combien_de_numeros_de_mobiles_francais_sont_des_nombres_premiers.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
// <div class="alert alert-block alert-info" style="margin-top: 20px">
// <a href="http://cocl.us/SC0101ENNotebookadd"><img src = "https://ibm.box.com/shared/static/kr8cc5o4va552yrgo83j54mishtgzj2n.png" width = 750, align = "center"></a>
//
//
//
//
// <a href="https://www.bigdatauniversity.com"><img src = "https://ibm.box.com/shared/static/ugcqz6ohbvff804xp84y4kqnvvk3bq1g.png" width = 300, align = "center"></a>
//
// <h1 align=center><font size = 5>Scala 101</font></h1>
// # Module 2 Basic Object Oriented Programming
// ## Table of contents
//
// <div class="alert alert-block alert-info" style="margin-top: 20px">
// <li><a href="#ref1">Classes </a>
// <li><a href="#ref2">Immutable and Mutable Fields </a>
// <li><a href="#ref3">Methods </a>
// <li><a href="#ref4">Default and Named Arguments </a>
// <li><a href="#ref5">Objects </a>
//
//
//
// <p></p>
// Estimated Time Needed: <strong>30 min</strong>
// </div>
// <a id="ref1"></a>
//
// # 2.1 Classes
// ### By the end of this section you should be able to:
//
// <ul>
// <li> Create a class in Scala </li>
// <li> Create a class with parameters </li>
// <li> Create an instance of a class (aka object) </li>
// </ul>
//
// ### Create a new class hello:
// We can create a class "Hello" as follows
class Hello
// We can create an instance of the class of type "Hello":
new Hello()
// This output is the location of the instance of the class inside the JVM , Hello() is Sometimes called the class constructor.
// Lets call a method that converts the instance to a string. We will go more in to methods later on:
(new Hello() ).toString()
// ## The body of a class
// The body of the class is contained in two curly brackets as shown below:
//
// <center> <font size="10" color="green">class </font> <font size="10" color="blue">Hello</font><font size="10" >{</font><font size="10" color="red"> Body of a Class </font><font size="10" >}</font></center>
// Anything in the body will run when an instance of the class is created, For example, we can create the class hello, in the body we can put a statement in the class to print out “hello” :
class Hello {print("hello")}
// When we create a new instance of the class hello, the new object runs the statement and prints out "hello"
new Hello()
// ## Class Parameters
// You can pass values into a class using parameters, parameters are placed inside the parentheses following the class name as shown here:
// <center> <font size="10" color="green">class </font> <font size="10" color="blue">Hello</font><font size="10" ><font size="10" >(</font><font size="10" color="red"> Parameters </font><font size="10" > <font size="10" >)</font>{</font><font size="10" color="red"> Body of a Class </font><font size="10" >}</font></center>
//
// You must specify the type of the parameters, for example we have the class "Hello", the parameter is a message of type string. We can use the parameter in the body of the class but the parameter is not visible outside the class
// +
class Hello( message:String){
println(message)
}
// -
// We can use the parameter in the body of the class, for example we print out the value of "message". When we create an instance of the class we will not have access to the parameter outside the instance:
new Hello("what up")
// We do not have access to the parameters:
(new Hello("what up")).message
// If a class has parameter values and we create an instance of that class without any parameter values we get an error:
new Hello()
// class parameters are not accessible
// <div class="alert alert-danger alertdanger" style="margin-top: 20px">
// <h1> Question 2.1: </h1>
// <p>
// Create an instance of the class Hello, set the parameter message to “Hi” .</b>
// </p>
// </div>
new Hello("Hi")
// <div align="right">
// <a href="#q1a" class="btn btn-default" data-toggle="collapse">Click here for the solution</a>
//
// </div>
// <div id="q1a" class="collapse">
// ```
// new Hello("Hi")
// ```
// </div>
// <a id="ref2"></a>
//
// # 2.2 Immutable and Mutable Fields
// ### By the end of this section you should be able to:
//
// <ul>
// <li>Describe the difference between mutable and immutable fields </li>
// <li>create fields in Scala classes</li>
// <li>describe the difference between class parameters and fields </li>
// <li>outline how to promote class parameters to fields </li>
// </ul>
//
// A Field is a value inside an instance of a class, it represents the state of the instance. Unlike parameters fields are inside the body of the class and accessible outside the instance of a class unless specified.
// ### Immutable fields
// Immutable fields cannot be changed, we use the “val” keyword to indicate the field is immutable. We can create a class hello, with the field “message” in the body:
class Hello1{val message="hello"}
// We can create an instance of the object Hello1 and access the field “message “ by using the dot notation :
(new Hello1).message
// Mutable fields can be changed, we use the “var” keyword to indicate the field is mutable. We can create a class hello, with the field “message” in the body:
class Hello2{var message="hello"}
// We can create an instance of the object Hello2 and access the field “message “ by using the dot notation :
val hello2= new Hello2
hello2.message
// We can change the field value:
hello2.message="good bye"
// Now the field value has changed for the instance of the class:
hello2.message
// If we tried something similar for the class Hello1 we would get an error because the field is a value and we cannot change a "val". Lets create an instance of class hello1,and assign it to the variable H1:
val H1=new Hello1()
H1.message
// If we try and change the field we get an error
hello1.message="good bye"
// ### Promoting class Parameters to become Fields
// you can convert a parameter a field by adding the key word “val” to the parameter in the class constructor (in this case, we are adding the "val" inside the round parenthesis, not the curly parenthesis)
class Hello(val message: String)
// Let’s create a new instance of the class with the parameter:
val hello=new Hello("Hello, world!")
// Because we have “val” key word we have access to the parameter :
hello.message
// <div class="alert alert-danger alertdanger" style="margin-top: 20px">
// <h1> Question 2.1: </h1>
// <p>
// The class person uses the persons birthday as a parameter, in the class body the variable year is the present year of 2017 and the age is the persons age , create an instants of class person and call it bob. Set the parameter to 1990. and show the field of age.</b>
// </p>
// </div>
class person( BirthYear:Int){
var year=2017
var age=year-BirthYear
}
var bob = new person(1990)
bob.age
// <div align="right">
// <a href="#q2a" class="btn btn-default" data-toggle="collapse">Click here for the solution</a>
//
// </div>
// <div id="q2a" class="collapse">
// ```
// var Bob=new person(1990)
// Bob.age
// or
//
// val Bob=new person(1990)
// Bob.age
//
//
// ```
// </div>
// <div class="alert alert-danger alertdanger" style="margin-top: 20px">
// <h1> Question 2.2: </h1>
// <p>
// Can you change the field year and what happens to age?
// </p>
// </div>
// <div align="right">
// <a href="#q2b" class="btn btn-default" data-toggle="collapse">Click here for the solution</a>
//
// </div>
// <div id="q2b" class="collapse">
// ```
// Yes the field is a variable, nothing happens to age as the fields change only when the object is created:
// Bob.year=20000
// Bob.age
// ```
// </div>
// <a id="ref3"> </a>
// ## 2.3 METHODS
// ### By the end of this section you should be able to:
//
// <ul>
// <li>Implement methods </li>
// <li>Described evaluation order of methods vs fields </li>
// <li>Outline how index notation works </li>
// </ul>
// Methods do work on the instance of the class, they may or may not take parameters, and they may return values. We create a method using the "def" key word.
// For example, we can create a method that returns the string "Hello":
def hello="Hello"
hello
// Next, we define the method echo, the method inputs a string and returns the output. In this case, we define the input and output type explicitly.
def echo(message: String):String=message
// We can call the method by providing it an argument:
echo("Hey")
// The method changes if we provide it a new argument:
echo("Hello")
// You can change the value of a method after the object has been created.We can create the class "hey" with the method "SaySomthing":
class hey{def SaySomthing(Somthing:String):String=Somthing}
// We can create a new object and call a method with an argument of 'Hello':
var Hi=new hey()
Hi.SaySomthing("Hello")
//
// If we call the method with different parameters we get a different result.
Hi.SaySomthing("Se Ya")
// You can change the value of a method after the object has been created
// <div class="alert alert-danger alertdanger" style="margin-top: 20px">
// <h1> Question 2.3: </h1>
// <p>
// We have modified the class person and replaced the field age with a method , create an instant of class person and call it bob. Set the parameter to 1990. Call the method of age.
//
// </p>
// </div>
class person( BirthYear:Int){
var year=2017
def age=year-BirthYear
}
var bob=new person(1990)
bob.age
// <div align="right">
// <a href="#q3a" class="btn btn-default" data-toggle="collapse">Click here for the solution</a>
//
// </div>
// <div id="q3a" class="collapse">
// ```
// var Bob=new person(1990)
// Bob.age
// ```
// </div>
// <div class="alert alert-danger alertdanger" style="margin-top: 20px">
// <h1> Question 2.4: </h1>
// <p>
// If we change the field year what will happen to the method age?
// </p>
// </div>
// <div align="right">
// <a href="#q3b" class="btn btn-default" data-toggle="collapse">Click here for the solution</a>
//
// </div>
// <div id="q3b" class="collapse">
// ```
// If we change the field year and call the method age the output will change. Unlike the previous version of "person" age is a now a method. Methods change when ever they are called; fields are only called when the object is created. For example:
//
// Bob.year=1991
// Bob.age
// ```
// </div>
// ## Infix Notaion
// If the method only takes one argument we can use Infix notation we can call a method with no dot or parentheses. For example we can split a string using the split method.
"Infix Notaion".split(" ")
// Equivalently using Infix notation
"Infix Notaion" split " "
// <a id="ref4"> </a>
// ## 2.4 Default and Named Arguments
// ### By the end of this section you should be able to:
//
// <ul>
// <li>Utilize default argument values in Scala class constructors </li>
// <li>Leverage named arguments to only pass certain values </li>
// </ul>
//
//
// We can create classes with default parameters, for example, we can create a method name, which has two input parameters. The first and last, the method concatenates the value of the strings. We can set default values for the parameters to be empty strings .
def Name(first:String="",last:String=""):String=first+" "+last
// We can call the method as follows:
Name("Rob","Roy")
// If we don't include all the parameters we don't get an error
Name("Johon")
// The parameters are called in the same order as they are called in the class constructor. You can be explicitly and reference the name of the parameter in the method call.
Name(last="Barker",first="Bob")
// Similarly, we can only pass one argument using the name of the parameter.
Name(last="Kubrick")
// <div class="alert alert-danger alertdanger" style="margin-top: 20px">
// <h1> Question 2.5: </h1>
// <p>
// Create an object of class name, with the first name "Joei" and Last name "JoeJoe"
// </p>
// </div>
Name("Joei","JoeJoe")
// <div align="right">
// <a href="#q34" class="btn btn-default" data-toggle="collapse">Click here for the solution</a>
//
// </div>
// <div id="q34" class="collapse">
// ```
// Name("Joe","JoeJoe")
//
// Bob.year=1991
// Bob.age
// ```
// <a id="ref5"> </a>
//
// # 2.5 OBJECTS
// ### Singlton object
// ### By the end of this section you should be able to:##
//
// <ul>
// <li> Understand singleton objects in Scala </li>
// <li>Describe the difference between a class and an object </li>
// </ul>
//
// In Scala we can create an object directly using the "object" key word. For example, we can create the object Hello as follows:
object Hello{def message ="Hello!"}
// Unlike a class we do not need to create the object, we can call it directly:
Hello.message
// We can create a class Time,that converts minutes to hours :
// +
class Time(FullHours :Int, PartialHours:Int){println(FullHours+"."+PartialHours+" Hours")}
// -
// We can then create an object Hello that creates instances of the class "Time" using the method "GetHours"
// +
object Hello{
val OneHourInMinutes:Int=60
def GetHours(minutes:Int)=new Time(minutes/OneHourInMinutes, minutes%OneHourInMinutes )
}
// -
// You can apply the method "GetHours" from the object to perform the conversion
Hello.GetHours(64)
Hello.GetHours(50)
// <div class="alert alert-danger alertdanger" style="margin-top: 20px">
// <h1> Question 2.6: </h1>
// <p>
// Create the object "Hello1", the object is identical to "Hello" but call the rename the method "GetHours" to the method "apply"
// </p>
// </div>
object Hello1{ val OneHourInMinutes:Int=60
def apply(minutes:Int)=new Time(minutes/OneHourInMinutes, minutes%OneHourInMinutes )
}
// <div align="right">
// <a href="#q26" class="btn btn-default" data-toggle="collapse">Click here for the solution</a>
//
// </div>
// <div id="q26" class="collapse">
// ```
// object Hello1{
// val OneHourInMinutes:Int=60
//
// def apply(minutes:Int)=new Time(minutes/OneHourInMinutes, minutes%OneHourInMinutes )
//
// }
//
//
// ```
// <div class="alert alert-danger alertdanger" style="margin-top: 20px">
// <h1> Question 2.7: </h1>
// <p>
// Convert 64 minutes to hours using object "Hello1" and the method apply.
