code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] hide_input=false
# # Creating your own dataset from Google Images
#
# *by: <NAME> and <NAME>. Inspired by [<NAME>](https://www.pyimagesearch.com/2017/12/04/how-to-create-a-deep-learning-dataset-using-google-images/)*
# + [markdown] hide_input=true
# In this tutorial we will see how to easily create an image dataset through Google Images. **Note**: You will have to repeat these steps for any new category you want to Google (e.g once for dogs and once for cats).
# -
# ## Get a list of URLs
# ### Search and scroll
# Go to [Google Images](http://images.google.com) and search for the images you are interested in. The more specific you are in your Google Search, the better the results and the less manual pruning you will have to do.
#
# Scroll down until you see a button that says 'Show more results'. All the images you scrolled past are now available to download. To get more, click on the button. Then continue scrolling until you cannot scroll anymore. The maximum number of images Google Images shows is 700.
# ### Download into file
# Now you must run some Javascript code in your browser which will save the URLs of all the images you want for you dataset.
#
# Press <kbd>Ctrl</kbd><kbd>Shift</kbd><kbd>J</kbd> in Windows/Linux and <kbd>Cmd</kbd><kbd>Opt</kbd><kbd>J</kbd> in Mac, and a small window the javascript 'Console' will appear. That is where you will paste the JavaScript commands.
#
# You will need to get the urls of each of the images. You can do this by running the following commands:
#
# ```javascript
# urls = Array.from(document.querySelectorAll('.rg_di .rg_meta')).map(el=>JSON.parse(el.textContent).ou);
# window.open('data:text/csv;charset=utf-8,' + escape(urls.join('\n')));
# ```
# ### Create directory and upload urls file into your server
# + hide_input=false
from fastai import *
from fastai.vision import *
# -
# Choose an appropriate name for your labeled images. You can run these steps multiple times to grab different labels.
folder = 'black'
file = 'urls_black.txt'
folder = 'teddys'
file = 'urls_teddys.txt'
folder = 'grizzly'
file = 'urls_grizzly.txt'
# You will need to run this line once per each category.
path = Path('data/bears')
dest = path/folder
dest.mkdir(parents=True, exist_ok=True)
# Finally, upload your urls file. You just need to press 'Upload' in your working directory and select your file, then click 'Upload' for each of the displayed files.
#
# 
# ## Download images
# Now you will need to download you images from their respective urls.
#
# fast.ai has a function that allows you to do just that. You just have to specify the urls filename and the destination folder and this function will download and save all images than can be opened. If they have some problem in being opened, they will not be saved.
#
# Let's download our images! Notice you can choose a maximum number of images to be downloaded. In this case we will not download all the urls.
#
# You will need to run this line once for every category.
download_images(path/file, dest, max_pics=200)
# Good! Let's take a look at some of our pictures then.
classes = ['teddys','grizzly','black']
for c in classes:
print(c)
verify_images(path/c, delete=True, max_workers=8)
# ## View data
np.random.seed(42)
data = ImageDataBunch.from_folder(path, train=".", valid_pct=0.2,
ds_tfms=get_transforms(), size=224, num_workers=4).normalize(imagenet_stats)
data.classes
data.show_batch(rows=3, figsize=(7,8))
data.classes, data.c, len(data.train_ds), len(data.valid_ds)
# ## Train model
learn = create_cnn(data, models.resnet34, metrics=error_rate)
learn.fit_one_cycle(4)
learn.save('stage-1')
learn.unfreeze()
learn.lr_find()
learn.recorder.plot()
learn.fit_one_cycle(2, max_lr=slice(1e-5,1e-4))
learn.save('stage-2')
# ## Interpretation
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_confusion_matrix()
interp.plot_top_losses(2)
|
nbs/dl1/lesson2-download.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from collections import defaultdict
import pandas as pd
import pprint
# +
g = defaultdict(list)
marry = pd.read_csv('data/marriages.csv')
kids = pd.read_csv('data/children.csv')
chars = pd.read_csv('data/characters.csv')
# -
for i, row in marry.iterrows():
if pd.isna(row.MarryOptions):
continue
options = row.MarryOptions.split('-')
for o in options:
g[row.Character].append(o)
g[o].append(row.Character)
for i, row in kids.iterrows():
g[row.Child].append(row.Main)
g[row.Main].append(row.Child)
for i, row in chars.iterrows():
options = row.OptionsList.split('-')
for o in options:
g[row.Name].append(o)
def find_path(graph, start, end, path=[]):
path = path + [start]
if start == end:
return path
if not graph.get(start, None):
return None
for node in graph[start]:
if node not in path:
newpath = find_path(graph, node, end, path)
if newpath: return newpath
return None
def find_all_paths(graph, start, end, path=[]):
path = path + [start]
if start == end:
return [path]
if not graph.get(start, None):
return []
paths = []
for node in graph[start]:
if node not in path:
newpaths = find_all_paths(graph, node, end, path)
for newpath in newpaths:
paths.append(newpath)
return paths
def find_shortest_path(graph, start, end, path=[]):
path = path + [start]
if start == end:
return path
if not graph.get(start, None):
return None
shortest = None
for node in graph[start]:
if node not in path:
newpath = find_shortest_path(graph, node, end, path)
if newpath:
if not shortest or len(newpath) < len(shortest):
shortest = newpath
return shortest
find_path(g, 'Lucina', 'Pegasus Knight')
find_shortest_path(g, 'Lucina', 'Pegasus Knight')
find_all_paths(g, 'Chrom', 'Pegasus Knight')
pprint.pprint(g)
|
graphing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# hide
# all_tutorial
! [ -e /content ] && pip install -Uqq mrl-pypi # upgrade mrl on colab
# # Small Molecule Property Optimization - Penalized LogP
#
# >Small molecule penalized logP optimization
# ## Penalized LogP Optimization
#
# This notebook shows how to optimize a generative model with respect to [Penalized LogP](http://proceedings.mlr.press/v70/kusner17a) score. This is a standard benchmark in many generative design papers
# ## Performance Notes
#
# The workflow in this notebook is more CPU-constrained than GPU-constrained due to the need to evaluate samples on CPU. If you have a multi-core machine, it is recommended that you uncomment and run the `set_global_pool` cells in the notebook. This will trigger the use of multiprocessing, which will result in 2-4x speedups.
#
# This notebook may run slow on Collab due to CPU limitations.
#
# If running on Collab, remember to change the runtime to GPU
# +
import sys
sys.path.append('..')
from mrl.imports import *
from mrl.core import *
from mrl.chem import *
from mrl.templates.all import *
from mrl.torch_imports import *
from mrl.torch_core import *
from mrl.layers import *
from mrl.dataloaders import *
from mrl.g_models.all import *
from mrl.vocab import *
from mrl.policy_gradient import *
from mrl.train.all import *
from mrl.model_zoo import *
from mrl.combichem import *
# -
# ## Agent
#
# Here we create the model we want to optimize. We will use the `LSTM_LM_Small_ZINC_NC` - a LSTM-based language model pretrained on part of the ZINC database without chirality. Training without chirality prevents a form of mode collapse where the model converges to generating different isomers of the same compound.
#
# Whether or not to use a model trained with or without chirality depends on the reward function you are trying to optimize. You should use a model with chirality if your reward function handles chirality in a meaningful way. Specically this means your reward function should give different scores to different isomers. This difference should relate to a real aspect of the propety predicted (ie affinity of different isomers) rather than being a spurious feature learned by the model (this happens surprisingly often).
#
# Penalized LogP score isn't impacted by chirality, so using a non-chiral model makes sense. To be technical, Penalized LogP includes a SA score component which is influenced by the number of stereocenters in a molecule, but this does not result in different isomers getting different scores.
agent = LSTM_LM_Small_ZINC_NC(drop_scale=0.3, opt_kwargs={'lr':5e-5})
# ## Template
#
# Here we create our template.
#
# We set the following hard filters:
# - `ValidityFilter`: screens for vaid compounds
# - `SingleCompoundFilter`: screens for single compounds
#
# We set the following soft filters:
# - `PenalizedLogPFilter`: evaluates the Penalized LogP score of a compound. By passing `score=PassThroughScore()`, this filter simply returns the Penalized LogP score
# +
template = Template([ValidityFilter(),
SingleCompoundFilter(),
],
[PenalizedLogPFilter(None, None, score=PassThroughScore())],
fail_score=-1., log=False)
template_cb = TemplateCallback(template, prefilter=True)
# -
# ## Reward
#
# We are only optimizing towards Penaized LogP, which is contained in our template. For this reason, we don't have any additional score terms
# ## Loss Function
#
# We will use the `PPO` policy gradient algorithm
# +
pg = PPO(0.99,
0.5,
lam=0.95,
v_coef=0.5,
cliprange=0.3,
v_cliprange=0.3,
ent_coef=0.01,
kl_target=0.03,
kl_horizon=3000,
scale_rewards=True)
loss = PolicyLoss(pg, 'PPO',
value_head=ValueHead(256),
v_update_iter=2,
vopt_kwargs={'lr':1e-3})
# -
# ## Samplers
#
# We create the following samplers:
# - `sampler1 ModelSampler`: this samples from the main model
# - `sampler2 ModelSampler`: this samples from the baseline model
# - `sampler3 LogSampler`: this samples high scoring samples from the log
# - `sampler4 CombichemSampler`: this sampler runs combichem generation on the top scoring compounds. The combination of generative models with combichem greatly accelerates finding high scoring compounds
# +
gen_bs = 1500
sampler1 = ModelSampler(agent.vocab, agent.model, 'live', 400, 0., gen_bs)
sampler2 = ModelSampler(agent.vocab, agent.base_model, 'base', 400, 0., gen_bs)
sampler3 = LogSampler('samples', 'rewards', 10, 98, 100)
mutators = [
ChangeAtom(['6', '7', '8', '9']),
AppendAtomSingle(['C', 'N', 'O', 'F', 'Cl', 'Br']),
AppendAtomsDouble(['C', 'N', 'O']),
AppendAtomsTriple(),
DeleteAtom(),
ChangeBond(),
InsertAtomSingle(['C', 'N', 'O']),
InsertAtomDouble(['C', 'N']),
InsertAtomTriple(),
AddRing(),
ShuffleNitrogen(10)
]
mc = MutatorCollection(mutators)
crossovers = [FragmentCrossover()]
cbc = CombiChem(mc, crossovers, template=template, rewards=[],
prune_percentile=70, max_library_size=400, log=True, p_explore=0.2)
sampler4 = CombichemSampler(cbc, 20, 98, 0.2, 1, 'rewards', 'combichem')
samplers = [sampler1, sampler2, sampler3, sampler4]
# -
# ## Other Callbacks
#
# We add the following callbacks:
# - `supervised_cb`: every 20 batches, this callback grabs the top 2% of samples from the log and runs supervised training with these samples
# - `live_max`: prints the maximum score from `sampler1` each batch
# - `live_p90`: prints the top 10% score from `sampler1` each batch
# +
supervised_cb = SupervisedCB(agent, 20, 0.5, 98, 5e-4, 64, epochs=5)
live_max = MaxCallback('rewards', 'live')
live_p90 = PercentileCallback('rewards', 'live', 90)
cbs = [supervised_cb, live_p90, live_max]
# -
# ## Environment
#
# We create our environment with the objects assembled so far
env = Environment(agent, template_cb, samplers=samplers, rewards=[], losses=[loss],
cbs=cbs)
# ## Train
set_global_pool(min(10, os.cpu_count()))
env.fit(128, 150, 400, 25)
env.log.plot_metrics()
subset = env.log.df[env.log.df.rewards>23.6]
draw_mols(to_mols(subset.samples.values), legends=[f"{i:.5f}" for i in subset.rewards.values])
# Note that penalized LogP is strongly influenced by the size of the molecule generated. For this reason it's not a very good benchmark. Nonetheless it is very common in literature. Increasing the maximum sequence length from `150` (what we used in `Environment.fit` above) to something like `200` will result in higher penalized logP scores for the same amount of training
|
nbs/tutorials/tutorials.small_molecule.plogp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
n=int(input('Digite um numero: '))
tot = 0
for i in range(1,n+1):
if n%i == 0:
print('\033[33m', end=" ")
tot += 1
else:
print('\033[31m', end=" ")
print('{}'.format(i), end=" ")
print('\n\033[mO numero {} foi divisivel {} vezes'.format(n,tot))
if tot == 2:
print('É um numero PRIMO')
|
.ipynb_checkpoints/EX052 - Números Primos-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# [**HackerEarth: Machine Learning Challenge - Adopt a buddy**](https://www.hackerearth.com/challenges/competitive/hackerearth-machine-learning-challenge-pet-adoption/)
#
# **Problem Statement**
#
# A leading pet adoption agency is planning to create a virtual tour experience for their customers showcasing all animals that are available in
# their shelter. To enable this tour experience, you are required to build a Machine Learning model that determines type and breed of the
# animal based on its physical attributes and other factors.
#
# Data Description:
#
# The data folder consists of 2 CSV files
# • **train.csv** - 18834 x 11
# • **test.csv** - 8072 x 9
#
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 5GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# +
# Import useful libraries
import time
import re
import string
from numpy import mean
from numpy import set_printoptions
from sklearn.feature_selection import SelectKBest, f_classif
import matplotlib.pyplot as plt
plt.style.use('seaborn-darkgrid')
import seaborn as sns
from sklearn.model_selection import train_test_split, RandomizedSearchCV, StratifiedKFold, KFold, GridSearchCV
from sklearn.metrics import f1_score
from sklearn.preprocessing import LabelEncoder, StandardScaler
from sklearn.utils.multiclass import type_of_target
from catboost import CatBoostClassifier
from collections import Counter
import warnings
warnings.filterwarnings('ignore')
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
# Read dataset
train_data = pd.read_csv('/kaggle/input/hackerearth-ml-challenge-adopt-a-buddy/train.csv')
test_data = pd.read_csv('/kaggle/input/hackerearth-ml-challenge-adopt-a-buddy/test.csv')
train_data.columns = train_data.columns.str.lower().str.strip().str.replace(' ', '_').str.replace('(', '').str.replace(')', '')
test_data.columns = test_data.columns.str.lower().str.strip().str.replace(' ', '_').str.replace('(', '').str.replace(')', '')
# -
print('Train Data Shape: ', train_data.shape)
print('Test Data Shape: ', test_data.shape)
train_data.head()
train_data.isnull().sum()
# # Exploratory Data Analysis
# +
# See the distribution of outcome 1: breed_category
sns.countplot(x = 'breed_category',data = train_data)
sns.despine()
# -
# **It appears that breed category 2 is a minority class. We can use Smote from imblearn to oversample this class**
print(train_data.breed_category.value_counts())
print(train_data.pet_category.value_counts())
# +
# See the distribution of outcome 2: pet_category
sns.countplot(x = 'pet_category',data = train_data)
sns.despine()
# -
# **It appears that pet category 0 and 4 are among minority classes. We can use Smote from imblearn to oversample these classes**
# ### Analysing other features in combined train and test data
# +
train_data['type'] = 'train'
test_data['type'] = 'test'
master_data = pd.concat([train_data, test_data])
master_data['issue_date'] = pd.to_datetime(master_data['issue_date'], dayfirst = True)
master_data['listing_date'] = pd.to_datetime(master_data['listing_date'].apply(lambda x: x.split(' ')[0]), dayfirst = True)
# +
# Relation between length and breed category
plt.figure(figsize = (8, 6))
sns.boxplot(x = 'breed_category',y = 'lengthm',data = master_data)
plt.show()
# +
# Relation between length and pet category
plt.figure(figsize = (8, 6))
sns.boxplot(x = 'pet_category',y = 'lengthm',data = master_data)
plt.show()
# +
# See the distribution of outcome 2: pet_category
plt.figure(figsize = (8, 6))
sns.countplot(x = 'condition',data = master_data)
sns.despine()
# -
# We can treat the missing class - nan as a separate category itself
# +
# See the distribution of outcome 1: breed_category
plt.figure(figsize = (22, 5))
sns.countplot(x = 'color_type',data = master_data)
plt.xticks(rotation = 90)
plt.show()
# -
# **More than half of the features are among miority classes, we can extract first word to create new color feature to reduce the classes here**
plt.figure(figsize = (8, 6))
sns.scatterplot(x = master_data['lengthm'], y = master_data['heightcm']/100)
plt.show()
# 1. **Both the features length and height can be dropped which led to improvement of score by 1**
# 2. **Note: If using dimension features - convert to same unit (m/cm)**
plt.figure(figsize = (8, 5))
sns.distplot(master_data['lengthm'])
plt.show()
plt.figure(figsize = (8, 5))
df = master_data[['lengthm','heightcm']]
df['lengthcm'] = df['lengthm']*100
df[['lengthcm','heightcm']].boxplot()
plt.show()
# +
# Correlation matrix
plt.figure(figsize = (11, 10))
#plt.subplots(figsize=(10,8))
sns.heatmap(master_data.corr(), annot = True)
# -
# ## Insights
#
# * **Drop the pet dimension columns and x2 if required as they are weakly correlated and don't have enough predicting power for the 2 target variables**
# ### Feature Engineering - New features
# +
master_data['days_to_reach'] = master_data['listing_date'] - master_data['issue_date']
master_data['days_to_reach'] = master_data['days_to_reach'].apply(lambda x: int(str(x).split(' ')[0]))
master_data['age'] = master_data['days_to_reach'] / 365
master_data['age'] = master_data['age'].abs()
# Mapping for condition of pets
condition = {0.0: 'A', 1.0: 'B', 2.0: 'C'}
master_data['condition'] = master_data['condition'].map(condition)
master_data['condition'] = master_data['condition'].astype(str)
# Convert height to cms
master_data['heightm'] = master_data['heightcm'] / 100
master_data = master_data.drop(['heightcm'], axis = 1)
#length_mean = master_data['lengthm'].mean()
master_data.loc[(master_data['lengthm'] == 0), 'lengthm'] = 0.005
#master_data['len_to_height'] = master_data['lengthm']/master_data['heightm']
master_data['color_type'] = master_data['color_type'].apply(lambda x: x.lower())
master_data.head()
# +
# 2 records exist where the listing date is less than the issue date, convert them to positive
master_data.loc[(master_data['days_to_reach'] <= 0), 'days_to_reach'] = master_data.loc[(master_data['days_to_reach'] < 0), 'days_to_reach'] * -1
# +
# Generate master color feature from the available color_type
master_data['master_color'] = master_data['color_type'].apply(lambda x: x.split(' ')[0])
master_data['species'] = master_data['color_type'].apply(lambda x: x.split(' ')[1] if len(x.split(' ')) == 2 else x.split(' ')[0])
# +
# Generate time features - e.g. Quarter
master_data['issue_qtr'] = master_data['issue_date'].dt.quarter
master_data['list_qtr'] = master_data['listing_date'].dt.quarter
master_data['issue_yr'] = master_data['issue_date'].dt.year
master_data['list_yr'] = master_data['listing_date'].dt.year
master_data['issue_mth'] = master_data['issue_date'].dt.month
master_data['list_mth'] = master_data['listing_date'].dt.month
master_data['issue_weekend'] = master_data['issue_date'].apply(lambda x: 1 if x.dayofweek in [5, 6] else 0)
master_data['list_weekend'] = master_data['listing_date'].apply(lambda x: 1 if x.dayofweek in [5, 6] else 0)
master_data.head()
# +
# Get numerical columns
cat_cols = ['condition', 'color_type', 'master_color', 'species']
numerical_cols = master_data.columns[~master_data.columns.isin(cat_cols + ['pet_id', 'issue_date', 'listing_date', 'type', 'breed_category', 'pet_category'])].tolist()
numerical_cols
# -
# ## Standard Scaling the data
ss = StandardScaler()
master_data[numerical_cols] = ss.fit_transform(master_data[numerical_cols])
# ## Label Encoding only done to find out kBest parameters using ANOVA score
#
# * **Uncomment and run below cell only to find out kBest columns & then run entire script without label encoding the feature columns**
# +
#le = LabelEncoder()
#for col in cat_cols:
# master_data[col] = le.fit_transform(master_data[col])
#master_data[numerical_cols + cat_cols] = ss.fit_transform(master_data[numerical_cols + cat_cols])
# -
train_data.columns
# ### Separating the data back to train and test
# +
# Separate train and test data
train_data = master_data.loc[master_data['type'] == 'train']
test_data = master_data.loc[master_data['type'] == 'test']
train_data['breed_category'] =train_data['breed_category'].astype(str)
train_data['pet_category'] =train_data['pet_category'].astype(str)
testIDs = test_data['pet_id']
train_data = train_data.drop(['pet_id', 'issue_date', 'listing_date', 'type'], axis = 1)
for col in ['breed_category', 'pet_category']:
train_data[col] = train_data[col].apply(lambda x: np.float16(x))
train_data[col] = train_data[col].apply(lambda x: np.int8(x))
testData = test_data.drop(['issue_date', 'listing_date', 'type', 'x2', 'breed_category', 'pet_category'], axis = 1)
test_data = test_data.drop(['pet_id', 'issue_date', 'listing_date', 'type', 'x2', 'breed_category', 'pet_category'], axis = 1)
train_data = train_data[['condition', 'color_type', 'lengthm', 'x1', 'days_to_reach', 'age', 'heightm', 'master_color', 'species', 'issue_qtr', 'list_qtr',
'breed_category', 'issue_yr', 'list_yr', 'issue_mth', 'list_mth', 'issue_weekend', 'list_weekend', 'pet_category']]
train_data.head()
# -
test_data.head()
# ## Selecting KBest features using Anova F test
#
# * Uncomment below 2 cells only for kBest features
# ## Part 1: Pet Category
train_data_1 = train_data.copy()
"""
X1 = train_data_1.drop(['breed_category', 'pet_category'],axis = 1).values
y1 = train_data_1['pet_category'].values
for num_feats in range(1, 9):
print('Using {} features:'.format(num_feats))
test = SelectKBest(score_func = f_classif, k = num_feats)
fit = test.fit(X1, y1)
# summarize scores
set_printoptions(precision = 0)
for i in fit.scores_:
print(i)
print(fit.scores_)
features = fit.transform(X1)
# summarize selected features
print(features[0:num_feats + 1, :])
"""
# #### It appears we can drop length and height features as they carry lowest Anova F score
# ## Part 2: Breed Category
"""
X2 = train_data_1.drop(['breed_category'],axis = 1).values
y2 = train_data_1['breed_category'].values
for num_feats in range(1, 10):
print('Using {} features:'.format(num_feats))
test = SelectKBest(score_func = f_classif, k = num_feats)
fit = test.fit(X2, y2)
# summarize scores
set_printoptions(precision = 0)
for i in fit.scores_:
print(i)
print(fit.scores_)
features = fit.transform(X2)
# summarize selected features
print(features[0:num_feats + 1, :])
"""
# +
train_data_1 = train_data.drop(['lengthm', 'heightm'], axis = 1)
test_data_1 = test_data.copy()
test_data_1 = test_data.drop(['lengthm', 'heightm'], axis = 1)
# +
X = train_data_1.drop(['breed_category', 'pet_category'],axis = 1).values
y_1 = train_data_1['pet_category'].values
_cat_indices_ = [0, 1, 5, 6]
#_cat_indices_ = [0, 4, 5]
type_of_target(y_1)
# -
# # Model Building
# ## CatBoost Model
# ### Part 1: Predict the pet_category target class
#
# Predict the target variable **pet_category** which is independent of the **breed_category** class
#
# --> Model: CatBoost
# --> KFold splits: 6
# +
# Catboost for pet_category
kfold, scores = KFold(n_splits = 6, shuffle = True, random_state = 22), list()
for train, test in kfold.split(X):
X_train, X_test = X[train], X[test]
y_train, y_test = y_1[train], y_1[test]
model = CatBoostClassifier(random_state = 22, max_depth = 6, n_estimators = 1000, verbose = 1000, l2_leaf_reg = 1)
model.fit(X_train, y_train, cat_features = _cat_indices_)
preds = model.predict(X_test)
score = f1_score(y_test, preds, average = 'weighted')
scores.append(score)
print('Validation f1_score:', score)
print("Average Validation f1_score: ", sum(scores)/len(scores))
# -
y_Preds_1 = model.predict(test_data_1.values)
pet_cat = pd.DataFrame(data = {'pet_id': testIDs, 'pet_category': y_Preds_1.ravel()})
pet_cat.head()
test_data_1 = testData.merge(pet_cat, on = 'pet_id', how = 'left')
test_data_1 = test_data_1.drop(['pet_id', 'heightm', 'lengthm'], axis = 1)
# ### Part 2: Predict the breed_category target class
#
# Predict the target variable **breed_category** which is dependent of the **pet_category** class
#
# --> Model: CatBoost
# --> KFold splits: 6
# +
X = train_data_1.drop(['breed_category', 'issue_yr', 'list_yr', 'issue_mth', 'list_mth', 'issue_weekend', 'list_weekend'],axis = 1).values
y_2 = train_data_1['breed_category'].values
_cat_indices_ = [0, 1, 5, 6, 9]
#_cat_indices_ = [0, 4, 5, 8]
type_of_target(y_2)
# +
# Catboost for breed_category
kfold, scores = KFold(n_splits = 6, shuffle = True, random_state = 22), list()
for train, test in kfold.split(X):
X_train, X_test = X[train], X[test]
y_train, y_test = y_2[train], y_2[test]
model_1 = CatBoostClassifier(random_state = 22, max_depth = 8, n_estimators = 1000, verbose = 1000)
model_1.fit(X_train, y_train, cat_features = _cat_indices_)
preds = model_1.predict(X_test)
score = f1_score(y_test, preds, average = 'weighted')
scores.append(score)
print('Validation f1_score:', score)
print("Average Validation f1_score: ", sum(scores)/len(scores))
# -
# ## Getting Predictions
# +
y_Preds_2 = model_1.predict(test_data_1.drop(['issue_yr', 'list_yr', 'issue_mth', 'list_mth', 'issue_weekend', 'list_weekend'], axis = 1))
submission = pd.DataFrame(data = {'pet_id': testIDs, 'breed_category': y_Preds_2.ravel(), 'pet_category': y_Preds_1.ravel()})
submission.to_csv('HE_adopt_a_buddy_final_v1.csv', index = False)
submission.head()
# -
# # F1Score of 91.4 using CatBoostClassifier
|
Rank 12 Solution - Adopt A Buddy/rank-12-hacker-earth-adopt-a-buddy-f1score-91-4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Model a Galaxy Cluster
# ## Notebook for generating an example galaxy cluster model.
#
# This notebook goes through the steps to generate model data for galaxy cluster weak lensing observables. In particular, we define a galaxy cluster model that follows and NFW distribution and generate various profiles for the model (mass density, convergence, shear, etc.), which we plot. Note, a full pipeline to measure a galaxy cluster weak lensing mass requires fitting the observed (or mock) data to a model.
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# Imports specific to clmm
# +
import os
os.environ['CLMM_MODELING_BACKEND'] = 'ccl' # here you may choose ccl, nc (NumCosmo) or ct (cluster_toolkit)
import clmm
import clmm.theory as m
from clmm import Cosmology
# -
# Make sure we know which version we're using
clmm.__version__
# Define a cosmology using astropy
cosmo = Cosmology(H0 = 70.0, Omega_dm0 = 0.27 - 0.045, Omega_b0 = 0.045, Omega_k0 = 0.0)
# Define the galaxy cluster model. Here, we choose parameters that describe the galaxy cluster model, including the mass definition, concentration, and mass distribution. For the mass distribution, we choose a distribution that follows an NFW profile.
density_profile_parametrization = 'nfw'
mass_Delta = 200
cluster_mass = 1.e15
cluster_concentration = 4
z_cl = 1.
z_source = 2.
alpha = [2, -0.5]
# Quick test of all theory functionality
r3d = np.logspace(-2, 2, 100)
rho = m.compute_3d_density(r3d, mdelta=cluster_mass, cdelta=cluster_concentration,
z_cl=z_cl, cosmo=cosmo)
Sigma = m.compute_surface_density(r3d, cluster_mass, cluster_concentration, z_cl, cosmo=cosmo,
delta_mdef=mass_Delta,
halo_profile_model=density_profile_parametrization)
DeltaSigma = m.compute_excess_surface_density(r3d, cluster_mass, cluster_concentration, z_cl, cosmo=cosmo,
delta_mdef=mass_Delta,
halo_profile_model=density_profile_parametrization)
Sigmac = m.compute_critical_surface_density(cosmo, z_cluster=z_cl, z_source=z_source)
gammat = m.compute_tangential_shear(r3d, mdelta=cluster_mass, cdelta=cluster_concentration, z_cluster=z_cl,
z_source=z_source, cosmo=cosmo, delta_mdef=mass_Delta,
halo_profile_model=density_profile_parametrization,
z_src_model='single_plane')
kappa = m.compute_convergence(r3d, mdelta=cluster_mass, cdelta=cluster_concentration,
z_cluster=z_cl, z_source=z_source,
cosmo=cosmo, delta_mdef=mass_Delta,
halo_profile_model=density_profile_parametrization,
z_src_model='single_plane')
gt = m.compute_reduced_tangential_shear(r3d, mdelta=cluster_mass, cdelta=cluster_concentration,
z_cluster=z_cl, z_source=z_source, cosmo=cosmo,
delta_mdef=mass_Delta,
halo_profile_model=density_profile_parametrization,
z_src_model='single_plane')
mu = m.compute_magnification(r3d, mdelta=cluster_mass, cdelta=cluster_concentration,
z_cluster=z_cl, z_source=z_source, cosmo=cosmo,
delta_mdef=mass_Delta,
halo_profile_model=density_profile_parametrization,
z_src_model='single_plane')
mu_bias = m.compute_magnification_bias(r3d, alpha=alpha, mdelta=cluster_mass, cdelta=cluster_concentration,
z_cluster=z_cl, z_source=z_source, cosmo=cosmo,
delta_mdef=mass_Delta,
halo_profile_model=density_profile_parametrization,
z_src_model='single_plane')
# Plot the predicted profiles
def plot_profile(r, profile_vals, profile_label='rho', label = ''):
plt.loglog(r, profile_vals, label=label)
plt.xlabel('r [Mpc]', fontsize='xx-large')
plt.ylabel(profile_label, fontsize='xx-large')
plot_profile(r3d, rho, '$\\rho_{\\rm 3d}$')
plot_profile(r3d, Sigma, '$\\Sigma_{\\rm 2d}$')
plot_profile(r3d, DeltaSigma, '$\\Delta\\Sigma_{\\rm 2d}$')
plot_profile(r3d, kappa, '$\\kappa$')
plot_profile(r3d, gammat, '$\\gamma_t$')
plot_profile(r3d, gt, '$g_t$')
plot_profile(r3d, mu, '$\mu$')
# +
plot_profile(r3d, mu_bias[0]-1, profile_label = '$\delta_{\mu}$', label = '$\\alpha$ =' + str(alpha[0]))
plot_profile(r3d, mu_bias[1]-1, '$\delta_{\mu}$', label = '$\\alpha$ =' + str(alpha[1]))
plt.legend(fontsize='xx-large')
plt.yscale('linear')
plt.grid()
plt.ylim(-3,5)
# +
# The 2-halo term excess surface density is only implemented for the CCL and NC backends
# An error will be raised if using the CT backend instead
DeltaSigma_2h = m.compute_excess_surface_density_2h(r3d, z_cl, cosmo=cosmo, halobias=0.3)
plot_profile(r3d, DeltaSigma_2h, '$\\Delta\\Sigma_{\\rm 2h}$')
# +
# The 2-halo term excess surface density is only implemented for the CCL and NC backends
# An error will be raised if using the CT backend instead
Sigma_2h = m.compute_surface_density_2h(r3d, z_cl, cosmo=cosmo, halobias=0.3)
plot_profile(r3d, Sigma_2h, '$\\Sigma_{\\rm 2h}$')
# -
# ## Side note regarding the Einasto profile (CCL and NC backends only)
#
# The Einasto profile is supported by both the CCL and NumCosmo backends. In CCL, the value of the Einasto slope
# is not a free parameter and depends on cosmology, redshift and halo mass. In NumCosmo, the default value is $\alpha=0.25$ but can be set to any other value. This is a source of confusion that needs to be handled. In the meantime, the verbose option allows to print the value of $\alpha$ that is being used, as follows:
# +
rho = m.compute_3d_density(r3d, mdelta=cluster_mass, cdelta=cluster_concentration,
z_cl=z_cl, cosmo=cosmo, halo_profile_model='einasto',
verbose=True)
# With the NC backend, you may set the slope of the Einasto profile to the value of your choosing
# rho = m.compute_3d_density(r3d, mdelta=cluster_mass, cdelta=cluster_concentration,
# z_cl=z_cl, cosmo=cosmo, halo_profile_model='einasto', alpha_ein=0.1,
# verbose=True)
plot_profile(r3d, rho, '$\\rho_{\\rm 3d}$')
# -
|
examples/demo_theory_functionality.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 2.1 Birthdays and Zeller's algorithm.
#
# Zeller's algorithm computes the day of the week on which a given date will fall (or fell). Use the description of Zeller's algorithm, given below, to write a Python script that computes the day of the week on which the user"s birthday fell in the year they were born and prints the result to the screen.
#
# Zeller's algorithm:
# - We first define the following variables (which each stores a non-negative integer):
# - `month`: This variable corresponds to the month of the year (ranging from 3 to 14), with March having the value 3, April the value 4, …, December the value 12, and January and February (of the following year) being counted as months 13 and 14. Note that if `month == 13` or `month == 14`, then you should adjust `year` such that `year = year - 1`.
# - `year`: This variable corresponds to the year of the century (e.g., `year == 89` for the year 1989; `year = 5` for the year 2005).
# - `yy`: This variable corresponds to the last 2 digits of `year`. For examples, `yy == 89` for the year 1989 and `yy == 5` for the year 2005.
# - `century`: This variable corresponds to the first 2 digits of `year`. For examples, `century == 19` for the year 1989 and `century == 20` for the year 2005.
# - `day`: This variable corresponds to the day of the month, i.e., an integer values between 1 and 31.
# - The day of the week is then calculated by
# - first computing `resultant` using the following formula: $$\tt{resultant}=\left({\tt day}+\left\lfloor \frac{13\left({\tt month}+1\right)}{5}\right\rfloor +{\tt yy}+\left\lfloor \frac{{\tt yy}}{4}\right\rfloor+\left\lfloor \frac{{\tt century}}{4}\right\rfloor -2\times{\tt century}\right) \mod 7.$$
# - Followed by matching `resultant` value to its corresponding values under `Day of the Week` column in the following table:
#
# <center>
#
# | Resultant | Day of the Week |
# |-----------|-----------------|
# | 0 | Saturday |
# | 1 | Sunday |
# | 2 | Monday |
# | 3 | Tuesday |
# | 4 | Wednesday |
# | 5 | Thursday |
# | 6 | Friday |
#
# </center>
#
# The problem:
# - Have the user input their birth date.
# - Use Zeller's algorithm to compute the day of the week on which they were born.
# - Print out that day.
#
# Example interaction:
#
# >Date of Birth? 20<br>
# >Month of Birth? 5<br>
# >Year of Birth? 2005<br>
# >You were born on Friday.<br>
# +
import math
days = ['Saturday', 'Sunday', 'Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday']
date = int(input("Please input your date of birth in DD form: "))
month = int(input("Please input your month of birth in MM form: "))
year = str(input("Please input your year of birth in YYYY form: "))
if month <= 2:
month += 12
yy = (year[2: ])
century = (year[ :2])
resultant = int((date + (13*(month+1))/5 + int(yy) + int(yy)/4 + (int(century)/4) - 2*int(century))%7)
print("You were born on", days[resultant])
# -
# # Assignment 2.2 Rock, Paper, Scissors, Lizard, Spock
#
# In this exercise, you are going to further practice using conditionals (`if`, `elif`, `else`). You will write a small program that will determine the result of a Rock, Paper, Scissors, Lizard, Spock game.
#
# For the rules of the game, please which the following clip from "The Big Bang Theory":
# [https://www.youtube.com/watch?v=_PUEoDYpUyQ]
#
# [](https://youtu.be/_PUEoDYpUyQ "Everything Is AWESOME")
#
#
# The problem:
# - Write a program that will first request the input for Player 1 and Player 2's choices.
# - Next, the program will compare the inputs and print out the result of the game (i.e., who the winner is).
#
# Example interaction:
#
# >Player 1? <br>
# >rock<br>
# >Player 2?<br>
# >scissors<br>
# >Player 1 wins<br>
# +
a = input("Player 1, rock, paper, scissors, lizard or spock?: ")
b = input("Player 2, rock, paper, scissors, lizard or spock?: ")
if a == 'scissors' and b == 'paper':
print("Player 1 wins")
if a == 'paper' and b == 'rock':
print("Player 1 wins")
if a == 'rock' and b == 'lizard':
print("Player 1 wins")
if a == 'lizard' and b == 'spock':
print("Player 1 wins")
if a == 'spock' and b == 'scissors':
print("Player 1 wins")
if a == 'scissors' and b == 'lizard':
print("Player 1 wins")
if a == 'lizard' and b == 'paper':
print("Player 1 wins")
if a == 'paper' and b == 'spock':
print("Player 1 wins")
if a == 'spock' and b == 'rock':
print("Player 1 wins")
if a == 'rock' and b == 'scissors':
print("Player 1 wins")
if a == 'paper' and b == 'scissors':
print("Player 2 wins")
if a == 'rock' and b == 'paper':
print("Player 2 wins")
if a == 'lizard' and b == 'rock':
print("Player 2 wins")
if a == 'spock' and b == 'lizard':
print("Player 2 wins")
if a == 'scissors' and b == 'spock':
print("Player 2 wins")
if a == 'lizard' and b == 'scissors':
print("Player 2 wins")
if a == 'paper' and b == 'lizard':
print("Player 2 wins")
if a == 'spock' and b == 'paper':
print("Player 2 wins")
if a == 'rock' and b == 'spock':
print("Player 2 wins")
if a == 'scissors' and b == 'rock':
print("Player 2 wins")
# -
# # Assignment 2.3 The $i$th prime number
#
# Your program should print the $i$th prime number where $i$ is a user defined input. You may use the calculator on the following website to check your answer:
# [http://www.bigprimes.net/archive/prime/]
#
#
# The problem:
# - Request an integer input from the user.
# - Compute the $i$-th prime number, such that $i$ is the integer input by the user above.
# - Print the $i$-th prime number
#
# Example interaction:
#
# > Input i, where $p_i$ is the $i$th prime number: <br>
# > 101<br>
# > The 101th prime number is : <br>
# > 547 <br>
# +
i = int(input("Input i, where 𝑝𝑖 is the 𝑖th prime number: "))
prime_validation = True
prime = 3
counter = 0
while True:
isPrime = True
for u in range(2, int(prime**.5+1)):
if prime%u == 0:
isPrime = False
break
if isPrime:
counter+=1
if counter == i-1:
break
prime += 1
print(prime)
# -
|
Assignment-2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# <img src="images/rapids-logo.png" width="500"/>
# <br/>
# <br/>
# <h1> Introduction </h1>
# <br/>
#
# The RAPIDS suite of software libraries gives you the freedom to execute end-to-end data science and analytics pipelines entirely on GPUs. It relies on NVIDIA® CUDA® primitives for low-level compute optimization, but exposes that GPU parallelism and high-bandwidth memory speed through user-friendly Python interfaces.
#
# RAPIDS also focuses on common data preparation tasks for analytics and data science. This includes a familiar DataFrame API that integrates with a variety of machine learning algorithms for end-to-end pipeline accelerations without paying typical serialization costs. RAPIDS also includes support for multi-node, multi-GPU deployments, enabling vastly accelerated processing and training on much larger dataset sizes.
# -------
# <h1> Benefits of Using Rapids </h1>
# <br/>
# <img src="images/rapids_benefits.png">
# <p style="text-align: center"> image source: rapid.ai</p>
# <br/>
# <h1> Performance at Scale </h1>
# <br/>
# <img src="images/rapids-end-to-end-performance-chart-oss-page-r4.svg" width='700px'>
# <p style="text-align: center"> image source: rapid.ai</p>
# <br/>
# ------
#
# <h1> prerequisites </h1>
# <br/>
# <br/>
#
# <table style="text-align: center; width: 70%">
# <tr>
# <th>PreReq</th>
# <th>Version</th>
# </tr>
# <tr>
# <td>Ubuntu</td>
# <td>16.04 / 18.04</td>
# </tr>
# <tr>
# <td>CUDA</td>
# <td>9.2 +</td>
# </tr>
# <tr>
# <td>NVIDIA driver</td>
# <td>396.44 +</td>
# </tr>
# </table>
#
# ----
#
# <h2>Installation using Conda</h2>
#
# RAPIDS is currently available in Conda package manager (documentation say it will be available in PIP package manager soon).
#
# Type the following command in the terminal:
#
# ```bash
# $ conda install -c nvidia -c rapidsai -c numba -c conda-forge -c defaults cudf=0.3.0
# ```
#
# **Note:** This conda installation only applies to Linux and Python versions 3.5/3.6.
#
# You can create and activate a development environment using the conda commands:
#
# ```bash
# # create the conda environment (assuming in base `cudf` directory)
# $ conda env create --name cudf_dev --file conda/environments/dev_py35.yml
# # activate the environment
# $ source activate cudf_dev
# # when not using default arrow version 0.10.0, run
# $ conda install -c nvidia -c rapidsai -c numba -c conda-forge -c defaults pyarrow=$ARROW_VERSION
# ```
# -----
|
.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# dictionary with mixed keys
my_dict = {'name': 'John', 1: [2, 4, 3], 0: 'apple', 'hey_to_the_world': 'good moring, world!'}
print(my_dict)
|
benchbook/test_dict_issue.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Weighted Generalized Linear Models
import numpy as np
import pandas as pd
import statsmodels.formula.api as smf
import statsmodels.api as sm
# ## Weighted GLM: Poisson response data
#
# ### Load data
#
# In this example, we'll use the affair dataset using a handful of exogenous variables to predict the extra-marital affair rate.
#
# Weights will be generated to show that `freq_weights` are equivalent to repeating records of data. On the other hand, `var_weights` is equivalent to aggregating data.
print(sm.datasets.fair.NOTE)
# Load the data into a pandas dataframe.
data = sm.datasets.fair.load_pandas().data
# The dependent (endogenous) variable is ``affairs``
data.describe()
data[:3]
# In the following we will work mostly with Poisson. While using decimal affairs works, we convert them to integers to have a count distribution.
data["affairs"] = np.ceil(data["affairs"])
data[:3]
(data["affairs"] == 0).mean()
np.bincount(data["affairs"].astype(int))
# ## Condensing and Aggregating observations
#
# We have 6366 observations in our original dataset. When we consider only some selected variables, then we have fewer unique observations. In the following we combine observations in two ways, first we combine observations that have values for all variables identical, and secondly we combine observations that have the same explanatory variables.
# ### Dataset with unique observations
#
# We use pandas's groupby to combine identical observations and create a new variable `freq` that count how many observation have the values in the corresponding row.
data2 = data.copy()
data2['const'] = 1
dc = data2['affairs rate_marriage age yrs_married const'.split()].groupby('affairs rate_marriage age yrs_married'.split()).count()
dc.reset_index(inplace=True)
dc.rename(columns={'const': 'freq'}, inplace=True)
print(dc.shape)
dc.head()
# ### Dataset with unique explanatory variables (exog)
#
# For the next dataset we combine observations that have the same values of the explanatory variables. However, because the response variable can differ among combined observations, we compute the mean and the sum of the response variable for all combined observations.
#
# We use again pandas ``groupby`` to combine observations and to create the new variables. We also flatten the ``MultiIndex`` into a simple index.
# +
gr = data['affairs rate_marriage age yrs_married'.split()].groupby('rate_marriage age yrs_married'.split())
df_a = gr.agg(['mean', 'sum','count'])
def merge_tuple(tpl):
if isinstance(tpl, tuple) and len(tpl) > 1:
return "_".join(map(str, tpl))
else:
return tpl
df_a.columns = df_a.columns.map(merge_tuple)
df_a.reset_index(inplace=True)
print(df_a.shape)
df_a.head()
# -
# After combining observations with have a dataframe `dc` with 467 unique observations, and a dataframe `df_a` with 130 observations with unique values of the explanatory variables.
print('number of rows: \noriginal, with unique observations, with unique exog')
data.shape[0], dc.shape[0], df_a.shape[0]
# ## Analysis
#
# In the following, we compare the GLM-Poisson results of the original data with models of the combined observations where the multiplicity or aggregation is given by weights or exposure.
#
#
# ### original data
glm = smf.glm('affairs ~ rate_marriage + age + yrs_married',
data=data, family=sm.families.Poisson())
res_o = glm.fit()
print(res_o.summary())
res_o.pearson_chi2 / res_o.df_resid
# ### condensed data (unique observations with frequencies)
#
# Combining identical observations and using frequency weights to take into account the multiplicity of observations produces exactly the same results. Some results attribute will differ when we want to have information about the observation and not about the aggregate of all identical observations. For example, residuals do not take ``freq_weights`` into account.
glm = smf.glm('affairs ~ rate_marriage + age + yrs_married',
data=dc, family=sm.families.Poisson(), freq_weights=np.asarray(dc['freq']))
res_f = glm.fit()
print(res_f.summary())
res_f.pearson_chi2 / res_f.df_resid
# ### condensed using ``var_weights`` instead of ``freq_weights``
#
# Next, we compare ``var_weights`` to ``freq_weights``. It is a common practice to incorporate ``var_weights`` when the endogenous variable reflects averages and not identical observations.
# I do not see a theoretical reason why it produces the same results (in general).
#
# This produces the same results but ``df_resid`` differs the ``freq_weights`` example because ``var_weights`` do not change the number of effective observations.
#
glm = smf.glm('affairs ~ rate_marriage + age + yrs_married',
data=dc, family=sm.families.Poisson(), var_weights=np.asarray(dc['freq']))
res_fv = glm.fit()
print(res_fv.summary())
# Dispersion computed from the results is incorrect because of wrong ``df_resid``.
# It is correct if we use the original ``df_resid``.
res_fv.pearson_chi2 / res_fv.df_resid, res_f.pearson_chi2 / res_f.df_resid
# ### aggregated or averaged data (unique values of explanatory variables)
#
# For these cases we combine observations that have the same values of the explanatory variables. The corresponding response variable is either a sum or an average.
#
# #### using ``exposure``
#
# If our dependent variable is the sum of the responses of all combined observations, then under the Poisson assumption the distribution remains the same but we have varying `exposure` given by the number of individuals that are represented by one aggregated observation.
#
# The parameter estimates and covariance of parameters are the same with the original data, but log-likelihood, deviance and Pearson chi-squared differ
glm = smf.glm('affairs_sum ~ rate_marriage + age + yrs_married',
data=df_a, family=sm.families.Poisson(), exposure=np.asarray(df_a['affairs_count']))
res_e = glm.fit()
print(res_e.summary())
res_e.pearson_chi2 / res_e.df_resid
# #### using var_weights
#
# We can also use the mean of all combined values of the dependent variable. In this case the variance will be related to the inverse of the total exposure reflected by one combined observation.
glm = smf.glm('affairs_mean ~ rate_marriage + age + yrs_married',
data=df_a, family=sm.families.Poisson(), var_weights=np.asarray(df_a['affairs_count']))
res_a = glm.fit()
print(res_a.summary())
# ### Comparison
#
# We saw in the summary prints above that ``params`` and ``cov_params`` with associated Wald inference agree across versions. We summarize this in the following comparing individual results attributes across versions.
#
# Parameter estimates `params`, standard errors of the parameters `bse` and `pvalues` of the parameters for the tests that the parameters are zeros all agree. However, the likelihood and goodness-of-fit statistics, `llf`, `deviance` and `pearson_chi2` only partially agree. Specifically, the aggregated version do not agree with the results using the original data.
#
# **Warning**: The behavior of `llf`, `deviance` and `pearson_chi2` might still change in future versions.
#
# Both the sum and average of the response variable for unique values of the explanatory variables have a proper likelihood interpretation. However, this interpretation is not reflected in these three statistics. Computationally this might be due to missing adjustments when aggregated data is used. However, theoretically we can think in these cases, especially for `var_weights` of the misspecified case when likelihood analysis is inappropriate and the results should be interpreted as quasi-likelihood estimates. There is an ambiguity in the definition of ``var_weights`` because they can be used for averages with correctly specified likelihood as well as for variance adjustments in the quasi-likelihood case. We are currently not trying to match the likelihood specification. However, in the next section we show that likelihood ratio type tests still produce the same result for all aggregation versions when we assume that the underlying model is correctly specified.
results_all = [res_o, res_f, res_e, res_a]
names = 'res_o res_f res_e res_a'.split()
pd.concat([r.params for r in results_all], axis=1, keys=names)
pd.concat([r.bse for r in results_all], axis=1, keys=names)
pd.concat([r.pvalues for r in results_all], axis=1, keys=names)
pd.DataFrame(np.column_stack([[r.llf, r.deviance, r.pearson_chi2] for r in results_all]),
columns=names, index=['llf', 'deviance', 'pearson chi2'])
# ### Likelihood Ratio type tests
#
# We saw above that likelihood and related statistics do not agree between the aggregated and original, individual data. We illustrate in the following that likelihood ratio test and difference in deviance aggree across versions, however Pearson chi-squared does not.
#
# As before: This is not sufficiently clear yet and could change.
#
# As a test case we drop the `age` variable and compute the likelihood ratio type statistics as difference between reduced or constrained and full or unconstraint model.
# #### original observations and frequency weights
glm = smf.glm('affairs ~ rate_marriage + yrs_married',
data=data, family=sm.families.Poisson())
res_o2 = glm.fit()
#print(res_f2.summary())
res_o2.pearson_chi2 - res_o.pearson_chi2, res_o2.deviance - res_o.deviance, res_o2.llf - res_o.llf
glm = smf.glm('affairs ~ rate_marriage + yrs_married',
data=dc, family=sm.families.Poisson(), freq_weights=np.asarray(dc['freq']))
res_f2 = glm.fit()
#print(res_f2.summary())
res_f2.pearson_chi2 - res_f.pearson_chi2, res_f2.deviance - res_f.deviance, res_f2.llf - res_f.llf
# #### aggregated data: ``exposure`` and ``var_weights``
#
# Note: LR test agrees with original observations, ``pearson_chi2`` differs and has the wrong sign.
glm = smf.glm('affairs_sum ~ rate_marriage + yrs_married',
data=df_a, family=sm.families.Poisson(), exposure=np.asarray(df_a['affairs_count']))
res_e2 = glm.fit()
res_e2.pearson_chi2 - res_e.pearson_chi2, res_e2.deviance - res_e.deviance, res_e2.llf - res_e.llf
glm = smf.glm('affairs_mean ~ rate_marriage + yrs_married',
data=df_a, family=sm.families.Poisson(), var_weights=np.asarray(df_a['affairs_count']))
res_a2 = glm.fit()
res_a2.pearson_chi2 - res_a.pearson_chi2, res_a2.deviance - res_a.deviance, res_a2.llf - res_a.llf
# ### Investigating Pearson chi-square statistic
#
# First, we do some sanity checks that there are no basic bugs in the computation of `pearson_chi2` and `resid_pearson`.
res_e2.pearson_chi2, res_e.pearson_chi2, (res_e2.resid_pearson**2).sum(), (res_e.resid_pearson**2).sum()
res_e._results.resid_response.mean(), res_e.model.family.variance(res_e.mu)[:5], res_e.mu[:5]
(res_e._results.resid_response**2 / res_e.model.family.variance(res_e.mu)).sum()
res_e2._results.resid_response.mean(), res_e2.model.family.variance(res_e2.mu)[:5], res_e2.mu[:5]
(res_e2._results.resid_response**2 / res_e2.model.family.variance(res_e2.mu)).sum()
(res_e2._results.resid_response**2).sum(), (res_e._results.resid_response**2).sum()
# One possible reason for the incorrect sign is that we are subtracting quadratic terms that are divided by different denominators. In some related cases, the recommendation in the literature is to use a common denominator. We can compare pearson chi-squared statistic using the same variance assumption in the full and reduced model.
#
# In this case we obtain the same pearson chi2 scaled difference between reduced and full model across all versions. (Issue [#3616](https://github.com/statsmodels/statsmodels/issues/3616) is intended to track this further.)
((res_e2._results.resid_response**2 - res_e._results.resid_response**2) / res_e2.model.family.variance(res_e2.mu)).sum()
((res_a2._results.resid_response**2 - res_a._results.resid_response**2) / res_a2.model.family.variance(res_a2.mu)
* res_a2.model.var_weights).sum()
((res_f2._results.resid_response**2 - res_f._results.resid_response**2) / res_f2.model.family.variance(res_f2.mu)
* res_f2.model.freq_weights).sum()
((res_o2._results.resid_response**2 - res_o._results.resid_response**2) / res_o2.model.family.variance(res_o2.mu)).sum()
# ## Remainder
#
# The remainder of the notebook just contains some additional checks and can be ignored.
np.exp(res_e2.model.exposure)[:5], np.asarray(df_a['affairs_count'])[:5]
res_e2.resid_pearson.sum() - res_e.resid_pearson.sum()
res_e2.mu[:5]
res_a2.pearson_chi2, res_a.pearson_chi2, res_a2.resid_pearson.sum(), res_a.resid_pearson.sum()
((res_a2._results.resid_response**2) / res_a2.model.family.variance(res_a2.mu) * res_a2.model.var_weights).sum()
((res_a._results.resid_response**2) / res_a.model.family.variance(res_a.mu) * res_a.model.var_weights).sum()
((res_a._results.resid_response**2) / res_a.model.family.variance(res_a2.mu) * res_a.model.var_weights).sum()
res_e.model.endog[:5], res_e2.model.endog[:5]
res_a.model.endog[:5], res_a2.model.endog[:5]
res_a2.model.endog[:5] * np.exp(res_e2.model.exposure)[:5]
res_a2.model.endog[:5] * res_a2.model.var_weights[:5]
from scipy import stats
stats.chi2.sf(27.19530754604785, 1), stats.chi2.sf(29.083798806764687, 1)
res_o.pvalues
print(res_e2.summary())
print(res_e.summary())
print(res_f2.summary())
print(res_f.summary())
|
examples/notebooks/glm_weights.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Trimming
# See If the sequencing compony done that for you,
# If not, we can use [Trimmomatic](http://www.usadellab.org/cms/?page=trimmomatic) to handle the quality control part
# ## FastQC
# observe the sequencing quality
/home/junyuchen/Lab/Phage-SOP/Result/fastqc
fastqc /home/junyuchen/Lab/Phage-SOP/rawdata/jtshen-2020-06-13/CD1382_FDSW202399938-1r_1.clean.fq /home/junyuchen/Lab/Phage-SOP/rawdata/jtshen-2020-06-13/CD1382_FDSW202399938-1r_2.clean.fq -o /home/junyuchen/Lab/Phage-SOP/Result/fastqc -t 16
fastqc /home/junyuchen/Lab/Phage-SOP/rawdata/jtshen-2020-06-13/BT_FDSW202399939-1r_1.clean.fq /home/junyuchen/Lab/Phage-SOP/rawdata/jtshen-2020-06-13/BT_FDSW202399939-1r_2.clean.fq -o /home/junyuchen/Lab/Phage-SOP/Result/fastqc -t 16
/home/LDlab/JuntaoSHEN/PhageGenomeNGSdata
scp -r LDlab@10.10.1.134:/home/LDlab/JuntaoSHEN/PhageGenomeNGSdata/H101SC20051899.rar ./
import os
inputDir = "/home/junyuchen/Lab/Phage-SOP/rawdata/jtshen-2020-06-13"
for subdir, dirs, files in os.walk(inputDir):
R1 = ""
R2 = ""
outputFilePath = ""
R1List = []
R2List = []
outFileList = []
for file in files:
if file.endswith("_1.clean.fq"):
R1 = os.path.join(subdir, file)
R2 = os.path.join(subdir, file[:-10]+"2.clean.fq")
R1List.append(R1)
print(R1)
print(R2)
R2List.append(R2)
sampleStr = os.path.splitext(file)[0][:-8]
#outputFilePath = os.path.join(ouputDir, sampleStr)
#outFileList.append(outputFilePath)
sampleStr
R1List
R2List
# /mnt/data2/LD_lab/JunyuChen/Phage/Assembly-2020-06-13
|
Anno/Utility_PreProcess.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import glob
import pandas as pd
os.chdir("./Tableau")
extension = 'csv'
all_filenames = [i for i in glob.glob('*.{}'.format(extension))]
#combine all files
combined_csv = pd.concat([pd.read_csv(f) for f in all_filenames])
#export to csv
combined_csv.to_csv("combined_csv.csv", index=False, encoding='utf-8-sig')
combined_csv
|
Citibike combined CSV.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Importing Libraries
# Four libraries were used in this implementation
# 1. **pandas** for data manipulation and analysis(importing the datasets and combining them).
# 2. **numpy** for adding support for large, multi-dimensional arrays and matrices, along with a large collection of high-level mathematical functions to operate on these arrays (for processing the TMDB dataset)
# 3. **sklearn** for the Term Frequency Inverse Document Frequency which we use for calculating cosine similarity
# 4. **suprise** for the SVD machine learning algorithm used in coming up with the list of movie recommendations
# + pycharm={"name": "#%%\n"}
import pandas as pd
import numpy as np
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import linear_kernel
from surprise import Reader, Dataset, SVD
from surprise.model_selection import cross_validate
# -
# ## Importing the dataset(s)
# The whole dataset was sourced from [TMDB](https://www.themoviedb.org). Since TMDB does not offer a way to acquire the whole dataset at once, a series of api requests were sent to generate the following csv files which are then combined to form the dataset.
#
#
# ### 1. ratings
# The `ratings` dataset is 4 columned csv files which contains the individual movie ratings as well as the user who gave the rating and the timestamp at which the rating was given. This dataset is crucial because we are going to use it when recommending movies to a particular user. Through *collaborative filtering*, we will recommend movies to a user based on prior ratings that the user has given to other movies. Here is a snippet of the dataset
#
# 
#
# ### 2. metadata
# The `metadata` dataset contains multiple columns all of which contain important details about the individual movies, from the title of the movie, to the genre to the crew. The data in this dataset is crucial when we want to apply *content-based filtering* whereby we will recommend similar movies based on their description (movie synopsis) so a user will get movies that have a similar plot line to the one he/she has selected. Here is a snippet of the dataset
#
# 
#
# ### 3. id_mappings
# The `id_mappings` dataset contains the id mappings between the `movieId` and the `tmdbid`. The data in this dataset is crucial because it helps us combine the `ratings` dataset and the `metadata` dataset so that we can build our hybrid recommendation system. Here is a snippet of the dataset
#
# 
#
#
#
#
#
# + pycharm={"name": "#%%\n"}
ratings = pd.read_csv('../Data/movies_ratings.csv')
metadata = pd.read_csv('../Data/movies_metadata.csv')
id_mappings = pd.read_csv('../Data/movie_id_mappings.csv')[['movieId', 'tmdbId']]
# -
# ## Preparing the dataset
#
# The dataset needs to be cleaned up and prepared before we can make use of it.
# - The first thing we need to do is to make sure that we have dealt with all missing values in cells by replacing them with an empty string.
# - We also need to concatenate the overview and the tagline columns to form the description column which we will use in calculating the *cosine similarity*
# - Lastly, we need to create our `indices map` dataframe that contains the correct mappings between movie_id and tmdbid
#
# + pycharm={"name": "#%%\n"}
metadata['tagline'] = metadata['tagline'].fillna('')
metadata['description'] = metadata['overview'] + metadata['tagline']
metadata['description'] = metadata['description'].fillna('')
metadata['year'] = pd.to_datetime(metadata['release_date'], errors='coerce').apply(lambda x: str(x).split('-')[0] if x != np.nan else np.nan)
vote_averages = metadata[metadata['vote_average'].notnull()]['vote_average'].astype('int')
vote_counts = metadata[metadata['vote_count'].notnull()]['vote_count'].astype('int')
titles = metadata['title']
id_mappings['tmdbId'] = id_mappings['tmdbId'].apply(lambda s: int(s) if s.is_integer() else np.nan)
id_mappings.columns = ['movieId', 'id']
id_map = id_mappings.merge(metadata[['title', 'id']], on='id').set_index('title')
indices_map = id_map.set_index('id')
# -
# ## Demographic Filtering (Popularity-based recommender)
# They offer generalized recommendations to every user, based on movie popularity and/or genre. The System recommends the same movies to users with similar demographic features. Since each user is different , this approach is considered to be too simple. The basic idea behind this system is that movies that are more popular and critically acclaimed will have a higher probability of being liked by the average audience.
#
# Weighted Rating (WR) = $(\frac{v}{v + m} . R) + (\frac{m}{v + m} . C)$
#
# where,
#
# - v is the number of votes for the movie
# - m is the minimum votes required to be listed in the chart
# - R is the average rating of the movie
# - C is the mean vote across the whole report
#
# The next step is to determine an appropriate value for m, the minimum votes required to be listed in the chart. We will use 95th percentile as our cutoff. In other words, for a movie to feature in the charts, it must have more votes than at least 95% of the movies in the list.
# + pycharm={"name": "#%%\n"}
m = vote_counts.quantile(0.95)
weighted_dataset = metadata[(metadata['vote_count'] >= m) & (metadata['vote_count'].notnull()) & (metadata['vote_average'].notnull())][['title', 'year', 'vote_count', 'vote_average', 'popularity', 'genres']]
weighted_dataset['vote_count'] = weighted_dataset['vote_count'].astype('int')
weighted_dataset['vote_average'] = weighted_dataset['vote_average'].astype('int')
def weighted_rating(x):
C = vote_averages.mean()
v = x['vote_count']
R = x['vote_average']
return (v/(v+m) * R) + (m/(m+v) * C)
weighted_dataset['wr'] = weighted_dataset.apply(weighted_rating, axis=1)
weighted_dataset = weighted_dataset.sort_values('wr', ascending=False).head(250)
# -
#
# + pycharm={"name": "#%%\n"}
weighted_dataset.head(15)
# -
# ## Content-based Filtering with Cosine Similarity
#
# Cosine similarity measures the similarity between two vectors of an inner product space. It is measured by the cosine of the angle between two vectors and determines whether two vectors are pointing in roughly the same direction. It is described mathematically as the division between the dot product of vectors and the product of the euclidean norms or magnitude of each vector.
#
# cosine similarity = $ S_C(A,B) := cos(\theta) = \frac{A . B}{||A|| ||B||} = \frac{\sum^n_{i=1}A_i B_i}{\sqrt{\sum^n_{i=1}A^2_i} \sqrt{\sum^n_{i=1}B^2_i}}$
#
# Cosine Similarity is a value that is bound by a constrained range of 0 and 1.
# The similarity measurement is a measure of the cosine of the angle between the two non-zero vectors A and B.
# Suppose the angle between the two vectors was 90 degrees. In that case, the cosine similarity will have a value of 0; this means that the two vectors are orthogonal or perpendicular to each other.
# As the cosine similarity measurement gets closer to 1, then the angle between the two vectors A and B is smaller.
#
#
# This scenario involves the requirement of identifying the similarity between pairs of movies and is a good use case for the utilisation of cosine similarity as a quantification of the measurement of similarity between two objects.
# Quantification of the similarity between two movies can be obtained by converting the description within of the movie into a vectorised form of representation.
# The vector representations of the movies can then be used within the cosine similarity formula to obtain a quantification of similarity.
# In the scenario described above, the cosine similarity of 1 implies that the two movies are exactly alike and a cosine similarity of 0 would point to the conclusion that there are no similarities between the two movies.
#
# ### How we are using cosine similarity
# We are applying cosine similarity by looking for movies that have a similar description plot and then returning a matrix of these movies.
# The cosine similarity is calculated using the Term Frequency Inverse Document Frequency (TF-IDF) which is powerful algorithm for transforming text into a meaningful representation of numbers which is used to fit cosine similarity algorithm.
#
# + pycharm={"name": "#%%\n"}
def get_recommendations_using_cosine_similarity(title):
tf = TfidfVectorizer(analyzer='word',ngram_range=(1, 2),min_df=0, stop_words='english')
tfidf_matrix = tf.fit_transform(metadata['description'])
indices = pd.Series(metadata.index, index=metadata['title'])
cosine_similarity = linear_kernel(tfidf_matrix, tfidf_matrix)
movie_index = indices[title]
similar_movies = list(enumerate(cosine_similarity[int(movie_index)]))
similar_movies = sorted(similar_movies, key=lambda x: x[1], reverse=True)
similar_movies = similar_movies[1:20]
return [i[0] for i in similar_movies]
# -
#
# #### Top 15 movies based on their cosine similarity
# + pycharm={"name": "#%%\n"}
titles.iloc[get_recommendations_using_cosine_similarity('The Godfather')].head(15)
# -
# ## Collaborative Filtering with SVD
# The Singular Value Decomposition (SVD), a method from linear algebra that has been generally used as a dimensionality reduction technique in machine learning. SVD is a matrix factorisation technique, which reduces the number of features of a dataset by reducing the space dimension from N-dimension to K-dimension (where K<N). In the context of the recommender system, the SVD is used as a collaborative filtering technique. It uses a matrix structure where each row represents a user, and each column represents an item. The elements of this matrix are the ratings that are given to items by users.
#
# The factorisation of this matrix is done by the singular value decomposition. It finds factors of matrices from the factorisation of a high-level (user-item-rating) matrix. The singular value decomposition is a method of decomposing a matrix into three other matrices as given below:
# <br/>
#
# $A = U\Sigma V^{T}$
#
# Where A is a m x n utility matrix, U is a m x r orthogonal left singular matrix, which represents the relationship between users and latent factors, S is a r x r diagonal matrix, which describes the strength of each latent factor and V is a r x n diagonal right singular matrix, which indicates the similarity between items and latent factors. The latent factors here are the characteristics of the items, for example, the genre of the movie. The SVD decreases the dimension of the utility matrix A by extracting its latent factors. It maps each user and each item into a r-dimensional latent space. This mapping facilitates a clear representation of relationships between users and items.
# ## Creating the training set
# Our SVD model will be trained using the `ratings` dataset.
# + pycharm={"name": "#%%\n"}
reader = Reader()
X_train = Dataset.load_from_df(ratings[['userId', 'movieId', 'rating']], reader)
# -
# ## Training the SVD model on the Training set
# We will build our SVD model using the training set data
# + pycharm={"name": "#%%\n"}
svd = SVD()
trainset = X_train.build_full_trainset()
svd.fit(trainset)
# -
# ### Evaluating the SVD model
# + pycharm={"name": "#%%\n"}
cross_validate(svd, X_train, measures=['RMSE', 'MAE'])
# -
# ### Hybrid Recommendation System
# + pycharm={"name": "#%%\n"}
def recommend_movies(user_id, title):
movie_indices = get_recommendations_using_cosine_similarity(title)
movies = metadata.iloc[movie_indices][['title', 'vote_count', 'vote_average', 'release_date', 'id']]
movies['est'] = movies['id'].apply(lambda x: svd.predict(user_id, indices_map.loc[x]['movieId']).est)
movies = movies.sort_values('est', ascending=False)
return movies.head(10)
# -
# ## Testing the model on example data
# Here we call the `recommend_movies` function and provide it with a a title of a movie as well as the user_id of the current user
# + pycharm={"name": "#%%\n"}
recommend_movies(user_id=6, title='The Matrix')
# + [markdown] pycharm={"name": "#%% md\n"}
#
|
Model/svd.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
from pydarknet.libdarknet import Libdarknet
import ctypes
import os
self = Libdarknet()
self
self.lib
self.exists
lib = self.lib
# +
# Relative files are **relative to the .darknet directory!
metadata = self.get_metadata("cfg/coco.data")
network = self.load_network("cfg/yolov3.cfg", "weights/yolov3.weights", 0)
img = self.load_image_color("data/dog.jpg")
# -
from pydarknet.libdarknet.structs import Image, Detection
# +
self.lib.network_predict_image.argtypes = [ctypes.c_void_p, Image]
self.lib.network_predict_image.restype = ctypes.POINTER(ctypes.c_float)
self.lib.network_predict_image(network, img)
# -
net_output = self.network_predict_image(network, img)
net_output[0]
self.get_network_boxes(network, img)
num = ctypes.c_int(0)
pnum = ctypes.pointer(num)
dets = self.lib.get_network_boxes(network, img.w, img.h, 0.5, 0.5, None, 0, pnum)
for idx in range(num.value):
det=dets[idx]
print(det.objectness, det.prob[0])
det.classes
thresh=0.5
heir_thresh=0.5
# +
self.lib.get_network_boxes.argtypes = [
ctypes.c_void_p,
ctypes.c_int,
ctypes.c_int,
ctypes.c_float,
ctypes.c_float,
ctypes.POINTER(ctypes.c_int),
ctypes.c_int,
ctypes.POINTER(ctypes.c_int),
]
self.lib.get_network_boxes.restype = ctypes.POINTER(Detection)
num = ctypes.c_int(0)
pnum = ctypes.pointer(num)
dets = self.lib.get_network_boxes(network, img.w, img.h, thresh, heir_thresh, None, 0, pnum)
# -
dets
pnum[0]
num.value
num = pnum[0]
num
det=dets[4]
det
det.classes
from pydarknet.libdarknet.structs import Detection
nms=0.5
self.lib.do_nms_sort.argtypes = [ctypes.POINTER(Detection), ctypes.c_int, ctypes.c_int, ctypes.c_float]
self.lib.do_nms_sort(dets, num, metadata.classes, nms);
dets[0]
dets[:num-1]
d = dets[0]
dets[num-1].bbox
d.classes
d.objectness
d.sort_class
dets[0:num-1]
dets
pnum
dets = self.get_network_boxes(network, img)
dets
self.free_image(img)
|
Development/InitialDevelopment/module_libdarknet-Copy4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Legal Studies 190 - Data, Prediction, and Law
#
# Welcome to our class! This introductory notebook will reviews concepts that you may already be familiar with from Data 8 or similar courses. The basic strategies and tools for data analysis covered in this notebook will be the foundations of this class. It will cover an overview of our software and some programming concepts.
#
# ## Table of Contents
#
# 1 - [Computing Environment](#computing environment)
#
# 2 - [Coding Concepts](#programming concepts)
#
# 1 - [Python Basics](#python basics)
#
# 2 - [Pandas](#tables)
# ## Our Computing Environment, Jupyter notebooks <a id='computing environment'></a>
# This webpage is called a Jupyter notebook. A notebook is a place to write programs and view their results.
#
# ### Text cells
# In a notebook, each rectangle containing text or code is called a *cell*.
#
# Text cells (like this one) can be edited by double-clicking on them. They're written in a simple format called [Markdown](http://daringfireball.net/projects/markdown/syntax) to add formatting and section headings. You don't need to learn Markdown, but you might want to.
#
# After you edit a text cell, click the "run cell" button at the top that looks like ▶| to confirm any changes. (Try not to delete the instructions of the lab.)
# **Understanding Check 1** This paragraph is in its own text cell. Try editing it so that this sentence is the last sentence in the paragraph, and then click the "run cell" ▶| button . This sentence, for example, should be deleted. So should this one.
# ### Code cells
# Other cells contain code in the Python 3 language. Running a code cell will execute all of the code it contains.
#
# To run the code in a code cell, first click on that cell to activate it. It'll be highlighted with a little green or blue rectangle. Next, either press ▶| or hold down the `shift` key and press `return` or `enter`.
#
# The fundamental building block of Python code is an expression. Cells can contain multiple lines with multiple expressions. When you run a cell, the lines of code are executed in the order in which they appear. Every `print` expression prints a line. Run the next cell and notice the order of the output.
print("First this line is printed,")
print("and then this one.")
print("\N{WAVING HAND SIGN}, \N{EARTH GLOBE ASIA-AUSTRALIA}!")
# Don't be scared if you see a "Kernel Restarting" message! Your data and work will still be saved. Once you see "Kernel Ready" in a light blue box on the top right of the notebook, you'll be ready to work again. You should rerun any cells with imports, variables, and loaded data.
# <img src="images/kernel.png">
# ### Writing Jupyter notebooks
# You can use Jupyter notebooks for your own projects or documents. When you make your own notebook, you'll need to create your own cells for text and code.
#
# To add a cell, click the + button in the menu bar. It'll start out as a text cell. You can change it to a code cell by clicking inside it so it's highlighted, clicking the drop-down box next to the restart (⟳) button in the menu bar, and choosing "Code".
# ### Errors
# Python is a language, and like natural human languages, it has rules. It differs from natural language in two important ways:
# 1. The rules are *simple*. You can learn most of them in a few weeks and gain reasonable proficiency with the language in a semester.
# 2. The rules are *rigid*. If you're proficient in a natural language, you can understand a non-proficient speaker, glossing over small mistakes. A computer running Python code is not smart enough to do that.
#
# Whenever you write code, you'll make mistakes. When you run a code cell that has errors, Python will sometimes produce error messages to tell you what you did wrong.
#
# Errors are okay; even experienced programmers make many errors. When you make an error, you just have to find the source of the problem, fix it, and move on.
#
# We have made an error in the next cell. Run it and see what happens.
print("This line is missing something."
SOLUTION: print("This line is missing something.")
# You should see something like this (minus our annotations):
#
# <img src="images/error.jpg"/>
#
# The last line of the error output attempts to tell you what went wrong. The *syntax* of a language is its structure, and this `SyntaxError` tells you that you have created an illegal structure. "`EOF`" means "end of file," so the message is saying Python expected you to write something more (in this case, a right parenthesis) before finishing the cell.
#
# There's a lot of terminology in programming languages, but you don't need to know it all in order to program effectively. If you see a cryptic message like this, you can often get by without deciphering it. (Of course, if you're frustrated, feel free to ask a friend or post on the class Piazza.)
#
# **Understanding Check 2** Try to fix the code above so that you can run the cell and see the intended message instead of an error.
# ## Programming Concepts <a id='programming concepts'></a>
#
# Now that you are comfortable with our computing environment, we are going to be moving into more of the fundamentals of Python, but first, run the cell below to ensure all the libraries needed for this notebook are installed.
# ### Part 1: Python basics <a id='python basics'></a>
# Before getting into the more advanced analysis techniques that will be required in this course, we need to cover a few of the foundational elements of programming in Python.
# #### A. Expressions
# The departure point for all programming is the concept of the __expression__. An expression is a combination of variables, operators, and other Python elements that the language interprets and acts upon. Expressions act as a set of instructions to be fed through the interpreter, with the goal of generating specific outcomes. See below for some examples of basic expressions.
# +
# Examples of expressions:
#addition
print(2 + 2)
#string concatenation
print('me' + ' and I')
#you can print a number with a string if you cast it
print("me" + str(2))
#exponents
print(12 ** 2)
# -
# You will notice that only the last line in a cell gets printed out. If you want to see the values of previous expressions, you need to call `print` on that expression. Try adding `print` statements to some of the above expressions to get them to display.
# #### Data Types
#
# In Python, all things have a type. In the above example, you saw saw *integers* (positive and negative whole numbers) and *strings* (sequences of characters, often thought of as words or sentences). We denote strings by surrounding the desired value with quotes. For example, "Data Science" and "2017" are strings, while `bears` and `2020` (both without quotes) are not strings (`bears` without quotes would be interpreted as a variable). You'll also be using decimal numbers in Python, which are called *floats* (positive and negative decimal numbers).
#
# You'll also often run into *booleans*. They can take on one of two values: `True` or `False`. Booleans are often used to check conditions; for example, we might have a list of dogs, and we want to sort them into small dogs and large dogs. One way we could accomplish this is to say either `True` or `False` for each dog after seeing if the dog weighs more than 15 pounds.
#
# We'll soon be going over additional data types. Below is a table that summarizes the information in this section:
# |Variable Type|Definition|Examples|
# |-|-|-|
# |Integer|Positive and negative whole numbers|`42`, `-10`, `0`|
# |Float|Positive and negative decimal numbers|`73.9`, `2.4`, `0.0`|
# |String|Sequence of characters|`"Go Bears!"`, `"variables"`|
# |Boolean|True or false value|`True`, `False`|
#
# #### B. Variables
# In the example below, `a` and `b` are Python objects known as __variables__. We are giving an object (in this case, an `integer` and a `float`, two Python data types) a name that we can store for later use. To use that value, we can simply type the name that we stored the value as. Variables are stored within the notebook's environment, meaning stored variable values carry over from cell to cell.
# +
a = 4
b = 10/5
# Notice that 'a' retains its value.
print(a)
a + b
# -
# #### Question 1: Variables
# See if you can write a series of expressions that creates two new variables called __x__ and __y__ and assigns them values of __10.5__ and __7.2__. Then assign their product to the variable __combo__ and print it.
# Fill in the missing lines to complete the expressions.
#SOLUTION
x = 10.5
y = 7.2
combo = x * y
print(combo)
# Check to see if the value you get for **combo** is what you expect it to be.
# #### C. Lists
# The next topic is particularly useful in the kind of data manipulation that you will see throughout this class. The following few cells will introduce the concept of __lists__ (and their counterpart, `numpy arrays`). Read through the following cell to understand the basic structure of a list.
#
# A list is an ordered collection of objects. They allow us to store and access groups of variables and other objects for easy access and analysis. Check out this [documentation](https://www.tutorialspoint.com/python/python_lists.htm) for an in-depth look at the capabilities of lists.
#
# To initialize a list, you use brackets. Putting objects separated by commas in between the brackets will add them to the list.
# +
# an empty list
lst = []
print(lst)
# reassigning our empty list to a new list
lst = [1, 3, 6, 'lists', 'are' 'fun', 4]
print(lst)
#lists in python are zero-indexed so the indices for lst are 0,1,2,3,4,5 and 6
example = lst[2]
print(example)
#list slicing: This line will store the first (inclusive) through fourth (exclusive) elements of lst as a new list
#called lst_2:
lst_2 = lst[1:4]
lst_2
# -
# It is important to note that when you store a list to a variable, you are actually storing the **pointer** to the list. That means if you assign your list to another variable, and you change the elements in your other variable, then you are changing the same data as in the original list.
a = [1,2,3] #original list
b = a #b now points to list a
b[0] = 4
print(a[0]) #return 4 since we modified the first element of the list pointed to by a and b
# #### Question 2: Lists
# Build a list of length 10 containing whatever elements you'd like. Then, slice it into a new list of length five using a index slicing. Finally, assign the last element in your sliced list to the given variable and print it.
# +
### Fill in the ellipses to complete the question.
my_list = [3,5,2,7,8,3,0,9,7,6]
my_list_sliced = my_list[:5] #last index is exclusive/not included in the slicing
last_of_sliced = my_list_sliced[len(my_list_sliced)-1]
print(last_of_sliced)
# -
# Lists can also be operated on with a few built-in analysis functions. These include `min` and `max`, among others. Lists can also be concatenated together. Find some examples below.
# +
# A list containing six integers.
a_list = [1, 6, 4, 8, 13, 2]
# Another list containing six integers.
b_list = [4, 5, 2, 14, 9, 11]
print('Max of a_list:', max(a_list))
print('Min of b_list:', min(a_list))
# Concatenate a_list and b_list:
c_list = a_list + b_list
print('Concatenated:', c_list)
# -
# #### D. Numpy Arrays
# Closely related to the concept of a list is the array, a nested sequence of elements that is structurally identical to a list. Arrays, however, can be operated on arithmetically with much more versatility than regular lists. For the purpose of later data manipulation, we'll access arrays through [Numpy](https://docs.scipy.org/doc/numpy/reference/routines.html), which will require an import statement.
#
# Now run the next cell to import the numpy library into your notebook, and examine how numpy arrays can be used.
import numpy as np
# +
# Initialize an array of integers 0 through 9.
example_array = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
# This can also be accomplished using np.arange
example_array_2 = np.arange(10)
print('Undoubled Array:', example_array_2)
# Double the values in example_array and print the new array.
double_array = example_array*2
print('Doubled Array:', double_array)
# -
# This behavior differs from that of a list. See below what happens if you multiply a list.
example_list = [1, 2, 3, 4, 5, 6, 7, 8, 9]
example_list * 2
# Notice that instead of multiplying each of the elements by two, multiplying a list and a number returns that many copies of that list. This is the reason that we will sometimes use Numpy over lists. Other mathematical operations have interesting behaviors with lists that you should explore on your own.
# #### E. Looping
# [Loops](https://www.tutorialspoint.com/python/python_loops.htm) are often useful in manipulating, iterating over, or transforming large lists and arrays. The first type we will discuss is the __for loop__. For loops are helpful in traversing a list and performing an action at each element. For example, the following code moves through every element in example_array, adds it to the previous element in example_array, and copies this sum to a new array.
#
# It's important to note that "element" is an arbitrary variable name used to represent whichever index value the loop is currently operating on. We can change the variable name to whatever we want and achieve the same result, as long as we stay consistent.
# +
new_list = []
for element in example_array:
new_element = element + 5
new_list.append(new_element)
print(new_list)
#iterate using list indices rather than elements themselves
for i in range(len(example_array)):
example_array[i] = example_array[i] + 5
example_array
# -
# #### Other types of loops
# The __while loop__ repeatedly performs operations until a conditional is no longer satisfied. A conditional is a [boolean expression](https://en.wikipedia.org/wiki/Boolean_expression), that is an expression that evaluates to `True` or `False`.
#
# In the below example, an array of integers 0 to 9 is generated. When the program enters the while loop on the subsequent line, it notices that the maximum value of the array is less than 50. Because of this, it adds 1 to the fifth element, as instructed. Once the instructions embedded in the loop are complete, the program refers back to the conditional. Again, the maximum value is less than 50. This process repeats until the the fifth element, now the maximum value of the array, is equal to 50, at which point the conditional is no longer true and the loop breaks.
# +
while_array = np.arange(10) # Generate our array of values
print('Before:', while_array)
while(max(while_array) < 50): # Set our conditional
while_array[4] += 1 # Add 1 to the fifth element if the conditional is satisfied
print('After:', while_array)
# -
# #### Question 3: Loops
# In the following cell, partial steps to manipulate an array are included. You must fill in the blanks to accomplish the following: <br>
# 1. Iterate over the entire array, checking if each element is a multiple of 5
# 2. If an element is not a multiple of 5, add 1 to it repeatedly until it is
# 3. Iterate back over the list and print each element.
#
# > Hint: To check if an integer `x` is a multiple of `y`, use the modulus operator `%`. Typing `x % y` will return the remainder when `x` is divided by `y`. Therefore, (`x % y != 0`) will return `True` when `y` __does not divide__ `x`, and `False` when it does.
# +
# Make use of iterators, range, length, while loops, and indices to complete this question.
#SOLUTIONS
question_3 = np.array([12, 31, 50, 0, 22, 28, 19, 105, 44, 12, 77])
for i in range(len(question_3)):
while(question_3[i] % 5 != 0):
question_3[i] = question_3[i] + 1
for element in question_3:
print(element)
# -
# The following cell should return `True` if your code is correct.
answer = np.array([15, 35, 50, 0, 25, 30, 20, 105, 45, 15, 80])
question_3 == answer
# #### F. Functions!
# Functions are useful when you want to repeat a series of steps on multiple different objects, but don't want to type out the steps over and over again. Many functions are built into Python already; for example, you've already made use of `len()` to retrieve the number of elements in a list. You can also write your own functions, and at this point you already have the skills to do so.
#
#
# Functions generally take a set of __parameters__ (also called inputs), which define the objects they will use when they are run. For example, the `len()` function takes a list or array as its parameter, and returns the length of that list.
#
# Let's look at a function that takes two parameters, compares them somehow, and then returns a boolean value (`True` or `False`) depending on the comparison. The `is_multiple` function below takes as parameters an integer `m` and an integer `n`, checks if `m` is a multiple of `n`, and returns `True` if it is. Otherwise, it returns `False`.
#
# `if` statements, just like `while` loops, are dependent on boolean expressions. If the conditional is `True`, then the following indented code block will be executed. If the conditional evaluates to `False`, then the code block will be skipped over. Read more about `if` statements [here](https://www.tutorialspoint.com/python/python_if_else.htm).
def is_multiple(m, n):
if (m % n == 0):
return True
else:
return False
is_multiple(12, 4)
is_multiple(12, 7)
# **Sidenote:** Another way to write `is_multiple` is below, think about why it works.
#
# def is_multiple(m, n):
# return m % n == 0
#
# Since functions are so easily replicable, we can include them in loops if we want. For instance, our `is_multiple` function can be used to check if a number is prime! See for yourself by testing some possible prime numbers in the cell below.
# +
# Change possible_prime to any integer to test its primality
# NOTE: If you happen to stumble across a large (> 8 digits) prime number, the cell could take a very, very long time
# to run and will likely crash your kernel. Just click kernel>interrupt if it looks like it's caught.
possible_prime = 9999991
for i in range(2, possible_prime):
if (is_multiple(possible_prime, i)):
print(possible_prime, 'is not prime')
break
if (i >= possible_prime/2):
print(possible_prime, 'is prime')
break
# -
# ### Part 2: Pandas <a id='tables'></a>
#
# We will be using Pandas tables for much of this class to organize and sort through tabular data. [Pandas](http://pandas.pydata.org/pandas-docs/stable/api.html) is a library that is used for manipulating tabular data. It has a user-friendly API, and can be used to answer difficult questions in relatively few commands. Like we did with `numpy`, we will have to import `pandas`.
from pandas import *
# #### Creating DataFrames
# When dealing with a collection of things with multiple attributes, it can be useful to put the data in a _dataframe_. DataFrames are a nice way of organizing data in a 2-dimensional data set. The `head(n)` function outputs the first n rows and by default, the first 5 rows. For example, take a look at the table below.
pandas.read_csv('../data/anes/ANES_legalst123.csv').head(5)
# This table is from the Incident Record-Type File of the NCVS. See page 31 of the codebook (on bCourses) for a description of the survey. To create this table, we have drawn the data from the path `data/anes`, stored in a file called `ANES_legalst123.csv`. In general, to import data from a `.csv` file, we write **`pandas.read_table("file_name")`.** Information in `.csv`'s are separated by commas, and are what are typically used with the `pandas` package.
#
# We can also create our own DataFrames from scratch without having to import data from another file. Let's say we have two arrays, one with a list of fruits, and another with a list of their price at the Berkeley Student Food Collective. Then, we can create a new `DataFrame` with each of these arrays as columns with the `with_columns` method:
fruit_names = ['Apple', 'Orange', 'Banana']
fruit_prices = [1, 0.75, 0.5]
fruit_table = pandas.DataFrame(data = {
"Fruit": fruit_names,
"Price ($)": fruit_prices
})
fruit_table
# The **`with_columns`** method takes in pairs of column labels and arrays, and creates a new DataFrame with each array as a column of the DataFrame. Finally, to create a new dataframe (with no columns or rows), we simply write
empty_table = DataFrame()
empty_table
# We typically start off with empty tables when we need to add rows inside for loops, which we'll see later.
# ## Accessing Values
# Often, it is useful to access only the rows, columns, or values related to our analysis. We'll look at several ways to cut down our table into smaller, more digestible parts.
#
# Let's go back to our table of incidents.
# ** Exercise 1 **
#
# Below, assign a variable named `anes` to the data from the `ANES_legalst123.csv` file with the path `../data/anes/`, then display the table. (Hint: use the `read_table` function from the previous section and don't forget about the parameter `delimiter`). We will take a closer look at the ANES data in Lab 3.
# +
# YOUR CODE HERE
anes = pandas.read_csv('../data/anes/ANES_legalst123.csv')
#the head function selects the first n rows or default value of 5 rows
anes.head(5)
# -
# Notice that not all of the rows are displayed--in fact, there are over 10000 rows in the DataFrame! By default, we are shown the first 10 rows.
#
# However, let's say we wanted to grab only the first _five_ rows of this DataFrame. We can do this by using the **`loc`** function; it takes in a list or range of numbers, and creates a new DataFrame with rows from the original DataFrame whose indices are given in the array or range. Remember that in Python, indices start at 0! Below are a few examples:
anes.iloc[[1, 3, 5]] # Takes rows with indices 1, 3, and 5 (the 2nd, 4th, and 6th rows)
anes.iloc[[7]] # Takes the row with index 7 (8th row)
anes.iloc[np.arange(7)] # Takes the row with indices 0, 1, ... 6
# Similarly, we can also choose to display certain columns of the DataFrame. There are two methods to accomplish this, and both methods take in lists of either column indices or column labels:
# - Insert the names of the columns as a list in the DataFrame
# - The **`drop`** method creates a new DataFrame with all columns _except_ those indicated by the parameters (i.e. the parameters are dropped).
#
# Some examples:
anes.loc[:, ["V161188", "V161204x"]].head() # Selects only "V161188" and "V161204x" columns
incidents.drop(incidents.columns[[0, 1]], axis=1).head() # Drops the columns with indices 0 and 1
incidents.iloc[[1,2,3,5], [1,68]] # Select only columns with indices 1 and 68,
# then only the rows with indices 1, 2, 3, 5
# ** Exercise 2**
#
# To make sure you understand the `loc`, `iloc`, and `drop` functions, try selecting the columns V4002 to V4008 with only the first 3 rows:
# YOUR CODE HERE
incidents.iloc[1:4, 4:10]
# Finally, the `loc` function in the DataFrame can be modified so instead of only choosing certain rows or columns you can give conditions for the selected columns or rows:
# - A column label
# - A condition that each row should match
#
# In other words, we call the select rows as so: `DataFrame_name.loc[DataFrame_name["column_name'] filter]`.
#
#
# Here are some examples of selection:
#
# The variable `V162365` indicates whether or not their is discrimmination against Christians. A value of 1 corresponds to the score of discrimmination. The below query will find all variables of the election where the discrimmination against Christians has a score of 1.
incidents.loc[incidents["V162365"] == 1]
# The variable `V161233x` corresponds to the death penalty. The variable takes values between 1 and 10. With the following where statement, we'll find the variables where the value of death penalty score is between 1 and 10.
anes.loc[anes["V161233x"].isin(np.arange(1, 10))]
# #### Attributes
# Using the methods that we have learned, we can now dive into calculating statistics from data in tables. Two useful _attributes_ (variables, not methods!) of tables are **`index`** and **`columns`**. They store the rows and the columns in a given table, respectively. For example:
num_variables = len(anes.index)
print("Number of rows: ", num_variables)
num_attributes = len(anes.columns)
print("Numbers of columns: ", num_attributes)
# Notice that we do _not_ put `()` after `num_rows` and `num_columns`, as we did for other methods.
# #### Sorting
#
# It can be very useful to sort our DataFrames according to some column. The `sort` function does exactly that; it takes the column that you want to sort by. By default, the `sort_values` function sorts the table in _ascending_ order of the data in the column indicated; however, you can change this by setting the optional parameter `ascending=False`.
