code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Example: CanvasXpress scatter2d Chart No. 10
#
# This example page demonstrates how to, using the Python package, create a chart that matches the CanvasXpress online example located at:
#
# https://www.canvasxpress.org/examples/scatter2d-10.html
#
# This example is generated using the reproducible JSON obtained from the above page and the `canvasxpress.util.generator.generate_canvasxpress_code_from_json_file()` function.
#
# Everything required for the chart to render is included in the code below. Simply run the code block.
# +
from canvasxpress.canvas import CanvasXpress
from canvasxpress.js.collection import CXEvents
from canvasxpress.render.jupyter import CXNoteBook
cx = CanvasXpress(
render_to="scatter2d10",
data={
"y": {
"vars": [
"S1",
"S2",
"S3",
"S4",
"S5",
"S6",
"S7",
"S8",
"S9",
"S10",
"S11",
"S12"
],
"smps": [
"Concentration",
"V1"
],
"data": [
[
0.0009,
172
],
[
0.0018,
177
],
[
0.0037,
160
],
[
0.0073,
166
],
[
0.0146,
211
],
[
0.0293,
248
],
[
0.0586,
269
],
[
0.117,
283
],
[
0.234,
298
],
[
0.469,
314
],
[
0.938,
328
],
[
1.88,
316
]
]
}
},
config={
"decorations": {
"nlfit": [
{
"type": "cst",
"param": [
"164",
"313",
0.031,
-1.5,
1.2e-06,
1.9
],
"label": "Custom Fit"
},
{
"param": [
"164",
"313",
0.031,
1.5,
1.2e-06,
1.9
],
"type": "reg",
"label": "Regular Fit"
}
]
},
"graphType": "Scatter2D",
"setMaxY": 350,
"setMinY": 100,
"showDecorations": True,
"theme": "CanvasXpress",
"xAxis": [
"Concentration"
],
"xAxisTransform": "log10",
"xAxisTransformTicks": False,
"yAxis": [
"V1"
],
"yAxisExact": True
},
width=613,
height=613,
events=CXEvents(),
after_render=[],
other_init_params={
"version": 35,
"events": False,
"info": False,
"afterRenderInit": False,
"noValidate": True
}
)
display = CXNoteBook(cx)
display.render(output_file="scatter2d_10.html")
|
tutorials/notebook/cx_site_chart_examples/scatter2d_10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### MaskRCNN training. Balloon dataset
# +
import os
os.chdir('..')
import tensorflow as tf
from samples.balloon import balloon
from preprocess import preprocess
from preprocess import augmentation as aug
from training import train_model
from model import mask_rcnn_functional
from common.utils import tf_limit_gpu_memory
# -
# %load_ext watermark
# %watermark
# %watermark --iversions
tf_limit_gpu_memory(tf, 4500)
from common.config import CONFIG
CONFIG.update({'class_dict': {'balloon': 1, 'background': 0},
'num_classes': 2,
'epochs': 30,
},
)
CONFIG.update({'meta_shape': (1 + 3 + 3 + 4 + 1 + CONFIG['num_classes']),})
CONFIG
model = mask_rcnn_functional(config=CONFIG)
CONFIG['training']
CONFIG['backbone']
# +
base_dir = os.getcwd().replace('src', 'balloon')
train_dir = os.path.join(base_dir, 'train')
val_dir = os.path.join(base_dir, 'val')
train_dataset = balloon.BalloonDataset(images_dir=train_dir,
class_key='object',
classes_dict=CONFIG['class_dict'],
augmentation=aug.get_training_augmentation(
image_size=CONFIG['img_size'],
normalize=CONFIG['normalization']
),
json_annotation_key=None,
**CONFIG
)
val_dataset = balloon.BalloonDataset(images_dir=val_dir,
class_key='object',
classes_dict=CONFIG['class_dict'],
augmentation=aug.get_validation_augmentation(
image_size=CONFIG['img_size'],
normalize=CONFIG['normalization']
),
json_annotation_key=None,
**CONFIG
)
# -
train_model(model,
train_dataset=train_dataset,
val_dataset=val_dataset,
config=CONFIG,
weights_path=None)
|
src/notebooks/example_training_balloon.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Ntyw6658nVwV" executionInfo={"status": "ok", "timestamp": 1647238288383, "user_tz": -540, "elapsed": 3759, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}}
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
import tensorflow as tf
from tensorflow.keras import Sequential,datasets
from tensorflow.keras.layers import Dense,Embedding,SimpleRNN,LSTM,GRU,Dropout
from tensorflow.keras.preprocessing import sequence
from sklearn.preprocessing import MinMaxScaler, StandardScaler
from sklearn.model_selection import train_test_split
# + colab={"base_uri": "https://localhost:8080/"} id="Lh6mkxCOnciZ" executionInfo={"status": "ok", "timestamp": 1647238302549, "user_tz": -540, "elapsed": 10860, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}} outputId="3991701b-9d59-40b7-d35b-d9c769d71972"
(X_train,y_train),(X_test,y_test) = datasets.imdb.load_data()
print(X_train.shape,y_train.shape)
print(X_test.shape,y_test.shape)
# + id="xQDHybNJrCUq" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1647238308177, "user_tz": -540, "elapsed": 349, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}} outputId="97b814e2-db81-476e-fc1a-feaf84b56cad"
print(X_train[0][:5], X_train[0].__len__())
# + id="F2UqBXrYrD23" colab={"base_uri": "https://localhost:8080/", "height": 339} executionInfo={"status": "ok", "timestamp": 1647238310951, "user_tz": -540, "elapsed": 889, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}} outputId="a8f7cf0f-f567-4275-c202-af2e1456ac6a"
sns.distplot(a=[len(x) for x in X_train])
# + id="3STYHAJdrGS-" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1647238314366, "user_tz": -540, "elapsed": 449, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}} outputId="8a57760a-13e5-4b7b-f0ca-0eff4272d2d0"
# 정수 4부터가 실제 IMDB 리뷰 데이터셋에서 빈도수가 가장 높은 영단어를 나타낸다.
words = datasets.imdb.get_word_index() # 저장된 값에 +3을 해야 실제 맵핑되는 정수
index_to_word = {}
for i,v in words.items():
index_to_word[v+3] = i
# + id="dmhDLGRUrGw1" colab={"base_uri": "https://localhost:8080/", "height": 127} executionInfo={"status": "ok", "timestamp": 1647238316443, "user_tz": -540, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}} outputId="d3dd39fc-caea-48fc-fc1d-778fb6a672f3"
# index to word test
' '.join([index_to_word.get(i,'?') for i in X_train[0]])
# + id="qNyBvOPVrHlT" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1647238318527, "user_tz": -540, "elapsed": 285, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}} outputId="b935d517-5cb1-437b-a596-f6f1635ca48d"
word_lens = pd.Series([len(x) for x in X_train])
print(word_lens.mean())
print(word_lens.median())
print(word_lens.value_counts().max())
# + id="RNgdfZX9rE_S" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1647238320548, "user_tz": -540, "elapsed": 284, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}} outputId="b752524b-1703-434c-aab5-424246c5b5ff"
set(y_train)
# + id="4cJpnO2XnfiV" executionInfo={"status": "ok", "timestamp": 1647238323110, "user_tz": -540, "elapsed": 869, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}}
X_train_with_pad = sequence.pad_sequences(X_train,maxlen=250)
X_test_with_pad = sequence.pad_sequences(X_test,maxlen=250)
# + id="QRWTOVHdngiX" executionInfo={"status": "ok", "timestamp": 1647238324812, "user_tz": -540, "elapsed": 1, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}}
def build_model(model_name):
base = Sequential()
base.add(Embedding(input_dim=10000,output_dim=128))
if model_name == 'LSTM':
base.add(LSTM(units=64,return_sequences = True))
base.add(LSTM(units=64))
elif model_name =='RNN':
base.add(SimpleRNN(units=64,return_sequences=True))
base.add(SimpleRNN(units=64))
elif model_name == 'GRU':
base.add(GRU(units=64,return_sequences = True))
base.add(GRU(units=64))
base.add(Dense(32,activation='relu'))
base.add(Dropout(rate=0.5))
base.add(Dense(1,activation='sigmoid'))
base.compile(optimizer='adam',loss='binary_crossentropy',metrics=['acc'])
base.summary()
return base
# + colab={"base_uri": "https://localhost:8080/"} id="xmgt64i6nhO5" outputId="4d1c39d2-ca52-4b5c-ea86-c19cd3448a20" executionInfo={"status": "ok", "timestamp": 1647244202070, "user_tz": -540, "elapsed": 5873047, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}}
options = {
'x':X_train_with_pad,
'y':y_train,
'batch_size':32,
'epochs':10,
'validation_split':0.1
}
rnn = build_model('RNN').fit(**options)
lstm = build_model('LSTM').fit(**options)
gru = build_model('GRU').fit(**options)
# + id="QWuy706gnis8" executionInfo={"status": "ok", "timestamp": 1647244448801, "user_tz": -540, "elapsed": 317, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}}
def train_val_plot(history,rg):
rng = range(rg)
fig,axes = plt.subplots(nrows=1,ncols=2,figsize=(20,8))
axes[0].plot(rng,history['loss'][:rg],label='train')
axes[0].plot(rng,history['val_loss'][:rg],label='validation')
axes[0].set_title('LOSS')
axes[0].legend()
axes[1].plot(rng,history['acc'][:rg],label='train')
axes[1].plot(rng,history['val_acc'][:rg],label='validation')
axes[1].set_title('ACC')
axes[1].legend()
plt.plot()
# + id="UulPiSnEnkkQ" colab={"base_uri": "https://localhost:8080/", "height": 499} executionInfo={"status": "ok", "timestamp": 1647244451888, "user_tz": -540, "elapsed": 1114, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}} outputId="2d4b0131-59a9-4aa4-bb2a-9e8f8cac9f31"
train_val_plot(rnn.history,10)
# + id="9eUUHkIirMY_" colab={"base_uri": "https://localhost:8080/", "height": 499} executionInfo={"status": "ok", "timestamp": 1647244455993, "user_tz": -540, "elapsed": 1033, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}} outputId="eece2eee-ce62-43dc-9758-a0603e402b42"
train_val_plot(lstm.history,10)
# + id="S5iC-bHrrOsP" colab={"base_uri": "https://localhost:8080/", "height": 499} executionInfo={"status": "ok", "timestamp": 1647244459688, "user_tz": -540, "elapsed": 1152, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14747709315698264138"}} outputId="787f63f4-1220-4023-b852-e17e42a625ad"
train_val_plot(gru.history,10)
|
source/NLP_imdb_movie.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Fake PII data: Exploratory data analysis
#
# This notebook is used to verify the different fake entities before and after the creation of a synthetic dataset / augmented dataset. First part looks at the generation details and stats, second part evaluates the created synthetic dataset after it has been generated.
# +
import pandas as pd
from presidio_evaluator.data_generator.extensions import generate_iban, generate_ip_addresses, generate_SSNs, \
generate_company_names, generate_url, generate_roles, generate_titles, generate_nationality, generate_nation_man, \
generate_nation_woman, generate_nation_plural, generate_title
from presidio_evaluator.data_generator import FakeDataGenerator, read_synth_dataset
from collections import Counter
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# 1. Evaluate generation logic and the fake PII bank used during generation
df = pd.read_csv("../presidio_evaluator/data_generator/raw_data/FakeNameGenerator.com_3000.csv",encoding="utf-8")
generator = FakeDataGenerator(fake_pii_df=df,
templates=None,
dictionary_path=None,
ignore_types={"IP_ADDRESS", 'US_SSN', 'URL','ADDRESS'})
pii_df = generator.prep_fake_pii(df)
for (name, series) in pii_df.iteritems():
print(name)
print("Unique values: {}".format(len(series.unique())))
print(series.value_counts())
print("\n**************\n")
# +
from wordcloud import WordCloud
def series_to_wordcloud(series):
freqs = series.value_counts()
wordcloud = WordCloud(background_color='white',width=800,height=400).generate_from_frequencies(freqs)
fig = plt.figure(figsize=(16, 8))
plt.suptitle("{} word cloud".format(series.name))
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
# -
series_to_wordcloud(pii_df.FIRST_NAME)
series_to_wordcloud(pii_df.LAST_NAME)
series_to_wordcloud(pii_df.COUNTRY)
series_to_wordcloud(pii_df.ORGANIZATION)
series_to_wordcloud(pii_df.CITY)
# 2. Evaluate different entities in the synthetic dataset after creation
synth = read_synth_dataset("../data/generated_train_November 12 2019.json")
sentences_only = [(sample.full_text,sample.metadata) for sample in synth]
sentences_only[2]
print("Proportions of female vs. male based samples:")
Counter([sentence[1]['Gender'] for sentence in sentences_only])
print("Proportion of lower case samples:")
Counter([sentence[1]['Lowercase'] for sentence in sentences_only])
print("Proportion of nameset across samples:")
Counter([sentence[1]['NameSet'] for sentence in sentences_only])
# +
def get_entity_values_from_sample(sample,entity_types):
name_entities = [span.entity_value for span in sample.spans if span.entity_type in entity_types]
return name_entities
names = [get_entity_values_from_sample(sample,['PERSON','FIRST_NAME','LAST_NAME']) for sample in synth]
names = [item for sublist in names for item in sublist]
series_to_wordcloud(pd.Series(names,name='PERSON, FIRST_NAME, LAST_NAME'))
# -
countries = [get_entity_values_from_sample(sample,['LOCATION']) for sample in synth]
countries = [item for sublist in countries for item in sublist]
series_to_wordcloud(pd.Series(countries,name='LOCATION'))
orgs = [get_entity_values_from_sample(sample,['ORGANIZATION']) for sample in synth]
orgs = [item for sublist in orgs for item in sublist]
series_to_wordcloud(pd.Series(orgs,name='ORGANIZATION'))
|
notebooks/PII EDA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
# Licensed under the MIT License.
# 
# # Training Pipeline - Automated ML
# _**Training many models using Automated Machine Learning**_
#
# ---
#
# This notebook demonstrates how to train and register 11,973 models using Automated Machine Learning. We will utilize the [ParallelRunStep](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.parallel_run_step.parallelrunstep?view=azure-ml-py) to parallelize the process of training 11,973 models. For this notebook we are using an orange juice sales dataset to predict the orange juice quantity for each brand and each store. For more information about the data refer to the Data Preparation Notebook.
#
# <span style="color:red"><b>NOTE: There are limits on how many runs we can do in parallel per workspace, and we currently recommend to set the parallelism to maximum of 20 runs per experiment per workspace. If users want to have more parallelism and increase this limit they might encounter Too Many Requests errors (HTTP 429). </b></span>
# <span style="color:red"><b> Please ensure you have the latest version of the SDK to ensure AutoML dependencies are consistent.</b></span>
# +
# #!pip install --upgrade azureml-sdk[automl]
# -
# Also install the pipeline.steps package that is needed for parallel run step
# +
# # !pip install --upgrade azureml-pipeline-steps
# -
# ### Prerequisites
# At this point, you should have already:
#
# 1. Created your AML Workspace using the [00_Setup_AML_Workspace notebook](../../00_Setup_AML_Workspace.ipynb)
# 2. Run [01_Data_Preparation.ipynb](../../01_Data_Preparation.ipynb) to create the dataset
# ## 1.0 Set up workspace, datastore, experiment
# +
import azureml.core
from azureml.core import Workspace, Datastore
import pandas as pd
# set up workspace
ws= Workspace.from_config()
# Take a look at Workspace
ws.get_details()
# set up datastores
dstore = ws.get_default_datastore()
output = {}
output['SDK version'] = azureml.core.VERSION
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Default datastore name'] = dstore.name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
# -
# ### Choose an experiment
# +
from azureml.core import Experiment
experiment = Experiment(ws, 'manymodels-training-pipeline')
print('Experiment name: ' + experiment.name)
# -
# ## 2.0 Call the registered filedataset
# We use 11,973 datasets and ParallelRunStep to build 11,973 time-series to predict the quantity of each store brand.
# Each dataset represents a brand's 2 years orange juice sales data that contains 7 columns and 122 rows.
# You will need to register the datasets in the Workspace first. We did so in the [data preparation notebook](../../01_Data_Preparation.ipynb).
#
# The registered 'oj_data_small_train' file dataset contains the first 10 csv files and 'oj_data_train' contains all 11,973 csv files. You can choose to pass either filedatasets_10_models_input or filedatasets_all_models_inputs in the ParallelRunStep.
#
# We recommend to **start with filedatasets_10_models** and make sure everything runs successfully, then scale up to filedatasets_all_models.
# +
from azureml.core.dataset import Dataset
filedst_10_models = Dataset.get_by_name(ws, name='oj_data_small_train')
filedst_10_models_input = filedst_10_models.as_named_input('train_10_models')
# filedst_all_models = Dataset.get_by_name(ws, name='oj_data_train')
# filedst_all_models_inputs = filedst_all_models.as_named_input('train_all_models')
# -
# ## 3.0 Build the training pipeline
# Now that the dataset, WorkSpace, and datastore are set up, we can put together a pipeline for training.
# ### Set up environment for ParallelRunStep
# [Environment](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.environment.environment?view=azure-ml-py) defines a collection of resources that we will need to run our pipelines. We configure a reproducible Python environment for our training script.
from scripts.helper import get_automl_environment
train_env = get_automl_environment()
# ### Choose a compute target
# Currently ParallelRunConfig only supports AMLCompute. You can change to a different compute cluster if one fails.
#
# This is the compute target we will pass into our ParallelRunConfig.
# +
from azureml.core.compute import ComputeTarget
from azureml.core.compute import AmlCompute
# Choose a name for your cluster.
amlcompute_cluster_name = "cpucluster"
found = False
# Check if this compute target already exists in the workspace.
cts = ws.compute_targets
if amlcompute_cluster_name in cts and cts[amlcompute_cluster_name].type == 'AmlCompute':
found = True
print('Found existing compute target.')
compute = cts[amlcompute_cluster_name]
if not found:
print('Creating a new compute target...')
provisioning_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D13_V2',
min_nodes=2,
max_nodes=20)
# Create the cluster.
compute = ComputeTarget.create(ws, amlcompute_cluster_name, provisioning_config)
print('Checking cluster status...')
# Can poll for a minimum number of nodes and for a specific timeout.
# If no min_node_count is provided, it will use the scale settings for the cluster.
compute.wait_for_completion(show_output = True, min_node_count = None, timeout_in_minutes = 20)
# For a more detailed view of current AmlCompute status, use get_status().
# -
# ## Train
#
# This dictionary defines the [AutoML settings](https://docs.microsoft.com/en-us/python/api/azureml-train-automl-client/azureml.train.automl.automlconfig.automlconfig?view=azure-ml-py#parameters), for this forecasting task we add the name of the time column and the maximum forecast horizon.
#
# |Property|Description|
# |-|-|
# |**task**|forecasting|
# |**primary_metric**|This is the metric that you want to optimize.<br> Forecasting supports the following primary metrics <br><i>spearman_correlation</i><br><i>normalized_root_mean_squared_error</i><br><i>r2_score</i><br><i>normalized_mean_absolute_error</i>|
# |**blacklist_models**|Models in blacklist won't be used by AutoML. All supported models can be found at [here](https://docs.microsoft.com/en-us/python/api/azureml-train-automl-client/azureml.train.automl.constants.supportedmodels.forecasting?view=azure-ml-py).|
# |**iterations**|Number of models to train. This is optional but provides customer with greater control.|
# |**iteration_timeout_minutes**|Maximum amount of time in minutes that the model can train. This is optional and depends on the dataset. We ask customer to explore a bit to get approximate times for training the dataset. For OJ dataset we set it 20 minutes|
# |**experiment_timeout_hours**|Maximum amount of time in hours that the experiment can take before it terminates.|
# |**label_column_name**|The name of the label column.|
# |**n_cross_validations**|Number of cross validation splits. Rolling Origin Validation is used to split time-series in a temporally consistent way.|
# |**enable_early_stopping**|Flag to enable early termination if the score is not improving in the short term.|
# |**time_column_name**|The name of your time column.|
# |**max_horizon**|The number of periods out you would like to predict past your training data. Periods are inferred from your data.|
# |**grain_column_names**|The column names used to uniquely identify timeseries in data that has multiple rows with the same timestamp.|
# |**group_column_names**|The names of columns used to group your models. For timeseries, the groups must not split up individual time-series. That is, each group must contain one or more whole time-series.|
# |**drop_column_names**|The names of columns to drop for forecasting tasks.|
# |**track_child_runs**|Flag to disable tracking of child runs. Only best run (metrics and model) is tracked if the flag is set to False.|
# +
import logging
from scripts.helper import write_automl_settings_to_file
automl_settings = {
"task" : 'forecasting',
"primary_metric" : 'normalized_root_mean_squared_error',
"iteration_timeout_minutes" : 10, # This needs to be changed based on the dataset. We ask customer to explore how long training is taking before settings this value
"iterations" : 15,
"experiment_timeout_hours" : 1,
"label_column_name" : 'Quantity',
"n_cross_validations" : 3,
"verbosity" : logging.INFO,
"debug_log": 'automl_oj_sales_debug.txt',
"time_column_name": 'WeekStarting',
"max_horizon" : 20,
"group_column_names": ['Store', 'Brand'],
"grain_column_names": ['Store', 'Brand'],
"drop_column_names": ['Revenue']
}
write_automl_settings_to_file(automl_settings)
# -
# ### Set up ParallelRunConfig
# [ParallelRunConfig](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.parallel_run_config.parallelrunconfig) is configuration for parallel run step. You will need to determine the number of workers and nodes appropriate for your use case. The process_count_per_node is based off the number of cores of the compute VM. The node_count will determine the number of master nodes to use, increasing the node count will speed up the training process.
#
#
# * <b>node_count</b>: The number of compute nodes to be used for running the user script. We recommend to start with 3 and increase the node_count if the training time is taking too long.
#
# * <b>process_count_per_node</b>: The number of processes per node.
#
# * <b>run_invocation_timeout</b>: The run() method invocation timeout in seconds. The timeout should be set to maximum training time of one AutoML run(with some buffer), by default it's 60 seconds.
#
# <span style="color:red"><b>NOTE: There are limits on how many runs we can do in parallel per workspace, and we currently recommend to set the parallelism to maximum of 20 runs per experiment per workspace. If users want to have more parallelism and increase this limit they might encounter Too Many Requests errors (HTTP 429). </b></span>
#
# +
from scripts.helper import build_parallel_run_config
# PLEASE MODIFY the following three settings based on your compute and experiment timeout.
node_count=2
process_count_per_node=6
run_invocation_timeout=3700 # this timeout(in seconds) is inline with AutoML experiment timeout or (no of iterations * iteration timeout)
parallel_run_config = build_parallel_run_config(train_env, compute, node_count, process_count_per_node, run_invocation_timeout)
# -
# ### Set up ParallelRunStep
# This [ParallelRunStep](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.parallelrunstep?view=azure-ml-py) is the main step in our pipeline. First, we set up the output directory and define the Pipeline's output name. The datastore that stores the pipeline's output data is Workspace's default datastore.
# +
from azureml.pipeline.core import PipelineData
training_output_name = "training_output"
output_dir = PipelineData(name=training_output_name,
datastore=dstore)
# -
# We specify the following parameters:
#
# * <b>name</b>: We set a name for our ParallelRunStep.
#
# * <b>parallel_run_config</b>: We then pass the previously defined ParallelRunConfig.
#
# * <b>allow_reuse</b>: Indicates whether the step should reuse previous results when re-run with the same settings.
#
# * <b>inputs</b>: We are going to use the registered FileDataset that we called earlier in the Notebook. _inputs_ points to a registered file dataset in AML studio that points to a path in the blob container. The number of files in that path determines the number of models will be trained in the ParallelRunStep.
#
# * <b>output</b>: The output directory we just defined. A PipelineData object that corresponds to the output directory.
#
# * <b>models</b>: Zero or more model names already registered in the Azure Machine Learning model registry.
#
# <span style="color:red"><b>Please upgrade azureml-pipeline-steps(>=1.6.0) if the following fails.</b></span>
# +
from azureml.pipeline.steps import ParallelRunStep
parallel_run_step = ParallelRunStep(
name="many-models-training",
parallel_run_config=parallel_run_config,
allow_reuse = False,
inputs=[filedst_10_models_input], # train 10 models
#inputs=[filedst_all_models_inputs], # switch to this inputs if train all 11,973 models
output=output_dir,
#arguments=['--retrain_failed_models', 'True'], # Uncomment this if you want to retrain only failed models
)
# -
# ## 4.0 Run the training pipeline
# ### Submit the pipeline to run
# Next we submit our pipeline to run. The whole training pipeline takes about 1h 11m using a Standard_D13_V2 VM with our current ParallelRunConfig setting.
# +
from azureml.pipeline.core import Pipeline
#from azureml.widgets import RunDetails
pipeline = Pipeline(workspace=ws, steps=parallel_run_step)
run = experiment.submit(pipeline)
#RunDetails(run).show()
# -
# You can run the folowing command if you'd like to monitor the training process in jupyter notebook. It will stream logs live while training.
#
# **Note**: This command may not work for Notebook VM, however it should work on your local laptop.
run.wait_for_completion(show_output=True)
# Succesfully trained, registered Automated ML models.
# ## 5.0 Review outputs of the training pipeline
# The training pipeline will train and register models to the Workspace. You can review trained models in the Azure Machine Learning Studio under 'Models'.
# If there are any issues with training, you can go to 'many-models-training' run under the pipeline run and explore logs under 'Logs'.
# You can look at the stdout and stderr output under logs/user/worker/<ip> for more details
#
# ## 6.0 Get list of AutoML runs along with registered model names and tags
# The following code snippet will iterate through all the automl runs for the experiment and list the details.
#
# **Framework** - AutoML, **Dataset** - input data set, **Run** - AutoML run id, **Status** - AutoML run status, **Model** - Registered model name, **Tags** - Tags for model, **StartTime** - Start time, **EndTime** - End time, **ErrorType** - ErrorType, **ErrorCode** - ErrorCode, **ErrorMessage** - Error Message
# +
from scripts.helper import get_training_output
import os
training_results_name = "training_results"
training_file = get_training_output(run, training_results_name, training_output_name)
all_columns = ["Framework", "Dataset", "Run", "Status", "Model", "Tags", "StartTime", "EndTime" , "ErrorType", "ErrorCode", "ErrorMessage" ]
df = pd.read_csv(training_file, delimiter=" ", header=None, names=all_columns)
training_csv_file = "training.csv"
df.to_csv(training_csv_file)
print("Training output has", df.shape[0], "rows. Please open", os.path.abspath(training_csv_file), "to browse through all the output.")
# -
# ## 7.0 Publish and schedule the pipeline (Optional)
# ### 7.1 Publish the pipeline
#
# Once you have a pipeline you're happy with, you can publish a pipeline so you can call it programmatically later on. See this [tutorial](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-create-your-first-pipeline#publish-a-pipeline) for additional information on publishing and calling pipelines.
# +
# published_pipeline = pipeline.publish(name = 'automl_train_many_models',
# description = 'train many models',
# version = '1',
# continue_on_step_failure = False)
# -
# ### 7.2 Schedule the pipeline
# You can also [schedule the pipeline](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-schedule-pipelines) to run on a time-based or change-based schedule. This could be used to automatically retrain models every month or based on another trigger such as data drift.
# +
# from azureml.pipeline.core import Schedule, ScheduleRecurrence
# training_pipeline_id = published_pipeline.id
# recurrence = ScheduleRecurrence(frequency="Month", interval=1, start_time="2020-01-01T09:00:00")
# recurring_schedule = Schedule.create(ws, name="automl_training_recurring_schedule",
# description="Schedule Training Pipeline to run on the first day of every month",
# pipeline_id=training_pipeline_id,
# experiment_name=experiment.name,
# recurrence=recurrence)
# -
# ## 8.0 Bookkeeping of workspace (Optional)
# ### 8.1 Cancel any runs that are running
#
# To cancel any runs that are still running in a given experiment.
# +
# from scripts.helper import cancel_runs_in_experiment
# failed_experiment = 'Please modify this and enter the experiment name'
# # Please note that the following script cancels all the currently running runs in the experiment
# cancel_runs_in_experiment(ws, failed_experiment)
# -
# ## Next Steps
#
# Now that you've trained and scored the models, move on to [03_AutoML_Forecasting_Pipeline.ipynb](../03_AutoML_Forecasting_Pipeline/03_AutoML_Forecasting_Pipeline.ipynb) to make forecasts with your models.
|
Automated_ML/02_AutoML_Training_Pipeline/02_AutoML_Training_Pipeline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from pathlib import Path
testfolder = Path().resolve().parent.parent / 'bifacial_radiance' / 'TEMP'
# Another option using relative address; for some operative systems you might need '/' instead of '\'
# testfolder = os.path.abspath(r'..\..\bifacial_radiance\TEMP')
print ("Your simulation will be stored in %s" % testfolder)
# -
from bifacial_radiance import *
demo = RadianceObj('bifacial_example',str(testfolder))
demo.setGround(0.30) # This prints available materials.
epwfile = demo.getEPW(lat = 37.5, lon = -77.6) # This location corresponds to Richmond, VA.
metdata = demo.readWeatherFile(epwfile)
demo.gendaylit(metdata,8) # Noon, June 17th (timepoint # 4020)\
# +
module_type = 'Bi60'
numcellsx = 6
numcellsy = 12
xcell = 0.156
ycell = 0.156
xcellgap = 0.02
ycellgap = 0.02
torquetube = True
diameter = 0.15
tubetype = 'round'
material = 'Metal_Grey'
xgap = 0.1
ygap = 0
zgap = 0.05
numpanels = 1
axisofrotationTorqueTube = False
glass = True
cellLevelModuleParams = {'numcellsx': numcellsx, 'numcellsy':numcellsy,
'xcell': xcell, 'ycell': ycell, 'xcellgap': xcellgap, 'ycellgap': ycellgap}
mymodule = demo.makeModule(name=module_type, torquetube=torquetube, diameter=diameter, tubetype=tubetype, material=material,
xgap=xgap, ygap=ygap, zgap=zgap, numpanels=numpanels,
cellLevelModuleParams=cellLevelModuleParams,
axisofrotationTorqueTube=axisofrotationTorqueTube, glass=glass, z=0.0002)
sceneDict = {'tilt':25,'pitch':5.5,'hub_height':1.0,'azimuth':90, 'nMods': 20, 'nRows': 1, originx=0, originy=0}
scene = demo.makeScene(module_type,sceneDict)
octfile = demo.makeOct(demo.getfilelist())
# -
sceneDict = {'tilt':25,'pitch':5.5,'hub_height':1.0,'azimuth':90, 'nMods': 20, 'nRows': 10, 'originx':0, 'originy':0}
scene = demo.makeScene(module_type,sceneDict)
octfile = demo.makeOct(demo.getfilelist())
# + active=""
# Advanced Rendering:
#
# My workflow for going from oct file to png is:
# rvu -> rpict -> pcond -> pfilt -> ra_tiff -> convert
# In detail:
# 1. Use rvu to view the oct file
# rvu 1axis_07_01_08.oct
# use aim and origin to move around, zoom in/out, etc
# save a view file with view render.vf
# 2. Run rpict to render the image to hdr. This is the time consuming step. It takes between 1 and 3 hours depending on how complex the geometry is.
# rpict -x 4800 -y 4800 -i -ps 1 -dp 530 -ar 964 -ds 0.016 -dj 1 -dt 0.03 -dc 0.9 -dr 5 -st 0.12 -ab 5 -aa 0.11 -ad 5800 -as 5800 -av 25 25 25 -lr 14 -lw 0.0002 -vf render.vf bifacial_example.oct > render.hdr
# 3. Run pcond to mimic human visual response:
# pcond -h render.hdr > render.pcond.hdr
# 4. Resize and adjust exposure with pfilt
# pfilt -e +0.2 -x /4 -y /4 render.pcond.hdr > render.pcond.pfilt.hdr
# 5. Convert hdr to tif
# ra_tiff render.pcond.pfilt.hdr render.tif
# 6. Convert tif to png with imagemagick convert utility
# convert render.tif render.png
# 7. Annotate the image with convert
# convert render.png -fill black -gravity South -annotate +0+5 'Created with NREL bifacial_radiance https://github.com/NREL/bifacial_radiance' render_annotated.png
#
# -
analysis = AnalysisObj(octfile, demo.basename)
frontscan, backscan = analysis.moduleAnalysis(scene)
results = analysis.analysis(octfile, demo.basename, frontscan, backscan)
load.read1Result('results\irr_bifacial_example.csv')
frontscan, backscan = analysis.moduleAnalysis(scene, frontsurfaceoffset=0.02, backsurfaceoffset = 0.02)
results = analysis.analysis(octfile, demo.basename, frontscan, backscan)
load.read1Result('results\irr_bifacial_example.csv')
|
docs/tutorials/(development) Glass Test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python3.8
# ---
# # Azure ML Compute Python SDK
#
# description: overview of the AML Compute Python SDK
# + tags=["create workspace"]
from azureml.core import Workspace
ws = Workspace.from_config()
ws
# -
# ## Introduction to AmlCompute
#
# Azure Machine Learning Compute is managed compute infrastructure that allows the user to easily create single to multi-node compute of the appropriate VM Family. It is created **within your workspace region** and is a resource that can be used by other users in your workspace. It autoscales by default to the max_nodes, when a job is submitted, and executes in a containerized environment packaging the dependencies as specified by the user.
#
# Since it is managed compute, job scheduling and cluster management are handled internally by Azure Machine Learning service.
#
# For more information on Azure Machine Learning Compute, please read [this article](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)
#
# **Note**: As with other Azure services, there are limits on certain resources (for eg. AmlCompute quota) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
# +
from azureml.core.compute import ComputeTarget, AmlCompute
AmlCompute.supported_vmsizes(workspace=ws)
# AmlCompute.supported_vmsizes(workspace = ws, location='southcentralus')
# -
# ### Provision as a persistent compute target (Basic)
#
# You can provision a persistent AmlCompute resource by simply defining two parameters thanks to smart defaults. By default it autoscales from 0 nodes and provisions dedicated VMs to run your job in a container. This is useful when you want to continously re-use the same target, debug it between jobs or simply share the resource with other users of your workspace.
#
# * `vm_size`: VM family of the nodes provisioned by AmlCompute. Simply choose from the supported_vmsizes() above
# * `max_nodes`: Maximum nodes to autoscale to while running a job on AmlCompute
#
# You can also specify additional properties or change defaults while provisioning AmlCompute using a more advanced configuration. This is useful when you want a dedicated cluster of 4 nodes (for example you can set the min_nodes and max_nodes to 4), or want the compute to be within an existing VNet in your subscription.
#
# In addition to `vm_size` and `max_nodes`, you can specify:
# * `min_nodes`: Minimum nodes (default 0 nodes) to downscale to while running a job on AmlCompute
# * `vm_priority`: Choose between 'dedicated' (default) and 'lowpriority' VMs when provisioning AmlCompute. Low Priority VMs use Azure's excess capacity and are thus cheaper but risk your run being pre-empted
# * `idle_seconds_before_scaledown`: Idle time (default 120 seconds) to wait after run completion before auto-scaling to min_nodes
# * `vnet_resourcegroup_name`: Resource group of the **existing** VNet within which AmlCompute should be provisioned
# * `vnet_name`: Name of VNet
# * `subnet_name`: Name of SubNet within the VNet
# * `admin_username`: Name of Admin user account which will be created on all the nodes of the cluster
# * `admin_user_password`: Password that you want to set for the user account above
# * `admin_user_ssh_key`: SSH Key for the user account above. You can specify either a password or an SSH key or both
# * `remote_login_port_public_access`: Flag to enable or disable the public SSH port. If you dont specify, AmlCompute will smartly close the port when deploying inside a VNet
# * `identity_type`: Compute Identity type that you want to set on the cluster, which can either be SystemAssigned or UserAssigned
# * `identity_id`: Resource ID of identity in case it is a UserAssigned identity, optional otherwise
#
# + tags=[]
from random import randint
from azureml.core.compute import AmlCompute, ComputeTarget
# name must be unique within a workspace
ct_name = f"ct-{str(randint(10000, 99999))}-concept"
if ct_name in ws.compute_targets:
ct = ws.compute_targets[ct_name]
ct.delete()
ct.wait_for_completion(show_output=True, is_delete_operation=True)
compute_config = AmlCompute.provisioning_configuration(
vm_size="STANDARD_D2_V2", max_nodes=4
)
ct = ComputeTarget.create(ws, ct_name, compute_config)
ct.wait_for_completion(show_output=True)
# -
# get_status() gets the latest status of the AmlCompute target
ct.get_status().serialize()
# list_nodes() gets the list of nodes on the cluster with status, IP and associated run
ct.list_nodes()
# +
# update() takes in the min_nodes, max_nodes and idle_seconds_before_scaledown and updates the AmlCompute target
# ct.update(min_nodes=1)
# ct.update(max_nodes=10)
# ct.update(idle_seconds_before_scaledown=300)
# ct.update(min_nodes=2, max_nodes=4, idle_seconds_before_scaledown=600)
# -
# delete() is used to deprovision and delete the AmlCompute target. Useful if you want to re-use the compute name
ct.delete()
ct.wait_for_completion(show_output=True, is_delete_operation=True)
|
concepts/compute/azureml-compute-api.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/xvdp/nerf/blob/master/tiny_nerf_pytorch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="UtwhxHG5dD_H"
# # Tiny NeRF, a Pytorch translation
#
# Step by step translation to pytorch
# ## differences between tf and pytorch that need to be minded
# * tf.math.cumprod(exclusive=True) # not existent in pytorch, need to zero and roll
# * i,j=tf.meshgrid('xy') -> j,i=torch.meshgrid()
# * Keras Dense and torch.nn.Linear are similar with transposed weights, by default initializes w as glorot uniform and b to zero.
# * pytorch does not do automatic gpu conversion
#
#
# This is a simplied version of the method presented in *NeRF: Representing Scenes as Neural Radiance Fields for View Synthesis*
#
# [Project Website](http://www.matthewtancik.com/nerf)
#
# [arXiv Paper](https://arxiv.org/abs/2003.08934)
#
# [Full Code](github.com/bmild/nerf)
#
# Components not included in the notebook
#
# 5D input including view directions
# Hierarchical Sampling
# + colab={"base_uri": "https://localhost:8080/"} id="apbXlzVbc4E_" outputId="6c52d9a1-ca4b-43ed-acec-89ec19ed5a7c"
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
if IN_COLAB:
# %tensorflow_version 1.x
import os, sys
import tensorflow as tf
tf.compat.v1.enable_eager_execution()
from tqdm import tqdm_notebook as tqdm
import numpy as np
import matplotlib.pyplot as plt
# + id="PWGsTSRveSge"
#xvdp
import torch
import torch.nn as nn
safelog10 = lambda x: 0 if not x else np.log10(np.abs(x))
sround = lambda x, d=1: np.round(x, max((-np.floor(safelog10(x)).astype(int) + d), 0))
# + id="fd78sD6nd-Gw"
if not os.path.exists('tiny_nerf_data.npz'):
# !wget https://people.eecs.berkeley.edu/~bmild/nerf/tiny_nerf_data.npz
# + [markdown] id="YHaK0qRtec4p"
# ## Load Input Images and Poses
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="FEFbWmjXecME" outputId="c404b068-c4f7-4550-9a29-5eae7578b402"
data = np.load('tiny_nerf_data.npz')
images = data['images']
poses = data['poses']
focal = data['focal']
H, W = images.shape[1:3]
print(images.shape, poses.shape, focal)
testimg, testpose = images[101], poses[101]
images = images[:100,...,:3]
poses = poses[:100]
plt.imshow(testimg)
plt.show()
# + id="KZyi0zLhe5Ji"
def posenc(x, L_embed=6):
"""no difference with tf"""
rets = [x]
for i in range(L_embed):
for fn in [torch.sin, torch.cos]:
rets.append(fn(2.**i * x))
return torch.cat(rets, -1)
L_embed = 6
embed_fn = posenc
def get_rays(H, W, focal, c2w):
"""meshgroid: note instead of i,j -> j, i"""
c2wt = torch.as_tensor(c2w)
jt, it = torch.meshgrid(torch.arange(W, dtype=torch.float32), torch.arange(H, dtype=torch.float32))
dirst = torch.stack([(it-W*.5)/focal, -(jt-H*.5)/focal, -torch.ones_like(it)], -1)
rays_dt = torch.sum(dirst.view(*dirst.shape[:2], 1, -1) * c2wt[:3,:3], -1)
rays_ot = c2wt[:3,-1] * torch.ones_like(rays_dt)
return rays_ot, rays_dt
# + id="ae-z0cQJfF4t"
class nerf(nn.Module):
"""
"""
def __init__(self, D=8, W=256, L_embed=6):
super(nerf, self).__init__()
channels = 3 + 3*2*L_embed
self.lin0 = nn.Linear(in_features=channels, out_features=W)
self.relu = nn.ReLU()
block0 = []
for i in range(1,5):
block0 += [nn.Linear(W, W), self.relu]
self.block0 = nn.Sequential(*block0)
block1 = [nn.Linear(W+channels, W), self.relu]
for i in range(2,4):
block1 += [nn.Linear(W, W), self.relu]
self.block1 = nn.Sequential(*block1)
self.fc = nn.Linear(W,4)
# initialization needs to be explicit, pytorch default differs from keras
for m in self.modules():
if isinstance(m, nn.Linear):
nn.init.constant_(m.bias, 0)
nn.init.xavier_uniform_(m.weight, gain=1.0)
def forward(self, x):
x = torch.cat([self.block0(self.relu(self.lin0(x))), x], -1)
return self.fc(self.block1(x))
# + id="p4Z4vXHqfzWQ"
def exclusive_cumprod(x):
"""torch cumprod does not have a exclusive=True option
concatenate ones and roll last value out.
"""
dim=-1
size = list(x.shape)#
size[dim] = 1
ones = torch.ones(size, dtype=x.dtype, layout=x.layout, device=x.device)
return torch.cat([ones, torch.cumprod(x, -1)[...,:-1]], dim=dim)
# + id="hNXssRYCgpwz"
def render_rays(network_fn, rays_o, rays_d, near, far, N_samples, rand=False, device="cuda"):
# sample template
z_vals = torch.linspace(near, far, N_samples)
# jitter sample for each ray
if rand:
z_vals = z_vals + torch.rand(list(rays_o.shape[:-1]) + [N_samples]) * (far-near)/N_samples
# project directions to global jittered samples
pts = rays_o[...,None,:] + rays_d[...,None,:] * z_vals[...,:,None]
# embed, global x,y,z samples per ray to sin(x), cos(x), sin(2x), cos(2x), ...cos(x*2**i), sin(y)...
pts_flat = embed_fn(pts.reshape(-1,3)).to(device=device)
# run model in batches
step = 32*1024
raw = torch.cat([network_fn(pts_flat[i:i+step])
for i in range(0, len(pts_flat), step)]).view(list(pts.shape[:-1]) + [4])
# Compute opacities and colors, model outputs rgba predictions
sigma_a = nn.ReLU()(raw[...,3])
rgb = nn.Sigmoid()(raw[...,:3])
# Do volume rendering
# depth steps per ray
dists = torch.cat([z_vals[..., 1:] - z_vals[..., :-1],
torch.full(z_vals[...,:1].shape, 1e10)], -1).to(device=device)
# alpha values
alpha = 1.-torch.exp(-sigma_a * dists)
weights = alpha * exclusive_cumprod(1.-alpha + 1e-10)
# sum up rgb values along ray
rgb_map = torch.sum(weights[...,None] * rgb, -2)
depth_map = torch.sum(weights * z_vals.to(device=device), -1)
acc_map = torch.sum(weights, -1)
return rgb_map, depth_map, acc_map
# + id="UkNc9K0uh87u"
device = "cuda"
model = nerf(D=8, W=256, L_embed=6)
N_samples = 64
model.to(device=device)
optimizer = torch.optim.Adam(model.parameters(), lr=5e-4)
N_iters = 1000
psnrs = []
iternums = []
losses = []
i_plot = 100
import time
# + [markdown] id="Mx9PMfLmiNra"
# ## simplified training loop
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="0TMCFbariJwL" outputId="cacc8d28-ded8-49d5-d1e7-552a2af2d546"
t = time.time()
for i in range(1000):
img_i = np.random.randint(images.shape[0])
target = torch.tensor(images[img_i]).to(device=device)
pose = torch.as_tensor(poses[img_i])
rays_o, rays_d = get_rays(H, W, focal, pose)
rgb, depth, acc = render_rays(model, rays_o, rays_d, near=2., far=6., N_samples=N_samples, rand=True,
device=device)
optimizer.zero_grad()
loss = ((rgb - target)**2).mean()
loss.backward()
optimizer.step()
print(i, sround(loss.item(),2), " ", end="\r")
if i%i_plot == 0:
with torch.no_grad():
rays_o, rays_d = get_rays(H, W, focal, torch.as_tensor(testpose))
rgb, depth, acc = render_rays(model, rays_o, rays_d, near=2., far=6., N_samples=N_samples)
rgb = rgb.cpu().clone().detach()
losses.append(((rgb - torch.tensor(testimg))**2).mean())
iternums.append(i)
plt.figure(figsize=(15,5))
plt.subplot(131)
plt.title(f"iter {i} loss {sround(loss.item(),2)} time {int(time.time()-t)}s")
plt.imshow(rgb.numpy())
plt.axis("off")
plt.subplot(132)
plt.title("depth")
plt.imshow(depth.cpu().clone().detach().numpy())
plt.axis("off")
plt.subplot(133)
plt.title("acc")
plt.imshow(acc.cpu().clone().detach().numpy())
plt.axis("off")
plt.tight_layout()
plt.show()
plt.figure(figsize=(15,5))
plt.subplot(121)
plt.title("loss")
plt.plot(iternums, losses)
plt.yscale("log")
plt.grid()
plt.subplot(122)
plt.title("psrn")
plt.plot(-10. *np.log10(np.array([l.item() for l in losses])))
plt.grid()
plt.show()
# + [markdown] id="nIVoLRbBxKkI"
# ## Inspect rendering
# + id="KD0AsfkDxqVv"
def depth_samples(near, far, N_samples, rand=True, verbose=False):
# sample template
z_vals = torch.linspace(near, far, N_samples)
# jitter sample for each ray
if rand:
z_vals = z_vals + torch.rand(list(rays_o.shape[:-1]) + [N_samples]) * (far-near)/N_samples
if verbose:
print("jittered depth sampling, z_vals\n", z_vals)
return z_vals
def project_samples(origin, direction, samples, verbose=False):
pts = origin[...,None,:] + direction[...,None,:] * samples[...,:,None]
if verbose:
print("projected directions\n", pts.shape)
return pts
def render_rays_verb(network_fn, rays_o, rays_d, near, far, N_samples, rand=False, device="cuda",verbose=[0,0,1]):
z_vals = depth_samples(near, far, N_samples, rand=rand, verbose=verbose[0])
# global jittered samples
pts = project_samples(rays_o, rays_d, z_vals, verbose=verbose[1])
# embed, global x,y,z samples per ray to sin(x), cos(x), sin(2x), cos(2x), ...cos(x*2**i), sin(y)...
pts_flat = embed_fn(pts.reshape(-1,3)).to(device=device)
if verbose[2]:
print("embeded samples\n", pts_flat)
# run model in batches
step = 32*1024
raw = torch.cat([network_fn(pts_flat[i:i+step])
for i in range(0, len(pts_flat), step)]).view(list(pts.shape[:-1]) + [4])
# Compute opacities and colors, model outputs rgba predictions
sigma_a = nn.ReLU()(raw[...,3])
rgb = nn.Sigmoid()(raw[...,:3])
# Do volume rendering
# depth steps per ray
dists = torch.cat([z_vals[..., 1:] - z_vals[..., :-1],
torch.full(z_vals[...,:1].shape, 1e10)], -1).to(device=device)
# alpha values
alpha = 1.-torch.exp(-sigma_a * dists)
weights = alpha * exclusive_cumprod(1.-alpha + 1e-10)
# sum up rgb values along ray
rgb_map = torch.sum(weights[...,None] * rgb, -2)
depth_map = torch.sum(weights * z_vals.to(device=device), -1)
acc_map = torch.sum(weights, -1)
return rgb_map, depth_map, acc_map
# + colab={"base_uri": "https://localhost:8080/"} id="th3pJOlCxbow" outputId="38b585e4-cf2e-4feb-8dbc-e586c5fc4c69"
rays_o, rays_d = get_rays(H, W, focal, torch.as_tensor(testpose))
rays_o.shape, rays_d.shape
# + [markdown] id="FM-jXn-zyI3O"
# ## compare single rays
# + colab={"base_uri": "https://localhost:8080/"} id="lsFa0hxxxzGA" outputId="5a0db8b1-384b-47b1-f72a-219b8b7a00e9"
## origin is the same for all rays
rayo = rays_o[:1, :1, :]
rayd0 = rays_d[:1, :1, :]
rayd1 = rays_d[50:51, 50:51, :]
rayo, rayd0, rayd1
# + colab={"base_uri": "https://localhost:8080/"} id="5M26zb1Ew0JW" outputId="d6575ed9-839e-494b-e747-c81c15435335"
rayd = rayd1
print("from", rayo, "along", rayd)
with torch.no_grad():
rgb0, depth0, acc0 = render_rays_verb(model, rayo, rayd, near=2., far=6., N_samples=N_samples, verbose=[0,1,1])
# + id="NHjgzUbX6hfw"
z_vals = depth_samples(near=2., far=6., N_samples=N_samples, rand=True, verbose=False)
# global jittered samples
pts = project_samples(rayo, rayd, z_vals, verbose=False)
# + colab={"base_uri": "https://localhost:8080/"} id="7JWHTxM878nT" outputId="f9b122ff-6eaf-40c9-a2b0-f43ee46dd0a1"
z_vals = depth_samples(near=2., far=6., N_samples=N_samples, rand=True, verbose=False)
print(z_vals.shape)
z_vals= z_vals[[0,50,99], 50:51, :]
rayd2 = rays_d[[0,50,99], 50:51, :]
rayo2 = rays_o[[0,50,99], 50:51, :]
pts = project_samples(rayo2, rayd2, z_vals, verbose=False)
raytext = ["0,50", "50,50", "99,50"]
pts.shape
# + colab={"base_uri": "https://localhost:8080/"} id="KlhSUNWU7VE4" outputId="d4a03883-b6d2-4afe-8a1f-88c00fe94407"
# ! pip install plotly
# + colab={"base_uri": "https://localhost:8080/"} id="2y6OKKEp7oPm" outputId="16a80d48-cea4-4e0d-ae82-c0f3c952c5b4"
pts.shape
# + colab={"base_uri": "https://localhost:8080/"} id="-L-P48rYEmSA" outputId="7f407e36-7e5b-4b62-ab0f-f6d931ac8dae"
pts_flat = embed_fn(pts.reshape(-1,3))#.to(device=device)
pts_flat.shape
# + [markdown] id="nNFFyosiJpbi"
# # Visualize sampling of ray subset
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="vBDt01SM3Pmz" outputId="a9521eef-ede2-4ae3-8cf2-2b70721f6abc"
import plotly.graph_objs as go
def scatter_points(points, size=2, text=""):
return go.Scatter3d(
x=points[:, 0],
y=points[:, 1],
z=points[:, 2],
mode='markers+text',
marker=dict(size=size),
text=text,
textposition="top center"
)
data= [scatter_points(rays_o[0,0].cpu().detach().view(-1,3).numpy(), size=10, text=["camera"])]
data += [ scatter_points(pt.cpu().detach().view(-1,3).numpy(), text=[f"ray[{raytext[i]}]"]) for i, pt in enumerate(pts)]
fig = go.Figure(data)
fig.update_layout(scene_dragmode='orbit', title='Depth Sampling in NeRF')
fig.show()
# + [markdown] id="7fC2CnlsJxyW"
# # Visualize positional embedding
# + colab={"base_uri": "https://localhost:8080/"} id="YqKFSEx0FBcl" outputId="f90c322f-03da-4e3a-fa9f-54811bde5d21"
pts_flat0 = embed_fn(pts[0].reshape(-1,3))#.to(device=device)
print("sample ray", tuple(pts[0].shape))
print("embedded ray", tuple(pts_flat0.shape))
print("flat points: x, sin(x*2**0),...,sin(x*2**5): 6*2*3", 6*2*3 + 3)
# + colab={"base_uri": "https://localhost:8080/"} id="5gmir0KvGlNt" outputId="3b7b37d9-fa7d-4fc1-813f-dfea6b224b23"
print("ray, x", pts[0][...,0])
# + colab={"base_uri": "https://localhost:8080/", "height": 334} id="h_itaKXVGy6F" outputId="7c20bf84-8756-4a31-f695-ff9cdb8e69c9"
def plot_ray(x, name="x"):
plt.plot(x, label=f"jittered sampled ray {name}", linewidth="5")
plt.plot(np.sin(x), label="ray sin(x)", linewidth="2")
plt.plot(np.cos(x), label="ray cos(x)", linewidth="2")
plt.plot(np.sin(2*x), label="ray sin(2x)", linewidth="1")
plt.plot(np.cos(2*x), label="ray cos(2x)", linewidth="1")
plt.plot(np.sin(5*x), label="ray sin(5x)", linewidth="0.5")
plt.plot(np.cos(5*x), label="ray cos(5x)", linewidth="0.5")
plt.xlabel("samples")
plt.ylabel(name)
plt.grid()
plt.legend()
plt.figure(figsize=(20,5))
plt.subplot(131)
x = pts[0][...,0].view(-1).clone().detach().numpy()
plot_ray(x)
plt.subplot(132)
y = pts[0][...,1].view(-1).clone().detach().numpy()
plot_ray(y, "y")
plt.subplot(133)
z = pts[0][...,2].view(-1).clone().detach().numpy()
plot_ray(z, "z")
plt.show()
# + [markdown] id="gYJGmFZMJ5af"
# # render 3 pixels, 2 corners (transparent), center (yellow)
# + colab={"base_uri": "https://localhost:8080/"} id="jAg9wYFAKITN" outputId="90d855fb-38af-4010-82f1-fbde0940c4dd"
# network inputs, 64 samples, 3 channels + 6 sin channels + 6 cos channels
pts_flat.shape
# + id="yo1Dz0YnJ4Nf"
# run model on 3 rays
raw = model(pts_flat.cuda()).view(list(pts.shape[:-1]) + [4])
# Compute opacities and colors, model outputs rgba predictions
sigma_a = nn.ReLU()(raw[...,3])
rgb = nn.Sigmoid()(raw[...,:3])
# + colab={"base_uri": "https://localhost:8080/"} id="s5oGx9rRKzJr" outputId="031dd794-f81e-434b-c98d-d216b37f96bf"
sigma_a.shape, rgb.shape
# + [markdown] id="Lt9xR3p6MF_z"
# ## model result: alphas are clamped, 0 = translarent
# + colab={"base_uri": "https://localhost:8080/"} id="tK-C1_jEQdI7" outputId="a3c37445-ccd6-4f17-8e2b-97b57494fdfe"
dists = torch.cat([z_vals[..., 1:] - z_vals[..., :-1],
torch.full(z_vals[...,:1].shape, 1e10)], -1)#.to(device=device)
dists.device
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="2SK4Bt5uK75L" outputId="b542a7e2-d64e-4cce-8c78-c394a9f19fac"
sigmas = sigma_a.cpu().clone().detach()
alphas = 1.-torch.exp(-sigmas * dists)
weights = alphas * exclusive_cumprod(1.-alphas + 1e-10).cpu().clone().detach()
plt.figure(figsize=(15,4))
plt.title("sigma and alpha for 3 rays")
plt.subplot(121)
plt.title("exponent of output alpha")
for i, alpha in enumerate(alphas):
plt.plot(alpha.view(-1).numpy(), label=f"ray[{raytext[i]}] sum(alpha) = {round(alpha.sum().item(),1)}")
plt.grid()
plt.legend()
plt.tight_layout()
plt.subplot(122)
plt.title("weights: cumprod of normalized output alpha")
for i, alpha in enumerate(weights):
plt.plot(alpha.view(-1).numpy(), label=f"ray[{raytext[i]}] sum(alpha) = {round(alpha.sum().item(),1)}")
plt.grid()
plt.legend()
plt.tight_layout()
plt.show()
# + [markdown] id="earhGgcTNruE"
# ## model result: rgbs are added over sigmoid, rgb on non transparent pixel
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="f0g7t3voSOdi" outputId="505d511f-58a3-48ab-e14f-fbdf059aa43d"
# Resulting color of pixel at [50,50]
rgb_map = torch.sum(weights[1][...,None].cpu() * rgb[1].cpu(), -2).clone().detach().numpy()
plt.imshow(rgb_map.reshape(1,1,3))
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="slmUMNKeTlib" outputId="ac6aacb0-e58d-488a-d394-58c6fd233db3"
wgts = weights[1][...,None].cpu() * rgb[1].cpu().clone().detach().numpy().reshape(64,3)
wgts.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 327} id="BkCBbPVuM2yT" outputId="022ec8e3-8652-4ccc-d781-ad2b09121506"
rgbs = rgb.cpu().clone().detach()[1]
wgts = weights[1][...,None].cpu() * rgb[1].cpu().clone().detach().numpy()#.reshape(64,3)
rgbs.shape, rgbs[...,0].shape
channels = ['r','g','b']
plt.figure(figsize=(15,4))
for i, channel in enumerate(channels):
plt.plot(rgbs[0][...,i], label=channel, color=channel)
plt.plot(wgts[0][...,i], label=f"{channel} wghts", color=channel, linestyle="--")
plt.grid()
plt.legend()
# plt.axis("off")
plt.tight_layout()
plt.show()
plt.figure(figsize=(15,1))
plt.imshow(rgbs.numpy())
plt.axis("off")
plt.tight_layout()
plt.show()
|
tiny_nerf_pytorch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from pandas import DataFrame
import matplotlib.pyplot as plt
# get weekly score
week_df = pd.read_csv("../data/Never Ending_week1_score.csv", encoding = "ISO-8859-1")
week_df
# -
# sort weekly score by name
week_df.sort_values(['Team Name'], ascending=[True], inplace=True)
week_df
total_df = pd.read_csv("../data/Never Ending_week0_score.csv", encoding = "ISO-8859-1")
total_df
# sort score score by name
total_df.sort_values(['Team Name'], ascending=[True], inplace=True)
total_df
names = week_df['Team Name'].tolist()
week_scores = week_df['Total'].tolist()
total_scores = total_df['Total'].tolist()
print(names, week_scores, total_scores)
# +
pos = list(range(1, len(df['Team Name'])+1))
print(pos)
width = 0.3
# Plotting the bars
fig, ax = plt.subplots(figsize=(20,12))
# Create a bar with week score,
# in position pos,
plt.bar([p + width for p in pos],
week_scores,
# of width
width,
# with alpha 0.5
alpha=0.5,
# with color
color='#EE3224',
edgecolor='red',
# with label the first value in first_name
label='Week')
# Create a bar with mid_score data,
# in position pos + some width buffer,
plt.bar([p + 2*width for p in pos],
#using df['mid_score'] data,
total_scores,
# of width
width,
# with alpha 0.5
alpha=0.2,
# with color
color='#F78F1E',
edgecolor='#000000',
# with label the second value in first_name
label='Season')
# Set the y axis label
ax.set_ylabel('Score')
# Set the chart's title
ax.set_title('Week 1')
# Set the position of the x ticks
ax.set_xticks([p + 1.5 * width for p in pos])
# Set the labels for the x ticks
ax.set_xticklabels(names, rotation=60)
# Setting the x-axis and y-axis limits
plt.xlim(min(pos)-width, max(pos)+width*4)
plt.ylim(0, 180 )
# Adding the legend and showing the plot
plt.legend(['Week', 'Season'], loc='upper right')
plt.grid()
plt.show()
# -
|
notebooks/week_bar_chart.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Super-Convergence Learning Rate Schedule (TensorFlow Backend)
# In this example we will implement super-convergence learning rate (LR) schedule (https://arxiv.org/pdf/1708.07120.pdf) and test it on a CIFAR10 image classification task. Super-covergence is a phenomenon where neural networks can be trained an order of magnitude faster than with standard training methods. The paper proposes a LR schedule which incorporates two parts: a LR range test to find the appropriate LR range and a cyclical LR schedule that uses the obtained information.
# +
import tempfile
import fastestimator as fe
from fastestimator.architecture.tensorflow import ResNet9
from fastestimator.dataset.data.cifair10 import load_data
from fastestimator.op.numpyop.meta import Sometimes
from fastestimator.op.numpyop.multivariate import HorizontalFlip, PadIfNeeded, RandomCrop
from fastestimator.op.numpyop.univariate import CoarseDropout, Normalize, Onehot
from fastestimator.op.tensorop.loss import CrossEntropy
from fastestimator.op.tensorop.model import ModelOp, UpdateOp
from fastestimator.trace.adapt import LRScheduler
from fastestimator.trace.io import BestModelSaver
from fastestimator.trace.metric import Accuracy
from fastestimator.util.util import Suppressor
import matplotlib.pyplot as plt
# + tags=["parameters"]
# Parameters
epochs=24
batch_size=128
lr_epochs=100
train_steps_per_epoch=None
save_dir=tempfile.mkdtemp()
# -
# ## Network Architecture and Data Pipeline
# We will use almost the same image classification configuration of the other Apphub example: [CIFAR10 Fast](../../image_classification/cifar10_fast/cifar10_fast.ipynb) including network architecture and data pipeline. The only difference is that we use SGD optimizer instead of Adam because author of the paper specially pointed out the incompatibility between Adam optimizer and super-convergence.
# +
# prepare dataset
train_data, test_data = load_data()
pipeline = fe.Pipeline(
train_data=train_data,
eval_data=test_data,
batch_size=batch_size,
ops=[
Normalize(inputs="x", outputs="x", mean=(0.4914, 0.4822, 0.4465), std=(0.2471, 0.2435, 0.2616)),
PadIfNeeded(min_height=40, min_width=40, image_in="x", image_out="x", mode="train"),
RandomCrop(32, 32, image_in="x", image_out="x", mode="train"),
Sometimes(HorizontalFlip(image_in="x", image_out="x", mode="train")),
CoarseDropout(inputs="x", outputs="x", mode="train", max_holes=1),
Onehot(inputs="y", outputs="y", mode="train", num_classes=10, label_smoothing=0.2)
])
# prepare network
model = fe.build(model_fn=ResNet9, optimizer_fn="sgd")
network = fe.Network(ops=[
ModelOp(model=model, inputs="x", outputs="y_pred"),
CrossEntropy(inputs=("y_pred", "y"), outputs="ce"),
UpdateOp(model=model, loss_name="ce")
])
# -
# ## LR Range Test
# The preparation of the super-convergence schedule is to search the suitable LR range. The process is training the target network with a linearly increasing LR and observing the validation accuracy. Generally, the accuracy will keep increase until at some certain point when the LR get too high and start making training diverge. The very LR of that moment is the "maximum LR".
#
# To run the test we need to implement the trace to record the maximum LR. After running the training with linear increaseing LR, we will get the maximum LR.
#
# <img src="./typical_lr.PNG" alt="drawing" width="400"/>
# [The typical learning rate and metircs plot from https://arxiv.org/pdf/1708.07120.pdf]
# +
def linear_increase(step, min_lr=0.0, max_lr=6.0, num_steps=1000):
lr = step / num_steps * (max_lr - min_lr) + min_lr
return lr
traces = [
Accuracy(true_key="y", pred_key="y_pred"),
LRScheduler(model=model, lr_fn=lambda step: linear_increase(step))
]
# prepare estimator
LR_range_test = fe.Estimator(pipeline=pipeline,
network=network,
epochs=lr_epochs,
traces=traces,
train_steps_per_epoch=10,
log_steps=10)
# run the LR_range_test this
print("Running LR range testing... It will take a while")
with Suppressor():
summary = LR_range_test.fit("LR_range_test")
# -
# Let's plot the accuracy vs LR graph and see the maximum LR.
acc_steps = [step for step in summary.history["eval"]["accuracy"].keys()]
acc_values = [acc for acc in summary.history["eval"]["accuracy"].values()]
best_step, best_acc = max(summary.history["eval"]["accuracy"].items(), key=lambda k: k[1])
lr_max = summary.history["train"]["model_lr"][best_step]
lr_values = [summary.history["train"]["model_lr"][x] for x in acc_steps]
assert len(lr_values) == len(acc_values)
plt.plot(lr_values, acc_values)
plt.plot(lr_max,
best_acc,
'o',
color='r',
label="Best Acc={}, LR={}".format(best_acc, lr_max))
plt.xlabel("Learning Rate")
plt.ylabel("Evaluation Accuracy")
plt.legend(loc='upper left', frameon=False)
# ## Super-Convergence LR Schedule
#
# Once we get the maximum LR, the minimum LR can be computed by dividing it by 40. Although this number is set to 4 in the paragraph of the original paper, it falls in range of [4, 40] in its experiment section. We empirically found 40 is the best value for this task.
#
# The LR change has 3 phases:
# 1. increase LR from minimum LR to maximum LR at 0~45% of training process
# 2. decrase LR from maximum LR to minimum LR at 45%~90% of training process
# 3. decrase LR from minimum LR to 0 at 90%~100% of training process
#
# <img src="./lr_schedule.PNG" alt="drawing" width="400"/>
# +
lr_min = lr_max / 40
mid = int(epochs * 0.45 * len(train_data) / batch_size)
end = int(epochs * len(train_data) / batch_size)
def super_schedule(step):
if step < mid:
lr = step / mid * (lr_max - lr_min) + lr_min # linear increase from lr_min to lr_max
elif mid <= step < mid * 2:
lr = lr_max - (step - mid) / mid * (lr_max - lr_min) # linear decrease from lr_max to lr_min
else:
lr = max(lr_min - (step - 2 * mid) / (end - 2 * mid) * lr_min, 0) # linear decrease from lr_min to 0
return lr
# -
# Before we start the main training, the model needs to be reinitialized. Therefore we re-instantiate the same network and plug the new LR scheduler in the estimator.
# +
# reinitialize the model
model = fe.build(model_fn=ResNet9, optimizer_fn="sgd")
network = fe.Network(ops=[
ModelOp(model=model, inputs="x", outputs="y_pred"),
CrossEntropy(inputs=("y_pred", "y"), outputs="ce"),
UpdateOp(model=model, loss_name="ce")
])
traces = [
Accuracy(true_key="y", pred_key="y_pred"),
BestModelSaver(model=model, save_dir=save_dir, metric="accuracy", save_best_mode="max"),
LRScheduler(model=model, lr_fn=lambda step: super_schedule(step))
]
# prepare estimator
main_train = fe.Estimator(pipeline=pipeline,
network=network,
epochs=epochs,
traces=traces,
train_steps_per_epoch=train_steps_per_epoch)
main_train.fit()
# -
# ## Result Discussion
# The result of it might not be super impressive when comparing with original example [CIFAR10 Fast](../../image_classification/cifar10_fast/cifar10_fast.ipynb). But please be aware that the example has its own LR schedules which is specially tuned on that configuration (plus that scheduler is also cyclical LR schedule).
|
apphub/lr_controller/super_convergence/super_convergence.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
from bs4 import BeautifulSoup
from selenium import webdriver
import pandas as pd
import time
import random
import datetime
import sys
import re
import warnings
warnings.filterwarnings("ignore")
# +
def strQ2B(ustring):
"""全角转半角"""
rstring = ""
for uchar in ustring:
inside_code=ord(uchar)
if inside_code == 12288: #全角空格直接转换
inside_code = 32
elif (inside_code >= 65281 and inside_code <= 65374): #全角字符(除空格)根据关系转化
inside_code -= 65248
rstring += chr(inside_code)
return rstring
def strB2Q(ustring):
"""半角转全角"""
rstring = ""
for uchar in ustring:
inside_code=ord(uchar)
if inside_code == 32: #半角空格直接转化
inside_code = 12288
elif inside_code >= 32 and inside_code <= 126: #半角字符(除空格)根据关系转化
inside_code += 65248
rstring += chr(inside_code)
return rstring
# -
etp_toggle = 'ETF'
# etp_toggle = 'LIP'
# +
# read in code list and create file
index_code_hk_stocks_pd = pd.read_excel('code_list_hk_all_' + etp_toggle + 's_v20181024.xlsx', header=None)
index_code_hk_stocks_ls = index_code_hk_stocks_pd[0].tolist()
columns_ls = ['index_code_hk', 'etf_name', 'listing_date', 'exp_ratio', 'suspend_status', 'suspend_date', 'replication_method', 'outstanding_shares', 'prev_close', 'lot_size']
results_filename = 'hkex-data-' + etp_toggle + '-v%s-supplement-2.csv' % datetime.date.today().strftime('%Y%m%d')
results_header_df = pd.DataFrame(columns = columns_ls)
results_header_df.to_csv(results_filename, header=True)
print(results_filename + " created.")
# -
results_header_df
# +
for i in [85, 91, 126, 143, 179, 180]:
# for i in range(len(index_code_hk_stocks_ls)):
tic = datetime.datetime.now()
try:
index_code_hk = index_code_hk_stocks_ls[i]
index_page = 'https://sc.hkex.com.hk/TuniS/www.hkex.com.hk/market-data/securities-prices/exchange-traded-products/exchange-traded-products-quote?sym=' + str(index_code_hk) + '&sc_lang=zh-cn'
driver = webdriver.PhantomJS()
driver.get(index_page)
soup = BeautifulSoup(driver.page_source)
etf_name = soup.find_all('p')[0].get_text()
etf_name = etf_name if not None else ''
etf_name = etf_name[:etf_name.rfind('(')].strip()
etf_name = strQ2B(etf_name)
etf_name = etf_name.replace('—', '-')
listing_date = soup.find_all("td", "ano col_listingdate")[0].get_text()
listing_year = listing_date[listing_date.find("年")-4:listing_date.find("年")]
listing_month = listing_date[listing_date.find("年")+1:listing_date.find("月")]
listing_day = listing_date[listing_date.find("月")+1:listing_date.find("日")]
listing_date_output = str(listing_year + '/' + listing_month + '/' + listing_day)
exp_ratio = soup.find_all('td', 'ano col_managefee')[0].get_text()
exp_ratio = exp_ratio if not None else ''
exp_ratio = exp_ratio[:exp_ratio.find('%')]
suspend_tag = soup.find_all('span', 'susico')[0]
suspend = 'suspend'
if suspend in str(suspend_tag):
suspend_status = 1
suspend_date = soup.find_all("dt", "ico_data col_aum_date")[0].get_text()
suspend_year = suspend_date[suspend_date.find("年")-4:suspend_date.find("年")]
suspend_month = suspend_date[suspend_date.find("年")+1:suspend_date.find("月")]
suspend_day = suspend_date[suspend_date.find("月")+1:suspend_date.find("日")]
suspend_date_output = str(suspend_year + '/' + suspend_month + '/' + suspend_day)
else:
suspend_status = 0
suspend_date_output = '-'
replication_method = soup.find_all("td", "col_replicatemethod")[0].get_text()
num_issued_shares = soup.find_all("td", "ano col_issued_shares")[0].get_text()
num_issued_shares = num_issued_shares[:num_issued_shares.find("(")].strip().replace(",", "")
prev_close = soup.find_all("dt", "ico_data col_prevcls")[0].get_text()
lot_size = soup.find_all("dt", "ico_data col_lotsize")[0].get_text()
df_tmp = pd.DataFrame([[index_code_hk, etf_name, listing_date_output, exp_ratio, suspend_status, suspend_date_output, replication_method, num_issued_shares, prev_close, lot_size]], columns = columns_ls)
df_tmp.to_csv(results_filename, header=False, mode='a')
# sys.stdout.flush()
except:
print(index_code_hk, ': error -------------------------')
sys.stdout.flush()
toc = datetime.datetime.now()
print(i+1, '/', len(index_code_hk_stocks_ls), ' ', 'processed, time elapsed: ', toc - tic)
sys.stdout.flush()
rand = random.randint(2,10)
print("About to sleep %d s" % rand)
sys.stdout.flush()
time.sleep(rand)
print("======= Finished =======")
# +
# test for new elements
index_page = 'https://sc.hkex.com.hk/TuniS/www.hkex.com.hk/market-data/securities-prices/exchange-traded-products/exchange-traded-products-quote?sym=2800&sc_lang=zh-cn'
driver = webdriver.PhantomJS()
driver.get(index_page)
soup = BeautifulSoup(driver.page_source)
|
ETF-data-hk-stock-etf-info-v20181015.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### General parameters
# +
# The name of the batch or the name of the forlder were programs are
batchName = 'batch_demo'
# The ID of the first progam to be generated, serves as a seed for the random generator
first_program_id = 2020
# Number of functions to be generated
number_of_functions = 15
# Number of different schedules to generate for each program
nb_schedule_per_function = 32
# Path to the directory containing the programs
data_path = './data/'+batchName+'/programs/'
# Number of nodes in the cluster, Each node will do the job on a portion of the programs
nb_nodes = 10
# Path of the temporary files folder
tmp_files_dir = './time_measurement/'
# Path to the script that compile generators
generator_script = './compile_tiramisu_code.py'
# Path of to the script that compiles wrappers
wrappers_script = './compile_tiramisu_wrappers.py'
# Path to the script that execute the compiled wrappers
execute_script = './execute_programs.py'
# Path to where to store the logs of the jobs
log_path = tmp_files_dir + "log_"+batchName+"/"
# Path where to strore the resulting dataset file
result_dataset_file = tmp_files_dir+'/processed_datasets/dataset_'+batchName+'.pkl'
#please edit the paths accordingly in compile_tiramisu_code.py, compile_tiramisu_wrappers.py, execute programs.py
# -
import pickle
from pathlib import Path
import numpy as np
import pandas as pd
import dill
from os import listdir
import json
from tqdm import tqdm
import re
import subprocess
import random
import time
from multiprocessing import Pool
from shutil import rmtree
import sys
sys.path.append("TiramisuCodeGenerator")
from TiramisuCodeGenerator import code_generator
from data_gen_utils import cluster_utilities, annotation_utilities, dataset_utilities
# ## 1. Generate synthetic Tiramisu programs
# +
# Generate programs using the program generator
code_generator.generate_programs(seed=first_program_id, number_of_functions=number_of_functions, nb_schedule_per_function=nb_schedule_per_function, batchName=batchName, output_dir = data_path)
cluster_utils = cluster_utilities(data_path, generator_script, wrappers_script, execute_script, log_path, batchName, nb_nodes, tmp_files_dir)
# Create the list of programs that are in the given path
cluster_utils.generate_prog_list()
# -
# ## 2. Generators' Compilation and Execution
## Generate job files for compiling Tiramisu code
cluster_utils.generate_compile_jobs()
# #### Submit generators' compilation and execution jobs
cluster_utils.submit_compile_jobs()
# +
# print(cluster_utils.compile_jobs_ids)
# -
cluster_utils.check_compile_progress()
# ## 3. Wrappers' Compilation
# Generate compile_wrappers and execute job files
cluster_utils.generate_wrapper_jobs()
# #### Submit wrapper's compilation jobs
cluster_utils.submit_wrapper_compilation_jobs()
# +
# print(cluster_utils.wrap_jobs_ids)
# -
cluster_utils.check_wrapper_compilation_progress()
# ## 4. Execute Programs
cluster_utils.generate_execution_slurm_script()
# #### Submit execution jobs
#submit exec jobs
cluster_utils.submit_execution_jobs()
print(cluster_utils.exec_jobs_ids)
cluster_utils.check_execution_progress()
# ## 5. Generate programs' annotation files
annot_utils= annotation_utilities()
annot_utils.generate_json_annotations(str(cluster_utils.data_path))
# annot_utils.generate_json_annotations_parallel(str(data_path),nb_threads=8)
# ## 6. Save execution results into a dataset file
dataset_utils = dataset_utilities()
dataset_utils.save_pkl_dataset(str(cluster_utils.data_path),
cluster_utils.tmp_files_dir+'results_'+cluster_utils.batchName+'/parts/',
result_dataset_file)
# ## 7. Test the resulting dataset (optional)
df = dataset_utils.get_dataset_df(result_dataset_file)
df
|
utils/SyntheticProgramsGenerator/Dataset_generator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
import PIL.Image
import numpy as np
int(None)
pil = PIL.Image.open("imgs/test1.jpg")
np.array(pil.convert('LA'))
arr = np.array(np.array(pil.convert('L')))
import matplotlib.pyplot as plt
# %matplotlib inline
hist = pil.convert('L').histogram()
fig, ax = plt.subplots(figsize=(10,int(pil.size[1]/pil.size[0]*10)))
ax.bar(range(256),height=hist,width=1.2)
ax.axis("off")
fig.savefig("d",'png')
pil.size
plt.bar(range(256),height=hist,width=0.8)
PIL.Image.fromarray(((arr>67)*255).astype(np.uint8))
from skimage.filters import roberts, sobel, prewitt, gaussian, median
arr
roberts(arr/255.)
PIL.Image.fromarray(((sobel(arr))*255.).astype(np.uint8))
PIL.Image.fromarray(((prewitt(arr))*255.).astype(np.uint8))
PIL.Image.fromarray(((median(arr))).astype(np.uint8))
median(arr)
PIL.Image.fromarray((np.abs(
prewitt(arr) - sobel(arr)
)*255.).astype(np.uint8))
np.abs(prewitt(arr) - sobel(arr))*255.
550/2.
2870-275
from skimage.morphology import erosion, dilation, opening, closing, white_tophat,binary_erosion
from skimage.morphology import disk
erosion(arr,disk(1))
opening(arr,disk(1))
PIL.Image.fromarray(((binary_erosion(arr,disk(1)))).astype(np.uint8))
a = 3
print(exec("a"))
int("3.3")
binary = arr >67
PIL.Image.fromarray(((binary*255.)).astype(np.uint8))
from skimage.morphology import medial_axis, skeletonize
skel, distance = medial_axis(binary, return_distance=True)
# +
# skel = skeletonize(binary, method='lee')
# -
import matplotlib.pyplot as plt
# %matplotlib inline
plt.imshow(skel)
distance
plt.imshow(distance, cmap=plt.cm.gray)
|
explore.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.11 64-bit (''mrcnn'': conda)'
# name: python3
# ---
# # Mask R-CNN - Train on Shapes Dataset
#
#
# This notebook shows how to train Mask R-CNN on your own dataset. To keep things simple we use a synthetic dataset of shapes (squares, triangles, and circles) which enables fast training. You'd still need a GPU, though, because the network backbone is a Resnet101, which would be too slow to train on a CPU. On a GPU, you can start to get okay-ish results in a few minutes, and good results in less than an hour.
#
# The code of the *Shapes* dataset is included below. It generates images on the fly, so it doesn't require downloading any data. And it can generate images of any size, so we pick a small image size to train faster.
# +
import os
import sys
import random
import math
import re
import time
import numpy as np
import cv2
import matplotlib
import matplotlib.pyplot as plt
# Root directory of the project
ROOT_DIR = os.path.abspath("../../")
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn.config import Config
from mrcnn import utils
import mrcnn.model as modellib
from mrcnn import visualize
from mrcnn.model import log
# %matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
# Local path to trained weights file
COCO_MODEL_PATH = os.path.join(ROOT_DIR, "mask_rcnn_coco.h5")
# Download COCO trained weights from Releases if needed
if not os.path.exists(COCO_MODEL_PATH):
utils.download_trained_weights(COCO_MODEL_PATH)
# -
# ## Configurations
# +
class ShapesConfig(Config):
"""Configuration for training on the toy shapes dataset.
Derives from the base Config class and overrides values specific
to the toy shapes dataset.
"""
# Give the configuration a recognizable name
NAME = "shapes"
# Train on 1 GPU and 8 images per GPU. We can put multiple images on each
# GPU because the images are small. Batch size is 8 (GPUs * images/GPU).
GPU_COUNT = 1
IMAGES_PER_GPU = 8
# Number of classes (including background)
NUM_CLASSES = 1 + 3 # background + 3 shapes
# Use small images for faster training. Set the limits of the small side
# the large side, and that determines the image shape.
IMAGE_MIN_DIM = 128
IMAGE_MAX_DIM = 128
# Use smaller anchors because our image and objects are small
RPN_ANCHOR_SCALES = (8, 16, 32, 64, 128) # anchor side in pixels
# Reduce training ROIs per image because the images are small and have
# few objects. Aim to allow ROI sampling to pick 33% positive ROIs.
TRAIN_ROIS_PER_IMAGE = 32
# Use a small epoch since the data is simple
STEPS_PER_EPOCH = 100
# use small validation steps since the epoch is small
VALIDATION_STEPS = 5
config = ShapesConfig()
config.display()
# -
# ## Notebook Preferences
def get_ax(rows=1, cols=1, size=8):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Change the default size attribute to control the size
of rendered images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
# ## Dataset
#
# Create a synthetic dataset
#
# Extend the Dataset class and add a method to load the shapes dataset, `load_shapes()`, and override the following methods:
#
# * load_image()
# * load_mask()
# * image_reference()
class ShapesDataset(utils.Dataset):
"""Generates the shapes synthetic dataset. The dataset consists of simple
shapes (triangles, squares, circles) placed randomly on a blank surface.
The images are generated on the fly. No file access required.
"""
def load_shapes(self, count, height, width):
"""Generate the requested number of synthetic images.
count: number of images to generate.
height, width: the size of the generated images.
"""
# Add classes
self.add_class("shapes", 1, "square")
self.add_class("shapes", 2, "circle")
self.add_class("shapes", 3, "triangle")
# Add images
# Generate random specifications of images (i.e. color and
# list of shapes sizes and locations). This is more compact than
# actual images. Images are generated on the fly in load_image().
for i in range(count):
bg_color, shapes = self.random_image(height, width)
self.add_image("shapes", image_id=i, path=None,
width=width, height=height,
bg_color=bg_color, shapes=shapes)
def load_image(self, image_id):
"""Generate an image from the specs of the given image ID.
Typically this function loads the image from a file, but
in this case it generates the image on the fly from the
specs in image_info.
"""
info = self.image_info[image_id]
bg_color = np.array(info['bg_color']).reshape([1, 1, 3])
image = np.ones([info['height'], info['width'], 3], dtype=np.uint8)
image = image * bg_color.astype(np.uint8)
for shape, color, dims in info['shapes']:
image = self.draw_shape(image, shape, dims, color)
return image
def image_reference(self, image_id):
"""Return the shapes data of the image."""
info = self.image_info[image_id]
if info["source"] == "shapes":
return info["shapes"]
else:
super(self.__class__).image_reference(self, image_id)
def load_mask(self, image_id):
"""Generate instance masks for shapes of the given image ID.
"""
info = self.image_info[image_id]
shapes = info['shapes']
count = len(shapes)
mask = np.zeros([info['height'], info['width'], count], dtype=np.uint8)
for i, (shape, _, dims) in enumerate(info['shapes']):
mask[:, :, i:i+1] = self.draw_shape(mask[:, :, i:i+1].copy(),
shape, dims, 1)
# Handle occlusions
occlusion = np.logical_not(mask[:, :, -1]).astype(np.uint8)
for i in range(count-2, -1, -1):
mask[:, :, i] = mask[:, :, i] * occlusion
occlusion = np.logical_and(occlusion, np.logical_not(mask[:, :, i]))
# Map class names to class IDs.
class_ids = np.array([self.class_names.index(s[0]) for s in shapes])
return mask.astype(np.bool), class_ids.astype(np.int32)
def draw_shape(self, image, shape, dims, color):
"""Draws a shape from the given specs."""
# Get the center x, y and the size s
x, y, s = dims
if shape == 'square':
cv2.rectangle(image, (x-s, y-s), (x+s, y+s), color, -1)
elif shape == "circle":
cv2.circle(image, (x, y), s, color, -1)
elif shape == "triangle":
points = np.array([[(x, y-s),
(x-s/math.sin(math.radians(60)), y+s),
(x+s/math.sin(math.radians(60)), y+s),
]], dtype=np.int32)
cv2.fillPoly(image, points, color)
return image
def random_shape(self, height, width):
"""Generates specifications of a random shape that lies within
the given height and width boundaries.
Returns a tuple of three valus:
* The shape name (square, circle, ...)
* Shape color: a tuple of 3 values, RGB.
* Shape dimensions: A tuple of values that define the shape size
and location. Differs per shape type.
"""
# Shape
shape = random.choice(["square", "circle", "triangle"])
# Color
color = tuple([random.randint(0, 255) for _ in range(3)])
# Center x, y
buffer = 20
y = random.randint(buffer, height - buffer - 1)
x = random.randint(buffer, width - buffer - 1)
# Size
s = random.randint(buffer, height//4)
return shape, color, (x, y, s)
def random_image(self, height, width):
"""Creates random specifications of an image with multiple shapes.
Returns the background color of the image and a list of shape
specifications that can be used to draw the image.
"""
# Pick random background color
bg_color = np.array([random.randint(0, 255) for _ in range(3)])
# Generate a few random shapes and record their
# bounding boxes
shapes = []
boxes = []
N = random.randint(1, 4)
for _ in range(N):
shape, color, dims = self.random_shape(height, width)
shapes.append((shape, color, dims))
x, y, s = dims
boxes.append([y-s, x-s, y+s, x+s])
# Apply non-max suppression wit 0.3 threshold to avoid
# shapes covering each other
keep_ixs = utils.non_max_suppression(np.array(boxes), np.arange(N), 0.3)
shapes = [s for i, s in enumerate(shapes) if i in keep_ixs]
return bg_color, shapes
# +
# Training dataset
dataset_train = ShapesDataset()
dataset_train.load_shapes(500, config.IMAGE_SHAPE[0], config.IMAGE_SHAPE[1])
dataset_train.prepare()
# Validation dataset
dataset_val = ShapesDataset()
dataset_val.load_shapes(50, config.IMAGE_SHAPE[0], config.IMAGE_SHAPE[1])
dataset_val.prepare()
# -
# Load and display random samples
image_ids = np.random.choice(dataset_train.image_ids, 4)
for image_id in image_ids:
image = dataset_train.load_image(image_id)
mask, class_ids = dataset_train.load_mask(image_id)
visualize.display_top_masks(image, mask, class_ids, dataset_train.class_names)
# ## Create Model
# Create model in training mode
model = modellib.MaskRCNN(mode="training", config=config,
model_dir=MODEL_DIR)
# +
# Which weights to start with?
init_with = "coco" # imagenet, coco, or last
if init_with == "imagenet":
model.load_weights(model.get_imagenet_weights(), by_name=True)
elif init_with == "coco":
# Load weights trained on MS COCO, but skip layers that
# are different due to the different number of classes
# See README for instructions to download the COCO weights
model.load_weights(COCO_MODEL_PATH, by_name=True,
exclude=["mrcnn_class_logits", "mrcnn_bbox_fc",
"mrcnn_bbox", "mrcnn_mask"])
elif init_with == "last":
# Load the last model you trained and continue training
model.load_weights(model.find_last(), by_name=True)
# -
a = model.keras_model.layers[1]
a.t
# ## Training
#
# Train in two stages:
# 1. Only the heads. Here we're freezing all the backbone layers and training only the randomly initialized layers (i.e. the ones that we didn't use pre-trained weights from MS COCO). To train only the head layers, pass `layers='heads'` to the `train()` function.
#
# 2. Fine-tune all layers. For this simple example it's not necessary, but we're including it to show the process. Simply pass `layers="all` to train all layers.
# Train the head branches
# Passing layers="heads" freezes all layers except the head
# layers. You can also pass a regular expression to select
# which layers to train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE,
epochs=1,
layers='heads')
# Fine tune all layers
# Passing layers="all" trains all layers. You can also
# pass a regular expression to select which layers to
# train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE / 10,
epochs=2,
layers="all")
# +
# Save weights
# Typically not needed because callbacks save after every epoch
# Uncomment to save manually
# model_path = os.path.join(MODEL_DIR, "mask_rcnn_shapes.h5")
# model.keras_model.save_weights(model_path)
# -
# ## Detection
# +
class InferenceConfig(ShapesConfig):
GPU_COUNT = 1
IMAGES_PER_GPU = 1
inference_config = InferenceConfig()
# Recreate the model in inference mode
model = modellib.MaskRCNN(mode="inference",
config=inference_config,
model_dir=MODEL_DIR)
# Get path to saved weights
# Either set a specific path or find last trained weights
# model_path = os.path.join(ROOT_DIR, ".h5 file name here")
model_path = model.find_last()
# Load trained weights
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
# +
# Test on a random image
image_id = random.choice(dataset_val.image_ids)
original_image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
log("original_image", original_image)
log("image_meta", image_meta)
log("gt_class_id", gt_class_id)
log("gt_bbox", gt_bbox)
log("gt_mask", gt_mask)
visualize.display_instances(original_image, gt_bbox, gt_mask, gt_class_id,
dataset_train.class_names, figsize=(8, 8))
# +
results = model.detect([original_image], verbose=1)
r = results[0]
visualize.display_instances(original_image, r['rois'], r['masks'], r['class_ids'],
dataset_val.class_names, r['scores'], ax=get_ax())
# -
# ## Evaluation
# +
# Compute VOC-Style mAP @ IoU=0.5
# Running on 10 images. Increase for better accuracy.
image_ids = np.random.choice(dataset_val.image_ids, 10)
APs = []
for image_id in image_ids:
# Load image and ground truth data
image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
molded_images = np.expand_dims(modellib.mold_image(image, inference_config), 0)
# Run object detection
results = model.detect([image], verbose=0)
r = results[0]
# Compute AP
AP, precisions, recalls, overlaps =\
utils.compute_ap(gt_bbox, gt_class_id, gt_mask,
r["rois"], r["class_ids"], r["scores"], r['masks'])
APs.append(AP)
print("mAP: ", np.mean(APs))
# -
|
samples/shapes/train_shapes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sweetpand/Algorithms/blob/master/FirstMissingPositive41.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="u77F2zhHVH7r" colab_type="text"
# Given an unsorted integer array, find the smallest missing positive integer.
#
# Input: [1,2,0]
# Output: 3
#
# Input: [3,4,-1,1]
# Output: 2
#
# Input: [7,8,9,11,12]
# Output: 1
# + id="SNko5AAvRHgO" colab_type="code" colab={}
def firstMissingPositive_Step1_2(A):
length=len(A)
#base Case
if 1 not in A:
return 1
elif length==1:
return 2
#replace negative intgers and numbers greater than length with 1
for i in range(length):
if A[i] <=0 or A[i]>length:
A[i]=1
return(A)
# + id="mBHnzh2UROby" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="014d8efa-a684-4a06-8ada-d2c36517fa5c"
A=[3,4,-1,1,-2]
firstMissingPositive_Step1_2(A)
# + id="KFb_Y0kCRZ_p" colab_type="code" colab={}
def firstMissingPositive(A):
length=len(A)
#base Case
if 1 not in A:
return 1
elif length==1:
return 2
#replace negative intgers and numbers greater than length with 1
for i in range(length):
if A[i] <=0 or A[i]>length:
A[i]=1
#Lets use index as hash key and number as sign decector =>
# if num [i] = +ve => i is misisng
# If num[i] = -ve => i is in Array
for i in range(length):
temp=abs(A[i])
#handle duplicates do it only once
if temp == length:
A[0]=-abs(A[0])
else:
A[temp]=-abs(A[temp])
#index of the first positive number is equal to first missing positive
for i in range(1,length):
if A[i] > 0:
return i
if A[0]>0:
return length
# + id="oye1WD8nRuRH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="abbf3c41-f184-43fd-ae58-dab0b0634ba7"
A=[3,4,-1,1]
firstMissingPositive(A)
# + id="G1IppdYHR2zO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="66fa58f1-bae8-4d24-fce8-0937541f7108"
A=[1,2,0]
firstMissingPositive(A)
# + id="kzjcsvBmR6Tt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4cb8123b-28c4-4bc1-e630-a68f91469018"
A=[7,8,9,11,12]
firstMissingPositive(A)
# + [markdown] id="e4w_U9w1R-ly" colab_type="text"
# Time Complexity: O(N).
#
# space Complexity: O(1)
#
# + [markdown] id="P0a9cGOb-bmO" colab_type="text"
# Solution using extra space: O(N) time and space.
#
#
# If the length of the nums array is N, then the first missing positive will be between 1 to N+1. Think Why N+1? We can have in the array 1 to N.
# Take an temp array of size N and for any number x in nums such that 1<=x<=N, mark temp[x-1]. Then simply walk the temp array and report the first unmarked index.
# + id="aM34v4Rz9-JQ" colab_type="code" colab={}
class Solution(object):
def firstMissingPositive(nums):
"""
:type nums: List[int]
:rtype: int
"""
temp, N = [None]*len(nums), len(nums)
for x in nums:
if 1<=x<=N:
temp[x-1] = x
for i in range(N):
if temp[i] == None:
return i+1
return N+1
# + id="wabRzCHPQ03j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="06498adb-ebe9-4af8-c150-3d69a90b598c"
A=[3,4,-1,1,-2]
Solution.firstMissingPositive(A)
# + [markdown] id="bCijzaWu-ulI" colab_type="text"
# Optimized solution with O(1) Space
#
# Simply traverse the nums array and put any number within [1, N] in their right place. For example if 2 is in that input, then put 2 at index 1.
# Now traverse this "shuffled" array again. You expect 1 at 0th index. Otherwise it is missing. Then you expect 2 at 1st index and so on.
# Above idea can be a little tricky. What about cases like [1] and [1,1] - i.e. 1 is in its place or there are duplicates - we need to advance pointer regardless.
#
# + id="ijhEwnEH-gff" colab_type="code" colab={}
class Solution(object):
def firstMissingPositive(nums):
"""
:type nums: List[int]
:rtype: int
"""
N, i = len(nums), 0
while i < N:
while 1<=nums[i]<=N:
idx_expected = nums[i]-1
if nums[i] == nums[idx_expected]:
break
nums[i], nums[idx_expected] = nums[idx_expected], nums[i]
i = i + 1
for i in range(N):
if nums[i] != i+1:
return i+1
return N+1
# + [markdown] id="1cM4IwdIVGmk" colab_type="text"
#
# + id="riWdGDwKRDfA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d6c98a60-ab29-439d-ca75-3bbf02e7577a"
A=[3,4,-1,1,-2]
Solution.firstMissingPositive(A)
# + id="JZ6KGwiWRGEW" colab_type="code" colab={}
|
FirstMissingPositive41.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# The BioGraph format is a compressed index of sequencing data that holds a rapidly querable representation of reads. The BioGraph has two main components. The Seqset and the Readmap. Briefly, the Seqset is the compressed index of every kmer from the foward and reverse complement of all reads in the sequencing experiment. Each of these kmers is called a SeqsetEntry or simply an entry. The Readmap is a lookup of what entries correspond to original input reads and keep information such as the read's length or if the read has a mate-pair.
#
# This tutorial is an introduction to basic queries and functionality of the BioGraphSDK.
#
# # Setup
#
# To follow along with this tutorial, the BioGraphSDK should have been installed and the `bgtools install_tests` have been run. For information on how to do this, see **Spiral Genetics’ BioGraph Toolkit and SDK User Guide** in the _Installation_ section
#
# # About the Data
#
# A handful of SNPs, indels, and SVs were randomly simulated in the "J02459.1 Enterobacteria
# phage lambda, complete genome". These variants were then given genotypes mocking a family
# trio. The sample genomes for the proband, mother, and father can be found in
# `references/samples/` as well a vcf in `variants/family.vcf`.
# For each sample, 150bp paired-end reads were simulated using [mason](http://www.seqan.de/apps/mason/).
#
# For the examples, we use the lambdaToyData. Point to the benchmark data or the results from a run of `bgtools install_tests` to follow along.
#
# # Opening a BioGraph
#
# To start, let's look at the single sample proband_lambda.bg by opening it as a `BioGraph` object and looking at some of its metadata.
# Update the `LAMBDA_PATH` variable to point to the root directory where converted lambdaToyData exists.
# +
LAMBDA_PATH = "/scratch/lambda/lambdaToyData/benchmark"
import os
import biograph as bgsdk
#Load the BioGraph
my_bg = bgsdk.BioGraph(os.path.join(LAMBDA_PATH, "proband_lambda.bg"))
ref = bgsdk.Reference(os.path.join(LAMBDA_PATH, "ref_lambda"))
print("BioGraph Version", my_bg.metadata.version)
print("AccessionId:", my_bg.metadata.accession_id)
print("BioGraphId:", my_bg.metadata.biograph_id)
print("Sample Info", my_bg.metadata.samples)
# -
# AccessionId is a user-specified sample identifier for the data.
# BioGraphId is an internal sha1 that's used to identify the sample for merging.
# Sample Info is a dictionary holding Key/Value of AccesionId to BioGraphId.
# We also looked at a couple propeties of the reads contained within the BioGraph.
#
# The AccessionIds are most useful for BioGraphs comprising multiple, merged samples. We'll cover details about merged BioGraphs in a later section.
# # Seqsets and Readmaps
#
# BioGraphs contain two main data structures: Seqsets and Readmaps. A readmap lists information about all reads that were seen. A Seqset contains a list of all sequences and subsequences of bases present in any readmap.
#
# The Seqset data structure is designed for efficient searching of arbitrary sequences, and effecient traversal of overlapping sequences.
#
# The Readmap data structure tracks which Seqset entries have reads that support them, and associated data about those reads such as pairing data.
# # Using a Seqset
#
# You can use the "find" method to look up a seqset entry by sequence, and the "sequence" method to look up the sequence associated with a seqset entry. However, try to avoid using these methods in performance critical code paths; it is much more efficient to use the seqset traversal methods such as "push_front" and "pop_front". In fact, "find" uses "push_front" internally, and "sequence" uses "pop_front" internally.
#
# Look up a seqset entry by sequence.
seqset = my_bg.seqset
print("This seqset has %d entries" % (seqset.size()))
query_seq = bgsdk.Sequence("GTAATCTTTTAAT" + "T" + "TTAAATAAGTTA")
missing_query_seq = bgsdk.Sequence("GTAATCTTTTAAT" + "C" + "TTAAATAAGTTA")
for seq in [query_seq, missing_query_seq]:
entry = seqset.find(seq)
if entry:
print("Sequence %s is present in the seqset" % (entry.sequence()))
else:
print("Sequence %s is not present in the seqset" % (seq))
# However, it is much more efficient to use the seqset traversal primitives to find neighboring entries if you're not looking up a whole sequence of bases from scratch.
seq = bgsdk.Sequence("AATCTTTTAATTTTAAATAAGTTA")
entry = seqset.find(seq)
for base in "TGATC":
seq = base + seq
entry = entry.push_front(base)
if entry:
print("%s exists in the seqset" % (seq))
else:
print("%s does not exist in the seqset" % (seq))
break
# # Using a Readmap
# If your BioGraph only contains a single sample, you will not have to specify the accession ID when opening the readmap. However, BioGraphs can contain multiple samples; see the section on Merged BioGraphs for details.
# +
my_rm = my_bg.open_readmap()
# Print general statistics about the readmap
print("Readmap contains %d reads and %d bases" % (my_rm.get_read_count(), my_rm.get_num_bases()))
p = my_rm.get_pair_stats()
print("\tPaired\tUnpaired")
print("Reads\t{p.paired_reads}\t{p.unpaired_reads}".format(p=p))
print("Bases\t{p.paired_bases}\t{p.unpaired_bases}".format(p=p))
print("Including both forward and reverse direction reads, readmap contains %d entries" % (my_rm.size()))
# -
# # Looking up reads
#
# If you've found a seqset entry and you'd like to see what reads relate to it, there are two methods available.
#
# One method lets you find all reads that are entirely contained within a seqset entry, and the other lets you find all reads that start with a given seqset entry and are entirely contained within it.
#
# Once you have a specific read, you can use "get_seqset_entry" to translate it back to its seqset entry.
# Find all reads that start with the given seqset entry and are entirely contained within it
query_seq = bgsdk.Sequence("AAAGAAGATTTCCAATAATCAGAACAAGTCGGCTCCTGTTTAGTTACGAGCGACATTGCTCCGTGTATTCACTCGTTGGAATGAATACACAGTGCAGTGTTTATTCTGTTATTTATGCCAAAAATAAAGGCCACTATCAGGCAGCTTTGT")
print("Original sequence:")
print(query_seq)
print("")
entry = seqset.find(query_seq)
for read in my_rm.get_prefix_reads(entry):
if read.is_original_orientation():
print("Found a read of length %d. This is how it appeared in the input data:" % (len(read)))
print(read.get_seqset_entry().sequence())
else:
print("Found a read of length %d. It appeared in the input data as its reverse complement:" % (len(read)))
print(read.get_seqset_entry().sequence().rev_comp())
if read.has_mate():
print(" The preceding read has a mate: %s" % (read.get_mate().get_seqset_entry().sequence()))
else:
print(" The preceding read was not part of a pair, however its reverse complement is: %s" % (read.get_rev_comp().get_seqset_entry().sequence()))
print("")
# Find all reads that contain a given seqset entry anywhere in the read.
query_seq = bgsdk.Sequence("CTTCCCTCTCCCCCAAATAAAAAGGCCTGCGATTACCAGCAGGCCTGTTATTAGCTCAGTAATGTAGATGGTCATCTTTTAACTCCATATACCGCCAATACCCGTTTCATCGCGGCACTCTGGCGACACTCCTTAAAAAC")
display_offset = 11
print("%s%s" % (" "*display_offset, query_seq))
print("")
entry = seqset.find(query_seq)
for (offset, read) in my_rm.get_reads_containing(entry):
print("%s%s" % (" "*(display_offset - offset), read.get_seqset_entry().sequence()))
# # Merged BioGraphs
#
# Merged BioGraphs can have multiple samples present in the BioGraph. In this case, the seqset will contain the union of all sequences contained in any input sample, and there will be a separate readmap for each sample.
#
# For these examples, we'll use the `family_lambda.bg` and take a look at all the properties.
my_bg = bgsdk.BioGraph(os.path.join(LAMBDA_PATH, "family_lambda.bg"))
print("AccessionId:", my_bg.metadata.accession_id)
print("BioGraphId:", my_bg.metadata.biograph_id)
print("Number of Samples", len(my_bg.metadata.samples))
for sample in my_bg.metadata.samples:
print("-- Opening Sample", sample, "--")
readmap = my_bg.open_readmap(sample)
print(" Number of Reads:", readmap.get_read_count())
print(" Number of Bases:", readmap.get_num_bases())
# Once you open a specific readmap, you can use all the same seqset and readmap calls in a merged BioGraph as you can use in a BioGraph with only one sample.