// </p>
// </div>
// <div align="right">
// <a href="#q27" class="btn btn-default" data-toggle="collapse">Click here for the solution</a>
//
// </div>
// <div id="q27" class="collapse">
//
// Hello1.apply(64)
//
//
//
// <div class="alert alert-danger alertdanger" style="margin-top: 20px">
// <h1> Question 2.8: </h1>
// <p>
// What happens if you call the object Hello1(64)
// </p>
// </div>
// <div align="right">
// <a href="#q28" class="btn btn-default" data-toggle="collapse">Click here for the solution</a>
//
// </div>
// <div id="q28" class="collapse">
//
// It turns out that apply is a special method, such that hello1.apply(64) equals hello1(64)
//
//
//
//
// <hr>
// Copyright © 2017 [cognitiveclass.ai](cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
// ## About the Authors:
//
//
// [<NAME>]( https://www.linkedin.com/in/joseph-s-50398b136/) has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
//
//
| Scala Programming for Data Science/Scala 101/Module 1: Introduction/Module_2__Basic_Objects_and_Classes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pydev
# language: python
# name: pydev
# ---
#
# ## Install Java, Spark, and Findspark
#
#
# !apt-get install openjdk-8-jdk-headless -qq > /dev/null
# !wget -q http://www-us.apache.org/dist/spark/spark-2.3.2/spark-2.3.2-bin-hadoop2.7.tgz
# !tar xf spark-2.3.2-bin-hadoop2.7.tgz
# !pip install -q findspark
# ## Set Environmental Variables
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = "/content/spark-2.3.2-bin-hadoop2.7"
# ## Find Spark and start session
# +
import findspark
findspark.init()
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName("ins_hash").getOrCreate()
# -
| 07-Extra-Content/Big-Data-Google-Colab/day-2/Activities/06-Ins_Pyspark_NLP_HashingTF/Unsolved/nlp_hashingTF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#from dask.distributed import Client
from dask.distributed import Client, LocalCluster
cluster = LocalCluster()
cluster.scale(4)
client = Client(cluster)
#client = Client('192.168.0.12:8786') # Connect to distributed cluster and override default
#df.x.sum().compute() # This now runs on the distributed system
import dask.dataframe as dd
# %time df = dd.read_csv("large/midyear_population_age_country_code.csv")
#df.x.sum().compute()
#df = dd.read_csv('s3://dask-data/nyc-taxi/2015/*.csv',
# parse_dates=['tpep_pickup_datetime', 'tpep_dropoff_datetime'],
# storage_options={'anon': True})
#dd = client.persist(dd)
df = client.persist(df)
#progress(df)
client
# -
| akash_4-worker-dask-csv-read.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Anne-va/Linear-Algebra-58051/blob/main/Week1/LinAlg_58051_Ebdane_Python_Programming.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="g6SwPoRKiSQ1"
name = input("Enter student name: ")
program = input("Enter student program: ")
prelims = float(input("Enter student prelim grade: "))
midterms = float(input("Enter student midtrem grade: "))
finals = float(input("Enter student final grade: "))
SG = float(0)
prelims = prelims*0.3
midterms = midterms*0.3
finals = finals*0.4
SG = prelims + midterms + finals
happy, lol, sad = ("\U0001F600","\U0001F606","\U0001F62D")
if (SG > 70):
print("SG: {:.2f} ".format(SG), "\U0001F600")
if (SG == 70):
print("SG: {:.2f} ".format(SG), "\U0001F606")
if (SG < 70):
print("SG: {:.2f} ".format(SG), "\U0001F62D")
| Week1/LinAlg_58051_Ebdane_Python_Programming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
gesinputs = pd.read_hdf('/data/jls/GaiaDR2/spectro/GES_input.hdf5')
from astropy.table import Table
ges = Table.read('/data/jls/GaiaDR2/spectro/GES_distances_withPRIOR.hdf5')
def input_output(inputs, data, a, title):
fltr = data['flag']==0
fltr &= data['log10_age']>-1.
plt.sca(a[0])
plt.scatter(np.log10(inputs['teff'][fltr]),inputs['logg'][fltr],
c=inputs['fe_h'][fltr],s=1,cmap=plt.cm.viridis, alpha=0.4, vmin=-2., vmax=0.5);
plt.xlabel(r'$\log_{10}T_\mathrm{eff}$')
plt.ylabel(r'$\log g$')
plt.annotate(title,xy=(0.05,0.95),ha='left',va='top',xycoords='axes fraction',fontsize=16)
plt.colorbar(label='[M/H]')
plt.gca().invert_yaxis()
plt.gca().invert_xaxis()
plt.xlim(4.,3.4)
plt.ylim(5.5,-0.5)
plt.sca(a[1])
plt.scatter(data['log10_teff'][fltr],data['logg'][fltr],c=data['Z'][fltr],s=1,
cmap=plt.cm.viridis, alpha=0.4, vmin=-2., vmax=0.5);
plt.gca().invert_yaxis()
plt.gca().invert_xaxis()
plt.xlabel(r'$\log_{10}T_\mathrm{eff}$')
plt.ylabel(r'$\log g$')
plt.colorbar(label='[M/H]')
plt.xlim(4.,3.4)
plt.ylim(5.5,-0.5)
plt.tight_layout()
f, a=plt.subplots(1,2,figsize=[10.,4.])
input_output(gesinputs[ges['flag']==0], ges[ges['flag']==0], a, 'GES')
def giant(data):
return data['logg']<(3.8-.5*(data['log10_teff']-3.7))
from matplotlib.colors import LogNorm
from plotting_general import running_median
f,a=plt.subplots(1,3,figsize=[14.,3.])
plim=[0.1,97.]
plim2=[5.,95.]
ges['teff']=np.power(10.,ges['log10_teff'])
for i,(t,t2) in enumerate(zip(['teff', 'logg', 'Z'],['teff','logg','fe_h'])):
plt.sca(a[i])
# plt.hexbin(gesinputs[t2],ges[t]-gesinputs[t2],C=gesinputs['e_'+t2],
# vmin=np.nanpercentile(gesinputs['e_'+t2],0.5),vmax=np.nanpercentile(gesinputs['e_'+t2],99.5))
plt.hist2d(gesinputs[t2],ges[t]-gesinputs[t2],
range=[np.nanpercentile(gesinputs[t2],plim),np.nanpercentile(ges[t]-gesinputs[t2],plim2)],
bins=40,norm=LogNorm())
r = running_median(gesinputs[t2],ges[t]-gesinputs[t2],nbins=30,percentiles=plim)
plt.errorbar(r[0],r[1],.5*(r[3]-r[2])/np.sqrt(r[4]))
r = running_median(gesinputs[t2][giant(ges)],(ges[t]-gesinputs[t2])[giant(ges)],
nbins=30,percentiles=plim)
plt.errorbar(r[0],r[1],.5*(r[3]-r[2])/np.sqrt(r[4]))
r = running_median(gesinputs[t2][~giant(ges)],(ges[t]-gesinputs[t2])[~giant(ges)],
nbins=30,percentiles=plim)
plt.errorbar(r[0],r[1],.5*(r[3]-r[2])/np.sqrt(r[4]))
plt.ylim(*np.nanpercentile(ges[t]-gesinputs[t2],plim2))
plt.xlabel(t)
plt.ylabel(r'$\Delta$'+t)
plt.tight_layout()
f,a=plt.subplots(1,3,figsize=[10.,3.])
plt.sca(a[0])
plt.hist(gesinputs['e_teff'],range=[0.,400.],histtype='step',lw=4,bins=30,label='Input')
# plt.hist(gesinputs['teff_val'],range=[0.,400.],histtype='step',lw=4,bins=30)
plt.hist(ges['log10_teff_err']*np.log(10.)*np.power(10.,ges['log10_teff']),
range=[0.,400.],histtype='step',lw=4,bins=30,label='Output');
plt.legend(ncol=2,bbox_to_anchor=(2.6,1.4))
plt.xlabel('teff error / K')
plt.sca(a[1])
plt.hist(gesinputs['e_logg'],range=[0.,0.5],histtype='step',lw=4,bins=30)
plt.hist(ges['logg_err'],range=[0.,0.5],histtype='step',lw=4,bins=30);
plt.xlabel('logg error')
plt.sca(a[2])
plt.hist(gesinputs['e_fe_h'],range=[0.,0.4],histtype='step',lw=4,bins=30)
plt.hist(ges['Z_err'],range=[0.,0.4],histtype='step',lw=4,bins=30);
plt.xlabel('[M/H] error')
plt.tight_layout()
| notebooks/GES.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:eu-west-1:470317259841:image/datascience-1.0
# ---
# # Amazon SageMaker Autopilot Candidate Definition Notebook
#
# This notebook was automatically generated by the AutoML job **automl-banking-17-12-50-55**.
# This notebook allows you to customize the candidate definitions and execute the SageMaker Autopilot workflow.
#
# The dataset has **21** columns and the column named **y** is used as
# the target column. This is being treated as a **BinaryClassification** problem. The dataset also has **2** classes.
# This notebook will build a **[BinaryClassification](https://en.wikipedia.org/wiki/Binary_classification)** model that
# **maximizes** the "**ACCURACY**" quality metric of the trained models.
# The "**ACCURACY**" metric provides the percentage of times the model predicted the correct class.
#
# As part of the AutoML job, the input dataset has been randomly split into two pieces, one for **training** and one for
# **validation**. This notebook helps you inspect and modify the data transformation approaches proposed by Amazon SageMaker Autopilot. You can interactively
# train the data transformation models and use them to transform the data. Finally, you can execute a multiple algorithm hyperparameter optimization (multi-algo HPO)
# job that helps you find the best model for your dataset by jointly optimizing the data transformations and machine learning algorithms.
#
# <div class="alert alert-info"> 💡 <strong> Available Knobs</strong>
# Look for sections like this for recommended settings that you can change.
# </div>
#
#
# ---
#
# ## Contents
#
# 1. [Sagemaker Setup](#Sagemaker-Setup)
# 1. [Downloading Generated Candidates](#Downloading-Generated-Modules)
# 1. [SageMaker Autopilot Job and Amazon Simple Storage Service (Amazon S3) Configuration](#SageMaker-Autopilot-Job-and-Amazon-Simple-Storage-Service-(Amazon-S3)-Configuration)
# 1. [Candidate Pipelines](#Candidate-Pipelines)
# 1. [Generated Candidates](#Generated-Candidates)
# 1. [Selected Candidates](#Selected-Candidates)
# 1. [Executing the Candidate Pipelines](#Executing-the-Candidate-Pipelines)
# 1. [Run Data Transformation Steps](#Run-Data-Transformation-Steps)
# 1. [Multi Algorithm Hyperparameter Tuning](#Multi-Algorithm-Hyperparameter-Tuning)
# 1. [Model Selection and Deployment](#Model-Selection-and-Deployment)
# 1. [Tuning Job Result Overview](#Tuning-Job-Result-Overview)
# 1. [Model Deployment](#Model-Deployment)
#
# ---
# ## Sagemaker Setup
#
# Before you launch the SageMaker Autopilot jobs, we'll setup the environment for Amazon SageMaker
# - Check environment & dependencies.
# - Create a few helper objects/function to organize input/output data and SageMaker sessions.
# **Minimal Environment Requirements**
#
# - Jupyter: Tested on `JupyterLab 1.0.6`, `jupyter_core 4.5.0` and `IPython 6.4.0`
# - Kernel: `conda_python3`
# - Dependencies required
# - `sagemaker-python-sdk>=2.40.0`
# - Use `!pip install sagemaker==2.40.0` to download this dependency.
# - Kernel may need to be restarted after download.
# - Expected Execution Role/permission
# - S3 access to the bucket that stores the notebook.
# ### Downloading Generated Modules
# Download the generated data transformation modules and an SageMaker Autopilot helper module used by this notebook.
# Those artifacts will be downloaded to **automl-banking-17-12-50-55-artifacts** folder.
# +
# !mkdir -p automl-banking-17-12-50-55-artifacts
# !aws s3 sync s3://sagemaker-eu-west-1-071908484098/sagemaker/autopilot-dm/output/automl-banking-17-12-50-55/sagemaker-automl-candidates/automl-banking-17-12-50-55-pr-1-f9acee0f25144a1693ba74fd9b3ce04/generated_module automl-banking-17-12-50-55-artifacts/generated_module --only-show-errors
# !aws s3 sync s3://sagemaker-eu-west-1-071908484098/sagemaker/autopilot-dm/output/automl-banking-17-12-50-55/sagemaker-automl-candidates/automl-banking-17-12-50-55-pr-1-f9acee0f25144a1693ba74fd9b3ce04/notebooks/sagemaker_automl automl-banking-17-12-50-55-artifacts/sagemaker_automl --only-show-errors
import sys
sys.path.append("automl-banking-17-12-50-55-artifacts")
# -
# ### SageMaker Autopilot Job and Amazon Simple Storage Service (Amazon S3) Configuration
#
# The following configuration has been derived from the SageMaker Autopilot job. These items configure where this notebook will
# look for generated candidates, and where input and output data is stored on Amazon S3.
LOCAL_AUTOML_JOB_NAME
# +
from sagemaker_automl import uid, AutoMLLocalRunConfig
# Where the preprocessed data from the existing AutoML job is stored
# 定义 BASE_AUTOML_JOB 的名字和配置
BASE_AUTOML_JOB_NAME = 'automl-banking-17-12-50-55'
BASE_AUTOML_JOB_CONFIG = {
'automl_job_name': BASE_AUTOML_JOB_NAME,
'automl_output_s3_base_path': 's3://sagemaker-eu-west-1-071908484098/sagemaker/autopilot-dm/output/automl-banking-17-12-50-55',
'data_transformer_image_repo_version': '2.5-1-cpu-py3',
'algo_image_repo_versions': {'xgboost': '1.3-1-cpu-py3', 'linear-learner': 'training-cpu'},
'algo_inference_image_repo_versions': {'xgboost': '1.3-1-cpu-py3', 'linear-learner': 'inference-cpu'}
}
# Path conventions of the output data storage path from the local AutoML job run of this notebook
# 定义 LOCAL_AUTOML_JOB的名字和配置
LOCAL_AUTOML_JOB_NAME = 'automl-ban-notebook-run-{}'.format(uid())
LOCAL_AUTOML_JOB_CONFIG = {
'local_automl_job_name': LOCAL_AUTOML_JOB_NAME,
'local_automl_job_output_s3_base_path': 's3://sagemaker-eu-west-1-071908484098/sagemaker/autopilot-dm/output/automl-banking-17-12-50-55/{}'.format(LOCAL_AUTOML_JOB_NAME),
'data_processing_model_dir': 'data-processor-models',
'data_processing_transformed_output_dir': 'transformed-data',
'multi_algo_tuning_output_dir': 'multi-algo-tuning'
}
# 定义 AUTOML_LOCAL_RUN 配置
AUTOML_LOCAL_RUN_CONFIG = AutoMLLocalRunConfig(
role='arn:aws:iam::071908484098:role/service-role/AmazonSageMaker-ExecutionRole-20220308T154036',
base_automl_job_config=BASE_AUTOML_JOB_CONFIG,
local_automl_job_config=LOCAL_AUTOML_JOB_CONFIG,
security_config={'EnableInterContainerTrafficEncryption': False, 'VpcConfig': {}})
AUTOML_LOCAL_RUN_CONFIG.display()
# -
# ## Candidate Pipelines
#
# The `AutoMLLocalRunner` keeps track of selected candidates and automates many of the steps needed to execute feature engineering and tuning steps.