#
# Below is an example using the variable, `V161212`, which is the environmental budget.
monetary_loss = anes.loc[anes["V161212"].isin(np.arange(1, 100))]
monetary_loss.sort_values(by=['V161212']) # Sort table by value of property taken in ascending order
# The above code will sort the DataFrame by the column `V161212` from least to greatest. Below, we'll sort it from greatest to least.
monetary_loss.sort_values(by='V161212', ascending=False) # Sort table by value of property taken in descending order (highest at top)
# ## Summary
# As a summary, here are the functions we learned about during this notebook:
#
# |Name|Example|Purpose|
# |-|-|-|
# |`DataFrame`|`DataFrame()`|Create an empty DataFrame, usually to extend with data|
# |`pandas.read_table`|`pandas.read_table("my_data.csv")`|Create a DataFrame from a data file|
# |`with_columns`|`tbl = pandas.DataFrame({"N": np.arange(5), "2*N": np.arange(0, 10, 2)})`|Create a copy of a DataFrame with more columns|
# |`column`|`tbl[["N"]]`|Create an array containing the elements of a column|
# |`sort`|`tbl.sort(["N"])`|Create a copy of a DataFrame sorted by the values in a column|
# |`loc`|`tbl.loc[tbl["N"] > 10]`|Create a copy of a DataFrame with only the rows that match some *predicate*|
# |`index`|`len(tbl.index)`|Compute the number of rows in a DataFrame|
# |`columns`|`len(tbl.columns)`|Compute the number of columns in a DataFrame|
# |`loc`|`tbl.loc["N"]`|Create a copy of a DataFrame with only some of the columns|
# |`drop`|`tbl.drop(columns=["2*N"])`|Create a copy of a DataFrame without some of the columns|
# |`iloc`|`tbl.iloc(np.arange(0, 6, 2))`|Create a copy of the DataFrame with only the rows whose indices are in the given array|
# Some materials this notebook were taken from [Data 8](http://data8.org/), [CS 61A](http://cs61a.org/), and [DS Modules](http://data.berkeley.edu/education/modules) lessons.
|
LEGALST-123/Lab1/lab1_solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="j5duG0Is_knE" outputId="13523ec7-6871-4257-9931-9485245ace21"
pip install tensorflow
# + id="tgRoccRu_nk1"
pip install -q tensorflow tensorflow-datasets
# + id="KvIM5hIz_stE"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
from keras.layers import Dropout,Flatten,Conv2D
import tensorflow_datasets as tfds
# + colab={"base_uri": "https://localhost:8080/"} id="9xlMtEK4ALIo" outputId="6c2a51dc-e1ab-4a69-d3e8-534ec92c82a8"
tfds.list_builders()
# + colab={"base_uri": "https://localhost:8080/"} id="t9pr-MjcAW4P" outputId="30258215-4b2f-4cb7-e185-d170e5aac3dc"
builder=tfds.builder("rock_paper_scissors")
info=builder.info
info
# + id="8J_AX1RbAjY0"
ds_train=tfds.load(name="rock_paper_scissors",split="train")
ds_test=tfds.load(name="rock_paper_scissors",split="test")
# + colab={"base_uri": "https://localhost:8080/"} id="vKz_o4lxA3Vl" outputId="2377f2f8-4673-4cf4-ce9d-fff8ca09033c"
ds_train
# + colab={"base_uri": "https://localhost:8080/"} id="G4R98fNJBZfK" outputId="c229fb09-4ee2-40bc-ae7b-366a74858bab"
info
# + colab={"base_uri": "https://localhost:8080/", "height": 569} id="EX0m8o7vBn_K" outputId="d992529b-b9bb-4d2c-dfc6-ca21c4de33e8"
fig=tfds.show_examples(info,ds_train)
# + id="HYeICviXBt2e"
# + id="fz9sRiXFIX9c"
train_images=np.array([example["image"].numpy()[:,:,0]for example in ds_train])
train_leabels=np.array([example["label"].numpy() for example in ds_train])
test_images=np.array([example["image"].numpy()[:,:,0] for example in ds_test])
test_labels=np.array([example["label"].numpy() for example in ds_test])
# + colab={"base_uri": "https://localhost:8080/"} id="FuJikcxyIYCD" outputId="214aa540-8ec5-4729-ec57-159eaada37a8"
print(type(train_images))
train_images.shape
# + colab={"base_uri": "https://localhost:8080/"} id="xgNRFE4lIYFa" outputId="1d42e125-390c-4e9f-c667-edc61913c1eb"
train_images[0].shape
# + id="BDNuAMPsIYID"
train_images=train_images.reshape(2520,300,300,1)
test_images=test_images.reshape(372,300,300,1)
# + colab={"base_uri": "https://localhost:8080/"} id="8KJ7lZaGIYLV" outputId="a673ef2b-ba08-440d-a3f4-3a0502ccb0f0"
print(train_images)
# + id="DL9vG9aNIYOF"
train_images=train_images.astype("float32")
# + id="17cV_NHeKTVV"
test_images=test_images.astype("float32")
# + colab={"base_uri": "https://localhost:8080/"} id="6-XL0iN8KZ9O" outputId="a782f3b2-609c-4116-fb86-3770771848ea"
print(train_images[0])
# + id="YwHaVB-pKc4-"
train_images=train_images/255
test_images=test_images/255
# + colab={"base_uri": "https://localhost:8080/"} id="r9qncu0SKjb9" outputId="e756b156-a801-49af-f765-2da830b9897e"
print(train_images[0])
# + colab={"base_uri": "https://localhost:8080/"} id="waIMqrEwKmAD" outputId="2404a56b-1ebd-459a-af61-2eda3334be05"
train_images.shape
# + colab={"base_uri": "https://localhost:8080/"} id="Whx3Hgv3K1cd" outputId="c82febae-1fe7-425f-b784-41d786cba87c"
test_images.shape
# + id="AjPRBoNlK33G"
from keras.models import Sequential
# + id="UioOlkXDK_Sf"
from keras.layers import Dense,Dropout,Flatten,Conv2D,MaxPool2D,AveragePooling2D
# + id="hVA8iQAyLJfJ"
from keras.optimizers import Adam
from tensorflow.keras import activations
model=Sequential()
# + id="TRxwlDC2LWZS"
model.add(Flatten())
model.add(Dense(512,input_shape=(300,300,1),activation='relu'))
model.add(Dense(216,activation='relu'))
model.add(Dense(64,activation="softmax"))
model.compile(optimizer="adam",loss="SparseCategoricalCrossentropy",metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} id="pMFahoBzLouk" outputId="3093f2fd-6bb4-4d2a-a149-647ac4ad75d5"
model.fit(train_images,train_labels,epochs=5,batch_size=32)
# + colab={"base_uri": "https://localhost:8080/"} id="4bT9OnvM4GKH" outputId="10bf2467-721d-4a01-a9bd-770066068571"
model.evaluate(test_images,test_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="Z2HRqEnb4ef4" outputId="c467a2a6-24c2-4a59-8bfa-2ed1017b94cf"
model=Sequential()
model.add(AveragePooling2D(6,3,input_shape=(300,300,1)))
model.add(Conv2D(64,3,activation="relu"))
model.add(Conv2D(32,3,activation="relu"))
model.add(MaxPool2D(2,2))
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(128,activation="relu"))
model.add(Dense(3,activation="softmax"))
model.compile(optimizer="adam",loss="SparseCategoricalCrossentropy",metrics=["accuracy"])
model.fit(train_images,train_labels,epochs=5,batch_size=32)
model.evaluate(test_images,test_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="HuBz-OVI409p" outputId="13714eb9-ebcb-4722-dc91-b936584202d2"
model.evaluate(test_images,test_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="5ILzX2UZ-EJv" outputId="02f928e2-28de-4509-d59b-bf4dab4d92a2"
pip install -U keras-tuner
# + colab={"base_uri": "https://localhost:8080/"} id="vVbYWJYl-KpC" outputId="8c4f629e-be53-44e6-c046-6099a35bd3cb"
from kerastuner.tuners import RandomSearch
def build_model(hp):
model = keras.Sequential()
model.add(keras.layers.AveragePooling2D(6,3,input_shape=(300,300,1)))
for i in range(hp.Int("Conv Layers", min_value=0, max_value=3)):
model.add(keras.layers.Conv2D(hp.Choice(f"layer_{i}_filters", [16,32,64]), 3, activation='relu'))
model.add(keras.layers.MaxPool2D(2,2))
model.add(keras.layers.Dropout(0.5))
model.add(keras.layers.Flatten())
model.add(keras.layers.Dense(hp.Choice("Dense layer", [64, 128, 256, 512, 1024]), activation='relu'))
model.add(keras.layers.Dense(3, activation='softmax'))
model.compile(optimizer='adam',
loss=keras.losses.SparseCategoricalCrossentropy(),
metrics=['accuracy'])
return model
tuner = RandomSearch(
build_model,
objective='val_accuracy',
max_trials=32,
)
tuner.search(train_images, train_labels, validation_data=(test_images, test_labels), epochs=10, batch_size=32)
# + id="y_yJWODe-lrB"
best_model = tuner.get_best_models()[0]
# + colab={"base_uri": "https://localhost:8080/"} id="hruKKhsaDJ8m" outputId="4081c46f-7b9d-4049-a4fc-c69b29472ee0"
best_model.evaluate(test_images, test_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="i1MDPXvxDLeF" outputId="e7c0207c-b3b2-4d8f-8e64-3ce7ba8ac65b"
best_model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="5KjQVzI7DNRs" outputId="99b3a022-0ca4-4e4e-adc0-96d382f46744"
tuner.results_summary()
# + id="Mt4SyqZ2DPun"
best_model.save('mymodel')
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="B1UkA_zvDbfI" outputId="44c3a79a-a800-4b17-9ad4-40bef51e9c2f"
image=train_images[0].reshape(300,300)
plt.imshow(image,cmap='Greys_r')
# + colab={"base_uri": "https://localhost:8080/"} id="QjzX65R0DkPR" outputId="f1aae783-9016-4d62-9aa2-e42590e7c59e"
result = best_model.predict(np.array([train_images[0]]))
print(result)
# + colab={"base_uri": "https://localhost:8080/"} id="PU2K8gvjDo7i" outputId="d65ee25a-9a8e-49c5-a525-b47624e02c50"
result=best_model.predict(np.array([train_images[1]]))
print(result)
# + colab={"base_uri": "https://localhost:8080/"} id="2gnq3AfHDxMH" outputId="4a713c51-26da-49b1-cb0f-485508afa628"
predicted_value = np.argmax(result)
print(predicted_value)
# + id="PfMlHgQgDy87"
|
Rock_Paper_Scissors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Frames an Visualization
#
# Setup our data fetching function again
# +
import pandas as pd
def get_nba_data(endpt, params, return_url=False):
## endpt: https://github.com/seemethere/nba_py/wiki/stats.nba.com-Endpoint-Documentation
## params: dictionary of parameters: i.e., {'LeagueID':'00'}
from pandas import DataFrame
from urllib.parse import urlencode
import json
useragent = "\"Mozilla/5.0 (Macintosh; Intel Mac OS X 10_11_2) AppleWebKit/601.3.9 (KHTML, like Gecko) Version/9.0.2 Safari/601.3.9\""
dataurl = "\"" + "http://stats.nba.com/stats/" + endpt + "?" + urlencode(params) + "\""
# for debugging: just return the url
if return_url:
return(dataurl)
# jsonstr = !wget -q -O - --user-agent={useragent} {dataurl}
data = json.loads(jsonstr[0])
h = data['resultSets'][0]['headers']
d = data['resultSets'][0]['rowSet']
return(DataFrame(d, columns=h))
# -
# ## Get one player's shot data
# +
params = {'PlayerID':'201939',
'PlayerPosition':'',
'Season':'2016-17',
'ContextMeasure':'FGA',
'DateFrom':'',
'DateTo':'',
'GameID':'',
'GameSegment':'',
'LastNGames':'0',
'LeagueID':'00',
'Location':'',
'Month':'0',
'OpponentTeamID':'0',
'Outcome':'',
'Period':'0',
'Position':'',
'RookieYear':'',
'SeasonSegment':'',
'SeasonType':'Regular Season',
'TeamID':'0',
'VsConference':'',
'VsDivision':''}
shotdata = get_nba_data('shotchartdetail', params)
shotdata.head()
# -
# ## Drawing the court
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
## code is from http://savvastjortjoglou.com/nba-shot-sharts.html
def draw_court(ax=None, color='black', lw=1, outer_lines=False):
from matplotlib.patches import Circle, Rectangle, Arc
from matplotlib.pyplot import gca
# If an axes object isn't provided to plot onto, just get current one
if ax is None:
ax = gca()
# Create the various parts of an NBA basketball court
# Create the basketball hoop
# Diameter of a hoop is 18" so it has a radius of 9", which is a value
# 7.5 in our coordinate system
hoop = Circle((0, 0), radius=7.5, linewidth=lw, color=color, fill=False)
# Create backboard
backboard = Rectangle((-30, -7.5), 60, 0, linewidth=lw, color=color)
# The paint
# Create the outer box 0f the paint, width=16ft, height=19ft
outer_box = Rectangle((-80, -47.5), 160, 190, linewidth=lw, color=color,
fill=False)
# Create the inner box of the paint, widt=12ft, height=19ft
inner_box = Rectangle((-60, -47.5), 120, 190, linewidth=lw, color=color,
fill=False)
# Create free throw top arc
top_free_throw = Arc((0, 142.5), 120, 120, theta1=0, theta2=180,
linewidth=lw, color=color, fill=False)
# Create free throw bottom arc
bottom_free_throw = Arc((0, 142.5), 120, 120, theta1=180, theta2=0,
linewidth=lw, color=color, linestyle='dashed')
# Restricted Zone, it is an arc with 4ft radius from center of the hoop
restricted = Arc((0, 0), 80, 80, theta1=0, theta2=180, linewidth=lw,
color=color)
# Three point line
# Create the side 3pt lines, they are 14ft long before they begin to arc
corner_three_a = Rectangle((-219, -47.5), 0, 140, linewidth=lw,
color=color)
corner_three_b = Rectangle((219, -47.5), 0, 140, linewidth=lw, color=color)
# 3pt arc - center of arc will be the hoop, arc is 23'9" away from hoop
# I just played around with the theta values until they lined up with the
# threes
three_arc = Arc((0, 0), 475, 475, theta1=22.5, theta2=157.5, linewidth=lw,
color=color)
# Center Court
center_outer_arc = Arc((0, 422.5), 120, 120, theta1=180, theta2=0,
linewidth=lw, color=color)
center_inner_arc = Arc((0, 422.5), 40, 40, theta1=180, theta2=0,
linewidth=lw, color=color)
# List of the court elements to be plotted onto the axes
court_elements = [hoop, backboard, outer_box, inner_box, top_free_throw,
bottom_free_throw, restricted, corner_three_a,
corner_three_b, three_arc, center_outer_arc,
center_inner_arc]
if outer_lines:
# Draw the half court line, baseline and side out bound lines
outer_lines = Rectangle((-250, -47.5), 500, 470, linewidth=lw,
color=color, fill=False)
court_elements.append(outer_lines)
# Add the court elements onto the axes
for element in court_elements:
ax.add_patch(element)
return ax
plt.figure(figsize=(12,11))
draw_court(outer_lines=True)
plt.xlim(-300,300)
plt.ylim(-100,500)
plt.show()
# -
# ## Drawing a scatter plot
# +
plt.figure(figsize=(12,11))
plt.scatter(shotdata.LOC_X, shotdata.LOC_Y)
draw_court(outer_lines=True)
plt.xlim(300,-300)
plt.ylim(-100,500)
plt.show()
# -
# ## Seaborn Package
#
# Seaborn has some statistical functionality built-in: e.g., [linear regression model](https://seaborn.pydata.org/tutorial/regression.html#functions-to-draw-linear-regression-models). Refer to PDSH for a section on [Seaborn plotting package](https://jakevdp.github.io/PythonDataScienceHandbook/04.14-visualization-with-seaborn.html)
#
# Another example is to use Seaborn to draw marginal density functions.
# +
# create our jointplot
joint_shot_chart = sns.jointplot(x="LOC_X", y="LOC_Y", data=shotdata, stat_func=None,
kind='scatter', space=0, alpha=0.5)
joint_shot_chart.fig.set_size_inches(12,11)
# A joint plot has 3 Axes, the first one called ax_joint
# is the one we want to draw our court onto and adjust some other settings
ax = joint_shot_chart.ax_joint
draw_court(ax)
# Adjust the axis limits and orientation of the plot in order
# to plot half court, with the hoop by the top of the plot
ax.set_xlim(-250,250)
ax.set_ylim(422.5, -47.5)
# Get rid of axis labels and tick marks
ax.set_xlabel('')
ax.set_ylabel('')
ax.tick_params(labelbottom=False, labelleft=False)
# Add a title
ax.set_title('<NAME>, 2016-17, FGA',
y=1.2, fontsize=18)
# Add Data Scource and Author
ax.text(-250,445,'Data Source: stats.nba.com'
'\nAuthor: <NAME> (<EMAIL>)',
fontsize=12)
plt.show()
# -
# Another example is to use Seaborn to bin the court into hex-shaped bins.
#
# __Exercise__: Change the plot kind to 'hex'
# ## More about the data
#
# Glossary of terms: https://stats.nba.com/help/glossary/
#
# - FGA: Field Goals Attempted
# - FGM: Field Goals Made
#
# Pandas allows for computing summary statistics similar to tidyverse. The general approach is based on a similar concept.
#
# ## Split-apply-combine
#
# The `groupby` function can be used to create summaries of different groupings. This is often referred to as the split-apply-combine approach.
#
# 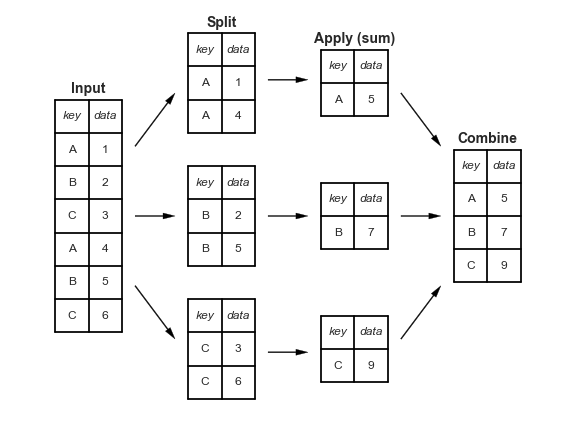
#
# > - The *split* step involves breaking up and grouping a ``DataFrame`` depending on the value of the specified key.
# > - The *apply* step involves computing some function, usually an aggregate, transformation, or filtering, within the individual groups.
# > - The *combine* step merges the results of these operations into an output array.
#
# Recall what the shot data looks like
shotdata.head()
# Let's inspect what the groupped data frame looks like
for r, d in shotdata.groupby('SHOT_ZONE_RANGE'):
print("***", r)
print(d.head())
# ### Split by `SHOT_ZONE_RANGE` column, Apply `mean()`, Combine
shotdata.groupby('SHOT_ZONE_RANGE')['SHOT_MADE_FLAG'].mean()
|
05-Data-Frame-and-Visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 📃 Solution for Exercise M1.02
#
# The goal of this exercise is to fit a similar model as in the previous
# notebook to get familiar with manipulating scikit-learn objects and in
# particular the `.fit/.predict/.score` API.
# Let's load the adult census dataset with only numerical variables
import pandas as pd
adult_census = pd.read_csv("../datasets/adult-census-numeric.csv")
data = adult_census.drop(columns="class")
target = adult_census["class"]
# In the previous notebook we used `model = KNeighborsClassifier()`. All
# scikit-learn models can be created without arguments, which means that you
# don't need to understand the details of the model to use it in scikit-learn.
#
# One of the `KNeighborsClassifier` parameters is `n_neighbors`. It controls
# the number of neighbors we are going to use to make a prediction for a new
# data point.
#
# What is the default value of the `n_neighbors` parameter? Hint: Look at the
# help inside your notebook `KNeighborsClassifier?` or on the [scikit-learn
# website](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsClassifier.html)
# The default value for `n_neighbors` is 5
# Create a `KNeighborsClassifier` model with `n_neighbors=50`
# +
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=50)
# -
# Fit this model on the data and target loaded above
model.fit(data, target)
# Use your model to make predictions on the first 10 data points inside the
# data. Do they match the actual target values?
first_data_values = data.iloc[:10]
first_predictions = model.predict(first_data_values)
first_predictions
first_target_values = target.iloc[:10]
first_target_values
number_of_correct_predictions = (
first_predictions == first_target_values).sum()
number_of_predictions = len(first_predictions)
print(
f"{number_of_correct_predictions}/{number_of_predictions} "
"of predictions are correct")
# Compute the accuracy on the training data.
model.score(data, target)
# Now load the test data from `"../datasets/adult-census-numeric-test.csv"` and
# compute the accuracy on the test data.
# +
adult_census_test = pd.read_csv("../datasets/adult-census-numeric-test.csv")
data_test = adult_census_test.drop(columns="class")
target_test = adult_census_test["class"]
model.score(data_test, target_test)
# -
# Looking at the previous notebook, the accuracy seems slightly higher with
# `n_neighbors=50` than with `n_neighbors=5` (the default value).
|
notebooks/02_numerical_pipeline_sol_00.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] school_cell_uuid="0e4c08e3a79e4b1e8f0744218c2b070d" id="i7IUKfM_q4RW"
# # KoNLPy 한국어 처리 패키지
# + [markdown] school_cell_uuid="143ed59c3e9e4786b58c52996198dfcf" id="Kyf0HEffq4RX"
# KoNLPy(코엔엘파이라고 읽는다)는 한국어 정보처리를 위한 파이썬 패키지이다.
# + school_cell_uuid="b6fe73ca07fd47c7a255b0db442b1502" id="Mt6NhMAUq4RY" colab={"base_uri": "https://localhost:8080/", "height": 398} executionInfo={"status": "ok", "timestamp": 1608091525061, "user_tz": -540, "elapsed": 6175, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="331ce70f-6074-49f2-a393-61f9e4eaaafd"
import warnings
warnings.simplefilter("ignore")
import pandas as pd
import matplotlib.pyplot as plt
# !pip install konlpy
# !pip install WordCloud
pd.set_option('display.max_rows', 80)
plt.rcParams["font.family"] = "NanumGothicCoding"
import konlpy
konlpy.__version__
# + [markdown] school_cell_uuid="240ff9b1ad014f91b179335673cc2005" id="dxVsnoriq4RY"
# ## 한국어 말뭉치
# + [markdown] school_cell_uuid="df3f1c0c41c444339d1dc70c3e586fa2" id="l2i8Oms9q4RZ"
# KoNLPy에서는 대한민국 헌법 말뭉치인 `kolaw`와 국회법안 말뭉치인 `kobill`을 제공한다. 각 말뭉치가 포함하는 파일의 이름은 `fields` 메서드로 알 수 있고 `open` 메서드로 해당 파일의 텍스트를 읽어들인다.
# + school_cell_uuid="7f80a49dfe8b46d486f0db8fe011e301" id="NZALCZZRq4RZ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091525061, "user_tz": -540, "elapsed": 6167, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="016838b6-94b1-4985-81cf-68b04ddb7658"
from konlpy.corpus import kolaw
kolaw.fileids()
# + school_cell_uuid="e1d24940656f4f9f9e6032f7a14056da" id="R5CDowM1q4RZ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091525062, "user_tz": -540, "elapsed": 6160, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="a56f7488-1d4d-4492-f869-a92175c8dec4"
c = kolaw.open('constitution.txt').read()
print(c[:40])
# + school_cell_uuid="34851c3c80d344688cdcb98a3826f325" id="mbbp9rqrq4Ra" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091525063, "user_tz": -540, "elapsed": 6154, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="1a87503f-6586-41d7-8953-61f78291ad89"
from konlpy.corpus import kobill
kobill.fileids()
# + school_cell_uuid="ab2c5be9e19646e8a8cfb97681a115e3" id="9wmwojA8q4Ra" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091525063, "user_tz": -540, "elapsed": 6148, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="d2b5cc9c-60c9-44c9-cdae-e024f13adc2d"
d = kobill.open('1809890.txt').read()
print(d[:40])
# + [markdown] school_cell_uuid="97879e3ed2f34d72b34d1ca880bd6aef" id="p2Al_Y5iq4Ra"
# ## 형태소 분석
# + [markdown] school_cell_uuid="ab1539cc28a240ceb304e9e256573219" id="Cw-DkKrSq4Ra"
# KoNLPy는 다음과 같은 다양한 형태소 분석, 태깅 라이브러리를 파이썬에서 쉽게 사용할 수 있도록 모아놓았다.
#
# * Hannanum: 한나눔. KAIST Semantic Web Research Center 개발.
# * http://semanticweb.kaist.ac.kr/hannanum/
# * Kkma: 꼬꼬마. 서울대학교 IDS(Intelligent Data Systems) 연구실 개발.
# * http://kkma.snu.ac.kr/
# * Komoran: 코모란. Shineware에서 개발.
# * https://github.com/shin285/KOMORAN
# * Mecab: 메카브. 일본어용 형태소 분석기를 한국어를 사용할 수 있도록 수정.
# * https://bitbucket.org/eunjeon/mecab-ko
# * Open Korean Text: 오픈 소스 한국어 분석기. 과거 트위터 형태소 분석기.
# * https://github.com/open-korean-text/open-korean-text
#
# 여기에서는 한나눔, 꼬꼬마, 오픈코리안텍스트 형태소만 예제로 포함하였다.
# + school_cell_uuid="9f10ef1f00f942499939fd3ea672c27f" id="-7ZudM0dq4Rb" executionInfo={"status": "ok", "timestamp": 1608091531725, "user_tz": -540, "elapsed": 12805, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}}
from konlpy.tag import *
hannanum = Hannanum()
kkma = Kkma()
komoran = Komoran()
# mecab = Mecab()
okt = Okt()
# + [markdown] school_cell_uuid="64f3dca5edac4f9085eb91993e846246" id="RjiVxBCLq4Rb"
# 이 클래스들은 다음과 같은 메서드를 공통적으로 제공한다.
#
# * `nouns` : 명사 추출
# * `morphs` : 형태소 추출
# * `pos` : 품사 부착
# + [markdown] school_cell_uuid="fd522673d75e4e7cb5d5f0d340a0d6a5" id="yG6ZGLqpq4Rb"
# ### 명사 추출
# + [markdown] school_cell_uuid="5f60d18cf7ed4c19a069017f142b69ec" id="JADmt0kLq4Rb"
# 문자열에서 명사만 추출하려면 `noun` 명령을 사용한다.
# + school_cell_uuid="d187899ebc5640368a6d33effb710e1d" id="pSrpFVQ3q4Rb" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091534320, "user_tz": -540, "elapsed": 15395, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="635e2095-5a48-40bb-a2b3-f150748ec29a"
hannanum.nouns(c[:40])
# + school_cell_uuid="8828336032f44002af47390b09442426" id="_kUIPvJKq4Rc" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091552216, "user_tz": -540, "elapsed": 33285, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="f4839cc0-a5d8-466b-b319-fab4e6895142"
kkma.nouns(c[:40])
# + school_cell_uuid="d51307e7b39e489fa0a6440806c0cb62" id="qozTPhkfq4Rc" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091552217, "user_tz": -540, "elapsed": 33280, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="50ca83b7-be79-44d1-f5e4-bb90b32df0e3"
# komoran은 빈줄이 있으면 에러가 남
komoran.nouns("\n".join([s for s in c[:40].split("\n") if s]))
# + school_cell_uuid="c40e74f66b3b46f9a3ef73caeea7c641" id="F1hjxhPNq4Rc" executionInfo={"status": "ok", "timestamp": 1608091552218, "user_tz": -540, "elapsed": 33276, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}}
# mecab.nouns(c[:40])
# + school_cell_uuid="c40e74f66b3b46f9a3ef73caeea7c641" id="cZvQOhpsq4Rc" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091565625, "user_tz": -540, "elapsed": 46678, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="514473f6-9df0-42d8-eecc-81c1f0193905"
okt.nouns(c[:40])
# + [markdown] school_cell_uuid="ee183964b2094d64ad17673fe835b0ac" id="8muRwqw-q4Rd"
# ### 형태소 추출
# + [markdown] school_cell_uuid="7d61541c732f4ded9b2d500f0f86312e" id="ZqHistCKq4Rd"
# 명사 뿐 아니라 모든 품사의 형태소를 알아내려면 `morphs`라는 명령을 사용한다.
# + school_cell_uuid="6626659da2234af89ac5b0d5cf9040f1" id="VzpmIYtCq4Rd" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091565625, "user_tz": -540, "elapsed": 46671, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="79bba2e5-411c-4219-f882-966b9f5a4cb0"
hannanum.morphs(c[:40])
# + school_cell_uuid="e54ef02c9ad4433caee92c401ab78965" id="VxHnfL72q4Rd" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091565626, "user_tz": -540, "elapsed": 46667, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="a3e630af-19cd-4c02-819a-d6352f6296fa"
kkma.morphs(c[:40])
# + school_cell_uuid="373b0e41c5c9473ab13ed438ffc09ce9" id="uf4jg__3q4Rd" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091565626, "user_tz": -540, "elapsed": 46661, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="4c9dbf37-e3a6-4952-af5c-38ff6317ef1a"
# komoran은 빈줄이 있으면 에러가 남
komoran.morphs("\n".join([s for s in c[:40].split("\n") if s]))
# + school_cell_uuid="bd4bb9aecf334af5a8bf9c80cbc31d41" id="xioyZAvTq4Re" executionInfo={"status": "ok", "timestamp": 1608091565628, "user_tz": -540, "elapsed": 46657, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}}
# mecab.morphs(c[:40])
# + school_cell_uuid="bd4bb9aecf334af5a8bf9c80cbc31d41" id="QB--9xY_q4Re" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091565628, "user_tz": -540, "elapsed": 46651, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="a50e0822-5b0e-498d-db8f-e0c3ee425876"
okt.morphs(c[:40])
# + [markdown] school_cell_uuid="ea7e1c31c110485f9016f6f29fbf4148" id="hueX9jLxq4Re"
# ### 품사 부착
# + [markdown] school_cell_uuid="babc095b565549e3b9ba71766746f8f6" id="HY85Xewfq4Re"
# `pos` 명령을 사용하면 품사 부착을 한다.
#
# 한국어 품사 태그세트로는 "21세기 세종계획 품사 태그세트"를 비롯하여 다양한 품사 태그세트가 있다. 형태소 분석기마다 사용하는 품사 태그가 다르므로 각 형태소 분석기에 대한 문서를 참조한다.
# + school_cell_uuid="45070e395aa64359b661f7edd2c5de45" id="vABHzlr2q4Re" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091566024, "user_tz": -540, "elapsed": 47041, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="ef366ba2-18ab-4e0c-a3e4-dd6604c146fc"
hannanum.pos(c[:40])
# + school_cell_uuid="e6501f0aeae14c1cbc9eb562fda0ae94" id="S_L9Grz3q4Rf" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091566025, "user_tz": -540, "elapsed": 47036, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="d4a415d8-7d76-45fa-a3a4-c5eca9cb4620"
kkma.pos(c[:40])
# + school_cell_uuid="93d4cc04eb7e4e59a67a33e83bd7b392" id="L9e29c-Pq4Rf" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091566025, "user_tz": -540, "elapsed": 47031, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="153b37fb-6237-4b3a-b857-a2d2dcda9cff"
# komoran은 빈줄이 있으면 에러가 남
komoran.pos("\n".join([s for s in c[:40].split("\n") if s]))
# + school_cell_uuid="e6501f0aeae14c1cbc9eb562fda0ae94" id="DGliHnrnq4Rf" executionInfo={"status": "ok", "timestamp": 1608091566026, "user_tz": -540, "elapsed": 47027, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}}
# mecab.pos(c[:40])
# + school_cell_uuid="e6501f0aeae14c1cbc9eb562fda0ae94" id="uoz6-XCjq4Rf" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091566026, "user_tz": -540, "elapsed": 47022, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="5e94c95c-1d93-48e3-e56b-f3e1c35ba9d2"
okt.pos(c[:40])
# + [markdown] school_cell_uuid="40599a8a056f4de6b03dbf9e58fdda18" id="Cf6_8Ldeq4Rg"
# 부착되는 품사 태그의 기호와 의미는 `tagset` 속성으로 확인할 수 있다.
# + school_cell_uuid="f8b427e35f7946298b71c4c90635757e" id="uBaR40qtq4Rg" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608091566027, "user_tz": -540, "elapsed": 47017, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="00f96a03-a893-4cba-ab75-d7ea6b80ab92"
okt.tagset
# + school_cell_uuid="324ee36b8ea246f0b445bf8dffbc92d3" id="qix1C17Vq4Rg" colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"status": "ok", "timestamp": 1608091568973, "user_tz": -540, "elapsed": 49956, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="2b83fcde-145b-466b-9766-6268813e5958"
tagsets = pd.DataFrame()
N = 67
tagsets["Hannanum-기호"] = list(hannanum.tagset.keys()) + list("*" * (N - len(hannanum.tagset)))
tagsets["Hannanum-품사"] = list(hannanum.tagset.values()) + list("*" * (N - len(hannanum.tagset)))
tagsets["Kkma-기호"] = list(kkma.tagset.keys()) + list("*" * (N - len(kkma.tagset)))
tagsets["Kkma-품사"] = list(kkma.tagset.values()) + list("*" * (N - len(kkma.tagset)))
tagsets["Komoran-기호"] = list(komoran.tagset.keys()) + list("*" * (N - len(komoran.tagset)))
tagsets["Komoran-품사"] = list(komoran.tagset.values()) + list("*" * (N - len(komoran.tagset)))
# tagsets["Mecab-기호"] = list(mecab.tagset.keys()) + list("*" * (N - len(mecab.tagset)))
# tagsets["Mecab-품사"] = list(mecab.tagset.values()) + list("*" * (N - len(mecab.tagset)))
tagsets["OKT-기호"] = list(okt.tagset.keys()) + list("*" * (N - len(okt.tagset)))
tagsets["OKT-품사"] = list(okt.tagset.values()) + list("*" * (N - len(okt.tagset)))
tagsets
# + [markdown] id="5VtDvJ206CE2"
# # 한글 폰트 설치
# + colab={"base_uri": "https://localhost:8080/"} id="uyfbCpyk6Btr" executionInfo={"status": "ok", "timestamp": 1608091571025, "user_tz": -540, "elapsed": 52002, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="7fa3401c-608f-4016-b2ef-8044f2e0f403"
import matplotlib.font_manager as fm
sys_font=fm.findSystemFonts()
print(f"sys_font number: {len(sys_font)}")
nanum_font = [f for f in sys_font if 'Nanum' in f]
print(f"nanum_font number: {len(nanum_font)}")
# !apt install fonts-nanum-coding -qq
sys_font=fm.findSystemFonts()
print(f"sys_font number: {len(sys_font)}")
nanum_font = [f for f in sys_font if 'Nanum' in f]
print(f"nanum_font number: {len(nanum_font)}")
# + colab={"base_uri": "https://localhost:8080/"} id="CwGOT7JUAtrX" executionInfo={"status": "ok", "timestamp": 1608091571026, "user_tz": -540, "elapsed": 51997, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="6a92936c-4770-436d-e1b1-e8373a484962"
import matplotlib.pyplot as plt
def current_font():
print(f"설정 폰트 글꼴: {plt.rcParams['font.family']}, 설정 폰트 사이즈: {plt.rcParams['font.size']}") # 파이썬 3.6 이상 사용가능하다
current_font()
path = '/usr/share/fonts/truetype/nanum/NanumGothicCoding.ttf' # 설치된 나눔글꼴중 원하는 녀석의 전체 경로를 가져오자
font_name = fm.FontProperties(fname=path, size=10).get_name()
print(font_name)
fm._rebuild()
plt.rc('font', family=font_name)
current_font()
# + id="gMtm9l5_A-kF" executionInfo={"status": "ok", "timestamp": 1608091571718, "user_tz": -540, "elapsed": 52684, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}}
import matplotlib as mpl
fm._rebuild()
!
mpl.rcParams['axes.unicode_minus'] = False
# + [markdown] school_cell_uuid="b7034fbcbebb4195a0a3fe7aab6809ae" id="cw-sR97Tq4Rg"
# koNLPy의 형태소 분석기와 NLTK의 Text 클래스를 결합하여 NLTK 기능을 사용할 수도 있다.
# - 코랩에서 오작동 할 수 있음. 오작동 하면 "Restart runtime and run all cells in notebook"
# + school_cell_uuid="b794bec4540b40f4816f32921d7df992" id="l7y-KviOq4Rg" colab={"base_uri": "https://localhost:8080/", "height": 566} executionInfo={"status": "ok", "timestamp": 1608091576236, "user_tz": -540, "elapsed": 57197, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="1e7fd832-2e3f-4ce7-9d0d-27daee0282aa"
from nltk import Text
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (14,10)
kolaw = Text(okt.nouns(c), name="kolaw")
kolaw.plot(30)
plt.show()
# + school_cell_uuid="39f7801606294220ad1e04a45fa43183" id="zMvDThFUq4Rh" colab={"base_uri": "https://localhost:8080/", "height": 482} executionInfo={"status": "ok", "timestamp": 1608091580167, "user_tz": -540, "elapsed": 61119, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}} outputId="df4c9acc-1456-48af-97a2-9f516c14f03e"
from wordcloud import WordCloud
# font_path parameter를 지원하는 함수의 경우
# %cd /content/
font_path = '/content/NanumGothicCoding-Bold.ttf'
![ ! -f {font_path} ]&& wget https://fonts.google.com/download?family=Nanum%20Gothic%20Coding -O /content/NanumGothicCoding.zip
![ ! -f {font_path} ]&& unzip /content/NanumGothicCoding.zip
wc = WordCloud(width = 1000, height = 600, background_color="white", font_path=font_path)
plt.imshow(wc.generate_from_frequencies(kolaw.vocab()))
plt.axis("off")
plt.show()
# + id="3ZH0DXwNrovR" executionInfo={"status": "ok", "timestamp": 1608091580168, "user_tz": -540, "elapsed": 61112, "user": {"displayName": "\ub0a8\uc911\uad6c", "photoUrl": "", "userId": "02914767085172164696"}}
|
2.NLP_and_Preprocessing/KoNLPyPackage.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
from statsmodels.tsa.stattools import adfuller
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from statsmodels.tsa.arima_model import ARMA
import pandas as pd
def perform_adf_test(series):
result = adfuller(series)
print('ADF Statistic: %f' % result[0])
print('p-value: %f' % result[1])
# # Original Series
ts = pd.read_csv('original_series.csv')
ts.index = np.arange(1,len(ts)+1)
# +
plt.figure(figsize=(10,4))
plt.plot(ts)
plt.xticks(np.arange(0,78,6), fontsize=14)
plt.xlabel('Hours Since Published', fontsize=16)
plt.yticks(np.arange(0,50000,10000), fontsize=14)
plt.ylabel('Views', fontsize=16)
# -
# # Original Series: $v_t$
#
# # (1) Normalize ($v_t \rightarrow n_t$): $n_t = \frac{v_t - \mu}{\sigma}$
#
# # (2) Exponentiate ($n_t \rightarrow e_t$): $e_t = e^{n_t}$
#
# # (3) First Difference ($e_t \rightarrow d_t$): $d_t = e_t - e_{t-1}$
#
# # $d_t = e^{\frac{v_t - \mu}{\sigma}} - e^{\frac{v_{t-1} - \mu}{\sigma}}$
#
# # ------------------------------------------------------------
# # (1) Normalize
# +
mu = np.mean(ts).iloc[0]
sigma = np.std(ts).iloc[0]
norm_ts = (ts - mu) / sigma
# +
plt.figure(figsize=(10,4))
plt.plot(norm_ts)
plt.xticks(np.arange(0,78,6), fontsize=14)
plt.xlabel('Hours Since Published', fontsize=16)
plt.yticks(np.arange(-3,2), fontsize=14)
plt.ylabel('Norm. Views', fontsize=16)
plt.axhline(0, color='k', linestyle='--')
# -
# # (2) Exponentiate
exp_ts = np.exp(norm_ts)
# +
plt.figure(figsize=(10,4))
plt.plot(exp_ts)
plt.xticks(np.arange(0,78,6), fontsize=14)
plt.xlabel('Hours Since Published', fontsize=16)
plt.yticks(np.arange(0,3.5,.5), fontsize=14)
plt.ylabel('Exp. Norm. Views', fontsize=16)
# -
perform_adf_test(exp_ts)
# # (3) First Difference
diff_ts = exp_ts.diff().dropna()
# +
plt.figure(figsize=(10,4))
plt.plot(diff_ts)
plt.xticks(np.arange(0,78,6), fontsize=14)
plt.xlabel('Hours Since Published', fontsize=16)
plt.yticks(np.arange(-0.2,0.3,.1), fontsize=14)
plt.ylabel('First Diff. \nExp. Norm. Views', fontsize=16)
# -
perform_adf_test(diff_ts)
# # Fit AR Model
plot_pacf(diff_ts)
plt.show()
plot_acf(diff_ts)
plt.show()
#create the model
model = ARMA(diff_ts, order=(4,1))
model_fit = model.fit()
# # Predict Out 3 Hours
# +
prediction_info = model_fit.forecast(3)
predictions = prediction_info[0]
lower_bound = prediction_info[2][:,0]
upper_bound = prediction_info[2][:,1]
# +
plt.figure(figsize=(10,4))
plt.plot(diff_ts)
plt.xticks(np.arange(0,78,6), fontsize=14)
plt.xlabel('Hours Since Published', fontsize=16)
plt.yticks(np.arange(-0.2,0.3,.1), fontsize=14)
plt.ylabel('First Diff. \nExp. Norm. Views', fontsize=16)
plt.plot(np.arange(len(ts)+1, len(ts)+4), predictions, color='g')
plt.fill_between(np.arange(len(ts)+1, len(ts)+4), lower_bound, upper_bound, color='g', alpha=0.1)
# -
# # Undo Transformations: ($\hat{d}_{t+1} \rightarrow \hat{v}_{t+1}$)
#
# # $\hat{v}_{t+1} = \sigma \ln(\hat{d}_{t+1} + e^{\frac{v_t - \mu}{\sigma}}) + \mu$
def undo_transformations(predictions, series, mu, sigma):
first_pred = sigma*np.log(predictions[0] + np.exp((series.iloc[-1]-mu)/sigma)) + mu
orig_predictions = [first_pred]
for i in range(len(predictions[1:])):
next_pred = sigma*np.log(predictions[i+1] + np.exp((orig_predictions[-1]-mu)/sigma)) + mu
orig_predictions.append(next_pred)
return np.array(orig_predictions).flatten()
orig_preds = undo_transformations(predictions, ts, mu, sigma)
orig_lower_bound = undo_transformations(lower_bound, ts, mu, sigma)
orig_upper_bound = undo_transformations(upper_bound, ts, mu, sigma)
# +
plt.figure(figsize=(10,4))
plt.plot(ts)
plt.xticks(np.arange(0,78,6), fontsize=14)
plt.xlabel('Hours Since Published', fontsize=16)
plt.yticks(np.arange(0,50000,10000), fontsize=14)
plt.ylabel('Views', fontsize=16)
plt.plot(np.arange(len(ts)+1, len(ts)+4), orig_preds, color='g')
plt.fill_between(np.arange(len(ts)+1, len(ts)+4), orig_lower_bound, orig_upper_bound, color='g', alpha=0.1)
# +
plt.figure(figsize=(10,4))
plt.plot(ts)
plt.xticks(np.arange(0,78), fontsize=14)
plt.xlabel('Hours Since Published', fontsize=16)
plt.yticks(np.arange(40000,46000,1000), fontsize=14)
plt.ylabel('Views', fontsize=16)
plt.plot(np.arange(len(ts)+1, len(ts)+4), orig_preds, color='g')
plt.fill_between(np.arange(len(ts)+1, len(ts)+4), orig_lower_bound, orig_upper_bound, color='g', alpha=0.1)
plt.xlim(64,76)
plt.ylim(40000, 45000)
# -
|
time-series-analysis-anomalies/Undo Stationary Transformations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Neural Networks: Application
#
# Welcome to Course 4's second assignment! In this notebook, you will:
#
# - Implement helper functions that you will use when implementing a TensorFlow model
# - Implement a fully functioning ConvNet using TensorFlow
#
# **After this assignment you will be able to:**
#
# - Build and train a ConvNet in TensorFlow for a classification problem
#
# We assume here that you are already familiar with TensorFlow. If you are not, please refer the *TensorFlow Tutorial* of the third week of Course 2 ("*Improving deep neural networks*").
# ### <font color='darkblue'> Updates to Assignment <font>
#
# #### If you were working on a previous version
# * The current notebook filename is version "1a".
# * You can find your work in the file directory as version "1".
# * To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#
# #### List of Updates
# * `initialize_parameters`: added details about tf.get_variable, `eval`. Clarified test case.
# * Added explanations for the kernel (filter) stride values, max pooling, and flatten functions.
# * Added details about softmax cross entropy with logits.
# * Added instructions for creating the Adam Optimizer.
# * Added explanation of how to evaluate tensors (optimizer and cost).
# * `forward_propagation`: clarified instructions, use "F" to store "flatten" layer.
# * Updated print statements and 'expected output' for easier visual comparisons.
# * Many thanks to <NAME> (mentor for the deep learning specialization) for his suggestions on the assignments in this course!
# ## 1.0 - TensorFlow model
#
# In the previous assignment, you built helper functions using numpy to understand the mechanics behind convolutional neural networks. Most practical applications of deep learning today are built using programming frameworks, which have many built-in functions you can simply call.
#
# As usual, we will start by loading in the packages.
# +
import math
import numpy as np
import h5py
import matplotlib.pyplot as plt
import scipy
from PIL import Image
from scipy import ndimage
import tensorflow as tf
from tensorflow.python.framework import ops
from cnn_utils import *
# %matplotlib inline
np.random.seed(1)
# -
# Run the next cell to load the "SIGNS" dataset you are going to use.
# Loading the data (signs)
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
# As a reminder, the SIGNS dataset is a collection of 6 signs representing numbers from 0 to 5.
#
# <img src="images/SIGNS.png" style="width:800px;height:300px;">
#
# The next cell will show you an example of a labelled image in the dataset. Feel free to change the value of `index` below and re-run to see different examples.
# Example of a picture
index = 6
plt.imshow(X_train_orig[index])
print ("y = " + str(np.squeeze(Y_train_orig[:, index])))
# In Course 2, you had built a fully-connected network for this dataset. But since this is an image dataset, it is more natural to apply a ConvNet to it.
#
# To get started, let's examine the shapes of your data.
X_train = X_train_orig/255.
X_test = X_test_orig/255.
Y_train = convert_to_one_hot(Y_train_orig, 6).T
Y_test = convert_to_one_hot(Y_test_orig, 6).T
print ("number of training examples = " + str(X_train.shape[0]))
print ("number of test examples = " + str(X_test.shape[0]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
conv_layers = {}
# ### 1.1 - Create placeholders
#
# TensorFlow requires that you create placeholders for the input data that will be fed into the model when running the session.
#
# **Exercise**: Implement the function below to create placeholders for the input image X and the output Y. You should not define the number of training examples for the moment. To do so, you could use "None" as the batch size, it will give you the flexibility to choose it later. Hence X should be of dimension **[None, n_H0, n_W0, n_C0]** and Y should be of dimension **[None, n_y]**. [Hint: search for the tf.placeholder documentation"](https://www.tensorflow.org/api_docs/python/tf/placeholder).
# +
# GRADED FUNCTION: create_placeholders
def create_placeholders(n_H0, n_W0, n_C0, n_y):
"""
Creates the placeholders for the tensorflow session.
Arguments:
n_H0 -- scalar, height of an input image
n_W0 -- scalar, width of an input image
n_C0 -- scalar, number of channels of the input
n_y -- scalar, number of classes
Returns:
X -- placeholder for the data input, of shape [None, n_H0, n_W0, n_C0] and dtype "float"
Y -- placeholder for the input labels, of shape [None, n_y] and dtype "float"
"""
### START CODE HERE ### (≈2 lines)
X = tf.placeholder(dtype='float',shape=[None, n_H0, n_W0, n_C0])
Y = tf.placeholder(dtype='float',shape=[None, n_y])
### END CODE HERE ###
return X, Y
# -
X, Y = create_placeholders(64, 64, 3, 6)
print ("X = " + str(X))
print ("Y = " + str(Y))
# **Expected Output**
#
# <table>
# <tr>
# <td>
# X = Tensor("Placeholder:0", shape=(?, 64, 64, 3), dtype=float32)
#
# </td>
# </tr>
# <tr>
# <td>
# Y = Tensor("Placeholder_1:0", shape=(?, 6), dtype=float32)
#
# </td>
# </tr>
# </table>
# ### 1.2 - Initialize parameters
#
# You will initialize weights/filters $W1$ and $W2$ using `tf.contrib.layers.xavier_initializer(seed = 0)`. You don't need to worry about bias variables as you will soon see that TensorFlow functions take care of the bias. Note also that you will only initialize the weights/filters for the conv2d functions. TensorFlow initializes the layers for the fully connected part automatically. We will talk more about that later in this assignment.
#
# **Exercise:** Implement initialize_parameters(). The dimensions for each group of filters are provided below. Reminder - to initialize a parameter $W$ of shape [1,2,3,4] in Tensorflow, use:
# ```python
# W = tf.get_variable("W", [1,2,3,4], initializer = ...)
# ```
# #### tf.get_variable()
# [Search for the tf.get_variable documentation](https://www.tensorflow.org/api_docs/python/tf/get_variable). Notice that the documentation says:
# ```
# Gets an existing variable with these parameters or create a new one.
# ```
# So we can use this function to create a tensorflow variable with the specified name, but if the variables already exist, it will get the existing variable with that same name.
#
# +
# GRADED FUNCTION: initialize_parameters
def initialize_parameters():
"""
Initializes weight parameters to build a neural network with tensorflow. The shapes are:
W1 : [4, 4, 3, 8]
W2 : [2, 2, 8, 16]
Note that we will hard code the shape values in the function to make the grading simpler.
Normally, functions should take values as inputs rather than hard coding.
Returns:
parameters -- a dictionary of tensors containing W1, W2
"""
tf.set_random_seed(1) # so that your "random" numbers match ours
### START CODE HERE ### (approx. 2 lines of code)
W1 = tf.get_variable('W1',shape=[4,4,3,8], initializer=tf.contrib.layers.xavier_initializer(seed=0))
W2 = tf.get_variable('W2',shape=[2,2,8,16], initializer=tf.contrib.layers.xavier_initializer(seed=0))
### END CODE HERE ###
parameters = {"W1": W1,
"W2": W2}
return parameters
# -
tf.reset_default_graph()
with tf.Session() as sess_test:
parameters = initialize_parameters()
init = tf.global_variables_initializer()
sess_test.run(init)
print("W1[1,1,1] = \n" + str(parameters["W1"].eval()[1,1,1]))
print("W1.shape: " + str(parameters["W1"].shape))
print("\n")
print("W2[1,1,1] = \n" + str(parameters["W2"].eval()[1,1,1]))
print("W2.shape: " + str(parameters["W2"].shape))
# ** Expected Output:**
#
# ```
# W1[1,1,1] =
# [ 0.00131723 0.14176141 -0.04434952 0.09197326 0.14984085 -0.03514394
# -0.06847463 0.05245192]
# W1.shape: (4, 4, 3, 8)
#
#
# W2[1,1,1] =
# [-0.08566415 0.17750949 0.11974221 0.16773748 -0.0830943 -0.08058
# -0.00577033 -0.14643836 0.24162132 -0.05857408 -0.19055021 0.1345228
# -0.22779644 -0.1601823 -0.16117483 -0.10286498]
# W2.shape: (2, 2, 8, 16)
# ```
# ### 1.3 - Forward propagation
#
# In TensorFlow, there are built-in functions that implement the convolution steps for you.
#
# - **tf.nn.conv2d(X,W, strides = [1,s,s,1], padding = 'SAME'):** given an input $X$ and a group of filters $W$, this function convolves $W$'s filters on X. The third parameter ([1,s,s,1]) represents the strides for each dimension of the input (m, n_H_prev, n_W_prev, n_C_prev). Normally, you'll choose a stride of 1 for the number of examples (the first value) and for the channels (the fourth value), which is why we wrote the value as `[1,s,s,1]`. You can read the full documentation on [conv2d](https://www.tensorflow.org/api_docs/python/tf/nn/conv2d).
#
# - **tf.nn.max_pool(A, ksize = [1,f,f,1], strides = [1,s,s,1], padding = 'SAME'):** given an input A, this function uses a window of size (f, f) and strides of size (s, s) to carry out max pooling over each window. For max pooling, we usually operate on a single example at a time and a single channel at a time. So the first and fourth value in `[1,f,f,1]` are both 1. You can read the full documentation on [max_pool](https://www.tensorflow.org/api_docs/python/tf/nn/max_pool).
#
# - **tf.nn.relu(Z):** computes the elementwise ReLU of Z (which can be any shape). You can read the full documentation on [relu](https://www.tensorflow.org/api_docs/python/tf/nn/relu).
#
# - **tf.contrib.layers.flatten(P)**: given a tensor "P", this function takes each training (or test) example in the batch and flattens it into a 1D vector.
# * If a tensor P has the shape (m,h,w,c), where m is the number of examples (the batch size), it returns a flattened tensor with shape (batch_size, k), where $k=h \times w \times c$. "k" equals the product of all the dimension sizes other than the first dimension.
# * For example, given a tensor with dimensions [100,2,3,4], it flattens the tensor to be of shape [100, 24], where 24 = 2 * 3 * 4. You can read the full documentation on [flatten](https://www.tensorflow.org/api_docs/python/tf/contrib/layers/flatten).
#
# - **tf.contrib.layers.fully_connected(F, num_outputs):** given the flattened input F, it returns the output computed using a fully connected layer. You can read the full documentation on [full_connected](https://www.tensorflow.org/api_docs/python/tf/contrib/layers/fully_connected).
#
# In the last function above (`tf.contrib.layers.fully_connected`), the fully connected layer automatically initializes weights in the graph and keeps on training them as you train the model. Hence, you did not need to initialize those weights when initializing the parameters.
#
#
# #### Window, kernel, filter
# The words "window", "kernel", and "filter" are used to refer to the same thing. This is why the parameter `ksize` refers to "kernel size", and we use `(f,f)` to refer to the filter size. Both "kernel" and "filter" refer to the "window."
# **Exercise**
#
# Implement the `forward_propagation` function below to build the following model: `CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED`. You should use the functions above.
#
# In detail, we will use the following parameters for all the steps:
# - Conv2D: stride 1, padding is "SAME"
# - ReLU
# - Max pool: Use an 8 by 8 filter size and an 8 by 8 stride, padding is "SAME"
# - Conv2D: stride 1, padding is "SAME"
# - ReLU
# - Max pool: Use a 4 by 4 filter size and a 4 by 4 stride, padding is "SAME"
# - Flatten the previous output.
# - FULLYCONNECTED (FC) layer: Apply a fully connected layer without an non-linear activation function. Do not call the softmax here. This will result in 6 neurons in the output layer, which then get passed later to a softmax. In TensorFlow, the softmax and cost function are lumped together into a single function, which you'll call in a different function when computing the cost.
# +
# GRADED FUNCTION: forward_propagation
def forward_propagation(X, parameters):
"""
Implements the forward propagation for the model:
CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED
Note that for simplicity and grading purposes, we'll hard-code some values
such as the stride and kernel (filter) sizes.
Normally, functions should take these values as function parameters.
Arguments:
X -- input dataset placeholder, of shape (input size, number of examples)
parameters -- python dictionary containing your parameters "W1", "W2"
the shapes are given in initialize_parameters
Returns:
Z3 -- the output of the last LINEAR unit
"""
# Retrieve the parameters from the dictionary "parameters"
W1 = parameters['W1']
W2 = parameters['W2']
### START CODE HERE ###
# CONV2D: stride of 1, padding 'SAME'
Z1 = tf.nn.conv2d(X,W1, strides = [1,1,1,1], padding = 'SAME')
# RELU
A1 = tf.nn.relu(Z1)
# MAXPOOL: window 8x8, stride 8, padding 'SAME'
P1 = tf.nn.max_pool(A1, ksize = [1,8,8,1], strides = [1,8,8,1], padding = 'SAME')
# CONV2D: filters W2, stride 1, padding 'SAME'
Z2 = tf.nn.conv2d(P1,W2, strides = [1,1,1,1], padding = 'SAME')
# RELU
A2 = tf.nn.relu(Z2)
# MAXPOOL: window 4x4, stride 4, padding 'SAME'
P2 = tf.nn.max_pool(A2, ksize = [1,4,4,1], strides = [1,4,4,1], padding = 'SAME')
# FLATTEN
F = tf.contrib.layers.flatten(P2)
# FULLY-CONNECTED without non-linear activation function (not not call softmax).
# 6 neurons in output layer. Hint: one of the arguments should be "activation_fn=None"
Z3 = tf.contrib.layers.fully_connected(F, 6, activation_fn=None)
### END CODE HERE ###
return Z3
# +
tf.reset_default_graph()
with tf.Session() as sess:
np.random.seed(1)
X, Y = create_placeholders(64, 64, 3, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
init = tf.global_variables_initializer()
sess.run(init)
a = sess.run(Z3, {X: np.random.randn(2,64,64,3), Y: np.random.randn(2,6)})
print("Z3 = \n" + str(a))
# -
# **Expected Output**:
#
# ```
# Z3 =
# [[-0.44670227 -1.57208765 -1.53049231 -2.31013036 -1.29104376 0.46852064]
# [-0.17601591 -1.57972014 -1.4737016 -2.61672091 -1.00810647 0.5747785 ]]
# ```
# ### 1.4 - Compute cost
#
# Implement the compute cost function below. Remember that the cost function helps the neural network see how much the model's predictions differ from the correct labels. By adjusting the weights of the network to reduce the cost, the neural network can improve its predictions.
#
# You might find these two functions helpful:
#
# - **tf.nn.softmax_cross_entropy_with_logits(logits = Z, labels = Y):** computes the softmax entropy loss. This function both computes the softmax activation function as well as the resulting loss. You can check the full documentation [softmax_cross_entropy_with_logits](https://www.tensorflow.org/api_docs/python/tf/nn/softmax_cross_entropy_with_logits).
# - **tf.reduce_mean:** computes the mean of elements across dimensions of a tensor. Use this to calculate the sum of the losses over all the examples to get the overall cost. You can check the full documentation [reduce_mean](https://www.tensorflow.org/api_docs/python/tf/reduce_mean).
#
# #### Details on softmax_cross_entropy_with_logits (optional reading)
# * Softmax is used to format outputs so that they can be used for classification. It assigns a value between 0 and 1 for each category, where the sum of all prediction values (across all possible categories) equals 1.
# * Cross Entropy is compares the model's predicted classifications with the actual labels and results in a numerical value representing the "loss" of the model's predictions.
# * "Logits" are the result of multiplying the weights and adding the biases. Logits are passed through an activation function (such as a relu), and the result is called the "activation."
# * The function is named `softmax_cross_entropy_with_logits` takes logits as input (and not activations); then uses the model to predict using softmax, and then compares the predictions with the true labels using cross entropy. These are done with a single function to optimize the calculations.
#
# ** Exercise**: Compute the cost below using the function above.
# +
# GRADED FUNCTION: compute_cost
def compute_cost(Z3, Y):
"""
Computes the cost
Arguments:
Z3 -- output of forward propagation (output of the last LINEAR unit), of shape (number of examples, 6)
Y -- "true" labels vector placeholder, same shape as Z3
Returns:
cost - Tensor of the cost function
"""
### START CODE HERE ### (1 line of code)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits = Z3, labels = Y))
### END CODE HERE ###
return cost
# +
tf.reset_default_graph()
with tf.Session() as sess:
np.random.seed(1)
X, Y = create_placeholders(64, 64, 3, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
cost = compute_cost(Z3, Y)
init = tf.global_variables_initializer()
sess.run(init)
a = sess.run(cost, {X: np.random.randn(4,64,64,3), Y: np.random.randn(4,6)})
print("cost = " + str(a))
# -
# **Expected Output**:
# ```
# cost = 2.91034
# ```
# ## 1.5 Model
#
# Finally you will merge the helper functions you implemented above to build a model. You will train it on the SIGNS dataset.
#
# **Exercise**: Complete the function below.
#
# The model below should:
#
# - create placeholders
# - initialize parameters
# - forward propagate
# - compute the cost
# - create an optimizer
#
# Finally you will create a session and run a for loop for num_epochs, get the mini-batches, and then for each mini-batch you will optimize the function. [Hint for initializing the variables](https://www.tensorflow.org/api_docs/python/tf/global_variables_initializer)
# #### Adam Optimizer
# You can use `tf.train.AdamOptimizer(learning_rate = ...)` to create the optimizer. The optimizer has a `minimize(loss=...)` function that you'll call to set the cost function that the optimizer will minimize.
#
# For details, check out the documentation for [Adam Optimizer](https://www.tensorflow.org/api_docs/python/tf/train/AdamOptimizer)
# #### Random mini batches
# If you took course 2 of the deep learning specialization, you implemented `random_mini_batches()` in the "Optimization" programming assignment. This function returns a list of mini-batches. It is already implemented in the `cnn_utils.py` file and imported here, so you can call it like this:
# ```Python
# minibatches = random_mini_batches(X, Y, mini_batch_size = 64, seed = 0)
# ```
# (You will want to choose the correct variable names when you use it in your code).
# #### Evaluating the optimizer and cost
#
# Within a loop, for each mini-batch, you'll use the `tf.Session` object (named `sess`) to feed a mini-batch of inputs and labels into the neural network and evaluate the tensors for the optimizer as well as the cost. Remember that we built a graph data structure and need to feed it inputs and labels and use `sess.run()` in order to get values for the optimizer and cost.
#
# You'll use this kind of syntax:
# ```
# output_for_var1, output_for_var2 = sess.run(
# fetches=[var1, var2],
# feed_dict={var_inputs: the_batch_of_inputs,
# var_labels: the_batch_of_labels}
# )
# ```
# * Notice that `sess.run` takes its first argument `fetches` as a list of objects that you want it to evaluate (in this case, we want to evaluate the optimizer and the cost).
# * It also takes a dictionary for the `feed_dict` parameter.
# * The keys are the `tf.placeholder` variables that we created in the `create_placeholders` function above.
# * The values are the variables holding the actual numpy arrays for each mini-batch.
# * The sess.run outputs a tuple of the evaluated tensors, in the same order as the list given to `fetches`.
#
# For more information on how to use sess.run, see the documentation [tf.Sesssion#run](https://www.tensorflow.org/api_docs/python/tf/Session#run) documentation.
# +
# GRADED FUNCTION: model
def model(X_train, Y_train, X_test, Y_test, learning_rate = 0.009,
num_epochs = 100, minibatch_size = 64, print_cost = True):
"""
Implements a three-layer ConvNet in Tensorflow:
CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED
Arguments:
X_train -- training set, of shape (None, 64, 64, 3)
Y_train -- test set, of shape (None, n_y = 6)
X_test -- training set, of shape (None, 64, 64, 3)
Y_test -- test set, of shape (None, n_y = 6)
learning_rate -- learning rate of the optimization
num_epochs -- number of epochs of the optimization loop
minibatch_size -- size of a minibatch
print_cost -- True to print the cost every 100 epochs
Returns:
train_accuracy -- real number, accuracy on the train set (X_train)
test_accuracy -- real number, testing accuracy on the test set (X_test)
parameters -- parameters learnt by the model. They can then be used to predict.
"""
ops.reset_default_graph() # to be able to rerun the model without overwriting tf variables
tf.set_random_seed(1) # to keep results consistent (tensorflow seed)
seed = 3 # to keep results consistent (numpy seed)
(m, n_H0, n_W0, n_C0) = X_train.shape
n_y = Y_train.shape[1]
costs = [] # To keep track of the cost
# Create Placeholders of the correct shape
### START CODE HERE ### (1 line)
X, Y = create_placeholders(n_H0, n_W0, n_C0, n_y)
### END CODE HERE ###
# Initialize parameters
### START CODE HERE ### (1 line)
parameters = initialize_parameters()
### END CODE HERE ###
# Forward propagation: Build the forward propagation in the tensorflow graph
### START CODE HERE ### (1 line)
Z3 = forward_propagation(X, parameters)
### END CODE HERE ###
# Cost function: Add cost function to tensorflow graph
### START CODE HERE ### (1 line)
cost = compute_cost(Z3, Y)
### END CODE HERE ###
# Backpropagation: Define the tensorflow optimizer. Use an AdamOptimizer that minimizes the cost.
### START CODE HERE ### (1 line)
optimizer = tf.train.AdamOptimizer(learning_rate).minimize(cost)
### END CODE HERE ###
# Initialize all the variables globally
init = tf.global_variables_initializer()
# Start the session to compute the tensorflow graph
with tf.Session() as sess:
# Run the initialization
sess.run(init)
# Do the training loop
for epoch in range(num_epochs):
minibatch_cost = 0.
num_minibatches = int(m / minibatch_size) # number of minibatches of size minibatch_size in the train set
seed = seed + 1
minibatches = random_mini_batches(X_train, Y_train, minibatch_size, seed)
for minibatch in minibatches:
# Select a minibatch
(minibatch_X, minibatch_Y) = minibatch
"""
# IMPORTANT: The line that runs the graph on a minibatch.
# Run the session to execute the optimizer and the cost.
# The feedict should contain a minibatch for (X,Y).
"""
### START CODE HERE ### (1 line)
_ , temp_cost = sess.run([optimizer, cost], feed_dict={X: minibatch_X, Y: minibatch_Y})
### END CODE HERE ###
minibatch_cost += temp_cost / num_minibatches
# Print the cost every epoch
if print_cost == True and epoch % 5 == 0:
print ("Cost after epoch %i: %f" % (epoch, minibatch_cost))
if print_cost == True and epoch % 1 == 0:
costs.append(minibatch_cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
# Calculate the correct predictions
predict_op = tf.argmax(Z3, 1)
correct_prediction = tf.equal(predict_op, tf.argmax(Y, 1))
# Calculate accuracy on the test set
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print(accuracy)
train_accuracy = accuracy.eval({X: X_train, Y: Y_train})
test_accuracy = accuracy.eval({X: X_test, Y: Y_test})
print("Train Accuracy:", train_accuracy)
print("Test Accuracy:", test_accuracy)
return train_accuracy, test_accuracy, parameters
# -
# Run the following cell to train your model for 100 epochs. Check if your cost after epoch 0 and 5 matches our output. If not, stop the cell and go back to your code!
_, _, parameters = model(X_train, Y_train, X_test, Y_test)
# **Expected output**: although it may not match perfectly, your expected output should be close to ours and your cost value should decrease.
#
# <table>
# <tr>
# <td>
# **Cost after epoch 0 =**
# </td>
#
# <td>
# 1.917929
# </td>
# </tr>
# <tr>
# <td>
# **Cost after epoch 5 =**
# </td>
#
# <td>
# 1.506757
# </td>
# </tr>
# <tr>
# <td>
# **Train Accuracy =**
# </td>
#
# <td>
# 0.940741
# </td>
# </tr>
#
# <tr>
# <td>
# **Test Accuracy =**
# </td>
#
# <td>
# 0.783333
# </td>
# </tr>
# </table>
# Congratulations! You have finished the assignment and built a model that recognizes SIGN language with almost 80% accuracy on the test set. If you wish, feel free to play around with this dataset further. You can actually improve its accuracy by spending more time tuning the hyperparameters, or using regularization (as this model clearly has a high variance).
#
# Once again, here's a thumbs up for your work!
fname = "images/thumbs_up.jpg"
image = np.array(ndimage.imread(fname, flatten=False))
my_image = scipy.misc.imresize(image, size=(64,64))
plt.imshow(my_image)
|
Convolutional Neural Networks/Convolution_model_Application_v1a.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ceus
# language: python
# name: ceus
# ---
# # Generate tessellation diagram
#
# Computational notebook 01 for **Morphological tessellation as a way of partitioning space: Improving consistency in urban morphology at the plot scale**.
#
#
# <NAME>., <NAME>., <NAME>. and <NAME>. (2020) _‘Morphological tessellation as a way of partitioning space: Improving consistency in urban morphology at the plot scale’_, Computers, Environment and Urban Systems, 80, p. 101441. doi: [10.1016/j.compenvurbsys.2019.101441](http://doi.org/10.1016/j.compenvurbsys.2019.101441).
#
# Archived version of this repository is stored at the University of Strathclyde KnowledgeBase at DOI [10.15129/c766db26-3fa8-45c6-8218-098d529571fc](https://doi.org/10.15129/c766db26-3fa8-45c6-8218-098d529571fc).
#
#
# Contact: <EMAIL>
#
# Date: 27/03/2020
#
# Note: notebook has been cleaned and released retroactively. It is likely that different versions of packages were initially used, but we made sure that the results remained unaltered.
#
# ---
# **Description**
#
# This notebook generates diagrams illustrating the principles of morphological tessellation used to prepare figures 2, 3 and 4. Figures were later post-processed in Illustrator.
import geopandas as gpd
import momepy as mm
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import scipy as sp
import pandas as pd
from scipy.spatial import Voronoi, voronoi_plot_2d
from random import random
import shapely
from shapely.wkt import loads
from shapely.geometry import Polygon, Point
from tqdm import tqdm
gpd.__version__, mm.__version__, matplotlib.__version__, sns.__version__, np.__version__, sp.__version__, shapely.__version__
polys = [
Polygon([(10, 10), (20, 10), (20, 20), (10, 20)]),
Polygon([(30, 10), (40, 10), (40, 20), (30, 20)]),
Polygon([(10, 30), (20, 30), (20, 40), (10, 40)]),
Polygon([(20, 30), (30, 30), (30, 40), (20, 40)]),
Polygon([(30, 30), (40, 30), (40, 40), (30, 40)]),
]
gdf = gpd.GeoDataFrame(geometry=polys)
gdf['uID'] = mm.unique_id(gdf)
retain = gdf.copy()
sns.set_style('whitegrid')
sns.set_context(context='paper', font_scale=1, rc=None)
f, ax = plt.subplots(figsize=(10, 10))
gdf.plot(ax=ax)
sns.despine(left=True, bottom=True)
#plt.savefig('tesdiag_0.svg')
limit = Polygon([(0, 0), (50, 0), (50, 50), (0, 50)])
f, ax = plt.subplots(figsize=(10, 10))
gdf.plot(ax=ax)
gpd.GeoDataFrame(geometry=[limit.boundary]).plot(ax=ax, edgecolor='red')
sns.despine(left=True, bottom=True)
#plt.savefig('tesdiag_1.svg')
shrink = 1
polys = ["Polygon", "MultiPolygon"]
print("Bufferring geometry...")
gdf["geometry"] = gdf.geometry.apply(
lambda g: g.buffer(-shrink, cap_style=2, join_style=2) if g.type in polys else g
)
f, ax = plt.subplots(figsize=(10, 10))
gdf.plot(ax=ax)
sns.despine(left=True, bottom=True)
#plt.savefig('tesdiag_2.svg')
segment = 2
# +
def _densify(geom, segment):
"""
Returns densified geoemtry with segments no longer than `segment`.
"""
from osgeo import ogr
poly = geom
wkt = geom.wkt # shapely Polygon to wkt
geom = ogr.CreateGeometryFromWkt(wkt) # create ogr geometry
geom.Segmentize(segment) # densify geometry by set metres
geom.CloseRings() # fix for GDAL 2.4.1 bug
wkt2 = geom.ExportToWkt() # ogr geometry to wkt
new = loads(wkt2) # wkt to shapely Polygon
return new
gdf["geometry"] = gdf["geometry"].apply(_densify, segment=segment)
# +
def _point_array(objects, unique_id):
"""
Returns lists of points and ids based on geometry and unique_id.
"""
points = []
ids = []
for idx, row in tqdm(objects.iterrows(), total=objects.shape[0]):
if row["geometry"].type in ["Polygon", "MultiPolygon"]:
poly_ext = row["geometry"].boundary
else:
poly_ext = row["geometry"]
if poly_ext is not None:
if poly_ext.type == "MultiLineString":
for line in poly_ext:
point_coords = line.coords
row_array = np.array(point_coords[:-1]).tolist()
for i, a in enumerate(row_array):
points.append(row_array[i])
ids.append(row[unique_id])
elif poly_ext.type == "LineString":
point_coords = poly_ext.coords
row_array = np.array(point_coords[:-1]).tolist()
for i, a in enumerate(row_array):
points.append(row_array[i])
ids.append(row[unique_id])
else:
raise Exception("Boundary type is {}".format(poly_ext.type))
return points, ids
points, ids = _point_array(gdf, 'uID')
# -
pts = [Point(p) for p in points]
pts = gpd.GeoDataFrame(geometry=pts)
f, ax = plt.subplots(figsize=(10, 10))
pts.plot(ax=ax)
sns.despine(left=True, bottom=True)
#plt.savefig('tesdiag_3.svg')
hull = limit.buffer(100)
hull = _densify(hull, 10)
hull_array = np.array(hull.boundary.coords).tolist()
for i, a in enumerate(hull_array):
points.append(hull_array[i])
ids.append(-1)
voronoi_diagram = Voronoi(np.array(points))
# +
def _regions(voronoi_diagram, unique_id, ids, crs):
"""
Generate GeoDataFrame of Voronoi regions from scipy.spatial.Voronoi.
"""
# generate DataFrame of results
regions = pd.DataFrame()
regions[unique_id] = ids # add unique id
regions["region"] = voronoi_diagram.point_region # add region id for each point
# add vertices of each polygon
vertices = []
for region in regions.region:
vertices.append(voronoi_diagram.regions[region])
regions["vertices"] = vertices
# convert vertices to Polygons
polygons = []
for region in tqdm(regions.vertices, desc="Vertices to Polygons"):
if -1 not in region:
polygons.append(Polygon(voronoi_diagram.vertices[region]))
else:
polygons.append(None)
# save polygons as geometry column
regions["geometry"] = polygons
# generate GeoDataFrame
regions_gdf = gpd.GeoDataFrame(regions.dropna(), geometry="geometry")
regions_gdf = regions_gdf.loc[
regions_gdf["geometry"].length < 1000000
] # delete errors
regions_gdf = regions_gdf.loc[
regions_gdf[unique_id] != -1
] # delete hull-based cells
regions_gdf.crs = crs
return regions_gdf
regions_gdf = _regions(voronoi_diagram, 'uID', ids, crs=gdf.crs)
# -
f, ax = plt.subplots(figsize=(10, 10))
regions_gdf.plot(ax=ax, alpha=.6, edgecolor='white', linewidth=1)
pts.plot(ax=ax)
ax.set_xlim(-10, 60)
ax.set_ylim(-10, 60)
sns.despine(left=True, bottom=True)
#plt.savefig('tesdiag_4.svg')
morphological_tessellation = regions_gdf[['uID', "geometry"]].dissolve(
by='uID', as_index=False
)
f, ax = plt.subplots(figsize=(10, 10))
morphological_tessellation.plot(ax=ax, alpha=.6, edgecolor='white', linewidth=1)
pts.plot(ax=ax)
ax.set_xlim(-10, 60)
ax.set_ylim(-10, 60)
gpd.GeoDataFrame(geometry=[limit.boundary]).plot(ax=ax, edgecolor='red')
sns.despine(left=True, bottom=True)
#plt.savefig('tesdiag_5.svg')
morphological_tessellation = gpd.clip(morphological_tessellation, limit)
f, ax = plt.subplots(figsize=(10, 10))
morphological_tessellation.plot(ax=ax, alpha=.6, edgecolor='white', linewidth=1)
retain.plot(ax=ax)
ax.set_xlim(-10, 60)
ax.set_ylim(-10, 60)
gpd.GeoDataFrame(geometry=[limit.boundary]).plot(ax=ax, edgecolor='red')
sns.despine(left=True, bottom=True)
#plt.savefig('tesdiag_6.svg')
# # Saw-like diagram
polys = [
Polygon([(10, 30), (20, 30), (20, 40), (10, 40)]),
Polygon([(20, 31), (30, 31), (30, 41), (20, 41)]),
Polygon([(30, 30), (40, 30), (40, 40), (30, 40)]),
]
gdf = gpd.GeoDataFrame(geometry=polys)
gdf['uID'] = mm.unique_id(gdf)
retain = gdf.copy()
f, ax = plt.subplots(figsize=(10, 10))
gdf.plot(ax=ax)
sns.despine(left=True, bottom=True)
#plt.savefig('sawdiag_0.svg')
polys = ["Polygon", "MultiPolygon"]
print("Bufferring geometry...")
shrink = 0.3
gdf["geometry"] = gdf.geometry.apply(
lambda g: g.buffer(-shrink, cap_style=2, join_style=2) if g.type in polys else g
)
f, ax = plt.subplots(figsize=(10, 10))
gdf.plot(ax=ax)
sns.despine(left=True, bottom=True)
#plt.savefig('sawdiag_1.svg')
gdf["geometry"] = gdf["geometry"].apply(_densify, segment=segment)
points, ids = _point_array(gdf, 'uID')
pts = [Point(p) for p in points]
pts = gpd.GeoDataFrame(geometry=pts)
f, ax = plt.subplots(figsize=(10, 10))
pts.plot(ax=ax)
sns.despine(left=True, bottom=True)
#plt.savefig('sawdiag_2.svg')
hull = limit.buffer(100)
hull = _densify(hull, 10)
hull_array = np.array(hull.boundary.coords).tolist()
for i, a in enumerate(hull_array):
points.append(hull_array[i])
ids.append(-1)
voronoi_diagram = Voronoi(np.array(points))
regions_gdf = _regions(voronoi_diagram, 'uID', ids, crs=gdf.crs)
f, ax = plt.subplots(figsize=(10, 10))
regions_gdf.plot(ax=ax, alpha=.6, edgecolor='white', linewidth=1)
pts.plot(ax=ax)
ax.set_xlim(9, 41)
ax.set_ylim(29, 42)
sns.despine(left=True, bottom=True)
#plt.savefig('sawdiag_3.svg')
morphological_tessellation = regions_gdf[['uID', "geometry"]].dissolve(
by='uID', as_index=False
)
f, ax = plt.subplots(figsize=(10, 10))
morphological_tessellation.plot(ax=ax, alpha=.6, edgecolor='white', linewidth=1)
retain.plot(ax=ax, alpha=.5)
ax.set_xlim(9, 41)
ax.set_ylim(29, 42)
sns.despine(left=True, bottom=True)
#plt.savefig('sawdiag_4.svg')
shrink = 0.4
segment = 0.5
polys = ["Polygon", "MultiPolygon"]
print("Bufferring geometry...")
gdf["geometry"] = gdf.geometry.apply(
lambda g: g.buffer(-shrink, cap_style=2, join_style=2) if g.type in polys else g
)
gdf["geometry"] = gdf["geometry"].apply(_densify, segment=segment)
points, ids = _point_array(gdf, 'uID')
hull = limit.buffer(100)
hull = _densify(hull, 10)
hull_array = np.array(hull.boundary.coords).tolist()
for i, a in enumerate(hull_array):
points.append(hull_array[i])
ids.append(-1)
voronoi_diagram = Voronoi(np.array(points))
regions_gdf = _regions(voronoi_diagram, 'uID', ids, crs=gdf.crs)
morphological_tessellation = regions_gdf[['uID', "geometry"]].dissolve(
by='uID', as_index=False
)
f, ax = plt.subplots(figsize=(10, 10))
morphological_tessellation.plot(ax=ax, alpha=.6, edgecolor='white', linewidth=1)
retain.plot(ax=ax, alpha=.5)
ax.set_xlim(9, 41)
ax.set_ylim(29, 42)
sns.despine(left=True, bottom=True)
#plt.savefig('sawdiag_5.svg')
# ## Voronoi tessellation illustration
points = np.array(
[[random(), random()] for _ in range(15)])
vor = Voronoi(points)
voronoi_plot_2d(vor)
#plt.savefig("voro_allpts.svg")
|
01_Draw_tessellation_diagrams.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Having a look at the data
# +
# imports
import os
import numpy as np
import warnings # ignore future warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
import pandas as pd
from astropy.table import Table
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm import tnrange, tqdm_notebook
from collections import Counter, OrderedDict
from operator import itemgetter
import cesium
from cesium.time_series import TimeSeries
import cesium.featurize
import schwimmbad
# fron sklearn
import sklearn
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.decomposition import PCA
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import confusion_matrix
import scipy.stats as spstat
# sns.set_palette("Reds")
# %matplotlib inline
# -
DATA_FOLDER = '../../data/'
data_files = ['sample_submission.csv', 'test_set.csv', 'test_set_metadata-csv',
'test_set_sample.csv', 'training_set.csv', 'training_set_metadata.csv']
# read the test sample
test_sample = pd.read_csv(DATA_FOLDER+'test_set_sample.csv', engine='python')
# read the test sample
train = pd.read_csv(DATA_FOLDER+'training_set.csv', engine='python')
train_metadata = pd.read_csv(DATA_FOLDER+'training_set_metadata.csv', engine='python')
test_metadata = pd.read_csv(DATA_FOLDER+'test_set_metadata.csv', engine='python')
test_metadata.describe()
train_metadata.describe()
a = np.array(train['object_id'].value_counts().index)
a
ids_array = np.sort(a)
for id_ in ids_array:
# +
extragal = train_metadata['hostgal_specz'] != 0.
g = sns.jointplot(train_metadata['hostgal_specz'][extragal],\
train_metadata['hostgal_photoz'][extragal], kind='hex',\
xlim=(-0.01, 3.01), ylim=(-0.01,3.01), height=8)
outliers = np.abs(train_metadata['hostgal_specz'] - train_metadata['hostgal_photoz']) > 0.1
fig = g.fig
fig.axes[0].scatter(train_metadata['hostgal_specz'][outliers],\
train_metadata['hostgal_photoz'][outliers], color='C8', alpha=0.3)
fig.tight_layout()
# -
print(train_metadata['target'].value_counts())
train_metadata.head()
# +
metafilename = DATA_FOLDER+'training_set_metadata.csv'
train_metadata = Table.read(metafilename, format='csv')
lcfilename = DATA_FOLDER+'training_set.csv'
lcdata = Table.read(lcfilename, format='csv')
pbmap = OrderedDict([(0,'u'), (1,'g'), (2,'r'), (3,'i'), (4, 'z'), (5, 'Y')])
# it also helps to have passbands associated with a color
pbcols = OrderedDict([(0,'blueviolet'), (1,'green'), (2,'red'),\
(3,'orange'), (4, 'black'), (5, 'brown')])
pbnames = list(pbmap.values())
# +
nobjects = len(train_metadata)
tsdict = OrderedDict()
for i in tnrange(nobjects, desc='Building Timeseries'):
row = train_metadata[i]
thisid = row['object_id']
target = row['target']
meta = {'z':row['hostgal_photoz'],\
'zerr':row['hostgal_photoz_err'],\
'mwebv':row['mwebv']}
ind = (lcdata['object_id'] == thisid)
thislc = lcdata[ind]
pbind = [(thislc['passband'] == pb) for pb in pbmap]
t = [thislc['mjd'][mask].data for mask in pbind ]
m = [thislc['flux'][mask].data for mask in pbind ]
e = [thislc['flux_err'][mask].data for mask in pbind ]
tsdict[thisid] = TimeSeries(t=t, m=m, e=e,\
label=target, name=thisid, meta_features=meta,\
channel_names=pbnames )
del lcdata
# -
# +
features_to_use = ["amplitude",
"percent_beyond_1_std",
"maximum",
"max_slope",
"median",
"median_absolute_deviation",
"percent_close_to_median",
"minimum",
"skew",
"std",
"weighted_average"]
# -
sns.countplot(train_metadata['target'])
test_sample.head()
# # LightCurve analysis
train.head()
# +
array_train = train.values
train.head()
train_objects_ids = np.sort(np.array(train['object_id'].value_counts().index))
# -
array_train[0:5]
train_objects_ids.shape
# train.loc('object_id'==[0])
# for object_id in train_object_ids:
object_id = train_objects_ids[0]
train.loc[train['object_id'] == object_id].groupby(['object_id','passband'])
#.groupby(['object_id','passband']).aggregate(np.sum)
objects = np.zeros
for i, object_id in enumerate(train_object_ids):
array_train[]
class LightCurve(object):
'''Light curve object for PLAsTiCC formatted data'''
_passbands = OrderedDict([(0,'C4'),\
(1,'C2'),\
(2,'C3'),\
(3,'C1'),\
(4,'k'),\
(5,'C5')])
_pbnames = ['u','g','r','i','z','y']
def __init__(self, filename):
'''Read in light curve data'''
self.DFlc = Table.read(filename, format='ascii.csv')
self.filename = filename.replace('.csv','')
self._finalize()
# this is some simple code to demonstrate how to calculate features on these multiband light curves
# we're not suggesting using these features specifically
# there also might be additional pre-processing you do before computing anything
# it's purely for illustration
def _finalize(self):
'''Store individual passband fluxes as object attributes'''
# in this example, we'll use the weighted mean to normalize the features
weighted_mean = lambda flux, dflux: np.sum(flux*(flux/dflux)**2)/np.sum((flux/dflux)**2)
# define some functions to compute simple descriptive statistics
normalized_flux_std = lambda flux, wMeanFlux: np.std(flux/wMeanFlux, ddof = 1)
normalized_amplitude = lambda flux, wMeanFlux: (np.max(flux) - np.min(flux))/wMeanFlux
normalized_MAD = lambda flux, wMeanFlux: np.median(np.abs((flux - np.median(flux))/wMeanFlux))
beyond_1std = lambda flux, wMeanFlux: sum(np.abs(flux - wMeanFlux) > np.std(flux, ddof = 1))/len(flux)
for pb in self._passbands:
ind = self.DFlc['passband'] == pb
pbname = self._pbnames[pb]
if len(self.DFlc[ind]) == 0:
setattr(self, f'{pbname}Std', np.nan)
setattr(self, f'{pbname}Amp', np.nan)
setattr(self, f'{pbname}MAD', np.nan)
setattr(self, f'{pbname}Beyond', np.nan)
setattr(self, f'{pbname}Skew', np.nan)
continue
f = self.DFlc['flux'][ind]
df = self.DFlc['flux_err'][ind]
m = weighted_mean(f, df)
# we'll save the measurements in each passband to simplify access.
setattr(self, f'{pbname}Flux', f)
setattr(self, f'{pbname}FluxUnc', df)
setattr(self, f'{pbname}Mean', m)
# compute the features
std = normalized_flux_std(f, df)
amp = normalized_amplitude(f, m)
mad = normalized_MAD(f, m)
beyond = beyond_1std(f, m)
skew = spstat.skew(f)
# and save the features
setattr(self, f'{pbname}Std', std)
setattr(self, f'{pbname}Amp', amp)
setattr(self, f'{pbname}MAD', mad)
setattr(self, f'{pbname}Beyond', beyond)
setattr(self, f'{pbname}Skew', skew)
# we can also construct features between passbands
pbs = list(self._passbands.keys())
for i, lpb in enumerate(pbs[0:-1]):
rpb = pbs[i+1]
lpbname = self._pbnames[lpb]
rpbname = self._pbnames[rpb]
colname = '{}Minus{}'.format(lpbname, rpbname.upper())
lMean = getattr(self, f'{lpbname}Mean', np.nan)
rMean = getattr(self, f'{rpbname}Mean', np.nan)
col = -2.5*np.log10(lMean/rMean) if lMean> 0 and rMean > 0 else -999
setattr(self, colname, col)
# this is a simple routine to visualize a light curve
# it can plot vs the MJD array of the light curve
# or vs an optional `phase` array that you pass
def plot_multicolor_lc(self, phase=None):
'''Plot the multiband light curve'''
fig, ax = plt.subplots(figsize=(8,6))
if phase is None:
phase = []
if len(phase) != len(self.DFlc):
phase = self.DFlc['mjd']
xlabel = 'MJD'
else:
xlabel = 'Phase'
for i, pb in enumerate(self._passbands):
pbname = self._pbnames[pb]
ind = self.DFlc['passband'] == pb
if len(self.DFlc[ind]) == 0:
continue
ax.errorbar(phase[ind],
self.DFlc['flux'][ind],
self.DFlc['flux_err'][ind],
fmt = 'o', color = self._passbands[pb], label = f'{pbname}')
ax.legend(ncol = 4, frameon = True)
ax.set_xlabel(f'{xlabel}', fontsize='large')
ax.set_ylabel('Flux', fontsize='large')
fig.suptitle(self.filename, fontsize='x-large')
fig.tight_layout(rect=[0, 0, 1, 0.97])
def get_features(self):
'''Return all the features for this object'''
variables = ['Std', 'Amp', 'MAD', 'Beyond', 'Skew']
feats = []
for i, pb in enumerate(self._passbands):
pbname = self._pbnames[pb]
feats += [getattr(self, f'{pbname}{x}', np.nan) for x in variables]
return feats
|
core/luca_data_analysis/luca_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from pulp import *
# +
d = {'Supply_Region':['USA', 'Germany', 'Japan', 'Brazil', 'India'], 'Dmd':[2719.6,84.1,1676.8,145.4,156.4]}
v = {'Supply_Region':['USA', 'Germany', 'Japan', 'Brazil', 'India'],
'USA':[6,13,20,12,22],'Germany':[13,6,14,14,13],'Japan':[20,14,3,21,10],
'Brazil':[12,14,21,8,23], 'India':[17,13,9,21,8]}
f = {'Supply_Region':['USA', 'Germany', 'Japan', 'Brazil', 'India'],
'Low_Cap':[6500,4980,6230,3230,2110], 'High_Cap':[9500,7270,9100,4730,3080]}
p = {'Supply_Region':['USA', 'Germany', 'Japan', 'Brazil', 'India'],
'Low_Cap':[500,500,500,500,500], 'High_Cap':[1500,1500,1500,1500,1500]}
demand = pd.DataFrame(data = d)
demand = demand.set_index('Supply_Region')
var_cost = pd.DataFrame(data = v)
var_cost = var_cost.set_index('Supply_Region')
fix_cost = pd.DataFrame(data = f)
fix_cost = fix_cost.set_index('Supply_Region')
cap = pd.DataFrame(data = p)
cap = cap.set_index('Supply_Region')
print(fix_cost)
print(cap)
print(demand)
# -
# Initialize, Define Decision Vars., and Objective Function
model = LpProblem("Capacitated Plant Location Model", LpMinimize)
loc = ['USA', 'Germany', 'Japan', 'Brazil', 'India']
size = ['Low_Cap','High_Cap']
x = LpVariable.dicts("production", [(i,j) for i in loc for j in loc],
lowBound=0, upBound=None, cat='Continuous')
y = LpVariable.dicts("plant",
[(i,s) for s in size for i in loc], cat='Binary')
# +
model += (lpSum([ fix_cost.loc[i,s] * y[(i,s)] for s in size for i in loc ])
+ lpSum([ var_cost.loc[i,j] * x[(i,j)] for i in loc for j in loc ]))
# Define the constraints
for j in loc:
model += lpSum([x[(i, j)] for i in loc]) == demand.loc[j,'Dmd']
for i in loc:
model += lpSum([x[(i, j)] for j in loc]) <= lpSum([cap.loc[i,s] * y[i,s]
for s in size])
# +
model.solve()
print(LpStatus[model.status])
o = [{'prod':'{} to {}'.format(i,j),'quant':x[(i,j)].varValue} for i in loc for j in loc]
print(pd.DataFrame(o))
o = [{'loc':i, 'lc':y[(i,size[0])].varValue, 'hc':y[(i,size[1])].varValue} for i in loc]
print(pd.DataFrame(o))
print("Objective = $", value(model.objective))
o = [{'name':name, 'shadow_price':c.pi, 'slack':c.slack} for name, c in model.constraints.items()]
print(pd.DataFrame(o))
# -
model
|
PULP/tutorial/4.34 Capacitated plant location p3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > Note: All code used in this notebook is contained in the [notebooks/beaconrunner2050](https://github.com/barnabemonnot/beaconrunner/tree/master/notebooks/beaconrunner2050) folder of the Beacon runner repo, and does not use current eth2.0 specs. Most of the content here remains applicable.
#
# ## TL;DR
#
# - We improve upon our second [_Beacon Runner 2049_](../beaconrunner2049/br2049.html), an economics-focused simulation environment for eth2.
