code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Анализ данных на Python
#
# ### Семинар 6. Множества и словари. Задачи.
mySet = set()
myDict = dict()
myList = [1, 1, 1, 2, 3]
set(myList)
s = 'ab' # неизменяемый тип
s[1] = 'c'
myTuple = (1, 2, 3)
myTuple[1] = 3
15
d = {1: 'abc', (2, 3): 1, 'key': [1, 2, 3]}
d[1]
d[(2, 3)]
d['key']
d.values()
d.keys()
d.items()
d = {1: {'a': 1, 'b':2}}
d[1]
d[1]['b']
# +
def test_problem(func, test_data):
for inputs, true_answer in test_data:
answer = func(inputs)
assert answer == true_answer, f'Expected {true_answer}, got {answer}. Input: {inputs}'
print("OK!")
def test_problem_13(func, test_data):
for inputs, true_answer in test_data:
answer = func(*inputs)
assert answer == true_answer, f'Expected {true_answer}, got {answer}. Input: {inputs}'
print("OK!")
# -
# ## Задачка 1: магазин
# Вам предостоит обработать базу данных о продажах некоторого интернет-магазина. База данных представляет собой набор кортежей, в каждом кортеже три элемента: (Покупатель, товар, количество), где Покупатель — имя покупателя (строка без пробелов), товар — название товара (строка без пробелов), количество — количество приобретенных единиц товара.
# Создайте словарь, ключами которого являются имена покупателей, а значениями — словари, ключами которых являются названия товаров, а значениями — количество единиц этого товара, которые купил этот покупатель.
# Напишите функцию aggregate, принимающую некоторое количество набор кортежей из базы данных и возвращающую сводную информацию в виде словаря.
input_ = [("Petrov","pens",5), ("Petrov","pens",6), ("Ivanov","marker",3), ("Ivanov","paper",7),
("Petrov","envelope",20), ("Ivanov","envelope",5)]
# +
from collections import defaultdict
dd = defaultdict(dict)
dd["Petrov"]['pens']= 5
dd
# -
d1 = {}
d1['Petrov']['pens'] = 5
d1
# +
dd = defaultdict(dict)
for name, item, num in input_:
if item in dd[name]:
dd[name][item] += num
else:
dd[name][item] = num
# -
dd
# +
d = {}
for tup in input_:
if tup[0] in d:
if tup[1] in d[tup[0]]:
d[tup[0]][tup[1]] += tup[2]
else:
d[tup[0]][tup[1]] = tup[2]
else:
d[tup[0]] = {}
d[tup[0]][tup[1]] = tup[2]
# -
d
'Ivanov' in d
'Petrov' in d
'Sidorov' in d
def aggregate(names):
d = {}
for tup in names:
if tup[0] in d:
if tup[1] in d[tup[0]]:
d[tup[0]][tup[1]] += tup[2]
else:
d[tup[0]][tup[1]] = tup[2]
else:
d[tup[0]] = {}
d[tup[0]][tup[1]] = tup[2]
return d
AGG_TESTS_DATA = [
([("Petrov","pens",5), ("Ivanov","marker",3), ("Ivanov","paper",7),
("Petrov","envelope",20), ("Ivanov","envelope",5)],
{'Petrov': {'pens': 5, 'envelope': 20},
'Ivanov': {'marker': 3, 'paper': 7, 'envelope': 5}}),
([("Ivanov","aaa",1), ("Petrov","aaa",2), ("Sidorov","aaa",3), ("Ivanov","aaa",6),
("Petrov","aaa",7), ("Sidorov","aaa",8), ("Ivanov","bbb",3), ("Petrov","bbb",7),
("Sidorov","aaa",345), ("Ivanov","ccc",45), ("Petrov","ddd",34),
("Ziborov","eee",234), ("Ivanov","aaa",45)],
{'Ivanov': {'aaa': 52, 'bbb': 3, 'ccc': 45},
'Petrov': {'aaa': 9, 'bbb': 7, 'ddd': 34},
'Sidorov': {'aaa': 356},
'Ziborov': {'eee': 234}})
]
test_problem(aggregate, AGG_TESTS_DATA)
# ## Задачка 2: одинаковые строки
#
# Напишите функцию, которая приенимает на вход две строки и возвращает True, если перестановкой символов можно получить вторую строку из первой.
#
# ```
# s1 = 'aab'
# s2 = 'ab
# ```
s1, s2 = ('foo', 'oof')
s3, s4 = ('aab', 'ab')
s1
s2
# решение 1
set(s1) == set(s2)
# решение 2
sorted(list(s1)) == sorted(list(s2))
sorted(list(s3)) == sorted(list(s4))
sorted(list(s2))
# +
# решение 3
# -
from collections import Counter
Counter(s1) == Counter(s2)
Counter(s1)
Counter(s2)
# +
# решение 4
# -
s1
def myCounter(string):
d = {}
for letter in string:
if letter in d:
d[letter] += 1
else:
d[letter] = 1
return d
myCounter(s1)
myCounter(s2)
myCounter(s1) == myCounter(s2)
def is_isomorphic(a, b):
if Counter(a) == Counter(b):
return True
else:
return False
# +
TEST_DATA = [
(('foo', 'bar'), False),
(('foo', 'oof'), True),
(('a', 'a'), True),
(('aaa', 'aaab'), False),
(('140', '041'), True),
]
test_problem_13(is_isomorphic, TEST_DATA)
# -
# ## Задачка 3. Парковка
#
# Напишите функцию, которая принимает на вход словарь, где ключом является индекс парковочного места, а значением True/False (занято/свободно). Вторым аргументом функция должна принимать индекс места, которое хочется занять.
# Функция должна возращать True, если место можно занять и модифицировать словарь, занимая место. Функция возвращает False, если место уже занято.
def parking_lot(data, index):
if data[index] == False: # место свободно
data[index] = True # заняли место
return True
else: # место занято
return False
# PEP8
def parking_lot(data, index):
if not data[index]: # место свободно
data[index] = True # заняли место
return True
else: # место занято
return False
if True:
print('x')
if False:
print('x')
not False
# +
parking = {0: False, 1: False, 2: False, 3: False}
TEST_DATA = [
((parking, 0), True),
((parking, 0), False),
((parking, 1), True),
((parking, 3), True),
]
test_problem_13(parking_lot, TEST_DATA)
# -
# ## Задачка 4. Последовательность ДНК.
#
# Последовательность ДНК содержит четыре вида символов: 'A', 'C', 'G', и 'T'.
# Напишите функцию, которая будет возращать все подпоследовательности длины 10, которые встретились чаще одного раза.
s = 'AAAAACCCCCAAAAACCCCCCAAAAAGGGTTT'
len(s) - 10 + 1
d = {} # ключи – подпоследовательности длины 10, значения – сколько раз она встретилась
for i in range(len(s) - 10 + 1):
if s[i:i+10] in d:
d[s[i:i+10]] += 1
else:
d[s[i:i+10]] = 1
# решение через цикл
answer = []
for k, v in d.items():
if v > 1:
answer.append(k)
# решение через dictionary comprehension
{k:v for k, v in d.items() if v > 1}
{k:v for k, v in d.items() if v > 1}.keys()
answer
[int(x) for x in input().split() if int(x) % 2 == 0]
d.items()
d
def repeated_dna(s):
d = {} # ключи – подпоследовательности длины 10, значения – сколько раз она встретилась
for i in range(len(s) - 10 + 1):
if s[i:i+10] in d:
d[s[i:i+10]] += 1
else:
d[s[i:i+10]] = 1
filtered_dict = {k:v for k, v in d.items() if v > 1}
return list(filtered_dict.keys())
# +
TEST_DATA = [
('AAAAACCCCCAAAAACCCCCCAAAAAGGGTTT', ['AAAAACCCCC','CCCCCAAAAA']),
('AAAAAAAAAAAAA', ['AAAAAAAAAA']),
]
test_problem(repeated_dna, TEST_DATA)
|
sem06_dict/2011_sem06_set_dict.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''buddhalight'': conda)'
# name: python3
# ---
# # Mastering PyTorch
#
# Edited by <NAME>
#
# This document was edited to understand better for beginner of torch and basic concepts of deep learning base on original document below.
#
# https://pytorch.org/tutorials/beginner/nn_tutorial.html
#
# # WHAT IS TORCH.NN REALLY?
# by <NAME>, fast.ai. Thanks to <NAME> and <NAME>.
#
# We recommend running this tutorial as a notebook, not a script. To download the notebook (.ipynb) file, click the link at the top of the page.
#
# PyTorch provides the elegantly designed modules and classes torch.nn , torch.optim , Dataset , and DataLoader to help you create and train neural networks. In order to fully utilize their power and customize them for your problem, you need to really understand exactly what they’re doing. To develop this understanding, we will first train basic neural net on the MNIST data set without using any features from these models; we will initially only use the most basic PyTorch tensor functionality. Then, we will incrementally add one feature from torch.nn, torch.optim, Dataset, or DataLoader at a time, showing exactly what each piece does, and how it works to make the code either more concise, or more flexible.
#
# **This tutorial assumes you already have PyTorch installed, and are familiar with the basics of tensor operations.** (If you’re familiar with Numpy array operations, you’ll find the PyTorch tensor operations used here nearly identical).
#
# MNIST data setup
# We will use the classic MNIST dataset, which consists of black-and-white images of hand-drawn digits (between 0 and 9).
#
# We will use pathlib for dealing with paths (part of the Python 3 standard library), and will download the dataset using requests. We will only import modules when we use them, so you can see exactly what’s being used at each point.
# # MNIST data setup
#
# We will use the classic MNIST dataset, which consists of black-and-white images of hand-drawn digits (between 0 and 9).
#
# We will use pathlib for dealing with paths (part of the Python 3 standard library), and will download the dataset using requests. We will only import modules when we use them, so you can see exactly what’s being used at each point.
# +
from pathlib import Path
import requests
DATA_PATH = Path("data")
PATH = DATA_PATH / "mnist"
PATH.mkdir(parents=True, exist_ok=True)
URL = "https://github.com/pytorch/tutorials/raw/master/_static/"
FILENAME = "mnist.pkl.gz"
if not (PATH / FILENAME).exists():
content = requests.get(URL + FILENAME).content
(PATH / FILENAME).open("wb").write(content)
# -
# This dataset is in numpy array format, and has been stored using pickle, a python-specific format for serializing data.
# +
import pickle
import gzip
with gzip.open((PATH / FILENAME).as_posix(), 'rb') as f:
((x_train, y_train), (x_valid, y_valid), _) = pickle.load(f, encoding="latin-1")
# -
# Each image is 28 x 28, and is being stored as a flattened row of length 784 (=28x28). Let’s take a look at one; we need to reshape it to 2d first.
# +
from matplotlib import pyplot
import numpy as np
pyplot.imshow(x_train[0].reshape((28, 28)), cmap="gray")
print(x_train.shape)
# -
# PyTorch uses torch.tensor, rather than numpy arrays, so we need to convert our data.
# ### Take a Moment!
#
# ### Python map() function
#
# ```map()``` function returns a map object(which is an iterator) of the results after applying the given function to each item of a given iterable (list, tuple etc.)
#
# **Syntax** :
#
# > ```map(fun, iter)```
#
# **Parameters** :
#
# > fun : It is a function to which map passes each element of given iterable.
#
# > iter : It is a iterable which is to be mapped.
#
# **NOTE** : You can pass one or more iterable to the map() function.
#
# **Returns** :
#
# > Returns a list of the results after applying the given function
# to each item of a given iterable (list, tuple etc.)
#
# **NOTE** : The returned value from map() (map object) then can be passed to functions like list() (to create a list), set() (to create a set) .
#
# ### Take a Moment!
# ### ```torch.tensor()``` function
#
# > ```torch.tensor(data, *, dtype=None, device=None, requires_grad=False, pin_memory=False)``` → Tensor
#
# Constructs a tensor with data.
#
# **Parameters**
# > **data (```array_like```)** – Initial data for the tensor. Can be a list, tuple, NumPy ndarray, scalar, and other types.
#
# **Keyword Arguments**
# - **dtype (```torch.dtype```, optional)** – the desired data type of returned tensor. Default: ***if None, infers data type from data.***
#
# - **device** (```torch.device```, optional) – the desired device of returned tensor. Default: if None, uses the current device for the default tensor type (see torch.set_default_tensor_type()). device will be the CPU for CPU tensor types and the current CUDA device for CUDA tensor types.
#
# - **requires_grad** (bool, optional) – If autograd should record operations on the returned tensor. Default: False.
#
# - **pin_memory** (bool, optional) – If set, returned tensor would be allocated in the pinned memory. Works only for CPU tensors. Default: False.
#
# ---
#
# ### TORCH.TENSOR datatype
# A ```torch.Tensor``` is a multi-dimensional matrix containing elements of a single **data type**.
# +
import torch
x_train, y_train, x_valid, y_valid = map(
torch.tensor, (x_train, y_train, x_valid, y_valid)
)
n, c = x_train.shape
print(x_train, y_train)
print(x_train.shape)
print(x_train.min(), y_train.max())
# -
# # Neural net from scratch (no torch.nn)
# Let’s first create a model using nothing but PyTorch tensor operations. We’re assuming you’re already familiar with the basics of neural networks. (If you’re not, you can learn them at course.fast.ai).
#
# PyTorch provides methods to create random or zero-filled tensors, which we will use to create our weights and bias for a simple linear model. These are just regular tensors, with one very special addition: we tell PyTorch that they require a gradient. This causes PyTorch to record all of the operations done on the tensor, so that it can calculate the gradient during back-propagation automatically!
#
# For the weights, we set requires_grad **after** the initialization, since we don’t want that step included in the gradient. (Note that a trailing _ in PyTorch signifies that the operation is performed in-place.)
#
# **NOTE**
#
# We are initializing the weights here with Xavier initialisation (by multiplying with 1/sqrt(n)).
#
# ### Take a Moment!
# ### TORCH.TENSOR.REQUIRES_GRAD_
#
# ```Tensor.requires_grad_```(requires_grad=True) → Tensor
#
# Change if autograd should record operations on this tensor: sets this tensor’s requires_grad attribute in-place. Returns this tensor.
#
# requires_grad_()’s main use case is to tell autograd to begin recording operations on a Tensor tensor. If tensor has requires_grad=False (because it was obtained through a DataLoader, or required preprocessing or initialization), tensor.requires_grad_() makes it so that autograd will begin to record operations on tensor.
#
# Parameters
# **requires_grad** (bool) – If autograd should record operations on this tensor. Default: True.
#
# Example:
#
# ```
# # Let's say we want to preprocess some saved weights and use
#
# # the result as new weights.
#
# saved_weights = [0.1, 0.2, 0.3, 0.25]
# loaded_weights = torch.tensor(saved_weights)
#
# weights = preprocess(loaded_weights) # some function
# weights
# tensor([-0.5503, 0.4926, -2.1158, -0.8303])
# ```
#
# ```
# # Now, start to record operations done to weights
#
# weights.requires_grad_()
# out = weights.pow(2).sum()
# out.backward()
# weights.grad
# tensor([-1.1007, 0.9853, -4.2316, -1.6606])
# ```
# +
import math
weights = torch.randn(784, 10) / math.sqrt(784)
weights.requires_grad_()
bias = torch.zeros(10, requires_grad=True)
# -
# Thanks to PyTorch’s ability to calculate gradients automatically, we can use any standard Python function (or callable object) as a model! So let’s just write a plain matrix multiplication and broadcasted addition to create a simple linear model. We also need an activation function, so we’ll write log_softmax and use it. Remember: although PyTorch provides lots of pre-written loss functions, activation functions, and so forth, you can easily write your own using plain python. PyTorch will even create fast GPU or vectorized CPU code for your function automatically.
# +
def log_softmax(x):
return x - x.exp().sum(-1).log().unsqueeze(-1)
def model(xb):
return log_softmax(xb @ weights + bias)
# -
# In the above, the @ stands for the dot product operation. We will call our function on one batch of data (in this case, 64 images). This is one forward pass. Note that our predictions won’t be any better than random at this stage, since we start with random weights.
# +
bs = 64
xb = x_train[0:bs]
preds = model(xb)
preds[0], preds.shape
print(preds[0], preds.shape)
# -
# As you see, the preds tensor contains not only the tensor values, but also a gradient function. We’ll use this later to do backprop.
#
# Let’s implement negative log-likelihood to use as the loss function (again, we can just use standard Python):
# ### Take a Moment!
#
# ```preds[range(target.shape[0]), target]```
#
# This is an indices itself to index of an specific matrix or a vector.
#
# so, if ```np.array([range(target.shape[0]), target])``` or just as a list like ```[range(target.shape[0]), target]``` will only show the itself, as
#
# ```
# [range(0, 64), tensor([5, 0, 4, 1, 9, 2, 1, 3, 1, 4, 3, 5, 3, 6, 1, 7, 2, 8, 6, 9, 4, 0, 9, 1,
# 1, 2, 4, 3, 2, 7, 3, 8, 6, 9, 0, 5, 6, 0, 7, 6, 1, 8, 7, 9, 3, 9, 8, 5,
# 9, 3, 3, 0, 7, 4, 9, 8, 0, 9, 4, 1, 4, 4, 6, 0])]```
#
#
# +
def nll(input, target):
return -input[range(target.shape[0]), target].mean()
loss_func = nll
# -
# Let’s check our loss with our random model, so we can see if we improve after a backprop pass later.
yb = y_train[0:bs]
print(loss_func(preds, yb))
# Let’s also implement a function to calculate the accuracy of our model. For each prediction, if the index with the largest value matches the target value, then the prediction was correct.
def accuracy(out, yb):
preds = torch.argmax(out, dim=1)
return (preds == yb).float().mean()
# Let’s check the accuracy of our random model, so we can see if our accuracy improves as our loss improves.
#
#
print(accuracy(preds, yb))
# We can now run a training loop. For each iteration, we will:
#
# select a mini-batch of data (of size bs)
# use the model to make predictions
# calculate the loss
# loss.backward() updates the gradients of the model, in this case, weights and bias.
# We now use these gradients to update the weights and bias. We do this within the torch.no_grad() context manager, **because we do not want these actions to be recorded for our next calculation of the gradient.** You can read more about how PyTorch’s Autograd records operations here. https://pytorch.org/docs/stable/notes/autograd.html
#
# We then set the gradients to zero, so that we are ready for the next loop. Otherwise, our gradients would record a running tally of all the operations that had happened (i.e. loss.backward() adds the gradients to whatever is already stored, rather than replacing them).
#
# **TIP**
#
# You can use the standard python debugger to step through PyTorch code, allowing you to check the various variable values at each step. Uncomment set_trace() below to try it out.
# +
from IPython.core.debugger import set_trace
lr=0.5
epochs = 2
for epoch in range(epochs):
for i in range((n - 1) // bs + 1):
# set_trace()
start_i = i *bs
end_i = start_i + bs
xb = x_train[start_i:end_i]
yb = y_train[start_i:end_i]
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward() ## take backward() since loss return the torch.Tensor
with torch.no_grad():
weights -= weights.grad * lr
bias -= bias.grad * lr
weights.grad.zero_()
bias.grad.zero_()
# -
# ### Take a Moment!
#
# ### NO_GRAD
# CLASS ```torch.no_grad```
# Context-manager that disabled gradient calculation.
#
# Disabling gradient calculation is useful for inference, when you are sure that you will not call Tensor.backward(). It will reduce memory consumption for computations that would otherwise have requires_grad=True.
#
# In this mode, the result of every computation will have requires_grad=False, even when the inputs have requires_grad=True.
#
# This context manager is thread local; it will not affect computation in other threads.
#
# Also functions as a decorator. (Make sure to instantiate with parenthesis.)
#
#
# ### TORCH.TENSOR.BACKWARD
#
# ```Tensor.backward(gradient=None, retain_graph=None, create_graph=False, inputs=None)```
# Computes the gradient of current tensor w.r.t. graph leaves.
#
# The graph is differentiated using the chain rule. If the tensor is non-scalar (i.e. its data has more than one element) and requires gradient, the function additionally requires specifying gradient. It should be a tensor of matching type and location, that contains the gradient of the differentiated function w.r.t. self.
#
# This function accumulates gradients in the leaves - you might need to zero .grad attributes or set them to None before calling it. See Default gradient layouts for details on the memory layout of accumulated gradients.
#
# **NOTE**
#
# If you run any forward ops, create gradient, and/or call backward in a user-specified CUDA stream context, see Stream semantics of backward passes.
#
# #### Parameters
# - **gradient** (Tensor or None) – Gradient w.r.t. the tensor. If it is a tensor, it will be automatically converted to a Tensor that does not require grad unless create_graph is True. None values can be specified for scalar Tensors or ones that don’t require grad. If a None value would be acceptable then this argument is optional.
#
# - **retain_graph** (bool, optional) – If False, the graph used to compute the grads will be freed. Note that in nearly all cases setting this option to True is not needed and often can be worked around in a much more efficient way. Defaults to the value of create_graph.
#
# - **create_graph** (bool, optional) – If True, graph of the derivative will be constructed, allowing to compute higher order derivative products. Defaults to False.
#
# - **inputs** (sequence of Tensor) – Inputs w.r.t. which the gradient will be accumulated into .grad. All other Tensors will be ignored. If not provided, the gradient is accumulated into all the leaf Tensors that were used to compute the attr::tensors. All the provided inputs must be leaf Tensors.
# That’s it: we’ve created and trained a minimal neural network (in this case, a logistic regression, since we have no hidden layers) entirely from scratch!
#
# Let’s check the loss and accuracy and compare those to what we got earlier. We expect that the loss will have decreased and accuracy to have increased, and they have.
# ### Take a Moment!
#
# Why this is logit regression?
#
# --- add
#
#
print(loss_func(model(xb), yb), accuracy(model(xb), yb))
# ## Using torch.nn.functional
# We will now refactor our code, so that it does the same thing as before, only we’ll start taking advantage of PyTorch’s nn classes to make it more concise and flexible. At each step from here, we should be making our code one or more of: shorter, more understandable, and/or more flexible.
#
# The first and easiest step is to make our code shorter by replacing our hand-written activation and loss functions with those from torch.nn.functional (which is generally imported into the namespace F by convention). This module contains all the functions in the torch.nn library (whereas other parts of the library contain classes). As well as a wide range of loss and activation functions, you’ll also find here some convenient functions for creating neural nets, such as pooling functions. (There are also functions for doing convolutions, linear layers, etc, but as we’ll see, these are usually better handled using other parts of the library.)
#
# If you’re using negative log likelihood loss and log softmax activation, then Pytorch provides **a single function F.cross_entropy that combines the two. So we can even remove the activation function from our model.**
# +
import torch.nn.functional as F
loss_func = F.cross_entropy
def model(xb):
return xb @ weights + bias
# -
# Note that we no longer call ```log_softmax``` in the ```model``` function. Let's confirm that our loss and accuracy are the same as before:
# ### Take a Moment!
#
# Before, we defined the **model** that the form of dot producted result in the log_softmax,
# and give it to the loss function that is the negative likelihood with the yb.
#
# However the ```cross_entropy``` above contains the log_softmax in it, as we can just define the model as ```xb @ weights + bias```.
print(loss_func(model(xb), yb), accuracy(model(xb), yb))
# ## Refactor using nn.Module
#
# Next up, we’ll use nn.Module and nn.Parameter, for a clearer and more concise training loop. We subclass nn.Module (which itself is a class and able to keep track of state). In this case, we want to create a class that holds our weights, bias, and method for the forward step. nn.Module has a number of attributes and methods (such as .parameters() and .zero_grad()) which we will be using.
#
# **NOTE**
#
# nn.Module (uppercase M) is a PyTorch specific concept, and is a class we’ll be using a lot. nn.Module is not to be confused with the Python concept of a (lowercase m) module, which is a file of Python code that can be imported.
#
# +
from torch import nn
class Mnist_Logistic(nn.Module):
def __init__(self):
super().__init__()
self.weights = nn.Parameter(torch.randn(784, 10) / math.sqrt(784))
self.bias = nn.Parameter(torch.zeros(10))
def forward(self, xb):
return xb @ self.weights + self.bias
# -
# Since we're now using an object instead of just using a function, we first have to instantiate our model:
model = Mnist_Logistic()
# Now we can calculate the loss in the same way as before. Note that ```nn.Module``` objects are used as if they are functions (i.e they are ***callable***), but behind the scenes Pytorch will call our ```forward``` method automatically.
print(loss_func(model(xb), yb))
# Previously for our training loop we had to update the values for each parameter by name, and manually zero out the grads for each parameter separately, like this:
with torch.no_grad():
weights -= weights.grad * lr
bias -= bias.grad * lr
weights.grad.zero_()
bias.grad.zero_()
# Now we can take advantage of model.parameters() and model.zero_grad() (which are both defined by PyTorch for nn.Module) to make those steps more concise and less prone to the error of forgetting some of our parameters, particularly if we had a more complicated model:
# +
# with torch.no_grad():
# for p in model.parameters(): p -= p.grad * lr
# model.zero_grad()
# -
# We’ll wrap our little training loop in a fit function so we can run it again later.
# +
def fit():
for epoch in range(epochs):
for i in range((n - 1) // bs + 1):
start_i = i * bs
end_i = start_i + bs
xb = x_train[start_i: end_i]
yb = y_train[start_i: end_i]
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
with torch.no_grad():
for p in model.parameters():
p -= p.grad * lr
model.zero_grad()
fit()
# -
# Let’s double-check that our loss has gone down:
#
#
print(loss_func(model(xb), yb))
# ## Refactor using nn.Linear
#
# We continue to refactor our code. Instead of manually defining and initializing self.weights and self.bias, and calculating xb @ self.weights + self.bias, we will instead use the Pytorch class nn.Linear for a linear layer, which does all that for us. Pytorch has many types of predefined layers that can greatly simplify our code, and often makes it faster too.
class Mnist_Logistic(nn.Module):
def __init__(self):
super().__init__()
self.lin = nn.Linear(784, 10)
def forward(self, xb):
return self.lin(xb)
# We instantiate our model and calculate the loss in the same way as before:
#
#
model = Mnist_Logistic()
print(loss_func(model(xb), yb))
# We are still able to use our same fit method as before.
fit()
print(loss_func(model(xb), yb))
# ## Refactor using optim
#
# Pytorch also has a package with various optimization algorithms, torch.optim. We can use the step method from our optimizer to take a forward step, instead of manually updating each parameter.
#
# This will let us replace our previous manually coded optimization step:
#
# ```
# with torch.no_grad():
# for p in model.parameters(): p -= p.grad * lr
# model.zero_grad()
# ```
#
# and instead use just:
#
# ```
#
# opt.step()
# opt.zero_grad()
#
# ```
#
# (```optim.zero_grad()``` resets the gradient to 0 and we need to call it before computing the gradient for the next minbatch.)
from torch import optim
# We'll define a little function to create our model and optimizer so we can reuse it in the future.
# +
def get_model():
model = Mnist_Logistic()
return model, optim.SGD(model.parameters(), lr=lr)
model, opt = get_model()
x_train_set = x_train[0: bs]
y_train_set = y_train[0: bs]
print(loss_func(model(x_train_set), y_train_set))
# print(loss_func(model(xb), yb)) => on tutorial => but the shape of xb is (16, 784) since the xb resigned by iteration of epochs
# print(50000 % 64) === 16
for epoch in range(epochs):
for i in range((n - 1) // bs + 1):
start_i = i * bs
end_i = start_i + bs
xb = x_train[start_i:end_i]
yb = y_train[start_i:end_i]
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
opt.step()
opt.zero_grad()
print(loss_func(model(x_train_set), y_train_set))
# output will return similar loss every playback since we load the model all the time on the top.
# -
# ## Refactor using Dataset
#
# PyTorch has an abstract Dataset ```class```. A Dataset can be anything that has a ```__len__``` function (called by Python’s standard ```len``` function) and a ```__getitem__``` function as a way of indexing into it. This tutorial walks through a nice example of creating a custom ```FacialLandmarkDataset``` class as a subclass of ```Dataset```.
#
# PyTorch’s TensorDataset is a Dataset wrapping tensors. By defining a length and way of indexing, this also gives us a way to iterate, index, and slice along the first dimension of a tensor. This will make it easier to access both the independent and dependent variables in the same line as we train.
from torch.utils.data import TensorDataset
# Both x_train and y_train can be combined in a single TensorDataset, which will be easier to iterate over and slice.
#
train_ds = TensorDataset(x_train, y_train)
# Previously, we had to iterate through minibatches of x and y values separately:
#
# ```
# xb = x_train[start_i:end_i]
# yb = y_train[start_i:end_i]
# ```
#
# Now, we can do these two steps together:
#
# ```xb, yb = train_ds[i*bs + i*bs+bs]```
# +
model, opt = get_model()
for epoch in range(epochs):
for i in range((n - 1) // bs + 1):
xb, yb = train_ds[i * bs: i * bs + bs]
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
opt.step()
opt.zero_grad()
print(loss_func(model(xb), yb))
# -
# ## Refactor using DataLoader
#
# Pytorch’s ```DataLoader``` is responsible for managing batches. You can create a ```DataLoader``` from any ```Dataset```. DataLoader makes it easier to iterate over batches. Rather than having to use ```train_ds[i*bs : i*bs+bs]```, the DataLoader gives us each minibatch automatically.
#
#
#
#
# +
from torch.utils.data import DataLoader
train_ds = TensorDataset(x_train, y_train)
train_dl = DataLoader(train_ds, batch_size = bs)
# -
# Previously, our loop iterated over batches (xb, yb) like this:
#
# ```
# for i in range((n-1)//bs + 1):
# xb,yb = train_ds[i*bs : i*bs+bs]
# pred = model(xb)
# ```
# Now, our loop is much cleaner, as (xb, yb) are loaded automatically from the data loader:
# ```
# for xb,yb in train_dl:
# pred = model(xb)
# ```
# ### Take a Moment!
#
# **Refactor Process**
#
# ```
# xb = x_train[start_i:end_i]
# yb = y_train[start_i:end_i]
# ```
#
# After using ```torch.utils.data.TensorDataset```
#
# ```
# xb, yb = train_ds[i*bs + i*bs+bs]
# ```
#
# after using ```torch.utils.data.DataLoader```
#
# ```
# for xb,yb in train_dl:
# pred = model(xb)
# ```
#
#
# +
model, opt = get_model()
for epoch in range(epochs):
for xb, yb in train_dl:
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
opt.step()
opt.zero_grad()
print(loss_func(model(xb), yb))
# -
# Thanks to Pytorch’s nn.Module, nn.Parameter, Dataset, and DataLoader, our training loop is now dramatically smaller and easier to understand. Let’s now try to add the basic features necessary to create effective models in practice.
# ## Add Validation
#
# In section 1, we were just trying to get a reasonable training loop set up for use on our training data. In reality, you always should also have a validation set, in order to identify if you are overfitting.
#
# Shuffling the training data is important to prevent correlation between batches and overfitting. On the other hand, the validation loss will be identical whether we shuffle the validation set or not. Since shuffling takes extra time, it makes no sense to shuffle the validation data.
#
# We’ll use a batch size for the validation set that is twice as large as that for the training set. This is because the validation set does not need backpropagation and thus takes less memory (it doesn’t need to store the gradients). We take advantage of this to use a larger batch size and compute the loss more quickly.
# +
train_ds = TensorDataset(x_train, y_train)
train_dl = DataLoader(train_ds, batch_size = bs, shuffle= True)
valid_ds = TensorDataset(x_valid, y_valid)
valid_dl = DataLoader(valid_ds, batch_size= bs* 2)
# -
# We will calculate and print the validation loss at the end of each epoch.
#
# (Note that we always call model.train() before training, and model.eval() before inference, because these are used by layers such as nn.BatchNorm2d and nn.Dropout to ensure appropriate behaviour for these different phases.)
# +
model, opt = get_model()
for epoch in range(epochs):
model.train()
for xb, yb in train_dl:
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
opt.step()
opt.zero_grad()
model.eval()
with torch.no_grad():
valid_loss = sum(loss_func(model(xb), yb) for xb, yb in valid_dl)
print(epoch, valid_loss / len(valid_dl))
# -
# ## Create fit() and get_data()
#
# We’ll now do a little refactoring of our own. Since we go through a similar process twice of calculating the loss for both the training set and the validation set, let’s make that into its own function, loss_batch, which computes the loss for one batch.
#
# We pass an optimizer in for the training set, and use it to perform backprop. For the validation set, we don’t pass an optimizer, so the method doesn’t perform backprop.
def loss_batch(model, loss_func, xb, yb, opt=None):
loss = loss_func(model(xb), yb)
if opt is not None:
loss.backward()
opt.step()
opt.zero_grad()
return loss.item(), len(xb)
# ```fit``` runs the necessary operations to train our model and compute the training and validation losses for each epoch.
#
# ### Take a Moment
#
# ```loss.item()```
#
# since loss is a tuple of tensor(loss_value, grad_fn), write ```loss.item()``` only returns the value of loss.
#
# i.e)
# loss : tensor(0.3109, grad_fn=<NllLossBackward>) loss.item() : 0.31085121631622314
#
# ### Take a Moment!
#
# **List generator comprehension**
#
# if just write ```func for in``` without any wraps like ```loss_batch(model, loss_func, xb, yb) for xb, yb in valid_dl```
#
# the error occurs
# ```
# File "<ipython-input-332-9c7cab139303>", line 12
# print('just', loss_batch(model, loss_func, xb, yb) for xb, yb in valid_dl)
# ^
# SyntaxError: Generator expression must be parenthesized
# ```
# **StackOverflow**
#
# "It is in the form of a list or generator comprehension, but without the wrapping [] for the list or () for the generator." https://stackoverflow.com/questions/61945870/generator-expression-must-be-parenthesized-on-a-constraint
#
# ---
#
# ** *[] Method**
#
#
# Using * in front of a list expands out the members as individual arguments. So, the following two function calls will be equivalent:
# ```
# my_function(*[1, 2, 3])
# my_function(1, 2, 3)
# ```
# Obviously, the first one is not very useful if you already know the precise number of arguments. It becomes more useful with a comprehension like you are using, where is is not clear how many items will be in the list.
# https://stackoverflow.com/questions/45825344/understanding-in-python-passed-to-agg-in-pyspark
#
# ---
# **At Below**
#
# ```
# print('without: ', (loss_batch(model, loss_func, xb, yb) for xb, yb in valid_dl))
#
# print()
#
# print('with: ', *[loss_batch(model, loss_func, xb, yb) for xb, yb in valid_dl])
# ```
# with: (0.36518675088882446, 128) (0.4869171977043152, 128) ... lots of items... (0.0817655399441719, 16)
#
# without: <generator object fit.<locals>.<genexpr> at 0x000002CCC6FC9430>
#
# +
import numpy as np
def fit(epochs, model, loss_func, opt, train_dl, valid_dl):
for epoch in range(epochs):
model.train()
for xb, yb in train_dl:
loss_batch(model, loss_func, xb, yb, opt)
model.eval()
with torch.no_grad():
losses, nums = zip(
*[loss_batch(model, loss_func, xb, yb) for xb, yb in valid_dl]
)
val_loss = np.sum(np.multiply(losses, nums)) / np.sum(nums)
print(epoch, val_loss)
# -
# ```get_data``` returns dataloaders for the training and validation seets.
def get_data(train_ds, valid_ds, bs):
return (
DataLoader(train_ds, batch_size = bs, shuffle=True),
DataLoader(valid_ds, batch_size = bs * 2)
)
# Now, out whole process of obtaining the data loaders and fitting the model can be run in 3 lines of code:
# + tags=[]
train_dl, valid_dl = get_data(train_ds, valid_ds, bs)
model, opt = get_model()
fit(epochs, model, loss_func, opt, train_dl, valid_dl)
# -
# You can use these basic 3 lines of code to train a wide variety of models. Let’s see if we can use them to train a convolutional neural network (CNN)!
# ## Switch to CNN
#
# We are now going to build our neural network with three convolutional layers. Because none of the functions in the previous section assume anything about the model form, we’ll be able to use them to train a CNN without any modification.
#
# We will use Pytorch’s predefined ```Conv2d``` class as our convolutional layer. We define a CNN with 3 convolutional layers. Each convolution is followed by a ReLU. At the end, we perform an average pooling. (Note that ```view``` is PyTorch’s version of numpy’s ```reshape```)
# +
class Mnist_CNN(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(1, 16, kernel_size = 3, stride = 2, padding = 1)
self.conv2 = nn.Conv2d(16, 16, kernel_size = 3, stride = 2, padding =1)
self.conv3 = nn.Conv2d(16, 10, kernel_size = 3, stride = 2, padding =1)
def forward(self, xb):
xb = xb.view(-1, 1 ,28, 28)
xb = F.relu(self.conv1(xb))
xb = F.relu(self.conv2(xb))
xb = F.relu(self.conv3(xb))
xb = F.avg_pool2d(xb, 4)
return xb.view(-1, xb.size(1))
lr = 0.1
# -
# Momentum is a variation on stochastic gradient descent that takes previous updates into account as well and generally leads to faster training.
# + tags=[]
model = Mnist_CNN()
opt = optim.SGD(model.parameters(), lr= lr, momentum=0.9)
fit(epochs, model, loss_func, opt, train_dl, valid_dl)
# -
# ## nn.Sequential
#
# ```torch.nn``` has another handy class we can use to simplify our code: Sequential . A ```Sequential``` object runs each of the modules contained within it, in a sequential manner. This is a simpler way of writing our neural network.
#
# To take advantage of this, we need to be able to easily define a custom layer from a given function. For instance, PyTorch doesn’t have a view layer, and we need to create one for our network. ```Lambda will``` create a layer that we can then use when defining a network with ```Sequential```.
# ### Take a Moment!
#
# don't misunderstand!
#
# the subclass that supered the ```nn.Module``` automatically excute ```self.forward()``` when called.
#
# therefore, we can customize as a subclass of ```nn.Module``` as a various excutable function.
# +
class Lambda(nn.Module):
def __init__(self, func):
super().__init__()
self.func = func
def forward(self, x):
return self.func(x)
def preprocess(x):
return x.view(-1, 1, 28, 28)
# -
# The model created with ```Sequential``` is simply:
# +
model = nn.Sequential(
Lambda(preprocess),
nn.Conv2d(1, 16, kernel_size = 3, stride=2, padding = 1),
nn.ReLU(),
nn.Conv2d(16, 16, kernel_size=3, stride = 2, padding=1),
nn.ReLU(),
nn.Conv2d(16, 10, kernel_size = 3, stride=2, padding=1),
nn.ReLU(),
nn.AvgPool2d(4),
Lambda(lambda x: x.view(x.size(0), -1)),
)
opt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)
fit(epochs, model, loss_func, opt, train_dl, valid_dl)
# -
# ## Wrapping DataLoader
#
# Our CNN is fairly concise, but it only works with MNIST, because:
# It assumes the input is a 28*28 long vector
# It assumes that the final CNN grid size is 4*4 (since that’s the average
# pooling kernel size we used)
#
# Let’s get rid of these two assumptions, so our model works with any 2d single channel image. First, we can remove the initial Lambda layer by moving the data preprocessing into a generator:
# ### Take a Moment!
#
# ```def __len__(self)``` and ```def __iter__(self)``` overwrite the basic method of python class.
#
# it seems to make able to work len function and iterable for working mini-batching of dl (data loader)
#
# ---
#
# ### Python Iterators
#
# An iterator is an object that contains a countable number of values.
#
# An iterator is an object that can be iterated upon, meaning that you can traverse through all the values.
#
# Technically, in Python, an iterator is an object which implements the iterator protocol, which consist of the methods __iter__() and __next__().
#
# **Iterator vs Iterable**
#
# Lists, tuples, dictionaries, and sets are all iterable objects. They are iterable containers which you can get an iterator from.
#
# All these objects have a iter() method which is used to get an iterator: Tuple, List, String etc...
#
# ---
# ### Python iter() Function
#
# Create an iterator object, and print the items:
# +
def preprocess(x, y):
return x.view(-1, 1, 28, 28), y
class WrappedDataLoader:
def __init__(self, dl, func):
self.dl = dl
self.func = func
def __len__(self):
return len(self.dl)
def __iter__(self):
batches = iter(self.dl)
for b in batches:
yield (self.func(*b))
train_dl, valid_dl = get_data(train_ds, valid_ds, bs)
train_dl = WrappedDataLoader(train_dl, preprocess)
valid_dl = WrappedDataLoader(valid_dl, preprocess)
# -
# Next, we can replace nn.AvgPool2d with nn.AdaptiveAvgPool2d, which allows us to define the size of the output tensor we want, rather than the input tensor we have. As a result, our model will work with any size input.
# +
model = nn.Sequential(
nn.Conv2d(1, 16, kernel_size = 3, stride = 2, padding = 1),
nn.ReLU(),
nn.Conv2d(16, 16, kernel_size = 3, stride = 2, padding = 1),
nn.ReLU(),
nn.Conv2d(16, 10, kernel_size = 3, stride = 2, padding = 1),
nn.ReLU(),
nn.AdaptiveAvgPool2d(1),
Lambda(lambda x: x.view(x.size(0), -1)),
)
opt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)
# -
# Let's try it out:
fit(epochs, model, loss_func, opt, train_dl, valid_dl)
# ## Using your GPU
#
# If you’re lucky enough to have access to a CUDA-capable GPU (you can rent one for about $0.50/hour from most cloud providers) you can use it to speed up your code. First check that your GPU is working in Pytorch:
print(torch.cuda.is_available())
# And then create a device object for it:
dev = torch.device("cuda") if torch.cuda.is_available() else torch.device("cpu")
# Let's update ```preprocess``` to move batches to the GPU:
# +
def preprocess(x, y):
return x.view(-1, 1, 28, 28).to(dev), y.to(dev)
train_dl, valid_dl = get_data(train_ds, valid_ds, bs)
train_dl = WrappedDataLoader(train_dl, preprocess)
valid_dl = WrappedDataLoader(valid_dl, preprocess)
# -
# Finally, we can move our model to the GPU.
model.to(dev)
opt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)
# You should find it runs faster now:
fit(epochs, model, loss_func, opt, train_dl, valid_dl)
# # Closing thoughts
# We now have a general data pipeline and training loop which you can use for training many types of models using Pytorch. To see how simple training a model can now be, take a look at the ***mnist_sample*** sample notebook.
#
# Of course, there are many things you’ll want to add, such as data augmentation, hyperparameter tuning, monitoring training, transfer learning, and so forth. These features are available in the fastai library, which has been developed using the same design approach shown in this tutorial, providing a natural next step for practitioners looking to take their models further.
#
# We promised at the start of this tutorial we’d explain through example each of ```torch.nn```, ```torch.optim```, ```Dataset```, and ```DataLoader```. So let’s summarize what we’ve seen:
#
# **torch.nn**
#
# - ```Module```: creates a callable which behaves like a function, but can also contain state(such as neural net layer weights). It knows what Parameter (s) it contains and can zero all their gradients, loop through them for weight updates, etc.
# - ```Parameter```: a wrapper for a tensor that tells a Module that it has weights that need updating during backprop. Only tensors with the requires_grad attribute set are updated
# - ```functional```: a module(usually imported into the F namespace by convention) which contains activation functions, loss functions, etc, as well as non-stateful versions of layers such as convolutional and linear layers.
#
# **```torch.optim```**: Contains optimizers such as SGD, which update the weights of Parameter during the backward step
#
# **```Dataset```**: An abstract interface of objects with a __len__ and a __getitem__, including classes provided with Pytorch such as TensorDataset
#
# **```DataLoader```**: Takes any Dataset and creates an iterator which returns batches of data.
|
3. Learning-Pytorch/1.3. what_is_torch.nn_really.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="l661l2MGzKN_" colab_type="code" colab={}
# !mkdir -p "/content/drive/My Drive/Colab Notebooks/matrix/matrix_2"
# + id="gE5yBGCZzref" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 132} outputId="16f69a07-cdcd-4fd2-e3d9-112b49c6607e" executionInfo={"status": "error", "timestamp": 1583403084321, "user_tz": -60, "elapsed": 4106, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}}
# cd "/content/drive/My Drive/Colab Notebooks/matrix_2"
#c19cbf7a1e67c3a08a1ef21cd3a66c0841ae18a8
# + id="yngbQNU3zvaB" colab_type="code" colab={}
# ls
# + id="K1z-INjM0Z11" colab_type="code" outputId="3d04f3da-548f-4ee2-efef-31c2c8ecd49b" executionInfo={"status": "ok", "timestamp": 1583403086752, "user_tz": -60, "elapsed": 1382, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd "drive/My Drive/Colab Notebooks/matrix/matrix_2/"
# + id="N3uM4xtd3Cm7" colab_type="code" outputId="d1cd50a5-790e-4313-a4f0-2bb6794c2b50" executionInfo={"status": "ok", "timestamp": 1583403093641, "user_tz": -60, "elapsed": 6614, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# !pwd
# + id="Hqmqdqkx0pE6" colab_type="code" colab={}
#{}@
GITHUB_TOKEN = ""
GITHUB_URL = "https://{}@github.com/eb-art/cars_forecasting.git".format(GITHUB_TOKEN)
# + id="tsQJ-2va0-M0" colab_type="code" outputId="9a0233a9-9bab-4aee-878b-577343416b0a" executionInfo={"status": "ok", "timestamp": 1583403145864, "user_tz": -60, "elapsed": 5996, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 54}
# !git clone $GITHUB_URL
# + id="qjlvj7-O9cjI" colab_type="code" outputId="24eac396-4b19-4b22-c185-75dce17ba3b3" executionInfo={"status": "ok", "timestamp": 1583403156110, "user_tz": -60, "elapsed": 3230, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# ls
# + id="1x9XE7lT9et1" colab_type="code" outputId="53b08578-1784-4ece-89d0-9501e7df6b58" executionInfo={"status": "ok", "timestamp": 1583403157327, "user_tz": -60, "elapsed": 383, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 54}
# cd cars_forecasting/
# + id="MGDJ8a1r1Gye" colab_type="code" colab={}
# !mkdir data
# + id="8oDN7FqQ15Se" colab_type="code" colab={}
# !echo -e "*\n!.gitignore" > data/.gitignore
#zignoruj wszystko (*) w katalogu oprócz (!)
# + id="mybcJnjX4yW0" colab_type="code" colab={}
# #!git init
# + id="M7E43qiv2Vi8" colab_type="code" colab={}
# !git add data/.gitignore
# + id="hAxt6xa_2ZA3" colab_type="code" outputId="40daa649-f9eb-40d6-f024-96f16fe0725c" executionInfo={"status": "ok", "timestamp": 1583131072940, "user_tz": -60, "elapsed": 2102, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 245}
# !git commit -m "add data directory"
# + id="-PmK6wfB4EOS" colab_type="code" colab={}
# !git config --global user.email "<EMAIL>"
# !git config --global user.name "<NAME>"
# + id="_B9cIncbLP78" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 156} outputId="748eeb4e-e2eb-45d5-8af1-44445d18b93b" executionInfo={"status": "ok", "timestamp": 1583403178684, "user_tz": -60, "elapsed": 7024, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}}
# !git commit -m 'add data visualization'
# + id="tASnsY0u5O7s" colab_type="code" outputId="ee6b0985-8e40-418b-f0a7-909fa0b7f5c2" executionInfo={"status": "ok", "timestamp": 1583131138165, "user_tz": -60, "elapsed": 1907, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 104}
# !git commit -m 'add data directory'
# + id="v0k4ElAG4Vx3" colab_type="code" outputId="161b4353-0241-4630-de69-1efdd835059d" executionInfo={"status": "ok", "timestamp": 1583403195055, "user_tz": -60, "elapsed": 3125, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 52}
# !git push -u origin master
# + id="Q-EKS8h45yd3" colab_type="code" colab={}
# #!git pull# origin master
# + id="48U5wcCC5FaS" colab_type="code" outputId="e3a980cd-107c-4af0-d397-46127b4a182e" executionInfo={"status": "ok", "timestamp": 1583131290022, "user_tz": -60, "elapsed": 767, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 54}
# cd data
# + id="iOCC84Vz-mTs" colab_type="code" outputId="e85ccfce-d242-4b5a-aecf-0e37f83a31af" executionInfo={"status": "ok", "timestamp": 1583131419003, "user_tz": -60, "elapsed": 4089, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 141}
# !curl -L http://bit.ly/dw_car_data -o car.h5
# + id="orSvlVKt_FEE" colab_type="code" outputId="41fde423-c377-46df-afa8-e6c7ea99d601" executionInfo={"status": "ok", "timestamp": 1583131457691, "user_tz": -60, "elapsed": 1628, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls
# + id="Iqa9Ykzb_PBT" colab_type="code" outputId="91664f17-fd4a-4a89-93a9-e180ca3dd32b" executionInfo={"status": "ok", "timestamp": 1583131468563, "user_tz": -60, "elapsed": 2293, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 52}
# !ls -lh
# + id="v8YGrw1nAG67" colab_type="code" outputId="7840f18d-f05e-4450-c320-814a1f47fbf3" executionInfo={"status": "ok", "timestamp": 1583131700536, "user_tz": -60, "elapsed": 2353, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# ls
# + id="fuchkGqDALBT" colab_type="code" outputId="913676af-6258-4a4e-9bf1-1429f0855a5f" executionInfo={"status": "ok", "timestamp": 1583131707745, "user_tz": -60, "elapsed": 852, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 54}
# cd cars_forecasting/
# + id="3yzynmtN_Rlk" colab_type="code" colab={}
import pandas as pd
# + id="QQE_BsXn_UWR" colab_type="code" colab={}
df = pd.read_hdf('data/car.h5')
# + id="pZ9V7n-d_YaO" colab_type="code" outputId="b424af54-c4f3-4a4d-9c3d-48c6d0b16a66" executionInfo={"status": "ok", "timestamp": 1583131571580, "user_tz": -60, "elapsed": 7121, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 297}
# !pip install --upgrade tables
# + id="pg-97buoAU2W" colab_type="code" outputId="1d53d003-693a-4b94-b248-2a4e370e1fd2" executionInfo={"status": "ok", "timestamp": 1583131746861, "user_tz": -60, "elapsed": 401, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df.shape
# + id="MHNMH4W5_pks" colab_type="code" outputId="d03a7165-001f-42bc-9ce0-8ce5c6ac8211" executionInfo={"status": "ok", "timestamp": 1583131758122, "user_tz": -60, "elapsed": 655, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02481780017115552185"}} colab={"base_uri": "https://localhost:8080/", "height": 585}
df.sample(5)
# + id="LFAWjmmaAYsJ" colab_type="code" colab={}
|
Day_1_car_prices_forecasting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:softlearning-sim-new] *
# language: python
# name: conda-env-softlearning-sim-new-py
# ---
# +
import glob
import matplotlib.pyplot as plt
import numpy as np
import os
import pickle
import sys
import copy
import matplotlib
# %matplotlib inline
sys.path.append("..")
from demo_2_awac import och_2_awac
import adept_envs
import gym
# Load in the data
all_paths = 'demo_list_2elements_PLAYSTYLE_June14.pkl'
paths = pickle.load(open(all_paths, 'rb'))
import itertools
def check_goal_completion(curr_pos):
max_objs = np.array([0.17, 0.6])
min_objs = np.array([0.08, 0.2])
init_bitflips = np.array([0, 0])
curr_bitflips = init_bitflips.copy()
for j, pos_idx in enumerate([0, 2]):
if curr_pos[pos_idx] > max_objs[j]:
curr_bitflips[j] = 1
elif curr_pos[pos_idx] < min_objs[j]:
curr_bitflips[j] = 0
new_idx = 2*curr_bitflips[0] + curr_bitflips[1]
return new_idx
input_x = []
for path in paths:
input_x.append(path['observations'][:-100])
input_x = np.concatenate(input_x, axis=0)[:, 2:6]
input_y = []
start_idxs = []
end_idxs = []
# Get labels
prev_idx = -1
curr_idx = -1
window = 5
state_final_goal_nextgoal_tuples = []
continuous_paths = []
curr_continuous = []
idx_vals = [[] for _ in range(4)]
for j, path in enumerate(paths):
curr_idx = check_goal_completion(path['observations'][0, 2:6])
if curr_idx != prev_idx and prev_idx != -1:
continuous_paths.append(curr_continuous)
curr_continuous = []
curr_continuous.append(path)
prev_idx = check_goal_completion(path['observations'][-1, 2:6])
idx_vals[prev_idx].append(path['observations'][-1, 2:6])
continuous_paths.append(curr_continuous)
# -
for path in continuous_paths[0]:
print(check_goal_completion(path['observations'][0, 2:6]))
delta_skip = 3
state_nextgoal_finalgoal_tuples = []
for curr_continuous in continuous_paths:
for j, path in enumerate(curr_continuous):
leftover_skips = min(len(curr_continuous) - j, delta_skip)
for k in range(len(path['observations']) - 100):
for skips in range(leftover_skips):
curr_ng = check_goal_completion(path['observations'][-1, 2:6])
final_ng = check_goal_completion(curr_continuous[j + skips]['observations'][-1, 2:6])
snf_tuple = (path['observations'][k][:9], curr_ng, final_ng)
state_nextgoal_finalgoal_tuples.append(snf_tuple)
state_nextgoal_finalgoal_tuples
# +
input_x = []
for snf_tuple in state_nextgoal_finalgoal_tuples:
input_x.append([np.concatenate([snf_tuple[0][2:6], [snf_tuple[2]]])])
input_x = np.concatenate(input_x, axis=0)
input_y = []
for snf_tuple in state_nextgoal_finalgoal_tuples:
input_y.append(snf_tuple[1])
input_y = np.array(input_y)
# -
input_x.shape
# +
from torch.utils.data import TensorDataset, DataLoader
from rlkit.torch.networks import ConcatMlp, Mlp
import torch
import torch.nn as nn
input_size = 5
num_goals = 4
batch_size = 32
goal_predictor = Mlp(
input_size=input_size,
output_size=num_goals,
hidden_sizes=(256, 256, 256),
)
optimizer = torch.optim.Adam(goal_predictor.parameters())
idxs_data = np.array(range(len(input_x)))
np.random.shuffle(idxs_data)
num_train = int(0.9*len(idxs_data))
input_x_train = input_x[idxs_data[:num_train]]
input_x_test = input_x[idxs_data[num_train:]]
input_y_train = input_y[idxs_data[:num_train]]
input_y_test = input_y[idxs_data[num_train:]]
# TODO: Check the syntax
train_ds = TensorDataset(torch.Tensor(input_x_train), torch.Tensor(input_y_train).long())
test_ds = TensorDataset(torch.Tensor(input_x_test), torch.Tensor(input_y_test).long())
trainloader = torch.utils.data.DataLoader(train_ds, batch_size=batch_size, shuffle=True)
criterion = nn.CrossEntropyLoss()
# +
num_epochs = 10
for epoch in range(num_epochs): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = goal_predictor(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 100 == 0: # print every 2000 mini-batches
print('[%d, %5d] loss: %.8f' %
(epoch + 1, i + 1, running_loss / 2000))
running_loss = 0.0
outputs = goal_predictor(torch.Tensor(input_x_test))
accuracy = np.sum(np.argmax(outputs.detach().numpy(), axis=1) == np.array(input_y_test, dtype=np.int32))/len(input_x_test)
# print('TEST %f \n'%accuracy*100)
print('Finished Training')
# +
start = 3
curr_s = idx_vals[start][np.random.randint(len(idx_vals[start]))]
curr_g = 1
pl = []
print("STARTING AT %d"%start)
for k in range(100):
start = 1
curr_s = idx_vals[start][np.random.randint(len(idx_vals[start]))]
curr_g = 2
print("============")
for j in range(5):
o = np.concatenate([curr_s, [curr_g]])
o = torch.Tensor(o)
prediction = torch.nn.Softmax()(goal_predictor(o)).detach().numpy()
next_idx = np.random.choice(range(4), p =prediction)
print("GOING TO %d"%next_idx)
curr_s = idx_vals[next_idx][np.random.randint(len(idx_vals[next_idx]))]
if next_idx == curr_g:
break
pl.append(j + 1)
np.array(pl).mean()
# -
torch.save(goal_predictor, 'BC_model_2elems_window3.pkl')
dat = torch.load('BC_model_2elems_window3.pkl')
goal_predictor.load_state_dict(dat.state_dict())
|
third_party/rlkit_library/notebooks/train_goalBC_sim_relabeled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.11 64-bit (''tf2'': conda)'
# name: python3
# ---
# <i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
#
# <i>Licensed under the MIT License.</i>
# # Neural Collaborative Filtering (NCF)
#
# This notebook serves as an introduction to Neural Collaborative Filtering (NCF), which is an innovative algorithm based on deep neural networks to tackle the key problem in recommendation — collaborative filtering — on the basis of implicit feedback.
# ## 0 Global Settings and Imports
# +
import sys
import os
import shutil
import papermill as pm
import scrapbook as sb
import pandas as pd
import numpy as np
import tensorflow as tf
tf.get_logger().setLevel('ERROR') # only show error messages
from recommenders.utils.timer import Timer
from recommenders.models.ncf.ncf_singlenode import NCF
from recommenders.models.ncf.dataset import Dataset as NCFDataset
from recommenders.datasets import movielens
from recommenders.datasets.python_splitters import python_chrono_split
from recommenders.evaluation.python_evaluation import (rmse, mae, rsquared, exp_var, map_at_k, ndcg_at_k, precision_at_k,
recall_at_k, get_top_k_items)
from recommenders.utils.constants import SEED as DEFAULT_SEED
print("System version: {}".format(sys.version))
print("Pandas version: {}".format(pd.__version__))
print("Tensorflow version: {}".format(tf.__version__))
# + tags=["parameters"]
# top k items to recommend
TOP_K = 10
# Select MovieLens data size: 100k, 1m, 10m, or 20m
MOVIELENS_DATA_SIZE = '100k'
# Model parameters
EPOCHS = 100
BATCH_SIZE = 256
SEED = DEFAULT_SEED # Set None for non-deterministic results
# -
# ## 1 Matrix factorization algorithm
#
# NCF is new neural matrix factorization model, which ensembles Generalized Matrix Factorization (GMF) and Multi-Layer Perceptron (MLP) to unify the strengths of linearity of MF and non-linearity of MLP for modelling the user–item latent structures. NCF can be demonstrated as a framework for GMF and MLP, which is illustrated as below:
# <img src="https://recodatasets.z20.web.core.windows.net/images/NCF.svg?sanitize=true">
#
# This figure shows how to utilize latent vectors of items and users, and then how to fuse outputs from GMF Layer (left) and MLP Layer (right). We will introduce this framework and show how to learn the model parameters in following sections.
#
# ### 1.1 The GMF model
#
# In ALS, the ratings are modeled as follows:
#
# $$\hat { r } _ { u , i } = q _ { i } ^ { T } p _ { u }$$
#
# GMF introduces neural CF layer as the output layer of standard MF. In this way, MF can be easily generalized
# and extended. For example, if we allow the edge weights of this output layer to be learnt from data without the uniform constraint, it will result in a variant of MF that allows varying importance of latent dimensions. And if we use a non-linear function for activation, it will generalize MF to a non-linear setting which might be more expressive than the linear MF model. GMF can be shown as follows:
#
# $$\hat { r } _ { u , i } = a _ { o u t } \left( h ^ { T } \left( q _ { i } \odot p _ { u } \right) \right)$$
#
# where $\odot$ is element-wise product of vectors. Additionally, ${a}_{out}$ and ${h}$ denote the activation function and edge weights of the output layer respectively. MF can be interpreted as a special case of GMF. Intuitively, if we use an identity function for aout and enforce h to be a uniform vector of 1, we can exactly recover the MF model.
#
# ### 1.2 The MLP model
#
# NCF adopts two pathways to model users and items: 1) element-wise product of vectors, 2) concatenation of vectors. To learn interactions after concatenating of users and items latent features, the standard MLP model is applied. In this sense, we can endow the model a large level of flexibility and non-linearity to learn the interactions between $p_{u}$ and $q_{i}$. The details of MLP model are:
#
# For the input layer, there is concatention of user and item vectors:
#
# $$z _ { 1 } = \phi _ { 1 } \left( p _ { u } , q _ { i } \right) = \left[ \begin{array} { c } { p _ { u } } \\ { q _ { i } } \end{array} \right]$$
#
# So for the hidden layers and output layer of MLP, the details are:
#
# $$
# \phi _ { l } \left( z _ { l } \right) = a _ { o u t } \left( W _ { l } ^ { T } z _ { l } + b _ { l } \right) , ( l = 2,3 , \ldots , L - 1 )
# $$
#
# and:
#
# $$
# \hat { r } _ { u , i } = \sigma \left( h ^ { T } \phi \left( z _ { L - 1 } \right) \right)
# $$
#
# where ${ W }_{ l }$, ${ b }_{ l }$, and ${ a }_{ out }$ denote the weight matrix, bias vector, and activation function for the $l$-th layer’s perceptron, respectively. For activation functions of MLP layers, one can freely choose sigmoid, hyperbolic tangent (tanh), and Rectifier (ReLU), among others. Because of binary data task, the activation function of the output layer is defined as sigmoid $\sigma(x)=\frac{1}{1+e^{-x}}$ to restrict the predicted score to be in (0,1).
#
#
# ### 1.3 Fusion of GMF and MLP
#
# To provide more flexibility to the fused model, we allow GMF and MLP to learn separate embeddings, and combine the two models by concatenating their last hidden layer. We get $\phi^{GMF}$ from GMF:
#
# $$\phi _ { u , i } ^ { G M F } = p _ { u } ^ { G M F } \odot q _ { i } ^ { G M F }$$
#
# and obtain $\phi^{MLP}$ from MLP:
#
# $$\phi _ { u , i } ^ { M L P } = a _ { o u t } \left( W _ { L } ^ { T } \left( a _ { o u t } \left( \ldots a _ { o u t } \left( W _ { 2 } ^ { T } \left[ \begin{array} { c } { p _ { u } ^ { M L P } } \\ { q _ { i } ^ { M L P } } \end{array} \right] + b _ { 2 } \right) \ldots \right) \right) + b _ { L }\right.$$
#
# Lastly, we fuse output from GMF and MLP:
#
# $$\hat { r } _ { u , i } = \sigma \left( h ^ { T } \left[ \begin{array} { l } { \phi ^ { G M F } } \\ { \phi ^ { M L P } } \end{array} \right] \right)$$
#
# This model combines the linearity of MF and non-linearity of DNNs for modelling user–item latent structures.
#
# ### 1.4 Objective Function
#
# We define the likelihood function as:
#
# $$P \left( \mathcal { R } , \mathcal { R } ^ { - } | \mathbf { P } , \mathbf { Q } , \Theta \right) = \prod _ { ( u , i ) \in \mathcal { R } } \hat { r } _ { u , i } \prod _ { ( u , j ) \in \mathcal { R } ^{ - } } \left( 1 - \hat { r } _ { u , j } \right)$$
#
# Where $\mathcal{R}$ denotes the set of observed interactions, and $\mathcal{ R } ^ { - }$ denotes the set of negative instances. $\mathbf{P}$ and $\mathbf{Q}$ denotes the latent factor matrix for users and items, respectively; and $\Theta$ denotes the model parameters. Taking the negative logarithm of the likelihood, we obatain the objective function to minimize for NCF method, which is known as [binary cross-entropy loss](https://en.wikipedia.org/wiki/Cross_entropy):
#
# $$L = - \sum _ { ( u , i ) \in \mathcal { R } \cup { \mathcal { R } } ^ { - } } r _ { u , i } \log \hat { r } _ { u , i } + \left( 1 - r _ { u , i } \right) \log \left( 1 - \hat { r } _ { u , i } \right)$$
#
# The optimization can be done by performing Stochastic Gradient Descent (SGD), which is described in the [Surprise SVD deep dive notebook](../02_model/surprise_svd_deep_dive.ipynb). Our SGD method is very similar to the SVD algorithm's.
# ## 2 TensorFlow implementation of NCF
#
# We will use the MovieLens dataset, which is composed of integer ratings from 1 to 5.
#
# We convert MovieLens into implicit feedback, and evaluate under our *leave-one-out* evaluation protocol.
#
# You can check the details of implementation in `recommenders/models/ncf`
#
# ## 3 TensorFlow NCF movie recommender
#
# ### 3.1 Load and split data
#
# To evaluate the performance of item recommendation, we adopted the leave-one-out evaluation.
#
# For each user, we held out his/her latest interaction as the test set and utilized the remaining data for training. We use `python_chrono_split` to achieve this. And since it is too time-consuming to rank all items for every user during evaluation, we followed the common strategy that randomly samples 100 items that are not interacted by the user, ranking the test item among the 100 items. Our test samples will be constructed by `NCFDataset`.
# +
df = movielens.load_pandas_df(
size=MOVIELENS_DATA_SIZE,
header=["userID", "itemID", "rating", "timestamp"]
)
df.head()
# -
train, test = python_chrono_split(df, 0.75)
# ### 3.2 Functions of NCF Dataset
#
# Dataset Class for NCF, where important functions are:
#
# `negative_sampling()`, sample negative user & item pair for every positive instances, with parameter `n_neg`.
#
# `train_loader(batch_size, shuffle=True)`, generate training batch with `batch_size`, also we can set whether `shuffle` this training set.
#
# `test_loader()`, generate test batch by every positive test instance, (eg. \[1, 2, 1\] is a positive user & item pair in test set (\[userID, itemID, rating\] for this tuple). This function returns like \[\[1, 2, 1\], \[1, 3, 0\], \[1,6, 0\], ...\], ie. following our *leave-one-out* evaluation protocol.
data = NCFDataset(train=train, test=test, seed=SEED)
# ### 3.3 Train NCF based on TensorFlow
# The NCF has a lot of parameters. The most important ones are:
#
# `n_factors`, which controls the dimension of the latent space. Usually, the quality of the training set predictions grows with as n_factors gets higher.
#
# `layer_sizes`, sizes of input layer (and hidden layers) of MLP, input type is list.
#
# `n_epochs`, which defines the number of iteration of the SGD procedure.
# Note that both parameter also affect the training time.
#
# `model_type`, we can train single `"MLP"`, `"GMF"` or combined model `"NCF"` by changing the type of model.
#
# We will here set `n_factors` to `4`, `layer_sizes` to `[16,8,4]`, `n_epochs` to `100`, `batch_size` to 256. To train the model, we simply need to call the `fit()` method.
model = NCF (
n_users=data.n_users,
n_items=data.n_items,
model_type="NeuMF",
n_factors=4,
layer_sizes=[16,8,4],
n_epochs=EPOCHS,
batch_size=BATCH_SIZE,
learning_rate=1e-3,
verbose=10,
seed=SEED
)
# +
with Timer() as train_time:
model.fit(data)
print("Took {} seconds for training.".format(train_time.interval))
# -
# ## 3.4 Prediction and Evaluation
# ### 3.4.1 Prediction
#
# Now that our model is fitted, we can call `predict` to get some `predictions`. `predict` returns an internal object Prediction which can be easily converted back to a dataframe:
# +
predictions = [[row.userID, row.itemID, model.predict(row.userID, row.itemID)]
for (_, row) in test.iterrows()]
predictions = pd.DataFrame(predictions, columns=['userID', 'itemID', 'prediction'])
predictions.head()
# -
# ### 3.4.2 Generic Evaluation
# We remove rated movies in the top k recommendations
# To compute ranking metrics, we need predictions on all user, item pairs. We remove though the items already watched by the user, since we choose not to recommend them again.
