code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exporting data from BigQuery to Google Cloud Storage
#
# In this notebook, we export BigQuery data to GCS so that we can reuse our Keras model that was developed on CSV data.
# Uncomment the following line if you are running the notebook locally:
# !sudo chown -R jupyter:jupyter /home/jupyter/training-data-analyst
# +
# #%load_ext google.cloud.bigquery
# +
import os
from google.cloud import bigquery
# -
# Change the following cell as necessary:
# +
# Change with your own bucket and project below:
BUCKET = "<BUCKET>"
PROJECT = "<PROJECT>"
OUTDIR = "gs://{bucket}/taxifare/data".format(bucket=BUCKET)
os.environ['BUCKET'] = BUCKET
os.environ['OUTDIR'] = OUTDIR
os.environ['PROJECT'] = PROJECT
# -
# ## Create BigQuery tables
# If you haven not already created a BigQuery dataset for our data, run the following cell:
# +
bq = bigquery.Client(project = PROJECT)
dataset = bigquery.Dataset(bq.dataset("taxifare"))
try:
bq.create_dataset(dataset)
print("Dataset created")
except:
print("Dataset already exists")
# -
# Let's create a table with 1 million examples.
#
# Note that the order of columns is exactly what was in our CSV files.
# +
# %%bigquery
CREATE OR REPLACE TABLE taxifare.feateng_training_data AS
SELECT
(tolls_amount + fare_amount) AS fare_amount,
pickup_datetime,
pickup_longitude AS pickuplon,
pickup_latitude AS pickuplat,
dropoff_longitude AS dropofflon,
dropoff_latitude AS dropofflat,
passenger_count*1.0 AS passengers,
'unused' AS key
FROM `nyc-tlc.yellow.trips`
WHERE ABS(MOD(FARM_FINGERPRINT(CAST(pickup_datetime AS STRING)), 1000)) = 1
AND
trip_distance > 0
AND fare_amount >= 2.5
AND pickup_longitude > -78
AND pickup_longitude < -70
AND dropoff_longitude > -78
AND dropoff_longitude < -70
AND pickup_latitude > 37
AND pickup_latitude < 45
AND dropoff_latitude > 37
AND dropoff_latitude < 45
AND passenger_count > 0
# -
# Make the validation dataset be 1/10 the size of the training dataset.
# +
# %%bigquery
CREATE OR REPLACE TABLE taxifare.feateng_valid_data AS
SELECT
(tolls_amount + fare_amount) AS fare_amount,
pickup_datetime,
pickup_longitude AS pickuplon,
pickup_latitude AS pickuplat,
dropoff_longitude AS dropofflon,
dropoff_latitude AS dropofflat,
passenger_count*1.0 AS passengers,
'unused' AS key
FROM `nyc-tlc.yellow.trips`
WHERE ABS(MOD(FARM_FINGERPRINT(CAST(pickup_datetime AS STRING)), 10000)) = 2
AND
trip_distance > 0
AND fare_amount >= 2.5
AND pickup_longitude > -78
AND pickup_longitude < -70
AND dropoff_longitude > -78
AND dropoff_longitude < -70
AND pickup_latitude > 37
AND pickup_latitude < 45
AND dropoff_latitude > 37
AND dropoff_latitude < 45
AND passenger_count > 0
# -
# ## Export the tables as CSV files
# + language="bash"
#
# echo "Deleting current contents of $OUTDIR"
# gsutil -m -q rm -rf $OUTDIR
#
# echo "Extracting training data to $OUTDIR"
# bq --location=US extract \
# --destination_format CSV \
# --field_delimiter "," --noprint_header \
# taxifare.feateng_training_data \
# $OUTDIR/taxi-train-*.csv
#
# echo "Extracting validation data to $OUTDIR"
# bq --location=US extract \
# --destination_format CSV \
# --field_delimiter "," --noprint_header \
# taxifare.feateng_valid_data \
# $OUTDIR/taxi-valid-*.csv
#
# gsutil ls -l $OUTDIR
# -
# !gsutil cat gs://$BUCKET/taxifare/data/taxi-train-000000000000.csv | head -2
# Copyright 2019 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
|
courses/machine_learning/deepdive2/building_production_ml_systems/solutions/0_export_data_from_bq_to_gcs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import tensorflow as tf
# ### Load the Corpus
# ##### Get book names
# +
import glob
book_filenames = sorted(glob.glob("/data/*.txt"))
print("Found {} books".format(len(book_filenames)))
# -
# ##### Combine books into a string
# +
import codecs
corpus_raw = u""
for filename in book_filenames:
with codecs.open(filename, 'r', 'utf-8') as book_file:
corpus_raw += book_file.read()
print("Corpus is {} characters long".format(len(corpus_raw)))
# -
# ### Process Corpus
# ##### Create lookup tables
def create_lookup_tables(text):
"""
Create lookup tables for vocab
:param text: The GOT text split into words
:return: A tuple of dicts (vocab_to_int, int_to_vocab)
"""
vocab = set(text)
int_to_vocab = {key: word for key, word in enumerate(vocab)}
vocab_to_int = {word: key for key, word in enumerate(vocab)}
return vocab_to_int, int_to_vocab
# ##### Tokenize punctuation
def token_lookup():
"""
Generate a dict to map punctuation into a token
:return: dictionary mapping puncuation to token
"""
return {
'.': '||period||',
',': '||comma||',
'"': '||quotes||',
';': '||semicolon||',
'!': '||exclamation-mark||',
'?': '||question-mark||',
'(': '||left-parentheses||',
')': '||right-parentheses||',
'--': '||emm-dash||',
'\n': '||return||'
}
# ##### Process and save data
# +
import pickle
token_dict = token_lookup()
for token, replacement in token_dict.items():
corpus_raw = corpus_raw.replace(token, ' {} '.format(replacement))
corpus_raw = corpus_raw.lower()
corpus_raw = corpus_raw.split()
vocab_to_int, int_to_vocab = create_lookup_tables(corpus_raw)
corpus_int = [vocab_to_int[word] for word in corpus_raw]
pickle.dump((corpus_int, vocab_to_int, int_to_vocab, token_dict), open('preprocess.p', 'wb'))
# -
# # Build the Network
# ### Batch the Data
def get_batches(int_text, batch_size, seq_length):
"""
Return batches of input and target data
:param int_text: text with words replaced by their ids
:param batch_size: the size that each batch of data should be
:param seq_length: the length of each sequence
:return: batches of data as a numpy array
"""
words_per_batch = batch_size * seq_length
num_batches = len(int_text)//words_per_batch
int_text = int_text[:num_batches*words_per_batch]
y = np.array(int_text[1:] + [int_text[0]])
x = np.array(int_text)
x_batches = np.split(x.reshape(batch_size, -1), num_batches, axis=1)
y_batches = np.split(y.reshape(batch_size, -1), num_batches, axis=1)
batch_data = list(zip(x_batches, y_batches))
return np.array(batch_data)
# ### Hyperparameters
num_epochs = 10000
batch_size = 512
rnn_size = 512
num_layers = 3
keep_prob = 0.7
embed_dim = 512
seq_length = 30
learning_rate = 0.001
save_dir = './save'
# ### Build the Graph
train_graph = tf.Graph()
with train_graph.as_default():
# Initialize input placeholders
input_text = tf.placeholder(tf.int32, [None, None], name='input')
targets = tf.placeholder(tf.int32, [None, None], name='targets')
lr = tf.placeholder(tf.float32, name='learning_rate')
# Calculate text attributes
vocab_size = len(int_to_vocab)
input_text_shape = tf.shape(input_text)
# Build the RNN cell
lstm = tf.contrib.rnn.BasicLSTMCell(num_units=rnn_size)
drop_cell = tf.contrib.rnn.DropoutWrapper(lstm, output_keep_prob=keep_prob)
cell = tf.contrib.rnn.MultiRNNCell([drop_cell] * num_layers)
# Set the initial state
initial_state = cell.zero_state(input_text_shape[0], tf.float32)
initial_state = tf.identity(initial_state, name='initial_state')
# Create word embedding as input to RNN
embed = tf.contrib.layers.embed_sequence(input_text, vocab_size, embed_dim)
# Build RNN
outputs, final_state = tf.nn.dynamic_rnn(cell, embed, dtype=tf.float32)
final_state = tf.identity(final_state, name='final_state')
# Take RNN output and make logits
logits = tf.contrib.layers.fully_connected(outputs, vocab_size, activation_fn=None)
# Calculate the probability of generating each word
probs = tf.nn.softmax(logits, name='probs')
# Define loss function
cost = tf.contrib.seq2seq.sequence_loss(
logits,
targets,
tf.ones([input_text_shape[0], input_text_shape[1]])
)
# Learning rate optimizer
optimizer = tf.train.AdamOptimizer(learning_rate)
# Gradient clipping to avoid exploding gradients
gradients = optimizer.compute_gradients(cost)
capped_gradients = [(tf.clip_by_value(grad, -1., 1.), var) for grad, var in gradients if grad is not None]
train_op = optimizer.apply_gradients(capped_gradients)
# ### Train the Network
# +
import time
pickle.dump((seq_length, save_dir), open('params.p', 'wb'))
batches = get_batches(corpus_int, batch_size, seq_length)
num_batches = len(batches)
start_time = time.time()
with tf.Session(graph=train_graph) as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(num_epochs):
state = sess.run(initial_state, {input_text: batches[0][0]})
for batch_index, (x, y) in enumerate(batches):
feed_dict = {
input_text: x,
targets: y,
initial_state: state,
lr: learning_rate
}
train_loss, state, _ = sess.run([cost, final_state, train_op], feed_dict)
time_elapsed = time.time() - start_time
print('Epoch {:>3} Batch {:>4}/{} train_loss = {:.3f} time_elapsed = {:.3f} time_remaining = {:.0f}'.format(
epoch + 1,
batch_index + 1,
len(batches),
train_loss,
time_elapsed,
((num_batches * num_epochs)/((epoch + 1) * (batch_index + 1))) * time_elapsed - time_elapsed))
# save model every 10 epochs
if epoch % 10 == 0:
saver = tf.train.Saver()
saver.save(sess, save_dir)
print('Model Trained and Saved')
# -
# ### Checkpoint
# +
import tensorflow as tf
import numpy as np
import pickle
corpus_int, vocab_to_int, int_to_vocab, token_dict = pickle.load(open('preprocess.p', mode='rb'))
seq_length, save_dir = pickle.load(open('params.p', mode='rb'))
# -
# # Generate GOT Text
# ### Pick a Random Word
def pick_word(probabilities, int_to_vocab):
"""
Pick the next word with some randomness
:param probabilities: Probabilites of the next word
:param int_to_vocab: Dictionary of word ids as the keys and words as the values
:return: String of the predicted word
"""
return np.random.choice(list(int_to_vocab.values()), 1, p=probabilities)[0]
# ### Load the Graph and Generate
# +
gen_length = 1000
prime_words = 'daenerys'
loaded_graph = tf.Graph()
with tf.Session(graph=loaded_graph) as sess:
# Load the saved model
loader = tf.train.import_meta_graph(save_dir + '.meta')
loader.restore(sess, save_dir)
# Get tensors from loaded graph
input_text = loaded_graph.get_tensor_by_name('input:0')
initial_state = loaded_graph.get_tensor_by_name('initial_state:0')
final_state = loaded_graph.get_tensor_by_name('final_state:0')
probs = loaded_graph.get_tensor_by_name('probs:0')
# Sentences generation setup
gen_sentences = prime_words.split()
prev_state = sess.run(initial_state, {input_text: np.array([[1 for word in gen_sentences]])})
# Generate sentences
for n in range(gen_length):
# Dynamic Input
dyn_input = [[vocab_to_int[word] for word in gen_sentences[-seq_length:]]]
dyn_seq_length = len(dyn_input[0])
# Get Prediction
probabilities, prev_state = sess.run(
[probs, final_state],
{input_text: dyn_input, initial_state: prev_state})
pred_word = pick_word(probabilities[dyn_seq_length-1], int_to_vocab)
gen_sentences.append(pred_word)
# Remove tokens
chapter_text = ' '.join(gen_sentences)
for key, token in token_dict.items():
chapter_text = chapter_text.replace(' ' + token.lower(), key)
print(chapter_text)
# -
# # Save a Chapter
# ### Cleanup Data a Bit
# +
chapter_text = ' '.join(gen_sentences)
for key, token in token_dict.items():
chapter_text = chapter_text.replace(' ' + token.lower(), key)
chapter_text = chapter_text.replace('\n ', '\n')
chapter_text = chapter_text.replace('( ', '(')
chapter_text = chapter_text.replace(' ”', '”')
capitalize_words = ['lannister', 'stark', 'lord', 'ser', 'tyrion', 'jon', '<NAME>', 'daenerys', 'targaryen', 'cersei', 'jaime', 'arya', 'sansa', 'bran', 'rikkon', 'joffrey',
'khal', 'drogo', 'gregor', 'clegane', 'kings landing', 'winterfell', 'the mountain', 'the hound', 'ramsay', 'bolton', 'melisandre', 'shae', 'tyrell',
'margaery', 'sandor', 'hodor', 'ygritte', 'brienne', 'tarth', 'petyr', 'baelish', 'eddard', 'greyjoy', 'theon', 'gendry', 'baratheon', 'baraTheon',
'varys', 'stannis', 'bronn', 'jorah', 'mormont', 'martell', 'oberyn', 'catelyn', 'robb', 'loras', 'missandei', 'tommen', 'robert', 'lady', 'donella', 'redwyne'
'myrcella', 'samwell', 'tarly', 'grey worm', 'podrick', 'osha', 'davos', 'seaworth', 'jared', '<NAME>', 'rickard', 'yoren', 'meryn', 'trant', 'king', 'queen',
'aemon']
for word in capitalize_words:
chapter_text = chapter_text.replace(word, word.lower().title())
# -
# ### Save File
# +
import os
version_dir = './generated-book-v1'
if not os.path.exists(version_dir):
os.makedirs(version_dir)
num_chapters = len([name for name in os.listdir(version_dir) if os.path.isfile(os.path.join(version_dir, name))])
next_chapter = version_dir + '/chapter-' + str(num_chapters + 1) + '.md'
with open(next_chapter, "w") as text_file:
text_file.write(chapter_text)
|
vows-generator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# seaborn中的sns库可以画出很好看的图
# 试用sns.pairplot()画出基于 “iris”数据集各两对属性的图
# +
import seaborn as sns
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
sns.set(style="ticks", color_codes=True)
iris = sns.load_dataset("iris")
g = sns.pairplot(iris, hue="species")
sns.plt.show()
# -
iris
# +
"""get the data corrspoding to 'versicolor' and 'setosa' from iris data"""
s = iris['species']
start = s[s == 'virginica'].index[0]
end = s[s == 'virginica'].index[-1]
df1 = iris.loc[start:end, :]
start = s[s == 'setosa'].index[0]
end = s[s == 'setosa'].index[-1]
df2 = iris.loc[start:end, :]
df = pd.concat([df1,df2],ignore_index=True)
# +
#10-fold cross validation
from sklearn.cross_validation import cross_val_score
from sklearn.linear_model import LogisticRegression
from sklearn import model_selection
from sklearn.metrics import classification_report
#get the test_data and label_data from df
X = df.loc[:,:'petal_width']
y = df.loc[:,'species']
model = LogisticRegression() #choose the logistictRegression
cross_val_score(model,X,y,cv=10)
# -
X_train, X_test, y_train, y_test = model_selection.train_test_split(X, y,test_size=0.5, random_state=42)
model = LogisticRegression()
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
print(classification_report(y_test, y_pred))
# 为何正确率是百分之一百
|
unit3/ch3.4 10-fold and LOO validation assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/IEwaspbusters/KopuruVespaCompetitionIE/blob/main/Competition_subs/2021-04-28_submit/batch_LARVAE/HEX.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # XGBoost Years: Prediction with Mario's Cluster Variable and selected Weather Variables (according to Feature importance), including GridSearchCV (without Population, since it is reflected in Cluster).
# ## Import the Data & Modules
# + colab={"base_uri": "https://localhost:8080/"} id="rt-Jj2BjesTz" outputId="171cecde-0242-4aaf-82b8-0e3458dff994"
# Base packages -----------------------------------
import pandas as pd
import numpy as np
import shap
import warnings
# Data Viz -----------------------------------
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (15, 10) # to set figure size when ploting feature_importance
# XGBoost -------------------------------
import xgboost as xgb
from xgboost import XGBRegressor, plot_importance, plot_tree # built-in function to plot features ordered by their importance & tree
# SKLearn -----------------------------------------
from sklearn import preprocessing # scaling data
from sklearn.model_selection import GridSearchCV
# + code_folding=[]
# Function that checks if final Output is ready for submission or needs revision
def check_data(HEX):
def template_checker(HEX):
submission_df = (HEX["CODIGO MUNICIPIO"].astype("string")+HEX["NOMBRE MUNICIPIO"]).sort_values().reset_index(drop=True)
template_df = (template["CODIGO MUNICIPIO"].astype("string")+template["NOMBRE MUNICIPIO"]).sort_values().reset_index(drop=True)
check_df = pd.DataFrame({"submission_df":submission_df,"template_df":template_df})
check_df["check"] = check_df.submission_df == check_df.template_df
if (check_df.check == False).any():
pd.options.display.max_rows = 112
return check_df.loc[check_df.check == False,:]
else:
return "All Municipality Names and Codes to be submitted match the Template"
print("Submission form Shape is", HEX.shape)
print("Number of Municipalities is", HEX["CODIGO MUNICIPIO"].nunique())
print("The Total 2020 Nests' Prediction is", int(HEX["NIDOS 2020"].sum()))
assert HEX.shape == (112, 3), "Error: Shape is incorrect."
assert HEX["CODIGO MUNICIPIO"].nunique() == 112, "Error: Number of unique municipalities is correct."
return template_checker(HEX)
# + id="9MLidG_FwhYB"
# Importing datasets from GitHub as Pandas Dataframes
queen_train = pd.read_csv("../Feeder_years/WBds03_QUEENtrainYears.csv", encoding="utf-8") #2018+2019 test df
queen_predict = pd.read_csv("../Feeder_years/WBds03_QUEENpredictYears.csv", encoding="utf-8") #2020 prediction df
queen_clusters = pd.read_csv("../auxiliary_files/WBds_CLUSTERSnests.csv",sep=",")
template = pd.read_csv("../../../Input_open_data/ds01_PLANTILLA-RETO-AVISPAS-KOPURU.csv",sep=";", encoding="utf-8")
# -
# ## Further Clean the Data
# ### Filter according to assumptions (Clusters & Relevant Municipalities)
# +
# Adding cluster labels
queen_train = pd.merge(queen_train, queen_clusters, how = 'left', left_on = 'municip_code', right_on = 'municip_code')
queen_predict = pd.merge(queen_predict, queen_clusters, how = 'left', left_on = 'municip_code', right_on = 'municip_code')
# +
# Remove the Municipalities to which we did not assign a Cluster, since there was not reliable data for us to predict -> Bilbao
queen_train = queen_train.loc[queen_train.municip_code != 48020,:].copy()
queen_predict = queen_predict.loc[queen_predict.municip_code != 48020,:].copy()
# -
# ### Arrange data into a features matrix and target vector
# +
# selecting the train X & y variables
# Y will be the response variable (filter for the number of wasp nests - waspbust_id)
y_train = queen_train.NESTS
# X will be the explanatory variables. Remove response variable and non desired categorical columns such as (municip code, year, etc...)
X_train = queen_train.iloc[:,4:-10].drop(["station_code"],axis=1).copy()
X_train["cluster"] = queen_train.Cluster.copy()
# We want to predict our response variable (number of nests in 2020). Remove response variable and non desired categorical columns such as (municip code, year, etc...)
X_test = queen_predict.iloc[:,4:-10].drop("station_code",axis=1).copy()
X_test["cluster"] = queen_predict.Cluster.copy()
# -
# Check if the shape of the features and their labels match or if there are errors raised
# +
# Perform checks of features labels & their shapes
assert X_test.shape[1] == X_train.shape[1], "Error: Number of columns do not match!"
assert (X_test.columns == X_train.columns).any(), "Error: Columns labels do not match"
assert y_train.shape == (222,), "Error: y shape is incorrect!"
# -
# ## Model Step 1: Finding out the Relevant Variables
# ### Scale the Data for better Regressions
# +
# Scale the datasets using MinMaxScaler
X_train_scaled = preprocessing.minmax_scale(X_train) # this creates a numpy array
X_train_scaled = pd.DataFrame(X_train_scaled,index=X_train.index,columns=X_train.columns) # create a Pandas Dataframe == X
# -
# #### Choose a class of model by importing the appropriate estimator class
# selecting the XGBoost model and fitting with the train data
model = XGBRegressor(random_state=0, objective="reg:squarederror")
# ### Use GridSearchCV to find out the best hyperparameters for our XGBoost model with our Fitted Data
# +
# Use GridSearchCV that will automatically split the data and give us the best estimator by:
#1) Establishing hyperparameters to change
param_grid = {
"learning_rate": [0.01],
"max_depth": [3],
"gamma" : [0],
"max_delta_step" : [3],
"min_child_weight": [6],
"subsample": [0.8],
"colsample_bytree": [0.2],
"reg_lambda" : [1],
"reg_alpha" : [1],
"n_estimators": [5000],
"scale_pos_weight" : [3]
}
warnings.filterwarnings(action='ignore', category=UserWarning)
grid = GridSearchCV(model, param_grid, cv=3)
#2) Fitting the model with our desired data and check for best results
grid.fit(X_train_scaled, y_train)
#) Retrieve the summary of GridSearchCV for analysis
print(F"The number homogeneous splits conducted by GridSearchCV are: {grid.n_splits_}.")
print(F"The best hyperparameters found were: {grid.best_params_}.")
print(F"The best score found was: {grid.best_score_}.")
# +
# Reset warnigns to default (this is used to suppred a warning message from XGBoost model and avoid converting X_train to numpy to keep features name)
warnings.filterwarnings(action='default', category=UserWarning)
# -
# #### Fit the model to your data by calling the `.fit()` method of the model instance using the best Hyperparameters
# +
# selecting the XGBoost model according to the best hyperparameters found in GridSearchCV and fitting with the train data
model = grid.best_estimator_
model.fit(X_train_scaled, y_train)
# -
# #### Selecting the Relevant Variables and filtering according to the results
# +
# Plot the Relevant Variables in order to filter the relevant ones per Cluster
# In built plot method from XGBoost
xgb.plot_importance(model,height=0.5,xlabel="F-Score",ylabel="Feature Importance",grid=False)
plt.show()
plt.figure(figsize=(15,7))
# Feature importance from model.feature_importances built-in attribute
sorted_idx = model.feature_importances_.argsort()
plt.subplot(1,3,1)
plt.barh(X_train_scaled.columns[sorted_idx], model.feature_importances_[sorted_idx])
plt.xlabel("Xgboost Feature Importance", fontsize=13)
# Shap library Summaries
# Shap values
explainer = shap.TreeExplainer(model)
shap_values = explainer.shap_values(X_train_scaled)
plt.subplot(1,3,2)
shap.summary_plot(shap_values, X_train_scaled, plot_type="bar", plot_size=None, show=False)
plt.subplot(1,3,3)
shap.summary_plot(shap_values, X_train_scaled, plot_size=None, show=False)
plt.subplots_adjust(wspace=10.0)
plt.tight_layout()
plt.show()
#fig, ax = plt.subplots(figsize=(30, 30))
xgb.plot_tree(model, num_trees=-1)
plt.show()
# +
# selecting the XGBoost model and fitting with the train data without the irrelevant variables
X_train = queen_train.loc[:,["colonies_amount","food_fruit","weath_days_rain","food_txakoli","Cluster"]].copy()
X_test = queen_predict.loc[:,["colonies_amount","food_fruit","weath_days_rain","food_txakoli","Cluster"]].copy()
# -
# ## Model Step 2: Prediction of 2020 Nests
#
# **Note: Variables are no longer scaled**
# ### Scale the Data for better Regressions
# +
# Scale the datasets using MinMaxScaler
# Train set
X_train_scaled = preprocessing.minmax_scale(X_train) # this creates a numpy array
X_train = pd.DataFrame(X_train_scaled,index=X_train.index,columns=X_train.columns) # create a Pandas Dataframe == X
# Test set
X_test_scaled = preprocessing.minmax_scale(X_test) # this creates a numpy array
X_test = pd.DataFrame(X_test_scaled,index=X_test.index,columns=X_test.columns) # create a Pandas Dataframe == X
# -
# ### Choose a class of model by importing the appropriate estimator class
# +
# selecting the XGBoost model and fitting with the train data
model = XGBRegressor(random_state=0, objective="reg:squarederror")
# -
# ### Use GridSearchCV to find out the best hyperparameters for our XGBoost model with our Fitted Data
# +
# Use GridSearchCV that will automatically split the data and give us the best estimator by:
#1) Establishing hyperparameters to change
param_grid = {
"learning_rate": [0.2],
"n_estimators": [1000],
"max_depth": [6],
"gamma" : [0.5],
"scale_pos_weight" : [1.5],
"max_delta_step" : [1],
"min_child_weight": [7],
"subsample": [0.7],
"colsample_bytree": [0.7],
"reg_alpha" : [1],
"reg_lambda" : [2]
}
warnings.filterwarnings(action='ignore', category=UserWarning)
grid = GridSearchCV(model, param_grid, cv=3)
#2) Fitting the model with our desired data and check for best results
grid.fit(X_train, y_train)
#) Retrieve the summary of GridSearchCV for analysis
print(F"The number homogeneous splits conducted by GridSearchCV are: {grid.n_splits_}.")
print(F"The best hyperparameters found were: {grid.best_params_}.")
print(F"The best score found was: {grid.best_score_}.")
# +
# Reset warnigns to default (this is used to suppred a warning message from XGBoost model and avoid converting X_train to numpy to keep features name)
warnings.filterwarnings(action='default', category=UserWarning)
# -
# ### Fit the model to your data by calling the `.fit()` method of the model instance
# +
# selecting the XGBoost model according to the best hyperparameters found in GridSearchCV and re-fitting with the train data with the appropriate variables
model = grid.best_estimator_
model.fit(X_train, y_train)
# -
# ### Tree visualization
fig, ax = plt.subplots(figsize=(30, 30))
xgb.plot_tree(model, num_trees=-1, ax=ax)
plt.show()
# ### Apply the model to new data:
#
# - For supervised learning, predict labels for unknown data using the `.predict()` method
# +
# make a prediction
prediction_2020 = model.predict(X_test)
# -
# ## Add Each Cluster Predictions to the original DataFrame and Save it as a `.csv file`
# Create a new Column with the 2020 prediction
queen_predict["nests_2020"] = prediction_2020
# +
# Create a new DataFrame with the Municipalities to insert manualy
HEX_aux = pd.DataFrame({"CODIGO MUNICIPIO":[48020],\
"NOMBRE MUNICIPIO":["Bilbao"],\
"NIDOS 2020":[0]})
# + id="Z3PcQ4UnACCA"
HEX = queen_predict.loc[:,["municip_code","municip_name_x","nests_2020"]].round() # create a new Dataframe for Kopuru submission
HEX.columns = ["CODIGO MUNICIPIO","NOMBRE MUNICIPIO","NIDOS 2020"] # change column names to Spanish (Decidata template)
HEX = HEX.append(HEX_aux, ignore_index=True) # Add rows of municipalities to add manually
# +
# Final check
check_data(HEX)
# +
# reset max_rows to default values (used in function to see which rows did not match template)
pd.reset_option("max_rows")
# + id="uiPq7zXi0STt"
# Save the new dataFrame as a .csv in the current working directory on Windows
HEX.to_csv("WaspBusters_20210519_XGyears_scaledX.csv", index=False)
|
B_Submissions_Kopuru_competition/2021-06-09_submit FINAL/XGBoost_XGeirinhoost/workerbee05_HEX_XGyears_BASE_scaledX.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > **Tip**: Welcome to the Investigate a Dataset project! You will find tips in quoted sections like this to help organize your approach to your investigation. Before submitting your project, it will be a good idea to go back through your report and remove these sections to make the presentation of your work as tidy as possible. First things first, you might want to double-click this Markdown cell and change the title so that it reflects your dataset and investigation.
#
# # Project: Investigate a Dataset "TMDb Movie Data"
#
# ## Table of Contents
# <ul>
# <li><a href="#intro">Introduction</a></li>
# <li><a href="#wrangling">Data Wrangling</a></li>
# <li><a href="#eda">Exploratory Data Analysis</a></li>
# <li><a href="#conclusions">Conclusions</a></li>
# </ul>
# <a id='intro'></a>
# ## Introduction
#
# > **Tip**: In this section of the report, provide a brief introduction to the dataset you've selected for analysis. At the end of this section, describe the questions that you plan on exploring over the course of the report. Try to build your report around the analysis of at least one dependent variable and three independent variables. If you're not sure what questions to ask, then make sure you familiarize yourself with the dataset, its variables and the dataset context for ideas of what to explore.
#
# > If you haven't yet selected and downloaded your data, make sure you do that first before coming back here. In order to work with the data in this workspace, you also need to upload it to the workspace. To do so, click on the jupyter icon in the upper left to be taken back to the workspace directory. There should be an 'Upload' button in the upper right that will let you add your data file(s) to the workspace. You can then click on the .ipynb file name to come back here.
# ## Introduction
# This data set contains information about 10.8K movies collected from The Movie Database (TMDb), including user ratings and revenue, release date and other data in 21 Columns
# +
# Use this cell to set up import statements for all of the packages that you
# plan to use.
# Remember to include a 'magic word' so that your visualizations are plotted
# inline with the notebook. See this page for more:
# http://ipython.readthedocs.io/en/stable/interactive/magics.html
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from datetime import datetime as dt
% matplotlib inline
# import data and check first 5 columns
df = pd.read_csv ("tmdb-movies.csv")
df.head()
# -
# ### After reviewing the data, there are a lot of questions to be asked as below:-
#
# 1- Who are the directors with top numbers of movies
#
# 2- What is the change in movie production numbers over years
#
# 3- What is the relationship between Budget and Revenue
#
# 4- What is the runtime movie distribution
#
# 5- Which month has the most average revenue
#
# 6- What is the relationship between budget and revenue over years
#
# 7- Which Month has the biggest number of movies released
#
# 8- What are the top 5 movies in terms of budget, revenue and popularity
#
# 9- Which movies have the highest profit
# <a id='wrangling'></a>
# ## Data Wrangling
#
# > **Tip**: In this section of the report, you will load in the data, check for cleanliness, and then trim and clean your dataset for analysis. Make sure that you document your steps carefully and justify your cleaning decisions.
#
# ### General Properties
# First we need to clean the data, so to check rows and columns for data that should be dropped or cleaned.
# Load your data and print out a few lines. Perform operations to inspect data
# types and look for instances of missing or possibly errant data.
#data already loaded before
#checking data shape
df.shape
# ### Check for duplicated lines
df.duplicated().sum()
# ### Check Data for more detailes
df.info()
df.describe()
df.dtypes
df.hist(figsize=(10,10));
#
# #### Data Wrangling Conclusion
#
#
# By the end of data Wrangling, below actions should be done in data cleaning
#
# 1- Remove duplicated lines
#
# 2- Remove unneeded columns
#
# 3- Remove lines with non-valid data ( like zero screen budget or revenue or runtime)
#
# 4- Remove lines with NA Values
#
# 5- Change release_date to date instead of strings
#
#
# > **Tip**: You should _not_ perform too many operations in each cell. Create cells freely to explore your data. One option that you can take with this project is to do a lot of explorations in an initial notebook. These don't have to be organized, but make sure you use enough comments to understand the purpose of each code cell. Then, after you're done with your analysis, create a duplicate notebook where you will trim the excess and organize your steps so that you have a flowing, cohesive report.
#
# > **Tip**: Make sure that you keep your reader informed on the steps that you are taking in your investigation. Follow every code cell, or every set of related code cells, with a markdown cell to describe to the reader what was found in the preceding cell(s). Try to make it so that the reader can then understand what they will be seeing in the following cell(s).
#
# ### Data Cleaning
# ## 1- Remove duplicated lines
df.drop_duplicates(inplace = True)
# ## 2- Remove unneeded columns
df.drop(['id', 'imdb_id', 'cast', 'homepage', 'tagline', 'keywords', 'overview','production_companies'], axis = 1, inplace = True)
df.info()
# ## 3- Remove lines with non-valid data
df = df[df['budget'] != 0]
df = df[df['revenue'] != 0]
df.info()
# ## 4- Remove lines with NA Values
df.dropna(axis = 0 , inplace = True,subset = ['director'])
df.info()
# ## 5- Change release_date to date
df.release_date = pd.to_datetime(df['release_date'])
df.info()
# #### Note: More than 6K lines was with missing budget or revenue data
# +
# After discussing the structure of the data and any problems that need to be
# cleaned, perform those cleaning steps in the second part of this section.
# -
# <a id='eda'></a>
# ## Exploratory Data Analysis
#
# > **Tip**: Now that you've trimmed and cleaned your data, you're ready to move on to exploration. Compute statistics and create visualizations with the goal of addressing the research questions that you posed in the Introduction section. It is recommended that you be systematic with your approach. Look at one variable at a time, and then follow it up by looking at relationships between variables.
#
#
df.describe()
# ### 1- Who are the directors with top numbers of movies
# +
# Use this, and more code cells, to explore your data. Don't forget to add
# Markdown cells to document your observations and findings.
df['director'].value_counts().sort_values(ascending=False).head(10).plot(kind = 'pie', figsize= (10,10));
plt.xlabel('Runtime of Movies',size= 15);
plt.ylabel('Number of Movies',size= 15);
plt.title('Runtime distribution of movies',size= 20);
# -
df.groupby('director')['revenue'].mean().sort_values(ascending=False).head()
df['director'].value_counts().head(10)
# #### <NAME> is the top director with 27 Movies, then clint eastwood, data is represented in the piechart
# ### 2- What is the change in movie production numbers over years
# +
year_ind = df['release_year'].value_counts().sort_index();
plt.figure(figsize=(9,6))
plt.plot(year_ind);
plt.title('Movies per year',size= 20);
plt.xlabel('Year',size= 15);
plt.ylabel('Number of movies',size= 15);
# -
# #### number of movies release increase over time incrementally
# ### 3- What is the relationship between Budget and Revenue
# +
def corfun(column1, column2):
return df[column1].corr(df[column2])
corfun('budget','revenue')
# -
df.plot(x='budget',y='revenue',kind='scatter',figsize=(8,8));
plt.title('Budget vs Revenue',size= 20)
plt.xlabel('Budget in 100 millions',size= 15);
plt.ylabel('Revenue in billions',size= 15);
# ##### There is a Positive relationship between budget and revenue appearing in the scatter chart and in the Correlation equation above (.688)
# ### 4- What is the runtime movie distribution
# +
plt.figure(figsize=(7,4))
plt.xlabel('Runtime of Movies',size= 15)
plt.ylabel('Number of Movies',size= 15)
plt.title('Runtime distribution of movies',size= 20)
plt.hist(df['runtime'], rwidth = .8, bins =20)
plt.show()
# +
plt.figure(figsize=(9,7))
sns.boxplot(df['runtime'])
plt.xlabel('Runtime of Movies',size= 15)
plt.ylabel('Number of Movies',size= 15)
plt.title('Runtime distribution of movies',size= 20)
plt.show()
# -
corfun('runtime','revenue')
# #### 50% of movies are between 95 to 119 Min
# #### there is no significant correlation between runtime and revenue
# ## 5- Which month has the most average revenue
# +
df['movie_month']=df['release_date'].dt.month
revenue_month = df.groupby('movie_month')['revenue'].mean()
df2=pd.DataFrame()
months=['Jan','Feb','Mar','Apr','May','Jun','Jul','Aug','Sep','Oct','Nov','Dec']
df2['Months']=months
df2['Revenues'] = list(revenue_month.get_values())
df2.plot(x='Months',y='Revenues',kind='bar',figsize=(8,8));
plt.title('Average revenue by month',size=20);
plt.ylabel('Revenue in 100 million',size=15);
plt.xlabel('Months',size=15);
# -
# #### May and June are the top months in terms of average revenue then Nov and Dec, there might be a correlation with the holiday's seasons
# ### 6- What is the relationship between budget and revenue over years
# +
rev_per_year = df.groupby('release_year')['revenue'].sum()
bud_per_year = df.groupby('release_year')['budget'].sum()
df3=pd.DataFrame()
years =df.release_year.unique()
df3['Years']=years
df3['Revenues'] = list(rev_per_year.get_values())
df3['Budget'] = list(bud_per_year.get_values())
df3.plot(x='Years',y=['Revenues','Budget'],kind='bar',figsize=(10,10));
plt.title('Budget Vs Revenue per year',size=20);
plt.ylabel('Revenue & Budget',size=15);
plt.xlabel('Years',size=15);
# -
# #### the postive correlation between budget and revenue is clear over years
# ### 7- Which Month has the biggest number of movies released
numreleases_month= df.movie_month.value_counts().sort_index()
df4=pd.DataFrame()
df4['Months']=months
df4['Number of releases']=numreleases_month.get_values()
df4.plot(x='Months',y='Number of releases',kind='bar');
plt.title('Months vs number of movie releases', size = 20);
plt.ylabel('Number of movie releases', size = 15);
plt.xlabel('Month', size = 15);
# #### Sep is the highest number of movie releases over time, then dec in the second place
# ### 8- What are the top 5 movies in terms of budget, revenue and popularity
# +
df.nlargest(5,'revenue')
# -
df.nlargest(5,'budget')
df.nlargest(5,'popularity')
# ## 9- Which movies have the highest profit
df['profit']=df['revenue']-df['budget']
df.nlargest(5,'profit')
corfun('profit','revenue')
# #### The higher the profit the higher the revenue, relationship is clear with 97.9% positive correlation
# #### it's notable that Jurassic World appear as one of the top five movies in terms of popularity, revenue and profit
# <a id='conclusions'></a>
# ## Conclusions
#
# > **Tip**: Finally, summarize your findings and the results that have been performed. Make sure that you are clear with regards to the limitations of your exploration. If you haven't done any statistical tests, do not imply any statistical conclusions. And make sure you avoid implying causation from correlation!
#
# > **Tip**: Once you are satisfied with your work here, check over your report to make sure that it is satisfies all the areas of the rubric (found on the project submission page at the end of the lesson). You should also probably remove all of the "Tips" like this one so that the presentation is as polished as possible.
#
# ## Submitting your Project
#
# > Before you submit your project, you need to create a .html or .pdf version of this notebook in the workspace here. To do that, run the code cell below. If it worked correctly, you should get a return code of 0, and you should see the generated .html file in the workspace directory (click on the orange Jupyter icon in the upper left).
#
# > Alternatively, you can download this report as .html via the **File** > **Download as** submenu, and then manually upload it into the workspace directory by clicking on the orange Jupyter icon in the upper left, then using the Upload button.
#
# > Once you've done this, you can submit your project by clicking on the "Submit Project" button in the lower right here. This will create and submit a zip file with this .ipynb doc and the .html or .pdf version you created. Congratulations!
# ## Conclusions
#
# After checking the data, we reached somed solide facts, and another conclusions that might be useful
#
# During the given period, top directors with highest numbers of movies are ( <NAME>, <NAME>, <NAME>, <NAME>, <NAME>)
# while top directors in terms of revenue are (<NAME>|<NAME>,<NAME>|<NAME>, <NAME>,<NAME>, <NAME>)
# Which indicates that number of movies per director doesn't imply higher revenue
#
#
# Number of movies is increasing over time, which indicates the success of the industry, even if there are few failures in some movies revenue.
#
#
# The correlation between budget and revenue is positive with correlation coefficient of 68.8%, yet when we add the profit variable we will find that the 5 top movies in terms of profit made, and not on the top list of high budget movies.
#
# May and June are the top months in terms of average revenue then Nov and Dec, there might be a correlation with the holiday's seasons, this might needs more search for the core reason, while the highest month of movie releases is Sep, that implys that number of movies here is not related to higher revenue, which support that there is another reason that boots movie sales during the months mentioned above.
#
#
# ### Limitation
# after cleaning the data, we dropped more than 50% of the rows to have a clean dataframe, which implies a problem in collecting data or in recording it.
# the conclusion we reached is according to what we got, which might be changed in case rest of the data was filled in a right way
#
# Also data might not be update to date, so this suggests that trends or outcomes might be changed by time
#
# Another problem is the currency was not mentioned in the revenue or budget, which will be a problem if there were different currencies, as most of the analysis will be wrong
#
# A lot of movies was related to more than one genre, separating gernes and counting movies twice will create a duplication in data, while creating new genere for every combition would make a lot of genres, which will make the findings inaccurate.
from subprocess import call
call(['python', '-m', 'nbconvert', 'Investigate_a_Dataset.ipynb'])
|
Investigate_a_Dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:eu-west-1:470317259841:image/datascience-1.0
# ---
# # From Unlabeled Data to a Deployed Machine Learning Model: A SageMaker Ground Truth Demonstration for Object Detection
#
# 1. [Introduction](#Introduction)
# 2. [Run a Ground Truth labeling job (time: about 4h)](#Run-a-Ground-Truth-labeling-job)
# 1. [Prepare the data](#Prepare-the-data)
# 2. [Specify the category](#Specify-the-categories)
# 3. [Create the instruction template](#Create-the-instruction-template)
# 4. [Create a private team to test your task ](#Create-a-private-team-to-test-your-task-[OPTIONAL])
# 5. [Define pre-built lambda functions for use in the labeling job](#Define-pre-built-lambda-functions-for-use-in-the-labeling-job)
# 6. [Submit the Ground Truth job request](#Submit-the-Ground-Truth-job-request)
# 1. [Verify your task using a private team ](#Verify-your-task-using-a-private-team-[OPTIONAL])
# 7. [Monitor job progress](#Monitor-job-progress)
# 3. [Analyze Ground Truth labeling job results (time: about 20min)](#Analyze-Ground-Truth-labeling-job-results)
# 1. [Postprocess the output manifest](#Postprocess-the-output-manifest)
# 2. [Plot class histograms](#Plot-class-histograms)
# 3. [Plot annotated images](#Plot-annotated-images)
# 1. [Plot a small output sample](#Plot-a-small-output-sample)
# 2. [Plot the full results](#Plot-the-full-results)
# 4. [Compare Ground Truth results to standard labels (time: about 5min)](#Compare-Ground-Truth-results-to-standard-labels)
# 1. [Compute accuracy](#Compute-accuracy)
# 2. [Plot correct and incorrect annotations](#Plot-correct-and-incorrect-annotations)
# 5. [Train an object detector using Ground Truth labels (time: about 15min)](#Train-an-image-classifier-using-Ground-Truth-labels)
# 6. [Deploy the Model (time: about 20min)](#Deploy-the-Model)
# 1. [Create Model](#Create-Model)
# 2. [Batch Transform](#Batch-Transform)
# 7. [Review](#Review)
# # Introduction
#
# This sample notebook takes you through an end-to-end workflow to demonstrate the functionality of SageMaker Ground Truth. We'll start with an unlabeled image data set, acquire bounding boxes for objects in the images using SageMaker Ground Truth, analyze the results, train an object detector, host the resulting model, and, finally, use it to make predictions. Before you begin, we highly recommend you start a Ground Truth labeling job through the AWS Console first to familiarize yourself with the workflow. The AWS Console offers less flexibility than the API, but is simple to use.
#
# #### Cost and runtime
# You can run this demo in two modes:
#
# 1. Set `RUN_FULL_AL_DEMO = False` in the next cell to label only 100 images. This should cost \$26. **Since Ground Truth's auto-labeling feature only kicks in for datasets of 1000 images or more, this cheaper version of the demo will not use it. Some of the analysis plots might look awkward, but you should still be able to see good results on the human-annotated 100 images.**
#
# 1. Set `RUN_FULL_AL_DEMO = True` in the next cell to label 1000 images. This should cost about $200 given the current [Ground Truth pricing scheme](https://aws.amazon.com/sagemaker/groundtruth/pricing/). In order to reduce the cost, we will use Ground Truth's auto-labeling feature. Auto-labeling uses computer vision to learn from human responses and automatically create bounding boxes for the easiest images at a cheap price. The total end-to-end runtime should be about 6h.
#
#
# #### Prerequisites
# To run this notebook, you can simply execute each cell in order. To understand what's happening, you'll need:
# * An S3 bucket you can write to -- please provide its name in the following cell. The bucket must be in the same region as this SageMaker Notebook instance. You can also change the `EXP_NAME` to any valid S3 prefix. All the files related to this experiment will be stored in that prefix of your bucket.
# * The S3 bucket that you use for this demo must have a CORS policy attached. To learn more about this requirement, and how to attach a CORS policy to an S3 bucket, see [CORS Permission Requirement](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-cors-update.html).
# * Familiarity with Python and [numpy](http://www.numpy.org/).
# * Basic familiarity with [AWS S3](https://docs.aws.amazon.com/s3/index.html).
# * Basic understanding of [AWS Sagemaker](https://aws.amazon.com/sagemaker/).
# * Basic familiarity with [AWS Command Line Interface (CLI)](https://aws.amazon.com/cli/) -- ideally, you should have it set up with credentials to access the AWS account you're running this notebook from.
#
# This notebook has only been tested on a SageMaker notebook instance. The runtimes given are approximate. We used an `ml.m4.xlarge` instance in our tests. However, you can likely run it on a local instance by first executing the cell below on SageMaker and then copying the `role` string to your local copy of the notebook.
# +
# %matplotlib inline
import os
from collections import namedtuple
from collections import defaultdict
from collections import Counter
from datetime import datetime
import itertools
import base64
import glob
import json
import random
import time
import imageio
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import shutil
from matplotlib.backends.backend_pdf import PdfPages
from sklearn.metrics import confusion_matrix
import boto3
import botocore
import sagemaker
from urllib.parse import urlparse
BUCKET = "<YOUR_BUCKET_NAME>"
EXP_NAME = "ground-truth-od-full-demo" # Any valid S3 prefix.
RUN_FULL_AL_DEMO = False # See 'Cost and Runtime' in the Markdown cell above!
# -
# ## Create a private team to test your task
#
# This step requires you to use the AWS Console.
#
# We will create a `private workteam` and add only one user (you) to it. Then, we will modify the Ground Truth API job request to send the task to that workforce. You will then be able to see your annotation job exactly as the public annotators would see it. You could even annotate the whole dataset yourself!
#
# To create a private team:
# 1. Go to `AWS Console > Amazon SageMaker > Labeling workforces`
# 2. Click "Private" and then "Create private team".
# 3. Enter the desired name for your private workteam.
# 4. Select "Create a new Amazon Cognito user group" and click "Create private team."
# 5. The AWS Console should now return to `AWS Console > Amazon SageMaker > Labeling workforces`.
# 6. Click on "Invite new workers" in the "Workers" tab.
# 7. Enter your own email address in the "Email addresses" section and click "Invite new workers."
# 8. Click on your newly created team under the "Private teams" tab.
# 9. Select the "Workers" tab and click "Add workers to team."
# 10. Select your email and click "Add workers to team."
# 11. The AWS Console should again return to `AWS Console > Amazon SageMaker > Labeling workforces`. Your newly created team should be visible under "Private teams". Next to it you will see an `ARN` which is a long string that looks like `arn:aws:sagemaker:region-name-123456:workteam/private-crowd/team-name`. Copy this ARN into the cell below.
# 12. You should get an email from `<EMAIL>` that contains your workforce username and password.
# 13. In `AWS Console > Amazon SageMaker > Labeling workforces > Private`, click on the URL under `Labeling portal sign-in URL`. Use the email/password combination from the previous step to log in (you will be asked to create a new, non-default password).
#
# That's it! This is your private worker's interface. When we create a verification task in [Verify your task using a private team](#Verify-your-task-using-a-private-team-[OPTIONAL]) below, your task should appear in this window. You can invite your colleagues to participate in the labeling job by clicking the "Invite new workers" button.
#
# The [SageMaker Ground Truth documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-workforce-management-private.html) has more details on the management of private workteams.
private_workteam_arn = "<YOUR_PRIVATE_WORKTEAM_ARN>""
# Make sure the bucket is in the same region as this notebook.
role = sagemaker.get_execution_role()
region = boto3.session.Session().region_name
s3 = boto3.client("s3")
bucket_region = s3.head_bucket(Bucket=BUCKET)["ResponseMetadata"]["HTTPHeaders"][
"x-amz-bucket-region"
]
assert (
bucket_region == region
), "Your S3 bucket {} and this notebook need to be in the same region.".format(BUCKET)
# # Run a Ground Truth labeling job
#
# **This section should take about 4 hours to complete.**
#
# We will first run a labeling job. This involves several steps: collecting the images we want annotated, creating instructions, and writing a labeling job specification. In addition, we highly recommend that you run a (free) mock job using a private workforce before you submit any job to the public workforce. This notebook will explain how to do that as an optional step. Using a public workforce, this section should take about 4 hours. However, this will vary depending on the availability of workers.
#
# ### Prepare the data
# We will first download images and labels of a subset of the [Google Open Images Dataset](https://storage.googleapis.com/openimages/web/index.html). These labels were [carefully verified](https://storage.googleapis.com/openimages/web/factsfigures.html). Later, we will compare Ground Truth annotations to these labels. Our dataset will consist of images of various species of bird.
#
# If you chose `RUN_FULL_AL_DEMO = False`, then we will choose a subset of 30 images from this dataset. This is a diverse dataset of interesting images, and it should be fun for the human annotators to work with. You are free to ask the annotators to annotate any images you wish as long as the images do not contain adult content. In this case, you must adjust the labeling job request this job produces; please check the Ground Truth documentation.
#
# We will copy these images to our local `BUCKET` and create a corresponding *input manifest*. The input manifest is a formatted list of the S3 locations of the images we want Ground Truth to annotate. We will upload this manifest to our S3 `BUCKET`.
#
# #### Disclosure regarding the Open Images Dataset V4:
# Open Images Dataset V4 is created by Google Inc. We have not modified the images or the accompanying annotations. You can obtain the images and the annotations [here](https://storage.googleapis.com/openimages/web/download.html). The annotations are licensed by Google Inc. under [CC BY 4.0](https://creativecommons.org/licenses/by/2.0/) license. The images are listed as having a [CC BY 2.0](https://creativecommons.org/licenses/by/2.0/) license. The following paper describes Open Images V4 in depth: from the data collection and annotation to detailed statistics about the data and evaluation of models trained on it.
#
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>.
# *The Open Images Dataset V4: Unified image classification, object detection, and visual relationship detection at scale.* arXiv:1811.00982, 2018. ([link to PDF](https://arxiv.org/abs/1811.00982))
# +
# Download and process the Open Images annotations.
# !wget https://storage.googleapis.com/openimages/2018_04/test/test-annotations-bbox.csv
# !wget https://storage.googleapis.com/openimages/2018_04/bbox_labels_600_hierarchy.json
with open("bbox_labels_600_hierarchy.json", "r") as f:
hierarchy = json.load(f)
CLASS_NAME = "Bird"
CLASS_ID = "/m/015p6"
# Find all the subclasses of the desired image class (e.g. 'swans' and 'pigeons' etc if CLASS_NAME=='Bird').
good_subclasses = set()
def get_all_subclasses(hierarchy, good_subtree=False):
if hierarchy["LabelName"] == CLASS_ID:
good_subtree = True
if good_subtree:
good_subclasses.add(hierarchy["LabelName"])
if "Subcategory" in hierarchy:
for subcat in hierarchy["Subcategory"]:
get_all_subclasses(subcat, good_subtree=good_subtree)
return good_subclasses
good_subclasses = get_all_subclasses(hierarchy)
# Find an appropriate number of images with at least one bounding box in the desired category
if RUN_FULL_AL_DEMO:
n_ims = 1000
else:
n_ims = 30
fids2bbs = defaultdict(list)
# Skip images with risky content.
skip_these_images = ["251d4c429f6f9c39", "065ad49f98157c8d"]
with open("test-annotations-bbox.csv", "r") as f:
for line in f.readlines()[1:]:
line = line.strip().split(",")
img_id, _, cls_id, conf, xmin, xmax, ymin, ymax, *_ = line
if img_id in skip_these_images:
continue
if cls_id in good_subclasses:
fids2bbs[img_id].append([CLASS_NAME, xmin, xmax, ymin, ymax])
if len(fids2bbs) == n_ims:
break
# Copy the images to our local bucket.
s3 = boto3.client("s3")
for img_id_id, img_id in enumerate(fids2bbs.keys()):
if img_id_id % 100 == 0:
print("Copying image {} / {}".format(img_id_id, n_ims))
copy_source = {"Bucket": "open-images-dataset", "Key": "test/{}.jpg".format(img_id)}
s3.copy(copy_source, BUCKET, "{}/images/{}.jpg".format(EXP_NAME, img_id))
print("Done!")
# Create and upload the input manifest.
manifest_name = "input.manifest"
with open(manifest_name, "w") as f:
for img_id_id, img_id in enumerate(fids2bbs.keys()):
img_path = "s3://{}/{}/images/{}.jpg".format(BUCKET, EXP_NAME, img_id)
f.write('{"source-ref": "' + img_path + '"}\n')
s3.upload_file(manifest_name, BUCKET, EXP_NAME + "/" + manifest_name)
# -
# After running the cell above, you should be able to go to `s3://BUCKET/EXP_NAME/images` in the [S3 console](https://console.aws.amazon.com/s3/) and see 1000 images (or 100 if you have set `RUN_FULL_AL_DEMO = False`). We recommend you inspect these images! You can download them to a local machine using the AWS CLI.
# ## Specify the categories
#
# To run an object detection labeling job, you must decide on a set of classes the annotators can choose from. At the moment, Ground Truth only supports annotating one OD class at a time. In our case, the singleton class list is simply `["Bird"]`. To work with Ground Truth, this list needs to be converted to a .json file and uploaded to the S3 `BUCKET`.
# +
CLASS_LIST = [CLASS_NAME]
print("Label space is {}".format(CLASS_LIST))
json_body = {"labels": [{"label": label} for label in CLASS_LIST]}
with open("class_labels.json", "w") as f:
json.dump(json_body, f)
s3.upload_file("class_labels.json", BUCKET, EXP_NAME + "/class_labels.json")
# -
# You should now see `class_labels.json` in `s3://BUCKET/EXP_NAME/`.
# ## Create the instruction template
#
# Part or all of your images will be annotated by human annotators. It is **essential** to provide good instructions. Good instructions are:
# 1. Concise. We recommend limiting verbal/textual instruction to two sentences and focusing on clear visuals.
# 2. Visual. In the case of object detection, we recommend providing several labeled examples with different numbers of boxes.
#
# When used through the AWS Console, Ground Truth helps you create the instructions using a visual wizard. When using the API, you need to create an HTML template for your instructions. Below, we prepare a very simple but effective template and upload it to your S3 bucket.
#
# NOTE: If you use any images in your template (as we do), they need to be publicly accessible. You can enable public access to files in your S3 bucket through the S3 Console, as described in [S3 Documentation](https://docs.aws.amazon.com/AmazonS3/latest/user-guide/set-object-permissions.html).
#
# #### Testing your instructions
# **It is very easy to create broken instructions.** This might cause your labeling job to fail. However, it might also cause your job to complete with meaningless results if, for example, the annotators have no idea what to do or the instructions are misleading. At the moment the only way to test the instructions is to run your job in a private workforce. This is a way to run a mock labeling job for free. We describe how in [Verify your task using a private team [OPTIONAL]](#Verify-your-task-using-a-private-team-[OPTIONAL]).
#
# It is helpful to show examples of correctly labeled images in the instructions. The following code block produces several such examples for our dataset and saves them in `s3://BUCKET/EXP_NAME/`.
# +
# Plot sample images.
def plot_bbs(ax, bbs, img):
'''Add bounding boxes to images.'''
ax.imshow(img)
imh, imw, _ = img.shape
for bb in bbs:
xmin, xmax, ymin, ymax = bb
xmin *= imw
xmax *= imw
ymin *= imh
ymax *= imh
rec = plt.Rectangle((xmin, ymin), xmax-xmin, ymax-ymin, fill=None, lw=4, edgecolor='blue')
ax.add_patch(rec)
plt.figure(facecolor='white', dpi=100, figsize=(3, 7))
plt.suptitle('Please draw a box\n around each {}\n like the examples below.\n Thank you!'.format(CLASS_NAME), fontsize=15)
for fid_id, (fid, bbs) in enumerate([list(fids2bbs.items())[idx] for idx in [1, 3]]):
# !aws s3 cp s3://open-images-dataset/test/{fid}.jpg .
img = imageio.imread(fid + '.jpg')
bbs = [[float(a) for a in annot[1:]] for annot in bbs]
ax = plt.subplot(2, 1, fid_id+1)
plot_bbs(ax, bbs, img)
plt.axis('off')
plt.savefig('instructions.png', dpi=60)
with open('instructions.png', 'rb') as instructions:
instructions_uri = base64.b64encode(instructions.read()).decode('utf-8').replace('\n', '')
# +
from IPython.core.display import HTML, display
def make_template(test_template=False, save_fname="instructions.template"):
template = r"""<script src="https://assets.crowd.aws/crowd-html-elements.js"></script>
<crowd-form>
<crowd-bounding-box
name="boundingBox"
src="{{{{ task.input.taskObject | grant_read_access }}}}"
header="Dear Annotator, please draw a tight box around each {class_name} you see (if there are more than 8 birds, draw boxes around at least 8). Thank you!"
labels="{labels_str}"
>
<full-instructions header="Please annotate each {class_name}.">
<ol>
<li><strong>Inspect</strong> the image</li>
<li><strong>Determine</strong> if the specified label is/are visible in the picture.</li>
<li><strong>Outline</strong> each instance of the specified label in the image using the provided “Box” tool.</li>
</ol>
<ul>
<li>Boxes should fit tight around each object</li>
<li>Do not include parts of the object are overlapping or that cannot be seen, even though you think you can interpolate the whole shape.</li>
<li>Avoid including shadows.</li>
<li>If the target is off screen, draw the box up to the edge of the image.</li>
</ul>
</full-instructions>
<short-instructions>
<img src="data:image/png;base64,{instructions_uri}" style="max-width:100%">
</short-instructions>
</crowd-bounding-box>
</crowd-form>
""".format(
class_name=CLASS_NAME,
instructions_uri=instructions_uri,
labels_str=str(CLASS_LIST)
if test_template
else "{{ task.input.labels | to_json | escape }}",
)
with open(save_fname, "w") as f:
f.write(template)
make_template(test_template=True, save_fname="instructions.html")
make_template(test_template=False, save_fname="instructions.template")
s3.upload_file("instructions.template", BUCKET, EXP_NAME + "/instructions.template")
# -
# You should now be able to find your template in `s3://BUCKET/EXP_NAME/instructions.template`.
# ## Define pre-built lambda functions for use in the labeling job
#
# Before we submit the request, we need to define the ARNs for four key components of the labeling job: 1) the workteam, 2) the annotation consolidation Lambda function, 3) the pre-labeling task Lambda function, and 4) the machine learning algorithm to perform auto-annotation. These functions are defined by strings with region names and AWS service account numbers, so we will define a mapping below that will enable you to run this notebook in any of our supported regions.
#
# See the official documentation for the available ARNs:
# * [Documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/API_HumanTaskConfig.html#SageMaker-Type-HumanTaskConfig-PreHumanTaskLambdaArn) for available pre-human ARNs for other workflows.
# * [Documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/API_AnnotationConsolidationConfig.html#SageMaker-Type-AnnotationConsolidationConfig-AnnotationConsolidationLambdaArn) for available annotation consolidation ANRs for other workflows.
# * [Documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/API_LabelingJobAlgorithmsConfig.html#SageMaker-Type-LabelingJobAlgorithmsConfig-LabelingJobAlgorithmSpecificationArn) for available auto-labeling ARNs for other workflows.
# +
# Specify ARNs for resources needed to run an object detection job.
ac_arn_map = {
"us-west-2": "081040173940",
"us-east-1": "432418664414",
"us-east-2": "266458841044",
"eu-west-1": "568282634449",
"eu-central-1": '203001061592',
"ap-northeast-1": "477331159723",
}
prehuman_arn = "arn:aws:lambda:{}:{}:function:PRE-BoundingBox".format(region, ac_arn_map[region])
acs_arn = "arn:aws:lambda:{}:{}:function:ACS-BoundingBox".format(region, ac_arn_map[region])
labeling_algorithm_specification_arn = "arn:aws:sagemaker:{}:027400017018:labeling-job-algorithm-specification/object-detection".format(
region
)
workteam_arn = "arn:aws:sagemaker:{}:394669845002:workteam/public-crowd/default".format(region)
# -
# ## Submit the Ground Truth job request (Only one workshop participant should run this)
#
# ### Make sure this section is only run by a single workshop participant
# The API starts a Ground Truth job by submitting a request. The request contains the
# full configuration of the annotation task, and allows you to modify the fine details of
# the job that are fixed to default values when you use the AWS Console. The parameters that make up the request are described in more detail in the [SageMaker Ground Truth documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/API_CreateLabelingJob.html).
#
# After you submit the request, you should be able to see the job in your AWS Console, at `Amazon SageMaker > Labeling Jobs`.
# You can track the progress of the job there. This job will take several hours to complete. If your job
# is larger (say 100,000 images), the speed and cost benefit of auto-labeling should be larger.
#
# ### Verify your task using a private team
# If you chose to follow the steps in [Create a private team](#Create-a-private-team-to-test-your-task-[OPTIONAL]), you can first verify that your task runs as expected. To do this:
# 1. Set VERIFY_USING_PRIVATE_WORKFORCE to True in the cell below.
# 2. Run the next two cells. This will define the task and submit it to the private workforce (you).
# 3. After a few minutes, you should be able to see your task in your private workforce interface [Create a private team](#Create-a-private-team-to-test-your-task-[OPTIONAL]).
# Please verify that the task appears as you want it to appear.
# 4. If everything is in order, change `VERIFY_USING_PRIVATE_WORKFORCE` to `False` and rerun the cell below to start the real annotation task!
# +
VERIFY_USING_PRIVATE_WORKFORCE = True
USE_AUTO_LABELING = True
task_description = "Dear Annotator, please draw a box around each {}. Thank you!".format(CLASS_NAME)
task_keywords = ["image", "object", "detection"]
task_title = "Please draw a box around each {}.".format(CLASS_NAME)
job_name = "ground-truth-od-demo-" + str(int(time.time()))
human_task_config = {
"AnnotationConsolidationConfig": {
"AnnotationConsolidationLambdaArn": acs_arn,
},
"PreHumanTaskLambdaArn": prehuman_arn,
"MaxConcurrentTaskCount": 200, # 200 images will be sent at a time to the workteam.
"NumberOfHumanWorkersPerDataObject": 5, # We will obtain and consolidate 5 human annotations for each image.
"TaskAvailabilityLifetimeInSeconds": 21600, # Your workteam has 6 hours to complete all pending tasks.
"TaskDescription": task_description,
"TaskKeywords": task_keywords,
"TaskTimeLimitInSeconds": 300, # Each image must be labeled within 5 minutes.
"TaskTitle": task_title,
"UiConfig": {
"UiTemplateS3Uri": "s3://{}/{}/instructions.template".format(BUCKET, EXP_NAME),
},
}
if not VERIFY_USING_PRIVATE_WORKFORCE:
human_task_config["PublicWorkforceTaskPrice"] = {
"AmountInUsd": {
"Dollars": 0,
"Cents": 3,
"TenthFractionsOfACent": 6,
}
}
human_task_config["WorkteamArn"] = workteam_arn
else:
human_task_config["WorkteamArn"] = private_workteam_arn
ground_truth_request = {
"InputConfig": {
"DataSource": {
"S3DataSource": {
"ManifestS3Uri": "s3://{}/{}/{}".format(BUCKET, EXP_NAME, manifest_name),
}
},
"DataAttributes": {
"ContentClassifiers": ["FreeOfPersonallyIdentifiableInformation", "FreeOfAdultContent"]
},
},
"OutputConfig": {
"S3OutputPath": "s3://{}/{}/output/".format(BUCKET, EXP_NAME),
},
"HumanTaskConfig": human_task_config,
"LabelingJobName": job_name,
"RoleArn": role,
"LabelAttributeName": "category",
"LabelCategoryConfigS3Uri": "s3://{}/{}/class_labels.json".format(BUCKET, EXP_NAME),
}
if USE_AUTO_LABELING and RUN_FULL_AL_DEMO:
ground_truth_request["LabelingJobAlgorithmsConfig"] = {
"LabelingJobAlgorithmSpecificationArn": labeling_algorithm_specification_arn
}
sagemaker_client = boto3.client("sagemaker")
sagemaker_client.create_labeling_job(**ground_truth_request)
# -
# ## Monitor job progress
# A Ground Truth job can take a few hours to complete (if your dataset is larger than 10000 images, it can take much longer than that!). One way to monitor the job's progress is through AWS Console. In this notebook, we will use Ground Truth output files and Cloud Watch logs in order to monitor the progress.
#
# You can re-evaluate the next cell repeatedly. It sends a `describe_labeling_job` request which should tell you whether the job is completed or not. If it is, then 'LabelingJobStatus' will be 'Completed'.
sagemaker_client = boto3.client("sagemaker")
sagemaker_client.describe_labeling_job(LabelingJobName=job_name)["LabelingJobStatus"]
# The next cell extracts detailed information on how your job is doing. You can re-evaluate it at any time. It should give you:
# * The number of human and machine-annotated images across the iterations of your labeling job.
# * The training curves of any neural network training jobs launched by Ground Truth **(only if you are running with `RUN_FULL_AL_DEMO=True`)**.
# * The cost of the human- and machine-annotated labels.
#
# To understand the pricing, study [this document](https://aws.amazon.com/sagemaker/groundtruth/pricing/) carefully. In our case, each human label costs `$0.08 + 5 * $0.036 = $0.26` and each auto-label costs `$0.08`. If you set `RUN_FULL_AL_DEMO=True`, there is also the added cost of using SageMaker instances for neural net training and inference during auto-labeling. However, this should be insignificant compared to the other costs.
#
# If `RUN_FULL_AL_DEMO==True`, then the job will proceed in multiple iterations.
# * Iteration 1: Ground Truth will send out 10 images as 'probes' for human annotation. If these are successfully annotated, proceed to Iteration 2.
# * Iteration 2: Send out a batch of `MaxConcurrentTaskCount - 10` (in our case, 190) images for human annotation to obtain an active learning training batch.
# * Iteration 3: Send out another batch of 200 images for human annotation to obtain an active learning validation set.
# * Iteration 4a: Train a neural net to do auto-labeling. Auto-label as many data points as possible.
# * Iteration 4b: If there is any data leftover, send out at most 200 images for human annotation.
# * Repeat Iteration 4a and 4b until all data is annotated.
#
# If `RUN_FULL_AL_DEMO==False`, only Iterations 1 and 2 will happen.
# +
HUMAN_PRICE = 0.26
AUTO_PRICE = 0.08
try:
os.makedirs('od_output_data/', exist_ok=False)
except FileExistsError:
shutil.rmtree('od_output_data/')
S3_OUTPUT = boto3.client('sagemaker').describe_labeling_job(LabelingJobName=job_name)[
'OutputConfig']['S3OutputPath'] + job_name
# Count number of human annotations in each class each iteration.
# !aws s3 cp {S3_OUTPUT + '/annotations/consolidated-annotation/consolidation-response'} od_output_data/consolidation-response --recursive --quiet
consolidated_nboxes = defaultdict(int)
consolidated_nims = defaultdict(int)
consolidation_times = {}
consolidated_cost_times = []
obj_ids = set()
for consolidated_fname in glob.glob('od_output_data/consolidation-response/**', recursive=True):
if consolidated_fname.endswith('json'):
iter_id = int(consolidated_fname.split('/')[-2][-1])
# Store the time of the most recent consolidation event as iteration time.
iter_time = datetime.strptime(consolidated_fname.split('/')[-1], '%Y-%m-%d_%H:%M:%S.json')
if iter_id in consolidation_times:
consolidation_times[iter_id] = max(consolidation_times[iter_id], iter_time)
else:
consolidation_times[iter_id] = iter_time
consolidated_cost_times.append(iter_time)
with open(consolidated_fname, 'r') as f:
consolidated_data = json.load(f)
for consolidation in consolidated_data:
obj_id = consolidation['datasetObjectId']
n_boxes = len(consolidation['consolidatedAnnotation']['content'][
'category']['annotations'])
if obj_id not in obj_ids:
obj_ids.add(obj_id)
consolidated_nims[iter_id] += 1
consolidated_nboxes[iter_id] += n_boxes
total_human_labels = sum(consolidated_nims.values())
# Count the number of machine iterations in each class each iteration.
# !aws s3 cp {S3_OUTPUT + '/activelearning'} od_output_data/activelearning --recursive --quiet
auto_nboxes = defaultdict(int)
auto_nims = defaultdict(int)
auto_times = {}
auto_cost_times = []
for auto_fname in glob.glob('od_output_data/activelearning/**', recursive=True):
if auto_fname.endswith('auto_annotator_output.txt'):
iter_id = int(auto_fname.split('/')[-3])
with open(auto_fname, 'r') as f:
annots = [' '.join(l.split()[1:]) for l in f.readlines()]
auto_nims[iter_id] += len(annots)
for annot in annots:
annot = json.loads(annot)
time_str = annot['category-metadata']['creation-date']
auto_time = datetime.strptime(time_str, '%Y-%m-%dT%H:%M:%S.%f')
n_boxes = len(annot['category']['annotations'])
auto_nboxes[iter_id] += n_boxes
if iter_id in auto_times:
auto_times[iter_id] = max(auto_times[iter_id], auto_time)
else:
auto_times[iter_id] = auto_time
auto_cost_times.append(auto_time)
total_auto_labels = sum(auto_nims.values())
n_iters = max(len(auto_times), len(consolidation_times))
# Get plots for auto-annotation neural-net training.
def get_training_job_data(training_job_name):
logclient = boto3.client('logs')
log_group_name = '/aws/sagemaker/TrainingJobs'
log_stream_name = logclient.describe_log_streams(logGroupName=log_group_name,
logStreamNamePrefix=training_job_name)['logStreams'][0]['logStreamName']
train_log = logclient.get_log_events(
logGroupName=log_group_name,
logStreamName=log_stream_name,
startFromHead=True
)
events = train_log['events']
next_token = train_log['nextForwardToken']
while True:
train_log = logclient.get_log_events(
logGroupName=log_group_name,
logStreamName=log_stream_name,
startFromHead=True,
nextToken=next_token
)
if train_log['nextForwardToken'] == next_token:
break
events = events + train_log['events']
mAPs = []
for event in events:
msg = event['message']
if 'Final configuration' in msg:
num_samples = int(msg.split('num_training_samples\': u\'')[1].split('\'')[0])
elif 'validation mAP <score>=(' in msg:
mAPs.append(float(msg.split('validation mAP <score>=(')[1][:-1]))
return num_samples, mAPs
# training_data = !aws s3 ls {S3_OUTPUT + '/training/'} --recursive
training_sizes = []
training_mAPs = []
training_iters = []
for line in training_data:
if line.split('/')[-1] == 'model.tar.gz':
training_job_name = line.split('/')[-3]
n_samples, mAPs = get_training_job_data(training_job_name)
training_sizes.append(n_samples)
training_mAPs.append(mAPs)
training_iters.append(int(line.split('/')[-5]))
plt.figure(facecolor='white', figsize=(14, 5), dpi=100)
ax = plt.subplot(131)
total_human = 0
total_auto = 0
for iter_id in range(1, n_iters + 1):
cost_human = consolidated_nims[iter_id] * HUMAN_PRICE
cost_auto = auto_nims[iter_id] * AUTO_PRICE
total_human += cost_human
total_auto += cost_auto
plt.bar(iter_id, cost_human, width=.8, color='C0',
label='human' if iter_id==1 else None)
plt.bar(iter_id, cost_auto, bottom=cost_human,
width=.8, color='C1', label='auto' if iter_id==1 else None)
plt.title('Total annotation costs:\n\${:.2f} human, \${:.2f} auto'.format(
total_human, total_auto))
plt.xlabel('Iter')
plt.ylabel('Cost in dollars')
plt.legend()
plt.subplot(132)
plt.title('Total annotation counts:\nHuman: {} ims, {} boxes\nMachine: {} ims, {} boxes'.format(
sum(consolidated_nims.values()), sum(consolidated_nboxes.values()), sum(auto_nims.values()), sum(auto_nboxes.values())))
for iter_id in consolidated_nims.keys():
plt.bar(iter_id, auto_nims[iter_id], color='C1', width=.4, label='ims, auto' if iter_id==1 else None)
plt.bar(iter_id, consolidated_nims[iter_id],
bottom=auto_nims[iter_id], color='C0', width=.4, label='ims, human' if iter_id==1 else None)
plt.bar(iter_id + .4, auto_nboxes[iter_id], color='C1', alpha=.4, width=.4, label='boxes, auto' if iter_id==1 else None)
plt.bar(iter_id + .4, consolidated_nboxes[iter_id],
bottom=auto_nboxes[iter_id], color='C0', width=.4, alpha=.4, label='boxes, human' if iter_id==1 else None)
tick_labels_boxes = ['Iter {}, boxes'.format(iter_id + 1) for iter_id in range(n_iters)]
tick_labels_images = ['Iter {}, images'.format(iter_id + 1) for iter_id in range(n_iters)]
tick_locations_images = np.arange(n_iters) + 1
tick_locations_boxes = tick_locations_images + .4
tick_labels = np.concatenate([[tick_labels_boxes[idx], tick_labels_images[idx]] for idx in range(n_iters)])
tick_locations = np.concatenate([[tick_locations_boxes[idx], tick_locations_images[idx]] for idx in range(n_iters)])
plt.xticks(tick_locations, tick_labels, rotation=90)
plt.legend()
plt.ylabel('Count')
if len(training_sizes) > 0:
plt.subplot(133)
plt.title('Active learning training curves')
plt.grid(True)
cmap = plt.get_cmap('coolwarm')
n_all = len(training_sizes)
for iter_id_id, (iter_id, size, mAPs) in enumerate(zip(training_iters, training_sizes, training_mAPs)):
plt.plot(mAPs, label='Iter {}, auto'.format(iter_id + 1), color=cmap(iter_id_id / max(1, (n_all-1))))
plt.legend()
plt.xlabel('Training epoch')
plt.ylabel('Validation mAP')
plt.tight_layout()
# -
# # Analyze Ground Truth labeling job results (can be run by all participants)
# **This section should take about 20 minutes to complete.**
#
# Once the job has finished, we can analyze the results. Evaluate the following cell and verify the output is `'Completed'` before continuing.
job_name = "ground-truth-od-demo-1626694327"
sagemaker_client.describe_labeling_job(LabelingJobName=job_name)["LabelingJobStatus"]
# The plots in the [Monitor job progress](#Monitor-job-progress) section form part of the analysis. In this section, we will gain additional insights into the results, which are contained in the output manifest. You can find the location of the output manifest under `AWS Console > SageMaker > Labeling Jobs > [name of your job]`. We will obtain it programmatically in the cell below.
#
# ## Postprocess the output manifest
# Now that the job is complete, we will download the output manifest manfiest and postprocess it to create a list of `output_images` with the results. Each entry in the list will be a `BoxedImage` object that contains information about the image and the bounding boxes created by the labeling jobs.
# +
# Load the output manifest's annotations.
OUTPUT_MANIFEST = "s3://{}/{}/output/{}/manifests/output/output.manifest".format(
BUCKET, EXP_NAME, job_name
)
# !aws s3 cp {OUTPUT_MANIFEST} 'output.manifest'
with open("output.manifest", "r") as f:
output = [json.loads(line.strip()) for line in f.readlines()]
# Retrieve the worker annotations.
# !aws s3 cp {S3_OUTPUT + '/annotations/worker-response'} od_output_data/worker-response --recursive --quiet
# Find the worker files.
worker_file_names = glob.glob("od_output_data/worker-response/**/*.json", recursive=True)
# +
from ground_truth_od import BoundingBox, WorkerBoundingBox, GroundTruthBox, BoxedImage
# Create data arrays.
confidences = np.zeros(len(output))
# Find the job name the manifest corresponds to.
keys = list(output[0].keys())
metakey = keys[np.where([("-metadata" in k) for k in keys])[0][0]]
jobname = metakey[:-9]
output_images = []
consolidated_boxes = []
# Extract the data.
for datum_id, datum in enumerate(output):
image_size = datum["category"]["image_size"][0]
box_annotations = datum["category"]["annotations"]
uri = datum["source-ref"]
box_confidences = datum[metakey]["objects"]
human = int(datum[metakey]["human-annotated"] == "yes")
# Make image object.
image = BoxedImage(id=datum_id, size=image_size, uri=uri)
# Create bounding boxes for image.
boxes = []
for i, annotation in enumerate(box_annotations):
box = BoundingBox(image_id=datum_id, boxdata=annotation)
box.confidence = box_confidences[i]["confidence"]
box.image = image
box.human = human
boxes.append(box)
consolidated_boxes.append(box)
image.consolidated_boxes = boxes
# Store if the image is human labeled.
image.human = human
# Retrieve ground truth boxes for the image.
oid_boxes_data = fids2bbs[image.oid_id]
gt_boxes = []
for data in oid_boxes_data:
gt_box = GroundTruthBox(image_id=datum_id, oiddata=data, image=image)
gt_boxes.append(gt_box)
image.gt_boxes = gt_boxes
output_images.append(image)
# Iterate through the json files, creating bounding box objects.
for wfn in worker_file_names:
image_id = int(wfn.split("/")[-2])
image = output_images[image_id]
with open(wfn, "r") as worker_file:
annotation = json.load(worker_file)
answers = annotation["answers"]
for answer in answers:
wid = answer["workerId"]
wboxes_data = answer["answerContent"]["boundingBox"]["boundingBoxes"]
for boxdata in wboxes_data or []:
box = WorkerBoundingBox(image_id=image_id, worker_id=wid, boxdata=boxdata)
box.image = image
image.worker_boxes.append(box)
# Get the human- and auto-labeled images.
human_labeled = [img for img in output_images if img.human]
auto_labeled = [img for img in output_images if not img.human]
# -
# ## Plot annotated images
# In any data science task, it is crucial to plot and inspect the results to check they make sense. In order to do this, we will
# 1. Download the input images that Ground Truth annotated.
# 2. Separate images annotated by humans from those annoted via the auto-labeling mechanism.
# 3. Plot images in the human/auto-annotated classes.
#
# We will download the input images to a `LOCAL_IMAGE_DIR` you can choose in the next cell. Note that if this directory already contains images with the same filenames as your Ground Truth input images, we will not re-download the images.
#
# If your dataset is large and you do not wish to download and plot **all** the images, simply set `DATASET_SIZE` to a small number. We will pick a random subset of your data for plotting.
# +
LOCAL_IMG_DIR = '/tmp' # Replace with the name of a local directory to store images.
assert LOCAL_IMG_DIR != '<< choose a local directory name to download the images to >>', 'Please provide a local directory name'
DATASET_SIZE = len(output_images) # Change this to a reasonable number if your dataset is larger than 10K images.
image_subset = np.random.choice(output_images, DATASET_SIZE, replace=False)
for img in image_subset:
target_fname = os.path.join(
LOCAL_IMG_DIR, img.uri.split('/')[-1])
if not os.path.isfile(target_fname):
# !aws s3 cp {img.uri} {target_fname}
# -
# ### Plot a small output sample to understand the labeling mechanism
# The following cell will create two figures. The first plots `N_SHOW` images as annotated by humans. The first column shows the original bounding boxes produced by the human labelers working on Amazon Mechanical Turk. The second column shows the result of combining these boxes to produce a consolidated label, which is the final output of Ground Truth for the human-labeled images. Finally, the third column shows the "true" bounding boxes according to the Open Images Dataset for reference.
#
# The second plots `N_SHOW` images as annotated by the auto-labeling mechanism. In this case, there is no consolidation phase, so only the auto-labeled image and the "true" label are displayed.
#
# By default, `N_SHOW = 5`, but feel free to change this to any small number.
# +
N_SHOW = 5
# Find human and auto-labeled images in the subset.
human_labeled_subset = [img for img in image_subset if img.human]
auto_labeled_subset = [img for img in image_subset if not img.human]
# Show examples of each
fig, axes = plt.subplots(N_SHOW, 3, figsize=(9, 2 * N_SHOW), facecolor="white", dpi=100)
fig.suptitle("Human-labeled examples", fontsize=24)
axes[0, 0].set_title("Worker labels", fontsize=14)
axes[0, 1].set_title("Consolidated label", fontsize=14)
axes[0, 2].set_title("True label", fontsize=14)
for row, img in enumerate(np.random.choice(human_labeled_subset, size=N_SHOW)):
img.download(LOCAL_IMG_DIR)
img.plot_worker_bbs(axes[row, 0])
img.plot_consolidated_bbs(axes[row, 1])
img.plot_gt_bbs(axes[row, 2])
if auto_labeled_subset:
fig, axes = plt.subplots(N_SHOW, 2, figsize=(6, 2 * N_SHOW), facecolor="white", dpi=100)
fig.suptitle("Auto-labeled examples", fontsize=24)
axes[0, 0].set_title("Auto-label", fontsize=14)
axes[0, 1].set_title("True label", fontsize=14)
for row, img in enumerate(np.random.choice(auto_labeled_subset, size=N_SHOW)):
img.download(LOCAL_IMG_DIR)
img.plot_consolidated_bbs(axes[row, 0])
img.plot_gt_bbs(axes[row, 1])
else:
print("No images were auto-labeled.")
# -
# ### Plot the resulting bounding boxes to a pdf
# Finally, we plot the results to two large pdf files. You can adjust the number of `rows_per_page` and `columns_per_page` if you would like. With the default settings, the pdfs will display 25 images per page. Each page will contain images annotated either by human annotators or by the auto-labeling mechanism. The first, `ground-truth-od-confidence.pdf`, contains images sorted by the confidence Ground Truth has in its prediction. The second, `ground-truth-od-miou.pdf`, contains the same images, but sorted by the quality of the annotations compared to the standard labels from the Open Images Dataset. See the [Compare Ground Truth results to standard labels](#Compare-Ground-Truth-results-to-standard-labels) section for more details.
#
# We will only plot 10 each of the human- and auto-annotated images. You can set `N_SHOW` to another number if you want to only plot more of the images.
# +
"""Create pdfs with images sorted by miou and confidence."""
N_SHOW = 10
# Created, sort list of imgs and mious.
h_img_mious = [(img, img.compute_iou_bb()) for img in human_labeled]
a_img_mious = [(img, img.compute_iou_bb()) for img in auto_labeled]
h_img_mious.sort(key=lambda x: x[1], reverse=True)
a_img_mious.sort(key=lambda x: x[1], reverse=True)
# Create, sort the images by confidence.
h_img_confs = [(img, img.compute_img_confidence()) for img in human_labeled]
a_img_confs = [(img, img.compute_img_confidence()) for img in auto_labeled]
h_img_confs.sort(key=lambda x: x[1], reverse=True)
a_img_confs.sort(key=lambda x: x[1], reverse=True)
# Define number of rows, columns per page.
rows_per_page = 5
columns_per_page = 5
n_per_page = rows_per_page * columns_per_page
def title_page(title):
"""Create a page with only text."""
plt.figure(figsize=(10, 10), facecolor="white", dpi=100)
plt.text(0.1, 0.5, s=title, fontsize=20)
plt.axis("off")
pdf.savefig()
plt.close()
def page_loop(mious, axes, worker=False):
"""Loop over a single image page of the output pdf."""
for i, row in enumerate(axes):
for j, ax in enumerate(row):
img_idx = n_per_page * page + rows_per_page * i + j
# Break out of loop if all the images are plotted.
if img_idx >= min(N_SHOW, len(mious)):
return
img, miou = mious[img_idx]
img.download(LOCAL_IMG_DIR)
if worker:
img.plot_worker_bbs(ax, img_kwargs={"aspect": "auto"}, box_kwargs={"lw": 0.5})
else:
img.plot_gt_bbs(
ax, img_kwargs={"aspect": "auto"}, box_kwargs={"edgecolor": "C2", "lw": 0.5}
)
img.plot_consolidated_bbs(
ax, img_kwargs={"aspect": "auto"}, box_kwargs={"edgecolor": "C1", "lw": 0.5}
)
# Create pdfs for the images sorted by confidence and by mIoU.
mode_metrics = (
("mIoU", (("Worker", h_img_mious), ("Consolidated human", h_img_mious), ("Auto", a_img_mious))),
(
"confidence",
(("Worker", h_img_confs), ("Consolidated human", h_img_confs), ("Auto", a_img_confs)),
),
)
for mode, labels_metrics in mode_metrics:
pdfname = f"ground-truth-od-{mode}.pdf"
with PdfPages(pdfname) as pdf:
title_page("Images labeled by SageMaker Ground Truth\n" f"and sorted by {mode}")
print(f"Plotting images sorted by {mode}...")
# Show human- and auto-labeled images.
for label, metrics in labels_metrics:
worker = label == "Worker"
if worker:
title_page("Original worker labels")
else:
title_page(f"{label} labels in orange,\n" "Open Image annotations in green")
n_images = min(len(metrics), N_SHOW)
n_pages = (n_images - 1) // n_per_page + 1
print(f"Plotting {label.lower()}-labeled images...")
for page in range(n_pages):
print(f"{page*n_per_page}/{n_images}")
fig, axes = plt.subplots(rows_per_page, columns_per_page, dpi=125)
page_loop(metrics, axes, worker=worker)
for ax in axes.ravel():
ax.axis("off")
# Find the max/min mIoU or confidence on each page.
metrics_page = metrics[page * n_per_page : min((page + 1) * n_per_page, n_images)]
max_metric = metrics_page[0][1]
min_metric = metrics_page[-1][1]
fig.suptitle(f"{mode} range: [{max_metric:1.3f}, {min_metric:1.3f}]")
pdf.savefig()
plt.close()
print("Done.")
# -
# # Compare Ground Truth results to standard labels
#
# **This section should take about 5 minutes to complete.**
#
# Sometimes we have an alternative set of data labels available.
# For example, the Open Images data has already been carefully annotated by a professional annotation workforce.
# This allows us to perform additional analysis that compares Ground Truth labels to the standard labels.
# When doing so, it is important to bear in mind that any image labels created by humans
# will most likely not be 100% accurate. For this reason, it is better to think of labeling accuracy as
# "adherence to a particular standard / set of labels" rather than "how good (in absolute terms) are the Ground Truth labels."
#
# ## Compute mIoUs for images in the dataset
# The following cell plots a histogram of the mean intersections-over-unions (mIoUs) between labels produced by Ground Truth and reference labels from the Open Images Dataset. The intersection over union, also known as the [Jaccard index](https://en.wikipedia.org/wiki/Jaccard_index), of two bounding boxes is a measure of their similarity. Because each image can contain multiple bounding boxes, we take the mean of the IoUs to measure the success of the labeling for that image.
# +
"""Plot the distribution of mIoUs by image in the dataset."""
h_mious = np.array([h_img_miou[1] for h_img_miou in h_img_mious])
a_mious = np.array([a_img_miou[1] for a_img_miou in a_img_mious])
xvals = np.linspace(0, 1, 17)
xticks = np.linspace(0, 1, 5)
plt.figure(figsize=(12, 5), dpi=300, facecolor="white")
plt.hist([h_mious, a_mious], rwidth=0.8, edgecolor="k", bins=xvals, label=["Human", "Auto"])
plt.xticks(xticks)
plt.title(
f"{len(h_mious)} human-labeled images with mIoU {np.mean(h_mious):.2f}\n{len(a_mious)} auto-labeled images with mIoU {np.mean(a_mious):.2f}"
)
plt.ylabel("Number of images")
plt.xlabel("mIoU")
plt.legend();
# -
# ## Visualize the results
# It is useful to see what corresponds to a good or bad mIoU in practice. The following cell displays images with the highest and lowest mIoUs vs the standard labels for both the human- and auto-labeled images. As before, the Ground Truth bounding boxes are in blue and the standard boxes are in lime green.
#
# In our example run, the images with the lowest mIoUs demonstrated that Ground Truth can sometimes outperform standard labels. In particular, many of the standard labels for this dataset contain only one large bounding box despite the presence of many small objects in the image.
# +
# Sort the images by mIoU.
h_img_mious.sort(key=lambda x: x[1], reverse=True)
a_img_mious.sort(key=lambda x: x[1], reverse=True)
# Plot images and mIoUs for human- vs auto-labeling.
if a_img_mious:
labels = ("Human", "Auto")
both_img_mious = (h_img_mious, a_img_mious)
else:
labels = ("Human",)
both_img_mious = (h_img_mious,)
for label, all_img_mious in zip(labels, both_img_mious):
# Do the highest and lowest mious
tb_img_mious = (all_img_mious[:6], all_img_mious[-6:])
titles = ("highest", "lowest")
for img_mious, title in zip(tb_img_mious, titles):
# Make a figure with six images.
fig, axes = plt.subplots(2, 3, figsize=(12, 4), dpi=100, facecolor="white")
for (img, miou), ax in zip(img_mious, axes.ravel()):
img.download(LOCAL_IMG_DIR)
img.plot_consolidated_bbs(ax, box_kwargs={"lw": 1.5, "color": "blue"})
img.plot_gt_bbs(ax, box_kwargs={"lw": 1, "color": "lime"})
ax.set_title(f"mIoU: {miou:1.3f}")
ax.axis("off")
fig.suptitle(f"{label}-labeled images with the {title} mIoUs", fontsize=16)
fig.tight_layout(rect=[0, 0, 1, 0.95])
# -
# ## Understand the relationship between confidence and annotation quality
#
# During both human- and auto-labeling, Ground Truth produces confidence scores associated with its labels. These scores are used internally by Ground Truth in various ways. As an example, the auto-labeling mechanism will only ouput an annotation for an image when the confidence passes a dynamically-generated threshold.
#
# In practice, Ground Truth is often used to annotate entirely new datasets for which there are no standard labels. The following cells show how the confidence acts as a proxy for the true quality of the annotations.
# +
"""Plot the mIoUs vs the confidences."""
from scipy import stats
import pandas as pd
import numpy as np
from ground_truth_od import group_miou
fig, (ax1, ax2) = plt.subplots(1, 2, dpi=100, facecolor="white", figsize=(12, 5))
if RUN_FULL_AL_DEMO:
label_confs_colors = (("Human", h_img_confs, "C0"), ("Auto", a_img_confs, "C1"))
else:
label_confs_colors = (("Human", h_img_confs, "C0"),)
ax1.set_title("mIoU vs confidence with regression lines")
ax1.set_xlabel("Confidence")
ax1.set_ylabel("mIoU")
for label, img_confs, color in label_confs_colors:
confs = [img_conf[1] for img_conf in img_confs]
mious = [img_conf[0].compute_iou_bb() for img_conf in img_confs]
# Compute regression line.
slope, intercept, *_ = stats.linregress(confs, mious)
xs = np.array((0, 1))
# Plot points and line.
ax1.plot(confs, mious, ".", label=label, color=color)
ax1.plot(xs, slope * xs + intercept, color=color, lw=3)
ax1.set_xlim([-0.05, 1.05])
ax1.set_ylim([-0.05, 1.05])
ax1.legend()
# Compute the mIoU of subsets of the images based on confidence level.
if RUN_FULL_AL_DEMO:
labels_imgs = (("Human", human_labeled), ("Auto", auto_labeled))
else:
labels_imgs = (("Human", human_labeled),)
deciles = np.linspace(0, 0.9, 10)
mious_deciles = {}
for label, imgs in labels_imgs:
# Find thresholds of confidences for deciles.
confs = np.array([img.compute_img_confidence() for img in imgs])
thresholds = pd.Series(confs).quantile(deciles)
# Select images with confidence greater than thresholds.
mious = []
for decile in deciles:
img_subset = [img for img in imgs if img.compute_img_confidence() > thresholds[decile]]
# Compute mious.
mious.append(group_miou(img_subset))
# Save the results.
mious_deciles[label] = mious
# Create pots
ax2.plot(100 - deciles * 100, mious, label=label)
ax2.set_ylabel("mIoU")
ax2.set_title("Effect of increasing confidence thresholds")
ax2.set_xlabel("Top x% of images by confidence")
ax2.set_xlim([105, 5])
ax2.set_xticks(np.linspace(100, 10, 10))
ax2.legend()
ax2.grid()
# -
# Once again, it is good to see some concrete examples. The next cell displays several of the human- and auto-labeled images with the highest confidence scores across the dataset.
# +
"""Plot the images with the highest confidences."""
# Sort the images by confidence.
h_img_confs = [(img, img.compute_img_confidence()) for img in human_labeled]
a_img_confs = [(img, img.compute_img_confidence()) for img in auto_labeled]
h_img_confs.sort(key=lambda x: x[1], reverse=True)
a_img_confs.sort(key=lambda x: x[1], reverse=True)
# Do both the human- and auto-labeled images.
label_confs = (("human", h_img_confs), ("auto", a_img_confs))
for label, img_confs in label_confs:
plt.figure(facecolor="white", figsize=(15, 4), dpi=100)
plt.suptitle(
f"Top-5 confidence {label}-labels (orange) and corresponding "
"Open Images annotations (green)"
)
for img_id, (img, conf) in enumerate(img_confs[:5]):
img.download(LOCAL_IMG_DIR)
ax = plt.subplot(1, 5, img_id + 1)
img.plot_gt_bbs(ax, box_kwargs={"edgecolor": "C2", "lw": 3})
img.plot_consolidated_bbs(ax, box_kwargs={"edgecolor": "C1", "lw": 3})
# -
print("mIoU for the whole dataset: ", group_miou(output_images))
print("mIoU for human-labeled images: ", group_miou(human_labeled))
print("mIoU for auto-labeled images: ", group_miou(auto_labeled))
# ### See how the number of objects in the image affects confidence
#
# The next cell produces two subplots:
# * The left subplot shows the counts of images with different numbers of objects in the image on a log scale. Notice that humans are assigned to label more of the images with many boxes.
#
# * The right subplot shows how the confidence associated with an image decreases as the number of objects in the image increases.
# +
# Compute the number of boxes per image and create a histogram.
nboxes_human = np.array([img.n_consolidated_boxes() for img in human_labeled])
nboxes_auto = np.array([img.n_consolidated_boxes() for img in auto_labeled])
max_boxes = max(
nboxes_auto.max() if nboxes_auto.size != 0 else 0,
nboxes_human.max() if nboxes_human.size != 0 else 0,
)
n_boxes = np.arange(0, max_boxes + 2)
# Find mean confidences by number of boxes.
h_confs_by_n = []
a_confs_by_n = []
# Do human and auto.
for labeled, mean_confs in ((human_labeled, h_confs_by_n), (auto_labeled, a_confs_by_n)):
for n_box in n_boxes:
h_img_n = [img for img in labeled if img.n_consolidated_boxes() == n_box]
mean_conf = np.mean([img.compute_img_confidence() for img in h_img_n])
mean_confs.append(mean_conf)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(14, 4), facecolor="white", dpi=100)
ax1.hist([nboxes_human, nboxes_auto], n_boxes, label=["Human", "Auto"], align="left")
ax1.set_xlabel("Bounding boxes in image")
ax1.set_title("Image counts vs number of bounding boxes")
ax1.set_yscale("log")
ax1.set_ylabel("Number of images")
ax1.legend()
# Find where we have nonzero box counts.
h_not_nan = np.logical_not(np.isnan(h_confs_by_n))
a_not_nan = np.logical_not(np.isnan(a_confs_by_n))
# Plot.
ax2.set_title("Image confidences vs number of bounding boxes")
ax2.plot(n_boxes[h_not_nan], np.array(h_confs_by_n)[h_not_nan], "D", color="C0", label="Human")
ax2.plot(n_boxes[a_not_nan], np.array(a_confs_by_n)[a_not_nan], "D", color="C1", label="Auto")
ax2.set_xlabel("Bounding boxes in image")
ax2.set_ylabel("Mean image confidence")
ax2.legend();
# -
# # Review
#
# Let's recap what we accomplished. First we started with an unlabeled dataset (technically, the dataset was previously labeled by the authors of the dataset, but we discarded the original labels for the purposes of this demonstration). Next, we created a SageMake Ground Truth labeling job and generated new labels for all of the images in our dataset. Then we analyzed the labeling job results.
#
# The next section is optional and shows how to train a machine learning model with the resulting labels.
# # OPTIONAL: Train an object detection model using Ground Truth labels
#
# At this stage, we have fully labeled our dataset and we can train a machine learning model to perform object detection. We'll do so using the **augmented manifest** output of our labeling job - no additional file translation or manipulation required! For a more complete description of the augmented manifest, see our other [example notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/ground_truth_labeling_jobs/object_detection_augmented_manifest_training/object_detection_augmented_manifest_training.ipynb).
#
# **NOTE:** Object detection is a complex task, and training neural networks to high accuracy requires large datasets and careful hyperparameter tuning. The following cells illustrate how to train a neural network using a Ground Truth output augmented manifest, and how to interpret the results. However, we shouldn't expect a network trained on 100 or 1000 images to do a phenomenal job on unseen images!
#
# First, we'll split our augmented manifest into a training set and a validation set using an 80/20 split and save the results to files that the model will use during training.
# +
with open("output.manifest", "r") as f:
output = [json.loads(line) for line in f.readlines()]
# Shuffle output in place.
np.random.shuffle(output)
dataset_size = len(output)
train_test_split_index = round(dataset_size * 0.8)
train_data = output[:train_test_split_index]
validation_data = output[train_test_split_index:]
num_training_samples = 0
with open("train.manifest", "w") as f:
for line in train_data:
f.write(json.dumps(line))
f.write("\n")
num_training_samples += 1
with open("validation.manifest", "w") as f:
for line in validation_data:
f.write(json.dumps(line))
f.write("\n")
# -
# Next, we'll upload these manifest files to the previously defined S3 bucket so that they can be used in the training job.
# !aws s3 cp train.manifest s3://{BUCKET}/{EXP_NAME}/train.manifest
# !aws s3 cp validation.manifest s3://{BUCKET}/{EXP_NAME}/validation.manifest
# ## Setup
#
# Here we define S3 file paths for input and output data, the training image containing the object detection algorithm, and instantiate a SageMaker session.
# +
import re
from sagemaker import get_execution_role
from time import gmtime, strftime
role = get_execution_role()
sess = sagemaker.Session()
s3 = boto3.resource("s3")
training_image = sagemaker.amazon.amazon_estimator.get_image_uri(
boto3.Session().region_name, "object-detection", repo_version="latest"
)
augmented_manifest_filename_train = "train.manifest"
augmented_manifest_filename_validation = "validation.manifest"
bucket_name = BUCKET
s3_prefix = EXP_NAME
s3_output_path = "s3://{}/groundtruth-od-augmented-manifest-output".format(
bucket_name
) # Replace with your desired output directory.
# +
# Defines paths for use in the training job request.
s3_train_data_path = "s3://{}/{}/{}".format(
bucket_name, s3_prefix, augmented_manifest_filename_train
)
s3_validation_data_path = "s3://{}/{}/{}".format(
bucket_name, s3_prefix, augmented_manifest_filename_validation
)
print("Augmented manifest for training data: {}".format(s3_train_data_path))
print("Augmented manifest for validation data: {}".format(s3_validation_data_path))
# +
augmented_manifest_s3_key = s3_train_data_path.split(bucket_name)[1][1:]
s3_obj = s3.Object(bucket_name, augmented_manifest_s3_key)
augmented_manifest = s3_obj.get()["Body"].read().decode("utf-8")
augmented_manifest_lines = augmented_manifest.split("\n")
num_training_samples = len(
augmented_manifest_lines
) # Compute number of training samples for use in training job request.
# Determine the keys in the training manifest and exclude the meta data from the labling job.
attribute_names = list(json.loads(augmented_manifest_lines[0]).keys())
attribute_names = [attrib for attrib in attribute_names if "meta" not in attrib]
# +
try:
if attribute_names == ["source-ref", "XXXX"]:
raise Exception(
"The 'attribute_names' variable is set to default values. Please check your augmented manifest file for the label attribute name and set the 'attribute_names' variable accordingly."
)
except NameError:
raise Exception(
"The attribute_names variable is not defined. Please check your augmented manifest file for the label attribute name and set the 'attribute_names' variable accordingly."
)
# Create unique job name
job_name_prefix = "ground-truthod-demo"
timestamp = time.strftime("-%Y-%m-%d-%H-%M-%S", time.gmtime())
model_job_name = job_name_prefix + timestamp
training_params = {
"AlgorithmSpecification": {
# NB. This is one of the named constants defined in the first cell.
"TrainingImage": training_image,
"TrainingInputMode": "Pipe",
},
"RoleArn": role,
"OutputDataConfig": {"S3OutputPath": s3_output_path},
"ResourceConfig": {"InstanceCount": 1, "InstanceType": "ml.p3.2xlarge", "VolumeSizeInGB": 50},
"TrainingJobName": model_job_name,
"HyperParameters": { # NB. These hyperparameters are at the user's discretion and are beyond the scope of this demo.
"base_network": "resnet-50",
"use_pretrained_model": "1",
"num_classes": "1",
"mini_batch_size": "1",
"epochs": "30",
"learning_rate": "0.001",
"lr_scheduler_step": "",
"lr_scheduler_factor": "0.1",
"optimizer": "sgd",
"momentum": "0.9",
"weight_decay": "0.0005",
"overlap_threshold": "0.5",
"nms_threshold": "0.45",
"image_shape": "300",
"label_width": "350",
"num_training_samples": str(num_training_samples),
},
"StoppingCondition": {"MaxRuntimeInSeconds": 86400},
"InputDataConfig": [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "AugmentedManifestFile", # NB. Augmented Manifest
"S3Uri": s3_train_data_path,
"S3DataDistributionType": "FullyReplicated",
# NB. This must correspond to the JSON field names in your augmented manifest.
"AttributeNames": attribute_names,
}
},
"ContentType": "application/x-recordio",
"RecordWrapperType": "RecordIO",
"CompressionType": "None",
},
{
"ChannelName": "validation",
"DataSource": {
"S3DataSource": {
"S3DataType": "AugmentedManifestFile", # NB. Augmented Manifest
"S3Uri": s3_validation_data_path,
"S3DataDistributionType": "FullyReplicated",
# NB. This must correspond to the JSON field names in your augmented manifest.
"AttributeNames": attribute_names,
}
},
"ContentType": "application/x-recordio",
"RecordWrapperType": "RecordIO",
"CompressionType": "None",
},
],
}
print("Training job name: {}".format(model_job_name))
print(
"\nInput Data Location: {}".format(
training_params["InputDataConfig"][0]["DataSource"]["S3DataSource"]
)
)
# -
# Now we create the SageMaker training job.
# +
client = boto3.client(service_name="sagemaker")
client.create_training_job(**training_params)
# Confirm that the training job has started
status = client.describe_training_job(TrainingJobName=model_job_name)["TrainingJobStatus"]
print("Training job current status: {}".format(status))
# -
# To check the progess of the training job, you can repeatedly evaluate the following cell. When the training job status reads `'Completed'`, move on to the next part of the tutorial.
client = boto3.client(service_name="sagemaker")
print(
"Training job status: ",
client.describe_training_job(TrainingJobName=model_job_name)["TrainingJobStatus"],
)
print(
"Secondary status: ",
client.describe_training_job(TrainingJobName=model_job_name)["SecondaryStatus"],
)
training_info = client.describe_training_job(TrainingJobName=model_job_name)
# # Deploy the Model
#
# Now that we've fully labeled our dataset and have a trained model, we want to use the model to perform inference.
#
# Object detection only supports encoded .jpg and .png image formats as inference input for now. The output is in JSON format, or in JSON Lines format for batch transform.