# - Validators are placed on an [asynchronous](https://decentralizedthoughts.github.io/2019-06-01-2019-5-31-models/) P2P network, with each validator storing their current view of the chain, incoming blocks and attestations in a `Store` object, as defined in the [specs](https://github.com/ethereum/eth2.0-specs/blob/dev/specs/phase0/fork-choice.md#store).
# - Validator behaviours are fully modular and can be plugged into the simulation as long as they follow a simple API, to produce rich agent-based models. Try it yourself!
#
# ---
#
# We want to understand how validator behaviours map to rewards, penalties and chain outcomes. Ideally, validators who are rational are also honest, i.e., they run the eth2 protocol the way it "should" be run. But apart from how incentives are designed, there is no guarantee that this will indeed the case. And as we will see, honesty may not always have a unique instantiation.
#
# In this notebook, we improve upon the [first](../beaconrunner/br.html) and [second](../beaconrunner2049/br2049.html) Beacon Runners by introducing a more full-fledged simulation environment. If you haven't read the previous two, that's OK! Let's catch up now.
#
# ## _Previously, on..._
#
# In the [first notebook](../beaconrunner/br.html), we introduced the possibility of "wrapping" the current specs in a [cadCAD](https://github.com/BlockScience/cadCAD) simulation environment. We defined simple _strategies_ for validators that allowed them to produce blocks and attest too. The implementation was centralised in the sense that all validators shared a common view of the chain at all times -- a situation akin to being on a network with _perfect information_ and _zero latency_.
#
# The natural next step was to relax this assumption, and allow different views of the chain to coexist. In the simplest case, these views have an empty intersection: this is the case when the network is perfectly _partitioned_, and each side of the partition works independently. [We explored in the second notebook](../beaconrunner2049/br2049.html) how the _inactivity leak_, which decreases the stake of inactive validators, eventually allows for finalisation to resume. But what if this intersection is not empty? In other words, what if some validators see both sides of the partition? More generally, what if each validator has their own view of the chain, governed by the messages they have received from other validators on the network?
#
# These are the conditions we explore here. They are sufficient to represent a realistic p2p network, where validators receive updates from each other after some (random) delay. We'll reuse the network model introduced in the previous notebook, reintroduced in the next section with a brief introduction to the validator API.
#
# ## Getting started
#
# [Once again](../beaconrunner2049/br2049.html), we import the specs loaded with a custom configuration file, `fast`, where epochs are only 4 slots long (for demonstration purposes).
# +
import specs
import importlib
from eth2spec.config.config_util import prepare_config
from eth2spec.utils.ssz.ssz_impl import hash_tree_root
prepare_config(".", "fast.yaml")
importlib.reload(specs)
# -
# We import our network library, seen in [network.py](https://github.com/ethereum/beaconrunner/tree/master/notebooks/beaconrunner2050/network.py), as well as a library of helper functions for our Beacon Runners, [brlib.py](https://github.com/ethereum/beaconrunner/tree/master/notebooks/beaconrunner2050/brlib.py). Open them up! The code is not that scary.
import network as nt
import brlib
# Now on to the new stuff. We've moved `honest_attest` and `honest_propose` to a new [validatorlib.py](https://github.com/ethereum/beaconrunner/tree/master/notebooks/beaconrunner2050/validatorlib.py) file. This file also defines a very important class, the `BRValidator`, intended to be an abstract superclass to custom validator implementations. `BRValidator` comes packaged with a `Store`, a nifty little helper class defined in the [specs](https://github.com/ethereum/eth2.0-specs/blob/dev/specs/phase0/fork-choice.md#store) and a bunch more logic to record past actions and current parameters. We'll get to them in a short while.
#
# We intend `BRValidator` to be an abstract superclass, meaning that though it is not supposed to be instantiated, it is friendly to inheritance. Subclasses of `BRValidator` inherit its attributes and methods, and are themselves intended to follow a simple API. Subclasses of `BRValidator` must expose a `propose()` and an `attest()` method which return, respectively, a block or an attestation when queried (or `None` when they are shy and don't want to return anything yet). We provide an example implementation in [ASAPValidator.py](https://github.com/ethereum/beaconrunner/tree/master/notebooks/beaconrunner2050/ASAPValidator.py), a very nice validator who always proposes and attests as soon as they can, and honestly too.
import validatorlib as vlib
from ASAPValidator import *
# Let's talk about cadCAD once more. Our simulations are now stochastic, since the latency of the network means that some updates are random. cadCAD makes it easy to organise and run any number of instances as well as define the steps that take place in each instance. But our simulation state is pretty large: there are _n_ validators and for each validator, a certain amount of data to keep track of, including chain states and current unincluded blocks and attestations. So we are using [radCAD](https://github.com/cadCAD-edu/radCAD), a cadCAD extension which supports disabling deep copies of the state at each step, in addition to performance improvements.
# We can now import the radCAD classes and methods we'll use here.
# +
from radcad import Model, Simulation, Experiment
from radcad.engine import Engine, Backend
import pandas as pd
# -
# Are we all set? It seems so!
#
# ## Discovering the validator and network APIs
#
# We'll start slow, as we have done in previous notebooks, before moving to a bigger simulation. We loaded a specs configuration with 4 slots per epoch, so we'll instantiate 4 `ASAPValidator`s, one to attest in each slot.
#
# ### Genesis
#
# First, we obtain a genesis state with 4 deposits registered. Second, we instantiate our validators from this state. A `Store` is created in each of them that records the genesis state root and a couple other things. Finally we ask our validators to skip the genesis block -- it is a special block at slot 0 that no one is supposed to suggest. The first block from a validator is expected at slot 1.
genesis_state = brlib.get_genesis_state(4, seed="riggerati")
validators = [ASAPValidator(genesis_state, i) for i in range(4)]
brlib.skip_genesis_block(validators)
# Note that the current store time is exactly `SECONDS_PER_SLOT` ahead of `genesis_time` (in our configuration, and the current canonical specs, 12 seconds). We've fast-forwarded beyond the first block at 0 to the start of slot 1.
print("Genesis time =", validators[0].store.genesis_time, "seconds")
print("Store time =", validators[0].store.time, "seconds")
print("Current slot =", validators[0].data.slot)
# Let's now reuse the network we had in [the second notebook](../beaconrunner2049/br2049.html). The four validators are arranged along a chain, `0` peering with `1`, `1` peering with `2`, and `2` peering with `3`. We create information sets (who peers with who) to represent the chain.
#
# 
# +
set_a = nt.NetworkSet(validators=list([0,1]))
set_b = nt.NetworkSet(validators=list([1,2]))
set_c = nt.NetworkSet(validators=list([2,3]))
net = nt.Network(validators = validators, sets = list([set_a, set_b, set_c]))
# -
# ### Proposer duties
#
# When we instantiate new validators, as we have done with `ASAPValidator(genesis_state, validator_index)`, their constructor preloads a few things. First, each validator checks their proposer duties for all slots of the current epoch.
proposer_views = [(validator_index, validator.data.current_proposer_duties) \
for validator_index, validator in enumerate(net.validators)]
proposer_views
# The array above shows for each validator index (0, 1, 2, 3) whether they are expected to propose a block in either of the 4 slots. Notice that the randomness means the same validator could be called twice in an epoch. This is distinct from attestation duties, where each validator is expected to attest once, and only once, in each epoch.
#
# Since we are at slot 1, we see that validator 1 is expected to propose here. Let's ping them by calling their `propose` method, which expects a dictionary of "known items": blocks and attestations communicated over the network which may not yet have been included in the chain.
#
# Let's take a brief look at the `propose` method of the ASAP validator:
#
# ```python
# def propose(self, known_items) -> Optional[specs.SignedBeaconBlock]:
# # Not supposed to propose for current slot
# if not self.data.current_proposer_duties[self.data.slot % specs.SLOTS_PER_EPOCH]:
# return None
#
# # Already proposed for this slot
# if self.data.last_slot_proposed == self.data.slot:
# return None
#
# # honest propose
# return vlib.honest_propose(self, known_items)
#
# ```
# Each validator has a `data` attribute, its "internals", maintained and updated by the `BRValidator` class. By exposing the current slot, whether the validator is supposed to propose (in `current_proposer_duties`) and whether the validator proposed something already (`last_slot_proposed`), one can build fairly sophisticated strategies already.
#
# Validator 1 is the first to do anything here, so we'll leave the `known_items` attributes empty.
block = net.validators[1].propose({ "attestations": [], "blocks": [] })
print("There are", len(block.message.body.attestations), "attestations in the block")
# Unsurprisingly, the new block produced does not contain any attestations.
#
# Now validator 1 communicates its block to the information sets it belongs to. Since it belongs to $\{ 0, 1 \}$ and $\{ 1, 2 \}$, validators 0 and 2 receive the block at this point.
#
# 
nt.disseminate_block(net, 1, block)
# In addition to the `data` attribute, our validators maintain a `store` which records current beacon chain states. We can access the blocks in those states or the states themselves from the hash of the block. Let's check if validators 0 and 3 have recorded the current block.
# +
block_root = hash_tree_root(block.message)
try:
net.validators[0].store.blocks[block_root]
print("0: there is a block")
except KeyError: print("0: no block")
try:
net.validators[3].store.blocks[block_root]
print("3: there is a block")
except KeyError: print("3: no block")
# -
# This confirms that validator 3 has not seen the block yet. In the next network update, triggered when `update_network` is called on the current `network` object, validator 2 communicates the block to validator 3. But let's not do that just yet, and instead fast-forward a little more to slot number 2.
for validator in net.validators:
validator.forward_by(specs.SECONDS_PER_SLOT)
print("Validator 0 says this is slot number", net.validators[0].data.slot)
# ### Attester duties
#
# Let's check who is expected to attest at slot 2. Our `BRValidator` superclass records the slot of the current epoch where validators are expected to attest (their _committee slot_), in the `current_attest_slot` attribute of their `data`. In general, computing attester or proposer duties is expensive, so we try to cache it when we can and recompute it only when necessary.
committee_slots = [validator.data.current_attest_slot for validator in net.validators]
pd.DataFrame({ "validator_index": [0, 1, 2, 3], "committee_slot": committee_slots})
# According to the schedule here, at slot 2, validator 2 is expected to attest. Let's check what items they currently know about.
known_items = nt.knowledge_set(net, 2)
print("Validator 2 knows", len(known_items["blocks"]), "block")
print("This block was proposed by validator", known_items["blocks"][0].item.message.proposer_index,
"in slot", known_items["blocks"][0].item.message.slot)
# Validator 2 knows about the block that validator 1 sent in slot 1! All is well here. Validator 2's attestation will set this block as the current head of the chain and the heat goes on.
attestation = net.validators[2].attest(known_items)
print(attestation)
# Woah, what happened here? Validator 2 refused to attest.
#
# Let's back up a bit and see why. Validator 2 is expected to attest during slot 2. Honest validators however are supposed to leave a bit of time for block proposers to communicate their blocks. We are indeed in slot 2, but we are early into it, at the very start. Meanwhile, slots last for about 12 seconds and validators are only expected to attest a third of the way into the slot, i.e., 4 seconds in. This leaves 4 seconds for the block proposer of slot 2 to produce their block and communicate it (in reality, a bit more since producers can start producing before the end of the previous slot, at the risk of missing incoming attestations).
#
# We can also look at the `attest` code in `ASAPValidator` to see this:
#
# ```python
# def attest(self, known_items) -> Optional[specs.Attestation]:
# # Not the moment to attest
# if self.data.current_attest_slot != self.data.slot:
# return None
#
# # Too early in the slot
# if (self.store.time - self.store.genesis_time) % specs.SECONDS_PER_SLOT < 4:
# return None
#
# # Already attested for this slot
# if self.data.last_slot_attested == self.data.slot:
# return None
#
# # honest attest
# return vlib.honest_attest(self, known_items)
# ```
#
# Alright. Let's assume that no one wants to propose anything yet for this slot. We'll forward everyone by 4 seconds and see if validator 2 is ready to attest then.
for validator in net.validators:
validator.forward_by(4)
print("Validator 2 says this is slot number", net.validators[2].data.slot)
print("Time is now", net.validators[2].store.time)
print("We are now",
(net.validators[2].store.time - net.validators[2].store.genesis_time) % specs.SECONDS_PER_SLOT, "seconds into the slot")
# Ready to attest now?
attestation = net.validators[2].attest(known_items)
print(attestation)
# Yes! Validator 2 returned a well-formed attestation. ASAP validators are in a hurry, sure, but not in such a hurry that they would attest too early in the slot.
#
# _What are the dangers of attesting too early?_ Simply put, validators are rewarded for attesting to the correct head of the chain from the perspective of their committee slot. Suppose a validator is expected to attest for slot 2. They send their attestation out before receiving the block for slot 2. In their attestation, they attested for the head of the chain they knew about, i.e., the block at slot 1. Later on, the block for slot 2 is included in the canonical chain. The head in the attestation is now incorrect!
#
# 
#
# _But shouldn't a validator attest as late as possible then?_ We are dipping our toes in the waters of game theory here. I like it. Maybe! Though attesting too late means that the reward obtained for being included early decreases, and if you are _really_ too late, like an epoch late, then you cannot be included at all. So pick your poison here.
#
# We'll define (or you can try it yourself!) a different validator behaviour, attesting _as soon as a block is received in the slot, or slightly before the end of the slot if no block comes in_. This is much unlike the current validator, who attests four seconds in no matter what. Let's look at this in [a different notebook](../thunderdome/thunderdome.html)!
# We need to forward a bit more for validators to record the new attestation. By default, validators ignore incoming attestations for the slot they are currently in. This is because an attestation for slot 2 can _at the earliest_ be included in a block for slot 3. So let's jump to slot 3 by forwarding by 8 seconds.
# +
for validator in net.validators:
validator.forward_by(8)
print("Validator 2 says this is slot number", net.validators[2].data.slot)
# -
# Let's have validator 2 disseminate their attestation. In the next section we'll see how other validators react to it.
#
# 
nt.disseminate_attestations(net, [(2, attestation)])
# ### Final state
#
# We'll check the state of each validator in turn. The `store` records in its `latest_messages` attribute the latest message received from every other validator (message being "attestation" here). This is the _LMD_ of _LMD_-GHOST, Latest Message-Driven fork choice!
print(net.validators[0].store.latest_messages)
# Validator 0 has an empty `latest_messages` attribute. Remember that validator 0 is not peering with validator 2. Since the network was not updated, the recent attestation from validator 2 did not make its way to validator 0.
print(net.validators[1].store.latest_messages)
# Validator 1 has seen the attestation from validator 2, since they are peering together. This makes sense.
print(net.validators[2].store.latest_messages)
# Obviously, validator 2 also knows about its own attestation.
print(net.validators[3].store.latest_messages)
# Hmm, this is trickier. Validator 3 received validator 2's attestation, since they are peering together. But why isn't it showing here in the `latest_messages`?
#
# The reason is simple: validator 2's attestation vouches for _validator 1's block_ as the current head of the chain. But validator 3 doesn't yet know about this block! From the point of view of validator 3, the attestation might as well be vouching for an nonexistent head. In our `net` object, the attestation is recorded as "known" by validator 3, but it cannot participate in validator 3's fork choice, until validator 3 knows about validator 1's block.
#
# Now that we have some intuition for what’s going on behind the scenes, let’s take a look at a larger-scale simulation!
# ## Simulating a complete chain
#
# First up, we need to reload our libraries as we'll be using a different specs configuration; the `medium` config now has 16 slots per epoch (see the [second notebook](../beaconrunner2049/br2049.html) where we used the same configuration).
# +
prepare_config(".", "medium.yaml")
importlib.reload(specs)
importlib.reload(nt)
importlib.reload(brlib)
importlib.reload(vlib)
from ASAPValidator import *
# -
# We'll start with 100 validators, divided into two sets, with a small overlap.
# +
num_validators = 100
genesis_state = brlib.get_genesis_state(num_validators)
validators = [ASAPValidator(genesis_state.copy(), validator_index) for validator_index in range(num_validators)]
brlib.skip_genesis_block(validators)
set_a = nt.NetworkSet(validators=list(range(0, int(num_validators * 2 / 3.0))))
set_b = nt.NetworkSet(validators=list(range(int(num_validators / 2.0), num_validators)))
network = nt.Network(validators = validators, sets=list([set_a, set_b]))
print("Set A = ", set_a)
print("Set B = ", set_b)
# -
# Notice that validators 50 to 65 belong to both sets. If the intersection was completely empty, we'd be back to the partition case we saw in the [previous notebook](../beaconrunner2049/br2049.html).
#
# Same as before, we set our `initial_conditions` to only contain the `network` object we just defined.
initial_conditions = {
'network': network
}
# How does the simulation proceed? We change the rules significantly here. In previous notebooks, we kept the pattern one simulation step = one slot. But to model the effects of network latency or timeliness of validator moves, this is not fine-grained enough. A `frequency` parameter (in Hertz) controls _how many times per second_ we update the simulation.
#
# Here is what happens at each update:
#
# 1. **(Policy)** All validators are queried to check if they want to attest at this time.
# 2. **(State update)** If attestations were made, we disseminate them over the network.
# 3. **(Policy)** All validators are queried to check if they want to propose a block at this time.
# 4. **(State update)** If blocks were proposed, we disseminate them over the network.
# 5. **(State update)** We call `tick` to move the clock by one step (= a second if `frequency` is 1, a tenth of a second if `frequency` is 10 etc). When `tick` moves the clock past the start of a new slot, validators update their internals, checking for instance their new attester or proposer duties if this tick coincides with a new epoch.
#
# Whenever `tick` is called, we also check whether we want the network to update or not, by flipping a biased coin. By "updating the network", we mean "peers exchange messages". In the chain example above, with 4 validators arranged as 0 <-> 1 <-> 2 <-> 3, it takes two network updates for a message from validator 3 to reach validator 0 (when validator 3 sends their message, we assume that it reaches all their peers instantly).
#
# The update frequency of the network is represented by the `network_update_rate` simulation parameter, also in Hertz. A `network_update_rate` of 1 means that messages spread one step further on the network each second.
params = {
"frequency": [1],
"network_update_rate": [1]
}
# As usual, we have our partial state update blocks which represent the substep in the simulation steps.
psubs = [
{
'policies': {
'action': brlib.attest_policy # step 1
},
'variables': {
'network': brlib.disseminate_attestations # step 2
}
},
{
'policies': {
'action': brlib.propose_policy # step 3
},
'variables': {
'network': brlib.disseminate_blocks # step 4
}
},
{
'policies': {
},
'variables': {
'network': brlib.tick # step 5
}
},
]
# We'll now set the parameters for our run. We want to run it for `number_slots`, meaning that we need `steps` timesteps, as given by the formula below. Notice that we feed our `parameters` dictionary to the `M` key of `simulation_parameters`. This exposes the `frequency` and `network_update_rate` parameters to all state update functions in our simulation (here we only use it for `tick`, which updates the clock of all validators and potentially the network too).
# +
number_epochs = 6
number_slots = number_epochs * specs.SLOTS_PER_EPOCH
steps = number_slots * specs.SECONDS_PER_SLOT * params["frequency"][0]
print("will simulate", number_epochs, "epochs (", number_slots, "slots ) at frequency", vlib.frequency, "moves/second")
print("total", steps, "simulation steps")
# -
# One last thing: we discussed before the use of a cadCAD fork that doesn't record a complete copy of the simulation state at each step. This is critical because when we set a very high `frequency`, during many steps nothing really happens: no one is proposing or attesting, but we still should ping validators to check if they want to do either. Recording the full state every step is quite wasteful! So instead, we'll define _observers_, or _metrics_: functions of the state that record a simple value at each step, such as the average balance of validators or the current slot. Let's write an observer for the current slot first:
current_slot = lambda s: s["network"].validators[0].data.slot
# That was quite easy. Our state only includes the `network` object and since we assume all validators share a clock (or at least they are all synced to the same time) any validator's current slot will do.
#
# Now let's think about how to get the average balance. Of course, this depends on _which_ beacon chain state we are looking at. Each validator maintains their own current state, which is made up of all the blocks and attestations they have seen until now. All validators may not agree on the current balances of everyone! In the extreme case of a partition, which we discussed in the [previous notebook](../beaconrunner2049/br2049.html), the two sides of the partition had completely different accounts of the current distribution.
#
# 
# There is no one single correct answer. If we believe our network is fully connected (i.e., no partition) with reasonable latency, it follows that all validators eventually receive any given message in a reasonable amount of time. Under these assumptions, to get a good idea of the distribution (of average balances) over all validators, it's probably enough to sample the distribution of any one validator.
# +
from eth2 import gwei_to_eth
def average_balance(state):
validator = state["network"].validators[0]
head = specs.get_head(validator.store)
current_state = validator.store.block_states[head]
current_epoch = specs.get_current_epoch(current_state)
number_validators = len(current_state.balances)
return gwei_to_eth(float(sum(current_state.balances)) / float(number_validators))
# -
# We now have a couple of custom functions to add our two observers, `current_slot` and `average_balance`, to the simulation proceedings. In the background, we record the current slot and the average balance in the state of the simulation, so we need to add them to the initial conditions as well as to the state update blocks defined above.
# +
from cadCADsupSUP import *
observers = {
"current_slot": current_slot,
"average_balance": average_balance
}
observed_ic = get_observed_initial_conditions(initial_conditions, observers)
observed_psubs = get_observed_psubs(psubs, observers)
# -
# Let's run it!
# +
# %%capture
model = Model(
initial_state=observed_ic,
state_update_blocks=observed_psubs,
params=params,
)
simulation = Simulation(model=model, timesteps=steps, runs=1)
experiment = Experiment([simulation])
experiment.engine = Engine(deepcopy=False, backend=Backend.SINGLE_PROCESS)
result = experiment.run()
df = pd.DataFrame(result)
# -
# This takes a little time (despite a lot of caching behind the scenes), but executes and returns a simulation transcript with our observers, in `df`.
df.head()
# Let's plot the average balance over time, taking slots as our time unit.
df.plot("current_slot", "average_balance")
# It increases! Remember that our epochs are 16 slots long here. Validators are behaving well, the network latency is small enough that no message is delayed too much, it's all good!
# ## Arrival
#
# So what have we done here?
#
# - We have a group of validators who are all individually keeping track of their view of the beacon state.
# - Validators are on a network and communicate with each other, keeping track of known (but perhaps unincluded) attestations and blocks.
# - Whenever something changes, e.g., the beginning of a new slot/epoch or a new block/attestation received on-the-wire (as in, from the p2p network), validators update their internals to parameterise their strategies.
# - Each step, the simulation pings all validators and asks whether they want to propose or attest at this point in time. Based on their current data and past actions, they either return a block and/or an attestation, or nothing.
#
# Now that we have this nice playground for validators to roam around, we are getting close to a full-fledged _agent-based model_. But we need more agents! Who expects here that all validators will be ASAP always? And is ASAP the only "good" behaviour? Probably not!
#
# We'll use the framework developed in this notebook to explore these questions, with a series of smaller "case studies" looking at specific questions. Note that our simulation environment is still incomplete: validators should do more than just proposing and attesting.
#
# - Up until now we assumed proposers were taking on the responsibility of aggregating their known attestations to record them in their proposed blocks. In eth2, aggregators are functionally different, and are validators chosen randomly for each slot.
# - We also haven't given our validators the power to slash malicious validators. With a few tweaks here and there we can do that simply enough.
# + active=""
# Beacon Runner 2050: An agent-based model of eth2
# + active=""
#
# + active=""
# // References + footnotes
#
# // Authors
# let authorData = ["barnabe"];
# + active=""
# Many thanks once more to <NAME> for his edits and suggestions; <NAME> and Protolambda for their help and inputs.
|
notebooks/beaconrunner2050/br2050.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Normalization Quiz
# Use what you've learned in the last video to normalize case in the following text and remove punctuation!
#
# **Note: All solution notebooks can be found by clicking on the Jupyter icon on the top left of this workspace.**
text = "The first time you see The Second Renaissance it may look boring. Look at it at least twice and definitely watch part 2. It will change your view of the matrix. Are the human people the ones who started the war ? Is AI a bad thing ?"
print(text)
# ### Case Normalization
# Convert to lowercase
text = text.lower()
print(text)
# ### Punctuation Removal
# Use the `re` library to remove punctuation with a regular expression (regex). Feel free to refer back to the video or Google to get your regular expression. You can learn more about regex [here](https://docs.python.org/3/howto/regex.html).
# Remove punctuation characters
import re
text = re.sub(r"[^a-zA-Z0-9]"," ",text)
print(text)
|
Data Engineering/NLP Pipleines/normalization_practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pyemma.msm as msm
import msmtools.generation as msmgen
import msmtools.analysis as msmana
import pyemma.coordinates as coor
import matplotlib.pylab as plt
#import anca
# %pylab inline
plt.style.use('ggplot')
def assign(X, cc):
T = X.shape[0]
I = np.zeros((T),dtype=int)
for t in range(T):
dists = X[t] - cc
dists = dists ** 2
I[t] = np.argmin(dists)
return I
P = np.array([[0.99, 0.01],
[0.01, 0.99]]);
T = 50000
means = [np.array([-1,1]), np.array([1,-1])];
widths = [np.array([0.3,2]),np.array([0.3,2])];
# continuous trajectory
X = np.zeros((T, 2))
# hidden trajectory
dtraj = msmgen.generate_traj(P, T)
for t in range(T):
s = dtraj[t]
X[t,0] = widths[s][0] * numpy.random.randn() + means[s][0]
X[t,1] = widths[s][1] * numpy.random.randn() + means[s][1]
dtraj.shape
plt.plot(dtraj[0:500]);
plt.figure(figsize=(4,7))
plt.scatter(X[:,0], X[:,1], marker = 'o', color=[0.6,0.6,0.6])
# ## Spatial Decorrelation of Order 2 (SD2)
#
# Parameters:
#
# data – a 3n x T data matrix (number 3 is due to the x,y,z coordinates for each atom). Maybe a numpy
# array or a matrix where,
#
# n: size of the protein
#
# T: number of snapshots of MD trajectory
#
# m – dimensionality of the subspace we are interested in; Default value is None, in which case m = n
# verbose – print information on progress. Default is true.
#
# Returns:
#
# A 3n x m matrix U (NumPy matrix type), such that Y = U * data is a 2nd order spatially whitened coordinates extracted from the 3n x T data matrix. If m is omitted, U is a square 3n x 3n matrix.
#from anca.decorrelation import SD2
import SD2
(Y, S, B, U) = SD2.SD2(X, m=2);
# ## Spatial Decorrelation Module of Order 4 (SD4)
#
# Parameters:
#
# Y -- an mxT spatially whitened matrix (m dimensionality of subspace, T snapshots). May be a numpy
# array or a matrix where
# m -- dimensionality of the subspace we are interested in. Defaults to None, in
# which case m=n.
# T -- number of snapshots of MD trajectory
#
# U -- whitening matrix obtained after doing the PCA analysis on m components
# of real data
#
# verbose -- print info on progress. Default is True.
#
# Returns:
#
# W -- a separating matrix for spatial decorrelation of order 4
#from anca.decorrelation import SD4
import SD4
W = SD4.SD4(Y, m=2, U=U)
def draw_arrow(a, v, color):
plt.arrow(0, 0, a*v[0], a*v[1], color=color, width=0.02, linewidth=3)
# +
plt.figure(figsize=(4,7))
scatter(X[:,0], X[:,1], marker = 'o', color=[0.6,0.6,0.6])
plt.arrow(0, 0, 7*U[0,0], 12*U[0,1], color='red', width=0.02, linewidth=3);
plt.text(-0.0, 6.5, 'SD2', color='red', fontsize=20, fontweight='bold', rotation='horizontal')
plt.arrow(0, 0, 3*W[0,0], 9*W[0,1], color='orange', width=0.02, linewidth=3);
plt.text(1.5, 3.5, 'SD4', color='orange', fontsize=20, fontweight='bold', rotation='horizontal')
# -
YSD4 = W.dot(Y);
# +
hist(3*Y[0,:].T, bins=50, histtype='step', linewidth=3, label='SD2', color='blue')
hist(4*YSD4[0,:].T, bins=50, histtype='step', linewidth=3, label='SD4', color='red')
xlabel('essential coordinate (1st principal or independent component)')
ylabel('projected histogram')
legend()
|
Examples/Double Well Spatial Decorrelation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TensorFlow: Static Graphs
#
# PyTorch autograd looks a lot like TensorFlow: in both frameworks we define a computational graph, and use automatic differentiation to compute gradients. The biggest difference between the two is that TensorFlow's computational graphs are *static* and PyTorch uses *dynamic* computational graphs.
#
# In TensorFlow, we define the computational graph once and then execute the same graph over and over again, possibly feeding different input data to the graph. In PyTorch, each forward pass defines a new computational graph.
#
# Static graphs are nice because you can optimize the graph up front; for example a framework might decide to fuse some graph operations for efficiency, or to come up with a strategy for distributing the graph across many GPUs or many machines. If you are reusing the same graph over and over, then this potentially costly up-front optimization can be amortized as the same graph is rerun over and over.
#
# One aspect where static and dynamic graphs differ is control flow. For some models we may wish to perform different computation for each data point; for example a recurrent network might be unrolled for different numbers of time steps for each data point; this unrolling can be implemented as a loop. With a static graph the loop construct needs to be a part of the graph; for this reason TensorFlow provides operators such as *tf.scan* for embedding loops into the graph. With dynamic graphs the situation is simpler: since we build graphs on-the-fly for each example, we can use normal imperative flow control to perform computation that differs for each input.
#
# To contrast with the PyTorch autograd example above, here we use TensorFlow to fit a simple two-layer net.
# +
import tensorflow as tf
import numpy as np
# First we set up the computational graph:
# N is batch size; D_in is input dimension;
# H is hidden dimension; D_out is output dimension.
N, D_in, H, D_out = 64, 1000, 100, 10
# Create placeholders for the input and target data; these will be filled
# with real data when we execute the graph.
x = tf.placeholder(tf.float32, shape=(None, D_in))
y = tf.placeholder(tf.float32, shape=(None, D_out))
# Create Variables for the weights and initialize them with random data.
# A TensorFlow Variable persists its value across executions of the graph.
w1 = tf.Variable(tf.random_normal((D_in, H)))
w2 = tf.Variable(tf.random_normal((H, D_out)))
# Forward pass: Compute the predicted y using operations on TensorFlow Tensors.
# Note that this code does not actually perform any numeric operations; it
# merely sets up the computational graph that we will later execute.
h = tf.matmul(x, w1)
h_relu = tf.maximum(h, tf.zeros(1))
y_pred = tf.matmul(h_relu, w2)
# Compute loss using operations on TensorFlow Tensors
loss = tf.reduce_sum((y - y_pred) ** 2.0)
# Compute gradient of the loss with respect to w1 and w2.
grad_w1, grad_w2 = tf.gradients(loss, [w1, w2])
# Update the weights using gradient descent. To actually update the weights
# we need to evaluate new_w1 and new_w2 when executing the graph. Note that
# in TensorFlow the the act of updating the value of the weights is part of
# the computational graph; in PyTorch this happens outside the computational
# graph.
learning_rate = 1e-6
new_w1 = w1.assign(w1 - learning_rate * grad_w1)
new_w2 = w2.assign(w2 - learning_rate * grad_w2)
# Now we have built our computational graph, so we enter a TensorFlow session to
# actually execute the graph.
with tf.Session() as sess:
# Run the graph once to initialize the Variables w1 and w2.
sess.run(tf.global_variables_initializer())
# Create numpy arrays holding the actual data for the inputs x and targets
# y
x_value = np.random.randn(N, D_in)
y_value = np.random.randn(N, D_out)
for _ in range(500):
# Execute the graph many times. Each time it executes we want to bind
# x_value to x and y_value to y, specified with the feed_dict argument.
# Each time we execute the graph we want to compute the values for loss,
# new_w1, and new_w2; the values of these Tensors are returned as numpy
# arrays.
loss_value, _, _ = sess.run([loss, new_w1, new_w2],
feed_dict={x: x_value, y: y_value})
print(loss_value)
|
pytorch/05-net-tensorflow.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Regression Challenge
#
# Predicting the selling price of a residential property depends on a number of factors, including the property age, availability of local amenities, and location.
#
# In this challenge, you will use a dataset of real estate sales transactions to predict the price-per-unit of a property based on its features. The price-per-unit in this data is based on a unit measurement of 3.3 square meters.
#
# > **Citation**: The data used in this exercise originates from the following study:
# >
# > *<NAME>., & <NAME>. (2018). Building real estate valuation models with comparative approach through case-based reasoning. Applied Soft Computing, 65, 260-271.*
# >
# > It was obtained from the UCI dataset repository (<NAME>. and <NAME>. (2019). [UCI Machine Learning Repository](http://archive.ics.uci.edu/ml). Irvine, CA: University of California, School of Information and Computer Science).
#
# ## Review the data
#
# Let's hit the ground running by importing the data and viewing the first few rows.
#
# +
# Load the core tidyverse and tidymodels in your current R session
suppressPackageStartupMessages({
library(tidyverse)
library(tidymodels)
})
# Read the csv file into a tibble
estate_data <- read_csv(file = "https://raw.githubusercontent.com/MicrosoftDocs/ml-basics/master/challenges/data/real_estate.csv",
show_col_types = FALSE)
# Print the first 10 rows of the data
estate_data %>%
slice_head(n = 10)
# -
# The data consists of the following variables:
#
# - **transaction_date** - the transaction date (for example, 2013.250=2013 March, 2013.500=2013 June, etc.)
#
# - **house_age** - the house age (in years)
#
# - **transit_distance** - the distance to the nearest light rail station (in meters)
#
# - **local_convenience_stores** - the number of convenience stores within walking distance
#
# - **latitude** - the geographic coordinate, latitude
#
# - **longitude** - the geographic coordinate, longitude
#
# - **price_per_unit** house price of unit area (3.3 square meters)
#
# Your challenge is to explore and prepare the data, identify predictive features that will help predict the `price_per_unit` label, and train a regression model that achieves the lowest *Root Mean Square Error* (RMSE) you can achieve (which must be less than *7*) when evaluated against a test subset of data.
#
# ### View the label distribution
#
# Let's start our analysis of the data by examining a few key descriptive statistics. We can use the `summarytools::descr()` function to neatly and quickly summarize the numeric features as well as the *rentals* label column.
#
# +
# Load summary tools library
library(summarytools)
# Obtain summary stats for feature and label columns
estate_data %>%
# Summary stats
descr(order = "preserve",
stats = c("mean", "sd", "min", "q1", "med", "q3", "max"),
round.digits = 6)
# -
# The statistics reveal some information about the distribution of the data in each of the numeric fields, including the number of observations (there are 414 records), the mean, standard deviation, minimum and maximum values, and the quantile values (the threshold values for 25%, 50% - which is also the median, and 75% of the data).
#
# From this, we can see that the mean number of price per unit is around 38. There's a comparatively *small standard deviation*, indicating *not much variance* in the prices per unit.
#
# We might get a clearer idea of the distribution of price values by visualizing the data.
#
# +
library(patchwork)
# Plot a histogram
theme_set(theme_light())
hist_plt <- estate_data %>%
ggplot(mapping = aes(x = price_per_unit)) +
geom_histogram(bins = 100, fill = "midnightblue", alpha = 0.7) +
# Add lines for mean and median
geom_vline(aes(xintercept = mean(price_per_unit), color = "Mean"),
linetype = "dashed", size = 1.3) +
geom_vline(aes(xintercept = median(price_per_unit), color = "Median"),
linetype = "dashed", size = 1.3) +
xlab("") +
ylab("Frequency") +
scale_color_manual(name = "", values = c(Mean = "red", Median = "yellow")) +
theme(legend.position = c(0.9, 0.9), legend.background = element_blank())
# Plot a box plot
box_plt <- estate_data %>%
ggplot(aes(x = price_per_unit, y = 1)) +
geom_boxplot(fill = "#E69F00", color = "gray23", alpha = 0.7) +
# Add titles and labels
xlab("Price_per_unit") +
ylab("")
# Combine plots using patchwork syntax
(hist_plt / box_plt) +
plot_annotation(title = "Price Distribution",
theme = theme(
plot.title = element_text(hjust = 0.5)))
# -
# What can we observe from the boxplot? Yes, outliers.
#
# ### Remove outliers
#
# We are now set to begin writing some code ourselves 🙂. Let's begin by dealing with outliers. An outlier is a data point that differs significantly from other observations.
#
# **Question 1.**
#
# Starting with the `estate_data` dataset, `filter` to create a subset that contains observations where `price_per_unit` is less than *70*.
#
# Fill in the placeholder `....` with the right code.
# Narrow down to observations whose price_per_unit is less than 70
estate_data <- estate_data %>%
....
# Test your answer:
#
. <- ottr::check("tests/Question 1.R")
# Now let's take a look at the distribution without the outliers.
#
#
# +
# Plot a histogram
theme_set(theme_light())
hist_plt <- estate_data %>%
ggplot(mapping = aes(x = price_per_unit)) +
geom_histogram(bins = 100, fill = "midnightblue", alpha = 0.7) +
# Add lines for mean and median
geom_vline(aes(xintercept = mean(price_per_unit), color = "Mean"),
linetype = "dashed", size = 1.3) +
geom_vline(aes(xintercept = median(price_per_unit), color = "Median"),
linetype = "dashed", size = 1.3) +
xlab("") +
ylab("Frequency") +
scale_color_manual(name = "", values = c(Mean = "red", Median = "yellow")) +
theme(legend.position = c(0.9, 0.9), legend.background = element_blank())
# Plot a box plot
box_plt <- estate_data %>%
ggplot(aes(x = price_per_unit, y = 1)) +
geom_boxplot(fill = "#E69F00", color = "gray23", alpha = 0.7) +
# Add titles and labels
xlab("Price_per_unit") +
ylab("")
# Combine plots using patchwork syntax
(hist_plt / box_plt) +
plot_annotation(title = "Price Distribution",
theme = theme(
plot.title = element_text(hjust = 0.5)))
# -
# Much better 🤩! What can we say about the distribution of the price?
#
# ### View numeric correlations
#
# We can now start to look for relationships between the *features* and the *label* we want to be able to predict.
#
# The *correlation* statistic, *r*, is a value between -1 and 1 that indicates the strength of a linear relationship.
#
# For numeric feature and label columns, we can create scatter plots that show the intersection of the feature and label values.
#
# **Question 2.**
#
# Starting with the `estate_data` dataset, in a piped sequence:
#
# - `pivot_longer` the data (increase the number of rows and decrease the number of columns) such that all the existing column names except price_per_unit, now fall under a new column name called `features` and their corresponding values under a new column name `values`
#
# - group the data by `features`
#
# - add a new column `corr_coef` which calculates the correlation between `values` and `price_per_unit` (hint: the function used for calculating correlation in R is `cor()`)
#
# Fill in the placeholder `....` with the right code.
# +
# Pivot numeric features to a long format
numeric_features_long <- estate_data %>%
pivot_....(!price_per_unit, names_to = "....", values_to = "....") %>%
# Group by features
....(features) %>%
# Calculate correlation coefficient between values and price_per_unit
mutate(corr_coef = ....) %>%
# Modifies the feature column to also include corr_coef
mutate(features = paste(features, "vs price, r = ",
round(corr_coef, 2), sep = "")) %>%
ungroup()
# Print the first few rows of the data
numeric_features_long %>%
slice_head(n = 10)
# -
# Test your answer:
#
. <- ottr::check("tests/Question 2.R")
# Fantastic! Now let's use a scatter plot to investigate whether there is any linear relationship between our predictors and outcome variables.
#
#
# Plot a scatter plot for each feature
numeric_features_long %>%
ggplot(aes(x = values, y = price_per_unit, color = features)) +
geom_point(alpha = 0.7, show.legend = F) +
facet_wrap(~ features, scales = "free") +
paletteer::scale_color_paletteer_d("ggthemes::excel_Parallax")
# Take a moment and go through the scatter plot. How does the correlation between these features and the price vary?
#
# ### View categorical features
#
# Now let's compare the categorical features to the label. We'll do this by creating box plots that show the distribution of rental counts for each category.
#
# `Transaction_date` and `local_convenience_stores` seem to be discrete values - so might work better if treated as categorical features. Let' get right into it.
#
# **Question 3.**
#
# Starting with the `estate_data` dataset, in a piped sequence:
#
# - only keep columns `transaction_date`, `local_convenience_stores` and `price_per_unit`
#
# - encode columns `transaction_date` and `local_convenience_stores` as categorical (factor)
#
# - `pivot_longer` the data (increase the number of rows and decrease the number of columns) such that all the existing column names except price_per_unit now fall under a new column name called `features` and their corresponding values under a new column name `values`
#
# Fill in the placeholder `....` with the right code.
# +
# Pivot categorical features to a long format
cat_features_long <- estate_data %>%
....(transaction_date, ...., ....) %>%
# Encode transaction_date & local_convenience_stores features
# from numeric to categorical
mutate(....) %>%
pivot_longer(....)
# Print some observations
cat_features_long %>%
slice_head(n = 10)
# -
# Test your answer:
#
. <- ottr::check("tests/Question 3.R")
# Perfect! Now, for our categorical features, boxplots can be a great way of visualising how the price per unit varies within the levels of the categorical feature.
#
#
# Plot a box plot for each feature
cat_features_long %>%
ggplot() +
geom_boxplot(aes(x = values, y = price_per_unit, fill = features),
alpha = 0.7, show.legend = F) +
facet_wrap(~ features, scales = "free") +
scale_fill_viridis_d() +
theme(panel.grid = element_blank(),
axis.text.x = element_text(angle = 90))
# Take a moment and interpret the graphics. How does the price vary with these features?
#
# ## Split the data into training and test sets.
#
# Now that we've explored the data, it's time to use it to train a regression model that uses the features we've identified as *potentially predictive* to predict the `price_per_unit` label.
#
# `Transaction_date` doesn't seem to be very predictive, so we'll omit it.
#
# Let's begin by splitting the data set such that some goes to training and some goes for validation. This enables us to evaluate how well the model performs in order to get a better estimate of how your models will perform on new data.
#
# **Question 4.**
#
# In this section:
#
# - Make a split specification of `estate_data` such that *70%* goes to training and the rest goes to testing. Save this to a variable name `estate_split`
#
# - Extract the training and testing sets from `estate_split` and save them in `estate_train` and `estate_test` variable names respectively.
#
# Fill in the placeholder `....` with the right code.
# +
# Set seed to ensure reproducibility and consistency of outputs
set.seed(2056)
# Load the tidymodels package
library(tidymodels)
# Split 70% of the data for training and the rest for tesing
estate_split <- estate_data %>%
initial_split(....)
# Extract the train and test data in each split
estate_train <- ....(estate_split)
estate_test <- ....(estate_split)
# Print the number of observations in each split
cat("Training Set", nrow(estate_train), "rows",
"\nTest Set", nrow(estate_test), "rows")
# -
# Test your answer:
#
. <- ottr::check("tests/Question 4.R")
# Great progress 💪! Now let's train some models.
#
# ## Train a regression model
#
# ### Preprocess data using recipes
#
# Often before fitting a model, we may want to reformat the predictor values to make them easier for a model to use effectively. This includes transformations and encodings of the data to best represent their important characteristics. In R,this is done using a `recipe`.
#
# A recipe is an object that defines a series of steps for data processing.
#
# **Question 5.**
#
# In this section, specify a recipe, `estate_recipe`, that will:
#
# - Remove the `transaction_date` feature
#
# - Transform `local_convenience_stores` feature into categorical (factor)
#
# - Center and scale all numeric predictors
#
# Fill in the placeholder `....` with the right code.
# +
# Create a preprocessing recipe
estate_recipe <- ....(price_per_unit ~ ., data = estate_train) %>%
# Specify the removal of transaction_date
step_rm(....) %>%
# Specify the encoding of local_convenience_stores as categorical
step_mutate(
local_convenience_stores = ....) %>%
# Specify the normalization of numeric features
....(all_numeric_predictors())
# Print recipe
estate_recipe
# -
# Test your answer:
#
. <- ottr::check("tests/Question 5.R")
# Fantastic! We have the data processing in order. Now, let's make a model specification. In this solution, we'll try out a random forest model which applies an averaging function to multiple decision tree models for a better overall model.
#
# **Question 6.**
#
# Create a random forest model specification, `rf_spec`, which uses the `randomForest` package as its engine and then set the mode to `regression`.
#
# Fill in the placeholder `....` with the right code.
# Build a random forest model specification
rf_spec <- rand_forest() %>%
# Specify engine
.... %>%
# Specify mode
set_mode("....")
# Test your answer:
#
. <- ottr::check("tests/Question 6.R")
# ### Create a modeling workflow
#
# The *workflows* package allows the user to bind modeling and preprocessing objects together. You can then fit the entire workflow to the data, so that the model encapsulates all of the preprocessing steps as well as the algorithm.
#
# **Question 7.**
#
# Components of a `workflow()` go together like LEGO blocks. In this section, create a workflow container and then add the preprocessing information from our recipe and then add the model specification to be trained.
#
# Fill in the placeholder `....` with the right code.
# Create a workflow that bundles a recipe and model specification
rf_workflow <- workflow() %>%
# Add a recipe
add_recipe(....) %>%
# Add a model specification
....
# Print workflow
rf_workflow
# Test your answer:
#
. <- ottr::check("tests/Question 7.R")
# Now that we have everything (recipe + model specification) wrapped together nicely in a workflow, we are ready to train a model. Workflows have a `fit()` method that can be used to train a model.
#
#
# +
# For reproducibility
set.seed(2056)
# Train a random forest model
rf_workflow_fit <- rf_workflow %>%
fit(data = estate_train)
# Print out the fitted workflow
rf_workflow_fit
# -
# Excellent! So we now have a trained random forest model; but is it any good? Let's evaluate its performance! We'll do this by making predictions on the `test data` and then evaluate some performance metrics based on the actual outcomes.
#
# **Question 8.**
#
# - We'll evaluate the model performance based on the *rmse* and *rsq* metrics. Use the `metric_set()` function to combine these metric functions together into a new function, `eval_metrics`, that calculates all of them at once.
#
# - Generate predictions for the test data and then bind them to the test set. Rename the column containing predictions from `.pred` to `predictions`.
#
# Fill in the placeholder `....` with the right code.
# +
# Create a metric set
eval_metrics <- ....(rmse, ....)
# Make and bind predictions to test data
results <- rf_workflow_fit %>%
....
# -
# Test your answer:
#
. <- ottr::check("tests/Question 8.R")
# Awesome work! You have just used your trained model to make predictions on the test set.
#
# How well did the model predict the prices per unit? Let's find out by looking at the metrics.
#
# +
# Evaluate the model
rf_metrics <- eval_metrics(data = results,
truth = price_per_unit,
estimate = predictions)
# Plot predicted vs actual
rf_plt <- results %>%
ggplot(mapping = aes(x = price_per_unit, y = predictions)) +
geom_point(color = "darkorchid", size = 1.6) +
# overlay regression line
geom_smooth(method = "lm", color = "black", se = F) +
ggtitle("Price per unit predictions") +
xlab("Actual Labels") +
ylab("Predicted Labels") +
theme(plot.title = element_text(hjust = 0.5))
# Return evaluations
list(metrics = rf_metrics, evaluation_plot = rf_plt)
# -
# How do you think the model performed? What do the values for `rsq` and `rmse` tell you? Please refer to the corresponding module for this notebook if you may need help answering these questions.
#
# ## Use the Trained Model
#
# Save your trained model, and then use it to predict the price-per-unit for the following real estate transactions:
#
# | **transaction_date** | **house_age** | **transit_distance** | **local_convenience_stores** | **latitude** | **longitude** |
# |----------------------|---------------|----------------------|------------------------------|--------------|---------------|
# | 2013.167 | 16.2 | 289.3248 | 5 | 24.98203 | 121.54348 |
# | 2013.000 | 13.6 | 4082.015 | 0 | 24.94155 | 121.50381 |
#
#
library(here)
# Save trained workflow
saveRDS(rf_workflow_fit, "rf_price_model.rds")
# In this way, we can load it whenever we need it, and use it to predict labels for new data. This is often called *scoring* or *inferencing*.
#
#
# +
# Create a tibble for the new real estate samples
new_data <- tibble(
transaction_date = c(2013.167, 2013.000),
house_age = c(16.2, 13.6),
transit_distance = c(289.3248, 4082.015),
local_convenience_stores = c(5, 0),
latitude = c(24.98203, 24.94155),
longitude = c(121.54348, 121.50381))
# Print out new data
new_data
# -
# Now that we have our data, let's load the saved model and make predictions.
#
#
# +
# Load the model into the current R session
loaded_model <- readRDS("rf_price_model.rds")
# Make predictions
predictions <- loaded_model %>%
augment(new_data = new_data)
predictions
# -
# Congratulations for completing this challenge! In this notebook, you:
#
# - Explored the data set to understand the relationships between the predictors and outcomes
# - Preprocessed the data using recipes to make them easier for a model to use effectively.
# - Made a random forest model specification.
# - Bundles a recipe and model specification into a workflow.
# - Trained a model.
# - Made predictions on test set and evaluated the model performance.
# - Saved the model, loaded it and then used it to predict labels for new data.
#
# Fantastic job for coming this far 👏! Feeling adventurous? Then, be sure to try out other regression models and tune some hyperparameters while at it.
#
# See you in our next module as we explore the realm of *classification* models!
#
# Happy Learning,
#
# [Eric](https://twitter.com/ericntay), Gold Microsoft Learn Student Ambassador.
#
|
full/intro-regression-R-tidymodels/solution/Challenge-regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
from torch import nn
import matplotlib.pyplot as plt
import context
from ccn import Constraint, ConstraintsGroup
from shapes import HalfPlane
from experiment import Experiment
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print(device)
# -
# ### Define the shapes
# +
A = HalfPlane(1, -1, 0)
B = HalfPlane(+1, +1, -1)
C = B | -A
shapes = [A, B, C]
fig, ax = plt.subplots(1, len(shapes))
for i, shape in enumerate(shapes):
shape.plot(ax[i], full=True)
plt.show()
constraints = [ConstraintsGroup([Constraint('n2 :- 0 n1')])]
# -
# ### Define the neural network
# +
class NeuralNetwork(nn.Module):
def __init__(self):
super(NeuralNetwork, self).__init__()
self.linear = nn.Sequential(
nn.Linear(2, 4),
nn.Tanh(),
nn.Linear(4, 3),
nn.Sigmoid()
)
def forward(self, x):
x = self.linear(x)
return x
model = NeuralNetwork()
print(model)
# -
# ### Train and test the model
experiment = Experiment('halfplanes', model, shapes, constraints)
experiment.run(500)
experiment.save(dir='./models/')
|
experiments/HalfPlanes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bert_serving.client import BertClient
bc = BertClient()
import torch
bc.encode(['Jagadish is a student of IITKgp'])
import os
os.environ['KMP_DUPLICATE_LIB_OK']='True'
import pandas as po
# !ls
# ## Best Score
# Pearson = 0.8621
# mem_dim = 100
# hidden_dim = 200
# lr = 0.03
# wd = 0.001
# epochs = 10
# seed = 1947
import numpy as np
a = np.ndarray([3,5])
torch.from_numpy(a)
torch.tensor(1).float()
# !python treelstm/main.py --lr 0.03 --wd 0.001 --optim adagrad --batchsize 25 --seed 1947 --epochs 1 --hidden_dim 200 --mem_dim 100 --model alldep
(torch.rand([768]).view(1,768)).shape
results = po.DataFrame(columns = ['hidden_dim', 'mem_dim', 'lr', 'pearson', 'spearman', 'mse'])
results.to_csv('checkpoints/mdep/Results.csv', index = False)
# !python treelstm/main.py --lr 0.05 --wd 0.0001 --opti m adagrad --batchsize 25 --freeze_embed --epochs 1 --hidden_dim 60 --mem_dim 100
# +
dict = {'hidden_dim': 50,
'lr': 0.05,
'pearson': 0.5673,
'spearman': 0.5711,
'mse': 0.6944,
'mem_dim': 150
}
results = results.append(dict, ignore_index=True)
results.to_csv('checkpoints/mdep/Results.csv', index = False)
# -
results=po.read_csv('checkpoints/mdep/Results.csv')
results
mse = results.iloc[-1]['mse']
mse
# +
from hyperopt import fmin,tpe,hp, STATUS_OK
best_mse = 1000
def f(space):
global best_h_d, best_m_d, best_lr, best_mse
hidden_dim = space['hidden_dim']
lr = space['lr']
mem_dim = space['mem_dim']
# !python treelstm/main.py --wd 0.0001 --optim adagrad --batchsize 25 --freeze_embed --epochs 1 --hidden_dim hidden_dim --mem_dim mem_dim
results=po.read_csv('checkpoints/mdep/Results.csv')
mse = results.iloc[-1]['mse']
if (mse < best_mse):
best_m_d = m_d
best_h_d = h_d
best_lr = lr
best_mse = mse
print('best_mse = {}'.format(best_mse))
print('best_h_d = {}'.format(h_d))
print('best_lr = {}'.format(lr))
print('best_m_d = {}'.format(m_d))
return {'loss': mse, 'status': STATUS_OK }
space = {
'hidden_dim': hp.quniform('hidden_dim', 10, 200,10),
'mem_dim': hp.quniform('mem_dim',10,200,10),
'lr': hp.quniform('lr',0.01,1,0.01),
}
best = fmin(fn=f,space=space,algo=tpe.suggest,max_evals=150)
# -
space['hidden_dim']
|
Tuning_BERT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Asynchronous Programming with OpenACC
# This lab is intended for Fortran programmers. If you prefer to use C/C++, click [this link.](../C/Lab7_C.ipynb)
#
# Don't forget to check out additional [OpenACC resources](https://www.openacc.org/resources) and join our [OpenACC Slack Channel](https://www.openacc.org/community#slack) to share your experience and get more help from the community.
# ---
# ## Introduction
#
# Asynchronous programming is a programming technique such that two or more unrelated operations can occur independently or even at the same time without immediate synchronization. In OpenACC, when we refer to asynchronous programming we are generally referring to performing accelerated computation while simultaneously performing data transfers between host and device, enqueuing additional work to keep the device busy, performing unrelated work on the host CPU, or even sending work to two different devices simultaneously (more on this in a future lab!). The goal for this lab is to use the OpenACC **async** clause to speedup a code by *overlapping* our compute with our data movement.
# ---
#
# ## Analyze the Code
#
# The code we will be using for this lab is an image filtering code. Click the following links to view the code before we begin analyzing it.
#
# [main.cpp](/edit/Fortran/main.cpp)
# [filter.F90](/edit/Fortran/NoBlocking/filter.F90)
#
# All of our changes will be done in *filter.F90*. We will start off with a very simple parallelization of the *blur5* function, and by the end of the lab have a more optimized version using OpenACC **async**.
# ---
# ## OpenACC Async Clause
#
# We use the **async** clause to specify code that should be run *asynchronously*. The async clause can be added to our *parallel* or *kernels* regions, and to our *enter data*, *exit data*, and *update* directives. The easiest way to explain how the async clause works is to first explain what happens when we don't use it.
#
# When no async clause is used, the host thread will pause at the end of the OpenACC region (which could be any of the directives we mentioned above). The host thread will stay paused until that region is finished. For example, if we have a loop inside of a parallel region, then that loop will be executed on the device. The host thread will wait until the kernels region is done, which means that it will wait until that loop is done running on the device. Here is a simple example:
#
# ```fortran
# sum = 0
# # !$acc parallel loop reduction(+:sum)
# do i = 1, 100
# sum = sum + i
# end do
#
# WRITE(*,*) 'Summation determined as', sum
# ```
#
# We are expecting the host thread to wait for the parallel loop to finish before it tries to print the value of *sum*. Next, consider if we add the async clause to our parallel loop:
#
# ```fortran
# sum = 0
# # !$acc parallel loop reduction(+:sum) async
# do i = 1, 100
# sum = sum + i
# end do
#
# WRITE(*,*) 'Summation determined as', sum
# ```
#
# The host thread will **not wait** for the parallel loop to finish. This means that instead of printing the correct sum value, it will most likely print 0. This example is to show that the async clause should not always be used, as it is often important that we wait for our parallel loops, and for our data movement to finish before continuing host code. However, in some codes (like the one we will work on shortly) we can exploit the OpenACC asynchronous behavior to improve performance.
#
# ### Benefits of Using Async
#
# Here are the primary benefits we can expect when using async in our code:
#
# * We can execute host and device code simultaneously. We can launch our device code with *async*, and while that executes we can go back to the host to continue unrelated (non-device dependent) code.
# * We can *queue up* multiple device kernel launches so that they execute back-to-back, which in some cases can reduce overhead associated with launching device kernels.
# * **We can perform device computation at the same time as data movement between host and device.** This is the optimization we will be applying to our code in this lab, and is the most general use case of async
#
# ---
# ## Async Example
#
# Let's look at the following code example:
#
# ```fortran
# real, dimension(1:N) :: A\n",
# real, dimension(1:M) :: B\n",
# # !$acc enter data create(A(:), B(:))\n",
#
# # ! Compute Loop 1
# # !$acc parallel loop present(A)
# do i = 1, N
# A(i) = ...
# end do
#
# # ! Update 1
# # !$acc update self(A(:))
#
# # ! Compute Loop 2
# # !$acc parallel loop present(B)
# do i = 1, M
# B(i) = ...
# end do
#
# # ! Update 2
# # !$acc update self(B(:))
# ```
#
# This code has two main restrictions: *Compute Loop 1* must finish before *Update 1* starts, and *Compute Loop 2* must finish before *Update 2* starts. This means that there is no reason to wait between *Update 1* and *Compute Loop 2*. We can use the **async clause** on *Update 1*. This will allow the code to continue onto *Compute Loop 2* without needing to wait for *Update 1* to finish.
#
# ```fortran
# # ! Update 1
# # !$acc update self(A(:)) async
# ```
#
# Another way we can accomplish this is by placing *Compute Loop 1*/*Update 1* and *Compute Loop 2*/*Update 2* into separate **queues**.
# ---
# ## OpenACC Queues
#
# Under-the-hood whenever we use the async clause, we are adding some *work* to a **queue**. Work that is in different queues can execute *asynchronously*, and work that is in the same queue will execute *sequentially* (one after the other). When we use async, we are able to specify a queue number. If no queue number is specified, then a default will automatically be used. Additionally, if there is no async clause present, the work will still be placed in a (different) default queue, known as the synchronous queue. There is an overhead associated with launching new queues, so the goal is to use the minimum number that you need, and reuse queues when possible. Let's first look at an example without using async.
#
# 
#
# Next, let's add some async and see how the work will be distributed into queues.
#
# 
#
# Since Loop1/Update1 and Loop2/Update2 are in separate queues, they will execute independently of each other. The queues are still limited by the capabilities of the device, but generally this means that if one queue is doing computation, and the other is doing data movement they can occur at the same time.
#
# We have one more problem that we have to address before working on the code.
# ---
# ## OpenACC Wait
#
# Let's consider the following code:
#
# ```fortran
# # !$acc parallel loop async(1)
# do i = 1, 100
# A(i) = 1.0
# end do
#
# # !$acc update self(A(1:100)) async(1)
#
# # !$acc parallel loop async(2)
# do i = 1, 100
# B(i) = 2.0
# end do
#
# # !$acc update self(B(0:100)) async(2)
#
# # ! Back to Host Code
# call print_arrays(A,B)
# ```
#
# We want to perform the loops and data movement asynchronously for the performance benefit, but we need to make sure we pause and wait for everything to finish up before executing that printing loop on the host. We can accomplish this by using the **OpenACC wait directive**. The syntax is as follows:
#
# **\!$acc wait(*queue*)**
#
# This will pause the host until the specified queue is finished. If no queue is specified, then it will wait for all queues to finish. Adding it to the above code would look like this:
#
# ```fortran
# # !$acc parallel loop async(1)
# do i = 1, 100
# A(i) = 1.0
# end do
#
# # !$acc update self(A(1:100)) async(1)
#
# # !$acc parallel loop async(2)
# do i = 1, 100
# B(i) = 2.0
# end do
#
# # !$acc update self(B(0:100)) async(2)
#
# # !$acc wait
#
# # ! Back to Host Code
# call print_arrays(A,B)
# }
# ```
#
# Let's look at our visual example from earlier with the added **wait** directive:
#
# 
# ---
# ## Optimizing Our Code
#
# In our code, we apply a simple filter to each pixel of an image. Each pixel of the image can be computed independently from each other. Our goal is to break up the computation of the image into *blocks*, and compute each block independently. Blocking the image in this manner allows us to alternate between computing a block and transferring the data of a block. Then, we can optimize it with async.
#
# If you followed the lecture slides associated with this lab, then you should be familiar with a code called *mandelbrot*. This code implements a similar concept that we are trying to achieve; here is a visualization:
#
# 
#
# Here we have an image that we are breaking up (along the Y direction) into several blocks. Then we want to use async to overlap the computation of the blocks, and the data transfer back to the host. Here is a diagram of what we hope to achieve.
#
# 
#
# ---
#
# Let's run the code and get the baseline performance. Also, if you would like to view the "before image", you may so so [here.](/view/C/costarica.jpg)
# !make clean && make
# ---
# ### Applying Blocking
#
# The first step that we will do is to apply **blocking** to the code. By blocking the code, we allow us to alternate between compute and data transfer, and will eventually let us apply the async clause.
#
# Edit the code file with [this link.](/edit/Fortran/filter.F90), and try to apply blocking to the **blur5** function. There is also blur5_serial and blur5_parallel which we will use to measure performance, so do not edit those.
#
# Blocking the code is not always easy. If you are feeling stuck, here are some hints:
#
# * First decide how many blocks you want to break it up into. You can choose a hard-coded number, or try to pick a number based off the size of the image.
# * Create a new outer block loop. This loop should go from 0 -> number_blocks
# * In the block loop, compute a lower and upper bound for your **y** loop. For example, if my image was height 100, and I have 4 blocks. For block0 I would have lower=1, upper=25. For block1 I would have lower=26, upper=50. For block2 I would have lower=51, upper=75. And lastly for block 3 I would have lower=76, upper=100.
# * Change the **y** loop to run from lower_bound -> upper_bound, instead of 0 -> image_height.
# * Move the update directives into the block loop, and change them to only update the data that you need for that block. Pay special attention to the imgData array and how it is used, you need to read offsetted values based on the size of the filter.
#
# Here is some pseudo-code to give you an idea of what to try:
#
# ```
# for block from 0 -> numBlocks {
#
# lower = block * rowsPerBlock + 1
# upper = MIN(lower + rowsPerBlock, h)
# lowerDataBound = MAX(lower-(filtersize/2), 1)
# upperDataBound = MIN(upper+(filtersize/2), h)
#
# # !$acc update device( inData(:,:,lowerDataBound:upperDataBound) )
#
# # !$acc parallel loop
# do y from lower -> upper
# do x from 0 -> WIDTH
# < Compute Pixel >
# enddo
# enddo
#
# # !$acc update self( outData(:,:,lower:upper) )
#
# }
# ```
#
# When you want to give it a try, run the following block:
# !make clean && make
# There isn't a single clear-cut solution on how to apply the blocking, but if you want to view our solution to compare, you can do so [here.](/edit/Fortran/Solution/filter_no_pipeline.F90) We have all of the steps that we took that lead for our final blocked solution. Also, if you would like to see the image that the code creates, you can view it [here.](view/C/out.jpg)
# ---
# ### Applying Async
# Now try adding async to the code and see if you can achieve a performance increase. Some hints to get you started:
#
# * You must first compute a block of the image before starting the data transfer. Thus, each block compute/transfer should share a queue, and different blocks should be in different queues. (you do use something like async(block%2) to achieve alternating blocks)
# * You need to ensure that you wait for all the blocks to finish before continuing onto the rest of the host code, so you will need the wait directive.
#
# Edit the code file with [this link.](/edit/Fortran/filter.F90)
# Re-run the code when you want to test your changes, and if you are stuck we will have the solution below! And after running, if you would like to see the output image again, use [this link.](/view/Fortran/out.jpg)
# !make clean && make
# ### Solutions
#
# If you would like to check the solution to this lab, click the [following link.](/edit/Fortran/Solution/filter.F90) This file contains the base version of blur5, alongside all of the steps taken to block the code, and the final async solution.
# ---
# ## Bonus Task
#
# We hinted earlier to another use of OpenACC async earlier - reducing the launch overhead of our device kernels. There is an OpenACC video tutorial done by <NAME> from PGI, and the 6th part of tutorial covers this other use of OpenACC async. This is a [link to that video.](https://youtu.be/voOcd1bNHIA)
# ---
#
# ## Post-Lab Summary
#
# If you would like to download this lab for later viewing, it is recommend you go to your browsers File menu (not the Jupyter notebook file menu) and save the complete web page. This will ensure the images are copied down as well.
#
# You can also execute the following cell block to create a zip-file of the files you've been working on, and download it with the link below.
# + language="bash"
# rm -f openacc_files.zip
# zip -r openacc_files.zip *
# -
# **After** executing the above zip command, you should be able to download the zip file [here](files/openacc_files.zip)
|
indevelopment/module7/English/Fortran/README.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Memanggil Library Pandas
import pandas as pd
import numpy as np
# # DataFrame & Series
# +
# Series
number_list = pd.Series([1,2,3,4,5,6])
print("Series:")
print(number_list)
# DataFrame
matrix = [[1,2,3],
['a','b','c'],
[3,4,5],
['d',4,6]]
matrix_list = pd.DataFrame(matrix)
print("DataFrame:")
print(matrix_list)
# -
# # Atribut DataFrame & Series
# +
# Series
number_list = pd.Series([1,2,3,4,5,6])
# DataFrame
matrix_list = pd.DataFrame([[1,2,3],
['a','b','c'],
[3,4,5],
['d',4,6]])
# [1] attribute .info()
print("[1] attribute .info()")
print(matrix_list.info())
# [2] attribute .shape
print("\n[2] attribute .shape")
print(" Shape dari number_list:", number_list.shape)
print(" Shape dari matrix_list:", matrix_list.shape)
# [3] attribute .dtypes
print("\n[3] attribute .dtypes")
print(" Tipe data number_list:", number_list.dtypes)
print(" Tipe data matrix_list:", matrix_list.dtypes)
# [4] attribute .astype()
print("\n[4] attribute .astype()")
print(" Konversi number_list ke str:", number_list.astype("str"))
print(" Konversi matrix_list ke str:", matrix_list.astype("str"))
# -
# [5] attribute .copy()
print("[5] attribute .copy()")
num_list = number_list.copy()
print(" Copy number_list ke num_list:", num_list)
mtr_list = matrix_list.copy()
print(" Copy matrix_list ke mtr_list:", mtr_list)
# [6] attribute .to_list()
print("[6] attribute .to_list()")
print(number_list.to_list())
# [7] attribute .unique()
print("[7] attribute .unique()")
print(number_list.unique())
# [8] attribute .index
print("[8] attribute .index")
print(" Index number_list:", number_list.index)
print(" Index matrix_list:", matrix_list.index)
# [9] attribute .columns
print("[9] attribute .columns")
print(" Column matrix_list:", matrix_list.columns)
# [10] attribute .loc
print("[10] attribute .loc")
print(" .loc[0:1] pada number_list:", number_list.loc[0:1])
print(" .loc[0:1] pada matrix_list:", matrix_list.loc[0:1])
# [11] attribute .iloc
print("[11] attribute .iloc")
print(" iloc[0:1] pada number_list:", number_list.iloc[0:1])
print(" iloc[0:1] pada matrix_list:", matrix_list.iloc[0:1])
# # Creating Series & Dataframe from List
# +
# Creating series from list
ex_list = ['a',1,3,5,'c','d']
ex_series = pd.Series(ex_list)
print(ex_series)
# Creating dataframe from list of list
ex_list_of_list = [[1, 'a', 'b', 'c'],
[2.5, 'd', 'e', 'f'],
[5, 'g', 'h', 'i'],
[7.5, 'j', 10.5, 'l']]
index = ['dq', 'lab', 'kar', 'lan']
cols = ['float', 'char', 'obj', 'char']
ex_df = pd.DataFrame(ex_list_of_list, index=index, columns=cols)
print(ex_df)
# -
# # Creating Series & Dataframe from Dictionary
# +
# Creating series from dictionary
dict_series = {'1' : 'a',
'2' : 'b',
'3' : 'c'}
ex_series = pd.Series(dict_series)
print(ex_series)
# Creating dataframe from dictionary
df_series = {'1': ['a','b','c'],
'2': ['b','c','d'],
'4': [2,3,'z']}
ex_df = pd.DataFrame(df_series)
print(ex_df)
# -
# # Creating Series & Dataframe from Numpy Array
# +
# Creating series from numpy array (1D)
arr_series = np.array([1,2,3,4,5,6,6,7])
ex_series = pd.Series(arr_series)
print(ex_series)
# Creating dataframe from numpy array (2D)
arr_df = np.array([[1,2,3,5],
[5,6,7,8],
['a','b','c',10]])
ex_df = pd.DataFrame(arr_df)
print(ex_df)
# -
# # Read Dataset - CSV dan TSV
# +
# File CSV
df_csv = pd.read_csv("https://storage.googleapis.com/dqlab-dataset/sample_csv.csv")
print(df_csv.head(3)) # Menampilkan 3 data teratas
# File TSV
df_tsv = pd.read_csv("https://storage.googleapis.com/dqlab-dataset/sample_tsv.tsv", sep='\t')
print(df_tsv.head(3)) # Menampilkan 3 data teratas
# -
# # Read Dataset - Excel
# File xlsx dengan data di sheet "test"
df_excel = pd.read_excel("https://storage.googleapis.com/dqlab-dataset/sample_excel.xlsx", sheet_name="test")
print(df_excel.head(4)) # Menampilkan 4 data teratas
# # Read Dataset - JSON
# File JSON
url = "https://storage.googleapis.com/dqlab-dataset/covid2019-api-herokuapp-v2.json"
df_json = pd.read_json(url)
print(df_json.head(10)) # Menampilkan 10 data teratas
# # Head & Tail
# +
# Baca file sample_csv.csv
df = pd.read_csv("https://storage.googleapis.com/dqlab-dataset/sample_csv.csv")
# Tampilkan 3 data teratas
print("Tiga data teratas:\n", df.head(3))
# Tampilkan 3 data terbawah
print("Tiga data terbawah:\n", df.tail(3))
# -
# # Indexing
# +
# Baca file TSV sample_tsv.tsv
df = pd.read_csv("https://storage.googleapis.com/dqlab-dataset/sample_tsv.tsv", sep="\t")
# Index dari df
print("Index : ", df.index)
# Column dari df
print("Columns : ", df.columns)
# +
# Set multi index df
df_x = df.set_index(['order_date', 'city','customer_id'])
# Print nama dan level dari multi index
for name, level in zip(df_x.index.names, df_x.index.levels):
print(name,':',level)
# -
# Cetak data frame awal
print("Dataframe awal:\n", df)
# Set index baru
df.index = ["Pesanan ke-" + str(i) for i in range(1, 11)]
# Cetak data frame dengan index baru
print("Dataframe dengan index baru:\n", df)
# Cetak data frame untuk 8 data teratas
print("Dataframe:\n", df.head(8))
# # Slicing
# +
# Baca file sample_csv.csv
df = pd.read_csv("https://storage.googleapis.com/dqlab-dataset/sample_csv.csv")
# Slice langsung berdasarkan kolom
df_slice = df.loc[(df["customer_id"] == "18055") &
(df["product_id"].isin(["P0029","P0040","P0041","P0116","P0117"]))
]
print("Slice langsung berdasarkan kolom:\n", df_slice)
# -
# Set index dari df sesuai instruksi
df = df.set_index(["order_date", "order_id","product_id"])
# Slice sesuai intruksi
df_slice = df.loc[("2019-01-01",1612339,["P2154","P2159"]),:]
print("Slice df:\n", df_slice)
# # Transforming
# +
# Baca file sample_csv.csv
df = pd.read_csv("https://storage.googleapis.com/dqlab-dataset/sample_csv.csv")
# Tampilkan tipe data
print("Tipe data df:\n", df.dtypes)
# Ubah tipe data kolom order_date menjadi datetime
df["order_date"] = pd.to_datetime(df["order_date"])
# Tampilkan tipe data df setelah transformasi
print("\nTipe data df setelah transformasi:\n", df.dtypes)
# +
# Tampilkan tipe data
print("Tipe data df:\n", df.dtypes)
# Ubah tipe data kolom quantity menjadi tipe data numerik float
df["quantity"] = pd.to_numeric(df["quantity"], downcast="float")
# Ubah tipe data kolom city menjadi tipe data category
df["city"] = df["city"].astype("category")
# Tampilkan tipe data df setelah transformasi
print("\nTipe data df setelah transformasi:\n", df.dtypes)
# +
# Cetak 5 baris teratas kolom brand
print("Kolom brand awal:\n", df["brand"].head())
# Gunakan method apply untuk merubah isi kolom menjadi lower case
df["brand"] = df["brand"].apply(lambda x: x.lower())
# Cetak 5 baris teratas kolom brand
print("Kolom brand setelah apply:\n", df["brand"].head())
# Gunakan method map untuk mengambil kode brand yaitu karakter terakhirnya
df["brand"] = df["brand"].map(lambda x: x[-1])
# Cetak 5 baris teratas kolom brand
print("Kolom brand setelah map:\n", df["brand"].head())
# +
# number generator, set angka seed menjadi suatu angka, bisa semua angka, supaya hasil random nya selalu sama ketika kita run
np.random.seed(1234)
# create dataframe 3 baris dan 4 kolom dengan angka random
df_tr = pd.DataFrame(np.random.rand(3,4))
# Cetak dataframe
print("Dataframe:\n", df_tr)
# Cara 1 dengan tanpa define function awalnya, langsung pake fungsi anonymous lambda x
df_tr1 = df_tr.applymap(lambda x: x**2+3*x+2)
print("\nDataframe - cara 1:\n", df_tr1)
# Cara 2 dengan define function
def qudratic_fun(x):
return x**2+3*x+2
df_tr2 = df_tr.applymap(qudratic_fun)
print("\nDataframe - cara 2:\n", df_tr2)
# -
# # Inspeksi Missing Value
# +
# Baca file "public data covid19 jhu csse eu.csv"
df = pd.read_csv("https://storage.googleapis.com/dqlab-dataset/CHAPTER%204%20-%20missing%20value%20-%20public%20data%20covid19%20.csv")
# Cetak info dari df
print(df.info())
# Cetak jumlah missing value di setiap kolom
mv = df.isna().sum()
print("\nJumlah missing value per kolom:\n", mv)
# -
# # Treatment untuk Missing Value
# +
# Cetak ukuran awal dataframe
print("Ukuran awal df: %d baris, %d kolom." % df.shape)
# Drop kolom yang seluruhnya missing value dan cetak ukurannya
df = df.dropna(axis=1, how="all")
print("Ukuran df setelah buang kolom dengan seluruh data missing: %d baris, %d kolom." % df.shape)
# Drop baris jika ada satu saja data yang missing dan cetak ukurannya
df = df.dropna(axis=0, how="any")
print("Ukuran df setelah dibuang baris yang memiliki sekurangnya 1 missing value: %d baris, %d kolom." % df.shape)
# +
# Cetak unique value pada kolom province_state
print("Unique value awal:\n", df["province_state"].unique())
# Ganti missing value dengan string "unknown_province_state"
df["province_state"] = df["province_state"].fillna("unknown_province_state")
# Cetak kembali unique value pada kolom province_state
print("Unique value setelah fillna:\n", df["province_state"].unique())
# +
# Cetak nilai mean dan median awal
print("Awal: mean = %f, median = %f." % (df["active"].mean(), df["active"].median()))
# Isi missing value kolom active dengan median
df_median = df["active"].fillna(df["active"].median())
# Cetak nilai mean dan median awal setelah diisi dengan median
print("Fillna median: mean = %f, median = %f." % (df_median.mean(), df_median.median()))
# Isi missing value kolom active dengan mean
df_mean = df["active"].fillna(df["active"].mean())
# Cetak nilai mean dan median awal setelah diisi dengan mean
print("Fillna mean: mean = %f, median = %f." % (df_mean.mean(), df_mean.median()))
# +
# Data
ts = pd.Series({
"2020-01-01":9,
"2020-01-02":np.nan,
"2020-01-05":np.nan,
"2020-01-07":24,
"2020-01-10":np.nan,
"2020-01-12":np.nan,
"2020-01-15":33,
"2020-01-17":np.nan,
"2020-01-16":40,
"2020-01-20":45,
"2020-01-22":52,
"2020-01-25":75,
"2020-01-28":np.nan,
"2020-01-30":np.nan
})
# Isi missing value menggunakan interpolasi linier
ts = ts.interpolate()
# Cetak time series setelah interpolasi linier
print("Setelah diisi missing valuenya:\n", ts)
# -
# # Project dari Andra
# +
# 1. Baca dataset
print("[1] BACA DATASET")
df = pd.read_csv("https://storage.googleapis.com/dqlab-dataset/retail_raw_test.csv", low_memory=False)
print(" Dataset:\n", df.head())
print(" Info:\n", df.info())
# 2. Ubah tipe data
print("\n[2] UBAH TIPE DATA")
df["customer_id"] = df["customer_id"].apply(lambda x: x.split("'")[1]).astype("int64")
df["quantity"] = df["quantity"].apply(lambda x: x.split("'")[1]).astype("int64")
df["item_price"] = df["item_price"].apply(lambda x: x.split("'")[1]).astype("int64")
print(" Tipe data:\n", df.dtypes)
# 3. Transform "product_value" supaya bentuknya seragam dengan format "PXXXX", assign ke kolom baru "product_id", dan drop kolom "product_value", jika terdapat nan gantilah dengan "unknown"
print("\n[3] TRANSFORM product_value MENJADI product_id")
# Buat fungsi
import math
def impute_product_value(val):
if math.isnan(val):
return "unknown"
else:
return 'P' + '{:0>4}'.format(str(val).split('.')[0])
# Buat kolom "product_id"
df["product_id"] = df["product_value"].apply(lambda x: impute_product_value(x))
# Hapus kolom "product_value"
df.drop(["product_value"], axis=1, inplace=True)
# Cetak 5 data teratas
print(df.head())
# 4. Tranform order_date menjadi value dengan format "YYYY-mm-dd"
print("\n[4] TRANSFORM order_date MENJADI FORMAT YYYY-mm-dd")
months_dict = {
"Jan":"01",
"Feb":"02",
"Mar":"03",
"Apr":"04",
"May":"05",
"Jun":"06",
"Jul":"07",
"Aug":"08",
"Sep":"09",
"Oct":"10",
"Nov":"11",
"Dec":"12"
}
df["order_date"] = pd.to_datetime(df["order_date"].apply(lambda x: str(x)[-4:] + "-" + months_dict[str(x)[:3]] + "-" + str(x)[4:7]))
print(" Tipe data:\n", df.dtypes)
# 5. Mengatasi data yang hilang di beberapa kolom
print("\n[5] HANDLING MISSING VALUE")
# Kolom "city" dan "province" masih memiliki missing value, nilai yang hilang di kedua kolom ini diisi saja dengan "unknown"
df[["city","province"]] = df[["city","province"]].fillna("unknown")
# Kolom brand juga masih memiliki missing value, Ganti value NaN menjadi "no_brand"
df["brand"] = df["brand"].fillna("no_brand")
# Cek apakah masih terdapat missing value di seluruh kolom
print(" Info:\n", df.info())
# 6. Membuat kolom baru "city/province" dengan menggabungkan kolom "city" dan kolom "province" dan delete kolom asalnya
print("\n[6] MEMBUAT KOLOM BARU city/province")
df["city/province"] = df["city"] + "/" + df["province"]
# drop kolom "city" dan "province" karena telah digabungkan
df.drop(["city","province"], axis=1, inplace=True)
# Cetak 5 data teratas
print(df.head())
# 7. Membuat hierarchical index yang terdiri dari kolom "city/province", "order_date", "customer_id", "order_id", "product_id"
print("\n[7] MEMBUAT HIERACHICAL INDEX")
df = df.set_index(["city/province","order_date","customer_id","order_id","product_id"])
# urutkanlah berdasarkan index yang baru
df = df.sort_index()
# Cetak 5 data teratas
print(df.head())
# 8. Membuat kolom "total_price" yang formula nya perkalian antara kolom "quantity" dan kolom "item_price"
print("\n[8] MEMBUAT KOLOM total_price")
df["total_price"] = df["quantity"] * df["item_price"]
# Cetak 5 data teratas
print(df.head())
# 9. Slice dataset agar hanya terdapat data bulan Januari 2019
print("\n[9] SLICE DATASET UNTUK BULAN JANUARI 2019 SAJA")
idx = pd.IndexSlice
df_jan2019 = df.loc[idx[:, "2019-01-01":"2019-01-31"], :]
print("Dataset akhir:\n", df_jan2019)
# END OF PROJECT
# -
|
My Class/Python/Fundamental/Data Manipulation with Pandas - Part 1/Data Manipulation with Pandas - Part 1.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# ## Reinforcement Learning (Delta Rule)
# ##### *Packages and Configurations*
using Plots, LaTeXStrings
cd(dirname(@__FILE__))
# ω -> ω + ϵδu
# δ = r - ν
# ν : The expected reward (in this problem it's either 1 or 0), r : The actual reward,
# ϵ : Learning rate (= 0.05), u : presence or absence of the stimuli(in this problem it always equals 1).
# 200 trials, ω₀ = 0.0, α : probability of recieving the reward.
function DeltaRule(trials, ω, ϵ, u, r, ν , ωList)
for i in 1:trials
ν = ω * u
δ = r - ν
ω += ϵ * δ * u
push!(ωList, ω)
end
return ωList
end
# +
ϵ=0.05; ω₀=0.0; u=1.0; trials=200;
ν = 0
ω = ω₀
ωList=[]
ωList = DeltaRule(trials/2, ω, ϵ, u, 1, ν , ωList)
ωList = DeltaRule(trials/2, ωList[end], ϵ, u, 0, ν , ωList)
scatter(ωList, ms=3, color=:black, title=L"\alpha=1", xlabel=L"trial\ number", ylabel=L"\omega", dpi=400, legend=false)
savefig("../../Fundamentals-of-Neuroscience_Fall1400/Reinforcement_Learning/Figs/A.pdf")
# -
# ##### *α =1*
function DeltaRuleP(trials, ω, ϵ, u, ν , ωList, α)
for i in 1:trials
if α > rand()
r=1
ν = ω * u
δ = r - ν
ω += ϵ * δ * u
push!(ωList, ω)
else
r=0
ν = ω * u
δ = r - ν
ω += ϵ * δ * u
push!(ωList, ω)
end
end
return ωList
end
# +
ϵ=0.05; ω₀=0.0; u=1.0; trials=200
ν = 0
ω = ω₀
ωList=[]
ωList = DeltaRuleP(trials, ω, ϵ, u, ν , ωList, 1)
#ωList = [ωList1 ; ωList2]
scatter(ωList, ms=3, color=:black, title=L"\alpha=1.0", xlabel=L"trial\ number", ylabel=L"\omega", dpi=400, legend=false)
savefig("../../Fundamentals-of-Neuroscience_Fall1400/Reinforcement_Learning/Figs/B.1.pdf")
# -
# ##### *α = 0.25*
# +
ϵ=0.05; ω₀=0.0; u=1.0; trials=200
ν = 0
ω = ω₀
ωList=[]
ωList = DeltaRuleP(trials, ω, ϵ, u, ν , ωList, 0.25)
#ωList = [ωList1 ; ωList2]
scatter(ωList, ms=3, color=:black, title=L"\alpha=0.25", xlabel=L"trial\ number", ylabel=L"\omega", dpi=400, legend=false)
savefig("../../Fundamentals-of-Neuroscience_Fall1400/Reinforcement_Learning/Figs/B.2.pdf")
# -
# ##### *α = 0.75*
ν = 0
ω = ω₀
ωList=[]
ωList = DeltaRuleP(trials, ω, ϵ, u, ν , ωList, 0.75)
scatter(ωList, ms=3, color=:black, title=L"\alpha=0.75", xlabel=L"trial\ number", ylabel=L"\omega", dpi=400, legend=false)
savefig("../../Fundamentals-of-Neuroscience_Fall1400/Reinforcement_Learning/Figs/B.3.pdf")
# ##### *α = 0.5*
# +
ϵ=0.05; ω₀=0.0; u=1.0; trials=200;
ν = 0
ω = ω₀
ωList = []
ωList = DeltaRuleP(trials, ω, ϵ, u, ν , ωList, 0.5)
p1=scatter(ωList, ms=3, color=:black, title=L"\alpha=0.5", xlabel=L"trial\ number", ylabel=L"\omega", dpi=400, legend=false)
# -
# ##### *reward = 0.5*
# +
ϵ=0.05; ω₀=0.0; u=1.0; trials=200;
ν = 0
ω = ω₀
ωList=[]
ωList = DeltaRule(trials, ω, ϵ, u, 0.5, ν , ωList)
p2 = scatter(ωList, ms=3, color=:black, title=L"reward=0.5", xlabel=L"trial\ number", ylabel=L"\omega", dpi=400, legend=false)
plot(p1, p2, layout=(1,2))
savefig("../../Fundamentals-of-Neuroscience_Fall1400/Reinforcement_Learning/Figs/E.pdf")
|
Reinforcement_Learning/Reinforcement_Learning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.utils import shuffle
import numpy as np
import os
#os.sys.path
# +
import cv2
import utils
#This file is aim to process all the pictures in the training folder.
# -
def read_image(path):
image = cv2.imread(path)
image = cv2.resize(image, (utils.IMAGE_WIDTH, utils.IMAGE_HEIGHT))
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
normalizer_image = image / 255.0 - 0.5
return normalizer_image
# +
# The Path of the dataset
training=True
traffic_light_dir = "traffic_light_images/"
if training:
red = traffic_light_dir + "training/red/"
yellow = traffic_light_dir + "training/yellow/"
green = traffic_light_dir + "training/green/"
else:
red = traffic_light_dir + "test/red/"
yellow = traffic_light_dir + "test/yellow/"
green = traffic_light_dir + "test/green/"
# +
images = []
labels = []
image_name = []
for f in os.listdir(red):
images.append(read_image(red + f))
labels.append(utils.RED)
image_name.append(f)
for f in os.listdir(yellow):
images.append(read_image(yellow + f))
labels.append(utils.YELLOW)
image_name.append(f)
for f in os.listdir(green):
images.append(read_image(green + f))
labels.append(utils.GREEN)
image_name.append(f)
# -
shuffle(np.array(images), np.array(labels), np.array(image_name))
|
Traffic_light_class_Image_process .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# <img src="../../images/qiskit-heading.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# ## _*The Bernstein-Vazirani Algorithm*_
#
# In this tutorial, we introduce the [Bernstein-Vazirani algorithm](http://epubs.siam.org/doi/abs/10.1137/S0097539796300921), which is one of the earliest algorithms demonstrating the power of quantum computing. Despite its simplicity, it is often used and is the inspiration for many other quantum algorithms even today; it is the basis of the power of the short-depth quantum circuits, as in [Bravyi et al.](https://arxiv.org/abs/1704.00690) that uses its non-oracular version, or in [Linke et al.](http://www.pnas.org/content/114/13/3305.full) that uses it to test the performance of the quantum processors (see also the [talk by Ken Brown](https://www.youtube.com/watch?v=eHV9LTiePrQ) at the ThinkQ 2017 conference). Here, we show the implementation of the Bernstein-Vazirani algorithm **without using entanglement** based on [Du et al.](https://arxiv.org/abs/quant-ph/0012114) on Qiskit and test it on IBM Q systems.
#
# The latest version of this notebook is available on https://github.com/qiskit/qiskit-tutorial.
#
# ***
# ### Contributors
# <NAME>
# ## Introduction <a id='introduction'></a>
#
# The Bernstein-Vazirani algorithm deals with finding a hidden integer $a \in \{0,1\}^n$ from an oracle $f_a$ that returns a bit $a \cdot x \equiv \sum_i a_i x_i \mod 2$ upon receiving an input $x \in \{0,1\}^n$. A classical oracle returns $f_a(x) = a \cdot x \mod 2$ given an input $x$. Meanwhile, a quantum oracle behaves similarly but can be queried with superposition of input $x$'s.
#
# Classically, the hidden integer $a$ can be revealed by querying the oracle with $x = 1, 2, \ldots, 2^i, \ldots, 2^{n-1}$, where each query reveals the $i$-th bit of $a$ (or, $a_i$). For example, with $x=1$ one can obtain the least significant bit of $a$, and so on. This turns out to be an optimal strategy; any classical algorithm that finds the hidden integer with high probability must query the oracle $\Omega(n)$ times. However, given a corresponding quantum oracle, the hidden integer can be found with only $1$ query using the Bernstein-Vazirani algorithm.
# ## The Algorithm
#
# The Bernstein-Vazirani algorithm to find the hidden integer is very simple: start from a $|0\rangle$ state, apply Hadamard gates, query the oracle, apply Hadamard gates, and measure. The correctness of the algorithm is best explained by looking at the transformation of a quantum register $|a \rangle$ by $n$ Hadamard gates, each applied to the qubit of the register. It can be shown that
#
# $$
# |a\rangle \xrightarrow{H^{\otimes n}} \frac{1}{\sqrt{2^n}} \sum_{x\in \{0,1\}^n} (-1)^{a\cdot x}|x\rangle.
# $$
#
# In particular, when we start with a quantum register $|0\rangle$ and apply $n$ Hadamard gates to it, we have the familiar quantum superposition as below
#
# $$
# |0\rangle \xrightarrow{H^{\otimes n}} \frac{1}{\sqrt{2^n}} \sum_{x\in \{0,1\}^n} |x\rangle,
# $$
#
# which is slightly different from the Hadamard transform of the reqister $|a \rangle$ by the phase $(-1)^{a\cdot x}$.
#
# Now, the quantum oracle $f_a$ returns $1$ on input $x$ such that $a \cdot x \equiv 1 \mod 2$, and returns $0$ otherwise. This means we have the following transformation:
#
# $$
# |x \rangle \left(|0\rangle - |1\rangle \right) \xrightarrow{f_a} | x \rangle \left(|0 \oplus f_a(x) \rangle - |1 \oplus f_a(x) \rangle \right) = (-1)^{a\cdot x} |x \rangle \left(|0\rangle - |1\rangle \right).
# $$
#
# Notice that the second register $|0\rangle - |1\rangle$ in the above does not change and can be omitted for simplicity. In short, the oracle can be used to create $(-1)^{a\cdot x}|x\rangle$ from the input $|x \rangle$. In this tutorial, we follow [Du et al.](https://arxiv.org/abs/quant-ph/0012114) to generate a circuit for a quantum oracle without the need of an ancilla qubit (often used in the standard quantum oracle).
#
# The algorithm to reveal the hidden integer follows naturally by querying the quantum oracle $f_a$ with the quantum superposition obtained from the Hadamard transformation of $|0\rangle$. Namely,
#
# $$
# |0\rangle \xrightarrow{H^{\otimes n}} \frac{1}{\sqrt{2^n}} \sum_{x\in \{0,1\}^n} |x\rangle \xrightarrow{f_a} \frac{1}{\sqrt{2^n}} \sum_{x\in \{0,1\}^n} (-1)^{a\cdot x}|x\rangle.