# # Using a Reference
#
# Since BioGraph files are reference agnostic, there is no information about a reference inside of it. Instead, information between the BioGraph and a Reference are pieced together on the fly. For example, let's use the lambda reference to find all of the reads that contain a portion of its chromosome.
my_ref = bgsdk.Reference(os.path.join(LAMBDA_PATH, "ref_lambda"))
ref_range = my_ref.make_range("lambda", 19575, 19600)
print("Looking for reads containing %s:%d-%d (%s)" % (ref_range.chromosome,
ref_range.start,
ref_range.end,
ref_range.sequence))
entry = my_bg.seqset.find(ref_range.sequence)
for sample in my_bg.metadata.samples:
readmap = my_bg.open_readmap(sample)
reads = readmap.get_reads_containing(entry)
print("%d matching reads found in %s" % (len(list(reads)), sample))
# Additionally, we can query the Reference for locations where a sequence exists. Note that this is more of a mapping than a proper alignment. This means that we report all of the places in the reference a sequence exactly matches to the direct strand. There currently is no method to do a 'fuzzy' match like what's needed for traditional alignment operations.
lookup = my_ref.find("CGTGCTGTC")
print(lookup.matches, "matches to reference found.")
print("chrom start end")
for i in range(lookup.matches):
mat = lookup.get_match(i)
print(mat.chromosome, mat.start, mat.end)
|
python/jupyter/biograph/BioGraph_SDK_Tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/tirthpatel7498/Classroom-Bot/blob/master/ECE542_Hw03a_CNN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-xwitz2wBmmG"
# # **Exploring Convolutional Neural Networks**
#
# In this notebook, we will explore some of the basic tools for regularization and hyper-parameter tuning for neural networks.
#
#
# + id="6NcBPjVCdMgK" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="5ca23435-546d-47c2-c04e-f60b8ed47ba3"
# %tensorflow_version 1.x
import numpy as np
import matplotlib.pyplot as plt
import keras
from keras.datasets import cifar10
from keras.utils import to_categorical
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Activation, BatchNormalization, Dropout
from tensorflow.keras import regularizers
from keras.optimizers import SGD, Adam, RMSprop
from sklearn.model_selection import GridSearchCV
from keras.wrappers.scikit_learn import KerasClassifier
# + [markdown] id="X-3WETNQiamR"
# ## Load and Splitting Data
#
# We Will make use of the CIFAR10 dataset.
# + id="e10W6I2QbDH6"
# Loading train and test dataset
(trainingX, trainingY), (testX, testY) = cifar10.load_data()
labels = {0:"airplane", 1:"automobile", 2:"bird", 3:"cat", 4:"deer", 5:"dog", 6:"frog", 7:"horse", 8:"ship", 9:"truck"}
# Keeping 20% for Validation
valid_set_pc = 0.2
split= int((len(trainingX))*(1-valid_set_pc))
validX = trainingX[split:]
validY = trainingY[split:]
trainX = trainingX[:split]
trainY = trainingY[:split]
# + id="Jh9Zdh4xOQyU" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="9e270c43-bd46-4d61-81fb-5a41fc28ba35"
# Showing a samples from the dataset
sampleID = 100
plt.imshow(trainX[sampleID])
print(labels[trainY[sampleID][0]])
# + id="ds8L26hSbsfr"
# Convert from integers to floats
trainingX = trainingX.astype('float32')
trainX = trainX.astype('float32')
validX = validX.astype('float32')
testX = testX.astype('float32')
# Normalize to range 0-1
trainingX = trainingX / 255.0
trainX = trainX / 255.0
validX = validX / 255.0
testX = testX / 255.0
# One-hot encoding of outputs
trainingY = to_categorical(trainingY)
trainY = to_categorical(trainY)
validY = to_categorical(validY)
testY = to_categorical(testY)
# + [markdown] id="Oejdojf2E99q"
# ## Training a Base CNN Model
# + id="2HOohn5l4RoF" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="62c8db5b-e51a-45e0-c12b-f33575d4bcff"
# Defining Base CNN Model
def define_base_model():
model = Sequential()
model.add(Conv2D(256, (3, 3), activation='relu', padding='same', input_shape=(32, 32, 3)))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(256, (3, 3), activation='relu', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(256, (3, 3), activation='relu', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(Flatten())
model.add(Dense(1024, activation='relu'))
model.add(Dense(10, activation='softmax'))
return model
# Creating an instance
base_model = define_base_model()
base_model.summary()
base_model.compile(optimizer=SGD(), loss='categorical_crossentropy', metrics=['accuracy'])
# + id="50IbgnrtcEf-" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="f9b9b225-f679-4a28-93cd-89fd9e193f16"
# Training base model
base_history = base_model.fit(trainX, trainY, epochs=15, validation_data=(validX, validY), verbose=1)
# + id="psvubBDOsJYH" colab={"base_uri": "https://localhost:8080/", "height": 573} outputId="ffcb4d6b-e8ca-4643-e78c-e257a503a866"
# Defining a function for plotting training and validation learning curves
def plot_history(history):
# plot loss
plt.title('Loss')
plt.plot(history.history['loss'], color='blue', label='train')
plt.plot(history.history['val_loss'], color='red', label='test')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'])
plt.show()
# plot accuracy
plt.title('Accuracy')
plt.plot(history.history['accuracy'], color='blue', label='train')
plt.plot(history.history['val_accuracy'], color='red', label='test')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'])
plt.show()
plot_history(base_history)
# + [markdown] id="Qrtm3IE2jyGN"
# # **[Task 1] Comparing Optimizers**
#
# The goal of of this section is to compare the performance of various optimizers: SGD with momentum, RMS Prop and Adam. Make sure you complete the following steps:
#
# 1. Train the network with SGB with momentum
# 2. Train the network with RMS Prop
# 3. Train the network with Adam
# 4. Create two plots showing accuracy for the training and validation sets
# 5. Comment on the performance of each optimizer. Which one had the steepest learning curve? Which one gave the final best performance?
#
#
#
# + id="WAImdmh5kCQ2" colab={"base_uri": "https://localhost:8080/", "height": 1542} outputId="9163507e-6b7c-4bac-c077-f96024f4f454"
# TODO - Training model with momentum
momentum_model = define_base_model()
momentum_model.summary()
momentum_model.compile(optimizer=SGD(learning_rate=0.01, momentum=0.9), loss='categorical_crossentropy', metrics=['accuracy'])
momentum_history = momentum_model.fit(trainX, trainY, epochs=15, validation_data=(validX, validY), verbose=1)
plot_history(momentum_history)
# + id="Cyi0YBgz2l1w" colab={"base_uri": "https://localhost:8080/", "height": 1542} outputId="62058be0-489d-4633-8a75-0a672a55348f"
# TODO - Training model with RMS Prop
rms_model = define_base_model()
rms_model.summary()
rms_model.compile(optimizer=RMSprop(learning_rate=0.001), loss='categorical_crossentropy', metrics=['accuracy'])
rms_history = rms_model.fit(trainX, trainY, epochs=15, validation_data=(validX, validY), verbose=1)
plot_history(rms_history)
# + id="xHWhlGUYZ68s" colab={"base_uri": "https://localhost:8080/", "height": 1542} outputId="d217f50d-8534-493b-a136-405c3d66fdd9"
# TODO - Training model with Adam
adam_model = define_base_model()
adam_model.summary()
adam_model.compile(optimizer=Adam(learning_rate=0.001), loss='categorical_crossentropy', metrics=['accuracy'])
adam_history = adam_model.fit(trainX, trainY, epochs=15, validation_data=(validX, validY), verbose=1)
plot_history(adam_history)
# + id="hRXknd9oTt4o" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="a6c7aa4e-6e09-4048-d0fe-966b89126c76"
# TODO - Plotting training accuracy
plt.title('Training Accuracy')
plt.plot(base_history.history['accuracy'], label='base train')
plt.plot(momentum_history.history['accuracy'], label='momentum train')
plt.plot(rms_history.history['accuracy'], label='rms train')
plt.plot(adam_history.history['accuracy'], label='adam train')
plt.legend()
plt.show()
# + [markdown] id="8vBjDkEFZSyH"
# [TODO - Add your observations]
#
#
# + [markdown] id="7JSTT5hZk4pX"
# # **[Task 2] Applying Standard Regularization**
#
# The goal of this section is to compare the effect of different normalization approaches including Batch normalization and Dropout. Make sure to complete the following steps:
#
# 1. Create a new model by adding a batch normalization layer after each convolutional layer and between the dense layers of the base model. Train it using the Adam optimizer.
# 2. Create a new model by adding dropout after each convolutional layer and between the dense layers of the base model with a rate equal to $0.2$. Train it using the Adam optimizer.
# 3. Create two plots showing accuracy for the training and validation sets comparing the regularized versions against the base model trained with the Adam optimizer.
# 4. Comment on the performance of each model. Which one had the steepest learning curve? Which one gave the final best performance?
#
#
#
# + id="dkfTDI205hGq" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="31affe6d-0876-4bc3-887c-d68cbedcb5b1"
# TODO - Defining Batch Normalization model and training it
def define_base_model_BN():
model = Sequential()
model.add(Conv2D(256, (3, 3), activation='relu', padding='same', input_shape=(32, 32, 3)))
model.add(BatchNormalization())
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(256, (3, 3), activation='relu', padding='same'))
model.add(BatchNormalization())
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(256, (3, 3), activation='relu', padding='same'))
model.add(BatchNormalization())
model.add(MaxPooling2D((2, 2)))
model.add(Flatten())
model.add(Dense(1024, activation='relu'))
model.add(BatchNormalization())
model.add(Dense(10, activation='softmax'))
return model
adam_model_BN = define_base_model_BN()
adam_model_BN.summary()
adam_model_BN.compile(optimizer=Adam(learning_rate=0.001), loss='categorical_crossentropy', metrics=['accuracy'])
adam_history_BN = adam_model_BN.fit(trainX, trainY, epochs=15, validation_data=(validX, validY), verbose=1)
# + id="04NtpB2tfMcG" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="d9157c80-5a9a-42d4-9df9-ada91387face"
# TODO - Defining dropout regularized model and training it
def define_base_model_Dropout():
model = Sequential()
model.add(Conv2D(256, (3, 3), activation='relu', padding='same', input_shape=(32, 32, 3)))
model.add(Dropout(0.2))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(256, (3, 3), activation='relu', padding='same'))
model.add(Dropout(0.2))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(256, (3, 3), activation='relu', padding='same'))
model.add(Dropout(0.2))
model.add(MaxPooling2D((2, 2)))
model.add(Flatten())
model.add(Dense(1024, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
return model
adam_model_Dropout = define_base_model_Dropout()
adam_model_Dropout.summary()
adam_model_Dropout.compile(optimizer=Adam(learning_rate=0.001), loss='categorical_crossentropy', metrics=['accuracy'])
adam_history_Dropout = adam_model_Dropout.fit(trainX, trainY, epochs=15, validation_data=(validX, validY), verbose=1)
# + id="FUduWXzv16rV" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="e9a8120f-4445-42e7-d615-2d29455d47ab"
# TODO - Plotting training accuracy
plt.plot(adam_history_BN.history['accuracy'], label='Model with BN')
plt.plot(adam_history_Dropout.history['accuracy'], label='Model with Dropout')
# + [markdown] id="zFURxqaLf1PQ"
# [TODO - Add your observations]
# + [markdown] id="J1REJkBwpEDd"
# # **[Task 3] Hyperparameter Tuning**
#
# The objective of this section is to perform hyperparameter tuning of the network using a grid search. Make sure to complete these steps:
#
# 1. Perform a grid search for the optimal hyperparameters for the model with dropout regularization and Adam optimizer using the dropout values [0.1, 0.2, 0.3] and the learning rate values [0.1, 0.01, 0.001].
# 2. Display the results for the best model on the test dataset.
# + id="M5HMnq-8D8l_" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="55302d42-0c9e-4f11-baa5-42275fd8d60a"
# TODO - Defining final model for optimization
def define_model_Dropout(d):
model = Sequential()
model.add(Conv2D(256, (3, 3), activation='relu', padding='same', input_shape=(32, 32, 3)))
model.add(Dropout(d))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(256, (3, 3), activation='relu', padding='same'))
model.add(Dropout(d))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(256, (3, 3), activation='relu', padding='same'))
model.add(Dropout(d))
model.add(MaxPooling2D((2, 2)))
model.add(Flatten())
model.add(Dense(1024, activation='relu'))
model.add(Dropout(d))
model.add(Dense(10, activation='softmax'))
return model
dropout=[0.1, 0.2, 0.3]
lr = [0.1, 0.01, 0.001]
best_dropout = 0.1
best_lr = 0.1
max = 0
for d in dropout:
model = define_model_Dropout(d)
for l in lr:
model.compile(optimizer=Adam(learning_rate=0.001), loss='categorical_crossentropy', metrics=['accuracy'])
model_history = model.fit(trainX, trainY, epochs=15, validation_data=(validX, validY), verbose=1)
val_acc = model_history.history['val_accuracy']
if val_acc[-1] > max:
best_dropout = d
best_lr = l
# + id="EGltGG4Ayj17" colab={"base_uri": "https://localhost:8080/"} outputId="875c859e-324c-481b-9dc4-67fbb2fdfd80"
# TODO - Displaying best parameters found
print("Best parameters are-> Dropout: ", best_dropout, " Learning rate: ", best_lr)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="E2zHNJvNmxaE" outputId="64d8529e-689d-49f5-af41-d7b61080ff7b"
final_model = model = define_model_Dropout(best_dropout)
final_model.compile(optimizer=Adam(learning_rate=best_lr), loss='categorical_crossentropy', metrics=['accuracy'])
final_model_history = final_model.fit(trainX, trainY, epochs=15, validation_data=(validX, validY), verbose=1)
result = final_model.evaluate(testX, testY)
print(result)
|
ECE542_Hw03a_CNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # KFServing Deployment Pipeline For CIFAR10 Image Recognition
#
# In this example (and Kale Kubeflow pipeline) we create a deployment pipeline for a pretrained CIFAR10 image model.
# The following steps will be run:
#
# * Setup Minio client
# * Download and test CIFAR10 model and train an Anchors Images explainer on it. Save both model and explainer to Minio.
# * Deploy model and explainer using KFServing and test
# * Train outlier detector
# * Train drift detector
# * Deploy knative eventing display to show asynchronous results from outlier and drift detectors.
# * Deploy outlier detector and test
# * deploy drift detector and test
#
# ### Setup
#
# You will need a kubeflow cluster >= 1.0 with
#
# * Knative eventing
# * Seldon >= 1.2.2
# * KFServing >= 0.3.0
#
# ### Kubeflow Jupyter Notebook Server
#
# To run this notebook inside kubeflow. Create a Jupyter notebook server using the image `seldonio/jupyter-lab-alibi-kale:0.11`
#
# ### GCP Setup
#
# On GCP If you use Kale to save this notebook as a pipeline you will need to add the storage_class of the `VolumeOp` to `nfs-client` if you have followed the steps to create a NFS RWX PV on GCP. e.g.:
#
# ```
# marshal_vop = dsl.VolumeOp(
# name="kale-marshal-volume",
# resource_name="kale-marshal-pvc",
# storage_class="nfs-client",
# modes=dsl.VOLUME_MODE_RWM,
# size="1Gi"
# )
# ```
#
#
#
# + tags=["imports"]
import numpy as np
from sklearn.ensemble import RandomForestClassifier
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from alibi.explainers import AnchorImage
from alibi.datasets import fetch_adult
from minio import Minio
from minio.error import ResponseError
from joblib import dump, load
import dill
from subprocess import run, Popen, PIPE
from alibi_detect.utils.data import create_outlier_batch
from alibi_detect.utils.fetching import fetch_tf_model
import json
import logging
import matplotlib.pyplot as plt
import tensorflow as tf
tf.keras.backend.clear_session()
from tensorflow.keras.layers import Conv2D, Conv2DTranspose, Dense, Layer, Reshape, InputLayer
from tqdm import tqdm
from alibi_detect.models.losses import elbo
from alibi_detect.od import OutlierVAE
from alibi_detect.utils.fetching import fetch_detector
from alibi_detect.utils.perturbation import apply_mask
from alibi_detect.utils.saving import save_detector, load_detector
from alibi_detect.utils.visualize import plot_instance_score, plot_feature_outlier_image
import time
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
# + [markdown] tags=[]
# ## Setup Pipeline Paramers
#
# The following global variables can be set. These will be used as Pipeline parameters.
# + tags=["pipeline-parameters"]
MINIO_HOST="minio-service.kubeflow:9000"
MINIO_ACCESS_KEY="minio"
MINIO_SECRET_KEY="minio123"
MINIO_MODEL_BUCKET="seldon"
CIFAR10_MODEL_PATH="tfserving/cifar10/model"
EXPLAINER_MODEL_PATH="tfserving/cifar10/explainer"
OUTLIER_MODEL_PATH="tfserving/cifar10/outlier"
DRIFT_MODEL_PATH="tfserving/cifar10/drift"
DEPLOY_NAMESPACE="admin"
TRAIN_OUTLIER_DETECTOR=False
TRAIN_DRIFT_DETECTOR=False
# + tags=["functions"]
def get_minio():
return Minio(MINIO_HOST,
access_key=MINIO_ACCESS_KEY,
secret_key=MINIO_SECRET_KEY,
secure=False)
# + tags=["block:setup"]
minioClient = get_minio()
buckets = minioClient.list_buckets()
for bucket in buckets:
print(bucket.name, bucket.creation_date)
# + tags=[]
if not minioClient.bucket_exists(MINIO_MODEL_BUCKET):
minioClient.make_bucket(MINIO_MODEL_BUCKET)
# + [markdown] tags=[]
# ## Test and save Model
#
# For simplicity we will use a pretrained Resnet32 CIFAR10 tensorflow model
# + tags=["block:train_model_and_explainer", "prev:setup"]
model = fetch_tf_model('cifar10', 'resnet32')
# + tags=[]
train, test = tf.keras.datasets.cifar10.load_data()
X_train, y_train = train
X_test, y_test = test
X_train = X_train.astype('float32') / 255
X_test = X_test.astype('float32') / 255
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
# + tags=[]
class_names = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
# + [markdown] tags=[]
# Test model locally.
# + tags=[]
idx = 1
X = X_test[idx].reshape(1, 32, 32, 3)
plt.imshow(X.reshape(32, 32, 3))
plt.axis('off')
plt.show()
print("class:",class_names[y_test[idx][0]])
print("prediction:",class_names[model.predict(X_test[idx:idx+1])[0].argmax()])
# + tags=[]
modelfilepath="resnet"
tf.saved_model.save(model, modelfilepath)
# + tags=[]
from os import listdir
from os.path import isfile, join
model_filepath="resnet"
print(get_minio().fput_object(MINIO_MODEL_BUCKET, f"{CIFAR10_MODEL_PATH}/1/saved_model.pb", modelfilepath+"/saved_model.pb"))
variable_filepath = modelfilepath+"/variables"
onlyfiles = [f for f in listdir(variable_filepath) if isfile(join(variable_filepath, f))]
for filename in onlyfiles:
print(filename)
print(get_minio().fput_object(MINIO_MODEL_BUCKET, f"{CIFAR10_MODEL_PATH}/1/variables/{filename}", join(variable_filepath, filename)))
# + [markdown] tags=[]
# ## Train Explainer
# + tags=["block:"]
def predict_fn(x):
return model.predict(x)
# + tags=["block:"]
image_shape = (32, 32, 3)
segmentation_fn = 'slic'
kwargs = {'n_segments': 5, 'compactness': 20, 'sigma': .5}
explainer = AnchorImage(predict_fn, image_shape, segmentation_fn=segmentation_fn,
segmentation_kwargs=kwargs, images_background=None)
# + tags=[]
idx=0
image = X_test[0]
np.random.seed(0)
explanation = explainer.explain(image, threshold=.95, p_sample=.5, tau=0.25)
# + tags=[]
X = X_test[idx].reshape(1, 32, 32, 3)
plt.imshow(X.reshape(32, 32, 3))
plt.axis('off')
plt.show()
print("class:",class_names[y_test[idx][0]])
print("prediction:",class_names[model.predict(X_test[idx:idx+1])[0].argmax()])
# + tags=[]
plt.imshow(explanation["anchor"])
# + tags=[]
with open("explainer.dill", "wb") as dill_file:
dill.dump(explainer, dill_file)
dill_file.close()
print(get_minio().fput_object(MINIO_MODEL_BUCKET, f"{EXPLAINER_MODEL_PATH}/explainer.dill", 'explainer.dill'))
# + [markdown] tags=[]
# ## Train Outlier Detector
#
# For further details and extended notebook see [Alibi-Detect Documentation](https://docs.seldon.io/projects/alibi-detect/en/stable/). These steps were derived from [Alibi-Detect CIFAR10 Example](https://docs.seldon.io/projects/alibi-detect/en/stable/examples/od_vae_cifar10.html)
# + tags=["block:train_outlier_detector", "prev:train_model_and_explainer"]
import logging
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
tf.keras.backend.clear_session()
from tensorflow.keras.layers import Conv2D, Conv2DTranspose, Dense, Layer, Reshape, InputLayer
from tqdm import tqdm
from alibi_detect.models.losses import elbo
from alibi_detect.od import OutlierVAE
from alibi_detect.utils.fetching import fetch_detector
from alibi_detect.utils.perturbation import apply_mask
from alibi_detect.utils.saving import save_detector, load_detector
from alibi_detect.utils.visualize import plot_instance_score, plot_feature_outlier_image
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
# + tags=[]
if TRAIN_OUTLIER_DETECTOR:
latent_dim = 1024
encoder_net = tf.keras.Sequential(
[
InputLayer(input_shape=(32, 32, 3)),
Conv2D(64, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2D(128, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2D(512, 4, strides=2, padding='same', activation=tf.nn.relu)
])
decoder_net = tf.keras.Sequential(
[
InputLayer(input_shape=(latent_dim,)),
Dense(4*4*128),
Reshape(target_shape=(4, 4, 128)),
Conv2DTranspose(256, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2DTranspose(64, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2DTranspose(3, 4, strides=2, padding='same', activation='sigmoid')
])
# initialize outlier detector
od = OutlierVAE(threshold=.015, # threshold for outlier score
score_type='mse', # use MSE of reconstruction error for outlier detection
encoder_net=encoder_net, # can also pass VAE model instead
decoder_net=decoder_net, # of separate encoder and decoder
latent_dim=latent_dim,
samples=2)
# train
od.fit(X_train,
loss_fn=elbo,
cov_elbo=dict(sim=.05),
epochs=50,
verbose=True)
else:
od = load_detector("/home/models/samples/od/cifar10")
# + tags=[]
idx = 8
X = X_train[idx].reshape(1, 32, 32, 3)
X_recon = od.vae(X)
plt.imshow(X.reshape(32, 32, 3))
plt.axis('off')
plt.show()
plt.imshow(X_recon.numpy().reshape(32, 32, 3))
plt.axis('off')
plt.show()
# + tags=[]
X = X_train[:500]
print(X.shape)
od_preds = od.predict(X,
outlier_type='instance', # use 'feature' or 'instance' level
return_feature_score=True, # scores used to determine outliers
return_instance_score=True)
print(list(od_preds['data'].keys()))
target = np.zeros(X.shape[0],).astype(int) # all normal CIFAR10 training instances
labels = ['normal', 'outlier']
plot_instance_score(od_preds, target, labels, od.threshold)
# + tags=[]
from alibi_detect.utils.saving import save_detector, load_detector
from os import listdir
from os.path import isfile, join
filepath="cifar10outlier"
save_detector(od, filepath)
onlyfiles = [f for f in listdir(filepath) if isfile(join(filepath, f))]
for filename in onlyfiles:
print(filename)
print(get_minio().fput_object(MINIO_MODEL_BUCKET, f"{OUTLIER_MODEL_PATH}/{filename}", join(filepath, filename)))
filepath="cifar10outlier/model"
onlyfiles = [f for f in listdir(filepath) if isfile(join(filepath, f))]
for filename in onlyfiles:
print(filename)
print(get_minio().fput_object(MINIO_MODEL_BUCKET, f"{OUTLIER_MODEL_PATH}/model/{filename}", join(filepath, filename)))
# + [markdown] tags=[]
# ## Train a Drift Detector
# + tags=["block:train_drift_detector", "prev:train_model_and_explainer"]
import matplotlib.pyplot as plt
import numpy as np
import os
import tensorflow as tf
from tensorflow.keras.layers import Conv2D, Dense, Flatten, InputLayer, Reshape
from alibi_detect.cd import KSDrift
from alibi_detect.cd.preprocess import uae, hidden_output
from alibi_detect.models.resnet import scale_by_instance
from alibi_detect.utils.fetching import fetch_tf_model, fetch_detector
from alibi_detect.utils.prediction import predict_batch
from alibi_detect.utils.saving import save_detector, load_detector
from alibi_detect.datasets import fetch_cifar10c, corruption_types_cifar10c
# + tags=[]
tf.random.set_seed(0)
if True:
np.random.seed(0)
n_test = X_test.shape[0]
idx = np.random.choice(n_test, size=n_test // 2, replace=False)
idx_h0 = np.delete(np.arange(n_test), idx, axis=0)
X_ref,y_ref = X_test[idx], y_test[idx]
X_h0, y_h0 = X_test[idx_h0], y_test[idx_h0]
print(X_ref.shape, X_h0.shape)
# define encoder
encoding_dim = 32
encoder_net = tf.keras.Sequential(
[
InputLayer(input_shape=(32, 32, 3)),
Conv2D(64, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2D(128, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2D(512, 4, strides=2, padding='same', activation=tf.nn.relu),
Flatten(),
Dense(encoding_dim,)
]
)
# initialise drift detector
p_val = .05
cd = KSDrift(
p_val=p_val, # p-value for K-S test
X_ref=X_ref, # test against original test set
preprocess_fn=uae, # UAE for dimensionality reduction
preprocess_kwargs={'encoder_net': encoder_net, 'batch_size': 128},
alternative='two-sided' # other options: 'less', 'greater'
)
else:
cd = load_detector("/home/models/samples/cd/cifar10")
# + tags=[]
from alibi_detect.utils.saving import save_detector, load_detector
from os import listdir
from os.path import isfile, join
filepath="cifar10Drift"
save_detector(cd, filepath)
onlyfiles = [f for f in listdir(filepath) if isfile(join(filepath, f))]
for filename in onlyfiles:
print(filename)
print(get_minio().fput_object(MINIO_MODEL_BUCKET, f"{DRIFT_MODEL_PATH}/{filename}", join(filepath, filename)))
filepath="cifar10Drift/model"
onlyfiles = [f for f in listdir(filepath) if isfile(join(filepath, f))]
for filename in onlyfiles:
print(filename)
print(get_minio().fput_object(MINIO_MODEL_BUCKET, f"{DRIFT_MODEL_PATH}/model/{filename}", join(filepath, filename)))
# + [markdown] tags=[]
# ## Deploy KFServing Model
# + tags=["block:deploy_model", "prev:train_model_and_explainer"]
secret=f"""apiVersion: v1
kind: Secret
metadata:
name: cifar10-kf-secret
namespace: {DEPLOY_NAMESPACE}
annotations:
serving.kubeflow.org/s3-endpoint: {MINIO_HOST} # replace with your s3 endpoint
serving.kubeflow.org/s3-usehttps: "0" # by default 1, for testing with minio you need to set to 0
type: Opaque
stringData:
awsAccessKeyID: {MINIO_ACCESS_KEY}
awsSecretAccessKey: {MINIO_SECRET_KEY}
"""
with open("secret.yaml","w") as f:
f.write(secret)
run("kubectl apply -f secret.yaml", shell=True)
# -
secret = f"""apiVersion: v1
kind: Secret
metadata:
name: seldon-init-container-secret
namespace: {DEPLOY_NAMESPACE}
type: Opaque
stringData:
AWS_ACCESS_KEY_ID: {MINIO_ACCESS_KEY}
AWS_SECRET_ACCESS_KEY: {MINIO_SECRET_KEY}
AWS_ENDPOINT_URL: http://{MINIO_HOST}
USE_SSL: "false"
"""
with open("secret.yaml","w") as f:
f.write(secret)
run("cat secret.yaml | kubectl apply -f -", shell=True)
# + tags=[]
sa = f"""apiVersion: v1
kind: ServiceAccount
metadata:
name: minio-kf-sa
namespace: {DEPLOY_NAMESPACE}
secrets:
- name: cifar10-kf-secret
"""
with open("sa.yaml","w") as f:
f.write(sa)
run("kubectl apply -f sa.yaml", shell=True)
# + tags=[]
from kubernetes import client
from kfserving import KFServingClient
from kfserving import constants
from kfserving import utils
from kfserving import V1alpha2EndpointSpec
from kfserving import V1alpha2PredictorSpec
from kfserving import V1alpha2ExplainerSpec
from kfserving import V1alpha2AlibiExplainerSpec
from kfserving import V1alpha2TensorflowSpec
from kfserving import V1alpha2InferenceServiceSpec
from kfserving import V1alpha2InferenceService
from kfserving import V1alpha2Logger
from kubernetes.client import V1ResourceRequirements
api_version = constants.KFSERVING_GROUP + '/' + constants.KFSERVING_VERSION
default_endpoint_spec = V1alpha2EndpointSpec(
predictor=V1alpha2PredictorSpec(
service_account_name='minio-kf-sa',
tensorflow=V1alpha2TensorflowSpec(
storage_uri='s3://'+MINIO_MODEL_BUCKET+'/'+ CIFAR10_MODEL_PATH,
resources=V1ResourceRequirements(
requests={'cpu':'100m','memory':'1Gi'},
limits={'cpu':'100m', 'memory':'1Gi'})),
logger=V1alpha2Logger(
mode='all'
)),
explainer=V1alpha2ExplainerSpec(
service_account_name='minio-kf-sa',
alibi=V1alpha2AlibiExplainerSpec(
type='AnchorImages',
storage_uri='s3://'+MINIO_MODEL_BUCKET+'/'+ EXPLAINER_MODEL_PATH,
resources=V1ResourceRequirements(
requests={'cpu':'100m','memory':'1Gi'},
limits={'cpu':'100m', 'memory':'1Gi'}))))
isvc = V1alpha2InferenceService(api_version=api_version,
kind=constants.KFSERVING_KIND,
metadata=client.V1ObjectMeta(
name='kf-cifar10', namespace=DEPLOY_NAMESPACE),
spec=V1alpha2InferenceServiceSpec(default=default_endpoint_spec))
# + tags=[]
KFServing = KFServingClient()
KFServing.create(isvc)
# + tags=[]
KFServing.get('kf-cifar10', namespace=DEPLOY_NAMESPACE, watch=True, timeout_seconds=240)
# + [markdown] tags=[]
# ## Test Model and Explainer
# + tags=["block:test_model_and_explainer", "prev:deploy_model"]
def test_model():
idx=10
test_example=X_test[idx:idx+1].tolist()
payload='{"instances":'+f"{test_example}"+' }'
cmd=f"""curl -v -d '{payload}' \
-H "Host: kf-cifar10.admin.example.com" \
-H "Content-Type: application/json" \
http://kfserving-ingressgateway.istio-system/v1/models/kf-cifar10:predict
"""
ret = Popen(cmd, shell=True,stdout=PIPE)
raw = ret.stdout.read().decode("utf-8")
print(raw)
res=json.loads(raw)
arr=np.array(res["predictions"])
X = X_test[idx].reshape(1, 32, 32, 3)
plt.imshow(X.reshape(32, 32, 3))
plt.axis('off')
plt.show()
print("class:",class_names[y_test[idx][0]])
print("prediction:",class_names[arr[0].argmax()])
ok = False
while not ok:
try:
test_model()
ok = True
except:
print("Failed calling model, sleeping")
time.sleep(2)
# + [markdown] tags=[]
# Make an explanation request
# + tags=[]
idx=1
test_example=X_test[idx:idx+1].tolist()
payload='{"instances":'+f"{test_example}"+' }'
cmd=f"""curl -v -d '{payload}' \
-H "Host: kf-cifar10.admin.example.com" \
-H "Content-Type: application/json" \
http://kfserving-ingressgateway.istio-system/v1/models/kf-cifar10:explain
"""
ret = Popen(cmd, shell=True,stdout=PIPE)
raw = ret.stdout.read().decode("utf-8")
res=json.loads(raw)
plt.imshow(np.array(explanation["anchor"]))
# + [markdown] tags=[]
# ## Deploy KNative Eventing Event Display
# + tags=["block:deploy_event_display", "prev:train_drift_detector", "prev:train_outlier_detector", "prev:test_model_and_explainer"]
event_display=f"""apiVersion: apps/v1
kind: Deployment
metadata:
name: event-display
namespace: {DEPLOY_NAMESPACE}
spec:
replicas: 1
selector:
matchLabels: &labels
app: event-display
template:
metadata:
labels: *labels
spec:
containers:
- name: helloworld-go
# Source code: https://github.com/knative/eventing-contrib/tree/master/cmd/event_display
image: gcr.io/knative-releases/knative.dev/eventing-contrib/cmd/event_display@sha256:f4628e97a836c77ed38bd3b6fd3d0b06de4d5e7db6704772fe674d48b20bd477
---
kind: Service
apiVersion: v1
metadata:
name: event-display
namespace: {DEPLOY_NAMESPACE}
spec:
selector:
app: event-display
ports:
- protocol: TCP
port: 80
targetPort: 8080
---
apiVersion: eventing.knative.dev/v1alpha1
kind: Trigger
metadata:
name: cifar10-outlier-display
namespace: {DEPLOY_NAMESPACE}
spec:
broker: default
filter:
attributes:
type: org.kubeflow.serving.inference.outlier
subscriber:
ref:
apiVersion: v1
kind: Service
name: event-display
---
apiVersion: eventing.knative.dev/v1alpha1
kind: Trigger
metadata:
name: cifar10-drift-display
namespace: {DEPLOY_NAMESPACE}
spec:
broker: default
filter:
attributes:
type: org.kubeflow.serving.inference.drift
subscriber:
ref:
apiVersion: v1
kind: Service
name: event-display
"""
with open("event_display.yaml","w") as f:
f.write(event_display)
run("kubectl apply -f event_display.yaml", shell=True)
# + tags=[]
run(f"kubectl rollout status -n {DEPLOY_NAMESPACE} deploy/event-display -n {DEPLOY_NAMESPACE}", shell=True)
# + [markdown] tags=[]
# ## Deploy KFServing Outlier Detector
# + tags=["block:deploy_outlier_detector", "prev:deploy_event_display"]
outlier_yaml=f"""apiVersion: serving.knative.dev/v1
kind: Service
metadata:
name: cifar10-outlier
namespace: {DEPLOY_NAMESPACE}
spec:
template:
metadata:
annotations:
autoscaling.knative.dev/minScale: "1"
spec:
containers:
- image: seldonio/alibi-detect-server:1.2.1
imagePullPolicy: IfNotPresent
args:
- --model_name
- cifar10od
- --protocol
- tensorflow.http
- --storage_uri
- s3://{MINIO_MODEL_BUCKET}/{OUTLIER_MODEL_PATH}
- --reply_url
- http://default-broker
- --event_type
- org.kubeflow.serving.inference.outlier
- --event_source
- org.kubeflow.serving.cifar10od
- OutlierDetector
envFrom:
- secretRef:
name: seldon-init-container-secret
"""
with open("outlier.yaml","w") as f:
f.write(outlier_yaml)
run("kubectl apply -f outlier.yaml", shell=True)
# + tags=[]
trigger_outlier_yaml=f"""apiVersion: eventing.knative.dev/v1alpha1
kind: Trigger
metadata:
name: cifar10-outlier-trigger
namespace: {DEPLOY_NAMESPACE}
spec:
filter:
sourceAndType:
type: org.kubeflow.serving.inference.request
subscriber:
ref:
apiVersion: serving.knative.dev/v1
kind: Service
name: cifar10-outlier
"""
with open("outlier_trigger.yaml","w") as f:
f.write(trigger_outlier_yaml)
run("kubectl apply -f outlier_trigger.yaml", shell=True)
# + tags=[]
run(f"kubectl rollout status -n {DEPLOY_NAMESPACE} deploy/$(kubectl get deploy -l serving.knative.dev/service=cifar10-outlier -o jsonpath='{{.items[0].metadata.name}}' -n {DEPLOY_NAMESPACE})", shell=True)
# + [markdown] tags=[]
# ## Test KFServing Outlier Detection
# + tags=["block:test_oulier_detection", "prev:deploy_outlier_detector"]
idx = 1
X = X_train[idx:idx+1]
# + tags=[]
np.random.seed(0)
X_mask, mask = apply_mask(X.reshape(1, 32, 32, 3),
mask_size=(10,10),
n_masks=1,
channels=[0,1,2],
mask_type='normal',
noise_distr=(0,1),
clip_rng=(0,1))
# + tags=[]
def predict():
test_example=X_mask.tolist()
payload='{"instances":'+f"{test_example}"+' }'
cmd=f"""curl -v -d '{payload}' \
-H "Host: kf-cifar10.admin.example.com" \
-H "Content-Type: application/json" \
http://kfserving-ingressgateway.istio-system/v1/models/kf-cifar10:predict
"""
ret = Popen(cmd, shell=True,stdout=PIPE)
raw = ret.stdout.read().decode("utf-8")
print(raw)
res=json.loads(raw)
arr=np.array(res["predictions"])
plt.imshow(X_mask.reshape(32, 32, 3))
plt.axis('off')
plt.show()
print("class:",class_names[y_train[idx][0]])
print("prediction:",class_names[arr[0].argmax()])
# + tags=[]
def get_outlier_event_display_logs():
cmd=f"kubectl logs $(kubectl get pod -l app=event-display -o jsonpath='{{.items[0].metadata.name}}' -n {DEPLOY_NAMESPACE}) -n {DEPLOY_NAMESPACE}"
ret = Popen(cmd, shell=True,stdout=PIPE)
res = ret.stdout.read().decode("utf-8").split("\n")
data= []
for i in range(0,len(res)):
if res[i] == 'Data,':
j = json.loads(json.loads(res[i+1]))
if "is_outlier"in j["data"].keys():
data.append(j)
if len(data) > 0:
return data[-1]
else:
return None
j = None
while j is None:
predict()
print("Waiting for outlier logs, sleeping")
time.sleep(2)
j = get_outlier_event_display_logs()
print(j)
print("Outlier",j["data"]["is_outlier"]==[1])
# + [markdown] tags=[]
# ## Deploy KFServing Drift Detector
# + tags=["block:deploy_drift_detector", "prev:test_oulier_detection"]
drift_yaml=f"""apiVersion: serving.knative.dev/v1
kind: Service
metadata:
name: cifar10-drift
namespace: {DEPLOY_NAMESPACE}
spec:
template:
metadata:
annotations:
autoscaling.knative.dev/minScale: "1"
spec:
containers:
- image: seldonio/alibi-detect-server:1.2.2-dev
imagePullPolicy: IfNotPresent
args:
- --model_name
- cifar10cd
- --protocol
- tensorflow.http
- --storage_uri
- s3://{MINIO_MODEL_BUCKET}/{DRIFT_MODEL_PATH}
- --reply_url
- http://default-broker
- --event_type
- org.kubeflow.serving.inference.drift
- --event_source
- org.kubeflow.serving.cifar10cd
- DriftDetector
- --drift_batch_size
- '500'
envFrom:
- secretRef:
name: seldon-init-container-secret
"""
with open("drift.yaml","w") as f:
f.write(drift_yaml)
run("kubectl apply -f drift.yaml", shell=True)
# + tags=[]
trigger_outlier_yaml=f"""apiVersion: eventing.knative.dev/v1alpha1
kind: Trigger
metadata:
name: cifar10-drift-trigger
namespace: {DEPLOY_NAMESPACE}
spec:
filter:
sourceAndType:
type: org.kubeflow.serving.inference.request
subscriber:
ref:
apiVersion: serving.knative.dev/v1
kind: Service
name: cifar10-drift
"""
with open("outlier_trigger.yaml","w") as f:
f.write(trigger_outlier_yaml)
run("kubectl apply -f outlier_trigger.yaml", shell=True)
# + tags=[]
run(f"kubectl rollout status -n {DEPLOY_NAMESPACE} deploy/$(kubectl get deploy -l serving.knative.dev/service=cifar10-drift -o jsonpath='{{.items[0].metadata.name}}' -n {DEPLOY_NAMESPACE})", shell=True)
# + [markdown] tags=[]
# ## Test KFServing Drift Detector
# + tags=["block:test_drift_detector", "prev:deploy_drift_detector"]
def show(X):
plt.imshow(X.reshape(32, 32, 3))
plt.axis('off')
plt.show()
# + tags=[]
from alibi_detect.datasets import fetch_cifar10c, corruption_types_cifar10c
corruption = ['motion_blur']
X_corr, y_corr = fetch_cifar10c(corruption=corruption, severity=5, return_X_y=True)
X_corr = X_corr.astype('float32') / 255
# + tags=[]
show(X_corr[0])
show(X_corr[1])
show(X_corr[2])
# + tags=[]
def predict(X):
test_example=X.tolist()
payload='{"instances":'+f"{test_example}"+' }'
with open("payload.json","w") as f:
f.write(payload)
cmd=f"""curl -d @./payload.json \
-H "Host: kf-cifar10.admin.example.com" \
-H "Content-Type: application/json" \
http://kfserving-ingressgateway.istio-system/v1/models/kf-cifar10:predict
"""
run(cmd, shell=True)
# + tags=[]
def get_drift_event_display_logs():
cmd=f"kubectl logs $(kubectl get pod -l app=event-display -o jsonpath='{{.items[0].metadata.name}}' -n {DEPLOY_NAMESPACE}) -n {DEPLOY_NAMESPACE}"
ret = Popen(cmd, shell=True,stdout=PIPE)
res = ret.stdout.read().decode("utf-8").split("\n")
data= []
for i in range(0,len(res)):
if res[i] == 'Data,':
j = json.loads(json.loads(res[i+1]))
if "is_drift" in j["data"].keys():
data.append(j)
if len(data) > 0:
return data[-1]
else:
return None
j = None
for i in range(0,1000,50):
X = X_corr[i:i+50]
predict(X)
print("Waiting for drift logs, sleeping")
time.sleep(2)
j = get_drift_event_display_logs()
if j is not None:
break
print(j)
print("Drift",j["data"]["is_drift"]==1)
# + [markdown] tags=[]
# ## Clean up
# + tags=["skip"]
run(f"kubectl delete inferenceservice kf-cifar10 -n {DEPLOY_NAMESPACE}", shell=True)
run(f"kubectl delete ksvc cifar10-outlier -n {DEPLOY_NAMESPACE}", shell=True)
run(f"kubectl delete ksvc cifar10-drift -n {DEPLOY_NAMESPACE}", shell=True)
run(f"kubectl delete trigger --all -n {DEPLOY_NAMESPACE}", shell=True)
run(f"kubectl delete sa minio-kf-sa -n {DEPLOY_NAMESPACE}", shell=True)
run(f"kubectl delete secret seldon-init-container-secret -n {DEPLOY_NAMESPACE}", shell=True)
run(f"kubectl delete secret cifar10-kf-secret -n {DEPLOY_NAMESPACE}", shell=True)
run(f"kubectl delete deployment event-display -n {DEPLOY_NAMESPACE}", shell=True)
run(f"kubectl delete svc event-display -n {DEPLOY_NAMESPACE}", shell=True)
# + tags=[]
|
samples/contrib/e2e-outlier-drift-explainer/kfserving/kfserving_e2e_cifar10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Collection for Hype-Machine
# #### Setup and Dependancies
# +
import os
import sys
import json
import math
import datetime as dt
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LinearRegression
from nltk.metrics import *
from linearmodels import PanelOLS
from linearmodels import RandomEffects
import warnings
import requests
from bs4 import BeautifulSoup as bs
from selenium import webdriver
import time
from datetime import datetime, timedelta
import re
import ast
warnings.filterwarnings('ignore')
# %matplotlib inline
# -
# #### Directories and Names
# +
rep16_candidates = [
("Jeb", "Bush"),
("Ben", "Carson"),
("Chris", "Christie"),
("Ted", "Cruz"),
("Carly", "Fiorina"),
("Jim", "Gilmore"),
("Lindsey", "Graham"),
("Mike", "Huckabee"),
("Bobby", "Jindal"),
("John", "Kasich"),
("George", "Pataki"),
("Randal", "Paul"),
("Rick", "Perry"),
("Marco", "Rubio"),
("Rick", "Santorum"),
("Donald", "Trump"),
("Scott", "Walker")
]
dem20_candidates = [
('Michael', 'Bennet'),
('Joe', 'Biden'),
('Corey', 'Booker'),
('Steve', 'Bullock'),
('Pete', 'Buttigieg'),
('Julian', 'Castro'),
('Bill', '<NAME>'),
('John', 'Delaney'),
('Tulsi', 'Gabbard'),
('Kirsten', 'Gillibrand'),
('Kamala', 'Harris'),
('John', 'Hickenlooper'),
('Jay', 'Inslee'),
('Amy', 'Klobuchar'),
('Beto', 'Orourke'),
('Andrew', 'Yang'),
('Bernie', 'Sanders'),
('Eric', 'Swalwell'),
('Elizabeth', 'Warren'),
('Marianne', 'Williamson'),
('Andrew', 'Yang')
]
# make 2016 republican fec folders
rep16_fec_path = os.path.join("..","data","fec_new","2016")
if(not os.path.isdir(rep16_fec_path)):
os.mkdir(rep16_fec_path)
rep16_fec_path = os.path.join(rep16_fec_path,"republican")
if(not os.path.isdir(rep16_fec_path)):
os.mkdir(rep16_fec_path)
for candid in rep16_candidates:
candid_path = os.path.join(rep16_fec_path,candid[1].lower())
if(not os.path.isdir(candid_path)):
os.mkdir(candid_path)
# make 2020 democrat fec folders
dem20_fec_path = os.path.join("..","data","fec_new","2020")
if(not os.path.isdir(dem20_fec_path)):
os.mkdir(dem20_fec_path)
dem20_fec_path = os.path.join(dem20_fec_path,"democrat")
if(not os.path.isdir(dem20_fec_path)):
os.mkdir(dem20_fec_path)
for candid in dem20_candidates:
candid_path = os.path.join(dem20_fec_path,candid[1].lower())
if(not os.path.isdir(candid_path)):
os.mkdir(candid_path)
# polls folders
rep16_polls_path = os.path.join("..","data","polls")
if(not os.path.isdir(rep16_polls_path)):
os.mkdir(rep16_polls_path)
rep16_polls_path = os.path.join(rep16_polls_path,"2016")
if(not os.path.isdir(rep16_polls_path)):
os.mkdir(rep16_polls_path)
dem20_polls_path = os.path.join("..","data","polls","2020")
if(not os.path.isdir(dem20_polls_path)):
os.mkdir(dem20_polls_path)
# gdelt folders
rep16_gdelt_path = os.path.join("..","data","gdelt")
if(not os.path.isdir(rep16_gdelt_path)):
os.mkdir(rep16_gdelt_path)
rep16_gdelt_path = os.path.join(rep16_gdelt_path,"2016")
if(not os.path.isdir(rep16_gdelt_path)):
os.mkdir(rep16_gdelt_path)
rep16_gdelt_path = os.path.join(rep16_gdelt_path,"republican")
if(not os.path.isdir(rep16_gdelt_path)):
os.mkdir(rep16_gdelt_path)
dem20_gdelt_path = os.path.join(os.path.join("..","data","gdelt","2020"))
if(not os.path.isdir(dem20_gdelt_path)):
os.mkdir(dem20_gdelt_path)
dem20_gdelt_path = os.path.join(dem20_gdelt_path,"democrat")
if(not os.path.isdir(dem20_gdelt_path)):
os.mkdir(dem20_gdelt_path)
# -
# ## Collect NLP Data
# +
nlp_raw = {}
#nlp_dir = os.path.join("..","data","candidate_aggregation")
nlp_dir = os.path.join("..","data","candidate_aggregation 2")
for ii,fname in enumerate(os.listdir(nlp_dir)):
tmp_df = pd.read_csv(os.path.join(nlp_dir, fname))
candidate = os.path.splitext(fname)[0]
nlp_raw[candidate] = {}
for ii,row in tmp_df.iterrows():
daily_nlp = {}
nlp_list = ast.literal_eval(row["topic_titles"])
for tup in nlp_list:
daily_nlp[tup[0]] = tup[1]
nlp_raw[candidate][row["day"]] = daily_nlp
dem20_nlp = pd.DataFrame.from_dict(
{(i,j): nlp_raw[i][j] for i in nlp_raw.keys() for j in nlp_raw[i].keys()},
orient='index').fillna(0)
dem20_nlp = dem20_nlp.reset_index()
dem20_nlp = dem20_nlp.rename({"level_0":"candidate", "level_1":"date"}, axis="columns")
dem20_nlp["date"] = pd.to_datetime(dem20_nlp["date"], format='%Y-%m-%d')
dem20_nlp.head()
# -
# ## Collect Polling Data
# #### 2016 Polling Data
# There are very few good aggregates of 2016 primary polls, and for consistency we'd like to use 538 as a data source if possible. We pulled the code below from [a github repository from sgodfrey66](https://github.com/sgodfrey66/Polls_and_press/blob/master/code/Poll_data_2016.ipynb). The code scrapes [this 538 webpage](https://projects.fivethirtyeight.com/election-2016/national-primary-polls/republican/).
# +
class Scrape538PollData:
# Attributes of the data retrieval
url = 'https://projects.fivethirtyeight.com/election-2016/national-primary-polls/republican/'
chrome_driver = os.path.join(os.getcwd(),"..","data","polls","chromedriver")
status_code_ = None
html_ = None
soup_ = None
candidates_ = []
polls_ = []
# Initialization method
def __init__(self, url = None, chrome_driver = None):
# If url != None then reset self.url
if url:
self.url = url
# If chrome_driver != None reset self.chrome_driver
if chrome_driver:
self.chrome_driver = chrome_driver
# method to collect data from posts
def collect_page_data(self):
# Set the selenium driver
try:
driver = webdriver.Chrome(self.chrome_driver)
driver.get(self.url)
except:
self.status_code_ = driver.error_handler.check_response
raise ValueError('Error retrieving the web page; see WebExceptionError for details.')
# Find the read more polls button and click it
driver.find_element_by_css_selector('.more-polls').click()
# Return the html of the new page
self.html_ = driver.page_source.encode('utf-8')
# Return the soup version of the page
self.soup_ = bs(self.html_, 'lxml')
# method to collect data from posts
def extract_polls(self):
# Check to see that self.soup_ has data, if not run collect_page_data
if self.soup_ == None or len(self.soup_) == 0:
self.collect_page_data()
# Find the list of candidates
table = self.soup_.find('table')
self.candidates_ = [c.text for c in table.find_all('th', {'class':'th th-rotate'})]
# Find the poll data
body = self.soup_.find('tbody')
polls = body.find_all('tr',{'class': 't-row'})
# For each poll extract information related to the poll
for poll in polls:
pl_d = {}
pl_d['dates'] = poll.find('td', {'class':'t-dates'}).text
pl_d['pollster_url'] = poll.find('a', href = True)
pl_d['pollster'] = poll.find('td', {'class': 't-pollster t-left-margin'}).text
pl_d['sample'] = poll.find('td', {'class': 't-sample t-left-margin t-right-align only-full'}).text
pl_d['weight'] = poll.find('td', \
{'class': 't-weight t-left-margin t-right-margin double-l-margin t-right-border-dark'}).text
try:
pl_d['leader'] = poll.find('td', \
{'class':'t-leader t-left-margin t-right-margin only-full color-text-rep'}).text
except:
pl_d['leader'] = ''
# Get the odds for each candidate except for the last candidate in the table
for i, odds in enumerate(poll.find_all('td', {'class':'t-center-align td-cand-odds td-block t-right-border'})):
# This tag is present if a value exists in the poll for that candidate
if odds.find('div', {'class':'t-cand-odds heat-map-blocks'}):
pl_d[self.candidates_[i]] = float(odds.text.replace('%','').strip())/100
# Get the odds for the last candidate
odds = poll.find('td', {'class':'t-center-align td-cand-odds td-block'})
# This tag is present if a value exists in the poll for that candidate
if odds.find('div', {'class':'t-cand-odds heat-map-blocks'}):
pl_d[self.candidates_[len(self.candidates_) - 1]] = float(odds.text.replace('%','').strip())/100
self.polls_.append(pl_d)
'''
# Walk through DataFrame to assign dates
def assign_dates(df = None):
# Loop through each row in this DataFrame to create
# a datetime object for the beginnging and end of the poll
start_year = '2016'
end_year = '2016'
for idx in df.index:
# Find the months and set them equal to start and end month
months = re.findall(r'[\w]{3}',df.loc[idx,'dates'])
if len(months)==1:
start_month = months[0]
end_month = months[0]
else:
start_month = months[0]
end_month = months[1]
# Find the months and set them equal to start and end month
dates = re.findall(r'[\d]{1,}',df.loc[idx,'dates'])
if len(dates)==1:
start_date = dates[0]
end_date = dates[0]
else:
start_date = dates[0]
end_date = dates[1]
# Figure out if the year needs to be changed
if start_month=='Dec' and end_month=='Jan':
start_year = '2015'
end_year = '2016'
elif start_month=='Jan' and end_month=='Jan':
start_year = '2016'
end_year = '2016'
elif start_month=='Dec' and end_month=='Dec':
start_year = '2015'
end_year = '2015'
start_time=start_year+' '+start_month+' '+start_date
end_time=end_year+' '+end_month+' '+end_date
dt_start = datetime.strptime(start_time,'%Y %b %d')
dt_end = datetime.strptime(end_time,'%Y %b %d')
df.loc[idx,'start_time'] = dt_start
df.loc[idx,'end_time'] = dt_end
'''
# Instantiate and a poll object and get poll data
rep16_polls = Scrape538PollData()
rep16_polls.extract_polls()
# -
# The scraped data needs munged to be compatible with the 2020 dataset.
# +
def date_range_str_to_tuple(arg):
arg = re.sub('[^0-9a-zA-Z]+', ' ', arg)
arg = arg.split()
if(len(arg)==3):
mm = arg[0]
dd = arg[2]
elif(len(arg)==4):
mm = arg[2]
dd = arg[3]
return (dd,mm)
# drop columns and filter data
rep16_polls_df = pd.DataFrame(rep16_polls.polls_)
rep16_polls_df = rep16_polls_df.loc[rep16_polls_df["pollster"]=="Morning Consult"]
rep16_polls_df = rep16_polls_df.drop(["leader","pollster_url","weight","sample","pollster"], axis=1)
rep16_polls_df = rep16_polls_df.rename({'dates':'date'}, axis='columns')
# date conversion
yy = 2016
for index, row in rep16_polls_df.iterrows():
dd, mm = date_range_str_to_tuple(row["date"])
if(mm=="Dec"):
yy = 2015
rep16_polls_df.loc[index, "date"] = datetime.strptime("{:2d} {:s} {:02d}".format(yy,mm,int(dd)), "%Y %b %d")
rep16_polls_df = rep16_polls_df.set_index("date")
rep16_polls_df.to_csv(os.path.join(rep16_polls_path,"president_primary_polls.csv"))
rep16_polls_df.head()
# -
# #### 2020 Polling Data
# 538 aggregates the 2020 polling data [here](https://data.fivethirtyeight.com/) under "Latest Polls", which makes it easy to import.
dem20_polls_df = pd.read_csv(os.path.join(dem20_polls_path,"president_primary_polls.csv"))
dem20_polls_df = dem20_polls_df.rename({"created_at":"date"}, axis='columns')
dem20_polls_df = dem20_polls_df.loc[dem20_polls_df["cycle"]==2020]
dem20_polls_df = dem20_polls_df.loc[dem20_polls_df["stage"]=="primary"]
dem20_polls_df = dem20_polls_df.loc[dem20_polls_df["party"]=="DEM"]
dem20_polls_df = dem20_polls_df.loc[dem20_polls_df["pollster"]=="Morning Consult"]
dem20_polls_df = dem20_polls_df.groupby(by=["candidate_name","date"])["pct"].mean()
dem20_polls_df = dem20_polls_df.unstack(level="candidate_name")
dem20_polls_df.head()
dem20_polls_df.columns
# ## Collect Donnation Data
# #### 2016 Donnations
# find the path to each fec file, store paths in a nested dict
rep16_fec_file_map = {}
for cand_dir in os.listdir(rep16_fec_path):
if(cand_dir[0]!="."):
rep16_fec_file_map[cand_dir] = {}
for csv_path in os.listdir(os.path.join(rep16_fec_path,cand_dir)):
if(csv_path.find("schedule_a")>=0):
rep16_fec_file_map[cand_dir]["donations"] = \
os.path.join(rep16_fec_path,cand_dir,csv_path)
elif(csv_path.find("schedule_b")>=0):
rep16_fec_file_map[cand_dir]["spending"] = \
os.path.join(rep16_fec_path,cand_dir,csv_path)
print(json.dumps(rep16_fec_file_map, indent=4))
# #### 2020 Donnations
# find the path to each fec file, store paths in a nested dict
dem20_fec_file_map = {}
for cand_dir in os.listdir(dem20_fec_path):
if(cand_dir[0]!="."):
dem20_fec_file_map[cand_dir] = {}
for csv_path in os.listdir(os.path.join(dem20_fec_path,cand_dir)):
if(csv_path.find("schedule_a")>=0):
dem20_fec_file_map[cand_dir]["donations"] = \
os.path.join(dem20_fec_path,cand_dir,csv_path)
elif(csv_path.find("schedule_b")>=0):
dem20_fec_file_map[cand_dir]["spending"] = \
os.path.join(dem20_fec_path,cand_dir,csv_path)
print(json.dumps(dem20_fec_file_map, indent=4))
# ## Collect Cable TV Mentions
# Uses the [gdelt TV API](https://blog.gdeltproject.org/gdelt-2-0-television-api-debuts/) to collect mentions of primary cadidates on cable television.
# #### Generic API Request
def cable_mentions(candidate_list, start_year=2019, end_year=2019):
# api setup
Market = ['National']
tv_api_base = "https://api.gdeltproject.org/api/v2/tv/"
tv_api_params = {
"mode":"timelinevol",
"format":"csv",
"STARTDATETIME":"{:04d}0101000000".format(start_year),
"ENDDATETIME":"{:04d}1231115959".format(end_year)
}
tv_api_params_string = "&".join(["{:s}={:s}".format(key,val) for key,val in tv_api_params.items()])
# send requests and combine into dataframe
data = pd.DataFrame()
for ii, candid in enumerate(candidate_list):
last_name = candid[1]
full_name = " ".join(candid)
print("\r{:0.0f}% {:s}{:s}".format(100*ii/len(candidate_list), full_name, " "*20), end="")
for location in Market:
api_call = "{:s}tv?query=%22{:s}%22%20market:%22{:s}%22&{:s}".format(
tv_api_base,
last_name,
location,
tv_api_params_string
)
try:
temp_data = pd.read_csv(api_call)
temp_data['Candidate'] = full_name
temp_data['Market'] = location
data = data.append(temp_data, ignore_index = True)
except Exception as e:
print(e, end="")
print("\r100% {:s}".format(" "*50))
# fix date column name
find_colname_str = "date"
for colname in data.columns:
if(colname[:len(find_colname_str)].lower()==find_colname_str):
found_colname_str = colname
data.rename({found_colname_str:find_colname_str.title()},axis="columns", inplace=True)
data["Date"] = pd.to_datetime(data["Date"], format='%Y-%m-%d')
# mung tv data
data = data[["Candidate","Date","Series","Value"]]
tv_mentions = data.groupby(["Candidate","Date"])["Value"].mean()
tv_mentions = tv_mentions.unstack(level="Candidate")
return tv_mentions
# #### 2016 Mentions
rep16_mentions = cable_mentions(rep16_candidates, 2015, 2016)
rep16_mentions.to_csv(os.path.join(rep16_gdelt_path,"tv_mentions.csv"))
# #### 2020 Mentions
dem20_mentions = cable_mentions(dem20_candidates)
dem20_mentions.to_csv(os.path.join(dem20_gdelt_path,"tv_mentions.csv"))
# ## Combining Datasets
# #### 2016 Name Mapping
rep16_name_mapping = {}
poll_names = rep16_polls_df.columns
media_names = list(rep16_mentions.columns)
for candid in rep16_fec_file_map.keys():
# poll data
comparison_scores = [edit_distance(x[-len(candid):].lower(), candid.lower()) for x in poll_names]
val, idx = min((val, idx) for (idx, val) in enumerate(comparison_scores))
poll_name_map = poll_names[idx]
# media data
comparison_scores = [edit_distance(x[-len(candid):].lower(), candid.lower()) for x in media_names]
val, idx = min((val, idx) for (idx, val) in enumerate(comparison_scores))
media_name_map = media_names[idx]
# output
rep16_name_mapping[candid] = (poll_name_map, media_name_map)
print("{:s} -> {:s} (score: {:d})".format(candid, str(rep16_name_mapping[candid]), val))
# #### 2020 Name Mapping
# +
dem20_name_mapping = {}
dem20_name_mapping_scores = {}
poll_names = dem20_polls_df.columns
media_names = list(dem20_mentions.columns)
nlp_names = list(dem20_nlp["candidate"].unique())
for candid in dem20_fec_file_map.keys():
# poll data
comparison_scores = [edit_distance(x[-len(candid):].lower(), candid.lower()) for x in poll_names]
poll_score, idx = min((val, idx) for (idx, val) in enumerate(comparison_scores))
poll_name_map = poll_names[idx]
# media data
comparison_scores = [edit_distance(x[-len(candid):].lower(), candid.lower()) for x in media_names]
media_score, idx = min((val, idx) for (idx, val) in enumerate(comparison_scores))
media_name_map = media_names[idx]
# nlp data
comparison_scores = [edit_distance(x[-len(candid):].lower(), candid.lower()) for x in nlp_names]
nlp_score, idx = min((val, idx) for (idx, val) in enumerate(comparison_scores))
nlp_name_map = nlp_names[idx]
# output
dem20_name_mapping[candid] = (poll_name_map, media_name_map, nlp_name_map)
dem20_name_mapping_scores[candid] = (poll_score, media_score, nlp_score)
# special case
dem20_name_mapping["ryan"] = ("<NAME>", "", "")
dem20_name_mapping["biden"] = ("<NAME>.", dem20_name_mapping["biden"][1], dem20_name_mapping["biden"][2])
# output
print("fec_name -> polling_name (edit_distance), media_name (edit_distance), nlp_name (edit distance)")
for candid in sorted(dem20_name_mapping.keys()):
print("{:s} -> {:s} ({:d}), {:s} ({:d}), {:s} ({:d})".format(
candid,
dem20_name_mapping[candid][0],
dem20_name_mapping_scores[candid][0],
dem20_name_mapping[candid][1],
dem20_name_mapping_scores[candid][1],
dem20_name_mapping[candid][2],
dem20_name_mapping_scores[candid][2]))
# -
# #### Joins and Normalization
# +
def make_normalized_col(df, col_name):
new_col_name = "{:s}_normalized".format(col_name)
daily_sums = df.groupby("date")[col_name].sum()
dates = df.index.get_level_values('date')
df[new_col_name] = daily_sums.loc[dates].values
df[new_col_name] = df[col_name]/df[new_col_name]
def read_donnation_csv(path, candid):
df = pd.read_csv(path)
df["contribution_receipt_date"] = pd.to_datetime(df["contribution_receipt_date"]).dt.date
df = df.loc[df["entity_type"]=="IND"]
df = df.loc[np.logical_and(df["contribution_receipt_amount"]>0,df["contribution_receipt_amount"]<=2800)]
# donation count
df_count = df.groupby(by="contribution_receipt_date")["contribution_receipt_amount"].count()
df_count.name = "individual_donations"
# donation sum
df_sum = df.groupby(by="contribution_receipt_date")["contribution_receipt_amount"].sum()
df_sum.name = "individual_donation_amount"
# donations under $200
df = df.loc[df["contribution_receipt_amount"]<=200]
df_small = df.groupby(by="contribution_receipt_date")["contribution_receipt_amount"].count()
df_small.name = "small_donor_count"
df = pd.concat([df_count, df_sum, df_small], axis=1)
df["candidate"] = candid
return df
def combine_campaign(
fec_paths,
polls,
tv_mentions,
name_mapping,
nlp = None):
dataset = pd.DataFrame()
for candid in fec_paths.keys():
if("donations" in fec_paths[candid].keys()):
# donnations data
df = read_donnation_csv(fec_paths[candid]["donations"], candid)
# polling data
if(name_mapping[candid][0] in polls.columns):
candid_polls = polls[name_mapping[candid][0]]
candid_polls.name = "polling_percentage"
candid_polls.index = pd.to_datetime(candid_polls.index).date
df = df.join(candid_polls, how="left")
df["polling_percentage"] = df["polling_percentage"].interpolate(limit_direction='both').fillna(0.0)
df = df.rename_axis("date").reset_index()
# process media dataset
df["date"] = pd.to_datetime(df["date"], format='%Y-%m-%d')
if(name_mapping[candid][1] in tv_mentions.columns):
candid_tv = tv_mentions[name_mapping[candid][1]]
candid_tv.name = "tv_mentions"
df = df.merge(candid_tv, how="left", left_on="date", right_index=True)
# process nlp data
if(nlp is not None):
nlp_candid = nlp.loc[nlp["candidate"]==name_mapping[candid][2]]
nlp_candid = nlp_candid.drop("candidate", axis=1)
df = df.merge(nlp_candid, how="left", left_on="date", right_on="date")
# append to main df
dataset = dataset.append(df)
# munge data
#dataset = dataset.reset_index()
#dataset = dataset.rename(columns={"index": "date"})
dataset = dataset.set_index(["candidate","date"])
dataset["individual_donations"] = dataset["individual_donations"].fillna(0.0)
make_normalized_col(dataset, "individual_donations")
dataset["individual_donation_amount"] = dataset["individual_donation_amount"].fillna(0.0)
make_normalized_col(dataset, "individual_donation_amount")
dataset["small_donor_count"] = dataset["small_donor_count"].fillna(0.0)
make_normalized_col(dataset, "small_donor_count")
dataset["tv_mentions"] = dataset["tv_mentions"].fillna(0.0)
make_normalized_col(dataset, "tv_mentions")
dataset["polling_percentage"] = dataset["polling_percentage"].fillna(0.0)
make_normalized_col(dataset, "polling_percentage")
return dataset
def weekly_aggregate(df):
df['date'] = pd.to_datetime(df['date']) - pd.to_timedelta(7, unit='d')
df = df.groupby(['candidate', pd.Grouper(key='date', freq='W-TUE')]).mean()
df = df.reset_index()
return df
# +
dem20_df = combine_campaign(dem20_fec_file_map, dem20_polls_df, dem20_mentions, dem20_name_mapping, dem20_nlp)
dem20_weekly_df = weekly_aggregate(dem20_df.reset_index())
# write data
dem20_df.to_csv("../data/dem20_dataset.csv")
dem20_df.to_pickle("../data/dem20_dataset.pkl")
dem20_weekly_df.to_csv("../data/dem20_weekly_dataset.csv")
# check data
dem20_df.head(15)
# +
rep16_df = combine_campaign(rep16_fec_file_map, rep16_polls_df, rep16_mentions, rep16_name_mapping)
# write data
rep16_df.to_csv("../data/rep16_dataset.csv")
rep16_df.to_pickle("../data/rep16_dataset.pkl")
rep16_df.head(15)
# -
|
src/Data_Collection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
df = pd.read_csv('data/camh/simdata_long_n100.csv')
# -
df = df.drop(['time'],axis=1)
df.describe()
df.fillna(0)
df.head(20)
df.describe()
df.loc[df.id=='sub69']
# +
max_timestamps = 0
for id in df.id.values:
if df.loc[df.id == id].iloc[:, 1].shape[0] > max_timestamps:
max_timestamps = df.loc[df.id == id].iloc[:, 1].shape[0]
# -
max_timestamps
# +
import numpy as np
tt = np.arange(max_timestamps)
# -
arr = np.array([1,2,3])
np.stack([arr for _ in range(5)], axis=0)
vals = df.loc[df.id == 'sub1'].iloc[:, 1:].values
dup = np.stack([vals[-1] for _ in range(max_timestamps-vals.shape[0])], axis=0)
np.vstack((vals, dup)).shape
masks = vals
masks
masks[~np.isnan(masks)]=1
masks[np.isnan(masks)]=0
masks
# +
import torch
patient = 'sub1'
tt = torch.tensor(tt).to("cpu")
vals = torch.tensor(vals).to("cpu").to(dtype=torch.float32)
masks = torch.tensor(masks).to("cpu")
# -
vals
dataset_obj = []
dataset_obj.append((patient,tt,vals,masks))
dataset_obj
import torch
b = torch.arange(3 * 3*4 ).view(3, 3,4)
b
torch.sum(b, (0, 2))
29*15
# +
import numpy as np
import h5py
a = np.random.random(size=(100,20)) # or some such
b = np.random.random(size=(100,20)) # or some such
# -
data_to_write
np.stack((a,b)).shape
with h5py.File('name-of-file.h5', 'w') as hf:
hf.create_dataset("name-of-dataset", data=np.stack((a,b)))
with h5py.File('name-of-file.h5', 'r') as hf:
data = hf['name-of-dataset'][:]
data.shape
|
test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import librosa
import os
import tensorflow as tf
import numpy as np
from tqdm import tqdm
wav_files = [f for f in os.listdir('./data') if f.endswith('.wav')]
text_files = [f for f in os.listdir('./data') if f.endswith('.txt')]
inputs, targets = [], []
for (wav_file, text_file) in tqdm(zip(wav_files, text_files), total = len(wav_files),ncols=80):
path = './data/' + wav_file
try:
y, sr = librosa.load(path, sr = None)
except:
continue
inputs.append(
librosa.feature.mfcc(
y = y, sr = sr, n_mfcc = 40, hop_length = int(0.05 * sr)
).T
)
with open('./data/' + text_file) as f:
targets.append(f.read())
# +
inputs = tf.keras.preprocessing.sequence.pad_sequences(
inputs, dtype = 'float32', padding = 'post'
)
chars = list(set([c for target in targets for c in target]))
num_classes = len(chars) + 1
idx2char = {idx: char for idx, char in enumerate(chars)}
char2idx = {char: idx for idx, char in idx2char.items()}
targets = [[char2idx[c] for c in target] for target in targets]
# +
def pad_sentence_batch(sentence_batch, pad_int):
padded_seqs = []
seq_lens = []
max_sentence_len = max([len(sentence) for sentence in sentence_batch])
for sentence in sentence_batch:
padded_seqs.append(sentence + [pad_int] * (max_sentence_len - len(sentence)))
seq_lens.append(len(sentence))
return padded_seqs, seq_lens
def sparse_tuple_from(sequences, dtype=np.int32):
indices = []
values = []
for n, seq in enumerate(sequences):
indices.extend(zip([n] * len(seq), range(len(seq))))
values.extend(seq)
indices = np.asarray(indices, dtype=np.int64)
values = np.asarray(values, dtype=dtype)
shape = np.asarray([len(sequences), np.asarray(indices).max(0)[1] + 1], dtype=np.int64)
return indices, values, shape
# +
def attention(inputs, attention_size):
hidden_size = inputs.shape[2].value
w_omega = tf.Variable(
tf.random_normal([hidden_size, attention_size], stddev = 0.1)
)
b_omega = tf.Variable(tf.random_normal([attention_size], stddev = 0.1))
u_omega = tf.Variable(tf.random_normal([attention_size], stddev = 0.1))
with tf.name_scope('v'):
v = tf.tanh(tf.tensordot(inputs, w_omega, axes = 1) + b_omega)
vu = tf.tensordot(v, u_omega, axes = 1, name = 'vu')
alphas = tf.nn.softmax(vu, name = 'alphas')
output = inputs * tf.expand_dims(alphas, -1)
return output, alphas
class Model:
def __init__(
self,
num_layers,
size_layers,
learning_rate,
num_features,
dropout = 1.0,
):
self.X = tf.placeholder(tf.float32, [None, None, num_features])
self.Y = tf.sparse_placeholder(tf.int32)
seq_lens = tf.count_nonzero(
tf.reduce_sum(self.X, -1), 1, dtype = tf.int32
)
self.label = tf.placeholder(tf.int32, [None, None])
self.Y_seq_len = tf.placeholder(tf.int32, [None])
def cells(size, reuse = False):
return tf.contrib.rnn.DropoutWrapper(
tf.nn.rnn_cell.LSTMCell(
size,
initializer = tf.orthogonal_initializer(),
reuse = reuse,
),
state_keep_prob = dropout,
output_keep_prob = dropout,
)
features = self.X
for n in range(num_layers):
(out_fw, out_bw), (
state_fw,
state_bw,
) = tf.nn.bidirectional_dynamic_rnn(
cell_fw = cells(size_layers),
cell_bw = cells(size_layers),
inputs = features,
sequence_length = seq_lens,
dtype = tf.float32,
scope = 'bidirectional_rnn_%d' % (n),
)
features = tf.concat((out_fw, out_bw), 2)
features, _ = attention(features, size_layers)
logits = tf.layers.dense(features, num_classes)
time_major = tf.transpose(logits, [1, 0, 2])
decoded, log_prob = tf.nn.ctc_beam_search_decoder(time_major, seq_lens)
decoded = tf.to_int32(decoded[0])
self.preds = tf.sparse.to_dense(decoded)
self.cost = tf.reduce_mean(
tf.nn.ctc_loss(
self.Y,
time_major,
seq_lens,
ignore_longer_outputs_than_inputs = True,
)
)
self.optimizer = tf.train.AdamOptimizer(
learning_rate = learning_rate
).minimize(self.cost)
preds = self.preds[:, :tf.reduce_max(self.Y_seq_len)]
masks = tf.sequence_mask(self.Y_seq_len, tf.reduce_max(self.Y_seq_len), dtype=tf.float32)
preds = tf.pad(preds, [[0, 0], [0, tf.reduce_max(self.Y_seq_len)]])
y_t = tf.cast(preds, tf.int32)
self.prediction = tf.boolean_mask(y_t, masks)
mask_label = tf.boolean_mask(self.label, masks)
correct_pred = tf.equal(self.prediction, mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# +
tf.reset_default_graph()
sess = tf.InteractiveSession()
size_layers = 128
learning_rate = 1e-3
num_layers = 2
batch_size = 32
epoch = 50
model = Model(num_layers, size_layers, learning_rate, inputs.shape[2])
sess.run(tf.global_variables_initializer())
# -
for e in range(epoch):
pbar = tqdm(
range(0, len(inputs), batch_size), desc = 'minibatch loop')
for i in pbar:
batch_x = inputs[i : min(i + batch_size, len(inputs))]
y = targets[i : min(i + batch_size, len(inputs))]
batch_y = sparse_tuple_from(y)
batch_label, batch_len = pad_sentence_batch(y, 0)
_, cost, accuracy = sess.run(
[model.optimizer, model.cost, model.accuracy],
feed_dict = {model.X: batch_x, model.Y: batch_y,
model.label: batch_label, model.Y_seq_len: batch_len},
)
accuracy = sess.run(model.accuracy, feed_dict = {model.X: batch_x[: 1],
model.label: batch_label[: 1],
model.Y_seq_len: batch_len[: 1]})
pbar.set_postfix(cost = cost, accuracy = np.mean(accuracy))
# +
import random
random_index = random.randint(0, len(targets) - 1)
batch_x = inputs[random_index : random_index + 1]
print(
'real:',
''.join(
[idx2char[no] for no in targets[random_index : random_index + 1][0]]
),
)
batch_y = sparse_tuple_from(targets[random_index : random_index + 1])
pred = sess.run(model.preds, feed_dict = {model.X: batch_x})[0]
print('predicted:', ''.join([idx2char[no] for no in pred]))
# -
|
speech-to-text/6.birnn-attention-ctc-beam.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# name: python385jvsc74a57bd02db524e06e9f5f4ffedc911c917cb75e12dbc923643829bf417064a77eb14d37
# ---
# # TASK 1
# ### By <NAME>
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn import preprocessing
from sklearn import metrics
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
data = pd.read_csv("Student_scores_dataset.csv")
data.head(5)
data.info()
data.describe()
data.plot(x='Hours', y='Scores', style='o')
plt.title('Hours vs Percentage')
plt.xlabel('Hours Studied')
plt.ylabel('Percentage Score')
plt.show()
# *Data Preparation*
x = data.iloc[:,:1].values
y = data.iloc[:,1].values
# *Algorithm*
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.2, random_state = 60)
linear=LinearRegression()
linear.fit(x_train, y_train)
print("Training done")
c = float(linear.coef_[0])
d = float(linear.intercept_)
line = c*x + d
plt.plot(x,line)
plt.scatter(x,y)
plt.ylabel('Score')
plt.xlabel('Number of Hours')
plt.title('Regression Line')
plt.show()
# *Making Prediction*
print(x_test)
ypred = linear.predict(x_test)
dataframe = pd.DataFrame({'Actual': y_test, 'Predicted': ypred})
dataframe
hours = 9.25
own_pred = linear.predict(np.array([hours]).reshape(-1,1))
print("No of Hours = {}".format(hours))
print("Predicted Score = {}".format(own_pred[0]))
# ### Evaluation of the model
print('Mean Absolute Error:',metrics.mean_absolute_error(y_test, ypred))
print("Mean Squared Error:",metrics.mean_squared_error(y_test, ypred))
|
Task1/Linear_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import researchpy as rp
import seaborn as sns
import statsmodels.api as sm
from statsmodels.formula.api import ols
import statsmodels.stats.multicomp
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
df = pd.read_csv('upload_mse_harmonic_ANOVA.txt', header=0, sep="\t")
#df.drop('metric', axis=1, inplace=True)
df.head()
# unpivot data
#df = pd.melt(df.reset_index(), id_vars=['embeddings', 'model','clusters', 'metric'], value_vars=['fold_1','fold_2','fold_3','fold_4','fold_5'])
df = pd.melt(df.reset_index(), id_vars=['embeddings', 'model','clusters'], value_vars=['hm_fold1','hm_fold2','hm_fold3','hm_fold4','hm_fold5'])
# +
# drop metric and variable columns since they're not useful
#df.drop('metric',axis=1,inplace=True)
df.drop('variable',axis=1,inplace=True)
# rename value to actual metric (our dependent variable)
df.rename(columns={'value':'map_mrr'}, inplace=True)
# -
# show general statistics
rp.summary_cont(df['map_mrr'])
# filter out MFH, Cosine
#df = df.loc[df.model.isin(['Baseline', 'Asymmetric', 'Neighbour']),:]
df = df.loc[df.model.isin(['Baseline', 'MFH']),:]
rp.summary_cont(df.groupby(['model']))['map_mrr']
# separate the embeddings
w2v = df.loc[df['embeddings']=='w2v', :]
ft = df.loc[df['embeddings']=='ft', :]
glove = df.loc[df['embeddings']=='glove', :]
# +
def anova_table(aov):
aov['mean_sq'] = aov[:]['sum_sq']/aov[:]['df']
aov['eta_sq'] = aov[:-1]['sum_sq']/sum(aov['sum_sq'])
aov['omega_sq'] = (aov[:-1]['sum_sq']-(aov[:-1]['df']*aov['mean_sq'][-1]))/(sum(aov['sum_sq'])+aov['mean_sq'][-1])
cols = ['sum_sq', 'mean_sq', 'df', 'F', 'PR(>F)', 'eta_sq', 'omega_sq']
aov = aov[cols]
return aov
def print_model(mod):
print(f"Overall model F({mod.df_model: .0f},{mod.df_resid: .0f}) = {mod.fvalue: .3f}, p = {mod.f_pvalue: .10f}")
# -
w2v.shape
# # Evaluate w2v results
# +
# Fits the model with the interaction term
# This will also automatically include the main effects for each factor
w2v_model = ols('map_mrr ~ C(clusters)*C(model)', w2v).fit()
# Seeing if the overall model is significant
print(f"Overall model F({w2v_model.df_model: .0f},{w2v_model.df_resid: .0f}) = {w2v_model.fvalue: .3f}, p = {w2v_model.f_pvalue: .10f}")
# -
w2v_model.summary()
# +
# Creates the ANOVA table
res = sm.stats.anova_lm(w2v_model, typ= 1)
anova_table(res)
#We found that the effect of the interaction of cluster and model was insignificant. F(4, 36)=1.92, p > 0.05.
#We redo the analysis, this time testing for the effect of cluster and model in isolation.
# +
# we remove the interaction term and analyse the indepedent variables separately
w2v_model2 = ols('map_mrr ~ C(clusters)+ C(model)', w2v).fit()
print(f"Overall model F({w2v_model2.df_model: .0f},{w2v_model2.df_resid: .0f}) = {w2v_model2.fvalue: .3f}, p = {w2v_model2.f_pvalue: .8f}")
# -
w2v_model2.summary()
res2 = sm.stats.anova_lm(w2v_model2, typ= 1)
anova_table(res2)
# After refitting the ANOVA model on the combination of cluster and model, very weak statistical evidence
# was found that the effect of the regularised models is significant.
# However, the effect of clusters was significant on the harmonic mean of MAP and MRR, F(2,40)=44.342, p << 0.05
# omega^2 = 0.637
# +
# employ Tukey's HSD to find evidence of effect in model
mc = statsmodels.stats.multicomp.MultiComparison(w2v['map_mrr'], w2v['clusters'])
mc_results = mc.tukeyhsd()
print(mc_results)
# post-hoc analysis revealed that increasing the clusters from 1 to 10, 1 to 25 and 10 to 25 all had a positive effect
# on the MAP_MRR harmonic mean when training an MSE model on word2vec features.
# -
# # Evaluate GloVe
# +
glove_model = ols('map_mrr ~ C(clusters)*C(model)', glove).fit()
# Seeing if the overall model is significant
print(f"Overall model F({glove_model.df_model: .0f},{glove_model.df_resid: .0f}) = {glove_model.fvalue: .3f}, p = {glove_model.f_pvalue: .10f}")
# +
glove_model.summary()
# the overall interaction combination model was insignificant, F(8, 45) = 2.149, p < 0.05.
# +
# Creates the ANOVA table
res = sm.stats.anova_lm(glove_model, typ= 1)
anova_table(res)
# we tested the combination of clusters and model on the effect of mrr_map harmonic mean.
# the overall 2-way combination ANOVA was insignificant.
# +
glove_model2 = ols('map_mrr ~ C(clusters)', glove).fit()
print(f"Overall model F({glove_model2.df_model: .0f},{glove_model2.df_resid: .0f}) = {glove_model2.fvalue: .3f}, p = {glove_model2.f_pvalue: .8f}")
# we fitted another ANOVA model, this time exploring the effect of clusters along.
# This was significant, F(2, 42)=9.748, p < 0.05; omega^2 = 0.280
# -
res2 = sm.stats.anova_lm(glove_model2, typ= 1)
anova_table(res2)
# +
mc = statsmodels.stats.multicomp.MultiComparison(glove['map_mrr'], glove['clusters'])
mc_results = mc.tukeyhsd()
print(mc_results)
# Post-hoc analysis revealed that increasing clusters from 1 to 10 had a significantly negative effect
# on the map_mrr harmonic mean, while increasing clusters from 10 to 25 had a positive effect.
# However, increasing the clusters from 1 to 25 had no effect on the results.
# -
# # Evaluate fastText results
# +
ft_model = ols('map_mrr ~ C(clusters)*C(model)', ft).fit()
# Seeing if the overall model is significant
print(f"Overall model F({ft_model.df_model: .0f},{ft_model.df_resid: .0f}) = {ft_model.fvalue: .3f}, p = {ft_model.f_pvalue: .10f}")
# +
ft_model.summary()
# the interaction combination overall model was significant on the map_nrr, F(8, 36) = 3.556, p<0.05.
# +
# Creates the ANOVA table
res = sm.stats.anova_lm(ft_model, typ= 1)
anova_table(res)
# Inspecting the ANOVA table, however revealed that neither the interaction term nor the model term
# significant. We retested a fresh ANOVA, focusing on just the cluster term.
# +
ft_model2 = ols('map_mrr ~ C(clusters)', ft).fit()
print(f"Overall model F({ft_model2.df_model: .0f},{ft_model2.df_resid: .0f}) = {ft_model2.fvalue: .3f}, p = {ft_model2.f_pvalue: .8f}")
res2 = sm.stats.anova_lm(ft_model2, typ= 1)
print (res2)
anova_table(res2)
# There was a significant effect of clusters on the map_mrr score, F(2, 42) = 16.016, p << 0.05., omega^2=0.40
# +
mc = statsmodels.stats.multicomp.MultiComparison(ft['map_mrr'], ft['clusters'])
mc_results = mc.tukeyhsd()
print(mc_results)
# post-hoc analysis reveals that that increasing clusters from 1 to 10, and from 1 to 25
# has a negative effect on MAP_MRR while increasing clusters from 10 to 25 has a positive negligible effect.
# -
# # Evaluate difference in embeddings using only baseline
all_embeddings = df.loc[df.model=='Baseline',:]
all_embeddings.head()
#all_embeddings.loc[, ['embeddings','clusters']]
# +
all_model = ols('map_mrr ~ C(embeddings)*C(clusters)', all_embeddings).fit()
# Seeing if the overall model is significant
print(f"Overall model F({all_model.df_model: .0f},{all_model.df_resid: .0f}) = {all_model.fvalue: .3f}, p = {all_model.f_pvalue: .10f}")
# +
all_model.summary()
# Once we could safely ignore the effect of the regularised models, we investigated the interaction of
# embeddings and cluster.s
# The overal ANOVA was significant F(8. 36) = 19.733, p << 0.05
# -
# Creates the ANOVA table
res = sm.stats.anova_lm(all_model, typ= 1)
anova_table(res)
sns.catplot(x="embeddings", y="map_mrr", hue="clusters", kind="point", data=all_embeddings);
# +
# interaction effect is significant. We decided to Rerun ANOVA investigating simple effect. That is we
# run three models, checking the influence of the embeddings whilst controlling for 1, 10 and 25 clusters
# respectively
# embeddings | c = 1
# embeddings | c = 10
# embeddings | c = 25
# +
cl_1 = all_embeddings.loc[all_embeddings.clusters == 1,]
cl_1_model = ols('map_mrr ~ C(embeddings)', cl_1).fit()
# Seeing if the overall model is significant
print(f"Overall model F({cl_1_model.df_model: .0f},{cl_1_model.df_resid: .0f}) = {cl_1_model.fvalue: .3f}, p = {cl_1_model.f_pvalue: .10f}")
cl_1_model.summary()
# +
res = sm.stats.anova_lm(cl_1_model, typ= 1)
anova_table(res)
# the embeddings had a significant effect on single cluster, baseline models, F(2, 12)=52.376, p < 0.05, omega^2 = 0.876
# +
mc = statsmodels.stats.multicomp.MultiComparison(cl_1['map_mrr'], cl_1['embeddings'])
mc_results = mc.tukeyhsd()
print(mc_results)
# post-hoc analysis reveals that fastText yields a higher map_mrr than either GloVe or word2vec and that GloVE
# returns a higher score than word2vec.
# +
cl_10 = all_embeddings.loc[all_embeddings.clusters == 10,]
cl_10_model = ols('map_mrr ~ C(embeddings)', cl_10).fit()
# Seeing if the overall model is significant
print(f"Overall model F({cl_10_model.df_model: .0f},{cl_10_model.df_resid: .0f}) = {cl_10_model.fvalue: .3f}, p = {cl_10_model.f_pvalue: .10f}")
cl_10_model.summary()
# +
res = sm.stats.anova_lm(cl_10_model, typ= 1)
anova_table(res)
# the Embeddings were also significant with respect to a 10-cluster model F(2, 12)=9.471, p < 0.05, omega^2 = 0.530
# +
mc = statsmodels.stats.multicomp.MultiComparison(cl_10['map_mrr'], cl_10['embeddings'])
mc_results = mc.tukeyhsd()
print(mc_results)
# Once again fastText-trained models yield a significantly higher map_mrr than either GloVe or word2vec
# but in this context word2vec vectors has only a marginally stronger effect on map_mrr than GloVe.
# +
cl_25 = all_embeddings.loc[all_embeddings.clusters == 25,]
cl_25_model = ols('map_mrr ~ C(embeddings)', cl_25).fit()
# Seeing if the overall model is significant
print(f"Overall model F({cl_25_model.df_model: .0f},{cl_25_model.df_resid: .0f}) = {cl_25_model.fvalue: .3f}, p = {cl_25_model.f_pvalue: .10f}")
cl_25_model.summary()
# +
res = sm.stats.anova_lm(cl_25_model, typ= 1)
anova_table(res)
# Lastly, embeddings have a medium effect on 25-cluster models too, F(2,12)=11.406, p<0.05, omega^2=0.58
# +
mc = statsmodels.stats.multicomp.MultiComparison(cl_25['map_mrr'], cl_25['embeddings'])
mc_results = mc.tukeyhsd()
print(mc_results)
# we already know from a previous test that the fastText model's performance is inhibited by training more cluster
# Indeed, we only detected that fastText has a bigger influence on GloVe but there was no detectable difference
# between fastText and word2vec and GloVe and word2vec.
# There is strong empirical evidence which suggests that - on our chosen combined dataset and metrics -
# training a single-cluster, baseline model on fastText is the way to go. This challenges
# Ustalov et al.'s assertion that regularisation has an effect on MSE model performance. We were not able to
# confirm that even on the same word2vec embeddings they used in their experiment. However, we do not run
# exhaustive grid-search tests to tune the lambda regularisation parameter, instead setting to 1., which is
# their chosen regularisation weight for re-projected regularisation.
# Fu et. al's finding were re-confirmed, despite us chaning the setup in several ways: we recast their original
# problem as hypernym discovery; evaluated on information retrieval metrics; cross-validated the models on a
# different dataset. Cluster size was certainly found to improve performance on word2vec embeddings,
# although we didn't test larger cluster sizes.
# -
# # Analyse performance of MSE against baseline
# +
base_w2v = df.loc[(df.embeddings=='w2v'),]#&(df.clusters==1),]
w2v_naive_model = ols('map_mrr ~ C(model)*C(clusters)', base_w2v).fit()
print_model(w2v_naive_model)
# -
w2v_naive_model.summary()
res = sm.stats.anova_lm(w2v_naive_model, typ= 1)
anova_table(res)
mc = statsmodels.stats.multicomp.MultiComparison(base_w2v['map_mrr'], base_w2v['clusters'])
mc_results = mc.tukeyhsd()
print(mc_results)
|
ANOVA Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Xmf_JRJa_N8C" colab_type="text"
# <table align="center">
# <td align="center"><a target="_blank" href="http://introtodeeplearning.com">
# <img src="http://introtodeeplearning.com/images/colab/mit.png" style="padding-bottom:5px;" />
# Visit MIT Deep Learning</a></td>
# <td align="center"><a target="_blank" href="https://colab.research.google.com/github/aamini/introtodeeplearning/blob/master/lab2/Part1_MNIST.ipynb">
# <img src="http://introtodeeplearning.com/images/colab/colab.png?v2.0" style="padding-bottom:5px;" />Run in Google Colab</a></td>
# <td align="center"><a target="_blank" href="https://github.com/aamini/introtodeeplearning/blob/master/lab2/Part1_MNIST.ipynb">
# <img src="http://introtodeeplearning.com/images/colab/github.png" height="70px" style="padding-bottom:5px;" />View Source on GitHub</a></td>
# </table>
#
# # Copyright Information
# + id="gKA_J7bdP33T" colab_type="code" colab={}
# Copyright 2020 MIT 6.S191 Introduction to Deep Learning. All Rights Reserved.
#
# Licensed under the MIT License. You may not use this file except in compliance
# with the License. Use and/or modification of this code outside of 6.S191 must
# reference:
#
# © MIT 6.S191: Introduction to Deep Learning
# http://introtodeeplearning.com
#
# + [markdown] id="Cm1XpLftPi4A" colab_type="text"
# # Laboratory 2: Computer Vision
#
# # Part 1: MNIST Digit Classification
#
# In the first portion of this lab, we will build and train a convolutional neural network (CNN) for classification of handwritten digits from the famous [MNIST](http://yann.lecun.com/exdb/mnist/) dataset. The MNIST dataset consists of 60,000 training images and 10,000 test images. Our classes are the digits 0-9.
#
# First, let's download the course repository, install dependencies, and import the relevant packages we'll need for this lab.
# + id="RsGqx_ai_N8F" colab_type="code" colab={}
# Import Tensorflow 2.0
# %tensorflow_version 2.x
import tensorflow as tf
# !pip install mitdeeplearning
import mitdeeplearning as mdl
import matplotlib.pyplot as plt
import numpy as np
import random
from tqdm import tqdm
# Check that we are using a GPU, if not switch runtimes
# using Runtime > Change Runtime Type > GPU
print(tf.config.list_physical_devices)
assert len(tf.config.list_physical_devices('GPU')) > 0
# + [markdown] id="HKjrdUtX_N8J" colab_type="text"
# ## 1.1 MNIST dataset
#
# Let's download and load the dataset and display a few random samples from it:
# + id="p2dQsHI3_N8K" colab_type="code" colab={}
mnist = tf.keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
train_images = (np.expand_dims(train_images, axis=-1)/255.).astype(np.float32)
train_labels = (train_labels).astype(np.int64)
test_images = (np.expand_dims(test_images, axis=-1)/255.).astype(np.float32)
test_labels = (test_labels).astype(np.int64)
# + [markdown] id="5ZtUqOqePsRD" colab_type="text"
# Our training set is made up of 28x28 grayscale images of handwritten digits.
#
# Let's visualize what some of these images and their corresponding training labels look like.
# + id="bDBsR2lP_N8O" colab_type="code" colab={}
plt.figure(figsize=(10,10))
random_inds = np.random.choice(60000,36)
for i in range(36):
plt.subplot(6,6,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
image_ind = random_inds[i]
plt.imshow(np.squeeze(train_images[image_ind]), cmap=plt.cm.binary)
plt.xlabel(train_labels[image_ind])
# + [markdown] id="V6hd3Nt1_N8q" colab_type="text"
# ## 1.2 Neural Network for Handwritten Digit Classification
#
# We'll first build a simple neural network consisting of two fully connected layers and apply this to the digit classification task. Our network will ultimately output a probability distribution over the 10 digit classes (0-9). This first architecture we will be building is depicted below:
#
# 
#
# + [markdown] id="rphS2rMIymyZ" colab_type="text"
# ### Fully connected neural network architecture
# To define the architecture of this first fully connected neural network, we'll once again use the Keras API and define the model using the [`Sequential`](https://www.tensorflow.org/api_docs/python/tf/keras/models/Sequential) class. Note how we first use a [`Flatten`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Flatten) layer, which flattens the input so that it can be fed into the model.
#
# In this next block, you'll define the fully connected layers of this simple work.
# + id="MMZsbjAkDKpU" colab_type="code" colab={}
def build_fc_model():
fc_model = tf.keras.Sequential([
# First define a Flatten layer
tf.keras.layers.Flatten(),
# '''TODO: Define the activation function for the first fully connected (Dense) layer.'''
tf.keras.layers.Dense(128, activation= 'relu'),
# '''TODO: Define the second Dense layer to output the classification probabilities'''
tf.keras.layers.Dense(10, activation='softmax')
])
return fc_model
model = build_fc_model()
# + [markdown] id="VtGZpHVKz5Jt" colab_type="text"
# As we progress through this next portion, you may find that you'll want to make changes to the architecture defined above. **Note that in order to update the model later on, you'll need to re-run the above cell to re-initialize the model. **
# + [markdown] id="mVN1_AeG_N9N" colab_type="text"
# Let's take a step back and think about the network we've just created. The first layer in this network, `tf.keras.layers.Flatten`, transforms the format of the images from a 2d-array (28 x 28 pixels), to a 1d-array of 28 * 28 = 784 pixels. You can think of this layer as unstacking rows of pixels in the image and lining them up. There are no learned parameters in this layer; it only reformats the data.
#
# After the pixels are flattened, the network consists of a sequence of two `tf.keras.layers.Dense` layers. These are fully-connected neural layers. The first `Dense` layer has 128 nodes (or neurons). The second (and last) layer (which you've defined!) should return an array of probability scores that sum to 1. Each node contains a score that indicates the probability that the current image belongs to one of the handwritten digit classes.
#
# That defines our fully connected model!
# + [markdown] id="gut8A_7rCaW6" colab_type="text"
#
#
# ### Compile the model
#
# Before training the model, we need to define a few more settings. These are added during the model's [`compile`](https://www.tensorflow.org/api_docs/python/tf/keras/models/Sequential#compile) step:
#
# * *Loss function* — This defines how we measure how accurate the model is during training. As was covered in lecture, during training we want to minimize this function, which will "steer" the model in the right direction.
# * *Optimizer* — This defines how the model is updated based on the data it sees and its loss function.
# * *Metrics* — Here we can define metrics used to monitor the training and testing steps. In this example, we'll look at the *accuracy*, the fraction of the images that are correctly classified.
#
# We'll start out by using a stochastic gradient descent (SGD) optimizer initialized with a learning rate of 0.1. Since we are performing a categorical classification task, we'll want to use the [cross entropy loss](https://www.tensorflow.org/api_docs/python/tf/keras/metrics/sparse_categorical_crossentropy).
#
# You'll want to experiment with both the choice of optimizer and learning rate and evaluate how these affect the accuracy of the trained model.
# + id="Lhan11blCaW7" colab_type="code" colab={}
'''TODO: Experiment with different optimizers and learning rates. How do these affect
the accuracy of the trained model? Which optimizers and/or learning rates yield
the best performance?'''
model.compile(optimizer=tf.keras.optimizers.SGD(learning_rate=1e-1),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# + [markdown] id="qKF6uW-BCaW-" colab_type="text"
# ### Train the model
#
# We're now ready to train our model, which will involve feeding the training data (`train_images` and `train_labels`) into the model, and then asking it to learn the associations between images and labels. We'll also need to define the batch size and the number of epochs, or iterations over the MNIST dataset, to use during training.
#
# In Lab 1, we saw how we can use `GradientTape` to optimize losses and train models with stochastic gradient descent. After defining the model settings in the `compile` step, we can also accomplish training by calling the [`fit`](https://www.tensorflow.org/api_docs/python/tf/keras/models/Sequential#fit) method on an instance of the `Model` class. We will use this to train our fully connected model
#
# + id="EFMbIqIvQ2X0" colab_type="code" colab={}
# Define the batch size and the number of epochs to use during training
BATCH_SIZE = 64
EPOCHS = 5
model.fit(train_images, train_labels, batch_size=BATCH_SIZE, epochs=EPOCHS)
# + [markdown] id="W3ZVOhugCaXA" colab_type="text"
# As the model trains, the loss and accuracy metrics are displayed. With five epochs and a learning rate of 0.01, this fully connected model should achieve an accuracy of approximatley 0.97 (or 97%) on the training data.
# + [markdown] id="oEw4bZgGCaXB" colab_type="text"
# ### Evaluate accuracy on the test dataset
#
# Now that we've trained the model, we can ask it to make predictions about a test set that it hasn't seen before. In this example, the `test_images` array comprises our test dataset. To evaluate accuracy, we can check to see if the model's predictions match the labels from the `test_labels` array.
#
# Use the [`evaluate`](https://www.tensorflow.org/api_docs/python/tf/keras/models/Sequential#evaluate) method to evaluate the model on the test dataset!
# + id="VflXLEeECaXC" colab_type="code" colab={}
'''TODO: Use the evaluate method to test the model!'''
test_loss, test_acc = model.evaluate(test_images, test_labels)
print('Test accuracy:', test_acc)
# + [markdown] id="yWfgsmVXCaXG" colab_type="text"
# You may observe that the accuracy on the test dataset is a little lower than the accuracy on the training dataset. This gap between training accuracy and test accuracy is an example of *overfitting*, when a machine learning model performs worse on new data than on its training data.
#
# What is the highest accuracy you can achieve with this first fully connected model? Since the handwritten digit classification task is pretty straightforward, you may be wondering how we can do better...
#
# 
# + [markdown] id="baIw9bDf8v6Z" colab_type="text"
# ## 1.3 Convolutional Neural Network (CNN) for handwritten digit classification
# + [markdown] id="_J72Yt1o_fY7" colab_type="text"
# As we saw in lecture, convolutional neural networks (CNNs) are particularly well-suited for a variety of tasks in computer vision, and have achieved near-perfect accuracies on the MNIST dataset. We will now build a CNN composed of two convolutional layers and pooling layers, followed by two fully connected layers, and ultimately output a probability distribution over the 10 digit classes (0-9). The CNN we will be building is depicted below:
#
# 
# + [markdown] id="EEHqzbJJAEoR" colab_type="text"
# ### Define the CNN model
#
# We'll use the same training and test datasets as before, and proceed similarly as our fully connected network to define and train our new CNN model. To do this we will explore two layers we have not encountered before: you can use [`keras.layers.Conv2D` ](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Conv2D) to define convolutional layers and [`keras.layers.MaxPool2D`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/MaxPool2D) to define the pooling layers. Use the parameters shown in the network architecture above to define these layers and build the CNN model.
# + id="vec9qcJs-9W5" colab_type="code" colab={}
def build_cnn_model():
cnn_model = tf.keras.Sequential([
# TODO: Define the first convolutional layer
tf.keras.layers.Conv2D(24, kernel_size=3, activation='relu'),
# TODO: Define the first max pooling layer
tf.keras.layers.MaxPool2D(pool_size=(3,3), strides=2),
# TODO: Define the second convolutional layer
tf.keras.layers.Conv2D(36, kernel_size=3, activation='relu'),
# TODO: Define the second max pooling layer
tf.keras.layers.MaxPool2D(pool_size=(2,2), strides=2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
# TODO: Define the last Dense layer to output the classification
# probabilities. Pay attention to the activation needed a probability
# output
tf.keras.layers.Dense(10, activation='softmax')
])
return cnn_model
cnn_model = build_cnn_model()
# Initialize the model by passing some data through
cnn_model.predict(train_images[[0]])
# Print the summary of the layers in the model.
print(cnn_model.summary())
# + [markdown] id="kUAXIBynCih2" colab_type="text"
# ### Train and test the CNN model
#
# Now, as before, we can define the loss function, optimizer, and metrics through the `compile` method. Compile the CNN model with an optimizer and learning rate of choice:
# + id="vheyanDkCg6a" colab_type="code" colab={}
'''TODO: Define the compile operation with your optimizer and learning rate of choice'''
cnn_model.compile(optimizer=tf.keras.optimizers.SGD(learning_rate=0.01), loss='sparse_categorical_crossentropy', metrics=['accuracy']) # TODO
# + [markdown] id="U19bpRddC7H_" colab_type="text"
# As was the case with the fully connected model, we can train our CNN using the `fit` method via the Keras API.
# + id="YdrGZVmWDK4p" colab_type="code" colab={}
'''TODO: Use model.fit to train the CNN model, with the same batch_size and number of epochs previously used.'''
cnn_model.fit(train_images, train_labels, BATCH_SIZE, EPOCHS)
# + [markdown] id="pEszYWzgDeIc" colab_type="text"
# Great! Now that we've trained the model, let's evaluate it on the test dataset using the [`evaluate`](https://www.tensorflow.org/api_docs/python/tf/keras/models/Sequential#evaluate) method:
# + id="JDm4znZcDtNl" colab_type="code" colab={}
'''TODO: Use the evaluate method to test the model!'''
test_loss, test_acc = cnn_model.evaluate(test_images, test_labels)
print('Test accuracy:', test_acc)
# + [markdown] id="2rvEgK82Glv9" colab_type="text"
# What is the highest accuracy you're able to achieve using the CNN model, and how does the accuracy of the CNN model compare to the accuracy of the simple fully connected network? What optimizers and learning rates seem to be optimal for training the CNN model?