#
# This section involves several steps:
# 1. Create Model: Create model for the training output
# 2. Batch Transform: Create a transform job to perform batch inference.
# 3. Host the model for realtime inference: Create an inference endpoint and perform realtime inference.
# ## Create Model
# +
timestamp = time.strftime("-%Y-%m-%d-%H-%M-%S", time.gmtime())
model_name = "groundtruth-demo-od-model" + timestamp
print(model_name)
model_data = training_info["ModelArtifacts"]["S3ModelArtifacts"]
print(model_data)
primary_container = {
"Image": training_image,
"ModelDataUrl": model_data,
}
create_model_response = sagemaker_client.create_model(
ModelName=model_name, ExecutionRoleArn=role, PrimaryContainer=primary_container
)
print(create_model_response["ModelArn"])
# -
# ## Batch Transform
# We now create a SageMaker Batch Transform job using the model created above to perform batch prediction.
#
# ### Download Test Data
# First, let's download a test image that has been held out from the training and validation data.
# Find a bird not in the images labeled by Ground Truth.
img_ids = {img.filename.split(".")[0] for img in output_images}
with open("test-annotations-bbox.csv", "r") as f:
for line in f.readlines()[1:]:
line = line.strip().split(",")
img_id, _, cls_id, conf, xmin, xmax, ymin, ymax, *_ = line
if img_id in skip_these_images:
continue
if cls_id in good_subclasses:
# Skip the first several images
if str(img_id) not in img_ids:
test_bird = img_id
break
# +
from IPython.display import Image
test_image = test_bird + ".jpg"
os.system(f"wget https://s3.amazonaws.com/open-images-dataset/test/{test_image}")
Image(test_image)
# +
batch_input = "s3://{}/{}/test/".format(BUCKET, EXP_NAME)
print(test_image)
# !aws s3 cp $test_image $batch_input
# +
timestamp = time.strftime("-%Y-%m-%d-%H-%M-%S", time.gmtime())
batch_job_name = "object-detection-model" + timestamp
request = {
"TransformJobName": batch_job_name,
"ModelName": model_name,
"MaxConcurrentTransforms": 16,
"MaxPayloadInMB": 6,
"BatchStrategy": "SingleRecord",
"TransformOutput": {
"S3OutputPath": "s3://{}/{}/{}/output/".format(BUCKET, EXP_NAME, batch_job_name)
},
"TransformInput": {
"DataSource": {"S3DataSource": {"S3DataType": "S3Prefix", "S3Uri": batch_input}},
"ContentType": "application/x-image",
"SplitType": "None",
"CompressionType": "None",
},
"TransformResources": {"InstanceType": "ml.p2.xlarge", "InstanceCount": 1},
}
print("Transform job name: {}".format(batch_job_name))
# +
sagemaker_client = boto3.client("sagemaker")
sagemaker_client.create_transform_job(**request)
print("Created Transform job with name: ", batch_job_name)
while True:
response = sagemaker_client.describe_transform_job(TransformJobName=batch_job_name)
status = response["TransformJobStatus"]
if status == "Completed":
print("Transform job ended with status: " + status)
break
if status == "Failed":
message = response["FailureReason"]
print("Transform failed with the following error: {}".format(message))
raise Exception("Transform job failed")
time.sleep(30)
# -
# ### Inspect the results
#
# The following cell plots the predicted bounding boxes for our example image. You'll notice that inside the function `get_predictions`, we filter the output to only include bounding boxes with a confidence score above a certain threshold (in this case, 0.2). This is because the object detection model we have trained always ouputs a fixed number of box candidates, and we must include a cutoff to eliminate the spurious results.
# +
s3_client = boto3.client("s3")
batch_output = "s3://{}/{}/{}/output/".format(BUCKET, EXP_NAME, batch_job_name)
def list_objects(s3_client, bucket, prefix):
response = s3_client.list_objects(Bucket=bucket, Prefix=prefix)
objects = [content["Key"] for content in response["Contents"]]
return objects
def get_predictions(s3_client, bucket, prefix):
filename = prefix.split("/")[-1]
s3_client.download_file(bucket, prefix, filename)
with open(filename) as f:
data = json.load(f)
predictions = data["prediction"]
# Return only the predictions with confidence above the threshold of 0.2.
return [prediction for prediction in predictions if prediction[1] > 0.2]
def make_predicted_image(predictions, img_id, uri):
"""Maked a BoxedImage object with output of batch/realtime prediction.
Args:
predictions: list, output of get_predictions.
uri: str, s3 uri of input image.
Returns:
BoxedImage object with predicted bounding boxes.
"""
img = BoxedImage(id=img_id, uri=uri)
img.download(".")
imread_img = img.imread()
imh, imw, *_ = imread_img.shape
# Create boxes.
for batch_data in batch_boxes_data:
class_id, confidence, xmin, ymin, xmax, ymax = batch_data
boxdata = {
"class_id": class_id,
"height": (ymax - ymin) * imh,
"width": (xmax - xmin) * imw,
"left": xmin * imw,
"top": ymin * imh,
}
box = BoundingBox(boxdata=boxdata, image_id=img.id)
img.consolidated_boxes.append(box)
return img
inputs = list_objects(s3_client, BUCKET, urlparse(batch_input).path.lstrip("/"))
print("Input: " + str(inputs[:2]))
outputs = list_objects(s3_client, BUCKET, urlparse(batch_output).path.lstrip("/"))
print("Output: " + str(outputs[:2]))
# Download prediction results.
batch_boxes_data = get_predictions(s3_client, BUCKET, outputs[0])
batch_uri = f"s3://{BUCKET}/{inputs[0]}"
batch_img = make_predicted_image(batch_boxes_data, "BatchTest", batch_uri)
# Plot the image and predicted boxes.
fig, ax = plt.subplots()
batch_img.plot_consolidated_bbs(ax)
# -
# ## Realtime Inference
#
# We now host the model with an endpoint and perform realtime inference.
#
# This section involves several steps:
#
# 1. Create endpoint configuration - Create a configuration defining an endpoint.
# 2. Create endpoint - Use the configuration to create an inference endpoint.
# 3. Perform inference - Perform inference on some input data using the endpoint.
# 4. Clean up - Delete the endpoint and model
#
# ### Create Endpoint Configuration
# +
timestamp = time.strftime("-%Y-%m-%d-%H-%M-%S", time.gmtime())
endpoint_config_name = job_name_prefix + "-epc" + timestamp
endpoint_config_response = sagemaker_client.create_endpoint_config(
EndpointConfigName=endpoint_config_name,
ProductionVariants=[
{
"InstanceType": "ml.m4.xlarge",
"InitialInstanceCount": 1,
"ModelName": model_name,
"VariantName": "AllTraffic",
}
],
)
print("Endpoint configuration name: {}".format(endpoint_config_name))
print("Endpoint configuration arn: {}".format(endpoint_config_response["EndpointConfigArn"]))
# -
# ### Create Endpoint
#
# The next cell creates an endpoint that can be validated and incorporated into production applications. This takes about 10 minutes to complete.
# +
timestamp = time.strftime("-%Y-%m-%d-%H-%M-%S", time.gmtime())
endpoint_name = job_name_prefix + "-ep" + timestamp
print("Endpoint name: {}".format(endpoint_name))
endpoint_params = {
"EndpointName": endpoint_name,
"EndpointConfigName": endpoint_config_name,
}
endpoint_response = sagemaker_client.create_endpoint(**endpoint_params)
print("EndpointArn = {}".format(endpoint_response["EndpointArn"]))
# get the status of the endpoint
response = sagemaker_client.describe_endpoint(EndpointName=endpoint_name)
status = response["EndpointStatus"]
print("EndpointStatus = {}".format(status))
# wait until the status has changed
sagemaker_client.get_waiter("endpoint_in_service").wait(EndpointName=endpoint_name)
# print the status of the endpoint
endpoint_response = sagemaker_client.describe_endpoint(EndpointName=endpoint_name)
status = endpoint_response["EndpointStatus"]
print("Endpoint creation ended with EndpointStatus = {}".format(status))
if status != "InService":
raise Exception("Endpoint creation failed.")
# -
# ### Perform inference
#
# The following cell transforms the image into the appropriate format for realtime prediction, submits the job, receives the prediction from the endpoint, and plots the result.
# +
with open(test_image, "rb") as f:
payload = f.read()
payload = bytearray(payload)
client = boto3.client("sagemaker-runtime")
response = client.invoke_endpoint(
EndpointName=endpoint_name, ContentType="application/x-image", Body=payload
)
result = response["Body"].read()
result = json.loads(result)
predictions = [prediction for prediction in result["prediction"] if prediction[1] > 0.2]
realtime_uri = batch_uri
realtime_img = make_predicted_image(predictions, "RealtimeTest", realtime_uri)
# Plot the realtime prediction.
fig, ax = plt.subplots()
realtime_img.download(".")
realtime_img.plot_consolidated_bbs(ax)
# -
# ### Clean up
#
# Finally, let's clean up and delete this endpoint.
boto3.client(service_name="sagemaker").delete_endpoint(EndpointName=endpoint_name)
# # Review
#
# We covered a lot of ground in this notebook! Let's recap what we accomplished. First we started with an unlabeled dataset (technically, the dataset was previously labeled by the authors of the dataset, but we discarded the original labels for the purposes of this demonstration). Next, we created a SageMake Ground Truth labeling job and generated new labels for all of the images in our dataset. Then we split this file into a training set and a validation set and trained a SageMaker object detection model. Next, we trained a new model using these Ground Truth results and submitted a batch job to label a held-out image from the original dataset. Finally, we created a hosted model endpoint and used it to make a live prediction for the same held-out image.
|
ground_truth_object_detection_tutorial/object_detection_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Coloring bars
#
# Demo of using color params for barchart-based visualizers
# +
import numpy as np
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import LabelEncoder
from sklearn.datasets import make_classification
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import StratifiedKFold
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.preprocessing import LabelEncoder, OrdinalEncoder
from yellowbrick.features import Rank1D
from yellowbrick.datasets import load_game
from yellowbrick.datasets import load_hobbies
from yellowbrick.datasets import load_concrete
from yellowbrick.datasets import load_occupancy
from yellowbrick.text import FreqDistVisualizer
from yellowbrick.model_selection import CVScores
from yellowbrick.features import FeatureImportances
from yellowbrick.contrib.missing import MissingValuesBar
from yellowbrick.target import ClassBalance, FeatureCorrelation
# -
# ## Rank1D
#
# has a `color` param now
# +
X, y = load_concrete()
visualizer = Rank1D(algorithm='shapiro', color=["cadetblue"])
visualizer.fit(X, y)
visualizer.transform(X)
visualizer.poof()
# -
# ## FreqDistVisualizer
#
# has a `color` param now
# +
corpus = load_hobbies()
y = LabelEncoder().fit_transform(corpus.target)
vectorizer = CountVectorizer(stop_words='english')
docs = vectorizer.fit_transform(corpus.data)
features = vectorizer.get_feature_names()
visualizer = FreqDistVisualizer(
features=features, orient='v', size=(600, 300), color=["crimson"]
)
visualizer.fit(docs)
visualizer.poof()
# -
# ## CVScores
#
# has a `color` param now
# +
X, y = load_occupancy()
cv = StratifiedKFold(n_splits=12, random_state=42)
visualizer = CVScores(
GaussianNB(), cv=cv, scoring='f1_weighted', color="goldenrod"
)
visualizer.fit(X, y)
visualizer.poof()
# -
# ## FeatureImportances
#
# has a `colors` and a `colormap` param now;
# `colors` is for `stack==False` and `colormap` is for `stack==True`
# +
X, y = load_occupancy()
model = RandomForestClassifier(n_estimators=10)
colors = ["lightpink", "pink", "hotpink", "crimson", "orchid"]
viz = FeatureImportances(model, colors=colors)
viz.fit(X, y)
viz.poof()
# +
X, y = load_game()
X = OrdinalEncoder().fit_transform(X)
y = LabelEncoder().fit_transform(y)
model = LogisticRegression(multi_class="auto", solver="liblinear")
viz = FeatureImportances(model, stack=True, relative=False, colormap="viridis")
viz.fit(X, y)
viz.poof()
# -
# ## MissingValuesBar
# +
# Make a classification dataset
X, y = make_classification(
n_samples=400, n_features=10, n_informative=2, n_redundant=3,
n_classes=2, n_clusters_per_class=2, random_state=854
)
# Assign NaN values
X[X > 1.5] = np.nan
features = ["Feature {}".format(str(n)) for n in range(10)]
visualizer = MissingValuesBar(features=features, color="lime")
visualizer.fit(X)
visualizer.poof()
# -
# ## ClassBalance
# +
X, y = load_game()
visualizer = ClassBalance(
labels=["draw", "loss", "win"],
colormap="copper"
)
visualizer.fit(y)
visualizer.poof()
# -
# ## FeatureCorrelation
# +
X, y = load_concrete(return_dataset=True).to_pandas()
# Create a list of the feature names
features = [
"cement", "slag", "ash", "water", "splast", "coarse", "fine", "age"
]
# Instantiate the visualizer
visualizer = FeatureCorrelation(labels=features, color="rebeccapurple")
visualizer.fit(X, y)
visualizer.poof()
|
examples/rebeccabilbro/barchart_colors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
from scipy.stats import norm
import statsmodels.api as sm
import pandas as pd
# +
# Aufgabe 5.2
# Calculate n', the equivalent number of independent samples,
# for the two sets of minimum air temperature in Table A1
# Ithaca Tmin
# Canadaigua Tmin
# +
# A1 Daily Precipitation and Temperature (F) Observations
# at Ithaca and Canadaigua, New York, for January 1987
Tmin_Ithaca = np.array([19,25,22,-1,4,14,21,22,23,27,29,25,29,15,29,24,0,2,26,17,19,9,20,-6,-13,-13,-11,-4,-4,11,23])
Tmin_Canadaigua = np.array([28,28,26,19,16,24,26,24,24,29,29,27,31,26,38,23,13,14,28,19,19,17,22,2,4,5,7,8,14,14,23]) # Canandaigua T max
print(Tmin_Ithaca.shape)
print(Tmin_Canadaigua.shape)
# -
plt.plot(Tmin_Ithaca)
plt.plot(Tmin_Canadaigua)
# +
#np.vectorize(res.autocorr)(np.arange(0,len(res)-1))
rho_lag_cndg=pd.Series(sm.tsa.acf(Tmin_Canadaigua, nlags=5,fft=False))
rho_lag_ithc=pd.Series(sm.tsa.acf(Tmin_Ithaca, nlags=5,fft=False))
print(rho_lag_cndg)
plt.plot(rho_lag_cndg)
print(rho_lag_ithc)
plt.plot(rho_lag_ithc)
# +
n_eff_itch=Tmin_Ithaca.shape[0]*(1-rho_lag_ithc[1])/(1+rho_lag_ithc[1])
n_eff_cndg=Tmin_Canadaigua.shape[0]*(1-rho_lag_cndg[1])/(1+rho_lag_cndg[1])
print('n: ',Tmin_Canadaigua.shape[0])
print('lag-1 Autocorrelation rho(1): ',rho_lag_ithc[1])
print('n_eff @ Ithaca: ',n_eff_itch)
print('n Canadaigua: ',Tmin_Canadaigua.shape[0])
print('lag-1 Autocorrelation rho(1): ',rho_lag_cndg[1])
print('n_eff @ Canadaigua: ',n_eff_cndg)
# -
|
Chapter_5/Wilks-Aufgabe - 5.2 - Effective Length of paired data - 01-MAR-2021.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tweepy
import json
import csv
from datetime import date
from datetime import datetime
import time
secrets = {
"consumer_key":"XXXX",
"consumer_secret":"XXXX",
"access_token":"XXXX",
"access_token_secret":"XXXX"
}
# +
consumer_key = secrets['consumer_key']
consumer_secret = secrets['consumer_secret']
access_token = secrets['access_token']
access_token_secret = secrets['access_token_secret']
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
api = tweepy.API(auth,wait_on_rate_limit=True)
# +
queryList = [
"#detox#suco",
"#suco#natural",
"#zeroalcool",
"semalcool",
"#noalcohol",
"#sonaagua",
"#suco",
"#refrigerante"
]
size = 1000000
language = "pt"
start_date = "2015-04-03"
data = []
for query in queryList:
for tweet in tweepy.Cursor(api.search,q=query,count=size,lang=language,since=start_date).items():
data.append(tweet)
print(tweet.created_at, tweet.text)
# -
|
data-extraction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pachterlab/MBGBLHGP_2019/blob/master/notebooks/merge/split_runtime.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="7LrGhLCQUf6W" outputId="392d7dd4-e836-4ad8-9c13-9d6308ed68af" colab={"base_uri": "https://localhost:8080/", "height": 453}
# install kallisto and bustools and seqkit
# !git clone https://github.com/pachterlab/MBGBLHGP_2019.git
# !cp MBGBLHGP_2019/notebooks/merge/kallisto /usr/local/bin/
# !cp MBGBLHGP_2019/notebooks/merge/bustools /usr/local/bin/
# !wget https://github.com/shenwei356/seqkit/releases/download/v0.13.2/seqkit_linux_amd64.tar.gz && tar -xvf seqkit_linux_amd64.tar.gz && cp seqkit /usr/local/bin/
# + id="nQd5MhOVcTT_"
# !mkdir -p split2 split4 split8 split10 full ref/split2 ref/split4 ref/split8 ref/split10 ref/full
# + id="aiWXEt32cBk7" outputId="c610187f-cf83-4900-ca0c-b501089ab363" colab={"base_uri": "https://localhost:8080/", "height": 277}
# download the transcriptome
# !wget ftp://ftp.ensembl.org/pub/release-101/fasta/homo_sapiens/cdna/Homo_sapiens.GRCh38.cdna.all.fa.gz
# + id="9CuzJy04d9_R"
# download fastqs
# !wget --quiet -O r1.fastq.gz https://caltech.box.com/shared/static/6boju5zerptobm51fkbq5zwmchjhhk92.gz
# !wget --quiet -O r2.fastq.gz https://caltech.box.com/shared/static/adme7zu1y8nz4ng2ph5wjbei6unvy093.gz
# + [markdown] id="o4gm7TSDgHG8"
# ## Full transcriptome
# + id="nKIa8F6TdH4A" outputId="bda0bbca-f839-4186-b28f-d97e302f81e8" colab={"base_uri": "https://localhost:8080/", "height": 280}
# !kallisto index -i ref/full/index.idx Homo_sapiens.GRCh38.cdna.all.fa.gz
# + id="uYxLIuCQgM8i"
# !zcat Homo_sapiens.GRCh38.cdna.all.fa.gz | awk '(NR-1)%2==0{print $1, $4}' OFS="\t"| tr -d "gene:" | tr -d ">" > ref/full/t2g.txt
# + [markdown] id="BBekqijZgKSP"
# ## Split transcriptome
# + id="8zWVcJiicWkl" outputId="b536d629-6ede-4897-d941-31a39a8ab652" colab={"base_uri": "https://localhost:8080/", "height": 104}
# !seqkit split -p 2 Homo_sapiens.GRCh38.cdna.all.fa.gz
# !mv Homo_sapiens.GRCh38.cdna.all.fa.gz.split/ ref/split2/
# + id="CEyOazK_A5mp" outputId="f69466d5-f629-4410-daed-ae9e7b782ffe" colab={"base_uri": "https://localhost:8080/", "height": 139}
# !seqkit split -p 4 Homo_sapiens.GRCh38.cdna.all.fa.gz
# !mv Homo_sapiens.GRCh38.cdna.all.fa.gz.split/ ref/split4/
# + id="XLXk_kxpBE-4" outputId="7c0dfc4b-29b1-4cda-bbfb-2e530a85491e" colab={"base_uri": "https://localhost:8080/", "height": 208}
# !seqkit split -p 8 Homo_sapiens.GRCh38.cdna.all.fa.gz
# !mv Homo_sapiens.GRCh38.cdna.all.fa.gz.split/ ref/split8/
# + id="tvWr33F6xO-8" outputId="9533f86f-ed7c-4f63-a159-6ba6da949b7e" colab={"base_uri": "https://localhost:8080/", "height": 280}
# !seqkit split -p 10 Homo_sapiens.GRCh38.cdna.all.fa.gz
# !mv Homo_sapiens.GRCh38.cdna.all.fa.gz.split/ ref/split10/
# + [markdown] id="Sl610JL6BK_3"
# # Build the indices
# + id="za7Hc8KCc4SG" outputId="15cde4dc-96ed-4885-8580-617dcb87273c" colab={"background_save": true, "base_uri": "https://localhost:8080/", "height": 188}
# !cd ref/split2/Homo_sapiens.GRCh38.cdna.all.fa.gz.split/ && for f in *.gz; do kallisto index -i $(echo $f | cut -d"." -f5).idx $f; done
# + id="Lfti40LVCj0X" outputId="4766e768-7ae1-4d49-9426-84cce5203fe6" colab={"background_save": true}
# !cd ref/split4/Homo_sapiens.GRCh38.cdna.all.fa.gz.split/ && for f in *.gz; do kallisto index -i $(echo $f | cut -d"." -f5).idx $f; done
# + id="YFF-PjVBDxV6" outputId="88daf3d5-c761-4609-f468-6234f32a1778" colab={"background_save": true}
# !cd ref/split8/Homo_sapiens.GRCh38.cdna.all.fa.gz.split/ && for f in *.gz; do kallisto index -i $(echo $f | cut -d"." -f5).idx $f; done
# + id="Pg2LRWxaxpAY" outputId="e317d6e7-ca1f-4a49-cfef-0b3a1150d67f" colab={"base_uri": "https://localhost:8080/", "height": 35}
# !echo Homo_sapiens.GRCh38.cdna.all.part_010.fa.gz | cut -d"." -f5
# + id="FRR5O_9Oxhhf" outputId="e072ef33-3d23-4083-e5b7-64fcac04d7ea" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !cd ref/split10/Homo_sapiens.GRCh38.cdna.all.fa.gz.split/ && for f in *.gz; do kallisto index -i $(echo $f | cut -d"." -f5).idx $f; done
# + id="RQVY4FElenBM" colab={"background_save": true}
# !mv Homo_sapiens.GRCh38.cdna.all.fa.gz ref/full
# + [markdown] id="f_L_u_Fel1OG"
# ## Full alignment
# + id="JDAl_pTnmFyH" outputId="b42456ca-402e-4e1c-deb6-ff3aed84ff21" colab={"background_save": true}
# %%time
# kallisto bus on full guy
# !time kallisto bus -x 10xv2 -i ref/full/index.idx -o full/o \
# r1.fastq.gz \
# r2.fastq.gz
# + id="teVg7bzMEkZc"
# !rm -rf ./full
# + id="rsP4dWxrL4n3"
# !mkdir split2/o1 split2/o2
# + id="O9lSfgUCL8h1" outputId="e2a7b40a-9f88-4e8c-9a9a-c164bed1d747" colab={"background_save": true, "base_uri": "https://localhost:8080/", "height": 379}
# %%time
# !for d in ./split2/o*; do kallisto bus -k -n -x 10xv2 -o $d \
# -i ref/split2/Homo_sapiens.GRCh38.cdna.all.fa.gz.split/part_00$(echo $d | cut -c11).idx \
# r1.fastq.gz \
# r2.fastq.gz; done
# + id="HLkIN-U3EiVc"
# !rm -rf ./split2
# + id="flsY7ZPcMFq0" colab={"background_save": true}
# !mkdir split4/o1 split4/o2 split4/o3 split4/o4
# + id="YmmVkhEDMFYv" outputId="ac0f1580-557b-4eb1-c0f5-d8f9e2798203" colab={"base_uri": "https://localhost:8080/", "height": 713}
# %%time
# !for d in ./split4/o*; do kallisto bus -k -n -x 10xv2 -o $d \
# -i ref/split4/Homo_sapiens.GRCh38.cdna.all.fa.gz.split/part_00$(echo $d | cut -c11).idx \
# r1.fastq.gz \
# r2.fastq.gz; done
# + id="EBvmaXSdEfI8"
# !rm -rf ./split4
# + id="4WF6F-3Ml9Nx"
# !mkdir split8/o1 split8/o2 split8/o3 split8/o4 split8/o5 split8/o6 split8/o7 split8/o8
# + id="fIp88RvYp_lj" outputId="75633e0d-7fa5-4202-c178-a0f0dc4c39fa" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %%time
# !for d in ./split8/o*; do kallisto bus -k -n -x 10xv2 -o $d \
# -i ref/split8/Homo_sapiens.GRCh38.cdna.all.fa.gz.split/part_00$(echo $d | cut -c11).idx \
# r1.fastq.gz \
# r2.fastq.gz; done
# + id="-f8-WZFO1m7b" outputId="79545434-ac9a-4802-c78b-cec4a6a8766b" colab={"base_uri": "https://localhost:8080/", "height": 173}
# !mkdir split10/o1 split10/o2 split10/o3 split10/o4 split10/o5 split10/o6 split10/o7 split10/o8 split10/o9 split10/o0
# + id="Fknjz2x_2Tzt"
# to make our lives easier we call index 10 index 0
# !mv ref/split10/Homo_sapiens.GRCh38.cdna.all.fa.gz.split/part_010.idx ref/split10/Homo_sapiens.GRCh38.cdna.all.fa.gz.split/part_000.idx
# + id="aXTjReOH1uRh" outputId="2a432092-c683-4aa0-f2c6-b2ea2d451118" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %%time
# !for d in ./split10/o*; do kallisto bus -k -n -x 10xv2 -o $d \
# -i ref/split10/Homo_sapiens.GRCh38.cdna.all.fa.gz.split/part_00$(echo $d | cut -c12).idx \
# r1.fastq.gz \
# r2.fastq.gz; done
# + id="SVgQE1gaB5xm"
# !rm -rf ./split8
# + id="OJnVOLCjMXIB"
num_i = [1, 2, 4, 8, 10]
times = [7*60+ 6, 18*60+ 17, 36*60+ 48, 1*60*60 +19*60 + 26, 1*60*60 + 25*60 + 9]
# + id="Un1ke1xzxoOF"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os
import string
from scipy import stats
from scipy.io import mmread
from collections import defaultdict
from sklearn.linear_model import LinearRegression
def nd(arr):
return np.asarray(arr).reshape(-1)
def yex(ax):
lims = [
np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes
]
# now plot both limits against eachother
ax.plot(lims, lims, 'k-', alpha=0.75, zorder=0)
ax.set_aspect('equal')
ax.set_xlim(lims)
ax.set_ylim(lims)
return ax
fsize=15
plt.rcParams.update({'font.size': fsize})
# %config InlineBackend.figure_format = 'retina'
# + id="P_uKSXSvBRPE" outputId="5b2ac661-1904-43b5-b997-377a67866fb1" colab={"base_uri": "https://localhost:8080/", "height": 615}
fig,ax = plt.subplots(figsize=(10,10))
x = nd(num_i)
y = nd(times)/60
X, Y = x.reshape(-1,1), y.reshape(-1,1)
lr = LinearRegression().fit(X, Y)
ax.scatter(x, y, color="k")
ax.plot(X, lr.predict(X), color="k", label=f"{lr.coef_[0][0]:.2f} min/split")
kwd = {
"xlabel": "Number of indices",
"ylabel": "Minutes"
}
ax.set(**kwd)
ax.legend()
plt.savefig("./graph.png",bbox_inches='tight', dpi=300)
fig.show()
# + id="etfnhjBLDJjL" outputId="6e9f0e29-3135-4d08-c64c-0dfaece6bca6" colab={"base_uri": "https://localhost:8080/", "height": 87}
# ! ls -lht ./ref/full
# + id="Vr4Bwkn3GDg6" outputId="3c087201-75bc-42ae-d055-82320dc39de8" colab={"base_uri": "https://localhost:8080/", "height": 52}
# !ls -lht ./ref/split2/*/*.idx
# + id="9lc-PZP3GK9k" outputId="d407ac6e-86d3-40b1-a424-f14531856003" colab={"base_uri": "https://localhost:8080/", "height": 87}
# !ls -lht ./ref/split4/*/*.idx
# + id="Cr06TwZMGYej" outputId="b78a99b9-17e5-477a-da31-ac139cba6736" colab={"base_uri": "https://localhost:8080/", "height": 156}
# !ls -lht ./ref/split8/*/*.idx
# + id="Laok4dWnGZUG" outputId="82157a75-2631-4bea-84ee-99a81a723e72" colab={"base_uri": "https://localhost:8080/", "height": 191}
# !ls -lht ./ref/split10/*/*.idx
# + id="fI-x1dkjgd4C"
|
notebooks/merge/split_runtime.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # WeatherPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import os
import json
import random
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import time
import requests
import datetime as dt # to put a datestamp on the outputs
from pprint import pprint
from scipy.stats import linregress
# Import API key
from api_keys import weather_api_key
from api_keys import g_key
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
"""" Output File (CSV)
Create file name output_data in starter_code for csv file
Create file name Images for saving plots"""
output_data_file = "output_data/cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
# -
# ## Generate Cities List
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(low=-90.000, high=90.000, size=1500)
lngs = np.random.uniform(low=-180.000, high=180.000, size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
""" Print the city count to confirm sufficient count
number of cities change (when i rerun program) because of randonly selection of lat and long
same thing will happen for all cells"""
len(cities)
# +
#create url
url = "http://api.openweathermap.org/data/2.5/weather?"
units = "imperial"
query_url = f"{url}appid={weather_api_key}&units={units}&q="
# Create empty lists to store the data for each field
city_id_list = []
city_name_list = []
country_list = []
lng_list = []
lat_list = []
temp_list = []
humidity_list = []
clouds_list = []
wind_speed_list = []
# Loop through cities
for city in cities:
# Get the JSON response from the OpenWeather API
response_json = requests.get(query_url + city).json()
# Start the error handling
# Retrieve the items for each field and append them to the empty lists
try:
city_id = response_json['id']
city_id_list.append(city_id)
city_name = response_json['name']
city_name_list.append(city_name)
country_name = response_json['sys']['country']
country_list.append(country_name)
lng = response_json['coord']['lon']
lng_list.append(lng)
lat = response_json['coord']['lat']
lat_list.append(lat)
temp = response_json['main']['temp']
temp_list.append(temp)
humidity = response_json['main']['humidity']
humidity_list.append(humidity)
clouds = response_json['clouds']['all']
clouds_list.append(clouds)
wind_speed = response_json['wind']['speed']
wind_speed_list.append(wind_speed)
print(f"City Name: {city}, City ID: {city_id}")
# Print error message and move on to the next city if there is an error
except:
print("That record does not exist, searching for next record...")
# -
# ### Convert Raw Data to DataFrame
# * Export the city data into a .csv.
# * Display the DataFrame
cities_df = pd.DataFrame({"City ID": city_id_list, "City": city_name_list,
"Country": country_list, "Lat": lat_list, "Lng": lng_list,
"Temperature": temp_list, "Humidity": humidity_list, "Clouds": clouds_list,
"Wind Speed": wind_speed_list})
cities_df.head()
# Export to .csv file
cities_df.to_csv(output_data_file, index_label='City_ID')
#describe function will give basic statistic values
#check the max humidity level for code
cities_df.describe()
# ## Inspect the data and remove the cities where the humidity > 100%.
# ----
# Skip this step if there are no cities that have humidity > 100%.
# +
#max humidity level is 100.
# +
# Get the indices of cities that have humidity over 100%.
#max humidity level is 100.
# -
# Make a new DataFrame equal to the city data to drop all humidity outliers by index.
# Passing "inplace=False" will make a copy of the city_data DataFrame, which we call "clean_city_data".
# ## Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# ## Latitude vs. Temperature Plot
""" refer cities_df data frame
check unit of variable
Images is the path way for saving picture ,created before in starter_code folder
plt.savefig should be before plt.show
"""
x_values = cities_df["Lat"]
y_values = cities_df["Temperature"]
plt.scatter(x_values,y_values,alpha=.75, edgecolors="k")
plt.title('Temperatures at Latitude Positions')
plt.xlabel('Latitude')
plt.ylabel('Temperature (F)')
plt.savefig("Images/lat_temp.png")
plt.show()
# Scatter plot shows when latitude is between -20 to +20 temperature is high and when latitude change from 20 to 80 temperature is decreasing.
# ## Latitude vs. Humidity Plot
# +
#check unit of variable
x_values = cities_df["Lat"]
y_values = cities_df["Humidity"]
plt.scatter(x_values,y_values,alpha=.75, edgecolors="k")
plt.title('Humidity at Latitude Positions')
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
plt.savefig("Images/lat_humidity.png")
plt.show()
# -
# There is clear correlation between lat and humidity.
# ## Latitude vs. Cloudiness Plot
# +
x_values = cities_df["Lat"]
y_values = cities_df["Clouds"]
plt.scatter(x_values,y_values, alpha=.75, edgecolors="k")
plt.title('Cloudiness at Latitude Positions')
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
plt.savefig("Images/lat_clouds.png")
plt.show()
# -
# There is clear correlation between lat and clouds.
# ## Latitude vs. Wind Speed Plot
# +
#unit of wind speed mph
x_values = cities_df["Lat"]
y_values = cities_df["Wind Speed"]
plt.scatter(x_values,y_values, alpha=.75, edgecolors="k")
plt.title('Wind Speeds at Latitude Positions')
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (mph)')
plt.savefig("Images/lat_wind.png")
plt.show()
# -
# There is clear correlation between lat and wind speed.
# ## Linear Regression
# +
"""Create Northern and Southern Hemisphere DataFrames
mask funtion will allows us to sort data depend on lat. since latitude of equador is zero ,
lat>0 will give data for northern hemisphere
~mask(lat<0) function will give us southern hemisphere """
mask = cities_df['Lat'] > 0
northern_hemisphere = cities_df[mask]
southern_hemisphere = cities_df[~mask]
# -
# #### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
#create the linear regression and check the spot for formula and units for label if applicable
#use str for any rounding
#similar codes homework 5
x_values = northern_hemisphere["Lat"]
y_values = northern_hemisphere["Temperature"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values, alpha=.75, edgecolors="k")
plt.title('Temperatures at Latitude Positions (Northern Hemisphere)')
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(50,95),fontsize=12,color="red")
plt.xlabel('Latitude')
plt.ylabel('Temperature (F)')
plt.savefig("Images/lat_temp_north.png")
plt.show()
print(rvalue)
# -
# r=-08889 that means there is strong correlation between temperature and latitude.
# #### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
x_values = southern_hemisphere["Lat"]
y_values = southern_hemisphere["Temperature"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values, alpha=.75, edgecolors="k")
plt.title('Temperatures at Latitude Positions (Southern Hemisphere)')
plt.plot(x_values,regress_values,"r-")
#for annotation look at the scatter plot and choose the best coordinate(x,y) for clear visualization
plt.annotate(line_eq,(-50,95),fontsize=12,color="red")
plt.xlabel('Latitude')
plt.ylabel('Temperature (F)')
plt.savefig("Images/lat_temp_south.png")
plt.show()
print(rvalue)
# -
# r=.638 that means there is moderate correlation between temperature and latitude.
#
# #### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# +
x_values = northern_hemisphere["Lat"]
y_values = northern_hemisphere["Humidity"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values, alpha=.75, edgecolors="k")
plt.title('Humidity at Latitude Positions (Northern Hemisphere)')
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(50,20),fontsize=13,color="red")
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
plt.savefig("Images/lat_humidity_north")
plt.show()
print(rvalue)
# -
# r=.236 shows there is weak correlation between latitude and humudity.
# #### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# +
x_values = southern_hemisphere["Lat"]
y_values = southern_hemisphere["Humidity"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
print(rvalue)
plt.scatter(x_values,y_values, alpha=.75, edgecolors="k")
plt.title('Humidity at Latitude Positions (Southern Hemisphere)')
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(-30,20),fontsize=12,color="red")
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
plt.savefig("Images/lat_humudity_south")
plt.show()
print(rvalue)
# -
# r=.271 shows there is weak correlation between latitude and humudity.
# #### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# +
x_values = northern_hemisphere["Lat"]
y_values = northern_hemisphere["Clouds"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values, alpha=.75, edgecolors="k")
plt.title('Cloudiness at Latitude Positions (Northern Hemisphere)')
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(20,62),fontsize=12,color="red")
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
plt.savefig("Images/lat_cloudiness_north")
plt.show()
print(rvalue)
# -
# r=.346 shows there is weak correlation between latitude and cloudiness.
# #### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# +
x_values = southern_hemisphere["Lat"]
y_values = southern_hemisphere["Clouds"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values, alpha=.75, edgecolors="k")
plt.title('Cloudiness at Latitude Positions (Southern Hemisphere)')
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(-55,50),fontsize=12,color="red")
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
plt.savefig("Images/lat_cloudiness_south")
plt.show()
print(rvalue)
# -
# r=.153 shows there is weak correlation between latitude and cloudiness.
# #### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# +
x_values = northern_hemisphere["Lat"]
y_values = northern_hemisphere["Wind Speed"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values, alpha=.75, edgecolors="k")
plt.title('Wind Speed at Latitude Positions (Northern Hemisphere)')
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(10,30),fontsize=12,color="red")
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (mph)')
plt.savefig("Images/lat_windspeed_north")
plt.show()
print(rvalue)
# -
# there is weak correlation between latitude and wind speed.
# #### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# +
x_values = southern_hemisphere["Lat"]
y_values = southern_hemisphere["Wind Speed"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values,alpha=.75, edgecolors="k")
plt.title('Wind Speed at Latitude Positions (Southern Hemisphere)')
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(-55,40),fontsize=12,color="red")
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (mph)')
plt.savefig("Images/lat_windspeed_south")
plt.show()
print(rvalue)
# +
#all done
|
starter_code/WeatherPy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import HTML
def css_styling():
sheet = '../css/custom.css'
styles = open(sheet, "r").read()
return HTML(styles)
css_styling()
# # Python OOP 1: Basics and Initialisation
# This exercise is designed to motivate the use of object oriented programming in scientific computation via a simplified case using images. Your task is to program the Python `classes` which reads in the data from a file, manipulates the data and plots the image.
#
import os
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import numpy as np
import sys
# %matplotlib inline
# ## Portable Greymap ( `.pgm`) Format
# You have been provided with some image files i.e. `img1, .. img4` in the `data` directory in *portable greymap* (.pgm) format. Although admittedly a primitive image format, `.pgm` files are simple to manipulate as they contain only one pixel grey value per $x, y$ pixel in the image: the scale runs from 0 (black) to 255 (white). This represents a common task in scientific computing where you must read in some field data on a grid of points. **You are provided with the code to read and reshape this data from a file!**
#
# Here's a demonstration of a greymap image that might be stored in `.pgm` format using a simple gradient of white to black - it is displayed here using a contour plot:
# +
Nx = 72
Ny = 72
img_x = np.linspace(1, 0, Nx)
img_y = np.linspace(1, 0, Ny)
X, Y = np.meshgrid(img_x, img_y)
# Generate the gradient image - this could be stored in .pgm format!
img_z = (X+Y) * 255*0.5
print(img_z)
fig = plt.figure()
ax = fig.add_subplot(111, aspect='equal')
ax.contourf(img_x, img_y, img_z, 20, cmap=cm.Greys_r)
ax.set_xlabel('x')
ax.set_ylabel('y')
# -
# ## Task breakdown
#
# Create a `SquareImage` class which reads (portable greymap, '.pgm') image data from a file. The class should implement the following:
#
#
# 1. The Initialiser method `__init__(self, ...)`, which takes a string `filename` as an argument and
# - stores $Nx$ and $Ny$ as *instance attributes*, both equal to 72 (this isn't robust, but simplifies the exercise)
#
# - calculates and stores $x$ and $y$ as *instance attrbutes*. These are again the regularly spaced pixel $x$ and $y$ values from 1 to 0 (image colour values in pgm files start from top right pixel) - use linspace from 1 to 0 with $Nx$ and $Ny$ points respectively
#
# - Read and store image grey levels in `filename` as an *instance attribute*, $z$. The line for extracting this data from the files is the same as before,
# >`np.loadtxt(filename, skiprows=4).reshape(self.Nx, self.Ny)`
#
# 2. An `add_image` method which takes an `image` argument, and adds the z component of image to `self.z`
# - don't forget to add `self` as the first argument! Instance methods require us to be specific
#
# 3. A `plot` method which takes no extra arguments, and plots the current instance attributes $z$ as a contour vs $x$ and $y$ (also instance attributes). As this exercise is not testing your matplotlib, we provide the code for the function here:
#
# > `fig = plt.figure()
# > ax = fig.add_subplot(111, aspect='equal')
# > ax.contourf(self.x, self.y, self.z, cmap=cm.Greys_r)
# > plt.show()`
#
# +
# Implement the class here:
class SquareImage(object):
def __init__(self, filename=None):
# To simplify this exercise, set the size of the image to a constant
# (Each image was specifically written to have size 72 by 72)
self.Nx = self.Ny = 72
self.x = np.linspace(1, 0, self.Nx)
self.y = np.linspace(1, 0, self.Ny)
if filename is None:
self.z = np.zeros([self.Nx, self.Ny])
else:
self.z = np.loadtxt(filename, skiprows=4).reshape(self.Nx, self.Ny)
def add_image(self, image):
"""Add the z values of another 72 by 72 image image to this instance"""
# Could add a defensive check here
assert(np.shape(image.z) == (self.Nx, self.Ny)), 'Image sizes are not equal!'
# Add the image z value to self:
self.z += image.z
def plot(self):
"""Plots the contour z against x and y coordinates"""
fig = plt.figure()
ax = fig.add_subplot(111, aspect='equal')
ax.contourf(self.x, self.y, self.z, cmap=cm.Greys_r)
plt.show()
# -
# The image file names
names = ['img1.pgm', 'img2.pgm', 'img3.pgm', 'img4.pgm']
files = [os.path.join('data', name) for name in names]
# Instantiate the class and plot each picture.
imgs = []
for f in files:
image = SquareImage(f)
print(image)
imgs.append(image) # objects are first class instances: add to a list
image.plot()
# ## Extension
#
# Now that we can read in image data from a file, extend your `SquareImages` class above so that if the filename is `None` (python keyword), we store the $z$ attribute as `np.zeros([Nx, Ny])`.
#
# * This will require an if statement, e.g.:
# > `if filename is None`
# > store z as zeros
# > `else`
# > read and store z data
#
# * The default `filename` argument should be `None`, so that SquareImage() produces an 'empty' image.
# Create an 'empty' SquareImage
combined = SquareImage()
print(combined.z)
# Now use the `add_image` method of the empty image to add on the contents of all other image in the list of `imgs`
# +
# Loop over the list of images
for image in imgs:
combined.add_image(image)
# Plot
combined.plot()
|
soln/01-Classes_basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# This is a notebook to explore opSim outputs in different ways, mostly useful to supernova analysis. We will look at the opsim output called (`minion_1016`)[https://confluence.lsstcorp.org/display/SIM/Summary+Table+Column+Descriptions].
# Other relevant OpSim outputs are listed on a [confluence page](https://confluence.lsstcorp.org/display/SIM/Operations+Simulator+Benchmark+Surveys). All of the newer outputs are in the format of sqlite databases (zipped up to save space), while older OpSim versions are in ASCII files. The database table which I think has all of the information we will need is the `Summary` Table. The quantities in the columns of this table are described (here)[https://confluence.lsstcorp.org/display/SIM/Summary+Table+Column+Descriptions]. The OpSim outputs are official products that the LSST project provides. This notebook will demonstrate a way of exploring these outputs (which is by no means official or project supplied).
#
# ``Note:`` The `Summary` Table is only about a GB in size, and has ~ million rows. For newish laptops, this is a small table and can be easily read into memory. For more constrained memory systems or computations that require a lot of memory, this may be a bad thing to do.
#
# ``Gotcha``: The column `obsHistID` is unique identifier of OpSim observations. Very often, you might end up with a table with multiple rows with the same obsHistID, but with other columns (like propID) having different values. These are not different observations, and a SN light curve corresponding to these observations should include only one of these.
# ### A bit more information about the columns from a SN perspective
#
# The LSST observations are over roughly half of the sky (~20000 sq degrees, or ~`2\pi`). This is covered by
# overlapping pointings of the telescope each covering (~10 sq degrees). These pointings are currently thought of as being located on a grid along with dithers. The grid locations of the pointings are assigned a unique integer called `fieldID` which is associated with its location :(`fieldRa`, `fieldDec` ). The actual location of the pointings (including dithers) are in (`ditheredRA`, `ditheredDec`). There are a number of columns that hold quantities of similar information, and the exact definitions of each quantity is provided in the [description](https://confluence.lsstcorp.org/display/SIM/Summary+Table+Column+Descriptions). For most purposes `filtSkyBrightness` and `FWHMeff` rather than the other similar looking quantities are probably recommended. `PropID` (also described there) refers to proposals under which the observation was taken. For the basic SN purposes, we will want the proposals (Wide Fast Deep (WFD)) and (Deep Drilling Fields (DDF)). The propID for these quantities differs from one opsim output to another. To find out what these are, you can use OpSim utils as in this notebook or look at the `PROPOSAL` table (5 lines) like this : and pick the integers corresponding to DDcosmology1.conf and Universal ...
# ```
# SELECT * FROM PROPOSAL;
# 362|../conf/survey/GalacticPlaneProp.conf|WL|27692624|enigma|1189
# 363|../conf/survey/SouthCelestialPole-18.conf|WL|27692368|enigma|1189
# 364|../conf/survey/Universal-18-0824B.conf|WLTSS|27692112|enigma|1189
# 365|../conf/survey/NorthEclipticSpur-18c.conf|WLTSS|27692240|enigma|1189
# 366|../conf/survey/DDcosmology1.conf|WLTSS|29065424|enigma|1189
# ```
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# Required packages sqlachemy, pandas (both are part of anaconda distribution, or can be installed with a python installer)
# One step requires the LSST stack, can be skipped for a particular OPSIM database in question
import opsimsummary as oss
import opsimsummary.summarize_opsim as so
from sqlalchemy import create_engine
import pandas as pd
print so.__file__
# +
# This step requires LSST SIMS package MAF. The main goal of this step is to set DD and WFD to integer keys that
# label an observation as Deep Drilling or for Wide Fast Deep.
# If you want to skip this step, you can use the next cell by uncommenting it, and commenting out this cell, if all you
# care about is the database used in this example. But there is no guarantee that the numbers in the cell below will work
# on other versions of opsim database outputs
from lsst.sims.maf import db
from lsst.sims.maf.utils import opsimUtils
# +
# DD = 56
# WFD = 54
# -
# ## Read in OpSim output for modern versions: (sqlite formats)
# Here we will use the opsim output minion_1016
# I have downloaded this database, unzipped and use the variable dbname to point to its location
# Change dbname to point at your own location of the opsim output
dbname = '/Users/rbiswas/data/LSST/OpSimData/minion_1016_sqlite.db'
opsdb = db.OpsimDatabase(dbname)
propID, propTags = opsdb.fetchPropInfo()
DD = propTags['DD'][0]
WFD = propTags['WFD'][0]
print("The propID for the Deep Drilling Field {0:2d}".format(DD))
print("The propID for the Wide Fast Deep Field {0:2d}".format(WFD))
# ## Read in the OpSim DataBase into a pandas dataFrame
# Here we will read the opsim database into a `pandas.DataFrame`
engine = create_engine('sqlite:///' + dbname)
# The opsim database is a large file (approx 4.0 GB), but still possible to read into memory on new computers. You usually only need the Summary Table, which is about 900 MB. If you are only interested in the Deep Drilling Fields, you can use the read_sql_query to only select information pertaining to Deep Drilling Observations. This has a memory footprint of about 40 MB.
# Obviously, you can reduce this further by narrowing down the columns to those of interest only. For the entire Summary Table, this step takes a few minutes on my computer.
# If you are going to do the read from disk step very often, you can further reduce the time used by storing the output on disk as a hdf5 file and reading that into memory
# We will look at three different Summaries of OpSim Runs. A summary of the
# 1. Deep Drilling fields: These are the observations corresponding to propID of the variable DD above, and are restricted to a handful of fields
# 2. WFD (Main) Survey: These are the observations corresponding to the propID of the variables WFD
# 3. Combined Survey: These are observations combining DEEP and WFD in the DDF. Note that this leads to duplicate observations which must be subsequently dropped.
# +
# Load to a dataframe
# Summary = pd.read_hdf('storage.h5', 'table') # This loads the table from a hdf file which I store as intermediate result, this is extremely quick
# Summary = pd.read_sql_table('Summary', engine, index_col='obsHistID'), #'loads all of the summary table'
# EnigmaDeep = pd.read_sql_query('SELECT * FROM SUMMARY WHERE PROPID is 366', engine) # loads only the DDF
# -
# If we knew ahead of time the proposal ID, then we could have done this quicker using
OpSim_combined = pd.read_sql_query('SELECT * FROM SUMMARY WHERE PROPID in ({0}, {1})'.format(DD, WFD), engine, index_col='obsHistID')
OpSim_combined.head()
# This could have duplicates unlike in the case of the OpSim Deep. This is because there are now two proposal IDs both of which may correspond to the same observation. We can check that this is indeed the case by:
len(OpSim_combined) == len(OpSim_combined.index.unique())
# Dropping the duplicate pointings can be done in the following way. The reset_index() makes 'obsHistID' an ordinary column rather than an index, `drop_duplicates` drops duplicate rows where
OpSim_combined.reset_index().drop_duplicates(subset='obsHistID', inplace=True)
OpSim_combined.head()
len(OpSim_combined) == len(OpSim_combined.index.unique())
OpSim_Deep = pd.read_sql_query('SELECT * FROM SUMMARY WHERE PROPID is ' + str(DD), engine, index_col='obsHistID')
# We can also sub-select this from the all-encompassing Summay Table. This can be done in two way:
OpSim_combined.propID.unique()
OpSim_Deep.propID.unique()
# Get fieldID closest to ra, dec
fieldIDFromRADec = oss.fieldID(OpSim_combined, np.radians(190.), np.radians(-83.0))
print fieldIDFromRADec
# ## Some properties of the OpSim Outputs
# ### Construct our Summary
OpSimDeepSummary = so.SummaryOpsim(OpSim_Deep)
OpSimCombinedSummary = so.SummaryOpsim(OpSim_combined)
# #### Plot the location of deep fields
fig = plt.figure(figsize=(10, 5))
ax = fig.add_subplot(111, projection='mollweide');
fig = OpSimDeepSummary.showFields(ax=fig.axes[0], marker='o', s=40)
# ## How often do multiple DDF fields get observed ?
DDF_fieldIDs = OpSim_Deep.fieldID.unique()
DDF_fieldIDs
grouped = OpSim_Deep[['night', 'fieldID']].groupby(['night'])
fig, ax = plt.subplots()
grouped.agg({'fieldID': lambda x: x.unique().size}).hist(ax=ax)
fig.savefig('DeepOnly_uniqueDDFFields.png')
# What about if we count WFD visits to these fields ?
OpSimCombined_DDF_Fields = OpSim_combined.query('fieldID in @DDF_fieldIDs')
len(OpSimCombined_DDF_Fields)
OpSimCombined_DDF_Fields.fieldID.unique()
combinedGrouped = OpSimCombined_DDF_Fields.groupby('night')
fig, ax = plt.subplots()
combinedGrouped.agg({'fieldID': lambda x: x.unique().size}).hist(ax=ax)
fig.savefig('Combined_DDFvisits.png')
# #### Example plotting of a single field
fig = plt.figure(figsize=(10, 5))
ax = fig.add_subplot(111, projection='mollweide');
opsimFieldSummary = so.SummaryOpsim(OpSim_combined.query('fieldID==347'))
fig = opsimFieldSummary.showFields(ax=fig.axes[0], marker='o', s=40)
# ### Plot all fields
OpSimCombinedSummary.showFields(ax=ax, marker='o', color='r', s=8)
fieldIDFromRADec = oss.fieldID(OpSim_Deep, np.radians(53.), np.radians(-28.))
print fieldIDFromRADec
fieldIDFromRADec = oss.fieldID(OpSim_Deep, np.radians(0.), np.radians(-45.))
print fieldIDFromRADec
fieldIDFromRADec = oss.fieldID(OpSim_combined, np.radians(53.), np.radians(-28.))
print fieldIDFromRADec
fieldIDFromRADec = oss.fieldID(OpSim_combined, np.radians(85.7), np.radians(-14.4))
print fieldIDFromRADec
fieldList = OpSimCombinedSummary.fieldIds
# ## Obtain a list of filters, mjds, depths, and field ra, dec for deep fields
OpSim_combined.columns
DDF_fieldIDs
selectedFields = OpSim_combined.query('fieldID in @DDF_fieldIDs')[['expMJD', 'propID', 'filter', 'fieldRA', 'fieldDec', 'fieldID', 'fiveSigmaDepth']]#.head()
selectedFields.head()
selectedFields.to_hdf('DDF_Fields_Info.hdf', 'summaryInfo')
# CHECK
read_table = pd.read_hdf('DDF_Fields_Info.hdf', 'summaryInfo')
read_table.head()
from pandas.util.testing import assert_frame_equal
assert_frame_equal(read_table, selectedFields)
# #### First Season
# We can visualize the cadence during the first season using the cadence plot for a particular field: The following plot shows how many visits we have in different filters on a particular night:
firstSeasonDeep = OpSimDeepSummary.cadence_plot(fieldID=1427, observedOnly=False, sql_query='night < 366')
firstSeasonCombined = OpSimCombinedSummary.cadence_plot(fieldID=1427, observedOnly=False, sql_query='night < 366')
firstSeasonCombined[0].savefig('minion_1427.pdf')
firstSeason_main[0].savefig('minion_1430.pdf')
firstSeason = OpSimDeepSummary.cadence_plot(fieldID=744, observedOnly=False, sql_query='night < 732',
nightMin=0, nightMax=732)
tenCadence = OpSimCombinedSummary.cadence_plot(fieldID=fieldList[2000], observedOnly=False, sql_query='night < 3500', nightMax=3500)
# Suppose we have a supernova with a peak around a particular MJD of 49540, and we want to see what the observations happened around it:
SN = OpSimDeepSummary.cadence_plot(summarydf=OpSimDeepSummary.df, fieldID=1427, #racol='fieldRA', deccol='fieldDec',
observedOnly=False, mjd_center=59640., mjd_range=[-30., 50.])
# ax = plt.gca()
# ax.axvline(49540, color='r', lw=2.)
# ax.xaxis.get_major_formatter().set_useOffset(False)
SN[0].savefig('SN_observaton.pdf')
# # Scratch
SN_matrix.sum(axis=1).sum()
EnigmaDeep.query('fieldID == 744 and expMJD < 49590 and expMJD > 49510').expMJD.size
SN_matrix[SN_matrix > 0.5] = 1
SN_matrix.sum().sum()
len(SN_matrix.sum(axis=1).dropna())
nightlySN_matrix = SN_matrix.copy(deep=True)
nightlySN_matrix[SN_matrix > 0.5] =1
nightlySN_matrix.sum(axis=1).dropna().sum()
nightlySN_matrix.sum(axis=1).dropna().size
nightlySN_matrix.sum(ax)
|
example/ExploringOpSimOutputs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dynamic Programming - Fibonacci
# ### Fibonacci Number
#
# The Fibonacci numbers, commonly denoted F(n) form a sequence, called the Fibonacci sequence, such that each number is the sum of the two preceding ones, starting from 0 and 1. That is,
#
# ```
# F(0) = 0, F(1) = 1
# F(n) = F(n - 1) + F(n - 2), for n > 1.
# ```
#
# Source: https://leetcode.com/problems/fibonacci-number/
#
# Given n, calculate F(n).
#
# Example 1:
#
# ```
# Input: n = 2
# Output: 1
# Explanation: F(2) = F(1) + F(0) = 1 + 0 = 1.
# ```
#
# Example 2:
#
# ```
# Input: n = 3
# Output: 2
# Explanation: F(3) = F(2) + F(1) = 1 + 1 = 2.
# ```
#
# Example 3:
#
# ```
# Input: n = 4
# Output: 3
# Explanation: F(4) = F(3) + F(2) = 2 + 1 = 3.
# ```
class Solution:
def fib(self, n):
'''
'''
if n == 0:
return 0
if n == 1:
return 1
# DP: initializaiton
# (n+1) is needed
F = []
for i in range(n+1):
F.append(None)
F[0] = 0
F[1] = 1
# DP substructure
for i in range(2, n+1):
F[i] = F[i-1] + F[i-2]
return F[-1]
solver = Solution()
solver.fib(4)
# ### N-th Tribonacci Number
#
# The Tribonacci sequence Tn is defined as follows:
#
# T0 = 0, T1 = 1, T2 = 1, and Tn+3 = Tn + Tn+1 + Tn+2 for n >= 0.
#
# Given n, return the value of Tn.
#
# Source: https://leetcode.com/problems/n-th-tribonacci-number/
#
# Example 1:
#
# ```
# Input: n = 4
# Output: 4
# Explanation:
# T_3 = 0 + 1 + 1 = 2
# T_4 = 1 + 1 + 2 = 4
# ```
#
# Example 2:
#
# ```
# Input: n = 25
# Output: 1389537
# ```
class Solution:
def tribonacci(self, n):
'''
'''
if n == 0:
return 0
if n == 1:
return 1
if n == 2:
return 1
# DP: initialization
T = []
for i in range(n+1):
T.append(None)
T[0] = 0
T[1] = 1
T[2] = 1
# DP": substructure
for i in range(3, n+1):
T[i] = T[i-3] + T[i-2] + T[i-1]
return T[-1]
solver = Solution()
solver.tribonacci(25)
# ### Climbing Stairs (frog jump)
# You are climbing a staircase. It takes n steps to reach the top.
#
# Each time you can either climb 1 or 2 steps. In how many distinct ways can you climb to the top?
#
# Example 1:
#
# ```
# Input: n = 2
# Output: 2
# Explanation: There are two ways to climb to the top.
# 1. 1 step + 1 step
# 2. 2 steps
# ```
#
# Example 2:
#
# ```
# Input: n = 3
# Output: 3
# Explanation: There are three ways to climb to the top.
# 1. 1 step + 1 step + 1 step
# 2. 1 step + 2 steps
# 3. 2 steps + 1 step
# ```
# +
class Solution:
def climbStairs(self, n):
'''
'''
if n == 1:
return 1
if n == 2:
return 2
# DP: initialization
step = []
for i in range(n):
step.append(None)
step[0] = 1
step[1] = 2
# DP: substructure
for i in range(2, n):
step[i] = step[i-1] + step[i-2]
return step[-1]
# -
solver = Solution()
solver.climbStairs(3)
# ### Min Cost Climbing Stairs (frog jump with cost)
#
# You are given an integer array cost where cost[i] is the cost of ith step on a staircase. Once you pay the cost, you can either climb one or two steps.
#
# You can either start from the step with index 0, or the step with index 1.
#
# Return the minimum cost to reach the top of the floor.
#
# Example 1:
#
# ```
# Input: cost = [10,15,20]
# Output: 15
# Explanation: You will start at index 1.
# - Pay 15 and climb two steps to reach the top.
# The total cost is 15.
# ```
#
# Example 2:
#
# ```
# Input: cost = [1,100,1,1,1,100,1,1,100,1]
# Output: 6
# Explanation: You will start at index 0.
# - Pay 1 and climb two steps to reach index 2.
# - Pay 1 and climb two steps to reach index 4.
# - Pay 1 and climb two steps to reach index 6.
# - Pay 1 and climb one step to reach index 7.
# - Pay 1 and climb two steps to reach index 9.
# - Pay 1 and climb one step to reach the top.
# The total cost is 6.
# ```
class Solution:
def minCostClimbingStairs(self, cost):
'''
'''
L = len(cost)
# DP: initialization
payment = []
for i in range(L+1):
payment.append(None)
payment[0] = 0
payment[1] = cost[0]
# DP: substructure
# finish with 2-step jump v.s. 1-step jump
cost = [0,] + cost
for i in range(2, L+1):
payment[i] = min(payment[i-2]+cost[i-2], payment[i-1]+cost[i-1])
return payment[-1]
solver = Solution()
solver.minCostClimbingStairs([1,100,1,1,1,100,1,1,100,1])
# ### House Robber
#
# You are a professional robber planning to rob houses along a street. Each house has a certain amount of money stashed, the only constraint stopping you from robbing each of them is that adjacent houses have security systems connected and it will automatically contact the police if two adjacent houses were broken into on the same night.
#
# Given an integer array nums representing the amount of money of each house, return the maximum amount of money you can rob tonight without alerting the police.
#
# Source: https://leetcode.com/problems/house-robber/
#
# Example 1:
#
# ```
# Input: nums = [1,2,3,1]
# Output: 4
# Explanation: Rob house 1 (money = 1) and then rob house 3 (money = 3).
# Total amount you can rob = 1 + 3 = 4.
# ```
#
# Example 2:
#
# ```
# Input: nums = [2,7,9,3,1]
# Output: 12
# Explanation: Rob house 1 (money = 2), rob house 3 (money = 9) and rob house 5 (money = 1).
# Total amount you can rob = 2 + 9 + 1 = 12.
# ```
class Solution:
def rob(self, nums):
'''
'''
L = len(nums)
if L == 1:
return nums[0]
rob_gain = []
for i in range(L+1):
rob_gain.append(None)
rob_gain[0] = 0
rob_gain[1] = nums[0]
nums = [0,]+nums
for i in range(2, L+1):
rob_gain[i] = max(rob_gain[i-2] + nums[i], rob_gain[i-1])
return rob_gain[-1]
solver = Solution()
solver.rob([1,2,3,1])
# ### House Robber II
#
# You are a professional robber planning to rob houses along a street. Each house has a certain amount of money stashed. All houses at this place are arranged in a circle. That means the first house is the neighbor of the last one. Meanwhile, adjacent houses have a security system connected, and it will automatically contact the police if two adjacent houses were broken into on the same night.
#
# Given an integer array nums representing the amount of money of each house, return the maximum amount of money you can rob tonight without alerting the police.
#
# Source: https://leetcode.com/problems/house-robber-ii/
#
# Example 1:
#
# ```
# Input: nums = [2,3,2]
# Output: 3
# Explanation: You cannot rob house 1 (money = 2) and then rob house 3 (money = 2), because they are adjacent houses.
# ```
#
# Example 2:
#
# ```
# Input: nums = [1,2,3,1]
# Output: 4
# Explanation: Rob house 1 (money = 1) and then rob house 3 (money = 3).
# Total amount you can rob = 1 + 3 = 4.
# ```
#
# Example 3:
#
# ```
# Input: nums = [1,2,3]
# Output: 3
# ```
class Solution:
def rob(self, nums):
'''
'''
# It is similar to that of the previous rob problem, but with two optimizations,
# one excludes the first house, the other exclude the last house
L = len(nums)
if L == 1:
return nums[0]
rob_head = []
rob_tail = []
for i in range(L):
rob_head.append(None)
rob_tail.append(None)
nums_head = nums[:-1].copy() # if head, dont rob tail
nums_head = [0,] + nums_head
rob_head[0] = 0
rob_head[1] = nums_head[1]
nums_tail = nums[1:].copy() # if tail, dont rob head
nums_tail = [0,] + nums_tail
rob_tail[0] = 0
rob_tail[1] = nums_tail[1]
for i in range(2, L):
rob_head[i] = max(rob_head[i-2]+nums_head[i], rob_head[i-1])
rob_tail[i] = max(rob_tail[i-2]+nums_tail[i], rob_tail[i-1])
return max(rob_head[-1], rob_tail[-1])
solver = Solution()
solver.rob([2,3,2])
solver.rob([1,2,3,1])
solver.rob([1,2,3])
# **What I have learned**
#
# * When confused by the DP substructure, take the "finishing step" as an example and generalize it by using "`i`, `i-1`, `i-2`".
|
Python/Dynamic_Programming_Fibonacci.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ``solarposition.py`` tutorial
#
# This tutorial needs your help to make it better!
#
# Table of contents:
# 1. [Setup](#Setup)
# 2. [SPA output](#SPA-output)
# 2. [Speed tests](#Speed-tests)
#
# This tutorial has been tested against the following package versions:
# * pvlib 0.3.0
# * Python 3.5.1
# * IPython 4.1
# * Pandas 0.18.0
#
# It should work with other Python and Pandas versions. It requires pvlib > 0.3.0 and IPython > 3.0.
#
# Authors:
# * <NAME> (@wholmgren), University of Arizona. July 2014, July 2015, March 2016
# ## Setup
# +
import datetime
# scientific python add-ons
import numpy as np
import pandas as pd
# plotting stuff
# first line makes the plots appear in the notebook
# %matplotlib inline
import matplotlib.pyplot as plt
# finally, we import the pvlib library
import pvlib
# -
import pvlib
from pvlib.location import Location
# ## SPA output
tus = Location(32.2, -111, 'US/Arizona', 700, 'Tucson')
print(tus)
golden = Location(39.742476, -105.1786, 'America/Denver', 1830, 'Golden')
print(golden)
golden_mst = Location(39.742476, -105.1786, 'MST', 1830, 'Golden MST')
print(golden_mst)
berlin = Location(52.5167, 13.3833, 'Europe/Berlin', 34, 'Berlin')
print(berlin)
times = pd.date_range(start=datetime.datetime(2014,6,23), end=datetime.datetime(2014,6,24), freq='1Min')
times_loc = times.tz_localize(tus.pytz)
times
# +
pyephemout = pvlib.solarposition.pyephem(times_loc, tus.latitude, tus.longitude)
spaout = pvlib.solarposition.spa_python(times_loc, tus.latitude, tus.longitude)
pyephemout['elevation'].plot(label='pyephem')
pyephemout['apparent_elevation'].plot(label='pyephem apparent')
spaout['elevation'].plot(label='spa')
plt.legend(ncol=2)
plt.title('elevation')
print('pyephem')
print(pyephemout.head())
print('spa')
print(spaout.head())
# +
plt.figure()
pyephemout['elevation'].plot(label='pyephem')
spaout['elevation'].plot(label='spa')
(pyephemout['elevation'] - spaout['elevation']).plot(label='diff')
plt.legend(ncol=3)
plt.title('elevation')
plt.figure()
pyephemout['apparent_elevation'].plot(label='pyephem apparent')
spaout['elevation'].plot(label='spa')
(pyephemout['apparent_elevation'] - spaout['elevation']).plot(label='diff')
plt.legend(ncol=3)
plt.title('elevation')
plt.figure()
pyephemout['apparent_zenith'].plot(label='pyephem apparent')
spaout['zenith'].plot(label='spa')
(pyephemout['apparent_zenith'] - spaout['zenith']).plot(label='diff')
plt.legend(ncol=3)
plt.title('zenith')
plt.figure()
pyephemout['apparent_azimuth'].plot(label='pyephem apparent')
spaout['azimuth'].plot(label='spa')
(pyephemout['apparent_azimuth'] - spaout['azimuth']).plot(label='diff')
plt.legend(ncol=3)
plt.title('azimuth')
# +
pyephemout = pvlib.solarposition.pyephem(times.tz_localize(golden.tz), golden.latitude, golden.longitude)
spaout = pvlib.solarposition.spa_python(times.tz_localize(golden.tz), golden.latitude, golden.longitude)
pyephemout['elevation'].plot(label='pyephem')
pyephemout['apparent_elevation'].plot(label='pyephem apparent')
spaout['elevation'].plot(label='spa')
plt.legend(ncol=2)
plt.title('elevation')
print('pyephem')
print(pyephemout.head())
print('spa')
print(spaout.head())
# +
pyephemout = pvlib.solarposition.pyephem(times.tz_localize(golden.tz), golden.latitude, golden.longitude)
ephemout = pvlib.solarposition.ephemeris(times.tz_localize(golden.tz), golden.latitude, golden.longitude)
pyephemout['elevation'].plot(label='pyephem')
pyephemout['apparent_elevation'].plot(label='pyephem apparent')
ephemout['elevation'].plot(label='ephem')
plt.legend(ncol=2)
plt.title('elevation')
print('pyephem')
print(pyephemout.head())
print('ephem')
print(ephemout.head())
# +
loc = berlin
pyephemout = pvlib.solarposition.pyephem(times.tz_localize(loc.tz), loc.latitude, loc.longitude)
ephemout = pvlib.solarposition.ephemeris(times.tz_localize(loc.tz), loc.latitude, loc.longitude)
pyephemout['elevation'].plot(label='pyephem')
pyephemout['apparent_elevation'].plot(label='pyephem apparent')
ephemout['elevation'].plot(label='ephem')
ephemout['apparent_elevation'].plot(label='ephem apparent')
plt.legend(ncol=2)
plt.title('elevation')
print('pyephem')
print(pyephemout.head())
print('ephem')
print(ephemout.head())
# +
loc = berlin
times = pd.DatetimeIndex(start=datetime.date(2015,3,28), end=datetime.date(2015,3,29), freq='5min')
pyephemout = pvlib.solarposition.pyephem(times.tz_localize(loc.tz), loc.latitude, loc.longitude)
ephemout = pvlib.solarposition.ephemeris(times.tz_localize(loc.tz), loc.latitude, loc.longitude)
pyephemout['elevation'].plot(label='pyephem')
pyephemout['apparent_elevation'].plot(label='pyephem apparent')
ephemout['elevation'].plot(label='ephem')
plt.legend(ncol=2)
plt.title('elevation')
plt.figure()
pyephemout['azimuth'].plot(label='pyephem')
ephemout['azimuth'].plot(label='ephem')
plt.legend(ncol=2)
plt.title('azimuth')
print('pyephem')
print(pyephemout.head())
print('ephem')
print(ephemout.head())
# +
loc = berlin
times = pd.DatetimeIndex(start=datetime.date(2015,3,30), end=datetime.date(2015,3,31), freq='5min')
pyephemout = pvlib.solarposition.pyephem(times.tz_localize(loc.tz), loc.latitude, loc.longitude)
ephemout = pvlib.solarposition.ephemeris(times.tz_localize(loc.tz), loc.latitude, loc.longitude)
pyephemout['elevation'].plot(label='pyephem')
pyephemout['apparent_elevation'].plot(label='pyephem apparent')
ephemout['elevation'].plot(label='ephem')
plt.legend(ncol=2)
plt.title('elevation')
plt.figure()
pyephemout['azimuth'].plot(label='pyephem')
ephemout['azimuth'].plot(label='ephem')
plt.legend(ncol=2)
plt.title('azimuth')
print('pyephem')
print(pyephemout.head())
print('ephem')
print(ephemout.head())
# +
loc = berlin
times = pd.DatetimeIndex(start=datetime.date(2015,6,28), end=datetime.date(2015,6,29), freq='5min')
pyephemout = pvlib.solarposition.pyephem(times.tz_localize(loc.tz), loc.latitude, loc.longitude)
ephemout = pvlib.solarposition.ephemeris(times.tz_localize(loc.tz), loc.latitude, loc.longitude)
pyephemout['elevation'].plot(label='pyephem')
pyephemout['apparent_elevation'].plot(label='pyephem apparent')
ephemout['elevation'].plot(label='ephem')
plt.legend(ncol=2)
plt.title('elevation')
plt.figure()
pyephemout['azimuth'].plot(label='pyephem')
ephemout['azimuth'].plot(label='ephem')
plt.legend(ncol=2)
plt.title('azimuth')
print('pyephem')
print(pyephemout.head())
print('ephem')
print(ephemout.head())
# -
pyephemout['elevation'].plot(label='pyephem')
pyephemout['apparent_elevation'].plot(label='pyephem apparent')
ephemout['elevation'].plot(label='ephem')
ephemout['apparent_elevation'].plot(label='ephem apparent')
plt.legend(ncol=2)
plt.title('elevation')
plt.xlim(pd.Timestamp('2015-06-28 02:00:00+02:00'), pd.Timestamp('2015-06-28 06:00:00+02:00'))
plt.ylim(-10,10)
# ## Speed tests
times_loc = times.tz_localize(loc.tz)
# +
# %%timeit
pyephemout = pvlib.solarposition.pyephem(times_loc, loc.latitude, loc.longitude)
#ephemout = pvlib.solarposition.ephemeris(times, loc)
# +
# %%timeit
#pyephemout = pvlib.solarposition.pyephem(times, loc)
ephemout = pvlib.solarposition.ephemeris(times_loc, loc.latitude, loc.longitude)
# +
# %%timeit
#pyephemout = pvlib.solarposition.pyephem(times, loc)
ephemout = pvlib.solarposition.get_solarposition(times_loc, loc.latitude, loc.longitude,
method='nrel_numpy')
# -
# This numba test will only work properly if you have installed numba.
# +
# %%timeit
#pyephemout = pvlib.solarposition.pyephem(times, loc)
ephemout = pvlib.solarposition.get_solarposition(times_loc, loc.latitude, loc.longitude,
method='nrel_numba')
# -
# The numba calculation takes a long time the first time that it's run because it uses LLVM to compile the Python code to machine code. After that it's about 4-10 times faster depending on your machine. You can pass a ``numthreads`` argument to this function. The optimum ``numthreads`` depends on your machine and is equal to 4 by default.
# +
# %%timeit
#pyephemout = pvlib.solarposition.pyephem(times, loc)
ephemout = pvlib.solarposition.get_solarposition(times_loc, loc.latitude, loc.longitude,
method='nrel_numba', numthreads=16)
# +
# %%timeit
ephemout = pvlib.solarposition.spa_python(times_loc, loc.latitude, loc.longitude,
how='numba', numthreads=16)
# -
|
docs/tutorials/solarposition.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gendered perspectives on character.
#
import csv, math
import pandas as pd
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
import statsmodels.api as sm
from adjustText import adjust_text
# %matplotlib inline
data = pd.read_csv('chartable.tsv', sep = '\t')
lexicon = pd.read_csv('lexicon.tsv', sep = '\t')
vocab = set(lexicon.word[0:1800])
def dunnings(vectora, vectorb):
''' Calculates a version of Dunning's log-likelihood,
differing from generic Dunning's in two ways: first,
we take an *additional* log (the log of log-likelihood),
in order to highlight subtle differences near the origin.
Second, we give a sign to the metric in order to identify
the direction of overrepresentation.
NOTE: the decision to take an additional log is an interpretive
choice made *explicitly in order to highlight* a specific aspect of
the results.
'''
assert len(vectora) == len(vectorb)
veclen = len(vectora)
totala = np.sum(vectora)
totalb = np.sum(vectorb)
totalboth = totala + totalb
dunningvector = np.zeros(veclen)
for i in range(veclen):
if vectora[i] == 0 or vectorb[i] == 0:
continue
# Cause you know you're going to get div0 errors.
try:
probI = (vectora[i] + vectorb[i]) / totalboth
probnotI = 1 - probI
expectedIA = totala * probI
expectedIB = totalb * probI
expectedNotIA = totala * probnotI
expectedNotIB = totalb * probnotI
expected_table = np.array([[expectedIA, expectedNotIA],
[expectedIB, expectedNotIB]])
actual_table = np.array([[vectora[i], (totala - vectora[i])],
[vectorb[i], (totalb - vectorb[i])]])
G = np.sum(actual_table * np.log(actual_table / expected_table))
# take an additional log to make it visualizable
# NOTE: this step makes a big difference; it is not present in
# generic Dunnings.
G = math.log(G)
# We're going to use a signed version of Dunnings, so features where
# B is higher than expected will be negative.
if expectedIB > vectorb[i]:
G = -G
dunningvector[i] = G
except:
pass
# There are a million ways to get a div-by-zero or log-zero error
# in that calculation. I could check them all, or just do this.
# The vector was initialized with zeroes, which are the default
# value I want for failed calculations anyhow.
return dunningvector
data.head()
# +
# I'm simply going to fix some grammatical issues, for cosmetic reasons
# If you prefer not to, you can delete this. It makes little substantive
# difference.
themap = {'was-seeing': 'was-seen', 'was-saw': 'was-seen', 'was-see': 'was-seen',
'was-tell': 'was-told', 'was-marry': 'was-married', 'was-love': 'was-loved',
'was-ask': 'was-asked'}
def fixgrammar(aword):
if aword in themap:
return themap[aword]
else:
return aword
data['word'] = data['word'].apply(fixgrammar)
vocab = vocab.intersection(set(data.word))
# +
def sumwords(df):
grouped = df.groupby('word')
wordsums = grouped.aggregate(np.sum)
return wordsums.wordct
def sumchars(df):
grouped = df.groupby('word')
wordsums = grouped.aggregate(np.sum)
return wordsums.charct
def dunningsforauthgender(data, authgender):
perspectiveonmen = data[(data.authgender == authgender) & (data.chargender == 'm')]
perspectiveonwomen = data[(data.authgender == authgender) & (data.chargender == 'f')]
mascwords = sumwords(perspectiveonmen).rename('m')
femwords = sumwords(perspectiveonwomen).rename('f')
comparison = pd.concat([femwords, mascwords], axis = 1)
comparison.fillna(0, inplace = True)
gvals = dunnings(comparison.m, comparison.f)
comparison = comparison.assign(g = gvals)
comparison.sort_values(by = 'g', inplace = True)
return comparison
def duncharsforauthgender(data, authgender):
perspectiveonmen = data[(data.authgender == authgender) & (data.chargender == 'm')]
perspectiveonwomen = data[(data.authgender == authgender) & (data.chargender == 'f')]
mascwords = sumchars(perspectiveonmen).rename('m')
femwords = sumchars(perspectiveonwomen).rename('f')
comparison = pd.concat([femwords, mascwords], axis = 1)
comparison.fillna(0, inplace = True)
gvals = dunnings(comparison.m, comparison.f)
comparison = comparison.assign(g = gvals)
comparison.sort_values(by = 'g', inplace = True)
return comparison
f_perspective = dunningsforauthgender(data, 'f')
m_perspective = dunningsforauthgender(data, 'm')
gforf = f_perspective.g.rename('f')
gform = m_perspective.g.rename('m')
twoperspectives = pd.concat([gforf, gform], axis = 1)
twoperspectives.fillna(0, inplace = True)
twoperspectives['word'] = twoperspectives.index
twoperspectives.head()
# -
def plot_gender_means(words, frametoplot, adjust=True, title=False, label_outliers=True, normalize_axes=False, save=False):
df = frametoplot.loc[(frametoplot.word.isin(words)),:]
ax0 = df.plot('m', 'f', kind='scatter', s=50, figsize=(15,15))
ax0.plot(0, 0, 'r+', ms=15, mew=2, label='(0,0) Gender Neutral')
texts = []
for x, y, s in zip(df['m'], df['f'], df['word']):
if label_outliers:
if (((x < - 4.3) & (y > 4.3)) |((x > 4.3) & (y < -4.3))):
texts.append(plt.text(x, y, s, size = 11))
if (((x < -8) & (y < -8)) |((x > 8.5) & (y > 8.5))):
texts.append(plt.text(x, y, s, size=11))
if adjust:
adjust_text(texts, arrowprops=dict(arrowstyle="-", color='k', lw=0.5))
plt.legend(loc='best')
plt.ylabel('m <-- Diff for women writers 1800-2000 --> f', fontsize=16, )
plt.xlabel('m <-- Diff for male writers 1800-2000 --> f', fontsize=16)
if title:
plt.title(title)
if normalize_axes:
ylim = ax0.get_ylim()
xlim = ax0.get_xlim()
new_low = min(xlim[0], ylim[0])
new_hi = max(xlim[1], ylim[1])
plt.ylim(new_low, new_hi)
plt.xlim(new_low, new_hi)
if save:
if title:
plt.savefig('./{}.png'.format(title))
else:
plt.savefig('./genderDiffMeans.png')
plt.show()
plot_gender_means(vocab, twoperspectives)
# +
# Let's write that to disk for R visualization
outframe = twoperspectives.loc[(twoperspectives.word.isin(vocab)), : ]
things2plot = {'said', 'was-married', 'was-told', 'seemed',
'tone', 'expression', 'said-poetry', 'kisses', 'kissed',
'was-seen', 'words', 'meant', 'wife', 'sword', 'grinned',
'pipe', 'pocket', 'said-sir', 'beard', 'pipe', 'horse',
'said-oh', 'hair', 'breasts', 'aunt', 'was-loved',
'husband', 'dress', 'mother', 'eyes', 'ear', 'feet', 'head', 'mind',
'heard', 'hungry', 'mind', 'remembered',
'saw', 'stomach', 'thought', 'throat', 'wondered'}
def map2label(word):
global things2plot
if word in things2plot:
return word
else:
return ''
outframe = outframe.assign(label = outframe.word.apply(map2label))
outframe.to_csv('data4r.csv', index = False)
# -
obj = twoperspectives[(twoperspectives.m < -3) & (twoperspectives.f > 3)]
obj
# +
twentieth = data[data.year > 1950]
f_perspective = dunningsforauthgender(twentieth, 'f')
m_perspective = dunningsforauthgender(twentieth, 'm')
gforf = f_perspective.g.rename('f')
gform = m_perspective.g.rename('m')
perspective20 = pd.concat([gforf, gform], axis = 1)
perspective20.fillna(0, inplace = True)
perspective20['word'] = perspective20.index
plot_gender_means(vocab, perspective20)
# +
nineteenth = data[data.year < 1900]
f_perspective = dunningsforauthgender(nineteenth, 'f')
m_perspective = dunningsforauthgender(nineteenth, 'm')
gforf = f_perspective.g.rename('f')
gform = m_perspective.g.rename('m')
perspective19 = pd.concat([gforf, gform], axis = 1)
perspective19.fillna(0, inplace = True)
perspective19['word'] = perspective19.index
plot_gender_means(vocab, perspective19)
# -
|
genderedperspectives/gendered_perspectives.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import required packages
from fastai2.tabular.all import *
from helpers import *
# path to project directory
path = Path('./')
# read in training dataset
train_df = pd.read_csv(path/'data/train_v4.csv', index_col=0, dtype={'season':str})
# ## Neural network model
#
# Fastai have an neural network architecture that can be applied to tabular data, including for time series. In practice this creates embeddings for categorical features, combines them with continuous variables and passes this through a number of fully connected layers to produce a prediction for the dependent variable. They also include a number of tricks and default parameters that should produce decent performance.
#
# Once again we'll first create some lag features.
# add a bunch of player lag features
lag_train_df, player_lag_vars = player_lag_features(train_df, ['total_points'], ['all', 10])#, 1, 2, 3, 4, 5, 10])
# And go ahead and create our training set.
#
# The gameweeks and seasons are ordered, so we want to have them as ordered categories with the correct order (2016-17, 2017-18, etc.). And for this model we need to make the dependent variable a float.
# set validaton point/length and categorical/continuous variables
valid_season = '1920'
valid_gw = 20
valid_len = 6
cat_vars = ['gw', 'season', 'position', 'team', 'opponent_team', 'was_home']
cont_vars = ['minutes']
dep_var = ['total_points']
# we want to set gw and season as ordered categorical variables
# need lists with ordered categories
ordered_gws = list(range(1,39))
ordered_seasons = ['1617', '1718', '1819', '1920']
# +
# set as categories with correct order
lag_train_df['gw'] = lag_train_df['gw'].astype('category')
lag_train_df['season'] = lag_train_df['season'].astype('category')
lag_train_df['gw'].cat.set_categories(ordered_gws, ordered=True, inplace=True)
lag_train_df['season'].cat.set_categories(ordered_seasons, ordered=True, inplace=True)
# -
# dependent variable needs to be float
lag_train_df['total_points'] = lag_train_df['total_points'].astype('float64')
# +
# create dataset with adjusted post-validation lag numbers
lag_train_df, train_idx, valid_idx = create_lag_train(lag_train_df,
cat_vars, cont_vars, player_lag_vars, dep_var,
valid_season, valid_gw, valid_len)
lag_train_df[player_lag_vars] = lag_train_df[player_lag_vars].fillna(0)
# -
# take a look at the dataframe
lag_train_df.info()
# create train/valid splits
splits = (list(train_idx), list(valid_idx))
valid_idx
lag_train_df[cat_vars + ['player']].nunique()
cat_vars + ['player']
cont_vars + player_lag_vars
# processors - categorify categorical variables and normalize continuous variables
# fill missing not used because new teams are almost certainly well below the league average
procs=[Categorify, Normalize]
to_nn = TabularPandas(lag_train_df, procs, cat_vars + ['player'], cont_vars + player_lag_vars,
splits=splits, y_names=dep_var)
# dls = to_nn.dataloaders()
dls = to_nn.dataloaders()
dls.show_batch()
# set range of predictions - minimum to current max
max_y = np.max(lag_train_df['total_points'])
min_y = np.min(lag_train_df['total_points'])
y_range = (-1, max_y)
learn = tabular_learner(dls, y_range=y_range, layers=[1000,500], ps=[0.001,0.01], emb_drop=0.04,
n_out=1, loss_func=F.mse_loss, metrics=rmse)
learn.lr_find()
# learn.fit_one_cycle(8, 1e-2, wd=0.2)
learn.fit_one_cycle(8, 1.2e-3, wd=0.2)
preds,targs = learn.get_preds()
r_mse(preds,targs)
# find validation cut point - index for 19-20 season second half (gw 19)
valid_start = train_df[(train_df['gw'] > 18) & (train_df['season'] == '1920')].index.min()
valid_end = train_df.index.max()
# +
# set validation dataset based on indexes
train_idx = range(valid_start)
valid_idx = range(valid_start, valid_end)
splits = (list(train_idx), list(valid_idx))
valid_idx
# -
# set categorical and continuous variables for tabular learner
cat_vars = ['player', 'gw', 'position', 'team', 'opponent_team', 'was_home', 'season']
cont_vars = ['minutes', 'relative_market_value_team', 'relative_market_value_opponent_team']
# set dependent variable
dep_var = 'total_points'
train_df[cat_vars].nunique()
# processors - categorify categorical variables and normalize continuous variables
# fill missing not used because new teams are almost certainly well below the league average
procs=[Categorify, Normalize]
to_nn = TabularPandas(train_df, procs, cat_vars, cont_vars,
splits=splits, y_names=dep_var)
# dls = to_nn.dataloaders()
dls = to_nn.dataloaders()
dls.show_batch()
# set range of predictions - minimum to current max
max_y = np.max(train_df['total_points'])
min_y = np.min(train_df['total_points'])
y_range = (-1, max_y)
learn = tabular_learner(dls, y_range=y_range, layers=[1000,500], ps=[0.001,0.01], emb_drop=0.04,
n_out=1, loss_func=F.mse_loss, metrics=rmse)
learn.lr_find()
learn.fit_one_cycle(5, 1e-2, wd=0.2)
def r_mse(pred,y): return round(math.sqrt(((pred-y)**2).mean()), 6)
preds,targs = learn.get_preds()
r_mse(preds,targs)
preds
targs
|
.ipynb_checkpoints/04_fpl_predict_fastai2_tabular-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plotting and Functions
# This notebook will work trough how to plot data and how to define functions. Throughout the lecture we will take a few moments to plot different functions and see how they depend on their parameters
# ## Plotting in Python: Matplot
import matplotlib.pyplot as plt
import numpy as np
import scipy as sp
# Pyplot is a powerful plotting library that can be used to make publication quaility plots. It is also useful for quikly plotting the results of a calcualtion.
#
# This is a quick demonstration of its use
#
# Note: when you call al library `import matplotlib.pyplot as plt` the way that use it is to do the following `plt.function()` where `function()` is whatever you are trying to call from the library
# Define x and y values for some function
x = [i for i in range(20)]
y1 = [i**2 for i in x]
y2 = [i**3 for i in x]
# The methods used above to make the lists is considered very *pythonic*. It works the same as a loop, but outputs all the results into a list. The left-hand most argument is what the list elements will be and the right hand side is the the way the loop will work.
# When you use pyplot to make a plot, you can add more than one data set to the figure until you render the plot. Once you render the plot it resets
plt.plot(x,y1)
plt.plot(x,y2)
plt.xlabel('X', fontsize=24)
plt.ylabel('Y', fontsize=24)
plt.legend(['Quadratic', 'Cubic'], loc=0)
plt.show()
# We can call also use numpy fucntions to make our plots. Numpy is a very powerful math library
# linspace will make a list of values from initial to final with however many increments you want
# this example goes from 0-2.5 with 20 increments
x=np.linspace(0,1.0,20)
print(x)
exp_func=np.exp(-2*np.pi*x)
print(exp_func)
plt.plot(x,exp_func, color="black")
plt.xlabel('x', fontsize=24)
plt.ylabel("y(x)", fontsize=24)
plt.show()
# All aspects of the plot can be changed. The best way to figure out what you want to do is to go to the Matplotlib gallery and choose an image that looks like what you are trying to do.
#
# https://matplotlib.org/gallery/index.html
# ### Example: Scatter plot with histograms
# +
import numpy as np
#Fixing random state for reproducibility
np.random.seed(19680801)
# the random data
x = np.random.randn(1000)
y = np.random.randn(1000)
# definitions for the axes
left, width = 0.1, 0.65
bottom, height = 0.1, 0.65
spacing = 0.005
rect_scatter = [left, bottom, width, height]
rect_histx = [left, bottom + height + spacing, width, 0.2]
rect_histy = [left + width + spacing, bottom, 0.2, height]
# start with a rectangular Figure
plt.figure(figsize=(8, 8))
ax_scatter = plt.axes(rect_scatter)
ax_scatter.tick_params(direction='in', top=True, right=True)
ax_histx = plt.axes(rect_histx)
ax_histx.tick_params(direction='in', labelbottom=False)
ax_histy = plt.axes(rect_histy)
ax_histy.tick_params(direction='in', labelleft=False)
# the scatter plot:
ax_scatter.scatter(x, y)
# now determine nice limits by hand:
binwidth = 0.25
lim = np.ceil(np.abs([x, y]).max() / binwidth) * binwidth
ax_scatter.set_xlim((-lim, lim))
ax_scatter.set_ylim((-lim, lim))
bins = np.arange(-lim, lim + binwidth, binwidth)
ax_histx.hist(x, bins=bins)
ax_histy.hist(y, bins=bins, orientation='horizontal')
ax_histx.set_xlim(ax_scatter.get_xlim())
ax_histy.set_ylim(ax_scatter.get_ylim())
plt.show()
# -
# I don't have to be an expert in making that kind of plot. I just have to understand and guess enough to figure out. I also google things I don't know
#
# https://www.google.com/search?client=firefox-b-1-d&q=pyplot+histogram+change+color
#
# https://stackoverflow.com/questions/42172440/python-matplotlib-histogram-color?rq=1
#
# https://matplotlib.org/examples/color/named_colors.html
#
# Then I can make small changes to have the plot look how I want it to look
#
# Notice below I changed
#
# `ax_scatter.scatter(x, y, color="purple")`,
#
# `ax_histx.hist(x, bins=bins, color = "skyblue")`,
#
# `ax_histy.hist(y, bins=bins, orientation='horizontal', color="salmon")`
# +
#Fixing random state for reproducibility
np.random.seed(19680801)
# the random data
x = np.random.randn(1000)
y = np.random.randn(1000)
# definitions for the axes
left, width = 0.1, 0.65
bottom, height = 0.1, 0.65
spacing = 0.005
rect_scatter = [left, bottom, width, height]
rect_histx = [left, bottom + height + spacing, width, 0.2]
rect_histy = [left + width + spacing, bottom, 0.2, height]
# start with a rectangular Figure
plt.figure(figsize=(8, 8))
ax_scatter = plt.axes(rect_scatter)
ax_scatter.tick_params(direction='in', top=True, right=True)
ax_histx = plt.axes(rect_histx)
ax_histx.tick_params(direction='in', labelbottom=False)
ax_histy = plt.axes(rect_histy)
ax_histy.tick_params(direction='in', labelleft=False)
# the scatter plot:
ax_scatter.scatter(x, y, color="purple")
# now determine nice limits by hand:
binwidth = 0.25
lim = np.ceil(np.abs([x, y]).max() / binwidth) * binwidth
ax_scatter.set_xlim((-lim, lim))
ax_scatter.set_ylim((-lim, lim))
bins = np.arange(-lim, lim + binwidth, binwidth)
ax_histx.hist(x, bins=bins, color = "skyblue")
ax_histy.hist(y, bins=bins, orientation='horizontal', color="salmon")
ax_histx.set_xlim(ax_scatter.get_xlim())
ax_histy.set_ylim(ax_scatter.get_ylim())
plt.show()
# -
# Notice how I changed the colors on the plot based off of what I found on the stack exchange. The way to solve issues in the course and computational work is to google them.
# ## Plotting Exersice 1
# Find a plot from the gallery that you like. Then make some sort of noticable change to it.
# +
import matplotlib.pyplot as plt
import numpy as np
# unit area ellipse
rx, ry = 3., 1.
area = rx * ry * np.pi
theta = np.arange(0, 2 * np.pi + 0.01, 0.1)
verts = np.column_stack([rx / area * np.cos(theta), ry / area * np.sin(theta)])
x, y, s, c = np.random.rand(4, 30)
s *= 10**2.
fig, ax = plt.subplots()
ax.scatter(x, y, s, c, marker=verts)
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
# unit area ellipse
rx, ry = 5., 2.
area = rx * ry * np.pi
theta = np.arange(4, 5 * np.pi + 0.01, 0.1)
verts = np.column_stack([rx / area * np.cos(theta), ry / area * np.sin(theta)])
x, y, s, c = np.random.rand(4, 60)
s *= 10**4.
fig, ax = plt.subplots()
ax.scatter(x, y, s, c, marker=verts)
plt.show()
# -
# ## Plotting Exersice 2
# Plot a the following functions on the same plot from $ -2\pi $ to $2\pi$
#
# $$ \sin(2\pi x+\pi)$$
# $$ \cos(2\pi x+\pi)$$
# $$\sin(2\pi x+\pi)+\cos(2\pi x+\pi)$$
# This might be useful:
# https://docs.scipy.org/doc/numpy/reference/generated/numpy.sin.html
# https://docs.scipy.org/doc/numpy/reference/generated/numpy.cos.html#numpy.cos
import matplotlib.pylab as plt
x = np.linspace(-2*np.pi, 2*np.pi, 201)
plt.plot(x, np.sin(2*np.pi*x+np.pi), color="red")
plt.plot(x, np.cos(2*np.pi*x+np.pi), color="blue")
plt.plot(x, np.sin(2*np.pi*x+np.pi)+np.cos(2*np.pi*x+np.pi), color="yellow")
plt.xlabel('Angle [rad]')
plt.ylabel('F(x)')
plt.axis('tight')
plt.show()
# # Lecture plots
# Periodically during lecture we will take a pause to plot some of the interesting functions that we use in class.
# ## Classical wavefunctions
#
# The following plot shows the the spacial component of the standard wavefunction with a wavelength of $\lambda=\text{1.45 m}$ and a relative amplitude of $A=1$ when the time, $t=0$ and the phase $\phi=1.0$.
##SKIP
x=np.linspace(0,3.0,100)
sinx=np.sin(2*np.pi*x+0+1)
plt.plot(x,sinx, color="black")
plt.xlabel('x', fontsize=24)
plt.ylabel("y(x)", fontsize=24)
plt.show()
# Make a new figure where you plot the same wave function at three time points in the future. Assume the frequency is $\nu=.1 \text{ ms / s} $ Use a different color for each plot
#SKIP
import numpy
x=numpy.linspace(0,3.0,100)
sinx1=np.sin(2*np.pi*x+3+0.1)
sinx2=np.sin(2*np.pi*x+6+0.1)
sinx3=np.sin(2*np.pi*x+9+0.1)
plt.plot(x,sinx1, color="black")
plt.plot(x,sinx2, color="red")
plt.plot(x,sinx3, color="yellow")
plt.xlabel('x', fontsize=24)
plt.ylabel("y(x)", fontsize=24)
plt.show()
# ## Orthogonality
# Graphically show that the the following two functions are orthogonal on the interval $-3\pi$ to $3\pi$
# $$ \sin(x) \text{ and } \cos(3x)$$
#
# Plot both functions together, then plot the product of both functions and explain why it is orthogonal
import matplotlib.pylab as plt
x = np.linspace(-3*np.pi, 3*np.pi, 201)
plt.plot(x, np.sin(x), color="red")
plt.plot(x, np.cos(3*x), color="blue")
plt.xlabel('Angle [rad]')
plt.ylabel('F(x)')
plt.axis('tight')
plt.show()
import matplotlib.pylab as plt
x = np.linspace(-3*np.pi, 3*np.pi, 201)
prod=np.sin(x)*np.cos(3*x)
plt.plot(x, prod, color="green")
plt.xlabel('Angle [rad]')
plt.ylabel('F(x)')
plt.axis('tight')
plt.show()
prod=np.sin(x)*np.cos(3*x)
prod=np.sin(x)*np.cos(3*x)
x = np.linspace(-3*np.pi, 3*np.pi, 201)
np.trapz(prod,x)
# +
#The two given functions and the product of both functions are orthogonal because when you take the dot product of the two functions you get a zero back and they are at right angles to one another.
# -
# Use the numpy trapezoid rule integrator to show the the two functions are orthogonal
# `np.trapz(y,x)`
#
# https://docs.scipy.org/doc/numpy/reference/generated/numpy.trapz.html
# +
# Example code
#x=numpy.linspace(0,1.0,20)
#exp_func=np.exp(-2*np.pi*x)
#np.trapz(exp_func,x)
# -
prod=np.sin(x)*np.cos(3*x)
x = np.linspace(-3*np.pi, 3*np.pi, 201)
exp_func=prod
np.trapz(exp_func,x)
|
9-19-2019 - Lecture Notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Gauravds435/colabtools/blob/master/PCA.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="zD16MpsmFa8N"
import pandas as pd
import numpy as np
from google.colab import files
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 75} id="4QnJpL8mIGmO" outputId="d8020f12-90bc-43ec-c24c-562284fe1184"
uploaded = files.upload()
# + colab={"base_uri": "https://localhost:8080/"} id="MCSMpurDISUx" outputId="ea411d74-0cb9-4500-d1b1-4aa95290e85b"
for fn in uploaded.keys():
print("User uploaded file'{name}' with length {length} bytes".format(name = fn, length=len(uploaded[fn])))
# + colab={"base_uri": "https://localhost:8080/"} id="wiLElZHSIUHo" outputId="282812dc-26a6-4c96-c2d3-d6fa17152bff"
uploaded
# + id="x7nCIYU4JKc-"
import io
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="ze_CH-mRIYCI" outputId="b27c2b25-09aa-4931-f905-50925bff737b"
df = pd.read_csv(io.StringIO(uploaded["amsPrediction - Sheet1.csv"].decode("utf-8")))
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="aipXmn3wJVvj" outputId="c4376de7-e310-4116-a93d-f1b3391887a9"
df.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="amPI4nX7Jg0L" outputId="50df27da-f569-487b-8800-4789d3de9f6c"
endog = df["ESE"].head()
exog = df[["MSE","Attendance", "HRS"]].head()
print(exog)
# + colab={"base_uri": "https://localhost:8080/"} id="t7HlKCx7KBG8" outputId="1ee5055d-e14d-4c37-b132-99b60a71ca94"
X = exog.to_numpy()
Y = endog.to_numpy()
print(X)
# + id="xMAAUXaIKNRt"
from numpy import array
from numpy import mean
from numpy import cov
from numpy.linalg import eig
# + colab={"base_uri": "https://localhost:8080/"} id="qXM5_UdLK7ud" outputId="02b4c69d-9695-49c2-8dab-d4670ae46759"
M = mean(X.T, axis = 1)
print(M)
# + colab={"base_uri": "https://localhost:8080/"} id="Di0tgErSLeR4" outputId="ceded9a0-5e84-4592-d0a6-80a0ad26190e"
C = X-M
print(C)
# + colab={"base_uri": "https://localhost:8080/"} id="VuhrB2-rLkbl" outputId="35c7eddc-41a7-4322-9793-58999d5cf771"
V = cov(C.T)
print(V)
# + colab={"base_uri": "https://localhost:8080/"} id="rsmDrfZyLp6b" outputId="66d0af87-416f-4f50-b8d8-0b4e8daba494"
values, vectors = eig(V)
print(values)
print(vectors)
# + colab={"base_uri": "https://localhost:8080/"} id="Sme5vzJAcW7v" outputId="e7770b2d-415d-46b7-d6e5-26a480efd626"
P = vectors.T.dot(C.T)
print(P.T)
# + id="8D0bYRCWcscb"
from sklearn.decomposition import PCA
pca = PCA(n_components=1)
principalcomponents = pca.fit_transform(X)
principalDf = pd.DataFrame(data = principalcomponents, columns = ["principal Component 1"])
# + colab={"base_uri": "https://localhost:8080/"} id="dJCiKYdaduer" outputId="0b1986d8-c03d-45f7-d6ea-032278fd2de0"
import statsmodels.api as sm
exog1 = sm.add_constant(principalDf[["principal Component 1"]])
print(exog1)
# + id="jdN0o91zduXs"
from sklearn.linear_model import LinearRegression
# + id="idPhPMtQeLCR"
lr = LinearRegression()
# + id="a-Minm2WeNm3"
model = lr.fit(exog1, endog)
# + colab={"base_uri": "https://localhost:8080/"} id="bKS-Lp1Ge3Df" outputId="e05235bf-7e07-41a0-e3e9-6d3beb1c4e88"
lr.coef_
# + colab={"base_uri": "https://localhost:8080/"} id="i3l8KLFPe9Gq" outputId="025959ba-a6ad-4b55-c7c4-394d3326ec89"
lr.intercept_
# + id="AF_CZF4CfCnD"
import math
def RSE(y_true, y_predicted):
y_true = np.array(y_true)
y_predicted = np.array(y_predicted)
RSS = np.sum(np.square(y_true-y_predicted))
rse = math.sqrt(RSS/(len(y_true)-2))
return rse
# + id="cgy7aQWtf-mu"
ypred = lr.predict(exog1)
# + colab={"base_uri": "https://localhost:8080/"} id="Z4-SMJuogEJX" outputId="06178a1c-2b05-4bbc-d4c7-9d88bd3788c5"
re = RSE(endog, ypred)
print(re)
|
PCA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # In-Class Coding Lab: Web Services and APIs
#
# ### Overview
#
# The web has long evolved from user-consumption to device consumption. In the early days of the web when you wanted to check the weather, you opened up your browser and visited a website. Nowadays your smart watch / smart phone retrieves the weather for you and displays it on the device. Your device can't predict the weather. It's simply consuming a weather based service.
#
# The key to making device consumption work are API's (Application Program Interfaces). Products we use everyday like smartphones, Amazon's Alexa, and gaming consoles all rely on API's. They seem "smart" and "powerful" but in actuality they're only interfacing with smart and powerful services in the cloud.
#
# API consumption is the new reality of programming; it is why we cover it in this course. Once you undersand how to conusme API's you can write a program to do almost anything and harness the power of the internet to make your own programs look "smart" and "powerful."
#
# This lab covers how to properly use consume web service API's with Python. Here's what we will cover.
#
# 1. Understading requests and responses
# 1. Proper error handling
# 1. Parameter handling
# 1. Refactoring as a function
#
# Run this to make sure you have the pre-requisites!
# !pip install -q requests
# ## Part 1: Understanding Requests and responses
#
# In this part we learn about the Python requests module. http://docs.python-requests.org/en/master/user/quickstart/
#
# This module makes it easy to write code to send HTTP requests over the internet and handle the responses. It will be the cornerstone of our API consumption in this course. While there are other modules which accomplish the same thing, `requests` is the most straightforward and easiest to use.
#
# We'll begin by importing the modules we will need. We do this here so we won't need to include these lines in the other code we write in this lab.
# start by importing the modules we will need
import requests
import json
# ### The request
#
# As you learned in class and your assigned readings, the HTTP protocol has **verbs** which consititue the type of request you will send to the remote resource, or **url**. Based on the url and request type, you will get a **response**.
#
# The following line of code makes a **get** request (that's the HTTP verb) to Google's Geocoding API service. This service attempts to convert the address (in this case `Syracuse University`) into a set of coordinates global coordinates (Latitude and Longitude), so that location can be plotted on a map.
#
url = 'https://nominatim.openstreetmap.org/search?q=Hinds+Hall+Syracuse+University&format=json'
response = requests.get(url)
# ### The response
#
# The `get()` method returns a `Response` object variable. I called it `response` in this example but it could be called anything.
#
# The HTTP response consists of a *status code* and *body*. The status code lets you know if the request worked, while the body of the response contains the actual data.
#
response.ok # did the request work?
response.text # what's in the body of the response, as a raw string
# ### Converting responses into Python object variables
#
# In the case of **web site url's** the response body is **HTML**. This should be rendered in a web browser. But we're dealing with Web Service API's so...
#
# In the case of **web API url's** the response body could be in a variety of formats from **plain text**, to **XML** or **JSON**. In this course we will only focus on JSON format because as we've seen these translate easily into Python object variables.
#
# Let's convert the response to a Python object variable. I this case it will be a Python dictionary
geodata = response.json() # try to decode the response from JSON format
geodata # this is now a Python object variable
# With our Python object, we can now walk the list of dictionary to retrieve the latitude and longitude
#
lat = geodata[0]['lat']
lon =geodata[0]['lon']
print(lat, lon)
# In the code above we "walked" the Python list of dictionary to get to the location
#
# - `geodata` is a list
# - `geodata[0]` is the first item in that list, a dictionary
# - `geodata[0]['lat']` is a dictionary key which represents the latitude
# - `geodata[0]['lon']` is a dictionary key which represents the longitude
#
# It should be noted that this process will vary for each API you call, so its important to get accustomed to performing this task. You'll be doing it quite often.