# +
with Timer() as test_time:
users, items, preds = [], [], []
item = list(train.itemID.unique())
for user in train.userID.unique():
user = [user] * len(item)
users.extend(user)
items.extend(item)
preds.extend(list(model.predict(user, item, is_list=True)))
all_predictions = pd.DataFrame(data={"userID": users, "itemID":items, "prediction":preds})
merged = pd.merge(train, all_predictions, on=["userID", "itemID"], how="outer")
all_predictions = merged[merged.rating.isnull()].drop('rating', axis=1)
print("Took {} seconds for prediction.".format(test_time.interval))
# +
eval_map = map_at_k(test, all_predictions, col_prediction='prediction', k=TOP_K)
eval_ndcg = ndcg_at_k(test, all_predictions, col_prediction='prediction', k=TOP_K)
eval_precision = precision_at_k(test, all_predictions, col_prediction='prediction', k=TOP_K)
eval_recall = recall_at_k(test, all_predictions, col_prediction='prediction', k=TOP_K)
print("MAP:\t%f" % eval_map,
"NDCG:\t%f" % eval_ndcg,
"Precision@K:\t%f" % eval_precision,
"Recall@K:\t%f" % eval_recall, sep='\n')
# -
# ### 3.4.3 "Leave-one-out" Evaluation
#
# We implement the functions to repoduce the leave-one-out evaluation protocol mentioned in original NCF paper.
#
# For each item in test data, we randomly samples 100 items that are not interacted by the user, ranking the test item among the 101 items (1 positive item and 100 negative items). The performance of a ranked list is judged by **Hit Ratio (HR)** and **Normalized Discounted Cumulative Gain (NDCG)**. Finally, we average the values of those ranked lists to obtain the overall HR and NDCG on test data.
#
# We truncated the ranked list at 10 for both metrics. As such, the HR intuitively measures whether the test item is present on the top-10 list, and the NDCG accounts for the position of the hit by assigning higher scores to hits at top ranks.
#
# **Note 1:** In exact leave-one-out evaluation protocol, we select only one of the latest items interacted with a user as test data for each user. But in this notebook, to compare with other algorithms, we select latest 25% dataset as test data. So this is an artificial "leave-one-out" evaluation only showing how to use `test_loader` and how to calculate metrics like the original paper. You can reproduce the real leave-one-out evaluation by changing the way of splitting data.
#
# **Note 2:** Because of sampling 100 negative items for each positive test item,
# +
k = TOP_K
ndcgs = []
hit_ratio = []
for b in data.test_loader():
user_input, item_input, labels = b
output = model.predict(user_input, item_input, is_list=True)
output = np.squeeze(output)
rank = sum(output >= output[0])
if rank <= k:
ndcgs.append(1 / np.log(rank + 1))
hit_ratio.append(1)
else:
ndcgs.append(0)
hit_ratio.append(0)
eval_ndcg = np.mean(ndcgs)
eval_hr = np.mean(hit_ratio)
print("HR:\t%f" % eval_hr)
print("NDCG:\t%f" % eval_ndcg)
# -
# ## 3.5 Pre-training
#
# To get better performance of NeuMF, we can adopt pre-training strategy. We first train GMF and MLP with random initializations until convergence. Then use their model parameters as the initialization for the corresponding parts of NeuMF’s parameters. Please pay attention to the output layer, where we concatenate weights of the two models with
#
# $$h ^ { N C F } \leftarrow \left[ \begin{array} { c } { \alpha h ^ { G M F } } \\ { ( 1 - \alpha ) h ^ { M L P } } \end{array} \right]$$
#
# where $h^{GMF}$ and $h^{MLP}$ denote the $h$ vector of the pretrained GMF and MLP model, respectively; and $\alpha$ is a
# hyper-parameter determining the trade-off between the two pre-trained models. We set $\alpha$ = 0.5.
# ### 3.5.1 Training GMF and MLP model
# `model.save`, we can set the `dir_name` to store the parameters of GMF and MLP
model = NCF (
n_users=data.n_users,
n_items=data.n_items,
model_type="GMF",
n_factors=4,
layer_sizes=[16,8,4],
n_epochs=EPOCHS,
batch_size=BATCH_SIZE,
learning_rate=1e-3,
verbose=10,
seed=SEED
)
# +
with Timer() as train_time:
model.fit(data)
print("Took {} seconds for training.".format(train_time.interval))
model.save(dir_name=".pretrain/GMF")
# +
model = NCF (
n_users=data.n_users,
n_items=data.n_items,
model_type="MLP",
n_factors=4,
layer_sizes=[16,8,4],
n_epochs=EPOCHS,
batch_size=BATCH_SIZE,
learning_rate=1e-3,
verbose=10,
seed=SEED
)
# +
with Timer() as train_time:
model.fit(data)
print("Took {} seconds for training.".format(train_time.interval))
model.save(dir_name=".pretrain/MLP")
# -
# ### 3.5.2 Load pre-trained GMF and MLP model for NeuMF
# `model.load`, we can set the `gmf_dir` and `mlp_dir` to store the parameters for NeuMF.
# +
model = NCF (
n_users=data.n_users,
n_items=data.n_items,
model_type="NeuMF",
n_factors=4,
layer_sizes=[16,8,4],
n_epochs=EPOCHS,
batch_size=BATCH_SIZE,
learning_rate=1e-3,
verbose=10,
seed=SEED
)
model.load(gmf_dir=".pretrain/GMF", mlp_dir=".pretrain/MLP", alpha=0.5)
# +
with Timer() as train_time:
model.fit(data)
print("Took {} seconds for training.".format(train_time.interval))
# -
# ### 3.5.3 Compare with not pre-trained NeuMF
#
# You can use beforementioned evaluation methods to evaluate the pre-trained `NCF` Model. Usually, we will find the performance of pre-trained NCF is better than the not pre-trained.
# +
with Timer() as test_time:
users, items, preds = [], [], []
item = list(train.itemID.unique())
for user in train.userID.unique():
user = [user] * len(item)
users.extend(user)
items.extend(item)
preds.extend(list(model.predict(user, item, is_list=True)))
all_predictions = pd.DataFrame(data={"userID": users, "itemID":items, "prediction":preds})
merged = pd.merge(train, all_predictions, on=["userID", "itemID"], how="outer")
all_predictions = merged[merged.rating.isnull()].drop('rating', axis=1)
print("Took {} seconds for prediction.".format(test_time.interval))
# +
eval_map2 = map_at_k(test, all_predictions, col_prediction='prediction', k=TOP_K)
eval_ndcg2 = ndcg_at_k(test, all_predictions, col_prediction='prediction', k=TOP_K)
eval_precision2 = precision_at_k(test, all_predictions, col_prediction='prediction', k=TOP_K)
eval_recall2 = recall_at_k(test, all_predictions, col_prediction='prediction', k=TOP_K)
print("MAP:\t%f" % eval_map2,
"NDCG:\t%f" % eval_ndcg2,
"Precision@K:\t%f" % eval_precision2,
"Recall@K:\t%f" % eval_recall2, sep='\n')
# -
# Record results with papermill for tests
sb.glue("map", eval_map)
sb.glue("ndcg", eval_ndcg)
sb.glue("precision", eval_precision)
sb.glue("recall", eval_recall)
sb.glue("map2", eval_map2)
sb.glue("ndcg2", eval_ndcg2)
sb.glue("precision2", eval_precision2)
sb.glue("recall2", eval_recall2)
# ### 3.5.4 Delete pre-trained directory
# +
save_dir = ".pretrain"
if os.path.exists(save_dir):
shutil.rmtree(save_dir)
print("Did \'%s\' exist?: %s" % (save_dir, os.path.exists(save_dir)))
# -
# ### Reference:
# 1. <NAME>, <NAME>, <NAME>, <NAME>, <NAME> & <NAME>, Neural Collaborative Filtering, 2017, https://arxiv.org/abs/1708.05031
#
# 2. Official NCF implementation [Keras with Theano]: https://github.com/hexiangnan/neural_collaborative_filtering
#
# 3. Other nice NCF implementation [Pytorch]: https://github.com/LaceyChen17/neural-collaborative-filtering
|
examples/02_model_hybrid/ncf_deep_dive.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Load the data
import pandas as pd
X = pd.read_csv('../data/tree_class_feats.csv')
y = pd.read_csv('../data/tree_class_target.csv')
# Split the dataset into training set and test set with a 80-20 ratio
from sklearn.model_selection import train_test_split
seed = 1
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=seed)
# +
# Define your model
from keras.models import Sequential
from keras.layers import Dense, Activation
import numpy as np
from tensorflow import random
np.random.seed(seed)
random.set_seed(seed)
model_1 = Sequential()
model_1.add(Dense(16, activation='relu', input_dim=X_train.shape[1]))
model_1.add(Dense(8, activation='relu'))
model_1.add(Dense(4, activation='relu'))
model_1.add(Dense(1, activation='sigmoid'))
# Choose the loss function to be binary cross entropy and the optimizer to be SGD for training the model
model_1.compile(optimizer='sgd', loss='binary_crossentropy')
# train the model
history = model_1.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=300, batch_size=50, verbose=0, shuffle=False)
# +
# import require packages for plotting
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
# plot training error and validation error
matplotlib.rcParams['figure.figsize'] = (10.0, 8.0)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.ylim(0,1)
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train loss', 'validation loss'], loc='upper right')
# +
#Define your model with early stopping on test error
from keras.callbacks import EarlyStopping
np.random.seed(seed)
random.set_seed(seed)
model_2 = Sequential()
model_2.add(Dense(16, activation='relu', input_dim=X_train.shape[1]))
model_2.add(Dense(8, activation='relu'))
model_2.add(Dense(4, activation='relu'))
model_2.add(Dense(1, activation='sigmoid'))
# Choose the loss function to be binary cross entropy and the optimizer to be SGD for training the model
model_2.compile(optimizer='sgd', loss='binary_crossentropy')
# define the early stopping callback
es_callback = EarlyStopping(monitor='val_loss', mode='min')
# train the model
history=model_2.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=300, batch_size=50, callbacks=[es_callback], verbose=0, shuffle=False)
# -
# plot training error and test error
matplotlib.rcParams['figure.figsize'] = (10.0, 8.0)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.ylim(0,1)
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train loss', 'validation loss'], loc='upper right')
# +
#Define your model with early stopping on test error with patience=10
from keras.callbacks import EarlyStopping
np.random.seed(seed)
random.set_seed(seed)
model_3 = Sequential()
model_3.add(Dense(16, activation='relu', input_dim=X_train.shape[1]))
model_3.add(Dense(8, activation='relu'))
model_3.add(Dense(4, activation='relu'))
model_3.add(Dense(1, activation='sigmoid'))
# Choose the loss function to be binary cross entropy and the optimizer to be SGD for training the model
model_3.compile(optimizer='sgd', loss='binary_crossentropy')
# define the early stopping callback
es_callback = EarlyStopping(monitor='val_loss', mode='min', patience=10)
# train the model
history=model_3.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=300, batch_size=50, callbacks=[es_callback], verbose=0, shuffle=False)
# -
# plot training error and test error
matplotlib.rcParams['figure.figsize'] = (10.0, 8.0)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.ylim(0,1)
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train loss', 'validation loss'], loc='upper right')
|
Exercise02/Exercise02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="IDuphEV0ufdm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 151} outputId="acd571e1-e7b5-48cd-bdfc-4ac8047b2778"
# !pip install download
# + id="JdBMQ9sOutJ5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="aebfd3fe-e034-46a9-8878-aefcd673f94d"
from __future__ import absolute_import,division,print_function,unicode_literals
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import os
from datetime import datetime
import pandas as pd
from download import download
mpl.rcParams['figure.figsize'] = (8,6)
mpl.rcParams['axes.grid'] = False
print("Import Succesfull")
# + id="8gMd2suEuw3B" colab_type="code" colab={}
def parse(x):
return datetime.strptime(x, '%m/%d/%Y')
# + id="G-gfJoCou5HW" colab_type="code" colab={}
df = pd.read_csv('https://raw.githubusercontent.com/srivatsan88/YouTubeLI/master/dataset/electricity_consumption.csv', parse_dates=['Bill_Date'], date_parser=parse)
# + id="NyOcyTYwvLz7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 403} outputId="fb4d5aad-390b-4429-8d03-ce81b24db9e2"
print("Rows :", df.shape[0])
print("Columns :", df.shape[1])
print("\n Features \n", df.columns.to_list())
print("\n Missing Values \n", df.isnull().any())
print("\n Unique Values \n", df.nunique())
# + id="ermdkrPQxKlK" colab_type="code" colab={}
bill_df = df.set_index('Bill_Date')
# + id="IqRdvaGXxR6G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="282c6aa4-5462-4780-b27a-fa7733071fbe"
bill_df.head(2)
# + id="OMtHn9sqxYDy" colab_type="code" colab={}
bill_2018 = bill_df['2016':'2018'][['Billed_amount']]
# + id="kj1qpBcfxlFI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="66d4a6d6-f160-4d5c-fe84-c1372914b972"
bill_2018
# + [markdown] id="qjWO78FpxyMI" colab_type="text"
# #Simple Moving Average
#
#
# The Simple Moving Average (SMA) is calculated by adding the price of an instrument over a number of time periods and then dividing the sum by the number of time periods. The SMA is basically the average price of the given time period, with equal weighting given to the price of each period.
#
# In financial applications a simple moving average (SMA) is the unweighted mean of the previous n data. However, in science and engineering, the mean is normally taken from an equal number of data on either side of a central value. This ensures that variations in the mean are aligned with the variations in the data rather than being shifted in time.
#
# Mathmatically (t+(t-1)+(t-2)+...+(t-n))/n
#
# Moving Average cant be a great tool when, the data is not stationary and fluctuating.
#
# + id="e2L_NVky0Gyx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 655} outputId="0aca7c70-ba1f-44de-926a-7187ab31b05a"
bill_2018['Billed_amount'].rolling(window=3).mean()
# + id="_XvUSM_O15xx" colab_type="code" colab={}
bill_2018['ma_rolling_3']= bill_2018['Billed_amount'].rolling(window=3).mean().shift(1)
# + id="OJk4F7CT2Frd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="f677f9b3-41e3-46a3-8e1a-7bc068e25ec1"
bill_2018
# + id="_19Pd1565wQF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 418} outputId="7f304382-b2de-426f-ee19-42d51ce7cb84"
bill_2018.plot()
# + [markdown] id="SArAHy0p83tS" colab_type="text"
# #Weighted Moving Average
#
# Weighted moving averages can find trens sooner than SMA, on the other hand its complex as we need to assign the weights manually.
# + id="jLSYPHZ49Eq-" colab_type="code" colab={}
def wma(weights):
def calc(x):
return (weights*x).mean()
return calc
# + id="knwCJ-izA6z7" colab_type="code" colab={}
#The weights should add up to the window value
bill_2018['wma_rolling_3']= bill_2018['Billed_amount'].rolling(window = 3).apply(wma(np.array([0.5,1,1.5]))).shift(3)
# + id="pG9xj7EFBZAk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="412a766b-d491-4610-b4ba-2b87463a81ed"
bill_2018
# + id="B-FaANn2B_tQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 418} outputId="6a13efc9-222a-4baf-eaab-73509369c567"
bill_2018.plot()
# + [markdown] id="XC3-UvxuCi86" colab_type="text"
# #Exponential Moving Average
#
# It adopts quickly to the data point changes ,and we dont have to decide the weights manually.
#
# α denote a "smoothing constant" (a number between 0 and 1).
#
# + id="VGOZ7HmhCmPt" colab_type="code" colab={}
bill_2018['ewm_window_3']= bill_2018['Billed_amount'].ewm(span = 3,adjust = False,min_periods = 0).mean().shift(1)
# + id="pR80M8cvDIDz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3c5fdd1f-21ff-449b-aea4-15ec9c6e2603"
bill_2018
# + [markdown] id="naSz8eogD-E5" colab_type="text"
# #Exponential Smoothing
#
# It Requires a parameter called alpha aka Smoothing parameter
#
# Larger value of alpha means the model is paying attention to the newer values,smaller value of alpha mean model is giving importance to history value.
#
# + id="shJSXaLnEHJq" colab_type="code" colab={}
bill_2018['esm_windiw_3_7'] = bill_2018['Billed_amount'].ewm(alpha = 0.7 , adjust = False ,min_periods = 3).mean().shift(1)
# + id="0ubzbg_sFjWM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 418} outputId="07226999-cb93-49d6-b242-04e51adb0837"
bill_2018.plot()
# + id="a5X43xIzFq0k" colab_type="code" colab={}
bill_2018['esm_windiw_3_3'] = bill_2018['Billed_amount'].ewm(alpha = 0.3 , adjust = False ,min_periods = 3).mean().shift(1)
# higher alpha takes , most recent values ,
# lower alpha means , more historical data
# + id="F4qYAD3uGhLm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 418} outputId="c3f09aee-b52d-4be1-ef39-d5cb00c7d926"
bill_2018[['Billed_amount','esm_windiw_3_7','esm_windiw_3_3']].plot()
# + id="7XgZxFhxHIUg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="174d2de1-abb3-4094-869f-d1fa04358514"
bill_2018
# + [markdown] id="mpG4CnRUGtwi" colab_type="text"
# #Evaluation
# + id="x7oLHVQpGtbw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e752bc6f-9941-4598-978d-fa020233e0b1"
((bill_2018['Billed_amount']-bill_2018['ma_rolling_3'])**2).mean()**0.5
# + id="RVt-LkU-GtYZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a5d7c707-21c1-4402-e724-9f36a9780467"
((bill_2018['Billed_amount']-bill_2018['wma_rolling_3'])**2).mean()**0.5
# + id="WxfYvakJHy0i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7b8471a2-e016-4d9d-df56-bb1100ef1fe5"
((bill_2018['Billed_amount']-bill_2018['ewm_window_3'])**2).mean()**0.5
# + id="cABjjDZcH8nL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="54aaff1b-c744-4360-c86a-526747fbd147"
((bill_2018['Billed_amount']-bill_2018['esm_windiw_3_7'])**2).mean()**0.5
# + id="I1ino_ZDIGxv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="06097d83-92a2-4fdb-8108-33c26d7e248a"
((bill_2018['Billed_amount']-bill_2018['esm_windiw_3_3'])**2).mean()**0.5
|
Time_series_analysis/ts_moving_avg.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table width="100%"> <tr>
# <td style="background-color:#ffffff;">
# <a href="https://qsoftware.lu.lv/index.php/qworld/" target="_blank"><img src="..\images\qworld.jpg" width="35%" align="left"> </a></td>
# <td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
# prepared by <NAME> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
# </td>
# </tr></table>
# <table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# <h2> Coin Flip: A Probabilistic Bit </h2>
#
# [Watch Lecture](https://youtu.be/uGKHEsVcSEs)
#
# <h3> A fair coin </h3>
#
# A coin has two sides: <i>Head</i> and <i>Tail</i>.
#
# After flipping a coin, we can get a Head or a Tail. We can represent these two cases by a single bit:
# <ul>
# <li> 0 represents Head </li>
# <li> 1 represents Tail </li>
# </ul>
# <h3> Flipping a fair coin </h3>
#
# If our coin is fair, then the probabilities of getting a Head and a Tail are equal:
#
# $ p= \dfrac{1}{2} = 0.5 $.
#
# Flipping a fair coin can be represented as an operator:
# <ul>
# <li> $ FairCoin(Head) = \frac{1}{2} Head + \frac{1}{2}Tail $ </li>
# <li> $ FairCoin(Tail) = \frac{1}{2} Head + \frac{1}{2}Tail $ </li>
# </ul>
# $
# FairCoin = \begin{array}{c|cc} & \mathbf{Head} & \mathbf{Tail} \\ \hline \mathbf{Head} & \dfrac{1}{2} & \dfrac{1}{2} \\ \mathbf{Tail} & \dfrac{1}{2} & \dfrac{1}{2} \end{array}
# $
#
# Or, by using 0 and 1:
#
# $
# FairCoin = \begin{array}{c|cc} & \mathbf{0} & \mathbf{1} \\ \hline \mathbf{0} & \dfrac{1}{2} & \dfrac{1}{2} \\ \mathbf{1} & \dfrac{1}{2} & \dfrac{1}{2} \end{array}
# $
# <h3> Task 1: Simulating FairCoin in Python</h3>
#
# Flip a fair coin 100 times. Calculate the total number of heads and tails, and then check the ratio of the number of heads and the number of tails.
#
# Do the same experiment 1000 times.
#
# Do the same experiment 10,000 times.
#
# Do the same experiment 100,000 times.
#
# Do your results get close to the ideal case (the numbers of heads and tails are equal)?
# +
from random import randrange
#
# you may use method 'randrange' for this task
# randrange(n) return a value from {0,1,...,n-1} randomly
#
#
# your solution is here
#
# -
# <a href="B06_Coin_Flip_Solutions.ipynb#task1">click for our solution</a>
# <h3> Flipping a biased coin </h3>
#
# Our coin may have a bias.
#
# For example, the probability of getting head is greater than the probability of getting tail.
#
# Here is an example:
#
# $
# BiasedCoin = \begin{array}{c|cc} & \mathbf{Head} & \mathbf{Tail} \\ \hline \mathbf{Head} & 0.6 & 0.6 \\ \mathbf{Tail} & 0.4 & 0.4 \end{array}
# $
#
# Or, by using 0 and 1 as the states:
#
# $
# BiasedCoin = \begin{array}{c|cc} & \mathbf{0} & \mathbf{1} \\ \hline \mathbf{0} & 0.6 & 0.6\\ \mathbf{1} & 0.4 & 0.4 \end{array}
# $
# <h3> Task 2: Simulating BiasedCoin in Python</h3>
#
# Flip the following biased coin 100 times. Calculate the total numbers of heads and tails, and then check the ratio of the number of heads and the number of tails.
#
# $
# BiasedCoin = \begin{array}{c|cc} & \mathbf{Head} & \mathbf{Tail} \\ \hline \mathbf{Head} & 0.6 & 0.6 \\ \mathbf{Tail} & 0.4 & 0.4 \end{array}
# $
#
#
# Do the same experiment 1000 times.
#
# Do the same experiment 10,000 times.
#
# Do the same experiment 100,000 times.
#
# Do your results get close to the ideal case $ \mypar{ \dfrac{ \mbox{# of heads} }{ \mbox{# of tails} } = \dfrac{0.6}{0.4} = 1.50000000 } $?
# +
#
# you may use method 'randrange' for this task
# randrange(n) return a value from {0,1,...,n-1} randomly
#
#
# your solution is here
#
# -
# <a href="B06_Coin_Flip_Solutions.ipynb#task2">click for our solution</a>
# <h3> Programming a biased coin [extra] </h3>
#
# We use a simple method to create a biased coin.
#
# First, we pick a range for the precision of probabilities, say $ N $, as $ N = 11, 101, 1001, \mbox{ or }, 10^k+1 $ for some $ k > 4 $.
#
# Second, we pick the bias, say $ B $, as an integer between 0 and $ N $.
#
# We fix $ N $ and $ B $.
#
# Third, we pick a random integer between 0 and $ N $:
# <ul>
# <li> if it is less than $ B $, we say "Head" and </li>
# <li> if it is equal to $ B $ or greater than $ B $, we say "Tail" </li>
# </ul>
#
# In this way, we can have a biased coin "landing" head with probability $ \frac{B}{N} $.
#
# Remark that we pick $ N = 10^k+1 $. In this way, the coin cannot be fair if $ B $ is an integer. ($ \frac{10^k+1}{2} $ is not an integer.)
# <h3> Task 3 </h3>
#
# Write a function to implement the described biased coin,
#
# The inputs are integers $ N >0 $ and $ 0 \leq B < N $.
#
# The output is either "Head" or "Tail".
def biased_coin(N,B):
from random import randrange
#
# your solution is here
#
# <a href="B06_Coin_Flip_Solutions.ipynb#task3">click for our solution</a>
# <h3> Task 4</h3>
#
# We use the biased coin defined in Task 3.
#
# (You may use the one given in the solution.)
#
# We pick $ N $ as 101.
#
# We determine $ B $ randomly, and we do not check its value immediately.
#
# The task is to guess the bias by using the biased coin at most 500 times.
#
# Check the real bias, and calculate the error.
from random import randrange
N = 101
B = randrange(100)
#
# your solution is here
#
# <a href="B06_Coin_Flip_Solutions.ipynb#task4">click for our solution</a>
|
bronze/.ipynb_checkpoints/B06_Coin_Flip-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "ddba1d47-1f8d-483d-8d80-b9d493276c9e", "showTitle": false, "title": ""}
def resize(path):
dirs = os.listdir(path)
for item in dirs:
if os.path.isfile(path+item):
im = Image.open(path+item)
f, e = os.path.splitext(path+item)
imResize = im.resize((299,299), Image.ANTIALIAS)
imResize = imResize.convert('RGB')
imResize.save(f+".jpg", 'JPEG', quality=90)
if (e !=".jpg"):
os.remove(path+item)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5ccace78-598c-48b5-ad6e-2a82cfc0b50c", "showTitle": false, "title": ""}
from PIL import Image
import os, sys
path_normal = '/dbfs/FileStore/tables/X-Ray_Image_DataSet/No_findings/'
path_covid = '/dbfs/FileStore/tables/X-Ray_Image_DataSet/Covid-19/'
path_viral = '/dbfs/FileStore/tables/X-Ray_Image_DataSet/Pneumonia/'
resize(path_normal)
resize(path_covid)
resize(path_viral)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "23e01dfc-835e-48c6-8ae0-675e4e235a2d", "showTitle": false, "title": ""}
import tensorflow as tf
import pyspark.sql.functions as f
import sparkdl as dl
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "3a1f80bc-6da3-4094-9d38-ed26217608e8", "showTitle": false, "title": ""}
from sparkdl.image import imageIO
normal_df= imageIO.readImagesWithCustomFn('./FileStore/tables/X-Ray_Image_DataSet/No_findings', decode_f=imageIO.PIL_decode).withColumn('label', f.lit(0))
viral_df= imageIO.readImagesWithCustomFn('./FileStore/tables/X-Ray_Image_DataSet/Pneumonia', decode_f=imageIO.PIL_decode).withColumn('label', f.lit(1))
covid_df= imageIO.readImagesWithCustomFn('./FileStore/tables/X-Ray_Image_DataSet/Covid-19', decode_f=imageIO.PIL_decode).withColumn('label', f.lit(2))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "9f70ffc8-df50-43c3-ac88-5b5f48a68019", "showTitle": false, "title": ""}
normal_train, normal_test= normal_df.randomSplit([0.8,0.2]) # use larger training sets (e.g. [0.6, 0.4] for non-community edition clusters)
covid_train, covid_test= covid_df.randomSplit([0.8,0.2]) # use larger training sets (e.g. [0.6, 0.4] for non-community edition clusters)
viral_train, viral_test= viral_df.randomSplit([0.8,0.2])
train_df = normal_train.unionAll(covid_train).unionAll(viral_train)
test_df = normal_test.unionAll(covid_test).unionAll(viral_test)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "8f7637e1-6d72-43db-b6f3-dd2d9ade3abc", "showTitle": false, "title": ""}
display(test_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "3019a468-3afe-4a00-a488-65ca4e142e66", "showTitle": false, "title": ""}
from pyspark.ml.classification import LogisticRegression
from pyspark.ml.classification import RandomForestClassifier
#from pyspark.ml.classification import GBTClassifier
from pyspark.ml.classification import DecisionTreeClassifier
from sklearn.model_selection import GridSearchCV
from pyspark.ml import Pipeline
from sparkdl import DeepImageFeaturizer
featurizer = DeepImageFeaturizer(inputCol="image", outputCol="features", modelName="InceptionV3")
lr = LogisticRegression(maxIter=20, regParam=0.05, elasticNetParam=0.3, labelCol="label")
rf = RandomForestClassifier(labelCol="label", featuresCol="features")
#gbt = GBTClassifier(labelCol="label", featuresCol="features", maxIter=10)
dt = DecisionTreeClassifier(labelCol="label", maxDepth=15)
p = Pipeline(stages=[featurizer, lr])
p_model = p.fit(train_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4ae2b9bf-c0f2-4477-9c26-9b414a52b5c9", "showTitle": false, "title": ""}
p_model.stages[1].write().overwrite().save('lr')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "28abcfe2-7452-4be7-bc0f-dad0d4043196", "showTitle": false, "title": ""}
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
tested_df = p_model.transform(test_df)
evaluator = MulticlassClassificationEvaluator(metricName="accuracy")
print("Test set accuracy = " + str(evaluator.evaluate(tested_df.select("prediction", "label"))))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "48ef04d3-81f0-4479-883b-6144bab1c149", "showTitle": false, "title": ""}
from pyspark.sql.types import DoubleType
from pyspark.sql.functions import expr
def _p1(v):
return float(v.array[1])
p1 = udf(_p1, DoubleType())
df = tested_df.withColumn("p_1", p1(tested_df.probability))
wrong_df = df.orderBy(expr("abs(p_1 - label)"), ascending=False)
display(wrong_df.select("image.origin", "p_1", "label").limit(10))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "8bd6c9ab-e7e9-4de4-bb4a-4fc2696d39c1", "showTitle": false, "title": ""}
import matplotlib.pyplot as plt
import numpy as np
import itertools
def plot_confusion_matrix(cm, classes,normalize=True,title='Confusion matrix',cmap=plt.cm.GnBu):
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),horizontalalignment="center", color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c82dc19f-219b-4f20-9f49-7d34b86e2cda", "showTitle": false, "title": ""}
from sklearn.metrics import confusion_matrix
y_true = tested_df.select("label")
y_true = y_true.toPandas()
y_pred = tested_df.select("prediction")
y_pred = y_pred.toPandas()
cnf_matrix = confusion_matrix(y_true, y_pred,labels=range(3))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "a4b88c3d-dbe0-40b0-8175-de07d499b726", "showTitle": false, "title": ""}
display(plot_confusion_matrix(cnf_matrix, classes=["normal","viral","covid"]))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "bc3e1787-79ed-4fb9-8a02-f80b8032f790", "showTitle": false, "title": ""}
from sklearn.metrics import classification_report
target_names = ["Class {}".format(i) for i in ["normal","viral","covid"]]
print(classification_report(y_true, y_pred, target_names = target_names))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0f3e548b-d7f6-4dbe-9dc9-ecf24eee2056", "showTitle": false, "title": ""}
|
pySpark/Multiclass classification_Alt_Last_Layer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_pytorch_p36
# language: python
# name: conda_pytorch_p36
# ---
# # Predicting Boston Housing Prices
#
# ## Using XGBoost in SageMaker (Batch Transform)
#
# _Deep Learning Nanodegree Program | Deployment_
#
# ---
#
# As an introduction to using SageMaker's Low Level Python API we will look at a relatively simple problem. Namely, we will use the [Boston Housing Dataset](https://www.cs.toronto.edu/~delve/data/boston/bostonDetail.html) to predict the median value of a home in the area of Boston Mass.
#
# The documentation reference for the API used in this notebook is the [SageMaker Developer's Guide](https://docs.aws.amazon.com/sagemaker/latest/dg/)
#
# ## General Outline
#
# Typically, when using a notebook instance with SageMaker, you will proceed through the following steps. Of course, not every step will need to be done with each project. Also, there is quite a lot of room for variation in many of the steps, as you will see throughout these lessons.
#
# 1. Download or otherwise retrieve the data.
# 2. Process / Prepare the data.
# 3. Upload the processed data to S3.
# 4. Train a chosen model.
# 5. Test the trained model (typically using a batch transform job).
# 6. Deploy the trained model.
# 7. Use the deployed model.
#
# In this notebook we will only be covering steps 1 through 5 as we just want to get a feel for using SageMaker. In later notebooks we will talk about deploying a trained model in much more detail.
# Make sure that we use SageMaker 1.x
# !pip install sagemaker==1.72.0
# ## Step 0: Setting up the notebook
#
# We begin by setting up all of the necessary bits required to run our notebook. To start that means loading all of the Python modules we will need.
# +
# %matplotlib inline
import os
import time
from time import gmtime, strftime
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.datasets import load_boston
import sklearn.model_selection
# -
# In addition to the modules above, we need to import the various bits of SageMaker that we will be using.
# +
import sagemaker
from sagemaker import get_execution_role
from sagemaker.amazon.amazon_estimator import get_image_uri
# This is an object that represents the SageMaker session that we are currently operating in. This
# object contains some useful information that we will need to access later such as our region.
session = sagemaker.Session()
# This is an object that represents the IAM role that we are currently assigned. When we construct
# and launch the training job later we will need to tell it what IAM role it should have. Since our
# use case is relatively simple we will simply assign the training job the role we currently have.
role = get_execution_role()
# -
# ## Step 1: Downloading the data
#
# Fortunately, this dataset can be retrieved using sklearn and so this step is relatively straightforward.
boston = load_boston()
# ## Step 2: Preparing and splitting the data
#
# Given that this is clean tabular data, we don't need to do any processing. However, we do need to split the rows in the dataset up into train, test and validation sets.
# +
# First we package up the input data and the target variable (the median value) as pandas dataframes. This
# will make saving the data to a file a little easier later on.
X_bos_pd = pd.DataFrame(boston.data, columns=boston.feature_names)
Y_bos_pd = pd.DataFrame(boston.target)
# We split the dataset into 2/3 training and 1/3 testing sets.
X_train, X_test, Y_train, Y_test = sklearn.model_selection.train_test_split(X_bos_pd, Y_bos_pd, test_size=0.33)
# Then we split the training set further into 2/3 training and 1/3 validation sets.
X_train, X_val, Y_train, Y_val = sklearn.model_selection.train_test_split(X_train, Y_train, test_size=0.33)
# -
# ## Step 3: Uploading the data files to S3
#
# When a training job is constructed using SageMaker, a container is executed which performs the training operation. This container is given access to data that is stored in S3. This means that we need to upload the data we want to use for training to S3. In addition, when we perform a batch transform job, SageMaker expects the input data to be stored on S3. We can use the SageMaker API to do this and hide some of the details.
#
# ### Save the data locally
#
# First we need to create the test, train and validation csv files which we will then upload to S3.
# This is our local data directory. We need to make sure that it exists.
data_dir = '../data/boston'
if not os.path.exists(data_dir):
os.makedirs(data_dir)
# +
# We use pandas to save our test, train and validation data to csv files. Note that we make sure not to include header
# information or an index as this is required by the built in algorithms provided by Amazon. Also, for the train and
# validation data, it is assumed that the first entry in each row is the target variable.
X_test.to_csv(os.path.join(data_dir, 'test.csv'), header=False, index=False)
pd.concat([Y_val, X_val], axis=1).to_csv(os.path.join(data_dir, 'validation.csv'), header=False, index=False)
pd.concat([Y_train, X_train], axis=1).to_csv(os.path.join(data_dir, 'train.csv'), header=False, index=False)
# -
# ### Upload to S3
#
# Since we are currently running inside of a SageMaker session, we can use the object which represents this session to upload our data to the 'default' S3 bucket. Note that it is good practice to provide a custom prefix (essentially an S3 folder) to make sure that you don't accidentally interfere with data uploaded from some other notebook or project.
# +
prefix = 'boston-xgboost-LL'
test_location = session.upload_data(os.path.join(data_dir, 'test.csv'), key_prefix=prefix)
val_location = session.upload_data(os.path.join(data_dir, 'validation.csv'), key_prefix=prefix)
train_location = session.upload_data(os.path.join(data_dir, 'train.csv'), key_prefix=prefix)
# -
# ## Step 4: Train and construct the XGBoost model
#
# Now that we have the training and validation data uploaded to S3, we can construct a training job for our XGBoost model and build the model itself.
#
# ### Set up the training job
#
# First, we will set up and execute a training job for our model. To do this we need to specify some information that SageMaker will use to set up and properly execute the computation. For additional documentation on constructing a training job, see the [CreateTrainingJob API](https://docs.aws.amazon.com/sagemaker/latest/dg/API_CreateTrainingJob.html) reference.
# +
# We will need to know the name of the container that we want to use for training. SageMaker provides
# a nice utility method to construct this for us.
container = get_image_uri(session.boto_region_name, 'xgboost')
# We now specify the parameters we wish to use for our training job
training_params = {}
# We need to specify the permissions that this training job will have. For our purposes we can use
# the same permissions that our current SageMaker session has.
training_params['RoleArn'] = role
# Here we describe the algorithm we wish to use. The most important part is the container which
# contains the training code.
training_params['AlgorithmSpecification'] = {
"TrainingImage": container,
"TrainingInputMode": "File"
}
# We also need to say where we would like the resulting model artifacts stored.
training_params['OutputDataConfig'] = {
"S3OutputPath": "s3://" + session.default_bucket() + "/" + prefix + "/output"
}
# We also need to set some parameters for the training job itself. Namely we need to describe what sort of
# compute instance we wish to use along with a stopping condition to handle the case that there is
# some sort of error and the training script doesn't terminate.
training_params['ResourceConfig'] = {
"InstanceCount": 1,
"InstanceType": "ml.m4.xlarge",
"VolumeSizeInGB": 5
}
training_params['StoppingCondition'] = {
"MaxRuntimeInSeconds": 86400
}
# Next we set the algorithm specific hyperparameters. You may wish to change these to see what effect
# there is on the resulting model.
training_params['HyperParameters'] = {
"max_depth": "5",
"eta": "0.2",
"gamma": "4",
"min_child_weight": "6",
"subsample": "0.8",
"objective": "reg:linear",
"early_stopping_rounds": "10",
"num_round": "200"
}
# Now we need to tell SageMaker where the data should be retrieved from.
training_params['InputDataConfig'] = [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": train_location,
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "csv",
"CompressionType": "None"
},
{
"ChannelName": "validation",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": val_location,
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "csv",
"CompressionType": "None"
}
]
# -
# ### Execute the training job
#
# Now that we've built the dictionary object containing the training job parameters, we can ask SageMaker to execute the job.
# +
# First we need to choose a training job name. This is useful for if we want to recall information about our
# training job at a later date. Note that SageMaker requires a training job name and that the name needs to
# be unique, which we accomplish by appending the current timestamp.
training_job_name = "boston-xgboost-" + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
training_params['TrainingJobName'] = training_job_name
# And now we ask SageMaker to create (and execute) the training job
training_job = session.sagemaker_client.create_training_job(**training_params)
# -
# The training job has now been created by SageMaker and is currently running. Since we need the output of the training job, we may wish to wait until it has finished. We can do so by asking SageMaker to output the logs generated by the training job and continue doing so until the training job terminates.
session.logs_for_job(training_job_name, wait=True)
# ### Build the model
#
# Now that the training job has completed, we have some model artifacts which we can use to build a model. Note that here we mean SageMaker's definition of a model, which is a collection of information about a specific algorithm along with the artifacts which result from a training job.
# +
# We begin by asking SageMaker to describe for us the results of the training job. The data structure
# returned contains a lot more information than we currently need, try checking it out yourself in
# more detail.
training_job_info = session.sagemaker_client.describe_training_job(TrainingJobName=training_job_name)
model_artifacts = training_job_info['ModelArtifacts']['S3ModelArtifacts']
# +
# Just like when we created a training job, the model name must be unique
model_name = training_job_name + "-model"
# We also need to tell SageMaker which container should be used for inference and where it should
# retrieve the model artifacts from. In our case, the xgboost container that we used for training
# can also be used for inference.
primary_container = {
"Image": container,
"ModelDataUrl": model_artifacts
}
# And lastly we construct the SageMaker model
model_info = session.sagemaker_client.create_model(
ModelName = model_name,
ExecutionRoleArn = role,
PrimaryContainer = primary_container)
# -
# ## Step 5: Testing the model
#
# Now that we have fit our model to the training data, using the validation data to avoid overfitting, we can test our model. To do this we will make use of SageMaker's Batch Transform functionality. In other words, we need to set up and execute a batch transform job, similar to the way that we constructed the training job earlier.
#
# ### Set up the batch transform job
#
# Just like when we were training our model, we first need to provide some information in the form of a data structure that describes the batch transform job which we wish to execute.
#
# We will only be using some of the options available here but to see some of the additional options please see the SageMaker documentation for [creating a batch transform job](https://docs.aws.amazon.com/sagemaker/latest/dg/API_CreateTransformJob.html).
# +
# Just like in each of the previous steps, we need to make sure to name our job and the name should be unique.
transform_job_name = 'boston-xgboost-batch-transform-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
# Now we construct the data structure which will describe the batch transform job.
transform_request = \
{
"TransformJobName": transform_job_name,
# This is the name of the model that we created earlier.
"ModelName": model_name,
# This describes how many compute instances should be used at once. If you happen to be doing a very large
# batch transform job it may be worth running multiple compute instances at once.
"MaxConcurrentTransforms": 1,
# This says how big each individual request sent to the model should be, at most. One of the things that
# SageMaker does in the background is to split our data up into chunks so that each chunks stays under
# this size limit.
"MaxPayloadInMB": 6,
# Sometimes we may want to send only a single sample to our endpoint at a time, however in this case each of
# the chunks that we send should contain multiple samples of our input data.
"BatchStrategy": "MultiRecord",
# This next object describes where the output data should be stored. Some of the more advanced options which
# we don't cover here also describe how SageMaker should collect output from various batches.
"TransformOutput": {
"S3OutputPath": "s3://{}/{}/batch-bransform/".format(session.default_bucket(),prefix)
},
# Here we describe our input data. Of course, we need to tell SageMaker where on S3 our input data is stored, in
# addition we need to detail the characteristics of our input data. In particular, since SageMaker may need to
# split our data up into chunks, it needs to know how the individual samples in our data file appear. In our
# case each line is its own sample and so we set the split type to 'line'. We also need to tell SageMaker what
# type of data is being sent, in this case csv, so that it can properly serialize the data.
"TransformInput": {
"ContentType": "text/csv",
"SplitType": "Line",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": test_location,
}
}
},
# And lastly we tell SageMaker what sort of compute instance we would like it to use.
"TransformResources": {
"InstanceType": "ml.m4.xlarge",
"InstanceCount": 1
}
}
# -
# ### Execute the batch transform job
#
# Now that we have created the request data structure, it is time to ask SageMaker to set up and run our batch transform job. Just like in the previous steps, SageMaker performs these tasks in the background so that if we want to wait for the transform job to terminate (and ensure the job is progressing) we can ask SageMaker to wait of the transform job to complete.
transform_response = session.sagemaker_client.create_transform_job(**transform_request)
transform_desc = session.wait_for_transform_job(transform_job_name)
# ### Analyze the results
#
# Now that the transform job has completed, the results are stored on S3 as we requested. Since we'd like to do a bit of analysis in the notebook we can use some notebook magic to copy the resulting output from S3 and save it locally.
transform_output = "s3://{}/{}/batch-bransform/".format(session.default_bucket(),prefix)
# !aws s3 cp --recursive $transform_output $data_dir
# To see how well our model works we can create a simple scatter plot between the predicted and actual values. If the model was completely accurate the resulting scatter plot would look like the line $x=y$. As we can see, our model seems to have done okay but there is room for improvement.
Y_pred = pd.read_csv(os.path.join(data_dir, 'test.csv.out'), header=None)
plt.scatter(Y_test, Y_pred)
plt.xlabel("Median Price")
plt.ylabel("Predicted Price")
plt.title("Median Price vs Predicted Price")
# ## Optional: Clean up
#
# The default notebook instance on SageMaker doesn't have a lot of excess disk space available. As you continue to complete and execute notebooks you will eventually fill up this disk space, leading to errors which can be difficult to diagnose. Once you are completely finished using a notebook it is a good idea to remove the files that you created along the way. Of course, you can do this from the terminal or from the notebook hub if you would like. The cell below contains some commands to clean up the created files from within the notebook.
# +
# First we will remove all of the files contained in the data_dir directory
# !rm $data_dir/*
# And then we delete the directory itself
# !rmdir $data_dir
# -
|
Tutorials/Boston Housing - XGBoost (Batch Transform) - Low Level.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" deletable=true editable=true id="mHF9VCProKJN"
# # AI Explanations: Explaining a tabular data model
#
# + [markdown] colab_type="text" deletable=true editable=true id="hZzRVxNtH-zG"
# ## Overview
#
# In this tutorial we will perform the following steps:
#
# 1. Build and train a Keras model.
# 1. Export the Keras model as a TF 1 SavedModel and deploy the model on Cloud AI Platform.
# 1. Compute explainations for our model's predictions using Explainable AI on Cloud AI Platform.
# + [markdown] colab_type="text" deletable=true editable=true id="iN69d4D9Flrh"
# ### Dataset
#
# The dataset used for this tutorial was created from a BigQuery Public Dataset: [London Bike Dataset](https://data.london.gov.uk/).
# + [markdown] colab_type="text" deletable=true editable=true id="Su2qu-4CW-YH"
# ### Objective
#
# The goal is to train a model using the Keras Sequential API that predicts the duration of a bike ride given the weekday, weather conditions, and start and stop station of the bike.
#
# This tutorial focuses more on deploying the model to AI Explanations than on the design of the model itself. We will be using preprocessed data for this lab.
#
# + [markdown] colab_type="text" deletable=true editable=true id="TSy-f05IO4LB"
# ### Setup
# + cellView="both" colab={} colab_type="code" deletable=true editable=true id="4qxwBA4RM9Lu"
import os
PROJECT_ID = "" # TODO: your PROJECT_ID here.
os.environ["PROJECT_ID"] = PROJECT_ID
# + colab={} colab_type="code" deletable=true editable=true id="bTxmbDg1I0x1"
BUCKET_NAME = "" # TODO: your BUCKET_NAME here.
REGION = "us-central1"
os.environ[
"BUCKET_NAME"
] = PROJECT_ID # Replace your BUCKET_NAME, if needed. You can leave it as is!
os.environ["REGION"] = REGION
# + [markdown] colab_type="text" deletable=true editable=true id="fsmCk2dwJnLZ"
# Run the following cell to create your Cloud Storage bucket if it does not already exist.
# + colab={} colab_type="code" deletable=true editable=true id="160PRO3aJqLD" language="bash"
# exists=$(gsutil ls -d | grep -w gs://${BUCKET_NAME}/)
#
# if [ -n "$exists" ]; then
# echo -e "Bucket gs://${BUCKET_NAME} already exists."
#
# else
# echo "Creating a new GCS bucket."
# gsutil mb -l ${REGION} gs://${BUCKET_NAME}
# echo -e "\nHere are your current buckets:"
# gsutil ls
# fi
# -
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, we create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
# +
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] colab_type="text" id="PyxoF-iqqD1t"
# ### Import libraries
#
# Import the libraries for this tutorial. This tutorial has been tested with **TensorFlow versions 2.3**.
# + colab={} colab_type="code" id="MEDlLSWK15UL"
import tensorflow as tf
import pandas as pd
import explainable_ai_sdk
# + [markdown] colab_type="text" deletable=true editable=true id="aRVMEU2Qshm4"
# ## Download and preprocess the data
#
# In this section you'll download the data to train your model from a public GCS bucket. The original data is from the BigQuery datasets linked above. For your convenience, we've joined the London bike and NOAA weather tables, done some preprocessing, and provided a subset of that dataset here.
#
# + colab={} colab_type="code" id="v7HLNsvekxvz"
# Copy the data to your notebook instance
# ! gsutil cp 'gs://explanations_sample_data/bike-data.csv' ./
# + [markdown] colab_type="text" deletable=true editable=true id="8zr6lj66UlMn"
# ### Read the data with Pandas
#
# You'll use Pandas to read the data into a `DataFrame` and then do some additional pre-processing.
# + colab={} colab_type="code" deletable=true editable=true id="Icz22E69smnD"
data = pd.read_csv("bike-data.csv")
# Shuffle the data
data = data.sample(frac=1, random_state=2)
# Drop rows with null values
data = data[data["wdsp"] != 999.9]
data = data[data["dewp"] != 9999.9]
# Rename some columns for readability
data = data.rename(columns={"day_of_week": "weekday"})
data = data.rename(columns={"max": "max_temp"})
data = data.rename(columns={"dewp": "dew_point"})
# Drop columns you won't use to train this model
data = data.drop(
columns=[
"start_station_name",
"end_station_name",
"bike_id",
"snow_ice_pellets",
]
)
# Convert trip duration from seconds to minutes so it's easier to understand
data["duration"] = data["duration"].apply(lambda x: float(x / 60))
# + colab={} colab_type="code" id="vxZryg4xmdy0"
# Preview the first 5 rows of training data
data.head()
# -
# Next, you will separate the data into features ('data') and labels ('labels').
# Save duration to its own DataFrame and remove it from the original DataFrame
labels = data["duration"]
data = data.drop(columns=["duration"])
# ### Split data into train and test sets
#
# You'll split your data into train and test sets using an 80 / 20 train / test split.
# +
# Use 80/20 train/test split
train_size = int(len(data) * 0.8)
print("Train size: %d" % train_size)
print("Test size: %d" % (len(data) - train_size))
# Split your data into train and test sets
train_data = data[:train_size]
train_labels = labels[:train_size]
test_data = data[train_size:]
test_labels = labels[train_size:]
# + [markdown] colab_type="text" id="kV_NEAQwwH0e"
# ## Build, train, and evaluate our model with Keras
#
# This section shows how to build, train, evaluate, and get local predictions from a model by using the Keras [Sequential API](https://www.tensorflow.org/guide/keras/sequential_model). The model will takes your 10 features as input and predict the trip duration in minutes.
# -
# **TODO: Build a simple keras sequential model with three dense layers for your structured data**
# + colab={} colab_type="code" id="HCQFzd_YdwLX"
# Build your model
model = #TODO: Keras sequential code goes here
# + colab={} colab_type="code" id="UvAcjSUcs_l7"
# Compile the model and see a summary
model.compile(loss="mean_squared_logarithmic_error", optimizer="adam")
model.summary()
# + [markdown] colab_type="text" id="GcOkuHPVwjiM"
# ### Create an input data pipeline with tf.data
#
# Per best practices, we will use `tf.Data` to create our input data pipeline. Our data is all in an in-memory dataframe, so we will use `tf.data.Dataset.from_tensor_slices` to create our pipeline.
# + colab={} colab_type="code" id="ZUu9wFklwmm6"
batch_size = 256
epochs = 3
input_train = tf.data.Dataset.from_tensor_slices(train_data)
output_train = tf.data.Dataset.from_tensor_slices(train_labels)
input_train = input_train.batch(batch_size).repeat()
output_train = output_train.batch(batch_size).repeat()
train_dataset = tf.data.Dataset.zip((input_train, output_train))
# + [markdown] colab_type="text" id="l98aRzfPwo5e"
# ### Train the model
#
# Now we train the model. We will specify a number of epochs which to train the model and tell the model how many steps to expect per epoch.
# + colab={} colab_type="code" id="h1x_8CR0wtRs"
# This will take about a minute to run
# To keep training time short, you're not using the full dataset
model.fit(
train_dataset, steps_per_epoch=train_size // batch_size, epochs=epochs
)
# -
# ### Evaluate the trained model locally
# Run evaluation
results = model.evaluate(test_data, test_labels)
print(results)
# + colab={} colab_type="code" id="bIh6uds2x2tr"
# Send test instances to model for prediction
predict = model.predict(test_data[:5])
# -
# Preview predictions on the first 5 examples from your test dataset
for i, val in enumerate(predict):
print("Predicted duration: {}".format(round(val[0])))
print("Actual duration: {} \n".format(test_labels.iloc[i]))
# + [markdown] colab_type="text" deletable=true editable=true id="gAO6-zv6osJ8"
# ## Export the model as a TF 2.x SavedModel
#
# When using TensorFlow 2.x, you export the model as a `SavedModel` and load it into Cloud Storage.
# + colab={} colab_type="code" id="fbvzBm1lji7b"
export_path = "gs://" + BUCKET_NAME + "/explanations/mymodel"
model.save(export_path)
print(export_path)
# + [markdown] colab_type="text" id="-f8elyM8KMNX"
# Use TensorFlow's `saved_model_cli` to inspect the model's SignatureDef. We'll use this information when we deploy our model to AI Explanations in the next section.
# + colab={} colab_type="code" id="yFg5r-7s1BKr"
# ! saved_model_cli show --dir $export_path --all
# + [markdown] colab_type="text" id="y270ZNinycoy"
# ## Deploy the model to AI Explanations
#
# In order to deploy the model to Explanations, you need to generate an `explanations_metadata.json` file and upload this to the Cloud Storage bucket with your SavedModel. Then you'll deploy the model using `gcloud`.
# + [markdown] colab_type="text" id="cUdUVjjGbvQy"
# ### Prepare explanation metadata
#
# In order to deploy this model to AI Explanations, you need to create an explanation_metadata.json file with information about your model inputs, outputs, and baseline. You can use the [Explainable AI SDK](https://pypi.org/project/explainable-ai-sdk/) to generate most of the fields.
#
# The value for `input_baselines` tells the explanations service what the baseline input should be for your model. Here you're using the median for all of your input features. That means the baseline prediction for this model will be the trip duration your model predicts for the median of each feature in your dataset.
#
# Since this model accepts a single numpy array with all numerical feature, you can optionally pass an `index_feature_mapping` list to AI Explanations to make the API response easier to parse. When you provide a list of feature names via this parameter, the service will return a key / value mapping of each feature with its corresponding attribution value.
# + colab={} colab_type="code" id="UolAW3lcVTGl"
# Print the names of your tensors
print("Model input tensor: ", model.input.name)
print("Model output tensor: ", model.output.name)
# + colab={} colab_type="code" id="qpZiW9Cq6IY4"
from explainable_ai_sdk.metadata.tf.v2 import SavedModelMetadataBuilder
builder = SavedModelMetadataBuilder(export_path)
builder.set_numeric_metadata(
model.input.name.split(":")[0],
input_baselines=[train_data.median().values.tolist()],
index_feature_mapping=train_data.columns.tolist(),
)
builder.save_metadata(export_path)
# + [markdown] colab_type="text" id="rT3iG5pDdrHi"
# Since this is a regression model (predicting a numerical value), the baseline prediction will be the same for every example we send to the model. If this were instead a classification model, each class would have a different baseline prediction.
# + [markdown] colab_type="text" id="J6MKKy6Xb2MT"
# ### Create the model
# +
import datetime
MODEL = "bike" + datetime.datetime.now().strftime("%d%m%Y%H%M%S")
# -
# **TODO: Create a model using the gcloud command. Enable logging and make sure to pass region to the command**
# Create the model if it doesn't exist yet (you only need to run this once)
! #TODO: gcloud command goes here
# ### Create the model version
#
# Creating the version will take ~5-10 minutes. Note that your first deploy could take longer.
# + colab={} colab_type="code" id="S2OaOycmb4o0"
# Each time you create a version the name should be unique
VERSION = "v1"
# -
# **TODO: Write the gcloud comman to create a new model version with explanations. You need to use gcloud beta on AI platform**
# + colab={} colab_type="code" id="0bwCxEr5b8BP"
# Create the version with gcloud
explain_method = "integrated-gradients"
! #TODO: Your gcloud command goes here
# -
# Make sure the model deployed correctly. State should be `READY` in the following log
# ! gcloud ai-platform versions describe $VERSION --model $MODEL --region $REGION
# + [markdown] colab_type="text" deletable=true editable=true id="JzevJps9IOcU"
# ## Get predictions and explanations
#
# Now that your model is deployed, you can use the AI Platform Prediction API to get feature attributions. You'll pass it a single test example here and see which features were most important in the model's prediction. Here you'll use the [Explainable AI SDK](https://pypi.org/project/explainable-ai-sdk/) to get your prediction and explanation. You can also use `gcloud`.
# + [markdown] colab_type="text" id="CJ-2ErWJDvcg"
# ### Format your explanation request
#
# To make your AI Explanations request, you need to create a JSON object with your test data for prediction.
# -
# Format data for prediction to your model
prediction_json = {
model.input.name.split(":")[0]: test_data.iloc[0].values.tolist()
}
# ### Send the explain request
#
# You can use the Explainable AI SDK to send explanation requests to your deployed model.
# + colab={} colab_type="code" id="D_PR2BcHD40-"
remote_ig_model = explainable_ai_sdk.load_model_from_ai_platform(
project=PROJECT_ID, model=MODEL, version=VERSION, region=REGION
)
ig_response = remote_ig_model.explain([prediction_json])
# + [markdown] colab_type="text" id="0nKR8RelNnkK"
# ### Understanding the explanations response
#
# First, let's look at the trip duration your model predicted and compare it to the actual value.
# + colab={} colab_type="code" id="825KoNgHR-tv"
attr = ig_response[0].get_attribution()
predicted = round(attr.example_score, 2)
print("Predicted duration: " + str(predicted) + " minutes")
print("Actual duration: " + str(test_labels.iloc[0]) + " minutes")
# -
# Next let's look at the feature attributions for this particular example. Positive attribution values mean a particular feature pushed your model prediction up by that amount, and vice versa for negative attribution values.
ig_response[0].visualize_attributions()
# +
# The above graph is missing because ig_response[0].get_attribution()
# does not fill `_values_dict` when the model is coming from AI Platform.
# below is a workaround, which redefines the Attribution with values_dict:
import numpy as np
import IPython
from explainable_ai_sdk.common import attribution
from xai_tabular_widget import TabularWidget
test_data_dict = dict(test_data.iloc[0])
for key, item in test_data_dict.items():
test_data_dict[key] = np.array([item], dtype=np.float32)
raw_attribution = ig_response[0].get_attribution()
attribution = attribution.Attribution(
output_name=raw_attribution.output_name,
baseline_score=raw_attribution.baseline_score,
example_score=raw_attribution.example_score,
values_dict=test_data_dict,
attrs_dict=raw_attribution.attrs_dict,
label_index=raw_attribution.label_index,
processed_attrs_dict=raw_attribution._get_attributions_dict(),
approx_error=raw_attribution.approx_error,
label_name=raw_attribution.label_name,
)
target_label_attr = attribution.to_json(include_input_values=True)
widget = TabularWidget()
def input_to_widget():
widget.load_data_from_json(target_label_attr)
widget.on_trait_change(input_to_widget, "ready")
IPython.display.display(widget)
# -
# ## Check your explanations and baselines
#
# To better make sense of the feature attributions you're getting, you should compare them with your model's baseline. In most cases, the sum of your attribution values + the baseline should be very close to your model's predicted value for each input. Also note that for regression models, the `baseline_score` returned from AI Explanations will be the same for each example sent to your model. For classification models, each class will have its own baseline.
#
# In this section you'll send 10 test examples to your model for prediction in order to compare the feature attributions with the baseline. Then you'll run each test example's attributions through two sanity checks in the `sanity_check_explanations` method.
# Prepare 10 test examples to your model for prediction
pred_batch = []
for i in range(10):
pred_batch.append(
{model.input.name.split(":")[0]: test_data.iloc[i].values.tolist()}
)
test_response = remote_ig_model.explain(pred_batch)
# In the function below you perform two sanity checks for models using Integrated Gradient (IG) explanations and one sanity check for models using Sampled Shapley.
def sanity_check_explanations(
example, mean_tgt_value=None, variance_tgt_value=None
):
passed_test = 0
total_test = 1
# `attributions` is a dict where keys are the feature names
# and values are the feature attributions for each feature
attr = example.get_attribution()
baseline_score = attr.baseline_score
# sum_with_baseline = np.sum(attribution_vals) + baseline_score
predicted_val = attr.example_score
# Sanity check 1
# The prediction at the input is equal to that at the baseline.
# Please use a different baseline. Some suggestions are: random input, training
# set mean.
if abs(predicted_val - baseline_score) <= 0.05:
print("Warning: example score and baseline score are too close.")
print("You might not get attributions.")
else:
passed_test += 1
# Sanity check 2 (only for models using Integrated Gradient explanations)
# Ideally, the sum of the integrated gradients must be equal to the difference
# in the prediction probability at the input and baseline. Any discrepency in
# these two values is due to the errors in approximating the integral.
if explain_method == "integrated-gradients":
total_test += 1
want_integral = predicted_val - baseline_score
got_integral = sum(attr.post_processed_attributions.values())
if abs(want_integral - got_integral) / abs(want_integral) > 0.05:
print("Warning: Integral approximation error exceeds 5%.")
print(
"Please try increasing the number of integrated gradient steps."
)
else:
passed_test += 1
print(passed_test, " out of ", total_test, " sanity checks passed.")
for response in test_response:
sanity_check_explanations(response)
# ## Understanding AI Explanations with the What-If Tool
#
# In this section you'll use the [What-If Tool](https://pair-code.github.io/what-if-tool/) to better understand how your model is making predictions. See the cell below the What-if Tool for visualization ideas.
# The What-If-Tool expects data with keys for each feature name, but your model expects a flat list. The functions below convert data to the format required by the What-If Tool.
# +
# This is the number of data points you'll send to the What-if Tool
WHAT_IF_TOOL_SIZE = 500
from witwidget.notebook.visualization import WitWidget, WitConfigBuilder
def create_list(ex_dict):
new_list = []
for i in feature_names:
new_list.append(ex_dict[i])
return new_list
def example_dict_to_input(example_dict):
return {"dense_input": create_list(example_dict)}
from collections import OrderedDict
wit_data = test_data.iloc[:WHAT_IF_TOOL_SIZE].copy()
wit_data["duration"] = test_labels[:WHAT_IF_TOOL_SIZE]
wit_data_dict = wit_data.to_dict(orient="records", into=OrderedDict)
# +
config_builder = (
WitConfigBuilder(wit_data_dict)
.set_ai_platform_model(
PROJECT_ID, MODEL, VERSION, adjust_example=example_dict_to_input
)
.set_target_feature("duration")
.set_model_type("regression")
)
WitWidget(config_builder)
# -
# ### What-If Tool visualization ideas
#
# On the x-axis, you'll see the predicted trip duration for the test inputs you passed to the What-If Tool. Each circle represents one of your test examples. If you click on a circle, you'll be able to see the feature values for that example along with the attribution values for each feature.
#
# * You can edit individual feature values and re-run prediction directly within the What-If Tool. Try changing `distance`, click **Run inference** and see how that affects the model's prediction
# * You can sort features for an individual example by their attribution value, try changing the sort from the attributions dropdown
# * The What-If Tool also lets you create custom visualizations. You can do this by changing the values in the dropdown menus above the scatter plot visualization. For example, you can sort data points by inference error, or by their similarity to a single datapoint.
# ## Cleaning up
# +
# Delete model version resource
# ! gcloud ai-platform versions delete $VERSION --quiet --model $MODEL --region $REGION
# Delete model resource
# ! gcloud ai-platform models delete $MODEL --region $REGION --quiet
# -
# ## What's next?
#
# To learn more about AI Explanations or the What-if Tool, check out the resources here.
#
# * [AI Explanations documentation](cloud.google.com/ml-engine/docs/ai-explanations)
# * [Documentation for using the What-if Tool with Cloud AI Platform models ](https://cloud.google.com/ml-engine/docs/using-what-if-tool)
# * [What-If Tool documentation and demos](https://pair-code.github.io/what-if-tool/)
# * [Integrated gradients paper](https://arxiv.org/abs/1703.01365)
|
notebooks/ml_fairness_explainability/explainable_ai/labs/xai_structured_caip.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3 64-bit
# name: python37364bit0d1bc8e257174fbe9ca52dee52151f48
# ---
# +
import os
import glob
import pandas as pd
from graphviz import Digraph, Graph
import json
# +
currDIR = os.getcwd() + "\\\\"
def getShape(gender):
if gender == "M":
return "rect"
else:
return "circle"
fileNameList = [f for f in glob.glob("data/*.csv")]
print(fileNameList)
familyNameDict = {}
for name in fileNameList:
familyNameDict[(name.replace("data\\", "").replace(".csv", ""))] = pd.read_csv(currDIR + name, encoding="UTF-8")
# familiesList.append(pd.read_csv(currDIR + name, encoding="UTF-8"))
# +
# #df = pd.read_csv("data/F1.csv")
#print(df)
selfNode = familyNameDict["F1"]
print(selfNode)
familyName = fileNameList[0].replace("data\\", "").replace(".csv", "")
# fam = Graph(comment=familyName, format="png")
# fam.node("{}".format(str(familyName + selfNode)), selfNode.Relationship, shape=getShape(selfNode[1]))
# fam.render("fam.gv")
# -
|
test3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:cta] *
# language: python
# name: conda-env-cta-py
# ---
# # Intensity Resolution Definition
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
def calculate_requirement_curve(pe, nsb, window_width, electronic_noise, miscal, enf):
"""
Equation for calculating the Goal and Requirement curves, as used in the CTA requirement
Parameters
----------
pe : ndarray
Number of photoelectrons (p.e.)
nsb : float
NSB rate (MHz)
window_width : float
Integration window width (ns)
electronic_noise : float
Charge Stddev due to integrated electronic noise (p.e.)
miscal : float
Multiplicative errors of the gain.
enf : float
Excess noise factor.
"""
var_noise = nsb * window_width + electronic_noise**2
var_enf = (1 + enf)**2 * pe
var_miscal = (miscal * pe)**2
sigma_q = np.sqrt(var_noise + var_enf + var_miscal)
return sigma_q / pe
def calculate_requirement_nominal_nsb(pe):
return calculate_requirement_curve(
pe,
nsb=0.125,
window_width=15,
electronic_noise=0.87,
miscal=0.1,
enf=0.2,
)
def calculate_requirement_high_nsb(pe):
return calculate_requirement_curve(
pe,
nsb=1.25,
window_width=15,
electronic_noise=0.87,
miscal=0.1,
enf=0.2,
)
# +
x, y = np.loadtxt("IntensityRes.txt", unpack=True)
plt.plot(x, y)
plt.xscale("log")
plt.yscale("log")
ph = x
requirement_pde = 0.25
pe = ph * requirement_pde
req_nominal_nsb = calculate_requirement_nominal_nsb(pe)
plt.plot(ph, req_nominal_nsb)
np.testing.assert_allclose(y, req_nominal_nsb, rtol=1e-5)
# +
x, y = np.loadtxt("IntensityResHighNSB.txt", unpack=True)
plt.plot(x, y)
plt.xscale("log")
plt.yscale("log")
ph = x
requirement_pde = 0.25
pe = ph * requirement_pde
req_high_nsb = calculate_requirement_high_nsb(pe)
plt.plot(ph, req_high_nsb)
np.testing.assert_allclose(y, req_high_nsb, rtol=1e-5)
# -
# The underlying formula for the requirement curves are demonstrated here. The formula used here defines the Intensity Resolution at an intensity $I$ as the Charge Resolution at a charge of $I \times \epsilon_{PDE}$, where a nominal PDE of $\epsilon_{PDE} = 0.25$ is used.