# + [markdown] id="xsoS7CPDCaXH" colab_type="text"
# ### Make predictions with the CNN model
#
# With the model trained, we can use it to make predictions about some images. The [`predict`](https://www.tensorflow.org/api_docs/python/tf/keras/models/Sequential#predict) function call generates the output predictions given a set of input samples.
#
# + id="Gl91RPhdCaXI" colab_type="code" colab={}
predictions = cnn_model.predict(test_images)
# + [markdown] id="x9Kk1voUCaXJ" colab_type="text"
# With this function call, the model has predicted the label for each image in the testing set. Let's take a look at the prediction for the first image in the test dataset:
# + id="3DmJEUinCaXK" colab_type="code" colab={}
predictions[0]
# + [markdown] id="-hw1hgeSCaXN" colab_type="text"
# As you can see, a prediction is an array of 10 numbers. Recall that the output of our model is a probability distribution over the 10 digit classes. Thus, these numbers describe the model's "confidence" that the image corresponds to each of the 10 different digits.
#
# Let's look at the digit that has the highest confidence for the first image in the test dataset:
# + id="qsqenuPnCaXO" colab_type="code" colab={}
'''TODO: identify the digit with the highest confidence prediction for the first
image in the test dataset. '''
prediction = np.argmax(predictions[0])
print(prediction)
# + [markdown] id="E51yS7iCCaXO" colab_type="text"
# So, the model is most confident that this image is a "???". We can check the test label (remember, this is the true identity of the digit) to see if this prediction is correct:
# + id="Sd7Pgsu6CaXP" colab_type="code" colab={}
print("Label of this digit is:", test_labels[0])
plt.imshow(test_images[0,:,:,0], cmap=plt.cm.binary)
# + [markdown] id="ygh2yYC972ne" colab_type="text"
# It is! Let's visualize the classification results on the MNIST dataset. We will plot images from the test dataset along with their predicted label, as well as a histogram that provides the prediction probabilities for each of the digits:
# + id="HV5jw-5HwSmO" colab_type="code" cellView="both" colab={}
#@title Change the slider to look at the model's predictions! { run: "auto" }
image_index = 50 #@param {type:"slider", min:0, max:100, step:1}
plt.subplot(1,2,1)
mdl.lab2.plot_image_prediction(image_index, predictions, test_labels, test_images)
plt.subplot(1,2,2)
mdl.lab2.plot_value_prediction(image_index, predictions, test_labels)
# + [markdown] id="kgdvGD52CaXR" colab_type="text"
# We can also plot several images along with their predictions, where correct prediction labels are blue and incorrect prediction labels are red. The number gives the percent confidence (out of 100) for the predicted label. Note the model can be very confident in an incorrect prediction!
# + id="hQlnbqaw2Qu_" colab_type="code" colab={}
# Plots the first X test images, their predicted label, and the true label
# Color correct predictions in blue, incorrect predictions in red
num_rows = 5
num_cols = 4
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
mdl.lab2.plot_image_prediction(i, predictions, test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
mdl.lab2.plot_value_prediction(i, predictions, test_labels)
# + [markdown] id="k-2glsRiMdqa" colab_type="text"
# ## 1.4 Training the model 2.0
#
# Earlier in the lab, we used the [`fit`](https://www.tensorflow.org/api_docs/python/tf/keras/models/Sequential#fit) function call to train the model. This function is quite high-level and intuitive, which is really useful for simpler models. As you may be able to tell, this function abstracts away many details in the training call, and we have less control over training model, which could be useful in other contexts.
#
# As an alternative to this, we can use the [`tf.GradientTape`](https://www.tensorflow.org/api_docs/python/tf/GradientTape) class to record differentiation operations during training, and then call the [`tf.GradientTape.gradient`](https://www.tensorflow.org/api_docs/python/tf/GradientTape#gradient) function to actually compute the gradients. You may recall seeing this in Lab 1 Part 1, but let's take another look at this here.
#
# We'll use this framework to train our `cnn_model` using stochastic gradient descent.
# + id="Wq34id-iN1Ml" colab_type="code" colab={}
# Rebuild the CNN model
cnn_model = build_cnn_model()
batch_size = 12
loss_history = mdl.util.LossHistory(smoothing_factor=0.95) # to record the evolution of the loss
plotter = mdl.util.PeriodicPlotter(sec=2, xlabel='Iterations', ylabel='Loss', scale='semilogy')
optimizer = tf.keras.optimizers.SGD(learning_rate=1e-2) # define our optimizer
if hasattr(tqdm, '_instances'): tqdm._instances.clear() # clear if it exists
for idx in tqdm(range(0, train_images.shape[0], batch_size)):
# First grab a batch of training data and convert the input images to tensors
(images, labels) = (train_images[idx:idx+batch_size], train_labels[idx:idx+batch_size])
images = tf.convert_to_tensor(images, dtype=tf.float32)
# GradientTape to record differentiation operations
with tf.GradientTape() as tape:
#'''TODO: feed the images into the model and obtain the predictions'''
logits = cnn_model(images)
#'''TODO: compute the categorical cross entropy loss
loss_value = tf.keras.backend.sparse_categorical_crossentropy(labels, logits) # TODO
loss_history.append(loss_value.numpy().mean()) # append the loss to the loss_history record
plotter.plot(loss_history.get())
# Backpropagation
'''TODO: Use the tape to compute the gradient against all parameters in the CNN model.
Use cnn_model.trainable_variables to access these parameters.'''
grads = tape.gradient(loss_value, cnn_model.trainable_variables)
optimizer.apply_gradients(zip(grads, cnn_model.trainable_variables))
# + [markdown] id="3cNtDhVaqEdR" colab_type="text"
# ## 1.5 Conclusion
# In this part of the lab, you had the chance to play with different MNIST classifiers with different architectures (fully-connected layers only, CNN), and experiment with how different hyperparameters affect accuracy (learning rate, etc.). The next part of the lab explores another application of CNNs, facial detection, and some drawbacks of AI systems in real world applications, like issues of bias.
|
lab2/Part1_MNIST.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import gym
import matplotlib.pyplot as plt
# %matplotlib inline
# All gym installed enviroments
gym.envs.registry.all()
# Lets start with simple one
env = gym.make("CartPole-v0")
# env start with reset function which returns initial observation
obs = env.reset()
print(obs)
# 
# number of posible actions
env.action_space.n
# number of observation that returns
env.observation_space
# get random action from posible actions
env.action_space.sample()
env.reset()
for _ in range(1000):
# display the game
env.render()
# take a random action
observation, reward, done, info = env.step(env.action_space.sample())
# Stop if the game is finished
if done:
break
# Atari enviroments
env = gym.make("Breakout-v0")
# this time observation is screen capture (pixel values)
obs = env.reset()
plt.imshow(obs)
plt.show()
obs.shape
# Posible actions
print(env.action_space.n)
print(env.env.get_action_meanings())
# Preprocessing the observation
# Crop the image
obs = obs[40:200,10:150]
# Downsample
obs = obs[::2,::2]
# rgb2gray
obs = obs.sum(axis=2)
# make it binary
obs[obs != 0] =1
obs.shape
plt.imshow(obs, cmap='gray')
plt.show()
|
openai_gym/Understanding Gym enviroments.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import Modules
import ipywidgets as widgets
from IPython.display import display
import numpy as np
import matplotlib.pyplot as plt
import os
from scipy.optimize import curve_fit
# %matplotlib widget
from datetime import datetime
import definitions as defs
# # Define definitions for widgets
# +
#Reset button
def reset_values(b):
"""Reset the interactive plots to inital values."""
for i, p in enumerate(start_parameter):
interactive_plot.children[i].value = p
#display of the initial guess
def print_out(center, sigma, scale):
print(center,sigma,scale)
# Fit button
def run_fit(button):
global params, pcov
if numbers.value == '1':
params, pcov = curve_fit(defs.log_norm_func1, d[index_left:index_right+1], y_values[index_left:index_right+1], p0=parameter_set, sigma = y_error[index_left:index_right+1], method ='trf')
elif numbers.value == '2':
params, pcov = curve_fit(defs.log_norm_func2, d[index_left:index_right+1], y_values[index_left:index_right+1], p0=parameter_set, sigma = y_error[index_left:index_right+1], method ='trf')
elif numbers.value == '3':
params, pcov = curve_fit(defs.log_norm_func3, d[index_left:index_right+1], y_values[index_left:index_right+1], p0=parameter_set, sigma = y_error[index_left:index_right+1], method ='trf')
elif numbers.value == '4':
params, pcov = curve_fit(defs.log_norm_func4, d[index_left:index_right+1], y_values[index_left:index_right+1], p0=parameter_set, sigma = y_error[index_left:index_right+1], method ='trf')
elif numbers.value == '5':
params, pcov = curve_fit(defs.log_norm_func5, d[index_left:index_right+1], y_values[index_left:index_right+1], p0=parameter_set, sigma = y_error[index_left:index_right+1], method ='trf')
with fit_results:
if x50 == []:
for i in range(int(numbers.value)):
x50.append(np.exp(params[0+3*i]))
sigma_s.append(np.exp(params[1+3*i]))
if numbers.value == '1':
print('Median1/nm: %.1f, Sigma: %.2f' %(x50[0], sigma_s[0]))
textstr = '\n'.join((
'$d_{50}/nm$\t $\sigma_g$',
'$%.1f$\t \t $%.2f$' %(x50[0], sigma_s[0])))
fit_lines[0].set_ydata(defs.log_norm_func1(x_fit, *params))
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[0:3]), facecolor="red", alpha=0.5)
elif numbers.value == '2':
print('Median1/nm: %.1f, Sigma: %.2f' %(x50[0], sigma_s[0]))
print('Median2/nm: %.1f, Sigma: %.2f' %(x50[1], sigma_s[1]))
textstr = '\n'.join((
'$d_{50}/nm$\t $\sigma_g$',
'$%.1f$\t \t $%.2f$' %(x50[0], sigma_s[0]),
'$%.1f$\t \t $%.2f$' %(x50[1], sigma_s[1])))
fit_lines[0].set_ydata(defs.log_norm_func2(x_fit, *params))
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[0:3]), facecolor="red", alpha=0.5)
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[3:6]), facecolor="yellow", alpha=0.5)
elif numbers.value == '3':
print('Median1/nm: %.1f, Sigma: %.2f' %(x50[0], sigma_s[0]))
print('Median2/nm: %.1f, Sigma: %.2f' %(x50[1], sigma_s[1]))
print('Median3/nm: %.1f, Sigma: %.2f' %(x50[2], sigma_s[2]))
textstr = '\n'.join((
'$d_{50}/nm$\t $\sigma_g$',
'$%.1f$\t \t $%.2f$' %(x50[0], sigma_s[0]),
'$%.1f$\t \t $%.2f$' %(x50[1], sigma_s[1]),
'$%.1f$\t \t $%.2f$' %(x50[2], sigma_s[2])))
fit_lines[0].set_ydata(defs.log_norm_func3(x_fit, *params))
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[0:3]), facecolor="red", alpha=0.5)
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[3:6]), facecolor="yellow", alpha=0.5)
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[6:9]), facecolor="cyan", alpha=0.5)
elif numbers.value == '4':
print('Median1/nm: %.1f, Sigma: %.2f' %(x50[0], sigma_s[0]))
print('Median2/nm: %.1f, Sigma: %.2f' %(x50[1], sigma_s[1]))
print('Median3/nm: %.1f, Sigma: %.2f' %(x50[2], sigma_s[2]))
print('Median4/nm: %.1f, Sigma: %.2f' %(x50[3], sigma_s[3]))
textstr = '\n'.join((
'$d_{50}/nm$\t $\sigma_g$',
'$%.1f$\t \t $%.2f$' %(x50[0], sigma_s[0]),
'$%.1f$\t \t $%.2f$' %(x50[1], sigma_s[1]),
'$%.1f$\t \t $%.2f$' %(x50[2], sigma_s[2]),
'$%.1f$\t \t $%.2f$' %(x50[3], sigma_s[3])))
fit_lines[0].set_ydata(defs.log_norm_func4(x_fit, *params))
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[0:3]), facecolor="red", alpha=0.5)
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[3:6]), facecolor="yellow", alpha=0.5)
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[6:9]), facecolor="cyan", alpha=0.5)
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[9:]), facecolor="magenta", alpha=0.5)
elif numbers.value == '5':
print('Median1/nm: %.1f, Sigma: %.2f' %(x50[0], sigma_s[0]))
print('Median2/nm: %.1f, Sigma: %.2f' %(x50[1], sigma_s[1]))
print('Median3/nm: %.1f, Sigma: %.2f' %(x50[2], sigma_s[2]))
print('Median4/nm: %.1f, Sigma: %.2f' %(x50[3], sigma_s[3]))
print('Median5/nm: %.1f, Sigma: %.2f' %(x50[4], sigma_s[4]))
textstr = '\n'.join((
'$d_{50}/nm$\t $\sigma_g$',
'$%.1f$\t \t $%.2f$' %(x50[0], sigma_s[0]),
'$%.1f$\t \t $%.2f$' %(x50[1], sigma_s[1]),
'$%.1f$\t \t $%.2f$' %(x50[2], sigma_s[2]),
'$%.1f$\t \t $%.2f$' %(x50[3], sigma_s[3]),
'$%.1f$\t \t $%.2f$' %(x50[4], sigma_s[4])))
fit_lines[0].set_ydata(defs.log_norm_func5(x_fit, *params))
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[0:3]), facecolor="red", alpha=0.5)
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[3:6]), facecolor="yellow", alpha=0.5)
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[6:9]), facecolor="cyan", alpha=0.5)
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[9:12]), facecolor="magenta", alpha=0.5)
ax2.fill_between(x_fit,defs.log_norm_func1(x_fit,*params[12:]), facecolor="chocolate", alpha=0.5)
ax2.text(0.755, 0.82, textstr, transform=ax2.transAxes, fontsize=10,verticalalignment='top', bbox={'facecolor': 'white', 'boxstyle':'round','pad':0.3,'edgecolor':'black', 'lw':0.8})
else:
fit_results.clear_output()
print('You fitted the data! Restart block to fit again!')
# Save button
def save_data(button):
if saved.outputs == ():
if not os.path.exists(os.path.join('results/')):
os.makedirs(os.path.join('results/'))
plt.savefig(os.path.join('results/', file_name +'_'+ date +'_fit_run'+str(select.value)+'.png'))
with saved:
print('Data saved!')
if not os.path.exists(os.path.join('results/', file_name + '_'+ date +'_results.txt')):
with open(os.path.join('results/', file_name + '_'+ date +'_results.txt'), 'w+') as exports:
if numbers.value == '1':
exports.write('Run' + '\t' + 'd50_1' + '\t' + 'Sigma1' + '\n')
exports.write(str(select.value) + '\t' + str(np.round(x50[0],2)) + '\t' + str(np.round(sigma_s[0],2)) + '\n' )
if numbers.value == '2':
exports.write('Run' + '\t' + 'd50_1' + '\t' + 'Sigma1' + '\t' + 'd50_2' + '\t' + 'Sigma2' + '\n')
exports.write(str(select.value) + '\t' + str(np.round(x50[0],2)) + '\t' + str(np.round(sigma_s[0],2)) + '\t' + str(np.round(x50[1],2)) + '\t' + str(np.round(sigma_s[1],2)) + '\n' )
if numbers.value == '3':
exports.write('Run' + '\t' + 'd50_1' + '\t' + 'Sigma1' + '\t' + 'd50_2' + '\t' + 'Sigma2'+ '\t' + 'd50_3' + '\t' + 'Sigma3'+ '\n')
exports.write(str(select.value) + '\t' + str(np.round(x50[0],2)) + '\t' + str(np.round(sigma_s[0],2)) + '\t' + str(np.round(x50[1],2)) + '\t' + str(np.round(sigma_s[1],2))+ '\t' + str(np.round(x50[2],2)) + '\t' + str(np.round(sigma_s[2],2)) + '\n' )
if numbers.value == '4':
exports.write('Run' + '\t' + 'd50_1' + '\t' + 'Sigma1' + '\t' + 'd50_2' + '\t' + 'Sigma2'+ '\t' + 'd50_3' + '\t' + 'Sigma3'+ '\t' + 'd50_4' + '\t' + 'Sigma4' + '\n')
exports.write(str(select.value) + '\t' + str(np.round(x50[0],2)) + '\t' + str(np.round(sigma_s[0],2)) + '\t' + str(np.round(x50[1],2)) + '\t' + str(np.round(sigma_s[1],2))+ '\t' + str(np.round(x50[2],2)) + '\t' + str(np.round(sigma_s[2],2))+ '\t' + str(np.round(x50[3],2)) + '\t' + str(np.round(sigma_s[3],2)) + '\n' )
if numbers.value == '5':
exports.write('Run' + '\t' + 'd50_1' + '\t' + 'Sigma1' + '\t' + 'd50_2' + '\t' + 'Sigma2'+ '\t' + 'd50_3' + '\t' + 'Sigma3'+ '\t' + 'd50_4' + '\t' + 'Sigma4'+ '\t' + 'd50_5' + '\t' + 'Sigma5' + '\n')
exports.write(str(select.value) + '\t' + str(np.round(x50[0],2)) + '\t' + str(np.round(sigma_s[0],2)) + '\t' + str(np.round(x50[1],2)) + '\t' + str(np.round(sigma_s[1],2))+ '\t' + str(np.round(x50[2],2)) + '\t' + str(np.round(sigma_s[2],2))+ '\t' + str(np.round(x50[3],2)) + '\t' + str(np.round(sigma_s[3],2))+ '\t' + str(np.round(x50[4],2)) + '\t' + str(np.round(sigma_s[4],2)) + '\n' )
else:
with open(os.path.join('results/', file_name + '_'+ date +'_results.txt'), 'a') as exports:
if numbers.value == '1':
exports.write(str(select.value) + '\t' + str(np.round(x50[0],2)) + '\t' + str(np.round(sigma_s[0],2)) + '\n' )
if numbers.value == '2':
exports.write(str(select.value) + '\t' + str(np.round(x50[0],2)) + '\t' + str(np.round(sigma_s[0],2)) + '\t' + str(np.round(x50[1],2)) + '\t' + str(np.round(sigma_s[1],2)) + '\n' )
if numbers.value == '3':
exports.write(str(select.value) + '\t' + str(np.round(x50[0],2)) + '\t' + str(np.round(sigma_s[0],2)) + '\t' + str(np.round(x50[1],2)) + '\t' + str(np.round(sigma_s[1],2))+ '\t' + str(np.round(x50[2],2)) + '\t' + str(np.round(sigma_s[2],2)) + '\n' )
if numbers.value == '4':
exports.write(str(select.value) + '\t' + str(np.round(x50[0],2)) + '\t' + str(np.round(sigma_s[0],2)) + '\t' + str(np.round(x50[1],2)) + '\t' + str(np.round(sigma_s[1],2))+ '\t' + str(np.round(x50[2],2)) + '\t' + str(np.round(sigma_s[2],2))+ '\t' + str(np.round(x50[3],2)) + '\t' + str(np.round(sigma_s[3],2)) + '\n' )
if numbers.value == '5':
exports.write(str(select.value) + '\t' + str(np.round(x50[0],2)) + '\t' + str(np.round(sigma_s[0],2)) + '\t' + str(np.round(x50[1],2)) + '\t' + str(np.round(sigma_s[1],2))+ '\t' + str(np.round(x50[2],2)) + '\t' + str(np.round(sigma_s[2],2))+ '\t' + str(np.round(x50[3],2)) + '\t' + str(np.round(sigma_s[3],2))+ '\t' + str(np.round(x50[4],2)) + '\t' + str(np.round(sigma_s[4],2)) + '\n' )
else:
with saved:
saved.clear_output()
print('Data already saved!')
# -
# # Load Data
#
# <font color='red'>Choose the path, date of measurement and file name. Look up the number of header lines, data points and data sets.</font>
# +
'''This part has to be updated to the current measurement file'''
#path = 'm:/Messdaten/SMPS/SiO2_standard/kriss_301-04-002'
#date = '2020-11-17'
#file_name ='kriss_301-04-002_002'
file_name = 'example_data'
now = datetime.now()
date =now.strftime("%Y-%m-%d-%Y")
header = 34
datapoints = 43
no_data_sets = 3
'''Up to here'''
#with open(os.path.join(path,date +'/','Exports/', file_name + '.txt')) as data_file:
with open(os.path.join(file_name + '.txt')) as data_file:
data = np.loadtxt(data_file,unpack=True, skiprows = header, max_rows=datapoints)
d = data[0] #define colum of diameter
data_used = data[1:] #define respective data sets; 0 is the diameter
#normalization to total counts
for j in range(data_used.__len__()):
summed = 0
for i in range(len(d)):
summed = summed+ data_used[j][i]
data_used[j][:]= (data_used[j][:]/summed)
sets=[] #list for the number of data sets
for i in range(len(data_used)):
sets.append(i+1)
# -
# # Interactive Plot for Setting Parameters
#
# <font color='red'>Determine roughly the values for each peak, set the numbers of peaks and the fitting range.</font>
# +
start_parameter=[np.log(np.mean(d)), 0.06, 1,(d[0],d[-1])] # initial parameters
x_dummies = np.linspace(d[0],d[-1],200)
# Interactive plot
def interactive_plot(center, sigma, scale, sli_range):
log_func[0].set_ydata(defs.log_norm_func1(x_dummies, center, sigma, scale))
left_bound.set_xdata(sli_range[0])
right_bound.set_xdata(sli_range[1])
fig, ax = plt.subplots()
data_plot = ax.bar(d, data_used[0], width = d/40)
log_func = ax.plot(x_dummies, defs.log_norm_func1(x_dummies,*start_parameter[0:3]), color ='tab:orange')
left_bound =ax.axvline(d[0], color ='tab:red')
right_bound =ax.axvline(d[-1], color ='tab:red')
ax.set_xlim(0.9*d[0],1.1*d[-1])
#ax.set_ylim(0,np.amax(data_used[0][:]))
interactive_plot = widgets.interactive(interactive_plot,sli_range =widgets.FloatRangeSlider(value=[d[0], d[-1]],min=d[0],max=d[-1],step=0.5,description='Range',readout_format='.1f'), center=widgets.FloatSlider(min = 1.0, max = 5.0, step = 0.01, value = start_parameter[0],description='Center'), sigma=widgets.FloatSlider(min = 0.01, max = 0.5, step = 0.01, value = start_parameter[1],description='Sigma'), scale=widgets.FloatSlider(min = 0.01, max = 2, value = start_parameter[2],step = 0.01,description='Scale'))
numbers = widgets.Dropdown(options=['1', '2', '3', '4', '5'], value='1', description='# peaks:')
reset_button = widgets.Button(description = "Reset")
reset_button.on_click(reset_values)
display(interactive_plot)
display(widgets.HBox([reset_button,numbers]))
center = interactive_plot.children[0]
sigma = interactive_plot.children[1]
scale = interactive_plot.children[2]
out = widgets.interactive_output(print_out, {'center': center, 'sigma':sigma,'scale':scale})
display(out)
# -
# # Get Set of Start Parameters for Fitting
#
# <font color='red'>Copy the values to the respective peak position.</font>
def get_parameter(number):
if number == '1':
parameter_set = [3.11, 0.04, 0.51]
elif number == '2':
parameter_set = [3.11, 0.04, 0.51, 3.39, 0.04, 0.32]
elif number == '3':
parameter_set = [3.11, 0.04, 0.51, 3.39, 0.04, 0.32, 3.56, 0.04, 0.1]
elif number == '4':
parameter_set = [3.11, 0.04, 0.51, 3.39, 0.04, 0.32, 3.56, 0.04, 0.1, 3.7, 0.06, 0.06]
elif number == '5':
parameter_set = [3.11, 0.04, 0.51, 3.39, 0.04, 0.32, 3.56, 0.04, 0.1, 3.7, 0.05, 0.06, 3.82, 0.04, 0.03]
else:
print('Wrong input')
return parameter_set
parameter_set = get_parameter(numbers.value)
# # Select Data
select = widgets.Dropdown(options=sets, description='Data set:', value = sets[0])
display(select)
# # Fit Log Norm Function to Data
# +
#clear lists
x50 = []
sigma_s = []
# define the fitting range
d_range = [interactive_plot.kwargs['sli_range'][0],interactive_plot.kwargs['sli_range'][1]]
index_left = (np.abs(d_range[0]-d)).argmin()
index_right = (np.abs(d_range[1]-d)).argmin()
x_fit = np.linspace(d[index_left],d[index_right],200)
#set y values and calculate error in y
y_values = data_used[select.value-1]
y_error = 1/np.sqrt(y_values)
#plot data and fits
fig2, ax2 = plt.subplots()
data_plot2 = ax2.bar(d, data_used[select.value-1], label ='Data', width = d/40)
if numbers.value == '1':
fit_lines = ax2.plot(x_fit, defs.log_norm_func1(x_fit, *parameter_set), '-', label='fit', color = 'tab:orange')
if numbers.value == '2':
fit_lines = ax2.plot(x_fit, defs.log_norm_func2(x_fit, *parameter_set), '-', label='fit', color = 'tab:orange')
if numbers.value == '3':
fit_lines = ax2.plot(x_fit, defs.log_norm_func3(x_fit, *parameter_set), '-', label='fit', color = 'tab:orange')
if numbers.value == '4':
fit_lines = ax2.plot(x_fit, defs.log_norm_func4(x_fit, *parameter_set), '-', label='fit', color = 'tab:orange')
if numbers.value == '5':
fit_lines = ax2.plot(x_fit, defs.log_norm_func5(x_fit, *parameter_set), '-', label='fit', color = 'tab:orange')
ax2.set_ylim(0,np.amax(y_values)+0.05*np.amax(y_values))
ax2.set_xlim(0.9*d[0],1.1*d[-1])
ax2.set_xlabel('Diameter/nm')
ax2.set_ylabel('Fraction')
ax2.legend(edgecolor='black')
fit_results = widgets.Output()
run_fit_button = widgets.Button(description = "Fit!",button_style ='success')
run_fit_button.on_click(run_fit)
saved = widgets.Output()
save_button = widgets.Button(description = 'Save!', button_style = 'warning')
save_button.on_click(save_data)
display(widgets.HBox([run_fit_button, fit_results]))
display(widgets.HBox([save_button, saved]))
# -
|
Interactive_fit_main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# *Accompanying code examples of the book "Introduction to Artificial Neural Networks and Deep Learning: A Practical Guide with Applications in Python" by [<NAME>](https://sebastianraschka.com). All code examples are released under the [MIT license](https://github.com/rasbt/deep-learning-book/blob/master/LICENSE). If you find this content useful, please consider supporting the work by buying a [copy of the book](https://leanpub.com/ann-and-deeplearning).*
#
# Other code examples and content are available on [GitHub](https://github.com/rasbt/deep-learning-book). The PDF and ebook versions of the book are available through [Leanpub](https://leanpub.com/ann-and-deeplearning).
# # Appendix G - TensorFlow Basics
# %load_ext watermark
# %watermark -a '<NAME>' -d -p tensorflow,numpy
# ### Table of Contents
#
#
# - TensorFlow in a Nutshell
# - Installation
# - Computation Graphs Variables
# - Placeholder Variables
# - Saving and Restoring Models
# - Naming TensorFlow Objects
# - CPU and GPU
# - TensorBoard
#
# This appendix offers a brief overview of TensorFlow, an open-source library for numerical computation and deep learning. This section is intended for readers who want to gain a basic overview of this library before progressing through the hands-on sections that are concluding the main chapters.
#
# The majority of *hands-on* sections in this book focus on TensorFlow and its Python API, assuming that you have TensorFlow >=1.2 installed if you are planning to execute the code sections shown in this book.
#
# In addition to glancing over this appendix, I recommend the following resources from TensorFlow's official documentation for a more in-depth coverage on using TensorFlow:
#
# - **[Download and setup instructions](https://www.tensorflow.org/get_started/os_setup)**
# - **[Python API documentation](https://www.tensorflow.org/api_docs/python/)**
# - **[Tutorials](https://www.tensorflow.org/tutorials/)**
# - **[TensorBoard, an optional tool for visualizing learning](https://www.tensorflow.org/how_tos/summaries_and_tensorboard/)**
# ## TensorFlow in a Nutshell
# At its core, TensorFlow is a library for efficient multidimensional array operations with a focus on deep learning. Developed by the Google Brain Team, TensorFlow was open-sourced on November 9th, 2015. And augmented by its convenient Python API layer, TensorFlow has gained much popularity and wide-spread adoption in industry as well as academia.
#
# TensorFlow shares some similarities with NumPy, such as providing data structures and computations based on multidimensional arrays. What makes TensorFlow particularly suitable for deep learning, though, are its primitives for defining functions on tensors, the ability of parallelizing tensor operations, and convenience tools such as automatic differentiation.
#
# While TensorFlow can be run entirely on a CPU or multiple CPUs, one of the core strength of this library is its support of GPUs (Graphical Processing Units) that are very efficient at performing highly parallelized numerical computations. In addition, TensorFlow also supports distributed systems as well as mobile computing platforms, including Android and Apple's iOS.
#
# But what is a *tensor*? In simplifying terms, we can think of tensors as multidimensional arrays of numbers, as a generalization of scalars, vectors, and matrices.
#
# 1. Scalar: $\mathbb{R}$
# 2. Vector: $\mathbb{R}^n$
# 3. Matrix: $\mathbb{R}^n \times \mathbb{R}^m$
# 4. 3-Tensor: $\mathbb{R}^n \times \mathbb{R}^m \times \mathbb{R}^p$
# 5. ...
#
# When we describe tensors, we refer to its "dimensions" as the *rank* (or *order*) of a tensor, which is not to be confused with the dimensions of a matrix. For instance, an $m \times n$ matrix, where $m$ is the number of rows and $n$ is the number of columns, would be a special case of a rank-2 tensor. A visual explanation of tensors and their ranks is given is the figure below.
#
# 
#
# ## Installation
# Code conventions in this book follow the Python 3.x syntax, and while the code examples should be backward compatible to Python 2.7, I highly recommend the use of Python >=3.5.
#
# Once you have your Python Environment set up ([Appendix - Python Setup]), the most convenient ways for installing TensorFlow are via `pip` or `conda` -- the latter only applies if you have the Anaconda/Miniconda Python distribution installed, which I prefer and recommend.
#
# Since TensorFlow is under active development, I recommend you to consult the official "[Download and Setup](https://www.tensorflow.org/get_started/os_setup)" documentation for detailed installation instructions to install TensorFlow on you operating system, macOS, Linux, or Windows.
#
# ## Computation Graphs
# In contrast to other tools such as NumPy, the numerical computations in TensorFlow can be categorized into two steps: a construction step and an execution step. Consequently, the typical workflow in TensorFlow can be summarized as follows:
#
# - Build a computational graph
# - Start a new *session* to evaluate the graph
# - Initialize variables
# - Execute the operations in the compiled graph
#
# Note that the computation graph has no numerical values before we initialize and evaluate it. To see how this looks like in practice, let us set up a new graph for computing the column sums of a matrix, which we define as a constant tensor (`reduce_sum` is the TensorFlow equivalent of NumPy's `sum` function).
#
# +
import tensorflow as tf
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.constant([[1., 2.],
[3., 4.],
[5., 6.]], dtype=tf.float32)
col_sum = tf.reduce_sum(tf_x, axis=0)
print('tf_x:\n', tf_x)
print('\ncol_sum:\n', col_sum)
# -
# As we can see from the output above, the operations in the graph are represented as `Tensor` objects that require an explicit evaluation before the `tf_x` matrix is populated with numerical values and its column sum gets computed.
#
# Now, we pass the graph that we created earlier to a new, active *session*, where the graph gets compiled and evaluated:
# +
with tf.Session(graph=g) as sess:
mat, csum = sess.run([tf_x, col_sum])
print('mat:\n', mat)
print('\ncsum:\n', csum)
# -
# Note that if we are only interested in the result of a particular operation, we don't need to `run` its dependencies -- TensorFlow will automatically take care of that. For instance, we can directly fetch the numerical values of `col_sum_times_2` in the active session without explicitly passing `col_sum` to `sess.run(...)` as the following example illustrates:
# +
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.constant([[1., 2.],
[3., 4.],
[5., 6.]], dtype=tf.float32)
col_sum = tf.reduce_sum(tf_x, axis=0)
col_sum_times_2 = col_sum * 2
with tf.Session(graph=g) as sess:
csum_2 = sess.run(col_sum_times_2)
print('csum_2:\n', csum_2)
# -
# ## Variables
# Variables are constructs in TensorFlow that allows us to store and update parameters of our models in the current session during training. To define a "variable" tensor, we use TensorFlow's `Variable()` constructor, which looks similar to the use of `constant` that we used to create a matrix previously. However, to execute a computational graph that contains variables, we must initialize all variables in the active session first (using `tf.global_variables_initializer()`), as illustrated in the example below.
#
# +
import tensorflow as tf
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]], dtype=tf.float32)
x = tf.constant(1., dtype=tf.float32)
# add a constant to the matrix:
tf_x = tf_x + x
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
result = sess.run(tf_x)
print(result)
# -
# Now, let us do an experiment and evaluate the same graph twice:
# +
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
result = sess.run(tf_x)
result = sess.run(tf_x)
print(result)
# -
# As we can see, the result of running the computation twice did not affect the numerical values fetched from the graph. To update or to assign new values to a variable, we use TensorFlow's `assign` operation. The function syntax of `assign` is `assign(ref, val, ...)`, where '`ref`' is updated by assigning '`value`' to it:
#
# +
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]], dtype=tf.float32)
x = tf.constant(1., dtype=tf.float32)
update_tf_x = tf.assign(tf_x, tf_x + x)
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
result = sess.run(update_tf_x)
result = sess.run(update_tf_x)
print(result)
# -
# As we can see, the contents of the variable `tf_x` were successfully updated twice now; in the active session we
#
# - initialized the variable `tf_x`
# - added a constant scalar `1.` to `tf_x` matrix via `assign`
# - added a constant scalar `1.` to the previously updated `tf_x` matrix via `assign`
#
# Although the example above is kept simple for illustrative purposes, variables are an
# important concept in TensorFlow, and we will see throughout the chapters, they are
# not only useful for updating model parameters but also for saving and loading
# variables for reuse.
# ## Placeholder Variables
#
# Another important concept in TensorFlow is the use of placeholder variables,
# which allow us to feed the computational graph with numerical values in an active session at runtime.
#
# In the following example, we will define a computational graph that performs a
# simple matrix multiplication operation. First, we define a placeholder variable
# that can hold 3x2-dimensional matrices. And after initializing the placeholder
# variable in the active session, we will use a dictionary, `feed_dict` we feed
# a NumPy array to the graph, which then evaluates the matrix multiplication operation.
#
# +
import tensorflow as tf
import numpy as np
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.placeholder(dtype=tf.float32,
shape=(3, 2))
output = tf.matmul(tf_x, tf.transpose(tf_x))
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
np_ary = np.array([[3., 4.],
[5., 6.],
[7., 8.]])
feed_dict = {tf_x: np_ary}
print(sess.run(output,
feed_dict=feed_dict))
# -
# Throughout the main chapters, we will make heavy use of placeholder variables,
# which allow us to pass our datasets to various learning algorithms
# in the computational graphs.
#
# ## Saving and Loading Variables
# Training deep neural networks requires a lot of computations and computational resources, and in practice, it would be infeasible to retrain our model each time we start a new TensorFlow session before we can use it to make predictions. In this section, we will go over the basics of saving and re-using the results of our TensorFlow models.
#
# The most convenient way to store the main components of our model is to use TensorFlows `Saver` class (`tf.train.Saver()`). To see how it works, let us reuse the simple example from the [Variables](#variables) section, where we added a constant `1.` to all elements in a 3x2 matrix:
#
# +
import tensorflow as tf
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]], dtype=tf.float32)
x = tf.constant(1., dtype=tf.float32)
update_tf_x = tf.assign(tf_x, tf_x + x)
# initialize a Saver, which gets all variables
# within this computation graph context
saver = tf.train.Saver()
# -
# Now, after we initialized the graph above, let us execute its operations in a new session:
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
result = sess.run(update_tf_x)
saver.save(sess, save_path='./my-model.ckpt')
# Notice the `saver.save` call above, which saves all variables in the graph to "checkpoint" files bearing the prefix `my-model.ckpt` in our local directory (`'./'`). Since we didn't specify which variables we wanted to save when we instantiated a `tf.train.Saver()`, it saved all variables in the graph by default -- here, we only have one variable, `tf_x`. Alternatively, if we are only interested in keeping particular variables, we can specify this by feeding `tf.train.Saver()` a dictionary or list of these variables upon instantiation. For example, if our graph contained more than one variable, but we were only interested in saving `tf_x`, we could instantiate a `saver` object as `tf.train.Saver([tf_x])`.
#
# After we executed the previous code example, we should find the three `my-model.ckpt` files (in binary format) in our local directory:
#
# - `my-model.ckpt.data-00000-of-00001`
# - `my-model.ckpt.index`
# - `my-model.ckpt.meta`
#
# The file `my-model.ckpt.data-00000-of-00001` saves our main variable values, the `.index` file keeps track of the data structures, and the `.meta` file describes the structure of our computational graph that we executed.
#
# Note that in our simple example above, we just saved our variable one single time. However, in real-world applications, we typically train models over multiple iterations or epochs, and it is useful to create intermediate checkpoint files during training so that we can pick up where we left off in case we need to interrupt our session or encounter unforeseen technical difficulties. For instance, by using the `global_step` parameter, we could save our results after each 10th iteration by making the following modification to our code:
#
# +
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]], dtype=tf.float32)
x = tf.constant(1., dtype=tf.float32)
update_tf_x = tf.assign(tf_x, tf_x + x)
# initialize a Saver, which gets all variables
# within this computation graph context
saver = tf.train.Saver()
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(100):
result = sess.run(update_tf_x)
if not epoch % 10:
saver.save(sess,
save_path='./my-model-multiple_ckpts.ckpt',
global_step=epoch)
# -
# After we executed this code we find five `my-model.ckpt` files in our local directory:
#
# - `my-model.ckpt-50 {.data-00000-of-00001, .ckpt.index, .ckpt.meta}`
# - `my-model.ckpt-60 {.data-00000-of-00001, .ckpt.index, .ckpt.meta}`
# - `my-model.ckpt-70 {.data-00000-of-00001, .ckpt.index, .ckpt.meta}`
# - `my-model.ckpt-80 {.data-00000-of-00001, .ckpt.index, .ckpt.meta}`
# - `my-model.ckpt-90 {.data-00000-of-00001, .ckpt.index, .ckpt.meta}`
#
# Although we saved our variables ten times, the `saver` only keeps the five most recent checkpoints by default to save storage space. However, if we want to keep more than five recent checkpoint files, we can provide an optional argument `max_to_keep=n` when we initialize the `saver`, where `n` is an integer specifying the number of the most recent checkpoint files we want to keep.
#
# Now that we learned how to save TensorFlow `Variable`s, let us see how we can restore them. Assuming that we started a fresh computational session, we need to specify the graph first. Then, we can use the `saver`'s `restore` method to restore our variables as shown below:
#
# +
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]], dtype=tf.float32)
x = tf.constant(1., dtype=tf.float32)
update_tf_x = tf.assign(tf_x, tf_x + x)
# initialize a Saver, which gets all variables
# within this computation graph context
saver = tf.train.Saver()
with tf.Session(graph=g) as sess:
saver.restore(sess, save_path='./my-model.ckpt')
result = sess.run(update_tf_x)
print(result)
# -
# Notice that the returned values of the `tf_x` `Variable` are now increased by a constant of two, compared to the values in the computational graph. The reason is that we ran the graph one time before we saved the variable,
#
# ```python
# with tf.Session(graph=g) as sess:
# sess.run(tf.global_variables_initializer())
# result = sess.run(update_tf_x)
#
# # save the model
# saver.save(sess, save_path='./my-model.ckpt')
# ```
#
# and we ran it a second time when after we restored the session.
#
#
# Similar to the example above, we can reload one of our checkpoint files by providing the desired checkpoint suffix (here: `-90`, which is the index of our last checkpoint):
#
#
with tf.Session(graph=g) as sess:
saver.restore(sess, save_path='./my-model-multiple_ckpts.ckpt-90')
result = sess.run(update_tf_x)
print(result)
# In this section, we merely covered the basics of saving and restoring TensorFlow models. If you want to learn more, please take a look at the official [API documentation](https://www.tensorflow.org/api_docs/python/tf/train/Saver) of TensorFlow's `Saver` class.
#
# ## Naming TensorFlow Objects
# When we create new TensorFlow objects like `Variables`, we can provide an optional argument for their `name` parameter -- for example:
#
# ```python
# tf_x = tf.Variable([[1., 2.],
# [3., 4.],
# [5., 6.]],
# name='tf_x_0',
# dtype=tf.float32)
# ```
#
# Assigning names to `Variable`s explicitly is not a requirement, but I personally recommend making it a habit when building (more) complex models. Let us walk through a scenario to illustrate the importance of naming variables, taking the simple example from the previous section and add new variable `tf_y` to the graph:
#
# +
import tensorflow as tf
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]], dtype=tf.float32)
tf_y = tf.Variable([[7., 8.],
[9., 10.],
[11., 12.]], dtype=tf.float32)
x = tf.constant(1., dtype=tf.float32)
update_tf_x = tf.assign(tf_x, tf_x + x)
saver = tf.train.Saver()
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
result = sess.run(update_tf_x)
saver.save(sess, save_path='./my-model.ckpt')
# -
# The variable `tf_y` does not do anything in the code example above; we added it for illustrative purposes, as we will see in a moment. Now, let us assume we started a new computational session and loaded our saved my-model into the following computational graph:
#
# +
g = tf.Graph()
with g.as_default() as g:
tf_y = tf.Variable([[7., 8.],
[9., 10.],
[11., 12.]], dtype=tf.float32)
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]], dtype=tf.float32)
x = tf.constant(1., dtype=tf.float32)
update_tf_x = tf.assign(tf_x, tf_x + x)
saver = tf.train.Saver()
with tf.Session(graph=g) as sess:
saver.restore(sess, save_path='./my-model.ckpt')
result = sess.run(update_tf_x)
print(result)
# -
# Unless you paid close attention on how we initialized the graph above, this result above surely was not the one you expected. What happened? Intuitively, we expected our session to `print`
#
# ```python
# [[ 3. 4.]
# [ 5. 6.]
# [ 7. 8.]]
# ```
# The explanation behind this unexpected `result` is that we reversed the order of `tf_y` and `tf_x` in the graph above. TensorFlow applies a default naming scheme to all operations in the computational graph, unless we use do it explicitly via the `name` parameter -- or in other words, we confused TensorFlow by reversing the order of two similar objects, `tf_y` and `tf_x`.
#
# To circumvent this problem, we could give our variables specific names -- for example, `'tf_x_0'` and `'tf_y_0'`:
#
# +
import tensorflow as tf
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]],
name='tf_x_0',
dtype=tf.float32)
tf_y = tf.Variable([[7., 8.],
[9., 10.,
]],
name='tf_y_0',
dtype=tf.float32)
x = tf.constant(1., dtype=tf.float32)
update_tf_x = tf.assign(tf_x, tf_x + x)
saver = tf.train.Saver()
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
result = sess.run(update_tf_x)
saver.save(sess, save_path='./my-model.ckpt')
# -
# Then, even if we flip the order of these variables in a new computational graph, TensorFlow knows which values to use for each variable when loading our model -- assuming we provide the corresponding variable names:
# +
g = tf.Graph()
with g.as_default() as g:
tf_y = tf.Variable([[7., 8.],
[9., 10.,
]],
name='tf_y_0',
dtype=tf.float32)
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]],
name='tf_x_0',
dtype=tf.float32)
x = tf.constant(1., dtype=tf.float32)
update_tf_x = tf.assign(tf_x, tf_x + x)
saver = tf.train.Saver()
with tf.Session(graph=g) as sess:
saver.restore(sess, save_path='./my-model.ckpt')
result = sess.run(update_tf_x)
print(result)
# -
# ## CPU and GPU
# Please note that all code examples in this book, and all TensorFlow operations in general, can be executed on a CPU. If you have a GPU version of TensorFlow installed, TensorFlow will automatically execute those operations that have GPU support on GPUs and use your machine's CPU, otherwise.
# However, if you wish to define your computing device manually, for instance, if you have the GPU version installed but want to use the main CPU for prototyping, we can run an active section on a specific device using the `with` context as follows
#
# with tf.Session() as sess:
# with tf.device("/gpu:1"):
#
# where
#
# - "/cpu:0": The CPU of your machine.
# - "/gpu:0": The GPU of your machine, if you have one.
# - "/gpu:1": The second GPU of your machine, etc.
# - etc.
#
# You can get a list of all available devices on your machine via
#
# from tensorflow.python.client import device_lib
#
# device_lib.list_local_devices()
#
# For more information on using GPUs in TensorFlow, please refer to the GPU documentation at https://www.tensorflow.org/how_tos/using_gpu/.
#
#
#
# ```python
# with tf.Session() as sess:
# with tf.device("/gpu:1"):
#
#
#
#
# from tensorflow.python.client import device_lib
#
# device_lib.list_local_devices()
# ```
# Another good way to check whether your current TensorFlow session runs on a GPU is to execute
#
# ```python
# >>> import tensorflow as tf
# >>> tf.test.gpu_device_name()
# ```
# In your current Python session. If a GPU is available to TensorFlow, it will return a non-empty string; for example, `'/gpu:0'`. Otherwise, if now GPU can be found, the function will return an empty string.
# ## Control Flow
# It is important to discuss TensorFlow's control flow mechanics, the way it handles control statements such as `if/else` and `while`-loops. Control flow in TensorFlow is not a complicated topic, but it can be quite unintuitive at first and a common pitfall for beginners -- especially, in the context of how control flow in Python is handled.
#
# To explain control flow in TensorFlow in a practical manner, let us consider a simple example first. The following graph is meant to add the value `1.0` to a placeholder variable `x` if `addition=True` and subtract `1.0` from `x` otherwise:
# +
import tensorflow as tf
addition = True
g = tf.Graph()
with g.as_default() as g:
x = tf.placeholder(dtype=tf.float32, shape=None)
if addition:
y = x + 1.
else:
y = x - 1.
# -
# Now, let us create a new session and execute the graph by feeding a `1.0` to the placeholder. If everything works as expected, the session should return the value `2.0` since `addition=True` and `1.0 + 1.0 = 2.0`:
# +
with tf.Session(graph=g) as sess:
result = sess.run(y, feed_dict={x: 1.})
print('Result:\n', result)
# -
# The previous session call clearly yielded the resulted we expected. Next, let us set `addition=False` to also check the other scenario, that is, subtracting `1.0` from `x`:
# +
addition = False
with tf.Session(graph=g) as sess:
result = sess.run(y, feed_dict={x: 1.})
print('Result:\n', result)
# -
# It appears that the session did return the same value that it returned when `addition` was set to `True`. Why did this happen? The explanation for this is that the `if/else` statements in the previous code only apply to the graph construction step. Or in other words, we created a graph by visiting the code contained under the `if` statement, and since TensorFlow graphs are static, we have no way of running the code under the `else` statement -- except for setting `addition=False` and creating a new graph.
# However, we do not have to create a new graph each time we want to include control statements -- TensorFlow implements a variety of helper functions that help with control flow inside a graph. For instance, to accomplish the little exercise of conditionally adding or subtracting a one from the placeholder variable `x`, we could use `tf.cond` as follows:
# +
addition = True
g = tf.Graph()
with g.as_default() as g:
addition = tf.placeholder(dtype=tf.bool, shape=None)
x = tf.placeholder(dtype=tf.float32, shape=None)
y = tf.cond(addition,
true_fn=lambda: tf.add(x, 1.),
false_fn=lambda: tf.subtract(x, 1.))
# -
# The basic use of `tf.cond` for conditional execution comes with three important arguments: a condition to check (here, if addition is `True` or `False`), a function that gets executed if the condition is `True` (`true_fn`) and a function that gets executed if the condition is `False` (`false_fn`), respectively.
#
# Next, let us repeat the little exercise from earlier and see if toggling the `addition` value between `True` and `False` affects the conditional execution that is now part of the graph:
# +
with tf.Session(graph=g) as sess:
result = sess.run(y, feed_dict={addition:True,
x: 1.})
print('Result:\n', result)
# +
with tf.Session(graph=g) as sess:
result = sess.run(y, feed_dict={addition:False,
x: 1.})
print('Result:\n', result)
# -
# Finally, we get the expected results
#
# - "1.0 + 1.0 = 2.0" if `addition=True`
# - "1.0 - 1.0 = 0.0" if `addition=False`
#
# While this section provides you with the most important concept behind control flow in Python versus TensorFlow, there are many control statements (and logical operators) that we have not covered. Since the use of other control statements is analogous to `tf.cond`, I recommend you to visit [TensorFlow's API documentation](https://www.tensorflow.org/api_guides/python/control_flow_ops), which provides an overview of all the different operators for control flow and links to useful examples.
# ## TensorBoard
# TensorBoard is one of the coolest features of TensorFlow, which provides us with a suite of tools to visualize our computational graphs and operations before and during runtime. Especially, when we are implementing large neural networks, our graphs can be quite complicated, and TensorBoard is only useful to visually track the training cost and performance of our network, but it can also be used as an additional tool for debugging our implementation. In this section, we will go over the basic concepts of TensorBoard, but make sure you also check out the [official documentation](https://www.tensorflow.org/how_tos/summaries_and_tensorboard/) for more details.
#
# To visualize a computational graph via TensorBoard, let us create a simple graph with two `Variable`s, the tensors `tf_x` and `tf_y` with shape `[2, 3]`. The first operation is to add these two tensors together. Second, we transpose `tf_x` and multiply it with `tf_y`:
# +
import tensorflow as tf
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]],
name='tf_x_0',
dtype=tf.float32)
tf_y = tf.Variable([[7., 8.],
[9., 10.],
[11., 12.]],
name='tf_y_0',
dtype=tf.float32)
output = tf_x + tf_y
output = tf.matmul(tf.transpose(tf_x), output)
# -
# If we want to visualize the graph via TensorBoard, we need to instantiate a new `FileWriter` object in our session, which we provide with a `logdir` and the graph itself. The `FileWriter` object will then write a [protobuf](https://developers.google.com/protocol-buffers/docs/overview) file to the `logdir` path that we can load into TensorBoard:
#
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
# create FileWrite object that writes the logs
file_writer = tf.summary.FileWriter(logdir='logs/1', graph=g)
result = sess.run(output)
print(result)
# If you installed TensorFlow via `pip`, the `tensorboard` command should be available from your command line terminal. So, after running the preceding code examples for defining the grap and running the session, you just need to execute the command `tensorboard --logdir logs/1`. You should then see an output similar to the following:
#
# Desktop Sebastian{$$} tensorboard --logdir logs/1
# Starting TensorBoard b'41' on port 6006
# (You can navigate to http://xxx.xxx.x.xx:6006)
#
# Copy and paste the `http` address from the terminal and open it in your favorite web browser to open the TensorBoard window. Then, click on the `Graph` tab at the top, to visualize the computational graph as shown in the figure below:
#
# 
#
# In our TensorBoard window, we can now see a visual summary of our computational graph (as shown in the screenshot above). The dark-shaded nodes labeled as `tf_x_0` and `tf_y_0` are the two variables we initializes, and following the connective lines, we can track the flow of operations. We can see the graph edges that are connecting `tf_x_0` and `tf_y_0` to an `add` node, with is the addition we defined in the graph, followed by the multiplication with the transpose of and the result of `add`.
#
# Next, we are introducing the concept of `name_scope`s, which lets us organize different parts in our graph. In the following code example, we are going to take the initial code snippets and add `with tf.name_scope(...)` contexts as follows:
#
#
# +
# Graph visualization with name scopes
g = tf.Graph()
with g.as_default() as g:
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]],
name='tf_x_0',
dtype=tf.float32)
tf_y = tf.Variable([[7., 8.],
[9., 10.],
[11., 12.]],
name='tf_y_0',
dtype=tf.float32)
# add custom name scope
with tf.name_scope('addition'):
output = tf_x + tf_y
# add custom name scope
with tf.name_scope('matrix_multiplication'):
output = tf.matmul(tf.transpose(tf_x), output)
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
file_writer = tf.summary.FileWriter(logdir='logs/2', graph=g)
result = sess.run(output)
print(result)
# -
# After executing the code example above, quit your previous TensorBoard session by pressing `CTRL+C` in the command line terminal and launch a new TensorBoard session via `tensorboard --logdir logs/2`. After you refreshed your browser window, you should see the following graph:
#
#
# 
#
# Comparing this visualization to our initial one, we can see that our operations have been grouped into our custom name scopes. If we double-click on one of these name scope summary nodes, we can expand it and inspect the individual operations in more details as shown for the `matrix_multiplication` name scope in the screenshot below:
#
# 
#
# So far, we have only been looking at the computational graph itself. However, TensorBoard implements many more useful features. In the following example, we will make use of the "Scalar" and "Histogram" tabs. The "Scalar" tab in TensorBoard allows us to track scalar values over time, and the "Histogram" tab is useful for displaying the distribution of value in our tensor `Variable`s (for instance, the model parameters during training). For simplicity, let us take our previous code snippet and modify it to demonstrate the capabilities of TensorBoard:
#
#
# +
# Graph visualization and variable inspection
g = tf.Graph()
with g.as_default() as g:
some_value = tf.placeholder(dtype=tf.int32,
shape=None,
name='some_value')
tf_x = tf.Variable([[1., 2.],
[3., 4.],
[5., 6.]],
name='tf_x_0',
dtype=tf.float32)
tf_y = tf.Variable([[7., 8.],
[9., 10.],
[11., 12.]],
name='tf_y_0',
dtype=tf.float32)
with tf.name_scope('addition'):
output = tf_x + tf_y
with tf.name_scope('matrix_multiplication'):
output = tf.matmul(tf.transpose(tf_x), output)
with tf.name_scope('update_tensor_x'):
tf_const = tf.constant(2., shape=None, name='some_const')
update_tf_x = tf.assign(tf_x, tf_x * tf_const)
# create summaries
tf.summary.scalar(name='some_value', tensor=some_value)
tf.summary.histogram(name='tf_x_values', values=tf_x)
# merge all summaries into a single operation
merged_summary = tf.summary.merge_all()
# -
# Notice that we added an additional `placeholder` to the graph which later receives a scalar value from the session. We also added a new operation that updates our `tf_x` tensor by multiplying it with a constant `2.`:
#
# ```python
# with tf.name_scope('update_tensor_x'):
# tf_const = tf.constant(2., shape=None, name='some_const')
# update_tf_x = tf.assign(tf_x, tf_x * tf_const)
# ```
#
# Finally, we added the lines
#
# ```python
# # create summaries
# tf.summary.scalar(name='some_value', tensor=some_value)
# tf.summary.histogram(name='tf_x_values', values=tf_x)
# ```
#
# at the end of our graph. These will create the "summaries" of the values we want to display in TensorBoard later. The last line of our graph is
#
# ```python
# merged_summary = tf.summary.merge_all()
# ```
#
# which summarizes all the `tf.summary` calls to one single operation, so that we only have to fetch one variable from the graph when we execute the session. When we executed the session, we simply fetched this merged summary from `merged_summary` as follows:
#
# ```python
# result, summary = sess.run([update_tf_x, merged_summary],
# feed_dict={some_value: i})
# ```
#
# Next, let us add a `for`-loop to our session that runs the graph five times, and feeds the counter of the `range` iterator to the `some_value` `placeholder` variable:
#
#
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
# create FileWrite object that writes the logs
file_writer = tf.summary.FileWriter(logdir='logs/3', graph=g)
for i in range(5):
# fetch the summary from the graph
result, summary = sess.run([update_tf_x, merged_summary],
feed_dict={some_value: i})
# write the summary to the log
file_writer.add_summary(summary=summary, global_step=i)
file_writer.flush()
# The two lines at the end of the preceding code snippet,
#
# ```python
# file_writer.add_summary(summary=summary, global_step=i)
# file_writer.flush()
# ```
#
# will write the summary data to our log file and the `flush` method updates TensorBoard. Executing `flush` explicitely is usually not necessary in real-world applications, but since the computations in our graph are so simple and "cheap" to execute, TensorBoard may not fetch the updates in real time.
# To visualize the results, quit your previous TensorBoard session (via `CTRL+C`) and execute `tensorboard --logdir logs/3` from the command line. In the TensorBoard window under the tab "Scalar," you should now see an entry called "some_value_1," which refers to our `placeholder` in the graph that we called `some_value`. Since we just fed it the iteration index of our `for`-loop, we expect to see a linear graph with the iteration index on the *x*- and *y*-axis:
#
# 
#
# Keep in mind that this is just a simple demonstration of how `tf.summary.scalar` works. For instance, more useful applications include the tracking of the training loss and the predictive performance of a model on training and validation sets throughout the different training rounds or epochs.
#
# Next, let us go to the "Distributions" tab:
#
#
# 
#
# The "Distributions" graph above shows us the distribution of values in `tf_x` for each step in the `for`-loop. Since we doubled the value in the tensor after each `for`-loop iteration, we the distribution graph grows wider over time.
#
# Finally, let us head over to the "Histograms" tab, which provides us with an individual histogram for each `for`-loop step that we can scroll through. Below, I selected the 3rd `for`-loop step that highlights the histogram of values in `tf_x` during this step:
#
#
# 
#
#
# Since TensorBoard is such a highly visual tool with graph and data exploration in mind, I highly recommend you to take it for a test drive and explore it interactively. Also, the are several features that we haven't covered in this simple introduction to TensorBoard, so be sure to check out the [official documentation](https://www.tensorflow.org/how_tos/summaries_and_tensorboard/) for more information.
|
code/_old-material/appendix_tensorflow-basics/appendix_g_tensorflow-basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/tomassams/tdt4173-machine-learning-project/blob/main/project_notebook_cnn.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="wsU1QHVNqj_I"
#
# ----
#
# <div align="center">
#
# <p align="center">
#
# <h1> TDT4173 Project Assignment </h1>
#
#
# <h1> A comparison of KNN and CNN for image classification in the gastrointestinal tract</h1>
#
# <h2> CNN Implementation </h2>
#
# </p>
#
# </div>
#
# ----
#
# + [markdown] id="P09dqkvnrG8e"
# *TDT4173 Machine Learning - NTNU, Fall 2020*
#
# This notebook is one of two notebooks containing code used in the group project assignment. It is intended for use in Google Colab, and should run without issues there. However, most cells and commands can likely also run locally with minor adjustments. Things like CLI commands in the beginning (wget, unzip, etc) might be different based on your operating system.
#
# The dataset used is the [Kvasir](https://datasets.simula.no/kvasir/) v2 dataset, containing classified images from the gastrointestinal tract.
#
# Note that some parts are computation heavy - e.g. model training in this notebook, and is preferably ran with GPU support. Colab environments are provided with GPU resources, this can be set in "Runtime => Change runtime type" in the top menu.
# + [markdown] id="4yhdyBawJmSx"
# ## 1 Setup and prerequisites
# + [markdown] id="d2hO4Ntlsr1j"
# If any of these imports return an error, you might have to install them with pip. It can be done in the command line, or by executing it in a notebook code cell. An export of the environment requirements can be found in **requirements.txt**.
# + id="pQPjrnUEePW7"
"""
If the notebook is running in Colab, we want to download models and results
when they are ready so they don't get lost if Colab times out.
This global variable keeps track of that.
"""
RunningInCOLAB = 'google.colab' in str(get_ipython())
if RunningInCOLAB:
from google.colab import files
# + id="y9yv5011yCUy"
# %matplotlib inline
from matplotlib import pyplot as plt
import os
import shutil
import pandas as pd
import numpy as np
import tensorflow as tf
from tensorflow import keras
from sklearn.metrics import confusion_matrix, classification_report
# + id="EeEEA_YxvP3m"
"""
Verify we are running the correct Tensorflow version
"""
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
# + id="e4SgSfFivZg5" colab={"base_uri": "https://localhost:8080/"} outputId="12407588-2102-4b7e-ee1a-3d60bcb3a48e"
"""
Verify that we are using a GPU runtime
If there is no GPU found, you can activate this in the Colab settings at "Runtime => Change runtime type".
If you are running this locally, GPU support might require setup of CUDA/CUDNN beforehand.
It is highly recommended to use a GPU while training the CNN models.
"""
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
raise SystemError('GPU device not found')
print('Found GPU at: {}'.format(device_name))
# + id="af3jUE8dgScS"
"""
Set up our project folder structure
"""
def create_folder_unless_exists(path):
if not os.path.exists(path):
os.makedirs(path)
create_folder_unless_exists('data/raw')
create_folder_unless_exists('data/zip')
create_folder_unless_exists('models')
create_folder_unless_exists('results/cnn_simple')
create_folder_unless_exists('results/cnn_resnet_tl')
create_folder_unless_exists('results/cnn_resnet_tl_tuned')
# + id="I45YsjIQgr4q" colab={"base_uri": "https://localhost:8080/"} outputId="f89f6a81-0837-4e4d-8416-10c0d0bfa5ea"
"""
Download and unzip the dataset
1. Downloads the zip-file with wget
2. Unzips it into the root folder
3. Moves it into the ./data/raw/ folder
4. Moves the original zip file to the ./data/zip/ folder
If you are not running in a Colab environment, you might need to do this manually.
Folder structure should look like this after:
|-- project root
| |--> ...
| |--> ...
| |--> data
| |--> zip
| |--> kvasir-dataset-v2.zip
| |--> raw
| |--> dyed-lifted-polyps
| |--> dyed-resection-margins
| |--> esophagitis
| |--> normal-cecum
| |--> normal-pylorus
| |--> normal-z-line
| |--> polyps
| |--> ulcerative-colitis
"""
# !wget "https://datasets.simula.no/kvasir/data/kvasir-dataset-v2.zip"
# !unzip -qq kvasir-dataset-v2.zip
# !cp kvasir-dataset-v2/* data/raw -r
# !cp kvasir-dataset-v2.zip data/zip -r
# !rm kvasir-dataset-v2 -rf
# !rm kvasir-dataset-v2.zip
# + id="woWgZDDlZc_u"
"""
A helper utility to generate a pretty confusion matrix
Citiation
---------
http://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html
"""
def plot_confusion_matrix(cm,
target_names,
title='Confusion matrix',
cmap=None,
normalize=True,
save_path=None):
"""
given a sklearn confusion matrix (cm), make a nice plot
Arguments
---------
cm: confusion matrix from sklearn.metrics.confusion_matrix
target_names: given classification classes such as [0, 1, 2]
the class names, for example: ['high', 'medium', 'low']
title: the text to display at the top of the matrix
cmap: the gradient of the values displayed from matplotlib.pyplot.cm
see http://matplotlib.org/examples/color/colormaps_reference.html
plt.get_cmap('jet') or plt.cm.Blues
normalize: If False, plot the raw numbers
If True, plot the proportions
Usage
-----
plot_confusion_matrix(cm = cm, # confusion matrix created by
# sklearn.metrics.confusion_matrix
normalize = True, # show proportions
target_names = y_labels_vals, # list of names of the classes
title = best_estimator_name) # title of graph
"""
import matplotlib.pyplot as plt
import numpy as np
import itertools
accuracy = np.trace(cm) / np.sum(cm).astype('float')
misclass = 1 - accuracy
if cmap is None:
cmap = plt.get_cmap('Blues')
plt.figure(figsize=(10, 8))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
# plt.title(title)
plt.colorbar()
plt.rcParams.update({'font.size': 18})
if target_names is not None:
tick_marks = np.arange(len(target_names))
plt.xticks(tick_marks, target_names, rotation=45, fontsize=18, ha="right")
plt.yticks(tick_marks, target_names, fontsize=18)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 1.5 if normalize else cm.max() / 2
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
if normalize:
plt.text(j, i, "{:0.4f}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
else:
plt.text(j, i, "{:,}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.autoscale()
plt.ylabel('True label', fontsize=16)
plt.xlabel('Predicted label\naccuracy={:0.4f}; misclass={:0.4f}'.format(accuracy, misclass), fontsize=16)
if save_path:
plt.savefig(save_path, bbox_inches='tight')
plt.show()
# + id="qbswis9FBHG-"
"""
Utility function to plot, save and download the confusion matrix
"""
def display_confusion_matrix(y_pred, y_true, target_names, save_path=None):
cm = confusion_matrix(y_true, y_pred)
plot_confusion_matrix(cm, target_names, normalize=False, save_path=save_path)
if RunningInCOLAB and save_path:
files.download(save_path)
"""
Utility function to print, save and download the classification report
"""
def display_classification_report(y_pred, y_true, target_names, save_path=None):
cr = classification_report(y_true, y_pred, target_names=target_names, output_dict=True)
df = pd.DataFrame(cr).transpose()
df = df.round(2)
print(df)
if save_path:
df.to_csv(save_path, sep=';')
if RunningInCOLAB and save_path:
files.download(save_path)
# + id="3FD2lR9QywgA"
"""
Simple helper utility to plot training history metrics against validation
"""
def plot_history(history):
# loss vs val_loss
plt.plot(history.history['loss'], label='loss')
plt.plot(history.history['val_loss'], label = 'val_loss')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.ylim([0, 1])
plt.legend(loc='right')
plt.show()
# acc vs val_acc
plt.plot(history.history['accuracy'], label='accuracy')
plt.plot(history.history['val_accuracy'], label = 'val_accuracy')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.ylim([0, 1])
plt.legend(loc='right')
plt.show()
# + id="aEMxkkm3ghiv"
"""
Even though Keras has provided an ImageDataGenerator which contains a
train/validation splitter, we actually want a train/validation/test split,
so lets do this manually.
"""
def get_files_from_folder(path):
files = os.listdir(path)
return np.asarray(files)
def split_image_dataset(raw_path, processed_path, split_ratio):
# make a copy from raw to processed
shutil.copytree(raw_path, processed_path)
# split files into train/test inside processed
train_path = os.path.join(processed_path, 'train')
test_path = os.path.join(processed_path,'test')
# get dirs
_, dirs, _ = next(os.walk(processed_path))
# calculates how many train data per class
data_counter_per_class = np.zeros((len(dirs)))
for i in range(len(dirs)):
path = os.path.join(processed_path, dirs[i])
files = get_files_from_folder(path)
data_counter_per_class[i] = len(files)
test_counter = np.round(data_counter_per_class * (1 - split_ratio))
print(f"Total data per class: {data_counter_per_class}")
print(f"Test instances per class: {test_counter}")
# transfers files
for i in range(len(dirs)):
files = get_files_from_folder(os.path.join(processed_path, dirs[i]))
train_path = os.path.join(processed_path, 'train' ,dirs[i])
test_path = os.path.join(processed_path, 'test' ,dirs[i])
if not os.path.exists(train_path):
os.makedirs(train_path)
if not os.path.exists(test_path):
os.makedirs(test_path)
# moves data
for j in range(int(data_counter_per_class[i])):
src = os.path.join(processed_path, dirs[i], files[j])
if j < test_counter[i]:
dst = os.path.join(processed_path, 'test', dirs[i], files[j])
else:
dst = os.path.join(processed_path, 'train', dirs[i], files[j])
shutil.move(src, dst)
os.removedirs(os.path.join(processed_path, dirs[i]))
# + id="4vDdEMlpmo0_" colab={"base_uri": "https://localhost:8080/"} outputId="643f9a40-7ccd-494f-f76a-da8eea2a0323"
"""
Split the dataset into 80/20 train/test and put it into 'data/processed'
"""
split_image_dataset('data/raw', 'data/processed', split_ratio=0.80)
# + id="V5OYeaSKeupJ"
"""
Parameters that apply for all models and datagenerators
"""
train_path = 'data/processed/train'
test_path = 'data/processed/test'
batch_size = 32
target_w = 256
target_h = 256
target_size = (target_w, target_h)
target_dims = 3 # (RGB channels)
# + [markdown] id="-PhNz-VZvMvQ"
# ## 2 Convolutional Neural Network
# + [markdown] id="Yu-Q2Ii26w3p"
# ### 2.1 Preprocessing
# + id="x2JRpOsS3r9u" colab={"base_uri": "https://localhost:8080/"} outputId="d22daf29-f4ed-4a21-915e-20581244fbae"
"""
Set up datagenerators for CNN model
Datagenerators handle image augmentation (shear, zoom, horizontal flips) as well as rescaling.
The datagenerator also has a built-in training/validation splitter, which creates a new 80/20 split.
In addition to that, we want a separate generator with our test set.
The dataset will be split as follows:
- Training set (64% - 80% of 80)
- Validation set (16% - 20% of 80)
- Test set (20%)
"""
"""
Datagenerator for training set
"""
cnn_train_idg = keras.preprocessing.image.ImageDataGenerator(rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
validation_split=0.2) # If defined, will split based on 'subset' in flow_from_directory()
cnn_train_datagen = cnn_train_idg.flow_from_directory(
train_path,
target_size=target_size,
batch_size=batch_size,
class_mode='sparse',
shuffle=True,
subset='training') # Marks this as the training data
cnn_val_datagen = cnn_train_idg.flow_from_directory(
train_path,
target_size=target_size,
batch_size=batch_size,
class_mode='sparse',
shuffle=False,
subset='validation') # Marks this as the validation data
"""
Datagenerator for test set
"""
cnn_test_idg = keras.preprocessing.image.ImageDataGenerator(rescale=1./255)
cnn_test_datagen = cnn_test_idg.flow_from_directory(
test_path,
target_size=target_size,
batch_size=batch_size,
class_mode='sparse',
shuffle=False)
# + [markdown] id="MSHfIPQg6yXs"
# ### 2.2 Build model
# + id="PSIhSXjXwCId"
"""
Function to build a simple CNN model layer by layer
"""
def build_simple_cnn():
model = keras.models.Sequential([
keras.layers.Conv2D(32, (3, 3), activation='relu', input_shape=(target_w, target_h, target_dims)),
keras.layers.MaxPooling2D((2, 2)),
keras.layers.Conv2D(64, (3, 3), activation='relu'),
keras.layers.MaxPooling2D((2, 2)),
keras.layers.Conv2D(64, (3, 3), activation='relu')
])
model.add(keras.layers.Flatten())
model.add(keras.layers.Dense(64, activation='relu'))
model.add(keras.layers.Dense(8, activation='softmax'))
model.summary()
return model
# + id="ufTUvzwxwC-r" colab={"base_uri": "https://localhost:8080/"} outputId="0c1929d1-909e-4a33-824c-a2c041ee4e82"
"""
Actually build and compile the model
"""
simple_cnn = build_simple_cnn()
simple_cnn.compile(
optimizer=keras.optimizers.Adam(0.001),
loss='sparse_categorical_crossentropy',
metrics=[ 'accuracy' ]
)
# + [markdown] id="zPLcJZfI6z9y"
# ### 2.3 Train model
# + id="pHaV8x1TwJzJ" colab={"base_uri": "https://localhost:8080/", "height": 697} outputId="117f4913-ec44-4d48-e7cf-f972c828e5a3"
"""
Start fitting the model
"""
cnn_history = simple_cnn.fit(cnn_train_datagen,
epochs=20,
verbose=1,
validation_data=cnn_val_datagen)
simple_cnn.save('models/cnn_simple.hdf5')
if RunningInCOLAB:
files.download('models/cnn_simple.hdf5')
# + [markdown] id="Cv9zSMXn61fS"
# ### 2.4 Evaluate model
# + id="H9d3P5lWHxaV" colab={"base_uri": "https://localhost:8080/"} outputId="16dd64dc-f6e3-4798-bcc0-e99d37040258"
"""
Use the model to predict on the unseen test set.
"""
cnn_predictions = simple_cnn.predict(cnn_test_datagen, verbose=1)
# + id="wqGeYo_Vc8K4" colab={"base_uri": "https://localhost:8080/", "height": 808} outputId="5c6b5220-c8fe-451d-9d41-d4656a0b751a"
"""
Evaluate the prediction results.
Displays a Confusion Matrix and a Classification Report.
These are also saved to the 'results' folder.
"""
unique_labels = list(cnn_test_datagen.class_indices.keys())
y_pred = np.argmax(cnn_predictions, axis=-1)
y_true = cnn_test_datagen.classes[cnn_test_datagen.index_array]
display_confusion_matrix(
y_pred=y_pred,
y_true=y_true,
target_names=unique_labels,
save_path='results/cnn_simple/cnn_simple_confusion_matrix.png'
)
display_classification_report(
y_pred=y_pred,
y_true=y_true,
target_names=unique_labels,
save_path='results/cnn_simple/cnn_simple_classification_report.csv'
)
# + [markdown] id="HHDlpspb6pPT"
# ## 3 ResNet
# + [markdown] id="CRMp-xYP67FX"
# ### 3.1 Preprocessing
# + id="KcZGN4CBfDYr" colab={"base_uri": "https://localhost:8080/"} outputId="82454d17-f9b3-4b31-ad2a-48fd1bd9639c"
"""
Set up datagenerators for ResNet50 model
Datagenerators handle image augmentation (shear, zoom, horizontal flips) as well as rescaling.
They also have built-in utility to replicate ResNet's original preprocessing steps.
The datagenerator also has a built-in training/validation splitter, which creates a new 80/20 split.
In addition to that, we want a separate generator with our test set.
The dataset will be split as follows:
- Training set (64% - 80% of 80)
- Validation set (16% - 20% of 80)
- Test set (20%)
"""
"""
Training set
"""
rn_train_idg = keras.preprocessing.image.ImageDataGenerator(preprocessing_function=keras.applications.resnet50.preprocess_input,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
validation_split=0.2) # If defined, will split based on 'subset' in flow_from_directory()
rn_train_datagen = rn_train_idg.flow_from_directory(
train_path,
target_size=target_size,
batch_size=batch_size,
class_mode='sparse',
shuffle=True,
subset='training') # Marks this as the training data
rn_val_datagen = rn_train_idg.flow_from_directory(
train_path,
target_size=target_size,
batch_size=batch_size,
class_mode='sparse',
shuffle=False,
subset='validation') # Marks this as the validation data
"""
Test set
"""
rn_test_idg = keras.preprocessing.image.ImageDataGenerator(preprocessing_function=keras.applications.resnet50.preprocess_input)
rn_test_datagen = rn_test_idg.flow_from_directory(
test_path,
target_size=target_size,
batch_size=batch_size,
class_mode='sparse',
shuffle=False)
# + [markdown] id="AICO664i68Ss"
# ### 3.2 Build model
# + id="iKFCDlBUJUQY"
"""
Function to build the ResNet50 model
"""
def build_transfer_learning_model():
# The ResNet50 is used as a base model
base_model = keras.applications.resnet.ResNet50(include_top=False,
weights='imagenet',
input_shape=(target_w, target_h, target_dims))
# We need our BatchNormalization layers to be trainable, the rest should be frozen
for layer in base_model.layers:
if isinstance(layer, keras.layers.BatchNormalization):
layer.trainable = True
else:
layer.trainable = False
# Assemble our final model
model = keras.models.Sequential([
base_model,
keras.layers.GlobalAveragePooling2D(),
keras.layers.Dropout(0.5),
keras.layers.Dense(8, activation="softmax")
])
model.summary()
return model
# + id="rxAveAwA5Cx_" colab={"base_uri": "https://localhost:8080/"} outputId="f3933dd0-c9b9-4da2-bec8-e6bb64b2c23d"
"""
Build and compile our ResNet50 model
This model consists of frozen layers where only the BatchNormalization and output Dense layers are trainable.
We will unfreeze and fine-tune the entire model later after the first fitting.
"""
rn_model = build_transfer_learning_model()
rn_model.compile(
optimizer=keras.optimizers.Adam(0.0001),
loss='sparse_categorical_crossentropy',
metrics=[ 'accuracy' ]
)
# + [markdown] id="yxdEj7QR69vS"
# ### 3.3 Train model
# + id="lWceWlIx5FLk" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="be9639c8-e4c4-40c4-e6d2-02ccd87597f2"
"""
Start training our ResNet50 model
This will also save the trained model file to our 'models' directory.
"""
rn_history = rn_model.fit(rn_train_datagen,
epochs=30,
verbose=1,
validation_data=rn_val_datagen)
rn_model.save('models/cnn_resnet_tl.hdf5')
if RunningInCOLAB:
files.download('models/cnn_resnet_tl.hdf5')
# + id="_c6R6N1l4Gef" colab={"base_uri": "https://localhost:8080/", "height": 549} outputId="9673ad2f-ce78-4b31-d7ca-c85b5db2cffa"
"""
Examine the training process
"""
plot_history(rn_history)
# + [markdown] id="awTNrEYR6_NO"
# ### 3.4 Evaluate model
# + id="lfDf5H2j5LNi" colab={"base_uri": "https://localhost:8080/"} outputId="4097ce75-cffb-41bc-faff-1017324efb7d"
"""
Use the model to predict on the unseen test set.
"""
rn_predictions = rn_model.predict(rn_test_datagen, verbose=1)
# + id="pSKz_A1nbQHU" colab={"base_uri": "https://localhost:8080/", "height": 808} outputId="696279f0-f9b9-4d7e-b0f2-5ab0e9e07cfd"
"""
Evaluate the prediction results.
Displays a Confusion Matrix and a Classification Report.
These are also saved to the 'results' folder.
"""
unique_labels = list(rn_test_datagen.class_indices.keys())
y_pred = np.argmax(rn_predictions, axis=-1)
y_true = rn_test_datagen.classes[rn_test_datagen.index_array]
display_confusion_matrix(
y_pred=y_pred,
y_true=y_true,
target_names=unique_labels,
save_path='results/cnn_resnet_tl/cnn_resnet_tl_confusion_matrix.png'
)
display_classification_report(
y_pred=y_pred,
y_true=y_true,
target_names=unique_labels,
save_path='results/cnn_resnet_tl/cnn_resnet_tl_classification_report.csv'
)
# + [markdown] id="5-oE9A4s7AcZ"
# ### 3.5 Unfreeze and re-train model
# + id="BuWToEG4wdVa" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="917c6451-3d05-475d-a4f4-c5a9d526361c"
"""
Unfreeze all layers
"""
rn_model.trainable = True
"""
Compile model with a lower learning rate
"""
rn_model.compile(
optimizer=keras.optimizers.Adam(0.000001), # Low learning rate
loss='sparse_categorical_crossentropy',
metrics=[ 'accuracy' ],
)
"""
Resume training
"""
rn_history_ft = rn_model.fit(rn_train_datagen,
epochs=30,
verbose=1,
validation_data=rn_val_datagen)
rn_model.save('models/cnn_resnet_tl_tuned.hdf5')
if RunningInCOLAB:
files.download('models/cnn_resnet_tl_tuned.hdf5')
# + id="Uj94xyxfK1yA" colab={"base_uri": "https://localhost:8080/", "height": 587} outputId="037f256e-7b27-4c14-a623-fe144ea51d09"
"""
Examine the training process
"""
plot_history(rn_history_ft)
# + [markdown] id="5de5L5nS7Dux"
# ### 3.6 Final model evaluation
# + id="PW-0jqW-xGNn" colab={"base_uri": "https://localhost:8080/"} outputId="27b3eb7a-9da1-447c-c596-ab0a4f19f41e"
"""
Use the model to predict on the unseen test set.
"""
rn_predictions = rn_model.predict(rn_test_datagen, verbose=1)
# + id="-7Rt5T0m_P15" colab={"base_uri": "https://localhost:8080/", "height": 808} outputId="e2c107b5-0519-4035-f932-f6f2557dae15"
"""
Evaluate the prediction results.
Displays a Confusion Matrix and a Classification Report.
These are also saved to the 'results' folder.
"""
unique_labels = list(rn_test_datagen.class_indices.keys())
y_pred = np.argmax(rn_predictions, axis=-1)
y_true = rn_test_datagen.classes[rn_test_datagen.index_array]
display_confusion_matrix(
y_pred=y_pred,
y_true=y_true,
target_names=unique_labels,
save_path='results/cnn_resnet_tl_tuned/cnn_resnet_tl_tuned_confusion_matrix.png'
)
display_classification_report(
y_pred=y_pred,
y_true=y_true,
target_names=unique_labels,
save_path='results/cnn_resnet_tl_tuned/cnn_resnet_tl_tuned_classification_report.csv'
)
|
project_notebook_cnn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
data = pd.read_csv('Company_Data.csv')
data.head()
data
data.columns
data.describe
df = pd.DataFrame(data)
df
type(data)
# create dummies for Shelveloc
dummies = pd.get_dummies(df['ShelveLoc']).rename(columns=lambda x: 'ShelveLoc_' + str(x))
# bring the dummies back into the original dataset
df = pd.concat([df, dummies], axis=1)
print(df)
df.head()
df = df.drop('ShelveLoc', 1)
df.head()
# create dummies for pitching team, batting team, pitcher id, batter id
dummies = pd.get_dummies(df['Urban']).rename(columns=lambda x: 'Urban_' + str(x))
# bring the dummies back into the original dataset
df = pd.concat([df, dummies], axis=1)
print(df)
# create dummies for US
dummies = pd.get_dummies(df['US']).rename(columns=lambda x: 'US_' + str(x))
# bring the dummies back into the original dataset
df = pd.concat([df, dummies], axis=1)
print(df)
df = df.drop('Urban', 1)
df = df.drop('US', 1)
df.head()
row_indexes=df[df['Sales']>=10].index
df.loc[row_indexes,'sales']="yes"
row_indexes=df[df['Sales']<10].index
df.loc[row_indexes,'sales']="no"
df.head()
df = df.drop('Sales', 1)
df.head()
colnames = list(df.columns)
colnames
predictors = colnames[:14]
target = colnames[14]
from sklearn import preprocessing
target
import numpy as np
from sklearn.model_selection import train_test_split
train,test = train_test_split(df,test_size = 0.2)
from sklearn.tree import DecisionTreeClassifier
help(DecisionTreeClassifier)
model = DecisionTreeClassifier(criterion = 'entropy')
model.fit(train[predictors],train[target])
preds = model.predict(test[predictors]) # predicting on test data set
pd.Series(preds).value_counts() # getting the count of each category
pd.crosstab(test[target],preds)
np.mean(preds==test.sales)
|
DecisionTree_CompanyData.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import cv2 as cv
import matplotlib.pyplot as plt
img = np.zeros((512,512,3), np.uint8)
# Draw a diagonal blue line with thickness of 5 px
cv.line(img,(0,0),(511,511),(255,0,0),5)
plt.imshow(img)
plt.show()
cv.rectangle(img,(384,0),(510,128),(0,255,0),3)
plt.imshow(img)
plt.show()
cv.circle(img,(447,63), 63, (0,0,255), -1)
plt.imshow(img)
plt.show()
cv.ellipse(img,(256,256),(100,50),0,0,180,255,-1)
plt.imshow(img)
plt.show()
pts = np.array([[10,5],[20,30],[70,20],[50,10]], np.int32)
pts = pts.reshape((-1,1,2))
cv.polylines(img,[pts],True,(0,255,255))
plt.imshow(img)
plt.show()
font = cv.FONT_HERSHEY_SIMPLEX
cv.putText(img,'OpenCV',(10,500), font, 4,(255,255,255),2,cv.LINE_AA)
plt.imshow(img)
plt.show()
|
opencv/Gui Features in OpenCV/Drawing Functions in OpenCV.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Check which Lipids are ACM dependent
# - Compare lipids found in Supernatant and Cells depending whether ACM as used or not
# ### Included libraries
import pandas as pd
from scipy.stats import ttest_ind
import numpy as np
from statsmodels.stats.multitest import multipletests
from matplotlib import cm
from matplotlib import pylab as plt
from matplotlib.lines import Line2D
import seaborn as sns
# ### Functions and definitions
# +
#define classes of lipids e.g. PC = Phosphatidylcholines
types_of_Lipids = ['CE','Cer','DAG','LPC','LPE','PC','PE','PI','PS','SM','TAG']
#colormap (20 unique colors)
cmap = cm.get_cmap('tab20')
#assign for each class of lipid a unique color
lipid_color = {}
for i,l in enumerate(types_of_Lipids):
lipid_color[l] = cmap(i)
# -
# ### Main Code
#Load the actual lipid results
LipidData = pd.read_excel('../data/Report_MEC025_LIPID01_jb.xlsx' ,header=2, na_values='<LOD')
#extract the lipids
columns = LipidData.columns
Lipids = columns[7:]
print (Lipids)
# +
#make analysis both for the cells and the supernatant experiment (where lipids are found)
specimens = ['cells','supernatant']
#make both analysis
for spec in specimens:
#extract WT control and WT with addition of ACM
data = LipidData.loc[(LipidData['Specimen'] == spec) & ((LipidData['Experiment'] == 'WT_C') | (LipidData['Experiment'] == 'WT_ACM'))]
#remove entries that have no values
data = data.dropna(axis=1,how='all')
#remaining lipids contains all valid columns (=lipids)
remaining_Lipids = data.columns.values[7:]
# The next 4 variables contain the (statistical )results for the individiual lipids (comparisan WT control and ACM)
# foldchanges = foldchanges
# significance = student_ttest
# calculated_lipids = name of lipid
# color = color for lipidclass (see before)
####
foldchanges = []
significance = []
calculated_lipids = []
color = []
# go through all valid lipids
for Lipid in remaining_Lipids:
# extract results for control and ACM
WT_C_values = data.loc[data['Experiment'] == 'WT_C'][Lipid]
WT_ACM_values = data.loc[data['Experiment'] == 'WT_ACM'][Lipid]
#drop rows with no values
WT_C_values = WT_C_values.dropna()
WT_ACM_values = WT_ACM_values.dropna()
#only calculate statistics if valid results found for both groups
if len(WT_C_values) > 0 and len(WT_ACM_values) > 0:
#calculate the statistics (see before)
significance.append(ttest_ind(WT_C_values,WT_ACM_values)[1])
foldchanges.append(WT_ACM_values.mean()/WT_C_values.mean())
calculated_lipids.append(Lipid)
color.append(lipid_color[Lipid.split(' ')[0]])
# correct pValues according to <NAME> (FDR)
pValues_Corrected = multipletests(significance,alpha=0.05,method='fdr_bh')[1]
pValues_Corrected = [-np.log10(p) for p in pValues_Corrected]
#transform the foldchanges to log2
foldchanges = [np.log2(f) for f in foldchanges]
####
# START MAKING PLOT ACM DEPENDENCY WT
# Scatter plot showing for each individual lipid the foldchange (x-axis) and pvalue (y-axis)
####
# Make result plot for differences upon ACM treatment (for wildtype)
plt.title('WT_ACM / WT_C - ' + spec)
plt.scatter(foldchanges,pValues_Corrected,c=color, alpha = 0.4)
plt.axhline(-np.log10(0.05), color= 'grey', ls='--')
plt.xlabel('log2[Fold Change ACM/Control]')
plt.ylabel('-log[PValue]')
# Make a legend showing the colors to lipid classes
legend_elements = []
for key in lipid_color:
legend_elements.append(Line2D([0], [0], marker='o', color='w', label=key,
markerfacecolor=lipid_color[key], markersize=10))
# Make actual plot
plt.legend(handles=legend_elements, loc='upper right',prop={'size': 5})
plt.savefig('../results/Lipid_ACM_Dependency/ACM_Dependency_'+spec+'.pdf')
plt.close()
####
# END PLOT ACM DEPENDENCY WT
####
###
# MAKE PLOT ALL LIPIDS FOLDCHANGE
####
# sort the foldchanges for the individual lipids (from largest to smalest)
foldchanges_sort, calculated_lipids_sort = zip(*sorted(zip(foldchanges, calculated_lipids),reverse=True))
foldchanges_sort, colors_sort = zip(*sorted(zip(foldchanges, color),reverse=True))
foldchanges_sort = [[x] for x in foldchanges_sort]
# Make plot showing the individual lipid foldchanges (largest top, smallest bottom)
# No error bar (as it consists already of too many individual bars)
plt.title('WT_ACM / WT_C - ' + spec)
sns.barplot(data=foldchanges_sort,orient='h',palette=colors_sort,linewidth=None)
plt.legend(handles=legend_elements, loc='right',prop={'size': 5})
plt.xlabel('log2[Fold Change ACM/Control]')
plt.yticks(range(0,len(calculated_lipids_sort)),calculated_lipids_sort,rotation=0,size=1.8)
plt.savefig('../results/Lipid_ACM_Dependency/ACM_Dependency_Barplot_AllLipids_'+spec+'.pdf')
plt.close()
###
# END PLOT ALL LIPIDS FOLDCHANGE
####
###
# MAKE PLOT LIPID GROUPs FOLDCHANGE
####
# Create dictionary for the individual lipid groups
Lipid_Group_Results = {}
for l in types_of_Lipids:
Lipid_Group_Results[l] = []
#add the individual lipids to the previosuly defined dictionary
for fc,lipid in zip(foldchanges,calculated_lipids):
Lipid_Group_Results[lipid.split(' ')[0]].append(fc)
# result lists that will contain the results for the whole lipid grups (same as before)
Lipid_Groups = []
Foldchanges_Groups = []
Foldchanges_Groups_Means = []
Colors_Groups = []
#go through all lipid groups
for key in Lipid_Group_Results:
#calculate only results if the lipid groups contains 1 or more lipids
if len(Lipid_Group_Results[key]) != 0:
Lipid_Groups.append(key)
Foldchanges_Groups.append(Lipid_Group_Results[key])
Foldchanges_Groups_Means.append(np.mean(Lipid_Group_Results[key]))
Colors_Groups.append(lipid_color[key])
#sort the foldchanges again from largest to smallest
_, Foldchanges_Groups = zip(*sorted(zip(Foldchanges_Groups_Means, Foldchanges_Groups),reverse=True))
_, Colors_Groups = zip(*sorted(zip(Foldchanges_Groups_Means, Colors_Groups),reverse=True))
_, Lipid_Groups = zip(*sorted(zip(Foldchanges_Groups_Means, Lipid_Groups),reverse=True))
#make bar plot with SEM as error bar
plt.title('WT_ACM / WT_C - ' + spec)
sns.barplot(data=Foldchanges_Groups,orient='h',palette=Colors_Groups, ci=68, capsize=0.3, errwidth=0.7,errcolor='black', linewidth=1, edgecolor='black')
plt.legend(handles=legend_elements, loc='right',prop={'size': 5})
plt.xlabel('log2[Fold Change ACM/Control] (+/- SEM)')
plt.yticks(range(0,len(Lipid_Groups)),Lipid_Groups,rotation=0,size=10)
plt.savefig('../results/Lipid_ACM_Dependency/ACM_Dependency_Barplot_LipidGroups_'+spec+'.pdf')
plt.close()
###
# END PLOT LIPID GROUPs FOLDCHANGE
####
|
code/ACM_Dependency.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Which GPU to use
# +
multiGPU = False
whichGPU = 0
# whichGPU = 1
# Select which GPU to use
if(multiGPU):
from keras.utils.training_utils import multi_gpu_model
else:
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
# The GPU id to use, usually either "0" or "1"
os.environ["CUDA_VISIBLE_DEVICES"] = str(whichGPU)
# # Do other imports now...
# -
# # Load all the functions
# %run -i 'arena.py'
# # General Parameters
# +
import math
# What data to use
tableBase = '4PpKk'
convertStates = False
# Interactive (just in general if one is asked for confirmations, set to False if on autopilot over night f.x.)
askForConfirmation = False
# NN parameters
filters = [16,32,32,64,128,128,128]
filterShape = [2,2,2,2,2,2,2]
batch_size = 256
optimizer = 'Adadelta'
useBatchNorm = False
num_classes = 3
input_shape = (4,8,8)
### DON'T MODIFY BELOW ###
# Generate dataset variables
fileName = tableBase + '.hdf5'
dataSetName = tableBase + '_onlyLegal'
if not convertStates:
dataSetName = tableBase + '_onlyLegal_fullStates'
dataSetWdlName = tableBase + '_Wdl_onlyLegal_3Values'
# Number of Pieces
nPi = int(dataSetName[0])
nPa = nPi - 2
nWPa = math.ceil(nPa/2)
# -
# # Experiment 1
# Bengio methood 3n4 with freeze
# ### Exp 1 Paramters
# +
# %run -i 'arena.py'
# Parameters
sourceNet = '103' # trained on 3pc from scratch
# sourceNet = '107' # trained on 4pc from scratch
freeze = True
resSaveFile = '3n4freeze'
epochs = 10
averageOver = 10
expDescrBaseName = "Bengio 3n4 - freeze = {} - average over {} runs".format(str(freeze), averageOver)
saveEveryRun = True # save stuff in results dir
saveWeightsCheckpoints = False # save chkp in results dit
saveTensorboardLogs = True # save logs in ./logs dir
resID = '---NORESID---' # used when not saving data, but fitModel() still needs a resID
fractionOfDataToUse = 1
plotDuringTraining = False
loadWeights = False
askForConfirmation = False
saveDir = 'bengioResults'
resSaveFile = resSaveFile + '-{}runAverage'.format(averageOver)
resSaveFileFullPath = saveDir + '/' + str(resSaveFile) + '.pkl'
# -
# ### Create model and load data
# +
# prepare save file
if not os.path.exists(resSaveFileFullPath):
print("Save file doesn't exists, creating...\n")
save_obj(saveDir, resSaveFile, [])
else:
print("Save file exists...\n")
# load data
X_train, X_test, y_train, y_test = loadData()
# create model
model, nnStr = createModel()
layersCount = len(model.layers)
# load old results
results = load_obj(saveDir, resSaveFile)
# initialize variables wrt old results
startTrainingAtLayer = len(results)
print("\nStarting/restarting TL at {} transfered layers".format(startTrainingAtLayer))
# -
# ### Train
# +
# %run -i 'arena.py'
for copyFirstNLayers in range(startTrainingAtLayer, layersCount):
print('\n\n')
print('==========================================================================================')
print('= =')
print('= Currently transfering first {} layers, out of {} ='.format(copyFirstNLayers, layersCount - 1))
print('= =')
print('==========================================================================================')
print()
# check if we are at the flatten layer, and skip it if so
if copyFirstNLayers == layersCount - 1:
copyFirstNLayers += 1
# train and average results
accumulatedScore = 0
for a in range(averageOver):
# set experement description test
expDescr = expDescrBaseName + '__copyLayers_{}__average_{}_of_{}'.format(copyFirstNLayers, a+1, averageOver)
# save current averagePosition to tmp file
with open(saveDir + '/' + str(resSaveFile) + '_currentPosition.txt','w') as file:
if copyFirstNLayers == layersCount:
location = copyFirstNLayers - 1
else:
location = copyFirstNLayers
file.write('Layers Transfered: {} out of {} \nInner avg loop position: {} out of {}'.format(location, layersCount-1, a+1, averageOver))
# load Model layers
model = loadNFirstLayers(model, sourceNet, copyFirstNLayers , freeze)
# Prepare save dir
if saveEveryRun:
resID = genNextResultsDir(model)
# train
fitHistory, logDir = trainModel(resID, model, saveWeightsCheckpoints, saveTensorboardLogs)
# score and save results
score = calcScore(model)
if saveEveryRun:
saveTrainResults(resID, model, logDir, score, copyFirstNLayers)
# update Return
accumulatedScore += score[1]
# append averaged results for one set of layers
results.append(accumulatedScore/averageOver)
# save old results to checkpoints dir
dateTime = time.strftime('%Y-%m-%d-%H:%M:%S', time.localtime())
src = saveDir + '/' + str(resSaveFile) + '.txt'
dest = saveDir + '/checkpoints/' + str(resSaveFile) + dateTime + '.txt'
if os.path.exists(src):
shutil.move(src, dest)
# save results
save_obj(saveDir, resSaveFile, results)
with open(saveDir + '/' + str(resSaveFile) + '.txt','w') as file:
file.write(str(results))
# to load:
# results = load_obj('temp','3n4.txt')
print('\n Final Results: {}'.format(results))
# -
# # Experiment 2
# Bengio methood 4n4 with freeze
time.sleep(60*60*4)
# ### Exp 2 Paramters
# +
# %run -i 'arena.py'
# Parameters
# sourceNet = '103' # trained on 3pc from scratch
sourceNet = '107' # trained on 4pc from scratch
freeze = True
resSaveFile = '4n4freeze'
epochs = 10
averageOver = 1
expDescrBaseName = "Bengio 4n4 - freeze = {} - average over {} runs".format(str(freeze), averageOver)
saveEveryRun = True # save stuff in results dir
saveWeightsCheckpoints = False # save chkp in results dit
saveTensorboardLogs = True # save logs in ./logs dir
resID = '---NORESID---' # used when not saving data, but fitModel() still needs a resID
fractionOfDataToUse = 1
plotDuringTraining = False
loadWeights = False
askForConfirmation = False
saveDir = 'bengioResults'
resSaveFile = resSaveFile + '-{}runAverage'.format(averageOver)
resSaveFileFullPath = saveDir + '/' + str(resSaveFile) + '.pkl'
# -
# ### Create model and load data
# +
# prepare save file
if not os.path.exists(resSaveFileFullPath):
print("Save file doesn't exists, creating...\n")
save_obj(saveDir, resSaveFile, [])
else:
print("Save file exists...\n")
# load data
X_train, X_test, y_train, y_test = loadData()
# create model
model, nnStr = createModel()
layersCount = len(model.layers)
# load old results
results = load_obj(saveDir, resSaveFile)
# initialize variables wrt old results
startTrainingAtLayer = len(results)
print("\nStarting/restarting TL at {} transfered layers".format(startTrainingAtLayer))
# -
# ### Train
# +
# %run -i 'arena.py'
for copyFirstNLayers in range(startTrainingAtLayer, layersCount):
print('\n\n')
print('==========================================================================================')
print('= =')
print('= Currently transfering first {} layers, out of {} ='.format(copyFirstNLayers, layersCount - 1))
print('= =')
print('==========================================================================================')
print()
# check if we are at the flatten layer, and skip it if so
if copyFirstNLayers == layersCount - 1:
copyFirstNLayers += 1
# train and average results
accumulatedScore = 0
for a in range(averageOver):
# set experement description test
expDescr = expDescrBaseName + '__copyLayers_{}__average_{}_of_{}'.format(copyFirstNLayers, a+1, averageOver)
# save current averagePosition to tmp file
with open(saveDir + '/' + str(resSaveFile) + '_currentPosition.txt','w') as file:
if copyFirstNLayers == layersCount:
location = copyFirstNLayers - 1
else:
location = copyFirstNLayers
file.write('Layers Transfered: {} out of {} \nInner avg loop position: {} out of {}'.format(location, layersCount-1, a+1, averageOver))
# load Model layers
model = loadNFirstLayers(model, sourceNet, copyFirstNLayers , freeze)
# Prepare save dir
if saveEveryRun:
resID = genNextResultsDir(model)
# train
fitHistory, logDir = trainModel(resID, model, saveWeightsCheckpoints, saveTensorboardLogs)
# score and save results
score = calcScore(model)
if saveEveryRun:
saveTrainResults(resID, model, logDir, score, copyFirstNLayers)
# update Return
accumulatedScore += score[1]
# append averaged results for one set of layers
results.append(accumulatedScore/averageOver)
# save old results to checkpoints dir
dateTime = time.strftime('%Y-%m-%d-%H:%M:%S', time.localtime())
src = saveDir + '/' + str(resSaveFile) + '.txt'
dest = saveDir + '/checkpoints/' + str(resSaveFile) + dateTime + '.txt'
if os.path.exists(src):
shutil.move(src, dest)
# save results
save_obj(saveDir, resSaveFile, results)
with open(saveDir + '/' + str(resSaveFile) + '.txt','w') as file:
file.write(str(results))
# to load:
# results = load_obj('temp','3n4.txt')
print('\n Final Results: {}'.format(results))
# -
# # Experiment 3
# Expansion learning
|
mainCode/1.old/TL-GPU1.old.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
import pyomo.environ as pyo
# ## Read the data from csv using pandas
csv_file_path = "./delivery_costs.csv"
df = pd.read_csv(csv_file_path, index_col=0)
df
N = list(df.index.map(str))
M = list(df.columns.map(str))
d = {(r, c):df.loc[r,c] for r in N for c in M}
P = 2
# ## Create the mode
# +
model = pyo.ConcreteModel(name = "(WL)")
model.x = pyo.Var(N, M, bounds=(0, 1))
model.y = pyo.Var(N, within=pyo.Binary)
def obj_rule(model):
return sum(d[n,m]*model.x[n,m] for n in N for m in M)
model.obj = pyo.Objective(rule=obj_rule)
def one_per_cust_rule(model, m):
return sum(model.x[n,m] for n in N) == 1
model.one_per_cust = pyo.Constraint(M, rule=one_per_cust_rule)
def warehouse_active_rule(model, n, m):
return model.x[n,m] <= model.y[n]
model.warehouse_active = pyo.Constraint(N, M, rule=warehouse_active_rule)
def num_warehouses_rule(model):
return sum(model.y[n] for n in N) <= P
model.num_warehouses = pyo.Constraint(rule=num_warehouses_rule)
# -
# ## Solve
solver = pyo.SolverFactory('glpk')
res = solver.solve(model)
# +
# model.pprint()
# -
# produce nicely formatted output
for wl in N:
if pyo.value(model.y[wl]) > 0.5:
customers = [str(cl) for cl in M if pyo.value(model.x[wl, cl] > 0.5)]
print(f"{wl} + serves customers: {customers}")
else:
print(str(wl)+": do not build")
|
wl_csv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# reload packages
# %load_ext autoreload
# %autoreload 2
# ### Choose GPU (this may not be needed on your computer)
# %env CUDA_DEVICE_ORDER=PCI_BUS_ID
# %env CUDA_VISIBLE_DEVICES=1
# ### load packages
from tfumap.umap import tfUMAP
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tqdm.autonotebook import tqdm
import umap
import pandas as pd
# ### Load dataset
from tensorflow.keras.datasets import mnist
# +
# load dataset
(train_images, Y_train), (test_images, Y_test) = mnist.load_data()
X_train = (train_images/255.).astype('float32')
X_test = (test_images/255.).astype('float32')
X_train = X_train.reshape((len(X_train), np.product(np.shape(X_train)[1:])))
X_test = X_test.reshape((len(X_test), np.product(np.shape(X_test)[1:])))
# subset a validation set
n_valid = 10000
X_valid = X_train[-n_valid:]
Y_valid = Y_train[-n_valid:]
X_train = X_train[:-n_valid]
Y_train = Y_train[:-n_valid]
# flatten X
X_train_flat = X_train.reshape((len(X_train), np.product(np.shape(X_train)[1:])))
X_test_flat = X_test.reshape((len(X_test), np.product(np.shape(X_test)[1:])))
X_valid_flat= X_valid.reshape((len(X_valid), np.product(np.shape(X_valid)[1:])))
print(len(X_train), len(X_valid), len(X_test))
# -
from tfumap.paths import ensure_dir, MODEL_DIR
# +
output_dir = MODEL_DIR/'projections'/ 'mnist' / 'direct'
ensure_dir(output_dir)
loss_df_direct = pd.read_pickle(output_dir / 'loss_df.pickle')
loss_df_direct
plt.plot(loss_df_direct.wall_time - loss_df_direct.wall_time[0], loss_df_direct.val)
# -
output_dir = MODEL_DIR/'projections'/ 'mnist' / 'network'
ensure_dir(output_dir)
loss_df = pd.read_pickle(output_dir / 'loss_df.pickle')
loss_df
plt.plot(loss_df.wall_time - loss_df.wall_time[0], loss_df.val)
# #### get umap learn
from umap import UMAP
embedder = UMAP(verbose=True)
z_umap = embedder.fit_transform(X_train_flat)
from datetime import datetime
start_time = datetime.strptime('13:00:31', '%H:%M:%S')
end_time = datetime.strptime('13:01:14', '%H:%M:%S')
umap_learn_training_time = (end_time - start_time).total_seconds()
umap_learn_training_time
# ### compute loss for UMAP learn
embedder.graph_
for each edge in umap graph
randomly sample 5 other edges
compute distances
from tfumap.umap import get_graph_elements
graph, epochs_per_sample, head, tail, weight, n_vertices = get_graph_elements(
embedder.graph_, embedder.n_epochs
)
max_sample_repeats_per_epoch = 25
batch_size = 1000
def batch_epoch_edges(edges_to, edges_from):
""" permutes and batches edges for epoch
"""
# compute the number of batches in one epoch
n_batches = int(len(edges_to) / batch_size)
# permute list of edges
permutation_mask = np.random.permutation(len(edges_to))[
: n_batches * batch_size
]
to_all = tf.reshape(
tf.gather(edges_to, permutation_mask), (n_batches, batch_size)
)
from_all = tf.reshape(
tf.gather(edges_from, permutation_mask), (n_batches, batch_size)
)
# return a tensorflow dataset of one epoch's worth of batches
return tf.data.Dataset.from_tensor_slices((to_all, from_all))
def create_edge_iterator(head, tail, weight):
""" create an iterator for edges
"""
# set the maximum number of times each edge should be repeated per epoch
epochs_per_sample = np.clip(
(weight / np.max(weight)) * max_sample_repeats_per_epoch,
1,
max_sample_repeats_per_epoch,
).astype("int")
edges_to_exp, edges_from_exp = (
np.array([np.repeat(head, epochs_per_sample.astype("int"))]),
np.array([np.repeat(tail, epochs_per_sample.astype("int"))]),
)
edge_iter = tf.data.Dataset.from_tensor_slices((edges_to_exp, edges_from_exp))
edge_iter = edge_iter.repeat()
edge_iter = edge_iter.map(batch_epoch_edges)
edge_iter = edge_iter.prefetch(buffer_size=10)
return iter(edge_iter), np.shape(edges_to_exp)[1]
# create iterator for data/edges
edge_iter, n_edges_per_epoch = create_edge_iterator(head, tail, weight)
from tfumap.umap import convert_distance_to_probability, compute_cross_entropy
def compute_umap_loss(batch_to, batch_from):
"""
compute the cross entropy loss for learning embeddings
Parameters
----------
batch_to : tf.int or tf.float32
Either X or the index locations of the embeddings for verticies (to)
batch_from : tf.int or tf.float32
Either X or the index locations of the embeddings for verticies (from)
Returns
-------
ce_loss : tf.float
cross entropy loss for UMAP
embedding_to : tf.float
embeddings for verticies (to)
embedding_from : tf.float
embeddings for verticies (from)
"""
# get the embeddings
embedding_to = tf.gather(z_umap, batch_to)
embedding_from = tf.gather(z_umap, batch_from)
# get negative samples
embedding_neg_to = tf.repeat(embedding_to, embedder.negative_sample_rate, axis=0)
repeat_neg = tf.repeat(embedding_from, embedder.negative_sample_rate, axis=0)
embedding_neg_from = tf.gather(
repeat_neg, tf.random.shuffle(tf.range(tf.shape(repeat_neg)[0]))
)
# distances between samples
distance_embedding = tf.concat(
[
tf.norm(embedding_to - embedding_from, axis=1),
tf.norm(embedding_neg_to - embedding_neg_from, axis=1),
],
axis=0,
)
# convert probabilities to distances
probabilities_distance = convert_distance_to_probability(
distance_embedding, embedder._a, embedder._b
)
# treat positive samples as p=1, and negative samples as p=0
probabilities_graph = tf.concat(
[tf.ones(embedding_to.shape[0]), tf.zeros(embedding_neg_to.shape[0])],
axis=0,
)
# cross entropy loss
(attraction_loss, repellant_loss, ce_loss) = compute_cross_entropy(
probabilities_graph,
probabilities_distance,
repulsion_strength=embedder.repulsion_strength,
)
return ce_loss, embedding_to, embedding_from
edge_epoch = next((iter(edge_iter)))
ce_loss_list = []
for batch_to, batch_from in tqdm(edge_epoch):
(ce_loss, _, _) = compute_umap_loss(
batch_to, batch_from
)
ce_loss_list.append(np.mean(ce_loss.numpy()))
umap_learn_loss = np.mean(ce_loss_list)
umap_learn_loss
time_d_mask
# +
import seaborn as sns
pal20c = sns.color_palette('tab20c', 20)
sns.palplot(pal20c)
hue_dict = {
"umap-learn": list(np.array(pal20c[3])**4),
"direct": pal20c[2],
"AE": pal20c[1],
"autoencoder": pal20c[1],
"network": pal20c[0],
'vae': pal20c[8],
'ae_only': pal20c[9],
"pca": pal20c[12],
"PCA": pal20c[12],
"TSNE": pal20c[4],
"parametric-tsne":pal20c[5],
"network-cpu": pal20c[2],
}
sns.palplot([hue_dict[i] for i in hue_dict.keys()])
# -
from tfumap.paths import FIGURE_DIR, save_fig
# +
max_time = 1000#120
fig, ax = plt.subplots(figsize=(4,2), dpi=300)
time_d = loss_df.wall_time.values - loss_df.wall_time[0]
time_d_mask = time_d<max_time
val_d = loss_df.val.values[time_d_mask]
time_d = time_d[time_d_mask]
ax.plot(time_d,val_d, lw = 3, label='P. UMAP', color=hue_dict['network'])
time_d = loss_df_direct.wall_time.values - loss_df_direct.wall_time[0]
time_d_mask = time_d<max_time
val_d = loss_df_direct.val.values[time_d_mask]
time_d = time_d[time_d_mask]
ax.plot(time_d,val_d, lw = 3, label='UMAP (TF)', color=hue_dict['direct'], ls='dashed')
ax.scatter([umap_learn_training_time], umap_learn_loss, color=hue_dict['umap-learn'], label='UMAP (UMAP-learn)')
ax.set_ylim([0, 0.5])
ax.legend()
ax.set_ylabel('Cross Entropy')
ax.set_xlabel('Time (S)')
ax.set_xscale('log')
#ax.set_yscale('log')
save_fig(FIGURE_DIR/'mnist-training-time', save_pdf=True, dpi=300)
# -
|
notebooks/dataset-projections/mnist/mnist-compare-embedding-time.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/poverty149/Speech-Emotion-Recognition/blob/main/speechemotionrecognitionrnn.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + papermill={"duration": 8.792978, "end_time": "2022-01-22T15:34:00.397970", "exception": false, "start_time": "2022-01-22T15:33:51.604992", "status": "completed"} tags=[] id="d5230fbd"
#Importing Libraries
import numpy as np
import pandas as pd
import os
import librosa
import wave
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.neural_network import MLPClassifier
from sklearn.metrics import accuracy_score
import keras
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential
from keras.layers import *
from tensorflow.keras.optimizers import RMSprop
# + colab={"base_uri": "https://localhost:8080/"} id="-RUfT6T9Az_0" outputId="333e9fc6-1b7e-4430-c709-c47cb9978cfd"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="gLQ2MFTwBCwJ" outputId="a5042a31-f704-4620-814b-aee3b81cae2e"
# cd drive/Mydrive
# + colab={"base_uri": "https://localhost:8080/"} id="brCWwygR_IDR" outputId="b235b668-d65b-4ee2-b5a8-569f588adfac"
# !unzip "archive.zip"
# + papermill={"duration": 0.021147, "end_time": "2022-01-22T15:34:00.433480", "exception": false, "start_time": "2022-01-22T15:34:00.412333", "status": "completed"} tags=[] id="972d6afd"
def extract_mfcc(wav_file_name):
#This function extracts mfcc features and obtain the mean of each dimension
y, sr = librosa.load(wav_file_name,duration=3
,offset=0.5)
#set n_mfcc = 40
mfccs = np.mean(librosa.feature.mfcc(y=y, sr=sr, n_mfcc=40).T,axis=0)
return mfccs
# + colab={"base_uri": "https://localhost:8080/"} id="9Rn5Z_IyB9R7" outputId="a4b48ad0-01f0-4707-f147-ab91545f1d18"
# !unzip -u "/content/drive/My Drive/archive.zip" -d "/content/NewFolder"
# + colab={"base_uri": "https://localhost:8080/"} id="dOdvFvRJCSL3" outputId="f58265bd-8262-4c70-9b71-56c085a0ba24"
# %cd /content
# + papermill={"duration": 615.031963, "end_time": "2022-01-22T15:44:15.477826", "exception": false, "start_time": "2022-01-22T15:34:00.445863", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="4cd41dff" outputId="a4eb8f96-96ea-4ec0-bb39-378414df655e"
##### load full radvess speech dataset #####
radvess_speech_labels = [] # to save extracted label/file
ravdess_speech_data = [] # to save extracted features/file
for dirname, _, filenames in os.walk('NewFolder/'):
for filename in filenames:
# print(os.path.join(dirname, filename))
radvess_speech_labels.append(int(filename[7:8]) - 1) # the index 7 and 8 of the file name represent the emotion label
wav_file_name = os.path.join(dirname, filename)
ravdess_speech_data.append(extract_mfcc(wav_file_name)) # extract MFCC features/file
print("Finish Loading the Dataset.")