#
# One final thing to address. What is the type of `lat` and `lon`?
type(lat), type(lon)
# Bummer they are strings. we want them to be floats so we will need to parse the strings with the `float()` function:
lat = float(geodata[0]['lat'])
lon = float(geodata[0]['lon'])
print("Latitude: %f, Longitude: %f" % (lat, lon))
# ### Now You Try It!
#
# Walk the `geodata` object variable and reteieve the value under the key `display_name` and the key `bounding_box`
# todo:
# retrieve the place_id put in a variable
# retrieve the formatted_address put it in a variable
# print both of them out
display_name = geodata[0]['display_name']
bounding_box = geodata[0]['boundingbox']
print('Display name: %s\nBounding box: %s' %(display_name, bounding_box))
# ## Part 2: Parameter Handling
#
# In the example above we hard-coded "Hinds Hall Syracuse University" into the request:
# ```
# url = 'https://nominatim.openstreetmap.org/search?q=Hinds+Hall+Syracuse+University&format=json'
# ```
# A better way to write this code is to allow for the input of any location and supply that to the service. To make this work we need to send parameters into the request as a dictionary. This way we can geolocate any address!
#
# You'll notice that on the url, we are passing **key-value pairs** the key is `q` and the value is `Hinds+Hall+Syracuse+University`. The other key is `format` and the value is `json`. Hey, Python dictionaries are also key-value pairs so:
url = 'https://nominatim.openstreetmap.org/search' # base URL without paramters after the "?"
search = 'Hinds Hall Syracuse University'
options = { 'q' : search, 'format' : 'json'}
response = requests.get(url, params = options)
geodata = response.json()
coords = { 'lat' : float(geodata[0]['lat']), 'lng' : float(geodata[0]['lon']) }
print("Search for:", search)
print("Coordinates:", coords)
print("%s is located at (%f,%f)" %(search, coords['lat'], coords['lng']))
# ### Looking up any address
#
# RECALL: For `requests.get(url, params = options)` the part that says `params = options` is called a **named argument**, which is Python's way of specifying an optional function argument.
#
# With our parameter now outside the url, we can easily re-write this code to work for any location! Go ahead and execute the code and input `Queens, NY`. This will retrieve the coordinates `(40.728224,-73.794852)`
url = 'https://nominatim.openstreetmap.org/search' # base URL without paramters after the "?"
search = input("Enter a loacation to Geocode: ")
options = { 'q' : search, 'format' : 'json'}
response = requests.get(url, params = options)
geodata = response.json()
coords = { 'lat' : float(geodata[0]['lat']), 'lng' : float(geodata[0]['lon']) }
print("Search for:", search)
print("Coordinates:", coords)
print("%s is located at (%f,%f)" %(search, coords['lat'], coords['lng']))
# ### So useful, it should be a function!
#
# One thing you'll come to realize quickly is that your API calls should be wrapped in functions. This promotes **readability** and **code re-use**. For example:
# +
def get_coordinates(search):
url = 'https://nominatim.openstreetmap.org/search' # base URL without paramters after the "?"
options = { 'q' : search, 'format' : 'json'}
response = requests.get(url, params = options)
geodata = response.json()
coords = { 'lat' : float(geodata[0]['lat']), 'lng' : float(geodata[0]['lon']) }
return coords
# main program here:
location = input("Enter a location: ")
coords = get_coordinates(location)
print("%s is located at (%f,%f)" %(location, coords['lat'], coords['lng']))
# -
# ### Other request methods
#
# Not every API we call uses the `get()` method. Some use `post()` because the amount of data you provide it too large to place on the url.
#
# An example of this is the **Text-Processing.com** sentiment analysis service. http://text-processing.com/docs/sentiment.html This service will detect the sentiment or mood of text. You give the service some text, and it tells you whether that text is positive, negative or neutral.
# 'you suck' == 'negative'
url = 'http://text-processing.com/api/sentiment/'
options = { 'text' : 'you suck'}
response = requests.post(url, data = options)
sentiment = response.json()
sentiment
# 'I love cheese' == 'positive'
url = 'http://text-processing.com/api/sentiment/'
options = { 'text' : 'I love cheese'}
response = requests.post(url, data = options)
sentiment = response.json()
sentiment
# In the examples provided we used the `post()` method instead of the `get()` method. the `post()` method has a named argument `data` which takes a dictionary of data. The key required by **text-processing.com** is `text` which hold the text you would like to process for sentiment.
#
# We use a post in the event the text we wish to process is very long. Case in point:
tweet = "<NAME> isn't voluntarily leaving the Apprentice, he was fired by his bad (pathetic) ratings, not by me. Sad end to a great show"
url = 'http://text-processing.com/api/sentiment/'
options = { 'text' : tweet }
response = requests.post(url, data = options)
sentiment = response.json()
sentiment
# ### Now You Try It!
#
# Use the above example to write a program which will input any text and print the sentiment using this API!
# todo write code here
text = input("Enter text: ")
url = 'http://text-processing.com/api/sentiment/'
options = {'text' : text}
response = requests.post(url, data = options)
sentiment = response.json()
if sentiment['label'] == 'neg':
print('Sentiment: negative')
elif sentiment['label'] == 'pos':
print('Sentiment: positive')
else:
print('Sentiment: neutral')
#
# ## Part 3: Proper Error Handling (In 3 Simple Rules)
#
# When you write code that depends on other people's code from around the Internet, there's a lot that can go wrong. Therefore we perscribe the following advice:
#
# ```
# Assume anything that CAN go wrong WILL go wrong
# ```
#
# ### Rule 1: Don't assume the internet 'always works'
#
# The first rule of programming over a network is to NEVER assume the network is available. You need to assume the worst. No WiFi, user types in a bad url, the remote website is down, etc.
#
# We handle this in the `requests` module by catching the `requests.exceptions.RequestException` Here's an example:
# +
url = "http://this is not a website"
try:
response = requests.get(url) # throws an exception when it cannot connect
# internet is broken
except requests.exceptions.RequestException as e:
print("ERROR: Cannot connect to ", url)
print("DETAILS:", e)
# -
# ### Rule 2: Don't assume the response you get back is valid
#
# Assuming the internet is not broken (Rule 1) You should now check for HTTP response 200 which means the url responded successfully. Other responses like 404 or 501 indicate an error occured and that means you should not keep processing the response.
#
# Here's one way to do it:
# +
url = 'http://www.syr.edu/mikeisawesum' # this should 404
try:
response = requests.get(url)
if response.ok: # same as response.status_code == 200
data = response.text
else: # Some other non 200 response code
print("There was an Error requesting:", url, " HTTP Response Code: ", response.status_code)
# internet is broken
except requests.exceptions.RequestException as e:
print("ERROR: Cannot connect to ", url)
print("DETAILS:", e)
# -
# ### Rule 2a: Use exceptions instead of if else in this case
#
# Personally I don't like to use `if ... else` to handle an error. Instead, I prefer to instruct `requests` to throw an exception of `requests.exceptions.HTTPError` whenever the response is not ok. This makes the code you write a little cleaner.
#
# Errors are rare occurences, and so I don't like error handling cluttering up my code.
#
# +
url = 'http://www.syr.edu/mikeisawesum' # this should 404
try:
response = requests.get(url) # throws an exception when it cannot connect
response.raise_for_status() # throws an exception when not 'ok'
data = response.text
# response not ok
except requests.exceptions.HTTPError as e:
print("ERROR: Response from ", url, 'was not ok.')
print("DETAILS:", e)
# internet is broken
except requests.exceptions.RequestException as e:
print("ERROR: Cannot connect to ", url)
print("DETAILS:", e)
# -
# ### Rule 3: Don't assume the data you get back is the data you expect.
#
# And finally, do not assume the data arriving the the `response` is the data you expected. Specifically when you try and decode the `JSON` don't assume that will go smoothly. Catch the `json.decoder.JSONDecodeError`.
# +
url = 'http://www.syr.edu' # this is HTML, not JSON
try:
response = requests.get(url) # throws an exception when it cannot connect
response.raise_for_status() # throws an exception when not 'ok'
data = response.json() # throws an exception when cannot decode json
# cannot decode json
except json.decoder.JSONDecodeError as e:
print("ERROR: Cannot decode the response into json")
print("DETAILS", e)
# response not ok
except requests.exceptions.HTTPError as e:
print("ERROR: Response from ", url, 'was not ok.')
print("DETAILS:", e)
# internet is broken
except requests.exceptions.RequestException as e:
print("ERROR: Cannot connect to ", url)
print("DETAILS:", e)
# -
# ### Now You try it!
#
# Using the last example above, write a program to input a location, call the `get_coordinates()` function, then print the coordindates. Make sure to handle all three types of exceptions!!!
#
# +
import requests
import json
def get_coordinates_using_google(location):
options = { 'q' : location, 'format' : 'json'}
try:
response = requests.get(url, params = options)
geodata = response.json()
lat = float(geodata[0]['lat'])
lon = float(geodata[0]['lon'])
coords = [lat, lon]
return coords
except requests.exceptions.HTTPError as e:
print("Error: Response was not ok")
return e
except requests.exceptions.RequestException as e:
print("Error: Could not connect to site")
return e
except json.decoder.JSONDecodeError as e:
print("Error: Could not decode the response into JSON")
return e
# todo write code here to input a location, look up coordinates, and print
# it should handle errors!!!
url = 'https://nominatim.openstreetmap.org/search'
location = input("Enter location: ")
coords = get_coordinates_using_google(location)
try:
print(coords)
except:
print('Details: ', e)
# -
|
content/lessons/11/Class-Coding-Lab/CCL-Web-Services-And-APIs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (tensorflow)
# language: python
# name: rga
# ---
# # T81-558: Applications of Deep Neural Networks
# **Module 3: Introduction to TensorFlow**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# # Module 3 Material
#
# * Part 3.1: Deep Learning and Neural Network Introduction [[Video]](https://www.youtube.com/watch?v=zYnI4iWRmpc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_03_1_neural_net.ipynb)
# * Part 3.2: Introduction to Tensorflow and Keras [[Video]](https://www.youtube.com/watch?v=PsE73jk55cE&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_03_2_keras.ipynb)
# * Part 3.3: Saving and Loading a Keras Neural Network [[Video]](https://www.youtube.com/watch?v=-9QfbGM1qGw&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_03_3_save_load.ipynb)
# * Part 3.4: Early Stopping in Keras to Prevent Overfitting [[Video]](https://www.youtube.com/watch?v=m1LNunuI2fk&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_03_4_early_stop.ipynb)
# * **Part 3.5: Extracting Weights and Manual Calculation** [[Video]](https://www.youtube.com/watch?v=7PWgx16kH8s&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_03_5_weights.ipynb)
# # Part 3.5: Extracting Keras Weights and Manual Neural Network Calculation
#
# In this section we will build a neural network and analyze it down the individual weights. We will train a simple neural network that learns the XOR function. It is not hard to simply hand-code the neurons to provide an [XOR function](https://en.wikipedia.org/wiki/Exclusive_or); however, for simplicity, we will allow Keras to train this network for us. We will just use 100K epochs on the ADAM optimizer. This is massive overkill, but it gets the result, and our focus here is not on tuning. The neural network is small. Two inputs, two hidden neurons, and a single output.
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation
import numpy as np
# Create a dataset for the XOR function
x = np.array([
[0,0],
[1,0],
[0,1],
[1,1]
])
y = np.array([
0,
1,
1,
0
])
# Build the network
# sgd = optimizers.SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
done = False
cycle = 1
while not done:
print("Cycle #{}".format(cycle))
cycle+=1
model = Sequential()
model.add(Dense(2, input_dim=2, activation='relu'))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(x,y,verbose=0,epochs=10000)
# Predict
pred = model.predict(x)
# Check if successful. It takes several runs with this small of a network
done = pred[0]<0.01 and pred[3]<0.01 and pred[1] > 0.9 and pred[2] > 0.9
print(pred)
# -
pred[3]
# The output above should have two numbers near 0.0 for the first and forth spots (input [[0,0]] and [[1,1]]). The middle two numbers should be near 1.0 (input [[1,0]] and [[0,1]]). These numbers are in scientific notation. Due to random starting weights, it is sometimes necessary to run the above through several cycles to get a good result.
#
# Now that the neural network is trained, lets dump the weights.
# Dump weights
for layerNum, layer in enumerate(model.layers):
weights = layer.get_weights()[0]
biases = layer.get_weights()[1]
for toNeuronNum, bias in enumerate(biases):
print(f'{layerNum}B -> L{layerNum+1}N{toNeuronNum}: {bias}')
for fromNeuronNum, wgt in enumerate(weights):
for toNeuronNum, wgt2 in enumerate(wgt):
print(f'L{layerNum}N{fromNeuronNum} -> L{layerNum+1}N{toNeuronNum} = {wgt2}')
# If you rerun this, you probably get different weights. There are many ways to solve the XOR function.
#
# In the next section, we copy/paste the weights from above and recreate the calculations done by the neural network. Because weights can change with each training, the weights used for the below code came from this:
#
# ```
# 0B -> L1N0: -1.2913415431976318
# 0B -> L1N1: -3.021530048386012e-08
# L0N0 -> L1N0 = 1.2913416624069214
# L0N0 -> L1N1 = 1.1912699937820435
# L0N1 -> L1N0 = 1.2913411855697632
# L0N1 -> L1N1 = 1.1912697553634644
# 1B -> L2N0: 7.626241297587034e-36
# L1N0 -> L2N0 = -1.548777461051941
# L1N1 -> L2N0 = 0.8394404649734497
# ```
# +
input0 = 0
input1 = 1
hidden0Sum = (input0*1.3)+(input1*1.3)+(-1.3)
hidden1Sum = (input0*1.2)+(input1*1.2)+(0)
print(hidden0Sum) # 0
print(hidden1Sum) # 1.2
hidden0 = max(0,hidden0Sum)
hidden1 = max(0,hidden1Sum)
print(hidden0) # 0
print(hidden1) # 1.2
outputSum = (hidden0*-1.6)+(hidden1*0.8)+(0)
print(outputSum) # 0.96
output = max(0,outputSum)
print(output) # 0.96
# -
|
t81_558_class_03_5_weights.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import numba
import matplotlib.pyplot as plt
import sympy as sym
plt.style.use('presentation.mplstyle')
# #%matplotlib notebook
def d2np(d):
names = []
numbers = ()
dtypes = []
for item in d:
names += item
if type(d[item]) == float:
numbers += (d[item],)
dtypes += [(item,float)]
if type(d[item]) == int:
numbers += (d[item],)
dtypes += [(item,int)]
if type(d[item]) == np.ndarray:
numbers += (d[item],)
dtypes += [(item,np.float64,d[item].shape)]
return np.array([numbers],dtype=dtypes)
# -
# ### Fortescue
# +
alpha = np.exp(2.0/3*np.pi*1j)
A_0a = np.array([[1, 1, 1],
[1, alpha**2, alpha],
[1, alpha, alpha**2]])
A_a0 = 1/3* np.array([[1, 1, 1],
[1, alpha, alpha**2],
[1, alpha**2, alpha]])
# -
# ### Voltage source
# +
theta = np.deg2rad(20.0)
V_zero = 0.0*np.exp(1j*0.0)
V_neg = 20.0*np.exp(1j*0.0)
V_pos =400.0/np.sqrt(3)*np.exp(1j*theta)
V_zpn = np.array([[V_zero],[V_pos],[V_neg]])
V_abc = A_0a @ V_zpn
# -
# ### Control inputs
# +
L = 500e-6
R = 0.01
omega = 2.0*np.pi*50.0
w = omega
v_dc = 800.0
V_012 = A_a0 @ V_abc
v_z = V_012[0,0]
v_p = V_012[1,0]
v_n = V_012[2,0]
# -
# ### PLL
theta_pll = np.angle(v_p)
# ### Park
# +
v_dq_z = v_z
v_dq_p = v_p*np.exp(-1j*theta_pll)*np.sqrt(2)
v_dq_n = v_n*np.exp( 1j*theta_pll)*np.sqrt(2)
v_d_z = v_dq_z.real # ??
v_q_z = v_dq_z.imag # ??
v_d_p = v_dq_p.imag
v_q_p = v_dq_p.real
v_d_n = v_dq_n.imag
v_q_n = v_dq_n.real
# -
# ### References
# +
p_ref = 0.6e6
q_ref = 0.2e6
pq_ref = np.array([p_ref,q_ref,0,0]).reshape(4,1)
i2p=3/2*np.array([[ v_d_p, v_q_p, v_d_n, v_q_n], # i_d_p
[-v_q_p, v_d_p,-v_q_n, v_d_n], # i_q_p
[-v_q_n, v_d_n, v_q_p,-v_d_p], # i_d_n
[ v_d_n, v_q_n, v_d_p, v_q_p]]) # i_q_n
p2i=np.linalg.inv(i2p)
i_dq_pn = p2i@pq_ref
i_d_p_ref = 100.0
i_q_p_ref = 0.0
i_d_n_ref = 0.0
i_q_n_ref = 0.0
i_d_p_ref = i_dq_pn[0,0]
i_q_p_ref = i_dq_pn[1,0]
i_d_n_ref = i_dq_pn[2,0]
i_q_n_ref = i_dq_pn[3,0]
mode = 'p_cte'
if mode == 'p_pos_i_n_0':
i_d_p_ref = -(0.666666666666667*p_ref*v_d_p*(v_d_n**2 + v_d_p**2 + v_q_n**2 + v_q_p**2) + 0.666666666666667*q_ref*v_q_p*(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2))/(v_d_n**4 + 2.0*v_d_n**2*v_q_n**2 - v_d_p**4 - 2.0*v_d_p**2*v_q_p**2 + v_q_n**4 - v_q_p**4)
i_q_p_ref = 0.666666666666667*(-p_ref*v_q_p*(v_d_n**2 + v_d_p**2 + v_q_n**2 + v_q_p**2) + q_ref*v_d_p*(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2))/(v_d_n**4 + 2.0*v_d_n**2*v_q_n**2 - v_d_p**4 - 2.0*v_d_p**2*v_q_p**2 + v_q_n**4 - v_q_p**4)
i_d_n_ref = 0
i_q_n_ref = 0
if mode == 'q_cte':
i_d_p_ref = 0.666666666666667*(p_ref*v_d_p*(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2) + q_ref*v_q_p*(v_d_n**2 + v_d_p**2 + v_q_n**2 + v_q_p**2))/(v_d_n**4 + 2.0*v_d_n**2*v_q_n**2 - v_d_p**4 - 2.0*v_d_p**2*v_q_p**2 + v_q_n**4 - v_q_p**4)
i_q_p_ref = 0.666666666666667*(p_ref*v_q_p*(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2) - q_ref*v_d_p*(v_d_n**2 + v_d_p**2 + v_q_n**2 + v_q_p**2))/(v_d_n**4 + 2.0*v_d_n**2*v_q_n**2 - v_d_p**4 - 2.0*v_d_p**2*v_q_p**2 + v_q_n**4 - v_q_p**4)
i_d_n_ref = 0.666666666666667*(p_ref*v_d_n*(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2) - q_ref*v_q_n*(v_d_n**2 + v_d_p**2 + v_q_n**2 + v_q_p**2))/(v_d_n**4 + 2.0*v_d_n**2*v_q_n**2 - v_d_p**4 - 2.0*v_d_p**2*v_q_p**2 + v_q_n**4 - v_q_p**4)
i_q_n_ref = 0.666666666666667*(p_ref*v_q_n*(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2) + q_ref*v_d_n*(v_d_n**2 + v_d_p**2 + v_q_n**2 + v_q_p**2))/(v_d_n**4 + 2.0*v_d_n**2*v_q_n**2 - v_d_p**4 - 2.0*v_d_p**2*v_q_p**2 + v_q_n**4 - v_q_p**4)
if mode == 'pq_cte': # Lipo
i_d_p_ref = 0.666666666666667*(-p_ref*v_d_p + q_ref*v_q_p)/(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2)
i_q_p_ref = -(0.666666666666667*p_ref*v_q_p + 0.666666666666667*q_ref*v_d_p)/(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2)
i_d_n_ref = 0.666666666666667*(p_ref*v_d_n + q_ref*v_q_n)/(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2)
i_q_n_ref = 0.666666666666667*(p_ref*v_q_n - q_ref*v_d_n)/(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2)
if mode == 'p_cte':
i_d_p_ref = -(0.666666666666667*p_ref*v_d_p*(v_d_n**2 + v_d_p**2 + v_q_n**2 + v_q_p**2) + 0.666666666666667*q_ref*v_q_p*(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2))/(v_d_n**4 + 2.0*v_d_n**2*v_q_n**2 - v_d_p**4 - 2.0*v_d_p**2*v_q_p**2 + v_q_n**4 - v_q_p**4)
i_q_p_ref = 0.666666666666667*(-p_ref*v_q_p*(v_d_n**2 + v_d_p**2 + v_q_n**2 + v_q_p**2) + q_ref*v_d_p*(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2))/(v_d_n**4 + 2.0*v_d_n**2*v_q_n**2 - v_d_p**4 - 2.0*v_d_p**2*v_q_p**2 + v_q_n**4 - v_q_p**4)
i_d_n_ref = 0.666666666666667*(p_ref*v_d_n*(v_d_n**2 + v_d_p**2 + v_q_n**2 + v_q_p**2) - q_ref*v_q_n*(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2))/(v_d_n**4 + 2.0*v_d_n**2*v_q_n**2 - v_d_p**4 - 2.0*v_d_p**2*v_q_p**2 + v_q_n**4 - v_q_p**4)
i_q_n_ref = 0.666666666666667*(p_ref*v_q_n*(v_d_n**2 + v_d_p**2 + v_q_n**2 + v_q_p**2) + q_ref*v_d_n*(v_d_n**2 - v_d_p**2 + v_q_n**2 - v_q_p**2))/(v_d_n**4 + 2.0*v_d_n**2*v_q_n**2 - v_d_p**4 - 2.0*v_d_p**2*v_q_p**2 + v_q_n**4 - v_q_p**4)
if mode == 'z_mode':
I_p_ref = np.conj((p_ref+1j*q_ref)/v_p)/3/np.sqrt(3)
Z_p = v_p/I_p_ref
I_n_ref = np.conj((p_ref+1j*q_ref)/v_n)/3/np.sqrt(3)
Z_n = v_n/I_n_ref
i_d_p_ref = ((v_q_p + 1j*v_d_p)/Z_p).imag
i_q_p_ref = ((v_q_p + 1j*v_d_p)/Z_p).real
i_d_n_ref = ((v_q_n + 1j*v_d_n)/Z_n).imag
i_q_n_ref = ((v_q_n + 1j*v_d_n)/Z_n).real
# -
# ### Control
# +
#L*did = e_d - R*i_d - w*L*i_q - v_d
#L*diq = e_q - R*i_q + w*L*i_d - v_q
eta_d_p = 2.0/v_dc*(R*i_d_p_ref + L*w*i_q_p_ref + v_d_p)
eta_q_p = 2.0/v_dc*(R*i_q_p_ref - L*w*i_d_p_ref + v_q_p)
eta_d_n = 2.0/v_dc*(R*i_d_n_ref + L*w*i_q_n_ref + v_d_n)
eta_q_n = 2.0/v_dc*(R*i_q_n_ref - L*w*i_d_n_ref + v_q_n)
eta_dq_p = eta_q_p + 1j*eta_d_p
e_dq_p = v_dc/2.0*eta_dq_p # phase-neutral peak value
eta_dq_n = eta_q_n + 1j*eta_d_n
e_dq_n = v_dc/2.0*eta_dq_n # phase-neutral peak value
# -
# ### Modulation
# +
e_p = e_dq_p *np.exp( 1j*theta_pll)/np.sqrt(2) # phase-neutral RMS value
e_n = e_dq_n *np.exp(-1j*theta_pll)/np.sqrt(2) # phase-neutral RMS value
e_z = 0.0
#e_n = 0.0
e_012 = np.array([e_z,e_p,e_n]).reshape(3,1)
e_abc = A_0a @ e_012
# -
# ### Plant
# +
Z_1 = R +1j *L*omega
Z_2 = Z_1
Z_0 = Z_1
Z_012 = np.diag([Z_0,Z_1,Z_2])
Z_abc = A_0a @ Z_012 @ A_a0
Y_abc = np.linalg.inv(Z_abc)
I_abc = Y_abc @ (e_abc-V_abc)
I_abc
# -
V_abc.T @ np.conj(I_abc)
# +
I_012 = A_a0 @ I_abc
i_dq_z_out = I_012[0] ## ???
i_dq_p_out = I_012[1]*np.exp(-1j*theta_pll)*np.sqrt(2)
i_dq_n_out = I_012[2]*np.exp( 1j*theta_pll)*np.sqrt(2)
i_d_p = i_dq_p_out.imag
i_q_p = i_dq_p_out.real
i_d_n = i_dq_n_out.imag
i_q_n = i_dq_n_out.real
print(i_d_p_ref,i_d_p)
print(i_q_p_ref,i_q_p)
print(i_d_n_ref,i_d_n)
print(i_q_n_ref,i_q_n)
# -
# ## Fisix
# +
p_cte_ref = 1.5*i_d_n*v_d_n + 1.5*i_d_p*v_d_p + 1.5*i_q_n*v_q_n + 1.5*i_q_p*v_q_p
p_cos_ref = 1.5*i_d_n*v_d_p + 1.5*i_d_p*v_d_n + 1.5*i_q_n*v_q_p + 1.5*i_q_p*v_q_n
p_sin_ref = 1.5*i_d_n*v_q_p - 1.5*i_d_p*v_q_n - 1.5*i_q_n*v_d_p + 1.5*i_q_p*v_d_n
q_cte_ref = -1.5*i_d_n*v_q_n - 1.5*i_d_p*v_q_p + 1.5*i_q_n*v_d_n + 1.5*i_q_p*v_d_p
q_cos_ref = -1.5*i_d_n*v_q_p - 1.5*i_d_p*v_q_n + 1.5*i_q_n*v_d_p + 1.5*i_q_p*v_d_n
q_sin_ref = 1.5*i_d_n*v_d_p - 1.5*i_d_p*v_d_n + 1.5*i_q_n*v_q_p - 1.5*i_q_p*v_q_n
lhs = ['p_cte_ref','p_cos_ref','p_sin_ref','q_cte_ref','q_cos_ref','q_sin_ref']
rhs = [p_cte_ref,p_cos_ref,p_sin_ref,q_cte_ref,q_cos_ref,q_sin_ref]
for lh,rh in zip(lhs,rhs):
print('{:s}_ref = {:s}'.format(str(lh) ,str(sym.simplify(rh))))
# -
# ### From phasor to time
# +
t = np.linspace(0.0,0.04-0.04/1000,1000)
v_a = (np.exp(1j*w*t)*V_abc[0]).real*np.sqrt(2)
v_b = (np.exp(1j*w*t)*V_abc[1]).real*np.sqrt(2)
v_c = (np.exp(1j*w*t)*V_abc[2]).real*np.sqrt(2)
i_a = (np.exp(1j*w*t)*I_abc[0]).real*np.sqrt(2)
i_b = (np.exp(1j*w*t)*I_abc[1]).real*np.sqrt(2)
i_c = (np.exp(1j*w*t)*I_abc[2]).real*np.sqrt(2)
v_a_p = (np.exp(1j*(w*t-np.pi/2))*V_abc[0]).real*np.sqrt(2)
v_b_p = (np.exp(1j*(w*t-np.pi/2))*V_abc[1]).real*np.sqrt(2)
v_c_p = (np.exp(1j*(w*t-np.pi/2))*V_abc[2]).real*np.sqrt(2)
#i_a = i_a_p + i_a_n
#i_b = i_c_p + i_c_n
#i_c = i_b_p + i_b_n
p = v_a*i_a + v_b*i_b + v_c*i_c
q = (i_a*(v_b-v_c) + i_b*(v_c-v_a) + i_c*(v_a-v_b))/np.sqrt(3)
q_lipo = v_a_p*i_a + v_b_p*i_b + v_c_p*i_c
#q = (i_a*(v_c-v_b) + i_b*(v_a-v_c) + i_c*(v_b-v_a))/np.sqrt(3)
# -
I_abc
# +
I_zpn = A_a0 @ I_abc
V_zpn = A_a0 @ V_abc
I_p = I_zpn[1]
I_n = I_zpn[2]
V_p = V_zpn[1]
V_n = V_zpn[2]
w = 2.0*np.pi*50.0
i_alpha_p = (np.exp( 1j*w*t)*I_p).imag*np.sqrt(2)
i_beta_p = (np.exp( 1j*w*t)*I_p).real*np.sqrt(2)
i_alpha_n = (np.exp(-1j*w*t)*I_n).imag*np.sqrt(2)
i_beta_n = (np.exp(-1j*w*t)*I_n).real*np.sqrt(2)
v_alpha_p = (np.exp( 1j*w*t)*V_p).imag*np.sqrt(2)
v_beta_p = (np.exp( 1j*w*t)*V_p).real*np.sqrt(2)
v_alpha_n = (np.exp(-1j*w*t)*V_n).imag*np.sqrt(2)
v_beta_n = (np.exp(-1j*w*t)*V_n).real*np.sqrt(2)
v_alpha_p_lipo = (-1j*np.exp( 1j*w*t)*V_p).imag*np.sqrt(2)
v_beta_p_lipo = (-1j*np.exp( 1j*w*t)*V_p).real*np.sqrt(2)
v_alpha_n_lipo = (1j*np.exp(-1j*w*t)*V_n).imag*np.sqrt(2)
v_beta_n_lipo = (1j*np.exp(-1j*w*t)*V_n).real*np.sqrt(2)
i_alpha = i_alpha_p + i_alpha_n
i_beta = i_beta_p + i_beta_n
v_alpha = v_alpha_p + v_alpha_n
v_beta = v_beta_p + v_beta_n
v_alpha_lipo = v_alpha_p_lipo + v_alpha_n_lipo
v_beta_lipo = v_beta_p_lipo + v_beta_n_lipo
#Clark = 2/3*[[1/np.sqrt(2),1/np.sqrt(2),1/np.sqrt(2)],
# [1,-0.5,-0.5]
# [0,-np.sqrt(3)/2,np.sqrt(3)/2]]
#i_oab = np.array([0.0,i_alpha,i_beta])
#v_oab = np.array([0.0,v_alpha,v_beta])
inv_Clark=np.linalg.inv(Clark)
def oab2abc(alpha,beta):
N_t = len(alpha)
abc = np.zeros((3,N_t))
for it in range():
abc[:,it] = Clark
#for
#v_abc = np.lianlg.solve(Clark,v_oab)
p = 3/2*(i_alpha*v_alpha + i_beta*v_beta)
q = 3/2*(v_alpha*i_beta - v_beta*i_alpha)
q_lipo = 3/2*(i_alpha*v_alpha_lipo + i_beta*v_beta_lipo)
# +
fig, axes = plt.subplots(nrows=3, ncols=1, figsize=(8, 4), sharex = True)
axes[0].plot(t, v_alpha)
axes[0].plot(t, v_beta)
axes[1].plot(t, i_alpha)
axes[1].plot(t, i_beta)
axes[2].plot(t, p/1000)
axes[2].plot(t, q/1000)
axes[2].plot(t, q_lipo/1000)
print('p = ',np.average(p))
print('q = ',np.average(q))
print('q_lipo = ',np.average(q_lipo))
print('i_alpha_max = ',np.max(abs(i_alpha)))
print('i_beta_max = ',np.max(abs(i_beta)))
# + active=""
# Lipo
#
# Figure 1
#
# p = 500000.0
# q = 1400000.0
# q_lipo = 200000.0
# i_alpha_max = 8080.75866864
# i_beta_max = 1538.33671853
#
# -
# ### Reference following check
# +
I_012 = A_a0 @ I_abc
i_dq_z_out = I_012[0]*np.exp(-1j*theta_pll)*np.sqrt(2)
i_dq_p_out = I_012[1]*np.exp(-1j*theta_pll)*np.sqrt(2)
i_dq_n_out = I_012[2]*np.exp(-1j*theta_pll)*np.sqrt(2)
i_d_p = i_dq_p_out.imag
i_q_p = i_dq_p_out.real
i_d_n = i_dq_n_out.imag
i_q_n = i_dq_n_out.real
print(i_d_p_ref,i_dq_p_out.real)
print(i_q_p_ref,i_dq_p_out.imag)
print(i_d_n_ref,i_dq_n_out.real)
print(i_q_n_ref,i_dq_n_out.imag)
p_cte_ref = 1.5*i_d_n*v_d_n + 1.5*i_d_p*v_d_p + 1.5*i_q_n*v_q_n + 1.5*i_q_p*v_q_p
p_cos_ref = 1.5*i_d_n*v_d_p + 1.5*i_d_p*v_d_n + 1.5*i_q_n*v_q_p + 1.5*i_q_p*v_q_n
p_sin_ref =-1.5*i_d_n*v_q_p + 1.5*i_d_p*v_q_n + 1.5*i_q_n*v_d_p - 1.5*i_q_p*v_d_n
q_cte_ref = 1.5*i_d_n*v_q_n + 1.5*i_d_p*v_q_p - 1.5*i_q_n*v_d_n - 1.5*i_q_p*v_d_p
q_cos_ref = 1.5*i_d_n*v_q_p + 1.5*i_d_p*v_q_n - 1.5*i_q_n*v_d_p - 1.5*i_q_p*v_d_n
q_sin_ref = 1.5*i_d_n*v_d_p - 1.5*i_d_p*v_d_n + 1.5*i_q_n*v_q_p - 1.5*i_q_p*v_q_n
# Lipo
p_cte_ref = 1.5*i_d_n*v_d_n + 1.5*i_d_p*v_d_p + 1.5*i_q_n*v_q_n + 1.5*i_q_p*v_q_p
p_cos_ref = 1.5*i_d_n*v_d_p + 1.5*i_d_p*v_d_n + 1.5*i_q_n*v_q_p + 1.5*i_q_p*v_q_n
p_sin_ref = -1.5*i_d_n*v_q_p + 1.5*i_d_p*v_q_n + 1.5*i_q_n*v_d_p - 1.5*i_q_p*v_d_n
q_cte_ref = -1.5*i_d_n*v_q_n + 1.5*i_d_p*v_q_p + 1.5*i_q_n*v_d_n - 1.5*i_q_p*v_d_p
q_cos_ref = 1.5*i_d_n*v_q_p - 1.5*i_d_p*v_q_n - 1.5*i_q_n*v_d_p + 1.5*i_q_p*v_d_n
q_sin_ref = 1.5*i_d_n*v_d_p + 1.5*i_d_p*v_d_n + 1.5*i_q_n*v_q_p + 1.5*i_q_p*v_q_n
lhs = ['p_cte_ref','p_cos_ref','p_sin_ref','q_cte_ref','q_cos_ref','q_sin_ref']
rhs = [p_cte_ref,p_cos_ref,p_sin_ref,q_cte_ref,q_cos_ref,q_sin_ref]
for lh,rh in zip(lhs,rhs):
print('{:s}_ref = {:s}'.format(str(lh) ,str(sym.simplify(rh))))
# -
# ### Positive sequence calculation
Z = R +1j *L*omega
I_pos = (e_p - v_p)/Z
I_pos
S =V_abc.T @ np.conj(I_abc)
S
I_012 = A_a0 @ I_abc
I_012*np.sqrt(2)
import sympy as sym
# +
v_d_p,v_q_p,v_d_n,v_q_n = sym.symbols('v_d_p,v_q_p,v_d_n,v_q_n')
i2p = sym.Matrix([[ v_d_p, v_q_p, v_d_n, v_q_n],
[-v_q_p, v_d_p,-v_q_n, v_d_n],
[-v_q_n, v_d_n, v_q_p,-v_d_p],
[ v_d_n, v_q_n, v_d_p, v_q_p]])
p2i = sym.simplify(i2p.inv())
# -
sym.simplify(p2i)
# +
theta = np.deg2rad(0.0)
phi = np.deg2rad(90.0)
V_zero = 0.0*np.exp(1j*0.0)
V_neg =100.0*np.exp(1j*0.0)
V_pos =231.0*np.exp(1j*theta)
V_012 = np.array([[V_zero],[V_pos],[V_neg]])
V_abc = A_0a @ V_012
I_zero = 0.0*np.exp(1j*0.0)
I_neg = 0.0*np.exp(1j*(theta+phi))
I_pos = 10.0*np.exp(1j*(theta-phi))
s_012 = 500e3
sin_012 = 0.0
cos_012 = 0.0
I_pos = (V_neg*sin_012 - V_pos*s_012)/(3*(V_neg**2 - V_pos**2))
I_neg = (V_neg*s_012 - V_pos*sin_012)/(3*(V_neg**2 - V_pos**2))
#I_pos = (-V_neg*sin_012 + V_pos*s_012)/(3*(V_neg**2 + V_pos**2))
#I_neg = ( V_neg*s_012 + V_pos*sin_012)/(3*(V_neg**2 + V_pos**2))
#I = 1j
#I_pos = 0.333333333333333*(V_neg*sin_012 - V_pos*s_012*(1.0 + I))/(V_neg**2*(1.0 - I) - V_pos**2*(1.0 + I))
#I_neg = 0.333333333333333*(V_neg*s_012*(1.0 - I) - V_pos*sin_012)/(V_neg**2*(1.0 - I) - V_pos**2*(1.0 + I))
#I_pos = 0.333333333333333*(V_neg*sin_012 - V_pos*s_012*(1.0 - I))/(V_neg**2*(1.0 + I) - V_pos**2*(1.0 - I))
#I_neg = 0.333333333333333*(V_neg*s_012*(1.0 + I) - V_pos*sin_012)/(V_neg**2*(1.0 + I) - V_pos**2*(1.0 - I))
#I_pos = 0.333333333333333*(I*V_neg*cos_012 + V_pos*s_012)/(V_neg**2 + V_pos**2)
#I_neg = 0.333333333333333*(V_neg*s_012 - I*V_pos*cos_012)/(V_neg**2 + V_pos**2)
#I_pos= (0.166666666666667 - 0.166666666666667*I)*(V_neg*(cos_012 + sin_012) - V_pos*s_012*(1.0 + I))/(V_neg**2 - V_pos**2)
#I_neg= (0.166666666666667 - 0.166666666666667*I)*(V_neg*s_012*(1.0 + I) - V_pos*(cos_012 + sin_012))/(V_neg**2 - V_pos**2)
#I_neg = (cos_012 + sin_012)/(6*V_pos)
#I_pos = (-V_neg*(cos_012 + sin_012) + 2*V_pos*s_012)/(6*V_pos**2)
I_pos = np.conj(s_012/(3*V_pos))
I_neg = -V_neg*I_pos/(V_pos)
I_012 = np.array([[I_zero],[I_pos],[I_neg]])
I_abc = A_0a @ I_012
# +
v_abc = (np.exp(1j*2.0*np.pi*50.0*t)*V_abc).real*np.sqrt(2)
i_abc = (np.exp(1j*2.0*np.pi*50.0*t)*I_abc).real*np.sqrt(2)
p = np.sum(v_abc * i_abc, axis=0)
q = -((v_abc[1]- v_abc[2]) * i_abc[0] + (v_abc[2]- v_abc[0]) * i_abc[1] + (v_abc[0]- v_abc[1]) * i_abc[2] )/np.sqrt(3)
# +
fig, axes = plt.subplots(nrows=3, ncols=1, figsize=(8, 6), sharex = True)
axes[0].plot(t, v_abc[0,:])
axes[0].plot(t, v_abc[1,:])
axes[0].plot(t, v_abc[2,:])
axes[1].plot(t, i_abc[0,:])
axes[1].plot(t, i_abc[1,:])
axes[1].plot(t, i_abc[2,:])
axes[2].plot(t, p/1000)
axes[2].plot(t, q/1000)
# -
3*V_pos*I_pos
3*V_neg*I_neg
s_012 = 3*V_pos*I_pos + 3*V_neg*I_neg
s_012
sin_012 = 3*V_pos*I_neg + 3*V_neg*I_pos
cos_012 = 3*V_pos*I_neg - 3*V_neg*I_pos
print(sin_012,cos_012)
# +
s_012,sin_012,cos_012,V_pos,I_pos,V_neg,I_neg = sym.symbols('s_012,sin_012,cos_012,V_pos,I_pos,V_neg,I_neg ')
sin_012_ = 3*V_pos*I_neg + 3*V_neg*I_pos
cos_012_ = 3*V_pos*I_neg - 3*V_neg*I_pos
eq1 = -s_012 + 3*V_pos*I_pos + 3*V_neg*I_neg
eq2 = sin_012-sin_012_ - cos_012+cos_012_
sym.solve([eq1,eq2],[I_pos,I_neg])
# -
I_pos
# ## Control Fisix
# +
from sympy.functions import re,im
v_d_p,v_q_p,v_d_n,v_q_n,i_d_p,i_q_p,i_d_n,i_q_n,wt = sym.symbols('v_d_p,v_q_p,v_d_n,v_q_n,i_d_p,i_q_p,i_d_n,i_q_n,wt',real=True)
p_ref,q_ref = sym.symbols('p_ref,q_ref',real=True)
exp_p = sym.cos( wt)+1j*sym.sin( wt)
exp_n = sym.cos(-wt)+1j*sym.sin(-wt)
v_dq_p = v_q_p + 1j*v_d_p
v_dq_n = v_q_n + 1j*v_d_n
i_dq_p = i_q_p + 1j*i_d_p
i_dq_n = i_q_n + 1j*i_d_n
s = 3/2*(v_dq_p*exp_p + v_dq_n*exp_n)*sym.conjugate(i_dq_p*exp_p + i_dq_n*exp_n)
s = sym.simplify(sym.factor(sym.expand(s)))
# -
p = sym.collect(re(s),[sym.cos(2*wt),sym.sin(2*wt)])
q = sym.collect(im(s),[sym.cos(2*wt),sym.sin(2*wt)])
p_cos = p.diff(sym.cos(2*wt))
p_sin = p.diff(sym.sin(2*wt))
p_cte = sym.simplify(p - p_cos*sym.cos(2*wt) - p_sin*sym.sin(2*wt))
q_cos = q.diff(sym.cos(2*wt))
q_sin = q.diff(sym.sin(2*wt))
q_cte = sym.simplify(q - q_cos*sym.cos(2*wt) - q_sin*sym.sin(2*wt))
lhs = ['p_cte','p_cos','p_sin','q_cte','q_cos','q_sin']
rhs = [p_cte,p_cos,p_sin,q_cte,q_cos,q_sin]
for lh,rh in zip(lhs,rhs):
print('{:s}_ref = {:s}'.format(str(lh) ,str(sym.simplify(rh))))
# ### References for p constant
sol = sym.solve([p_cte-p_ref,q_cte-q_ref,p_cos,p_sin],[i_d_p,i_q_p,i_d_n,i_q_n])
for item in [i_d_p,i_q_p,i_d_n,i_q_n]:
print('{:s}_ref = {:s}'.format(str(item) ,str(sym.simplify(sol[item]))))
# ### References for q constant
sol = sym.solve([p_cte-p_ref,q_cte-q_ref,q_cos,q_sin],[i_d_p,i_q_p,i_d_n,i_q_n])
for item in [i_d_p,i_q_p,i_d_n,i_q_n]:
print('{:s}_ref = {:s}'.format(str(item) ,str(sym.simplify(sol[item]))))
# ### References for p and q constant
sol = sym.solve([p_cte-p_ref,q_cte-q_ref,p_cos,q_sin],[i_d_p,i_q_p,i_d_n,i_q_n])
for item in [i_d_p,i_q_p,i_d_n,i_q_n]:
print('{:s}_ref = {:s}'.format(str(item) ,str(sym.simplify(sol[item]))))
sym.simplify(p_cos-q_sin)
sym.simplify(p_sin-q_cos)
# ### Lipo
# +
import sympy as sym
from sympy.functions import re,im
v_d_p,v_q_p,v_d_n,v_q_n,i_d_p,i_q_p,i_d_n,i_q_n,wt = sym.symbols('v_d_p,v_q_p,v_d_n,v_q_n,i_d_p,i_q_p,i_d_n,i_q_n,wt',real=True)
p_ref,q_ref = sym.symbols('p_ref,q_ref',real=True)
exp_p = sym.cos( wt)+1j*sym.sin( wt)
exp_n = sym.cos(-wt)+1j*sym.sin(-wt)
v_dq_p = v_d_p + 1j*v_q_p
v_dq_n = v_d_n + 1j*v_q_n
i_dq_p = i_d_p + 1j*i_q_p
i_dq_n = i_d_n + 1j*i_q_n
s = 3/2*(exp_p*v_dq_p + exp_n*v_dq_n)*sym.conjugate(exp_p*i_dq_p + exp_n*i_dq_n)
s = sym.simplify(sym.factor(sym.expand(s)))
t = 3/2*(-1j*exp_p*v_dq_p + 1j*exp_n*v_dq_n)*sym.conjugate(exp_p*i_dq_p + exp_n*i_dq_n)
t = sym.simplify(sym.factor(sym.expand(t)))
p = sym.collect(re(s),[sym.cos(2*wt),sym.sin(2*wt)])
q = sym.collect(re(t),[sym.cos(2*wt),sym.sin(2*wt)])
p_cos = p.diff(sym.cos(2*wt))
p_sin = p.diff(sym.sin(2*wt))
p_cte = sym.simplify(p - p_cos*sym.cos(2*wt) - p_sin*sym.sin(2*wt))
q_cos = q.diff(sym.cos(2*wt))
q_sin = q.diff(sym.sin(2*wt))
q_cte = sym.simplify(q - q_cos*sym.cos(2*wt) - q_sin*sym.sin(2*wt))
lhs = ['p_cte','p_cos','p_sin','q_cte','q_cos','q_sin']
rhs = [p_cte,p_cos,p_sin,q_cte,q_cos,q_sin]
for lh,rh in zip(lhs,rhs):
print('{:s}_ref = {:s}'.format(str(lh) ,str(sym.simplify(rh))))
# -
p
sol = sym.solve([p_cte-p_ref,q_cte-q_ref,p_cos,p_sin],[i_d_p,i_q_p,i_d_n,i_q_n])
for item in [i_d_p,i_q_p,i_d_n,i_q_n]:
print('{:s}_ref = {:s}'.format(str(item) ,str(sym.simplify(sol[item]))))
q
p_cte_ref = 1.5*i_d_n*v_d_n + 1.5*i_d_p*v_d_p + 1.5*i_q_n*v_q_n + 1.5*i_q_p*v_q_p
p_cos_ref = 1.5*i_d_n*v_d_p + 1.5*i_d_p*v_d_n + 1.5*i_q_n*v_q_p + 1.5*i_q_p*v_q_n
p_sin_ref = -1.5*i_d_n*v_q_p + 1.5*i_d_p*v_q_n + 1.5*i_q_n*v_d_p - 1.5*i_q_p*v_d_n
q_cte_ref = -1.5*i_d_n*v_q_n + 1.5*i_d_p*v_q_p + 1.5*i_q_n*v_d_n - 1.5*i_q_p*v_d_p
q_cos_ref = 1.5*i_d_n*v_q_p - 1.5*i_d_p*v_q_n - 1.5*i_q_n*v_d_p + 1.5*i_q_p*v_d_n
q_sin_ref = 1.5*i_d_n*v_d_p + 1.5*i_d_p*v_d_n + 1.5*i_q_n*v_q_p + 1.5*i_q_p*v_q_n
# +
## Lipo con dq según fisix
# +
import sympy as sym
from sympy.functions import re,im
v_d_p,v_q_p,v_d_n,v_q_n,i_d_p,i_q_p,i_d_n,i_q_n,wt = sym.symbols('v_d_p,v_q_p,v_d_n,v_q_n,i_d_p,i_q_p,i_d_n,i_q_n,wt',real=True)
p_ref,q_ref = sym.symbols('p_ref,q_ref',real=True)
exp_p = sym.cos( wt)+1j*sym.sin( wt)
exp_n = sym.cos(-wt)+1j*sym.sin(-wt)
v_dq_p = v_q_p + 1j*v_d_p
v_dq_n = v_q_n + 1j*v_d_n
i_dq_p = i_q_p + 1j*i_d_p
i_dq_n = i_q_n + 1j*i_d_n
s = 3/2*(exp_p*v_dq_p + exp_n*v_dq_n)*sym.conjugate(exp_p*i_dq_p + exp_n*i_dq_n)
s = sym.simplify(sym.factor(sym.expand(s)))
t = 3/2*(-1j*exp_p*v_dq_p + 1j*exp_n*v_dq_n)*sym.conjugate(exp_p*i_dq_p + exp_n*i_dq_n)
t = sym.simplify(sym.factor(sym.expand(t)))
p = sym.collect(re(s),[sym.cos(2*wt),sym.sin(2*wt)])
q = sym.collect(re(t),[sym.cos(2*wt),sym.sin(2*wt)])
p_cos = p.diff(sym.cos(2*wt))
p_sin = p.diff(sym.sin(2*wt))
p_cte = sym.simplify(p - p_cos*sym.cos(2*wt) - p_sin*sym.sin(2*wt))
q_cos = q.diff(sym.cos(2*wt))
q_sin = q.diff(sym.sin(2*wt))
q_cte = sym.simplify(q - q_cos*sym.cos(2*wt) - q_sin*sym.sin(2*wt))
lhs = ['p_cte','p_cos','p_sin','q_cte','q_cos','q_sin']
rhs = [p_cte,p_cos,p_sin,q_cte,q_cos,q_sin]
for lh,rh in zip(lhs,rhs):
print('{:s}_ref = {:s}'.format(str(lh) ,str(sym.simplify(rh))))
# -
sol = sym.solve([p_cte-p_ref,q_cte-q_ref,p_cos,p_sin],[i_d_p,i_q_p,i_d_n,i_q_n])
for item in [i_d_p,i_q_p,i_d_n,i_q_n]:
print('{:s}_ref = {:s}'.format(str(item) ,str(sym.simplify(sol[item]))))
# +
Clark = sym.Matrix([[1.0/sym.sqrt(2.0),1.0/sym.sqrt(2.0),1.0/sym.sqrt(2.0)],[1.0,-1.0/2.0,-1.0/2.0],[0,-sym.sqrt(3.0)/2.0,sym.sqrt(3.0)/2.0]])
# -
import numpy as np
Clark = 2/3*np.array([[1/np.sqrt(2), 1/np.sqrt(2),1/np.sqrt(2)],
[ 1, -0.5, -0.5],
[ 0,-np.sqrt(3)/2,np.sqrt(3)/2]])
inv_Clark = np.linalg.inv(Clark)
pasar al tiempo con seq. pos y neg
inv_Clark
|
models/vsc_phasor_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-tf-2-gpu] *
# language: python
# name: conda-env-.conda-tf-2-gpu-py
# ---
# +
import sys, os
# Insert the PATH to the directory of this notebook here
# ROOT_PATH must end with /
ROOT_PATH = None
sys.path.append(ROOT_PATH)
if ROOT_PATH is None:
raise Exception("Please specify the ROOT_PATH")
import experiment
import tensorflow as tf
import numpy as np
import imp
experiment = imp.reload(experiment)
run_experiment = experiment.run_experiment
exp_prefix = 'protein'
seqlen = 500
n_seqs = 512
n_emissions = 19
em_lr = 0.01
n_gt_iter = 10
nl_dict = {3: [1, 2, 3], 5: [2, 3, 5], 10: [2, 3, 5, 10]}
gt_seqs = dict()
experiment_directories = dict()
failed_exps = []
for n_hidden_states in nl_dict.keys():
for l in nl_dict[n_hidden_states]:
for i in range(n_gt_iter):
exp_params = {'dataset_ident': 'protein',
'dataset_params': {'len_cut': seqlen, 'total_perc': 0.002},
'train_perc': 0.5,
'standard_params' : {'n_hidden_states': n_hidden_states,
'em_iter': 40
},
'standard_log_config': {'gamma_after_estep': False,
'gamma_after_mstep': False,
'samples_after_estep': True,
'samples_after_mstep': True,
},
'dense_params': {'verbose': False,
'em_iter': 40,
'n_hidden_states': n_hidden_states,
'mstep_config': {'l_uz':l,
'l_vw': l,
'scaling': l*n_emissions*n_hidden_states*seqlen*n_seqs,
'trainables': 'uvwzz0',
'em_lr': em_lr,
'initializer': tf.initializers.random_normal(0., 1.),
'em_optimizer': tf.train.AdamOptimizer(em_lr),
'em_epochs': 25,
'cooc_epochs': 10000,
'cooc_optimizer': tf.train.AdamOptimizer(0.001),
}
},
'dense_log_config': {'gamma_after_estep': False,
'gamma_after_mstep': False,
'samples_after_estep': True,
'samples_after_mstep': True,
},
'dense_opt_schemes': ('em','cooc'),
'compare_to_fair_standard': True,
'fair_standard_log_config': {'gamma_after_estep': False,
'gamma_after_mstep': False,
'samples_after_estep': True,
'samples_after_mstep': True,
}
}
try:
directory = run_experiment('dataset_sequences', '%s/protein_i=%d_n=%d_m=%d_l=%d' % (exp_prefix, i, n_hidden_states, n_emissions, l),
exp_params)
experiment_directories[(i, n_hidden_states, l)] = directory
except Exception as e:
print(e)
failed_exps.append((i, n_hidden_states, l))
np.save('%s%s/experiment_directories' % (ROOT_PATH, exp_prefix), experiment_directories)
|
code_dense_hmm/start_protein_experiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
import sys
import os
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
# +
project_path = os.path.abspath(os.path.join('..'))
if project_path not in sys.path:
sys.path.append(f'{project_path}/src/visualizations/')
from covid_data_viz import CovidDataViz
# -
# # Goal
# My goal is to visualize various aspect of the `COVID-19` pandemic.
# # Data sources
#
# In this project I use data from the following sources:
# - https://github.com/CSSEGISandData/COVID-19 - JHU CSSE COVID-19 Data.
# - https://datahub.io/JohnSnowLabs/country-and-continent-codes-list - country codes and continents.
# # Data loading
cdv = CovidDataViz()
# # Fancy plot
#
# Visual for repo readme.
def create_plot(width, height, dpi, period, step,
fontsize, fontfamily, n_clabels,
countries, suffix, spinewidth,
titleweight='normal'):
# Data prep
df = cdv.data['Confirmed chg'][countries].copy()
df = df.rename(columns={'United Kingdom': 'UK'})
countries = df.columns.to_list()
df = df.rolling(period)
df = df.mean()
df = df.dropna()
df = df.to_numpy()
df = df.astype(float)
df = df.transpose()
df = np.sqrt(df)
# Plot
size = (width / dpi, height / dpi)
plt.figure(figsize=size, dpi=dpi)
plt.imshow(df, aspect='auto', interpolation='nearest')
plt.set_cmap('hot')
# Plot pines
ax = plt.gca()
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(spinewidth)
# Plot labels
xticks = range(df.shape[1])[::step]
xlabels = list(cdv.data['Confirmed chg']['Date'])[period:]
xlabels = [x.strftime(format='%Y-%m') for x in xlabels]
xlabels = xlabels[::step]
yticks = range(len(countries))
ylabels = countries
plt.yticks(ticks=yticks, labels=ylabels, fontsize=fontsize,
family=fontfamily, verticalalignment='center')
plt.xticks(ticks=xticks, labels=xlabels, rotation=45,
fontsize=fontsize, family=fontfamily,
horizontalalignment='center')
ax.tick_params(width=spinewidth, color='black')
# Colorbar
cticks = np.round(np.linspace(0, np.max(df), 6), -1)
cticks = cticks.astype(np.int)
clabels = np.power(cticks, 2)
cticks = sorted(set(cticks))
clabels = np.power(cticks, 2)
clabels = [int((round(x, -3)) / 1000) for x in clabels]
clabels = [str(x) + 'k' for x in clabels]
cbar = plt.colorbar()
cbar.set_ticks(cticks)
cbar.set_ticklabels(clabels)
cbar.ax.tick_params(labelsize=fontsize, width=spinewidth)
for l in cbar.ax.yaxis.get_ticklabels():
l.set_family(fontfamily)
cbar.outline.set_linewidth(spinewidth)
plt.title('New COVID-19 cases', fontsize=fontsize + 1,
family=fontfamily, weight=titleweight)
plt.tight_layout()
plt.savefig(fname=f'../img/covid_tiles_{suffix}.png',
bbox_inches='tight')
plt.show()
# +
countries = ['Germany',
'France',
'Italy',
'Spain',
'United Kingdom',
'Russia',
'India',
'Brazil',
'US',
'Poland',
'Mexico']
countries = sorted(countries)
# -
create_plot(width=625,
height=375,
dpi=200,
period=7,
step=30,
fontsize=4.5,
fontfamily='serif',
n_clabels=6,
countries=countries,
suffix='portfolio_readme',
spinewidth=0.5)
create_plot(width=1000,
height=600,
dpi=300,
period=7,
step=30,
fontsize=3,
fontfamily='serif',
n_clabels=6,
countries=countries,
suffix='project_readme',
spinewidth=1/3,
titleweight='bold')
|
covid-19/notebooks/1.3-exploratory_analysis_fancy_plot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # An analysis of imdb scores as it relates to TV Series longevity.
#
# The audience for this analysis is content creators, programmers, and deal makers. If one can find a relationship between imdb and series longevity, this data can inform greenlight decisions and save potentially millions of dollars on rights and production costs.
#
# To shed light into whether this data can inform decision makeers, it aims to answer the following questions:
# * Is there a relationship between imdb scores and a television series age?
# * Is there a relationship between a greater than average rating early in a series and its age?
# * Is there a relationship between imdb scores and a series ending over time?
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(rc={'figure.figsize':(11.7,8.27)})
# -
# ## ETL
# The three files provided by imdb are titles, episodes, and ratings. This section imports the file, performs transformations and
# saves the files. Use the transformed files in the analysis to save processing time.
# * Titles file (title.basics.tsv) contains movies and television title data identified by tconst.
# * Episode file (title.episode.tsv) contains episode data that links to series via the parentTconst
# * Ratings file (title.ratings.tsv) contains the ratings for each title linked by tconst
# There is an anomaly in this file that requires use of the python engine instead of C
titles = pd.read_csv('./title.basics.tsv', delimiter='\t+', encoding='ISO-8859-1', engine='python')
episodes = pd.read_csv('./title.episode.tsv', delimiter='\t', encoding='ISO-8859-1')
ratings = pd.read_csv('./title.ratings.tsv', delimiter='\t', encoding='ISO-8859-1')
# For each file:
# * Change the imdb null value to pandas null value
# * Convert years and minutes to numeric types
titles = titles.replace('\\N',np.nan)
titles[["startYear",
"endYear",
"runtimeMinutes"]] = titles[["startYear",
"endYear",
"runtimeMinutes"]].apply(pd.to_numeric)
episodes = episodes.replace('\\N',np.nan)
ratings = ratings.replace('\\N', np.nan)
# The group by is used to get the value for tv series and episodes. Then the data is split between series and episodes. I filtered the series by 1990 and newer to have a more manageably size dataset. I chose 1990 roughly as the Seignfeld epic, but any epic can be used (e.g., Cheers).
# Get list of title types to select Programs/Series and Episodes
titles[["tconst", "titleType"]].groupby('titleType').count()
tvSeries1990 = titles.loc[ (titles['titleType'] == "tvSeries") & (titles['startYear'] >= 1990)]
tvEpisodes = titles.loc[(titles['titleType'] == "tvEpisode")]
ratings.head()
# Join TV series data with the corresponding ratings
tvSeries1990 = tvSeries1990.merge(ratings, how='left', on='tconst')
# Turn genres column, a comma separated list of genres, into a dummy binary categorical variable
tvSeries1990 = pd.concat([tvSeries1990,tvSeries1990['genres'].str.get_dummies(sep=',')])
tvSeries1990.head()
# Merge the TV Episode dataset from the titles file with the episode data in the episodes file. This gives a complete picture of the episodes
tvEpisodesFull = tvEpisodes.merge(episodes, how='inner', on='tconst')
tvEpisodesFull.head()
# Merge episode data with the parent series data to have a data that includes the series and episode data in one. Merge the ratings data. Yes, I got lazy and didn't rename the \_x and \_y fields
tvEpisodesFull2 = tvEpisodesFull.merge(
tvSeries1990,
how='inner',
left_on='parentTconst',
right_on='tconst')
tvEpisodesFull2 = tvEpisodesFull2[["tconst_x",
"titleType_x",
"primaryTitle_x",
"originalTitle_x",
"isAdult_x",
"startYear_x",
"endYear_x",
"runtimeMinutes_x",
"genres_x",
"parentTconst",
"seasonNumber",
"episodeNumber"]]
tvEpisodesFull2 = tvEpisodesFull2.merge(ratings, how='left', left_on='tconst_x', right_on='tconst')
tvEpisodesFull2 = pd.concat([tvEpisodesFull2, tvEpisodesFull2['genres_x'].str.get_dummies(sep=',')])
tvEpisodesFull2.head()
# Save the prepared data assets as new csvs. Use these when preforming the analysis.
tvSeries1990.to_csv('tvSeries1990.csv', index=True)
tvEpisodesFull2.to_csv('tvEpisodes.csv', index=True)
# ## Data Analysis
tvSeries1990 = pd.read_csv('tvSeries1990.csv')
tvEpisodesFull2 = pd.read_csv('tvEpisodes.csv')
tvEpisodesFull2.loc[tvEpisodesFull2['parentTconst'] == 'tt0298685'].sort_values(['seasonNumber', 'episodeNumber'])
# ### Question: What is the distribution of number of seasons per series title?
# Answer: There is a long tail of number of seasons with an elbow at 50 seasons. There a couple of House Hunters series (tt0795129,tt0369117) that have an anomalous amount of seasons for the number of years it has been on the air. A google search reveals the longest running program is the Tonight Show with almost 70 seasons https://www.mentalfloss.com/article/646554/longest-running-tv-shows.
# +
uniqueSeasons = (tvEpisodesFull2.loc[tvEpisodesFull2['parentTconst'] != 'tt0369117']
.groupby('parentTconst')
.agg(unique_seasons=('seasonNumber','nunique'))
)
(uniqueSeasons
.sort_values(by='unique_seasons', ascending=False)
.head(100)
.plot(figsize=(12,3))
.bar(color='b')
)
# -
# Cut at the elbow
sns.boxplot(y = 'unique_seasons',
data = uniqueSeasons.loc[uniqueSeasons["unique_seasons"] <= 60])
(uniqueSeasons.loc[uniqueSeasons['unique_seasons'] <= 30]
.sort_values(by='unique_seasons', ascending=False)
.head(500)
.plot(figsize=(12,3))
.bar(color='b')
)
#tvEpisodesFull2.loc[tvEpisodesFull2['parentTconst'] == 'tt0369117'].sort_values(by='seasonNumber', ascending=False)
tvEpisodesFull2.loc[tvEpisodesFull2['parentTconst'] == 'tt0795129'].sort_values(by='seasonNumber', ascending=False)
#tvSeries1990.loc[tvSeries1990['tconst'] == 'tt0369117']
# ### Question: Is there a relationship between TV Series age and imdb rating?
# Answer: This analysis does not show any conclusive relationship; however, the range does tighten. That being said, the boxplot shows perhaps a decrease in ratings as the the series ages.
# Calculate the age of the series title from start date to end date. If there is no end date assume it is still on the air and use current year
tvSeries1990['age'] = tvSeries1990.apply(
lambda x: 2021 - x['startYear'] if x['endYear'] == np.nan else x['endYear'] - x['startYear'],
axis=1)
sns.boxplot(y = 'age',
data = tvSeries1990[['age']])
# Not surprisingly, there is no relationship between age and rating
tvSeries1990[['averageRating','age']].plot.scatter(x='age', y='averageRating')
sns.set(rc={'figure.figsize':(11.7,8.27)})
sns.boxplot(x = 'age', y = 'averageRating',
data = tvSeries1990[['age',
'averageRating']])
# ### Question: Do TV Series that receive a higher than average rating in seasons 1 and/or 2 have a longer life?
# Answer: This analysis does not reveal a relationship much like the previous question. However, the very long running shows show higher than average 1st and 2nd season ratings.
#
# Todo:
# * A regression analysis can reveal a causual relationship.
# * More complicated season aggregation comparisons (for example: season 1 + season 2)
# +
group_by_season = (tvEpisodesFull2
.groupby(['parentTconst','seasonNumber'])
.agg(avg_rating_by_season=('averageRating','mean'))
#.reset_index()
#.head()
[['avg_rating_by_season']]
)
group_by_season = group_by_season.reset_index()
group_by_season.head()
# -
grp_season_age=group_by_season.merge(
tvSeries1990[["tconst","age"]],
how='inner',
left_on='parentTconst',
right_on='tconst')
grp_season_age[grp_season_age['age'].isna()][['parentTconst']]
(grp_season_age[grp_season_age['seasonNumber']=='1'][['avg_rating_by_season','age']]
.plot.scatter(x='age', y='avg_rating_by_season')
)
(grp_season_age[grp_season_age['seasonNumber']=='2'][['avg_rating_by_season','age']]
.plot.scatter(x='age', y='avg_rating_by_season')
)
sns.boxplot(x='age', y='avg_rating_by_season',
data=grp_season_age[grp_season_age['seasonNumber']=='1'][['avg_rating_by_season','age']])
sns.boxplot(x='age', y='avg_rating_by_season',
data=grp_season_age[grp_season_age['seasonNumber']=='2'][['avg_rating_by_season','age']])
# ### Question: Is there a relationship between imdb scores and a series ending over time?
# Answer: From this analysis the answer is inconclusive; however, further analysis could reveal a relationship. Keep executing this cell to plot the ratings by season for a random 20 TV series.
#
# Todo:
# * Filter out series that are very short (e.g., < 5 seasons).
# * Filter out series that are missing many values.
# +
grp_season_age_pivot = pd.pivot_table(
grp_season_age[['avg_rating_by_season','parentTconst','seasonNumber']],
values='avg_rating_by_season',
index='seasonNumber',
columns='parentTconst',
aggfunc=np.mean)
grp_season_age_pivot.index = pd.to_numeric(
grp_season_age_pivot.index,
errors='coerce')
grp_season_age_pivot = grp_season_age_pivot.sort_index()
import random
rand_start = random.randrange(0, len(grp_season_age_pivot.columns)-20)
fig = sns.lineplot(data=grp_season_age_pivot.iloc[0:50, rand_start:rand_start+20]) # limit to 50 seasons
plt.xlabel("Season Number")
plt.ylabel("Rating")
plt.title("Ratings by Season")
plt.show(fig)
# -
# ## Acknowledgements
# * https://gist.github.com/Towhidn/ea3f8ad18116a2203d14b44620fe8d21
# * https://datasets.imdbws.com/
|
Python Data Analysis Project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Oinarrizko datu sarrera eta irteera
#
# Aurreko ataletan ikusi dugunez, `print()` funtzioa pantailatik informazioa erakusteko erabiliko dugu. Atal honetan, sarrera-irteerako oinarrizko beste funtzio bat ere aztertuko dugu, `input()` funtzioa hain zuzen ere (bi funtzio hauek eta aurreko ataletan ikusitako beste guztiak, Python-en [*Built-in Functions*](https://docs.python.org/3/library/functions.html) multzoan definitzen dira).
# + [markdown] slideshow={"slide_type": "slide"}
# ## `print()` funtzioa
#
# Informazioa testu moduan erakusteko balio duen funtzioa da. Objektu sorta bat jaso eta bakoitzaren balioa testu moduan erakutsiko du:
# + slideshow={"slide_type": "-"}
a = 1
b = 3.4
c = "kaixo"
print(a,b,c)
# + [markdown] slideshow={"slide_type": "-"}
# `print()` funtzioari ematen dizkiogun objektuak karaktere kate bilakatuko dira `str()` funtzioaren bidez, ondoren testu hoiek pantailatik erakusteko. Funtzioak objektuak jasoko dituela esan badugu ere, funtzioa erabiltzean espresioak erabil ditzakegu (funtzioari emango zaion objektua, espresioaren emaitza izango da):
# + slideshow={"slide_type": "-"}
print(a*4, b>=2, c+"?")
# + [markdown] slideshow={"slide_type": "slide"}
# Python-eko funtzioen argumentuek defektuzko balioak izan ditzakete. `print()` funtzioak halako lau argumentu ditu, bere konportamoldea aldatzeko erabil daitezkeenak (ezer adierazi ezean, defektuzko balioa izango dute):
# + slideshow={"slide_type": "-"}
help(print)
# + [markdown] slideshow={"slide_type": "-"}
# * `sep` → balioen artean gehitutako karattere katea (defektuz, hutsunea).
# * `end` → amaieran gehitutako karaktere katea (defektuz, lerro berri bat).
# * `file` → *non* idatzi (defektuz, irteera estandarra).
# * `flush` → *flushing*-a behartu ala ez.
# + [markdown] slideshow={"slide_type": "slide"}
# `print()` funtzioa bitxia da, argumentu kopuru mugagabea duelako. Hau dela eta `sep`, `end`, `file` edo ta `flush` argumentuei beste balio bat ematea nahi badugu, beren izena erabili beharko dugu (hau beti egin daiteke):
# + slideshow={"slide_type": "-"}
print(a*4, b>=2, c+"?", sep=" <--> ", end="\nTHE END\n")
print("EPA")
# + [markdown] slideshow={"slide_type": "-"}
# Argumentuen izena jartzeak edozein ordenetan adierazteko aukera ere ematen digu:
# + slideshow={"slide_type": "-"}
print(a*4, b>=2, c+"?", end="\nTHE END\n", sep=" <--> ")
# + [markdown] slideshow={"slide_type": "slide"}
# Izen bidez adierazitako argumentuei *keyword* argumentu deritzo, eta beti amaieran azaldu behar dira:
# + slideshow={"slide_type": "-"}
print(end="\nTHE END\n", sep=" <--> ", a*4, b>=2, c+"?")
# + [markdown] slideshow={"slide_type": "slide"}
# ## `input()` funtzioa
#
# `input()` funtzioak sistemaren sarrera estandarra erabiliko du, erabiltzailearengandik informazioa jaso ahal izateko teklatuaren bidez. Funtzoio honek exekuzioa geldiarazten du, erabiltzaileak *Return* (*Enter*) tekla sakatu arte. Orduan, erabiltzaileak idatzitako testua bueltatuko du
# + slideshow={"slide_type": "-"}
a = input()
print("Ados,",a,"idatzi duzu")
# + [markdown] slideshow={"slide_type": "slide"}
# `input()` funtzioak defektuzko `''` balioa duen `prompt` argumentua du:
# + slideshow={"slide_type": "-"}
help(input)
# + [markdown] slideshow={"slide_type": "-"}
# * `prompt` → pantailatik erakutsiko den mezua (defektuz, hutsa).
#
# Argumentu honekin, erabiltzaileari jakinaraziko diogu bere zain gaudela:
# + slideshow={"slide_type": "-"}
a = input("Idatzi balio bat: ")
print("Ados,",a,"idatzi duzu")
# + [markdown] slideshow={"slide_type": "slide"}
# Beti kontutan izan beharreko bi gauza:
# 1. `input()` funtzioak bueltatzen duenarekin **ZERBAIT** egin behar da (**gorde**, adibidez).
# 1. `input()` funtzioak **KARAKTERE KATE** bat bueltatzen du (**ez da zenbaki bat**).
# + slideshow={"slide_type": "-"}
a = input("Idatzi balio bat: ")
print("Jasotako", a, "balioa", type(a), "motakoa da")
print("a * 2 :" , a*2)
a = int(a)
print("Orain", a, "balioa", type(a), "motakoa da")
print("a * 2 :" , a*2)
# + [markdown] slideshow={"slide_type": "-"}
# <table border="0" width="100%" style="margin: 0px;">
# <tr>
# <td style="text-align:left"><a href="Eragileak.ipynb">< < Eragileak < <</a></td>
# <td style="text-align:right"><a href="Kontrol egiturak.ipynb">> > Kontrol egiturak > ></a></td>
# </tr>
# </table>
|
Gardenkiak/Programazioa/.ipynb_checkpoints/Oinarrizko datu sarrera eta irteera-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="qdJKkQrye4H6" outputId="8723b5b3-583d-4b1e-cff5-9da3f251adc5" colab={"base_uri": "https://localhost:8080/", "height": 35}
from numpy import load
import pickle
from google.colab import drive
drive.mount('/content/drive')
# + id="KvI2618ke-IV" outputId="4353c30a-d8ba-4913-8b6b-b3fddbf267d5" colab={"base_uri": "https://localhost:8080/", "height": 126}
train_X = load('/content/drive/My Drive/Dataset-FakeNews/train_X.npy')
train_X_new = load('/content/drive/My Drive/Dataset-FakeNews/train_X_new.npy')
train_Y = load('/content/drive/My Drive/Dataset-FakeNews/train_Y.npy')
val_X = load('/content/drive/My Drive/Dataset-FakeNews/val_X.npy')
val_X_new = load('/content/drive/My Drive/Dataset-FakeNews/val_X_new.npy')
val_Y = load('/content/drive/My Drive/Dataset-FakeNews/val_Y.npy')
print(train_X.shape)
print(train_X_new.shape)
print(train_Y.shape)
print(val_X.shape)
print(val_X_new.shape)
print(val_Y.shape)
# + id="GYRR_rw_ohcD" outputId="52a59ba6-f78a-4468-f4f5-59f3101bd34e" colab={"base_uri": "https://localhost:8080/", "height": 35}
print(train_X_new[0][0])
# + id="D2TliHdXkpTZ"
import seaborn as sn
import pandas as pd
import matplotlib.pyplot as plt
def plotCM(cMatrix, N,label = []):
df_cm = pd.DataFrame(cMatrix, range(N),range(N))
#plt.figure(figsize = (8,8))
plt.figure(figsize = (10,7))
cmap = sn.cm.rocket_r
if label:
sn.heatmap(df_cm.round(2), annot=True,fmt='g',cmap = cmap, xticklabels = label, yticklabels=label)
else:
sn.heatmap(df_cm.round(2), annot=True,fmt='g',cmap = cmap, )
# + id="uJM4rME5fzQ4"
from sklearn.linear_model import LogisticRegression
reg = LogisticRegression(max_iter=200).fit(train_X, train_Y,)
# + id="9B2yPWIMf6AM" outputId="2a4bd708-3d60-4425-a234-baa3a3e62578" colab={"base_uri": "https://localhost:8080/", "height": 53}
print("Train Accuracy = ", reg.score(train_X, train_Y))
print("Val Accuracy = ", reg.score(val_X, val_Y))
filename = '/content/drive/My Drive/AAAI CONSTRAINT/Fake News/W2V-LR.pickle'
pickle.dump(reg, open(filename, 'wb'))
# + id="SCy_GSDVjby1" outputId="aca319d6-8502-4e38-8d5a-90a99819f428" colab={"base_uri": "https://localhost:8080/", "height": 428}
pred = reg.predict(val_X)
N = 2 # num_classes
confMatrix = [ [0 for y in range(N)] for x in range(N)]
for x,y in zip(val_Y,pred):
confMatrix[x][y] += 1
plotCM(confMatrix,N,['real','fake'])
# + id="939aRB8sVWFG" outputId="66b49d11-b464-4f66-c9cf-093baf2b7f76" colab={"base_uri": "https://localhost:8080/", "height": 182}
from sklearn.metrics import classification_report
y_true = val_Y
y_pred = pred
target_names = ['Real','Fake']
print(classification_report(y_true, y_pred, target_names=target_names))
# + id="KI0WtO2Mf7mN" outputId="b3efb56c-61b2-4dd0-8952-d4b5f31f90af" colab={"base_uri": "https://localhost:8080/", "height": 201}
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.svm import SVC
clf = make_pipeline(StandardScaler(), SVC(gamma='auto'))
clf.fit(train_X, train_Y)
# + id="t39etyL4f94_" outputId="5def9fa3-91a4-44ab-e0ca-37a2d38bc480" colab={"base_uri": "https://localhost:8080/", "height": 54}
print("Train Accuracy = ",clf.score(train_X,train_Y))
print("Val Accuracy = ", clf.score(val_X,val_Y))
filename = '/content/drive/My Drive/AAAI CONSTRAINT/Fake News/W2V-SVM.pickle'
pickle.dump(clf, open(filename, 'wb'))
# + id="REHsHg2XmaSI" outputId="524465d0-056f-4a22-95c4-518a89cd9cec" colab={"base_uri": "https://localhost:8080/", "height": 428}
pred = clf.predict(val_X)
N = 2 # num_classes
confMatrix = [ [0 for y in range(N)] for x in range(N)]
for x,y in zip(val_Y,pred):
confMatrix[x][y] += 1
plotCM(confMatrix,N,['real','fake'])
arr = []
for i in range(len(pred)):
if pred[i] != val_Y[i]:
arr.append(i)
# + id="o8ZBF5zmVclL" outputId="eae2b87b-8568-4df0-eaa2-935a44a506e3" colab={"base_uri": "https://localhost:8080/", "height": 182}
from sklearn.metrics import classification_report
y_true = val_Y
y_pred = pred
target_names = ['Real','Fake']
print(classification_report(y_true, y_pred, target_names=target_names))
# + id="jo1y_BKUf_Lq" outputId="5a4c228c-48f7-4b17-99b1-c0f8d45920a9" colab={"base_uri": "https://localhost:8080/", "height": 164}
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(max_depth=11, random_state=1, n_estimators=100)
clf.fit(train_X, train_Y)
# + id="igC8qELqgAho" outputId="6a25b080-e6ff-4a82-913c-13844a930371" colab={"base_uri": "https://localhost:8080/", "height": 54}
print("Train Accuracy = ",clf.score(train_X,train_Y))
print("Val Accuracy = ", clf.score(val_X,val_Y))
filename = '/content/drive/My Drive/AAAI CONSTRAINT/Fake News/W2V-RF.pickle'
pickle.dump(clf, open(filename, 'wb'))
# + id="kRBjZGSim5xk" outputId="8a6e49ef-5fd2-474b-d759-af21889a355e" colab={"base_uri": "https://localhost:8080/", "height": 428}
pred = clf.predict(val_X)
N = 2 # num_classes
confMatrix = [ [0 for y in range(N)] for x in range(N)]
for x,y in zip(val_Y,pred):
confMatrix[x][y] += 1
plotCM(confMatrix,N,['real','fake'])
# + id="MRZ7U8FO4B95" outputId="c37526d4-1343-407a-b1c3-ae1e8f27cdef" colab={"base_uri": "https://localhost:8080/", "height": 182}
from sklearn.metrics import classification_report
y_true = val_Y
y_pred = pred
target_names = ['Real','Fake']
print(classification_report(y_true, y_pred, target_names=target_names))
# + id="jNbdSl8OFD7r"
train_path = '/content/drive/My Drive/AAAI CONSTRAINT/Fake News/Constraint_English_Train - Sheet1.csv'
val_path = '/content/drive/My Drive/AAAI CONSTRAINT/Fake News/Constraint_English_Val - Sheet1.csv'
# + id="QRgzvW1MFpWP" outputId="736fe54c-fba5-44f8-f224-f55d9c8e2e6e" colab={"base_uri": "https://localhost:8080/", "height": 206}
val_df = pd.read_csv(val_path)
val_df.head()
# + id="rCdDYwW_FtKu" outputId="3d214fd5-8e05-4eee-aa1d-f0ab8b86c9a8" colab={"base_uri": "https://localhost:8080/", "height": 1000}
mis = []
for x in arr:
mis.append((val_df['label'][x],val_df['tweet'][x]))
print(arr)
for x in mis:
print(x[0], [x[1]])
# + id="oZwPU6gjIMUg"
# + [markdown] id="kUjqZvykMRNd"
# Many Tweets had URLs, on inspection -
#
# * images or videos attached came as url on opening it takes you to the tweet
# * many tweets had 2 urls, 1 had tweet image/video other had an article outside twitter(news,blog etc)
# * In a lot of cases truth of tweet is determined by the url example fake - 57, 28, real - 67
#
# Also among the misclassified tweets i decided to search with keywords. Searching for Virus found 3 misclassified all are real classified as fake. There are other similar keywords mainly breath, cough, symptoms along with covid.
# When searching for similar tweets in the dataset i found lot of fake tweets talking about breath test which had similar keywords. Where as, lot of real tweets had the keyword mask in it. combination of these words without context is causing the issue.
# + id="uT7z-QQUMXAQ"
|
Fake News/Notebooks/Fake_News_Results.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Python Flujos de Control
#
# Hasta ahora hemos visto cómo ejecutar un programa secuencialmente, pero ¿y si queremos que cambien los outputs del programa en función de ciertas condiciones, o si queremos que tome otros caminos en caso de encontrar errores?. Todo esto lo podremos hacer con los flujos de control. Sentencias que encontrarás en todos los lenguajes de programación.
#
# 1. [Sintaxis de línea](#1.-Sintaxis-de-línea)
# 1. [if/elif/else](#2.-if/elif/else)
# 2. [Bucle for](#3.-Bucle-for)
# 3. [Bucle while](#4.-Bucle-while)
# 4. [Break/continue](#5.-Break/continue)
# 5. [Try/except](#6.-Try/except)
# 10. [Resumen](#7.-Resumen)
# ## 1. Sintaxis de línea
# La manera en la que Python encapsula todo el código que va dentro de un flujo de control como `if` o `for` es diferente a como se suele hacer en otros lenguajes, donde se declara el flujo de control, y todo lo que va dentro se rodea de llaves `{}` o paréntesis `()`. Con Python no. En Python simplemente hay que añadir una tabulación a cada línea de código que vaya dentro del flujo de control.
#
#
# > ```Python
# > for condiciones:
# > Código dentro de este bucle
# > ```
#
#
# Si lo dejamos fuera, este código se ejecutará secuencialmente después de que corra el for
#
# > ```Python
# > for condiciones:
# >
# > Código fuera de este bucle
# > ```
#
# Forma genérica:
# > ```Python
# > for condiciones:
# > 1ª línea código dentro de este bucle
# > 2ª línea código dentro de este bucle
# > 3ª línea código dentro de este bucle
# > ...
# > última línea código dentro de este bucle
# >
# > Código fuera de este bucle (esta línea no se ejecutará dentro de este bucle, sino al salir de él)
# > ```
#
# Veamos un ejemplo. Tenemos una lista de numeros, y queremos ver cuales son enteros. Para ello los recorremos con un `for` (vermos más en profundiad en este notebook). Vamos iternando uno a uno cada elemento. Luego mediante un `if` comprobamos si es entero. Fíjate que todo lo que va dentro del `for` lleva una tabulación y lo que va dentro del `if` lleva dos tabulaciones, ya que sus sentencias van tanto dentro del `if`, como dentro del `for`.
# +
numeros = [4,6,8.0,10.0]
for num in numeros:
if type(num) == int:
print("El numero", num, "es un entero")
print("Estoy en el bucle")
# -
# <table align="left">
# <tr><td width="80"><img src="./error.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>ERRORES ¿Qué ocurre si nos olvidamos de tabular?</h3>
#
# </td></tr>
# </table>
# +
numeros = [4,6,8.0,10.0]
for num in numeros:
if type(num) == int:
print("El numero", num, "es un entero")
# -
# Ojo, el error no ha dado en el `if`, sino en el `for`. Te señala lo que hay inmediatamente despues de los dos puntos del `for`, ya que considera que ahí debería haber una tabulación. No la hay, y por eso salta el error.
# ### Sintaxis
# Por tanto, toda sentencia `if`, `for`, `while`, `try`, declaración de funciones, de clases, llevan dos puntos. Y después de los dos puntos, tabulado, va todo el contenido de ese bloque. **Siempre**.
#
# Si pones los dos puntos y le das a enter, Python automáticamente te tabula todo lo que vayas a escribir a continuación.
if 1 == 1:
# Ya va tabulado
# ### ¿Tabulaciones o espacios?
# Cuidado con las tabulaciones ya que cuando pasamos de un editor de Python a otro, o cuando ejecutamos un corrector de estilo sobre nuestro código, hay veces que las tabulaciones dan problemas. Es por ello que muchos programadores en vez de usar tabulaciones, los sustituyen por 4 espacios.
#
# Este es el problema cuando no se normaliza o estandariza algo. Que cada progrmador usa lo que considera y después hay conflictos cuando pasamos de un IDE a otro. Este asunto lleva años sobre la mesa por lo que ahora la mayordía de IDEs no suelen tener problemas.
#
# Entonces, ¿qué usamos? Lo más cómodo es añadir una tabulación, pero lo más correcto son espacios. En Jupyter esto es bastante transparente para nosotros ya que cuando añadimos una tabulación, realmente Jupyter lo traduce a 4 espacios, por lo que no debería ser un tema preocupante
#
# Es tal la discusión que hasta le [han dedicado una escena en la serie Silicon Valley](https://www.youtube.com/watch?v=ussOk-ilK_8)
# ## 2. if/elif/else
# En función de lo que valgan unas condiciones booleanas, ejecutaremos unas líneas de código, u otras. La sintaxis es muy sencilla:
#
#
# > ```Python
# > if condiciones:
# > Si se cumplen las condiciones, ejecuta este código
# > else:
# > Si no, ejecutas estre otro código
# > ```
#
# Veamos un ejemplo
# +
mi_nota_examen = 7
if mi_nota_examen < 5:
print("A septiembre :(")
print("Estudia un poco")
else:
print("Aprobado!")
# -
# Únicamente se ejecuta la parte de código que consigue un `True` en la condición. `print("Aprobado!")` sólo se imprimirá por pantalla si la nota es mayor o igual a 5.
#
# Vamos a ver otro ejemplo. Ahora quiero un poco más de granularidad en la nota, con bienes, notables y tal
# +
mi_nota_de_examen = 7
if mi_nota_de_examen < 5 :
print("A septiembre :(")
elif mi_nota_de_examen < 6 :
print("Suficiente")
elif mi_nota_de_examen < 7 :
print("Bien")
elif mi_nota_de_examen < 9:
print("Notable")
else:
print("Sobresaliente")
# -
# **IMPORTANTE**. Todos los `ifs` se ejecutan secuencialmente. Por eso, en este caso no es necesario acotar tanto la nota:
#
# * Primero comprueba si es menor de 5, de ser así, suspenso.
# * Ya sabemos que es mayor o igual a 5
# * En la siguiente condición comprueba si es menor que 6, es decir, entre 5 y 6. Si es `False`, seguimos a la siguiente condición. Ahora bien, si es `True`, ejecutamos únicamente ese código y nos olvidamos de todo lo demás. Ya puede haber 150 condiciones, que si la primera es `True`, el resto es como si no existiese.
#
# Fíjate que la sintaxis es bastante intuitiva. `if` una condición, dos puntos y me ejecutas todo lo que hay aqui dentro, `elif` (acorta el `else if`), si se cumple esta otra condición, se ejecuta este otro código. Y si no se cumple ninguna de estas, ejecuta lo que haya en el `else`.
# <table align="left">
# <tr><td width="80"><img src="../../imagenes/ejercicio.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>Ejercicio if/else</h3>
#
# El ejemplo de las notas está muy bien, pero demasiado sencillo. ¿Qué pasa si la nota es mayor de 10 o menor que 0? No parece una nota correcta. En programación hay que anticiparse a los errores. Reescribe el código para tener en cuenta esos casos, cuya nota tendrás que catalogarla como "Nota errónea"
#
# </td></tr>
# </table>
# ## 3. Bucle for
# Gracias a los bucles podemos ejecutar código repetitivo, de manera bastante automática. Son muy útiles para que nuestro código no sea redundante, y también para aplicar operaciones cuando manejamos colecciones. Ya descubrirás que tienen muchísimas aplicaciones.
#
# La sintaxis de los bucles `for` es la siguiente:
#
# > ```Python
# > for var_ejecucion in limites ejecución:
# > ```
# > ```
# >
# > codigo del for...
# > ```
#
#
# * **Límites de ejecución**: La cantidad de veces que queremos que se ejecute un `for`. Por ejemplo del 1 al 10. Esto es así porque si no se ejecutarían hasta el infinito. Y además, tienen una variable de ejecución que se va actualizando. Primero valdría 1, luego 2...así hasta 10.
#
#
# * **Variable de ejecución**: dentro del for habrá una variable que se irá actualizando con cada ejecución. Si se ejecuta 10 veces, primero la variable valdrá 1, luego 2, y así hasta 10.
#
# Mejor vemos un ejemplo para entenderlo. Tienes las notas de tres alumnos en una lista, y quieres imprimir por pantalla las notas
# +
variable = 10
for variable in [1, 2, 3]:
print(variable)
print(variable)
# +
notas = [3, 4, 4]
print(notas[0])
print(notas[1])
print(notas[2])
# -
# Genial, pero qué ocurre si ahora tienes 30 notas, o simplemente quieres que tu programa no dependa de cuántas notas tienes, unas veces son 30, otras 20...
# +
notas_clase = [3, 4, 4, 6, 7, 3, 4, 5, 6, 7, 8, 9, 3]
len(notas_clase)
# -
# <table align="left">
# <tr><td width="80"><img src="../../imagenes/ejercicio.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>Ejercicio if/else</h3>
#
# Ahora imagínate que tienes todas esas notas, ¿cómo harías para imprimirlas todas? ¿Y unas que cumplan ciertas características?
#
# Recorre la lista elemento a elemento e imprime:
# - Si la nota es aprobada: la calificación y la traducción a letra (aprobado, notable..., como hemos hecho en el ejercicio del if). Ej.: "Tu calificación es Notable, has sacado un 5".
# - Si la nota es suspenso: Solo imprime que la calificación es suspenso
#
# </td></tr>
# </table>
# Pero no solo podemos iterar conjuntos de números, también podemos iterar sobre listas de strings:
# +
# Texto
dias_semana = ["L", "M", "X", "J", "V", "S", "D"]
for i in dias_semana:
print(i)
# -
# Todo objeto que sea **iterable**, lo podrás recorrer en un `for`. Los objetos iterables son aquellos que tienen un número contable de elementos, como es el caso de una lista. Veremos los iterables más en detalle en las colecciones.
# Los Strings son iterables
for letra in "Python":
print(letra)
# ### Función range
# Es muy común usar la función `range()` en las condiciones de un bucle. Esta función puede funcionar con un único argumento numérico y su output es un **iterable**, comprendido entre el 0 y el número introducido como argumento.
#
# Verás en [la documentación](https://www.w3schools.com/python/ref_func_range.asp) que `range()` tiene más posibilidades, combinando sus argumentos.
for num in range(5):
print(num)
print(range(3))
print(type(range(3)))
# En ocasiones nos interesa iterar sobre la posición que tiene cada elemento dentro de un iterable. Para ello podemos combinar `range` con `len` dentro de las condiciones del bucle
# +
colores = ["rojo", "verde", "azul"]
for i in range(len(colores)):
print(i)
colores[i] = "amarillo"
print(colores[i])
print(colores)
print("\n")
# -
colores = ["rojo", "verde", "azul"]
print(colores[0])
colores[0] = "camion"
colores
# ### Función enumerate
# ¿Y si dentro del bucle necesitamos tanto el elemento del iterable, como su índice? En [la documentación](https://www.w3schools.com/python/ref_func_enumerate.asp) verás que puedes elegir desde qué elemento de la lista quieres empezar.
# +
names = ["Pedro", "Mariano", "<NAME>"]
for cosa, cosa2 in enumerate(names):
print(f"Nombre {cosa}: {cosa2}")
# -
list(range(len(names)))
for i in range(len(names)):
try:
names.pop(i)
print(names)
except:
print("Hubo error")
# <table align="left">
# <tr><td width="80"><img src="../../imagenes/error.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>ERRORES en los rangos</h3>
#
# </td></tr>
# </table>
# Mucho cuidado al escribir las condiciones del bucle. Lo primero, porque podríamos tener condiciones infinitas de ejecución que ni nosotros, ni nuestro ordenador lo deseamos. Y lo segundo porque si intentamos acceder a un índice de nuestro iterable que no existe, saltará un error. Veamos ejemplo
# +
names = ["Pedro", "Mariano", "<NAME>"]
for i in range(4):
print(names[i])
# -
# Es por ello que se recomienda dejar el código lo más "en automático" posible. Poner en el range la longitud del iterable no es una buena práctica, ¿Y si mañana el iterable tiene menos nombres? saltará error. ¿Y si tiene más? No los tendremos en cuenta en el for. Por ello es mejor usar `len`.
# <table align="left">
# <tr><td width="80"><img src="../../imagenes/ejercicio.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>Ejercicio bucle for</h3>
#
# Recorre la siguiente lista con un for, imprime únicamente los elementos múltiplos de 3, así como el índice que tengan dentro de la lista
#
# </td></tr>
# </table>
# <table align="left">
# <tr><td width="80"><img src="./ejercicio.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>Ejercicio bucle for 2</h3>
#
# Crea una lista con los 10 primeros múltiplos de 3. Recórrela 1 a 1 e imprímelo por pantalla si es múltiplo de 2, en cuyo caso también deberás ver si es múltiplo de 5, especificando en cada caso que sea múltiplo de ese número.
#
# </td></tr>
# </table>
# <table align="left">
# <tr><td width="80"><img src="./ejercicio.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>Ejercicio bucle for 3</h3>
#
# Crea una lista con 5 colores. Añade 3 colores (de los que ya has puesto en la lista original) en las posiciones 2, 4 y 6 (posiciones de la lista final). Recorre la lista uno a uno e indica si está en los elementos anteriores a él o no.
#
# PISTA: Utiliza un bucle for dentro de otro bucle for para
#
# </td></tr>
# </table>
colores = ['rojo', 'azul', 'amarillo', 'verde']
# ## 4. Bucle while
# Se trata de otra manera de implementar un bucle en programación. Los bucles tienen que ir siempre limitados. En el caso del `for`, le poníamos un número concreto de ejecuciones, según el iterable que estuviésemos recorriendo. Para el `while` es algo diferente. Tiene una **condición de ejecución**, que mientras que se cumpla (`True`), seguirá ejecutando una y otra vez. Por otro lado, el bucle tiene una **variable de ejecucón**, al igual que en el `for`, que se irá actualizando con cada vuelta, y es esa variable la que determina cuándo acaba el bucle.
#
# Veamos un ejemplo
# +
i = 0
while(i > 0 and i < 10):
print(i)
i = i + 1
# -
# La manera más habitual de implementar estos bucles es:
# 1. Declaro la **variable de ejecución fuera del bucle**
# 2. Establezco una **condición de ejecución** para determinar cuándo queremos que se pare el bucle.
# 3. **Actualizo la variable de ejecución** en cada iteración del bucle.
#
# **Cuidado** con estos bucles ya que es muy fácil olvidarnos de actualiza la variable de ejecución, o equivocarnos en la condición de ejecución. Si esto ocurre el código se quedará corriendo hasta que detengamos el kernel (botón *interrupt the kernel*, arriba al lado del Run)
# ## 5. Break/continue
# Son dos sentencias que podemos usar dentro de los bucles para evitar ejecutar código de más.
#
# ### Break
# Se usa cuando queremos salir del bucle forzadamente. Imagina que eres una tienda y estás buscando con un for si al menos uno de los pedidos era un abrigo. Si has tenido 1000 pedidos, vas a tener que iterar sobre todos y mediante un `if`, comprobar si es un abrigo. Ahora bien, si el abrigo es el primer elemento de la lista, el `for` va a recorrer igualmente los otros 999 elementos, cuando no es necesario. Con un `break` podremos salirnos del bucle y continuar con el programa
# +
# Texto
dias_semana = ["L", "M", "X", "J", "V", "S", "D"]
for i in dias_semana:
if i == "X":
print("Miércoles encontrado")
break
print(i)
# -
for elemento in ["hola", "strings", "texto"]:
for val in elemento:
if val == "i":
break
print(val)
# ### Continue
# Esta sentencia se usa dentro de un bucle para indicarle que continue con el siguiente elemento del iterable. Al igual que con el `break`, nos sirve para evitar que se ejecute código de más. Volviendo al ejemplo anterior, si después de comprobar que tenemos un abrigo, hay 200 líneas más de código que se utiliza en otros casos, con un `continue` evitamos que se ejecute todo eso, hacemos lo que tengamos que hacer con el abrigo, y le decimos al bucle que pase al siguiente elemento, e ignore el resto del código.
for val in "stringstring":
if val == "i":
continue
print(val)
# ## 6. Try/except
# ¿Qué ocurre cuando hay un error en nuestro código? Se para toda la ejecución. Por muy buenos programadores que seamos, hay que contar siempre con que puede haber errores. Podemos llegar a controlarlos con sentencias `if/else`, por ejemplo si no sabemos muy bien los tipos de los datos, `if type(data) == float:` haces algo con floats, `else` haces otra cosa con otro tipo de datos, pero lo mejor es usar `try/except`.
#
# Ahora bien, si intuimos que el comportamiento de nuestro código puede ser algo impredecible, en programación podemos usar las sentencias `try/except` para capturar ese error, tomar decisiones, y que el código pueda continuar ejecutándose.