# $$
#
# Because the inverse of the $n$ Hadamard gates is again the $n$ Hadamard gates, we can obtain $a$ by
#
# $$
# \frac{1}{\sqrt{2^n}} \sum_{x\in \{0,1\}^n} (-1)^{a\cdot x}|x\rangle \xrightarrow{H^{\otimes n}} |a\rangle.
# $$
#
# ## The (Inner-Product) Oracle <a id='oracle'></a>
#
# Here, we describe how to build the oracle used in the Bernstein-Vazirani algorithm. The oracle is also referred to as the [inner-product oracle](https://arxiv.org/pdf/quant-ph/0108095.pdf) (while the oracle of the Grover search is known as the Equivalence, or EQ, oracle). Notice that it transforms $|x\rangle$ into $(-1)^{a\cdot x} |x\rangle$. Clearly, we can observe that
#
# $$
# (-1)^{a\cdot x} = (-1)^{a_1 x_1} \ldots (-1)^{a_ix_i} \ldots (-1)^{a_nx_n} = \prod_{i: a_i = 1} (-1)^{x_i}.
# $$
#
# Therefore, the inner-product oracle can be realized by the following unitary transformation, which is decomposable as single-qubit unitaries:
#
# $$
# O_{f_a} = O^1 \otimes O^2 \otimes \ldots \otimes O^i \otimes \ldots \otimes O^n,
# $$
# where $O^i = (1 - a_i)I + a_i Z$, where $Z$ is the Pauli $Z$ matrix and $I$ is the identity matrix for $a_i \in \{0,1\}$.
#
# Notice that we start from a separable quantum state $|0\rangle$ and apply a series of transformations that are separable (i.e., can be described by unitaries acting on a single qubit): Hadamard gates to each qubit, followed by the call to the *decomposable* quantum oracle as [Du et al.](https://arxiv.org/abs/quant-ph/0012114), and another Hadamard gate. Hence, there is no entanglement created during the computation. This is in contrast with the circuit at [Linke et al.](http://www.pnas.org/content/114/13/3305.full) that used CNOT gates to realize the oracle and an ancilla qubit to store the answer of the oracle.
# ## The Circuit <a id="circuit"></a>
#
# We now implement the Bernstein-Vazirani algorithm with Qiskit by first preparing the environment.
# +
#initialization
import sys, getpass
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
# importing Qiskit
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister,
from qiskit import available_backends, execute, register, get_backend
try:
sys.path.append("../../") # go to parent dir
import Qconfig
qx_config = {
"APItoken": Qconfig.APItoken,
"url": Qconfig.config['url']}
print('Qconfig loaded from %s.' % Qconfig.__file__)
except:
APItoken = getpass.getpass('Please input your token and hit enter: ')
qx_config = {
"APItoken": APItoken,
"url":"https://quantumexperience.ng.bluemix.net/api"}
print('Qconfig.py not found in qiskit-tutorial directory; Qconfig loaded using user input.')
# import basic plot tools
from qiskit.tools.visualization import plot_histogram
# -
# We first set the number of qubits used in the experiment, and the hidden integer $a$ to be found by the Bernstein-Vazirani algorithm. The hidden integer $a$ determines the circuit for the quantum oracle.
# +
nQubits = 16 # number of physical qubits
a = 101 # the hidden integer whose bitstring is 1100101
# make sure that a can be represented with nQubits
a = a % 2**(nQubits)
# -
# We then use Qiskit to program the Bernstein-Vazirani algorithm.
# +
# Creating registers
# qubits for querying the oracle and finding the hidden integer
qr = QuantumRegister(nQubits)
# for recording the measurement on qr
cr = ClassicalRegister(nQubits)
circuitName = "BernsteinVazirani"
bvCircuit = QuantumCircuit(qr, cr)
# Apply Hadamard gates before querying the oracle
for i in range(nQubits):
bvCircuit.h(qr[i])
# Apply barrier so that it is not optimized by the compiler
bvCircuit.barrier()
# Apply the inner-product oracle
for i in range(nQubits):
if (a & (1 << i)):
bvCircuit.z(qr[i])
else:
bvCircuit.iden(qr[i])
# Apply barrier
bvCircuit.barrier()
#Apply Hadamard gates after querying the oracle
for i in range(nQubits):
bvCircuit.h(qr[i])
# Measurement
for i in range(nQubits):
bvCircuit.measure(qr[i], cr[i])
# -
# ## Experiment with Simulators
#
# We can run the above circuit on the simulator.
# +
# use local simulator
backend = "local_qasm_simulator"
shots = 1000
results = execute(bvCircuit, backend=backend, shots=shots).result()
answer = results.get_counts()
plot_histogram(answer)
# -
# We can see that the result of the measurement is the binary representation of the hidden integer $a$.
#
# ## Experiment with Real Devices
#
# We can run the circuit on the real device as below.
# +
#to enable sleep
import time
#connect to remote API to be able to use remote simulators and real devices
register(qx_config['APItoken'], qx_config['url'])
print("Available backends:", available_backends())
# to run on remote simulator
backend = "ibmq_qasm_simulator"
# uncomment below to run on ibmqx5
# backend = "ibmqx5"
shots = 1000
if get_backend(backend).status["operational"] is True:
job_exp = execute(bvCircuit, backend=backend, shots=shots)
lapse = 0
interval = 10
while not job_exp.done:
print('Status @ {} seconds'.format(interval * lapse))
print(job_exp.status)
time.sleep(interval)
lapse += 1
print(job_exp.status)
results = job_exp.result()
answer = results.get_counts(bvCircuit)
threshold = int(0.03 * shots) #the threshold of plotting significant measurements
filteredAnswer = {k: v for k,v in answer.items() if v >= threshold} #filter the answer for better view of plots
removedCounts = np.sum([ v for k,v in answer.items() if v < threshold ]) #number of counts removed
filteredAnswer['other_bitstring'] = removedCounts #the removed counts is assigned to a new index
plot_histogram(filteredAnswer)
print(filteredAnswer)
# -
# We indeed see that the outcome is the binary representation of the hidden integer $a$ with high probability.
|
reference/algorithms/bernstein_vazirani.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pylab
# +
file0 = "/home/keithchow/data/recordings/sorted/" + \
"FM_RBDS_20200226_21-59-50__fc_106.9000_MHz_fs_600.0000_ksps.32cf"
signal = np.fromfile(file0, dtype=np.complex64)
pylab.specgram(signal, Fs=600e3)
pylab.show()
# -
# # RDS
# 1187.5 bits/s
# BPSK
# 57 kHz subcarrier
# # HD Radio (NRSC-5)
# OFDM signal
#
# +/- 70 kHz past the 200 kHz analog signal +/- 30 kHz guard band
#
# Subcarrier Spacing = 1488375 / 4096
# Cyclic Prefix = 7 / 128
# OFDM symbol duration = (135 / 128) * (4096 / 1488375)
#
|
communications/FM_Radio.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os, os.path
import numpy as np
import pandas as pd
import data_structures as ds
import setup_analysis as sa
import support_functions as sf
import importlib
import time
import warnings
import matplotlib.pyplot as plt
importlib.reload(ds)
importlib.reload(sa)
importlib.reload(sf)
import model_afolu as ma
importlib.reload(ma)
df_fake_data_base_2 = df_fake_data_base.copy()
df_fake_data_base_2 = df_fake_data_base_2[df_fake_data_base_2["time_period"] < 9]
# +
df_fake_data_base_3 = df_fake_data_base_2.copy()
df_fake_data_base_3 = df_fake_data_base_3[~df_fake_data_base_3["time_period"].isin([3, 4])]
df_fake_data_base_3b = df_fake_data_base_3.copy()
df_fake_data_base_3["strategy_id"] = 0
df_fake_data_base_3b["strategy_id"] = 1
#df_fake_data_base_3 = pd.concat([df_fake_data_base_3, df_fake_data_base_3b], axis = 0).reset_index(drop = True)
#df_fake_data_base_3
# +
#sa.model_attributes.check_projection_input_df(df_fake_data_base_3)#[[x for x in df_fake_data_base_3.columns if x != sa.model_attributes.dim_time_period]])[1]
# -
##
importlib.reload(sa)
# +
importlib.reload(ds)
importlib.reload(ma)
#pd.DataFrame(a2).to_csv(os.path.join("example_transition_matrix.csv"), index = None, encoding = "UTF-8")
a2 = pd.read_csv(os.path.join(sa.dir_ref, "example_transition_matrix.csv"))
a2 = np.array(a2);
df_fake_data_base = pd.read_csv(os.path.join(sa.dir_ref, "fake_data", "fake_data_afolu.csv"))
df_fake_data_high_veg = pd.read_csv(os.path.join(sa.dir_ref, "fake_data", "fake_data_afolu-high_veg.csv"))
df_fake_data_high_veg_norealloc = pd.read_csv(os.path.join(sa.dir_ref, "fake_data", "fake_data_afolu-high_veg_norealloc.csv"))
df_fake_data_high_veg_allrealloc = pd.read_csv(os.path.join(sa.dir_ref, "fake_data", "fake_data_afolu-high_veg_allrealloc.csv"))
df_fake_data_high_yield = pd.read_csv(os.path.join(sa.dir_ref, "fake_data", "fake_data_afolu-high_yields.csv"))
model_afolu = ma.AFOLU(sa.model_attributes)
df_out_base = model_afolu.project(df_fake_data_base)
df_out_high_veg = model_afolu.project(df_fake_data_high_veg)
df_out_high_veg_norealloc = model_afolu.project(df_fake_data_high_veg_norealloc)
df_out_high_veg_allrealloc = model_afolu.project(df_fake_data_high_veg_allrealloc)
df_fake_data_high_yield_test = df_fake_data_high_yield.iloc[0:10].copy()
df_fake_data_high_yield_test["time_period"].iloc[9] = 13
df_out_high_yield = model_afolu.project(df_fake_data_high_yield_test)
#df_emissions_conv, arr_emissions_conv, arr_land_use, arrs_land_conv = model_afolu.project(df_fake_data)
#df_afolu = model_afolu.project(df_fake_data)
# -
#df_out_high_yield[[x df_fake_data_high_yield_testfor x in df_out_high_yield.columns if "lvst" in x]]
df_out_high_yield
# +
cols = [x for x in df_out.columns if "area_agrc" in x]
cg = "area_lndu_grasslands_ha"
fig, ax = plt.subplots(2, 2, figsize = (20, 20))
l_1 = 1500000
for col in cols:
x = np.array(df_out_base["time_period"])
y = np.array(df_out_base[col])
y2 = np.array(df_out_base[cg])
ax[0, 0].plot(x, y, label = col.replace("area_agrc_crops_", "").replace("_ha", ""))
if col == cols[0]:
ax[0, 0].plot(x, y2, c = "black", linewidth = 3, label = "grassland")
ax[0, 0].set_title("Base (0.5 realloc factor)")
ax[0, 0].legend()
ax[0, 0].set_ylim((0, l_1))
x = np.array(df_out_high_veg["time_period"])
y = np.array(df_out_high_veg[col])
y2 = np.array(df_out_high_veg[cg])
ax[0, 1].plot(x, y)
if col == cols[0]:
ax[0, 1].plot(x, y2, c = "black", linewidth = 3, label = "grassland")
ax[0, 1].set_title("High Veg (0.5 realloc factor)")
ax[0, 1].set_ylim((0, l_1))
x = np.array(df_out_high_veg_norealloc["time_period"])
y = np.array(df_out_high_veg_norealloc[col])
y2 = np.array(df_out_high_veg_norealloc[cg])
ax[1, 0].plot(x, y)
if col == cols[0]:
ax[1, 0].plot(x, y2, c = "black", linewidth = 3, label = "grassland")
ax[1, 0].set_title("High Veg (0 realloc factor)")
ax[1, 0].set_ylim((0, l_1))
x = np.array(df_out_high_yield["time_period"])
y = np.array(df_out_high_yield[col])
y2 = np.array(df_out_high_yield[cg])
ax[0, 0].plot(x, y, alpha = 0.5, linewidth = 1)
if col == cols[0]:
ax[0, 0].plot(x, y2, c = "black", linewidth = 1.5, alpha = 0.4)
ax[0, 0].set_title("High Yield (0.5 realloc factor)")
ax[0, 0].set_ylim((0, l_1))
x = np.array(df_out_high_veg_allrealloc["time_period"])
y = np.array(df_out_high_veg_allrealloc[col])
y2 = np.array(df_out_high_veg_allrealloc[cg])
ax[1, 1].plot(x, y)
if col == cols[0]:
ax[1, 1].plot(x, y2, c = "black", linewidth = 3, label = "grassland")
ax[1, 1].set_title("High Veg (1 realloc factor)")
ax[1, 1].set_ylim((0, l_1))
plt.show()
# -
model_afolu.mat_ef[0][2,:]*model_afolu.arrs_land_conv[0][2,:]
# +
#list(zip(df_out.columns, list(df_out.iloc[20,:])))
df_in = df_out.copy()
list_subsectors = model_afolu.required_base_subsectors
for subsector in list_subsectors:#self.required_base_subsectors:
vars_subsec = model_afolu.model_attributes.dict_model_variables_by_subsector[subsector]
# add subsector abbreviation
fld_nam = model_afolu.model_attributes.get_subsector_attribute(subsector, "abv_subsector")
fld_nam = f"emission_co2e_subsector_total_{fld_nam}"
flds_add = []
for var in vars_subsec:
var_type = model_afolu.model_attributes.get_variable_attribute(var, "variable_type").lower()
gas = model_afolu.model_attributes.get_variable_characteristic(var, model_afolu.model_attributes.varchar_str_emission_gas)
if (var_type == "output") and gas:
flds_add += model_afolu.model_attributes.dict_model_variables_to_variables[var]
# check for missing fields; notify
missing_fields = [x for x in flds_add if x not in df_in.columns]
if len(missing_fields) > 0:
str_mf = print_setdiff(set(df_in.columns), set(flds_add))
str_mf = f"Missing fields {str_mf}.%s"
if stop_on_missing_fields_q:
raise ValueError(str_mf%(" Subsector emission totals will not be added."))
else:
warnings.warn(str_mf%(" Subsector emission totals will exclude these fields."))
# -
df_in[flds_add].sum(axis = 1)
# +
# check for internal variables and add if necessary; note, this can be defined for different variables (see model attributes)
df_afolu_trajectories = df_fake_data.copy()
model_afolu.model_attributes.manage_pop_to_df(df_afolu_trajectories, "add")
df_afolu_trajectories.sort_values(by = [model_afolu.model_attributes.dim_time_period], inplace = True)
# check that all required fields are contained—assume that it is ordered by time period
model_afolu.check_df_fields(df_afolu_trajectories)
## CATEGORY INITIALIZATION
pycat_agrc = model_afolu.model_attributes.get_subsector_attribute("Agriculture", "pycategory_primary")
pycat_frst = model_afolu.model_attributes.get_subsector_attribute("Forest", "pycategory_primary")
pycat_lndu = model_afolu.model_attributes.get_subsector_attribute("Land Use", "pycategory_primary")
pycat_lvst = model_afolu.model_attributes.get_subsector_attribute("Livestock", "pycategory_primary")
# attribute tables
attr_agrc = model_afolu.model_attributes.dict_attributes[pycat_agrc]
attr_frst = model_afolu.model_attributes.dict_attributes[pycat_frst]
attr_lndu = model_afolu.model_attributes.dict_attributes[pycat_lndu]
attr_lvst = model_afolu.model_attributes.dict_attributes[pycat_lvst]
## FIELD INITIALIZATION
# get the gdp and total population fields
field_gdp = model_afolu.model_attributes.build_varlist("Economy", variable_subsec = model_afolu.modvar_econ_gdp)[0]
field_pop = model_afolu.model_attributes.build_varlist("General", variable_subsec = model_afolu.modvar_gnrl_pop_total)[0]
## ECON/GNRL VECTOR AND ARRAY INITIALIZATION
# get some vectors
vec_gdp = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_econ_gdp, False, return_type = "array_base")#np.array(df_afolu_trajectories[field_gdp])
vec_pop = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_gnrl_pop_total, False, return_type = "array_base")
vec_gdp_per_capita = vec_gdp/vec_pop
# growth rates
vec_rates_gdp = vec_gdp[1:]/vec_gdp[0:-1] - 1
vec_rates_gdp_per_capita = vec_gdp_per_capita[1:]/vec_gdp_per_capita[0:-1] - 1
## OUTPUT INITIALIZATION
df_out = [df_afolu_trajectories[model_afolu.required_dimensions].copy()]
# common indices
cat_lndu_ind_pstr = model_afolu.model_attributes.dict_attributes["cat_landuse"].get_key_value_index("grasslands")
cat_lndu_ind_crop = model_afolu.model_attributes.dict_attributes["cat_landuse"].get_key_value_index("croplands")
# area of the country
area = float(model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_gnrl_area, return_type = "array_base")[0])
# get the initial distribution of land
vec_modvar_lndu_initial_frac = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_lndu_initial_frac, return_type = "array_base")[0]
vec_modvar_lndu_initial_area = vec_modvar_lndu_initial_frac*area
model_afolu.vec_modvar_lndu_initial_area = vec_modvar_lndu_initial_area
model_afolu.mat_trans_unadj, model_afolu.mat_ef = model_afolu.get_markov_matrices(df_afolu_trajectories)
# get some land use variables, including the land use reallocation factor
vec_lndu_yield_realloc_factor = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_lndu_reallocation_factor, False, "array_base", var_bounds = (0, 1))
###########################
# CALCULATE DEMANDS #
###########################
## livestock demands (calculated exogenously)
# variables requried to estimate demand
vec_modvar_lvst_pop_init = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_lvst_pop_init, True, "array_base")[0]
fields_lvst_elas = model_afolu.model_attributes.switch_variable_category("Livestock", model_afolu.modvar_lvst_elas_lvst_demand, "demand_elasticity_category")
arr_lvst_elas_demand = np.array(df_afolu_trajectories[fields_lvst_elas])
# get the "vegetarian" factor and use to estimate livestock pop
vec_lvst_demscale = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_lvst_frac_eating_red_meat, False, "array_base", var_bounds = (0, np.inf))
#vec_lvst_demscale = np.arange(36)*0.015
arr_lvst_dem_pop = model_afolu.project_per_capita_demand(vec_modvar_lvst_pop_init, vec_pop, vec_rates_gdp_per_capita, arr_lvst_elas_demand, vec_lvst_demscale, int)
# get weights for allocating grazing area and feed requirement to animals - based on first year only
vec_lvst_base_graze_weights = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_lvst_dry_matter_consumption, True, "array_base")[0]
vec_lvst_feed_allocation_weights = (vec_modvar_lvst_pop_init*vec_lvst_base_graze_weights)/np.dot(vec_modvar_lvst_pop_init, vec_lvst_base_graze_weights)
# get information used to calculate carrying capacity of land
vec_lvst_carry_capacity_scale = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_lvst_carrying_capacity_scalar, False, "array_base", var_bounds = (0, np.inf))
## agricultural demands
# variables required for demand
arr_agrc_elas_crop_demand = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_agrc_elas_crop_demand_income, False, "array_base")
arr_agrc_frac_feed = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_agrc_frac_animal_feed, False, "array_base")
# get initial cropland area
area_agrc_cropland_init = area*vec_modvar_lndu_initial_frac[cat_lndu_ind_crop]
vec_agrc_frac_cropland_area = model_afolu.check_cropland_fractions(df_afolu_trajectories)[0]
vec_agrc_cropland_area = area_agrc_cropland_init*vec_agrc_frac_cropland_area
# estimate yield capacity
arr_agrc_yf = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_agrc_yf, True, "array_base")
vec_agrc_yield_init = arr_agrc_yf[0]*vec_agrc_cropland_area
# split into yield for livestock feed (responsive to changes in livestock population) and yield for consumption and export (not)
vec_agrc_yield_init_lvstfeed = vec_agrc_yield_init*arr_agrc_frac_feed[0]
vec_agrc_yield_init_nonlvstfeed = vec_agrc_yield_init - vec_agrc_yield_init_lvstfeed
# project ag demand for crops that are driven by gdp/capita - set demand scalar for crop demand (increases based on reduction in red meat demand) - depends on how many people eat red meat (vec_lvst_demscale)
vec_agrc_diet_exchange_scalar = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_lndu_vdes, False, "array_base", var_bounds = (0, np.inf))
vec_agrc_demscale = vec_lvst_demscale + vec_agrc_diet_exchange_scalar - vec_lvst_demscale*vec_agrc_diet_exchange_scalar
arr_agrc_nonfeeddem_yield = model_afolu.project_per_capita_demand(vec_agrc_yield_init_nonlvstfeed, vec_pop, vec_rates_gdp_per_capita, arr_agrc_elas_crop_demand, vec_agrc_demscale, float)
# array gives the total yield of crop type i allocated to livestock type j at time 0
arr_lndu_yield_i_reqd_lvst_j_init = np.outer(vec_agrc_yield_init_lvstfeed, vec_lvst_feed_allocation_weights)
model_afolu.project_integrated_land_use(
vec_modvar_lndu_initial_area,
model_afolu.mat_trans_unadj,
model_afolu.mat_ef,
arr_agrc_nonfeeddem_yield,
arr_agrc_yf,
arr_lndu_yield_i_reqd_lvst_j_init,
arr_lvst_dem_pop,
vec_agrc_frac_cropland_area,
vec_lndu_yield_realloc_factor,
vec_modvar_lvst_pop_init,
vec_lvst_feed_allocation_weights,
vec_lvst_carry_capacity_scale
)
# -
# +
# project land use
vec_modvar_lndu_initial_frac = model_afolu.model_attributes.get_standard_variables(df_afolu_trajectories, model_afolu.modvar_lndu_initial_frac, return_type = "array_base")[0]
vec_initial_area = vec_modvar_lndu_initial_area
arrs_transitions = model_afolu.mat_trans_unadj
arrs_efs = model_afolu.mat_ef
# new vars to add to arg
arr_dem_lvst = arr_lvst_dem_pop
arr_agrc_nonfeeddem_yield = arr_agrc_nonfeeddem_yield
arr_lndu_yield_by_lvst = arr_lndu_yield_i_reqd_lvst_j_init
vec_agrc_frac_cropland_area = vec_agrc_frac_cropland_area
vec_lndu_yrf = vec_lndu_yield_realloc_factor
vec_lvst_pop_init = vec_modvar_lvst_pop_init
vec_lvst_pstr_weights = vec_lvst_feed_allocation_weights
vec_lvst_scale_cc = vec_lvst_carry_capacity_scale
def project_integrated_land_use(self,
vec_initial_area: np.ndarray,
arrs_transitions: np.ndarray,
arrs_efs: np.ndarray,
arr_dem_lvst: np.ndarray,
arr_agrc_nonfeeddem_yield: np.ndarray,
arr_lndu_yield_by_lvst: np.ndarray,
vec_agrc_frac_cropland_area: np.ndarray,
vec_lndu_yrf: np.ndarray,
vec_lvst_pop_init: np.ndarray,
vec_lvst_pstr_weights: np.ndarray,
vec_lvst_scale_cc: np.ndarray
):
t0 = time.time()
# check shapes
model_afolu.check_markov_shapes(arrs_transitions, "arrs_transitions")
model_afolu.check_markov_shapes(arrs_efs, "arrs_efs")
# get attributes
pycat_agrc = model_afolu.model_attributes.get_subsector_attribute("Agriculture", "pycategory_primary")
attr_agrc = model_afolu.model_attributes.dict_attributes[pycat_agrc]
pycat_lndu = model_afolu.model_attributes.get_subsector_attribute("Land Use", "pycategory_primary")
attr_lndu = model_afolu.model_attributes.dict_attributes[pycat_lndu]
pycat_lvst = model_afolu.model_attributes.get_subsector_attribute("Livestock", "pycategory_primary")
attr_lvst = model_afolu.model_attributes.dict_attributes[pycat_lvst]
# set some commonly called attributes and indices in arrays
m = attr_lndu.n_key_values
ind_crop = attr_lndu.get_key_value_index("croplands")
ind_pstr = attr_lndu.get_key_value_index("grasslands")
# initialize variables
arr_dem_lvst_gr = np.cumprod(arr_dem_lvst/arr_dem_lvst[0], axis = 0)
vec_lvst_cc_init = vec_lvst_pop_init/(vec_initial_area[ind_pstr]*vec_lvst_pstr_weights)
# intilize output arrays, including land use, land converted, emissions, and adjusted transitions
arr_agrc_frac_cropland = np.array([vec_agrc_frac_cropland_area for k in range(model_afolu.n_time_periods)])
arr_agrc_net_import_increase = np.zeros((model_afolu.n_time_periods, attr_agrc.n_key_values))
arr_emissions_conv = np.zeros((model_afolu.n_time_periods, attr_lndu.n_key_values))
arr_land_use = np.zeros((model_afolu.n_time_periods, attr_lndu.n_key_values))
arr_lvst_net_import_increase = np.zeros((model_afolu.n_time_periods, attr_lvst.n_key_values))
arrs_land_conv = np.zeros((model_afolu.n_time_periods, attr_lndu.n_key_values, attr_lndu.n_key_values))
arrs_transitions_adj = np.zeros(arrs_transitions.shape)
arrs_yields_per_livestock = np.array([arr_lndu_yield_i_reqd_lvst_j_init for k in range(model_afolu.n_time_periods)])
# modify input arrays for loop
# initialize running matrix of land use and iteration index i
x = vec_initial_area
i = 0
while i < model_afolu.n_time_periods:
# check emission factor index
i_ef = i if (i < len(arrs_efs)) else len(arrs_efs) - 1
if i_ef != i:
print(f"No emission factor matrix found for time period {model_afolu.time_periods[i]}; using the matrix from period {len(arrs_efs) - 1}.")
# check transition matrix index
i_tr = i if (i < len(arrs_transitions)) else len(arrs_transitions) - 1
if i_tr != i:
print(f"No transition matrix found for time period {model_afolu.time_periods[i]}; using the matrix from period {len(arrs_efs) - 1}.")
# calculate the unadjusted land use areas (projected to time step i + 1)
area_crop_cur = x[ind_crop]
area_pstr_cur = x[ind_pstr]
vec_agrc_cropland_area_cur = area_crop_cur*arr_agrc_frac_cropland[i]
# LIVESTOCK - calculate carrying capacities, demand used for pasture reallocation, and net surplus
vec_lvst_cc_cur = vec_lvst_scale_cc[i]*vec_lvst_cc_init
vec_lvst_prod_cur = vec_lvst_cc_cur*area_pstr_cur*vec_lvst_pstr_weights
vec_lvst_net_surplus = np.nan_to_num(arr_dem_lvst[i] - vec_lvst_prod_cur)
vec_lvst_reallocation = vec_lvst_net_surplus*vec_lndu_yrf[i] # demand for livestock met by reallocating land
vec_lvst_net_import_increase = vec_lvst_net_surplus - vec_lvst_reallocation # demand for livestock met by increasing net imports (neg => net exports)
# calculate required increase in transition probabilities
area_lndu_pstr_increase = sum(np.nan_to_num(vec_lvst_reallocation/vec_lvst_cc_cur))
scalar_lndu_pstr = (area_pstr_cur + area_lndu_pstr_increase)/np.dot(x, arrs_transitions[i_tr][:, ind_pstr])
# AGRICULTURE - calculate demand increase in crops, which is a function of gdp/capita (exogenous) and livestock demand (used for feed)
vec_agrc_feed_dem_yield = sum((arr_lndu_yield_by_lvst*arr_dem_lvst_gr[i]).transpose())
vec_agrc_dem_cropareas = (arr_agrc_nonfeeddem_yield[i] + vec_agrc_feed_dem_yield)/arr_agrc_yf[i]
vec_agrc_net_surplus_cropland_area_cur = vec_agrc_dem_cropareas - vec_agrc_cropland_area_cur
vec_agrc_reallocation = vec_agrc_net_surplus_cropland_area_cur*vec_lndu_yrf[i]
# get surplus yield (increase to net imports)
vec_agrc_net_imports_increase_yield = (vec_agrc_net_surplus_cropland_area_cur - vec_agrc_reallocation)*arr_agrc_yf[i]
vec_agrc_cropareas_adj = vec_agrc_cropland_area_cur + vec_agrc_reallocation
scalar_lndu_crop = sum(vec_agrc_cropareas_adj)/np.dot(x, arrs_transitions[i_tr][:, ind_crop])
# adjust the transition matrix
trans_adj = model_afolu.adjust_transition_matrix(arrs_transitions[i_tr], {(ind_pstr, ): scalar_lndu_pstr, (ind_crop, ): scalar_lndu_crop})
# calculate final land conversion and emissions
arr_land_conv = (trans_adj.transpose()*x.transpose()).transpose()
vec_emissions_conv = sum((trans_adj*arrs_efs[i_ef]).transpose()*x.transpose())
# update arrays
rng_agrc = list(range(i*attr_agrc.n_key_values, (i + 1)*attr_agrc.n_key_values))
np.put(arr_agrc_net_import_increase, rng_agrc, np.round(vec_agrc_net_imports_increase_yield), 2)
np.put(arr_agrc_frac_cropland, rng_agrc, vec_agrc_cropareas_adj/sum(vec_agrc_cropareas_adj))
# non-ag arrays
rng_put = np.arange(i*attr_lndu.n_key_values, (i + 1)*attr_lndu.n_key_values)
np.put(arr_land_use, rng_put, x)
np.put(arr_emissions_conv, rng_put, vec_emissions_conv)
arr_lvst_net_import_increase[i] = np.round(vec_lvst_net_import_increase).astype(int)
arrs_land_conv[i] = arr_land_conv
arrs_transitions_adj[i] = trans_adj
# update land use vector
x = np.matmul(x, trans_adj)
i += 1
return (
arr_agrc_frac_cropland,
arr_agrc_net_import_increase,
arr_emissions_conv,
arr_land_use,
arr_lvst_net_import_increase,
arrs_land_conv,
arrs_transitions_adj,
arrs_yields_per_livestock
)
t1 = time.time()
t_elapse = round(t1 - t0, 2)
print(f"Land use projection complete in {t_elapse} seconds.")
#return arr_emissions_conv, arr_land_use, arrs_land_conv
# -
arr_emissions_conv
np.cumprod((arr_dem_lvst[0:3]/arr_dem_lvst[0])[:, 2])
"""
a1 = np.round(arrs_transitions[i_tr], 5)
a2 = np.round(model_afolu.adjust_transition_matrix(arrs_transitions[i_tr], {(ind_pstr, ): scalar_lndu_pstr}), 5)
print("base:")
print(a1[:, ind_pstr])
print("\nAdjust with 0.5:")
print(a2[:, ind_pstr])
""";
arr_lvst_dem_gr
# +
#arrs_land_conv
#sum(arr_lvst_rqd_land.transpose())
ab = [1, 2, 5, 9, -1]
ab.index(5)
m = model_afolu.model_attributes.dict_attributes["cat_landuse"].n_key_values
# indices in arrays
ind_pstr = model_afolu.model_attributes.dict_attributes["cat_landuse"].get_key_value_index("grasslands")
ind_crop = model_afolu.model_attributes.dict_attributes["cat_landuse"].get_key_value_index("croplands")
# get totals that need to be scaled
vec_prelim_pstr = arr_land_use[1:, ind_pstr]
# -
# +
#inds = [x for x in range(len)]
np.round((arrs_land_conv[0].transpose()/sum(arrs_land_conv[0].transpose())).transpose(), 3)
# -
model_afolu.model_attributes.dict_attributes["cat_landuse"].key_values
# +
class AFOLU:
def __init__(self, attributes):
self.model_attributes = attributes
self.required_dimensions = self.get_required_dimensions()
self.required_subsectors, self.required_base_subsectors = self.get_required_subsectors()
self.required_variables, self.output_variables = self.get_afolu_input_output_fields()
## set some model fields to connect to the attribute tables
# agricultural model variables
self.modvar_agrc_area_prop = "Cropland Area Proportion"
self.modvar_agrc_area_crop = "Crop Area"
self.modvar_agrc_ef_ch4 = ":math:\\text{CH}_4 Crop Activity Emission Factor"
self.modvar_agrc_ef_n2o = ":math:\\text{CO}_2 Crop Activity Emission Factor"
self.modvar_agrc_ef_co2 = ":math:\\text{N}_2\\text{O} Crop Activity Emission Factor"
self.modvar_agrc_elas_crop_demand_income = "Crop Demand Income Elasticity"
self.modvar_agrc_emissions_ch4_crops = ":math:\\text{CH}_4 Emissions from Crop Activity"
self.modvar_agrc_emissions_co2_crops = ":math:\\text{CO}_2 Emissions from Crop Activity"
self.modvar_agrc_emissions_n2o_crops = ":math:\\text{N}_2\\text{O} Emissions from Crop Activity"
self.modvar_agrc_net_imports = "Crop Surplus Demand"
self.modvar_agrc_yf = "Crop Yield Factor"
self.modvar_agrc_yield = "Crop Yield"
# forest model variables
self.modvar_frst_elas_wood_demand = "Elasticity of Wood Products Demand to Value Added"
self.modvar_frst_ef_fires = "Forest Fire Emission Factor"
self.modvar_frst_ef_ch4 = "Forest Methane Emissions"
self.modvar_frst_emissions_sequestration = ":math:\\text{CO}_2 Emissions from Forest Sequestration"
self.modvar_frst_emissions_methane = ":math:\\text{CH}_4 Emissions from Forests"
self.modvar_frst_sq_co2 = "Forest Sequestration Emission Factor"
# land use model variables
self.modvar_lndu_area_by_cat = "Land Use Area"
self.modvar_lndu_ef_co2_conv = ":math:\\text{CO}_2 Land Use Conversion Emission Factor"
self.modvar_lndu_emissions_conv = ":math:\\text{CO}_2 Emissions from Land Use Conversion"
self.modvar_lndu_emissions_ch4_from_wetlands = ":math:\\text{CH}_4 Emissions from Wetlands"
self.modvar_lndu_emissions_n2o_from_pastures = ":math:\\text{N}_2\\text{O} Emissions from Pastures"
self.modvar_lndu_emissions_co2_from_pastures = ":math:\\text{CO}_2 Emissions from Pastures"
self.modvar_lndu_initial_frac = "Initial Land Use Area Proportion"
self.modvar_lndu_ef_ch4_boc = "Land Use BOC :math:\\text{CH}_4 Emission Factor"
self.modvar_lndu_ef_n2o_past = "Land Use Pasture :math:\\text{N}_2\\text{O} Emission Factor"
self.modvar_lndu_ef_co2_soilcarb = "Land Use Soil Carbon :math:\\text{CO}_2 Emission Factor"
self.modvar_lndu_prob_transition = "Unadjusted Land Use Transition Probability"
# livestock model variables
self.modvar_lvst_carrying_capacity_scalar = "Carrying Capacity Scalar"
self.modvar_lvst_dry_matter_consumption = "Daily Dry Matter Consumption"
self.modvar_lvst_ef_ch4_ef = ":math:\\text{CH}_4 Enteric Fermentation Emission Factor"
self.modvar_lvst_ef_ch4_mm = ":math:\\text{CH}_4 Manure Management Emission Factor"
self.modvar_lvst_ef_n2o_mm = ":math:\\text{N}_2\\text{O} Manure Management Emission Factor"
self.modvar_lvst_elas_lvst_demand = "Elasticity of Livestock Demand to GDP per Capita"
self.modvar_lvst_emissions_ch4_ef = ":math:\\text{CH}_4 Emissions from Livestock Enteric Fermentation"
self.modvar_lvst_emissions_ch4_mm = ":math:\\text{CH}_4 Emissions from Livestock Manure"
self.modvar_lvst_emissions_n2o_mm = ":math:\\text{N}_2\\text{O} Emissions from Livestock Manure"
self.modvar_lvst_frac_meat_import = "Fraction of Meat Consumption from Imports"
self.modvar_lvst_meat_demand_scalar = "Red Meat Demand Scalar"
self.modvar_lvst_net_imports = "Livestock Surplus Demand"
self.modvar_lvst_pop = "Livestock Head Count"
self.modvar_lvst_pop_init = "Initial Livestock Head Count"
# economy and general variables
self.modvar_econ_gdp = "GDP"
self.modvar_econ_va = "Value Added"
self.modvar_gnrl_area = "Area of Country"
self.modvar_gnrl_occ = "National Occupation Rate"
self.modvar_gnrl_subpop = "Population"
self.modvar_gnrl_pop_total = "Total Population"
## MISCELLANEOUS VARIABLES
self.time_periods, self.n_time_periods = self.get_time_periods()
# TEMP:SET TO DERIVE FROM ATTRIBUTE TABLES---
self.cat_lu_crop = "croplands"
self.cat_lu_grazing = "grasslands"
self.varchar_str_emission_gas = "$EMISSION-GAS$"
self.varchar_str_unit_mass = "$UNIT-MASS$"
## FUNCTIONS FOR MODEL ATTRIBUTE DIMENSIONS
def check_df_fields(self, df_afolu_trajectories):
check_fields = self.required_variables
# check for required variables
if not set(check_fields).issubset(df_afolu_trajectories.columns):
set_missing = list(set(check_fields) - set(df_afolu_trajectories.columns))
set_missing = sf.format_print_list(set_missing)
raise KeyError(f"AFOLU projection cannot proceed: The fields {set_missing} are missing.")
def get_required_subsectors(self):
subsectors = list(sf.subset_df(self.model_attributes.dict_attributes["abbreviation_subsector"].table, {"sector": ["AFOLU"]})["subsector"])
subsectors_base = subsectors.copy()
subsectors += ["Economy", "General"]
return subsectors, subsectors_base
def get_required_dimensions(self):
## TEMPORARY - derive from attributes later
required_doa = [self.model_attributes.dim_time_period]
return required_doa
def get_afolu_input_output_fields(self):
required_doa = [self.model_attributes.dim_time_period]
required_vars, output_vars = self.model_attributes.get_input_output_fields(self.required_subsectors)
return required_vars + self.get_required_dimensions(), output_vars
# define a function to clean up code
def get_standard_variables(self, df_in, modvar, override_vector_for_single_mv_q: bool = False, return_type: str = "data_frame"):
flds = self.model_attributes.dict_model_variables_to_variables[modvar]
flds = flds[0] if ((len(flds) == 1) and not override_vector_for_single_mv_q) else flds
valid_rts = ["data_frame", "array_base", "array_units_corrected"]
if return_type not in valid_rts:
vrts = sf.format_print_list(valid_rts)
raise ValueError(f"Invalid return_type in get_standard_variables: valid types are {vrts}.")
# initialize output, apply various common transformations based on type
out = df_in[flds]
if return_type != "data_frame":
out = np.array(out)
if return_type == "array_units_corrected":
out *= self.get_scalar(modvar, "total")
return out
def get_time_periods(self):
pydim_time_period = self.model_attributes.get_dimensional_attribute("time_period", "pydim")
time_periods = self.model_attributes.dict_attributes[pydim_time_period].key_values
return time_periods, len(time_periods)
## STREAMLINING FUNCTIONS
# convert an array to a varibale out dataframe
def array_to_df(self, arr_in, modvar: str, include_scalars = False) -> pd.DataFrame:
# get subsector and fields to name based on variable
subsector = self.model_attributes.dict_model_variable_to_subsector[modvar]
fields = self.model_attributes.build_varlist(subsector, variable_subsec = modvar)
scalar_em = 1
scalar_me = 1
if include_scalars:
# get scalars
gas = sa.model_attributes.get_variable_characteristic(modvar, self.varchar_str_emission_gas)
mass = sa.model_attributes.get_variable_characteristic(modvar, self.varchar_str_unit_mass)
# will conver ch4 to co2e e.g. + kg to MT
scalar_em = 1 if not gas else self.model_attributes.get_gwp(gas.lower())
scalar_me = 1 if not mass else self.model_attributes.get_mass_equivalent(mass.lower())
# raise error if there's a shape mismatch
if len(fields) != arr_in.shape[1]:
flds_print = sf.format_print_list(fields)
raise ValueError(f"Array shape mismatch for fields {flds_print}: the array only has {arr_in.shape[1]} columns.")
return pd.DataFrame(arr_in*scalar_em*scalar_me, columns = fields)
# some scalars
def get_scalar(self, modvar: str, return_type: str = "total"):
valid_rts = ["total", "gas", "mass"]
if return_type not in valid_rts:
tps = sf.format_print_list(valid_rts)
raise ValueError(f"Invalid return type '{return_type}' in get_scalar: valid types are {tps}.")
# get scalars
gas = self.model_attributes.get_variable_characteristic(modvar, self.varchar_str_emission_gas)
scalar_gas = 1 if not gas else self.model_attributes.get_gwp(gas.lower())
#
mass = self.model_attributes.get_variable_characteristic(modvar, self.varchar_str_unit_mass)
scalar_mass = 1 if not mass else self.model_attributes.get_mass_equivalent(mass.lower())
if return_type == "gas":
out = scalar_gas
elif return_type == "mass":
out = scalar_mass
elif return_type == "total":
out = scalar_gas*scalar_mass
return out
# loop over a dictionary of simple variables that map an emission factor () to build out
def get_simple_input_to_output_emission_arrays(self, df_ef: pd.DataFrame, df_driver: pd.DataFrame, dict_vars: dict, variable_driver: str):
"""
NOTE: this only works w/in subsector
"""
df_out = []
subsector_driver = self.model_attributes.dict_model_variable_to_subsector[variable_driver]
for var in dict_vars.keys():
subsector_var = self.model_attributes.dict_model_variable_to_subsector[var]
if subsector_driver != subsector_driver:
warnings.warn(f"In get_simple_input_to_output_emission_arrays, driver variable '{variable_driver}' and emission variable '{var}' are in different sectors. This instance will be skipped.")
else:
# get emissions factor fields and apply scalar using get_standard_variables
arr_ef = np.array(self.get_standard_variables(df_ef, var, True, "array_units_corrected"))
# get the emissions driver array (driver must h)
arr_driver = np.array(df_driver[sa.model_attributes.build_target_varlist_from_source_varcats(var, variable_driver)])
df_out.append(self.array_to_df(arr_driver*arr_ef, dict_vars[var]))
return df_out
# add subsector emission totals
def add_subsector_emissions_aggregates(self, df_in: pd.DataFrame, stop_on_missing_fields_q: bool = False):
# loop over base subsectors
for subsector in model_afolu.required_base_subsectors:
vars_subsec = self.model_attributes.dict_model_variables_by_subsector[subsector]
# add subsector abbreviation
fld_nam = sa.model_attributes.get_subsector_attribute(subsector, "abv_subsector")
fld_nam = f"emission_co2e_subsector_total_{fld_nam}"
flds_add = []
for var in vars_subsec:
var_type = self.model_attributes.get_variable_attribute(var, "variable_type").lower()
gas = self.model_attributes.get_variable_characteristic(var, "$EMISSION-GAS$")
if (var_type == "output") and gas:
flds_add += self.model_attributes.dict_model_variables_to_variables[var]
# check for missing fields; notify
missing_fields = [x for x in flds_add if x not in df_in.columns]
if len(missing_fields) > 0:
str_mf = print_setdiff(set(df_in.columns), set(flds_add))
str_mf = f"Missing fields {str_mf}.%s"
if stop_on_missing_fields_q:
raise ValueError(str_mf%(" Subsector emission totals will not be added."))
else:
warnings.warn(str_mf%(" Subsector emission totals will exclude these fields."))
keep_fields = [x for x in flds_add if x in df_in.columns]
print(keep_fields)
df_in[fld_nam] = df_in[keep_fields].sum(axis = 0)
######################################
# SUBSECTOR SPECIFIC FUNCTIONS #
######################################
### AGRICULTURE
def check_cropland_fractions(self, df_in, thresh_for_correction: float = 0.01):
arr = self.get_standard_variables(df_in, self.modvar_agrc_area_prop, True, "array_base")
totals = sum(arr.transpose())
m = max(np.abs(totals - 1))
if m > thresh_for_correction:
raise ValueError(f"Invalid crop areas found in check_cropland_fractions. The maximum fraction total was {m}; the maximum allowed deviation from 1 is {thresh_for_correction}.")
else:
arr = (arr.transpose()/totals).transpose()
return arr
### LAND USE
## check the shape of transition/emission factor matrices sent to project_land_use
def check_markov_shapes(self, arrs: np.ndarray, function_var_name:str):
# get land use info
pycat_lndu = self.model_attributes.get_subsector_attribute("Land Use", "pycategory_primary")
attr_lndu = self.model_attributes.dict_attributes[pycat_lndu]
if len(arrs.shape) < 3:
raise ValueError(f"Invalid shape for array {function_var_name}; the array must be a list of square matrices.")
elif arrs.shape[1:3] != (attr_lndu.n_key_values, attr_lndu.n_key_values):
raise ValueError(f"Invalid shape of matrices in {function_var_name}. They must have shape ({attr_lndu.n_key_values}, {attr_lndu.n_key_values}).")
## get the transition and emission factors matrices from the data frame
def get_markov_matrices(self, df_ordered_trajectories, thresh_correct = 0.0001):
"""
- assumes that the input data frame is ordered by time_period
- thresh_correct is used to decide whether or not to correct the transition matrix (assumed to be row stochastic) to sum to 1; if the abs of the sum is outside this range, an error will be thrown
- fields_pij and fields_efc will be properly ordered by categories for this transformation
"""
fields_pij = self.model_attributes.dict_model_variables_to_variables[self.modvar_lndu_prob_transition]
fields_efc = self.model_attributes.dict_model_variables_to_variables[self.modvar_lndu_ef_co2_conv]
sf.check_fields(df_ordered_trajectories, fields_pij + fields_efc)
pycat_landuse = self.model_attributes.get_subsector_attribute("Land Use", "pycategory_primary")
n_categories = len(self.model_attributes.dict_attributes[pycat_landuse].key_values)
# fetch arrays of transition probabilities and co2 emission factors
arr_pr = np.array(df_ordered_trajectories[fields_pij])
arr_pr = arr_pr.reshape((self.n_time_periods, n_categories, n_categories))
arr_ef = np.array(df_ordered_trajectories[fields_efc])
arr_ef = arr_ef.reshape((self.n_time_periods, n_categories, n_categories))
return arr_pr, arr_ef
## project land use
def project_land_use(self, vec_initial_area: np.ndarray, arrs_transitions: np.ndarray, arrs_efs: np.ndarray):
t0 = time.time()
# check shapes
self.check_markov_shapes(arrs_transitions, "arrs_transitions")
self.check_markov_shapes(arrs_efs, "arrs_efs")
# get land use info
pycat_lndu = self.model_attributes.get_subsector_attribute("Land Use", "pycategory_primary")
attr_lndu = self.model_attributes.dict_attributes[pycat_lndu]
# intilize the land use and conversion emissions array
shp_init = (self.n_time_periods, attr_lndu.n_key_values)
arr_land_use = np.zeros(shp_init)
arr_emissions_conv = np.zeros(shp_init)
arrs_land_conv = np.zeros((self.n_time_periods, attr_lndu.n_key_values, attr_lndu.n_key_values))
# running matrix Q_i; initialize as identity. initialize running matrix of land use are
Q_i = np.identity(attr_lndu.n_key_values)
x = vec_initial_area
i = 0
while i < self.n_time_periods:
# check emission factor index
i_ef = i if (i < len(arrs_efs)) else len(arrs_efs) - 1
if i_ef != i:
print(f"No emission factor matrix found for time period {self.time_periods[i]}; using the matrix from period {len(arrs_efs) - 1}.")
# check transition matrix index
i_tr = i if (i < len(arrs_transitions)) else len(arrs_transitions) - 1
if i_tr != i:
print(f"No transition matrix found for time period {self.time_periods[i]}; using the matrix from period {len(arrs_efs) - 1}.")
# calculate land use, conversions, and emissions
vec_land_use = np.matmul(vec_initial_area, Q_i)
vec_emissions_conv = sum((arrs_transitions[i_tr] * arrs_efs[i_ef]).transpose()*x.transpose())
arr_land_conv = (arrs_transitions[i_tr].transpose()*x.transpose()).transpose()
# update matrices
rng_put = np.arange(i*attr_lndu.n_key_values, (i + 1)*attr_lndu.n_key_values)
np.put(arr_land_use, rng_put, vec_land_use)
np.put(arr_emissions_conv, rng_put, vec_emissions_conv)
np.put(arrs_land_conv, np.arange(i*attr_lndu.n_key_values**2, (i + 1)*attr_lndu.n_key_values**2), arr_land_conv)
# update transition matrix and land use matrix
Q_i = np.matmul(Q_i, arrs_transitions[i_tr])
x = vec_land_use
i += 1
t1 = time.time()
t_elapse = round(t1 - t0, 2)
print(f"Land use projection complete in {t_elapse} seconds.")
return arr_emissions_conv, arr_land_use, arrs_land_conv
## LIVESTOCK
def reassign_pops_from_proj_to_carry(self, arr_lu_derived, arr_dem_based):
"""
Before assigning net imports, there are many non-grazing animals to consider (note that these animals are generally not emission-intensive animals)
Due to 0 graze area, their estimated population is infinite, or stored as a negative
We assign their population as the demand-estimated population
"""
if arr_lu_derived.shape != arr_dem_based.shape:
raise ValueError(f"Error in reassign_pops_from_proj_to_carry: array dimensions do not match: arr_lu_derived = {arr_lu_derived.shape}, arr_dem_based = {arr_dem_based.shape}.")
cols = np.where(arr_lu_derived[0] < 0)[0]
n_row, n_col = arr_lu_derived.shape
for w in cols:
rng = np.arange(w*n_row, (w + 1)*n_row)
np.put(arr_lu_derived.transpose(), rng, arr_dem_based[:, w])
return arr_lu_derived
####################################
### ###
### PRIMARY MODEL FUNCTION ###
### ###
####################################
def project(self, df_afolu_trajectories):
"""
- AFOLU.project takes a data frame (ordered by time series) and returns a data frame of the same order
- designed to be parallelized or called from command line via __main__ in run_afolu.py
"""
## CHECKS
# check for internal variables and add if necessary; note, this can be defined for different variables (see model attributes)
self.model_attributes.manage_pop_to_df(df_afolu_trajectories, "add")
df_afolu_trajectories.sort_values(by = [self.model_attributes.dim_time_period], inplace = True)
# check that all required fields are contained—assume that it is ordered by time period
self.check_df_fields(df_afolu_trajectories)
## CATEGORY INITIALIZATION
pycat_agrc = self.model_attributes.get_subsector_attribute("Agriculture", "pycategory_primary")
pycat_frst = self.model_attributes.get_subsector_attribute("Forest", "pycategory_primary")
pycat_lndu = self.model_attributes.get_subsector_attribute("Land Use", "pycategory_primary")
pycat_lvst = self.model_attributes.get_subsector_attribute("Livestock", "pycategory_primary")
# attribute tables
attr_agrc = self.model_attributes.dict_attributes[pycat_agrc]
attr_frst = self.model_attributes.dict_attributes[pycat_frst]
attr_lndu = self.model_attributes.dict_attributes[pycat_lndu]
attr_lvst = self.model_attributes.dict_attributes[pycat_lvst]
## FIELD INITIALIZATION
# get the gdp and total population fields
field_gdp = self.model_attributes.build_varlist("Economy", variable_subsec = self.modvar_econ_gdp)[0]
field_pop = self.model_attributes.build_varlist("General", variable_subsec = self.modvar_gnrl_pop_total)[0]
## ECON/GNRL VECTOR AND ARRAY INITIALIZATION
# get some vectors
vec_gdp = self.get_standard_variables(df_afolu_trajectories, self.modvar_econ_gdp, False, return_type = "array_base")#np.array(df_afolu_trajectories[field_gdp])
vec_pop = self.get_standard_variables(df_afolu_trajectories, self.modvar_gnrl_pop_total, False, return_type = "array_base")
vec_gdp_per_capita = vec_gdp/vec_pop
# growth rates
vec_rates_gdp = vec_gdp[1:]/vec_gdp[0:-1] - 1
vec_rates_gdp_per_capita = vec_gdp_per_capita[1:]/vec_gdp_per_capita[0:-1] - 1
## OUTPUT INITIALIZATION
df_out = [df_afolu_trajectories[self.required_dimensions].copy()]
##################
# LAND USE #
##################
# area of the country
area = float(self.get_standard_variables(df_afolu_trajectories, self.modvar_gnrl_area, return_type = "array_base")[0])
## LU MARKOV
# get the initial distribution of land
vec_modvar_lndu_initial_frac = self.get_standard_variables(df_afolu_trajectories, self.modvar_lndu_initial_frac, return_type = "array_base")[0]
vec_modvar_lndu_initial_area = vec_modvar_lndu_initial_frac*area
self.vec_modvar_lndu_initial_area = vec_modvar_lndu_initial_area
self.mat_trans, self.mat_ef = self.get_markov_matrices(df_afolu_trajectories)
# get land use projections (np arrays) - note, arrs_land_conv returns a list of matrices for troubleshooting
arr_lndu_emissions_conv, arr_land_use, arrs_land_conv = self.project_land_use(vec_modvar_lndu_initial_area, *self.get_markov_matrices(df_afolu_trajectories))
# scale emissions
arr_lndu_emissions_conv *= self.get_scalar(self.modvar_lndu_ef_co2_conv, "total")
df_lndu_emissions_conv = self.array_to_df(arr_lndu_emissions_conv, self.modvar_lndu_emissions_conv)
df_land_use = self.array_to_df(arr_land_use, self.modvar_lndu_area_by_cat)
# add to output data frame
df_out.append(df_lndu_emissions_conv)
df_out.append(df_land_use)
## EXISTENCE EMISSIONS FOR OTHER LANDS, INCLUDING AG ACTIVITY ON PASTURES
# dictionary variables mapping emission factor variables to output variables
dict_modvars_lndu_simple_efs = {
self.modvar_lndu_ef_n2o_past: self.modvar_lndu_emissions_n2o_from_pastures,
self.modvar_lndu_ef_co2_soilcarb: self.modvar_lndu_emissions_co2_from_pastures,
self.modvar_lndu_ef_ch4_boc: self.modvar_lndu_emissions_ch4_from_wetlands
}
# add to output dataframe
df_out += self.get_simple_input_to_output_emission_arrays(df_afolu_trajectories, df_land_use, dict_modvars_lndu_simple_efs, self.modvar_lndu_area_by_cat)
##################
# FORESTRY #
##################
# get ordered fields from land use
fields_lndu_forest_ordered = [self.model_attributes.matchstring_landuse_to_forests + x for x in self.model_attributes.dict_attributes[pycat_frst].key_values]
arr_area_frst = np.array(df_land_use[self.model_attributes.build_varlist("Land Use", variable_subsec = self.modvar_lndu_area_by_cat, restrict_to_category_values = fields_lndu_forest_ordered)])
# get different variables
arr_frst_ef_sequestration = self.get_standard_variables(df_afolu_trajectories, self.modvar_frst_sq_co2, True, "array_units_corrected")
arr_frst_ef_methane = self.get_standard_variables(df_afolu_trajectories, self.modvar_frst_ef_ch4, True, "array_units_corrected")
# build output variables
df_out += [
self.array_to_df(-1*arr_area_frst*arr_frst_ef_sequestration, self.modvar_frst_emissions_sequestration),
self.array_to_df(arr_area_frst*arr_frst_ef_methane, self.modvar_frst_emissions_methane)
]
## NEEDED: FOREST FIRES (ADD HERE)
## NEEDED: WOOD PRODUCTS (ADD HERE)
#####################
# AGRICULTURE #
#####################
# get area of cropland
field_crop_array = self.model_attributes.build_varlist("Land Use", variable_subsec = self.modvar_lndu_area_by_cat, restrict_to_category_values = [self.cat_lu_crop])[0]
vec_cropland_area = np.array(df_land_use[field_crop_array])
# fraction of cropland represented by each crop
arr_agrc_frac_cropland_area = self.check_cropland_fractions(df_afolu_trajectories)
arr_agrc_crop_area = (arr_agrc_frac_cropland_area.transpose()*vec_cropland_area.transpose()).transpose()
# area-corrected emission factors
arr_agrc_ef_ch4 = self.get_standard_variables(df_afolu_trajectories, self.modvar_agrc_ef_ch4, True, "array_units_corrected")
arr_agrc_ef_co2 = self.get_standard_variables(df_afolu_trajectories, self.modvar_agrc_ef_co2, True, "array_units_corrected")
arr_agrc_ef_n2o = self.get_standard_variables(df_afolu_trajectories, self.modvar_agrc_ef_n2o, True, "array_units_corrected")
# estimate yield capacity
arr_agrc_yf = self.get_standard_variables(df_afolu_trajectories, self.modvar_agrc_yf, True, "array_base")
arr_yield = arr_agrc_yf*arr_agrc_crop_area
# estimate demand for crops (used in CBA)
arr_agrc_elas_crop_demand = self.get_standard_variables(df_afolu_trajectories, self.modvar_agrc_elas_crop_demand_income, False, "array_base")
arr_agrc_yield_dem_scale_proj = (vec_rates_gdp_per_capita.transpose()*arr_agrc_elas_crop_demand[0:-1].transpose()).transpose()
arr_agrc_yield_dem_scale_proj = np.cumprod(1 + arr_agrc_yield_dem_scale_proj, axis = 0)
arr_agrc_yield_dem_scale_proj = np.concatenate([np.ones((1,len(arr_agrc_yield_dem_scale_proj[0]))), arr_agrc_yield_dem_scale_proj])
# estimate net imports (surplus demand)
arr_agrc_net_imports = arr_agrc_yield_dem_scale_proj*arr_yield[0] - arr_yield
# add to output dataframe
df_out += [
self.array_to_df(arr_agrc_crop_area, self.modvar_agrc_area_crop),
self.array_to_df(arr_yield, self.modvar_agrc_yield),
self.array_to_df(arr_agrc_ef_ch4, self.modvar_agrc_emissions_ch4_crops),
self.array_to_df(arr_agrc_ef_co2, self.modvar_agrc_emissions_co2_crops),
self.array_to_df(arr_agrc_ef_n2o, self.modvar_agrc_emissions_n2o_crops),
self.array_to_df(arr_agrc_net_imports, self.modvar_agrc_net_imports)
]
###################
# LIVESTOCK #
###################
# get area of grassland/pastures
field_lvst_graze_array = self.model_attributes.build_varlist("Land Use", variable_subsec = self.modvar_lndu_area_by_cat, restrict_to_category_values = [self.cat_lu_grazing])[0]
vec_lvst_graze_area = np.array(df_land_use[field_lvst_graze_array])
# get weights for allocating grazing area to animals - based on first year only
vec_lvst_base_graze_weights = self.get_standard_variables(df_afolu_trajectories, self.modvar_lvst_dry_matter_consumption, True, "array_base")[0]
vec_modvar_lvst_pop_init = self.get_standard_variables(df_afolu_trajectories, self.modvar_lvst_pop_init, True, "array_base")[0]
vec_lvst_grassland_allocation_weights = (vec_modvar_lvst_pop_init*vec_lvst_base_graze_weights)/np.dot(vec_modvar_lvst_pop_init, vec_lvst_base_graze_weights)
# estimate the total area used for grazing, then get the number of livestock/area
arr_lvst_graze_area = np.outer(vec_lvst_graze_area, vec_lvst_grassland_allocation_weights)
vec_lvst_carry_capacity_scale = self.get_standard_variables(df_afolu_trajectories, self.modvar_lvst_carrying_capacity_scalar, False, "array_base")
vec_lvst_carry_capacity = vec_modvar_lvst_pop_init/arr_lvst_graze_area[0]
arr_lvst_carry_capacity = np.outer(vec_lvst_carry_capacity_scale, vec_lvst_carry_capacity)
# estimate the total number of livestock that are raised, then get emission factors
arr_lvst_pop = np.array(arr_lvst_carry_capacity*arr_lvst_graze_area).astype(int)
arr_lvst_emissions_ch4_ef = self.get_standard_variables(df_afolu_trajectories, self.modvar_lvst_ef_ch4_ef, True, "array_units_corrected")
arr_lvst_emissions_ch4_mm = self.get_standard_variables(df_afolu_trajectories, self.modvar_lvst_ef_ch4_mm, True, "array_units_corrected")
arr_lvst_emissions_n2o_mm = self.get_standard_variables(df_afolu_trajectories, self.modvar_lvst_ef_n2o_mm, True, "array_units_corrected")
# estimate demand for livestock (used in CBA)
fields_lvst_elas = self.model_attributes.switch_variable_category("Livestock", self.modvar_lvst_elas_lvst_demand, "demand_elasticity_category")
arr_lvst_elas_demand = np.array(df_afolu_trajectories[fields_lvst_elas])
# get the demand scalar, then apply to the initial population
arr_lvst_dem_scale_proj = (vec_rates_gdp_per_capita.transpose()*arr_lvst_elas_demand[0:-1].transpose()).transpose()
arr_lvst_dem_scale_proj = np.cumprod(1 + arr_lvst_dem_scale_proj, axis = 0)
arr_lvst_dem_scale_proj= np.concatenate([np.ones((1,len(arr_lvst_dem_scale_proj[0]))), arr_lvst_dem_scale_proj])
arr_lvst_dem_pop = np.array(arr_lvst_dem_scale_proj*vec_modvar_lvst_pop_init).astype(int)
# clean the population and grab net imports
arr_lvst_pop = self.reassign_pops_from_proj_to_carry(arr_lvst_pop, arr_lvst_dem_pop)
arr_lvst_net_imports = arr_lvst_dem_pop - arr_lvst_pop
# add to output dataframe
df_out += [
self.array_to_df(arr_lvst_emissions_ch4_ef*arr_lvst_pop, self.modvar_lvst_emissions_ch4_ef),
self.array_to_df(arr_lvst_emissions_ch4_mm*arr_lvst_pop, self.modvar_lvst_emissions_ch4_mm),
self.array_to_df(arr_lvst_emissions_n2o_mm*arr_lvst_pop, self.modvar_lvst_emissions_n2o_mm),
self.array_to_df(arr_lvst_pop, self.modvar_lvst_pop),
self.array_to_df(arr_lvst_net_imports, self.modvar_lvst_net_imports)
]
df_out = pd.concat(df_out, axis = 1).reset_index(drop = True)
self.add_subsector_emissions_aggregates(df_out, False)
return df_out
model_afolu = AFOLU(sa.model_attributes)
#df_emissions_conv, arr_emissions_conv, arr_land_use, arrs_land_conv = model_afolu.project(df_fake_data)
df_afolu = model_afolu.project(df_fake_data)
# -
# +
if True and "tmp":
print("tmp")
if "this":
print("n")
# -
#importlib.reload(sa)
np.where(np.array(df_afolu[["net_imports_lvst_mules", "net_imports_lvst_goats"]])[1] <= 850)[0]
#model_afolu.get_standard_variables(df_afolu_trajectories, self.modvar_frst_sq_co2, True, "array_units_corrected")
df_afolu
importlib.reload(sa)
x = np.array(df_afolu["area_lu_forests_primary_ha"])
x[1:]/x[0:-1]
x
v0 = model_afolu.vec_modvar_lndu_initial_area
print(v0)
v1 = np.matmul(v0.transpose(), model_afolu.mat_trans[0])
v1
v2 = np.matmul(v1.transpose(), model_afolu.mat_trans[1])
v2
# +
df_afolu
modvar = model_afolu.modvar_lndu_ef_n2o_past
mass = sa.model_attributes.get_variable_characteristic(modvar, "$UNIT-MASS$")
scalar_mass = 1 if not mass else sa.model_attributes.get_mass_equivalent(mass.lower())
gas = sa.model_attributes.get_variable_characteristic(modvar, "$EMISSION-GAS$")
scalar_gas = 1 if not gas else sa.model_attributes.get_gwp(gas.lower())
scalar_mass*scalar_gas
model_afolu.get_scalar(modvar, "total")
# +
importlib.reload(sa)
sa.model_attributes.dict_model_variable_to_category_restriction[model_afolu.modvar_lndu_ef_n2o_past]
sa.model_attributes.separate_varreq_dict_for_outer("Land Use", "key_varreqs_all", ("$CAT-LANDUSE-I$", "$CAT-LANDUSE-J$"))#, variable_type = var_type)
sa.model_attributes.dict_model_variables_by_subsector["Land Use"]#[model_afolu.modvar_lndu_ef_n2o_past]
#get_variable_attribute(model_afolu.modvar_lndu_ef_n2o_past, "categories")
#sa.model_attributes.dict_varreqs["partial_category_af_lndu"].field_maps
# +
importlib.reload(sa)
#sa.model_attributes.dict_model_variables_by_subsector
#sa.model_attributes.get_variable_attribute(":math:\\text{CO}_2 Emissions from Land Use Conversion", "variable_type")
sa.model_attributes.get_categories(":math:\\text{CO}_2 Emissions from Land Use Conversion")
sa.model_attributes.get_categories(model_afolu.modvar_lndu_ef_n2o_past)
# function to build a variable using an ordered set of categories associated with another variable
def build_target_varlist_from_source_varcats(self, modvar_source: str, modvar_target: str):
# get source categories
cats_source = self.get_categories(modvar_source)
# build the target variable list using the source categories
subsector_target = self.dict_model_variable_to_subsector[modvar_target]
vars_target = self.build_varlist(subsector_target, variable_subsec = modvar_target, restrict_to_category_values = cats_source)
return vars_target
# +
sa.model_attributes.build_target_varlist_from_source_varcats(model_afolu.modvar_lndu_ef_n2o_past, model_afolu.modvar_lndu_area_by_cat)
#model_afolu.get_scalar(model_afolu.modvar_lndu_ef_co2_conv, "mass")
#model_afolu.model_attributes.get_variable_characteristic(model_afolu.modvar_lndu_ef_co2_conv, "$UNIT-MASS$")
# -
x = np.array([50, 10, 81.4, 14])
x2 = np.array([60, 0.9, 81.4, 14])
#x2 = np.array([x for i in range(50)])
#np.reshape(np.repeat(x, 10), (4, 10)).transpose()
x0 = np.zeros((10, 4))
#np.put(x0, x, [2])
np.put(x0, np.arange(4, 8), x)
np.put(x0, np.arange(36, 40), x2)
x0
# +
l_mats = model_afolu.get_markov_matrices(df_fake_data)[0]
t0 = time.time()
mat_it = np.identity(len(l_mats[0]))
for i in range(len(l_mats)):
mat_it = np.matmul(mat_it, l_mats[i])
t1 = time.time()
print(t1 - t0)
# -
#importlib.reload(sa)
#sa.model_attributes.dict_varreqs["cat_landuse"]#.field_maps#["variable_to_variable_schema"]#dict_attributes["emission_gas"].field_maps#_requirements.field_maps#["analytical_parameter_to_default_value"]#dict_model_variables_to_variables
#sa.model_attributes.dict_varreqs[sa.model_attributes.get_subsector_attribute("Livestock", "key_varreqs_partial")].field_maps
#df_fake_data[[x for x in df_fake_data if "pij_forests_primary" in x]]
sa.model_attributes.dict_attributes["emission_gas"].field_maps
#df_fake_data[sa.model_attributes.dict_model_variables_to_variables["Unadjusted Land Use Transition Probability"]]
# + jupyter={"outputs_hidden": true}
#importlib.reload(sa)
cats = sa.model_attributes.dict_attributes["cat_livestock"].key_values
repls = [sa.model_attributes.dict_attributes["cat_livestock"].field_maps["cat_livestock_to_demand_elasticity_category"][x].replace("`", "") for x in cats]
sa.model_attributes.build_varlist("Livestock", model_afolu.modvar_lvst_elas_lvst_demand, repls, {model_afolu.modvar_lvst_elas_lvst_demand: repls})
sa.model_attributes.dict_variable_to_subsector
# +
#sa.model_attributes.dict_model_variables_to_variables[model_afolu.modvar_lvst_elas_lvst_demand]
importlib.reload(sa)
sa.model_attributes.build_varlist("Livestock", model_afolu.modvar_lvst_elas_lvst_demand, repls)
sa.model_attributes.check_category_restrictions(repls, sa.model_attributes.dict_attributes["cat_livestock"])
subsector = "Livestock"
category_ij_tuple = ("$CAT-LIVESTOCK-I$", "$CAT-LIVESTOCK-J$")
variable_subsec = model_afolu.modvar_lvst_elas_lvst_demand
# -
#importlib.reload(sa)
sa.model_attributes.switch_variable_category("Livestock", model_afolu.modvar_lvst_elas_lvst_demand, "demand_elasticity_category")
sa.model_attributes.switch_variable_category("IPPU", "Value Added", "value_added_category")#, cats_to_switch = ["minerals", "electronics"])
with open("/Users/jsyme/Desktop/tmp.txt", "r") as fltmp:
vl = fltmp.readlines()
vl = [x.split(" =")[0].replace("\t", "").replace(" ", "") for x in vl]
for k in vl:
print(k)
# +
model_afolu.get_required_variables(True).to_csv(os.path.join(sa.dir_out, "tmp_vars_for_afolu2.csv"), encoding = "UTF-8", index = None)
# +
model_afolu = AFOLU(sa.model_attributes)
#model_afolu.get_required_variables(True)
df_cur = pd.read_csv(os.path.join(sa.dir_out, "tmp_afolu_vars.csv"))
df_reqd = model_afolu.get_required_variables(True)
df_reqd[~df_reqd["variable"].isin(df_cur["variable"])].reset_index(drop = True).to_csv(os.path.join(sa.dir_out, "tmp_vars_for_afolu_comps_new.csv"), encoding = "UTF-8", index = None)
# -
df_reqd
# +
importlib.reload(sa)
model_afolu = AFOLU(sa.model_attributes)
model_afolu.get_required_variables(True).to_csv(os.path.join(sa.dir_out, "tmp_vars_for_afolu2.csv"), encoding = "UTF-8", index = None)
if False:
df_fake_data = pd.read_csv(os.path.join(sa.dir_out, "fake_data_afolu.csv"))
df_fake_data[sa.model_attributes.build_varlist("Economy", "Gross Domestic Product")[0]] = df_fake_data[sa.model_attributes.build_varlist("Economy", "Value Added")].sum(axis = 1)
# note that categories are sorted properly when using build_varlist
[sa.model_attributes.build_varlist("Livestock", x) for x in sa.model_attributes.dict_variables_by_subsector["Livestock"]]
# -
df_fake_data = pd.read_csv(os.path.join(sa.dir_out, "fake_data_afolu.csv"))
# add some vars
df_fake_data[sa.model_attributes.build_varlist("Economy", "Gross Domestic Product")[0]] = df_fake_data[sa.model_attributes.build_varlist("Economy", "Value Added")].sum(axis = 1)
df_fake_data["population_total"] = df_fake_data[sa.model_attributes.build_varlist("General", "Population")].sum(axis = 1).astype(int)
model_afolu.project(df_fake_data)
sa.model_attributes.build_varlist("Land Use", ":math:\\text{CO}_2 Land Use Conversion Emission Factor")
#sa.model_attributes.dict_varreqs["category_af_lndu"].table["variable"].iloc[2]
[x for x in df_fake_data.columns if "forest" in x]
# +
sa.model_attributes.
# -
#
#
#
|
python/build_afolu.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="OqSmFS3lZj9Y"
# # <center> Video Image Data </center>
# #### CMSE 495 Ford Group
#
# This tutorial teaches the user how to input a video file, such a mp4 and convert each frame of the video into a jpeg image using python, primarily in a Jupyter notebook.
# + [markdown] id="GQIJxk_kdjT1"
# <b> Environment Setup (Makefile):</b>
# - Use the command 'make innit' automatically set up the environment for you.
#
# <b> Environment Setup (Manual):</b>
# - Set up new environment using pip/conda (Conda Recommended). Use command
#
# <code> conda create -n envs python=3.10 </code>
#
# - Activate your new environment. Use command
#
# <code> conda activate envs</code>
#
# - Install the requisite packages.Use command
#
# <code> pip install opencv-python</code> or,
#
# <code> conda install -c conda-forge opencv</code>
#
# <b> Usage Instructions:</b>
#
# - The example call shows the format in which this func may be used.
#
# + [markdown] id="f0aUW4PLdobE"
# This process uses 2 packages called [os](https://docs.python.org/3/library/os.html) and [cv2](https://pypi.org/project/opencv-python/). Os provides miscellaneous operating system interfaces such as opening and reading the files.
# + [markdown] id="qClPlbKaduK5"
# <b> ADD </b> An example question and visualization that that the data or tool can answer (this should be unique)
# + id="PCbpVR-HZzmt"
import cv2
import os
import glob
def video_to_frames(file_path, directory_path, greyscale = False):
'''This function will change a video file to a frames'''
#opening the video
vidcap = cv2.VideoCapture(file_path)
dirname = directory_path
os.makedirs(dirname, exist_ok=True)
#capturing a frame as well as a boolean value representing whether an image was properly opened
success,image = vidcap.read()
count = 0
while success:
#this is specifically for foam_segmented.avi
if greyscale:
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
#writing the image to a the directory path that was specified,
#if the path specified does not exist then it will be created
#this finctionality was added so that the images could be stored in a separate folder
#example of output file names: 1.jpg, 2.jpg, 3.jpg, and so on
cv2.imwrite(os.path.join(dirname, str(count)+".jpg"), image)
success,image = vidcap.read()
count += 1
#All the frames will be added in order
cv2.waitKey(1)
#releasing the threads
vidcap.release()
# + [markdown] id="x6DuN40gcSGC"
# Example call for the function above
#
# avi_frames('./ford_data/2D_xy.avi', "./frames/2D_xy",False)
# + id="-N-hJD11jJjo"
# Making a Video From Frames
def frames_to_video(directory_path, fps, width, height):
fourcc = cv2.VideoWriter_fourcc(*'mp4v')
video = cv2.VideoWriter('video.avi', fourcc, fps, (width, height))
num_frames = len([name for name in os.listdir(directory_path) if os.path.isfile(name)])
for j in range(num_frames):
img = cv2.imread(str(j) + '.jpg')
video.write(img)
cv2.destroyAllWindows()
video.release()
# + [markdown] id="bnfzxPNJeZVS"
# **The code below will put the image arrays into a list.** This snippet of code utilizes glob but packages like os can also be used.
# + id="mLbqPW7SeJd-"
path = glob.glob("./*.jpg")
images = []
for img in path:
n = cv2.imread(img)
images.append(n)
# + [markdown] id="GIq-_h4wdxWM"
# <b> References:</b>
# - https://docs.conda.io/projects/conda/en/latest/user-guide/tasks/manage-environments.html
# - https://pypi.org/project/opencv-python/
# + id="hXCCqo-kkptN"
|
Ford_Video_Analysis.ipynb
|
# +
from itertools import islice
import numpy as np
import holoviews as hv
from holoviews import opts
from holoviews.plotting.links import RangeToolLink
hv.extension('bokeh')
# -
# This demo demonstrates how to link two plots using the RangeToolLink along both axes. This can be useful to get an overview and a detailed view of some data at the same time.
# ## Declare data
# +
def mandelbrot_generator(h,w, maxit, bounds):
"Generator that yields the mandlebrot set."
(l,b,r,t) = bounds
y,x = np.ogrid[b:t : h*1j, l:r:w*1j]
c = x+y*1j
z = c
divtime = maxit + np.zeros(z.shape, dtype=int)
for i in range(maxit):
z = z**2 + c
diverge = z*np.conj(z) > 2**2
div_now = diverge & (divtime==maxit)
divtime[div_now] = i
z[diverge] = 2
yield divtime
def mandelbrot(h,w, n, maxit, bounds):
"Returns the mandelbrot set computed to maxit"
iterable = mandelbrot_generator(h,w, maxit, bounds)
return next(islice(iterable, n, None))
bounds = (-2,-1.4,0.8,1.4)
mbset = mandelbrot(800, 800, 45, 46, bounds)
mbset_image = hv.Image(mbset, bounds=bounds)
# -
# ## Declare plot
# Having declared an ``Image`` of the Mandelbrot set we make a smaller and larger version of it. The smaller ``source`` will serve as an overview containing the ``RangeTool`` which allows selecting the region to show in the larger ``target`` plot. We can control which axes should be linked to the ``RangeTool`` with the axes parameter on the ``RangeToolLink``:
# +
src = mbset_image.clone(link=False).opts(width=300, height=300, padding=0.1)
tgt = mbset_image.opts(width=500, height=500, xaxis=None, yaxis=None, clone=True)
# Declare a RangeToolLink between the x- and y-axes of the two plots
RangeToolLink(src, tgt, axes=['x', 'y'])
(tgt + src).opts(
opts.Image(default_tools=[]),
opts.Layout(shared_axes=False))
|
examples/gallery/demos/bokeh/image_range_tool.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #Fine-tuning a Pretrained Network for Color Constancy
#
# In this example, we'll explore a common approach that is particularly useful in real-world applications: take a pre-trained Caffe network and fine-tune the parameters on your custom data.
#
# The upside of such approach is that, since pre-trained networks are learned on a large set of images, the intermediate layers capture the "semantics" of the general visual appearance. Think of it as a very powerful feature that you can treat as a black box. On top of that, only a few layers will be needed to obtain a very good performance of the data.
# First, we will need to prepare the data. This involves the following parts:
# (1) Get the ImageNet ilsvrc pretrained model with the provided shell scripts.
# (2) Download a subset of the overall Flickr style dataset for this demo.
# (3) Compile the downloaded Flickr dataset into a database that Caffe can then consume.
# +
import os
os.chdir('..')
import sys
sys.path.insert(0, './python')
import caffe
import numpy as np
from pylab import *
# %matplotlib inline
# -
# Let's show what is the difference between the fine-tuning network and the original caffe model.
# For your record, if you want to train the network in pure C++ tools, here is the command:
#
# <code>
# build/tools/caffe train \
# -solver models/finetune_flickr_style/solver.prototxt \
# -weights models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel \
# -gpu 0
# </code>
#
# However, we will train using Python in this example.
# +
niter = 200
# losses will also be stored in the log
train_loss = np.zeros(niter)
scratch_train_loss = np.zeros(niter)
caffe.set_device(0)
caffe.set_mode_gpu()
# We create a solver that fine-tunes from a previously trained network.
solver = caffe.SGDSolver('models/finetune_flickr_style_2/solver.prototxt')
solver.net.copy_from('models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel')
# For reference, we also create a solver that does no finetuning.
scratch_solver = caffe.SGDSolver('models/finetune_flickr_style_2/solver.prototxt')
# We run the solver for niter times, and record the training loss.
for it in range(niter):
solver.step(1) # SGD by Caffe
scratch_solver.step(1)
# store the train loss
train_loss[it] = solver.net.blobs['loss'].data
scratch_train_loss[it] = scratch_solver.net.blobs['loss'].data
if it % 10 == 0:
print 'iter %d, finetune_loss=%f, scratch_loss=%f' % (it, train_loss[it], scratch_train_loss[it])
print 'done'
# -
# Let's look at the training loss produced by the two training procedures respectively.
plot(np.vstack([train_loss, scratch_train_loss]).T)
# Notice how the fine-tuning procedure produces a more smooth loss function change, and ends up at a better loss. A closer look at small values, clipping to avoid showing too large loss during training:
plot(np.vstack([train_loss, scratch_train_loss]).clip(0, 4).T)
# Let's take a look at the testing accuracy after running 200 iterations. Note that we are running a classification task of 5 classes, thus a chance accuracy is 20%. As we will reasonably expect, the finetuning result will be much better than the one from training from scratch. Let's see.
test_iters = 10
accuracy = 0
scratch_accuracy = 0
for it in arange(test_iters):
solver.test_nets[0].forward()
accuracy += solver.test_nets[0].blobs['accuracy'].data
scratch_solver.test_nets[0].forward()
scratch_accuracy += scratch_solver.test_nets[0].blobs['accuracy'].data
accuracy /= test_iters
scratch_accuracy /= test_iters
print 'Accuracy for fine-tuning:', accuracy
print 'Accuracy for training from scratch:', scratch_accuracy
print np.shape(solver.test_nets[0])
# Huzzah! So we did finetuning and it is awesome. Let's take a look at what kind of results we are able to get with a longer, more complete run of the style recognition dataset. Note: the below URL might be occassionally down because it is run on a research machine.
#
# http://demo.vislab.berkeleyvision.org/
|
examples/testColorConstancy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data structures
#
# ## Introduction
#
# This is a getting started tutorial for Gammapy.
#
# In this tutorial we will use the [Second Fermi-LAT Catalog of High-Energy Sources (3FHL) catalog](http://fermi.gsfc.nasa.gov/ssc/data/access/lat/3FHL/), corresponding event list and images to learn how to work with some of the central Gammapy data structures.
#
# We will cover the following topics:
#
# * **Sky maps**
# * We will learn how to handle image based data with gammapy using a Fermi-LAT 3FHL example image. We will work with the following classes:
# - `~gammapy.maps.WcsNDMap`
# - [astropy.coordinates.SkyCoord](http://astropy.readthedocs.io/en/latest/coordinates/index.html)
# - [numpy.ndarray](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ndarray.html)
#
# * **Event lists**
# * We will learn how to handle event lists with Gammapy. Important for this are the following classes:
# - `~gammapy.data.EventList`
# - [astropy.table.Table](http://docs.astropy.org/en/stable/api/astropy.table.Table.html)
#
# * **Source catalogs**
# * We will show how to load source catalogs with Gammapy and explore the data using the following classes:
# - `~gammapy.catalog.SourceCatalog`, specifically `~gammapy.catalog.SourceCatalog3FHL`
# - [astropy.table.Table](http://docs.astropy.org/en/stable/api/astropy.table.Table.html)
#
# * **Spectral models and flux points**
# * We will pick an example source and show how to plot its spectral model and flux points. For this we will use the following classes:
# - `~gammapy.modeling.models.SpectralModel`, specifically the `~gammapy.modeling.models.PowerLaw2SpectralModel`
# - `~gammapy.estimators.FluxPoints`
# - [astropy.table.Table](http://docs.astropy.org/en/stable/api/astropy.table.Table.html)
#
# ## Setup
#
# **Important**: to run this tutorial the environment variable `GAMMAPY_DATA` must be defined and point to the directory on your machine where the datasets needed are placed. To check whether your setup is correct you can execute the following cell:
#
#
# +
import os
path = os.path.expandvars("$GAMMAPY_DATA")
if not os.path.exists(path):
raise Exception("gammapy-data repository not found!")
else:
print("Great your setup is correct!")
# -
# In case you encounter an error, you can un-comment and execute the following cell to continue. But we recommend to set up your environment correctly as described in [getting started](https://docs.gammapy.org/dev/getting-started/index.html#download-tutorials) after you are done with this notebook.
# +
# os.environ['GAMMAPY_DATA'] = os.path.join(os.getcwd(), '..')
# -
# Now we can continue with the usual IPython notebooks and Python imports:
# %matplotlib inline
import matplotlib.pyplot as plt
import astropy.units as u
from astropy.coordinates import SkyCoord
# ## Maps
#
# The `~gammapy.maps` package contains classes to work with sky images and cubes.
#
# In this section, we will use a simple 2D sky image and will learn how to:
#
# * Read sky images from FITS files
# * Smooth images
# * Plot images
# * Cutout parts from images
# +
from gammapy.maps import Map
gc_3fhl = Map.read("$GAMMAPY_DATA/fermi-3fhl-gc/fermi-3fhl-gc-counts.fits.gz")
# -
# The image is a `~gammapy.maps.WcsNDMap` object:
gc_3fhl
# The shape of the image is 400 x 200 pixel and it is defined using a cartesian projection in galactic coordinates.
#
# The ``geom`` attribute is a `~gammapy.maps.WcsGeom` object:
gc_3fhl.geom
# Let's take a closer look a the `.data` attribute:
gc_3fhl.data
# That looks familiar! It just an *ordinary* 2 dimensional numpy array, which means you can apply any known numpy method to it:
print(f"Total number of counts in the image: {gc_3fhl.data.sum():.0f}")
# To show the image on the screen we can use the ``plot`` method. It basically calls [plt.imshow](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.imshow), passing the `gc_3fhl.data` attribute but in addition handles axis with world coordinates using [astropy.visualization.wcsaxes](https://docs.astropy.org/en/stable/visualization/wcsaxes/) and defines some defaults for nicer plots (e.g. the colormap 'afmhot'):
gc_3fhl.plot(stretch="sqrt");
# To make the structures in the image more visible we will smooth the data using a Gaussian kernel.
gc_3fhl_smoothed = gc_3fhl.smooth(kernel="gauss", width=0.2 * u.deg)
gc_3fhl_smoothed.plot(stretch="sqrt");
# The smoothed plot already looks much nicer, but still the image is rather large. As we are mostly interested in the inner part of the image, we will cut out a quadratic region of the size 9 deg x 9 deg around Vela. Therefore we use `~gammapy.maps.Map.cutout` to make a cutout map:
# + nbsphinx-thumbnail={"tooltip": "Introduction to basic data structures handling."}
# define center and size of the cutout region
center = SkyCoord(0, 0, unit="deg", frame="galactic")
gc_3fhl_cutout = gc_3fhl_smoothed.cutout(center, 9 * u.deg)
gc_3fhl_cutout.plot(stretch="sqrt");
# -
# For a more detailed introduction to `~gammapy.maps`, take a look a the [maps.ipynb](../api/maps.ipynb) notebook.
#
# ### Exercises
#
# * Add a marker and circle at the position of `Sag A*` (you can find examples in [astropy.visualization.wcsaxes](https://docs.astropy.org/en/stable/visualization/wcsaxes/)).
# ## Event lists
#
# Almost any high level gamma-ray data analysis starts with the raw measured counts data, which is stored in event lists. In Gammapy event lists are represented by the `~gammapy.data.EventList` class.
#
# In this section we will learn how to:
#
# * Read event lists from FITS files
# * Access and work with the `EventList` attributes such as `.table` and `.energy`
# * Filter events lists using convenience methods
#
# Let's start with the import from the `~gammapy.data` submodule:
from gammapy.data import EventList
# Very similar to the sky map class an event list can be created, by passing a filename to the `~gammapy.data.EventList.read()` method:
events_3fhl = EventList.read(
"$GAMMAPY_DATA/fermi-3fhl-gc/fermi-3fhl-gc-events.fits.gz"
)
# This time the actual data is stored as an [astropy.table.Table](http://docs.astropy.org/en/stable/api/astropy.table.Table.html) object. It can be accessed with `.table` attribute:
events_3fhl.table
# You can do *len* over event_3fhl.table to find the total number of events.
len(events_3fhl.table)
# And we can access any other attribute of the `Table` object as well:
events_3fhl.table.colnames
# For convenience we can access the most important event parameters as properties on the `EventList` objects. The attributes will return corresponding Astropy objects to represent the data, such as [astropy.units.Quantity](http://docs.astropy.org/en/stable/api/astropy.units.Quantity.html), [astropy.coordinates.SkyCoord](http://docs.astropy.org/en/stable/api/astropy.coordinates.SkyCoord.html) or [astropy.time.Time](http://docs.astropy.org/en/stable/api/astropy.time.Time.html#astropy.time.Time) objects:
events_3fhl.energy.to("GeV")
events_3fhl.galactic
# events_3fhl.radec
events_3fhl.time
# In addition `EventList` provides convenience methods to filter the event lists. One possible use case is to find the highest energy event within a radius of 0.5 deg around the vela position:
# +
# select all events within a radius of 0.5 deg around center
from gammapy.utils.regions import SphericalCircleSkyRegion
region = SphericalCircleSkyRegion(center, radius=0.5 * u.deg)
events_gc_3fhl = events_3fhl.select_region(region)
# sort events by energy
events_gc_3fhl.table.sort("ENERGY")
# and show highest energy photon
events_gc_3fhl.energy[-1].to("GeV")
# -
# ### Exercises
#
# * Make a counts energy spectrum for the galactic center region, within a radius of 10 deg.
# ## Source catalogs
#
# Gammapy provides a convenient interface to access and work with catalog based data.
#
# In this section we will learn how to:
#
# * Load builtins catalogs from `~gammapy.catalog`
# * Sort and index the underlying Astropy tables
# * Access data from individual sources
#
# Let's start with importing the 3FHL catalog object from the `~gammapy.catalog` submodule:
from gammapy.catalog import SourceCatalog3FHL
# First we initialize the Fermi-LAT 3FHL catalog and directly take a look at the `.table` attribute:
fermi_3fhl = SourceCatalog3FHL()
fermi_3fhl.table
# This looks very familiar again. The data is just stored as an [astropy.table.Table](http://docs.astropy.org/en/stable/api/astropy.table.Table.html#astropy.table.Table) object. We have all the methods and attributes of the `Table` object available. E.g. we can sort the underlying table by `Signif_Avg` to find the top 5 most significant sources:
#
#
# +
# sort table by significance
fermi_3fhl.table.sort("Signif_Avg")
# invert the order to find the highest values and take the top 5
top_five_TS_3fhl = fermi_3fhl.table[::-1][:5]
# print the top five significant sources with association and source class
top_five_TS_3fhl[["Source_Name", "ASSOC1", "ASSOC2", "CLASS", "Signif_Avg"]]
# -
# If you are interested in the data of an individual source you can access the information from catalog using the name of the source or any alias source name that is defined in the catalog:
# +
mkn_421_3fhl = fermi_3fhl["3FHL J1104.4+3812"]
# or use any alias source name that is defined in the catalog
mkn_421_3fhl = fermi_3fhl["Mkn 421"]
print(mkn_421_3fhl.data["Signif_Avg"])
# -
# ### Exercises
#
# * Try to load the Fermi-LAT 2FHL catalog and check the total number of sources it contains.
# * Select all the sources from the 2FHL catalog which are contained in the Galactic Center region. The methods `~gammapy.maps.WcsGeom.contains()` and `~gammapy.catalog.SourceCatalog.positions` might be helpful for this. Add markers for all these sources and try to add labels with the source names. The function [ax.text()](http://matplotlib.org/api/_as_gen/matplotlib.axes.Axes.text.html#matplotlib.axes.Axes.text) might be also helpful.
# * Try to find the source class of the object at position ra=68.6803, dec=9.3331
#
# ## Spectral models and flux points
#
# In the previous section we learned how access basic data from individual sources in the catalog. Now we will go one step further and explore the full spectral information of sources. We will learn how to:
#
# * Plot spectral models
# * Compute integral and energy fluxes
# * Read and plot flux points
#
# As a first example we will start with the Crab Nebula:
crab_3fhl = fermi_3fhl["Crab Nebula"]
crab_3fhl_spec = crab_3fhl.spectral_model()
print(crab_3fhl_spec)
# The `crab_3fhl_spec` is an instance of the `~gammapy.modeling.models.PowerLaw2SpectralModel` model, with the parameter values and errors taken from the 3FHL catalog.
#
# Let's plot the spectral model in the energy range between 10 GeV and 2000 GeV:
ax_crab_3fhl = crab_3fhl_spec.plot(
energy_bounds=[10, 2000] * u.GeV, energy_power=0
)
# We assign the return axes object to variable called `ax_crab_3fhl`, because we will re-use it later to plot the flux points on top.
#
# To compute the differential flux at 100 GeV we can simply call the model like normal Python function and convert to the desired units:
crab_3fhl_spec(100 * u.GeV).to("cm-2 s-1 GeV-1")
# Next we can compute the integral flux of the Crab between 10 GeV and 2000 GeV:
crab_3fhl_spec.integral(energy_min=10 * u.GeV, energy_max=2000 * u.GeV).to(
"cm-2 s-1"
)
# We can easily convince ourself, that it corresponds to the value given in the Fermi-LAT 3FHL catalog:
crab_3fhl.data["Flux"]
# In addition we can compute the energy flux between 10 GeV and 2000 GeV:
crab_3fhl_spec.energy_flux(energy_min=10 * u.GeV, energy_max=2000 * u.GeV).to(
"erg cm-2 s-1"
)
# Next we will access the flux points data of the Crab:
print(crab_3fhl.flux_points)
# If you want to learn more about the different flux point formats you can read the specification [here](https://gamma-astro-data-formats.readthedocs.io/en/latest/spectra/flux_points/index.html).