# + papermill={"duration": 0.049962, "end_time": "2022-01-22T15:44:15.551538", "exception": false, "start_time": "2022-01-22T15:44:15.501576", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="8097b928" outputId="2e14a7c8-5d87-4757-9dd2-fd96cfc7b014"
#### convert data and label to array
ravdess_speech_data_array = np.asarray(ravdess_speech_data) # convert the input to an array
ravdess_speech_label_array = np.array(radvess_speech_labels)
ravdess_speech_label_array.shape # get tuple of array dimensions
#### make categorical labels
labels_categorical = to_categorical(ravdess_speech_label_array) # converts a class vector (integers) to binary class matrix
ravdess_speech_data_array.shape
labels_categorical.shape
print(ravdess_speech_label_array.shape, ravdess_speech_data_array.shape, labels_categorical.shape)
# + papermill={"duration": 0.024463, "end_time": "2022-01-22T15:44:15.596169", "exception": false, "start_time": "2022-01-22T15:44:15.571706", "status": "completed"} tags=[] id="f5e0147e"
x_train,x_test,y_train,y_test= train_test_split(np.array(ravdess_speech_data_array),labels_categorical, test_size=0.20, random_state=9)
# + papermill={"duration": 0.0216, "end_time": "2022-01-22T15:44:15.630828", "exception": false, "start_time": "2022-01-22T15:44:15.609228", "status": "completed"} tags=[] id="2841fe4a"
# Initialize the Multi Layer Perceptron Classifier
model=MLPClassifier(alpha=0.01, batch_size=22, epsilon=1e-08, hidden_layer_sizes=(150,), learning_rate='adaptive', max_iter=400)
# + papermill={"duration": 6.407834, "end_time": "2022-01-22T15:44:22.051992", "exception": false, "start_time": "2022-01-22T15:44:15.644158", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="7feb9346" outputId="aa3739d4-27a5-48da-bc0d-ac7a7d2dd49b"
# Train the model
model.fit(x_train,y_train)
# + papermill={"duration": 0.028507, "end_time": "2022-01-22T15:44:22.094638", "exception": false, "start_time": "2022-01-22T15:44:22.066131", "status": "completed"} tags=[] id="6135d89b"
# Predict for the test set
y_pred=model.predict(x_test)
# + papermill={"duration": 0.046747, "end_time": "2022-01-22T15:44:22.167105", "exception": false, "start_time": "2022-01-22T15:44:22.120358", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="be49f075" outputId="c504f7be-0bf8-44d0-f84d-c229f26a537a"
# Calculate the accuracy of our model
accuracy=accuracy_score(y_true=y_test, y_pred=y_pred)
# Print the accuracy
print("Accuracy: {:.2f}%".format(accuracy*100))
# + papermill={"duration": 0.028884, "end_time": "2022-01-22T15:44:22.223066", "exception": false, "start_time": "2022-01-22T15:44:22.194182", "status": "completed"} tags=[] id="ec9cea05"
# Split the training, validating, and testing sets
number_of_samples = ravdess_speech_data_array.shape[0]
training_samples = int(number_of_samples * 0.6)
validation_samples = int(number_of_samples * 0.4)
test_samples = int(number_of_samples * 0.01)
# + papermill={"duration": 0.02874, "end_time": "2022-01-22T15:44:22.266737", "exception": false, "start_time": "2022-01-22T15:44:22.237997", "status": "completed"} tags=[] id="173eecf8"
# Define the LSTM model
def create_model_LSTM():
model = Sequential()
model.add(LSTM(128, return_sequences=False, input_shape=(40, 1)))
model.add(Dense(64))
model.add(Dropout(0.3))
model.add(Activation('relu'))
model.add(Dense(32))
model.add(Dropout(0.3))
model.add(Activation('relu'))
model.add(Dense(8))
model.add(Activation('softmax'))
# Configures the model for training
model.compile(loss='categorical_crossentropy', optimizer='Adam', metrics=['accuracy'])
return model
# + papermill={"duration": 444.151265, "end_time": "2022-01-22T15:51:46.438435", "exception": false, "start_time": "2022-01-22T15:44:22.287170", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="a2ad6f9c" outputId="f5f4b492-17d2-449b-d840-5f571235aea6"
### train using LSTM model
##import os
os.environ['KMP_WARNINGS'] = 'off'
model_A = create_model_LSTM()
history = model_A.fit(np.expand_dims(x_train,-1), y_train, validation_data=(np.expand_dims(x_test, -1), y_test),
epochs=150, batch_size = 32)
# + papermill={"duration": 2.10379, "end_time": "2022-01-22T15:51:50.298753", "exception": false, "start_time": "2022-01-22T15:51:48.194963", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 295} id="5d98a71d" outputId="6bf9c1bb-3b26-4f27-d496-178767c0dc8f"
### loss plots using LSTM model
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'ro', label='Training Loss')
plt.plot(epochs, val_loss, 'b', label='Validation Loss')
plt.title('Training and Validation Loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + papermill={"duration": 1.977786, "end_time": "2022-01-22T15:51:54.047285", "exception": false, "start_time": "2022-01-22T15:51:52.069499", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 295} id="6ef1eb82" outputId="b211aa37-5cd6-43e6-b8d3-c5bb8d8d2376"
### accuracy plots using LSTM model
plt.clf()
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
plt.plot(epochs, acc, 'ro', label='Training Accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation Accuracy')
plt.title('Training and Validation Accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
# + papermill={"duration": 1.780464, "end_time": "2022-01-22T15:51:57.573965", "exception": false, "start_time": "2022-01-22T15:51:55.793501", "status": "completed"} tags=[] id="c01a00ce"
model_A.save_weights("Model_LSTM.h5")
# + papermill={"duration": 1.793427, "end_time": "2022-01-22T15:52:01.218158", "exception": false, "start_time": "2022-01-22T15:51:59.424731", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="839bfc65" outputId="8777e48c-c4d5-40dc-f45c-c809c096efaa"
np.expand_dims(x_train,-1).shape
|
speechemotionrecognitionrnn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div><img src='../../img/logo.png' width="30%">
# # Tutorial Part 1
# boxsimu is a simple modelling framework based on the principle of a system of discrete boxes interacting with each other. boxsimu offers a simple interface to define a system, its components and properties. The system definition is done by instantiating several classes of the boxsimu package like ```Fluid```, ```Variable```, ```Flow```, ```Flux```, ```Process```, ```Reaction```, and ```Box```. The temporal evolution of a system defined in boxsimu can then easily be simulated and the result of the simulation can be visualized or further investigated.
# After a short primer on what mathematical modelling is and how boxsimu fits in there, this tutorial will lead you through the installation of boxsimu. After the installion, it will briefly explain the basic structure of a system-definition in boxsimu. After that, you should have a basic understanding of the most important classes of boxsimu and you're ready to dive into a first example!
# ### Introduction
# #### Mathematical modelling
# In order to understand the dynamics of a system, it is often useful to create a (mathematical) model thereof. A system can be almost anything, e.g. a mountain lake, the Atlantic ocean, the atmosphere, a machine, a company, or even the entire Earth System. A (mathematical) model represents such a system while neglecting a big amount of complexity. Thus a (mathematical) model tries to explain the fundamental system behaviour as simply as possible.
# Schwarzenbach et al. (2003) uses the following defintion of a system:
# > ''A model is an imitation of reality that stresses those aspects that are assumed to be important and omits all properties considered to be nonessential''.
# Mathematical modelling of a system is a challenging and complex activity. Depending on the system an analytical solution can even be nonexistent. As a consequence one is forced to simulate/solve such systems numerically. This is where boxsimu comes into play:
# > boxsimu allows the user to easily define systems of simple to intermediate complexity and to simulate their temporal evolution. All that can be done on a high level of abstraction, meaning the user doesn't have to bother about the representation of the system with computer code.
# #### Limitations of boxsimu
# At the current status of development of boxsimu, the numerical solver is not very efficient. As a consequence large and complexe systems with a lot of variables, boxes, and processes/reactions will be solved slowly.
# ### Installation
# #### Dependencies
# **Python**:
# boxsimu was only tested with Python 3.6. Therefore boxsimu may not work as expected with older versions of Python.
# The following python packages are required by boxsimu:
#
# **Scientifc packages:**<br>
# - **numpy** (1.10 or newer): Fundamental package for scientifc computing, offers powerful and very fast matrix operations.
# - **matplotlib** (1.5 or newer): Plotting library with publication quality figures.
# - **jupyter** (4.0 or newer): Offers a browser-based IDE.
# - **pandas** (0.15 or newer): Allows spreadsheet-like calculation.
# - **pint** (0.8.1 or newer): Physical units-framework for python.
#
# **Other packages:**<br>
# - **attrdict** (1.2 or newer): Extension of the python dict class which allows to access dict items also as instance attributes.
# - **svgwrite** (1.1 or newer): Generation of SVG graphics.
# - **dill** (2.7 or newer): Extended pickeling of python objects.
# All of the above listed python package are available in the official python repository and thus can be installed using pip. Thus, if a package is missing on your system just use:
# ```bash
# pip install <package name>
# ```
# Additionally an up-to-date C compiler like gcc is needed, since some computational critical parts of boxsimu are written in Cython, which has to be compiled into binary code on your computer.
# #### Installation using pip
# boxsimu can easily be installed using pip. On your console type in
# ```bash
# pip install boxsimu
# ```
#
# this should automatically compile all Cython files and copy all source files into the right directory. If the installation fails check if all of the above mentioned dependencies are met.
# #### Installation from source
# Alternatively to the installation via pip, boxsimu can also be installed from source. For this, download the most recent source code from github {{URL}} or clone the repository. Afterwards open a system console and change into the directory where you downloaded boxsimu. Then execute the following commands:
# > \$ cd boxsimu <br>
# \$ python setup.py
# ## boxsimu Overview
# A system is defined by instantiating the class BoxModelSystem and passing instances of the classes Box, Flow, and Flux to it.
# The instance of the class BoxModelSystem contains all information on the whole system while its components (Boxes, Flows, and Fluxes) store information on the distribution of Fluids and Variables in the system and how the different compartements (boxes) exchange these quantities. The basic structure of a BoxModelSystem instance is graphically shown below:
# <img src='../../img/BoxModelSystemVisualization.png' width="60%">
# In this diagram an instance of BoxModelSystem is shown that contains two boxes: 'Box 1' and 'Box 2'. Both boxes contain the same fluid ('Fluid 1') and two instances of the class Variable ('A' and 'B'). Additionally both boxes can contain an arbitrary number of independet processes and reactions of these variables. Finally, the boxes exchange mass of the variables and the fluid via fluxes and flows (the difference between a flow and a flux will be explained further down).
# A BoxModelSystem can contain an arbitrary number of boxes, however, the more boxes there are the slower the system's simulation will progress. Similarly the number of fluxes, flows, and variables, processes, reactions within boxes is only limited in regard of computational power. In contrast, every box can only contain zero or one instance of the class Fluid.
# The most important class that a user of BoxSimu is interacting with are:
# - **BoxModelSystem**: Contains the whole system that a user wants to simulate and investigate. Most often this is the last class that is instantiated since instance of the other classes are needed as arguments for the BoxModelSystem-constructor. Once an instance of BoxModelSystem is built, its temporal evolution can be simulated using the instance's 'solve' method.
# - **Box**: Represents a compartement of the system. Contains information about the mass of variables and fluid within the represented compartement and which processes and reaction take place. Additionally for every box an 'environmental condition' (instance of the class Condition) can be defined. These conditions (e.g. the pH or the temperature within the box) in turn can then be used by user-defined rate functions (dynamic rates of processes/reactions/flows/fluxes/fluid-densities that can be dependent on the condition of the box).
# - **Flow**: Represent a transport of a fluid between two boxes. This flow can passively transport variables (e.g. a river that passively transport a lot of different chemical substances) but doesn't have to (e.g. evaporation from a lake into the atmosphere where almost pure water is transported).
# - **Flux**: Represents a transport of a variable that is not associated with a flow of a fluid. Examples of fluxes are for example: sedimentation of particles in a sea/ocean/atmosphere.
# - **Variable**: Variables are tracer within the system which time evolution is simulated. Thus variables are quantites within the system that are of direct or indirect interst to the user.
# - **Process**: Represents sink or source mechanisms of a certain variable.
# - **Reaction**: Represents transformations between different variables. There is no mass conservation constraint. Thus a ''transformation'' of approximately 1kg of phosphate into 114kg of phytoplankton (approx. Redfield ratio) is a valid reaction.
# A system is most easily defined following these steps:
# - Define all instances of the classes Variable and Fluid
# - Define all instances of the classes Reaction and Process
# - Define all instances of the class Box
# - Define all instances of the classes Flow and Flux
# - Define the instance of the class BoxModelSystem
# Given this first fundamental understanding of boxsimu classes and their sequence of appearence within a system-definition, we jump directly into a first example!
# ## A simple Lake 1-Box Model:
# ### Model description
# Our first system consists of a freshwater lake that only has one inflow and one outflow. We want to simulate how the concentration of phosphate in this lake evolves over time. In order to do that we assume the inflow to have a constant concentration of phosphate (PO4) while the outflow has the same concentration of PO4 as the (well-mixed) lake itself. The volume/mass of lake-water is constant over time.
#
# In the following we have a simple depiction of the lake system and important parameters thereof:
# <img src='../../img/1D-lake-model2.png' width="50%">
# ### Analytical solution
# Before we use boxsimu to simulate this system we can solve the govering equations of this system analytically in order to validate the output of boxsimu.
# So lets start with the defintion of all needed variables and their physical units:
#
# Variables (symbols are consistent with the figure above):
# - $V$ = Volume of the lake [m^3] ($V$ is constant)
# - $m$ = Mass of phosphate in the lake [kg]
# - $C$ = $\frac{m}{V}$ = Concentration of phosphate in the lake [kg/m^3]
# - $C_0$ = Concentration of phosphate in the lake at the begining of the simulation (t=0) [s]
# - $C_{in}$ = Concentration of phosphate in the inflow [kg/m^3]
# - $Q$ = Volumetric water flow rate of the Inflow and Outflow [m^3/s]
# - $t$ = Time [d]
#
# - $k_w$ = $\frac{Q}{V}$ = Specific flow rate [1/d]
#
# Assumptions:
# - $C_{in}$ = const.
# - $Q$ = const.
# - $V$ = const.
#
#
# Based on the definition of our system given above we can set up the following differential equation:
#
# $\frac{dm}{dt} = \sum sources - \sum sinks = Q \cdot C_{in} - Q \cdot C$.
#
# The rate of change of the phosphate concentration in the lake is given by the sum of all source terms minus the sum of all sink terms. In this case the only source term is the inflow of phosphate from the river upstream, while the only sink term is the outflow of phosphate through the river downstream.
# Where $Q \cdot C_{in}$ and $Q \cdot C$ represents the phosphate-gain and phosphate-loss of the lake per time, respectively.
#
# Next, we devide both sides by the volume of the lake ($V$) and use the specific flow rate $k_w$ on the right hand side (r.h.s.):
#
# $\frac{1}{V}\frac{dm}{dt} = k_w \cdot C_{in} - k_w \cdot C = k_w (C_{in} - C)$
#
# Now, since the volume of the box is constant, we can incorporate the volume into the time-derivative
# and end up with:
#
# $\frac{1}{V}\frac{dm}{dt} = \frac{d(m/V)}{dt} = \frac{dC}{dt} = k_w (C_{in} - C)$
#
# The solution of this linear, inhomogene, ordinary differential equation is:
#
# $C(t) = (C_0 - C_{in}) e^{-k_wt} + C_{in}$
#
# where we also used the initial condition $C(t=0) = C_0$.
#
# The solution is plotted below for a system with the following parameters:
# - $V = 10^7m^3$
# - $Q = 10^5\frac{m^3}{d}$
# - $C_0 = 10^{-2}\frac{kg}{m^3}$
# - $C_{in} = 3 \cdot 10^{-1}\frac{kg}{m^3}$
# We define a function that calculates and returns the concentration of phosphate at a time $t$. We also vectorize this function using numpy in order to be able to apply it on arrays. The resulting array is then plotted as a function of time:
# +
import matplotlib.pyplot as plt
import numpy as np
@np.vectorize
def C(t):
V = 1e7
Q = 1e5
C0 = 1e-2
Cin = 3e-1
kw = Q/V
return (C0-Cin)*np.exp(-kw*t) + Cin
t = np.linspace(0, 8e2, 1000)
c_phosphate = C(t)
plt.plot(t, c_phosphate)
plt.xlabel('Time [days]')
plt.ylabel('PO4 concentration [kg/m^3]')
# -
# We can see that the system reaches a steady-state after about $400$ days.
#
# Now we want to use boxsimu to simulate this system...
# ### boxsimu
# Since boxsimu accepts some quantities only in certain units (dimensionalities) we first have to calculate the system parameters in the right dimensionalities/units:
# - The amount of fluid inside a box has to be given in mass units: $m_{water} = V \cdot \rho = 10^7\,m^3 \cdot 10^3\,\frac{kg}{m^3} = 10^{10}kg$
# - The inital amount of phosphate also has to be given in mass units: $m(t=0) = V \cdot C_0 = 10^7\,m^3 \cdot 10^{-2}\,\frac{kg}{m^3} = 10^5\,kg$
# - The flow rate has to be given in units of mass per time: $J = Q \cdot \rho = 10^5\,\frac{m^3}{d} \cdot 10^3\,\frac{kg}{m^3} = 10^8\,\frac{kg}{d}$
# Now we define these parameters as python variables. But before we can do that we have to import boxsimu (if not already happened) and the instance of pint.UnitRegistry that is used by boxsimu (boxsimu.ur).
# > If you use boxsimu you have to use the UnitRegistry from boxsimu, since different UnitRegistry instances are incompatible. Thus use <br>
# ```python
# from boxsimu import ur
# ```
# instead of importing the UnitRegistry directly from the pint libary.
import boxsimu
from boxsimu import ur
# Now you are able to specify the units of a python variable by simply multiplying by pint units:
length = 3 * ur.meter
# Pint units can easily be transformed to other units of the same physical dimensionality:
length.to(ur.km) # transforms length to kilometer
# Now lets define the system parameters calculated above:
m_water = 1e10 * ur.kg
m_0 = 1e5 * ur.kg
flow_rate = 1e8 * ur.kg / ur.day
# Next we define the boxsimu system!
# In the following you see the complete model defintion in boxsimu:
# +
# FLUIDS
freshwater = boxsimu.Fluid('freshwater', rho=1000*ur.kg/ur.meter**3)
# VARIABLES
po4 = boxsimu.Variable('po4')
# PROCESSES
# No processes in this system
# REACTIONS
# No reactions in this system
# BOXES
lake = boxsimu.Box(
name='lake',
description='Little Lake',
fluid=freshwater.q(m_water),
variables=[po4.q(m_0)],
)
# FLOWS
inflow = boxsimu.Flow(
name='Inflow',
source_box=None,
target_box=lake,
rate=flow_rate,
tracer_transport=True,
concentrations={po4: 3e-1 * ur.gram / ur.kg},
)
outflow = boxsimu.Flow(
name='Outflow',
source_box=lake,
target_box=None,
rate=flow_rate,
tracer_transport=True,
)
# FLUXES
# No fluxes in this system
# BOXMODELSYSTEM
system = boxsimu.BoxModelSystem(
name='lake_system',
description='Simple Lake Box Model',
boxes=[lake,],
flows=[inflow, outflow,],
)
# -
# ### Code overview
# We will go through this code line by line.
# But before we dive into the details, have a quick look at the sequence of definitions of boxsimu classes:
#
# In a first step, instances of the classes **Fluid** and **Variable** are instantiated.
# A fluid represents the solvent of a box in which variables can be dissolved (e.g. the fluid of a lake is water, in which a myrade of chemicals can be dissolved).
# A box can, but doesn't have to be filled with a fluid. This allows to also simulate quantites that are not typically dissolved (e.g. to simulate a population of animals).
# Variables are quantities of the system that are of interest to the user and that are simulated. A variable defined for one box, flow, flux, process, or reaction will automatically be created for all boxes.
#
# At the time of instantiation (creation of the instance) instances of the classes Fluid and Variable are not quantified, that means the instances contain no information about the mass of fluid or variable they represent. However, before they are passed to an instance of the class Box they need to be quantified. This is done by calling the method ''q(mass)'' for a fluid or variable.
#
# In our simple lake system the fluids and variables are:
# - Fluid: The freshwater that is present in the lake (see line 2 in the code).
# - Variable: Phosphate whose concentration we want to analyse as a function of time (see line 5 in the code).
#
# Next, if needed, the classes **Process** and **Reaction** are instantiated (created). However, in this simple lake system there are no processes and no reactions.
#
# The created instances of Fluid, Variable, Process, and Reaction are then assigned to instances of the class **Box** that represent a compartement of the system (see lines 14-19 in the code).
#
# In a next step we have to define how the boxes in our system interact with each other and with the environement (outside the system). That means how the boxes are exchaning mass of fluids and variables. Therefore we instantiate the classes **Flow** and **Flux** (see lines 22-40 in the code).
# The difference between a flow and a flux is that flows represent a transport of fluid that again can (but doesn't have to) passively transport dissolved variables, whereas fluxes represent a transport of variables that is not associated with any fluid transport.
#
# Finally we create an instance of the class **BoxModelSystem** that glues together all elements of the system (see lines 43-46 in the following code).
# ### Code Line by Line
# Lets go through this code line by line:
# **Line 2**:
# ```python
# freshwater = boxsimu.Fluid('freshwater', rho=1000*ur.kg/ur.meter**3)
# ```
# An instance of the class ```Fluid```, representing the freshwater of the lake, is created.
# The class ```Fluid``` has the following signature:
# ```python
# Fluid(name, rho, description=None)
# ```
#
# - **name**: String which also has to be a valid python variable name since it is internally used to identify specific fluid instances. Thus, 'fluid123' or 'fluid_123' would be valid, while 'fluid 123' or '123fluid' would not be valid name arguments.
# - **rho**: Pint quantity or a user-defined function that returns a pint quantity. You find more information about user-defined functions in the second part of the tutorial.
# - **description**: String which gives a short description of the fluid.
#
# (For further information on the class ```Fluid``` confront the documentation: DOCUMENTATION_URL)
# **Line 5**:
# ```python
# po4 = boxsimu.Variable('po4')
# ```
#
# An instance of the class ```Variable``` is instantiated (created). This variable represents the substance we are interested in: phosphate. Every system needs to define at least one variable since its the temporal evolution of these variables that is simulated with the boxsimu-solver.
#
# The class ```Variable``` has the following signature:
# ```python
# Variable(name, molar_mass=None, mobility=True, description=None)
# ```
#
# - **name**: String which also has to be a valid python variable name since it is internally used to identify specific fluid instances. Thus, 'variable123' or 'variable_123' would be valid, while 'variable 123' or '123variable' would not be valid name arguments.
# - **molar_mass**: Pint quantity that specifies the molar mass of the substance that is represented by the variable instance. Must be of dimensionality [M/N] (e.g. mol/kg).
# - **mobility**: Boolean or function that returns boolean. Specifies if the variable is dissolved in a fluid (if one is present) and therefore it is transported by fluid flow.
# - **description**: String which gives a short description of the variable.
# **Line 14-19**:
# ```python
# lake = boxsimu.Box(
# name='lake',
# name_long='<NAME>',
# fluid=freshwater.q(m_water),
# variables=[po4.q(m_0)],
# )
# ```
#
# An instance of the class ```Box``` is instantiated (created). A box represents a compartement (region/part of the system which is somehow separated or at least distinguishable from the rest of the system.
#
# The class ```Box``` has the following signature:
# ```python
# Box(name, description, fluid=None, condition=None,
# variables=None, processes=None, reactions=None)
# ```
#
# - **name**: String which also has to be a valid python variable name since it is internally used to identify specific box instances. Thus, 'box123' or 'box_123' would be valid, while 'box 123' or '123box' would not be valid name arguments.
# - **description**: String which gives a short description of the box.
# - **fluid**: Instance of the class ```Fluid``` or None. Specifies the fluid that is present within a box. If None is given, the box is assumed to not contain a fluid at all.
# - **condition**: Instance of the class ```boxsimu.Condition```. Specifies the condition within the box (e.g. the pH, the temperature, and more many more properties of a box that remain constant).
# - **variables**: List of quantified (!) instances of the class ```Variable```. Specifies the variables that are present within the box. The given variables must be quantified - that means a variable's quantified (alias **q**) must have been called.
# - **processes**: List of instances of the class ```Process```. Specifies the processes that are running within the box.
# - **reactions**: List of instances of the class ```Reaction```. Specifies the reactions that are running within the box.
# **Line 22-37**:
# ```python
# inflow = boxsimu.Flow(
# name='Inflow',
# source_box=None,
# target_box=lake,
# rate=flow_rate,
# tracer_transport=True,
# concentrations={po4: 3e-4},
# )
#
# outflow = boxsimu.Flow(
# name='Outflow',
# source_box=lake,
# target_box=None,
# rate=flow_rate,
# tracer_transport=True,
# )
# ```
#
# Two instances of the class ```Flow``` are instantiated. A flow represents a transport of a fluid from one box to another, or an exchange of fluid-mass with the environment of the system (the ''the outside''). The first flow (```inflow```) represents a river that is flowing into the lake, and has an associated phosphate concentratin of $3\cdot10^{-4}\frac{kg}{kg}$. The second flow represent the outflow of the lake.
#
# The class ```Flow``` has the following signature:
#
# ```python
# Flow(name, source_box, target_box, rate,
# tracer_transport=True, concentrations={}):
# ```
#
# - **name**: String which also has to be a valid python variable name since it is internally used to identify specific flow instances. Thus, 'flow123' or 'flow_123' would be valid, while 'flow 123' or '123flow' would not be valid name arguments.
# - **source_box**: Instance of the class ```Box``` or None. Specifies where the flow originates. If source_box is set to None, the flow is assumed to come from outside the system.
# - **target_box**: Instance of the class ```Box``` or None. Specifies where the flow ends. If target_box is set to None, the flow is assumed to go outside the system.
# - **rate**: Pint Quantity or function that returns Pint Quantity of dimensionality [M/T]. Defines the mass-transport per time of the flow.
# - **tracer_transport**: Boolean. Specifies whether variables are passively transported by this flow.
# - **concentrations**: Dict of Variables and associated concentrations (Pint Quantity of dimensionality [M/M]). Specifies the concentration of variables within the flow. This attribute can only be set for flows that originate form outside the system (flows that originate from a box within the system always have concentrations equal to the concentration of the box itself).
# **Line 43-48**:
# ```python
# system = boxsimu.BoxModelSystem(
# name='lake_system',
# description='Simple Lake Box Model',
# boxes=[lake,],
# flows=[inflow, outflow,],
# )
# ```
# An instance of the class ```BoxModelSystem``` is instantiated. A BoxModelSystem represents the system that will be simulated with boxsimu.
#
# The class ```BoxModelSystem``` has the following signature:
#
# ```python
# BoxModelSystem
# ```
# ### Solving/Simulating the system
# The system defined in boxsimu can now easily be solved using the solve method of the BoxModelSystem class:
sol = system.solve(800*ur.day, 1*ur.day)
# The simulation output is accessible as a pandas dataframe: ```Solution.df```. In our example we can retrieve the model output as follows:
sol.df[:10]
# Example: The first 10 timesteps of the po4 mass in the box lake can be accessed by:
sol.df[('lake', 'po4')].head(10)
# The output can now be visualized using several methods of the class ```Solution```. In the following just a few examples are given.
sol.plot_variable_concentration(po4)
# ## References
# - Schwarzenbach, <NAME>., <NAME>, and <NAME>. "Environmental Organic Chemistry. 2003, Hoboken." 808-811.
# ## Add custom css
from IPython.core.display import HTML
#import urllib2
#HTML( urllib2.urlopen('http://bit.ly/1Bf5Hft').read() )
HTML(open('../costum_jupyter_look.css', 'r').read())
|
examples/tutorial/tutorial_part1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imoport necessary modules
# +
import numpy as np
import sep
import astropy.io
from astropy.io import fits
# %matplotlib inline
import matplotlib.pyplot as plt
from matplotlib import rcParams
# Set image dimensions with rcParams
rcParams['figure.figsize'] = [10., 8.]
# -
# # Open fits file and extract data
# +
#fits image declaration and array conversion
hdu_list = fits.open('hlsp_hudf12_hst_wfc3ir_udfmain_f105w_v1.0_drz.fits')
data = hdu_list[0].data
data = data.byteswap().newbyteorder()
# -
# # Display image
# +
#image view
m, s = np.mean(data), np.std(data)
plt.imshow(data, interpolation='nearest', cmap='gray', vmin=m-s, vmax=m+s, origin='lower')
plt.colorbar();
plt.savefig('f105w_fig_1')
# -
# # Extract background from image data
#measure spacially varying background on the image
bkg = sep.Background(data)
# # Display global background level and rms
#'global' mean noise of image background
print(bkg.globalback)
print(bkg.globalrms)
#evaluate background as 2-d array maintaining size of original image
bkg_image = bkg.back()
# # Display image of separated background
# +
#view the background
plt.imshow(bkg_image, interpolation='nearest', cmap='gray', origin='lower')
plt.colorbar();
plt.savefig('f105w_fig_2')
# -
#evaluate background noise as 2-d array maintaining size of original image
bkg_rms = bkg.rms()
# # Display image of background rms
# +
#view the background noise
plt.imshow(bkg_rms, interpolation='nearest', cmap='gray', origin='lower')
plt.colorbar();
plt.savefig('f105w_fig_3')
# -
# # Separate background from image data
#subtraction of background noise
data_sub = data - bkg
# # Extract the objects from the image
# Define source of objects, set threshold of object detection, set variance for detection threshold
objects = sep.extract(data_sub, 1.5, err=bkg.globalrms)
# # List the number of detected objects from the array
# Display amount of objects detected as the length of the array
len(objects)
# # Show the background-subtracted image and create shapes to highlight each detected object
# +
from matplotlib.patches import Ellipse
#plot background subtracted image
fig, ax = plt.subplots()
m, s = np.mean(data_sub), np.std(data_sub)
im = ax.imshow(data_sub, interpolation='nearest', cmap='gray', vmin=m-s, vmax=m+s, origin='lower')
#plot an ellipse for each object
for i in range(len(objects)):
e = Ellipse(xy = (objects['x'][i], objects['y'][i]),
width = 6*objects['a'][i],
height = 6*objects['b'][i],
angle = objects['theta'][i] * 180. / np.pi)
e.set_facecolor('none')
e.set_edgecolor('red')
ax.add_artist(e)
plt.savefig('f105w_fig_4')
# -
# # Display the parameters extracted from the objects array
#available fields
objects.dtype.names
# # Extract the location and flux data from each object
flux, fluxerr, flag = sep.sum_circle(data_sub, objects['x'], objects['y'], 3.0, err=bkg.globalrms, gain=1.0)
# # Create loop to display the flux from the first 10 detected objects
#shows the first ten object results
for i in range(10):
print("object {:d}: flux = {:f} +/- {:f}".format(i, flux[i], fluxerr[i]))
# # Convert flux data to AB magnitude, and create histogram to plot data
# +
data = hdu_list[0]
PHOTPLAM = data.header['PHOTPLAM']
PHOTFLAM = data.header['PHOTFLAM']
#convert flux to ABMAG
abmag_zpt = -2.5 * np.log10(PHOTFLAM) - 21.10 - 5 * np.log10(PHOTPLAM) + 18.692
abmag = -2.5 * np.log10(np.fabs(flux)) + abmag_zpt
#create histogram of ABMAG flux conversions
plt.hist(abmag, range=[15,40], bins=300)
plt.savefig('f105w_fig_5')
plt.show()
# -
|
Final_project_HDF_f105w_evaluation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Broadcast Joins
#
# Joining two (or more) data sources is an important and elementary operation in an relation algebra, like Spark. But actually the implementation is not trivial, especially for distributed systems like Spark. The main challenge is to physically bring together all records that need to be joined from both data sources onto a single machine, otherwise they cannot be merged. This means that data needs to be exchanged over the network, which is complex and slower than local access.
#
# Depending on the size of the DataFrames to be joined, different strategies can be used. Spark supports two different join implementations:
# * Shuffle join - will shuffle both DataFrames over the network to ensure that matching records end up on the same machine
# * Broadcast join - will provide a copy of one DataFrames to all machines of the network
#
# While shuffle joins can work with arbitrary large data sets, a broadcast join always requires that the broadcast DataFrame completely fits into memory on all machines. But it can be much faster when the DataFrame is small enoguh.
#
# ### Weather Example
#
# Again we will investigate into the different join types with our weather example.
# # 1 Load Data
#
# First we load the weather data, which consists of the measurement data and some station metadata.
storageLocation = "s3://dimajix-training/data/weather"
# ## 1.1 Load Measurements
#
# Measurements are stored in multiple directories (one per year). But we will limit ourselves to a single year in the analysis to improve readability of execution plans.
# +
from functools import reduce
from pyspark.sql.functions import *
# Read in all years, store them in an Python array
raw_weather_per_year = [
spark.read.text(storageLocation + "/" + str(i)).withColumn("year", lit(i))
for i in range(2003, 2015)
]
# Union all years together
raw_weather = reduce(lambda l, r: l.union(r), raw_weather_per_year)
# -
# Use a single year to keep execution plans small
raw_weather = spark.read.text(storageLocation + "/2003").withColumn("year", lit(2003))
# ### Extract Measurements
#
# Measurements were stored in a proprietary text based format, with some values at fixed positions. We need to extract these values with a simple SELECT statement.
weather = raw_weather.select(
col("year"),
substring(col("value"), 5, 6).alias("usaf"),
substring(col("value"), 11, 5).alias("wban"),
substring(col("value"), 16, 8).alias("date"),
substring(col("value"), 24, 4).alias("time"),
substring(col("value"), 42, 5).alias("report_type"),
substring(col("value"), 61, 3).alias("wind_direction"),
substring(col("value"), 64, 1).alias("wind_direction_qual"),
substring(col("value"), 65, 1).alias("wind_observation"),
(substring(col("value"), 66, 4).cast("float") / lit(10.0)).alias("wind_speed"),
substring(col("value"), 70, 1).alias("wind_speed_qual"),
(substring(col("value"), 88, 5).cast("float") / lit(10.0)).alias("air_temperature"),
substring(col("value"), 93, 1).alias("air_temperature_qual"),
)
# ## 1.2 Load Station Metadata
#
# We also need to load the weather station meta data containing information about the geo location, country etc of individual weather stations.
stations = spark.read.option("header", True).csv(storageLocation + "/isd-history")
# # 2 Standard Joins
#
# Per defaulkt Spark will automatically decide which join implementation to use (broadcast or hash exchange). In order to see the differences, we disable this automatic optimization and later we will explicitly instruct Spark how to perform a join.
#
# With the automatic optimization disabled, all joins will be performed as hash exchange joins if not told otherwise.
spark.conf.set("spark.sql.adaptive.enabled", False)
spark.conf.set("spark.sql.autoBroadcastJoinThreshold", -1)
# ## 2.1 Original Execution Plan
#
# Let us have a look at the execution plan of the join.
df = weather.join(
stations, (weather.usaf == stations.USAF) & (weather.wban == stations.WBAN)
)
df.explain(True)
# ### Remarks
#
# As said before, the join is a `SortMergeJoin` requiring a hash exchange shuffle operation. The join has the following steps:
# 1. Filter away `NULL` values (this is an inner join)
# 2. Repartition both DataFrames according to the join columns (`Exchange hashpartitioning`) with the same number of partitions each
# 3. Sort each partition of both DataFrames independently
# 4. Perform SortMergeJoin of both DataFrames by merging two according partitions from both DataFrames
#
# This is a rather expensive operation, since it requires a repartitioning over network of both DataFrames.
# ## 2.2 Explicit Broadcast Joins
#
# Now let us perform the logically same join operation, but this time using a *broadcast join* (also called *mapside join*). We can instruct Spark to broadcast a DataFrame to all worker nodes by using the `broadcast` function. This actually serves as a hint and returns a new DataFrame which is marked to be broadcasted in `JOIN` operations.
df = weather.join(
broadcast(stations), (weather.usaf == stations.USAF) & (weather.wban == stations.WBAN)
)
df.explain(True)
# ### Remarks
#
# Now the execution plan looks significantly differnt. The stations metadata DataFrame is now broadcast to all worker nodes (still a network operation), but the measurement DataFrame does not require any repartitioning or shuffling any more. The broadcast join operation now is executed in three steps:
# * Filter `NULL` values again
# * Broadcast station metadata to all Spark executors
# * Perform `BroadcastHashJoin`
#
# A broadcast operation often makes sense in similar cases where you have large fact tables (measurements, purchase orders etc) and smaller lookup tables.
# ## 2.3 Automatic Broadcast Joins
#
# Per default Spark automatically determines which join strategy to use depending on the size of the DataFrames. This mechanism works fine when reading data from disk, but will not work after non-trivial transformations like `JOIN`s or grouped aggregations. In these cases Spark has no idea how large the results will be, but the execution plan has to be fixed before the first transformation is executed. In these cases (if by domain knowledge) you know that certain DataFrames will be small, an explicit `broadcast()` will still help.
# ### Reenable automatic broadcast
#
# In order to re-enable Sparks default mechanism for selecting the `JOIN` strategy, we simply need to unset the configuration variable `spark.sql.autoBroadcastJoinThreshold`.
spark.conf.unset("spark.sql.autoBroadcastJoinThreshold")
# ### Inspect automatic execution plan
df = weather.join(
stations, (weather.usaf == stations.USAF) & (weather.wban == stations.WBAN)
)
df.explain(True)
# ### Remarks
#
# Since the stations metadata table is relatively small, Spark automatically decides to use a broadcast join again.
|
pyspark-advanced/jupyter-joins/Broadcast Joins - Full.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### The purpose of this notebook to convert the JSON output of “bitcoinabuse” crawler to the data frame and give a quick look into it.
#
# Import modules
import pandas
import json
import glob
# Import data
iles = glob.glob('../data/bitcoinabuse crawler data/json/*')
# Temporary data holders
DATA = list()
err = list()
# Convert the data
for file in files:
data = open(file,'r').read()
for con in data.split('\n'):
con = con.replace("\'", "\"")
con = con.replace('u"', '"')
try:
DATA.append(json.loads(con))
except:
err.append(file)
# Sample data view
DATA
|
notebooks/bitcoinabuse data analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exercise 3.01: Mall Customer Segmentation - Understanding the Data
import numpy as np, pandas as pd
import matplotlib.pyplot as plt, seaborn as sns
# %matplotlib inline
data0 = pd.read_csv("Mall_Customers.csv")
data0.head()
data0.info()
# #### Renaming columns
data0.rename({'Annual Income (k$)':'Income', 'Spending Score (1-100)':'Spend_score'}, axis=1, inplace=True)
data0.head()
data0.describe()
# ## Exercise 3.02: Traditional Segmentation of Mall Customers
data0.Income.plot.hist()
plt.xlabel('Income')
plt.show()
# 2. Create a new column ‘Cluster’ to have the values ‘Low Income’, ‘Moderate Income’, ‘High earners’ for customers with income in the ranges < 50, 50 – 90, and >= 90 respectively, using the code below.
data0['Cluster'] = np.where(data0.Income >= 90, 'High earners', \
np.where(data0.Income < 50, \
'Low Income', 'Moderate Income'))
data0.groupby('Cluster')['Income'].describe()
# ## Exercise 3.03: Standardizing Customer Data
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
cols_to_scale = ['Age', 'Income', 'Spend_score']
data_scaled = data0.copy()
data_scaled[cols_to_scale] = scaler.fit_transform(data0[cols_to_scale])
data_scaled[cols_to_scale].describe()
# ### Exercise 3.4: Calculating distance between customers
sel_cols = ['Income', 'Spend_score']
cust3 = data_scaled[sel_cols].head(3)
cust3
from scipy.spatial.distance import cdist
cdist(cust3, cust3, metric='euclidean')
np.sqrt((-1.739+1.739)**2 + (-0.4348-1.1957)**2)
# ## Exercise 3.05: k-means Clustering on Mall Customers
cluster_cols = ['Income', 'Spend_score']
data_scaled[cluster_cols].head(3)
# #### Visualize the data using a scatterplot
data_scaled.plot.scatter(x='Income', y='Spend_score')
plt.show()
from sklearn.cluster import KMeans
model = KMeans(n_clusters=5, random_state=42)
# +
model.fit(data_scaled[cluster_cols])
data_scaled['Cluster'] = model.predict(data_scaled[cluster_cols])
# -
data_scaled.head(3)
# ### Visualizing the clusters
# +
markers = ['x', '*', '.','|','_']
for clust in range(5):
temp = data_scaled[data_scaled.Cluster == clust]
plt.scatter(temp.Income, temp.Spend_score, marker=markers[clust], label="Cluster "+str(clust))
plt.xlabel('Income')
plt.ylabel('Spend_score')
plt.legend()
plt.show()
# -
# ### Understanding and describing the clusters
data0['Cluster'] = data_scaled.Cluster
data0.groupby('Cluster')['Income', 'Spend_score'].mean().plot.bar()
plt.show()
# ### Exercise 3.6: Dealing with High-Dimensional Data
# 1. Create a list ‘cluster_cols’ containing the columns, ‘Age’, ‘Income’ and ‘Spend_score’ that will be used for clustering. Print the first 3 rows of the dataset for these columns.
cluster_cols = ['Age', 'Income', 'Spend_score']
data_scaled[cluster_cols].head(3)
# 2. Perform k-means clustering specifying 4 clusters using the scaled features. Specify random_state as 42. Assign the clusters to the ‘Cluster’ column.
model = KMeans(n_clusters=4, random_state=42)
model.fit(data_scaled[cluster_cols])
data_scaled['Cluster'] = model.predict(data_scaled[cluster_cols])
# 3. Using PCA on the scaled columns, create new columns 'pc1' and 'pc2' containing the data for PC1 and PC2 respectively.
# +
from sklearn import decomposition
pca = decomposition.PCA(n_components=2)
pca_res = pca.fit_transform(data_scaled[cluster_cols])
data_scaled['pc1'] = pca_res[:,0]
data_scaled['pc2'] = pca_res[:,1]
# -
# 4. Visualize the clusters by using different markers and colours for the clusters on a scatterplot between 'pc1' and 'pc2' using the code below.
# +
markers = ['x', '*', 'o','|']
for clust in range(4):
temp = data_scaled[data_scaled.Cluster == clust]
plt.scatter(temp.pc1, temp.pc2, marker=markers[clust], label="Cluster "+str(clust))
plt.xlabel('PC1')
plt.ylabel('PC2')
plt.legend()
plt.show()
# -
# 5. Understanding the clusters
# a. Print the avg. values of the original features used for clustering against the four clusters.
# b. For a visual analysis, plot the average values for these features against the clusters.
# c. Which features are the most differentiated for the clusters?
#
data0['Cluster'] = data_scaled.Cluster
data0.groupby('Cluster')[['Age', 'Income', 'Spend_score']].mean()
data0.groupby('Cluster')[['Age', 'Income', 'Spend_score']].mean().plot.bar()
plt.show()
|
Chapter03/Exercise3.01-3.06.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ngl\_resum: a package to resum non-global logarithms at leading logarithmic accuracy
# If you use the package ngl\_resum, please cite [doi:10.1007/JHEP09(2020)029](https://inspirehep.net/literature/1798660).
# In this documentation we show some features of ngl\_resum. In particular, we want to visit each of the classes defined in the module and explain their main purposes. We suggest this notebook to be used in Binder:
# [](https://mybinder.org/v2/gh/MarcelBalsiger/ngl_resum/master?filepath=%2Fdocs%2Fnglresum.ipynb)
# To have this example working as a jupyter notebook, one needs to have the packages numpy, physt and - obviously - ngl\_resum installed. The easiest way to do this is to use <code>pip install ngl_resum</code>. Details may be found here: [https://packaging.python.org/tutorials/installing-packages/#use-pip-for-installing](https://packaging.python.org/tutorials/installing-packages/#use-pip-for-installing)
# ## Imports
# We start by importing the package ngl\_resum and numpy:
import ngl_resum as ngl
import numpy as np
# ## FourVector
# We start with the <code>FourVector</code> class. As its name suggests, this class is used to describe fourvectors and contains some information specifically used in collider physics. To instantiate a <code>FourVector</code>, we have to feed all four components of it. Let us define <code>fvA</code> with energy $e$ <code>energyFvA</code> and momenta $p_i$ <code>iMomFvA</code>:
energyFvA=6.9
xMomFvA=4.2
yMomFvA=3.5
zMomFvA=1.2
fvA=ngl.FourVector(energyFvA,xMomFvA,yMomFvA,zMomFvA)
print(fvA)
# ### Attributes
# Of course, we can access the four individual coordinates:
print("energy: ",fvA.e,"\nx-mom.: ",fvA.px,\
"\ny-mom.: ",fvA.py,"\nz-mom.: ",fvA.pz)
# In some cases, it may become useful to have the momentum vector displayed as a numpy-array:
print("numpy-array: ",fvA.vec)
# The two angles $\theta$ (angle between the z-axis or beam-axis)and $\phi$ (angle in the x-y-plane) of the three-vector are attributes:
print("theta: ",fvA.theta,"\nphi: ",fvA.phi)
# We can also access the mass and the velocity of the particle:
print("mass: ",fvA.m,"\nvelocity: ",fvA.beta)
# The length of the spatial vector $\sqrt{p_x^2+p_y^2+p_z^2}$ can also be accessed:
print("np.sqrt(px*px+py*py+pz*pz): ",fvA.absSpace)
# Collider-physics specific attributes are the transverse energy $E_T$, the transverse momentum $p_T$, the rapidity $y=\frac{1}{2}\ln\frac{e+p_z}{e-p_z}$ and the pseudorapidity $\eta=-\ln\left(\tan\frac{\theta}{2}\right)$:
print("transverse energy: ",fvA.eT,"\ntransverse momentum: ",fvA.pT,\
"\nrapidity: ",fvA.rap,"\npseudorapidity: ",fvA.pseudorap)
# ### Operations
# Let us instantiate another <code>FourVector</code>:
fvB=ngl.FourVector(5,4,3,0)
# We can now add and subtract the fourvectors with the usual operators:
print("fvA+fvB:",fvA+fvB,"\nfvA-fvB: ",fvA-fvB)
# The multiplication operator $*$ can be used to give a scalar product of two fourvectors or as the multiplication of a scalar. Note that we use te mostly-minus metric (+ - - -):
print("fvA*fvB:",fvA*fvB,"\n10*fvA: ",10*fvA)
# The division by a scalar does work, too:
print("fvA/10:",fvA/10)
# ### Methods
# To measure the squared angular distance $\Delta R^2=\Delta\phi^2+\Delta\eta^2=|\phi_A-\phi_B|^2+|\eta_A-\eta_B|^2$ between two fourvectors, we can use
print("deltaR^2:",fvA.R2(fvB)," or ",fvB.R2(fvA))
# while the cosine of the spatial angle between two fourvectors can be accessed by
print("cosTheta:",fvA.costheta(fvB)," or ",fvB.costheta(fvA))
# We can check, whether the <code>FourVector</code> is massive or massless. A <code>FourVector</code> <code>a</code> is treated as massive (or time-like), if <code>a*a</code> is larger than $10^{-7}$, otherwise it is treated as massless (or light-like):
fvA.isMassive()
fvA.isMassless()
fvB.isMassive()
fvB.isMassless()
# We can check whether two FourVectors are the same. We consider two FourVectors <code>a</code> and <code>b</code> to be the same, if <code>(a-b).e^2+(a-b).px^2+(a-b).py^2+(a-b).pz^2</code> is smaller than $10^{-10}$ to account for rounding errors. To apply this check, we can use
fvA.isSame(fvB)
fvA.isSame(fvA+ngl.FourVector(0.000000001,0.000000001,0.000000001,0))
# One last method of the <code>FourVector</code> class is going to be the tensor product fvA$_\mu$fvB$_\nu$ (which is probably not going to be used that often), given by
print("fvA.metric.fvB:\n",fvA.tensorProd(fvB))
# ## Boost
# The <code>Boost</code> class takes care of the boosting procedure as described in Section 3.1 [doi:10.1007/JHEP09(2020)029](https://inspirehep.net/literature/1798660). It takes two arbitrary momentum vectors from the lab frame and creates the boost from lab frame to the frame where these two fourvectors are back-to-back alongside the z-axis. For transparency we take the first two vectors of the event from Appendix A of above article, as given in (A.3):
p1=ngl.FourVector(504.7,125.6,82.44,-450.4)
u1=p1/p1.e
u2=ngl.FourVector(1,0,0,-1)
# Now to get the boost accounting for the boost from the lab frame to the frame where <code>u1</code> is alongside the positive z-axis and <code>u2</code> alongside the negative z-axis, we just have to instantiate a <code>Boost</code> with the two fourvectors as arguments:
bst=ngl.Boost(u1,u2)
# ### Attributes
# We have access to each single transformation $X$, $B$ and $Z$ as defined in Section 3.1 and thoroughly explained with an example in the pages after (A.3) of [doi:10.1007/JHEP09(2020)029](https://inspirehep.net/literature/1798660). Note, that each of these matrices are 4x4 numpy arrays.
# We start out with the boost $X$, which is a rotation that puts the added two initial fourvectors along the x-axis:
np.dot(bst.X,(u1+u2).vec)
# Now we apply the boost $B$, which removes the spatial component of above fourvector:
np.dot(bst.B,np.dot(bst.X,(u1+u2).vec))
# Finally, we apply a second rotation $Z$ that puts the two initial vectors alongside the z-axis, with <code>u1</code> in the positive direction:
np.dot(bst.Z,np.dot(bst.B,np.dot(bst.X,u1.vec)))
np.dot(bst.Z,np.dot(bst.B,np.dot(bst.X,u2.vec)))
# As we can see, the two fourvectors are now nicely aligned back-to-back. The full boost is also stored in an attribute:
u1prime=np.dot(bst.LABtoCMS,u1.vec)
u1prime
# The same goes for the inverse boost:
u1BoostedBack=np.dot(bst.CMStoLAB,u1prime)
u1BoostedBack
u1BoostedBack-u1.vec
# ### Methods
# Of course, the numpy arrays are not easy to handle as we have to keep in mind which variable is a <code>FourVector</code> and which one a numpy array. We have a shortcut to erase this problem. We can take the boost and apply it on any <code>FourVector</code> and get back the <code>FourVector</code> in the new frame as follows:
fvu1prime=bst.boostLABtoCMS(u1)
fvu1prime
fvu1BoostedBack=bst.boostCMStoLAB(fvu1prime)
fvu1BoostedBack
fvu1BoostedBack.isSame(u1)
# ## Hist
# The <code>Hist</code> class acts as an adapter to the physt package and is immensly based on it. It accounts for the histograms $R(t)$ that is the result of the resummation (see (4.3) of [doi:10.1007/JHEP09(2020)029](https://inspirehep.net/literature/1798660)).
# When initializing a <code>Hist</code>, we at least need to provide a number of bins <code>nbins</code> and a maximal value for $t$, <code>tmax</code>:
nbins=10
tmax=0.1
hst=ngl.Hist(nbins, tmax)
hst
# Another possibility is to also calculate an error estimate of each bin:
hstErr=ngl.Hist(nbins,tmax,errorHistCalc=True)
hstErr
# Compared to the rest of the classes we switch the order and postpone the discussion of the attributes to after the methods and operations. This is due to the fact that the methods are mainly used to populate the histograms to avoid looking at a wall of zeroes in the discussion of the attributes.
# For the sake of streamlinedness, we will stop discussing the more involved case of the error estimation at this point. The discussion thereof is quite involved and provides little insight. We will look at the extraction of the error from the histogram later.
# ### Methods
# We have two functions that can be used to set a whole histogram to zero or to one:
hst.setOne()
hst
hst.setZero()
hst
# To populate the histogram, we can use <code>addToBin</code> by specifying the value of <code>t</code> at which we add a weight <code>w</code>:
tVal=0.053
w=0.696969
hst.addToBin(tVal,w)
hst
# ### Operators
# Let us start with a disclaimer - we assume the two histograms to be initialized with the same number of bin <code>nbins</code> and the same maximal $t$ value <code>tmax</code>. If this is not the case, anything might happen.
# To show the use of some operators, we will populate two histograms with some random numbers:
hst1=ngl.Hist(nbins, tmax)
hst2=ngl.Hist(nbins, tmax)
for i in range(0,50):
hst1.addToBin(tmax*np.random.random_sample(),np.random.random_sample())
hst2.addToBin(tmax*np.random.random_sample(),np.random.random_sample())
# Let us have a look at how these historgams are populated:
hst1
hst2
# We can add and subtract the histograms. This sums (or subtracts) the entry of each bin:
hst1+hst2
hst1-hst2
# The multiplication of two Hists multiplies the entry of each bin, and we can also multiply with a scalar (which multiplicates each entry by the scalar):
hst1*hst2
10*hst1
# Division by a scalar is possible as well:
hst1/10
# ### Attributes
# Let us finally look at how to access the data in <code>Hist</code>. The Histograms certainly knows about the number of bins and maximal $t$:
print("nbins: ",hst1.nbins,"\ntmax: ",hst1.tmax)
# To access the entries of the histogram, we can do so:
hst1.entries
# The bin values can be read out as well. As it is sometimes useful to have the lower or upper bin boundary or the central value, we have created access to all of them:
print("central bin values: ",hst1.centerBinValue)
print("lower bin boundary: ",hst1.lowerBinBoundary)
print("upper bin boundary: ",hst1.upperBinBoundary)
# Now let us have a quick glance at the error estimations of the bins. Note, that while the representation of the histogram comes with the error itself, due to intricacies of the error computation when multiplying histograms (which is used in the showering procedure), we are only able to access the squared of the error estimate. To illustrate this, let us first populize the <code>hstErr</code> from above and show its representation:
for i in range(0,50):
hstErr.addToBin(tmax*np.random.random_sample(),np.random.random_sample())
hstErr
# Now we will access the squared error of the bins:
hstErr.squaredError
# This might seem a little bit odd. For more insight into the error handling we refer to the documentation of the example codes.
# ## Event
# Now we get to the core of the package. An instance of <code>Event</code> contains all the reevant information to start one showering. We will not go too deep into details here, but mainly refer to the two example codes. To instantiate an <code>Event</code>, one can either feed it
# * a dipole (using the <code>feedDipole</code> parameter), or
# * an <code>pylhe.LHEEvent</code> read-in via <code>pylhe.readLHE</code> (using the <code>eventFromFile</code> parameter)
#
# To each of those we have an example code.While we will not explain every attribute and method of this class in detail, we still want to give an overview of some intricacies.
# First of all we want to discuss the feedDipole feature. Via <code>Event(feedDipole=dipole)</code> we can set up the showering of one single dipole, consisting of two fourVectors in an array:
leg1=ngl.FourVector(1,0,0,0.5)
leg2=ngl.FourVector(1,0,0,-0.5)
dipole=[leg1,leg2]
evDip=ngl.Event(feedDipole=dipole)
# This feature is straightforward enough. We have set up this dipole for showering.
# The other feature of <code>Event</code> is the more intricate showering of an event read in from a .lhe-file. After feeding the <code>pylhe.LHEEvent</code> into the parameter <code>eventFromFile</code> we have two additional options, namely
# * whether we want to form the color-connected dipoles between the incoming and outgoing particles of the event or between the incoming and intermediate ones, and
# * whether we also want to account for the decay dipoles between the intermediate and the outgoing particles.
#
# An example where we shower both the dipoles formed by the incoming-intermediate particles and the intermediate-outgoing particles is given for example in Section 5 of [doi:10.1007/JHEP09(2020)029](https://inspirehep.net/literature/1798660) .
# To keep this documentation simple, we will not actually read in a .lhe-file, and therefore can not go hands-on here. If you want to play around with this feature, we suggest you move to the example code.
# To instantiate an <code>Event</code> by an <code>event=pylhe.LHEEvent</code>, we have to use <br><code>evLHE=ngl.Event(eventFromFile="pylhe.LHEEvent")</code>.<br>
# It sets up the <code>Event</code> with the default case of color-sorting the incoming and outgoing particles. To set up the showering of the incoming-intermediate particle dipoles, we have to use <br><code>evLHE=ngl.Event(eventFromFile=pylhe.LHEEvent, productionDipoles='intermediate')</code>,<br>
# and if we not only want to have the production dipoles showered, but also the dipoles associated to the decay, we have to use <br><code>evLHE=ngl.Event(eventFromFile=pylhe.LHEEvent,productionDipoles='intermediate',decayDipoles=True)</code>.<br>
# Note that you will probably seldom use these additional features and most oftenly only have to use <code>evLHE=ngl.Event(eventFromFile="pylhe.LHEEvent")</code>, except if you work with top quarks.
# Note that if you instantiate an <code>Event</code> using a <code>pylhe.LHEEvent</code>, you have access to the weight as well as an array of the <code>FourVector</code> of each kind of particles in the form of an attribute. You can access the weight of the event via
# <code>ev=ngl.Event(eventFromFile="pylhe.LHEEvent")</code><br>
# <code>ev.weight</code><br>
# If you want the fourvectors of all incoming up-type quarks and antiquarks, you can access them via <br>
# <code>ev=ngl.Event(eventFromFile="pylhe.LHEEvent")</code><br>
# <code>ev.incomingUp</code>.<br>
# In the same way, you can access <code>ev.statusType</code>, with <code>status</code> being <code>incoming</code>,<code>intermediate</code> or <code>outgoing</code>, and <code>Type</code> being <code>Down</code>, <code>Up</code>, <code>Strange</code>, <code>Charm</code>, <code>Bottom</code>, <code>Top</code>, <code>Electron</code>, <code>ENeutrino</code>, <code>Muon</code>, <code>MNeutrino</code>, <code>Tau</code>, <code>TNeutrino</code>, <code>Gluon</code>, <code>Photon</code>, <code>ZBoson</code>, <code>WBoson</code> or <code>Higgs</code>.
# ## OutsideRegion
# Just as the <code>Event</code> class, <code>OutsideRegion</code> is very specific to the observable you are considering. Its whole purpose is to tell the <code>Shower</code>, whether a <code>FourVector</code> is pointing into the region where it gets vetoed. The nomenclature of **outside** comes from the textbook example of the interjet energy flow, where radiation that is not inside the jets gets vetoed.
# We can initiate the <code>OutsideRegion</code> with or without an <code>Event</code>. Whether you should feed it an <code>Event</code> or not depends on whether the region where you want to veto radiation depends on the distribution of the outgoing particles. We have one example code each for the usage with and without an <code>Event</code>.
# An instance of <code>OutsideRegion</code> doesn't do anything. It containes the stub of a method <code>outside(self,v)</code> which needs to be implemented by you. To do so, you need to write a method
def _outside(self,v):
#
# Code that checks whether v is a FourVector
# landing outside.
#
retVal=True # if v outside
retVal=False # if v not outside
return (retVal)
# and - after creating an instance of OutsideRegion
outsideRegion=ngl.OutsideRegion()
# exchange the stub of <code>outside(self,v)</code> of your instance <code>outsideRegion</code> to your method by invoking
outsideRegion.outside = _outside.__get__(outsideRegion,ngl.OutsideRegion)
# For more details we refer to the two example codes which show the handling of the <code>OutsideRegion</code> class.
# ## Shower
# Let us now get to the core of our resummation precedure, the showering of an <code>Event</code>. To instantiate a <code>Shower</code>, we feed it the following parameters (most of which come with a default choice):
# +
# event: Event,
# outsideRegion: OutsideRegion,
# nsh: int=50,
# nbins: int=100,
# tmax: float=0.1,
# cut: float=5.0,
# fixedOrderExpansion: bool=True,
# virtualSubtracted: bool=False
# -
# Of these parameters, <code>event</code> and <code>outsideRegion</code> unsurprisingly contain the <code>Event</code> to shower with the respective <code>OutsideRegion</code> under consideration. The number of showerings you want to apply on the <code>event</code> is fed in via <code>nsh</code> (<code>50</code> by default). To create the <code>Hist</code> which will eventually be the result of the resummation, we can change the <code>nbins</code> (<code>100</code> by default) and <code>tmax</code> (<code>0.1</code> by default). We can also change the collinear cutoff <code>cut</code> (<code>5.0</code> by default), which corresponds to $\eta_{max}$ as discussed in (A.14) of [doi:10.1007/JHEP09(2020)029](https://inspirehep.net/literature/1798660). Finally, we can decide whether or not we want to calculate the first two expansion parameters of the resummation as given in (4.3) of [doi:10.1007/JHEP09(2020)029](https://inspirehep.net/literature/1798660) in <code>fixedOrderExpansion</code> (<code>True</code> by default). The last option is <code>virtualSubtracted</code> will most likely have to be turned off (as is the default), it is used to subtract the global one-loop part from the soft anomalous dimension as discussed in (3.5) of [doi:10.1007/JHEP04(2019)020](https://inspirehep.net/literature/1717208).
# Instead of going through the details of the showering precedure we refer to Appendix A of [doi:10.1007/JHEP09(2020)029](https://inspirehep.net/literature/1798660), where this is explained in a very detailed fashion.
|
docs/nglresum.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Variables
# The understanding of variables in programming languages is a must!
#
# Variables exist in (almost) every programming language. However, the term has a different meaning in programming
# compared to mathematics. In programming, a variable is a name to which a value can be assigned. From a (overly
# simplistic) technical point of view, a variable is space in the memory that has a certain value at a point in time.
#
# ## Using variables
# It is possible to put (or assign or write) a value into the variable. Vice versa, it is possible to read the currently
# stored value out of the variable. (Important: After the value of the variable is read, the value in the variable is
# still available and unchanged!)
#
# The value of a variable can be set using the assignment operator `=`. For example, the assertion `a = 2` writes the
# value `2` into the variable `a`. If in subsequent statements, variable `a` is used, the value from the variable is
# taken. For example in the following code cell, first the value `2` is written in to the variable `a`, then the value `3`
# is written into cell `b`. In the final statement, the values are read from variable `a` and `b` and multiplied.
#
# Note: In mathematics, `a = 2` is an *assertion*, which is either true or false. In programming, `a = 2` is an
# *instruction*, which describes the assignment of the value `2` to the variable `a`.
a = 2
b = 3
a * b
# ### Several assignments in a sequence
# The value of a variable can also be changed by an assignment, if assignments take place one after the other. In the
# example below, there is the variable `a`. First, the value `2` is assigned to `a`, then the value `3` is assigned. This
# second assignment deletes the `2`, the memory is just occupied with the new value `3`. The `2` is forgotten.
a = 2
a = 3
a
# ### Assigning multiple values to multiple variables
# Of course, it is possible to have more than one variable in a Python programme. You can assign values to those variables
# as you have already seen.
# +
a = 1
b = 2
c = 3
b
# -
# ### A variable on both sides of the assignment
# Assignments can look strange (or even wrong) through the eyes of mathematics. This is the case, for example, when a
# variable is on both sides of the assignment. In the example below, the variable `a` is increased by `10`. To be more
# precise: First, the value of the variable `a` is read, the `10` is added, and the result is written back into the memory
# location named `a` at the end. This happens to be the same memory location as before.
a = 25
a = a + 10
a
# ### Accessing a variable in read mode
# In the examples above you have seen, that variables can be on the right side of the `=`. These variables are accessed in
# *read mode*. The value is written into the variables on the left side. This variable is accessed in *write mode*. A
# variable can only be accessed in read mode if a value has been assigned to the variable beforehand. Otherwise there is
# an error message. Execute the next cell and try to understand the error message.
a = unknown_variable
# #### Small Excursion: Cells in notebooks are connected to each other
# When a cell in a notebook has been successfully executed, the results are also known in other cells of the notebook.
# - First execute the first of the following two cells. An error message will appear.
# - Then execute the second cell and then the first cell again. This time everything should work.
# - If you want to repeat the whole process, you must reset the output of all cells in the "Kernel" menu at the top. Then
# the variables will *forget* their current values.
a = new_variable + 10
a
new_variable = 20
# ## Sequentialization of assignments
# Sometimes it happens that several operations are applied to one variable. These can also be sequentialized with the help
# of the construct above. This sometimes helps to make a programme better readable. For example, the assignment `a = 5 * 3 * 7 + (2 - 10 * 3)`
# can be written as follows: (When sequentializing, one must be careful not to violate any bracket rule or the like).
a = 5
a = a * 3
a = a * 7
a = a + (2 - 10 * 3)
a
# ## Only one variable on the left side of an assignment
# On the right-hand side of an assignment, there can be complex expressions which themselves contain variables (e.g. a = b
# ** 2 + c ** 2). On the left side, on the other hand, there can only be the one variable. If you execute the following
# construct, an error message will occur.
# +
b = 3
c = 4
a ** 2 = b ** 2 + c ** 2
a
# -
# You have certainly recognised Pythagoras' theorem in the cell above. But how do you then obtain the length of a if b and
# c are given? The calculation can be done in two steps as follows.
# +
b = 3
c = 4
a = b ** 2 + c ** 2
a = a ** 0.5
a
# -
# ## Variable Names
# There are a number of rules for variable names in Python. A variable name must always begin with a letter or an
# underscore `_`. This can be followed by any number of letters, digits and underscores.
#
# Variable names may not contain special characters or spaces. Thus, for example, the following variable names are valid:
name = "David"
surname = "Bowie"
account_balance = -2000
_new_balance = 1000
# In contrast, the following variable names are invalid. Execute the cell and check the error message.
1_konto = 1234
email@fh = '<EMAIL>'
# In Python, variable names are case sensitive. This means that in a Python program, `name` and `Name` represent different
# variables. This is illustrated in the following example.
# +
name = "Joey"
Name = "Ramone"
name + " " + Name
# -
# Finally, there are a number of *reserved keywords* in Python that must not be used as variable names. These keywords
# have a special meaning in Python, which you will learn about in the course of the lecture. Example of reserved keywords
# are `and`, `while` or `if`. Execute the following cell and have a look at the error message.
if = 42
# ## Conventions for Variable Names
# In the Python community, lower case is preferred for variable names. Thus, `name` is used instead of `Name`. Variable
# names that consist of several words are separated by an underscore `_`, e.g. `account_number` or
# `minimum_account_balance`.
#
# Not every allowed variable name is a good variable name! A good programming style (not only in Python) is characterised
# by the fact that a program is easy for a human to understand.
#
# > Any fool can write code that a computer can understand. Good programmers write code that humans can understand.<br>
# > <NAME>, 2008.
#
# For this reason, you should use variable names that have a meaning.
#
# - `new_account_balance` is better than `nab`
# - `car_length` is better than `length`
#
# Make your programs readable by using good variable names so that you will still be able to understand it in a year's
# time.
a = 5
b = 7
a = b
print(a)
a = 5
b = 7
b = a
print(a)
print(b)
# ## More Details about Variables
#
# In Python, a *variable* is created in an assignment the first time it is used. Variables do not have to be declared
# first, as in some other programming languages.
# A *variable* can be on both the right and left side of the assignment operator at the same time.
a = 10
a = a + 1
print(a)
# In programming the statement in the middle means, that the value in the variable `a` is increased by `1`. Sometimes
# the term *Increment* is used. This construction is used quite often. Thus Python offers a short way of writing this
# statement: `a += 1`. Yes, from a mathematics point of view this looks really weird but it simply means that the value
# stored in `a` is increased by `1`.
a = 1
a += 1
print(a)
# The same short way of writing works with other operators and values:
a = 10
a *= 2
a -= 100
print(a)
# # Exercise
# Calculate the volume and surface area of a cube (side lengths a, b, c equal) or a cuboid (side lengths a, b, c
# different). Proceed as follows:
# - Define reasonable names for the variables representing the side length of the cube and assign values to them.
# - Calculate the area of the cube and assign the result to the variable cube_area.
# - Calculate the volume of the cube and assign the result to the variable cube_volume.
# - Output the result of the calculation using the following statements: `print(cube_area)` and `print(cube_volume)`
#
# Repeat these steps for the cuboid.
|
week_1/week_1_unit_2_variab_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Counterfactual emissions scenarios
#
# A SIMPLE EMISSIONS DECOMPOSITION MODEL OF COUNTERFACTUAL EMISSIONS PATHWAYS from the article "<NAME>., <NAME>., <NAME>., and <NAME>. (2022). From low-growth to post-growth scenarios of climate mitigation. (In peer-review)".
#
# # Introduction
#
# The urgency of climate change mitigation has been recognised at the international governance stage since 1990. However, the general agreement on the problem has yet to translate into an effective action to curb global emissions, which have grown by 60% since 1990. As a result of increasing emissions, we are in a state of accelerated global heating and rapidly approaching the turning point beyond which it will be theoretically impossible to stabilize the global warming between 1.5 °C and 2 °C, as pledged by the Paris Agreement. To meet the Paris Agreement goals we need to keep total carbon emissions within the carbon budget limit, which is estimated at 340 GtCO2 for 1.5 °C and at 1090 GtCO2 for 2 °C of global warming. Staying within those carbon budgets will require a far-reaching and an unprecedented transformation of our economies, lifestyles, and energy systems (IPCC, 2018). In other words, adequate climate mitigation action requires a decisive break with the historical pathway of development.
#
# In this exercise, we will explore by how much the key emissions drivers would need to change to stabilize global warming below 1.5 °C, had we started to mitigate earlier than 2022. By exploring alternative histories of mitigation (counterfactual scenarios), we will analysie how the conditions for accomplishing sufficient emission reductions have changed due to increasing emissions over time.
# # Possible exercises
#
# Get acquainted with the model by trying to answer these questions
#
# 1. Find what would be the necessary rate of annual emissions reductions for staying below 1.5 degrees if we started reducing emissions this year (2022). How would this rate change, had we started mitigation earlier?
#
# 2. Which of the emissions drivers has been historically the most important?
#
# 3. By how much would the emissions have changed, had the global economic growth from 2007 to 2021 increased by two-fold (as the IMF has been projecting before the 2008 financial crisis), if we leave other assumptions unchanged?
#
# 4. If you set all the mitigation levers (all the parameters) to the most ambitious values starting in 2016 (the year after the Paris Agreement was signed), would that be enough to stabilize global warming below 1.5 degrees? How significant is the gap, if there is one?
#
# 5. By how much should we have raised the mitigation action in 1991, had we wanted to accomplish the ambition of the Toronto Conference for CO2 emissions reduction of 20% by 2005.
#
# # Ideas for the Assignment
#
# Come up with a research question related to the climate impacts and/or necessary mitigation measures to reduce them. Design a counterfactual scenario to address this question. Think of an alternative trajectory of development and write down a short narrative description of your scenario. In the narrative description, provide qualitative information concerning the changes for all of the emissions drivers. Argue about the feasibility of the scenario. Select the starting year and the parameters of emissions drivers consistently with your narrative. You can design multiple scenarios and compare the results.
#
# Research question example:
#
# By how much should we have raised the mitigation action in 1991, had we wanted to accomplish the mitigation advice of the Toronto Conference for CO2 emissions reduction of 20% by 2005.To address this question, we explore the parameters related to the efficient use of energy and the decarbonisation of energy generation, but do not assume any changes to population and economic growth.
#
#-----IMPORTING LIBRARIES-----
import numpy as np
import pandas as pd
from scipy.interpolate import CubicSpline
import matplotlib.pyplot as plt
# +
#-----IMPORTING HISTORICAL DATA-----
df = pd.read_excel('data/counterfactual_scenarios_data.xlsx', 'Historical data')
df.head()
CO2_emissions = df['CO2 Emissions (GtCO2)'].to_numpy() # Global CO2 emissions from antropogenic activities [GtCO2]
Final_energy = df['Final Energy (EJ)'].to_numpy() * 10**9 # Final energy consumption [GJ]
GDP = df['GDP ($)'].to_numpy() # Gross Domestic Product [$]
Population = df['Population '].to_numpy() # Population [Number of people]
Low_carbon_penetration = df['Low carbon penetration (%)'].to_numpy() # Share of Low-carbon energy in Final energy consumption [from 0% to 100%]
# -
# # Emissions Drivers
#
# The original IPAT equation relates environmental impacts to population, affluence, and technology, as shown in Equation 1.
#
#
# $$ Impact = Population\cdot Affluence\cdot Technology \; \; \; (1) $$
#
#
# In 1990s, Yoichi Kaya adapted the IPAT equation for the analysis of the underlying drivers (factors) of anthropogenic CO2 emissions from energy.(Kaya, 1990) The KAYA equation relates CO2 to population, per-capita gross domestic product (GDPpc), carbon intensity of energy consumption (CIec), and energy intensity of gross domestic product (EI), as shown in Equations 2 and 3. Energy intensity of GDP can be interpreted as a measure of energy efficiency of the economy. Carbon intensity can be interpreted as the carbon footprint of our system of energy generation.
#
#
# $$ CO_{2}= Population\cdot \frac{GDP}{Population}\cdot \frac{CO_{2}}{Energy}\cdot \frac{Energy}{GDP} \; \; \; (2) $$
#
#
#
#
# $$ CO_{2}= Population\cdot GDPpc\cdot CIec\cdot EI \; \; \; (3) $$
#
#
# While Equation 3 allows us to analyse the dependence of CO2 emissions on the underlying emissions factors, it is a bit difficult to relate to the mitigation policy which is commonly framed around the transition to low-carbon energy sources. To represent this, we decompose the carbon intensity of energy into two factors, which are: the share of low-carbon energy, and the carbon intensity of energy from fossil fuels. Here, low-carbon energy which includes all the energy generated from the renewables, nuclear energy and biomass which are assumed to generate zero direct carbon emissions. Carbon intensity of energy from fossil fuels describes the carbon footprint of part of the energy system that is not low-carbon. Carbon intensity of energy from fossil fuels changes if there is a substitution between different fossil fuel sources, for example, if there is a switch from coal to gas.
#
# $$ CO_{2}= Population\cdot GDPpc\cdot (1-Lowcarbon_{\%})\cdot \frac{CO_{2}}{Energy}\cdot \frac{Energy}{GDP} \; \; \; (4) $$
#
#
#-----DEFINING EMISSIONS DRIVERS-----
Energy_intensity = Final_energy/GDP # Energy intensity of Gross Domestic Product [GJ/$]
Carbon_intensity_FF = CO2_emissions * 10**9 / (Final_energy * (1 - 0.01 * Low_carbon_penetration)) # Carbon intensity of energy [tCO2/GJ]
GDP_per_capita = GDP / Population # Global Gross Domestic Product per capita [$/capita]
Low_carbon_energy = Final_energy * Low_carbon_penetration * 0.01 # Final energy consumption from low-carbon sources (renewables & nuclear) [GJ]
FF_energy = Final_energy - Low_carbon_penetration # Final energy consumption from fossil fuel sources [GJ]
Carbon_intensity_tot = CO2_emissions / Final_energy*10**9 # Carbon intensity of total final energy [tCO2/GJ]
# # Designing a counterfactual scenario
#
# We calculate the average annual change of each of the emissions drivers (y) over from 1990 to 2021 (𝛥t), as shown in Equation 5. We use historical changes of emissions drivers as a benchmark to which we will compare the counterfactual scenarios.
#
# $$ r_{y}= (\frac{y_{t+\Delta t}}{y_{t}})^{\frac{1}{\Delta t}}-1 \; \; \; (5) $$
#
#
# We now provide the underlying assumptions of the counterfactual scenarios. We do so by defining the values of the emissions drivers multipliers (m), which relate the assumed annual change in counterfactual scenarios to the average historical annual change, as shown in Equation 6. For example, by assuming the multiplier for low-carbon energy of 1.2, you assume a 20% faster deployment of low-carbon energy. The multiplier value of 1.0 (default) assumes average annual change in the counterfactual correspond to the historical values.
#
# $$ r_{y,counterfactual}= m_{y}\cdot r_{y,historical} \; \; \; (6) $$
#
# Values of emissions drivers in counterfactual scenario are then calculated as:
#
# $$ y_{counterfactual}(t+1)=y_{counterfactual}(t)\cdot (1 + m_{y} \cdot r_{y,historical}) \; \; \; (7) $$
#
# You can design your own scenario by adjusting the following parameters:
#
# - Average population growth
#
# - Economic growth expressed in GDPpc
#
# - Growth in low carbon energy
#
# - Energy efficiency improvements in the economy
#
# - Carbon intensity improvements of fossil fuels
#
# - Year when the counterfactual scenario starts to deviate from the historical pathway
#
# We provide some guidelines for the choice of parameters in Table 1, but you can also explore scenarios outside the range (just provide justification in the assignments, and try not to crash the model). In addition, you can also select the start year when your scenario begins to deviate from the historical values. The chosen set of assumptions will form a unique counterfactual scenario of emissions. The prepared script of the model will compare your counterfactual scenario to the historical values (see example in Figure 3).
#
# 
#
#-----HISTORICAL AVERAGE CHANGES from 1990 to 2021-----
Avg_Energy_intensity_improvement = 100*((Energy_intensity[-1]/Energy_intensity[0])**(1/31)-1) # Change in Energy intensity
Avg_Carbon_intensity_FF_improvement = 100*((Carbon_intensity_FF[-1]/Carbon_intensity_FF[0])**(1/31)-1) # Change in Carbon intensity of fossil fuel sources (Carbon intensity can change due to fuel switching, for example from coal to natural gas
Avg_GDP_per_capita_growth = 100*((GDP_per_capita[-1]/GDP_per_capita[0])**(1/31)-1) # Growth in GDP per capita
Avg_low_carbon_growth = 100*((Low_carbon_energy[-1]/Low_carbon_energy[0])**(1/31)-1) # Growth in Low carbon energy
Population_growth = 100*((Population[-1]/Population[0])**(1/31)-1) # Population growth
# +
#----COUNTERFACTURAL ASSUMPTIONS-----
Multiplicator_Energy_intensity = 1 # [0 - no change, 1 - historical value, 2 - twice as fast (ambitious), 3 - very ambitiou, >3 (difficult to achieve)] - > By how much (faster/slower) was energy intensity to change compared to the historical value of 1.06% per year
counterfactual_intensity_improvement = Multiplicator_Energy_intensity*Avg_Energy_intensity_improvement
Multiplicator_Carbon_intensity = 1 # [-1 - transition to a coal-powered world, 0 - no change, 1 - historical value, 5 - switching from coal to gas (ambitious)] - > By how much (faster/slower) was carbon intensity of fossil fuels to change compared to the historical value of 0.06% per year
counterfactual_carbon_intensity_FF_improvement = Multiplicator_Carbon_intensity*Avg_Carbon_intensity_FF_improvement
Multiplicator_GDP_per_capita = 1 # [-1 -> deep degrowth scenario, -0.5 -> slow degrowth, 0 -> steady-state economy, 0.5 -> secular stagnation, 1 - historical value, 2 - fast GDP growth] - > By how much (faster/slower) was GDP per capita to change compared to the historical value of 1.41% per year
counterfactual_GDP_per_capita_growth = Multiplicator_GDP_per_capita*Avg_GDP_per_capita_growth
Multiplicator_low_carbon = 1 # [1 - historical value (slow deployment of renewables), 1.5 -> 50% faster deployment (moderately ambitiou), 2-> 100% faster deployment (very ambitious)] - > By how much (faster/slower) was growth of low-carbon energy compared to the historical value of 1.93% per year
counterfactual_low_carbon_growth = Multiplicator_low_carbon*Avg_low_carbon_growth
Multiplicator_population = 1 # [0.5 -> Empty planet scenario, 1 - historical value, 1.5 -> 50% faster population growth] - > By how much (faster/slower) was population growth compared to the historical value of 1.28% per year
counterfactual_population_growth = Multiplicator_population*Population_growth
# +
#-----TIME RANGE-----
year1 = np.arange(1990, 2021+1, 1) # Our data are from 1990 to 2021
year2 = np.arange(2021, 2051, 1)
first_year = 1990 # Begining of the time series
start_year = 2016 # What year do you want to deviate from the historical trajectory?
start = start_year - first_year
# -
#-----EMISSIONS FACTORS IN COUNTERFACTURAL SCENARIOS-----
counterfactual_CO2_emissions = np.zeros(32)
counterfactual_Energy_intensity = np.zeros(32)
counterfactual_Low_carbon_penetration = np.zeros(32)
counterfactual_Low_carbon_energy = np.zeros(32)
counterfactual_Final_energy = np.zeros(32)
counterfactual_FF_energy = np.zeros(32)
counterfactual_Carbon_intensity_FF = np.zeros(32)
counterfactual_GDP_per_capita = np.zeros(32)
counterfactual_Population = np.zeros(32)
#-----COUNTERFACTUAL SCENARIO IS THE SAME AS THE HISTORICAL SCENARIO BEFORE THE START YEAR-----
counterfactual_Energy_intensity[0:start] = Energy_intensity[0:start]
counterfactual_Final_energy[0:start] = Final_energy[0:start]
counterfactual_Low_carbon_energy[0:start] = Low_carbon_energy[0:start]
counterfactual_FF_energy[0:start] = FF_energy[0:start]
counterfactual_Low_carbon_penetration[0:start] = counterfactual_Low_carbon_energy[0:start] / counterfactual_Final_energy[0:start]
counterfactual_Carbon_intensity_FF[0:start] = Carbon_intensity_FF[0:start]
counterfactual_CO2_emissions[0:start] = CO2_emissions[0:start]
counterfactual_GDP_per_capita[0:start] = GDP_per_capita[0:start]
counterfactual_Population[0:start] = Population[0:start]
#-----IPAT EQUATION FOR THE MODELLING OF COUNTERFACTUAL SCENARIOS-----
for j in np.arange(start-1, 31, 1):
if Multiplicator_Energy_intensity == 1:
counterfactual_Energy_intensity[j+1] = Energy_intensity[j+1]
else:
counterfactual_Energy_intensity[j+1] = counterfactual_Energy_intensity[j]*(1 + 0.01*counterfactual_intensity_improvement);
if Multiplicator_low_carbon == 1:
counterfactual_Low_carbon_energy[j+1] = Low_carbon_energy[j+1]
else:
counterfactual_Low_carbon_energy[j+1] = counterfactual_Low_carbon_energy[j]*(1 + 0.01*counterfactual_low_carbon_growth);
if Multiplicator_GDP_per_capita == 1:
counterfactual_GDP_per_capita[j+1] = GDP_per_capita[j+1]
else:
counterfactual_GDP_per_capita[j+1] = counterfactual_GDP_per_capita[j]*(1 + 0.01*counterfactual_GDP_per_capita_growth);
if Multiplicator_population == 1:
counterfactual_Population[j+1] = Population[j+1]
else:
counterfactual_Population[j+1] = counterfactual_Population[j]*(1 + 0.01*counterfactual_population_growth);
if Multiplicator_Carbon_intensity == 1:
counterfactual_Carbon_intensity_FF[j+1] = Carbon_intensity_FF[j+1]
else:
counterfactual_Carbon_intensity_FF[j+1] = counterfactual_Carbon_intensity_FF[j]*(1 + 0.01*counterfactual_carbon_intensity_FF_improvement)
counterfactual_Final_energy[j+1] = counterfactual_Energy_intensity[j+1]*counterfactual_GDP_per_capita[j+1]*counterfactual_Population[j+1]
counterfactual_FF_energy[j+1] = counterfactual_Final_energy[j+1] - counterfactual_Low_carbon_energy[j+1]
counterfactual_Low_carbon_penetration[j+1] = counterfactual_Low_carbon_energy[j+1]/counterfactual_Final_energy[j+1]
counterfactual_CO2_emissions[j+1] = (counterfactual_Energy_intensity[j+1]
*(1 - counterfactual_Low_carbon_penetration[j+1])*counterfactual_Carbon_intensity_FF[j+1]
*counterfactual_GDP_per_capita[j+1]*counterfactual_Population[j+1]*10**(-9))
# # Coupling and Decoupling analysis
#
# Coupling describes how closely two select variables are connected over time. In our case, we explore the links between the GDP per capita, and the ‘impact variables’ which are the CO2 emissions and energy consumption. Hyper-coupling happens when CO2 emissions (energy consumption) grow faster than the economy. Relative decoupling is when the emissions (energy consumption) grow slower than the economy. Absolute decoupling is when CO2 emissions (energy consumption) decrease while the economy continues to grow. The three modes of (de)coupling are shown in the following figure:
#
# 
# ## Decomposition analysis
#
# CO2 emissions can be related to the product of emissions drivers, as described Equations 1‒4. A change in emissions then depends on the change of emissions drivers. To find how much of the change in emissions can be attributed to each of the drivers, we need to perform the decomposition analysis, as shown in Equation 5.
#
# $$ ΔCO_{2}= dCI' + dEI' + dGDP' \; \; \; (8) $$
#
# Decomposition analysis requires some knowledge of linear algebra which we will not explain here in detail. For those who are interested, we decompose the emissions between the drivers from Equation 2 using the Shapley value decomposition approach (Ang et al., 2003), which is a commonly used method in simple decomposition analysis studies (<NAME> and Moutinho, 2013; Štreimikiene and Balezentis, 2016). For the sake of simplicity, we aggregate the drivers of population and GDPpc into GDP.
#
#-----DECOMPOSITION FACTORS IN A COUNTERFACTUAL SCENARIO------
Carbon_intensity_counterfactual_tot = counterfactual_CO2_emissions/counterfactual_Final_energy;
GDP_counterfactual = counterfactual_GDP_per_capita*counterfactual_Population;
Energy_intensity_counterfactual = counterfactual_Final_energy/GDP_counterfactual;
# +
#-----ANNUAL GROWTH RATES-----
Growth_GDP_historical = 100*(GDP_per_capita[1:]/GDP_per_capita[:-1] - 1)
Growth_Energy_historical = 100*(Final_energy[1:]/Final_energy[:-1] - 1)
Growth_CO2_historical = 100*(CO2_emissions[1:]/CO2_emissions[:-1] - 1)
Growth_GDP_counterfactual = 100*(counterfactual_GDP_per_capita[1:]/counterfactual_GDP_per_capita[:-1] - 1)
Growth_Energy_counterfactual = 100*(counterfactual_Final_energy[1:]/counterfactual_Final_energy[:-1] - 1)
Growth_CO2_counterfactual = 100*(counterfactual_CO2_emissions[1:]/counterfactual_CO2_emissions[:-1] - 1)
# +
#-----DIFFERENCE IN TOTAL ANTROPOGENIC EMISSIONS BETWEEN REAL HISTORY AND THE COUNTERFACTUAL SCENARIO-----
Diff_counterfactual = np.sum((CO2_emissions[start-1:] - counterfactual_CO2_emissions[start-1:]))
#-----1.5 DEGREE COMPATIBLE PATHWAY FROM 2021 UNDER THE HISTORICAL SCENARIO-----
Emissions_15 = [CO2_emissions[31], 1/2*CO2_emissions[31], 1/4*CO2_emissions[31], 1/8*CO2_emissions[31]] # we need to halve carbon emissions every decade
Emissions_15_interp = CubicSpline([2021, 2030, 2040, 2050], Emissions_15)(np.arange(2021, 2051, 1))
# +
#-----FINDING THE 1.5 DEGREE COMPATIBLE PATHWAY FROM 2021 UNDER THE COUNTERFACTUAL SCENARIO-----
r = np.arange(-0.15, 0.01, 0.001)
aux = np.zeros(160);
for t in np.arange(0, 160, 1):
aux[t] = sum(counterfactual_CO2_emissions[31]*(1+r[t])**np.arange(1, 60, 1)) - np.sum(Emissions_15_interp) - Diff_counterfactual
minVal = np.min(np.abs(aux))
i = np.where(np.abs(aux) == minVal)[0][0]
counterfactual_Emissions_15 = counterfactual_CO2_emissions[31]*(1+r[i])**np.arange(0, 60, 1)
# -
# # Model Outputs
#
# The implementation of the model in the Jupyter Notebook offers the following tools for analysis and data visualisations of the modelling outputs:
#
# a) Visualisation of emissions pathways and data on the average mitigation rates in the scenarios.
#
# b) Visualisation of emissions factors of the historical data and the counterfactual scenario.
#
# c) The map of the decoupling space.
#
# d) Decomposition analysis for the three emissions factors described in Equation 3.
#
# # a) Emissions pathways and neccessary emissions reductions
#
# Here you can compare the historal pathway and the counterfactual scenario with regards to annual emissions reductions from 2022 onwards that are neccessary to stay stabilize global warming below 1.5 degrees (dashed line in the figure).
100*((pd.Series(counterfactual_CO2_emissions)[31]/pd.Series(counterfactual_CO2_emissions)[start])**(1/(31-start))-1) # Average annual change in emissions in the counterfactual scenario from the start year to 2021
100*r[i] # Neccessary average change in the emissions in the counterfactual scenario from 2022
100*((pd.Series(Emissions_15_interp)[29]/pd.Series(CO2_emissions)[29])**(1/29)-1) # Neccessary average change in the emissions in the historical pathway from 2022
plt.plot(year1, CO2_emissions, label='Historical emissions', color = [0.337, 0.706, 0.914])
plt.plot(year1, counterfactual_CO2_emissions, label='Counterfactual scenario', color = [204/255, 121/255, 167/255])
plt.plot(year2, Emissions_15_interp, label='Current 1.5 degree pathway', linestyle = '--', color = [0.337, 0.706, 0.914])
plt.plot(year2, counterfactual_Emissions_15[0:30], label='Countrafactual 1.5 degree pathway', linestyle = '--', color = [204/255, 121/255, 167/255])
plt.xlabel('Year')
plt.ylabel('CO2 emissions (GtCO2)')
plt.title('Emissions Scenarios')
plt.legend()
# # b) Emissions Factors
#
# Here you can see the emissions factors in the historical dataset and compare it to the counterfactual scenario.
# +
#-----EMISSIONS FACTORS AND AUXILIARY VARIABLES-----
fig, axs = plt.subplots(3, 2, figsize=(10, 10))
axs = [item for sublist in axs for item in sublist]
plot_vars = [
(Population / 10**9, counterfactual_Population / 10**9, '(Billion)', 'Population'),
(Low_carbon_energy/ 10**9, counterfactual_Low_carbon_energy/ 10**9, 'Final energy (EJ)', 'Low-carbon energy'),
(Low_carbon_penetration, 100*counterfactual_Low_carbon_penetration, '(%)', 'Low-carbon energy share'),
(Final_energy/10**9, counterfactual_Final_energy/10**9, 'Final energy (EJ)', 'Energy consumption'),
(GDP_per_capita, counterfactual_GDP_per_capita, 'GDP per capita ($)', 'Affluence'),
(Carbon_intensity_FF, counterfactual_Carbon_intensity_FF, '(GtCO2/GJ)', 'Carbon intensity of fossil fuels'),
]
for ax, (x1, x2, label, title) in zip(axs, plot_vars):
ax.plot(year1, x1, label='Historical scenario')
ax.plot(year1, x2, label='Counterfactual scenario')
ax.legend()
ax.set_title(title);
ax.set_ylabel(label);
ax.set_xlabel('Year');
plt.tight_layout()
# -
# # Decoupling
#
# See if your assumptions for the counterfactual scenario imply a relative or an absolute decoupling between the GDP and emissions, and the GDP and energy.
# +
#-----DECOUPLING ANALYSIS-----
#-----Annual growth rates-----
Growth_GDP_historical = 100*(GDP_per_capita[1:]/GDP_per_capita[:-1] - 1)
Growth_Energy_historical = 100*(Final_energy[1:]/Final_energy[:-1] - 1)
Growth_CO2_historical = 100*(CO2_emissions[1:]/CO2_emissions[:-1] - 1)
Growth_GDP_counterfactual = 100*(counterfactual_GDP_per_capita[1:]/counterfactual_GDP_per_capita[:-1] - 1)
Growth_Energy_counterfactual = 100*(counterfactual_Final_energy[1:]/counterfactual_Final_energy[:-1] - 1)
Growth_CO2_counterfactual = 100*(counterfactual_CO2_emissions[1:]/counterfactual_CO2_emissions[:-1] - 1)
fig, axs = plt.subplots(2, 2, figsize=(10, 10))
axs = [item for sublist in axs for item in sublist]
plot_vars = [
(Growth_GDP_historical, Growth_Energy_historical, 'GDP per capita growth', 'Energy growth', 'Energy-GDP decoupling (historical)'),
(Growth_GDP_counterfactual, Growth_Energy_counterfactual, 'GDP per capita growth', 'Energy growth', 'Energy-GDP decoupling (counterfactual)'),
(Growth_GDP_historical, Growth_CO2_historical, 'GDP per capita growth', 'Emissions growth', 'Emissions-GDP decoupling (historical)'),
(Growth_GDP_counterfactual, Growth_CO2_counterfactual, 'GDP per capita growth', 'Emissions growth', 'Emissions-GDP decoupling (counterfactual)'),
]
for ax, (x1, x2, label1, label2, title) in zip(axs, plot_vars):
ax.plot(x1, x2, label=title, marker="o", linestyle="-")
ax.plot(np.arange(-6.0, 6.4, 0.2), np.arange(-6.0, 6.4, 0.2), color = ([0, 0, 0]));
ax.plot(np.arange(-6.0, 6.4, 0.2), np.zeros(62), color = ([0, 0, 0]), linestyle="--");
ax.plot(np.zeros(60), np.arange(-6.0, 6.0, 0.2), color = ([0, 0, 0]), linestyle="--");
ax.legend()
ax.set_title(title);
ax.set_ylabel(label2);
ax.set_xlabel(label1);
plt.tight_layout()
# -
# # Decomposition
#
# Find which emissions factors define the counterfactual emissions pathway from 1990 to 2021. Find if your mitigation plan and/or a different socio-economic development have succeed at holding the emissions in check!