#
# La sintaxis es la siguiente:
#
# > ```Python
# > try:
# > Código que puede contener errores
# > except:
# > Qué hacer si nos encontramos con un error
# > ```
10/0
# +
try:
"alshhfndas".replace()
except:
print("ERROR")
print("Continua el programa")
# -
# Hay un error en el código, pero no para el programa.
#
# Podemos ser un poco más específicos con los errores, y en función del tipo de error que nos de, tomaremos diferentes caminos
# +
try:
print(variable_)
except NameError:
print("El codigo tiene errores porque la variable 'variable_' no existe")
print(NameError) # Tendremos tambien disponible el error, por si queremos manipularlo
# -
# ## 7. Resumen
# +
# If/elif/else
mi_nota_de_examen = 7
if mi_nota_de_examen < 5 :
print("A septiembre :(")
elif mi_nota_de_examen < 6 :
print("Suficiente")
elif mi_nota_de_examen < 7 :
print("Bien")
elif mi_nota_de_examen < 9 :
print("Notable")
else:
print("Sobresaliente")
# Bucle for
dias_semana = ["Lunes", "Martes", "Miércoles", "Jueves", "Viernes", "Sábado", "Domingo"]
for dia in dias_semana:
print(dia)
# Bucle while
i = 0
while(i < 5):
print(i)
i = i + 1
# Break y continue
for val in "string":
if val == "i":
break
print(val)
print("Fin")
# Try/except
try:
print(variable_)
except NameError:
print("El codigo tiene errores porque la variable 'variable_' no existe")
print("Continuo con el programa")
|
Bloque 1 - Ramp-Up/05_Python/01_Flujos de control/01_Python flujos de control.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sayakpaul/Action-Recognition-in-TensorFlow/blob/main/Data_Preparation_UCF101.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="PYgy38H_-WdA"
# ## Collect Data
# + id="oKxtdcjgjuy2"
# !wget -q --no-check-certificate https://www.crcv.ucf.edu/data/UCF101/UCF101.rar
# !wget -q --no-check-certificate https://www.crcv.ucf.edu/data/UCF101/UCF101TrainTestSplits-RecognitionTask.zip
# + id="LZGjbGEUlIwc"
# %%capture
# !unrar e UCF101.rar data/
# !unzip -qq UCF101TrainTestSplits-RecognitionTask.zip
# + [markdown] id="qtHh7Q9o-YdY"
# ## Imports
# + id="KqTOScR1i7UP"
from imutils import paths
from tqdm import tqdm
import pandas as pd
import numpy as np
import shutil
import cv2
import os
# + [markdown] id="gRlcc2Ph-arE"
# ## Metadata Loading
# + id="_rzBFXN4i7UR" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="6877e904-d3ba-45af-e782-7411e573a757"
# Open the .txt file which have names of training videos
f = open("ucfTrainTestlist/trainlist01.txt", "r")
temp = f.read()
videos = temp.split('\n')
# Create a dataframe having video names
train = pd.DataFrame()
train['video_name'] = videos
train = train[:-1]
train.head()
# + id="WLhPitHyi7US" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="8c663359-b1ac-44d7-a59c-51fd74fffb79"
# Open the .txt file which have names of test videos
with open("ucfTrainTestlist/testlist01.txt", "r") as f:
temp = f.read()
videos = temp.split("\n")
# Create a dataframe having video names
test = pd.DataFrame()
test["video_name"] = videos
test = test[:-1]
test.head()
# + [markdown] id="v7wK62Oi-lSk"
# ## Utility Functions
# + id="xCrOOcY_i7US"
def extract_tag(video_path):
return video_path.split("/")[0]
def separate_video_name(video_name):
return video_name.split("/")[1]
def rectify_video_name(video_name):
return video_name.split(" ")[0]
def move_videos(df, output_dir):
if not os.path.exists(output_dir):
os.mkdir(output_dir)
for i in tqdm(range(df.shape[0])):
videoFile = df['video_name'][i].split("/")[-1]
videoPath = os.path.join("data", videoFile)
shutil.copy2(videoPath, output_dir)
print()
print(f"Total videos: {len(os.listdir(output_dir))}")
# + [markdown] id="OtyGiF7L-nnr"
# ## DataFrame Preparation
# + id="v-lNa682i7US" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="49636796-ea5f-445c-f014-1b79c9dd534e"
train["tag"] = train["video_name"].apply(extract_tag)
train["video_name"] = train["video_name"].apply(separate_video_name)
train.head()
# + id="nVhz28EIi7UT" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="7024466d-479f-447e-d47c-d65679192e19"
train["video_name"] = train["video_name"].apply(rectify_video_name)
train.head()
# + id="5QnYrt_xi7UT" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="bd1e1784-a722-4b39-c26b-9641eb44839e"
test["tag"] = test["video_name"].apply(extract_tag)
test["video_name"] = test["video_name"].apply(separate_video_name)
test.head()
# + [markdown] id="dKo0rwC5-qa5"
# ## Filtering Top-n Actions
# + id="C0Cipx6Wi7UT" colab={"base_uri": "https://localhost:8080/"} outputId="92cbfe45-4d64-422f-d846-f88151e63225"
n = 10
topNActs = train["tag"].value_counts().nlargest(n).reset_index()["index"].tolist()
train_new = train[train["tag"].isin(topNActs)]
test_new = test[test["tag"].isin(topNActs)]
train_new.shape, test_new.shape
# + id="0HeihX0_i7UU"
train_new = train_new.reset_index(drop=True)
test_new = test_new.reset_index(drop=True)
# + [markdown] id="Q0BtXtt8-vMU"
# ## Move Top-n Action Videos
# + id="5vjduxM_xEoB" colab={"base_uri": "https://localhost:8080/"} outputId="cea7dba5-164e-4d7f-e743-70a33a4db5dc"
move_videos(train_new, "train")
move_videos(test_new, "test")
# + id="7_9w-5ArxWuc"
train_new.to_csv("train.csv", index=False)
test_new.to_csv("test.csv", index=False)
# + [markdown] id="_qbrwlIX-xqJ"
# ## Serialization
# + id="7GUzA8d1xxJR"
# !tar cf ucf101_top10.tar.gz train test train.csv test.csv
# + id="aagZH1iLx99b" colab={"base_uri": "https://localhost:8080/"} outputId="5429c06f-e970-4b4e-a2ad-be030ea8a2d3"
from google.colab import drive
drive.mount('/content/drive')
# + id="vjFRFB2Dx_aO"
# !cp ucf101_top10.tar.gz /content/drive/MyDrive
|
Data_Preparation_UCF101.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: AutoEq (Python 3.7)
# language: python
# name: autoeq
# ---
import os
import sys
from glob import glob
import numpy as np
import pandas as pd
import scipy
import matplotlib.pyplot as plt
from tabulate import tabulate
sys.path.insert(1, os.path.realpath(os.path.join(sys.path[0], os.pardir, os.pardir)))
from frequency_response import FrequencyResponse
from constants import ROOT_DIR
with open('frequencies.csv', 'r', encoding='utf-8') as fh:
onear_frequencies = [float(x) for x in fh.read().strip().split('\n')[::-1]]
print(onear_frequencies)
with open('inear_frequencies.csv', 'r', encoding='utf-8') as fh:
inear_frequencies = [float(x) for x in fh.read().strip().split('\n')[::-1]]
print(inear_frequencies)
# +
inear_frequencies = [20.0, 21.2, 22.4, 23.6, 25.0, 26.5, 28.0, 30.0, 31.5, 33.5, 35.5, 37.5, 40.0, 42.5, 45.0, 47.5, 50.0, 53.0, 56.0, 60.0, 63.0, 67.0, 71.0, 75.0, 80.0, 85.0, 90.0, 95.0, 100.0, 106.0, 112.0, 118.0, 125.0, 132.0, 140.0, 150.0, 160.0, 170.0, 180.0, 190.0, 200.0, 212.0, 224.0, 236.0, 250.0, 265.0, 280.0, 300.0, 315.0, 335.0, 355.0, 375.0, 400.0, 425.0, 450.0, 475.0, 500.0, 530.0, 560.0, 600.0, 630.0, 670.0, 710.0, 750.0, 800.0, 850.0, 900.0, 950.0, 1000.0, 1060.0, 1120.0, 1180.0, 1250.0, 1320.0, 1400.0, 1500.0, 1600.0, 1700.0, 1800.0, 1900.0, 2000.0, 2120.0, 2240.0, 2360.0, 2500.0, 2650.0, 2800.0, 3000.0, 3150.0, 3350.0, 3550.0, 3750.0, 4000.0, 4250.0, 4500.0, 4750.0, 5000.0, 5300.0, 5600.0, 6000.0, 6300.0, 6700.0, 7100.0, 7500.0, 8000.0, 8500.0, 9000.0, 9500.0, 10000.0, 10600.0, 11200.0, 11800.0, 12500.0, 13200.0, 14000.0, 15000.0, 16000.0, 17000.0, 18000.0, 19000.0, 20000.0]
inear_frequencies = np.array(inear_frequencies)
def inear_score(fr):
fr = fr.copy()
fr.interpolate(inear_frequencies)
sl = np.logical_and(fr.frequency >= 20, fr.frequency <= 10000)
x = fr.frequency[sl]
xm = np.mean(x)
y = fr.error[sl]
ym = np.mean(y)
slope, _, _, _, _ = scipy.stats.linregress(np.log(x), y)
mean = np.mean(np.abs(fr.error[np.logical_and(fr.frequency >= 40, fr.frequency <= 10000)]))
std = np.std(y)
score = 100.0795 - 8.5 * std - 6.796 * np.abs(slope) - 3.475 * mean
# TODO: score and std differs from oratory1990 PDFs, could be Harman in-ear 2017-1 target
return score, slope, mean, std, fr.error
onear_frequencies = [20.0, 21.0, 22.0, 24.0, 25.0, 27.0, 28.0, 30.0, 32.0, 34.0, 36.0, 38.0, 40.0, 43.0, 45.0, 48.0, 50.0, 53.0, 56.0, 60.0, 63.0, 67.0, 71.0, 75.0, 80.0, 85.0, 90.0, 95.0, 100.0, 106.0, 112.0, 118.0, 125.0, 132.0, 140.0, 150.0, 160.0, 170.0, 180.0, 190.0, 200.0, 212.0, 224.0, 236.0, 250.0, 265.0, 280.0, 300.0, 315.0, 335.0, 355.0, 375.0, 400.0, 425.0, 450.0, 475.0, 500.0, 530.0, 560.0, 600.0, 630.0, 670.0, 710.0, 750.0, 800.0, 850.0, 900.0, 950.0, 1000.0, 1060.0, 1120.0, 1180.0, 1250.0, 1320.0, 1400.0, 1500.0, 1600.0, 1700.0, 1800.0, 1900.0, 2000.0, 2120.0, 2240.0, 2360.0, 2500.0, 2650.0, 2800.0, 3000.0, 3150.0, 3350.0, 3550.0, 3750.0, 4000.0, 4250.0, 4500.0, 4750.0, 5000.0, 5300.0, 5600.0, 6000.0, 6300.0, 6700.0, 7100.0, 7500.0, 8000.0, 8500.0, 9000.0, 9500.0, 10000.0, 10600.0, 11200.0, 11800.0, 12500.0, 13200.0, 14000.0, 15000.0, 16000.0, 17000.0, 18000.0, 19000.0, 20000.0]
onear_frequencies = np.array(onear_frequencies)
def onear_score(fr):
fr = fr.copy()
fr.interpolate(onear_frequencies)
sl = np.logical_and(fr.frequency >= 50, fr.frequency <= 10000)
x = fr.frequency[sl]
xm = np.mean(x)
y = fr.error[sl]
ym = np.mean(y)
slope, _, _, _, _ = scipy.stats.linregress(np.log(x), y)
std = np.std(y)
mean = np.mean(np.abs(y))
score = 114.490443008238 - 12.62 * std - 15.5163857197367 * np.abs(slope)
return score, slope, mean, std, fr.error
# -
harman_oe = FrequencyResponse.read_from_csv(os.path.join(ROOT_DIR, 'compensation', 'harman_over-ear_2018.csv'))
onear = []
errs = []
names = []
for fp in glob(os.path.join(ROOT_DIR, 'measurements', 'oratory1990', 'data', 'onear', '*', '*.csv')):
fr = FrequencyResponse.read_from_csv(fp)
fr.compensate(harman_oe, bass_boost_gain=0.0)
score, slope, mean, std, err = onear_score(fr)
onear.append([fr.name, f'{score:.0f}', f'{slope:.2f}', f'{mean:.2f}', f'{std:.2f}'])
errs.append(np.concatenate([[std, slope, score], err[::-1]]))
names.append(fr.name)
# TODO: ignore samples
errs = np.vstack(errs)
pd.DataFrame(errs.transpose(), columns=names).to_csv('onear_errs.csv', header=True, index=False)
onear = sorted(onear, key=lambda x: float(x[1]), reverse=True)
onear_table = tabulate(
onear, headers=['Model', 'Score', 'Slope', 'Mean', 'STD'], tablefmt='orgtbl'
).replace('+', '|').replace('|-', '|:')
print(onear_table)
harman_ie = FrequencyResponse.read_from_csv(os.path.join(ROOT_DIR, 'compensation', 'harman_in-ear_2019v2.csv'))
inear = []
errs = []
names = []
for fp in glob(os.path.join(ROOT_DIR, 'measurements', 'oratory1990', 'data', 'inear', '*', '*.csv')):
fr = FrequencyResponse.read_from_csv(fp)
fr.compensate(harman_ie, bass_boost_gain=0.0)
score, slope, mean, std, err = inear_score(fr)
inear.append([fr.name, f'{score:.0f}', f'{slope:.2f}', f'{mean:.2f}', f'{std:.2f}'])
errs.append(np.concatenate([[std, slope, mean, score], err[::-1]]))
names.append(fr.name)
# TODO: ignore samples
errs = np.vstack(errs)
pd.DataFrame(errs.transpose(), columns=names).to_csv('inear_errs.csv', header=True, index=False)
|
research/preference_score/preference_score.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DETECTING RBG PART IN AN IMAGE
# importing package/libraries
import matplotlib.pyplot as plt
import numpy as np
from PIL import Image
pil = Image.open(r".\images\opencv.jpg")
pil
# Converting image to numpy array
arr = np.array(pil)
arr
#to get demension of matrix(M X N) and 3 is the thickness or layers.
arr.shape
#display image
plt.imshow(arr)
# ### Slicing and detecting red,green and blue part in image.
# #### 0 is used for RED
# #### 1 is used for BLUE
# #### 2 is used for GREEN.
red = arr[:,:,0]
plt.imshow(red)
green = arr[:,:,1]
plt.imshow(green)
blue = arr[:,:,2]
plt.imshow(blue)
|
Image filters/detect_rgb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Importing
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import requests
from bs4 import BeautifulSoup
import re
import pandas as pd
import os
import time
import datetime
import csv
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from __future__ import division
# # Reading the 3 workouts dfs to clean
apr14_workouts = pd.read_csv('workouts_csv')
apr7_workouts = pd.read_csv('workouts_csv_2')
apr6_workouts = pd.read_csv('workouts_csv_3')
allworkouts_uncleaned = pd.concat([apr14_workouts, apr7_workouts, apr6_workouts], axis=0)
allworkouts_uncleaned.to_csv('allworkouts_uncleaned')
# + active=""
# allworkouts_uncleaned = pd.read_csv('allworkouts_uncleaned')
# -
len(allworkouts_uncleaned)
allworkouts_uncleaned.sample(10)
# +
allworkouts_clean = allworkouts_uncleaned.drop('Unnamed: 0', axis=1).drop('Course', axis=1).drop('Note', axis=1 ).drop('Rank', axis=1).drop('Track',axis=1)
# -
allworkouts_clean
# +
Practice_days = allworkouts_clean.groupby(['Horse Name']).Date.count().reset_index()
#Practice_days = allworkouts_clean.groupby((['Horse Name']).Date isin '\17')
#Practice_counts = allworkouts_clean.groupby(['Horse Name']).Distance.value_counts()
#allworkouts_clean3 = allworkouts_clean2[(allworkouts_clean2.Distance == "4F") | (allworkouts_clean2.Distance == "5F")
# -
Practice_days.head(10) ###Combine into final data frame.
# +
#allworkouts_clean = allworkouts_clean.filter['5F','4F']
#allworkouts_clean = (allworkouts_clean['Distance'] == '5F') or (allworkouts_clean['Distance'] == '4F')
#FiveF = allworkouts_clean['Distance'] != "2F","3F","6F"
#FourF = allworkouts_clean['Distance'] == "4F"
#allworkouts_clean = allworkouts_clean[FiveF]
# -
# ### Creating df from workouts df of number of practice days
allworkouts_clean2 = allworkouts_clean.groupby(['Horse Name', "Distance"]).Time_tenths_second.agg([np.mean]).reset_index()
allworkouts_clean3 = allworkouts_clean2[(allworkouts_clean2.Distance == "4F") | (allworkouts_clean2.Distance == "5F") | (allworkouts_clean2.Distance == "3F")]
allworkouts_clean3.head(10)
# +
allworkouts_clean4 = allworkouts_clean3.pivot(index='Horse Name', columns='Distance', values='mean').reset_index()
#allworkouts_clean4 = allworkouts_clean3.unstack(level=-1)
# -
allworkouts_clean4.head(10)
allworkouts_clean4.to_csv('workout_means')
# # Combining final DF
#Merging all the workout dataframes
apr14_final = pd.read_csv('final_merged')
apr7_final = pd.read_csv('final_merged2')
apr6_final = pd.read_csv('final_merged3')
finalmerged_uncleaned = pd.concat([apr14_final, apr7_final, apr6_final], axis=0)
finalmerged_uncleaned.to_csv('final_Times.csv')
finalmerged_uncleaned = pd.read_csv('final_Times.csv').drop('Unnamed: 0', axis=1)
#Adding the percentage of the time the horse places in a race.
finalmerged_uncleaned["PercentWin"] = (finalmerged_uncleaned["Firsts"] + finalmerged_uncleaned["Seconds"]+ finalmerged_uncleaned["Thirds"])/ (finalmerged_uncleaned["Starts"])
finalmerged_uncleaned.head(2)
#Merging the final times df with the #of practice days df
finalmerged_clean = pd.merge(finalmerged_uncleaned,Practice_days,on='Horse Name', how = 'left')
finalmerged_clean.head(2)
#Merging the merged data frame, with the workouts df
finalmerged_clean = pd.merge(finalmerged_clean,allworkouts_clean4,on='Horse Name', how = 'left')
finalmerged_clean.head(2)
#Renaming the columns so they don't confuse the model later.
finalmerged_clean = finalmerged_clean.rename(columns={'3F':'ThreeF'}).rename(columns={'4F':'FourF'}).rename(columns={'5F':'FiveF'}).fillna(0)
finalmerged_clean
finalmerged_clean.shape
finalmerged_clean.to_csv('finalmerged_clean')
|
Horse Race Data - Race time prediction/Step2_DataCleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
import warnings
warnings.filterwarnings("ignore")
tf.set_random_seed(1)
np.random.seed(1)
LR = 0.01
BATCH_SIZE = 32
# fake data
x = np.linspace(-1, 1, 100)[:, np.newaxis] # shape (100, 1)
noise = np.random.normal(0, 0.1, size=x.shape)
y = np.power(x, 2) + noise # shape (100, 1) + some noise
# plot dataset
plt.scatter(x, y)
plt.show()
# default network
class Net:
def __init__(self, opt, **kwargs):
self.x = tf.placeholder(tf.float32, [None, 1])
self.y = tf.placeholder(tf.float32, [None, 1])
l = tf.layers.dense(self.x, 20, tf.nn.relu)
out = tf.layers.dense(l, 1)
self.loss = tf.losses.mean_squared_error(self.y, out)
self.train = opt(LR, **kwargs).minimize(self.loss)
# different nets
net_SGD = Net(tf.train.GradientDescentOptimizer)
net_Momentum = Net(tf.train.MomentumOptimizer, momentum=0.9)
net_RMSprop = Net(tf.train.RMSPropOptimizer)
net_Adam = Net(tf.train.AdamOptimizer)
nets = [net_SGD, net_Momentum, net_RMSprop, net_Adam]
sess = tf.Session()
sess.run(tf.global_variables_initializer())
losses_his = [[], [], [], []] # record loss
# training
for step in range(300): # for each training step
index = np.random.randint(0, x.shape[0], BATCH_SIZE)
b_x = x[index]
b_y = y[index]
for net, l_his in zip(nets, losses_his):
_, l = sess.run([net.train, net.loss], {net.x: b_x, net.y: b_y})
l_his.append(l) # loss recoder
# plot loss history
labels = ['SGD', 'Momentum', 'RMSprop', 'Adam']
for i, l_his in enumerate(losses_his):
plt.plot(l_his, label=labels[i])
plt.legend(loc='best')
plt.xlabel('Steps')
plt.ylabel('Loss')
plt.ylim((0, 0.2))
plt.show()
|
TensorflowTUT2/304_optimizer.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # Inference with Tensorflow Lite
//
// In this tutorial, you learn how to load an existing TensorFlow Lite model and use it to run a prediction task.
//
//
// ## Preparation
//
// This tutorial requires the installation of Java Kernel. For more information on installing the Java Kernel, see the [README](https://github.com/deepjavalibrary/djl/blob/master/jupyter/README.md).
// +
// // %mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
// %maven ai.djl:api:0.16.0
// %maven ai.djl:model-zoo:0.16.0
// %maven ai.djl.tflite:tflite-engine:0.16.0
// %maven org.slf4j:slf4j-simple:1.7.32
// Use secondary engine to help pre-processing and post-processing
// %maven ai.djl.pytorch:pytorch-engine:0.16.0
// -
import java.awt.image.*;
import java.nio.file.*;
import ai.djl.*;
import ai.djl.inference.*;
import ai.djl.ndarray.*;
import ai.djl.modality.*;
import ai.djl.modality.cv.*;
import ai.djl.modality.cv.util.*;
import ai.djl.modality.cv.transform.*;
import ai.djl.modality.cv.translator.*;
import ai.djl.repository.zoo.*;
import ai.djl.translate.*;
import ai.djl.training.util.*;
import ai.djl.util.*;
// ## Step 1: Load your Tensorflow Lite mode from DJL model zoo
Criteria<Image, Classifications> criteria = Criteria.builder()
.setTypes(Image.class, Classifications.class)
.optEngine("TFLite")
.optFilter("dataset", "aiyDish")
.build();
ZooModel<Image, Classifications> model = criteria.loadModel();
// ## Step 2: Create a Predictor
Predictor<Image, Classifications> predictor = model.newPredictor();
// ## Step 3: Load image for classification
// +
var img = ImageFactory.getInstance().fromUrl("https://resources.djl.ai/images/sachertorte.jpg");
img.getWrappedImage()
// -
// ## Step 4: Run inference
// +
Classifications classifications = predictor.predict(img);
classifications
// -
// ## Summary
//
// Now, you can load Tensorflow Lite model and run inference.
//
|
jupyter/tensorflow_lite/inference_with_tensorflow_lite.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment2 Day6
import math
class cone():
def __init__(self,radius,height):
self.radius = radius
self.height = height
def Volume(self):
return math.pi*(self.radius**2)*(self.height/3)
def SurfaceArea(self,a):
if a == 'base':
return math.pi *(self.radius**2)
elif a == 'side':
return math.pi * self.radius * (math.sqrt((self.radius**2)+(self.height**2)))
c = cone(8,7)
c.Volume()
c.SurfaceArea('base')
c.SurfaceArea('side')
|
Assignment2D6.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Constrained clustering with noisy constraints
# In this example, we add some constraint noise to the problem, which simulates an imperfect oracle. Any constrained clustering method must account for errors in the constraint set, especially with the prominence of crowdsourced solutions for annotation.
#
# There has been almost no coverage of dealing with imperfect oracles in the literature. Aside from this work, only "Constrained clustering with imperfect oracles" [Zhu 2015] exists. As opposed with the method of Zhu, our ImperfectOracles tool very explicitly detects constraints which interfere with fellow constraints, and automatically determines the number of constraints that should be removed. Thus, it is appropriate to apply even to perfect constraint sets.
#
# The ImperfectOracles tool functions by creating an ensemble of reasonable clusterings of the data. Constraints which are mutually satisfied are said to vote for one another. Constraints present in the same clusters are said to be relevant. A measure called "trust" is the fraction of the relevant, trusted vote that a constraint gets from its peers. The trust is calculated with a fixed point update based on this recursive definition.
# +
import numpy as np
import matplotlib.pyplot as plt
import sklearn.datasets as ds
from robustclust import get_constraints, \
plot_constraints, \
remove_constraints
# -
# First, make some data and make some constraints. We can add random constraint errors by flipping some constraint values (ML to CL, CL to ML).
# +
N, n_clusters, n_constraints, err_rate = (300, 4, 200, 0.0)
data, labels = ds.make_blobs(n_samples=N, n_features=2, centers=n_clusters)
constraint_mat, _ = get_constraints(
data,
labels,
method='mmffqs',
n_constraints=n_constraints,
err_rate=err_rate)
plot_constraints(data, constraint_mat=constraint_mat)
# -
# Remove errant constraints
keep_idx = remove_constraints(data, constraint_mat, n_clusters=n_clusters, threshold=0.999)
print(constraint_mat.shape)
print(np.mean(keep_idx))
# The results are not very good, we need to clean the constraint set. Use the ImperfectOracle class.
# +
io = IO(data=data,
constraintMat=constraintMat,
n_clusters=Nclusters)
keepInd = io.remove_constraints()
plt.figure()
io.plot_removal(labels, keepInd)
plt.tight_layout()
plt.show()
# -
# After cleaning the constraint set, let's try turning constraints to labels again.
# +
newConstraintMat = constraintMat[keepInd,:]
ctl2 = CTL(data=data,
constraintMat=newConstraintMat,
n_clusters=Nclusters)
ctl2.fit_constrained()
plt.figure()
cc.plot_labels(data)
cc.plot_labels(data[ctl2.constrainedSamps], ctl2.labelSet)
plt.show()
# -
# We now have turned the noisy constraint set into a very useful set of labels, with no additional information.
|
examples/constrained_clustering_with_noise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# ## Activation function
# +
def sigmoid(z):
return 1/(1 + np.exp(-z))
def sigmoid_d(z):
return sigmoid(z) * (1 - sigmoid(z))
# -
# ### Cost funtion
# +
def cost(activation, target):
return 0.5 * (target - activation) ** 2
def cost_d(activation, target):
return (target - activation)
# -
# ### Simple ANN with 1 hidden layer
#
# Here we manually do backpropagation to learn weights for a simple nn with 30 hidden weights that will attempt to classify between 0 and 1 in the mnist dataset
# +
input_dim = 28 * 28
hidden_dim = 30
out_dim = 10
hidden_weights = np.random.normal(0, 1, (input_dim, hidden_dim))
out_weights = np.random.normal(0, 1, (hidden_dim, out_dim))
hidden_biases = np.zeros((1, hidden_dim))
out_biases = np.zeros((1, out_dim))
# -
def forward(x):
"""Compute a full forward pass of this small network"""
hidden_activations = sigmoid(np.dot(x, hidden_weights) + hidden_biases)
print('hidden a', hidden_activations.shape)
out_activations = sigmoid(np.dot(hidden_activations, out_weights) + out_biases)
print('out_a', out_activations.shape)
return out_activations
def backpropagate(x, y):
# Do a forward pass but cache the activations and z's
hidden_z = np.dot(x, hidden_weights) + hidden_biases
hidden_a = sigmoid(hidden_z)
out_z = np.dot(hidden_a, out_weights) + out_biases
out_a = sigmoid(out_z)
correct = np.count_nonzero(np.argmax(out_a, 1) == np.argmax(y, 1))
# Now do the backwards pass
# Output layer
out_error = cost_d(out_a, y) * sigmoid_d(out_z)
# Partial derivatives
change_out_biases = np.sum(out_error, 0)
change_out_weights = np.dot(hidden_a.T, out_error)
# Hidden layer
# Backpropagate the error
hidden_error = np.dot(out_weights, out_error.T).T * sigmoid_d(hidden_z)
# Get the partial derivatives
change_hidden_biases = np.sum(hidden_error, 0)
change_hidden_weights = np.dot(x.T, hidden_error)
return correct, change_out_weights, change_out_biases, change_hidden_weights, change_hidden_biases
# ### Get the MNIST dataset
# And just grab the 0 and 1 images
from data_loaders import MNISTLoader
mloader = MNISTLoader()
train_data, train_labels = mloader.get_training_set()
# +
num_samples = train_labels.shape[0]
# get images as vectors
x = train_data.reshape(num_samples, -1)/255
print(x[0].max())
# one-hot encode labels
y = np.zeros((num_samples, 10))
y[np.arange(num_samples),train_labels] = 1
# -
# ### Stochastic gradient descent
#
# Use SGD to train this small network for a few epochs and watch our training accuracy
learning_rate = 1e-2
batch_size = 5
batch_starts = np.arange(batch_size, num_samples, batch_size)
idxes = np.arange(0, num_samples, 1)
num_epochs = 1
for epoch in range(num_epochs):
np.random.shuffle(idxes)
batches = np.split(idxes, batch_starts)
total_correct = 0
for batch in batches:
correct, d_ow, d_ob, d_hw, d_hb = backpropagate(x[batch], y[batch])
total_correct += correct
out_weights += learning_rate * (1/batch_size) * d_ow
out_biases += learning_rate * (1/batch_size) * d_ob
hidden_weights += learning_rate * (1/batch_size) * d_hw
hidden_biases += learning_rate * (1/batch_size) * d_hb
print(f'{total_correct} / {num_samples}')
plt.imshow(x[50280].reshape(28, 28))
50000/60000
|
nn/dev_notebooks/dev_simple_mnist_network_vectorised_batches.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.0
# language: julia
# name: julia-0.6
# ---
# # Parameter estimation of Lotka Volterra model using optimisation methods
using ParameterizedFunctions, OrdinaryDiffEq, DiffEqParamEstim
using BlackBoxOptim, NLopt, Plots, RecursiveArrayTools, QuadDIRECT
gr(fmt=:png)
loc_bounds = Tuple{Float64, Float64}[(0, 5), (0, 5), (0, 5), (0, 5)]
glo_bounds = Tuple{Float64, Float64}[(0, 10), (0, 10), (0, 10), (0, 10)]
loc_init = [1,0.5,3.5,1.5]
glo_init = [5,5,5,5]
f = @ode_def_nohes LotkaVolterraTest begin
dx = a*x - b*x*y
dy = -c*y + d*x*y
end a b c d
u0 = [1.0,1.0] #initial values
tspan = (0.0,10.0)
p = [1.5,1.0,3.0,1,0] #parameters used, these need to be estimated from the data
tspan = (0.0, 30.0) # sample of 3000 observations over the (0,30) timespan
prob = ODEProblem(f, u0, tspan,p)
tspan2 = (0.0, 3.0) # sample of 3000 observations over the (0,30) timespan
prob_short = ODEProblem(f, u0, tspan2,p)
dt = 30.0/3000
tf = 30.0
tinterval = 0:dt:tf
t = collect(tinterval)
h = 0.01
M = 300
tstart = 0.0
tstop = tstart + M * h
tinterval_short = 0:h:tstop
t_short = collect(tinterval_short)
#Generate Data
data_sol_short = solve(prob_short,Tsit5(),saveat=t_short,reltol=1e-9,abstol=1e-9)
data_short = convert(Array, data_sol_short)
data_sol = solve(prob,Tsit5(),saveat=t,reltol=1e-9,abstol=1e-9)
data = convert(Array, data_sol)
# #### Plot of the solution
# ##### Short Solution
p1 = plot(data_sol_short)
# ##### Longer Solution
p2 = plot(data_sol)
# ### Local Solution from the short data set
obj_short = build_loss_objective(prob_short,Tsit5(),L2Loss(t_short,data_short),tstops=t_short)
res1 = bboptimize(obj_short;SearchRange = glo_bounds, MaxSteps = 7e3)
# Lower tolerance could lead to smaller fitness (more accuracy)
obj_short = build_loss_objective(prob_short,Tsit5(),L2Loss(t_short,data_short),tstops=t_short,reltol=1e-9)
res1 = bboptimize(obj_short;SearchRange = glo_bounds, MaxSteps = 7e3)
# Change in tolerance makes it worse
obj_short = build_loss_objective(prob_short,Vern9(),L2Loss(t_short,data_short),tstops=t_short,reltol=1e-9,abstol=1e-9)
res1 = bboptimize(obj_short;SearchRange = glo_bounds, MaxSteps = 7e3)
# using the moe accurate Vern9() reduces the fitness marginally and leads to some increase in time taken
# # Using NLopt
# #### Global Optimisation first
obj_short = build_loss_objective(prob_short,Vern9(),L2Loss(t_short,data_short),tstops=t_short,reltol=1e-9,abstol=1e-9)
opt = Opt(:GN_ORIG_DIRECT_L, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[10.0,10.0,10.0,10.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,glo_init)
opt = Opt(:GN_CRS2_LM, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[10.0,10.0,10.0,10.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,glo_init)
opt = Opt(:GN_ISRES, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[10.0,10.0,10.0,10.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,glo_init)
opt = Opt(:GN_ESCH, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[10.0,10.0,10.0,10.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,glo_init)
# Now local optimization algorithms are used to check the global ones, these use the local constraints, different intial values and time step
opt = Opt(:LN_BOBYQA, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
opt = Opt(:LN_NELDERMEAD, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
opt = Opt(:LD_SLSQP, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
opt = Opt(:LN_COBYLA, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
opt = Opt(:LN_NEWUOA_BOUND, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
opt = Opt(:LN_PRAXIS, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
opt = Opt(:LN_SBPLX, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
opt = Opt(:LD_MMA, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
opt = Opt(:LD_TNEWTON_PRECOND_RESTART, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj_short.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
# ## Now the longer problem is solved for a global solution
#
# Vern9 solver with reltol=1e-9 and abstol=1e-9 is used and the dataset is increased to 3000 observations per variable with the same integration time step of 0.01.
obj = build_loss_objective(prob,Vern9(),L2Loss(t,data),tstops=t,reltol=1e-9,abstol=1e-9)
res1 = bboptimize(obj;SearchRange = glo_bounds, MaxSteps = 4e3)
opt = Opt(:GN_ORIG_DIRECT_L, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[10.0,10.0,10.0,10.0])
min_objective!(opt, obj.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,glo_init)
opt = Opt(:GN_CRS2_LM, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[10.0,10.0,10.0,10.0])
min_objective!(opt, obj.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 20000)
@time (minf,minx,ret) = NLopt.optimize(opt,glo_init)
opt = Opt(:GN_ISRES, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[10.0,10.0,10.0,10.0])
min_objective!(opt, obj.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 50000)
@time (minf,minx,ret) = NLopt.optimize(opt,glo_init)
opt = Opt(:GN_ESCH, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[10.0,10.0,10.0,10.0])
min_objective!(opt, obj.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 20000)
@time (minf,minx,ret) = NLopt.optimize(opt,glo_init)
opt = Opt(:LN_BOBYQA, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
opt = Opt(:LN_NELDERMEAD, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj.cost_function2)
xtol_rel!(opt,1e-9)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
opt = Opt(:LD_SLSQP, 4)
lower_bounds!(opt,[0.0,0.0,0.0,0.0])
upper_bounds!(opt,[5.0,5.0,5.0,5.0])
min_objective!(opt, obj.cost_function2)
xtol_rel!(opt,1e-12)
maxeval!(opt, 10000)
@time (minf,minx,ret) = NLopt.optimize(opt,loc_init)
# #### Using QuadDIRECT
obj_short = build_loss_objective(prob_short,Tsit5(),L2Loss(t_short,data_short),tstops=t_short)
lower = [0.0,0.0,0.0,0.0]
upper = [5.0,5.0,5.0,5.0]
splits = ([0.0,1.0,3.0],[0.0,1.0,3.0],[0.0,1.0,3.0],[0.0,1.0,3.0])
root, x0 = analyze(obj_short,splits,lower,upper)
minimum(root)
obj = build_loss_objective(prob,Vern9(),L2Loss(t,data),tstops=t,reltol=1e-9,abstol=1e-9)
lower = [0.0,0.0,0.0,0.0]
upper = [10.0,10.0,10.0,10.0]
splits = ([0.0,3.0,6.0],[0.0,3.0,6.0],[0.0,3.0,6.0],[0.0,3.0,6.0])
root, x0 = analyze(obj,splits,lower,upper)
minimum(root)
# #### Parameter estimation on the longer sample proves to be extremely challenging for some of the global optimizers. A few give the accurate values, BlacBoxOptim also performs quite well while others seem to struggle with accuracy a lot.
# # Conclusion
# In general we observe that lower tolerance lead to higher accuracy but too low tolerance could affect the convergance time drastically. Also fitting a shorter timespan seems to be easier in comparision (quite intutively). NLOpt methods seem to give great accuracy in the shorter problem with a lot of the algorithms giving 0 fitness, BBO performs very well on it with marginal change with `tol` values. In case of global optimization of the longer problem there is some difference in the perfomance amongst the algorithms with `LD_SLSQP` `GN_ESCH` `GN_ISRES` `GN_ORIG_DIRECT_L` performing among the worse, BBO also gives a bit high fitness in comparison. QuadDIRECT gives accurate results in the case of the shorter problem but doesn't perform very well in the longer problem case.
|
ParameterEstimation/LotkaVolterraParameterEstimation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Writing the Client Node
import rospy
from rospy_tutorials.srv import *
# The client code for calling services is also simple. For clients you don't have to call `init_node()`. We first call:
rospy.wait_for_service('add_two_ints')
# This is a convenience method that blocks until the service named `add_two_ints` is available. Next we create a handle for calling the service:
add_two_ints = rospy.ServiceProxy('add_two_ints', AddTwoInts)
# We can use this handle just like a normal function and call it:
try:
print( add_two_ints(5, 3) )
except rospy.ServiceException, e:
print "Service call failed: %s"%e
# Because we've declared the type of the service to be `AddTwoInts`, it does the work of generating the `AddTwoIntsRequest` object for you (you're free to pass in your own instead). The return value is an `AddTwoIntsResponse` object. If the call fails, a `rospy.ServiceException` may be thrown, so you should setup the appropriate `try/except` block.
|
Notebooks/core_tutorials/Writing the Client Node in Python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Voronoi Graphs
# + inputHidden=false outputHidden=false
# Make the relevant imports including Voronoi methods
import numpy as np
from scipy.spatial import Voronoi, voronoi_plot_2d
import matplotlib.pyplot as plt
# %matplotlib inline
# -
plt.rcParams["figure.figsize"] = [12, 12]
# +
# Recreate the figure above for a new set of random points
points = np.random.randint(50, size=(50, 2))
graph = Voronoi(points)
voronoi_plot_2d(graph)
plt.show()
# + inputHidden=false outputHidden=false
# Read in the obstacle data
filename = 'colliders.csv'
data = np.loadtxt(filename, delimiter=',', dtype='Float64', skiprows=2)
# -
# If you want to use the prebuilt bresenham method
# Import the Bresenham package
from bresenham import bresenham
# Here you'll modify the `create_grid()` method from a previous exercise
# In this new function you'll record obstacle centres and
# create a Voronoi graph around those points
def create_grid_and_edges(data, drone_altitude, safety_distance):
"""
Returns a grid representation of a 2D configuration space
along with Voronoi graph edges given obstacle data and the
drone's altitude.
"""
# minimum and maximum north coordinates
north_min = np.floor(np.min(data[:, 0] - data[:, 3]))
north_max = np.ceil(np.max(data[:, 0] + data[:, 3]))
# minimum and maximum east coordinates
east_min = np.floor(np.min(data[:, 1] - data[:, 4]))
east_max = np.ceil(np.max(data[:, 1] + data[:, 4]))
# given the minimum and maximum coordinates we can
# calculate the size of the grid.
north_size = int(np.ceil((north_max - north_min)))
east_size = int(np.ceil((east_max - east_min)))
# Initialize an empty grid
grid = np.zeros((north_size, east_size))
# Center offset for grid
north_min_center = np.min(data[:, 0])
east_min_center = np.min(data[:, 1])
# Define a list to hold Voronoi points
points = []
# Populate the grid with obstacles
for i in range(data.shape[0]):
north, east, alt, d_north, d_east, d_alt = data[i, :]
if alt + d_alt + safety_distance > drone_altitude:
obstacle = [
int(north - d_north - safety_distance - north_min_center), # min north
int(north + d_north + safety_distance - north_min_center), # max north
int(east - d_east - safety_distance - east_min_center), # min east
int(east + d_east + safety_distance - east_min_center), # max east
]
grid[obstacle[0]:obstacle[1], obstacle[2]:obstacle[3]] = 1
# add center of obstacles to points list
points.append([north - north_min, east - east_min])
# create a voronoi graph based on location of obstacle centres
graph = Voronoi(points)
# TODO: check each edge from graph.ridge_vertices for collision
edges = []
for v in graph.ridge_vertices:
v1 = graph.vertices[v[0]].astype(np.int)
v2 = graph.vertices[v[1]].astype(np.int)
valid = True
cells = bresenham(v1[0], v1[1], v2[0], v2[1])
for c in cells:
if c[0] < 0 or c[1] < 0 or c[0] >= grid.shape[0] or c[1] >= grid.shape[1]:
valid = False
break
if grid[c[0], c[1]] == 1:
valid = False
break
if valid:
edges.append((v1, v2))
return grid, edges
# Define a flying altitude (feel free to change this)
drone_altitude = 5
safety_distance = 3
grid, edges = create_grid_and_edges(data, drone_altitude, safety_distance)
print('Found %5d edges' % len(edges))
# +
# equivalent to
# plt.imshow(np.flip(grid, 0))
# Plot it up!
plt.imshow(grid, origin='lower', cmap='Greys')
# Stepping through each edge
for e in edges:
p1 = e[0]
p2 = e[1]
plt.plot([p1[1], p2[1]], [p1[0], p2[0]], 'b-')
plt.xlabel('EAST')
plt.ylabel('NORTH')
plt.show()
# -
|
Course02/Voronoi.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Accessing MODIS snow data with the Planetary Computer STAC API
#
# The planetary computer hosts two snow-related MODIS 6.1 products:
#
# - Snow cover daily (10A1)
# - Snow cover 8-day (10A2)
#
# For more information about the products themselves, check out the Data Pages at the [bottom of this document](#data-pages).
# ### Environment setup
#
# This notebook works with or without an API key, but you will be given more permissive access to the data with an API key.
# The Planetary Computer Hub is pre-configured to use your API key.
import odc.stac
import planetary_computer
from pystac_client import Client
import rich.table
# ### Query for available data
#
# MODIS is a global dataset with a variety of products available within each larger category (vegetation, snow, fire, temperature, and reflectance).
# The [MODIS group](https://planetarycomputer.microsoft.com/dataset/group/modis) contains a complete listing of available collections.
# Each collection's id is in the format `modis-{product}-061`, where `product` is the MODIS product id.
# The `-061` suffix indicates that all of the MODIS collections are part of the [MODIS 6.1 update](https://atmosphere-imager.gsfc.nasa.gov/documentation/collection-61).
#
# We'll look at the snow cover around the Mount of the Holy Cross in Colorado and how it progresses throughout the winter, using the daily snow product (10A1).
# +
longitude = -106.481687
latitude = 39.466829
mount_of_the_holy_cross = [longitude, latitude]
geometry = {
"type": "Point",
"coordinates": mount_of_the_holy_cross,
}
datetimes = [
"2020-11",
"2020-12",
"2021-01",
"2021-02",
"2021-03",
"2021-04",
"2021-05",
"2021-06",
]
items = dict()
catalog = Client.open("https://planetarycomputer-staging.microsoft.com/api/stac/v1")
for datetime in datetimes:
print(f"Fetching {datetime}")
search = catalog.search(
collections=["modis-10A1-061"],
intersects=geometry,
datetime=datetime,
)
item = search.get_all_items()[0]
items[datetime] = planetary_computer.sign(item)
print(items)
# -
# ### Available assets
#
# Each item has several available assets, including the original HDF file and a Cloud-optimized GeoTIFF of each subdataset.
t = rich.table.Table("Key", "Title")
for key, asset in items["2020-11"].assets.items():
t.add_row(key, asset.title)
t
# ### Loading the snow cover data
#
# For this example, we'll visualize and compare the snow cover month to month.
# Let's grab each snow cover COG and load them into an xarray using [odc-stac](https://github.com/opendatacube/odc-stac).
# We'll also apply the scaling as defined by the `raster:bands` extension.
# The MODIS coordinate reference system is a [sinusoidal grid](https://modis-land.gsfc.nasa.gov/MODLAND_grid.html), which means that views in a naïve XY raster look skewed.
# For visualization purposes, we reproject to a [spherical Mercator projection](https://wiki.openstreetmap.org/wiki/EPSG:3857) for intuitive, north-up visualization.
#
# The NDSI Snow Cover values are defined as:
#
# ```
# 0–100: NDSI snow cover
# 200: missing data
# 201: no decision
# 211: night
# 237: inland water
# 239: ocean
# 250: cloud
# 254: detector saturated
# 255: fill
# ```
#
# We want to mask out all numbers greater than 100.
bbox = [longitude - 0.5, latitude - 0.5, longitude + 0.5, latitude + 0.5]
data = odc.stac.load(
items.values(),
crs="EPSG:3857",
bbox=bbox,
bands="NDSI_Snow_Cover",
resolution=500,
)
data = data.where(data <= 100, drop=True)
data
# ### Displaying the data
#
# Let's display the snow cover for each month.
# +
g = data["NDSI_Snow_Cover"].plot.imshow(
col="time", vmin=0, vmax=100, col_wrap=4, size=4
)
months = data["NDSI_Snow_Cover"].time.to_pandas().dt.strftime("%B %Y")
for ax, month in zip(g.axes.flat, months):
ax.set_title(month)
# -
# You'll notice there's a lot of missing values due to masking, probably due to clouds.
# More sophisticated analysis would use the `NDSI_Snow_Cover_Basic_QA` and other bands to merge information from multiple scenes, or pick the best scenes for analysis.
#
# ### Data pages
#
# These pages include links to the user guides:
#
# - MOD10A1: https://nsidc.org/data/MOD10A1/versions/61
# - MOD10A2: https://nsidc.org/data/MOD10A2/versions/61
|
datasets/modis/modis-snow-example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tushare
import tushare as ts
token = '<KEY>'
ts_client = ts.pro_api(token)
# +
df = ts_client.daily(ts_code='600000.SH', start_date='20180701', end_date='20180718')
# -
print(df)
|
Document/Notebook/tushare.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Partitioning a subset of Wikidata
#
# This notebook illustrates how to partition a Wikidata KGTK edges file.
#
# Parameters are set up in the first cell so that we can run this notebook in batch mode. Example invocation command:
#
# ```
# papermill partition-wikidata.ipynb partition-wikidata.out.ipynb \
# -p wikidata_input_path /data3/rogers/kgtk/gd/kgtk_public_graphs/cache/wikidata-20201130/data/all.tsv.gz \
# -p wikidata_parts_path /data3/rogers/kgtk/gd/kgtk_public_graphs/cache/wikidata-20201130/parts \
# ```
#
# Here is a sample of the records that might appear in the input KGTK file:
# ```
# id node1 label node2 rank node2;wikidatatype lang
# Q1-P1036-418bc4-78f5a565-0 Q1 P1036 "113" normal external-id
# Q1-P1343-Q19190511-ab132b87-0 Q1 P1343 Q19190511 normal wikibase-item
# Q1-P18-92a7b3-0dcac501-0 Q1 P18 "Hubble ultra deep field.jpg" normal commonsMedia
# Q1-P2386-cedfb0-0fdbd641-0 Q1 P2386 +880000000000000000000000Q828224 normal quantity
# Q1-P580-a2fccf-63cf4743-0 Q1 P580 ^-13798000000-00-00T00:00:00Z/3 normal time
# Q1-P920-47c0f2-52689c4e-0 Q1 P920 "LEM201201756" normal string
# Q1-P1343-Q19190511-ab132b87-0-P805-Q84065667-0 Q1-P1343-Q19190511-ab132b87-0 P805 Q84065667 wikibase-item
# Q1-P1343-Q88672152-5080b9e2-0-P304-5724c3-0 Q1-P1343-Q88672152-5080b9e2-0 P304 "13-36" string
# Q1-P2670-Q18343-030eb87e-0-P1107-ce87f8-0 Q1-P2670-Q18343-030eb87e-0 P1107 +0.70 quantity
# Q1-P793-Q273508-1900d69c-0-P585-a2fccf-0 Q1-P793-Q273508-1900d69c-0 P585 ^-13798000000-00-00T00:00:00Z/3 time
# P10-alias-en-282226-0 P10 alias 'gif'@en
# P10-description-en P10 description 'relevant video. For images, use the property P18. For film trailers, qualify with \"object has role\" (P3831)=\"trailer\" (Q622550)'@en en
# P10-label-en P10 label 'video'@en en
# Q1-addl_wikipedia_sitelink-19e42a-0 Q1 addl_wikipedia_sitelink http://enwikiquote.org/wiki/Universe en
# Q1-addl_wikipedia_sitelink-19e42a-0-language-0 Q1-addl_wikipedia_sitelink-19e42a-0 sitelink-language en en
# Q1-addl_wikipedia_sitelink-19e42a-0-site-0 Q1-addl_wikipedia_sitelink-19e42a-0 sitelink-site enwikiquote en
# Q1-addl_wikipedia_sitelink-19e42a-0-title-0 Q1-addl_wikipedia_sitelink-19e42a-0 sitelink-title "Universe" en
# Q1-wikipedia_sitelink-5e459a-0 Q1 wikipedia_sitelink http://en.wikipedia.org/wiki/Universe en
# Q1-wikipedia_sitelink-5e459a-0-badge-Q17437798 Q1-wikipedia_sitelink-5e459a-0 sitelink-badge Q17437798 en
# Q1-wikipedia_sitelink-5e459a-0-language-0 Q1-wikipedia_sitelink-5e459a-0 sitelink-language en en
# Q1-wikipedia_sitelink-5e459a-0-site-0 Q1-wikipedia_sitelink-5e459a-0 sitelink-site enwiki en
# Q1-wikipedia_sitelink-5e459a-0-title-0 Q1-wikipedia_sitelink-5e459a-0 sitelink-title "Universe" en
# ```
# Here are some contraints on the contents of the input file:
# - The input file starts with a KGTK header record.
# - In addition to the `id`, `node1`, `label`, and `node2` columns, the file may contain the `node2;wikidatatype` column.
# - The `node2;wikidatatype` column is used to partition claims by Wikidata property datatype.
# - If it does not exist, it will be created during the partitioning process and populated using `datatype` relationships.
# - If it does exist, any empty values in the column will be populated using `datatype` relationships.
# - The `id` column must contain a nonempty value.
# - The first section of an `id` value must be the `node` value for the record.
# - The qualifier extraction operations depend upon this constraint.
# - In addition to the claims and qualifiers, the input file is expected to contain:
# - English language labels for all property entities appearing in the file.
# - The input file ought to contain the following:
# - claims records,
# - qualifier records,
# - alias records in appropriate languages,
# - description records in appropriate languages,
# - label records in appropriate languages, and
# - sitelink records in appropriate languages.
# - `datatype` records that map Wikidata property entities to Wikidata property datatypes. These records are required if the input file does not contain the `node2;wikidatatype` column.
# - Additionally, this script provides for the appearance of `type` records in the input file.
# - `type` records that list all `entityId` values and identify them as properties or items. These records provides a correctness check on the operation of `kgtk import-wikidata`, and may be deprecated in the future.
# - The input file is assumed to be unsorted. If it is already sorted on the (`id` `node1` `label` `node2`) columns , then set the `presorted` parameter to `True` to shorten the execution time of this script.
# ### Parameters for invoking the notebook
#
# | Parameter | Description | Default |
# | --------- | ----------- | ------- |
# | `wikidata_input_path` | A folder containing the Wikidata KGTK edges to partition. | '/data4/rogers/elicit/cache/datasets/wikidata-20200803/data/all.tsv.gz' |
# | `wikidata_parts_path` | A folder to receive the partitioned Wikidata files, such as `part.wikibase-item.tsv.gz` | '/data4/rogers/elicit/cache/datasets/wikidata-20200803/parts' |
# | `temp_folder_path` | A folder that may be used for temporary files. | wikidata_parts_path + '/temp' |
# | `gzip_command` | The compression command for sorting. | 'pigz' (Note: use version 2.4 or later)|
# | `kgtk_command` | The kgtk commmand. | 'time kgtk' |
# | `kgtk_options` | The kgtk commmand options. | '--debug --timing' |
# | `kgtk_extension` | The file extension for generated KGTK files. Appending `.gz` implies gzip compression. | 'tsv.gz' |
# | `presorted` | When True, the input file is already sorted on the (`id` `node1` `label` `node2`) columns. | 'False' |
# | `sort_extras` | Extra parameters for the sort program. The default specifies a path for temporary files. Other useful parameters include '--buffer-size' and '--parallel'. | '--parallel 24 --buffer-size 30% --temporary-directory ' + temp_folder_path |
# | `use_mgzip` | When True, use the mgzip program where appropriate for faster compression. | 'True' |
# | `verbose` | When True, produce additional feedback messages. | 'True' |
#
# Note: if `pigz` version 2.4 (or later) is not available on your system, use `gzip`.
#
# + tags=["parameters"]
# Parameters
wikidata_input_path = '/data3/rogers/kgtk/gd/kgtk_public_graphs/cache/wikidata-20201130/data/all.tsv.gz'
wikidata_parts_path = '/data3/rogers/kgtk/gd/kgtk_public_graphs/cache/wikidata-20201130/parts'
temp_folder_path = wikidata_parts_path + '/temp'
gzip_command = 'pigz'
kgtk_command = 'time kgtk'
kgtk_options = '--debug --timing'
kgtk_extension = 'tsv.gz'
presorted = 'False'
sort_extras = '--parallel 24 --buffer-size 30% --temporary-directory ' + temp_folder_path
use_mgzip = 'True'
verbose = 'True'
# -
print('wikidata_input_path = %s' % repr(wikidata_input_path))
print('wikidata_parts_path = %s' % repr(wikidata_parts_path))
print('temp_folder_path = %s' % repr(temp_folder_path))
print('gzip_command = %s' % repr(gzip_command))
print('kgtk_command = %s' % repr(kgtk_command))
print('kgtk_options = %s' % repr(kgtk_options))
print('kgtk_extension = %s' % repr(kgtk_extension))
print('presorted = %s' % repr(presorted))
print('sort_extras = %s' % repr(sort_extras))
print('use_mgzip = %s' % repr(use_mgzip))
print('verbose = %s' % repr(verbose))
# ### Create working folders and empty them
# !mkdir {wikidata_parts_path}
# !mkdir {temp_folder_path}
# !rm {wikidata_parts_path}/*.tsv {wikidata_parts_path}/*.tsv.gz
# !rm {temp_folder_path}/*.tsv {temp_folder_path}/*.tsv.gz
# ### Sort the Input Data Unless Presorted
# Sort the input data file by (id, node1, label, node2).
# This may take a while.
if presorted.lower() == "true":
print('Using a presorted input file %s.' % repr(wikidata_input_path))
partition_input_file = wikidata_input_path
else:
print('Sorting the input file %s.' % repr(wikidata_input_path))
partition_input_file = wikidata_parts_path + '/all.' + kgtk_extension
# !{kgtk_command} {kgtk_options} sort2 --verbose={verbose} --gzip-command={gzip_command} \
# --input-file {wikidata_input_path} \
# --output-file {partition_input_file} \
# --columns id node1 label node2 \
# --extra "{sort_extras}"
# ### Partition the Claims, Qualifiers, and Entity Data
# Split out the entity data (alias, description, label, and sitelinks) and additional metadata (datatype, type). Separate the qualifiers from the claims.
#
# !{kgtk_command} {kgtk_options} filter --verbose={verbose} --use-mgzip={use_mgzip} --first-match-only \
# --input-file {partition_input_file} \
# -p '; datatype ;' -o {wikidata_parts_path}/metadata.property.datatypes.{kgtk_extension} \
# -p '; alias ;' -o {wikidata_parts_path}/aliases.{kgtk_extension} \
# -p '; description ;' -o {wikidata_parts_path}/descriptions.{kgtk_extension} \
# -p '; label ;' -o {wikidata_parts_path}/labels.{kgtk_extension} \
# -p '; addl_wikipedia_sitelink,wikipedia_sitelink ;' \
# -o {wikidata_parts_path}/sitelinks.{kgtk_extension} \
# -p '; sitelink-badge,sitelink-language,sitelink-site,sitelink-title ;' \
# -o {wikidata_parts_path}/sitelinks.qualifiers.{kgtk_extension} \
# -p '; type ;' -o {wikidata_parts_path}/metadata.types.{kgtk_extension} \
# --reject-file {temp_folder_path}/claims-and-qualifiers.sorted-by-id.{kgtk_extension}
# ### Sort the claims and qualifiers on Node1
# Sort the combined claims and qualifiers file by the node1 column.
# This may take a while.
# !{kgtk_command} {kgtk_options} sort2 --verbose={verbose} --gzip-command={gzip_command} \
# --input-file {temp_folder_path}/claims-and-qualifiers.sorted-by-id.{kgtk_extension} \
# --output-file {temp_folder_path}/claims-and-qualifiers.sorted-by-node1.{kgtk_extension}\
# --columns node1 \
# --extra "{sort_extras}"
# ### Split the claims and qualifiers
# If row A's node1 value matches some other row's id value, the then row A is a qualifier.
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {temp_folder_path}/claims-and-qualifiers.sorted-by-node1.{kgtk_extension} \
# --filter-file {temp_folder_path}/claims-and-qualifiers.sorted-by-id.{kgtk_extension} \
# --output-file {temp_folder_path}/qualifiers.sorted-by-node1.{kgtk_extension}\
# --reject-file {temp_folder_path}/claims.sorted-by-node1.{kgtk_extension}\
# --input-keys node1 \
# --filter-keys id
# ### Sort the claims by ID
# Sort the split claims by id, node1, label, node2.
# This may take a while.
# !{kgtk_command} {kgtk_options} sort2 --verbose={verbose} --gzip-command={gzip_command} \
# --input-file {temp_folder_path}/claims.sorted-by-node1.{kgtk_extension} \
# --output-file {temp_folder_path}/claims.no-datatype.{kgtk_extension}\
# --columns id node1 label node2 \
# --extra "{sort_extras}"
# ### Merge the Wikidata Property Datatypes into the claims
# Merge the Wikidata Property Datatypes into the claims row as node2;wikidatatype. This column will be used to partition the claims by Wikidata Property Datatype ina later step. If the claims file already has a node2;wikidatatype column, lift only when that column has an empty value.
#
# !{kgtk_command} {kgtk_options} lift --verbose={verbose} --use-mgzip={use_mgzip} \
# --input-file {temp_folder_path}/claims.no-datatype.{kgtk_extension} \
# --columns-to-lift label \
# --overwrite False \
# --label-file {wikidata_parts_path}/metadata.property.datatypes.{kgtk_extension}\
# --label-value datatype \
# --output-file {wikidata_parts_path}/claims.{kgtk_extension}\
# --columns-to-write 'node2;wikidatatype'
# ### Sort the qualifiers by ID
# Sort the split qualifiers by id, node1, label, node2.
# This may take a while.
# !{kgtk_command} {kgtk_options} sort2 --verbose={verbose} --gzip-command={gzip_command} \
# --input-file {temp_folder_path}/qualifiers.sorted-by-node1.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.{kgtk_extension}\
# --columns id node1 label node2 \
# --extra "{sort_extras}"
# ### Extract the English aliases, descriptions, labels, and sitelinks.
# Aliases, descriptions, and labels are extracted by selecting rows where the `node2` value ends in the language suffix for English (`@en`) in a KGTK language-qualified string. This is an abbreviated pattern; a more general pattern would include the single quotes used to delimit a KGTK language-qualified string. If `kgtk import-wikidata` has executed properly, the abbreviated pattern should be sufficient.
#
# Sitelink rows do not have a language-specific marker in the `node2` value. We use the `lang` column to provide the language code for English ('en'). The `lang` column is an additional column created by `kgtk import-wikidata`.
# !{kgtk_command} {kgtk_options} filter --verbose={verbose} --use-mgzip={use_mgzip} --regex \
# --input-file {wikidata_parts_path}/aliases.{kgtk_extension} \
# -p ';; ^.*@en$' -o {wikidata_parts_path}/aliases.en.{kgtk_extension}
# !{kgtk_command} {kgtk_options} filter --verbose={verbose} --use-mgzip={use_mgzip} --regex \
# --input-file {wikidata_parts_path}/descriptions.{kgtk_extension} \
# -p ';; ^.*@en$' -o {wikidata_parts_path}/descriptions.en.{kgtk_extension}
# !{kgtk_command} {kgtk_options} filter --verbose={verbose} --use-mgzip={use_mgzip} --regex \
# --input-file {wikidata_parts_path}/labels.{kgtk_extension} \
# -p ';; ^.*@en$' -o {wikidata_parts_path}/labels.en.{kgtk_extension}
# !{kgtk_command} {kgtk_options} filter --verbose={verbose} --use-mgzip={use_mgzip} \
# --input-file {wikidata_parts_path}/sitelinks.qualifiers.{kgtk_extension} \
# -p '; sitelink-language ; en' -o {temp_folder_path}/sitelinks.language.en.{kgtk_extension}
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/sitelinks.{kgtk_extension} \
# --filter-on {temp_folder_path}/sitelinks.language.en.{kgtk_extension} \
# --output-file {wikidata_parts_path}/sitelinks.en.{kgtk_extension} \
# --input-keys id \
# --filter-keys node1
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/sitelinks.qualifiers.{kgtk_extension} \
# --filter-on {temp_folder_path}/sitelinks.language.en.{kgtk_extension} \
# --output-file {wikidata_parts_path}/sitelinks.qualifiers.en.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys node1
# ### Partition the claims by Wikidata Property Datatype
# Wikidata has two names for each Wikidata property datatype: the name that appears in the JSON dump file, and the name that appears in the TTL dump file. `kgtk import-wikidata` currently imports rows from Wikikdata JSON dump files, and these are the names that appear below.
#
# The `part.other` file catches any records that have an unknown Wikidata property datatype. Additional Wikidata property datatypes may occur when processing from certain Wikidata extensions.
# !{kgtk_command} {kgtk_options} filter --verbose={verbose} --use-mgzip={use_mgzip} --first-match-only \
# --input-file {wikidata_parts_path}/claims.{kgtk_extension} \
# --obj 'node2;wikidatatype' \
# -p ';; commonsMedia' -o {wikidata_parts_path}/claims.commonsMedia.{kgtk_extension} \
# -p ';; external-id' -o {wikidata_parts_path}/claims.external-id.{kgtk_extension} \
# -p ';; geo-shape' -o {wikidata_parts_path}/claims.geo-shape.{kgtk_extension} \
# -p ';; globe-coordinate' -o {wikidata_parts_path}/claims.globe-coordinate.{kgtk_extension} \
# -p ';; math' -o {wikidata_parts_path}/claims.math.{kgtk_extension} \
# -p ';; monolingualtext' -o {wikidata_parts_path}/claims.monolingualtext.{kgtk_extension} \
# -p ';; musical-notation' -o {wikidata_parts_path}/claims.musical-notation.{kgtk_extension} \
# -p ';; quantity' -o {wikidata_parts_path}/claims.quantity.{kgtk_extension} \
# -p ';; string' -o {wikidata_parts_path}/claims.string.{kgtk_extension} \
# -p ';; tabular-data' -o {wikidata_parts_path}/claims.tabular-data.{kgtk_extension} \
# -p ';; time' -o {wikidata_parts_path}/claims.time.{kgtk_extension} \
# -p ';; url' -o {wikidata_parts_path}/claims.url.{kgtk_extension} \
# -p ';; wikibase-form' -o {wikidata_parts_path}/claims.wikibase-form.{kgtk_extension} \
# -p ';; wikibase-item' -o {wikidata_parts_path}/claims.wikibase-item.{kgtk_extension} \
# -p ';; wikibase-lexeme' -o {wikidata_parts_path}/claims.wikibase-lexeme.{kgtk_extension} \
# -p ';; wikibase-property' -o {wikidata_parts_path}/claims.wikibase-property.{kgtk_extension} \
# -p ';; wikibase-sense' -o {wikidata_parts_path}/claims.wikibase-sense.{kgtk_extension} \
# --reject-file {wikidata_parts_path}/claims.other.{kgtk_extension}
# ### Partition the qualifiers
# Extract the qualifier records for each of the Wikidata property datatype partition files.
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.commonsMedia.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.commonsMedia.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.external-id.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.external-id.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.geo-shape.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.geo-shape.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.globe-coordinate.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.globe-coordinate.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.math.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.math.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.monolingualtext.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.monolingualtext.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.musical-notation.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.musical-notation.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.quantity.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.quantity.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.string.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.string.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.tabular-data.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.tabular-data.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.time.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.time.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.url.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.url.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.wikibase-form.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.wikibase-form.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.wikibase-item.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.wikibase-item.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.wikibase-lexeme.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.wikibase-lexeme.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.wikibase-property.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.wikibase-property.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
# !{kgtk_command} {kgtk_options} ifexists --verbose={verbose} --use-mgzip={use_mgzip} --presorted \
# --input-file {wikidata_parts_path}/qualifiers.{kgtk_extension} \
# --filter-on {wikidata_parts_path}/claims.wikibase-sense.{kgtk_extension} \
# --output-file {wikidata_parts_path}/qualifiers.wikibase-sense.{kgtk_extension} \
# --input-keys node1 \
# --filter-keys id
|
examples/partition-wikidata.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Keras Callbacks and Functional API
# +
from tensorflow.keras.datasets import cifar10
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation, Dropout
from tensorflow.keras.optimizers import SGD, RMSprop
from tensorflow.keras.callbacks import EarlyStopping, TensorBoard, ModelCheckpoint
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# +
(X_train_t, y_train), (X_test_t, y_test) = cifar10.load_data()
X_train_t = X_train_t.astype('float32') / 255.
X_test_t = X_test_t.astype('float32') / 255.
X_train = X_train_t.reshape(len(X_train_t), 32*32*3)
X_test = X_test_t.reshape(len(X_test_t), 32*32*3)
# -
print("Training set:")
print("Tensor images shape:\t", X_train_t.shape)
print("Flat images shape:\t", X_train.shape)
print("Labels shape:\t\t", y_train.shape)
plt.figure(figsize=(15, 4))
for i in range(0, 8):
plt.subplot(1, 8, i+1)
plt.imshow(X_train[i].reshape(32, 32, 3))
plt.title(y_train[i, 0])
# ## Callbacks on a simple model
# +
outpath='/tmp/tensorboard/cifar/'
early_stopper = EarlyStopping(monitor='val_accuracy', patience=10)
tensorboard = TensorBoard(outpath, histogram_freq=1)
checkpointer = ModelCheckpoint(outpath+'weights_epoch_{epoch:02d}_val_accuracy_{val_accuracy:.2f}.hdf5',
monitor='val_acc')
# +
model = Sequential([
Dense(1024, activation='relu', input_dim=3072),
Dense(512, activation='relu'),
Dense(10, activation='softmax')
])
model.compile(loss='sparse_categorical_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
# -
model.fit(X_train, y_train,
batch_size=128,
epochs=10,
verbose=1,
validation_split=0.1,
callbacks=[early_stopper,
tensorboard,
checkpointer])
import os
sorted(os.listdir(outpath))
# Now check the tensorboard.
#
# - If using provided instance, just browse to: `http://<your-domain>/tensorboard/`
#
# - If using local, open a terminal, activate the environment and run:
# ```
# tensorboard --logdir=/tmp/tensorboard/cifar/
# ```
# then open a browser at `localhost:6006`
#
# You should see something like this:
#
# 
#
# If you can't access port 6006, you can use Local Tunel. Open another terminal and run [localtunnel](https://localtunnel.github.io/www/) on port 6006:
# ```
# lt --port 6006
# ```
# 3. Go to the url provided
#
# You should see something like this.
#
# > TIP: if you get an error `lt: command not found` install localtunnel as:
# ```
# sudo npm install -g localtunnel
# ```
# ## Exercise 1: Keras functional API
#
# We'e built a model using the `Sequential API` from tensorflow.keras. Keras also offers a [functional API](https://keras.io/getting-started/functional-api-guide/). This API is the way to go for defining complex models, such as multi-output models, directed acyclic graphs, or models with shared layers.
#
# Can you rewrite the model above using the functional API?
from tensorflow.keras.layers import Input
from tensorflow.keras.models import Model
# + tags=["solution", "empty"]
inputs = Input(shape=(3072,))
x = Dense(1024, activation='relu')(inputs)
x = Dense(512, activation='relu')(x)
predictions = Dense(10, activation='softmax')(x)
model = Model(inputs=inputs, outputs=predictions)
# +
model.compile(loss='sparse_categorical_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
model.fit(X_train, y_train,
batch_size=128,
epochs=10,
verbose=1,
validation_split=0.1)
# Final test evaluation
score = model.evaluate(X_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# -
# ## Exercise 2: Convolutional Model with Functional API
#
# The above model is a very simple fully connected deep neural network. As we have seen, Convolutional Neural Networks are much more powerful when dealing with images. The original data has shape:
#
# (N_images, Height, Width, Channels)
#
# Can you write a convolutional model using the functional API?
#
# Bonus points if you use callbacks to write data to tensorboard.
from tensorflow.keras.layers import Dense, Dropout, Activation
from tensorflow.keras.layers import Conv2D, MaxPool2D, AveragePooling2D, Flatten
outpath_conv = outpath.replace('cifar', 'cifar_conv')
tensorboard_conv = TensorBoard(outpath_conv, histogram_freq=1)
# + tags=["solution", "empty"]
inputs = Input(shape=(32, 32, 3))
x = Conv2D(32, (3, 3),
padding='same',
activation='relu')(inputs)
x = Conv2D(32, (3, 3), activation='relu')(x)
x = MaxPool2D(pool_size=(2, 2))(x)
x = Conv2D(64, (3, 3), padding='same', activation='relu')(x)
x = Conv2D(64, (3, 3), activation='relu')(x)
x = MaxPool2D(pool_size=(2, 2))(x)
x = Flatten()(x)
x = Dense(512, activation='relu')(x)
predictions = Dense(10, activation='softmax')(x)
model = Model(inputs=inputs, outputs=predictions)
model.compile(loss='sparse_categorical_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
# Fit your model
model.fit(X_train_t, y_train,
batch_size=32,
epochs=10,
validation_split=0.1,
shuffle=True,
verbose=1,
callbacks=[tensorboard_conv]
)
# Final test evaluation
score = model.evaluate(X_test_t, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# -
# ## Exrcise 3: Discuss with the person next to you
#
# 1. What are the pros/cons of the sequential API?
# - What are the pros/cons of the functional API?
# - What are the key differences between a Fully connected and a Convolutional neural network?
# - What is a dropout layer? How does it work? Why does it help?
#
|
solutions_do_not_open/Lab_15_DL Keras Callbacks and Functional API_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Time-frequency beamforming using LCMV
#
#
# Compute LCMV source power [1]_ in a grid of time-frequency windows and
# display results.
#
# References
# ----------
# .. [1] Dalal et al. Five-dimensional neuroimaging: Localization of the
# time-frequency dynamics of cortical activity.
# NeuroImage (2008) vol. 40 (4) pp. 1686-1700
#
#
# +
# Author: <NAME> <<EMAIL>>
#
# License: BSD (3-clause)
import mne
from mne import compute_covariance
from mne.datasets import sample
from mne.event import make_fixed_length_events
from mne.beamformer import tf_lcmv
from mne.viz import plot_source_spectrogram
print(__doc__)
data_path = sample.data_path()
raw_fname = data_path + '/MEG/sample/sample_audvis_raw.fif'
noise_fname = data_path + '/MEG/sample/ernoise_raw.fif'
event_fname = data_path + '/MEG/sample/sample_audvis_raw-eve.fif'
fname_fwd = data_path + '/MEG/sample/sample_audvis-meg-eeg-oct-6-fwd.fif'
subjects_dir = data_path + '/subjects'
label_name = 'Aud-lh'
fname_label = data_path + '/MEG/sample/labels/%s.label' % label_name
# -
# Read raw data, preload to allow filtering
#
#
# +
raw = mne.io.read_raw_fif(raw_fname, preload=True)
raw.info['bads'] = ['MEG 2443'] # 1 bad MEG channel
# Pick a selection of magnetometer channels. A subset of all channels was used
# to speed up the example. For a solution based on all MEG channels use
# meg=True, selection=None and add grad=4000e-13 to the reject dictionary.
# We could do this with a "picks" argument to Epochs and the LCMV functions,
# but here we use raw.pick_types() to save memory.
left_temporal_channels = mne.read_selection('Left-temporal')
raw.pick_types(meg='mag', eeg=False, eog=False, stim=False, exclude='bads',
selection=left_temporal_channels)
reject = dict(mag=4e-12)
# Re-normalize our empty-room projectors, which should be fine after
# subselection
raw.info.normalize_proj()
# Setting time limits for reading epochs. Note that tmin and tmax are set so
# that time-frequency beamforming will be performed for a wider range of time
# points than will later be displayed on the final spectrogram. This ensures
# that all time bins displayed represent an average of an equal number of time
# windows.
tmin, tmax = -0.5, 0.75 # s
tmin_plot, tmax_plot = -0.3, 0.5 # s
# Read epochs. Note that preload is set to False to enable tf_lcmv to read the
# underlying raw object.
# Filtering is then performed on raw data in tf_lcmv and the epochs
# parameters passed here are used to create epochs from filtered data. However,
# reading epochs without preloading means that bad epoch rejection is delayed
# until later. To perform bad epoch rejection based on the reject parameter
# passed here, run epochs.drop_bad(). This is done automatically in
# tf_lcmv to reject bad epochs based on unfiltered data.
event_id = 1
events = mne.read_events(event_fname)
epochs = mne.Epochs(raw, events, event_id, tmin, tmax, proj=True,
baseline=None, preload=False, reject=reject)
# Read empty room noise, preload to allow filtering, and pick subselection
raw_noise = mne.io.read_raw_fif(noise_fname, preload=True)
raw_noise.info['bads'] = ['MEG 2443'] # 1 bad MEG channel
raw_noise.pick_types(meg='mag', eeg=False, eog=False, stim=False,
exclude='bads', selection=left_temporal_channels)
raw_noise.info.normalize_proj()
# Create artificial events for empty room noise data
events_noise = make_fixed_length_events(raw_noise, event_id, duration=1.)
# Create an epochs object using preload=True to reject bad epochs based on
# unfiltered data
epochs_noise = mne.Epochs(raw_noise, events_noise, event_id, tmin, tmax,
proj=True, baseline=None,
preload=True, reject=reject)
# Make sure the number of noise epochs is the same as data epochs
epochs_noise = epochs_noise[:len(epochs.events)]
# Read forward operator
forward = mne.read_forward_solution(fname_fwd)
# Read label
label = mne.read_label(fname_label)
# -
# Time-frequency beamforming based on LCMV
#
#
# +
# Setting frequency bins as in Dalal et al. 2008 (high gamma was subdivided)
freq_bins = [(4, 12), (12, 30), (30, 55), (65, 299)] # Hz
win_lengths = [0.3, 0.2, 0.15, 0.1] # s
# Setting the time step
tstep = 0.05
# Setting the whitened data covariance regularization parameter
data_reg = 0.001
# Subtract evoked response prior to computation?
subtract_evoked = False
# Calculating covariance from empty room noise. To use baseline data as noise
# substitute raw for raw_noise, epochs.events for epochs_noise.events, tmin for
# desired baseline length, and 0 for tmax_plot.
# Note, if using baseline data, the averaged evoked response in the baseline
# period should be flat.
noise_covs = []
for (l_freq, h_freq) in freq_bins:
raw_band = raw_noise.copy()
raw_band.filter(l_freq, h_freq, n_jobs=1, fir_design='firwin')
epochs_band = mne.Epochs(raw_band, epochs_noise.events, event_id,
tmin=tmin_plot, tmax=tmax_plot, baseline=None,
proj=True)
noise_cov = compute_covariance(epochs_band, method='shrunk', rank=None)
noise_covs.append(noise_cov)
del raw_band # to save memory
# Computing LCMV solutions for time-frequency windows in a label in source
# space for faster computation, use label=None for full solution
stcs = tf_lcmv(epochs, forward, noise_covs, tmin, tmax, tstep, win_lengths,
freq_bins=freq_bins, subtract_evoked=subtract_evoked,
reg=data_reg, label=label, rank=None)
# Plotting source spectrogram for source with maximum activity.
# Note that tmin and tmax are set to display a time range that is smaller than
# the one for which beamforming estimates were calculated. This ensures that
# all time bins shown are a result of smoothing across an identical number of
# time windows.
plot_source_spectrogram(stcs, freq_bins, tmin=tmin_plot, tmax=tmax_plot,
source_index=None, colorbar=True)
|
0.17/_downloads/793e913bab4b20ad5f4863ca669c17f8/plot_tf_lcmv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# *Python Machine Learning 3rd Edition* by [<NAME>](https://sebastianraschka.com), Packt Publishing Ltd. 2019
#
# Code Repository: https://github.com/rasbt/python-machine-learning-book-3rd-edition
#
# Code License: [MIT License](https://github.com/rasbt/python-machine-learning-book-3rd-edition/blob/master/LICENSE.txt)
# # Python Machine Learning - Code Examples
# # Chapter 10 - Predicting Continuous Target Variables with Regression Analysis
# Note that the optional watermark extension is a small IPython notebook plugin that I developed to make the code reproducible. You can just skip the following line(s).
# %load_ext watermark
# %watermark -a "<NAME>" -u -d -v -p numpy,pandas,matplotlib,sklearn,mlxtend
# *The use of `watermark` is optional. You can install this IPython extension via "`pip install watermark`". For more information, please see: https://github.com/rasbt/watermark.*
# The mlxtend package (http://rasbt.github.io/mlxtend/), which contains a few useful functions on top of scikit-learn and matplotloib, can be installed via
#
# conda install mlxtend
#
# or
#
# pip install mlxtend
# <br>
# <br>
# ### Overview
# - [Introducing regression](#Introducing-linear-regression)
# - [Simple linear regression](#Simple-linear-regression)
# - [Exploring the Housing Dataset](#Exploring-the-Housing-Dataset)
# - [Loading the Housing dataset into a data frame](Loading-the-Housing-dataset-into-a-data-frame)
# - [Visualizing the important characteristics of a dataset](#Visualizing-the-important-characteristics-of-a-dataset)
# - [Implementing an ordinary least squares linear regression model](#Implementing-an-ordinary-least-squares-linear-regression-model)
# - [Solving regression for regression parameters with gradient descent](#Solving-regression-for-regression-parameters-with-gradient-descent)
# - [Estimating the coefficient of a regression model via scikit-learn](#Estimating-the-coefficient-of-a-regression-model-via-scikit-learn)
# - [Fitting a robust regression model using RANSAC](#Fitting-a-robust-regression-model-using-RANSAC)
# - [Evaluating the performance of linear regression models](#Evaluating-the-performance-of-linear-regression-models)
# - [Using regularized methods for regression](#Using-regularized-methods-for-regression)
# - [Turning a linear regression model into a curve - polynomial regression](#Turning-a-linear-regression-model-into-a-curve---polynomial-regression)
# - [Modeling nonlinear relationships in the Housing Dataset](#Modeling-nonlinear-relationships-in-the-Housing-Dataset)
# - [Dealing with nonlinear relationships using random forests](#Dealing-with-nonlinear-relationships-using-random-forests)
# - [Decision tree regression](#Decision-tree-regression)
# - [Random forest regression](#Random-forest-regression)
# - [Summary](#Summary)
# <br>
# <br>
from IPython.display import Image
# %matplotlib inline
# # Introducing linear regression
# ## Simple linear regression
Image(filename='images/10_01.png', width=500)
# ## Multiple linear regression
Image(filename='images/10_15.png', width=500)
# <br>
# <br>
# # Exploring the Housing dataset
# ## Loading the Housing dataset into a data frame
# Description, which was previously available at: [https://archive.ics.uci.edu/ml/datasets/Housing](https://archive.ics.uci.edu/ml/datasets/Housing)
#
# Attributes:
#
# <pre>
# 1. CRIM per capita crime rate by town
# 2. ZN proportion of residential land zoned for lots over
# 25,000 sq.ft.
# 3. INDUS proportion of non-retail business acres per town
# 4. CHAS Charles River dummy variable (= 1 if tract bounds
# river; 0 otherwise)
# 5. NOX nitric oxides concentration (parts per 10 million)
# 6. RM average number of rooms per dwelling
# 7. AGE proportion of owner-occupied units built prior to 1940
# 8. DIS weighted distances to five Boston employment centres
# 9. RAD index of accessibility to radial highways
# 10. TAX full-value property-tax rate per $10,000
# 11. PTRATIO pupil-teacher ratio by town
# 12. B 1000(Bk - 0.63)^2 where Bk is the proportion of blacks
# by town
# 13. LSTAT % lower status of the population
# 14. MEDV Median value of owner-occupied homes in $1000s
# </pre>
# +
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/rasbt/'
'python-machine-learning-book-3rd-edition/'
'master/ch10/housing.data.txt',
header=None,
sep='\s+')
df.columns = ['CRIM', 'ZN', 'INDUS', 'CHAS',
'NOX', 'RM', 'AGE', 'DIS', 'RAD',
'TAX', 'PTRATIO', 'B', 'LSTAT', 'MEDV']
df.head()
# -
# <hr>
#
# ### Note:
#
#
# You can find a copy of the housing dataset (and all other datasets used in this book) in the code bundle of this book, which you can use if you are working offline or the UCI server at https://archive.ics.uci.edu/ml/machine-learning-databases/housing/housing.data is temporarily unavailable. For instance, to load the housing dataset from a local directory, you can replace the lines
# df = pd.read_csv('https://archive.ics.uci.edu/ml/'
# 'machine-learning-databases'
# '/housing/housing.data',
# sep='\s+')
# in the following code example by
# df = pd.read_csv('./housing.data',
# sep='\s+')
# <br>
# <br>
# ## Visualizing the important characteristics of a dataset
import matplotlib.pyplot as plt
from mlxtend.plotting import scatterplotmatrix
# +
cols = ['LSTAT', 'INDUS', 'NOX', 'RM', 'MEDV']
scatterplotmatrix(df[cols].values, figsize=(10, 8),
names=cols, alpha=0.5)
plt.tight_layout()
#plt.savefig('images/10_03.png', dpi=300)
plt.show()
# +
import numpy as np
from mlxtend.plotting import heatmap
cm = np.corrcoef(df[cols].values.T)
hm = heatmap(cm, row_names=cols, column_names=cols)
# plt.savefig('images/10_04.png', dpi=300)
plt.show()
# -
# <br>
# <br>
# # Implementing an ordinary least squares linear regression model
# ...
# ## Solving regression for regression parameters with gradient descent
class LinearRegressionGD(object):
def __init__(self, eta=0.001, n_iter=20):
self.eta = eta
self.n_iter = n_iter
def fit(self, X, y):
self.w_ = np.zeros(1 + X.shape[1])
self.cost_ = []
for i in range(self.n_iter):
output = self.net_input(X)
errors = (y - output)
self.w_[1:] += self.eta * X.T.dot(errors)
self.w_[0] += self.eta * errors.sum()
cost = (errors**2).sum() / 2.0
self.cost_.append(cost)
return self
def net_input(self, X):
return np.dot(X, self.w_[1:]) + self.w_[0]
def predict(self, X):
return self.net_input(X)
X = df[['RM']].values
y = df['MEDV'].values
# +
from sklearn.preprocessing import StandardScaler
sc_x = StandardScaler()
sc_y = StandardScaler()
X_std = sc_x.fit_transform(X)
y_std = sc_y.fit_transform(y[:, np.newaxis]).flatten()
# -
lr = LinearRegressionGD()
lr.fit(X_std, y_std)
plt.plot(range(1, lr.n_iter+1), lr.cost_)
plt.ylabel('SSE')
plt.xlabel('Epoch')
#plt.tight_layout()
#plt.savefig('images/10_05.png', dpi=300)
plt.show()
def lin_regplot(X, y, model):
plt.scatter(X, y, c='steelblue', edgecolor='white', s=70)
plt.plot(X, model.predict(X), color='black', lw=2)
return
# +
lin_regplot(X_std, y_std, lr)
plt.xlabel('Average number of rooms [RM] (standardized)')
plt.ylabel('Price in $1000s [MEDV] (standardized)')
#plt.savefig('images/10_06.png', dpi=300)
plt.show()
# -
print('Slope: %.3f' % lr.w_[1])
print('Intercept: %.3f' % lr.w_[0])
num_rooms_std = sc_x.transform(np.array([[5.0]]))
price_std = lr.predict(num_rooms_std)
print("Price in $1000s: %.3f" % sc_y.inverse_transform(price_std))
# <br>
# <br>
# ## Estimating the coefficient of a regression model via scikit-learn
from sklearn.linear_model import LinearRegression
slr = LinearRegression()
slr.fit(X, y)
y_pred = slr.predict(X)
print('Slope: %.3f' % slr.coef_[0])
print('Intercept: %.3f' % slr.intercept_)
# +
lin_regplot(X, y, slr)
plt.xlabel('Average number of rooms [RM]')
plt.ylabel('Price in $1000s [MEDV]')
#plt.savefig('images/10_07.png', dpi=300)
plt.show()
# -
# **Normal Equations** alternative:
# +
# adding a column vector of "ones"
Xb = np.hstack((np.ones((X.shape[0], 1)), X))
w = np.zeros(X.shape[1])
z = np.linalg.inv(np.dot(Xb.T, Xb))
w = np.dot(z, np.dot(Xb.T, y))
print('Slope: %.3f' % w[1])
print('Intercept: %.3f' % w[0])
# -
# <br>
# <br>
# # Fitting a robust regression model using RANSAC
# +
from sklearn.linear_model import RANSACRegressor
ransac = RANSACRegressor(LinearRegression(),
max_trials=100,
min_samples=50,
loss='absolute_loss',
residual_threshold=5.0,
random_state=0)
ransac.fit(X, y)
inlier_mask = ransac.inlier_mask_
outlier_mask = np.logical_not(inlier_mask)
line_X = np.arange(3, 10, 1)
line_y_ransac = ransac.predict(line_X[:, np.newaxis])
plt.scatter(X[inlier_mask], y[inlier_mask],
c='steelblue', edgecolor='white',
marker='o', label='Inliers')
plt.scatter(X[outlier_mask], y[outlier_mask],
c='limegreen', edgecolor='white',
marker='s', label='Outliers')
plt.plot(line_X, line_y_ransac, color='black', lw=2)
plt.xlabel('Average number of rooms [RM]')
plt.ylabel('Price in $1000s [MEDV]')
plt.legend(loc='upper left')
#plt.savefig('images/10_08.png', dpi=300)
plt.show()
# -
print('Slope: %.3f' % ransac.estimator_.coef_[0])
print('Intercept: %.3f' % ransac.estimator_.intercept_)
# <br>
# <br>
# # Evaluating the performance of linear regression models
# +
from sklearn.model_selection import train_test_split
X = df.iloc[:, :-1].values
y = df['MEDV'].values
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.3, random_state=0)
# +
slr = LinearRegression()
slr.fit(X_train, y_train)
y_train_pred = slr.predict(X_train)
y_test_pred = slr.predict(X_test)
# +
import numpy as np
import scipy as sp
ary = np.array(range(100000))
# -
# %timeit np.linalg.norm(ary)
# %timeit sp.linalg.norm(ary)
# %timeit np.sqrt(np.sum(ary**2))
# +
plt.scatter(y_train_pred, y_train_pred - y_train,
c='steelblue', marker='o', edgecolor='white',
label='Training data')
plt.scatter(y_test_pred, y_test_pred - y_test,
c='limegreen', marker='s', edgecolor='white',
label='Test data')
plt.xlabel('Predicted values')
plt.ylabel('Residuals')
plt.legend(loc='upper left')
plt.hlines(y=0, xmin=-10, xmax=50, color='black', lw=2)
plt.xlim([-10, 50])
plt.tight_layout()
# plt.savefig('images/10_09.png', dpi=300)
plt.show()
# +
from sklearn.metrics import r2_score
from sklearn.metrics import mean_squared_error
print('MSE train: %.3f, test: %.3f' % (
mean_squared_error(y_train, y_train_pred),
mean_squared_error(y_test, y_test_pred)))
print('R^2 train: %.3f, test: %.3f' % (
r2_score(y_train, y_train_pred),
r2_score(y_test, y_test_pred)))
# -
# <br>
# <br>
# # Using regularized methods for regression
# +
from sklearn.linear_model import Lasso
lasso = Lasso(alpha=0.1)
lasso.fit(X_train, y_train)
y_train_pred = lasso.predict(X_train)
y_test_pred = lasso.predict(X_test)
print(lasso.coef_)
# -
print('MSE train: %.3f, test: %.3f' % (
mean_squared_error(y_train, y_train_pred),
mean_squared_error(y_test, y_test_pred)))
print('R^2 train: %.3f, test: %.3f' % (
r2_score(y_train, y_train_pred),
r2_score(y_test, y_test_pred)))
# Ridge regression:
from sklearn.linear_model import Ridge
ridge = Ridge(alpha=1.0)
# LASSO regression:
from sklearn.linear_model import Lasso
lasso = Lasso(alpha=1.0)
# Elastic Net regression:
from sklearn.linear_model import ElasticNet
elanet = ElasticNet(alpha=1.0, l1_ratio=0.5)
# <br>
# <br>
# # Turning a linear regression model into a curve - polynomial regression
# +
X = np.array([258.0, 270.0, 294.0,
320.0, 342.0, 368.0,
396.0, 446.0, 480.0, 586.0])\
[:, np.newaxis]
y = np.array([236.4, 234.4, 252.8,
298.6, 314.2, 342.2,
360.8, 368.0, 391.2,
390.8])
# +
from sklearn.preprocessing import PolynomialFeatures
lr = LinearRegression()
pr = LinearRegression()
quadratic = PolynomialFeatures(degree=2)
X_quad = quadratic.fit_transform(X)
# +
# fit linear features
lr.fit(X, y)
X_fit = np.arange(250, 600, 10)[:, np.newaxis]
y_lin_fit = lr.predict(X_fit)
# fit quadratic features
pr.fit(X_quad, y)
y_quad_fit = pr.predict(quadratic.fit_transform(X_fit))
# plot results
plt.scatter(X, y, label='Training points')
plt.plot(X_fit, y_lin_fit, label='Linear fit', linestyle='--')
plt.plot(X_fit, y_quad_fit, label='Quadratic fit')
plt.xlabel('Explanatory variable')
plt.ylabel('Predicted or known target values')
plt.legend(loc='upper left')
plt.tight_layout()
#plt.savefig('images/10_11.png', dpi=300)
plt.show()
# -
y_lin_pred = lr.predict(X)
y_quad_pred = pr.predict(X_quad)
print('Training MSE linear: %.3f, quadratic: %.3f' % (
mean_squared_error(y, y_lin_pred),
mean_squared_error(y, y_quad_pred)))
print('Training R^2 linear: %.3f, quadratic: %.3f' % (
r2_score(y, y_lin_pred),
r2_score(y, y_quad_pred)))
# <br>
# <br>
# ## Modeling nonlinear relationships in the Housing Dataset
# +
X = df[['LSTAT']].values
y = df['MEDV'].values
regr = LinearRegression()
# create quadratic features
quadratic = PolynomialFeatures(degree=2)
cubic = PolynomialFeatures(degree=3)
X_quad = quadratic.fit_transform(X)
X_cubic = cubic.fit_transform(X)
# fit features
X_fit = np.arange(X.min(), X.max(), 1)[:, np.newaxis]
regr = regr.fit(X, y)
y_lin_fit = regr.predict(X_fit)
linear_r2 = r2_score(y, regr.predict(X))
regr = regr.fit(X_quad, y)
y_quad_fit = regr.predict(quadratic.fit_transform(X_fit))
quadratic_r2 = r2_score(y, regr.predict(X_quad))
regr = regr.fit(X_cubic, y)
y_cubic_fit = regr.predict(cubic.fit_transform(X_fit))
cubic_r2 = r2_score(y, regr.predict(X_cubic))
# plot results
plt.scatter(X, y, label='Training points', color='lightgray')
plt.plot(X_fit, y_lin_fit,
label='Linear (d=1), $R^2=%.2f$' % linear_r2,
color='blue',
lw=2,
linestyle=':')
plt.plot(X_fit, y_quad_fit,
label='Quadratic (d=2), $R^2=%.2f$' % quadratic_r2,
color='red',
lw=2,
linestyle='-')
plt.plot(X_fit, y_cubic_fit,
label='Cubic (d=3), $R^2=%.2f$' % cubic_r2,
color='green',
lw=2,
linestyle='--')
plt.xlabel('% lower status of the population [LSTAT]')
plt.ylabel('Price in $1000s [MEDV]')
plt.legend(loc='upper right')
#plt.savefig('images/10_12.png', dpi=300)
plt.show()
# -
# Transforming the dataset:
# +
X = df[['LSTAT']].values
y = df['MEDV'].values
# transform features
X_log = np.log(X)
y_sqrt = np.sqrt(y)
# fit features
X_fit = np.arange(X_log.min()-1, X_log.max()+1, 1)[:, np.newaxis]
regr = regr.fit(X_log, y_sqrt)
y_lin_fit = regr.predict(X_fit)
linear_r2 = r2_score(y_sqrt, regr.predict(X_log))
# plot results
plt.scatter(X_log, y_sqrt, label='Training points', color='lightgray')
plt.plot(X_fit, y_lin_fit,
label='Linear (d=1), $R^2=%.2f$' % linear_r2,
color='blue',
lw=2)
plt.xlabel('log(% lower status of the population [LSTAT])')
plt.ylabel('$\sqrt{Price \; in \; \$1000s \; [MEDV]}$')
plt.legend(loc='lower left')
plt.tight_layout()
#plt.savefig('images/10_13.png', dpi=300)
plt.show()
# -
# <br>
# <br>
# # Dealing with nonlinear relationships using random forests
# ...
# ## Decision tree regression
# +
from sklearn.tree import DecisionTreeRegressor
X = df[['LSTAT']].values
y = df['MEDV'].values
tree = DecisionTreeRegressor(max_depth=3)
tree.fit(X, y)
sort_idx = X.flatten().argsort()
lin_regplot(X[sort_idx], y[sort_idx], tree)
plt.xlabel('% lower status of the population [LSTAT]')
plt.ylabel('Price in $1000s [MEDV]')
#plt.savefig('images/10_14.png', dpi=300)
plt.show()
# -
# <br>
# <br>
# ## Random forest regression
# +
X = df.iloc[:, :-1].values
y = df['MEDV'].values
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.4, random_state=1)
# +
from sklearn.ensemble import RandomForestRegressor
forest = RandomForestRegressor(n_estimators=1000,
criterion='mse',
random_state=1,
n_jobs=-1)
forest.fit(X_train, y_train)
y_train_pred = forest.predict(X_train)
y_test_pred = forest.predict(X_test)
print('MSE train: %.3f, test: %.3f' % (
mean_squared_error(y_train, y_train_pred),
mean_squared_error(y_test, y_test_pred)))
print('R^2 train: %.3f, test: %.3f' % (
r2_score(y_train, y_train_pred),
r2_score(y_test, y_test_pred)))
# +
plt.scatter(y_train_pred,
y_train_pred - y_train,
c='steelblue',
edgecolor='white',
marker='o',
s=35,
alpha=0.9,
label='Training data')
plt.scatter(y_test_pred,
y_test_pred - y_test,
c='limegreen',
edgecolor='white',
marker='s',
s=35,
alpha=0.9,
label='Test data')
plt.xlabel('Predicted values')
plt.ylabel('Residuals')
plt.legend(loc='upper left')
plt.hlines(y=0, xmin=-10, xmax=50, lw=2, color='black')
plt.xlim([-10, 50])
plt.tight_layout()
#plt.savefig('images/10_15.png', dpi=300)
plt.show()
# -
# <br>
# <br>
# # Summary
# ...
# ---
#
# Readers may ignore the next cell.
# ! python ../.convert_notebook_to_script.py --input ch10.ipynb --output ch10.py
|
ch10/ch10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ML_venv
# name: ml_venv
# ---
# # MLP Test 1
#
# Exploração e resumo sobre **_Multi-Layer Perceptron_** com base na documentação do [scikit-learn](https://scikit-learn.org/stable/modules/neural_networks_supervised.html).
#
# ## Multi-layer Perceptron
#
# Possui esse nome por consistir de multiplas camadas de **Perceptron**.
#
# 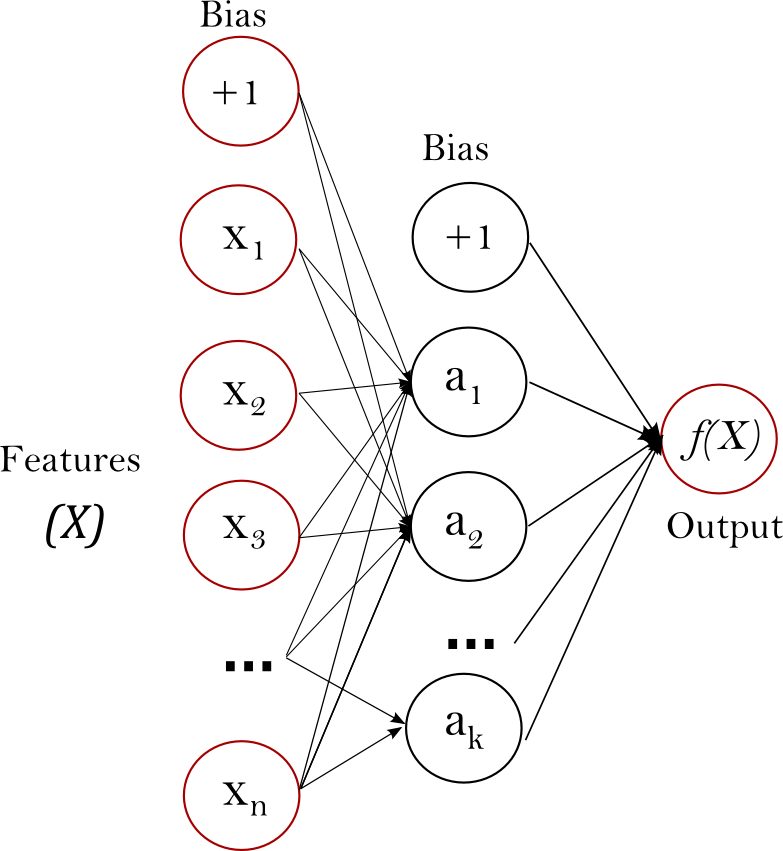
#
# Com base na imagem, destacam-se 3 partes principais:
#
# - _input layer_: contendo os dados de entrada;
#
# - _hidden layer_: contendo até multiplas camadas de Perceptrons, responsáveis pelo processamento dos dados de entrada;
#
# - _output layer_: responsáveis por receberem os dados processados e com isso gerar o dado de saída.
#
# O módulo contém atributos públicos `coefs_` e `intercepts_`:
#
# - `coefs_` é uma lista com as matrizes de peso, onde a matriz de índice _i_ representa os pesos entre as camadas _i_ e _i + 1_;
#
# - `intercepts_` é uma lista com vetores de _bias_ de cada neurônio, onde o vetor no ìndice _i_ contém os _bias_ da camada _i + 1_.
#
# ### Vantagens
#
# - Capacidade de aprender modelos não lineares.
# - Capacidade de aprender modelos em tempo real usando `partial_fit` (método).
#
# ### Desvantagens
#
# - Possui uma função de perda não-convexa com mais de um mínimo local, então, diferentes pesos aleatórios de inicialização podem levar a diferentes acurácias de validação.
# - Requer o ajuste de vários hiperparâmetros.
# - É sensível a escala das features.
#
# ## Classificação
#
# A Classe `MLPClassifier` implementa **MLP** com treino usando o algoritmo _Backpropagation_.
#
# ## Importando bibliotecas
# +
import numpy as np
import pandas as pd
from sklearn.datasets import load_iris
from sklearn.preprocessing import StandardScaler
from sklearn.neural_network import MLPClassifier
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.metrics import classification_report, confusion_matrix, plot_confusion_matrix
# +
# Carregando dataset
iris_dataset = load_iris()
X, y = load_iris(return_X_y=True)
# Separando dados
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1, test_size=0.3)
# Preprocessando dados
sc = StandardScaler().fit(X_train)
X_train_prepro = sc.transform(X_train)
X_test_prepro = sc.transform(X_test)
# -
# Dados de treino puros
pd.DataFrame(data=X_train, columns=iris_dataset.feature_names).describe()
# Dados de treino após pré-processamento
pd.DataFrame(data=X_train_prepro, columns=iris_dataset.feature_names).describe()
# +
# MLPClassifier?
# +
# Instanciando classificador
classifier = MLPClassifier(
hidden_layer_sizes=(4, 4),
random_state=1
).fit(X_train_prepro, y_train)
y1_pred = classifier.predict(X_test_prepro)
print(f'classes: {classifier.classes_}')
print(classification_report(
y_true=y_test,
y_pred=y1_pred,
target_names=iris_dataset.target_names
))
plot_confusion_matrix(
estimator=classifier,
X=X_test_prepro,
y_true=y_test,
display_labels=iris_dataset.target_names,
normalize='true'
)
# +
# Instanciando classificador
classifier = MLPClassifier(
hidden_layer_sizes=(4, 4, 4),
random_state=1
).fit(X_train_prepro, y_train)
y2_pred = classifier.predict(X_test_prepro)
print(f'classes: {classifier.classes_}')
print(classification_report(
y_true=y_test,
y_pred=y2_pred,
target_names=iris_dataset.target_names
))
plot_confusion_matrix(
estimator=classifier,
X=X_test_prepro,
y_true=y_test,
display_labels=iris_dataset.target_names,
normalize='true'
)
# +
# Instanciando classificador
classifier = MLPClassifier(
hidden_layer_sizes=(20),
random_state=1
).fit(X_train_prepro, y_train)
y3_pred = classifier.predict(X_test_prepro)
print(f'classes: {classifier.classes_}')
print(classification_report(
y_true=y_test,
y_pred=y3_pred,
target_names=iris_dataset.target_names
))
plot_confusion_matrix(
estimator=classifier,
X=X_test_prepro,
y_true=y_test,
display_labels=iris_dataset.target_names,
normalize='true'
)
# +
# Instanciando classificador
classifier = MLPClassifier(
hidden_layer_sizes=(50, 50),
random_state=1
).fit(X_train_prepro, y_train)
y4_pred = classifier.predict(X_test_prepro)
print(f'classes: {classifier.classes_}')
print(classification_report(
y_true=y_test,
y_pred=y4_pred,
target_names=iris_dataset.target_names
))
plot_confusion_matrix(
estimator=classifier,
X=X_test_prepro,
y_true=y_test,
display_labels=iris_dataset.target_names,
normalize='true'
)
# +
# Instanciando classificador
classifier = MLPClassifier(
hidden_layer_sizes=(100, 100),
random_state=1
).fit(X_train_prepro, y_train)
y5_pred = classifier.predict(X_test_prepro)
print(f'classes: {classifier.classes_}')
print(classification_report(
y_true=y_test,
y_pred=y5_pred,
target_names=iris_dataset.target_names
))
plot_confusion_matrix(
estimator=classifier,
X=X_test_prepro,
y_true=y_test,
display_labels=iris_dataset.target_names,
normalize='true'
)
|
ML/f00-sklearn_explore/03-mlp/01-test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import relevant libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# # Step 1: Read the file and display columns.
#
# +
# load the dataset
crew_df = pd.read_csv('../Dataset/ship_info.csv')
# check the dataset
print(crew_df.shape)
crew_df.head()
# -
# Above we observe that there are 158 rows and nine columns in the dataset. The columns are described as:
#
# - `Ship_name`: unique ship name
# - `Cruise_line`: The company that operates the cruise ship
# - `Age`: The number of years the ship has been in operation
# - `Tonnage`: The size or carrying capacity of the ships measured in tons.
# - `passengers`: The number of passengers in the ship
# - `length`: The length of the ships
# - `cabins`: Number of cabins the ship has
# - `passenger_density`: shows how densely occupied the ship is (higher density shows that there is high compactness of passengers in the ship)
# - `crew`: show the number of people who work or operate on the ship.
#
# We observe from the above also that some column values are continuous which are suppose to be discrete. For example, the `passengers`, `cabins` and `crew` columns are measured in continuous scale whereas, they should be in discrete values since there can't be `6.94` passengers, `3.55` cabins and `3.55` crew members on a ship. For this reason, we'll round the values in these columns to the nearest whole number.
# +
# round the passengers, cabins and crew columns to the nearest whole number
# and convert them to a discrete integer
crew_df[['passengers', 'cabins', 'crew']] = crew_df[['passengers', 'cabins', 'crew']].round().astype('int64')
# check
crew_df.head()
# -
# The dataframe looks better.
#
# # Step 2: Calculate basic statistics of the data
crew_df.describe(include='all')
crew_df.info()
# From the above outputs, we observe the following:
#
# - There are no null values across all columns in the dataset
# - Two columns are non-numeric, while four are discrete numeric columns and 3 are continuous numeric columns
# - There are 138 unique ships in the dataset. The total number of ships is 158, indicating that the dataset does not have exclusively unique ships.
# - There are 20 cruise lines, indicating that there are 20 different companies that provide cruise ship services out of which `Royal_Carribbean` company owns 20 of such ships.
# - The average age of ships in the dataset is 15 years and over 50% of ships have been in use for 14 years or more.
# - The average tonnage of ships in the dataset is 71.3 tonnes, but the standard deviation shows that the tonnage capacity of ships in the dataset varies largely and this is also evident from the minimum tonnage of 2.3 tonnes and the maximum tonnage of 220 tonnes. This invariably means that the sizes of ships in the dataset varies greatly as som ships are over 100 times the size of others.
# - Some ships in the dataset have a carrying capacity of just 1 passenger, and one crew member, while some have as much as 54 passengers and 21 crew members.
# - On the average, ships in the dataset have about 9 cabins. Although a particular ship has none at all and 50% of the ships have 10 or more cabins.
#
# # Step 3 (A): Select columns that will be probably important to predict crew size.
#
# To accomplish this task, we'll need to drop the `ship_name` column because, it contains the unique ship names, with just a few ships occuring more than once in the dataset.
#
# Since the aim is to predict `crew` size, we can be sure that the `ship_name` plays no role in determining the size of crew for ships.
#
# Another categorical column is the `Cruise_line` column which has 20 unique values within it. We'll use a boxplot to show the crew size for the various cruise lines
# +
# create a boxplot showing the variation of crew size for each cruise line
crew_df.boxplot('crew', 'Cruise_line', rot = 90, figsize = (9, 4))
plt.show()
# -
# The above boxplot shows that some cruise lines have just one crew member, while some have over 15 crew members. There is no pattern from the boxplots as regards to `cruise_line` and `crew` size, therefore, we'll also drop the `cruise_line` column.
# +
# drop the `Ship_name` and `Cruise_line` columns
crew_df.drop(['Ship_name', 'Cruise_line'], axis = 1, inplace = True)
#check
crew_df.head()
# -
# Having dropped the categorical columns beacuse they show no relationships with the `crew` size, we'll perform a correlation between all other numeric columns and the crew column and also drop columns that correlates poorly to the crew size.
crew_df.corr()
# Above, we find thatall variables correlates strongly with crew size except the `passenger_density` column which has a ver weak negative relationship crew size. We'll therefore drop this column and extract the other columns are plausible features.
# +
# drop passenger density column
crew_df.drop('passenger_density', axis = 1, inplace = True)
# check the dataset
crew_df.head()
# -
# # Step 3 (B): Create training and testing sets
#
# Next, we'll create training and testing sets from the datset we have left. Our train set will consist of 60% of the entire dataset, while the test set will hold the remaining 40 percent of the dataset.
# +
# split the dataset into features and target variable
# get features (x) as numpy array
x = crew_df.drop('crew', axis = 1).values
# get target variable (y) as numpy array
y = crew_df['crew'].values
# check
print(x.shape)
y.shape
# +
# import module for splitting dataset.
from sklearn.model_selection import train_test_split
# instantiate train_test_split, specifying its parameters
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state = 42, test_size = 0.4)
# check
print(f'Training set for feature variables is of the shape {x_train.shape} \n')
print(f'Test set for feature variables is of the shape {x_test.shape} \n')
print(f'Training set for target variable is of the shape {y_train.shape} \n')
print(f'Test set for target variable is of the shape {y_test.shape}')
# -
# # Step 4: Build a machine learning model to predict the crew size.
#
# Next, we'll build a model to predict crew size given the set of feature variables. We'll do this using the Lasso regression model, which will output a features plot afterwards.
#
# Before proceeding we'll want to get the best alpha value for the lasso regressor. For this, we'll use a grid search to search for the alpha constant that gives the best score.
# +
# import relevant modules
from sklearn.pipeline import Pipeline
from sklearn.linear_model import Lasso
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import GridSearchCV
# outline the steps to be used in our pipeline
steps = [('scaler', StandardScaler()),
('lasso', Lasso())]
# create the pipeline that will standardize and resgress on the features
pipeline = Pipeline(steps)
# specify the parameters to be used by the regressor
parameters = {'lasso__alpha': np.linspace(0.001, 0.1, 20)}
# Instantiate GridSearchCV with the pipeline and hyperparameter space and use 5 fold cv
lasso_cv = GridSearchCV(pipeline, param_grid = parameters, cv = 5)
# train the model by fitting it on the training set
lasso_cv.fit(x_train, y_train)
# print the lasso regression best parameter, r2 and best score
print("Tuned Lasso Regression Parameters: {}".format(lasso_cv.best_params_))
print('Best Score is: {}'.format(lasso_cv.best_score_))
# -
# It seems an alpha parameter of `0.1` yielded the best score of `0.89` for our model.
# Next, we'll use this parameter to get best lasso coefficients and extract features that best predicts the `crew` size.
# +
# instantiate the standard scaler
ss = StandardScaler()
# apply the scaler to the dataset
x_scaled = ss.fit_transform(x)
# split the dataset into training and test sets using the scaled data
x_train, x_test, y_train, y_test = train_test_split(x_scaled, y, random_state = 42, test_size = 0.4)
# rerun lasso with the tuned hyper parameter
lasso_tunned = Lasso(alpha = 0.1)
# fit the tunned lasso regressor to the dataset
lasso_tunned.fit(x_train, y_train)
# get the lasso coefficients
lasso_coef = lasso_tunned.fit(x_train, y_train).coef_
# predict on the test set
y_pred = lasso_tunned.predict(x_test)
# print the lasso R squared
print('Tunned lasso R squared score is: {}'.format(lasso_tunned.score(x_test, y_test)))
# get the names of our feature variables
names = crew_df.drop('crew', axis = 1).columns
# plot the lasso coefficients
plt.plot(range(len(names)), lasso_coef)
# set the x-axis names
plt.xticks(range(len(names)), names, rotation= 60)
# set the y-axis label
plt.ylabel('coefficients')
# display the plot
plt.show()
# -
# Having standardized the features in the dataset, the lasso R squared shows that about 94% percent of variations in `crew` of ships is determined by the feature variables.
#
# The coefficients plot above shows that the most prominent feature in predicting `crew` size is the number of `cabins` in the ship, followed by the `tonnage` capacity of the ship. The `length` of the ship also contributed but to rather smaller extent, whereas `Age` of the ship and number of `passengers` do not significantly contribute to determining the `crew` size of ships.