#
# There are two equivalent formula which therefore describe the Fractional Intensity Resolution:
#
# $$\frac{\sigma_{I_T}}{I_T} = \frac{1}{I_T} \sqrt{\frac{\sum_{i=0}^N (I_{M_i} - I_T)^2}{N}}$$
#
# Where $I_{M_i}$ are individual measurements of the intensity in photons of a true intensity $I_T$, and
#
# $$\frac{\sigma_{I_T=\frac{Q_T}{\epsilon_{PDE}}}}{Q_T} = \frac{1}{Q_T} \sqrt{\frac{\sum_{i=0}^N (Q_{M_i} - Q_T)^2}{N}}$$
#
# Where $Q_{M_i}$ are individual measurements of the charge (p.e.) of a true charge $Q_T$. The equivalence is demonstrated below difference between the two definitions is explored below:
# +
amplitude_pe = 50
charge_pe = np.random.normal(amplitude_pe, 10, 100000)
res_pe = charge_pe.std()
amplitude_ph = amplitude_pe / requirement_pde
charge_ph = charge_pe / requirement_pde
res_ph = charge_ph.std()
print(f"Charge Resolution at Q = {amplitude_pe} p.e. is {res_pe/amplitude_pe:.2f}")
print(f"Intensity Resolution at I={amplitude_ph} photons using Equation 1 is {res_ph / amplitude_ph:.2f}")
print(f"Intensity Resolution at I={amplitude_ph} photons using Equation 2 is {res_pe / amplitude_pe:.2f}")
# -
|
d210127_cr_calculators/resolution_definition.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## <center> Real Time Stock Price Prediction</center>
# Stock Market Prediction, the aim is to predict the future value of the financial stocks of a company. The recent trend in stock market prediction technologies is the use of machine learning which makes predictions based on the values of current stock market indices by training on their previous values.
# +
#import the libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import datetime as dt
import pandas_datareader.data as web
from matplotlib import style
import statsmodels.api as sm
from statsmodels.tsa.seasonal import seasonal_decompose
from statsmodels.tsa.arima_model import ARIMA
from sklearn.metrics import mean_squared_error
from bokeh.plotting import figure, output_notebook, show
# -
df=web.DataReader('AAPL',data_source="yahoo",start="3-1-15",end="25-10-19")
df.reset_index()
df.isnull().sum()
df.info()
df.describe()
df.rename(columns={"Adj Close":"Adj_close"},inplace=True)
plt.figure(figsize=(16,8))
plt.subplot(1,2,1);
plt.plot(df.Open.values, color='red', label='Open')
plt.plot(df.Close.values, color='green', label='Close')
plt.plot(df.Low.values, color='blue', label='low')
plt.plot(df.High.values, color='black', label='high')
plt.plot(df.Adj_close.values,color='yellow',label="adj_close")
plt.title('stock price')
plt.xlabel('time [days]')
plt.ylabel('volume')
plt.legend(loc='best')
plt.subplot(1,2,2);
plt.plot(df.Volume.values, color='black', label='volume')
plt.title('stock volume')
plt.xlabel('time [days]')
plt.ylabel('volume')
# +
sm.tsa.seasonal_decompose(df.Close.values,freq=30).plot()
result = sm.tsa.stattools.adfuller(df.Close)
plt.show()
#decomposition = seasonal_decompose(df.Close.values, freq=30)
# -
df.shape
train = df[:900]
test = df[900:]
# +
train_ar = train['Close'].values
test_ar = test['Close'].values
# https://machinelearningmastery.com/arima-for-time-series-forecasting-with-python/
history = [x for x in train_ar]
print(type(history))
predictions = list()
for t in range(len(test_ar)):
model = ARIMA(history, order=(5,1,0))
model_fit = model.fit(disp=0)
output = model_fit.forecast()
yhat = output[0]
predictions.append(yhat)
obs = test_ar[t]
history.append(obs)
error = mean_squared_error(test_ar, predictions)
print('Testing Mean Squared Error: %.3f' % error)
# -
plt.figure(figsize=(12,5))
plt.plot(df['Close'], 'green', color='blue', label='Training Data')
plt.plot(test.index, predictions, color='green', marker='o', linestyle='dashed',
label='Predicted Price')
plt.plot(test.index, test['Close'], color='red', label='Actual Price')
plt.title('Close Prices Prediction')
plt.xlabel('Dates')
plt.ylabel('Prices')
plt.legend()
plt.figure(figsize=(12,7))
plt.plot(test.index, predictions, color='green', marker='.', linestyle='dashed',
label='Predicted Price')
plt.plot(test.index, test['Close'], color='red', label='Actual Price')
plt.title('Close Prices Prediction')
plt.xlabel('Dates')
plt.ylabel('Prices')
plt.legend()
plt.figure(figsize=(15, 7))
plt.plot(df.index,df["Close"])
# +
output_notebook()
# create figure
p = figure(plot_width = 400, plot_height = 400)
# -
p.line(df.index,df.Close, line_width = 2, color = "green")
# +
show(p)
# -
# #### Conclusion:
# One technique have been utilized in this paper: ARIMA, on the Yahoo finance dataset. the technique have shown an improvement in the accuracy of predictions, thereby yielding positive results. Use of recently introduced machine learning technique in the prediction of stocks have yielded promising results and thereby marked the use of them in profitable exchange schemes. It has led to the conclusion that it is possible to predict stock market with more accuracy and efficiency using machine learning technique.
|
final stock price prediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from multiprocessing import Pool
import time
import sys
# +
dims = 2
dx = 1
minx = 0
maxx = 100
def isvalid(x):
R = 15**2
valid = np.sum((np.array(x)-50)**2)<R
return valid
def inspace(x):
for xd in x:
if xd<minx or xd>maxx:
return False
return True
# -
def explore_dim(dim,dx,valid_solns,computed_solns):
valid_solns0 = set(valid_solns)
for v in valid_solns0:
xm = list(v)
xm[dim] -= dx
xm = tuple(xm)
xp = list(v)
xp[dim] += dx
xp = tuple(xp)
xes = [xm,xp]
for x in xes:
if inspace(x):
if x not in computed_solns:
computed_solns.add(x)
if isvalid(x):
valid_solns.add(x)
explore_dim(dim,dx,valid_solns,computed_solns)
def explore_both(dx,valid_solns,computed_solns):
valid_solns1 = set(valid_solns)
computed_solns1 = set(computed_solns)
valid_solns2 = set(valid_solns)
computed_solns2 = set(computed_solns)
explore_dim(0,dx,valid_solns1,computed_solns1)
explore_dim(1,dx,valid_solns2,computed_solns2)
valid_solns.update(valid_solns1,valid_solns2)
computed_solns.update(computed_solns1,computed_solns2)
# +
valid_solns = {(50,50)}
computed_solns = set(valid_solns)
dx = 4
N = 8
for i in range(N):
explore_both(dx,valid_solns,computed_solns)
for x in valid_solns:
plt.plot(x[0],x[1],'.',color=plt.cm.rainbow(i/N),zorder=1/(i+1))
plt.plot(x[0],x[1],'.',color=plt.cm.rainbow(i/N),zorder=1/(i+1),label='iter={}'.format(i+1))
plt.xlim(minx,maxx)
plt.ylim(minx,maxx)
plt.legend()
# -
# # Multiprocessing
from multiprocessing import Process, Queue
# +
def explore_pos(x,dim,dx,valid_solns,computed_solns):
xp = list(x)
xp[dim] += dx
xp = tuple(xp)
if not inspace(xp):
return
if xp in computed_solns:
return
computed_solns.add(xp)
if not isvalid(xp):
return
valid_solns.add(xp)
explore_pos(xp,dim,dx,valid_solns,computed_solns)
def explore_neg(x,dim,dx,valid_solns,computed_solns):
xm = list(x)
xm[dim] -= dx
xm = tuple(xm)
if not inspace(xm):
return
if xm in computed_solns:
return
computed_solns.add(xm)
if not isvalid(xm):
return
valid_solns.add(xm)
explore_neg(xm,dim,dx,valid_solns,computed_solns)
def explore_dim(dim,dx,valid_solns,computed_solns):
valid_solns0 = set(valid_solns)
for v in valid_solns0:
explore_pos(v,dim,dx,valid_solns,computed_solns)
for v in valid_solns0:
explore_neg(v,dim,dx,valid_solns,computed_solns)
def multi_explore(dim,dx,valid_solns,computed_solns, q):
explore_dim(dim,dx,valid_solns,computed_solns)
q.put([valid_solns,computed_solns])
def run_parallel(dims, dxs, valid_solns, computed_solns):
qs = []
ps = []
for dim in dims:
qs.append(Queue())
ps.append(Process(target=multi_explore, args=(dim,dxs[dim],valid_solns,computed_solns,qs[dim])))
ps[dim].start()
for dim in dims:
out = qs[dim].get() # prints "[42, None, 'hello']"
valid_solns.update(out[0])
computed_solns.update(out[1])
ps[dim].join()
def iter_parallel(dims,dxs,valid_solns,computed_solns, Nmax):
N = 0
done = False
while(not done and N<Nmax):
N +=1
Ncomp = len(computed_solns)
run_parallel(dims,dxs,valid_solns,computed_solns)
if Ncomp == len(computed_solns):
done=True
print('done after {} iters'.format(N))
print(len(valid_solns),len(computed_solns))
return valid_solns,computed_solns
# +
dims = 2
dx = 1
minx = 0
maxx = 100
def isvalid(x):
R = 2**2
valid = np.sum((np.array(x)-50)**2)<R
return valid
def inspace(x):
for xd in x:
if xd<minx or xd>maxx:
return False
return True
# +
Ndim = 6
valid_solns = {tuple([50]*Ndim)}
computed_solns = set(valid_solns)
dims = list(range(Ndim))
dxs = tuple([1]*Ndim)
Nmax = 10
vout, cout = iter_parallel(dims, dxs, valid_solns, computed_solns, Nmax)
# -
valid_solns
# # Testing
Ndim = 1
valid_solns = {tuple([50]),tuple([54])}
computed_solns = set(valid_solns)
explore_pos(tuple([54]*Ndim),0,2,valid_solns,computed_solns)
print(computed_solns)
print(valid_solns)
|
Build Function to Explore Space.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# We have some models that are trained on a subset of the data (e.g. the one that was trained on 100 proteases for 100,000 iterations). Let's check the generalization error on those.
# +
from pin import pin
import pandas as pd
import json
import pickle as pkl
# -
with open('../experiments/outputs/100-graphs_10000-iters_wbs.pkl', 'rb') as f:
wb = pkl.load(f)
wb['layer0_GraphConvLayer']['weights'].min()
data = pd.read_csv('../data/hiv_data/hiv-protease-data-expanded.csv', index_col=0)
data.head()
with open('../data/batch_summary.json') as f:
model_data = json.load(f)
model_data['projects'][0]
# +
# Make the model.
n_graphs = 8
def make_protein_graphs(project, seqid):
"""
Custom function for this script to parallelize the making of protein
graphs over individual cores.
"""
p = pin.ProteinInteractionNetwork('../data/batch_models/{0}/model_01.pdb'
.format(project))
p.graph['project'] = project
p.graph['input_shape'] = p.nodes(data=True)[0][1]['features'].shape
p.graph['seqid'] = seqid
return p
project_id = 10 # change this number to play around with it.
project = model_data['projects'][project_id]['code']
p = pin.ProteinInteractionNetwork('../data/batch_models/{0}/model_01.pdb'.format(project))
p.graph['project'] = project
p.graph['input_shape'] = p.nodes(data=True)[project_id][1]['features'].shape
p.graph['seqid'] = model_data['projects'][project_id]['title']
# -
p.graph
from graphfp.layers import GraphConvLayer, FingerprintLayer, LinearRegressionLayer
from graphfp.flatten import flatten
from graphfp.utils import batch_sample
# +
input_shape = p.graph['input_shape']
layers = [GraphConvLayer(kernel_shape=(input_shape[1], input_shape[1])),
FingerprintLayer(shape=(input_shape)),
LinearRegressionLayer(shape=(input_shape, 1)),
]
# -
graphs = [p]
batch_size = 1
samp_graphs, samp_inputs = batch_sample(graphs, input_shape, batch_size)
def predict(wb_struct, inputs, graphs):
"""
Makes predictions by running the forward pass over all of the layers.
Parameters:
===========
- wb_struct: a dictionary of weights and biases stored for each layer.
- inputs: the input data matrix. should be one row per graph.
- graphs: a list of all graphs.
"""
curr_inputs = inputs
for i, layer in enumerate(layers):
# print(type(wb_struct))
wb = wb_struct['layer{0}_{1}'.format(i, layer)]
curr_inputs = layer.forward_pass(wb, curr_inputs, graphs)
return curr_inputs
preds = predict(wb, samp_inputs, samp_graphs)
preds
data[data.seqid == p.graph['seqid']]['FPV']
# Clearly there is overfitting going on.
|
notebooks/predictions.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# Lists are the R objects which contain elements of different types like − numbers, strings, vectors and another list inside it. A list can also contain a matrix or a function as its elements. List is created using list() function.
#
# **Creating a List**
#
# Following is an example to create a list containing strings, numbers, vectors and a logical values.
# 
list_data <- list("Red", "Green", c(21,32,11), TRUE, 51.23, 119.1)
list_data
# **Naming List Elements**
# The list elements can be given names and they can be accessed using these names.
# Create a list containing a vector, a matrix and a list.
list_data <- list(10,c("Jan","Feb","Mar"), matrix(c(3,9,5,1,-2,8), nrow = 2),
list("green",12.3))
list_data
# +
# Give names to the elements in the list.
names(list_data) <- c("1st Quarter", "A_Matrix", "A Inner list")
# Show the list.
list_data
# -
# **Accessing List Elements**
#
# Elements of the list can be accessed by the index of the element in the list. In case of named lists it can also be accessed using the names.
# Create a list containing a vector, a matrix and a list.
list_data <- list(c("Jan","Feb","Mar"), matrix(c(3,9,5,1,-2,8), nrow = 2),list("green",12.3))
list_data
# +
# Give names to the elements in the list.
names(list_data) <- c("1st Quarter", "A_Matrix", "A Inner list")
# Access the first element of the list.
print(list_data[1])
# -
# Access the thrid element. As it is also a list, all its elements will be printed.
print(list_data[3])
# Access the list element using the name of the element.
print(list_data$A_Matrix)
# **Manipulating List Elements**
#
# We can add, delete and update list elements as shown below. We can add and delete elements only at the end of a list. But we can update any element.
# Create a list containing a vector, a matrix and a list.
list_data <- list(c("Jan","Feb","Mar"), matrix(c(3,9,5,1,-2,8), nrow = 2),
list("green",12.3))
list_data
# +
# Give names to the elements in the list.
names(list_data) <- c("1st Quarter", "A_Matrix", "A Inner list")
# Add element at the end of the list.
list_data[4] <- "New element"
print(list_data[4])
# +
# Remove the last element.
list_data[4] <- NULL
# Print the 4th Element.
print(list_data[4])
# +
# Update the 3rd Element.
list_data[3] <- "updated element"
print(list_data[3])
# -
# **Merging Lists**
#
# You can merge many lists into one list by placing all the lists inside one list() function.
# +
# Create two lists.
list1 <- list(1,2,3)
list2 <- list("Sun","Mon","Tue")
# Merge the two lists By using Vectore.
merged.list <- c(list1,list2)
# Print the merged list.
merged.list
# -
# `Converting List to Vector`
# A list can be converted to a vector so that the elements of the vector can be used for further manipulation. All the arithmetic operations on vectors can be applied after the list is converted into vectors. To do this conversion, we use the unlist() function. It takes the list as input and produces a vector.
# +
# Create lists.
list1 <- list(1:5)
print(list1)
list2 <-list(10:14)
print(list2)
# +
# Convert the lists to vectors.
v1 <- unlist(list1)
v2 <- unlist(list2)
print(v1)
print(v2)
# -
# Now add the vectors
result <- v1+v2
print(result)
|
6. R - Lists.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ml
# language: python
# name: ml
# ---
import pandas as pd
import matplotlib.pyplot as plt
from imgaug import augmenters as iaa
from segmentation_models import Unet, Linknet
from segmentation_models import get_preprocessing
from keras.optimizers import Adam
from keras.callbacks import ModelCheckpoint, TensorBoard, ReduceLROnPlateau
from keras.utils.training_utils import multi_gpu_model
from keras.models import Model
from .utils import *
# +
# Fold 1.
train_df = pd.read_csv("jsrt/jsrt_fold1/train.csv")
val_df = pd.read_csv("jsrt/jsrt_fold1/val.csv")
test_df = pd.read_csv("jsrt/jsrt_fold1/test.csv")
backbone = "resnext50"
preprocessing_fn = get_preprocessing(backbone)
shape = 512
# -
seq = iaa.Sequential([
iaa.Fliplr(0.5),
iaa.OneOf([
iaa.Affine(
scale={"x": (0.8, 1.2), "y": (0.8, 1.2)},
rotate=(-15, 15),
shear=(-10, 10),
),
iaa.Affine(
translate_percent={"x": (-0.1, 0.1), "y": (-0.1, 0.1)},
shear=(-10, 10),
)
])
], random_order=True)
for i in range(10):
show_augm(i, train_df, seq,preprocessing_fn)
val_images, val_masks = load_val(val_df, shape, preprocessing_fn)
model = Unet(backbone_name="resnext50", encoder_weights="imagenet", classes=6)
model.summary()
optimizer = Adam(lr=0.001, decay=1e-7, beta_1=0.9, beta_2=0.999)
model.compile(optimizer=optimizer, loss=bc_dice_loss, metrics=[dice_coefficient, "binary_accuracy"])
batch_size = 4
epochs = 30
callbacks = [
ModelCheckpoint("backup/epoch_{epoch:02d}.hdf5", monitor="val_dice_coef", mode="max", save_weights_only=True, save_best_only=False, verbose=1),
TensorBoard(log_dir="logs", batch_size=batch_size),
ReduceLROnPlateau(monitor="val_dice_coef", factor=0.4, patience=2, verbose=1, mode="max", min_lr=0.000000001),
]
model.fit_generator(generator(batch_size, shape, train_df, seq, preprocessing_fn), validation_data=(val_images, val_masks), steps_per_epoch=500, epochs=epochs, callbacks=callbacks)
model.save_weights("backup/final.hdf5")
model.load_weights("backup/" + sorted(os.listdir("backup"))[-1])
test_images, test_masks = load_test(test_df)
test_results = model.predict(np.array(test_images))
test_results[test_results >= 0.5] = 1
test_results[test_results < 0.5] = 0
print("Lungs: " + str(hard_dice(test_results[:, :, :, 0], test_masks[:, :, :, 0]))
print("Heart: " + str(hard_dice(test_results[:, :, :, 2], test_masks[:, :, :, 0]))
print("Clavicles: " + str(hard_dice(test_results[:, :, :, 3], test_masks[:, :, :, 0]))
print("Lungs: " + str(iou(test_results[:, :, :, 0], test_masks[:, :, :, 0]))
print("Heart: " + str(iou(test_results[:, :, :, 2], test_masks[:, :, :, 0]))
print("Clavicles: " + str(iou(test_results[:, :, :, 3], test_masks[:, :, :, 0]))
# +
layer_outputs = [layer.output for layer in model.layers[-16:]]
activation_model = Model(inputs=model.input, outputs=layer_outputs)
activations = activation_model.predict(np.expand_dims(np.array(test_images[0]), 0))
def display_activation(activations, col_size, row_size, act_index):
activation = activations[act_index]
activation_index=0
fig, ax = plt.subplots(row_size, col_size, figsize=(row_size*2.5,col_size*1.5))
for row in range(0,row_size):
for col in range(0,col_size):
ax[row][col].imshow(activation[0, :, :, activation_index], cmap='gray')
activation_index += 1
# +
plt.figure(figsize=(30, 30))
rows = 4
cols = 4
for i in range(16):
subplot = plt.subplot(rows, cols, i + 1)
subplot.axis("off")
subplot.imshow(activations[-3][0,...,i], cmap="hot")
plt.show()
# +
plt.figure(figsize=(30, 30))
rows = 1
cols = 6
for i in range(6):
subplot = plt.subplot(rows, cols, i + 1)
subplot.axis("off")
subplot.imshow(activations[-2][0,...,i], cmap="hot")
plt.show()
# -
for image, mask in zip(test_images, test_results):
mask[mask >= 0.5] = 255
mask[mask < 0.5] = 0
mask = cv2.cvtColor(mask.astype(np.uint8), cv2.COLOR_GRAY2RGB)
mask[:, :, 1] = 0
mask[:, :, 2] = 0
cv2.addWeighted(mask, 0.4, image, 0.6, 0, image)
# +
plt.figure(figsize=(30, 30))
rows = 6
cols = 4
for i in range(len(test_images[:24])):
subplot = plt.subplot(rows, cols, i + 1)
subplot.axis("off")
subplot.imshow(test_images[i])
plt.show()
# -
|
keras-unet-linknet/unet_fold1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/wesleybeckner/data_science_foundations/blob/main/notebooks/extras/X1_Thinking_Data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8302b110-f682-45eb-ab0a-9c256ebfe6a1"
# # Data Science Foundations <br> X2: Airbnb
#
# **Instructor**: <NAME>
#
# **Contact**: <EMAIL>
#
# ---
#
# <br>
#
# Today we are going to take our newfound knowledge from the course, and practice how we can leverage data to build predictive models. We'll start with a feature engineering problem on some dummy data. This will get us thinking creatively about problem solving. We will then pivot over to an [Airbnb dataset](https://www.kaggle.com/dgomonov/new-york-city-airbnb-open-data/code). After performing some general, exploratory data analysis, we will solve the following business case: Airbnb is interested in using historical list prices from their airbnb hosts, to make pricing suggestions to new hosts. How can we use this existing datset to assist with this price listing suggestion?
#
# <br>
#
# ---
# + [markdown] id="005ab658-621d-410b-b44f-cbd665552ba3"
# ## Prepare Environment and Import Data
# + id="cad88b06-668a-4560-a303-738f2e4e3b9e"
# basic packages
import pandas as pd
import numpy as np
import random
import copy
# visualization packages
import matplotlib.pyplot as plt
import plotly.express as px
import seaborn as sns; sns.set()
import graphviz
# stats packages
import scipy.stats as stats
from scipy.spatial.distance import cdist
import statsmodels.api as sm
from statsmodels.formula.api import ols
from statsmodels.stats.outliers_influence import variance_inflation_factor
# sklearn preprocessing
from sklearn.preprocessing import OneHotEncoder, StandardScaler, PolynomialFeatures
from sklearn.decomposition import PCA
from sklearn.impute import SimpleImputer
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
# sklearn modeling
from sklearn.neighbors import KNeighborsRegressor
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor, AdaBoostClassifier, GradientBoostingClassifier
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn.mixture import GaussianMixture
# sklearn evaluation
from sklearn.metrics import mean_squared_error, r2_score, accuracy_score
from sklearn.model_selection import GridSearchCV, cross_val_score
# + [markdown] id="275c3ab6-0348-45f9-9561-b2b5d187793a"
# ## Warm Up
# + [markdown] id="6be40a8e-5c7e-451a-978c-da16cd1b2841"
# Add aditional feature(s) to `X` to predict `y` with a linear classifier (e.g. logistic regression)
# + id="491a8522-c4f3-40a0-b205-37d3021f0001" outputId="347acc0a-290d-4093-a166-5b744005bcf2"
from sklearn.datasets import make_circles
X, y = make_circles(random_state=42, noise=.01)
relabel = dict(zip([0,1,2,3],[0,1,0,1]))
y = np.vectorize(relabel.get)(y)
plt.scatter(X[:,0], X[:,1], c=y, cmap='viridis')
# + [markdown] id="4ff00110-ff0c-4f80-be2b-4aa22110a9a7"
# and now predict
# + id="5d20f91d-fdc4-4725-9bad-33f5bf18dd70"
# consider using
# LogisticRegression()
# r2_score
# + [markdown] id="2c62ffa6-9995-4b06-bf5d-fc3f28ff8168"
# ## Exploratory Data Analysis
# + [markdown] id="62a366ab-3a44-4b39-b569-dcf920a05015"
# which columns are numerical, string; which contain nans/nulls; what is the correlation between features
# + id="41c49b64-910b-4ccf-8e8f-3a55cf38c1a2"
airbnb = pd.read_csv("https://raw.githubusercontent.com/wesleybeckner/datasets/main/datasets/airbnb/AB_NYC_2019.csv")
# + id="ba79dab1-3881-47d4-891f-e86d6f53621b"
# recall these attributes
# shape dtypes
# and these methods
# head() tail() isnull() sum() nunique() copy() select_dtypes() describe() drop()
# + [markdown] id="44a60c76-dfa7-4643-bd86-238a15f7472a"
# what visualizations would be useful to make?
# + id="d1c0a022-6b4f-41c8-8631-58fb71323578" outputId="7e34cd05-16dc-4ff6-903c-aceb5cea71b7"
plt.figure(figsize=(10,6))
sns.scatterplot(x=airbnb.longitude,y=airbnb.latitude,hue=airbnb.neighbourhood_group)
# + [markdown] id="265d19ea-251f-448b-b75e-3de82921c096"
# ## Feature Engineering
#
# Say we want to predict `price`, using an ML model. How would you build your features?
#
# Based on the number of null values, what would you do with the `last_review` and `reviews_per_month` column?
# + id="11b99d05-a3d7-4009-af6e-5d14986d2d85"
X = airbnb.copy()
# + [markdown] id="2abb4566-4ba8-4c89-99c7-21459fa72ae2"
# How will we deal with the categorical features?
# + id="63bad0f2-0542-4ea1-8292-1a0cf3c4ef5e" outputId="d0b49439-0bb4-4223-eb16-468f7995c104"
# Recall
# OneHotEncoder()
X_cat = X.select_dtypes(include='object')
display(X_cat.head())
print(X_cat.nunique())
# + id="fcf3ec49-80e5-47f8-9b06-a3bdd061ae4c" outputId="a233ee1f-0443-4a44-f235-9656d33ad2c1"
X_num = X.select_dtypes(exclude='object')
X_num.head()
# + [markdown] id="1aed4997-4633-493e-9e7c-6f43483d75c9"
# ## Feature Transformation
#
# What features do you think will cause the most problems if untransformed?
#
# Scale and Center all but the target variable, price
# + id="d2d19696-afe5-4fae-929a-cca5fbbc3f1d"
# Recall
# StandardScaler() df.pop()
# + [markdown] id="d89113a6-0d40-46b0-a0ad-1e8ea471f621"
# ## Model Baseline
# + id="c1b2ce33-2ab3-4d97-90f3-44c4e3b0e003"
# Recall
# X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.8, random_state=42)
# LinearRegression()
# r2_score()
# + [markdown] id="4d090f38-16ef-414f-9569-950e87be0245"
# try a model that captures non-linear relationships
# + id="8cf0ab3b-a318-4e71-b5c5-a8d226fa48ff"
# Recall
# RandomForestRegressor()
# + [markdown] id="f8502dd9-c243-4b10-945a-1dd13523f1c4"
# both of these results from the `LinearRegression` and `RandomForest` models should indicate something to you (think back to [Model Selection and Validation](https://wesleybeckner.github.io/data_science_foundations/S3_Model_Selection_and_Validation/))
# + [markdown] id="dd45db4e-9043-439b-9236-a8b78db3106a"
# ## Additional Strategies
#
# After this first pass, what are some additional strategies to consider for improving the model?
|
notebooks/extras/X2_Airbnb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Softmax Sklearn CIFAR10
# +
# import the necessary packages
from sklearn.preprocessing import LabelBinarizer
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
from sklearn import datasets
import matplotlib.pyplot as plt
import numpy as np
# +
from keras.datasets import cifar10
# load the training and testing data, scale it into the range [0, 1],
# then reshape the design matrix
print("[INFO] loading CIFAR-10 data...")
((trainX, trainY), (testX, testY)) = cifar10.load_data()
trainX = trainX.reshape((trainX.shape[0], 3072))
testX = testX.reshape((testX.shape[0], 3072))
# +
# append the bias dimension of ones (i.e. bias trick) so that our SVM
# only has to worry about optimizing a single weight matrix W.
trainX = np.hstack([trainX, np.ones((trainX.shape[0], 1))])
testX = np.hstack([testX, np.ones((testX.shape[0], 1))])
print(trainX.shape, testX.shape)
# -
from sklearn.linear_model import SGDClassifier
model = SGDClassifier(loss="log", penalty='l2', alpha=2.5e4, eta0=5e-7, n_jobs=-1)
# +
classes = ['plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
labelBinarizer = LabelBinarizer()
classes = labelBinarizer.fit_transform(classes)
batch_size = 200
num_iters = 1000
num_train = 50000
for it in range(num_iters):
X_batch = None
y_batch = None
#########################################################################
# TODO: #
# Sample batch_size elements from the training data and their #
# corresponding labels to use in this round of gradient descent. #
# Store the data in X_batch and their corresponding labels in #
# y_batch; after sampling X_batch should have shape (batch_size, dim) #
# and y_batch should have shape (batch_size,) #
# #
# Hint: Use np.random.choice to generate indices. Sampling with #
# replacement is faster than sampling without replacement. #
#########################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
batch_idx = np.random.choice(num_train, batch_size)
X_batch = trainX[batch_idx]
Y_batch = trainY[batch_idx]
model.partial_fit(X_batch, Y_batch.ravel(), classes=classes)
# -
# evaluate the classifier
acc = model.score(testX, testY.ravel())
print("[INFO] accuracy: {:.2f}%".format(acc * 100))
# Well Done
|
notebooks/deep-learning-for-computer-vision-with-python/109_Softmax_Sklearn_CIFAR10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # RNN for Classifying Names
# In this notebook we are building and training a basic character-level RNN to classify
# words. A character-level RNN reads words as a series of characters -
# outputting a prediction and "hidden state" at each step, feeding its
# previous hidden state into each next step. We take the final prediction
# to be the output, i.e. which class the word belongs to.
# ### Preparing the Data
#
# Download the data in folder `data/names` from GitHub.
#
# Included in the ``data/names`` directory are 18 text files named as
# ``[Language].txt``. Each file contains a bunch of names, one name per
# line, mostly romanized (but we still need to convert from Unicode to
# ASCII).
# +
import string
import unicodedata
# these is the vocabulary we will use
all_letters = string.ascii_letters + " .,;'"
n_letters = len(all_letters)
print(f"Vocab is of size {n_letters} and contains:", all_letters)
# +
# we convert anything into ascii
def unicodeToAscii(s):
return ''.join(
c for c in unicodedata.normalize('NFD', s)
if unicodedata.category(c) != 'Mn'
and c in all_letters
)
print(unicodeToAscii('Ślusàrski'))
print(unicodeToAscii('Heute ist es schön heiß'))
# +
from io import open
import glob
import os
# Read a file and split into lines
def readLines(filename):
lines = open(filename, encoding='utf-8').read().strip().split('\n')
return [unicodeToAscii(line) for line in lines]
all_categories = []
X = []
y = []
for filename in glob.glob('data/names/*.txt'):
category = os.path.splitext(os.path.basename(filename))[0]
all_categories.append(category)
lines = readLines(filename)
for line in lines:
X.append(line)
y.append(category)
n_categories = len(all_categories)
n_categories, len(X)
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
print("Train data points:", len(X_train))
# -
# Turning Names into Tensors
# --------------------------
#
# Now that we have all the names organized, we need to turn them into
# Tensors to make any use of them.
#
# To represent a single letter, we use a "one-hot vector" of size
# ``<1 x n_letters>``. A one-hot vector is filled with 0s except for a 1
# at index of the current letter, e.g. ``"b" = <0 1 0 0 0 ...>``.
#
# To make a word we join a bunch of those into a 2D matrix
# ``<line_length x 1 x n_letters>``.
#
# That extra 1 dimension is because PyTorch assumes everything is in
# batches - we're just using a batch size of 1 here.
#
#
#
# +
import torch
def letterToTensor(letter):
tensor = torch.zeros(1, n_letters)
index = all_letters.find(letter)
tensor[0][index] = 1
return tensor
print(letterToTensor('J'))
# +
# Turn a line into a <line_length x 1 x n_letters>,
# or an array of one-hot letter vectors
def lineToTensor(line):
tensor = torch.zeros(len(line), 1, n_letters)
for i, letter in enumerate(line):
index = all_letters.find(letter)
tensor[i][0][index] = 1
return tensor
print(lineToTensor('Jones').size())
# +
def categoryToTensor(category):
index = all_categories.index(category)
return torch.tensor([index], dtype=torch.long)
categoryToTensor("Korean")
# -
# Creating the Network
# ====================
#
# Before autograd, creating a recurrent neural network in Torch involved
# cloning the parameters of a layer over several timesteps. The layers
# held hidden state and gradients which are now entirely handled by the
# graph itself. This means you can implement a RNN in a very "pure" way,
# as regular feed-forward layers.
#
# This RNN module is just 2 linear layers which operate on an input and hidden state, with
# a LogSoftmax layer after the output.
# You can see the architecture here: https://i.imgur.com/Z2xbySO.png
# +
import torch.nn as nn
class RNN(nn.Module):
def __init__(self, input_size, output_size):
super(RNN, self).__init__()
self.hidden_size = 128 # number of hidden layer size
self.i2h = nn.Linear(input_size + self.hidden_size, self.hidden_size)
self.i2o = nn.Linear(input_size + self.hidden_size, output_size)
def forward(self, x, hidden):
combined = torch.cat((x, hidden), 1)
hidden = self.i2h(combined)
output = self.i2o(combined)
return output, hidden
def initHidden(self):
return torch.zeros(1, self.hidden_size)
# -
# To run a step of this network we need to pass an input (in our case, the
# Tensor for the current letter) and a previous hidden state (which we
# initialize as zeros at first). We'll get back the output (probability of
# each language) and a next hidden state (which we keep for the next
# step).
#
#
#
# +
rnn = RNN(n_letters, n_categories)
x = letterToTensor('A')
hidden = torch.zeros(1, 128)
output, next_hidden = rnn(x, hidden)
print(torch.softmax(output, 1))
# -
# As you can see the output is a ``<1 x n_categories>`` Tensor, where
# every item is the likelihood of that category (higher is more likely).
#
#
#
# Task 1: Training the Network
# --------------------
#
# Finish the following training function to train the RNN on the training data set.
# +
import math
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(rnn.parameters(), lr=0.005)
for epoch in range(1, 10):
print("Training epoch:", epoch)
# iterate through all names in X_train
# for every name:
# init the hidden layer of the rnn
# insert the name character by character into the rnn and compute the final output
# note: you need to carry on the hidden state in every time step
# define the loss on the last output of the rnn and the category (=label)
# backpropagate the loss and take an optimizer step
# -
# ### Task 2: Evaluating the Results
#
# Evaluate the accuarcy of the RNN on the test data.
# ### Task 3: Running on User Input
#
# Write a function that takes an abritrary names as input and outputs the top 3 categories of the RNN for the input.
#
|
7_RNNs_ClassifyNames.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JimKing100/DS-Unit-2-Applied-Modeling/blob/master/DS_Sprint_Challenge_7a.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Kz9V5lVFMo86" colab_type="text"
# _Lambda School Data Science, Unit 2_
#
# # Applied Modeling Sprint Challenge: Predict Chicago food inspections 🍔
# + [markdown] id="yWOjtM9iMo87" colab_type="text"
# For this Sprint Challenge, you'll use a dataset with information from inspections of restaurants and other food establishments in Chicago from January 2010 to March 2019.
#
# [See this PDF](https://data.cityofchicago.org/api/assets/BAD5301B-681A-4202-9D25-51B2CAE672FF) for descriptions of the data elements included in this dataset.
#
# According to [Chicago Department of Public Health — Food Protection Services](https://www.chicago.gov/city/en/depts/cdph/provdrs/healthy_restaurants/svcs/food-protection-services.html), "Chicago is home to 16,000 food establishments like restaurants, grocery stores, bakeries, wholesalers, lunchrooms, mobile food vendors and more. Our business is food safety and sanitation with one goal, to prevent the spread of food-borne disease. We do this by inspecting food businesses, responding to complaints and food recalls."
# + [markdown] id="VvBYdx2xMo88" colab_type="text"
# #### Your challenge: Predict whether inspections failed
#
# The target is the `Fail` column.
#
# - When the food establishment failed the inspection, the target is `1`.
# - When the establishment passed, the target is `0`.
# + [markdown] id="3YMtu4LaF8Jq" colab_type="text"
# #### Run this cell to install packages in Colab:
# + id="vWyiJKQgF6ax" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3f2aa327-3642-470e-96e1-935d9d62b070"
import sys
if 'google.colab' in sys.modules:
# Install packages in Colab
# !pip install category_encoders==2.*
# !pip install eli5
# !pip install pandas-profiling==2.*
# !pip install pdpbox
# !pip install shap
# + [markdown] id="Na5IvRq1Mo89" colab_type="text"
# #### Run this cell to load the data:
# + id="Yyc7zftTMo89" colab_type="code" colab={}
import pandas as pd
train_url = 'https://drive.google.com/uc?export=download&id=13_tP9JpLcZHSPVpWcua4t2rY44K_s4H5'
test_url = 'https://drive.google.com/uc?export=download&id=1GkDHjsiGrzOXoF_xcYjdzBTSjOIi3g5a'
train = pd.read_csv(train_url)
test = pd.read_csv(test_url)
assert train.shape == (51916, 17)
assert test.shape == (17306, 17)
# + [markdown] id="8EhbzqgfMo9A" colab_type="text"
# ### Part 1: Preprocessing
#
# You may choose which features you want to use, and whether/how you will preprocess them. If you use categorical features, you may use any tools and techniques for encoding.
#
# _To earn a score of 3 for this part, find and explain leakage. The dataset has a feature that will give you an ROC AUC score > 0.90 if you process and use the feature. Find the leakage and explain why the feature shouldn't be used in a real-world model to predict the results of future inspections._
#
# ### Part 2: Modeling
#
# **Fit a model** with the train set. (You may use scikit-learn, xgboost, or any other library.) Use cross-validation or do a three-way split (train/validate/test) and **estimate your ROC AUC** validation score.
#
# Use your model to **predict probabilities** for the test set. **Get an ROC AUC test score >= 0.60.**
#
# _To earn a score of 3 for this part, get an ROC AUC test score >= 0.70 (without using the feature with leakage)._
#
#
# ### Part 3: Visualization
#
# Make visualizations for model interpretation. (You may use any libraries.) Choose two of these types:
#
# - Permutation Importances
# - Partial Dependence Plot, 1 feature isolation
# - Partial Dependence Plot, 2 features interaction
# - Shapley Values
#
# _To earn a score of 3 for this part, make all four of these visualization types._
# + [markdown] id="muEIQ4EPGVH_" colab_type="text"
# ## Part 1: Preprocessing
#
# > You may choose which features you want to use, and whether/how you will preprocess them. If you use categorical features, you may use any tools and techniques for encoding.
# + id="lnJVkBYqy2qZ" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestClassifier
from sklearn.pipeline import make_pipeline
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_error
# + id="Ao1PDJtpIWwp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 581} outputId="682bfa24-d5dc-4414-a14b-82f43eb68d5a"
train.head()
# + id="SDbB_srwyLgk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="54eb2760-1d06-4022-db7d-cd858f53e93c"
# Check for nulls
train.isnull().sum()
# + id="fvIhYnR2ysHm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="4e5725ec-9c34-4303-9ed5-beae4b2e09c7"
# Explore features
for col in sorted(train.columns):
if train[col].nunique() < 12:
sns.catplot(x=col, y='Fail', data = train, kind = 'bar', color = 'grey')
plt.show()
# + id="lBOH263WzfCk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="d1ae2ba2-ab7e-429a-8933-5ac8e4c7fc74"
# Wrangle the data for train and test
def engineer_features(X):
# Convert date_recorded to datetime
X['Inspection Date'] = pd.to_datetime(X['Inspection Date'], infer_datetime_format=True)
# Extract components from date_recorded and drop the original column
# X['year_inspection'] = X['Inspection Date'].dt.year
# X['month_inspection'] = X['Inspection Date'].dt.month
# X['day_inspection'] = X['Inspection Date'].dt.day
X = X.drop(columns='Inspection Date')
X = X.drop(columns='AKA Name')
X = X.drop(columns='Location')
X = X.drop(columns='City')
X = X.drop(columns='State')
X = X.drop(columns='Violations')
return X
train = engineer_features(train)
test = engineer_features(test)
print(train.shape)
train.head()
# + [markdown] id="nB8CQnExGqzr" colab_type="text"
# ## Part 2: Modeling
#
# > **Fit a model** with the train set. (You may use scikit-learn, xgboost, or any other library.) Use cross-validation or do a three-way split (train/validate/test) and **estimate your ROC AUC** validation score.
# >
# > Use your model to **predict probabilities** for the test set. **Get an ROC AUC test score >= 0.60.**
# + id="YQ3hD2G1IbVH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4f4cf890-1eb7-4957-d30a-7e23b9433d23"
# Split training data into training and validation sets 80/20
train, val = train_test_split(train, train_size=0.80, test_size=0.20, random_state=42)
print(train.shape, val.shape, test.shape)
# + id="ITeb0tAo1Nw0" colab_type="code" colab={}
# Encode and fit a Random Forest Model - Optimization done at end and value used here
target = 'Fail'
features = train.columns.drop(target)
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
X_test = test[features]
y_test = test[target]
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='mean'),
RandomForestClassifier(n_estimators=100, max_depth=2, random_state=42, verbose=1)
)
# + id="-UvyipN21wRV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="a9c3cabc-9c90-4365-d309-5e6a6ad835a0"
# Get validation score
pipeline.fit(X_train, y_train)
print ('Validation Accuracy', pipeline.score(X_val, y_val))
# + id="rRNRLsli18IA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="9c324fe2-c31a-4939-9630-9b23adf5bf85"
from sklearn.metrics import roc_auc_score
y_pred_proba = pipeline.predict_proba(X_val)[:, 1]
roc_auc_score(y_val, y_pred_proba)
# + id="JyLrakhH2F4C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="193f1d55-c379-475f-8ad5-9f7d4d1baa1f"
# Plot ROC curve
import matplotlib.pyplot as plt
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_val==1, y_pred_proba)
plt.plot(fpr, tpr)
plt.title('ROC curve')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate');
# + id="4CJZKjqC77ci" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="4d342b40-e519-4386-e3c5-0c0b5d3893d6"
y_pred_proba = pipeline.predict_proba(X_test)[:, 1]
roc_auc_score(y_test, y_pred_proba)
# + id="uC7cIDTd4FgZ" colab_type="code" colab={}
# from sklearn.ensemble import RandomForestRegressor
# from sklearn.model_selection import RandomizedSearchCV
# # Number of trees in random forest
# n_estimators = [int(x) for x in np.linspace(start = 200, stop = 500, num = 10)]
# # Number of features to consider at every split
# max_features = ['auto', 'sqrt']
# # Maximum number of levels in tree
# max_depth = [int(x) for x in np.linspace(10, 50, num = 11)]
# max_depth.append(None)
# # Minimum number of samples required to split a node
# min_samples_split = [2, 5, 10]
# # Minimum number of samples required at each leaf node
# min_samples_leaf = [1, 2, 4]
# # Method of selecting samples for training each tree
# bootstrap = [True, False]
# # Create the random grid
# random_grid = {'n_estimators': n_estimators,
# 'max_features': max_features,
# 'max_depth': max_depth,
# 'min_samples_split': min_samples_split,
# 'min_samples_leaf': min_samples_leaf,
# 'bootstrap': bootstrap}
# print(random_grid)
# pipeline = make_pipeline (
# ce.OrdinalEncoder(),
# SimpleImputer(strategy='mean'),
# RandomizedSearchCV(estimator = RandomForestRegressor(),
# param_distributions = random_grid,
# n_iter = 5,
# verbose=2,
# random_state=42,
# n_jobs = -1)
# )
# pipeline.fit(X_train, y_train)
# pd.set_option('display.max_rows', 200)
# model = pipeline.named_steps['randomizedsearchcv']
# best = pd.Series(model.best_params_)
# print(best)
# + [markdown] id="u98nLGBTMo9s" colab_type="text"
# ## Part 3: Visualization
#
# > Make visualizations for model interpretation. (You may use any libraries.) Choose two of these types:
# >
# > - Permutation Importances
# > - Partial Dependence Plot, 1 feature isolation
# > - Partial Dependence Plot, 2 features interaction
# > - Shapley Values
# + id="_Q379i5CIeKY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="c80083c8-1f73-445e-8c80-791b9ced1b5c"
import eli5
from eli5.sklearn import PermutationImportance
transformers = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='mean')
)
X_train_transformed = transformers.fit_transform(X_train)
X_val_transformed = transformers.fit_transform(X_val)
model = RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
model.fit(X_train_transformed, y_train)
# + id="ZxMPgeEs6n8m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="be5a238f-b70b-43f0-e37b-cf796515c824"
permuter = PermutationImportance(
model,
scoring='accuracy',
n_iter=2,
random_state=42
)
permuter.fit(X_val_transformed, y_val)
feature_names = X_val.columns.tolist()
eli5.show_weights(
permuter,
top=None,
feature_names = feature_names
)
# + id="GGIyjt-3AJBo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="d0e04d45-b9a3-4238-be00-f1c368dc9918"
X_test.head()
# + id="FfTJoDG523Lp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 193} outputId="349c3b5a-72bc-4d08-f569-3eac6980068a"
import shap
row = X_test.iloc[[1]]
explainer = shap.TreeExplainer(model)
row_process = transformers.transform(row)
shap_values = explainer.shap_values(row_process)
shap.initjs()
shap.force_plot(
base_value=explainer.expected_value[1],
shap_values=shap_values[1],
features=row
)
|
DS_Sprint_Challenge_7a.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib notebook
import numpy as np
import math
import scipy as sp
import copy
import os
import matplotlib.pyplot as plt
from libwallerlab.projects.motiondeblur import blurkernel
import bluranalysis as analysis
# plt.style.use('deblur')
# -
# ## Blur Len vs Beta
# +
# blur_len = np.arange(1, 100)
# beta = np.arange(0.1,1.0, 0.01)
# image = []
# for _len in blur_len:
# for _beta in beta:
# image.append(analysis.getOptimalDnf(_len, _beta))
# plt.figure()
# plt.imshow(np.asarray(np.log10(image)).reshape(len(blur_len), len(beta)), vmin=0, vmax=2)
# -
# ## Show kernel and padded kernel in frequency domain
# +
x = np.zeros(100)
x[5] = 1
x[19] =1
x[14] =1
x_padded =np.pad(x, (0,100), mode='constant')
plt.figure()
plt.plot(np.abs(np.fft.fft(x)) ** 2, label='original')
plt.plot(np.abs(np.fft.fft(x_padded)) ** 2, label='padded')
plt.legend()
plt.xlabel('Fourier Coefficient')
plt.ylabel('Magnitude')
plt.title('Effect of Zero-padding')
plt.tight_layout()
print(analysis.calcCondNumFromKernel(x))
print(analysis.calcCondNumFromKernel(x_padded))
print(analysis.calcDnfFromKernel(x))
print(analysis.calcDnfFromKernel(x_padded))
# -
# ## Pulse Length and DNF
# +
kernel_len_list = np.arange(11,1000)
n_pulses = 10
dnf_list = []
for kernel_len in kernel_len_list:
dnf_list.append(analysis.getOptimalDnf(kernel_len, n_pulses=n_pulses, n_tests=100))
plt.figure()
plt.plot(dnf_list)
plt.xlabel('Sequence Length')
plt.ylabel('DNF')
plt.title('Effect of Sequence Length on DNF, %d pulses' % n_pulses)
plt.tight_layout()
# +
kernel_len_list = np.arange(51,1000)
n_pulses = 50
dnf_list = []
for kernel_len in kernel_len_list:
dnf_list.append(analysis.getOptimalDnf(kernel_len, n_pulses=n_pulses, n_tests=100))
plt.figure()
plt.plot(dnf_list)
plt.xlabel('Sequence Length')
plt.ylabel('DNF')
plt.title('Effect of Sequence Length on DNF, %d pulses' % n_pulses)
plt.tight_layout()
# -
# ## DNF vs Pulse Count
# +
pulse_count_list = np.arange(3,1500)
dnf_list = []
for pulse_count in pulse_count_list:
dnf_list.append(analysis.getOptimalDnf(pulse_count * 2, n_pulses=pulse_count, n_tests=100))
# Perform log fit
coeffs = np.polyfit(np.log10(pulse_count_list), dnf_list, 1)
y = coeffs[0] * np.log10(pulse_count_list) + coeffs[1]
def func_powerlaw(x, m):
return np.sqrt(2) * x**m - 1
sol1, _ = sp.optimize.curve_fit(func_powerlaw, pulse_count_list, dnf_list, maxfev=2000 )
# sol1 = [0.6116, np.sqrt(2), -1]
yp = func_powerlaw(pulse_count_list, sol1[0])
plt.figure(figsize=(12,5))
plt.plot(pulse_count_list, dnf_list, label='Calculated DNF')
plt.plot(pulse_count_list, yp, label='Power Law Fit')
plt.xlabel('Pulse Count')
plt.ylabel('DNF')
plt.title('Effect of Pulse Count on DNF')
plt.legend()
plt.tight_layout()
# -
# ## Try with $3\times$ pulse count
# +
pulse_count_list = np.arange(3,500)
n_pulses = 50
dnf_list = []
for pulse_count in pulse_count_list:
dnf_list.append(analysis.getOptimalDnf(pulse_count * 3, n_pulses=pulse_count, n_tests=100))
# Perform log fit
coeffs = np.polyfit(np.log10(pulse_count_list), dnf_list, 1)
y = coeffs[0] * np.log10(pulse_count_list) + coeffs[1]
def func_powerlaw(x, m):
return np.sqrt(2) * x**m - 1
sol1, _ = sp.optimize.curve_fit(func_powerlaw, pulse_count_list, dnf_list, maxfev=2000 )
# sol1 = [0.6116, np.sqrt(2), -1]
yp = func_powerlaw(pulse_count_list, sol1[0])
plt.figure(figsize=(12,5))
plt.plot(pulse_count_list, dnf_list, label='Calculated DNF')
plt.plot(pulse_count_list, yp, label='Power Law Fit')
plt.xlabel('Pulse Count')
plt.ylabel('DNF')
plt.title('Effect of Pulse Count on DNF')
plt.legend()
plt.tight_layout()
# -
# ## What does the SNR vs n_pulses curve look like?
# +
N = np.arange(3,500)
c = 10
snr_strobed = np.sqrt(c)
f = func_powerlaw(pulse_count_list, sol1[0])
snr_imaging = np.sqrt(N * c)
snr_dnf = snr_imaging / f
plt.figure()
plt.plot(pulse_count_list, snr_dnf, label='SNR Improvement')
plt.plot(pulse_count_list, snr_strobed * np.ones_like(snr_dnf), label='Baseline (Strobed)')
# plt.plot(pulse_count_list, snr_0, label='Imaging SNR')
plt.legend()
plt.xlabel('Number of Pulses')
plt.ylabel('SNR')
plt.tight_layout()
# +
# signal_photons = 100
# noise_var = 1000
# G = np.sqrt(1 + noise_var / signal_photons)
# print(G)
# -
# # What if you add signal-independent noise?
# +
N = np.arange(3,500)
c = 1000
var_dependent = N * c
var_independent = 0
# Calculate DNF
f = func_powerlaw(pulse_count_list, sol1[0])
# Calculate SNR for astrobed and coded illum
snr_strobed = c / np.sqrt(c + var_independent)
snr_coded = c * N / (f * np.sqrt(N * c + var_independent))
plt.figure()
plt.plot(pulse_count_list, snr_coded, label='Decoded SNR')
plt.plot(pulse_count_list, snr_strobed * np.ones_like(snr_coded), label='Baseline (Strobed) SNR')
# plt.plot(pulse_count_list, snr_0, label='Imaging SNR')
plt.legend()
plt.xlabel('Number of Pulses')
plt.ylabel('SNR')
plt.xlim((0,300))
plt.tight_layout()
# -
# ## Plot SNR of Strobed and Coded Illumination Under Different Amounts of Readout Noise
# +
N = np.arange(3,500)
c = 1000
var_dependent = N * c
var_independent = 500
# Calculate DNF as a function of N
dnf_list = func_powerlaw(pulse_count_list, sol1[0])
# Create variance list
var_independent_list = np.arange(0, 10000, 100)
plt.figure(figsize=(9,7))
snr_strobed_list, snr_coded_list = [], []
for var_independent in var_independent_list:
for dnf in dnf_list:
# Calculate SNR for astrobed and coded illum
snr_strobed = c / np.sqrt(c + var_independent)
snr_coded = c * N / (f * np.sqrt(N * c + var_independent))
snr_strobed_list.append(snr_strobed)
snr_coded_list.append(snr_coded)
snr_strobed_image = np.asarray(snr_strobed_list).reshape((len(var_independent_list), len(dnf_list)))
snr_coded_image = np.asarray(snr_coded_list).reshape((len(var_independent_list), len(dnf_list)))
# -
# ## Plot SNR of Strobed and Coded Illumination Under Different Amounts of Readout Noise
# +
N = pulse_count_list
c = 1000
var_dependent = N * c
var_independent = 500
# Calculate DNF
f = func_powerlaw(pulse_count_list, sol1[0])
plt.figure(figsize=(9,7))
for index, var_independent in enumerate([0, 500, 1000, 5000]):
plt.subplot(411 + index)
# Calculate SNR for astrobed and coded illum
snr_strobed = c / np.sqrt(c + var_independent)
snr_coded = c * N / (f * np.sqrt(N * c + var_independent))
plt.plot(pulse_count_list, snr_coded, label='Decoded SNR', lw=3)
plt.plot(pulse_count_list, snr_strobed * np.ones_like(snr_coded), label='Baseline (Strobed) SNR', lw=3)
# plt.plot(pulse_count_list, snr_0, label='Imaging SNR')
if index ==0:
plt.legend()
plt.xlabel('Number of Pulses')
plt.ylabel('SNR')
plt.xlim((0,300))
plt.title('Signal-Independent Noise Variance: %d counts' % var_independent)
plt.tight_layout()
|
notebooks/publications/old/fig5_dnf_analysis_and_illum_opt.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Rkh-DMFHRIuF"
# # Demo of pre-trained anime character identification
# + id="uj4VvGUD0v9j"
# ! pip install git+https://github.com/kosuke1701/AnimeCV.git
# + id="LW9XwO591FKf"
# !wget https://github.com/kosuke1701/AnimeCV/releases/download/0111_best_randaug/0111_best_randaug.zip
# !unzip 0111_best_randaug
# + id="YCaMRmrW3fn-"
# Face detection module
from animecv.object_detection import FaceDetector_EfficientDet
from animecv.util import load_image
detector = FaceDetector_EfficientDet(coef=2, use_cuda=True)
# + id="5vBSVgTz1O-5"
# Character face encoder
import animecv
from animecv.module import ImageBBEncoder, Similarity
from torchvision import transforms
torch_model = animecv.general.OML_ImageFolder_Pretrained("0111_best_randaug")
transform = [
transforms.Resize((224,224)), # 0206_seresnet152 uses input size of 256
transforms.ToTensor(),
transforms.Normalize(
mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225]
)
]
transform = transforms.Compose(transform)
encoder = ImageBBEncoder(torch_model, post_trans=transform, scale=1.0)
encoder.to("cuda")
threshold = 0.65 # Threshold of dot-product of embeddings which is determined so that the model's FPR becomes 0.22.
# Use following threshold when using different models.
# 0206_resnet18: 0.601
# 0206_resnet152: 0.645
# 0206_seresnet152: 0.656
# + id="SLLV-6C61mKu"
from google.colab import files
import IPython
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Upload your image here. Two images are required.
#
# ここで画像をアップロードしてください。合計2枚の画像をアップロードします。
# + id="4JGIv8Jq1quV"
uploaded = list(files.upload())
image1 = uploaded[0]
IPython.display.Image(image1, width=300)
# + id="5L8n59rK5my8"
uploaded = list(files.upload())
image2 = uploaded[0]
IPython.display.Image(image2, width=300)
# + id="uSFAdQt99AMY"
images = [load_image(image1), load_image(image2)]
face_bbox = detector.detect(images)
face_embs, lst_i_img, lst_i_bbox = encoder.encode(images, face_bbox)
face_embs = face_embs.detach().cpu().numpy()
cropped_images = []
for i_img, i_bbox in zip(lst_i_img, lst_i_bbox):
xmin, ymin, xmax, ymax = face_bbox[i_img][i_bbox]["coordinates"]
crop_img = images[i_img].crop((xmin, ymin, xmax, ymax))
if min(crop_img.size) == 0:
continue
cropped_images.append(crop_img)
n_img = len(cropped_images)
print(f"Detected {n_img} faces.")
for i_img, img in enumerate(cropped_images):
ax = plt.subplot(1, n_img, i_img+1)
ax.imshow(np.array(img))
plt.show()
print("Similarity of each face pair. Rows and columns correspond to each image.")
for i_img in range(n_img):
line = []
for j_img in range(n_img):
sim = np.dot(face_embs[i_img], face_embs[j_img])
label = "SAME" if sim > threshold else "DIFF"
line.append(f"{sim:.3f}/{label}")
print("\t".join(line))
|
examples/demo_oml.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# install py2neo to connect python to the platform Neo4j
# #!pip install py2neo
# -
# Import the essentials to create a graph
from py2neo import Graph, Node, Relationship
# Set the environnement and the connection between the browser and the notebook
graph = Graph(uri="bolt://localhost:7687", auth=("julien", "julien"))
# Set an easy example
alice = Node("person",name="alice")
bob = Node("person",name="bob")
alice_knows_bob = Relationship(alice,"KNOWS",bob)
graph.create(alice_knows_bob)
# MATCH (n) Return n
# +
nicole = Node("Person", name="Nicole", age=24)
drew = Node("Person", name="Drew", age=20)
mtdew = Node("Drink", name="Mountain Dew", calories=9000)
cokezero = Node("Drink", name="Coke Zero", calories=0)
coke = Node("Manufacturer", name="Coca Cola")
pepsi = Node("Manufacturer", name="Pepsi")
graph.create(nicole | drew | mtdew | cokezero | coke | pepsi)
# +
from scripts.vis import draw
options = {"Person": "name", "Drink": "name", "Manufacturer": "name"}
draw(graph, options)
# -
from pyvis.network import Network
net = Network(notebook=True)
net.add_node(1, label="Facebook") # node id = 1 and label = Node 1
net.add_node(2, label="isLocated") # node id and label = 2
net.add_node(3, label="Bordeaux") # node id and label = 2
net.add_edge(1,2, value = 2)
net.add_edge(2,3, value = 2, title="test")
#net.enable_physics(True)
net.show_buttons(filter_=['physics'])
net.show("mygraph.html")
|
JupyNeo4j/KnowledgeRepresentationEntities.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import torch.utils.data
from torch.autograd import Variable
import torch.nn as nn
import torch.optim as optim
import numpy as np
import h5py
from data_utils import get_data
import matplotlib.pyplot as plt
# +
# Load data from all .mat files, combine them, eliminate EOG signals, shuffle and
# seperate training data, validation data and testing data.
# Also do mean subtraction on x.
data = get_data('../project_datasets',num_validation=100, num_test=50)
for k in data.keys():
print('{}: {} '.format(k, data[k].shape))
# -
# class flatten to connect to FC layer
class Flatten(nn.Module):
def forward(self, x):
N, C, H = x.size() # read in N, C, H
return x.view(N, -1)
# +
# turn x and y into torch type tensor
N_train, C_train, H_train = data.get('X_train').shape
N_val, C_val, H_val = data.get('X_val').shape
N_test, C_test, H_test = data.get('X_test').shape
dtype = torch.FloatTensor
X_train = Variable(torch.Tensor(data.get('X_train'))).type(dtype)
y_train = Variable(torch.Tensor(data.get('y_train'))).type(torch.LongTensor)
X_val = Variable(torch.Tensor(data.get('X_val'))).type(dtype)
y_val = Variable(torch.Tensor(data.get('y_val'))).type(torch.LongTensor)
X_test = Variable(torch.Tensor(data.get('X_test'))).type(dtype)
y_test = Variable(torch.Tensor(data.get('y_test'))).type(torch.LongTensor)
# +
# set up sequential model
model = nn.Sequential(
nn.Conv1d(22, 10, kernel_size=12, stride=4),
nn.ReLU(inplace=True),
nn.Dropout(p=0.5),
nn.BatchNorm1d(num_features=10),
nn.MaxPool1d(kernel_size=4, stride=4),
Flatten(),
nn.Linear(620, 20),
nn.Dropout(p=0.5),
nn.ReLU(inplace=True),
nn.Linear(20, 4)
)
model.type(dtype)
loss_fn = nn.CrossEntropyLoss().type(dtype)
optimizer = optim.Adam(model.parameters(), lr=1e-4)
# +
# train through several iterations
num_epoch = 300
batch_size = 80
step = np.arange(0,N_train,batch_size)
#step = np.append(step,N_train) #discard some data
loss_his = []
train_accu_his = []
val_accu_his = []
for epoch in range(num_epoch):
for t in range(step.shape[0]-1):
# calculate training loss
y_train_pred = model(X_train[step[t]:step[t+1],:,:])
loss = loss_fn(y_train_pred, y_train[step[t]:step[t+1]])
# backpropagation
model.zero_grad()
loss.backward()
# update parameters
optimizer.step()
# calculate predicted value for validation
y_val_pred = model(X_val)
# training loss
print('Epoch ', epoch, ', loss is ', loss.data.numpy())
_, y_pred = torch.max(y_train_pred,1)
loss_his.append(loss.data.numpy())
# training accuracy
train_accu = np.mean(y_pred.data.numpy() ==
y_train.data[step[t]:step[t+1]].numpy())
print('Training accuracy', train_accu)
train_accu_his.append(train_accu)
# validation accuracy
_, y_pred = torch.max(y_val_pred,1)
val_accu = np.mean(y_pred.data.numpy() == y_val.data.numpy())
print('Validation accuracy', val_accu, '\n')
val_accu_his.append(val_accu)
# +
# plot training and validation history
plt.subplot(2, 1, 1)
plt.plot(loss_his, 'o')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.subplot(2,1,2)
plt.plot(train_accu_his, '-o')
plt.plot(val_accu_his, '-o')
plt.legend(['train','val'], loc='upper left')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.show()
# +
# test set
y_test_pred = model(X_test)
_, y_pred = torch.max(y_test_pred,1)
test_accu = np.mean(y_pred.data.numpy() == y_test.data.numpy())
print('Test accuracy', test_accu, '\n')
# -
|
project_cnn_run_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: maowei
# language: python
# name: maowei
# ---
# +
import pandas as pd
import os
from datetime import datetime
import utils.data_utils as data_utils
from collections import defaultdict
LONG_TERM_ACTIONS = ['walking', 'eating', 'smoking', 'discussion']
ACTIONS = data_utils.define_actions('all')
ACTIONS.sort(key=len)
# +
def csv_iter():
for f in os.listdir('./checkpoint/test'):
try:
datetime.strptime(f[-23:-4], "%d-%m-%Y-%H:%M:%S")
if f.split('.')[-1] == 'csv':
yield f
except:
pass
def extend_df(df, long_term=True):
if long_term:
actions = LONG_TERM_ACTIONS
time_lens = [560, 1000]
else:
actions = ACTIONS
time_lens = [80, 160, 320, 400]
columns_mapping = defaultdict(list)
for time_len in time_lens:
for action_name in actions:
columns_mapping['3d' + str(time_len)].append(action_name + '3d' + str(time_len))
columns_mapping['usingfulltestset_3d' + str(time_len)].append('usingfulltestset_' + action_name + '3d' + str(time_len))
for key in columns_mapping:
df[key] = df[columns_mapping[key]].mean(axis=1)
return list(columns_mapping.keys())
def get_summary_df(merged):
cols = list(set([i[:-5] for i in merged.columns]))
new_cols = []
for col in cols:
col_name = [c for c in merged.columns if c.startswith(col)]
new_cols.append(col+'_mean')
new_cols.append(col+'_std')
merged[col+'_mean'] = merged[col_name].mean(axis=1)
merged[col+'_std'] = merged[col_name].std(axis=1)
return merged[new_cols]
# +
df_lst = []
mini_df_lst = []
for i, f in enumerate(csv_iter()):
df = pd.read_csv(os.path.join('checkpoint/test', "./", f))
df = df.sort_values(by='v_3d')
df_lst.append(df[extend_df(df, LONG_TERM_ACTIONS)].add_suffix('_run' + str(i)))
mini_df_lst.append(df[extend_df(df, LONG_TERM_ACTIONS)].head(1).add_suffix('_run' + str(i)).reset_index(drop=True))
merged = pd.concat(mini_df_lst, axis=1)
# -
get_summary_df(merged)
mini_df_lst[0]
mini_df_lst[1]
mini_df_lst[2]
mini_df_lst[3]
mini_df_lst[4]
|
analyse_csv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/konstin/ColabFold/blob/main/AlphaFold2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="G4yBrceuFbf3"
# #ColabFold: AlphaFold2 w/ MMseqs2
#
# <img src="https://raw.githubusercontent.com/sokrypton/ColabFold/main/.github/ColabFold_Marv_Logo_Small.png" height="256" align="right" style="height:256px">
#
# Easy to use AlphaFold2 protein structure [(Jumper et al. 2021)](https://www.nature.com/articles/s41586-021-03819-2) and complex [(Evans et al. 2021)](https://www.biorxiv.org/content/10.1101/2021.10.04.463034v1) prediction using multiple sequence alignments generated through MMseqs2. For details, refer to our manuscript:
#
# [<NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>. ColabFold - Making protein folding accessible to all.
# *bioRxiv*, 2021](https://www.biorxiv.org/content/10.1101/2021.08.15.456425v2)
#
# - This notebook provides basic functionality for **protein structure (Alphafold2)** and **complex prediction (Alphafold2-multimer)**. Advanced features such as recycles, sampling, ... ->[advanced notebook](https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/beta/AlphaFold2_advanced.ipynb).