# +
Ci_driver = np.zeros(32)
Energy_driver = np.zeros(32)
GDP_driver = np.zeros(32)
Ci_driver_counterfactual = np.zeros(32)
Energy_driver_counterfactual = np.zeros(32)
GDP_driver_counterfactual = np.zeros(32)
Carbon_intensity_tot_counterfactual = counterfactual_CO2_emissions/counterfactual_Final_energy*10**9
GDP_counterfactual = counterfactual_GDP_per_capita*counterfactual_Population
Energy_intensity_counterfactual = counterfactual_Final_energy/GDP_counterfactual
for t in np.arange(0, 31, 1):
Ci_driver[t+1] = 1/3*(Carbon_intensity_tot[t+1]*Energy_intensity[t]*GDP[t] - Carbon_intensity_tot[t]*Energy_intensity[t]*GDP[t]) + 1/6*(Carbon_intensity_tot[t+1]*Energy_intensity[t+1]*GDP[t] - Carbon_intensity_tot[t]*Energy_intensity[t+1]*GDP[t] + Carbon_intensity_tot[t+1]*Energy_intensity[t]*GDP[t+1] - Carbon_intensity_tot[t]*Energy_intensity[t]*GDP[t+1]) + 1/3*(Carbon_intensity_tot[t+1]*Energy_intensity[t+1]*GDP[t+1] - Carbon_intensity_tot[t]*Energy_intensity[t+1]*GDP[t+1])
Energy_driver[t+1] = 1/3*(Carbon_intensity_tot[t]*Energy_intensity[t+1]*GDP[t] - Carbon_intensity_tot[t]*Energy_intensity[t]*GDP[t]) + 1/6*(Carbon_intensity_tot[t+1]*Energy_intensity[t+1]*GDP[t] - Carbon_intensity_tot[t+1]*Energy_intensity[t]*GDP[t] + Carbon_intensity_tot[t]*Energy_intensity[t+1]*GDP[t+1] - Carbon_intensity_tot[t]*Energy_intensity[t]*GDP[t+1]) + 1/3*(Carbon_intensity_tot[t+1]*Energy_intensity[t+1]*GDP[t+1] - Carbon_intensity_tot[t+1]*Energy_intensity[t]*GDP[t+1])
GDP_driver[t+1] = 1/3*(Carbon_intensity_tot[t]*Energy_intensity[t]*GDP[t+1] - Carbon_intensity_tot[t]*Energy_intensity[t]*GDP[t]) + 1/6*(Carbon_intensity_tot[t+1]*Energy_intensity[t]*GDP[t+1] - Carbon_intensity_tot[t+1]*Energy_intensity[t]*GDP[t] + Carbon_intensity_tot[t]*Energy_intensity[t+1]*GDP[t+1] - Carbon_intensity_tot[t]*Energy_intensity[t+1]*GDP[t]) + 1/3*(Carbon_intensity_tot[t+1]*Energy_intensity[t+1]*GDP[t+1] - Carbon_intensity_tot[t+1]*Energy_intensity[t+1]*GDP[t])
Ci_driver_counterfactual[t+1] = 1/3*(Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t] - Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t]) + 1/6*(Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t] - Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t] + Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t+1] - Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t+1]) + 1/3*(Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t+1] - Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t+1])
Energy_driver_counterfactual[t+1] = 1/3*(Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t] - Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t]) + 1/6*(Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t] - Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t] + Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t+1] - Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t+1]) + 1/3*(Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t+1] - Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t+1])
GDP_driver_counterfactual[t+1] = 1/3*(Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t+1] - Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t]) + 1/6*(Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t+1] - Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t]*GDP_counterfactual[t] + Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t+1] - Carbon_intensity_tot_counterfactual[t]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t]) + 1/3*(Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t+1] - Carbon_intensity_tot_counterfactual[t+1]*Energy_intensity_counterfactual[t+1]*GDP_counterfactual[t])
Carbon_Intensity_contribution = np.sum(np.arange(32,0,-1)*Ci_driver)/10**9
Energy_Intensity_contribution = np.sum(np.arange(32,0,-1)*Energy_driver)/10**9
GDP_contribution = np.sum(np.arange(32,0,-1)*GDP_driver)/10**9
Norm_start = 31*CO2_emissions[start-1]
Net_change = np.sum(100*[GDP_contribution/Norm_start, Carbon_Intensity_contribution/Norm_start, Energy_Intensity_contribution/Norm_start])
Net_sum = GDP_contribution + Carbon_Intensity_contribution + Energy_Intensity_contribution
Carbon_Intensity_contribution_counterfactual = np.sum(np.arange(32,0,-1)*Ci_driver_counterfactual)/10**9
Energy_Intensity_contribution_counterfactual = np.sum(np.arange(32,0,-1)*Energy_driver_counterfactual)/10**9
GDP_contribution_counterfactual = np.sum(np.arange(32,0,-1)*GDP_driver_counterfactual)/10**9
Norm_start_counterfactual = 31*counterfactual_CO2_emissions[start]
Net_change_counterfactual = np.sum(100*[GDP_contribution_counterfactual/Norm_start_counterfactual, Carbon_Intensity_contribution_counterfactual/Norm_start_counterfactual, Energy_Intensity_contribution_counterfactual/Norm_start_counterfactual])
Net_sum_counterfactual = GDP_contribution_counterfactual + Carbon_Intensity_contribution_counterfactual + Energy_Intensity_contribution_counterfactual
df = pd.DataFrame(index=['Historical', 'Counterfactual'],
data={'GDP per-capita': [GDP_contribution, GDP_contribution_counterfactual],
'Carbon intensity': [Carbon_Intensity_contribution, Carbon_Intensity_contribution_counterfactual],
'Energy intensity': [Energy_Intensity_contribution, Energy_Intensity_contribution_counterfactual]})
ax = df.plot(kind="bar", stacked=True)
ax.set_title('Emissions Drivers');
ax.set_ylabel('Cumulative emissions (GtCO2)');
plt.hlines(GDP_contribution + Carbon_Intensity_contribution + Energy_Intensity_contribution , xmin=-0.25, xmax=0.25, color = ([0, 0, 0]))
plt.hlines(GDP_contribution_counterfactual + Carbon_Intensity_contribution_counterfactual + Energy_Intensity_contribution_counterfactual , xmin=0.75, xmax=1.25, color = ([0, 0, 0]))
plt.show()
df = pd.DataFrame(index=['Historical', 'Counterfactual'],
data={'GDP per-capita': [GDP_contribution/Norm_start, GDP_contribution_counterfactual/Norm_start_counterfactual],
'Carbon intensity': [Carbon_Intensity_contribution/Norm_start, Carbon_Intensity_contribution_counterfactual/Norm_start_counterfactual],
'Energy intensity': [Energy_Intensity_contribution/Norm_start, Energy_Intensity_contribution_counterfactual/Norm_start_counterfactual]})
ax = df.plot(kind="bar", stacked=True)
ax.set_title('Emissions Drivers');
ax.set_ylabel('Cumulative emissions (%)');
plt.hlines(GDP_contribution/Norm_start + Carbon_Intensity_contribution/Norm_start + Energy_Intensity_contribution/Norm_start , xmin=-0.25, xmax=0.25, color = ([0, 0, 0]))
plt.hlines(GDP_contribution_counterfactual/Norm_start_counterfactual + Carbon_Intensity_contribution_counterfactual/Norm_start_counterfactual + Energy_Intensity_contribution_counterfactual/Norm_start_counterfactual , xmin=0.75, xmax=1.25, color = ([0, 0, 0]))
plt.show()
# -
# # Literature
#
# 1. <NAME>., <NAME>., <NAME>., 2003. Perfect decomposition techniques in energy and environmental analysis. Energy Policy 31, 1561–1566. https://doi.org/10.1016/S0301-4215(02)00206-9
#
# 2. IPCC, 2018. Special Report on Global Warming of 1.5 C, Global Warming of 1.5 °C: An IPCC Special Report on the impacts of global warming of 1.5°C above pre-industrial levels and related global greenhouse gas emission pathways, in the context of strengthening the global response to the threat of climate change.
#
# 3. <NAME>., 1990. Impact of Carbon Dioxide emission control on GNP growth: Interpretation of proposed scenarios, IPCC Energy and Industry Subgroup, Response Strategies Working Group.
#
# 4. <NAME>., <NAME>., 2013. Decomposition analysis and Innovative Accounting Approach for energy-related CO2 (carbon dioxide) emissions intensity over 1996-2009 in Portugal. Energy 57, 775–787. https://doi.org/10.1016/j.energy.2013.05.036
#
# 5. <NAME>., <NAME>., 2016. Kaya identity for analysis of the main drivers of GHG emissions and feasibility to implement EU “20-20-20” targets in the Baltic States. Renew. Sustain. Energy Rev. 58, 1108–1113. https://doi.org/10.1016/j.rser.2015.12.311
#
#
|
03_Counterfactual_Scenarios.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 0.0. IMPORTS
# +
import math
import numpy as np
import pandas as pd
import inflection
import seaborn as sns
from matplotlib import pyplot as plt
from IPython.core.display import HTML
from IPython.display import Image
# + [markdown] heading_collapsed=true
# ## 0.1. Helper Functions
# + hidden=true
def jupyter_settings():
# %matplotlib inline
# %pylab inline
plt.style.use( 'bmh' )
plt.rcParams['figure.figsize'] = [25, 12]
plt.rcParams['font.size'] = 24
display( HTML( '<style>.container { width:100% !important; }</style>') )
pd.options.display.max_columns = None
pd.options.display.max_rows = None
pd.set_option( 'display.expand_frame_repr', False )
sns.set()
# + hidden=true
jupyter_settings()
# + [markdown] heading_collapsed=true
# ## 0.2. Loading data
# + hidden=true
df_sales_raw = pd.read_csv( '../data/train.csv', low_memory=False )
df_store_raw = pd.read_csv( '../data/store.csv', low_memory=False )
# merge
df_raw = pd.merge( df_sales_raw, df_store_raw, how='left', on='Store' )
# + [markdown] heading_collapsed=true
# # 1.0. PASSO 01 - DESCRICAO DOS DADOS
# + hidden=true
df1 = df_raw.copy()
# + [markdown] heading_collapsed=true hidden=true
# ## 1.1. Rename Columns
# + hidden=true
cols_old = ['Store', 'DayOfWeek', 'Date', 'Sales', 'Customers', 'Open', 'Promo', 'StateHoliday', 'SchoolHoliday',
'StoreType', 'Assortment', 'CompetitionDistance', 'CompetitionOpenSinceMonth',
'CompetitionOpenSinceYear', 'Promo2', 'Promo2SinceWeek', 'Promo2SinceYear', 'PromoInterval']
snakecase = lambda x: inflection.underscore( x )
cols_new = list( map( snakecase, cols_old ) )
# rename
df1.columns = cols_new
# + [markdown] heading_collapsed=true hidden=true
# ## 1.2. Data Dimensions
# + hidden=true
print( 'Number of Rows: {}'.format( df1.shape[0] ) )
print( 'Number of Cols: {}'.format( df1.shape[1] ) )
# + [markdown] heading_collapsed=true hidden=true
# ## 1.3. Data Types
# + hidden=true
df1['date'] = pd.to_datetime( df1['date'] )
df1.dtypes
# + [markdown] heading_collapsed=true hidden=true
# ## 1.4. Check NA
# + hidden=true
df1.isna().sum()
# + [markdown] heading_collapsed=true hidden=true
# ## 1.5. Fillout NA
# + hidden=true
df1.sample()
# + hidden=true
#competition_distance
df1['competition_distance'] = df1['competition_distance'].apply( lambda x: 200000.0 if math.isnan( x ) else x )
#competition_open_since_month
df1['competition_open_since_month'] = df1.apply( lambda x: x['date'].month if math.isnan( x['competition_open_since_month'] ) else x['competition_open_since_month'], axis=1 )
#competition_open_since_year
df1['competition_open_since_year'] = df1.apply( lambda x: x['date'].year if math.isnan( x['competition_open_since_year'] ) else x['competition_open_since_year'], axis=1 )
#promo2_since_week
df1['promo2_since_week'] = df1.apply( lambda x: x['date'].week if math.isnan( x['promo2_since_week'] ) else x['promo2_since_week'], axis=1 )
#promo2_since_year
df1['promo2_since_year'] = df1.apply( lambda x: x['date'].year if math.isnan( x['promo2_since_year'] ) else x['promo2_since_year'], axis=1 )
#promo_interval
month_map = {1: 'Jan', 2: 'Fev', 3: 'Mar', 4: 'Apr', 5: 'May', 6: 'Jun', 7: 'Jul', 8: 'Aug', 9: 'Sep', 10: 'Oct', 11: 'Nov', 12: 'Dec'}
df1['promo_interval'].fillna(0, inplace=True )
df1['month_map'] = df1['date'].dt.month.map( month_map )
df1['is_promo'] = df1[['promo_interval', 'month_map']].apply( lambda x: 0 if x['promo_interval'] == 0 else 1 if x['month_map'] in x['promo_interval'].split( ',' ) else 0, axis=1 )
# + hidden=true
df1.isna().sum()
# + [markdown] heading_collapsed=true hidden=true
# ## 1.6. Change Data Types
# + hidden=true
# competiton
df1['competition_open_since_month'] = df1['competition_open_since_month'].astype( int )
df1['competition_open_since_year'] = df1['competition_open_since_year'].astype( int )
# promo2
df1['promo2_since_week'] = df1['promo2_since_week'].astype( int )
df1['promo2_since_year'] = df1['promo2_since_year'].astype( int )
# + [markdown] heading_collapsed=true hidden=true
# ## 1.7. Descriptive Statistics
# + hidden=true
num_attributes = df1.select_dtypes( include=['int64', 'float64'] )
cat_attributes = df1.select_dtypes( exclude=['int64', 'float64', 'datetime64[ns]'] )
# + [markdown] heading_collapsed=true hidden=true
# ### 1.7.1. Numerical Atributes
# + hidden=true
# Central Tendency - mean, meadina
ct1 = pd.DataFrame( num_attributes.apply( np.mean ) ).T
ct2 = pd.DataFrame( num_attributes.apply( np.median ) ).T
# dispersion - std, min, max, range, skew, kurtosis
d1 = pd.DataFrame( num_attributes.apply( np.std ) ).T
d2 = pd.DataFrame( num_attributes.apply( min ) ).T
d3 = pd.DataFrame( num_attributes.apply( max ) ).T
d4 = pd.DataFrame( num_attributes.apply( lambda x: x.max() - x.min() ) ).T
d5 = pd.DataFrame( num_attributes.apply( lambda x: x.skew() ) ).T
d6 = pd.DataFrame( num_attributes.apply( lambda x: x.kurtosis() ) ).T
# concatenar
m = pd.concat( [d2, d3, d4, ct1, ct2, d1, d5, d6] ).T.reset_index()
m.columns = ['attributes', 'min', 'max', 'range', 'mean', 'median', 'std', 'skew', 'kurtosis']
m
# + hidden=true
sns.distplot( df1['competition_distance'], kde=False )
# + [markdown] heading_collapsed=true hidden=true
# ### 1.7.2. Categorical Atributes
# + hidden=true
cat_attributes.apply( lambda x: x.unique().shape[0] )
# + hidden=true
aux = df1[(df1['state_holiday'] != '0') & (df1['sales'] > 0)]
plt.subplot( 1, 3, 1 )
sns.boxplot( x='state_holiday', y='sales', data=aux )
plt.subplot( 1, 3, 2 )
sns.boxplot( x='store_type', y='sales', data=aux )
plt.subplot( 1, 3, 3 )
sns.boxplot( x='assortment', y='sales', data=aux )
# -
# # 2.0. PASSO 02 - FEATURE ENGINEERING
df2 = df1.copy()
# + [markdown] heading_collapsed=true
# ## 2.1. Mapa Mental de Hipoteses
# + hidden=true
Image( 'img/MindMapHypothesis.png' )
# + [markdown] heading_collapsed=true
# ## 2.2. Criacao das Hipoteses
# + [markdown] heading_collapsed=true hidden=true
# ### 2.2.1. Hipoteses Loja
# + [markdown] hidden=true
# **1.** Lojas com número maior de funcionários deveriam vender mais.
#
# **2.** Lojas com maior capacidade de estoque deveriam vender mais.
#
# **3.** Lojas com maior porte deveriam vender mais.
#
# **4.** Lojas com maior sortimentos deveriam vender mais.
#
# **5.** Lojas com competidores mais próximos deveriam vender menos.
#
# **6.** Lojas com competidores à mais tempo deveriam vendem mais.
# + [markdown] heading_collapsed=true hidden=true
# ### 2.2.2. Hipoteses Produto
# + [markdown] hidden=true
# **1.** Lojas que investem mais em Marketing deveriam vender mais.
#
# **2.** Lojas com maior exposição de produto deveriam vender mais.
#
# **3.** Lojas com produtos com preço menor deveriam vender mais.
#
# **5.** Lojas com promoções mais agressivas ( descontos maiores ), deveriam vender mais.
#
# **6.** Lojas com promoções ativas por mais tempo deveriam vender mais.
#
# **7.** Lojas com mais dias de promoção deveriam vender mais.
#
# **8.** Lojas com mais promoções consecutivas deveriam vender mais.
# + [markdown] heading_collapsed=true hidden=true
# ### 2.2.3. Hipoteses Tempo
# + [markdown] hidden=true
# **1.** Lojas abertas durante o feriado de Natal deveriam vender mais.
#
# **2.** Lojas deveriam vender mais ao longo dos anos.
#
# **3.** Lojas deveriam vender mais no segundo semestre do ano.
#
# **4.** Lojas deveriam vender mais depois do dia 10 de cada mês.
#
# **5.** Lojas deveriam vender menos aos finais de semana.
#
# **6.** Lojas deveriam vender menos durante os feriados escolares.
# + [markdown] heading_collapsed=true
# ## 2.3. Lista Final de Hipóteses
# + [markdown] hidden=true
# **1.** Lojas com maior sortimentos deveriam vender mais.
#
# **2.** Lojas com competidores mais próximos deveriam vender menos.
#
# **3.** Lojas com competidores à mais tempo deveriam vendem mais.
# + [markdown] hidden=true
# **4.** Lojas com promoções ativas por mais tempo deveriam vender mais.
#
# **5.** Lojas com mais dias de promoção deveriam vender mais.
#
# **7.** Lojas com mais promoções consecutivas deveriam vender mais.
# + [markdown] hidden=true
# **8.** Lojas abertas durante o feriado de Natal deveriam vender mais.
#
# **9.** Lojas deveriam vender mais ao longo dos anos.
#
# **10.** Lojas deveriam vender mais no segundo semestre do ano.
#
# **11.** Lojas deveriam vender mais depois do dia 10 de cada mês.
#
# **12.** Lojas deveriam vender menos aos finais de semana.
#
# **13.** Lojas deveriam vender menos durante os feriados escolares.
#
# + [markdown] heading_collapsed=true
# ## 2.4. Feature Engineering
# + hidden=true
# year
df2['year'] = df2['date'].dt.year
# month
df2['month'] = df2['date'].dt.month
# day
df2['day'] = df2['date'].dt.day
# week of year
df2['week_of_year'] = df2['date'].dt.weekofyear
# year week
df2['year_week'] = df2['date'].dt.strftime( '%Y-%W' )
# competition since
df2['competition_since'] = df2.apply( lambda x: datetime.datetime( year=x['competition_open_since_year'], month=x['competition_open_since_month'],day=1 ), axis=1 )
df2['competition_time_month'] = ( ( df2['date'] - df2['competition_since'] )/30 ).apply( lambda x: x.days ).astype( int )
# promo since
df2['promo_since'] = df2['promo2_since_year'].astype( str ) + '-' + df2['promo2_since_week'].astype( str )
df2['promo_since'] = df2['promo_since'].apply( lambda x: datetime.datetime.strptime( x + '-1', '%Y-%W-%w' ) - datetime.timedelta( days=7 ) )
df2['promo_time_week'] = ( ( df2['date'] - df2['promo_since'] )/7 ).apply( lambda x: x.days ).astype( int )
# assortment
df2['assortment'] = df2['assortment'].apply( lambda x: 'basic' if x == 'a' else 'extra' if x == 'b' else 'extended' )
# state holiday
df2['state_holiday'] = df2['state_holiday'].apply( lambda x: 'public_holiday' if x == 'a' else 'easter_holiday' if x == 'b' else 'christmas' if x == 'c' else 'regular_day' )
# -
# # 3.0. PASSO 03 - FILTRAGEM DE VARIÁVEIS
# Check Point
df3 = df2.copy()
# ## 3.1. Filtragem das Linhas
#
#Filtramos as linhas que não possuem informações de vendas - Open diferente de zero e Sales Maior que zero
df3 = df3[(df3['open'] != 0) & (df3['sales'] > 0)]
# ## 3.2. Selecao das Colunas
# Dropamos as colunas que não temos no momento da predição(planejamento)
cols_drop = ['customers', 'open', 'promo_interval', 'month_map']
df3 = df3.drop( cols_drop, axis=1 )
|
m03_v01_store_sales_prediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="VhYTQjkrIU_0"
# !wget https://data.vision.ee.ethz.ch/cvl/clic/mobile_train_2020.zip
# !unzip mobile_train_2020.zip
# !wget https://data.vision.ee.ethz.ch/cvl/clic/mobile_valid_2020.zip
# !unzip mobile_valid_2020.zip
# !wget http://data.vision.ee.ethz.ch/cvl/DIV2K/DIV2K_train_HR.zip
# !unzip DIV2K_train_HR.zip
# !wget http://data.vision.ee.ethz.ch/cvl/DIV2K/DIV2K_valid_HR.zip
# !unzip DIV2K_valid_HR.zip
# + id="ykCHhRiKgcy3" outputId="5e337b42-9d05-422f-b699-4aa57121a64d" colab={"base_uri": "https://localhost:8080/"}
import glob
import numpy as np
import cv2
# %tensorflow_version 1.x
import tensorflow as tf
tf.compat.v1.graph_util.extract_sub_graph
import keras
import os
x_valid = np.empty((161,512,768,3)).astype(np.float32)
count = 0
for image_path in glob.glob("../content/valid/*.png"):
image = cv2.imread(image_path).astype(np.float32)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
if (image.shape[0] >= 512 and image.shape[1] >=768):
image = image[0:0+512, 0:0+768]
x_valid[count] = image/255
count+=1
for image_path in glob.glob("../content/DIV2K_valid_HR/*.png"):
image = cv2.imread(image_path).astype(np.float32)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
if (image.shape[0] >= 512 and image.shape[1] >=768):
image = image[0:0+512, 0:0+768]
x_valid[count] = image/255
count+=1
x_train = np.empty((1832,512,768,3)).astype(np.float32)
count = 0
for image_path in glob.glob("../content/train/*.png"):
image = cv2.imread(image_path).astype(np.float32)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
if (image.shape[0] >= 512 and image.shape[1] >= 768):
image = image[0:0+512, 0:0+768]
x_train[count] = image/255
count+=1
for image_path in glob.glob("../content/DIV2K_train_HR/*.png"):
image = cv2.imread(image_path).astype(np.float32)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
if (image.shape[0] >= 512 and image.shape[1] >= 768):
image = image[0:0+512, 0:0+768]
x_train[count] = image/255
count+=1
# + id="Z0jIc35dICFC" outputId="780315ae-38f4-4cd1-8c45-3ec2f654a1d8" colab={"base_uri": "https://localhost:8080/"}
tf.compat.v1.graph_util.extract_sub_graph
from keras.layers import Input, Conv2D, Conv2DTranspose, MaxPooling2D, UpSampling2D
from keras.models import Model
from keras.callbacks import ModelCheckpoint
input_img = Input(shape=(512, 768, 3))
x = Conv2D(32, [3,3], strides=(1,1), activation="relu")(input_img)
x = Conv2D(32, [3,3], strides=(2,2), activation="relu")(x)
x = Conv2D(64, [3,3], strides=(1,1), activation="relu")(x)
x = Conv2D(64, [3,3], strides=(2,2), activation="relu")(x)
encoded = Conv2D(32, [3,3], strides=(2,2), activation="relu")(x)
encoder = Model(input_img, encoded)
latentInputs = Input(shape=(62, 94, 32))
y = Conv2DTranspose(32, [3,3], strides= (2,2), activation = "relu")(latentInputs)
y = Conv2DTranspose(64, [3,3], strides= (2,2), activation = "relu")(y)
y = Conv2DTranspose(64, [3,3], strides= (1,1), activation = "relu")(y)
y = Conv2DTranspose(32, [3,3], strides= (2,2), activation = "relu")(y)
y = Conv2DTranspose(32, [3,3], strides= (1,1), activation = "relu")(y)
decoded = Conv2DTranspose(3, [4,4], activation = "relu")(y)
decoder = Model(latentInputs, decoded)
autoencoder = Model(input_img, decoder(encoder(input_img)))
#autoencoder = Model(input_img, decoded)
autoencoder.compile(optimizer='adam', loss='mse', metrics=['accuracy'])
encoder.compile(optimizer='adam', loss='mse', metrics=['accuracy'])
# + id="RJ2Y_jhSaER8" outputId="66af13c2-7e52-46bf-bd02-52f033a88348" colab={"base_uri": "https://localhost:8080/"}
print(encoder.summary())
print(decoder.summary())
print(autoencoder.summary())
# + id="kMNcYSw9Z_gW" outputId="a213bc31-9906-43a8-8c71-6fafd0ea1c7c" colab={"base_uri": "https://localhost:8080/"}
filepath="k_model.h5"
checkpoint = ModelCheckpoint(filepath, monitor='val_accuracy', verbose=1, save_best_only=True)
callbacks_list = [checkpoint]
H = autoencoder.fit(x_train, x_train,
epochs=25,
batch_size=16,
shuffle=True,
validation_data=(x_valid, x_valid),
callbacks = callbacks_list)
# + id="RYHHPEDHoOlL"
a = []
while(1):
a.append("1")
# + id="t1_QwSGoTJ9f"
encoder.save("encoder_weights.h5")
decoder.save("decoder_weights.h5")
# + id="grNThX3eVRXq" outputId="117d25b2-c18a-41f5-dee8-503c2b21a153" colab={"base_uri": "https://localhost:8080/", "height": 296}
import matplotlib.pyplot as plt
N = np.arange(0, 50)
plt.style.use("seaborn-colorblind")
plt.figure()
plt.plot(N, H.history["loss"], label="train_loss")
plt.plot(N, H.history["val_loss"], label="val_loss")
#plt.title("Training and validation loss")
plt.xlabel("Epoch #")
plt.ylabel("Loss")
plt.legend(loc="lower left")
# + id="UHGWAWcNVq-k" outputId="eeea9b56-5887-47cc-9995-b63e27deb533" colab={"base_uri": "https://localhost:8080/", "height": 296}
plt.figure()
plt.plot(N, H.history["accuracy"], label="train_accuracy")
plt.plot(N, H.history["val_accuracy"], label="val_accuracy")
#plt.title("Training and validation accuracy")
plt.xlabel("Epoch #")
plt.ylabel("Accuracy")
plt.legend(loc="lower left")
# + id="52W3Lm7ls93T" outputId="eab436c4-8062-4e7e-8c61-8dcbc3fe4ae8" colab={"base_uri": "https://localhost:8080/", "height": 54}
print(H.history)
# + id="eU14EHWDmSPi" outputId="442d4850-efb0-47f0-86b7-6de4c773ac79" colab={"base_uri": "https://localhost:8080/", "height": 269}
import imageio
import glob
kodak = np.empty((18, 512, 768, 3)).astype(np.float32)
count = 0
for image_path in glob.glob("../content/*.png"):
image = cv2.imread(image_path).astype(np.float32)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = image[0:0+512, 0:0+768]
kodak[count] = image/255
plt.imshow(kodak[count])
count+=1
# + id="ikzKMmABZzgU"
decoded_imgs = autoencoder.predict(kodak)
# + id="XFYo_OMFd2s7" outputId="c22cd271-87f5-4cd3-e5e8-a43eee041d7e" colab={"base_uri": "https://localhost:8080/", "height": 768}
from skimage.measure import compare_ssim as ssim
from skimage.measure import compare_psnr as psnr
avg_ssim = []
avg_psnr = []
for i in range(18):
for k in range(10):
if k==0:
sum_psnr = psnr(kodak[i], decoded_imgs[i], data_range=None)
else:
sum_psnr = sum_psnr + psnr(kodak[i], decoded_imgs[i], data_range=None)
avg_psnr.append(sum_psnr/10)
print(avg_psnr[i])
print("ssim")
for i in range(18):
for k in range(10):
if k==0:
sum_ssim = ssim(kodak[i], decoded_imgs[i], multichannel=True)
else:
sum_ssim = sum_ssim + ssim(kodak[i], decoded_imgs[i], multichannel=True)
avg_ssim.append(sum_ssim/10)
print(avg_ssim[i])
# + id="NZPjZmDUgOgr" outputId="ae323b0b-c249-460e-9483-bd652ab0d945" colab={"base_uri": "https://localhost:8080/", "height": 555}
plt.figure(1)
plt.imshow(kodak[0])
plt.figure(2)
plt.imshow(decoded_imgs[0])
|
CAE_Implementation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3 (tf-gpu)
# language: python
# name: tf-gpu
# ---
# +
import traci
import pandas as pd
import random
import time
import pickle
log = pickle.load(open('./output/bus_ids.pkl', 'rb'))
sumo_cmd = ["/opt/ohpc/pub/apps/sumo/bin/sumo"]
sumo_config = ["-c", "./Sim/network-bus-background.sumocfg"]
sumo_cmd.extend(sumo_config)
# +
start_t = time.time()
traci.start(sumo_cmd, label=str(random.randint(10000, 50000)))
while traci.simulation.getTime() <= 32400:
step = int(traci.simulation.getTime())
if step % 100 == 0:
print(step)
veh_ids = traci.vehicle.getIDList()
for veh in veh_ids:
if veh in log:
edge = traci.vehicle.getRoadID(veh)
log[veh].append({'step': step, 'speed': traci.vehicle.getSpeed(veh), 'edge': edge,
'distance': traci.vehicle.getDistance(veh), 'fuel_consumption': traci.vehicle.getFuelConsumption(veh),
'position': traci.vehicle.getPosition(veh), 'waiting_time': traci.vehicle.getWaitingTime(veh),
'accumulated_waiting_time': traci.vehicle.getAccumulatedWaitingTime(veh)})
traci.simulationStep()
finish_t = time.time()
# -
pickle.dump(log, open('output/bus_data_normal_traffic.pkl', 'wb'))
print("Simulation time: %d" % (finish_t - start_t))
|
background_traffic_elimination/log-bus-data-in-normal-simultion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
dfeda = pd.read_csv("../EDA_New.csv")
dfeda['condition'].unique()
dfeda['condition'].replace('time pressure', 'interruption', inplace=True)
dfeda['condition'].replace('interruption', 1, inplace=True)
dfeda['condition'].replace('no stress', 0, inplace=True)
a = dfeda[dfeda['condition']==0].reset_index()[5000:20001].reset_index()['SC']
sns.set_context("paper", rc={"lines.linewidth": 2.5})
sns.set_palette("binary_d")
fig, ax = plt.subplots()
ax.tick_params('x', labelbottom=False, bottom=False)
ax.tick_params()
g = sns.lineplot(data=a)
g.set_ylim(0, 0.12)
plt.ylabel('Electrodermal Activity')
plt.xlabel('Sequential Data-Points')
a = pd.concat([dfeda[dfeda['condition']==1].reset_index()['SC'][40175:40200], dfeda[dfeda['condition']==1].reset_index()['SC'][40375:40400], dfeda[dfeda['condition']==1].reset_index()['SC'][50280:50300]], axis=0)
a = a.reset_index()
plt.gca().set_xticklabels(['']*100000)
fig, ax = plt.subplots()
# ax.tick_params('x', labelbottom=False, bottom=False)
g = sns.lineplot(data=a['SC'])
g.set_ylim(0, 0.12)
g.set_xlim(0, 60)
plt.ylabel('Electrodermal Activity')
plt.xlabel('Sequential Data-Points')
plt.show()
sns.set_context("paper", rc={"lines.linewidth": 2.5})
sns.set_palette("binary_d")
fig, ax = plt.subplots()
# ax.tick_params('x', labelbottom=False, bottom=False)
g = sns.lineplot(data=dfeda[dfeda['condition']==1].reset_index().iloc[40379:40400].reset_index()['SC'])
g.set_ylim(0, 0.12)
g.set_xlim(0, 20)
plt.ylabel('Electrodermal Activity')
plt.xlabel('Sequential Data-Points')
plt.show()
|
SWELL-KW/SWELL-KW_Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h4>Hidden factors Latent models</h4>
#
#
# <p></p>
# https://blog.insightdatascience.com/explicit-matrix-factorization-als-sgd-and-all-that-jazz-b00e4d9b21ea
# <p></p>
# Notation for this note is here:
# http://yifanhu.net/PUB/cf.pdf
# <p></p>
# <p>There is a progression for applying deep learning. 1) apply to the linear situation.
# This may be more inefficient but you don't have the same problems with inverting square matricies
# as in normal solvers. You minimize the error instead. There can be issues with this approach. The bigger
# step is when y-Ax can be replaced by a nonlinearity and solve via DNN. </p>
#
# +
import os
import numpy as np
import pandas as pd
np.random.seed(0)
#get movielens dataset
#curl -O http://files.grouplens.org/datasets/movielens/ml-100k.zip
#unzip ml-100k.zip
# #cd ml-100k/
os.chdir("/Users/dc/DeepLearning/hinton/ml-100k/")
names = ['user_id', 'item_id', 'rating', 'timestamp']
df = pd.read_csv('u.data', sep='\t', names=names)
df.head()
n_users = df.user_id.unique().shape[0]
n_items = df.item_id.unique().shape[0]
ratings = np.zeros((n_users, n_items))
for row in df.itertuples():
ratings[row[1]-1, row[2]-1] = row[3]
ratings
print (str(n_users) + ' users')
print (str(n_items) + ' items')
sparsity = float(len(ratings.nonzero()[0]))
sparsity /= (ratings.shape[0] * ratings.shape[1])
sparsity *= 100
print ('Sparsity: {:4.2f}%'.format(sparsity))
def train_test_split(ratings):
test = np.zeros(ratings.shape)
train = ratings.copy()
for user in range(ratings.shape[0]):
test_ratings = np.random.choice(ratings[user, :].nonzero()[0],
size=10,
replace=False)
train[user, test_ratings] = 0.
test[user, test_ratings] = ratings[user, test_ratings]
# Test and training are truly disjoint
assert(np.all((train * test) == 0))
return train, test
train, test = train_test_split(ratings)
print("train shape:", train.shape," test.shape:",test.shape)
# -
# <h4>Matrix factorization</h4>
# <p>The non deep learning way</p>
# Assumptions:
# <p>
# <li>We have a single matrix of user/products where the rows are users and the columns are products/movies.
# A 0 indicated the user did not like the product a 1 indicates like product. </li>
# <li>Each user described with k features. A feature can be a list of actors in a movie</li>
# <li>Each item can be described a set of k features. A feature can be a list of actors</li>
# <li>If we multiply user_featurs*item_featues this is an approximation for a user rating of a movie</li>
# </p>
# We do not specify the features beforehand. We pick an integer value for k and learn the features.
# The modeling of the user/product matrix into 2 separate matrices is expressed as 2 matricies,
# u for user and p for products:
# $z_{ji} = u_j^T \cdot p_i=\sum\limits_{k=0}^N u_{jk}p_{ik}$
# <p></p>
# The u and p matricies are latent vectors and the k features are called latent factors.
# <p></p>
# OK this is all cool, where is the minimization/maximation formula so I can take a gradient?
# <p></p>
# Minimize the difference between the ragings in teh dataset and predictions.
# <p></p>
# $L = \sum\limits_{j,i}(z_{ji}-u_{j}^Tp_{i})^2$
# <p></p>
# Adding in L_2 normalization gives us:
# <p></p>
# $L = \sum\limits_{j,i}(z_{ji}-u_{j}^Tp_{i})^2 + \lambda_j {\lVert {u_j} \rVert} ^2 +\lambda_i{\lVert {p_i} \rVert}^2$
# <p></p>
# $\frac{\partial L}{\partial u_j} = 2 \sum (z_{ji}-u_j^Tp_i)\frac{\partial(z_{ui} - u_j^Tp_i)}{\partial u_j}+ 2 \lambda_j {\lVert {u_j} \rVert}$
# <p></p>
# $= 2 \sum (z_{ji}-u_j^Tp_i){ (-p_i)}+ 2 \lambda_j {\lVert {u_j} \rVert}$
# <p></p>
# $0= 2 \sum (z_{ji}-u_j^Tp_i){ (-p_i)}+ 2 \lambda_j {\lVert {u_j} \rVert}$
# <p></p>
# $0= \sum (z_{ji}-u_j^Tp_i){ (-p_i)}+ \lambda_j {\lVert {u_j} \rVert}$
# <p></p>
# <p></p>
# +
from numpy.linalg import solve
class ExplicitMF():
def __init__(self,
ratings,
n_factors=40,
item_reg=0.0,
user_reg=0.0,
verbose=False):
"""
Train a matrix factorization model to predict empty
entries in a matrix. The terminology assumes a
ratings matrix which is ~ user x item
Params
======
ratings : (ndarray)
User x Item matrix with corresponding ratings
n_factors : (int)
Number of latent factors to use in matrix
factorization model
item_reg : (float)
Regularization term for item latent factors
user_reg : (float)
Regularization term for user latent factors
verbose : (bool)
Whether or not to printout training progress
"""
self.ratings = ratings
self.n_users, self.n_items = ratings.shape
self.n_factors = n_factors
self.item_reg = item_reg
self.user_reg = user_reg
self._v = verbose
def als_step(self,
latent_vectors,
fixed_vecs,
ratings,
_lambda,
type='user'):
"""
One of the two ALS steps. Solve for the latent vectors
specified by type.
"""
if type == 'user':
# Precompute
YTY = fixed_vecs.T.dot(fixed_vecs)
lambdaI = np.eye(YTY.shape[0]) * _lambda
for u in range(latent_vectors.shape[0]):
latent_vectors[u, :] = solve((YTY + lambdaI),
ratings[u, :].dot(fixed_vecs))
elif type == 'item':
# Precompute
XTX = fixed_vecs.T.dot(fixed_vecs)
lambdaI = np.eye(XTX.shape[0]) * _lambda
for i in range(latent_vectors.shape[0]):
latent_vectors[i, :] = solve((XTX + lambdaI),
ratings[:, i].T.dot(fixed_vecs))
return latent_vectors
def train(self, n_iter=10):
""" Train model for n_iter iterations from scratch."""
# initialize latent vectors
self.user_vecs = np.random.random((self.n_users, self.n_factors))
self.item_vecs = np.random.random((self.n_items, self.n_factors))
self.partial_train(n_iter)
def partial_train(self, n_iter):
"""
Train model for n_iter iterations. Can be
called multiple times for further training.
"""
ctr = 1
while ctr <= n_iter:
if ctr % 10 == 0 and self._v:
print ('\tcurrent iteration: {}'.format(ctr))
self.user_vecs = self.als_step(self.user_vecs,
self.item_vecs,
self.ratings,
self.user_reg,
type='user')
self.item_vecs = self.als_step(self.item_vecs,
self.user_vecs,
self.ratings,
self.item_reg,
type='item')
ctr += 1
def predict_all(self):
""" Predict ratings for every user and item. """
predictions = np.zeros((self.user_vecs.shape[0],
self.item_vecs.shape[0]))
for u in range(self.user_vecs.shape[0]):
for i in range(self.item_vecs.shape[0]):
predictions[u, i] = self.predict(u, i)
return predictions
def predict(self, u, i):
""" Single user and item prediction. """
return self.user_vecs[u, :].dot(self.item_vecs[i, :].T)
def calculate_learning_curve(self, iter_array, test):
"""
Keep track of MSE as a function of training iterations.
Params
======
iter_array : (list)
List of numbers of iterations to train for each step of
the learning curve. e.g. [1, 5, 10, 20]
test : (2D ndarray)
Testing dataset (assumed to be user x item).
The function creates two new class attributes:
train_mse : (list)
Training data MSE values for each value of iter_array
test_mse : (list)
Test data MSE values for each value of iter_array
"""
iter_array.sort()
self.train_mse =[]
self.test_mse = []
iter_diff = 0
for (i, n_iter) in enumerate(iter_array):
if self._v:
print ('Iteration: {}'.format(n_iter))
if i == 0:
self.train(n_iter - iter_diff)
else:
self.partial_train(n_iter - iter_diff)
predictions = self.predict_all()
self.train_mse += [get_mse(predictions, self.ratings)]
self.test_mse += [get_mse(predictions, test)]
if self._v:
print ('Train mse: ' + str(self.train_mse[-1]))
print ('Test mse: ' + str(self.test_mse[-1]))
iter_diff = n_iter
# +
from sklearn.metrics import mean_squared_error
def get_mse(pred, actual):
# Ignore nonzero terms.
pred = pred[actual.nonzero()].flatten()
actual = actual[actual.nonzero()].flatten()
return mean_squared_error(pred, actual)
MF_ALS = ExplicitMF(train, n_factors=40, \
user_reg=0.0, item_reg=0.0)
iter_array = [1, 2, 5, 10, 25, 50, 100]
MF_ALS.calculate_learning_curve(iter_array, test)
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
def plot_learning_curve(iter_array, model):
plt.plot(iter_array, model.train_mse, \
label='Training', linewidth=5)
plt.plot(iter_array, model.test_mse, \
label='Test', linewidth=5)
plt.xticks(fontsize=16);
plt.yticks(fontsize=16);
plt.xlabel('iterations', fontsize=30);
plt.ylabel('MSE', fontsize=30);
plt.legend(loc='best', fontsize=20);
plot_learning_curve(iter_array, MF_ALS)
# +
MF_ALS = ExplicitMF(train, n_factors=40, \
user_reg=30., item_reg=30.)
iter_array = [1, 2, 5, 10, 25, 50, 100]
MF_ALS.calculate_learning_curve(iter_array, test)
plot_learning_curve(iter_array, MF_ALS)
# +
latent_factors = [5, 10, 20, 40, 80]
regularizations = [0.1, 1., 10., 100.]
regularizations.sort()
iter_array = [1, 2, 5, 10, 25, 50, 100]
best_params = {}
best_params['n_factors'] = latent_factors[0]
best_params['reg'] = regularizations[0]
best_params['n_iter'] = 0
best_params['train_mse'] = np.inf
best_params['test_mse'] = np.inf
best_params['model'] = None
for fact in latent_factors:
print ('Factors: {}'.format(fact))
for reg in regularizations:
print ('Regularization: {}'.format(reg))
MF_ALS = ExplicitMF(train, n_factors=fact, \
user_reg=reg, item_reg=reg)
MF_ALS.calculate_learning_curve(iter_array, test)
min_idx = np.argmin(MF_ALS.test_mse)
if MF_ALS.test_mse[min_idx] < best_params['test_mse']:
best_params['n_factors'] = fact
best_params['reg'] = reg
best_params['n_iter'] = iter_array[min_idx]
best_params['train_mse'] = MF_ALS.train_mse[min_idx]
best_params['test_mse'] = MF_ALS.test_mse[min_idx]
best_params['model'] = MF_ALS
print ('New optimal hyperparameters')
print (pd.Series(best_params))
best_als_model = best_params['model']
plot_learning_curve(iter_array, best_als_model)
# -
#add SGD
class ExplicitMF():
def __init__(self,
ratings,
n_factors=40,
learning='sgd',
item_fact_reg=0.0,
user_fact_reg=0.0,
item_bias_reg=0.0,
user_bias_reg=0.0,
verbose=False):
"""
Train a matrix factorization model to predict empty
entries in a matrix. The terminology assumes a
ratings matrix which is ~ user x item
Params
======
ratings : (ndarray)
User x Item matrix with corresponding ratings
n_factors : (int)
Number of latent factors to use in matrix
factorization model
learning : (str)
Method of optimization. Options include
'sgd' or 'als'.
item_fact_reg : (float)
Regularization term for item latent factors
user_fact_reg : (float)
Regularization term for user latent factors
item_bias_reg : (float)
Regularization term for item biases
user_bias_reg : (float)
Regularization term for user biases
verbose : (bool)
Whether or not to printout training progress
"""
self.ratings = ratings
self.n_users, self.n_items = ratings.shape
self.n_factors = n_factors
self.item_fact_reg = item_fact_reg
self.user_fact_reg = user_fact_reg
self.item_bias_reg = item_bias_reg
self.user_bias_reg = user_bias_reg
self.learning = learning
if self.learning == 'sgd':
self.sample_row, self.sample_col = self.ratings.nonzero()
self.n_samples = len(self.sample_row)
self._v = verbose
def als_step(self,
latent_vectors,
fixed_vecs,
ratings,
_lambda,
type='user'):
"""
One of the two ALS steps. Solve for the latent vectors
specified by type.
"""
if type == 'user':
# Precompute
YTY = fixed_vecs.T.dot(fixed_vecs)
lambdaI = np.eye(YTY.shape[0]) * _lambda
for u in range(latent_vectors.shape[0]):
latent_vectors[u, :] = solve((YTY + lambdaI),
ratings[u, :].dot(fixed_vecs))
elif type == 'item':
# Precompute
XTX = fixed_vecs.T.dot(fixed_vecs)
lambdaI = np.eye(XTX.shape[0]) * _lambda
for i in range(latent_vectors.shape[0]):
latent_vectors[i, :] = solve((XTX + lambdaI),
ratings[:, i].T.dot(fixed_vecs))
return latent_vectors
def train(self, n_iter=10, learning_rate=0.1):
""" Train model for n_iter iterations from scratch."""
# initialize latent vectors
self.user_vecs = np.random.normal(scale=1./self.n_factors,\
size=(self.n_users, self.n_factors))
self.item_vecs = np.random.normal(scale=1./self.n_factors,
size=(self.n_items, self.n_factors))
if self.learning == 'als':
self.partial_train(n_iter)
elif self.learning == 'sgd':
self.learning_rate = learning_rate
self.user_bias = np.zeros(self.n_users)
self.item_bias = np.zeros(self.n_items)
self.global_bias = np.mean(self.ratings[np.where(self.ratings != 0)])
self.partial_train(n_iter)
def partial_train(self, n_iter):
"""
Train model for n_iter iterations. Can be
called multiple times for further training.
"""
ctr = 1
while ctr <= n_iter:
if ctr % 10 == 0 and self._v:
print ('\tcurrent iteration: {}'.format(ctr))
if self.learning == 'als':
self.user_vecs = self.als_step(self.user_vecs,
self.item_vecs,
self.ratings,
self.user_fact_reg,
type='user')
self.item_vecs = self.als_step(self.item_vecs,
self.user_vecs,
self.ratings,
self.item_fact_reg,
type='item')
elif self.learning == 'sgd':
self.training_indices = np.arange(self.n_samples)
np.random.shuffle(self.training_indices)
self.sgd()
ctr += 1
def sgd(self):
for idx in self.training_indices:
u = self.sample_row[idx]
i = self.sample_col[idx]
prediction = self.predict(u, i)
e = (self.ratings[u,i] - prediction) # error
# Update biases
self.user_bias[u] += self.learning_rate * \
(e - self.user_bias_reg * self.user_bias[u])
self.item_bias[i] += self.learning_rate * \
(e - self.item_bias_reg * self.item_bias[i])
#Update latent factors
self.user_vecs[u, :] += self.learning_rate * \
(e * self.item_vecs[i, :] - \
self.user_fact_reg * self.user_vecs[u,:])
self.item_vecs[i, :] += self.learning_rate * \
(e * self.user_vecs[u, :] - \
self.item_fact_reg * self.item_vecs[i,:])
def predict(self, u, i):
""" Single user and item prediction."""
if self.learning == 'als':
return self.user_vecs[u, :].dot(self.item_vecs[i, :].T)
elif self.learning == 'sgd':
prediction = self.global_bias + self.user_bias[u] + self.item_bias[i]
prediction += self.user_vecs[u, :].dot(self.item_vecs[i, :].T)
return prediction
def predict_all(self):
""" Predict ratings for every user and item."""
predictions = np.zeros((self.user_vecs.shape[0],
self.item_vecs.shape[0]))
for u in range(self.user_vecs.shape[0]):
for i in range(self.item_vecs.shape[0]):
predictions[u, i] = self.predict(u, i)
return predictions
def calculate_learning_curve(self, iter_array, test, learning_rate=0.1):
"""
Keep track of MSE as a function of training iterations.
Params
======
iter_array : (list)
List of numbers of iterations to train for each step of
the learning curve. e.g. [1, 5, 10, 20]
test : (2D ndarray)
Testing dataset (assumed to be user x item).
The function creates two new class attributes:
train_mse : (list)
Training data MSE values for each value of iter_array
test_mse : (list)
Test data MSE values for each value of iter_array
"""
iter_array.sort()
self.train_mse =[]
self.test_mse = []
iter_diff = 0
for (i, n_iter) in enumerate(iter_array):
if self._v:
print ('Iteration: {}'.format(n_iter))
if i == 0:
self.train(n_iter - iter_diff, learning_rate)
else:
self.partial_train(n_iter - iter_diff)
predictions = self.predict_all()
self.train_mse += [get_mse(predictions, self.ratings)]
self.test_mse += [get_mse(predictions, test)]
if self._v:
print ('Train mse: ' + str(self.train_mse[-1]))
print ('Test mse: ' + str(self.test_mse[-1]))
iter_diff = n_iter
MF_SGD = ExplicitMF(train, 40, learning='sgd', verbose=True)
iter_array = [1, 2, 5, 10, 25, 50, 100, 200]
MF_SGD.calculate_learning_curve(iter_array, test, learning_rate=0.001)
# +
plot_learning_curve(iter_array, MF_SGD)
iter_array = [1, 2, 5, 10, 25, 50, 100, 200]
learning_rates = [1e-5, 1e-4, 1e-3, 1e-2]
best_params = {}
best_params['learning_rate'] = None
best_params['n_iter'] = 0
best_params['train_mse'] = np.inf
best_params['test_mse'] = np.inf
best_params['model'] = None
for rate in learning_rates:
print ('Rate: {}'.format(rate))
MF_SGD = ExplicitMF(train, n_factors=40, learning='sgd')
MF_SGD.calculate_learning_curve(iter_array, test, learning_rate=rate)
min_idx = np.argmin(MF_SGD.test_mse)
if MF_SGD.test_mse[min_idx] < best_params['test_mse']:
best_params['n_iter'] = iter_array[min_idx]
best_params['learning_rate'] = rate
best_params['train_mse'] = MF_SGD.train_mse[min_idx]
best_params['test_mse'] = MF_SGD.test_mse[min_idx]
best_params['model'] = MF_SGD
print ('New optimal hyperparameters')
print (pd.Series(best_params))
# +
iter_array = [1, 2, 5, 10, 25, 50, 100, 200]
latent_factors = [5, 10, 20, 40, 80]
regularizations = [0.001, 0.01, 0.1, 1.]
regularizations.sort()
best_params = {}
best_params['n_factors'] = latent_factors[0]
best_params['reg'] = regularizations[0]
best_params['n_iter'] = 0
best_params['train_mse'] = np.inf
best_params['test_mse'] = np.inf
best_params['model'] = None
for fact in latent_factors:
print ('Factors: {}'.format(fact))
for reg in regularizations:
print ('Regularization: {}'.format(reg))
MF_SGD = ExplicitMF(train, n_factors=fact, learning='sgd',\
user_fact_reg=reg, item_fact_reg=reg, \
user_bias_reg=reg, item_bias_reg=reg)
MF_SGD.calculate_learning_curve(iter_array, test, learning_rate=0.001)
min_idx = np.argmin(MF_SGD.test_mse)
if MF_SGD.test_mse[min_idx] < best_params['test_mse']:
best_params['n_factors'] = fact
best_params['reg'] = reg
best_params['n_iter'] = iter_array[min_idx]
best_params['train_mse'] = MF_SGD.train_mse[min_idx]
best_params['test_mse'] = MF_SGD.test_mse[min_idx]
best_params['model'] = MF_SGD
print ('New optimal hyperparameters')
print (pd.Series(best_params))
# -
3http://blog.ethanrosenthal.com/2017/06/20/matrix-factorization-in-pytorch/
#PYTORCH
# +
import numpy as np
from scipy.sparse import rand as sprand
import torch
from torch.autograd import Variable
# Make up some random explicit feedback ratings
# and convert to a numpy array
n_users = 1000
n_items = 1000
ratings = sprand(n_users, n_items,
density=0.01, format='csr')
ratings.data = (np.random.randint(1, 5,
size=ratings.nnz)
.astype(np.float64))
ratings = ratings.toarray()
# -
class MatrixFactorization(torch.nn.Module):
def __init__(self, n_users, n_items, n_factors=20):
super().__init__()
self.user_factors = torch.nn.Embedding(n_users,
n_factors,
sparse=True)
self.item_factors = torch.nn.Embedding(n_items,
n_factors,
sparse=True)
def forward(self, user, item):
return (self.user_factors(user) * self.item_factors(item)).sum(1)
# +
model = MatrixFactorization(n_users, n_items, n_factors=20)
loss_func = torch.nn.MSELoss()
optimizer = torch.optim.SGD(model.parameters(),
lr=1e-6) # learning rate
# Sort our data
rows, cols = ratings.nonzero()
p = np.random.permutation(len(rows))
rows, cols = rows[p], cols[p]
for row, col in zip(*(rows, cols)):
# Turn data into variables
rating = Variable(torch.FloatTensor([ratings[row, col]]))
row = Variable(torch.LongTensor([np.long(row)]))
col = Variable(torch.LongTensor([np.long(col)]))
# Predict and calculate loss
prediction = model.forward(row, col)
loss = loss_func(prediction, rating)
# Backpropagate
loss.backward()
# Update the parameters
optimizer.step()
# -
class BiasedMatrixFactorization(torch.nn.Module):
def __init__(self, n_users, n_items, n_factors=20):
super().__init__()
self.user_factors = torch.nn.Embedding(n_users,
n_factors,
sparse=True)
self.item_factors = torch.nn.Embedding(n_items,
n_factors,
sparse=True)
self.user_biases = torch.nn.Embedding(n_users,
1,
sparse=True)
self.item_biases = torch.nn.Embedding(n_items,
1,
sparse=True)
def forward(self, user, item):
pred = self.user_biases(user) + self.item_biases(item)
pred += (self.user_factors(user) * self.item_factors(item)).sum(1)
return pred
|
hinton/hidden_factors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AnkurMali/IST597_Spring_2022/blob/main/IST597_MLP_withkeras_collab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="v0hRDnJGY3bH"
import os
import numpy as np
import time
import tensorflow as tf
np.random.seed(1234)
tf.random.set_seed(1234)
# + id="6dv_IcvjZJz6"
size_input = 32
size_hidden = 128
size_output = 1
number_of_train_examples = 1000
number_of_test_examples = 300
# + id="ANBNiazKZMxe"
X_train = np.random.randn(number_of_train_examples , size_input)
y_train = np.random.randn(number_of_train_examples)
X_test = np.random.randn(number_of_test_examples, size_input)
y_test = np.random.randn(number_of_test_examples)
# + id="p29oB9imZQC0"
# Split dataset into batches
train_ds = tf.data.Dataset.from_tensor_slices((X_train, y_train)).batch(16)
test_ds = tf.data.Dataset.from_tensor_slices((X_test, y_test)).batch(4)
# + id="DpGETHIzZj1T"
class MLP(tf.keras.Model):
def __init__(self, size_input, size_hidden, size_output, device=None):
super(MLP, self).__init__()
"""
size_input: int, size of input layer
size_hidden: int, size of hidden layer
size_output: int, size of output layer
device: str or None, either 'cpu' or 'gpu' or None. If None, the device to be used will be decided automatically during Eager Execution
"""
# self.size_input = 32
# self.size_hidden = 128
# self.size_output = 1
# self.device = 'gpu'
self.size_input, self.size_hidden, self.size_output, self.device =\
size_input, size_hidden, size_output, device
# Initialize weights between input layer and hidden layer
self.W1 = tf.Variable(tf.random.normal([self.size_input, self.size_hidden]))
# Initialize biases for hidden layer
self.b1 = tf.Variable(tf.random.normal([1, self.size_hidden]))
# Initialize weights between hidden layer and output layer
self.W2 = tf.Variable(tf.random.normal([self.size_hidden, self.size_output]))
# Initialize biases for output layer
self.b2 = tf.Variable(tf.random.normal([1, self.size_output]))
# Define variables to be updated during backpropagation
self.MLP_variables = [self.W1, self.W2, self.b1, self.b2]
def forward(self, X):
"""
forward pass
X: Tensor, inputs
"""
if self.device is not None:
with tf.device('gpu:0' if self.device=='gpu' else 'cpu'):
self.y = self.compute_output(X)
else:
self.y = self.compute_output(X)
return self.y
def loss(self, y_pred, y_true):
'''
y_pred - Tensor of shape (batch_size, size_output)
y_true - Tensor of shape (batch_size, size_output)
'''
y_true_tf = tf.cast(tf.reshape(y_true, (-1, self.size_output)), dtype=tf.float32)
y_pred_tf = tf.cast(y_pred, dtype=tf.float32)
return tf.losses.mean_squared_error(y_true_tf, y_pred_tf)
def backward(self, X_train, y_train):
"""
backward pass
"""
optimizer = tf.keras.optimizers.SGD(learning_rate=1e-4)
with tf.GradientTape() as tape:
predicted = self.forward(X_train)
current_loss = self.loss(predicted, y_train)
grads = tape.gradient(current_loss, self.MLP_variables)
optimizer.apply_gradients(zip(grads, self.MLP_variables))
def compute_output(self, X):
"""
Custom method to obtain output tensor during forward pass
"""
# Cast X to float32
X_tf = tf.cast(X, dtype=tf.float32)
#Remember to normalize your dataset before moving forward
# Compute values in hidden layer
what = tf.matmul(X_tf, self.W1) + self.b1
hhat = tf.nn.relu(what)
# Compute output
output = tf.matmul(hhat, self.W2) + self.b2
#Now consider two things , First look at inbuild loss functions if they work with softmax or not and then change this
#Second add tf.Softmax(output) and then return this variable
return output
# + colab={"base_uri": "https://localhost:8080/"} id="cwg_5g_pZ56Z" outputId="8efb0a28-6177-4b95-9a2a-6fdfc0ce9700"
# Set number of epochs
NUM_EPOCHS = 10
# Initialize model using GPU
#mlp_on_gpu = MLP()
mlp_on_gpu = MLP(size_input, size_hidden, size_output, device='cpu')
time_start = time.time()
for epoch in range(NUM_EPOCHS):
loss_total_gpu = tf.zeros([1,1], dtype=tf.float32)
lt = 0
train_ds = tf.data.Dataset.from_tensor_slices((X_train, y_train)).shuffle(25, seed=epoch*(1234)).batch(20)
for inputs, outputs in train_ds:
preds = mlp_on_gpu.forward(inputs)
loss_total_gpu = loss_total_gpu + mlp_on_gpu.loss(preds, outputs)
lt = lt + mlp_on_gpu.loss(preds, outputs)
mlp_on_gpu.backward(inputs, outputs)
print('Number of Epoch = {} - Average MSE:= {}'.format(epoch + 1, np.sum(loss_total_gpu) / X_train.shape[0]))
time_taken = time.time() - time_start
print('\nTotal time taken (in seconds): {:.2f}'.format(time_taken))
|
week3/IST597_MLP_withkeras_collab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import scipy.optimize as opt
import sklearn.linear_model
import sklearn.model_selection
# Load the dataset
# + tags=["outputPrepend"]
from sklearn.datasets import fetch_20newsgroups
data = fetch_20newsgroups()
text, label = data['data'][0], data['target_names'][data['target'][0]]
print(list(data)) #see list of attributes in this class.
print('---')
print(list(data.target_names)) # list 20 class under this dataset.
print('---')
print(len(data['target'])) #no of rows = 11314
# -
# Use `HashingVectorizer` to encode the text into sparse features:
# +
from sklearn.feature_extraction.text import HashingVectorizer
vectorizer = HashingVectorizer(n_features=5000, binary=True)
features = vectorizer.fit_transform(data['data'])
targets = data['target']
newfeatures = features.todense()
print(targets) #todense return a matrix.
# -
# features have a dimension of 11314 samples * 5000 features
print(newfeatures.shape)
print(targets.shape)
print(type(newfeatures))
print(type(targets))
# Use the K-Fold cross-validation to split the dataset into training and test parts:
Kfolddata = sklearn.model_selection.KFold(n_splits=10)
# Experiment with different models (L1, L2, ...)
# +
scores = []
classification_model = sklearn.linear_model.SGDClassifier(loss='log', penalty='l1')
for train_index, test_index in Kfolddata.split(newfeatures):
print(newfeatures.shape, ' ' , targets.shape)
X_train, X_test, y_train, y_test = newfeatures[train_index], newfeatures[test_index], targets[train_index], targets[test_index]
classification_model.fit(X_train, y_train)
scores.append(classification_model.score(X_test, y_test))
print(np.mean(scores))
#100features, 100 folds: 0.19037571805620246
#1000 features, 10 folds : 0.6530836715374306
# -
#Simpler methods
classification_modelL1 = sklearn.linear_model.SGDClassifier(loss='squared_loss', penalty='l1', alpha=0.0001)
sklearn.model_selection.cross_val_score(classification_modelL1, newfeatures, targn_jobs=-1)
mean(avg_score)
#5000 features, 5 folds, 0.0001 alpha : 0.7206110055547025
#1000 features, 5 folds, 0.001 alpha : 0.2530836715374306
#1000 features, 5 folds, 0.00001 alpha :0.4664129796483858
from statistics import mean
classification_modelL2 = sklearn.linear_model.SGDClassifier(loss='log', penalty='l2', alpha=0.00001)
avg_score = sklearn.model_selection.cross_val_score(classification_modelL2, newfeatures, targets, cv=5)
mean(avg_score)
#5000 features, 5 folds, 0.0001 alpha : 0.8168636814194283
#1000 features, 5 folds, 0.00001 alpha : 0.7180491690998038
#5000 features, 5 folds, 0.00001 alpha : 0.8568143271238033
from statistics import mean
classification_modelL2 = sklearn.linear_model.SGDClassifier(loss='log', penalty='elasticnet', alpha=0.00001)
avg_score = sklearn.model_selection.cross_val_score(classification_modelL2, newfeatures, targets, cv=5)
mean(avg_score)
#5000 features, 5 folds, 0.0001 alpha : 0.801838400627009
#1000 features, 5 folds, 0.00001 alpha : 0.7206110055547025
#5000 features, 5 folds, 0.00001 alpha : 0.8538985478537797
# + [markdown] slideshow={"slide_type": "fragment"}
# What model worked best?
# -
# L2 and elasticnet works similar, L1 is significantly weaker
|
machine-learning/HSE-AML-HW-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
#
# ===========================================================================
# Motor imagery decoding from EEG data using the Common Spatial Pattern (CSP)
# ===========================================================================
#
# Decoding of motor imagery applied to EEG data decomposed using CSP.
# Here the classifier is applied to features extracted on CSP filtered signals.
#
# See https://en.wikipedia.org/wiki/Common_spatial_pattern and [1]_. The EEGBCI
# dataset is documented in [2]_. The data set is available at PhysioNet [3]_.
#
# References
# ----------
#
# .. [1] <NAME>. The quantitative extraction and topographic mapping
# of the abnormal components in the clinical EEG. Electroencephalography
# and Clinical Neurophysiology, 79(6):440--447, December 1991.
# .. [2] <NAME>., <NAME>., <NAME>., <NAME>.,
# <NAME>. (2004) BCI2000: A General-Purpose Brain-Computer Interface
# (BCI) System. IEEE TBME 51(6):1034-1043.
# .. [3] <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>,
# <NAME>, <NAME>, <NAME>, <NAME>. (2000) PhysioBank,
# PhysioToolkit, and PhysioNet: Components of a New Research Resource for
# Complex Physiologic Signals. Circulation 101(23):e215-e220.
#
# +
# Authors: <NAME> <<EMAIL>>
#
# License: BSD (3-clause)
import numpy as np
import matplotlib.pyplot as plt
from sklearn.pipeline import Pipeline
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.model_selection import ShuffleSplit, cross_val_score
from mne import Epochs, pick_types, events_from_annotations
from mne.channels import read_layout
from mne.io import concatenate_raws, read_raw_edf
from mne.datasets import eegbci
from mne.decoding import CSP
print(__doc__)
# #############################################################################
# # Set parameters and read data
# avoid classification of evoked responses by using epochs that start 1s after
# cue onset.
tmin, tmax = -1., 4.
event_id = dict(hands=2, feet=3)
subject = 1
runs = [6, 10, 14] # motor imagery: hands vs feet
raw_fnames = eegbci.load_data(subject, runs)
raw = concatenate_raws([read_raw_edf(f, preload=True) for f in raw_fnames])
# strip channel names of "." characters
raw.rename_channels(lambda x: x.strip('.'))
# Apply band-pass filter
raw.filter(7., 30., fir_design='firwin', skip_by_annotation='edge')
events, _ = events_from_annotations(raw, event_id=dict(T1=2, T2=3))
picks = pick_types(raw.info, meg=False, eeg=True, stim=False, eog=False,
exclude='bads')
# Read epochs (train will be done only between 1 and 2s)
# Testing will be done with a running classifier
epochs = Epochs(raw, events, event_id, tmin, tmax, proj=True, picks=picks,
baseline=None, preload=True)
epochs_train = epochs.copy().crop(tmin=1., tmax=2.)
labels = epochs.events[:, -1] - 2
# -
# Classification with linear discrimant analysis
#
#
# +
# Define a monte-carlo cross-validation generator (reduce variance):
scores = []
epochs_data = epochs.get_data()
epochs_data_train = epochs_train.get_data()
cv = ShuffleSplit(10, test_size=0.2, random_state=42)
cv_split = cv.split(epochs_data_train)
# Assemble a classifier
lda = LinearDiscriminantAnalysis()
csp = CSP(n_components=4, reg=None, log=True, norm_trace=False)
# Use scikit-learn Pipeline with cross_val_score function
clf = Pipeline([('CSP', csp), ('LDA', lda)])
scores = cross_val_score(clf, epochs_data_train, labels, cv=cv, n_jobs=1)
# Printing the results
class_balance = np.mean(labels == labels[0])
class_balance = max(class_balance, 1. - class_balance)
print("Classification accuracy: %f / Chance level: %f" % (np.mean(scores),
class_balance))
# plot CSP patterns estimated on full data for visualization
csp.fit_transform(epochs_data, labels)
layout = read_layout('EEG1005')
csp.plot_patterns(epochs.info, layout=layout, ch_type='eeg',
units='Patterns (AU)', size=1.5)
# -
# Look at performance over time
#
#
# +
sfreq = raw.info['sfreq']
w_length = int(sfreq * 0.5) # running classifier: window length
w_step = int(sfreq * 0.1) # running classifier: window step size
w_start = np.arange(0, epochs_data.shape[2] - w_length, w_step)
scores_windows = []
for train_idx, test_idx in cv_split:
y_train, y_test = labels[train_idx], labels[test_idx]
X_train = csp.fit_transform(epochs_data_train[train_idx], y_train)
X_test = csp.transform(epochs_data_train[test_idx])
# fit classifier
lda.fit(X_train, y_train)
# running classifier: test classifier on sliding window
score_this_window = []
for n in w_start:
X_test = csp.transform(epochs_data[test_idx][:, :, n:(n + w_length)])
score_this_window.append(lda.score(X_test, y_test))
scores_windows.append(score_this_window)
# Plot scores over time
w_times = (w_start + w_length / 2.) / sfreq + epochs.tmin
plt.figure()
plt.plot(w_times, np.mean(scores_windows, 0), label='Score')
plt.axvline(0, linestyle='--', color='k', label='Onset')
plt.axhline(0.5, linestyle='-', color='k', label='Chance')
plt.xlabel('time (s)')
plt.ylabel('classification accuracy')
plt.title('Classification score over time')
plt.legend(loc='lower right')
plt.show()
|
stable/_downloads/65f9e13513d4b308c2019298009b3efd/plot_decoding_csp_eeg.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from add_parent_path import add_parent_path
with add_parent_path(1):
from clinical_sectionizer import TextSectionizer
# -
text = """
FINDINGS: Compared to the prior days study, there is stable appearance of the
right parietal intraparenchymal hemorrhage with surrounding edema. At the
superior margin of the parenchymal hemorrhage there is a rounded heterogeneous
focus which could represent a metastatic lesion. An additional 2mm hyperdense
focus, possibly hemorrhage, is noted in the posteromedial margin of the left
thalamus, with surroundng edema. Low-attenuation foci seen in both basal
ganglia and insular regions are consistent with chronic lacunar infarcts.
There is no shift of midline structures. The ventricles are stable in
appearance. The osseous and soft tissue structures are unremarkable.
IMPRESSION: Stable appearance of right parietal lobe and left thalamic
hemorrhages, which are concerning for hemorrhagic metastasis in this patient
with known metastatic lung carcinoma to the brain."""
# #### By default section detection is done ignoring case (`re.I`)
# +
sectionizer = TextSectionizer(patterns=None)
rad_patterns = [{'section_title': 'impression',
'pattern':'impression:'},]
sectionizer.add(rad_patterns)
sections = sectionizer(text)
# -
# #### Text is split into two parts: before and after section
len(sections)
sections[1]
# #### Create regular expression without `re.I` flag
# +
sectionizer = TextSectionizer(patterns=None)
rad_patterns = [{'section_title': 'impression',
'pattern':'impression:'},]
sectionizer.add(rad_patterns, cflags=[])
sections = sectionizer(text)
# -
# #### Section not detected
#
# Only one section in result
len(sections)
|
notebooks/with_compile_flags.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from datetime import datetime
from typing import List
from pydantic import BaseModel
from pdcst.repository import PodcastInDb, EpisodeInDb, FeedInDb
# +
updated1 = datetime(2022, 1, 1)
feed1 = FeedInDb(url="https://example.com/rss.xml", updated=updated1)
podcast1 = PodcastInDb(feed=feed1, title="Python Podcast")
updated2 = datetime(2022, 2, 1)
feed2 = FeedInDb(url="https://foobar.com/atom.xml", updated=updated2)
podcast2 = PodcastInDb(feed=feed2, title="Foo Bar")
# -
podcast1.json()
podcast.dict()
class PodcastList(BaseModel):
__root__: List[PodcastInDb]
pl = PodcastList(__root__=[podcast1, podcast2])
pl.json()
pl.json(exclude={"episodes"})
list(pl)
for index, i in enumerate(range(10), 1):
print(index, i)
|
notebooks/serialization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center>
# <img src="../../img/ods_stickers.jpg">
# ## Открытый курс по машинному обучению
# <center>Автор материала: <NAME>, Levka.
# ## <center>Обзор библиотеки для генерации временных признаков tsfresh</center>
# ### <center>Time Series FeatuRe Extraction based on Scalable Hypothesis tests</center>
# Библиотека используется для извлечения признаков из временных рядов. Практически все признаки, которые могут прийти вам в голову, уже внесены в расчёт этой библиотеки и нет никакого смысла создавать их самому, когда это можно сделать парой строчек кода из библиотеки.
#
# Извлечённые признаки могут быть использованы для описания или кластеризации временных рядов. Также их можно использовать для задач классификации/регрессии на временных рядах.
# <img src="http://tsfresh.readthedocs.io/en/latest/_images/introduction_ts_exa_features.png" width="700" height="600">
# ### Процесс расчёта признаков состоит из двух этапов:
# - Расчёт всех возможных признаков
#
# ```python
# from tsfresh import extract_features
# extracted_features = extract_features(timeseries, column_id="id", column_sort="time")
# ```
# - Отбор релевантных признаков и удаление константных/нулевых признаков
#
# ```python
# from tsfresh import select_features
# from tsfresh.utilities.dataframe_functions import impute
#
# impute(extracted_features) # удаление константных признаков
# features_filtered = select_features(extracted_features, y) # отбор признаков
# ```
# ### Процедура отбора признаков
# #### Стадия 1
# Расчёт признаков
# #### Стадия 2
# Проверка на значимость каждого признака, расчёт p-value
# #### Стадия 3
# Поправка на множественную проверку гипотез Бенджамини-Иекутиели
# <img src="http://tsfresh.readthedocs.io/en/latest/_images/feature_extraction_process_20160815_mc_1.png" width="700" height="600">
# ### Приведём пример генерации признаков на основе датасета Human Activity Recognition
# +
import matplotlib.pylab as plt
# %matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from tsfresh import (extract_features, extract_relevant_features,
select_features)
from tsfresh.examples.har_dataset import (download_har_dataset,
load_har_classes, load_har_dataset)
from tsfresh.feature_extraction import ComprehensiveFCParameters
from tsfresh.utilities.dataframe_functions import impute
# -
# **Загрузка и отрисовка данных**
download_har_dataset()
df = load_har_dataset()
plt.title('accelerometer reading')
plt.plot(df.iloc[0,:])
plt.show()
# **Извлечение признаков**
# расчёт только определённого набора параметров, заданного в ComprehensiveFCParameters
extraction_settings = ComprehensiveFCParameters()
# переформируем данные 500 первых показаний сенсоров column-wise, как этого требует формат библиотеки
N = 500
master_df = pd.DataFrame({0: df[:N].values.flatten(),
1: np.arange(N).repeat(df.shape[1])})
master_df.head()
X = extract_features(master_df, column_id=1, impute_function=impute, default_fc_parameters=extraction_settings)
"Число рассчитанных признаков: {}.".format(X.shape[1])
# **Обучение классификатора**
y = load_har_classes()[:N]
y.shape
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.2)
cl = DecisionTreeClassifier()
cl.fit(X_train, y_train)
print(classification_report(y_test, cl.predict(X_test)))
# **Отберём признаки для каждого класса отдельно и решим задачу бинарной классификации**
# +
relevant_features = set()
for label in y.unique():
y_train_binary = y_train == label
X_train_filtered = select_features(X_train, y_train_binary)
print("Number of relevant features for class {}: {}/{}".format(label, X_train_filtered.shape[1], X_train.shape[1]))
relevant_features = relevant_features.union(set(X_train_filtered.columns))
# -
len(relevant_features)
# Мы уменьшили количество признаков с 794 до 264.
X_train_filtered = X_train[list(relevant_features)]
X_test_filtered = X_test[list(relevant_features)]
X_train_filtered.shape, X_test_filtered.shape
cl = DecisionTreeClassifier()
cl.fit(X_train_filtered, y_train)
print(classification_report(y_test, cl.predict(X_test_filtered)))
# Качество модели практически не изменилось, однако модель стала намного проще.
# **Сравнение с классификатором на стандартных признаках**
X_1 = df.iloc[:N,:]
X_1.shape
X_train, X_test, y_train, y_test = train_test_split(X_1, y, test_size=.2)
cl = DecisionTreeClassifier()
cl.fit(X_train, y_train)
print(classification_report(y_test, cl.predict(X_test)))
# Как видимо, качество модели значительно улучшилось по сравнению с наивным классификатором.
|
jupyter_russian/tutorials/tsfresh_Levka.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example of processing UI changes to pandas changes
# This notebook explores how a JSON file returned from the User Interface could be used to make changes to the panda dataframes that are used as the primary input into the optimizer model.
#
# At the time of writing it was clear that the UI would be unable to generate test data to build this facility, so we have assumed that the output files from the UI will look similar to those that were passed to it i.e. they will be in a hierarchical JSON format.
import workforce_pandas as wfpd
wfpd.sheets
# As the return data is quite large, it is more convenient to play with it as a file. It is recommended that a JSON response is captured from the main data requests APIs (provider_profile, geo_profile) and then modified in an online JSON editor to add, change and remove information. This will provide a good test file.
#
# In this example we are going to look at provider/type to service mapping and understand what changes (if any) have been made in the user interface.
#
# The instruction below loads the JSON file as a Pyton data dictionary for manipulation.
# +
import json
with open('my_output2.json') as json_data:
my_dict = json.load(json_data)
# -
# The format of the resulting dictionary is shown in the cell below:
my_dict
# To navigate the JSON structure, it will be useful to have a list of the provider types abbreviations; these can be extracted quickly from the provider list dataframe.
provider_list = wfpd.dataframes["provider_list"]["provider_abbr"]
list(provider_list)
# The data we are going to compare against is stored in the service characteristics dataframe:
total_wage = wfpd.dataframes["service_characteristics"]
total_wage
# The routine below cycles through the dataframe one row at a time. For each column that corresponds to a profession type, it reads the existing value for suitability.
#
# As the JSON only contains the information with valid entries and a number of arrays, the parsing is not necessarily straightforward. The nested loops below basically parses the JSON tree using the values from the base dataframe, detecting dead ends in the JSON data structure and then moving on.
#
# This routine successfully identifies:
# When suitabilityX
not_defined_in_json = "not defined in json"
for row_index, row in total_wage.iterrows():
#print (row['min_f2f_time'], row['max_f2f_time'])
print ("==========================================================")
print (str(row['svc_category'] +"/"+ str(row['svc_desc'])))
for column in total_wage[provider_list]:
#score = my_dict.get('response').get(column).get('services:')[0].get(row['svc_category'])[0].get(row['svc_desc'].get('score'))
new_value = not_defined_in_json
old_value = row[column]
#print (type(old_value),old_value)
#print (column + " old value : " + str(old_value))
category_size = int(len(my_dict.get('response').get(column).get('services:')))
#print ("Category: " + str(category_size))
for n in range(category_size):
in_json1 = my_dict.get('response').get(column).get('services:')[n].get(row['svc_category'])
if in_json1 != None :
description_size = int(len(my_dict.get('response').get(column).get('services:')[n].get(row['svc_category'])))
#print ("Svc desc: " + str(description_size
for m in range(description_size):
in_json2 = my_dict.get('response').get(column).get('services:')[n].get(row['svc_category'])[m]
if in_json2 != None :
#print(in_json2)
in_json3 = my_dict.get('response').get(column).get('services:')[n].get(row['svc_category'])[m].get(row['svc_desc'])
if in_json3 != None :
new_value = my_dict.get('response').get(column).get('services:')[n].get(row['svc_category'])[m].get(row['svc_desc']).get('score')
string = "Comparison: " + column + " : " + str(old_value) + " -> " + str(new_value)
#print (string)
#print ("Changing " + column + ": " + str(old_value) + " to " + str(new_value))
print ("Changing " + column + ": " + str(old_value) + " to " + str(new_value))
response = my_dict.get('response')
for item in provider_list:
print ("=========================="+ str(item) +"=========================================================")
services = response.get(item).get('services:')
for item in services:
print ("=========================="+ str(item) +"=========================================================")
print (str(item))
|
models/4.1.1 Model Manipulation/.ipynb_checkpoints/Changing Model State in Response to UI Changes-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Numpy Array Basics - Boolean Selection
import sys
print(sys.version)
import numpy as np
print(np.__version__)
npa = np.arange(20)
npa
# Now i’m going to introduce some new notation but you’re going to be seeing this a lot in pandas. It’s similar to a dictionary but we’re performing boolean selection.
#
# Boolean selection is not so different from filtering or using list comprehensions like we did in the last selection.
#
# let's start with a worked example.
#
# Let's get all values that are divisible by 2 in our list
[x for x in npa if x % 2 == 0]
list(filter(lambda x: x % 2 ==0, npa))
# You can see how we did that with the list comprehension and a filter, now let’s do it with numpy.
npa % 2 == 0
#
# It’s an interesting notation but what the result of what we're getting isn't really so different. We're basically just getting the boolean value of the result of each value in the array.
# so how might we complete the filter? Easy, we just treat it like a dictionary and query our original array for those values that are true.
npa[npa % 2 == 0]
# Now you might ask yourself why things are done this way and now we’re starting going to get into the efficiency of the operation. And for datasets of reasonable size this is typically orders of magnitude. Let me show you very quickly before we move on to different boolean selections.
#
np2 = np.arange(20000)
# %timeit [x for x in np2 if x % 2 == 0]
# %timeit np2[np2 % 2 == 0]
# We can see that it is orders of magnitude faster than our original list comprehension.
npa
# Now here’s an exercise, try to do the same thing but get all numbers from that array that are greater than 10. Go ahead and pause and try it out.
npa > 10
npa[npa > 10]
# How about greater than 15 or less than 5?
npa > 15 or npa < 5
npa > 15 | npa < 5
# Instinct is to use “or” but we can’t use that. We have to use the bar in place of or and wrap them up in parenthesis.
#
# now we can filter down with this expression.
(npa > 15) | (npa < 5)
#
# Now that is basically boolean selection, we are querying data that we want to from an array. This is an extremely powerful concept that will come up over and over again and don’t worry if you don’t understand it completely just understand how it works.
npa[(npa > 15) | (npa < 5)]
# on a final note, for boolean selection to occur, you just have to pass in a list with the same length as the original list and has boolean values. Let me show you quickly.
np3 = np.array([True for x in range(20)])
npa[np3]
|
3 - NumPy Basics/3-1 Numpy Array Basics - Boolean Selection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# ### Load csv data as DataFrame
names_df = pd.read_csv("../DF/", sep="\t")
names_df.head(n=3)
names_df.sample(n=3)
# #### header param
names_df = pd.read_csv("name.basics_sample_500.tsv", sep="\t", header=)
prices_ds = pd.Series([1.5, 2, 2.5, 3])
prices_ds
|
LIVE/labs/pandas/DF/imdb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Estimating COVID-19's $R_t$ in Real-Time
# <NAME> - April 12
#
# Adapted for South Africa - Vukosi Marivate April 14
#
# In any epidemic, $R_t$ is the measure known as the effective reproduction number. It's the number of people who become infected per infectious person at time $t$. The most well-known version of this number is the basic reproduction number: $R_0$ when $t=0$. However, $R_0$ is a single measure that does not adapt with changes in behavior and restrictions.
#
# As a pandemic evolves, increasing restrictions (or potential releasing of restrictions) change $R_t$. Knowing the current $R_t$ is essential. When $R>1$, the pandemic will spread through the entire population. If $R_t<1$, the pandemic will grow to some fixed number less than the population. The lower $R_t$, the more manageable the situation. The value of $R_t$ helps us (1) understand how effective our measures have been controlling an outbreak and (2) gives us vital information about whether we should increase or reduce restrictions based on our competing goals of economic prosperity and human safety. [Well-respected epidemiologists argue](https://www.nytimes.com/2020/04/06/opinion/coronavirus-end-social-distancing.html) that tracking $R_t$ is the only way to manage through this crisis.
#
# Yet, today, to my knowledge there is no real-time tracking of $R_t$ in United States. In fact, the only real-time measure I've seen has been for [Hong Kong](https://covid19.sph.hku.hk/dashboard). More importantly, it is not useful to understand $R_t$ at a national level. Instead, to manage this crisis effectively, we need a local (state, county and/or city) level granularity of $R_t$.
#
# What follows is a solution to this problem at the US State level. It's a modified version of a solution created by [Bettencourt & Ribeiro 2008](https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0002185) to estimate real-time $R_t$ using a Bayesian approach. While I have stayed true to most of their process, my solution differs in an important way that I will call out clearly.
#
# If you have questions, comments, or improvments feel free to get in touch: [<EMAIL>](mailto:<EMAIL>). And if it's not entirely clear, I'm not an epidemiologist. At the same time, data is data, and statistics are statistics and this is based on work by well-known epidemiologists so calibrate accordingly. In the meantime, I hope you can learn something new as I did by reading through this example. Feel free to take this work and apply it elsewhere – internationally or to counties in the United States.
# +
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
from matplotlib.dates import date2num, num2date
from matplotlib import dates as mdates
from matplotlib import ticker
from matplotlib.colors import ListedColormap
from matplotlib.patches import Patch
from scipy import stats as sps
from scipy.interpolate import interp1d
from IPython.display import clear_output
FILTERED_REGIONS = [
'Virgin Islands',
'American Samoa',
'Northern Mariana Islands',
'Guam',
'Puerto Rico']
# %config InlineBackend.figure_format = 'retina'
# -
# ## Bettencourt & Ribeiro's Approach
#
# Every day, we learn how many more people have COVID-19. This new case count gives us a clue about the current value of $R_t$. We also, figure that the value of $R_t$ today is related to the value of $R_{t-1}$ (yesterday's value) and every previous value of $R_{t-m}$ for that matter.
#
# With these insights, the authors use [Bayes' rule](https://en.wikipedia.org/wiki/Bayes%27_theorem) to update their beliefs about the true value of $R_t$ based on how many new cases have been reported each day.
#
# This is Bayes' Theorem as we'll use it:
#
# $$ P(R_t|k)=\frac{P(R_t)\cdot\mathcal{L}(R_t|k)}{P(k)} $$
#
# This says that, having seen $k$ new cases, we believe the distribution of $R_t$ is equal to:
#
# - The __prior__ beliefs of the value of $P(R_t)$ without the data ...
# - times the __likelihood__ of $R_t$ given that we've seen $k$ new cases ...
# - divided by the probability of seeing this many cases in general.
#
# Importantly, $P(k)$ is a constant, so the numerator is proportional to the posterior. Since all probability distributions sum to 1.0, we can ignore $P(k)$ and normalize our posterior to sum to 1.0:
#
# $$ P(R_t|k) \propto P(R_t) \cdot \mathcal{L}(R_t|k) $$
#
# This is for a single day. To make it iterative: every day that passes, we use yesterday's conclusion (ie. posterior) $P(R_{t-1}|k_{t-1})$ to be today's prior $P(R_t)$ so on day two:
#
# $$ P(R_2|k) \propto P(R_0)\cdot\mathcal{L}(R_2|k_2)\cdot\mathcal{L}(R_1|k_1) $$
#
# And more generally:
#
# $$ P(R_t|k_t) \propto P(R_0) \cdot {\displaystyle \prod^{T}_{t=0}}\mathcal{L}(R_t|k_t) $$
#
# With a uniform prior $P(R_0)$, this reduces to:
#
# $$ P(R_t|k_t) \propto {\displaystyle \prod^{T}_{t=0}}\mathcal{L}\left(R_t|k_t\right) $$
# ### My Proposed Modification
#
# This works fine, but it suffers from an issue: the posterior on any given day is equally influenced by the distant past as much as the recent day. For epidemics that have $R_t>1$ for a long time and then become under control ($R_t<1$), the posterior gets stuck. It cannot forget about the many days where $R_t>1$ so eventually $P(R_t|k)$ asymptotically approaches 1 when we know it's well under 1. The authors note this in the paper as a footnote. Unfortunately this won't work for us. __The most critical thing to know is when we've dipped below the 1.0 threshold!__
#
# So, I propose to only incorporate the last $m$ days of the likelihood function. By doing this, the algorithm's prior is built based on the recent past which is a much more useful prior than the entire history of the epidemic. So this simple, but important change leads to the following:
#
# $$ P(R_t|k_t) \propto {\displaystyle \prod^{T}_{t=T-m}}\mathcal{L}\left(R_t|k_t\right) $$
#
# While this takes the last $m$ priors into account equally, you can decide to apply a windowing function (such as an exponential) to favor recent priors over more distant.
# ### Choosing a Likelihood Function $\mathcal{L}\left(R_t|k_t\right)$
#
# A likelihood function function says how likely a value of $R_t$ is given an observed number of new cases $k$.
#
# Any time you need to model 'arrivals' over some time period of time, statisticians like to use the [Poisson Distribution](https://en.wikipedia.org/wiki/Poisson_distribution). Given an average arrival rate of $\lambda$ new cases per day, the probability of seeing $k$ new cases is distributed according to the Poisson distribution:
#
# $$P(k|\lambda) = \frac{\lambda^k e^{-\lambda}}{k!}$$
# +
# Column vector of k
k = np.arange(0, 70)[:, None]
# Different values of Lambda
lambdas = [10, 20, 30, 40]
# Evaluated the Probability Mass Function (remember: poisson is discrete)
y = sps.poisson.pmf(k, lambdas)
# Show the resulting shape
print(y.shape)
# -
# > __Note__: this was a terse expression which makes it tricky. All I did was to make $k$ a column. By giving it a column for $k$ and a 'row' for lambda it will evaluate the pmf over both and produce an array that has $k$ rows and lambda columns. This is an efficient way of producing many distributions all at once, and __you will see it used again below__!
# +
fig, ax = plt.subplots()
ax.set(title='Poisson Distribution of Cases\n $p(k|\lambda)$')
plt.plot(k, y,
marker='o',
markersize=3,
lw=0)
plt.legend(title="$\lambda$", labels=lambdas);
# -
# The Poisson distribution says that if you think you're going to have $\lambda$ cases per day, you'll probably get that many, plus or minus some variation based on chance.
#
# But in our case, we know there have been $k$ cases and we need to know what value of $\lambda$ is most likely. In order to do this, we fix $k$ in place while varying $\lambda$. __This is called the likelihood function.__
#
# For example, imagine we observe $k=20$ new cases, and we want to know how likely each $\lambda$ is:
# +
k = 20
lam = np.linspace(1, 45, 90)
likelihood = pd.Series(data=sps.poisson.pmf(k, lam),
index=pd.Index(lam, name='$\lambda$'),
name='lambda')
likelihood.plot(title=r'Likelihood $L\left(\lambda|k_t\right)$');
# -
# This says that if we see 20 cases, the most likely value of $\lambda$ is (not surprisingly) 20. But we're not certain: it's possible lambda was 21 or 17 and saw 20 new cases by chance alone. It also says that it's unlikely $\lambda$ was 40 and we saw 20.
#
# Great. We have $\mathcal{L}\left(\lambda_t|k_t\right)$ which is parameterized by $\lambda$ but we were looking for $\mathcal{L}\left(R_t|k_t\right)$ which is parameterized by $R_t$. We need to know the relationship between $\lambda$ and $R_t$
# ### Connecting $\lambda$ and $R_t$
#
# __The key insight to making this work is to realize there's a connection between $R_t$ and $\lambda$__. [The derivation](https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0002185) is beyond the scope of this notebook, but here it is:
#
# $$ \lambda = k_{t-1}e^{\gamma(R_t-1)}$$
#
# where $\gamma$ is the reciprocal of the serial interval ([about 4 days for COVID19](https://wwwnc.cdc.gov/eid/article/26/6/20-0357_article)). Since we know every new case count on the previous day, we can now reformulate the likelihood function as a Poisson parameterized by fixing $k$ and varying $R_t$.
#
# $$ \lambda = k_{t-1}e^{\gamma(R_t-1)}$$
#
# $$\mathcal{L}\left(R_t|k\right) = \frac{\lambda^k e^{-\lambda}}{k!}$$
#
# ### Evaluating the Likelihood Function
#
# To contiue our example, let's imagine a sample of new case counts $k$. What is the likelihood of different values of $R_t$ on each of those days?
# +
k = np.array([20, 40, 55, 90])
# We create an array for every possible value of Rt
R_T_MAX = 12
r_t_range = np.linspace(0, R_T_MAX, R_T_MAX*100+1)
# Gamma is 1/serial interval
# https://wwwnc.cdc.gov/eid/article/26/6/20-0357_article
GAMMA = 1/4
# Map Rt into lambda so we can substitute it into the equation below
# Note that we have N-1 lambdas because on the first day of an outbreak
# you do not know what to expect.
lam = k[:-1] * np.exp(GAMMA * (r_t_range[:, None] - 1))
# Evaluate the likelihood on each day and normalize sum of each day to 1.0
likelihood_r_t = sps.poisson.pmf(k[1:], lam)
likelihood_r_t / np.sum(likelihood_r_t, axis=0)
# Plot it
ax = pd.DataFrame(
data = likelihood_r_t,
index = r_t_range
).plot(
title='Likelihood of $R_t$ given $k$',
xlim=(0,7)
)
ax.legend(labels=k[1:], title='New Cases')
ax.set_xlabel('$R_t$');
# -
# You can see that each day we have a independent guesses for $R_t$. The goal is to combine the information we have about previous days with the current day. To do this, we use Bayes' theorem.
#
# ### Performing the Bayesian Update
#
# To perform the Bayesian update, we need to multiply the likelihood by the prior (which is just the previous day's likelihood) to get the posteriors. Let's do that using the cumulative product of each successive day:
# +
posteriors = likelihood_r_t.cumprod(axis=1)
posteriors = posteriors / np.sum(posteriors, axis=0)
columns = pd.Index(range(1, posteriors.shape[1]+1), name='Day')
posteriors = pd.DataFrame(
data = posteriors,
index = r_t_range,
columns = columns)
ax = posteriors.plot(
title='Posterior $P(R_t|k)$',
xlim=(0,7)
)
ax.legend(title='Day')
ax.set_xlabel('$R_t$');
# -
# Notice how on Day 1, our posterior matches Day 1's likelihood from above? That's because we have no information other than that day. However, when we update the prior using Day 2's information, you can see the curve has moved left, but not nearly as left as the likelihood for Day 2 from above. This is because Bayesian updating uses information from both days and effectively averages the two. Since Day 3's likelihood is in between the other two, you see a small shift to the right, but more importantly: a narrower distribution. We're becoming __more__ confident in our believes of the true value of $R_t$.
#
# From these posteriors, we can answer important questions such as "What is the most likely value of $R_t$ each day?"
most_likely_values = posteriors.idxmax(axis=0)
# We can also obtain the [highest density intervals](https://www.sciencedirect.com/topics/mathematics/highest-density-interval) for $R_t$:
#
# > Note: I apologize in advance for the clunky brute force HDI algorithm. Please let me know if there are better ones out there.
# +
def highest_density_interval(pmf, p=.95):
# If we pass a DataFrame, just call this recursively on the columns
if(isinstance(pmf, pd.DataFrame)):
return pd.DataFrame([highest_density_interval(pmf[col]) for col in pmf],
index=pmf.columns)
cumsum = np.cumsum(pmf.values)
best = None
for i, value in enumerate(cumsum):
for j, high_value in enumerate(cumsum[i+1:]):
if (high_value-value > p) and (not best or j<best[1]-best[0]):
best = (i, i+j+1)
break
low = pmf.index[best[0]]
high = pmf.index[best[1]]
return pd.Series([low, high], index=['Low', 'High'])
hdi = highest_density_interval(posteriors, p=.95)
# -
# Finally, we can plot both the most likely values for $R_t$ and the HDIs over time. This is the most useful representation as it shows how our beliefs change with every day.
# +
ax = most_likely_values.plot(marker='o',
label='Most Likely',
title=f'$R_t$ by day',
c='k',
markersize=4)
ax.fill_between(hdi.index,
hdi['Low'],
hdi['High'],
color='k',
alpha=.1,
lw=0,
label='HDI')
ax.legend();
# -
# We can see that the most likely value of $R_t$ changes with time and the highest-density interval narrows as we become more sure of the true value of $R_t$ over time. Note that since we only had four days of history, I did not apply my windowing modification to this sample. Next, however, we'll turn to a real-world application where this modification is necessary.
# # Real-World Application to South African Data
#
# ### Setup
#
# Load South Africa data from the DSFSI data repo
url = 'https://raw.githubusercontent.com/dsfsi/covid19za/master/data/covid19za_provincial_cumulative_timeline_confirmed.csv'
df = pd.read_csv(url,
parse_dates=['date'],
squeeze=True).sort_index()
# Taking a look at the full country, we need to start the analysis when there are a consistent number of cases each day. Find the last zero new case day and start on the day after that.
#
# Also, case reporting is very erratic based on testing backlogs, etc. To get the best view of the 'true' data we can, I've applied a gaussian filter to the time series. This is obviously an arbitrary choice, but you'd imagine the real world process is not nearly as stochastic as the actual reporting.
# +
province_name = 'total'
def prepare_cases(cases):
new_cases = cases.diff()
smoothed = new_cases.rolling(7,
win_type='gaussian',
min_periods=1,
center=True).mean(std=2).round()
zeros = smoothed.index[smoothed.eq(0)]
if len(zeros) == 0:
idx_start = 0
else:
last_zero = zeros.max()
idx_start = smoothed.index.get_loc(last_zero) + 1
smoothed = smoothed.iloc[idx_start:]
original = new_cases.loc[smoothed.index]
return original, smoothed
cases = pd.Series(df[province_name].values,index=df['date'])
original, smoothed = prepare_cases(cases)
original.plot(title=f"South Africa: {province_name} New Cases per Day",
c='k',
linestyle=':',
alpha=.5,
label='Actual',
legend=True,
figsize=(600/72, 400/72))
ax = smoothed.plot(label='Smoothed',
legend=True)
ax.get_figure().set_facecolor('w')
# -
# ### Running the Algorithm
# Just like the example before, we create lambda based on the previous day's counts from all values of $R_t$. Unlike the previous example, I now evaluate the __log__ of the Poisson. Why? It makes windowing easier.
#
# Since $\log{ab}=\log{a}+\log{b}$, we can do a rolling sum over the last $m$ periods and then exponentiate to get the rolling product of the original values. This does not change any of the numbers – it's just a convenience.
# +
def get_posteriors(sr, window=7, min_periods=1):
lam = sr[:-1].values * np.exp(GAMMA * (r_t_range[:, None] - 1))
# Note: if you want to have a Uniform prior you can use the following line instead.
# I chose the gamma distribution because of our prior knowledge of the likely value
# of R_t.
# prior0 = np.full(len(r_t_range), np.log(1/len(r_t_range)))
prior0 = np.log(sps.gamma(a=3).pdf(r_t_range) + 1e-14)
likelihoods = pd.DataFrame(
# Short-hand way of concatenating the prior and likelihoods
data = np.c_[prior0, sps.poisson.logpmf(sr[1:].values, lam)],
index = r_t_range,
columns = sr.index)
# Perform a rolling sum of log likelihoods. This is the equivalent
# of multiplying the original distributions. Exponentiate to move
# out of log.
posteriors = likelihoods.rolling(window,
axis=1,
min_periods=min_periods).sum()
posteriors = np.exp(posteriors)
# Normalize to 1.0
posteriors = posteriors.div(posteriors.sum(axis=0), axis=1)
return posteriors
posteriors = get_posteriors(smoothed)
# -
# ### The Result
#
# Below you can see every day (row) of the posterior distribution plotted simultaneously. The posteriors start without much confidence (wide) and become progressively more confident (narrower) about the true value of $R_t$
# +
ax = posteriors.plot(title=f'South Africa: {province_name} - Daily Posterior for $R_t$',
legend=False,
lw=1,
c='k',
alpha=.3,
xlim=(0.4,4))
ax.set_xlabel('$R_t$');
# -
# ### Plotting in the Time Domain with Credible Intervals
# Since our results include uncertainty, we'd like to be able to view the most likely value of $R_t$ along with its highest-density interval.
# +
# Note that this takes a while to execute - it's not the most efficient algorithm
hdis = highest_density_interval(posteriors)
most_likely = posteriors.idxmax().rename('ML')
# Look into why you shift -1
result = pd.concat([most_likely, hdis], axis=1)
result.tail()
# +
def plot_rt(result, ax, province_name):
ax.set_title(f"{province_name}")
# Colors
ABOVE = [1,0,0]
MIDDLE = [1,1,1]
BELOW = [0,0,0]
cmap = ListedColormap(np.r_[
np.linspace(BELOW,MIDDLE,25),
np.linspace(MIDDLE,ABOVE,25)
])
color_mapped = lambda y: np.clip(y, .5, 1.5)-.5
index = result['ML'].index.get_level_values('date')
values = result['ML'].values
# Plot dots and line
ax.plot(index, values, c='k', zorder=1, alpha=.25)
ax.scatter(index,
values,
s=40,
lw=.5,
c=cmap(color_mapped(values)),
edgecolors='k', zorder=2)
# Aesthetically, extrapolate credible interval by 1 day either side
lowfn = interp1d(date2num(index),
result['Low'].values,
bounds_error=False,
fill_value='extrapolate')
highfn = interp1d(date2num(index),
result['High'].values,
bounds_error=False,
fill_value='extrapolate')
extended = pd.date_range(start=pd.Timestamp('2020-03-01'),
end=index[-1]+pd.Timedelta(days=1))
ax.fill_between(extended,
lowfn(date2num(extended)),
highfn(date2num(extended)),
color='k',
alpha=.1,
lw=0,
zorder=3)
ax.axhline(1.0, c='k', lw=1, label='$R_t=1.0$', alpha=.25);
# Formatting
ax.xaxis.set_major_locator(mdates.MonthLocator())
ax.xaxis.set_major_formatter(mdates.DateFormatter('%b'))
ax.xaxis.set_minor_locator(mdates.DayLocator())
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_formatter(ticker.StrMethodFormatter("{x:.1f}"))
ax.yaxis.tick_right()
ax.spines['left'].set_visible(False)
ax.spines['bottom'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.margins(0)
ax.grid(which='major', axis='y', c='k', alpha=.1, zorder=-2)
ax.margins(0)
ax.set_ylim(0.0,3.5)
ax.set_xlim(pd.Timestamp('2020-03-01'), result.index.get_level_values('date')[-1]+pd.Timedelta(days=1))
fig.set_facecolor('w')
fig, ax = plt.subplots(figsize=(600/72,400/72))
plot_rt(result, ax, province_name)
ax.set_title(f'Real-time $R_t$ for South Africa: {province_name}')
ax.set_ylim(.5,3.5)
ax.xaxis.set_major_locator(mdates.WeekdayLocator())
ax.xaxis.set_major_formatter(mdates.DateFormatter('%b %d'))
# -
# ### Repeat the Process for Every Province
# +
results = {}
provinces_to_process = list(df.columns.values[2:-2])
provinces_to_process.append('total')
for province_name in provinces_to_process:
clear_output(wait=True)
print(f'Processing {province_name}')
cases = pd.Series(df[province_name].values,index=df['date'])
new, smoothed = prepare_cases(cases)
print('\tGetting Posteriors')
try:
posteriors = get_posteriors(smoothed)
except:
display(cases)
print('\tGetting HDIs')
hdis = highest_density_interval(posteriors)
print('\tGetting most likely values')
most_likely = posteriors.idxmax().rename('ML')
result = pd.concat([most_likely, hdis], axis=1)
results[province_name] = result#.droplevel(0)
clear_output(wait=True)
print('Done.')