#
# ### Step 5: Calculate the Pearson correlation coefficient for the training set and testing data sets.
#
# We first convert the numpy arrays to dataframes as follows:
# +
# convert the test_features to dataframe
features_test_df = pd.DataFrame(x_test, columns = ['age', 'tonnage', 'passengers', 'length', 'cabins'])
# convert the test target to series
target_test_df = pd.Series(y_test)
# convert the train_features to dataframe
features_train_df = pd.DataFrame(x_train, columns = ['age', 'tonnage', 'passengers', 'length', 'cabins'])
# convert the train_target to series
target_train_df = pd.Series(y_train)
# concactenate the features and target varibles for the training set
training_set = pd.concat([features_train_df, target_train_df], axis = 1)
# rename the columns of the training set
training_set.rename(columns = {'age': 'Age', 'tonnage': 'Tonnage',
'passengers': 'Passengers', 'length': 'Length',
'cabins': 'Cabins', 0: 'Crew'}, inplace = True)
# check that we have a well defined training dataset
training_set.head()
# +
# concactenate the features and target varibles for the test set
test_set = pd.concat([features_test_df, target_test_df], axis = 1)
# rename the columns of the test set
test_set.rename(columns = {'age': 'Age', 'tonnage': 'Tonnage',
'passengers': 'Passengers', 'length': 'Length',
'cabins': 'Cabins', 0: 'Crew'}, inplace = True)
# check that we have a well defined test dataset
test_set.head()
# +
# Plot correlations of the training set
print('CORRELATIONS OF TRAINING SET \n \n', training_set.corr(), '\n \n')
# Plot correlations of the test set
print('CORRELATIONS OF THE TEST SET \n \n', test_set.corr())
# -
# From the above correlations we found the following:
#
# - All the features we extracted for the training set have strong and positive relationships with the `Crew` size, except for `Age` of ships, which has a weak negative relationship with crew size.
# - All the features for the test dataset also have very strong positive relationship with the `Crew` size, except for the `Age` of ships, which has a negative relationship with crew size.
#
# Conclusively, older ships generally tend to have fewer crews members than newer ships.
|
Weekly_Challenge/Predicting_crew_size.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py27]
# language: python
# name: conda-env-py27-py
# ---
# # Generative Adversarial Network
# Practise for implementing a basic GAN
# ## Theory
# - two nets fight together, generator tries to fool the discrimeter with fake examples, discrimeter net tries to figure out wheter the generator is lying
# - generator loss: $-\log (D(G(z))$, maximize the score given by the discrimeter
# - discriminator loss: $-[\log(D(x) + \log(1 - D(G(z))]$, maximize the probability from real samples $x$ and minimize probabilities from fake examples $G(z)$
# 
from IPython.display import Image
Image(url='http://www.timzhangyuxuan.com/static/images/project_DCGAN/structure.png')
from IPython.display import Image
Image(url='http://www.timzhangyuxuan.com/static/images/project_DCGAN/structure.png', embed=True)
#
# ## Geneate MINIST pictures with GAN
# ** MNIST Dataset Overview**
#
# This example is using MNIST handwritten digits. The dataset contains 60,000 examples for training and 10,000 examples for testing. The digits have been size-normalized and centered in a fixed-size image (28x28 pixels) with values from 0 to 1. For simplicity, each image has been flattened and converted to a 1-D numpy array of 784 features (28*28).
#
# 
# # Implementation
# ## Import modules
# +
from __future__ import division, print_function, absolute_import
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
# %matplotlib inline
# -
# ## Import dataset
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/", one_hot=True)
# ## Hyper parameters
# +
num_steps = 70000
batch_size = 128
learning_rate = 2e-4
image_dim = 28*28
gen_hidden_dim = 256
disc_hidden_dim = 256
input_z = 100 # input of generator
def init_weight(shape):
return tf.random_normal(shape=shape, stddev = 1. / tf.sqrt(shape[0] / 2.0) )
# +
# Weights
weights = {'gen_hidden1':tf.Variable(init_weight([input_z, gen_hidden_dim])),
'gen_out':tf.Variable(init_weight([gen_hidden_dim, image_dim])),
'disc_hidden1':tf.Variable(init_weight([image_dim, disc_hidden_dim])),
'disc_out':tf.Variable(init_weight([disc_hidden_dim, 1]))
}
bias = {'gen_hidden1':tf.Variable(tf.zeros([gen_hidden_dim])),
'gen_out':tf.Variable(tf.zeros([image_dim])),
'disc_hidden1':tf.Variable(tf.zeros([disc_hidder_dim])),
'disc_out':tf.Variable(tf.zeros([1]))
}
gen_input = tf.placeholder(tf.float32, [None, input_z], name='gen_input')
disc_input = tf.placeholder(tf.float32, [None, image_dim], name='disc_input')
gen_var_list = [weights['gen_hidden1'], bias['gen_hidden1'], weights['gen_out'], bias['gen_out']]
disc_var_list = [weights['disc_hidden1'], bias['disc_hidden1'], weights['disc_out'], bias['disc_out']]
# -
# ## Network architecture
# +
def generator(x):
hidden_layer = tf.matmul(x, weights['gen_hidden1'])
hidden_layer = tf.add(hidden_layer, bias['gen_hidden1'])
hidden_layer = tf.nn.relu(hidden_layer)
out_layer = tf.matmul(hidden_layer, weights['gen_out'])
out_layer = tf.add(out_layer, bias['gen_out'])
out_layer = tf.nn.sigmoid(out_layer)
return out_layer
def discriminator(disc_input):
hidden_layer = tf.matmul(disc_input, weights['disc_hidden1'])
hidden_layer = tf.add(hidden_layer, bias['disc_hidden1'])
hidden_layer = tf.nn.relu(hidden_layer)
out_layer = tf.matmul(hidden_layer, weights['disc_out'])
out_layer = tf.add(out_layer, bias['disc_out'])
out_layer = tf.nn.sigmoid(out_layer)
return out_layer
# -
# ## Construct model
gen_out = generator(gen_input)
disc_real = discriminator(disc_input)
disc_fake = discriminator(gen_out)
# ## Loss
# +
gen_loss = -tf.reduce_mean(tf.log(disc_fake))
disc_loss = -tf.reduce_mean(tf.log(disc_real) + tf.log(1. - disc_fake))
# Optimizer
gen_opt = tf.train.AdadeltaOptimizer(
learning_rate).minimize(gen_loss, var_list=gen_var_list)
disc_opt = tf.train.AdamOptimizer(learning_rate).minimize(
disc_loss, var_list=disc_var_list)
# -
# ## Training
# + run_control={"marked": true}
init = tf.global_variables_initializer()
loss_array = []
sess = tf.Session()
sess.run(init)
for step in range(1, num_steps+1):
batch_x, _ = mnist.train.next_batch(batch_size)
z = np.random.uniform(-1., 1., size=[batch_size, input_z])
_, _, g_loss, d_loss = sess.run([gen_opt, disc_opt, gen_loss, disc_loss], feed_dict={
gen_input: z, disc_input: batch_x})
if step % 10 == 0 or step== 1:
loss_array.append([step, g_loss, d_loss])
if step % 2000 == 0 or step == 1:
print ("step:{}, generator loss:{}, discriminator loss:{}".format(step, g_loss, d_loss))
# +
# plot
loss = np.array(loss_array)
plt.plot(loss[:,0], loss[:,1],'r', loss[:,0], loss[:,2], 'b')
plt.xlabel('Steps')
plt.ylabel('Loss')
plt.title('Generator loss & discriminator loss')
plt.legend(('generator loss', 'discriminator loss'))
plt.show()
# -
# ## Test
# +
# Testing
# Generate images from noise, using the generator network.
n = 6
canvas = np.empty((28 * n, 28 * n))
for i in range(n):
# Noise input.
z = np.random.uniform(-1., 1., size=[n, input_z])
# Generate image from noise.
g = sess.run(gen_out, feed_dict={gen_input: z})
# Reverse colours for better display
g = -1 * (g - 1)
for j in range(n):
# Draw the generated digits
canvas[i * 28:(i + 1) * 28, j * 28:(j + 1) * 28] = g[j].reshape([28, 28])
plt.figure(figsize=(n, n))
plt.imshow(canvas, origin="upper", cmap="gray")
plt.show()
|
Notebook/GAN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 一、锯齿波的傅里叶展开
# +
# inline 模式下,matplotlib 绘制的图会直接输出到 notebook 内
# 但是 inline 模式有很多缺陷(不支持 animation,不能交互)
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
PI2 = 2*np.pi
PI3 = 3*np.pi
PI4 = 4*np.pi
# -
# ## 1.1 锯齿波函数的定义与图形
#
# $$
# \begin{equation}
# f(x)=
# \begin{cases}
# \frac{1}{2}(\pi - x) & 0 < x \leq 2 \pi \\
# f(x + 2 \pi) & \text{otherwise}
# \end{cases}
# \end{equation}
# $$
# +
# 锯齿波图形
def f(x): # 函数定义
if x <= 0:
return f(x + PI2)
elif x > 2*np.pi:
return f(x - PI2)
else:
return 1/2 * (np.pi - x)
X = np.arange(-PI3, PI3, 0.02) # x 范围
Y = list(map(f, X)) # 计算对应的 y 值
fig, axes = plt.subplots(dpi=100)
axes.grid()
axes.plot(X, Y) # 绘图
# -
# ## 1.2 傅里叶展开
#
# 这是一个奇函数,展开为正弦级数为 $\sum_{n=1}^{\infty} \frac{\sin nx}{n}$,
# 下面开始绘制级数叠加过程图
# +
# 这里使用矩阵方法,同时计算 n 项的值。因为这种方式要比单独计算每一项要快。
n_max = 300 # 展开到第 n 项
N = np.arange(1,n_max)
X = np.arange(-PI3, PI3, 0.001) # x 范围
NX = np.outer(N, X) # 计算 N 和 X 的外积,所得矩阵的每一行,即为对应 n 值的 nx 向量。
NY = np.array(np.sin(NX) / np.matrix(N).transpose()) # 计算对应的 Y(矩阵),n 应被对应的 sin(nx) 所除,所以 N 需要转置
Serise = np.cumsum(NY, axis=0) # 计算级数,不是原地计算,因此不会影响到 NY
# -
# ## 1.3 动画
# +
# 1. 使用 matplotlib 的 animation。略麻烦,但是交互性高。
# %matplotlib notebook
from matplotlib.lines import Line2D
import matplotlib.animation as animation
fig, (ax1, ax2) = plt.subplots(1, 2, dpi=100)
ax1.grid()
ax2.grid()
def animate(Y):
ax1.plot(X,Y)
return ax2.plot(X,Y, "black")
ani = animation.FuncAnimation(
fig, animate, frames=Serise, interval=80, blit=True)
fig.show()
# +
# 2. 使用 jupyterlab 的 ipywidgets 插件进行交互
import ipywidgets as widgets
# %matplotlib inline
def update_lines(i):
# inline 模式下,fig好像是一经显示就不能修改的,因此每次都要创建新的
fig, axes = plt.subplots(dpi=150)
axes.grid()
axes.plot(X, Serise[i], 'red')
widgets.interact(update_lines, i=(1,100,1))
# -
|
jupyter-notebook/digital singal processing/Fourier expansion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Principal Component Analysis
#
# by <NAME> and <NAME>
#
# Part of the Quantopian Lecture Series:
#
# * [www.quantopian.com/lectures](https://www.quantopian.com/lectures)
# * [https://github.com/quantopian/research_public](https://github.com/quantopian/research_public)
#
# ---
#
# Applications in many fields, such as image processing, bioinformatics, and quantitative finance, involve large-scale data. Both the size and complexity of this data can make the computations required for analysis practically infeasible. Principal Component Analysis (PCA) is a classical method for dimension reduction. It uses the first several **principal components**, statistical features that explain most of the variation of a $m \times n$ data matrix $\mathbf{X}$, to describe the large-scale data matrix $\mathbf{X}$ economically.
from numpy import linalg as LA
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# We will introduce PCA with an image processing example. A grayscale digital image can be represented by a matrix, whose $(i,j)^{th}$ entry corresponds to the measurement of gray
# scale at the $(i,j)^{th}$ pixel. The following gray-scale image has $200 \times 200$ pixels, though it can be changed on the fly. We store it in a matrix $\mathbf{X}$. The number of rows of the $\mathbf{X}$ is $200$, and the number of columns of $\mathbf{X}$ is $200$.
def generate_test_image(m,n):
X = np.zeros((m,n))
# generate a rectangle
X[25:80,25:80] = 1
# generate a triangle
for i in range(25, 80, 1):
X[i+80:160, 100+i-1] = 2
# generate a circle
for i in range(0,200,1):
for j in range(0,200,1):
if ((i - 135)*(i - 135) +(j - 53)*(j - 53) <= 900):
X[i, j] = 3
return X
X = generate_test_image(200,200)
# We start with a simple checkboard pattern, add some random normal noise, and add a gradient.
imgplot = plt.imshow(X, cmap='gray')
plt.title('Original Test Image');
m = X.shape[0] # num of rows
n = X.shape[1] # num of columns
# Set each row as a variable, with observations in the columns. Denote the covariance matrix of $\mathbf{X}$ as $\mathbf{C}$, where the size of $\mathbf{C}$ is $m \times m$. $\mathbf{C}$ is a matrix whose $(i,j)^{th}$ entry is the covariance between the $i^{th}$ row and $j^{th}$ row of the matrix $\mathbf{X}$.
X = np.asarray(X, dtype=np.float64)
C = np.cov(X)
np.linalg.matrix_rank(C)
# Performing principal component analysis decomposes the matrix $\mathbf{C}$ into:
#
# $$\mathbf{C} = \mathbf{L}\mathbf{P}\mathbf{L}^{\top},$$
#
# where $\mathbf{P}$ is a diagonal matrix $\mathbf{P}=\text{diag}(\lambda_1,\lambda_2,\dots,\lambda_m)$, with $\lambda_1 \geq \lambda_1 \geq \dots \lambda_m \geq 0$ being the eigenvalues of matrix $\mathbf{C}$. The matrix $\mathbf{L}$ is an orthogonal matrix, consisting the eigenvectors of matrix $\mathbf{C}$.
P, L = LA.eigh(C)
# The function `LA.eigh` lists the eigenvalues from small to large in $P$. Let us change the order first to list them from largest to smallest and make sure that $\mathbf{L}\mathbf{P}\mathbf{L}^{\top}==\mathbf{C}$.
P = P[::-1]
L = L[:,::-1]
np.allclose(L.dot(np.diag(P)).dot(L.T), C)
# Here we plot all of the eigenvalues:
plt.semilogy(P, '-o')
plt.xlim([1, P.shape[0]])
plt.xlabel('eigenvalue index')
plt.ylabel('eigenvalue in a log scale')
plt.title('Eigenvalues of Covariance Matrix');
# The $i^{th}$ **principal component** is given as $i^{th}$ row of $\mathbf{V}$,
#
# $$\mathbf{V} =\mathbf{L}^{\top} \mathbf{X}.$$
#
V = L.T.dot(X)
V.shape
# If we multiply both sides on the left by $\mathbf{L}$, we get the following:
#
# $$\mathbf{L}\mathbf{L}^{\top} \mathbf{X}= \mathbf{L}\mathbf{V}.$$
#
# The matrix $\mathbf{L}$ is the set of eigenvectors from a covariance matrix , so $\mathbf{L}\mathbf{L}^{\top} = \mathbf{I}$ and $\mathbf{L}\mathbf{L}^{\top}\mathbf{X} = \mathbf{X}$. The relationship among matrices of $\mathbf{X}$, $\mathbf{L}$, and $\mathbf{V}$ can be expressed as
#
# $$\mathbf{X} = \mathbf{L}\mathbf{V}.$$
#
#
# To approximate $\mathbf{X}$, we use $k$ eigenvectors that have largest eigenvalues:
#
# $$\mathbf{X} \approx \mathbf{L[:, 1:k]}\mathbf{L[:, 1:k]}^{\top} \mathbf{X}.$$
#
# Denote the approximated $\mathbf{X}$ as $\tilde{\mathbf{X}} = \mathbf{L[:, 1:k]}\mathbf{L[:, 1:k]}^{\top} \mathbf{X}$. When $k = m $, the $\tilde{\mathbf{X}}$ should be same as $\mathbf{X}$.
k = 200
X_tilde = L[:,0:k-1].dot(L[:,0:k-1].T).dot(X)
np.allclose(X_tilde, X)
plt.imshow(X_tilde, cmap='gray')
plt.title('Approximated Image with full rank');
# The proportion of total variance due to the $i^{th}$ principal component is given by the ratio $\frac{\lambda_i}{\lambda_1 + \lambda_2 + \dots \lambda_m}.$ The sum of proportion of total variance should be $1$. As we defined, $\lambda_i$ is $i^{th}$ entry of $\mathbf{P}$,
#
# $$\sum_{i}\frac{P_i}{\text{trace}(P)} = 1$$
#
# Where the trace$(P)$ is the sum of the diagonal of $P$.
(P/P.sum()).sum()
plt.plot((P/P.sum()).cumsum(), '-o')
plt.title('Cumulative Sum of the Proportion of Total Variance')
plt.xlabel('index')
plt.ylabel('Proportion');
# Recall the number of principal components is denoted as $k$. Let $k$ be $10, 20, 30, 60$ as examples and take a look at the corresponding approximated images.
X_tilde_10 = L[:,0:10-1].dot(V[0:10-1,:])
X_tilde_20 = L[:,0:20-1].dot(V[0:20-1,:])
X_tilde_30 = L[:,0:30-1].dot(V[0:30-1,:])
X_tilde_60 = L[:,0:60-1].dot(V[0:60-1,:])
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2, 2, figsize=(12, 12))
ax1.imshow(X_tilde_10, cmap='gray')
ax1.set(title='Approximated Image with k = 10')
ax2.imshow(X_tilde_20, cmap='gray')
ax2.set(title='Approximated Image with k = 20')
ax3.imshow(X_tilde_30, cmap='gray')
ax3.set(title='Approximated Image with k = 30')
ax4.imshow(X_tilde_60, cmap='gray')
ax4.set(title='Approximated Image with k = 60');
# The number of variables in $X$ is $200$. When reducing the dimension to $k=60$, which uses half of the principal components, the approximated image is close to the original one.
#
# Moving forward, we do not have to do PCA by hand. Luckly, [scikit-learn](http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html) has an implementation that we can use. Next, let us show an example in quantitative finance using sklearn.
# ## PCA on a Portfolio
# Construct a portfolio with 10 stocks, IBM, MSFT, FB, T, INTC, ABX, NEM, AU, AEM, GFI. 5 of them are technology related and 5 of them are gold mining companies.
#
# In this case, there are 10 variables (companies), and each column is a variable.
# +
symbol = ['IBM','MSFT', 'FB', 'T', 'INTC', 'ABX','NEM', 'AU', 'AEM', 'GFI']
start = "2015-09-01"
end = "2016-11-01"
portfolio_returns = get_pricing(symbol, start_date=start, end_date=end, fields="price").pct_change()[1:]
# +
from sklearn.decomposition import PCA
num_pc = 2
X = np.asarray(portfolio_returns)
[n,m] = X.shape
print 'The number of timestamps is {}.'.format(n)
print 'The number of stocks is {}.'.format(m)
pca = PCA(n_components=num_pc) # number of principal components
pca.fit(X)
percentage = pca.explained_variance_ratio_
percentage_cum = np.cumsum(percentage)
print '{0:.2f}% of the variance is explained by the first 2 PCs'.format(percentage_cum[-1]*100)
pca_components = pca.components_
# -
# Notice that the grand bulk of the variance of the returns of these assets can be explained by the first two principal components.
#
# Now we collect the first two principal components and plot their contributions.
# +
x = np.arange(1,len(percentage)+1,1)
plt.subplot(1, 2, 1)
plt.bar(x, percentage*100, align = "center")
plt.title('Contribution of principal components',fontsize = 16)
plt.xlabel('principal components',fontsize = 16)
plt.ylabel('percentage',fontsize = 16)
plt.xticks(x,fontsize = 16)
plt.yticks(fontsize = 16)
plt.xlim([0, num_pc+1])
plt.subplot(1, 2, 2)
plt.plot(x, percentage_cum*100,'ro-')
plt.xlabel('principal components',fontsize = 16)
plt.ylabel('percentage',fontsize = 16)
plt.title('Cumulative contribution of principal components',fontsize = 16)
plt.xticks(x,fontsize = 16)
plt.yticks(fontsize = 16)
plt.xlim([1, num_pc])
plt.ylim([50,100]);
# -
# From these principal components we can construct "statistical risk factors", similar to more conventional common risk factors. These should give us an idea of how much of the portfolio's returns comes from some unobservable statistical feature.
factor_returns = X.dot(pca_components.T)
factor_returns = pd.DataFrame(columns=["factor 1", "factor 2"],
index=portfolio_returns.index,
data=factor_returns)
factor_returns.head()
# The factor returns here are an analogue to the principal component matrix $\mathbf{V}$ in the image processing example.
factor_exposures = pd.DataFrame(index=["factor 1", "factor 2"],
columns=portfolio_returns.columns,
data = pca.components_).T
factor_exposures
# The factor exposures are an analogue to the eigenvector matrix $\mathbf{L}$ in the image processing example.
labels = factor_exposures.index
data = factor_exposures.values
# +
plt.subplots_adjust(bottom = 0.1)
plt.scatter(
data[:, 0], data[:, 1], marker='o', s=300, c='m',
cmap=plt.get_cmap('Spectral'))
plt.title('Scatter Plot of Coefficients of PC1 and PC2')
plt.xlabel('factor exposure of PC1')
plt.ylabel('factor exposure of PC2')
for label, x, y in zip(labels, data[:, 0], data[:, 1]):
plt.annotate(
label,
xy=(x, y), xytext=(-20, 20),
textcoords='offset points', ha='right', va='bottom',
bbox=dict(boxstyle='round,pad=0.5', fc='yellow', alpha=0.5),
arrowprops=dict(arrowstyle = '->', connectionstyle='arc3,rad=0')
);
# -
# Creating statistical risk factors allows us to further break down the returns of a portfolio to get a better idea of the risk. This can be used as an additional step after performance attribution with more common risk factors, such as those in the [Quantopian Risk Model](https://www.quantopian.com/risk-model), to try to account for additional unknown risks.
#
#
# ## References:
# - <NAME>., 2010. *Numerical linear algebra and applications*. Siam.
# - <NAME>., <NAME>. and <NAME>., 2007. *Quantitative equity portfolio management: modern techniques and applications*. CRC Press.
# *This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, Quantopian, Inc. has not taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information, believed to be reliable, available to Quantopian, Inc. at the time of publication. Quantopian makes no guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
|
quantopian/lectures/PCA/notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tensorflow-gpu
# language: python
# name: tensorflow-gpu
# ---
# ## Problem 4 - Batch Augmentation, Cutout Regularization 20 points
import tensorflow as tf
# import keras
from tensorflow.keras.layers import Dense, Conv2D, BatchNormalization, Activation
from tensorflow.keras.layers import AveragePooling2D, Input, Flatten
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.callbacks import ModelCheckpoint, LearningRateScheduler
from tensorflow.keras.callbacks import ReduceLROnPlateau
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.regularizers import l2
from tensorflow.keras import backend as K
from tensorflow.keras.models import Model
from tensorflow.keras.datasets import cifar10
import numpy as np
from albumentations import Cutout
import matplotlib.pyplot as plt
import cv2
# ### 1.Explain cutout regularization and its advantages compared to simple dropout (as argued in the paper by DeVries et al) in your own words. Select any 2 images from CIFAR10 and show how does these images look after applying cutout. Use a square-shaped fixed size zero-mask to a random location of each image and generate its cutout version. Refer to the paper by DeVries et al (Section 3) and associated github repository. (2+4)
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
cutout = Cutout(num_holes=1, max_h_size=7, max_w_size=7, fill_value=0, p=1)
plt.imshow(cutout(image=x_train[0])['image'])
plt.imshow(cutout(image=x_train[1])['image'])
# ### 2.Using CIFAR10 datasest and Resnet-44 we will first apply simple data augmentation as in He et al. (look at Section 4.2 of He et al.) and train the model with batch size 64. Note that testing is always done with original images. Plot validation error vs number of training epochs. (4)
# Training parameters for ResNet-44
batch_size = 64
epochs = 100
# data_augmentation = True
num_classes = 10
n = 3
depth = n * 6 + 2
# +
# Input image dimensions.
input_shape = x_train.shape[1:]
# Normalize data.
x_train = x_train.astype('float32') / 255
x_test = x_test.astype('float32') / 255
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
print('y_train shape:', y_train.shape)
# Convert class vectors to binary class matrices.
y_train = tf.keras.utils.to_categorical(y_train, num_classes)
y_test = tf.keras.utils.to_categorical(y_test, num_classes)
# -
def data_augmentation(img, pad_width, flip=True):
'''
padding some pixels on each side, and randomly sample a crop from the padded image or its horizontal flip.
'''
if flip == True:
img = np.rot90(img ,int(np.random.rand()*4))
pad_img = np.pad(img,((pad_width,pad_width),(pad_width,pad_width),(0,0)))
rand_x = int(np.random.rand()*pad_width*2)
rand_y = int(np.random.rand()*pad_width*2)
return pad_img[rand_x:rand_x+img.shape[0], rand_y:rand_y+img.shape[1],...]
x_train_aug = np.zeros((x_train.shape[0]*4, x_train.shape[1], x_train.shape[2], x_train.shape[3]))
y_train_aug = np.zeros((y_train.shape[0]*4, y_train.shape[1]))
for i in range(x_train_aug.shape[0]):
x_train_aug[i] = data_augmentation(x_train[i%x_train.shape[0]], 4)
y_train_aug[i] = y_train[i%x_train.shape[0]]
def lr_schedule(epoch):
"""Learning Rate Schedule
Learning rate is scheduled to be reduced after 80, 120, 160, 180 epochs.
Called automatically every epoch as part of callbacks during training.
# Arguments
epoch (int): The number of epochs
# Returns
lr (float32): learning rate
"""
lr = 1e-3
if epoch > 180:
lr *= 0.5e-3
elif epoch > 160:
lr *= 1e-3
elif epoch > 120:
lr *= 1e-2
elif epoch > 80:
lr *= 1e-1
print('Learning rate: ', lr)
return lr
# +
def resnet_layer(inputs,
num_filters=16,
kernel_size=3,
strides=1,
activation='relu',
batch_normalization=True,
conv_first=True):
"""2D Convolution-Batch Normalization-Activation stack builder
# Arguments
inputs (tensor): input tensor from input image or previous layer
num_filters (int): Conv2D number of filters
kernel_size (int): Conv2D square kernel dimensions
strides (int): Conv2D square stride dimensions
activation (string): activation name
batch_normalization (bool): whether to include batch normalization
conv_first (bool): conv-bn-activation (True) or
bn-activation-conv (False)
# Returns
x (tensor): tensor as input to the next layer
"""
conv = Conv2D(num_filters,
kernel_size=kernel_size,
strides=strides,
padding='same',
kernel_initializer='he_normal',
kernel_regularizer=l2(1e-4))
x = inputs
if conv_first:
x = conv(x)
if batch_normalization:
x = BatchNormalization()(x)
if activation is not None:
x = Activation(activation)(x)
else:
if batch_normalization:
x = BatchNormalization()(x)
if activation is not None:
x = Activation(activation)(x)
x = conv(x)
return x
def resnet_v1(input_shape, depth, num_classes=10):
"""ResNet Version 1 Model builder [a]
Stacks of 2 x (3 x 3) Conv2D-BN-ReLU
Last ReLU is after the shortcut connection.
At the beginning of each stage, the feature map size is halved (downsampled)
by a convolutional layer with strides=2, while the number of filters is
doubled. Within each stage, the layers have the same number filters and the
same number of filters.
Features maps sizes:
stage 0: 32x32, 16
stage 1: 16x16, 32
stage 2: 8x8, 64
The Number of parameters is approx the same as Table 6 of [a]:
ResNet20 0.27M
ResNet32 0.46M
ResNet44 0.66M
ResNet56 0.85M
ResNet110 1.7M
# Arguments
input_shape (tensor): shape of input image tensor
depth (int): number of core convolutional layers
num_classes (int): number of classes (CIFAR10 has 10)
# Returns
model (Model): Keras model instance
"""
if (depth - 2) % 6 != 0:
raise ValueError('depth should be 6n+2 (eg 20, 32, 44 in [a])')
# Start model definition.
num_filters = 16
num_res_blocks = int((depth - 2) / 6)
inputs = Input(shape=input_shape)
x = resnet_layer(inputs=inputs)
# Instantiate the stack of residual units
for stack in range(3):
for res_block in range(num_res_blocks):
strides = 1
if stack > 0 and res_block == 0: # first layer but not first stack
strides = 2 # downsample
y = resnet_layer(inputs=x,
num_filters=num_filters,
strides=strides)
y = resnet_layer(inputs=y,
num_filters=num_filters,
activation=None)
if stack > 0 and res_block == 0: # first layer but not first stack
# linear projection residual shortcut connection to match
# changed dims
x = resnet_layer(inputs=x,
num_filters=num_filters,
kernel_size=1,
strides=strides,
activation=None,
batch_normalization=False)
x = tf.keras.layers.add([x, y])
x = Activation('relu')(x)
num_filters *= 2
# Add classifier on top.
# v1 does not use BN after last shortcut connection-ReLU
x = AveragePooling2D(pool_size=8)(x)
y = Flatten()(x)
outputs = Dense(num_classes,
activation='softmax',
kernel_initializer='he_normal')(y)
# Instantiate model.
model = Model(inputs=inputs, outputs=outputs)
return model
# -
model = resnet_v1(input_shape=input_shape, depth=depth)
model.compile(loss='categorical_crossentropy',
optimizer=Adam(lr=lr_schedule(0)),
metrics=['accuracy'])
# +
lr_scheduler = LearningRateScheduler(lr_schedule)
lr_reducer = ReduceLROnPlateau(factor=np.sqrt(0.1),
cooldown=0,
patience=5,
min_lr=0.5e-6)
callbacks = [lr_reducer, lr_scheduler]
# +
import albumentations
from albumentations import (Blur,Flip,ShiftScaleRotate,GridDistortion,ElasticTransform,PadIfNeeded,
HueSaturationValue,Transpose,RandomBrightnessContrast,CLAHE,RandomCrop,
CoarseDropout,Normalize,ToFloat,OneOf,Compose,Cutout)
from tensorflow import keras
import tensorflow.keras.backend as K
class HeGenerator(keras.utils.Sequence):
def __init__(self, image_filenames, labels,
batch_size=64, mix=False, augment=True):
self.image_filenames = image_filenames
self.labels = labels
self.batch_size = batch_size
self.is_mix = mix
self.is_augment = augment
if self.is_augment:
self.generator = Compose([Flip(p=0.5),
PadIfNeeded(40, 40, border_mode=0),
RandomCrop(32, 32, always_apply=False, p=1.0)])
else:
self.generator = Compose([ToFloat(max_value=255.0,p=1.0)],p=1.0)
def __len__(self):
return int(np.ceil(len(self.image_filenames)/self.batch_size))
def mix_up(self,x,y):
original_index = np.arange(x.shape[0])
new_index = np.arange(x.shape[0])
np.random.shuffle(new_index)
beta = np.random.beta(0.2, 0.4)
mix_x = beta * x[original_index] + (1 - beta) * x[new_index]
mix_y = beta * y[original_index] + (1 - beta) * y[new_index]
return mix_x, mix_y
def __getitem__(self,index):
batch_x = self.image_filenames[index*self.batch_size:(index+1)*self.batch_size]
batch_y = self.labels[index*self.batch_size:(index+1)*self.batch_size]
new_images = []
new_labels = []
for image_name,label in zip(batch_x,batch_y):
image = image_name
img = self.generator(image=image)['image']
new_images.append(img)
new_labels.append(label)
new_images = np.array(new_images)
new_labels = np.array(new_labels)
if self.is_mix:
new_images, new_labels = self.mix_up(new_images, new_labels)
return new_images, new_labels
# -
train_generator = HeGenerator(x_train, y_train, batch_size=batch_size)
train_generator.__getitem__(0)[0].shape
plt.imshow(train_generator.__getitem__(0)[0][0])
# Fit the model on the batches generated by datagen.flow().
history = model.fit_generator(train_generator,
validation_data=(x_test, y_test),
epochs=epochs, verbose=1,
callbacks=callbacks)
valid_error = [1 - i for i in history.history['val_accuracy']]
epoch = [i for i in range(0, epochs)]
# Score trained model.
scores = model.evaluate(x_test, y_test, verbose=1)
plt.title('validation error VS epoch')
plt.xlabel("epoch")
plt.ylabel("validation error")
plt.plot(epoch, valid_error)
# ### 3.Next use cutout for data augmentation in Resnet-44 as in Hoffer et al. and train the model and use the same set-up in your experiments. Plot validation error vs number of epochs for different values of M (2,4,8,16,32) where M is the number of instances generated from an input sample after applying cutout M times effectively increasing the batch size to M * B, where B is the original batch size (before applying cutout augmentation). You will obtain a figure similar to Figure 3(a) in the paper by Hoffer et al. Also compare the number of epochs and wallclock time to reach 94% accuracy for different values of M. Do not run any experiment for more than 100 epochs. If even after 100 epochs of training you did not achieve 94% then just report the accuracy you obtain and the corresponding wallclock time to train for 100 epochs. Before attempting this question it is advisable to read paper by Hoffer et al. and especially Section 4.1. (5+5)
# +
import albumentations
from albumentations import (Blur,Flip,ShiftScaleRotate,GridDistortion,ElasticTransform,PadIfNeeded,
HueSaturationValue,Transpose,RandomBrightnessContrast,CLAHE,RandomCrop,
CoarseDropout,Normalize,ToFloat,OneOf,Compose,Cutout)
from tensorflow import keras
import tensorflow.keras.backend as K
class MyGenerator(keras.utils.Sequence):
def __init__(self, image_filenames, labels,
batch_size=64, M=16, mix=False, augment=True):
self.image_filenames = image_filenames
self.labels = labels
self.batch_size = batch_size
self.is_mix = mix
self.is_augment = augment
self.M = M
if self.is_augment:
# self.generator = Compose([Blur(),Flip(),Transpose(),ShiftScaleRotate(),
# RandomBrightnessContrast(),HueSaturationValue(),
# CLAHE(),GridDistortion(),ElasticTransform(),CoarseDropout(),
# ToFloat(max_value=255.0,p=1.0)],p=1.0)
self.generator = Compose([Flip(p=0.5),
PadIfNeeded(40, 40, border_mode=0),
RandomCrop(32, 32, always_apply=False, p=1.0),
Cutout(num_holes=1, max_h_size=16, max_w_size=16, fill_value=0, p=0.5)])
else:
self.generator = Compose([ToFloat(max_value=255.0,p=1.0)],p=1.0)
def __len__(self):
return int(np.ceil(len(self.image_filenames)/self.batch_size))
def mix_up(self,x,y):
original_index = np.arange(x.shape[0])
new_index = np.arange(x.shape[0])
np.random.shuffle(new_index)
beta = np.random.beta(0.2, 0.4)
mix_x = beta * x[original_index] + (1 - beta) * x[new_index]
mix_y = beta * y[original_index] + (1 - beta) * y[new_index]
return mix_x, mix_y
def __getitem__(self,index):
batch_x = self.image_filenames[index*self.batch_size:(index+1)*self.batch_size]
batch_y = self.labels[index*self.batch_size:(index+1)*self.batch_size]
new_images = []
new_labels = []
for image_name,label in zip(batch_x,batch_y):
image = image_name
for j in range(0, self.M): # Add M new generate batches
img = self.generator(image=image)['image']
new_images.append(img)
new_labels.append(label)
new_images = np.array(new_images)
new_labels = np.array(new_labels)
if self.is_mix:
new_images, new_labels = self.mix_up(new_images, new_labels)
return new_images, new_labels
# -
train_generator = MyGenerator(x_train, y_train, batch_size=batch_size, M=1)
train_generator.__getitem__(0)[0].shape
plt.imshow(train_generator.__getitem__(0)[0][0])
def lr_schedule(epoch):
"""Learning Rate Schedule
Learning rate is scheduled to be reduced after 80, 120, 160, 180 epochs.
Called automatically every epoch as part of callbacks during training.
# Arguments
epoch (int): The number of epochs
# Returns
lr (float32): learning rate
"""
lr = 1e-3
if epoch > 180:
lr *= 0.5e-3
elif epoch > 160:
lr *= 1e-3
elif epoch > 120:
lr *= 1e-2
elif epoch > 80:
lr *= 1e-1
print('Learning rate: ', lr)
return lr
import time
class LossHistory(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.totaltime = time.time()
def on_train_end(self, logs={}):
self.totaltime = time.time() - self.totaltime
lr_scheduler = LearningRateScheduler(lr_schedule)
lr_reducer = ReduceLROnPlateau(factor=np.sqrt(0.1),
cooldown=0,
patience=5,
min_lr=0.5e-6)
time_history = LossHistory()
callbacks = [lr_reducer, lr_scheduler, time_history]
train_generator = MyGenerator(x_train, y_train, batch_size=batch_size, M=1) # Set M = 1
model = resnet_v1(input_shape=input_shape, depth=depth)
model.compile(loss='categorical_crossentropy',
optimizer=Adam(lr=lr_schedule(0)),
metrics=['accuracy'])
history = model.fit_generator(train_generator,
validation_data=(x_test, y_test),
epochs=epochs, verbose=1,
callbacks=callbacks)
totaltime_M1 = time_history.totaltime
val_error_M1 = [1 - i for i in history.history['val_accuracy']]
print(totaltime_M1)
lr_scheduler = LearningRateScheduler(lr_schedule)
lr_reducer = ReduceLROnPlateau(factor=np.sqrt(0.1),
cooldown=0,
patience=5,
min_lr=0.5e-6)
time_history = LossHistory()
callbacks = [lr_reducer, lr_scheduler, time_history]
train_generator = MyGenerator(x_train, y_train, batch_size=batch_size, M=2) # Set M = 2
model = resnet_v1(input_shape=input_shape, depth=depth)
model.compile(loss='categorical_crossentropy',
optimizer=Adam(lr=lr_schedule(0)),
metrics=['accuracy'])
history = model.fit_generator(train_generator,
validation_data=(x_test, y_test),
epochs=epochs, verbose=1,
callbacks=callbacks)
totaltime_M2 = time_history.totaltime
val_error_M2 = [1 - i for i in history.history['val_accuracy']]
print(totaltime_M2)
plt.title('validation error VS epoch')
plt.xlabel("epoch")
plt.ylabel("validation error")
plt.plot(epoch, valid_error)
plt.plot(epoch, val_error_M1)
plt.plot(epoch, val_error_M2)
plt.legend(["ResNet-44 baseline","M = 1","M = 2"],loc='upper right')
|
Assignment3/question4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D4-MachineLearning/W1D4_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="jsETFMEqs3j3"
# # Neuromatch Academy: Week 1, Day 4, Tutorial 1
# # Machine Learning: GLMs
#
# + [markdown] colab_type="text" id="bm1XgN5i8hna"
# In this tutorial you will learn about Generalized Linear Models (GLMs), which are a fundamental framework for supervised learning.
#
# The objective is to model retinal ganglion cell spike trains. First with a Linear-Gaussian GLM (also known as ordinary least-squares regression model) and then with a Poisson GLM (aka "Linear-Nonlinear-Poisson" model).
#
# This tutorial is designed to run with retinal ganglion cell spike train data from [Uzzell & Chichilnisky 2004](https://journals.physiology.org/doi/full/10.1152/jn.01171.2003?url_ver=Z39.88-2003&rfr_id=ori:rid:crossref.org&rfr_dat=cr_pub%20%200pubmed).
#
# *Acknowledgements:*
#
# - We thank <NAME> for providing the dataset. Please note that it is provided for tutorial purposes only, and should not be distributed or used for publication without express permission from the author (<EMAIL>).
# - We thank <NAME>, much of this tutorial is inspired by exercises asigned in his 'Statistical Modeling and Analysis of Neural Data' class.
# + [markdown] colab_type="text" id="3oe5ZZYgL5gv"
# # Setup
# Run these cells to get the tutorial started
# + cellView="both" colab={} colab_type="code" id="ZgdrFHYF8hnH"
import numpy as np
import matplotlib.pyplot as plt
from scipy.io import loadmat
from scipy.optimize import minimize
# + cellView="form" colab={} colab_type="code" id="fV2zkxVJuc-L"
#@title Plot setup
fig_w, fig_h = 8, 6
plt.rcParams.update({'figure.figsize': (fig_w, fig_h)})
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# + cellView="form" colab={} colab_type="code" id="Rj8dcuCENOZs"
#@title Helper functions
def plot_stim_and_spikes(stim, spikes, dt, nt=120):
"""Show time series of stim intensity and spike counts.
Args:
stim (1D array): vector of stimulus intensities
spikes (1D array): vector of spike counts
dt (number): duration of each time step
nt (number): number of time steps to plot
"""
timepoints = np.arange(120)
time = timepoints * dt_stim
f, (ax_stim, ax_spikes) = plt.subplots(
nrows=2, sharex=True, figsize=(8, 5),
)
ax_stim.plot(time, stim[timepoints])
ax_stim.set_ylabel('Stimulus intensity')
ax_spikes.plot(time, spikes[timepoints])
ax_spikes.set_xlabel('Time (s)')
ax_spikes.set_ylabel('Number of spikes')
f.tight_layout()
def plot_glm_matrices(X, y, nt=50):
"""Show X and Y as heatmaps.
Args:
X (2D array): Design matrix.
y (1D or 2D array): Target vector.
"""
Y = np.c_[y] # Ensure Y is 2D and skinny
f, (ax_y, ax_x) = plt.subplots(
ncols=2,
figsize=(4, 6),
sharey=True,
gridspec_kw=dict(width_ratios=(1, 6)),
)
ax_y.pcolormesh(Y[:nt], cmap="magma")
ax_x.pcolormesh(X[:nt], cmap="coolwarm")
ax_y.set(
title="Y (Spikes)",
ylabel="Time point",
)
ax_x.set(
title="X (Lagged stimulus)",
xlabel="Time lag",
xticks=[],
)
ax_y.invert_yaxis()
f.tight_layout()
def plot_spike_filter(theta, dt, **kws):
"""Plot estimated weights based on time lag model.
Args:
theta (1D array): Filter weights, not including DC term.
dt (number): Duration of each time bin.
kws: Pass additional keyword arguments to plot()
"""
d = len(theta)
t = np.arange(-d + 1, 1) * dt
ax = plt.gca()
ax.plot(t, theta, marker="o", **kws)
ax.axhline(0, color=".2", linestyle="--", zorder=1)
ax.set(
xlabel="Time before spike (s)",
ylabel="Filter weight",
)
def plot_spikes_with_prediction(
spikes, predicted_spikes, dt, nt=50, t0=120, **kws):
"""Plot actual and predicted spike counts.
Args:
spikes (1D array): Vector of actual spike counts
predicted_spikes (1D array): Vector of predicted spike counts
dt (number): Duration of each time bin.
nt (number): Number of time bins to plot
t0 (number): Index of first time bin to plot.
kws: Pass additional keyword arguments to plot()
"""
t = np.arange(t0, t0 + nt) * dt
f, ax = plt.subplots()
lines = ax.stem(t, spikes[:nt], use_line_collection=True)
plt.setp(lines, color=".5")
lines[-1].set_zorder(1)
kws.setdefault("linewidth", 3)
yhat, = ax.plot(t, predicted_spikes[:nt], **kws)
ax.set(
xlabel="Time (s)",
ylabel="Spikes",
)
ax.yaxis.set_major_locator(plt.MaxNLocator(integer=True))
ax.legend([lines[0], yhat], ["Spikes", "Predicted"])
# + cellView="form" colab={} colab_type="code" id="0bZa5Y5lLElv"
#@title Data retrieval
import os
data_filename = 'RGCdata.mat'
if data_filename not in os.listdir():
# !wget -qO $data_filename https://osf.io/mzujs/download
# + [markdown] colab_type="text" id="qmTMUaRWLq5e"
# -----
#
# + [markdown] colab_type="text" id="Ve3vEEP6uCgh"
# ## Linear-Gaussian GLM
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 518} colab_type="code" id="hzCW7VcxuElj" outputId="a2ecc7ad-fc1c-485e-e489-4450f6ebb2cb"
#@title Video: General linear model
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="oOHqjvDyrE8", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# + [markdown] colab_type="text" id="8iZQlSSz8hnd"
# ### Load retinal ganglion cell activity data
#
# In this exercise we will use data from an experiment studying retinal ganglion cell (RGC) responses to a "full-field flicker" binary white noise stimulus. In this experiment, a screen randomly alternated between two luminance values while spikes were recorded from 4 RGCs. The dataset includes the luminance of the screen on each refresh (~120Hz) and the number of spikes each neuron emitted in that time bin.
#
# The file `RGCdata.mat` contains three variablies:
#
# - `Stim`, the stimulus intensity at each time point. It is an array with shape $T \times 1$, where $T=144051$.
#
# - `SpCounts`, the binned spike counts for 2 ON cells, and 2 OFF cells. It is a $144051 \times 4$ array, and each column has counts for a different cell.
#
# - `dtStim`, the size of a single time bin (in seconds), which is needed for computing model output in units of spikes / s. The stimulus frame rate is given by `1 / dtStim`.
#
# Because these data were saved in MATLAB, where everything is a matrix, we will also process the variables to more Pythonic representations (1D arrays or scalars, where appropriate) as we load the data.
# + colab={} colab_type="code" id="G02xDLg08hnk"
data = loadmat('RGCdata.mat') # loadmat is a function in scipy.io
dt_stim = data['dtStim'].item() # .item extracts a scalar value
# Extract the stimulus intensity
stim = data['Stim'].squeeze() # .squeeze removes dimensions with 1 element
# Extract the spike counts for one cell
cellnum = 2
spikes = data['SpCounts'][:, cellnum]
# Don't use all of the timepoints in the dataset, for speed
keep_timepoints = 20000
stim = stim[:keep_timepoints]
spikes = spikes[:keep_timepoints]
# + [markdown] colab_type="text" id="DgdE1ihWAS83"
# Use the `plot_stim_and_spikes` helper function to visualize the changes in stimulus intensities and spike counts over time.
# + colab={"base_uri": "https://localhost:8080/", "height": 367} colab_type="code" id="VH6X4cbrOhNA" outputId="1e071780-e324-4f6f-acd6-33e637c1cd0b"
plot_stim_and_spikes(stim, spikes, dt_stim)
# + [markdown] colab_type="text" id="vhDwPb0E8hoA"
# ### Exercise: Create design matrix
#
# Our goal is to predict the cell's activity from the stimulus intensities preceding it. That will help us understand how RGCs process information over time. To do so, we first need to create the *design matrix* for this model, which organizes the stimulus intensities in matrix form such that the $i$th row has the stimulus frames preceding timepoint $i$.
#
# In this exercise, we will create the design matrix $X$ using $d=25$ time lags. That is, $X$ should be a $T \times d$ matrix. $d = 25$ (about 200 ms) is a choice we're making based on our prior knowledge of the temporal window that influences RGC responses. In practice, you might not know the right duration to use.
#
# The last entry in row `t` should correspond to the stimulus that was shown at time `t`, the entry to the left of it should contain the value that was show one time bin earlier, etc. Specifically, $X_{ij}$ will be the stimulus intensity at time $i + d - 1 - j$.
#
# Assume values of `stim` are 0 for the time lags prior to the first timepoint in the dataset. (This is known as "zero-padding", so that the design matrix has the same number of rows as the response vectors in `spikes`.)
#
# Your tasks are to
#
# - make a zero-padded version of the stimulus
# - initialize an empty design matrix with the correct shape
# - fill in each row of the design matrix, using the stimulus information
#
# To visualize your design matrix (and the corresponding vector of spike counts), we will plot a "heatmap", which encodes the numerical value in each position of the matrix as a color. The helper functions include some code to do this.
# + colab={} colab_type="code" id="wnOc5lK4Z74o"
def make_design_matrix(stim, d=25):
"""Create time-lag design matrix from stimulus intensity vector.
Args:
stim (1D array): Stimulus intensity at each time point.
d (number): Number of time lags to use.
Returns
X (2D array): GLM design matrix with shape T, d
"""
#####################################################################
# Fill in missing code (...) and then remove
raise NotImplementedError("Complete the make_design_matrix function")
#####################################################################
# Create version of stimulus vector with zeros before onset
padded_stim = ...
# Construct a matrix where each row has the d frames of
# the stimulus proceeding and including timepoint t
T = ... # Total number of timepoints
X = ...
for t in range(T):
X[t] = ...
return X
# Uncomment and run after completing `make_design_matrix`
# X = make_design_matrix(stim)
# plot_glm_matrices(X, spikes, nt=50)
# + colab={"base_uri": "https://localhost:8080/", "height": 432} colab_type="code" id="Wpzp2rxCabUb" outputId="c300adb4-83f7-4dcb-f637-b8dc33d30d3c"
# to_remove solution
def make_design_matrix(stim, d=25):
"""Create time-lag design matrix from stimulus intensity vector.
Args:
stim (1D array): Stimulus intensity at each time point.
d (number): Number of time lags to use.
Returns
X (2D array): GLM design matrix with shape T, d
"""
padded_stim = np.concatenate([np.zeros(d - 1), stim])
T = len(stim)
X = np.zeros((T, d))
for t in range(T):
X[t] = padded_stim[t:t + d]
return X
with plt.xkcd():
X = make_design_matrix(stim)
plot_glm_matrices(X, spikes, nt=50)
# + [markdown] colab_type="text" id="QxFwPdWn8hoV"
# ### Fit Linear-Gaussian regression model
#
# First, we will use the design matrix to compute the maximum likelihood estimate for a linear-Gaussian GLM (aka "general linear model"). The maximum likelihood estimate of $\theta$ in this model can be solved analytically using the equation you learned about on Day 3:
#
# $$\hat \theta = (X^TX)^{-1}X^Ty$$
#
# Before we can apply this equation, we need to augment the design matrix to account for the mean of $y$, because the spike counts are all $\geq 0$. We do this by adding a constant column of 1's to the design matrix, which will allow the model to learn an additive offset weight. We will refer to this additional weight as $b$ (for bias), although it is alternatively known as a "DC term" or "intercept".
# + colab={} colab_type="code" id="qbHiZvChkyv4"
# Build the full design matrix
y = spikes
constant = np.ones_like(y)
X = np.column_stack([constant, make_design_matrix(stim)])
# Get the MLE weights for the LG model
all_theta = np.linalg.inv(X.T @ X) @ X.T @ y
theta_lg = all_theta[1:]
# + [markdown] colab_type="text" id="6Ce0QArC8hoZ"
# Plot the resulting maximum likelihood filter estimate (just the 25-element weight vector $\theta$ on the stimulus elements, not the DC term $b$).
# + colab={"base_uri": "https://localhost:8080/", "height": 277} colab_type="code" id="2BOMuZHFmEka" outputId="83946327-214e-4378-f948-d79995d9c6fc"
plot_spike_filter(theta_lg, dt_stim)
# + [markdown] colab_type="text" id="N8ViEJyIpV3-"
# ---
#
# ### Exercise: Predict spike counts with Linear-Gaussian model
#
# Now we are going to put these pieces together and write a function that outputs a predicted spike count for each timepoint using the stimulus information.
#
# Your steps should be:
#
# - Create the complete design matrix
# - Obtain the MLE weights ($\hat \theta$)
# - Compute $\hat y = X\hat \theta$
# + colab={} colab_type="code" id="9eeX7EMqq3mk"
def predict_spike_counts_lg(stim, spikes, d=25):
"""Compute a vector of predicted spike counts given the stimulus.
Args:
stim (1D array): Stimulus values at each timepoint
spikes (1D array): Spike counts measured at each timepoint
d (number): Number of time lags to use.
Returns:
yhat (1D array): Predicted spikes at each timepoint.
"""
#####################################################################
# Fill in missing code (...) and then remove
raise NotImplementedError(
"Complete the predict_spike_counts_lg function"
)
#####################################################################
# Create the design matrix
...
# Get the MLE weights for the LG model
...
# Compute predicted spike counts
yhat = ...
return yhat
# Uncomment and run after completing the function to plot the prediction
# predicted_counts = predict_spike_counts_lg(stim, spikes)
# plot_spikes_with_prediction(spikes, predicted_counts, dt_stim)
# + colab={"base_uri": "https://localhost:8080/", "height": 321} colab_type="code" id="PazkvLjCryTY" outputId="f947c76c-b45f-4a55-e3b1-5350c1f4a119"
# to_remove solution
def predict_spike_counts_lg(stim, spikes, d=25):
"""Compute a vector of predicted spike counts given the stimulus.
Args:
stim (1D array): Stimulus values at each timepoint
spikes (1D array): Spike counts measured at each timepoint
d (number): Number of time lags to use.
Returns:
yhat (1D array): Predicted spikes at each timepoint.
"""
y = spikes
constant = np.ones_like(y)
X = np.column_stack([constant, make_design_matrix(stim)])
theta = np.linalg.inv(X.T @ X) @ X.T @ y
yhat = X @ theta
return yhat
predicted_counts = predict_spike_counts_lg(stim, spikes)
with plt.xkcd():
plot_spikes_with_prediction(spikes, predicted_counts, dt_stim)
# + [markdown] colab_type="text" id="LhS1zfbV8hor"
# Is this a good model? The prediction line more-or-less follows the bumps in the spikes, but it never predicts as many spikes as are actually observed. And, more troublingly, it's predicting *negative* spikes for some time points.
#
# The Poisson GLM will help to address these failures.
#
#
# ### Bonus challenge
#
# The "spike-triggered average" falls out as a subcase of the linear Gaussian GLM: $\mathrm{STA} = X^T y \,/\, \textrm{sum}(y)$, where $y$ is the vector of spike counts of the neuron. In the LG GLM, the term $(X^TX)^{-1}$ corrects for potential correlation between the regressors. Because the experiment that produced these data used a white note stimulus, there are no such correlations. Therefore the two methods are equivalent. (How would you check the statement about no correlations?)
# + [markdown] colab_type="text" id="ax5n9J648hov"
# ## Linear-Nonlinear-Poisson GLM
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 518} colab_type="code" id="_sp9v00rygPw" outputId="a070662f-4ade-46ed-e401-e12e5af83efc"
#@title Video: Generalized linear model
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="eAd2ILUrPyE", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# + [markdown] colab_type="text" id="vQr090AR2j1R"
# ### Nonlinear optimization with `scipy.optimize`
#
# When we used the Linear-Gaussian model, we were able to obtain the maximum likelihood estimate for the parameter vector in a single line of code, because there is an analytical solution for that model. In the more general case, we don't have an analytical solution. Instead, we need to apply a nonlinear optimization algorithm to find the parameter values that minimize some *objective function*.
#
# Note: when using this approach to perform maximum likelihood estimation, the objective function should return the *negative* log likelihood, because optimization algorithms are written with the convention that minimization is your goal.
#
# The `scipy.optimize` module has a powerful function called [`minimize`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html) that provides a generic interface to a large number of optimization algorithms. The way it works is that you pass an objective function object and an "initial guess" for the parameter values. It then returns an dictionary that includes the minimum function value, the parameters that give this minimum, and other information.
#
# Let's see how this works with a simple example.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ewE5w7QW5sfh" outputId="a0b8798e-b28e-4161-e68f-767ae529cc3f"
f = np.square
res = minimize(f, x0=2) # Imported from scipy.optimize in a hidden cell
print(
f"Minimum value: {res['fun']:.4g}",
f"at x = {res['x']}",
)
# + [markdown] colab_type="text" id="N3dgrYlQ7DCb"
# When minimizing a $f(x) = x^2$, we get a minimum value of $f(x) \approx 0$ when $x \approx 0$. The algorithm doesn't return exactly $0$, because it stops when it gets "close enough" to a minimum. You can change the `tol` parameter to control how it defines "close enough".
#
# A point about the code bears emphasis. The first argument to `minimize` is not just a number or a string but a *function*. Here, we used `np.square`. Take a moment to make sure you understand what's going on here, because it's a bit unusual, and it will be important for the exercise you're going to do in a moment.
#
# In this example, we started at $x_0 = 2$. Let's try different values for the starting point:
# + colab={"base_uri": "https://localhost:8080/", "height": 277} colab_type="code" id="onBX1pq-7ul5" outputId="6583b942-1fcb-43ef-c912-b6d8c366bd34"
f = np.square
start_points = -1, 1.5
xx = np.linspace(-2, 2, 100)
plt.plot(xx, f(xx), color=".2")
plt.xlabel("$x$")
plt.ylabel("$f(x)$")
for i, x0 in enumerate(start_points):
res = minimize(f, x0)
plt.plot(x0, f(x0), "o", color=f"C{i}", ms=10, label=f"Start {i}")
plt.plot(res["x"].item(), res["fun"], "x", c=f"C{i}", ms=10, mew=2, label=f"End {i}")
plt.legend()
# + [markdown] colab_type="text" id="Ri_KY6fN8ZMP"
# The three runs started at different points (the dots), but they each ended up at roughly the same place (the cross): $f(x_\textrm{final}) \approx 0$. Let's see what happens if we use a different function:
# + colab={"base_uri": "https://localhost:8080/", "height": 277} colab_type="code" id="GCytz2Gt8xSS" outputId="8cafd2bf-8454-4f76-bfcf-8e3522e8af4f"
g = lambda x: x / 5 + np.cos(x)
start_points = -.5, 1.5
xx = np.linspace(-4, 4, 100)
plt.plot(xx, g(xx), color=".2")
plt.xlabel("$x$")
plt.ylabel("$f(x)$")
for i, x0 in enumerate(start_points):
res = minimize(g, x0)
plt.plot(x0, g(x0), "o", color=f"C{i}", ms=10, label=f"Start {i}")
plt.plot(res["x"].item(), res["fun"], "x", color=f"C{i}", ms=10, mew=2, label=f"End {i}")
plt.legend()
# + [markdown] colab_type="text" id="GqVaWzUE9WH7"
# Unlike $f(x) = x^2$, $g(x) = \frac{x}{5} + \cos(x)$ is not *convex*. That means that the final position of the minimization algorithm depends on the starting point. In practice, one way to deal with this would be to try a number of different starting points and then use the parameters that give the minimum value value across all runs. But we won't worry about that for now.
# + [markdown] colab_type="text" id="8xMcD_hf8how"
# ### Exercise: Fitting the Poisson GLM and prediction spikes
#
# In this exercise, we will use [`scipy.optimize.minimize`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html) to compute maximum likelihood estimates for the filter weights in the Poissson GLM model with an exponential nonlinearity (LNP: Linear-Nonlinear-Poisson).
#
# In practice, this will involve filling out two functions.
#
# - The first should be an *objective function* that takes a design matrix, a spike count vector, and a vector of parameters. It should return a negative log likelihood.
# - The second function should take `stim` and `spikes`, build the design matrix and then use `minimize` internally, and return the MLE parameters.
#
# What should the objective function look like? We want it to return
#
# $$-\log \mathcal{L} = -\log P(y \mid X, \theta).$$
#
# In the Poisson GLM,
#
# $$
# \log P(y \mid X, \theta) = \sum_t \log P(y_t \mid \mathbf{x_t},\theta)
# $$
#
# and
#
# $$
# P(y_t \mid \mathbf{x_t}, \theta) \
# = \frac{\lambda^{y_t}\exp(-\lambda)}{y_t!}
# $$
#
# with $$\lambda = \exp(\theta^T \mathbf{x_t}).$$
#
# So we can get the log likelihood for all the data with
#
# $$
# \log \mathcal{L} = \sum_t y_t \log(\theta^T \mathbf{x_t}) - \theta^T \mathbf{x_t}
# $$.
#
# *Tip: Starting with a loop is the most obvious way to implement this equation, but it will be also be slow. Can you get the log likelihood for all trials using matrix operations?*
# + colab={} colab_type="code" id="ac3z2RvXROzp"
def neg_log_lik_lnp(theta, X, y):
"""Return -loglike for the Poisson GLM model.
Args:
theta (1D array): Parameter vector.
X (2D array): Full design matrix.
y (1D array): Data values.
Returns:
number: Negative log likelihood.
"""
#####################################################################
# Fill in missing code (...) and then remove
raise NotImplementedError("Complete the neg_log_lik_lnp function")
#####################################################################
# Compute the Poisson log likeliood
log_lik = ...
return ...
def fit_lnp(X, y, d=25):
"""Obtain MLE parameters for the Poisson GLM.
Args:
X (2D array): Full design matrix.
y (1D array): Data values.
d (number): Number of time lags to use.
Returns:
1D array: MLE parameters
"""
#####################################################################
# Fill in missing code (...) and then remove
raise NotImplementedError("Complete the fit_lnp function")
#####################################################################
# Build the design matrix
y = spikes
constant = np.ones_like(y)
X = np.column_stack([constant, make_design_matrix(stim)])
# Use a random vector of weights to start (mean 0, sd .2)
x0 = np.random.normal(0, .2, d + 1)
# Find parameters that minmize the negative log likelihood function
res = minimize(..., args=(X, y))
return ...
# Uncomment and run when the functions are ready
# theta_lnp = fit_lnp(X, spikes)
# plot_spike_filter(theta_lg[1:], dt_stim, color=".5", label="LG")
# plot_spike_filter(theta_lnp[1:], dt_stim, label="LNP")
# plt.legend(loc="upper left");
# + colab={"base_uri": "https://localhost:8080/", "height": 321} colab_type="code" id="udyGHYlkHpkM" outputId="305fc735-b5b5-4d07-a9c2-8d453e9e362c"
# to_remove solution
def neg_log_lik_lnp(theta, X, y):
"""Return -loglike for the Poisson GLM model.
Args:
theta (1D array): Parameter vector.
X (2D array): Full design matrix.
y (1D array): Data values.
Returns:
number: Negative log likelihood.
"""
rate = np.exp(X @ theta)
loglik = np.log(rate) @ y - rate.sum()
return -loglik
def fit_lnp(stim, spikes, d=25):
"""Obtain MLE parameters for the Poisson GLM.
Args:
stim (1D array): Stimulus values at each timepoint
spikes (1D array): Spike counts measured at each timepoint
d (number): Number of time lags to use.
Returns:
1D array: MLE parameters
"""
y = spikes
constant = np.ones_like(spikes)
X = np.column_stack([constant, make_design_matrix(stim)])
x0 = np.random.normal(0, .2, d + 1)
res = minimize(neg_log_lik_lnp, x0, args=(X, y))
return res["x"]
theta_lnp = fit_lnp(stim, spikes)
with plt.xkcd():
plot_spike_filter(theta_lg[1:], dt_stim, color=".5", label="LG")
plot_spike_filter(theta_lnp[1:], dt_stim, label="LNP")
plt.legend(loc="upper left");
# + [markdown] colab_type="text" id="EvqdySzYTTKu"
# Plotting the LG and LNP weights together, we see that they are broadly similar, but the LNP weights are generally larger. What does that mean for the model's ability to *predict* spikes? To see that, let's finish the exercise by filling out the `predict_spike_counts_lnp` function:
# + colab={} colab_type="code" id="i1JhwAkXSXOK"
def predict_spike_counts_lnp(stim, spikes, theta=None, d=25):
"""Compute a vector of predicted spike counts given the stimulus.
Args:
stim (1D array): Stimulus values at each timepoint
spikes (1D array): Spike counts measured at each timepoint
theta (1D array): Filter weights; estimated if not provided.
d (number): Number of time lags to use.
Returns:
yhat (1D array): Predicted spikes at each timepoint.
"""
###########################################################################
# Fill in missing code (...) and then remove
raise NotImplementedError("Complete the predict_spike_counts_lnp function")
###########################################################################
y = spikes
constant = np.ones_like(spikes)
X = np.column_stack([constant, make_design_matrix(stim)])
if theta is None: # Allow pre-cached weights, as fitting is slow
theta = fit_lnp(X, y, d)
yhat = ...
return yhat
# Uncomment and run when predict_spike_counts_lnp is complete
# yhat = predict_spike_counts_lnp(stim, spikes, theta_lnp)
# plot_spikes_with_prediction(spikes, yhat, dt_stim)
# + colab={"base_uri": "https://localhost:8080/", "height": 285} colab_type="code" id="4_h1I69HN5DB" outputId="521c45c1-1d0d-4c2b-ce32-0a2285acfb53"
# to_remove solution
def predict_spike_counts_lnp(stim, spikes, theta=None, d=25):
"""Compute a vector of predicted spike counts given the stimulus.
Args:
stim (1D array): Stimulus values at each timepoint
spikes (1D array): Spike counts measured at each timepoint
theta (1D array): Filter weights; estimated if not provided.
d (number): Number of time lags to use.
Returns:
yhat (1D array): Predicted spikes at each timepoint.
"""
y = spikes
constant = np.ones_like(spikes)
X = np.column_stack([constant, make_design_matrix(stim)])
if theta is None:
theta = fit_lnp(X, y, d)
yhat = np.exp(X @ theta)
return yhat
yhat = predict_spike_counts_lnp(stim, spikes, theta_lnp)
with plt.xkcd():
plot_spikes_with_prediction(spikes, yhat, dt_stim)
# + [markdown] colab_type="text" id="bufTaOcdTtnm"
# We see that the LNP model does a better job of fitting the actual spiking data. Importantly, it never predicts negative spikes!
#
# *Bonus:* Our statement that the LNP model "does a better job" is qualitative and based mostly on the visual appearance of the plot. But how would you make this a quantitative statement?
# + [markdown] colab_type="text" id="50ybKB24SzCb"
# ## Summary
#
# In this first tutorial, we used two different models to learn something about how retinal ganglion cells respond to a flickering white noise stimulus. We learned how to construct a design matrix that we could pass to different GLMs, and we found that the Linear-Nonlinear-Poisson (LNP) model allowed us to predict spike rates better than a simple Linear-Gaussian (LG) model.
#
# In the next tutorial, we'll extend these ideas further. We'll meet yet another GLM — logistic regression — and we'll learn how to ensure good model performance with large, high-dimensional datasets.
|
tutorials/W1D4_MachineLearning/W1D4_Tutorial1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/indahpuspitaa17/TensorFlow-Data-and-Deployment/blob/main/rest_simple.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="MhoQ0WE77laV"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="_ckMIh7O7s6D"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="jYysdyb-CaWM"
# # Train and serve a TensorFlow model with TensorFlow Serving
# + [markdown] id="E6FwTNtl3S4v"
# **Warning: This notebook is designed to be run in a Google Colab only**. It installs packages on the system and requires root access. If you want to run it in a local Jupyter notebook, please proceed with caution.
#
# Note: You can run this example right now in a Jupyter-style notebook, no setup required! Just click "Run in Google Colab"
#
# <div class="devsite-table-wrapper"><table class="tfo-notebook-buttons" align="left">
# <tr><td><a target="_blank" href="https://www.tensorflow.org/tfx/tutorials/serving/rest_simple">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/tfx/blob/master/docs/tutorials/serving/rest_simple.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png">Run in Google Colab</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/tfx/blob/master/docs/tutorials/serving/rest_simple.ipynb">
# <img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">View source on GitHub</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/tfx/docs/tutorials/serving/rest_simple.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a></td>
# </tr></table></div>
# + [markdown] id="FbVhjPpzn6BM"
# This guide trains a neural network model to classify [images of clothing, like sneakers and shirts](https://github.com/zalandoresearch/fashion-mnist), saves the trained model, and then serves it with [TensorFlow Serving](https://www.tensorflow.org/serving/). The focus is on TensorFlow Serving, rather than the modeling and training in TensorFlow, so for a complete example which focuses on the modeling and training see the [Basic Classification example](https://github.com/tensorflow/docs/blob/master/site/en/r1/tutorials/keras/basic_classification.ipynb).
#
# This guide uses [tf.keras](https://github.com/tensorflow/docs/blob/master/site/en/r1/guide/keras.ipynb), a high-level API to build and train models in TensorFlow.
# + id="FWkuJabJSKGB"
import sys
# Confirm that we're using Python 3
assert sys.version_info.major is 3, 'Oops, not running Python 3. Use Runtime > Change runtime type'
# + id="dzLKpmZICaWN" colab={"base_uri": "https://localhost:8080/"} outputId="31cd1144-67ac-49bd-d9ab-aa6597986c14"
# TensorFlow and tf.keras
print("Installing dependencies for Colab environment")
# !pip install -Uq grpcio==1.26.0
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
import os
import subprocess
print('TensorFlow version: {}'.format(tf.__version__))
# + [markdown] id="5jAk1ZXqTJqN"
# ## Create your model
# + [markdown] id="yR0EdgrLCaWR"
# ### Import the Fashion MNIST dataset
#
# This guide uses the [Fashion MNIST](https://github.com/zalandoresearch/fashion-mnist) dataset which contains 70,000 grayscale images in 10 categories. The images show individual articles of clothing at low resolution (28 by 28 pixels), as seen here:
#
# <table>
# <tr><td>
# <img src="https://tensorflow.org/images/fashion-mnist-sprite.png"
# alt="Fashion MNIST sprite" width="600">
# </td></tr>
# <tr><td align="center">
# <b>Figure 1.</b> <a href="https://github.com/zalandoresearch/fashion-mnist">Fashion-MNIST samples</a> (by Zalando, MIT License).<br/>
# </td></tr>
# </table>
#
# Fashion MNIST is intended as a drop-in replacement for the classic [MNIST](http://yann.lecun.com/exdb/mnist/) dataset—often used as the "Hello, World" of machine learning programs for computer vision. You can access the Fashion MNIST directly from TensorFlow, just import and load the data.
#
# Note: Although these are really images, they are loaded as NumPy arrays and not binary image objects.
# + id="7MqDQO0KCaWS" colab={"base_uri": "https://localhost:8080/"} outputId="93cd4642-b4cd-4220-f315-176a16a7527f"
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# scale the values to 0.0 to 1.0
train_images = train_images / 255.0
test_images = test_images / 255.0
# reshape for feeding into the model
train_images = train_images.reshape(train_images.shape[0], 28, 28, 1)
test_images = test_images.reshape(test_images.shape[0], 28, 28, 1)
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
print('\ntrain_images.shape: {}, of {}'.format(train_images.shape, train_images.dtype))
print('test_images.shape: {}, of {}'.format(test_images.shape, test_images.dtype))
# + [markdown] id="PDu7OX8Nf5PY"
# ### Train and evaluate your model
#
# Let's use the simplest possible CNN, since we're not focused on the modeling part.
# + id="LTNN0ANGgA36" colab={"base_uri": "https://localhost:8080/"} outputId="fd8f5f34-885e-4449-bd1a-007b949a8fdf"
model = keras.Sequential([
keras.layers.Conv2D(input_shape=(28,28,1), filters=8, kernel_size=3,
strides=2, activation='relu', name='Conv1'),
keras.layers.Flatten(),
keras.layers.Dense(10, name='Dense')
])
model.summary()
testing = False
epochs = 5
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[keras.metrics.SparseCategoricalAccuracy()])
model.fit(train_images, train_labels, epochs=epochs)
test_loss, test_acc = model.evaluate(test_images, test_labels)
print('\nTest accuracy: {}'.format(test_acc))
# + [markdown] id="AwGPItyphqXT"
# ## Save your model
#
# To load our trained model into TensorFlow Serving we first need to save it in [SavedModel](https://www.tensorflow.org/versions/r1.15/api_docs/python/tf/saved_model) format. This will create a protobuf file in a well-defined directory hierarchy, and will include a version number. [TensorFlow Serving](https://www.tensorflow.org/tfx/guide/serving) allows us to select which version of a model, or "servable" we want to use when we make inference requests. Each version will be exported to a different sub-directory under the given path.
# + id="0w5Rq8SsgWE6" colab={"base_uri": "https://localhost:8080/"} outputId="3ce1a3de-5cd6-45ac-8557-9d36309cec01"
# Fetch the Keras session and save the model
# The signature definition is defined by the input and output tensors,
# and stored with the default serving key
import tempfile
MODEL_DIR = tempfile.gettempdir()
version = 1
export_path = os.path.join(MODEL_DIR, str(version))
print('export_path = {}\n'.format(export_path))
tf.keras.models.save_model(
model,
export_path,
overwrite=True,
include_optimizer=True,
save_format=None,
signatures=None,
options=None
)
print('\nSaved model:')
# !ls -l {export_path}
# + [markdown] id="FM7B_RuDYoIj"
# ## Examine your saved model
#
# We'll use the command line utility `saved_model_cli` to look at the [MetaGraphDefs](https://www.tensorflow.org/versions/r1.15/api_docs/python/tf/MetaGraphDef) (the models) and [SignatureDefs](../signature_defs) (the methods you can call) in our SavedModel. See [this discussion of the SavedModel CLI](https://github.com/tensorflow/docs/blob/master/site/en/r1/guide/saved_model.md#cli-to-inspect-and-execute-savedmodel) in the TensorFlow Guide.
# + id="LU4GDF_aYtfQ" colab={"base_uri": "https://localhost:8080/"} outputId="558c3dd0-de4b-42d7-b380-191f36e26df8"
# !saved_model_cli show --dir {export_path} --all
# + [markdown] id="lSPWuegUb7Eo"
# That tells us a lot about our model! In this case we just trained our model, so we already know the inputs and outputs, but if we didn't this would be important information. It doesn't tell us everything, like the fact that this is grayscale image data for example, but it's a great start.
# + [markdown] id="DBgsyhytS6KD"
# ## Serve your model with TensorFlow Serving
#
# **Warning: If you are running this NOT on a Google Colab,** following cells
# will install packages on the system with root access. If you want to run it in
# a local Jupyter notebook, please proceed with caution.
#
# ### Add TensorFlow Serving distribution URI as a package source:
#
# We're preparing to install TensorFlow Serving using [Aptitude](https://wiki.debian.org/Aptitude) since this Colab runs in a Debian environment. We'll add the `tensorflow-model-server` package to the list of packages that Aptitude knows about. Note that we're running as root.
#
# Note: This example is running TensorFlow Serving natively, but [you can also run it in a Docker container](https://www.tensorflow.org/tfx/serving/docker), which is one of the easiest ways to get started using TensorFlow Serving.
# + id="v2hF_ChoOrEd"
import sys
# We need sudo prefix if not on a Google Colab.
if 'google.colab' not in sys.modules:
SUDO_IF_NEEDED = 'sudo'
else:
SUDO_IF_NEEDED = ''
# + id="EWg9X2QHlbGS" colab={"base_uri": "https://localhost:8080/"} outputId="fbd020e6-c4bf-457e-e3e1-98d3ca74c83c"
# This is the same as you would do from your command line, but without the [arch=amd64], and no sudo
# You would instead do:
# # echo "deb [arch=amd64] http://storage.googleapis.com/tensorflow-serving-apt stable tensorflow-model-server tensorflow-model-server-universal" | sudo tee /etc/apt/sources.list.d/tensorflow-serving.list && \
# # curl https://storage.googleapis.com/tensorflow-serving-apt/tensorflow-serving.release.pub.gpg | sudo apt-key add -
# !echo "deb http://storage.googleapis.com/tensorflow-serving-apt stable tensorflow-model-server tensorflow-model-server-universal" | {SUDO_IF_NEEDED} tee /etc/apt/sources.list.d/tensorflow-serving.list && \
# curl https://storage.googleapis.com/tensorflow-serving-apt/tensorflow-serving.release.pub.gpg | {SUDO_IF_NEEDED} apt-key add -
# !{SUDO_IF_NEEDED} apt update
# + [markdown] id="W1ZVp_VOU7Wu"
# ### Install TensorFlow Serving
#
# This is all you need - one command line!
# + id="ygwa9AgRloYy" colab={"base_uri": "https://localhost:8080/"} outputId="7af79013-a70d-4317-b005-44275e0d4ff4"
# !{SUDO_IF_NEEDED} apt-get install tensorflow-model-server
# + [markdown] id="k5NrYdQeVm52"
# ### Start running TensorFlow Serving
#
# This is where we start running TensorFlow Serving and load our model. After it loads we can start making inference requests using REST. There are some important parameters:
#
# * `rest_api_port`: The port that you'll use for REST requests.
# * `model_name`: You'll use this in the URL of REST requests. It can be anything.
# * `model_base_path`: This is the path to the directory where you've saved your model.
#
# + id="aUgp3vUdU5GS"
os.environ["MODEL_DIR"] = MODEL_DIR
# + id="kJDhHNJVnaLN" colab={"base_uri": "https://localhost:8080/"} outputId="5faa74e3-b65c-4190-b724-bdb1d59ac12a" magic_args="--bg " language="bash"
# nohup tensorflow_model_server \
# --rest_api_port=8501 \
# --model_name=fashion_model \
# --model_base_path="${MODEL_DIR}" >server.log 2>&1
#
# + id="IxbeiOCUUs2z" colab={"base_uri": "https://localhost:8080/"} outputId="0613d18c-1b59-461e-bbee-f89357c742fa"
# !tail server.log
# + [markdown] id="vwg1JKaGXWAg"
# ## Make a request to your model in TensorFlow Serving
#
# First, let's take a look at a random example from our test data.
# + id="Luqm_Jyff9iR" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="cb652f75-27b6-4fc8-8376-0171a56325dd"
def show(idx, title):
plt.figure()
plt.imshow(test_images[idx].reshape(28,28))
plt.axis('off')
plt.title('\n\n{}'.format(title), fontdict={'size': 16})
import random
rando = random.randint(0,len(test_images)-1)
show(rando, 'An Example Image: {}'.format(class_names[test_labels[rando]]))
# + [markdown] id="TKnEHeTrbh3L"
# Ok, that looks interesting. How hard is that for you to recognize? Now let's create the JSON object for a batch of three inference requests, and see how well our model recognizes things:
# + id="2dsD7KQG1m-R" colab={"base_uri": "https://localhost:8080/"} outputId="c0c9a131-1315-400e-cd94-ae6db0bc08de"
import json
data = json.dumps({"signature_name": "serving_default", "instances": test_images[0:3].tolist()})
print('Data: {} ... {}'.format(data[:50], data[len(data)-52:]))
# + [markdown] id="ReQd4QESIwXN"
# ### Make REST requests
# + [markdown] id="iT3J-lHrhOYQ"
# #### Newest version of the servable
#
# We'll send a predict request as a POST to our server's REST endpoint, and pass it three examples. We'll ask our server to give us the latest version of our servable by not specifying a particular version.
# + id="vGvFyuIzW6n6" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="07efa302-765d-4544-e7e7-f2baaa81946b"
# docs_infra: no_execute
# !pip install -q requests
import requests
headers = {"content-type": "application/json"}
json_response = requests.post('http://localhost:8501/v1/models/fashion_model:predict', data=data, headers=headers)
predictions = json.loads(json_response.text)['predictions']
show(0, 'The model thought this was a {} (class {}), and it was actually a {} (class {})'.format(
class_names[np.argmax(predictions[0])], np.argmax(predictions[0]), class_names[test_labels[0]], test_labels[0]))
# + [markdown] id="YJH8LtM4XELp"
# #### A particular version of the servable
#
# Now let's specify a particular version of our servable. Since we only have one, let's select version 1. We'll also look at all three results.
# + id="zRftRxeR1tZx" colab={"base_uri": "https://localhost:8080/", "height": 869} outputId="c2be3f67-4753-4396-94a7-a4e4b2e09e19"
# docs_infra: no_execute
headers = {"content-type": "application/json"}
json_response = requests.post('http://localhost:8501/v1/models/fashion_model/versions/1:predict', data=data, headers=headers)
predictions = json.loads(json_response.text)['predictions']
for i in range(0,3):
show(i, 'The model thought this was a {} (class {}), and it was actually a {} (class {})'.format(
class_names[np.argmax(predictions[i])], np.argmax(predictions[i]), class_names[test_labels[i]], test_labels[i]))
|
4-Advanced-Deployment-Scenarios-with-TensorFlow/rest_simple.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # multiplot tutorial
#
# Although the forthcoming inline plots are static, running this code in a Python shell will produce interactive matplotlib windows.
import pandas as pd
import numpy as np
import scipy.signal as signal
from multiplot import PandasPlot, NumpyPlot
# %matplotlib inline
# Generate a set of sample signals.
# +
samp_freq = 1000 # Hz
duration = 5 # seconds
first_signal_freq =1 # Hz
signals = []
labels = []
for x in xrange(1,6):
signal_freq = first_signal_freq * x
time_points = np.arange(0, duration, 1/float(samp_freq))
sig = np.sin(2 * np.pi * signal_freq * time_points)
sig_label = "Ch %d" %(x-1)
labels.append(sig_label)
signals.append(sig)
df = pd.DataFrame(np.transpose(signals), columns=labels)
nump = np.array(signals)
# -
# Note that PandasPlot expects a DataFrame where each series is a column, whereas NumpyPlot expects an array where each series is a row.
print 'DataFrame: ', df.shape
print 'Numpy array: ', nump.shape
PandasPlot(df)
NumpyPlot(nump, labels=labels) # if labels aren't supplied, 'Ch x' labels are auto-generated
# ### Reduce number of channels displayed at once
PandasPlot(df, num_display_chans=2)
# ### Reduce number of samples displayed at once
PandasPlot(df, num_display_samps=2000)
# ### Highlight segments of the signals
# +
highlights = {'Ch 0': [[2000, 3000]],
'Ch 2': [[1000, 2000], [3000, 4000]],
'Ch 4': [[2000, 3000]]}
PandasPlot(df, highlights=highlights)
|
tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as graph
import numpy as np
from numpy.fft import fft2, ifft2
import math
import cv2
from scipy.signal import gaussian, convolve2d
# +
def blur(img, kernel_size = 3):
dummy = np.copy(img)
h = np.eye(kernel_size) / kernel_size
dummy = convolve2d(dummy, h, mode = 'valid')
return dummy
def gaussian_kernel(kernel_size = 3):
h = gaussian(kernel_size, kernel_size / 3).reshape(kernel_size, 1)
h = np.dot(h, h.transpose())
h /= np.sum(h)
return h
def wiener_filter(img, kernel, K):
kernel /= np.sum(kernel)
dummy = np.copy(img)
dummy = fft2(dummy)
kernel = fft2(kernel, s = img.shape)
kernel = np.conj(kernel) / (np.abs(kernel) ** 2 + K)
dummy = dummy * kernel
dummy = np.abs(ifft2(dummy))
return dummy
# -
image = cv2.imread('./data/1.jpg')
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
img_h = image.shape[0]
img_w = image.shape[1]
graph.figure()
graph.xlabel("Original Image")
graph.gray()
graph.imshow(image) # 显示原图像
# +
kernel = gaussian_kernel(5)
filtered_img = wiener_filter(image, kernel, K = 10)
graph.figure()
graph.xlabel("Wiener Image")
graph.gray()
graph.imshow(filtered_img) # 显示原图像
|
deblurring_test/wiener_deblur_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Title: RP- Spatial Accessibility of COVID-19 Healthcare Resources in Illinois
# ---
#
# **Reproduction of**: Rapidly measuring spatial accessibility of COVID-19 healthcare resources: a case study of Illinois, USA
#
# Original study *by* <NAME>., <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>. 2020. Rapidly measuring spatial accessibility of COVID-19 healthcare resources: a case study of Illinois, USA. International Journal of Health Geographics 19 (1):1–17. DOI:[10.1186/s12942-020-00229-x](https://ij-healthgeographics.biomedcentral.com/articles/10.1186/s12942-020-00229-x).
#
# Reproduction Authors: <NAME>, <NAME>, and <NAME>
# With contributions from <NAME>, <NAME>, and the Spring 2021 Open Source GIScience class at Middlebury
#
# Reproduction Materials Available at: [github.com/HEGSRR/RPr-Kang-2020](https://github.com/HEGSRR/RPr-Kang-2020)
#
# Created: `2021-06-01`
# Revised: `2021-08-19`
# ### Original Data
# To perform the ESFCA method, three types of data are required, as follows: (1) road network, (2) population, and (3) hospital information. The road network can be obtained from the [OpenStreetMap Python Library, called OSMNX](https://github.com/gboeing/osmnx). The population data is available on the [American Community Survey](https://data.census.gov/cedsci/deeplinks?url=https%3A%2F%2Ffactfinder.census.gov%2F&tid=GOVSTIMESERIES.CG00ORG01). Lastly, hosptial information is also publically available on the [Homelanad Infrastructure Foundation-Level Data](https://hifld-geoplatform.opendata.arcgis.com/datasets/hospitals?geometry=-94.504%2C40.632%2C-80.980%2C43.486).
# ### Introduction
#
# to be written. a draft can be found in docs/report/preanalysis.md
# ### Materials and Methods
# to be written.
# ### Deviatons from & Improvements to the Original Code
#
# to be written
# ### Modules
# Import necessary libraries to run this model.
# See `requirements.txt` for the library versions used for this analysis.
# +
# Import modules
import numpy as np
import pandas as pd
import geopandas as gpd
import networkx as nx
import osmnx as ox
import re
from shapely.geometry import Point, LineString, Polygon
import matplotlib.pyplot as plt
from tqdm import tqdm
import multiprocessing as mp
import folium
import itertools
import os
import time
import warnings
import IPython
from IPython.display import display, clear_output
warnings.filterwarnings("ignore")
# -
# ## Check Directories
#
# Because we have restructured the repository for replication, we need to check our working directory and make necessary adjustments.
# Check working directory
os.getcwd()
# Use to set work directory properly
if os.path.basename(os.getcwd()) == 'code':
os.chdir('../../')
os.getcwd()
# ## Load and Visualize Data
#
# ### Population and COVID-19 Cases Data by County
#
# *'Cases' column is coming in as 'Unnamed_0' --> easy to rename but this probably should be reportede to the original authors*
# If you would like to use the data generated from the pre-processing scripts, use the following code:
#
# ```py
# covid_data = gpd.read_file('./data/raw/public/Pre-Processing/covid_pre-processed.shp')
# atrisk_data = gpd.read_file('./data/raw/public/Pre-Processing/atrisk_pre-processed.shp')
# ```
# Read in at risk population data
atrisk_data = gpd.read_file('./data/raw/public/PopData/Chicago_Tract.shp')
atrisk_data.head()
# Read in covid case data
covid_data = gpd.read_file('./data/raw/public/PopData/Chicago_ZIPCODE.shp')
covid_data['cases'] = covid_data['cases']
covid_data.head()
# ### Load Hospital Data
#
# Note that 999 is treated as a "NULL"/"NA" so these hospitals are filtered out. This data contains the number of ICU beds and ventilators at each hospital.
# Read in hospital data
hospitals = gpd.read_file('./data/raw/public/HospitalData/Chicago_Hospital_Info.shp')
hospitals.head()
# ### Generate and Plot Map of Hospitals
# +
# Plot hospital data
m = folium.Map(location=[41.85, -87.65], tiles='cartodbpositron', zoom_start=10)
for i in range(0, len(hospitals)):
folium.CircleMarker(
location=[hospitals.iloc[i]['Y'], hospitals.iloc[i]['X']],
popup="{}{}\n{}{}\n{}{}".format('Hospital Name: ',hospitals.iloc[i]['Hospital'],
'ICU Beds: ',hospitals.iloc[i]['Adult ICU'],
'Ventilators: ', hospitals.iloc[i]['Total Vent']),
radius=5,
color='blue',
fill=True,
fill_opacity=0.6,
legend_name = 'Hospitals'
).add_to(m)
legend_html = '''<div style="position: fixed; width: 20%; heigh: auto;
bottom: 10px; left: 10px;
solid grey; z-index:9999; font-size:14px;
"> Legend<br>'''
m
# -
# ### Load and Plot Hexagon Grids (500-meter resolution)
# Read in and plot grid file for Chicago
grid_file = gpd.read_file('./data/raw/public/GridFile/Chicago_Grid.shp')
grid_file.plot(figsize=(8,8))
# ### Load the Road Network
#
# If `Chicago_Network_Buffer.graphml` does not already exist, this cell will query the road network from OpenStreetMap.
#
# Each of the road network code blocks may take a few mintues to run.
# %%time
# Read in Chicago street network (pull from OSMNX drive if it doesn't already exist)
if not os.path.exists("data/raw/private/Chicago_Network_Buffer.graphml"):
print("Loading Chicago road network from OpenStreetMap. Please wait...", flush=True)
G = ox.graph_from_place('Chicago', network_type='drive', buffer_dist=24140.2) # pulling the drive network the first time will take a while
print("Saving Chicago road network to raw/private/Chicago_Network_Buffer.graphml. Please wait...", flush=True)
ox.save_graphml(G, 'raw/private/Chicago_Network_Buffer.graphml')
print("Data saved.")
else:
print("Loading Chicago road network from raw/private/Chicago_Network_Buffer.graphml. Please wait...", flush=True)
G = ox.load_graphml('raw/private/Chicago_Network_Buffer.graphml', node_type=str)
print("Data loaded.")
# ### Plot the Road Network
# %%time
ox.plot_graph(G)
# #### Check speed limit values
#
# Display all the unique speed limit values and count how many network edges (road segments) have each value.
# We will compare this to our cleaned network later.
# +
# %%time
# Turn nodes and edges into geodataframes
nodes, edges = ox.graph_to_gdfs(G, nodes=True, edges=True)
# Get unique counts of road segments for each speed limit
print(edges['maxspeed'].value_counts())
print(len(edges))
# -
# ### network_setting function
#
# Cleans the OSMNX network to work better with drive-time analysis.
#
# First, we remove all nodes with 0 outdegree because any hospital assigned to such a node would be unreachable from everywhere. Next, we remove small (under 10 node) *strongly connected components* to reduce erroneously small ego-centric networks. Lastly, we ensure that the max speed is set and in the correct units before calculating time.
#
# Args:
#
# * network: OSMNX network for the spatial extent of interest
#
# Returns:
#
# * OSMNX network: cleaned OSMNX network for the spatial extent
def network_setting(network):
_nodes_removed = len([n for (n, deg) in network.out_degree() if deg ==0])
network.remove_nodes_from([n for (n, deg) in network.out_degree() if deg ==0])
for component in list(nx.strongly_connected_components(network)):
if len(component)<10:
for node in component:
_nodes_removed+=1
network.remove_node(node)
for u, v, k, data in tqdm(G.edges(data=True, keys=True),position=0):
if 'maxspeed' in data.keys():
speed_type = type(data['maxspeed'])
if (speed_type==str):
# Add in try/except blocks to catch maxspeed formats that don't fit Kang et al's cases
try:
if len(data['maxspeed'].split(','))==2:
data['maxspeed_fix']=float(data['maxspeed'].split(',')[0])
elif data['maxspeed']=='signals':
data['maxspeed_fix']=30.0 # drive speed setting as 35 miles
else:
data['maxspeed_fix']=float(data['maxspeed'].split()[0])
except:
data['maxspeed_fix']=30.0 #miles
else:
try:
data['maxspeed_fix']=float(data['maxspeed'][0].split()[0])
except:
data['maxspeed_fix']=30.0 #miles
else:
data['maxspeed_fix']=30.0 #miles
data['maxspeed_meters'] = data['maxspeed_fix']*26.8223 # convert mile to meter
data['time'] = float(data['length'])/ data['maxspeed_meters']
print("Removed {} nodes ({:2.4f}%) from the OSMNX network".format(_nodes_removed, _nodes_removed/float(network.number_of_nodes())))
print("Number of nodes: {}".format(network.number_of_nodes()))
print("Number of edges: {}".format(network.number_of_edges()))
return(network)
# ### Preprocess the Network using network_setting
# %%time
# G, hospitals, grid_file, pop_data = file_import (population_dropdown.value, place_dropdown.value)
G = network_setting(G)
# Create point geometries for each node in the graph, to make constructing catchment area polygons easier
for node, data in G.nodes(data=True):
data['geometry']=Point(data['x'], data['y'])
# Modify code to react to processor dropdown (got rid of file_import function)
# #### Re-check speed limit values
#
# Display all the unique speed limit values and count how many network edges (road segments) have each value.
# Compare to the previous results.
# +
# %%time
## Get unique counts for each road network
# Turn nodes and edges in geodataframes
nodes, edges = ox.graph_to_gdfs(G, nodes=True, edges=True)
# Count
print(edges['maxspeed_fix'].value_counts())
print(len(edges))
# -
# ## "Helper" Functions
#
# The functions below are needed for our analysis later, let's take a look!
#
# ### hospital_setting
#
# Finds the nearest network node for each hospital.
#
# Args:
#
# * hospital: GeoDataFrame of hospitals
# * G: OSMNX network
#
# Returns:
#
# * GeoDataFrame of hospitals with info on nearest network node
def hospital_setting(hospitals, G):
# Create an empty column
hospitals['nearest_osm']=None
# Append the neaerest osm column with each hospitals neaerest osm node
for i in tqdm(hospitals.index, desc="Find the nearest network node from hospitals", position=0):
hospitals['nearest_osm'][i] = ox.get_nearest_node(G, [hospitals['Y'][i], hospitals['X'][i]], method='euclidean') # find the nearest node from hospital location
print ('hospital setting is done')
return(hospitals)
# ### pop_centroid
#
# Converts geodata to centroids
#
# Args:
#
# * pop_data: a GeodataFrame
# * pop_type: a string, either "pop" for general population or "covid" for COVID-19 case data
#
# Returns:
#
# * GeoDataFrame of centroids with population data
def pop_centroid (pop_data, pop_type):
pop_data = pop_data.to_crs({'init': 'epsg:4326'})
# If pop is selected in dropdown, select at risk pop where population is greater than 0
if pop_type =="pop":
pop_data=pop_data[pop_data['OverFifty']>=0]
# If covid is selected in dropdown, select where covid cases are greater than 0
if pop_type =="covid":
pop_data=pop_data[pop_data['cases']>=0]
pop_cent = pop_data.centroid # it make the polygon to the point without any other information
# Convert to gdf
pop_centroid = gpd.GeoDataFrame()
i = 0
for point in tqdm(pop_cent, desc='Pop Centroid File Setting', position=0):
if pop_type== "pop":
pop = pop_data.iloc[i]['OverFifty']
code = pop_data.iloc[i]['GEOID']
if pop_type =="covid":
pop = pop_data.iloc[i]['cases']
code = pop_data.iloc[i].ZCTA5CE10
pop_centroid = pop_centroid.append({'code':code,'pop': pop,'geometry': point}, ignore_index=True)
i = i+1
return(pop_centroid)
# ### djikstra_cca_polygons
#
# Function written by <NAME> + <NAME>. It is a more efficient way to calculate distance-weighted catchment areas for each hospital. The algorithm runs quicker than the original one ("calculate_catchment_area"). It first creaets a dictionary (with a node and its corresponding drive time from the hospital) of all nodes within a 30 minute drive time (using single_cource_dijkstra_path_length function). From here, two more dictionaries are constructed by querying the original one. From this dictionaries, single part convex hulls are created for each drive time interval and appended into a single list (one list with 3 polygon geometries). Within the list, the polygons are differenced from each other to produce three catchment areas.
#
# Args:
# * G: cleaned network graph *with node point geometries attached*
# * nearest_osm: A unique nearest node ID calculated for a single hospital
# * distances: 3 distances (in drive time) to calculate catchment areas from
# * distance_unit: unit to calculate (time)
#
# Returns:
# * A list of 3 diffrenced (not-overlapping) catchment area polygons (10 min poly, 20 min poly, 30 min poly)
def dijkstra_cca_polygons(G, nearest_osm, distances, distance_unit = "time"):
'''
Before running: must assign point geometries to street nodes
# create point geometries for the entire graph
for node, data in G.nodes(data=True):
data['geometry']=Point(data['x'], data['y'])
'''
## CREATE DICTIONARIES
# create dictionary of nearest nodes
nearest_nodes_30 = nx.single_source_dijkstra_path_length(G, nearest_osm, distances[2], distance_unit) # creating the largest graph from which 10 and 20 minute drive times can be extracted from
# extract values within 20 and 10 (respectively) minutes drive times
nearest_nodes_20 = dict()
nearest_nodes_10 = dict()
for key, value in nearest_nodes_30.items():
if value <= 20:
nearest_nodes_20[key] = value
if value <= 10:
nearest_nodes_10[key] = value
## CREATE POLYGONS FOR 3 DISTANCE CATEGORIES (10 min, 20 min, 30 min)
# 30 MIN
# If the graph already has a geometry attribute with point data,
# this line will create a GeoPandas GeoDataFrame from the nearest_nodes_30 dictionary
points_30 = gpd.GeoDataFrame(gpd.GeoSeries(nx.get_node_attributes(G.subgraph(nearest_nodes_30), 'geometry')))
# This line converts the nearest_nodes_30 dictionary into a Pandas data frame and joins it to points
# left_index=True and right_index=True are options for merge() to join on the index values
points_30 = points_30.merge(pd.Series(nearest_nodes_30).to_frame(), left_index=True, right_index=True)
# Re-name the columns and set the geodataframe geometry to the geometry column
points_30 = points_30.rename(columns={'0_x':'geometry','0_y':'z'}).set_geometry('geometry')
# Create a convex hull polygon from the points
polygon_30 = gpd.GeoDataFrame(gpd.GeoSeries(points_30.unary_union.convex_hull))
polygon_30 = polygon_30.rename(columns={0:'geometry'}).set_geometry('geometry')
# 20 MIN
# Select nodes less than or equal to 20
points_20 = points_30.query("z <= 20")
# Create a convex hull polygon from the points
polygon_20 = gpd.GeoDataFrame(gpd.GeoSeries(points_20.unary_union.convex_hull))
polygon_20 = polygon_20.rename(columns={0:'geometry'}).set_geometry('geometry')
# 10 MIN
# Select nodes less than or equal to 10
points_10 = points_30.query("z <= 10")
# Create a convex hull polygon from the points
polygon_10 = gpd.GeoDataFrame(gpd.GeoSeries(points_10.unary_union.convex_hull))
polygon_10 = polygon_10.rename(columns={0:'geometry'}).set_geometry('geometry')
# Create empty list and append polygons
polygons = []
# Append
polygons.append(polygon_10)
polygons.append(polygon_20)
polygons.append(polygon_30)
# Clip the overlapping distance ploygons (create two donuts + hole)
for i in reversed(range(1, len(distances))):
polygons[i] = gpd.overlay(polygons[i], polygons[i-1], how="difference")
return polygons
# ### hospital_measure_acc (adjusted to incorporate dijkstra_cca_polygons)
#
# Measures the effect of a single hospital on the surrounding area. (Uses `dijkstra_cca_polygons`)
#
# Args:
#
# * \_thread\_id: int used to keep track of which thread this is
# * hospital: Geopandas dataframe with information on a hospital
# * pop_data: Geopandas dataframe with population data
# * distances: Distances in time to calculate accessibility for
# * weights: how to weight the different travel distances
#
# Returns:
#
# * Tuple containing:
# * Int (\_thread\_id)
# * GeoDataFrame of catchment areas with key stats
def hospital_measure_acc (_thread_id, hospital, pop_data, distances, weights):
# Create polygons
polygons = dijkstra_cca_polygons(G, hospital['nearest_osm'], distances)
# Calculate accessibility measurements
num_pops = []
for j in pop_data.index:
point = pop_data['geometry'][j]
# Multiply polygons by weights
for k in range(len(polygons)):
if len(polygons[k]) > 0: # To exclude the weirdo (convex hull is not polygon)
if (point.within(polygons[k].iloc[0]["geometry"])):
num_pops.append(pop_data['pop'][j]*weights[k])
total_pop = sum(num_pops)
for i in range(len(distances)):
polygons[i]['time']=distances[i]
polygons[i]['total_pop']=total_pop
polygons[i]['hospital_icu_beds'] = float(hospital['Adult ICU'])/polygons[i]['total_pop'] # proportion of # of beds over pops in 10 mins
polygons[i]['hospital_vents'] = float(hospital['Total Vent'])/polygons[i]['total_pop'] # proportion of # of beds over pops in 10 mins
polygons[i].crs = { 'init' : 'epsg:4326'}
polygons[i] = polygons[i].to_crs({'init':'epsg:32616'})
print('\rCatchment for hospital {:4.0f} complete'.format(_thread_id), end=" ", flush=True)
return(_thread_id, [ polygon.copy(deep=True) for polygon in polygons ])
# ### measure_acc_par
#
# Parallel implementation of accessibility measurement.
#
# Args:
#
# * hospitals: Geodataframe of hospitals
# * pop_data: Geodataframe containing population data
# * network: OSMNX street network
# * distances: list of distances to calculate catchments for
# * weights: list of floats to apply to different catchments
# * num\_proc: number of processors to use.
#
# Returns:
#
# * Geodataframe of catchments with accessibility statistics calculated
# +
def hospital_acc_unpacker(args):
return hospital_measure_acc(*args)
# WHERE THE RESULTS ARE POOLED AND THEN REAGGREGATED
def measure_acc_par (hospitals, pop_data, network, distances, weights, num_proc = 4):
catchments = []
for distance in distances:
catchments.append(gpd.GeoDataFrame())
pool = mp.Pool(processes = num_proc)
hospital_list = [ hospitals.iloc[i] for i in range(len(hospitals)) ]
results = pool.map(hospital_acc_unpacker, zip(range(len(hospital_list)), hospital_list, itertools.repeat(pop_data), itertools.repeat(distances), itertools.repeat(weights)))
pool.close()
results.sort()
results = [ r[1] for r in results ]
for i in range(len(results)):
for j in range(len(distances)):
catchments[j] = catchments[j].append(results[i][j], sort=False)
return catchments
# -
# ### overlap_calc
#
# Calculates and aggregates accessibility statistics for one catchment on our grid file.
#
# Args:
#
# * \_id: thread ID
# * poly: GeoDataFrame representing a catchment area
# * grid_file: a GeoDataFrame representing our grids
# * weight: the weight to applied for a given catchment
# * service_type: the service we are calculating for: ICU beds or ventilators
#
# Returns:
#
# * Tuple containing:
# * thread ID
# * Counter object (dictionary for numbers) with aggregated stats by grid ID number
# +
from collections import Counter
def overlap_calc(_id, poly, grid_file, weight, service_type):
value_dict = Counter()
if type(poly.iloc[0][service_type])!=type(None):
value = float(poly[service_type])*weight
intersect = gpd.overlay(grid_file, poly, how='intersection')
intersect['overlapped']= intersect.area
intersect['percent'] = intersect['overlapped']/intersect['area']
intersect=intersect[intersect['percent']>=0.5]
intersect_region = intersect['id']
for intersect_id in intersect_region:
try:
value_dict[intersect_id] +=value
except:
value_dict[intersect_id] = value
return(_id, value_dict)
def overlap_calc_unpacker(args):
return overlap_calc(*args)
# -
# ### overlapping_function
#
# Calculates how all catchment areas overlap with and affect the accessibility of each grid in our grid file.
#
# Args:
#
# * grid_file: GeoDataFrame of our grid
# * catchments: GeoDataFrame of our catchments
# * service_type: the kind of care being provided (ICU beds vs. ventilators)
# * weights: the weight to apply to each service type
# * num\_proc: the number of processors
#
# Returns:
#
# * Geodataframe - grid\_file with calculated stats
def overlapping_function (grid_file, catchments, service_type, weights, num_proc = 4):
grid_file[service_type]=0
pool = mp.Pool(processes = num_proc)
acc_list = []
for i in range(len(catchments)):
acc_list.extend([ catchments[i][j:j+1] for j in range(len(catchments[i])) ])
acc_weights = []
for i in range(len(catchments)):
acc_weights.extend( [weights[i]]*len(catchments[i]) )
results = pool.map(overlap_calc_unpacker, zip(range(len(acc_list)), acc_list, itertools.repeat(grid_file), acc_weights, itertools.repeat(service_type)))
pool.close()
results.sort()
results = [ r[1] for r in results ]
service_values = results[0]
for result in results[1:]:
service_values+=result
for intersect_id, value in service_values.items():
grid_file.loc[grid_file['id']==intersect_id, service_type] += value
return(grid_file)
# ### normalization
#
# Normalizes our result (Geodataframe) for a given resource (res).
def normalization (result, res):
result[res]=(result[res]-min(result[res]))/(max(result[res])-min(result[res]))
return result
# ### file_import
#
# Imports all files we need to run our code and pulls the Illinois network from OSMNX if it is not present (will take a while).
#
# **NOTE:** even if we calculate accessibility for just Chicago, we want to use the Illinois network (or at least we should not use the Chicago network) because using the Chicago network will result in hospitals near but outside of Chicago having an infinite distance (unreachable because roads do not extend past Chicago).