# +
from sagemaker_automl import AutoMLInteractiveRunner, AutoMLLocalCandidate
# 创建一个 AutoMLInteractiveRunner 实例
automl_interactive_runner = AutoMLInteractiveRunner(AUTOML_LOCAL_RUN_CONFIG)
# -
# ### Generated Candidates
#
# The SageMaker Autopilot Job has analyzed the dataset and has generated **5** machine learning
# pipeline(s) that use **2** algorithm(s). Each pipeline contains a set of feature transformers and an
# algorithm.
#
# <div class="alert alert-info"> 💡 <strong> Available Knobs</strong>
#
# 1. The resource configuration: instance type & count
# 1. Select candidate pipeline definitions by cells
# 1. The linked data transformation script can be reviewed and updated. Please refer to the [README.md](./automl-banking-17-12-50-55-artifacts/generated_module/README.md) for detailed customization instructions.
# </div>
# **[dpp0-xgboost](automl-banking-17-12-50-55-artifacts/generated_module/candidate_data_processors/dpp0.py)**: This data transformation strategy first transforms 'numeric' features using [RobustImputer (converts missing values to nan)](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/impute/base.py), 'categorical' features using [ThresholdOneHotEncoder](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/encoders.py), 'datetime' features using [DateTimeVectorizer](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/feature_extraction/date_time.py). It merges all the generated features and applies [RobustStandardScaler](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/data.py). The
# transformed data will be used to tune a *xgboost* model. Here is the definition:
# 设置automl的candidate, 设置数据和算法的配置
automl_interactive_runner.select_candidate({
"data_transformer": {
"name": "dpp0",
"training_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
"volume_size_in_gb": 50
},
"transform_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
},
"transforms_label": True,
"transformed_data_format": "text/csv",
"sparse_encoding": False
},
"algorithm": {
"name": "xgboost",
"training_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
},
}
})
# **[dpp1-xgboost](automl-banking-17-12-50-55-artifacts/generated_module/candidate_data_processors/dpp1.py)**: This data transformation strategy first transforms 'numeric' features using [RobustImputer](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/impute/base.py), 'categorical' features using [ThresholdOneHotEncoder](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/encoders.py), 'datetime' features using [DateTimeVectorizer](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/feature_extraction/date_time.py). It merges all the generated features and applies [RobustPCA](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/decomposition/robust_pca.py) followed by [RobustStandardScaler](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/data.py). The
# transformed data will be used to tune a *xgboost* model. Here is the definition:
# 设置automl的candidate, 设置数据和算法的配置
automl_interactive_runner.select_candidate({
"data_transformer": {
"name": "dpp1",
"training_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
"volume_size_in_gb": 50
},
"transform_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
},
"transforms_label": True,
"transformed_data_format": "text/csv",
"sparse_encoding": False
},
"algorithm": {
"name": "xgboost",
"training_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
},
}
})
# **[dpp2-linear-learner](automl-banking-17-12-50-55-artifacts/generated_module/candidate_data_processors/dpp2.py)**: This data transformation strategy first transforms 'numeric' features using [combined RobustImputer and RobustMissingIndicator](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/impute/base.py) followed by [QuantileExtremeValuesTransformer](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/base.py), 'categorical' features using [ThresholdOneHotEncoder](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/encoders.py), 'datetime' features using [DateTimeVectorizer](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/feature_extraction/date_time.py) followed by [RobustImputer](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/impute/base.py). It merges all the generated features and applies [RobustPCA](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/decomposition/robust_pca.py) followed by [RobustStandardScaler](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/data.py). The
# transformed data will be used to tune a *linear-learner* model. Here is the definition:
# 设置automl的candidate, 设置数据和算法的配置
automl_interactive_runner.select_candidate({
"data_transformer": {
"name": "dpp2",
"training_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
"volume_size_in_gb": 50
},
"transform_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
},
"transforms_label": True,
"transformed_data_format": "application/x-recordio-protobuf",
"sparse_encoding": False
},
"algorithm": {
"name": "linear-learner",
"training_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
},
}
})
# **[dpp3-xgboost](automl-banking-17-12-50-55-artifacts/generated_module/candidate_data_processors/dpp3.py)**: This data transformation strategy first transforms 'numeric' features using [RobustImputer (converts missing values to nan)](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/impute/base.py), 'categorical' features using [ThresholdOneHotEncoder](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/encoders.py), 'datetime' features using [DateTimeVectorizer](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/feature_extraction/date_time.py). It merges all the generated features and applies [RobustStandardScaler](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/data.py). The
# transformed data will be used to tune a *xgboost* model. Here is the definition:
# 设置automl的candidate, 设置数据和算法的配置
automl_interactive_runner.select_candidate({
"data_transformer": {
"name": "dpp3",
"training_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
"volume_size_in_gb": 50
},
"transform_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
},
"transforms_label": True,
"transformed_data_format": "text/csv",
"sparse_encoding": False
},
"algorithm": {
"name": "xgboost",
"training_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
},
}
})
# **[dpp4-xgboost](automl-banking-17-12-50-55-artifacts/generated_module/candidate_data_processors/dpp4.py)**: This data transformation strategy first transforms 'numeric' features using [RobustImputer (converts missing values to nan)](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/impute/base.py), 'categorical' features using [ThresholdOneHotEncoder](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/encoders.py), 'datetime' features using [DateTimeVectorizer](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/feature_extraction/date_time.py). It merges all the generated features and applies [RobustStandardScaler](https://github.com/aws/sagemaker-scikit-learn-extension/blob/master/src/sagemaker_sklearn_extension/preprocessing/data.py). The
# transformed data will be used to tune a *xgboost* model. Here is the definition:
# 设置automl的candidate, 设置数据和算法的配置
automl_interactive_runner.select_candidate({
"data_transformer": {
"name": "dpp4",
"training_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
"volume_size_in_gb": 50
},
"transform_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
},
"transforms_label": True,
"transformed_data_format": "application/x-recordio-protobuf",
"sparse_encoding": True
},
"algorithm": {
"name": "xgboost",
"training_resource_config": {
"instance_type": "ml.m5.4xlarge",
"instance_count": 1,
},
}
})
# ### Selected Candidates
#
# You have selected the following candidates (please run the cell below and click on the feature transformer links for details):
automl_interactive_runner.display_candidates()
# The feature engineering pipeline consists of two SageMaker jobs:
#
# 1. Generated trainable data transformer Python modules like [dpp0.py](automl-banking-17-12-50-55-artifacts/generated_module/candidate_data_processors/dpp0.py), which has been downloaded to the local file system
# 2. A **training** job to train the data transformers
# 3. A **batch transform** job to apply the trained transformation to the dataset to generate the algorithm compatible data
#
# The transformers and its training pipeline are built using open sourced **[sagemaker-scikit-learn-container][]** and **[sagemaker-scikit-learn-extension][]**.
#
# [sagemaker-scikit-learn-container]: https://github.com/aws/sagemaker-scikit-learn-container
# [sagemaker-scikit-learn-extension]: https://github.com/aws/sagemaker-scikit-learn-extension
# ## Executing the Candidate Pipelines
#
# Each candidate pipeline consists of two steps, feature transformation and algorithm training.
# For efficiency first execute the feature transformation step which will generate a featurized dataset on S3
# for each pipeline.
#
# After each featurized dataset is prepared, execute a multi-algorithm tuning job that will run tuning jobs
# in parallel for each pipeline. This tuning job will execute training jobs to find the best set of
# hyper-parameters for each pipeline, as well as finding the overall best performing pipeline.
#
# ### Run Data Transformation Steps
#
# Now you are ready to start execution all data transformation steps. The cell below may take some time to finish,
# feel free to go grab a cup of coffee. To expedite the process you can set the number of `parallel_jobs` to be up to 10.
# Please check the account limits to increase the limits before increasing the number of jobs to run in parallel.
# 运行训练, parallel_jobs=10
automl_interactive_runner.fit_data_transformers(parallel_jobs=10)
# ### Multi Algorithm Hyperparameter Tuning
#
# Now that the algorithm compatible transformed datasets are ready, you can start the multi-algorithm model tuning job
# to find the best predictive model. The following algorithm training job configuration for each
# algorithm is auto-generated by the AutoML Job as part of the recommendation.
#
# <div class="alert alert-info"> 💡 <strong> Available Knobs</strong>
#
# 1. Hyperparameter ranges
# 2. Objective metrics
# 3. Recommended static algorithm hyperparameters.
#
# Please refers to [Xgboost tuning](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost-tuning.html) and [Linear learner tuning](https://docs.aws.amazon.com/sagemaker/latest/dg/linear-learner-tuning.html) for detailed explanations of the parameters.
# </div>
# The AutoML recommendation job has recommended the following hyperparameters, objectives and accuracy metrics for
# the algorithm and problem type:
# +
# 定义 METRICS
ALGORITHM_OBJECTIVE_METRICS = {
'xgboost': 'validation:accuracy',
'linear-learner': 'validation:binary_classification_accuracy',
}
# 设置超参
# xgboost: 超参
# objective:
# eval_metric: 评价函数
# _kfold:
# _num_cv_round:
# linear-learner 超参: https://docs.aws.amazon.com/sagemaker/latest/dg/ll_hyperparameters.html
# predictor_type: 目标变量的类型
# ml_application:
# loss_function: 损失函数
# reporting_metrics: 监控指标
# eval_metric: 评价函数
# kfold:
# num_cv_rounds:
# prediction_storage_mode:
STATIC_HYPERPARAMETERS = {
'xgboost': {
'objective': 'binary:logistic',
'eval_metric': 'accuracy,f1_binary,auc',
'_kfold': 5,
'_num_cv_round': 3,
},
'linear-learner': {
'predictor_type': 'binary_classifier',
'ml_application': 'linear_learner',
'loss_function': 'SoftmaxCrossEntropyLoss',
'reporting_metrics': 'binary_classification_accuracy,binary_f_beta,roc_auc_score',
'eval_metric': 'binary_classification_accuracy',
'kfold': 5,
'num_cv_rounds': 3,
'prediction_storage_mode': 'store_cv_avg_predictions',
},
}
# -
# The following tunable hyperparameters search ranges are recommended for the Multi-Algo tuning job:
# +
from sagemaker.parameter import CategoricalParameter, ContinuousParameter, IntegerParameter
# 设置模型超参: 参考: https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost_hyperparameters.html
# num_round: 迭代次数
# max_depth 最大深度,
# eta 学习率,
# gamma 分裂最小loss Gamma指定了节点分裂所需的最小损失函数下降值, 这个参数的值越大,算法越保守
# min_child_weight 决定最小叶子节点样本权重和 参数用于避免过拟合。当它的值较大时,可以避免模型学习到局部的特殊样本。 但是如果这个值过高,会导致欠拟合,
# subsample: 这个参数控制对于每棵树,随机采样的比例。 减小这个参数的值,算法会更加保守,避免过拟合。但是,如果这个值设置得过小,它可能会导致欠拟合.
# colsample_bytree: 构建每棵树时列的子样本比率。
# lambda: 权重的 L2 正则化项。增加这个值会使模型更加保守。
# alpha: 权重的 L1 正则化项。增加这个值会使模型更加保守
# linear-learner 超参
# mini_batch_size: 最一批次的数据大小
# wd: 权重衰减参数,也称为L2正则化参数。
# learning_rate: 学习率
ALGORITHM_TUNABLE_HYPERPARAMETER_RANGES = {
'xgboost': {
'num_round': IntegerParameter(64, 1024, scaling_type='Logarithmic'),
'max_depth': IntegerParameter(2, 8, scaling_type='Logarithmic'),
'eta': ContinuousParameter(1e-3, 1.0, scaling_type='Logarithmic'),
'gamma': ContinuousParameter(1e-6, 64.0, scaling_type='Logarithmic'),
'min_child_weight': ContinuousParameter(1e-6, 32.0, scaling_type='Logarithmic'),
'subsample': ContinuousParameter(0.5, 1.0, scaling_type='Linear'),
'colsample_bytree': ContinuousParameter(0.3, 1.0, scaling_type='Linear'),
'lambda': ContinuousParameter(1e-6, 2.0, scaling_type='Logarithmic'),
'alpha': ContinuousParameter(1e-6, 2.0, scaling_type='Logarithmic'),
},
'linear-learner': {
'mini_batch_size': IntegerParameter(128, 512, scaling_type='Linear'),
'wd': ContinuousParameter(1e-12, 1e-2, scaling_type='Logarithmic'),
'learning_rate': ContinuousParameter(1e-6, 1e-2, scaling_type='Logarithmic'),
},
}
# -
# #### Prepare Multi-Algorithm Tuner Input
#
# To use the multi-algorithm HPO tuner, prepare some inputs and parameters. Prepare a dictionary whose key is the name of the trained pipeline candidates and the values are respectively:
#
# 1. Estimators for the recommended algorithm
# 2. Hyperparameters search ranges
# 3. Objective metrics
# 设置多个算法微调参数
multi_algo_tuning_parameters = automl_interactive_runner.prepare_multi_algo_parameters(
objective_metrics=ALGORITHM_OBJECTIVE_METRICS,
static_hyperparameters=STATIC_HYPERPARAMETERS,
hyperparameters_search_ranges=ALGORITHM_TUNABLE_HYPERPARAMETER_RANGES)
# Below you prepare the inputs data to the multi-algo tuner:
# 准备inputs数据
multi_algo_tuning_inputs = automl_interactive_runner.prepare_multi_algo_inputs()
# #### Create Multi-Algorithm Tuner
#
# With the recommended Hyperparameter ranges and the transformed dataset, create a multi-algorithm model tuning job
# that coordinates hyper parameter optimizations across the different possible algorithms and feature processing strategies.