# -
# ### Plot All South Africa Provinces
# +
ncols = 5
nrows = int(np.ceil(len(results) / ncols))
# fig, axes = plt.subplots(nrows=nrows, ncols=ncols, figsize=(15, nrows*3))
fig, axes = plt.subplots(nrows=nrows, ncols=ncols, figsize=(15, nrows*3))
for i, (province_name, result) in enumerate(results.items()):
plot_rt(result, axes.flat[i], province_name)
fig.tight_layout()
fig.set_facecolor('w')
# -
# ### Export Data to CSV
# +
overall = None
for province_name, result in results.items():
r = result.copy()
r.index = pd.MultiIndex.from_product([[province_name], result.index])
if overall is None:
overall = r
else:
overall = pd.concat([overall, r])
overall.sort_index(inplace=True)
# Uncomment this line if you'd like to export
# overall.to_csv('data/rt.csv')
# -
# ### Standings
# +
# # As of 4/12
# no_lockdown = [
# 'North Dakota',
# 'South Dakota',
# 'Nebraska',
# 'Iowa',
# 'Arkansas'
# ]
# partial_lockdown = [
# 'Utah',
# 'Wyoming',
# 'Oklahoma'
# ]
FULL_COLOR = [.7,.7,.7]
NONE_COLOR = [179/255,35/255,14/255]
PARTIAL_COLOR = [.5,.5,.5]
ERROR_BAR_COLOR = [.3,.3,.3]
# +
filtered = overall.index.get_level_values(0).isin(FILTERED_REGIONS)
mr = overall.loc[~filtered].groupby(level=0)[['ML', 'High', 'Low']].last()
def plot_standings(mr, figsize=None, title='Most Recent $R_t$ by Province'):
if not figsize:
figsize = (10,4) #((15.9/50)*len(mr)+.1,2.5)
fig, ax = plt.subplots(figsize=figsize)
ax.set_title(title)
err = mr[['Low', 'High']].sub(mr['ML'], axis=0).abs()
bars = ax.bar(mr.index,
mr['ML'],
width=.825,
color=FULL_COLOR,
ecolor=ERROR_BAR_COLOR,
capsize=2,
error_kw={'alpha':.5, 'lw':1},
yerr=err.values.T)
# for bar, state_name in zip(bars, mr.index):
# if state_name in no_lockdown:
# bar.set_color(NONE_COLOR)
# if state_name in partial_lockdown:
# bar.set_color(PARTIAL_COLOR)
labels = mr.index.to_series()
ax.set_xticklabels(labels, rotation=90, fontsize=11)
ax.margins(0)
ax.set_ylim(0,2.)
ax.axhline(1.0, linestyle=':', color='k', lw=1)
leg = ax.legend(handles=[
Patch(label='Full', color=FULL_COLOR),
Patch(label='Partial', color=PARTIAL_COLOR),
Patch(label='None', color=NONE_COLOR)
],
title='Lockdown',
ncol=3,
loc='upper left',
columnspacing=.75,
handletextpad=.5,
handlelength=1)
leg._legend_box.align = "left"
fig.set_facecolor('w')
return fig, ax
mr.sort_values('ML', inplace=True)
plot_standings(mr);
# -
mr.sort_values('High', inplace=True)
plot_standings(mr);
show = mr[mr.High.le(1.1)].sort_values('ML')
fig, ax = plot_standings(show, title='Likely Under Control');
show = mr[mr.Low.ge(1.05)].sort_values('Low')
fig, ax = plot_standings(show, title='Likely Not Under Control');
ax.get_legend().remove()
|
notebooks/Realtime R0.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://qworld.net" target="_blank" align="left"><img src="../qworld/images/header.jpg" align="left"></a>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# $ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
# $ \newcommand{\greenbit}[1] {\mathbf{{\color{green}#1}}} $
# $ \newcommand{\bluebit}[1] {\mathbf{{\color{blue}#1}}} $
# $ \newcommand{\redbit}[1] {\mathbf{{\color{red}#1}}} $
# $ \newcommand{\brownbit}[1] {\mathbf{{\color{brown}#1}}} $
# $ \newcommand{\blackbit}[1] {\mathbf{{\color{black}#1}}} $
# <font style="font-size:28px;" align="left"><b> <font color="blue"> Solutions for </font>Superposition and Measurement </b></font>
# <br>
# _prepared by <NAME>_
# <br><br>
# <a id="task3"></a>
# <h3> Task 3</h3>
#
# Repeat the second experiment with the following modifications.
#
# Start in state $ \ket{1} $.
#
# Apply a Hadamard gate.
#
# Make a measurement.
#
# If the measurement outcome is 0, stop.
#
# Otherwise, apply a second Hadamard, and then make a measurement.
#
# Execute your circuit 1000 times.
#
# Calculate the expected values of observing '0' and '1', and then compare your result with the simulator result.
# <h3> Solution </h3>
# +
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
# define a quantum register with a single qubit
q = QuantumRegister(1)
# define a classical register with a single bit
c = ClassicalRegister(1,"c")
# define a quantum circuit
qc = QuantumCircuit(q,c)
# start in state |1>
qc.x(q[0])
# apply the first Hadamard
qc.h(q[0])
# the first measurement
qc.measure(q,c)
# apply the second Hadamard if the measurement outcome is 1
qc.h(q[0]).c_if(c,1)
# the second measurement
qc.measure(q[0],c)
# draw the circuit
display(qc.draw(output="mpl"))
# -
# We expect to see outcome '0' and '1' with frequency %75 and %25, respectively.
# +
# execute the circuit 1000 times in the local simulator
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=1000)
counts = job.result().get_counts(qc)
print(counts)
# -
# <a id="task4"></a>
# <h3> Task 4</h3>
#
# Design the following quantum circuit.
#
# Start in state $ \ket{0} $.
#
# Repeat 3 times:
# if the classical bit is 0:
# apply a Hadamard operator
# make a measurement
#
# Execute your circuit 1000 times.
#
# Calculate the expected values of observing '0' and '1', and then compare your result with the simulator result.
# <h3> Solution </h3>
# +
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
# define a quantum register with a single qubit
q = QuantumRegister(1,"q")
# define a classical register with a single bit
c = ClassicalRegister(1,"c")
# define a quantum circuit
qc = QuantumCircuit(q,c)
qc.x(q[0])
for i in range(3):
qc.h(q[0]).c_if(c,0)
qc.measure(q,c)
# draw the circuit
qc.draw(output="mpl")
# -
# We start in state $ \ket{0} $. Thus, the first Hadamard and measurement are implemented.
#
# Out of 1000, we expect to observe 500 '0' and 500 '1'.
#
# If the classical bit is 1, then there will be no further Hadamard operator, and so the quantum register will always be in state $ \ket{1} $ and so all measurements results will be 1.
#
# If the classical bit is 0, then another Hadamard applied followed by a measuement.
#
# Thus, out ouf 1000, we expect to observe 250 '0' and 750 '1'.
#
# Similarly, after the third control, we expect to observe 125 '0' and 875 '1'.
# +
# execute the circuit 1000 times in the local simulator
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=10280)
counts = job.result().get_counts(qc)
print(counts)
# -
# <a id="task5"></a>
# <h3> Task 5</h3>
#
# Design the following randomly created quantum circuit.
#
# Start in state $ \ket{0} $.
#
# apply a Hadamard operator
# make a measurement
# REPEAT 4 times:
# randomly pick x in {0,1}
# if the classical bit is x:
# apply a Hadamard operator
# make a measurement
#
# Draw your circuit, and guess the expected frequency of observing '0' and '1' if the circuit is executed 10000 times.
#
# Then, execute your circuit 10000 times, and compare your result with the simulator result.
#
# Repeat execution a few more times.
# <h3> Solution </h3>
#
# We can calculate the frequencies iteratively by python.
# +
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
# import randrange for random choices
from random import randrange
# define a quantum register with a single qubit
q = QuantumRegister(1)
# define a classical register with a single bit
c = ClassicalRegister(1,"c")
# define a quantum circuit
qc = QuantumCircuit(q,c)
shot = 10000
observe = [0,0]
qc.h(q[0])
qc.measure(q,c)
observe = [shot/2,shot/2]
for i in range(4):
x = randrange(2)
if x==0:
observe[0] = observe[0] / 2
observe[1] = observe[1] + observe[0]
else:
observe[1] = observe[1] / 2
observe[0] = observe[0] + observe[1]
qc.h(q[0]).c_if(c,x)
qc.measure(q,c)
# draw the circuit
display(qc.draw(output="mpl"))
# +
print('0:',round(observe[0]),'1:',round(observe[1]))
# execute the circuit 10000 times in the local simulator
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=shot)
counts = job.result().get_counts(qc)
print(counts)
|
quantum-with-qiskit/Q36_Superposition_and_Measurement_Solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Perlin 2d Example
# ## To run this example install the following packages
#
# ```
# # !pip install perlin-cupy
# # !pip install matplotlib
# # !pip install numpy
# ```
# ## Import packages
# +
import matplotlib.pyplot as plt
import numpy as np
from perlin_cupy import (
generate_perlin_noise_2d, generate_fractal_noise_2d
)
# -
# ## Generate Perlin Noise
np.random.seed(0)
noise = generate_perlin_noise_2d((256, 256), (8, 8))
plt.imshow(noise.get(), cmap='gray', interpolation='lanczos')
plt.colorbar()
plt.show()
# ## Generate Perlin Noise Fractal
np.random.seed(0)
noise = generate_fractal_noise_2d((256, 256), (8, 8), 5)
plt.figure()
plt.imshow(noise.get(), cmap='gray', interpolation='lanczos')
plt.colorbar()
plt.show()
|
docs/examples/perlin 2d example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Traffic Sign Recognition**
#
# **Build a Traffic Sign Recognition Project**
#
# This is another project of the Self-Driving cars nanodegree at Udacity.
#
# [//]: # 'Image References'
# [classes_chart]: ./examples/classes_chart.png 'Classes chart'
# [processed_img]: ./examples/processed_img.png 'Processing'
# [de_signs]: ./examples/traffic_imgs.png 'Traffic Signs'
# [image8]: ./examples/placeholder.png 'Traffic Sign 5'
#
#
# ### Data Set Summary & Exploration
#
# #### 1. Dataset summary
#
# I used the python and pandas library to calculate summary statistics of the traffic
# signs data set:
#
# | Description | Value |
# | :----------------------------------------------------- | :---------------- |
# | Input | 32x32x3 RGB image |
# | The size of training set is | 34799 |
# | The size of the validation set is | 4410 |
# | The size of test set is | 12630 |
# | The shape of a traffic sign image is | (32, 32, 3) |
# | The number of unique classes/labels in the data set is | 43 |
#
# #### 2. Dataset distribution
#
# Here is an exploratory visualization of the data set. It is a bar chart showing how the data is distributed over the classes
#
# ![alt text][classes_chart]
#
# ### Design and Test a Model Architecture
#
# #### 1. Preprocess
#
# As a first step, I decided to convert the images to grayscale to reduce the amount of inputs by reducing channels of the image.
#
# Here is an example of a traffic sign image before and after grayscaling.
#
# ![alt text][processed_img]
#
# I normalized the image data to maintain data near to 0 and simplify the training process
#
# The difference between the original data set and the augmented data set is the following ...
#
# #### 2. Architecture
#
# My final model consisted of the following layers:
#
# | Layer | Description |
# | :-------------- | :------------------------------------------ |
# | Input | 32x32x1 RGB image |
# | Convolution 5x5 | 1x1 stride, valid padding, outputs 28x28x16 |
# | RELU | droupout prob 0.8 after if training |
# | Max pooling | 5x5 stride, outputs 14x14x16 |
# | Convolution 5x5 | 1x1 stride, valid padding, outputs 10x10x64 |
# | RELU | droupout prob 0.8 after if training |
# | Max pooling | 5x5 stride, outputs 5x5x |
# | Convolution 2x2 | 1x1 stride, valid padding, outputs 4x4x120 |
# | RELU | droupout prob 0.8 after if training |
# | Max pooling | 2x2 stride, outputs 2x2x120 |
# | Flattening | output 480 |
# | Fully connected | output 120 |
# | Fully connected | output 43 (classes) |
# | Softmax | |
#
# #### 3. How you trained my model.
#
# To train the model, I used an adam optimizer and apply 128 samples batch size. To improve the accuracy I repeat the training fo 30 ephocs with a 0.001 lerning rate value.
#
# #### 4. Approach
#
# My final model results were:
#
# - training set accuracy of 0.996
# - validation set accuracy of 0.957
# - test set accuracy of 0.946
#
# The first architecture was the LeNet but was not enough to get more than 0.89 accuracy.
# I tried to use the YUV color space as in this [paper](http://yann.lecun.com/exdb/publis/pdf/sermanet-ijcnn-11.pdf). But with my actual architecture was still under 0.85~0.87 of accuracy. By adding another convolutional layer the accuracy increase. After that I added a dropout to make the recognition more flexible. The accuracy rise the 0.93 accuracy araound the 20th ephoc but i decided leave 30 to consolidate the pattern.
#
#
# ### Test a Model on New Images
#
# #### 1. Choose five German traffic signs on the web
#
# Here are five German traffic signs that I found on Google Maps moving around Hamburg by taking sreenshots.
#
# ![alt text][de_signs]
#
# The prediction was pretty accurate.
#
# #### 2. Model's predictions
#
# Here are the results of the prediction:
#
# | Image | Prediction |
# | :------------------: | :------------------: |
# | Yield | Yield |
# | Keep right | Keep right |
# | Road work | Road work |
# | No entry | No entry |
# | Speed limit (30km/h) | Speed limit (30km/h) |
#
# The model was able to correctly guess 5 of the 5 traffic signs, which gives an accuracy of 100%.
#
# #### 3. Probabilities of predictions
#
# The code for making predictions on my final model is located in the 11th cell of the Ipython notebook.
#
# For the first image, the model is has no doubt, and also others images returns similar values on probabilities.
#
# | Probability | Prediction |
# | :---------- | :------------------------------------------- |
# | 1.000000 | Yield |
# | 0.000000 | No vehicles |
# | 0.000000 | No passing for vehicles over 3.5 metric tons |
# | 0.000000 | Right-of-way at the next intersection |
# | 0.000000 | Ahead only |
#
# But let's se the last one that introduce a little uncertainty on the value of the speed limit.
#
# | Probability | Prediction |
# | :---------- | :--------------------------------------- |
# | 0.911139 | Speed limit (30km/h) |
# | 0.077434 | Speed limit (50km/h) |
# | 0.005195 | Speed limit (20km/h) |
# | 0.001936 | Vehicles over 3.5 metric tons prohibited |
# | 0.001237 | Speed limit (80km/h) |
#
#
|
README.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rodrigowe1988/Data-Science-na-Pratica/blob/main/Limpeza_de_dados%2C_outliers_e_valores_ausentes.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="2bcpjHZO4w3t"
# # Limpeza de dados, outliers e valores ausentes
#
# Neste projeto, vamos focar em uma habilidade básica, mas essencial de um Cientista de Dados. Saber lidar com valores ausentes, lidar com *outliers*, transformar os dados. Essas são algumas das atividades que fazem parte do checklist de todos os projetos que executamos.
#
#
# Esses processos podem ter um impacto enorme nos resultados, desde a parte da análise dos dados até os modelos de *Machine Learning*.
# <p align="center"><img src="https://image.freepik.com/free-vector/group-analysts-working-graphs_1262-21249.jpg
# "></p>
#
#
# Aqui, veremos o que buscar entender nos dados, os passos a dar, como identificar, e tratar *outliers*, as melhores práticas e métodos para lidar com dados ausentes, e por fim, converter os dados para os formatos necessários.
#
# + [markdown] id="HkCMRDLF5UvU"
# ## O Que Analisar?
#
# Primeiro de tudo, precisamos entender o que são bons dados, para que saibamos os passos que precisamos tomar para ter o melhor conjunto de dados possível, a partir dos dados que estamos utilizando.
#
# * Tipo de Dados
# * Coluna de data em formato `datetime`, coluna de valores monetários em `float`.
# * *Range* dos Dados
# * Meses de 1 a 12, dias do mês de 1 a 31.
# * Dados Obrigatórios
# * Algumas colunas não podem estar vazias
# * Dados Únicos
# * CPF, RG, CNPJ, ID de usuário.
# * Dados Categóricos
# * Gênero Masculino ou Feminino.
# * Padrões regulares
# * (61) 9 8765-4321
# * Validade Entre os Campos
# * Data de saída não ser anterior à data de entrada.
#
# + id="2u6rYOSl4c6E"
# importando os pacotes necessários
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# configurando a visualização
sns.set_style()
# %matplotlib inline
# + [markdown] id="gwWHHJQMC_R-"
# ## Os Dados
#
# Utilizaremos mais de um conjunto de dados nesse projeto, mas o primeiro deles será do mesmo grupo usado no Projeto do Módulo 2 do curso Data Science na Prática.
#
# Nesse projeto, analisamos dados sobre a violência no Rio de Janeiro.
#
# <p align="center"><img src="https://image.freepik.com/free-vector/brazilian-carnival-concept-with-dancing-people-nature_1284-27444.jpg
# ", width="50%"></p>
#
#
# Esse conjunto de dados é interessante pois está organizado em números de crimes por mês, desde janeiro de 1991, mas alguns dos crimes só começaram a ser registrados algum tempo depois, como veremos abaixo.
# + id="9-42E02p6bt9" colab={"base_uri": "https://localhost:8080/", "height": 275} outputId="ba625db1-89ee-4551-bd68-c29d69698b7b"
# importando os dados
df = pd.read_csv('https://raw.githubusercontent.com/carlosfab/curso_data_science_na_pratica/master/modulo_02/violencia_rio.csv', sep=',')
# verificando as dimensões
print('Dimensões do Dataset',
'\nVariáveis: ',df.shape[1], "\n"
'Entradas: ', df.shape[0])
# verificando as primeiras entradas
df.head()
# + id="EayvbHidJLEb" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="87e64823-0ee6-445d-cb65-fdce12c7dde2"
# verificando o final do dataset
df.tail()
# + id="fUmzdSwa9Sc0" colab={"base_uri": "https://localhost:8080/"} outputId="0e1f0927-3de1-443f-d3af-bfeae37bf359"
# modificando o tipo
df.lesao_corp_morte[343].astype('int64')
# + id="htk9-VHy8BBf" colab={"base_uri": "https://localhost:8080/"} outputId="92271d06-986b-4a52-d995-8d197a9c04eb"
# pandas usa floats quando tem dados ausentes
df.dtypes
# + [markdown] id="OKfC7GRqB5vl"
# ## Como lidar com Dados Ausentes
#
# A resposta é: Depende!
#
# Que tipo de dado está ausente? Em qual proporção? De forma aleatória? Todos esses são aspectos que precisamos levar em consideração ao tratar dados ausentes.
#
# Tomando os dados do RJ como exemplo, vemos que temos algumas colunas com quase todos os dados ausentes, mas qual o motivo disso? De onde esses dados são extraídos e qual o processo de coleta deles? Dados ausentes implicam algum significado?
#
# Nesse caso, nossa teoria mais predominante é que os dados não começaram a ser registrados até uma determinada data, e após isso, a coleta foi feita de forma efetiva.
# + id="FsyTQC3WD5WK" colab={"base_uri": "https://localhost:8080/"} outputId="178b41e7-1f54-4ee4-b8d4-557c1e6cc58d"
# mostrando a quantidade de dados ausentes por variável
(df.isnull().sum()).sort_values(ascending=False)
# + id="9kDe0AxF3hbh" colab={"base_uri": "https://localhost:8080/", "height": 999} outputId="7802a944-10f2-4f64-eeca-e47327250f20"
# dados ausentes por ano
df.set_index('vano').isna().sum(level=0)
# + id="cqME7vVEEPvI" colab={"base_uri": "https://localhost:8080/", "height": 906} outputId="7c225738-b8d5-45d6-fa1c-2852da7db31c"
# visualizando
df[273:300]
# + [markdown] id="ZidjjcglHXzM"
# ## Tratando os Dados
#
# No caso desse conjunto de dados específico, o ideal é analisar os dados apenas do período em que se tem dados. Especialmente em algumas variáveis onde o volume é muito grande, qualquer tipo de preenchimento poderia enviesar os dados de forma que a análise deixasse de ser relevante.
#
# Para outros casos, podemos considerar as seguintes hipóteses:
#
# * Excluir
# * Se os dados ausentes estão em pequeno número,ocorrem aleatoriamente, e a ausência não carrega significado, é melhor excluir a linha. No caso da coluna, se ainda for possível analisar alguma parte dela, use-a, como é o caso aqui. Mas para algumas situações, o ideal é excluir a coluna.
#
# * Preencher
# * Preencher as entradas com dados ausentes com valores estatísticos como a média, mediana, moda ou zeros.
# * A média é mais útil quando a distribuição dos dados é normal. Em dados com distribuição mais enviesada (*skewed*), a mediana é uma solução mais robusta, pois ela é menos sensível a outliers.
# * Uma `Regressão Linear` também pode ser útil, apesar de sensível a outliers, podem nos ajudar a inserir valores que nos ajudem.
# * Indetificar a entrada ausente com algum valor que indique isso pode ser mais informativo, quando a ausência representa valor. Por exemplo, em dados numéricos preencher com zero, e em categóricos criar uma categoria "Desconhecido". Atenção, pois os zeros não podem ser levados em consideração em análises estatísticas.
# + [markdown] id="3V8xTTjogf1w"
# ## Tratando Outliers
#
# <p align="center"><img src="https://miro.medium.com/max/18000/1*2c21SkzJMf3frPXPAR_gZA.png
# ", width="50%"></p>
#
#
# *Outliers* são pontos discrepantes, que estão destoando do padrão do conjunto de dados.
#
# É muito importante conseguir identificar e tratar esses outliers, pois eles podem nos mostrar uma imagem incorreta dos nossos dados.
#
# Podemos identificar um outlier de diversas formas, entre elas podemos citar:
#
# * IQR Score
# * Boxplots
# * Scatter plots
# * Z-Score
#
# Vamos ver na prática o processo completo de limpeza de dados, tratando dados ausentes, convertendo os dados para o formato correto, e tratando dos outliers.
# + [markdown] id="gezBu5ZKRNRC"
# ## Airbnb - NYC
#
# Para esse estudo, faremos o tratamento dos dados do Airbnb referentes à cidade de Nova Iorque.
#
# <center><img alt="New York City" width="50%" src="https://image.freepik.com/free-vector/future-metropolis-downtown-modern-city-business-center-cartoon-background_33099-1466.jpg"></center>
# + id="d7LSSj19As1R" colab={"base_uri": "https://localhost:8080/", "height": 428} outputId="1c7c4a50-fd2c-46e1-bc29-be8aee650a34"
# importando os dados
df_nyc = pd.read_csv('https://raw.githubusercontent.com/rafaelnduarte/eds_outliers/master/nyc.csv', index_col=0)
# verificando as dimensões
print('Dimensões do Dataset',
'\nVariáveis: ',df_nyc.shape[1], "\n"
'Entradas: ', df_nyc.shape[0])
# verificando as primeiras entradas
df_nyc.head()
# + id="PW9hb1MBDYlP" colab={"base_uri": "https://localhost:8080/"} outputId="4a74ee33-a586-416b-d64d-74ed83e5d635"
# verificando os tipos
df_nyc.dtypes
# + id="yH6CdFHYB3Bt" colab={"base_uri": "https://localhost:8080/"} outputId="c922127b-9cf5-482f-daea-5efa15a93ea7"
# transformando as colunas
df_nyc[['price','latitude', 'longitude']] = df_nyc[['price','latitude', 'longitude']].astype('object')
# verificando o resultado;l
df_nyc.dtypes
# + id="7M0vkoedDcs7" colab={"base_uri": "https://localhost:8080/"} outputId="cf948062-a011-4284-d15e-45f085c47790"
# verificando dados ausentes
(df_nyc.isnull().sum()).sort_values(ascending=False)
# + [markdown] id="5P7YMj0jSDHq"
# Seguindo o que falamos anteriormente, colunas com baixo poder preditivo e grande quantidade de dados faltantes podem ser excluídas.
#
# Em relação às outras entradas, estão em poucas entradas, não parecem ter algum tipo de relação entre elas, e não parecem ter grande poder preditivo. Mais uma vez, vamos fazer a exclusão. Porém, aqui vamos excluir as entradas.
# + id="s5Kv4vjWt1pD"
# excluindo colunas com dados faltantes
df_nyc.drop(columns=['reviews_per_month', 'last_review'], inplace=True)
# excluindo entradas com dados faltantes
df_nyc.dropna(axis=0, inplace=True)
# convertendo os tipos de dados
df_nyc[['price','latitude', 'longitude']] = df_nyc[['price','latitude', 'longitude']].astype('float')
# + [markdown] id="ElwypmbGSfOS"
# Feita a limpeza, é hora de conferir os resultados.
# + id="kPGqXFB1uMEn" colab={"base_uri": "https://localhost:8080/"} outputId="e16c7b4e-33cc-4007-de9a-15eeca2f819d"
# verificando o resultado
(df_nyc.isnull().sum()).sort_values(ascending=False)
# + [markdown] id="kQ7EMdQ6xsAz"
# ## Limpando Outliers
#
# Para tratar dos outliers desse conjunto de dados, iremos analisar a distribuição estatística, plotar boxplots e calcular os limites utilizando a regra do IQR Score.
#
# Primeiramente, vamos lembrar o que é o IQR.
#
# O IQR é calculado subtraindo o Terceiro Quartil (75%) pelo Primeiro Quartil (25%).
#
# # IQR = Q3 - Q1
#
# Vamos dar uma olhada nos nossos dados e ver o que identificamos.
# + id="00UrR7dBvjCn" colab={"base_uri": "https://localhost:8080/", "height": 317} outputId="a3037dfb-e547-4db3-82d0-0284e9915604"
# verificando a distribuição estatística
df_nyc.describe().round(1)
# + [markdown] id="zGkdeB_eXaSs"
# Aqui, algumas coisas já chamam a nossa atenção, como por exemplo:
#
# * A variável `price` tem o mínimo em 0.
# * Lembrando que a variável `price` trata do preço da diária dos imóveis em moeda local (USD), estamos vendo que o Q3 está em 175 dólares, mas o máximo está em 10 mil dórales. Claramente, há outliers por aqui.
# * A variável `minimum_nights` tem como seu máximo o valor 1250, sendo que o Q3 está em 6. Claramente temos outliers nessa variável.
# * As variáveis `number_of_reviews`, `calculated_host_listings_count` e `availability_365` também podem conter outliers, mas não vamos nos preocupar com elas agora.
# + id="5N-LxhFtu3vG" colab={"base_uri": "https://localhost:8080/", "height": 871} outputId="6e39f06c-8402-49f7-8453-6ea13db3dda4"
# verificando as distribuições
df_nyc.hist(figsize=(20,15), grid=False);
# + [markdown] id="0Kr88iOlYils"
# Verificando os histogramas, conseguimos ver claramente que temos outliers presentes. Para tratá-los vamos seguir os seguintes passos:
#
# * Definir o Q1 e Q3 para as variáveis que serão limpas.
# * Calcular o IQR para as variáveis.
# * Definir o limite superior e inferior para cortar os outliers.
# * Remover os outliers.
#
# + id="G6fuoxkI23Ss" colab={"base_uri": "https://localhost:8080/"} outputId="b148fc51-55e1-47bc-efe7-672a93005f4e"
# identificando os outliers para a variável price
q1_price = df_nyc.price.quantile(.25)
q3_price = df_nyc.price.quantile(.75)
IQR_price = q3_price - q1_price
print('IQR da variável price: ', IQR_price)
# definindo os limites
sup_price = q3_price + 1.5 * IQR_price
inf_price = q1_price - 1.5 * IQR_price
print('Limite superior de price: ', sup_price)
print('Limite inferior de price: ', inf_price)
# + [markdown] id="KEWG6kfaY_Z7"
# Aqui podemos ver que, apesar de não termos outliers na parte inferior, continuamos tendo valores iguais a zero, que precisam ser tratados.
#
# Vamos plotar um boxplot para visualizarmos a diferença feita pela limpeza.
# + id="XGvT72qpHUOP" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="69d90dad-eb5d-4eb5-8a46-f0163990ca41"
# verificando o conjunto original
fig, ax = plt.subplots(figsize=(15,3))
df_nyc.price.plot(kind='box', vert=False);
ax.set_title('Dataset Original - price')
plt.show()
print("O dataset possui {} colunas".format(df_nyc.shape[0]))
print("{} Entradas acima de 335.5".format(len(df_nyc[df_nyc.price > 335.5])))
print("Representam {:.2f}% do dataset".format((len(df_nyc[df_nyc.price > 335.5]) / df_nyc.shape[0])*100))
# + id="TFlvqW6w5iJC" colab={"base_uri": "https://localhost:8080/"} outputId="f6a0de52-19ac-48f6-ee71-f26b0b416ab3"
# identificando os outliers para a variável minimum_nights
q1_minimum_nights = df_nyc.minimum_nights.quantile(.25)
q3_minimum_nights = df_nyc.minimum_nights.quantile(.75)
IQR_minimum_nights = q3_minimum_nights - q1_minimum_nights
print('IQR da variável minimum_nights: ', IQR_minimum_nights)
# definindo os limites
sup_minimum_nights = q3_minimum_nights + 1.5 * IQR_minimum_nights
inf_minimum_nights = q1_minimum_nights - 1.5 * IQR_minimum_nights
print('Limite superior de minimum_nights: ', sup_minimum_nights)
print('Limite inferior de minimum_nights: ', inf_minimum_nights)
# + id="Hr_FUZ3mHV1o" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="4b9841e3-0e74-4d06-eb42-66d1ca2cb163"
# verificando o conjunto original
fig, ax = plt.subplots(figsize=(15,3))
df_nyc.minimum_nights.plot(kind='box', vert=False);
ax.set_title('Dataset Original - minimum_nights')
plt.show()
print("O dataset possui {} colunas".format(df_nyc.shape[0]))
print("{} Entradas acima de 335.5".format(len(df_nyc[df_nyc.minimum_nights > 12.0])))
print("Representam {:.2f}% do dataset".format((len(df_nyc[df_nyc.minimum_nights > 12.0]) / df_nyc.shape[0])*100))
# + id="TWc_6lXt6csi" colab={"base_uri": "https://localhost:8080/"} outputId="2a4d8eb0-670d-40a2-e607-8ab406f1e150"
# limpando o dataset
df_clean = df_nyc.copy()
df_clean.drop(df_clean[df_clean.price > 335.5].index, axis=0, inplace=True)
df_clean.drop(df_clean[df_clean.price == 0.0].index, axis=0, inplace=True)
df_clean.drop(df_clean[df_clean.minimum_nights > 12].index, axis=0, inplace=True)
print('Shape antes da limpeza: ', df_nyc.shape)
print('Shape após a limpeza: ',df_clean.shape)
# + id="28OVII13EKrq" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="d83988e4-cf39-45a3-de30-4196611d79fe"
# plotando novamente o boxplot original
fig, ax = plt.subplots(figsize=(15,3))
df_nyc.price.plot(kind='box', vert=False);
ax.set_title('Dataset Original - price')
plt.show()
print("O dataset possui {} colunas".format(df_nyc.shape[0]))
print("{} Entradas acima de 335.5".format(len(df_nyc[df_nyc.price > 335.5])))
print("Representam {:.2f}% do dataset".format((len(df_nyc[df_nyc.price > 335.5]) / df_nyc.shape[0])*100))
# + id="jaKGqOm3Dgrf" colab={"base_uri": "https://localhost:8080/", "height": 261} outputId="ea6a49ef-227e-48a6-80ba-6ccad73d05b1"
# verificando o conjunto limpo
fig, ax = plt.subplots(figsize=(15,3))
df_clean.price.plot(kind='box', vert=False);
ax.set_title('Dataset Limpo - price')
plt.show()
print("Tamanho original: {} colunas".format(df_nyc.shape[0]))
print("Tamanho atual: {} colunas".format(df_clean.shape[0]))
# + [markdown] id="W7Gkhc6ybC-a"
# Aqui podemos ver o resultado da limpeza.
#
# O boxplot mostra alguns dados como outliers, entretanto, esses estão sendo calculados em relação ao novo dataset, e nossa limpeza levou em consideração os quartis do dado original.
#
# Para garantirmos que não estamos lidando com outliers que vão prejudicar nossa análise, vamos checar os histogramas novamente.
#
# + id="8eUuufyf3qL6" colab={"base_uri": "https://localhost:8080/", "height": 870} outputId="83ccf849-e2e7-44b9-e028-bccac075e9fa"
# verificando as distribuições
df_clean.hist(figsize=(20,15), grid=False);
# + id="gipucX3o4bqE" colab={"base_uri": "https://localhost:8080/", "height": 317} outputId="6e094a24-a3df-4ef9-8297-fd45a55cfb32"
# verificando a distribuição estatística dos dados limpos
df_clean.describe().round(1)
# + [markdown] id="qipTpXZqbjwZ"
# Agora conseguimos ter uma ideia bem melhor da distribuição dos nossos dados.
#
# Alguns destaques:
#
# * A mediana da variável `price` foi pouquíssimo afetada pela limpeza dos outliers, mostrando mais uma vez a robustez desse atributo como solução para dados ausentes.
# * Agora, temos dados que respeitam as regras definidas no início do notebook, onde vimos o que são bons dados.
# * A média da variável `price` foi reduzida drásticamente, enfatizando a sensibilidade desse atributo em relação aos outliers.
# + [markdown] id="oE4qoumRcnNi"
# ## Informações Importantes
#
# * Modelos lineares são mais sensíveis aos outliers. Ao trabalhar com modelos desse tipo é essencial que o trabalho com outliers seja feito com bastante atenção. (Linear Regression, Logistic Regression)
#
# * Modelos baseados em árvores de decisão são menos sensívels a outliers. (Decision Trees, Random Forest, SVM, KNN)
#
# * Técnicas de pré-processamento como Normalização e Padronização ajudam em relação aos outliers.
|
Limpeza_de_dados,_outliers_e_valores_ausentes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["remove_cell"]
# # Hybrid quantum-classical Neural Networks with PyTorch and Qiskit
# -
# Machine learning (ML) has established itself as a successful interdisciplinary field which seeks to mathematically extract generalizable information from data. Throwing in quantum computing gives rise to interesting areas of research which seek to leverage the principles of quantum mechanics to augment machine learning or vice-versa. Whether you're aiming to enhance classical ML algorithms by outsourcing difficult calculations to a quantum computer or optimise quantum algorithms using classical ML architectures - both fall under the diverse umbrella of quantum machine learning (QML).
#
# In this chapter, we explore how a classical neaural network can be partially quantized to create a hybrid quantum-classical neural network. We will code up a simple example which integrates qiskit with a state-of-the-art open source software package - [PyTorch](https://pytorch.org/). The purpose of this example is to demonstrate the ease of integrating Qiskit with existing ML tools and to encourage ML practitioners to explore what is possible with quantum computing.
# ## How does it work?
# <img src="hybridnetwork.png" width="800"/>
#
# **Fig.1** Illustrates the framework we will construct in this chapter. Ultimately, we will create a hybrid quantum-classical neural network that seeks to classify hand drawn digits. Note that the edges shown in this image are all directed downward; however, the directionality is not visually indicated.
# ### Preliminaries:
# The background presented here on classical neural networks is included to establish relevant ideas and shared terminology; however, it is still extremely high-level. __If you'd like to dive one step deeper into classical neural networks, see the well made video series by youtuber__ [3Blue1Brown](https://youtu.be/aircAruvnKk). Alternatively, if you are already familiar with classical networks, you can [skip to the next section](#quantumlayer).
#
# ###### Neurons and Weights
# A neural network is ultimately just an elaborate function that is built by composing smaller building blocks called neurons. A ***neuron*** is typically a simple, easy-to-compute, and nonlinear function that maps one or more inputs to a single real number. The single output of a neuron is typically copied and fed as input into other neurons. Graphically, we represent neurons as nodes in a graph and we draw directed edges between nodes to indicate how the output of one neuron will be used as input to other neurons. It's also important to note that each edge in our graph is often associated with a scalar-value called a [***weight***](https://en.wikipedia.org/wiki/Artificial_neural_network#Connections_and_weights). The idea here is that each of the inputs to a neuron will be multiplied by a different scalar before being collected and processed into a single value. The objective when training a neural network consists primarily of choosing our weights such that the network behaves in a particular way.
#
# ###### Feed Forward Neural Networks
# It is also worth noting that the particular type of neural network we will concern ourselves with is called a **[feed-forward neural network (FFNN)](https://en.wikipedia.org/wiki/Feedforward_neural_network)**. This means that as data flows through our neural network, it will never return to a neuron it has already visited. Equivalently, you could say that the graph which describes our neural network is a **[directed acyclic graph (DAG)](https://en.wikipedia.org/wiki/Directed_acyclic_graph)**. Furthermore, we will stipulate that neurons within the same layer of our neural network will not have edges between them.
#
# ###### IO Structure of Layers
# The input to a neural network is a classical (real-valued) vector. Each component of the input vector is multiplied by a different weight and fed into a layer of neurons according to the graph structure of the network. After each neuron in the layer has been evaluated, the results are collected into a new vector where the i'th component records the output of the i'th neuron. This new vector can then treated as input for a new layer, and so on. We will use the standard term ***hidden layer*** to describe all but the first and last layers of our network.
#
# ## So how does quantum enter the picture? <a id='quantumlayer'> </a>
#
# To create a quantum-classical neural network, one can implement a hidden layer for our neural network using a parameterized quantum circuit. By "parameterized quantum circuit", we mean a quantum circuit where the rotation angles for each gate are specified by the components of a classical input vector. The outputs from our neural network's previous layer will be collected and used as the inputs for our parameterized circuit. The measurement statistics of our quantum circuit can then be collected and used as inputs for the following layer. A simple example is depicted below:
#
# <img src="neuralnetworkQC.png" width="800"/>
#
# Here, $\sigma$ is a [nonlinear function](https://en.wikipedia.org/wiki/Activation_function) and $h_i$ is the value of neuron $i$ at each hidden layer. $R(h_i)$ represents any rotation gate about an angle equal to $h_i$ and $y$ is the final prediction value generated from the hybrid network.
#
# ### What about backpropagation?
# If you're familiar with classical ML, you may immediately be wondering *how do we calculate gradients when quantum circuits are involved?* This would be necessary to enlist powerful optimisation techniques such as **[gradient descent](https://en.wikipedia.org/wiki/Gradient_descent)**. It gets a bit technical, but in short, we can view a quantum circuit as a black box and the gradient of this black box with respect to its parameters can be calculated as follows:
#
# <img src="quantumgradient.png" width="800"/>
#
# where $\theta$ represents the parameters of the quantum circuit and $s$ is a macroscopic shift. The gradient is then simply the difference between our quantum circuit evaluated at $\theta+s$ and $\theta - s$. Thus, we can systematically differentiate our quantum circuit as part of a larger backpropogation routine. This closed form rule for calculating the gradient of quantum circuit parameters is known as **[the parameter shift rule](https://arxiv.org/pdf/1905.13311.pdf)**.
# # Let's code!
#
# ### Imports
# First, we import some handy packages that we will need, including Qiskit and PyTorch.
import numpy as np
import torch
from torch.autograd import Function
import torch.optim as optim
import torch.nn as nn
import torch.nn.functional as F
import torchvision
from torchvision import datasets, transforms
from qiskit import QuantumRegister, QuantumCircuit, ClassicalRegister, execute
from qiskit.circuit import Parameter
from qiskit import Aer
from matplotlib import pyplot as plt
# %matplotlib inline
# ### Tensors to lists
# Next we create an additional function that converts a tensor to a list in Python. This is needed to connect Qiskit and PyTorch objects. In particular, we will use this function to convert tensors produced by PyTorch to a list, such that they can be fed into quantum circuits in Qiskit.
def to_numbers(tensor_list):
num_list = []
for tensor in tensor_list:
num_list += [tensor.item()]
return num_list
# ### Create a "quantum class" with Qiskit
# We can conveniently put our Qiskit quantum functions into a class. First, we specify how many trainable quantum parameters and how many shots we wish to use in our quantum circuit. In this example, we will keep it simple and use a 1-qubit circuit with one trainable quantum parameter $\theta$. We hard code the circuit for simplicity and use a $RY-$rotation by the angle $\theta$ to train the output of our circuit. The circuit looks like this:
#
# <img src="1qubitcirc.png" width="400"/>
#
# In order to measure the output in the $z-$basis, we create a Python function to obtain the $\sigma_z$ expectation. Lastly, we create a "bind" function to convert our parameter to a list and run the circuit on the Aer simulator. We will see later how this all ties into the hybrid neural network.
class QiskitCircuit():
# Specify initial parameters and the quantum circuit
def __init__(self,shots):
self.theta = Parameter('Theta')
self.shots = shots
def create_circuit():
qr = QuantumRegister(1,'q')
cr = ClassicalRegister(1,'c')
ckt = QuantumCircuit(qr,cr)
ckt.h(qr[0])
ckt.barrier()
ckt.ry(self.theta,qr[0])
ckt.barrier()
ckt.measure(qr,cr)
return ckt
self.circuit = create_circuit()
def N_qubit_expectation_Z(self, counts, shots, nr_qubits):
expects = np.zeros(nr_qubits)
for key in counts.keys():
perc = counts[key]/shots
check = np.array([(float(key[i])-1/2)*2*perc for i in range(nr_qubits)])
expects += check
return expects
def bind(self, parameters):
[self.theta] = to_numbers(parameters)
self.circuit.data[2][0]._params = to_numbers(parameters)
def run(self, i):
self.bind(i)
backend = Aer.get_backend('qasm_simulator')
job_sim = execute(self.circuit,backend,shots=self.shots)
result_sim = job_sim.result()
counts = result_sim.get_counts(self.circuit)
return self.N_qubit_expectation_Z(counts,self.shots,1)
# ### Create a "quantum-classical class" with PyTorch
# Now that our quantum circuit is defined, we can create the functions needed for backpropagation using PyTorch. [The forward and backward passes](http://www.ai.mit.edu/courses/6.034b/backprops.pdf) contain elements from our Qiskit class. The backward pass directly computes the analytical gradients using the finite difference formula we introduced above.
class TorchCircuit(Function):
@staticmethod
def forward(ctx, i):
if not hasattr(ctx, 'QiskitCirc'):
ctx.QiskitCirc = QiskitCircuit(shots=100)
exp_value = ctx.QiskitCirc.run(i[0])
result = torch.tensor([exp_value]) # store the result as a torch tensor
ctx.save_for_backward(result, i)
return result
@staticmethod
def backward(ctx, grad_output):
s = np.pi/2
forward_tensor, i = ctx.saved_tensors
# Obtain paramaters
input_numbers = to_numbers(i[0])
gradient = []
for k in range(len(input_numbers)):
input_plus_s = input_numbers
input_plus_s[k] = input_numbers[k] + s # Shift up by s
exp_value_plus = ctx.QiskitCirc.run(torch.tensor(input_plus_s))[0]
result_plus_s = torch.tensor([exp_value_plus])
input_minus_s = input_numbers
input_minus_s[k] = input_numbers[k] - s # Shift down by s
exp_value_minus = ctx.QiskitCirc.run(torch.tensor(input_minus_s))[0]
result_minus_s = torch.tensor([exp_value_minus])
gradient_result = (result_plus_s - result_minus_s)
gradient.append(gradient_result)
result = torch.tensor([gradient])
return result.float() * grad_output.float()
# Putting this all together
# We will create a simple hybrid neural network to classify images of two types of digits (0 or 1) from the [MNIST dataset](http://yann.lecun.com/exdb/mnist/). We first load MNIST and filter for pictures containing 0's and 1's. These will serve as inputs for our neural network to classify.
#
# ### Data loading and preprocessing
# +
transform = torchvision.transforms.Compose([torchvision.transforms.ToTensor()]) # transform images to tensors/vectors
mnist_trainset = datasets.MNIST(root='./data', train=True, download=True, transform=transform)
labels = mnist_trainset.targets # get the labels for the data
labels = labels.numpy()
idx1 = np.where(labels == 0) # filter on zeros
idx2 = np.where(labels == 1) # filter on ones
# Specify number of datapoints per class (i.e. there will be n pictures of 1 and n pictures of 0 in the training set)
n=100
# concatenate the data indices
idx = np.concatenate((idx1[0][0:n],idx2[0][0:n]))
# create the filtered dataset for our training set
mnist_trainset.targets = labels[idx]
mnist_trainset.data = mnist_trainset.data[idx]
train_loader = torch.utils.data.DataLoader(mnist_trainset, batch_size=1, shuffle=True)
# -
# The data will consist of images belonging to two classes: 0 and 1. An example image from both classes looks like this:
#
# <img src="MNISTplot.png" width="100"/>
#
# So far, we have loaded the data and coded a class that creates our quantum circuit which contains 1 trainable parameter. This quantum parameter will be inserted into a classical neural network along with the other classical parameters to form the hybrid neural network. We also created backward and forward pass functions that allow us to do backpropagation and optimise our neural network. Lastly, we need to specify our neural network architecture such that we can begin to train our parameters using optimisation techniques provided by PyTorch.
#
#
# ### Creating the hybrid neural network
# We can use a neat PyTorch pipeline to create a neural network architecture. The network will need to be compatible in terms of its dimensionality when we insert the quantum layer (i.e. our quantum circuit). Since our quantum in this example contains 1 parameter, we must ensure the network condenses neurons down to size 1. We create a network consisting of 3 hidden layers with 320, 50 and 1 neurons respectively. The value of the last neuron is fed as the parameter $\theta$ into our quantum circuit. The circuit measurement then serves as the final prediction for 0 or 1 as provided by a $\sigma_z$ measurement. The measurement outcomes are -1 which implies a predicted label of 0 and 1 which implies a predicted label of 1.
# +
qc = TorchCircuit.apply
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 10, kernel_size=5)
self.conv2 = nn.Conv2d(10, 20, kernel_size=5)
self.conv2_drop = nn.Dropout2d()
self.h1 = nn.Linear(320, 50)
self.h2 = nn.Linear(50, 1)
def forward(self, x):
x = F.relu(F.max_pool2d(self.conv1(x), 2))
x = F.relu(F.max_pool2d(self.conv2_drop(self.conv2(x)), 2))
x = x.view(-1, 320)
x = F.relu(self.h1(x))
x = F.dropout(x, training=self.training)
x = self.h2(x)
x = qc(x)
x = (x+1)/2 # Normalise the inputs to 1 or 0
x = torch.cat((x, 1-x), -1)
return x
# -
# ### Training the network
# We now have all the ingredients to train our hybrid network! We can specify any [PyTorch optimiser](https://pytorch.org/docs/stable/optim.html), [learning rate](https://en.wikipedia.org/wiki/Learning_rate) and [cost/loss function](https://en.wikipedia.org/wiki/Loss_function) in order to train over multiple epochs. In this instance, we use the [Adam optimiser](https://arxiv.org/abs/1412.6980), a learning rate of 0.001 and the [negative log-likelihood loss function](https://pytorch.org/docs/stable/_modules/torch/nn/modules/loss.html).
# +
network = Net()
optimizer = optim.Adam(network.parameters(), lr=0.001)
epochs = 30
loss_list = []
for epoch in range(epochs):
total_loss = []
target_list = []
for batch_idx, (data, target) in enumerate(train_loader):
target_list.append(target.item())
optimizer.zero_grad()
output = network(data)
loss = F.nll_loss(output, target)
loss.backward()
optimizer.step()
total_loss.append(loss.item())
loss_list.append(sum(total_loss)/len(total_loss))
print(loss_list[-1])
# Normalise the loss between 0 and 1
for i in range(len(loss_list)):
loss_list[i] += 1
# -
# Plot the loss per epoch
plt.plot(loss_list)
plt.title("Hybrid NN Training Convergence")
plt.xlabel('Training Iterations')
plt.ylabel('Loss')
# # What now?
#
# While it is totally possible to create hybrid neural networks, does this actually have any benefit? In fact, the classical layers of this network train perfectly fine (in fact, better) without the quantum layer. Furthermore, you may have noticed that the quantum layer we trained here **generates no entanglement**, and will therefore continue to be classically simulable as we scale up this particular architecture. This means that if you hope to achieve a quantum advantage using hybrid nerural networks, you'll need to start by extending this code to include a more sophisticated quantum layer.
#
#
# The point of this exercise was to get you thinking about integrating techniques from ML and quantum computing in order to investigate if there is indeed some element of interest - and thanks to PyTorch and Qiskit, this becomes a little bit easier.
|
content/ch-machine-learning/machine-learning-qiskit-pytorch.ipynb
|
# + [markdown] deletable=true editable=true
# Largest Rectangle in Bar Chart puzzle
# + deletable=true editable=true
>./ (* >:@i.@#) >./&> +/;.1 each <"1 |: 0, (2 3 4 2 5 1) #"0 (1)
# + deletable=true editable=true
[ largest_rectangle =. 13 : '>./ (* >:@i.@#) >./&> +/;.1 each <"1 |: 0, y #"0 (1)'
# + deletable=true editable=true
largest_rectangle 2 3 4 2 5 1
# + [markdown] deletable=true editable=true
# String division - https://codegolf.stackexchange.com/questions/129259/divide-a-string
# Test cases
#
# "Hello, world!", 4 -> (["Hel", "lo,", " wo", "rld"], "!") ("!" is the remainder)
# "Hello, world!", 5 -> (["He", "ll", "o,", " w", "or"], "ld!")
# "ABCDEFGH", 2 -> (["ABCD", "EFGH"], "") (no remainder; optional "")
# "123456789", 5 -> (["1", "2", "3", "4", "5"], "6789")
# "ALABAMA", 3 -> (["AL", "AB", "AM"], "A")
# "1234567", 4 -> (["1", "2", "3", "4"], "567")
# -
test1 =. test2 =. 'Hello, world!'
test3 =. 'ABCDEFGH'
test4 =. '123456789'
test5 =. 'ALABAMA'
test6 =. '1234567'
(0 5 #: #) test2
13 : '(0 , x) #: # y'
5 ((0,[)#:[:#]) test2
5 (] $~ [ , (<.@%~#)) test2
4 ((] $~ [ , (<.@%~#)) ; (] {.~ 0 - (|#) ) ) test1
chunk =. (]$~[,(<.@%~#));]{.~0-(|#)
5 chunk test2
4 chunk test1
5 chunk test2
2 chunk test3
5 chunk test4
3 chunk test5
4 chunk test6
chunk
|
Puzzles.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:metis] *
# language: python
# name: conda-env-metis-py
# ---
# +
import os
import re
import glob
import time
import numpy as np
import pandas as pd
pd.set_option("display.max_columns", None)
from bs4 import BeautifulSoup
from selenium import webdriver
import matplotlib.pyplot as plt
import seaborn as sns
# +
# file not available locally on the GitHub repo, must be downloaded from http://www.robesafe.uah.es/personal/eduardo.romera/uah-driveset/#download
data_path = 'UAH-DRIVESET-v1/'
files = glob.glob(data_path+'*/*/SEMANTIC_ONLINE*.txt', recursive=True)
# -
def get_class(drive_class):
drive_class = re.sub(r'[0-9]', '', drive_class)
if drive_class == 'NORMAL':
return drive_class
else:
return 'NOT NORMAL'
# +
df = pd.DataFrame()
for i,file in enumerate(files):
drive_info = file.split('/')[2].split('-')
drive_date = drive_info[0]
dist_km = drive_info[1][:-2]
driver = drive_info[2]
drive_class = get_class(drive_info[3])
road_type = drive_info[4]
row = {'date_time_raw':drive_date, 'total_dist':dist_km, 'driver_id':driver, 'road_type':road_type, 'class_normal':drive_class}
df = df.append(row, ignore_index=True)
df['date_time_clean'] = pd.to_datetime(df['date_time_raw']).dt.round('30min')
df['date_time_clean'] = pd.to_datetime(df['date_time_clean']).dt.strftime('%Y-%m-%d %I:%M %p')
df
# -
def rendering(url):
driver = webdriver.Chrome('/usr/local/bin/chromedriver') # run ChromeDriver
driver.get(url) # load the web page from the URL
time.sleep(3) # wait for the web page to load
render = driver.page_source # get the page source HTML
driver.quit() # quit ChromeDriver
return render # return the page source HTML
df.info()
cols = ['temp_F', 'dp_F', 'humidity_pct', 'wind_direction', 'wind_speed_mph', 'wind_gust_mph', 'pressure_in', 'precipitation_in', 'condition']
for c in cols:
df[c] = np.nan
def get_wunderground(i, row):
drive_date = row['date_time_clean'][:10]
drive_time = row['date_time_clean'][11:].lstrip('0')
search_url = f'http://www.wunderground.com/history/daily/es/madrid/LEMD/date/{drive_date}'
wunderground_page = rendering(search_url)
wunderground_soup = BeautifulSoup(wunderground_page, 'html.parser')
soup_container = wunderground_soup.find('lib-city-history-observation')
soup_data = soup_container.find_all('tr')
for j,dat in enumerate(soup_data[1:]):
for k,td in enumerate(dat.find_all('td', class_='ng-star-inserted')):
tmp = td.text
if k == 0:
row_time = tmp
if (k != 0) and (row_time == drive_time):
df.loc[i, cols[k-1]] = tmp.split(u'\xa0°')[0]
while len(df[df[cols[0]].isna()]) != 0:
for i, row in df.iterrows():
get_wunderground(i, row)
df
df.to_csv('UAH-DRIVESET-weather.csv')
|
scrape_wunderground.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os, sys, string
from collections import Counter
textFiles = [f for f in os.listdir('.') if '.txt' in f] # get list of every file in current directory that ends with .txt
rawCorpus=[]
for f in textFiles:
file = open(f,'rt', encoding='utf-8', errors='replace') # open each text file for reading
print (f" Reading from: '{f}' . . .")
rawCorpus.append(file.read()
.replace('. . .','.')
.replace('!',' .') # substitue space period for ! mark to have a simple token to end a sentence
.replace('"',' ')
.replace('#',' ')
.replace('$',' ')
.replace('%',' ')
.replace('&',' ')
.replace('\\',' ')
.replace('\' ',' ') # only remove ' if it has a space before or after meaning it is used as a quote
.replace(' \'',' ') # but leave it in if it is inside a word as a contraction
.replace('(',' ')
.replace(')',' ')
.replace('*',' ')
.replace('+',' ')
.replace(',',' ')
.replace('-',' ')
.replace('. ',' . ') # add a space so the period becomes a token to end a sentence
.replace('/',' ')
.replace(':',' ')
.replace(';',' ')
.replace('<',' ')
.replace('=',' ')
.replace('>',' ')
.replace('?',' .') # substitue space period for ? mark to have a simple token to end a sentence
.replace('@',' ')
.replace('[',' ')
.replace('\\',' ')
.replace(']',' ')
.replace('^',' ')
.replace('_',' ') # remove all unwanted punctuation
.replace('`',' ')
.replace('{',' ')
.replace('|',' ')
.replace('}',' ')
.replace('~',' ')
.replace('0',' ') # remove all digits
.replace('1',' ')
.replace('2',' ')
.replace('3',' ')
.replace('4',' ')
.replace('5',' ')
.replace('6',' ')
.replace('7',' ')
.replace('8',' ')
.replace('9',' '))
file.close()
corpus = ' '.join(rawCorpus)
print(len(corpus))
tokens=corpus.split()
output = {}
n=1
for i in range(len(tokens) - n + 1):
g = ' '.join(tokens[i:i+n])
output.setdefault(g, 0)
output[g] += 1
COUNTS = Counter(output)
print(f"\n The most common unigrams are: {(COUNTS.most_common(10))}")
f=open("unigramfile-smaller.dat","w+", encoding='utf-8', errors='replace')
f.write(str(sum(COUNTS.values())))
f.write(str(COUNTS.most_common(3000))) #trying to keep file size at about 50 k for this sample
f.close()
output = {}
n=2
for i in range(len(tokens) - n + 1):
g = ' '.join(tokens[i:i+n])
output.setdefault(g, 0)
output[g] += 1
COUNTS = Counter(output)
print(f"\n The most common bigrams are: {(COUNTS.most_common(10))}")
f=open("bigramfile-smaller.dat","w+", encoding='utf-8', errors='replace')
f.write(str(sum(COUNTS.values())))
f.write(str(COUNTS.most_common(2700))) #trying to keep file size at about 50 k for this sample
f.close()
output = {}
n=3
for i in range(len(tokens) - n + 1):
g = ' '.join(tokens[i:i+n])
output.setdefault(g, 0)
output[g] += 1
COUNTS = Counter(output)
print(f"\n The most common trigrams are: {(COUNTS.most_common(10))}")
f=open("trigramfile-smaller.dat","w+", encoding='utf-8', errors='replace')
f.write(str(sum(COUNTS.values())))
f.write(str(COUNTS.most_common(2300))) #trying to keep file size at about 50 k for this sample
f.close()
output = {}
n=4
for i in range(len(tokens) - n + 1):
g = ' '.join(tokens[i:i+n])
output.setdefault(g, 0)
output[g] += 1
COUNTS = Counter(output)
print(f"\n The most common quadgrams are: {(COUNTS.most_common(10))}")
f=open("quadgramfile-smaller.dat","w+", encoding='utf-8', errors='replace')
f.write(str(sum(COUNTS.values())))
f.write(str(COUNTS.most_common(1900))) #trying to keep file size at about 50 k for this sample
f.close()
# -
|
Phase 4 generate text/Generate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Searching for best models by tuning params
# ## Libraries
import pandas as pd
import numpy as np
import sklearn
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
np.random.seed(123)
# ## Reading processed data (script 'data-preprocessing')
data = pd.read_csv('../processed_data/out.csv')
# ## Dividing into train and test
# To divide set so in both parts with have simmilar amounts of big and small credits we need to put them into groups:
# (in traditional splits, randomization makes results appear very uneven)
# +
from sklearn import preprocessing
x = data[['credit_amount']].values.astype(float)
min_max_scaler = preprocessing.MinMaxScaler()
x_scaled = min_max_scaler.fit_transform(x)
data['amount_groups'] = x_scaled
bins = [0.0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8]
data['amount_groups'] = np.digitize(data['amount_groups'], bins)
unique, counts = np.unique(data['amount_groups'], return_counts=True)
dict(zip(unique, counts))
# -
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data.drop(['customer_type', 'amount_groups'], axis=1), data.customer_type, test_size=0.20, stratify = data[['amount_groups', 'customer_type']])
# ## Needed functions
# ### Encoding
import category_encoders as ce
class Error(Exception):
pass
class NonMatchingLengthsError(Error):
pass
def multiEnc(X_train, X_test, target_train, cols, encodings):
"""
Lista znaków do "encodings":
d - backward difference
n - base N
b - binary
c - cat boost
# - hashing
h - helmert
j - James-Stein
l - leave one out
m - m-estimate
1 - one-hot
o - ordinal
p - polynomial
s - sum coding
t - target encoding
w - weight of evidence
"""
ce_map = {"d": ce.backward_difference.BackwardDifferenceEncoder,
"n": ce.basen.BaseNEncoder,
"b": ce.binary.BinaryEncoder,
"c": ce.cat_boost.CatBoostEncoder,
"#": ce.hashing.HashingEncoder,
"h": ce.helmert.HelmertEncoder,
"j": ce.james_stein.JamesSteinEncoder,
"l": ce.leave_one_out.LeaveOneOutEncoder,
"m": ce.m_estimate.MEstimateEncoder,
"1": ce.one_hot.OneHotEncoder,
"o": ce.ordinal.OrdinalEncoder,
"p": ce.polynomial.PolynomialEncoder,
"s": ce.sum_coding.SumEncoder,
"t": ce.target_encoder.TargetEncoder,
"w": ce.woe.WOEEncoder}
try:
if len(cols)!=len(encodings):
raise(NonMatchingLengthsError)
except NonMatchingLengthsError:
print("Lengths do not match")
return None
e=0
for c in cols:
if X_train[c].dtypes=='object':
enc=ce_map[encodings[e]](cols=c)
enc=enc.fit(X_train, target_train)
X_train=enc.transform(X_train)
X_test=enc.transform(X_test)
e=e+1
return (X_train, X_test)
# ### Models creation
# +
from abc import (ABC,
abstractmethod,
abstractproperty)
from typing import Any
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import GradientBoostingClassifier
from xgboost import XGBClassifier # Inna paczka niż sklearn!
import xgboost as xgb
class Builder(ABC):
@abstractproperty
def product(self) -> None:
pass
class Product():
def __init__(self) -> None:
self.parts = []
def add(self, part: Any) -> None:
self.parts.append(part)
def list_parts(self):
return self.parts
class ConcreteBuilder(Builder):
def __init__(self) -> None:
self.reset()
def reset(self) -> None:
self._product = Product()
@property
def product(self) -> Product:
product = self._product
self.reset()
return product
def _add_model(self, model_type: str, params: dict):
if model_type == 'logistic regression':
self._logistic_regression(params)
if model_type == 'decision tree':
self._decision_tree(params)
if model_type == 'svm':
self._svm(params)
if model_type == 'naive bayes':
self._naive_bayes(params)
if model_type == 'random forest':
self._random_forest(params)
if model_type == 'ada boost':
self._ada_boost(params)
if model_type == 'gradient boost':
self._gradient_boost(params)
if model_type == 'xgboost':
self._xgboost(params)
def _logistic_regression(self, params: dict):
lr = LogisticRegression(**params)
return self._product.add(lr)
def _decision_tree(self, params: dict):
dt = DecisionTreeClassifier(**params)
dt_params = {'criterion': dt.criterion}
return self._product.add(dt)
def _svm(self, params: dict):
svm = SVC(**params)
return self._product.add(svm)
def _naive_bayes(self, params: dict):
nb = GaussianNB(**params)
return self._product.add(nb)
def _random_forest(self, params: dict):
rf = RandomForestClassifier(**params)
return self._product.add(rf)
def _ada_boost(self, params: dict):
ada = AdaBoostClassifier(**params)
return self._product.add(ada)
def _gradient_boost(self, params: dict):
gb= GradientBoostingClassifier(**params)
return self._product.add(gb)
def _xgboost(self, params: dict):
xg=XGBClassifier(**params)
return self._product.add(xg)
class Director:
def __init__(self) -> None:
self._builder = None
@property
def builder(self) -> Builder:
return self._builder
@builder.setter
def builder(self, builder: Builder) -> None:
self._builder = builder
def add_model(self, model_type, params):
return self.builder._add_model(model_type, params)
def add_all_models(self):
self.add_model('logistic regression', {})
self.add_model('decision tree', {})
self.add_model('svm', {})
self.add_model('naive bayes', {})
self.add_model('random forest', {})
self.add_model('ada boost', {})
self.add_model('gradient boost', {})
self.add_model('xgboost', {})
return builder.product.list_parts()
def get_all_models(self, metric_name: str = ''):
parts = builder.product.list_parts()
return parts
# -
director = Director()
builder = ConcreteBuilder()
director.builder = builder
# ### Business metrics to rate models (simmilar to F1, but more focused on earnings for the bank)
ir_loan = 0.13
lgd = 0.38
def calculateEarningsLosses(X_test, y_pred, y_test):
'''
As declared, takes test data and predicted classes and calculates:
- earnings made by following prediction
- losses made by following prediction
- earnings omited by following prediction
- losses omited by following prediction
'''
amounts = X_test['credit_amount']
balance_all = y_test.apply(lambda x: ir_loan if x==1 else -lgd) * amounts
earnings_made = balance_all.iloc[np.logical_and(y_test==1, y_pred==1).array].sum()
earnings_omitted = balance_all.iloc[np.logical_and(y_test==1, y_pred==0).array].sum()
losses_made = balance_all.iloc[np.logical_and(y_test==0, y_pred==1).array].sum()
losses_omitted = balance_all.iloc[np.logical_and(y_test==0, y_pred==0).array].sum()
results = pd.DataFrame(columns=['Earnings made', 'Earnings omitted', 'Losses made', 'Losses omitted'])
results.loc[0] = [earnings_made, earnings_omitted, losses_made, losses_omitted]
final_balance = earnings_made + losses_made
max_income = balance_all.iloc[(y_test==1).array].sum()
perc_of_max_income = final_balance/max_income
return (results, final_balance, perc_of_max_income)
# ## Search for best type of encoding
columns_enc = ['checking_account_status', 'credit_history', 'purpose', 'savings', 'present_employment',
'personal', 'other_debtors', 'property', 'other_installment_plans',
'housing', 'job', 'telephone']
def encoding_list_gen(nominal, ordinal):
enc = [ordinal, nominal, nominal, ordinal, nominal, nominal, nominal, nominal, nominal, nominal, nominal, nominal]
return enc
# we group columns as ordinal and nominal to encode them with the same type
#enc = ['j', '1', '1', 'j', '1', '1', '1', '1', '1', '1', '1', '1']
X_test.columns
# Previous attempts have shown, that XGB, GradientBoosting, RandomForest and SVM are the classifiers to aim at. In following cells we will evaluate those three algorithms with different encoding types. Because most of them are tree based, OneHot will be used only to test SVM.
# ### Testing function
# +
from typing import List
from sklearn.metrics import f1_score
def compare_models(models_list: List, X_train, y_train, X_test, y_test, categorical_variables, encoding_list):
results = dict()
df_train, df_test = multiEnc(X_train, X_test, y_train, categorical_variables, encoding_list)
for model in models_list:
training = model.fit(df_train, y_train)
score = training.score(df_test, y_test)
f1 = f1_score(y_test, model.predict(df_test))
_, _, business = calculateEarningsLosses(X_test, model.predict(df_test), y_test)
results[model] = (['score:', score], ['f1:', f1], ['business:', business])
return results
# -
def compare_encoders(model, X_train, y_train, X_test, y_test, columns_enc, enc_nominal, enc_ordinal):
results = pd.DataFrame(columns = enc_nominal, index = enc_ordinal)
results_f1 = pd.DataFrame(columns = enc_nominal, index = enc_ordinal)
results_bus = pd.DataFrame(columns = enc_nominal, index = enc_ordinal)
for nom in enc_nominal:
for ordi in enc_ordinal:
encoding_list = encoding_list_gen(nom, ordi)
df_train, df_test = multiEnc(X_train, X_test, y_train, columns_enc, encoding_list)
training = model.fit(df_train, y_train)
score = training.score(df_test, y_test)
f1 = f1_score(y_test, model.predict(df_test))
_, _, business = calculateEarningsLosses(X_test, model.predict(df_test), y_test)
results.loc[ordi, nom] = score
results_f1.loc[ordi, nom] = f1
results_bus.loc[ordi, nom] = business
return (results, results_f1, results_bus)
director.add_model('gradient boost', {})
director.add_model('xgboost', {})
director.add_model('random forest', {})
director.add_model('svm', {})
models = director.get_all_models()
enc_nominal = ['l', 'j', 'm'] # encoding types to check for nominal values
enc_ordinal = ['p', 'h', 'd', 'l', 'j'] # encoding types to check for ordinal values
# we don't check any encodings leading to information loss
# ### Gradient boosting encoders fit
(score, f1, buss) = compare_encoders(models[0], X_train, y_train, X_test, y_test, columns_enc, enc_nominal, enc_ordinal)
score
f1
buss
# ### XGB encoders fit
(score, f1, buss) = compare_encoders(models[1], X_train, y_train, X_test, y_test, columns_enc, enc_nominal, enc_ordinal = ['l', 'j'])
# XGB has some issues with ordinal encoders so we omit them
score
f1
buss
# ### Random Forest encoders fit
(score, f1, buss) = compare_encoders(models[2], X_train, y_train, X_test, y_test, columns_enc, enc_nominal, enc_ordinal)
score
f1
buss
# ### SVM encoders fit
enc_nominal.append('o')
(score, f1, buss) = compare_encoders(models[3], X_train, y_train, X_test, y_test, columns_enc, enc_nominal, enc_ordinal)
score
f1
buss
# ### Results
# According to score and f1 score all of the algorithms perform well. However most important aspect for potenital client is how much money it would make them. In that case only two algorithms have some potential in improving which are:
# **Gradient boosting** with encoding: ordinal: **polynominal** encoding, nominal: **James-Stein** (all performed the same, I just picked that one)
# **Random Forest** with encoding: ordinal: **backward difference**, nominal: **m-estimate**
# Rest algorithms present very low income levels or even lossess.
# ## Tuning parameters for selected algorithms
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
# Just scoring for bussiness
def bussScore(y, y_pred, X):
amounts = X['credit_amount']
balance_all = y.apply(lambda x: ir_loan if x==1 else -lgd) * amounts
earnings_made = balance_all.iloc[np.logical_and(y==1, y_pred==1).array].sum()
losses_made = balance_all.iloc[np.logical_and(y==0, y_pred==1).array].sum()
final_balance = earnings_made + losses_made
max_income = balance_all.iloc[(y==1).array].sum()
perc_of_max_income = final_balance/max_income
return perc_of_max_income
# Making our own scorer for parameter tuning
b_scorer = make_scorer(bussScore, greater_is_better=True)
# Setting encodings
gboost_enc = encoding_list_gen('j', 'p')
rforest_enc = encoding_list_gen('m', 'b')
# ### Gradient boosting tuning
parameters = {
"loss":["deviance"],
"learning_rate": [0.1, 0.15, 0.2],
"min_samples_split": [0.01, 0.03, 0.05],
"min_samples_leaf": [0.01, 0.02, 0.03],
"max_depth":[3,5,8],
"max_features":["log2","sqrt"],
"criterion": ["friedman_mse", "mae"],
"subsample":[0.8, 0.85, 0.9, 1.0],
"n_estimators":[10, 100]
}
df_train, df_test = multiEnc(X_train, X_test, y_train, columns_enc, gboost_enc)
grid = GridSearchCV(estimator=models[0], param_grid = parameters, scoring = 'f1', cv=4, n_jobs=-1)
grid_result = grid.fit(df_train, y_train)
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
# These are the best parameters to be used for Grid Search. Let's check their bussiness scorer.
best_model_gboost = grid_result.best_estimator_
best_model_gboost.score(df_test, y_test)
bussScore(y_test, best_model_gboost.predict(df_test), df_test)
# ### Random Forest tuning
parameters = {
"min_samples_split": [0.01, 0.03, 0.05],
"min_samples_leaf": [0.01, 0.02, 0.03],
"max_depth":[3,5,8],
"max_features":["log2","sqrt"],
"criterion": ["gini", "mae"],
"n_estimators":[10, 100, 150],
"ccp_alpha":[0.0, 0.01, 0.1]
}
df_train, df_test = multiEnc(X_train, X_test, y_train, columns_enc, rforest_enc)
grid = GridSearchCV(estimator=models[2], param_grid = parameters, scoring = 'f1', cv=4, n_jobs=-1)
grid_result = grid.fit(df_train, y_train)
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
best_model_rforest = grid_result.best_estimator_
best_model_rforest.score(df_test, y_test)
bussScore(y_test, best_model_rforest.predict(df_test), df_test)
# ## Saving results
# +
def save_final_data(df_train, y_train, df_test, y_test):
df_train.to_csv('../final_data/df_train.csv', index=False)
df_test.to_csv('../final_data/df_test.csv', index=False)
y_test.to_csv('../final_data/y_test.csv', index=False)
y_train.to_csv('../final_data/y_train.csv', index=False)
save_final_data(df_train, y_train, df_test, y_test)
# +
import pickle
def save_model(model, filename: str):
pickle.dump(model, open('../models/' + str(filename) + '.sav', 'wb'))
# -
save_model(best_model_gboost, 'gradient_boost_best')
save_model(best_model_rforest, 'random_forest_best')
|
Projekty/Projekt1/Grupa1/JakubowskiKorbinSlapek/scripts/params-tuning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 特征缩放
#
# 对于任何基于距离的机器学习模型(正则化回归方法、神经网络,现在是 KMeans ),你都需要缩放数据。
#
# 如果你的数据有一些尺度完全不同的特征,则将会对 KMeans 的聚类结果有很大的影响。
#
# 在本 Notebook 中,你会看到第一手的资料。 首先,让我们导入必要的库。
# +
import numpy as np
import pandas as pd
from sklearn.cluster import KMeans
import matplotlib.pyplot as plt
from sklearn import preprocessing as p
# %matplotlib inline
plt.rcParams['figure.figsize'] = (16, 9)
import helpers2 as h
import tests as t
# Create the dataset for the notebook
data = h.simulate_data(200, 2, 4)
df = pd.DataFrame(data)
df.columns = ['height', 'weight']
df['height'] = np.abs(df['height']*100)
df['weight'] = df['weight'] + np.random.normal(50, 10, 200)
# -
# `1.` 接下来,查看数据来熟悉它。 这个数据集有个两列,它被存储在变量 df 中。 了解当前数据的分布以及可视化这些点可能很有用。
# +
#Take a look at the data
# +
#use this cell if you would like as well
# -
# 现在我们已经有了一个数据集,让我们看看一些用于缩放数据的选项, 以及如何缩放数据。我们将讨论两种非常常见的特征缩放类型:
#
#
# **I. MinMaxScaler**
#
# 在某些情况下,把你的数据看作是百分比数是有用的,即它们与最大值相比的百分比。 在这些情况下,你需要使用 **MinMaxScaler**。
#
#
# **II. StandardScaler**
#
# 另一种非常流行的特征缩放类型是缩放数据使其具有均值为 0、方差为 1的分布。在这些情况下,你需要使用 **StandardScaler**。
#
# 使用 **StandardScaler** 可能更适合此数据。但是,为了在python中练习特征缩放方法,我们将执行这两类缩放操作。
#
# `2.` 首先让我们用 **StandardScaler** 变换来拟合这个数据集。 我将做这个,以便你可以看到如何用 sklearn 进行数据预处理。
df_ss = p.StandardScaler().fit_transform(df) # Fit and transform the data
# +
df_ss = pd.DataFrame(df_ss) #create a dataframe
df_ss.columns = ['height', 'weight'] #add column names again
plt.scatter(df_ss['height'], df_ss['weight']); # create a plot
# -
# `3.` 现在轮到你了。 请尝试将 **MinMaxScaler** 变换拟合到此数据集。 你应该能够从前面的示例得到帮助。
# +
# fit and transform
# +
#create a dataframe
#change the column names
#plot the data
# -
# `4.` 现在让我们来看看 KMeans 是如何根据不同的数据缩放,对数据集进行不同的分组。 当数据缩放不同时,最终你是否得到了不同的聚类?
# +
def fit_kmeans(data, centers):
'''
INPUT:
data = the dataset you would like to fit kmeans to (dataframe)
centers = the number of centroids (int)
OUTPUT:
labels - the labels for each datapoint to which group it belongs (nparray)
'''
kmeans = KMeans(centers)
labels = kmeans.fit_predict(data)
return labels
labels = fit_kmeans(df, 10) #fit kmeans to get the labels
# Plot the original data with clusters
plt.scatter(df['height'], df['weight'], c=labels, cmap='Set1');
# +
#plot each of the scaled datasets
# +
#another plot of the other scaled dataset
# -
# 请在这里写下你的回答!
|
Unsupervised Learning/01 Clustering/Feature_Scaling-zh.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# dr. <NAME> (www.ladisk.si, <EMAIL>)
from pyTrigger import RingBuffer2D, pyTrigger
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# # Showcase of the package pyTrigger
#
# Jan 2018, dr. <NAME> (www.ladisk.si, <EMAIL>)
# Lets prepare some data (impuls):
T = 1.5
N = 1000
w = 0.1
t, dt = np.linspace(-T/2, T/2, N, endpoint=False, retstep=True)
x = np.cos(2*np.pi*t/(2*w*T))
x[np.logical_or(-w*T*0.5>t, t>w*T*0.5)] = 0.
np.random.seed(0)
data = np.array([x + 0.05*(np.random.rand(N)-0.5),
np.roll(x,50) + 0.05*(np.random.rand(N)-0.5)]).T
plt.plot(t,data[:,0], label='Channel 0')
plt.plot(t,data[:,1], label='Channel 1')
plt.legend();
# ## Big chunk example
# Prepare the trigger object:
pt = pyTrigger(rows=300, channels=2, trigger_channel=0, trigger_level=0.5,
trigger_type='up', presamples=50)
pt.add_data(data)
triggered_data = pt.get_data()
len(triggered_data)
plt.plot(t[:len(triggered_data)], triggered_data[:,0], label='Channel 0')
plt.plot(t[:len(triggered_data)], triggered_data[:,1], label='Channel 1')
plt.legend();
# ## Streaming example
pt = pyTrigger(rows=300, channels=2, trigger_channel=0, trigger_level=0.5,
trigger_type='up', presamples=50)
for d in data.reshape((-1, 100, 2)):
if not pt.finished:
pt.add_data(d)
print(f'Triggering started: {pt.triggered}, # of rows to acquire: {pt.rows_left}')
pt.finished
triggered_from_stream = pt.get_data()
plt.plot(t[:len(triggered_from_stream)], triggered_from_stream[:,0], label='Channel 0')
plt.plot(t[:len(triggered_from_stream)], triggered_from_stream[:,1], label='Channel 1')
plt.legend();
|
Showcase - pyTrigger.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .rs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Rust - nixpkgs
// language: Rust
// name: rust_nixpkgs
// ---
// +
let foo = 1336 + 1;
println!("Hello world!");
foo
// -
|
example/Rust/simple.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# <div class="contentcontainer med left" style="margin-left: -50px;">
# <dl class="dl-horizontal">
# <dt>Title</dt> <dd> HeatMap Tap stream example</dd>
# <dt>Description</dt> <dd>A linked streams example demonstrating how use Tap stream on a HeatMap. The data contains the incidence of measles across US states by year and week (obtained from [Project Tycho](http://www.tycho.pitt.edu/)). The HeatMap represents the mean measles incidence per year. On tap the Histogram on the right will generate a Histogram of the incidences for each week in the selected year and state.</dd>
# <dt>Backends</dt> <dd> Bokeh</dd>
# <dt>Tags</dt> <dd> streams, tap, interactive</dd>
# </dl>
# </div>
# +
import pandas as pd
import numpy as np
import holoviews as hv
from holoviews import opts
hv.extension('bokeh', width=90)
# +
# Declare dataset
df = pd.read_csv('http://assets.holoviews.org/data/diseases.csv.gz', compression='gzip')
dataset = hv.Dataset(df, vdims=('measles','Measles Incidence'))
# Declare HeatMap
heatmap = hv.HeatMap(dataset.aggregate(['Year', 'State'], np.mean),
label='Measles Incidence').select(Year=(1928, 2002))
# Declare Tap stream with heatmap as source and initial values
posxy = hv.streams.Tap(source=heatmap, x=1951, y='New York')
# Define function to compute histogram based on tap location
def tap_histogram(x, y):
return hv.Curve(dataset.select(State=y, Year=int(x)), kdims='Week',
label='Year: %s, State: %s' % (x, y))
tap_dmap = hv.DynamicMap(tap_histogram, streams=[posxy])
(heatmap + tap_dmap).opts(
opts.Curve(framewise=True, height=500, line_color='black', width=375, yaxis='right'),
opts.HeatMap(cmap='RdBu_r', fontsize={'xticks': '6pt'}, height=500,
logz=True, tools=['hover'], width=700, xrotation=90)
)
# -
# <center><img src="https://assets.holoviews.org/gifs/examples/streams/bokeh/heatmap_tap.gif" width=600></center>
|
examples/reference/streams/bokeh/Tap.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# # CS::APEX (Algorithm Prototyper and EXperimentor for Cognitive Systems)
#
# CS::APEX is a framework based on synchronous dataflow and event-based message passing that
# aims to speed up prototyping of new robotic algorithms using visual programming aspects.
#
# Example workflow: (click for better quality)
#
# [](https://youtu.be/sNkHnQhNXuU)
#
#
# ## Dataflow Graph and Core Features
#
# Calculations are represented by a nodes in a directed graph with data flowing on
# the directed edges.This execution graph is manipulated using a simple graphical user
# interface that allows spawning and deleting nodes, adding and removing edges and
# visualizating data in the stream. To speed up the prototyping process,
# other features like undo/redo mechanisms and profiling are implemented as well.
# Finally there exists an easy to use parameter system that generates UI controls
# for each block and allows parameter tuning and optimization.
#
#
# ## Functionality via Plug-ins
#
# The framework itself does not provide any predefined computation nodes and does
# not depend on specific message definitions or node types.
# These details are instead implemented in plug-in libraries that extend
# the functionality of the whole system.
#
#
# ## Application to Robotics
#
# The framework is targeted toward use in robotics and is fully compatible with
# [ROS](http://wiki.ros.org/). Configurations generated using the GUI can
# directly be deployed on any ROS-based robotic system.
#
#
# ## Tutorials and more Information
#
# For more information, please refer to the Wiki at
# https://github.com/cogsys-tuebingen/csapex/wiki
#
# the official website at
# http://www.ra.cs.uni-tuebingen.de/forschung/apex/
#
# or contact the maintainers via email.
#
#
# ## Dependencies
#
# We rely on the [catkin](http://wiki.ros.org/catkin) build system
# developed in the [ROS](http://wiki.ros.org/) ecosystem. The core framework
# is independent of ROS, however.
#
# Required for a build are:
# - Linux system (tested with Ubuntu)
# - C++11 compatible compiler (g++, clang++)
# - Qt5 (on Ubuntu: qt5-default libqt5svg5-dev)
# - libraries:
# - boost (program_options, filesystem, system, regex)
# - [classloader](https://github.com/ros/class_loader)
# - TinyXML (on Ubuntu: libtinyxml-dev)
# - yaml-cpp (on Ubuntu: libyaml-cpp-dev)
#
#
# These dependencies can be installed via rosdep (see below.)
#
#
# ## Basic Installation
#
# To get the cs::APEX framework and a set of core plugins, perform the following:
# +
cd ~
mkdir -p ws/csapex/src
cd ws/csapex/src
git clone https://github.com/cogsys-tuebingen/csapex.git
cd csapex/plugins
git clone https://github.com/cogsys-tuebingen/csapex_core_plugins.git
cd ../..
mkdir libs
cd libs
git clone https://github.com/cogsys-tuebingen/cslibs_vision.git
git clone https://github.com/cogsys-tuebingen/cslibs_laser_processing.git
git clone https://github.com/cogsys-tuebingen/cslibs_arff.git
git clone https://github.com/cogsys-tuebingen/cslibs_indexed_storage.git
cd ../..
rosdep install -y -r -i --from-paths src
catkin_make
# -
# ## Creating Documenation
#
# To create the documentation, run
doxygen doc/Doxyfile
# This will generate the documentation at
doc/html/index.html
# ## Contributions
#
# All contributions are welcome, please refer to the CONTRIBUTING.md file.
|
README.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="9b3e43f6f3787243de855b9416a0b5697be6d116"
# # Titanic challenge part 2
# In this kernel, we will be covering all of the steps required to train, tune and assess a random forest model.
#
# [**Part 1**](https://www.kaggle.com/jamesleslie/titanic-eda-wrangling-imputation/notebook) of this series dealt with the pre-processing and manipulation of the data. This notebook will make use of the data sets that were created in the first part.
#
# We will do each of the following:
# - train and test default RF model
# - introduce cross-validation for model training
# - use grid search to optimize hyperparameters
# - submit our predictions for the test set
#
# [**Part 3**](https://www.kaggle.com/jamesleslie/titanic-neural-network-for-beginners/notebook) of this challenge involves fitting and tuning a **neural network** to make predictions.
# + [markdown] _cell_guid="969e5a36-8f64-4129-ba03-7fd19b314ca9" _uuid="cc4b58927770beab136969e526dbbd69a9cc21c8"
# # Table of Contents:
#
# - **1. [Load packages and data](#loading)**
# - **2. [Pre-processing](#pre-processing)**
# - **3. [Random Forest](#random-forest)**
# - **3.1. [Train/test split](#train-test)**
# - **3.2. [Cross-validation](#cv)**
# - **3.3. [Grid search](#grid-search)**
# - **4. [Submit predictions](#submission)**
# + [markdown] _cell_guid="ec3a4ceb-2397-45b3-aafe-a8ffde879888" _uuid="de8369a5716eb80519979ef773ecbb135f66e4b9"
# <a id="loading"></a>
# # 1. Load packages and data
# We will be using the train and test sets that we created in [part 1](https://www.kaggle.com/jamesleslie/titanic-eda-wrangling-imputation/notebook) of this series.
#
# You can find the dataset [here](https://www.kaggle.com/jamesleslie/titanic-cleaned-data).
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _kg_hide-output=true _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib.pyplot import rcParams
import os
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV, cross_val_score
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score, f1_score, precision_score, recall_score
# %matplotlib inline
rcParams['figure.figsize'] = 10,8
sns.set(style='whitegrid', palette='muted',
rc={'figure.figsize': (15,10)})
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
# print(os.listdir("../input"))
# + _uuid="a36589b28132214f8f50534c4eb61a5a66e52bfe"
#print(os.listdir("../input/titanic-cleaned-data"))
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _kg_hide-output=true _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# Load data as Pandas dataframe
train = pd.read_csv('train_clean.csv', )
test = pd.read_csv('test_clean.csv')
df = pd.concat([train, test], axis=0, sort=True)
# + _uuid="bafdb419c8b7f571dbbb9c0b63fd3c52c66c3c1b"
df.head()
# + _uuid="d8f24917231bd2484e4c2a4896f4fbbdcc3e89ab"
def display_all(df):
with pd.option_context("display.max_rows", 1000, "display.max_columns", 1000):
display(df)
display_all(df.describe(include='all').T)
# + _uuid="ca279aca8e6064618de4b7badb8d94366628cf2b"
df['Survived'].value_counts()
# + [markdown] _cell_guid="b1fc54e6-9655-4a05-b147-e2dfe206c7d0" _uuid="732fc7427f32d790561d03b4a6d870d2a7c67013"
# <a id="pre-processing"></a>
# # 2. Encode categorical variables
# We need to convert all categorical variables into numeric format. The categorical variables we will be keeping are `Embarked`, `Sex` and `Title`.
#
# The `Sex` variable can be encoded into single 1-or-0 column, but the other variables will need to be [one-hot encoded](https://hackernoon.com/what-is-one-hot-encoding-why-and-when-do-you-have-to-use-it-e3c6186d008f). Regular label encoding assigns some category labels higher numerical values. This implies some sort of scale (Embarked = 1 is not **more** than Embarked = 0 - it's just _different_). One Hot Encoding avoids this problem.
#
# We will assume that there is some ordinality in the `Pclass` variable, so we will leave that as a single column.
# + _uuid="7d6d38029e57b2b7eecd8978b4f3b9ab2bbf79d9"
sns.countplot(x='Pclass', data=df, palette='hls', hue='Survived')
plt.xticks(rotation=45)
plt.show()
# + _uuid="3b5e6978df59b98a614297659371823a4a5dbbcc"
sns.countplot(x='Sex', data=df, palette='hls', hue='Survived')
plt.xticks(rotation=45)
plt.show()
# + _uuid="74e4e4d6a937c94fda45dcd135c8f2936707c8d6"
sns.countplot(x='Embarked', data=df, palette='hls', hue='Survived')
plt.xticks(rotation=45)
plt.show()
# + _uuid="e4350c9dbdfedb4a92d266b30daa3e0e35f3f322"
# convert to category dtype
df['Sex'] = df['Sex'].astype('category')
# convert to category codes
df['Sex'] = df['Sex'].cat.codes
# + _cell_guid="ddf8a643-1c1d-4036-818c-0d230a05310d" _uuid="911939a11430319dbda5a2c533c6b8c077a46766"
# subset all categorical variables which need to be encoded
categorical = ['Embarked', 'Title']
for var in categorical:
df = pd.concat([df,
pd.get_dummies(df[var], prefix=var)], axis=1)
del df[var]
# + _uuid="ecfe8fb0453e91f3a8cf80c45ed8ad4a406e0e0c"
# drop the variables we won't be using
df.drop(['Cabin', 'Name', 'Ticket', 'PassengerId'], axis=1, inplace=True)
# + _uuid="416083bad67fefd62cfee27da0f60bdb57a9540b"
df.head()
# + [markdown] _cell_guid="bc50ae8f-a1d7-48b6-9771-e50b4e8e169b" _uuid="bbead52ea8dd6e9cba5da969375c7829a34b87d4"
# <a id="random-forest"></a>
# # 3. Random Forest
# Now, all that is left is to feed our data that has been cleaned, encoded and scaled to a random forest.
# <a id="train-test"></a>
# ## 3.1. Train/test split
# But first, we need to separate *data_df* back into *train* and *test* sets.
# + _uuid="c3fffdde68114df12c2a1871d8501cae18a2e2ce"
train = df[pd.notnull(df['Survived'])]
X_test = df[pd.isnull(df['Survived'])].drop(['Survived'], axis=1)
# + [markdown] _uuid="26ac216b3215e4b030177916fd5ea79f9ed63bfc"
# ### Validation set
# Since we can't use our test set to assess our model (it doesn't have any labels), we will create a separte 'validation set'. We will use this set to test how our model generalises to unseen data.
# + _uuid="94f5bbc7429ec311e8a6eced5d7ed651ef2cba8f"
X_train, X_val, y_train, y_val = train_test_split(
train.drop(['Survived'], axis=1),
train['Survived'],
test_size=0.2, random_state=42)
# + _uuid="4282fce18e666cd315d47c7aca63e4d03c224146"
for i in [X_train, X_val, X_test]:
print(i.shape)
# + [markdown] _cell_guid="edcc8edb-a50b-4ec3-9958-06c081fbcd68" _uuid="75f56351057f33dcee93e207c4b3c461fcb65fae"
# ### Create Random Forest model
# We will first make a random forest model, using all of the default parameters.
# > Note: set the `random_state` to 42 for reproducibility
# + _uuid="92ff8b5cdbc88dae948d8df9a2219615e674bb09"
rf = RandomForestClassifier(random_state=42)
# + [markdown] _cell_guid="34a52736-364d-4f77-87c1-b432a8cc6834" _uuid="1d96485d799ad3e3850dfc5656680dec1b62ae95"
# ### Train model
# Now, let's train the model on our training set.
# + _cell_guid="af48be46-7e2a-4816-95e7-b54489e19c1c" _uuid="aa8ceed05ab371e22807e332024f822695916912"
rf.fit(X_train, y_train)
# + [markdown] _uuid="71217231ff4f11b9eea2ae0f46470b672d820c7b"
# ### Test model
# + _uuid="0f868bd1d23a3ed93d268f8517b1ef09fc285fa5"
accuracy_score(y_val, rf.predict(X_val))
# + [markdown] _uuid="602d97ad434eb9e87faa76cfaf49de88c08df7e5"
# <a id="cv"></a>
# ## 3.2. Cross-validation
# Keeping a separate validation set means that we have less data on which to train our model. Cross-validation allows us to train our model on _all_ of the data, while still assessing its performance on unseen data.
#
# K-folds cross validation is the process of creating *k* different train/validate splits in the data and training the model *k* times.
#
# 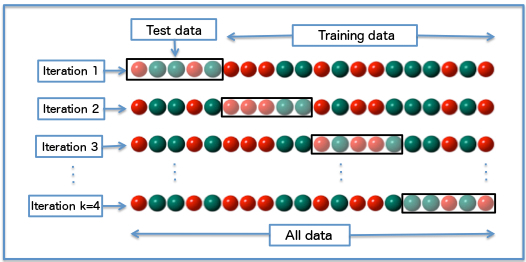
#
# In the image above, k=4. This means that the model will be trained 4 times, each time using 1/4 of the data for validation. In this way, each of the four 'folds' takes one turn sitting out from training and is used as the validation set.
#
# Let's combine our train and validation sets back into one training set, and then use cross-validation to assess our model:
# + _uuid="ca04b98bc96895fdbef13f2ce225e26fb0f374f3"
X_train = pd.concat([X_train, X_val])
y_train = pd.concat([y_train, y_val])
# + _uuid="ee3cc7c3270c30eb4e80fabe2623e504c3bc26c7"
X_train.shape
# + [markdown] _uuid="f0ba4b6d6382ea13815008d87736f2e1880d1d0d"
# Now we have all of training data again. Let's fit a model to it, and assess its accuracy using 5-fold cross-validation:
# + _uuid="f99d7b624572cf6826493d4c387304ca2a374e5d"
rf = RandomForestClassifier(n_estimators=10, random_state=42)
cross_val_score(rf, X_train, y_train, cv=5)
# + _uuid="b24804581b6e9c16d8dd4c212fa57f9e171a981e"
cross_val_score(rf, X_train, y_train, cv=5).mean()
# + [markdown] _uuid="62df755aaa3130891edc76c01e53e6fd911bb55f"
# Here, our CV score is slightly lower than our previous single validation score. Taking a look at the scores for each of the folds, the score does seem to vary slightly.
#
# Cross-validation has the added advantage of being a more robust measure of model accuracy than single validation.
# > Note: the method we used initially is actually just 1-fold cross-validation
# + [markdown] _uuid="06d18e0e5e48f5cf1781f6f91a99ad21c6af11f4"
# <a id="grid-search"></a>
# ## 3.3. Hyperparameter tuning
# Our first model didn't do too badly! It scored over 80% on the CV score. However, we didn't put any thought into our choice of hyperparameters, we simply went with the defaults.
#
# Take a look at the various parameters by using the `help()` function:
# + _kg_hide-output=false _uuid="8e6c2fb1f54d4868c37f2d25b80f3c6fb2d00d50"
# help(RandomForestClassifier)
# + [markdown] _uuid="0b220d582e6c2210a166807855d273efb8fb73ea"
# It is hard to know the best values for each of these hyperparameters without first _trying_ them out. If we wanted to know the best value for the `n_estimators` parameter, we could fit a few models, each with a different value, and see which one tests the best.
#
# **Grid search** allows us to do this for multiple parameters simultaneously. We will select a few different parameters that we want to tune, and for each one we will provide a few different values to try out. Then grid search will fit models to every possible combination of these parameter values and use **cross-validation** to assess the performance in each case.
#
# Furthermore, since we are using CV, we don't need to keep a separate validation set.
# + [markdown] _uuid="eabfa8a3dbb1dd061e642e8263bcf98feae6e65e"
# ### 3.2.1. Number of estimators and max depth
# We will start by tuning the `n_estimators` (number of trees in the forest) and the `max_depth` (how deep each tree grows) parameters.
#
# The first step that we need to do is to define the grid of parameters over which to search:
# + _uuid="aee8c8a76611d4e075179b2191b4da5324c472bd"
# create the grid
n_estimators = [10, 100, 1000, 2000]
max_depth = [None, 5, 10, 20]
param_grid = dict(n_estimators=n_estimators, max_depth=max_depth)
# + [markdown] _uuid="2767ce9f3dae7a78fabf83fb0e57e07f50e7c1b4"
# We have set out a total of $4 \times 4 = 16$ models over which to search. Grid search uses cross-validation on each of the models, so if we use 3-folds cross-validation, that will leave us with 48 different fits to try out. (You can see how the number of fits can grow pretty quickly as we increase the number of parameters!)
#
# The good news is that SkLearn's grid search allows us to run the job in parallel. Including the `n_jobs=-1` argument below let's grid search run on all of the available cores on the host machine.
# + _uuid="aee8c8a76611d4e075179b2191b4da5324c472bd"
# create the default model
rf = RandomForestClassifier(random_state=42)
# search the grid
grid = GridSearchCV(estimator=rf,
param_grid=param_grid,
cv=3,
verbose=2,
n_jobs=-1)
grid_result = grid.fit(X_train, y_train)
# + [markdown] _uuid="3a27f7fae51580b6c21e63304b36f15550fdaa7d"
# Now let's take a look at the results of the grid search.
#
# We can get the best performing model directly from `grid_result`:
# + _uuid="fd49612793fb6dc3b7d1349609730d8d7252c24c"
grid_result.best_estimator_
# + [markdown] _uuid="b838096425ad229c47184e1bdf38cfaf97c77041"
# Or just the best parameters:
# + _uuid="d53d9749a0bf5512b07d74b52f3af42ef4664efe"
grid_result.best_params_
# + [markdown] _uuid="f7b709a2647359ad2489cf91bfe9774c68c069fc"
# Or the best score:
# + _uuid="414c18d0f81e23676d44fd003711af3714960151"
grid_result.best_score_
# + [markdown] _uuid="660e0f9fa4b44a90d957f9961ebd5f5370e3309b"
# But let's take a look at all of the models so we can make a more informed decision
# + _uuid="aee8c8a76611d4e075179b2191b4da5324c472bd"
# summarize results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
means = grid_result.cv_results_['mean_test_score']
stds = grid_result.cv_results_['std_test_score']
params = grid_result.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# + [markdown] _uuid="186635a5b21616833ab080a6d311d20345845acc"
# ### 3.2.2. Leaf size
# The `min_samples_leaf` argument controls the size of the leaves in the trees.
#
# We will set out the grid in a similar manner as before, only this time we will use the `max_depth` and `n_estimators` parameters that we found above.
# + _uuid="04e78698779d2862174346da739ff9bc262f13fa"
# create the grid
leaf_samples = [1, 2, 3, 4, 5, 6]
param_grid = dict(min_samples_leaf=leaf_samples)
# create the model with new max_depth and n_estimators
rf = grid_result.best_estimator_
# search the grid
grid = GridSearchCV(estimator=rf,
param_grid=param_grid,
cv=3,
verbose=2,
n_jobs=-1)
grid_result = grid.fit(X_train, y_train)
# + _uuid="3a7ab234020ad2a55643950392c17c4202a65dc6"
# summarize results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
means = grid_result.cv_results_['mean_test_score']
stds = grid_result.cv_results_['std_test_score']
params = grid_result.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# + [markdown] _uuid="2f55ba2987ca48df93dcff646787394999421556"
# ### 3.2.3. To bag or not to bag
# Bootstrap aggregating (or bagging) is a special case of the random forest where we bootstrap (sample with replacement) from the n training obersvations to create a new training set of size n for each tree. Furthermore, each tree considers all variables when making each split.
#
# We can use grid search to determine if bootstrapping will be an appropriate method to use.
# + _uuid="730a03c80e66e543693c39eaa9b906185ee2689c"
# create the grid
max_features = [5, 8, 10, 12, None]
bootstrap = [True, False]
param_grid = dict(max_features=max_features, bootstrap=bootstrap)
# create the model with new leaf size
rf = grid_result.best_estimator_
# search the grid
grid = GridSearchCV(estimator=rf,
param_grid=param_grid,
cv=3,
verbose=2,
n_jobs=-1)
grid_result = grid.fit(X_train, y_train)
# + _uuid="12d05efb91691a1f7468834e84c3436bfce63071"
# summarize results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
means = grid_result.cv_results_['mean_test_score']
stds = grid_result.cv_results_['std_test_score']
params = grid_result.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# + [markdown] _cell_guid="5ccf334a-8a72-45e5-b565-cc516639e087" _uuid="ae291559b10db2f627a87f62a553d15da43ada31"
# <a id="submission"></a>
# ## 4. Make Predictions on Test Set
# Finally, we can attempt to predict which passengers in the test set survived.
# + _uuid="4398cc0ae4581a3722f2e10d3dac944baf8c8d47"
rf = grid_result.best_estimator_
# + _uuid="cd64287cf95c931d94e8c2263ce64e33e580c50f"
# test our CV score
cross_val_score(rf, X_train, y_train, cv=5).mean()
# + _uuid="ce685f9df90375c19422e45f82417a1eaa6a01d9"
test['Survived'] = rf.predict(X_test)
# + _uuid="c644df967bacf87fb867b194a626b8684dffc90f"
solution = test[['PassengerId', 'Survived']]
solution['Survived'] = solution['Survived'].apply(int)
# + _uuid="40f53edf0a58f57b9dc450070beba6efef051dda"
solution.head(10)
# + [markdown] _cell_guid="0f646038-2eb3-4a77-94f5-a3abc6c3be1f" _uuid="0a9b9f4189c03b253c9140c4a93c9f453508a6ba"
# ## Output Final Predictions
# + _cell_guid="04e4be98-5955-43ab-a355-be65580a1162" _uuid="7dc52ff626a2620c8607c25d85bd8952f049690b"
solution.to_csv("Random_Forest_Solution.csv", index=False)
# -
|
Titanic-random-forest-grid-search (1).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="YFNYzLMu4Diy"
# ### Importing required libraries
#
#
#
# + id="6178afbb"
import pandas as pd
import numpy as np
import tensorflow as tf
from tensorboard.plugins import projector
# + [markdown] id="S0UE7m6Y4W9_"
# ### Reading CSV files that contains lables and its vectors
# + id="8301cd82"
### Here we are reading body parts and its 200 dimension vectors for sake of example.
body_parts = pd.read_csv(r'body_part_vectors_sme.csv', header=None)
# + id="716446ce"
### Get labels from data frame
body_parts_list = list(body_parts[0].values)
# + id="f7db045d"
### get the length of labels or no. of items in dataframe or csv file.
body_parts_length = len(body_parts_list)
dimensions = 200
# + id="f113d017"
### Slice the vectors out from the data frame.
vector_df = body_parts[body_parts.columns[1:]]
# + id="5094a45e"
### Transform the vectors dataframe into numpy array
numpy_vector = np.zeros((body_parts_length,dimensions))
for i in range(body_parts_length):
numpy_vector[i] = vector_df.iloc[i,:]
# + id="f9043058"
### Search for location where we can store tsv file for generating metadata. For simplicity, we can make folder "projector" and inside that we can keep all files related to it.
### It will write labels into metadata file.
with open(r'C:\Users\Downloads\projector\prefix_metadata.tsv', 'w+') as file_metadata:
for body_part in body_parts_list:
file_metadata.write(body_part + '\n')
# + id="21ebd9bf"
# sess = tf.InteractiveSession()
### Get the tensors out of numpy array
with tf.device("/cpu:0"):
embedding = tf.Variable(numpy_vector, trainable=False, name = 'prefix_embedding')
# + id="7338f556"
# tf.global_variables_initializer().run()
# saver = tf.train.Saver()
### Generate the checkpoint file and save it in projector folder.
checkpoint = tf.train.Checkpoint(embedding=embedding)
checkpoint.save(r'C:\Users\Downloads\projector\embedding.ckpt')
# + id="647e5131"
### Call visualize embeddings by integrating metadata file and embeddings.
### And follow the instruction below to generate embeddings.
config = projector.ProjectorConfig()
embed= config.embeddings.add()
embedding.tensor_name = "prefix_embedding"
embed.metadata_path = r'C:\Users\Downloads\projector\prefix_metadata.tsv'
projector.visualize_embeddings(r'C:\Users\Downloads\projector', config)
# + [markdown] id="a1c0984f"
# #### Instructions
#
# Step 1 - Run all above cells to generate ckpt (embeddings) and metadata files
#
# Step 2 - Make sure in the projector folder, you have these files.
#
# Step 3 - Open CMD, type pip show tensorflow
#
# Step 4 - Go to site packages -> Tensorboard
#
# Step 5 - Run -> python main.py --logdir=r'C:\Users\Downloads\projector' in which your metadata and ckpt files is there.