#
# No we can check again the underlying astropy data structure by accessing the `.table` attribute:
crab_3fhl.flux_points.to_table(sed_type="dnde", formatted=True)
# Finally let's combine spectral model and flux points in a single plot and scale with `energy_power=2` to obtain the spectral energy distribution:
ax = crab_3fhl_spec.plot(energy_bounds=[10, 2000] * u.GeV, energy_power=2)
crab_3fhl.flux_points.plot(ax=ax, sed_type="dnde", energy_power=2);
# ### Exercises
#
# * Plot the spectral model and flux points for PKS 2155-304 for the 3FGL and 2FHL catalogs. Try to plot the error of the model (aka "Butterfly") as well. Note this requires the [uncertainties package](https://pythonhosted.org/uncertainties/) to be installed on your machine.
#
# ## What next?
#
# This was a quick introduction to some of the high level classes in Astropy and Gammapy.
#
# * To learn more about those classes, go to the API docs (links are in the introduction at the top).
# * To learn more about other parts of Gammapy (e.g. Fermi-LAT and TeV data analysis), check out the other tutorial notebooks.
# * To see what's available in Gammapy, browse the Gammapy docs or use the full-text search.
# * If you have any questions, ask on the mailing list.
|
docs/tutorials/starting/overview.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import random
import numpy as np
import pandas as pd
import importlib as imp
import matplotlib.pyplot as plt
from tqdm import tqdm, tqdm_notebook
import warnings
warnings.simplefilter('ignore')
pd.options.display.max_columns = 100
# **article**
# http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.457.8027&rep=rep1&type=pdf
from utils import tools, visualize, common
bname = 'benchmarks/B/B-n35-k5.vrp'
problem = tools.get_problem(bname)
# +
tools = imp.reload(tools)
visualize = imp.reload(visualize)
common = imp.reload(common)
visualize.visualize_problem(problem)
for k, v in problem.items():
if k !='locations' and k !='dists':
print(k,"->", v)
plt.savefig('output/images/problem'+bname[-10:]+'.png', dpi=400)
# -
common = imp.reload(common)
solution = common.generate_solution(problem, alpha=0.01, betta=50, verbose=False)
print('Is feasible? {}'.format(common.check_solution(problem, solution, verbose=True)))
sol_cost = common.compute_solution(problem, solution)
print('Solution cost:', sol_cost)
solution
# +
from algorithm import bee_colony, local_search, neighbor_operator
bee_colony = imp.reload(bee_colony)
local_search = imp.reload(local_search)
neighbor_operator = imp.reload(neighbor_operator)
common = imp.reload(common)
ABC = bee_colony.BeeColony(problem)
ABC.set_params(
n_epoch=200,
n_initials=30,
n_onlookers=20,
search_limit=50
)
# %time abc_solution = ABC.solve(alpha=0.1, delta=0.01, gen_alpha=0.01, gen_betta=25)
# -
new_cost = common.compute_solution(problem, abc_solution)
print('Is feasible?',common.check_solution(problem, abc_solution, verbose=True))
print('Is capacity?',common.check_capacity_criteria(problem, abc_solution, verbose=False))
print('ABC cost: {}'.format(new_cost))
print('Optimal cost: {}'.format(problem['optimal']))
visualize = imp.reload(visualize)
visualize.visualize_fitness(ABC.history)
plt.savefig('output/images/history_'+bname[-10:]+'.png', dpi=400)
plt.figure(figsize=(9,4))
plt.plot(ABC.history_alpha, label='alpha')
plt.legend()
plt.grid()
plt.show()
visualize.visualize_problem(problem, solution, figsize=(13,6))
plt.savefig('output/images/beforeABC_'+bname[-10:]+'.png', dpi=400)
visualize.visualize_problem(problem, abc_solution, figsize=(13,6))
depots = list(filter(lambda i: abc_solution[i]==0, range(len(abc_solution))))
for i, route in enumerate(common.get_routes(abc_solution)):
print('Route #{}'.format(i+1),route)
# len(abc_solution), len(np.unique(abc_solution))
plt.savefig('output/images/afterABC_'+bname[-10:]+'.png', dpi=400)
# ## B-benchmarks
# +
# %%time
import glob
from datetime import datetime
tools = imp.reload(tools)
bee_colony = imp.reload(bee_colony)
local_search = imp.reload(local_search)
common = imp.reload(common)
neighbor_operator = imp.reload(neighbor_operator)
b_benchmarks = glob.glob('benchmarks/B/*')
info_dct = dict(benchmark=[],
n_locations=[],
n_trucks=[],
capacity=[],
optimal_cost=[],
ABC_cost=[],
ABC_time=[],
is_feasible=[],
error=[],
abc_solution=[],
abc_epochs=[],
abc_employers=[],
abc_onlookers=[],
abc_search_limit=[])
for itr, benchmark in enumerate(b_benchmarks[:]):
problem = tools.get_problem(benchmark)
bench_name = benchmark[benchmark.index("\\")+1:]
print('#{}'.format(itr), bench_name,'...')
ABC = bee_colony.BeeColony(problem)
ABC.set_params(
n_epoch=300,
n_initials=problem['n_locations'],
n_onlookers=5,
search_limit=problem['n_locations']
)
start_time = datetime.now()
alpha = problem['n_locations'] / 100
delta = 0.01
gen_alpha = 0.5
gen_betta = problem['n_locations']
abc_solution = ABC.solve(alpha=alpha, delta=delta, gen_betta=gen_betta)
end_time = (datetime.now() - start_time).total_seconds()
abc_cost = common.compute_solution(problem, abc_solution)
is_feasible = common.check_solution(problem, abc_solution, verbose=True)
error = (abc_cost - problem['optimal']) / problem['optimal']
print('epoch:',ABC.n_epoch, 'initials:', ABC.n_initials,'search_limit:', ABC.search_limit)
info_dct['benchmark'] += [bench_name]
info_dct['n_locations'] += [problem['n_locations']]
info_dct['n_trucks'] += [problem['n_trucks']]
info_dct['capacity'] += [problem['capacity']]
info_dct['optimal_cost'] += [problem['optimal']]
info_dct['ABC_cost'] += [abc_cost]
info_dct['ABC_time'] += [end_time]
info_dct['is_feasible'] += [is_feasible]
info_dct['error'] += [error]
info_dct['abc_solution'] += [abc_solution]
info_dct['abc_epochs'] += [ABC.n_epoch]
info_dct['abc_employers'] += [ABC.n_initials]
info_dct['abc_onlookers'] += [ABC.n_onlookers]
info_dct['abc_search_limit'] += [ABC.search_limit]
print(alpha, delta, gen_betta)
if not is_feasible:
print('Does not feasible...')
break
tools.write_solution(abc_solution, abc_cost, filename='B_test/'+bench_name[:-4]+'.sol')
# -
b_stat = pd.DataFrame.from_dict(info_dct)
columns = ['benchmark', 'n_locations', 'n_trucks', 'capacity', 'optimal_cost', 'ABC_cost', 'ABC_time', 'error', 'is_feasible']
print(b_stat[columns].to_markdown())
# | | benchmark | n_locations | n_trucks | capacity | optimal_cost | ABC_cost | ABC_time | error | is_feasible |
# |---:|:--------------|--------------:|-----------:|-----------:|---------------:|-----------:|-----------:|----------:|:--------------|
# | 0 | B-n31-k5.vrp | 31 | 5 | 100 | 672 | 706.569 | 20.9377 | 0.0514423 | True |
# | 1 | B-n34-k5.vrp | 34 | 5 | 100 | 788 | 808.974 | 23.4713 | 0.0266171 | True |
# | 2 | B-n35-k5.vrp | 35 | 5 | 100 | 955 | 996.195 | 24.5341 | 0.0431363 | True |
# | 3 | B-n38-k6.vrp | 38 | 6 | 100 | 805 | 820.224 | 28.2701 | 0.0189118 | True |
# | 4 | B-n39-k5.vrp | 39 | 5 | 100 | 549 | 567.277 | 26.7929 | 0.0332911 | True |
# | 5 | B-n41-k6.vrp | 41 | 6 | 100 | 829 | 947.016 | 30.4575 | 0.142359 | True |
# | 6 | B-n43-k6.vrp | 43 | 6 | 100 | 742 | 777.761 | 33.5126 | 0.0481955 | True |
# | 7 | B-n44-k7.vrp | 44 | 7 | 100 | 909 | 985.969 | 36.6416 | 0.0846748 | True |
# | 8 | B-n45-k5.vrp | 45 | 5 | 100 | 751 | 796.818 | 32.5534 | 0.0610096 | True |
# | 9 | B-n45-k6.vrp | 45 | 6 | 100 | 678 | 768.834 | 37.3276 | 0.133974 | True |
# | 10 | B-n50-k7.vrp | 50 | 7 | 100 | 741 | 763.865 | 41.6256 | 0.0308568 | True |
# | 11 | B-n50-k8.vrp | 50 | 8 | 100 | 1312 | 1354.85 | 44.7541 | 0.0326566 | True |
# | 12 | B-n51-k7.vrp | 51 | 7 | 100 | 1032 | 1124.62 | 42.9064 | 0.0897509 | True |
# | 13 | B-n52-k7.vrp | 52 | 7 | 100 | 747 | 818.84 | 43.2389 | 0.0961716 | True |
# | 14 | B-n56-k7.vrp | 56 | 7 | 100 | 707 | 792.316 | 47.4659 | 0.120674 | True |
# | 15 | B-n57-k7.vrp | 57 | 7 | 100 | 1153 | 1555.21 | 66.0018 | 0.348837 | True |
# | 16 | B-n57-k9.vrp | 57 | 9 | 100 | 1598 | 1740.66 | 57.6039 | 0.0892725 | True |
# | 17 | B-n63-k10.vrp | 63 | 10 | 100 | 1496 | 1775.97 | 75.6478 | 0.187143 | True |
# | 18 | B-n64-k9.vrp | 64 | 9 | 100 | 861 | 1082.98 | 75.0857 | 0.257812 | True |
# | 19 | B-n66-k9.vrp | 66 | 9 | 100 | 1316 | 1611.2 | 82.5796 | 0.224317 | True |
# | 20 | B-n67-k10.vrp | 67 | 10 | 100 | 1032 | 1206.82 | 86.4656 | 0.169402 | True |
# | 21 | B-n68-k9.vrp | 68 | 9 | 100 | 1272 | 1442.81 | 67.5301 | 0.134288 | True |
# | 22 | B-n78-k10.vrp | 78 | 10 | 100 | 1221 | 1602.17 | 93.2046 | 0.312182 | True |
# ## A-benchmarks
#
# +
# %%time
tools = imp.reload(tools)
bee_colony = imp.reload(bee_colony)
local_search = imp.reload(local_search)
common = imp.reload(common)
neighbor_operator = imp.reload(neighbor_operator)
b_benchmarks = glob.glob('benchmarks/A/*')
info_dct = dict(benchmark=[],
n_locations=[],
n_trucks=[],
capacity=[],
optimal_cost=[],
ABC_cost=[],
ABC_time=[],
is_feasible=[],
error=[],
abc_solution=[],
abc_epochs=[],
abc_employers=[],
abc_onlookers=[],
abc_search_limit=[])
for itr, benchmark in enumerate(b_benchmarks[:]):
problem = tools.get_problem(benchmark)
bench_name = benchmark[benchmark.index("\\")+1:]
print('#{}'.format(itr), bench_name,'...')
ABC = bee_colony.BeeColony(problem)
ABC.set_params(
n_epoch=400,
n_initials=problem['n_locations'],
n_onlookers=10,
search_limit=problem['n_locations']
)
start_time = datetime.now()
alpha = problem['n_locations'] / 80
delta = 0.05
gen_alpha = 0.3
gen_betta = problem['n_locations']
abc_solution = ABC.solve(alpha=alpha, delta=delta, gen_betta=gen_betta)
end_time = (datetime.now() - start_time).total_seconds()
abc_cost = common.compute_solution(problem, abc_solution)
is_feasible = common.check_solution(problem, abc_solution, verbose=True)
error = (abc_cost - problem['optimal']) / problem['optimal']
print('epoch:',ABC.n_epoch, 'initials:', ABC.n_initials,'search_limit:', ABC.search_limit,'\n\n')
info_dct['benchmark'] += [bench_name]
info_dct['n_locations'] += [problem['n_locations']]
info_dct['n_trucks'] += [problem['n_trucks']]
info_dct['capacity'] += [problem['capacity']]
info_dct['optimal_cost'] += [problem['optimal']]
info_dct['ABC_cost'] += [abc_cost]
info_dct['ABC_time'] += [end_time]
info_dct['is_feasible'] += [is_feasible]
info_dct['error'] += [error]
info_dct['abc_solution'] += [abc_solution]
info_dct['abc_epochs'] += [ABC.n_epoch]
info_dct['abc_employers'] += [ABC.n_initials]
info_dct['abc_onlookers'] += [ABC.n_onlookers]
info_dct['abc_search_limit'] += [ABC.search_limit]
print(alpha, delta, gen_betta)
if not is_feasible:
print('Does not feasible...')
break
tools.write_solution(abc_solution, abc_cost, filename='A/'+bench_name[:-4]+'.sol')
# -
a_stat = pd.DataFrame.from_dict(info_dct)
print(a_stat[columns].to_markdown())
# | | benchmark | n_locations | n_trucks | capacity | optimal_cost | ABC_cost | ABC_time | error | is_feasible |
# |---:|:--------------|--------------:|-----------:|-----------:|---------------:|-----------:|-----------:|----------:|:--------------|
# | 0 | A-n32-k5.vrp | 32 | 5 | 100 | 784 | 793.689 | 34.4186 | 0.0123585 | True |
# | 1 | A-n33-k5.vrp | 33 | 5 | 100 | 661 | 677.849 | 33.94 | 0.0254898 | True |
# | 2 | A-n33-k6.vrp | 33 | 6 | 100 | 742 | 789.651 | 35.5225 | 0.0642203 | True |
# | 3 | A-n34-k5.vrp | 34 | 5 | 100 | 778 | 890.053 | 34.7679 | 0.144027 | True |
# | 4 | A-n36-k5.vrp | 36 | 5 | 100 | 799 | 902.63 | 39.6765 | 0.129699 | True |
# | 5 | A-n37-k5.vrp | 37 | 5 | 100 | 669 | 722.61 | 40.3947 | 0.0801341 | True |
# | 6 | A-n37-k6.vrp | 37 | 6 | 100 | 949 | 1002.95 | 41.3557 | 0.0568444 | True |
# | 7 | A-n38-k5.vrp | 38 | 5 | 100 | 730 | 796.663 | 39.2311 | 0.0913186 | True |
# | 8 | A-n39-k5.vrp | 39 | 5 | 100 | 822 | 917.818 | 41.4462 | 0.116567 | True |
# | 9 | A-n39-k6.vrp | 39 | 6 | 100 | 831 | 1039.23 | 46.8593 | 0.250572 | True |
# | 10 | A-n44-k6.vrp | 44 | 6 | 100 | 937 | 987.494 | 51.036 | 0.0538888 | True |
# | 11 | A-n45-k6.vrp | 45 | 6 | 100 | 944 | 1209.34 | 55.1767 | 0.281077 | True |
# | 12 | A-n45-k7.vrp | 45 | 7 | 100 | 1146 | 1270.99 | 62.4989 | 0.109066 | True |
# | 13 | A-n46-k7.vrp | 46 | 7 | 100 | 914 | 1030.53 | 73.3458 | 0.127491 | True |
# | 14 | A-n48-k7.vrp | 48 | 7 | 100 | 1073 | 1195.6 | 71.0007 | 0.114257 | True |
# | 15 | A-n53-k7.vrp | 53 | 7 | 100 | 1010 | 1151.61 | 82.1195 | 0.140208 | True |
# | 16 | A-n54-k7.vrp | 54 | 7 | 100 | 1167 | 1331.5 | 86.6397 | 0.140958 | True |
# | 17 | A-n55-k9.vrp | 55 | 9 | 100 | 1073 | 1147.89 | 98.8293 | 0.0697974 | True |
# | 18 | A-n60-k9.vrp | 60 | 9 | 100 | 1354 | 1502.62 | 118.18 | 0.109762 | True |
# | 19 | A-n61-k9.vrp | 61 | 9 | 100 | 1034 | 1398.01 | 129.125 | 0.352039 | True |
# | 20 | A-n62-k8.vrp | 62 | 8 | 100 | 1288 | 1517.37 | 119.281 | 0.178084 | True |
# | 21 | A-n63-k10.vrp | 63 | 10 | 100 | 1314 | 1515.19 | 127.642 | 0.153113 | True |
# | 22 | A-n63-k9.vrp | 63 | 9 | 100 | 1616 | 1959.36 | 131.052 | 0.212478 | True |
# | 23 | A-n64-k9.vrp | 64 | 9 | 100 | 1401 | 1664.51 | 122.621 | 0.188088 | True |
# | 24 | A-n65-k9.vrp | 65 | 9 | 100 | 1174 | 1558.45 | 133.355 | 0.327473 | True |
# | 25 | A-n69-k9.vrp | 69 | 9 | 100 | 1159 | 1434.53 | 154.796 | 0.237731 | True |
# | 26 | A-n80-k10.vrp | 80 | 10 | 100 | 1763 | 2203.57 | 184.733 | 0.249898
|
VehicleRoutingProblem.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # # Testing Thresholding
# +
import numpy as np
import cv2
from tkinter import Tk, filedialog
# let user choose file, returns image
def open_image():
root = Tk()
root.fileName = filedialog.askopenfilename(
initialdir="./samples",
title="Select an Image",
filetypes=((".jpg", "*.jpg"), (".png", "*.png"), ("all files", "*.*")),
)
fname = root.fileName
image = cv2.imread(fname)
return image
def thresholding(image, thresh):
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
blurred = cv2.GaussianBlur(gray, (9, 9), 0)
retval, th = cv2.threshold(blurred, 0, 255, cv2.THRESH_BINARY + cv2.THRESH_OTSU)
return th
# +
thresh_val = 40
def change_thresh(new_thresh):
global thresh_val
thresh_val = new_thresh
open_image()
cv2.namedWindow("Thresholding", cv2.WINDOW_AUTOSIZE)
cv2.createTrackbar("Thresh value", "Thresholding", thresh_val, 200, change_thresh)
while True:
thresholded = thresholding(linearized_image, sigma)
cv2.imshow("Thresholding", thresholded)
key = cv2.waitKey(1)
if key == 27:
break
# -
|
notebooks/.ipynb_checkpoints/test-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Metrics
# * **MAE** - Mean Absolute Error - easy to understand
# * **MSE** - Mean Squared Error - more popular - it punishes larger error
# * **RMSE** - Root Mean Squared Error - even more popular - it's interpretable in y units
true_output = [100, 50, 30, 20]
pred_output = [90, 50, 50, 30]
# ### Mean Absolute Error
from sklearn import metrics
# +
print((10 + 0 + 20 + 10) / 4.)
# Or
print(metrics.mean_absolute_error(true_output, pred_output))
# -
# ### Mean Squared Error
# +
print((10 ** 2 + 0 ** 2 + 20 ** 2 + 10 ** 2) / 4.)
# Or
print(metrics.mean_squared_error(true_output, pred_output))
# -
# ### Root Mean Squared Error
import numpy as np
# +
print(np.sqrt((10 ** 2 + 0 ** 2 + 20 ** 2 + 10 ** 2) / 4.))
# Or
print(np.sqrt(metrics.mean_squared_error(true_output, pred_output)))
|
numpy/metrics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # GO enrichment with `goenrich`
#
# In the following we will perform a gene ontology enrichment analysis using 'goenrich' and the example data provided by the published [`WebGestalt`](http://bioinfo.vanderbilt.edu/webgestalt/) tool.
#
# ### 1. run `WebGestalt`
#
# First we can run `WebGestalt` on their website using the example data provided (`final_sig_file_XXX.tsv`). I wrote a small script `wegestalt.py` to parse their weird output format into something more convenient.
import webgestalt
web = webgestalt.read('../db/webgestalt_example_results/files/final_sig_file_1439064580.tsv')
web.head()
# ### 2. parse example into pandas
#
# We need the `geneinfo` table in order to map the ids from `GeneID` to `Symbol`
import pandas as pd
geneinfo = pd.read_table('../db/Homo_sapiens.gene_info.gz', comment='#', usecols=[1,2], names=['GeneID', 'Symbol'])
interesting = pd.read_table('../db/interestingGenes.txt', names=['Symbol']).merge(geneinfo)
reference = pd.read_table('../db/referenceGenes.txt', names=['Symbol']).merge(geneinfo)
# ### 3. build the ontology
#
# Now we can move on to use `goenrich` to build the ontology DAG. Additionally we need the set of all GO annotation from `gene2go`.
import goenrich
O = goenrich.obo.ontology('../db/go-basic.obo')
gene2go = goenrich.read.gene2go('../db/gene2go.gz')
# ### 4. build the background-set and propagate
#
# Using a small helper functon we can generate a background set from the `pandas.DataFrames` we parsed before. We use the background set to annotate our ontology. Because of the DAG structure of the ontology we can do so by first perfoming a topological sort on the ontology.
import goenrich.tools as tools
background = tools.generate_background(gene2go, reference, 'GO_ID', 'GeneID')
goenrich.enrich.propagate(O, background, 'reference')
# ### 5. run the query
#
# Using the set of interesting genes as query we can perform our analysis. Additional options for filtering the categories we would like to test on can be passed to `analyze`. One might choose to extract small (`min_category_size`), too specific (`max_category_depth`) or too big/generic (`max_category_size`) categories.
#
# The underlying test is the hypergeometric test `hypergeom.sf(x, M, n, N)`. All $p$-values are corrected for multiple testing using the Benjamini-Hochberge method before a significance cut-off of $\alpha=0.05$ is applied.
query = set(interesting['GeneID'])
options = { 'min_category_size' : 2, 'max_category_size' : 100000,
'max_category_depth' : 100000 }
enrichment = goenrich.enrich.analyze(O, query, 'reference', **options).dropna().sort_values('q')
enrichment.head()
# ### 6. comparison to `WebGestalt`
#
# We can first look at the category sizes and intersections
#
# `Webgestalt`: `C`, `O`
#
# `goenrich`: `n`, `x`
df = pd.merge(enrichment, web)
df[['C', 'O', 'n', 'x']].head()
# we can see come differences which might be due to different version of the `gene2go` annotation file, the ontology tree, or artifacts of the id mapping.
#
# To quantify to consistency of both predictions we can therefore look at the correlation between the two
# %matplotlib inline
import seaborn as sns
sns.corrplot(df[['C', 'O', 'rawP', 'n', 'x', 'p']], method='spearman')
# The resulting correlation between `WebGestalt` and `goenrich` $p$-values is $0.92$
|
examples/go_enrichment_with_goenrich.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Ak2-OoRHr4FF" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive')
# + id="xWEEN_SWSR_C" colab_type="code" colab={}
import os
par = "/content/drive/My Drive"
directory = "Gdrive-Colab-Upload"
path = os.path.join(par, directory)
os.mkdir(path)
# + id="DUsHVL-pallB" colab_type="code" colab={}
import requests
import re
def getFilename_fromCd(cd):
if not cd:
return None
fname = re.findall('filename=(.+)', cd)
if len(fname) == 0:
return None
return fname[0]
url = input("Enter Download Link : ")
if url.find('/'):
filename = url.rsplit('/', 1)[1]
else:
filename = getFilename_fromCd(r.headers.get('content-disposition'))
if filename is None:
filename = input("Enter Filename : ")
r = requests.get(url, allow_redirects=True)
print("\n",filename)
print(r.headers.get('content-type'))
size = r.headers.get('content-length')
size = int(size)/1048576
print(size, "MB")
size = int(size)/1024
print(size, "GB")
open("/content/drive/My Drive/Gdrive-Colab-Upload/"+filename, 'wb').write(r.content)
|
Google_Drive_Uploader.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
# Q3 API
from layers import dense
epochs = 20
batch_size = 500
learning_rate = 0.01
mnist = np.load('mnist.npz')
# split training data (60,000) into 55,000 for train and 5,000 for validate images
train, x_test = mnist['x_train'], mnist['x_test']
train = train.reshape((60000, 784))/255
x_train = train[:55000].copy()
x_val = train[55000:].copy()
x_test = x_test.reshape((10000, 784))/255
# labels
train_labels = mnist['y_train']
y_train = np.eye(10)[train_labels[:55000]]
y_val = np.eye(10)[train_labels[55000:]]
y_test = np.eye(10)[mnist['y_test']]
# input
x_p = tf.placeholder(tf.float32, [None, 784])
# output
y_p = tf.placeholder(tf.float32, [None, 10])
hidden1 = dense(x=x_p, in_length=784, neurons=300, activation=tf.nn.relu, layer_name='Layer_1', dev=0.08)
hidden2 = dense(x=hidden1, in_length=300, neurons=100, activation=tf.nn.relu, layer_name='Layer_2', dev=0.08)
output = dense(x=hidden2, in_length=100, neurons=10, activation=tf.nn.softmax, layer_name='Layer_Output')
y_clipped = tf.clip_by_value(output, 1e-10, 0.9999999)
cross_entropy = -tf.reduce_mean(tf.reduce_sum(y_p * tf.log(y_clipped)+ (1 - y_p) * tf.log(1 - y_clipped), axis=1))
optimiser = tf.train.GradientDescentOptimizer(learning_rate=learning_rate).minimize(cross_entropy)
labels = tf.argmax(y_p, 1)
predictions = tf.argmax(output, 1)
acc, acc_op = tf.metrics.accuracy(labels, predictions)
conmat = tf.confusion_matrix(labels, predictions)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
sess.run(tf.local_variables_initializer())
avg_loss = []
validate_accuracy = []
total_batches = x_train.shape[0] // batch_size
# Training
for e in range(epochs):
avg_loss.append(0.0)
for b in range(total_batches):
start = b*batch_size
end = (b+1)*batch_size
batch = sess.run([optimiser, cross_entropy],
feed_dict={x_p: x_train[start:end], y_p: y_train[start:end]})
avg_loss[e] += batch[1] / total_batches
# Validation
accuracy = sess.run(acc_op,
feed_dict={x_p: x_val, y_p: y_val})
validate_accuracy.append(accuracy)
print("Epoch: {:2d}".format(e + 1), "train_loss =", "{:.4f}".format(avg_loss[e]), "validate_accuracy =", "{:.4f}".format(validate_accuracy[e]))
# Testing
test_accuracy, confusion_mat = sess.run([acc_op, conmat],
feed_dict={x_p:x_test, y_p:y_test})
print('Testing Accuracy:', test_accuracy)
print('Confusion Matrix:', confusion_mat)
tf.io.write_graph(sess.graph_def, 'graphs/', 'mnist-v2.pbtxt')
np.savetxt('mnistv2-conmat.txt', confusion_mat, fmt='%4d', delimiter=' & ', newline='\\\\\ \hline\n')
plt.xlabel('Epoch')
plt.ylabel('Cross Entropy Loss')
plt.plot(avg_loss[None:])
plt.show()
plt.xlabel('Epoch')
plt.ylabel('Validation Accuracy')
plt.plot(validate_accuracy)
plt.show()
True_positives = np.diag(confusion_mat)
False_positives = np.sum(confusion_mat, axis=1) - True_positives
False_negatives = np.sum(confusion_mat, axis=0) - True_positives
Precision = True_positives / (True_positives + False_positives)
print("Precision:", Precision)
Recall = True_positives / (True_positives + False_negatives)
print("\nRecall:", Recall)
F_scores = (2*Precision*Recall) / (Recall+Precision)
print("\nF_scores:", F_scores)
plt.plot(Precision, label='Precision')
plt.plot(Recall, label='Recall')
plt.plot(F_scores, label='F Scores')
plt.ylabel('Score')
plt.xlabel('Class')
plt.legend()
plt.show()
|
Assignments/Assignment_1/Q3/Q3_mnist_v2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Approximate solvers for the Euler equations of gas dynamics
# In this chapter we discuss approximate solvers for the one-dimensional Euler equations:
#
# \begin{align}
# \rho_t + (\rho u)_x & = 0 \\
# (\rho u)_t + (\rho u^2 + p)_x & = 0 \\
# E_t + ((E+p)u)_x & = 0.
# \end{align}
#
# As in [Euler_equations.ipynb](Euler_equations.ipynb), we focus on the case of an ideal gas, for which the total energy is given by
#
# \begin{align} \label{EA:EOS}
# E = \frac{p}{\gamma-1} + \frac{1}{2}\rho u^2.
# \end{align}
# ## Roe solver
# We first derive a Roe solver for the Euler equations, following the same approach as in [shallow_water_approximate_solvers.ipynb](shallow_water_approximate_solvers.ipynb). Namely, we assume that $\hat{A} = f'(\hat{q})$ for some average state $\hat{q}$, and impose the condition of conservation:
#
# \begin{align} \label{EA:cons}
# f'(\hat{q}) (q_r - q_l) & = f(q_r) - f(q_l).
# \end{align}
#
# We will need the following quantities:
#
# \begin{align}
# q & = \begin{pmatrix} \rho \\ \rho u \\ E \end{pmatrix}, \ \ \ \ \ \ f(q) = \begin{pmatrix} \rho u \\ \rho u^2 + p \\ H u \rho \end{pmatrix}, \\
# f'(\hat{q}) & = \begin{pmatrix}
# 0 & 1 & 0 \\
# \frac{\gamma-3}{2}\hat{u}^2 & (3-\gamma)\hat{u} & \gamma-1 \\
# \frac{\gamma-1}{2}\hat{u}^3 - \hat{u}\hat{H} & \hat{H} - (\gamma-1)\hat{u}^2 & \gamma \hat{u} \end{pmatrix}.
# \end{align}
#
# Here $H = \frac{E+p}{\rho}$ is the enthalpy. We have rewritten most expressions involving $E$ in terms of $H$ because it simplifies the derivation that follows. We now solve (\ref{EA:cons}) to find $\hat{u}$ and $\hat{H}$. It turns out that, for the case of a polytropic ideal gas, the average density $\hat{\rho}$ plays no role in the Roe solver.
#
# The first equation of (\ref{EA:cons}) is an identity, satisfied independently of our choice of $\hat{q}$. The second equation is (using (\ref{EA:EOS}))
#
# \begin{align}
# \frac{\gamma-3}{2}\hat{u}^2 (\rho_r - \rho_l) + (3-\gamma)\hat{u}(\rho_r u_r - \rho_l u_l) + (\gamma-1)\left( \frac{p_r-p_l}{\gamma-1} + \frac{1}{2}(\rho_r u_r^2 - \rho_l u_l^2) \right) & = \rho_r u_r^2 - \rho_l u_l^2 + p_r - p_l,
# \end{align}
#
# which simplifies to a quadratic equation for $\hat{u}$:
#
# \begin{align} \label{EA:u_quadratic}
# (\rho_r - \rho_l)\hat{u}^2 - 2(\rho_r u_r - \rho_l u_l) \hat{u} + (\rho_r u_r^2 - \rho_l u_l^2) & = 0,
# \end{align}
#
# with roots
#
# \begin{align}
# \hat{u}_\pm & = \frac{\rho_r u_r - \rho_l u_l \mp \sqrt{\rho_r \rho_l} (u_l - u_r)}{\rho_r - \rho_l} = \frac{\sqrt{\rho_r} u_r \pm \sqrt{\rho_l} u_l}{\sqrt{\rho_r}\pm\sqrt{\rho_l}}
# \end{align}
#
# Notice that this is identical to the Roe average of the velocity for the shallow water equations, if we replace the density $\rho$ with depth $h$. As before, we choose the root $u_+$ since it is well defined for all values of $\rho_r, \rho_l$.
# Next we find $\hat{H}$ by solving the last equation of (\ref{EA:cons}), which reads
# \begin{align}
# \left( \frac{\gamma-1}{2}\hat{u}^3 - \hat{u}\hat{H} \right)(\rho_r - \rho_l) + \left( \hat{H} - (\gamma-1)\hat{u}^2 \right)(\rho_r u_r - \rho_l u_l) + \gamma \hat{u}(E_r - E_l) & = H_r u_r \rho_r - H_l u_l \rho_l.
# \end{align}
# We can simplify this using the equality $\gamma E = \rho H + \frac{\gamma-1}{2}\rho u^2$ and solve for $\hat{H}$ to find
# \begin{align}
# \hat{H} & = \frac{\rho_r H_r (u_r - \hat{u}_+) - \rho_l H_l (u_l - \hat{u}_+)}{\rho_r u_r - \rho_l u_l - \hat{u}_\pm(\rho_r -\rho_l)} \\
# & = \frac{\rho_r H_r (u_r - \hat{u}_+) - \rho_l H_l (u_l - \hat{u}_+)}{\pm\sqrt{\rho_r \rho_l}(u_r-u_l)} \\
# & = \frac{\rho_r H_r - \rho_l H_l \mp\sqrt{\rho_r \rho_l}(H_r - H_l)}{\rho_r - \rho_l} \\
# & = \frac{\sqrt{\rho_r}H_r \pm \sqrt{\rho_l} H_l}{\sqrt{\rho_r}\pm\sqrt{\rho_l}}.
# \end{align}
# Once more, we take the plus sign in the final expression for $\hat{H}$.
# To implement the Roe solver, we also need the eigenvalues and eigenvectors of the averaged flux Jacobian $f'(\hat{q})$. These are just the eigenvalues of the true Jacobian, evaluated at the averaged state:
# \begin{align}
# \lambda_1 & = \hat{u} - \hat{c}, & \lambda_2 & = \hat{u} & \lambda_3 & = \hat{u} + \hat{c},
# \end{align}
# \begin{align}
# r_1 & = \begin{bmatrix} 1 \\ \hat{u}-\hat{c} \\ \hat{H}-\hat{u}\hat{c}\end{bmatrix} &
# r_2 & = \begin{bmatrix} 1 \\ \hat{u} \\ \frac{1}{2}\hat{u}^2 \end{bmatrix} &
# r_3 & = \begin{bmatrix} 1 \\ \hat{u}+\hat{c} \\ \hat{H}+\hat{u}\hat{c}\end{bmatrix}.
# \end{align}
# Here $\hat{c} = \sqrt{(\gamma-1)(\hat{H}-\hat{u}^2/2)}$.
#
# Solving the system of equations
# \begin{align}
# q_r - q_l & = \sum_{p=1}^3 {\mathcal W}_p = \sum_{p=1}^3 \alpha_p r_p
# \end{align}
# for the wave strengths gives
# \begin{align}
# \alpha_2 & = \delta_1 + (\gamma-1)\frac{\hat{u}\delta_2 - \delta_3}{\hat{c}^2} \\
# \alpha_3 & = \frac{\delta_2 + (\hat{c}-\hat{u})\delta_1 - \hat{c}\alpha_2}{2\hat{c}} \\
# \alpha_1 & = \delta_1 - \alpha_2 - \alpha_3,
# \end{align}
# where $\delta = q_r - q_l$. We now have everything we need to implement the Roe solver.
# + tags=["hide"]
# %matplotlib inline
# + tags=["hide"]
# %config InlineBackend.figure_format = 'svg'
import numpy as np
from exact_solvers import Euler
from utils import riemann_tools
from ipywidgets import interact
from ipywidgets import widgets
State = Euler.Primitive_State
# +
def roe_averages(q_l, q_r, gamma=1.4):
rho_sqrt_l = np.sqrt(q_l[0])
rho_sqrt_r = np.sqrt(q_r[0])
p_l = (gamma-1.)*(q_l[2]-0.5*(q_l[1]**2)/q_l[0])
p_r = (gamma-1.)*(q_r[2]-0.5*(q_r[1]**2)/q_r[0])
denom = rho_sqrt_l + rho_sqrt_r
u_hat = (q_l[1]/rho_sqrt_l + q_r[1]/rho_sqrt_r)/denom
H_hat = ((q_l[2]+p_l)/rho_sqrt_l + (q_r[2]+p_r)/rho_sqrt_r)/denom
c_hat = np.sqrt((gamma-1)*(H_hat-0.5*u_hat**2))
return u_hat, c_hat, H_hat
def Euler_roe(q_l, q_r, gamma=1.4):
"""
Approximate Roe solver for the Euler equations.
"""
rho_l = q_l[0]
rhou_l = q_l[1]
u_l = rhou_l/rho_l
rho_r = q_r[0]
rhou_r = q_r[1]
u_r = rhou_r/rho_r
u_hat, c_hat, H_hat = roe_averages(q_l, q_r, gamma)
delta = q_r - q_l
s1 = u_hat - c_hat
s2 = u_hat
s3 = u_hat + c_hat
alpha2 = (gamma-1.)/c_hat**2 *((H_hat-u_hat**2)*delta[0]+u_hat*delta[1]-delta[2])
alpha3 = (delta[1] + (c_hat - u_hat)*delta[0] - c_hat*alpha2) / (2.*c_hat)
alpha1 = delta[0] - alpha2 - alpha3
r1 = np.array([1., u_hat-c_hat, H_hat - u_hat*c_hat])
r2 = np.array([1., u_hat, 0.5*u_hat**2])
q_l_star = q_l + alpha1*r1
q_r_star = q_l_star + alpha2*r2
states = np.column_stack([q_l,q_l_star,q_r_star,q_r])
speeds = [s1, s2, s3]
wave_types = ['contact','contact', 'contact']
def reval(xi):
rho = (xi<s1)*states[0,0] + (s1<=xi)*(xi<s2)*states[0,1] + (s2<=xi)*(xi<s3)*states[0,2] + (s3<=xi)*states[0,3]
mom = (xi<s1)*states[1,0] + (s1<=xi)*(xi<s2)*states[1,1] + (s2<=xi)*(xi<s3)*states[1,2] + (s3<=xi)*states[1,3]
E = (xi<s1)*states[2,0] + (s1<=xi)*(xi<s2)*states[2,1] + (s2<=xi)*(xi<s3)*states[2,2] + (s3<=xi)*states[2,3]
return rho, mom, E
return states, speeds, reval, wave_types
# -
# ### Examples
#
# Let's compare the Roe approximation to the exact solution. As a first example, we use the Sod shock tube.
def compare_solutions(left_state, right_state, solvers=['Exact','HLL']):
q_l = np.array(Euler.primitive_to_conservative(*left_state))
q_r = np.array(Euler.primitive_to_conservative(*right_state))
outputs = []
states = {}
for solver in solvers:
if solver.lower() == 'exact':
outputs.append(Euler.exact_riemann_solution(q_l,q_r))
if solver.lower() == 'hll':
outputs.append(Euler_hll(q_l, q_r))
states['hll'] = outputs[-1][0]
if solver.lower() == 'roe':
outputs.append(Euler_roe(q_l, q_r))
states['roe'] = outputs[-1][0]
plot_function = riemann_tools.make_plot_function([val[0] for val in outputs],
[val[1] for val in outputs],
[val[2] for val in outputs],
[val[3] for val in outputs],
solvers,
layout='vertical',
variable_names=Euler.primitive_variables,
derived_variables=Euler.cons_to_prim)
interact(plot_function, t=widgets.FloatSlider(min=0,max=0.9,step=0.1));
return states
# +
left_state = State(Density = 3.,
Velocity = 0.,
Pressure = 3.)
right_state = State(Density = 1.,
Velocity = 0.,
Pressure = 1.)
states = compare_solutions(left_state,right_state,solvers=['Exact','Roe'])
# -
Euler.phase_plane_plot(left_state,right_state,approx_states=states['roe'])
# Recall that in the true solution the middle wave is a contact discontinuity and carries only a jump in the density. For that reason the three-dimensional phase space plot is generally shown projected onto the pressure-velocity plane as shown above: The two intermediate states in the true solution have the same pressure and velocity, and so are denoted by a single Middle state in the phase plane plot.
#
# The Roe solver, on the other hand, generates a middle wave that carries a jump in all 3 variables and there are two green dots appearing in the plot above for the two middle states (though the pressure jump is quite small in this example). For a Riemann problem like this one with zero initial velocity on both sides, the Roe average velocity must also be zero, so the middle wave is stationary; this is of course not typically true in the exact solution, even when $u_l=u_r=0$.
# Here is a second example. Experiment with the initial states to explore how the Roe solution compares to the exact solution.
# +
left_state = State(Density = 0.1,
Velocity = 0.,
Pressure = 0.1)
right_state = State(Density = 1.,
Velocity = 1.,
Pressure = 1.)
states = compare_solutions(left_state,right_state,solvers=['Exact','Roe'])
# -
Euler.phase_plane_plot(left_state,right_state,approx_states=states['roe'])
# ### Single-shock solution
# Next we demonstrate the exactness property of the Roe solver by applying it to a case where the left and right states are connected by a single shock wave.
# +
M = 2. # Mach number of the shock wave
gamma = 1.4
mu = 2*(M**2-1)/(M*(gamma+1.))
right_state = State(Density = 1.,
Velocity = 0.,
Pressure = 1.)
c_r = np.sqrt(gamma*right_state.Pressure/right_state.Density)
rho_l = right_state.Density * M/(M-mu)
p_l = right_state.Pressure * ((2*M**2-1)*gamma+1)/(gamma+1)
u_l = mu*c_r
left_state = State(Density = rho_l,
Velocity = u_l,
Pressure = p_l)
states = compare_solutions(left_state,right_state,solvers=['Exact','Roe'])
# -
Euler.phase_plane_plot(left_state,right_state,approx_states=states['roe'])
# It is evident that the solution consists of a single right-going shock. The exact solution cannot be seen because it coincides exactly with the Roe solution. The path of the shock in the first plot also cannot be seen since it is plotted under the path of the rightmost Roe solution wave. The two solutions differ only in the wave speeds predicted for the other two waves, but since these waves have zero strength this makes no difference.
# ### Transonic rarefactions and an entropy fix
# Here is an example of a Riemann problem whose solution includes a transonic 2-rarefaction:
# +
left_state = State(Density = 0.1,
Velocity = -2.,
Pressure = 0.1)
right_state = State(Density = 1.,
Velocity = -1.,
Pressure = 1.)
states = compare_solutions(left_state,right_state,solvers=['Exact','Roe'])
# -
# Notice that in the exact solution, the right edge of the rarefaction travels to the right. In the Roe solution, all waves travel to the left. As in the case of the shallow water equations, here too this behavior can lead to unphysical solutions when this approximate solver is used in a numerical discretization. In order to correct this, we can split the single wave into two when a transonic rarefaction is present, in a way similar to what is done in the shallow water equations. We do not go into details here.
# ## HLLE Solver
#
# Recall that an HLL solver uses only two waves with a constant state between them. The Euler equations are our first example for which the number of waves in the true solution is larger than the number of waves in the approximate solution. As one might expect, this leads to noticeable inaccuracy in solutions produced by the solver.
#
# The left-going wave speed is chosen to be the minimum of the Roe speed for the 1-wave and the characterstic speed $\lambda^1$ in the left state $q_\ell$. The right-going wave speed is chosen to be the maximum of the Roe speed for the 3-wave and the characteristic speed $\lambda^3$ in the right state $q_r$. Effectively, this means that
# \begin{align}
# s_1 & = \min(u_l - c_l, \hat{u}-\hat{c}) \\
# s_2 & = \max(u_r + c_r, \hat{u}+\hat{c})
# \end{align}
#
# Recall that once we have chosen these two wave speeds, conservation dictates the value of the intermediate state:
# \begin{align} \label{SWA:hll_middle_state}
# q_m = \frac{f(q_r) - f(q_l) - s_2 q_r + s_1 q_l}{s_1 - s_2}.
# \end{align}
def Euler_hll(q_l, q_r, gamma=1.4):
"""HLL approximate solver for the Euler equations."""
rho_l = q_l[0]
rhou_l = q_l[1]
u_l = rhou_l/rho_l
rho_r = q_r[0]
rhou_r = q_r[1]
u_r = rhou_r/rho_r
E_r = q_r[2]
E_l = q_l[2]
u_hat, c_hat, H_hat = roe_averages(q_l, q_r, gamma)
p_r = (gamma-1.) * (E_r - rho_r*u_r**2/2.)
p_l = (gamma-1.) * (E_l - rho_l*u_l**2/2.)
H_r = (E_r+p_r) / rho_r
H_l = (E_l+p_l) / rho_l
c_r = np.sqrt((gamma-1.)*(H_r-u_r**2/2.))
c_l = np.sqrt((gamma-1.)*(H_l-u_l**2/2.))
s1 = min(u_l-c_l,u_hat-c_hat)
s2 = max(u_r+c_r,u_hat+c_hat)
rho_m = (rhou_r - rhou_l - s2*rho_r + s1*rho_l)/(s1-s2)
rhou_m = (rho_r*u_r**2 - rho_l*u_l**2 + p_r - p_l - s2*rhou_r + s1*rhou_l)/(s1-s2)
E_m = ( u_r*(E_r+p_r) - u_l*(E_l+p_l) -s2*E_r + s1*E_l)/(s1-s2)
q_m = np.array([rho_m, rhou_m, E_m])
states = np.column_stack([q_l,q_m,q_r])
speeds = [s1, s2]
wave_types = ['contact','contact']
def reval(xi):
rho = (xi<s1)*rho_l + (s1<=xi)*(xi<=s2)*rho_m + (s2<xi)*rho_r
mom = (xi<s1)*rhou_l + (s1<=xi)*(xi<=s2)*rhou_m + (s2<xi)*rhou_r
E = (xi<s1)*E_l + (s1<=xi)*(xi<=s2)*E_m + (s2<xi)*E_r
return rho, mom, E
return states, speeds, reval, wave_types
# ### Examples
# +
left_state = State(Density = 3.,
Velocity = 0.,
Pressure = 3.)
right_state = State(Density = 1.,
Velocity = 0.,
Pressure = 1.)
states = compare_solutions(left_state, right_state, solvers=['Exact','HLL'])
# -
Euler.phase_plane_plot(left_state,right_state,approx_states=states['hll'])
# ### Preservation of positivity
# Just as we saw in the case of the shallow water equations, the Roe solver (or any linearized solver) for the Euler equations fails to preserve positivity of the pressure and/or density in some situations. Here is one example.
# +
left_state = State(Density = 1.,
Velocity = -5.,
Pressure = 1.)
right_state = State(Density = 1.,
Velocity = 1.,
Pressure = 1.)
states = compare_solutions(left_state, right_state, solvers=['Exact', 'Roe'])
# -
# As we can see, in this example each Roe solver wave moves much more slowly than the leading edge of the corresponding true rarefaction. In order to maintain conservation, this implies that the middle Roe state must have lower density than the true middle state. This leads to a negative density. Note that the velocity and pressure take huge values in the intermediate state.
#
# The HLL solver, on the other hand, guarantees positivity of the density and pressure. Since the HLL wave speed in the case of a rarefaction is always the speed of the leading edge of the true rarefaction, and since the HLL solution is conservative, the density in a rarefaction will always be at least as great as that of the true solution. This can be seen clearly in the example below.
# +
left_state = State(Density = 1.,
Velocity = -10.,
Pressure = 1.)
right_state = State(Density = 1.,
Velocity = 1.,
Pressure = 1.)
states = compare_solutions(left_state, right_state, solvers=['Exact', 'HLL']);
# -
Euler.phase_plane_plot(left_state,right_state,approx_states=states['hll'])
|
Euler_approximate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + hidden="all" school_cell_uuid="53b7ddb44ee046ab99439ca159abdf5e"
# %config InlineBackend.figure_format = 'png'
# + [markdown] school_cell_uuid="c8e79538701e46cf82e42042707ca962"
# # Scikit-Learn 패키지의 샘플 데이터 - classification용
# + [markdown] school_cell_uuid="b34ae27707c944ebaf99b84c56338e11"
# ## Iris Dataset
# + [markdown] school_cell_uuid="a6bf4c8f0ca64d60bb26bd664f8e5b74" slideshow={"slide_type": "slide"}
# #### `load_iris()`
# * https://en.wikipedia.org/wiki/Iris_flower_data_set
# * R.A Fisher의 붓꽃 분류 연구
# * 관찰 자료
# * 꽃받침 길이(Sepal Length)
# * 꽃받침 폭(Sepal Width)
# * 꽃잎 길이(Petal Length)
# * 꽃잎 폭(Petal Width)
# * 종
# * setosa
# * versicolor
# * virginica
# + school_cell_uuid="f9b43e0f7557436f802b49725128d02c" slideshow={"slide_type": "slide"}
from sklearn.datasets import load_iris
iris = load_iris()
print(iris.DESCR)
# + school_cell_uuid="4e7f0b2c60914938a0eae3a1d0ed9b6b" slideshow={"slide_type": "slide"}
df = pd.DataFrame(iris.data, columns=iris.feature_names)
sy = pd.Series(iris.target, dtype="category")
sy = sy.cat.rename_categories(iris.target_names)
df['species'] = sy
df.tail()
# + school_cell_uuid="7a612dd7b3484f85bdbf919eae63a3b0"
sns.pairplot(df, hue="species")
plt.show()
# + [markdown] school_cell_uuid="be864bccf0d444d7aa239ee712ed424c"
# ## 뉴스 그룹 텍스트
# + [markdown] school_cell_uuid="2279aeb8cd3343cda5bc9e0dd21ed03e"
# #### `fetch_20newsgroups()`: 20 News Groups text
# + school_cell_uuid="c4c606873e174bd6810ad28e7ac7bcae"
from sklearn.datasets import fetch_20newsgroups
newsgroups = fetch_20newsgroups(subset='all')
print(newsgroups.description)
print(newsgroups.keys())
# + school_cell_uuid="0c6063e4d1f542f0aa1fb0317896be95"
from pprint import pprint
pprint(list(newsgroups.target_names))
# + school_cell_uuid="a9230e0ecdbe431ba2d2fcfe513bfe47"
print(newsgroups.data[1])
print("=" * 80)
print(newsgroups.target_names[newsgroups.target[1]])
# + [markdown] school_cell_uuid="1d6be724fae44e0990347cda62acd4b8"
# ## Olivetti faces
# + [markdown] school_cell_uuid="38b1f59d2adc48318bd6cc64686c8508"
# #### `fetch_olivetti_faces()`
#
# * 얼굴 인식 이미지
# + school_cell_uuid="a0c9849ab798438395a77b59391aa149"
from sklearn.datasets import fetch_olivetti_faces
olivetti = fetch_olivetti_faces()
print(olivetti.DESCR)
print(olivetti.keys())
# + school_cell_uuid="567e6ecdbb544f79a50278b52ecd3efc"
N=2; M=5;
fig = plt.figure(figsize=(8,5))
plt.subplots_adjust(top=1, bottom=0, hspace=0, wspace=0.05)
klist = np.random.choice(range(len(olivetti.data)), N * M)
for i in range(N):
for j in range(M):
k = klist[i*M+j]
ax = fig.add_subplot(N, M, i*M+j+1)
ax.imshow(olivetti.images[k], cmap=plt.cm.bone);
ax.grid(False)
ax.xaxis.set_ticks([])
ax.yaxis.set_ticks([])
plt.title(olivetti.target[k])
plt.tight_layout()
plt.show()
# + [markdown] school_cell_uuid="72bacc171c7c456c968ea1dde1f51c2f"
# ## Labeled Faces in the Wild (LFW)
# + [markdown] school_cell_uuid="acdf16122d0446339db545ca3138f6d0"
# #### `fetch_lfw_people()`
#
# * 유명인 얼굴 이미지
#
# * Parameters
#
# * funneled : boolean, optional, default: True
# * Download and use the funneled variant of the dataset.
# * resize : float, optional, default 0.5
# * Ratio used to resize the each face picture.
# * min_faces_per_person : int, optional, default None
# * The extracted dataset will only retain pictures of people that have at least min_faces_per_person different pictures.
# * color : boolean, optional, default False
# * Keep the 3 RGB channels instead of averaging them to a single gray level channel. If color is True the shape of the data has one more dimension than than the shape with color = False.
#
# + school_cell_uuid="fdbf54d8ba7548e6a15d8399afd5ad32"
from sklearn.datasets import fetch_lfw_people
lfw_people = fetch_lfw_people(min_faces_per_person=70, resize=0.4)
print(lfw_people.DESCR)
print(lfw_people.keys())
# + school_cell_uuid="4e6c53037fe145639f447f6c85e0ef00"
N=2; M=5;
fig = plt.figure(figsize=(8,5))
plt.subplots_adjust(top=1, bottom=0, hspace=0.1, wspace=0.05)
klist = np.random.choice(range(len(lfw_people.data)), N * M)
for i in range(N):
for j in range(M):
k = klist[i*M+j]
ax = fig.add_subplot(N, M, i*M+j+1)
ax.imshow(lfw_people.images[k], cmap=plt.cm.bone);
ax.grid(False)
ax.xaxis.set_ticks([])
ax.yaxis.set_ticks([])
plt.title(lfw_people.target_names[lfw_people.target[k]])
plt.tight_layout()
plt.show()
# + [markdown] school_cell_uuid="81f7380ce53b49978edd454954fc9658"
# #### `fetch_lfw_pairs()`
#
# * 얼굴 이미지 Pair
# * 동일 인물일 수도 있고 아닐 수도 있음
# + school_cell_uuid="a75f65f58e7e4ae3aed0e5bd75f4ab05"
from sklearn.datasets import fetch_lfw_pairs
lfw_pairs = fetch_lfw_pairs(resize=0.4)
print(lfw_pairs.DESCR)
print(lfw_pairs.keys())
# + school_cell_uuid="9ca7265bd367415faff6105924ee05e4"
N=2; M=5;
fig = plt.figure(figsize=(8,5))
plt.subplots_adjust(top=1, bottom=0, hspace=0.01, wspace=0.05)
klist = np.random.choice(range(len(lfw_pairs.data)), M)
for j in range(M):
k = klist[j]
ax1 = fig.add_subplot(N, M, j+1)
ax1.imshow(lfw_pairs.pairs [k][0], cmap=plt.cm.bone);
ax1.grid(False)
ax1.xaxis.set_ticks([])
ax1.yaxis.set_ticks([])
plt.title(lfw_pairs.target_names[lfw_pairs.target[k]])
ax2 = fig.add_subplot(N, M, j+1 + M)
ax2.imshow(lfw_pairs.pairs [k][1], cmap=plt.cm.bone);
ax2.grid(False)
ax2.xaxis.set_ticks([])
ax2.yaxis.set_ticks([])
plt.tight_layout()
plt.show()
# + [markdown] school_cell_uuid="4c0b604d748d458481a93538cadfbbc8"
# ## Digits Handwriting Image
# + [markdown] school_cell_uuid="e1f3cea40cf44734a760bc3dc0b70cd9"
# #### `load_digits()`
#
# * 숫자 필기 이미지
#
# + school_cell_uuid="c816f3101b4c4dbdb3d23a0ab0526bc1"
from sklearn.datasets import load_digits
digits = load_digits()
print(digits.DESCR)
print(digits.keys())
# + school_cell_uuid="fb3aa96223204271a15ed154919c3020"
N=2; M=5;
fig = plt.figure(figsize=(10,5))
plt.subplots_adjust(top=1, bottom=0, hspace=0, wspace=0.05)
for i in range(N):
for j in range(M):
k = i*M+j
ax = fig.add_subplot(N, M, k+1)
ax.imshow(digits.images[k], cmap=plt.cm.bone, interpolation="none");
ax.grid(False)
ax.xaxis.set_ticks([])
ax.yaxis.set_ticks([])
plt.title(digits.target_names[k])
plt.tight_layout()
plt.show()
# + [markdown] school_cell_uuid="fb19417bfbd24919bfd226ab77825e09"
# ## mldata.org repository
# + [markdown] school_cell_uuid="83abd689c1334e858b39a83b09ac3f6c"
# #### `fetch_mldata()`
#
# * http://mldata.org
# * public repository for machine learning data, supported by the PASCAL network
# * 홈페이지에서 data name 을 검색 후 key로 이용
# + [markdown] school_cell_uuid="5cad627db73b4288b7258dda5352ed3c" slideshow={"slide_type": "slide"}
# ##### MNIST 숫자 필기인식 자료
# * https://en.wikipedia.org/wiki/MNIST_database
# * Mixed National Institute of Standards and Technology (MNIST) database
# * 0-9 필기 숫자 이미지
# * 28x28 pixel bounding box
# * anti-aliased, grayscale levels
# * 60,000 training images and 10,000 testing images
#
# + school_cell_uuid="f8790ce123074ac3b099adc757f1c1d1" slideshow={"slide_type": "slide"}
from sklearn.datasets.mldata import fetch_mldata
mnist = fetch_mldata('MNIST original')
mnist.keys()
# + school_cell_uuid="f2c07ec8d8174b25a95d9f1edfd50fed" slideshow={"slide_type": "slide"}
N=2; M=5;
fig = plt.figure(figsize=(8,5))
plt.subplots_adjust(top=1, bottom=0, hspace=0, wspace=0.05)
klist = np.random.choice(range(len(mnist.data)), N * M)
for i in range(N):
for j in range(M):
k = klist[i*M+j]
ax = fig.add_subplot(N, M, i*M+j+1)
ax.imshow(mnist.data[k].reshape(28, 28), cmap=plt.cm.bone, interpolation="nearest");
ax.grid(False)
ax.xaxis.set_ticks([])
ax.yaxis.set_ticks([])
plt.title(mnist.target[k])
plt.tight_layout()
plt.show()
|
13. Scikit-Learn, Statsmodel/04. Scikit-Learn 패키지의 샘플 데이터 - classification용.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend", "outputPrepend"]
import numpy as np
import matplotlib.pyplot as plt
import math
def cost(y,y1):
ans=0
for i in range(len(y)):
ans+=((y[i]*np.math.log(y1[i],np.exp(1)))+(1-y[i])*np.math.log((1-y1[i]),np.exp(1)))
return -1*ans
def normalise_scale(y):
l=[]
for i in range(len(y)):
l.append(((y[i]-min(y))/(max(y)-min(y)))*(1-0)+0)
return l
def sigmoid(z):
return (1/(1+np.exp(-z)))
def calculate_y(x,w):
ans=[]
for j in range(len(x)):
y=0
y+=w[0]
for i in range(1,len(w)):
y+=w[i]*x[j][i-1]
ans.append(y)
return sigmoid(np.asarray(ans))
def gradient_descent(x,y,learning_rate,stopping):
temp=0
w=np.random.random(len(x[0])+1)*np.random.randint(1,5,1)
y1=calculate_y(x,w)
print(y,y1)
while(abs(cost(y,y1)-temp)>stopping):
temp=cost(y,y1)
w[0]-=learning_rate*-1*np.sum(y-y1)
for i in range(1,len(w)):
w[i]-=learning_rate*-1*np.sum((y-y1)*(x[:,i-1]))
print(w,cost(y,y1))
y1=calculate_y(x,w)
return w
def logistic_regressor(x,y,learning_rate,stopping):
w=gradient_descent(x,y,learning_rate,stopping)
print(w)
y1=calculate_y(x,w)
return y1,w
x1=np.zeros((4,2))
x1[1,1]+=1
x1[2,0]+=1
x1[3,0]+=1
x1[3,1]+=1
y=np.zeros(4)
for i in range(len(x1)):
if(x1[i,0]==1 or x1[i,1]==1):
y[i]=1
else:
y[i]=0
#for i in range(len(x1)):
# x1[i]=normalise_scale(x1[i])
#y=normalise_scale(y)
print(x1,y)
y1,w=logistic_regressor(x1,y,0.1,0.00001)
# +
for i in range(len(x1)):
if(y[i]==1):
plt.scatter(x1[i,0],x1[i,1],c='r')
else:
plt.scatter(x1[i,0],x1[i,1],c='b')
print(w,y1)
x=[0,1]
z=[]
for i in range(len(x)):
z.append((0.5-w[0]-(x[i]*w[1]))/w[2])
plt.plot(x,z)
# -
|
Logistic Regression/Logistic Regression using gradient descent.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="VYNA79KmgvbY" colab_type="text"
# Copyright 2018 The Dopamine Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# + [markdown] id="emUEZEvldNyX" colab_type="text"
# # Dopamine: How to create and train a custom agent
#
# This colab demonstrates how to create a variant of a provided agent (Example 1) and how to create a new agent from
# scratch (Example 2).
#
# Run all the cells below in order.
# + id="Ckq6WG-seC7F" colab_type="code" cellView="form" colab={}
# @title Install necessary packages.
# !pip install --upgrade --no-cache-dir dopamine-rl
# !pip install cmake
# !pip install atari_py
# !pip install gin-config
# + id="WzwZoRKxdFov" colab_type="code" cellView="form" colab={}
# @title Necessary imports and globals.
import numpy as np
import os
from dopamine.agents.dqn import dqn_agent
from dopamine.discrete_domains import run_experiment
from dopamine.colab import utils as colab_utils
from absl import flags
import gin.tf
BASE_PATH = '/tmp/colab_dope_run' # @param
GAME = 'Asterix' # @param
# + id="EFY3tTITHugq" colab_type="code" cellView="form" colab={}
# @title Load baseline data
# !gsutil -q -m cp -R gs://download-dopamine-rl/preprocessed-benchmarks/* /content/
experimental_data = colab_utils.load_baselines('/content')
# + [markdown] id="bidurBV0djGi" colab_type="text"
# ## Example 1: Train a modified version of DQN
# Asterix is one of the standard agents provided with Dopamine.
# The purpose of this example is to demonstrate how one can modify an existing agent. The modification
# we are doing here (choosing actions randomly) is for illustrative purposes: it will clearly perform very
# poorly.
# + id="PUBRSmX6dfa3" colab_type="code" colab={}
# @title Create an agent based on DQN, but choosing actions randomly.
LOG_PATH = os.path.join(BASE_PATH, 'random_dqn', GAME)
class MyRandomDQNAgent(dqn_agent.DQNAgent):
def __init__(self, sess, num_actions):
"""This maintains all the DQN default argument values."""
super(MyRandomDQNAgent, self).__init__(sess, num_actions)
def step(self, reward, observation):
"""Calls the step function of the parent class, but returns a random action.
"""
_ = super(MyRandomDQNAgent, self).step(reward, observation)
return np.random.randint(self.num_actions)
def create_random_dqn_agent(sess, environment, summary_writer=None):
"""The Runner class will expect a function of this type to create an agent."""
return MyRandomDQNAgent(sess, num_actions=environment.action_space.n)
random_dqn_config = """
import dopamine.discrete_domains.atari_lib
import dopamine.discrete_domains.run_experiment
atari_lib.create_atari_environment.game_name = '{}'
atari_lib.create_atari_environment.sticky_actions = True
run_experiment.Runner.num_iterations = 200
run_experiment.Runner.training_steps = 10
run_experiment.Runner.max_steps_per_episode = 100
""".format(GAME)
gin.parse_config(random_dqn_config, skip_unknown=False)
# Create the runner class with this agent. We use very small numbers of steps
# to terminate quickly, as this is mostly meant for demonstrating how one can
# use the framework.
random_dqn_runner = run_experiment.TrainRunner(LOG_PATH, create_random_dqn_agent)
# + id="WuWFGwGHfkFp" colab_type="code" colab={}
# @title Train MyRandomDQNAgent.
print('Will train agent, please be patient, may be a while...')
random_dqn_runner.run_experiment()
print('Done training!')
# + id="IknanILXX4Zz" colab_type="code" colab={}
# @title Load the training logs.
random_dqn_data = colab_utils.read_experiment(LOG_PATH, verbose=True)
random_dqn_data['agent'] = 'MyRandomDQN'
random_dqn_data['run_number'] = 1
experimental_data[GAME] = experimental_data[GAME].merge(random_dqn_data,
how='outer')
# + id="mSOVFUKN-kea" colab_type="code" outputId="c7053a43-9f59-4817-ee0a-b3c074b509b1" colab={"base_uri": "https://localhost:8080/", "height": 512}
# @title Plot training results.
import seaborn as sns
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(16,8))
sns.tsplot(data=experimental_data[GAME], time='iteration', unit='run_number',
condition='agent', value='train_episode_returns', ax=ax)
plt.title(GAME)
plt.show()
# + [markdown] id="8T0yfWPw-7QZ" colab_type="text"
# ## Example 2: Train an agent built from scratch.
# The purpose of this example is to demonstrate how one can create an agent from scratch. The agent
# created here is meant to demonstrate the bare minimum functionality that is expected from agents. It is
# selecting actions in a very suboptimal way, so it will clearly do poorly.
# + id="1kgV__YU-_ET" colab_type="code" colab={}
# @title Create a completely new agent from scratch.
LOG_PATH = os.path.join(BASE_PATH, 'sticky_agent', GAME)
class StickyAgent(object):
"""This agent randomly selects an action and sticks to it. It will change
actions with probability switch_prob."""
def __init__(self, sess, num_actions, switch_prob=0.1):
self._sess = sess
self._num_actions = num_actions
self._switch_prob = switch_prob
self._last_action = np.random.randint(num_actions)
self.eval_mode = False
def _choose_action(self):
if np.random.random() <= self._switch_prob:
self._last_action = np.random.randint(self._num_actions)
return self._last_action
def bundle_and_checkpoint(self, unused_checkpoint_dir, unused_iteration):
pass
def unbundle(self, unused_checkpoint_dir, unused_checkpoint_version,
unused_data):
pass
def begin_episode(self, unused_observation):
return self._choose_action()
def end_episode(self, unused_reward):
pass
def step(self, reward, observation):
return self._choose_action()
def create_sticky_agent(sess, environment, summary_writer=None):
"""The Runner class will expect a function of this type to create an agent."""
return StickyAgent(sess, num_actions=environment.action_space.n,
switch_prob=0.2)
sticky_config = """
import dopamine.discrete_domains.atari_lib
import dopamine.discrete_domains.run_experiment
atari_lib.create_atari_environment.game_name = '{}'
atari_lib.create_atari_environment.sticky_actions = True
run_experiment.Runner.num_iterations = 200
run_experiment.Runner.training_steps = 10
run_experiment.Runner.max_steps_per_episode = 100
""".format(GAME)
gin.parse_config(sticky_config, skip_unknown=False)
# Create the runner class with this agent. We use very small numbers of steps
# to terminate quickly, as this is mostly meant for demonstrating how one can
# use the framework.
sticky_runner = run_experiment.TrainRunner(LOG_PATH, create_sticky_agent)
# + id="gQt3t_IS_Gku" colab_type="code" colab={}
# @title Train StickyAgent.
print('Will train sticky agent, please be patient, may be a while...')
sticky_runner.run_experiment()
print('Done training!')
# + id="oom0wB0A_Qb8" colab_type="code" colab={}
# @title Load the training logs.
sticky_data = colab_utils.read_experiment(log_path=LOG_PATH, verbose=True)
sticky_data['agent'] = 'StickyAgent'
sticky_data['run_number'] = 1
experimental_data[GAME] = experimental_data[GAME].merge(sticky_data,
how='outer')
# + id="DqsagPbb_Xjm" colab_type="code" outputId="1d263334-e476-4f76-88df-28d0b6a271ae" colab={"base_uri": "https://localhost:8080/", "height": 512}
# @title Plot training results.
import seaborn as sns
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(16,8))
sns.tsplot(data=experimental_data[GAME], time='iteration', unit='run_number',
condition='agent', value='train_episode_returns', ax=ax)
plt.title(GAME)
plt.show()
|
dopamine/colab/agents.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Misc code
# <hr>
# +
# create dataframe based on PROPANE DF
df = propane_df.drop(['invoice','cost per gal','cost','Environmental Compliance Fee',], axis = 1).copy()
df.head(1)
# Creates an Initial temperature reading
# based on avg temperature of the 30 days before 1st propane data entered
initial_temp_reading = round( temp_df.loc[ # iloc statement
(df['date'][0] - pd.Timedelta(weeks=4)) : # start date
(df['date'][0] - pd.Timedelta(days=1)) # end date
]['T2M_RANGE'].mean(),2 )
initial_temp_reading
#
avg_temp_list = [initial_temp_reading]
n_days = [12]
n_date = 12
# var for loop
# number of times for loop
end_of_range = len( df['date'] ) -1
#
for n in range(0, end_of_range ):
if n_date >= 10:
break
# assigns start date
date_start = df['date'][n]
# assigns end date
date_end = df['date'][n + 1]
# n of days between [date_start to date_end]
n_days.append( int(str( date_end - date_start ).split(' ')[0]) )
# slices datetime and Calcualtes MEAN for 'ts'
avg_temp_list.append( round( temp_df.loc[ date_start: (date_end - pd.Timedelta(days=1)) ]['ts'].mean(),2 ) )
# print TEMP for debuging
temp = temp_df.loc[ date_start: (date_end - pd.Timedelta(days=1)) ]['ts'].mean()
print(f'number {n}, date {date_start} to {date_end} Temp Mean {temp}.')
print()
# Examine avg_temp_list list
np.array(avg_temp_list)
# join .mean columns
df['avg_temp'] = np.array(temp_average)
df['days'] = np.array(n_days)
df
# -
|
code/misc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dates in timeseries models
# + jupyter={"outputs_hidden": false}
import pandas as pd
import matplotlib.pyplot as plt
import statsmodels.api as sm
from statsmodels.tsa.ar_model import AutoReg, ar_select_order
plt.rc("figure", figsize=(16,8))
plt.rc("font", size=14)
# -
# ## Getting started
# + jupyter={"outputs_hidden": false}
data = sm.datasets.sunspots.load()
# -
# Right now an annual date series must be datetimes at the end of the year.
# + jupyter={"outputs_hidden": false}
from datetime import datetime
dates = pd.date_range('1700-1-1', periods=len(data.endog), freq="A-DEC")
# -
# ## Using Pandas
#
# Make a pandas TimeSeries or DataFrame
# + jupyter={"outputs_hidden": false}
data.endog.index = dates
endog = data.endog
endog
# -
# Instantiate the model
# + jupyter={"outputs_hidden": false}
selection_res = ar_select_order(endog, 9, old_names=False, seasonal=True, period=11)
pandas_ar_res = selection_res.model.fit()
# -
# Out-of-sample prediction
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
pred = pandas_ar_res.predict(start='2005', end='2027')
print(pred)
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
fig = pandas_ar_res.plot_predict(start='2005', end='2027')
|
examples/notebooks/tsa_dates.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#import BeautifulSoup, Pandas, Requests/Splinter
# -
import requests as req
import time
import os
import pandas as pd
# import pymongo
from bs4 import BeautifulSoup as bs
from splinter import Browser
from selenium import webdriver
from flask import Flask, render_template, redirect
# from flask_pymongo import PyMongo
# Point out chromedriver directory
executable_path = {'executable_path': 'chromedriver.exe'}
browser = Browser('chrome', **executable_path, headless = False)
# Get latest news from NASA website
news_url = 'https://mars.nasa.gov/news/'
browser.visit(news_url)
html = browser.html
# Parse HTML with Beautiful Soup
soup = bs(html, 'html.parser')
# Obtain the latest news title and paragraph text
article = soup.find('div', class_='list_text')
news_title = soup.find("div", class_="content_title").text
news_p = soup.find("div", class_="description").text
print(news_title)
print(news_p)
#JPL Mars Space Images - Featured Image
#visit url and extract JPL Featured space image
images_url = "https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars"
browser.visit(images_url)
html = browser.html
images_soup = bs(html, 'html.parser')
#Retrieve full size image link
full_size_image_path = images_soup.find_all('img')[3]["src"]
full_size_image_url = f'http://www.jpl.nasa.gov{full_size_image_path}'
print(full_size_image_url)
# +
# Mars weather
weather_url = 'https://twitter.com/marswxreport?lang=en'
browser.visit(weather_url)
time.sleep(5)
weather_html = browser.html
weather_soup = bs(weather_html, 'html.parser')
# +
mars_weather = weather_soup.find('section', attrs={"aria-labelledby":"accessible-list-0"})
mars_weather = mars_weather.find_all('span')
mars_w = []
for x in mars_weather:
if len(x.get_text())>100:
mars_w.append(x.get_text())
mars_w
# -
# Mars facts page
url = "https://space-facts.com/mars/"
# Scrape the table of mars facts using pandas
mars_facts = pd.read_html(url)
mars_facts
# extract description and value from the table and put the information in as data frame
mars_facts_df = tables[2]
mars_facts_df.columns = ["Description", "Value"]
mars_facts_df
# Use Pandas to convert the data to a HTML table string.
mars_html_table = mars_facts_df.to_html()
print(mars_html_table)
### Mars Hemispheres
hemispheres_url = 'https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
browser.visit(hemispheres_url)
hemispheres_html = browser.html
hemispheres_soup = bs(hemispheres_html, 'html.parser')
# +
# Mars hemispheres products data
all_mars_hemispheres = hemispheres_soup.find('div', class_='collapsible results')
mars_hemispheres = all_mars_hemispheres.find_all('div', class_='item')
hemisphere_image_urls = []
url = 'https://astrogeology.usgs.gov'
# Iterate through each hemisphere data
for i in mars_hemispheres:
# Collect Title information
hemisphere = i.find('div', class_="description")
title = hemisphere.h3.text
# Collect image link by browsing to hemisphere page
hemisphere_link = hemisphere.a["href"]
browser.visit(url + hemisphere_link)
image_html = browser.html
image_soup = bs(image_html, 'html.parser')
image_link = image_soup.find('div', class_='downloads')
image_url = image_link.find('li').a['href']
# Save both the image url string for the full resolution hemisphere image, and the Hemisphere title containing the hemisphere name.
image_dict = {}
image_dict['title'] = title
image_dict['img_url'] = image_url
# Append the dictionary with the image url string and the hemisphere title to a list. This list will contain one dictionary for each hemisphere.
hemisphere_image_urls.append(image_dict)
print(hemisphere_image_urls)
# -
Mars_information ={
"news_title": news_title,
"news_p": news_p,
"featured_image_url": full_size_image_url,
"mars_weather": Mars_weather,
"fact_table": str(mars_html_table),
"hemisphere_images": hemisphere_image_urls
}
print(Mars_information)
|
.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from collections import namedtuple
import datetime
import numpy as np
import spacepy.plot as spp
import spacepy.toolbox as tb
import pymc as mc
import tqdm
# +
x = mc.Uniform('x', 0, 1)
y = mc.Uniform('y', 0, 1)
# @mc.deterministic(plot=True)
# def pi(x=x, y=y):
# inside = (x**2 + y**2) <= 1
# outside = (x**2 + y**2) > 1
# return 4*inside/outside
model = mc.MCMC((x, y))
# -
model.sample(50000, burn=0, burn_till_tuned=False)
mc.Matplot.plot(model)
# +
inside = np.sum(x.trace()**2 + y.trace()**2 < 1)
total = len(x.trace())
pi = 4*inside/total
print('PI: 4*{0}/{1}={2}'.format(inside, total, pi))
# -
# # Wrap all this is a loop to see how the $\sqrt{N}$ decrease of error
# +
dat = namedtuple('Pi', 'niter error')
data = []
for i in tqdm.tqdm(tb.logspace(100, 1e7, 30)[::-1]):
x = mc.Uniform('x', 0, 1)
y = mc.Uniform('y', 0, 1)
model = mc.MCMC((x, y))
model.sample(i, burn=10, progress_bar=False)
inside = np.sum(x.trace()**2 + y.trace()**2 < 1)
total = len(x.trace())
pi = 4*inside/total
data.append(dat(i, np.abs((pi-np.pi)/np.pi)))
# print(i, pi, np.abs((pi-np.pi)/np.pi))
# -
its = [v.niter for v in data]
err = [v.error for v in data]
spp.plt.loglog(its, err, '.-')
spp.plt.plot(its, 1/np.sqrt(its)*0.4)
spp.plt.xlabel('Number of draws')
spp.plt.ylabel('Percent error')
|
Integration/Calculate Pi.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %load_ext autoreload
# %autoreload 2
# -
frames_df = pd.read_csv("/home/romet/projects/ut/wp4/nvidia-e2e/data_extract/temp/2021-05-28-15-17-19_e2e_sulaoja_20_30/frames.csv")
# +
N_WAYPOINTS = 5
for i in np.arange(1, N_WAYPOINTS+1):
print(i)
vehicle_x = frames_df["position_x"]
vehicle_y = frames_df["position_y"]
wp_global_x = frames_df["position_x"].shift(-i)
wp_global_y = frames_df["position_y"].shift(-i)
yaw = frames_df["yaw"]
wp_local_x = (wp_global_x - vehicle_x)*np.cos(yaw) + (wp_global_y - vehicle_y)*np.sin(yaw)
wp_local_y = -(wp_global_x - vehicle_x)*np.sin(yaw) + (wp_global_y - vehicle_y)*np.cos(yaw)
frames_df[f"x_{i}_offset"] = wp_local_x
frames_df[f"y_{i}_offset"] = wp_local_y
# -
frames_df.head(10)
vehicle_pose = np.array([t0["position_x"], t0["position_y"], t0["yaw"]])
vehicle_pose
# +
from math import sin, cos
from numpy.linalg import inv
transform = np.array([[cos(yaw), -sin(yaw)], [sin(yaw), cos(yaw)]])
inv_transform = inv(transform)
inv_transform
# -
waypoints_global = np.array([[t0["position_x_1"], t0["position_y_1"]],
[t0["position_x_2"], t0["position_y_2"]],
[t0["position_x_3"], t0["position_y_3"]]])
waypoints_global.T
def global_to_local(vehicel_pose, waypoint_global):
x_global = (waypoint_global[0] - vehicel_pose[0])*cos(vehicle_pose[2]) + (waypoint_global[1] - vehicle_pose[1])*sin(vehicle_pose[2])
y_global = -(waypoint_global[0] - vehicel_pose[0])*sin(vehicle_pose[2]) + (waypoint_global[1] - vehicle_pose[1])*cos(vehicle_pose[2])
return (x_global, y_global)
global_to_local(vehicle_pose, [t0["position_x_1"], t0["position_y_1"]])
global_to_local(vehicle_pose, [t0["position_x_2"], t0["position_y_2"]])
global_to_local(vehicle_pose, [t0["position_x_3"], t0["position_y_3"]])
np.matmul(inv_transform, (waypoints_global-vehicle_pose[0:2]).T).T
|
notebooks/checkpoints.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Been trying to download SQL server management studio but the process has been a mess. So I'm gonna have to settle for SQLite3. Doesn't seem like a problem right now, I don't think I'll be dealing with too much data just yet. Alright, this is just a basic test run.
import sqlite3, pandas as pd
# Blah blah blah, import packages
con=sqlite3.connect('C:\\Users\\Jonathan\\Desktop\\database\\first.db')
cursor = con.cursor()
# Awesome, so I was able to connect without any complains. Alright, here's what happened:
# * Tried whole path. Didn't work. Tried again with double backslahes and it suddenly works. Thanks stack overflow.
# * I found this to be really helpful. https://www.youtube.com/watch?v=JVAWKVpdb4Y If you want to dive in, this got me hitting the ground the fastest. Props to this guy.
# +
my_query = """
SELECT *
FROM BostonHousing
LIMIT 5
"""
for row in cursor.execute(my_query):
print row
# -
# Not really liking this format here...
df = pd.read_sql(my_query,con)
df.head()
df.columns = df.ix[0].as_matrix() #df.columns.values is often a copy. set it on df.columns to change column names
df = df.reindex(df.index.drop(0)).reset_index()
df = df[df.columns.values[1:]]
df.head()
# That's much better. Turns out I didn't check off a box when importing the csv into SQLite3.
|
Scratch_Pad/SQLite3 first steps.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:adh]
# language: python
# name: conda-env-adh-py
# ---
import os
from bs4 import BeautifulSoup
import pandas as pd
import itertools
## Read the XML files
def extract_from_xml(search_roots, filenames):
metadata = []
matched_words = []
for filename in filenames:
with open(filename) as fn:
xml_data = BeautifulSoup(fn, 'xml')
meta_dict = {meta['name']: meta.text.strip() for meta in xml_data.metadata.find_all('meta')}
#meta_dict['Bookname'] = dirname
#meta_dict['Filename'] = filename
metadata.append(meta_dict)
# loop over words and match with the searched words
# To do: do not include roots that are not within the search set
for word in xml_data.morphology_analysis.find_all('word'):
roots = set([a.get('root', '') for a in word.find_all('analysis')])
if not set(search_roots).isdisjoint(roots):
matched_words.append((filename, word.attrs, [a.attrs for a in word.find_all('analysis')]))
# # Put all results in a dataframe
df_total = pd.DataFrame()
for filename, word_dict, analyses in matched_words:
df_analyses = pd.DataFrame(analyses)
df_analyses['Filename'] = os.path.basename(filename)
for att in word_dict:
df_analyses[att] = word_dict[att]
df_total = df_total.append(df_analyses)
return metadata, df_total
## Read the csv files
def extract_from_csv(search_roots, filenames):
df_total = pd.DataFrame()
for i in range(len(filenames)):
if i%1000==0:
print(i)
filename = filenames[i]
df_sub = pd.read_csv(filename, index_col=0)
df_sub = df_sub[df_sub.root.isin(search_roots)]
df_total = df_total.append(df_sub)
return df_total
# +
from lxml import etree
from tqdm import tqdm
import pandas as pd
def analyzer_xml2df2(fname, filter_roots=None, filter_stems=None, filter_words=None):
result = []
# Extract the words
context = etree.iterparse(fname, events=('end', ), tag=('word'))
for event, elem in tqdm(context):
word = elem.attrib['value']
if word != '' and (filter_words is None or word in filter_words):
roots = []
stems = []
for a in elem.getchildren():
if a.tag == 'analysis':
try:
roots.append(a.attrib['root'])
except:
pass
try:
stems.append(a.attrib['stem'])
except:
pass
roots = list(set(roots))
stems = list(set(stems))
if len(roots) == 0:
roots.append('NOANALYSIS')
if len(stems) == 0:
stems.append('NOANALYSIS')
if filter_roots is None or not set(filter_roots).isdisjoint(set(roots)):
if filter_stems is None or not set(filter_stems).isdisjoint(set(stems)):
result.append({'word': elem.attrib['value'], 'proposed_root': '\\'.join(roots), 'proposed_stem': '\\'.join(stems)})
# make iteration over context fast and consume less memory
#https://www.ibm.com/developerworks/xml/library/x-hiperfparse
elem.clear()
while elem.getprevious() is not None:
del elem.getparent()[0]
return pd.DataFrame(result)
# -
# ## Free reasoning
# +
stems_dict = {
u'اجتهاد': 'free',
u'مجتهد': 'free',
u'مقلد': 'following',
u'تقليد': 'following'
}
stems_list = stems_dict.keys()
# +
fpath = '/media/sf_VBox_Shared/Arabic/Fiqh/2018-09-18-Al-Khalil/'
df = pd.DataFrame()
for fname in os.listdir(fpath):
print(fname)
df_sub = analyzer_xml2df2(os.path.join(fpath, fname), filter_stems=stems_list)
df_sub['fname'] = fname
df = df.append(df_sub)
# -
fname_out = '/media/sf_VBox_Shared/Arabic/Analyses/fiqh-reasoning-raw.csv'
df.to_csv(fname_out, index=False)
# +
# Retrieve the stem we were originally interested in
def get_original_stem(stems):
l = set(stems.split('\\'))
overlap = l.intersection(set(stems_list))
if len(overlap)>0:
return list(overlap)[0]
else:
return None
df['stem'] = df.proposed_stem.map(get_original_stem)
# -
df['category'] = df['stem'].map(lambda s: stems_dict[s])
df['BookURI'] = df['fname'].map(lambda s: s.replace('.xml', ''))
counts_category = df.groupby(['BookURI', 'category']).size().unstack().fillna(0)
counts_category.head()
# merge with meta data:
metadata_fname = '/media/sf_VBox_Shared/Arabic/fiqh_corpus/Meta/Metadata_Fiqh.csv'
metadata = pd.read_csv(metadata_fname, index_col=1)
metadata['century_n'] = metadata.Century.str.extract('([0-9]*)')
metadata.columns
metadata_columns = ['AuthorAKA', 'AuthorBORNH', 'AuthorBORNC', 'AuthorDIEDH', 'AuthorDIEDC', 'Century', 'School', 'Geographical_area', 'Number_of_tokens', 'century_n']
metadata = metadata[metadata_columns]
counts_category[metadata_columns] = metadata[metadata_columns]
counts_long = df.groupby(['BookURI', 'stem', 'category']).size()
counts_long.head()
counts_long_merged = pd.merge(pd.DataFrame(counts_long, columns=['count']).reset_index(),
metadata.reset_index(), left_on='BookURI', right_on='BookURI')
fname_out = '/media/sf_VBox_Shared/Arabic/Analyses/fiqh-reasoning-aggregated.csv'
counts_category.to_csv(fname_out)
fname_out = '/media/sf_VBox_Shared/Arabic/Analyses/fiqh-reasoning-aggregated-long.csv'
counts_long_merged.to_csv(fname_out, index=False)
# # Extract senses
senses_roots = set('''سمع
بصر
لمس
شمم
ذوق'''.split('\n'))
## from XML
filepath = '/media/sf_VBox_Shared/Arabic/indices/20180424/merged/'
xml_file_names = itertools.chain.from_iterable([[os.path.join(d, f) for f in fnames] for d, dnames, fnames in os.walk(filepath)])
metadata, matched_words = extract_from_xml(senses_roots, list(xml_file_names)[:20])
# from CSV
filepath = '/media/sf_VBox_Shared/Arabic/Fiqh/Fiqh-Alkhalil-csv/csv'
csv_file_names = [os.path.join(filepath, fn) for fn in os.listdir(filepath)]
df_total = extract_from_csv(senses_roots, list(csv_file_names))
df_total.shape
df_total['root'].value_counts()
# +
senses_dict = {
u'بصر': 'see',
u'سمع': 'hear',
u'لمس': 'touch',
u'شمم': 'smell',
u'ذوق': 'taste'
}
df_total['sense'] = [senses_dict[s] for s in df_total['root']]
# -
# ## Merged with metadata
# +
metadata_fields = ['BookURI', 'Century', 'AuthorNAME', 'AuthorGeographicalArea', 'AuthorBORNH', 'AuthorBORNC', 'AuthorDIEDH', 'AuthorDIEDC', 'BookSUBJ', 'NumberOfTokens']
metadata_new = pd.read_csv('/media/sf_VBox_Shared/Arabic/Fiqh/merged_metadata.csv')
metadata_new['Bookname'] = metadata_new.filename_old.str.extract('(.*)\.txt', expand=False)
#metadata_merged = metadata_df['Bookname'].reset_index().merge(metadata_new, left_on='Bookname', right_on='Bookname', how='left')
metadata_merged = metadata_new[['Bookname']+metadata_fields].copy()
metadata_merged.columns
# -
df_merged = df_total.merge(metadata_merged, left_on='title', right_on='Bookname', how='left').drop(['Bookname', 'title'], axis=1)
df_merged.to_csv('/media/sf_VBox_Shared/Arabic/Analyses/senses_fiqh.csv', index=False)
tr_dict = {s['root']: s['tr_root'] for i, s in df_total[['root', 'tr_root']].drop_duplicates().iterrows()}
tr_dict
# +
# Also prepare aggregated csv
df_agg = df_total.groupby(['title', 'sense']).size().unstack(fill_value=0)
#df_agg.columns = [u'{} ({})'.format(c, tr_dict[c]) for c in df_agg.columns]
df_agg_merged = df_agg.reset_index().merge(metadata_merged, left_on='title', right_on='Bookname', how='left').drop(['Bookname'], axis=1)
senses_cols = df_agg.columns
senses_cols_relative = [c+'_p' for c in df_agg.columns]
df_agg_merged[senses_cols_relative] = df_agg_merged.apply(lambda r: r[senses_cols]/r['NumberOfTokens'], axis=1)
df_agg_merged.to_csv('/media/sf_VBox_Shared/Arabic/Analyses/senses_fiqh_agg.csv')
# -
|
notebooks/ExtractRoots.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Q3. [3 points] Build a model predicting turnout (LogisticRegression)
# +
# importing libraries
from sklearn.tree import DecisionTreeClassifier # Import Decision Tree Classifier
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split # Import train_test_split function
from sklearn import metrics #Import scikit-learn metrics module for accuracy calculation
import pandas as pd
import numpy as np
from sklearn import preprocessing
import seaborn as sns
# -
FX_indicators_2020_df = pd.read_csv('FX_indicators_2020.csv')
FX_indicators_2020_df
FX_indicators_2020_df.describe()
FX_indicators_2020_df.dtypes
FX_indicators_2020_df.info()
print(FX_indicators_2020_df.shape) # find shape of dataframe
print(FX_indicators_2020_df.isnull().sum()) # check if any nulls
FX_indicators_2020_df.isnull().any()
FX_indicators_2020_df.drop(columns = ['CAND1S', 'CAND2S', 'MESSAGE'], axis=1, inplace=True)
FX_indicators_2020_df = FX_indicators_2020_df.drop(FX_indicators_2020_df.loc[:, 'COMM_LT10':'ED_4COL'].columns, axis = 1)
FX_indicators_2020_df = FX_indicators_2020_df.drop(FX_indicators_2020_df.loc[:, 'H_AFDLN3P':'VG_08'].columns, axis = 1)
FX_indicators_2020_df = FX_indicators_2020_df.drop(FX_indicators_2020_df.loc[:, 'VG_12':'MSG_B'].columns, axis = 1)
FX_indicators_2020_df = FX_indicators_2020_df.drop(FX_indicators_2020_df.loc[:, 'CAND1_UND':'MOVED_ARMB'].columns, axis = 1)
FX_indicators_2020_df
FX_indicators_2020_df.VG_10
print(FX_indicators_2020_df.shape) # find shape of dataframe
print(FX_indicators_2020_df.isnull().sum()) # check if any nulls
FX_indicators_2020_df.dropna(inplace=True)
FX_indicators_2020_df
print(FX_indicators_2020_df.shape) # find shape of dataframe
print(FX_indicators_2020_df.isnull().sum()) # check if any nulls
FX_indicators_2020_df_new = pd.get_dummies(FX_indicators_2020_df)
FX_indicators_2020_df_new
X = FX_indicators_2020_df_new.drop(columns = ['VG_14_DV_Y','VG_14_DV_N', 'D2_N', 'D2_Y', 'R2_N', 'R2_Y', 'D3_N', 'D3_Y',
'R3_N', 'R3_Y','I3_N', 'I3_Y'])
y = FX_indicators_2020_df_new['VG_14_DV_Y']
X
y[0:100]
from sklearn.model_selection import train_test_split
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=1) # 70% training and 30% test
logreg = LogisticRegression()
logreg = logreg.fit(X_train,y_train)
#Predict the response for test dataset
y_pred = logreg.predict(X_test)
y_pred
# +
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# -
# Predict for 1 observation
logreg.predict(X_test.iloc[0].values.reshape(1, -1))
# Predict for multiple observations
logreg.predict(X_test[0:100])
# The score method returns the accuracy of the model
score = logreg.score(X_test, y_test)
print(score)
from sklearn.metrics import confusion_matrix #confusion_matrix
confusion_matrix(y_test, y_pred)
# +
from sklearn.metrics import classification_report # classification_report
print(classification_report(y_test, y_pred))
# -
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
import matplotlib.pyplot as plt
logit_roc_auc = roc_auc_score(y_test, logreg.predict(X_test))
fpr, tpr, thresholds = roc_curve(y_test, logreg.predict_proba(X_test)[:,1])
plt.figure()
plt.plot(fpr, tpr, label='Logistic Regression (area = %0.2f)' % logit_roc_auc)
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.savefig('Log_ROC')
plt.show()
from sklearn.metrics import plot_roc_curve # roc_curve
plot_roc_curve(logreg, X_test, y_test)
prob = logreg.predict_proba(X_test[0:100])
prob
|
Assignment 2 APA track and TESU Masters/Q3. [3 points] Build a model predicting turnout (LogisticRegression).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Datatypes
# ---
#
# [Reference](http://nbviewer.jupyter.org/github/lfapython/pybasics/blob/master/References/1.Datatypes.ipynb)
# ## Problems
# ---
#
# ### Calculate mean of these numbers:
# 5, 16, 18, 32, 12, 3, 8, 17, 14, 13
5 + 16 + 18 + 32 + 12 + 3 + 8 + 17 + 14 + 13
138/10
# ### Which of the above numbers are factors of 738
738%18
738%12
# ### Find percentage of 738 given by above numbers ( use decimal numbers as well )
.5 * 738
16 / 100 * 738
.16 * 738
.17 * 738
17/100 * 738
# - Parenthesis
# - Exponential
# - Multiplication and Division
# - Addition and Substraction
#
# - Left to Right
17 * 738/100
17*738
12546/100
# ## Smallest divisor
#
# write a program to find smallest divisor of a given number.
number = 18
number % 2
number % 3
number % 4
# **%** is remainder operator, above number **18** is perfectly divisible by **2** and **3** while not by **4**
num = input("Enter a number")
# *Much like __print__, __input__ is also a function, but it does exactly opposite of __print__. It get's some data from us*
#
# *What we enter inside the box, is stored in variable __num__*
num
# *The stored data is always string*
# repitition operator
num * 2
# Here is what happens:
#
# 1. __/__ is a division operator, ie we are trying to divide num by __2__
# 2. It gave us __TypeError__, it means we are trying to do something _stupid_.
# - Like in this case trying to divide a string, remember __'25'__ above in quotes.
#
# Trying to do **num / 2** is same as **'25' / 2** like below,
# ###### What happens if we tried to divide num by 2 ?
num / 2
# So what to do ?
#
# 1. First we need to convert that string to a number
# 2. Then divide the number by 2
int(num)
int(num) / 2
# *__int__ converts a string to a integer*
float(num)
# *or in this case, __float__ converts a string to a float*
#
# **But...**
int('a')
# *You cannot convert a "alphabets" to a number this way,*
#
# *Every thing inside __"__ or __'__ is a string.*
# *Let's get to our problem now*
#
# we have a number stored in __num__
num
num = int(num)
num
# *We convert given string of number into a integer now, and stored that in same name ie __num__*
#
# *We can reuse a name/variable, *
#
# Let's finish our problem
num % 2
num % 3
num % 4
num % 5
number = 25
number % 2
number % 3
number % 4
number % 5
# *For __25__ our smallest divisor is __5__*
# ## String
city = 'kathmandu'
city[0]
capital
# 0-based indexing
city[4]
city = 'kathmandu city'
city[9]
""
# __''__ is empty string
city[14]
city.index('u')
"Kathmandu City".index("u")
city.index('a')
city.upper()
city.capitalize()
city
some_char = city[5]
some_char
some_char.upper()
some_char = city[5].upper()
some_char
# ## Solve
#
# If the Roman alpabet has 26 letters, ordered as an alphabet **ABCDEFGHIJKLMNOPQSTUVWXYZ**, then an arbitrary word can be translated by rotating every letter by **13=26/2** places. Thus, **A** becomes **N**, **B** becomes **O** etc. Finally, letter **Z** is translated to **M**.
alphabets = 'ABCDEFGHIJKLMNOPQRSTUVWXYZ'
given = input("Enter a alphabet: ")
given = given.upper()
current = alphabets.index(given)
result = current + 13
alphabets[result]
# *Review line by line*
#
# ```python
# 1. alphabets = 'ABCDEFGHIJKLMNOPQRSTUVWXYZ'
# 2. given = input("Enter a alphabet: ")
# 3. given = given.upper()
# 4. current = alphabets.index(given)
# 5. result = current + 13
# 6. alphabets[result]
# ```
#
# 1. we are storing all uppercase characters in a string named __alphabets__
# 2. we are getting any letter from user
# 3. if user enters lowercase letter we convert it to uppercase
# 4. we find assigned number of that letter
# - in every string a number is assigned to characters in an incremental order
# - i.e _A_ is assigned _0_, _B_ _1_ ... _Z_ = _25_
# - those numbers are called index
# 5. we added _13_ to that _index_ as required.
# 6. we access the letter in the calculated index _result_
# - i.e in case of _b_ it is _1 + 13_ = _14_
# - _alphabets[14]_ means give us the letter/character whose index is _14_
# *It gave use result for B, C, D and so on but what will it give if we passed Z ?*
alphabets = 'ABCDEFGHIJKLMNOPQRSTUVWXYZ'
given = input("Enter a alphabet: ")
given = given.upper()
current = alphabets.index(given)
result = current + 13
alphabets[result]
# *So what happened here, in case of __Z__*
#
# 4. index of _Z_ is _25_
# 5. we added _25 + 13_ = _38_
# 6. we tried to access _alphabets[38]_
# - but _alphabets_ has index only from _0_ to _25_
#
# *This means we need "0" if result is "26" "1" for "27"*
alphabets = 'ABCDEFGHIJKLMNOPQRSTUVWXYZ'
given = input("Enter a alphabet: ")
given = given.upper()
current = alphabets.index(given)
result = (current + 13) - (current // 13) * 26
alphabets[result]
# *What did we changed*
#
# 1. we substract _26_ from _result_ if _current_ is more than _12_ ie _result_ will be more than _25_.
# - we divided _current_ with _13_ which will result in either _0_ or _1_ since it is integer division and maximum index is _25_.