#
# <div class="alert alert-info"> 💡 <strong> Available Knobs</strong>
#
# 1. Tuner strategy: [Bayesian](https://en.wikipedia.org/wiki/Hyperparameter_optimization#Bayesian_optimization), [Random Search](https://en.wikipedia.org/wiki/Hyperparameter_optimization#Random_search)
# 2. Objective type: `Minimize`, `Maximize`, see [optimization](https://en.wikipedia.org/wiki/Mathematical_optimization)
# 3. Max Job size: the max number of training jobs HPO would be launching to run experiments. Note the default value is **250**
# which is the default of the managed flow.
# 4. Parallelism. Number of jobs that will be executed in parallel. Higher value will expedite the tuning process.
# Please check the account limits to increase the limits before increasing the number of jobs to run in parallel
# 5. Please use a different tuning job name if you re-run this cell after applied customizations.
# </div>
# +
from sagemaker.tuner import HyperparameterTuner
base_tuning_job_name = "{}-tuning".format(AUTOML_LOCAL_RUN_CONFIG.local_automl_job_name)
# 创建一个 tuner 超参
tuner = HyperparameterTuner.create(
base_tuning_job_name=base_tuning_job_name,
strategy='Bayesian',
objective_type='Maximize',
max_parallel_jobs=7,
max_jobs=250,
**multi_algo_tuning_parameters,
)
# -
# #### Run Multi-Algorithm Tuning
#
# Now you are ready to start running the **Multi-Algo Tuning** job. After the job is finished, store the tuning job name which you use to select models in the next section.
# The tuning process will take some time, please track the progress in the Amazon SageMaker Hyperparameter tuning jobs console.
# +
from IPython.display import display, Markdown
# Run tuning
tuner.fit(inputs=multi_algo_tuning_inputs, include_cls_metadata=None)
tuning_job_name = tuner.latest_tuning_job.name
display(
Markdown(f"Tuning Job {tuning_job_name} started, please track the progress from [here](https://{AUTOML_LOCAL_RUN_CONFIG.region}.console.aws.amazon.com/sagemaker/home?region={AUTOML_LOCAL_RUN_CONFIG.region}#/hyper-tuning-jobs/{tuning_job_name})"))
# Wait for tuning job to finish
tuner.wait()
# -
# ## Model Selection and Deployment
#
# This section guides you through the model selection process. Afterward, you construct an inference pipeline
# on Amazon SageMaker to host the best candidate.
#
# Because you executed the feature transformation and algorithm training in two separate steps, you now need to manually
# link each trained model with the feature transformer that it is associated with. When running a regular Amazon
# SageMaker Autopilot job, this will automatically be done for you.
# ### Tuning Job Result Overview
#
# The performance of each candidate pipeline can be viewed as a Pandas dataframe. For more interactive usage please
# refers to [model tuning monitor](https://docs.aws.amazon.com/sagemaker/latest/dg/automatic-model-tuning-monitor.html).
# +
from pprint import pprint
from sagemaker.analytics import HyperparameterTuningJobAnalytics
SAGEMAKER_SESSION = AUTOML_LOCAL_RUN_CONFIG.sagemaker_session
SAGEMAKER_ROLE = AUTOML_LOCAL_RUN_CONFIG.role
# 设置TuningJobAnalytics的超参
tuner_analytics = HyperparameterTuningJobAnalytics(
tuner.latest_tuning_job.name, sagemaker_session=SAGEMAKER_SESSION)
# 获取 dataframe
df_tuning_job_analytics = tuner_analytics.dataframe()
# Sort the tuning job analytics by the final metrics value
# 通过 FinalObjectiveValue对数据进行排序
df_tuning_job_analytics.sort_values(
by=['FinalObjectiveValue'],
inplace=True,
ascending=False if tuner.objective_type == "Maximize" else True)
# Show detailed analytics for the top 20 models
# 显示分析的前20条数据
df_tuning_job_analytics.head(20)
# -
# The best training job can be selected as below:
#
# <div class="alert alert-info"> 💡 <strong>Tips: </strong>
# You could select alternative job by using the value from `TrainingJobName` column above and assign to `best_training_job` below
# </div>
# +
attached_tuner = HyperparameterTuner.attach(tuner.latest_tuning_job.name, sagemaker_session=SAGEMAKER_SESSION)
best_training_job = attached_tuner.best_training_job()
print("Best Multi Algorithm HPO training job name is {}".format(best_training_job))
# -
# ### Linking Best Training Job with Feature Pipelines
#
# Finally, deploy the best training job to Amazon SageMaker along with its companion feature engineering models.
# At the end of the section, you get an endpoint that's ready to serve online inference or start batch transform jobs!
# Deploy a [PipelineModel](https://sagemaker.readthedocs.io/en/stable/pipeline.html) that has multiple containers of the following:
#
# 1. Data Transformation Container: a container built from the model we selected and trained during the data transformer sections
# 2. Algorithm Container: a container built from the trained model we selected above from the best HPO training job.
# 3. Inverse Label Transformer Container: a container that converts numerical intermediate prediction value back to non-numerical label value.
#
# Get both best data transformation model and algorithm model from best training job and create an pipeline model:
# +
from sagemaker.estimator import Estimator
from sagemaker import PipelineModel
from sagemaker_automl import select_inference_output
# Get a data transformation model from chosen candidate
best_candidate = automl_interactive_runner.choose_candidate(df_tuning_job_analytics, best_training_job)
best_data_transformer_model = best_candidate.get_data_transformer_model(role=SAGEMAKER_ROLE, sagemaker_session=SAGEMAKER_SESSION)
# Our first data transformation container will always return recordio-protobuf format
# 在 protobuf recordIO 格式中,SageMaker 将数据集中的每个观察转换为二进制表示形式(一组 4 字节浮点数),然后将其加载到 protobuf 值字段
best_data_transformer_model.env["SAGEMAKER_DEFAULT_INVOCATIONS_ACCEPT"] = 'application/x-recordio-protobuf'
# Add environment variable for sparse encoding
if best_candidate.data_transformer_step.sparse_encoding:
best_data_transformer_model.env["AUTOML_SPARSE_ENCODE_RECORDIO_PROTOBUF"] = '1'
# Get a algo model from chosen training job of the candidate
algo_estimator = Estimator.attach(best_training_job)
best_algo_model = algo_estimator.create_model(**best_candidate.algo_step.get_inference_container_config())
# Final pipeline model is composed of data transformation models and algo model and an
# inverse label transform model if we need to transform the intermediates back to non-numerical value
model_containers = [best_data_transformer_model, best_algo_model]
if best_candidate.transforms_label:
model_containers.append(best_candidate.get_data_transformer_model(
transform_mode="inverse-label-transform",
role=SAGEMAKER_ROLE,
sagemaker_session=SAGEMAKER_SESSION))
# This model can emit response ['predicted_label', 'probability', 'labels', 'probabilities']. To enable the model to emit one or more
# of the response content, pass the keys to `output_key` keyword argument in the select_inference_output method.
model_containers = select_inference_output("BinaryClassification", model_containers, output_keys=['predicted_label'])
# 创建一个PipelineModel模型
pipeline_model = PipelineModel(
name="AutoML-{}".format(AUTOML_LOCAL_RUN_CONFIG.local_automl_job_name),
role=SAGEMAKER_ROLE,
models=model_containers,
vpc_config=AUTOML_LOCAL_RUN_CONFIG.vpc_config)
# -
# ### Deploying Best Pipeline
#
# <div class="alert alert-info"> 💡 <strong> Available Knobs</strong>
#
# 1. You can customize the initial instance count and instance type used to deploy this model.
# 2. Endpoint name can be changed to avoid conflict with existing endpoints.
#
# </div>
#
# Finally, deploy the model to SageMaker to make it functional.
# 部署模型
pipeline_model.deploy(initial_instance_count=1,
instance_type='ml.m5.2xlarge',
endpoint_name=pipeline_model.name,
wait=True)
# Congratulations! Now you could visit the sagemaker
# [endpoint console page](https://eu-west-1.console.aws.amazon.com/sagemaker/home?region=eu-west-1#/endpoints) to find the deployed endpoint (it'll take a few minutes to be in service).
#
# <div class="alert alert-warning">
# <strong>To rerun this notebook, delete or change the name of your endpoint!</strong> <br>
# If you rerun this notebook, you'll run into an error on the last step because the endpoint already exists. You can either delete the endpoint from the endpoint console page or you can change the <code>endpoint_name</code> in the previous code block.
# </div>
| lab4/AutopilotCandidateDefinitionNotebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
from ipywidgets import interactive
import matplotlib
def flat_fun(x):
output = 1.6*np.ones_like(x)
output[x<-0.8] = x[x<-0.8]+2.4
output[x>4] = x[x>4]-2.4
return output
def pt1(x,x_0,k,y_0,L):
return ((x-x_0)+(L/2))+y_0
def pt2(x,x_0,k,L):
return L/(np.exp((-k)*(x-x_0)) + 1.)
# return np.tanh(k*(x-x_0)/((1.-(x-x_0)**2)))
def test(x,x_0,k,L):
return (-k)*(x-x_0)**3
def sigmoid(x,x_0,k,y_0,L):
return pt1(x,x_0,k,y_0,L)-pt2(x,x_0,k,L)
def f(x_0a=1.6,x_0b=1.6,L=0.8,k=4.,y_0=1.6):
yl,yu = -4,8
# yl,yu = -2,2
inputs = np.linspace(-5,8,100)
#inputs = np.linspace(-2,2,100)
fig = plt.figure(figsize=(12,8))
gs = matplotlib.gridspec.GridSpec(2,2)
ax = fig.add_subplot(gs[0,:])
ax.plot(inputs,flat_fun(inputs))
ax.plot(inputs,pt1(inputs,x_0a,k,y_0,L))
ax.set_ylim([yl,yu])
ax = fig.add_subplot(gs[1,0])
ax.plot(inputs,sigmoid(inputs,x_0a,k,y_0,L))
ax.plot(inputs,pt1(inputs,x_0a,k,y_0,L))
ax.plot(inputs,pt2(inputs,x_0a,k,L))
# ax.plot(inputs,test(inputs,x_0,k,L))
ax.set_ylim([-4,8])
ax = fig.add_subplot(gs[1,1])
ax.plot(inputs,flat_fun(inputs))
ax.plot(inputs,sigmoid(inputs,x_0a,k,y_0,L))
ax.plot(inputs,sigmoid(inputs,x_0b,k,y_0,L))
ax.set_ylim([yl,yu])
interactive_plot = interactive(f, x_0a =(-2,6,0.1), x_0b =(-2,6,0.1),L=(0.5,5,0.1),k=(.1,5,0.01),y_0=(0,4,0.1))
output = interactive_plot.children[-1]
output.layout.height = '1000px'
interactive_plot
| .ipynb_checkpoints/speed_sigmoid-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pymaceuticals Inc.
# ---
#
# ### Analysis
# * Overall, it is clear that Capomulin is a viable drug regimen to reduce tumor growth.
# * Capomulin had the most number of mice complete the study, with the exception of Remicane, all other regimens observed a number of mice deaths across the duration of the study.
# * There is a strong correlation between mouse weight and tumor volume, indicating that mouse weight may be contributing to the effectiveness of any drug regimen.
# * There was one potential outlier within the Infubinol regimen. While most mice showed tumor volume increase, there was one mouse that had a reduction in tumor growth in the study.
# !pip install pandas
# !pip install matplotlib
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
# Study data files
mouse_metadata_path = "Mouse_metadata.csv"
study_results_path = "Study_results.csv"
# Read the mouse data and the study results
mouse_metadata_df = pd.read_csv(mouse_metadata_path)
study_results_df = pd.read_csv(study_results_path)
# Combine the data into a single dataset.
#study_results = study_results.dropna(axis=1)
#merge_mouse_study = mouse_methdata.merge(study_results, on = 'Mouse ID')
merge_mouse_study_df = pd.merge(mouse_metadata_df, study_results_df, on="Mouse ID")
merge_mouse_study_df.head()
# Display the data table for preview
#merge_mouse_study_df.to_csv("output.csv", index=False)
#merge_mouse_study_df.to_csv.head()
# +
# Checking the number of mice.
print (f'Number of mice:', merge_mouse_study_df['Mouse ID'].nunique())
# -
# Getting the duplicate mice by ID number that shows up for Mouse ID and Timepoint.
data_duplicate_mouse = merge_mouse_study_df.loc[merge_mouse_study_df.duplicated(subset=['Mouse ID', 'Timepoint']), 'Mouse ID'].unique()
data_duplicate_mouse
# Optional: Get all the data for the duplicate mouse ID.
dat_dup_mouse = merge_mouse_study_df.loc[merge_mouse_study_df['Mouse ID'] == "g989", :]
dat_dup_mouse
# Create a clean DataFrame by dropping the duplicate mouse by its ID.
clean_mouseDF = merge_mouse_study_df[merge_mouse_study_df['Mouse ID'].isin(dat_dup_mouse) == False]
clean_mouseDF.head()
# Checking the number of mice in the clean DataFrame.
print (f'Number of mice:', clean_mouseDF['Mouse ID'].nunique())
# ## Summary Statistics
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Use groupby and summary statistical methods to calculate the following properties of each drug regimen:
# mean, median, variance, standard deviation, and SEM of the tumor volume.
# Assemble the resulting series into a single summary dataframe.
#data_1 = merge_mouse_study_df
mean_data=merge_mouse_study_df['Tumor Volume (mm3)'].groupby(merge_mouse_study_df['Drug Regimen']).mean()
median_data=merge_mouse_study_df['Tumor Volume (mm3)'].groupby(merge_mouse_study_df['Drug Regimen']).median()
var_data=merge_mouse_study_df['Tumor Volume (mm3)'].groupby(merge_mouse_study_df['Drug Regimen']).var()
std_data=merge_mouse_study_df['Tumor Volume (mm3)'].groupby(merge_mouse_study_df['Drug Regimen']).std()
sem_data=merge_mouse_study_df['Tumor Volume (mm3)'].groupby(merge_mouse_study_df['Drug Regimen']).sem()
#Create the table:
drugregimen_table = pd.DataFrame ({"Mean Tumor Volume": mean_data,
"Median Tumor Volume": median_data,
"Tumor Volume Variance": var_data,
"Tumor Volume Std. Dev.": std_data,
"Tumor Volume Std. Err.": sem_data})
drugregimen_table
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Using the aggregation method, produce the same summary statistics in a single line.