#
# Old versions: [v1.0-alpha](https://colab.research.google.com/github/sokrypton/ColabFold/blob/v1.0-alpha/AlphaFold2.ipynb), [v1.1-permultimer](https://colab.research.google.com/github/sokrypton/ColabFold/blob/v1.1-premultimer/AlphaFold2.ipynb)
#
# For more details, see **<a href="#Instructions">bottom</a>** of the notebook and checkout the **[ColabFold GitHub](https://github.com/sokrypton/ColabFold)**.
# + id="kOblAo-xetgx" cellView="form"
#@title Input protein sequence, then hit `Runtime` -> `Run all`
from google.colab import files
import os.path
import re
import hashlib
def add_hash(x,y):
return x+"_"+hashlib.sha1(y.encode()).hexdigest()[:5]
query_sequence = 'PIAQIHILEGRSDEQKETLIREVSEAISRSLDAPLTSVRVIITEMAKGHFGIGGELASK' #@param {type:"string"}
#@markdown - Use `:` to specify inter-protein chainbreaks for **modeling complexes** (supports homo- and hetro-oligomers). For example **PI...SK:PI...SK** for a mono-dimer
# remove whitespaces
query_sequence = "".join(query_sequence.split())
jobname = 'test5' #@param {type:"string"}
# remove whitespaces
jobname = "".join(jobname.split())
jobname = re.sub(r'\W+', '', jobname)
jobname = add_hash(jobname, query_sequence)
with open(f"{jobname}.csv", "w") as text_file:
text_file.write(f"id,sequence\n{jobname},{query_sequence}")
queries_path=f"{jobname}.csv"
# number of models to use
#@markdown ---
#@markdown ### Advanced settings
msa_mode = "MMseqs2 (UniRef+Environmental)" #@param ["MMseqs2 (UniRef+Environmental)", "MMseqs2 (UniRef only)","single_sequence","custom"]
model_type = "auto" #@param ["auto", "AlphaFold2-ptm", "AlphaFold2-multimer"]
#@markdown - auto = protein structure prediction using "AlphaFold2-ptm" and complex prediction "AlphaFold-multimer". For complexes "AlphaFold-multimer" and "AlphaFold-ptm" can be used.
num_models = 5 #@param [1,2,3,4,5] {type:"raw"}
num_recycles = 3 #@param [1,3,6,12,24,48] {type:"raw"}
use_msa = True if msa_mode.startswith("MMseqs2") else False
use_env = True if msa_mode == "MMseqs2 (UniRef+Environmental)" else False
use_amber = False #@param {type:"boolean"}
use_templates = False #@param {type:"boolean"}
#@markdown ---
#@markdown ### Experimental options
save_to_google_drive = False #@param {type:"boolean"}
#@markdown ---
#@markdown Don't forget to hit `Runtime` -> `Run all` after updating the form.
with open(f"{jobname}.log", "w") as text_file:
text_file.write("num_models=%s\n" % num_models)
text_file.write("use_amber=%s\n" % use_amber)
text_file.write("use_msa=%s\n" % use_msa)
text_file.write("msa_mode=%s\n" % msa_mode)
text_file.write("use_templates=%s\n" % use_templates)
# decide which a3m to use
if use_msa:
a3m_file = f"{jobname}.a3m"
elif msa_mode == "custom":
a3m_file = f"{jobname}.custom.a3m"
if not os.path.isfile(a3m_file):
custom_msa_dict = files.upload()
custom_msa = list(custom_msa_dict.keys())[0]
header = 0
import fileinput
for line in fileinput.FileInput(custom_msa,inplace=1):
if line.startswith(">"):
header = header + 1
if not line.rstrip():
continue
if line.startswith(">") == False and header == 1:
query_sequence = line.rstrip()
print(line, end='')
os.rename(custom_msa, a3m_file)
queries_path=a3m_file
print(f"moving {custom_msa} to {a3m_file}")
else:
a3m_file = f"{jobname}.single_sequence.a3m"
with open(a3m_file, "w") as text_file:
text_file.write(">1\n%s" % query_sequence)
if save_to_google_drive:
from pydrive.drive import GoogleDrive
from pydrive.auth import GoogleAuth
from google.colab import auth
from oauth2client.client import GoogleCredentials
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
print("You are logged into Google Drive and are good to go!")
# + id="iccGdbe_Pmt9" pycharm={"name": "#%%\n"} cellView="form"
#@title Install dependencies
# %%bash -s $use_amber $use_msa $use_templates
set -e
USE_AMBER=$1
USE_MSA=$2
USE_TEMPLATES=$3
if [ ! -f COLABFOLD_READY ]; then
# install dependencies
pip install -q biopython dm-haiku ml-collections py3Dmol
# Trick for dev stage because otherwise pip won't install newer git versions
pip uninstall -y -q colabfold
pip install -q "colabfold[alphafold] @ git+https://github.com/konstin/ColabFold"
touch COLABFOLD_READY
fi
# download libraries for interfacing with MMseqs2 API
if [ ${USE_MSA} == "True" ] || [ ${USE_TEMPLATES} == "True" ]; then
if [ ! -f MMSEQ2_READY ]; then
apt-get -qq -y update 2>&1 1>/dev/null
apt-get -qq -y install jq curl zlib1g gawk 2>&1 1>/dev/null
touch MMSEQ2_READY
fi
fi
# setup conda
if [ ${USE_AMBER} == "True" ] || [ ${USE_TEMPLATES} == "True" ]; then
if [ ! -f CONDA_READY ]; then
wget -qnc https://repo.anaconda.com/miniconda/Miniconda3-latest-Linux-x86_64.sh
bash Miniconda3-latest-Linux-x86_64.sh -bfp /usr/local 2>&1 1>/dev/null
rm Miniconda3-latest-Linux-x86_64.sh
touch CONDA_READY
fi
fi
# setup template search
if [ ${USE_TEMPLATES} == "True" ] && [ ! -f HH_READY ]; then
conda install -y -q -c conda-forge -c bioconda kalign3=3.2.2 hhsuite=3.3.0 python=3.7 2>&1 1>/dev/null
touch HH_READY
fi
# setup openmm for amber refinement
if [ ${USE_AMBER} == "True" ] && [ ! -f AMBER_READY ]; then
conda install -y -q -c conda-forge openmm=7.5.1 python=3.7 pdbfixer 2>&1 1>/dev/null
wget -qnc https://raw.githubusercontent.com/deepmind/alphafold/main/docker/openmm.patch
(cd /usr/local/lib/python3.7/site-packages; patch -s -p0 < /content/openmm.patch)
wget -qnc https://git.scicore.unibas.ch/schwede/openstructure/-/raw/7102c63615b64735c4941278d92b554ec94415f8/modules/mol/alg/src/stereo_chemical_props.txt
touch AMBER_READY
fi
# + id="_sztQyz29DIC" cellView="form"
#@title Run Prediction
import sys
from colabfold.download import download_alphafold_params, default_data_dir
from colabfold.utils import setup_logging
from colabfold.batch import get_queries, run, set_model_type
from pathlib import Path
# For some reason we need that to get pdbfixer to import
if use_amber and '/usr/local/lib/python3.7/site-packages/' not in sys.path:
sys.path.insert(0, '/usr/local/lib/python3.7/site-packages/')
result_dir="."
setup_logging(Path(".").joinpath("log.txt"))
queries, is_complex = get_queries(queries_path)
model_type = set_model_type(is_complex, model_type)
download_alphafold_params(model_type, Path("."))
run(
queries=queries,
result_dir=result_dir,
use_templates=use_templates,
use_amber=use_amber,
msa_mode=msa_mode,
model_type=model_type,
num_models=num_models,
num_recycles=num_recycles,
model_order=[1, 2, 3, 4, 5],
is_complex=is_complex,
data_dir=Path("."),
keep_existing_results=False,
recompile_padding=1.0,
rank_mode="auto",
pair_mode="unpaired+paired",
stop_at_score=float(100),
)
# + id="KK7X9T44pWb7" cellView="form"
#@title Display 3D structure {run: "auto"}
import py3Dmol
import glob
import matplotlib.pyplot as plt
rank_num = 1 #@param ["1", "2", "3", "4", "5"] {type:"raw"}
color = "lDDT" #@param ["chain", "lDDT", "rainbow"]
show_sidechains = False #@param {type:"boolean"}
show_mainchains = False #@param {type:"boolean"}
jobname_prefix = ".custom" if msa_mode == "custom" else ""
if use_amber:
pdb_filename = f"{jobname}{jobname_prefix}_relaxed_model_*_rank_{rank_num}.pdb"
else:
pdb_filename = f"{jobname}{jobname_prefix}_unrelaxed_model_*_rank_{rank_num}.pdb"
pdb_file = glob.glob(pdb_filename)
def show_pdb(rank_num=1, show_sidechains=False, show_mainchains=False, color="lDDT"):
model_name = f"rank_{rank_num}"
if use_amber:
pdb_filename = f"{jobname}{jobname_prefix}_relaxed_{model_name}.pdb"
else:
pdb_filename = f"{jobname}{jobname_prefix}_unrelaxed_{model_name}.pdb"
view = py3Dmol.view(js='https://3dmol.org/build/3Dmol.js',)
view.addModel(open(pdb_file[0],'r').read(),'pdb')
if color == "lDDT":
view.setStyle({'cartoon': {'colorscheme': {'prop':'b','gradient': 'roygb','min':50,'max':90}}})
elif color == "rainbow":
view.setStyle({'cartoon': {'color':'spectrum'}})
elif color == "chain":
for n,chain,color in zip(range(homooligomer),list("ABCDEFGH"),
["lime","cyan","magenta","yellow","salmon","white","blue","orange"]):
view.setStyle({'chain':chain},{'cartoon': {'color':color}})
if show_sidechains:
BB = ['C','O','N']
view.addStyle({'and':[{'resn':["GLY","PRO"],'invert':True},{'atom':BB,'invert':True}]},
{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.addStyle({'and':[{'resn':"GLY"},{'atom':'CA'}]},
{'sphere':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.addStyle({'and':[{'resn':"PRO"},{'atom':['C','O'],'invert':True}]},
{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
if show_mainchains:
BB = ['C','O','N','CA']
view.addStyle({'atom':BB},{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.zoomTo()
return view
show_pdb(rank_num,show_sidechains, show_mainchains, color).show()
# + id="11l8k--10q0C" cellView="form"
#@title Plots {run: "auto"}
from IPython.display import display, HTML
import base64
from html import escape
# see: https://stackoverflow.com/a/53688522
def image_to_data_url(filename):
ext = filename.split('.')[-1]
prefix = f'data:image/{ext};base64,'
with open(filename, 'rb') as f:
img = f.read()
return prefix + base64.b64encode(img).decode('utf-8')
pae = image_to_data_url(f"{jobname}{jobname_prefix}_PAE.png")
cov = image_to_data_url(f"{jobname}{jobname_prefix}_coverage.png")
plddt = image_to_data_url(f"{jobname}{jobname_prefix}_plddt.png")
display(HTML(f"""
<style>
img {{
float:left;
}}
.full {{
max-width:100%;
}}
.half {{
max-width:50%;
}}
@media (max-width:640px) {{
.half {{
max-width:100%;
}}
}}
</style>
<div style="max-width:90%; padding:2em;">
<h1>Plots for {escape(jobname)}</h1>
<img src="{pae}" class="full" />
<img src="{cov}" class="half" />
<img src="{plddt}" class="half" />
</div>
"""))
# + id="33g5IIegij5R" cellView="form"
#@title Package and download results
#@markdown If you are having issues downloading the result archive, try disabling your adblocker and run this cell again. If that fails click on the little folder icon to the left, navigate to file: `jobname.result.zip`, right-click and select \"Download\" (see [screenshot](https://pbs.twimg.com/media/E6wRW2lWUAEOuoe?format=jpg&name=small)).
if msa_mode == "custom":
print("Don't forget to cite your custom MSA generation method.")
# !zip -FSr $jobname".result.zip" $jobname".log" $jobname".a3m" $jobname*"relaxed_model_"*".pdb" $jobname*"_coverage_lDDT.png" "cite.bibtex" $jobname*".png"
files.download(f"{jobname}.result.zip")
if save_to_google_drive == True and drive:
uploaded = drive.CreateFile({'title': f"{jobname}.result.zip"})
uploaded.SetContentFile(f"{jobname}.result.zip")
uploaded.Upload()
print(f"Uploaded {jobname}.result.zip to Google Drive with ID {uploaded.get('id')}")
# + [markdown] id="UGUBLzB3C6WN"
# # Instructions <a name="Instructions"></a>
# **Quick start**
# 1. Paste your protein sequence(s) in the input field.
# 2. Press "Runtime" -> "Run all".
# 3. The pipeline consists of 5 steps. The currently running steps is indicated by a circle with a stop sign next to it.
#
# **Result zip file contents**
#
# 1. PDB formatted structures sorted by avg. pIDDT and complexes are sorted by pTMscore. (unrelaxed and relaxed if `use_amber` is enabled).
# 2. Plots of the model quality.
# 3. Plots of the MSA coverage.
# 4. Parameter log file.
# 5. A3M formatted input MSA.
# 6. BibTeX file with citations for all used tools and databases.
#
# At the end of the job a download modal box will pop up with a `jobname.result.zip` file. Additionally, if the `save_to_google_drive` option was selected, the `jobname.result.zip` will be uploaded to your Google Drive.
#
# **MSA generation for complexes**
#
# For the complex prediction we use unpaired and paired MSAs. Unpaired MSA is generated the same way as for the protein structures prediction by searching the UniRef100 and environmental sequences three iterations each.
#
# The paired MSA is generated by searching the UniRef100 database and pairing the best hits sharing the same NCBI taxonomical identifier (=species or sub-species). We only pair sequences if all of the query sequences are present for the respective taxonomical identifier.
#
# **Using a custom MSA as input**
#
# To predict the structure with a custom MSA (A3M formatted): (1) Change the msa_mode: to "custom", (2) Wait for an upload box to appear at the end of the "Input Protein ..." box. Upload your A3M. The first fasta entry of the A3M must be the query sequence without gaps.
#
# As an alternative for MSA generation the [HHblits Toolkit server](https://toolkit.tuebingen.mpg.de/tools/hhblits) can be used. After submitting your query, click "Query Template MSA" -> "Download Full A3M". Download the A3M file and upload it in this notebook.
#
# **Comparision to the full AlphaFold2 and Alphafold2 colab**
#
# This notebook replaces the homology detection and MSA pairing of AlphaFold2 with MMseqs2. For a comparision against the [AlphaFold2 Colab](https://colab.research.google.com/github/deepmind/alphafold/blob/main/notebooks/AlphaFold.ipynb) and the full [AlphaFold2](https://github.com/deepmind/alphafold) system read our [preprint](https://www.biorxiv.org/content/10.1101/2021.08.15.456425v1).
#
# **Troubleshooting**
# * Check that the runtime type is set to GPU at "Runtime" -> "Change runtime type".
# * Try to restart the session "Runtime" -> "Factory reset runtime".
# * Check your input sequence.
#
# **Known issues**
# * Google Colab assigns different types of GPUs with varying amount of memory. Some might not have enough memory to predict the structure for a long sequence.
# * Your browser can block the pop-up for downloading the result file. You can choose the `save_to_google_drive` option to upload to Google Drive instead or manually download the result file: Click on the little folder icon to the left, navigate to file: `jobname.result.zip`, right-click and select \"Download\" (see [screenshot](https://pbs.twimg.com/media/E6wRW2lWUAEOuoe?format=jpg&name=small)).
#
# **Limitations**
# * Computing resources: Our MMseqs2 API can handle ~20-50k requests per day.
# * MSAs: MMseqs2 is very precise and sensitive but might find less hits compared to HHblits/HMMer searched against BFD or Mgnify.
# * We recommend to additionally use the full [AlphaFold2 pipeline](https://github.com/deepmind/alphafold).
#
# **Description of the plots**
# * **Number of sequences per position** - We want to see at least 30 sequences per position, for best performance, ideally 100 sequences.
# * **Predicted lDDT per position** - model confidence (out of 100) at each position. The higher the better.
# * **Predicted Alignment Error** - For homooligomers, this could be a useful metric to assess how confident the model is about the interface. The lower the better.
#
# **Bugs**
# - If you encounter any bugs, please report the issue to https://github.com/sokrypton/ColabFold/issues
#
#
# **Acknowledgments**
# - We thank the AlphaFold team for developing an excellent model and open sourcing the software.
#
# - [Söding Lab](https://www.mpibpc.mpg.de/soeding) for providing the computational resources for the MMseqs2 server
#
# - <NAME> for helping to bechmark the ColabFold's Alphafold-multimer support
#
# - [<NAME>](https://github.com/dkoes) for his awesome [py3Dmol](https://3dmol.csb.pitt.edu/) plugin, without whom these notebooks would be quite boring!
#
# - Do-Yoon Kim for creating the ColabFold logo.
#
# - A colab by <NAME> ([@sokrypton](https://twitter.com/sokrypton)), <NAME> ([@milot_mirdita](https://twitter.com/milot_mirdita)) and <NAME> ([@thesteinegger](https://twitter.com/thesteinegger)).
#
|
AlphaFold2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''py38-hotel'': conda)'
# name: python388jvsc74a57bd046e16dc5765237a024e1f11c34a9a67322fdcdedd6cc2d0be7272ca72acaf593
# ---
# +
import pandas as pd
df = pd.read_csv('data/train.csv')
# -
df.hotel_id.value_counts()[:50]
|
hotels.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SARIMAX: Introduction
# This notebook replicates examples from the Stata ARIMA time series estimation and postestimation documentation.
#
# First, we replicate the four estimation examples http://www.stata.com/manuals13/tsarima.pdf:
#
# 1. ARIMA(1,1,1) model on the U.S. Wholesale Price Index (WPI) dataset.
# 2. Variation of example 1 which adds an MA(4) term to the ARIMA(1,1,1) specification to allow for an additive seasonal effect.
# 3. ARIMA(2,1,0) x (1,1,0,12) model of monthly airline data. This example allows a multiplicative seasonal effect.
# 4. ARMA(1,1) model with exogenous regressors; describes consumption as an autoregressive process on which also the money supply is assumed to be an explanatory variable.
#
# Second, we demonstrate postestimation capabilities to replicate http://www.stata.com/manuals13/tsarimapostestimation.pdf. The model from example 4 is used to demonstrate:
#
# 1. One-step-ahead in-sample prediction
# 2. n-step-ahead out-of-sample forecasting
# 3. n-step-ahead in-sample dynamic prediction
# %matplotlib inline
import numpy as np
import pandas as pd
from scipy.stats import norm
import statsmodels.api as sm
import matplotlib.pyplot as plt
from datetime import datetime
import requests
from io import BytesIO
# Register converters to avoid warnings
pd.plotting.register_matplotlib_converters()
plt.rc("figure", figsize=(16,8))
plt.rc("font", size=14)
# ### ARIMA Example 1: Arima
#
# As can be seen in the graphs from Example 2, the Wholesale price index (WPI) is growing over time (i.e. is not stationary). Therefore an ARMA model is not a good specification. In this first example, we consider a model where the original time series is assumed to be integrated of order 1, so that the difference is assumed to be stationary, and fit a model with one autoregressive lag and one moving average lag, as well as an intercept term.
#
# The postulated data process is then:
#
# $$
# \Delta y_t = c + \phi_1 \Delta y_{t-1} + \theta_1 \epsilon_{t-1} + \epsilon_{t}
# $$
#
# where $c$ is the intercept of the ARMA model, $\Delta$ is the first-difference operator, and we assume $\epsilon_{t} \sim N(0, \sigma^2)$. This can be rewritten to emphasize lag polynomials as (this will be useful in example 2, below):
#
# $$
# (1 - \phi_1 L ) \Delta y_t = c + (1 + \theta_1 L) \epsilon_{t}
# $$
#
# where $L$ is the lag operator.
#
# Notice that one difference between the Stata output and the output below is that Stata estimates the following model:
#
# $$
# (\Delta y_t - \beta_0) = \phi_1 ( \Delta y_{t-1} - \beta_0) + \theta_1 \epsilon_{t-1} + \epsilon_{t}
# $$
#
# where $\beta_0$ is the mean of the process $y_t$. This model is equivalent to the one estimated in the statsmodels SARIMAX class, but the interpretation is different. To see the equivalence, note that:
#
# $$
# (\Delta y_t - \beta_0) = \phi_1 ( \Delta y_{t-1} - \beta_0) + \theta_1 \epsilon_{t-1} + \epsilon_{t} \\
# \Delta y_t = (1 - \phi_1) \beta_0 + \phi_1 \Delta y_{t-1} + \theta_1 \epsilon_{t-1} + \epsilon_{t}
# $$
#
# so that $c = (1 - \phi_1) \beta_0$.
# +
# Dataset
wpi1 = requests.get('https://www.stata-press.com/data/r12/wpi1.dta').content
data = pd.read_stata(BytesIO(wpi1))
data.index = data.t
# Set the frequency
data.index.freq="QS-OCT"
# Fit the model
mod = sm.tsa.statespace.SARIMAX(data['wpi'], trend='c', order=(1,1,1))
res = mod.fit(disp=False)
print(res.summary())
# -
# Thus the maximum likelihood estimates imply that for the process above, we have:
#
# $$
# \Delta y_t = 0.0943 + 0.8742 \Delta y_{t-1} - 0.4120 \epsilon_{t-1} + \epsilon_{t}
# $$
#
# where $\epsilon_{t} \sim N(0, 0.5257)$. Finally, recall that $c = (1 - \phi_1) \beta_0$, and here $c = 0.0943$ and $\phi_1 = 0.8742$. To compare with the output from Stata, we could calculate the mean:
#
# $$\beta_0 = \frac{c}{1 - \phi_1} = \frac{0.0943}{1 - 0.8742} = 0.7496$$
#
# **Note**: This value is virtually identical to the value in the Stata documentation, $\beta_0 = 0.7498$. The slight difference is likely down to rounding and subtle differences in stopping criterion of the numerical optimizers used.
# ### ARIMA Example 2: Arima with additive seasonal effects
#
# This model is an extension of that from example 1. Here the data is assumed to follow the process:
#
# $$
# \Delta y_t = c + \phi_1 \Delta y_{t-1} + \theta_1 \epsilon_{t-1} + \theta_4 \epsilon_{t-4} + \epsilon_{t}
# $$
#
# The new part of this model is that there is allowed to be a annual seasonal effect (it is annual even though the periodicity is 4 because the dataset is quarterly). The second difference is that this model uses the log of the data rather than the level.
#
# Before estimating the dataset, graphs showing:
#
# 1. The time series (in logs)
# 2. The first difference of the time series (in logs)
# 3. The autocorrelation function
# 4. The partial autocorrelation function.
#
# From the first two graphs, we note that the original time series does not appear to be stationary, whereas the first-difference does. This supports either estimating an ARMA model on the first-difference of the data, or estimating an ARIMA model with 1 order of integration (recall that we are taking the latter approach). The last two graphs support the use of an ARMA(1,1,1) model.
# +
# Dataset
data = pd.read_stata(BytesIO(wpi1))
data.index = data.t
data.index.freq="QS-OCT"
data['ln_wpi'] = np.log(data['wpi'])
data['D.ln_wpi'] = data['ln_wpi'].diff()
# +
# Graph data
fig, axes = plt.subplots(1, 2, figsize=(15,4))
# Levels
axes[0].plot(data.index._mpl_repr(), data['wpi'], '-')
axes[0].set(title='US Wholesale Price Index')
# Log difference
axes[1].plot(data.index._mpl_repr(), data['D.ln_wpi'], '-')
axes[1].hlines(0, data.index[0], data.index[-1], 'r')
axes[1].set(title='US Wholesale Price Index - difference of logs');
# +
# Graph data
fig, axes = plt.subplots(1, 2, figsize=(15,4))
fig = sm.graphics.tsa.plot_acf(data.iloc[1:]['D.ln_wpi'], lags=40, ax=axes[0])
fig = sm.graphics.tsa.plot_pacf(data.iloc[1:]['D.ln_wpi'], lags=40, ax=axes[1])
# -
# To understand how to specify this model in statsmodels, first recall that from example 1 we used the following code to specify the ARIMA(1,1,1) model:
#
# ```python
# mod = sm.tsa.statespace.SARIMAX(data['wpi'], trend='c', order=(1,1,1))
# ```
#
# The `order` argument is a tuple of the form `(AR specification, Integration order, MA specification)`. The integration order must be an integer (for example, here we assumed one order of integration, so it was specified as 1. In a pure ARMA model where the underlying data is already stationary, it would be 0).
#
# For the AR specification and MA specification components, there are two possibilities. The first is to specify the **maximum degree** of the corresponding lag polynomial, in which case the component is an integer. For example, if we wanted to specify an ARIMA(1,1,4) process, we would use:
#
# ```python
# mod = sm.tsa.statespace.SARIMAX(data['wpi'], trend='c', order=(1,1,4))
# ```
#
# and the corresponding data process would be:
#
# $$
# y_t = c + \phi_1 y_{t-1} + \theta_1 \epsilon_{t-1} + \theta_2 \epsilon_{t-2} + \theta_3 \epsilon_{t-3} + \theta_4 \epsilon_{t-4} + \epsilon_{t}
# $$
#
# or
#
# $$
# (1 - \phi_1 L)\Delta y_t = c + (1 + \theta_1 L + \theta_2 L^2 + \theta_3 L^3 + \theta_4 L^4) \epsilon_{t}
# $$
#
# When the specification parameter is given as a maximum degree of the lag polynomial, it implies that all polynomial terms up to that degree are included. Notice that this is *not* the model we want to use, because it would include terms for $\epsilon_{t-2}$ and $\epsilon_{t-3}$, which we do not want here.
#
# What we want is a polynomial that has terms for the 1st and 4th degrees, but leaves out the 2nd and 3rd terms. To do that, we need to provide a tuple for the specification parameter, where the tuple describes **the lag polynomial itself**. In particular, here we would want to use:
#
# ```python
# ar = 1 # this is the maximum degree specification
# ma = (1,0,0,1) # this is the lag polynomial specification
# mod = sm.tsa.statespace.SARIMAX(data['wpi'], trend='c', order=(ar,1,ma)))
# ```
#
# This gives the following form for the process of the data:
#
# $$
# \Delta y_t = c + \phi_1 \Delta y_{t-1} + \theta_1 \epsilon_{t-1} + \theta_4 \epsilon_{t-4} + \epsilon_{t} \\
# (1 - \phi_1 L)\Delta y_t = c + (1 + \theta_1 L + \theta_4 L^4) \epsilon_{t}
# $$
#
# which is what we want.
# Fit the model
mod = sm.tsa.statespace.SARIMAX(data['ln_wpi'], trend='c', order=(1,1,(1,0,0,1)))
res = mod.fit(disp=False)
print(res.summary())
# ### ARIMA Example 3: Airline Model
#
# In the previous example, we included a seasonal effect in an *additive* way, meaning that we added a term allowing the process to depend on the 4th MA lag. It may be instead that we want to model a seasonal effect in a multiplicative way. We often write the model then as an ARIMA $(p,d,q) \times (P,D,Q)_s$, where the lowercase letters indicate the specification for the non-seasonal component, and the uppercase letters indicate the specification for the seasonal component; $s$ is the periodicity of the seasons (e.g. it is often 4 for quarterly data or 12 for monthly data). The data process can be written generically as:
#
# $$
# \phi_p (L) \tilde \phi_P (L^s) \Delta^d \Delta_s^D y_t = A(t) + \theta_q (L) \tilde \theta_Q (L^s) \epsilon_t
# $$
#
# where:
#
# - $\phi_p (L)$ is the non-seasonal autoregressive lag polynomial
# - $\tilde \phi_P (L^s)$ is the seasonal autoregressive lag polynomial
# - $\Delta^d \Delta_s^D y_t$ is the time series, differenced $d$ times, and seasonally differenced $D$ times.
# - $A(t)$ is the trend polynomial (including the intercept)
# - $\theta_q (L)$ is the non-seasonal moving average lag polynomial
# - $\tilde \theta_Q (L^s)$ is the seasonal moving average lag polynomial
#
# sometimes we rewrite this as:
#
# $$
# \phi_p (L) \tilde \phi_P (L^s) y_t^* = A(t) + \theta_q (L) \tilde \theta_Q (L^s) \epsilon_t
# $$
#
# where $y_t^* = \Delta^d \Delta_s^D y_t$. This emphasizes that just as in the simple case, after we take differences (here both non-seasonal and seasonal) to make the data stationary, the resulting model is just an ARMA model.
#
# As an example, consider the airline model ARIMA $(2,1,0) \times (1,1,0)_{12}$, with an intercept. The data process can be written in the form above as:
#
# $$
# (1 - \phi_1 L - \phi_2 L^2) (1 - \tilde \phi_1 L^{12}) \Delta \Delta_{12} y_t = c + \epsilon_t
# $$
#
# Here, we have:
#
# - $\phi_p (L) = (1 - \phi_1 L - \phi_2 L^2)$
# - $\tilde \phi_P (L^s) = (1 - \phi_1 L^12)$
# - $d = 1, D = 1, s=12$ indicating that $y_t^*$ is derived from $y_t$ by taking first-differences and then taking 12-th differences.
# - $A(t) = c$ is the *constant* trend polynomial (i.e. just an intercept)
# - $\theta_q (L) = \tilde \theta_Q (L^s) = 1$ (i.e. there is no moving average effect)
#
# It may still be confusing to see the two lag polynomials in front of the time-series variable, but notice that we can multiply the lag polynomials together to get the following model:
#
# $$
# (1 - \phi_1 L - \phi_2 L^2 - \tilde \phi_1 L^{12} + \phi_1 \tilde \phi_1 L^{13} + \phi_2 \tilde \phi_1 L^{14} ) y_t^* = c + \epsilon_t
# $$
#
# which can be rewritten as:
#
# $$
# y_t^* = c + \phi_1 y_{t-1}^* + \phi_2 y_{t-2}^* + \tilde \phi_1 y_{t-12}^* - \phi_1 \tilde \phi_1 y_{t-13}^* - \phi_2 \tilde \phi_1 y_{t-14}^* + \epsilon_t
# $$
#
# This is similar to the additively seasonal model from example 2, but the coefficients in front of the autoregressive lags are actually combinations of the underlying seasonal and non-seasonal parameters.
#
# Specifying the model in statsmodels is done simply by adding the `seasonal_order` argument, which accepts a tuple of the form `(Seasonal AR specification, Seasonal Integration order, Seasonal MA, Seasonal periodicity)`. The seasonal AR and MA specifications, as before, can be expressed as a maximum polynomial degree or as the lag polynomial itself. Seasonal periodicity is an integer.
#
# For the airline model ARIMA $(2,1,0) \times (1,1,0)_{12}$ with an intercept, the command is:
#
# ```python
# mod = sm.tsa.statespace.SARIMAX(data['lnair'], order=(2,1,0), seasonal_order=(1,1,0,12))
# ```
# +
# Dataset
air2 = requests.get('https://www.stata-press.com/data/r12/air2.dta').content
data = pd.read_stata(BytesIO(air2))
data.index = pd.date_range(start=datetime(data.time[0], 1, 1), periods=len(data), freq='MS')
data['lnair'] = np.log(data['air'])
# Fit the model
mod = sm.tsa.statespace.SARIMAX(data['lnair'], order=(2,1,0), seasonal_order=(1,1,0,12), simple_differencing=True)
res = mod.fit(disp=False)
print(res.summary())
# -
# Notice that here we used an additional argument `simple_differencing=True`. This controls how the order of integration is handled in ARIMA models. If `simple_differencing=True`, then the time series provided as `endog` is literally differenced and an ARMA model is fit to the resulting new time series. This implies that a number of initial periods are lost to the differencing process, however it may be necessary either to compare results to other packages (e.g. Stata's `arima` always uses simple differencing) or if the seasonal periodicity is large.
#
# The default is `simple_differencing=False`, in which case the integration component is implemented as part of the state space formulation, and all of the original data can be used in estimation.
# ### ARIMA Example 4: ARMAX (Friedman)
#
# This model demonstrates the use of explanatory variables (the X part of ARMAX). When exogenous regressors are included, the SARIMAX module uses the concept of "regression with SARIMA errors" (see http://robjhyndman.com/hyndsight/arimax/ for details of regression with ARIMA errors versus alternative specifications), so that the model is specified as:
#
# $$
# y_t = \beta_t x_t + u_t \\
# \phi_p (L) \tilde \phi_P (L^s) \Delta^d \Delta_s^D u_t = A(t) +
# \theta_q (L) \tilde \theta_Q (L^s) \epsilon_t
# $$
#
# Notice that the first equation is just a linear regression, and the second equation just describes the process followed by the error component as SARIMA (as was described in example 3). One reason for this specification is that the estimated parameters have their natural interpretations.
#
# This specification nests many simpler specifications. For example, regression with AR(2) errors is:
#
# $$
# y_t = \beta_t x_t + u_t \\
# (1 - \phi_1 L - \phi_2 L^2) u_t = A(t) + \epsilon_t
# $$
#
# The model considered in this example is regression with ARMA(1,1) errors. The process is then written:
#
# $$
# \text{consump}_t = \beta_0 + \beta_1 \text{m2}_t + u_t \\
# (1 - \phi_1 L) u_t = (1 - \theta_1 L) \epsilon_t
# $$
#
# Notice that $\beta_0$ is, as described in example 1 above, *not* the same thing as an intercept specified by `trend='c'`. Whereas in the examples above we estimated the intercept of the model via the trend polynomial, here, we demonstrate how to estimate $\beta_0$ itself by adding a constant to the exogenous dataset. In the output, the $beta_0$ is called `const`, whereas above the intercept $c$ was called `intercept` in the output.
# +
# Dataset
friedman2 = requests.get('https://www.stata-press.com/data/r12/friedman2.dta').content
data = pd.read_stata(BytesIO(friedman2))
data.index = data.time
data.index.freq = "QS-OCT"
# Variables
endog = data.loc['1959':'1981', 'consump']
exog = sm.add_constant(data.loc['1959':'1981', 'm2'])
# Fit the model
mod = sm.tsa.statespace.SARIMAX(endog, exog, order=(1,0,1))
res = mod.fit(disp=False)
print(res.summary())
# -
# ### ARIMA Postestimation: Example 1 - Dynamic Forecasting
#
# Here we describe some of the post-estimation capabilities of statsmodels' SARIMAX.
#
# First, using the model from example, we estimate the parameters using data that *excludes the last few observations* (this is a little artificial as an example, but it allows considering performance of out-of-sample forecasting and facilitates comparison to Stata's documentation).
# +
# Dataset
raw = pd.read_stata(BytesIO(friedman2))
raw.index = raw.time
raw.index.freq = "QS-OCT"
data = raw.loc[:'1981']
# Variables
endog = data.loc['1959':, 'consump']
exog = sm.add_constant(data.loc['1959':, 'm2'])
nobs = endog.shape[0]
# Fit the model
mod = sm.tsa.statespace.SARIMAX(endog.loc[:'1978-01-01'], exog=exog.loc[:'1978-01-01'], order=(1,0,1))
fit_res = mod.fit(disp=False, maxiter=250)
print(fit_res.summary())
# -
# Next, we want to get results for the full dataset but using the estimated parameters (on a subset of the data).
mod = sm.tsa.statespace.SARIMAX(endog, exog=exog, order=(1,0,1))
res = mod.filter(fit_res.params)
# The `predict` command is first applied here to get in-sample predictions. We use the `full_results=True` argument to allow us to calculate confidence intervals (the default output of `predict` is just the predicted values).
#
# With no other arguments, `predict` returns the one-step-ahead in-sample predictions for the entire sample.
# In-sample one-step-ahead predictions
predict = res.get_prediction()
predict_ci = predict.conf_int()
# We can also get *dynamic predictions*. One-step-ahead prediction uses the true values of the endogenous values at each step to predict the next in-sample value. Dynamic predictions use one-step-ahead prediction up to some point in the dataset (specified by the `dynamic` argument); after that, the previous *predicted* endogenous values are used in place of the true endogenous values for each new predicted element.
#
# The `dynamic` argument is specified to be an *offset* relative to the `start` argument. If `start` is not specified, it is assumed to be `0`.
#
# Here we perform dynamic prediction starting in the first quarter of 1978.
# Dynamic predictions
predict_dy = res.get_prediction(dynamic='1978-01-01')
predict_dy_ci = predict_dy.conf_int()
# We can graph the one-step-ahead and dynamic predictions (and the corresponding confidence intervals) to see their relative performance. Notice that up to the point where dynamic prediction begins (1978:Q1), the two are the same.
# +
# Graph
fig, ax = plt.subplots(figsize=(9,4))
npre = 4
ax.set(title='Personal consumption', xlabel='Date', ylabel='Billions of dollars')
# Plot data points
data.loc['1977-07-01':, 'consump'].plot(ax=ax, style='o', label='Observed')
# Plot predictions
predict.predicted_mean.loc['1977-07-01':].plot(ax=ax, style='r--', label='One-step-ahead forecast')
ci = predict_ci.loc['1977-07-01':]
ax.fill_between(ci.index, ci.iloc[:,0], ci.iloc[:,1], color='r', alpha=0.1)
predict_dy.predicted_mean.loc['1977-07-01':].plot(ax=ax, style='g', label='Dynamic forecast (1978)')
ci = predict_dy_ci.loc['1977-07-01':]
ax.fill_between(ci.index, ci.iloc[:,0], ci.iloc[:,1], color='g', alpha=0.1)
legend = ax.legend(loc='lower right')
# -
# Finally, graph the prediction *error*. It is obvious that, as one would suspect, one-step-ahead prediction is considerably better.
# +
# Prediction error
# Graph
fig, ax = plt.subplots(figsize=(9,4))
npre = 4
ax.set(title='Forecast error', xlabel='Date', ylabel='Forecast - Actual')
# In-sample one-step-ahead predictions and 95% confidence intervals
predict_error = predict.predicted_mean - endog
predict_error.loc['1977-10-01':].plot(ax=ax, label='One-step-ahead forecast')
ci = predict_ci.loc['1977-10-01':].copy()
ci.iloc[:,0] -= endog.loc['1977-10-01':]
ci.iloc[:,1] -= endog.loc['1977-10-01':]
ax.fill_between(ci.index, ci.iloc[:,0], ci.iloc[:,1], alpha=0.1)
# Dynamic predictions and 95% confidence intervals
predict_dy_error = predict_dy.predicted_mean - endog
predict_dy_error.loc['1977-10-01':].plot(ax=ax, style='r', label='Dynamic forecast (1978)')
ci = predict_dy_ci.loc['1977-10-01':].copy()
ci.iloc[:,0] -= endog.loc['1977-10-01':]
ci.iloc[:,1] -= endog.loc['1977-10-01':]
ax.fill_between(ci.index, ci.iloc[:,0], ci.iloc[:,1], color='r', alpha=0.1)
legend = ax.legend(loc='lower left');
legend.get_frame().set_facecolor('w')
|
examples/notebooks/statespace_sarimax_stata.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/krakowiakpawel9/machine-learning-bootcamp/blob/master/unsupervised/03_association_rules/02_apriori.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="C80St0V-_VIi" colab_type="text"
# ### scikit-learn
# Strona biblioteki: [https://scikit-learn.org](https://scikit-learn.org)
#
# Dokumentacja/User Guide: [https://scikit-learn.org/stable/user_guide.html](https://scikit-learn.org/stable/user_guide.html)
#
# Podstawowa biblioteka do uczenia maszynowego w języku Python.
#
# Aby zainstalować bibliotekę scikit-learn, użyj polecenia poniżej:
# ```
# # # !pip install scikit-learn
# ```
# Aby zaktualizować do najnowszej wersji bibliotekę scikit-learn, użyj polecenia poniżej:
# ```
# # # !pip install --upgrade scikit-learn
# ```
# Kurs stworzony w oparciu o wersję `0.22.1`
#
# ### Spis treści:
# 1. [Import bibliotek](#0)
# 2. [Załadownaie danych](#1)
# 3. [Przygotowanie danych](#2)
# 4. [Kodowanie transakcji](#3)
# 5. [Algorytm Apriori](#4)
#
#
#
# + [markdown] id="dCogYvFX_YD4" colab_type="text"
# ### <a name='0'></a> Import bibliotek
# + id="h1UzDFRtN3rJ" colab_type="code" colab={}
import pandas as pd
pd.set_option('display.float_format', lambda x: f'{x:.2f}')
# + [markdown] id="Gfy_mLSlSQZj" colab_type="text"
# ### <a name='1'></a> Załadownaie danych
# + id="iMjFzTjHhiOP" colab_type="code" outputId="b9ea4911-0b52-40ad-c3d5-f18911ecb47d" colab={"base_uri": "https://localhost:8080/", "height": 394}
# !wget https://storage.googleapis.com/esmartdata-courses-files/ml-course/products.csv
# !wget https://storage.googleapis.com/esmartdata-courses-files/ml-course/orders.csv
# + id="MeLHR8d-Orh4" colab_type="code" outputId="3b848fb1-d9fe-4bc9-a477-491b8afb2622" colab={"base_uri": "https://localhost:8080/", "height": 203}
products = pd.read_csv('products.csv', usecols=['product_id', 'product_name'])
products.head()
# + id="GjtwaJ8COwLt" colab_type="code" outputId="d1b0956e-0dbe-423e-d6f8-d28fa20a32e5" colab={"base_uri": "https://localhost:8080/", "height": 203}
orders = pd.read_csv('orders.csv', usecols=['order_id', 'product_id'])
orders.head()
# + [markdown] id="BU5EtQt9Sai_" colab_type="text"
# ### <a name='2'></a> Przygotowanie danych
# + id="QuhRw60QPJxp" colab_type="code" outputId="2e5940bf-53ce-4b43-c6b8-9ce9b613daca" colab={"base_uri": "https://localhost:8080/", "height": 203}
data = pd.merge(orders, products, how='inner', on='product_id', sort=True)
data = data.sort_values(by='order_id')
data.head()
# + id="DCdXwfd4PmPE" colab_type="code" outputId="aa2bebb4-8e91-4536-c209-f0835954e47e" colab={"base_uri": "https://localhost:8080/", "height": 295}
data.describe()
# + id="BwozcaSZR41L" colab_type="code" outputId="16df1332-27ca-430e-e6e9-3d9ba624d9e9" colab={"base_uri": "https://localhost:8080/", "height": 223}
# rozkład produktów
data['product_name'].value_counts()
# + id="k_VXYgJ_S-Ox" colab_type="code" outputId="dbf6042a-b78c-4317-8bf6-04b40f33d77b" colab={"base_uri": "https://localhost:8080/", "height": 34}
# liczba transakcji
data['order_id'].nunique()
# + id="MoqM_XDBTg4g" colab_type="code" outputId="92816132-4199-45f4-871e-02936a8c6563" colab={"base_uri": "https://localhost:8080/", "height": 240}
transactions = data.groupby(by='order_id')['product_name'].apply(lambda name: ','.join(name))
transactions
# + id="w7OSUdo4UDDC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 240} outputId="c90f576a-866a-4293-ce53-5f217cac15a3"
transactions = transactions.str.split(',')
transactions
# + [markdown] id="6z-LfUIPShPp" colab_type="text"
# ### <a name='3'></a> Kodowanie transakcji
# + id="Cy9FhFYUXLlz" colab_type="code" outputId="17c7b4f8-86a2-452c-c255-2eb56eed352c" colab={"base_uri": "https://localhost:8080/", "height": 51}
from mlxtend.preprocessing import TransactionEncoder
encoder = TransactionEncoder()
encoder.fit(transactions)
transactions_encoded = encoder.transform(transactions, sparse=True)
transactions_encoded
# + id="z9uIysohYm7S" colab_type="code" outputId="7ede8ae4-b6ac-4bf2-d740-bcdc9a1c7378" colab={"base_uri": "https://localhost:8080/", "height": 608}
transactions_encoded_df = pd.DataFrame(transactions_encoded.toarray(), columns=encoder.columns_)
transactions_encoded_df
# + [markdown] id="fC3HKl0GSsV4" colab_type="text"
# ### <a name='4'></a> Algorytm Apriori
# + id="Yw3mMi8JX0gL" colab_type="code" outputId="d3169c8b-3550-450f-97ce-f04dbaff6d48" colab={"base_uri": "https://localhost:8080/", "height": 357}
from mlxtend.frequent_patterns import apriori, association_rules
supports = apriori(transactions_encoded_df, min_support=0.01, use_colnames=True, n_jobs=-1)
supports = supports.sort_values(by='support', ascending=False)
supports.head(10)
# + id="w9YR0_x-YeFo" colab_type="code" outputId="0e2a0119-311d-49dc-913f-caaeb09a2d5a" colab={"base_uri": "https://localhost:8080/", "height": 511}
rules = association_rules(supports, metric='confidence', min_threshold=0)
rules = rules.iloc[:, [0, 1, 4, 5, 6]]
rules = rules.sort_values(by='lift', ascending=False)
rules.head(15)
# + id="bni0nMj7fHF1" colab_type="code" colab={}
|
unsupervised/03_association_rules/02_apriori.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TO-DO LIST
# - Label Smoothing
# - https://www.kaggle.com/chocozzz/train-cassava-starter-using-label-smoothing
# - https://www.kaggle.com/c/siim-isic-melanoma-classification/discussion/173733
#
# - Class Imbalance
#
# - SWA / SWAG
#
# - Augmentation
# - https://www.kaggle.com/sachinprabhu/pytorch-resnet50-snapmix-train-pipeline
import os
print(os.listdir("./input/"))
# + papermill={"duration": 0.664524, "end_time": "2020-11-23T13:32:47.332411", "exception": false, "start_time": "2020-11-23T13:32:46.667887", "status": "completed"} tags=[]
package_paths = [
'./input/pytorch-image-models/pytorch-image-models-master', #'../input/efficientnet-pytorch-07/efficientnet_pytorch-0.7.0'
'./input/pytorch-gradual-warmup-lr-master'
]
import sys;
for pth in package_paths:
sys.path.append(pth)
# from warmup_scheduler import GradualWarmupScheduler
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 2.173722, "end_time": "2020-11-23T13:32:49.521795", "exception": false, "start_time": "2020-11-23T13:32:47.348073", "status": "completed"} tags=[]
from glob import glob
from sklearn.model_selection import GroupKFold, StratifiedKFold
import cv2
from skimage import io
import torch
from torch import nn
import os
from datetime import datetime
import time
import random
import cv2
import torchvision
from torchvision import transforms
import pandas as pd
import numpy as np
from tqdm import tqdm
import matplotlib.pyplot as plt
from torch.utils.data import Dataset,DataLoader
from torch.utils.data.sampler import SequentialSampler, RandomSampler
from torch.cuda.amp import autocast, GradScaler
from torch.nn.modules.loss import _WeightedLoss
import torch.nn.functional as F
import timm
from adamp import AdamP
import sklearn
import warnings
import joblib
from sklearn.metrics import roc_auc_score, log_loss
from sklearn import metrics
import warnings
import cv2
#from efficientnet_pytorch import EfficientNet
from scipy.ndimage.interpolation import zoom
##SWA
from torch.optim.swa_utils import AveragedModel, SWALR, update_bn
from torch.optim.lr_scheduler import CosineAnnealingLR
# + papermill={"duration": 0.026635, "end_time": "2020-11-23T13:32:49.570638", "exception": false, "start_time": "2020-11-23T13:32:49.544003", "status": "completed"} tags=[]
CFG = {
'fold_num': 5,
'seed': 719,
'model_arch': 'tf_efficientnet_b4_ns',
'img_size': 512,
'epochs': 7,
'train_bs': 9,
'valid_bs': 16,
'T_0': 10,
'lr': 4e-4,
'min_lr': 3e-5,
'weight_decay':1e-6,
'num_workers': 4,
'accum_iter': 2, # suppoprt to do batch accumulation for backprop with effectively larger batch size
'verbose_step': 1,
'device': 'cuda:0',
'target_size' : 5,
'smoothing' : 0.2,
'swa_start_epoch' : 2,
## Following four are related to FixMatch
'mu' : 2,
'T' : 1, # temperature
'lambda_u' : 1.,
'threshold' : 0.85,
##
'debug' : False
}
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" papermill={"duration": 0.057123, "end_time": "2020-11-23T13:32:49.643710", "exception": false, "start_time": "2020-11-23T13:32:49.586587", "status": "completed"} tags=[]
train = pd.read_csv('./input/cassava-leaf-disease-classification/train.csv')
delete_id = ['2947932468.jpg', '2252529694.jpg', '2278017076.jpg']
train = train[~train['image_id'].isin(delete_id)].reset_index(drop=True)
train.head()
# + [markdown] papermill={"duration": 0.016085, "end_time": "2020-11-23T13:32:49.720073", "exception": false, "start_time": "2020-11-23T13:32:49.703988", "status": "completed"} tags=[]
# > We could do stratified validation split in each fold to make each fold's train and validation set looks like the whole train set in target distributions.
# + papermill={"duration": 0.032053, "end_time": "2020-11-23T13:32:49.768481", "exception": false, "start_time": "2020-11-23T13:32:49.736428", "status": "completed"} tags=[]
submission = pd.read_csv('./input/cassava-leaf-disease-classification/sample_submission.csv')
submission.head()
# + [markdown] papermill={"duration": 0.015931, "end_time": "2020-11-23T13:32:49.801027", "exception": false, "start_time": "2020-11-23T13:32:49.785096", "status": "completed"} tags=[]
# # Helper Functions
# + papermill={"duration": 0.315262, "end_time": "2020-11-23T13:32:50.132792", "exception": false, "start_time": "2020-11-23T13:32:49.817530", "status": "completed"} tags=[]
def seed_everything(seed):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
def get_img(path):
im_bgr = cv2.imread(path)
im_rgb = im_bgr[:, :, ::-1]
#print(im_rgb)
return im_rgb
# + [markdown] papermill={"duration": 0.021311, "end_time": "2020-11-23T13:32:50.174973", "exception": false, "start_time": "2020-11-23T13:32:50.153662", "status": "completed"} tags=[]
# # Dataset
# + papermill={"duration": 0.064816, "end_time": "2020-11-23T13:32:50.261340", "exception": false, "start_time": "2020-11-23T13:32:50.196524", "status": "completed"} tags=[]
def rand_bbox(size, lam):
W = size[0]
H = size[1]
cut_rat = np.sqrt(1. - lam)
cut_w = np.int(W * cut_rat)
cut_h = np.int(H * cut_rat)
# uniform
cx = np.random.randint(W)
cy = np.random.randint(H)
bbx1 = np.clip(cx - cut_w // 2, 0, W)
bby1 = np.clip(cy - cut_h // 2, 0, H)
bbx2 = np.clip(cx + cut_w // 2, 0, W)
bby2 = np.clip(cy + cut_h // 2, 0, H)
return bbx1, bby1, bbx2, bby2
class CassavaDataset(Dataset):
def __init__(self, df, data_root,
transforms=None,
output_label=True,
):
super().__init__()
self.df = df.reset_index(drop=True).copy()
self.transforms = transforms
self.data_root = data_root
self.output_label = output_label
self.labels = self.df['label'].values
def __len__(self):
return self.df.shape[0]
def __getitem__(self, index: int):
# get labels
if self.output_label:
target = self.labels[index]
img = get_img("{}/{}".format(self.data_root, self.df.loc[index]['image_id']))
if self.transforms:
img = self.transforms(image=img)['image']
if self.output_label == True:
return img, target
else:
return img
# + [markdown] papermill={"duration": 0.02183, "end_time": "2020-11-23T13:32:50.304795", "exception": false, "start_time": "2020-11-23T13:32:50.282965", "status": "completed"} tags=[]
# # Define Train\Validation Image Augmentations
# -
from albumentations.core.transforms_interface import DualTransform
# from albumentations.augmentations import functional as F
class GridMask(DualTransform):
"""GridMask augmentation for image classification and object detection.
Author: <NAME>
Email: <EMAIL>
2020/01/29
Args:
num_grid (int): number of grid in a row or column.
fill_value (int, float, lisf of int, list of float): value for dropped pixels.
rotate ((int, int) or int): range from which a random angle is picked. If rotate is a single int
an angle is picked from (-rotate, rotate). Default: (-90, 90)
mode (int):
0 - cropout a quarter of the square of each grid (left top)
1 - reserve a quarter of the square of each grid (left top)
2 - cropout 2 quarter of the square of each grid (left top & right bottom)
Targets:
image, mask
Image types:
uint8, float32
Reference:
| https://arxiv.org/abs/2001.04086
| https://github.com/akuxcw/GridMask
"""
def __init__(self, num_grid=3, fill_value=0, rotate=0, mode=0, always_apply=False, p=0.5):
super(GridMask, self).__init__(always_apply, p)
if isinstance(num_grid, int):
num_grid = (num_grid, num_grid)
if isinstance(rotate, int):
rotate = (-rotate, rotate)
self.num_grid = num_grid
self.fill_value = fill_value
self.rotate = rotate
self.mode = mode
self.masks = None
self.rand_h_max = []
self.rand_w_max = []
def init_masks(self, height, width):
if self.masks is None:
self.masks = []
n_masks = self.num_grid[1] - self.num_grid[0] + 1
for n, n_g in enumerate(range(self.num_grid[0], self.num_grid[1] + 1, 1)):
grid_h = height / n_g
grid_w = width / n_g
this_mask = np.ones((int((n_g + 1) * grid_h), int((n_g + 1) * grid_w))).astype(np.uint8)
for i in range(n_g + 1):
for j in range(n_g + 1):
this_mask[
int(i * grid_h) : int(i * grid_h + grid_h / 2),
int(j * grid_w) : int(j * grid_w + grid_w / 2)
] = self.fill_value
if self.mode == 2:
this_mask[
int(i * grid_h + grid_h / 2) : int(i * grid_h + grid_h),
int(j * grid_w + grid_w / 2) : int(j * grid_w + grid_w)
] = self.fill_value
if self.mode == 1:
this_mask = 1 - this_mask
self.masks.append(this_mask)
self.rand_h_max.append(grid_h)
self.rand_w_max.append(grid_w)
def apply(self, image, mask, rand_h, rand_w, angle, **params):
h, w = image.shape[:2]
mask = F.rotate(mask, angle) if self.rotate[1] > 0 else mask
mask = mask[:,:,np.newaxis] if image.ndim == 3 else mask
image *= mask[rand_h:rand_h+h, rand_w:rand_w+w].astype(image.dtype)
return image
def get_params_dependent_on_targets(self, params):
img = params['image']
height, width = img.shape[:2]
self.init_masks(height, width)
mid = np.random.randint(len(self.masks))
mask = self.masks[mid]
rand_h = np.random.randint(self.rand_h_max[mid])
rand_w = np.random.randint(self.rand_w_max[mid])
angle = np.random.randint(self.rotate[0], self.rotate[1]) if self.rotate[1] > 0 else 0
return {'mask': mask, 'rand_h': rand_h, 'rand_w': rand_w, 'angle': angle}
@property
def targets_as_params(self):
return ['image']
def get_transform_init_args_names(self):
return ('num_grid', 'fill_value', 'rotate', 'mode')
# + papermill={"duration": 0.590042, "end_time": "2020-11-23T13:32:50.916225", "exception": false, "start_time": "2020-11-23T13:32:50.326183", "status": "completed"} tags=[]
from albumentations import (
HorizontalFlip, VerticalFlip, IAAPerspective, ShiftScaleRotate, CLAHE, RandomRotate90,
Transpose, ShiftScaleRotate, Blur, OpticalDistortion, GridDistortion, HueSaturationValue,
IAAAdditiveGaussianNoise, GaussNoise, MotionBlur, MedianBlur, IAAPiecewiseAffine, RandomResizedCrop,
IAASharpen, IAAEmboss, RandomBrightnessContrast, Flip, OneOf, Compose, Normalize, Cutout, CoarseDropout, ShiftScaleRotate, CenterCrop, Resize
)
from albumentations.pytorch import ToTensorV2
def get_train_transforms():
return Compose([
OneOf([
Resize(CFG['img_size'], CFG['img_size'], p=1.),
CenterCrop(CFG['img_size'], CFG['img_size'], p=1.),
RandomResizedCrop(CFG['img_size'], CFG['img_size'], p=1.)
], p=1.),
Transpose(p=0.5),
HorizontalFlip(p=0.5),
VerticalFlip(p=0.5),
ShiftScaleRotate(p=0.5),
HueSaturationValue(hue_shift_limit=0.2, sat_shift_limit=0.2, val_shift_limit=0.2, p=0.5),
RandomBrightnessContrast(brightness_limit=(-0.1,0.1), contrast_limit=(-0.1, 0.1), p=0.5),
Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225], max_pixel_value=255.0, p=1.0),
CoarseDropout(p=0.5),
GridMask(num_grid=3, p=0.5),
ToTensorV2(p=1.0),
], p=1.)
def get_valid_transforms():
return Compose([
CenterCrop(CFG['img_size'], CFG['img_size'], p=1.),
Resize(CFG['img_size'], CFG['img_size']),
Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225], max_pixel_value=255.0, p=1.0),
ToTensorV2(p=1.0),
], p=1.)
def get_inference_transforms():
return Compose([
OneOf([
Resize(CFG['img_size'], CFG['img_size'], p=1.),
CenterCrop(CFG['img_size'], CFG['img_size'], p=1.),
RandomResizedCrop(CFG['img_size'], CFG['img_size'], p=1.)
], p=1.),
Transpose(p=0.5),
HorizontalFlip(p=0.5),
VerticalFlip(p=0.5),
Resize(CFG['img_size'], CFG['img_size']),
Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225], max_pixel_value=255.0, p=1.0),
ToTensorV2(p=1.0),
], p=1.)