# - if _current_ value is _13_ or more _current // 13_ will be _1_ and _1 * 26_ will be 26.
# - if _current_ value is _12_ or less _current // 13_ will be _0_ and _0 * 26_ will be _0_ and result will be _current + 13_.
#
# We will output result in nice way
current
(current + 13) - (current // 13) * 26
15//13
13//13
12//13
alphabets = 'ABCDEFGHIJKLMNOPQRSTUVWXYZ'
given = input("Enter a alphabet: ")
given = given.upper()
current = alphabets.index(given)
result = (current + 13) - (current // 13) * 26
print('You have given "' + given + '" for which result is "' + alphabets[result] + '".')
# Above we have created a longer string by adding multiple small string together with addition__( + )__ operator.
#
# *Joining multiple string to form single string is called __concatenation__.*
'You have given "' + given + '" for which result is "' + alphabets[result] + '".'
'you have ' + 'some file '
'you have ', 'some file '
print('You have given "', given, '" for which result is "', alphabets[result], '".')
'The sum of 2 and 5 is ' + 7
'The sum of 2 and 5 is ' + str(7)
# ## Simple Problems
#
# - [http://codingbat.com/prob/p182144](http://codingbat.com/prob/p182144)
# +
# Hi, Bye -> HiByeByeHi
# we are defining names for each words
first_word = 'Hi'
second_word = 'Bye'
final_word = first_word + second_word * 2 + first_word
print(final_word)
# -
# _when **\*** is used with string, it repeates the given string __times__ the given number._
print("Result is '{}'".format(final_word))
# *See. reference on [string formatting]().*
# ### Alphabets have values too
ord("a")
ord("ज")
chr(2348)
# ### Practice
#
# [codeabbey.com](http://www.codeabbey.com/)
#
# [projecteuler.net](https://projecteuler.net/)
#
# [practicepython.org](http://www.practicepython.org/)
|
Sessions/1.Datatypes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://maltem.com/wp-content/uploads/2020/04/LOGO_MALTEM.png" style="float: left; margin: 20px; height: 55px">
#
# <br>
# <br>
# <br>
# <br>
#
# # Lab 3_01: Statistical Modeling and Model Validation
#
# > Authors: <NAME>, <NAME>
#
# ---
# ## Objective
# The goal of this lab is to guide you through the modeling workflow to produce the best model you can. In this lesson, you will follow all best practices when slicing your data and validating your model.
# ## Imports
# +
# Import everything you need here.
# You may want to return to this cell to import more things later in the lab.
# DO NOT COPY AND PASTE FROM OUR CLASS SLIDES!
# Muscle memory is important!
import pandas as pd
from scipy.stats import ttest_ind
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score, mean_squared_error
import matplotlib.pyplot as plt
import statsmodels.api as sm
# %matplotlib inline
# -
# ## Read Data
# The `citibike` dataset consists of Citi Bike ridership data for over 224,000 rides in February 2014.
# Read in the citibike data in the data folder in this repository.
citibike = pd.read_csv('../data/citibike_feb2014.csv')
# ## Explore the data
# Use this space to familiarize yourself with the data.
#
# Convince yourself there are no issues with the data. If you find any issues, clean them here.
# Check first five rows.
citibike.head()
# Check datatypes and numbers of non-null values.
citibike.info()
# Summarize all variables.
citibike.describe(include='all')
# Check for missing values. (This is easier to read
# than the .info() output.)
citibike.isnull().sum()
# ## Is average trip duration different by gender?
#
# Conduct a hypothesis test that checks whether or not the average trip duration is different for `gender=1` and `gender=2`. Be sure to specify your null and alternative hypotheses, and to state your conclusion carefully and correctly!
# $$
# \begin{eqnarray*}
# &H_0:& \mu_1 = \mu_2 \\
# &H_A:& \mu_1 \neq \mu_2
# \end{eqnarray*}
# $$
#
# We will conduct this test assuming $\alpha=0.05$.
ttest_ind(citibike[citibike['gender'] == 1]['tripduration'],
citibike[citibike['gender'] == 2]['tripduration'])
# **Answer**: Our $t$-statistic is -5.93 and our $p$-value is very, very small. Because $p$ is smaller than $\alpha$, we reject our null hypothesis and accept that $\mu_1 \neq \mu_2$. This means that we accept that the average trip duration is different for `gender=1` and `gender=2`.
# ## What numeric columns shouldn't be treated as numeric?
# **Answer:** The `start station id`, `end station id`, and `bikeid` columns are all categorical in nature (e.g. adding two of these ID numbers together would be meaningless). These are technically integers, but should not be treated that way.
# ## Dummify the `start station id` Variable
# Before dummifying, let's see how many columns we should create.
len(set(citibike['start station id']))
# How many columns are there in the original data?
len(citibike.columns)
# Let's dummy the data.
citibike = pd.get_dummies(citibike,
columns=['start station id'],
drop_first=True)
# How many columns are there now?
len(citibike.columns)
# +
# 329 unique values + 15 original columns = 344.
# We dropped the `start station id` variable: 344 - 1 = 343.
# We set `drop_first = True`: 343 - 1 = 342.
# We got the right number of columns in our output!
# Let's check out our data to make sure it looks like we did this right.
citibike.head()
# -
# ## Engineer a feature called `age` that shares how old the person would have been in 2014 (at the time the data was collected).
#
# - Note: you will need to clean the data a bit.
# First attempt.
citibike['age'] = 2014 - citibike['birth year']
# +
# We got an error! Somewhere, there's a string.
# Check the values in the birth year column.
citibike['birth year'].value_counts()
# +
# Can we just pull out the strings?
# Iterate through all unique values in birth year column.
for i in set(citibike['birth year']):
# Try typecasting each value to be an integer.
try:
int(i)
# If it gives you an error (so it can't be
# printed as an integer), print the value.
except:
print(i)
# -
# How many values of "\N" are there?
citibike[citibike['birth year'] == '\N'].shape[0]
# How many values of "\N" are there?
# We got an error - it interprets \ as an escape character.
# We need to use the escape character twice!
citibike[citibike['birth year'] == '\\N'].shape[0]
# +
# There's 6,717 values, which is just under 3% of the rows.
# Let's replace "\N" with np.nan.
citibike.loc[citibike['birth year'] == '\\N','birth year'] = np.nan
# -
# Did we successfully do this?
citibike.isnull().sum()
# Now let's try creating our age column.
citibike['age'] = citibike['birth year'].map(lambda x: 2014 - int(x),
na_action = 'ignore')
# Let's check to see if age and birth year seem to match up.
citibike['age'].hist();
citibike['birth year'].dropna().astype(int).hist();
# Yes, birth year is a mirror image of age.
# ## Split your data into train/test data
#
# Look at the size of your data. What is a good proportion for your split? **Justify your answer.**
#
# Use the `tripduration` column as your `y` variable.
#
# For your `X` variables, use `age`, `usertype`, `gender`, and the dummy variables you created from `start station id`. (Hint: You may find the Pandas `.drop()` method helpful here.)
#
# **NOTE:** When doing your train/test split, please use random seed 123.
# Because usertype is a column of strings, we must
# dummy that column as well.
citibike = pd.get_dummies(citibike,
columns=['usertype'],
drop_first=True)
X_train, X_test, y_train, y_test = train_test_split(citibike.dropna().drop(columns=['tripduration', 'birth year', 'bikeid',
'end station longitude', 'end station latitude',
'end station name', 'end station id',
'start station longitude', 'start station latitude',
'start station name', 'starttime', 'stoptime']),
citibike.dropna()['tripduration'],
test_size=0.2,
random_state=123)
X_train.head()
X_test.shape
# **Answer**: The more data we train on, the better it will usually perform! I used `test_size = 0.2` because we have lots of data. This leaves a lot of data (about 43,600 rows!) in our test set to still evaluate our model.
# ## Fit a Linear Regression model in `sklearn` predicting `tripduration`.
# +
# Step 1. Instantiate the model.
model = LinearRegression()
# Step 2. Fit the model on the training data.
model.fit(X_train, y_train)
# Step 3. Generate predictions.
preds = model.predict(X_test)
# -
# ## Evaluate your model
# Look at some evaluation metrics for **both** the training and test data.
# - How did your model do? Is it overfit, underfit, or neither?
# - Does this model outperform the baseline? (e.g. setting $\hat{y}$ to be the mean of our training `y` values.)
# +
# Check the MSE on the training and testing sets.
print(f'MSE on testing set: {mean_squared_error(y_train, model.predict(X_train))}')
print(f'MSE on training set: {mean_squared_error(y_test, preds)}')
# +
# Check the R^2 on the training and testing sets.
print(f'R^2 on testing set: {r2_score(y_train, model.predict(X_train))}')
print(f'R^2 on training set: {r2_score(y_test, preds)}')
# -
# **Answer**: Based on the MSE, our model is performing far worse on the testing set than on the training set, which means that our model is likely overfit to the data.
#
# Based on the $R^2$, our model is explaining approximately zero variance in the $Y$ data. Our model is probably quite bad.
# +
plt.figure(figsize = (12, 9))
# Examine the relationship between observed and predicted values.
plt.scatter(y_test, preds)
# Line showing perfect predictions.
plt.plot([0, max(max(y_test),max(preds))],
[0, max(max(y_test),max(preds))],
linestyle = '--')
plt.title('Predicted values are quite small,\nbut true values are spread out!', fontsize = 24)
plt.xlabel('True Values', fontsize = 16)
plt.ylabel('Predicted Values', fontsize = 16);
# -
print(f'MSE of baseline model: {mean_squared_error(y_test, [np.mean(y_train)] * len(y_test))}')
print(f'R^2 of baseline model: {r2_score(y_test, [np.mean(y_train)] * len(y_test))}')
# **Answer**: Based on the above information, I conclude that my model is both overfit to the data and a bad model.
# - Our MSE and $R^2$ comparing our observed `y_test` values to the average `y_train` value are better than the MSE and $R^2$ on the more complex model we've fit on the training dataset!
# - I might try removing features to improve the fit of the model.
# ## Fit a Linear Regression model in `statsmodels` predicting `tripduration`.
# Remember, we need to add a constant in statsmodels!
X_train = sm.add_constant(X_train)
model_sm = sm.OLS(y_train, X_train).fit()
# ## Using the `statsmodels` summary, test whether or not `age` has a significant effect when predicting `tripduration`.
# - Be sure to specify your null and alternative hypotheses, and to state your conclusion carefully and correctly **in the context of your model**!
model_sm.summary()
# $$
# \begin{eqnarray*}
# &H_0:& \beta_{age} = 0 \\
# &H_A:& \beta_{age} \neq 0
# \end{eqnarray*}
# $$
#
# We will conduct this test assuming $\alpha=0.05$.
#
# **Answer**: The $p$-value for `age` (found in the `model_sm.summary()` table) is less than 0.001, which means that $p < \alpha$ and we will reject $H_0$. This means we accept our alternative hypothesis, $H_A$, and accept that `age` is a significant predictor of `tripduration`.
# ## Citi Bike is attempting to market to people who they think will ride their bike for a long time. Based on your modeling, what types of individuals should Citi Bike market toward?
# **Answer:** Based on the two hypothesis tests we've run, `age` and `gender` are significant predictors of `tripduration`. If we look at the coefficients for `age` and `gender`, both coefficients are positive, indicating that as `age` and `gender` increase, `tripduration` increases. Based on this alone, we should market toward individuals of older age who identify as `gender=2`. (We should consult a data dictionary to figure out what `2` means, but there isn't one here!)
#
# However, our model performance is quite bad! Our predicted values aren't close to our observed values, and our $R^2$ values are terrible. We may want to iterate on our model and try to improve it before using it to make any serious decisions.
|
Notebook/Lab-regression-and-model-validation/solution-code/solution-code.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="P9bJCDjdlgG6" colab_type="text"
# # **Spit some [tensor] flow**
#
# We need to learn the intricacies of tensorflow to master deep learning
#
# `Let's get this over with`
#
#
# + id="aQwc0re5mFld" colab_type="code" outputId="20ca1a48-1899-4e6b-b1ac-f43db9e3a4c1" colab={"base_uri": "https://localhost:8080/", "height": 34}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
import cv2
print(tf.__version__)
# + [markdown] id="0KL1p_roy9zM" colab_type="text"
# ## Look at the following equations
#
# Well now we're going to go into the details of the implementation;
#
# ```
# x_pred_t = w0 + w1 * x_t-1 + w2 * x_t-2 + w3 * x_t-3 + w4 * x_t-4
#
# x_pred_t+1 = w0 + w1 * x_pred_t + w2 * x_t-1 + w3 * x_t-2 + w4 * x_t-3
# ```
#
# This is an autoregressive model now.
#
# ```
# x_pred_5 = w0 + w1 * x_4 + w2 * x_3 + w3 * x_2 + w4 * x_1
#
# x_pred_6 = w0 + w1 * x_pred_5 + w2 * x_4 + w3 * x_3 + w4 * x_2
#
# ```
#
# We know that the predictions at time T in an RNN depend on all the previous times. Thus, indirectly, y(t) depends on x(t), x(t-1), x(t-2),....., x(2), x(1)
#
# To optimize the weights, we must take the derivative of the equation containing the weights; however, there will be numerous W_input_to_hidden weights in the network.
#
#
# The RNN keeps nesting the older timestamps; the derivatives use multiplication and chain rule in composite functions, thus, the more older the timestamp, the more its gradient vanishes.
#
# ## So how do GRU's work?
#
# Throwback to the RNN equation:
#
# h(t) = activation(W(x,h) x(t) + W(h,h) h(t-1) + b(h))
#
# GRU's calulate two other things to calculate h(t):
#
# - update gate vector z(t)
#
# - reset gate vector r(t)
#
# z(t) = sigmoid(W(x,z) x(t) + W(h,z) h(t-1) + b(z))
#
# r(t) = sigmoid(W(x,r) x(t) + W(h,r) h(t-1) + b(r))
#
# ```
# h(t) = (1 - z(t)) * h(t-1) +
#
# z(t) tanh(W(x,h) x(t) + W(h,h) (r(t) * h(t-1)) + b(h))
# ```
#
# (*) ==== element wise multiplication
#
# z(t), r(t), h(t) ===== Size U
#
# ## Z(t)
#
# Should we take the new value of h(t)? or keep h(t-1)?
#
# - z(t) close to 0, take h(t-1)
#
# - z(t) close to 1, take h(t)
#
# So:
#
# ```
# h(t) = (1 - z(t)) * h(t-1) +
#
# z(t) tanh(W(x,h) x(t) + W(h,h) (r(t) * h(t-1)) + b(h))
# ```
#
# Becomes
#
# ```
# h(t) = (factor keep h(t-1)) * h(t-1) +
#
# (discard h(t-1)) * RNN(x(t), h(t-1))
# ```
#
# ## R(t)
#
# Change the value of h(t-1)
#
# - r(t) close to 0, zero value h(t-1)
#
# - r(t) close to 1, keep value h(t-1)
#
#
# ## So how do LSTM's work?
#
# ### https://towardsdatascience.com/
#
# We add another state to the mix, the cell state c(t)
#
# we add three different neurons:
#
# forget neuron = f(t)
#
# This gate decides which information should be thrown away or kept. Input from h(t-1) and x(t) is passed through this gate; and it uses sigmoid to either forget (0) or remember (1) it.
#
#
# input gate neuron = i(t)
#
# We use this to update the cell state. We pass the h(t-1) and x(t) to the sigmoid function. This will decide which values will be updated in the cell state.
#
# output gate neuron = o(t)
#
# The output gate decides what the next hidden state h(t) should be. Remember that the hidden state contains information on previous inputs. The hidden state is also used for predictions.
#
# First, we pass the previous hidden state and the current input into a sigmoid function.
#
# Then we pass the newly modified cell state to the tanh function.
#
# We multiply the tanh output with the sigmoid output to decide what information the hidden state should carry.
#
# The output is the hidden state.
#
# The new cell state and the new hidden is then carried over to the next time step.
#
# ```
#
# f(t) = sigmoid ( W(x,f) x(t) + W(h, f)h(t-1) + b(f) )
#
# i(t) = sigmoid ( W(x,i) x(t) + W(h, i)h(t-1) + b(i) )
#
# o(t) = sigmoid ( W(x,o) x(t) + W(h, o)h(t-1) + b(o) )
#
#
# c(t) = f(t) * c(t-1) +
#
# i(t) * tanh ( W(x,c) x(t) + W(h,c) h(t-1) + b(c) )
#
#
# h(t) = o(t) * tanh( c(t) )
#
# ```
#
#
#
# + id="CaBxIWkog_i-" colab_type="code" colab={}
from tensorflow.keras.layers import Input, Dropout, Dense, Flatten, SimpleRNN, GRU
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import SGD, Adam, Adamax
# + [markdown] id="ofiCPRlY3MUV" colab_type="text"
# ## Let's get some synthetic data
# + id="PKsiBvQv3PKy" colab_type="code" outputId="84315fb6-873f-4dab-aa7e-cce1f5004b8b" colab={"base_uri": "https://localhost:8080/", "height": 295}
n = 1001
x = 0.1 * np.arange(1,n)
data = np.cos(x) + np.random.randn(n-1) * 0.2
# Visualising the dataset
plt.plot(x, data, color = 'red')
plt.title('X Plot')
plt.xlabel('X')
plt.ylabel('y')
plt.show()
# + id="7FMT9DUV_mzN" colab_type="code" outputId="af971516-2690-4e38-9878-19dbcd616ea6" colab={"base_uri": "https://localhost:8080/", "height": 51}
T = 40
D = 1
X = []
y = []
print(len(data))
print("The last sample would be: " + str(len(data)) + " - " + str(T) + " = " + str((len(data)-T)) )
# + id="s6NII-fuCTkl" colab_type="code" colab={}
for i in range(len(data) - T):
x = data[i:i+T]
X.append(x)
y_temp = data[i+T]
y.append(y_temp)
# + id="0nB4lR65DC9i" colab_type="code" colab={}
# Since we need an N x T x D input
X = np.array(X).reshape(-1, T, D)
y = np.array(y)
# + id="j6TzIr6pC8C5" colab_type="code" outputId="62ced6b3-f65b-4495-d567-d864fd156b94" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(X.shape)
print(y.shape)
N, T, D = X.shape
# + id="9Ob8G90SDwfb" colab_type="code" colab={}
i_layer = Input(shape = (T, D))
h_layer = SimpleRNN(10)(i_layer)
o_layer = Dense(1)(h_layer)
model = Model(i_layer, o_layer)
model.compile(loss = 'mse',
optimizer = Adam(lr = 0.1))
# + id="A936E2iuElkz" colab_type="code" outputId="9f837072-7822-40f1-81f3-aeb803d216e4" colab={"base_uri": "https://localhost:8080/", "height": 1000}
index = -N//2
report = model.fit(X[:index], y[:index], epochs=50, validation_data=(X[index:], y[index:]))
# + id="8xVbWzu2BTjT" colab_type="code" outputId="549fc993-76ac-4681-a489-8b38a7c83144" colab={"base_uri": "https://localhost:8080/", "height": 284}
plt.plot(report.history['loss'], label='training_loss')
plt.plot(report.history['val_loss'], label='validation_loss')
plt.legend()
# + id="IuarNdxvFvEW" colab_type="code" colab={}
y_test = y[index:]
y_pred = []
# + id="qWRNknWnF3U4" colab_type="code" colab={}
X_end = X[index]
while len(y_pred) < len(y_test):
pred = model.predict(X_end.reshape(1, -1))[0,0]
y_pred.append(pred)
X_end = np.roll(X_end, -1)
X_end[-1] = pred
# + id="i8dR2ZYJHt_A" colab_type="code" outputId="b91dd233-fbda-4b7a-f4e1-a41d6bce1516" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(len(y_pred))
print(len(y_test))
# + id="ckP1_2XYHi0h" colab_type="code" outputId="5a4cd325-04ad-4c49-c845-3d56e3cdce90" colab={"base_uri": "https://localhost:8080/", "height": 282}
plt.plot(y_test, label='y_test')
plt.plot(y_pred, label='y_pred')
plt.legend()
# + id="HC3bV6Lsa-ai" colab_type="code" outputId="5d85d04f-0cec-4177-debd-658c70662544" colab={"base_uri": "https://localhost:8080/", "height": 282}
y_test_single = y[index:]
y_pred_single = []
i = index
while len(y_pred_single) < len(y_test_single):
pred = model.predict(X[i].reshape(1,T, D))[0,0]
i+=1
y_pred_single.append(pred)
plt.plot(y_test_single, label='y_test_single')
plt.plot(y_pred_single, label='y_pred_singe')
plt.legend()
# + [markdown] id="4rp827FMc-Uy" colab_type="text"
# ## This is bad, the single prediction shows that our model just copies the previous value, this is bad bad bad
#
# But we're in too deep in the world of deep learning, we must find a way! Let's fix our window
#
# + id="OrSGJwUldECo" colab_type="code" outputId="233f3d4d-6af0-4811-b8cc-fb99239762e7" colab={"base_uri": "https://localhost:8080/", "height": 1000}
T = 40
D = 1
X = []
y = []
print(len(data))
print("The last sample would be: " + str(len(data)) + " - " + str(T) + " = " + str((len(data)-T)) )
for i in range(len(data) - T):
x = data[i:i+T]
X.append(x)
y_temp = data[i+T]
y.append(y_temp)
X = np.array(X).reshape(-1, T, D)
y = np.array(y)
i_layer = Input(shape = (T, D))
h_layer = GRU(10, activation='tanh')(i_layer)
o_layer = Dense(1)(h_layer)
model = Model(i_layer, o_layer)
model.compile(loss = 'mse',
optimizer = Adam(lr = 0.2))
index = -N//4
report = model.fit(X[:index], y[:index], epochs=50, validation_data=(X[index:], y[index:]))
plt.plot(report.history['loss'], label='training_loss')
plt.plot(report.history['val_loss'], label='validation_loss')
plt.legend()
# + id="kNsp-ogydhfQ" colab_type="code" outputId="7fa2f279-2a59-485d-bada-92692603e2a2" colab={"base_uri": "https://localhost:8080/", "height": 283}
y_test = y[index:]
y_pred = []
X_end = X[index]
while len(y_pred) < len(y_test):
pred = model.predict(X_end.reshape(1, -1))[0,0]
y_pred.append(pred)
X_end = np.roll(X_end, -1)
X_end[-1] = pred
plt.plot(y_test, label='y_test')
plt.plot(y_pred, label='y_pred')
plt.legend()
# + id="y0tLu-XNj3qh" colab_type="code" outputId="54fa43c5-901a-4566-cccc-009935a3d06f" colab={"base_uri": "https://localhost:8080/", "height": 283}
y_test_single = y[index:]
y_pred_single = []
i = index
while len(y_pred_single) < len(y_test_single):
pred = model.predict(X[i].reshape(1,T, D))[0,0]
i+=1
y_pred_single.append(pred)
plt.plot(y_test_single, label='y_test_single')
plt.plot(y_pred_single, label='y_pred_singe')
plt.legend()
# + id="_9hHMWhNsRrY" colab_type="code" outputId="dea5785d-4b51-4207-dfa4-3484a93edaef" colab={"base_uri": "https://localhost:8080/", "height": 1000}
T = 100
D = 1
X = []
y = []
print(len(data))
print("The last sample would be: " + str(len(data)) + " - " + str(T) + " = " + str((len(data)-T)) )
for i in range(len(data) - T):
x = data[i:i+T]
X.append(x)
y_temp = data[i+T]
y.append(y_temp)
X = np.array(X).reshape(-1, T, D)
y = np.array(y)
i_layer = Input(shape = (T, D))
h_layer = GRU(10, activation='tanh')(i_layer)
o_layer = Dense(1)(h_layer)
model = Model(i_layer, o_layer)
model.compile(loss = 'mse',
optimizer = Adam(lr = 0.2))
index = -N//4
report = model.fit(X[:index], y[:index], epochs=50, validation_data=(X[index:], y[index:]))
plt.plot(report.history['loss'], label='training_loss')
plt.plot(report.history['val_loss'], label='validation_loss')
plt.legend()
# + id="GvZgWolzsfW0" colab_type="code" outputId="66984ec3-e10a-46eb-cf63-3eca1186f4d9" colab={"base_uri": "https://localhost:8080/", "height": 283}
y_test = y[index:]
y_pred = []
X_end = X[index]
while len(y_pred) < len(y_test):
pred = model.predict(X_end.reshape(1, -1))[0,0]
y_pred.append(pred)
X_end = np.roll(X_end, -1)
X_end[-1] = pred
plt.plot(y_test, label='y_test')
plt.plot(y_pred, label='y_pred')
plt.legend()
# + id="mT8NwdRDt7gO" colab_type="code" outputId="c0d4beeb-2320-4e0c-f981-a4cd44b06c7b" colab={"base_uri": "https://localhost:8080/", "height": 283}
y_test_single = y[index:]
y_pred_single = []
i = index
while len(y_pred_single) < len(y_test_single):
pred = model.predict(X[i].reshape(1,T, D))[0,0]
i+=1
y_pred_single.append(pred)
plt.plot(y_test_single, label='y_test_single')
plt.plot(y_pred_single, label='y_pred_singe')
plt.legend()
|
Tensorflow_2X_Notebooks/Demo45_RNNGatedRecurrentUnit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Thank you for helping our study
#
#
# <a href="#/slide-1-0" class="navigate-right" style="background-color:blue;color:white;padding:10px;margin:2px;font-weight:bold;">Continue with the lesson</a>
#
# Throughout this lesson you will see reminders, like the one below, to ensure that all participants understand that they are in a voluntary research study.
#
# ### Reminder
#
# <font size="+1">
#
# By continuing with this lesson you are granting your permission to take part in this research study for the Hour of Cyberinfrastructure: Developing Cyber Literacy for GIScience project. In this study, you will be learning about cyberinfrastructure and related concepts using a web-based platform that will take approximately one hour per lesson. Participation in this study is voluntary.
#
# Participants in this research must be 18 years or older. If you are under the age of 18 then please exit this webpage or navigate to another website such as the Hour of Code at https://hourofcode.com, which is designed for K-12 students.
#
# If you are not interested in participating please exit the browser or navigate to this website: http://www.umn.edu. Your participation is voluntary and you are free to stop the lesson at any time.
#
# For the full description please navigate to this website: <a href="gateway-1.ipynb">Gateway Lesson Research Study Permission</a>.
#
# </font>
# + hide_input=true init_cell=true slideshow={"slide_type": "skip"} tags=["Hide"]
# This code cell starts the necessary setup for Hour of CI lesson notebooks.
# First, it enables users to hide and unhide code by producing a 'Toggle raw code' button below.
# Second, it imports the hourofci package, which is necessary for lessons and interactive Jupyter Widgets.
# Third, it helps hide/control other aspects of Jupyter Notebooks to improve the user experience
# This is an initialization cell
# It is not displayed because the Slide Type is 'Skip'
from IPython.display import HTML, IFrame, Javascript, display
from ipywidgets import interactive
import ipywidgets as widgets
from ipywidgets import Layout
import getpass # This library allows us to get the username (User agent string)
# import package for hourofci project
import sys
sys.path.append('../../supplementary') # relative path (may change depending on the location of the lesson notebook)
import hourofci
# load javascript to initialize/hide cells, get user agent string, and hide output indicator
# hide code by introducing a toggle button "Toggle raw code"
HTML('''
<script type="text/javascript" src=\"../../supplementary/js/custom.js\"></script>
<style>
.output_prompt{opacity:0;}
</style>
<input id="toggle_code" type="button" value="Toggle raw code">
''')
# + [markdown] slideshow={"slide_type": "slide"}
# # Introduction and History
#
# In this section we will cover the history of computation and computing and why the need for the word cyberinfrastructure arose.
#
# - For a long time (thousands of years) people talked about computation and computers.
# - Then in 2003, the National Science Foundation decided that a new word was needed to talk about the infrastructure that was used to support the creation of knowledge: **cyberinfrastructure**.
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Why?
#
# To understand what happened, let’s look a bit at the history of computation and computers.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Early History of Computation
#
# <table>
# <tr style="background-color:transparent">
# <td style="padding-right:50px; width:700px">
# <ul>
# <li>During the time of the Roman empire (about 2,000 years ago), calculating where stones thrown by a catapult would land was an important mathematical problem.</li>
# <li>Here is an example of a catapult, used by Rome and by countries that invaded Rome.</li>
# </ul>
# </td>
# <td>
# <img src='supplementary/catapult.png', width="450"/>
# <font size="-1">Image credit: <a href="https://commons.wikimedia.org/wiki/File:Mang2.png">Wikimedia</a></font>
# </td>
# </tr>
# </table>
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Run some simulations of your own!
#
# - To get a sense for the number of parameters you need to consider to estimate the distance a boulder (payload ) is thrown by a catapult, you can run a simulation of a catapult.
# - Try it a few times!
#
# + hide_input=false slideshow={"slide_type": "slide"} tags=["Init", "Hide"]
IFrame("supplementary/catapult.html", width=984, height=700)
# + [markdown] slideshow={"slide_type": "slide"}
# ### An early “computation device”
#
# <table>
# <tr style="background-color:transparent">
# <td style="padding-right:50px; width:700px">
# <ul>
# <li>Archimedes was an important mathematician who made many discoveries.</li>
# <li>He lived from 287 to 212 BC.</li>
# <li>He was valuable to the army of Carthage (which was at war with Rome during Archimedes lifetime) because he was very good at calculating where stones thrown by a catapult would land. This was a calculation that had essentially one number as <b>output</b>: how far the stone would go.</li>
# </ul>
# </td>
# <td>
# <img src="supplementary/Archimedes.jpg" width="300"/>
# <font size="-1">Image credit: <a href="https://commons.wikimedia.org/wiki/File:Domenico-Fetti_Archimedes_1620.jpg">(Wikimedia)</a></font>
# </td>
# </tr>
# </table>
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Let's figure out how quickly Archimedes could calculate the distance a boulder is going to fly
# Once a catapult is built, there are really just two parameters you can adjust that impact the distance:
# 1. Mass of the projectile
# 2. Amount of tension
#
# These two parameters are called **inputs** that will determine the **output**, which is distance a boulder will fly.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Let's figure out how quickly Archimedes could calculate the distance a boulder is going to fly
#
# Inputs and outputs are simply **data** that Archimedes can use and produce as a human calculator to calculate distance. Let's get a little more precise in how we represent this data. Computers are based on the binary number system, which means they use 0's and 1's. One **bit** is either a 0 or a 1. One **byte** is 8 bits.
#
# | Binary Number | Decimal Number |
# |------|--------|
# |00000000 | 0 |
# |00000001 | 1 |
# |00000010 | 2 |
# |00001010 | 10 |
# |00010000 | 16 |
# |11111111 | 255 |
#
# Two bytes (or 16 bits) can store numbers up to 65,536. Four bytes (or 32 bits) can store numbers up to 4,294,967,295! See more about binary numbers [here](https://en.wikipedia.org/wiki/Binary_number). If we can store more numbers, then we can be more precise.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## How do bits relate to precision?
#
# Let's take a look at the following image. If Archimedes is trying to hit the target that is in the firing range, then he needs to have small enough units to be able to communicate the location. If he has only 1 bit, then he can either fire the maximum distance (1) or half the maximum distance (0). If he has 4 bits, then he has 16 different locations that can be calculated. This is similar to the difference between measuring distance using kilometers versus meters. If the target is 1,400 meters away, then 1km is too short, but 2 km is too far.
#
# <img src="supplementary/catapult-bits.png">
# + [markdown] slideshow={"slide_type": "slide"}
# ## Let's figure out how quickly Archimedes could calculate the distance a boulder is going to fly
#
# Let’s figure that maybe people were really precise with their measurements and the two input values might take four bytes each, and the one output value might take four bytes as well...
#
# So that means that we have:
# * 2 * 4 bytes of **input**
# * 1 * 4 bytes of **output**
#
# We can use these to figure out the **Input/Output Rate** or **I/O Rate**, which is how quickly we can accept input, run the calculations, and produce output. What do you think the "I/O rate" of Archimedes might have been?
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Well, we don't really know ....
#
# because we don’t know how quickly Archimedes could run his calculations. Perhaps he approximated his results. He was, after all, trying to help soldiers crush other soldiers with rocks; he wasn’t doing brain surgery.
#
# But let’s figure maybe 1 calculation in 5 minutes and 4 Bytes of I/O per minute, tops, was the I/O rate for Archimedes as a human calculator. If that is the case it would take him two minutes to get the input location from a soldier (2 * 4 bytes), five minutes to run his calculations, and another minute to communicate the distance to the catapult launcher (1 * 4 bytes). So his I/O rate would be approximately 8 minutes to input, calculate, and output 12 bytes.
# + [markdown] slideshow={"slide_type": "fragment"}
# Now let's compare Archimedes to others types of calculating machines...
# + [markdown] slideshow={"slide_type": "slide"}
# ## Later there were “calculating machines”
#
# - These are mechanical devices that performed calculations
# - The Chinese Abacus was very practical and skilled people using it were very fast
# - The Arithmometer, manufactured and sold in 1851, was the first commercially successful calculating machine for office use
#
# <table>
# <tr style="background-color:transparent">
# <td style="padding-right:50px; width:50%">
# <figure>
# <div>
# <img src="https://upload.wikimedia.org/wikipedia/commons/a/af/Abacus_6.png" width="250"/>
# <footer><small><!-- copyright noice --></small></footer>
# </div>
# </figure>
# <font size="-1">Image credit: [Wikimedia](https://commons.wikimedia.org/wiki/File:Abacus_6.png)</font>
# </td>
# <td>
# <figure>
# <div>
# <img src="https://upload.wikimedia.org/wikipedia/commons/5/59/Arithmometre.jpg" width="300"/>
# <footer><small><!-- copyright noice --></small></footer>
# </div>
# </figure>
# <font size="-1">Image credit: [Wikipedia](https://en.wikipedia.org/wiki/Arithmometer)</font>
# </td>
# </tr>
# </table>
#
#
#
#
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Wait a moment...!
#
# You're about to see a virtual Chinese abacus with a hexadecimal numeral system. Hexadecimal numeral system is simply a numeral system that has 16 base digits (instead of 10 for decimal).
# The hexadecimal base digits are 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, a, b, c, d, e, f which an "f" is equivalent to 15 in decimal system (see <a href = https://en.wikipedia.org/wiki/Hexadecimal>here</a> for more details).
#
# The abacus has an upper deck with two beads, each is worth 5, and the bottom has five, each worth 1.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Try a virtual Abacus simulator and see how fast you can make it go!
# + hide_input=true slideshow={"slide_type": "-"} tags=["Init", "Hide"]
# Auto-run
IFrame("supplementary/abacus.html", width=600, height=475)
# + [markdown] slideshow={"slide_type": "slide"}
# > In 2012, <NAME>, a 22-year-old abacus instructor from Japan completed the task of calculating 10 sums of 10 10-digit numbers each in three minutes and 11 seconds. ( [Recordholders.org](http://www.recordholders.org/en/events/worldcup/2012/results.html); [theguardian.com](https://www.theguardian.com/science/alexs-adventures-in-numberland/2012/oct/10/mental-calculation-world-cup) )
#
# ### What is the calculation rate and I/O rate that the winner achieved?
#
# - The input was 100 integers each of which could be represented by 2 bytes (200 bytes total)
# - The output was 10 integers each of which could be represented by 2 bytes (20 bytes total)
# - The number of seconds was 191
# - Each sum of 10 integers took 9 additions, so there were a total of 900 mathematical **operations**
# - The total calculation rate was approximately (900 operations / 191 seconds) or ~4.7 operations per second
# - So the total I/O rate was something like 220 bytes/191 seconds or between 1 and 2 bytes per second
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## The first fully electronic computer
#
# <table>
# <tr style="background-color:transparent">
# <td style="padding-right:50px; width:700px; text-align: left;">
# <ul>
# <li>The Z3, invented by <NAME>, in Berlin, Germany in 1941.</li>
# <li>It could accept any program and took about 1 second per addition and 3 seconds per multiplication of a 22 bit number. Faster than an abacus but not much!</li>
# <li>We’re not quite sure what the I/O rates were - but input was with a keyboard and output with lights, so … pretty slow.</li>
# </ul>
# </td>
# <td>
# <figure>
# <div>
# <img src="https://upload.wikimedia.org/wikipedia/commons/4/4c/Z3_Deutsches_Museum.JPG" width="400"/>
# <footer><small><!-- copyright noice --></small></footer>
# </div>
# </figure>
#
# <font size="-1">Image credit: [Wikimedia](https://upload.wikimedia.org/wikipedia/commons/4/4c/Z3_Deutsches_Museum.JPG)</font>
# </td>
# </tr>
# </table>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Supercomputers!
#
# In the 1970s and 1980s there were lots of different labels for computers. Mainframe computers, minicomputers, workstations. The label “supercomputers” was invented for several reasons:
# - This word distinguished the most powerful computers on earth from "ordinary" computers
# - It sounds cool
#
# ### What makes a supercomputer "super"?
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## What makes a supercomputer "super"?
# - There is no fixed and agreed on definition. The general idea is that a supercomputer is one of the most powerful computers around
# - One of the general characteristics of supercomputers is that they break computational problems up into many parts and work on those problems in **parallel** – many different processors each analyzing a part of a problem
# - If you have a computer and it costs more than \$1,000,000 and you want to call it “super” then go ahead!
#
# *Side note:* To use supercomputers effectively to solve problems, it is important to learn about **parallel computing** so make sure to check out the beginner lesson on parallel computing if you are interested!
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## The First Supercomputer
#
# <table>
# <tr style="background-color:transparent">
# <td style="padding-right:50px; width:700px; text-align: left;">
# <ul>
# <li>The first supercomputer which was both “super” and called a supercomputer was the Control Data 6600, designed by <NAME>. The first system was delivered to a commercial customer in 1964.</li>
# <li>This supercomputer could do 3,000 calculations per second.</li>
# <li>I/O Rate. Because of the way the system was designed, it’s tricky to calculate an input rate. But output was with a teletype, so output was no more than about 10 characters per second.</li>
# </ul>
# </td>
# <td>
# <figure>
# <div>
# <img src="https://upload.wikimedia.org/wikipedia/commons/c/c4/CDC_6600.jc.jpg" width="400"/>
# <footer><small><!-- copyright noice --></small></footer>
# </div>
# </figure>
#
# <font size="-1">Image credit: [Wikipedia](https://upload.wikimedia.org/wikipedia/commons/c/c4/CDC_6600.jc.jpg)</font>
# </td>
# </tr>
# </table>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Today’s fastest supercomputer
#
#
# <table>
# <tr style="background-color:transparent">
# <td style="padding-right:50px; width:700px; text-align: left;">
# <ul>
# <li>The fastest (unclassified) supercomputer in the world as of Summer 2020 is called Summit, at Oak Ridge National Labs in the US.</li>
# <li>It has achieved a calculation speed of 148,600,000,000,000 calculations per second. (Calculations are measured now in “FLOPS” - floating point operations per second - and that’s 148 PetaFLOPS. Or, in round numbers, really fast. </li>
# <li>I/O rate is 100 GigaBytes per second from an external network (that’s 1,000,000,000 Bytes per second). Local output - to a local file system - is 2.5 TByte / second, or 2,500,000,000,000 Bytes per second.</li>
# </ul>
# </td>
# <td>
# <figure>
# <div>
# <img src="https://upload.wikimedia.org/wikipedia/commons/b/b4/Summit_%28supercomputer%29.jpg" width="400"/>
# <footer><small><!-- copyright noice --></small></footer>
# </div>
# </figure>
#
# <font size="-1">Image credit: [Wikipedia](https://en.wikipedia.org/wiki/Summit_(supercomputer)#/media/File:Summit_(supercomputer).jpg)</font>
# </td>
# </tr>
# </table>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Supercomputers help people do cool stuff
#
#
# <table style="width:90%">
# <tr style="background-color:transparent">
# <td style="padding-right:50px; width:650px; text-align: left;">
# Many discoveries were made with supercomputers, including:
# <ul>
# <li>Calculate the mass of subatomic particles</li>
# <li>Simulate how suns form</li>
# <li>Simulate how tornadoes form</li>
# <li>Solving the four color problem. This is a mathematical problem that is hundreds of years old. The problem is this: prove that any map can be colored with four colors and no two adjoining countries on the map will be colored the same. Supercomputers were used to solve this problem in the mid 1970s. See the illustration</li>
# </ul>
# </td>
# <td>
# <figure>
# <div>
# <img src="https://upload.wikimedia.org/wikipedia/commons/8/8a/Four_Colour_Map_Example.svg" width="350"/>
# <footer><small><!-- copyright noice --></small></footer>
# </div>
# </figure>
#
# Image credit: https://commons.wikimedia.org/wiki/File:Four_Colour_Map_Example.svg
# </td>
# </tr>
# </table>
#
#
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Can you solve the four color problem?
# + hide_input=true slideshow={"slide_type": "-"} tags=["Init", "Hide"]
IFrame("supplementary/fourcolor.html", width="970", height="730")
# + [markdown] slideshow={"slide_type": "slide"}
# ## Let's look back: what was the evolution of I/O rates of computing devices?
# In early computing devices, up to early supercomputers, the ratio of Calculation / IO was very high. This is called the Compute / Bandwidth ratio now.
#
# | Device | Year | Calculation Rate (measured in Floating Point Operations per Second) | I/O Rate |
# | :-: | :-: | :-: | :-: |
# | Archimedes | 0 | <1 | 1 Byte / Minute
# | Abacus | 1000 | 5 | 1 Byte / Second
# | Z3 | 1941 | ⅓ to 1 operation per second | Not sure - but slow
# | Control Data 6600 | 1943 | 3,000 | 10 Bytes / second
# | Summit - the fastest supercomputer in the world | 2018 | 148,600,000,000,000 | In, across a network: 1,000,000,000 Bytes / second <br/>Out, locally to a file system: 2,500,000,000,000 Bytes / second
#
# + [markdown] slideshow={"slide_type": "slide"}
# <figure>
# <div>
# <img src="supplementary/congratulations.png" width="400"/>
# <footer><small><!-- copyright notice --></small></footer>
# </div>
# </figure>
#
# ## You now understand three very important concepts
# 1. The early history of computation
# 2. I/O Rate, the speed at which data is input and output
# 3. Calculation Rate (now measured in Floating Point Operations per Second or FLOP)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Next
#
# Let's learn about cyberinfrastructure and how even supercomputers are enough for science.
#
# <a href="cyberinfrastructure-3.ipynb">Click here to move to the next section to learn more!</a>
|
beginner-lessons/cyberinfrastructure/cyberinfrastructure-2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dual Momentum GEM
#
# (optimize lookback)
# +
import pandas as pd
import matplotlib.pyplot as plt
import datetime
from talib.abstract import *
import pinkfish as pf
import strategy
# format price data
pd.options.display.float_format = '{:0.2f}'.format
# %matplotlib inline
# -
# set size of inline plots
'''note: rcParams can't be in same cell as import matplotlib
or %matplotlib inline
%matplotlib notebook: will lead to interactive plots embedded within
the notebook, you can zoom and resize the figure
%matplotlib inline: only draw static images in the notebook
'''
plt.rcParams["figure.figsize"] = (10, 7)
# Some global data
# +
symbols = {'SP500' : 'SPY', 'BONDS' : 'AGG', 'EXUS' : 'EFA', 'T-BILL': 'BIL'}
capital = 10000
start = datetime.datetime(1900, 1, 1)
end = datetime.datetime.now()
# num months lookback
lookback = None
use_cache = True
# -
# Define Optimizations
# +
# pick one
optimize_lookback = True
# define high low trade periods ranges
if optimize_lookback:
Xs = range(3, 18+1, 1)
Xs = [str(X) for X in Xs]
# -
# Run Strategy
strategies = pd.Series(dtype=object)
for X in Xs:
print("{0}".format(X), end=" ")
if optimize_lookback:
strategies[X] = strategy.Strategy(symbols, capital, start, end, lookback=int(X),
margin=1, use_cache=True, use_absolute_mom = True)
strategies[X].run()
strategies[X].rlog, strategies[X].tlog, strategies[X].dbal = strategies[X].get_logs()
strategies[X].stats = strategies[X].get_stats()
# Summarize results
# +
metrics = ('annual_return_rate',
'max_closed_out_drawdown',
'drawdown_annualized_return',
'drawdown_recovery',
'best_month',
'worst_month',
'sharpe_ratio',
'sortino_ratio',
'monthly_std',
'pct_time_in_market',
'total_num_trades',
'pct_profitable_trades',
'avg_points')
df = strategy.summary(strategies, metrics)
df
# -
# Bar graphs
strategy.plot_bar_graph(df, 'annual_return_rate')
strategy.plot_bar_graph(df, 'sharpe_ratio')
strategy.plot_bar_graph(df, 'max_closed_out_drawdown')
# Run Benchmark
s = strategies[Xs[0]]
benchmark = pf.Benchmark('SPY', capital, s.start, s.end, use_adj=True)
benchmark.run()
benchmark.tlog, benchmark.dbal = benchmark.get_logs()
benchmark.stats = benchmark.get_stats()
# Equity curve
# +
if optimize_lookback: Y = '3'
pf.plot_equity_curve(strategies[Y].dbal, benchmark=benchmark.dbal)
# -
def plot_equity_curves(strategies):
"""
Plot Equity Curve: multiple equity curves on same plot
Arguement is daily balance.
"""
fig = plt.figure(figsize=(16,12))
axes = fig.add_subplot(111, ylabel='Portfolio value in $')
for strategy in strategies:
axes.plot(strategy.dbal['close'], label=strategy.lookback)
plt.legend(loc='best')
plot_equity_curves(strategies[3:10:2])
|
examples/momentum-gem/optimize.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: csci566Assg3
# language: python
# name: csci566asg3
# ---
# +
from platform import python_version
import tensorflow as tf
print(tf.test.is_gpu_available())
print(python_version())
# -
import os
import numpy as np
from os import listdir
from PIL import Image
import time
import tensorflow as tf
from tensorflow.keras import layers,models,optimizers
from keras import backend as K
import matplotlib.pyplot as plt
# +
path1="datasets/ofg_family/"
path2="datasets/TSKinFace_Data/TSKinFace_cropped/"
randomiser = np.random.RandomState(123)
# +
def generate_image_1(family_dir):
dic={}
sub=[a for a in listdir(path1+"/"+family_dir)]
for ele in sub:
if ele == '.DS_Store':
continue;
mypath = path1+"/"+family_dir+"/"+ele+"/"
onlyfiles = [mypath+f for f in listdir(mypath)]
addr = randomiser.choice(onlyfiles)
original_img = np.array(Image.open(addr).resize((64,64),Image.ANTIALIAS))
if ele[0].lower()=='f':
dic['father'] = original_img
elif ele[0].lower()=='m':
dic['mother'] = original_img
elif ele.lower()=='child_male':
dic['child'] = original_img
dic['gender']=np.zeros((original_img.shape))
elif ele.lower()=='child_female':
dic['child'] = original_img
dic['gender'] = np.ones((original_img.shape))
return [dic['father'],dic['mother'],dic['gender'],dic['child']]
def generate_image_2(family_dir, family_number, gender):
dic={}
sub = ["F" , "M", gender]
family_pth = path2+"/"+family_dir+"/" + family_dir + "-" + str(family_number) + "-"
for ele in sub:
addr = family_pth+ele+".jpg"
original_img = np.array(Image.open(addr).resize((64,64),Image.ANTIALIAS))
if ele =='F':
dic['father'] = original_img
elif ele == 'M':
dic['mother'] = original_img
elif ele == 'S':
dic['child'] = original_img
dic['gender']=np.zeros((original_img.shape))
elif ele == 'D':
dic['child'] = original_img
dic['gender'] = np.ones((original_img.shape))
return [dic['father'],dic['mother'],dic['gender'],dic['child']]
def generate_batch(families_batch):
np_images=[]
for family in families_batch:
if(len(family) == 3):
res = generate_image_2(family[0], family[1], family[2])
elif(len(family) == 1):
res = generate_image_1(family[0])
if( res != None):
np_images.append(res)
return np_images
# +
for r, d, f in os.walk(path1):
all_families = d
break
all_families = [[family] for family in all_families]
for i in range(285):
all_families.append(['FMS', i+1, 'S'])
for i in range(274):
all_families.append(['FMD', i+1, 'D'])
for i in range(228):
all_families.append(['FMSD', i+1, 'D'])
all_families.append(['FMSD', i+1, 'S'])
randomiser.shuffle(all_families)
train_families = all_families[:-500]
test_families = all_families[-500:]
# -
OUTPUT_CHANNELS = 3
# +
mean = 0.
std_dev = 0.02
lr = 0.0001
b1 = 0.5
sd_random_normal_init = 0.02
EPOCHS = 20
batch = 125
# -
def gen_downsample_parent(filters, size, apply_batchnorm=True, apply_dropout=False):
initializer = tf.random_normal_initializer(mean, std_dev)
result = tf.keras.Sequential()
result.add(
tf.keras.layers.Conv2D(filters, size, strides=2, padding='same',
kernel_initializer=initializer,
use_bias=False))
if apply_batchnorm:
result.add(tf.keras.layers.BatchNormalization())
result.add(tf.keras.layers.ELU())
if apply_dropout:
result.add(tf.keras.layers.Dropout(rate = 0.5))
return result
def gen_downsample_noise(filters, size, apply_batchnorm=True):
initializer = tf.random_normal_initializer(mean, std_dev)
result = tf.keras.Sequential()
result.add(
tf.keras.layers.Conv2DTranspose(filters, size, strides=2, padding='same',
kernel_initializer=initializer,
use_bias=False))
if apply_batchnorm:
result.add(tf.keras.layers.BatchNormalization())
result.add(tf.keras.layers.ELU())
return result
def gen_upsample(filters, size,apply_batchnorm = False):
initializer = tf.random_normal_initializer(mean, std_dev)
result = tf.keras.Sequential()
result.add(
tf.keras.layers.Conv2DTranspose(filters, size, strides=2,
padding='same',
kernel_initializer=initializer,
use_bias=False))
if apply_batchnorm:
result.add(tf.keras.layers.BatchNormalization())
result.add(tf.keras.layers.ELU())
return result
# +
def EncoderNN():
down_stack_parent = [
gen_downsample_parent(32,4,apply_batchnorm=True, apply_dropout=True),
gen_downsample_parent(64,4,apply_batchnorm=True, apply_dropout=False)
]
down_stack_noise =[
# z = 4x4x64
gen_downsample_noise(64,4,apply_batchnorm=True), #8x8x64
gen_downsample_noise(32,4,apply_batchnorm=True) #16x16x32
]
final_conv =[
gen_upsample(32,4 ,apply_batchnorm = True)
]
initializer = tf.random_normal_initializer(mean, sd_random_normal_init)
last = tf.keras.layers.Conv2DTranspose(OUTPUT_CHANNELS, 4,
strides=2,
padding='same',
kernel_initializer=initializer,
activation='tanh')
concat = tf.keras.layers.Concatenate()
father = tf.keras.layers.Input(shape=(64,64,3))
mother = tf.keras.layers.Input(shape=(64,64,3))
# create noise 4x4x64
# noise = tf.random.normal((1,4,4,64),mean=0.0,stddev=1.0,dtype=tf.dtypes.float32)
# print(noise.shape)
x1 = father
for down in down_stack_parent:
x1 = down(x1)
# print(x1.shape)
x2 = mother
for down in down_stack_parent:
x2 = down(x2)
# print(x2.shape)
# n = noise
# for down in down_stack_noise:
# print(n.shape)
# n = down(n)
# print(n.shape)
# print("NOISE STACK DONE")
final = concat([x1,x2])
final = final_conv[0](final)
final = last(final)
return tf.keras.Model(inputs=[father, mother], outputs=final)
# + active=""
# family_data = generate_image(all_families[700])
# noise = tf.random.normal((4,4,64),mean=0.0,stddev=1.0,dtype=tf.dtypes.float32)
# inp = np.array([family_data[0],family_data[1]])
# inp.shape
# -
encoder_optimizer = tf.keras.optimizers.Adam(learning_rate = lr, beta_1=b1)
def tensor_to_array(tensor1):
return tensor1.numpy()
# +
def train_encoder(father_batch, mother_batch, target_batch, b_size):
with tf.GradientTape() as enc_tape:
gen_outputs = encoder([father_batch, mother_batch], training=True)
diff = tf.abs(target_batch - gen_outputs)
flatten_diff = tf.reshape(diff, (b_size, 64*64*3))
encoder_loss_batch = tf.reduce_mean(flatten_diff, axis=1)
encoder_loss = tf.reduce_mean(encoder_loss_batch)
print("ENCODER_LOSS: ",tensor_to_array(encoder_loss))
#calculate gradients
encoder_gradients = enc_tape.gradient(encoder_loss,encoder.trainable_variables)
#apply gradients on optimizer
encoder_optimizer.apply_gradients(zip(encoder_gradients,encoder.trainable_variables))
# -
def fit_encoder(train_ds, epochs, test_ds, batch):
losses=np.array([])
for epoch in range(epochs):
print("______________________________EPOCH %d_______________________________"%(epoch+1))
start = time.time()
for i in range(len(train_ds)//batch):
batch_data = np.asarray(generate_batch(train_ds[i*batch:(i+1)*batch]))
batch_data = batch_data / 255 * 2 -1
print("Generated batch", batch_data.shape)
X_Father_train = tf.convert_to_tensor(batch_data[:,0],dtype =tf.float32)
X_Mother_train = tf.convert_to_tensor(batch_data[:,1],dtype =tf.float32)
Y_train = tf.convert_to_tensor(batch_data[:,3],dtype =tf.float32)
train_encoder(X_Father_train, X_Mother_train, Y_train,batch)
print("Trained for batch %d/%d"%(i+1,(len(train_ds)//batch)))
print("______________________________TRAINING COMPLETED_______________________________")
# +
train_dataset = all_families[:-500]
test_dataset = all_families[-500:]
encoder = EncoderNN()
with tf.device('/cpu:0'):
fit_encoder(train_dataset, EPOCHS, test_dataset,batch)
# +
f_no = 560
family_data = generate_image(all_families[f_no][0], all_families[f_no][1], all_families[f_no][2])
inp = [family_data[0],family_data[1]]
inp = tf.cast(inp, tf.float32)
father_inp = inp[0][tf.newaxis,...]
mother_inp = inp[1][tf.newaxis,...]
with tf.device('/cpu:0'):
gen_output = encoder([father_inp, mother_inp], training=True)
temp = gen_output.numpy()
plt.imshow(np.squeeze(temp))
# print(temp)
print(np.amin(temp))
print(np.amax(temp))
# +
# family_data = generate_image(all_families[126])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=True)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# family_data = generate_image(all_families[126])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=True)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# family_data = generate_image(all_families[0])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=True)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# family_data = generate_image(all_families[126])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=True)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# family_data = generate_image(all_families[126])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=True)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# family_data = generate_image(all_families[126])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=True)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# family_data = generate_image(all_families[126])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=True)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# # epochs = 10 bs 250 lr 0.002 mean = 0.02
# family_data = generate_image(all_families[700])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=False)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# # epoch 10 batch = 250 mean 0.04 lr 1e-4 beta_1= 0.5
# family_data = generate_image(all_families[700])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=False)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# # epoch 10 batch = 250 mean 0.02 lr 0.0005
# family_data = generate_image(all_families[700])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=False)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# # epoch 20 batch = 400
# family_data = generate_image(all_families[700])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=False)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# # epochs : 10 batch size 250
# family_data = generate_image(all_families[700])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=False)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# +
# #epochs = 4 batch_size : 250
# family_data = generate_image(all_families[700])
# inp = [family_data[0],family_data[1]]
# inp = tf.cast(inp, tf.float32)
# with tf.device('/cpu:0'):
# gen_output = encoder(inp, training=False)
# temp = gen_output.numpy()
# plt.imshow(np.squeeze(temp))
# # print(temp)
# print(np.amin(temp))
# print(np.amax(temp))
# + active=""
# # epochs = 1
# batch = 500
#
# for epoch in range(epochs):
# # print("Epoch ", epoch , " .....")
# for i in range(len(train_families)//batch):
# batch_data = np.asarray(generate_batch(train_families[i*batch:(i+1)*batch]))
# batch_data = batch_data / 255 * 2 -1
# # print("Generated batch", batch_data.shape)
#
# X_train = tf.convert_to_tensor(concat([batch_data[:,0],batch_data[:,1]]),dtype =tf.float32)
# # print("Batch converted to tensor")
#
# Y_train = batch_data[:,3]
# history = encoder.fit(X_train, Y_train, batch_size=batch)
# print("Training DONE!")
# + active=""
# family_data = generate_image(all_families[1])
# inp = concat([family_data[0],family_data[1]])
# inp = tf.cast(inp, tf.float32)
# gen_output = encoder(inp[tf.newaxis,...], training=False)
# temp = gen_output.numpy()
# temp = np.squeeze(temp)
# plt.imshow(temp)
#
# print(np.amin(temp))
# print(np.amax(temp))
# print(encoder.trainable_weights[0][0][0][0])
# 0.00196027
# # plt.imshow((gen_output[0]+1)/2)
# -
def disc_downsample_parent_target(filters, size, apply_batchnorm=True):
initializer = tf.random_normal_initializer(mean, std_dev)
result = tf.keras.Sequential()
result.add(
tf.keras.layers.Conv2D(filters, size, strides=2, padding='same',
kernel_initializer=initializer,
use_bias=False))
if apply_batchnorm:
result.add(tf.keras.layers.BatchNormalization())
result.add(tf.keras.layers.LeakyReLU(alpha = 0.2))
return result
def disc_loss(filters, size,apply_batchnorm = False):
initializer = tf.random_normal_initializer(mean, std_dev)
result = tf.keras.Sequential()
result.add(
tf.keras.layers.Conv2D(filters, size, strides=2,
padding='same',
kernel_initializer=initializer,
use_bias=False))
if apply_batchnorm:
result.add(tf.keras.layers.BatchNormalization())
result.add(tf.keras.layers.LeakyReLU(alpha = 0.2))
return result
def Discriminator():
father = tf.keras.layers.Input(shape=(64,64,3))
mother = tf.keras.layers.Input(shape=(64,64,3))
target = tf.keras.layers.Input(shape=(64,64,3))
down_stack_parent_target = [
disc_downsample_parent_target(32,4,apply_batchnorm=False),
disc_downsample_parent_target(64,4,apply_batchnorm=True)
]
down_stack_combined =[
disc_loss(192,4,apply_batchnorm=True),
disc_loss(256,4,apply_batchnorm=False)
]
initializer = tf.random_normal_initializer(mean, sd_random_normal_init)
last = tf.keras.layers.Conv2D(1, 4, strides=1,padding='same',
kernel_initializer=initializer) # linear layer
concat = tf.keras.layers.Concatenate()
x1 = father
for down in down_stack_parent_target:
x1 = down(x1)
x2 = mother
for down in down_stack_parent_target:
x2 = down(x2)
x3 = target
for down in down_stack_parent_target:
x3 = down(x3)
combined = concat([x1,x2,x3])
# combined is 1x16x16x192
x4 = combined
for down in down_stack_combined:
x4 = down(x4)
output = last(x4) #4X4
# print(output.shape)
return tf.keras.Model(inputs=[father,mother,target], outputs=output)
# +
#discriminator = Discriminator()
# +
# family_data = generate_image(all_families[126])
# p1 = tf.cast(family_data[0], tf.float32)
# p2 = tf.cast(family_data[1], tf.float32)
# c = tf.cast(family_data[2], tf.float32)
# discriminator = Discriminator()
# with tf.device('/cpu:0'):
# disc_out = discriminator(inputs = [p1,p2,c], training=True)
# -
LAMBDA = 0.1
loss_object = tf.keras.losses.BinaryCrossentropy()
def tensor_to_array(tensor1):
return tensor1.numpy()
def discriminator_loss(disc_real_output, disc_generated_output):
# real_loss = loss_object(tf.ones_like(disc_real_output,dtype=tf.float32), disc_real_output) #L1 loss
real_loss = tf.reduce_mean(tf.abs(tf.ones_like(disc_real_output) - disc_real_output))
generated_loss = tf.reduce_mean(tf.abs(tf.zeros_like(disc_generated_output) - disc_generated_output))
# generated_loss = loss_object(tf.zeros_like(disc_generated_output,dtype=tf.float32), disc_generated_output)
total_disc_loss = real_loss + generated_loss
print("D real loss: {}, D fake loss: {}".format(real_loss, generated_loss))
return total_disc_loss
def generator_loss(disc_generated_output, gen_output, target):
# gan_loss = loss_object(tf.ones_like(disc_generated_output,dtype=tf.float32), disc_generated_output)
gan_loss = tf.reduce_mean(tf.abs(tf.ones_like(disc_generated_output) - disc_generated_output))
# mean absolute error
l1_loss = tf.reduce_mean(tf.abs(target - gen_output))
total_gen_loss = l1_loss #+ LAMBDA * gan_loss +
print("Reconstruction loss: {}, GAN loss: {}".format(l1_loss, gan_loss))
return total_gen_loss
generator_optimizer = tf.keras.optimizers.Adam(lr, beta_1=b1)
discriminator_optimizer = tf.keras.optimizers.Adam(lr, beta_1=b1)
# +
def train_step(father_batch, mother_batch, target_batch,b_size):
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
gen_outputs = encoder([father_batch, mother_batch], training=True)
print("Generated outputs",gen_outputs.shape)
disc_real_output = discriminator([father_batch, mother_batch, target_batch], training=True)
print("disc_real_output ", disc_real_output.shape)
disc_generated_output = discriminator([father_batch, mother_batch, gen_outputs], training=True)
print("disc_generated_output ", disc_generated_output.shape)
gen_loss = generator_loss(disc_generated_output, gen_outputs, target_batch)
disc_loss = discriminator_loss(disc_real_output, disc_generated_output)
# inp_batch = tf.unstack(inp_batch)
# target_batch = tf.unstack(target_batch)
# gen_loss =tf.Variable(0,dtype='float32')
# disc_loss =tf.Variable(0,dtype='float32')
# for idx,inputs in enumerate(inp_batch):
# gen_output = encoder(inputs, training=True)
# # gen_output shape is 1x64x64x3
# disc_real_output = discriminator([inputs[0],inputs[1],target_batch[idx]], training=True)
# # print(disc_real_output)
# disc_generated_output = discriminator([inputs[0],inputs[1],tf.squeeze(gen_output)], training=True)
# # print(disc_generated_output)
# gen_loss = gen_loss + generator_loss(disc_generated_output, gen_output, target_batch[idx])
# disc_loss = disc_loss + discriminator_loss(disc_real_output, disc_generated_output)
# gen_loss/=b_size
# disc_loss/=b_size
print("GEN_LOSS",tensor_to_array(gen_loss))
print("DISC_LOSS",tensor_to_array(disc_loss))
generator_gradients = gen_tape.gradient(gen_loss,encoder.trainable_variables)
discriminator_gradients = disc_tape.gradient(disc_loss,discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(generator_gradients,
encoder.trainable_variables))
discriminator_optimizer.apply_gradients(zip(discriminator_gradients,
discriminator.trainable_variables))
# -
def fit(train_ds, epochs, test_ds,batch):
for epoch in range(epochs):
print("______________________________EPOCH %d_______________________________"%(epoch))
start = time.time()
for i in range(len(train_ds)//batch):
batch_data = np.asarray(generate_batch(train_ds[i*batch:(i+1)*batch]))
print("Generated batch", batch_data.shape)
X_father_train = tf.convert_to_tensor(batch_data[:,0],dtype =tf.float32)
X_mother_train = tf.convert_to_tensor(batch_data[:,1],dtype =tf.float32)
# print("Xtrain",X_train.shape)
# print("Batch converted to tensor")
Y_train = tf.convert_to_tensor(batch_data[:,3],dtype =tf.float32)
train_step(X_father_train, X_mother_train, Y_train, batch)
print("Trained for batch %d/%d"%(i+1,(len(train_ds)//batch)))
family_data = generate_image(all_families[700])
inp = [family_data[0],family_data[1]]
inp = tf.cast(inp, tf.float32)
father_inp = inp[0][tf.newaxis,...]
mother_inp = inp[1][tf.newaxis,...]
gen_output = encoder([father_inp, mother_inp], training=True)
print(tf.reduce_min(gen_output))
print(tf.reduce_max(gen_output))
plt.figure()
plt.imshow(gen_output[0,...])
plt.show()
print("______________________________TRAINING COMPLETED_______________________________")
checkpoint.save(file_prefix = checkpoint_prefix)
concat = tf.keras.layers.Concatenate()
train_dataset = all_families[:-500]
test_dataset = all_families[-500:]
encoder = EncoderNN()
discriminator = Discriminator()
checkpoint_dir = './checkpoint'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=encoder,
discriminator=discriminator)
with tf.device('/cpu:0'):
fit(train_dataset, EPOCHS, test_dataset,batch)
family_data = generate_image(all_families[1])
inp = [family_data[0],family_data[1]]
inp = tf.cast(inp, tf.float32)
with tf.device('/cpu:0'):
gen_output = encoder(inp, training=True)
print(tf.reduce_min(gen_output))
print(tf.reduce_max(gen_output))
plt.imshow(gen_output[0,...])
|
DCGAN-TSkin_Old.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Crossentropy method
#
# This notebook will teach you to solve reinforcement learning problems with crossentropy method.
# +
import gym
import numpy as np, pandas as pd
env = gym.make("Taxi-v2")
env.reset()
env.render()
# +
n_states = env.observation_space.n
n_actions = env.action_space.n
print("n_states=%i, n_actions=%i"%(n_states, n_actions))
# -
# # Create stochastic policy
#
# This time our policy should be a probability distribution.
#
# ```policy[s,a] = P(take action a | in state s)```
#
# Since we still use integer state and action representations, you can use a 2-dimensional array to represent the policy.
#
# Please initialize policy __uniformly__, that is, probabililities of all actions should be equal.
#
policy = np.ones([n_states,n_actions])/np.ones([n_states,n_actions]).sum(axis=1,keepdims=True)
assert type(policy) in (np.ndarray,np.matrix)
assert np.allclose(policy,1./n_actions)
assert np.allclose(np.sum(policy,axis=1), 1)
# # Play the game
#
# Just like before, but we also record all states and actions we took.
def generate_session(policy,t_max=10**4):
"""
Play game until end or for t_max ticks.
:param policy: an array of shape [n_states,n_actions] with action probabilities
:returns: list of states, list of actions and sum of rewards
"""
states,actions = [],[]
total_reward = 0.
s = env.reset()
for t in range(t_max):
a = np.random.choice(list(range(n_actions)),p=policy[s])# <sample action from policy (hint: use np.random.choice)>
new_s, r, done, info = env.step(a)
#Record state, action and add up reward to states,actions and total_reward accordingly.
states.append(s)
actions.append(a)
total_reward += r
s = new_s
if done:
break
return states, actions, total_reward
s,a,r = generate_session(policy)
assert type(s) == type(a) == list
assert len(s) == len(a)
assert type(r) in [float,np.float]
# +
#let's see the initial reward distribution
import matplotlib.pyplot as plt
# %matplotlib inline
sample_rewards = [generate_session(policy,t_max=1000)[-1] for _ in range(200)]
plt.hist(sample_rewards,bins=20);
plt.vlines([np.percentile(sample_rewards, 50)], [0], [100], label="50'th percentile", color='green')
plt.vlines([np.percentile(sample_rewards, 90)], [0], [100], label="90'th percentile", color='red')
plt.legend()
# -
# ### Crossentropy method steps (2pts)
def select_elites(states_batch,actions_batch,rewards_batch,percentile=50):
"""
Select states and actions from games that have rewards >= percentile
:param states_batch: list of lists of states, states_batch[session_i][t]
:param actions_batch: list of lists of actions, actions_batch[session_i][t]
:param rewards_batch: list of rewards, rewards_batch[session_i][t]
:returns: elite_states,elite_actions, both 1D lists of states and respective actions from elite sessions
Please return elite states and actions in their original order
[i.e. sorted by session number and timestep within session]
If you're confused, see examples below. Please don't assume that states are integers (they'll get different later).
"""
reward_threshold = np.percentile(rewards_batch,percentile)#<Compute minimum reward for elite sessions. Hint: use np.percentile>
elite_states = [state for i in range(len(rewards_batch)) if rewards_batch[i]>= reward_threshold for state in states_batch[i] ]
elite_actions = [action for i in range(len(rewards_batch)) if rewards_batch[i] >= reward_threshold for action in actions_batch[i]]
return elite_states,elite_actions
# +
states_batch = [
[1,2,3], #game1
[4,2,0,2], #game2
[3,1] #game3
]
actions_batch = [
[0,2,4], #game1
[3,2,0,1], #game2
[3,3] #game3
]
rewards_batch = [
3, #game1
4, #game2
5, #game3
]
test_result_0 = select_elites(states_batch, actions_batch, rewards_batch, percentile=0)
test_result_40 = select_elites(states_batch, actions_batch, rewards_batch, percentile=30)
test_result_90 = select_elites(states_batch, actions_batch, rewards_batch, percentile=90)
test_result_100 = select_elites(states_batch, actions_batch, rewards_batch, percentile=100)
assert np.all(test_result_0[0] == [1, 2, 3, 4, 2, 0, 2, 3, 1]) \
and np.all(test_result_0[1] == [0, 2, 4, 3, 2, 0, 1, 3, 3]),\
"For percentile 0 you should return all states and actions in chronological order"
assert np.all(test_result_40[0] == [4, 2, 0, 2, 3, 1]) and \
np.all(test_result_40[1] ==[3, 2, 0, 1, 3, 3]),\
"For percentile 30 you should only select states/actions from two first"
assert np.all(test_result_90[0] == [3,1]) and \
np.all(test_result_90[1] == [3,3]),\
"For percentile 90 you should only select states/actions from one game"
assert np.all(test_result_100[0] == [3,1]) and\
np.all(test_result_100[1] == [3,3]),\
"Please make sure you use >=, not >. Also double-check how you compute percentile."
print("Ok!")
# -
def update_policy(elite_states,elite_actions):
"""
Given old policy and a list of elite states/actions from select_elites,
return new updated policy where each action probability is proportional to
policy[s_i,a_i] ~ #[occurences of si and ai in elite states/actions]
Don't forget to normalize policy to get valid probabilities and handle 0/0 case.
In case you never visited a state, set probabilities for all actions to 1./n_actions
:param elite_states: 1D list of states from elite sessions
:param elite_actions: 1D list of actions from elite sessions
"""
new_policy = np.zeros([n_states,n_actions]) #* 1/n_actions
#<Your code here: update probabilities for actions given elite states & actions>
for s,a in zip(elite_states,elite_actions):
new_policy[s][a] += 1
default_p = 1/n_actions
for i in range(new_policy.shape[0]):
sa_sum = new_policy[i].sum()
if sa_sum == 0:
new_policy[i] = np.ones(n_actions) * default_p
else:
for j in range(new_policy.shape[1]):
if new_policy[i][j] > 0:
new_policy[i][j]/= sa_sum
#Don't forget to set 1/n_actions for all actions in unvisited states.
return new_policy
# +
elite_states, elite_actions = ([1, 2, 3, 4, 2, 0, 2, 3, 1], [0, 2, 4, 3, 2, 0, 1, 3, 3])
new_policy = update_policy(elite_states,elite_actions)
assert np.isfinite(new_policy).all(), "Your new policy contains NaNs or +-inf. Make sure you don't divide by zero."
assert np.all(new_policy>=0), "Your new policy can't have negative action probabilities"
assert np.allclose(new_policy.sum(axis=-1),1), "Your new policy should be a valid probability distribution over actions"
reference_answer = np.array([
[ 1. , 0. , 0. , 0. , 0. ],
[ 0.5 , 0. , 0. , 0.5 , 0. ],
[ 0. , 0.33333333, 0.66666667, 0. , 0. ],
[ 0. , 0. , 0. , 0.5 , 0.5 ]])
assert np.allclose(new_policy[:4,:5],reference_answer)
print("Ok!")
# -
# # Training loop
# Generate sessions, select N best and fit to those.
# +
from IPython.display import clear_output
def show_progress(batch_rewards, log, percentile, reward_range=[-990,+10]):
"""
A convenience function that displays training progress.
No cool math here, just charts.
"""
mean_reward, threshold = np.mean(batch_rewards), np.percentile(batch_rewards, percentile)
log.append([mean_reward,threshold])
clear_output(True)
print("mean reward = %.3f, threshold=%.3f"%(mean_reward, threshold))
plt.figure(figsize=[8,4])
plt.subplot(1,2,1)
plt.plot(list(zip(*log))[0], label='Mean rewards')
plt.plot(list(zip(*log))[1], label='Reward thresholds')
plt.legend()
plt.grid()
plt.subplot(1,2,2)
plt.hist(batch_rewards,range=reward_range);
plt.vlines([np.percentile(batch_rewards, percentile)], [0], [100], label="percentile", color='red')
plt.legend()
plt.grid()
plt.show()
# -
#reset policy just in case
policy = np.ones([n_states, n_actions]) / n_actions
# +
n_sessions = 350 #sample this many sessions
percentile = 10 #take this percent of session with highest rewards
learning_rate = 0.7 #add this thing to all counts for stability
log = []
for i in range(100):
# %time sessions = [generate_session(policy) for _ in range(n_sessions)]#[<generate a list of n_sessions new sessions>]
batch_states,batch_actions,batch_rewards = zip(*sessions)
elite_states, elite_actions = select_elites(batch_states,batch_actions,batch_rewards,percentile)#<select elite states/actions>
new_policy = update_policy(elite_states,elite_actions)#<compute new policy>
policy = learning_rate * new_policy + (1-learning_rate) * policy
#display results on chart
show_progress(batch_rewards, log, percentile)
# -
# ### Reflecting on results
#
# You may have noticed that the taxi problem quickly converges from <-1000 to a near-optimal score and then descends back into -50/-100. This is in part because the environment has some innate randomness. Namely, the starting points of passenger/driver change from episode to episode.
#
# In case CEM failed to learn how to win from one distinct starting point, it will siply discard it because no sessions from that starting point will make it into the "elites".
#
# To mitigate that problem, you can either reduce the threshold for elite sessions (duct tape way) or change the way you evaluate strategy (theoretically correct way). You can first sample an action for every possible state and then evaluate this choice of actions by running _several_ games and averaging rewards.
# ### Submit to coursera
from submit import submit_taxi
submit_taxi(generate_session, policy, <EMAIL>, <TOKEN>)
|
week1_intro/crossentropy_method.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py37] *
# language: python
# name: conda-env-py37-py
# ---
# +
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append('../')
sys.path.append('../RunAnova')
from RunAnova import OneWayAnova
import pandas as pd
# +
resp_var = 'age'
treatment = 'company'
experiment = {'NASA' : [18,19,20,21,22,23,18,19,20,21],
'Tesla' : [18,20,16,20,21,20,18,19,17,13],
'Orange' : [21,22,17,18,22,19,21,20,18,23]}
exp_df = pd.DataFrame(experiment)
exp_df.index.name = "observation"
exp_df
# -
anova_proc = OneWayAnova(exp_df, resp_var=resp_var, treatments=treatment)
anova_proc.show_distribution()
anova_proc.compute_indep_ci(0.95, show=True)
anova_proc.normality_test(test_type="qqplot")
anova_proc.normality_test(test_type="shapiro")
anova_proc.normality_test(test_type="levene")
anova_proc.run_anova(simple=False)
|
notebooks/example_oneway_cie_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=[]
import tensorflow as tf
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.layers import Dropout, Dense, Dropout, Input, GlobalAveragePooling1D
import tensorflow.keras as keras
from util import DynamicPadding
from encoder import Encoder
import numpy as np
# %load_ext autoreload
# %autoreload 2
# %load_ext tensorboard
import warnings
warnings.filterwarnings('ignore')
# -
#
# ## Load Data
#
# We take the IMBD data, and sort the training data according to the length of the sequence. Sorting introduces more uniform batch sizes w.r.t. the sequence length which reduces training time considerably if combined with dynamical padding. Furthermore, we crop sequences beyond 200 tokens.
#
# + tags=[]
vocab_size = 20000 # Only consider the top 20k words
maxlen = 200 # Only consider the first 200 words of each movie review
(x_train, y_train), (x_val, y_val) = keras.datasets.imdb.load_data(num_words=vocab_size)
print(len(x_train), "Training sequences")
print(len(x_val), "Validation sequences")
# sort training data w.r.t. the sequence length
seq_length = [len(x) for x in x_train]
permuted_indicies = np.argsort(seq_length)
x_train, y_train = x_train[permuted_indicies], y_train[permuted_indicies]
# crop sequences
x_train = [x[:maxlen] for x in x_train]
x_val = [x[:maxlen] for x in x_val]
# -
# ## Dynamical Padding
#
# We overwrite the Keras Sequence class to support dynamical padding which pads batches only and therefore reduce sequence length. This speeds up training because Transformers training time growths quadratically with the sequence legth. See also [<NAME>](https://towardsdatascience.com/divide-hugging-face-transformers-training-time-by-2-or-more-21bf7129db9q-21bf7129db9e) contribution for further details.
# dump the data into the Dynamic Padding batch loader
train = DynamicPadding(x_train, y_train, batch_size=64)
test = DynamicPadding(x_val, y_val, batch_size=64)
# ## Build the Model
#
# Build a Classifier by using a single encoding layer. The architecture is adopted from the official [Keras example](https://keras.io/examples/nlp/text_classification_with_transformer/) by <NAME>.
# +
embed_dim = 32 # Embedding size for each token
num_heads = 2 # Number of attention heads
ff_dim = 32 # Hidden layer size in feed forward network inside transformer
inputs = Input(shape=(maxlen,))
encoder_embedding = Encoder(vocab_size + 1, maxlen, embed_dim, num_heads, ffn_units=ff_dim, encoders=1)
x = encoder_embedding(inputs)
x = GlobalAveragePooling1D()(x)
x = Dropout(0.1)(x)
x = Dense(20, activation="relu")(x)
x = Dropout(0.1)(x)
outputs = Dense(1, activation='sigmoid')(x)
model = keras.Model(inputs=inputs, outputs=outputs)
# -
# ## Compile and train model
# + tags=[]
adam_opt = Adam(0.001, beta_1=0.9, beta_2=0.98, epsilon=1e-9)
# save the model after each epoch
callbacks = [
tf.keras.callbacks.ModelCheckpoint(
filepath='./imbd_model/prst_model_{epoch}',
save_freq='epoch',
),
tf.keras.callbacks.TensorBoard(
log_dir='./imbd_logs',
profile_batch=0,
)
]
# define relevant metrics
metrics = [
tf.keras.metrics.AUC(),
tf.keras.metrics.Precision(),
tf.keras.metrics.Recall()
]
# compile model
model.compile(
optimizer=adam_opt,
loss="binary_crossentropy",
metrics=metrics,
# run_eagerly=True
)
history = model.fit(
train, validation_data=test, callbacks=callbacks, epochs=2, verbose=1
)
|
imbd_encoder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Quick Start with HydroPy!
# ### an example of incorporating HydroCloud into a Jupyter Notebook.
# *NOTE:* [HydroPy](https://github.com/stijnvanhoey/hydropy) is not required to use HydroCloud in a notebook.
# +
# Import the libraries that we'll be using
import numpy as np
import pandas as pd
import hydropy as hp
# Set the notebook to plot graphs in the output cells.
# %matplotlib inline
# -
# ## Load USGS data into a dataframe
# Use HydroCloud.org to find a stream gauge to investigate.
# Click on the red points to find the site number.
from IPython.display import HTML
HTML('<iframe src=https://hydrocloud.org/ width=700 height=400></iframe>')
# +
# Create a Pandas dataframe using the USGS daily discharge for Herring Run.
herring = hp.get_usgs('01585200', 'dv', '2011-01-01', '2016-01-01')
# List the first few values from the top of the dataframe.
herring.head()
# -
# Calculate some basic statistics for the dataframe.
herring.describe()
# +
# For more advanced analysis, use the HydroAnalysis class.
my_analysis = hp.HydroAnalysis(herring)
# Plot discharge on a logarithmic scale for the Y axis.
my_analysis.plot(figsize=(16,6), logy=True)
# +
## Finding Help
# -
# Use help() to learn more about a particular function.
help(hp.get_usgs)
# # Learn More!
# To learn more about hydropy, read the [documentation](https://readthedocs.org/projects/hydropy/), visit us on [github](https://github.com/stijnvanhoey/hydropy), or try out more [notebooks](Additional_Features.ipynb)!
|
resources/example-Jupyter-Notebook.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Dyalog APL
# language: apl
# name: dyalog-kernel
# ---
# An operator takes 1 or 2 operands (which are usually functions) as arguments, and derives a function which itself can either be monadic or dyadic.
# `/` (reduce) is a monadic operator which derives an ambivalent function. ambivalent meaning that it can be used both monadically and dyadically. It is called reduce because it always *reduces* the rank of its argument by 1.
#
# Reduction on a vector is straightforward; `F/a b c d e...` is equivalent to `⊂a F b F c F d F e...`. The result of a reduction will have the shape of the argument excluding the last axis.
+/3 1 4 1 5
3 + 1 + 4 + 5
×/(1 2 3)(4 5 6)(7 8 9) ⍝ The result is enclosed, because the rank must be 0
# Since functions in APL are right-associative, this has an effect on reduce. For example `-/` is alternating sum.
-/1 2 3 4
1-2-3-4
# For higher rank arrays reduce will reduce along the last axis.
3 4⍴⍳12
+/3 4⍴⍳12
# The twin of `/`, `⌿` reduces along the first axis, i.e the columns of a matrix.
+⌿3 4⍴⍳12
# If you need to reduce across some other axis, like the second in a rank-3 array, you can use `f/[axis]`. `f/[1]` is the same as `f⌿`.