# -
# ## Bar and Pie Charts
# +
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pandas.
drug_groups = merge_mouse_study_df.groupby('Drug Regimen')
measurments_ = drug_groups['Mouse ID'].count()
measurments_
#or
#merge_mouse_study_df['Mouse ID'].value_counts()
measurments_ = measurments_.drop(measurments_.index[3])
measurments_chart = measurments_.plot(kind="bar", title="Measurments of Drug Regimens")
measurments_chart.set_xlabel("Drug Regimen")
measurments_chart.set_ylabel("Total Number of Measurments")
plt.show()
plt.tight_layout()
# -
# +
# Generate a bar plot showing the total number of measurements taken on each drug regimen using using pyplot.
drug_groups = merge_mouse_study_df.groupby('Drug Regimen')
measurments_ = drug_groups['Mouse ID'].count()
measurments_
#Data Set
x_axis = measurments_
y_axis = drug_groups
#plt.bar=(drug_groups, measurments_)
plt.title = ('Measurments of Drug Regimens')
plt.xlabel = ('Drug Regimen')
plt.ylabel = ('Total Number of Measurments')
plt.xlim(-0.75, len(drug_groups)- 1)
plt.ylim(0, max(measurments_) + 1)
plt.bar(x_axis, measurments_, facecolor="red", alpha = 0.75, align = "center")
#plt.xticks(tick_locations, drug_groups)
plt.show()
# +
# Generate a pie plot showing the distribution of female versus male mice using pandas.
sex = ['Male', 'Female']
sex_count = merge_mouse_study_df['Sex'].value_counts()
pie_chart = sex_count.plot(kind="pie")
mf_pie = sex_count.plot(kind="pie", y='Sex', title=("Female and Male Measurments"))
plt.ylabel("Sex")
plt.show()
# +
# Generate a pie plot showing the distribution of female versus male mice using pyplot
sex = ['Male', 'Female']
sex_count = [51, 49]
fig = plt.figure
plt.pie(sex_count, labels = sex)
plt.ylabel("Sex")
plt.show()
# -
# ## Quartiles, Outliers and Boxplots
# +
# Calculate the final tumor volume of each mouse across four of the treatment regimens:
# Capomulin, Ramicane, Infubinol, and Ceftamin
treatments = ["Capomulin", "Ramicane", "Infubinol", "Ceftamin"]
#data['Timepoint'] = merge_mouse_study_df['Timepoint'].astype(float)
#data = merge_mouse_study_df['Timepoint'].astype(float)
capomulin_data = merge_mouse_study_df.loc[merge_mouse_study_df['Drug Regimen'] =='Capomulin',:]
ramicane_data = merge_mouse_study_df.loc[merge_mouse_study_df['Drug Regimen'] =='Ramicane',:]
infubinol_data = merge_mouse_study_df.loc[merge_mouse_study_df['Drug Regimen'] =='Infubinol',:]
ceftamin_data = merge_mouse_study_df.loc[merge_mouse_study_df['Drug Regimen'] =='Ceftamin',:]
# Start by getting the last (greatest) timepoint for each mouse.
greatest_capomulin_data = capomulin_data.groupby('Mouse ID').max()['Timepoint']
greatest_ramicane_data = ramicane_data.groupby('Mouse ID').max()['Timepoint']
greatest_infubinol_data = infubinol_data.groupby('Mouse ID').max()['Timepoint']
greatest_ceftamin_data = ceftamin_data.groupby('Mouse ID').max()['Timepoint']
# Merge this group df with the original dataframe to get the tumor volume at the last timepoint
capomulin_df = pd.DataFrame(greatest_capomulin_data)
ramicane_df = pd.DataFrame(greatest_ramicane_data)
infubinol_df = pd.DataFrame(greatest_infubinol_data)
ceftamin_df = pd.DataFrame(greatest_ceftamin_data)
capomulin_merge = pd.merge(capomulin_df, merge_mouse_study_df, on = ("Mouse ID", "Timepoint"), how = "left")
ramicane_merge = pd.merge(ramicane_df, merge_mouse_study_df, on = ("Mouse ID", "Timepoint"), how = "left")
infubinol_merge = pd.merge(infubinol_df, merge_mouse_study_df, on = ("Mouse ID", "Timepoint"), how = "left")
ceftamin_merge = pd.merge(ceftamin_df, merge_mouse_study_df, on = ("Mouse ID", "Timepoint"), how = "left")
# +
# Put treatments into a list for for loop (and later for plot labels)
treatments = ['Capomulin', 'Ramicane', 'Infubinol', 'Ceftamin']
# Create empty list to fill with tumor vol data (for plotting)
tumor_vol_data = [vol for vol in merge_mouse_study_df['Tumor Volume (mm3)']]
# Calculate the IQR and quantitatively determine if there are any potential outliers.
capomulin_tumors = capomulin_merge['Tumor Volume (mm3)']
ramicane_tumors = ramicane_merge['Tumor Volume (mm3)']
infubinol_tumors = infubinol_merge['Tumor Volume (mm3)']
ceframin_tumors = ceftamin_merge['Tumor Volume (mm3)']
# Locate the rows which contain mice on each drug and get the tumor volumes
# add subset
# Determine outliers using upper and lower bounds
#C = CAPOMULIN
c_quartiles = capomulin_tumors.quantile([.25, .5, .75])
c_lowerqt = c_quartiles[.25]
c_upperqt = c_quartiles[.75]
c_iqr = c_upperqt-c_lowerqt
c_lower_bound = c_lowerqt - (1.5*c_iqr)
c_upper_bound = c_upperqt + (1.5*c_iqr)
#R = RAMICANE
r_quartiles = ramicane_tumors.quantile([.25, .5, .75])
r_lowerqt = r_quartiles[.25]
r_upperqt = r_quartiles[.75]
r_iqr = r_upperqt-r_lowerqt
r_lower_bound = r_lowerqt - (1.5*r_iqr)
r_upper_bound = r_upperqt + (1.5*r_iqr)
#i = INFUBINOL
i_quartiles = infubinol_tumors.quantile([.25, .5, .75])
i_lowerqt = i_quartiles[.25]
i_upperqt = i_quartiles[.75]
i_iqr = i_upperqt-i_lowerqt
i_lower_bound = i_lowerqt - (1.5*i_iqr)
i_upper_bound = i_upperqt + (1.5*i_iqr)
#CE-CEFRAMIN
ce_quartiles = ceframin_tumors.quantile([.25, .5, .75])
ce_lowerqt = ce_quartiles[.25]
ce_upperqt = ce_quartiles[.75]
ce_iqr = ce_upperqt-ce_lowerqt
ce_lower_bound = ce_lowerqt - (1.5*ce_iqr)
ce_upper_bound = ce_upperqt + (1.5*ce_iqr)
print(f"Capomulin")
print(f"Lower quartile of Capomulin tumors = {c_lowerqt}")
print(f"Upper quartile of Capomulin tumors = {c_upperqt}")
print(f"Interquartile range of Capomulin tumors = {c_iqr}")
print(f"Values below {c_lower_bound} could be outliers.")
print(f"Values above {c_upper_bound} could be outliers.")
print(f"Ramicane")
print(f"Lower quartile of Ramicane tumors = {r_lowerqt}")
print(f"Upper quartile of Ramicane tumors = {r_upperqt}")
print(f"Interquartile range of Ramicane tumors = {r_iqr}")
print(f"Values below {r_lower_bound} could be outliers.")
print(f"Values above {r_upper_bound} could be outliers.")
print(f"Infubinol")
print(f"Lower quartile of Infubinol tumors = {i_lowerqt}")
print(f"Upper quartile of Infubinol tumors = {i_upperqt}")
print(f"Interquartile range of Infubinol tumors = {i_iqr}")
print(f"Values below {i_lower_bound} could be outliers.")
print(f"Values above {i_upper_bound} could be outliers.")
print(f"Ceframin")
print(f"Lower quartile of Ceframin tumors = {ce_lowerqt}")
print(f"Upper quartile of Ceframin tumors = {ce_upperqt}")
print(f"Interquartile range of Ceframin tumors = {ce_iqr}")
print(f"Values below {ce_lower_bound} could be outliers.")
print(f"Values above {ce_upper_bound} could be outliers.")
# +
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest.
#means = [s.MEDV.mean() for s in samples]
boxplot_treatments = [capomulin_tumors, ramicane_tumors, infubinol_tumors, ceframin_tumors]
labels_treatments = ['Capomulin', 'Ramicane', 'Infubinol', 'Ceftamin']
fig, ax1 = plt.subplots()
plt.boxplot(boxplot_treatments, labels=labels_treatments, widths=.5, notch=False, vert=True)
#ax1.set_xlim(0, len(means)+1)
ax1.set_xlabel("Treatments")
ax1.set_ylabel("Final Tumor Volume (mm3)")
#plt.grid()
plt.show()
# -
# ## Line and Scatter Plots
# +
# Generate a line plot of tumor volume vs. time point for a mouse treated with Capomulin.
mouse_tdata = capomulin_data.loc[capomulin_data['Mouse ID'] == "s185",:]
days= mouse_tdata["Timepoint"]
tvolume = mouse_tdata["Tumor Volume (mm3)"]
fig1, ax1 = plt.subplots()
plt.plot(days, tvolume, color="blue")
plt.title("Capomulin treatment of mouse s185")
plt.xlabel("Timepoint(days)")
#plt.xticks(np.arange(min(days), max(days)+1, 1.0))
plt.ylabel("Tumor Volume (mm3)")
plt.show()
# +
# Generate a scatter plot of average tumor volume vs. mouse weight for the Capomulin regimen
fig1, ax1 = plt.subplots()
#Where do u get data?
tumor_data = capomulin_data.groupby(['Mouse ID']).mean()
#x_values = ["Weight (g)"]
#y_values = ["Tumor Volume (mm3)"]
marker_size=20
plt.scatter(tumor_data['Weight (g)'],tumor_data['Tumor Volume (mm3)'], color = "red")
#Find correct name of merge of datas.
plt.xlabel('Weight (g)')
plt.ylabel('Average Tumor Volume (mm3)')
plt.show()
# -
# ## Correlation and Regression
# +
# Calculate the correlation coefficient and linear regression model.
# for mouse weight and average tumor volume for the Capomulin regimen.
#corr_model = st.linregress(capomulin_average['Weight (g)'],capomulin_average['Tumor Volume (mm3)'])
#x_values = capomulin_data["Weight (g)"] #Weight
#y_values = capomulin_data["Tumor Volume (mm3)"] #Avg tumor volume WHERE IS AVG TUMOR VOLUME?