# + [markdown] papermill={"duration": 0.024452, "end_time": "2020-11-23T13:32:50.962106", "exception": false, "start_time": "2020-11-23T13:32:50.937654", "status": "completed"} tags=[]
# # Model
# + papermill={"duration": 0.033239, "end_time": "2020-11-23T13:32:51.017593", "exception": false, "start_time": "2020-11-23T13:32:50.984354", "status": "completed"} tags=[]
class CassvaImgClassifier(nn.Module):
def __init__(self, model_arch, n_class, pretrained=False):
super().__init__()
self.model = timm.create_model(model_arch, pretrained=pretrained)
n_features = self.model.classifier.in_features
self.model.classifier = nn.Linear(n_features, n_class)
def forward(self, x):
x = self.model(x)
return x
# + [markdown] papermill={"duration": 0.024452, "end_time": "2020-11-23T13:32:50.962106", "exception": false, "start_time": "2020-11-23T13:32:50.937654", "status": "completed"} tags=[]
# # For FixMatch Unlabeled DataLoader
# -
#######
o = os.listdir('./input/cassava-disease/all/')
o = np.array([o]).T
label_col = np.ones_like(o)
o = np.concatenate((o,label_col),axis=1)
unlabeled = pd.DataFrame(o,columns=['image_id','label'])
unlabeled.head()
# unlabeled = train
# +
import PIL
import PIL.ImageOps
import PIL.ImageEnhance
import PIL.ImageDraw
from PIL import Image
PARAMETER_MAX = 10
def AutoContrast(img, **kwarg):
return PIL.ImageOps.autocontrast(img)
def Brightness(img, v, max_v, bias=0):
v = _float_parameter(v, max_v) + bias
return PIL.ImageEnhance.Brightness(img).enhance(v)
def Color(img, v, max_v, bias=0):
v = _float_parameter(v, max_v) + bias
return PIL.ImageEnhance.Color(img).enhance(v)
def Contrast(img, v, max_v, bias=0):
v = _float_parameter(v, max_v) + bias
return PIL.ImageEnhance.Contrast(img).enhance(v)
def Cutout(img, v, max_v, bias=0):
if v == 0:
return img
v = _float_parameter(v, max_v) + bias
v = int(v * min(img.size))
return CutoutAbs(img, v)
def CutoutAbs(img, v, **kwarg):
w, h = img.size
x0 = np.random.uniform(0, w)
y0 = np.random.uniform(0, h)
x0 = int(max(0, x0 - v / 2.))
y0 = int(max(0, y0 - v / 2.))
x1 = int(min(w, x0 + v))
y1 = int(min(h, y0 + v))
xy = (x0, y0, x1, y1)
# gray
color = (127, 127, 127)
img = img.copy()
PIL.ImageDraw.Draw(img).rectangle(xy, color)
return img
def Equalize(img, **kwarg):
return PIL.ImageOps.equalize(img)
def Identity(img, **kwarg):
return img
def Invert(img, **kwarg):
return PIL.ImageOps.invert(img)
def Posterize(img, v, max_v, bias=0):
v = _int_parameter(v, max_v) + bias
return PIL.ImageOps.posterize(img, v)
def Rotate(img, v, max_v, bias=0):
v = _int_parameter(v, max_v) + bias
if random.random() < 0.5:
v = -v
return img.rotate(v)
def Sharpness(img, v, max_v, bias=0):
v = _float_parameter(v, max_v) + bias
return PIL.ImageEnhance.Sharpness(img).enhance(v)
def ShearX(img, v, max_v, bias=0):
v = _float_parameter(v, max_v) + bias
if random.random() < 0.5:
v = -v
return img.transform(img.size, PIL.Image.AFFINE, (1, v, 0, 0, 1, 0))
def ShearY(img, v, max_v, bias=0):
v = _float_parameter(v, max_v) + bias
if random.random() < 0.5:
v = -v
return img.transform(img.size, PIL.Image.AFFINE, (1, 0, 0, v, 1, 0))
def Solarize(img, v, max_v, bias=0):
v = _int_parameter(v, max_v) + bias
return PIL.ImageOps.solarize(img, 256 - v)
def SolarizeAdd(img, v, max_v, bias=0, threshold=128):
v = _int_parameter(v, max_v) + bias
if random.random() < 0.5:
v = -v
img_np = np.array(img).astype(np.int)
img_np = img_np + v
img_np = np.clip(img_np, 0, 255)
img_np = img_np.astype(np.uint8)
img = Image.fromarray(img_np)
return PIL.ImageOps.solarize(img, threshold)
def TranslateX(img, v, max_v, bias=0):
v = _float_parameter(v, max_v) + bias
if random.random() < 0.5:
v = -v
v = int(v * img.size[0])
return img.transform(img.size, PIL.Image.AFFINE, (1, 0, v, 0, 1, 0))
def TranslateY(img, v, max_v, bias=0):
v = _float_parameter(v, max_v) + bias
if random.random() < 0.5:
v = -v
v = int(v * img.size[1])
return img.transform(img.size, PIL.Image.AFFINE, (1, 0, 0, 0, 1, v))
def _float_parameter(v, max_v):
return float(v) * max_v / PARAMETER_MAX
def _int_parameter(v, max_v):
return int(v * max_v / PARAMETER_MAX)
# +
class RandAugmentMC(object):
def __init__(self, n, m):
assert n >= 1
assert 1 <= m <= 10
self.n = n
self.m = m
self.augment_pool = fixmatch_augment_pool()
def __call__(self, img):
ops = random.choices(self.augment_pool, k=self.n)
for op, max_v, bias in ops:
v = np.random.randint(1, self.m)
if random.random() < 0.5:
img = op(img, v=v, max_v=max_v, bias=bias)
img = CutoutAbs(img, int(CFG['img_size']*0.5))
return img
def fixmatch_augment_pool():
# FixMatch paper
augs = [(AutoContrast, None, None),
(Brightness, 0.9, 0.05),
(Color, 0.9, 0.05),
(Contrast, 0.9, 0.05),
(Equalize, None, None),
(Identity, None, None),
(Posterize, 4, 4),
(Rotate, 30, 0),
(Sharpness, 0.9, 0.05),
(ShearX, 0.3, 0),
(ShearY, 0.3, 0),
(Solarize, 256, 0),
(TranslateX, 0.3, 0),
(TranslateY, 0.3, 0)]
return augs
class TransformFixMatch(object):
def __init__(self, mean, std):
self.weak = transforms.Compose([
transforms.RandomHorizontalFlip(),
transforms.RandomCrop(size=CFG['img_size'],
padding=int(CFG['img_size']*0.125),
padding_mode='reflect')])
self.strong = transforms.Compose([
transforms.RandomHorizontalFlip(),
transforms.RandomCrop(size=CFG['img_size'],
padding=int(CFG['img_size']*0.125),
padding_mode='reflect'),
RandAugmentMC(n=2, m=10)])
self.normalize = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=mean, std=std)])
def __call__(self, x):
weak = self.weak(x)
strong = self.strong(x)
return self.normalize(weak), self.normalize(strong)
class CassavaDataset_ul(Dataset):
def __init__(self, df, data_root,
transforms=None,
output_label=True,
):
super().__init__()
self.df = df.reset_index(drop=True).copy()
self.transforms = transforms
self.data_root = data_root
self.output_label = output_label
self.labels = self.df['label'].values
def __len__(self):
return self.df.shape[0]
def __getitem__(self, index: int):
# get labels
if self.output_label:
target = self.labels[index]
img = Image.open("{}/{}".format(self.data_root, self.df.loc[index]['image_id']))
if self.transforms:
img = self.transforms(img)
if self.output_label == True:
return img, target
else:
return img
# +
from torch.utils.data import RandomSampler
######################## 바꿔주자!!! 2019 데이터셋으로
# unlabeled_dataset = CassavaDataset_ul(unlabeled, './input/cassava-disease/all', transforms=TransformFixMatch(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]))
unlabeled_dataset = CassavaDataset_ul(unlabeled, './input/cassava-disease/all/', transforms=TransformFixMatch(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]))
train_loader_ul = torch.utils.data.DataLoader(
unlabeled_dataset,
sampler = RandomSampler(unlabeled_dataset),
batch_size=CFG['train_bs'] * CFG['mu'],
pin_memory=False,
drop_last=True,
num_workers=CFG['num_workers'],
)
def interleave(x, size):
s = list(x.shape)
return x.reshape([-1, size] + s[1:]).transpose(0, 1).reshape([-1] + s[1:])
def de_interleave(x, size):
s = list(x.shape)
return x.reshape([size, -1] + s[1:]).transpose(0, 1).reshape([-1] + s[1:])
# train_loader_ul = iter(train_loader_ul)
# (inputs_u_w, inputs_u_s), _ = train_loader_ul.next()
# print(len(inputs_u_s), len(inputs_u_w))
# + [markdown] papermill={"duration": 0.021054, "end_time": "2020-11-23T13:32:51.059722", "exception": false, "start_time": "2020-11-23T13:32:51.038668", "status": "completed"} tags=[]
# # Training APIs
# + papermill={"duration": 0.061685, "end_time": "2020-11-23T13:32:51.144150", "exception": false, "start_time": "2020-11-23T13:32:51.082465", "status": "completed"} tags=[]
def prepare_dataloader(df, trn_idx, val_idx, data_root='./input/cassava-leaf-disease-classification/train_images/'):
# from catalyst.data.sampler import BalanceClassSampler
train_ = df.loc[trn_idx,:].reset_index(drop=True)
valid_ = df.loc[val_idx,:].reset_index(drop=True)
train_ds = CassavaDataset(train_, data_root, transforms=get_train_transforms(), output_label=True)
valid_ds = CassavaDataset(valid_, data_root, transforms=get_valid_transforms(), output_label=True)
train_loader = torch.utils.data.DataLoader(
train_ds,
batch_size=CFG['train_bs'],
pin_memory=False,
drop_last=True,###
shuffle=True,
num_workers=CFG['num_workers'],
#sampler=BalanceClassSampler(labels=train_['label'].values, mode="downsampling")
)
val_loader = torch.utils.data.DataLoader(
valid_ds,
batch_size=CFG['valid_bs'],
num_workers=CFG['num_workers'],
shuffle=False,
pin_memory=False,
)
return train_loader, val_loader
def train_one_epoch(epoch, model, loss_fn, optimizer, train_loader, unlabeled_trainloader, device, scheduler=None, swa_scheduler=None, schd_batch_update=False):
model.train()
t = time.time()
running_loss = None
# pbar = tqdm(enumerate(train_loader), total=len(train_loader))
for step, (imgs, image_labels) in enumerate(train_loader):
imgs = imgs.float()
image_labels = image_labels.to(device).long()
try:
(inputs_u_s, inputs_u_w), _ = unlabeled_iter.next()
except:
unlabeled_iter = iter(unlabeled_trainloader)
(inputs_u_s, inputs_u_w), _ = unlabeled_iter.next()
inputs = interleave(
torch.cat((imgs, inputs_u_w, inputs_u_s)), 2*CFG['mu']+1).contiguous().to(device)
with autocast():
image_preds = model(inputs) #output = model(input)
logits = de_interleave(image_preds, 2*CFG['mu']+1)
logits_x = logits[:CFG['train_bs']]
logits_u_w, logits_u_s = logits[CFG['train_bs']:].chunk(2)
del logits
Lx = loss_fn(logits_x, image_labels)
pseudo_label = torch.softmax(logits_u_w.detach()/CFG['T'], dim=-1)
max_probs, targets_u = torch.max(pseudo_label, dim=-1)
mask = max_probs.ge(CFG['threshold']).float()
# Lu = (F.cross_entropy(logits_u_s, targets_u, reduction='none') * mask).mean()
Lu = (loss_fn(logits_u_s, targets_u, reduction='none')*mask).mean()
loss = Lx + CFG['lambda_u'] * Lu
scaler.scale(loss).backward()
if running_loss is None:
running_loss = loss.item()
else:
running_loss = running_loss * .99 + loss.item() * .01
if ((step + 1) % CFG['accum_iter'] == 0) or ((step + 1) == len(train_loader)):
scaler.step(optimizer)
scaler.update()
optimizer.zero_grad()
if scheduler is not None and schd_batch_update:
scheduler.step()
# if ((step + 1) % CFG['verbose_step'] == 0) or ((step + 1) == len(train_loader)):
# description = f'epoch {epoch} loss: {running_loss:.4f}'
# print(description)
# pbar.set_description(description)
if scheduler is not None and not schd_batch_update:
if epoch >= CFG['swa_start_epoch']:
swa_scheduler.step()
else:
scheduler.step()
def valid_one_epoch(epoch, model, loss_fn, val_loader, device, scheduler=None, schd_loss_update=False):
model.eval()
t = time.time()
loss_sum = 0
sample_num = 0
image_preds_all = []
image_targets_all = []
# pbar = tqdm(enumerate(val_loader), total=len(val_loader))
for step, (imgs, image_labels) in enumerate(val_loader):
imgs = imgs.to(device).float()
image_labels = image_labels.to(device).long()
image_preds = model(imgs) #output = model(input)
image_preds_all += [torch.argmax(image_preds, 1).detach().cpu().numpy()]
image_targets_all += [image_labels.detach().cpu().numpy()]
loss = loss_fn(image_preds, image_labels)
loss_sum += loss.item()*image_labels.shape[0]
sample_num += image_labels.shape[0]
# if ((step + 1) % CFG['verbose_step'] == 0) or ((step + 1) == len(val_loader)):
# description = f'epoch {epoch} loss: {loss_sum/sample_num:.4f}'
# pbar.set_description(description)
image_preds_all = np.concatenate(image_preds_all)
image_targets_all = np.concatenate(image_targets_all)
print('epoch = {}'.format(epoch+1), 'validation multi-class accuracy = {:.4f}'.format((image_preds_all==image_targets_all).mean()))
if scheduler is not None:
if schd_loss_update:
scheduler.step(loss_sum/sample_num)
else:
scheduler.step()
def inference_one_epoch(model, data_loader, device):
model.eval()
image_preds_all = []
# pbar = tqdm(enumerate(data_loader), total=len(data_loader))
with torch.no_grad():
for step, (imgs, image_labels) in enumerate(data_loader):
imgs = imgs.to(device).float()
image_preds = model(imgs) #output = model(input)
image_preds_all += [torch.softmax(image_preds, 1).detach().cpu().numpy()]
image_preds_all = np.concatenate(image_preds_all, axis=0)
return image_preds_all
# + papermill={"duration": 0.034873, "end_time": "2020-11-23T13:32:51.200704", "exception": false, "start_time": "2020-11-23T13:32:51.165831", "status": "completed"} tags=[]
# reference: https://www.kaggle.com/c/siim-isic-melanoma-classification/discussion/173733
class MyCrossEntropyLoss(_WeightedLoss):
def __init__(self, weight=None, reduction='mean'):
super().__init__(weight=weight, reduction=reduction)
self.weight = weight
self.reduction = reduction
def forward(self, inputs, targets):
lsm = F.log_softmax(inputs, -1)
if self.weight is not None:
lsm = lsm * self.weight.unsqueeze(0)
loss = -(targets * lsm).sum(-1)
if self.reduction == 'sum':
loss = loss.sum()
elif self.reduction == 'mean':
loss = loss.mean()
return loss
# -
# ====================================================
# Label Smoothing
# ====================================================
class LabelSmoothingLoss(nn.Module):
def __init__(self, classes, smoothing=0.0, dim=-1):
super(LabelSmoothingLoss, self).__init__()
self.confidence = 1.0 - smoothing
self.smoothing = smoothing
self.cls = classes
self.dim = dim
def forward(self, pred, target, reduction = 'mean'):
pred = pred.log_softmax(dim=self.dim)
with torch.no_grad():
true_dist = torch.zeros_like(pred)
true_dist.fill_(self.smoothing / (self.cls - 1))
true_dist.scatter_(1, target.data.unsqueeze(1), self.confidence)
if reduction == 'mean':
return torch.mean(torch.sum(-true_dist * pred, dim=self.dim))
else:
return torch.sum(-true_dist * pred, dim=self.dim)
# + [markdown] papermill={"duration": 0.020806, "end_time": "2020-11-23T13:32:51.243006", "exception": false, "start_time": "2020-11-23T13:32:51.222200", "status": "completed"} tags=[]
# # Main Loop
# -
from sklearn.metrics import accuracy_score
os.environ['CUDA_VISIBLE_DEVICES'] = '0' # specify GPUs locally
# +
# #debug
# train = pd.read_csv('./input/cassava-leaf-disease-classification/train_debug.csv')
# CFG['epochs']=7
# model_path = 'temporary'
# # !mkdir -p temporary
# -
model_path='v2_hwkim_fixmatch_2019_fast_thr085_bs9_mu2_7ep_CusSwa4'
# # !mkdir -p v2_hwkim_fixmatch_2019_fast_thr085_bs9_mu2_7ep_CusSwa4
if __name__ == '__main__':
for c in range(5):
train[c] = 0
folds = StratifiedKFold(n_splits=CFG['fold_num'], shuffle=True, random_state=CFG['seed']).split(np.arange(train.shape[0]), train.label.values)
for fold, (trn_idx, val_idx) in enumerate(folds):
if fold<3:
continue
print('Training with {} started'.format(fold))
print(len(trn_idx), len(val_idx))
train_loader, val_loader = prepare_dataloader(train, trn_idx, val_idx, data_root='./input/cassava-leaf-disease-classification/train_images/')
unlabeled_trainloader = train_loader_ul
device = torch.device(CFG['device'])
model = CassvaImgClassifier(CFG['model_arch'], train.label.nunique(), pretrained=True).to(device)
scaler = GradScaler()
optimizer = AdamP(model.parameters(), lr=CFG['lr'], weight_decay=CFG['weight_decay'])
scheduler = torch.optim.lr_scheduler.CosineAnnealingWarmRestarts(optimizer, T_0=CFG['swa_start_epoch']+1, T_mult=1, eta_min=CFG['min_lr'], last_epoch=-1)
swa_scheduler = SWALR(optimizer, swa_lr = CFG['min_lr'], anneal_epochs=1)
loss_tr = LabelSmoothingLoss(classes=CFG['target_size'], smoothing=CFG['smoothing']).to(device)
loss_fn = nn.CrossEntropyLoss().to(device)
for epoch in range(CFG['epochs']):
print(optimizer.param_groups[0]["lr"])
train_one_epoch(epoch, model, loss_tr, optimizer, train_loader, unlabeled_trainloader, device, scheduler=scheduler, swa_scheduler=swa_scheduler, schd_batch_update=False)
if epoch > CFG['swa_start_epoch']:
if epoch-1 == CFG['swa_start_epoch']:
swa_model = AveragedModel(model,device='cpu').to(device)
# update_bn(train_loader, swa_model, device=device)
else:
swa_model.update_parameters(model)
with torch.no_grad():
print('non swa')
valid_one_epoch(epoch, model, loss_fn, val_loader, device, scheduler=None, schd_loss_update=False)
if epoch > CFG['swa_start_epoch']:
print('swa')
valid_one_epoch(epoch, swa_model, loss_fn, val_loader, device, scheduler=None, schd_loss_update=False)
torch.save(model.state_dict(),'./'+model_path+'/{}_fold_{}_{}_{}'.format(CFG['model_arch'], fold, epoch, CFG['seed']))
del unlabeled_trainloader, model
with torch.no_grad():
# valid_one_epoch(epoch, swa_model, loss_fn, val_loader, device, scheduler=None, schd_loss_update=False)
torch.save(swa_model.module.state_dict(),'./'+model_path+'/noBN_swa_{}_fold_{}_{}'.format(CFG['model_arch'], fold, epoch))
# print('swa_BN')
# update_bn(train_loader, swa_model, device=device)
# valid_one_epoch(epoch, swa_model, loss_fn, val_loader, device, scheduler=None, schd_loss_update=False)
# torch.save(swa_model.state_dict(),'./'+model_path+'/BN_swa_{}_fold_{}_{}'.format(CFG['model_arch'], fold, epoch))
tst_preds = []
for tta in range(5):
tst_preds += [inference_one_epoch(swa_model, val_loader, device)]
train.loc[val_idx, [0, 1, 2, 3, 4]] = np.mean(tst_preds, axis=0)
del swa_model, optimizer, train_loader, val_loader, scaler, scheduler
torch.cuda.empty_cache()
train['pred'] = np.array(train[[0, 1, 2, 3, 4]]).argmax(axis=1)
print(accuracy_score(train['label'].values, train['pred'].values))
|
Cassava Leaf Disease Classification/code/10. FixMatch(SEDD_719).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Character level language model - Dinosaurus Island
#
# Welcome to Dinosaurus Island! 65 million years ago, dinosaurs existed, and in this assignment they are back. You are in charge of a special task. Leading biology researchers are creating new breeds of dinosaurs and bringing them to life on earth, and your job is to give names to these dinosaurs. If a dinosaur does not like its name, it might go berserk, so choose wisely!
#
# <table>
# <td>
# <img src="images/dino.jpg" style="width:250;height:300px;">
#
# </td>
#
# </table>
#
# Luckily you have learned some deep learning and you will use it to save the day. Your assistant has collected a list of all the dinosaur names they could find, and compiled them into this [dataset](dinos.txt). (Feel free to take a look by clicking the previous link.) To create new dinosaur names, you will build a character level language model to generate new names. Your algorithm will learn the different name patterns, and randomly generate new names. Hopefully this algorithm will keep you and your team safe from the dinosaurs' wrath!
#
# By completing this assignment you will learn:
#
# - How to store text data for processing using an RNN
# - How to synthesize data, by sampling predictions at each time step and passing it to the next RNN-cell unit
# - How to build a character-level text generation recurrent neural network
# - Why clipping the gradients is important
#
# We will begin by loading in some functions that we have provided for you in `rnn_utils`. Specifically, you have access to functions such as `rnn_forward` and `rnn_backward` which are equivalent to those you've implemented in the previous assignment.
# ## <font color='darkblue'>Updates</font>
#
# #### If you were working on the notebook before this update...
# * The current notebook is version "3a".
# * You can find your original work saved in the notebook with the previous version name ("v3")
# * To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#
# #### List of updates
# * Sort and print `chars` list of characters.
# * Import and use pretty print
# * `clip`:
# - Additional details on why we need to use the "out" parameter.
# - Modified for loop to have students fill in the correct items to loop through.
# - Added a test case to check for hard-coding error.
# * `sample`
# - additional hints added to steps 1,2,3,4.
# - "Using 2D arrays instead of 1D arrays".
# - explanation of numpy.ravel().
# - fixed expected output.
# - clarified comments in the code.
# * "training the model"
# - Replaced the sample code with explanations for how to set the index, X and Y (for a better learning experience).
# * Spelling, grammar and wording corrections.
import numpy as np
from utils import *
import random
import pprint
# ## 1 - Problem Statement
#
# ### 1.1 - Dataset and Preprocessing
#
# Run the following cell to read the dataset of dinosaur names, create a list of unique characters (such as a-z), and compute the dataset and vocabulary size.
data = open('dinos.txt', 'r').read()
data= data.lower()
chars = list(set(data))
data_size, vocab_size = len(data), len(chars)
print('There are %d total characters and %d unique characters in your data.' % (data_size, vocab_size))
#
# * The characters are a-z (26 characters) plus the "\n" (or newline character).
# * In this assignment, the newline character "\n" plays a role similar to the `<EOS>` (or "End of sentence") token we had discussed in lecture.
# - Here, "\n" indicates the end of the dinosaur name rather than the end of a sentence.
# * `char_to_ix`: In the cell below, we create a python dictionary (i.e., a hash table) to map each character to an index from 0-26.
# * `ix_to_char`: We also create a second python dictionary that maps each index back to the corresponding character.
# - This will help you figure out what index corresponds to what character in the probability distribution output of the softmax layer.
chars = sorted(chars)
print(chars)
char_to_ix = { ch:i for i,ch in enumerate(chars) }
ix_to_char = { i:ch for i,ch in enumerate(chars) }
pp = pprint.PrettyPrinter(indent=4)
pp.pprint(ix_to_char)
# ### 1.2 - Overview of the model
#
# Your model will have the following structure:
#
# - Initialize parameters
# - Run the optimization loop
# - Forward propagation to compute the loss function
# - Backward propagation to compute the gradients with respect to the loss function
# - Clip the gradients to avoid exploding gradients
# - Using the gradients, update your parameters with the gradient descent update rule.
# - Return the learned parameters
#
# <img src="images/rnn.png" style="width:450;height:300px;">
# <caption><center> **Figure 1**: Recurrent Neural Network, similar to what you had built in the previous notebook "Building a Recurrent Neural Network - Step by Step". </center></caption>
#
# * At each time-step, the RNN tries to predict what is the next character given the previous characters.
# * The dataset $\mathbf{X} = (x^{\langle 1 \rangle}, x^{\langle 2 \rangle}, ..., x^{\langle T_x \rangle})$ is a list of characters in the training set.
# * $\mathbf{Y} = (y^{\langle 1 \rangle}, y^{\langle 2 \rangle}, ..., y^{\langle T_x \rangle})$ is the same list of characters but shifted one character forward.
# * At every time-step $t$, $y^{\langle t \rangle} = x^{\langle t+1 \rangle}$. The prediction at time $t$ is the same as the input at time $t + 1$.
# ## 2 - Building blocks of the model
#
# In this part, you will build two important blocks of the overall model:
# - Gradient clipping: to avoid exploding gradients
# - Sampling: a technique used to generate characters
#
# You will then apply these two functions to build the model.
# ### 2.1 - Clipping the gradients in the optimization loop
#
# In this section you will implement the `clip` function that you will call inside of your optimization loop.
#
# #### Exploding gradients
# * When gradients are very large, they're called "exploding gradients."
# * Exploding gradients make the training process more difficult, because the updates may be so large that they "overshoot" the optimal values during back propagation.
#
# Recall that your overall loop structure usually consists of:
# * forward pass,
# * cost computation,
# * backward pass,
# * parameter update.
#
# Before updating the parameters, you will perform gradient clipping to make sure that your gradients are not "exploding."
#
# #### gradient clipping
# In the exercise below, you will implement a function `clip` that takes in a dictionary of gradients and returns a clipped version of gradients if needed.
# * There are different ways to clip gradients.
# * We will use a simple element-wise clipping procedure, in which every element of the gradient vector is clipped to lie between some range [-N, N].
# * For example, if the N=10
# - The range is [-10, 10]
# - If any component of the gradient vector is greater than 10, it is set to 10.
# - If any component of the gradient vector is less than -10, it is set to -10.
# - If any components are between -10 and 10, they keep their original values.
#
# <img src="images/clip.png" style="width:400;height:150px;">
# <caption><center> **Figure 2**: Visualization of gradient descent with and without gradient clipping, in a case where the network is running into "exploding gradient" problems. </center></caption>
#
# **Exercise**:
# Implement the function below to return the clipped gradients of your dictionary `gradients`.
# * Your function takes in a maximum threshold and returns the clipped versions of the gradients.
# * You can check out [numpy.clip](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.clip.html).
# - You will need to use the argument "`out = ...`".
# - Using the "`out`" parameter allows you to update a variable "in-place".
# - If you don't use "`out`" argument, the clipped variable is stored in the variable "gradient" but does not update the gradient variables `dWax`, `dWaa`, `dWya`, `db`, `dby`.
# +
### GRADED FUNCTION: clip
def clip(gradients, maxValue):
'''
Clips the gradients' values between minimum and maximum.
Arguments:
gradients -- a dictionary containing the gradients "dWaa", "dWax", "dWya", "db", "dby"
maxValue -- everything above this number is set to this number, and everything less than -maxValue is set to -maxValue
Returns:
gradients -- a dictionary with the clipped gradients.
'''
dWaa, dWax, dWya, db, dby = gradients['dWaa'], gradients['dWax'], gradients['dWya'], gradients['db'], gradients['dby']
### START CODE HERE ###
# clip to mitigate exploding gradients, loop over [dWax, dWaa, dWya, db, dby]. (≈2 lines)
for gradient in [dWax, dWaa, dWya, db, dby]:
gradient=np.clip(gradient,-maxValue,maxValue,out=gradient)
### END CODE HERE ###
gradients = {"dWaa": dWaa, "dWax": dWax, "dWya": dWya, "db": db, "dby": dby}
return gradients
# -
# Test with a maxvalue of 10
maxValue = 10
np.random.seed(3)
dWax = np.random.randn(5,3)*10
dWaa = np.random.randn(5,5)*10
dWya = np.random.randn(2,5)*10
db = np.random.randn(5,1)*10
dby = np.random.randn(2,1)*10
gradients = {"dWax": dWax, "dWaa": dWaa, "dWya": dWya, "db": db, "dby": dby}
gradients = clip(gradients, maxValue)
print("gradients[\"dWaa\"][1][2] =", gradients["dWaa"][1][2])
print("gradients[\"dWax\"][3][1] =", gradients["dWax"][3][1])
print("gradients[\"dWya\"][1][2] =", gradients["dWya"][1][2])
print("gradients[\"db\"][4] =", gradients["db"][4])
print("gradients[\"dby\"][1] =", gradients["dby"][1])
#
# ** Expected output:**
#
# ```Python
# gradients["dWaa"][1][2] = 10.0
# gradients["dWax"][3][1] = -10.0
# gradients["dWya"][1][2] = 0.29713815361
# gradients["db"][4] = [ 10.]
# gradients["dby"][1] = [ 8.45833407]
# ```
# Test with a maxValue of 5
maxValue = 5
np.random.seed(3)
dWax = np.random.randn(5,3)*10
dWaa = np.random.randn(5,5)*10
dWya = np.random.randn(2,5)*10
db = np.random.randn(5,1)*10
dby = np.random.randn(2,1)*10
gradients = {"dWax": dWax, "dWaa": dWaa, "dWya": dWya, "db": db, "dby": dby}
gradients = clip(gradients, maxValue)
print("gradients[\"dWaa\"][1][2] =", gradients["dWaa"][1][2])
print("gradients[\"dWax\"][3][1] =", gradients["dWax"][3][1])
print("gradients[\"dWya\"][1][2] =", gradients["dWya"][1][2])
print("gradients[\"db\"][4] =", gradients["db"][4])
print("gradients[\"dby\"][1] =", gradients["dby"][1])
# ** Expected Output: **
# ```Python
# gradients["dWaa"][1][2] = 5.0
# gradients["dWax"][3][1] = -5.0
# gradients["dWya"][1][2] = 0.29713815361
# gradients["db"][4] = [ 5.]
# gradients["dby"][1] = [ 5.]
# ```
# ### 2.2 - Sampling
#
# Now assume that your model is trained. You would like to generate new text (characters). The process of generation is explained in the picture below:
#
# <img src="images/dinos3.png" style="width:500;height:300px;">
# <caption><center> **Figure 3**: In this picture, we assume the model is already trained. We pass in $x^{\langle 1\rangle} = \vec{0}$ at the first time step, and have the network sample one character at a time. </center></caption>
# **Exercise**: Implement the `sample` function below to sample characters. You need to carry out 4 steps:
#
# - **Step 1**: Input the "dummy" vector of zeros $x^{\langle 1 \rangle} = \vec{0}$.
# - This is the default input before we've generated any characters.
# We also set $a^{\langle 0 \rangle} = \vec{0}$
# - **Step 2**: Run one step of forward propagation to get $a^{\langle 1 \rangle}$ and $\hat{y}^{\langle 1 \rangle}$. Here are the equations:
#
# hidden state:
# $$ a^{\langle t+1 \rangle} = \tanh(W_{ax} x^{\langle t+1 \rangle } + W_{aa} a^{\langle t \rangle } + b)\tag{1}$$
#
# activation:
# $$ z^{\langle t + 1 \rangle } = W_{ya} a^{\langle t + 1 \rangle } + b_y \tag{2}$$
#
# prediction:
# $$ \hat{y}^{\langle t+1 \rangle } = softmax(z^{\langle t + 1 \rangle })\tag{3}$$
#
# - Details about $\hat{y}^{\langle t+1 \rangle }$:
# - Note that $\hat{y}^{\langle t+1 \rangle }$ is a (softmax) probability vector (its entries are between 0 and 1 and sum to 1).
# - $\hat{y}^{\langle t+1 \rangle}_i$ represents the probability that the character indexed by "i" is the next character.
# - We have provided a `softmax()` function that you can use.
# #### Additional Hints
#
# - $x^{\langle 1 \rangle}$ is `x` in the code. When creating the one-hot vector, make a numpy array of zeros, with the number of rows equal to the number of unique characters, and the number of columns equal to one. It's a 2D and not a 1D array.
# - $a^{\langle 0 \rangle}$ is `a_prev` in the code. It is a numpy array of zeros, where the number of rows is $n_{a}$, and number of columns is 1. It is a 2D array as well. $n_{a}$ is retrieved by getting the number of columns in $W_{aa}$ (the numbers need to match in order for the matrix multiplication $W_{aa}a^{\langle t \rangle}$ to work.
# - [numpy.dot](https://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html)
# - [numpy.tanh](https://docs.scipy.org/doc/numpy/reference/generated/numpy.tanh.html)
# #### Using 2D arrays instead of 1D arrays
# * You may be wondering why we emphasize that $x^{\langle 1 \rangle}$ and $a^{\langle 0 \rangle}$ are 2D arrays and not 1D vectors.
# * For matrix multiplication in numpy, if we multiply a 2D matrix with a 1D vector, we end up with with a 1D array.
# * This becomes a problem when we add two arrays where we expected them to have the same shape.
# * When two arrays with a different number of dimensions are added together, Python "broadcasts" one across the other.
# * Here is some sample code that shows the difference between using a 1D and 2D array.
import numpy as np
matrix1 = np.array([[1,1],[2,2],[3,3]]) # (3,2)
matrix2 = np.array([[0],[0],[0]]) # (3,1)
vector1D = np.array([1,1]) # (2,)
vector2D = np.array([[1],[1]]) # (2,1)
print("matrix1 \n", matrix1,"\n")
print("matrix2 \n", matrix2,"\n")
print("vector1D \n", vector1D,"\n")
print("vector2D \n", vector2D)
print("Multiply 2D and 1D arrays: result is a 1D array\n",
np.dot(matrix1,vector1D))
print("Multiply 2D and 2D arrays: result is a 2D array\n",
np.dot(matrix1,vector2D))
print("Adding (3 x 1) vector to a (3 x 1) vector is a (3 x 1) vector\n",
"This is what we want here!\n",
np.dot(matrix1,vector2D) + matrix2)
print("Adding a (3,) vector to a (3 x 1) vector\n",
"broadcasts the 1D array across the second dimension\n",
"Not what we want here!\n",
np.dot(matrix1,vector1D) + matrix2
)
# - **Step 3**: Sampling:
# - Now that we have $y^{\langle t+1 \rangle}$, we want to select the next letter in the dinosaur name. If we select the most probable, the model will always generate the same result given a starting letter.
# - To make the results more interesting, we will use np.random.choice to select a next letter that is likely, but not always the same.
# - Sampling is the selection of a value from a group of values, where each value has a probability of being picked.
# - Sampling allows us to generate random sequences of values.
# - Pick the next character's index according to the probability distribution specified by $\hat{y}^{\langle t+1 \rangle }$.
# - This means that if $\hat{y}^{\langle t+1 \rangle }_i = 0.16$, you will pick the index "i" with 16% probability.
# - You can use [np.random.choice](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.random.choice.html).
#
# Example of how to use `np.random.choice()`:
# ```python
# np.random.seed(0)
# probs = np.array([0.1, 0.0, 0.7, 0.2])
# idx = np.random.choice([0, 1, 2, 3] p = probs)
# ```
# - This means that you will pick the index (`idx`) according to the distribution:
#
# $P(index = 0) = 0.1, P(index = 1) = 0.0, P(index = 2) = 0.7, P(index = 3) = 0.2$.
#
# - Note that the value that's set to `p` should be set to a 1D vector.
# - Also notice that $\hat{y}^{\langle t+1 \rangle}$, which is `y` in the code, is a 2D array.
# ##### Additional Hints
# - [range](https://docs.python.org/3/library/functions.html#func-range)
# - [numpy.ravel](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ravel.html) takes a multi-dimensional array and returns its contents inside of a 1D vector.
# ```Python
# arr = np.array([[1,2],[3,4]])
# print("arr")
# print(arr)
# print("arr.ravel()")
# print(arr.ravel())
# ```
# Output:
# ```Python
# arr
# [[1 2]
# [3 4]]
# arr.ravel()
# [1 2 3 4]
# ```
#
# - Note that `append` is an "in-place" operation. In other words, don't do this:
# ```Python
# fun_hobbies = fun_hobbies.append('learning') ## Doesn't give you what you want
# ```
# - **Step 4**: Update to $x^{\langle t \rangle }$
# - The last step to implement in `sample()` is to update the variable `x`, which currently stores $x^{\langle t \rangle }$, with the value of $x^{\langle t + 1 \rangle }$.
# - You will represent $x^{\langle t + 1 \rangle }$ by creating a one-hot vector corresponding to the character that you have chosen as your prediction.
# - You will then forward propagate $x^{\langle t + 1 \rangle }$ in Step 1 and keep repeating the process until you get a "\n" character, indicating that you have reached the end of the dinosaur name.
# ##### Additional Hints
# - In order to reset `x` before setting it to the new one-hot vector, you'll want to set all the values to zero.
# - You can either create a new numpy array: [numpy.zeros](https://docs.scipy.org/doc/numpy/reference/generated/numpy.zeros.html)
# - Or fill all values with a single number: [numpy.ndarray.fill](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ndarray.fill.html)
# +
# GRADED FUNCTION: sample
def sample(parameters, char_to_ix, seed):
"""
Sample a sequence of characters according to a sequence of probability distributions output of the RNN
Arguments:
parameters -- python dictionary containing the parameters Waa, Wax, Wya, by, and b.
char_to_ix -- python dictionary mapping each character to an index.
seed -- used for grading purposes. Do not worry about it.
Returns:
indices -- a list of length n containing the indices of the sampled characters.
"""
# Retrieve parameters and relevant shapes from "parameters" dictionary
Waa, Wax, Wya, by, b = parameters['Waa'], parameters['Wax'], parameters['Wya'], parameters['by'], parameters['b']
vocab_size = by.shape[0]
n_a = Waa.shape[1]
### START CODE HERE ###
# Step 1: Create the a zero vector x that can be used as the one-hot vector
# representing the first character (initializing the sequence generation). (≈1 line)
x = np.zeros((vocab_size,1))
# Step 1': Initialize a_prev as zeros (≈1 line)
a_prev = np.zeros((n_a,1))
# Create an empty list of indices, this is the list which will contain the list of indices of the characters to generate (≈1 line)
indices = []
# idx is the index of the one-hot vector x that is set to 1
# All other positions in x are zero.
# We will initialize idx to -1
idx = -1
# Loop over time-steps t. At each time-step:
# sample a character from a probability distribution
# and append its index (`idx`) to the list "indices".
# We'll stop if we reach 50 characters
# (which should be very unlikely with a well trained model).
# Setting the maximum number of characters helps with debugging and prevents infinite loops.
counter = 0
newline_character = char_to_ix['\n']
while (idx != newline_character and counter != 50):
# Step 2: Forward propagate x using the equations (1), (2) and (3)
a = np.tanh(np.dot(Waa,a_prev)+np.dot(Wax,x)+b)
z = np.dot(Wya,a)+by
y = softmax(z)
# for grading purposes
np.random.seed(counter+seed)
# Step 3: Sample the index of a character within the vocabulary from the probability distribution y
# (see additional hints above)
idx = np.random.choice(range(len(y)),p=y.ravel())
# Append the index to "indices"
indices.append(idx)
# Step 4: Overwrite the input x with one that corresponds to the sampled index `idx`.
# (see additional hints above)
x = np.zeros((vocab_size,1))
x[idx] = 1
# Update "a_prev" to be "a"
a_prev = a
# for grading purposes
seed += 1
counter +=1
### END CODE HERE ###
if (counter == 50):
indices.append(char_to_ix['\n'])
return indices
# +
np.random.seed(2)
_, n_a = 20, 100
Wax, Waa, Wya = np.random.randn(n_a, vocab_size), np.random.randn(n_a, n_a), np.random.randn(vocab_size, n_a)
b, by = np.random.randn(n_a, 1), np.random.randn(vocab_size, 1)
parameters = {"Wax": Wax, "Waa": Waa, "Wya": Wya, "b": b, "by": by}
indices = sample(parameters, char_to_ix, 0)
print("Sampling:")
print("list of sampled indices:\n", indices)
print("list of sampled characters:\n", [ix_to_char[i] for i in indices])
# -
# ** Expected output:**
#
# ```Python
# Sampling:
# list of sampled indices:
# [12, 17, 24, 14, 13, 9, 10, 22, 24, 6, 13, 11, 12, 6, 21, 15, 21, 14, 3, 2, 1, 21, 18, 24, 7, 25, 6, 25, 18, 10, 16, 2, 3, 8, 15, 12, 11, 7, 1, 12, 10, 2, 7, 7, 11, 17, 24, 12, 13, 24, 0]
# list of sampled characters:
# ['l', 'q', 'x', 'n', 'm', 'i', 'j', 'v', 'x', 'f', 'm', 'k', 'l', 'f', 'u', 'o', 'u', 'n', 'c', 'b', 'a', 'u', 'r', 'x', 'g', 'y', 'f', 'y', 'r', 'j', 'p', 'b', 'c', 'h', 'o', 'l', 'k', 'g', 'a', 'l', 'j', 'b', 'g', 'g', 'k', 'q', 'x', 'l', 'm', 'x', '\n']
# ```
#
# * Please note that over time, if there are updates to the back-end of the Coursera platform (that may update the version of numpy), the actual list of sampled indices and sampled characters may change.
# * If you follow the instructions given above and get an output without errors, it's possible the routine is correct even if your output doesn't match the expected output. Submit your assignment to the grader to verify its correctness.
# ## 3 - Building the language model
#
# It is time to build the character-level language model for text generation.
#
#
# ### 3.1 - Gradient descent
#
# * In this section you will implement a function performing one step of stochastic gradient descent (with clipped gradients).
# * You will go through the training examples one at a time, so the optimization algorithm will be stochastic gradient descent.
#
# As a reminder, here are the steps of a common optimization loop for an RNN:
#
# - Forward propagate through the RNN to compute the loss
# - Backward propagate through time to compute the gradients of the loss with respect to the parameters
# - Clip the gradients
# - Update the parameters using gradient descent
#
# **Exercise**: Implement the optimization process (one step of stochastic gradient descent).
#
# The following functions are provided:
#
# ```python
# def rnn_forward(X, Y, a_prev, parameters):
# """ Performs the forward propagation through the RNN and computes the cross-entropy loss.
# It returns the loss' value as well as a "cache" storing values to be used in backpropagation."""
# ....
# return loss, cache
#
# def rnn_backward(X, Y, parameters, cache):
# """ Performs the backward propagation through time to compute the gradients of the loss with respect
# to the parameters. It returns also all the hidden states."""
# ...
# return gradients, a
#
# def update_parameters(parameters, gradients, learning_rate):
# """ Updates parameters using the Gradient Descent Update Rule."""
# ...
# return parameters
# ```
#
# Recall that you previously implemented the `clip` function:
#
# ```Python
# def clip(gradients, maxValue)
# """Clips the gradients' values between minimum and maximum."""
# ...
# return gradients
# ```
# #### parameters
#
# * Note that the weights and biases inside the `parameters` dictionary are being updated by the optimization, even though `parameters` is not one of the returned values of the `optimize` function. The `parameters` dictionary is passed by reference into the function, so changes to this dictionary are making changes to the `parameters` dictionary even when accessed outside of the function.
# * Python dictionaries and lists are "pass by reference", which means that if you pass a dictionary into a function and modify the dictionary within the function, this changes that same dictionary (it's not a copy of the dictionary).
# +
# GRADED FUNCTION: optimize
def optimize(X, Y, a_prev, parameters, learning_rate = 0.01):
"""
Execute one step of the optimization to train the model.
Arguments:
X -- list of integers, where each integer is a number that maps to a character in the vocabulary.
Y -- list of integers, exactly the same as X but shifted one index to the left.
a_prev -- previous hidden state.
parameters -- python dictionary containing:
Wax -- Weight matrix multiplying the input, numpy array of shape (n_a, n_x)
Waa -- Weight matrix multiplying the hidden state, numpy array of shape (n_a, n_a)
Wya -- Weight matrix relating the hidden-state to the output, numpy array of shape (n_y, n_a)
b -- Bias, numpy array of shape (n_a, 1)
by -- Bias relating the hidden-state to the output, numpy array of shape (n_y, 1)
learning_rate -- learning rate for the model.
Returns:
loss -- value of the loss function (cross-entropy)
gradients -- python dictionary containing:
dWax -- Gradients of input-to-hidden weights, of shape (n_a, n_x)
dWaa -- Gradients of hidden-to-hidden weights, of shape (n_a, n_a)
dWya -- Gradients of hidden-to-output weights, of shape (n_y, n_a)
db -- Gradients of bias vector, of shape (n_a, 1)
dby -- Gradients of output bias vector, of shape (n_y, 1)
a[len(X)-1] -- the last hidden state, of shape (n_a, 1)
"""
### START CODE HERE ###
# Forward propagate through time (≈1 line)
loss, cache = rnn_forward(X,Y,a_prev,parameters)
# Backpropagate through time (≈1 line)
gradients, a = rnn_backward(X,Y,parameters,cache)
# Clip your gradients between -5 (min) and 5 (max) (≈1 line)
gradients = clip(gradients,5)
# Update parameters (≈1 line)
parameters = update_parameters(parameters,gradients,learning_rate)
### END CODE HERE ###
return loss, gradients, a[len(X)-1]
# +
np.random.seed(1)
vocab_size, n_a = 27, 100
a_prev = np.random.randn(n_a, 1)
Wax, Waa, Wya = np.random.randn(n_a, vocab_size), np.random.randn(n_a, n_a), np.random.randn(vocab_size, n_a)
b, by = np.random.randn(n_a, 1), np.random.randn(vocab_size, 1)
parameters = {"Wax": Wax, "Waa": Waa, "Wya": Wya, "b": b, "by": by}
X = [12,3,5,11,22,3]
Y = [4,14,11,22,25, 26]
loss, gradients, a_last = optimize(X, Y, a_prev, parameters, learning_rate = 0.01)
print("Loss =", loss)
print("gradients[\"dWaa\"][1][2] =", gradients["dWaa"][1][2])
print("np.argmax(gradients[\"dWax\"]) =", np.argmax(gradients["dWax"]))
print("gradients[\"dWya\"][1][2] =", gradients["dWya"][1][2])
print("gradients[\"db\"][4] =", gradients["db"][4])
print("gradients[\"dby\"][1] =", gradients["dby"][1])
print("a_last[4] =", a_last[4])
# -
# ** Expected output:**
#
# ```Python
# Loss = 126.503975722
# gradients["dWaa"][1][2] = 0.194709315347
# np.argmax(gradients["dWax"]) = 93
# gradients["dWya"][1][2] = -0.007773876032
# gradients["db"][4] = [-0.06809825]
# gradients["dby"][1] = [ 0.01538192]
# a_last[4] = [-1.]
# ```
# ### 3.2 - Training the model
# * Given the dataset of dinosaur names, we use each line of the dataset (one name) as one training example.
# * Every 100 steps of stochastic gradient descent, you will sample 10 randomly chosen names to see how the algorithm is doing.
# * Remember to shuffle the dataset, so that stochastic gradient descent visits the examples in random order.
#
# **Exercise**: Follow the instructions and implement `model()`. When `examples[index]` contains one dinosaur name (string), to create an example (X, Y), you can use this:
#
# ##### Set the index `idx` into the list of examples
# * Using the for-loop, walk through the shuffled list of dinosaur names in the list "examples".
# * If there are 100 examples, and the for-loop increments the index to 100 onwards, think of how you would make the index cycle back to 0, so that we can continue feeding the examples into the model when j is 100, 101, etc.
# * Hint: 101 divided by 100 is zero with a remainder of 1.
# * `%` is the modulus operator in python.
#
# ##### Extract a single example from the list of examples
# * `single_example`: use the `idx` index that you set previously to get one word from the list of examples.
# ##### Convert a string into a list of characters: `single_example_chars`
# * `single_example_chars`: A string is a list of characters.
# * You can use a list comprehension (recommended over for-loops) to generate a list of characters.
# ```Python
# str = 'I love learning'
# list_of_chars = [c for c in str]
# print(list_of_chars)
# ```
#
# ```
# ['I', ' ', 'l', 'o', 'v', 'e', ' ', 'l', 'e', 'a', 'r', 'n', 'i', 'n', 'g']
# ```
# ##### Convert list of characters to a list of integers: `single_example_ix`
# * Create a list that contains the index numbers associated with each character.
# * Use the dictionary `char_to_ix`
# * You can combine this with the list comprehension that is used to get a list of characters from a string.
# * This is a separate line of code below, to help learners clarify each step in the function.
# ##### Create the list of input characters: `X`
# * `rnn_forward` uses the `None` value as a flag to set the input vector as a zero-vector.
# * Prepend the `None` value in front of the list of input characters.
# * There is more than one way to prepend a value to a list. One way is to add two lists together: `['a'] + ['b']`
# ##### Get the integer representation of the newline character `ix_newline`
# * `ix_newline`: The newline character signals the end of the dinosaur name.
# - get the integer representation of the newline character `'\n'`.
# - Use `char_to_ix`
# ##### Set the list of labels (integer representation of the characters): `Y`
# * The goal is to train the RNN to predict the next letter in the name, so the labels are the list of characters that are one time step ahead of the characters in the input `X`.
# - For example, `Y[0]` contains the same value as `X[1]`
# * The RNN should predict a newline at the last letter so add ix_newline to the end of the labels.
# - Append the integer representation of the newline character to the end of `Y`.
# - Note that `append` is an in-place operation.
# - It might be easier for you to add two lists together.
# +
# GRADED FUNCTION: model
def model(data, ix_to_char, char_to_ix, num_iterations = 35000, n_a = 50, dino_names = 7, vocab_size = 27):
"""
Trains the model and generates dinosaur names.
Arguments:
data -- text corpus
ix_to_char -- dictionary that maps the index to a character
char_to_ix -- dictionary that maps a character to an index
num_iterations -- number of iterations to train the model for
n_a -- number of units of the RNN cell
dino_names -- number of dinosaur names you want to sample at each iteration.
vocab_size -- number of unique characters found in the text (size of the vocabulary)
Returns:
parameters -- learned parameters
"""
# Retrieve n_x and n_y from vocab_size
n_x, n_y = vocab_size, vocab_size
# Initialize parameters
parameters = initialize_parameters(n_a, n_x, n_y)
# Initialize loss (this is required because we want to smooth our loss)
loss = get_initial_loss(vocab_size, dino_names)
# Build list of all dinosaur names (training examples).
with open("dinos.txt") as f:
examples = f.readlines()
examples = [x.lower().strip() for x in examples]
# Shuffle list of all dinosaur names
np.random.seed(0)
np.random.shuffle(examples)
# Initialize the hidden state of your LSTM
a_prev = np.zeros((n_a, 1))
# Optimization loop
for j in range(num_iterations):
### START CODE HERE ###
# Set the index `idx` (see instructions above)
idx = j%len(examples)
# Set the input X (see instructions above)
single_example = examples[idx]
single_example_chars = [c for c in single_example]
single_example_ix = [char_to_ix[c] for c in single_example_chars]
X = [None]+single_example_ix
# Set the labels Y (see instructions above)
ix_newline = [char_to_ix['\n']]
Y = X[1:]+ix_newline
# Perform one optimization step: Forward-prop -> Backward-prop -> Clip -> Update parameters
# Choose a learning rate of 0.01
curr_loss, gradients, a_prev = optimize(X,Y,a_prev,parameters,0.01)
### END CODE HERE ###
# Use a latency trick to keep the loss smooth. It happens here to accelerate the training.
loss = smooth(loss, curr_loss)
# Every 2000 Iteration, generate "n" characters thanks to sample() to check if the model is learning properly
if j % 2000 == 0:
print('Iteration: %d, Loss: %f' % (j, loss) + '\n')
# The number of dinosaur names to print
seed = 0
for name in range(dino_names):
# Sample indices and print them
sampled_indices = sample(parameters, char_to_ix, seed)
print_sample(sampled_indices, ix_to_char)
seed += 1 # To get the same result (for grading purposes), increment the seed by one.
print('\n')
return parameters
# -
# Run the following cell, you should observe your model outputting random-looking characters at the first iteration. After a few thousand iterations, your model should learn to generate reasonable-looking names.
parameters = model(data, ix_to_char, char_to_ix)
# ** Expected Output**
#
# The output of your model may look different, but it will look something like this:
#
# ```Python
# Iteration: 34000, Loss: 22.447230
#
# Onyxipaledisons
# Kiabaeropa
# Lussiamang
# Pacaeptabalsaurus
# Xosalong
# Eiacoteg
# Troia
# ```
# ## Conclusion
#
# You can see that your algorithm has started to generate plausible dinosaur names towards the end of the training. At first, it was generating random characters, but towards the end you could see dinosaur names with cool endings. Feel free to run the algorithm even longer and play with hyperparameters to see if you can get even better results. Our implementation generated some really cool names like `maconucon`, `marloralus` and `macingsersaurus`. Your model hopefully also learned that dinosaur names tend to end in `saurus`, `don`, `aura`, `tor`, etc.
#
# If your model generates some non-cool names, don't blame the model entirely--not all actual dinosaur names sound cool. (For example, `dromaeosauroides` is an actual dinosaur name and is in the training set.) But this model should give you a set of candidates from which you can pick the coolest!
#
# This assignment had used a relatively small dataset, so that you could train an RNN quickly on a CPU. Training a model of the english language requires a much bigger dataset, and usually needs much more computation, and could run for many hours on GPUs. We ran our dinosaur name for quite some time, and so far our favorite name is the great, undefeatable, and fierce: Mangosaurus!
#
# <img src="images/mangosaurus.jpeg" style="width:250;height:300px;">
# ## 4 - Writing like Shakespeare
#
# The rest of this notebook is optional and is not graded, but we hope you'll do it anyway since it's quite fun and informative.
#
# A similar (but more complicated) task is to generate Shakespeare poems. Instead of learning from a dataset of Dinosaur names you can use a collection of Shakespearian poems. Using LSTM cells, you can learn longer term dependencies that span many characters in the text--e.g., where a character appearing somewhere a sequence can influence what should be a different character much much later in the sequence. These long term dependencies were less important with dinosaur names, since the names were quite short.
#
#
# <img src="images/shakespeare.jpg" style="width:500;height:400px;">
# <caption><center> Let's become poets! </center></caption>
#
# We have implemented a Shakespeare poem generator with Keras. Run the following cell to load the required packages and models. This may take a few minutes.
from __future__ import print_function
from keras.callbacks import LambdaCallback
from keras.models import Model, load_model, Sequential
from keras.layers import Dense, Activation, Dropout, Input, Masking
from keras.layers import LSTM
from keras.utils.data_utils import get_file
from keras.preprocessing.sequence import pad_sequences
from shakespeare_utils import *
import sys
import io
# To save you some time, we have already trained a model for ~1000 epochs on a collection of Shakespearian poems called [*"The Sonnets"*](shakespeare.txt).
# Let's train the model for one more epoch. When it finishes training for an epoch---this will also take a few minutes---you can run `generate_output`, which will prompt asking you for an input (`<`40 characters). The poem will start with your sentence, and our RNN-Shakespeare will complete the rest of the poem for you! For example, try "Forsooth this maketh no sense " (don't enter the quotation marks). Depending on whether you include the space at the end, your results might also differ--try it both ways, and try other inputs as well.
#
# +
print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
model.fit(x, y, batch_size=128, epochs=1, callbacks=[print_callback])
# -
# Run this cell to try with different inputs without having to re-train the model
generate_output()
# The RNN-Shakespeare model is very similar to the one you have built for dinosaur names. The only major differences are:
# - LSTMs instead of the basic RNN to capture longer-range dependencies
# - The model is a deeper, stacked LSTM model (2 layer)
# - Using Keras instead of python to simplify the code
#
# If you want to learn more, you can also check out the Keras Team's text generation implementation on GitHub: https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py.
#
# Congratulations on finishing this notebook!
# **References**:
# - This exercise took inspiration from <NAME>'s implementation: https://gist.github.com/karpathy/d4dee566867f8291f086. To learn more about text generation, also check out Karpathy's [blog post](http://karpathy.github.io/2015/05/21/rnn-effectiveness/).
# - For the Shakespearian poem generator, our implementation was based on the implementation of an LSTM text generator by the Keras team: https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py
|
courses/Sequence Models/WEEK 1/Dinosaurus_Island_Character_level_language_model_final_v3a.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # First Deep Learning Model
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.datasets.samples_generator import make_circles
X, y = make_circles(n_samples=1000,
noise=0.1,
factor=0.2,
random_state=0)
X
X.shape
plt.figure(figsize=(5, 5))
plt.plot(X[y==0, 0], X[y==0, 1], 'ob', alpha=0.5)
plt.plot(X[y==1, 0], X[y==1, 1], 'xr', alpha=0.5)
plt.xlim(-1.5, 1.5)
plt.ylim(-1.5, 1.5)
plt.legend(['0', '1'])
plt.title("Blue circles and Red crosses")
from keras.models import Sequential
from keras.layers import Dense
from keras.optimizers import SGD
model = Sequential()
model.add(Dense(4, input_shape=(2,), activation='tanh'))
model.add(Dense(1, activation='sigmoid'))
model.compile(SGD(lr=0.5), 'binary_crossentropy', metrics=['accuracy'])
model.fit(X, y, epochs=20)
hticks = np.linspace(-1.5, 1.5, 101)
vticks = np.linspace(-1.5, 1.5, 101)
aa, bb = np.meshgrid(hticks, vticks)
ab = np.c_[aa.ravel(), bb.ravel()]
c = model.predict(ab)
cc = c.reshape(aa.shape)
plt.figure(figsize=(5, 5))
plt.contourf(aa, bb, cc, cmap='bwr', alpha=0.2)
plt.plot(X[y==0, 0], X[y==0, 1], 'ob', alpha=0.5)
plt.plot(X[y==1, 0], X[y==1, 1], 'xr', alpha=0.5)
plt.xlim(-1.5, 1.5)
plt.ylim(-1.5, 1.5)
plt.legend(['0', '1'])
plt.title("Blue circles and Red crosses")
|
course/1 First Deep Learning Model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Telen VanBarel construction to solve for roots.
# +
#Local imports
import TVB_Method.root_finder as rf
import TVB_Method.cheb_class as Cheb
#python imports
from matplotlib import pyplot as plt
import numpy as np
from scipy.io import loadmat
# + slideshow={"slide_type": "slide"}
# Enter the desired dim and degree.
deg = 7
dim = 3 # number of polys should equal degree so that the zero locus is
# discrete. (with probability 1)
# Create random Chebyshev polys of the desired the degree and dim.
polys = Cheb.polyList(deg,dim, 'random')
#find the roots
# %time zeros = rf.roots(polys)
rf.check_zeros(zeros,polys,tol=1e-8)
# +
# Enter the desired dim and degree.
deg = 10
dim = 3 # number of polys should equal degree so that the zero locus is
# discrete. (with probability 1)
# Create random Chebyshev polys of the desired the degree and dim.
polys = Cheb.polyList(deg,dim, 'random')
#find the roots
# %time zeros = rf.roots(polys)
rf.check_zeros(zeros,polys,tol=1e-8)
# +
# Use this cell to test the root finder.
# Enter the desired dim and degree.
deg = 30
dim = 2 # number of polys should equal degree so that the zero locus is
# discrete. (with probability 1)
# Create random Chebyshev polys of the desired the degree and dim.
polys = Cheb.polyList(deg,dim, 'random')
#find the roots
# %time zeros = rf.roots(polys)
rf.check_zeros(zeros,polys,tol=1e-8)
# -
# ## Compare TVB to Bezout in dim 2
# ### Run with TVB in Python and Bezout in Matlab
# ### Run with 8 gb of RAM and an i7 processor
domain = np.array([n for n in range(2,51)])
mat = loadmat("bezout-outer-times.mat")
Bezout_times = mat["times"][0]
TVB_times = np.load("tvb_times.npy")
# +
plt.plot(domain, TVB_times, 'b-', label="TVB")
plt.plot(domain, np.array(Bezout_times), 'g-', label="Bezout")
plt.legend(loc="upper left")
plt.xlabel("degree")
plt.ylabel("run time")
plt.show()
#plt.savefig('TvB-vs-Bezout2d.pdf')
|
CHEBYSHEV/DEMO.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: stm
# language: python
# name: stm
# ---
# +
import pickle
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import omicronscala
import spym
import xarray
import os
from pathlib import Path
from tqdm import tqdm
from multiprocessing import Pool
from functools import partial
def load_stm():
"""load stm dataset from cleaned version"""
with open('clean_stm.pkl','rb') as f:
df = pickle.load(f)
return df
def save_train_img(imgs_path, df, img_id):
"""helper to create training dataset, plot image from ID in 224x224 px"""
plt.ioff()
img = df.loc[img_id]
file = img.ImageOriginalName
ds = omicronscala.to_dataset(Path(imgs_path+file))
tf = ds.Z_Forward
tf.spym.align()
tf.spym.plane()
tf.spym.fixzero(to_mean=True)
fig = plt.figure(figsize=(4,4))
axis = plt.Axes(fig, [0., 0., 1., 1.])
axis.set_axis_off()
fig.add_axes(axis)
tf.plot(ax=axis, cmap='afmhot', add_colorbar=False)
Path('data/train').mkdir(parents=True, exist_ok=True)
plt.savefig('data/train/{}.png'.format(ID), aspect='auto', dpi=56)
plt.close()
def make_training_set(imgs_path, df):
"""sequential function that create training dataset for all images"""
for i in tqdm(range(0,len(df)), position=0, leave=True):
try:
img_id = df.iloc[i].name
imgTrain(imgs_path, stm, img_id)
except Exception as e:
print(e)
continue
def multicore_train(imgs_path, df, workers):
"""multi core training dataset creation"""
splits = np.array_split(df, workers)
pool = Pool(workers)
func = partial(make_training_set, imgs_path)
pool.map(func, splits)
pool.close()
pool.join()
def autocomplete_train(df,imgs_path):
"""checks training set, completes it if some images are missing"""
for i, row in df.iterrows():
if not os.path.isfile('{}/data/train/{}.png'.format(imgs_path,i)):
try:
imgTrain(path, stm, i)
except:
print(i)
# -
stm = load_stm()
path = "path_to_images"
n_cores = 24
multicore_train(stm, path, n_cores)
autocomplete_train(stm, path)
|
jupyter_notebooks/make_training_set_multicore.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # 02. Distributed PyTorch with Horovod
# In this tutorial, you will train a PyTorch model on the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset using distributed training via [Horovod](https://github.com/uber/horovod).
# ## Prerequisites
# * Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning (AML)
# * Go through the [00.configuration.ipynb](https://github.com/Azure/MachineLearningNotebooks/blob/master/00.configuration.ipynb) notebook to:
# * install the AML SDK
# * create a workspace and its configuration file (`config.json`)
# * Review the [tutorial](https://aka.ms/aml-notebook-pytorch) on single-node PyTorch training using the SDK
# +
# Check core SDK version number
import azureml.core
print("SDK version:", azureml.core.VERSION)
# -
# ## Initialize workspace
#
# Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`.
# +
from azureml.core.workspace import Workspace
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
# -
# ## Create a remote compute target
# You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) to execute your training script on. In this tutorial, you create an [Azure Batch AI](https://docs.microsoft.com/azure/batch-ai/overview) cluster as your training compute resource. This code creates a cluster for you if it does not already exist in your workspace.
#
# **Creation of the cluster takes approximately 5 minutes.** If the cluster is already in your workspace this code will skip the cluster creation process.
# +
from azureml.core.compute import ComputeTarget, BatchAiCompute
from azureml.core.compute_target import ComputeTargetException
# choose a name for your cluster
cluster_name = "gpucluster"
try:
compute_target = ComputeTarget(workspace=ws, name=cluster_name)
print('Found existing compute target.')
except ComputeTargetException:
print('Creating a new compute target...')
compute_config = BatchAiCompute.provisioning_configuration(vm_size='STANDARD_NC6',
autoscale_enabled=True,
cluster_min_nodes=0,
cluster_max_nodes=4)
# create the cluster
compute_target = ComputeTarget.create(ws, cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# Use the 'status' property to get a detailed status for the current cluster.
print(compute_target.status.serialize())
# -
# The above code creates a GPU cluster. If you instead want to create a CPU cluster, provide a different VM size to the `vm_size` parameter, such as `STANDARD_D2_V2`.
# ## Train model on the remote compute
# Now that we have the cluster ready to go, let's run our distributed training job.
# ### Create a project directory
# Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script and any additional files your training script depends on.
# +
import os
project_folder = './pytorch-distr-hvd'
os.makedirs(project_folder, exist_ok=True)
# -
# Copy the training script `pytorch_horovod_mnist.py` into this project directory.
import shutil
shutil.copy('pytorch_horovod_mnist.py', project_folder)
# ### Create an experiment
# Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed PyTorch tutorial.
# +
from azureml.core import Experiment
experiment_name = 'pytorch-distr-hvd'
experiment = Experiment(ws, name=experiment_name)
# -
# ### Create a PyTorch estimator
# The AML SDK's PyTorch estimator enables you to easily submit PyTorch training jobs for both single-node and distributed runs. For more information on the PyTorch estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-pytorch).
# +
from azureml.train.dnn import PyTorch
estimator = PyTorch(source_directory=project_folder,
compute_target=compute_target,
entry_script='pytorch_horovod_mnist.py',
node_count=2,
process_count_per_node=1,
distributed_backend='mpi',
use_gpu=True)
# -
# The above code specifies that we will run our training script on `2` nodes, with one worker per node. In order to execute a distributed run using MPI/Horovod, you must provide the argument `distributed_backend='mpi'`. Using this estimator with these settings, PyTorch, Horovod and their dependencies will be installed for you. However, if your script also uses other packages, make sure to install them via the `PyTorch` constructor's `pip_packages` or `conda_packages` parameters.
# ### Submit job
# Run your experiment by submitting your estimator object. Note that this call is asynchronous.
run = experiment.submit(estimator)
print(run.get_details())
# ### Monitor your run
# You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes.
from azureml.train.widgets import RunDetails
RunDetails(run).show()
# Alternatively, you can block until the script has completed training before running more code.
run.wait_for_completion(show_output=True) # this provides a verbose log
|
training/02.distributed-pytorch-with-horovod/02.distributed-pytorch-with-horovod.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#TextCNN" data-toc-modified-id="TextCNN-1"><span class="toc-item-num">1 </span>TextCNN</a></span><ul class="toc-item"><li><span><a href="#notes:" data-toc-modified-id="notes:-1.1"><span class="toc-item-num">1.1 </span>notes:</a></span></li></ul></li><li><span><a href="#LSTM" data-toc-modified-id="LSTM-2"><span class="toc-item-num">2 </span>LSTM</a></span></li></ul></div>
# + [markdown] id="a73ce9a4"
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#TextCNN" data-toc-modified-id="TextCNN-1"><span class="toc-item-num">1 </span>TextCNN</a></span><ul class="toc-item"><li><span><a href="#notes:" data-toc-modified-id="notes:-1.1"><span class="toc-item-num">1.1 </span>notes:</a></span></li></ul></li><li><span><a href="#LSTM" data-toc-modified-id="LSTM-2"><span class="toc-item-num">2 </span>LSTM</a></span></li></ul></div>
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 8643, "status": "ok", "timestamp": 1639083811240, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="5lXX-_poA-la" outputId="9e3bea65-1b7e-4d23-cda0-3f7a5f8ef3b7"
from google.colab import drive
drive.mount('/content/drive')
import os
os.chdir("/content/drive/MyDrive/Text-Classification/code")
# !pip install pyLDAvis
# !pip install gensim
# !pip install pandas==1.3.0
import nltk
nltk.download('punkt')
nltk.download('stopwords')
# + id="Y4ipJF1JA-mh"
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2257, "status": "ok", "timestamp": 1639083813489, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="8db79286" outputId="81ad274d-d6f0-429c-99fb-d0489424b1d1"
import numpy as np
from sklearn import metrics
from clustering_utils import *
from eda_utils import *
from nn_utils_keras import *
from sklearn.model_selection import train_test_split
from tensorflow.keras.utils import to_categorical
####################################
### string normalized
####################################
from gensim.utils import tokenize
from nltk.tokenize import word_tokenize
from gensim.parsing.preprocessing import remove_stopwords
def normal_string(x):
x = remove_stopwords(x)
# x = " ".join(preprocess_string(x))
x = " ".join(word_tokenize(x, preserve_line=False)).strip()
return x
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3186, "status": "ok", "timestamp": 1639083816667, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="e726484b" outputId="6f67d250-f6e7-450f-af3b-7285c885895c"
train, test = load_data()
train, upsampling_info = upsampling_train(train)
train_text, train_label = train_augmentation(train, select_comb=[['text'], ['reply', 'reference_one'], ['Subject', 'reference_one', 'reference_two']])
# train_text, train_label = train_augmentation(train, select_comb=None)
test_text, test_label = test['text'], test['label']
# test_text = test_text.apply(lambda x: normal_string(x))
# train_text = train_text.apply(lambda x: normal_string(x))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 5, "status": "ok", "timestamp": 1639083816667, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="f834796a" outputId="7d283b1e-a9e7-443a-dd95-ef17ec8d82d8"
####################################
### label mapper
####################################
labels = sorted(train_label.unique())
label_mapper = dict(zip(labels, range(len(labels))))
train_label = train_label.map(label_mapper)
test_label = test_label.map(label_mapper)
y_train = train_label
y_test = test_label
print(train_text.shape)
print(test_text.shape)
print(train_label.shape)
print(test_label.shape)
print(labels)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 13495, "status": "ok", "timestamp": 1639083830160, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="a380c6a4" outputId="dcfafd41-cfa0-414b-8991-8822a4574ddb"
####################################
### hyper params
####################################
filters = '"#$%&()*+,-/:;<=>@[\\]^_`{|}~\t\n0123465789!.?\''
MAX_NB_WORDS_ratio = 0.98
MAX_DOC_LEN_ratio = 0.999
MAX_NB_WORDS = eda_MAX_NB_WORDS(train_text, ratio=MAX_NB_WORDS_ratio, char_level=False, filters=filters)
MAX_DOC_LEN = eda_MAX_DOC_LEN(train_text, ratio=MAX_DOC_LEN_ratio, char_level=False, filters=filters)
# + id="a019380f"
from tensorflow.keras import optimizers
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.layers import Embedding, Dense, Conv1D, MaxPooling1D, Dropout, Activation, Input, Flatten, Concatenate, Lambda
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.utils import to_categorical
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from tensorflow import keras
import numpy as np
import pandas as pd
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
import os
# + [markdown] id="7e47544a"
# # TextCNN
# + [markdown] id="2db5da6d"
# ## notes:
# + colab={"base_uri": "https://localhost:8080/", "height": 764} executionInfo={"elapsed": 10300, "status": "ok", "timestamp": 1639083840453, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="1b020a50" outputId="be63679c-132b-4858-bebb-bbec79c54467"
####################################
### train val test split
####################################
X_train_val, y_train_val, X_test, y_test = train_text, train_label, test_text, test_label
X_train, x_val, y_train, y_val = train_test_split(X_train_val, y_train_val, test_size=0.2, stratify=y_train_val)
####################################
### preprocessor for NN input
####################################
processor = text_preprocessor(MAX_DOC_LEN, MAX_NB_WORDS, train_text, filters='"#$%&()*+,-/:;<=>@[\\]^_`{|}~\t\n0123465789')
X_train = processor.generate_seq(X_train)
x_val = processor.generate_seq(x_val)
X_test = processor.generate_seq(X_test)
# y_train = to_categorical(y_train)
# y_val = to_categorical(y_val)
# y_test = to_categorical(y_test)
print('Shape of x_tr: ' + str(X_train.shape))
print('Shape of y_tr: ' + str(y_train.shape))
print('Shape of x_val: ' + str(x_val.shape))
print('Shape of y_val: ' + str(y_val.shape))
print('Shape of X_test: ' + str(X_test.shape))
print('Shape of y_test: ' + str(y_test.shape))
info = pd.concat([y_train.value_counts(), y_val.value_counts(), y_val.value_counts()/y_train.value_counts(), y_train.value_counts()/y_train.size\
, y_test.value_counts(), y_test.value_counts()/y_test.size], axis=1)
info.index = labels
info.columns = ['tr_size', 'val_size', 'val_ratio', 'tr_prop', 'test_size', 'test_prop']
info
# + id="90736596"
# define Model for classification
def model_Create(FS, NF, EMB, MDL, MNW, PWV=None, optimizer='RMSprop', trainable_switch=True):
cnn_box = cnn_model_l2(FILTER_SIZES=FS, MAX_NB_WORDS=MNW, MAX_DOC_LEN=MDL, EMBEDDING_DIM=EMB,
NUM_FILTERS=NF, PRETRAINED_WORD_VECTOR=PWV, trainable_switch=trainable_switch)
# Hyperparameters: MAX_DOC_LEN
q1_input = Input(shape=(MDL,), name='q1_input')
encode_input1 = cnn_box(q1_input)
# half_features = int(len(FS)*NF/2)*10
x = Dense(384, activation='relu', name='half_features')(encode_input1)
x = Dropout(rate=0.3, name='dropout1')(x)
# x = Dense(256, activation='relu', name='dense1')(x)
# x = Dropout(rate=0.3, name='dropou2')(x)
x = Dense(128, activation='relu', name='dense2')(x)
x = Dropout(rate=0.3, name='dropout3')(x)
x = Dense(64, activation='relu', name='dense3')(x)
x = Dropout(rate=0.3, name='dropout4')(x)
pred = Dense(len(labels), activation='softmax', name='Prediction')(x)
model = Model(inputs=q1_input, outputs=pred)
model.compile(optimizer=optimizer,
loss=keras.losses.SparseCategoricalCrossentropy(),
metrics=[keras.metrics.SparseCategoricalAccuracy()])
return model
EMBEDDING_DIM = 200
# W2V = processor.w2v_pretrain(EMBEDDING_DIM, min_count=2, seed=1, cbow_mean=1,negative=5, window=20, workers=7) # pretrain w2v by gensim
# W2V = processor.load_glove_w2v(EMBEDDING_DIM) # download glove
W2V = None
trainable_switch = True
# + id="9821d103"
# Set hyper parameters
FILTER_SIZES = [2, 4,6,8]
# FILTER_SIZES = [2,3,4]
NUM_FILTERS = 64
# OPT = optimizers.Adam(learning_rate=0.005)
OPT = optimizers.RMSprop(learning_rate=0.0005) # 'RMSprop'
PWV = W2V
model = model_Create(FS=FILTER_SIZES, NF=NUM_FILTERS, EMB=EMBEDDING_DIM,
MDL=MAX_DOC_LEN, MNW=MAX_NB_WORDS+1, PWV=PWV,
optimizer=OPT, trainable_switch=trainable_switch)
# + id="87bcd69a"
# visual_textCNN(model)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2314708, "status": "ok", "timestamp": 1639090229043, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="cebc0600" outputId="3efafc37-6e0e-46bc-9dce-980bad8fc1a6"
BATCH_SIZE = 32 # 先在小的batch上train, 容易找到全局最优部分, 然后再到 大 batch 上train, 快速收敛到局部最优
NUM_EPOCHES = 50 # 20步以上
patience = 30
file_name = 'test'
BestModel_Name = file_name + 'Best_GS_3'
BEST_MODEL_FILEPATH = BestModel_Name
# model.load_weights(BestModel_Name) # 这样就能接着上次train
earlyStopping = EarlyStopping(monitor='val_sparse_categorical_accuracy', patience=patience, verbose=1, mode='max') # patience: number of epochs with no improvement on monitor : val_loss
checkpoint = ModelCheckpoint(BEST_MODEL_FILEPATH, monitor='val_sparse_categorical_accuracy', verbose=1, save_best_only=True, mode='max')
# history = model.fit(X_train, y_train, validation_data=(X_test,y_test), batch_size=BATCH_SIZE, epochs=NUM_EPOCHES, callbacks=[earlyStopping, checkpoint], verbose=1)
history = model.fit(X_train, y_train, validation_data=(x_val, y_val), batch_size=BATCH_SIZE, epochs=NUM_EPOCHES, callbacks=[earlyStopping, checkpoint], verbose=1)
model.load_weights(BestModel_Name)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 21053, "status": "ok", "timestamp": 1639090250094, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="a8e44226" outputId="8a2091f6-13f9-4ec6-81fb-5e8c30543f08"
#### classification Report
history_plot(history)
y_pred = model.predict(X_test)
# print(classification_report(y_test, np.argmax(y_pred, axis=1)))
print(classification_report(test_label, np.argmax(y_pred, axis=1), target_names=labels))
scores = model.evaluate(X_test, y_test, verbose=2)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
print( "\n\n\n")
# + id="2Fk4bV6JK_jW"
# + executionInfo={"elapsed": 2, "status": "ok", "timestamp": 1639090250095, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="DGb4iQJtK_kf"
# + executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1639090250096, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="156e307e"
# + [markdown] id="fbff5a0a"
# # LSTM
# + executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1639090250096, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="a176a6d2"
# from tensorflow.keras.layers import SpatialDropout1D, GlobalMaxPooling1D, GlobalMaxPooling2D
# def model_Create(FS, NF, EMB, MDL, MNW, PWV = None, optimizer='RMSprop', trainable_switch=True):
# model = Sequential()
# model.add(Embedding(input_dim=MNW, output_dim=EMB, embeddings_initializer='uniform', mask_zero=True, input_length=MDL))
# model.add(Flatten())
# # model.add(GlobalMaxPooling2D()) # downsampling
# # model.add(SpatialDropout1D(0.2))
# model.add(Dense(1024, activation='relu'))
# model.add(Dense(512, activation='relu'))
# model.add(Dense(128, activation='relu'))
# model.add(Dense(64, activation='relu'))
# # model.add(LSTM(100, dropout=0.2, recurrent_dropout=0.2))
# model.add(Dense(20, activation='softmax'))
# model.compile(optimizer=optimizer,
# loss=keras.losses.SparseCategoricalCrossentropy(from_logits=False),
# metrics=[keras.metrics.SparseCategoricalAccuracy()])
# return model
# model = model_Create(FS=FILTER_SIZES, NF=NUM_FILTERS, EMB=EMBEDDING_DIM,
# MDL=MAX_DOC_LEN, MNW=MAX_NB_WORDS+1, PWV=PWV, trainable_switch=trainable_switch)
# + executionInfo={"elapsed": 2, "status": "ok", "timestamp": 1639090250240, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="bd56494e"
# visual_textCNN(model)
# + executionInfo={"elapsed": 2, "status": "ok", "timestamp": 1639090250241, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="d4e1631d"
# EMBEDDING_DIM = 200
# # W2V = processor.w2v_pretrain(EMBEDDING_DIM, min_count=2, seed=1, cbow_mean=1,negative=5, window=20, workers=7) # pretrain w2v by gensim
# # W2V = processor.load_glove_w2v(EMBEDDING_DIM) # download glove
# trainable_switch = True
# W2V = None
# + executionInfo={"elapsed": 2, "status": "ok", "timestamp": 1639090250241, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="352e76ba"
# BATCH_SIZE = 64
# NUM_EPOCHES = 10 # patience=20
# patience = 30
# BestModel_Name = 'text_CNN.h5'
# BEST_MODEL_FILEPATH = BestModel_Name
# earlyStopping = EarlyStopping(monitor='val_sparse_categorical_accuracy', patience=patience, verbose=1, mode='max') # patience: number of epochs with no improvement on monitor : val_loss
# checkpoint = ModelCheckpoint(BEST_MODEL_FILEPATH, monitor='val_sparse_categorical_accuracy', verbose=1, save_best_only=True, mode='max')
# history = model.fit(X_train, y_train, validation_split=0.2, batch_size=BATCH_SIZE, epochs=NUM_EPOCHES, callbacks=[earlyStopping, checkpoint], verbose=1)
# model.load_weights(BestModel_Name)
# + executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1639090250242, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="0453d996"
# #### classification Report
# history_plot(history)
# y_pred = model.predict(X_test)
# # print(classification_report(y_test, np.argmax(y_pred, axis=1)))
# print(classification_report(test_label, np.argmax(y_pred, axis=1), target_names=labels))
# scores = model.evaluate(X_test, y_test, verbose=2)
# print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# print( "\n\n\n")
# + executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1639090250242, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="95a094bc"
# + executionInfo={"elapsed": 4, "status": "ok", "timestamp": 1639090250243, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="4d1dcb7d"
|
code/.ipynb_checkpoints/NN_based_models_v4-3-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Solution Notebook
# ## Problem: Given a list of tuples representing ranges, condense the ranges.
#
# Example: [(2, 3), (3, 5), (7, 9), (8, 10)] -> [(2, 5), (7, 10)]
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# ## Constraints
#
# * Are the tuples in sorted order?
# * No
# * Are the tuples ints?
# * Yes
# * Will all tuples have the first element less than the second?
# * Yes
# * Is there an upper bound on the input range?
# * No
# * Is the output a list of tuples?
# * Yes
# * Is the output a new array?
# * Yes
# * Can we assume the inputs are valid?
# * No, check for None
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# <pre>
# * None input -> TypeError
# * [] - []
# * [(2, 3), (7, 9)] -> [(2, 3), (7, 9)]
# * [(2, 3), (3, 5), (7, 9), (8, 10)] -> [(2, 5), (7, 10)]
# * [(2, 3), (3, 5), (7, 9), (8, 10), (1, 11)] -> [(1, 11)]
# * [(2, 3), (3, 8), (7, 9), (8, 10)] -> [(2, 10)]
# </pre>
# ## Algorithm
#
# * Sort the tuples based on start time
# * Check each adjacent tuple to see if they can be merged
#
# <pre>
# Case: * [(2, 3), (3, 8), (7, 9), (8, 10)] -> [(2, 10)]
#
# * Sort by start time (already sorted)
# * Add the first tuple to the merged_array
# * Loop through each item in sorted_array starting at index 1
# * If there is no overlap
# * Add the current item to merged_array
# * Else
# * Update the last item in merged_array
# * The end time will be the max of merged_array[-1][1] and sorted_array[i][1]
#
# Start:
# i
# 0 1 2 3
# sorted_array = [(2, 3), (3, 8), (7, 9), (8, 10)]
# merged_array = [(2, 3)]
#
# Overlap with (2, 3), (3, 8):
# i
# 0 1 2 3
# sorted_array = [(2, 3), (3, 8), (7, 9), (8, 10)]
# merged_array = [(2, 8)]
#
# Overlap with (2, 8), (7, 9):
# i
# 0 1 2 3
# sorted_array = [(2, 3), (3, 8), (7, 9), (8, 10)]
# merged_array = [(2, 9)]
#
# Overlap with (2, 9) (8, 10):
# i
# 0 1 2 3
# sorted_array = [(2, 3), (3, 8), (7, 9), (8, 10)]
# merged_array = [(2, 10)]
# </pre>
#
# Complexity:
# * Time: O(n log(n))
# * Space: O(n)
# ## Code
class Solution(object):
def merge_ranges(self, array):
if array is None:
raise TypeError('array cannot be None')
if not array:
return array
sorted_array = sorted(array)
merged_array = [sorted_array[0]]
for index, item in enumerate(sorted_array):
if index == 0:
continue
start_prev, end_prev = merged_array[-1]
start_curr, end_curr = item
if end_prev < start_curr:
# No overlap, add the entry
merged_array.append(item)
else:
# Overlap, update the previous entry's end value
merged_array[-1] = (start_prev, max(end_prev, end_curr))
return merged_array
# ## Unit Test
# +
# %%writefile test_merge_ranges.py
import unittest
class TestMergeRanges(unittest.TestCase):
def test_merge_ranges(self):
solution = Solution()
self.assertRaises(TypeError, solution.merge_ranges, None)
self.assertEqual(solution.merge_ranges([]), [])
array = [(2, 3), (7, 9)]
expected = [(2, 3), (7, 9)]
self.assertEqual(solution.merge_ranges(array), expected)
array = [(3, 5), (2, 3), (7, 9), (8, 10)]
expected = [(2, 5), (7, 10)]
self.assertEqual(solution.merge_ranges(array), expected)
array = [(2, 3), (3, 5), (7, 9), (8, 10), (1, 11)]
expected = [(1, 11)]
self.assertEqual(solution.merge_ranges(array), expected)
print('Success: test_merge_ranges')
def main():
test = TestMergeRanges()
test.test_merge_ranges()
if __name__ == '__main__':
main()
# -
# %run -i test_merge_ranges.py
|
online_judges/merge_ranges/merge_ranges_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Self-Driving Car Engineer Nanodegree
#
# ## Deep Learning
#
# ## Project: Build a Traffic Sign Recognition Classifier
#
# In this notebook is an implementation of Traffic Sign Classifier using LeNet neural network for the Udacity Self Drivig Car Nanodegree program.