2 3 4⍴⍳24
(+⌿2 3 4⍴⍳24)(+/[2]2 3 4⍴⍳24)(+/2 3 4⍴⍳24)
# As a dyadic function, `L f/ R` is a windowed reduction, i.e the `f`-reduction of each sliding window of size L in R.
2 +/3 1 4 1 5
3 +/ 3 1 4 1 5
# Windowed reduction does not change the rank.
2 +/ 3 4⍴⍳12
2 +⌿ 3 4⍴⍳12
# If the left argument is negative, the windows are reversed.
¯2 -/ 0 1 3 6 10 15
# Common uses of `/` are for sum with `+`, product with `×`, all with `∧`, and any with `∨`.
# `\` (scan) is a similar monadic operator. It reduces each prefix of the last axis. For a vector this means that `f\a b c d...` is `(f/a) (f/a b) (f/a b c) (f/a b c d) ...`
#
# A common use is for cumulative sum:
+\2 3 5 7 11
+\3 4⍴⍳12
# Similarly `\` also has a twin `⍀` who behaves as you might expect. Scan on first axis.
+⍀3 4⍴⍳12
|
Reduce and Scan.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Exercice 1:
# Write a Python class named square constructed by a length and two methods which will compute the area and the perimeter of the square.
class square():
def __init__(self,a):
self.length = a
def area(self):
return(self.length**2)
def perimeter(self):
return(self.length*4)
pass
carre = square(2)
print(carre.area())
print(carre.perimeter())
# ### Exercice 2:
# Write a python class rectangle that inherits from the square class.
class rectangle(square):
def __init__(self,a,b):
self.longueur=a
self.largeur=b
def area(self):
return(self.longueur*self.largeur)
def perimeter(self):
return(self.longueur*2+self.largeur*2)
pass
rec = rectangle(3,5)
print(rec.area())
print(rec.perimeter())
# ### Exercice 3:
# +
class SampleClass:
def __init__(self, a):
## private varibale in Python
self.__a = a
@property
def value(self):
return(self.__a)
@value.setter
def value(self,value):
self.__a=value
x = SampleClass(3)
print(x.value)
x.a = 23
print(x.a)
# -
# Use python decorators to make the above code works
|
exercices/part4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
np.random.seed(42)
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
from plotting import plot
# +
dataset = load_iris()
x = dataset.data[:, :2]
y = dataset.target
# +
colors = ["red", "green", "blue"]
for idx, point in enumerate(x):
plt.scatter(point[0], point[1], color=colors[y[idx]])
plt.show()
# -
class KMeans:
def __init__(self, n_clusters: int = 8, max_iter: int = 3_000):
pass
def fit(self, x: np.ndarray):
pass
def predict(self, x: np.ndarray):
pass
def score(self, x: np.ndarray):
pass
kmeans = KMeans(n_clusters=3, max_iter=1_000)
kmeans.fit(x)
y_pred = kmeans.predict(x)
|
Chapter10_Clustering/KMeansImplementation/KMeans_Implementation_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="rUWDxJYaXbGS"
# ปฏิบัติการครั้งที่ 3 กระบวนวิชา 229351 Statistical Learning for Data Science
#
# คำชี้แจง
#
# 1. ให้เริ่มทำปฏิบัติการจาก colab notebook ที่กำหนดให้ จากนั้นบันทึกเป็นไฟล์ *.pdf
# + [markdown] id="40DP82euz0nP"
# ###ในปฏิบัติการนี้เราจะฝึกการทำ PCA ด้วยสองวิธี คือ
# 1. หาด้วยการแยกส่วนประกอบของเมทริกซ์ผ่าน numpy
# 2. หาด้วยการใช้เครื่องมือที่มีมาให้ใน scikit-learn
# + [markdown] id="BFeBRlckOxSG"
# ดาวน์โหลดข้อมูลสัตว์ต่างๆ 50 ชนิดดังนี้
# ชนิดสัตว์: http://www.donlapark.cmustat.com/229351/data/classes.txt
# ตัวแปรต่างๆ: http://www.donlapark.cmustat.com/229351/data/predicates.txt
# ค่าของสัตว์แต่ละชนิด: http://www.donlapark.cmustat.com/229351/data/predicate-matrix-continuous.txt
# + id="m4HFWsz5j8sS"
#Download the files
# !pip install wget
# !python -m wget -o classes.txt http://www.donlapark.cmustat.com/229351/data/classes.txt
# !python -m wget -o predicate-matrix-continuous.txt http://www.donlapark.cmustat.com/229351/data/predicate-matrix-continuous.txt
# + id="5Tebr3-dBKxj"
import numpy as np
classes = np.genfromtxt('classes.txt',dtype='str')
classes[:5]
# + id="O_4Mr2OWaAae"
data = np.genfromtxt('predicate-matrix-continuous.txt')
data.shape
# + [markdown] id="DntRinCFiSMx"
# ####ใน code block ข้างล่างนี้ จงทำ PCA บนข้อมูลที่ได้มาให้เหลือเมทริกซ์ข้อมูลที่มีตัวแปรแค่ 2 ตัว โดยใช้ฟังก์ชัน $\texttt{np.linalg.eigh}$
# ####ดังนั้น เมทริกซ์ที่ได้ต้องมีขนาด 50x2
# + [markdown] id="H-aWhy49CquB"
# $$ \Sigma =
# \begin{pmatrix}
# \text{var}(X_1) & \text{cov}(X_1,X_2) & \cdots & \text{cov}(X_1,X_{85}) \\
# \text{cov}(X_2,X_1) & \text{var}(X_2) & \cdots & \text{cov}(X_2,X_{85}) \\
# \vdots & \vdots & \ddots & \vdots \\
# \text{cov}(X_{85},X_1) & \text{cov}(X_{85},X_1) & \cdots & \text{var}(X_{85}) \\
# \end{pmatrix}
# $$
# + id="-gCRXKb8fkuH"
data_c = data-np.mean(data, axis = 0)
#TODO: enter code here
#1. หา covariance matrix
#2. Decompose the covariance matrix UDU^T
#3. ดึง column ของ U ที่ประกอบไปด้วย eigenvector สองตัวที่มีค่า eigenvalue สูงที่สุด
#4. เอา data_c ไปทำ projection ทิศทางของ eigenvector ใน U
#Result: a (50,2) matrix
# + [markdown] id="refYgya0jlfh"
# ####OPTIONAL: ใน code block ข้างล่างนี้ จงทำ PCA บนข้อมูลที่ได้มาให้เหลือเมทริกซ์ข้อมูลที่มีตัวแปรแค่ 2 ตัว โดยใช้ $scikit-learn$
# ####พร้อมกับตรวจสอบว่าเมทริกซ์ที่ได้จากทั้งสองวิธีนี้มีค่าเท่ากัน
# + id="TWoz-qMICQsv"
from sklearn.decomposition import PCA
#TODO: enter code here
# + [markdown] id="kvxBs5Hlk5rZ"
# ####ในขั้นตอนสุดท้าย เราจะทำการพล็อตจุดของข้อมูลที่ได้จาก PCA ข้างบนพร้อมกับใส่คำกำกับว่าจุดไหนเป็นของสัตว์ชนิดใดโดยใช้ชื่อสัตว์จาก classes.txt
# + id="2srU_UiTbhS9"
# %matplotlib inline
from matplotlib import pyplot as plt
#กำหนดขนาดของรูป
plt.figure(figsize=(12,16))
#จงเติม argument ที่เหมาะสมในวงเล็บข้างล่างนี้
#รูปแบบของฟังก์ชันคือ plt.scatter(numpy array ของ x-coordinate, numpy array ของ y-coordinate)
plt.scatter( #TODO: 1st column of your 50x2 matrix, #TODO: 2nd column of your 50x2 matrix )
#for loop เพื่อใส่คำกำกับ (annotate) ชื่อสัตว์ของแต่ละจุด วนให้ครบสัตว์ทุกชนิดที่อยู่ใน classes
#ใส่ numpy array ที่ผ่านการทำ PCA แล้วลงในตำแหน่งที่ระบุเพื่อบอกพิกัดที่ต้องวางคำกำกับ
for i in range(50):
plt.annotate(
classes[i,1],
xy=(#TODO: YOUR_MATRIX[i,0],
#TODO YOUR_MATRIX[i,1]),
xytext=(5, -8),
textcoords='offset pixels')
plt.show()
# + [markdown] id="pp8J2lKI0Vea"
# ####จงตอบคำถามต่อไปนี้
#
# 1. จงระบุว่าสัตว์ต่างๆ ที่อยู่มุมขวาบนมีลักษณะใดที่คล้ายคลึงกัน
# 2. จงระบุว่าสัตว์ต่างๆ ที่อยู่มุมซ้ายบนมีลักษณะใดที่คล้ายคลึงกัน
# 3. จงระบุว่าสัตว์ต่างๆ ที่อยู่ด้านล่างมีลักษณะใดที่คล้ายคลึงกัน
# 4. จงหาว่ามีสัตว์กลุ่มใดอีกบ้างที่มีลักษณะคล้ายกันและอยู่ใกล้ๆ กันในแผนภาพข้างบน
# + [markdown] id="SvI3Rwjcpt6c"
#
|
Labs/229351-LAB03-02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# # A Seq2seq model for generating tweets
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
import numpy as np
import pandas as pd
import data_load_seq2seq_utils as s2s_util
import data_load_utils as util
from importlib import reload
util = reload(util)
s2s_util = reload(s2s_util)
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
tweets_orig = util.read_tweet_data('data/emojis_homemade.csv')
tweets_additional_0 = util.read_tweet_data('data/emojis_additional.csv')
tweets=pd.DataFrame.append(tweets_orig, tweets_additional_0)
# +
tweets = util.filter_tweets_min_count(tweets, min_count=1000)
tweets.reset_index()
tweets['text'] = util.filter_text_for_handles(tweets['text'])
# After the filtering, remember to append a \n character to each tweet
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
tweets.head()
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
tweets.shape
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# Define the set of characters that we'll use to encode our text data:
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# Create dicts for character/emoji to index conversion
chars_univ, chars_univ_idx = s2s_util.get_universal_chars_list()
emojis = sorted(list(set(tweets['emoji'])))
emoji_idx = dict((emoji, emojis.index(emoji)) for emoji in emojis)
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
TRAIN_SIZE = 2**19 # 8192 try 131072 = 2**18 for production
DEV_SIZE = 2**14 # 8192 try 8192 = 2**13 for production
TWEETS_PER_BATCH = 2048
MAX_TWEET_LENGTH = 160
n_train_batches = TRAIN_SIZE / TWEETS_PER_BATCH
n_dev_batches = DEV_SIZE / TWEETS_PER_BATCH
print ("n_train_batches:", n_train_batches)
print ("n_dev_batches:", n_dev_batches)
tweets_train = tweets.iloc[0:TRAIN_SIZE] # 8192 = 2**13
tweets_dev = tweets.iloc[TRAIN_SIZE:TRAIN_SIZE+DEV_SIZE] # 2048 = 2**11
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
tweets_train.shape
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
train_generator = s2s_util.xy_generator(tweets_train, emoji_indices=emoji_idx)
dev_generator = s2s_util.xy_generator(tweets_dev, emoji_indices=emoji_idx)
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
([emoj, x], y) = train_generator.__next__()
#e = emoj.reshape(64, 1, 111)
x.shape
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# Now we're going to use the algorithm from the Keras example of a seq2seq model.
# We'll supply the emoji to the encoder LSTM which will encode it into two state vectors,
# and the decoder LSTM will be trained on the tweets using teacher forcing.
#
#
#
# # Summary of the algorithm
#
# - We start with input sequences from a domain (e.g. English sentences)
# and corresponding target sequences from another domain
# (e.g. French sentences).
# - An encoder LSTM turns input sequences to 2 state vectors
# (we keep the last LSTM state and discard the outputs).
# - A decoder LSTM is trained to turn the target sequences into
# the same sequence but offset by one timestep in the future,
# a training process called "teacher forcing" in this context.
# Is uses as initial state the state vectors from the encoder.
# Effectively, the decoder learns to generate `targets[t+1...]`
# given `targets[...t]`, conditioned on the input sequence.
# - In inference mode, when we want to decode unknown input sequences, we:
# - Encode the input sequence into state vectors
# - Start with a target sequence of size 1
# (just the start-of-sequence character)
# - Feed the state vectors and 1-char target sequence
# to the decoder to produce predictions for the next character
# - Sample the next character using these predictions
# (we simply use argmax).
# - Append the sampled character to the target sequence
# - Repeat until we generate the end-of-sequence character or we
# hit the character limit.
#
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# # Building the model
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
from keras.models import Model
from keras.layers import Input, LSTM, Dense
ENCODER_HIDDEN_SIZE = 256
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# Define an input sequence and process it.
encoder_inputs = Input(shape=(None, len(emoji_idx)))
encoder = LSTM(ENCODER_HIDDEN_SIZE, return_state=True)
encoder_outputs, state_h, state_c = encoder(encoder_inputs)
# We discard `encoder_outputs` and only keep the states.
encoder_states = [state_h, state_c]
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# Set up the decoder, using `encoder_states` as initial state.
decoder_inputs = Input(shape=(None, len(chars_univ)))
# We set up our decoder to return full output sequences,
# and to return internal states as well. We don't use the
# return states in the training model, but we will use them in inference.
decoder_lstm = LSTM(ENCODER_HIDDEN_SIZE, return_sequences=True, return_state=True)
decoder_outputs, _, _ = decoder_lstm(decoder_inputs,
initial_state=encoder_states)
decoder_dense = Dense(len(chars_univ), activation='softmax')
decoder_outputs = decoder_dense(decoder_outputs)
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# Define the model that will turn
# `encoder_input_data` & `decoder_input_data` into `decoder_target_data`
model = Model([encoder_inputs, decoder_inputs], decoder_outputs)
# Run training
model.compile(optimizer='rmsprop', loss='categorical_crossentropy')
model.summary()
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
model.fit_generator(train_generator,
steps_per_epoch=n_train_batches,
epochs=100,
validation_data=dev_generator,
validation_steps=n_dev_batches,
verbose=1)
# Save model
model.save('emoji_s2s.h5')
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
encoder_model = Model(encoder_inputs, encoder_states)
decoder_state_input_h = Input(shape=(ENCODER_HIDDEN_SIZE,))
decoder_state_input_c = Input(shape=(ENCODER_HIDDEN_SIZE,))
decoder_states_inputs = [decoder_state_input_h, decoder_state_input_c]
decoder_outputs, state_h, state_c = decoder_lstm(
decoder_inputs, initial_state=decoder_states_inputs)
decoder_states = [state_h, state_c]
decoder_outputs = decoder_dense(decoder_outputs)
decoder_model = Model(
[decoder_inputs] + decoder_states_inputs,
[decoder_outputs] + decoder_states)
# -
reverse_chars_idx = dict(
(i, char) for char, i in chars_univ_idx.items())
def decode_sequence(input_seq):
# Encode the input as state vectors.
states_value = encoder_model.predict(input_seq)
# Generate empty target sequence of length 1.
target_seq = np.zeros((1, 1, len(chars_univ)))
# Populate the first character of target sequence with the start character.
target_seq[0, 0, chars_univ_idx['\n']] = 1.
# Sampling loop for a batch of sequences
# (to simplify, here we assume a batch of size 1).
stop_condition = False
decoded_sentence = ''
while not stop_condition:
output_tokens, h, c = decoder_model.predict(
[target_seq] + states_value)
# Sample a token
sampled_token_index = np.argmax(output_tokens[0, -1, :])
sampled_char = reverse_chars_idx[sampled_token_index]
decoded_sentence += sampled_char
# Exit condition: either hit max length
# or find stop character.
if (sampled_char == '\n' or
len(decoded_sentence) > MAX_TWEET_LENGTH):
stop_condition = True
# Update the target sequence (of length 1).
target_seq = np.zeros((1, 1, len(chars_univ)))
target_seq[0, 0, sampled_token_index] = 1.
# Update states
states_value = [h, c]
return decoded_sentence
def emoji_to_oh(emoji, emoji_idx):
emoji_arr = np.zeros(shape=(1, 1, len(emoji_idx)))
emoji_arr[0, 0, emoji_idx[emoji]] = 1
return emoji_arr
tweets_train.iloc[3].loc['emoji']
# +
# make one hot vector for emoji input
emoji = tweets_train.iloc[3].loc['emoji']
print (emoji)
#emoji_arr = np.zeros(shape=(1, 1, len(emoji_idx)))
#emoji_arr[0, 0, emoji_idx[emoji]] = 1
decode_sequence (emoji_to_oh(emoji, emoji_idx))
# -
for i in range (100):
emoji = tweets_train.iloc[i].loc['emoji']
generated_tweet = decode_sequence (emoji_to_oh(emoji, emoji_idx))
print (emoji, generated_tweet)
|
5 - seq2seq tweet generation model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pyportfolioopt
# language: python
# name: pyportfolioopt
# ---
# # Advanced MVO - custom objectives
#
# PyPortfolioOpt has implemented some of the most common objective functions (e.g `min_volatility`, `max_sharpe`, `max_quadratic_utility`, `efficient_risk`, `efficient_return`). However, sometimes yoy may have an idea for a different objective function.
#
# In this cookbook recipe, we cover:
#
# - Mininimising transaction costs
# - Custom convex objectives
# - Custom nonconvex objectives
#
# ## Acquiring data
#
# As discussed in the previous notebook, assets are an exogenous input (i.e you must come up with a list of tickers). We will use `yfinance` to download data for thesee tickers
import yfinance as yf
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
tickers = ["BLK", "BAC", "AAPL", "TM", "WMT",
"JD", "INTU", "MA", "UL", "CVS",
"DIS", "AMD", "NVDA", "PBI", "TGT"]
ohlc = yf.download(tickers, period="max")
prices = ohlc["Adj Close"]
prices.tail()
# ## Expected returns and risk models
#
# In this notebook, we will use James-Stein shrinkage and semicovariance (which only penalises downside risk).
import pypfopt
pypfopt.__version__
# +
from pypfopt import risk_models, expected_returns
from pypfopt import plotting
mu = expected_returns.capm_return(prices)
S = risk_models.semicovariance(prices)
# -
mu.plot.barh(figsize=(10,5));
plotting.plot_covariance(S, plot_correlation=True);
# ## Min volatility with a transaction cost objective
#
# Let's say that you already have a portfolio, and want to now optimise it. It could be quite expensive to completely reallocate, so you may want to take into account transaction costs. PyPortfolioOpt provides a simple objective to account for this.
#
# Note: this objective will not play nicely with `max_sharpe`.
# Pretend that you started with a default-weight allocation
initial_weights = np.array([1/len(tickers)] * len(tickers))
# +
from pypfopt import EfficientFrontier, objective_functions
ef = EfficientFrontier(mu, S)
# 1% broker commission
ef.add_objective(objective_functions.transaction_cost, w_prev=initial_weights, k=0.01)
ef.min_volatility()
weights = ef.clean_weights()
weights
# -
# Notice that many of the weights are 0.06667, i.e your original equal weight. In fact, the only change has been an allocation of AMD's weight to JD. If we lower the cost `k`, the allocation will change more:
ef = EfficientFrontier(mu, S)
ef.add_objective(objective_functions.transaction_cost, w_prev=initial_weights, k=0.001)
ef.min_volatility()
weights = ef.clean_weights()
weights
# The optimiser seems to really like JD. The reason for this is that it is highly anticorrelated to other assets (notice the dark column in the covariance plot). Hence, historically, it adds a lot of diversification. But it is dangerous to place too much emphasis on what happened in the past, so we may want to limit the asset weights.
#
# In addition, we notice that 4 stocks have now been allocated zero weight, which may be undesirable. Both of these problems can be fixed by adding an [L2 regularisation objective](https://pyportfolioopt.readthedocs.io/en/latest/EfficientFrontier.html#more-on-l2-regularisation).
ef = EfficientFrontier(mu, S)
ef.add_objective(objective_functions.transaction_cost, w_prev=initial_weights, k=0.001)
ef.add_objective(objective_functions.L2_reg)
ef.min_volatility()
weights = ef.clean_weights()
weights
# This has had too much of an evening-out effect. After all, if the resulting allocation is going to be so close to equal weights, we may as well stick with our initial allocation. We can reduce the strength of the L2 regularisation by reducing `gamma`:
ef = EfficientFrontier(mu, S)
ef.add_objective(objective_functions.transaction_cost, w_prev=initial_weights, k=0.001)
ef.add_objective(objective_functions.L2_reg, gamma=0.05) # default is 1
ef.min_volatility()
weights = ef.clean_weights()
weights
ef.portfolio_performance(verbose=True);
# This portfolio is now reasonably balanced, but also puts significantly more weight on JD.
pd.Series(weights).plot.pie(figsize=(10,10));
# ## Custom convex objectives
#
# PyPortfolioOpt comes with the following built-in objective functions, as of v1.2.1:
#
# - Portfolio variance (i.e square of volatility)
# - Portfolio return
# - Sharpe ratio
# - L2 regularisation (minimising this reduces nonzero weights)
# - Quadratic utility
# - Transaction cost model (a simple one)
#
# However, you may want have a different objective. If this new objective is **convex**, you can optimise a portfolio with the full benefit of PyPortfolioOpt's modular syntax, for example adding other constraints and objectives.
#
# To demonstrate this, we will minimise the **logarithmic-barrier** function suggested in the paper 60 Years of Portfolio Optimisation, by Kolm et al (2014):
#
# $$f(w, S, k) = w^T S w - k \sum_{i=1}^N \ln w$$
#
# We must first convert this mathematical objective into the language of cvxpy. Cvxpy is a powerful modelling language for convex optimisation problems. It is clean and easy to use, the only caveat is that objectives must be expressed with `cvxpy` functions, a list of which can be found [here](https://www.cvxpy.org/tutorial/functions/index.html).
# +
import cvxpy as cp
# Note: functions are minimised. If you want to maximise an objective, stick a minus sign in it.
def logarithmic_barrier_objective(w, cov_matrix, k=0.1):
log_sum = cp.sum(cp.log(w))
var = cp.quad_form(w, cov_matrix)
return var - k * log_sum
# -
# Once we have written the objective function, we can just use the `ef.convex_objective()` to minimise the objective.
ef = EfficientFrontier(mu, S, weight_bounds=(0.01, 0.2))
ef.convex_objective(logarithmic_barrier_objective, cov_matrix=S, k=0.001)
weights = ef.clean_weights()
weights
ef.portfolio_performance(verbose=True);
# This is compatible with all the constraints discussed in the previous recipe. Let's say that we want to limit JD's weight to 15%.
ef = EfficientFrontier(mu, S, weight_bounds=(0.01, 0.2))
jd_index = ef.tickers.index("JD") # get the index of JD
ef.add_constraint(lambda w: w[jd_index] <= 0.15)
ef.convex_objective(logarithmic_barrier_objective, cov_matrix=S, k=0.001)
weights = ef.clean_weights()
weights
# ## Custom nonconvex objectives
#
# In some cases, you may be trying to optimise for nonconvex objectives. Optimisation in general is a very hard problem, so please be aware that you may have mixed results in that case. Convex problems, on the other hand, are well understood and can be solved with nice theoretical guarantees.
#
# PyPortfolioOpt does offer some functionality for nonconvex optimisation, but it is not really encouraged. In particular, nonconvex optimisation is not compatible with PyPortfolioOpt's modular constraints API.
#
# As an example, we will use the Deviation Risk Parity objective from Kolm et al (2014). Because we are not using a convex solver, we don't have to define it using `cvxpy` functions.
def deviation_risk_parity(w, cov_matrix):
diff = w * np.dot(cov_matrix, w) - (w * np.dot(cov_matrix, w)).reshape(-1, 1)
return (diff ** 2).sum().sum()
ef = EfficientFrontier(mu, S, weight_bounds=(0.01, 0.12))
ef.nonconvex_objective(deviation_risk_parity, ef.cov_matrix)
weights = ef.clean_weights()
weights
# However, let's say we now want to enforce that JD has a weight of 10%. In the convex case, this would be as simple as:
#
# ```python
# ef.add_objective(lambda w: w[jd_index] == 0.10)
# ```
#
# But unfortunately, scipy does not allow for such intuitive syntax. You will need to rearrange your constraints to make them either `=0` or `<= 0`.
#
# ```python
# constraints = [
# # First constraint
# {"type": "eq", # equality constraint,
# "fun": lambda w: w[1] - 0.2}, # the equality functions are assumed to = 0
#
# # Second constraint
# {"type": "ineq", # inequality constraint
# "fun": lambda w: w[0] - 0.5} # inequality functions <= 0
# ]
# ```
#
# For more information, you can consult the [scipy docs](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html), but they aren't very helpful.
# +
ef = EfficientFrontier(mu, S, weight_bounds=(0.01, 0.12))
ef.nonconvex_objective(
deviation_risk_parity,
objective_args=S,
weights_sum_to_one=True,
constraints=[
{"type": "eq", "fun": lambda w: w[jd_index] - 0.10},
],
)
weights = ef.clean_weights()
weights
# -
# ## More examples of nonconvex objectives
#
# The scipy format is not intuitive and is hard to explain, so here are a bunch of examples (adapted from the tests). Some of these are actually convex, so you should use `convex_objective` instead.
# +
# Another example of deviation risk parity
def deviation_risk_parity(w, cov_matrix):
n = cov_matrix.shape[0]
rp = (w * (cov_matrix @ w)) / cp.quad_form(w, cov_matrix)
return cp.sum_squares(rp - 1 / n).value
ef = EfficientFrontier(mu, S)
ef.nonconvex_objective(deviation_risk_parity, ef.cov_matrix)
weights = ef.clean_weights()
weights
# -
# Deviation risk parity with weight bound on the first asset
ef = EfficientFrontier(mu, S)
ef.nonconvex_objective(deviation_risk_parity,
ef.cov_matrix,
constraints=[{"type":"eq", "fun":lambda w: w[0] - 0.1}])
weights = ef.clean_weights()
weights
# +
# Market-neutral efficient risk.
# Please use ef.efficient_risk() for anything serious.
target_risk = 0.19
ef = EfficientFrontier(mu, S, weight_bounds=(None, None))
# Weights sum to zero
weight_constr = {"type": "eq", "fun": lambda w: np.sum(w)}
# Portfolio vol less than target vol
risk_constr = {
"type": "eq",
"fun": lambda w: target_risk ** 2 - np.dot(w.T, np.dot(ef.cov_matrix, w)),
}
constraints = [weight_constr, risk_constr]
ef.nonconvex_objective(
lambda w, mu: -w.T.dot(mu), # min negative return i.e max return
objective_args=(ef.expected_returns),
weights_sum_to_one=False,
constraints=constraints,
)
weights = ef.clean_weights()
weights
# +
# Utility objective - you could actually use ef.max_quadratic_utility
ef = EfficientFrontier(mu, S)
def utility_obj(weights, mu, cov_matrix, k=1):
return -weights.dot(mu) + k * np.dot(weights.T, np.dot(cov_matrix, weights))
ef.nonconvex_objective(
utility_obj,
objective_args=(ef.expected_returns, ef.cov_matrix, 1)
# default is for weights to sum to 1
)
weights = ef.clean_weights()
weights
# -
ef.weights.sum()
# +
# Kelly objective with weight bounds on zeroth asset
def kelly_objective(w, e_returns, cov_matrix, k=3):
variance = np.dot(w.T, np.dot(cov_matrix, w))
objective = variance * 0.5 * k - np.dot(w, e_returns)
return objective
lower_bounds, upper_bounds = 0.01, 0.3
ef = EfficientFrontier(mu, S)
ef.nonconvex_objective(
kelly_objective,
objective_args=(ef.expected_returns, ef.cov_matrix, 1000),
constraints=[
{"type": "eq", "fun": lambda w: np.sum(w) - 1},
{"type": "ineq", "fun": lambda w: w[0] - lower_bounds},
{"type": "ineq", "fun": lambda w: upper_bounds - w[0]},
],
)
weights = ef.clean_weights()
weights
# -
|
cookbook/3-Advanced-Mean-Variance-Optimisation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pokemon
# ### Introduction:
#
# This time you will create the data.
#
#
#
# ### Step 1. Import the necessary libraries
import pandas as pd
# ### Step 2. Create a data dictionary
raw_data = {"name": ['Bulbasaur', 'Charmander','Squirtle','Caterpie'],
"evolution": ['Ivysaur','Charmeleon','Wartortle','Metapod'],
"type": ['grass', 'fire', 'water', 'bug'],
"hp": [45, 39, 44, 45],
"pokedex": ['yes', 'no','yes','no']
}
# ### Step 3. Assign it to a variable called
pokemon = pd.DataFrame(raw_data)
pokemon.head()
# ### Step 4. Ops...it seems the DataFrame columns are in alphabetical order. Place the order of the columns as name, type, hp, evolution, pokedex
pokemon = pokemon[['name', 'type', 'hp', 'evolution','pokedex']]
pokemon
# ### Step 5. Add another column called place, and insert what you have in mind.
pokemon['place'] = ['park','street','lake','forest']
pokemon
# ### Step 6. Present the type of each column
pokemon.dtypes
# ### BONUS: Create your own question and answer it.
|
08_Creating_Series_and_DataFrames/Pokemon/Exercises-with-solutions-and-code.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Decision boundary of label propagation versus SVM on the Iris dataset
#
#
# Comparison for decision boundary generated on iris dataset
# between Label Propagation and SVM.
#
# This demonstrates Label Propagation learning a good boundary
# even with a small amount of labeled data.
#
#
#
# +
print(__doc__)
# Authors: <NAME> <<EMAIL>>
# License: BSD
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn import svm
from sklearn.semi_supervised import label_propagation
rng = np.random.RandomState(0)
iris = datasets.load_iris()
X = iris.data[:, :2]
y = iris.target
# step size in the mesh
h = .02
y_30 = np.copy(y)
y_30[rng.rand(len(y)) < 0.3] = -1
y_50 = np.copy(y)
y_50[rng.rand(len(y)) < 0.5] = -1
# we create an instance of SVM and fit out data. We do not scale our
# data since we want to plot the support vectors
ls30 = (label_propagation.LabelSpreading().fit(X, y_30),
y_30)
ls50 = (label_propagation.LabelSpreading().fit(X, y_50),
y_50)
ls100 = (label_propagation.LabelSpreading().fit(X, y), y)
rbf_svc = (svm.SVC(kernel='rbf').fit(X, y), y)
# create a mesh to plot in
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
# title for the plots
titles = ['Label Spreading 30% data',
'Label Spreading 50% data',
'Label Spreading 100% data',
'SVC with rbf kernel']
color_map = {-1: (1, 1, 1), 0: (0, 0, .9), 1: (1, 0, 0), 2: (.8, .6, 0)}
for i, (clf, y_train) in enumerate((ls30, ls50, ls100, rbf_svc)):
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
plt.subplot(2, 2, i + 1)
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired)
plt.axis('off')
# Plot also the training points
colors = [color_map[y] for y in y_train]
plt.scatter(X[:, 0], X[:, 1], c=colors, edgecolors='black')
plt.title(titles[i])
plt.suptitle("Unlabeled points are colored white", y=0.1)
plt.show()
|
lab13/semi_supervised/plot_label_propagation_versus_svm_iris.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import nltk
message = "Hello, My name is Akash"
tokens = nltk.word_tokenize(message)
tokens
para = "Hello!, My name is Akash. I work at IDeaS"
para_tokens = nltk.sent_tokenize(para)
para_tokens
intro = "Hi Akash, How are you? It is raining here. The day temperature has significantly dropped. I am going to play STALKER at night. "
from nltk.tokenize import sent_tokenize
print(sent_tokenize(intro))
# +
#download_shell()
# -
print(nltk.pos_tag(tokens)) #nnp - nouns - vbz - verbs
print(nltk.ne_chunk(nltk.pos_tag(tokens)))
from nltk.tokenize import word_tokenize
print(word_tokenize(intro)) #treats punctuation as a word
for i in word_tokenize(intro):
print(i)
from nltk.corpus import stopwords
stop_words = stopwords.words('english')
stop_words
words = word_tokenize(intro)
filtered_words = []
for word in words:
if word not in stop_words:
filtered_words.append(word)
filtered_words
len(words)
len(filtered_words)
# +
#8 stop words removed
# +
#stop words do not matter , so we remove them in pre-processing.
# -
from nltk.stem import PorterStemmer #use stem to get parent word, used in search engines
ps = PorterStemmer()
for word in filtered_words:
print(ps.stem(word))
# +
#we could see it doesnt work well here.
# -
ps.stem('asking')
ps.stem('asks')
# +
#lemmetizing - similar word
# -
from nltk.stem import WordNetLemmatizer
lemmatizer= WordNetLemmatizer()
download_shell()
lemmatizer.lemmatize('better') #default is pos = n (noun)
lemmatizer.lemmatize('better',pos = 'a') #adjective
lemmatizer.lemmatize('runs',pos = 'v') #verb
lemmatizer.lemmatize('worse',pos = 'a')
# +
#n NOUN
#v VERB
#a ADJECTIVE
#s ADJECTIVE SATELLITE
#r ADVERB
# -
from nltk.corpus import state_union
speech = state_union.raw("2002-GWBush.txt") #read file as single string
tok = sent_tokenize(speech)
tok[5:15]
from nltk.corpus import wordnet
syns = wordnet.synsets("program")
print(syns)
syns[0].name()
syns[0].lemmas() #plan is chosen as most appropriate lemma
syns[0].lemmas()[0].name()
syns[0].definition()
syns[0].examples()
synonyms = []
antonyms = []
# +
for syn in wordnet.synsets("good"):
for l in syn.lemmas():
synonyms.append(l.name())
if l.antonyms():
antonyms.append(l.antonyms()[0].name())
# -
a = []
if a:
print("not empty")
else:
print("empty")
print(set(synonyms))
print(set(antonyms))
wordnet.synsets("trunk")
wordnet.synsets("trunk")[0].lemmas()
wordnet.synsets("trunk")[0].definition()
wordnet.synsets("trunk")[1].lemmas()
wordnet.synsets("trunk")[1].definition()
wordnet.synsets("trunk")[2].lemmas()
wordnet.synsets('boat')
wordnet.synsets('ship')
w1 = wordnet.synset("boat.n.01")
w1.definition()
w2 = wordnet.synset("ship.n.01")
w2.definition()
w1.wup_similarity(w2) #91 percent similar words
w1 = wordnet.synset("ship.n.01")
w2 = wordnet.synset("car.n.01")
w1.wup_similarity(w2) #69 percent similar words
w1 = wordnet.synset("ship.n.01")
w2 = wordnet.synset("cow.n.01")
w1.wup_similarity(w2) #27 percent similar words
message = "Hi, is it raining or is it cloudy?"
tokens = word_tokenize(message)
import nltk
fd = nltk.FreqDist(tokens)
fd
fd.most_common(3)
fd.plot()
|
.ipynb_checkpoints/nltk sendtex-checkpoint.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.3
# language: julia
# name: julia-1.0
# ---
using DifferentialEquations
using Random
using DataFrames
using Distributions
function continuous_time_SIR(β,γ,N,S0,I0,R0,tf)
#Temp Variables for Time, Susceptible Individuals, Infected Individuals, Recovery
t = 0
S = S0
I = I0
R = R0
#Array to store SIR and Time
ta= Float64[]
Sa= Float64[]
Ia= Float64[]
Ra= Float64[]
while t < tf
push!(ta,t)
push!(Sa,S)
push!(Ia,I)
push!(Ra,R)
pf1 = β*S*I
pf2 = γ*I
pf = pf1+pf2
dt = rand(Exponential(1/pf))
t = t+dt
if t>tf
break
end
ru = rand()
if ru<(pf1/pf)
S=S-1
I=I+1
else
I=I-1
R=R+1
end
end
results = DataFrame()
results[:time] = ta
results[:S] = Sa
results[:I] = Ia
results[:R] = Ra
return(results)
end
Random.seed!(42);
#continuous_time_SIR(Infected Rate, Recover Rate, Sample Size ,
#Initial Susceptible Individuals, Initial Infected Rate, Initial Recovery Rate, Max Time Increment)
sir_out = continuous_time_SIR(0.1/1000,0.05,1000,999,1,0,200);
head_size = 6
first(sir_out,head_size)
using StatPlots
@df sir_out plot(:time, [:S :I :R], xlabel="Time",ylabel="Number")
|
examples/epicookbook/notebooks/SimpleStochasticModels/ContinuousTimeSIR.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import parent
# +
import torch
from torch import nn
#image_A, image_B = (x[0].cuda() for x in next(zip(d1_t, d2_t)))
class RegistrationTransformer(nn.Module):
def __init__(self, size):
self.size = size
super(RegistrationTransformer, self).__init__()
feature_n = 120
self.position_embedding = nn.Parameter(
torch.randn(((size // 7) **2, 1, feature_n))
)
self.embed = nn.Conv2d(
1,
feature_n,
kernel_size=7,
padding=0,
stride=7,
).cuda()
self.decode = nn.ConvTranspose2d(
feature_n,
2,
kernel_size=7,
padding=0,
stride=7,
).cuda()
#torch.nn.init.zeros_(self.decode.weight)
#torch.nn.init.zeros_(self.decode.bias)
self.t = torch.nn.Transformer(d_model=120, dim_feedforward=256)
def forward(self, x, y):
x = self.embed_and_reshape(x)
y = self.embed_and_reshape(y)
out = self.t(y, x)
out = self.reshape_and_decode(out) / 20
return out
def embed_and_reshape(self, a):
a = self.embed(a)
a = a.reshape((a.shape[0], a.shape[1], a.shape[2] * a.shape[3]))
a = a.permute(2, 0, 1)
a = a + self.position_embedding
return a
def reshape_and_decode(self, seq):
a = seq.permute(1, 2, 0)
size = int(math.sqrt(a.shape[-1]))
a = a.reshape(a.shape[0], a.shape[1], size, size)
a = self.decode(a)
return a
#model = RegistrationTransformer(28)
#model.cuda()
#model(image_A, image_B).shape
# +
import parent
import data
import networks
import network_wrappers
import visualize
import train
import inverseConsistentNet
import numpy as np
import torch
import matplotlib.pyplot as plt
import random
import os
import pickle
import describe
import argparse
import math
batch_size = 128
data_size=63 * 2
d1, d2 = data.get_dataset_triangles(
"train", data_size=data_size, hollow=True, batch_size=batch_size
)
d1_t, d2_t = data.get_dataset_triangles(
"test", data_size=data_size, hollow=True, batch_size=batch_size
)
# +
lmbda = 150
random.seed(1)
torch.manual_seed(1)
torch.cuda.manual_seed(1)
np.random.seed(1)
print("=" * 50)
net = inverseConsistentNet.InverseConsistentNet(
network_wrappers.DownsampleNet(network_wrappers.FunctionFromVectorField(RegistrationTransformer(data_size // 2)), 2),
lambda x, y: torch.mean((x - y) ** 2),
lmbda,
)
input_shape = next(iter(d1))[0].size()
network_wrappers.assignIdentityMap(net, input_shape)
net.cuda()
optimizer = torch.optim.Adam(net.parameters(), lr=0.001)
net.train()
xs = []
# -
for _ in range(0,30):
y = np.array(train.train2d(net, optimizer, d1, d2, epochs=50))
xs.append(y)
x = np.concatenate(xs)
plt.title(
"Loss curve for " + type(net.regis_net).__name__ + " lambda=" + str(lmbda)
)
plt.plot(x[:, :3])
plt.savefig(describe.run_dir + f"loss.png")
plt.clf()
plt.title("Log # pixels with negative Jacobian per epoch")
plt.plot(x[:, 3])
# random.seed(1)
plt.savefig(describe.run_dir + f"lossj.png")
plt.clf()
with open(describe.run_dir + "loss.pickle", "wb") as f:
pickle.dump(x, f)
# torch.manual_seed(1)
# torch.cuda.manual_seed(1)
# np.random.seed(1)
image_A, image_B = (x[0].cuda() for x in next(zip(d1_t, d2_t)))
for N in range(3):
visualize.visualizeRegistration(
net,
image_A,
image_B,
N,
describe.run_dir + f"epoch{_:03}" + "case" + str(N) + ".png",
)
# +
random.seed(1)
torch.manual_seed(1)
torch.cuda.manual_seed(1)
np.random.seed(1)
image_A, image_B = (x[0].cuda() for x in next(zip(d1_t, d2_t)))
os.mkdir(describe.run_dir + "final/")
for N in range(30):
visualize.visualizeRegistrationCompact(net, image_A, image_B, N)
plt.savefig(describe.run_dir + f"final/{N}.png")
plt.clf()
torch.save(net.state_dict(), describe.run_dir + "network.trch")
torch.save(optimizer.state_dict(), describe.run_dir + "opt.trch")
# -
plt.imshow(net.regis_net.net.decode.weight.cpu().detach()[27][1])
model.t.d_model
# +
# model.t.forward?
# -
torch.Parameter
torch.randn((
# +
# nn.Transformer?
# -
|
notebooks/TransformerRegister.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import tensorflow as tf
from tf_explain.core import GradCAM,GradientsInputs
import glob
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# ### Load images
validation = pd.read_csv('pneumonia-validation-set.csv')
validation.tail()
neg_image_paths = validation['image_path'][:10].to_list()
pos_image_paths = validation['image_path'][-10:].to_list()
image_paths = neg_image_paths + pos_image_paths
len(image_paths)
def load_image(image_path):
# read the image from disk, decode it, resize it, and scale the
# pixels intensities to the range [0, 1]
image = tf.io.read_file(image_path)
image = tf.image.decode_png(image, channels=1)
image = tf.cast(image,dtype=tf.float32)
image = image / 255.0
return (image)
images = []
for i in range(len(image_paths)):
image = load_image(image_paths[i])
images.append(image)
plt.imshow(images[0].numpy(),cmap = 'gray')
plt.imshow(images[-1].numpy(),cmap = 'gray')
# ### GradCAM Implementation
# Load Model
model = tf.keras.models.load_model('pneu-model/')
# +
# model.summary() # conv5_block3_3_conv
# -
labels = [0] * 10 + [1] * 10
labels
grad_cam
from tf_explain.callbacks.grad_cam import GradCAMCallback
callbacks = [
GradCAMCallback(
validation_data=(images[:10], labels[:10]),
layer_name="conv5_block3_3_conv",
class_index=0,
)
]
model.evaluate(callbacks = )
|
004-GradCAM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.model_selection import train_test_split
# +
df = pd.read_csv("data/train_with_translations_clean.csv")
df = df[["clean_txt", "clean_txt_T1", "Intencion"]]
train_index, test_index = train_test_split(list(df.index), random_state=13571113)
train_df = df.iloc[train_index].reset_index().drop(columns="index")
test_df = df.iloc[test_index].reset_index().drop(columns="index")
thr = train_df["Intencion"].value_counts().values[0] // 2
cats_to_popu = [ k for k, v in dict(train_df["Intencion"].value_counts()).items() if v < thr]
train_da_df = train_df[train_df["Intencion"].isin(cats_to_popu)][["clean_txt_T1", "Intencion"]]
train_da_df.columns = ["clean_txt", "Intencion"]
train_df = train_df[["clean_txt", "Intencion"]]
train_df = pd.concat([train_df, train_da_df])
train_df.columns = ["Pregunta", "Intencion"]
test_df = test_df[["clean_txt", "Intencion"]]
test_df.columns = ["Pregunta", "Intencion"]
# -
test_df.to_csv("exp1/valid.csv", index=False)
train_df.to_csv("exp1/train.csv", index=False)
encode_cat = json.load(open(mapping_dict, "r"))
train_dataset['ENCODE_CAT'] = train_dataset['Intencion'].apply(lambda x: encode_cat(x))
test_dataset['ENCODE_CAT'] = test_dataset['Intencion'].apply(lambda x: encode_cat(x))
# ## Translate es --> ar --> es
import pandas as pd
from tqdm.auto import tqdm
from googletrans import Translator
translator = Translator()
df = pd.read_csv("data/train_with_translations_clean.csv")
df_test = pd.read_csv("data/test_with_translations_clean.csv")
sentences_es = list(df.Pregunta.values)
print(f'Amount of sentences {len(sentences_es)}')
sentences_es[:2]
sentences_es_test = list(df_test.Pregunta.values)
print(f'Amount of sentences {len(sentences_es_test)}')
sentences_es_test[:2]
# +
translations_ar = []
for sent in tqdm(sentences_es):
translation = translator.translate(sent, src="es", dest="ar").text
translations_ar.append(translation)
print(f'Amount sentences en: {len(translations_ar)}')
translations_es_back = []
for sent in tqdm(translations_ar):
translation = translator.translate(sent, src="ar", dest="es").text
translations_es_back.append(translation)
print(f'Amount sentences en: {len(translations_es_back)}')
df["Pregunta_T4_ar"] = translations_es_back
translations_ar = []
for sent in tqdm(sentences_es_test):
translation = translator.translate(sent, src="es", dest="ar").text
translations_ar.append(translation)
print(f'Amount sentences en: {len(translations_ar)}')
translations_es_back = []
for sent in tqdm(translations_ar):
translation = translator.translate(sent, src="ar", dest="es").text
translations_es_back.append(translation)
print(f'Amount sentences en: {len(translations_es_back)}')
df_test["Pregunta_T4_ar"] = translations_es_back
# -
# !pip install joblib
# +
from textacy.preprocess import preprocess_text as textacy_preprocess
import re
EMOJI_PATTERN = re.compile("["
u"\U0001F600-\U0001F64F" # emoticons
u"\U0001F300-\U0001F5FF" # symbols & pictographs
u"\U0001F680-\U0001F6FF" # transport & map symbols
u"\U0001F1E0-\U0001F1FF" # flags (iOS)
"]+", flags=re.UNICODE)
def remove_stopwords(text):
"""Remove stop words from list of tokenized words."""
new_words = [word for word in text.split() if word not in STOPWORDS_SET]
return ' '.join(new_words)
def remove_emoticons(text):
"""Remove emoticos."""
return EMOJI_PATTERN.sub('', text)
def to_lowercase(text):
"""To lowercase."""
return text.lower()
def preprocess_text(text, fix_unicode=True, lowercase=True,
no_urls=True, no_emails=True,
no_phone_numbers=True,
no_numbers=True, no_currency_symbols=True,
no_punct=True, no_accents=True):
"""Preprocess text."""
clean_text = ""
if text:
clean_text = remove_emoticons(text)
if lowercase:
clean_text = to_lowercase(clean_text)
# clean_text = remove_stopwords(clean_text)
clean_text = textacy_preprocess(clean_text, fix_unicode=fix_unicode,
lowercase=lowercase,
no_urls=no_urls, no_emails=no_emails,
no_phone_numbers=no_phone_numbers,
no_numbers=no_numbers,
no_currency_symbols=no_currency_symbols,
no_punct=no_punct,
no_accents=no_accents)
return clean_text
# -
df["clean_txt_T4_ar"] = df["Pregunta_T4_ar"].apply(lambda x: preprocess_text(x))
df_test["clean_txt_T4_ar"] = df_test["Pregunta_T4_ar"].apply(lambda x: preprocess_text(x))
df.head()
df_ = pd.read_csv("data/train_with_translations_clean.csv")
df_test_ = pd.read_csv("data/test_with_translations_clean.csv")
df_["clean_txt_T4_ar"] = df["clean_txt_T4_ar"]
df_test_["clean_txt_T4_ar"] = df_test["clean_txt_T4_ar"]
df_["Pregunta_T4_ar"] = df["Pregunta_T4_ar"]
df_test_["Pregunta_T4_ar"] = df_test["Pregunta_T4_ar"]
df_.to_csv("data/train_with_translations_clean.csv", index=False)
df_test_.to_csv("data/test_with_translations_clean.csv", index=False)
import seaborn as sns
columns = ["clean_txt_T1", "clean_txt_T2_fr", "clean_txt_T3_pt", "clean_txt_T4_ar"]
df = df_[columns + ["clean_txt", "Intencion"]]
df0 = df[["clean_txt", "Intencion"]].copy()
for col in columns:
thr = df0["Intencion"].value_counts().values[0] // 2
print(thr)
cats_to_popu = [ k for k, v in dict(df0["Intencion"].value_counts()).items() if v < thr]
print(f'Amount categories to populate : {len(cats_to_popu)}')
df1 = df[df["Intencion"].isin(cats_to_popu)][[col, "Intencion"]].copy()
df1.columns = ["clean_txt", "Intencion"]
df0 = pd.concat([df0, df1])
sns.lineplot(data=pd.DataFrame(list(df0.Intencion.value_counts().values)))
|
notebooks/EDA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="8b181d27-001" colab_type="text"
# #1. Install Dependencies
# First install the libraries needed to execute recipes, this only needs to be done once, then click play.
#
# + id="8b181d27-002" colab_type="code"
# !pip install git+https://github.com/google/starthinker
# + [markdown] id="8b181d27-003" colab_type="text"
# #2. Get Cloud Project ID
# To run this recipe [requires a Google Cloud Project](https://github.com/google/starthinker/blob/master/tutorials/cloud_project.md), this only needs to be done once, then click play.
#
# + id="8b181d27-004" colab_type="code"
CLOUD_PROJECT = 'PASTE PROJECT ID HERE'
print("Cloud Project Set To: %s" % CLOUD_PROJECT)
# + [markdown] id="8b181d27-005" colab_type="text"
# #3. Get Client Credentials
# To read and write to various endpoints requires [downloading client credentials](https://github.com/google/starthinker/blob/master/tutorials/cloud_client_installed.md), this only needs to be done once, then click play.
#
# + id="8b181d27-006" colab_type="code"
CLIENT_CREDENTIALS = 'PASTE CREDENTIALS HERE'
print("Client Credentials Set To: %s" % CLIENT_CREDENTIALS)
# + [markdown] id="8b181d27-007" colab_type="text"
# #4. Enter CM360 Segmentology Parameters
# CM360 funnel analysis using Census data.
# 1. Wait for <b>BigQuery->->->Census_Join</b> to be created.
# 1. Join the <a hre='https://groups.google.com/d/forum/starthinker-assets' target='_blank'>StarThinker Assets Group</a> to access the following assets
# 1. Copy <a href='https://datastudio.google.com/c/u/0/reporting/3673497b-f36f-4448-8fb9-3e05ea51842f/' target='_blank'>CM360 Segmentology Sample</a>. Leave the Data Source as is, you will change it in the next step.
# 1. Click Edit Connection, and change to <b>BigQuery->->->Census_Join</b>.
# 1. Or give these intructions to the client.
# Modify the values below for your use case, can be done multiple times, then click play.
#
# + id="8b181d27-008" colab_type="code"
FIELDS = {
'account': '',
'auth_read': 'user', # Credentials used for reading data.
'auth_write': 'service', # Authorization used for writing data.
'recipe_name': '', # Name of report, not needed if ID used.
'recipe_slug': '', # Name of Google BigQuery dataset to create.
'advertisers': [], # Comma delimited list of CM360 advertiser ids.
}
print("Parameters Set To: %s" % FIELDS)
# + [markdown] id="8b181d27-009" colab_type="text"
# #5. Execute CM360 Segmentology
# This does NOT need to be modified unless you are changing the recipe, click play.
#
# + id="8b181d27-010" colab_type="code"
from starthinker.util.project import project
from starthinker.script.parse import json_set_fields
USER_CREDENTIALS = '/content/user.json'
TASKS = [
{
'dataset': {
'description': 'Create a dataset for bigquery tables.',
'hour': [
4
],
'auth': 'user',
'dataset': {'field': {'name': 'recipe_slug','kind': 'string','description': 'Place where tables will be created in BigQuery.'}}
}
},
{
'bigquery': {
'auth': 'user',
'function': 'Pearson Significance Test',
'to': {
'dataset': {'field': {'name': 'recipe_slug','kind': 'string','order': 4,'default': '','description': 'Name of Google BigQuery dataset to create.'}}
}
}
},
{
'google_api': {
'auth': 'user',
'api': 'dfareporting',
'version': 'v3.4',
'function': 'accounts.get',
'kwargs': {
'id': {'field': {'name': 'account','kind': 'integer','order': 5,'default': '','description': 'Campaign Manager Account ID'}},
'fields': 'id,name'
},
'results': {
'bigquery': {
'auth': 'user',
'dataset': {'field': {'name': 'recipe_slug','kind': 'string','order': 4,'default': '','description': 'Name of Google BigQuery dataset to create.'}},
'table': 'CM360_Account'
}
}
}
},
{
'dcm': {
'auth': 'user',
'report': {
'filters': {
'dfa:advertiser': {
'values': {'field': {'name': 'advertisers','kind': 'integer_list','order': 6,'default': [],'description': 'Comma delimited list of CM360 advertiser ids.'}}
}
},
'account': {'field': {'name': 'account','kind': 'string','order': 5,'default': '','description': 'Campaign Manager Account ID'}},
'body': {
'name': {'field': {'name': 'recipe_name','kind': 'string','prefix': 'Segmentology ','description': 'The report name.','default': ''}},
'criteria': {
'dateRange': {
'kind': 'dfareporting#dateRange',
'relativeDateRange': 'LAST_90_DAYS'
},
'dimensions': [
{
'kind': 'dfareporting#sortedDimension',
'name': 'dfa:advertiserId'
},
{
'kind': 'dfareporting#sortedDimension',
'name': 'dfa:advertiser'
},
{
'kind': 'dfareporting#sortedDimension',
'name': 'dfa:campaignId'
},
{
'kind': 'dfareporting#sortedDimension',
'name': 'dfa:campaign'
},
{
'kind': 'dfareporting#sortedDimension',
'name': 'dfa:placementId'
},
{
'kind': 'dfareporting#sortedDimension',
'name': 'dfa:placement'
},
{
'kind': 'dfareporting#sortedDimension',
'name': 'dfa:zipCode'
}
],
'metricNames': [
'dfa:impressions',
'dfa:clicks',
'dfa:totalConversions'
]
},
'type': 'STANDARD',
'delivery': {
'emailOwner': False
},
'format': 'CSV'
}
}
}
},
{
'dcm': {
'auth': 'user',
'report': {
'account': {'field': {'name': 'account','kind': 'string','default': ''}},
'name': {'field': {'name': 'recipe_name','kind': 'string','order': 3,'prefix': 'Segmentology ','default': '','description': 'Name of report, not needed if ID used.'}}
},
'out': {
'bigquery': {
'auth': 'user',
'dataset': {'field': {'name': 'recipe_slug','kind': 'string','order': 4,'default': '','description': 'Name of Google BigQuery dataset to create.'}},
'table': 'CM360_KPI',
'header': True
}
}
}
},
{
'bigquery': {
'auth': 'user',
'from': {
'query': 'SELECT Id AS Partner_Id, Name AS Partner, Advertiser_Id, Advertiser, Campaign_Id, Campaign, Zip_Postal_Code AS Zip, SAFE_DIVIDE(Impressions, SUM(Impressions) OVER(PARTITION BY Advertiser_Id)) AS Impression_Percent, SAFE_DIVIDE(Clicks, Impressions) AS Click_Percent, SAFE_DIVIDE(Total_Conversions, Impressions) AS Conversion_Percent, Impressions AS Impressions FROM `{dataset}.CM360_KPI` CROSS JOIN `{dataset}.CM360_Account` ',
'parameters': {
'dataset': {'field': {'name': 'recipe_slug','kind': 'string','description': 'Place where tables will be created in BigQuery.'}}
},
'legacy': False
},
'to': {
'dataset': {'field': {'name': 'recipe_slug','kind': 'string','description': 'Place where tables will be written in BigQuery.'}},
'view': 'CM360_KPI_Normalized'
}
}
},
{
'census': {
'auth': 'user',
'normalize': {
'census_geography': 'zip_codes',
'census_year': '2018',
'census_span': '5yr'
},
'to': {
'dataset': {'field': {'name': 'recipe_slug','kind': 'string','order': 4,'default': '','description': 'Name of Google BigQuery dataset to create.'}},
'type': 'view'
}
}
},
{
'census': {
'auth': 'user',
'correlate': {
'join': 'Zip',
'pass': [
'Partner_Id',
'Partner',
'Advertiser_Id',
'Advertiser',
'Campaign_Id',
'Campaign'
],
'sum': [
'Impressions'
],
'correlate': [
'Impression_Percent',
'Click_Percent',
'Conversion_Percent'
],
'dataset': {'field': {'name': 'recipe_slug','kind': 'string','order': 4,'default': '','description': 'Name of Google BigQuery dataset to create.'}},
'table': 'CM360_KPI_Normalized',
'significance': 80
},
'to': {
'dataset': {'field': {'name': 'recipe_slug','kind': 'string','order': 4,'default': '','description': 'Name of Google BigQuery dataset to create.'}},
'type': 'view'
}
}
}
]
json_set_fields(TASKS, FIELDS)
project.initialize(_recipe={ 'tasks':TASKS }, _project=CLOUD_PROJECT, _user=USER_CREDENTIALS, _client=CLIENT_CREDENTIALS, _verbose=True, _force=True)
project.execute(_force=True)
|
colabs/cm360_segmentology.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.chdir('../../modules/')
import pickle
from util.util import *
from util.losses import *
from util.pred_blocks import ConvEncoder, GRUDecoder
from likelihood_predictor import PlastPredictor
from vae_generator import PlastVAEGen
import torch
import torch.nn as nn
import torch.nn.functional as F
from torchsummary import summary
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.cm as cm
# +
# %load_ext autoreload
# %autoreload 2
import IPython.display as Disp
np.set_printoptions(suppress=True)
# -
# Load Data
pl_ll = pd.read_pickle('../database/vae_pl.pkl')
org_ll = pd.read_pickle('../database/vae_org.pkl')
orgpl_test = pd.read_pickle('../database/vae_orgpl.pkl').to_numpy()
orgpl_sample = pd.read_pickle('../database/vae_orgpl.pkl').sample(n=500).to_numpy()
gdb_data = pd.read_pickle('../database/gdb_vae_test_1mil.pkl').to_numpy()
gdb_test = pd.read_pickle('../database/gdb_vae_test_1mil.pkl').sample(n=500).to_numpy()
all_data = pd.concat([pl_ll, org_ll]).to_numpy()
test_data = pd.concat([pl_ll, org_ll]).sample(n=500).to_numpy()
# ### Evaluating model reconstruction
char_weights_gdb = np.load('util/char_weights_gdb.npy')
char_weights_orgpl = np.load('util/char_weights_orgpl.npy')
with open('util/char_dict.pkl', 'rb') as f:
char_dict = pickle.load(f)
with open('util/org_dict.pkl', 'rb') as f:
org_dict = pickle.load(f)
params={'MAX_LENGTH': 180,
'BATCH_SIZE': 10,
'MODEL_CLASS': 'GRUGRU',
'ARCH_SIZE': 'large',
'CHAR_DICT': char_dict,
'ORG_DICT': org_dict,
'CHAR_WEIGHTS': char_weights_orgpl}
# +
ckpt_fn = 'latest_GRUGRU_transfer_1mil.ckpt'
pvg = PlastVAEGen(params=params, name='test')
pvg.load(os.path.join('checkpoints', ckpt_fn), transfer=False, predict_property=False)
pvg.initiate(orgpl_test)
# -
train_smiles = np.array(pvg.usable_smiles)[pvg.params['TRAIN_IDXS']]
val_smiles = np.array(pvg.usable_smiles)[pvg.params['VAL_IDXS']]
X_train = pvg.encoded[pvg.params['TRAIN_IDXS'],:]
X_val = pvg.encoded[pvg.params['VAL_IDXS'],:]
def get_smiles(idx, set='train', temp=0.1, encoded=False):
if set == 'train':
encoded_smile = X_train[idx,:]
in_smile = train_smiles[idx]
elif set == 'val':
encoded_smile = X_val[idx,:]
in_smile = val_smiles[idx]
decoded_smile_for_loss = pvg.predict(encoded_smile.unsqueeze(0).numpy())[0,:]
print(decoded_smile_for_loss.shape)
decoded_smile = F.softmax(torch.tensor(decoded_smile_for_loss), dim=0).numpy()
out_smile = decode_smiles(decoded_smile, pvg.params['ORG_DICT'], temp=temp)
if encoded:
return encoded_smile.unsqueeze(0), torch.tensor(decoded_smile_for_loss).unsqueeze(0)
else:
return in_smile, out_smile
for idx in range(10):
in_smile, out_smile = get_smiles(idx, set='val', temp=0.02)
print('-----Index {}-----'.format(idx+1))
print('Target ', ''.join(in_smile))
print('Output ', out_smile)
# Almost good enough!
enc, dec = get_smiles(0, set='train', temp=0.05, encoded=True)
ce_loss(enc, dec)
# +
def enc_to_class(x):
x = x.permute(0, 2, 1)
x = x.contiguous().view(-1, x.size(2))
_, x = torch.max(x, 1)
return x
def reform_dec(x):
x = x.permute(0, 2, 1)
x = x.contiguous().view(-1, x.size(2))
return x
def loss_per_char(enc, dec, verbose=False):
x = enc_to_class(enc)
y = reform_dec(dec)
loss = 0
losses = []
for i in range(y.shape[0]):
l = -y[i,x[i]]
exp_sum = 0
for j in range(y.shape[1]):
exp_sum += np.exp(y[i,j])
l += np.log(exp_sum)
losses.append(l)
if verbose:
print(l, x[i], F.softmax(y[i,:]).numpy()[x[i]])
loss += l
loss /= pvg.params['MAX_LENGTH']
return loss, losses
loss, losses = loss_per_char(enc, dec, verbose=False)
loss
# -
# ## Notes
# ### KL_BETA Introduction
#
# This loss function is not good enough. It provides too much reward for basically random guessing (has 1/num_char chance of guessing "right" for all null characters). Need to use cross entropy. Back to where we were before experiments with v2 and bce_loss.
#
# I think repeat vector may be a culprit in the models inability to generalize. The latent space is size batch_sizexlatent_size and is repeated along the sequence dimension aka if the sequence length is 60 then 60 copies of this latent space vector are sent to the GRU. Idk how the network is supposed to learn sequence when the input sequence has identical elements. Gonna try and add a nonlinearity between latent space and input of GRU.
# ### Adding Latent Non-Linearity
# The loss is the best that I've achieved so far with, on average, a greater than 20x chance of selecting better than random for both the training set and validation set. However, the model still cannot adequately reconstruct SMILES from the latent vector.
# ```
# -----Index 1-----
# Input CC1C2CN=C1OC1=C(NC(N)=N2)N=C(O)N1
# Target CC1CCCCCCCCCC=C(CC(O)=N2)N=C(O)N1___________________________
# -----Index 2-----
# Input CC12CCCC3=C(C=NN3)C(CO1)NC2C#C
# Target CC1CCCCCCCC(CCCC))C(C#N)C2=C=O______________________________
# -----Index 3-----
# Input CC1CC(CN1)C1NC2=NCC3CCC1C23O
# Target CC1CCCCCCCCCCCCCCCCCCCCCC24C________________________________
# -----Index 4-----
# Input CC(O)C(C)(C)C1C2OC(C2O)C(C)C1C
# Target CC1CCCCCC(CC)CCC=C(C#C)C(C)C1C______________________________
# -----Index 5-----
# Input CCCNCC1OC(C#N)C(N)C2COC1O2
# Target CC1CCCCCCCC(O)C(C)CCCCCOCC1_________________________________
# -----Index 6-----
# Input CC1=COC(=N)C(NC=N)=NC=CC(C=O)=N1
# Target CC1CCCCCCCCC(NC=O)=CCCCC(C=O)=N1____________________________
# -----Index 7-----
# Input CCCC1=C(CC(N)(CO)C#C)NN=C1C#N
# Target CC1CCCCCCCCCC(C)CCCC)CC=C(C#N_______________________________
# -----Index 8-----
# Input CC1(O)COCOC1C1CNC1CC=O
# Target CC1CCCCCCCCCC=CCCCCCCCC_____________________________________
# -----Index 9-----
# Input CC1C(O)C2(C)CC1C(N)=NC(CO)(C2)C#N
# Target CC1CCCCCC(=)C1=C(C)CCC(=O)(C1)C=N___________________________
# -----Index 10-----
# Input CC12CC(CC#N)CNCC1(CO)CNC=N2
# Target CC1CCCCCCCCCCCCCC(CO)CCC=N2_________________________________
# ```
# The model is actually doing a fairly good job predicting sequence except for a small group of characters towards the beginning. It thinks every SMILE should start with `CC1CCCC...`.
# +
# Code to calculate avg train and val losses per character position (takes a really long time)
# stride = 100
# train_losses = np.zeros((pvg.params['N_TRAIN'] // stride, pvg.params['MAX_LENGTH']))
# val_losses = np.zeros((pvg.params['N_TEST'] // stride, pvg.params['MAX_LENGTH']))
# for i in range(pvg.params['N_TRAIN']):
# j = 0
# if i % stride == 0:
# enc, dec = get_smiles(i, set='train', temp=0.05, encoded=True)
# loss, losses = loss_per_char(enc, dec, verbose=False)
# train_losses[j,:] = losses
# j += 1
# for i in range(pvg.params['N_TEST']):
# j = 0
# if i % stride == 0:
# enc, dec = get_smiles(i, set='test', temp=0.05, encoded=True)
# loss, losses = loss_per_char(enc, dec, verbose=False)
# val_losses[j,:] = losses
# j += 1
# avg_train_losses = np.mean(train_losses, axis=0)
# avg_val_losses = np.mean(val_losses, axis=0)
avg_train_losses = np.load('run_data/latent_nonlin/avg_train_losses.npy')
avg_val_losses = np.load('run_data/latent_nonlin/avg_val_losses.npy')
plt.plot(avg_train_losses, label='Train')
plt.plot(avg_val_losses, label='Validation')
plt.legend(loc='best')
plt.show()
# -
# The average loss per character doesn't really show any trends that would indicate the model predicts the same starting characters every time. It must have to do with the distribution of characters at each position in the training set.
# +
# # Code to generate character count dictionary (takes a while)
# char_dist = {}
# for k in pvg.params['CHAR_DICT'].keys():
# char_dist[k] = np.zeros((pvg.params['MAX_LENGTH']+1,))
# for smile in train_smiles:
# for i, char in enumerate(smile):
# char_dist[char][i] += 1
# char_dist[char][-1] += 1
# for j in range(i, pvg.params['MAX_LENGTH']):
# char_dist['_'][j] += 1
# char_dist['_'][-1] += 1
with open('run_data/latent_nonlin/char_dist.pkl', 'rb') as f:
char_dist = pickle.load(f)
# +
# # Code to generate validation character count dictionary (takes a really long time)
# val_char_dist = {}
# for k in pvg.params['CHAR_DICT'].keys():
# val_char_dist[k] = np.zeros((pvg.params['MAX_LENGTH']+1,))
# for idx in range(pvg.params['N_TRAIN']):
# in_smile, out_smile = get_smiles(idx, set='train', temp=0.05)
# for i, char in enumerate(out_smile):
# val_char_dist[char][i] += 1
# val_char_dist[char][-1] += 1
# for j in range(i, pvg.params['MAX_LENGTH']):
# val_char_dist['_'][j] += 1
# val_char_dist['_'][-1] += 1
with open('run_data/latent_nonlin/val_char_dist.pkl', 'rb') as f:
val_char_dist = pickle.load(f)
# -
all_chars = np.zeros((30,))
all_val_chars = np.zeros((30,))
char_labels = []
i = 0
for k, v in char_dist.items():
char_labels.append(k)
all_chars[i] = v[-1]
all_val_chars[i] = val_char_dist[k][-1]
i += 1
plt.bar(range(len(all_chars)), all_chars)
plt.plot(all_val_chars, c='purple', ls=':')
plt.xticks(range(len(all_chars)), labels=char_labels)
plt.yscale('log')
plt.show()
# +
# fig, axs = plt.subplots(10, 6, figsize=(30,20))
# plt.setp(axs, xticks=range(len(all_chars)), xticklabels=char_labels)
# for i, ax in enumerate(fig.axes):
# position_chars = np.zeros((30,))
# val_pos_chars = np.zeros((30,))
# j = 0
# for k, v in char_dist.items():
# position_chars[j] = v[i]
# val_pos_chars[j] = val_char_dist[k][i]
# j += 1
# ax.bar(range(len(position_chars)), position_chars)
# ax.plot(val_pos_chars, c='purple', ls=':')
# ax.set_title('Position {}'.format(i+1))
# ax.set_ylim([0,1e6])
# fig.tight_layout()
# plt.show()
fig, axs = plt.subplots(5, 5, figsize=(30,20))
plt.setp(axs, xticks=range(len(all_chars)), xticklabels=char_labels)
for i, ax in enumerate(fig.axes):
position_chars = np.zeros((30,))
val_pos_chars = np.zeros((30,))
j = 0
for k, v in char_dist.items():
position_chars[j] = v[i+4]
val_pos_chars[j] = val_char_dist[k][i+4]
j += 1
ax.bar(range(len(position_chars)), position_chars)
ax.plot(val_pos_chars, c='purple', ls=':')
ax.set_title('Position {}'.format(i+5))
ax.set_ylim([0,1e6])
fig.tight_layout()
plt.show()
# -
# The model predictions are fit well when the data is evenly distributed (or a single peak) but struggles when one class dominates while others are still present in smaller numbers. I also now realize how many extra unnecessary padding characters I've been adding. I think the obvious first step is to add weights to each class so the loss is reduced for classes that are dominating. This will not take character position into account so I may have to later account for the distribution within specific positions. I also switched from single characters to regex patterns so, for instance, `B` and `r` are no longer considered separate characters but rather a single `Br` token.
# ### Weighing Loss by Character Token
# Adding a reduction in importance to the loss generated from the most frequent characters had a vast improvement on the model's ability to reconstruct the initial sequence of tokens.
# ```
# -----Index 1-----
# Target CCC1CC(O)C(O)(C#C)C2(CO)NC=NC12______________
# Output CCCCC(C)CC(O)(C#C)C(CCO)NC=NC12______________
# -----Index 2-----
# Target CC1=C2C3C(C1)N3C1=C2C(O)=CC(C)=C1N___________
# Output CC1CCC3=C(N1)N1C1=C3C(O)CCC(C)=C1N___________
# -----Index 3-----
# Target CC12C(O)CCCC(N=CN)C1=CNC2=O__________________
# Output CCCCC(C)CCCC(N=CN)C1=CNC2=O__________________
# -----Index 4-----
# Target CC1C2CC2(CN1C=O)C(N)C1=COC=N1________________
# Output CC1CCCCC(CC1C=O)C(N)C1=COC=N1________________
# -----Index 5-----
# Target CC#CC12C(C)CN1CC(C)(C#C)C2(C)O_______________
# Output CC1CCCCC(C)C11CC(C)(OC=)C2(C)O_______________
# -----Index 6-----
# Target CC1C2CC2(O)C(C(C)=O)=C1CNC=N_________________
# Output CC1CCCCC(O)C(C(C)=O)=C1CNC=N_________________
# -----Index 7-----
# Target CC1CN(CC(O)(CO)C#N)C2=CON=C2C1_______________
# Output CCCCCCCC(O)(CO)C#N)C2=CON=C2N1_______________
# -----Index 8-----
# Target CC1NC2CC1(C)C1CN=C(CCC(N)C2)N1_______________
# Output CC1CCC2CC(C)C1CN=C(CCC(N)N=)S1_______________
# -----Index 9-----
# Target CC(C)(OC1=NC(CO)CN1)C(C=O)C(N)=N_____________
# Output CC((O)NC1=NC(CO=CN1)C(CNO)C(N)=O_____________
# -----Index 10-----
# Target CCNC1(CC(CC=C1)=NO)C(C)(O)CNC________________
# Output CC1CCCCC(CC=C1)=NO)C(C)(O)CNC_C______________
# ```
# Additionally, both the validation and training losses were continuing to decrease as of 180 epochs. Further gains may be seen by simply training the model for an additional period of time. Before I do any more hyper-parameter tuning I want to make sure the `ConvGRU` architecture is the best for this task. I plan on comparing it to a `GRUGRU` style architecture with different combinations of bi-directionality.
# Code to generate character count dictionary (takes a while)
char_dist = {}
for k in pvg.params['CHAR_DICT'].keys():
char_dist[k] = np.zeros((pvg.params['MAX_LENGTH']+1,))
for smile in train_smiles:
for i, char in enumerate(smile):
char_dist[char][i] += 1
char_dist[char][-1] += 1
for j in range(i, pvg.params['MAX_LENGTH']):
char_dist['_'][j] += 1
char_dist['_'][-1] += 1
# Code to generate validation character count dictionary (takes a really long time)
stride = 1000
val_char_dist = {}
for k in pvg.params['CHAR_DICT'].keys():
val_char_dist[k] = np.zeros((pvg.params['MAX_LENGTH']+1,))
for idx in range(0, pvg.params['N_TRAIN'], stride):
in_smile, out_smile = get_smiles(idx, set='train', temp=0.05)
out_smile = smi_tokenizer(out_smile)
for i, char in enumerate(out_smile):
val_char_dist[char][i] += 1
val_char_dist[char][-1] += 1
for j in range(i, pvg.params['MAX_LENGTH']):
val_char_dist['_'][j] += 1
val_char_dist['_'][-1] += 1
# +
all_chars = np.zeros((27,))
all_val_chars = np.zeros((27,))
char_labels = []
all_normalizer = 0
val_normalizer = 0
for k, v in char_dist.items():
all_normalizer += v[-1]
val_normalizer += val_char_dist[k][-1]
i = 0
for k, v in char_dist.items():
char_labels.append(k)
all_chars[i] = v[-1] / all_normalizer
all_val_chars[i] = val_char_dist[k][-1] / val_normalizer
i += 1
plt.bar(range(len(all_chars)), all_chars)
plt.plot(all_val_chars, c='purple', ls=':')
plt.xticks(range(len(all_chars)), labels=char_labels, rotation='vertical')
plt.tight_layout()
plt.show()
# +
fig, axs = plt.subplots(5, 5, figsize=(30,20))
plt.setp(axs, xticks=range(len(all_chars)), xticklabels=char_labels)
for i, ax in enumerate(fig.axes):
position_chars = np.zeros((27,))
val_pos_chars = np.zeros((27,))
all_normalizer = 0
val_normalizer = 0
for k, v in char_dist.items():
all_normalizer += v[i+4]
val_normalizer += val_char_dist[k][i+4]
j = 0
for k, v in char_dist.items():
position_chars[j] = v[i+4] / all_normalizer
val_pos_chars[j] = val_char_dist[k][i+4] / val_normalizer
j += 1
ax.bar(range(len(position_chars)), position_chars)
ax.plot(val_pos_chars, c='purple', ls=':')
ax.tick_params(labelrotation=90)
ax.set_title('Position {}'.format(i+5))
fig.tight_layout()
plt.show()
# -
# ### Exploring Model Architectures
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
model_classes = ['ConvGRU', 'ConvbiGRU', 'GRUGRU', 'biGRUGRU', 'biGRUbiGRU']
num_params = []
for model_class in model_classes:
max_len = 45
pvg = PlastVAEGen(params={'MAX_LENGTH': max_len,
'N_EPOCHS': 10,
'BATCH_SIZE': 10,
'MODEL_CLASS': model_class}, weigh_freq=False)
pvg.initiate(gdb_test)
num_params.append(count_parameters(pvg.network))
plt.bar(range(len(num_params)), num_params)
plt.xticks(range(len(num_params)), labels=model_classes)
plt.ylabel('Number of Parameters')
plt.show()
# GRUs are expensive relative to convolutional layers and even more so when bi-directionality is added. The additional model complexity should be considered when comparing the performance of these architectures.
max_len = 45
pvg = PlastVAEGen(params={'MAX_LENGTH': max_len,
'N_EPOCHS': 10,
'BATCH_SIZE': 10,
'MODEL_CLASS': 'biGRUbiGRU'}, weigh_freq=False)
pvg.initiate(gdb_test)
count_parameters(pvg.network.encoder), count_parameters(pvg.network.decoder)
# ### Transfer Learning with Plasticizers and PubChem Organics
# ### Live Model Performance
def loss_goals(m, nc):
x = m / (nc + m - 1)
return -np.log(x)
df = pd.read_csv('run_data/logs/log_reduced_len_beta0.1.txt')
df[df.data_type == 'train'].groupby('epoch').mean()['bce_loss'].plot(label='train_loss')
df[df.data_type == 'test'].groupby('epoch').mean()['bce_loss'].plot(label='val_loss')
plt.axhline(loss_goals(1, 30), c='red', ls=':', label='No better than random')
plt.axhline(loss_goals(5, 30), c='orange', ls=':', label='5x better than random')
plt.axhline(loss_goals(10, 30), c='yellow', ls=':', label='10x better than random')
plt.axhline(loss_goals(20, 30), c='black', ls=':', label='20x better than random')
plt.yscale('log')
plt.legend(loc='best')
plt.show()
df1 = pd.read_csv('run_data/logs/log_arch_ConvGRU.txt')
df2 = pd.read_csv('run_data/logs/log_arch_ConvbiGRU.txt')
df3 = pd.read_csv('run_data/logs/log_arch_GRUGRU.txt')
df4 = pd.read_csv('run_data/logs/log_arch_biGRUGRU.txt')
df5 = pd.read_csv('run_data/logs/log_arch_biGRUbiGRU.txt')
# +
colors = ['#005073', '#B86953', '#932191', '#90041F', '#0F4935']
plt.figure(figsize=(10,8))
ax = plt.subplot(111)
df1[df1.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='ConvGRU_train',
c=colors[0])
df2[df2.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='ConvbiGRU_train',
c=colors[1])
df3[df3.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='GRUGRU_train',
c=colors[2])
df4[df4.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='biGRUGRU_train',
c=colors[3])
df5[df5.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='biGRUbiGRU_train',
c=colors[4])
df1[df1.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='ConvGRU_test',
ls=':',
c=colors[0])
df2[df2.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='ConvbiGRU_test',
ls=':',
c=colors[1])
df3[df3.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='GRUGRU_test',
ls=':',
c=colors[2])
df4[df4.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='biGRUGRU_test',
ls=':',
c=colors[3])
df5[df5.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='biGRUbiGRU_test',
ls=':',
c=colors[4])
# df1[df1.data_type == 'test'].groupby('epoch').mean()['naive_loss'].plot(label='val_loss')
# ax.axhline(loss_goals(1, 27), c='red', ls=':', label='No better than random', alpha=0.5)
# ax.axhline(loss_goals(5, 27), c='orange', ls=':', label='5x better than random', alpha=0.5)
# ax.axhline(loss_goals(10, 27), c='yellow', ls=':', label='10x better than random', alpha=0.5)
# ax.axhline(loss_goals(20, 27), c='green', ls=':', label='20x better than random', alpha=0.5)
box = ax.get_position()
ax.set_position([box.x0, box.y0, box.width * 0.8, box.height])
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.yscale('log')
plt.ylabel('Total Loss', rotation='horizontal')
plt.show()
# -
df1 = pd.read_csv('run_data/logs/log_GRUGRU_pretrain_1mil.txt')
df2 = pd.read_csv('run_data/logs/log_GRUGRU_pretrain_5mil.txt')
df3 = pd.read_csv('run_data/logs/log_GRUGRU_pretrain_10mil.txt')
# df4 = pd.read_csv('run_data/logs/log_GRUGRU_transfer_1mil.txt')
# df5 = pd.read_csv('run_data/logs/log_GRUGRU_transfer_5mil.txt')
# +
colors = ['#005073', '#B86953', '#932191', '#90041F', '#0F4935']
plt.figure(figsize=(10,8))
ax = plt.subplot(111)
df1[df1.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='pretrain_1mil',
c=colors[0])
df2[df2.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='pretrain_5mil',
c=colors[1])
df3[df3.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='pretrain_10mil',
c=colors[2])
# df4[df4.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='transfer_train_1mil',
# c=colors[3])
# df5[df5.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='transfer_train_5mil',
# c=colors[4])
df1[df1.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='pretest_1mil',
ls=':',
c=colors[0])
df2[df2.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='pretest_5mil',
ls=':',
c=colors[1])
df3[df3.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='pretest_10mil',
ls=':',
c=colors[2])
# df4[df4.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='transfer_test_1mil',
# ls=':',
# c=colors[3])
# df5[df5.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='transfer_test_5mil',
# ls=':',
# c=colors[4])
# df1[df1.data_type == 'test'].groupby('epoch').mean()['naive_loss'].plot(label='val_loss')
# ax.axhline(loss_goals(1, 27), c='red', ls=':', label='No better than random', alpha=0.5)
# ax.axhline(loss_goals(5, 27), c='orange', ls=':', label='5x better than random', alpha=0.5)
# ax.axhline(loss_goals(10, 27), c='yellow', ls=':', label='10x better than random', alpha=0.5)
# ax.axhline(loss_goals(20, 27), c='green', ls=':', label='20x better than random', alpha=0.5)
box = ax.get_position()
ax.set_position([box.x0, box.y0, box.width * 0.8, box.height])
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.yscale('log')
plt.ylabel('Total Loss', rotation='horizontal')
plt.show()
# -
# df1 = pd.read_csv('run_data/logs/log_GRUGRU_prop_pred_1mil.txt')
# df2 = pd.read_csv('run_data/logs/log_GRUGRU_prop_pred_5mil.txt')
df3 = pd.read_csv('run_data/logs/log_GRUGRU_prop_pred_no_pretrain.txt')
df4 = pd.read_csv('run_data/logs/log_GRUGRU_transfer_1mil.txt')
df5 = pd.read_csv('run_data/logs/log_GRUGRU_transfer_5mil.txt')
# +
colors = ['#005073', '#B86953', '#932191', '#90041F', '#0F4935']
plt.figure(figsize=(10,8))
ax = plt.subplot(111)
# df1[df1.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='pretrain_1mil',
# c=colors[0])
# df2[df2.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='pretrain_5mil',
# c=colors[1])
df3[df3.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='no_pretrain',
c=colors[2])
df4[df4.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='pretrain_1mil',
c=colors[3])
df5[df5.data_type == 'train'].groupby('epoch').mean()['tot_loss'].plot(label='pretrain_5mil',
c=colors[4])
# df1[df1.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='pretest_1mil',
# ls=':',
# c=colors[0])
# df2[df2.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='pretest_5mil',
# ls=':',
# c=colors[1])
df3[df3.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='no_pretest',
ls=':',
c=colors[2])
df4[df4.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='pretest_1mil',
ls=':',
c=colors[3])
df5[df5.data_type == 'test'].groupby('epoch').mean()['tot_loss'].plot(label='pretest_5mil',
ls=':',
c=colors[4])
# df1[df1.data_type == 'test'].groupby('epoch').mean()['naive_loss'].plot(label='val_loss')
# ax.axhline(loss_goals(1, 27), c='red', ls=':', label='No better than random', alpha=0.5)
# ax.axhline(loss_goals(5, 27), c='orange', ls=':', label='5x better than random', alpha=0.5)
# ax.axhline(loss_goals(10, 27), c='yellow', ls=':', label='10x better than random', alpha=0.5)
# ax.axhline(loss_goals(20, 27), c='green', ls=':', label='20x better than random', alpha=0.5)
box = ax.get_position()
ax.set_position([box.x0, box.y0, box.width * 0.8, box.height])
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.yscale('log')
plt.ylabel('Total Loss', rotation='horizontal')
plt.show()
# -
# ### Testing New Models
char_weights_gdb = np.load('util/char_weights_gdb.npy')
char_weights_orgpl = np.load('util/char_weights_orgpl.npy')
with open('util/char_dict.pkl', 'rb') as f:
char_dict = pickle.load(f)
with open('util/org_dict.pkl', 'rb') as f:
org_dict = pickle.load(f)
remap_checkpoint('checkpoints/latest_GRUGRU_pretrain_1mil.ckpt', 'checkpoints/latest_GRUGRU_pretrain_1mil_large.ckpt', 'GRUGRU_pretrain_1mil_orgpl')
# Test Model
small_len = 45
big_len = 180
pvg = PlastVAEGen(params={'MAX_LENGTH': big_len,
'BATCH_SIZE': 10,
'MODEL_CLASS': 'GRUGRU',
'ARCH_SIZE': 'large',
'CHAR_DICT': char_dict,
'ORG_DICT': org_dict,
'CHAR_WEIGHTS': char_weights_orgpl},
name='test')
pvg.load('checkpoints/latest_GRUGRU_pretrain_1mil_large.ckpt', transfer=True, predict_property=True)
pvg.train(orgpl_test, epochs=10, save_best=False, save_last=False, log=False, log_latent=False, make_grad_gif=False)
pvg_big.load('checkpoints/latest_arch_GRUGRU_large.ckpt', transfer=True)
# +
# code for remapping small SMILES train to large?
ckpt_f = torch.load('checkpoints/latest_arch_GRUGRU.ckpt', map_location=torch.device('cpu'))
old_keys = ['encoder.conv2.weight', 'encoder.conv2.bias', 'encoder.conv3.weight', 'encoder.conv3.bias']
new_keys = ['encoder.conv2.0.weight', 'encoder.conv2.0.bias', 'encoder.conv3.0.weight', 'encoder.conv3.0.bias']
for ok, nk in zip(old_keys, new_keys):
ckpt_f['model_state_dict'][nk] = ckpt_f['model_state_dict'][ok]
ckpt_f['model_state_dict'].pop(ok)
torch.save(ckpt_f, 'checkpoints/latest_arch_GRUGRU_large.ckpt')
# -
# ```
# RuntimeError: Error(s) in loading state_dict for GRUGRU:
# Missing key(s) in state_dict: "encoder.conv2.0.weight", "encoder.conv2.0.bias", "encoder.conv3.0.weight", "encoder.conv3.0.bias".
# Unexpected key(s) in state_dict: "encoder.conv2.weight", "encoder.conv2.bias", "encoder.conv3.weight", "encoder.conv3.bias".
# ```
pvg_big.params['CHAR_DICT']
pl_ll.shape, org_ll.shape
upsample_ratio = int(org_ll.shape[0] / pl_ll.shape[0])
pl_ll = np.tile(pl_ll, (upsample_ratio, 1))
vae_pl_org = np.concatenate([pl_ll, org_ll])
df = pd.DataFrame(vae_pl_org, columns=['SMILES', 'll'])
df.to_pickle('../database/vae_orgpl.pkl')
gdb_char = ['Br', 'C', '1', '=', '2', '3', '4', 'N', '(', ')', '#', 'O', 'S', '5', 'F', 'Cl', 'I', '_']
pl_char = ['C', '(', ')', 'O', '=', '1', 'P', '2', '3', 'N', 'S', '4', 'Br', 'Cl', '5', '_']
org_char = ['C', 'O', '1', '=', 'N', '(', ')', '2', '3', '4', '5', '6', '7', '8', 'S', 'Cl', '9', '#', 'P', 'F', 'I', '_']
all_char = ['C', 'O', 'N', 'S', 'F', 'P', 'I', 'Cl', 'Br', '1', '2', '3', '4', '5', '6', '7', '8', '9', '#', '=', '(', ')', '_']
all_char_dict, all_ord_dict = {}, {}
for i, char in enumerate(list(all_char)):
all_char_dict[char] = i
all_ord_dict[i] = char
all_char_dict
vae_gdb = pd.read_pickle('../database/vae_gdb.pkl').to_numpy()
all_smiles = np.concatenate([vae_pl_org[:,0], vae_gdb[:,0]])
vae_gdb_test = vae_gdb[np.random.choice(np.arange(vae_gdb.shape[0]), size=1000, replace=False)]
all_smiles = [smi_tokenizer(smi) for smi in all_smiles]
char_weights_gdb = get_char_weights(all_smiles, {'NUM_CHAR': 23,
'MAX_LENGTH': 45,
'CHAR_DICT': all_char_dict})
char_weights_orgpl = get_char_weights(all_smiles, {'NUM_CHAR': 23,
'MAX_LENGTH': 180,
'CHAR_DICT': all_char_dict})
char_weights_gdb.shape, char_weights_orgpl.shape
np.save('util/char_weights_gdb.npy', char_weights_gdb)
np.save('util/char_weights_orgpl.npy', char_weights_orgpl)
with open('util/char_dict.pkl', 'wb') as f:
pickle.dump(all_char_dict, f, protocol=pickle.HIGHEST_PROTOCOL)
with open('util/org_dict.pkl', 'wb') as f:
pickle.dump(all_ord_dict, f, protocol=pickle.HIGHEST_PROTOCOL)
vae_gdb_1mil = vae_gdb[np.random.choice(np.arange(vae_gdb.shape[0]), size=int(1e6), replace=False)]
vae_gdb_5mil = vae_gdb[np.random.choice(np.arange(vae_gdb.shape[0]), size=int(5e6), replace=False)]
vae_gdb_10mil = vae_gdb[np.random.choice(np.arange(vae_gdb.shape[0]), size=int(1e7), replace=False)]
np.save('../database/vae_gdb_1mil.npy', vae_gdb_1mil)
np.save('../database/vae_gdb_5mil.npy', vae_gdb_5mil)
np.save('../database/vae_gdb_10mil.npy', vae_gdb_10mil)
|
notebooks/research_notebooks/vae_explore.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: navigating-stories
# language: python
# name: navigating-stories
# ---
# # Scrape Stories from HSPV NRW
#
# Scrape German Corona stories from HSPV NRW (University of Applied Sciences for the Police and Public Administration Nordrhrein Westfalen [German State]): https://www.hspv.nrw.de/services/corona-krise/corona-geschichten.
#
# The stories are written by staff and students of the university.
#
# The stories are stored in dictionaries with the following fields:
#
# - `link`: The link of the story page (string)
# - `title`: The title of the story page (string; is the same for all stories)
# - `author`: The author of the story (string)
# - `date`: The when the story was published (string)
# - `description`: The introductory text giving some background info on story (string)
# - `story_text`: The main text of the story (string)
#
# The web pages containing the stories are stored as `.html`. The notebook requires a folder at `DATA_DIR`.
# +
""" Scrape stories from HSPV NRW """
import os
from urllib.request import Request, urlopen
from bs4 import BeautifulSoup
# +
# Set URL of the main page containing the links to the story pages
URL = "https://www.hspv.nrw.de/nachrichten/artikel/corona-geschichten"
# Define story page ids
ID_STORIES = ["01", "02", "03", "04", "05", "06",
"07", "08", "09", "10", "11", "12", "13"]
# Set directory for storing web pages
DATA_DIR = "../hspvnrw.de/scraped/"
# -
def check_dir(data_dir):
""" Check if directory for saving web page exists """
if not os.path.isdir(data_dir):
os.makedirs(data_dir)
print(f"Created saving directory at {data_dir}")
def load_web_page(url, file_name, data_dir):
""" Check if web page can be loaded from disk;
otherwise fetch website and save as .html to disk """
check_dir(data_dir)
if os.path.exists(file_name):
with open(file_name, "r", encoding="utf-8") as file:
page = file.read()
print(f"Loaded web page from {file_name}")
else:
req = Request(url, headers={"User-Agent": "Mozilla/5.0"})
page = urlopen(req).read()
with open(file_name, "w", encoding="utf-8") as file:
file.write(page.decode())
print(f"Saved web page to {file_name}")
return page
def extract_text_from_url(url):
""" Extract text from story pages """
new_file_name = DATA_DIR + url.split("-")[-1] + ".html"
new_page = BeautifulSoup(load_web_page(
url, new_file_name, DATA_DIR), "html.parser")
new_title = new_page.title.string
new_author = new_page.find("meta", property="og:author")["content"]
new_date = new_page.find("time", attrs={"itemprop": "date"}).string
new_description = new_page.find(
"meta", property="og:description")["content"]
new_story_text = new_page.find(
"div", attrs={"class": "ce-textpic ce-h-left ce-v-above"}).text
new_doc = {
"link": url,
"title": new_title.split("#WirmeisterndieKrise ")[-1].split(" | HSPV NRW")[0],
"author": new_author,
"date": new_date,
"description": new_description,
"story_text": new_story_text
}
print(f"Extracted text from: {url}")
return new_doc
def extract_texts_from_url_ids(url, ids):
""" Wrapper to for list of story pages """
docs = []
for id in ids:
docs.append(extract_text_from_url(f"{url}-{id}"))
print("Done")
return docs
def print_doc(doc):
""" Print doc """
for field in doc.keys():
print(field + ": " + doc[field] + "\n")
# +
# Extract texts from links
docs = extract_texts_from_url_ids(URL, ID_STORIES)
# +
# Show example doc
print_doc(docs[0])
|
hspvnrw.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ash12hub/DS-Unit-2-Tree-Ensembles/blob/master/Ashwin_Raghav_Swamy_Decision_Tree_Classifier_CC.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="po219-YmmGr1" colab_type="text"
# # Decision Tree Classifier: Coding Challenge
#
# Decision trees are extremely intuitive ways to classify or label objects: you simply ask a series of questions designed to zero-in on the classification.
#
# For example, if you wanted to build a decision tree to classify an animal you come across while on a hike, you might construct the one shown here:
#
# <br>
#
# <img src=https://jakevdp.github.io/PythonDataScienceHandbook/figures/05.08-decision-tree.png></img>
#
# <br>
#
# For this coding challenge, we will be writing a decision tree classifier classifying fruit from scratch. To provide just the right level of challenge for each of you, I will let you choose your own difficulty:
#
# ## 1. Beginner 🌲
#
# Watch the video tutorial. Reproduce the code for the decision tree classifier provided [in this GitHub repo](https://github.com/random-forests/tutorials/blob/master/decision_tree.ipynb). Try to review the notebook once only or write individual functions in the classifier on your own. Make it as easy or as challenging as you wish!
#
#
#
# + id="peFRepyhmJYL" colab_type="code" outputId="ae6d0134-69fe-46a9-a4dc-b1d16e7e2236" colab={"base_uri": "https://localhost:8080/", "height": 321}
from IPython.lib.display import YouTubeVideo
YouTubeVideo('LDRbO9a6XPU')
# + id="6g_veP2Sqh0q" colab_type="code" colab={}
# STARTER CODE
training_data = [
['Green', 3, 'Apple'],
['Yellow', 3, 'Apple'],
['Red', 1, 'Grape'],
['Red', 1, 'Grape'],
['Yellow', 3, 'Lemon'],
]
# Column labels
header = ["color", "diameter", "label"]
# + id="MGIqxvhSmFiE" colab_type="code" colab={}
### YOUR CODE HERE ###
# + [markdown] id="unOFJXoLoHWs" colab_type="text"
# ## 2. Intermediate 🌲🌲
#
# Follow the video above and try writing the classifier with only minimal help from the code in the video.
# + id="Sy5QDLH9mm7w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1cdd69f1-d555-409d-ab9c-342c2862fd9b"
### YOUR CODE HERE ###
item_count = len(training_data)
item_count
# + id="s7ET3ocAw15f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="15f4be88-06e7-49eb-e5b1-b9f9c64ded67"
import pandas as pd;
import numpy as np;
data=pd.DataFrame(data = training_data, columns=header);
data
# + id="GBlPkAER1Lw0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="3b5992a6-ae76-4494-ee79-3693b4a021df"
data['color'].value_counts()
# + id="3oELIKs5xhT8" colab_type="code" colab={}
def get_unique_labels(data):
colors = data['color'].unique();
return colors;
# + id="IbHAOhZ604dV" colab_type="code" colab={}
def check_question(data, label, value):
true = data[data[label] == value]
false = data[data[label] != value]
return true, false;
# + id="3SnffiEc3X5I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="c0e54640-b3a7-4aef-91c4-4f84a2e7896e"
check_labels = get_unique_labels(data)
true, false = check_question(data, header[0], check_labels[0])
print(f'True:\n{true}\n\nFalse:\n{false}')
# + [markdown] id="rjvNeT0Powco" colab_type="text"
# ## 3. Advanced 🌲🌲🌲
#
# Write the algorithm with minimum help/guidance from the video only. Add additional improvements to the algorithm and document them - i.e. you can add handling for missing values or other features.