# (slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
#regress_values = x_values * slope + intercept
# line_eq = "y = " + str(round(slope,2)) + "x +" +str(round(incept,2))
#plt.scatter(x_values, y_values)
#plt.plot(x_values, regress_values, "r-")
#plt.annotate(line_eq,(6,10), fontsize = 15, color="red")
#plt.xlabel('Weight (g)')
#plt.ylabel('Average Tumor Volume (mm3)')
#plt.show()
# -
| Pymaceuticals/02-Homework_Matplotlib_Instructions_Pymaceuticals_CARB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# + language="sh"
#
# algorithm_name='sklearn-boston-housing-mme'
#
# account=$(aws sts get-caller-identity --query Account --output text)
#
# # Get the region defined in the current configuration (default to us-west-2 if none defined)
# region=$(aws configure get region)
#
# ecr_image="${account}.dkr.ecr.${region}.amazonaws.com/${algorithm_name}:latest"
#
# # If the repository doesn't exist in ECR, create it.
# aws ecr describe-repositories --repository-names "${algorithm_name}" > /dev/null 2>&1
#
# if [ $? -ne 0 ]
# then
# aws ecr create-repository --repository-name "${algorithm_name}" > /dev/null
# fi
#
# # Get the login command from ECR and execute it directly
# $(aws ecr get-login --region ${region} --no-include-email --registry-ids ${account})
#
# # Build the docker image locally with the image name and then push it to ECR
# # with the full image name.
#
# # First clear out any prior version of the cloned repo
# rm -rf sagemaker-scikit-learn-container/
#
# # Clone the sklearn container repo
# git clone --single-branch --branch mme https://github.com/aws/sagemaker-scikit-learn-container.git
# cd sagemaker-scikit-learn-container/
#
# # Build the "base" container image that encompasses the installation of the
# # scikit-learn framework and all of the dependencies needed.
# docker build -q -t sklearn-base:0.20-2-cpu-py3 -f docker/0.20-2/base/Dockerfile.cpu --build-arg py_version=3 .
#
# # Create the SageMaker Scikit-learn Container Python package.
# python setup.py bdist_wheel --universal
#
# # Build the "final" container image that encompasses the installation of the
# # code that implements the SageMaker multi-model container requirements.
# docker build -q -t ${algorithm_name} -f docker/0.20-2/final/Dockerfile.cpu .
#
# docker tag ${algorithm_name} ${ecr_image}
#
# docker push ${ecr_image}
# -
import sagemaker
# +
sess = sagemaker.Session()
bucket = sess.default_bucket()
role = sagemaker.get_execution_role()
prefix = 'sklearn-boston-housing-mme'
training = sess.upload_data(path='housing.csv', key_prefix=prefix + "/training")
output = 's3://{}/{}/output/'.format(bucket,prefix)
print(training)
print(output)
# +
from sagemaker.sklearn import SKLearn
jobs = {}
for test_size in [0.2, 0.1, 0.05]:
sk = SKLearn(entry_point='sklearn-boston-housing.py',
role=role,
train_instance_count=1,
train_instance_type='ml.m5.large',
output_path=output,
hyperparameters={
'normalize': True,
'test-size': test_size,
}
)
sk.fit({'training':training}, wait=False)
jobs[sk.latest_training_job.name] = {}
jobs[sk.latest_training_job.name]['test-size'] = test_size
# -
jobs
# +
import boto3
sm = boto3.client('sagemaker')
for j in jobs.keys():
job = sm.describe_training_job(TrainingJobName=j)
jobs[j]['artifact'] = job['ModelArtifacts']['S3ModelArtifacts']
jobs[j]['key'] = '/'.join(job['ModelArtifacts']['S3ModelArtifacts'].split('/')[3:8])
# -
jobs
# + magic_args="-s \"$bucket\" \"$prefix\"" language="sh"
# aws s3 rm --recursive s3://$1/$2/models
# +
s3 = boto3.client('s3')
for j in jobs.keys():
copy_source = { 'Bucket': bucket, 'Key': jobs[j]['key'] }
s3.copy_object(CopySource=copy_source, Bucket=bucket, Key=prefix+'/models/'+j+'.tar.gz')
response = s3.list_objects(Bucket=bucket, Prefix=prefix+'/models/')
for o in response['Contents']:
print(o['Key'])
# +
script = 'sklearn-boston-housing.py'
script_archive = 's3://{}/{}/source/source.tar.gz'.format(bucket, prefix)
print(script)
print(script_archive)
# + magic_args="-s \"$script\" \"$script_archive\"" language="sh"
# tar cvfz source.tar.gz $1
# aws s3 cp source.tar.gz $2
# +
account = boto3.client('sts').get_caller_identity()['Account']
region = boto3.Session().region_name
container = '{}.dkr.ecr.{}.amazonaws.com/{}:latest'.format(account, region, prefix)
print(container)
# +
response = sm.create_model(
ModelName = prefix,
ExecutionRoleArn = role,
Containers = [
{
'Image': container,
'ModelDataUrl': 's3://{}/{}/models/'.format(bucket, prefix),
'Mode': 'MultiModel',
'Environment': {
'SAGEMAKER_PROGRAM' : script,
'SAGEMAKER_SUBMIT_DIRECTORY' : script_archive
}
}
]
)
print(response)
# +
epc_name = prefix+'-epc'
response = sm.create_endpoint_config(
EndpointConfigName = epc_name,
ProductionVariants=[{
'InstanceType': 'ml.m5.large',
'InitialInstanceCount': 1,
'InitialVariantWeight': 1,
'ModelName': prefix,
'VariantName': 'variant-1'}
]
)
print(response)
# +
ep_name = prefix+'-ep'
response = sm.create_endpoint(
EndpointName=ep_name,
EndpointConfigName=epc_name)
print(response)
# -
sm.describe_endpoint(EndpointName=ep_name)
waiter = sm.get_waiter('endpoint_in_service')
waiter.wait(EndpointName=ep_name)
# +
import pandas as pd
import numpy as np
from io import BytesIO
data = pd.read_csv('housing.csv', delim_whitespace=True)
payload = data[:1].drop(['medv'], axis=1)
buffer = BytesIO()
np.save(buffer, payload.values)
# +
smrt = boto3.client('runtime.sagemaker')
for j in jobs.keys():
model_name=j+'.tar.gz'
print(model_name)
response = smrt.invoke_endpoint(
EndpointName=ep_name,
TargetModel=model_name,
Body=buffer.getvalue(),
ContentType='application/x-npy')
print(response['Body'].read())
# +
sk = SKLearn(entry_point='sklearn-boston-housing.py',
role=role,
train_instance_count=1,
train_instance_type='ml.m5.large',
output_path=output,
hyperparameters={
'normalize': False,
'test-size': 0.15,
}
)
sk.fit({'training':training})
# -
job = sm.describe_training_job(TrainingJobName=sk.latest_training_job.name)
job_name = sk.latest_training_job.name
artifact = job['ModelArtifacts']['S3ModelArtifacts']
# + magic_args="-s \"$artifact\" \"$bucket\" \"$prefix\" \"$job_name\"" language="sh"
# aws s3 cp $1 s3://$2/$3/models/$4.tar.gz
# aws s3 ls s3://$2/$3/models/
# +
model_name=job_name+'.tar.gz'
print(model_name)
response = smrt.invoke_endpoint(
EndpointName=ep_name,
TargetModel=model_name,
Body=buffer.getvalue(),
ContentType='application/x-npy')
print(response['Body'].read())
# -
sm.delete_endpoint(EndpointName=ep_name)
sm.delete_endpoint_config(EndpointConfigName=epc_name)
sm.delete_model(ModelName=prefix)
| sdkv1/ch13/multi_model/Scikit-learn on Boston Housing Dataset - Multi-Model Endpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Logistic Regression
# ---
# For us to utilize Logistic Regression in TensorFlow, we first need to import the required libraries. To do so, you can run the code cell below.
import tensorflow as tf
import pandas as pd
import numpy as np
import time
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# Next, we will load the dataset we are going to use. In this case, we are utilizing the <code>iris</code> dataset, which is inbuilt -- so there's no need to do any preprocessing and we can jump right into manipulating it. We separate the dataset into <i>xs</i> and <i>ys</i>, and then into training <i>xs</i> and <i>ys</i> and testing <i>xs</i> and <i>ys</i>, (pseudo)randomly.
# <h3>Understanding the Data</h3>
#
# <h4><code>Iris Dataset</code>:</h4>
# This dataset was introduced by British Statistician and Biologist Ronald Fisher, it consists of 50 samples from each of three species of Iris (Iris setosa, Iris virginica and Iris versicolor). In total it has 150 records under five attributes - petal length, petal width, sepal length, sepal width and species. <a href="https://archive.ics.uci.edu/ml/datasets/iris">Dataset source</a>
#
# Attributes
# Independent Variable
# <ul>
# <li>petal length</li>
# <li>petal width</li>
# <li>sepal length</li>
# <li>sepal width</li>
# </ul>
# Dependent Variable
# <ul>
# <li>Species
# <ul>
# <li>Iris setosa</li>
# <li>Iris virginica</li>
# <li>Iris versicolor</li>
# </ul>
# </li>
# </ul>
# <br>
iris = load_iris()
iris_X, iris_y = iris.data[:-1,:], iris.target[:-1]
iris_y= pd.get_dummies(iris_y).values
trainX, testX, trainY, testY = train_test_split(iris_X, iris_y, test_size=0.33, random_state=42)
# Now we define x and y. These placeholders will hold our iris data (both the features and label matrices), and help pass them along to different parts of the algorithm. You can consider placeholders as empty shells into which we insert our data. We also need to give them shapes which correspond to the shape of our data. Later, we will insert data into these placeholders by “feeding” the placeholders the data via a “feed_dict” (Feed Dictionary).
#
# <h3>Why use Placeholders?</h3>
#
# <ol>
# <li>This feature of TensorFlow allows us to create an algorithm which accepts data and knows something about the shape of the data without knowing the amount of data going in.</li>
# <li>When we insert “batches” of data in training, we can easily adjust how many examples we train on in a single step without changing the entire algorithm.</li>
# </ol>
# +
# numFeatures is the number of features in our input data.
# In the iris dataset, this number is '4'.
numFeatures = trainX.shape[1]
# numLabels is the number of classes our data points can be in.
# In the iris dataset, this number is '3'.
numLabels = trainY.shape[1]
# Placeholders
# 'None' means TensorFlow shouldn't expect a fixed number in that dimension
X = tf.placeholder(tf.float32, [None, numFeatures]) # Iris has 4 features, so X is a tensor to hold our data.
yGold = tf.placeholder(tf.float32, [None, numLabels]) # This will be our correct answers matrix for 3 classes.
# -
# <h3>Set model weights and bias</h3>
#
# Much like Linear Regression, we need a shared variable weight matrix for Logistic Regression. We initialize both <code>W</code> and <code>b</code> as tensors full of zeros. Since we are going to learn <code>W</code> and <code>b</code>, their initial value does not matter too much. These variables are the objects which define the structure of our regression model, and we can save them after they have been trained so we can reuse them later.
#
# We define two TensorFlow variables as our parameters. These variables will hold the weights and biases of our logistic regression and they will be continually updated during training.
#
# Notice that <code>W</code> has a shape of [4, 3] because we want to multiply the 4-dimensional input vectors by it to produce 3-dimensional vectors of evidence for the difference classes. <code>b</code> has a shape of [3] so we can add it to the output. Moreover, unlike our placeholders above which are essentially empty shells waiting to be fed data, TensorFlow variables need to be initialized with values, e.g. with zeros.
W = tf.Variable(tf.zeros([4, 3])) # 4-dimensional input and 3 classes
b = tf.Variable(tf.zeros([3])) # 3-dimensional output [0,0,1],[0,1,0],[1,0,0]
# +
#Randomly sample from a normal distribution with standard deviation .01
weights = tf.Variable(tf.random_normal([numFeatures,numLabels],
mean=0,
stddev=0.01,
name="weights"))
bias = tf.Variable(tf.random_normal([1,numLabels],
mean=0,
stddev=0.01,
name="bias"))
# -
# <h3>Logistic Regression model</h3>
#
# We now define our operations in order to properly run the Logistic Regression. Logistic regression is typically thought of as a single equation:
#
# $$
# ŷ =sigmoid(WX+b)
# $$
#
# However, for the sake of clarity, we can have it broken into its three main components:
# - a weight times features matrix multiplication operation,
# - a summation of the weighted features and a bias term,
# - and finally the application of a sigmoid function.
#
# As such, you will find these components defined as three separate operations below.
#
# Three-component breakdown of the Logistic Regression equation.
# Note that these feed into each other.
apply_weights_OP = tf.matmul(X, weights, name="apply_weights")
add_bias_OP = tf.add(apply_weights_OP, bias, name="add_bias")
activation_OP = tf.nn.sigmoid(add_bias_OP, name="activation")
# As we have seen before, the function we are going to use is the <i>logistic function</i> $(\frac{1}{1+e^{-Wx}})$, which is fed the input data after applying weights and bias. In TensorFlow, this function is implemented as the <code>nn.sigmoid</code> function. Effectively, this fits the weighted input with bias into a 0-100 percent curve, which is the probability function we want.
# <a id="ref3"></a>
# <h2>Training</h2>
#
# The learning algorithm is how we search for the best weight vector (${\bf w}$). This search is an optimization problem looking for the hypothesis that optimizes an error/cost measure.
#
# <b>What tell us our model is bad?</b>
# The Cost or Loss of the model, so what we want is to minimize that.
#
# <b>What is the cost function in our model?</b>
# The cost function we are going to utilize is the Squared Mean Error loss function.
#
# <b>How to minimize the cost function?</b>
# We can't use <b>least-squares linear regression</b> here, so we will use <a href="http://en.wikipedia.org/wiki/Gradient_descent">gradient descent</a> instead. Specifically, we will use batch gradient descent which calculates the gradient from all data points in the data set.
#
# <h3>Cost function</h3>
# Before defining our cost function, we need to define how long we are going to train and how should we define the learning rate.
# +
# Number of Epochs in our training
numEpochs = 700
# Defining our learning rate iterations (decay)
learningRate = tf.train.exponential_decay(learning_rate=0.0008,
global_step= 1,
decay_steps=trainX.shape[0],
decay_rate= 0.95,
staircase=True)
# +
#Defining our cost function - Squared Mean Error
cost_OP = tf.nn.l2_loss(activation_OP-yGold, name="squared_error_cost")
#Defining our Gradient Descent
training_OP = tf.train.GradientDescentOptimizer(learningRate).minimize(cost_OP)
# -
# Now we move on to actually running our operations. We will start with the operations involved in the prediction phase (i.e. the logistic regression itself).
#
# First, we need to initialize our weights and biases with zeros or random values via the inbuilt Initialization Op, <b>tf.initialize_all_variables()</b>. This Initialization Op will become a node in our computational graph, and when we put the graph into a session, then the Op will run and create the variables.