#
# You can find this project on [this github repo](https://github.com/Vladimir-Lazic/CarND-Traffic-Sign-Classifier-Project)
#
# Below I will adress each point in the [project rubric](https://review.udacity.com/#!/rubrics/481/view).
#
# The project files are:
#
# - `'traffic_sign_classifier_project.ipynb'` is a jupyter notebook containing the code
#
# - `'traffic_sign_classifier_project.html'` is the HTML export of the code
#
# - `'traffic_sign_classifier_project.pdf'` is the project writeup in pdf
#
# +
import pickle
import matplotlib.pyplot as plt
import random
import numpy as np
import csv
import warnings
from sklearn.utils import shuffle
import cv2
# Suppressing TensorFlow FutureWarings
with warnings.catch_warnings():
warnings.filterwarnings("ignore", category=FutureWarning)
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
# +
image_label_file = 'signnames.csv'
def parse_image_labels(input_csc_file):
reader = csv.reader(open(input_csc_file, 'r'))
retVal = {}
for row in reader:
key, value = row
if key == 'ClassId':
continue
retVal.update({int(key): value})
return retVal
# Parsing image label csv file
image_labels = parse_image_labels(image_label_file)
# -
# ## Data set
#
# In the two cells bellow several rubric points are addressed:
#
# - `'Dataset Summary'` in which I display the basic propertives of the dataset like the number of images, number of classes and image shape
#
# - `'Exploratory Visualization'` in which I display selected images of the dataset
#
# - `'Preprocessing'` where I apply preprocessing techniques to the dataset. The techniques I have implemented are dataset normalization and grayscale. My initial idea was to train the network using RGB images, so it was required that I normalize the values of image pixels in order to improve the the perscision of the network. However, after initial few tries I have opted againts this approach and went with the training the network on a grayscale input data set. This approach proved to be more effective in terms of achieving desired network percision.
#
# In the cell below I load the train, validation and tests set and apply the grayscaleing on each image for each set.
# +
"""
Functions for dataset exploration
"""
figsize_default = plt.rcParams['figure.figsize']
def samples_stat(features, labels):
h = [0 for i in range(len(labels))]
samples = {}
for idx, l in enumerate(labels):
h[l] += 1
if l not in samples:
samples[l] = features[idx]
return h, samples
def dataset_exploration(features, labels):
plt.rcParams['figure.figsize'] = (20.0, 20.0)
histo, samples = samples_stat(features, labels)
total_class = len(set(labels))
ncols = 4
nrows = 11
_, axes = plt.subplots(nrows=nrows, ncols=ncols)
class_idx = 0
for r in range(nrows):
for c in range(ncols):
a = axes[r][c]
a.axis('off')
if class_idx in samples:
a.imshow(samples[class_idx])
if class_idx in image_labels:
a.set_title("No.{} {}(#{})".format(class_idx, image_labels[class_idx], histo[class_idx]), fontsize=12)
class_idx += 1
plt.rcParams['figure.figsize'] = figsize_default
# +
training_file = './data_set/train.p'
validation_file = './data_set/valid.p'
testing_file = './data_set/test.p'
# Loading the data set
with open(training_file, mode='rb') as f:
train = pickle.load(f)
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
assert (len(X_train) == len(y_train))
assert (len(X_valid) == len(y_valid))
print()
print("Image Shape: {}".format(X_train[0].shape))
print()
print("Training Set: {} samples".format(len(X_train)))
print("Validation Set: {} samples".format(len(X_valid)))
print("Test Set: {} samples".format(len(X_test)))
def image_normalize(image):
image = np.divide(image, 255)
return image
def dataset_normalization(X_data):
X_normalized = X_data.copy()
num_examples = len(X_data)
for i in range(num_examples):
image = X_normalized[i]
normalized_image = image_normalize(image)
X_normalized[i] = normalized_image
return X_normalized
def dataset_grayscale(X_data):
X_grayscale = []
num_examples = len(X_data)
for i in range(num_examples):
image = X_data[i]
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
X_grayscale.append(gray.reshape(32, 32, 1))
return np.array(X_grayscale)
dataset_exploration(X_train, y_train)
print('Grayscaling training set')
X_train = dataset_grayscale(X_train)
X_valid = dataset_grayscale(X_valid)
X_test = dataset_grayscale(X_test)
assert (len(X_train) == len(y_train))
print("Grayscaled Training Set: {} samples".format(len(X_train)))
print("Grayscale Image Shape: {}".format(X_train[0].shape))
X_train, y_train = shuffle(X_train, y_train)
# -
# ## Neural Network
#
# In the two cells bellow several rubric points are addressed:
#
# - `'Model Architecture'` : For model architecture I have opted for standard LeNet neural network. The neural network takes an input shape of 32x32x1, which is a grayscaled image. The network contains two convolutional layers. The convolutional layers are a combination of convolution, relu activation function, max pool layer. After that we have three fully connected layers. After each respective layers I have implemented a dropout layer, for reducing network overfitting. The dropout layers have 2 different keep probabilities. One probability is used for the output of convolutional layer, and the other is used for the output of fully connected layers. The output of the network is an array of logits size 43. The network is implemented in lenet.py file.
#
# - `'Model Training'` : The training of the network was done on the input sample of the grayscaled images. The images have been preprocessed in the cells above. The network has been trained with a learing rate of 0.001 over 30 epoch with a bach size of 64. For optimizer I used Adam optimizer.
#
# - `'Solution Approach'` : After a serveral tried tunning the hyperparameters I found the best was to use a learning rate of 0.001 and the batch size of 64. For the number of epoch I found that 30 number of epoch gave the network enough tries to achieve desired validation accuracy without having to take too much time or run the risk of the network overfitting. After 30 epoch the validation accuracy that was achieved was 94.6%. After getting the desired validation accuracy the network was then introduced to the test set. On the first try running the network on the tests set the accuracy achieved was 93.2% which is a good inicator that the network was trained correctly and that it didn't overfit.
# +
from lenet import *
EPOCHS = 30
BATCH_SIZE = 64
x = tf.placeholder(tf.float32, (None, 32, 32, 1))
y = tf.placeholder(tf.int32, None)
one_hot_y = tf.one_hot(y, 43)
logits = LeNet(x)
# Training pipeline
rate = 0.001
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=one_hot_y, logits=logits)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate=rate)
training_operation = optimizer.minimize(loss_operation)
# Model evaluation
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# +
def evaluate(X_data, y_data, model='lenet'):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset + BATCH_SIZE], y_data[offset:offset + BATCH_SIZE]
accuracy = sess.run(accuracy_operation, feed_dict={x: batch_x,
y: batch_y,
keep_prob_conv: 1.0,
keep_prob: 1.0})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
def predict_single_label(x_image):
sess = tf.get_default_session()
logits_output = sess.run(tf.argmax(logits, 1),
feed_dict={
x: np.expand_dims(x_image, axis=0),
keep_prob_conv: 1.0,
keep_prob: 1.0})
classification_index = logits_output[0]
logits_return = logits_output.copy()
return image_labels[classification_index], classification_index, logits_return
def batch_predict(X_data, BATCH_SIZE=64):
num_examples = len(X_data)
batch_predict = np.zeros(num_examples, dtype=np.int32)
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x = X_data[offset:offset + BATCH_SIZE]
batch_predict[offset:offset + BATCH_SIZE] = sess.run(tf.argmax(logits, 1), feed_dict={x: batch_x, keep_prob_conv: 1.0,
keep_prob: 1.0})
return batch_predict
# +
saver = tf.train.Saver()
with tf.Session(config=tf.ConfigProto(log_device_placement=True)) as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print("Training...")
print()
for i in range(EPOCHS):
X_train, y_train = shuffle(X_train, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x,
y: batch_y,
keep_prob_conv: 1.0,
keep_prob: 0.7})
print("EPOCH {} ...".format(i + 1))
validation_accuracy = evaluate(X_valid, y_valid)
print("Validation Accuracy = {:.3f}".format(validation_accuracy))
print()
saver.save(sess, './model/lenet')
print("Model saved")
# -
# Check Test Accuracy
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('./model/.'))
test_accuracy = evaluate(X_test, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy))
# ## Model testing
#
# In the cells below I have addressed the rubric points regarding testing the model on new images.
#
# - `'Acquiring New Images'` : The model was tested on 5 new images of German Traffic signs found on the web and they are diplayed in the cell below. The images are converted to grayscale and resized to (32, 32, 1) size in order to be compatible with the trained model.
#
# - `'Performance on New Images'` : The model is evaluated on the new images found on the web. The results are displayed as a batch prediction and as single image. The model was able to recognize the new signs.
#
# - `'Model Certainty - Softmax Probabilities'` : In the very last cell I have displayed the top 5 probabilities for each of the new signs that the network has predicted. This is done using the tf.nn.top_k function, and the results are printed in the cell below.
# +
import matplotlib.image as mpimg
sign_1 = './new_images/nopassing.png' # label : 9
sign_2 = './new_images/keepleft.jpg' # label : 39
sign_3 = './new_images/70.jpg' # label : 4
sign_4 = './new_images/yield.png' # label : 13
sign_5 = './new_images/turnleft.jpg' # label : 33
new_signs = [sign_1, sign_2, sign_3, sign_4, sign_5]
sign_images = []
new_images_label = [9, 39, 4, 13, 34]
dst_size = (32,32)
for sign in new_signs:
image = cv2.imread(sign)
image = cv2.resize(image, dst_size)
sign_images.append(image)
fig = plt.figure(figsize=(150, 200))
ax = []
print('Original images')
for i in range(len(sign_images)):
ax.append(fig.add_subplot(32, 32, i+1))
plt.imshow(cv2.cvtColor(sign_images[i], cv2.COLOR_BGR2RGB))
images_grayscale = dataset_grayscale(sign_images)
# +
# Check bluk accuracy of new traffic signs
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('./model/.'))
test_accuracy = evaluate(images_grayscale, new_images_label)
print("Test Accuracy = {:.3f}".format(test_accuracy))
print('\n\nIndividual signs detection:\n\n')
# Check individual accuracy of new traffic signs
predicted_images = []
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('./model/.'))
for image in images_grayscale:
predicted_image, predicted_label, logits_result = predict_single_label(image)
print("Sign: " + predicted_image + ", label : " + str(predicted_label))
predicted_images.append(predicted_image)
# +
top_k = tf.nn.top_k(logits, k=5)
def top_five_outputs(x_image):
sess = tf.get_default_session()
top_k_output = sess.run(top_k,
feed_dict={
x: np.expand_dims(x_image, axis=0),
keep_prob_conv: 1.0,
keep_prob: 1.0})
return top_k_output
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('./model/.'))
for i in range(len(images_grayscale)):
top_five = top_five_outputs(images_grayscale[i])
print('\nFor predicted image : ' + predicted_images[i] + ' the models top five probabilities are: ')
for j in range(5):
label = top_five[1][0][j]
probability = str(top_five[0][0][j])
print("Label: " + image_labels[label] + " \nProbability of: " + probability + "%\n")
|
traffic_sign_classifier_project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
df_labels = pd.read_csv('train_labels.csv')
df_values = pd.read_csv('train_values.csv')
# -
df_labels.head()
df_values.head()
# merge the labels and values
merged_df = pd.merge(df_values, df_labels, how='inner', on='patient_id')
# +
#merged_df["heart_disease_present"]
# -
merged_df.plot(x="heart_disease_present", y="max_heart_rate_achieved", kind="bar")
merged_df.plot(x="heart_disease_present", y="sex", kind="scatter")
plt.hist(merged_df["heart_disease_present"])
# correlation matrix
merged_df.corr()
# binning your data
bins = [0, 19.50, 39.50, 59.60, 79.50, 9999]
labels = ['<20', '20-39', '40-59', '60-79', '80+']
merged_df["age_group"] = pd.cut(merged_df["age"], bins=bins, labels=labels)
merged_df.head()
# +
#drop NaN from heart disease present
merged_df_1 = merged_df.drop(merged_df.index[(merged_df.heart_disease_present.eq(0))])
merged_df_0 = merged_df.drop(merged_df.index[(merged_df.heart_disease_present.eq(1))])
# -
merged_df_0.head()
list(merged_df_1['resting_blood_pressure'])
merged_df_0.boxplot(column=["age"])
merged_df_1.plot(x="age", y="exercise_induced_angina", kind="scatter")
merged_df.plot(x="age", y="heart_disease_present", kind="scatter")
# +
#plt.hist(merged_df["heart_disease_present"])
plt.hist(merged_df["heart_disease_present"])
# -
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import numpy as np
# +
x = np.heart_disease_present
y = np.age_group
plt.plot(x, y, 'o', color='black');
# -
|
Trang/heart_disease_exploratory.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2-2 Intro Python Practice
# ## Lists
# <font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - Create Lists
# - Access items in a list
# - Add Items to the end of a list
# - Insert items into a list
# - Delete items from a list
# ## Create Lists
# [ ] create and populate list called days_of_week then print it
days_of_week = ["Sunday", "Monday", "Tuesday", "Wednesday", "Thursday", "Friday", "Saturday"]
print(days_of_week)
# [ ] after days_of_week is run above, print the days in the list at odd indexes 1,3,5
print(days_of_week[1])
print(days_of_week[3])
print(days_of_week[5])
# ## Phone letters
# 
# Create a list, **`phone_letters`**, where the index 0 - 9 contains the letters for keys 0 - 9.
#
# - 0 = ' ' (a space)
# - 1 = '' (empty)
# - 2 = 'ABC'
# - 3 = 'DEF'
# - etc...
#
# [ ] create and populate list called phone_letters then print it
phone_letters = [" ","","ABC","DEF","GHI","JKL","MNO","PQRS","TUV","WXYZ"]
print(phone_letters)
# ## Access Lists
# ### for the 2 cells below
# - Use days_of_week list created above
# - Run the cell above to make the list available
# +
# [ ] create a variable: day, assign day to "Tuesday" using days_of_week[]
# [ ] print day variable
day = days_of_week[2]
print(day)
# +
# PART 2
# [ ] assign day to days_of_week index = 5
# [ ] print day
day = days_of_week[5]
print(day)
# -
# ## Append and Insert items into a list
#
# ### Endsday, Midsday, Resterday
# #### for the exercises below
# - Use days_of_week list created above
# - Run the cell defining days_of_week above to make the list available
# +
# [ ] Make up a new day! - append an 8th day of the week to days_of_week
# [ ] print days_of_week
days_of_week.append("Funday")
print(days_of_week)
# -
# ### Question
# - What happens if you keep running the cell above?
# - How can you return to the initial state with the regular 7 days in days_of_week?
# +
# [ ] Make up another new day - insert a new day into the middle of days_of_week between Wed - Thurs
# [ ] print days_of_week
days_of_week.insert(3,"Myday")
print(days_of_week)
# +
# [ ] Extend the weekend - insert a day between Fri & Sat in the days_of_week list
# [ ] print days_of_week
days_of_week.insert(7,"Endday")
print(days_of_week)
# -
# ## Delete from a list
# ### `del` & `.pop()` some bad ideas
# #### exercises below assume days_of_week appended/inserted 3 extra days in previous exercises
# +
# [ ] print days_of_week
print(days_of_week)
# [ ] modified week is too long - pop() the last index of days_of_week & print .pop() value
print(days_of_week.pop())
# [ ] print days_of_week
print(days_of_week)
# -
# [ ] print days_of_week
print(days_of_week)
# [ ] delete (del) the new day added to the middle of the week
del days_of_week[3]
# [ ] print days_of_week
print(days_of_week)
# [ ] print days_of_week
print(days_of_week)
# [ ] programmers choice - pop() any day in days_of week & print .pop() value
print(days_of_week.pop(4))
# [ ] print days_of_week
print(days_of_week)
# ## Program: Letter to Number Function
# # TODO: insert video
# ### for the exercise below
# - Use phone_letters list created above
# - Run the cell above to make the list available
#
# #### recall unit 1 using **`in`** to search for a string in a string
# ```python
# if "e" in "open":
# print("e found")
# else:
# print("e not found")
# ```
#
# 
#
# ### create funtion let_to_num()
# - let_to_num() takes input of a single letter, space or empty string stored in an argument variable: letter
# - use `while key < 10:` to try numbers 0 - 9 as index for `phone_letters` ("key" = phone dial pad key)
# - check if `letter` variable is in the index of `phone_letters[key]`
# ```python
# key = 0
# while key < 10:
# if # Create Code: determine if letter is **`in`** any of the phone_letters[key] where key is the index 0 -9:
# return key
# else:
# key = key + 1
# return "Not Found"
# ```
# - return the number or "Not Found"
# - **call** let_to_num() to test the function so it prints the argument and return value with:
# - space
# - lowercase letter
# - different letter, uppercase
# - a number
#
# **Bonus**: create a special case to check if empty string (`""`) was submitted
# the problem is that an empty string will be found in all strings as
# ```python
# if "" in "ABC":
# ```
# is True, and is true for any phone_letters, but should `return 1`
# +
# [ ] create let_to_num()
phone_letters = [' ', '', 'ABC', 'DEF', 'GHI', 'JKL', 'MNO', 'PQRS', 'TUV', 'WXYZ']
def let_to_num():
letter = input("Enter a letter (or space):\n")
key = 0
while key < 10:
if letter.upper() in phone_letters[key]:
return key
else:
key = key + 1
return ("Not found")
print(let_to_num())
# -
# ## Challenge: reverse a string
# ### using
# - while
# - .pop()
# - insert()
#
# **`pop()`** the **first item** in the list and
# +
# [ ] Challenge: write the code for "reverse a string"
#not sure what to do here - seems like something may be missing from the instructions
# -
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
|
Python Fundamentals/Module_2_Practice_Python_Fundamentals.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>The median and average home values across all Boston Suburbs in dollars.</h1>
# <h3>July 16, 2017</h3>
# <h3><NAME></h3>
# 1. Title: Boston Housing Data
#
# 2. Sources:
# (a) Origin: This dataset was taken from the StatLib library which is
# maintained at Carnegie Mellon University.
# (b) Creator: <NAME>. and <NAME>. 'Hedonic prices and the
# demand for clean air', J. Environ. Economics & Management,
# vol.5, 81-102, 1978.
# (c) Date: July 7, 1993
#
# 3. Past Usage:
# - Used in Belsley, Kuh & Welsch, 'Regression diagnostics ...', Wiley,
# 1980. N.B. Various transformations are used in the table on
# pages 244-261.
# - Quinlan,R. (1993). Combining Instance-Based and Model-Based Learning.
# In Proceedings on the Tenth International Conference of Machine
# Learning, 236-243, University of Massachusetts, Amherst. Morgan
# Kaufmann.
#
# 4. Relevant Information:
#
# Concerns housing values in suburbs of Boston.
#
# 5. Number of Instances: 506
#
# 6. Number of Attributes: 13 continuous attributes (including "class"
# attribute "MEDV"), 1 binary-valued attribute.
#
# 7. Attribute Information:
#
# 1. CRIM per capita crime rate by town
# 2. ZN proportion of residential land zoned for lots over
# 25,000 sq.ft.
# 3. INDUS proportion of non-retail business acres per town
# 4. CHAS Charles River dummy variable (= 1 if tract bounds
# river; 0 otherwise)
# 5. NOX nitric oxides concentration (parts per 10 million)
# 6. RM average number of rooms per dwelling
# 7. AGE proportion of owner-occupied units built prior to 1940
# 8. DIS weighted distances to five Boston employment centres
# 9. RAD index of accessibility to radial highways
# 10. TAX full-value property-tax rate per $10,000
# 11. PTRATIO pupil-teacher ratio by town
# 12. B 1000(Bk - 0.63)^2 where Bk is the proportion of blacks
# by town
# 13. LSTAT % lower status of the population
# 14. MEDV Median value of owner-occupied homes in $1000's
#
# 8. Missing Attribute Values: None.
#
# sourced from University of California Irvine - Machine Learning Repository
#
# https://archive.ics.uci.edu/ml/machine-learning-databases/housing/housing.names.
# <h3>Analyze Data Set:</h3>
# CSV Data sourced from University of California Irvine - Machine Learning Repository
# https://archive.ics.uci.edu/ml/datasets/Housing
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import pandas as pd
housing_data = pd.read_csv('boston_housing_data.csv', delimiter="\s+", names=('CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO', 'B', 'LSTAT', 'MEDV'))
housing_data
# <h3>The median and average home values across all Boston Suburbs in dollars.</h3>
data_frame = pd.DataFrame(housing_data)
'Median value of all homes in $1000\'s is ${:0.2f}'.format(data_frame['MEDV'].median() * 1000)
'Mean value of all homes in $1000\'s is ${:0.2f}'.format(data_frame['MEDV'].mean() * 1000)
# <h3>The median home value of the suburb with the newest houses.</h3>
newest = housing_data.loc[housing_data['AGE'].idxmin()]
'Median value of newest homes in $1000\'s is ${:0.2f}'.format(newest['MEDV']*1000)
# <h3>The relationship between per-capita crime rate and the pupil-teacher ratio. Differentiate between whether or not the suburb is bounded by the Charles River.</h3>
bound = housing_data[housing_data['CHAS']==1].filter(['CRIM', 'PTRATIO'])
unbound = housing_data[housing_data['CHAS']==0].filter(['CRIM', 'PTRATIO'])
plt.scatter(
unbound['PTRATIO'],
unbound['CRIM'],
c='red',
label='Suburb not on the Charles River'
)
plt.scatter(
bound['PTRATIO'],
bound['CRIM'],
c='blue',
label='Suburbs on Charles River'
)
plt.xlabel('Pupil to Teacher ratio')
plt.ylabel('Crime Rate per Capita (%)')
plt.title('Crime rate versus pupil to teacher ratio, bounded by Charles River')
plt.legend(loc='upper left')
plt.show()
# <h3>The relationship between the proportion of black citizens and the distance to employment centers in Boston.</h3>
BK = (housing_data.B / (1000**(1/2))) + 0.63
plt.scatter(housing_data['DIS'], BK)
plt.xlabel('Weighted distance to employment center')
plt.ylabel('Proportion of Black people (%)')
plt.title("Relation of black citizens and distance to employment centers")
plt.show()
# <h3>The relationship between median value of owner-occuped homes and nitric oxide concentration along with median home value and the proportion of non-retail business (on the same plot).</h3>
plt.scatter(housing_data['MEDV'], housing_data['NOX'], c='blue', label='Nitric Oxides')
plt.scatter(housing_data['MEDV'], housing_data['INDUS'], c='red', label='Non-Retail Business')
plt.xlabel('Median value of homes')
plt.ylabel('Nitric Oxides Concentration Proportion of Non-Retail Business')
plt.legend(loc='upper right')
plt.title("N.O.X. concentration proportion of Non-Retail Business")
plt.show()
# <h3>The relationship between per capita crime rate and median value of homes.</h3>
plt.scatter(housing_data['MEDV'], housing_data['CRIM'], c='red', label='Non-Retail Business')
plt.xlabel('Median value of homes')
plt.ylabel('Crime Per Capita')
plt.legend(loc='upper right')
plt.title("Median value of homes in relation to Crime Rate per Capita")
plt.show()
|
analyze_data_set.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pybaseball
df = pybaseball.batting_stats_bref(2009)
data_dir = 'data'
fname = '2018_stats.csv'
data_path = data_dir + '/' + fname
print(data_path)
df.to_csv(data_path, index=False)
|
statsmerge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from __future__ import absolute_import
from __future__ import print_function
import numpy as np
np.random.seed(1337)
from keras.preprocessing import sequence
from keras.optimizers import SGD, RMSprop, Adagrad
from keras.utils import np_utils
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation
from keras.layers.embeddings import Embedding
from keras.layers.recurrent import LSTM, GRU
from keras.datasets import imdb
from keras import backend as K
from theano import function
# -
print("Loading data...")
max_features = 5000
(X_train, y_train), (X_test, y_test) = imdb.load_data(nb_words = max_features)
print(len(X_train), 'train sequences')
print(len(X_test), 'test sequences')
print("Pad sequences(samples x time)")
maxlen = 500
X_train = sequence.pad_sequences(X_train, maxlen = maxlen)
X_test = sequence.pad_sequences(X_test, maxlen = maxlen)
print("X_train shape:", X_train.shape)
print("X_test shape:", X_test.shape)
print("Build model..")
model = Sequential()
model.add(Embedding(max_features, 128, input_length = maxlen))
model.add(LSTM(128))
model.add(Dropout(0.5))
model.add(Dense(1))
model.add(Activation('sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer = 'adam',
metrics=["accuracy"])
print("Train..")
batch_size = 30
score = model.fit(X_train, y_train, batch_size = batch_size,
nb_epoch = 4, validation_data = (X_test, y_test))
# +
import matplotlib.pyplot as plt
plt.plot(score.history['acc'])
plt.plot(score.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(score.history['loss'])
plt.plot(score.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# -
X_train[0]
word_index = imdb.get_word_index()
index_word = {v:k for k,v in word_index.items()}
type(index_word.keys()[0])
index_word[0] = '0'
' '.join(index_word[w] for w in X_train[0])
X_train_words = []
for sentence in X_train:
X_train_words += [[index_word[w] for w in sentence if w != "0"]]
import gensim
import logging
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
w2v_model = gensim.models.Word2Vec(X_train_words, min_count=1)
X_train_words[0]
# w2v_model.wv[u'wonderful']
# w2v_model.wv['lucas']
# [index_word[w] for w in X_train[0]]
# w2v_model.wv.most_similar(positive=['woman', 'king'], negative=['man'])
w2v_model.wv.similarity('good', 'spielberg')
w2v_model.wv.similarity('good', 'tarantino')
from tempfile import mkstemp
fs, temp_path = mkstemp('word2vec_model_may9')
w2v_model.save(temp_path)
w2v_model.accuracy('questions-words.txt')
fs, temp_path = mkstemp('word2vec_model_may9')
w2v.model = gensim.models.Word2Vec.load('word2vec_model_may9')
from gensim.corpora import WikiCorpus
import pandas as pd
train = pd.read_csv( "labeledTrainData.tsv", header=0,
delimiter="\t", quoting=3 )
test = pd.read_csv( "testData.tsv", header=0, delimiter="\t", quoting=3 )
unlabeled_train = pd.read_csv( "unlabeledTrainData.tsv", header=0,
delimiter="\t", quoting=3 )
# Verify the number of reviews that were read (100,000 in total)
print (train["review"].size, test["review"].size, unlabeled_train["review"].size)
from bs4 import BeautifulSoup
import re
from nltk.corpus import stopwords
def review_to_wordlist( review, remove_stopwords=False ):
# Function to convert a document to a sequence of words,
# optionally removing stop words. Returns a list of words.
#
# 1. Remove HTML
review_text = BeautifulSoup(review).get_text()
#
# 2. Remove non-letters
review_text = re.sub("[^a-zA-Z]"," ", review_text)
#
# 3. Convert words to lower case and split them
words = review_text.lower().split()
#
# 4. Optionally remove stop words (false by default)
if remove_stopwords:
stops = set(stopwords.words("english"))
words = [w for w in words if not w in stops]
#
# 5. Return a list of words
return(words)
# +
# Download the punkt tokenizer for sentence splitting
import nltk.data
nltk.download()
# Load the punkt tokenizer
tokenizer = nltk.data.load('tokenizers/punkt/english.pickle')
# Define a function to split a review into parsed sentences
def review_to_sentences( review, tokenizer, remove_stopwords=False ):
# Function to split a review into parsed sentences. Returns a
# list of sentences, where each sentence is a list of words
#
# 1. Use the NLTK tokenizer to split the paragraph into sentences
raw_sentences = tokenizer.tokenize(review.strip())
#
# 2. Loop over each sentence
sentences = []
for raw_sentence in raw_sentences:
# If a sentence is empty, skip it
if len(raw_sentence) > 0:
# Otherwise, call review_to_wordlist to get a list of words
sentences.append( review_to_wordlist( raw_sentence, \
remove_stopwords ))
#
# Return the list of sentences (each sentence is a list of words,
# so this returns a list of lists
return sentences
# +
sentences = [] # Initialize an empty list of sentences
print("Parsing sentences from training set")
for review in train["review"]:
sentences += review_to_sentences(review, tokenizer)
print("Parsing sentences from unlabeled set")
for review in unlabeled_train["review"]:
sentences += review_to_sentences(review, tokenizer)
# -
# # Opções para o experimento
#
# ## Vetores
#
# * Usar vetores pre-treinados
# ([word2vec](http://mccormickml.com/2016/04/12/googles-pretrained-word2vec-model-in-python/)
# [GloVe](https://nlp.stanford.edu/projects/glove/))
# * Treinar vetores em dados do IMDB sem remover stop-words
# * Treinar vetores em outros corpora
#
# ## Intrinsic evaluation
#
# * Usar google questions
# * Criar questions baseadas no domínio (filmes, adjetivos, etc, personagens!!, diretores e atrizes, outras analogias, gênero e ator, ver distâncias --> ver a interface web para escolher as perguntas) <--------
# * Seguir modelo do artigo do [GloVe](https://nlp.stanford.edu/pubs/glove.pdf) (Pennington et al. 2014)
# * Use [20 News groups](http://scikit-learn.org/stable/datasets/twenty_newsgroups.html) from scikit
#
# ## Extrinsic evaluation
#
# * Sentiment analysis IMDB
# * Named entity recognition - [CoNLL 2013](https://cogcomp.cs.illinois.edu/page/resource_view/81); [reference](https://github.com/monikkinom/ner-lstm)
#
|
models/.ipynb_checkpoints/Untitled-checkpoint.ipynb
|
# Written by <NAME> for CoderDojo Twin Cities - www.coderdojotc.org
def write ( minecraft, text, material ):
letters = { 'A' : [[1,0,0,1],[1,0,0,1],[1,1,1,1],[1,0,0,1],[0,1,1,0]],
'B' : [[1,1,1,0],[1,0,0,1],[1,1,1,0],[1,0,0,1],[1,1,1,0]],
'C' : [[0,1,1,0],[1,0,0,1],[1,0,0,0],[1,0,0,1],[0,1,1,0]],
'D' : [[1,1,1,0],[1,0,0,1],[1,0,0,1],[1,0,0,1],[1,1,1,0]],
'E' : [[1,1,1,1],[1,0,0,0],[1,1,1,1],[1,0,0,0],[1,1,1,1]],
'F' : [[1,1,1,1],[1,0,0,0],[1,1,1,1],[1,0,0,0],[1,0,0,0]],
'G' : [[1,1,1,0],[1,0,0,1],[1,0,0,0],[1,0,1,1],[0,1,1,0]],
'H' : [[1,0,0,1],[1,0,0,1],[1,1,1,1],[1,0,0,1],[1,0,0,1]],
'I' : [[1,1,1],[0,1,0],[0,1,0],[0,1,0],[1,1,1]],
'J' : [[0,1,1,0],[1,0,0,1],[0,0,0,1],[0,0,0,1],[0,0,0,1]],
'K' : [[1,0,0,1],[1,0,1,0],[1,1,0,0],[1,0,1,0],[1,0,0,1]],
'L' : [[1,1,1,1],[1,0,0,0],[1,0,0,0],[1,0,0,0],[1,0,0,0]],
'M' : [[1,0,0,0,1],[1,0,0,0,1],[1,0,1,0,1],[1,1,0,1,1],[1,0,0,0,1]],
'N' : [[1,0,0,0,1],[1,0,0,1,1],[1,0,1,0,1],[1,1,0,0,1],[1,0,0,0,1]],
'O' : [[0,1,1,0],[1,0,0,1],[1,0,0,1],[1,0,0,1],[0,1,1,0]],
'P' : [[1,0,0,0],[1,0,0,0],[1,1,1,1],[1,0,0,1],[1,1,1,1]],
'Q' : [[0,0,1,1],[0,1,1,0],[1,0,0,1],[1,0,0,1],[0,1,1,0]],
'R' : [[1,0,0,1],[1,0,1,0],[1,1,1,1],[1,0,0,1],[1,1,1,1]],
'S' : [[1,1,1,0],[0,0,0,1],[0,1,1,0],[1,0,0,0],[0,1,1,1]],
'T' : [[0,1,0],[0,1,0],[0,1,0],[0,1,0],[1,1,1]],
'U' : [[0,1,1,0],[1,0,0,1],[1,0,0,1],[1,0,0,1],[1,0,0,1]],
'V' : [[0,0,1,0,0],[0,1,0,1,0],[0,1,0,1,0],[1,0,0,0,1],[1,0,0,0,1]],
'W' : [[0,1,0,1,0],[1,0,1,0,1],[1,0,1,0,1],[1,0,0,0,1],[1,0,0,0,1]],
'X' : [[1,0,0,0,1],[0,1,0,1,0],[0,0,1,0,0],[0,1,0,1,0],[1,0,0,0,1]],
'Y' : [[0,0,1,0,0],[0,0,1,0,0],[0,1,0,1,0],[1,0,0,0,1],[1,0,0,0,1]],
'Z' : [[1,1,1,1,1],[0,1,0,0,0],[0,0,1,0,0],[0,0,0,1,0],[1,1,1,1,1]],
' ' : [[0,0,0],[0,0,0],[0,0,0],[0,0,0],[0,0,0]]}
# Get the player's current position
pos = minecraft.player.getPos()
row = 0
kearning = 0
for letter in text.upper():
print letter
while row < len( letters[letter] ):
col = 0
while col < len( letters[letter][0] ):
# If a block should be printed in that row/column for a
# given letter, print it
if letters[letter][row][col]:
minecraft.setBlock( pos.x - col - kearning, pos.y + row + 1, pos.z, material )
col = col + 1
row = row + 1
# Reset the row and col for each letter
row = 0
# Adjust the spacing based on how big the letter is since this is
# not a fixed width font
kearning = kearning + len( letters[letter][0] ) + 1
|
classroom-code/examples/minecraft_letters.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Keyboard Shortcuts
# + language="html"
# <style>
# table { float: left; }
# table th, table td { font-size: 150%; }
# </style>
# -
# | Navigating cells ||
# |:---------|:------------------|
# | `up / j` | select cell above |
# | `down / k` | select cell below |
# | `y` | change cell to code mode |
# | `m` | change cell to markdown |
# | `enter` | enter edit mode |
# | `esc` | exit edit mode |
# | Managing cells ||
# |:---------------|:-------------------------|
# | `a` | insert cell above |
# | `b` | insert cell below |
# | `c` | copy cell |
# | `v` | paste cell |
# | `d,d` | delete dell|
# | `shift + (up / j)` | extend selection above |
# | `shift + (down / k)` | extend selection below |
# | `shift + m` | merge selected cells |
# | Executing cells ||
# |:---------------|:-----------------------|
# | `ctrl + enter` | run cell |
# | `shift + enter` | run cell, select below |
# | `alt + enter` | run cell, insert below |
# | Edit mode ||
# |:---------|:------------------|
# | `tab` | suggest code completion |
# | `ctrl + shift + -` | split cell in half |
# | `esc` | exit edit mode |
|
docs/notebooks/00 Keyboard Shortcuts.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="9w0LU6S7GGEr"
# # 算法分析
#
# **为啥要做算法分析?**
#
# - 我们需要有一种方式来谈论和选择不同的算法
#
# - 足够"粗糙"以避免架构, 语言, 编译器等等系统因素的干扰
# + [markdown] colab_type="text" id="fJqclpHgYoiE"
# ## 看一个问题
#
# 在有序列表s中找出x
# + [markdown] colab_type="text" id="zDKdlGP6bS9S"
# ### 定义第一种算法
#
# 从第一个开始依次往上找, 直到找到x, 否则x在队列中不存在.
#
# 这种办法叫 **简单查找**, 也叫**线性查找 (linear search)**
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="q_4C4xghZzuN"
def linear_search(x, sequence):
"""在sequence中查找x, 并返回其下标, 没有就返回空(python中叫做None, 其他语言如C++叫做NULL)
:param x: 待查找的数字
:param sequence: 需要找x的队列
:return: x的下标或者None
"""
for i, val in enumerate(sequence):
if val == x:
return i
return None
# + [markdown] colab_type="text" id="V0kAM7oAcqn2"
# 试验一下:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 1804, "status": "ok", "timestamp": 1526293867512, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-sQ952bLluIc/AAAAAAAAAAI/AAAAAAAAAAs/GbMfis6C_o8/s50-c-k-no/photo.jpg", "userId": "112844315788872411035"}, "user_tz": -480} id="8XoE6dLMcpRX" outputId="7531889c-c16a-4c51-d6d4-7f88371d8fc4"
# 定义队列s
s = list(range(10_000_000))
# 列出s中的前10项
print(*s[:5], sep=', ')
x = 1_000_000
idx = linear_search(x, s)
print("找到的答案为:", idx)
# + [markdown] colab_type="text" id="tAFF1XNsebQK"
# 这种办法的问题:
#
# 在每次检查中都只排除了一个数字, 如果想找的恰好是列表中最后一个, 那么你需要检查整个列表
# + [markdown] colab_type="text" id="kVIODRNqfET4"
# ### 更好的办法
#
# 下面是一种更好的办法, 从中间开始. 通过判断它与x的大小关系就能确定列表中一半的元素与x的大小关系, 从而每次检查都能排除当前队列中一半的元素, 从而快速锁定x该在的位置.
#
# 这种办法叫**二分查找 (binary search)**
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="2BOnPf8mdGiK"
def binary_search(x, sequence):
current_range_lower, current_range_upper = 0, len(sequence) - 1
center = int((current_range_lower + current_range_upper) / 2)
center_val = sequence[center]
while center_val != x:
if current_range_lower >= current_range_upper:
return None
if center_val > x:
current_range_upper = center - 1
else:
current_range_lower = center + 1
center = int((current_range_lower + current_range_upper) / 2)
center_val = sequence[center]
return center
# + [markdown] colab_type="text" id="ndHr6EDTjjDK"
# 试验一下新方法:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1667, "status": "ok", "timestamp": 1526294129548, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-sQ952bLluIc/AAAAAAAAAAI/AAAAAAAAAAs/GbMfis6C_o8/s50-c-k-no/photo.jpg", "userId": "112844315788872411035"}, "user_tz": -480} id="AkoG5uQ3iixn" outputId="3628b065-b6c4-4c43-8673-b3205e7969e1"
s = [i * 2 for i in range(1000000)]
idx = binary_search(999997, s)
print("找到的答案为:", idx)
# + [markdown] colab_type="text" id="sAVZGnZvk0As"
# 比较两种方法的结果
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 1612, "status": "ok", "timestamp": 1526294197093, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-sQ952bLluIc/AAAAAAAAAAI/AAAAAAAAAAs/GbMfis6C_o8/s50-c-k-no/photo.jpg", "userId": "112844315788872411035"}, "user_tz": -480} id="EYM9fMs-k6rU" outputId="2f6c5e05-207a-40cf-effc-ca6eaa5319b3"
# 制造一个100万长度的偶数序列
sequence = [i * 2 for i in range(1_000_000)]
# 随机创建一个查找任务
import random
x = random.randint(0, 2_000_000)
# 比较两个算法的结果
result_1 = linear_search(x, sequence)
result_2 = binary_search(x, sequence)
print("任务x={}:\n线性查找: {}, 二分查找: {}".format(x, result_1, result_2))
# + [markdown] colab_type="text" id="Q2m5xdS0l-Xl"
# 比较两种方法所消耗的时间
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 4201, "status": "ok", "timestamp": 1526294270963, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-sQ952bLluIc/AAAAAAAAAAI/AAAAAAAAAAs/GbMfis6C_o8/s50-c-k-no/photo.jpg", "userId": "112844315788872411035"}, "user_tz": -480} id="kpFLqd-wmEJD" outputId="d155684e-42f9-4eb2-9289-d9770c8d4b9f"
"""Jupyter notebook 中的magic commands"""
# %timeit \
# _ = linear_search(x, sequence)
# %timeit \
# _ = binary_search(x, sequence)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 1815, "status": "ok", "timestamp": 1526290781557, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-sQ952bLluIc/AAAAAAAAAAI/AAAAAAAAAAs/GbMfis6C_o8/s50-c-k-no/photo.jpg", "userId": "112844315788872411035"}, "user_tz": -480} id="MwXwuZXkpIo-" outputId="fa0bda06-ad09-4468-de2f-de55e7aab691"
"""常规实现方法"""
from datetime import datetime
# 第一种方法所消耗的时间
tick = datetime.now()
_ = linear_search(x, sequence)
tock = datetime.now()
time_1 = (tock - tick).total_seconds()
# 第二种方法所消耗的时间
tick = datetime.now()
_ = binary_search(x, sequence)
tock = datetime.now()
time_2 = (tock - tick).total_seconds()
print("任务x={}:\n线性查找: {}秒, 二分查找: {}秒".format(x, time_1, time_2))
# + [markdown] colab_type="text" id="4nLgiNINj01G"
# ## 评估算法
#
# 对于这个问题, 一旦保证算法是正确的, 那么重要的一步就是确定该算法将需要多少时间或空间等资源量的问题. 如果一个问题的求解算法需要长达一年时间, 那么这种算法就很难在应用中得到体现.
#
# 估计算法资源消耗所需的分析一般来说是一个理论问题, 我们永远无法准确地计算出实际所需的时间, 因此需要一套正式的系统构架
#
# ### 操作数
#
# 我们定义$T(n)$为算法执行的操作数, 其中$n$为算法中数据的规模.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="YSYPiDrgisFL"
a = 0 # 操作数=1
# 操作数=1000 * (1 + 1) = 2000
for i in range(1000):
a = 0
b = 1
# + [markdown] colab_type="text" id="RXs7Amd3tFpp"
# linear_search中问题的规模为列表的长度, 设为$n$. 那么linear_search的操作数:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="hNRFoWW5nOxY"
def linear_search(x, sequence):
for i, val in enumerate(sequence):
if val == x: # 操作数 = 2 (条件运算1 + 判断运算1)
return i # 操作数 = 1
return None # 操作数 = 1
# + [markdown] colab_type="text" id="Ooy6BK_ZuELQ"
# 总操作数如下:
# $$T_{LS}(n) \leq n(2+1) + 1 \approx 3n + 1$$
#
# ***思考*** 为什么这里是小于等于的关系?
# + [markdown] colab_type="text" id="2FPtS5kUubVn"
# binary_search的操作数为:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="3X9BqV91utzy"
def binary_search(x, sequence):
current_range_lower, current_range_upper = 0, len(sequence) - 1 # 计算长度1 + 赋值1 + 赋值1 = 3
center = int((current_range_lower + current_range_upper) / 2) # 加法1 + 除法1 + 取整1 + 赋值1 = 4
center_val = sequence[center] # 取值1 + 赋值1 = 2
while center_val != x:
if current_range_lower >= current_range_upper: # 条件计算1 + 判断1
return None # 返回1
if center_val > x: # 2
current_range_upper = center - 1 # 2
else:
current_range_lower = center + 1 # 2
center = int((current_range_lower + current_range_upper) / 2) # 4
center_val = sequence[center] # 2
return center # 1
# + [markdown] colab_type="text" id="HUCCYBs2vXtD"
# $$
# T_{BS}(n) = 3 + 4 + 2 + (2 + 1 + 2 + 2 + 2 + 4 + 2)log_2{n} + 1 = 15log_2n+10
# $$
#
# ***思考*** 为什么这里是$log_2n$
# + [markdown] colab_type="text" id="mr07LhE-wEks"
# ## 比较T(n)
#
# 在科学计算中一般使用2为对数的底, 但在程序设计中默认的对数运算为自然对数$log_en$, 所以需要使用到对数的换底公式
# $$
# log_ab={log_cb \over log_ca}
# $$
#
# 以n为自变量, 画出$T_{LS}(n)$与$T_{BS}(n)$
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 265} colab_type="code" executionInfo={"elapsed": 1715, "status": "ok", "timestamp": 1526290790438, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-sQ952bLluIc/AAAAAAAAAAI/AAAAAAAAAAs/GbMfis6C_o8/s50-c-k-no/photo.jpg", "userId": "112844315788872411035"}, "user_tz": -480} id="yRHLjtsVv_dF" outputId="f72e5a63-d949-42f0-ad58-c45510bdae1f"
# %matplotlib inline
from matplotlib import pyplot as plt
import numpy as np
def log(a, b):
return np.log(b) / np.log(a)
def T_LS(n):
return 3 * n + 1
def T_BS(n):
return 15 * log(2, n) + 10
n = np.array([i + 1 for i in range(1, 100, 1)])
t1 = T_LS(n)
t2 = T_BS(n)
plt.plot(n, t1, '#000000')
plt.plot(n, t2, '#ff0000')
plt.show()
# + [markdown] colab_type="text" id="-ehdY0E7yxtg"
# 我们发现当$n=20$时, t1 < t2, 当n=40时, t1 > t2. 且t1的增长速度远大于t2.
#
# ### 启示
#
# 单纯比较$T(n)$意义并不大, 而应该比较他们的**相对增长率**. 很显然, 从上面的例子中能看出二者的**相对增长率**有非常大的差别, 但缺乏一种系统的表达方式来描述这一差异.
# + [markdown] colab_type="text" id="zdHiWwd6GqK9"
# ## 时间复杂度 - The big O notation
#
#
# ### 一些数学定义
#
# 1. 如果存在正常数$c$和$n_0$使得当$N \geq n_0$时, $T(N) \leq cf(N)$, 则记
# $$T(N)=O(f(N))$$
#
# 2. 如果存在正常数$c$和$n_0$使得当$N \geq n_0$时, $T(N) \geq cf(N)$, 则记
# $$T(N)=\Omega(f(N))$$
#
# 3. 若$T(N)=O(f(N))$且$T(n)=\Omega(f(N))$, 则记
# $$T(N)=\Theta(f(N))$$
#
# **解读**
#
# 1. $T(N)$的增长率小于等于$f(N)$的增长率
#
# 2. $T(N)$的增长率大于等于$f(N)$的增长率
#
# 3. $T(N)$的增长率等于$f(N)$的增长率
#
# ### 一些例子
#
# * $T(n)=3n^2+4n+1=O(n^2)$
#
# * $T(n)=4(n+1) + \sqrt{n-1} + 10000=O(n)$
#
# * $T(n)=2log_2(3n)+log_3n=O(logn)$
#
# ### 一些规律总结
#
# * 如果$T(n)$是一个$k$次多项式, 那么$T(n)=\Theta(N^k)$
#
# * 如果$T_1(N)=O(f(N)), T_2(N)=O(g(N))$, 那么:
#
# * $T_1(N)+T_2(N)=max\{ O(f(N)), O(g(N)) \}$
# * $T_1(N) * T_2(N) = O(f(n) \times g(n))$
#
# * 对任意常数$k, log^kN=O(N)$
#
# 当某算法的操作数$T(n)=\Theta(f(n))$时, 我们称该算法的时间复杂度为$\Theta(f(n))$.
#
# 在实际使用和习惯中, 由于$T(n)=O(f(n))$含义为$f(n)$是$T(n)$的上限, 我们更多时候会将$\Theta$替换为$O$, 称该算法的时间复杂度为$O(f(n))$.
#
# 例如:
#
# 我们常说快速排序算法的时间复杂度为$O(n log{n})$, 实际上快速排序的$T(n)=\Theta(n log n)$
#
# ### 一句话求复杂度
#
# "消去常数, 只保留最高次项"
# + [markdown] colab_type="text" id="3rvXt0TpwOyb"
# ## The Master Theorem (a.k.a the Master Method) 主定理
#
# 在评估递归算法的时间复杂度时, 我们往往无法直接写出该算法的操作数函数$T(n)$, 但可以写成如下递推式:
#
# $$T(n) = aT({n \over b}) + O(n^d)$$
#
# ### 解读
#
# - a = 递归调用次数 (a ≥ 1)
# - b = 数据规模缩小速度 (b > 1)
# - d = 递归调用外对当前数据的处理相对于当前规模$n$的复杂度指数
# - a, b, d皆与$n$独立
#
# ### 公式
#
# $$
# T(n)=
# \begin{cases}
# O(n^dlogn)& ,\text{if } a=b^d\\
# O(n^d)& ,\text{if } a < b^d\\
# O(n^{log_ba})& ,\text{if } a > b^d
# \end{cases}
# $$
#
# ### 举例
#
# #### 二分查找
#
# $a=1, b=2, d=0, b^d=1=a$, 属于Case 1, $T(n)=O(n^dlogn)=O(n^0logn)=O(logn)$
#
# #### 快速排序
#
# $a = 2, b = 2, d = 1, b^d=2=a$, 属于Case 1, $T(n)=O(n^dlogn)=O(n^1logn)=O(nlogn)$
#
# #### 二叉树遍历
#
# $a = 2, b = 2, d = 0, b^d=1<a$, 属于Case 3, $T(n)=O(n^{log_ba})=O(n^{log_22})=O(n^1)=O(n)$
# + [markdown] colab_type="text" id="70LSEyPssKi2"
# ## 算法的比较
#
# 假设有两个算法, 其复杂度分别为$T_1(n)=O(f(n))$和$T_2(n)=O(g(n))$. 比较它们的时间效率方法如下:
#
# $$
# 相对增长率r=\lim_{n\to\infty} {f(n) \over g(n)}
# $$
#
# **注意** 当 $\lim_{n\to\infty} f(n) = \infty$且$\lim_{x\to\infty} g(n)=\infty$时, 需应用l'Hospital's rule.
#
# ### L' Hospital's Rule - 洛必达法则
#
# $$
# \lim_{n\to\infty} {f(n) \over g(n)} = \lim_{n\to\infty} {f'(n) \over g'(n)}
# $$
#
# 增长率$r$的范围所含意义如下:
#
# * $r = 0$: 算法1比2快
# * $r=c\neq0$: 两个算法复杂度一样
# * $r=\infty$: 算法1比2慢
# * $r$不收敛: 两个算法无关
|
lectures/lecture0.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import json
jsonfile="ebike-5min-ride.json"
#jsonfile="standard-5min-ride.json"
with open(jsonfile) as f:
dataraw=json.load(f)
# +
actions=[]
timestamps=[]
paths=[]
payloads=[]
for item in dataraw:
actions.append(item['action'])
timestamps.append(item['timestamp'])
paths.append(item['path'])
payloads.append(item['payload'])
df=pd.DataFrame(data={"actions":actions,"timestamps":timestamps,"paths":paths,"payloads":payloads})
#df.head(50)
# -
ps=df.paths.unique()
ps
# +
#df.loc[df['paths']=='rideService/speed',:]
# +
#df.loc[df["paths"]=="mobile/location",'payloads']
# +
rsspeedi=[]
userPoweri=[]
ridingDurationi=[]
ridingDistancei=[]
ascenti=[]
caloriesi=[]
heartRatei=[]
cadencei=[]
distancei=[]
averageSpeedi=[]
rangei=[]
tsi=[]
def state2arrays(state):
rsspeedi.append(state['rideService/speed'])
userPoweri.append(state['rideService/userPower'])
ridingDurationi.append(state['tourService/ridingDuration'])
ridingDistancei.append(state['tourService/ridingDistance'])
ascenti.append(state['tourService/ascent'])
caloriesi.append(state['tourService/calories'])
heartRatei.append(state['rideService/heartRate'])
cadencei.append(state['rideService/cadence'])
distancei.append(state['motor/distance'])
averageSpeedi.append(state['tourService/averageSpeed'])
rangei.append(state['motor/range'])
tsi.append(state['ts'])
state={}
state['oldts']=0
state['ts']=0
state['rideService/speed']=0
state['rideService/userPower']=0
state['tourService/ridingDuration']=0
state['tourService/ridingDistance']=0
state['tourService/ascent']=0
state['tourService/calories']=0
state['rideService/heartRate']=0
state['rideService/cadence']=0
state['motor/distance']=0
state['tourService/averageSpeed']=0
state['motor/range']=0
badpaths= ['rideService/heartRateAvailability',
'rideService/cadenceAvailability', 'mobile/locationAvailability',
'hub/motorInterfaceId', 'mobile/location']
for i,row in df.iterrows():
if row['paths'] not in badpaths:
state['oldts']=state['ts']
state['ts']=row['timestamps']
state[row['paths']]=row['payloads']
if state['oldts']!=state['ts']:
state2arrays(state)
# -
datastate={'rsspeed':rsspeedi,
'userPower':userPoweri,
'ridingDuration':ridingDurationi,
'ridingDistance':ridingDistancei,
'ascent':ascenti,
'calories':caloriesi,
'heartRate':heartRatei,
'cadence':cadencei,
'distance':distancei,
'averageSpeed':averageSpeedi,
'range':rangei,
'ts':tsi}
dfstate=pd.DataFrame(data=datastate)
dfstate.tail(50)
dfstate.shape
print(dfstate.columns)
dfstate.cadence.unique()
keepcols=['rsspeed', 'userPower', 'ridingDuration', 'ridingDistance', 'ascent',
'calories', 'heartRate', 'cadence', 'averageSpeed']
outdf=dfstate.loc[500:,keepcols]
outdf.to_csv("cobi_state_data2.csv",index=False, header=False)
outdf.describe()
rsspeedmax=10.
userPowermax=100.
ridingDurationmax=500.
ridingDistancemax=2000.
ascentmax=15.
caloriesmax=50.
heartratemax=150.
cadencemax=90.
averageSpeedmax=8.
|
examples/cobi_bike/processing_cobi_data.ipynb
|
/ -*- coding: utf-8 -*-
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: SQL
/ language: sql
/ name: SQL
/ ---
/ + [markdown] azdata_cell_guid="e1f157c0-51ef-4c65-8e09-3707be8fb303"
/ # BI Metrics in Power BI vs. SQL
/
/ The aim of this notebook is to demonstrate the savings we have in productivity when we use DAX in Power BI (or SSAS) to calculate complex metrics, instead of T-SQL (e.g. YoY and YTD metrics). For this reason we're going to use the [World Wide Importers DW](https://docs.microsoft.com/en-us/sql/samples/wide-world-importers-dw-install-configure?view=sql-server-ver15). I've installed SQL Server on Ubuntu using the instructions [here](https://docs.microsoft.com/en-us/sql/linux/quickstart-install-connect-ubuntu?view=sql-server-ver15), and Azure Data Studio using the instructions [here](https://docs.microsoft.com/en-us/sql/azure-data-studio/download-azure-data-studio?view=sql-server-ver15). Then imported the database as a .bak file, and created this notebook in Azure Data Studio. At the same time, I've used Power BI Desktop on a Windows VM, and imported all relevant tables there:
/ + azdata_cell_guid="6b7fc779-b59c-4b45-81ef-45b5c6bc58a6"
use WideWorldImportersDW
select schema_name(t.schema_id) as schema_name,
t.name as table_name
from sys.tables t
where schema_name(t.schema_id) IN ('Dimension','Fact')
order by t.schema_id,table_name;
/ + [markdown] azdata_cell_guid="aa8823a4-c27b-47b2-8f6f-2f2a956e878f"
/ Power BI will detect the relationships automatically, as there are foreign keys defined in the database. The part of the data model which is relevant to our example is the below:
/
/ 
/
/ We also need to mark the 'Dimension.Date' table as a 'Date' table in Power BI:
/
/ 
/
/ and sort months by month number:
/
/ 
/ + [markdown] azdata_cell_guid="e68cfc79-501a-4ecf-a441-5c93b4c49788"
/ To calculate YTD (Year To Date) sales per brand for each month of 2014, we'd write something like this:
/ + azdata_cell_guid="dd7c5c5c-0b3a-4b79-ac03-9a86618fbc78"
select
Color
,[Month]
,FORMAT(SUM(Revenue) OVER (PARTITION BY color ORDER BY [Month No] ), '##,##0.##') AS [YTD Sales per Color]
,FORMAT(Revenue,'##,##0.##') [Sales per Color]--for ref
FROM
(select P.[Color] Color
,D.[Calendar Month Label] [Month]
,D.[Calendar Month Number] [Month No]
,sum(F.[Total Excluding Tax]) Revenue
from
fact.Sale F
JOIN dimension.[Stock Item] P on F.[Stock Item Key]=P.[Stock Item Key]
JOIN dimension.[Date] D on F.[Invoice Date Key]=D.[Date]
where year([invoice date key])=2014
group by P.[Color], D.[Calendar Month Number],D.[Calendar Month Label]
) NESTED
order by color,[Month No]
/ + [markdown] azdata_cell_guid="808826f3-87a5-454d-8c74-dd0e75c55e5b"
/ If we'd like to so the same for 2015, we'd need to change the where clause. If we'd like to group for another attribute, we'd need to replace \[Stock Item\].\[Color\] with that attribute (e.g. \[Stock Item\].Brand). If the attribute belonged to another dimension, we'd have to change the query completely.
/
/ On the other hand, on Power BI, we'd only need to define this DAX measure: `Sales YTD = TOTALYTD([Sales],'Dimension Date'[Date])`
/
/ having defined measure `Sales = SUM('Fact Sale'[Total Excluding Tax])` to improve readability.