#
# Args:
#
# * pop_type: population type, either "pop" for general population or "covid" for COVID-19 cases
# * region: the region to use for our hospital and grid file ("Chicago" or "Illinois")
#
# Returns:
#
# * G: OSMNX network
# * hospitals: Geodataframe of hospitals
# * grid_file: Geodataframe of grids
# * pop_data: Geodataframe of population
def output_map(output_grid, base_map, hospitals, resource):
ax=output_grid.plot(column=resource, cmap='PuBuGn',figsize=(18,12), legend=True, zorder=1)
# Next two lines set bounds for our x- and y-axes because it looks like there's a weird
# Point at the bottom left of the map that's messing up our frame (Maja)
ax.set_xlim([314000, 370000])
ax.set_ylim([540000, 616000])
base_map.plot(ax=ax, facecolor="none", edgecolor='gray', lw=0.1)
hospitals.plot(ax=ax, markersize=10, zorder=1, c='blue')
# ### Run the model
#
# Below you can customize the input of the model:
#
# * Processor - the number of processors to use
# * Region - the spatial extent of the measure
# * Population - the population to calculate the measure for
# * Resource - the hospital resource of interest
# * Hospital - all hospitals or subset to check code
# +
import ipywidgets
from IPython.display import display
processor_dropdown = ipywidgets.Dropdown( options=[("1", 1), ("2", 2), ("3", 3), ("4", 4)],
value = 4, description = "Processor: ")
place_dropdown = ipywidgets.Dropdown( options=[("Chicago", "Chicago"), ("Illinois","Illinois")],
value = "Chicago", description = "Region: ")
population_dropdown = ipywidgets.Dropdown( options=[("Population at Risk", "pop"), ("COVID-19 Patients", "covid") ],
value = "pop", description = "Population: ")
resource_dropdown = ipywidgets.Dropdown( options=[("ICU Beds", "hospital_icu_beds"), ("Ventilators", "hospital_vents") ],
value = "hospital_icu_beds", description = "Resource: ")
hospital_dropdown = ipywidgets.Dropdown( options=[("All hospitals", "hospitals"), ("Subset", "hospital_subset") ],
value = "hospitals", description = "Hospital:")
display(processor_dropdown,place_dropdown,population_dropdown,resource_dropdown,hospital_dropdown)
# -
# ### Process population data
if population_dropdown.value == "pop":
pop_data = pop_centroid(atrisk_data, population_dropdown.value)
elif population_dropdown.value == "covid":
pop_data = pop_centroid(covid_data, population_dropdown.value)
distances=[10,20,30] # Distances in travel time
weights=[1.0, 0.68, 0.22] # Weights where weights[0] is applied to distances[0]
# Other weighting options representing different distance decays
# weights1, weights2, weights3 = [1.0, 0.42, 0.09], [1.0, 0.75, 0.5], [1.0, 0.5, 0.1]
# ### Process hospital data
# If you have already run this code and changed the Hospital selection, rerun the Load Hospital Data block.
# Set hospitals according to hospital dropdown
if hospital_dropdown.value == "hospital_subset":
hospitals = hospital_setting(hospitals[:1], G)
else:
hospitals = hospital_setting(hospitals, G)
resources = ["hospital_icu_beds", "hospital_vents"] # resources
# ### Visualize catchment areas for first hospital
# +
# Create point geometries for entire graph
for node, data in G.nodes(data=True):
data['geometry']=Point(data['x'], data['y'])
# Create catchment
poly = dijkstra_cca_polygons(G, hospitals['nearest_osm'][0], distances)
# Reproject polygons
for i in range(len(poly)):
poly[i].crs = { 'init' : 'epsg:4326'}
poly[i] = poly[i].to_crs({'init':'epsg:32616'})
# Reproject hospitals
# Possible to map from the hospitals data rather than creating hospital_subset?
hospital_subset = hospitals[:1].to_crs(epsg=32616)
fig, ax = plt.subplots(figsize=(12,8))
min_10 = poly[0].plot(ax=ax, color="royalblue", label="10 min drive")
min_20 = poly[1].plot(ax=ax, color="cornflowerblue", label="20 min drive")
min_30 = poly[2].plot(ax=ax, color="lightsteelblue", label="30 min drive")
hospital_subset.plot(ax=ax, color="red", legend=True, label = "hospital")
# Add legend
ax.legend()
# -
# ### Calculate hospital catchment areas
# %%time
catchments = measure_acc_par(hospitals, pop_data, G, distances, weights, num_proc=processor_dropdown.value)
# ### Calculate accessibility
# %%time
for j in range(len(catchments)):
catchments[j] = catchments[j][catchments[j][resource_dropdown.value]!=float('inf')]
result=overlapping_function(grid_file, catchments, resource_dropdown.value, weights, num_proc=processor_dropdown.value)
# %%time
result = normalization (result, resource_dropdown.value)
result.head()
# ### Results & Discussion
#
# to be written.
# ### Accessibility Map
# %%time
hospitals = hospitals.to_crs({'init': 'epsg:26971'})
result = result.to_crs({'init': 'epsg:26971'})
output_map(result, pop_data, hospitals, resource_dropdown.value)
# Classified Accessibility Outputs
# ### Conclusion
#
# to be written.
# ### References
#
# <NAME>., & <NAME>. (2009). An enhanced two-step floating catchment area (E2SFCA) method for measuring spatial accessibility to primary care physicians. Health & place, 15(4), 1100-1107.
|
procedure/code/03-COVID-19Acc-Reanalysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reading Questions
#
# **Paper**: A synthetic oscillatory network of transcriptional regulators. <NAME>, Leibler S. Nature. 2000 Jan 20;403(6767):335-8. [DOI Link](http://dx.doi.org/10.1038/35002125)
#
# 1. How was the repressilator network that Elowitz and Liebler explored and constructed? How did they monitor the behavior of the network in E. coli cells?
#
# 2. Elowitz and Liebler developed a mathematical model of the repressilator to understand the space of dynamical behaviors of the system. What sort of parameters did they consider important in their model? What sort of outcomes did their model predict?
#
# 3. Did the in vivo experiments that Elowitz and Liebler carried out agree with their modeling results? What other types of phenomena did Elowitz and Liebler observe in their experiments?
#
# 
# # Modeling the Repressilator
# Implement a model of the repressilator using the logic approximation framework we explored in class sessions 4 and 5.
# +
## define the differential equations for each gene here
# +
### define the simulations here
# +
## generate some plots here
# -
# ### Questions, Hybrid models
#
# 1. For what range of parameters do you get oscillatory behaviors vs is stable fixed points?
#
|
ode-modeling3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="I-NZXBybeUFS"
# # Importing the Required Libraries
# + id="3H2A9oApeNzU"
import cv2 # for using computer vision related functions
import numpy as np # for numerical computations on 2D image array
import pandas as pd # for dataset preparation for deep learning libraries
import matplotlib.pyplot as plt # for displaying image and plotting graph
# + id="BWQa96fBoYX-"
def gaussian_filter(img, mask_size = 5, sigma = 2):
offset = mask_size // 2
x, y = np.meshgrid(range(-offset, offset + 1), range(-offset, offset + 1))
gauss_filter = np.exp(-((x ** 2 + y ** 2) / (2 * sigma ** 2)))
gauss_filter /= gauss_filter.sum()
return cv2.filter2D(src = img, ddepth = -1, kernel = gauss_filter)
# + colab={"base_uri": "https://localhost:8080/", "height": 773} id="e-8Kn2h8eZNf" outputId="8b95f4d5-8189-4fb0-8786-31b2fc9c21f9"
img = cv2.imread("/content/drive/MyDrive/sem 8/CV/processed_shapes/shapes.png")
orig_img = img.copy()
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
threshed_img = cv2.threshold(gaussian_filter(gray, mask_size = 5, sigma = 10), 0, 1, cv2.THRESH_BINARY | cv2.THRESH_OTSU)[1]
plt.imshow(img), plt.show();
plt.imshow(gray, cmap = 'gray'), plt.show();
plt.imshow(threshed_img, cmap = 'binary_r'), plt.show();
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="42P8qPJRmtaa" outputId="599324a1-71d5-44df-99a8-b4da09404824"
edges = cv2.Canny(threshed_img, 0.2, 0.8)
plt.imshow(edges, cmap = 'gray');
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="HRzl-KKKhWRO" outputId="474e3366-0565-4c7a-8be9-3f174588f26c"
contours, hierarchy = cv2.findContours(edges.copy(), cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_NONE)
blank = np.zeros(threshed_img.shape)
cv2.drawContours(blank, contours, -1, (255,0,0), 1)
plt.imshow(blank, cmap = 'binary');
# + id="D7H3AcdvsafN"
# the different classes of our shapes
categories = ["circle", "square", "star", "triangle"]
# + id="aGnwesbBt2Le"
# # !pip install cPickle
import _pickle as cPickle
# load the gaussian model again
with open('/content/drive/MyDrive/sem 8/CV/processed_shapes/gauss-without-lda.pkl', 'rb') as fid:
clf_loaded = cPickle.load(fid)
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="l_q_wMngerP-" outputId="883dc16a-b50e-463e-f4c9-641804748467"
# obtaining the bounding box, extracting and saving the ROI (region of interest) font
font = cv2.FONT_HERSHEY_SIMPLEX
# org
org = (50, 50)
# fontScale
fontScale = 0.5
# Blue color in BGR
color = (255, 0, 0)
# Line thickness of 2 px
thickness = 2
ROI_number = 0
img = orig_img.copy()
for c in contours:
offset = 5
x,y,w,h = cv2.boundingRect(c)
x = x-offset
y = y-offset
w += 2*offset
h += 2*offset
cv2.rectangle(img, (x, y), (x + w, y + h), (36,255,12), 2)
ROI = cv2.resize(blank[y:y+h, x:x+w], (25,25), interpolation = cv2.INTER_AREA)
thres, ROI_thresh = cv2.threshold(ROI, 50, 255, cv2.THRESH_BINARY);
ROI_thresh = ROI_thresh/ROI_thresh.max()
pred = clf_loaded.predict([ROI_thresh.flatten()])
cv2.putText(img, categories[pred[0]], (x, y), font,
fontScale, color, thickness, cv2.LINE_AA)
plt.imshow(img);
|
CV_Project_Approach_2_Segmenting_Image.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### IMPORTING MODULES
# +
# Ignore the warnings
import warnings
warnings.filterwarnings('always')
warnings.filterwarnings('ignore')
# data visualisation and manipulation
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import style
import seaborn as sns
#configure
# sets matplotlib to inline and displays graphs below the corressponding cell.
% matplotlib inline
style.use('fivethirtyeight')
sns.set(style='whitegrid',color_codes=True)
#nltk
import nltk
#keras
import keras
from keras.preprocessing.text import one_hot,Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential
from keras.layers import Dense , Flatten ,Embedding,Input
from keras.models import Model
from keras.preprocessing.text import text_to_word_sequence
# -
# #### CREATING SAMPLE CORPUS OF DOCUMENTS ie TEXTS
# +
sample_text_1="one in hand is better than two in bush"
sample_text_2="bush was the president of US"
sample_text_3="India has only one president"
corp=[sample_text_1,sample_text_2,sample_text_3]
# -
# check output on each stage.
vocab_size=50
tokenizer=Tokenizer(num_words=vocab_size)
a=tokenizer.fit_on_texts(corp)
tokenizer.texts_to_sequences(corp)
# +
# the above piece of text is equiavalent to using the 'one_hot' function in Keras though the integer encodings seems to be different.
# -
encd_corp=[]
for doc in corp:
encd_corp.append(one_hot(doc,vocab_size))
print(encd_corp)
|
Keras Tokenizer practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Does it work on the KS? Let's check
# +
# General imports
import numpy as np
import torch
# DeepMoD stuff
from deepymod import DeepMoD
from deepymod.model.func_approx import NN, Siren
from deepymod.model.library import Library1D
from deepymod.model.constraint import LeastSquares
from deepymod.model.sparse_estimators import Threshold
from deepymod.training.sparsity_scheduler import TrainTestPeriodic, Periodic, TrainTest
from deepymod.data import Dataset
from deepymod.data.burgers import BurgersDelta
from deepymod.utils.logger import Logger
from deepymod.training.convergence import Convergence
from scipy.io import loadmat
from deepymod.analysis import load_tensorboard
if torch.cuda.is_available():
device = 'cuda'
else:
device = 'cpu'
# Settings for reproducibility
np.random.seed(42)
torch.manual_seed(0)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
# +
# Prepping data
data = loadmat('kuramoto_sivishinky.mat')
t = data['tt']
x = data['x']
u = data['uu']
# Normalizing data
t = (t - t.min())/(t.max()-t.min()) * 2 - 1
x = (x - x.min())/(x.max()-x.min()) * 2 - 1
x_grid, t_grid = np.meshgrid(x, t, indexing='ij')
# Limiting to non-chaotic part
lower_lim = 80
x_grid = x_grid[:, lower_lim:]
t_grid = t_grid[:, lower_lim:]
u = u[:, lower_lim:]
# # %%Making training data
X = np.concatenate((t_grid.reshape(-1, 1), x_grid.reshape(-1, 1)), axis=1)
y = u.reshape(-1, 1)
# Adding noise
noise_level = 0.05
y_noisy = y + noise_level * np.std(y, axis=0) * np.random.randn(*y.shape)
number_of_samples = 25000
# Into tensor
idx = np.random.permutation(y.shape[0])
X_train = torch.tensor(X[idx, :][:number_of_samples], dtype=torch.float32).to(device)
y_train = torch.tensor(y_noisy[idx, :][:number_of_samples], dtype=torch.float32).to(device)
# -
def train_SBL(model: DeepMoD,
data: torch.Tensor,
target: torch.Tensor,
optimizer,
extra_params,
sparsity_scheduler,
split = 0.8,
exp_ID: str = None,
log_dir: str = None,
max_iterations: int = 10000,
write_iterations: int = 25,
**convergence_kwargs) -> None:
"""Trains the DeepMoD model. This function automatically splits the data set in a train and test set.
Args:
model (DeepMoD): A DeepMoD object.
data (torch.Tensor): Tensor of shape (n_samples x (n_spatial + 1)) containing the coordinates, first column should be the time coordinate.
target (torch.Tensor): Tensor of shape (n_samples x n_features) containing the target data.
optimizer ([type]): Pytorch optimizer.
sparsity_scheduler ([type]): Decides when to update the sparsity mask.
split (float, optional): Fraction of the train set, by default 0.8.
exp_ID (str, optional): Unique ID to identify tensorboard file. Not used if log_dir is given, see pytorch documentation.
log_dir (str, optional): Directory where tensorboard file is written, by default None.
max_iterations (int, optional): [description]. Max number of epochs , by default 10000.
write_iterations (int, optional): [description]. Sets how often data is written to tensorboard and checks train loss , by default 25.
"""
logger = Logger(exp_ID, log_dir)
sparsity_scheduler.path = logger.log_dir # write checkpoint to same folder as tb output.
t, a, l = extra_params
# Splitting data, assumes data is already randomized
n_train = int(split * data.shape[0])
n_test = data.shape[0] - n_train
data_train, data_test = torch.split(data, [n_train, n_test], dim=0)
target_train, target_test = torch.split(target, [n_train, n_test], dim=0)
M = 10
N = data_train.shape[0]
threshold = torch.tensor(1e4).to(data.device)
alpha_threshold = torch.tensor(1e8).to(data.device)
# Training
convergence = Convergence(**convergence_kwargs)
for iteration in torch.arange(0, max_iterations):
# ================== Training Model ============================
prediction, time_derivs, thetas = model(data_train)
tau_ = torch.exp(t)
alpha_ = torch.min(torch.exp(a), alpha_threshold)
lambda_ = torch.min(torch.exp(l), 2 * threshold)
y = time_derivs[0]
X = thetas[0] / torch.norm(thetas[0], dim=0, keepdim=True)
p_MSE = N / 2 * (tau_ * torch.mean((prediction - target_train)**2, dim=0) - t + np.log(2*np.pi))
A = torch.diag(lambda_) + alpha_ * X.T @ X
mn = (lambda_ < threshold)[:, None] * (alpha_ * torch.inverse(A) @ X.T @ y)
E = alpha_ * torch.sum((y - X @ mn)**2) + mn.T @ torch.diag(lambda_) @ mn
p_reg = 1/2 * (E + torch.sum(torch.log(torch.diag(A)[lambda_ < threshold])) - (torch.sum(l[lambda_ < threshold]) + N * a) - N * np.log(2*np.pi))
MSE = torch.mean((prediction - target_train)**2, dim=0) # loss per output
Reg = torch.stack([torch.mean((dt - theta @ coeff_vector)**2)
for dt, theta, coeff_vector in zip(time_derivs, thetas, model.constraint_coeffs(scaled=False, sparse=True))])
loss = torch.sum(p_MSE + p_reg)
# Optimizer step
optimizer.zero_grad()
loss.backward()
optimizer.step()
if iteration % write_iterations == 0:
# ================== Validation costs ================
with torch.no_grad():
prediction_test = model.func_approx(data_test)[0]
MSE_test = torch.mean((prediction_test - target_test)**2, dim=0) # loss per output
# ====================== Logging =======================
_ = model.sparse_estimator(thetas, time_derivs) # calculating estimator coeffs but not setting mask
logger(iteration,
loss, MSE, Reg,
model.constraint_coeffs(sparse=True, scaled=True),
model.constraint_coeffs(sparse=True, scaled=False),
model.estimator_coeffs(),
MSE_test=MSE_test,
p_MSE = p_MSE,
p_reg = p_reg,
tau = tau_,
alpha=alpha_,
lambda_=lambda_,
mn=mn)
# ================== Sparsity update =============
# Updating sparsity
update_sparsity = sparsity_scheduler(iteration, torch.sum(MSE_test), model, optimizer)
if update_sparsity:
model.constraint.sparsity_masks = model.sparse_estimator(thetas, time_derivs)
# ================= Checking convergence
l1_norm = torch.sum(torch.abs(torch.cat(model.constraint_coeffs(sparse=True, scaled=True), dim=1)))
converged = convergence(iteration, l1_norm)
if converged:
break
logger.close(model)
t = torch.nn.Parameter(-torch.log(torch.var(y_train)).to(device))
a = torch.nn.Parameter(-torch.log(torch.var(y_train)).to(device))
l = torch.nn.Parameter(torch.zeros(12).to(device))
# +
# # %%Configuring model
network = Siren(2, [50, 50, 50, 50, 50, 50, 50, 50], 1) # Function approximator
library = Library1D(poly_order=1, diff_order=4) # Library function
estimator = Threshold(0.1)#PDEFIND(lam=1e-6, dtol=0.1) # Sparse estimator
constraint = LeastSquares() # How to constrain
model = DeepMoD(network, library, estimator, constraint).to(device) # Putting it all in the model
# %% Setting schedulers
sparsity_scheduler = TrainTestPeriodic(patience=8, delta=1e-5)#Periodic(initial_epoch=10000, periodicity=100) # Defining when to apply sparsity
optimizer = torch.optim.Adam([{'params':model.parameters(), 'betas':(0.999, 0.999), 'amsgrad':True, 'lr':0.00025}, {'params':[t, a, l], 'betas':(0.999, 0.999), 'amsgrad':True, 'lr':0.00025}]) # Defining optimizer
# -
train_SBL(model, X_train, y_train, optimizer, [t, a, l], sparsity_scheduler, exp_ID='KS', split=0.8, write_iterations=50, max_iterations=50000, delta=0.0, patience=200)
|
notebooks/Bayes/.ipynb_checkpoints/KS-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Selecting chains, residues, and atoms
# This tutorial shows how to select parts of a structure and apply different styles. It also shows how to add labels and how to zoom in on a selection.
import py3Dmol
# ## Instantiate viewer with a Hemoglobin structure
# This example also shows how to set the size of the viewer.
viewer = py3Dmol.view(query='pdb:5WOG', width=400, height=400)
viewer.show()
# ## Apply a style to a chain
# Here we select chain A and apply a cartoon style.
viewer.setStyle({'chain':'A'},{'cartoon': {'color': 'orange'}})
viewer.show()
# ## Apply a style to a list of chains and add labels
# Use a list to select to multiple chains.
# +
viewer.setStyle({'chain':['A','B']},{'cartoon': {'color': 'orange'}}) # alpha subunits of hemoglobin
viewer.addLabel('alpha subunits', {'fontColor':'orange', 'backgroundColor':'lightgray'},
{'chain': ['A','B']})
viewer.setStyle({'chain':['C','D']},{'cartoon': {'color': 'blue'}}) # beta subunits of hemoglobin
viewer.addLabel('beta subunits', {'fontColor':'blue', 'backgroundColor':'lightgray'},
{'chain': ['C','D']})
viewer.show()
# -
# # Apply style to specific residues by residue name (resn)
# Here we display all water molecules as spheres.
viewer.setStyle({'resn': 'HOH'}, {'sphere':{'radius':0.5}})
viewer.show()
# And turn waters off again by resetting their style.
viewer.setStyle({'resn': 'HOH'}, {})
viewer.show()
# ## Apply a style by residue name (resn)
# Show heme as spheres.
viewer.setStyle({'resn': 'HEM'},{'sphere': {'colorscheme': 'greenCarbon'}})
viewer.show()
# ## Apply a style to a specific residue by residue name (resn) on a specific chain
# Here we select the heme molecule in chain A and color its carbons red. We also adjust the stick radius. Finally, we zoom into a specific heme residue in chain A.
viewer.setStyle({'chain': 'A', 'resn': 'HEM'},{'stick': {'colorscheme': 'redCarbon', 'radius': 0.2}})
viewer.zoomTo({'chain': 'A', 'resn': 'HEM'})
viewer.show()
# # Apply style to a specific chain and residue by residue index (resi)
# In the following example we display the HIS-87 side chain that interacts with iron in heme.
viewer.setStyle({'chain': 'A', 'resi': '87'},{'stick': {'colorscheme': 'redCarbon'}})
viewer.show()
# # Apply style to an element (elem)
# We display the iron atom as a green sphere.
viewer.setStyle({'chain': 'A', 'elem': 'Fe'}, {'sphere': {'color': 'green'}})
viewer.show()
# # Add residue labels
# Finally, we add labels to HIS-87 and HEM.
viewer.addResLabels({'chain': 'A', 'resi': '87'})
viewer.addResLabels({'chain': 'A', 'resn': 'HEM'})
viewer.show()
# # Select binding site residues by proximity
# Here we select all residues within 5 A of the HEM in chain A.
# +
# reset styles and remove labels
viewer.removeAllLabels()
viewer.setStyle({'line':{}})
# select by distance
selection = {'resn':'HEM', 'chain':'A', 'byres':'true', 'expand': 5}
# set styles
viewer.setStyle(selection,{'stick':{'colorscheme':'orangeCarbon'}})
viewer.setStyle({'chain': 'A', 'resn': 'HEM'},{'sphere': {'colorscheme': 'lightgreenCarbon'}})
viewer.zoomTo(selection)
viewer.show()
# -
|
1-3D-visualization/3-Selection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
#
# # Template Matching
#
#
# We use template matching to identify the occurrence of an image patch
# (in this case, a sub-image centered on a single coin). Here, we
# return a single match (the exact same coin), so the maximum value in the
# ``match_template`` result corresponds to the coin location. The other coins
# look similar, and thus have local maxima; if you expect multiple matches, you
# should use a proper peak-finding function.
#
# The ``match_template`` function uses fast, normalized cross-correlation [1]_
# to find instances of the template in the image. Note that the peaks in the
# output of ``match_template`` correspond to the origin (i.e. top-left corner) of
# the template.
#
# .. [1] <NAME>, "Fast Normalized Cross-Correlation", Industrial Light and
# Magic.
#
#
#
# +
import numpy as np
import matplotlib.pyplot as plt
from skimage import data
from skimage.feature import match_template
from skimage.io import imread, imsave
from skimage.transform import rotate, rescale
from skimage.draw import polygon_perimeter
from glob import glob
import mpld3
mpld3.enable_notebook()
# -
def rotate_good(img, angle):
return 1 - rotate(1 - img, angle)
triangles = [imread(file_name, as_grey=True) for file_name in reversed(['trojkat-maly.png', 'trojkat.png', 'trojkat-duzy.png'])]
triangles_straight = [rotate_good(triangle, angle) for angle in [-90, 0, 90] for triangle in triangles] + [imread('hak.png', as_grey=True), imread('hak-maly.png', as_grey=True)]
triangles_skewed = [rotate_good(triangle, angle) for angle in [-45, 45] for triangle in triangles]
triangles = triangles_straight + triangles_skewed
def match(image):
def match_patterns(patterns):
THRESHOLD = 0.7
best_score = THRESHOLD
best_tid = None
best_x = 0
best_y = 0
best_result = None
for (tid, triangle) in enumerate(patterns):
result = match_template(image, triangle)
ij = np.unravel_index(np.argmax(result), result.shape)
x, y = ij[::-1]
score = result[y, x]
if score > best_score:
best_score = score
best_tid = tid
best_x = x
best_y = y
best_result = result
return (best_result, best_tid, best_x, best_y)
(best_result, best_tid, best_x, best_y) = match_patterns(triangles_straight)
if best_tid is None:
(best_result, best_tid, best_x, best_y) = match_patterns(triangles_skewed)
if best_tid is not None:
best_tid += len(triangles_straight)
if best_tid is not None:
new_image = image.copy()
triangle = triangles[best_tid]
for i in xrange(triangle.shape[0]):
for j in xrange(triangle.shape[1]):
new_image[best_y + i, best_x + j] = 1 - (1 - image[best_y + i, best_x + j]) * triangle[i, j]
return (new_image, best_result, best_tid, best_x, best_y)
else:
return None
def do_stuff(image):
infos = []
while True:
res = match(image)
if res is None:
break
(image, result, tid, x, y) = res
infos.append((tid, x, y))
#ax2.set_axis_off()
#plt.set_title('image')
# highlight matched region
image2 = image.copy()
for (tid, x, y) in infos:
h, w = triangles[tid].shape
(rr, cc) = polygon_perimeter([y, y + h, y + h, y], [x, x, x + w, x + w])
image2[rr, cc] = 0
#rect = plt.Rectangle((x, y), wcoin, hcoin, edgecolor='r', facecolor='none')
#ax2.add_patch(rect)
return image2
print glob('pics/*.png')
for file_name in glob('pics/*.png'):
print 'dupa1'
image = imread(file_name, as_grey=True)
print 'dupa2'
image2 = do_stuff(image)
print 'dupa3'
#print fig
#print file_name
#file_name = '.'.join(file_name.split('.')[:-1]) + '_plot.png'
#print file_name
#imsave(file_name, image2)
#print 'dupa4'
# +
(image, result, tid, x, y) = match(image)
triangle = triangles[tid]
print tid
fig = plt.figure(figsize=(8, 3))
ax1 = plt.subplot(1, 3, 1)
ax2 = plt.subplot(1, 3, 2, adjustable='box-forced')
ax3 = plt.subplot(1, 3, 3, sharex=ax2, sharey=ax2, adjustable='box-forced')
ax1.imshow(triangle, cmap=plt.cm.gray)
ax1.set_axis_off()
ax1.set_title('template')
ax2.imshow(image, cmap=plt.cm.gray)
ax2.set_axis_off()
ax2.set_title('image')
# highlight matched region
hcoin, wcoin = triangles[tid].shape
rect = plt.Rectangle((x, y), wcoin, hcoin, edgecolor='r', facecolor='none')
ax2.add_patch(rect)
ax3.imshow(result)
ax3.set_axis_off()
ax3.set_title('`match_template`\nresult')
# highlight matched region
ax3.plot(x, y, 'o', markeredgecolor='r', markerfacecolor='none', markersize=10)
plt.show()
# -
|
plot_template.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LSDDA: Hackathon #1
#
# ## Objective
# **Predict if a web page is fake (binary classification).**
#
# ## Data structure
# ### Target variable:
# **label**: label=0 webpage not fake | label=1 webpage is fake
# ### Features:
# - 24 features: categoricals and numericals:
#
# **Feature Name | Description**
# urlid | unique identifier for each url
# category1 | info category
# category1_score | info score
# avglinksize | Average number of words in each link
# commonlink_1 | 1 link sharing ratio
# commonlink_2 | 2 link sharing ratio
# commonlink_3 | 3 link sharing ratio
# commonlink_4 | 4 link sharing ratio
# compression_r | zip compression
# embed_r | embedded count
# frame1 | A page is frame-based (1)
# frame2 | Ratio of iframe
# hasDL | Contains url with domain
# html_r | html ratio
# image_r | image ratio
# not_news | not news label
# lengthyLD | alphanumeric characters
# linkws | Percentage on hyperlink
# news_fp | If front-page news
# non_markup_a | Page's text's number of alphanumeric characters
# numberL | Markups
# numwordsU | Words in url
# parametrizedLR | url contains parameters or has an attached event
# spelling_r | Ratio on spelling mistake
#
# ## Evaluation
# The classification metric that shall be used is [AUROC](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_auc_score.html).
import pandas as pd
import numpy as np
# # Load Train Data
# The test data is loaded the same way.
df = pd.read_csv('train.csv')
print('Shape:', df.shape)
df.head()
# # Have fun!
from IPython.display import HTML
HTML('<img src="https://media.giphy.com/media/o0vwzuFwCGAFO/giphy.gif">')
|
hackathon-2017-Binary-Classification/Hackathon #1 - README.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 02 Huia Experience Training
# # Setup
# ## Install Tensorflow 2 Nightly and other Libraries
# +
# #!pip install opencv-python
# #!pip install scipy
# #!pip install sklearn
# #!pip install pathlib
# #!pip install matplotlib
# #!pip install fastai=1.0.52
# #!conda install cudatoolkit=10.0
# #!pip install scikit-learn
# Tensorflow 2 Alpha
# #!pip install tensorflow-gpu==2.0.0-alpha0
# Tensorflow 2 Nightly
# #!pip install tf-nightly-gpu-2.0-preview --upgrade --force-reinstall
#tf addons only works with alpha for now, don't use
# #!pip install tensorflow-addons
#list versions of tensorflow related files
# !pip freeze | egrep 'tensor|tb|tf|numpy'
# clear tensorboard logs
# !rm -rf ./logs/*
# -
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
import tensorflow as tf
from tensorflow import keras
#import tensorflowjs as tfjs
import pathlib
import os
import random
#import tensorflow_addons as tfa # not used for now as it is incompatible with tf.data.Dataset
# enable logging to make sure we are running on the GPU
tf.debugging.set_log_device_placement(True)
# check tensorflow version
tf.__version__
# +
# clear any active session
tf.keras.backend.clear_session()
AUTOTUNE = tf.data.experimental.AUTOTUNE
BATCH_SIZE = 48
# -
# # Data
# We have to understand well our data as this is fundamental to achieve good results.
# +
root_path = pathlib.Path("./training_data/")
image_path = root_path /"images"
json_path = root_path /"json"
augmented_path = root_path/"augmented_imgs"
image_path,json_path ,augmented_path
# +
# lets set the random seed so we can reproduce our results
random.seed(7)
# get filenames from directories
all_image_paths = list(image_path.glob('*.png')) + list(augmented_path.glob('*.png'))
#all_json_paths = list(json_path.glob('*.json'))
all_image_paths = [str(path) for path in all_image_paths] # convert to strings
random.shuffle(all_image_paths) # randomize
# -
len(all_image_paths)#, len(all_json_paths)
all_image_paths[:20]
import re
# extract categories for classification
pat = r'/([^/]+)_\d+.png$'
all_image_labels = [str(re.search(pat,str(image)).group(1)).lower() for image in all_image_paths]
len(all_image_labels)
# +
import IPython.display as display
import matplotlib.pyplot as plt
# lets check samples of our images to see what they look like
for n in range(3):
image = random.choice(all_image_paths)
display.display(display.Image(str(image)))
print(f"file: {image}")
# -
# get unique classes
huia_person=[]
for label in all_image_labels:
if label not in huia_person:
huia_person.append(label)
huia_person = sorted(huia_person) # sort label list
huia_person
# put them in a dict for lookup
label_to_index = dict((name, index) for index,name in enumerate(huia_person))
label_to_index
# lets format it, so we can copy and paste the dict direclty into javascript :-)
print("POSE_CLASSES = {")
for index,name in enumerate(huia_person):
print("\t" +str(index)+": '"+name+"',")
print("}")
# +
# load data into tf data
img_raw = tf.io.read_file(all_image_paths[0])
img_tensor = tf.image.decode_image(img_raw)
print(img_tensor.shape)
print(img_tensor.dtype)
plt.imshow(img_tensor)
#img_raw.numpy()
# +
import numpy
from random import randint
import math
<EMAIL>
def preprocess_image(image):
#print("EagerMode:" + str(tf.executing_eagerly()))
# decode PNG
image = tf.image.decode_png(image, channels=3)
# data augmentation - doing statically via fastai, as tensorflow addons is not compatible with graph mode yet
# rotate random
#degrees = random.randint(-6,6)
#angle = degrees * math.pi / 180
#image = tfa.image.rotate(image,angle,interpolation='BILINEAR')
# comented as this is not supported in graph mode yet
# random crop - images are tensors of shape (500,640,3)
#crop_factor = (random.randint(0,20)/100) # generate numbers between 0.7 and 1.0
#new_width = int(640 * (1-crop_factor))
#new_height = int(500 * (1-crop_factor))
# resize_image_with_crop_or_pad
# image = tf.image.resize_image_with_crop_or_pad(image,new_height,new_width)
# commented as we are doing prepocessing on static files
# resize
image = tf.image.resize(image, [224, 224])
# normalize = convert to [-1:1]
offset = 127.5
image = (image-offset)/offset
return image
<EMAIL>
def load_and_preprocess_image(path):
image = tf.io.read_file(path)
return preprocess_image(image)
# -
# load data into tf data
img_tensor = load_and_preprocess_image(all_image_paths[0])
plt.imshow(img_tensor)
# +
path_ds = tf.data.Dataset.from_tensor_slices(all_image_paths)
path_ds
next(iter(path_ds))
# -
# force non performatic eager mode as tfa.image doesn't support graph mode yet
#image_ds = path_ds.map(lambda path: tf.py_function(func=load_and_preprocess_image,inp=[path],Tout=tf.float32))
image_ds = path_ds.map(map_func=load_and_preprocess_image,num_parallel_calls = AUTOTUNE)#.cache(filename='images_normalized')
all_image_labels_idx = [label_to_index[label] for label in all_image_labels]
label_ds = tf.data.Dataset.from_tensor_slices(tf.cast(all_image_labels_idx, tf.int64))
len(all_image_labels)
# +
image_label_ds = tf.data.Dataset.zip((image_ds, label_ds))
image_label_ds
image_count = len(all_image_labels)
type(image_label_ds)
# -
ds = image_label_ds #.cache()
ds = ds.apply(tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds = ds.batch(BATCH_SIZE).prefetch(buffer_size=AUTOTUNE)
ds
# we will use a pretrained mobilenet for transferlearning
mobilenet = tf.keras.applications.MobileNetV2(input_shape=(224,224,3),include_top=False,weights='imagenet')
mobilenet.trainable = False
mobilenet.summary()
image_batch, label_batch = next(iter(ds))
print(image_batch.shape)
label_batch[0].numpy()
image_batch[0].numpy()
# +
image_batch, label_batch = next(iter(ds))
feature_map_batch = mobilenet(image_batch)
print(feature_map_batch.shape)
# -
model = tf.keras.Sequential([
mobilenet,
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Dense(1024,activation='relu',bias_initializer=tf.keras.initializers.he_normal()),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Dense(1024,activation='relu',bias_initializer=tf.keras.initializers.he_normal()),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(512,activation='relu',bias_initializer=tf.keras.initializers.he_normal(),name='features'),
tf.keras.layers.Dense(len(huia_person),activation='softmax'),
])
model.summary()
# +
logit_batch = model(image_batch).numpy()
print("min logit:", logit_batch.min())
print("max logit:", logit_batch.max())
print()
print("Shape:", logit_batch.shape)
# -
len(model.trainable_variables)
model.summary()
steps_per_epoch=tf.math.ceil((len(all_image_paths)-838)/BATCH_SIZE).numpy()
steps_per_epoch
# +
# One Cycle https://www.kaggle.com/robotdreams/one-cycle-policy-with-keras
# import OneCycleLR
from datetime import datetime
now = datetime.now()
log_dir = "./logs/step_1_" + now.strftime("%Y%m%d-%H%M%S") + "/"
callbacks = []
# logging
tbCallback = tf.keras.callbacks.TensorBoard(log_dir=log_dir,histogram_freq=1)
tbCallback.set_model(model)
callbacks.append(tbCallback)
epochs = 20
max_lr = 3e-4
base_lr = max_lr/10
# max_m = 0.98
# base_m = 0.85
# cyclical_momentum = False
# augment = True
# cycles = 2.35
# iterations = round(len(all_image_paths)/BATCH_SIZE*epochs)
# iterations = list(range(0,iterations+1))
# step_size = len(iterations)/(cycles)
# clr = OneCycleLR.CyclicLR(base_lr=base_lr,
# max_lr=max_lr,
# step_size=step_size,
# max_m=max_m,
# base_m=base_m,
# cyclical_momentum=cyclical_momentum)
#callbacks.append(clr)
# class myCallback(tf.keras.callbacks.Callback):
# def on_epoch_end(self, epoch, logs={}):
# if(logs.get('val_accuracy')>0.9):
# print("\nReached 60% accuracy so cancelling training!")
# self.model.stop_training = True
# earlyStopping = tf.keras.callbacks.EarlyStopping(patience=2, monitor='loss')
# callbacks.append(earlyStopping)
# chkPoint = tf.keras.callbacks.ModelCheckpoint('./models.h5')
# chkPoint.set_model(model)
# callbacks.append(chkPoint)
# -
# skip 20% validation images
test_dataset = ds.take(838)
train_dataset = ds.skip(838)
mobilenet.trainable = False
model.compile(optimizer=tf.keras.optimizers.Adam(lr=max_lr, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=1e-07, amsgrad=False),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
start = datetime.now()
history = model.fit(train_dataset, epochs=epochs,steps_per_epoch=steps_per_epoch,verbose=1,validation_data=test_dataset,
callbacks=callbacks)
end = datetime.now()
print(f"total time of {end-start} for {epochs} epochs, tensorflow version={tf.__version__}")
# +
# list all data in history
print(history.history.keys())
# summarize history for accuracy
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# +
# #!pip install scikit-learn
# +
from datetime import datetime
import OneCycleLR
now = datetime.now()
log_dir = "./logs/step_2_" + now.strftime("%Y%m%d-%H%M%S") + "/"
callbacks = []
epochs = 50
max_lr = 3e-6
# base_lr = max_lr/10
# max_m = 0.98
# base_m = 0.85
# cyclical_momentum = False
# augment = True
# cycles = 2.35
# iterations = round(len(all_image_paths)/BATCH_SIZE*epochs)
# iterations = list(range(0,iterations+1))
# step_size = len(iterations)/(cycles)
# clr = OneCycleLR.CyclicLR(base_lr=base_lr,
# max_lr=max_lr,
# step_size=step_size,
# max_m=max_m,
# base_m=base_m,
# cyclical_momentum=cyclical_momentum)
#callbacks.append(clr)
# logging
tbCallback = tf.keras.callbacks.TensorBoard(log_dir=log_dir,histogram_freq=1)
tbCallback.set_model(model)
callbacks.append(tbCallback)
# -
mobilenet.trainable = True
model.compile(optimizer=tf.keras.optimizers.Adam(lr=max_lr, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=1e-07, amsgrad=False),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
history = model.fit(train_dataset, epochs=epochs,steps_per_epoch=steps_per_epoch,verbose=1,validation_data=test_dataset,
callbacks=callbacks)
# +
# list all data in history
print(history.history.keys())
# summarize history for accuracy
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.ylim(0, 1)
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.ylim(0, 1)
plt.show()
# -
model.save("models/huia_poses_final.h5") # 1.0
model
# # Convert to TENSORFLOW JS / Quantization
# !pip freeze
# #!pip install tensorflowjs==0.6.4 --force-reinstall
# +
# Tensorflow 2 Alpha has a bug exporting to TFJS, so we need to use a nightly version
import tensorflowjs as tfjs
import numpy as np
# model = tf.keras.models.load_model("models/huia_mob_224_final_one_cycle.h5")
# model.load_weights("models/huia_mob_224_final_one_cycle.h5")
#tfjs.converters.save_keras_model(model, "models_tfjs/huia_poses_final_q8ale",quantization_dtype=np.uint8)
tfjs.converters.save_keras_model(model, "models_tfjs/huia_poses_final_q16ale",quantization_dtype=np.uint16)
# run on command line, above commands are buggy at the moment
# # !tensorflowjs_converter \
# # --input_format=keras \
# # --output_format=tfjs_layers_model \
# # ./models/huia_poses_final.h5 \
# # ./models_tfjs/huia_poses_final_8b \
# # --quantization_bytes 1
# # !tensorflowjs_converter \
# # --input_format=keras \
# # --output_format=tfjs_layers_model \
# # ./models/huia_poses_final.h5 \
# # ./models_tfjs/huia_poses_final_16b \
# # --quantization_bytes 2
# # !tensorflowjs_converter \
# # --input_format=tfjs_layers_model \
# # --output_format=tfjs_layers_model \
# # ./tfjs_poses/model.json \
# # ./tfjs_poses_q1 \
# # --quantization_bytes 1
# tensorflowjs_converter \
# --input_format=tfjs_layers_model \
# --output_format=tfjs_layers_model \
# --quantization_bytes=1 \
# huia_poses_final_16b/model.json \
# huia_poses_final_16b8
# # copy tfjs model to 03_experience/static and change App.vue reference to load it
# -
model.summary()
tf.keras.utils.plot_model(model,'model.png',show_layer_names=False) #,show_shapes=True)
# # Tensorboard Visualization
# https://www.tensorflow.org/tensorboard/
import tensorboard as tb
# %load_ext tensorboard
# #%load_ext tensorboard.notebook
# %tensorboard --logdir ./logs
#notebook.list() # View open TensorBoard instances
#notebook.display(port=6006, height=1000)
# #!kill 25264
# # Test / Predict
# +
import numpy as np
sample = np.reshape(img_final,[1,224,224,3])
predict = int(model.predict_classes(sample))
predict
# -
[key for key,value in label_to_index.items() if value == predict]
label_to_index
# # UTILS
# ## Delete Images that are empty
# +
# during image capture some images are empty, so we automatically delete them
# open images and delete if they are empty
def remove_empty_imgs(imgpath):
for item in imgpath.iterdir():
im = imread(str(item), format='png')
if np.count_nonzero(im)==0:
print(item,np.count_nonzero(im))
os.remove(str(item))
#remove_empty_imgs(image_path)
# -
# ## Sync JSONs with Images
# +
# since we might delete unwanted images for training, this will also delete the json files
# sync json & images
def sync_json_images(json_paths, img_path):
for json in json_paths:
#print(json.stem, end=" ")
img = img_path/f"{json.stem}.png"
if not Path(img).exists():
print(f"{img} doesn't exist, deleting {json}")
Path(json).unlink()
#sync_json_images(all_json_paths, image_path)
# -
# # Static Data Augmentation
# Decided to use FASTAI to statically preprocess data augmentation, as tensorflow addons (0.3.1) still didn't support graph mode and is therefore not compatible with tf.data.Dataset mappings
#
#
#
# +
from fastai.vision import *
from fastai.metrics import error_rate
from random import randint
import pathlib
root_path = pathlib.Path("./training_data/")
save_fast = Path('./training_data/augmented_imgs/')
save_fast.mkdir(parents=True, exist_ok=True)
tfms = get_transforms(do_flip=False,
flip_vert=False,
max_rotate=6,
max_zoom=1.2,
max_lighting=None,
max_warp=0.2,
p_affine=0.2,
p_lighting=0)
image_path = root_path /"images"
all_image_paths = list(image_path.glob('*.png'))
def generate_augmented(qty):
for f in all_image_paths:
image = open_image(f)
for i in range(0,qty):
image_fast = image.apply_tfms(tfms[0])
save_name = str(save_fast) + '/' + f.stem + '99' + str(i) + f.suffix
print(save_name)
image_fast.save(save_name)
#generate 10 variations of each image
#generate_augmented(10)
# -
|
02_train/train_huia_poses_keras_tf2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# ## 栈
# >栈(stack),有些地方称为堆栈,是一种容器,可存入数据元素、访问元素、删除元素,
# >它的特点在于只能允许在容器的一端(称为栈顶端指标,英语:top)进行加入数据(英语:push)
# >和输出数据(英语:pop)的运算。没有了位置概念,保证任何时候可以访问、删除的元素都是
# >此前最后存入的那个元素,确定了一种默认的访问顺序。
# >由于栈数据结构只允许在一端进行操作,因而按照后进先出(LIFO, Last In First Out)的原理运作。
# >栈可以用顺序表实现,也可以用链表实现。
#
# ### 栈的结构示意图
# 
# + pycharm={"name": "#%% \u4ee3\u7801\u5b9e\u73b0\n", "is_executing": false}
class Stack(object):
"""栈"""
def __init__(self):
self.items = []
def is_empty(self):
"""判断是否为空"""
return self.size() == 0
def push(self, item):
"""加入元素"""
self.items.append(item)
def pop(self):
"""弹出元素"""
return self.items.pop()
def peek(self):
"""返回栈顶元素"""
if self.is_empty():
return None
else:
return self.items[-1]
def size(self):
"""返回栈的大小"""
return len(self.items)
# + pycharm={"name": "#%% \u6d4b\u8bd5\u6570\u636e\n", "is_executing": false}
if __name__ == "__main__":
stack = Stack()
stack.push(1)
stack.push(2)
stack.push(3)
stack.push(4)
print("栈的大小为:", stack.size())
print("栈顶元素为:", stack.peek())
print(stack.pop())
print(stack.pop())
print(stack.pop())
print(stack.pop())
print("栈的大小为:", stack.size())
print(stack.is_empty())
|
DataStructure/stack.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Python
#
# ## [Functional Programming](https://docs.python.org/3/howto/functional.html) with Python:
#
# + #### _lambda_ functions
# + #### _zip_
# + #### _map_
# + #### _filter_
# + #### _reduce_
import time
# ## lambda Functions
#
# When writing functional-style programs, you’ll often need little functions that act as predicates or that combine elements in some way. If there’s a Python built-in or a module function that’s suitable, you don’t need to define a new function at all, as in these examples:
#
# stripped_lines = [line.strip() for line in lines]
# existing_files = filter(os.path.exists, file_list)
#
# If the function you need doesn’t exist, you need to write it.
# One way to write small functions is to use the _lambda_ expression. lambda takes a number of parameters and an expression combining these parameters, and creates an anonymous function that returns the value of the expression:
#
# The pattern is:
#
# lambda < variables > : operation(< variables >)
# #### Examples
def adder(x, y):
return x + y
# +
# %%timeit
adder(3,4)
# -
adder_lambda = lambda x, y: x+y
# +
# %%timeit
adder_lambda(3,4)
# -
general_adder = lambda *x : sum(x)
general_adder(2,3,4,5)
square = lambda x: x**2
square(2)
def print_assign(name, value):
return name + '=' + str(value)
print_assign = lambda name, value: name + '=' + str(value)
print_assign('year',2020)
even = lambda x:True if x%2==0 else False
print(even(17))
print(even(16))
# #### Dictionary of functions
power = {'square': lambda x:x**2,
'cube': lambda x:x**3,
'fourth': lambda x:x**4
}
power
type(power["cube"])
# +
print(power['cube'](9))
print(power['square'](3))
print(power['fourth'](7))
# -
funcs = [lambda x:x**2, lambda x:x-1]
print(funcs)
[func(x) for func in funcs for x in [1,2,3]]
# ## Functional Tools: _zip_, _filter_, _map_ , _reduce_
# ## [_zip_](https://medium.com/techtofreedom/7-levels-of-using-the-zip-function-in-python-a4bd22ee8bcd)
#
# + #### _zip_ function returns a _zip_ object (which is an iterator) that will aggregate elements from two or more iterables.
# + #### You can use the resulting iterator to solve common tasks, like creating dictionaries.
sq1 = [1,2,3,4,5,6,7,8]
sq2 = ['a','b','c','d','e','f']
z = zip(sq1,sq2)
print(z)
next(z)
#z.__next__()
z = zip(sq1,sq2)
for t in z:
print(t)
z = zip(sq1,sq2)
list(z)
names = ['Leticia', 'Ana', 'Raquel']
grades = [8,9,10]
dic_grades = dict(zip(names,grades))
dic_grades
# #### Zip trims to the length of the smaller sequence
students = ['Diogo','Rafael','Gustavo','Deborah', 'Extra Student']
grades = [0,1,2,3]
new_dict_grades = dict(zip(students,grades))
print(new_dict_grades)
list1 = list(range(11))
list2 = list(range(1,30,2))
list3 = list(range(1,100,5))
print(list1)
print(list2)
print(list3)
zipped = list(zip(list1, list2, list3))
print(zipped)
# #### For unequal length sequences we can use zip_logest from itertools
# +
from itertools import zip_longest
id = [1, 2]
leaders = ['<NAME>', '<NAME>', '<NAME>', '<NAME>']
# -
long_record = zip_longest(id, leaders)
print(list(long_record))
long_record_2 = zip_longest(id, leaders, fillvalue='Top')
print(list(long_record_2))
# #### How to reverse a zip command?
t1 = ((1,2),(3,4),(4,5))
print(t1)
print(*t1)
print(*zipped)
unzipped = (zip(*zipped))
list(unzipped)
# #### Using Zip with comprehensions
sq1 = [1,2,3,4,5,6,7,8]
sq2 = ['a','b','c','d','e','f']
d3 = {x.upper():y for y,x in zip(sq1,sq2)}
d3
s1 = [x.lower() for x in d3.keys()]
s2 = [x for x in d3.values()]
print(s1)
print(s2)
d4 = {k:v for k,v in zip(s1,s2)}
d4
sq1 = [1,2,3,4,5,6,7,8]
sq2 = ['a','b','c','d','e','f']
sq3 = ['w','e','r','y']
z4 = zip(sq1,sq2,sq3)
print(list(z4))
# #### Using zip tp transpose a matrix
matrix = [[1, 2, 3], [1, 2, 3]]
matrix_T = [list(i) for i in zip(*matrix)]
print(matrix_T)
# ## _map_
#
# + #### _map_ function returns a map object (which is an iterator) of the results after applying the given function to each item of a given iterable (list, tuple etc.)
# + #### Maps are similar to [list comprehensions](https://stackoverflow.com/questions/1247486/list-comprehension-vs-map) but they create generators instead of lists
def my_function(x):
return x**10
print(my_function(4))
print(my_function(10))
seq6 = [3,7,9,1,5,7]
# +
# %%time
results = map(my_function, seq6)
print(list(results))
print(type(results))
# +
# %%time
results = [my_function(x) for x in seq6]
print(results)
print(type(results))
# +
# %%time
results = map(lambda x:x**10, seq6)
print(list(results))
print(type(results))
# -
# ## _reduce_
#
# + #### The reduce() function in Python takes in a function and a list as argument. The function is called with a lambda function and a list and a new reduced result is returned. This performs a repetitive operation over the pairs of the list. This is a part of functools module.
from functools import reduce
seq9 = [1,2,3,4,5,6,7,8,9,10]
multiply = reduce(lambda x,y:x*y, seq9)
multiply
seq10 = ['a','b','c','d','e','f','g']
concatenate = reduce(lambda x,y:x.upper()+y, seq10)
concatenate
list1 = list(range(11))
list2 = list(range(1,30,2))
list3 = list(range(1,100,5))
soma = reduce(lambda x,y:x+y**2,list1)
soma
soma2 = reduce(lambda x,y:x+y**2,list2)
soma2
soma3 = reduce(lambda x,y:x+y**2,list3)
soma3
import random
seq = [random.random() for x in range(10)]
print(seq)
max(seq)
compara = lambda x,y: x if x>=y else y
reduce(compara,seq)
# ## _filter_
#
# + #### The filter() method filters the given sequence with the help of a function that tests each element in the sequence to be true or not. This function must return a boolean value.
my_string = 'aAbRmmmTTTBfgHHrTEB'
resp = filter(lambda x:x.islower(), my_string)
print(list(resp))
print(type(resp))
resp = filter(lambda x: not x.islower(), my_string)
print(list(resp))
print(type(resp))
resp = filter(lambda x:x.isupper(), my_string)
print(list(resp))
print(type(resp))
list1 = [random.random() for x in range(10)]
bigger_than_dot4 = filter(lambda x:x>0.4,list1)
list(bigger_than_dot4)
# +
simplified_genesis = '''
In the beginning God created the heaven and the earth.
And the earth was without form, and void; and darkness
was upon the face of the deep. And the Spirit of God
moved upon the face of the waters. And God said,
Let there be light: and there was light.'''
simplified_genesis.split()[0:10]
# -
istitle = lambda x : x.istitle()
print(list(filter(istitle, simplified_genesis.split())))
print(list(filter(lambda x: x.istitle(), simplified_genesis.split())))
print(list(filter(str.istitle, simplified_genesis.split())))
#
# ## Implementing the builtin generators as ordinary functions:
# ## _zip_:
def my_zip(*sequences):
smaller = min([len(sequence) for sequence in sequences])
for i in range(smaller):
yield(tuple([sequence[i] for sequence in sequences]))
zipped = my_zip([1,2,3,5],[5,6,7],[3,2,5])
print(type(zipped))
print(list(zipped))
# ## _map_:
# +
## option 1 - create a generator
def my_map(func, sequence):
for element in (sequence):
yield(func(element))
# +
## option 2 - return a generator
def my_map(func, sequence):
mapped = (func(item) for item in sequence)
return mapped
# -
mapped = my_map(lambda x:x**2, [1,2,3,4,5])
type(mapped)
print(list(mapped))
# ## _filter_:
# +
## option 1 - create a generator
def my_filter(func_bool, sequence):
filtered = (item for item in sequence if func_bool(item))
for element in filtered:
yield(element)
# +
## option 2 - return a generator
def my_filter(func_bool, sequence):
filtered = (item for item in sequence if func_bool(item))
return filtered
# -
filtered = my_filter(lambda x:x%2==0, [1,2,3,4,5,6])
type(filtered)
print(list(filtered))
# ## _range_:
def my_range(*args):
start = 0
step = 1
if len(args) == 1:
end = args[0]
elif len(args) == 2:
start = args[0]
end = args[1]
elif len(args) == 3:
start = args[0]
end = args[1]
step = args[2]
else:
print('Too few or too many arguments')
return
while start < end:
yield start
start += step
print(list(my_range(10)))
print(list(my_range(2,10,2)))
# ## _reduce_:
def my_reduce(function, sequence):
result = sequence[0]
for i in range(len(sequence)-1):
result = function(result,sequence[i+1])
return result
my_reduce(lambda x,y:x+y, [1,2,3,4,5,6,7])
|
Notebooks/07_Functional_Programming.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GP Regression with LOVE for Fast Predictive Variances and Sampling
#
# ## Overview
#
# In this notebook, we demonstrate that LOVE (the method for fast variances and sampling introduced in this paper https://arxiv.org/abs/1803.06058) can significantly reduce the cost of computing predictive distributions. This can be especially useful in settings like small-scale Bayesian optimization, where predictions need to be made at enormous numbers of candidate points.
#
# In this notebook, we will train a KISS-GP model on the `skillcraft `UCI dataset, and then compare the time required to make predictions with each model.
#
# **NOTE**: The timing results reported in the paper compare the time required to compute (co)variances __only__. Because excluding the mean computations from the timing results requires hacking the internals of GPyTorch, the timing results presented in this notebook include the time required to compute predictive means, which are not accelerated by LOVE. Nevertheless, as we will see, LOVE achieves impressive speed-ups.
# +
import math
import torch
import gpytorch
import tqdm
from matplotlib import pyplot as plt
# Make plots inline
# %matplotlib inline
# -
# ### Loading Data
#
# For this example notebook, we'll be using the `elevators` UCI dataset used in the paper. Running the next cell downloads a copy of the dataset that has already been scaled and normalized appropriately. For this notebook, we'll simply be splitting the data using the first 40% of the data as training and the last 60% as testing.
#
# **Note**: Running the next cell will attempt to download a small dataset file to the current directory.
# +
import urllib.request
import os
from scipy.io import loadmat
from math import floor
# this is for running the notebook in our testing framework
smoke_test = ('CI' in os.environ)
if not smoke_test and not os.path.isfile('../elevators.mat'):
print('Downloading \'elevators\' UCI dataset...')
urllib.request.urlretrieve('https://drive.google.com/uc?export=download&id=1jhWL3YUHvXIaftia4qeAyDwVxo6j1alk', '../elevators.mat')
if smoke_test: # this is for running the notebook in our testing framework
X, y = torch.randn(100, 3), torch.randn(100)
else:
data = torch.Tensor(loadmat('../elevators.mat')['data'])
X = data[:, :-1]
X = X - X.min(0)[0]
X = 2 * (X / X.max(0)[0]) - 1
y = data[:, -1]
train_n = int(floor(0.8 * len(X)))
train_x = X[:train_n, :].contiguous()
train_y = y[:train_n].contiguous()
test_x = X[train_n:, :].contiguous()
test_y = y[train_n:].contiguous()
if torch.cuda.is_available():
train_x, train_y, test_x, test_y = train_x.cuda(), train_y.cuda(), test_x.cuda(), test_y.cuda()
# -
# LOVE can be used with any type of GP model, including exact GPs, multitask models and scalable approximations. Here we demonstrate LOVE in conjunction with KISS-GP, which has the amazing property of producing **constant time variances.**
#
# ## The KISS-GP + LOVE GP Model
#
# We now define the GP model. For more details on the use of GP models, see our simpler examples. This model uses a `GridInterpolationKernel` (SKI) with an Deep RBF base kernel. The forward method passes the input data `x` through the neural network feature extractor defined above, scales the resulting features to be between 0 and 1, and then calls the kernel.
#
# The Deep RBF kernel (DKL) uses a neural network as an initial feature extractor. In this case, we use a fully connected network with the architecture `d -> 1000 -> 500 -> 50 -> 2`, as described in the original DKL paper. All of the code below uses standard PyTorch implementations of neural network layers.
# +
class LargeFeatureExtractor(torch.nn.Sequential):
def __init__(self, input_dim):
super(LargeFeatureExtractor, self).__init__()
self.add_module('linear1', torch.nn.Linear(input_dim, 1000))
self.add_module('relu1', torch.nn.ReLU())
self.add_module('linear2', torch.nn.Linear(1000, 500))
self.add_module('relu2', torch.nn.ReLU())
self.add_module('linear3', torch.nn.Linear(500, 50))
self.add_module('relu3', torch.nn.ReLU())
self.add_module('linear4', torch.nn.Linear(50, 2))
class GPRegressionModel(gpytorch.models.ExactGP):
def __init__(self, train_x, train_y, likelihood):
super(GPRegressionModel, self).__init__(train_x, train_y, likelihood)
self.mean_module = gpytorch.means.ConstantMean()
self.covar_module = gpytorch.kernels.GridInterpolationKernel(
gpytorch.kernels.ScaleKernel(gpytorch.kernels.RBFKernel()),
grid_size=100, num_dims=2,
)
# Also add the deep net
self.feature_extractor = LargeFeatureExtractor(input_dim=train_x.size(-1))
def forward(self, x):
# We're first putting our data through a deep net (feature extractor)
# We're also scaling the features so that they're nice values
projected_x = self.feature_extractor(x)
projected_x = projected_x - projected_x.min(0)[0]
projected_x = 2 * (projected_x / projected_x.max(0)[0]) - 1
# The rest of this looks like what we've seen
mean_x = self.mean_module(projected_x)
covar_x = self.covar_module(projected_x)
return gpytorch.distributions.MultivariateNormal(mean_x, covar_x)
likelihood = gpytorch.likelihoods.GaussianLikelihood()
model = GPRegressionModel(train_x, train_y, likelihood)
if torch.cuda.is_available():
model = model.cuda()
likelihood = likelihood.cuda()
# -
# ### Training the model
#
# The cell below trains the GP model, finding optimal hyperparameters using Type-II MLE. We run 20 iterations of training using the `Adam` optimizer built in to PyTorch. With a decent GPU, this should only take a few seconds.
# +
training_iterations = 1 if smoke_test else 20
# Find optimal model hyperparameters
model.train()
likelihood.train()
# Use the adam optimizer
optimizer = torch.optim.Adam(model.parameters(), lr=0.1) # Includes GaussianLikelihood parameters
# "Loss" for GPs - the marginal log likelihood
mll = gpytorch.mlls.ExactMarginalLogLikelihood(likelihood, model)
def train():
iterator = tqdm.notebook.tqdm(range(training_iterations))
for i in iterator:
optimizer.zero_grad()
output = model(train_x)
loss = -mll(output, train_y)
loss.backward()
iterator.set_postfix(loss=loss.item())
optimizer.step()
# %time train()
# -
# ## Computing predictive variances (KISS-GP or Exact GPs)
#
# ### Using standard computaitons (without LOVE)
#
# The next cell gets the predictive covariance for the test set (and also technically gets the predictive mean, stored in `preds.mean`) using the standard SKI testing code, with no acceleration or precomputation.
#
# **Note:** Full predictive covariance matrices (and the computations needed to get them) can be quite memory intensive. Depending on the memory available on your GPU, you may need to reduce the size of the test set for the code below to run. If you run out of memory, try replacing `test_x` below with something like `test_x[:1000]` to use the first 1000 test points only, and then restart the notebook.
# +
import time
# Set into eval mode
model.eval()
likelihood.eval()
with torch.no_grad():
start_time = time.time()
preds = likelihood(model(test_x))
exact_covar = preds.covariance_matrix
exact_covar_time = time.time() - start_time
print(f"Time to compute exact mean + covariances: {exact_covar_time:.2f}s")
# -
# ### Using LOVE
#
# Next we compute predictive covariances (and the predictive means) for LOVE, but starting from scratch. That is, we don't yet have access to the precomputed cache discussed in the paper. This should still be faster than the full covariance computation code above.
#
# To use LOVE, use the context manager `with gpytorch.settings.fast_pred_var():`
#
# You can also set some of the LOVE settings with context managers as well. For example, `gpytorch.settings.max_root_decomposition_size(100)` affects the accuracy of the LOVE solves (larger is more accurate, but slower).
#
# In this simple example, we allow a rank 100 root decomposition, although increasing this to rank 20-40 should not affect the timing results substantially.
# +
# Clear the cache from the previous computations
model.train()
likelihood.train()
# Set into eval mode
model.eval()
likelihood.eval()
with torch.no_grad(), gpytorch.settings.fast_pred_var(), gpytorch.settings.max_root_decomposition_size(100):
start_time = time.time()
preds = model(test_x)
fast_time_no_cache = time.time() - start_time
# -
# The above cell additionally computed the caches required to get fast predictions. From this point onwards, unless we put the model back in training mode, predictions should be extremely fast. The cell below re-runs the above code, but takes full advantage of both the mean cache and the LOVE cache for variances.
with torch.no_grad(), gpytorch.settings.fast_pred_var():
start_time = time.time()
preds = likelihood(model(test_x))
fast_covar = preds.covariance_matrix
fast_time_with_cache = time.time() - start_time
print('Time to compute mean + covariances (no cache) {:.2f}s'.format(fast_time_no_cache))
print('Time to compute mean + variances (cache): {:.2f}s'.format(fast_time_with_cache))
# ### Compute Error between Exact and Fast Variances
#
# Finally, we compute the mean absolute error between the fast variances computed by LOVE (stored in fast_covar), and the exact variances computed previously.
#
# Note that these tests were run with a root decomposition of rank 10, which is about the minimum you would realistically ever run with. Despite this, the fast variance estimates are quite good. If more accuracy was needed, increasing `max_root_decomposition_size` would provide even better estimates.
mae = ((exact_covar - fast_covar).abs() / exact_covar.abs()).mean()
print(f"MAE between exact covar matrix and fast covar matrix: {mae:.6f}")
# ## Computing posterior samples (KISS-GP only)
#
# With KISS-GP models, LOVE can also be used to draw fast posterior samples. (The same does not apply to exact GP models.)
#
# ### Drawing samples the standard way (without LOVE)
#
# We now draw samples from the posterior distribution. Without LOVE, we accomlish this by performing Cholesky on the posterior covariance matrix. This can be slow for large covariance matrices.
# +
import time
num_samples = 20 if smoke_test else 20000
# Set into eval mode
model.eval()
likelihood.eval()
with torch.no_grad():
start_time = time.time()
exact_samples = model(test_x).rsample(torch.Size([num_samples]))
exact_sample_time = time.time() - start_time
print(f"Time to compute exact samples: {exact_sample_time:.2f}s")
# -
# ### Using LOVE
#
# Next we compute posterior samples (and the predictive means) using LOVE.
# This requires the additional context manager `with gpytorch.settings.fast_pred_samples():`.
#
# Note that we also need the `with gpytorch.settings.fast_pred_var():` flag turned on. Both context managers respond to the `gpytorch.settings.max_root_decomposition_size(100)` setting.
# +
# Clear the cache from the previous computations
model.train()
likelihood.train()
# Set into eval mode
model.eval()
likelihood.eval()
with torch.no_grad(), gpytorch.settings.fast_pred_var(), gpytorch.settings.max_root_decomposition_size(200):
# NEW FLAG FOR SAMPLING
with gpytorch.settings.fast_pred_samples():
start_time = time.time()
_ = model(test_x).rsample(torch.Size([num_samples]))
fast_sample_time_no_cache = time.time() - start_time
# Repeat the timing now that the cache is computed
with torch.no_grad(), gpytorch.settings.fast_pred_var():
with gpytorch.settings.fast_pred_samples():
start_time = time.time()
love_samples = model(test_x).rsample(torch.Size([num_samples]))
fast_sample_time_cache = time.time() - start_time
print('Time to compute LOVE samples (no cache) {:.2f}s'.format(fast_sample_time_no_cache))
print('Time to compute LOVE samples (cache) {:.2f}s'.format(fast_sample_time_cache))
# -
# ### Compute the empirical covariance matrices
#
# Let's see how well LOVE samples and exact samples recover the true covariance matrix.
# +
# Compute exact posterior covar
with torch.no_grad():
start_time = time.time()
posterior = model(test_x)
mean, covar = posterior.mean, posterior.covariance_matrix
exact_empirical_covar = ((exact_samples - mean).t() @ (exact_samples - mean)) / num_samples
love_empirical_covar = ((love_samples - mean).t() @ (love_samples - mean)) / num_samples
exact_empirical_error = ((exact_empirical_covar - covar).abs()).mean()
love_empirical_error = ((love_empirical_covar - covar).abs()).mean()
print(f"Empirical covariance MAE (Exact samples): {exact_empirical_error}")
print(f"Empirical covariance MAE (LOVE samples): {love_empirical_error}")
# -
|
examples/02_Scalable_Exact_GPs/Simple_GP_Regression_With_LOVE_Fast_Variances_and_Sampling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Neural Networks
# (example from sklearn and https://github.com/amirziai/deep-learning-coursera)
# ## The problem: determine if an image has a cat or not, given labeled data
# <table>
# <tr>
# <th>Cat</th>
# <td> <img src="images/cat1.jpg" width="150"></td>
# <td> <img src="images/cat2.png" width="150"></td>
# </tr>
# <tr>
# <th>Non Cat</th>
# <td> <img src="images/no_cat1.jpeg" width="150"></td>
# <td> <img src="images/no_cat2.jpg" width="150"></td>
# </tr>
# </table>
#
# +
import numpy as np
from sklearn import neural_network
from lr_utils import load_dataset
import matplotlib.pyplot as plt
# %matplotlib inline
# -
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
# # NORMALIZATION
# +
X_train = X_train_orig.reshape(X_train_orig.shape[0], -1)
X_test = X_test_orig.reshape(X_test_orig.shape[0], -1)
X_train = X_train/255.
X_test = X_test/255.
Y_train = Y_train_orig.T
Y_test = Y_test_orig.T
# -
index = 102
# print (X_train[index].reshape(()))
plt.imshow(X_train_orig[index])
print ("y = " + str(Y_train[index]) + ", it's a '" + classes[np.squeeze(Y_train[index])].decode("utf-8") + "' picture.")
# ## sklearn implementation
clf = neural_network.MLPClassifier(
solver='lbfgs',
alpha=1e-5,
hidden_layer_sizes=(5, 2),
random_state=1,
max_iter=1000)
clf.fit(X_train, Y_train.ravel())
predictions = clf.predict(X_train)
print ('Accuracy: %d ' % ((np.sum(Y_train.ravel() == predictions))/float(Y_train.size)*100))
# +
predictions = clf.predict(X_test)
print ('Accuracy: %d ' % ((np.sum(Y_test.ravel() == predictions))/float(Y_test.size)*100))
predictions, Y_test.ravel()
# -
# # What if we implement it?
#
# (This implementation is for a logistic regression classifier)
def sigmoid(z):
return 1.0 / (1.0 + np.exp(-z))
def initialize_with_zeros(dim):
w = np.zeros((dim, 1))
b = 0.0
return w, b
def propagate(w, b, X, Y):
m = X.shape[1]
# FORWARD PROPAGATION
A = sigmoid(np.dot(w.T, X) + b) # compute activation
cost = (-1.0 / m) * np.sum(Y * np.log(A) + (1.0 - Y) * np.log(1.0 - A)) # compute cost
# BACKWARD PROPAGATION
dw = (1.0 / m) * np.dot(X, (A - Y).T)
db = (1.0 / m) * np.sum(A - Y)
cost = np.squeeze(cost)
grads = {"dw": dw,
"db": db}
return grads, cost
def optimize(w, b, X, Y, num_iterations, learning_rate, print_cost = False):
costs = []
for i in range(num_iterations):
# Cost and gradient
grads, cost = propagate(w, b, X, Y)
if i % 100 == 0:
costs.append(cost)
dw = grads["dw"]
db = grads["db"]
w = w - learning_rate * dw
b = b - learning_rate * db
if print_cost and i % 100 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
params = {"w": w,
"b": b}
grads = {"dw": dw,
"db": db}
return params, grads, costs
def predict(w, b, X):
m = X.shape[1]
Y_prediction = np.zeros((1,m))
w = w.reshape(X.shape[0], 1)
A = sigmoid(np.dot(w.T, X) + b)
Y_prediction[A >= 0.5] = 1
Y_prediction[A < 0.5] = 0
return Y_prediction
def model(X_train, Y_train, X_test, Y_test, num_iterations = 2000, learning_rate = 0.5, print_cost = False):
w, b = initialize_with_zeros(X_train.shape[0])
parameters, grads, costs = optimize(w, b, X_train, Y_train, num_iterations, learning_rate, print_cost)
w = parameters["w"]
b = parameters["b"]
Y_prediction_test = predict(w, b, X_test)
Y_prediction_train = predict(w, b, X_train)
print("train accuracy: {} %".format(100 - np.mean(np.abs(Y_prediction_train - Y_train)) * 100))
print("test accuracy: {} %".format(100 - np.mean(np.abs(Y_prediction_test - Y_test)) * 100))
d = {"costs": costs,
"Y_prediction_test": Y_prediction_test,
"Y_prediction_train" : Y_prediction_train,
"w" : w,
"b" : b,
"learning_rate" : learning_rate,
"num_iterations": num_iterations}
return d
d = model(X_train.T, Y_train.ravel(), X_test.T, Y_test.ravel(), num_iterations = 5000, learning_rate = 0.001, print_cost = True)
# +
index = 30
num_px = X_train_orig[index].shape[0]
plt.imshow(X_test[index,:].reshape((num_px, num_px, 3)))
print ("y = " + str(Y_test[index,0]) + ", you predicted that it is a \"" + classes[int(d["Y_prediction_test"][0,index])].decode("utf-8") + "\" picture.")
# -
# # Keras
from keras.models import Sequential
from keras.layers import Dense, Dropout
X_train.shape
model = Sequential([
Dense(5, input_shape=(X_train.shape[1],), activation="relu"),
Dropout(0.5),
Dense(1, activation="sigmoid")
])
model.compile(
optimizer="sgd",
loss="binary_crossentropy",
metrics="acc"
)
model.summary()
model.fit(X_train, Y_train, batch_size=32, epochs=5, validation_data=(X_test, Y_test))
|
teorico/demo_10_neural_networks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="of3HGFCW2ii7"
# <a id='Q0'></a>
# <center><a target="_blank" href="http://www.propulsion.academy"><img src="https://drive.google.com/uc?id=1McNxpNrSwfqu1w-QtlOmPSmfULvkkMQV" width="200" style="background:none; border:none; box-shadow:none;" /></a> </center>
# <center> <h4 style="color:#303030"> Python for Data Science, Homework, template: </h4> </center>
# <center> <h1 style="color:#303030">Simplified Breast Cancer Selection</h1> </center>
# <p style="margin-bottom:1cm;"></p>
# <center style="color:#303030"><h4>Propulsion Academy, 2021</h4></center>
# <p style="margin-bottom:1cm;"></p>
#
# <div style="background:#EEEDF5;border-top:0.1cm solid #EF475B;border-bottom:0.1cm solid #EF475B;">
# <div style="margin-left: 0.5cm;margin-top: 0.5cm;margin-bottom: 0.5cm">
# <p><strong>Goal:</strong> Practice binary classification on Breast Cancer data</p>
# <strong> Sections:</strong>
# <a id="P0" name="P0"></a>
# <ol>
# <li> <a style="color:#303030" href="#SU">Set Up </a> </li>
# <li> <a style="color:#303030" href="#P1">Exploratory Data Analysis</a></li>
# <li> <a style="color:#303030" href="#P2">Modeling</a></li>
# </ol>
# <strong>Topics Trained:</strong> Binary Classification.
# </div>
# </div>
#
# <nav style="text-align:right"><strong>
# <a style="color:#00BAE5" href="https://monolith.propulsion-home.ch/backend/api/momentum/materials/intro-2-ds-materials/" title="momentum"> SIT Introduction to Data Science</a>|
# <a style="color:#00BAE5" href="https://monolith.propulsion-home.ch/backend/api/momentum/materials/intro-2-ds-materials/weeks/week2/day1/index.html" title="momentum">Week 2 Day 1, Applied Machine Learning</a>|
# <a style="color:#00BAE5" href="https://colab.research.google.com/drive/17X_OTM8Zqg-r4XEakCxwU6VN1OsJpHh7?usp=sharing" title="momentum"> Assignment, Classification of breast cancer cells</a>
# </strong></nav>
# + [markdown] id="TRFmWZYGJp5j"
# ## Submitted by <NAME> and <NAME>
# + [markdown] id="ckLGGhLpmYD8"
# <a id='SU' name="SU"></a>
# ## [Set up](#P0)
# + id="WzyvLIkXKczO" colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"status": "ok", "timestamp": 1618926803534, "user_tz": -300, "elapsed": 185096, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02813448518636787842"}} outputId="b6dc6c2c-50ca-4534-fd4a-1237797ef0ce"
# !sudo apt-get install build-essential swig
# !curl https://raw.githubusercontent.com/automl/auto-sklearn/master/requirements.txt | xargs -n 1 -L 1 pip install
# !pip install -U auto-sklearn
# !pip install -U matplotlib
# !pip install pipelineprofiler
# !pip install shap
# !pip install --upgrade plotly
# !pip3 install -U scikit-learn
# + id="Qc9gb4tAKpVu" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1618926867030, "user_tz": -300, "elapsed": 9651, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02813448518636787842"}} outputId="a901dbf9-6e70-4235-f993-9f498a2dbc4b"
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split, cross_val_score, KFold
from pandas_profiling import ProfileReport
import matplotlib.pyplot as plt
import plotly
plotly.__version__
import plotly.graph_objects as go
import plotly.io as pio
import plotly.express as px
from plotly.subplots import make_subplots
# your code here
from scipy import stats
from sklearn.preprocessing import LabelEncoder, StandardScaler, OneHotEncoder
from sklearn.impute import SimpleImputer
from sklearn import preprocessing
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score, ConfusionMatrixDisplay,mean_squared_error
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.pipeline import Pipeline
import time
from google.colab import files
from sklearn import set_config
from sklearn.compose import ColumnTransformer
import autosklearn.classification
import PipelineProfiler
import shap
import datetime
from joblib import dump
import logging
# + [markdown] id="RspLUVmbsTLB"
# **Connect** to your Google Drive
# + id="VcNDUjQZNxnV" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1618926903778, "user_tz": -300, "elapsed": 29402, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02813448518636787842"}} outputId="48a0efca-471a-4f11-ab19-8c7dda6348d3"
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + id="Kv5qB1cWNyL5" executionInfo={"status": "ok", "timestamp": 1618926907695, "user_tz": -300, "elapsed": 1686, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02813448518636787842"}}
data_path = "/content/drive/MyDrive/Introduction2DataScience/exercises/sit_w2d2_ml_engineering_assignment/data/raw/"
# + id="gYP1c2D6lvZp" executionInfo={"status": "ok", "timestamp": 1618926908869, "user_tz": -300, "elapsed": 2427, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02813448518636787842"}}
model_path = "/content/drive/MyDrive/Introduction2DataScience/exercises/sit_w2d2_ml_engineering_assignment/models/"
# + id="XRlYspr9l5RM" executionInfo={"status": "ok", "timestamp": 1618926908869, "user_tz": -300, "elapsed": 1901, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02813448518636787842"}}
timesstr = str(datetime.datetime.now()).replace(' ', '_')
# + id="g0fEkXLwl52w" executionInfo={"status": "ok", "timestamp": 1618926908870, "user_tz": -300, "elapsed": 1521, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02813448518636787842"}}
logging.basicConfig(filename=f"{model_path}explog_{timesstr}.log", level=logging.INFO)
# + [markdown] id="tiBsfiFvstdj"
# Please Download the data from [this source](https://drive.google.com/file/d/1af2YyHIp__OdpuUeOZFwmwOvCsS0Arla/view?usp=sharing), and upload it on your introduction2DS/data google drive folder.
# + [markdown] id="_TwNrMQVmX4S"
# <a id='P1' name="P1"></a>
# ## [Loading Data and Train-Test Split](#P0)
#
# + id="vhLQe4H-qqLH"
df = pd.read_csv(f"{data_path}data-breast-cancer.csv")
# + id="saTmF6mRukU8"
#encode the categrical column
encoder = LabelEncoder()
df['diagnosis'] = encoder.fit_transform(df['diagnosis'])
# + id="F8FOA9Izwa3E"
df.drop(['Unnamed: 32','id'], axis=1, inplace=True)
# + id="dEq5kR5Zma-D"
test_size = 0.2
random_state = 45
# + id="RSkI9tD0mk5p"
train, test = train_test_split(df, test_size=test_size, random_state=random_state)
# + id="OItQpNaZmoKV"
logging.info(f'train test split with test_size={test_size} and random state={random_state}')
# + id="d4z3n5crmsz2"
train.to_csv(f'{data_path}Breast_Cancer_Train.csv', index=False)
# + id="i6X_RVujmy0E"
train= train.copy()
# + id="FtewZj54m1tk"
test.to_csv(f'{data_path}Breast_Cancer_Test.csv', index=False)
# + id="Nau0uDh6nOhW"
test = test.copy()
# + [markdown] id="hfxPkrSZnRSu"
# <a id='P2' name="P2"></a>
# ## [Modelling](#P0)
# + id="Zlb4ji55nTfx"
X_train, y_train = train.iloc[:,1:], train['diagnosis']
# + id="B4-4PYCGncIN"
total_time = 600
per_run_time_limit = 30
# + id="IuyRg1Vtngs5"
automl = autosklearn.classification.AutoSklearnClassifier(
time_left_for_this_task=total_time,
per_run_time_limit=per_run_time_limit,
)
automl.fit(X_train, y_train)
# + id="DWZjwLqhnt7z"
logging.info(f'Ran autosklearn regressor for a total time of {total_time} seconds, with a maximum of {per_run_time_limit} seconds per model run')
# + id="HS_MuNWjnx1g"
dump(automl, f'{model_path}model{timesstr}.pkl')
# + id="HcqeouZXoGOH"
logging.info(f'Saved classification model at {model_path}model{timesstr}.pkl ')
# + id="kRsAKBIqoKzp"
logging.info(f'autosklearn model statistics:')
logging.info(automl.sprint_statistics())
# + id="ZSyu675wogna"
# profiler_data= PipelineProfiler.import_autosklearn(automl)
# PipelineProfiler.plot_pipeline_matrix(profiler_data)
# + [markdown] id="vbQLe2QdoUx6"
# <a id='P2' name="P2"></a>
# ## [Model Evluation and Explainability](#P0)
# + id="F6zGAX5Qor7Y"
X_test, y_test = train.iloc[:,1:], train['diagnosis']
# + [markdown] id="aZAUeqplo1ZH"
# Now, we can attempt to predict the diagnosis prediction from our test set. To do that, we just use the .predict method on the object "automl" that we created and trained in the last sections:
# + id="NWB2iQeoo0VX"
y_pred = automl.predict(X_test)
# + [markdown] id="vIqWbLbXpEZP"
# Let's now evaluate it using the mean_squared_error function from scikit learn:
# + id="YNnD8ZufpG9x"
logging.info(f"Mean Squared Error is {mean_squared_error(y_test, y_pred)}, \n R2 score is {automl.score(X_test, y_test)}")
# + [markdown] id="uLyeHqGepJJp"
# we can also plot the y_test vs y_pred scatter:
# + id="9gYcHatIpOU6"
df = pd.DataFrame(np.concatenate((X_test, y_test.to_numpy().reshape(-1,1), y_pred.reshape(-1,1)), axis=1))
# + id="eeAG9fW2pS0n"
df.columns = ['radius_mean', 'texture_mean', 'perimeter_mean',
'area_mean', 'smoothness_mean', 'compactness_mean', 'concavity_mean',
'concave points_mean', 'symmetry_mean', 'fractal_dimension_mean',
'radius_se', 'texture_se', 'perimeter_se', 'area_se', 'smoothness_se',
'compactness_se', 'concavity_se', 'concave points_se', 'symmetry_se',
'fractal_dimension_se', 'radius_worst', 'texture_worst',
'perimeter_worst', 'area_worst', 'smoothness_worst',
'compactness_worst', 'concavity_worst', 'concave points_worst',
'symmetry_worst', 'fractal_dimension_worst', 'Predicted Target','True Target']
# + id="x3LX60cBpqSh"
fig = px.scatter(df, x='Predicted Target', y='True Target')
fig.write_html(f"{model_path}residualfig_{timesstr}.html")
# + id="NPmIV1XCpsY8"
logging.info(f"Figure of residuals saved as {model_path}residualfig_{timesstr}.html")
# + [markdown] id="sSlAaezUpvP0"
# #### Model Explainability
# + id="NFgTt58cpw5o"
explainer = shap.KernelExplainer(model = automl.predict, data = X_test.iloc[:50, :], link = "identity")
# + id="g4ePoWyOpz-E"
# Set the index of the specific example to explain
X_idx = 0
shap_value_single = explainer.shap_values(X = X_test.iloc[X_idx:X_idx+1,:], nsamples = 100)
X_test.iloc[X_idx:X_idx+1,:]
# print the JS visualization code to the notebook
# shap.initjs()
shap.force_plot(base_value = explainer.expected_value,
shap_values = shap_value_single,
features = X_test.iloc[X_idx:X_idx+1,:],
show=False,
matplotlib=True
)
plt.savefig(f"{model_path}shap_example_{timesstr}.png")
logging.info(f"Shapley example saved as {model_path}shap_example_{timesstr}.png")
# + id="MWN2To1Xp4Xr"
shap_values = explainer.shap_values(X = X_test.iloc[0:50,:], nsamples = 100)
# + id="55AF67rzp8kB"
# print the JS visualization code to the notebook
# shap.initjs()
fig = shap.summary_plot(shap_values = shap_values,
features = X_test.iloc[0:50,:],
show=False)
plt.savefig(f"{model_path}shap_summary_{timesstr}.png")
logging.info(f"Shapley summary saved as {model_path}shap_summary_{timesstr}.png")
# + [markdown] id="eTpULQqLMPt3"
# --------------
# # End of This Notebook
|
notebooks/W2D1_Breast_Cancer_Solution_Simplified.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 4. Wstępne przetwarzanie danych
#
# Przed przystąpieniem do analizy dane należy odpowiednio przygotować. Rzeczywiste dane moga zawierać błedy wynikające z metody pomiaru lub niepoprawnej akwizycji. Niekóre wartości mogą być niezgodne ze zbiorem danych i naszą wiedzą o domenie problemu. W danych mogą występować brakujące wartości lub powtórzenia tych samych pomiarów. Niektóre metody analizy wymagają też odpowiedniego przygotowania danych, np. wykonanie standaryzacji lub zamianę ciągłych wartości na dyskretne (dyskretyzacja) lub dyskretnych - na ciągłe.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Dane Iris
#
# Dane Iris zawierają pomiary rozmiarów płatków (petal) oraz rozmiarów liści kielicha (sepal) dokonanych dla 3 odmian Irysów: Setosa, Virginica i Versicolor
#
# <img src="https://s3.amazonaws.com/assets.datacamp.com/blog_assets/Machine+Learning+R/iris-machinelearning.png" alt="drawing" width="400"/>
#
# Dane można porać z adresu https://www.is.umk.pl/~grochu/wdm/files/iris-data.csv lub z katalogu z danymi w reposytorium ``dane/iris-data.csv``
#
# + slideshow={"slide_type": "slide"}
import pandas as pd
iris = pd.read_csv('https://www.is.umk.pl/~grochu/wdm/files/iris-data.csv')
print('Ilość przypadków = %d' % (iris.shape[0]))
print('Ilość zmiennych = %d' % (iris.shape[1]))
iris.head() # wyswielimy pięc pierwszch wierszy
# + [markdown] slideshow={"slide_type": "slide"}
# ## Brakujące wartości
#
# Na pierwszy rzut oka wszystko wydaje się być w porządku. Niemniej jednak pierwsze co powiniśmy sprawdzić to to czy w pliku nie ma braków danych. W tabeli Pandas takie wartości są reprezentowane za pomocą wartości `null`. Sprawdzmy, czy mamy takie dane w pliku za pomocą metody `isnull()`
# -
iris.isnull().values.any()
# Wygląda na to, że w danych są brakujące wartości. Sprawdzmy ile ich jest w każdej ze zmiennych.
iris.isnull().sum()
# Zobaczmy także jakiego typu są zmienne.
iris.info()
# + [markdown] slideshow={"slide_type": "slide"}
# Pierwsza zmienna powinna zawierać wartości numeryczne a jest reprezentowana jako `object`. Naprawdopodobniej w pliku występuje pewna wartość, która nie jest liczbą, dlatego Pandas zaimportował te wartości do typu `object` (w postaci napisów). Spróbujmy zamienić tą zmienną na zmienną nymeryczną za pomocą metody `to_numeric()`. Argument `errors='coerce'` sprawia, że wszystkie problematyczne wartości zostaną zamienione na NaN.
# +
sepal_numeric = pd.to_numeric(iris['sepal_length_cm'], errors='coerce')
print('Typ zmiennej: %s' % sepal_numeric.dtype)
print(sepal_numeric)
import numpy as np
np.where(sepal_numeric.isna() == True) # indeksy brakujących wartości
# + [markdown] slideshow={"slide_type": "slide"}
# Spradzmy, co było problemem.
# -
print(iris.sepal_length_cm[sepal_numeric.isna()])
# Okazuje się, że dwie wartości w pliku zamiast liczby zawierały znak zapytania (`?`).
#
# Wstawmy poprawioną zmienną we właściwe miejsce w danych.
iris.sepal_length_cm = sepal_numeric
iris.info()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Uzupełnianie wartości brakujących
#
# Jeżeli zależy nam na zachowaniu pomiarów zawierających braki to możemy spróbować wypełnic je odpowiednimi wartościami, np. takimi, które wadają się najbardziej pradopodobne, tj. wartością średnią zmiennej lub wartością modalną.
# +
iris.sepal_length_cm = iris['sepal_length_cm'].fillna(sepal_numeric.mean())
iris.sepal_length_cm.isna().values.any()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Usuwanie wartości brakujących
#
# Jednak najczęściej będziemy chcieli się pozbyć pomiarów posiadających brakujące dane. W przypadku, gdy danych jest dostatecznie dużo nie będzie to miało istotnego wpływu na wynik analizy.
#
# Usuńmy przypadki, które posiadają braki za pomocą metody `dropna()`.
#
# +
print("Obecnie w danych jest %d przypadków" % iris.shape[0])
iris = iris.dropna() # usuwanie wierszy zawierających wartości NaN
print("Po usunięciu braków pozostało %d przypadków" % iris.shape[0])
iris.isna().any()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Wartości odstające i inna anomalie
#
#
# Wypiszmy podstawowe statystyki o danych i sprawdzmy czy występują tam inne anomalie.
# -
iris.describe()
# Dzieki tego typu tabelkom możemy sprawdzić podstawowe informacje o danych oraz sprawdzić czy pomiary nie przekraczają rozsądnych przedziałów dla poszczególnych cech.
#
# Cecha `sepal_length_cm` posiada podejrzanie małą wartość minimalną (0.055 cm), zaź wartość minimalna zmiennej `sepal_width_cm` jest ujemna (-1.0). Szerokość i wysokość powinny być wartościami większymi od 0. Najprawdopodobniej jest to wartość błędna i ten pomiar należy usunąć.
#
# Wartości odstające najwygodniej wykryć za pomocą wykresu skrzynkowego.
# +
import matplotlib.pyplot as plt
import seaborn as sb
sb.boxplot(data=iris)
# -
# Skrzynia określa zakres od 1 do 3 kwartyla z kreską środkową oznaczjąca medianę (wartość środkowa). Punkty leżące za "wąsami" są podejrzane, są to wartości odstające, nietypowe dla rozkładu.
#
# Potwierdza się, że dwie pierwsze zmienne posiadają odstające wartości.
#
# Przypadki odstające tj. takie które znajdują się poza zakresem w które wpada większość danych, nie muszą jednoznacznie być błędne. Niemniej jednak należy im się dobrze przyjrzeć. I bardzo rzadko możemy jednoznacznie stwierdzić, czy taka wartość odstająca była błędem pomiaru, błędem na poziomie akwizycji czy też wprowadzania danych, czy może faktycznie jest odstępstwem od normy w samym fenomenie, który obserwujemy.
#
#
# Zobaczmy najpierw, które przypadki posiadają wartość ujemną, gdyż one z pewnością są błędne.
iris[iris['sepal_width_cm'] < 0.0]
# Jest jeden taki przypadek - usuńmy go.
iris = iris[iris['sepal_width_cm'] > 0.0]
# + [markdown] slideshow={"slide_type": "slide"}
# Badając rozkład danych przyjrzyjmy się również rozkładowi danych dla każdej pary zmiennych.
# Grupy przypadków zaznaczmy różnymi kolorami.
# +
import seaborn as sb
import matplotlib.pyplot as plt
sb.pairplot(iris, hue='class')
plt.show()
# + [markdown] slideshow={"slide_type": "-"}
# Z tego wykresu możemy wysnuć od razu następujące wnioski:
# - wartości odstające w zmiennej `sepal_length_cm` są bardzo wyraźnie widoczne i dotyczą odmiany Iris-Versicolor,
# - domeana klasy (`class`) ma 5 unikalnych wartości, podczas gdy powinna mieć ich 3.
# + [markdown] slideshow={"slide_type": "slide"}
# Zbadajmy najpierw zmienną `class`. Jest to zmienna kategoryczna.
# +
# wypiszmy jakie posiaday unikalne wartośći w kolumnie class
print(iris['class'].unique())
iris['class'].value_counts()
# -
# Możemy zauważyć, że musiał nastąpić błąd w kodowaniu danych, podczas wprowadzania danych zostały wykonane dwie literówki. Poprawmy to.
# +
iris.loc[iris['class'] == 'versicolor', 'class'] = 'Iris-versicolor'
# podoby efekt osiągniemy z pomoca metody replace()
iris.loc[:, 'class'] = iris['class'].replace('Iris-setossa', 'Iris-setosa')
print(iris['class'].unique())
print(iris['class'].value_counts())
# + [markdown] slideshow={"slide_type": "slide"}
# ## Błędy w danych
#
# Przyjrzyjmy się też danym w zmiennej `sepal_length_cm` odmiany `Iris-versicolor`, które posiadają podejrzanie małe wartości, mniejsze od 2.5cm
# +
#dla ułatwienia najpier zapiszę sobie indeks do wykrajania dla tych danych
broken_slice_idx = (iris['class'] == 'Iris-versicolor') & (iris['sepal_length_cm'] < 1.0)
iris.loc[broken_slice_idx]
# + [markdown] slideshow={"slide_type": "-"}
# Możemy wrócić, do tabeli ze statysykami i zauważyć, że wartość `sepal_length_cm` wynosiła średnio 5.6 + 1.3.
# -
print("%.1f +- %.1f" % (iris.sepal_length_cm.mean() , iris.sepal_length_cm.std()))
# + [markdown] slideshow={"slide_type": "-"}
# Wygląda na to że podczas wprowadzania danych zostały pomylone jednostki miary, dane zostały wprowadzone w milimetrach zamiast w cm. Oczywiście to należałoby w jakiś sposób potwierdzić, ale na ten moment zmienmy jednostki dla tych danych.
# + slideshow={"slide_type": "slide"}
iris.loc[broken_slice_idx, 'sepal_length_cm'] *= 100.0
# -
sb.boxplot(data=iris)
sb.pairplot(iris, hue='class')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Duplikaty
#
#
# -
# Dane mogą zawierać powtarzające się pomiary, np. w wyniku nieuwagi pomiary jednego obiektu mogły zostać kilkukrotnie wpisane do bazy danych. W przypadku danych Irys nie mamy pewności, czy powtarzające się dane są wynikami uzyskanymi dla róznych kwiatów. Spróbujmy jednak poszukać powtarzających się wierszy i je usunąć.
#
# Wypiszmy najpierw wszystkie wiersze, które się duplikują.
iris_d = iris.duplicated(keep=False)
iris_d
iris[iris_d]
# + [markdown] slideshow={"slide_type": "slide"}
# Argument `keep='first'` metody `duplicated()` pozostawia pierwsze wystąpienie powtarzającego się wiersza i pozwala zaindeksować pozostałe duplikaty. Usuńmy wszytkie (oprócz pierwszewgo) powtarzające się pomiary.
# +
iris_d = iris.duplicated(keep='first')
print('Ilość powtarzających się przypadkóws = %d' % (iris_d.sum()))
print('Liczba przypadków przed selekcją = %d' % (iris.shape[0]))
iris = iris.drop_duplicates()
print('Liczba przypadków po odrzuceniu powtarzających się przypadków = %d' % (iris.shape[0]))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Standaryzacja danych
#
# Standaryzacja - normalizacja zmiennych zamieniająca średnią $\mu$ na 0 (centrowanie) a odchylenie standardowe $\sigma$ na 1
#
# $$
# z=\frac{x-\mu}{\sigma}
# $$
# +
iris_num = iris.select_dtypes(include=np.number)
iris_std = (iris_num - iris_num.mean()) / iris_num.std()
iris_std.plot(kind='box')
# pd.options.display.float_format = '{:,.2f}'.format
iris_std.describe()
# +
# dane odstające leżaca dalej niż 3 odchylenia standardowe
outliers = ((iris_std > 3) | (iris_std < -3)).any(axis=1)
iris[outliers]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Normalizacja
#
# Normaliacja wartości zmiennych w ustalonym zakresie, zawzyczaj $[-1, 1]$
#
# $$
# z= 2 \frac{x - x_{min}}{x_{max}-x_{min}} - 1
# $$
#
#
# +
iris_norm = 2 * (iris_num - iris_num.min()) / (iris_num.max() - iris_num.min()) - 1
iris_norm.plot(kind='box')
iris_norm.describe()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Próbkowanie (sampling)
# -
sample = iris.sample(n=10) # wybieramy losowo 10 próbek
sample
sample = iris.sample(frac=0.05, random_state=13) # wybieramy losowo 5% próbek z całego zbioru
sample
sample = iris.sample(frac=0.05, replace=True, random_state=13) # wybieramy losowo 5% próbek, ale ta sama próbka może być wybrana wiele razy
sample
# + [markdown] slideshow={"slide_type": "slide"}
# ## Dyskretyzacja danych
#
# Zamiana zmiennych ciągłych na dyskretne
# -
iris['sepal_length_cm'].hist(bins=10) # podział zbioru na 10 elementów
# +
bins = pd.cut(iris['sepal_length_cm'], 3) # podział zbioru na 3 elementy o równych odstępach (mniej-więcej)
bins.value_counts(sort=False)
# -
bins = pd.qcut(iris['sepal_length_cm'], 4, labels=['a', 'b', 'c', 'd']) # podział zbioru na 4 elementy po równo w każdym (mniej-więcej)
bins.value_counts(sort=False)
iris['sepal_length_size'] = pd.cut(iris['sepal_length_cm'], 3, labels=['small', 'medium', 'large'])
iris.groupby(iris['sepal_length_size']).mean()
iris.info()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Zamiana kategorycznych danych na zmienne numeryczne
# ### Kodowanie one-hot
#
# Kodowanie *one-hot* - zamiana wartości kategorycznych na wektor binarny $[0, 0, 1, 0, \ldots, 0]$
#
# -
class_one_hot = pd.get_dummies(iris['class'])
class_one_hot
# + [markdown] slideshow={"slide_type": "slide"}
# ### Mapowanie wartości kategorycznych na liczby
# +
size_map = {
'small' : 1,
'medium' : 2,
'large' : 3
}
size_data = iris['sepal_length_size'].map(size_map)
size_data
# + [markdown] slideshow={"slide_type": "slide"}
# ## Zadanie
#
# Wczytaj dane "Breast Cancer Wisconsin" i przeprowadź preprocesing zgodnie z podanymi poniżej wytycznymi.
#
# Adres danech: https://archive.ics.uci.edu/ml/machine-learning-databases/breast-cancer-wisconsin/breast-cancer-wisconsin.data
#
# Dane zawierają wartości opisujące cechy jąder komórkowych obecnych na obrazie uzyskanym przy badaniu piersi dla dwóch grup badanaych: `benign` (złożliwy), `malignat` (łagodny).
#
# Oto lista zmiennych:
#
# ```
# # Attribute Domain
# -- -----------------------------------------
# 1. Sample code number id number
# 2. Clump Thickness 1 - 10
# 3. Uniformity of Cell Size 1 - 10
# 4. Uniformity of Cell Shape 1 - 10
# 5. Marginal Adhesion 1 - 10
# 6. Single Epithelial Cell Size 1 - 10
# 7. Bare Nuclei 1 - 10
# 8. Bland Chromatin 1 - 10
# 9. Normal Nucleoli 1 - 10
# 10. Mitoses 1 - 10
# 11. Class: (2 for benign, 4 for malignant)
#
# ```
#
# Wszystkie istotne cechy posiadają wartości numeryczne, ostatnia zmienna zawiera informacje o 2 klasach.
#
# 1. Wczytaj zbiór danych używając Pandas spod adresu https://archive.ics.uci.edu/ml/machine-learning-databases/breast-cancer-wisconsin/breast-cancer-wisconsin.data. Dane są w formacie zgodnym z CSV (wartości oddzielone przecinkami). Zwróć uwagę na to, że plik nie posiada nagłowka. Dodaj nazwy zmiennych (kolumn) zgodne z listą podanych wyżej artybutów.
# -
# + [markdown] slideshow={"slide_type": "-"}
# 2. Pierwsza zmienna zawiera liczbę porządkową (ID), unikatową dla każdego badanego. Jest ona nieistotna dla analizy - usuń ją ze zbioru.
# -
# + [markdown] slideshow={"slide_type": "-"}
# 3. Wartości brakujące w pliku kodowane są za pomocą znaku zapytania (`?`). Sprawdź dla ilu badanych występują braki i w których zmiennych się pojawiają. Zastąp wartości brakujące wartością modalną (zob. funkcja [mode()](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.mode.html)). Wszystkie cechy (oprócz klasy) powinny być zmiennymi numerycznymi.
# -
# + [markdown] slideshow={"slide_type": "-"}
# 4. Sprawdź, czy zbiór danych zawiera przypadki odstające lub nietypowe. Spodzewamy się, że wszystkie zmienne mają wartości od 1 do 10.
# -
# + [markdown] slideshow={"slide_type": "-"}
# 5. Usuń ze zbioru przypadki odstające, których wartości zmiennych leżą poza przedziałem $(\bar{x}-3\cdot\sigma, \bar{x}+ 3\cdot\sigma)$, gdzie $\bar{x}$ to wartość średnia cechy, $\sigma$ to odchyenie standardowe.
# -
# + [markdown] slideshow={"slide_type": "-"}
# 6. Sprawdź czy dane zawierają powtarzające się pomiary i usuń ze zbioru danych duplikaty.
# -
|
04_Preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="tTfbwGrz2P3J"
# # Download Images and Preprocess
# + id="11oYDl342Uxf"
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="lYO-cVXjFbFW"
# Raw Image Data Download https://github.com/VisionLearningGroup/taskcv-2017-public/tree/master/classification
# + id="Ja4-eUc324ZN"
# #!wget http://csr.bu.edu/ftp/visda17/clf/validation.tar
# !tar xvf /content/gdrive/MyDrive/Projects/syn_real_gan/data/01_raw/validation.tar
'''
!wget http://csr.bu.edu/ftp/visda17/clf/train.tar
!tar xvf train.tar
!wget http://csr.bu.edu/ftp/visda17/clf/test.tar
!tar xvf test.tar
!wget https://raw.githubusercontent.com/VisionLearningGroup/taskcv-2017-public/master/classification/data/image_list.txt
'''
# + id="Phuw9g7IHEcA"
drive.flush_and_unmount()
# + [markdown] id="SAOQIO98tIWu"
# ## Data Stat
# + id="3QI4mR--juoZ"
pwd
# + id="DfTbl2ZbtHt3"
# ls
# + [markdown] id="P5rUYWynwoCH"
# Show number of files in each class.
# + id="VhmpyMawtN4H"
import os, os.path
# simple version for working with CWD
#dir = '/content/gdrive/MyDrive/Projects/syn_real_gan/data/01_raw'
dir = '/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate'
def print_num_files(dir):
file_count = 0
for name in os.listdir(dir):
if os.path.isfile(dir+'/'+name):
file_count += 1
elif os.path.isdir(dir+'/'+name):
print_num_files(dir+'/'+name)
print(f'{dir}:{file_count}')
print_num_files(dir)
# + id="-vD7XbaDWpqB"
import os,shutil
import random
from tqdm import tqdm
import cv2
def build_cycle_gan_dir(cls_name,target_dir):
os.makedirs(target_dir,exist_ok=True)
os.makedirs(os.path.join(target_dir,'trainA' ) ,exist_ok=True)
os.makedirs(os.path.join(target_dir, 'trainB' ) ,exist_ok=True)
os.makedirs(os.path.join(target_dir,'testA' ) ,exist_ok=True)
os.makedirs(os.path.join(target_dir,'testB' ) ,exist_ok=True)
raw_dir = '/content/gdrive/MyDrive/Projects/syn_real_gan/data/01_raw'
full_dir = '/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/5_classes'
B_dir = os.path.join(raw_dir,'validation',cls_name)
A_dir = os.path.join(raw_dir,'train',cls_name)
print(f'{cls_name} list dir ...')
A_dir_files = os.listdir(A_dir)
B_dir_files = os.listdir(B_dir)
print(f'{cls_name} shuffling ...')
random.shuffle(A_dir_files)
random.shuffle(B_dir_files)
def process_file(file_path,target_path,full_path):
im = cv2.imread(file_path)
sp = im.shape
if not (sp[0]/sp[1] < 0.4 or sp[0]/sp[1] > 2.5):
if not (sp[0] < 200 or sp[1] < 200):
shutil.copy(file_path,target_path)
shutil.copy(file_path,full_path)
print(f'{file_path}:{sp} moved')
return True
return False
count_train = 0
for f in tqdm(B_dir_files):
if process_file(os.path.join(B_dir,f),os.path.join(target_dir,'trainB'),os.path.join(full_dir,'trainB')):
count_train += 1
for f in tqdm(A_dir_files[:count_train]):
shutil.copy(os.path.join(A_dir,f),os.path.join(target_dir,'trainA'))
shutil.copy(os.path.join(A_dir,f),os.path.join(full_dir,'trainA'))
'''
for f in tqdm(A_dir_files[:200]):
shutil.copy(os.path.join(A_dir,f),os.path.join(target_dir,'trainA'))
for f in tqdm(B_dir_files[:200]):
shutil.copy(os.path.join(B_dir,f),os.path.join(target_dir,'trainB'))
for f in tqdm(A_dir_files[200:220]):
shutil.copy(os.path.join(A_dir,f),os.path.join(target_dir,'testA'))
for f in tqdm(B_dir_files[200:220]):
shutil.copy(os.path.join(B_dir,f),os.path.join(target_dir,'testB'))
'''
def move_train_to_test(cls_name,target_dir):
full_dir = '/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/5_classes'
A_dir = os.path.join(target_dir,'trainA')
B_dir = os.path.join(target_dir,'trainB')
A_test_dir = os.path.join(target_dir,'testA')
B_test_dir = os.path.join(target_dir,'testB')
full_A_dir = os.path.join(full_dir,'trainA')
full_B_dir = os.path.join(full_dir,'trainB')
full_A_test_dir = os.path.join(full_dir,'testA')
full_B_test_dir = os.path.join(full_dir,'testB')
print(f'{cls_name} list dir ...')