# +
# Create a tensorflow session
sess = tf.Session()
# Initialize our weights and biases variables.
init_OP = tf.global_variables_initializer()
# Initialize all tensorflow variables
sess.run(init_OP)
# -
# We also want some additional operations to keep track of our model's efficiency over time. We can do this like so:
# +
# argmax(activation_OP, 1) returns the label with the most probability
# argmax(yGold, 1) is the correct label
correct_predictions_OP = tf.equal(tf.argmax(activation_OP,1),tf.argmax(yGold,1))
# If every false prediction is 0 and every true prediction is 1, the average returns us the accuracy
accuracy_OP = tf.reduce_mean(tf.cast(correct_predictions_OP, "float"))
# Summary op for regression output
activation_summary_OP = tf.summary.histogram("output", activation_OP)
# Summary op for accuracy
accuracy_summary_OP = tf.summary.scalar("accuracy", accuracy_OP)
# Summary op for cost
cost_summary_OP = tf.summary.scalar("cost", cost_OP)
# Summary ops to check how variables (W, b) are updating after each iteration
weightSummary = tf.summary.histogram("weights", weights.eval(session=sess))
biasSummary = tf.summary.histogram("biases", bias.eval(session=sess))
# Merge all summaries
merged = tf.summary.merge([activation_summary_OP, accuracy_summary_OP, cost_summary_OP, weightSummary, biasSummary])
# Summary writer
writer = tf.summary.FileWriter("summary_logs", sess.graph)
# -
# Now we can define and run the actual training loop, like this:
# +
# Initialize reporting variables
cost = 0
diff = 1
epoch_values = []
accuracy_values = []
cost_values = []
# Training epochs
for i in range(numEpochs):
if i > 1 and diff < .0001:
print("change in cost %g; convergence."%diff)
break
else:
# Run training step
step = sess.run(training_OP, feed_dict={X: trainX, yGold: trainY})
# Report occasional stats
if i % 10 == 0:
# Add epoch to epoch_values
epoch_values.append(i)
# Generate accuracy stats on test data
train_accuracy, newCost = sess.run([accuracy_OP, cost_OP], feed_dict={X: trainX, yGold: trainY})
# Add accuracy to live graphing variable
accuracy_values.append(train_accuracy)
# Add cost to live graphing variable
cost_values.append(newCost)
# Re-assign values for variables
diff = abs(newCost - cost)
cost = newCost
#generate print statements
print("step %d, training accuracy %g, cost %g, change in cost %g"%(i, train_accuracy, newCost, diff))
# How well do we perform on held-out test data?
print("final accuracy on test set: %s" %str(sess.run(accuracy_OP,
feed_dict={X: testX,
yGold: testY})))
# -
# <b>Why don't we plot the cost to see how it behaves?</b>
# %matplotlib inline
plt.plot([np.mean(cost_values[i-50:i]) for i in range(len(cost_values))])
plt.show()
# Assuming no parameters were changed, you should reach a peak accuracy of 90% at the end of training, which is commendable. Try changing the parameters such as the length of training, and maybe some operations to see how the model behaves. Does it take much longer? How is the performance?
# ## Thanks for reading :)
# Created by [<NAME>](https://www.linkedin.com/in/saeedaghabozorgi/) and modified by [<NAME>](https://www.linkedin.com/in/kambojtarun/).
| Code/Logistic Regression/Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import pandas as pd
import os
from tensorflow.keras.models import load_model
# +
model = load_model('models/weights.best.basic_cnn.hdf5')
directory = 'models/basic_cnn_history'
##put all csvs into dataframes and into list
frames = [pd.read_csv(directory +'/' +path) for path in os.listdir(directory)]
### and separate dataframes
train_loss, val_loss, val_acc, train_acc = frames[0], frames[1], frames[2], frames[3]
# +
plt.title("Training and Validation Loss")
plt.plot(train_loss["Value"] , label='Training')
plt.plot(val_loss["Value"], label='Validation' )
plt.legend()
plt.xlabel("Epoch")
plt.ylabel("Loss")
# +
plt.title("Training and Validation Accuracy")
plt.plot(train_acc["Value"] , label='Training')
plt.plot(val_acc["Value"], label='Validation' )
plt.legend()
plt.xlabel("Epoch")
plt.ylabel("Accuracy")
# -
print("Best validation loss epoch: ", np.argmin(val_loss['Value'].values))
print("Best validation accuracy epoch: ", np.argmax(val_acc['Value'].values))
print("Best validation loss: ", val_loss.Value.min())
print("Best validation accuracy: ", val_acc.Value.max())
print('Top epochs for loss:')
val_loss.sort_values(by='Value')[:10]
# By the 175th epoch, the model had already reached 0.2316 validation loss. The lowest val loss model (379th epoch) offered a 0.0075 decrease in loss from this.
#
print("Best validation accuracies")
val_acc.sort_values(by='Value', ascending=False)[:10]
# Highest validation accuracy occured at the 466th epoch
# The model below gave the best performance yet, as of 2/26/2020. 72 epochs of training
#
# Average validation accuracy: 66.13%
#
# n_mels = 60
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.callbacks import ModelCheckpoint, TensorBoard, EarlyStopping
from tensorflow.keras.layers import Dense, Dropout, Activation, Flatten
from tensorflow.keras.layers import Convolution2D, Conv2D, MaxPooling2D, GlobalAveragePooling2D
from tensorflow.keras.regularizers import l2
from tensorflow.keras.optimizers import Adam
from tensorflow.keras import utils
from tensorflow.keras.metrics import AUC
num_rows = 60
num_columns = 174 #number of frames / max padding
num_channels = 1
num_labels = 10 #categories
# -
def get_conv_model():
model = Sequential()
model.add(Conv2D(filters=16, kernel_size=(2,2), kernel_regularizer=l2(0.0001),
input_shape=(num_rows, num_columns, num_channels), activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Conv2D(filters=32, kernel_size=(2,2), kernel_regularizer=l2(0.0001), activation='relu'))
model.add(Dropout(0.2))
model.add(Conv2D(filters=64, kernel_size=(3,3), kernel_regularizer=l2(0.0001), activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_labels, activation='softmax'))
model.compile(loss='categorical_crossentropy', metrics=['accuracy', AUC()], optimizer='adam')
return model
model = get_conv_model()
model.summary()
# +
#Importing tensorboard logs
val_acc_dict = {}
path = 'past_logs/acc_csvs_1'
for i, fpath in enumerate(sorted(os.listdir(path), key=lambda name: name[-28:])):
# val_acc_dict
print(fpath)
val_acc_dict[f"fold{i+1}"] = pd.read_csv(os.path.join(path,fpath))
# +
fig, ax = plt.subplots()
ax.set_title("Validation Accuracy")
for fold, df in val_acc_dict.items():
ax.plot(df.Value*100, linewidth=1.25, label=fold)
ax.set_xlabel('Epoch')
ax.set_ylabel('% Accuracy')
ax.legend(ncol=2)
# +
model_dict = {}
for i in range(1,11):
model_dict[i] = load_model('models/mels_basic_cnn_fold{}.hdf5'.format(i))
# -
from sklearn.metrics import confusion_matrix
labels = np.array(['dog_bark', 'children_playing', 'children_playing', ...,
'car_horn', 'car_horn', 'car_horn'], dtype='<U16')
labels
| notebooks/us_model_history_plotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Quantifiers
#
# This notebook focuses on the general usage of quantifiers rather than specific details of each quantifier, which can be found in the [documentation](https://notes.eonu.net/docs/sigment/).
#
# To best understand this notebook, it is recommended that you read the [Using Transformations](https://nbviewer.jupyter.org/github/eonu/sigment/blob/master/notebooks/Using%20Transformations.ipynb) notebook first.
#
# ---
#
# **Contents**:
#
# - [What are quantifiers?](#What-are-quantifiers?)
# - [Nested quantifiers](#Nested-quantifiers)
# - [Probabilistic quantification](#Probabilistic-quantification)
# - [Applying in random order](#Applying-in-random-order)
import matplotlib.pyplot as plt
from librosa import load
from sigment import *
from utils import plot
plt.style.use('ggplot')
# Load some sample audio
X, sr = load('assets/audio.wav', mono=False)
# ## What are quantifiers?
#
# Quantifiers (along with transformations) are one of the two core components of Sigment. They allow you to collectively apply multiple transformations to an audio signal, according to how the quantifier specifies the transformations should be applied.
#
# The most basic quantifier is the `Pipeline`, which simply applies all of the specified transformations in the pipeline in order:
# +
# Create an augmentation pipeline
aug = Pipeline([
TimeStretch(rate=0.8),
GaussianWhiteNoise(scale=(0.01, 0.015))
])
# Augment and plot the audio signal
plot(X, aug(X), sr=sr)
# -
# As mentioned in the [Using Transformations](https://nbviewer.jupyter.org/github/eonu/sigment/blob/master/notebooks/Using%20Transformations.ipynb) notebook, it is sometimes useful to use the `p` parameter in transformations, to introduce an element of randomness to a quantifier:
#
# ```python
# aug = Pipeline([
# TimeStretch(rate=0.8, p=0.4),
# GaussianWhiteNoise(scale=(0.01, 0.015), p=0.85)
# ])
# ```
#
# To control the randomness, rather than specifying the `random_state` argument for each transformation, you can specify a global random state as a parameter of the quantifier. Doing this will internally set the same random state object for each transformation:
#
# ```python
# aug = Pipeline([
# TimeStretch(rate=0.8, p=0.4),
# GaussianWhiteNoise(scale=(0.01, 0.015), p=0.85)
# ], random_state=0)
# ```
# ## Nested quantifiers
#
# Sigment supports the creation of more complex data augmentation pipelines by allowing quantifiers to be nested within each other.
#
# Suppose we wish to take the previous pipeline consisting of `TimeStretch` and `GaussianWhiteNoise` transformations, and add either a fade in or fade out transformation afterwards. We can nest the pipeline with a `OneOf` quantifier to do this:
# +
# Create an augmentation pipeline
aug = Pipeline([
TimeStretch(rate=0.8),
GaussianWhiteNoise(scale=(0.01, 0.015)),
OneOf([
LinearFade('in', fade_size=(0.1, 0.2)),
LinearFade('out', fade_size=(0.1, 0.2))
])
])
# Augment and plot the audio signal
plot(X, aug(X), sr=sr)
# -
# Once again, although all quantifiers and transforms accept a `random_state` argument, it is only necessary to include it in the outermost quantifier:
#
# ```python
# aug = Pipeline([
# TimeStretch(rate=0.8),
# GaussianWhiteNoise(scale=(0.01, 0.015)),
# OneOf([
# LinearFade('in', fade_size=(0.1, 0.2)),
# LinearFade('out', fade_size=(0.1, 0.2))
# ])
# ], random_state=0)
# ```
# ## Probabilistic quantification
#
# Unlike transformations, not all quantifiers (and therefore the transformations that fall under them) can be applied probabilistically by specifying a `p` argument.
#
# Only the `Sometimes` quantifier can be used to achieve this:
#
# ```python
# aug = Pipeline([
# TimeStretch(rate=0.8),
# GaussianWhiteNoise(scale=(0.01, 0.015)),
# Sometimes([
# OneOf([
# LinearFade('in', fade_size=(0.1, 0.2)),
# LinearFade('out', fade_size=(0.1, 0.2))
# ])
# ], p=0.65)
# ], random_state=0)
# ```
# ## Applying in random order
#
# In some cases, it might be desirable to randomize the order in which transformations defined under a quantifier are applied. Setting the `random_order` parameter (defined on all quantifiers) to `True` can be used to achieve this:
#
# ```python
# aug = Pipeline([
# TimeStretch(rate=0.8, p=0.4),
# GaussianWhiteNoise(scale=(0.01, 0.015), p=0.85)
# ], random_state=0, random_order=True)
# ```
#
# The `random_state` parameter can also be used to control the randomness of the `random_order` parameter.
| notebooks/Using Quantifiers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''herbert'': pipenv)'
# name: python37364bitherbertpipenvf409fddaf3f446fd8dcf7490c441f6bd
# ---
# # PoLitBert - Polish RoBERT'a model
#
# ## Preparation of vocabulary and encoding the data
#
# Used corpuses:
# * Wikipedia, Link:
# * Oscar
# * Polish Books
# Usefull resources
# * https://github.com/pytorch/fairseq/blob/master/examples/roberta/README.pretraining.md
# * https://github.com/musixmatchresearch/umberto/issues/2
# + pycharm={"name": "#%%\n"}
import csv
import sys
import datetime as dt
import os
from pathlib import Path
import re
from tqdm import tqdm
import mmap
# -
# ## Create vocabulary
#
# ### Prepare data for vocab
#
# Separate text file for training vocabulary has been created with one sentence per line.
# We used polish sentence tokenizer with [additional abbreviations](https://gist.github.com/ksopyla/f05fe2f48bbc9de895368b8a7863b5c3)
# typical for the Polish language.
# Sentencepiece model is capable of handling around 12.000.000 sentences, so larger files are not necessary.
# ### Train the BPE vocabulary model
#
# We used the [SentencePiece](https://github.com/google/sentencepiece) segmentation model trained from raw
# sentences with fixed final vocabulary size - 32K and 50K unique tokens.
#
# Training and segmentation can be done in two ways:
# - as a python module,
# - as a command-line tool.
#
# To use it as a command-line it should be installed from source, which is described in the
# [build the C++ version from source](https://github.com/google/sentencepiece#c-from-source) section of the documentation.
#
# #### Training SentencePiece vocab using command line
#
# * 32k vocab:
# ```
# spm_train \
# --input=./data/corpus_raw/corpus_books_wiki_12M_lines.txt \
# --max_sentence_length=4192\
# --model_prefix=./data/vocab/books_wikipedia_v32k_sen10M.spm.bpe \
# --vocab_size=32000 \
# --model_type=bpe \
# --shuffle_input_sentence=true \
# --input_sentence_size=10000000 \
# --bos_id=0 --eos_id=1 --pad_id=2 --unk_id=3
# ```
#
# * 50k vocab:
# ```
# spm_train \
# --input=./data/corpus_raw/corpus_books_wiki_12M_lines.txt \
# --max_sentence_length=4192\
# --model_prefix=./data/vocab/books_wikipedia_v50k_sen10M.spm.bpe \
# --vocab_size=50000 \
# --model_type=bpe \
# --shuffle_input_sentence=true \
# --input_sentence_size=10000000 \
# --bos_id=0 --eos_id=1 --pad_id=2 --unk_id=3
# ```
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Training SentencePiece vocab with Python module
#
# Below, for reference, an example of how to prepare a SP model if Python script is preferred.
# + pycharm={"name": "#%%\n"}
import sentencepiece as spm
vocab_size = 32000
model_type = "bpe"
iss = 10_000_000
data_file = './data/corpus_raw/corpus_books_wiki_12M_lines.txt'
tok_model = f"books_wikipedia_v32k_sen10M"
tok_model = os.path.abspath(f"./data/vocab/{tok_model}")
piece_options = ' --bos_id=0 --eos_id=1 --pad_id=2 --unk_id=3 --shuffle_input_sentence=true'
cmd = f"--input={data_file} --model_prefix={tok_model} --num_threads=4 --vocab_size={vocab_size} --input_sentence_size={iss}" + piece_options
print(cmd)
start = dt.datetime.now()
print(start)
spm.SentencePieceTrainer.train(cmd)
end = dt.datetime.now()
print(f"Created vocab of {vocab_size} tokens from {data_file}, took {end-start}.")