/
/ Then we could create any report we'd like with YTD sales, for any year, for any customer, for any product... That would be achieved by simple drag-and-drop, for any of these attributes and any visual. We could also add filters and groupings at will. For example two visuals showing YTD sales per color and brand could be the following:
/
/ 
/
/ 
/
/ Similarly, to calculate YoY (Year on Year) percentage sales increase between 2014 and 2013, we would do something like this:
/ + azdata_cell_guid="179e7ffa-191a-4a11-a0f3-c792c3d46e79"
select
Color
,(sales - lag1)/lag1 as [YoY% Sales]
FROM (
select
Color
,[Invoice Year]
,Revenue as Sales
,Lag(Revenue,1,0) OVER (PARTITION BY color ORDER BY [Invoice Year]) as Lag1
FROM (
select P.[Color] Color
,year(D.Date) as [Invoice Year]
,sum(F.[Total Excluding Tax]) Revenue
from
fact.Sale F
JOIN dimension.[Stock Item] P on F.[Stock Item Key]=P.[Stock Item Key]
JOIN dimension.[Date] D on F.[Invoice Date Key]=D.[Date]
where year([invoice date key]) IN (2013,2014)
group by P.[Color], year(D.Date)
) NESTED
) NESTED1
where [Invoice Year]=2014
order by color
/ + [markdown] azdata_cell_guid="79214fbd-729f-4306-b3e5-dbcbc51f38cb"
/ The DAX measure is also simple: `Sales YoY% = DIVIDE([Sales]-[Sales Last Year],[Sales Last Year])` also having defined `Sales Last Year = CALCULATE([Sales],SAMEPERIODLASTYEAR('Dimension Date'[date]))` for readability.
/
/ and can be used to create visuals involving additional attributes, without requiring other code:
/
/ 
/
/ 
/ + [markdown] azdata_cell_guid="3cd77a80-ea31-479e-ad94-35c31a97d0ba"
/ It should be clear by now that, using just a couple of DAX measures, we can create any sort of visualizations, filtering and grouping by any attribute we want. Even if we add dimensions and attributes in our model, DAX measures we have already created will work for the new attributes as well. In SQL we would need to create additional queries for each and every filter / group combination we might need.
/
/
/
/ Consequently, using the correct tool for the correct problem, we can save significantly amount of development time and, as a matter of fact, improve execution performance. At the same time we enable self service reporting, as the person who designs the report or performs an analysis using visualizations, does not need to be the same person who wites the DAX.
|
TSQL/BI Metrics in PoBI vs SQL.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R [conda env:miniconda3-so-co2-R]
# language: R
# name: conda-env-miniconda3-so-co2-R-r
# ---
# + [markdown] tags=[]
# # Process airborne PFP merge files
# - R Program to read in HIPPO, ORCAS, and ATom PFP merge files, massage time variables, add a strat flag, and subset
# - then filter aircraft data for strong local continental influences, subtract off NOAA in situ SPO, and write out flat text files
# - have to run process_aircraft_10s.ipynb first
# + tags=[]
library('ncdf4')
library('yaml')
# + tags=[]
project_tmpdir_obs = read_yaml('../_config_calc.yml')$project_tmpdir_obs
username = Sys.info()['user']
project_tmpdir_obs = gsub('\\{\\{env\\[\'USER\'\\]\\}\\}', username, project_tmpdir_obs)
# + tags=[]
# specify aircraft data file names
hippomergefile=paste(project_tmpdir_obs,'/aircraft-merge-products/HIPPO_noaa_flask_allparams_merge_insitu_20121129.tbl',sep='') # this is the official HIPPO merge product
atommergedir=paste(project_tmpdir_obs,'/aircraft-merge-products',sep='') # these are version 2.0 (21-08-26)
atommergefiles=c('MER-PFP_DC8_ATom-2.nc','MER-PFP_DC8_ATom-3.nc','MER-PFP_DC8_ATom-4.nc') # no ATom-1 PFP data
# no ORCAS PFP data
# -
# set strat flag cutoffs for use below based upon Jin et al., 2021 (https://doi.org/10.5194/acp-21-217-2021)
stratcoh2o=50
stratcoo3=150
stratcon2o=319
# read in global N2O for detrending aircraft N2O
glbn2ofile=url('ftp://aftp.cmdl.noaa.gov/products/trends/n2o/n2o_annmean_gl.txt')
hlines=61
glbn2o=read.table(glbn2ofile,skip=hlines,header=F,stringsAsFactors=F)
colnames(glbn2o)=c('year','n2o','unc')
# for HIPPO, need to interpolate prof variable from 10-sec merge - load from process_aircraft_10s.ipynb output before reading PFP data below
load('HIPPO_10s.RData')
hippomerge10s=hippomerge # 'hippomerge' reused below
hippo10sdt=ISOdatetime(hippomerge$year,hippomerge$mon,hippomerge$day,hippomerge$hour,hippomerge$min,hippomerge$sec,tz='UTC')
# # read in HIPPO file, calc strat flag, and subset
# + tags=[]
# read and add time variables
hippomerge=read.table(hippomergefile,header=T)
hippodt=strptime(paste(hippomerge[,"Year"],hippomerge[,"DOY"]),format='%Y %j',tz='UTC')+hippomerge[,"UTC"] # DOY is day of year of takeoff; UTC is seconds since midnight on day of takeoff
hippomerge$Month=as.POSIXlt(hippodt)$mon+1
hippomerge$Day=as.POSIXlt(hippodt)$mday
hippomerge$Hour=as.POSIXlt(hippodt)$hour
hippomerge$Min=as.POSIXlt(hippodt)$min
hippomerge$Sec=as.POSIXlt(hippodt)$sec
# -
# interpolate prof from 10 sec file
hippomerge$prof=approx(as.POSIXct(hippo10sdt),hippomerge10s$prof,as.POSIXct(hippodt),method='constant',f=0)$y
# add strat flag
hippomerge$strat=rep(0,nrow(hippomerge)) # 0 means trop
h2oref=hippomerge$H2Oppmv_vxl; h2oref[is.na(h2oref)]=hippomerge$H2O_UWV[is.na(h2oref)]
hippomerge$h2oref=h2oref # for output
h2oref[is.na(h2oref)]=0 # if H2O missing treat as if potentially strat
n2oref=hippomerge$N2O_CCG; n2oref[is.na(n2oref)]=hippomerge$N2O_QCLS[is.na(n2oref)]
n2oref=n2oref-(approx(glbn2o$year+0.5,glbn2o$n2o,hippomerge$Year+hippomerge$DOY/365)$y-glbn2o$n2o[glbn2o$year==2009])
hippomerge$n2oref=n2oref # for output
n2oref[is.na(n2oref)]=400 # if N2O missing do not use for filter
o3ref=hippomerge$O3_ppb; o3ref[is.na(o3ref)]=hippomerge$O3_UO3[is.na(o3ref)]
hippomerge$o3ref=o3ref # for output
o3ref[is.na(o3ref)]=0 # if O3 missing do not use for filter
hippomerge$strat[h2oref<stratcoh2o&(o3ref>stratcoo3|n2oref<stratcon2o|(o3ref==0&n2oref==400))]=1 # if either o3 or n2o criteria are met, or if both are missing, consider strat
hippomerge$strat[h2oref==0&o3ref==0&n2oref==400&hippomerge$GGALT<8000]=0 # if all 3 missing assume < 8 km is trop
# select columns
colsel=c('Year','Month','Day','Hour','Min','Sec','H.no','flt','prof','GGLAT','GGLON','GGALT','PSXC','THETA','CO2_CCG','CO2_QCLS','CO2_OMS','CO2_AO2','CH4_CCG','CH4_QCLS','SF6_CCG','strat','h2oref','n2oref','o3ref')
hippomerge=hippomerge[,is.element(colnames(hippomerge),colsel)]
hippomerge=hippomerge[,match(colsel,names(hippomerge))] # reorder
names(hippomerge)=c('year','mon','day','hour','min','sec','camp','flt','prof','lat','lon','alt','pressure','theta','co2','co2qcls','co2oms','co2ao2','ch4pfp','ch4qcls','sf6pfp','strat','h2oref','n2oref','o3ref') ## 'co2' = CO2_CCG = co2pfp
# # read in ATom files, calc strat flag, and subset
# + tags=[]
# read and add time variables
atomvar=c('time','Flight_Date','DLH-H2O/H2O_DLH','UCATS-H2O/H2O_UWV','UCATS-O3/O3_UCATS','QCLS-CH4-CO-N2O/N2O_QCLS','NOyO3-O3/O3_CL','MMS/G_ALT','RF','prof.no','MMS/P','MMS/POT','MMS/G_LAT','MMS/G_LONG',
'MMS/G_ALT','NOAA-Picarro/CO2_NOAA','QCLS-CO2/CO2_QCLS','AO2/CO2_AO2','CO2.X','NOAA-Picarro/CH4_NOAA','QCLS-CH4-CO-N2O/CH4_QCLS','PFP/CO2_PFP','PFP/CH4_PFP','PFP/N2O_PFP','PFP/SF6_CCGG_PFP')
atommerge=NULL
for(i in c(1:3)){
atomnc=nc_open(paste(atommergedir,'/',atommergefiles[i],sep=''))
count=length(ncvar_get(atomnc,'time'))
campdata=NULL
for(var in atomvar){
campdata=cbind(campdata,ncvar_get(atomnc,var))
}
campdata=cbind(campdata,rep(i+1,count)) # A.no
nc_close(atomnc)
atommerge=rbind(atommerge,campdata)
}
atommerge=data.frame(atommerge,stringsAsFactors=F)
names(atommerge)=c(gsub('.*/','',atomvar),'A.no')
atommerge$YYYYMMDD=atommerge$Flight_Date
atomdt=as.POSIXlt(ISOdatetime(2016,1,1,0,0,0,tz='UTC')+atommerge$time,tz='UTC')
atommerge$Year=atomdt$year+1900
atommerge$Month=as.POSIXlt(atomdt)$mon+1
atommerge$Day=as.POSIXlt(atomdt)$mday
atommerge$Hour=as.POSIXlt(atomdt)$hour
atommerge$Min=as.POSIXlt(atomdt)$min
atommerge$Sec=as.POSIXlt(atomdt)$sec
# +
# add strat flag
atommerge$strat=rep(0,nrow(atommerge)) # 0 means trop
h2oref=atommerge$H2O_DLH; h2oref[is.na(h2oref)]=atommerge$H2O_UWV[is.na(h2oref)]
atommerge$h2oref=h2oref # for output
h2oref[is.na(h2oref)]=0 # if H2O missing treat as if potentially strat
n2oref=atommerge$N2O_QCLS; n2oref[is.na(n2oref)]=atommerge$N2O_PFP[is.na(n2oref)]
n2oref=n2oref-(approx(glbn2o$year+0.5,glbn2o$n2o,atommerge$Year+atomdt$yday/365)$y-glbn2o$n2o[glbn2o$year==2009])
atommerge$n2oref=n2oref # for outptut
n2oref[is.na(n2oref)]=400 # if N2O missing do not use for filter
o3ref=atommerge$O3_CL; o3ref[is.na(o3ref)]=atommerge$O3_UCATS[is.na(o3ref)]
atommerge$o3ref=o3ref
o3ref[is.na(o3ref)]=0 # if O3 missing do not use for filter
atommerge$strat[h2oref<stratcoh2o&(o3ref>stratcoo3|n2oref<stratcon2o|(o3ref==0&n2oref==400))]=1 # if either o3 or n2o criteria are met, or if both are missing, consider strat
atommerge$strat[h2oref==0&o3ref==0&n2oref==400&atommerge$G_ALT<8000]=0 # if all 3 missing assume < 8 km is trop
# -
# select column
colsel=c('Year','Month','Day','Hour','Min','Sec','A.no','RF','prof.no','G_LAT','G_LONG','G_ALT','P','POT','CO2_PFP','CO2_NOAA','CO2_QCLS','CO2_AO2','CH4_PFP','CH4_NOAA','CH4_QCLS','SF6_CCGG_PFP','strat','h2oref','n2oref','o3ref')
atommerge=atommerge[,is.element(colnames(atommerge),colsel)]
atommerge=atommerge[,match(colsel,names(atommerge))] # reorder
names(atommerge)=c('year','mon','day','hour','min','sec','camp','flt','prof','lat','lon','alt','pressure','theta','co2','co2noaa','co2qcls','co2ao2','ch4pfp','ch4noaa','ch4qcls','sf6pfp','strat','h2oref','n2oref','o3ref') ## 'co2' = CO2_PFP
# + [markdown] tags=[]
# # Filter airborne PFP data and subtract SPO
# -
# read in NOAA in situ record from SPO
sponc=nc_open(paste(project_tmpdir_obs,'/obspack_co2_1_GLOBALVIEWplus_v6.0_2020-09-11/data/nc/co2_spo_surface-insitu_1_allvalid.nc',sep=''))
spoco2=data.frame(cbind(ncvar_get(sponc,'time_decimal'),t(ncvar_get(sponc,'time_components')),ncvar_get(sponc,'value')*1E6)) ; colnames(spoco2)=c('date','year','mon','day','hour','min','sec','co2')
qcflag=ncvar_get(sponc,'qcflag'); spoco2$co2[substr(qcflag,1,1)!='.']=NA; spoco2$co2[substr(qcflag,2,2)!='.']=NA
spodt=ISOdatetime(spoco2$year,spoco2$mon,spoco2$day,spoco2$hour,spoco2$min,spoco2$sec,tz='UTC')
# # HIPPO
# filter
ints=read.table(paste(project_tmpdir_obs,'/hippo_xsect_filt_datetime.txt',sep=''),header=T)
startdt=ISOdatetime(ints$startyear,ints$startmon,ints$startday,ints$starthour,ints$startmin,ints$startsec,tz='UTC')
stopdt=ISOdatetime(ints$stopyear,ints$stopmon,ints$stopday,ints$stophour,ints$stopmin,ints$stopsec,tz='UTC')
blfilt=rep(T,nrow(hippomerge))
for(i in c(1:nrow(ints))){
blfilt[difftime(hippodt,startdt[i])>=0&difftime(hippodt,stopdt[i])<=0]=F
}
hippodt=hippodt[blfilt]
hippomerge=hippomerge[blfilt,]
print(paste('Filtered ',sum(!blfilt),' of ',length(blfilt),' HIPPO obs (',round(sum(!blfilt)/length(blfilt)*100,1),'%)',sep=''))
# calculate differences
hippomerge$co2mspo=round(hippomerge$co2-approx(as.POSIXct(spodt),spoco2$co2,as.POSIXct(hippodt))$y,3) ## co2 = 'CO2_CCG'
hippomerge$co2mqcls=round(hippomerge$co2-hippomerge$co2qcls,3)
hippomerge$co2moms=round(hippomerge$co2-hippomerge$co2oms,3)
hippomerge$co2mao2=round(hippomerge$co2-hippomerge$co2ao2,3)
hippomerge$ch4mqcls=round(hippomerge$ch4pfp-hippomerge$ch4qcls,3)
# +
# write out
write(names(hippomerge),'../data/aircraft-obs/HIPPO_SO_mSPO_pfp.txt',ncol=ncol(hippomerge))
write(t(hippomerge),'../data/aircraft-obs/HIPPO_SO_mSPO_pfp.txt',ncol=ncol(hippomerge),append=T)
print(apply(!is.na(hippomerge),2,sum))
# -
# # ATom
# filter
ints=read.table(paste(project_tmpdir_obs,'/atom_xsect_filt_datetime.txt',sep=''),header=T)
startdt=ISOdatetime(ints$startyear,ints$startmon,ints$startday,ints$starthour,ints$startmin,ints$startsec,tz='UTC')
stopdt=ISOdatetime(ints$stopyear,ints$stopmon,ints$stopday,ints$stophour,ints$stopmin,ints$stopsec,tz='UTC')
blfilt=rep(T,nrow(atommerge))
for(i in c(1:nrow(ints))){
blfilt[difftime(atomdt,startdt[i])>=0&difftime(atomdt,stopdt[i])<=0]=F
}
atomdt=atomdt[blfilt]
atommerge=atommerge[blfilt,]
print(paste('Filtered ',sum(!blfilt),' of ',length(blfilt),' ATom obs (',round(sum(!blfilt)/length(blfilt)*100,1),'%)',sep=''))
# calculate differences
atommerge$co2mspo=round(atommerge$co2-approx(as.POSIXct(spodt),spoco2$co2,as.POSIXct(atomdt))$y,2) ## co2 = 'CO2_PFP'
atommerge$co2mqcls=round(atommerge$co2-atommerge$co2qcls,3)
atommerge$co2mao2=round(atommerge$co2-atommerge$co2ao2,3)
atommerge$co2mnoaa=round(atommerge$co2-atommerge$co2noaa,3)
atommerge$ch4mqcls=round(atommerge$ch4pfp-atommerge$ch4qcls,3)
atommerge$ch4mnoaa=round(atommerge$ch4pfp-atommerge$ch4noaa,3)
# +
# write out
write(names(atommerge),'../data/aircraft-obs/ATOM_SO_mSPO_pfp.txt',ncol=ncol(atommerge))
write(t(atommerge),'../data/aircraft-obs/ATOM_SO_mSPO_pfp.txt',ncol=ncol(atommerge),append=T)
print(apply(!is.na(atommerge),2,sum))
|
so-co2-airborne-obs/_prestage-obs/process_aircraft_pfp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GooPyCharts Demo Notebook
# ## Import GooPyCharts
# To cut down on syntax, import figure directly.
from gpcharts import figure
# ## Simple Line Graph
# Same graph as described in the readme.
fig1 = figure()
fig1.plot([8,7,6,5,4])
# ## Line Graph with Two Lines
# Another line graph, but with two dependent variables. Also customizing plot.
fig2 = figure(title='Two lines',xlabel='Days',ylabel='Count',height=600,width=600)
xVals = ['Mon','Tues','Wed','Thurs','Fri']
yVals = [[5,4],[8,7],[4,8],[10,10],[3,12]]
fig2.plot(xVals,yVals)
# ## DateTime Graph
# A graph with dates and times. Title is assigned afterwards, and data is given header information.
fig3 = figure()
fig3.title = 'Weather over Days'
fig3.ylabel = 'Temperature'
#modify size of graph
fig3.height = 800
fig3.width = 1000
# X datetime data can take either of the following formats: "yyyy-mm-dd HH:MM:SS" or "yyyy-mm-dd", but be consistent.
#xVals = ['Dates','2016-03-20 00:00:00','2016-03-21 00:00:00','2016-03-25 00:00:00','2016-04-01 00:00:00']
xVals = ['Dates','2016-03-20','2016-03-21','2016-03-25','2016-04-01']
yVals = [['Shakuras','Korhal','Aiur'],[10,30,40],[12,28,41],[15,34,38],[8,33,47]]
fig3.plot(xVals,yVals)
# ## A Log Scale Example
# Set "logScale=True" when calling plot (or plot_nb for notebooks) to plot the y axis in log scale.
fig4 = figure(title='Population Growth',ylabel='Population')
xVals = ['Year',1700,1800,1900,2000]
yVals = [['Gotham City', 'Central City'],[0,10],[100,200],[100000,500000],[5000000,10000000]]
fig4.plot(xVals,yVals,logScale=True)
# ## Scatter Plot
# Scatter plot arguments are the same as for normal line graph arguments, but use "scatter" (or "scatter_nb" for notebooks) to plot instead. Scatter plots also support trend lines. Set "trendline=True" in the arguments to get a trendline on your graph. Currently only a trendline for the first dependent variable is supported.
fig5 = figure('Strong Correlation')
fig5.scatter([1,2,3,4,5],[[1,5],[2,4],[3,3],[4,2],[5,1]],trendline=True)
# ## Bar Graph
# Simple horizontal bar graphs are supported. Use function "bar" (or "bar_nb" for notebooks).
fig6 = figure('Percent Alcohol Consumption')
fig6.bar(['Percentage','Beer','Wine','Liquor'],['Type',40,50,10])
# ## Histogram
# Simple histograms are also supported. Histograms take in 1 list of input. Use function "hist" (or "hist_nb" for notebooks).
fig7 = figure('Distribution',xlabel='value')
fig7.hist([1,1,1,1,1,1,1,1,1,1,1,2,2,2,2,2,2,2,3,3,3,3,4,4,5,6,7,8,8,8,8,8,9,9,9,10,11,12,13,13,13,13,14])
|
MachineLearning/gpcharts test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Recap
# So far, you have loaded your data and reviewed it with the following code. Run this cell to set up your coding environment where the previous step left off.
# +
# Code you have previously used to load data
import pandas as pd
# Path of the file to read
iowa_file_path = '../input/home-data-for-ml-course/train.csv'
home_data = pd.read_csv(iowa_file_path)
home_data.describe()
# -
# # Exercises
#
# Run the following code cell to set up code checking.
# Set up code checking
from learntools.core import binder
binder.bind(globals())
from learntools.machine_learning.ex3 import *
# ## Step 1: Specify Target
# Select the target variable, which corresponds to the sales price. Save this to a new variable called `y`. You'll need to print a list of the columns to find the name of the column you need.
#
# print the list of columns in the dataset to find the name of the prediction target
print(home_data.columns)
y = home_data.SalePrice
print(y.mean())
step_1.check()
# The lines below will show you a hint or the solution.
# step_1.hint()
step_1.solution()
# ## Step 2: Create X
# Now you will create the DataFrame called `X` holding the predictive features.
#
# Since you want only some columns from the original data, you'll first create a list with the names of the columns you want in `X`.
#
# You'll use just the following columns in the list (you can copy and paste the whole list to save some typing, though you'll still need to add quotes):
# * LotArea
# * YearBuilt
# * 1stFlrSF
# * 2ndFlrSF
# * FullBath
# * BedroomAbvGr
# * TotRmsAbvGrd
#
# After you've created that list of featurse, use it to create the DataFrame that you'll use when fitting the model.
# +
# Create the list of features below
feature_names = ["LotArea", "YearBuilt", "1stFlrSF", "2ndFlrSF",
"FullBath", "BedroomAbvGr", "TotRmsAbvGrd"]
# select data corresponding to features in feature_names
X = home_data[feature_names]
step_2.check()
# -
step_2.solution()
# ## Review Data
# Before building a model, take a quick look at your features to see if they seem sensible
# +
# Review data
# print description or statistics from X
#print(_)
# print the top few lines
#print(_)
# -
# ## Step 3: Specify and Fit Model
# Create a `DecisionTreeRegressor` and save it iowa_model. Ensure you've done the relevant import from sklearn to run this command.
#
# Then fit the model you just created using the data in `X` and `y` that you saved above.
from sklearn.tree import DecisionTreeRegressor
iowa_model = DecisionTreeRegressor(random_state=1)
print("Incorrect")
step_3.check()
iowa_model.fit(X,y)
step_3.check()
step_3.hint()
step_3.solution()
# ## Step 4: Make Predictions
# Make predictions with the model's `predict` command. Save the predictions as first_preds.
predictions = iowa_model.predict(X)
step_4.check()
# step_4.hint()
step_4.solution()
# ## Think About Your Results
#
# Use the head command to compare the top few predictions to the home values (in y) for those same homes. Anything surprising?
#
# You'll understand why this happened if you keep going.
#
# ## Keep Going
# You've built a decision tree model. It's natural to ask how accurate the model's predictions will be and how you can improve that. Learn how to do that with **[Model Validation](https://www.kaggle.com/dansbecker/model-validation)**.
#
# ---
# **[Course Home Page](https://www.kaggle.com/learn/machine-learning)**
#
#
#
|
learntools/machine_learning/nbs/ch3-testing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# +
import sys
#sys.path.insert(0, '/Users/cwilson/desktop/programming/jupyter_tings/healthyride/healthyride')
sys.path.insert(0, 'C:\\Users\\chwilson\\OneDrive - Turner Construction\\Desktop\\HealthyRide')
import pandas as pd
from plotting_funcs import plot_all
from weekday_analytics.weekday import group_weekdaydict_by_station
from trip import Trip
import json
# -
with open('../data.json') as f:
json_list = json.load(f)
#convert list of dicts to list of objects called "Trips"
Trip_list = []
for dict in json_list:
li = list(dict.values())
Trip_list.append(Trip(*li))
#test er out
print(Trip_list[420].toname)
print(Trip_list[420].starttime)
print(Trip_list[420].weekday)
neighborhood_weekday_dict = group_weekdaydict_by_station(Trip_list)
plot_all(neighborhood_weekday_dict)
|
weekday_analytics/weekday_analytics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Employing Naive Bayes Classifiers to determine demographic labels
# In this notebook we will begin the process of classifying our households, such that we might populate recommendations for each one based on their real attributes.
# To do so, I'd like to train models using the labeled data -- the households for which we already have demographic information. This classifies as Supervised Learning, because we have accurate labels already.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import dtcj
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import GaussianNB, MultinomialNB, BernoulliNB
from sklearn.preprocessing import MinMaxScaler, StandardScaler
# +
# loading data
demo = dtcj.load_demo()
merged = dtcj.load_merged()
hh_agg = dtcj.load_hh_agg(merged)
# + code_folding=[]
def test_Multinomial_NB():
# for each of our targets
for target in ['single_couple_family']:
print(target)
# define feature space;
# remove data-leaked features
# negative values to positive
# # multi-collinearity?
# add target column as the last column.
test_hh_agg = abs(hh_agg.drop(['R', 'F', 'M', 'RFM', 'RFM Bins'], axis=1))
df = test_hh_agg.merge(demo[['household_key', target]], on='household_key')
# set feature space and target variable
X = df.iloc[:, :-1].values
y = df.iloc[:, -1].values
# train test split the data -- employing stratify.
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y)
# hyperparameter optimization
for alpha in [0.005, 0.05, 0.5, 1, 10, 100, 1000, 100000, 1000000000]:
# Instatiate our model
mnbmodel = MultinomialNB(alpha=alpha)
# Fit our model
mnbmodel.fit(X_train, y_train)
# simple print for results
print('alpha:', alpha, 'score', mnbmodel.score(X_test, y_test))
test_Multinomial_NB() # this is calculating the target single_couple_family
# + code_folding=[]
def test_Gaussian_NB():
for target in ['income_50K+', 'age_45+']:
# define feature space; remove data-leaked features and multi-collinearity?
test_hh_agg = abs(hh_agg.drop(['R', 'F', 'M', 'RFM', 'RFM Bins'], axis=1))
df = test_hh_agg.merge(demo[['household_key', target]], on='household_key')
# set features and target
X = df.iloc[:, :-1].values
y = df.iloc[:, -1].values
# train test split
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y)
# scale the data
ss = StandardScaler().fit(X_train)
X_train = ss.transform(X_train)
X_test = ss.transform(X_test)
# hyperparameter optimization
# Instatiate our model
gnbmodel = GaussianNB()
# Fit our model
gnbmodel.fit(X_train, y_train)
# simple print for results
print(target, gnbmodel.score(X_test, y_test))
test_Gaussian_NB() # this is calculating the target income_50K+
# -
# + code_folding=[0]
def test_Bernoulli_NB():
for target in ['single', 'couple', 'has_kids']:
print(target)
# define feature space; remove data-leaked features and multi-collinearity?
test_hh_agg = abs(hh_agg.drop(['R', 'F', 'M', 'RFM', 'RFM Bins'], axis=1))
df = test_hh_agg.merge(demo[['household_key', target]], on='household_key')
# set features and target
X = df.iloc[:, :-1].values
y = df.iloc[:, -1].values
# train test split
X_train, X_test, y_train, y_test = train_test_split(X, y)
# scale the data
ss = StandardScaler().fit(X_train)
X_train = ss.transform(X_train)
X_test = ss.transform(X_test)
# hyperparameter optimization
for alpha in [0.005, 0.05, 0.5, 1, 10, 100, 1000, 100000, 1000000000]:
# Instatiate our model
bnbmodel = BernoulliNB(alpha=alpha)
# Fit our model
bnbmodel.fit(X_train, y_train)
# simple print for results
print('alpha:', alpha, 'score', bnbmodel.score(X_test, y_test))
test_Bernoulli_NB()
# -
|
2.3 -- Testing Naive Bayes for demographic groups.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/priyanshgupta1998/All_codes/blob/master/prac1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="lKHTpj0672rV" colab_type="code" colab={}
import pandas as pd
import random
# + id="eIdVP-vl72vC" colab_type="code" colab={}
data = pd.read_csv('/home/test.csv')
# + id="j21fFe6V72yO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5f5c109d-6f07-40e6-8618-67460a38c22a"
print(len(data))
# + id="sK9urPqV9F2J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="01406728-d405-4d40-d6ad-46903a62f3cf"
print(data.shape)
# + id="Nzn9e9Nc9F-Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7853b9bc-c142-47d6-fc24-67d9b53d9e30"
print(data.shape[0])
# + id="418Y8xe09T10" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6273e9e4-42e6-4208-affd-fbbdf6825341"
k = random.randint(0,1000)
k
# + id="2t-A6FSY9T5L" colab_type="code" colab={}
data['id'] = [random.randint(0,1000) for x in range(data.shape[0])]
# + id="NxIyW-lS9T-k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="94dfc4ba-8cf5-487e-a32d-9c30fe451a95"
print(data.shape) # one column is added in the dataframe
# + id="IQaqlx1E9T9A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 346} outputId="781f4720-8aec-4b64-b7dc-c835d3a03f90"
data.head(8)
# + id="zWVs6aL29F8x" colab_type="code" colab={}
# + id="lcS2VjFK9F68" colab_type="code" colab={}
# + id="_YMkxpB29F5b" colab_type="code" colab={}
|
prac1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.11 64-bit (''tf24'': venv)'
# name: python3
# ---
# ## Demo 3: HKR classifier on MNIST dataset
# [](https://colab.research.google.com/github/deel-ai/deel-lip/blob/master/doc/notebooks/demo3.ipynb)
#
# This notebook will demonstrate learning a binary task on the MNIST0-8 dataset.
# +
# pip install deel-lip -qqq
# +
import tensorflow as tf
from tensorflow.keras import backend as K
from tensorflow.python.keras.layers import Input, Flatten
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.metrics import binary_accuracy
from tensorflow.keras.models import Sequential
from deel.lip.layers import (
SpectralConv2D,
SpectralDense,
FrobeniusDense,
ScaledL2NormPooling2D,
)
from deel.lip.activations import MaxMin, GroupSort, GroupSort2, FullSort
from deel.lip.losses import HKR, KR, HingeMargin
# -
# ### data preparation
#
# For this task we will select two classes: 0 and 8. Labels are changed to {-1,1}, wich is compatible
# with the Hinge term used in the loss.
# +
from tensorflow.keras.datasets import mnist
# first we select the two classes
selected_classes = [0, 8] # must be two classes as we perform binary classification
def prepare_data(x, y, class_a=0, class_b=8):
"""
This function convert the MNIST data to make it suitable for our binary classification
setup.
"""
# select items from the two selected classes
mask = (y == class_a) + (
y == class_b
) # mask to select only items from class_a or class_b
x = x[mask]
y = y[mask]
x = x.astype("float32")
y = y.astype("float32")
# convert from range int[0,255] to float32[-1,1]
x /= 255
x = x.reshape((-1, 28, 28, 1))
# change label to binary classification {-1,1}
y[y == class_a] = 1.0
y[y == class_b] = -1.0
return x, y
# now we load the dataset
(x_train, y_train_ord), (x_test, y_test_ord) = mnist.load_data()
# prepare the data
x_train, y_train = prepare_data(
x_train, y_train_ord, selected_classes[0], selected_classes[1]
)
x_test, y_test = prepare_data(
x_test, y_test_ord, selected_classes[0], selected_classes[1]
)
# display infos about dataset
print(
"train set size: %i samples, classes proportions: %.3f percent"
% (y_train.shape[0], 100 * y_train[y_train == 1].sum() / y_train.shape[0])
)
print(
"test set size: %i samples, classes proportions: %.3f percent"
% (y_test.shape[0], 100 * y_test[y_test == 1].sum() / y_test.shape[0])
)
# -
# ### Build lipschitz Model
#
# Let's first explicit the paremeters of this experiment
# +
# training parameters
epochs = 10
batch_size = 128
# network parameters
activation = GroupSort # ReLU, MaxMin, GroupSort2
# loss parameters
min_margin = 1.0
alpha = 10.0
# -
# Now we can build the network.
# Here the experiment is done with a MLP. But `Deel-lip` also provide state of the art 1-Lipschitz convolutions.
K.clear_session()
# helper function to build the 1-lipschitz MLP
wass = Sequential(
layers=[
Input((28, 28, 1)),
Flatten(),
SpectralDense(32, GroupSort2(), use_bias=True),
SpectralDense(16, GroupSort2(), use_bias=True),
FrobeniusDense(1, activation=None, use_bias=False),
],
name="lipModel",
)
wass.summary()
optimizer = Adam(lr=0.001)
# as the output of our classifier is in the real range [-1, 1], binary accuracy must be redefined
def HKR_binary_accuracy(y_true, y_pred):
S_true = tf.dtypes.cast(tf.greater_equal(y_true[:, 0], 0), dtype=tf.float32)
S_pred = tf.dtypes.cast(tf.greater_equal(y_pred[:, 0], 0), dtype=tf.float32)
return binary_accuracy(S_true, S_pred)
wass.compile(
loss=HKR(
alpha=alpha, min_margin=min_margin
), # HKR stands for the hinge regularized KR loss
metrics=[
KR, # shows the KR term of the loss
HingeMargin(min_margin=min_margin), # shows the hinge term of the loss
HKR_binary_accuracy, # shows the classification accuracy
],
optimizer=optimizer,
)
# ### Learn classification on MNIST
#
# Now the model is build, we can learn the task.
wass.fit(
x=x_train,
y=y_train,
validation_data=(x_test, y_test),
batch_size=batch_size,
shuffle=True,
epochs=epochs,
verbose=1,
)
# As we can see the model reach a very decent accuracy on this task.
|
doc/notebooks/demo3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="zeHYl-s4UUnj" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623345417804, "user_tz": -120, "elapsed": 21374, "user": {"displayName": "\<PASSWORD>", "photoUrl": "", "userId": "08102834906078443726"}} outputId="e4269e72-60c2-447e-885c-ce7ca67a5081"
# This mounts your Google Drive to the Colab VM.
from google.colab import drive
drive.mount('/content/drive')
# TODO: Enter the foldername in your Drive where you have saved the unzipped
# assignment folder, e.g. 'cs231n/assignments/assignment1/'
FOLDERNAME = 'CS231N/assignment/assignment2/'
assert FOLDERNAME is not None, "[!] Enter the foldername."
# Now that we've mounted your Drive, this ensures that
# the Python interpreter of the Colab VM can load
# python files from within it.
import sys
sys.path.append('/content/drive/My Drive/{}'.format(FOLDERNAME))
# This downloads the CIFAR-10 dataset to your Drive
# if it doesn't already exist.
# %cd /content/drive/My\ Drive/$FOLDERNAME/cs231n/datasets/
# !bash get_datasets.sh
# %cd /content/drive/My\ Drive/$FOLDERNAME
# + [markdown] tags=["pdf-title"] id="gTCCVfF7UUnj"
# # Introduction to TensorFlow
#
# You've written a lot of code in this assignment to provide a whole host of neural network functionality. Dropout, Batch Norm, and 2D convolutions are some of the workhorses of deep learning in computer vision. You've also worked hard to make your code efficient and vectorized.
#
# For the last part of this assignment, though, we're going to leave behind your beautiful codebase and instead migrate to one of two popular deep learning frameworks: in this instance, TensorFlow (or PyTorch, if you choose to work with that notebook).
# + [markdown] tags=["pdf-ignore"] id="XpbBoo7cUUnk"
# ## Why do we use deep learning frameworks?
#
# * Our code will now run on GPUs! This will allow our models to train much faster. When using a framework like PyTorch or TensorFlow you can harness the power of the GPU for your own custom neural network architectures without having to write CUDA code directly (which is beyond the scope of this class).
# * In this class, we want you to be ready to use one of these frameworks for your project so you can experiment more efficiently than if you were writing every feature you want to use by hand.
# * We want you to stand on the shoulders of giants! TensorFlow and PyTorch are both excellent frameworks that will make your lives a lot easier, and now that you understand their guts, you are free to use them :)
# * Finally, we want you to be exposed to the sort of deep learning code you might run into in academia or industry.
#
# ## What is TensorFlow?
# TensorFlow is a system for executing computational graphs over Tensor objects, with native support for performing backpropogation for its Variables. In it, we work with Tensors which are n-dimensional arrays analogous to the numpy ndarray.
#
# ## How do I learn TensorFlow?
#
# TensorFlow has many excellent tutorials available, including those from [Google themselves](https://www.tensorflow.org/get_started/get_started).
#
# Otherwise, this notebook will walk you through much of what you need to do to train models in TensorFlow. See the end of the notebook for some links to helpful tutorials if you want to learn more or need further clarification on topics that aren't fully explained here.
#
# **Note:** This notebook is meant to teach you Tensorflow 2.x. Most examples on the web today are still in 1.x, so be careful not to confuse the two when looking up documentation.
# + [markdown] tags=["pdf-ignore"] id="Lvzh6zDzUUnk"
# # Table of Contents
#
# This notebook has 5 parts. We will walk through TensorFlow at **three different levels of abstraction**, which should help you better understand it and prepare you for working on your project.
#
# 1. Part I, Preparation: load the CIFAR-10 dataset.
# 2. Part II, Barebone TensorFlow: **Abstraction Level 1**, we will work directly with low-level TensorFlow graphs.
# 3. Part III, Keras Model API: **Abstraction Level 2**, we will use `tf.keras.Model` to define arbitrary neural network architecture.
# 4. Part IV, Keras Sequential + Functional API: **Abstraction Level 3**, we will use `tf.keras.Sequential` to define a linear feed-forward network very conveniently, and then explore the functional libraries for building unique and uncommon models that require more flexibility.
# 5. Part V, CIFAR-10 open-ended challenge: please implement your own network to get as high accuracy as possible on CIFAR-10. You can experiment with any layer, optimizer, hyperparameters or other advanced features.
#
# We will discuss Keras in more detail later in the notebook.
#
# Here is a table of comparison:
#
# | API | Flexibility | Convenience |
# |---------------|-------------|-------------|
# | Barebone | High | Low |
# | `tf.keras.Model` | High | Medium |
# | `tf.keras.Sequential` | Low | High |
# + [markdown] id="KFpz2-jUUUnl"
# # GPU
#
# You can manually switch to a GPU device on Colab by clicking `Runtime -> Change runtime type` and selecting `GPU` under `Hardware Accelerator`. You should do this before running the following cells to import packages, since the kernel gets restarted upon switching runtimes.
# + tags=["pdf-ignore"] id="Saq6wAn3UUnl" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623346102726, "user_tz": -120, "elapsed": 1806, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="ff3593d6-6894-4332-b226-cf4be5cfd671"
import os
import tensorflow as tf
import numpy as np
import math
import timeit
import matplotlib.pyplot as plt
# %matplotlib inline
USE_GPU = True
if USE_GPU:
device = '/device:GPU:0'
else:
device = '/cpu:0'
# Constant to control how often we print when training models.
print_every = 100
print('Using device: ', device)
# + [markdown] id="pJSjX9tVUUnl"
# # Part I: Preparation
#
# First, we load the CIFAR-10 dataset. This might take a few minutes to download the first time you run it, but after that the files should be cached on disk and loading should be faster.
#
# In previous parts of the assignment we used CS231N-specific code to download and read the CIFAR-10 dataset; however the `tf.keras.datasets` package in TensorFlow provides prebuilt utility functions for loading many common datasets.
#
# For the purposes of this assignment we will still write our own code to preprocess the data and iterate through it in minibatches. The `tf.data` package in TensorFlow provides tools for automating this process, but working with this package adds extra complication and is beyond the scope of this notebook. However using `tf.data` can be much more efficient than the simple approach used in this notebook, so you should consider using it for your project.
# + tags=["pdf-ignore"] id="lUZVbgIuUUnm" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623346135083, "user_tz": -120, "elapsed": 8115, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="571cd495-ea3d-461e-f297-d90bbbea8d10"
def load_cifar10(num_training=49000, num_validation=1000, num_test=10000):
"""
Fetch the CIFAR-10 dataset from the web and perform preprocessing to prepare
it for the two-layer neural net classifier. These are the same steps as
we used for the SVM, but condensed to a single function.
"""
# Load the raw CIFAR-10 dataset and use appropriate data types and shapes
cifar10 = tf.keras.datasets.cifar10.load_data()
(X_train, y_train), (X_test, y_test) = cifar10
X_train = np.asarray(X_train, dtype=np.float32)
y_train = np.asarray(y_train, dtype=np.int32).flatten()
X_test = np.asarray(X_test, dtype=np.float32)
y_test = np.asarray(y_test, dtype=np.int32).flatten()
# Subsample the data
mask = range(num_training, num_training + num_validation)
X_val = X_train[mask]
y_val = y_train[mask]
mask = range(num_training)
X_train = X_train[mask]
y_train = y_train[mask]
mask = range(num_test)
X_test = X_test[mask]
y_test = y_test[mask]
# Normalize the data: subtract the mean pixel and divide by std
mean_pixel = X_train.mean(axis=(0, 1, 2), keepdims=True)
std_pixel = X_train.std(axis=(0, 1, 2), keepdims=True)
X_train = (X_train - mean_pixel) / std_pixel
X_val = (X_val - mean_pixel) / std_pixel
X_test = (X_test - mean_pixel) / std_pixel
return X_train, y_train, X_val, y_val, X_test, y_test
# If there are errors with SSL downloading involving self-signed certificates,
# it may be that your Python version was recently installed on the current machine.
# See: https://github.com/tensorflow/tensorflow/issues/10779
# To fix, run the command: /Applications/Python\ 3.7/Install\ Certificates.command
# ...replacing paths as necessary.
# Invoke the above function to get our data.
NHW = (0, 1, 2)
X_train, y_train, X_val, y_val, X_test, y_test = load_cifar10()
print('Train data shape: ', X_train.shape)
print('Train labels shape: ', y_train.shape, y_train.dtype)
print('Validation data shape: ', X_val.shape)
print('Validation labels shape: ', y_val.shape)
print('Test data shape: ', X_test.shape)
print('Test labels shape: ', y_test.shape)
# + tags=["pdf-ignore"] id="swGYkqSVUUnm" executionInfo={"status": "ok", "timestamp": 1623346235155, "user_tz": -120, "elapsed": 325, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}}
class Dataset(object):
def __init__(self, X, y, batch_size, shuffle=False):
"""
Construct a Dataset object to iterate over data X and labels y
Inputs:
- X: Numpy array of data, of any shape
- y: Numpy array of labels, of any shape but with y.shape[0] == X.shape[0]
- batch_size: Integer giving number of elements per minibatch
- shuffle: (optional) Boolean, whether to shuffle the data on each epoch
"""
assert X.shape[0] == y.shape[0], 'Got different numbers of data and labels'
self.X, self.y = X, y
self.batch_size, self.shuffle = batch_size, shuffle
def __iter__(self):
N, B = self.X.shape[0], self.batch_size
idxs = np.arange(N)
if self.shuffle:
np.random.shuffle(idxs)
return iter((self.X[i:i+B], self.y[i:i+B]) for i in range(0, N, B))
train_dset = Dataset(X_train, y_train, batch_size=64, shuffle=True)
val_dset = Dataset(X_val, y_val, batch_size=64, shuffle=False)
test_dset = Dataset(X_test, y_test, batch_size=64)
# + id="Ca2zoDVTUUnm" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623346238275, "user_tz": -120, "elapsed": 326, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="7ae72242-718f-4297-ca03-51f539bb6d2f"
# We can iterate through a dataset like this:
for t, (x, y) in enumerate(train_dset):
print(t, x.shape, y.shape)
if t > 5: break
# + [markdown] tags=["pdf-ignore"] id="JRAQ4GRXUUnn"
# # Part II: Barebones TensorFlow
# TensorFlow ships with various high-level APIs which make it very convenient to define and train neural networks; we will cover some of these constructs in Part III and Part IV of this notebook. In this section we will start by building a model with basic TensorFlow constructs to help you better understand what's going on under the hood of the higher-level APIs.
#
# **"Barebones Tensorflow" is important to understanding the building blocks of TensorFlow, but much of it involves concepts from TensorFlow 1.x.** We will be working with legacy modules such as `tf.Variable`.
#
# Therefore, please read and understand the differences between legacy (1.x) TF and the new (2.0) TF.
#
# ### Historical background on TensorFlow 1.x
#
# TensorFlow 1.x is primarily a framework for working with **static computational graphs**. Nodes in the computational graph are Tensors which will hold n-dimensional arrays when the graph is run; edges in the graph represent functions that will operate on Tensors when the graph is run to actually perform useful computation.
#
# Before Tensorflow 2.0, we had to configure the graph into two phases. There are plenty of tutorials online that explain this two-step process. The process generally looks like the following for TF 1.x:
# 1. **Build a computational graph that describes the computation that you want to perform**. This stage doesn't actually perform any computation; it just builds up a symbolic representation of your computation. This stage will typically define one or more `placeholder` objects that represent inputs to the computational graph.
# 2. **Run the computational graph many times.** Each time the graph is run (e.g. for one gradient descent step) you will specify which parts of the graph you want to compute, and pass a `feed_dict` dictionary that will give concrete values to any `placeholder`s in the graph.
#
# ### The new paradigm in Tensorflow 2.0
# Now, with Tensorflow 2.0, we can simply adopt a functional form that is more Pythonic and similar in spirit to PyTorch and direct Numpy operation. Instead of the 2-step paradigm with computation graphs, making it (among other things) easier to debug TF code. You can read more details at https://www.tensorflow.org/guide/eager.
#
# The main difference between the TF 1.x and 2.0 approach is that the 2.0 approach doesn't make use of `tf.Session`, `tf.run`, `placeholder`, `feed_dict`. To get more details of what's different between the two version and how to convert between the two, check out the official migration guide: https://www.tensorflow.org/alpha/guide/migration_guide
#
# Later, in the rest of this notebook we'll focus on this new, simpler approach.
# + [markdown] tags=["pdf-ignore"] id="eILGb9i0UUnn"
# ### TensorFlow warmup: Flatten Function
#
# We can see this in action by defining a simple `flatten` function that will reshape image data for use in a fully-connected network.
#
# In TensorFlow, data for convolutional feature maps is typically stored in a Tensor of shape N x H x W x C where:
#
# - N is the number of datapoints (minibatch size)
# - H is the height of the feature map
# - W is the width of the feature map
# - C is the number of channels in the feature map
#
# This is the right way to represent the data when we are doing something like a 2D convolution, that needs spatial understanding of where the intermediate features are relative to each other. When we use fully connected affine layers to process the image, however, we want each datapoint to be represented by a single vector -- it's no longer useful to segregate the different channels, rows, and columns of the data. So, we use a "flatten" operation to collapse the `H x W x C` values per representation into a single long vector.
#
# Notice the `tf.reshape` call has the target shape as `(N, -1)`, meaning it will reshape/keep the first dimension to be N, and then infer as necessary what the second dimension is in the output, so we can collapse the remaining dimensions from the input properly.
#
# **NOTE**: TensorFlow and PyTorch differ on the default Tensor layout; TensorFlow uses N x H x W x C but PyTorch uses N x C x H x W.
# + tags=["pdf-ignore"] id="JFDyxw9ZUUnn" executionInfo={"status": "ok", "timestamp": 1623347087280, "user_tz": -120, "elapsed": 211, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}}
def flatten(x):
"""
Input:
- TensorFlow Tensor of shape (N, D1, ..., DM)
Output:
- TensorFlow Tensor of shape (N, D1 * ... * DM)
"""
N = tf.shape(x)[0]
return tf.reshape(x, (N, -1))
# + tags=["pdf-ignore-input"] id="bqlBpm9SUUnn" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623347122314, "user_tz": -120, "elapsed": 5461, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="41a767e7-5069-46de-f7f2-8f0286e0d459"
def test_flatten():
# Construct concrete values of the input data x using numpy
x_np = np.arange(24).reshape((2, 3, 4))
print('x_np:\n', x_np, '\n')
# Compute a concrete output value.
x_flat_np = flatten(x_np)
print('x_flat_np:\n', x_flat_np, '\n')
test_flatten()
# + [markdown] id="NficlBG3UUno"
# ### Barebones TensorFlow: Define a Two-Layer Network
# We will now implement our first neural network with TensorFlow: a fully-connected ReLU network with two hidden layers and no biases on the CIFAR10 dataset. For now we will use only low-level TensorFlow operators to define the network; later we will see how to use the higher-level abstractions provided by `tf.keras` to simplify the process.
#
# We will define the forward pass of the network in the function `two_layer_fc`; this will accept TensorFlow Tensors for the inputs and weights of the network, and return a TensorFlow Tensor for the scores.
#
# After defining the network architecture in the `two_layer_fc` function, we will test the implementation by checking the shape of the output.
#
# **It's important that you read and understand this implementation.**
# + tags=["pdf-ignore"] id="_mkAe9J7UUno" executionInfo={"status": "ok", "timestamp": 1623347210638, "user_tz": -120, "elapsed": 314, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}}
def two_layer_fc(x, params):
"""
A fully-connected neural network; the architecture is:
fully-connected layer -> ReLU -> fully connected layer.
Note that we only need to define the forward pass here; TensorFlow will take
care of computing the gradients for us.
The input to the network will be a minibatch of data, of shape
(N, d1, ..., dM) where d1 * ... * dM = D. The hidden layer will have H units,
and the output layer will produce scores for C classes.
Inputs:
- x: A TensorFlow Tensor of shape (N, d1, ..., dM) giving a minibatch of
input data.
- params: A list [w1, w2] of TensorFlow Tensors giving weights for the
network, where w1 has shape (D, H) and w2 has shape (H, C).
Returns:
- scores: A TensorFlow Tensor of shape (N, C) giving classification scores
for the input data x.
"""
w1, w2 = params # Unpack the parameters
x = flatten(x) # Flatten the input; now x has shape (N, D)
h = tf.nn.relu(tf.matmul(x, w1)) # Hidden layer: h has shape (N, H)
scores = tf.matmul(h, w2) # Compute scores of shape (N, C)
return scores
# + tags=["pdf-ignore-input"] id="55XBueB9UUno" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623347244965, "user_tz": -120, "elapsed": 2822, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="5f060dce-0680-4570-9e10-39c8caa0685b"
def two_layer_fc_test():
hidden_layer_size = 42
# Scoping our TF operations under a tf.device context manager
# lets us tell TensorFlow where we want these Tensors to be
# multiplied and/or operated on, e.g. on a CPU or a GPU.
with tf.device(device):
x = tf.zeros((64, 32, 32, 3))
w1 = tf.zeros((32 * 32 * 3, hidden_layer_size))
w2 = tf.zeros((hidden_layer_size, 10))
# Call our two_layer_fc function for the forward pass of the network.
scores = two_layer_fc(x, [w1, w2])
print(scores.shape)
two_layer_fc_test()
# + [markdown] id="4cx52pRuUUno"
# ### Barebones TensorFlow: Three-Layer ConvNet
# Here you will complete the implementation of the function `three_layer_convnet` which will perform the forward pass of a three-layer convolutional network. The network should have the following architecture:
#
# 1. A convolutional layer (with bias) with `channel_1` filters, each with shape `KW1 x KH1`, and zero-padding of two
# 2. ReLU nonlinearity
# 3. A convolutional layer (with bias) with `channel_2` filters, each with shape `KW2 x KH2`, and zero-padding of one
# 4. ReLU nonlinearity
# 5. Fully-connected layer with bias, producing scores for `C` classes.
#
# **HINT**: For convolutions: https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/nn/conv2d; be careful with padding!
#
# **HINT**: For biases: https://www.tensorflow.org/performance/xla/broadcasting
# + id="gdo_RFKlUUnp" executionInfo={"status": "ok", "timestamp": 1623347816374, "user_tz": -120, "elapsed": 331, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}}
def three_layer_convnet(x, params):
"""
A three-layer convolutional network with the architecture described above.
Inputs:
- x: A TensorFlow Tensor of shape (N, H, W, 3) giving a minibatch of images
- params: A list of TensorFlow Tensors giving the weights and biases for the
network; should contain the following:
- conv_w1: TensorFlow Tensor of shape (KH1, KW1, 3, channel_1) giving
weights for the first convolutional layer.
- conv_b1: TensorFlow Tensor of shape (channel_1,) giving biases for the
first convolutional layer.
- conv_w2: TensorFlow Tensor of shape (KH2, KW2, channel_1, channel_2)
giving weights for the second convolutional layer
- conv_b2: TensorFlow Tensor of shape (channel_2,) giving biases for the
second convolutional layer.
- fc_w: TensorFlow Tensor giving weights for the fully-connected layer.
Can you figure out what the shape should be?
- fc_b: TensorFlow Tensor giving biases for the fully-connected layer.
Can you figure out what the shape should be?
"""
conv_w1, conv_b1, conv_w2, conv_b2, fc_w, fc_b = params
scores = None
############################################################################
# TODO: Implement the forward pass for the three-layer ConvNet. #
############################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
N, H, W, C = x.shape
out = tf.nn.relu(tf.nn.conv2d(x,conv_w1,1,[[0, 0], [2,2], [2, 2], [0, 0]]) + conv_b1)
out = tf.nn.relu(tf.nn.conv2d(out,conv_w2,1,[[0, 0], [1,1], [1, 1], [0, 0]]) + conv_b2)
out = flatten(out)
scores = tf.matmul(out, fc_w) + fc_b
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
############################################################################
# END OF YOUR CODE #
############################################################################
return scores
# + [markdown] id="R_MKMwW0UUnp"
# After defing the forward pass of the three-layer ConvNet above, run the following cell to test your implementation. Like the two-layer network, we run the graph on a batch of zeros just to make sure the function doesn't crash, and produces outputs of the correct shape.
#
# When you run this function, `scores_np` should have shape `(64, 10)`.
# + id="barebones_output_shape" tags=["pdf-ignore-input"] colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623347818444, "user_tz": -120, "elapsed": 323, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="16480340-3732-446a-9dc6-85ba3e5ba4a3"
def three_layer_convnet_test():
with tf.device(device):
x = tf.zeros((64, 32, 32, 3))
conv_w1 = tf.zeros((5, 5, 3, 6))
conv_b1 = tf.zeros((6,))
conv_w2 = tf.zeros((3, 3, 6, 9))
conv_b2 = tf.zeros((9,))
fc_w = tf.zeros((32 * 32 * 9, 10))
fc_b = tf.zeros((10,))
params = [conv_w1, conv_b1, conv_w2, conv_b2, fc_w, fc_b]
scores = three_layer_convnet(x, params)
# Inputs to convolutional layers are 4-dimensional arrays with shape
# [batch_size, height, width, channels]
print('scores_np has shape: ', scores.shape)
three_layer_convnet_test()
# + [markdown] id="PF6-cHbHUUnp"
# ### Barebones TensorFlow: Training Step
#
# We now define the `training_step` function performs a single training step. This will take three basic steps:
#
# 1. Compute the loss
# 2. Compute the gradient of the loss with respect to all network weights
# 3. Make a weight update step using (stochastic) gradient descent.
#
#
# We need to use a few new TensorFlow functions to do all of this:
# - For computing the cross-entropy loss we'll use `tf.nn.sparse_softmax_cross_entropy_with_logits`: https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/nn/sparse_softmax_cross_entropy_with_logits
#
# - For averaging the loss across a minibatch of data we'll use `tf.reduce_mean`:
# https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/reduce_mean
#
# - For computing gradients of the loss with respect to the weights we'll use `tf.GradientTape` (useful for Eager execution): https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/GradientTape
#
# - We'll mutate the weight values stored in a TensorFlow Tensor using `tf.assign_sub` ("sub" is for subtraction): https://www.tensorflow.org/api_docs/python/tf/assign_sub
#
# + tags=["pdf-ignore"] id="JUpqOyoAUUnp" executionInfo={"status": "ok", "timestamp": 1623348142874, "user_tz": -120, "elapsed": 360, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}}
def training_step(model_fn, x, y, params, learning_rate):
with tf.GradientTape() as tape:
scores = model_fn(x, params) # Forward pass of the model
loss = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=scores)
total_loss = tf.reduce_mean(loss)
grad_params = tape.gradient(total_loss, params)
# Make a vanilla gradient descent step on all of the model parameters
# Manually update the weights using assign_sub()
for w, grad_w in zip(params, grad_params):
w.assign_sub(learning_rate * grad_w)
return total_loss
# + tags=["pdf-ignore"] id="aN68ChW9UUnp" executionInfo={"status": "ok", "timestamp": 1623348193984, "user_tz": -120, "elapsed": 201, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}}
def train_part2(model_fn, init_fn, learning_rate):
"""
Train a model on CIFAR-10.
Inputs:
- model_fn: A Python function that performs the forward pass of the model
using TensorFlow; it should have the following signature:
scores = model_fn(x, params) where x is a TensorFlow Tensor giving a
minibatch of image data, params is a list of TensorFlow Tensors holding
the model weights, and scores is a TensorFlow Tensor of shape (N, C)
giving scores for all elements of x.
- init_fn: A Python function that initializes the parameters of the model.
It should have the signature params = init_fn() where params is a list
of TensorFlow Tensors holding the (randomly initialized) weights of the
model.
- learning_rate: Python float giving the learning rate to use for SGD.
"""
params = init_fn() # Initialize the model parameters
for t, (x_np, y_np) in enumerate(train_dset):
# Run the graph on a batch of training data.
loss = training_step(model_fn, x_np, y_np, params, learning_rate)
# Periodically print the loss and check accuracy on the val set.
if t % print_every == 0:
print('Iteration %d, loss = %.4f' % (t, loss))
check_accuracy(val_dset, x_np, model_fn, params)
# + tags=["pdf-ignore"] id="rY-yM8jOUUnq" executionInfo={"status": "ok", "timestamp": 1623348219264, "user_tz": -120, "elapsed": 318, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}}
def check_accuracy(dset, x, model_fn, params):
"""
Check accuracy on a classification model, e.g. for validation.
Inputs:
- dset: A Dataset object against which to check accuracy
- x: A TensorFlow placeholder Tensor where input images should be fed
- model_fn: the Model we will be calling to make predictions on x
- params: parameters for the model_fn to work with
Returns: Nothing, but prints the accuracy of the model
"""
num_correct, num_samples = 0, 0
for x_batch, y_batch in dset:
scores_np = model_fn(x_batch, params).numpy()
y_pred = scores_np.argmax(axis=1)
num_samples += x_batch.shape[0]
num_correct += (y_pred == y_batch).sum()
acc = float(num_correct) / num_samples
print('Got %d / %d correct (%.2f%%)' % (num_correct, num_samples, 100 * acc))
# + [markdown] id="AG7HV7XZUUnq"
# ### Barebones TensorFlow: Initialization
# We'll use the following utility method to initialize the weight matrices for our models using Kaiming's normalization method.
#
# [1] He et al, *Delving Deep into Rectifiers: Surpassing Human-Level Performance on ImageNet Classification
# *, ICCV 2015, https://arxiv.org/abs/1502.01852
# + id="tHzQzux1UUnq" executionInfo={"status": "ok", "timestamp": 1623348311672, "user_tz": -120, "elapsed": 363, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}}
def create_matrix_with_kaiming_normal(shape):
if len(shape) == 2:
fan_in, fan_out = shape[0], shape[1]
elif len(shape) == 4:
fan_in, fan_out = np.prod(shape[:3]), shape[3]
return tf.keras.backend.random_normal(shape) * np.sqrt(2.0 / fan_in)
# + [markdown] id="35zbRAsPUUnq"
# ### Barebones TensorFlow: Train a Two-Layer Network
# We are finally ready to use all of the pieces defined above to train a two-layer fully-connected network on CIFAR-10.
#
# We just need to define a function to initialize the weights of the model, and call `train_part2`.
#
# Defining the weights of the network introduces another important piece of TensorFlow API: `tf.Variable`. A TensorFlow Variable is a Tensor whose value is stored in the graph and persists across runs of the computational graph; however unlike constants defined with `tf.zeros` or `tf.random_normal`, the values of a Variable can be mutated as the graph runs; these mutations will persist across graph runs. Learnable parameters of the network are usually stored in Variables.
#
# You don't need to tune any hyperparameters, but you should achieve validation accuracies above 40% after one epoch of training.
# + id="2XIpyxxqUUnq" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623348466795, "user_tz": -120, "elapsed": 3993, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="fe88d9a5-4896-404b-cc24-fa55c7ecd2eb"
def two_layer_fc_init():
"""
Initialize the weights of a two-layer network, for use with the
two_layer_network function defined above.
You can use the `create_matrix_with_kaiming_normal` helper!
Inputs: None
Returns: A list of:
- w1: TensorFlow tf.Variable giving the weights for the first layer
- w2: TensorFlow tf.Variable giving the weights for the second layer
"""
hidden_layer_size = 4000
w1 = tf.Variable(create_matrix_with_kaiming_normal((3 * 32 * 32, 4000)))
w2 = tf.Variable(create_matrix_with_kaiming_normal((4000, 10)))
return [w1, w2]
learning_rate = 1e-2
train_part2(two_layer_fc, two_layer_fc_init, learning_rate)
# + [markdown] id="PPWP_ypQUUnq"
# ### Barebones TensorFlow: Train a three-layer ConvNet
# We will now use TensorFlow to train a three-layer ConvNet on CIFAR-10.
#
# You need to implement the `three_layer_convnet_init` function. Recall that the architecture of the network is:
#
# 1. Convolutional layer (with bias) with 32 5x5 filters, with zero-padding 2
# 2. ReLU
# 3. Convolutional layer (with bias) with 16 3x3 filters, with zero-padding 1
# 4. ReLU
# 5. Fully-connected layer (with bias) to compute scores for 10 classes
#
# You don't need to do any hyperparameter tuning, but you should see validation accuracies above 43% after one epoch of training.
# + id="barebones_accuracy" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623348883199, "user_tz": -120, "elapsed": 6692, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="e5460eab-5773-445e-ea14-4f1db3893283"
def three_layer_convnet_init():
"""
Initialize the weights of a Three-Layer ConvNet, for use with the
three_layer_convnet function defined above.
You can use the `create_matrix_with_kaiming_normal` helper!
Inputs: None
Returns a list containing:
- conv_w1: TensorFlow tf.Variable giving weights for the first conv layer
- conv_b1: TensorFlow tf.Variable giving biases for the first conv layer
- conv_w2: TensorFlow tf.Variable giving weights for the second conv layer
- conv_b2: TensorFlow tf.Variable giving biases for the second conv layer
- fc_w: TensorFlow tf.Variable giving weights for the fully-connected layer
- fc_b: TensorFlow tf.Variable giving biases for the fully-connected layer
"""
params = None
############################################################################
# TODO: Initialize the parameters of the three-layer network. #
############################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
conv_w1 = tf.Variable(create_matrix_with_kaiming_normal((5,5,3,32)))
conv_b1 = tf.Variable(tf.zeros((32,)))
conv_w2 = tf.Variable(create_matrix_with_kaiming_normal((3,3,32,16)))
conv_b2 = tf.Variable(tf.zeros((16,)))
fc_w = tf.Variable(create_matrix_with_kaiming_normal((32*32*16,10)))
fc_b = tf.Variable(tf.zeros((10,)))
params = [conv_w1, conv_b1, conv_w2, conv_b2, fc_w, fc_b]
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
############################################################################
# END OF YOUR CODE #
############################################################################
return params
learning_rate = 3e-3
train_part2(three_layer_convnet, three_layer_convnet_init, learning_rate)
# + [markdown] tags=["pdf-ignore"] id="EYo_kwbKUUnr"
# # Part III: Keras Model Subclassing API
#
# Implementing a neural network using the low-level TensorFlow API is a good way to understand how TensorFlow works, but it's a little inconvenient - we had to manually keep track of all Tensors holding learnable parameters. This was fine for a small network, but could quickly become unweildy for a large complex model.
#
# Fortunately TensorFlow 2.0 provides higher-level APIs such as `tf.keras` which make it easy to build models out of modular, object-oriented layers. Further, TensorFlow 2.0 uses eager execution that evaluates operations immediately, without explicitly constructing any computational graphs. This makes it easy to write and debug models, and reduces the boilerplate code.
#
# In this part of the notebook we will define neural network models using the `tf.keras.Model` API. To implement your own model, you need to do the following:
#
# 1. Define a new class which subclasses `tf.keras.Model`. Give your class an intuitive name that describes it, like `TwoLayerFC` or `ThreeLayerConvNet`.
# 2. In the initializer `__init__()` for your new class, define all the layers you need as class attributes. The `tf.keras.layers` package provides many common neural-network layers, like `tf.keras.layers.Dense` for fully-connected layers and `tf.keras.layers.Conv2D` for convolutional layers. Under the hood, these layers will construct `Variable` Tensors for any learnable parameters. **Warning**: Don't forget to call `super(YourModelName, self).__init__()` as the first line in your initializer!
# 3. Implement the `call()` method for your class; this implements the forward pass of your model, and defines the *connectivity* of your network. Layers defined in `__init__()` implement `__call__()` so they can be used as function objects that transform input Tensors into output Tensors. Don't define any new layers in `call()`; any layers you want to use in the forward pass should be defined in `__init__()`.
#
# After you define your `tf.keras.Model` subclass, you can instantiate it and use it like the model functions from Part II.
#
# ### Keras Model Subclassing API: Two-Layer Network
#
# Here is a concrete example of using the `tf.keras.Model` API to define a two-layer network. There are a few new bits of API to be aware of here:
#
# We use an `Initializer` object to set up the initial values of the learnable parameters of the layers; in particular `tf.initializers.VarianceScaling` gives behavior similar to the Kaiming initialization method we used in Part II. You can read more about it here: https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/initializers/VarianceScaling
#
# We construct `tf.keras.layers.Dense` objects to represent the two fully-connected layers of the model. In addition to multiplying their input by a weight matrix and adding a bias vector, these layer can also apply a nonlinearity for you. For the first layer we specify a ReLU activation function by passing `activation='relu'` to the constructor; the second layer uses softmax activation function. Finally, we use `tf.keras.layers.Flatten` to flatten the output from the previous fully-connected layer.
# + tags=["pdf-ignore-input"] id="kvgkp9yuUUnr" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623349684288, "user_tz": -120, "elapsed": 331, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="6bf2cd29-faab-4b53-fac2-5c0faf47b9d9"
class TwoLayerFC(tf.keras.Model):
def __init__(self, hidden_size, num_classes):
super(TwoLayerFC, self).__init__()
initializer = tf.initializers.VarianceScaling(scale=2.0)
self.fc1 = tf.keras.layers.Dense(hidden_size, activation='relu',
kernel_initializer=initializer)
self.fc2 = tf.keras.layers.Dense(num_classes, activation='softmax',
kernel_initializer=initializer)
self.flatten = tf.keras.layers.Flatten()
def call(self, x, training=False):
x = self.flatten(x)
x = self.fc1(x)
x = self.fc2(x)
return x
def test_TwoLayerFC():
""" A small unit test to exercise the TwoLayerFC model above. """
input_size, hidden_size, num_classes = 50, 42, 10
x = tf.zeros((64, input_size))
model = TwoLayerFC(hidden_size, num_classes)
with tf.device(device):
scores = model(x)
print(scores.shape)
test_TwoLayerFC()
# + [markdown] id="Zs00jH37UUnr"
# ### Keras Model Subclassing API: Three-Layer ConvNet
# Now it's your turn to implement a three-layer ConvNet using the `tf.keras.Model` API. Your model should have the same architecture used in Part II:
#
# 1. Convolutional layer with 5 x 5 kernels, with zero-padding of 2
# 2. ReLU nonlinearity
# 3. Convolutional layer with 3 x 3 kernels, with zero-padding of 1
# 4. ReLU nonlinearity
# 5. Fully-connected layer to give class scores
# 6. Softmax nonlinearity
#
# You should initialize the weights of your network using the same initialization method as was used in the two-layer network above.