#
# **Extra stretch goal:** Use another dataset to classify your data.
# + id="ree8TxfhpAuQ" colab_type="code" colab={}
### YOUR CODE HERE ###
# + [markdown] id="w5zBJaCHpOK2" colab_type="text"
# ## 4. Wrap-Up 🏅
#
# Next steps:
#
# 1. Share your work with the class! I love seeing how you approached the problem and what cool solutions you came up with!
# 2. Submit your Colab to the GitHub repo for this week
# 3. Let me know how you enjoyed this spin on our traditional coding challenges!
#
# *You can provide feedback below:*
#
#
# <br>
# <br>
# <br>
#
|
Ashwin_Raghav_Swamy_Decision_Tree_Classifier_CC.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Optimise Read csv
#
# ``` {python}
# df.to_csv('pandasfile.csv', float_format='%g')
# ```
# +
from utils.config import Config
from utils.data import Data
import pandas as pd
import numpy as np
from glob import glob
import pathlib
# -
c = Config()
d = Data()
# # this is what it looks like for one chunk
path = c.get_datapath('18M_full_features/*.csv')
files = glob(path.as_posix())
files[0]
df1 = pd.read_csv(files[0])
df1_mem_usg = df1.memory_usage()
df1.info(memory_usage='deep')
# # Optimise reading floats
#
# this article is important in understading memory usage of pandas and data types [Pandas big data](https://www.dataquest.io/blog/pandas-big-data/)
df2 = pd.read_csv(files[0], nrows=10)
df2_dtypes = df2.dtypes
colnames = df2_dtypes.index
df2_floats = df2_dtypes[df2_dtypes == 'float64']
df2_new_types = ['float32' for i in df2_floats.values]
read_float_types = dict(zip(df2_floats.index, df2_new_types))
df3 = pd.read_csv(files[0], dtype=read_float_types)
df3.info(memory_usage='deep')
df3.head()
df1.head()
df4 = pd.read_csv(files[0], dtype=d.get_float_types(files[0]))
df4.info(memory_usage='deep')
df4.head()
df5 = df4[d.getDescriptorsColumnNames_C33()]
df5.info(memory_usage='deep')
df5.head()
pd.DataFrame(columns=['a','b','c'])
# # using the new function to read 18M features
df18m = d.get18M_features()
df18m.info(memory_usage='deep')
|
optimise-read_csv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DSM (BETA Phase!)
import functions
import FINE as fn
# ## Get basic energy system data
dsm_test_esM_ = functions.dsm_test_esM()
# ## Run without DSM for reference purposes
# +
"""
Given a one-node system with two generators, check whether the load and generation is shifted correctly in both
directions with and without demand side management.
"""
functions.run_esM_without_DSM()
# -
# ## Run with DSM (tFwd=tBwd=1)
# w/o & w/- time series aggregation
functions.run_esM_with_DSM(timeSeriesAggregation=False, tBwd=1, tFwd=1)
functions.run_esM_with_DSM(True, numberOfTypicalPeriods=23, tBwd=1, tFwd=1)
# ## Run with DSM (tFwd=2, tBwd=0)
functions.run_esM_with_DSM(timeSeriesAggregation=False, tBwd=0, tFwd=2)
# ## Running into trouble
#
# If tBwd + tFwd +1 is not divisor of total number of timesteps (24), infeasibilities can occur. These can be fixed with relaxing the dynamic state of charge constaints. Less notable for a larger amount of time steps!
functions.run_esM_with_DSM(timeSeriesAggregation=False, tBwd=2, tFwd=2)
|
examples/Demand_Side_Management/DSMtesting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Negative Indexing
li = [1,2,3,4,5]
li[0|]
li[0]
li[-1]
li[-2]
li[-3]
li[-5]
li[-8]
# Sequencing In List
li
li[1:5:1]
li[1:3:1]
li[1:5:2]
li
li[1:]
li[1::2]
li[:3]
li[-1:]
li[-3:-1]
li[3::-1]
li[3:1:-1]
li
li[3:0:-1]
li[3:-1:-1]
li[3:5:-1]
li[3::-1]
|
random/NegativeIndexingAndSequencingInList.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="edcaac7f"
#Imports
import numpy as np
import pandas as pd
import os
import matplotlib.pyplot as plt
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/"} id="72ca894c" outputId="50897fb2-6d51-475d-c115-5fdd3ebd6170"
#Getting the dataset
# !git clone https://github.com/alexeygrigorev/clothing-dataset-small.git ../data/raw/clothing-dataset-small/
# + id="c40d58d9"
train_path = '../data/raw/clothing-dataset-small/train/'
val_path = '../data/raw/clothing-dataset-small/validation/'
test_path = '../data/raw/clothing-dataset-small/test/'
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="1696442e" outputId="082fcd54-c692-4c23-e354-c0c9f3648f71"
import glob
from PIL import Image
import random
clothing_list = glob.glob('../data/raw/clothing-dataset-small/train/**/*.*', recursive=True)
Image.open(random.choice(clothing_list))
# + id="a7c54c95"
# !pip install deepfeatx --quiet
# + id="7f0deaac"
from deepfeatx.image import ImageFeatureExtractor
# + colab={"base_uri": "https://localhost:8080/"} id="2d03dafa" outputId="a4a9ee53-7d46-4a51-b60f-a516a7f823ac"
fe = ImageFeatureExtractor()
# + colab={"base_uri": "https://localhost:8080/"} id="f381977b" outputId="c0cd129a-69a2-4117-accf-5b319c08169c"
df = fe.extract_features_from_directory(train_path,
classes_as_folders=True,
export_class_names=True)
# -
val = fe.extract_features_from_directory(val_path,
classes_as_folders=True,
export_class_names=True)
test = fe.extract_features_from_directory(test_path,
classes_as_folders=True,
export_class_names=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 338} id="cf5e9d7e" outputId="c64c8ca7-13b8-4581-9b0c-a589577be1fd"
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="2242358e" outputId="d0c1d105-60dc-4bee-a810-660ea34d6b12"
df.shape
# + id="0c134146"
X_train = df.drop(['filepaths', 'classes'], axis=1)
y_train = df['classes']
# + id="71d37608"
X_val = val.drop(['filepaths', 'classes'], axis=1)
y_val = val['classes']
# -
X_test = test.drop(['filepaths', 'classes'], axis=1)
y_test = test['classes']
# + id="f1684d90"
from sklearn.linear_model import LogisticRegression
# + colab={"base_uri": "https://localhost:8080/"} id="e34182d2" outputId="a13cf12b-7888-4183-bbc9-121ac3098c63"
lr = LogisticRegression(max_iter=10000, random_state=42)
lr.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="8b44b77f" outputId="926de9e4-b47a-42c2-a0d4-0b2198a56472"
lr.score(X_test, y_test)
# + id="5569d41c"
from sklearn.metrics import classification_report
# + colab={"base_uri": "https://localhost:8080/"} id="3bc460aa" outputId="87aeb856-f951-4305-c348-34c0852bd21e"
y_pred = lr.predict(X_test)
print(classification_report(y_test, y_pred))
# + id="b6755d6b"
directory = '../models'
if not os.path.exists(directory):
os.makedirs(directory)
import pickle
pickle.dump(lr, open('../models/logistic_regression.pkl', 'wb'))
# + colab={"base_uri": "https://localhost:8080/"} id="7PBPVAEdvku_" outputId="e5befc38-8e2a-4701-b40d-091e09881f85"
import pickle
model = pickle.load(open('../models/logistic_regression.pkl', 'rb'))
# -
model.classes_
# + id="64513bb9"
import gradio as gr
from PIL import Image
def recognize_clothing(img):
#img=Image.fromarray(img)
vector = fe.img_to_vector(img)
label = lr.predict(vector)
confidence = lr.predict_proba(vector)
return label[0], confidence[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="93966e36" outputId="5f6619f7-2e5d-4a4c-d6a9-e0b781ec8fc6"
clothing_list = glob.glob('./clothing-dataset-small/train/**/*.*', recursive=True)
img=Image.open(random.choice(clothing_list))
img
# + id="29f9b008"
vector = fe.img_to_vector(img)
# + colab={"base_uri": "https://localhost:8080/"} id="3e7def12" outputId="379c1001-0eb1-47c3-d444-987a4402802f"
recognize_clothing(img)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="7111a112" outputId="eb5f5350-3bab-487a-d344-c9bbf777dd13"
gr.Interface(fn=recognize_clothing, inputs="image", outputs="text").launch(debug=True)
# + id="esCSa7QWzmYB"
|
notebooks/clothing_studies.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
#sys.path.append('../../')
sys.path.insert(1, '../../') # use local (git) copy
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import fmdtools.faultsim as fs
import fmdtools.resultdisp as rd
import quadpy
from IPython.display import HTML
from fmdtools.modeldef import SampleApproach
from fmdtools.modeldef import Model
# -
# ## Initial Model
#
#
# In our initial model, all we have is the flows, functions, and connections between them. These are set up in a model class as shown:
class Drone(Model):
def __init__(self, params={}):
super().__init__()
self.params=params
#add flows to the model
self.add_flow('Force_ST', {})
self.add_flow('Force_Lin', {})
self.add_flow('Force_GR' , {})
self.add_flow('Force_LG', {})
self.add_flow('EE_1', {})
self.add_flow('EEmot', {})
self.add_flow('EEctl', {})
self.add_flow('Ctl1', {})
self.add_flow('DOFs', {})
self.add_flow('Env1', {})
self.add_flow('Dir1', {})
#add functions to the model
flows=['EEctl', 'Force_ST']
self.add_fxn('StoreEE',['EE_1', 'Force_ST'])
self.add_fxn('DistEE', ['EE_1','EEmot','EEctl', 'Force_ST'])
self.add_fxn('AffectDOF',['EEmot','Ctl1','DOFs','Force_Lin'])
self.add_fxn('CtlDOF', ['EEctl', 'Dir1', 'Ctl1', 'DOFs', 'Force_ST'])
self.add_fxn('Planpath', ['EEctl', 'Env1','Dir1', 'Force_ST'])
self.add_fxn('Trajectory', ['Env1','DOFs','Dir1', 'Force_GR'] )
self.add_fxn('EngageLand',['Force_GR', 'Force_LG'])
self.add_fxn('HoldPayload',['Force_LG', 'Force_Lin', 'Force_ST'])
self.add_fxn('ViewEnv', ['Env1'])
self.construct_graph()
# ## Setting Node Positions
# As shown below, it can be difficult to make sense of a model structure using the default shell graph layout. We might instead want to see something that more closely approximates a flow chart of the system.
mdl = Drone()
rd.graph.show(mdl.graph)
# To set node positions, we can use ``rd.graph.set_pos()``, which lets one drag the nodes to their desired locations. If a model is sent to ``set_pos``, it will set those locations in the model going forward, though it is good practice to save node locations when one is done in the script or the model class file (they can be used as inputs to ``construct_graph()``.
# %matplotlib qt5
pos = rd.graph.set_pos(mdl)
pos
# We can use the same process to arrange the bipartite graph:
# %matplotlib qt5
pos = rd.graph.set_pos(mdl, gtype='bipartite')
# As shown, in a large model, the Bipartite graph is often easier to arrange to get a good layout. Since the model will be redefined several times going forward, we will use these positions to keep a consistent layout:
# +
bipartite_pos = {'StoreEE': [-1.067135163123663, 0.32466987344741055],
'DistEE': [-0.617149602161968, 0.3165981670924663],
'AffectDOF': [0.11827439153655106, 0.10792528450121897],
'CtlDOF': [-0.2636856982162134, 0.42422600969836144],
'Planpath': [-0.9347151173753852, 0.6943421719257798],
'Trajectory': [0.6180477286739998, 0.32930706399226856],
'EngageLand': [0.0015917696269229786, -0.2399760932810826],
'HoldPayload': [-0.8833099612826893, -0.247201580673997],
'ViewEnv': [0.5725955705698363, 0.6901513410348765],
'Force_ST': [-0.8925771348524384, -0.025638904424547027],
'Force_Lin': [-0.5530952425102891, -0.10380834289626095],
'Force_GR': [0.568921162299461, -0.22991830334765573],
'Force_LG': [-0.37244114591548894, -0.2355298479531287],
'EE_1': [-0.809433489993954, 0.319191761486317],
'EEmot': [-0.33469985340998853, 0.1307636433702345],
'EEctl': [-0.48751243650229525, 0.4852032717825657],
'Ctl1': [-0.06913038312848868, 0.2445174568603189],
'DOFs': [0.2606664304933561, 0.3243482171363975],
'Env1': [0.06157634305459603, 0.7099922980251693],
'Dir1': [-0.13617863906968142, 0.6037252153639261]}
graph_pos = {'StoreEE': [-1.0787279392101061, -0.06903523859088145],
'DistEE': [-0.361531174332526, -0.0935883732235363],
'AffectDOF': [0.36541282312106205, -0.09674444529230719],
'CtlDOF': [0.4664934329906758, 0.5822138245848214],
'Planpath': [-0.7095750728126631, 0.8482786785038505],
'Trajectory': [1.1006824683444765, -0.10423208715241583],
'EngageLand': [0.8423521094741182, -0.8813666134484857],
'HoldPayload': [-0.5857395187723944, -0.86974898769837],
'ViewEnv': [1.1035500215472247, 0.9373523025760659]}
# -
# %matplotlib inline
rd.graph.show(mdl, pos=graph_pos)
rd.graph.show(mdl, gtype='bipartite', pos=bipartite_pos)
# ## Network Model
#
# A network model can be used to compute network metrics and visualize network vulnerabilities.
# We can calculate network metrics using calc_aspl, calc_modularity, and calc_robustness_coefficient in the `networks` module.
# +
aspl = fs.networks.calc_aspl(mdl,gtype='normal')
q = fs.networks.calc_modularity(mdl,gtype='normal')
rc = fs.networks.calc_robustness_coefficient(mdl,gtype='normal')
print("ASPL: %.2f" % round(aspl, 2))
print("Modularity: %.2f" % round(q,2))
print("Robustness Coefficient: %.2f" % round(rc,2))
# -
# Next, we visualize network vulnerabilities using find_bridging_nodes and find_high_degree_nodes.
[bridging_nodes,fig_bridging_nodes,ax_bridging_nodes] = fs.networks.find_bridging_nodes(mdl,plot='on',gtype='normal', pos=graph_pos)
fig_bridging_nodes.savefig('bridgingnodes.pdf', format="pdf", bbox_inches = 'tight', pad_inches = 0)
[high_degree_nodes,fig_high_degree_nodes,ax_high_degree_nodes] = fs.networks.find_high_degree_nodes(mdl,plot='on',gtype='normal', pos=graph_pos,scale=1.5)
#fig_high_degree_nodes.subplots_adjust(left=0.2, right=1.3, top=1.0, bottom=0.2)
ax_high_degree_nodes.axis("off")
ax_high_degree_nodes.margins(0.08)
fig_high_degree_nodes.tight_layout()
fig_high_degree_nodes
fig_high_degree_nodes.savefig('highdegreenodes.pdf', format="pdf", bbox_inches = 'tight', pad_inches = 0.0)
# High degree nodes (along with their degrees) and bridging nodes are also obtainable as lists.
print('Bridging Nodes:',bridging_nodes)
print('High Degree Nodes:',high_degree_nodes)
# Finally, we can plot the degree distribution of the network using degree_dist.
fig = fs.networks.degree_dist(mdl,gtype='normal')
fig.savefig('degreedist.pdf', format="pdf", bbox_inches = 'tight', pad_inches = 0.0)
# The above analysis includes only function nodes. It is also possible to treat the bipartite graph (containing both functions and flows) as a unipartite-like graph and perform similar analysis on both function and flow nodes.
# +
aspl = fs.networks.calc_aspl(mdl,gtype='bipartite')
q = fs.networks.calc_modularity(mdl,gtype='bipartite')
rc = fs.networks.calc_robustness_coefficient(mdl,gtype='bipartite')
print("ASPL, functions and flows: %.2f" % round(aspl, 2))
print("Modularity, functions and flows: %.2f" % round(q,2))
print("Robustness Coefficient, functions and flows: %.2f" % round(rc,2))
# -
[bridging_nodes,fig_bridging_nodes,ax_bridging_nodes] = fs.networks.find_bridging_nodes(mdl,plot='on',gtype='bipartite', pos=bipartite_pos)
[high_degree_nodes,fig_high_degree_nodes,ax_high_degree_nodes] = fs.networks.find_high_degree_nodes(mdl,plot='on',gtype='bipartite', pos=bipartite_pos)
print('Bridging Nodes:',bridging_nodes)
print('High Degree Nodes:',high_degree_nodes)
fs.networks.degree_dist(mdl,gtype='bipartite')
# The SFF model can be simulated with options for simulation time, infection (failure) rate, and recovery (fix) rate. The start node can be selected or chosen randomly. Plotting includes an option for error bars. This models the system's response to a failure using an analogy of an epidemic spreading through a network.
fig=fs.networks.sff_model(mdl,gtype='normal',endtime=15,pi=.3,pr=.1,start_node='AffectDOF',error_bar_option='on')
fig.savefig('sff_model.pdf', format="pdf", bbox_inches = 'tight', pad_inches = 0)
fig=fs.networks.sff_model(mdl,gtype='normal',endtime=15,pi=.3,pr=.1,start_node='random',error_bar_option='on')
# ## Static Model
#
# In this demonstration, we will use a static representation of the system model to displaygraph views of fault scenarios and produce a static FMEA
#
# The static model is located in `drone_mdl_static.py`.
from drone_mdl_static import Drone as Drone_Static
# In design, it often helps to quantify the relative impact of fault scenarios. Here we produce a scenario-based FMEA to show which scenarios are most important in the model:
static_mdl = Drone_Static(params={'graph_pos':graph_pos, 'bipartite_pos':bipartite_pos})
endclasses, mdlhists = fs.propagate.single_faults(static_mdl)
reshists, diffs, summaries = rd.process.hists(mdlhists)
static_fmea = rd.tabulate.fullfmea(endclasses, summaries)
static_fmea.sort_values('expected cost', ascending=False)
print(static_fmea.sort_values('expected cost', ascending=False).to_latex())
# We can in turn visualize these faults on the graph representation of the system. Here we will focus on the break of one of the rotors in the AffectDOF function, the effects of which are shown below:
static_mdl = Drone_Static(params={'graph_pos':graph_pos, 'bipartite_pos':bipartite_pos})
endresults, resgraph, mdlhist = fs.propagate.one_fault(static_mdl,'AffectDOF', 'mechbreak', gtype='bipartite')
fig, ax = rd.graph.show(resgraph, pos = bipartite_pos, faultscen='AffectDOF: Mechbreak', time='NA', scale=1, gtype='bipartite', retfig=True)
fig.savefig('static_propagation.pdf', format="pdf", bbox_inches = 'tight', pad_inches = 0)
# ## Dynamic Model
#
# In the dynamic model, we add time ranges and dynamic behaviors to generate behavior-over-time graphs and dynamic/phase-based FMEAs.
#
# This model is located in `drone_mdl_dynamic.py`.
from drone_mdl_dynamic import Drone as Drone_Dynamic
# Here we can see how the system operates over time in the nominal case:
# Note: because of the complicated functions, the model must be re-instantiated for each function in order to work in this case
dynamic_mdl = Drone_Dynamic(params={'graph_pos':graph_pos, 'bipartite_pos':bipartite_pos})
endresults, resgraph, mdlhist = fs.propagate.nominal(dynamic_mdl)
rd.plot.mdlhistvals(mdlhist)
# As shown below, in the case of the break in the AffectDOF function, the system crashes:
dynamic_mdl = Drone_Dynamic(params={'graph_pos':graph_pos, 'bipartite_pos':bipartite_pos})
endresults, resgraph, mdlhist = fs.propagate.one_fault(dynamic_mdl,'AffectDOF', 'mechbreak', time=50)
# +
fig= rd.plot.mdlhistvals(mdlhist,'AffectDOF mechbreak', time=50, fxnflowvals={'Env1':['x','y','elev'], 'StoreEE':['soc']}, units=['m','m','m','%'],legend=False, returnfig=True, timelabel='time (s)')
ax = fig.axes[1]
ax.legend(['faulty', 'nominal'], loc='right')
fig.savefig("fault_behavior.pdf", format="pdf", bbox_inches = 'tight', pad_inches = 0)
# -
# Finally, we can see how the cost function of this scenario changes over time. As shown, when the fault is injected early, it has a lower cost because it crashes at the landing pad and not in a dangerous area. When it is injected at the end, the cost is minimal because the drone has already landed.
mdl_quad_comp = Drone_Dynamic(params={'graph_pos':graph_pos, 'bipartite_pos':bipartite_pos})
quad_comp_app = SampleApproach(mdl_quad_comp, faults=[('AffectDOF', 'mechbreak')],defaultsamp={'samp':'evenspacing','numpts':5})
quad_comp_endclasses, quad_comp_mdlhists = fs.propagate.approach(mdl_quad_comp, quad_comp_app, staged=True)
rd.plot.samplecost(quad_comp_app, quad_comp_endclasses, ('AffectDOF', 'mechbreak'))
fig = plt.gcf()
fig.savefig("cost_over_time.pdf", format="pdf", bbox_inches = 'tight', pad_inches = 0)
quad_comp_endclasses
# ## Hierarchical Model
#
# In the hierarchical model, we can use the simulation to compare system architectures. First by seeing how faults effect the behaviors in each architechture, then by seing how it affects the overall system resilience.
#
# This model is located in `drone_mdl_hierarchical.py`.
from drone_mdl_hierarchical import Drone as Drone_Hierarchical
# First, we can model how the quadrotor architecture behaves under faults--in this case, identically to the non-hierarchical model:
hierarchical_model = Drone_Hierarchical(params={'graph_pos':graph_pos, 'bipartite_pos':bipartite_pos,'arch':'quad'})
endresults, resgraph, mdlhist = fs.propagate.one_fault(hierarchical_model,'AffectDOF', 'RFmechbreak', time=50)
rd.plot.mdlhistvals(mdlhist,'AffectDOF mechbreak', time=50, fxnflowvals={'Env1':['x','y','elev'], 'StoreEE':['soc']}, legend=False)
# Then we can see how the octorotor architecture performs in the same case:
hierarchical_model = Drone_Hierarchical(params={'graph_pos':graph_pos, 'bipartite_pos':bipartite_pos,'arch':'oct'})
endresults, resgraph, mdlhist = fs.propagate.one_fault(hierarchical_model,'AffectDOF', 'RFmechbreak', time=50)
# +
fig= rd.plot.mdlhistvals(mdlhist,'AffectDOF: RFmechbreak', time=50, fxnflowvals={'Env1':['x','y','elev'], 'StoreEE':['soc']}, units=['m','m','m','%'], legend=False, returnfig=True)
ax = fig.axes[1]
ax.legend(['faulty', 'nominal'], loc='right')
fig.savefig("red_fault_behavior.pdf", format="pdf", bbox_inches = 'tight', pad_inches = 0)
# -
# As shown, the octorotor architecture enables the quadrotor to recover from the fault and land.
# Next, we can compare how each architecture mitigates the set of faults that originiate in each function:
# ### Quadcopter Resilience
#
# Here we quantify the expected costs of faults originiating in the quadcopter architecture:
mdl_quad = Drone_Hierarchical(params={'graph_pos':graph_pos, 'bipartite_pos':bipartite_pos,'arch':'quad'})
mdl_quad.fxns['AffectDOF'].faultmodes
quad_faults = [('AffectDOF', fault) for fault in list(mdl_quad.fxns['AffectDOF'].faultmodes.keys())]
quad_app = SampleApproach(mdl_quad, faults=quad_faults)
quad_endclasses, quad_mdlhists = fs.propagate.approach(mdl_quad, quad_app, staged=True)
quad_tab = rd.tabulate.simplefmea(quad_endclasses)
quad_tab.sort_values('expected cost', ascending=False)
# Based on this model, we can calculate some metrics that quantify how resilient the system was to the set of faults, such as the cost of resilience:
quad_res = sum(quad_tab['expected cost'])
quad_res
# The overall rate of crashes:
quad_crashes = quad_tab[quad_tab['cost']>100000]
quad_rate = sum(quad_crashes['rate'])
quad_rate
# The number of crashes:
quad_num_crashes = len(quad_crashes['rate'])
quad_num_crashes
# The percentage of crashes:
quad_perc_crashes = len(quad_crashes['rate'])/len(quad_tab['rate'])
quad_perc_crashes
# ### Octocopter Resilience
#
# Here we quantify the expected costs of faults originiating in the octocopter architecture:
mdl_oct = Drone_Hierarchical(params={'graph_pos':graph_pos, 'bipartite_pos':bipartite_pos,'arch':'oct'})
mdl_oct.fxns['AffectDOF'].faultmodes
oct_faults = [('AffectDOF', fault) for fault in list(mdl_oct.fxns['AffectDOF'].faultmodes.keys())]
oct_app = SampleApproach(mdl_oct, faults=oct_faults)
oct_endclasses, oct_mdlhists = fs.propagate.approach(mdl_oct, oct_app, staged=True)
oct_tab = rd.tabulate.simplefmea(oct_endclasses)
oct_tab.sort_values('expected cost', ascending=False)
# Based on this model, we can calculate some metrics that quantify how resilient the system was to the set of faults, such as the cost of resilience:
oct_res = sum(oct_tab['expected cost'])
oct_res
# The overall rate of crashes:
oct_crashes = oct_tab[oct_tab['cost']>100000]
oct_rate = sum(oct_crashes['rate'])
oct_rate
# Number of crashes:
oct_num_crashes = len(oct_crashes['rate'])
oct_num_crashes
# Percent of crashes:
oct_perc_crashes = len(oct_crashes['rate'])/len(oct_tab['rate'])
oct_perc_crashes
oct_crashes[35:]
|
multirotor example/fmdtools paper demonstration/Demonstration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.5 64-bit (''py37'': conda)'
# language: python
# name: python37564bitpy37conda071905c6724a47628f250a53a7223ba0
# ---
# +
import scipy.io as sio
import matplotlib.pyplot as plt
from matplotlib import cm as cm
import numpy as np
import time as tm
import random as rand
# Loading network data
mat = sio.loadmat('AlgorithmOutput/optimalNetwork.mat')
W = mat['A_norm_max_inh'].astype(float)
N = len(W)
V = np.load('Connectome/V_weights.npy').flatten()
# seed for random number generation
seed = 2000
np.random.seed(seed)
rand.seed(seed)
# simulate dynamics function
def simulation(time, V, W, external_input):
X = np.zeros([N, time + 1])
X[:, 0] = np.random.rand(N) * 2 - 1
for t in range(0, time):
X[:, t + 1] = np.tanh(V * external_input[t] + np.dot(W, X[:, t]))
return X
# plot information
def plot(X,inputt):
fig2 = plt.figure(figsize=(15,10),dpi=300)
fig2.patch.set_facecolor('xkcd:white')
ax4 = plt.subplot2grid((10,1), (0,0), colspan=1, rowspan=1)
cax4 = plt.xlim(0,time)
fig2.text(0.5,0.9,'Input',color='black',fontsize=25,ha="center", va="bottom")
ax4.set_facecolor('xkcd:white')
ax4.tick_params(axis=('x'), colors='white')
ax4.tick_params(axis=('y'), colors='white')
ax4.set_yticklabels([])
ax4.set_xticklabels([])
for spine in ax4.spines.values():
spine.set_edgecolor('white')
plt.plot(np.arange(0,time),inputt,color='black')
ax6 = plt.subplot2grid((10,1), (2,0), colspan=1, rowspan=2)
cax6 = plt.xlim(0,time)
fig2.text(0.5,0.75,'Neuron '+str(neuron_chosen),color='black',fontsize=25,ha="center", va="bottom")
ax6.set_facecolor('xkcd:white')
ax6.tick_params(axis=('x'), colors='black')
ax6.tick_params(axis=('y'), colors='black')
plt.plot(np.reshape(np.arange(0,time),(time,1)),np.transpose(X[neuron_chosen,:-1]),color='black',linewidth=2)
plt.xlabel('Time',fontsize=25)
ax6.xaxis.label.set_color('black')
ax6.tick_params(labelsize=15)
plt.ylim(-1,1)
ax6.spines['top'].set_color('white'); ax6.spines['right'].set_color('white')
ax6.spines['bottom'].set_color('white'); ax6.spines['left'].set_color('white')
ax5 = plt.subplot2grid((10,1), (5,0), colspan=1, rowspan=6)
cax5 = plt.imshow(X,cmap='jet',aspect='auto')
color_bar = plt.colorbar(orientation='horizontal',drawedges=False)
color_bar.outline.set_edgecolor('white')
color_bar.ax.tick_params(labelsize=15,color='black')
cbytick_obj = plt.getp(color_bar.ax.axes, 'xticklabels')
plt.setp(cbytick_obj, color='black')
plt.clim(-1,1)
ax5.set_yticklabels([])
ax5.set_facecolor('xkcd:white')
ax5.tick_params(axis=('x'), colors='black')
ax5.tick_params(axis=('y'), colors='black')
plt.ylabel('Neurons',fontsize=25)
ax5.xaxis.label.set_color('black')
ax5.yaxis.label.set_color('black')
ax5.tick_params(color='white',labelsize=15)
for spine in ax5.spines.values():
spine.set_edgecolor('white')
# apply inhibition randomly
def random_inhibition(A, n):
newA = np.abs(np.copy(A))
inhi_pos = rand.sample(range(0, len(A)), n)
for i in inhi_pos:
newA[i, :] = -newA[i, :]
return newA, inhi_pos
# check applied inhibition
def check_inhibition(A):
sumA = np.sum(A, 1)
return sum(sumA < 0)
# +
# parameters
time = 500
pulses_amount = 20 # pulses
wid = 0.01 # histogram width
duration = 8 # pulse duration
t0 = 15 # time before first pulse
t = 10 # time for 0 state
std = 2 # variation for pulses length in range [-2,2]
neuron_chosen = 200
# generate input
input_pulses = np.zeros(t0)
pulseirr_number = np.zeros(t0)
number = 1
while len(input_pulses) < time:
suplementary = rand.randrange(-std, std, 1)
input_pulses = np.append(input_pulses, np.ones(duration + suplementary))
input_pulses = np.append(input_pulses, np.zeros(t))
pulseirr_number = np.append(pulseirr_number, np.zeros(duration + suplementary))
pulseirr_number = np.append(pulseirr_number, number * np.ones(t))
number = number + 1
if len(input_pulses) > time:
aux = time - len(input_pulses)
input_pulses = input_pulses[:aux]
elif len(input_pulses) < time:
aux = time - len(input_pulses)
input_pulses = np.append(input_pulses, np.zeros(aux))
# computing dynamics for intraseries
iterations = 526
response_intraseries = np.zeros([pulses_amount, t, iterations])
out_pulsesirr = np.zeros((N, time + 1, iterations))
for j in range(0, iterations):
out_pulsesirr[:, :, j] = simulation(time, V, W, input_pulses)
for z in range(1, pulses_amount + 1):
indexes = np.where(pulseirr_number == z)[0] + 1
response_intraseries[z - 1, :, j] = out_pulsesirr[neuron_chosen, indexes, j]
# computing dynamics for interseries
iterations = 100
response_interseries = np.zeros([iterations, t, pulses_amount])
for j in range(0, iterations):
out_pulsesirr = simulation(time, V, W, input_pulses)
for p in range(0, pulses_amount):
indexes = np.where(pulseirr_number == p + 1)[0] + 1
response_interseries[j, :, p] = out_pulsesirr[neuron_chosen, indexes]
# normalizing intraseries dynamics
for i in range(response_intraseries.shape[2]):
for j in range(response_intraseries.shape[0]):
signal = response_intraseries[j, :, i]
signal = (signal - np.mean(signal)) / np.std(signal)
response_intraseries[j, :, i] = signal
# computing correlations for intraseries
intraseries_corr = []
for i in range(response_intraseries.shape[2]):
corr_mat = np.triu(np.corrcoef(response_intraseries[:, :, i]), 1).flatten()
index = np.where(corr_mat == 0)
corr_vec = np.delete(corr_mat, index).tolist()
intraseries_corr = intraseries_corr + corr_vec
intraseries_corr = np.asarray(intraseries_corr)
#normalizing dynamics and computing correlations for interseries
interseries_corr = []
for p in range(response_interseries.shape[2]):
npmean = np.mean(response_interseries[:, :, p],1).reshape(response_interseries[:, :, p].shape[0], 1)
npstd = np.std(response_interseries[:, :, p], 1).reshape(response_interseries[:, :, p].shape[0], 1)
norm_interseries = (response_interseries[:, :, p] - npmean) / npstd
corr_mat = np.triu(np.corrcoef(norm_interseries), 1).flatten()
index = np.where(corr_mat == 0)
interseries_corr = interseries_corr + np.delete(corr_mat, index).tolist()
# uncomment to save data for Visualizing Final Figuryes
# np.savetxt("ProgrammingFigures/fig5_intraseries_corr.csv", intraseries_corr, delimiter=",")
# np.savetxt("ProgrammingFigures/fig5_interseries_corr.csv", interseries_corr, delimiter=",")
# +
# computing distributions
response_intra_distribution, bins_intra = np.histogram(intraseries_corr, bins=np.arange(-1, 1 + wid, wid))
response_inter_distribution, bins_inter = np.histogram(interseries_corr, bins=np.arange(-1, 1 + wid, wid))
response_intra_distribution = response_intra_distribution / np.sum(response_intra_distribution)
response_inter_distribution = response_inter_distribution / np.sum(response_inter_distribution)
# showing distributions
fig1 = plt.figure(figsize=(15, 10))
ax2 = plt.subplot2grid((10, 1), (0, 0), colspan=1, rowspan=2)
input_external = plt.plot(input_pulses[:200], color='black')
plt.title('Input, pulse duration = ' + str(duration) + '$\pm' + str(std) + '$', fontsize=25)
ax2.spines['top'].set_color('w')
ax2.spines['right'].set_color('w')
ax2.xaxis.set_tick_params(labelsize=16)
ax2.yaxis.set_tick_params(labelsize=16)
ax1 = plt.subplot2grid((10, 1), (3, 0), colspan=1, rowspan=7)
bar_intra = plt.bar(bins_intra[:-1] + np.diff(bins_intra) / 2, response_intra_distribution, alpha=0.5, width=wid)
bar_inter = plt.bar(bins_inter[:-1] + np.diff(bins_inter) / 2, response_inter_distribution, alpha=0.5, width=wid)
ax1.tick_params(axis=('x'))
ax1.tick_params(axis=('y'))
ax1.xaxis.set_tick_params(labelsize=16)
ax1.yaxis.set_tick_params(labelsize=16)
mean_intra = np.mean(intraseries_corr)
mean_inter = np.mean(interseries_corr)
leg = plt.legend([bar_intra, bar_inter], ['Intraseries, $\mu$=' + str(round(mean_intra, 3)), 'Interseries, $\mu$=' + str(round(mean_inter, 2))], fontsize=23)
plt.title('Distribution of Correlations', fontsize=25)
plt.ylabel('Amount (normalized)', fontsize=23)
plt.xlabel('Correlation Coefficient', fontsize=23)
plt.xlim((0, 1))
# +
# computing distributions
intraseries_corr_abs = [i for i in intraseries_corr if i > 0]
interseries_corr_abs = [i for i in interseries_corr if i > 0]
[response_intra_distribution,bins_intra] = np.histogram(intraseries_corr_abs, bins=np.arange(0, 1 + wid, wid))
[response_inter_distribution,bins_inter] = np.histogram(interseries_corr_abs, bins=np.arange(0, 1 + wid, wid))
response_intra_distribution = response_intra_distribution / np.sum(response_intra_distribution)
response_inter_distribution = response_inter_distribution / np.sum(response_inter_distribution)
# computing cummulative distributions
CDF_intra = np.cumsum(response_intra_distribution)
CDF_inter = np.cumsum(response_inter_distribution)
# uncomment to save data for figure visualization
# np.savetxt("CDF_intra.csv", CDF_intra, delimiter=",")
# np.savetxt("CDF_inter.csv", CDF_inter, delimiter=",")
# np.savetxt("binsCDF.csv", bins_intra, delimiter=",")
# np.savetxt("ProgrammingFigures/intraseries_corr.csv", intraseries_corr, delimiter=",")
# np.savetxt("ProgrammingFigures/interseries_corr.csv", interseries_corr, delimiter=",")
#computing AUROC
AUROC = np.trapz(CDF_intra, CDF_inter)
AUref = np.trapz([0, 1],[0, 1])
fig1 = plt.figure(figsize=(15, 7))
ax3 = fig1.add_subplot(111)
cdf_intra = plt.plot(bins_intra[:-1] + np.diff(bins_intra) / 2, CDF_intra, label='Intraseries')
cdf_inter = plt.plot(bins_intra[:-1] + np.diff(bins_intra) / 2, CDF_inter, label='Interseries')
ax3.tick_params(axis=('x'))
ax3.tick_params(axis=('y'))
ax3.xaxis.set_tick_params(labelsize=16)
ax3.yaxis.set_tick_params(labelsize=16)
leg = plt.legend(fontsize=23)
plt.ylabel('CDF', fontsize=23)
plt.xlabel('Correlation Coefficient', fontsize=23)
plt.xlim((0, 1))
fig1 = plt.figure(figsize=(15, 7))
ax4 = fig1.add_subplot(111)
ROC = plt.plot(CDF_inter, CDF_intra, label='ROC, AUROC=' + str(round(AUROC, 4)))
cdf_inter = plt.plot([0, 1], [0, 1], 'k--', label='Reference, Area=' + str(round(AUref, 4)))
ax4.tick_params(axis=('x'))
ax4.tick_params(axis=('y'))
ax4.xaxis.set_tick_params(labelsize=16)
ax4.yaxis.set_tick_params(labelsize=16)
leg = plt.legend(fontsize=23)
plt.title('ROC', fontsize=25)
plt.ylabel('CDF Intraseries', fontsize=23)
plt.xlabel('CDF Interseries', fontsize=23)
plt.xlim((0, 1))
# -
|
Others/correlation_distributions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
# Calling DataFrame constructor
df = pd.DataFrame()
print(df)
# +
# list of strings
lst = ['Geeks', 'For', 'Geeks', 'is',
'portal', 'for', 'Geeks']
# Calling DataFrame constructor on list
df = pd.DataFrame(lst)
print(df)
# -
|
Python Pandas/DataFrame_basic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 以 Reverse Polish Notation 表示法计算算术表达式的值。
# 有效运算符为 +、-、*、/。
# 每个操作数可以是整数或另一个表达式。
# 注:两个整数之间的除法应向零截断。
# 给定的RPN表达式始终有效。
# 这意味着表达式将始终求值为结果,并且不会有任何除以零的操作。
#
#
# Example 1:
# Input: ["2", "1", "+", "3", "*"]
# Output: 9
# Explanation: ((2 + 1) * 3) = 9
#
# Example 2:
# Input: ["4", "13", "5", "/", "+"]
# Output: 6
# Explanation: (4 + (13 / 5)) = 6
#
# Example 3:
# Input: ["10", "6", "9", "3", "+", "-11", "*", "/", "*", "17", "+", "5", "+"]
# Output: 22
# Explanation:
# ((10 * (6 / ((9 + 3) * -11))) + 17) + 5
# = ((10 * (6 / (12 * -11))) + 17) + 5
# = ((10 * (6 / -132)) + 17) + 5
# = ((10 * 0) + 17) + 5
# = (0 + 17) + 5
# = 17 + 5
# = 22
# -
class Solution:
def evalRPN(self, tokens) -> int:
outputs = []
symbols = ['+', '-', '*', '/']
for val in tokens:
if val in symbols:
val_1 = int(outputs.pop())
val_2 = int(outputs.pop())
if val == '+':
val = val_1 + val_2
elif val == '-':
val = val_2 - val_1
elif val == '*':
val = val_1 * val_2
else:
val = int(val_2 / val_1)
outputs.append(str(val))
return int(outputs[0])
tokens_ = ["2", "1", "+", "3", "*"]
solution = Solution()
solution.evalRPN(tokens_)
|
Stack/0914/150. Evaluate Reverse Polish Notation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os.path
import pickle
import random
from wbtools.lib.nlp.text_preprocessing import get_documents_from_text
from sentence_transformers import SentenceTransformer, util
from matplotlib import pyplot as plt
import numpy as np
import random
import umap
from scipy import spatial
import sent2vec
from nltk import word_tokenize
from nltk.corpus import stopwords
from string import punctuation
from numpy import savetxt
from sklearn.utils import shuffle
# -
# ## Define constants
# + pycharm={"name": "#%%\n"}
min_sentence_length = 50
max_corpus_size = 10000
use_cached_embeddings = False
bio_sent_vec_model_location = "../biosentvec/model.bin"
# -
# ## Read positive and negative sentences from file
# + pycharm={"name": "#%%\n"}
def read_sentences_from_file(filepath, min_sent_length=20):
return [sentence for line in open(filepath) for sentence in get_documents_from_text(
text=line.strip(), split_sentences=True) if len(sentence) > min_sent_length]
def read_sentences_from_file_with_papid(filepath, min_sent_length=20):
return [(sent.split("\t")[0], s) for sent in open(filepath) if len(s:=sent.split("\t")[1]) > min_sent_length]
sent_expr_pattern = read_sentences_from_file("../extracted_sentences/sentences_exp_pattern.txt",
min_sentence_length)
sent_subcellloc = read_sentences_from_file("../extracted_sentences/sentences_exp_subcellloc.txt",
min_sentence_length)
neg_sent_otherexpr = read_sentences_from_file_with_papid("../extracted_sentences/neg_sentences_otherexpr.txt")
# -
# ## Clean sentences
# + pycharm={"name": "#%%\n"}
def clean_sentence(sentence):
sentence = sentence.replace('/', ' / ')
sentence = sentence.replace('.-', ' .- ')
sentence = sentence.replace('.', ' . ')
sentence = sentence.replace('\'', ' \' ')
tokens = [token for token in word_tokenize(sentence) if token not in punctuation]
sentence = ' '.join(tokens)
sentence = sentence.strip(' ').strip('.;,/-|').strip()
return sentence
def clean_sentences(sentences):
return list(set([clean_sentence(sentence) for sentence in sentences]))
def clean_sentences_with_papid(sentences_with_papid):
added_sentences = set()
ret_sent = []
for pap_id, sentence in sentences_with_papid:
sentence = clean_sentence(sentence)
if sentence not in added_sentences:
added_sentences.add(sentence)
ret_sent.append((pap_id, sentence))
return ret_sent
sent_expr_pattern_clean = clean_sentences(sent_expr_pattern)
sent_subcellloc_clean = clean_sentences(sent_subcellloc)
neg_sent_otherxpr_clean = clean_sentences_with_papid(neg_sent_otherexpr)
# -
# ## Sample sentences
# + pycharm={"name": "#%%\n"}
def get_random_sentence_subset(sentences, max_num_sentences):
corpus = sentences
random.shuffle(corpus)
return corpus[0:max_num_sentences]
if use_cached_embeddings and os.path.exists("../sentence_embeddings/corpus_expr_pattern.txt") and os.path.exists("../sentence_embeddings/corpus_subcellloc.txt") and os.path.exists("../sentence_embeddings/corpus_neg_otherexpr.txt"):
corpus_expr_pattern = [line.strip() for line in open("../sentence_embeddings/corpus_expr_pattern.txt")]
corpus_expr_subcellloc = [line.strip() for line in open("../sentence_embeddings/corpus_subcellloc.txt")]
corpus_neg_otherexpr = [line.strip() for line in open("../sentence_embeddings/corpus_neg_otherexpr.txt")]
else:
corpus_expr_pattern = get_random_sentence_subset(sent_expr_pattern_clean, max_corpus_size)
corpus_expr_subcellloc = get_random_sentence_subset(sent_subcellloc_clean, max_corpus_size)
corpus_neg_otherexpr = get_random_sentence_subset(neg_sent_otherxpr_clean, max_corpus_size)
with open("../sentence_embeddings/corpus_expr_pattern.txt", "w") as corpus_expr_pattern_file:
for sent in corpus_expr_pattern:
corpus_expr_pattern_file.write(sent + "\n")
with open("../sentence_embeddings/corpus_subcellloc.txt", "w") as corpus_expr_subcellloc_file:
for sent in corpus_expr_subcellloc:
corpus_expr_subcellloc_file.write(sent + "\n")
with open("../sentence_embeddings/corpus_neg_otherexpr.txt", "w") as corpus_neg_otherexpr_file:
for sent in corpus_neg_otherexpr:
corpus_neg_otherexpr_file.write(sent + "\n")
# -
# ## Extract sentence embeddings with SBERT
# + pycharm={"name": "#%%\n"}
if use_cached_embeddings and os.path.exists("../sentence_embeddings/corpus_embeddings_expr_pattern.pickle") and os.path.exists("../sentence_embeddings/corpus_embeddings_subcellloc.pickle") and os.path.exists("../sentence_embeddings/corpus_embeddings_neg_otherexpr.pickle"):
corpus_embeddings_expr_pattern = pickle.load(open("../sentence_embeddings/corpus_embeddings_expr_pattern.pickle", "rb"))
corpus_embeddings_subcellloc = pickle.load(open("../sentence_embeddings/corpus_embeddings_subcellloc.pickle", "rb"))
corpus_embeddings_neg_otherexpr = pickle.load(open("../sentence_embeddings/corpus_embeddings_neg_otherexpr.pickle", "rb"))
else:
embedder = SentenceTransformer('all-MiniLM-L6-v2')
corpus_embeddings_expr_pattern = embedder.encode(corpus_expr_pattern, convert_to_tensor=True)
corpus_embeddings_subcellloc = embedder.encode(corpus_expr_subcellloc, convert_to_tensor=True)
corpus_embeddings_neg_otherexpr = embedder.encode(corpus_neg_otherexpr, convert_to_tensor=True)
pickle.dump(corpus_embeddings_expr_pattern, open("../sentence_embeddings/corpus_embeddings_expr_pattern.pickle", "wb"))
pickle.dump(corpus_embeddings_subcellloc, open("../sentence_embeddings/corpus_embeddings_subcellloc.pickle", "wb"))
pickle.dump(corpus_embeddings_neg_otherexpr, open("../sentence_embeddings/corpus_embeddings_neg_otherexpr.pickle", "wb"))
# -
# ## Use UMAP to obtain a dataset that can be visualized in 2d
# + pycharm={"name": "#%%\n"}
umap_reducer = umap.UMAP(metric='cosine')
corpus_embeddings_expr_pattern_2d = umap_reducer.fit_transform(corpus_embeddings_expr_pattern)
corpus_embeddings_subcellloc_2d = umap_reducer.fit_transform(corpus_embeddings_subcellloc)
corpus_embeddings_neg_otherexpr_2d = umap_reducer.fit_transform(corpus_embeddings_neg_otherexpr)
# + pycharm={"name": "#%%\n"}
colors = ['g'] * len(corpus_embeddings_expr_pattern_2d) + ['y'] * len(corpus_embeddings_subcellloc_2d) + ['b'] * len(corpus_embeddings_neg_otherexpr_2d)
# + pycharm={"name": "#%%\n"}
all_corpora_2d = np.vstack((corpus_embeddings_expr_pattern_2d, corpus_embeddings_subcellloc_2d, corpus_embeddings_neg_otherexpr_2d))
# -
# ## UMAP visualization with different colors for each sentence category
# - expr_pattern = green
# - subcellloc = yellow
# - negative = blue
# + pycharm={"name": "#%%\n"}
fig_scatter = plt.figure(figsize=(20, 20))
plt.scatter(x=list(all_corpora_2d[:,0]), y=list(all_corpora_2d[:,1]), c=colors)
for i in range(len(all_corpora_2d)):
if i % 50 == 0:
plt.annotate(str(i), (all_corpora_2d[i,0], all_corpora_2d[i,1]))
# + pycharm={"name": "#%%\n"}
best_matches = util.semantic_search(query_embeddings=corpus_embeddings_neg_otherexpr, corpus_embeddings=corpus_embeddings_expr_pattern)
# + pycharm={"name": "#%%\n"}
[(i, corpus_id_score_dict) for i, values in enumerate(best_matches) for corpus_id_score_dict in values if corpus_id_score_dict['score'] > 0.8]
# + pycharm={"name": "#%%\n"}
1 - spatial.distance.cosine(corpus_embeddings_neg_otherexpr[394], corpus_embeddings_expr_pattern[2208])
# -
# # Extract sentence embeddings with BioSentVec
# + pycharm={"name": "#%%\n"}
model_path = bio_sent_vec_model_location
biosentvec_model = sent2vec.Sent2vecModel()
try:
biosentvec_model.load_model(model_path)
except Exception as e:
print(e)
print('model successfully loaded')
# + pycharm={"name": "#%%\n"}
corpus_embeddings_expr_pattern_bio = biosentvec_model.embed_sentences(corpus_expr_pattern)
corpus_embeddings_subcellloc_bio = biosentvec_model.embed_sentences(corpus_expr_subcellloc)
corpus_embeddings_neg_otherexpr_bio = biosentvec_model.embed_sentences(corpus_neg_otherexpr)
# + pycharm={"name": "#%%\n"}
umap_reducer = umap.UMAP(metric='cosine')
corpus_embeddings_expr_pattern_bio_2d = umap_reducer.fit_transform(corpus_embeddings_expr_pattern_bio)
corpus_embeddings_subcellloc_bio_2d = umap_reducer.fit_transform(corpus_embeddings_subcellloc_bio)
corpus_embeddings_neg_otherexpr_bio_2d = umap_reducer.fit_transform(corpus_embeddings_neg_otherexpr_bio)
# + pycharm={"name": "#%%\n"}
all_corpora_bio_2d = np.vstack((corpus_embeddings_expr_pattern_bio_2d, corpus_embeddings_subcellloc_bio_2d, corpus_embeddings_neg_otherexpr_bio_2d))
# -
# ## UMAP visualization
# - expr_pattern = green
# - subcellloc = yellow
# - negative = blue
# + pycharm={"name": "#%%\n"}
fig_scatter = plt.figure(figsize=(20, 20))
plt.scatter(x=list(all_corpora_bio_2d[:,0]), y=list(all_corpora_bio_2d[:,1]), c=colors)
# + pycharm={"name": "#%%\n"}
best_matches_bio = util.semantic_search(query_embeddings=corpus_embeddings_neg_otherexpr_bio, corpus_embeddings=corpus_embeddings_expr_pattern_bio)
# + pycharm={"name": "#%%\n"}
[(i, corpus_id_score_dict) for i, values in enumerate(best_matches_bio) for corpus_id_score_dict in values if corpus_id_score_dict['score'] > 0.8 and corpus_id_score_dict['corpus_id'] != i][0:10]
# -
# ## Calculate centroids
# + pycharm={"name": "#%%\n"}
centroid_exp_pattern_embeddings_bio = np.average(corpus_embeddings_expr_pattern_bio, 0)
centroid_subcellloc_embeddings_bio = np.average(corpus_embeddings_subcellloc_bio, 0)
# -
# ## Classification accuracy
# + pycharm={"name": "#%%\n"}
def get_stats(cosine_sim_thr, centroid_positive, embeddings_positive, embeddings_negative):
tp = len([(i, c) for i in range(len(embeddings_positive)) if (c := (1 - spatial.distance.cosine(centroid_positive, embeddings_positive[i]))) > cosine_sim_thr and not all(embeddings_positive[i] == 0)])
fn = len(embeddings_positive) - tp
fp = len([(i, c) for i in range(len(embeddings_negative)) if (c := (1 - spatial.distance.cosine(centroid_positive, embeddings_negative[i]))) > cosine_sim_thr and not all(embeddings_negative[i] == 0)])
tn = len(embeddings_negative) - fp
precision = tp / (tp + fp)
recall = tp / (tp + fn)
f1 = 2 * (precision * recall) / (precision + recall)
tpr = tp / (tp + fn)
fpr = fp / (fp + tn)
return f1, precision, recall, tpr, fpr
# + pycharm={"name": "#%%\n"}
stats_expr_pattern_list = [get_stats(i, centroid_exp_pattern_embeddings_bio, corpus_embeddings_expr_pattern_bio, corpus_embeddings_neg_otherexpr_bio) for i in [elem + 0.3 + i/100 for i, elem in enumerate([0] * 30)]]
stats_subcellloc_list = [get_stats(i, centroid_subcellloc_embeddings_bio, corpus_embeddings_subcellloc_bio, corpus_embeddings_neg_otherexpr_bio) for i in [elem + 0.3 + i/100 for i, elem in enumerate([0] * 30)]]
# -
# ## F1 measure analysis
# ### Expression Pattern
# + pycharm={"name": "#%%\n"}
plt.plot([elem + 0.3 + i/100 for i, elem in enumerate([0] * 30)], [stat[0] for stat in stats_expr_pattern_list])
# -
# ### Subcellular localization
# + pycharm={"name": "#%%\n"}
plt.plot([elem + 0.3 + i/100 for i, elem in enumerate([0] * 30)], [stat[0] for stat in stats_subcellloc_list])
# -
# ## Precision and recall of best threshold
# ### Expression Pattern
# + pycharm={"name": "#%%\n"}
print("Exp Pattern precision: " + str(get_stats(0.45, centroid_exp_pattern_embeddings_bio, corpus_embeddings_expr_pattern_bio, corpus_embeddings_neg_otherexpr_bio)[1]))
print("Exp pattern recall: " + str(get_stats(0.45, centroid_exp_pattern_embeddings_bio, corpus_embeddings_expr_pattern_bio, corpus_embeddings_neg_otherexpr_bio)[2]))
# -
# ### Subcellular localization
# + pycharm={"name": "#%%\n"}
print("Subcellular localization precision: " + str(get_stats(0.45, centroid_subcellloc_embeddings_bio, corpus_embeddings_subcellloc_bio, corpus_embeddings_neg_otherexpr_bio)[1]))
print("Subcellular localization recall: " + str(get_stats(0.45, centroid_subcellloc_embeddings_bio, corpus_embeddings_subcellloc_bio, corpus_embeddings_neg_otherexpr_bio)[2]))
# -
# ## ROC
# ### Exp pattern
# + pycharm={"name": "#%%\n"}
plt.plot([stat[4] for stat in stats_expr_pattern_list], [stat[3] for stat in stats_expr_pattern_list])
# -
# ### Subcell localization
# + pycharm={"name": "#%%\n"}
plt.plot([stat[4] for stat in stats_subcellloc_list], [stat[3] for stat in stats_subcellloc_list])
# -
# ## Save centroids to file
# + pycharm={"name": "#%%\n"}
savetxt("../sentence_embeddings/centroid_biosentvec_expr_pattern.csv", centroid_exp_pattern_embeddings_bio, delimiter=",")
savetxt("../sentence_embeddings/centroid_biosentvec_subcellloc.csv", centroid_subcellloc_embeddings_bio, delimiter=",")
# -
# ## Prepare validation set
# + pycharm={"name": "#%%\n"}
validation_nnc_high_with_papid = clean_sentences_with_papid(read_sentences_from_file_with_papid("../extracted_sentences/validation_nnc_high.txt", min_sent_length=min_sentence_length))
validation_nnc_med_with_papid = clean_sentences_with_papid(read_sentences_from_file_with_papid("../extracted_sentences/validation_nnc_med.txt", min_sent_length=min_sentence_length))
validation_nnc_low_with_papid = clean_sentences_with_papid(read_sentences_from_file_with_papid("../extracted_sentences/validation_nnc_low.txt", min_sent_length=min_sentence_length))
validation_nnc_neg_with_papid = clean_sentences_with_papid(read_sentences_from_file_with_papid("../extracted_sentences/validation_nnc_neg.txt", min_sent_length=min_sentence_length))
validation_all_clean = [*validation_nnc_high_with_papid, *validation_nnc_med_with_papid, *validation_nnc_low_with_papid, *validation_nnc_neg_with_papid]
random.shuffle(validation_all_clean)
validation_all_clean = validation_all_clean[0:1000]
# + pycharm={"name": "#%%\n"}
validation_all_clean_embeddings = biosentvec_model.embed_sentences([s[1] for s in validation_all_clean])
# + pycharm={"name": "#%%\n"}
validation_all_clean_cosine_dist_exp_pattern = []
for i, emb in enumerate(validation_all_clean_embeddings):
if any([feat > 0 for feat in emb]):
sim = 1 - spatial.distance.cosine(centroid_exp_pattern_embeddings_bio, emb)
validation_all_clean_cosine_dist_exp_pattern.append(sim)
else:
validation_all_clean_cosine_dist_exp_pattern.append("NA")
validation_all_clean_cosine_dist_subcellloc = []
for i, emb in enumerate(validation_all_clean_embeddings):
if any([feat > 0 for feat in emb]):
sim = 1 - spatial.distance.cosine(centroid_subcellloc_embeddings_bio, emb)
validation_all_clean_cosine_dist_subcellloc.append(sim)
else:
validation_all_clean_cosine_dist_subcellloc.append("NA")
# + pycharm={"name": "#%%\n"}
validation_set = [(validation_all_clean[i][0], validation_all_clean[i][1], validation_all_clean_cosine_dist_exp_pattern[i], validation_all_clean_cosine_dist_exp_pattern[i] > 0.45 if validation_all_clean_cosine_dist_exp_pattern[i] != "NA" else "NA", validation_all_clean_cosine_dist_subcellloc[i], validation_all_clean_cosine_dist_subcellloc[i] > 0.45 if validation_all_clean_cosine_dist_subcellloc[i] != "NA" else "NA") for i in range(len(validation_all_clean))]
# + pycharm={"name": "#%%\n"}
with open("../extracted_sentences/validation_set.csv", "w") as out_f_vs:
out_f_vs.write("PAPER_ID\tSENTENCE\tCOSINE_DISTANCE_EXP_PATTERN_CENTROID\tEXP_PATTERN_POSITIVE\tCOSINE_DISTANCE_SUBCELLLOC\tSUBCELLLOC_POSITIVE\n")
for line in validation_set:
out_f_vs.write("\t".join([str(col) for col in line]) + "\n")
# + pycharm={"name": "#%%\n"}
|
notebooks/sentence_classification_with_db_sentences.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# ## Create Azure Machine Learning datasets for Anomaly Detection
#
# Azure Machine Learning datasets can be extremely useful for your local or remote experiments. In this notebook, we will do the following things.
#
# 1. Configure workspace using credentials for Azure subscription
# 2. Download the dataset from ADLS Gen2
# 3. Upload the featured dataset into the default datastore in Azure
# 4. Register the featured dataset into Azure
#
# ## Configure workspace using credentials for Azure subscription
#
# As part of the setup you have already created a Workspace. To run AutoML, you also need to create an Experiment. An Experiment corresponds to a prediction problem you are trying to solve, while a Run corresponds to a specific approach to the problem.
# +
# Install the required package
# !pip install azure-storage-blob==2.1.0
# Import the libraries
from azureml.core import Workspace
# Importing user defined config
import config
# Import the subscription details as below to access the resources
subscription_id=config.subscription_id
resource_group=config.resource_group
workspace_name=config.workspace_name
try:
workspace = Workspace(subscription_id = subscription_id, resource_group = resource_group, workspace_name = workspace_name)
# write the details of the workspace to a configuration file to the notebook library
workspace.write_config()
print("Workspace configuration succeeded. Skip the workspace creation steps below")
except:
print("Workspace not accessible. Change your parameters or create a new workspace below")
# -
# ## Download the dataset from ADLS Gen2
# +
## setting up the credentials for ADLS Gen2
import os
from azure.storage.blob import BlockBlobService
# setting up blob storage configs
STORAGE_ACCOUNT_NAME = config.STORAGE_ACCOUNT_NAME
STORAGE_ACCOUNT_ACCESS_KEY = config.STORAGE_ACCOUNT_ACCESS_KEY
STORAGE_CONTAINER_NAME = "azureml-mfg"
blob_service = BlockBlobService(STORAGE_ACCOUNT_NAME, STORAGE_ACCOUNT_ACCESS_KEY)
output_file_path=os.path.join(os.getcwd(),"data", "mfg_anomaly_pdm.csv")
output_blob_file= "mfg_anomaly_pdm.csv"
# Create a project_folder if it doesn't exist
if not os.path.isdir('anomalydata'):
os.mkdir('anomalydata')
# uploading the csv to the ADLSGen2 storage container
blob_service.get_blob_to_path(STORAGE_CONTAINER_NAME, output_blob_file,output_file_path)
# -
# ## Upload the featured dataset into the default datastore in Azure
# +
#Uploading dataset to the Datastore
from sklearn import datasets
from azureml.core.dataset import Dataset
from scipy import sparse
import os
ds = workspace.get_default_datastore()
ds.upload(src_dir='./anomalydata', target_path='mfganomalydata', overwrite=True, show_progress=True)
final_df = Dataset.Tabular.from_delimited_files(path=ds.path('mfganomalydata/mfg_anomaly_pdm.csv'))
# -
# ## Register the featured dataset into Azure
#Registering the dataset in Azure ML
train_data_registered = final_df.register(workspace=workspace,
name='pdmanomalymfg',
description='Synapse Mfg data',
tags= {'type': 'Mfg', 'date':'2020'},
create_new_version=False)
|
Manufacturing/automation/artifacts/amlnotebooks/2a Machine Anomaly Detection Dataset Registration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# +
def load_city(city_name):
"""
Load all the information of a city into 5 dataframes
"""
lst=[]
for file in ['business', 'checkin', 'review', 'tip', 'user']:
f = open(f'yelp-all/{city_name}/{file}.json')
lst.append(pd.read_json(f, lines = True))
business = lst[0]
checkin = lst[1]
review = lst[2]
tip = lst[3]
user = lst[4]
return business, checkin, review, tip, user
def just_categories(business):
"""
Split the categories column of a dataframe into seperate categories with a seperate entry with combination category
and business_id as key.
"""
genres_m = business.apply(lambda row: pd.Series([row['business_id']] + row['categories'].lower().split(",")), axis=1)
stack_genres = genres_m.set_index(0).stack()
df_stack_genres = stack_genres.to_frame()
df_stack_genres['business_id'] = stack_genres.index.droplevel(1)
df_stack_genres.columns = ['category', 'business_id']
return df_stack_genres.reset_index()[['business_id', 'category']]
def split_data(data, d=0.75):
"""
Split data in a training and test set with a standard distrubution of 0.75
"""
np.random.seed(seed=5)
mask_test = np.random.rand(data.shape[0]) < d
mask_test = mask_test
return data[mask_test], data[~mask_test]
def handle_duplicates (reviews):
"""
returns the mean of businesses having multiple reviews by the same user
"""
try:
return reviews.groupby(['business_id', 'categories', 'user_id'])['stars'].mean().reset_index()
except:
return reviews.groupby(['business_id', 'user_id'])['stars'].mean().reset_index()
def join_business_reviews(business, reviews):
business = business.set_index('business_id')
reviews = reviews.set_index('business_id')
temp = reviews.join(business, rsuffix='business').reset_index()
return temp[['user_id', 'business_id', 'review_id', 'categories', 'stars']]
def rating_density(reviews):
"""
Compute the density of a dataset
"""
return number_of_ratings(reviews)/(number_of_businesses(reviews) * number_of_users(reviews))
def number_of_businesses(reviews):
"""
returns the number of unique businesses in a set of reviews
"""
return len(reviews['business_id'].unique())
def number_of_users(reviews):
"""
returns the number of unique users in a set of reviews
"""
return len(reviews['user_id'].unique())
def number_of_ratings(reviews):
"""
returns the number of ratings of a set of reviews
"""
return reviews.shape[0]
def print_properties(reviews):
print("Number of Businesses", number_of_businesses(reviews))
print("Number of Users", number_of_users(reviews))
print("Number of ratings", number_of_ratings(reviews))
print("Rating Density", rating_density(reviews))
# +
def pivot_categories(df):
"""
Create a one-hot encoded matrix for genres
"""
return df.pivot_table(index='business_id', columns='category', aggfunc='size', fill_value=0)
def pivot_ratings(df):
"""
Creates a utility matrix for user ratings for businesses
"""
return df.pivot(values='stars', columns='user_id', index='business_id')
def create_similarity_matrix_categories(matrix):
"""
Create a similarity matrix based on categories
"""
npu = matrix.values
m1 = npu @ npu.T
diag = np.diag(m1)
m2 = m1 / diag
m3 = np.minimum(m2, m2.T)
return pd.DataFrame(m3, index = matrix.index, columns = matrix.index)
# +
def predict_ratings(similarity, utility, to_predict):
"""Predicts the predicted rating for the input test data.
Arguments:
similarity -- a dataFrame that describes the similarity between items
utility -- a dataFrame that contains a rating for each user (columns) and each movie (rows).
If a user did not rate an item the value np.nan is assumed.
to_predict -- A dataFrame containing at least the columns movieId and userId for which to do the predictions
"""
# copy input (don't overwrite)
ratings_test_c = to_predict.copy()
# apply prediction to each row
ratings_test_c['predicted rating'] = to_predict.apply(lambda row: predict_ids(similarity, utility, row['user_id'], row['business_id']), axis=1)
return ratings_test_c[['business_id', 'user_id', 'stars', 'predicted rating']]
### Helper functions for predict_ratings_item_based ###
def predict_ids(similarity, utility, user_id, business_id):
# select right series from matrices and compute
if user_id in utility.columns and business_id in similarity.index:
return predict_vectors(utility.loc[:,user_id], similarity[business_id])
return 0
def predict_vectors(user_ratings, similarities):
# select only movies actually rated by user
relevant_ratings = user_ratings.dropna()
# select corresponding similairties
similarities_s = similarities[relevant_ratings.index]
# select neighborhood
similarities_s = similarities_s[similarities_s > 0.0]
relevant_ratings = relevant_ratings[similarities_s.index]
# if there's nothing left return a prediction of 0
norm = similarities_s.sum()
if(norm == 0):
return 0
# compute a weighted average (i.e. neighborhood is all)
return np.dot(relevant_ratings, similarities_s)/norm
def mse(predicted_ratings):
"""
Computes the means square error betweeen actual ratings
"""
diff = predicted_ratings['stars'] - predicted_ratings['predicted rating']
return (diff**2).mean()
def mean_center_columns(matrix):
for column in matrix.columns:
matrix[column] -= matrix[column].mean()
return matrix
# +
def cosine_similarity(matrix, id1, id2):
"""Compute cosine similarity"""
selected_features = matrix.loc[id1].notna() & matrix.loc[id2].notna()
# if no matching features, return 0
if not selected_features.any():
return 0
# get the features from the matrix
features1 = matrix.loc[id1][selected_features]
features2 = matrix.loc[id2][selected_features]
# return 1 for the diagonals and 0 if there are no matching features
if features1.equals(features2):
return 1
if features1.max() == 0 or features2.max() == 0:
return 0
return sum(features1 * features2)/((sum(features1**2)**0.5) * sum((features2**2))**0.5)
def create_similarity_matrix_cosine(matrix):
""" creates the similarity matrix based on cosine similarity """
similarity_matrix = pd.DataFrame(0, index=matrix.index, columns=matrix.index, dtype=float)
id1 = similarity_matrix.columns.values
for i in id1:
for j in id1:
similarity_matrix[i][j] = cosine_similarity(matrix, i, j)
return similarity_matrix
# -
def content_based_benchmark(city):
"""
Test the performance of our content-based system
"""
a, b, c, d, e = load_city(city)
print(city, "Content-based collaborative filtering")
print_properties(c)
review_business = join_business_reviews(a,c)
review_business_clean = handle_duplicates(review_business)
training, test = split_data(review_business_clean, d=0.9)
training_split = just_categories(training).drop_duplicates()
utility_categories = pivot_categories(training_split)
utility_ratings = pivot_ratings(review_business_clean)
similarity_categories = create_similarity_matrix_categories(utility_categories)
predictions = predict_ratings(similarity_categories, utility_ratings, test[['user_id', 'business_id', 'stars']])
print("MSE", mse(predictions), end='\n\n')
return predictions
def user_based_benchmark(city):
"""
Test the performance of our user-based system
"""
a, b, c, d, e = load_city(city)
print(city, "User-based collaborative filtering")
print_properties(c)
reviews_clean = handle_duplicates(c)
training, test = split_data(reviews_clean, d=0.9)
utility = mean_center_columns(pivot_ratings(training))
similarity = create_similarity_matrix_cosine(utility)
predictions = predict_ratings(similarity, utility, test[['user_id', 'business_id', 'stars']])
print("MSE", mse(predictions), end='\n\n')
return predictions
def item_based_benchmark(city):
"""
Test the performance of our user-based system
"""
a, b, c, d, e = load_city(city)
print(city, "Item-based collaborative filtering")
print_properties(c)
reviews_clean = handle_duplicates(c)
training, test = split_data(reviews_clean, d=0.9)
utility = mean_center_columns(pivot_ratings(training).T).T
similarity = create_similarity_matrix_cosine(utility)
predictions = predict_ratings(similarity, utility, test[['user_id', 'business_id', 'stars']])
print("MSE", mse(predictions), end='\n\n')
return predictions
for city in ['stouffville', 'sun city', 'westlake']:
content_based_benchmark(city)
user_based_benchmark(city)
item_based_benchmark(city)
|
Item_User_Content.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import print_function
import numpy as np
from tensorflow.keras.preprocessing import sequence
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Embedding, LSTM, Bidirectional
from tensorflow.keras.datasets import imdb
max_features = 200
# cut texts after this number of words
# (among top max_features most common words)
maxlen = 100
batch_size = 32
print('Loading data...')
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=max_features)
print(len(x_train), 'train sequences')
print(len(x_test), 'test sequences')
print('Pad sequences (samples x time)')
x_train = sequence.pad_sequences(x_train, maxlen=maxlen)
x_test = sequence.pad_sequences(x_test, maxlen=maxlen)
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
y_train = np.array(y_train)
y_test = np.array(y_test)
x_train
y_train
model = Sequential()
model.add(Embedding(max_features, 128, input_length=maxlen))
model.add(Bidirectional(LSTM(64)))
model.add(Dropout(0.5))
model.add(Dense(1, activation='sigmoid'))
# try using different optimizers and different optimizer configs
model.compile('adam', 'binary_crossentropy', metrics=['accuracy'])
print('Train...')
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=4,
validation_data=[x_test, y_test])
|
010-BiDirectional-LSTM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://github.com/pmservice/ai-openscale-tutorials/raw/master/notebooks/images/banner.png" align="left" alt="banner">
# # IBM Watson OpenScale and Batch Processing:<br>Remote Spark
# This notebook must be run in the Python 3.x runtime environment. It requires Watson OpenScale service credentials.
#
# The notebook configures Watson OpenScale to monitor the German Credit Risk model. Use the notebook to enable quality and drift monitoring and run on-demand evaluations. Before you can run the notebook, you must have the following resources:
#
# 1. The common configuration JSON and Drift Configuration archive generated by using the [common configuration notebook](https://github.com/IBM/watson-openscale-samples/blob/main/Cloud%20Pak%20for%20Data/Batch%20Support/Configuration%20generation%20for%20OpenScale%20batch%20subscription.ipynb).
# 2. Feedback, payload, and drifted transactions tables in an IBM DB2 storage that use the data description language statements (DDLs) that are generated as part of running the previous common configuration notebook.
# ## Contents
#
# 1. [Setup](#setup)
# 2. [Configure Watson OpenScale](#openscale)
# 3. [Set up a subscription](#subscription)
# 4. [Quality monitoring](#quality)
# 5. [Drift monitoring](#drift)
# # 1. Setup <a name="setup"></a>
# ## Package installation
#
# First import some of the packages you need to use. After you finish installing the following software packages, restart the kernel.
#
#
import warnings
warnings.filterwarnings('ignore')
# %env PIP_DISABLE_PIP_VERSION_CHECK=1
# !pip install ibm-watson-openscale --no-cache | tail -n 1
# !pip show ibm-watson-openscale
# ## Configure credentials
#
# Provide your IBM Watson OpenScale credentials in the following cell:
#
#
WOS_CREDENTIALS = {
"url": "<cluster-url>",
"username": "<username>",
"password": "<password>",
"instance_id": "<openscale instance id>"
}
# ## Specify model details
# ### Service provider and subscription metadata
# +
# Service Provider
SERVICE_PROVIDER_NAME = "<service-provider-name>"
SERVICE_PROVIDER_DESCRIPTION = "<service-provider-description>"
# Subscription
SUBSCRIPTION_NAME = "<subscription-name>"
SUBSCRIPTION_DESCRIPTION = "<subscription-description>"
# -
# ### Spark Cluster
#
# Make sure that the Apache Spark manager on the Spark cluster is running, and then provide the following details:
#
# - SPARK_ENGINE_ENDPOINT: _Endpoint URL where the Spark Manager Application is running_
# - SPARK_ENGINE_USERNAME: _Username to connect to Spark Manager Application_
# - SPARK_ENGINE_PASSWORD: _<PASSWORD>_
# - SPARK_ENGINE_NAME: _Custom display name for the Spark Manager Application_
# - SPARK_ENGINE_DESCRIPTION: _Custom description for the Spark Manager Application_
SPARK_ENGINE_NAME="<spark-engine-name>"
SPARK_ENGINE_DESCRIPTION="<spark-engine-description>"
SPARK_ENGINE_ENDPOINT="<spark-engine-endpoint>"
SPARK_ENGINE_ENDPOINT_USERNAME="<spark-engine-username>"
SPARK_ENGINE_ENDPOINT_PASSWORD="<spark-engine-password>"
# #### Provide Spark Resource Settings
#
# To configure how much of your Spark Cluster resources this job can consume, edit the following values:
#
#
# - max_num_executors: _Maximum Number of executors to launch for this session_
# - min_executors: _Minimum Number of executors to launch for this session_
# - executor_cores: _Number of cores to use for each executor_
# - executor_memory: _Amount of memory (in GBs) to use per executor process_
# - driver_cores: _Number of cores to use for the driver process_
# - driver_memory: _Amount of memory (in GBs) to use for the driver process_
spark_parameters = {
"max_num_executors": 2,
"min_num_executors": 1,
"executor_cores": 3,
"executor_memory": 2,
"driver_cores": 2,
"driver_memory": 2
}
# ### Storage Inputs
#
# Please enter a name and description for your JDBC Storage
#
# - JDBC_CONNECTION_NAME: _Custom display name for the JDBC Storage Connection_
# - JDBC_CONNECTION_DESCRIPTION: _Custom description for the JDBC Storage Connection_
#
# To connect to your JDBC storage, you must provide the following details:
#
# - JDBC_HOST: Hostname of the JDBC Connection
# - JDBC_PORT: Port of the JDBC Connection
# - JDBC_USE_SSL: Boolean Flag to indicate whether to use SSL while connecting.
# - JDBC_SSL_CERTIFICATE: SSL Certificate [Base64 encoded string] of the JDBC Connection. Ignored if JDBC_USE_SSL is False.
# - JDBC_DRIVER: Class name of the JDBC driver to use to connect.
# - JDBC_USERNAME: Username of the JDBC Connection
# - JDBC_PASSWORD: Password of the JDBC Connection
# - JDBC_DATABASE_NAME: Name of the Database to connect to.
# +
JDBC_CONNECTION_NAME = "<jdbc-connection-name>"
JDBC_CONNECTION_DESCRIPTION = "<jdbc-connection-description>"
JDBC_HOST = "<Hostname of the JDBC Connection>"
JDBC_PORT = "<Port of the JDBC Connection>"
JDBC_USE_SSL = "<Boolean Flag to indicate whether to use SSL while connecting.>"
JDBC_SSL_CERTIFICATE = "<SSL Certificate [Base64 encoded string] of the JDBC Connection. Ignored if JDBC_USE_SSL is False.>"
JDBC_DRIVER = "<Class name of the JDBC driver to use to connect.>"
JDBC_USERNAME = "<Username of the JDBC Connection>"
JDBC_PASSWORD = "<Password of the JDBC Connection>"
JDBC_DATABASE_NAME = "<Name of the Database to connect to.>"
# -
# ### Feedback table metadata
#
# The quality monitor stores metadata in the feedback table. To configure the quality monitor, you must provide the following details. To skip quality monitoring, run the following cell to initialize variables with the value of `None`.
#
# - FEEDBACK_SCHEMA_NAME: _Schema name where feedback table is present_
# - FEEDBACK_TABLE_NAME: _Name of the feedback table_
# +
#feedback
FEEDBACK_SCHEMA_NAME = None
FEEDBACK_TABLE_NAME = None
# -
# ### Payload and drift table metadata
#
# The drift monitor stores metadata in the payload and drift tables. To configure the drift monitor, you must provide the following details. To skip drift monitoring, run the following cell to initialize variables with the value of `None`.
#
# - PAYLOAD_SCHEMA_NAME: _Schema name where payload logging table is present_
# - PAYLOAD_TABLE_NAME: _Name of the payload logging table_
# - DRIFT_SCHEMA_NAME: _Schema name where drifted transactions table is present_
# - DRIFT_TABLE_NAME: _Name of the drifted transactions table_
#
# +
#payload logging
PAYLOAD_SCHEMA_NAME = None
PAYLOAD_TABLE_NAME = None
#drift
DRIFT_SCHEMA_NAME = None
DRIFT_TABLE_NAME = None
# -
# # 2. Configure Watson OpenScale <a name="openscale"></a>
# ### Import the required libraries and set up the Watson OpenScale client
# +
from ibm_cloud_sdk_core.authenticators import CloudPakForDataAuthenticator
from ibm_watson_openscale import *
from ibm_watson_openscale.supporting_classes.enums import *
from ibm_watson_openscale.supporting_classes import *
from ibm_watson_openscale.base_classes.watson_open_scale_v2 import *
authenticator = CloudPakForDataAuthenticator(
url=WOS_CREDENTIALS["url"],
username=WOS_CREDENTIALS["username"],
password=WOS_CREDENTIALS["password"],
disable_ssl_verification=True
)
wos_client = APIClient(authenticator=authenticator, service_url=WOS_CREDENTIALS["url"], service_instance_id=WOS_CREDENTIALS["instance_id"])
# -
# ### Display Watson OpenScale datamart details
wos_client.data_marts.show()
data_marts = wos_client.data_marts.list().result.data_marts
data_mart_id=data_marts[0].metadata.id
# ### Create a service provider
# +
# Delete existing service provider with the same name as provided
service_providers = wos_client.service_providers.list().result.service_providers
for provider in service_providers:
if provider.entity.name == SERVICE_PROVIDER_NAME:
wos_client.service_providers.delete(service_provider_id=provider.metadata.id)
break
# +
# Add Service Provider
added_service_provider_result = wos_client.service_providers.add(
name=SERVICE_PROVIDER_NAME,
description=SERVICE_PROVIDER_DESCRIPTION,
service_type=ServiceTypes.CUSTOM_MACHINE_LEARNING,
credentials={},
operational_space_id="production",
background_mode=False
).result
service_provider_id = added_service_provider_result.metadata.id
wos_client.service_providers.show()
# -
service_provide_details = wos_client.service_providers.get(service_provider_id=service_provider_id).result
print(service_provide_details)
# ### Create integrated systems for Spark Engine and JDBC Storage
# +
# Delete existing spark and jdbc integrated systems if present
integrated_systems = IntegratedSystems(wos_client).list().result.integrated_systems
for system in integrated_systems:
if system.entity.name in (SPARK_ENGINE_NAME, JDBC_CONNECTION_NAME):
print("Deleting integrated system {}".format(system.entity.name))
IntegratedSystems(wos_client).delete(integrated_system_id=system.metadata.id)
# -
# #### Spark Engine
# +
spark_engine_details = IntegratedSystems(wos_client).add(
name=SPARK_ENGINE_NAME,
description=SPARK_ENGINE_DESCRIPTION,
type="spark",
credentials={
"username": SPARK_ENGINE_ENDPOINT_USERNAME,
"password": <PASSWORD>ENDPOINT_PASSWORD
},
connection={
"endpoint": SPARK_ENGINE_ENDPOINT,
"location_type": "custom"
}
).result
spark_engine_id = spark_engine_details.metadata.id
print(spark_engine_details)
# -
# #### JDBC Storage
# +
jdbc_url = "jdbc:db2://{}:{}/{}".format(JDBC_HOST, JDBC_PORT, JDBC_DATABASE_NAME)
jdbc_connection_details = IntegratedSystems(wos_client).add(
name=JDBC_CONNECTION_NAME,
description=JDBC_CONNECTION_DESCRIPTION,
type="jdbc",
credentials={
"username": JDBC_USERNAME,
"password": <PASSWORD>
},
connection={
"location_type": "jdbc",
"jdbc_url": jdbc_url,
"db_driver": JDBC_DRIVER,
"use_ssl": JDBC_USE_SSL,
"certificate": JDBC_SSL_CERTIFICATE,
}
).result
jdbc_connection_id=jdbc_connection_details.metadata.id
print(jdbc_connection_details)
# -
# # 3. Set up a subscription <a name="subscription"></a>
# +
# Delete an existing subscription with the provided name
subscriptions = wos_client.subscriptions.list().result.subscriptions
for sub in subscriptions:
if sub.entity.deployment.name == SUBSCRIPTION_NAME:
wos_client.subscriptions.delete(subscription_id=sub.metadata.id)
break
# Display all subscriptions
wos_client.subscriptions.show()
# -
# ### Set subscription metadata
#
# In the following cell, type a path to the common configuration JSON file that you created by running the [common configuration notebook](https://github.com/IBM/watson-openscale-samples/blob/main/Cloud%20Pak%20for%20Data/Batch%20Support/Configuration%20generation%20for%20OpenScale%20batch%20subscription.ipynb). After you edit the path information, run the cell to set the asset details and properties, the deployment details, the analytics engine details, and to add the required tables as data sources.
# +
import uuid
common_configuration = None
with open("/path/to/dir/containing/common_config.json", "r") as fp:
common_configuration = json.load(fp).get("common_configuration")
if common_configuration is None:
print("Please provide the correct path to the common configuration JSON")
# Set asset details
asset = Asset(
asset_id=str(uuid.uuid4()),
url="",
name=SUBSCRIPTION_NAME,
asset_type=AssetTypes.MODEL,
input_data_type=InputDataType.STRUCTURED,
problem_type=ProblemType.BINARY_CLASSIFICATION
)
# Set deployment details
asset_deployment = AssetDeploymentRequest(
deployment_id=str(uuid.uuid4()),
name=SUBSCRIPTION_NAME,
description=SUBSCRIPTION_DESCRIPTION,
deployment_type="batch"
)
# Set asset properties
asset_properties_request = AssetPropertiesRequest(
label_column=common_configuration["label_column"],
probability_fields=[common_configuration["probability"]],
prediction_field=common_configuration["prediction"],
feature_fields=common_configuration["feature_columns"],
categorical_fields=common_configuration["categorical_columns"]
)
# Set analytics engine details
analytics_engine = AnalyticsEngine(
type="spark",
integrated_system_id=spark_engine_id,
parameters = spark_parameters
)
# Add selected tables as data sources
data_sources = []
if FEEDBACK_SCHEMA_NAME is not None and FEEDBACK_TABLE_NAME is not None:
feedback_data_source = DataSource(
type="feedback",
database_name=JDBC_DATABASE_NAME,
schema_name=FEEDBACK_SCHEMA_NAME,
table_name=FEEDBACK_TABLE_NAME,
connection=DataSourceConnection(
type="jdbc",
integrated_system_id=jdbc_connection_id
)
)
data_sources.append(feedback_data_source)
if PAYLOAD_SCHEMA_NAME is not None and PAYLOAD_TABLE_NAME is not None \
and DRIFT_SCHEMA_NAME is not None and DRIFT_TABLE_NAME is not None:
payload_logging_data_source = DataSource(
type="payload",
database_name=JDBC_DATABASE_NAME,
schema_name=PAYLOAD_SCHEMA_NAME,
table_name=PAYLOAD_TABLE_NAME,
connection=DataSourceConnection(
type="jdbc",
integrated_system_id=jdbc_connection_id
)
)
drifted_transactions_table_data_source = DataSource(
type="drift",
database_name=JDBC_DATABASE_NAME,
schema_name=DRIFT_SCHEMA_NAME,
table_name=DRIFT_TABLE_NAME,
connection=DataSourceConnection(
type="jdbc",
integrated_system_id=jdbc_connection_id
)
)
data_sources.append(payload_logging_data_source)
data_sources.append(drifted_transactions_table_data_source)
# +
# Adding the subscription
subscription_details = Subscriptions(wos_client).add(
data_mart_id=data_mart_id,
service_provider_id=service_provider_id,
asset=asset,
deployment=asset_deployment,
asset_properties=asset_properties_request,
analytics_engine=analytics_engine,
data_sources=data_sources).result
subscription_id = subscription_details.metadata.id
print(subscription_details)
# +
# Checking subscription status
wos_client.subscriptions.get(subscription_id).result.entity.status.state
# +
# Add training, output, and input data schemas to the subscription
training_data_schema_patch_document=[
JsonPatchOperation(op=OperationTypes.REPLACE, path='/asset_properties/training_data_schema', value=common_configuration["training_data_schema"])
]
input_data_schema_patch_document=[
JsonPatchOperation(op=OperationTypes.REPLACE, path='/asset_properties/input_data_schema', value=common_configuration["input_data_schema"])
]
output_data_schema_patch_document=[
JsonPatchOperation(op=OperationTypes.REPLACE, path='/asset_properties/output_data_schema', value=common_configuration["output_data_schema"])
]
wos_client.subscriptions.update(subscription_id=subscription_id, patch_document=training_data_schema_patch_document)
wos_client.subscriptions.update(subscription_id=subscription_id, patch_document=input_data_schema_patch_document)
wos_client.subscriptions.update(subscription_id=subscription_id, patch_document=output_data_schema_patch_document)
# +
# Check subscription status
wos_client.subscriptions.get(subscription_id).result.entity.status.state
# -
# # 4. Quality monitoring <a name="quality"></a>
# ### Enable the quality monitor
#
# In the following code cell, default values are set for the quality monitor. You can change the default values by updating the optional `min_feedback_data_size` attribute in the `parameters` dict and set the quality threshold in the `thresholds` list.
# +
import time
target = Target(
target_type=TargetTypes.SUBSCRIPTION,
target_id=subscription_id
)
parameters = {
"min_feedback_data_size": 2000000
}
thresholds = [{
"metric_id": "area_under_roc",
"type": "lower_limit",
"value": 0.8
}]
quality_monitor_details = wos_client.monitor_instances.create(
data_mart_id=data_mart_id,
monitor_definition_id=wos_client.monitor_definitions.MONITORS.QUALITY.ID,
target=target,
parameters=parameters,
thresholds=thresholds
).result
quality_monitor_instance_id = quality_monitor_details.metadata.id
print(quality_monitor_details)
# -
# ### Check monitor instance status
# +
quality_status = None
while quality_status not in ("active", "error"):
monitor_instance_details = wos_client.monitor_instances.get(monitor_instance_id=quality_monitor_instance_id).result
quality_status = monitor_instance_details.entity.status.state
if quality_status not in ("active", "error"):
print(datetime.utcnow().strftime('%H:%M:%S'), quality_status)
time.sleep(30)
print(datetime.utcnow().strftime('%H:%M:%S'), quality_status)
# -
monitor_instance_details = wos_client.monitor_instances.get(monitor_instance_id=quality_monitor_instance_id).result
print(monitor_instance_details)
# ### Run an on-demand evaluation
# +
# Check Quality monitor instance details
monitor_instance_details = wos_client.monitor_instances.get(monitor_instance_id=quality_monitor_instance_id).result
print(monitor_instance_details)
# +
# Trigger on-demand run
monitoring_run_details = wos_client.monitor_instances.run(monitor_instance_id=quality_monitor_instance_id).result
monitoring_run_id=monitoring_run_details.metadata.id
print(monitoring_run_details)
# +
# Check run status
quality_run_status = None
while quality_run_status not in ("finished", "error"):
monitoring_run_details = wos_client.monitor_instances.get_run_details(monitor_instance_id=quality_monitor_instance_id, monitoring_run_id=monitoring_run_id).result
quality_run_status = monitoring_run_details.entity.status.state
if quality_run_status not in ("finished", "error"):
print(datetime.utcnow().strftime("%H:%M:%S"), quality_run_status)
time.sleep(30)
print(datetime.utcnow().strftime("%H:%M:%S"), quality_run_status)
# -
# ### Display quality metrics
wos_client.monitor_instances.show_metrics(monitor_instance_id=quality_monitor_instance_id)
# # 5. Drift monitoring <a name="drift"></a>
# ### Enable the drift monitor
#
# In the following code cell, type a path to the drift configuration tar ball.
wos_client.monitor_instances.upload_drift_model(
model_path="/path/to/dir/containing/drift.tar.gz",
data_mart_id=data_mart_id,
subscription_id=subscription_id
).result
# In the following code cell, default values are set for the drift monitor. You can change the default values by updating the values in the parameters section. The `min_samples` parameter controls the number of records that triggers the drift monitor to run. The `drift_threshold` parameter sets the threshold in decimal format for the drift percentage to trigger an alert. The `train_drift_model` parameter controls whether to re-train the model based on the drift analysis.
#
#
# +
import time
target = Target(
target_type=TargetTypes.SUBSCRIPTION,
target_id=subscription_id
)
parameters = {
"min_samples": 5000,
"drift_threshold": 0.05,
"train_drift_model": False
}
drift_monitor_details = wos_client.monitor_instances.create(
data_mart_id=data_mart_id,
monitor_definition_id=wos_client.monitor_definitions.MONITORS.DRIFT.ID,
target=target,
parameters=parameters
).result
drift_monitor_instance_id = drift_monitor_details.metadata.id
print(drift_monitor_details)
# -
# ### Check monitor instance status
# +
drift_status = None
while drift_status not in ("active", "error"):
monitor_instance_details = wos_client.monitor_instances.get(monitor_instance_id=drift_monitor_instance_id).result
drift_status = monitor_instance_details.entity.status.state
if drift_status not in ("active", "error"):
print(datetime.utcnow().strftime('%H:%M:%S'), drift_status)
time.sleep(30)
print(datetime.utcnow().strftime('%H:%M:%S'), drift_status)
# -
# ### Run an on-demand evaluation
# +
# Check Drift monitor instance details
monitor_instance_details = wos_client.monitor_instances.get(monitor_instance_id=drift_monitor_instance_id).result
print(monitor_instance_details)
# +
# Trigger on-demand run
monitoring_run_details = wos_client.monitor_instances.run(monitor_instance_id=drift_monitor_instance_id).result
monitoring_run_id=monitoring_run_details.metadata.id
print(monitoring_run_details)
# +
# Check run status
drift_run_status = None
while drift_run_status not in ("finished", "error"):
monitoring_run_details = wos_client.monitor_instances.get_run_details(monitor_instance_id=drift_monitor_instance_id, monitoring_run_id=monitoring_run_id).result
drift_run_status = monitoring_run_details.entity.status.state
if drift_run_status not in ("finished", "error"):
print(datetime.utcnow().strftime("%H:%M:%S"), drift_run_status)
time.sleep(30)
print(datetime.utcnow().strftime("%H:%M:%S"), drift_run_status)
# -
# ### Display drift metrics
wos_client.monitor_instances.show_metrics(monitor_instance_id=drift_monitor_instance_id)
# ## Congratulations!
#
# You have finished the Batch demo for IBM Watson OpenScale using Remote Apache Spark. You can now view the [Watson OpenScale Dashboard](https://url-to-your-cp4d-cluster/aiopenscale). Click the tile for the **German Credit model** to see quality and drift monitors. Click the timeseries graph to get detailed information on transactions during a specific time window.
|
Cloud Pak for Data/Batch Support/4.0/Watson OpenScale and Batch Support with Remote Spark - JDBC.ipynb
|