A_dir_files = os.listdir(A_dir)
B_dir_files = os.listdir(B_dir)
print(f'{cls_name} shuffling ...')
random.shuffle(A_dir_files)
random.shuffle(B_dir_files)
for f in tqdm(A_dir_files[:50]):
shutil.move(os.path.join(A_dir,f),A_test_dir)
shutil.move(os.path.join(full_A_dir,f),full_A_test_dir)
for f in tqdm(B_dir_files[:50]):
shutil.move(os.path.join(B_dir,f),B_test_dir)
shutil.move(os.path.join(full_B_dir,f),full_B_test_dir)
# + id="wYkzJfc3ACyh"
build_cycle_gan_dir('car','/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/car')
build_cycle_gan_dir('motorcycle','/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/motorcycle')
build_cycle_gan_dir('horse','/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/horse')
build_cycle_gan_dir('aeroplane','/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/aeroplane')
build_cycle_gan_dir('plant','/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/plant')
# + id="8DDV9Y124iZG"
move_train_to_test('car','/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/car')
move_train_to_test('motorcycle','/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/motorcycle')
move_train_to_test('horse','/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/horse')
move_train_to_test('aeroplane','/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/aeroplane')
move_train_to_test('plant','/content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/plant')
# + id="XqO19j6MK9x3"
# rm -r /content/gdrive/MyDrive/Projects/syn_real_gan/data/02_intermediate/car/testA/*
# + id="Q0_3Jb_M5IrC"
|
scripts/CycleGAN_Download_Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
#export
from exp.nb_00 import *
import operator
def test(a,b,cmp,cname=None):
if cname is None: cname = cmp.__name__
assert cmp(a,b), f'{cname}:\n{a}\n{b}'
def test_eq(a,b): test(a,b,operator.eq, '==')
# -
test_eq(TEST, 'test')
# #!python run_notebook.py 01_matmul.ipynb # to run tests in console:
# +
#export
from pathlib import Path
from IPython.core.debugger import set_trace
# from fastai.torch_basics import *
# from fastai.data.external import *
from fastai.vision.all import *
import pickle, gzip, math, torch, matplotlib as mpl
import matplotlib.pyplot as plt
from torch import tensor
MNIST_URL = 'http://deeplearning.net/data/mnist/mnist.pkl'
# -
path = untar_data(URLs.MNIST)
path.ls()
(path/'training').ls()
files = get_image_files(path)
files
mnist = DataBlock(blocks=(ImageBlock(cls=PILImageBW), CategoryBlock),
get_items = get_image_files,
splitter = GrandparentSplitter('training', 'testing'),
get_y = parent_label)
dls = mnist.dataloaders(untar_data(URLs.MNIST))
dls.show_batch()
# +
# learn = cnn_learner(dls, resnet34, metrics=error_rate)
# learn.fine_tune(1)
# -
dls.train_ds
dls.valid_ds
type(dls.train_ds)
dls.train_ds[0]
dls.train_ds[0][0]
type(dls.train_ds[0][0])
dls.train_ds[0][1]
x_train, y_train, x_valid, y_valid = [], [], [], []
for elem in dls.train_ds:
x_train.append(elem[0])
y_train.append(elem[1])
for elem in dls.valid_ds:
x_valid.append(elem[0])
y_valid.append(elem[1])
def x_to_tensor(listed):
return tensor(list(map(array, listed)), dtype=torch.float32).reshape(len(listed), -1)
def y_to_tensor(listed):
return tensor(array(listed))
x_train = x_to_tensor(x_train)
x_valid = x_to_tensor(x_valid)
y_train = y_to_tensor(y_train)
y_valid = y_to_tensor(y_valid)
n,c = x_train.shape
x_train, x_train.shape, y_train, y_train.shape, y_train.min(), y_train.max()
x_valid.shape
assert n == y_train.shape[0] == 60000
test_eq(c, 28*28)
test_eq(y_train.min(), 0)
test_eq(y_train.max(), 9)
mpl.rcParams['image.cmap'] = 'gray'
img = x_train[0]
img.view(28,28).type()
plt.imshow(img.view(28,28));
# # Initial python model
weights = torch.randn(784,10)
bias = torch.zeros(10)
# ## Matrix multiplication
def matmul(a,b):
ar,ac = a.shape
br,bc = b.shape
assert ac == br
c = torch.zeros(ar,bc)
for i in range(ar):
for j in range(bc):
for k in range(ac):
c[i,j] += a[i,k] * b[k,j]
return c
m1 = x_valid[:5]
m2 = weights
m1.shape, m2.shape
# %time t1 = matmul(m1,m2)
t1.shape
len(x_train)
# ## Elementwise ops
a = tensor([10., 6, -4])
b = tensor([2., 8, 7])
a,b
(a < b).float().mean()
m = tensor([[1., 2, 3], [4,5,6], [7,8,9]]); m
(m*m).sum().sqrt()
# ## Elementwise matmul
def matmul(a,b):
ar,ac = a.shape
br,bc = b.shape
assert ac == br
c = torch.zeros(ar,bc)
for i in range(ar):
for j in range(bc):
c[i,j] = (a[i,:] * b[:,j]).sum()
return c
# %timeit -n 10 _=matmul(m1, m2)
665/1.13
#export
def near(a,b): return torch.allclose(a, b, rtol=1e-3, atol=1e-5)
def test_near(a,b): test(a,b,near)
test_near(t1, matmul(m1,m2))
# ## Broadcasting
c = tensor([10.,20,30]); c
m
m.shape, c.shape
t = c.expand_as(m)
t
t.storage()
t.stride(), t.shape
c.unsqueeze(0)
c.unsqueeze(1)
c[None]
c[:,None]
c[...,None]
c[None,...]
# ## matmul with broadcasting
m1.shape, m2.shape
def matmul(a,b):
ar,ac = a.shape
br,bc = b.shape
assert ac == br
c = torch.zeros(ar,bc)
for i in range(ar):
c[i] = (a[i,:,None] * b).sum(dim=0)
return c
# %timeit -n 10 _ = matmul(m1,m2)
test_near(t1, matmul(m1,m2))
# ## Einstein summation
def matmul(a,b): return torch.einsum('ik,kj->ij', a, b)
# %timeit -n 10 _ = matmul(m1,m2)
test_near(t1, matmul(m1,m2))
# ## pytorch op
# %timeit -n 10 _ = m1.matmul(m2)
# %timeit -n 10 _ = m1 @ m2
test_near(t1, m1@m2)
# # Export
# !python notebook2script.py 01_matmul.ipynb
|
nbs/dl2/selfmade/01_matmul.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Let's make a LogisticRegression
#
# **Didn't work very well**
import pandas as pd
import numpy as np
import requests
import json
df_matches_grouped = pd.read_pickle("../data/df_matches_grouped.pkl")
model_df = df_matches_grouped[["match_id", "radiant_1", "radiant_2", "radiant_3", "radiant_4", "radiant_5", "dire_1", "dire_2", "dire_3", "dire_4", "dire_5"]]
target_df = df_matches_grouped["radiant_win"]
model_df
# +
feature_set = ["radiant_1", "radiant_2", "radiant_3", "radiant_4", "radiant_5", "dire_1", "dire_2", "dire_3", "dire_4", "dire_5",]
radiant_features = (pd.get_dummies(model_df["radiant_1"], prefix="radiant")
+ pd.get_dummies(model_df["radiant_2"], prefix="radiant")
+ pd.get_dummies(model_df["radiant_3"], prefix="radiant")
+ pd.get_dummies(model_df["radiant_4"], prefix="radiant")
+ pd.get_dummies(model_df["radiant_5"], prefix="radiant"))
dire_features = (pd.get_dummies(model_df["dire_1"], prefix="dire")
+ pd.get_dummies(model_df["dire_2"], prefix="dire")
+ pd.get_dummies(model_df["dire_3"], prefix="dire")
+ pd.get_dummies(model_df["dire_4"], prefix="dire")
+ pd.get_dummies(model_df["dire_5"], prefix="dire"))
model_df = pd.concat([radiant_features, dire_features], axis=1)
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(model_df, target_df, test_size=0.1)
# +
from sklearn.linear_model import LogisticRegression
model = LogisticRegression()
model.fit(X_train, y_train)
# -
train_predictions = model.predict(X_train)
test_predictions = model.predict(X_test)
# +
X_train["predictions"] = train_predictions.tolist()
X_train["target"] = y_train
X_test["predictions"] = test_predictions.tolist()
X_test["target"] = y_test
# -
X_train.head(100)
# +
X_train["prediction_correct"] = np.where(X_train["predictions"] == X_train["target"], 1, 0)
X_train["prediction_correct"].sum() / len(X_train["prediction_correct"])
# +
X_test["prediction_correct"] = np.where(X_test["predictions"] == X_test["target"], 1, 0)
X_test["prediction_correct"].sum() / len(X_test["prediction_correct"])
|
notebooks/deprecated/logistic_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DAT210x - Programming with Python for DS
# ## Module3 - Lab3
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib
from mpl_toolkits.mplot3d import Axes3D
# %matplotlib notebook
# +
# Look pretty...
# matplotlib.style.use('ggplot')
plt.style.use('ggplot')
# -
# Load up the wheat seeds dataset into a dataframe. We've stored a copy in the Datasets directory.
df = pd.read_csv('C:\\Users\\ashish.r\\Documents\\GitHub\\DAT210x\\Module3\\Datasets\\wheat.data')
df.head()
# Create a new 3D subplot using figure `fig`, which we've defined for you below. Use that subplot to draw a 3D scatter plot using the `area`, `perimeter`, and `asymmetry` features. Be sure so use the optional display parameter `c='red'`, and also label your axes:
# +
fig = plt.figure()
ax = fig.add_subplot(111,projection='3d')
ax.set_xlabel('area')
ax.set_ylabel('perimeter')
ax.set_zlabel('asymmetry')
ax.scatter(df.area, df.perimeter, df.asymmetry, c='r', marker='.')
# -
# Create another 3D subplot using fig. Then use the subplot to graph a 3D scatter plot of the `width`, `groove`, and `length` features. Be sure so use the optional display parameter `c='green'`, and be sure to label your axes:
# +
fig = plt.figure()
ay = fig.add_subplot(111,projection='3d')
ay.set_xlabel('width')
ay.set_ylabel('groove')
ay.set_zlabel('length')
ay.scatter(df.width,df.groove, df.length, c='g', marker='.')
# -
# Finally, display the graphs:
plt.show()
|
Module3/Module3 - Lab3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 05. Install Chrome Extension(BQ Mate, Streak BigQuery Developer Tools)
# - 본 문서에서 알려드릴 내용은 Chrome Extension과 관련된 내용입니다!
# - BigQuery를 사용하면 다 좋은데, 제가 날리는 쿼리의 비용이 얼마일지 궁금할 때가 있습니다
# - 이것을 해결해주는 크롬 확장 프로그램이 바로 [BQ Mate](https://chrome.google.com/webstore/detail/bigquery-mate/nepgdloeceldecnoaaegljlichnfognh)입니다!
# - 또한 설치한 것은 [Streak BigQuery Developer Tools](https://chrome.google.com/webstore/detail/streak-bigquery-developer/lfmljmpmipdibdhbmmaadmhpaldcihgd)입니다! 이 친구는 센트 단위로 Cost를 보여줘서.. 설치했습니다
# - 2개 중 1개만 설치해도 무방합니다
# ### BQ Mate와 Streak BigQuery Developer Tools를 설치한 화면
# <img src="../images/014_install_chrome_extension_01.png" width="700" height="700">
# - 네모친 부분에 The cost will be around ~ 이 부분이 예상 금액입니다!
# <img src="../images/014_install_chrome_extension_02.png" width="700" height="700">
# - BQ Mate의 큰 장점 중 하나는 별도의 설정이 존재하는 것
# - 여기서 Find/Replace를 할 수 있고, Separator 설정도 할 수 있습니다!
# - 또한 default인 legacySQL에서 standardSQL로 변경할 수 있습니다
# - 단순한 기능이지만 도움이 되길 :)
# # BQ Mate 설명글
# - 현재 작동하지 않는 기능도 있습니다
# ---
# BigQuery Mate
# Companion tool for BigQuery UI used by myself and my peers at Viant Inc.
# (Vindico, Specific Media, Myspace, Xumo). It adds some functionality
# to BigQuery Native UI that is found helpful in day on day activity.
# BigQuery Mate Features: (see Help for details)
# 1. Control Nav Panel size - retired on Jan.31, 2016
# 2. Hide/Show Datasets Panel
# 3. Datasets Filtering
# 4. Datasets Count (Filtered/Total)
# 5. Monetary Cost estimation based on processed bytes
# 6. Keywords Auto-Capitalization
# 7. Comment/Uncomment Selection or Outside Selection
# 8. Support for Parametrized Query / Saved Query with Parameters
# 9. Tables Decoration / Manipulation
# 10. Find / Replace Functionality (with support for regular expressions)
# 11. Tabs Auto naming after query name (saved query, new query)
# 12. Navigator: list of all BQ sessions across your PC
# with quick info and ability to jump toSome Useful Links:
# New BigQuery Session
# Query Reference (Legacy SQL)
# Query Reference (Standard SQL)
# Google Cloud Status
# Ask on Stack Overflow
# BigQuery Mate on Trello
# 13. Query Outline
# 14. Extended Result Panel
# 15. F5 to execute query + Ctrl-F5 to run selection
# 16. Interactive Pivots / Charts of Query Result
# 17. Search / Filter Recent Queries
# 18. Tables Filtering
# 19. Add All Fields to Query Editor, with or w/o alias
# 20. Thousands separator for numeric fields
# 21. Contextual Help via F1 or Right-Click Menu
# 22. SQL Version Quick Switch (Click on SQL button or Alt-L) Recent!
# 23. Save Default SQL Version (Double CLick on SQL button or Shift-Alt-L) Recent!
# 24. Support for BigQuery Named Paramaters (@param). New!
# Magnus style parameters still supported (<var_param>)
# 25. Keyboard Shortcuts to view previous/next page - Ctrl-ArrowLeft, Ctrl-ArrowRight
# and first/last page - Ctrl-ArrowDown, Ctrl-ArrowUp
# Warning: Google Team is constantly working on improving their BigQuery UI. Sometimes this breaks
# features of the Tool. When this is a case, please calm down, stay tuned and check periorically
# for extension updates. I am trying to fix issues as soon as they are introduced and I see them!
|
tutorials/02-Utils/05. Install_Chrome_Extension.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DAT210x - Programming with Python for DS
# ## Module5- Lab4
# +
import math
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib
from sklearn import preprocessing
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans
# You might need to import more modules here..
# .. your code here ..
matplotlib.style.use('ggplot') # Look Pretty
c = ['red', 'green', 'blue', 'orange', 'yellow', 'brown']
# -
# You can experiment with these parameters:
PLOT_TYPE_TEXT = False # If you'd like to see indices
PLOT_VECTORS = True # If you'd like to see your original features in P.C.-Space
# ### Some Convenience Functions
def drawVectors(transformed_features, components_, columns, plt):
num_columns = len(columns)
# This function will project your *original* feature (columns)
# onto your principal component feature-space, so that you can
# visualize how "important" each one was in the
# multi-dimensional scaling
# Scale the principal components by the max value in
# the transformed set belonging to that component
xvector = components_[0] * max(transformed_features[:,0])
yvector = components_[1] * max(transformed_features[:,1])
## Visualize projections
# Sort each column by its length. These are your *original*
# columns, not the principal components.
important_features = { columns[i] : math.sqrt(xvector[i]**2 + yvector[i]**2) for i in range(num_columns) }
important_features = sorted(zip(important_features.values(), important_features.keys()), reverse=True)
print("Projected Features by importance:\n", important_features)
ax = plt.axes()
for i in range(num_columns):
# Use an arrow to project each original feature as a
# labeled vector on your principal component axes
plt.arrow(0, 0, xvector[i], yvector[i], color='b', width=0.0005, head_width=0.02, alpha=0.75, zorder=600000)
plt.text(xvector[i]*1.2, yvector[i]*1.2, list(columns)[i], color='b', alpha=0.75, zorder=600000)
return ax
def doPCA(data, dimensions=2):
model = PCA(n_components=dimensions, svd_solver='randomized', random_state=7)
model.fit(data)
return model
def doKMeans(data, num_clusters=2):
# TODO: Do the KMeans clustering here, passing in the # of clusters parameter
# and fit it against your data. Then, return a tuple containing the cluster
# centers and the labels.
#
# Hint: Just like with doPCA above, you will have to create a variable called
# `model`, which will be a SKLearn K-Means model for this to work.
# TODO: Use K-Means to try and find seven cluster centers in this df.
# Be sure to name your kmeans model `model` so that the printing works.
#
kmodel = KMeans(n_clusters=n_clusters)
kmodel.fit(data)
return kmodel.cluster_centers_, kmodel.labels_
# Load up the dataset. It may or may not have nans in it. Make sure you catch them and destroy them, by setting them to `0`. This is valid for this dataset, since if the value is missing, you can assume no money was spent on it.
df = pd.read_csv('./Datasets/Wholesale customers data.csv', header=0)
df = df.fillna(0)
#df.shape
#df.head(10)
df.dtypes
# As instructed, get rid of the `Channel` and `Region` columns, since you'll be investigating as if this were a single location wholesaler, rather than a national / international one. Leaving these fields in here would cause KMeans to examine and give weight to them:
slicecols = ['Fresh', 'Milk', 'Grocery', 'Frozen', 'Detergents_Paper', 'Delicassen']
df = df.loc[:, slicecols]
# Fix column names
fixcols = ['Fresh', 'Dairy', 'Grocery', 'Frozen', 'Household', 'Deli']
df.columns = fixcols
df.head(10)
# Before unitizing / standardizing / normalizing your data in preparation for K-Means, it's a good idea to get a quick peek at it. You can do this using the `.describe()` method, or even by using the built-in pandas `df.plot.hist()`:
df.plot.hist()
plt.show()
df.describe()
# Having checked out your data, you may have noticed there's a pretty big gap between the top customers in each feature category and the rest. Some feature scaling algorithms won't get rid of outliers for you, so it's a good idea to handle that manually---particularly if your goal is NOT to determine the top customers.
#
# After all, you can do that with a simple Pandas `.sort_values()` and not a machine learning clustering algorithm. From a business perspective, you're probably more interested in clustering your +/- 2 standard deviation customers, rather than the top and bottom customers.
#
# Remove top 5 and bottom 5 samples for each column:
drop = {}
for col in df.columns:
# Bottom 5
sort = df.sort_values(by=col, ascending=True)
if len(sort) > 5: sort=sort[:5]
for index in sort.index: drop[index] = True # Just store the index once
# Top 5
sort = df.sort_values(by=col, ascending=False)
if len(sort) > 5: sort=sort[:5]
for index in sort.index: drop[index] = True # Just store the index once
# Drop rows by index. We do this all at once in case there is a collision. This way, we don't end up dropping more rows than we have to, if there is a single row that satisfies the drop for multiple columns. Since there are 6 rows, if we end up dropping < 5*6*2 = 60 rows, that means there indeed were collisions:
print("Dropping {0} Outliers...".format(len(drop)))
df.drop(inplace=True, labels=drop.keys(), axis=0)
df.describe()
# ### What are you interested in?
# Depending on what you're interested in, you might take a different approach to normalizing/standardizing your data.
#
# You should note that all columns left in the dataset are of the same unit. You might ask yourself, do I even need to normalize / standardize the data? The answer depends on what you're trying to accomplish. For instance, although all the units are the same (generic money unit), the price per item in your store isn't. There may be some cheap items and some expensive one. If your goal is to find out what items people tend to buy together but you didn't "unitize" properly before running kMeans, the contribution of the lesser priced item would be dwarfed by the more expensive item. This is an issue of scale.
#
# For a great overview on a few of the normalization methods supported in SKLearn, please check out: https://stackoverflow.com/questions/30918781/right-function-for-normalizing-input-of-sklearn-svm
#
# Suffice to say, at the end of the day, you're going to have to know what question you want answered and what data you have available in order to select the best method for your purpose. Luckily, SKLearn's interfaces are easy to switch out so in the mean time, you can experiment with all of them and see how they alter your results.
#
# 5-sec summary before you dive deeper online:
# ### Normalization
# Let's say your user spend a LOT. Normalization divides each item by the average overall amount of spending. Stated differently, your new feature is = the contribution of overall spending going into that particular item: \$spent on feature / \$overall spent by sample.
# ### MinMax
# What % in the overall range of $spent by all users on THIS particular feature is the current sample's feature at? When you're dealing with all the same units, this will produce a near face-value amount. Be careful though: if you have even a single outlier, it can cause all your data to get squashed up in lower percentages.
#
# Imagine your buyers usually spend \$100 on wholesale milk, but today only spent \$20. This is the relationship you're trying to capture with MinMax. NOTE: MinMax doesn't standardize (std. dev.); it only normalizes / unitizes your feature, in the mathematical sense. MinMax can be used as an alternative to zero mean, unit variance scaling. [(sampleFeatureValue-min) / (max-min)] * (max-min) + min Where min and max are for the overall feature values for all samples.
# ### Back to The Assignment
# Un-comment just ***ONE*** of lines at a time and see how alters your results. Pay attention to the direction of the arrows, as well as their LENGTHS:
#T = preprocessing.StandardScaler().fit_transform(df)
#T = preprocessing.MinMaxScaler().fit_transform(df)
#T = preprocessing.MaxAbsScaler().fit_transform(df)
T = preprocessing.Normalizer().fit_transform(df)
#T = df # No Change
# Sometimes people perform PCA before doing KMeans, so that KMeans only operates on the most meaningful features. In our case, there are so few features that doing PCA ahead of time isn't really necessary, and you can do KMeans in feature space. But keep in mind you have the option to transform your data to bring down its dimensionality. If you take that route, then your Clusters will already be in PCA-transformed feature space, and you won't have to project them again for visualization.
# +
# Do KMeans
n_clusters = 3
centroids, labels = doKMeans(T, n_clusters)
# -
# Print out your centroids. They're currently in feature-space, which is good. Print them out before you transform them into PCA space for viewing
centroids
# Now that we've clustered our KMeans, let's do PCA, using it as a tool to visualize the results. Project the centroids as well as the samples into the new 2D feature space for visualization purposes:
display_pca = doPCA(T)
T = display_pca.transform(T)
CC = display_pca.transform(centroids)
# Visualize all the samples. Give them the color of their cluster label
fig = plt.figure()
ax = fig.add_subplot(111)
if PLOT_TYPE_TEXT:
# Plot the index of the sample, so you can further investigate it in your dset
for i in range(len(T)): ax.text(T[i,0], T[i,1], df.index[i], color=c[labels[i]], alpha=0.75, zorder=600000)
ax.set_xlim(min(T[:,0])*1.2, max(T[:,0])*1.2)
ax.set_ylim(min(T[:,1])*1.2, max(T[:,1])*1.2)
else:
# Plot a regular scatter plot
sample_colors = [ c[labels[i]] for i in range(len(T)) ]
ax.scatter(T[:, 0], T[:, 1], c=sample_colors, marker='o', alpha=0.2)
# Plot the Centroids as X's, and label them
ax.scatter(CC[:, 0], CC[:, 1], marker='x', s=169, linewidths=3, zorder=1000, c=c)
for i in range(len(centroids)):
ax.text(CC[i, 0], CC[i, 1], str(i), zorder=500010, fontsize=18, color=c[i])
# Display feature vectors for investigation:
if PLOT_VECTORS:
drawVectors(T, display_pca.components_, df.columns, plt)
# Add the cluster label back into the dataframe and display it:
df['label'] = pd.Series(labels, index=df.index)
df
plt.show()
|
Module5/Module5 - Lab4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install efficientnet -q
# !pip install iterative-stratification
import os
import tensorflow_addons as tfa
import efficientnet.tfkeras as efn
import numpy as np
import pandas as pd
from kaggle_datasets import KaggleDatasets
from sklearn.model_selection import train_test_split
import tensorflow as tf
from sklearn.model_selection import GroupKFold
import matplotlib.pyplot as plt
import glob
from tqdm import tqdm
from iterstrat.ml_stratifiers import MultilabelStratifiedKFold,MultilabelStratifiedShuffleSplit
import tensorflow as tf
import numpy as np
# +
# -
#fixed version
def map_loss(y_true,y_pred):
lss_fn = tfa.losses.SigmoidFocalCrossEntropy()
# y_true should be (BS,N_CLS)
bs = y_true.shape[0]
tp_mul = y_true * y_pred
y_pred_sort = tf.sort(y_pred,axis=-1,direction='DESCENDING',name=None)
#loss = tf.convert_to_tensor(np.zeros([1]),dtype=tf. float32)
loss = tf.convert_to_tensor(np.array([]),dtype=tf. float32)
for i in range(bs):
arr_nonzero=[]
nonzero=0
arr_pred = []
t_pred = tf.convert_to_tensor(np.array([0]),dtype=tf. float32)
t_nonzero = tf.convert_to_tensor(np.array([0]),dtype=tf. float32)
for k in tp_mul[i]:
if k>0:
arr_nonzero.append(k)
nonzero+=1
t_nonzero = t_nonzero + k
for k in y_pred_sort[i][:nonzero]:
arr_pred.append(k)
t_pred = t_pred + k
cor_num=0.
total_num=0.
if t_pred==t_nonzero:
cor_num=1.
total_num=1.
else:
cor_num=0.
total_num=1.
alp = 2-cor_num/total_num
#loss+=alp*lss_fn(y_true[i,:],y_pred[i,:])
loss = tf.experimental.numpy.append(loss, alp*lss_fn(y_true[i,:],y_pred[i,:]), axis=None)
loss = tf.reduce_mean(loss)
return loss
# +
def onehot(image,label):
CLASSES = 19
return image,tf.one_hot(label,CLASSES)
def cutmix(image, label, PROBABILITY = 1.0):
# input image - is a batch of images of size [n,dim,dim,3] not a single image of [dim,dim,3]
# output - a batch of images with cutmix applied
DIM = IMSIZE[IMS]
CLASSES = 19
imgs = []; labs = []
for j in range(AUG_BATCH):
# DO CUTMIX WITH PROBABILITY DEFINED ABOVE
P = tf.cast( tf.random.uniform([],0,1)<=PROBABILITY, tf.int32)
# CHOOSE RANDOM IMAGE TO CUTMIX WITH
k = tf.cast( tf.random.uniform([],0,AUG_BATCH),tf.int32)
# CHOOSE RANDOM LOCATION
x = tf.cast( tf.random.uniform([],0,DIM),tf.int32)
y = tf.cast( tf.random.uniform([],0,DIM),tf.int32)
b = tf.random.uniform([],0,1) # this is beta dist with alpha=1.0
WIDTH = tf.cast( DIM * tf.math.sqrt(1-b),tf.int32) * P
ya = tf.math.maximum(0,y-WIDTH//2)
yb = tf.math.minimum(DIM,y+WIDTH//2)
xa = tf.math.maximum(0,x-WIDTH//2)
xb = tf.math.minimum(DIM,x+WIDTH//2)
# MAKE CUTMIX IMAGE
one = image[j,ya:yb,0:xa,:]
two = image[k,ya:yb,xa:xb,:]
three = image[j,ya:yb,xb:DIM,:]
middle = tf.concat([one,two,three],axis=1)
img = tf.concat([image[j,0:ya,:,:],middle,image[j,yb:DIM,:,:]],axis=0)
imgs.append(img)
# MAKE CUTMIX LABEL
a = tf.cast(WIDTH*WIDTH/DIM/DIM,tf.float32)
if len(label.shape)==1:
lab1 = tf.one_hot(label[j],CLASSES)
lab2 = tf.one_hot(label[k],CLASSES)
else:
lab1 = label[j,]
lab2 = label[k,]
lab1 = tf.cast(lab1,tf.float32)
lab2 = tf.cast(lab2,tf.float32)
labs.append((1-a)*lab1 + a*lab2)
# RESHAPE HACK SO TPU COMPILER KNOWS SHAPE OF OUTPUT TENSOR (maybe use Python typing instead?)
image2 = tf.reshape(tf.stack(imgs),(AUG_BATCH,DIM,DIM,3))
label2 = tf.reshape(tf.stack(labs),(AUG_BATCH,CLASSES))
return image2,label2
def mixup(image, label, PROBABILITY = 1.0):
# input image - is a batch of images of size [n,dim,dim,3] not a single image of [dim,dim,3]
# output - a batch of images with mixup applied
DIM = IMSIZE[IMS]
CLASSES = 19
imgs = []; labs = []
for j in range(AUG_BATCH):
# DO MIXUP WITH PROBABILITY DEFINED ABOVE
P = tf.cast( tf.random.uniform([],0,1)<=PROBABILITY, tf.float32)
# CHOOSE RANDOM
k = tf.cast( tf.random.uniform([],0,AUG_BATCH),tf.int32)
a = tf.random.uniform([],0,1)*P # this is beta dist with alpha=1.0
# MAKE MIXUP IMAGE
img1 = image[j,]
img2 = image[k,]
imgs.append((1-a)*img1 + a*img2)
# MAKE CUTMIX LABEL
if len(label.shape)==1:
lab1 = tf.one_hot(label[j],CLASSES)
lab2 = tf.one_hot(label[k],CLASSES)
else:
lab1 = label[j,]
lab2 = label[k,]
lab1 = tf.cast(lab1,tf.float32)
lab2 = tf.cast(lab2,tf.float32)
labs.append((1-a)*lab1 + a*lab2)
# RESHAPE HACK SO TPU COMPILER KNOWS SHAPE OF OUTPUT TENSOR (maybe use Python typing instead?)
image2 = tf.reshape(tf.stack(imgs),(AUG_BATCH,DIM,DIM,3))
label2 = tf.reshape(tf.stack(labs),(AUG_BATCH,CLASSES))
return image2,label2
def transform(image,label):
# THIS FUNCTION APPLIES BOTH CUTMIX AND MIXUP
DIM = IMSIZE[IMS]
CLASSES = 19
SWITCH = 0.5
CUTMIX_PROB = 0.666
MIXUP_PROB = 0.666
# FOR SWITCH PERCENT OF TIME WE DO CUTMIX AND (1-SWITCH) WE DO MIXUP
image2, label2 = cutmix(image, label, CUTMIX_PROB)
image3, label3 = mixup(image, label, MIXUP_PROB)
imgs = []; labs = []
for j in range(AUG_BATCH):
P = tf.cast( tf.random.uniform([],0,1)<=SWITCH, tf.float32)
imgs.append(P*image2[j,]+(1-P)*image3[j,])
labs.append(P*label2[j,]+(1-P)*label3[j,])
# RESHAPE HACK SO TPU COMPILER KNOWS SHAPE OF OUTPUT TENSOR (maybe use Python typing instead?)
image4 = tf.reshape(tf.stack(imgs),(AUG_BATCH,DIM,DIM,3))
label4 = tf.reshape(tf.stack(labs),(AUG_BATCH,CLASSES))
return image4,label4
# +
def auto_select_accelerator():
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver()
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)
strategy = tf.distribute.experimental.TPUStrategy(tpu)
print("Running on TPU:", tpu.master())
except ValueError:
strategy = tf.distribute.get_strategy()
print(f"Running on {strategy.num_replicas_in_sync} replicas")
return strategy
def build_decoder(with_labels=True, target_size=(256, 256), ext='jpg'):
def decode(path):
file_bytes = tf.io.read_file(path)
if ext == 'png':
img = tf.image.decode_png(file_bytes, channels=3)
elif ext in ['jpg', 'jpeg']:
img = tf.image.decode_jpeg(file_bytes, channels=3)
else:
raise ValueError("Image extension not supported")
img = tf.cast(img, tf.float32) / 255.0
img = tf.image.resize(img, target_size)
return img
def decode_with_labels(path, label):
return decode(path), label
return decode_with_labels if with_labels else decode
def build_augmenter(with_labels=True):
def augment(img):
img = tf.image.random_flip_left_right(img)
img = tf.image.random_flip_up_down(img)
return img
def augment_with_labels(img, label):
return augment(img), label
return augment_with_labels if with_labels else augment
DIM =600
n_class = 19
def _parse_image_function(example_proto,augment = True):
image_feature_description = {
'image': tf.io.FixedLenFeature([], tf.string),
'label': tf.io.FixedLenFeature([], tf.string)
}
single_example = tf.io.parse_single_example(example_proto, image_feature_description)
image = tf.reshape( tf.io.decode_raw(single_example['image'],out_type=np.dtype('uint8')), (DIM,DIM, 3))
mask = tf.reshape(tf.io.decode_raw(single_example['label'],out_type=np.dtype('uint8')),[n_class])
image = tf.dtypes.cast(image, tf.float32)
mask = tf.dtypes.cast(mask, tf.float32)
image = image/255.
if augment: # https://www.kaggle.com/kool777/training-hubmap-eda-tf-keras-tpu
if tf.random.uniform(()) > 0.5:
image = tf.image.flip_left_right(image)
mask = tf.image.flip_left_right(mask)
if tf.random.uniform(()) > 0.4:
image = tf.image.flip_up_down(image)
mask = tf.image.flip_up_down(mask)
if tf.random.uniform(()) > 0.5:
image = tf.image.rot90(image, k=1)
mask = tf.image.rot90(mask, k=1)
if tf.random.uniform(()) > 0.45:
image = tf.image.random_saturation(image, 0.7, 1.3)
if tf.random.uniform(()) > 0.45:
image = tf.image.random_contrast(image, 0.8, 1.2)
return tf.cast(image, tf.float32),tf.cast(mask, tf.float32)
def load_dataset(filenames, ordered=False, augment = False):
AUTO = tf.data.experimental.AUTOTUNE
ignore_order = tf.data.Options()
if not ordered:
ignore_order.experimental_deterministic = False
dataset = tf.data.TFRecordDataset(filenames, num_parallel_reads=AUTO)
dataset = dataset.with_options(ignore_order)
dataset = dataset.map(lambda ex: _parse_image_function(ex, augment = augment), num_parallel_calls=AUTO)
return dataset
def build_dataset(paths, labels=None, bsize=128, cache=True,
decode_fn=None, augment_fn=None,
augment=True,augment_mixup_cutmix=False
, repeat=True, shuffle=1024,
cache_dir=""):
if cache_dir != "" and cache is True:
os.makedirs(cache_dir, exist_ok=True)
if decode_fn is None:
decode_fn = build_decoder(labels is not None)
if augment_fn is None:
augment_fn = build_augmenter(True)
AUTO = tf.data.experimental.AUTOTUNE
#slices = paths if labels is None else (paths, labels)
dset = load_dataset(paths)
#dset = tf.data.Dataset.from_tensor_slices(slices)
#dset = dset.map(decode_fn, num_parallel_calls=AUTO)
dset = dset.cache(cache_dir) if cache else dset
#dset = dset.map(transform, num_parallel_calls=AUTO) if augment else dset
dset = dset.map(augment_fn, num_parallel_calls=AUTO) if augment else dset
dset = dset.repeat() if repeat else dset
dset = dset.shuffle(shuffle) if shuffle else dset
dset = dset.batch(bsize)
dset = dset.map(transform,num_parallel_calls=AUTO) if augment_mixup_cutmix else dset
dset = dset.prefetch(AUTO)
return dset
# -
def count_data_items(filenames):
img_num = 0
for i in filenames:
img_num +=int(i.split('_')[-2])
return img_num
# +
strategy = auto_select_accelerator()
BATCH_SIZE = strategy.num_replicas_in_sync * 16
AUG_BATCH = BATCH_SIZE
# -
# + _kg_hide-output=true
# -
train_list=[]
valid_list=[]
# +
gs_csv = pd.read_csv('../input/hpagscsv/gs.csv')
val_index=[0] # decide which dataset to be the valid one
for i in gs_csv.index:
temp=gs_csv.loc[i,'URL']
if i not in val_index:
train_list.extend(tf.io.gfile.glob(i+'/*'))
else:
valid_list.extend(tf.io.gfile.glob(i+'/*'))
# 'gs://kds-db891f03f8787fad574c283c8f45d40c0072ebcdc9e44f8b821c3788' hpatrain-green-imagetfrec
#'gs://kds-38d7c7971dfd017bc732339d9814b3cdf2211a934a6c1423e5e7a80c' hpavalid-green-imagetfrec
#'gs://kds-3867054d379fba84653fcf956ef637375f645217ab4358a185785aaf' hpatrain-ex08-green-imagetfrec
#'gs://kds-45d6c0aafb360b188d1a74670585227dbcc2847490f9dec13e879934' hpatrain-ex-916-green-imagetfrec
#'gs://kds-47b27aa31455ce4713b8c540ab3bb52d07f01b5fe2bf4be64c673821' hpatrain-ex-1724-green-imagetfrec
#'gs://kds-833ca2c0d1e0e67d9c5b3ad7c0da7c3235f418ca65031f5e6df34fc3' hpatrain-ex-2532-green-imagetfrec
# 'gs://kds-ee5eb0076b7380bde326f8d748a8103f22ed7b4f309d43ddaadb5049' hpatrain-ex-3338-green-imagetfrec
# -
# +
IMSIZE = (224, 240, 260, 300, 380, 456, 528, 600)
IMS = 7
decoder = build_decoder(with_labels=True, target_size=(IMSIZE[IMS], IMSIZE[IMS]))
test_decoder = build_decoder(with_labels=False, target_size=(IMSIZE[IMS], IMSIZE[IMS]))
train_dataset = build_dataset(
train_paths, bsize=BATCH_SIZE,cache=False, decode_fn=decoder,augment_mixup_cutmix=True
)
valid_dataset = build_dataset(
valid_paths, bsize=BATCH_SIZE, decode_fn=decoder,
repeat=False, shuffle=False,cache=False, augment=False
)
# -
# +
try:
n_labels = 19
except:
n_labels = 1
with strategy.scope():
model = tf.keras.Sequential([
efn.EfficientNetB7(
input_shape=(IMSIZE[IMS], IMSIZE[IMS], 3),
weights='noisy-student',
include_top=False),
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(n_labels, activation='sigmoid')
])
model.compile(
optimizer=tfa.optimizers.Lookahead(tfa.optimizers.RectifiedAdam()),#tf.keras.optimizers.Adam(),
loss=tfa.losses.SigmoidFocalCrossEntropy(),#'binary_crossentropy',#tfa.losses.SigmoidFocalCrossEntropy(),
metrics=[tf.keras.metrics.AUC(multi_label=True)])
model.summary()
# -
colour = '_green'
#steps_per_epoch = train_paths.shape[0] // BATCH_SIZE
steps_per_epoch = count_data_items(train_paths) // BATCH_SIZE
checkpoint = tf.keras.callbacks.ModelCheckpoint(
f'model{colour}.h5', save_best_only=True, monitor='val_loss', mode='min')
lr_reducer = tf.keras.callbacks.ReduceLROnPlateau(
monitor="val_loss", patience=3, min_lr=1e-6, mode='min')
history = model.fit(
train_dataset,
epochs=30,
verbose=1,
callbacks=[checkpoint, lr_reducer],
steps_per_epoch=steps_per_epoch,
validation_data=valid_dataset)
hist_df = pd.DataFrame(history.history)
hist_df.to_csv(f'history{colour}.csv')
|
train/hpa-image-model-training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: p_profile
# language: python
# name: p_profile
# ---
import numpy as np
import pandas as pd
from pandas_profiling import ProfileReport
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
raw_data = pd.read_excel("data/Telco_customer_churn.xlsx")
raw_data.shape
raw_data.head().T
raw_data.info()
profile = ProfileReport(raw_data, title="Pandas Profiling Report", explorative=True)
# +
#profile.to_file("profiling_report.html")
# -
raw_data.describe()
# ## Check for missing values
raw_data.isnull().sum()
raw_data[raw_data['Churn Reason'].isnull()].head()
raw_data.loc[raw_data['Churn Reason'].isnull(), 'Churn Label'].unique()
# From above analysis, we see that only the column `Churn Reason` has missing values. Further examination of that column reveals that the values are missing only for those observations with `Churn Label` = No. Thus it represents the customers that are still with the company and it definitely makes sense for those customers to have null value for `Churn Reason` column.
#
# Hence it is concluded that no other treatement of null values are needed for this dataset.
# ## Check for duplicate values and outliers
len(raw_data['CustomerID'].unique())
raw_data.shape[0]
raw_data['Country'].unique()
raw_data['State'].unique()
len(raw_data['Zip Code'].unique())
raw_data['Zip Code'].min()
raw_data['Zip Code'].max()
# The above analysis shows that our data set include 7043 observations about unique customers from the California state of United States. This includes customers from various postal codes ranging between 90001 and 96161. Also, all the observations have valid data for columns `Country`, `State` and `Zip Code`.
raw_data[['Latitude', 'Longitude']].describe()
# The above statistics verifies that there are no outliers or typos in the `Latitude` and `Longitude` columns. The column `Lat Long` contains the same data in a combined form as a categorical variable.
# ### Analysis of binary variables
fig, axes = plt.subplots(2,2, figsize=(14,8))
sns.countplot(ax=axes[0,0], x='Gender', data=raw_data)
sns.countplot(ax=axes[0,1], x='Senior Citizen', data=raw_data)
sns.countplot(ax=axes[1,0], x='Partner', data=raw_data)
sns.countplot(ax=axes[1,1], x='Dependents', data=raw_data)
fig.suptitle('Check for outliers in binary variables')
# ### Analysis of categorical variables
fig, axes = plt.subplots(2,2, figsize=(14,8))
sns.countplot(ax=axes[0,0], x='Multiple Lines', data=raw_data)
sns.countplot(ax=axes[0,1], x='Internet Service', data=raw_data)
sns.countplot(ax=axes[1,0], x='Online Security', data=raw_data)
sns.countplot(ax=axes[1,1], x='Online Backup', data=raw_data)
fig, axes = plt.subplots(2,2, figsize=(14,8))
sns.countplot(ax=axes[0,0], x='Device Protection', data=raw_data)
sns.countplot(ax=axes[0,1], x='Tech Support', data=raw_data)
sns.countplot(ax=axes[1,0], x='Streaming TV', data=raw_data)
sns.countplot(ax=axes[1,1], x='Streaming Movies', data=raw_data)
sns.set()
raw_data[['Tenure Months', 'Monthly Charges', 'Churn Score', 'CLTV']].hist(figsize=(15, 10))
plt.subplots_adjust(hspace=0.5);
# ## Analysis of target variable
p = sns.catplot(x='Churn Value', y='Churn Score', data=raw_data, kind='box')
plt.show()
# +
fig, ax = plt.subplots(figsize=(8,5))
ax.hist(raw_data.loc[raw_data['Churn Value']==0, 'Churn Score'], histtype='step', label='churn = 0')
ax.hist(raw_data.loc[raw_data['Churn Value']==1, 'Churn Score'], histtype='step', label='churn = 1')
ax.set_ylabel("# of churns")
ax.legend()
# -
target_counts = raw_data['Churn Label'].value_counts(normalize=True)
target_counts = target_counts.rename('percent').reset_index()
target_counts['percent'] = target_counts['percent'] * 100
# +
g = sns.catplot(x='index', y='percent', data=target_counts, kind='bar')
for p in g.ax.patches:
txt = str(p.get_height().round(2)) + '%'
txt_x = p.get_x() + 0.3
txt_y = p.get_height() + 0.2
g.ax.text(txt_x,txt_y,txt)
plt.xlabel('Customer churn value')
plt.title('Distribution of target variable')
# -
target_counts
# ### Summary
#
# * The dataset includes 7043 observations about telecommunication customers from California. Out of the 7043 rows, 27% of the customers left the company in the end of Q3
# * No missing values in any columns except `Churn Reason`. This column has a value only for those customers who left the company.
#
#
# **Numerical columns**
# * Tenure Months, Monthly Charges, Churn Score, CLTV
# * No missing values or outliers in any of these columns
#
# **categorical columns**
# * Multiple Lines, Internet Service, Online Security, Online Backup, Device Protection, Tech Support, Streaming TV, Streaming Movies, Contract, Payment Method,
#
# **boolean columns**
# * Gender, Senior Citizen, Partner, Dependents, Phone Service, Paperless Billing, Churn Label, Churn Value
#
# **others**
# * City, Churn Reason
#
# Careful analysis of dataset verifies that dataset is clean and there are no outliers. Also, all the columns have the correct data type.
|
old/note books/CAP2_wrangling_v1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Cleaning Quiz: Udacity's Course Catalog
# It's your turn! Udacity's course catalog page has changed since the last video was filmed. One notable change is the introduction of schools.
#
# In this activity, you're going to perform similar actions with BeautifulSoup to extract the following information from each course listing on the page:
#
# The course name - e.g. "Data Analyst"
# The school the course belongs to - e.g. "School of Data Science"
# Note: All solution notebooks can be found by clicking on the Jupyter icon on the top left of this workspace.
# Step 1: Get text from Udacity's course catalog web page
# You can use the requests library to do this.
#
# Outputting all the javascript, CSS, and text may overload the space available to load this notebook, so we omit a print statement here.
# import statements
import requests
from bs4 import BeautifulSoup
# fetch web page
r=requests.get("https://www.udacity.com/courses/all")
# Step 2: Use BeautifulSoup to remove HTML tags
# Use "lxml" rather than "html5lib".
#
# Again, printing this entire result may overload the space available to load this notebook, so we omit a print statement here.
soup= BeautifulSoup(r.text, "lxml")
# Step 3: Find all course summaries
# Use the BeautifulSoup's find_all method to select based on tag type and class name. Just ike in the video, you can right click on the item, and click "Inspect" to view its html on a web page.
summaries = soup.find_all("li", {"class":"catalog-cards__list__item"} )
print('Number of Courses:', len(summaries))
# Step 4: Inspect the first summary to find selectors for the course name and school
# Tip: .prettify() is a super helpful method BeautifulSoup provides to output html in a nicely indented form! Make sure to use print() to ensure whitespace is displayed properly.
print(summaries[0].prettify())
# Look for selectors contain the the courses title and school name text you want to extract. Then, use the select_one method on the summary object to pull out the html with those selectors. Afterwards, don't forget to do some extra cleaning to isolate the names (get rid of unnecessary html), as you saw in the last video.
# Extract course title
summaries[0].select_one("h2").get_text().strip()
# Extract school
summaries[0].select_one("h3").get_text().strip()
# Step 5: Collect names and schools of ALL course listings¶
# Reuse your code from the previous step, but now in a loop to extract the name and school from every course summary in summaries!
courses = []
for summary in summaries:
# append name and school of each summary to courses list
title = summary.select_one("h2").get_text().strip()
school = summary.select_one("h3").get_text().strip()
courses.append((title, school))
# display results
print(len(courses), "course summaries found. Sample:")
courses[:20]
|
Pipelines/NLPpipelines/.ipynb_checkpoints/cleaning_practise-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: DESI master
# language: python
# name: desi-master
# ---
import os
import sv as SV
import numpy as np
import astropy.table as atable
from tqdm.notebook import tqdm, trange
from desitarget import cuts as desi_cuts
# -- plotting --
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams['text.usetex'] = True
mpl.rcParams['font.family'] = 'serif'
mpl.rcParams['axes.linewidth'] = 1.5
mpl.rcParams['axes.xmargin'] = 1
mpl.rcParams['xtick.labelsize'] = 'x-large'
mpl.rcParams['xtick.major.size'] = 5
mpl.rcParams['xtick.major.width'] = 1.5
mpl.rcParams['ytick.labelsize'] = 'x-large'
mpl.rcParams['ytick.major.size'] = 5
mpl.rcParams['ytick.major.width'] = 1.5
mpl.rcParams['legend.frameon'] = False
# !pip install corner --user
import corner as DFM
# +
ls_S_dir = '/global/cfs/cdirs/cosmo/data/legacysurvey/dr9/south/sweep/9.0'
ls_N_dir = '/global/cfs/cdirs/cosmo/data/legacysurvey/dr9/north/sweep/9.0'
#ls_S_dir = '/Users/chahah/data/bgs_cmxsv/sv_paper'
#ls_N_dir = '/Users/chahah/data/bgs_cmxsv/sv_paper'
cat_S = atable.Table.read(os.path.join(ls_S_dir, 'sweep-200p005-210p010.fits'))
cat_N = atable.Table.read(os.path.join(ls_N_dir, 'sweep-190p040-200p045.fits'))
# -
cat_S['PHOTSYS'] = 'S'
cat_N['PHOTSYS'] = 'N'
def BGS_quality_mask(targets, south=True):
''' apply BGS quality cut and imaging mask
'''
photsys_north, photsys_south, obs_rflux, gflux, rflux, zflux, \
w1flux, w2flux, gfiberflux, rfiberflux, zfiberflux, \
gfibertotflux, rfibertotflux, zfibertotflux, \
objtype, release, ra, dec, gfluxivar, rfluxivar, zfluxivar, w1fluxivar, \
gnobs, rnobs, znobs, gfracflux, rfracflux, zfracflux, \
gfracmasked, rfracmasked, zfracmasked, \
gfracin, rfracin, zfracin, gallmask, rallmask, zallmask, \
gsnr, rsnr, zsnr, w1snr, w2snr, dchisq, deltaChi2, maskbits, refcat = desi_cuts._prepare_optical_wise(objects=targets, mask=True)
gaia, pmra, pmdec, parallax, parallaxovererror, parallaxerr, gaiagmag, \
gaiabmag, gaiarmag, gaiaaen, gaiadupsource, Grr, gaiaparamssolved, \
gaiabprpfactor, gaiasigma5dmax, galb = desi_cuts._prepare_gaia(objects=targets, colnames=None)
bgs = np.ones_like(gnobs, dtype='?')
bgs &= (gnobs >= 1) & (rnobs >= 1) & (znobs >= 1)
bgs &= (gfluxivar > 0) & (rfluxivar > 0) & (zfluxivar > 0)
# ADM geometric masking cuts from the Legacy Surveys.
# BRIGHT & CLUSTER for BGS
bgs &= desi_cuts.imaging_mask(maskbits, bgsmask=True)
return bgs
qm_S = BGS_quality_mask(cat_S, south=True)
qm_N = BGS_quality_mask(cat_N, south=True)
# +
optw_S = desi_cuts._prepare_optical_wise(objects=cat_S, mask=True)
optw_N = desi_cuts._prepare_optical_wise(objects=cat_N, mask=True)
gaia_S = desi_cuts._prepare_gaia(objects=cat_S, colnames=None)
gaia_N = desi_cuts._prepare_gaia(objects=cat_N, colnames=None)
# +
rfib_S = 22.5 - 2.5 * np.log10(optw_S[9].clip(1e-16))
gmag_S = 22.5 - 2.5 * np.log10(optw_S[3].clip(1e-16))
rmag_S = 22.5 - 2.5 * np.log10(optw_S[4].clip(1e-16))
zmag_S = 22.5 - 2.5 * np.log10(optw_S[5].clip(1e-16))
w1_S = 22.5 - 2.5 * np.log10(optw_S[6].clip(1e-16))
G_rr_S = gaia_S[11]
rfib_N = 22.5 - 2.5 * np.log10(optw_N[9].clip(1e-16))
gmag_N = 22.5 - 2.5 * np.log10(optw_N[3].clip(1e-16))
rmag_N = 22.5 - 2.5 * np.log10(optw_N[4].clip(1e-16))
zmag_N = 22.5 - 2.5 * np.log10(optw_N[5].clip(1e-16))
w1_N = 22.5 - 2.5 * np.log10(optw_N[6].clip(1e-16))
G_rr_N = gaia_N[11]
# -
def BGS_sample(targets, south=True, targtype='bright'):
''' apply BGS quality cut and imaging mask
'''
photsys_north, photsys_south, obs_rflux, gflux, rflux, zflux, \
w1flux, w2flux, gfiberflux, rfiberflux, zfiberflux, \
gfibertotflux, rfibertotflux, zfibertotflux, \
objtype, release, ra, dec, gfluxivar, rfluxivar, zfluxivar, w1fluxivar, \
gnobs, rnobs, znobs, gfracflux, rfracflux, zfracflux, \
gfracmasked, rfracmasked, zfracmasked, \
gfracin, rfracin, zfracin, gallmask, rallmask, zallmask, \
gsnr, rsnr, zsnr, w1snr, w2snr, dchisq, deltaChi2, maskbits, refcat = desi_cuts._prepare_optical_wise(objects=targets, mask=True)
gaia, pmra, pmdec, parallax, parallaxovererror, parallaxerr, gaiagmag, \
gaiabmag, gaiarmag, gaiaaen, gaiadupsource, Grr, gaiaparamssolved, \
gaiabprpfactor, gaiasigma5dmax, galb = desi_cuts._prepare_gaia(objects=targets, colnames=None)
bgs = desi_cuts.isBGS(
gflux=gflux, rflux=rflux, zflux=zflux, w1flux=w1flux, w2flux=w2flux, rfibertotflux=rfibertotflux,
rfiberflux=rfiberflux, gnobs=gnobs, rnobs=rnobs, znobs=znobs,
gfluxivar=gfluxivar, rfluxivar=rfluxivar, zfluxivar=zfluxivar,
maskbits=maskbits, Grr=Grr, refcat=refcat, w1snr=w1snr, w2snr=w2snr, gaiagmag=gaiagmag,
objtype=objtype, primary=None, south=south, targtype=targtype)
return bgs
bgs_bright_S = BGS_sample(cat_S, south=True, targtype='bright')
bgs_bright_N = BGS_sample(cat_N, south=False, targtype='bright')
bgs_faint_S = BGS_sample(cat_S, south=True, targtype='faint')
bgs_faint_N = BGS_sample(cat_N, south=False, targtype='faint')
# +
fig = plt.figure(figsize=(15,6))
sub = fig.add_subplot(121)
DFM.hist2d(np.concatenate([rmag_S, rmag_N]), np.concatenate([rfib_S, rfib_N]),
range=[(14, 21), (16, 25)], color='k',
plot_density=False, plot_datapoints=True, no_fill_contours=True,
bins=200, smooth=True,
contour_kwargs={'linewidths': 1, 'linestyles': ':'},
ax=sub)
sub.scatter(rmag_S, rfib_S, c='gray', s=0.1, rasterized=True)
sub.scatter(rmag_N, rfib_N, c='gray', s=0.1, rasterized=True)
sub.text(0.05, 0.95, r'Legacy Survey', ha='left', va='top', transform=sub.transAxes, fontsize=30)
sub.text(14.5, 20.1, 'fiber-magnitude cut', rotation=35, fontsize=18)
sub.plot([14, 17.8], [19.1, 22.9], c='k', ls='--', lw=3)
sub.plot([17.8, 19.5], [22.9, 22.9], c='k', ls='--', lw=3)
sub.text(19.55, 17., '$r < 19.5$', rotation=270, fontsize=18)
sub.plot([19.5, 19.5], [16., 22.9], c='k', ls='--', lw=3)
sub.set_xlim(14, 21)
sub.set_ylim(16, 25)
sub.set_yticks([16, 18, 20, 22, 24])
sub = fig.add_subplot(122)
hb = sub.hexbin(rmag_S[bgs_bright_S], rfib_S[bgs_bright_S],
C=np.ones(np.sum(bgs_bright_S))/47,
reduce_C_function=np.sum, extent=(14, 20, 16, 24), gridsize=30,
edgecolors='k', linewidth=1., mincnt=2, cmap='Spectral_r')
sub.text(0.05, 0.95, r'BGS Bright', ha='left', va='top', transform=sub.transAxes, fontsize=30)
sub.plot([14, 17.8], [19.1, 22.9], c='k', ls='--', lw=3)
sub.plot([17.8, 19.5], [22.9, 22.9], c='k', ls='--', lw=3)
sub.plot([19.5, 19.5], [16., 22.9], c='k', ls='--', lw=3)
sub.set_xlim(14, 21)
sub.set_ylim(16, 25)
sub.set_yticks([16, 18, 20, 22, 24])
sub.set_yticklabels([])
bkgd = fig.add_subplot(111, frameon=False)
bkgd.set_xlabel(r'$r$ magnitude', labelpad=10, fontsize=25)
bkgd.set_ylabel(r'$r_{\rm fiber}$ magnitude', labelpad=10, fontsize=25)
bkgd.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
fig.subplots_adjust(wspace=0.1, right=0.8)
cbar_ax = fig.add_axes([0.85, 0.15, 0.02, 0.7])
cbar = fig.colorbar(hb, cax=cbar_ax)
cbar.ax.set_ylabel(r'target density [${\rm deg}^{-2}$]', labelpad=30, fontsize=25, rotation=270)
fig.savefig('figs/bgs_bright.pdf', bbox_inches='tight')
# -
fibcol_S = (zmag_S - w1_S) - 3/2.5 * (gmag_S - rmag_S) + 1.2
fibcol_N = (zmag_N - w1_N) - 3/2.5 * (gmag_N - rmag_N) + 1.2
# +
fig = plt.figure(figsize=(15,6))
sub = fig.add_subplot(121)
sub.scatter(rfib_S, fibcol_S, c='gray', s=0.1, rasterized=True)
sub.scatter(rfib_N - 0.04, fibcol_N, c='gray', s=0.1, rasterized=True)
DFM.hist2d(np.concatenate([rfib_S, rfib_N - 0.04]), np.concatenate([fibcol_S, fibcol_N]),
range=[(19.0, 22.5), (-1., 1.8)], color='k',
plot_density=False, plot_datapoints=True, no_fill_contours=True,
smooth=True,
contour_kwargs={'linewidths': 1, 'linestyles': ':'},
ax=sub)
sub.scatter([], [], c='gray', s=100, label='Legacy Survey')
sub.plot([20.79, 20.79], [-2, 0.], c='k', ls='--', lw=3)
sub.plot([20.79, 21.54], [0., 0.], c='k', ls='--', lw=3)
sub.plot([21.54, 21.54], [0., 2.], c='k', ls='--', lw=3)
sub.text(0.05, 0.95, r'Legacy Survey', ha='left', va='top', transform=sub.transAxes, fontsize=30)
sub.fill_between([19.2, 20.9], [1.4, 1.4], [1.7, 1.7], color='white', alpha=0.75)
sub.set_yticks([-1., -0.5, 0., 0.5, 1., 1.5])
sub.set_ylim(-1., 1.8)
#sub.set_xlabel(r'$r_{\rm fiber}$ magnitude', fontsize=25)
sub.set_xlim(19.0, 22.5)
sub.set_xticks([19., 20, 21, 22])
sub = fig.add_subplot(122)
hb = sub.hexbin(rfib_S[bgs_faint_S], fibcol_S[bgs_faint_S], C=(np.ones(len(bgs_faint_S))/47).astype(float),
reduce_C_function=np.sum, extent=(19., 22.5, -1., 1.5), gridsize=30, mincnt=5,
cmap='Spectral_r', edgecolors='k', linewidth=1.)
sub.plot([20.79, 20.79], [-2, 0.], c='k', ls='--', lw=3)
sub.plot([20.79, 21.54], [0., 0.], c='k', ls='--', lw=3)
sub.plot([21.54, 21.54], [0., 2.], c='k', ls='--', lw=3)
#sub.fill_between([19.2, 20.5], [1.3, 1.3], [1.7, 1.7], color='white', alpha=0.75)
#sub.legend(loc='upper left', bbox_to_anchor=(-0.035, 1.025), fontsize=20, handletextpad=-0.4)
sub.text(0.05, 0.95, r'BGS Faint', ha='left', va='top', transform=sub.transAxes, fontsize=30)
sub.set_yticks([-1., -0.5, 0., 0.5, 1., 1.5])
sub.set_ylim(-1., 1.8)
sub.set_xlim(19.0, 22.5)
sub.set_xticks([19., 20, 21, 22])
sub.set_yticklabels([])
bkgd = fig.add_subplot(111, frameon=False)
bkgd.set_xlabel(r'$r_{\rm fiber}$ magnitude', labelpad=10, fontsize=25)
bkgd.set_ylabel(r'$(z - w1) - 1.2 (g - r) + 1.2$', labelpad=10, fontsize=25)
bkgd.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
fig.subplots_adjust(wspace=0.1, right=0.8)
cbar_ax = fig.add_axes([0.85, 0.15, 0.02, 0.7])
cbar = fig.colorbar(hb, cax=cbar_ax)
cbar.ax.set_ylabel(r'target density [${\rm deg}^{-2}$]', labelpad=30, fontsize=25, rotation=270)
fig.savefig('figs/bgs_faint.pdf', bbox_inches='tight')
# +
# load compiled zsuccess rates to file
dat_dir = '/global/cscratch1/sd/chahah/bgs-cmxsv/sv-paper/'
#dat_dir = '/Users/chahah/data/bgs_cmxsv/sv_paper/'
sv1_gals = atable.Table.read(os.path.join(dat_dir, 'sv1.bgs_exps.efftime160_200.zsuccess.fits'))
sv3_gals = atable.Table.read(os.path.join(dat_dir, 'sv3.bgs_exps.efftime160_200.zsuccess.fits'))
main_gals = atable.Table.read(os.path.join(dat_dir, 'main.bgs_exps.efftime160_200.zsuccess.fits'))
# +
from desiutil.dust import ext_odonnell
sv1_gals = sv1_gals[(sv1_gals['PHOTSYS'] != '') & (sv1_gals['PHOTSYS'] != 'G')]
sv3_gals = sv3_gals[sv3_gals['PHOTSYS'] != '']
from speclite import filters
decamwise = filters.load_filters('decam2014-g', 'decam2014-r','decam2014-z', 'wise2010-W1', 'wise2010-W2')
bassmzlswise = filters.load_filters('BASS-g', 'BASS-r', 'MzLS-z','wise2010-W1', 'wise2010-W2')
RV = 3.1
sv1_mw_trans = np.array([10**(-0.4 * sv1_gals['EBV'][i] * RV * ext_odonnell(bassmzlswise.effective_wavelengths.value, Rv=RV)) if sv1_gals['PHOTSYS'][i] == 'N'
else 10**(-0.4 * sv1_gals['EBV'][i] * RV * ext_odonnell(decamwise.effective_wavelengths.value, Rv=RV)) for i in range(len(sv1_gals['EBV']))])
sv3_mw_trans = np.array([10**(-0.4 * sv3_gals['EBV'][i] * RV * ext_odonnell(bassmzlswise.effective_wavelengths.value, Rv=RV)) if sv3_gals['PHOTSYS'][i] == 'N'
else 10**(-0.4 * sv3_gals['EBV'][i] * RV * ext_odonnell(decamwise.effective_wavelengths.value, Rv=RV)) for i in range(len(sv3_gals['EBV']))])
# +
sv1_gmag = 22.5 - 2.5 * np.log10((sv1_gals['FLUX_G']/sv1_mw_trans[:,0]).clip(1e-16))
sv1_rmag = 22.5 - 2.5 * np.log10((sv1_gals['FLUX_R']/sv1_mw_trans[:,1]).clip(1e-16))
sv1_zmag = 22.5 - 2.5 * np.log10((sv1_gals['FLUX_Z']/sv1_mw_trans[:,2]).clip(1e-16))
sv1_rfib = 22.5 - 2.5 * np.log10((sv1_gals['FIBERFLUX_R']/sv1_mw_trans[:,1]).clip(1e-16))
sv1_w1 = 22.5 - 2.5 * np.log10((sv1_gals['FLUX_W1']/sv1_mw_trans[:,3]).clip(1e-16))
sv1_g_rr = sv1_gals['GAIA_PHOT_G_MEAN_MAG'] - 22.5 + 2.5*np.log10(sv1_gals['FLUX_R'])
sv3_gmag = 22.5 - 2.5 * np.log10((sv3_gals['FLUX_G']/sv3_mw_trans[:,0]).clip(1e-16))
sv3_rmag = 22.5 - 2.5 * np.log10((sv3_gals['FLUX_R']/sv3_mw_trans[:,1]).clip(1e-16))
sv3_zmag = 22.5 - 2.5 * np.log10((sv3_gals['FLUX_Z']/sv3_mw_trans[:,2]).clip(1e-16))
sv3_rfib = 22.5 - 2.5 * np.log10((sv3_gals['FIBERFLUX_R']/sv3_mw_trans[:,1]).clip(1e-16))
sv3_w1 = 22.5 - 2.5 * np.log10((sv3_gals['FLUX_W1']/sv3_mw_trans[:,3]).clip(1e-16))
sv3_g_rr = sv3_gals['GAIA_PHOT_G_MEAN_MAG'] - 22.5 + 2.5*np.log10(sv3_gals['FLUX_R'])
# +
from desitarget.sv1.sv1_targetmask import bgs_mask as sv1_bgs_mask
from desitarget.sv3.sv3_targetmask import bgs_mask as sv3_bgs_mask
sv1_bgs = sv1_gals['SV1_BGS_TARGET'] != 0
sv1_brgt = (sv1_gals['SV1_BGS_TARGET'] & sv1_bgs_mask['BGS_BRIGHT']) != 0
sv1_lowq = (sv1_gals['SV1_BGS_TARGET'] & sv1_bgs_mask['BGS_LOWQ']) != 0
sv1_fain = (sv1_gals['SV1_BGS_TARGET'] & sv1_bgs_mask['BGS_FAINT']) != 0
sv1_fain_ext = (sv1_gals['SV1_BGS_TARGET'] & sv1_bgs_mask['BGS_FAINT_EXT']) != 0
sv1_fibmag = (sv1_gals['SV1_BGS_TARGET'] & sv1_bgs_mask['BGS_FIBMAG']) != 0
sv3_bgs = sv3_gals['SV3_BGS_TARGET'] != 0
sv3_brgt = (sv3_gals['SV3_BGS_TARGET'] & sv3_bgs_mask['BGS_BRIGHT']) != 0
sv3_fain = (sv3_gals['SV3_BGS_TARGET'] & sv3_bgs_mask['BGS_FAINT']) != 0
# -
sv3_offset = np.zeros(len(sv3_gals['PHOTSYS']))
sv3_offset[sv3_gals['PHOTSYS'] == 'N'] = 0.04
# +
fig = plt.figure(figsize=(21,7))
sub = fig.add_subplot(131)
sub.scatter((gmag_S - zmag_S), G_rr_S, c='gray', s=0.1, rasterized=True)
sub.scatter((gmag_N - zmag_N), G_rr_N, c='gray', s=0.1, rasterized=True)
sub.scatter([], [], c='gray', s=100, label='LS')
sub.text(5.9, 0.69, 'galaxy', ha='right', va='bottom', fontsize=18)
sub.text(5.7, 0.5, 'star', ha='right', va='top', fontsize=18)
sub.axhline(0.6, color='k', linestyle='--', linewidth=3)
sub.text(0.95, 0.05, r'Legacy Survey', ha='right', va='bottom', transform=sub.transAxes, fontsize=30)
sub.set_xlim(-2., 6.)
sub.set_xticks([-2., 0., 2., 4., 6.])
sub.set_ylim(-3., 5.)
sub = fig.add_subplot(132)
sub.scatter((gmag_S - zmag_S), G_rr_S, c='gray', s=0.1, rasterized=True)
sub.scatter((gmag_N - zmag_N), G_rr_N, c='gray', s=0.1, rasterized=True)
sub.scatter([], [], c='gray', s=100, label='LS')
hb = sub.hexbin((sv1_gmag - sv1_zmag)[sv1_bgs], sv1_g_rr[sv1_bgs],
C=((sv1_gals['RR_SPECTYPE'] == 'STAR') | (sv1_gals['Z_TRUE'] < 0.00100069))[sv1_bgs].astype(float),
edgecolors='k', linewidth=1., reduce_C_function=np.mean, extent=(-2, 6, -2, 5), gridsize=30, vmin=0., vmax=0.2, mincnt=10, cmap='Spectral_r')#, alpha=0.75)
sub.axhline(0.6, color='k', linestyle='--', linewidth=3)
sub.text(0.95, 0.05, r'SV1', ha='right', va='bottom', transform=sub.transAxes, fontsize=30)
sub.set_xlim(-2., 6.)
sub.set_xticks([-2., 0., 2., 4., 6.])
sub.set_ylim(-3., 5.)
sub.set_yticklabels([])
sub = fig.add_subplot(133)
sub.scatter((gmag_S - zmag_S), G_rr_S, c='gray', s=0.1, rasterized=True)
sub.scatter((gmag_N - zmag_N), G_rr_N, c='gray', s=0.1, rasterized=True)
sub.scatter([], [], c='gray', s=100, label='Legacy Survey')
hb = sub.hexbin((sv3_gmag - sv3_zmag)[sv3_brgt | sv3_fain], sv3_g_rr[sv3_brgt | sv3_fain],
C=((sv3_gals['RR_SPECTYPE'] == 'STAR') & (sv3_gals['Z_TRUE'] < 0.00100069))[sv3_brgt | sv3_fain].astype(float),
edgecolors='k', linewidth=1., reduce_C_function=np.mean, extent=(-2, 6, -2, 5), gridsize=30, vmin=0., vmax=0.2, mincnt=10, cmap='Spectral_r')
#sub.scatter((sv3_gmag - sv3_rmag)[sv3_brgt | sv3_fain], sv3_g_rr[sv3_brgt | sv3_fain], c='r', s=1)
sub.axhline(0.6, color='k', linestyle='--', linewidth=3)
sub.text(0.95, 0.05, r'$1\%$', ha='right', va='bottom', transform=sub.transAxes, fontsize=30)
sub.set_xlim(-2., 6.)
sub.set_xticks([-2., 0., 2., 4., 6.])
sub.set_ylim(-3., 5.)
sub.set_yticklabels([])
bkgd = fig.add_subplot(111, frameon=False)
bkgd.set_xlabel(r'$r - z$', labelpad=10, fontsize=30)
bkgd.set_ylabel(r'$G_{\rm Gaia} - r_{\rm raw}$', labelpad=10, fontsize=30)
bkgd.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
fig.subplots_adjust(wspace=0.1, right=0.85)
cbar_ax = fig.add_axes([0.87, 0.15, 0.015, 0.7])
cbar = fig.colorbar(hb, cax=cbar_ax)
cbar.ax.set_ylabel(r'stellar contamination (redrock)', labelpad=30, fontsize=25, rotation=270)
fig.savefig('figs/star_galaxy_redrock.pdf', bbox_inches='tight')
# -
star_galaxy_fail = sv1_bgs & (sv1_g_rr > 0.6) & ((sv1_gals['RR_SPECTYPE'] == 'STAR') | (sv1_gals['Z_TRUE'] < 0.00100069))
print(np.sum(star_galaxy_fail))
for targ in ['BRIGHT', 'FAINT', 'FAINT_EXT', 'LOWQ', 'FIBMAG', 'BRIGHT_NORTH', 'BRIGHT_SOUTH', 'FAINT_NORTH', 'FAINT_SOUTH', 'FAINT_EXT_NORTH', 'FAINT_EXT_SOUTH']:
print(targ, np.sum(sv1_bgs & star_galaxy_fail & ((sv1_gals['SV1_BGS_TARGET'] & sv1_bgs_mask['BGS_%s' % targ]) != 0)))
star_galaxy_fail_bright = sv1_bgs & star_galaxy_fail & sv1_brgt & ~sv1_lowq
print(np.sum(star_galaxy_fail_bright))
sv1_gals[star_galaxy_fail_bright]
sv1_bgs_mask
print(np.sum(star_galaxy_fail & ~sv1_brgt & ~sv1_fain & ~sv1_lowq))
len(sv1_gals), len(sv1_gals[sv1_brgt | sv1_fain | sv1_lowq | sv1_fain_ext | sv1_fibmag])
np.sum(sv1_gals['SV1_BGS_TARGET'] != 0)
# +
sv1_brgt = (sv1_gals['SV1_BGS_TARGET'] & sv1_bgs_mask['BGS_BRIGHT']) != 0
sv1_lowq = (sv1_gals['SV1_BGS_TARGET'] & sv1_bgs_mask['BGS_LOWQ']) != 0
sv1_fain = (sv1_gals['SV1_BGS_TARGET'] & sv1_bgs_mask['BGS_FAINT']) != 0
# -
sv1_gals['Z_TRUE'][star_galaxy_fail]
|
doc/sv_paper/imaging_everest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What is CMS?
# <img src="images/cern_cms_endon.jpg" width="500"/>
# $$pp \rightarrow \chi \chi$$
# The <a href="http://en.wikipedia.org/wiki/Compact_Muon_Solenoid">CMS detector</a> is designed to measure the properties of particles produced in the collisions of high energy protons at the LHC.
#
# The CMS detector is made of many. many sub-detectors and when the particles are created in each proton-proton collision, they fly through these detectors and we are able to measure in which direction all these particles went.
# <img src="images/small_cms1.png" width="500"/>
# These types of displays of the detector can be challenging to understand and at the end of the day, it's not what we scientists actually analyze. We use this information to get the electric charge, energy, and momentum of these particles, and that's about it.
#
# <b>Let's go take a look at some of <i>that</i> data!</b>
# The first step is to import some helper functions. One is to get the collisions data out of the files, and the other is to display the particles that are produced in these collisions.
import tools.cms as cms
# Next, we will open the file and pull out the collision data. This will return a Python list of all the collisions in that file.
#
# You can use these data to visualize individual collisions or to perform a data analysis on <i>all</i> the collisions.
# +
filehandler = open('./data/small_cms_test_file.dat', 'r')
collisions = cms.get_collisions(filehandler)
number_of_collisions = len(collisions)
print("# of proton-proton collisions: {0}".format(number_of_collisions))
# -
# Let's take a look at some of these collisions!
# %matplotlib inline
cms.display_collision3D(collisions[3])
cms.display_collision3D(collisions[3])
cms.display_collision3D(collisions[6])
# Animate - NOT WORKING RIGHT NOW...
'''
from IPython.display import clear_output,display
import time
fig = plt.figure(figsize=(6,4))
for i in range(0,10):
cms.display_collision3D(collisions[i],fig=fig)
time.sleep(0.01)
clear_output(wait=True)
display(fig)
fig.clear()
''';
# What are we looking at here?
#
# * The red lines represent the protons colliding.
# * The other lines represent particles created in the collisions. The length of these lines tell us how much momentum (or energy) they have. The colors are different particles/object.
# * Blue - <a href="http://en.wikipedia.org/wiki/Muon">muons</a>
# * Orange - <a href="http://en.wikipedia.org/wiki/Jet_(particle_physics)">jets</a>
# * Green - <a href="http://en.wikipedia.org/wiki/Jet_(particle_physics)">electrons</a>
# * Gray - <a href="http://en.wikipedia.org/wiki/Photon">photons</a>
# You can also make plots of the properties of the particles.
# +
energies = []
for collision in collisions:
jets,muons,electrons,photons,met = collision
for jet in jets:
energy,px,py,pz,btag = jet
energies.append(energy)
plt.figure(figsize=(4,4))
h = plt.hist(energies)
# -
# So now you know how to play around with data from the CMS experiment at the Large Hadron Collider. What do you want to do next? :)
|
week-2/3-data-visualization/3. CMS Histgrams/A. CMS Experiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Catalyst example on table-data
# @DBusAI
# +
from collections import OrderedDict
import numpy as np
from matplotlib.pylab import plt
# %matplotlib inline
from sklearn.datasets.california_housing import fetch_california_housing
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import mean_squared_error
import torch
from torch import nn
import torch.nn.functional as F
from torch.utils.data import TensorDataset, DataLoader
from catalyst.dl import SupervisedRunner
from catalyst.dl.callbacks import SchedulerCallback
from catalyst.contrib.optimizers import Lookahead
from catalyst.utils import set_global_seed
# -
# ### Reproduce all
# Catalyst provides a special utils for research results reproducibility.
SEED=42
set_global_seed(SEED)
# ### Get some data
# In this tutorial we will use
# [California dataset](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_california_housing.html )<br>
# Also, we split all data: <b>75/25</b> - for training /validation
X, y = fetch_california_housing(return_X_y=True)
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=SEED)
# ### Dataset definition
#
# We have to normalize all X-data
mscl = StandardScaler()
x_train = mscl.fit_transform(x_train)
x_test = mscl.transform(x_test)
# And prepare PyTorch Datasets
train_ds = TensorDataset(torch.FloatTensor(x_train), torch.FloatTensor(y_train.reshape(-1,1)))
test_ds = TensorDataset(torch.FloatTensor(x_test), torch.FloatTensor(y_test.reshape(-1,1)))
# ### DataLoader definition
#
# We have to define bacth size and shuffle train data:
# +
batch = 120
train_dl = DataLoader(train_ds, batch_size=batch, shuffle=True, num_workers=2)
test_dl = DataLoader(test_ds, batch_size=batch, shuffle=False, num_workers=2)
# -
# Catalyst loader:
data = OrderedDict()
data['train'] = train_dl
data['valid'] = test_dl
# ### Define model
#
# Our Neural Network structure will be very simple. Just MLP with 40,20,1 linear layers. Also, default initialization.
class Net(nn.Module):
def __init__(self, num_features):
super(Net,self).__init__()
layers = [40, 20]
self.L1 = nn.Linear(num_features, layers[0])
torch.nn.init.xavier_uniform_(self.L1.weight)
torch.nn.init.zeros_(self.L1.bias)
self.L2 = nn.Linear(layers[0], layers[1])
torch.nn.init.xavier_uniform_(self.L2.weight)
torch.nn.init.zeros_(self.L2.bias)
self.L3 = nn.Linear(layers[1], 1)
torch.nn.init.xavier_uniform_(self.L3.weight)
torch.nn.init.zeros_(self.L3.bias)
def forward(self, x):
x = F.relu(self.L1(x))
x = F.relu(self.L2(x))
x = F.relu(self.L3(x))
return x
model = Net(x_train.shape[1])
# Default optimizer and <b>L2 loss</b>
optimizer = torch.optim.Adam(model.parameters(), lr=1e-2)
crit = nn.MSELoss()
# For model training we need SupervisedRunner and train method:
runner = SupervisedRunner()
runner.train(
model=model,
criterion=crit,
optimizer=optimizer,
loaders=data,
logdir="run",
num_epochs=20)
# ### Inference
#
# Inference part is mush easier: <br>
# <b>/checkpoints/best.pth</b> - is default dir for checkpoints<br>
# <b>run</b> - our logdir
predictions = runner.predict_loader(
data["valid"], resume=f"run/checkpoints/best.pth", verbose=True
)
# ### Results
#
# Let's calculate MSE error
mean_squared_error(y_test, predictions)
# ### Prediction Viz
#
# And finally - show scatterplot for our predictions
plt.scatter(y_test, predictions.flatten())
|
examples/notebooks/table-data-tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Host Annotable web app as Bokeh server
#
# Make a bokeh plot in a Jupyter notebook and serve it up to a remote browser.
#
# ToDo:
# ----
# - Make a basic `bokeh` plot servable
# - Make a `holoviews` plot servable
# - Make a `hvplot` with gridded data and a pen servable
# - Test whether doodles are synched back into the Notebook
# ## Setup
# The following requirements are needed for serving up Bokeh plots from JupyterHub
#
# `$ pip install nbserverproxy && jupyter serverextension enable --py nbserverproxy`
# ## Basic `bokeh` plot served from Jupyter Lab
# ## `holoviews` plot served from Jupyter Lab
import hvplot as hplt
import xarray as xr
import hvplot.xarray
import holoviews as hv
import geoviews as gv
import cartopy.crs as ccrs
import cartopy.feature as cf
from holoviews.streams import FreehandDraw
air_temp = xr.tutorial.open_dataset('air_temperature').load().air.isel(time=0)
print("Loaded dataset")
# +
coastlines = gv.feature.coastline
proj = ccrs.Orthographic(-90, 30)
air_plot = air_temp.hvplot.quadmesh(
'lon', 'lat', projection=proj, project=True, global_extent=True,
width=800, height=720, cmap='viridis', rasterize=True, dynamic=False)
print("Made plot elements")
# +
warm_front = gv.Path([]).opts(color='red', line_width=9)
warm_front_pen = FreehandDraw(source=warm_front)
warning_orange = gv.Polygons([]).opts(line_color='orange', line_width=9,
fill_color='orange', fill_alpha=0.6)
warning_orange_tool = FreehandDraw(source=warning_orange)
print("Made pen elements")
# +
plot = air_plot * coastlines * warm_front * warning_orange
print("Combined elements")
# -
# ## Deploy Jupyter notebook as `bokeh.server`
# +
doc = hv.renderer('bokeh').server_doc(plot)
doc.title = 'Annotable Bokeh App - From Jupyter Notebook'
print("Serving document")
# -
# To run this notebook as a Bokeh app, run the following in a terminal:
#
# `$ bokeh serve --show bokeh_server.ipynb`
# ## Deploy plot as a `bokeh.app`
renderer = hv.renderer('bokeh')
print(renderer)
app = renderer.app(air_plot * coastlines * warm_front)
print(app)
# +
from bokeh.server.server import Server
# server = Server({'/app': app}, port=0)
server = Server({'/app': app}, port=7777, allow_websocket_origin=['pangeo.informaticslab.co.uk'])
# -
server.start()
server.show('/app')
warm_front_pen.element.data
server.stop()
warm_front_pen.element.data
warm_front_pen.element
# ## Define annotable function
def make_annotable(plot, port=0, websocket_origin='pangeo.informaticslab.co.uk', url_path='annotable'):
import holoviews as hv
from bokeh.server.server import Server
import os
from IPython.core.display import display, HTML
import ipywidgets as widgets
renderer = hv.renderer('bokeh')
app = renderer.app(plot)
server = Server({f'/{url_path}': app}, port=port, allow_websocket_origin=[websocket_origin])
server.start()
prefix = os.environ['JUPYTERHUB_SERVICE_PREFIX']
url = f"https://{websocket_origin}{prefix}proxy/{server.port}/{url_path}"
display(HTML(f'<a href={url}>{url}</a>'))
stop_button = widgets.Button(description="Stop Annotable")
stop_button.on_click(lambda b: server.stop())
display(stop_button)
return server
annotable = make_annotable(plot)
annotable.start()
warm_front_pen.element()
warning_orange_tool.element()
annotable.stop()
|
bokeh_server.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ipywidgets based Dashboard
#
# This notebook contains a prototype dashboard built with [ipywidgets](https://ipywidgets.readthedocs.io). The dashboard definitely started to push the limits of `ipywidgets`, suggesting that for this type of dashboard, going straight to Dash or Panel would be a better move.
# +
# %load_ext autoreload
# %autoreload 2
from melbviz.prototype import PedestrianDemo
from melbviz.config import MELBVIZ_COUNTS_CSV_PATH, MELBVIZ_SENSOR_CSV_PATH, MELBVIZ_CLEANED_DATA_PATH
# -
if MELBVIZ_CLEANED_DATA_PATH.is_file():
print("loading dataset from Parquet")
demo = PedestrianDemo.from_parquet(MELBVIZ_CLEANED_DATA_PATH)
else:
print("loading abd cleaning dataset from CVS")
demo = PedestrianDemo.load(MELBVIZ_COUNTS_CSV_PATH, sensor_csv_path=MELBVIZ_SENSOR_CSV_PATH)
demo.prototype()
|
notebooks/demos/panel_prototype.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ##### Example use of the hyperopt to see the sensitivy of an algorith with respect ot their hyperparameters
# Here the use-case is a RandomForest algo and see the importance of different hyperaparameters based on the accuracy on the train and test set.
# Simple train/test split has been performed.
# +
from __future__ import print_function
from pprint import pprint
from time import time
import logging
from sklearn.datasets import fetch_20newsgroups
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.linear_model import SGDClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.metrics import classification_report, confusion_matrix
print(__doc__)
# Display progress logs on stdout
logging.basicConfig(level=logging.INFO,
format='%(asctime)s %(levelname)s %(message)s')
# #############################################################################
# Load some categories from the training set
categories = [
'alt.atheism',
'talk.religion.misc',
]
# Uncomment the following to do the analysis on all the categories
#categories = None
print("Loading 20 newsgroups dataset for categories:")
print(categories)
data = fetch_20newsgroups(subset='train', categories=categories)
print("%d documents" % len(data.filenames))
print("%d categories" % len(data.target_names))
print()
#############################################################################
# Define a pipeline combining a text feature extractor with a simple
# classifier
pipeline = Pipeline([
('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', SGDClassifier()),
])
# uncommenting more parameters will give better exploring power but will
# increase processing time in a combinatorial way
parameters = {
'clf__alpha': (0.00001, 0.000001),
'clf__penalty': ('l2', 'elasticnet'),
'clf__max_iter': (10, 50, 80, 150),
}
X = data.data
y = data.target
if __name__ == "__main__":
# multiprocessing requires the fork to happen in a __main__ protected
# block
# find the best parameters for both the feature extraction and the
# classifier
grid_search = GridSearchCV(pipeline, parameters, n_jobs=-1, verbose=1)
# print("Performing grid search...")
# print("pipeline:", [name for name, _ in pipeline.steps])
# print("parameters:")
# pprint(parameters)
# t0 = time()
# # grid_search.fit(data.data, data.target)
# grid_search.fit(X, y)
# print("done in %0.3fs" % (time() - t0))
# print()
# print("Best score: %0.3f" % grid_search.best_score_)
# print("Best parameters set:")
# best_parameters = grid_search.best_estimator_.get_params()
# for param_name in sorted(parameters.keys()):
# print("\t%s: %r" % (param_name, best_parameters[param_name]))
# -
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
X_train, X_test, y_train, y_test= train_test_split(X, y, random_state=42, test_size=0.3)
# X_valid, X_test, y_valid, y_test= train_test_split(X_test, y_test,
# random_state=42, test_size=0.5)
grid_search.fit(X_train, y_train)
cur_mod = grid_search.best_estimator_
pred = cur_mod.predict(X_test)
print(accuracy_score(y_test, pred))
# +
from sklearn.feature_selection import mutual_info_classif
from sklearn.feature_extraction.text import TfidfVectorizer
pip = Pipeline([
('vect', CountVectorizer()),
('tfidf', TfidfTransformer())])
tr = pip.fit_transform(X_train, y_train)
mi = mutual_info_classif(tr, y_train)
print(len(mi),tr.shape[1])
mi = mi/sum(mi)
#tr_valid = pip.transform(X_valid)
tr_test = pip.transform(X_test)
# +
from sklearn.base import BaseEstimator, ClassifierMixin, clone
from sklearn.utils import check_X_y, check_random_state, check_array
from sklearn.metrics import get_scorer
from sklearn.utils.validation import column_or_1d, check_is_fitted
from sklearn.multiclass import check_classification_targets
from sklearn.utils.metaestimators import if_delegate_has_method
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.preprocessing import MinMaxScaler
from sklearn.feature_selection import mutual_info_classif
import numpy as np
from sklearn.ensemble import RandomForestClassifier
class Liquid_Voter2(BaseEstimator, ClassifierMixin):
"""
Liquid Voter delegation mechanism. It builds a standard random forest but
votes according to the liquid voting mechanism of action. Currently, the competence
is expressed through the accuracy of the trees based on a validation split on the
train set. The alpha and cap parameters essentially control a) how much better does
one model to be with regards to another so as to become it's delegate and b) the maximum
capacity of votes one delegate can carry.
"""
def __init__(self,
ensemble=RandomForestClassifier(),
n_estimators=100,
random_state=42,
min_samples_leaf=2,
#competence = 'valid_acc',
valid_size = 0.1,
alpha = 0.2,
cap = 0.2,
metric='accuracy'):
self.n_estimators = n_estimators
self.random_state = check_random_state(random_state)
self.min_samples_leaf = min_samples_leaf
self.ensemble = ensemble.set_params(**{'random_state':self.random_state,
'n_estimators':self.n_estimators,
'min_samples_leaf':self.min_samples_leaf})
self.alpha = alpha
self.cap = cap
self.scoring = get_scorer(metric)
#self.competence = competence
self.valid_size = valid_size
self.delegation_map = {}
self.competence = []
def fit(self, X, y):
return self._fit(X, y)
def _validate_y(self, y):
y = column_or_1d(y, warn=True)
check_classification_targets(y)
self.classes_, y = np.unique(y, return_inverse=True)
self.n_classes_ = len(self.classes_)
return y
def _fit(self,X,y):
X, y = check_X_y(
X, y, ['csr', 'csc'], dtype=None, force_all_finite=False,
multi_output=True)
y = self._validate_y(y)
n_samples, self.n_features_ = X.shape
self.ensemble.fit(X,y)
return self
def _validate_y(self, y):
y = column_or_1d(y, warn=True)
check_classification_targets(y)
self.classes_, y = np.unique(y, return_inverse=True)
self.n_classes_ = len(self.classes_)
return y
def predict(self, X):
"""Predict class for X.
The predicted class of an input sample is computed as the class with
the highest mean predicted probability. If base estimators do not
implement a ``predict_proba`` method, then it resorts to voting.
Parameters
----------
X : {array-like, sparse matrix} of shape = [n_samples, n_features]
The training input samples. Sparse matrices are accepted only if
they are supported by the base estimator.
Returns
-------
y : array of shape = [n_samples]
The predicted classes.
"""
return self.predict_delegate_(X)
def predict_delegate_(self, X):
import collections
import operator
final_pred = []
for sample in xrange(X.shape[0]):
# Competence
self.competence = []
for est in self.ensemble.estimators_:
proba = est.predict_proba(X[sample,:].reshape(1, -1))[0][0]
#print(proba)
comp = abs(proba-0.5)*2
self.competence.append(comp)
#print(self.competence)
# Delegation
approves = {}
approved_by = {}
approved_by_length = {}
for i, sc_i in enumerate(self.competence):
k = np.copy(self.competence)
approves[i] = np.where(k - sc_i - self.alpha>0)[0].tolist()
approved_by[i] = np.where(k - sc_i + self.alpha<0)[0].tolist()
approved_by_length[i] = len(approved_by[i])
sorted_approved = sorted(approved_by_length.items(), key=operator.itemgetter(1))[::-1]
cap = int(self.n_estimators*self.cap)
cap_per_est = {}
for i in xrange(self.n_estimators):
cap_per_est[i] = cap
poss_nodes = set([i for i in xrange(self.n_estimators)])
#print(sorted_approved)
for est_id, _ in sorted_approved:
to_deleg = []
poss_deleg = list(poss_nodes.intersection(approved_by[est_id]))
if len(poss_deleg) <= cap_per_est[est_id] - 1:
to_deleg = poss_deleg
else:
to_deleg = np.random.choice(poss_deleg, cap_per_est[est_id] - 1).tolist()
to_deleg = to_deleg + [est_id]
cap_per_est[est_id] -= len(to_deleg)
poss_nodes = poss_nodes.difference(to_deleg)
self.delegation_map[est_id] = to_deleg
if len(poss_nodes) == 0:
break
# Prediction
sample_pred = []
for est_id, delegates in self.delegation_map.items():
n_votes = len(delegates)
est_predict = self.ensemble.estimators_[est_id].predict(X[sample,:].reshape(1, -1))[0]
sample_pred.extend([est_predict for i in xrange(n_votes)])
final_pred.append(collections.Counter(sample_pred).most_common()[0][0])
return np.array(final_pred)
lv = Liquid_Voter2(alpha=2, cap=0.1, n_estimators=100, min_samples_leaf=2)
lv.fit(tr,y_train)
pred = lv.predict(tr_test[:])
print(accuracy_score(y_test[:], pred))
# -
np.arange(1, 2)
# +
import pandas as pd
import numpy as np
from sklearn.metrics import accuracy_score
from hyperopt import hp, fmin, tpe, STATUS_OK, Trials
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
space = {
#'criterion': hp.choice( 'criterion', ( 'gini', 'entropy' )),
#'bootstrap': hp.choice( 'bootstrap', ( True, False )),
#'class_weight': hp.choice( 'class_weight', ( 'balanced', 'balanced_subsample', None )),
#'max_depth': hp.choice('max_depth', np.arange( 2, 80, dtype=int)),
#'max_features': hp.choice( 'mf', ( 'sqrt', 'log2', None )),
#'min_samples_split': hp.choice('min_samples_split', np.arange( 2, 20, dtype=int)),
'min_samples_leaf': hp.choice('min_samples_leaf', np.arange( 1, 3, dtype=int)),
'alpha': hp.uniform('alpha', 0.01, 0.8),
'cap': hp.uniform('cap', 0.01, 0.2)
#'n_estimators': hp.choice( 'n_estimators', [100,150, 200, 250, 300, 350,500]),
}
random_state = 42
# X_train, X_cv, y_train, y_cv = train_test_split(X_B,
# y,
# test_size=0.1,
# random_state=random_state,
# stratify=y)
def objective(space):
clf = Liquid_Voter2(random_state=random_state, **space)
clf.fit(tr, y_train)
pred = clf.predict(tr_test)
acc = accuracy_score(y_test, pred)
train_acc = accuracy_score(y_train, clf.predict(tr))
print("Train score: %0.3f -- Validation score: %0.3f"% (train_acc, acc))
to_return = {'loss':1-acc, 'train_loss':1-train_acc, 'status': STATUS_OK }
to_return.update(space)
return to_return
trials = Trials()
best = fmin(fn=objective,
space=space,
algo=tpe.suggest,
max_evals=100,
trials=trials)
print(best)
# -
# # Fetch the trials results
# And keep a separate Dataframe for the categorical variables
tt = [t['result'] for t in trials.trials]
dd = pd.DataFrame(tt)
# categorical_feats = ['criterion',
# 'class_weight',
# 'max_features',
# 'bootstrap']
# dd_cat = dd[categorical_feats+['loss']+['train_loss']]
# for cat in categorical_feats:
# del dd[cat]
del dd['status']
print(dd.shape)
print(dd.columns)
# # Pivot the table
# (this can be done more efficiently probably)
tr = []
for i, row in dd.iterrows():
cur = {}
cur['loss'] = row['loss']
train_loss = row['train_loss']
del row['loss']
del row['train_loss']
cur['set'] = "Test"
cur.update(row.to_dict())
tr.append(cur)
cur = {}
cur['loss'] = train_loss
cur['set'] = "Train"
cur.update(row.to_dict())
tr.append(cur)
dd2 = pd.DataFrame(tr)
dd2
# # Do the same for the categorical Dataframe
#
tr = []
for i, row in dd_cat.iterrows():
cur = {}
cur['loss'] = row['loss']
train_loss = row['train_loss']
del row['loss']
del row['train_loss']
cur['set'] = "Test"
cur.update(row.to_dict())
tr.append(cur)
cur = {}
cur['loss'] = train_loss
cur['set'] = "Train"
cur.update(row.to_dict())
tr.append(cur)
dd2_cat = pd.DataFrame(tr)
dd2_cat
# ### Plot the Results
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
dd2_melted = dd2.melt(['loss', 'set'])
g = sns.lmplot(x="value", y="loss", col="variable", hue='set', data=dd2_melted,sharex=False)
np.arange(0.01, 0.8)
# ### Plot the Results
# (Categorical Vars into boxplot)
# This needs some fixing for the empty labels
dd2_cat_melted = dd2_cat.melt(['loss', 'set'])
g = sns.catplot(x="value", y="loss", col='variable', hue='set',
data=dd2_cat_melted, sharex=False,
kind='box')
|
Hyperapameter Sensitivity Analysis for Tuning (RF example)-Copy1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Introduction To Markdown
#
# Jupyter has an implementation of markdown language that can be used in in markdown cells to create formatted text and media documentation in your notebook. LaTeX is also implemented to create high quality mathematical typeset.
#
# ## Headings
#
# # H1
# ## H2
# ### H3
# #### H4
# ##### H5
# ###### H6
#
# **Code:**
# ```markdown
# #H1
# ##H2
# ###H3
# ####H4
# #####H5
# ######H6
# ```
# **Code:**
# ```markdown
# Heading 1
# =========
#
# Heading 2
# ---------
# ```
# ## Font Styles
#
# **Bold Font** or __Bold Font__
#
# *Italic* or _Italic Font_
#
# ~~Scratched Text~~
#
# Markdown doesn't support underline. but you can do that using HTML <u>Text</u>
#
# **Code:**
# ```markdown
# **Bold Font** or __Bold Font__
#
# *Italic* or _Italic Font_
#
# ~~Scratched Text~~
#
# Markdown doesn't support underline. but you can do that using HTML <u>Text</u>
# ```
#
# ## Lists
#
# - item
# - item
# - subitem
# - subitem
# - item
#
#
# 1. item
# 2. item
# 1. sub item
# 2. sub item
# 3. item
#
# **Code:**
# ```markdown
# - item
# - item
# - subitem
# - subitem
# - item
#
#
# 1. item
# 2. item
# 1. sub item
# 2. sub item
# 3. item
# ```
# ## Links
#
# http://www.github.com/
#
# [Github](http://www.github.com/)
#
#
# **Code:**
# ```
# http://www.github.com/
#
# [Github](http://www.github.com/)
# ```
#
# ### Images
# 
#
# **Code:**
# ```markdown
# 
# ```
# ## Quotes
#
# > Why, oh why, Javascript??? Wars, famine, planetary destruction... I guess as a species, we deserve this abomination...
# >
# > [<NAME>](https://twitter.com/fperez_org)
#
# **Code:**
# ```
# > Why, oh why, Javascript??? Wars, famine, planetary destruction... I guess as a species, we deserve this abomination...