# + [markdown] id="a217c913"
# ### Links
#
# https://gist.github.com/lampts/026a4d6400b1efac9a13a3296f16e655
#
# https://www.tensorflow.org/tensorboard/tensorboard_projector_plugin
|
Tensorboard_Projection_From_Vectors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="aXehiGc3Kr2I"
# ##### Copyright 2020 The TensorFlow Hub Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + cellView="form" id="-6LKjmi8Ktoh"
#@title Copyright 2020 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + [markdown] id="MfBg1C5NB3X0"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/hub/tutorials/spice"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/hub/blob/master/examples/colab/spice.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/hub/blob/master/examples/colab/spice.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/hub/examples/colab/spice.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# <td>
# <a href="https://tfhub.dev/google/spice/2"><img src="https://www.tensorflow.org/images/hub_logo_32px.png" />See TF Hub model</a>
# </td>
# </table>
# + [markdown] id="sPQKw4x4bL8w"
# # Pitch Detection with SPICE
#
# This colab will show you how to use the SPICE model downloaded from TensorFlow Hub.
# + id="rfKwZlPnPwD1"
# !sudo apt-get install -q -y timidity libsndfile1
# + id="dYrIdOS8SW3b"
# All the imports to deal with sound data
# !pip install pydub numba==0.48 librosa music21
# + id="p09o78LGYdnz"
import tensorflow as tf
import tensorflow_hub as hub
import numpy as np
import matplotlib.pyplot as plt
import librosa
from librosa import display as librosadisplay
import logging
import math
import statistics
import sys
from IPython.display import Audio, Javascript
from scipy.io import wavfile
from base64 import b64decode
import music21
from pydub import AudioSegment
logger = logging.getLogger()
logger.setLevel(logging.ERROR)
print("tensorflow: %s" % tf.__version__)
#print("librosa: %s" % librosa.__version__)
# + [markdown] id="wHxox8hXc3w1"
# # The audio input file
# Now the hardest part: Record your singing! :)
#
# We provide four methods to obtain an audio file:
#
# 1. Record audio directly in colab
# 2. Upload from your computer
# 3. Use a file saved on Google Drive
# 4. Download the file from the web
#
# Choose one of the four methods below.
# + cellView="form" id="HaCAHOqiVu5B"
#@title [Run this] Definition of the JS code to record audio straight from the browser
RECORD = """
const sleep = time => new Promise(resolve => setTimeout(resolve, time))
const b2text = blob => new Promise(resolve => {
const reader = new FileReader()
reader.onloadend = e => resolve(e.srcElement.result)
reader.readAsDataURL(blob)
})
var record = time => new Promise(async resolve => {
stream = await navigator.mediaDevices.getUserMedia({ audio: true })
recorder = new MediaRecorder(stream)
chunks = []
recorder.ondataavailable = e => chunks.push(e.data)
recorder.start()
await sleep(time)
recorder.onstop = async ()=>{
blob = new Blob(chunks)
text = await b2text(blob)
resolve(text)
}
recorder.stop()
})
"""
def record(sec=5):
try:
from google.colab import output
except ImportError:
print('No possible to import output from google.colab')
return ''
else:
print('Recording')
display(Javascript(RECORD))
s = output.eval_js('record(%d)' % (sec*1000))
fname = 'recorded_audio.wav'
print('Saving to', fname)
b = b64decode(s.split(',')[1])
with open(fname, 'wb') as f:
f.write(b)
return fname
# + cellView="both" id="sBpWWkTzfUYR"
#@title Select how to input your audio { run: "auto" }
INPUT_SOURCE = 'https://storage.googleapis.com/download.tensorflow.org/data/c-scale-metronome.wav' #@param ["https://storage.googleapis.com/download.tensorflow.org/data/c-scale-metronome.wav", "RECORD", "UPLOAD", "./drive/My Drive/YOUR_MUSIC_FILE.wav"] {allow-input: true}
print('You selected', INPUT_SOURCE)
if INPUT_SOURCE == 'RECORD':
uploaded_file_name = record(5)
elif INPUT_SOURCE == 'UPLOAD':
try:
from google.colab import files
except ImportError:
print("ImportError: files from google.colab seems to not be available")
else:
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
uploaded_file_name = next(iter(uploaded))
print('Uploaded file: ' + uploaded_file_name)
elif INPUT_SOURCE.startswith('./drive/'):
try:
from google.colab import drive
except ImportError:
print("ImportError: files from google.colab seems to not be available")
else:
drive.mount('/content/drive')
# don't forget to change the name of the file you
# will you here!
gdrive_audio_file = 'YOUR_MUSIC_FILE.wav'
uploaded_file_name = INPUT_SOURCE
elif INPUT_SOURCE.startswith('http'):
# !wget --no-check-certificate 'https://storage.googleapis.com/download.tensorflow.org/data/c-scale-metronome.wav' -O c-scale.wav
uploaded_file_name = 'c-scale.wav'
else:
print('Unrecognized input format!')
print('Please select "RECORD", "UPLOAD", or specify a file hosted on Google Drive or a file from the web to download file to download')
# + [markdown] id="4S2BvIoDf9nf"
# # Preparing the audio data
#
# Now we have the audio, let's convert it to the expected format and then listen to it!
#
# The SPICE model needs as input an audio file at a sampling rate of 16kHz and with only one channel (mono).
#
# To help you with this part, we created a function (`convert_audio_for_model`) to convert any wav file you have to the model's expected format:
# + id="bQ1362i-JoFI"
# Function that converts the user-created audio to the format that the model
# expects: bitrate 16kHz and only one channel (mono).
EXPECTED_SAMPLE_RATE = 16000
def convert_audio_for_model(user_file, output_file='converted_audio_file.wav'):
audio = AudioSegment.from_file(user_file)
audio = audio.set_frame_rate(EXPECTED_SAMPLE_RATE).set_channels(1)
audio.export(output_file, format="wav")
return output_file
# + id="oL9pftZ2nPm9"
# Converting to the expected format for the model
# in all the input 4 input method before, the uploaded file name is at
# the variable uploaded_file_name
converted_audio_file = convert_audio_for_model(uploaded_file_name)
# + id="TslkX2AOZN0p"
# Loading audio samples from the wav file:
sample_rate, audio_samples = wavfile.read(converted_audio_file, 'rb')
# Show some basic information about the audio.
duration = len(audio_samples)/sample_rate
print(f'Sample rate: {sample_rate} Hz')
print(f'Total duration: {duration:.2f}s')
print(f'Size of the input: {len(audio_samples)}')
# Let's listen to the wav file.
Audio(audio_samples, rate=sample_rate)
# + [markdown] id="iBicZu5AgcpR"
# First thing, let's take a look at the waveform of our singing.
# + id="aAa2M3CLZcWW"
# We can visualize the audio as a waveform.
_ = plt.plot(audio_samples)
# + [markdown] id="J1eI0b8qgn08"
# A more informative visualization is the [spectrogram](https://en.wikipedia.org/wiki/Spectrogram), which shows frequencies present over time.
#
# Here, we use a logarithmic frequency scale, to make the singing more clearly visible.
#
# + id="fGR4UZtpZvWI"
MAX_ABS_INT16 = 32768.0
def plot_stft(x, sample_rate, show_black_and_white=False):
x_stft = np.abs(librosa.stft(x, n_fft=2048))
fig, ax = plt.subplots()
fig.set_size_inches(20, 10)
x_stft_db = librosa.amplitude_to_db(x_stft, ref=np.max)
if(show_black_and_white):
librosadisplay.specshow(data=x_stft_db, y_axis='log',
sr=sample_rate, cmap='gray_r')
else:
librosadisplay.specshow(data=x_stft_db, y_axis='log', sr=sample_rate)
plt.colorbar(format='%+2.0f dB')
plot_stft(audio_samples / MAX_ABS_INT16 , sample_rate=EXPECTED_SAMPLE_RATE)
plt.show()
# + [markdown] id="MGCzo_cjjH-7"
# We need one last conversion here. The audio samples are in int16 format. They need to be normalized to floats between -1 and 1.
# + id="dv4H4O1Xb8T8"
audio_samples = audio_samples / float(MAX_ABS_INT16)
# + [markdown] id="yTdo_TwljVUV"
# # Executing the Model
# Now is the easy part, let's load the model with **TensorFlow Hub**, and feed the audio to it.
# SPICE will give us two outputs: pitch and uncertainty
#
#
#
# + [markdown] id="xUptYSTAbc3I"
# **TensorFlow Hub** is a library for the publication, discovery, and consumption of reusable parts of machine learning models. It makes easy to use machine learning to solve your challenges.
#
# To load the model you just need the Hub module and the URL pointing to the model:
# + id="ri0A0DSXY_Yd"
# Loading the SPICE model is easy:
model = hub.load("https://tfhub.dev/google/spice/2")
# + [markdown] id="kQV5H6J4suMT"
# **Note:** An interesting detail here is that all the model urls from Hub can be used for download and also to read the documentation, so if you point your browser to that link you can read documentation on how to use the model and learn more about how it was trained.
# + [markdown] id="GUVICjIps9hI"
# With the model loaded, data prepared, we need 3 lines to get the result:
# + id="tP55fXBYcBhb"
# We now feed the audio to the SPICE tf.hub model to obtain pitch and uncertainty outputs as tensors.
model_output = model.signatures["serving_default"](tf.constant(audio_samples, tf.float32))
pitch_outputs = model_output["pitch"]
uncertainty_outputs = model_output["uncertainty"]
# 'Uncertainty' basically means the inverse of confidence.
confidence_outputs = 1.0 - uncertainty_outputs
fig, ax = plt.subplots()
fig.set_size_inches(20, 10)
plt.plot(pitch_outputs, label='pitch')
plt.plot(confidence_outputs, label='confidence')
plt.legend(loc="lower right")
plt.show()
# + [markdown] id="blJwFWR4kMul"
# Let's make the results easier to understand by removing all pitch estimates with low confidence (confidence < 0.9) and plot the remaining ones.
#
#
# + id="d1MRmcm2cEkM"
confidence_outputs = list(confidence_outputs)
pitch_outputs = [ float(x) for x in pitch_outputs]
indices = range(len (pitch_outputs))
confident_pitch_outputs = [ (i,p)
for i, p, c in zip(indices, pitch_outputs, confidence_outputs) if c >= 0.9 ]
confident_pitch_outputs_x, confident_pitch_outputs_y = zip(*confident_pitch_outputs)
fig, ax = plt.subplots()
fig.set_size_inches(20, 10)
ax.set_ylim([0, 1])
plt.scatter(confident_pitch_outputs_x, confident_pitch_outputs_y, )
plt.scatter(confident_pitch_outputs_x, confident_pitch_outputs_y, c="r")
plt.show()
# + [markdown] id="vNBZ7ZblkxOm"
# The pitch values returned by SPICE are in the range from 0 to 1. Let's convert them to absolute pitch values in Hz.
# + id="n-CnpKzmcQi9"
def output2hz(pitch_output):
# Constants taken from https://tfhub.dev/google/spice/2
PT_OFFSET = 25.58
PT_SLOPE = 63.07
FMIN = 10.0;
BINS_PER_OCTAVE = 12.0;
cqt_bin = pitch_output * PT_SLOPE + PT_OFFSET;
return FMIN * 2.0 ** (1.0 * cqt_bin / BINS_PER_OCTAVE)
confident_pitch_values_hz = [ output2hz(p) for p in confident_pitch_outputs_y ]
# + [markdown] id="24yK0a6HjCSZ"
# Now, let's see how good the prediction is: We will overlay the predicted pitches over the original spectrogram. To make the pitch predictions more visible, we changed the spectrogram to black and white.
# + id="L1kaAcX9rrDo"
plot_stft(audio_samples / MAX_ABS_INT16 ,
sample_rate=EXPECTED_SAMPLE_RATE, show_black_and_white=True)
# Note: conveniently, since the plot is in log scale, the pitch outputs
# also get converted to the log scale automatically by matplotlib.
plt.scatter(confident_pitch_outputs_x, confident_pitch_values_hz, c="r")
plt.show()
# + [markdown] id="NskqpiHLxq6V"
# # Converting to musical notes
#
# Now that we have the pitch values, let's convert them to notes!
# This is part is challenging by itself. We have to take into account two things:
# 1. the rests (when there's no singing)
# 2. the size of each note (offsets)
# + [markdown] id="KDOlm9PLTTjt"
# ### 1: Adding zeros to the output to indicate when there's no singing
# + id="9uSQ3bJmTZmo"
pitch_outputs_and_rests = [
output2hz(p) if c >= 0.9 else 0
for i, p, c in zip(indices, pitch_outputs, confidence_outputs)
]
# + [markdown] id="9fM0UwlsTt4w"
# ### 2: Adding note offsets
#
# When a person sings freely, the melody may have an offset to the absolute pitch values that notes can represent.
# Hence, to convert predictions to notes, one needs to correct for this possible offset.
# This is what the following code computes.
# + id="fsJu-P5ksdFW"
A4 = 440
C0 = A4 * pow(2, -4.75)
note_names = ["C", "C#", "D", "D#", "E", "F", "F#", "G", "G#", "A", "A#", "B"]
def hz2offset(freq):
# This measures the quantization error for a single note.
if freq == 0: # Rests always have zero error.
return None
# Quantized note.
h = round(12 * math.log2(freq / C0))
return 12 * math.log2(freq / C0) - h
# The ideal offset is the mean quantization error for all the notes
# (excluding rests):
offsets = [hz2offset(p) for p in pitch_outputs_and_rests if p != 0]
print("offsets: ", offsets)
ideal_offset = statistics.mean(offsets)
print("ideal offset: ", ideal_offset)
# + [markdown] id="K17It_qT2DtE"
# We can now use some heuristics to try and estimate the most likely sequence of notes that were sung.
# The ideal offset computed above is one ingredient - but we also need to know the speed (how many predictions make, say, an eighth?), and the time offset to start quantizing. To keep it simple, we'll just try different speeds and time offsets and measure the quantization error, using in the end the values that minimize this error.
# + id="eMUTI4L52ZHA"
def quantize_predictions(group, ideal_offset):
# Group values are either 0, or a pitch in Hz.
non_zero_values = [v for v in group if v != 0]
zero_values_count = len(group) - len(non_zero_values)
# Create a rest if 80% is silent, otherwise create a note.
if zero_values_count > 0.8 * len(group):
# Interpret as a rest. Count each dropped note as an error, weighted a bit
# worse than a badly sung note (which would 'cost' 0.5).
return 0.51 * len(non_zero_values), "Rest"
else:
# Interpret as note, estimating as mean of non-rest predictions.
h = round(
statistics.mean([
12 * math.log2(freq / C0) - ideal_offset for freq in non_zero_values
]))
octave = h // 12
n = h % 12
note = note_names[n] + str(octave)
# Quantization error is the total difference from the quantized note.
error = sum([
abs(12 * math.log2(freq / C0) - ideal_offset - h)
for freq in non_zero_values
])
return error, note
def get_quantization_and_error(pitch_outputs_and_rests, predictions_per_eighth,
prediction_start_offset, ideal_offset):
# Apply the start offset - we can just add the offset as rests.
pitch_outputs_and_rests = [0] * prediction_start_offset + \
pitch_outputs_and_rests
# Collect the predictions for each note (or rest).
groups = [
pitch_outputs_and_rests[i:i + predictions_per_eighth]
for i in range(0, len(pitch_outputs_and_rests), predictions_per_eighth)
]
quantization_error = 0
notes_and_rests = []
for group in groups:
error, note_or_rest = quantize_predictions(group, ideal_offset)
quantization_error += error
notes_and_rests.append(note_or_rest)
return quantization_error, notes_and_rests
best_error = float("inf")
best_notes_and_rests = None
best_predictions_per_note = None
for predictions_per_note in range(20, 65, 1):
for prediction_start_offset in range(predictions_per_note):
error, notes_and_rests = get_quantization_and_error(
pitch_outputs_and_rests, predictions_per_note,
prediction_start_offset, ideal_offset)
if error < best_error:
best_error = error
best_notes_and_rests = notes_and_rests
best_predictions_per_note = predictions_per_note
# At this point, best_notes_and_rests contains the best quantization.
# Since we don't need to have rests at the beginning, let's remove these:
while best_notes_and_rests[0] == 'Rest':
best_notes_and_rests = best_notes_and_rests[1:]
# Also remove silence at the end.
while best_notes_and_rests[-1] == 'Rest':
best_notes_and_rests = best_notes_and_rests[:-1]
# + [markdown] id="vMZbWA3aVqee"
# Now let's write the quantized notes as sheet music score!
#
# To do it we will use two libraries: [music21](http://web.mit.edu/music21/) and [Open Sheet Music Display](https://github.com/opensheetmusicdisplay/opensheetmusicdisplay)
#
# **Note:** for simplicity, we assume here that all notes have the same duration (a half note).
# + id="yVrk_IOIzpQR"
# Creating the sheet music score.
sc = music21.stream.Score()
# Adjust the speed to match the actual singing.
bpm = 60 * 60 / best_predictions_per_note
print ('bpm: ', bpm)
a = music21.tempo.MetronomeMark(number=bpm)
sc.insert(0,a)
for snote in best_notes_and_rests:
d = 'half'
if snote == 'Rest':
sc.append(music21.note.Rest(type=d))
else:
sc.append(music21.note.Note(snote, type=d))
# + cellView="both" id="CEleCWHtG2s4"
#@title [Run this] Helper function to use Open Sheet Music Display (JS code) to show a music score
from IPython.core.display import display, HTML, Javascript
import json, random
def showScore(score):
xml = open(score.write('musicxml')).read()
showMusicXML(xml)
def showMusicXML(xml):
DIV_ID = "OSMD_div"
display(HTML('<div id="'+DIV_ID+'">loading OpenSheetMusicDisplay</div>'))
script = """
var div_id = {{DIV_ID}};
function loadOSMD() {
return new Promise(function(resolve, reject){
if (window.opensheetmusicdisplay) {
return resolve(window.opensheetmusicdisplay)
}
// OSMD script has a 'define' call which conflicts with requirejs
var _define = window.define // save the define object
window.define = undefined // now the loaded script will ignore requirejs
var s = document.createElement( 'script' );
s.setAttribute( 'src', "https://cdn.jsdelivr.net/npm/opensheetmusicdisplay@0.7.6/build/opensheetmusicdisplay.min.js" );
//s.setAttribute( 'src', "/custom/opensheetmusicdisplay.js" );
s.onload=function(){
window.define = _define
resolve(opensheetmusicdisplay);
};
document.body.appendChild( s ); // browser will try to load the new script tag
})
}
loadOSMD().then((OSMD)=>{
window.openSheetMusicDisplay = new OSMD.OpenSheetMusicDisplay(div_id, {
drawingParameters: "compacttight"
});
openSheetMusicDisplay
.load({{data}})
.then(
function() {
openSheetMusicDisplay.render();
}
);
})
""".replace('{{DIV_ID}}',DIV_ID).replace('{{data}}',json.dumps(xml))
display(Javascript(script))
return
# + id="WTu4phq4WeAI"
# rendering the music score
showScore(sc)
print(best_notes_and_rests)
# + [markdown] id="fGPXm6Z83U2g"
# Let's convert the music notes to a MIDI file and listen to it.
#
# To create this file, we can use the stream we created before.
# + id="klYoWjgmPaod"
# Saving the recognized musical notes as a MIDI file
converted_audio_file_as_midi = converted_audio_file[:-4] + '.mid'
fp = sc.write('midi', fp=converted_audio_file_as_midi)
# + id="tz7Mj3Qx1lpR"
wav_from_created_midi = converted_audio_file_as_midi.replace(' ', '_') + "_midioutput.wav"
print(wav_from_created_midi)
# + [markdown] id="ahss5EOiWDDp"
# To listen to it on colab, we need to convert it back to wav. An easy way of doing that is using Timidity.
# + id="XmeJ-UITV2nq"
# !timidity $converted_audio_file_as_midi -Ow -o $wav_from_created_midi
# + [markdown] id="bnvwmyNj7kCC"
# And finally, listen the audio, created from notes, created via MIDI from the predicted pitches, inferred by the model!
#
# + id="qNLBB0zJV6vN"
Audio(wav_from_created_midi)
|
examples/colab/spice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/liviasantos08/Analise-de-Dados-com-Python/blob/main/Aula1_Pandas.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="kLMFILjBIE0R" outputId="aaf9cf8d-1b64-4e62-daf4-70324b11e11f"
print("Hello world")
# + id="tIgwFSkpId9w"
#importando a biblioteca panda
import pandas as pd
# + id="9nhTWzf_Skx0"
#armazenar os conjuntos de dados
df = pd.read_csv("/content/drive/MyDrive/datasets/Gapminder.csv",error_bad_lines=False,sep = ";")
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="DCyQO2AxTPRt" outputId="90473596-93ba-4aba-e046-3025352766eb"
#Visualiazando as 5 primeira linhas
df.head()
# + id="OfucO4UIUwn2"
#Renomeando as colunas
df = df.rename(columns={"country": "País", "continent":"Continente", "year":"Ano", "lifeExp":"Expectativa de vida", "pop":"Pop total", "gdpPercap": "Pib"})
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="nHnqEVjaqCUf" outputId="a8d7e6eb-d188-4976-9a17-8a1ba18d405e"
#as primeiras linhas do conjunto de dados
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="zfAb__8SqPf2" outputId="9e4ab2f8-9544-4811-8345-0ecb33cbd6b8"
#numero de linha e colunas
df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="8aWyOkqOqSDR" outputId="66494568-67fc-409c-9227-10fbaa787cdf"
#retorna as colunas
df.columns
# + colab={"base_uri": "https://localhost:8080/"} id="vydX0VpyqivX" outputId="8eec6a52-6aed-4725-9cf3-aa77f9c035a0"
#retorna os tipos da coluna
df.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="OlqBp5Irq7Ef" outputId="98819af2-1889-4a39-eff9-940a3aea6eba"
#retorna as últimas linhas
df.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="IuX1p9zIrGNM" outputId="1f2e3639-ed43-4d58-a0f0-694c3ea4609f"
#Retorna informação estatística
df.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="es8uOSP1rZfx" outputId="2bc9659d-33a8-4e0d-9d29-5efba6e42d68"
#fazer um filtro
df["Continente"].unique()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="lkxAnpvrsHGb" outputId="9ab3d880-956a-4b33-839f-148a7fcfd370"
#localização de dados
oceania= df.loc[df["Continente"] == "Oceania"]
oceania.head()
# + colab={"base_uri": "https://localhost:8080/"} id="3kxbHq1atAhL" outputId="88cfd84a-34ca-401c-c0ba-1872f70348d6"
#Agrupamento de dados: quantos países cada continente tem
df.groupby("Continente")["País"].nunique()
# + colab={"base_uri": "https://localhost:8080/"} id="Gkv-GOCGtZNe" outputId="38a96dcf-7129-48cd-92ac-488cd5794b74"
#Qual é a expectiva media de vida para cada ano
df.groupby("Ano")["Expectativa de vida"].mean()
# + colab={"base_uri": "https://localhost:8080/"} id="f9KC2HDzuX9T" outputId="c80cf736-587b-4a96-8924-72e74981cd38"
#A média do Pib
df["Pib"].mean()
# + colab={"base_uri": "https://localhost:8080/"} id="0UXxkQBDujN7" outputId="901d765c-ec49-43db-e2c2-e5d49de4967b"
#A soma do Pib
df["Pib"].sum()
# + colab={"base_uri": "https://localhost:8080/"} id="67EWYwkimc5F" outputId="9ce3c6cc-1fb1-4e32-8fc5-a905b7a1746e"
df.isnull().sum()
|
Aula1_Pandas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Bayesian parameter estimation using MCMC
#
# Authors: <NAME> and <NAME> <br>
# Last modified on 12/14/2018 by <NAME>
#
# In Bayesian parameter estimation, one is looking to estimate the posterior pdf of the parameter vector characterizing the model, based on available data. The posterior pdf is given by Bayes' theorem:
#
# $$ p(\theta \vert data) = \frac{p(data \vert \theta) p(\theta)}{p(data)} $$
#
# The posterior pdf of nonlinear non-Gaussian models is often intractable, approximations must be used. In particular, Markov Chain Monte Carlo methods are a very popular way to sample from the posterior pdf. This notebook illustrates how to perfrom Bayesian parameter estimation via MCMC in UQpy, using the BayesParameterEstimation class. At the end of this notebook, we also show how to quickly check your results using some simple Diagnostics tools implemented in UQpy.
import numpy as np
import matplotlib.pyplot as plt
from UQpy.Inference import *
from UQpy.RunModel import RunModel # required to run the quadratic model
from sklearn.neighbors import KernelDensity # for the plots
# Function to plot posterior pdf from samples
def pdf_from_kde(domain, samples1d):
bandwidth = 1.06 * np.std(samples1d) * samples1d.size ** (-1/5)
kde = KernelDensity(bandwidth=bandwidth).fit(samples1d.reshape((-1,1)))
log_dens = kde.score_samples(domain)
return np.exp(log_dens)
# ## Probability model
#
# In the following we learn the mean and covariance of a univariate gaussian distribution from data.
#
# First, for the sake of this example, we generate fake data from a gaussian distribution with mean 10 and standard deviation 1.
# +
np.random.seed(100)
mu, sigma = 10, 1 # true mean and standard deviation
data_1 = np.random.normal(mu, sigma, 100).reshape((-1, 1))
np.random.seed()
# plot the data and true distribution
count, bins, ignored = plt.hist(data_1, 30, density=True)
plt.plot(bins, 1/(sigma * np.sqrt(2 * np.pi)) * np.exp( - (bins - mu)**2 / (2 * sigma**2) ),
linewidth=2, color='r')
plt.title('data as histogram and true distribution to be estimated')
plt.show()
# -
# In a Bayesian setting, the definition of a prior pdf is a key point. The prior for the parameters must be defined in the model. Note that if no prior is given, an inproper, uninformative, prior is chosen, $p(\theta)=1$ for all $\theta$.
from UQpy.Distributions import JointInd, Uniform, Lognormal
p0 = Uniform(loc=0., scale=15)
p1 = Lognormal(s=1., loc=0., scale=1.)
prior = JointInd(marginals=[p0, p1])
# create an instance of class Model
candidate_model = InferenceModel(dist_object=Normal(loc=None, scale=None), nparams=2, prior=prior)
# Learn the unknown parameters using MCMC
from UQpy.SampleMethods import MH
bayes_estimator = BayesParameterEstimation(data=data_1, inference_model=candidate_model, sampling_class=MH,
nsamples=500, jump=10, nburn=10, seed=np.array([1.0, 0.2]))
# +
# print results
s = bayes_estimator.sampler.samples
plt.scatter(s[:,0],s[:,1])
plt.scatter(10, 1, marker = '+', label='true parameter')
plt.title('MCMC samples')
plt.legend()
plt.show()
fig, ax = plt.subplots(1,2, figsize=(10, 4))
domain = np.linspace(0, 15, 200)[:, np.newaxis]
pdf = pdf_from_kde(domain, s[:, 0])
ax[0].plot(domain, p0.pdf(domain), label='prior')
ax[0].plot(domain, pdf, label='posterior')
ax[0].set_title('posterior pdf of theta=mu')
ax[0].legend()
domain = np.linspace(0, 2, 200)[:, np.newaxis]
pdf = pdf_from_kde(domain, s[:, 1])
ax[1].plot(domain, p1.pdf(domain), label='prior')
ax[1].plot(domain, pdf, label='posterior')
ax[1].set_title('posterior pdf of theta=sigma')
ax[1].legend()
plt.show()
# -
# ## Regression model
#
# Here a model is defined that is of the form
#
# $$y=f(\theta) + \epsilon$$
#
# where f consists in running RunModel. In particular, here $f(\theta)=\theta_{0} x + \theta_{1} x^{2}$ is a regression model.
#
# First we generate synthetic data, and add some noise to it.
# +
# Generate data
from UQpy.Distributions import Normal
from UQpy.RunModel import RunModel
param_true = np.array([1.0, 2.0]).reshape((1, -1))
print('Shape of true parameter vector: {}'.format(param_true.shape))
h_func = RunModel(model_script='pfn_models.py', model_object_name='model_quadratic', vec=False,
var_names=['theta_0', 'theta_1'])
h_func.run(samples=param_true)
data_clean = np.array(h_func.qoi_list[0])
# -
# Add noise, use a RandomState for reproducible results
error_covariance = 1.
noise = Normal(loc=0., scale=np.sqrt(error_covariance)).rvs(nsamples=50, random_state=123).reshape((50, ))
data_3 = data_clean + noise
print('Shape of data: {}'.format(data_3.shape))
print(data_3[:4])
# +
from UQpy.Distributions import Normal, JointInd
p0 = Normal()
p1 = Normal()
prior = JointInd(marginals=[p0, p1])
inference_model = InferenceModel(nparams=2, runmodel_object=h_func, error_covariance=error_covariance,
prior=prior)
# +
from UQpy.SampleMethods import MH
proposal = JointInd([Normal(scale=0.1), Normal(scale=0.05)])
bayes_estimator = BayesParameterEstimation(
data=data_3, inference_model=inference_model, sampling_class=MH, nsamples=500, jump=10, nburn=0,
proposal=proposal, seed=[0.5, 2.5], verbose=True, random_state=456)
# +
s = bayes_estimator.sampler.samples
plt.scatter(s[:,0], s[:,1])
plt.scatter(1.0, 2.0, label='true value')
plt.title('MCMC samples')
plt.show()
fig, ax = plt.subplots(1,2, figsize=(10,4))
domain = np.linspace(-4, 4, 200)[:, np.newaxis]
pdf_ = pdf_from_kde(domain, s[:, 0])
ax[0].plot(domain, pdf_, label='posterior')
ax[0].plot(domain, p0.pdf(domain), label='prior')
ax[0].set_title('posterior pdf of theta_{1}')
domain = np.linspace(-4, 4, 200)[:, np.newaxis]
pdf_ = pdf_from_kde(domain, s[:, 1])
ax[1].plot(domain, pdf_, label='posterior')
ax[1].plot(domain, p1.pdf(domain), label='prior')
ax[1].set_title('posterior pdf of theta_{2}')
plt.show()
print(bayes_estimator.sampler.samples[:4])
# -
|
example/Inference/BayesParameterEstimation/Bayesian_parameter_estimation_MCMC.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Fine Tuning BERT For Sentiment Classification
#
# This notebook uses BERT to perform sentiment classification. A great tutorial for fine tuning BERT can be found here: https://skimai.com/fine-tuning-bert-for-sentiment-analysis/
#
# We may not always have access to GPU, in which case, training to defer back to using CPU.
#
# As a rule of thumb, reviews that are 3 stars and above are **positive**, and vice versa.
# +
# #!pip install transformers
# +
# #!pip install torch
# -
import gzip
import json
import matplotlib.pyplot as plt
import numpy as np
import re
import random
import pandas as pd
import seaborn as sns
import torch
from collections import Counter, defaultdict
from sklearn.dummy import DummyClassifier
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import BernoulliNB, ComplementNB, MultinomialNB
from sklearn.metrics import f1_score, classification_report, accuracy_score, confusion_matrix
from sklearn.model_selection import train_test_split, GridSearchCV
from transformers import BertTokenizer
from nltk.tokenize import sent_tokenize, word_tokenize
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
from tqdm import tqdm
torch.cuda.empty_cache()
# +
if torch.cuda.is_available():
device = torch.device("cuda")
print(f'There are {torch.cuda.device_count()} GPU(s) available.')
print('Device name:', torch.cuda.get_device_name(0))
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
# -
RANDOM_SEED = 33
reviews_sg = pd.read_pickle("assets/au_reviews.pkl")
reviews_sg.head()
reviews_sg['label'] = np.where(reviews_sg['rating'] >= 3, 0, 1)
reviews_sg.head()
# ## 1. Data Processing
reviews_sg.isnull().sum()
reviews_sg = reviews_sg.dropna()
df_proc = reviews_sg.copy()
df_proc.drop(columns=['date', 'rating', 'app'], inplace=True)
df_proc.head()
X = df_proc['review']
y = df_proc['label']
# We will split the dataset into `train`, and `dev` (at this point I don't see the point of using a test set, since I'm not doing cross validation). But ideally, I want the dev set to be the same across all different training sizes.
#
# We'll also try to stratify our dataset when we split.
X_train, X_dev, y_train, y_dev = train_test_split(X, y, test_size=0.2, random_state=RANDOM_SEED, stratify=y)
#X_test, X_dev, y_test, y_dev = train_test_split(X_test, y_test, test_size=0.5, random_state=RANDOM_SEED)
len(X_train)
len(X_dev)
df_train = pd.concat([X_train, y_train], axis=1)
df_train.head()
# Split again to get a subset of the train dataset. We only want 100k to train.
X_disc, X_train, y_disc, y_train = train_test_split(X_train, y_train,
test_size=100000, random_state=RANDOM_SEED, stratify=y_train)
len(X_train)
X_train.iloc[0]
X_dev.iloc[0]
# ## 2. BERT Tokenization
#
# It is a requirement to use BERT tokenization. Prior to that, the example notebook also provided a function for some basic text processing, albeit much less (according to the notebook, this is because BERT was trained with the entire sentences?).
#
#
def text_preprocessing(text):
"""
- Correct errors (eg. '&' to '&')
@param text (str): a string to be processed.
@return text (Str): the processed string.
"""
# Replace '&' with '&'
text = re.sub(r'&', '&', text)
# Remove trailing whitespace
text = re.sub(r'\s+', ' ', text).strip()
return text
# +
# Load the BERT tokenizer
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased', do_lower_case=True)
# Create a function to tokenize a set of texts
def preprocessing_for_bert(data):
"""Perform required preprocessing steps for pretrained BERT.
@param data (np.array): Array of texts to be processed.
@return input_ids (torch.Tensor): Tensor of token ids to be fed to a model.
@return attention_masks (torch.Tensor): Tensor of indices specifying which
tokens should be attended to by the model.
"""
# Create empty lists to store outputs
input_ids = []
attention_masks = []
# For every sentence...
for sent in data:
# `encode_plus` will:
# (1) Tokenize the sentence
# (2) Add the `[CLS]` and `[SEP]` token to the start and end
# (3) Truncate/Pad sentence to max length
# (4) Map tokens to their IDs
# (5) Create attention mask
# (6) Return a dictionary of outputs
encoded_sent = tokenizer.encode_plus(
text=text_preprocessing(sent), # Preprocess sentence
add_special_tokens=True, # Add `[CLS]` and `[SEP]`
max_length=MAX_LEN, # Max length to truncate/pad
truncation=True,
padding='max_length', # Pad sentence to max length
#return_tensors='pt', # Return PyTorch tensor
return_attention_mask=True # Return attention mask
)
# Add the outputs to the lists
input_ids.append(encoded_sent.get('input_ids'))
attention_masks.append(encoded_sent.get('attention_mask'))
# Convert lists to tensors
input_ids = torch.tensor(input_ids)
attention_masks = torch.tensor(attention_masks)
return input_ids, attention_masks
# -
# And apparently, we need to find the max length of our sentence to later feed into the function above.
# +
# Encode our concatenated data
encoded_reviews = [tokenizer.encode(review, add_special_tokens=True) for review in df_proc.review]
# Find the maximum length
max_len = max([len(sent) for sent in encoded_reviews])
print('Max length: ', max_len)
# +
# Specify `MAX_LEN`
MAX_LEN = 512
# Print sentence 0 and its encoded token ids
token_ids = list(preprocessing_for_bert([X.values[0]])[0].squeeze().numpy())
print('Original: ', X.values[0])
print('Token IDs: ', token_ids)
# Run function `preprocessing_for_bert` on the train set and the validation set
# Note that the function takes in a numpy array
print('Tokenizing data...')
train_inputs, train_masks = preprocessing_for_bert(X_train.values)
dev_inputs, dev_masks = preprocessing_for_bert(X_dev.values)
# -
# Now create a torch DataLoader class, which is meant to save memory during training and boost training speed.
# +
# Convert other data types to torch.Tensor
train_labels = torch.tensor(y_train.values)
dev_labels = torch.tensor(y_dev.values)
# For fine-tuning BERT, the authors recommend a batch size of 16 or 32.
batch_size = 16
# Create the DataLoader for our training set
train_data = TensorDataset(train_inputs, train_masks, train_labels)
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=batch_size)
# Create the DataLoader for our validation set
dev_data = TensorDataset(dev_inputs, dev_masks, dev_labels)
dev_sampler = SequentialSampler(dev_data)
dev_dataloader = DataLoader(dev_data, sampler=dev_sampler, batch_size=batch_size)
# -
# ## 3. Model Training
#
# BERT-base consists of 12 transformer layers, each transformer layer takes in a list of token embeddings, and produces the same number of embeddings with the same hidden size (or dimensions) on the output. The output of the final transformer layer of the [CLS] token is used as the features of the sequence to feed a classifier.
#
# The transformers library has the BertForSequenceClassification class which is designed for classification tasks. However, we will create a new class so we can specify our own choice of classifiers.
#
# Below we will create a BertClassifier class with a BERT model to extract the last hidden layer of the [CLS] token and a single-hidden-layer feed-forward neural network as our classifier.
# ### 3.1 Create Classifier
# +
# %%time
import torch
import torch.nn as nn
from transformers import BertModel
# Create the BertClassfier class
class BertClassifier(nn.Module):
"""Bert Model for Classification Tasks.
"""
def __init__(self, freeze_bert=False):
"""
@param bert: a BertModel object
@param classifier: a torch.nn.Module classifier
@param freeze_bert (bool): Set `False` to fine-tune the BERT model
"""
super(BertClassifier, self).__init__()
# Specify hidden size of BERT, hidden size of our classifier, and number of labels
D_in, H, D_out = 768, 50, 2
# Instantiate BERT model
self.bert = BertModel.from_pretrained('bert-base-uncased')
# Instantiate an one-layer feed-forward classifier
self.classifier = nn.Sequential(
nn.Linear(D_in, H),
nn.ReLU(),
#nn.Dropout(0.5),
nn.Linear(H, D_out)
)
# Freeze the BERT model
if freeze_bert:
for param in self.bert.parameters():
param.requires_grad = False
def forward(self, input_ids, attention_mask):
"""
Feed input to BERT and the classifier to compute logits.
@param input_ids (torch.Tensor): an input tensor with shape (batch_size,
max_length)
@param attention_mask (torch.Tensor): a tensor that hold attention mask
information with shape (batch_size, max_length)
@return logits (torch.Tensor): an output tensor with shape (batch_size,
num_labels)
"""
# Feed input to BERT
outputs = self.bert(input_ids=input_ids,
attention_mask=attention_mask)
# Extract the last hidden state of the token `[CLS]` for classification task
last_hidden_state_cls = outputs[0][:, 0, :]
# Feed input to classifier to compute logits
logits = self.classifier(last_hidden_state_cls)
return logits
# -
# ### 3.2 Create Optimizer
# To fine-tune our Bert Classifier, we need to create an optimizer. The authors recommend following hyper-parameters:
#
# - Batch size: 16 or 32
# - Learning rate (Adam): 5e-5, 3e-5 or 2e-5
# - Number of epochs: 2, 3, 4
#
# Huggingface provided the `run_glue.py` script, an examples of implementing the transformers library. In the script, the AdamW optimizer is used.
# +
from transformers import AdamW, get_linear_schedule_with_warmup
def initialize_model(epochs=4):
"""Initialize the Bert Classifier, the optimizer and the learning rate scheduler.
"""
# Instantiate Bert Classifier
bert_classifier = BertClassifier(freeze_bert=False)
# Tell PyTorch to run the model on GPU
bert_classifier.to(device)
# Create the optimizer
optimizer = AdamW(bert_classifier.parameters(),
lr=5e-5, # Default learning rate
eps=1e-8 # Default epsilon value
)
# Total number of training steps
total_steps = len(train_dataloader) * epochs
# Set up the learning rate scheduler
scheduler = get_linear_schedule_with_warmup(optimizer,
num_warmup_steps=0, # Default value
num_training_steps=total_steps)
return bert_classifier, optimizer, scheduler
# -
# ### 3.3 Create a Training Loop
# We will train our Bert Classifier for 4 epochs. In each epoch, we will train our model and evaluate its performance on the validation set. In more details, we will:
#
# Training:
# - Unpack our data from the dataloader and load the data onto the GPU
# - Zero out gradients calculated in the previous pass
# - Perform a forward pass to compute logits and loss
# - Perform a backward pass to compute gradients (loss.backward())
# - Clip the norm of the gradients to 1.0 to prevent "exploding gradients"
# - Update the model's parameters (optimizer.step())
# - Update the learning rate (scheduler.step())
#
# Evaluation:
# - Unpack our data and load onto the GPU
# - Forward pass
# - Compute loss and accuracy rate over the validation set
# The script below is commented with the details of our training and evaluation loop.
# +
import random
import time
# Specify loss function
loss_fn = nn.CrossEntropyLoss()
def set_seed(seed_value=33):
"""Set seed for reproducibility.
"""
random.seed(seed_value)
np.random.seed(seed_value)
torch.manual_seed(seed_value)
torch.cuda.manual_seed_all(seed_value)
def train(model, train_dataloader, val_dataloader=None, epochs=4, evaluation=False):
"""Train the BertClassifier model.
"""
# Start training loop
print("Start training...\n")
for epoch_i in range(epochs):
# =======================================
# Training
# =======================================
# Print the header of the result table
print(f"{'Epoch':^7} | {'Batch':^7} | {'Train Loss':^12} | {'Val Loss':^10} | {'Val Acc':^9} | {'Elapsed':^9}")
print("-"*70)
# Measure the elapsed time of each epoch
t0_epoch, t0_batch = time.time(), time.time()
# Reset tracking variables at the beginning of each epoch
total_loss, batch_loss, batch_counts = 0, 0, 0
# Put the model into the training mode
model.train()
# For each batch of training data...
for step, batch in enumerate(train_dataloader):
batch_counts +=1
# Load batch to GPU
b_input_ids, b_attn_mask, b_labels = tuple(t.to(device) for t in batch)
# Zero out any previously calculated gradients
model.zero_grad()
# Perform a forward pass. This will return logits.
logits = model(b_input_ids, b_attn_mask)
# Compute loss and accumulate the loss values
loss = loss_fn(logits, b_labels)
batch_loss += loss.item()
total_loss += loss.item()
# Perform a backward pass to calculate gradients
loss.backward()
# Clip the norm of the gradients to 1.0 to prevent "exploding gradients"
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
# Update parameters and the learning rate
optimizer.step()
scheduler.step()
# Print the loss values and time elapsed for every 200 batches
if (step % 200 == 0 and step != 0) or (step == len(train_dataloader) - 1):
# Calculate time elapsed for 20 batches
time_elapsed = time.time() - t0_batch
# Print training results
print(f"{epoch_i + 1:^7} | {step:^7} | {batch_loss / batch_counts:^12.6f} | {'-':^10} | {'-':^9} | {time_elapsed:^9.2f}")
# Reset batch tracking variables
batch_loss, batch_counts = 0, 0
t0_batch = time.time()
# Calculate the average loss over the entire training data
avg_train_loss = total_loss / len(train_dataloader)
print("-"*70)
# =======================================
# Evaluation
# =======================================
if evaluation == True:
# After the completion of each training epoch, measure the model's performance
# on our validation set.
dev_loss, dev_accuracy = evaluate(model, dev_dataloader)
# Print performance over the entire training data
time_elapsed = time.time() - t0_epoch
print(f"{epoch_i + 1:^7} | {'-':^7} | {avg_train_loss:^12.6f} | {dev_loss:^10.6f} | {dev_accuracy:^9.2f} | {time_elapsed:^9.2f}")
print("-"*70)
print("\n")
print("Training complete!")
def evaluate(model, dev_dataloader):
"""After the completion of each training epoch, measure the model's performance
on our validation set.
"""
# Put the model into the evaluation mode. The dropout layers are disabled during
# the test time.
model.eval()
# Tracking variables
dev_accuracy = []
dev_loss = []
# For each batch in our validation set...
for batch in dev_dataloader:
# Load batch to GPU
b_input_ids, b_attn_mask, b_labels = tuple(t.to(device) for t in batch)
# Compute logits
with torch.no_grad():
logits = model(b_input_ids, b_attn_mask)
# Compute loss
loss = loss_fn(logits, b_labels)
dev_loss.append(loss.item())
# Get the predictions
preds = torch.argmax(logits, dim=1).flatten()
# Calculate the accuracy rate
accuracy = (preds == b_labels).cpu().numpy().mean() * 100
dev_accuracy.append(accuracy)
# Compute the average accuracy and loss over the development set.
dev_loss = np.mean(dev_loss)
dev_accuracy = np.mean(dev_accuracy)
return dev_loss, dev_accuracy
# -
set_seed(33) # Set seed for reproducibility
bert_classifier, optimizer, scheduler = initialize_model(epochs=2)
train(bert_classifier, train_dataloader, dev_dataloader, epochs=2, evaluation=True)
# ### 3.4 Evaluation on Development Set
#
# The prediction step is similar to the evaluation step that we did in the training loop, but simpler. We will perform a forward pass to compute logits and apply softmax to calculate probabilities.
# +
import torch.nn.functional as F
def bert_predict(model, test_dataloader):
"""Perform a forward pass on the trained BERT model to predict probabilities
on the test set.
"""
# Put the model into the evaluation mode. The dropout layers are disabled during
# the test time.
model.eval()
all_logits = []
# For each batch in our test set...
for batch in test_dataloader:
# Load batch to GPU
b_input_ids, b_attn_mask = tuple(t.to(device) for t in batch)[:2]
# Compute logits
with torch.no_grad():
logits = model(b_input_ids, b_attn_mask)
all_logits.append(logits)
# Concatenate logits from each batch
all_logits = torch.cat(all_logits, dim=0)
# Apply softmax to calculate probabilities
probs = F.softmax(all_logits, dim=1).cpu().numpy()
return probs
# +
from sklearn.metrics import accuracy_score, roc_curve, auc
def evaluate_roc(probs, y_true):
"""
- Print AUC and accuracy on the test set
- Plot ROC
@params probs (np.array): an array of predicted probabilities with shape (len(y_true), 2)
@params y_true (np.array): an array of the true values with shape (len(y_true),)
"""
preds = probs[:, 1]
fpr, tpr, threshold = roc_curve(y_true, preds)
roc_auc = auc(fpr, tpr)
print(f'AUC: {roc_auc:.4f}')
# Get accuracy over the test set
y_pred = np.where(preds >= 0.5, 1, 0)
accuracy = accuracy_score(y_true, y_pred)
print(f'Accuracy: {accuracy*100:.2f}%')
# Plot ROC AUC
plt.title('Receiver Operating Characteristic')
plt.plot(fpr, tpr, 'b', label = 'AUC = %0.2f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# +
# Compute predicted probabilities on the test set
probs = bert_predict(bert_classifier, dev_dataloader)
# Evaluate the Bert classifier
evaluate_roc(probs, y_dev)
# -
# Get predictions from the probabilities
threshold = 0.5
y_dev_pred = np.where(probs[:, 1] > threshold, 1, 0)
score = accuracy_score(y_dev, y_dev_pred)
print("accuracy: %0.3f" % score)
print("classification report:")
print(classification_report(y_dev, y_dev_pred))
print("confusion matrix:")
print(confusion_matrix(y_dev, y_dev_pred))
print("Training Complete")
print()
torch.save(bert_classifier, 'bert_au_small')
saved_model = torch.load('bert_au_small')
# Making sure that the saved model works...
# +
# Compute predicted probabilities on the test set
probs_load_test = bert_predict(saved_model, dev_dataloader)
# Evaluate the Bert classifier
evaluate_roc(probs_load_test, y_dev)
# -
threshold = 0.5
y_dev_pred_load_test = np.where(probs_load_test[:, 1] > threshold, 1, 0)
score = accuracy_score(y_dev, y_dev_pred_load_test)
print("accuracy: %0.3f" % score)
print("classification report:")
print(classification_report(y_dev, y_dev_pred_load_test))
print("confusion matrix:")
print(confusion_matrix(y_dev, y_dev_pred_load_test))
print("Training Complete")
print()
# FINALLY.
torch.save(bert_classifier, 'bert_au_small.pth')
|
playground/mel/sentiment_classification_au_DL_100k_sampling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Coding exercises
# Exercises 1-3 are thought exercises that don't require coding.
#
# ## Exercise 4: Generate the data by running this cell
# This will give you a list of numbers to work with in the remaining exercises.
salaries = [844000.0, 758000.0, 421000.0, 259000.0, 511000.0, 405000.0, 784000.0, 303000.0, 477000.0, 583000.0, 908000.0, 505000.0, 282000.0, 756000.0, 618000.0, 251000.0, 910000.0, 983000.0, 810000.0, 902000.0, 310000.0, 730000.0, 899000.0, 684000.0, 472000.0, 101000.0, 434000.0, 611000.0, 913000.0, 967000.0, 477000.0, 865000.0, 260000.0, 805000.0, 549000.0, 14000.0, 720000.0, 399000.0, 825000.0, 668000.0, 1000.0, 494000.0, 868000.0, 244000.0, 325000.0, 870000.0, 191000.0, 568000.0, 239000.0, 968000.0, 803000.0, 448000.0, 80000.0, 320000.0, 508000.0, 933000.0, 109000.0, 551000.0, 707000.0, 547000.0, 814000.0, 540000.0, 964000.0, 603000.0, 588000.0, 445000.0, 596000.0, 385000.0, 576000.0, 290000.0, 189000.0, 187000.0, 613000.0, 657000.0, 477000.0, 90000.0, 758000.0, 877000.0, 923000.0, 842000.0, 898000.0, 923000.0, 541000.0, 391000.0, 705000.0, 276000.0, 812000.0, 849000.0, 895000.0, 590000.0, 950000.0, 580000.0, 451000.0, 660000.0, 996000.0, 917000.0, 793000.0, 82000.0, 613000.0, 486000.0]
print(salaries)
print(len(salaries))
# ## Exercise 5: Calculating statistics and verifying
# ### mean
from statistics import mean
print(mean(salaries))
# + [markdown] tags=[]
# ### median
# -
from statistics import median
print(median(salaries))
# ### mode
from statistics import mode
print(mode(salaries))
# ### sample variance
# Remember to use Bessel's correction.
from statistics import variance
print(variance(salaries))
# ### sample standard deviation
# Remember to use Bessel's correction.
from statistics import stdev
print(stdev(salaries))
# ## Exercise 6: Calculating more statistics
# ### range
# ### coefficient of variation
# ### interquartile range
# ### quartile coefficent of dispersion
# ## Exercise 7: Scaling data
# ### min-max scaling
# ### standardizing
# ## Exercise 8: Calculating covariance and correlation
# ### covariance
# ### Pearson correlation coefficient ($\rho$)
|
ch_01/exercises_stat library.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 6
# This homework is all about useful external libraries that are most common to use in astronomy research. The two most important libraries apart from scipy, numpy, and matplotlib are **astropy** and **pandas**. We explore the basics of these super versatile libraries.
# # Astropy (50 Points)
# ## CRAZY UNIT CONVERSION!!! (20 Points)
# As you take more astronomy classes, you will face more and more unit conversion problems - they are annoying. That's why astropy.units is very helpful. Let's do some practices here.
#
# The documentations for astropy.units and astropy.constants will very helpful to you.
#
# astropy.units documentation: https://docs.astropy.org/en/stable/units/
#
# astropy.constants documentation: https://docs.astropy.org/en/stable/constants/
#
# NOTE: In this problem, you MUST use astropy.constants when doing calculations involving fundamental constants. Also, you cannot look up values such as solar mass, earth mass, etc. Use the two packages solely.
# ### Problem 1) Speed of light (5 Points)
#
# What is the speed of light ($c$) in $pc/yr$?
# +
### Write your code here
import astropy.constants as cons
import astropy.units as u
cons.c.to(u.pc / u.yr)
# -
# ### Problem 2) Newton's 2nd Law (5 Points)
#
# Recall that NII states
# $$F =ma\,\,.$$
# Say a force of $97650134N$ is exerted on an object having a mass of $0.0071$ earth mass. What is the acceleration of the object in $AU/days^2$?
# +
### Write your code here
a = (97650134 * u.N) / (0.0071*u.kg) #a = F/m
a.to(u.AU / (u.d)**2)
# -
# ### Problem 3) Newton's Universal Law of Gravitation (10 Points)
#
# Recall that the gravitational acceleration due to an object with mass $m$ at a distance $r$ is given by
# $$a_g = \frac{Gm}{r^2}\,\,.$$
# What is the gravitational acceleration due to a planet of $3.1415926$ Jupiter-mass at a distance of $1.523AU$? Give your answer in $pc/yr^2$.
# +
### Write your code here
a = cons.G*(3.1415926*cons.M_jup)/(1.523*u.AU)**2
a.to(u.pc / (u.yr)**2)
# -
# ## Visualising Coordinate Transformation (30 Points)
# We introduced coordinate transformation using astropy, but maybe that was too astract to you, so let's use this problem as a way for you to visualise this process. Each part will be worth **5 Points**
#
# There are several things you need to do:
# 1. Open up the FITS file named 'clusters.fits' (this part of the code is written for you already)
#
#
# 2. Read it as a table using astropy.table (you will have to import the packages you need and write your own code from hereafter)
#
#
# 3. Plot the positions of all the objects in the table, COLOUR-CODED by their types (there is a column named 'CLASS'), with RA on the x-axis and DEC on the y-axis. You should see a curved trend with a huge dip in the middle.
#
#
# 4. Carry out a coordinate transformation from the ICRS coordinates to the galactic coordinates - there is a column named "DISTANCE" which you will need.
#
#
# 5. Now plot the position of all the objects in the galactic coordinates, with $\ell$ on the x-axis and $b$ on the y-axis; again, colour-code everything by their "CLASS". If you did everything correctly, you should see that the curve in the previous plot resembles a horizontal band.
#
#
# 6. Answer this question: What is that curved band in the first plot and the horizontal band in the second plot? Does it make sense that the band got straightened up? Why?
#
#
# Note: When you make your plots, please include the axis labels with units and the legend.
# +
from astropy.io import fits
#You will have to import other packages to complete this problem
###IMPORT YOUR OTHER PACKAGES HERE
from astropy.table import Table
from astropy.coordinates import SkyCoord
import matplotlib.pyplot as plt
import numpy as np
# +
fits_file = fits.open('clusters.fits')
#To read the fits file as a table, simply run the line: Table.read(fits_file)
#Although you will have to write up your code to get that Table function
### YOUR CODE HERE
data = Table.read(fits_file)
CLASS = np.array(data['CLASS'])
ra_data = np.array(data['RA'])
dec_data = np.array(data['DEC'])
print(np.unique(CLASS))
RA1,DEC1 = [], []
RA2,DEC2 = [], []
RA3,DEC3 = [], []
RA4,DEC4 = [], []
RA5,DEC5 = [], []
for i in range(len(ra_data)):
if CLASS[i] == ' NEBULA\n':
RA1.append(ra_data[i])
DEC1.append(dec_data[i])
elif CLASS[i] == ' UNIDENTIFIED\n':
RA2.append(ra_data[i])
DEC2.append(dec_data[i])
elif CLASS[i] == ' OPEN STAR CLUSTER\n':
RA3.append(ra_data[i])
DEC3.append(dec_data[i])
elif CLASS[i] == ' OB ASSOCIATION/HII REGION\n':
RA4.append(ra_data[i])
DEC4.append(dec_data[i])
else:
RA5.append(ra_data[i])
DEC5.append(dec_data[i])
plt.figure(figsize=(12,8))
plt.scatter(RA1,DEC1,s = 10, c = 'red', label = 'Nebula')
plt.scatter(RA2,DEC2,s = 10, c = 'pink', label = 'Unidentified')
plt.scatter(RA3,DEC3,s = 3, c = 'lightblue', label = 'Open Star Clusters')
plt.scatter(RA4,DEC4,s = 10, c = 'orange', label = 'OB Association/Hii Region')
plt.scatter(RA5,DEC5,s = 10, c = 'green', label = 'Extragalactic')
plt.xlabel('RA in Degrees')
plt.ylabel('DEC in Degrees')
plt.legend()
plt.title('ICRS Coordinates')
plt.show()
#################################################################
#################################################################
dist = np.array(data['DISTANCE'])
icrs = SkyCoord(ra=ra_data*u.deg, dec=dec_data*u.deg)
GAL = icrs.transform_to('galactic')
L_data = np.array(GAL.l)
B_data = np.array(GAL.b)
L1,B1 = [], []
L2,B2 = [], []
L3,B3 = [], []
L4,B4 = [], []
L5,B5 = [], []
for i in range(len(ra_data)):
if CLASS[i] == ' NEBULA\n':
L1.append(L_data[i])
B1.append(B_data[i])
elif CLASS[i] == ' UNIDENTIFIED\n':
L2.append(L_data[i])
B2.append(B_data[i])
elif CLASS[i] == ' OPEN STAR CLUSTER\n':
L3.append(L_data[i])
B3.append(B_data[i])
elif CLASS[i] == ' OB ASSOCIATION/HII REGION\n':
L4.append(L_data[i])
B4.append(B_data[i])
else:
L5.append(L_data[i])
B5.append(B_data[i])
plt.figure(figsize=(12,8))
plt.scatter(L1,B1 , s = 10, c = 'red', label = 'Nebula')
plt.scatter(L2,B2 , s = 10, c = 'pink', label = 'Unidentified')
plt.scatter(L3,B3 , s = 3, c = 'lightblue', label = 'Open Star Clusters')
plt.scatter(L4,B4 , s = 10, c = 'orange', label = 'OB Association/Hii Region')
plt.scatter(L5,B5 , s = 10, c = 'green', label = 'Extragalactic')
plt.xlabel('l in Degrees')
plt.ylabel('b in Degrees')
plt.title('Galactic Coordinates')
plt.legend()
plt.show()
# -
# (DOUBLE CLICK HERE TO ANSWER QUESTION 6):
#
# YOUR ANSWER:
# # Pandas (40 Points)
#
# One of the most efficient and easy to use libraries for importing data files. We will explore the basics here.
#
# Let's import some data that represents the position of a ball being thrown off the roof of Campbell Hall. Using some basic kinematics we can derive the following equation.
#
# $$y(t) = -\frac{1}{2} g t^2 + v_{0,y} t + y_0$$
#
# For this problem we need to import our position measurements from our fellow colleagues in our research group.
#
# <img src='diagram.jpeg' width="600" height="400">
# ## Problem 5 (5 Points)
#
# Your job for this problem is to simply read in the file named **"projectile.csv"** using the pandas library (DONT USE `numpy`). Print out your DataFrame so we can see what the data looks like as a table.
# +
###YOUR CODE HERE###
import numpy as np
import matplotlib.pyplot as plt
import scipy.optimize as fitter
import pandas as pd
data = pd.read_csv('projectile.csv')
data
# -
# ## Problem 6 (5 Points)
#
# Now load your DataFrame columns into numpy arrays and make a plot of Position vs. Time.
# +
###YOUR CODE HERE###
time = data['Time[s]']
position = data['Position[m]']
plt.figure(figsize=(12,8))
plt.plot(time, position, 'ro')
plt.title('Position vs. Time')
plt.xlabel("Time [s]")
plt.ylabel("Position [m]")
plt.show()
# -
# ## Problem 7 (10 Points)
#
# In the last problem set we learned how to curve fit a quadratic equation. The above equation is also a quadratic equation with respect to time. Use what we learned last week to fit a curve to the noisy data from our fellow researchers. Explicitly print out what the initial velocity $v_{0,y}$ and initial height $y_0$ are based on your curve fit along with their respective errors.
# +
###YOUR CODE HERE###
"""This solution is from physics 77"""
#we have to define our model with our needed parameters
def model_quad(x, a, b, c):
return a*x**2 + b*x + c
par0 = np.array([-2.5, 1.5, 100.0]) # initial guess for parameters
par, cov = fitter.curve_fit(model_quad, time, position, par0) #fitter.curve_fit takes in the model, x,y data, guess, and sigma
# par arrays contains the values of parameters. cov is the covariance matrix
# decode it now
a = par[0]
ea = np.sqrt(cov[0,0])
print('a={0:6.3f}+/-{1:5.3f}'.format(a,ea))
b = par[1]
eb = np.sqrt(cov[1,1])
print('b={0:6.3f}+/-{1:5.3f}'.format(b,eb))
c = par[2]
ec = np.sqrt(cov[2,2])
print('c={0:6.3f}+/-{1:5.3f}'.format(c,ec))
print("""\n Initial velocity in the y direction is going to be 13.298 m/s and the initial
height was 97.839 m""")
plt.figure(figsize=(12,8))
plt.plot(time, model_quad(time, a,b,c))
plt.plot(time, position, 'ro')
plt.title('Position vs. Time')
plt.xlabel("Time [s]")
plt.ylabel("Position [m]")
plt.show()
# -
# ## Problem 8 (10 Points)
#
# Alright now we have a model function that can fit the function as a function of time. create two lists/arrays of values using this function. One list's values should be time where we use `t = np.linspace(0,5,100)` to create the values and the other list should be your model's output after taking in all those times. (A list of the values you would normally plot)
#
# Once you have created your two lists of values, construct a pandas DataFrame using these lists. Your data frame should have two columns with 100 values each.
# +
###Your Code Here###
t = np.linspace(0,5,100)
new_position = model_quad(t, a,b,c)
DataFrame = pd.DataFrame({'time': t, 'position': new_position})
DataFrame
# -
# ## Problem 9 (10 Points)
#
# Last part of the problem set! This is basically one line of code. Export your new DataFrame to a csv file called **"trajectory.csv"**, this will be useful for your colleagues!
# +
###Your Code Here###
DataFrame.to_csv('trajectory.csv')
# -
|
Spring2021_DeCal_Material/Homework/Week7/.ipynb_checkpoints/HW 6 Solutions-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 【課題】量子ダイナミクスシミュレーション・続
#
# 第三回の実習では量子計算の並列性と、その顕著な利用法としての量子ダイナミクスシミュレーションを取り上げました。また、実機で計算を行う際の実用的な問題として、回路の最適化や測定エラーの緩和についても議論しました。この課題はその直接の延長です。
#
# ```{contents} 目次
# ---
# local: true
# ---
# ```
# $\newcommand{\ket}[1]{|#1\rangle}$
# $\newcommand{\plusket}{\ket{+}}$
# $\newcommand{\minusket}{\ket{-}}$
# ## 問題1: ハイゼンベルグモデル、X方向のスピン
# ### 問題
#
# 実習ではハイゼンベルグモデルのシミュレーションをし、各スピンの$Z$方向の期待値の時間発展を追いました。しかし、シミュレーションそのものは最終的なオブザーバブル(観測量)によらず成立するので、(ほぼ)同じ回路を用いて系の他の性質を調べることもできます。そこで、各スピンの$X$方向の期待値の時間発展を測定する回路を書き、実習時と同様に時間に対してプロットしてください。
#
# **ヒント**:
#
# [プロット用関数`plot_heisenberg_spins`](https://github.com/UTokyo-ICEPP/qc-workbook/blob/master/source/utils/dynamics.py)で厳密解のカーブを書くとき、追加の引数`spin_component='x'`を渡すと$X$方向のスピンのプロットに切り替わります。ただし、実験結果の`counts_list`は相応する測定の結果となっていなければいけません。具体的には、各スピンについて「0が測定される=スピンが+$X$を向いている、1が測定される=スピンが-$X$を向いている」という対応付けが必要です。)
# + tags=["raises-exception", "remove-output"]
# 必要なモジュールを先にインポート
import numpy as np
import matplotlib.pyplot as plt
from qiskit import QuantumCircuit, Aer, IBMQ, transpile
from qiskit.tools.monitor import job_monitor
from qiskit.providers.ibmq import least_busy
# このワークブック独自のモジュール
from utils.dynamics import plot_heisenberg_spins, bit_expectations_sv, bit_expectations_counts, insert_initial_counts
from utils.hamiltonian import make_hamiltonian, diagonalized_evolution
# + tags=["remove-output"]
n = 5
M = 10
omegadt = 0.1
shots = 100000
# Define the circuits
circuits = []
circuit = QuantumCircuit(n)
# Bit 0 in state 1/sqrt(2)(|0> + |1>)
circuit.h(0)
for istep in range(M):
for j in range(n - 1):
# ZZ
circuit.cx(j, j + 1)
circuit.rz(-omegadt, j + 1)
circuit.cx(j, j + 1)
# XX
circuit.h(j)
circuit.h(j + 1)
circuit.cx(j, j + 1)
circuit.rz(-omegadt, j + 1)
circuit.cx(j, j + 1)
circuit.h(j)
circuit.h(j + 1)
# YY
circuit.p(-np.pi / 2., j)
circuit.p(-np.pi / 2., j + 1)
circuit.h(j)
circuit.h(j + 1)
circuit.cx(j, j + 1)
circuit.rz(-omegadt, j + 1)
circuit.cx(j, j + 1)
circuit.h(j)
circuit.h(j + 1)
circuit.p(np.pi / 2., j)
circuit.p(np.pi / 2., j + 1)
# Copy of the circuit up to this point
snapshot = circuit.copy()
##################
### EDIT BELOW ###
##################
# Set up the observable for this snapshot
# #snapshot.?
##################
### EDIT ABOVE ###
##################
snapshot.measure_all()
circuits.append(snapshot)
qasm_simulator = Aer.get_backend('qasm_simulator')
circuits = transpile(circuits, backend=qasm_simulator)
sim_job = qasm_simulator.run(circuits, shots=shots)
sim_counts_list = sim_job.result().get_counts()
# Initial state as a statevector
initial_state = np.zeros(2 ** n, dtype=np.complex128)
initial_state[0:2] = np.sqrt(0.5)
plot_heisenberg_spins(sim_counts_list, n, initial_state, omegadt, add_theory_curve=True, spin_component='x')
# -
# **提出するもの**
#
# - 完成した回路のコードとシミュレーション結果によるプロット
# - 一般の方向のスピンの期待値を測定するためにはどうすればいいかの説明
# ### おまけ: スピン総和
#
# 注:これは量子コンピューティングというより物理の問題なので、興味のある方だけ考えてみてください。
#
# 上のハイゼンベルグモデルのシミュレーションで、初期状態の$X$, $Y$, $Z$方向のスピン期待値の全系での平均値$m_x$, $m_y$, $m_z$はそれぞれ
#
# $$
# m_x = \frac{1}{n} \sum_{j=0}^{n} \langle \sigma^{X}_j \rangle = \frac{1}{n} \\
# m_y = \frac{1}{n} \sum_{j=0}^{n} \langle \sigma^{Y}_j \rangle = 0 \\
# m_z = \frac{1}{n} \sum_{j=0}^{n} \langle \sigma^{Z}_j \rangle = \frac{n-1}{n}
# $$
#
# です。これらの平均値はどう時間発展するでしょうか。理論的議論をし、シミュレーションで数値的に確かめてください。
# ## 問題2: シュウィンガーモデル
#
# これまで扱ったような、スピンに関連する現象とは異なる物理モデルのシミュレーションをしましょう。空間1次元、時間1次元の時空における量子電磁力学の模型「シュウィンガーモデル」を考えます。
# ### シュウィンガーモデルの物理
#
# 簡単に物理の解説をします(ここは読み飛ばしても差し支えありません)。といっても、まともにゼロから解説をしたらあまりにも長くなってしまうので、かなり前提知識を仮定します。興味のある方は参考文献{cite}`shifman_schwinger,Martinez_2016`などを参照してください。特に{cite}`Martinez_2016`は実際にこれから実装する回路をイオントラップ型量子コンピュータで実行した論文です。
#
# 量子電磁力学とは量子場の理論の一種です。量子場の理論とは物質やその相互作用(力)をすべて量子力学的な「場」(時空中の各点に応じた値を持つ存在)で記述した理論で、素粒子論などで物質の根源的な性質を記述する際の基本言語です。量子場の理論において、一部の場を「物質場」とし、それに特定の対称性($U(1)$ゲージ対称性)を持たせると、「電荷」が生み出され、電荷を持った場の間の相互作用を媒介する「光子場」が生じます。電荷を持った場と光子場の振る舞いを記述するのが量子電磁力学です。
#
# 量子場の理論は「ラグランジアン」[^lagrangian]を指定すれば定まります。シュウィンガーモデルのラグランジアンは物質場(電子)$\psi$とゲージ場(光子)$A$からなり、
#
# ```{math}
# :label: schwinger_lagrangian
# \mathcal{L} = -\frac{1}{4g^2} F^{\mu\nu}F_{\mu\nu} + \bar{\psi} (i\gamma^{\mu}D_{\mu} - m) \psi
# ```
#
# です。ただし、これまでの物理系を扱った話と異なり、ここでは場の量子論の一般慣習に従って、光速$c$とプランク定数$\hbar$がともに1である単位系を使っています。
#
# 式{eq}`schwinger_lagrangian`の指数$\mu, \nu$は0(時間次元)か1(空間次元)の値を取ります。$\frac{1}{2g} F_{\mu\nu}$は$A$の強度テンソル(電場)で
#
# $$
# F_{\mu\nu} = \partial_{\mu} A_{\nu} - \partial_{\nu} A_{\mu}
# $$
#
# です。$\psi$は物質と反物質を表す2元スピノルで、$m$がその質量となります。$\{\gamma^0, \gamma^1\}$は2次元のクリフォード代数の表現です。
#
# このラグランジアンを元に、Kogut-Susskindの手法{cite}`PhysRevD.10.732`でモデルを空間格子(格子間隔$a$)上の場の理論に移すと、そのハミルトニアンは
#
# ```{math}
# :label: kogut_susskind_hamiltonian
# H = \frac{1}{2a} \bigg\{ -i \sum_{j=0}^{n-2} \left[ \Phi^{\dagger}_{j} e^{i\theta_{j}} \Phi_{j+1} + \Phi_{j} e^{-i\theta_{j}} \Phi^{\dagger}_{j+1} \right] + 2 J \sum_{j=0}^{n-2} L_{j}^2 + 2 \mu \sum_{j=0}^{n-1} (-1)^{j+1} \Phi^{\dagger}_{j} \Phi_{j} \bigg\}
# ```
#
# となります。ここで$J = g^2 a^2 / 2$, $\mu = m a$, また$\Phi_j$はサイト$j$上の(1元)物質場、$\theta_j$は$j$上のゲージ場、$L_j$は格子$j$と$j+1$間の接続上の電場です。
#
# Kogut-Susskindハミルトニアンにおける物質場はstaggered fermionsと呼ばれ、隣接サイトのうち片方が物質を、もう一方が反物質を表します。約束として、ここでは$j$が偶数のサイトを物質(電荷-1)に、奇数のサイトを反物質(電荷1)に対応付けます。一般に各サイトにおける物質の状態は、フェルミ統計に従って粒子が存在する・しないという2つの状態の重ね合わせです。サイト$j$の基底$\plusket_j$と$\minusket_j$を、$\Phi_j$と$\Phi^{\dagger}_j$が
#
# ```{math}
# :label: creation_annihilation
# \Phi_j \plusket_j = \minusket_j \\
# \Phi_j \minusket_j = 0 \\
# \Phi^{\dagger}_j \plusket_j = 0 \\
# \Phi^{\dagger}_j \minusket_j = \plusket_j
# ```
#
# と作用する状態と定めます。質量項の符号から、偶数サイトでは$\minusket$が粒子が存在する状態、$\plusket$が存在しない状態を表現し、奇数サイトでは逆に$\plusket$が粒子あり、$\minusket$が粒子なしを表すことがわかります。つまり、$\Phi^{\dagger}_j$と$\Phi_j$はサイト$j$における電荷の上昇と下降を引き起こす演算子です。
# ### ハミルトニアンを物質場のみで記述する
#
# $\newcommand{\mfrac}[2]{\genfrac{}{}{2pt}{0}{#1}{#2}}$
#
# このままのハミルトニアンではまだデジタルモデルが構築しにくいので、ゲージを固定して$\theta$と$L$を除いてしまいます[^another_approach]。まず$\Phi_j$を以下のように再定義します。
#
# $$
# \Phi_j \rightarrow \prod_{k=0}^{j-1} e^{-i\theta_{k}} \Phi_j.
# $$
#
# また、ガウスの法則から、サイト$j$の電荷$\rho_j$が同じサイトの電場の発散と等しいので、
#
# $$
# L_j - L_{j-1} = \rho_j \\
# \therefore L_j = \sum_{k=0}^{j} \rho_k
# $$
#
# となります。ただし、サイト0に系の境界の外から作用する電場はないもの($L_{-1} = 0$)としました。
#
# 質量項と同様にサイトの偶奇を考慮した電荷は
#
# $$
# \rho_k = \Phi_{k}^{\dagger} \Phi_{k} - (k+1 \bmod 2)
# $$
#
# なので、
#
# $$
# L_j = \sum_{k=0}^{j} \Phi_{k}^{\dagger} \Phi_{k} - \mfrac{j}{2} - 1
# $$
#
# となります。ここで太線の分数$\genfrac{}{}{2pt}{1}{j}{2}$は切り捨ての割り算$[j - (j \bmod 2)]/2$(Pythonでの`j // 2`と同等)です。この電場を式{eq}`kogut_susskind_hamiltonian`に代入して
#
# $$
# H = \frac{1}{2a} \left\{ -i \sum_{j=0}^{n-2} \left[ \Phi^{\dagger}_{j} \Phi_{j+1} + \Phi_j \Phi^{\dagger}_{j+1} \right] + 2J \sum_{j=0}^{n-2} \left[\sum_{k=0}^{j} \Phi_{k}^{\dagger} \Phi_{k} - \mfrac{j}{2} - 1 \right]^2 + 2\mu \sum_{j=0}^{n-1} (-1)^{j+1} \Phi^{\dagger}_{j} \Phi_{j} \right\}
# $$
#
# が得られます。
# ### ハミルトニアンをパウリ行列で表現する
#
# 最後に、$\plusket$と$\minusket$をスピン$\pm Z$の状態のようにみなして、$\Phi^{\dagger}_j\Phi_j$と$\Phi^{\dagger}_j\Phi_{j+1}$をパウリ行列で表現します。式{eq}`creation_annihilation`から
# 前者は
#
# $$
# \Phi^{\dagger}_j\Phi_j \rightarrow \frac{1}{2} (\sigma^Z_j + 1)
# $$
#
# と表現できることがわかります。一方、$\Phi^{\dagger}_j\Phi_{j+1}$に関しては、やや込み入った議論{cite}`PhysRevD.13.1043`の末、
#
# $$
# \Phi^{\dagger}_j\Phi_{j+1} \rightarrow i \sigma^+_j \sigma^-_{j+1}
# $$
#
# が正しい表現であることがわかっています。ここで、
#
# $$
# \sigma^{\pm} = \frac{1}{2}(\sigma^X \pm i \sigma^Y)
# $$
#
# です。ハミルトニアンには$\Phi_j\Phi^{\dagger}_{j+1} \rightarrow i \sigma^-_j \sigma^+_{j+1}$も登場するので、二つの項を合わせると
#
# $$
# \Phi^{\dagger}_{j} \Phi_{j+1} + \Phi_j \Phi^{\dagger}_{j+1} \rightarrow \frac{i}{2} (\sigma^X_j \sigma^X_{j+1} + \sigma^Y_j \sigma^Y_{j+1})
# $$
#
# となります。まとめると、
#
# $$
# H \rightarrow \frac{1}{4a} \left\{ \sum_{j=0}^{n-2} (\sigma^X_j \sigma^X_{j+1} + \sigma^Y_j \sigma^Y_{j+1}) + J \sum_{j=1}^{n-2} (n - j - 1) \sum_{k=0}^{j-1} \sigma^Z_k \sigma^Z_j + \sum_{j=0}^{n-1} \left[ (-1)^{j+1} \mu - J \mfrac{n-j}{2} \right] \sigma^Z_j \right\}
# $$
#
# です。ただし、計算過程で現れる定数項(恒等演算子に比例する項)は時間発展において系の状態に全体位相をかける作用しか持たないため、無視しました。
# ### 問題
#
# 上のシュウィンガーモデルのハミルトニアンによる時間発展シミュレーションを、$\plusket$と$\minusket$をそれぞれ$\ket{0}$と$\ket{1}$に対応させて、8ビット量子レジスタに対して実装してください。初期状態は真空、つまり$\ket{-+-+-+-+}$(右端がサイト$j=0$)とし、系全体の粒子数密度の期待値
#
# $$
# \nu = \left\langle \frac{1}{n} \sum_{j=0}^{n-1} \frac{1}{2} \left[(-1)^{j+1} \sigma^Z_j + 1\right] \right\rangle
# $$
#
# を時間の関数としてプロットしてください。余裕があれば、各サイトにおける粒子数、電荷、サイト間の電場などの期待値の時間変化も観察してみましょう。
#
# ハミルトニアンのパラメターは、$J = 1$, $\mu = 0.5$とします(他の$J$や$\mu$の値もぜひ試してみてください)。$\omega = 1/(2a)$とおき、鈴木・トロッター分解における時間ステップ$\Delta t$の大きさ$\omega \Delta t = 0.2$として、時間$\omega t = 2$までシミュレーションをします。
#
# **解説**:
#
# 偶数サイトでは$\plusket$が物質粒子の存在しない状態、奇数サイトでは$\minusket$が反物質粒子の存在しない状態を表すので、初期状態は粒子数密度0となります。しかし、場の量子論においては場の相互作用によって物質と反物質が対生成・対消滅を起こし、一般に系の粒子数の期待値は時間とともに変化します。
#
# **ヒント**:
#
# 上のハミルトニアンのパラメターの値は参考文献{cite}`Martinez_2016`と同一です。したがって、$n=4$, $\omega \Delta t = \pi/8$とすれば、論文中の図3aを再現できるはずです。答え合わせに使ってください。
#
# また、問題を解くためのヒントではありませんが、ハイゼンベルグモデルと同様にこのモデルでも対角化による厳密解を比較的簡単にプロットできるように道具立てがしてあります。下のコードのテンプレートでは、シミュレーション回路と厳密解を計算するためのハミルトニアンのパウリ行列分解だけ指定すれば、`plot_heisenberg_spins`と同様のプロットが作成されるようになっています。パウリ行列分解を指定するには、`paulis`と`coeffs`という二つのリストを作ります。これらのリストの長さはハミルトニアンの項数で、`paulis`の各要素は対応する項のパウリ行列のリスト、`coeffs`の各要素はその項にかかる係数にします。例えば
#
# $$
# H = 0.5 \sigma^X_0 \sigma^Y_1 I_2 + I_0 \sigma^Z_1 \sigma^X_2
# $$
#
# というハミルトニアンに対しては、
#
# ```{code-block} python
# paulis = [['x', 'y', 'i'], ['i', 'z', 'x']]
# coeffs = [0.5, 1.]
# ```
#
# です。
#
# [^lagrangian]: ここで「ラグランジアン」と呼ばれているのは本来「ラグランジアン密度」で、正しくはこれを空間積分したものがラグランジアンですが、素粒子論の文脈で「ラグランジアン」と言った場合はほぼ100%積分する前のものを指します。
# [^another_approach]: 参考文献{cite}`Shaw2020quantumalgorithms`では、別のアプローチで同じハミルトニアンの量子回路実装をしています。
# + tags=["raises-exception", "remove-output"]
def number_density(bit_exp):
particle_number = np.array(bit_exp) # shape (T, n)
# Particle number is 1 - (bit expectation) on odd sites
particle_number[:, fc00:db20:35b:7399::5] = 1. - particle_number[:, 1::2]
return np.mean(particle_number, axis=1)
n = 8 # number of sites
J = 1. # Hamiltonian J parameter
mu = 0.5 # Hamiltonian mu parameter
## Quantum circuit experiment
M = 10 # number of Trotter steps
omegadt = 0.2 # Trotter step size
shots = 100000
# Define the circuits
circuits = []
circuit = QuantumCircuit(n)
# Initial state = vacuum
circuit.x(range(1, n, 2))
for istep in range(M):
##################
### EDIT BELOW ###
##################
# #circuit.?
##################
### EDIT ABOVE ###
##################
circuits.append(circuit.measure_all(inplace=False))
# Run the circuits in the simulator
qasm_simulator = Aer.get_backend('qasm_simulator')
circuits = transpile(circuits, backend=qasm_simulator)
sim_job = qasm_simulator.run(circuits, shots=shots)
sim_counts_list = sim_job.result().get_counts()
## Numerical solution through diagonalization
# Construct the Hamiltonian
paulis = []
coeffs = []
##################
### EDIT BELOW ###
##################
paulis = [['i'] * n]
coeffs = None
##################
### EDIT ABOVE ###
##################
hamiltonian = make_hamiltonian(paulis, coeffs)
# Initial state as a statevector
initial_state = np.zeros(2 ** n, dtype=np.complex128)
vacuum_state_index = 0
for j in range(1, n, 2):
vacuum_state_index += (1 << j)
initial_state[vacuum_state_index] = 1.
## Plotting
# Plot the exact solution
time_points, statevectors = diagonalized_evolution(hamiltonian, initial_state, omegadt * M)
_, bit_exp = bit_expectations_sv(time_points, statevectors)
plt.plot(time_points, number_density(bit_exp))
# Plot the simulation results
time_points = np.linspace(0., omegadt * M, M + 1, endpoint=True)
insert_initial_counts(sim_counts_list, initial_state)
_, bit_exp = bit_expectations_counts(time_points, sim_counts_list, n)
plt.plot(time_points, number_density(bit_exp), 'o')
# + [markdown] tags=["remove-output"]
# **提出するもの**
#
# - 完成した回路のコードとシミュレーション結果によるプロット
# -
# ## 参考文献
#
# ```{bibliography}
# :filter: docname in docnames
# ```
|
source/ja/more_dynamics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Simple-ML-MODEL" data-toc-modified-id="Simple-ML-MODEL-1"><span class="toc-item-num">1 </span>Simple ML MODEL</a></span><ul class="toc-item"><li><span><a href="#Logistic-Regression" data-toc-modified-id="Logistic-Regression-1.1"><span class="toc-item-num">1.1 </span>Logistic Regression</a></span></li><li><span><a href="#Naive-Bayes" data-toc-modified-id="Naive-Bayes-1.2"><span class="toc-item-num">1.2 </span>Naive Bayes</a></span></li><li><span><a href="#Lasso-Regression" data-toc-modified-id="Lasso-Regression-1.3"><span class="toc-item-num">1.3 </span>Lasso Regression</a></span></li><li><span><a href="#Support-Vector-Machines" data-toc-modified-id="Support-Vector-Machines-1.4"><span class="toc-item-num">1.4 </span>Support Vector Machines</a></span></li></ul></li><li><span><a href="#Models-with-Train-Test-Split" data-toc-modified-id="Models-with-Train-Test-Split-2"><span class="toc-item-num">2 </span>Models with Train-Test Split</a></span><ul class="toc-item"><li><span><a href="#Logistic-Regression-(Train-Test-Split)" data-toc-modified-id="Logistic-Regression-(Train-Test-Split)-2.1"><span class="toc-item-num">2.1 </span>Logistic Regression (Train-Test Split)</a></span><ul class="toc-item"><li><span><a href="#Logistic-Regression-Accuracy-on-Train-data" data-toc-modified-id="Logistic-Regression-Accuracy-on-Train-data-2.1.1"><span class="toc-item-num">2.1.1 </span>Logistic Regression Accuracy on Train data</a></span></li><li><span><a href="#Logistic-Regression-Accuracy-on-Test-data" data-toc-modified-id="Logistic-Regression-Accuracy-on-Test-data-2.1.2"><span class="toc-item-num">2.1.2 </span>Logistic Regression Accuracy on Test data</a></span></li></ul></li><li><span><a href="#Support-Vector-Machine-(Train-Test-Split)" data-toc-modified-id="Support-Vector-Machine-(Train-Test-Split)-2.2"><span class="toc-item-num">2.2 </span>Support Vector Machine (Train-Test Split)</a></span><ul class="toc-item"><li><span><a href="#Support-Vector-Machine-Accuracy-on-Train-data" data-toc-modified-id="Support-Vector-Machine-Accuracy-on-Train-data-2.2.1"><span class="toc-item-num">2.2.1 </span>Support Vector Machine Accuracy on Train data</a></span></li><li><span><a href="#Support-Vector-Machine-Accuracy-on-Test-data" data-toc-modified-id="Support-Vector-Machine-Accuracy-on-Test-data-2.2.2"><span class="toc-item-num">2.2.2 </span>Support Vector Machine Accuracy on Test data</a></span></li></ul></li></ul></li></ul></div>
# -
# # Simple ML MODEL
import pickle
import random
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn import linear_model
from imblearn.over_sampling import SMOTE
from sklearn.datasets import make_classification
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler, scale
from sklearn.metrics import classification_report, confusion_matrix
sns.set_style('whitegrid')
random.seed(123)
data = pd.read_csv('Heart.csv', sep = ";")
data.sample(5)
data.info()
# Checking for duplicates
duplicate_data = data[data.duplicated()]
duplicate_data
# Counting the number of duplicate values
data.duplicated().sum()
# Data set length before removing duplicates
len(data)
# Removing duplicate data
data.drop_duplicates(keep='first', inplace=True)
# Data set length after removing duplicates
len(data)
# double checking for duplicates
duplicate_data = data[data.duplicated()]
duplicate_data
# Counting the number of duplicate values
data.duplicated().sum()
# Percentage of missing values
(data.isnull().sum() * 100 / data.shape[0]).sort_values(ascending=False)
cust_col = ["#1f77b4","#ff7f0e"] # own custom color scheme
sns.set_palette(cust_col) # set color scheme
total = len(data)
plt.figure(figsize=(7,5))
g = sns.countplot(x='target', data=data)
g.set_ylabel('Count', fontsize=14)
for p in g.patches:
height = p.get_height()
g.text(p.get_x()+p.get_width()/2.,
height + 1.5,
'{:1.2f}%'.format(height/total*100),
ha="center", fontsize=14, fontweight='bold')
plt.margins(y=0.1)
plt.title('Patients with Heart Disease');
plt.xlabel('Heart Disease');
plt.ylabel('Number of Patients');
plt.show()
# ## Logistic Regression
# # copy data to avoid modifying original data
lr_data = data.copy()
#split dataset in features and target variable
X = lr_data.loc[:, lr_data.columns != 'target']
y = lr_data.target
X_t = scale(X)
lr_model = LogisticRegression()
lr_model.fit(X_t,y)
print(classification_report(y, lr_model.predict(X_t)))
# save the model to disk
filename = 'Saved_Experiment_Models/LR_model_experiment.pkl'
pickle.dump(lr_model, open(filename, 'wb'))
# load the model from disk
loaded_model = pickle.load(open(filename, 'rb'))
data.head(2)
# Use Model for prediction (Predicting the Row 0)
loaded_model.predict(scale([[63, 1, 3, 145, 233, 1, 0, 150, 0, 2.3, 0, 0, 1]]))
# ## Naive Bayes
# # copy data to avoid modifying original data
nb_data = data.copy()
#split dataset in features and target variable
X = nb_data.loc[:, nb_data.columns != 'target']
y = nb_data.target # Target feature
X_t = scale(X)
from sklearn.naive_bayes import GaussianNB
nb_model = GaussianNB()
nb_model.fit(X_t,y)
print(classification_report(y, nb_model.predict(X_t)))
# save the model to disk
filename2 = 'Saved_Experiment_Models/NB_model_experiment.pkl'
pickle.dump(nb_model, open(filename2, 'wb'))
# load the model from disk
loaded_NBmodel = pickle.load(open(filename2, 'rb'))
# make prediction with loaded model
loaded_NBmodel.predict(scale([[63, 1, 3, 145, 233, 1, 0, 150, 0, 2.3, 0, 0, 1]]))
# ## Lasso Regression
# # copy data to avoid modifying original data
la_data = data.copy()
#split dataset in features and target variable
X = la_data.loc[:, nb_data.columns != 'target']
y = la_data.target # Target feature
X_t = scale(X)
from sklearn import linear_model
la_model = linear_model.Lasso(alpha=0.1)
la_model.fit(X_t,y)
la_cls = la_model.predict(X_t)
cls = np.array([1 if x>0.5 else 0 for x in la_cls])
print(classification_report(y, cls))
# ## Support Vector Machines
# Support Vector Machine is considered as a classification approach, but it can be applied to both classification and regression problems.
#
# More on SVM: [Wikipedia](https://en.wikipedia.org/wiki/Support-vector_machine)
# # copy data to avoid modifying original data
svm_data = data.copy()
#split dataset in features and target variable
X = svm_data.loc[:, svm_data.columns != 'target']
y = svm_data.target # Target feature
X_t = scale(X)
from sklearn import svm
svm_model = svm.SVC(kernel='linear')
svm_model.fit(X_t,y)
print(classification_report(y, svm_model.predict(X_t)))
# save the model to disk
filename4 = 'Saved_Experiment_Models/svm_model.pkl'
pickle.dump(svm_model, open(filename4, 'wb'))
# load the model from disk
loaded_SVMmodel = pickle.load(open(filename4, 'rb'))
# Use Model for prediction (Predicting the Row 0)
loaded_SVMmodel.predict(scale([[63, 1, 3, 145, 233, 1, 0, 150, 0, 2.3, 0, 0, 1]]))
# # Models with Train-Test Split
# - `Best Performing Models:` Logistic Regression and Support Vector Machine
from sklearn.model_selection import train_test_split
# ## Logistic Regression (Train-Test Split)
# # copy data to avoid modifying original data
lr2_data = data.copy()
#split dataset in features and target variable
X = lr2_data.loc[:, lr2_data.columns != 'target']
y = lr2_data.target
X_t = scale(X)
# 80% Training and 20% Testing
X_train, X_test, y_train, y_test = train_test_split(X_t, y, test_size=0.2,random_state=109)
lr2_model = LogisticRegression()
lr2_model.fit(X_train,y_train)
# ### Logistic Regression Accuracy on Train data
print(classification_report(y_train, lr2_model.predict(X_train)))
# ### Logistic Regression Accuracy on Test data
print(classification_report(y_test, lr2_model.predict(X_test)))
# ## Support Vector Machine (Train-Test Split)
# # copy data to avoid modifying original data
svm2_data = data.copy()
#split dataset in features and target variable
X = svm2_data.loc[:, svm2_data.columns != 'target']
y = svm2_data.target
X_t = scale(X)
# 80% Training and 20% Testing
X_train, X_test, y_train, y_test = train_test_split(X_t, y, test_size=0.2,random_state=109)
svm2_model = svm.SVC(kernel='linear')
svm2_model.fit(X_train,y_train)
# ### Support Vector Machine Accuracy on Train data
print(classification_report(y_train, svm2_model.predict(X_train)))
# ### Support Vector Machine Accuracy on Test data
print(classification_report(y_test, svm2_model.predict(X_test)))
|
Model_Experiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SageMaker Model Monitor - visualizing monitoring results
#
#
# The prebuilt container from SageMaker computes a variety of statistics and evaluates constraints out of the box. This notebook demonstrates how you can visualize them. You can grab the ProcessingJob arn from the executions behind a MonitoringSchedule and use this notebook to visualize the results.
#
# Let's import some python libraries that will be helpful for visualization
# +
from IPython.display import HTML, display
import json
import os
import boto3
import sagemaker
from sagemaker import session
from sagemaker.model_monitor import MonitoringExecution
from sagemaker.s3 import S3Downloader
# -
# ## Get Utilities for Rendering
#
# The functions for plotting and rendering distribution statistics or constraint violations are implemented in a `utils` file so let's grab that.
# +
# !wget https://raw.githubusercontent.com/awslabs/amazon-sagemaker-examples/master/sagemaker_model_monitor/visualization/utils.py
import utils as mu
# -
# ## Get Execution and Baseline details from Processing Job Arn
#
# Enter the ProcessingJob arn for an execution of a MonitoringSchedule below to get the result files associated with that execution
processing_job_arn = "FILL-IN-PROCESSING-JOB-ARN"
execution = MonitoringExecution.from_processing_arn(sagemaker_session=session.Session(), processing_job_arn=processing_job_arn)
exec_inputs = {inp['InputName']: inp for inp in execution.describe()['ProcessingInputs']}
exec_results = execution.output.destination
# +
baseline_statistics_filepath = exec_inputs['baseline']['S3Input']['S3Uri'] if 'baseline' in exec_inputs else None
execution_statistics_filepath = os.path.join(exec_results, 'statistics.json')
violations_filepath = os.path.join(exec_results, 'constraint_violations.json')
baseline_statistics = json.loads(S3Downloader.read_file(baseline_statistics_filepath)) if baseline_statistics_filepath is not None else None
execution_statistics = json.loads(S3Downloader.read_file(execution_statistics_filepath))
violations = json.loads(S3Downloader.read_file(violations_filepath))['violations']
# -
# ## Overview
#
# The code below shows the violations and constraint checks across all features in a simple table.
mu.show_violation_df(baseline_statistics=baseline_statistics, latest_statistics=execution_statistics, violations=violations)
# ## Distributions
#
# This section visualizes the distribution and renders the distribution statistics for all features
features = mu.get_features(execution_statistics)
feature_baselines = mu.get_features(baseline_statistics)
mu.show_distributions(features)
# ### Execution Stats vs Baseline
mu.show_distributions(features, feature_baselines)
|
sagemaker_model_monitor/visualization/.ipynb_checkpoints/SageMaker-Model-Monitor-Visualize-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Create a Target Forecast
# * **About 40 mins may be elapsed**
import boto3
from time import sleep
import pandas as pd
import json
import time
import pprint
import numpy as np
# Recover variables stored by other notebooks
# %store -r
session = boto3.Session(region_name=region)
forecast = session.client(service_name='forecast')
# ## Create Prophet and DeepAR+ Campaign
# Prophet
prophet_forecastName = project+'_prophet_algo_forecast' + target_suffix + suffix
prophet_create_forecast_response=forecast.create_forecast(
ForecastName=prophet_forecastName,
PredictorArn=target_prophet_predictorArn)
target_prophet_forecast_arn = prophet_create_forecast_response['ForecastArn']
forecast.describe_forecast(ForecastArn = target_prophet_forecast_arn)
# DeepAR+
deeparp_forecastName = project+'_deeparp_algo_forecast' + target_suffix + suffix
deeparp_create_forecast_response=forecast.create_forecast(
ForecastName=deeparp_forecastName,
PredictorArn=target_deepar_predictorArn)
target_deeparp_forecast_arn = deeparp_create_forecast_response['ForecastArn']
forecast.describe_forecast(ForecastArn = target_deeparp_forecast_arn)
# +
# %%time
# Check the Prophet status
while True:
createProphetStatus = forecast.describe_forecast(ForecastArn= target_prophet_forecast_arn)['Status']
createDeeparpStatus = forecast.describe_forecast(ForecastArn= target_deeparp_forecast_arn)['Status']
print("Prophet: ", createProphetStatus)
print("DeepARP: ", createProphetStatus)
if createProphetStatus != 'ACTIVE' and createProphetStatus != 'CREATE_FAILED':
sleep(60)
elif createDeeparpStatus != 'ACTIVE' and createDeeparpStatus != 'CREATE_FAILED':
sleep(60)
else:
break
# -
# ## Upload forecast results to S3
target_prophet_path = "s3://" + bucket_name + "/" + bucket_folder + "/prophet_" + target_suffix + suffix + "/"
target_prophet_job_name = "ProphetExport1" + target_suffix + suffix
create_forecast_export_job_prophet_response = forecast.create_forecast_export_job(
ForecastExportJobName = target_prophet_job_name,
ForecastArn = target_prophet_forecast_arn,
Destination={
"S3Config" : {
"Path": target_prophet_path,
"RoleArn": role_arn
}
})
TargetForecastProphetExportJobArn = create_forecast_export_job_prophet_response["ForecastExportJobArn"]
forecast.describe_forecast_export_job(ForecastExportJobArn = TargetForecastProphetExportJobArn)
target_deeparp_path = "s3://" + bucket_name + "/" + bucket_folder + "/deeparp_" + target_suffix + suffix + "/"
target_deeparp_job_name = "DeepARPExport1" + target_suffix + suffix
create_forecast_export_job_deepar_response = forecast.create_forecast_export_job(
ForecastExportJobName = target_deeparp_job_name,
ForecastArn = target_deeparp_forecast_arn,
Destination={
"S3Config" : {
"Path": target_deeparp_path,
"RoleArn": role_arn
}
})
TargetForecastDeeparExportJobArn = create_forecast_export_job_deepar_response["ForecastExportJobArn"]
forecast.describe_forecast_export_job(ForecastExportJobArn = TargetForecastDeeparExportJobArn)
# +
# %%time
# Check the Prophet status
while True:
createProphetStatus = forecast.describe_forecast_export_job(ForecastExportJobArn= TargetForecastProphetExportJobArn)['Status']
createDeeparpStatus = forecast.describe_forecast_export_job(ForecastExportJobArn= TargetForecastDeeparExportJobArn)['Status']
print("Prophet: ", createProphetStatus)
print("DeepARP: ", createProphetStatus)
if createProphetStatus != 'ACTIVE' and createProphetStatus != 'CREATE_FAILED':
sleep(60)
elif createDeeparpStatus != 'ACTIVE' and createDeeparpStatus != 'CREATE_FAILED':
sleep(60)
else:
break
# +
# %store target_prophet_forecast_arn
# %store target_deeparp_forecast_arn
# %store TargetForecastProphetExportJobArn
# %store TargetForecastDeeparExportJobArn
# -
|
WalmartSale/working/1.4.Create_Target_Forecast.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Test Your Algorithm
#
# ## Instructions
# 1. From the **Pulse Rate Algorithm** Notebook you can do one of the following:
# - Copy over all the **Code** section to the following Code block.
# - Download as a Python (`.py`) and copy the code to the following Code block.
# 2. In the bottom right, click the <span style="color:blue">Test Run</span> button.
#
# ### Didn't Pass
# If your code didn't pass the test, go back to the previous Concept or to your local setup and continue iterating on your algorithm and try to bring your training error down before testing again.
#
# ### Pass
# If your code passes the test, complete the following! You **must** include a screenshot of your code and the Test being **Passed**. Here is what the starter filler code looks like when the test is run and should be similar. A passed test will include in the notebook a green outline plus a box with **Test passed:** and in the Results bar at the bottom the progress bar will be at 100% plus a checkmark with **All cells passed**.
# 
#
# 1. Take a screenshot of your code passing the test, make sure it is in the format `.png`. If not a `.png` image, you will have to edit the Markdown render the image after Step 3. Here is an example of what the `passed.png` would look like
# 2. Upload the screenshot to the same folder or directory as this jupyter notebook.
# 3. Rename the screenshot to `passed.png` and it should show up below.
# 
# 4. Download this jupyter notebook as a `.pdf` file.
# 5. Continue to Part 2 of the Project.
# + edited=false gradable=true grader_id="nrtnppao4pm" udacity_user_query=""
import glob
import numpy as np
import scipy as sp
import scipy.io
import scipy.signal
import scipy.stats
def LoadTroikaDataset():
"""
Retrieve the .mat filenames for the troika dataset.
Review the README in ./datasets/troika/ to understand the organization of the .mat files.
Returns:
data_fls: Names of the .mat files that contain signal data
ref_fls: Names of the .mat files that contain reference data
<data_fls> and <ref_fls> are ordered correspondingly, so that ref_fls[5] is the
reference data for data_fls[5], etc...
"""
data_dir = "./datasets/troika/training_data"
data_fls = sorted(glob.glob(data_dir + "/DATA_*.mat"))
ref_fls = sorted(glob.glob(data_dir + "/REF_*.mat"))
return data_fls, ref_fls
def LoadTroikaDataFile(data_fl):
"""
Loads and extracts signals from a troika data file.
Usage:
data_fls, ref_fls = LoadTroikaDataset()
ppg, accx, accy, accz = LoadTroikaDataFile(data_fls[0])
Args:
data_fl: (str) filepath to a troika .mat file.
Returns:
numpy arrays for ppg, accx, accy, accz signals.
"""
data = sp.io.loadmat(data_fl)['sig']
return data[2:]
def AggregateErrorMetric(pr_errors, confidence_est):
"""
Computes an aggregate error metric based on confidence estimates.
Computes the MAE at 90% availability.
Args:
pr_errors: a numpy array of errors between pulse rate estimates and corresponding
reference heart rates.
confidence_est: a numpy array of confidence estimates for each pulse rate
error.
Returns:
the MAE at 90% availability
"""
# Higher confidence means a better estimate. The best 90% of the estimates
# are above the 10th percentile confidence.
percentile90_confidence = np.percentile(confidence_est, 10)
# Find the errors of the best pulse rate estimates
best_estim0tes = pr_errors[confidence_est >= percentile90_confidence]
# Return the mean absolute error
return np.mean(np.abs(best_estimates))
def Evaluate():
"""
Top-level function evaluation function.
Runs the pulse rate algorithm on the Troika dataset and returns an aggregate error metric.
Returns:
Pulse rate error on the Troika dataset. See AggregateErrorMetric.
"""
# Retrieve dataset files
data_fls, ref_fls = LoadTroikaDataset()
errs, confs = [], []
for data_fl, ref_fl in zip(data_fls, ref_fls):
# Run the pulse rate algorithm on each trial in the dataset
errors, confidence = RunPulseRateAlgorithm(data_fl, ref_fl)
errs.append(errors)
confs.append(confidence)
# Compute aggregate error metric
errs = np.hstack(errs)
confs = np.hstack(confs)
return AggregateErrorMetric(errs, confs)
def bandpass_filter(signal, fs):
"""
Runs a bandpass filter with butterworth algorithm.
Returns:
Bandpass filtered signal
"""
pass_band=(40/60.0, 240/60.0)
b, a = scipy.signal.butter(2, pass_band, btype='bandpass', fs=fs)
return scipy.signal.filtfilt(b, a, signal)
def fourier_transform(signal, fs):
"""
Run a Fourier Transform on a signal
Returns:
Freq and Magnitude of the signal after running the FFT
"""
freqs = np.fft.rfftfreq(2*len(signal), 1/fs)
fft = np.abs(np.fft.rfft(signal, 2*len(signal)))
return freqs, fft
def get_features(ppg, acc, fs):
"""
Get features in our data
Returns:
PPG and ACC: Peaks, Peaks Freqency, Frequency and Magnitude after running FFT
"""
# Fourier Transform the ppg signal
ppg_freqs, ppg_fft = fourier_transform(ppg, fs)
# Filter data
ppg_fft[ppg_freqs <= 70/60.0] = 0.0
ppg_fft[ppg_freqs >= 190/60.0] = 0.0
# Find peaks
ppg_peaks = sp.signal.find_peaks(ppg_fft, height=2000)[0]
ppg_peaks_f = ppg_freqs[ppg_peaks]
# Fourier Transform the acc signal
acc_freqs, acc_fft = fourier_transform(acc, fs)
# Filter data
acc_fft[acc_freqs <= 70/60.0] = 0.0
acc_fft[acc_freqs >= 190/60.0] = 0.0
# Find peaks
acc_peaks = sp.signal.find_peaks(acc_fft, height=None)[0]
acc_peaks_f = acc_freqs[acc_peaks]
return {'ppg': (ppg_peaks, ppg_peaks_f, ppg_freqs, ppg_fft), 'acc': (acc_peaks, acc_peaks_f, acc_freqs, acc_fft)}
def calculate_confidence(freqs, fft_mag, bpm_f):
"""
Calculate the confidence of the heart rate
Returns:
Confidence of the estimated heart rate
"""
window_f = 30/60
fundamental_freq_window = (freqs > bpm_f - window_f) & (freqs < bpm_f + window_f)
return np.sum(fft_mag[fundamental_freq_window])/ np.sum(fft_mag)
def estimate(ppg, acc, window_length_s, window_shift_s, fs):
"""
Estimate the heart rate
Returns:
The estimated heart rate and the confidence
"""
window_length = window_length_s * fs
window_shift = window_shift_s * fs
est_bpm = []
confidence = []
prev_est = 40/60
for i in range(0, len(ppg) - window_length, window_shift):
ppg_window = ppg[i:i+window_length]
acc_window = acc[i:i+window_length]
window_features = get_features(ppg_window, acc, fs)
ppg_peaks, ppg_peaks_f, ppg_freqs, ppg_fft = window_features['ppg']
acc_peaks, acc_peaks_f, acc_freqs, acc_fft = window_features['acc']
ppg_max = ppg_freqs[np.argmax(ppg_fft)]
acc_max = acc_freqs[np.argmax(acc_fft)]
k = 1
while np.abs(acc_max-ppg_max) <= 0.2 and k <=2:
k+=1
ppg_max = ppg_freqs[np.argsort(ppg_fft, axis=0)[-k]]
acc_max = acc_freqs[np.argsort(acc_fft, axis=0)[-k]]
est_bpm_f = ppg_max
prev_est = est_bpm_f
est_bpm.append(est_bpm_f*60)
confidence.append(calculate_confidence(ppg_freqs, ppg_fft, est_bpm_f))
return est_bpm, confidence
def RunPulseRateAlgorithm(data_fl, ref_fl):
# Load data using LoadTroikaDataFile
ppg, accx, accy, accz = LoadTroikaDataFile(data_fl)
bpm = sp.io.loadmat(ref_fl)['BPM0']
# Our data is sampled at 125Hz
fs = 125
# Bandpass Filter
ppg = bandpass_filter(ppg, fs)
accx = bandpass_filter(accx, fs)
accy = bandpass_filter(accy, fs)
accz = bandpass_filter(accz, fs)
# Calculate the magnitute of the accelerometers
acc = np.sqrt(accx**2 + accy**2 + accz**2)
# Windowing our ppg and acc signal to estimate
window_length_s = 8
window_shift_s = 2
# Compute pulse rate estimates and estimation confidence.
est_bpm, confidence = estimate(ppg, acc, window_length_s, window_shift_s, fs)
# Return per-estimate mean absolute error and confidence as a 2-tuple of numpy arrays.
errors = np.abs(np.diag(np.subtract(bpm, est_bpm)))
return errors, confidence
|
part-1-github/test/unit_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="CWMAYFVtl5Ah" outputId="26bbf00b-f048-4ee1-aa53-401b57c7225a"
# Mount google drive files
from google.colab import drive
drive.mount("/content/drive", force_remount=True)
# + id="WDoUkrQ6qlMF"
# Install packages
# !pip install git+https://github.com/seankmartin/NeuroChaT -q
# !pip install git+https://github.com/seankmartin/PythonUtils -q
# !pip install git+https://github.com/seankmartin/SIMURAN -q
# !pip install git+https://github.com/seankmartin/lfp_atn -q
# !pip install mne -q
# + id="h4RN5fHbqdMx"
# Import libraries
import os
import matplotlib.pyplot as plt
import simuran
from lfp_atn_simuran.analysis.frequency_analysis import powers
from tqdm.notebook import tqdm
# + id="3ONXcXauqLRx"
# Configuration
path_dir = "/content/drive/My Drive/NeuroScience/ATN_CA1"
temp_storage_location = "/content/drive/My Drive/NeuroScience/Temp"
index_location = os.path.join(temp_storage_location, "index.csv")
mapping_params_location = os.path.join(temp_storage_location, "mapping.py")
os.makedirs(os.path.dirname(index_location), exist_ok=True)
nc_loader_kwargs = {
"system": "Axona",
"pos_extension": ".pos"
}
clean_kwargs = {
"pick_property": "group",
"channels": ["LFP"],
}
all_params = {
# Cleaning params
"clean_method": "pick",
"clean_kwargs": clean_kwargs,
# Filtering params
"fmin": 1,
"fmax": 100,
"theta_min": 6,
"theta_max": 10,
"delta_min": 1.5,
"delta_max": 4,
# Plotting params
"psd_scale": "decibels",
"image_format": "png",
# Path setup
"cfg_base_dir" : "/content/drive/My Drive/NeuroScience/ATN_CA1",
# STA
"number_of_shuffles_sta": 5
}
# + id="BUAzLh9Gw48F"
# Write out the mapping file
def setup_signals():
"""Set up the signals (such as eeg or lfp)."""