# +
# Example segmentation usage:
# make segmenter instance and load the model file (m.model)
sp = spm.SentencePieceProcessor()
sp.load(f"{tok_model}.model")
# verify vocab size
print(sp.get_piece_size())
# encode: text => id
text = """Będąc młodym programistą (hoho), czytałem "Dziady" w 1983r."""
print(sp.encode_as_pieces(text))
# -
# ### Fairseq vocab
#
# Usage of sentencepiece the model's with fairseq requires changing the separator used in the dictionary.
# All _\t_ characters should be replaced with _whitespace_ in the vocab file.
for vocab_size in ("32k", "50k"):
vocab_file = f"./data/vocab/books_wikipedia_v{vocab_size}_sen10M.spm.bpe.vocab"
p = Path(vocab_file)
output_path = f"{p.with_suffix('')}_fair.vocab"
with open(output_path, 'w+') as output_file:
with open(vocab_file) as f:
text = f.read().replace('\t', ' ')
output_file.write(text)
# ### Encode data with sentence piece model
#
# Encoding prepared training and test datasets with SentencePiece tokenizer. Both, for 32k and 50k vocabularies.
#
# * 32k vocab:
#
# ```
# DATA_PATH=./data/wiki_books_oscar/
# VOCAB_SIZE=32k
#
# for SPLIT in test train ; do \
# spm_encode \
# --model=./data/vocab/books_wikipedia_v${VOCAB_SIZE}_sen10M.spm.bpe.model \
# --extra_options=bos:eos \
# --output_format=piece \
# < ${DATA_PATH}corpus_wiki_books_oscar_${SPLIT}.txt \
# > ${DATA_PATH}corpus_wiki_books_oscar_${SPLIT}_${VOCAB_SIZE}.txt.bpe
# done
# ```
#
# * 50k vocab:
#
# ```
# DATA_PATH=./data/wiki_books_oscar/
# VOCAB_SIZE=50k
#
# for SPLIT in test train ; do \
# spm_encode \
# --model=./data/vocab/books_wikipedia_v${VOCAB_SIZE}_sen10M.spm.bpe.model \
# --extra_options=bos:eos \
# --output_format=piece \
# < ${DATA_PATH}corpus_wiki_books_oscar_${SPLIT}.txt \
# > ${DATA_PATH}corpus_wiki_books_oscar_${SPLIT}_${VOCAB_SIZE}.txt.bpe
# done
# ```
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Data binarization with Fairseq
#
# ### Fairseq-preprocessing bpe encoded and splited data
#
# * Data processed with 32k vocab:
#
# ```
# DATA_PATH=./data/wiki_books_oscar/
# VOCAB_SIZE=32k
#
# fairseq-preprocess \
# --only-source \
# --srcdict ./vocab/books_wikipedia_v${VOCAB_SIZE}_sen10M.spm.bpe_fair.vocab \
# --trainpref ${DATA_PATH}corpus_wiki_books_oscar_train_vocab${VOCAB_SIZE}.txt.bpe \
# --validpref ${DATA_PATH}corpus_wiki_books_oscar_test_vocab${VOCAB_SIZE}.txt.bpe \
# --destdir ${DATA_PATH}vocab${VOCAB_SIZE} \
# --workers 8
# ```
#
# * Data processed with 50k vocab:
#
# ```
# DATA_PATH=./data/wiki_books_oscar/
# VOCAB_SIZE=50k
#
# fairseq-preprocess \
# --only-source \
# --srcdict ./vocab/books_wikipedia_v${VOCAB_SIZE}_sen10M.spm.bpe_fair.vocab \
# --trainpref ${DATA_PATH}corpus_wiki_books_oscar_train_vocab${VOCAB_SIZE}.txt.bpe \
# --validpref ${DATA_PATH}corpus_wiki_books_oscar_test_vocab${VOCAB_SIZE}.txt.bpe \
# --destdir ${DATA_PATH}vocab${VOCAB_SIZE} \
# --workers 8
# ```
| polish_roberta_vocab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Looking into the new eleanor hlsp http://adina.feinste.in/eleanor/getting_started/about.html. NB only sectors 1-13 done yet, come back to this later?
# +
import numpy as np
import matplotlib.pyplot as plt
import astropy.io.fits as fits
import os
import glob
from astropy.table import Table
from astropy.io import ascii
import astropy.units as u
import astropy.constants as const
from astropy.modeling import models, fitting
import lightkurve as lk
from astropy.timeseries import LombScargle
#matplotlib set up
# %matplotlib inline
from matplotlib import rcParams
rcParams["figure.figsize"] = (14, 5)
rcParams["font.size"] = 20
# -
import eleanor
from astropy.coordinates import SkyCoord
star = eleanor.Source(tic=259773610, sector=15)
data = eleanor.TargetData(star)
vis = eleanor.Visualize(data)
| old/.ipynb_checkpoints/eleanor_test-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="BqkGR-LB_-Bf"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
# sns.set_theme(style="darkgrid")
import scipy as sp
from sklearn.ensemble import RandomForestClassifier
from sklearn.impute import SimpleImputer
from datetime import datetime
import os
# -
exec(open("../../header.py").read())
# **Description**:
#
# The dataset contains medical informa-tion for 858 patients fromHospital Universitario de Caracas. There are 32 numericaland binary features including age, number of pregnancies, and use of IUD. The targetvariable isBiopsy, which is a binary variable
# # Import data
# + id="vXpomyt6FMz2"
# read in data and replace nans
data = pd.read_csv(raw_root('risk_factors_cervical_cancer.csv')).\
replace('?', np.nan)
# -
# # Clean data
# + id="V73VBEsDITTj"
# impute, default is mean
my_imputer = SimpleImputer()
data_final = pd.DataFrame(my_imputer.fit_transform(data))
data_final.columns= data.columns
# + id="cWk2pkalG5Rm"
# all the target variables, i only use Biopsy
targets = ['Hinselmann', 'Schiller','Citology','Biopsy']
# + id="tux7W3G4GlBM"
# split data
X = data_final.loc[:, ~data_final.columns.isin(targets)]
y = data_final['Biopsy']
# -
X
# # Save data
try:
os.mkdir(processed_root("cervical_cancer_risks"))
except FileExistsError:
print("Folder already exists")
X.to_csv(processed_root("cervical_cancer_risks/X.csv"), index = False)
y.to_csv(processed_root("cervical_cancer_risks/y.csv"), index = False)
| notebooks/00-data/01-cervical-cancer-risk.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import torch.nn as nn
import torch.optim as optim
import numpy as np
# +
class TreeNNLayer(nn.Module):
def __init__(self, layer_class, out_features, *in_layers, **kwargs):
super(__class__, self).__init__()
self.in_layers = in_layers
if len(in_layers):
self.has_in_layers = True
self.in_features = sum(map(lambda l: l.out_features, in_layers))
self.in_key = None
else:
self.has_in_layers = False
self.in_features = kwargs['in_features']
self.in_key = kwargs['in_key']
self.out_features = out_features
layer_args = {}
if 'layer_args' in kwargs:
layer_args = kwargs['layer_args']
# FIXME find a better way to know whether to pass in/out features
if getattr(nn, layer_class).__module__.endswith('activation'):
self.layer = getattr(nn, layer_class)(**layer_args)
else:
self.layer = getattr(nn, layer_class)(self.in_features, self.out_features, **layer_args)
def forward(self, X):
if not self.has_in_layers:
assert isinstance(X, dict)
return self.layer(X[self.in_key])
inputs = []
for layer in self.in_layers:
inputs.append(layer(X))
return self.layer(torch.cat(inputs, 1))
class TreeNN(nn.Module):
def __init__(self, loss_class, optimizer_class, lr, out_layer, loss_args = {}, optimizer_args = {}):
super(__class__, self).__init__()
self.out_layer = out_layer
self.loss = getattr(nn, loss_class)(**loss_args)
optimizer_args['lr'] = lr
self.optimizer = getattr(optim, optimizer_class)(self.parameters(), **optimizer_args)
def forward(self, X):
return self.out_layer(X)
def fit(self, X, Y):
Y_hat = self(X)
loss = self.loss(Y_hat, Y)
loss.backward()
self.optimizer.step()
self.zero_grad()
return loss.item()
# -
treenn = TreeNN(
'MSELoss', 'Adagrad', 0.1, TreeNNLayer('Linear', 1,
TreeNNLayer('ReLU', 8,
TreeNNLayer('Linear', 8, in_features = 4, in_key = 'foo')
),
TreeNNLayer('ReLU', 8,
TreeNNLayer('Linear', 8, in_features = 4, in_key = 'bar')
)
)
)
treenn
treenn.fit({
'foo': torch.tensor([1, 2, 3, 4], dtype=torch.float).view(1, -1),
'bar': torch.tensor([10, 20, 30, 40], dtype=torch.float).view(1, -1)
}, torch.tensor([5], dtype=torch.float).view(1, -1))
| TreeNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mrfoxie/colab/blob/master/Colab%20RDP/Colab%20RDP.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="iLh_9SkSut4u"
# # **Colab RDP** : Remote Desktop to Colab Instance
#
# Used Google Remote Desktop & Ngrok Tunnel
#
# > **Warning :** This notebook is against the Policy of Colab. Use it on your own risk
#
# Google Colab can give you Instance with 12GB of RAM and GPU for 12 hours (Max.). Anyone can use it to perform Heavy Tasks
#
# To use other similiar Notebooks use my Repository **[Colab Hacks](https://github.com/mrfoxie/colab)**
# + [markdown] id="cpktYY4Cwqxf"
# ### Before We Started
# - Create a Ngrok Account to configure tunnel
# - Don't use username : `root`
# + [markdown] id="RLkhifGWu_1X"
# ### Installing Colab SSH Library
# + id="Ks-xI7nV2aW5"
# ! pip install colab_ssh --upgrade
from colab_ssh import launch_ssh, init_git
# + [markdown] id="3LAjx5HyvUe0"
# ### Configure User
# + [markdown] id="iY0xW8IrvklB"
# Setting Username & Password
# + id="fMXfodDURPZB"
username = input("Enter username : ")
password = input("Enter password : ")
# + [markdown] id="EcUIE8VDvcCD"
# Create User & Set Password
# + id="uep_NskcRq4c"
# ! sudo useradd -m $username
# ! sudo adduser $username sudo
# ! echo '$username:$password' | sudo chpasswd
# ! sed -i 's/\/bin\/sh/\/bin\/bash/g' /etc/passwd
# + [markdown] id="mnK94LuVvt3a"
# ### Mount Google Drive
# + id="GpmseAlaYuwY"
from google.colab import drive
home='/home/'+username+'/drive'
drive.mount(home)
# + [markdown] id="70AUcQIPxB0e"
# ### Setting Chrome Remote Desktop
# + [markdown] id="3bo6Aq7xxkjT"
# Download Remote Desktop Installation Script
# + id="p_9JGChxxKDs"
# ! wget https://github.com/mrfoxie/colab/blob/main/Colab%20RDP/install.sh -P /home/$username
# + [markdown] id="zsJ8sPTfx3oE"
# Use the script
#
# - It install the required files and configure your instance
# - Wait at least 4-5 mintues to complete the process
# + id="1EsGY1VQ4mry"
# ! sudo chmod +x /home/$username/install.sh
# ! sudo /home/$username/install.sh
# + [markdown] id="zNOpEkSRvzDN"
# ### Run Ngrok Tunnel
# + id="tNtwEGhvSJMB"
print("Copy authtoken from https://dashboard.ngrok.com/auth")
ngrokToken = input("Enter Token : ")
launch_ssh(ngrokToken, password)
# + [markdown] id="IoG0YmUPxnsT"
# Connect to instance via SSH
# - Use the command below in your terminal
# - When ask for password give your password which is set previously
# + id="KjSebmRoSlOl"
print("ssh", username, end='@')
# !curl -s http://localhost:4040/api/tunnels | python3 -c \
# "import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'][6:].replace(':', ' -p '))"
# + [markdown] id="b2BgUYJj8mHU"
# If unable to Connect through SSH then Kill and Start NGROK Tunnel again
# + [markdown] id="VyDzoqrcyeo1"
# ### Authenticate Through [Remote Desktop Headless](http://remotedesktop.google.com/headless)
# - Open browser and go to http://remotedesktop.google.com/headless
# - Dont Download any file simply proceed directly to authenticate button
# - when you got a command copy and paste it to SSH terminal and complete the process by setting up a pin
# + [markdown] id="zbtNhA25zOXm"
# Now your process completes
#
# Visit [Remote Desktop](http://remotedesktop.google.com/access) at http://remotedesktop.google.com/headless to access your instance
# + [markdown] id="DNwQ66F5v4iu"
# ### Kill Ngrok Tunnel
# + id="9gMg9YWns-f0"
# ! killall ngrok
# + [markdown] id="Sbus2Qb62MnM"
# ### Sleep Colab
# Sleep Colab for 12 hours
# + id="3UGxFLrh25LU"
# ! sleep 43200
| Colab RDP/Colab RDP.ipynb |