# >
# > [<NAME>](https://twitter.com/fperez_org)
# ```
#
# ## Horizontal Line
# ---
#
# **Code:**
# ```markdown
# ---
# ```
# ## Tables
#
# | Tables | Are | Cool |
# | ------------- |:-------------:| -----:|
# | col 3 is | right-aligned | 1600 |
# | col 2 is | centered | 12 |
# | zebra stripes | are neat | 1 |
#
# **Code:**
#
# ```
# | Tables | Are | Cool |
# | ------------- |:-------------:| -----:|
# | col 3 is | right-aligned | 1600 |
# | col 2 is | centered | 12 |
# | zebra stripes | are neat | 1 |
# ```
# ## Code
#
# You can add in line code like this `import numpy as np`
#
# Or block code:
#
# Python Code:
# ```python
# x = 5
# print "%.2f" % x
# ```
#
# **Code:**
#
# <pre>
# Python Code:
# ```python
# x = 5
# print "%.2f" % x
# ```
#
# </pre>
#
#
#
#
#
#
#
#
# ### HTML
#
# <b>You</b> can <i>render</i> almost any <span style="color:red;">HTML</span> code you <u>like</u>.
#
# **Code:**
#
# ```
# <b>You</b> can <i>render</i> almost any <span style="color:red;">HTML</span> code you <u>like</u>.
# ```
#
#
# ## References
# - https://www.math.ubc.ca/~pwalls/math-python/jupyter/latex/
# - [LaTeX Wiki](http://en.wikibooks.org/wiki/LaTeX/Mathematics)
# - [Duke University, Department of Statistical Science](https://stat.duke.edu/resources/computing/latex)
# - [Equation Sheet](http://www.equationsheet.com/)
|
01 - Jupyter & Markdown/notebooks/01-Introduction to Markdown with Jupyter Notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/swampxx/sciann-examples/blob/main/sciann_burgers_equation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="CGD-ihQ5ELZv"
# Install sciann
# %%capture sciann_installation
# !pip install sciann
# + colab={"base_uri": "https://localhost:8080/"} id="yBncc3uWRP5m" outputId="3eefb89a-a350-4653-a2c8-a4243a534b83"
import numpy as np
import matplotlib.pyplot as plt
import sciann as sn
# %matplotlib inline
# + [markdown] id="5LOhU_gKREGK"
# We reproduce the Burgers Equation example which is illustraded in the SciANN paper.
#
# Burgers Equation:
#
# $u_{,t} + u u_{,x} - (0.01/\pi)u_{,xx} = 0, t \in [0,1], x \in [-1,1]$
#
# initial and boundary conditions $u(t=0, x)=-\sin (\pi x)$ and $u(t, x= \pm 1) = 0$, respectively.
# + id="cflo-SAnRao8"
# In this cell, we define the input variables (x, t)
# and the solution variable u
x = sn.Variable('x')
t = sn.Variable('t')
u = sn.Functional('u', [t, x], 8*[20], 'tanh')
# + id="O2scCAfLRbUT"
# the governing equation
L1 = sn.diff(u, t) + u * sn.diff(u, x) - (0.01 / np.pi) * sn.diff(u, x, order=2)
# initial and boundary conditions should be defined as continuous fucntions
TOL = 0.001 # tolerance value
# note that C1 is zero at all sampling points except t < 0
# if the result of the solution variable u is not zero,
# we penalize it with the loss term
C1 = (1-sn.sign(t - TOL)) * (u + sn.sin(np.pi*x))
C2 = (1-sn.sign(x - (-1+TOL))) * (u)
C3 = (1+sn.sign(x - ( 1-TOL))) * (u)
# + id="dI3ELjxnSpsI"
# define SciANN model
m = sn.SciModel([x, t], [L1, C1, C2, C3])
# + id="ByNYdnkoSwjp"
# generate input data
x_data, t_data = np.meshgrid(
np.linspace(-1, 1, 100),
np.linspace(0, 1, 100)
)
# + id="F4GhoMnqSy2V"
# training...
h = m.train([x_data, t_data], 4*['zero'], batch_size=256, epochs=10000, verbose=2)
# + id="zgr8UFY-S0mg"
# test data
x_test, t_test = np.meshgrid(
np.linspace(-1, 1, 200),
np.linspace(0, 1, 200)
)
# generate predictions for the test data
u_pred = u.eval(m, [x_test, t_test])
# + id="a5RlDL8TS3KS" colab={"base_uri": "https://localhost:8080/"} outputId="3935c527-a2bc-45ad-be39-0d20431c58bf"
# visualization
fig = plt.figure(figsize=(3, 4))
plt.pcolor(x_test, t_test, u_pred, cmap='seismic')
plt.xlabel('x')
plt.ylabel('t')
plt.colorbar()
|
sciann_burgers_equation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Activity #1: Basic Maps with cartopy
# import our usual things
# %matplotlib inline
import cartopy
import pandas as pd
import matplotlib.pyplot as plt
import ipywidgets
# lets make our maps a bit bigger for now
plt.rcParams["figure.dpi"] = 300
# * this is grabbing a "shape" file for frequently used data
# * there are a bunch of specific files you can grab here: https://github.com/nvkelso/natural-earth-vector/tree/master/zips
#
# +
# ok, lets start thinking about how to link this data to
# the actual readings of each detector
# first, lets read in the detector data
seismic = pd.read_csv("/Users/jillnaiman/Downloads/data_tohoku_norm_transpose.csv",
header = None)
# lets upload the locations of each of these sensors
# during the earthquake
locations = pd.read_csv("/Users/jillnaiman/Downloads/location.txt", delimiter="\t",
header = None, names = ["longitude", "latitude", "empty1", "empty2"])
# +
# we have 3 options: we can decrese the number of stations,
# or the number of time samples, or both
# for illustration purposes, lets do both
nstations = 300 # downsample to 300
ntimes = 1440 # factor of 10
import numpy as np
stationsIndex = np.random.choice(range(locations.shape[0]-1),
nstations, replace=False)
timesIndex = np.random.choice(range(seismic.shape[0]-1),
ntimes, replace=False)
# sort each
stationsIndex.sort()
timesIndex.sort()
# -
locations2 = locations.loc[stationsIndex]
seismic2 = seismic.loc[timesIndex,stationsIndex]
seismic2.shape, locations2.shape
# sweet
# +
# note, we can also do the above plot with bqplot as well:
import bqplot
# scales
x_sc = bqplot.LinearScale()
y_sc = bqplot.LinearScale()
# marks
lines = bqplot.Lines(x = seismic2.index.values,
y = seismic2.iloc[:,0],
scales = {'x': x_sc, 'y': y_sc})
# axes
x_ax = bqplot.Axis(scale = x_sc)
y_ax = bqplot.Axis(scale = y_sc, orientation = 'vertical')
# combine into figure
fig = bqplot.Figure(marks = [lines], axes = [x_ax, y_ax])
# create our slider using ipywidgets
slider = ipywidgets.IntSlider(min=0, max=nstations-1)
y_sc.min = -1.0
y_sc.max = 1.0
# create a linking function for slider & plot
def update_slider(event):
lines.y = seismic2.iloc[:,event['new']]
slider.observe(update_slider, 'value')
display(ipywidgets.VBox([slider, fig]))
# note that this is much more responsive now
# than we we did this ourselves
# bqplots ftw
# +
# # ok, so we are now super into linking THING A with THING B
# # so lets link our sesmic data with its location on the map
# # we can do this with cartopy & matplotlib
# @ipywidgets.interact(station = (0, nstations, 1),
# t = (0, ntimes, 1))
# def plot(station = 0, t = 0):
# fig = plt.figure(figsize=(10, 10))
# ax = fig.add_subplot(211,
# projection = cartopy.crs.LambertCylindrical())
# colors = seismic2.iloc[t]
# ax.scatter(locations2["longitude"],
# locations2["latitude"],
# transform = cartopy.crs.PlateCarree(),
# c = colors)
# ax.coastlines()
# ax = fig.add_subplot(212)
# ax.plot(seismic2.index.values, seismic2.iloc[:,station])
# ax.set_ylim(-1, 1)
# -
# # Activity #3: Info viz maps with bqplot
# +
# with bqplot
map_mark = bqplot.Map(scales={'projection': bqplot.AlbersUSA()})
fig = bqplot.Figure(marks=[map_mark], title='Basic Map Example')
fig
# +
# can make a statemap instead
#(1)
sc_geo = bqplot.AlbersUSA()
state_data = bqplot.topo_load('map_data/USStatesMap.json')
# (2)
def_tt = bqplot.Tooltip(fields=['id', 'name'])
states_map = bqplot.Map(map_data=state_data,
scales={'projection':sc_geo},
tooltip=def_tt)
# (2) grab interactions
states_map.interactions = {'click': 'select', 'hover': 'tooltip'}
# (3) grab data directly from map
# we could also grab from the state_data itself
from states_utils import get_ids_and_names
ids, state_names = get_ids_and_names(states_map)
# lets make into arrays for ease
#state_names =np.array(state_names)
#ids = np.array(ids)
state_names, ids
# into arrays
# (4) data
def get_data_value(change):
if change['owner'].selected is not None:
for i,s in enumerate(change['owner'].selected):
print(state_names[s == ids])
states_map.observe(get_data_value,'selected')
# (1)
fig=bqplot.Figure(marks=[states_map],
title='US States Map Example',
fig_margin={'top': 0, 'bottom': 0, 'left': 0, 'right': 0}) # try w/o first and see
fig
# -
# ## Adding in some data to link to our usa map
# lets add in some exprot data
comm = pd.read_csv('/Users/jillnaiman/Downloads/total_export.csv')
# +
comm.loc[comm['State'] == 'Alabama'].values
# we note that these are formatted as strings - this means we'll have to
# do some formatting when we plot data
# also, note that the state name is the first column and not a number
# we'll also have to take care of this too
# -
# grab years
years = list(comm.columns.values)
years = np.array(years[1:]) # get rid of state
# as numbers
years = years.astype('int')
years
# +
sc_geo = bqplot.AlbersUSA()
state_data = bqplot.topo_load('map_data/USStatesMap.json')
def_tt = bqplot.Tooltip(fields=['id', 'name'])
states_map = bqplot.Map(map_data=state_data, scales={'projection':sc_geo}, tooltip=def_tt)
states_map.interactions = {'click': 'select', 'hover': 'tooltip'}
fig=bqplot.Figure(marks=[states_map], title='US States Map Example',
fig_margin={'top': 0, 'bottom': 0, 'left': 0, 'right': 0})
# lets also make a line plot
# second, the lineplot
x_scl = bqplot.LinearScale()
y_scl = bqplot.LinearScale()
ax_xcl = bqplot.Axis(label='Year', scale=x_scl)
ax_ycl = bqplot.Axis(label='Total Export from State NA',
scale=y_scl,
orientation='vertical', side='left')
lines = bqplot.Lines(x = years, y = np.zeros(len(years)),
scales = {'x': x_scl, 'y': y_scl})
#print(lines)
fig_lines = bqplot.Figure(marks = [lines],
axes = [ax_ycl, ax_xcl],)
# let do something additive for all states selected
def get_data_value(change):
exports = np.zeros(len(years))
snames = ''
if change['owner'].selected is not None:
for i,s in enumerate(change['owner'].selected):
sn = state_names[s == ids][0]
snames += sn + ', '
# because of formatting, things are in arrays hence [0]
# also, take out state name hence [1:]
# NOTE! BQPLOT has misspelled massachussetts!
if sn == 'Massachusetts': sn = 'Massachussetts'
exports_in=comm.loc[comm['State'] == sn].values[0][1:]
# there are ","'s in exports we gotta take out
exports_in = np.array([exports_in[i].replace(',','') for i in range(len(exports_in))])
exports = np.add(exports, exports_in.astype('float64'))
lines.y = exports
ax_ycl.label='Total Export from ' + snames
else:
lines.y = np.zeros(len(exports))
ax_ycl.label='Total Export from NA'
states_map.observe(get_data_value,'selected')
# some formatting for vertical
#fig_lines.layout.max_height='250px'
#fig_lines.layout.min_width='800px'
#fig.layout.min_width='800px'
#ipywidgets.VBox([fig_lines,fig])
ipywidgets.HBox([fig,fig_lines])
# -
sn = 'Massachusetts'
sn = 'Massachussetts'
print(comm[comm['State'] == sn])
comm
state_names
comm['State'].index
import pandas as pd
buildings = pd.read_csv("/Users/jillnaiman/Downloads/building_inventory.csv",
na_values = {'Year Acquired': 0, 'Year Constructed': 0, 'Square Footage': 0})
import numpy as np
nsamples =100
dsm = np.random.choice(range(len(buildings)-1),nsamples,replace=False)
dsm
buildingsDS = buildings.loc[dsm]
len(buildingsDS)
# +
import bqplot
x_scl = bqplot.LinearScale()
y_scl = bqplot.LinearScale()
cd = buildings['Congress Dist']
an = buildings['Agency Name']
sf = buildings['Square Footage']
i,j = 0,0
cdNames = cd.unique()
anNames = an.unique()
mask = (cd.values == cdNames[i]) & (an.values == anNames[j])
ya = buildings['Year Acquired'][mask]
yaNames = ya.unique()
sfNames2 = [sf[mask][ya == yaNames[b]].sum() for b in range(len(yaNames)) ]
sfNames2 = np.array(sfNames2)
yfLine = bqplot.Lines(x=yaNames,
y=sfNames2,
colors=['Blue'],
scales={'x': x_scl, 'y': y_scl})
fig = bqplot.Figure(marks=[yfLine])
fig
# -
|
week08/spring2019_prep_notebook_week07_part1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pickle
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import argparse
sns.set(style="darkgrid")
from sklearn.metrics import classification_report
args= argparse.Namespace(
loc = '../data/'
)
# !ls ../data/runs
# # Global Variables
# ## Gold Labels
ENGLISH_GOLD = pd.read_csv(
os.path.join(args.loc,'gold','english_data.tsv'),
sep='\t'
)
HINDI_GOLD = pd.read_csv(
os.path.join(args.loc,'gold','hindi_data.tsv'),
sep='\t'
)
GERMAN_GOLD = pd.read_csv(
os.path.join(args.loc,'gold','german_data.tsv'),
sep='\t'
)
ENGLISH_GOLD.columns
HINDI_GOLD.columns
# ## Model Predictions
GERMAN_PREDS = {
'task_1': pd.read_csv(
os.path.join(args.loc,'runs','FalsePostive_german_task_1_run_2.tsv',),
sep='\t'
) ,
'task_2': pd.read_csv(
os.path.join(args.loc,'runs','FalsePostive_german_task_2_run_3.tsv',),
sep='\t'
) ,
}
ENGLISH_PREDS = {
'task_1': pd.read_csv(
os.path.join(args.loc,'runs','FalsePostive_english_task_1_run_1.tsv',),
sep='\t'
) ,
'task_2': pd.read_csv(
os.path.join(args.loc,'runs','FalsePostive_english_task_2_run_2.tsv',),
sep='\t'
) ,
'task_3': pd.read_csv(
os.path.join(args.loc,'runs','FalsePostive_english_task_3_run_1.tsv',),
sep='\t'
) ,
}
HINDI_PREDS = {
'task_1': pd.read_csv(
os.path.join(args.loc,'runs','FalsePostive_hindi_task_1_run_2.tsv',),
sep='\t'
) ,
'task_2': pd.read_csv(
os.path.join(args.loc,'runs','FalsePostive_hindi_task_2_run_1.tsv',),
sep='\t'
) ,
'task_3': pd.read_csv(
os.path.join(args.loc,'runs','FalsePostive_hindi_task_3_run_1.tsv',),
sep='\t'
) ,
}
# # Helper Functions
def get_analysis(
gold_df:pd.DataFrame,
pred_df:pd.DataFrame,
subtask:str,
):
"""
Args:
gold_df: DataFrame containing gold labels
preds_df: DataFrame containing model preds
subtask: 1,2 or 3
Returns:
analysis_df: DF containing the columns Text, True labels as 'true'
predicted labels as 'pred'
cross_tab: A pandas crosstab as confusion matrix
"""
pred_df.rename(
columns={'result':'pred'},
inplace=True,
)
# gold_df = gold_df[['text_id','text',f'task_{subtask}']]
analysis_df = pd.merge(
pred_df,
gold_df,
on='text_id',
)
analysis_df.rename(
columns={f'task_{subtask}': 'true'},
inplace=True
)
cross_tab = pd.crosstab(
analysis_df.true , #y_true
analysis_df.pred , #y_pred
rownames=['True'], colnames=['Predicted'], margins=True
)
return analysis_df, cross_tab
# +
def print_samples(
analysis_df:pd.DataFrame,
mistakes:bool=True,
num_samples:int=5
)-> None:
"""Prints the samples for analysis"""
with pd.option_context('display.max_colwidth', -1):
if mistakes: #print misclassifications
df = analysis_df[analysis_df.true != analysis_df.pred]
[['text','true','pred']]
else: #print correct classifications
df = analysis_df[analysis_df.true == analysis_df.pred]
[['text','true','pred']]
del df['text_id']
df = df[['text','pred','true']]
print(df.sample(num_samples))
print('\n',df['text'].map(len).describe())
# -
# # English Error Analysis
# ## Task 1
ENGLISH_PREDS[f'task_{1}']
task = 1
eng_task_1_analysis_df, eng_task_1_crosstab = get_analysis(
gold_df = ENGLISH_GOLD,
pred_df = ENGLISH_PREDS[f'task_{task}'],
subtask = task,
)
print(classification_report(
y_true = eng_task_1_analysis_df.true,
y_pred = eng_task_1_analysis_df.pred,
digits = 4,
))
print_samples(
analysis_df = eng_task_1_analysis_df,
mistakes =True,
num_samples=5
)
with sns.plotting_context('paper',font_scale=1.6):
f, ax = plt.subplots(figsize=(6, 6))
ax = sns.heatmap(
eng_task_1_crosstab,
annot=True,
fmt="d",
cmap= 'Blues',
annot_kws={"size": 25},
)
f.savefig(
'confmats/eng_1_confmat.pdf', format='pdf', dpi=600
)
# ## Task 2
task = 2
eng_task_2_analysis_df, eng_task_2_crosstab = get_analysis(
gold_df = ENGLISH_GOLD,
pred_df = ENGLISH_PREDS[f'task_{task}'],
subtask = task,
)
print(classification_report(
y_true = eng_task_2_analysis_df.true,
y_pred = eng_task_2_analysis_df.pred,
digits = 4,
))
print_samples(
analysis_df = eng_task_2_analysis_df,
mistakes =True,
num_samples=5
)
with sns.plotting_context('paper',font_scale=1.6):
f, ax = plt.subplots(figsize=(6, 6))
ax = sns.heatmap(
eng_task_2_crosstab,
annot=True,
fmt="d",
cmap= 'Blues',
annot_kws={"size": 23},
)
f.savefig(
'confmats/eng_2_confmat.pdf', format='pdf', dpi=600
)
# ## Task 3
task = 3
eng_task_3_analysis_df, eng_task_3_crosstab = get_analysis(
gold_df = ENGLISH_GOLD,
pred_df = ENGLISH_PREDS[f'task_{task}'],
subtask = task,
)
print(classification_report(
y_true = eng_task_3_analysis_df.true,
y_pred = eng_task_3_analysis_df.pred,
digits = 4,
))
print_samples(
analysis_df = eng_task_3_analysis_df,
mistakes =True,
num_samples=5
)
with sns.plotting_context('paper',font_scale=1.6):
f, ax = plt.subplots(figsize=(6, 6))
ax = sns.heatmap(
eng_task_3_crosstab,
annot=True,
fmt="d",
cmap= 'Blues',
annot_kws={"size": 25},
)
f.savefig(
'confmats/eng_3_confmat.pdf', format='pdf', dpi=600
)
# # Hindi Error Analysis
# ## TASK 1
task = 1
hin_task_1_analysis_df, hin_task_1_crosstab = get_analysis(
gold_df = HINDI_GOLD,
pred_df = HINDI_PREDS[f'task_{task}'],
subtask = task,
)
print(classification_report(
y_true = hin_task_1_analysis_df.true,
y_pred = hin_task_1_analysis_df.pred,
digits = 5,
))
print_samples(
analysis_df = hin_task_1_analysis_df,
mistakes =True,
num_samples=5
)
with sns.plotting_context('paper',font_scale=1.6):
f, ax = plt.subplots(figsize=(6, 6))
ax = sns.heatmap(
hin_task_1_crosstab,
annot=True,
fmt="d",
cmap= 'Blues',
annot_kws={"size": 25},
)
f.savefig(
'confmats/hin_1_confmat.pdf', format='pdf', dpi=600
)
# ## Task 2
task = 2
hin_task_2_analysis_df, hin_task_2_crosstab = get_analysis(
gold_df = HINDI_GOLD,
pred_df = HINDI_PREDS[f'task_{task}'],
subtask = task,
)
print_samples(
analysis_df = hin_task_2_analysis_df,
mistakes =True,
num_samples=5
)
with sns.plotting_context('paper',font_scale=1.6):
f, ax = plt.subplots(figsize=(6, 6))
ax = sns.heatmap(
hin_task_2_crosstab,
annot=True,
fmt="d",
cmap= 'Blues',
annot_kws={"size": 23},
)
f.savefig(
'confmats/hin_2_confmat.pdf', format='pdf', dpi=600
)
# ## Task 3
task = 3
hin_task_3_analysis_df, hin_task_3_crosstab = get_analysis(
gold_df = HINDI_GOLD,
pred_df = HINDI_PREDS[f'task_{task}'],
subtask = task,
)
print_samples(
analysis_df = hin_task_3_analysis_df,
mistakes =True,
num_samples=5
)
with sns.plotting_context('paper',font_scale=1.6):
f, ax = plt.subplots(figsize=(6, 6))
ax = sns.heatmap(
hin_task_3_crosstab,
annot=True,
fmt="d",
cmap= 'Blues',
annot_kws={"size": 25},
)
f.savefig(
'confmats/hin_3_confmat.pdf', format='pdf', dpi=600
)
# # German Error Analysis
# ## Task 1
#
task = 1
ger_task_1_analysis_df, ger_task_1_crosstab = get_analysis(
gold_df = GERMAN_GOLD,
pred_df = GERMAN_PREDS[f'task_{task}'],
subtask = task,
)
print(classification_report(
y_true = ger_task_1_analysis_df.true,
y_pred = ger_task_1_analysis_df.pred,
digits = 5,
))
print_samples(
analysis_df = ger_task_1_analysis_df,
mistakes =True,
num_samples=5
)
with sns.plotting_context('paper',font_scale=1.6):
f, ax = plt.subplots(figsize=(6, 6))
ax = sns.heatmap(
ger_task_1_crosstab,
annot=True,
fmt="d",
cmap= 'Blues',
annot_kws={"size": 25},
)
f.savefig(
'confmats/ger_1_confmat.pdf', format='pdf', dpi=600
)
# ## Task 2
task = 2
ger_task_2_analysis_df, ger_task_2_crosstab = get_analysis(
gold_df = GERMAN_GOLD,
pred_df = GERMAN_PREDS[f'task_{task}'],
subtask = task,
)
print_samples(
analysis_df = ger_task_2_analysis_df,
mistakes =True,
num_samples=5
)
with sns.plotting_context('paper',font_scale=1.6):
f, ax = plt.subplots(figsize=(6, 6))
ax = sns.heatmap(
ger_task_2_crosstab,
annot=True,
fmt="d",
cmap= 'Blues',
annot_kws={"size": 23},
)
f.savefig(
'confmats/ger_2_confmat.pdf', format='pdf', dpi=600
)
|
notebooks/HASOC Error Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Challenge PART 1 Get the Weather Description and Amount of Precipitation for Each City
#
# +
import numpy as np
import pandas as pd
# -
pip install citipy
from citipy import citipy
# Create a set of random latitude and longitude combinations.
lats = np.random.uniform(low=-90.000, high=90.000, size=1500)
lngs = np.random.uniform(low=-180.000, high=180.000, size=1500)
lat_lngs = zip(lats, lngs)
# Add the latitudes and longitudes to a list.
coordinates = list(lat_lngs)
# +
# Create a list for holding the cities.
cities = []
# Identify the nearest city for each latitude and longitude combination.
for coordinate in coordinates:
city = citipy.nearest_city(coordinate[0], coordinate[1]).city_name
# If the city is unique, then we will add it to the cities list.
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count.
len(cities)
# +
# Import the requests library.
import requests
from datetime import datetime
from config import weather_api_key
# -
print(weather_api_key)
# +
# Starting URL for Weather Map API Call.
#url = f"http://api.openweathermap.org/data/2.5/weather?units=Imperial&APPID=(weather_api_key)"
url = "http://api.openweathermap.org/data/2.5/weather?units=Imperial&APPID=" + weather_api_key
print(url)
# +
# list of city data
city_data = []
# Print the beginning of the logging.
print("Beginning Data Retrieval")
print("-" * 25)
# Create counters
record_count = 1
set_count = 1
# Loop through all the cities in the list.
for i, city in enumerate(cities):
# Group cities in sets of 50 for logging purposes.
if (i % 50 == 0 and i >= 50):
set_count += 1
record_count = 1
# Create endpoint URL with each city.
city_url = url + "&q=" + city.replace(" ","+")
print(city_url)
# Log the URL, record, and set numbers and the city.
print(f"Processing Record {record_count} of Set {set_count} | {city}")
# Add 1 to the record count.
record_count += 1
# try:
# # Parse the JSON and retrieve data.
# city_weather = requests.get(city_url).json()
# # Parse out the needed data.
# # city_lat = city_weather["coord"]["lat"]
# # city_lng = city_weather["coord"]["lon"]
# # city_max_temp = city_weather["main"]["temp_max"]
# # city_humidity = city_weather["main"]["humidity"]
# # city_clouds = city_weather["clouds"]["all"]
# # city_wind = city_weather["wind"]["speed"]
# # city_country = city_weather["sys"]["country"]
# # city_weather_description = city_weather["weather"][0]["description"]
# city_lat = city_weather["coord"]["lat"]
# city_lng = city_weather["coord"]["lon"]
# city_max_temp = city_weather["main"]["temp_max"]
# city_humidity = city_weather["main"]["humidity"]
# city_clouds = city_weather["clouds"]["all"]
# city_wind = city_weather["wind"]["speed"]
# city_country = city_weather["sys"]["country"]
# city_weather_description = city_weather["weather"][0]["description"]
# try:
# city_rain_inches = city_weather["rain"]["1h"]
# except KeyError:
# city_rain_inches = 0
# try:
# city_snow_inches = city_weather["snow"]["1h"]
# except KeyError:
# city_snow_inches = 0
# # Append the city information into city_data list.
# city_data.append(
# {
# "City": city,
# "Lat": city_lat,
# "Lng": city_lng,
# "Max Temp": city_max_temp,
# "Humidity": city_humidity,
# "Cloudiness": city_clouds,
# "Wind Speed": city_wind,
# "Country": city_country,
# "Date": city_date,
# "Current Description": city_weather_description,
# "Rain(inches)": city_rain_inches,
# "Snow(inches)": city_snow_inches
# })
# except:
# print("City not found. Skipping...")
# pass
try:
city_weather = requests.get(city_url).json()
# Parse out the max temp, humidity, and cloudiness
city_lat = city_weather["coord"]["lat"]
city_lng = city_weather["coord"]["lon"]
city_max_temp = city_weather["main"]["temp_max"]
city_humidity = city_weather["main"]["humidity"]
city_clouds = city_weather["clouds"]["all"]
city_wind = city_weather["wind"]["speed"]
city_country = city_weather["sys"]["country"]
city_weather_description = city_weather["weather"][0]["description"]
try:
city_rain_inches = city_weather["rain"]["1h"]
except KeyError:
city_rain_inches = 0
try:
city_snow_inches = city_weather["snow"]["1h"]
except KeyError:
city_snow_inches = 0
# Append the City information into city_data list
city_data.append({"City": city.title(),
"Lat": city_lat,
"Lng": city_lng,
"Max Temp": city_max_temp,
"Current Description": city_weather_description,
"Rain (inches)": city_rain_inches,
"Snow (inches)": city_snow_inches,
"Humidity": city_humidity,
"Cloudiness": city_clouds,
"Wind Speed": city_wind,
"Country": city_country})
# If an error is experienced, skip the city
except:
print("City not found. Skipping...")
pass
# Indicate that Data Loading is complete.
print("-" * 20)
print("Data Retrieval Complete")
print("-" * 20)
# -
len(city_data)
# convert list of dictionaries to a pandas Dataframe
city_data_df = pd.DataFrame(city_data)
city_data_df.head(10)
city_data_df.columns
new_column_order = sorted(city_data_df.columns.tolist())
city_data_df = city_data_df[new_column_order]
city_data_df
raining_cities = city_data_df.loc[city_data_df["Rain (inches)"]> 0]
raining_cities.head()
snowing_cities = city_data_df.loc[city_data_df["Snow (inches)"]> 0]
snowing_cities.head()
# Create the output file (CSV)
output_data_file = "data/WeatherPy_Database.csv"
city_data_df.to_csv(output_data_file, index_label="City_ID")
|
Weather_Database.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Instagram AGDS - NumPy implementation
import numpy as np
import pickle
import pandas as pd
#from tqdm import tqdm
with open("word_df_normalized.pickle", "rb") as f:
word_df = pickle.load(f)
word_df.head()
# +
# Convert word DataFrame to the dictionary
word_dict = word_df.to_dict('index')
i = 0
for k,v in word_dict.items():
print(f"{k}: {v}")
if i == 10:
break
i+= 1
# +
# Load the available influencers
arch_df = pd.read_csv("archetypes_pl_available.csv", index_col=0, header=0)
# Show the head of the DataFrame
arch_df.head()
# +
# Load the archetypes from the list
import os
import toml
import re
import itertools
from text_cleaner import *
class InfluencerNode(object):
def __init__(self, profile_name: str, db_directory: str, data_frame: pd.DataFrame):
self.username = profile_name
self.posts = []
self.hashtags = []
# Load the user data
try:
if os.path.exists(db_directory):
# Iterate over directories in categories:
category_dirs = os.listdir(db_directory)
for cat_dir in category_dirs:
user_dir = os.path.join(db_directory, cat_dir)
# Check if influencer username is present in this category
if profile_name in os.listdir(user_dir):
user_path = os.path.join(user_dir, profile_name)
# Get user data
with open(os.path.join(user_path, f"{profile_name}.toml"), "r") as f:
toml_file = toml.load(f)
# After reading .toml file, set up the node attributes
self.full_name: str = toml_file["full_name"]
self.biography: str = toml_file["biography"]
self.business_category: str = toml_file["business_category_name"]
self.followers: int = toml_file["followers"]
self.followees: int = toml_file["followees"]
self.mediacount: int = toml_file["mediacount"]
self.posts: list = []
# Red all posts into a list
for file in os.listdir(user_path):
if not file.endswith(".toml"):
with open(os.path.join(user_path, file), "r") as post_f:
self.posts.append(post_f.read())
self.hashtags = self.extract_hashtags()
# Post reedition
self.posts = [remove_stopwords(clean_up_text(post)) for post in self.posts]
# Set up archetype/character trait weights
self.traits = data_frame.loc[self.username].to_dict()
for k, v in self.traits.items():
setattr(self, k, v)
except FileNotFoundError as e:
print(e)
def extract_hashtags(self):
HASH_RE = re.compile(r"\#\w+")
out_list = []
for post in self.posts:
tmp = re.findall(HASH_RE, post)
out_list.append(list(set(tmp)))
out_list = list(itertools.chain.from_iterable(out_list))
return list(set(out_list))
# +
import copy
node_list = []
for i, row in tqdm(arch_df.iterrows()):
try:
tmp = InfluencerNode(i, "instagram_dataset/pl", arch_df)
node_list.append(tmp)
except:
pass
# -
print(f"List length: {len(node_list)}")
print(node_list[10].traits)
# + tags=[]
# Iterate over word dictionary to associate words with influencers
word_keys = word_dict.keys()
word_influencer_array = np.array([[word, []] for word in word_keys], dtype=object)
# -
for i, pair in tqdm(enumerate(word_influencer_array)):
word = pair[0]
inf_list = []
# Iterate over all the influencers
for node in node_list:
if word in node.hashtags:
inf_list.append((node.username, node.traits))
continue
if word in list(itertools.chain.from_iterable(node.posts)):
inf_list.append((node.username, node.traits))
word_influencer_array[i][1] = inf_list
# +
# Save the generated array as binary
np.save("word_influencer_graph.npy", word_influencer_array)
# Save the generated file as pickle
with open("word_influencer_graph.pickle", "wb") as f:
pickle.dump(word_influencer_array, f)
# -
# Map words to numpy array indices
word_map = {word: i for i, word in enumerate(list(word_keys))}
# Save map to pickle
with open("word_map.pickle", "wb") as f:
pickle.dump(word_map, f)
with open("word_influencer_graph.pickle", "rb") as f:
graph = pickle.load(f)
graph[:1]
|
instagram_analysis/archive/instagram_agds_numpy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic Python Revision
# ### Assigment question's
'''
Ques 1: Answer these 3 questions without typing code. Then type code to check your answer.
What is the value of the expression 4 * (6 + 5)
What is the value of the expression 4 * 6 + 5
What is the value of the expression4 * 6 + 5
Ques 2: What is the type of the result of the expression 3 + 1.5 + 4?
Ques 3: What would you use to find a number’s square root, as well as its square?
Ques 4: Given the string 'hello' give an index command that returns 'e’.
Ques 5: Reverse the string 'hello' using slicing
Ques 6: Given the string hello, give two methods of producing the letter 'o' using indexing.
Ques 7: Check if a list contains an element
Ques 8: priint the strings present in reverse order and items also inreversse manner
Ques 9: print("the homogeneos list in descendiing order
Ques 10: print( odd index number items present in list
'''
# answer 1
print(4*(6+5)) #44
print(4 * 6 + 5) #29
print(4 + 6 * 5) #34
# answer 2
print(3 + 1.5 + 4) # float
# answer 3
print(5 ** 0.5) # sqrt with power
# answer 4/5/6
str = "hello"
print(str[1])
print(str[::-1])
print(str[4])
print(str[-1])
# +
# create a lottery game
guess_num = 5
user_num = int(input("Guess any number: "))
if(user_num > guess_num):
print("lower down your guess")
elif(user_num < guess_num):
print("increases your guess value")
else:
print("You won Lottery!!!")
# -
# nested loop
# Q: you have to fetch all the item from list and then acess all charcter of item of list.
a = ["akash","mango","apple"]
for i in a:
for j in i:
print(j, end=" ")
# print even or odd
num = int(input("Enter num: "))
if num % 2 == 0:print("Even")
else:print("Odd")
# +
# vowel or not
a = ['a','e','i','o','u']
char = input("Enter character : ")
if char in a:
print("Its vowel")
else:
print("not a vowel ")
# -
a=15
b = int(input("guess the number"))
while b!=a:
print('wrong! try again')
b=int(input())
print('correct')
# ### lecture 2 Assignment
#
'''
Q1: Write a Python function to sum all the numbers in a list
Q2: Write a Python function that takes a list and returns a new list with unique elements of the first list
Q3: Write a Python program to print the even numbers from a given list
Q4: Write a function func1() such that it can accept a variable length of argument and print all arguments value
Q5: Write a function calculation() such that it can accept two variables and calculate the addition and subtraction of them. And also it must return both addition and subtraction in a single return call
Q6: Create a function showEmployee() in such a way that it should accept employee name, and its salary and display both. If the salary is missing in the function call assign default value 9000 to salary
'''
# sol 1
num_lis = [1,2,3,4,5,6,7,8,9,10]
print(sum(num_lis))
# +
# sol 2
def uniquelis(lis):
uni_lis = []
for i in lis:
if i not in uni_lis:
uni_lis.append(i)
return uni_lis
x = [1,1,2,2,3,3,4,4,5,5]
print(uniquelis(x))
# +
# sol 3
def evenNum(lis):
even_lis = []
for i in lis:
if i%2 == 0:
even_lis.append(i)
return even_lis
x = [1,2,3,4,5,6,7,8,9,10]
print(evenNum(x))
# +
# sol 4
def fact1(*arg):
for i in arg:
print(i)
fact1(1,2,3,4,8,2,6,9,2,63,5,5,2,2)
# +
# sol 5
def cal(a,b):
return a+b,a-b
x,y = cal(5,10)
print(x,y)
# +
# sol 6
def showEmployee(name,salary = 9999):
print(f"{name} earning is {salary}")
showEmployee("Akash",10000)
# -
# # Lecture 3
import pandas as pd
data = pd.Series([1,2,3,4,5,6])
data
# get index value
data.index
data.values
data = pd.Series([10,20,30,40,50,60,70],index=['a','b','c','d','e','f','g'])
data.index
data.values
data['c']
data['a':'c']
# Lec 3
a = (10,20,30,40,50,60,70)
df = pd.Series(a,dtype=float,index=range(1,8))
df
a = {'a':22,'b':23,'c':25}
df = pd.Series(a)
df
# ### DataFrame
# +
a = [1,2,3,4]
b = [5,6,7,8]
numSet = list(zip(a,b))
df = pd.DataFrame(numSet,columns = ['x','y'])
df
# -
df.to_csv("dataframe1.csv")
df_read = pd.read_csv("dataframe1.csv")
df_read
df_read['x']
#dic to pandas
d = {'a':[1,2],'b':[3,4]}
df = pd.DataFrame(d)
df
# ### Assignment
'''
Q1. Write a Pandas program to convert Series of lists to one Series.
Q2. Write a Pandas program to compare the elements of the two Pandas Series.
Q3. Sample Series:
Q4. Write a Pandas program to add, subtract, multiple and divide two Pandas Series.
'''
# sol1
data = pd.Series([[1,2],[3,4],[5,6],[7,8]])
data = data.apply(pd.Series)
data
# +
# sol2
a = pd.Series([1,2,3])
b = pd.Series([3,2,1])
print(a > b)
print(a < b)
print(a == b)
# +
# sol 3
x,y = pd.Series([2, 4, 6, 8, 10]), pd.Series([1, 3, 5, 7, 10])
print(x+y)
print(x-y)
print(x*y)
print(x/y)
# -
# # Lec 4
df = pd.read_csv("Dataset/students.csv",header = 0)
df
df1 = df.dropna()
df1
import pandas as pd
import numpy as np
data = pd.read_csv("Dataset/ign.csv")
data
# +
# remenber
# head
data.head() # first five record
data.head(15) # as much as you want
# tail
data.tail()
data.tail(15)
# +
# info
data.info() # complete descp
# describe
data.describe() # describe dataset by performing mathematical opr (only int/float type)
# -
# shape
data.shape # number of row and column
# columns
data.columns # in list of all column
# to count NaN value in dataset
data.isnull().sum()
# ### Indexing and Slicing of DataFrame
# +
ser = data["score_phrase"] # taking particular column
ser
# selected dataset
ser_slic = ser[1:6]
ser_slic
# +
# two col by name
col = data[["score_phrase","title"]]
col
# +
# two col by index iloc[:R,:C]
data.iloc[:,1]
data.iloc[1:10,1:5]
# +
# col by loc
data.loc[:,['url','genre']]
# random index col accessing
data.iloc[:,[2,5]]
# +
# particular row aand column selection
data.iloc[[34,89,100],[2,4,8,9]]
# -
data.head()
data.iloc[:,:]
# +
# filtering data
fil = data.query("score > 8")
fil
# +
# other method
fil = data[data.score > 9]
fil
# -
# 3rd way
fil = data[data["score"] > 9]
fil
data.score.unique()
data.score.unuique()
# # Assigmnet Lec 4
'''
Q1: Create a DataFrame using List:
Q2: How will you create a series from dict in Pandas?
Q3: How will you create an empty DataFrame in Pandas?
Q4: How will you add a column to a pandas DataFrame?
Q5: How to add an Index, row, or column to a Pandas DataFrame?
Q6: How to Delete Indices, Rows or Columns From a Pandas Data Frame?
Q7: How to Rename the Index or Columns of a Pandas DataFrame?
Q8: How to get the items of series A not present in series B?
Q9: How to get the items not common to both series A and series B?
Q10: How to get the minimum, 25th percentile, median, 75th, and max of a numeric series?
Q11: count the no. of game publish (acording to score_phrase)
Q12: count the games run on Playstation Vita
Q13: number of games start with Letter "L"
Q14: sort the Amazing games record in descending order (according to score)
Q15: count the games of different genre
Q16: sort the 2012 records of Puzzle genre game(according to score)
'''
import pandas as pd
# sol1
lis = [1,2,3,4,5,6]
datafram = pd.DataFrame(lis)
datafram
# Sol 2
dict = {'a':9,'b':8,'c':7,'d':6,'e':5}
ser = pd.Series(dict)
ser
# sol 3
emp = pd.DataFrame()
emp
# sol 4
lis = [[10,20,30],[40,50,60],[70,80,90]]
data = pd.DataFrame(lis,columns = ['a','b','c'])
data['d'] = [100,110,120]
data
# sol 5 # new row
lis = [[10,20,30],[40,50,60],[70,80,90]]
data = pd.DataFrame(lis)
new_data = [100,110,120]
data.loc[3] = new_data
data
# +
# sol 6 # new col
lis = [[10,20,30],[40,50,60],[70,80,90]]
data = pd.DataFrame(lis)
newdata = [100,110,120]
data = data.assign(age = newdata)
data
# -
# sol 6 drop row and col
lis = [[10,20,30],[40,50,60],[70,80,90]]
data = pd.DataFrame(lis)
data
data.drop([1],inplace = True) # row 1 droped
data
data.drop([2],axis = 1,inplace = True) # col 2 droped
data
# sol 7 rename
lis = [[10,20,30],[40,50,60],[70,80,90]]
data = pd.DataFrame(lis,columns = ['a','b','c'])
data
# +
# sol 8
a = pd.Series([2, 4, 8, 20, 10, 47, 99])
b = pd.Series([1, 3, 6, 4, 10, 99, 50])
res = a[~a.isin(b)]
print(res)
# -
# sol 9
a = pd.Series([2, 4, 8, 20, 10, 47, 99])
b = pd.Series([1, 3, 6, 4, 10, 99, 50])
# sol 10
a = pd.Series([2, 4, 8, 20, 10, 47, 99])
a.describe()
# sol 11
data = pd.read_csv("Dataset/ign.csv")
data
data.score_phrase.count()
data.platform.value_counts().PlayStationVita
# # Lec 5
# +
# concat // merging // joining {4 type}
# -
# #
|
Lec 5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using QAOA to solve a UD-MIS problem
# +
import numpy as np
import igraph
from itertools import combinations
import matplotlib.pyplot as plt
from pulser import Pulse, Sequence, Register
from pulser.simulation import Simulation
from pulser.devices import Chadoq2
from scipy.optimize import minimize
# -
# ## 1. Introduction
# In this tutorial, we illustrate how to solve the Maximum Independent Set (MIS) problem using the Quantum Approximate Optimization Algorithm procedure on a platform of Rydberg atoms in analog mode, using Pulser.
#
# For more details about this problem and how to encode it on a Rydberg atom quantum processor, see [Pichler, et al., 2018](https://arxiv.org/abs/1808.10816), [Henriet, 2020]( https://journals.aps.org/pra/abstract/10.1103/PhysRevA.101.012335) and [Dalyac, et al., 2020]( https://arxiv.org/abs/2012.14859).
# Consider an undirected graph composed of a set of vertices connected by unweighted edges. An independent set of this graph is a subset of vertices where any two elements of this subset are not connected by an edge. The Maximum Independent Set (MIS) corresponds to the largest of such subsets, and it is in general an NP-complete problem to determine the MIS of a graph.
#
# The MIS problem has several interesting applications, such as portfolio diversification in
# finance, or broadcast systems (wifi or cellular network) optimization.
# For example, assume an ensemble of identical radio transmitters over French cities that each have the same radius of transmission. It was quickly realized that two transmitters with close or equal frequencies could interfere with one another, hence the necessity to assign non-interfering frequencies to overlapping transmiting towers. Because of the limited amount of bandwith space, some towers have to be assigned the same or close frequencies. The MIS of a graph of towers indicate the maximum number of towers that can have close or equal given frequency (red points).
#
# <center>
# <img src="attachment:France_MIS.png" alt="MIS map France" width="650">
# </center>
# Here, we will show how to solve the MIS problem on Unit-Disk (UD) graphs with an excellent approximation.
# When looking for the MIS of a graph, we separate the nodes into two distinct classes: an independence one and the others. We can attribute a status $z$ to each node, where $z_i = 1$ if node $i$ is attributed to the independent set, and $z_i=0$ otherwise. The Maximum Independent Set corresponds to the minima of the following cost function:
#
# $$
# C(z_1,\dots,z_N) = -\sum_{i=1}^N z_i + U \sum_{\langle i,j \rangle}z_i z_j
# $$
#
# where $U \gg 1$ and $\langle i,j \rangle$ represents adjacent nodes (i.e. there is a link between node $i$ and $j$). In this cost function, we want to promote a maximal number of atoms to the $1$ state, but the fact that $U \gg 1$ strongly penalizes two adjacent vertices in state $1$. The minimum of $C(z_0,\dots,z_N)$ therefore corresponds to the maximum independent set of the graph.
# Interestingly, the operator $\hat{C}$ associated with the cost function of the previous equation can be natively realized on a neutral atom platform with some constraints on the graph edges. We map a ground state and a Rydberg state of each atom to a two-level system, where $|1 \rangle = |r \rangle$ is a Rydberg state and $|0 \rangle = |g \rangle$ is a ground state. An atom in a Rydberg state has an excited electron with a very high principal quantum number and therefore exhibits a huge electric dipole moment. As such, when two atoms are excited to Rydberg states, they exhibit a strong van der Waals interaction. Placing $N$ atoms at positions $\textbf{r}_j$ in a 2D plane, and coupling the ground state $|0\rangle$ to the Rydberg state $|1\rangle$ with a laser system enables the realization of the Hamiltonian :
#
# $$
# H= \sum_{i=1}^N \frac{\hbar\Omega}{2} \sigma_i^x - \sum_{i=1}^N \frac{\hbar \delta}{2} \sigma_i^z+\sum_{j<i}\frac{C_6}{|\textbf{r}_i-\textbf{r}_j|^{6}} n_i n_j.
# \label{eq:ising_Hamiltonian}
# $$
#
# Here, $\Omega$ and $\delta$ are respectively the Rabi frequency and detuning of the laser system and $\hbar$ is the reduced Planck constant. The first two terms of the equation govern the transition between states $|0\rangle$ and $|1 \rangle$ induced by the laser, while the third term represents the repulsive Van der Waals interaction between atoms in the $|0\rangle$ state. More precisely, $n_i = \frac 12 (\sigma_i
# ^z + 1)$ counts the number of Rydberg excitations at position $i$. The interaction strength between two atoms decays as $|\textbf{r}_i-\textbf{r}_j|^{-6}$.
# ### From a graph to an atomic register
# We now illustrate how one can use Pulser and a neutral-atom device to find the MIS of a UD-graph. Because the quantum platform is emulated in this notebook, we restrict the number of atoms to 5, just to show a proof-of-concept.
#
# A link in the graph corresponds to two atoms that are within the Rydberg Blockade Radius (RBR) of each other. The radius of RBR is directly linked to the Rabi frequency $\Omega$ and is obtained using `Chadoq2.rydberg_blockade_radius()`. In this notebook, $\Omega$ is fixed to a frequency of 1 rad/µs.
def pos_to_graph(pos):
rb = Chadoq2.rydberg_blockade_radius(1.)
g = igraph.Graph()
N = len(pos)
edges = [[m,n] for m,n in combinations(range(N), r=2) if np.linalg.norm(pos[m] - pos[n]) < rb]
g.add_vertices(N)
g.add_edges(edges)
return g
# Here, we create an atomic register with 5 atoms. We draw our register with half-radius circles around the atoms to highlight the crossing of each one's blockade radius (thus forming a linked graph of interacting qubits):
# +
pos = np.array([[0., 0.], [-4, -7], [4,-7], [8,6], [-8,6]])
G = pos_to_graph(pos)
qubits = dict(enumerate(pos))
reg = Register(qubits)
reg.draw(blockade_radius=Chadoq2.rydberg_blockade_radius(1.), draw_graph=True, draw_half_radius=True)
# -
# This graph has two maximal independent sets: $(1,3,4)$ and $(2,3,4)$, respectively `01011` and `00111` in binary.
# ## 2. Building the quantum loop
# Now, we must build the quantum part of the QAOA. All atoms are initially in the groundstate $|00\dots0\rangle$ of the `ground-rydberg`basis. We then apply $p$ layers of alternating non-commutative Hamiltonians. The first one, called the mixing Hamiltonian $H_M$, is realized by taking $\Omega = 1$ rad/µs, and $\delta = 0$ rad/µs in the Hamiltonian equation. The second Hamiltonian $H_c$ is realized with $\Omega = \delta = 1$ rad/µs. $H_M$ and $H_c$ are applied turn in turn with parameters $\tau$ and $t$ respectively. A classical optimizer is then used to estimate the optimal parameters.
#
# Instead of creating a new `Sequence` everytime the quantum loop is called, we are going to create a parametrized `Sequence` and give that to the quantum loop.
# +
LAYERS = 2
# Parametrized sequence
seq = Sequence(reg, Chadoq2)
seq.declare_channel('ch0','rydberg_global')
t_list = seq.declare_variable('t_list', size=LAYERS)
s_list = seq.declare_variable('s_list', size=LAYERS)
if LAYERS == 1:
t_list = [t_list]
s_list = [s_list]
for t, s in zip(t_list, s_list):
pulse_1 = Pulse.ConstantPulse(1000*t, 1., 0., 0)
pulse_2 = Pulse.ConstantPulse(1000*s, 1., 1., 0)
seq.add(pulse_1, 'ch0')
seq.add(pulse_2, 'ch0')
seq.measure('ground-rydberg')
# -
# Once we have the parameters that we want to apply, we use the `.build()` method to assign these values into a `assigned_seq` sequence. It is this sequence which is simulated every time the quantum loop is called. Here's an example of a sequence for some arbitrary parameters:
# Experimentally, we don't have access to the state vector $|\psi\rangle$. We therefore make it more realistic by taking samples from the state vector that results from running the simulation with `simul.run()`. This is done with the built-in method `results.sample_final_state()`, in which we add the measurement basis which was declared at the end of the sequence, and the number of samples desired. Currently, the repetition rate of the machine is $5$Hz.
def quantum_loop(parameters):
params = np.array(parameters)
t_params, s_params = np.reshape(params.astype(int), (2, LAYERS))
assigned_seq = seq.build(t_list=t_params, s_list=s_params)
simul = Simulation(assigned_seq, sampling_rate=.01)
results = simul.run()
count_dict = results.sample_final_state() #sample from the state vector
return count_dict
guess = {'t': np.random.uniform(8, 10, LAYERS),
's': np.random.uniform(1, 3, LAYERS)}
example_dict = quantum_loop(np.r_[guess['t'], guess['s']])
# We can then plot the distribution of the samples, to see the most frequent bitstrings sampled.
def plot_distribution(C):
C = dict(sorted(C.items(), key=lambda item: item[1], reverse=True))
indexes = ['01011', '00111'] # MIS indexes
color_dict = {key:'r' if key in indexes else 'g' for key in C}
plt.figure(figsize=(12,6))
plt.xlabel("bitstrings")
plt.ylabel("counts")
plt.bar(C.keys(), C.values(), width=0.5, color = color_dict.values())
plt.xticks(rotation='vertical')
plt.show()
plot_distribution(example_dict)
# The bitstrings `01011` and `00111` (in red) correspond to the two MIS of the graph. The goal of QAOA is to choregraph interferences between the basis states, in order to maximize the frequency of the MIS states.
# ## 3. Optimization
# We estimate the cost of a sampled state vector by making an average over the samples. This is done by taking the corresponding bitstring ${\bf z}=(z_1, \ldots, z_N)$ and calculating
#
# $$
# C({\bf z}) = - \sum_i z_i + \sum_{i\geq j} pA_{ij}z_iz_j = p\,({\bf z}^\top \cdot A^\textsf{U} \cdot {\bf z}) - |{\bf z}|_0,
# $$
#
# where $A^\textsf{U}$ is the upper triangular part of the adjacency matrix of the graph, $|\cdot|_0$ gives the sum of non-zero terms of the bitstring, and $p$ is the "penalty" introduced by the magnitude of the quadratic term.
#
# Determining the cost of a given bitstring takes polynomial time. The average estimate is then used in the classical loop to optimize the variational parameters $\tau$ and $t$.
# +
def get_cost_colouring(bitstring, G, penalty=10):
z = np.array(list(bitstring), dtype=int)
A = np.array(G.get_adjacency().data)
# Add penalty and bias:
cost = penalty*(z.T @ np.triu(A) @ z) - np.sum(z)
return cost
def get_cost(counter,G):
cost = sum(counter[key] * get_cost_colouring(key,G) for key in counter)
return cost / sum(counter.values()) # Divide by total samples
# -
get_cost_colouring('00111', G)
get_cost(example_dict, G)
def func(param,*args):
G = args[0]
C = quantum_loop(param)
cost = get_cost(C,G)
return cost
# ### QAOA for depth $p = 2$
# We now use a classical optimizer `minimize` in order to find the best variational parameters. This function takes as arguments `func`, the graph `G`and an initial `x0` point for the simplex in Nelder-Mead minimization.
res = minimize(func,
args=G,
x0=np.r_[guess['t'], guess['s']],
method='Nelder-Mead',
tol=1e-5,
options = {'maxiter': 100}
)
# We can now plot the sample that we woud obtain using the variational parameters `res.x`.
count_dict = quantum_loop(res.x)
plot_distribution(count_dict)
# QAOA is capable of finding good variational parameters $\tau$ and $t$. Now, sampling from this final state $|\psi(t_{f})\rangle$ will return both MISs of the graph with high probability. Note that listing all maximal independent sets of a graph is also NP, and can be used as a subroutine for solving many NP-complete graph problems.
|
tutorials/applications/Using QAOA to solve a MIS problem.ipynb
|
-- ---
-- jupyter:
-- jupytext:
-- text_representation:
-- extension: .hs
-- format_name: light
-- format_version: '1.5'
-- jupytext_version: 1.14.4
-- kernelspec:
-- display_name: Haskell
-- language: haskell
-- name: haskell
-- ---
:load DescriptiveStats
import Data.List
import Database.HDBC
import Database.HDBC.Sqlite3
import DescriptiveStats
db <- connectSqlite3 "../data/movies.sqlite3"
quickQuery db "SELECT avg(rating) FROM data" []
quickQuery db "SELECT median(rating) FROM data" []
quickQuery db "SELECT mode(rating) FROM data" []
ratingsAction <- quickQuery db "SELECT data.rating FROM data, items WHERE data.itemid=items.movieid AND items.action=1" []
length ratingsAction
readColumn = map fromSql
mean (readColumn ( (head . transpose) ratingsAction) :: [Double])
median (readColumn ( (head . transpose) ratingsAction) :: [Double])
ratingsDrama <- quickQuery db "SELECT data.rating FROM data, items WHERE data.itemid=items.movieid AND items.drama=1" []
mean (readColumn ( (head . transpose) ratingsDrama) :: [Double])
median (readColumn ( (head . transpose) ratingsDrama) :: [Double])
ratingsScifi <- quickQuery db "SELECT data.rating FROM data, items WHERE data.itemid=items.movieid AND items.scifi=1" []
mean (readColumn ( (head . transpose) ratingsScifi) :: [Double])
median (readColumn ( (head . transpose) ratingsScifi) :: [Double])
|
Chapter06/6.2/MovieLens-SQLite3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This notebook demos varios differential operators
# We start with our differential operators
# +
from dusk.script import *
@stencil
def gradient(f: Field[Edge], nx: Field[Edge], ny: Field[Edge], L: Field[Edge], A: Field[Cell], edge_orientation: Field[Cell > Edge],
f_x: Field[Cell], f_y: Field[Cell]):
with levels_downward:
f_x = sum_over(Cell > Edge, f * nx * L * edge_orientation) / A
f_y = sum_over(Cell > Edge, f * ny * L * edge_orientation) / A
@stencil
def divergence(u: Field[Edge], v: Field[Edge], nx: Field[Edge], ny: Field[Edge], L: Field[Edge], A: Field[Cell], edge_orientation: Field[Cell > Edge],
uv_div: Field[Cell]):
with levels_downward:
uv_div = sum_over(Cell > Edge, (u*nx + v*ny) * L * edge_orientation) / A
@stencil
def curl(u: Field[Edge], v: Field[Edge], nx: Field[Edge], ny: Field[Edge], dualL: Field[Edge], dualA: Field[Vertex], edge_orientation: Field[Vertex > Edge],
uv_curl: Field[Vertex]):
with levels_downward:
uv_curl = sum_over(Vertex > Edge, (u*nx + v*ny) * dualL * edge_orientation) / dualA
# -
# Then we can use dusk's Python API to convert the stencils to SIR. This API can also invoke dawn to compile SIR to C++:
from dusk.transpile import callables_to_pyast, pyast_to_sir, sir_to_json
with open("gradient.sir", "w+") as f:
sir = pyast_to_sir(callables_to_pyast([gradient]))
f.write(sir_to_json(sir))
with open("divergence.sir", "w+") as f:
sir = pyast_to_sir(callables_to_pyast([divergence]))
f.write(sir_to_json(sir))
with open("curl.sir", "w+") as f:
sir = pyast_to_sir(callables_to_pyast([curl]))
f.write(sir_to_json(sir))
# !dawn-opt gradient.sir | dawn-codegen -b naive-ico -o gradient_cxx-naive.cpp
# !dawn-opt divergence.sir | dawn-codegen -b naive-ico -o divergence_cxx-naive.cpp
# !dawn-opt curl.sir | dawn-codegen -b naive-ico -o curl_cxx-naive.cpp
# The generated C++ code also requires a driver which is already setup for this demo. With the driver code we can generate an executable `runner`:
# !make
# Now, its up to you which differentail operator you want to run and check. Simply launch `runner gradient`, `runner divergence` or `runner curl`
# !./runner gradient
# Besides ensuring the error norms L1, L2 and L infinity are small (they should all be well below 0.1), you can also have a look at some test functions and their differentials. Again, you can use `check gradient`, `check divergence` or `check curl`. Please make sure that you ran the appropriate differential operator beforhand using the `runner`
# %run checker.py gradient
|
content/differential_ops/differential_ops_solution.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
# **POZNÁMKA**: _Toto je demonštratívne riešenie optimalizačného problému a ide o druhú časť tutoriálu_ [Julia getting started](julia_getting_started.ipynb) s použítím Jupyter zápisníkov a Bindera.
#
# Binder je open-source projekt a Julia je v ňom už od raných začiatkov podporovaná.
#
# [Tento](https://mybinder.org/v2/gh/Dano-drevo/sop-julia-demo/master?filepath=SVM_jl.ipynb)(a iné) `Jupyter zápisníky` si môžete pozrieť a interaktívne spustiť [tu](https://mybinder.org/v2/gh/Dano-drevo/sop-julia-demo/master).
#
# # Support Vector Machine optimization problem
# ## Teoretický úvod
# V tomto zápisníku si naprogramujeme optimalizačný problém **Support Vector Machine(SVM)**, teda metódu podporných vektorov. Naša úloha je [klasifikovať](https://cs.wikipedia.org/wiki/Klasifikace_(um%C4%9Bl%C3%A1_inteligence)) dáta do 2 skupín na základe nášho vstupného trénovacieho datasetu, teda množiny, už takto oklasifikovaných dát. Túto úlohu z demonštratívnych dôvodov skonštruujeme v 2D.
# Predstavme si teda, že máme trénovací dataset, ktorý vieme jednoznačne rozdeliť na dve skupiny dát, povedzme s vlastnosťou $y_1$ a $y_2$. Celá táto úloha slúži na vytvorenie modelu, pomocou ktorého budeme môcť rôzne dáta oklasifikovať, teda rozhodnúť, či majú vlastnosť $y_1$ alebo $y_2$.
# Rozdelený dataset s vlastnosťami $y_1$ a $y_2$ v 2D môžeme interpretovať ako 2 množiny bodov v rovine:
# Príklad v $R^2$:<img src="images/img1.png" style="width: 400px;">
# Na klasifikáciu dát použijeme [lineárny klasifikátor](https://en.wikipedia.org/wiki/Linear_classifier) a pokúsime sa náš trénovací dataset rozdeliť nadrovinou(v 2D priamka), tak aby, čo najlepšie rozdeľovala dáta s vlastnosťou $y_1$ od dát s vlasnosťou $y_2$. Nadrovín, ktoré rozdeľujú dáta môže byť viac:
# <img src="images/img2.png" style="width: 400px;">
# Náš cieľ je teda nájsť rovnicu priamky, ktorá najlepšie oddeľuje tieto 2 skupiny bodov. Rovnica nadroviny $H_i$ je $\underline{\omega_i}\cdot\underline{x}+b_i=0$, kde $\underline{\omega_i}$ je normálový vektor danej nadroviny a $\underline{x}$ je ľubovolný bod z uvažovaného priestoru. Táto rovnica v 2D splýva s rovnicou priamky $ax+by+c=0$
# Nech existuje nami hľadaná nadrovina rozdeĺujúca náš dataset s rovnicou $\underline{\omega}\cdot\underline{x}+b=0$. Potom môžeme zadefinovať nadroviny na hraniciach $y_1$ množiny a $y_2$ množiny, ako:
# $$\underline{\omega_{y_1}}\cdot\underline{x}+b_{y_1}=\delta$$
# $$\underline{\omega_{y_2}}\cdot\underline{x}+b_{y_2}=-\delta$$
# Pre zjednodušenie môžeme za $\delta$ položiť $1$ a ilustrovať to na nasledujúcom obrázku:
# <img src="images/img3.png" style="width: 400px;">
# Po [odvodení](https://www.svm-tutorial.com/2015/06/svm-understanding-math-part-3/), ktoré pre krátkosť a zameranie tohto textu nebudeme uvádzať, zistíme, že šírka pásu oddeľujúceho tieto 2 nadroviny je $\frac{2}{||\omega||}$. Ak chceme maximalizovať šírku tohto pásu, musíme minimalizovať prevrátenú hodnotu, teda $\frac{1}{2}||\omega||$. Tak sa konečne dostávame k samotnej optimalizačnej úlohe.
# ## Formulácia problému
# Položme
# $$
# f(\underline{x}) =\begin{cases}
# 1 & \quad \text{ak } \underline{x}\in y_1 (\text{má vlastnosť }y_1)\\
# -1 & \quad \text{ak } \underline{x}\in y_2 (\text{má vlastnosť }y_2)\end{cases}
# $$
# čím matematicky vyjadríme príslušnosť bodu k množine s danou vlastnosťou. Potom môžeme náš optimalizačný problém pre $n$ bodov zadefinovať nasledovne:
# $${\text{minimize}} \quad\frac{1}{2}||\omega||^2$$
# $${\text{subject to}} \quad f(\underline{x_i})(\underline{\omega}\cdot\underline{x_i}+b) \ge 1\qquad for\ i = 1,\ldots,n$$
# Kvadrátom $\omega$ zaručíme konvexnosť problém, a teda jedinečné globálne minimum, $\underline{\text{ak existuje}}$. Nemáme totiž zaručenú existenciu riešenia. Tento model sa nazýva tzv. "_Hard Margin SVM_"
# Teraz už len ostáva to naprogramovať a nechať Juliu robiť svoju práci.
# ## Implementácia v Julii
# ### Vstupné dáta - trénovací dataset a jeho zobrazenie pomocou balíčku Plots
# Najprv vygenerujeme náš testovací dataset. Použijeme na to pomocnú funkciu zadefinovanú v inom Julia súbore. Všetky 3 pomocné funkcie použité v tomto deme si môžete pozrieť [tu](SVM_utils_jl.ipynb).
# +
include("SVM_utils.jl")
accurancy = 10e-2
y1_data_first = generate_2D_dataset(5, (0,4), (0, 5), accurancy)
y1_data_between = generate_2D_dataset(10, (5,15), (4, 8), accurancy)
y1_data_last = generate_2D_dataset(5, (16,20), (9, 15), accurancy )
y1_data = [y1_data_first; y1_data_between; y1_data_last]
y2_data_first = generate_2D_dataset(5, (0,4), (9, 15), accurancy)
y2_data_between = generate_2D_dataset(10, (5,15), (12, 19), accurancy)
y2_data_last = generate_2D_dataset(5, (16,20), (16, 18), accurancy )
y2_data = [y2_data_first; y2_data_between; y2_data_last]
# -
# **POZNÁMKA**: _Pozor, ak si skúšate vygenerovať nové vlastné dáta interaktívnym spustením tohto bloku kódu, je možné, že takto vygenerované dáta nebude možné lineárne rozdeliť a žiadne prípustné riešenie nebude existovať. Každopádne, aj v tom prípade je zaujímavé vidieť, čo sa stane a ako to dopadne s naším problémom. Eventuálne spúšťajte kód "kým sa to nepodarí"._
#
# Keď nami náhodne vygenerované dáta hodíme do grafu, vyzerá to takto:
# +
using Plots
gr() #choosing backend for plotting
x1_data = [y1_data[:,1] y2_data[:,1]]
x2_data = [y1_data[:,2] y2_data[:,2]]
labels = ["feature 'y1'" "feature 'y2'"]
x1_domain, x2_domain = get_minmax_vals([y1_data;y2_data])
# -
plot(x1_data, x2_data, xlims=x1_domain, ylims=x2_domain, label=labels, legendtitle="Features", seriestype=:scatter, title="Trénovací dataset")
# ### Formulácia problému s optimalizačnou knižnicou Convex
# Máme trénovací dataset, takže môžeme pristúpiť k formulácii problému.
# Ako prvé zadefinujeme **optimalizačné premenné** $\omega$ a $b$, aj s ich dimenziami, **účelovú funkciu** $\frac{1}{2}||\omega||^2$ a problém samotný:
# +
using Convex
omega = Variable(1,2)
b = Variable(1,1)
objective = sumsquares(omega)/2 #sumsquares(x) = ||x||^2 with Euclidian norm
problem = minimize(objective)
# -
# Teraz budeme iterovať cez body nášho datasetu a definovať **obmedzenie** $$f(\underline{x_i})(\underline{\omega}\cdot\underline{x_i}+b) \ge 1\qquad for\ i = 1,\ldots,n$$
# +
data = Dict(1 => y1_data, -1 => y2_data)
constraints = []
for (y, dataset) in data
points_number = size(dataset)[1]
for point_index in collect(1:points_number)
x = dataset[point_index, :]
problem.constraints += y*(omega*x+b) >= 1
end
end
# -
# ### Riešenie úlohy pomocou riešiča ECOS
# Teraz už len vyriešíme našu úlohu, a to pomocou riešiča $ECOS$.
# +
using ECOS
solve!(problem, ECOSSolver())
# -
# Môžeme sa pozrieť na výsledok problému, hodnotu účelovej funkcie a optimalizačných premenných:
println(problem.status, "\n\n", problem.optval, "\n\n", omega, "\n\n", b )
# ### Zhrnutie výsledkov
# Vyriešili sme náš optimalizačný problém a našli sme **optimálne riešenie**. Poďme naše výsledky demonštrovať na predchádzajúcom grafe a zakresliť doňho oddeľujúcu nami nájdenú nadrovinu.
hyperplane = create_2D_linear_function(omega.value, b.value)
line_x_data = collect(x1_domain[1]:0.01:x1_domain[2])
line_y_data = hyperplane.(line_x_data)
plot!(line_x_data, line_y_data, label=["Hyperplane"], seriestype=:line, title="Support Vector Machine")
# Ako vidíme, všetko funguje ako má a naše 2 skupiny bodov s vlastnosťami $y_1$ a $y_2$ sú jednoznačne oddelené našou nadrovinou. Keď teraz dostaneme za úlohu rozhodnúť, ktorú vlastnosť má nejaký bod z tohto priestoru, môžeme na to využiť tento model a priradiť mú vlastnosť podľa toho, na ktorej strane(priestoru oddeľovaného nadrovinou) sa nachádza.
# Úplne rovnaký postup pre nájdenie optimálnej nadroviny by sme mohli zvoliť pre **ľubovoľnú** dimenziu nielen 2D.
# Existenciu riešenia sme zaručenú nemali. Sú však variácie tohto problému, kde pomocou dodatočne zadefinovaných umelých premenných môžeme relaxovať optimalizačné obmedzenia a aj keď dáta nevieme lineárne separovať, stále vieme nájsť nadrovinu "kvázi" vyhovujúcu naším potrebám. Ide o tzv. "_Soft Margin SVM_"
# Sú rôzne variácie SVM. Body sa dajú separovať aj nelineárne, prípadne môžeme penalizovať isté faktory, čím dokážeme model lepšie napasovať na náš problém. Vo všeobecnosti má SVM rôznorodé aplikácie naprieč spektrom odvetví.
# ## Záver
# Opdorúčam, aby ste si pomenili parametre funkcií, poskúšali rôzne situácie a precvičili si tak Juliu. Prípadne model rozšírili na [soft-margin](https://www.saedsayad.com/support_vector_machine.htm) alebo si skúsili naprogramovať vlastnú optimalizačnú úlohu alebo skriptík v Julii. Ak uvažujete v pokračovaní v online precvičovaní Julie, vyskúšajte [JuliaBox](https://juliabox.com/), je zadarmo, len sa musíte prihlásiť, napríklad e-mailom.
|
SVM_jl.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="bwJ8PhXrr5OW" colab_type="text"
# # Bayesian optimization with `skopt`
#
# (based on scikit-optimize documentation https://scikit-optimize.github.io/notebooks/sklearn-gridsearchcv-replacement.html)
# + id="6mH_S_RfsRo-" colab_type="code" outputId="93d4f520-8024-448c-d3ca-be620e457be5" colab={"base_uri": "https://localhost:8080/", "height": 187}
# sklearn version fixed to avoid known skopt issue
# !pip install scikit-optimize scikit-learn==0.20.3
# + id="MnKNBiQTr5Oc" colab_type="code" colab={}
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
from skopt import BayesSearchCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
# + [markdown] id="YnkUBtXcww23" colab_type="text"
# ## Optimising a classifier
# + id="j5-QBPE_s6pg" colab_type="code" outputId="48beac76-e735-4ebd-b41c-d37f8e3055b4" colab={"base_uri": "https://localhost:8080/", "height": 75}
from sklearn.datasets import load_digits
X, y = load_digits(10, True)
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.75,
random_state=0)
# + [markdown] id="531c3cie1Dxf" colab_type="text"
# dimensions for parameters [list, shape=(n_dims,)]: List of search space dimensions. Each search dimension can be defined either as
#
# - a (lower_bound, upper_bound) tuple (for Real or Integer dimensions),
# - a (lower_bound, upper_bound, prior) tuple (for Real dimensions),
# - as a list of categories (for Categorical dimensions), or
# - an instance of a Dimension object (Real, Integer or Categorical).
# + id="h8UFVdNmyXS9" colab_type="code" colab={}
param_dist = {
"max_depth": (3, 10,),
"max_features": (1, 11),
"min_samples_split": <YOUR CODE>, # from 2 to 10
"min_samples_leaf": <YOUR CODE>, # from 1 to 10
"bootstrap": [True, False], # categorical valued parameter
"criterion": <YOUR CODE> # either "gini" or "entropy"
}
# + id="KVdB96xzzOcY" colab_type="code" colab={}
clf = RandomForestClassifier(n_estimators=20)
opt = BayesSearchCV(clf, param_dist, n_iter=10, return_train_score=True, cv=3)
# + id="Y24slK8AzUG6" colab_type="code" colab={}
opt.fit(X_train, y_train);
# + id="7aY681Q_xiPD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} outputId="38d12d49-6e3a-4ab8-e121-fd478c358a64"
print("val. score: %s" % opt.best_score_)
print("test score: %s" % opt.score(X_test, y_test))
# + id="ySuKwUxu1hHh" colab_type="code" colab={}
# Utility function to report best scores
import pandas as pd
def report(results, n_top=3):
res = pd.DataFrame(results)
res = res.sort_values(by=['mean_test_score'], ascending=False, axis=0)
res.reset_index(inplace = True, drop=True)
# a = res[['mean_test_score', 'std_test_score']]
for candidate in range(0, n_top):
print("Model with rank: {0}".format(candidate))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
res['mean_test_score'][candidate],
res['std_test_score'][candidate]))
print("Parameters: {0}".format(res['params'][candidate]))
print("")
# + id="Ip5ROztL1mPp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 261} outputId="3d12931d-0290-44be-ce58-d7fa78d97246"
report(opt.cv_results_)
# + [markdown] id="56dsimIm0KAt" colab_type="text"
# ## Task
# Optimise the neural net from the previous notebook via `BayesSearchCV`
#
# + id="NHyak9rb8NbZ" colab_type="code" colab={}
import torch
from torch import nn
import torch.nn.functional as F
from skorch import NeuralNetClassifier
torch.manual_seed(0);
# + id="QyVl7Ii38bEt" colab_type="code" colab={}
from sklearn.datasets import make_classification
X, y = make_classification(1000, 20, n_informative=10, n_classes=2, random_state=0)
X = X.astype(np.float32)
# + id="EL8NhGT48LMW" colab_type="code" colab={}
class ClassifierModule(nn.Module):
<CODE OF THE CLASSIFIER FROM NOTEBOOK i-1>
# + id="LXv7qEAh8iD-" colab_type="code" colab={}
net = NeuralNetClassifier(
ClassifierModule,
max_epochs=20,
lr=0.1,
device='cuda', # comment this to train with CPU
optimizer__momentum=0.9,
verbose=0
)
# + id="ZBEXtAIC8j0W" colab_type="code" colab={}
params = {
'lr': [0.05, 0.1],
'module__num_units': [10, 20, 30], # range from 10 to 50
'module__dropout': [0.1, 0.3], # range from 0.1 to 0.3
'optimizer__nesterov': [False, True],
}
# + id="E3lT0R1681FF" colab_type="code" colab={}
bs = BayesSearchCV(net, params, refit=False, cv=3, scoring='accuracy',
verbose=0, n_jobs=1, n_iter=10, return_train_score=True)
# + id="jSoqTdZr8717" colab_type="code" colab={}
bs.fit(X, y);
# + id="lcszRHhc8_qh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 261} outputId="f5ea4743-8783-4552-bc35-300a73b8cbf3"
report(bs.cv_results_)
|
3-bayesian-optimization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp quantification
# -
# # Quantification
#
# > Functions related to quantification
#hide
from nbdev.showdoc import *
# ## Label-free quantification
#
# Algorithms related to label-free quantifications are motivated by the [MaxLFQ paper](https://doi.org/10.1074/mcp.m113.031591). The main goal is to derive relative protein intensities that can be used for downstream analyses. In a first step, constant normalization coefficients are derived for each run. In a second step, pseudointensities are derived for each protein, such that differing conditions can be compared.
# ## Delayed Normalization
#
# Delayed normalization describes the process of normalizing the differences that occur from prefractionation as well as from sample handling. For each sample, a constant scaling factor is derived by minimizing the term
# $$H(\vec{N}) = \sum_{P \in peptides} \sum_{A,B \in sample pairs} |\frac{I(N_A, P, A)}{I(N_B, P, B)}|, $$
# with peptide intensities $I$, which are determined by the peptide $P$ the sample $A$ or $B$ and the normalization factors $N_A$, $N_B$. In principle H(N) quantifies the variation of peptides over the samples. Minimizing this variation gives appropriate scaling factors under the assumption that most peptides do not change between the samples. Peptide intensities for fractionated samples are described as the sum of the intensities over the fractions, with fraction-specific normalization factors. Therefore, calculation of the summed intensities is *delayed* until the normalization is finished.
# ### In Silico Test data
#
# To test the delayed normalization approach we create an in silico test dataset with a known ground truth. We therefore know, which systematic changes are between the samples and we employ different solvers to recover the normalization parameters.
# +
#export
import random
import numpy as np
import logging
def gaussian(mu: float, sigma: float, grid : np.ndarray) -> np.ndarray:
"""Calculates normally distributed probability densities along an input array.
Args:
mu (float): mean of ND.
sigma (float): standard deviation of ND.
grid (np.ndarray): input array np.int[:]. For each element of the array, the probability density is calculated.
Returns:
np.ndarray: probability density array, np.float[:].
"""
norm = 0.3989422804014327 / sigma
return norm * np.exp(-0.5 * ((grid - mu) / sigma) ** 2)
# +
#hide
def test_gaussian():
assert np.allclose(gaussian(10, 3, np.arange(5)), np.array([0.00051409, 0.00147728, 0.00379866, 0.00874063, 0.01799699]))
assert np.allclose(gaussian(1, 3, np.arange(6)), np.array([0.12579441, 0.13298076, 0.12579441, 0.10648267, 0.08065691, 0.05467002]))
test_gaussian()
# -
#export
def return_elution_profile(timepoint: float, sigma : float, n_runs : int) -> np.ndarray:
"""Simulates a gaussian elution profile.
Args:
timepoint (float): coordinate of the peak apex.
sigma (float): standard deviation of the gaussian.
n_runs (int): number of points along which the density is calculated.
Returns:
np.ndarray: probability density array, np.float[:].
"""
return gaussian(timepoint, sigma, np.arange(0, n_runs))
# +
#hide
def test_return_elution_profile():
assert np.allclose(return_elution_profile(10, 2, 6), np.array([7.43359757e-07, 7.99187055e-06, 6.69151129e-05, 4.36341348e-04,
2.21592421e-03, 8.76415025e-03]))
assert np.allclose(return_elution_profile(1, 5, 3), np.array([0.07820854, 0.07978846, 0.07820854]))
test_return_elution_profile()
# -
#export
def simulate_sample_profiles(n_peptides: int, n_runs: int, n_samples: int, threshold:float=0.2, use_noise:bool=True) -> [np.ndarray, np.ndarray]:
"""Generates random profiles to serve as test_data.
Args:
n_peptides (int): number of peptides to be simulated.
n_runs (int): number of runs to be simulated.
n_samples (int): number of samples to be simulated.
threshold (float, optional): threshold below which a simulated intensity will be discarded. Defaults to 0.2.
use_noise (bool, optional): add simulated noise to the profile values. Defaults to True.
Returns:
Tuple[np.ndarray, np.ndarray]: profiles: np.float[:,:,:] array containing the simulated profiles, true_normalization: np.float[:,:,:] array containing the ground truth.
"""
np.random.seed(42)
abundances = np.random.rand(n_peptides)*10e7
true_normalization = np.random.normal(loc=1, scale=0.1, size=(n_runs, n_samples))
true_normalization[true_normalization<0] = 0
true_normalization = true_normalization/np.max(true_normalization)
maxvals = np.max(true_normalization, axis=1)
elution_timepoints = random.choices(list(range(n_runs)), k=n_peptides)
profiles = np.empty((n_runs, n_samples, n_peptides))
profiles[:] = np.nan
for i in range(n_peptides):
elution_timepoint = elution_timepoints[i]
abundance = abundances[i]
profile = return_elution_profile(elution_timepoint, 1, n_runs)
profile = profile/np.max(profile)
profile = profile * abundance
elution_profiles = np.tile(profile, (n_samples, 1)).T
# Add Gaussian Noise
if use_noise:
noise = np.random.normal(1, 0.2, elution_profiles.shape)
noisy_profile = noise * elution_profiles
else:
noisy_profile = elution_profiles
normalized_profile = noisy_profile * true_normalization
normalized_profile[normalized_profile < threshold] = 0
normalized_profile[normalized_profile == 0] = np.nan
profiles[:,:,i] = normalized_profile
return profiles, true_normalization
# +
#hide
def test_simulate_sample_profiles():
#The function to be tested is a random generator so we only test the output format here
n_peptides = 2
n_runs = 5
n_samples = 10
profiles, true_normalization = simulate_sample_profiles(n_peptides, n_runs, n_samples)
assert profiles.shape == (n_runs, n_samples, n_peptides)
assert true_normalization.shape == (n_runs, n_samples)
assert np.all(profiles > 0)
assert np.all(true_normalization > 0)
test_simulate_sample_profiles()
# -
# ## Delayed Normalization
# +
#export
from numba import njit, prange
@njit
def get_peptide_error(profile: np.ndarray, normalization: np.ndarray) -> float:
"""Distance function for least squares optimization. Calculates the peptide ratios between samples. Smaller ratios mean better normalization.
Args:
profile (np.ndarray): peptide intensity values.
normalization (np.ndarray): per sample normalization factors.
Returns:
float: summed squared error.
"""
pep_ints = np.zeros(profile.shape[1])
normalized_profile = profile*normalization
for i in range(len(pep_ints)):
pep_ints[i] = np.nansum(normalized_profile[:,i])
pep_ints = pep_ints[pep_ints>0]
# Loop through all combinations
n = len(pep_ints)
error = 0
for i in range(n):
for j in range(i+1,n):
error += np.abs(np.log(pep_ints[i]/pep_ints[j]))**2
return error
# +
#hide
def test_get_peptide_error():
profile = np.ones((10, 10))
normalization = np.ones((10))
assert get_peptide_error(profile, normalization) == 0
normalization = np.arange((10))
assert np.allclose(get_peptide_error(profile, normalization), 37.24832444019646)
test_get_peptide_error()
# -
#export
def get_total_error(normalization: np.ndarray, profiles: np.ndarray) -> float:
"""Computes the summed peptide errors over the whole dataset.
Args:
normalization (np.ndarray): per sample normalization factors.
profiles (np.ndarray): peptide intensity profiles over the dataset.
Returns:
float: summed peptide error.
"""
normalization = normalization.reshape(profiles.shape[:2])
total_error = 0
for index in range(profiles.shape[2]):
total_error += get_peptide_error(profiles[:,:, index], normalization)
return total_error
# +
#hide
def test_get_total_error():
profiles = np.ones((10, 10, 4))
normalization = np.ones((10, 10))
assert get_total_error(normalization, profiles) == 0
normalization = np.array([np.arange(10) for i in range(10)])
assert np.allclose(get_total_error(normalization, profiles), 4*37.24832444019646)
test_get_total_error()
# -
# ## Benchmarking different optimiziers
# The normalization step is in principle a quadratic minimization of the normalization factors. Such minimization problems can be solved in various ways and a variety of approaches are realized in python community packages. We compare different solvers using our benchmarking set and uncover substantial differences in precision and runtime. We observe that the *Sequential Least Squares Quadratic Programming* (SLSQP) approach is a robust solution in our benchmarking, which gives substantial speed improvements.
# +
from scipy.optimize import minimize
from time import time
from scipy.optimize import least_squares
import pandas as pd
import warnings
n_peptides = 100
n_runs = 10
n_samples = 3
profiles, true_normalization = simulate_sample_profiles(n_peptides, n_runs, n_samples)
methods = ['L-BFGS-B', 'TNC', 'SLSQP','trf']
results = []
for method in methods:
start = time()
with warnings.catch_warnings():
warnings.filterwarnings("ignore", category=RuntimeWarning)
if method in ['trf']:
x0 = np.ones(profiles.shape[0] * profiles.shape[1])
bounds = (x0*0.1, x0)
res = least_squares(get_total_error, args = [profiles], bounds = bounds, x0 = x0*0.5, verbose=0, method = method)
else:
x0 = np.ones(profiles.shape[0] * profiles.shape[1])
bounds = [(0.1, 1) for _ in x0]
res = minimize(get_total_error, args = profiles , x0 = x0*0.5, bounds=bounds, method=method)
solution = res.x/np.max(res.x)
solution = solution.reshape(profiles.shape[:2])
end = time()
time_elapsed_min = (end-start)/60
optimality = get_total_error(solution, profiles) /get_total_error(x0, profiles)
optimality_ = get_total_error(solution, profiles) / get_total_error(true_normalization, profiles)
results.append((method, time_elapsed_min, optimality, optimality_))
pd.DataFrame(results, columns=['Method', 'Time Elapsed (min)','Error / Baseline Error','Error / Ground Truth'])
# +
#export
from scipy.optimize import minimize
import pandas as pd
import numpy as np
import warnings
def normalize_experiment_SLSQP(profiles: np.ndarray) -> np.ndarray:
"""Calculates normalization with SLSQP approach.
Args:
profiles (np.ndarray): peptide intensities.
Returns:
np.ndarray: normalization factors.
"""
x0 = np.ones(profiles.shape[0] * profiles.shape[1])
bounds = [(0.1, 1) for _ in x0]
res = minimize(get_total_error, args = profiles , x0 = x0*0.5, bounds=bounds, method='SLSQP', options={'disp': False} )
solution = res.x/np.max(res.x)
solution = solution.reshape(profiles.shape[:2])
return solution
# +
#hide
def test_normalize_experiment_SLSQP():
n_peptides = 15
n_runs = 5
n_samples = 20
profiles, true_normalization = simulate_sample_profiles(n_peptides, n_runs, n_samples)
x0 = np.ones(profiles.shape[0] * profiles.shape[1])
with warnings.catch_warnings():
warnings.filterwarnings("ignore", category=RuntimeWarning)
solution = normalize_experiment_SLSQP(profiles)
optimality = get_total_error(solution, profiles) / get_total_error(x0, profiles)
assert optimality < 1
test_normalize_experiment_SLSQP()
# -
#export
def normalize_experiment_BFGS(profiles: np.ndarray) -> np.ndarray:
"""Calculates normalization with BFGS approach.
Args:
profiles (np.ndarray): peptide intensities.
Returns:
np.ndarray: normalization factors.
"""
x0 = np.ones(profiles.shape[0] * profiles.shape[1])
bounds = [(0.1, 1) for _ in x0]
res = minimize(get_total_error, args = profiles , x0 = x0*0.5, bounds=bounds, method='L-BFGS-B', options={'disp': False} )
solution = res.x/np.max(res.x)
solution = solution.reshape(profiles.shape[:2])
return solution
# +
#hide
def test_normalize_experiment_BFGS():
n_peptides = 15
n_runs = 5
n_samples = 20
profiles, true_normalization = simulate_sample_profiles(n_peptides, n_runs, n_samples)
x0 = np.ones(profiles.shape[0] * profiles.shape[1])
with warnings.catch_warnings():
warnings.filterwarnings("ignore", category=RuntimeWarning)
solution = normalize_experiment_BFGS(profiles)
optimality = get_total_error(solution, profiles) / get_total_error(x0, profiles)
assert optimality < 1
test_normalize_experiment_BFGS()
# -
#export
def delayed_normalization(df: pd.DataFrame, field: str='int_sum', minimum_occurence:bool=None) -> [pd.DataFrame, np.ndarray]:
"""Returns normalization factors for given peptide intensities.
If the solver does not converge, the unnormalized data will be used.
Args:
df (pd.DataFrame): alphapept quantified features table.
field (str, optional): The column in df containing the quantitative peptide information (i.e. precursor intensities).
minimum_occurence (bool, optional): minimum number of replicates the peptide must be observed in. Defaults to None.
Returns:
[pd.DataFrame, np.ndarray]: pd.DataFrame: alphapept quantified features table extended with the normalized intensities, np.ndarray: normalized intensities
"""
files = np.sort(df['filename'].unique()).tolist()
n_files = len(files)
if 'fraction' not in df.keys():
df['fraction'] = [1 for x in range(len(df.index))]
fractions = np.sort(df['fraction'].unique()).tolist()
n_fractions = len(fractions)
df_max = df.groupby(['precursor','fraction','filename'])[field].max() #Maximum per fraction
prec_count = df_max.index.get_level_values('precursor').value_counts()
if not minimum_occurence:
minimum_occurence = np.percentile(prec_count[prec_count>1].values, 75) #Take the 25% best datapoints
logging.info('Setting minimum occurence to {}'.format(minimum_occurence))
shared_precs = prec_count[prec_count >= minimum_occurence]
precs = shared_precs.index.tolist()
n_profiles = len(precs)
selected_precs = df_max.loc[precs]
selected_precs = selected_precs.reset_index()
profiles = np.empty((n_fractions, n_files, n_profiles))
profiles[:] = np.nan
#get dictionaries
fraction_dict = {_:i for i,_ in enumerate(fractions)}
filename_dict = {_:i for i,_ in enumerate(files)}
precursor_dict = {_:i for i,_ in enumerate(precs)}
prec_id = [precursor_dict[_] for _ in selected_precs['precursor']]
frac_id = [fraction_dict[_] for _ in selected_precs['fraction']]
file_id = [filename_dict[_] for _ in selected_precs['filename']]
profiles[frac_id,file_id, prec_id] = selected_precs[field]
with warnings.catch_warnings():
warnings.filterwarnings("ignore", category=RuntimeWarning)
try:
normalization = normalize_experiment_SLSQP(profiles)
norm1d = np.ravel(normalization)
if sum((norm1d!=1))==0:
raise ValueError("optimization with SLSQP terminated at initial values. Trying BFGS")
except ValueError: # SLSQP error in scipy https://github.com/scipy/scipy/issues/11403
logging.info('Normalization with SLSQP failed. Trying BFGS')
normalization = normalize_experiment_BFGS(profiles)
norm1d = np.ravel(normalization)
if sum((norm1d!=1))==0:
logging.warn('No normalization factors could be determined. Continuing with non-normalized data.')
#intensity normalization: total intensity to remain unchanged
df[field+'_dn'] = df[field]*normalization[[fraction_dict[_] for _ in df['fraction']], [filename_dict[_] for _ in df['filename']]]
df[field+'_dn'] *= df[field].sum()/df[field+'_dn'].sum()
return df, normalization
# +
sample_data = {}
sample_data['precursor'] = ['Prec_1'] * 6 + ['Prec_2'] * 6 + ['Prec_3'] * 6
sample_data['fraction'] = [1,1,2]*6
sample_data['filename'] = ['A','A','A', 'B','B','B'] * 3
sample_data['int_sum'] = [0.6, 0.8, 0.6, 1.2, 1.6, 1.2] * 3
test_df = pd.DataFrame(sample_data)
test_df, normalization = delayed_normalization(test_df, field='int_sum', minimum_occurence=0)
display(pd.DataFrame(normalization))
display(test_df.head(6))
# +
#hide
def test_delayed_normalization():
sample_data = {}
sample_data['precursor'] = ['Prec_1'] * 6 + ['Prec_2'] * 6 + ['Prec_3'] * 6
sample_data['fraction'] = [1,1,2]*6
sample_data['filename'] = ['A','A','A', 'B','B','B'] * 3
sample_data['int_sum'] = [0.6, 0.8, 0.6, 1.2, 1.6, 1.2] * 3
test_df = pd.DataFrame(sample_data)
test_df, normalization = delayed_normalization(test_df, field='int_sum', minimum_occurence=0)
assert normalization.shape == (2,2)
test_delayed_normalization()
# -
# ## Constructing protein intensity profiles
# Protein intensity profiles are constructed for each protein individually. All possible protein fold changes between the samples are derived from the median peptide fold changes. Subsequently, pseudointensities are chosen such that the fold changes between the pseudointensities ideally reconstruct the actually observed fold changes. Similar to the delayed normalization, this is formulated as a quadratic minimization, which we solve with the SLSQP solver.
#
# Codewise, we start with simulating in-silico test data to serve as a ground-truth for assessing solvers for the optimization problem. For the algorithmic optimization, we define the function `get_protein_ratios` that allows to quickly calculate the protein ratios. Next, we define an error function `triangle_error` that we use for the optimization problem. Lastly, we have several wrapper functions to access the functions.
# ## In-silico test data
#
# Create a simulated input dataset of peptide intensities.
# +
#export
import numpy as np
import string
from time import time
import pandas as pd
np.random.seed(42)
def generate_dummy_data(n_sequences: int, n_samples: int, noise:bool=True, remove:bool= True, peptide_ratio:bool= True, abundance:bool=True, signal_level:int=100, noise_divider:int=10, keep:float=0.8) -> [pd.DataFrame, list, np.ndarray]:
"""Simulate an input dataset of peptide intensities.
Args:
n_sequences (int): number of peptides to simulate.
n_samples (int): number of samples to simulate.
noise (bool, optional): add random signal to distort the simulated intensity levels. Defaults to True.
remove (bool, optional): remove intensities (i.e. add missing values). Defaults to True.
peptide_ratio (bool, optional): simulate different peptide intensities. Defaults to True.
abundance (bool, optional): simulate different abundances for each sample (i.e. systematic shifts). Defaults to True.
signal_level (int, optional): signal level for simulated intensity. Defaults to 100.
noise_divider (int, optional): the factor through which the noise is divided (higher factor -> higher signal to noise). Defaults to 10.
keep (float, optional): aimed-at fraction of non-missing values, applies if 'remove' is set. Defaults to 0.8.
Returns:
[pd.DataFrame, list, np.ndarray]: pd.DataFrame: simulated dataset with peptide intensities, list: sample names: np.ndarray: shift factors of each sample
"""
species = ['P'+str(_) for _ in range(1,n_sequences+1)]
sample = [string.ascii_uppercase[_%26]+str(_//26) for _ in range(n_samples)]
if peptide_ratio:
peptide_ratio = np.random.rand(n_sequences)
peptide_ratio = peptide_ratio/np.sum(peptide_ratio)
else:
peptide_ratio = np.ones(n_sequences)
if abundance:
abundance_profile = np.random.rand(n_samples,1)
else:
abundance_profile = np.ones((n_samples,1))
original_signal = np.ones((n_samples, n_sequences))
noise_sim = (np.random.rand(n_samples, n_sequences)-0.5)/noise_divider
if noise:
noisy_signal = original_signal+noise_sim
noisy_signal = noisy_signal*signal_level*peptide_ratio*abundance_profile
else:
noisy_signal = original_signal*signal_level*peptide_ratio*abundance_profile
if remove:
#Remove points
keep_probability = keep #keep 60% of the points
to_remove = np.random.rand(n_samples, n_sequences)
to_remove = to_remove>=keep_probability
dummy_data = noisy_signal.copy()
dummy_data[to_remove] = 0
else:
dummy_data = noisy_signal
dummy_data = pd.DataFrame(dummy_data, index = sample, columns = species).T
ground_truth = abundance_profile.flatten()
ground_truth = ground_truth/np.max(ground_truth)
return dummy_data, sample, ground_truth
# +
#hide
def test_generate_dummy_data():
#The function to be tested is a random-generateor, so we test the output format here.
n_samples = 10
n_sequences = 5
dummy_data, sample, ground_truth = generate_dummy_data(n_samples, n_sequences)
assert dummy_data.shape == (n_samples, n_sequences)
assert len(sample) == n_sequences
assert len(ground_truth) == n_sequences
test_generate_dummy_data()
# -
# ## Determine pair-wise intenisty ratios
# The pair-wise protein ratios are determined from the median peptide ratio.
# +
#export
from numba import njit
@njit
def get_protein_ratios(signal: np.ndarray, column_combinations: list, minimum_ratios:int = 1) -> np.ndarray:
"""Calculates the protein ratios between samples for one protein.
Args:
signal (np.ndarray): np.array[:,:] containing peptide intensities for each sample.
column_combinations (list): list of all index combinations to compare (usually all sample combinations).
minimum_ratios (int, optional): minimum number of peptide ratios necessary to calculate a protein ratio. Defaults to 1.
Returns:
np.ndarray: np.array[:,:] matrix comparing the ratios for all column combinations.
"""
n_samples = signal.shape[1]
ratios = np.empty((n_samples, n_samples))
ratios[:] = np.nan
for element in column_combinations:
i = element[0]
j = element[1]
ratio = signal[:,j] / signal[:,i]
non_nan = np.sum(~np.isnan(ratio))
if non_nan >= minimum_ratios:
ratio_median = np.nanmedian(ratio)
else:
ratio_median = np.nan
ratios[j,i] = ratio_median
return ratios
# +
#hide
from itertools import combinations
from numba.typed import List
def test_get_protein_ratios():
n_samples = 5
n_peptides = 2
signal = np.ones((n_samples, n_peptides))
column_combinations = List([_ for _ in combinations(range(n_samples), 2)])
ratios = get_protein_ratios(signal, column_combinations)
assert ratios[1,0] == 1
signal[:,1]*=2
ratios = get_protein_ratios(signal, column_combinations)
assert ratios[1,0] == 2
#test_get_protein_ratios() #TODO: this test seems to break the CI
# -
# ## Error Function
# The error function evaluates the difference between the actual observed fold change and the fold change that is derived from the pseudointensities.
#export
@njit
def triangle_error(normalization: np.ndarray, ratios:np.ndarray) -> float:
"""Calculates the difference between calculated ratios and expected ratios.
Args:
normalization (np.ndarray): Used normalization.
ratios (np.ndarray): Peptide ratios.
Returns:
float: summed quadratic difference.
"""
int_matrix = np.repeat(normalization, len(normalization)).reshape((len(normalization), len(normalization))).transpose()
x = (np.log(ratios) - np.log(int_matrix.T) + np.log(int_matrix))**2
return np.nansum(x)
# +
#hide
def test_triangle_error():
n_samples = 5
n_peptides = 4
signal = np.ones((n_samples, n_peptides))
column_combinations = List([_ for _ in combinations(range(n_samples), 2)])
ratios = get_protein_ratios(signal, column_combinations)
x0 = np.ones(ratios.shape[1])
assert triangle_error(x0, ratios) == 0
signal[:,1]*=2
ratios = get_protein_ratios(signal, column_combinations)
x0 = np.ones(ratios.shape[1])
assert np.allclose(triangle_error(x0, ratios), 1.441359041754604)
#test_triangle_error() #TODO: this test seems to break the CI
# -
# ## Solver implementation
# As with the delayed normalization we implement multiple solvers from scipy.
# +
#export
## L-BFGS-B
from scipy.optimize import minimize, least_squares
def solve_profile(ratios: np.ndarray, method: str) -> [np.ndarray, bool]:
"""Calculates protein pseudointensities with a specified solver.
Args:
ratios (np.ndarray): np.array[:,:] matrix containing all estimated protein ratios between samples.
method (str): string specifying which solver to use.
Raises:
NotImplementedError: if the solver is not implemented.
Returns:
[np.ndarray, bool]: np.ndarray: the protein pseudointensities, bool: wether the solver was successful.
"""
if method not in ['L-BFGS-B', 'SLSQP', 'Powell', 'trust-constr','trf']:
raise NotImplementedError(method)
x0 = np.ones(ratios.shape[1])
bounds = [(min(np.nanmin(ratios), 1/np.nanmax(ratios)), 1) for _ in x0]
with warnings.catch_warnings():
warnings.filterwarnings("ignore", category=RuntimeWarning)
if method == 'trf':
bounds = (x0*0+0.01, x0)
res_wrapped = least_squares(triangle_error, args = [ratios] , x0 = x0, bounds=bounds, verbose=0, method = 'trf')
solution = res_wrapped.x
else:
res_wrapped = minimize(triangle_error, args = ratios , x0 = x0, bounds=bounds, method = method)
solution = res_wrapped.x
solution = solution/np.max(solution)
return solution, res_wrapped.success
# +
#hide
def test_solve_profile():
# The tested function is a wrapper for scipy, so we only test execution here
n_samples = 5
n_peptides = 2
signal = np.ones((n_samples, n_peptides))
column_combinations = List([_ for _ in combinations(range(n_samples), 2)])
ratios = get_protein_ratios(signal, column_combinations)
for method in ['L-BFGS-B', 'SLSQP', 'Powell', 'trust-constr','trf']:
solve_profile(ratios, method)
#test_solve_profile() #TODO: this test seems to break the CI
# -
# ## Solving single profiles
# +
#export
from numba.typed import List
from itertools import combinations
import pandas as pd
def protein_profile(files: list, minimum_ratios: int, chunk:tuple) -> (np.ndarray, np.ndarray, str):
"""Function to extract optimal protein ratios for a given input of peptides.
Note for the chunk argument: This construction is needed to call this function from a parallel pool.
Args:
files (list): A list of files for which the profile shall be extracted.
minimum_ratios (int): A minimum number of peptide ratios to be considered for optimization.
chunk: (tuple[pd.DataFrame, str]): A pandas dataframe with the peptide information and a string to identify the protein.
Returns:
np.ndarray: optimized profile
np.ndarray: profile w/o optimization
str: protein identifier
"""
grouped, protein = chunk
column_combinations = List()
[column_combinations.append(_) for _ in combinations(range(len(files)), 2)]
selection = grouped.unstack().T.copy()
selection = selection.replace(0, np.nan)
if not selection.shape[1] == len(files):
selection[[_ for _ in files if _ not in selection.columns]] = np.nan
selection = selection[files]
ratios = get_protein_ratios(selection.values, column_combinations, minimum_ratios)
retry = False
try:
solution, success = solve_profile(ratios, 'L-BFGS-B')
except ValueError:
retry = True
if retry or not success:
logging.info('Normalization with L-BFGS-B failed. Trying Powell')
solution, success = solve_profile(ratios, 'Powell')
pre_lfq = selection.sum().values
if not success or np.sum(~np.isnan(ratios)) == 0: # or np.sum(solution) == len(pre_lfq):
profile = np.zeros_like(pre_lfq)
if np.sum(np.isnan(ratios)) != ratios.size:
logging.info(f'Solver failed for protein {protein} despite available ratios:\n {ratios}')
else:
invalid = ((np.nansum(ratios, axis=1) == 0) & (np.nansum(ratios, axis=0) == 0))
total_int = pre_lfq.sum() * solution
total_int[invalid] = 0
profile = total_int * pre_lfq.sum() / np.sum(total_int) #Normalize inensity again
return profile, pre_lfq, protein
# +
import matplotlib.pyplot as plt
sample_data = {}
sample_data['precursor'] = ['Prec_1'] * 3 + ['Prec_2'] * 3 + ['Prec_3'] * 3
sample_data['filename'] = ['A','B','C'] * 3
sample_data['protein_group'] = ['X'] * 9
sample_data['int_sum'] = [0.6, 0.8, 1.0, 0.6, 1.2, 1.4, 1.6, 1.2, 1.8]
test_df = pd.DataFrame(sample_data)
display(test_df.head(6))
grouped = test_df.groupby(['protein_group','filename','precursor']).sum().loc['X']
files = ['A','B','C']
minimum_ratios = 1
chunk = (grouped, 'X')
if False: #TODO: this test seems to break the CI
profile, pre_lfq, protein = protein_profile(files, minimum_ratios, chunk)
plt.figure(figsize=(5,5))
plt.title('Protein ratio')
plt.plot(pre_lfq, 'o', label='before optimization')
plt.plot(profile, 'o', label='after optimization')
plt.legend()
plt.show()
# +
#hide
def test_protein_profile():
sample_data = {}
sample_data['precursor'] = ['Prec_1'] * 6 + ['Prec_2'] * 6 + ['Prec_3'] * 6
sample_data['fraction'] = [1,2,3]*6
sample_data['filename'] = ['A','A','A', 'B','B','B'] * 3
sample_data['protein_group'] = ['X'] * 18
sample_data['int_sum'] = [0.6, 0.8, 0.6, 1.2, 1.6, 1.2] * 3
test_df = pd.DataFrame(sample_data)
grouped = test_df.groupby(['protein_group','filename','precursor']).sum().loc['X']
files = ['A','B']
minimum_ratios = 1
chunk = (grouped, 'X')
profile, pre_lfq, protein = protein_profile(files, minimum_ratios, chunk)
assert np.allclose(profile.sum(), pre_lfq.sum())
sample_data = {}
sample_data['precursor'] = ['Prec_1'] * 2 + ['Prec_2'] * 2 + ['Prec_3'] * 2
sample_data['filename'] = ['A','B'] * 3
sample_data['protein_group'] = ['X'] * 6
sample_data['int_sum'] = [0.6, 0.8, 0.6, 1.2, 1.6, 1.2]
test_df = pd.DataFrame(sample_data)
grouped = test_df.groupby(['protein_group','filename','precursor']).sum().loc['X']
files = ['A','B']
minimum_ratios = 1
chunk = (grouped, 'X')
profile, pre_lfq, protein = protein_profile(files, minimum_ratios, chunk)
assert np.allclose(profile.sum(), pre_lfq.sum())
test_protein_profile()
# -
# ## Wrapper functions
#
# To be compatible with interface, we have three wrapper functions:
#
# * protein_profile_parallel: A wrapper that executes protein_profile in parallel
# * protein_profile_parallel_ap: A wrapper function to calculate protein ratios based on AlphaPept tabular data
# * protein_profile_prallalel_mq: A wrapper function to calculate protein ratios based on MaxQuant tabular data
# +
#export
import os
import alphapept.performance
from functools import partial
# This function invokes a parallel pool and has therfore no dedicated test in the notebook
def protein_profile_parallel(df: pd.DataFrame, minimum_ratios: int, field: str, callback=None) -> pd.DataFrame:
"""Derives LFQ intensities from the feature table.
Args:
df (pd.DataFrame): Feature table by alphapept.
minimum_ratios (int): Minimum number of peptide ratios necessary to derive a protein ratio.
field (str): The field containing the quantitative peptide information (i.e. precursor intensities).
callback ([type], optional): Callback function. Defaults to None.
Returns:
pd.DataFrame: table containing the LFQ intensities of each protein in each sample.
"""
unique_proteins = df['protein_group'].unique().tolist()
files = df['filename'].unique().tolist()
files.sort()
columnes_ext = [_+'_LFQ' for _ in files]
protein_table = pd.DataFrame(index=unique_proteins, columns=columnes_ext + files)
grouped = df[[field, 'filename','precursor','protein_group']].groupby(['protein_group','filename','precursor']).sum()
column_combinations = List()
[column_combinations.append(_) for _ in combinations(range(len(files)), 2)]
files = df['filename'].unique().tolist()
files.sort()
results = []
if len(files) > 1:
logging.info('Preparing protein table for parallel processing.')
split_df = []
for idx, protein in enumerate(unique_proteins):
split_df.append((grouped.loc[protein], protein))
if callback:
callback((idx+1)/len(unique_proteins)*1/5)
results = []
logging.info(f'Starting protein extraction for {len(split_df)} proteins.')
n_processes = alphapept.performance.set_worker_count(
worker_count=0,
set_global=False
)
with alphapept.performance.AlphaPool(n_processes) as p:
max_ = len(split_df)
for i, _ in enumerate(p.imap_unordered(partial(protein_profile, files, minimum_ratios), split_df)):
results.append(_)
if callback:
callback((i+1)/max_*4/5+1/5)
for result in results:
profile, pre_lfq, protein = result
protein_table.loc[protein, [_+'_LFQ' for _ in files]] = profile
protein_table.loc[protein, files] = pre_lfq
protein_table[protein_table == 0] = np.nan
protein_table = protein_table.astype('float')
else:
protein_table = df.groupby(['protein_group'])[field].sum().to_frame().reset_index()
protein_table = protein_table.set_index('protein_group')
protein_table.index.name = None
protein_table.columns=[files[0]]
if callback:
callback(1)
return protein_table
# +
#export
# This function invokes a parallel pool and has therfore no dedicated test in the notebook
def protein_profile_parallel_ap(settings: dict, df : pd.DataFrame, callback=None) -> pd.DataFrame:
"""Derives protein LFQ intensities from the alphapept quantified feature table
Args:
settings (dict): alphapept settings dictionary.
df (pd.DataFrame): alphapept feature table.
callback ([type], optional): [description]. Defaults to None.
Raises:
ValueError: raised in case of observed negative intensities.
Returns:
pd.DataFrame: table containing the LFQ intensities of each protein in each sample.
"""
minimum_ratios = settings['quantification']['lfq_ratio_min']
field = settings['quantification']['mode']
if field+'_dn' in df.columns:
field_ = field+'_dn'
else:
field_ = field
if df[field_].min() < 0:
raise ValueError('Negative intensity values present.')
protein_table = protein_profile_parallel(df, minimum_ratios, field_, callback)
return protein_table
# This function invokes a parallel pool and has therfore no dedicated test in the notebook
def protein_profile_parallel_mq(evidence_path : str, protein_groups_path: str, callback=None) -> pd.DataFrame:
"""Derives protein LFQ intensities from Maxquant quantified features.
Args:
evidence_path (str): path to the Maxquant standard output table evidence.txt.
protein_groups_path (str): path to the Maxquant standard output table proteinGroups.txt.
callback ([type], optional): [description]. Defaults to None.
Raises:
FileNotFoundError: if Maxquant files cannot be found.
Returns:
pd.DataFrame: table containing the LFQ intensities of each protein in each sample.
"""
logging.info('Loading files')
for file in [evidence_path, protein_groups_path]:
if not os.path.isfile(file):
raise FileNotFoundError(f'File {file} not found.')
evd = pd.read_csv(evidence_path, sep='\t')
ref = pd.read_csv(protein_groups_path, sep='\t')
experiments = evd['Raw file'].unique().tolist()
logging.info(f'A total of {len(experiments):,} files.')
protein_df = []
max_ = len(ref)
for i in range(max_):
investigate = ref.iloc[i]
evd_ids = [int(_) for _ in investigate['Evidence IDs'].split(';')]
subset = evd.loc[evd_ids].copy()
subset['protein_group'] = investigate['Protein IDs']
subset['filename'] = subset['Raw file']
subset['precursor'] = ['_'.join(_) for _ in zip(subset['Sequence'].values, subset['Charge'].values.astype('str'))]
protein_df.append(subset)
if callback:
callback((i+1)/len(ref))
logging.info(f'A total of {max_:,} proteins.')
df = pd.concat(protein_df)
df, normed = delayed_normalization(df, field ='Intensity')
protein_table = protein_profile_parallel(df, minimum_ratios=1, field='Intensity', callback=callback)
return protein_table
# -
#hide
from nbdev.export import *
notebook2script()
|
nbs/08_quantification.ipynb
|
-- ---
-- jupyter:
-- jupytext:
-- text_representation:
-- extension: .hs
-- format_name: light
-- format_version: '1.5'
-- jupytext_version: 1.14.4
-- kernelspec:
-- display_name: Haskell
-- language: haskell
-- name: haskell
-- ---
-- # 16. Reasoning about programs
-- ## 16.1 Equational reasoning
-- ## 16.2 Reasoning about Haskell
-- ## 16.3 Simple examples
-- ## 16.4 Induction on numbers
-- ## 16.6 Making append vanish
-- ## 16.7 Compiler correctness
-- +
data Expr = Val Int | Add Expr Expr
eval :: Expr -> Int
eval (Val n) = n
eval (Add x y) = eval x + eval y
-- +
type Stack = [Int]
type Code = [Op]
data Op = PUSH Int | ADD
deriving Show
-- -
exec :: Code -> Stack -> Stack
exec [] s = s
exec (PUSH n : c) s = exec c (n : s)
exec (ADD : c) (m : n : s) = exec c (n+m : s)
comp :: Expr -> Code
comp (Val n) = [PUSH n]
comp (Add x y) = comp x ++ comp y ++ [ADD]
-- +
e = Add (Add (Val 2) (Val 3)) (Val 4)
eval e
comp e
exec (comp e) []
-- -
-- ## 16.8 Chapter remarks
-- ## 16.9 Exercises
|
16 Reasoning about programs.ipynb
|