# The total number of signals in the recording
num_signals = 32
# What brain region each signal was recorded from
regions = ["CA1"] * 32
# If the wires were bundled, or any other kind of grouping existed
# If no grouping, grouping = [i for in range(num_signals)]
groups = ["LFP", "LFP", "LFP", "LFP"] + [i for i in range(num_signals - 4)]
# The sampling rate in Hz of each signal
sampling_rate = [250] * num_signals
# This just passes the information on
output_dict = {
"num_signals": num_signals,
"region": regions,
"group": groups,
"sampling_rate": sampling_rate,
}
return output_dict
def setup_units():
"""Set up the single unit data."""
# The number of tetrodes, probes, etc - any kind of grouping
num_groups = 8
# The region that each group belongs to
regions = ["CA1"] * num_groups
# A group number for each group, for example the tetrode number
groups = [1, 2, 3, 4, 9, 10, 11, 12]
output_dict = {
"num_groups": num_groups,
"region": regions,
"group": groups,
}
return output_dict
def setup_spatial():
"""Set up the spatial data."""
output_dict = {
"arena_size": "default",
}
return output_dict
def setup_loader():
"""
Set up the loader and keyword arguments for the loader.
See also
--------
simuran.loaders.loader_list.py
"""
# The type of loader to use, see simuran.loaders.loader_list.py for options
loader = "nc_loader"
output_dict = {
"loader": loader,
"loader_kwargs": nc_loader_kwargs,
}
return output_dict
load_params = setup_loader()
mapping = {
"signals": setup_signals(),
"units": setup_units(),
"spatial": setup_spatial(),
"loader": load_params["loader"],
"loader_kwargs": load_params["loader_kwargs"],
}
ph = simuran.ParamHandler(params=mapping)
ph.write(mapping_params_location)
# + colab={"base_uri": "https://localhost:8080/"} id="-p99fNtutp-v" outputId="8be58d64-7dbf-40b1-ff15-f240240e4ac9"
# Index the files if not already done
def add_mapping_to_df(input_df, **kwargs):
input_df["mapping"] = [os.path.basename(mapping_params_location)] * len(input_df)
return input_df
files_df = simuran.index_ephys_files(
path_dir,
loader_name="neurochat",
out_loc=index_location,
post_process_fn=add_mapping_to_df,
overwrite=False,
post_process_kwargs=None,
loader_kwargs=nc_loader_kwargs,
)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="A8a2i7Dayn2Y" outputId="d1dfc7cc-6cdf-4ae9-ffca-63396893860c"
# Inspect the files_df
files_df
# + id="93yGYbO1rXnh"
# Parse the recording information
rc = simuran.recording_container_from_df(
files_df,
base_dir=path_dir,
param_dir=temp_storage_location
)
# + colab={"base_uri": "https://localhost:8080/"} id="kQ5Unkvk-3oH" outputId="6ab28e87-505c-4c14-8252-469e7dfdb06e"
from skm_pyutils.py_log import get_default_log_loc
log_loc = get_default_log_loc("test.log")
print(log_loc)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["e73e1dbb445e44a3bf87a89501f457f1", "93e9da0706e0498798a224830b01d411", "7b6dc7b5488d4062b760f969741b0886", "375909c6042e4e46ac6c53352d25c139", "c0b7dbb6b8ab41b0982ad9916e7e64fe", "75ce3a74483e48999f2324b31b5f7e0a", "d702b14efc744b01a90fd4d7b3c8ef95", "38c041bb1f1f4b43af3720f13987405e"]} id="SXUqc_kXq3X_" outputId="5b70bd52-e759-43c3-adcd-fe399647b764"
# Analyse each recording in the container
ah = simuran.AnalysisHandler(handle_errors=True)
sm_figures = []
fn_kwargs = all_params
for r in rc:
for i in range(len(r.signals)):
r.signals[i].load()
fn_args = [r, path_dir, sm_figures]
ah.add_fn(powers, *fn_args, **fn_kwargs)
ah.run_all_fns(pbar="notebook")
# + colab={"base_uri": "https://localhost:8080/"} id="PZlxlY-n7o1x" outputId="b2b6bc06-08d4-4ab0-db94-1e9d90f43076"
# Save the analysis results
print(ah)
simuran.save_figures(sm_figures, temp_storage_location, verbose=True)
ah.save_results(os.path.join(temp_storage_location, "results.csv"))
# + id="9FQjafH2BQxb"
# To see errors, use this
# !ln -s /root/.skm_python /root/skm_python
|
lfp_atn_simuran/notebooks/ATNx_CA1_LFP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.13 ('pytorch')
# language: python
# name: python3
# ---
import torchvision
from model import *
from torch import nn
from torch.utils.data import DataLoader
from torch.utils.tensorboard import SummaryWriter
from torch.nn import Conv2d, MaxPool2d, Flatten, Linear, Sequential
# Loading Data
train_data = torchvision.datasets.CIFAR10("../P28-30_mock_project_practice/data/train", train=True, transform=torchvision.transforms.ToTensor(),download=True)
## training data has larger dataset which is used for train
test_data = torchvision.datasets.CIFAR10("../P28-30_mock_project_practice/data/test", train=False, transform=torchvision.transforms.ToTensor(),download=True)
## smaller dataset, only for test the trained result
# Data Length:
train_data_size = len(train_data)
test_data_size = len(test_data)
print("train data set length: {}".format(train_data_size))
print("test data set length: {}".format(test_data_size))
# Load data as batch:
batchsize = 64
train_dataloader = DataLoader(train_data, batch_size = batchsize)
test_dataloader = DataLoader(test_data, batch_size = batchsize)
# Load MODEL:
# +
class CIFAR10model(nn.Module):
def __init__(self):
super(CIFAR10model, self).__init__()
self.model1 = Sequential(
Conv2d(3, 32, 5, padding=2),
MaxPool2d(2),
Conv2d(32,32,5, padding=2),
MaxPool2d(2),
Conv2d(32,64,5, padding=2),
MaxPool2d(2),
Flatten(),
Linear(1024,64), # be careful, the last step is linearing 1024 data to 64 then linear to 10
Linear(64,10)
)
def forward(self, input):
output1 = self.model1(input)
return output1
CIFAR10Model = CIFAR10model()
if torch.cuda.is_available(): # if GPU is available -> use GPU for trainning
CIFAR10Model = CIFAR10Model.cuda()
# -
# Loss Function:
# +
loss_fn = nn.CrossEntropyLoss()
if torch.cuda.is_available(): # if GPU is available -> use GPU for trainning
loss_fn = loss_fn.cuda()
# -
# Optimizer:
Learning_Rate = 0.01
optimizer = torch.optim.SGD(CIFAR10Model.parameters(), lr = Learning_Rate)
# Trainning process:
total_train_step = 0
epoch = 5
writer = SummaryWriter("logs")
# +
for i in range(epoch):
print("No.{} round trainning".format(i+1))
# Train
for data in train_dataloader:
imgs, target = data
if torch.cuda.is_available(): # if GPU is available -> use GPU for trainning
imgs = imgs.cuda()
target = target.cuda()
output = CIFAR10Model(imgs)
loss = loss_fn(output, target)
optimizer.zero_grad()
loss.backward()
optimizer.step()
total_train_step += 1
if total_train_step % 500 ==0 :
print("trained {} times, loss: {}".format(total_train_step, loss.item()))
writer.add_scalar("train_loss", loss.item(), total_train_step)
# Test
with torch.no_grad(): # torch.no_grad() = This process no need to calculate gradient
total_accuracy = 0
total_test_step = 0
total_test_loss = 0
for data in test_dataloader:
imgs, targets = data
if torch.cuda.is_available(): # if GPU is available -> use GPU for trainning
imgs = imgs.cuda()
targets = targets.cuda()
outputs = CIFAR10Model(imgs)
loss = loss_fn(outputs, targets)
total_test_loss += loss.item()
total_test_step += 1
accuracy = (outputs.argmax(1) == targets).sum()/batchsize # Calculate the accuracy for a batch i.e. 64 images
total_accuracy += accuracy
if total_test_step % 50 ==0 :
print("test {} times, loss: {}".format(total_test_step, total_test_loss))
writer.add_scalar("test_loss", total_test_loss, total_test_step)
print("Accuracy: {}".format(total_accuracy/total_test_step))
writer.add_scalar("Accuracy", total_accuracy/total_test_step, total_test_step)
writer.close()
# -
# Save Result:
torch.save(CIFAR10Model, "CIFAR10Model_trained.pth")
print(len(targets))
print(len(target))
print(len(outputs))
print(len(output))
print(output)
|
P31-32_GPU_Train/GPU_Train_method_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Extract Parallel Texts from TED talks
#
# Derived / inspired by : [<NAME>'s GitHub](https://github.com/ajinkyakulkarni14/How-I-Extracted-TED-talks-for-parallel-Corpus-/blob/master/Ipython_notebook.ipynb).
import requests
from bs4 import BeautifulSoup
#import shutil
#import codecs
import os, glob
import csv
import time, random
def enlist_talk_names(url, dict_):
time.sleep( random.random()*5.0+5.0 )
r = requests.get(url)
print(" Got %d bytes from %s" % (len(r.text), url))
soup = BeautifulSoup(r.text, 'html.parser')
talks= soup.find_all("a", class_='')
for i in talks:
if i.attrs['href'].find('/talks/')==0 and dict_.get(i.attrs['href'])!=1:
dict_[i.attrs['href']]=1
return dict_
# +
all_talk_names={}
# Get all pages of talks (seems a bit abusive)
#for i in xrange(1,61):
# url='https://www.ted.com/talks?page=%d'%(i)
# all_talk_names=enlist_talk_names(url, all_talk_names)
# A specific seach term
#url='https://www.ted.com/talks?sort=newest&q=ai'
# Specific topics
url='https://www.ted.com/talks?sort=newest&topics[]=AI'
#url='https://www.ted.com/talks?sort=newest&topics[]=machine+learning'
#url='https://www.ted.com/talks?sort=newest&topics[]=mind'
#url='https://www.ted.com/talks?sort=newest&topics[]=mind&page=2'
all_talk_names=enlist_talk_names(url, all_talk_names)
len(all_talk_names)
# +
data_path = './data'
if not os.path.exists(data_path):
os.makedirs(data_path)
def extract_talk_languages(url, talk_name, language_list=['en', 'ko', 'ja']):
need_more_data=False
for lang in language_list:
talk_lang_file = os.path.join(data_path, talk_name+'-'+lang+'.csv')
if not os.path.isfile( talk_lang_file ) :
need_more_data=True
if not need_more_data:
print(" Data already retrieved for %s" % (url,))
return
time.sleep( random.random()*5.0+5.0 )
r = requests.get(url)
print(" Got %d bytes from %s" % (len(r.text), url))
if len(r.text)<1000: return # FAIL!
soup = BeautifulSoup(r.text, 'html.parser')
for i in soup.findAll('link'):
if i.get('href')!=None and i.attrs['href'].find('?language=')!=-1:
#print i.attrs['href']
lang=i.attrs['hreflang']
url_lang=i.attrs['href']
if not lang in language_list:
continue
talk_lang_file = os.path.join(data_path, talk_name+'-'+lang+'.csv')
if os.path.isfile( talk_lang_file ) :
continue
time.sleep( random.random()*5.0+5.0 )
r_lang = requests.get(url_lang)
print(" Lang[%s] : Got %d bytes" % (lang, len(r_lang.text), ))
if len(r.text)<1000: return # FAIL!
lang_soup = BeautifulSoup(r_lang.text, 'html.parser')
talk_data = []
for i in lang_soup.findAll('span',class_='talk-transcript__fragment'):
d = [ int( i.attrs['data-time'] ), i.text.replace('\n',' ') ]
talk_data.append(d)
with open(talk_lang_file, 'w') as csvfile:
writer = csv.writer(csvfile)
writer.writerow(['ts', 'txt'])
writer.writerows(talk_data)
if False:
# Now flatten out the talk_data into time_step order
talk_data_csv = [ ['ts']+language_list, ]
for ts in sorted(talk_data.keys(), key=int):
row = [ts] + [ talk_data[ts].get(lang, '') for lang in language_list]
talk_data_csv.append(row)
with open(os.path.join(data_path, talk_name+'.csv'), 'w') as csvfile:
writer = csv.writer(csvfile)
writer.writerows(talk_data_csv)
# -
for name in all_talk_names:
extract_talk_languages('https://www.ted.com'+name+'/transcript', name[7:])
#break
print("Finished extract_talk_languages for all_talk_names")
|
notebooks/work-in-progress/translation/1-TED-parallel-texts.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## 1. Load the data
# + slideshow={"slide_type": "fragment"}
# # %%bash
# pip install pandas
# + slideshow={"slide_type": "fragment"}
# Import statements
import pandas as pd
import os
# + slideshow={"slide_type": "fragment"}
pwd = "/Users/ayush/work/Zemanta-Data-Science-Summer-School"
os.chdir(pwd)
# -
df = pd.read_csv('data/labeledTrainData.tsv.zip', sep='\t')
# +
msk = np.random.rand(len(df)) < 0.5
train = df[msk]
# -
msk = np.random.rand(len(train)) < 0.8
train1 = train[msk]
test1 = train[~msk]
train1.to_csv("data/task_train.csv", index=False)
test1.to_csv("data/task_test_actual.csv", index=False)
print len(train1)
print len(test1)
print len(test)
test = test1[['id','review']]
test.to_csv("data/task_test.csv", index=False)
# + slideshow={"slide_type": "fragment"}
# Read the training data in pandas dataframe
# https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html
df_train = pd.read_csv("data/workshop_train.csv")
# -
from sklearn.model_selection import train_test_split
df_train, df_validate = train_test_split(df_train, test_size=0.2)
# + slideshow={"slide_type": "subslide"}
# Let's have a look at the dataset
df_train.head()
# + slideshow={"slide_type": "fragment"}
df_train.shape
# + slideshow={"slide_type": "fragment"}
df_train.columns
# + slideshow={"slide_type": "fragment"}
df_train = df_train[['sentiment','review']]
# + slideshow={"slide_type": "fragment"}
df_train.columns
# + slideshow={"slide_type": "subslide"}
df_train.head()
# + slideshow={"slide_type": "fragment"}
# Let's have a look at the reviews
df_train.review.values[1]
# + slideshow={"slide_type": "subslide"}
# Distribution of the examples
pd.value_counts(df_train.sentiment)
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Data Preprocessing & Visualization
# + slideshow={"slide_type": "fragment"}
# # %%bash
# pip install BeautifulSoup4
# pip install -U nltk
# + slideshow={"slide_type": "fragment"}
from bs4 import BeautifulSoup
# The package comes built-in with Python
import re
# http://www.nltk.org/install.html
import nltk
# + slideshow={"slide_type": "fragment"}
# Download text data sets, including stop words
# nltk.download()
# + slideshow={"slide_type": "subslide"}
from nltk.corpus import stopwords # Import the stop word list
stops = set(stopwords.words("english"))
# + slideshow={"slide_type": "fragment"}
# Pre-processing step
def text_to_words(raw_text):
# Function to convert a raw review to a string of words
# The input is a single string (a raw movie review), and
# the output is a single string (a preprocessed movie review)
# 1. Remove HTML
review_text = BeautifulSoup(raw_text).get_text()
# 2. Remove non-letters
# Find anything that is NOT a lowercase letter (a-z) or an upper case letter (A-Z), and replace it with a space
letters_only = re.sub("[^a-zA-Z]", " ", review_text)
# 3. Convert to lower case, split into individual words
words = letters_only.lower().split()
# 4. In Python, searching a set is much faster than searching
# a list, so convert the stop words to a set
# stops = set(stopwords.words("english"))
# There are many other things we could do to the data
# For example, Porter Stemming and Lemmatizing (both available in NLTK)
# would allow us to treat "messages", "message", and "messaging" as the same word
# which could certainly be useful.
# 5. Remove stop words
meaningful_words = [w for w in words if not w in stops]
#
# 6. Join the words back into one string separated by space,
# and return the result.
clear_text = " ".join(meaningful_words)
return clear_text
#validate_text_sentiment["Message"][i] = clear_text
#print i
#return( " ".join( meaningful_words ))
# + slideshow={"slide_type": "subslide"}
# Use the apply function in pandas to apply the function
# axis = 1 specifier means that the application is done at a row than a column level
df_train['cleaned_text'] = df_train.apply (lambda row: text_to_words(row.review), axis=1)
# + slideshow={"slide_type": "subslide"}
df_train.head()
# + slideshow={"slide_type": "fragment"}
df_train.to_csv("data/cleaned_train.csv",index=False)
# -
# ## Visualization
# + slideshow={"slide_type": "fragment"}
# # %%bash
# pip install wordcloud
# + slideshow={"slide_type": "fragment"}
# conda install -c conda-forge wordcloud
# http://amueller.github.io/word_cloud/index.html
from wordcloud import WordCloud,STOPWORDS
# Import plotting library
import matplotlib.pyplot as plt
# %matplotlib inline
# + slideshow={"slide_type": "fragment"}
def wordcloud_draw(data, color = 'black'):
words = ' '.join(data)
wordcloud = WordCloud(stopwords=STOPWORDS,
background_color=color,
width=2500,
height=2000
).generate(words)
plt.figure(1,figsize=(13, 13))
plt.imshow(wordcloud)
plt.axis('off')
plt.show()
# + slideshow={"slide_type": "subslide"}
print "Word cloud for positive words"
wordcloud_draw(df_train[(df_train.sentiment==1)]['cleaned_text'])
# + slideshow={"slide_type": "subslide"}
print "Word cloud for negative words"
wordcloud_draw(df_train[(df_train.sentiment==0)]['cleaned_text'], color = 'white')
# + [markdown] slideshow={"slide_type": "slide"}
# ## 3. Text Vectorization
# + slideshow={"slide_type": "subslide"}
# CountVectorizer is scikit-learn's bag of words tool
from sklearn.feature_extraction.text import CountVectorizer
# + slideshow={"slide_type": "fragment"}
# http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html
# analyzer = "word" specifies that the feature should be made of word
# To limit the size of the feature vectors, we should choose some maximum vocabulary size.
# Below, we use the 5000 most frequent words.
vectorizer = CountVectorizer(analyzer = "word", tokenizer = None, preprocessor = None, stop_words = None,
max_features = 5000)
# + slideshow={"slide_type": "fragment"}
# fit_transform() does two functions: First, it fits the model and learns the vocabulary;
# second, it transforms our training data into feature vectors.
# The input to fit_transform should be a list of strings.
train_data_features = vectorizer.fit_transform(df_train['cleaned_text'].values.astype('U'))
# + slideshow={"slide_type": "fragment"}
# The result of the above operation is a sparse matrix
train_data_features
# + slideshow={"slide_type": "fragment"}
print train_data_features.shape
# + slideshow={"slide_type": "fragment"}
# Numpy arrays are easy to work with, so convert the result to an array
train_data_features = train_data_features.toarray()
# + slideshow={"slide_type": "subslide"}
# Take a look at the words in the vocabulary
vocab = vectorizer.get_feature_names()
print vocab[0:10]
# + slideshow={"slide_type": "fragment"}
# We can also print the counts of each word in the vocabulary
import numpy as np
# Sum up the counts of each vocabulary word
dist = np.sum(train_data_features, axis=0)
print dist
# + slideshow={"slide_type": "fragment"}
# For each, print the vocabulary word and the number of times it appears in the training set
# for tag, count in zip(vocab, dist):
# print count, tag
# + slideshow={"slide_type": "fragment"}
print train_data_features[0]
# + slideshow={"slide_type": "notes"}
reverse_vocab = {v: k for k, v in vectorizer.vocabulary_.iteritems()}
# print reverse_vocab
# + slideshow={"slide_type": "notes"}
# # Convert vectors to text
# text = ''
# for i in range(0, len(vec)):
# if vec[i] != 0:
# text = text + ' ' + reverse_vocab.get(i)
# print text
# + slideshow={"slide_type": "notes"}
# vec = vectorizer.transform([df_train['cleaned_text'][0]]).toarray()
# # print df_train['cleaned_text'][0]
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Different techniques for Text Vectoriation
# -
# ### Bigram Model
# + slideshow={"slide_type": "fragment"}
bigram_vectorizer = CountVectorizer(ngram_range=(1, 2), token_pattern=r'\b\w+\b', min_df=1)
analyze = bigram_vectorizer.build_analyzer()
analyze('Bi-grams are cool!') == (['bi', 'grams', 'are', 'cool', 'bi grams', 'grams are', 'are cool'])
# + [markdown] slideshow={"slide_type": "subslide"}
# ### TfidfVectorizer
# + slideshow={"slide_type": "fragment"}
from sklearn.feature_extraction.text import TfidfTransformer
transformer = TfidfTransformer(smooth_idf=False)
# tfidf = transformer.fit_transform(df_train['cleaned_text'].values.astype('U'))
# + [markdown] slideshow={"slide_type": "slide"}
# # 5. Apply Machine Learning Algorithms
# + [markdown] slideshow={"slide_type": "subslide"}
# ## [Naive Bayes](http://scikit-learn.org/stable/modules/naive_bayes.html)
# + slideshow={"slide_type": "fragment"}
from sklearn.naive_bayes import GaussianNB
# + slideshow={"slide_type": "fragment"}
y_actual = df_train.sentiment.values
# + slideshow={"slide_type": "fragment"}
nb_model = GaussianNB().fit(train_data_features, y_actual)
# + slideshow={"slide_type": "subslide"}
# nb_model.predict(?)
# + slideshow={"slide_type": "fragment"}
df_test = pd.read_csv("data/workshop_train.csv")
# + slideshow={"slide_type": "fragment"}
df_test['cleaned_text'] = df_test.apply (lambda row: text_to_words(row.review), axis=1)
test_data_features = vectorizer.transform(df_test['cleaned_text'].values.astype('U'))
test_data_features = test_data_features.toarray()
# + slideshow={"slide_type": "fragment"}
predictions = nb_model.predict(test_data_features)
# + slideshow={"slide_type": "fragment"}
print predictions[0:10]
# + slideshow={"slide_type": "fragment"}
df_test['nb_prediction'] = predictions
# + slideshow={"slide_type": "fragment"}
# df_test.to_csv("results/test_prediction.csv", index=False)
# + [markdown] slideshow={"slide_type": "slide"}
# # Performance Metrics
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Accuracy
# + slideshow={"slide_type": "fragment"}
from sklearn.metrics import accuracy_score
y_pred = nb_model.predict(train_data_features)
y_true = df_train['sentiment']
# + slideshow={"slide_type": "fragment"}
print 'Accuracy of the algorithm is %f' %(accuracy_score(y_true, y_pred))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Precision
# + slideshow={"slide_type": "fragment"}
from sklearn.metrics import precision_score
print 'Precison of the algorithm is %f' %(precision_score(y_true, y_pred))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Recall
# + slideshow={"slide_type": "fragment"}
from sklearn.metrics import recall_score
print 'Recall of the algorithm is %f' %(recall_score(y_true, y_pred))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## F1-score
# + slideshow={"slide_type": "fragment"}
from sklearn.metrics import f1_score
print 'F1-score of the algorithm is %f' %(f1_score(y_true, y_pred))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Classification Report
# + slideshow={"slide_type": "fragment"}
from sklearn.metrics import classification_report
target_names = ['0', '1']
print(classification_report(y_true, y_pred, target_names=target_names))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Confusion Matrix
# + slideshow={"slide_type": "fragment"}
from sklearn.metrics import confusion_matrix
tn, fp, fn, tp = confusion_matrix(y_true, y_pred).ravel()
print tn, fp, fn, tp
# -
df_validate['cleaned_text'] = df_validate.apply (lambda row: text_to_words(row.review), axis=1)
validate_data_features = vectorizer.transform(df_validate['cleaned_text'].values.astype('U'))
validate_data_features = validate_data_features.toarray()
y_pred = nb_model.predict(validate_data_features)
y_true = df_validate.sentiment.values
print 'Accuracy of the algorithm is %f' %(accuracy_score(y_true, y_pred))
# + [markdown] slideshow={"slide_type": "slide"}
# ## 6. Cross-Validation & Hyperparameter Tuning
# + slideshow={"slide_type": "subslide"}
from sklearn.ensemble import RandomForestClassifier
# + slideshow={"slide_type": "fragment"}
# Random Forest classifier with 10 trees
forest = RandomForestClassifier(n_estimators = 10)
# + slideshow={"slide_type": "fragment"}
# 10-fold Cross-Validation using Grid Search
from sklearn.cross_validation import cross_val_score
from sklearn import metrics
# + slideshow={"slide_type": "fragment"}
# scoring = ['precision_macro', 'recall_macro', 'f1_macro']
param_grid = {'n_estimators': [10, 5]}
# + slideshow={"slide_type": "fragment"}
from sklearn.grid_search import GridSearchCV
grid_clf = GridSearchCV(forest, param_grid, cv=3, scoring='precision_macro')
# + slideshow={"slide_type": "fragment"}
grid_clf.fit(train_data_features, df_train['sentiment'].values)
# + slideshow={"slide_type": "fragment"}
# Get the best model
best_model = grid_clf.best_estimator_
# + slideshow={"slide_type": "fragment"}
# Get the value of the hyperparameters for best model
print grid_clf.best_params_
# -
grid_clf = GridSearchCV(forest, param_grid, cv=10, scoring='precision_macro', n_jobs = -1)
grid_clf.fit(train_data_features, df_train['sentiment'].values)
best_model = grid_clf.best_estimator_
# + slideshow={"slide_type": "fragment"}
predictions = best_model.predict(test_data_features)
# + slideshow={"slide_type": "fragment"}
df_test['rf_prediction'] = predictions
# + slideshow={"slide_type": "fragment"}
# df_test.to_csv("results/test_prediction.csv", index=False)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Model Ensembles
# + slideshow={"slide_type": "fragment"}
from sklearn import svm
from sklearn.ensemble import BaggingClassifier
# + slideshow={"slide_type": "fragment"}
# https://en.wikipedia.org/wiki/Radial_basis_function_kernel
svm_rbf = svm.SVC(C = 1, gamma = 0.001, kernel='rbf')
# + slideshow={"slide_type": "fragment"}
n_estimators = 10
svm_rbf_ensemble = BaggingClassifier(svm_rbf, max_samples=1.0 / n_estimators,
n_estimators=n_estimators)
# + slideshow={"slide_type": "fragment"}
model = svm_rbf_ensemble.fit(train_data_features, df_train["sentiment"])
# + slideshow={"slide_type": "fragment"}
predictions = model.predict(test_data_features)
# + slideshow={"slide_type": "fragment"}
df_test['svm_ensemble_prediction'] = predictions
# + slideshow={"slide_type": "fragment"}
# df_test.to_csv("results/test_prediction.csv", index=False)
|
sentiment_analysis_workshop_code.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=[]
import torch
print(torch.__version__)
# -
# Returns the default type of a tensor
torch.get_default_dtype()
# +
# Cannot set cuda tensor types to default dtype
# torch.set_default_dtype(torch.cuda.FloatTensor)
# Sets the default tensor type
torch.set_default_dtype(torch.half)
# -
# Create tensor from a given list
tensor = torch.Tensor([
[1,2,3],
[4,5,6]
])
# Checks variable is a tensor or not
torch.is_tensor(tensor)
# Returns number of elements
torch.numel(tensor)
# Create uninitialized tensor of a given size
torch.Tensor(2, 2)
# Creat tensor of a given size with random values
torch.rand(2, 3)
# Change the dtype of a tensor
torch.tensor([1,2,3]).type(torch.cuda.FloatTensor)
# Create a tensor of a given type
torch.cuda.DoubleTensor([1,2,3,4])
# Create a tensor of the given size filled with 1
ones_tensor = torch.ones(2, 3)
# Create a tensor of size of the passed tensor filled with zeros
torch.zeros_like(ones_tensor)
# Create and fill tensor with fill value
torch.full((2, 4), fill_value=5)
# Create 2d tensor of shape 5 x 5
# Ones are filled in the main diagonal
eye_tensor = torch.eye(5)
eye_tensor
# Returns the positions of non zero elements
torch.nonzero(eye_tensor)
# +
# Get
|
01-tensors-basic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_excel('tb.xlsx')
df
df['Balance'].sum()
df.info()
df['Acct_num_name'] = df['Account'].astype(str) + '_' + df['Name']
df
df.to_csv('new_tb.csv', index=False)
|
Accounting Internship/Loading TB/Load Trial Balance into Python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from fastai import * # Quick access to most common functionality
from fastai.text import * # Quick access to NLP functionality
# # Text example
# An example of creating a language model and then transfering to a classifier.
path = untar_data(URLs.IMDB)
path
# Open and view the independent and dependent variables:
df = pd.read_csv(path/'train.csv', header=None)
df.head()
classes = read_classes(path/'classes.txt')
classes[0], classes[1]
# Create a `DataBunch` for each of the language model and the classifier:
data_lm = TextLMDataBunch.from_csv(path)
data_clas = TextClasDataBunch.from_csv(path, vocab=data_lm.train_ds.vocab)
# [fast.ai](http://www.fast.ai/) has a pre-trained English model available that we can download.
datasets.download_wt103_model()
# We'll fine-tune the language model:
learn = RNNLearner.language_model(data_lm, pretrained_fnames=['lstm_wt103', 'itos_wt103'])
learn.unfreeze()
learn.fit(2, slice(1e-4,1e-2))
# Save our language model's encoder:
learn.save_encoder('enc')
# Fine tune it to create a classifier:
learn = RNNLearner.classifier(data_clas)
learn.load_encoder('enc')
learn.fit(3, 1e-3)
|
examples/text.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# %load_ext autoreload
# %autoreload 2
# +
import logging
logging.basicConfig(format="%(asctime)s [%(process)d] %(levelname)-8s "
"%(name)s,%(lineno)s\t%(message)s")
logging.getLogger().setLevel('INFO')
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook as tqdm
# -
# Read information to connect to the database and put it in environment variables
import os
with open('ENVVARS.txt') as f:
for line in f:
parts = line.split('=')
if len(parts) == 2:
os.environ[parts[0]] = parts[1].strip()
db_name = 'ticclat'
# db_name = 'ticclat_test'
os.environ['dbname'] = db_name
# +
from ticclat.ticclat_schema import Lexicon, Wordform, Anahash, Document, Corpus, WordformLink, WordformLinkSource, lexical_source_wordform, TextAttestation
from ticclat.dbutils import get_session, session_scope
Session = get_session(os.environ['user'], os.environ['password'], os.environ['dbname'])
# -
with session_scope(Session) as session:
print(session.get_bind())
# +
from sqlalchemy import Table, Column, BigInteger, Integer, Unicode
from sqlalchemy import select
from sqlalchemy import text
from sqlalchemy.sql import func, desc, and_
from sqlalchemy_views import CreateView
from ticclat.ticclat_schema import Base
class wordform_frequencies(Base):
__tablename__ = 'wordform_frequency'
wf_id = Column(BigInteger().with_variant(Integer, 'sqlite'), primary_key=True)
wordform_id = Column(BigInteger().with_variant(Integer, 'sqlite'))
wordform = Column(Unicode(255, convert_unicode=False), index=True)
frequency = Column(BigInteger())
with session_scope(Session) as session:
Base.metadata.create_all(session.get_bind())
# +
# %%time
from ticclat.sacoreutils import sql_insert
def iterate_results(result):
for row in tqdm(result.fetchall()):
yield {'wordform': row.wordform, 'wordform_id': row.wordform_id, 'frequency': row.freq}
with session_scope(Session) as session:
q = select([Wordform, func.sum(TextAttestation.frequency).label('freq')]).select_from(Wordform.__table__.join(TextAttestation)).group_by(Wordform.wordform_id)
#q = select([Wordform, Lexicon.lexicon_name]).select_from(lexical_source_wordform.join(Wordform).join(Lexicon))
r = session.execute(q)
sql_insert(session, wordform_frequencies, iterate_results(r))
# +
# %%time
from ticclat.dbutils import create_wf_frequencies_table
with session_scope(Session) as session:
create_word_frequencies_table(session)
# -
|
notebooks/create_wf_frequencies_table.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Zen Of Python
import this
# # Variables
# A name that is used to denote something or a value is called a variable. In python, variables can be declared and values can be assigned to it as follows,
x = 2
y = 5
xy = 'Hey'
print(x+y, xy)
# Multiple variables can be assigned with the same value.
x = y = 1
print(x,y)
# # Operators
# ## Arithmetic Operators
# | Symbol | Task Performed |
# |----|---|
# | + | Addition |
# | - | Subtraction |
# | / | division |
# | % | mod |
# | * | multiplication |
# | // | floor division |
# | ** | to the power of |
1+2
2-1
1*2
1/2 # if you are using python2 this operation will give result as '0'
15%10 # finding remainder when 15 is divided by 10
# Floor division is nothing but converting the result so obtained to the nearest integer.
2.8//2.0
# ## Relational Operators
# | Symbol | Task Performed |
# |----|---|
# | == | True, if it is equal |
# | != | True, if not equal to |
# | < | less than |
# | > | greater than |
# | <= | less than or equal to |
# | >= | greater than or equal to |
z = 1 # initializing value of z
z != 3 # check whether z is equal to 1 or not, if z is 1 then result is True else False.
z > 1
# ## Bitwise Operators
# | Symbol | Task Performed |
# |----|---|
# | & | Logical And |
# | l | Logical OR |
# | ^ | XOR |
# | ~ | Negate |
# | >> | Right shift |
# | << | Left shift |
a = 2 # binary representation of 2: 10
b = 3 # binary representarion of 3: 11
print(a & b)
print(bin(a&b))
5 >> 1
# 0000 0101 -> 5
#
# Shifting the digits by 1 to the right and zero padding
#
# 0000 0010 -> 2
5 << 1
# 0000 0101 -> 5
#
# Shifting the digits by 1 to the left and zero padding
#
# 0000 1010 -> 10
# # Built-in Functions
# Python comes loaded with pre-built functions
# ## Conversion from one system to another
# Conversion from hexadecimal to decimal is done by adding prefix **0x** to the hexadecimal value or vice versa by using built in **hex( )**, Octal to decimal by adding prefix **0** to the octal value or vice versa by using built in function **oct( )**.
hex(170)
0xAA
oct(8)
0o10
# **int( )** accepts two values when used for conversion, one is the value in a different number system and the other is its base. Note that input number in the different number system should be of string type.
print(int('0o10',8))
print(int('0xaa',16))
print(int('1010',2))
# **int( )** can also be used to get only the integer value of a float number or can be used to convert a number which is of type string to integer format. Similarly, the function **str( )** can be used to convert the integer back to string format
print(int(7.7)) # typecasting
print(int('7'))
# Also note that function **bin( )** is used for binary and **float( )** for decimal/float values. **chr( )** is used for converting ASCII to its alphabet equivalent, **ord( )** is used for the other way round.
chr(65)
ord('\t')
# ## Simplifying Arithmetic Operations
# **round( )** function rounds the input value to a specified number of places or to the nearest integer.
print(round(5.6231)) # by default round to the nearest integer
print(round(4.55892, 2))
# **complex( )** is used to define a complex number and **abs( )** outputs the absolute value of the same.
c =complex('5+2j')
print(c)
print(abs(c))
# **divmod(x,y)** outputs the quotient and the remainder in a __tuple__ in the format (quotient, remainder).
divmod(9,2)
# **isinstance( )** returns True, if the first argument is an instance of that class. Multiple classes can also be checked at once.
# +
print(isinstance(1, int))
print(isinstance(1.0,int))
# checks whether 0 is of the type int or float,
# if the first parameter is of any type mentioned in the second parameter
# then return true or else return false.
print(isinstance(0, (int,float) ))
# -
print(pow(3,3)) # pow(x,y) ==> means x raise to power y
# ## Accepting User Inputs
# **input( )** accepts input and stores it as a string. Hence, if the user inputs a integer, the code should convert the string to an integer and then proceed.
abc = int(input("Enter a Value:"))
# +
print(abc)
type(abc)
# -
# Note that **type( )** returns the format or the type of a variable or a number
|
01_Variables_Operators_DAC.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: AFEL analysis
# language: python
# name: venv
# ---
# # Knowledge Improvement and Post-test Activity
# We study here the relation that may exist between knowledge improvement and users' activities on didactalia and on the AFEL App after the knowledge post-test.
# # Imports & Constants
# ## Fuseki Endpoint : you might want to change it!
FUSEKI_DATASET = "evalafel" # Change according your own dataset name
SPARQL_ENDPOINT = "http://localhost:3030/%s/query" % FUSEKI_DATASET
# ## Imports, constant settings
from SPARQLWrapper import SPARQLWrapper, CSV
from rdflib import Literal
import numpy as np
import pandas as pd
from io import BytesIO
import gc
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import Normalizer
from scipy.stats import pearsonr
from sklearn.linear_model import LinearRegression
SPARQL_PREFIX = """
PREFIX xsd: <http://www.w3.org/2001/XMLSchema#>
PREFIX rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
PREFIX afl: <http://vocab.afel-project.eu/>
PREFIX extafl: <http://vocab.afel-project.eu/extension/>
PREFIX sch: <http://schema.org/>
"""
# Setting seaborn to have proper visualisation adapted to jupyter notebook
sns.set_context("notebook", font_scale=1.5, rc={"lines.linewidth": 2.5})
def sparql_to_dataframe(query, **kwargs):
sparql = SPARQLWrapper(SPARQL_ENDPOINT)
sparql.setQuery(query)
sparql.setReturnFormat(CSV)
res = sparql.query().convert()
try:
with BytesIO(res) as sio:
return pd.read_csv(sio, **kwargs)
finally:
del res
gc.collect()
# # Knowledge Increasment Extraction
def get_knowledge_cognitive_tests():
"""
Retrieve the results of the cognitives pre and post tests in geography and history
"""
query = SPARQL_PREFIX + """
SELECT ?userid ?questionID ?ratingValue
WHERE {
$action a sch:ChooseAction .
$action sch:endTime ?time .
$action sch:actionOption $rating .
$rating sch:ratingValue ?ratingValue .
$action sch:object $question .
$question sch:identifier ?questionID .
$question sch:isPartOf $questionnaire .
$rating sch:author $user .
$user afl:userName ?userid .
FILTER regex(?questionID, "^AFEL_2_KNOW") .
}
"""
return sparql_to_dataframe(query)
def process_know_stats(df_know_tests):
'''
from the results of test, compute two dataframes that provide min, mean& max score
of knowledge increasing for each user for both geopgraphy and history tests.
'''
def process_sub_test_df(sub_df):
# Compute relative id of questions
sub_df['questionID'] = sub_df.questionID.str.extract('AFEL_2_KNOW_(?:PRE|POST)_(?:GEO|HIST)_(.*)', expand=False)
# Compute pivot table to have questionID as columns
return sub_df.pivot(index='userid', columns='questionID', values='ratingValue')
# convert userid into simpler int userid
df_know_tests['userid'] = np.vectorize(int)(df_know_tests.userid.str.extract('project.afel\+(\d+)', expand=False))
# split test results into four independent dataframes
df_pre_geo = process_sub_test_df(df_know_tests.loc[df_know_tests.questionID.str.startswith('AFEL_2_KNOW_PRE_GEO'),:].copy())
df_pre_hist = process_sub_test_df(df_know_tests.loc[df_know_tests.questionID.str.startswith('AFEL_2_KNOW_PRE_HIST'),:].copy())
df_post_geo = process_sub_test_df(df_know_tests.loc[df_know_tests.questionID.str.startswith('AFEL_2_KNOW_POST_GEO'),:].copy())
df_post_hist = process_sub_test_df(df_know_tests.loc[df_know_tests.questionID.str.startswith('AFEL_2_KNOW_POST_HIST'),:].copy())
# Compute differences of scores
df_diff_geo = df_post_geo - df_pre_geo
df_diff_hist = df_post_hist - df_pre_hist
# Remove partial results
df_diff_geo.dropna(inplace=True)
df_diff_hist.dropna(inplace=True)
# Compute min, max and mean score for each user
df_know_geo_stats = df_diff_geo.apply(axis=1, func=lambda x: {'minKnow':x.min(), 'meanKnow':x.mean(), 'maxKnow':x.max(), 'stdKnow': x.std()},
result_type='expand')
df_know_hist_stats = df_diff_hist.apply(axis=1, func=lambda x: {'minKnow':x.min(), 'meanKnow':x.mean(), 'maxKnow':x.max(), 'stdKnow': x.std()},
result_type='expand')
# Assert that a user has not participated to both tests
assert len(set(df_know_geo_stats.index.values) & set(df_know_hist_stats.index.values)) == 0
return pd.concat([df_know_geo_stats, df_know_hist_stats])
df_know_stats = process_know_stats(get_knowledge_cognitive_tests())
# # Activities Extraction
def get_user_didactalia_activities():
query = SPARQL_PREFIX + """
SELECT ?userid ?location ?activityType ?artId ?actStartTime ?actEndTime
WHERE {
$activity a ?activityType .
$activity afl:user ?user .
$activity afl:artifact $artifact .
$activity afl:eventStartDate ?actStartTime .
$activity afl:eventEndDate ?actEndTime .
$activity sch:location ?location .
?user afl:userName ?userid .
$artifact afl:resourceID ?artId .
FILTER(?activityType IN (afl:ArtifactView, extafl:DidactaliaGamePlayed, extafl:RecommendedArtifactView, extafl:ScopeView)) .
FILTER(?actStartTime >= ?timePost) .
{
SELECT ?user (MAX(?timeTest) AS ?timePost)
WHERE {
$action a sch:ChooseAction .
$action sch:object $question .
$question a sch:Question.
$question sch:isPartOf $questionnaire .
$questionnaire a extafl:Questionnaire .
$action sch:agent $user .
$user a afl:User .
$action sch:endTime ?timeTest .
$questionnaire sch:identifier ?questionnaireId .
FILTER(REGEX(?questionnaireId, "AFEL_2_KNOW_(PRE|POST)_(HIST|GEO)")) .
}
GROUP BY ?user ?catQuest
}
}
"""
return sparql_to_dataframe(query, parse_dates=[3, 4])
df_activities = get_user_didactalia_activities()
# ### Convert usernames to ids
df_activities['userid'] = np.vectorize(int)(df_activities.userid.str.extract('project.afel\+(\d+)', expand=False))
# # Analyze number of activities and knowledge improvement
df_num_activities = df_activities.reset_index().loc[:, ['userid', 'index']]\
.groupby('userid').count()\
.rename(columns={'index':'numActivities'})
df_num_activities = pd.merge(df_num_activities, df_know_stats,
on='userid', validate='one_to_one')
# ### Distribution analysis
sns.distplot(df_num_activities.numActivities, hist=False, rug=True, color="r")
# ### Correlation analysis
g = sns.PairGrid(df_num_activities)
g = g.map(plt.scatter)
for depVar in ['minKnow', 'meanKnow', 'maxKnow', 'stdKnow']:
r, pval = pearsonr(df_num_activities[depVar], df_num_activities.numActivities)
indicator = '***' if pval < .05 else ''
print("%s: r = %.3f pval = %.3f %s" % (depVar, r, pval, indicator))
# # Analysis of number of activity of a certain type/location and knowledge improvement
df_activities['actFullType'] = \
np.vectorize(lambda x: 'DIDA_' if x == 'https://didactalia.net' else 'AFEL_')(df_activities.location)\
+ df_activities.activityType.str.split('/').str.get(-1)
df_details_activities = df_activities.reset_index().loc[:, ['userid', 'actFullType', 'index']]\
.groupby(['userid', 'actFullType']).count()\
.rename(columns={'index':'numActivities'})\
.reset_index()\
.pivot(index='userid', columns='actFullType', values='numActivities')\
.fillna(0)
df_details_activities = pd.merge(df_details_activities, df_know_stats, on='userid', validate='one_to_one')
# ### Correlation analysis
def correl_test(df, features):
for feature in features:
print("Correlation btwn %s and know" % feature)
for indic_know in ['meanKnow', 'minKnow', 'maxKnow', 'stdKnow']:
print(" - %s:" % indic_know)
res = pearsonr(df[indic_know], df[feature])
suff = '***' if res[1] < .05 else ''
print(" r = %.3f p-value = %.5f %s" % (res[0], res[1], suff))
df_details_activities.columns
correl_test(df_details_activities, ['AFEL_ArtifactView', 'AFEL_RecommendedArtifactView', 'AFEL_ScopeView',
'DIDA_ArtifactView', 'DIDA_DidactaliaGamePlayed'])
|
analysis/KnowledgeImprovement_and_Post-testActivity.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Initialisation Cell
from matplotlib import pyplot as plt
from IPython.display import display, HTML, Javascript
from math import *
import numpy as np
import numpy.testing as nt
# + [markdown] deletable=false editable=false nbgrader={"checksum": "e8bbe90c814ccb94902c0b68f412dace", "grade": false, "grade_id": "cell-7d967f1d8719468c", "locked": true, "schema_version": 1, "solution": false}
# # Optimisation II - Test 2 Deferred
#
#
# ## Instructions
#
# * You may only access https://moodle.ms.wits.ac.za/tests/ during the test and **NOT** normal moodle (i.e. https: //moodle.ms.wits.ac.za/moodle/ )
# * All machines are being logged and anyone caught logging into the regular Moodle listed above will be given **zero**.
# * Anyone caught trying to log into someone else's account will be given **zero**.
# * Anyone attempting to use brought in code, notes or any other materials will be given **zero**.
# * Read all the instructions carefully.
# * **DO NOT change the name of the notebook in any way!**
# * The test will run from 09:15 - 10:45 (this includes the extra time students). At 10:45 Moodle will no longer take submissions so ensure that you have uploaded your notebook by at least 11:55.
# * DO NOT rename the file. Simply answer all questions within the notebook and resubmit the file.
# * There are 95 marks available. There are 5 MCQ questions and 5 programming questions.
# + [markdown] deletable=false editable=false nbgrader={"checksum": "a0717fff2c107777fef6323bb2b71a7f", "grade": false, "grade_id": "cell-238e5ad89d42dd4d", "locked": true, "schema_version": 1, "solution": false}
# ## MCQ Questions
#
# If each of the MCQ questions there is only one correct answer. Once you have decided a answer in each question, uncomment out the response of your choosing from the correspond python function.
#
# There are 5 MCQ questions.
#
# ***
#
# ### Question 1
#
# Applying Newton's method on the quadratic function: $$f(\mathbf{x}) = \dfrac{1}{2}\mathbf{x}^TQ\mathbf{x} + \mathbf{x}^T \mathbf{b} + \mathbf{c},$$where $Q$ is a symmetric negative-definite matrix, will converge to the minimum in how many steps?
#
# * (a) 1
# * (b) Depends on the starting point
# * (c) Depends on the tolerance used
# * (d) Depends on the loops determined by the *for-loop*
# * (e) None of the above
# + deletable=false nbgrader={"checksum": "14fc8552f424a4b3a36c5247816a6fb4", "grade": false, "grade_id": "cell-f0ffb7bf7938c1ba", "locked": false, "schema_version": 1, "solution": true}
def question1():
"""
Uncomment your choice to the question
"""
#return 'a'
#return 'b'
#return 'c'
#return 'd'
#return 'e'
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "a36c50ce0b77d78a457cbc4eda961be3", "grade": true, "grade_id": "cell-5190bc15807307bd", "locked": true, "points": 0, "schema_version": 1, "solution": false}
assert(question1() in ['a', 'b', 'c', 'd', 'e'])
print('Function is returning a plausible response')
# + deletable=false editable=false nbgrader={"checksum": "71a1bc2bd3ad781684db6ed679600b65", "grade": true, "grade_id": "cell-73bc8f21efa81efd", "locked": true, "points": 3, "schema_version": 1, "solution": false}
# 3 Marks
# Hidden Evaluation of Answer
# + [markdown] deletable=false editable=false nbgrader={"checksum": "78f2e8770b7e5a823ef90dc3b52c0646", "grade": false, "grade_id": "cell-6ceae329b86b84ae", "locked": true, "schema_version": 1, "solution": false}
# ### Question 2
#
# Newton's method makes use of what when considering a search direction:
#
# * (a) No derivatives are used
# * (b) The first derivative is used
# * (c) Both the first and second derivatives are used
# * (d) The second derivative is used
# * (e) None of the above
# + deletable=false nbgrader={"checksum": "df238f1d997e52d9526a737891f3b26e", "grade": false, "grade_id": "cell-fad53ad658a04108", "locked": false, "schema_version": 1, "solution": true}
def question2():
"""
Uncomment your choice to the question
"""
#return 'a'
#return 'b'
#return 'c'
#return 'd'
#return 'e'
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "d9ad380896d38d6c7dcdb602b13ff67e", "grade": true, "grade_id": "cell-51e8d01b2a97ab11", "locked": true, "points": 0, "schema_version": 1, "solution": false}
assert(question2() in ['a', 'b', 'c', 'd', 'e'])
print('Function is returning a plausible response')
# + deletable=false editable=false nbgrader={"checksum": "4394f2ab5a1a4ae4e31ba338106ae2a6", "grade": true, "grade_id": "cell-c8e39f37cca5beba", "locked": true, "points": 3, "schema_version": 1, "solution": false}
# 3 Marks
# Hidden Evaluation of Answer
# + [markdown] deletable=false editable=false nbgrader={"checksum": "ea009167ade3cffbf206ec96bdeb77b1", "grade": false, "grade_id": "cell-fd42234117cd4902", "locked": true, "schema_version": 1, "solution": false}
# ### Question 3
#
# Given the Rosenbrock function:
# $$
# f(x)=100(x_2-x_1^2)^2+(1-x_1)^2.
# $$
# Applying Newton's method with a starting value of $\mathbf{x}^0= {0\choose 0}$ yields the next iterate $\mathbf{x}^1= {1\choose 0}$. What can you conclude about the performance of the algorithm:
#
# * (a) The algorithm is converging to a minimum
# * (b) Nothing, more iterations need to be performed in order for a patten to emerge
# * (c) Nothing, the patten of iterates is random
# * (d) The algorithm is diverging
# * (e) None of the above
# + deletable=false nbgrader={"checksum": "e70cb4642d1450a8ab67da8c703541bf", "grade": false, "grade_id": "cell-ec17e576119d50a1", "locked": false, "schema_version": 1, "solution": true}
def question3():
"""
Uncomment your choice to the question
"""
#return 'a'
#return 'b'
#return 'c'
#return 'd'
#return 'e'
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "1a9768527ba1c6c69f8d4ae2dcd8dccd", "grade": true, "grade_id": "cell-d9f98220c2b345ff", "locked": true, "points": 0, "schema_version": 1, "solution": false}
assert(question3() in ['a', 'b', 'c', 'd', 'e'])
print('Function is returning a plausible response')
# + deletable=false editable=false nbgrader={"checksum": "9dbd1b8cb1b0d0d6db1a9fc59352e94a", "grade": true, "grade_id": "cell-68f89d3eb95965db", "locked": true, "points": 3, "schema_version": 1, "solution": false}
# 3 Marks
# Hidden Evaluation of Answer
# + [markdown] deletable=false editable=false nbgrader={"checksum": "6575d0ccf6bd3d178eaa880a36e74a15", "grade": false, "grade_id": "cell-92ce152343e8dbe0", "locked": true, "schema_version": 1, "solution": false}
# ### Question 4
#
# Consider the Modified Newton Method. One major drawback of the method is:
#
# * (a) The method has poor convergence and is slow
# * (b) The method rarely works on non-quadratic functions
# * (c) The method is computational expensive due to computing the inverse Hessian per iterate
# * (d) The method can diverge even if $H^{k}$ is positive definite
# * (e) None of the above
# + deletable=false nbgrader={"checksum": "7ad6f1b676299307f9922d2ab9552e65", "grade": false, "grade_id": "cell-5276b192c0be54da", "locked": false, "schema_version": 1, "solution": true}
def question4():
"""
Uncomment your choice to the question
"""
#return 'a'
#return 'b'
#return 'c'
#return 'd'
#return 'e'
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "37b9c450a277f408b95f6cb3de8536a9", "grade": true, "grade_id": "cell-316ef23e7fc216f7", "locked": true, "points": 0, "schema_version": 1, "solution": false}
assert(question4() in ['a', 'b', 'c', 'd', 'e'])
print('Function is returning a plausible response')
# + deletable=false editable=false nbgrader={"checksum": "15a4f6b56b67b1bcea818c96428baff7", "grade": true, "grade_id": "cell-dd6f73c25a7dc1e4", "locked": true, "points": 3, "schema_version": 1, "solution": false}
# 3 Marks
# Hidden Evaluation of Answer
# + [markdown] deletable=false editable=false nbgrader={"checksum": "895f67545b4650ce0cc3ca23ecb6b184", "grade": false, "grade_id": "cell-f379eb1138166df9", "locked": true, "schema_version": 1, "solution": false}
# ### Question 5
#
# Random Walk and Downhill Simplex use which derivative(s) when computing a new iterate:
#
# * (a) First
# * (b) Second
# * (c) Second but First used in a line search
# * (d) None
# * (e) None of the above
# + deletable=false nbgrader={"checksum": "7d21dd6389d0ef1abeb61f165c8d4ddd", "grade": false, "grade_id": "cell-1561778c493f358f", "locked": false, "schema_version": 1, "solution": true}
def question5():
"""
Uncomment your choice to the question
"""
#return 'a'
#return 'b'
#return 'c'
#return 'd'
#return 'e'
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "5aac588d6c273bb56419dc5de9e8915f", "grade": true, "grade_id": "cell-a359c2e1235d8f2c", "locked": true, "points": 0, "schema_version": 1, "solution": false}
assert(question5() in ['a', 'b', 'c', 'd', 'e'])
print('Function is returning a plausible response')
# + deletable=false editable=false nbgrader={"checksum": "f54bc95926394ada670f5139b35febe6", "grade": true, "grade_id": "cell-f2a7af867c36da82", "locked": true, "points": 3, "schema_version": 1, "solution": false}
# 3 Marks
# Hidden Evaluation of Answer
# + [markdown] deletable=false editable=false nbgrader={"checksum": "0ecd85d9b4baf9cca0a5fafd4247bc67", "grade": false, "grade_id": "cell-5f92969ff18db909", "locked": true, "schema_version": 1, "solution": false}
# ## Programming Questions
#
# ### Question 1 [15 Marks]
#
# Converting the polar coordinate $(r, \theta)$ of a point to the Cartesian coordinates $(x, y)$ we have:
# \begin{eqnarray*}
# x = r\cos\theta,\\
# y = r\sin\theta,\\
# r = x^2 + y^2.
# \end{eqnarray*}
# Write a function `neighbourhood`, which takes as input a origin point `c` (i.e. ($x_0, y_0$), as a numpy array), `r` a radial distance and `n` scalar number of samples. The function should generate `n` samples in the neighbourhood around `c`. The samples should be generated randomly within the radius `r`.
# + deletable=false nbgrader={"checksum": "22d3a2993cb1969a091654225cefa6a8", "grade": false, "grade_id": "cell-a2b5fa6bc5b4d111", "locked": false, "schema_version": 1, "solution": true}
def neighbourhood(c, r, n):
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "5cdea93f813d86cfe331dd9122f2a52d", "grade": true, "grade_id": "cell-e763ce7265cca916", "locked": true, "points": 15, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# 15 Marks
import numpy.linalg as LA
c = [np.random.randint(10), np.random.randint(10)]
r = np.random.randint(10)
n = 2000
xtest, ytest = neighbourhood(c, r, n)
for i in range(n):
pt = np.array([xtest[i], ytest[i]])
dist = LA.norm(pt - c, 2)
if dist > r:
raise AssertionError('point exceeds maximum Euclidean distance')
print('Test case passed!!!')
# + [markdown] deletable=false editable=false nbgrader={"checksum": "9a6f5ede86c2cde940d92451a0b41759", "grade": false, "grade_id": "cell-a156b4eecb171c69", "locked": true, "schema_version": 1, "solution": false}
# ### Question 2 [15 Marks]
#
# Write a function `secant_minimiser`, which takes as inputs, the function `f`, its derivative `g`, initial guesses, `x0` and `x1`. It should also take in the scalar `tol`. The function should perform the secant method to determine the minima of `f` as well as the corresponding $x$ value associated with the minima. The third and final output should be the number of iterations required. You must use `abs(x1 - x0)` to control your tolerance.
#
# **Recall:** The updating formula for the *standard Secant Method for root finding* is:
# $$
# x_{n+1}=x_n-f(x_n)\dfrac{(x_n-x_{n-1})}{\left (f(x_n)-f(x_{n-1})
# \right)}. \nonumber
# $$
# + deletable=false nbgrader={"checksum": "259da871ee5594e3dd17f5df03194b09", "grade": false, "grade_id": "cell-584640c8cd53a6b4", "locked": false, "schema_version": 1, "solution": true}
def secant_min(f, g, x0, x1, tol):
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "8deb624d1de8749b354599f2633e6167", "grade": true, "grade_id": "cell-4c376330935755c5", "locked": true, "points": 2, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Unit test
# 2 Marks
f = lambda x: x**2 - 612
g = lambda x: 2*x
x0 = 10
x1 = 30
tol = 1e-5
secant_min(f, g, x0, x1, tol)
nt.assert_almost_equal(-612.0, secant_min(f, g, x0, x1, tol)[0])
print('Test case passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "574a1f7ff61cbf3b669686992ebf0283", "grade": true, "grade_id": "cell-554cff2e99a8a255", "locked": true, "points": 2, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Unit test
# 2 Marks
f = lambda x: x**2 - 612
g = lambda x: 2*x
x0 = 10
x1 = 30
tol = 1e-5
secant_min(f, g, x0, x1, tol)
nt.assert_almost_equal(0, secant_min(f, g, x0, x1, tol)[1])
print('Test case passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "792fd591323f61d6c37d213ff2a58318", "grade": true, "grade_id": "cell-716162a06c8107c2", "locked": true, "points": 2, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Unit test
# 2 Marks
f = lambda x: x**2 - 612
g = lambda x: 2*x
x0 = 10
x1 = 30
tol = 1e-5
secant_min(f, g, x0, x1, tol)
nt.assert_almost_equal(2, secant_min(f, g, x0, x1, tol)[2])
print('Test case passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "82bfb20fd8a4ac6fac2be681ed0ed95b", "grade": true, "grade_id": "cell-b59deab4e43649bc", "locked": true, "points": 5, "schema_version": 1, "solution": false}
# Do not delete this cell
# Hidden test
# 5 Marks
# + deletable=false editable=false nbgrader={"checksum": "f69ddc304860b9d1c481c7288e66049c", "grade": true, "grade_id": "cell-b31a54215541f045", "locked": true, "points": 4, "schema_version": 1, "solution": false}
# Do not delete this cell
# Hidden test
# 6 Marks
# + [markdown] deletable=false editable=false nbgrader={"checksum": "463aff21d4a8e650067ea888ea5607a5", "grade": false, "grade_id": "cell-7ce9a9b307742471", "locked": true, "schema_version": 1, "solution": false}
# ### Question 3 [20 Marks]
#
# Write a function `steepest_descent` which performs two variants of the method of steepest descent. The function should take in as inputs the function, `f`, its derivative `g`, an initial guess `x0`, a tolerance `tol`, a backtracking parameter `beta` and optional input `alpha`. If the optional input `alpha` is passed, then the function should perform the standard method of steepest descent with the given fixed step-size `alpha`. If no optional input is passed, then the function must perform the steepest descent with backtracking line search. You must use `beta` to find the acceptable `alpha` at each iterate. Both variants of the function should return the approximate minimum $x^*$, as well as the number of iterations required to achieve it. The logic to control your main while loops should be the `norm(g(x0), 2)`. In the case of backtracking, recall the while loop control:
# $$
# f(x - t\nabla f(x)) > f(x) - \dfrac{t}{2}\lVert \nabla f(x) \rVert^2.\nonumber
# $$
#
# **Note:** it would be advisable to limit your while limits in case of an infinite loop.
# + deletable=false nbgrader={"checksum": "46f480f67718981366e51dd1ee3b9e0a", "grade": false, "grade_id": "cell-d7f7eba75cff492e", "locked": false, "schema_version": 1, "solution": true}
def steepest_descent(f, g, x0, tol, beta, *alpha):
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "b52b6aecdb4909a87fa3fe7dc4af1268", "grade": true, "grade_id": "cell-a85b42d6c4291ddb", "locked": true, "points": 1, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Unit test
# 1 Marks
f1 = lambda x: 2*x[0]**2 + 3*x[1]**2
g1 = lambda x: np.array([4*x[0], 6*x[1]])
x0 = np.array([1, 1])
beta = 0.9
tol = 1e-7
alpha = 8/70
nt.assert_array_almost_equal([2.02246559e-08, 2.64493244e-15], steepest_descent(f1, g1, x0, tol, beta, alpha)[0])
print('Test case passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "c70bb7eb20071eb3c09dbe54dda75c2e", "grade": true, "grade_id": "cell-3947dd80c23ad8e2", "locked": true, "points": 1, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Unit test
# 1 Marks
f1 = lambda x: 2*x[0]**2 + 3*x[1]**2
g1 = lambda x: np.array([4*x[0], 6*x[1]])
x0 = np.array([1, 1])
beta = 0.9
tol = 1e-7
alpha = 8/70
assert(abs(steepest_descent(f1, g1, x0, tol, beta, alpha)[1] - 29) < 2)
print('Test case passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "543ed2ef6f78958352b489b9214e7033", "grade": true, "grade_id": "cell-aec54515523b6b0c", "locked": true, "points": 1, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Unit test
# 1 Marks
f1 = lambda x: 2*x[0]**2 + 3*x[1]**2
g1 = lambda x: np.array([4*x[0], 6*x[1]])
x0 = np.array([1, 1])
beta = 0.9
tol = 1e-7
alpha = 8/70
nt.assert_array_almost_equal([ 3.42518076e-09, -1.39964061e-10], steepest_descent(f1, g1, x0, tol, beta)[0])
print('Test case passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "bda5fa23b93a478b99951df789895375", "grade": true, "grade_id": "cell-570d58288c1b89cc", "locked": true, "points": 1, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Unit test
# 1 Marks
f1 = lambda x: 2*x[0]**2 + 3*x[1]**2
g1 = lambda x: np.array([4*x[0], 6*x[1]])
x0 = np.array([1, 1])
beta = 0.9
tol = 1e-7
alpha = 8/70
assert(abs(steepest_descent(f1, g1, x0, tol, beta)[1] - 11) < 2)
print('Test case passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "59b3fd53983607f92aae53bc42ef8b60", "grade": true, "grade_id": "cell-07ceaf2128b6a435", "locked": true, "points": 4, "schema_version": 1, "solution": false}
# Do not delete this cell
# Hidden test
# 4 Marks
# + deletable=false editable=false nbgrader={"checksum": "b57245ae1d936799671f11287dc37f1f", "grade": true, "grade_id": "cell-955e1736500943ad", "locked": true, "points": 4, "schema_version": 1, "solution": false}
# Do not delete this cell
# Hidden test
# 4 Marks
# + deletable=false editable=false nbgrader={"checksum": "c183b9647adb32e86ec62c1a622879b4", "grade": true, "grade_id": "cell-42ef9d86fc919c88", "locked": true, "points": 4, "schema_version": 1, "solution": false}
# Do not delete this cell
# Hidden test
# 4 Marks
# + deletable=false editable=false nbgrader={"checksum": "4e14435b1049ff60eeb495168ebf1379", "grade": true, "grade_id": "cell-2b5601dae5d9ab75", "locked": true, "points": 4, "schema_version": 1, "solution": false}
# Do not delete this cell
# Hidden test
# 4 Marks
# + [markdown] deletable=false editable=false nbgrader={"checksum": "bc4e46b7e34fecc47c3980c17ebd3a78", "grade": false, "grade_id": "cell-cba9b57ba02eee93", "locked": true, "schema_version": 1, "solution": false}
# ### Question 4 [15 Marks]
#
# Recall the algorithm for the DFP Method given below:
#
# 1. Set $k = 0, G^0 = I$ and compute $\mathbf{g}^k = g(\mathbf{x}^k)$.
# 2. Compute $\mathbf{d}^k$ from $\mathbf{d}^k = -G^k\mathbf{g}^k$.
# 3. Compute $\alpha = \alpha^k$ such that $f(\mathbf{x}^k + \mathbf{\alpha}^k\mathbf{d}^k),$ set $\mathbf{x}^{k+1} = \mathbf{x}^{k} + \alpha^k\mathbf{d}^k$.
# 4. Compute $\mathbf{g}^{k+1}$ such that $\mathbf{g}^{k+1} = g(\mathbf{x}^{k+1})$.
# 5. If $\lVert g^{k+1} \rVert \leq \epsilon$ ($\epsilon$ is a user supplied small number) then go to (9).
# 6. Compute $\delta^k$ and $\gamma^k$ such that $\delta^k = \mathbf{x}^{k+1} - \mathbf{x}^k$ and $\gamma = \mathbf{g}^{k+1} - \mathbf{g}^k$.
# 7. Compute $G^{k+1}$.
# 8. Set $k = k + 1$ and go to (2).
# 9. Set $\mathbf{x}^* = \mathbf{k+1}$, STOP.
#
# The updating rank-2 formula is given below:
# \begin{equation*}
# G^{k+1}=G^k+\dfrac{\delta^k \delta^{k^T}}{\delta^{k^T}\gamma^k} - \dfrac{G^k\gamma^k{\gamma^k}^TG^k}{{\gamma^k}^T G^k\gamma^k}
# \end{equation*}
#
# Finish the implementation seen below according to the above. The function should return an array of all approximations up to and including the final approximation, the number of iterations and the final $G^k$.
# + deletable=false nbgrader={"checksum": "c0e0004d82ef7a5743121af381f2f952", "grade": false, "grade_id": "cell-0806f3a1c2f5ad41", "locked": false, "schema_version": 1, "solution": true}
def dfp(f, g, x0, tol, beta):
import numpy.linalg as LA
G = np.eye(len(x0))
it = 0
app = np.array([x0])[0]
while LA.norm(g(x0)) > tol and it < 20:
d = -G.dot(g(x0))
alpha = 1.
while f(x0 + np.dot(alpha,d)) > f(x0) - (alpha/2)*(LA.norm(d)**2):
alpha = beta*alpha
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "63fb6a2784ec9b3dea040a54830abb6b", "grade": true, "grade_id": "cell-e86cfe738a38f123", "locked": true, "points": 2, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Unit test
# 2 Marks
f2 = lambda x: x[0] - x[1] + 2*x[0]**2 + 2*x[0]*x[1] + x[1]**2
g2 = lambda x: np.array([1 + 4*x[0] + 2*x[1], -1 + 2*x[0] + 2*x[1]])
x0 = np.array([0, 0])
beta = 0.9
tol = 1e-7
t1 = np.array([[ 0. , 0. ],
[-0.9 , 0.9 ],
[-0.93874205, 1.32616254],
[-0.99409587, 1.48281575],
[-0.99940883, 1.49828123],
[-0.99994089, 1.49982812],
[-0.99999409, 1.49998281],
[-0.99999941, 1.49999828],
[-0.99999994, 1.49999983],
[-1. , 1.5 ]])
t2 = 9
t3 = np.array([[ 0.50165299, -0.50078642],
[-0.50078642, 1.00037415]])
nt.assert_array_almost_equal(t1, dfp(f2, g2, x0, tol, beta)[0])
print('Test case passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "c57c078f9f299b501be60139d90e99c0", "grade": true, "grade_id": "cell-3d6a4b2a385d7dd7", "locked": true, "points": 2, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Unit test
# 2 Marks
nt.assert_almost_equal(t2, dfp(f2, g2, x0, tol, beta)[1])
print('Test case passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "bc1f1765fb5931bb3e6a48b7eaa2df7e", "grade": true, "grade_id": "cell-16352b9e48c7d127", "locked": true, "points": 2, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Unit test
# 2 Marks
nt.assert_array_almost_equal(t3, dfp(f2, g2, x0, tol, beta)[2])
print('Test case passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "0c1d50a308e34a82e3229448ca8e7962", "grade": true, "grade_id": "cell-e12ad5d8b017ea91", "locked": true, "points": 3, "schema_version": 1, "solution": false}
# Do not delete this cell
# Hidden test
# 3 Marks
# + deletable=false editable=false nbgrader={"checksum": "e03ef5e7b0a2103cff51b9863858e0c0", "grade": true, "grade_id": "cell-ff4164a0ac391296", "locked": true, "points": 3, "schema_version": 1, "solution": false}
# Do not delete this cell
# Hidden test
# 3 Marks
# + deletable=false editable=false nbgrader={"checksum": "f9f0dff0a0f5440f38a1dd9afd235e49", "grade": true, "grade_id": "cell-7287ae597fd7d60d", "locked": true, "points": 3, "schema_version": 1, "solution": false}
# Do not delete this cell
# Hidden test
# 3 Marks
# + [markdown] deletable=false editable=false nbgrader={"checksum": "86998bf995368ca9585b74ea0d9e6c5d", "grade": false, "grade_id": "cell-f5da6ac36bd016ac", "locked": true, "schema_version": 1, "solution": false}
# ### Question 5 [15 Marks]
#
# Write a function that takes in one input argument, $x$. If $x$ is an empty matrix, the function must return -1. If $x$ is a scalar it must return $0$. If $x$ is a vector it must return 1. Finally, if $x$ is none of these it must return 2. You may not use any builtin function such as `𝚒𝚜𝚜𝚌𝚊𝚕𝚊𝚛` etc.
#
# + deletable=false nbgrader={"checksum": "03a1b129c2caebc4c740483cb035dc92", "grade": false, "grade_id": "cell-9b713c7c9b0ae5c0", "locked": false, "schema_version": 1, "solution": true}
def classifier(x):
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "80f3125d249a161e2b06f6225bbe5ad2", "grade": true, "grade_id": "cell-b2db29d3b9799b48", "locked": true, "points": 1, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# 1 Mark
assert(2 == classifier(np.random.rand(2, 5)))
print('All Tests Passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "92d9e800fd7c8e1592aa5bfca62addd1", "grade": true, "grade_id": "cell-08c0ad32aef0fe62", "locked": true, "points": 1, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# 1 Mark
assert(1 == classifier(np.random.rand(1, 5)))
print('All Tests Passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "2c31a3e70e04bf94d8b5de82f09297ee", "grade": true, "grade_id": "cell-e6cacbe6fcfb8f67", "locked": true, "points": 2, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# 2 Mark
assert(-1 == classifier(np.array([])))
print('All Tests Passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "b66e4f9949e9f58790a1e3a2f5aaa7ef", "grade": true, "grade_id": "cell-ff00ef95e4c9439c", "locked": true, "points": 2, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# 2 Mark
assert(0 == classifier([1]))
print('All Tests Passed!!!')
# + deletable=false editable=false nbgrader={"checksum": "88b276430d5abbfd0e01161d9bf7feb5", "grade": true, "grade_id": "cell-8b529db97ede3e03", "locked": true, "points": 9, "schema_version": 1, "solution": false}
# Run this test cell to check your code
# Do not delete this cell
# Hidden cases
# 9 Mark
|
2018/release/Test3/test3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras.models import Model
from keras.layers import Dense
from keras.layers import Input, LSTM, GRU, Bidirectional, GlobalMaxPool1D, MaxPooling1D, Embedding
import numpy as np
from keras.layers import SimpleRNN # new!
import matplotlib.pyplot as plt
from keras.layers import SpatialDropout1D, Conv1D, GlobalMaxPooling1D # new!
from keras.callbacks import ModelCheckpoint
import os
from sklearn.metrics import roc_auc_score
import keras.backend as K
if len(K.tensorflow_backend._get_available_gpus()) > 0:
from keras.layers import CuDNNLSTM as LSTM
from keras.layers import CuDNNGRU as GRU
T = 8 ### T is the sequnces lenght
D = 2 ### D is the vector length dimensionality
M = 3 ### hidden layer size
X = np.random.randn(1, T, D)
X
# ### In the following feedforward we have 3 Dense nuerons so all the 8 (T= sequence length) gets multiplied to each of the nuerons and produces 24 parameters in the output.
def Feedforward():
input_ = Input(shape=(T, D))
rnn = Dense(M, activation='sigmoid')
x = rnn(input_)
model = Model(inputs=input_, outputs=x)
model.summary()
o= model.predict(X)
print("Feedforward output:", o)
print("Feedforward output.shape:", o.shape)
Feedforward()
# ### IN RNN if we have 3 (M=3 HIDDEN UNITS) when we pass in a sequence of 8 (T=8), it goes one by one into the network and when "return_state=True" alone, It will output the final value at T(7) (time stamp 7) one from each hidden nueron and the shape will be (1,3) and since return states is true is will also output h: the final hidden state for each of the hidden nueron (1,3) but if the return_sequnece parameter is True as well, then we will get an output vector at each time stamp from T(0) to T(7) (8 is the sequnce length) so T1 will have (1,3) from each hidden state all the way up to T(7) making the output shape to be (8,3)
def RNN1():
input_ = Input(shape=(T, D))
rnn = SimpleRNN(M,return_state=True )
x = rnn(input_)
model = Model(inputs=input_, outputs=x)
model.summary()
o, h= model.predict(X)
print("RNN o:", o)
print("RNN o.shape:", o.shape)
print("RNN h:", h)
print("RNN h:", h.shape)
RNN1()
def RNN2():
input_ = Input(shape=(T, D))
rnn = SimpleRNN(M)
x = rnn(input_)
model = Model(inputs=input_, outputs=x)
model.summary()
o= model.predict(X)
print("RNN o:", o)
print("RNN o.shape:", o.shape)
#print("RNN h:", h)
#print("RNN h:", h.shape)
RNN2()
def RNN3():
input_ = Input(shape=(T, D))
rnn = SimpleRNN(M,return_state=True, return_sequences=True )
x = rnn(input_)
model = Model(inputs=input_, outputs=x)
model.summary()
o, h= model.predict(X)
print("RNN3 o:", o)
print("RNN3 o.shape:", o.shape)
print("RNN3 h:", h)
print("RNN3 h:", h.shape)
RNN3()
def lstm1():
input_ = Input(shape=(T, D))
rnn = LSTM(M, return_state=True)
x = rnn(input_)
model = Model(inputs=input_, outputs=x)
model.summary()
o, h1, c1= model.predict(X)
print("o:", o)
print("o.shape:", o.shape)
print("h1:", h1)
print("h1:", h1.shape)
print("c1:", c1)
print("c1:", c1.shape)
lstm1()
def lstm2():
input_ = Input(shape=(T, D))
rnn = LSTM(M, return_state=True, return_sequences=True)
x = rnn(input_)
model = Model(inputs=input_, outputs=x)
model.summary()
o, h1, c1= model.predict(X)
print("o:", o)
print("o.shape:", o.shape)
print("h1:", h1)
print("h1:", h1.shape)
print("c1:", c1)
print("c1:", c1.shape)
lstm2()
def gru1():
input_ = Input(shape=(T, D))
rnn = GRU(M, return_state=True)
x = rnn(input_)
model = Model(inputs=input_, outputs=x)
o, h = model.predict(X)
print("o:", o)
print("h:", h)
gru1()
def gru2():
input_ = Input(shape=(T, D))
rnn = GRU(M, return_state=True, return_sequences=True)
x = rnn(input_)
model = Model(inputs=input_, outputs=x)
model.summary()
o, h = model.predict(X)
print("o:", o)
print("h:", h)
gru2()
def bidirectional():
input_ = Input(shape=(T, D))
rnn = Bidirectional(LSTM(M, return_state=True, return_sequences=True))
#rnn = Bidirectional(LSTM(M, return_state=True, return_sequences=True))
x = rnn(input_)
model = Model(inputs=input_, outputs=x)
o, h1, c1, h2, c2 = model.predict(X)
print("o:", o)
print("o.shape:", o.shape)
print("h1:", h1)
print("h1:", h1.shape)
print("c1:", c1)
print("c1:", c1.shape)
print("h2:", h2)
print("c2:", c2)
bidirectional()
def stacked():
input_ = Input(shape=(T, D))
rnn = LSTM(M, return_sequences=True)
x = rnn(input_)
rnn1 = LSTM(M,return_sequences=True)
y = rnn1(x)
pool=GlobalMaxPool1D()
z =pool(y)
k = Dense(2, activation="sigmoid")(z)
model3 = Model(inputs=input_, outputs=k)
model2 = Model(inputs=input_, outputs=y)
model = Model(inputs=input_, outputs=z)
o = model.predict(X)
print("o:", o)
print("o.shape:", o.shape)
o1 = model2.predict(X)
print("o1:", o1)
print("o.shape:", o1.shape)
o2 = model3.predict(X)
print("o2:", o2)
print("o2.shape:", o2.shape)
#print("h1:", h1)
#print("h1:", h1.shape)
#print("c1:", c1)
#print("c1:", c1.shape)
#print("h2:", h2)
#print("c2:", c2)
stacked()
# +
M=25
def conv1d():
input_ = Input(shape=(T, D))
rnn = LSTM(3, return_sequences=True)
x = rnn(input_)
conv = Conv1D(3, 2, strides=2, activation='relu')
z= conv(x)
pool=GlobalMaxPool1D()
k =pool(z)
m = Dense(1, activation="sigmoid")(k)
model_lstm = Model(inputs=input_, outputs=x)
o_lstm = model_lstm.predict(X)
print("o_lstm:", o_lstm)
print("o_lstm.shape:", o_lstm.shape)
model = Model(inputs=input_, outputs=z)
o = model.predict(X)
print("o:", o)
print("o.shape:", o.shape)
model1 = Model(inputs=input_, outputs=k)
o1 = model1.predict(X)
print("o1:", o1)
print("o1.shape:", o1.shape)
model2 = Model(inputs=input_, outputs=m)
o2 = model2.predict(X)
print("o2:", o2)
print("o2.shape:", o2.shape)
#print("h1:", h1)
#print("h1:", h1.shape)
#print("c1:", c1)
#print("c1:", c1.shape)
#print("h2:", h2)
#print("c2:", c2)
conv1d()
# +
def conv2d():
input_ = Input(shape=(T, D))
conv = Conv1D(8, 4, activation='relu')
z= conv(input_)
model = Model(inputs=input_, outputs=z)
o = model.predict(X)
print("o:", o)
print("o.shape:", o.shape)
conv2d()
# -
def conv2d():
input_ = Input(shape=(T, D))
conv = Conv1D(6, 2, strides=1, activation='relu')
z= conv(input_)
pool=MaxPooling1D(2)
k =pool(z)
conv1 = Conv1D(2,2, strides=1, activation='relu')
x= conv1(k)
model = Model(inputs=input_, outputs=z)
o = model.predict(X)
print("o:", o)
print("o.shape:", o.shape)
model_gmp = Model(inputs=input_, outputs=k)
o_gmp = model_gmp.predict(X)
print("o_gmp:", o_gmp)
print("o_gmp.shape:", o_gmp.shape)
model_conv1 = Model(inputs=input_, outputs=x)
o_conv1 = model_conv1.predict(X)
print("o_conv1:", o_conv1)
print("o_conv1.shape:", o_conv1.shape)
conv2d()
|
Deep-Learning-master 2/Shapes in Deep Learning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
# Extending PyVista {#extending_pyvista_example}
# =================
#
# A `pyvista.DataSet`{.interpreted-text role="class"}, such as
# `pyvista.PolyData`{.interpreted-text role="class"}, can be extended by
# users. For example, if the user wants to keep track of the location of
# the maximum point in the (1, 0, 1) direction on the mesh.
#
# There are two methods by which users can handle subclassing. One is
# directly managing the types objects. This may require checking types
# during filter operations.
#
# The second is automatic managing of types. Users can control whether
# user defined classes are nearly always used for particular types of
# DataSets.
#
# ::: {.note}
# ::: {.admonition-title}
# Note
# :::
#
# This is for advanced usage only. Automatic managing of types will not
# work in all situations, in particular when a builtin dataset is directly
# instantiated. See examples below.
# :::
#
# +
import numpy as np
import vtk
import pyvista
pyvista.set_plot_theme("document")
# -
# A user defined subclass of `pyvista.PolyData`{.interpreted-text
# role="class"}, `FooData` is defined. It includes a property to keep
# track of the point on the mesh that is furthest along in the (1, 0, 1)
# direction.
#
class FooData(pyvista.PolyData):
@property
def max_point(self):
"""Returns index of point that is furthest along (1, 0, 1) direction."""
return np.argmax(np.dot(self.points, (1.0, 0.0, 1.0)))
# Directly Managing Types
# =======================
#
# Now a `foo_sphere` object is created of type `FooData`. The index of the
# point and location of the point of interest can be obtained directly.
# The sphere has a radius of 0.5, so the maximum extent in the direction
# (1, 0, 1) is $0.5\sqrt{0.5}\approx0.354$
#
foo_sphere = FooData(pyvista.Sphere(theta_resolution=100, phi_resolution=100))
print("Original foo sphere:")
print(f"Type: {type(foo_sphere)}")
print(f"Maximum point index: {foo_sphere.max_point}")
print(f"Location of maximum point: {foo_sphere.points[foo_sphere.max_point, :]}")
# Using an inplace operation like
# `pyvista.DataSet.rotate_y`{.interpreted-text role="func"} does not
# affect the type of the object.
#
foo_sphere.rotate_y(90, inplace=True)
print("\nRotated foo sphere:")
print(f"Type: {type(foo_sphere)}")
print(f"Maximum point index: {foo_sphere.max_point}")
print(f"Location of maximum point: {foo_sphere.points[foo_sphere.max_point, :]}")
# However, filter operations can return different `DataSet` types
# including ones that differ from the original type. In this case, the
# `decimate <pyvista.PolyDataFilters.decimate>`{.interpreted-text
# role="func"} method returns a `pyvista.PolyData`{.interpreted-text
# role="class"} object.
#
print("\nDecimated foo sphere:")
decimated_foo_sphere = foo_sphere.decimate(0.5)
print(f"Type: {type(decimated_foo_sphere)}")
# It is now required to explicitly wrap the object into `FooData`.
#
decimated_foo_sphere = FooData(foo_sphere.decimate(0.5))
print(f"Type: {type(decimated_foo_sphere)}")
print(f"Maximum point index: {decimated_foo_sphere.max_point}")
print(f"Location of maximum point: {foo_sphere.points[foo_sphere.max_point, :]}")
# Automatically Managing Types
# ============================
#
# The default `pyvista.DataSet`{.interpreted-text role="class"} type can
# be set using `pyvista._wrappers`. In general, it is best to use this
# method when it is expected to primarily use the user defined class.
#
# In this example, all objects that would have been created as
# `pyvista.PolyData`{.interpreted-text role="class"} would now be created
# as a `FooData` object. Note, that the key is the underlying vtk object.
#
pyvista._wrappers['vtkPolyData'] = FooData
# It is no longer necessary to specifically wrap
# `pyvista.PolyData`{.interpreted-text role="class"} objects to obtain a
# `FooData` object.
#
foo_sphere = pyvista.Sphere(theta_resolution=100, phi_resolution=100)
print("Original foo sphere:")
print(f"Type: {type(foo_sphere)}")
print(f"Maximum point index: {foo_sphere.max_point}")
print(f"Location of maximum point: {foo_sphere.points[foo_sphere.max_point, :]}")
# Using an inplace operation like
# `rotate_y <pyvista.DataSet.rotate_y>`{.interpreted-text role="func"}
# does not affect the type of the object.
#
foo_sphere.rotate_y(90, inplace=True)
print("\nRotated foo sphere:")
print(f"Type: {type(foo_sphere)}")
print(f"Maximum point index: {foo_sphere.max_point}")
print(f"Location of maximum point: {foo_sphere.points[foo_sphere.max_point, :]}")
# Filter operations that return `pyvista.PolyData`{.interpreted-text
# role="class"} now return `FooData`
#
print("\nDecimated foo sphere:")
decimated_foo_sphere = foo_sphere.decimate(0.5)
print(f"Type: {type(decimated_foo_sphere)}")
print(f"Maximum point index: {decimated_foo_sphere.max_point}")
print(f"Location of maximum point: {foo_sphere.points[foo_sphere.max_point, :]}")
# Users can still create a native `pyvista.PolyData`{.interpreted-text
# role="class"} object, but using this method may incur unintended
# consequences. In this case, it is recommended to use the directly
# managing types method.
#
poly_object = pyvista.PolyData(vtk.vtkPolyData())
print(f"Type: {type(poly_object)}")
# catch error
try:
poly_object.rotate_y(90, inplace=True)
except TypeError:
print("This operation fails")
# Usage of `pyvista._wrappers` may require resetting the default value to
# avoid leaking the setting into cases where it is unused.
#
pyvista._wrappers['vtkPolyData'] = pyvista.PolyData
# For instances where a localized usage is preferred, a tear-down method
# is recommended. One example is a `try...finally` block.
#
try:
pyvista._wrappers['vtkPolyData'] = FooData
# some operation that sometimes raises an error
finally:
pyvista._wrappers['vtkPolyData'] = pyvista.PolyData
|
examples/99-advanced/extending-pyvista.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="oAuJOpPZ-ip6" colab_type="code" outputId="bb570bfa-6cbb-48c7-ce9c-2b2249486ef3" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/gdrive',force_remount=True)
# + id="G-Ar3fHP_NsG" colab_type="code" colab={}
# ! tar -xf '/content/gdrive/My Drive/Colab Notebooks/Selfie-dataset.tar.gz'
# + id="goVgHzju_c3s" colab_type="code" outputId="54c4627a-8161-43e6-8c7a-5bf42138d38b" colab={"base_uri": "https://localhost:8080/", "height": 34}
import tensorflow as tf
import keras
from keras.applications.resnet50 import ResNet50
from keras.preprocessing import image
from keras.applications.resnet50 import preprocess_input, decode_predictions
from keras.layers import GlobalAveragePooling2D, Dense, Dropout, Activation, Flatten
from keras.callbacks import ModelCheckpoint
from keras.models import Model
import numpy as np
import random
import cv2
from scipy import ndarray
import skimage as sk
from skimage import transform
from skimage import io
from skimage import util
from matplotlib import pyplot as plt
import matplotlib.pyplot as plt
import time
from keras.wrappers.scikit_learn import KerasClassifier
from keras.utils import np_utils
import matplotlib.gridspec as gridspec
from keras import backend as K
import cv2
from sklearn.preprocessing import LabelEncoder
import os
import time
import numpy as np
import pandas as pd
from sklearn.preprocessing import OneHotEncoder
# %matplotlib inline
import matplotlib.image as mpimg
# + id="GE_meS9A_k2e" colab_type="code" colab={}
data_frame = pd.read_csv("Selfie-dataset/selfie_dataset.txt",sep=" ")
data_frame.columns = ['image_name','popularity_score','partial_faces','is_female','baby','child','teenager','youth','middle_age','senior','white','black','asian','ovaal_face','round_face','heart_face','smiling','mouth_open','frowning','wearing_glasses','wearing_sunglasses','wearing_lipstick','tongue_out','duck_face','black_hair','blond_hair','brown_hair','red_hair','curly_hair','straight_hair','braid_hair','showing_cellphone','using_earphone','using_mirror','braces','wearing_hat','harsh_lighting','dim_lighting']
# + id="r2aYQ-Gp_l5s" colab_type="code" outputId="904475b1-5e19-4006-904d-9e1728051c41" colab={"base_uri": "https://localhost:8080/", "height": 119}
good_selfie_df = data_frame[['image_name','is_female','baby','child','teenager','youth','middle_age','senior']]
popularity_score = data_frame['popularity_score']
popularity_score.head()
# + id="UGn1BKar_vEa" colab_type="code" outputId="aedc2488-7088-4db9-a500-8383304b5e36" colab={"base_uri": "https://localhost:8080/", "height": 359}
good_selfie_df.head(10)
# + id="i_D-DVOr_0N1" colab_type="code" outputId="86555af5-aa96-41e6-e0d0-3e5beee4f039" colab={"base_uri": "https://localhost:8080/", "height": 359}
good_selfie_df.tail(10)
# + id="OiWxbkL7_1A1" colab_type="code" outputId="b05de3bb-78fd-4418-d6bd-15da43d7228d" colab={"base_uri": "https://localhost:8080/", "height": 776}
good_selfie_df.hist(figsize=(10,10))
# + id="qrtBZeDz_7zV" colab_type="code" outputId="a05a061e-202e-45fa-a136-3791cc98a01d" colab={"base_uri": "https://localhost:8080/", "height": 136}
good_selfie_df.loc[good_selfie_df['baby']== -1, 'baby'] = 0
good_selfie_df.loc[good_selfie_df['teenager']== -1, 'teenager'] = 0
good_selfie_df[good_selfie_df.is_female != 0]
good_selfie_df.loc[good_selfie_df['is_female']== -1, 'is_female'] = 0
good_selfie_df.loc[good_selfie_df['child']== -1, 'child'] = 0
good_selfie_df.loc[good_selfie_df['youth']== -1, 'youth'] = 0
good_selfie_df.loc[good_selfie_df['middle_age']== -1, 'middle_age'] = 0
good_selfie_df.loc[good_selfie_df['senior']== -1, 'senior'] = 0
good_selfie_df.head(10)
good_selfie_df = good_selfie_df[:5000]
popularity_score = popularity_score[:5000]
len(popularity_score)
# + id="q41sFRmiABwF" colab_type="code" colab={}
msk = np.random.rand(len(good_selfie_df)) < 0.85
train_df = good_selfie_df[msk]
train_popularity_score_df = popularity_score[msk]
test_df = good_selfie_df[~msk]
test_popularity_score_df = popularity_score[~msk]
# + id="UXQkmhT8ADWt" colab_type="code" outputId="c60a54e5-afe4-4d69-eb49-2020532c1b32" colab={"base_uri": "https://localhost:8080/", "height": 85}
print(len(train_df))
print(len(test_df))
print(len(train_popularity_score_df))
print(len(test_popularity_score_df))
# + id="fCq9M086AFgl" colab_type="code" outputId="a2a3e113-858d-4cf6-c776-a19608a224df" colab={"base_uri": "https://localhost:8080/", "height": 34}
imgs = []
y = []
for index,row in train_df.iterrows():
file_name = row['image_name']
image_path = 'Selfie-dataset/images/'+file_name+'.jpg'
image = mpimg.imread(image_path)
x = preprocess_input(image)
imgs.append(x)
imgs_data = np.array(imgs)
print(imgs_data.shape)
# + id="kuC0hYEdAJwB" colab_type="code" outputId="5942b4d4-d78a-4104-d317-0b4d91d64c90" colab={"base_uri": "https://localhost:8080/", "height": 51}
y_popularity_score_train = []
for row in train_popularity_score_df:
y_popularity_score_train.append(row)
y_np_popularity_score_train = np.array(y_popularity_score_train)
print(y_np_popularity_score_train.shape)
min_value = np.amin(y_np_popularity_score_train, axis=0)
max_value = np.amax(y_np_popularity_score_train, axis=0)
print(min_value,max_value)
# + id="V1sAKKWvCPIG" colab_type="code" outputId="85c71c6c-3296-42b9-9822-6c01d2e80408" colab={"base_uri": "https://localhost:8080/", "height": 51}
min_value = 1.656
max_value = 6.71399
fraction = (max_value-min_value)/3
print(fraction)
class_1 = min_value + fraction
class_2 = class_1 + fraction
class_3 = class_2 + fraction
print(class_1,class_2,class_3)
# + id="C86ipNEAjGZZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="b44a7e0c-f6fe-435f-fed8-bb72570b559e"
y_train= pd.qcut(y_np_popularity_score_train,3, labels=['Great_Selfie','Average_Selfie','Poor_Selfie'])
print(y_train)
# + id="LevRlgn6C7jC" colab_type="code" outputId="039ec2b8-b2f3-46f8-be66-610c2a348f2f" colab={"base_uri": "https://localhost:8080/", "height": 54}
selfie_state = ['Great_Selfie','Average_Selfie','Poor_Selfie']
y_train = []
for value in y_np_popularity_score_train:
if value<=class_1:
state = selfie_state[0]
elif value>class_1 and value<=class_2:
state = selfie_state[1]
else:
state = selfie_state[2]
y_train.append(state)
print(y_train[0:100])
# + id="clDR3z5CAOo2" colab_type="code" outputId="e7ec958e-a168-458f-89d2-dfc5f7a15af9" colab={"base_uri": "https://localhost:8080/", "height": 34}
imgs = []
for index, row in test_df.iterrows():
file_name = row['image_name']
image_path = 'Selfie-dataset/images/'+file_name+'.jpg'
image = mpimg.imread(image_path)
x = preprocess_input(image)
imgs.append(x)
imgs_test = np.array(imgs)
print(imgs_test.shape)
# + id="ZoL_88niFCMn" colab_type="code" outputId="7f428388-8485-4525-8ec3-9654cf7fbb3e" colab={"base_uri": "https://localhost:8080/", "height": 34}
y_popularity_score_test = []
for row in test_popularity_score_df:
y_popularity_score_test.append(row)
y_np_popularity_score_test = np.array(y_popularity_score_test)
y_np_popularity_score_test = np.array(y_np_popularity_score_test)
print(y_np_popularity_score_test.shape)
# + id="7SNUaFJYksH7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="937b3b6e-0e1a-4569-d165-87d7adf0850c"
y_test= pd.qcut(y_np_popularity_score_test,3, labels=['Great_Selfie','Average_Selfie','Poor_Selfie'])
print(y_test)
# + id="OKDpQywrp9ci" colab_type="code" colab={}
selfie_state = ['Great_Selfie','Average_Selfie','Poor_Selfie']
# + id="Xhkw07QfAjHr" colab_type="code" outputId="81e93bd3-fa99-4336-a959-de59885d866d" colab={"base_uri": "https://localhost:8080/", "height": 34}
selfie_state = ['Great_Selfie','Average_Selfie','Poor_Selfie']
y_test = []
for value in y_np_popularity_score_test:
if value<=class_1:
state = selfie_state[0]
elif value>class_1 and value<=class_2:
state = selfie_state[1]
else:
state = selfie_state[2]
y_test.append(state)
print(len(y_test))
# + id="LNQjmel-NaB6" colab_type="code" outputId="90480b27-fa0e-4a20-a237-129d1169a0dd" colab={"base_uri": "https://localhost:8080/", "height": 34}
# encode class values as integers
encoder = LabelEncoder()
encoder.fit(y_train)
encoded_Y = encoder.transform(y_train)
y_train = np_utils.to_categorical(encoded_Y)
print(y_train[100])
# + id="GqwVnPbHOPXZ" colab_type="code" outputId="46bad2cb-148e-4124-a59a-e4982ddc4f99" colab={"base_uri": "https://localhost:8080/", "height": 34}
# encode class values as integers
encoder = LabelEncoder()
encoder.fit(y_test)
encoded_Y = encoder.transform(y_test)
y_test = np_utils.to_categorical(encoded_Y)
print(y_test[1])
# + id="Nb9Q4zKJF14O" colab_type="code" outputId="9b42b41b-8a27-46de-f88c-cb185997a328" colab={"base_uri": "https://localhost:8080/", "height": 190}
onehotencoder = OneHotEncoder()
print(y_train[0])
y_train_enc = onehotencoder.fit_transform(y_train.reshape(-1,1)).toarray()
y_test_enc = onehotencoder.fit_transform(y_test.reshape(-1,1)).toarray()
# + id="gYp8a6qJGa3w" colab_type="code" outputId="a9434847-b65a-4db1-c4b3-61193024bdcb" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(y_train[100],y_train_enc[100],y_test[87],y_test_enc[87])
# + id="JPMOui8fGho4" colab_type="code" colab={}
def create_model():
num_output_unit = 3
model = ResNet50(weights='imagenet',include_top=False)
#model.summary()
last_layer = model.output
# add a global spatial average pooling layer
x = GlobalAveragePooling2D()(last_layer)
# add fully-connected & dropout layers
x = Dense(512, activation='relu',name='fc-1')(x)
x = Dropout(0.5)(x)
x = Dense(256, activation='relu',name='fc-2')(x)
x = Dropout(0.5)(x)
# a softmax layer for 4 classes
out_layer = Dense(num_output_unit, activation='softmax',name='output_layer')(x)
# this is the model we will train
custom_resnet_model2 = Model(inputs=model.input, outputs=out_layer)
#custom_resnet_model2.summary()
for layer in custom_resnet_model2.layers[:-6]:
layer.trainable = False
#custom_resnet_model2.layers[-1].trainable
return custom_resnet_model2
# + id="YpJ-9-VELpgC" colab_type="code" colab={}
def train():
custom_resnet_model2 = create_model()
custom_resnet_model2.compile(loss='categorical_crossentropy',optimizer='adam',metrics=['accuracy'])
ckpt_model = 'weights.best_h2.hdf5'
checkpoint = ModelCheckpoint(ckpt_model,
monitor='val_acc',
verbose=1,
save_best_only=True,
mode='max')
callbacks_list = [checkpoint]
# fit the model
t=time.time()
hist = custom_resnet_model2.fit(imgs_data,y_train,batch_size=32, epochs=12, verbose=1,callbacks=callbacks_list,validation_data=(imgs_test,y_test))
print("Training time: %s" % (t - time.time()))
(loss, accuracy) = custom_resnet_model2.evaluate(imgs_test,y_test, batch_size=10, verbose=1)
print("[INFO] loss={:.4f}, accuracy: {:.4f}%".format(loss,accuracy * 100))
# + id="QqQ8lPnnMjaV" colab_type="code" outputId="d50a9a2d-7e63-4b53-825b-d69aa12d46ce" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(y_train.shape)
#train()
# + id="skWMWPTBMa1G" colab_type="code" colab={}
def test_model_250():
# create the model
model = create_model()
# load model weights
model.load_weights('weights.best_h2.hdf5')
# predict values
predicted_values = model.predict(imgs_test[:250])
#print(predicted_values)
for values in predicted_values:
ind = np.argmax(values)
arr = [0,0,0]
arr[ind] = 1
print(arr)
# + id="KMnWbWEbpt0G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 4338} outputId="72d8ad78-991a-46c3-f56f-184643df9d32"
print(test_model_250())
# + id="kHGYSi8tOr8t" colab_type="code" outputId="971b100e-eec3-435a-a3ea-0424fca4512a" colab={"base_uri": "https://localhost:8080/", "height": 955}
train()
# + id="NivWtYJGOtCH" colab_type="code" outputId="e07077b6-8f8b-4449-a8bb-f8938ee061f1" colab={"base_uri": "https://localhost:8080/", "height": 88}
print(test_model('/content/gdrive/My Drive/Colab Notebooks/bad sel.jpg'))
# + id="N2urUy1GWLeb" colab_type="code" outputId="2e1a126b-a219-4056-a66e-c50be6be990b" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(imgs_test[1].shape)
# + id="a-OScJPtafW5" colab_type="code" colab={}
|
Selfie/popularity_score_qcut.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
total_minimax_games = 100
minimax_wins = 13
combi_minimax_wins = 87
total_rollouts_games = 117
rollouts_wins = 33
combi_rollouts_wins = 75
total_smart_games = 103
smart_wins = 2
combi_smart_wins = 101
total_probability_games = 92
probability_wins = 4
combi_probability_wins = 87
labels = ["minimax", "rollouts", "basic", "probability"]
data = [[combi_minimax_wins / total_minimax_games, combi_rollouts_wins / total_rollouts_games, combi_smart_wins/total_smart_games, combi_probability_wins/total_probability_games],
[minimax_wins / total_minimax_games, rollouts_wins / total_rollouts_games, smart_wins/total_smart_games, probability_wins/total_probability_games]]
# +
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
X = np.arange(len(labels)) + 0.2
ax.bar(X + 0.00, data[0], width = 0.4, label="Combi")
ax.bar(X + 0.4, data[1], width = 0.4, label="Client")
plt.tick_params(
axis='x',
which='both',
bottom=False,
top=False,
labelbottom=True)
ax.set_xticks(np.arange(len(labels))+0.4)
ax.set_xticklabels(labels)
ax.set_ylabel("Gewinnrate")
ax.set_xlabel("Client")
ax.legend()
# -
fig.savefig("chart_internal_testing.svg")
|
scripts/chart_internal.ipynb
|