#
# **Hint**: Refer to the documentation for `tf.keras.layers.Conv2D` and `tf.keras.layers.Dense`:
#
# https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/Conv2D
#
# https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/Dense
# + id="kAtD-lpmUUnr" executionInfo={"status": "ok", "timestamp": 1623350090409, "user_tz": -120, "elapsed": 204, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}}
class ThreeLayerConvNet(tf.keras.Model):
def __init__(self, channel_1, channel_2, num_classes):
super(ThreeLayerConvNet, self).__init__()
########################################################################
# TODO: Implement the __init__ method for a three-layer ConvNet. You #
# should instantiate layer objects to be used in the forward pass. #
########################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
initializer = tf.initializers.VarianceScaling(scale=2.0)
self.conv1 = tf.keras.layers.Conv2D(channel_1,5,padding='same',activation='relu')
self.conv2 = tf.keras.layers.Conv2D(channel_2,3,padding='same',activation='relu')
self.fc = tf.keras.layers.Dense(num_classes, activation='softmax',
kernel_initializer=initializer)
self.flatten = tf.keras.layers.Flatten()
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
########################################################################
# END OF YOUR CODE #
########################################################################
def call(self, x, training=False):
scores = None
########################################################################
# TODO: Implement the forward pass for a three-layer ConvNet. You #
# should use the layer objects defined in the __init__ method. #
########################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
scores = self.conv1(x)
scores = self.conv2(scores)
scores = self.flatten(scores)
scores = self.fc(scores)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
########################################################################
# END OF YOUR CODE #
########################################################################
return scores
# + [markdown] id="f9IO9zooUUnr"
# Once you complete the implementation of the `ThreeLayerConvNet` above you can run the following to ensure that your implementation does not crash and produces outputs of the expected shape.
# + id="keras_model_output_shape" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623350093199, "user_tz": -120, "elapsed": 326, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="f220285b-e582-4670-f8e7-687610ac359a"
def test_ThreeLayerConvNet():
channel_1, channel_2, num_classes = 12, 8, 10
model = ThreeLayerConvNet(channel_1, channel_2, num_classes)
with tf.device(device):
x = tf.zeros((64, 3, 32, 32))
scores = model(x)
print(scores.shape)
test_ThreeLayerConvNet()
# + [markdown] id="arjqNInHUUns"
# ### Keras Model Subclassing API: Eager Training
#
# While keras models have a builtin training loop (using the `model.fit`), sometimes you need more customization. Here's an example, of a training loop implemented with eager execution.
#
# In particular, notice `tf.GradientTape`. Automatic differentiation is used in the backend for implementing backpropagation in frameworks like TensorFlow. During eager execution, `tf.GradientTape` is used to trace operations for computing gradients later. A particular `tf.GradientTape` can only compute one gradient; subsequent calls to tape will throw a runtime error.
#
# TensorFlow 2.0 ships with easy-to-use built-in metrics under `tf.keras.metrics` module. Each metric is an object, and we can use `update_state()` to add observations and `reset_state()` to clear all observations. We can get the current result of a metric by calling `result()` on the metric object.
# + tags=["pdf-ignore"] id="nS2ltqw2UUns" executionInfo={"status": "ok", "timestamp": 1623350267987, "user_tz": -120, "elapsed": 325, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}}
def train_part34(model_init_fn, optimizer_init_fn, num_epochs=1, is_training=False):
"""
Simple training loop for use with models defined using tf.keras. It trains
a model for one epoch on the CIFAR-10 training set and periodically checks
accuracy on the CIFAR-10 validation set.
Inputs:
- model_init_fn: A function that takes no parameters; when called it
constructs the model we want to train: model = model_init_fn()
- optimizer_init_fn: A function which takes no parameters; when called it
constructs the Optimizer object we will use to optimize the model:
optimizer = optimizer_init_fn()
- num_epochs: The number of epochs to train for
Returns: Nothing, but prints progress during trainingn
"""
with tf.device(device):
# Compute the loss like we did in Part II
loss_fn = tf.keras.losses.SparseCategoricalCrossentropy()
model = model_init_fn()
optimizer = optimizer_init_fn()
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='train_accuracy')
val_loss = tf.keras.metrics.Mean(name='val_loss')
val_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='val_accuracy')
t = 0
for epoch in range(num_epochs):
# Reset the metrics - https://www.tensorflow.org/alpha/guide/migration_guide#new-style_metrics
train_loss.reset_states()
train_accuracy.reset_states()
for x_np, y_np in train_dset:
with tf.GradientTape() as tape:
# Use the model function to build the forward pass.
scores = model(x_np, training=is_training)
loss = loss_fn(y_np, scores)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
# Update the metrics
train_loss.update_state(loss)
train_accuracy.update_state(y_np, scores)
if t % print_every == 0:
val_loss.reset_states()
val_accuracy.reset_states()
for test_x, test_y in val_dset:
# During validation at end of epoch, training set to False
prediction = model(test_x, training=False)
t_loss = loss_fn(test_y, prediction)
val_loss.update_state(t_loss)
val_accuracy.update_state(test_y, prediction)
template = 'Iteration {}, Epoch {}, Loss: {}, Accuracy: {}, Val Loss: {}, Val Accuracy: {}'
print (template.format(t, epoch+1,
train_loss.result(),
train_accuracy.result()*100,
val_loss.result(),
val_accuracy.result()*100))
t += 1
# + [markdown] id="O2ZdifM3UUns"
# ### Keras Model Subclassing API: Train a Two-Layer Network
# We can now use the tools defined above to train a two-layer network on CIFAR-10. We define the `model_init_fn` and `optimizer_init_fn` that construct the model and optimizer respectively when called. Here we want to train the model using stochastic gradient descent with no momentum, so we construct a `tf.keras.optimizers.SGD` function; you can [read about it here](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/optimizers/SGD).
#
# You don't need to tune any hyperparameters here, but you should achieve validation accuracies above 40% after one epoch of training.
# + id="2inRV3l0UUns" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623350460652, "user_tz": -120, "elapsed": 6588, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="502ec882-2828-4a04-dc68-863c58713ede"
hidden_size, num_classes = 4000, 10
learning_rate = 1e-2
def model_init_fn():
return TwoLayerFC(hidden_size, num_classes)
def optimizer_init_fn():
return tf.keras.optimizers.SGD(learning_rate=learning_rate)
train_part34(model_init_fn, optimizer_init_fn)
# + [markdown] id="zR5idyyNUUns"
# ### Keras Model Subclassing API: Train a Three-Layer ConvNet
# Here you should use the tools we've defined above to train a three-layer ConvNet on CIFAR-10. Your ConvNet should use 32 filters in the first convolutional layer and 16 filters in the second layer.
#
# To train the model you should use gradient descent with Nesterov momentum 0.9.
#
# **HINT**: https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/optimizers/SGD
#
# You don't need to perform any hyperparameter tuning, but you should achieve validation accuracies above 50% after training for one epoch.
# + id="keras_model_accuracy" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623350636808, "user_tz": -120, "elapsed": 8246, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="32b09a1f-07c1-434b-f737-e0f79431b2b2"
learning_rate = 3e-3
channel_1, channel_2, num_classes = 32, 16, 10
def model_init_fn():
model = None
############################################################################
# TODO: Complete the implementation of model_fn. #
############################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
model = ThreeLayerConvNet(channel_1,channel_2,num_classes)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
############################################################################
# END OF YOUR CODE #
############################################################################
return model
def optimizer_init_fn():
optimizer = None
############################################################################
# TODO: Complete the implementation of model_fn. #
############################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
optimizer = tf.keras.optimizers.SGD(learning_rate, momentum=0.9,nesterov=True)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
############################################################################
# END OF YOUR CODE #
############################################################################
return optimizer
train_part34(model_init_fn, optimizer_init_fn)
# + [markdown] id="aHajkTLIUUnt"
# # Part IV: Keras Sequential API
# In Part III we introduced the `tf.keras.Model` API, which allows you to define models with any number of learnable layers and with arbitrary connectivity between layers.
#
# However for many models you don't need such flexibility - a lot of models can be expressed as a sequential stack of layers, with the output of each layer fed to the next layer as input. If your model fits this pattern, then there is an even easier way to define your model: using `tf.keras.Sequential`. You don't need to write any custom classes; you simply call the `tf.keras.Sequential` constructor with a list containing a sequence of layer objects.
#
# One complication with `tf.keras.Sequential` is that you must define the shape of the input to the model by passing a value to the `input_shape` of the first layer in your model.
#
# ### Keras Sequential API: Two-Layer Network
# In this subsection, we will rewrite the two-layer fully-connected network using `tf.keras.Sequential`, and train it using the training loop defined above.
#
# You don't need to perform any hyperparameter tuning here, but you should see validation accuracies above 40% after training for one epoch.
# + id="FjOANY-UUUnt" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623350944600, "user_tz": -120, "elapsed": 6736, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="c9e18606-f023-4d27-9314-23c4a21489b5"
learning_rate = 1e-2
def model_init_fn():
input_shape = (32, 32, 3)
hidden_layer_size, num_classes = 4000, 10
initializer = tf.initializers.VarianceScaling(scale=2.0)
layers = [
tf.keras.layers.Flatten(input_shape=input_shape),
tf.keras.layers.Dense(hidden_layer_size, activation='relu',
kernel_initializer=initializer),
tf.keras.layers.Dense(num_classes, activation='softmax',
kernel_initializer=initializer),
]
model = tf.keras.Sequential(layers)
return model
def optimizer_init_fn():
return tf.keras.optimizers.SGD(learning_rate=learning_rate)
train_part34(model_init_fn, optimizer_init_fn)
# + [markdown] id="CgwIy_1NUUnt"
# ### Abstracting Away the Training Loop
# In the previous examples, we used a customised training loop to train models (e.g. `train_part34`). Writing your own training loop is only required if you need more flexibility and control during training your model. Alternately, you can also use built-in APIs like `tf.keras.Model.fit()` and `tf.keras.Model.evaluate` to train and evaluate a model. Also remember to configure your model for training by calling `tf.keras.Model.compile.
#
# You don't need to perform any hyperparameter tuning here, but you should see validation and test accuracies above 42% after training for one epoch.
# + id="VNPZSxDJUUnt" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623351033569, "user_tz": -120, "elapsed": 7151, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="9287fbc9-6066-4633-c4e8-df0c704765c3"
model = model_init_fn()
model.compile(optimizer=tf.keras.optimizers.SGD(learning_rate=learning_rate),
loss='sparse_categorical_crossentropy',
metrics=[tf.keras.metrics.sparse_categorical_accuracy])
model.fit(X_train, y_train, batch_size=64, epochs=1, validation_data=(X_val, y_val))
model.evaluate(X_test, y_test)
# + [markdown] id="s9JQ1QVvUUnt"
# ### Keras Sequential API: Three-Layer ConvNet
# Here you should use `tf.keras.Sequential` to reimplement the same three-layer ConvNet architecture used in Part II and Part III. As a reminder, your model should have the following architecture:
#
# 1. Convolutional layer with 32 5x5 kernels, using zero padding of 2
# 2. ReLU nonlinearity
# 3. Convolutional layer with 16 3x3 kernels, using zero padding of 1
# 4. ReLU nonlinearity
# 5. Fully-connected layer giving class scores
# 6. Softmax nonlinearity
#
# You should initialize the weights of the model using a `tf.initializers.VarianceScaling` as above.
#
# You should train the model using Nesterov momentum 0.9.
#
# You don't need to perform any hyperparameter search, but you should achieve accuracy above 45% after training for one epoch.
# + id="keras_sequential_accuracy" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623351407368, "user_tz": -120, "elapsed": 9402, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="36fe1523-021e-4d45-d579-1d8f963508f1"
def model_init_fn():
model = None
############################################################################
# TODO: Construct a three-layer ConvNet using tf.keras.Sequential. #
############################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
initializer = tf.initializers.VarianceScaling(scale=2.0)
layers = [
tf.keras.layers.Conv2D(32,5,padding='same',activation='relu',input_shape=(32,32,3),kernel_initializer=initializer),
tf.keras.layers.Conv2D(32,5,padding='same',activation='relu',kernel_initializer=initializer),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(10,activation='softmax')
]
model = tf.keras.Sequential(layers)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
############################################################################
# END OF YOUR CODE #
############################################################################
return model
learning_rate = 5e-4
def optimizer_init_fn():
optimizer = None
############################################################################
# TODO: Complete the implementation of model_fn. #
############################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
optimizer = tf.keras.optimizers.SGD(learning_rate,0.9,True)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
############################################################################
# END OF YOUR CODE #
############################################################################
return optimizer
train_part34(model_init_fn, optimizer_init_fn)
# + [markdown] id="YalmGBkRUUnt"
# We will also train this model with the built-in training loop APIs provided by TensorFlow.
# + id="0d03a9u8UUnu" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623351418439, "user_tz": -120, "elapsed": 7131, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="1bb03168-2eeb-48ff-c38d-e8387a631eb9"
model = model_init_fn()
model.compile(optimizer='sgd',
loss='sparse_categorical_crossentropy',
metrics=[tf.keras.metrics.sparse_categorical_accuracy])
model.fit(X_train, y_train, batch_size=64, epochs=1, validation_data=(X_val, y_val))
model.evaluate(X_test, y_test)
# + [markdown] id="X_fxsfmjUUnu"
# ## Part IV: Functional API
# ### Demonstration with a Two-Layer Network
#
# In the previous section, we saw how we can use `tf.keras.Sequential` to stack layers to quickly build simple models. But this comes at the cost of losing flexibility.
#
# Often we will have to write complex models that have non-sequential data flows: a layer can have **multiple inputs and/or outputs**, such as stacking the output of 2 previous layers together to feed as input to a third! (Some examples are residual connections and dense blocks.)
#
# In such cases, we can use Keras functional API to write models with complex topologies such as:
#
# 1. Multi-input models
# 2. Multi-output models
# 3. Models with shared layers (the same layer called several times)
# 4. Models with non-sequential data flows (e.g. residual connections)
#
# Writing a model with Functional API requires us to create a `tf.keras.Model` instance and explicitly write input tensors and output tensors for this model.
# + tags=["pdf-ignore"] id="tqd4NtVHUUnu" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623351752558, "user_tz": -120, "elapsed": 228, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="cfd9a4e7-3a3a-4496-a7d6-7dcd72347c76"
def two_layer_fc_functional(input_shape, hidden_size, num_classes):
initializer = tf.initializers.VarianceScaling(scale=2.0)
inputs = tf.keras.Input(shape=input_shape)
flattened_inputs = tf.keras.layers.Flatten()(inputs)
fc1_output = tf.keras.layers.Dense(hidden_size, activation='relu',
kernel_initializer=initializer)(flattened_inputs)
scores = tf.keras.layers.Dense(num_classes, activation='softmax',
kernel_initializer=initializer)(fc1_output)
# Instantiate the model given inputs and outputs.
model = tf.keras.Model(inputs=inputs, outputs=scores)
return model
def test_two_layer_fc_functional():
""" A small unit test to exercise the TwoLayerFC model above. """
input_size, hidden_size, num_classes = 50, 42, 10
input_shape = (50,)
x = tf.zeros((64, input_size))
model = two_layer_fc_functional(input_shape, hidden_size, num_classes)
with tf.device(device):
scores = model(x)
print(scores.shape)
test_two_layer_fc_functional()
# + [markdown] id="BOs7P0zHUUnu"
# ### Keras Functional API: Train a Two-Layer Network
# You can now train this two-layer network constructed using the functional API.
#
# You don't need to perform any hyperparameter tuning here, but you should see validation accuracies above 40% after training for one epoch.
# + id="KYaXe_aSUUnu" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623351841990, "user_tz": -120, "elapsed": 7332, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="ebc8b910-1094-4217-b991-2bcb8ecae5da"
input_shape = (32, 32, 3)
hidden_size, num_classes = 4000, 10
learning_rate = 1e-2
def model_init_fn():
return two_layer_fc_functional(input_shape, hidden_size, num_classes)
def optimizer_init_fn():
return tf.keras.optimizers.SGD(learning_rate=learning_rate)
train_part34(model_init_fn, optimizer_init_fn)
# + [markdown] id="t_EUS-rvUUnu"
# # Part V: CIFAR-10 open-ended challenge
#
# In this section you can experiment with whatever ConvNet architecture you'd like on CIFAR-10.
#
# You should experiment with architectures, hyperparameters, loss functions, regularization, or anything else you can think of to train a model that achieves **at least 70%** accuracy on the **validation** set within 10 epochs. You can use the built-in train function, the `train_part34` function from above, or implement your own training loop.
#
# Describe what you did at the end of the notebook.
#
# ### Some things you can try:
# - **Filter size**: Above we used 5x5 and 3x3; is this optimal?
# - **Number of filters**: Above we used 16 and 32 filters. Would more or fewer do better?
# - **Pooling**: We didn't use any pooling above. Would this improve the model?
# - **Normalization**: Would your model be improved with batch normalization, layer normalization, group normalization, or some other normalization strategy?
# - **Network architecture**: The ConvNet above has only three layers of trainable parameters. Would a deeper model do better?
# - **Global average pooling**: Instead of flattening after the final convolutional layer, would global average pooling do better? This strategy is used for example in Google's Inception network and in Residual Networks.
# - **Regularization**: Would some kind of regularization improve performance? Maybe weight decay or dropout?
#
# ### NOTE: Batch Normalization / Dropout
# If you are using Batch Normalization and Dropout, remember to pass `is_training=True` if you use the `train_part34()` function. BatchNorm and Dropout layers have different behaviors at training and inference time. `training` is a specific keyword argument reserved for this purpose in any `tf.keras.Model`'s `call()` function. Read more about this here : https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/BatchNormalization#methods
# https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/Dropout#methods
#
# ### Tips for training
# For each network architecture that you try, you should tune the learning rate and other hyperparameters. When doing this there are a couple important things to keep in mind:
#
# - If the parameters are working well, you should see improvement within a few hundred iterations
# - Remember the coarse-to-fine approach for hyperparameter tuning: start by testing a large range of hyperparameters for just a few training iterations to find the combinations of parameters that are working at all.
# - Once you have found some sets of parameters that seem to work, search more finely around these parameters. You may need to train for more epochs.
# - You should use the validation set for hyperparameter search, and save your test set for evaluating your architecture on the best parameters as selected by the validation set.
#
# ### Going above and beyond
# If you are feeling adventurous there are many other features you can implement to try and improve your performance. You are **not required** to implement any of these, but don't miss the fun if you have time!
#
# - Alternative optimizers: you can try Adam, Adagrad, RMSprop, etc.
# - Alternative activation functions such as leaky ReLU, parametric ReLU, ELU, or MaxOut.
# - Model ensembles
# - Data augmentation
# - New Architectures
# - [ResNets](https://arxiv.org/abs/1512.03385) where the input from the previous layer is added to the output.
# - [DenseNets](https://arxiv.org/abs/1608.06993) where inputs into previous layers are concatenated together.
# - [This blog has an in-depth overview](https://chatbotslife.com/resnets-highwaynets-and-densenets-oh-my-9bb15918ee32)
#
# ### Have fun and happy training!
# + id="open_ended_accuracy" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623353289241, "user_tz": -120, "elapsed": 196472, "user": {"displayName": "\u7ae0\u4ef0\u5b8b", "photoUrl": "", "userId": "08102834906078443726"}} outputId="2bec1c7e-b881-425d-b8d4-f2812a196383"
class CustomConvNet(tf.keras.Model):
def __init__(self):
super(CustomConvNet, self).__init__()
############################################################################
# TODO: Construct a model that performs well on CIFAR-10 #
############################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
initializer = tf.initializers.VarianceScaling(scale=2.0)
self.conv1 = tf.keras.layers.Conv2D(128,3,padding='same',activation='relu',
kernel_initializer=initializer)
self.conv2 = tf.keras.layers.Conv2D(128,3,padding='same',activation='relu',
kernel_initializer=initializer)
self.pool2 = tf.keras.layers.MaxPooling2D(strides=(2,2))
self.conv3 = tf.keras.layers.Conv2D(128,3,padding='same',activation='relu',
kernel_initializer=initializer)
self.pool3 = tf.keras.layers.MaxPooling2D(strides=(2,2))
self.fc = tf.keras.layers.Dense(10, activation='softmax',
kernel_initializer=initializer)
self.flatten = tf.keras.layers.Flatten()
self.dropout = tf.keras.layers.Dropout(0.4)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
############################################################################
# END OF YOUR CODE #
############################################################################
def call(self, input_tensor, training=False):
############################################################################
# TODO: Construct a model that performs well on CIFAR-10 #
############################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
scores = self.conv1(input_tensor)
scores = self.dropout(scores)
scores = self.conv2(scores)
scores = self.pool2(scores)
scores = self.dropout(scores)
scores = self.conv3(scores)
scores = self.pool3(scores)
scores = self.dropout(scores)
scores = self.flatten(scores)
x = self.fc(scores)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
############################################################################
# END OF YOUR CODE #
############################################################################
return x
print_every = 700
num_epochs = 10
model = CustomConvNet()
def model_init_fn():
return CustomConvNet()
def optimizer_init_fn():
learning_rate = 1e-3
return tf.keras.optimizers.Adam(learning_rate)
train_part34(model_init_fn, optimizer_init_fn, num_epochs=num_epochs, is_training=True)
# + [markdown] tags=["pdf-inline"] id="MyrnWILaUUnv"
# ## Describe what you did
#
# In the cell below you should write an explanation of what you did, any additional features that you implemented, and/or any graphs that you made in the process of training and evaluating your network.
# + [markdown] tags=["pdf-inline"] id="YjtNgPVmUUnv"
# **Answer:**
#
#
|
assignment2/TensorFlow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import pandas as pd
# File to Load (Remember to Change These)
school_data_to_load = "Resources/schools_complete.csv"
student_data_to_load = "Resources/students_complete.csv"
# Read School and Student Data File and store into Pandas DataFrames
school_data = pd.read_csv(school_data_to_load)
student_data = pd.read_csv(student_data_to_load)
# Combine the data into a single dataset.
school_data_complete = pd.merge(student_data, school_data, how="left", on=["school_name", "school_name"])
# -
# ## District Summary
#
# * Calculate the total number of schools
#
# * Calculate the total number of students
#
# * Calculate the total budget
#
# * Calculate the average math score
#
# * Calculate the average reading score
#
# * Calculate the percentage of students with a passing math score (70 or greater)
#
# * Calculate the percentage of students with a passing reading score (70 or greater)
#
# * Calculate the percentage of students who passed math **and** reading (% Overall Passing)
#
# * Create a dataframe to hold the above results
#
# * Optional: give the displayed data cleaner formatting
# +
# Calculate the total number of schools
total_schools = len(school_data['school_name'].unique())
# Calculate the total number of students
total_students = len(student_data['student_name'])
#Calculate Total Budget
total_budget = school_data_complete['budget'].unique().sum()
#Calculate average math score
avg_math_score = school_data_complete['math_score'].mean()
# Calculate average reading score
avg_reading_score = school_data_complete['reading_score'].mean()
# Calculate the percentage of students with a passing math score (70 or greater)
students_math_with_above_70 = student_data.loc[(student_data['math_score']) >= 70].count()
percentage_students_math_with_above_70=students_math_with_above_70['math_score']/total_students * 100
# Calculate the percentage of students with a passing reading score (70 or greater)
students_reading_with_above_70 = school_data_complete.loc[(school_data_complete['reading_score']) >= 70].count()
percentage_students_reading_with_above_70 = students_reading_with_above_70['reading_score']/total_students * 100
# Calculate the percentage of students who passed math and reading (% Overall Passing)
total_pass_percentage = school_data_complete.loc[(school_data_complete['math_score'] >= 70) &
(school_data_complete['reading_score'] >= 70)
].count()
overall_pass_percentage = total_pass_percentage['Student ID']/total_students * 100
district_sumary = pd.DataFrame([{
"Total Schools": total_schools,
"Total Students": total_students,
"Total Budget": total_budget,
"Average Math Score": avg_math_score,
"Average Reading Score": avg_reading_score,
"% Passing Math": percentage_students_math_with_above_70,
"% Passing Reading": percentage_students_reading_with_above_70,
"% Overall Passing": overall_pass_percentage
}])
# formatting total students and total budget columns
district_sumary['Total Students'] = district_sumary['Total Students'].astype(int).map("{:,}".format)
district_sumary['Total Budget'] = district_sumary['Total Budget'].astype(float).map("${:,.2f}".format)
district_sumary
# -
# ## School Summary
# * Create an overview table that summarizes key metrics about each school, including:
# * School Name
# * School Type
# * Total Students
# * Total School Budget
# * Per Student Budget
# * Average Math Score
# * Average Reading Score
# * % Passing Math
# * % Passing Reading
# * % Overall Passing (The percentage of students that passed math **and** reading.)
#
# * Create a dataframe to hold the above results
# +
# create data series with only type value
grouped_school=school_data_complete.groupby(['school_name'])
# Total Students
total_students_by_school = school_data_complete.groupby('school_name')['student_name'].count()
# Total School Budget
total_budget_by_school = school_data.groupby(['school_name'])['budget'].sum()
# Per Student Budget
Per_Student_Budget=total_budget_by_school/total_students_by_school
#Average Math Score
average_math_score_by_school = student_data[['school_name','math_score']].groupby('school_name').mean()
#Average Reading Score
average_reading_score_by_school = student_data[['school_name','reading_score']].groupby('school_name').mean()
# % Passing Math
student_math_score_by_school = student_data.loc[(student_data['math_score']) >= 70].groupby('school_name').count()
Average_Math_Percentage_Score_by_school = student_math_score_by_school['math_score']/total_students_by_school * 100
# % Passing Reading
student_reading_score_by_school = student_data.loc[(student_data['reading_score']) >= 70].groupby('school_name').count()
Average_Reading_Percentage_Score_by_school = student_reading_score_by_school['reading_score']/total_students_by_school * 100
# % Overall Passing (The percentage of students that passed math and reading.)
total_pass_percentage_by_school = student_data.loc[(student_data['math_score'] >= 70) &
(student_data['reading_score'] >= 70)
].groupby('school_name').count()
overall_pass_percentage_by_school = total_pass_percentage_by_school['Student ID']/total_students_by_school * 100
# create a summary data frame
school_summary = pd.DataFrame({'School Type': grouped_school['type'].first(),
'Total Students': total_students_by_school,
"Total School Budget":total_budget_by_school,
'Per Student Budget': Per_Student_Budget,
'Average Math Score': average_math_score_by_school['math_score'],
'Average Reading Score': average_reading_score_by_school['reading_score'],
'% Passing Math': Average_Math_Percentage_Score_by_school,
'% Passing Reading': Average_Reading_Percentage_Score_by_school,
'% Overall Passing': overall_pass_percentage_by_school
})
#formatting
school_summary['Total School Budget']=school_summary['Total School Budget'].astype(float).map("${:,.2f}".format)
#school_summary['Per Student Budget']=school_summary['Per Student Budget'].map("${:,.2f}".format)
school_summary.style.format({'Per Student Budget':"${:,.2}",})
school_summary.index.name=None
school_summary
# -
# ## Top Performing Schools (By % Overall Passing)
# * Sort and display the top five performing schools by % overall passing.
school_summary.sort_values('% Overall Passing',ascending=False).head()
# ## Bottom Performing Schools (By % Overall Passing)
# * Sort and display the five worst-performing schools by % overall passing.
school_summary.sort_values('% Overall Passing',ascending=True).head()
# ## Math Scores by Grade
# * Create a table that lists the average Reading Score for students of each grade level (9th, 10th, 11th, 12th) at each school.
#
# * Create a pandas series for each grade. Hint: use a conditional statement.
#
# * Group each series by school
#
# * Combine the series into a dataframe
#
# * Optional: give the displayed data cleaner formatting
# +
avg_math_score_by_9th_grade = school_data_complete.loc[school_data_complete['grade']=='9th'].groupby('school_name')['math_score'].mean()
avg_math_score_by_10th_grade = school_data_complete.loc[school_data_complete['grade']=='10th'].groupby('school_name')['math_score'].mean()
avg_math_score_by_11th_grade = school_data_complete.loc[school_data_complete['grade']=='11th'].groupby('school_name')['math_score'].mean()
avg_math_score_by_12th_grade = school_data_complete.loc[school_data_complete['grade']=='12th'].groupby('school_name')['math_score'].mean()
Mathscores_summary = pd.DataFrame({
'9th': avg_math_score_by_9th_grade,
'10th': avg_math_score_by_10th_grade,
'11th': avg_math_score_by_11th_grade,
'12th': avg_math_score_by_12th_grade
})
Mathscores_summary
# -
# ## Reading Score by Grade
# * Perform the same operations as above for reading scores
# +
avg_reading_score_by_9th_grade = school_data_complete.loc[school_data_complete['grade']=='9th'].groupby('school_name')['reading_score'].mean()
avg_reading_score_by_10th_grade = school_data_complete.loc[school_data_complete['grade']=='10th'].groupby('school_name')['reading_score'].mean()
avg_reading_score_by_11th_grade = school_data_complete.loc[school_data_complete['grade']=='11th'].groupby('school_name')['reading_score'].mean()
avg_reading_score_by_12th_grade = school_data_complete.loc[school_data_complete['grade']=='12th'].groupby('school_name')['reading_score'].mean()
Reading_scores_summary = pd.DataFrame({
'9th': avg_reading_score_by_9th_grade,
'10th': avg_reading_score_by_10th_grade,
'11th': avg_reading_score_by_11th_grade,
'12th': avg_reading_score_by_12th_grade
})
Reading_scores_summary
# -
# ## Scores by School Spending
# * Create a table that breaks down school performances based on average Spending Ranges (Per Student). Use 4 reasonable bins to group school spending. Include in the table each of the following:
# * Average Math Score
# * Average Reading Score
# * % Passing Math
# * % Passing Reading
# * Overall Passing Rate (Average of the above two)
# +
bins = [0,584,629,644,674]
group_names=['<$584',"$585-629",'$630-644','$645-675']
Avg_Spending = school_summary.loc[:,['Average Math Score','Average Reading Score','% Passing Math','% Passing Reading','% Overall Passing']]
Avg_Spending['Spending Ranges (Per Student)'] = pd.cut(school_summary['Per Student Budget'],bins,labels=group_names,include_lowest=True)
Avg_Spending = Avg_Spending.groupby('Spending Ranges (Per Student)').mean()
Avg_Spending.style.format({
'Average Math Score':"{:,.2f}",
'Average Reading Score':"{:,.2f}",
'% Passing Math':"{:,.2f}",'% Passing Reading':"{:,.2f}",
'% Overall Passing':"{:,.2f}"})
# -
# ## Scores by School Size
# * Perform the same operations as above, based on school size.
# +
bins = [0,999.99,1999.99,9999]
group_names=['Small (<1000)',"Medium (1000-2000)",'Large (2000-5000)']
School_size = school_summary.loc[:,['Average Math Score','Average Reading Score','% Passing Math','% Passing Reading','% Overall Passing']]
School_size['School Size'] = pd.cut(school_summary['Total Students'],bins,labels=group_names,include_lowest=True)
School_size = School_size.groupby('School Size').mean()
School_size
# -
# ## Scores by School Type
# * Perform the same operations as above, based on school type
# +
School_Type = school_summary.loc[:,['School Type','Average Math Score','Average Reading Score','% Passing Math','% Passing Reading','% Overall Passing']]
School_Type = School_Type.groupby('School Type').mean()
School_Type
# -
|
PyCitySchools/PyCitySchools_starter.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py37-use_this_one
# language: python
# name: py37
# ---
# # Purpose
#
# Convert RGBA PNG image to a grayscale PNG image.
# +
from pathlib import Path
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from PIL import Image
# -
# # Get test image and look at Pillow and numpy image information
# +
test_file = Path("images") / "out0099_RGBA.png"
pil_img = Image.open(test_file)
print("PIL info")
print("size:", pil_img.size)
print("mode:", pil_img.mode)
print("")
np_img = np.array(pil_img)
print("numpy array info")
print("shape:", np_img.shape)
print("dtype:", np_img.dtype)
print("min, max:", np.min(np_img), np.max(np_img))
# -
# # Plots
# ## Pillow
fig, ax = plt.subplots()
ax.imshow(pil_img, cmap="gray", interpolation=None, vmin=0, vmax=255);
# ## Numpy array
fig, ax = plt.subplots()
ax.imshow(np_img, cmap="gray", interpolation=None, vmin=0, vmax=255);
# ## Individual numpy layers for R,G,B,A
for i in range(4):
fig, ax = plt.subplots()
ax.imshow(np_img[:,:,i], cmap="gray", interpolation=None, vmin=0, vmax=255)
ax.set_title(f"Image layer {i}");
np_img[400, 600]
# # Convert
#
# Just use the first (R) layer.
# +
new_img = Image.fromarray(np_img[:,:,0])
print("PIL info")
print("size:", new_img.size)
print("mode:", new_img.mode)
print("")
fig, ax = plt.subplots()
ax.imshow(new_img, cmap="gray", interpolation=None, vmin=0, vmax=255);
# -
# For information about Pillow image modes, see [documentation](https://pillow.readthedocs.io/en/stable/handbook/concepts.html#concept-modes):
#
# The mode of an image defines the type and depth of a pixel in the image. Each pixel uses the full range of the bit depth. So a 1-bit pixel has a range of 0-1, an 8-bit pixel has a range of 0-255 and so on. The current release supports the following standard modes:
#
# 1 (1-bit pixels, black and white, stored with one pixel per byte)
# L (8-bit pixels, black and white)
# P (8-bit pixels, mapped to any other mode using a color palette)
# RGB (3x8-bit pixels, true color)
# RGBA (4x8-bit pixels, true color with transparency mask)
# CMYK (4x8-bit pixels, color separation)
# YCbCr (3x8-bit pixels, color video format)
# Note that this refers to the JPEG, and not the ITU-R BT.2020, standard
# LAB (3x8-bit pixels, the L*a*b color space)
# HSV (3x8-bit pixels, Hue, Saturation, Value color space)
# I (32-bit signed integer pixels)
# F (32-bit floating point pixels)
#
|
python/images_convert_RGBA_to_gray.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS"
# # Can I define steps in separate files
# + [markdown] kernel="SoS"
# * **Difficulty level**: easy
# * **Time need to lean**: 10 minutes or less
# * **Key points**:
#
# + [markdown] kernel="SoS"
# ## Modularize SoS Workflow <a id="Modularize_SoS_Workflow"></a>
# + [markdown] kernel="SoS"
# SoS encourages verbatim inclusion of scripts in a single SoS Script (or Notebook) so that the entire workflow can be self-contained, easily readable and modifiable, can be easily versioned, archived and shared. Such workflows are robust to changes in dependent scripts or modulars and therefore more reproducible.
#
# However, there are certainly cases when it makes more sense to keep parts of the workflow outside of a SoS script, user cases include but not limited to
#
# 1. Parts of workflow that are meant to be the same across multiple workflows (e.g. a workflow to deliver and archive results).
# 2. Scripts that are long, or multi-file in nature, or functions that are designed to be reused by multiple scripts in the same or across SoS workflows.
# + [markdown] kernel="SoS"
# SoS provides a number of features that allow the inclusion, importing, or execution of external functions, scripts, or workflows. They are scattered around the documentation so here is a summary of what you can do to modularize your SoS workflow:
#
# |Method | Example | Pros | Cons | Comment|
# |---|---|---|---|--|
# |**utility modules and libraries** | `import my_module` in Python, `library(my_library)` in R etc | Most clean, and can be used for highly reusable components that worth the effort to formalize and maintain them. | It can be difficult to keep the modules backward compatible so changes in such modules might make a previous workflow non-reproducible. | It is usually not practical to rerun archived projects just to test compatibility of these modules. |
# |**include shared configurations or functions** | `python: input='utility.py'` | Easier to maintain than modules, include the same scripts across actions or workflows | Without proper wrapping and testing, such functions tend to be changed easily and break previous workflow | Suitable only for configuration and functions that do not need to be changed. |
# |**nested workflow**| `sos_run(source="file.sos")`| Allows the separation of a big workflow to multiple smaller workflows maintained in different SoS workflows. Very flexible because the workflows and sources can be determined programmatically.| The workflows should be logically and functionally separated so that changes in one workflow will not break the entire workflow. | Suitable for the creation of large workflows. |
# |**external commands**| `sh("sos run workflow")` in SoS or `!sos run workflow` in SoS Notebook | Similar to `sos_run` in SoS | Similar to `sos_run` in SoS | Useful for SoS notebooks with standardized sub-workflows |
# + [markdown] kernel="SoS"
# ## Further reading
#
# *
|
src/user_guide/external_scripts.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData]
# language: python
# name: conda-env-PythonData-py
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
la_df = pd.read_csv("LA_15-17_Merged.csv")
chi_df = pd.read_csv("Chicago_15-17_Merged.csv")
la_df
chi_df
chi_df["Event"].describe
plt.plot(la_df["Date"],la_df["Temp (F) avg"])
plt.plot(chi_df["Date"],chi_df["Temp (F) avg"],alpha=0.5)
plt.title("Temperature & Count of Crime Occurences")
plt.xlabel("Date")
plt.ylabel("Temperature")
plt.show()
plt.scatter(la_df["Temp (F) avg"], la_df["Total"],s=10,marker="x",alpha=0.2,label="LA")
plt.scatter(chi_df["Temp (F) avg"], chi_df["Total"],s=10,marker="+",alpha=0.2,label="Chicago")
plt.legend()
plt.title("Total Number of Crimes in Temperature")
plt.xlabel("Temperature (F)")
plt.ylabel("Total amount of Crime")
plt.show()
plt.scatter(la_df["Temp (F) avg"], la_df["Consensual"],s=10,marker="+")
plt.scatter(la_df["Temp (F) avg"], la_df["White Collar"],s=10,marker="x")
plt.scatter(la_df["Temp (F) avg"], la_df["Violent"],s=10,marker="x")
plt.scatter(la_df["Temp (F) avg"], la_df["Property"],s=10,marker="x")
plt.scatter(la_df["Temp (F) avg"], la_df["Disturbance"],s=10,marker="x")
plt.legend()
plt.show()
#plt.scatter(chi_df["Temp (F) avg"], chi_df["Consensual"],s=10,marker="+")
#plt.scatter(chi_df["Temp (F) avg"], chi_df["White Collar"],s=10,marker="x")
#plt.scatter(chi_df["Temp (F) avg"], chi_df["Violent"],s=10,marker="x")
#plt.scatter(chi_df["Temp (F) avg"], chi_df["Property"],s=10,marker="x")
#plt.scatter(chi_df["Temp (F) avg"], chi_df["Disturbance"],s=10,marker="x")
plt.legend()
plt.show()
chi_temp = chi_df.groupby("Temp (F) avg").mean()
plt.plot(chi_temp["Total"],linestyle="--")
plt.plot(chi_temp["Violent"])
plt.plot(chi_temp["Property"])
plt.plot(chi_temp["Disturbance"])
plt.plot(chi_temp["Consensual"])
plt.plot(chi_temp["White Collar"])
plt.legend()
plt.show()
plt.plot(chi_temp.index,[chi_temp["Violent"],chi_temp["Property"],chi_temp["Consensual"],
chi_temp["White Collar"],chi_temp["Disturbance"]])
plt.legend()
plt.show()
la_temp = la_df.groupby("Temp (F) avg").mean()
plt.plot(la_temp["Total"], linestyle="--")
plt.plot(la_temp["Violent"])
plt.plot(la_temp["Property"])
plt.plot(la_temp["Disturbance"])
plt.plot(la_temp["Consensual"])
plt.plot(la_temp["White Collar"])
plt.legend()
plt.show()
plt.stackplot(la_temp.index,[la_temp["Violent"],la_temp["Property"],la_temp["Consensual"],
la_temp["White Collar"],la_temp["Disturbance"]])
plt.legend()
plt.show()
# +
x1 = list(chi_df["Event"].value_counts().index)
chi_df["Clear"]=[x.find("Clear")>=0 in x for x in chi_df["Event"]]
chi_df["Thunder"]=[x.find("Thunder")>0 in x for x in chi_df["Event"]]
chi_df["Fog"]=[x.find("Fog")>=0 for x in chi_df["Event"]]
chi_df["Rain"]=[x.find("Rain")>=0 for x in chi_df["Event"]]
chi_df["Hail"]=[x.find("Hail")>=0 for x in chi_df["Event"]]
#chi_df.loc[chi_df["Thunder"]==True]
chi_df.head(15)
# -
chi_thunder = chi_df.loc[chi_df["Thunder"]==True]
#plt.scatter(chi_thunder["Temp (F) avg"],chi_thunder["Total"],s=5)
plt.scatter(chi_thunder["Temp (F) avg"],chi_thunder["Violent"],s=5)
plt.scatter(chi_thunder["Temp (F) avg"],chi_thunder["Property"],s=5)
plt.scatter(chi_thunder["Temp (F) avg"],chi_thunder["Consensual"],s=5)
plt.scatter(chi_thunder["Temp (F) avg"],chi_thunder["White Collar"],s=5)
plt.scatter(chi_thunder["Temp (F) avg"],chi_thunder["Disturbance"],s=5)
plt.legend()
plt.show()
# +
#chi_thunder = chi_df.loc[chi_df["Thunder"]==True]
chi_rain = chi_df.loc[chi_df["Rain"]==True]
chi_fog = chi_df.loc[chi_df["Fog"]==True]
chi_hail = chi_df.loc[chi_df["Hail"]==True]
#chi_clear = chi_df.loc[chi_df["Clear"]==True]
chi_w_dfs = [chi_thunder, chi_rain, chi_fog, chi_hail, chi_clear]
chi_w_labels = ["Thunder", "Rain", "Fog", "Hail", "Clear"]
for x in range(len(chi_w_dfs)):
df = chi_w_dfs[x]
l = chi_w_labels[x]
plt.scatter(df["Temp (F) avg"],df["Total"], s=5, label=l)
plt.legend()
plt.show()
# -
la_df["Year"] = la_df["Date"].map(lambda x: x[:4])
la_df.groupby("Year").sum()
chi_df["Year"] = chi_df["Date"].map(lambda x: x[:4])
chi_df.groupby("Year").sum()
la_std = la_df.groupby("Temp (F) avg").std().fillna(0)
plt.plot(la_std.index, la_temp["Total"],c="b")
plt.plot(la_std.index, la_temp["Total"]+la_std["Total"],c="b",linestyle="--",alpha=0.5)
plt.plot(la_std.index, la_temp["Total"]-la_std["Total"],c="b",linestyle="--",alpha=0.5)
plt.show()
la_df.loc[la_df["Temp (F) avg"]==48]
plt.plot(la_df["Date"],la_df["Total"],alpha=.8)
plt.plot(chi_df["Date"],chi_df["Total"],alpha=.8)
plt.xticks([])
plt.show()
plt.plot(chi_df["Date"],chi_df["Total"])
plt.xticks([])
plt.show()
crime_type_list = ["Violent", "Property", "White Collar", "Consensual", "Disturbance"]
bar_means=np.array([[x.mean()[y] for x in chi_w_dfs] for y in crime_type_list])
plt.bar(np.arange(5), bar_means[0])
plt.bar(np.arange(5), bar_means[1], bottom = bar_means[0])
plt.bar(np.arange(5), bar_means[2], bottom = sum(bar_means[:2]))
plt.bar(np.arange(5), bar_means[3], bottom = sum(bar_means[:3]))
plt.bar(np.arange(5), bar_means[4], bottom = sum(bar_means[:4]))
plt.xticks(np.arange(5),chi_w_labels)
plt.show()
|
Visualizations/Some_Plots_Kevin_v2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="G8sAZl707dlV"
# # ECE 225 Final Project - Music Chrod Progression Analysis
# + colab={} colab_type="code" id="u7677Phouw0O"
# Import packages
# %matplotlib inline
import os.path
import json
import glob
import numpy as np
from matplotlib import pyplot as plt
from collections import Counter
from matplotlib.ticker import FormatStrFormatter
# -
# Set Matplotlib default style
plt.rc('figure', titleweight='bold', dpi=100)
plt.rc('axes', labelweight='bold', linewidth=1.5, titleweight='bold')
plt.rc('xtick', direction='in')
plt.rc('ytick', direction='in')
# + [markdown] colab_type="text" id="d24i5Xfw7W9-"
# ## Data preprocessing
# + colab={} colab_type="code" id="mguXwVkUnXp0"
# Download and decompress dataset
if not os.path.exists("data/event"):
exec("wget -P data https://github.com/salu133445/music_chord_progression/raw/master/event.tar.gz")
exec("tar zxf data/event.tar.gz -C data/")
# + colab={} colab_type="code" id="4C07CQVJ7WUs"
# Constant
note_names = ['C', 'C#', 'D', 'D#', 'E', 'F', 'F#', 'G', 'G#', 'A', 'A#', 'B']
# + colab={} colab_type="code" id="oLwoXDjTw1bE"
# Load data
data, info = [], []
for filepath in glob.glob("data/event/*/*/*/*_symbol_nokey.json"):
with open(filepath) as f:
data.append(json.load(f))
with open("data/xml" + os.path.split(filepath[10:])[0] + "/song_info.json") as f:
info.append(json.load(f))
# -
# ## Metadata analysis
keys = []
bpms = []
beats_in_measures = []
for song in data:
keys.append(song['metadata']['key'])
bpms.append(int(song['metadata']['BPM']))
beats_in_measures.append(song['metadata']['beats_in_measure'])
count_keys = Counter(keys).most_common()
count_bpms = Counter(bpms).most_common()
count_beats_in_measures = Counter(beats_in_measures).most_common()
# Keys count
plt.bar(range(len(count_keys)), [x[1] * 100 / len(data) for x in count_keys])
labels = [x[0] for x in count_keys]
plt.xticks(range(len(count_keys)), labels=labels)
plt.xlabel('Key')
plt.ylabel('Frequency (%)')
plt.show()
# BPM count
plt.hist(bpms, np.arange(45,275,10))
plt.xticks(np.arange(40,280,20))
plt.xlabel('bpm (beat/minute)')
plt.ylabel('Count')
plt.show()
# Beats per measure count
plt.bar(range(len(count_beats_in_measures)), [x[1] * 100 / len(data) for x in count_beats_in_measures])
labels = [x[0] for x in count_beats_in_measures]
plt.xticks(range(len(count_beats_in_measures)), labels=labels)
plt.xlabel('# of beats per measure')
plt.ylabel('Frequency (%)')
plt.ylim(0, 100)
plt.show()
# + [markdown] colab_type="text" id="WMgEWIS38kK-"
# ## Chord-level analysis
# -
# Chord sequence without repeatness (e.g., CCFFGG -> CFG)
chord_sequences = []
song_infos = []
for song, song_info in zip(data, info):
c = [x for x in song['tracks']['chord'] if x is not None]
if not c:
continue
prev = c[0]
chord_sequence = [prev]
for now in c:
if (prev['root'] == now['root']) and (prev['quality'] == now['quality']):
continue
chord_sequence.append(now)
prev = now
chord_sequences.append(chord_sequence)
song_infos.append(song_info)
# + colab={} colab_type="code" id="bcifnVFLwC9k"
qualities = []
chords = []
for chord_sequence in chord_sequences:
for c in chord_sequence:
qualities.append(c['quality'])
chords.append(note_names[c['root']] + c['quality'])
n_chords = len(chords)
count_qualities = Counter(qualities).most_common()
count_chords = Counter(chords).most_common()
# -
# Chord quailty count
plt.bar(range(len(count_qualities)), [x[1] * 100 / n_chords for x in count_qualities])
labels = [x[0] for x in count_qualities]
labels[0] = 'M'
plt.xticks(range(len(count_qualities)), labels=labels)
plt.xlabel('Chord quality')
plt.ylabel('Frequency (%)')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 279} colab_type="code" id="Au-xHv214Dtr" outputId="b706eb2e-a2c3-4bd5-be82-ff19fd5c9e92"
# Chord count
count_chords_ = count_chords[:24]
plt.figure(figsize=(12, 3))
plt.bar(range(len(count_chords_)), [x[1] * 100 / n_chords for x in count_chords_])
plt.xticks(range(len(count_chords_)), labels=[x[0] for x in count_chords_])
plt.xlabel('Chord')
plt.ylabel('Frequency (%)')
plt.show()
# -
chord_transitions = []
chord_transition_pairs = np.zeros((15, 15))
top_chords = [x[0] for x in count_chords_[:15]]
for chord_sequence in chord_sequences:
for prev, now in zip(chord_sequence[:-1], chord_sequence[1:]):
prev_chord = note_names[prev['root']] + prev['quality']
now_chord = note_names[now['root']] + now['quality']
chord_transitions.append(
note_names[prev['root']] + prev['quality'] + '-' + note_names[now['root']] + now['quality'])
if prev_chord in top_chords and now_chord in top_chords:
chord_transition_pairs[top_chords.index(prev_chord), top_chords.index(now_chord)] += 1
n_chord_transitions = len(chord_transitions)
count_chord_transitions = Counter(chord_transitions).most_common()
# Chord transition count
count_chord_transitions_ = count_chord_transitions[:15]
plt.figure(figsize=(12, 3))
plt.bar(range(len(count_chord_transitions_)), [x[1] * 100 / n_chord_transitions for x in count_chord_transitions_])
plt.xticks(range(len(count_chord_transitions_)), labels=[x[0] for x in count_chord_transitions_])
plt.xlabel('Chord transition')
plt.ylabel('Frequency (%)')
plt.show()
plt.figure(figsize=(8, 8))
plt.imshow(chord_transition_pairs / chord_transition_pairs.sum(axis=1, keepdims=True), cmap='inferno')
plt.xticks(range(len(top_chords)), labels=top_chords)
plt.yticks(range(len(top_chords)), labels=top_chords)
plt.gca().xaxis.tick_top()
plt.colorbar()
plt.show()
three_grams = []
for chord_sequence in chord_sequences:
for prev, now, nex in zip(chord_sequence[:-2], chord_sequence[1:-1], chord_sequence[2:]):
three_grams.append(
note_names[prev['root']] + prev['quality'] + '-' + note_names[now['root']] + now['quality']
+ '-' + note_names[nex['root']] + nex['quality'])
n_three_grams = len(chord_transitions)
count_three_grams = Counter(three_grams).most_common()
# Chord three-gram count
count_three_grams_ = count_three_grams[:12]
plt.figure(figsize=(12, 3))
plt.bar(range(len(count_three_grams_)), [x[1] * 100 / n_chord_transitions for x in count_three_grams_])
plt.xticks(range(len(count_three_grams_)), labels=[x[0] for x in count_three_grams_])
plt.xlabel('Three-gram')
plt.ylabel('Frequency (%)')
plt.show()
four_grams = []
for chord_sequence in chord_sequences:
for prev, now, nex, nnex in zip(chord_sequence[:-3], chord_sequence[1:-2], chord_sequence[2:-1], chord_sequence[3:]):
if (prev['root'] == nex['root']) and (prev['quality'] == nex['quality']):
continue
if (now['root'] == nnex['root']) and (now['quality'] == nnex['quality']):
continue
four_grams.append(
note_names[prev['root']] + prev['quality']
+ '-' + note_names[now['root']] + now['quality']
+ '-' + note_names[nex['root']] + nex['quality']
+ '-' + note_names[nnex['root']] + nnex['quality'])
n_four_grams = len(chord_transitions)
count_four_grams = Counter(four_grams).most_common()
# Chord four-gram count
count_four_grams_ = count_four_grams[:10]
plt.figure(figsize=(12, 3))
plt.bar(range(len(count_four_grams_)), [x[1] * 100 / n_chord_transitions for x in count_four_grams_])
plt.xticks(range(len(count_four_grams_)), labels=[x[0] for x in count_four_grams_])
plt.xlabel('Four-gram')
plt.ylabel('Frequency (%)')
plt.show()
# ## Genre Analysis
all_genres = []
for x in info:
if x['genres']:
all_genres.extend(x['genres'])
count_genres = Counter(all_genres).most_common()
n_count_genres = len(all_genres)
top_genres = [x[0] for x in count_genres[:9]]
# Genre count
plt.figure(figsize=(12, 4))
plt.bar(range(len(count_genres)), [x[1] * 100 / n_count_genres for x in count_genres])
plt.xticks(range(len(count_genres)), labels=[x[0] for x in count_genres], rotation=90)
plt.xlabel('Genre')
plt.ylabel('Frequency (%)')
plt.show()
genre_chords = [[] for _ in range(len(top_genres))]
for (chord_sequence, song_info) in zip(chord_sequences, song_infos):
if song_info['genres']:
for g in song_info['genres']:
if g not in top_genres:
continue
for c in chord_sequence:
genre_chords[top_genres.index(g)].append(note_names[c['root']] + c['quality'])
n_genre_chords = [len(genre_chord) for genre_chord in genre_chords]
count_genre_chords = [Counter(genre_chord).most_common() for genre_chord in genre_chords]
# Chord count per genre
plt.figure(figsize=(15, 10))
plt.subplots_adjust(hspace=0.5)
for idx, (genre, count_genre_chord, n_genre_chord) in enumerate(zip(top_genres, count_genre_chords, n_genre_chords)):
plt.subplot(3, 3, idx + 1)
count_genre_chord_ = count_genre_chord[:12]
plt.bar(range(len(count_genre_chord_)), [x[1] * 100 / n_genre_chord for x in count_genre_chord_])
plt.xticks(range(len(count_genre_chord_)), labels=[x[0] for x in count_genre_chord_])
plt.yticks(np.arange(0, 17, 4))
plt.xlabel('Chord')
plt.ylabel('Frequency (%)')
plt.title(genre)
plt.show()
genre_chord_transitions = [np.zeros((10, 10)) for _ in range(len(top_genres))]
top_chords = [x[0] for x in count_chords[:10]]
for (chord_sequence, song_info) in zip(chord_sequences, song_infos):
if song_info['genres']:
for g in song_info['genres']:
if g not in top_genres:
continue
for prev, now in zip(chord_sequence[:-1], chord_sequence[1:]):
prev_chord = note_names[prev['root']] + prev['quality']
now_chord = note_names[now['root']] + now['quality']
if prev_chord in top_chords and now_chord in top_chords:
genre_chord_transitions[top_genres.index(g)][top_chords.index(prev_chord), top_chords.index(now_chord)] += 1
# Chord transition per genre
plt.figure(figsize=(12, 12))
for idx, (genre, genre_chord_transition) in enumerate(zip(top_genres, genre_chord_transitions)):
plt.subplot(3, 3, idx + 1)
plt.imshow(genre_chord_transition / genre_chord_transition.sum(axis=1, keepdims=True), cmap='inferno', vmin=0, vmax=0.6)
plt.title(genre)
plt.xticks(range(len(top_chords)), labels=top_chords)
plt.yticks(range(len(top_chords)), labels=top_chords)
plt.gca().xaxis.tick_top()
plt.colorbar()
plt.show()
|
music_chord_progression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hacker News post mining for show HN and Ask HN posts
#
# Hacker News is a site where user-submitted stories (known as "posts") are voted and commented upon.
#
# We have a data set for the posts of hacker newس. But note that it has been reduced from almost 300,000 rows to approximately 20,000 rows by removing all submissions that did not receive any comments, and then randomly sampling from the remaining submissions. you can download the data set from [here](https://www.kaggle.com/hacker-news/hacker-news-posts/home)
#
# We have to main questions
#
# 1- Ask HN or Show HN posts receive more comments on average?
# 2- The posts that created in specific time do they receive more comments on average?
#
# ## Summary of results:
#
# 3 AM is the best time to post a Ask HN post to get more comments and 11 AM to 1PM is the best time to post a Show HN post.
#
# ## opening the data set
#
#
# We start with opening and reading hacker_news.csv and showing first five rows of it.
#
#
#
#
# +
from csv import reader
def open_file(file_name):
opened_file = open(file_name, encoding = 'utf8')
read_file = reader(opened_file)
dataset = list(read_file)
return dataset[1:], dataset[0:1]
hn, hn_header = open_file("hacker_news.csv")
print(hn_header, hn[:4])
# -
# ## Categorizing the data set
#
# We divide the data set to three main groups
#
# 1- Ask HN posts
# 2- Show HN posts
# 3- Oher posts
#
# We put the rows with the title starting with Ask HN to ask_posts list, with Show HN to show_posts list and the others to other_posts list. Also we displayed the number of rows and some rows for sub data sets
#
# +
ask_posts = []
show_posts = []
other_posts = []
for row in hn:
title = row[1]
title = title.lower()
if title.startswith("ask hn"):
ask_posts.append(row)
elif title.startswith("show hn"):
show_posts.append(row)
else:
other_posts.append(row)
print(len(ask_posts))
print(len(show_posts))
print(len(other_posts))
print("\n")
print(ask_posts[0:2])
print("\n")
print(show_posts[0:2])
# -
# ## Average comments for ask and show posts
#
# In this section we are calculating average number of comments for per post for ask and show sub data sets
#
# Number of comments for each row is stored in the index 4
# We iterate over each sub data sets and calculate total number of comments for each sub data set then divide it to lengths of each sub data sets
#
# +
total_ask_comments = 0
total_show_comments = 0
for row in ask_posts:
num_comments = int(row[4])
total_ask_comments += num_comments
avg_ask_comments = total_ask_comments / len(ask_posts)
print("Ask posts average comments is equal to {}".format(avg_ask_comments))
for row in show_posts:
num_comments = int(row[4])
total_show_comments += num_comments
avg_show_comments = total_show_comments / len(show_posts)
print("Show posts average comments is equal to {}".format(avg_show_comments))
# -
# As you can see the average comments for ask posts is equal to 10.39 and for show posts is equal to 4.88. Therefore ask posts receives more comments on average
#
# Because of this we will focuses the remaining analysis just on these posts and we want to determine if ask posts created at a certain time are more likely to attract comments.
#
# In the code blow we convers the value of 'created_at' column for ask_posts sub data set to datetime.datetime object and we use hour method to only store hours for this column because we want to calculate post and comments for each hour of the day
#
# +
import datetime as dt
for row in ask_posts:
date_time = row[-1]
date_time = dt.datetime.strptime(date_time, "%m/%d/%Y %H:%M")
time_hour = date_time.hour
row[-1] = time_hour
# -
# ## Frequency tables
# In this part we create frequency table for number of posts for each hours a day and for number of comments for each hours a day.
#
# Keys for both of dictionaries are day's hours and values are number of posts for 'freq_hour_post' and number of comments for 'freq_hour_comments'
#
#
# +
freq_hour_post = {}
freq_hour_comments = {}
for row in ask_posts:
num_comments = int(row[4])
hour = row[-1]
if hour in freq_hour_post:
freq_hour_post[hour] += 1
else:
freq_hour_post[hour] = 1
if hour in freq_hour_comments:
freq_hour_comments[hour] += num_comments
else:
freq_hour_comments[hour] = num_comments
print(freq_hour_post)
print("\n")
print(freq_hour_comments)
# -
# After calculating frequency tables now is the time to calculate average number of comments per post for each hours of the day.
#
# We only need to divide the values of the 'freq_hour_comments' dictionary to values of the 'freq_hour_post' one by one for each corresponding key which are the day's hours.
#
# +
averge_comments_hour = []
for hour in freq_hour_comments:
averge_comments = freq_hour_comments[hour] / freq_hour_post[hour]
averge_comments_hour.append([hour, averge_comments])
print(averge_comments_hour)
# -
# To display better view of the above results we first changes the columns of the averge_comments_hour. In this case we have hours at first column then the corresponding average in the next column. With this we can use sorted () faction to sort this list to have a better view of our results.
#
# After that by using datetime.strptime method we create time object for each hours column then by using strftime method we transform it to our desired string format which is for example: 14:00.
#
# +
swap_averge_comments_hour = []
for row in averge_comments_hour:
date_time_p = dt.datetime.strptime(str(row[0]), "%H")
time_string = date_time_p.strftime("%H:%M")
swap_averge_comments_hour.append([row[1], time_string])
#print(swap_averge_comments_hour)
sorted_swap = sorted(swap_averge_comments_hour, reverse = True)
print("Top 6 hourse for ask posts comments average")
template = "{time}: {average:.2f} average comments per post"
for i in range(5):
print(template.format(time = sorted_swap[i][1], average = sorted_swap[i][0] ))
# -
# As we can see clearly in the results above. The ask posts which posted at 3 PM have the most comments and after that is 1 PM post. Therefore if we want to have the most comments for our Ask HN post we need to post it in 3PM Eastern Time in the US (based on the data set [documentation](https://www.kaggle.com/hacker-news/hacker-news-posts/home).
#
# ## Analysis the average points for ask and show posts
#
# In this section we will repeat somehow previous works but this time for average points for each hours a day and each two types of posts Ask HN and Show HN.
#
# First we will calculate average points for Ask and Show HN posts as in blew code. The index for points in the dataset is 3.
#
# +
total_ask_points = 0
total_show_points = 0
for row in ask_posts:
num_points = int(row[3])
total_ask_points += num_points
avg_ask_points = total_ask_points / len(ask_posts)
print("Ask posts average points is equal to {}".format(avg_ask_points))
for row in show_posts:
num_points = int(row[3])
total_show_points += num_points
avg_show_points = total_show_points / len(show_posts)
print("Show posts average points is equal to {}".format(avg_show_points))
# -
# As it's clear average points for Show HN posts is higher than ask posts then we are going to calculate average points for each days hour for Show HN data set. We will reaped previous steps for comments average. Therefore we will not explain every step explicitly (just the heading)
#
# #### Transforming 'created_at' column for show hn sub data set to datetime hour object
#
for row in show_posts:
date_time = row[-1]
date_time = dt.datetime.strptime(date_time, "%m/%d/%Y %H:%M")
time_hour = date_time.hour
row[-1] = time_hour
# #### Creating frequency tables for number of posts and points for each hours a day
# +
freq_hour_post_show = {}
freq_hour_points = {}
for row in show_posts:
num_points = int(row[3])
hour = row[-1]
if hour in freq_hour_post_show:
freq_hour_post_show[hour] += 1
else:
freq_hour_post_show[hour] = 1
if hour in freq_hour_points:
freq_hour_points[hour] += num_points
else:
freq_hour_points[hour] = num_points
print(freq_hour_post_show)
print("\n")
print(freq_hour_points)
# -
# #### Average points per post for each hours a day
# +
averge_points_hour = []
for hour in freq_hour_points:
averge_points = freq_hour_points[hour] / freq_hour_post_show[hour]
averge_points_hour.append([hour, averge_points])
print(averge_points_hour)
# -
# #### Modifing the data to have better view of results
# +
swap_averge_points_hour = []
for row in averge_points_hour:
date_time_p = dt.datetime.strptime(str(row[0]), "%H")
time_string = date_time_p.strftime("%H:%M")
swap_averge_points_hour.append([row[1], time_string])
sorted_swap_points = sorted(swap_averge_points_hour, reverse = True)
print("Top 6 hourse for show posts points average")
template_2 = "{time}: {points:.2f} average points per post"
for i in range(5):
print(template_2.format(time = sorted_swap_points[i][1], points = sorted_swap_points[i][0] ))
# -
# As we can see in the above average number of points per post for 12:00 is equal to 20.91 following by 11 AM with 19.26 and 1PM with 17.02 average points per posts. This shows us the best time to post a ask show post to get a higher points is 12:00 Eastern Time in the US (based on the data set [documentation](https://www.kaggle.com/hacker-news/hacker-news-posts/home).
#
# But because average points result for 12:00, 11:00 and 13:00 is not so different we can conclude that we can post our Show HN post from 11AM to 1PM.
#
# ## Conclusion
#
# In this project we analyzed the Hacker news web site data set. We calculated the average number of comments per post for each hour's day for Ask HN posts and average points per posts for each hours a day for Show HN posts. The result was if we want o to posts Ask HN posts and get more comments the best time is 3 PM. But if we want to post Show HN post to get more points the best time is from 11AM to 1PM.
#
|
HackerNews.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ###This notebook covers:
# ###Lesson 2A-L1 Images as Functions - Lesson 2C-L3 Aliasing
# #####Import Libraries and Dependencies
# +
import cv2
import os
import matplotlib.pyplot as plt
# %matplotlib inline
IMG = 'imgs/'
# -
# #####Some Helper Functions
def show_img(img):
"""
Function takes an image, and shows the image using pyplot.
The image is shown in RGB
"""
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
plt.imshow(img)
# #####Load and Display an Image
img_path = os.path.join(IMG, 'sea.jpeg')
img = cv2.imread(img_path)
# What is the size of the Image?
print img.shape
# What is the class of the image?
print img.dtype
show_img(img)
# #####Inspect Image Values
# My image has three channels: Red, Green, and Blue. That's why when selecting the 50th row and 100th column, three values are returned.
img[50,100]
plt.plot(img[1500,:,2])
# TODO: Extract a 2D slice between rows 101 to 103 and columns 201 to 203 (inclusive)
extraction = img[101:103,201:203]
# #####Crop Image
cropped = img[1700:,300:,:]
show_img(cropped)
# Size of Cropped Image
cropped.shape
# #####Color Planes
img_green = img[:,:,1]
plt.imshow(img_green, cmap='gray')
# #####Adding Pixels
# Load the Dog Image
dog_path = os.path.join(IMG, 'dog.jpeg')
dog = cv2.imread(dog_path)
#dog = cv2.resize
dog = cv2.resize(dog, (img.shape[1], img.shape[0]))
show_img(dog)
summed = dog + img
|
.ipynb_checkpoints/Lesson 2A-L1-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="7je53AV_Spq9"
# [](https://colab.research.google.com/github/japan-medical-ai/medical-ai-course-materials/blob/master/notebooks/Image_Segmentation.ipynb)
#
# # 実践編: MRI画像のセグメンテーション
#
# 画像を対象とした深層学習の応用技術には様々なものがあります.例えば,画像の中の個別の物体の周りを矩形で囲むようにして検出する**物体検出**や,画像内で個別物体が占める領域を認識する**画像セグメンテーション**などがあります.
#
# **物体検出**は,対象物体の**「種類」と「位置」を認識する技術**であるといえます.
#
# 
# (上図:物体検出の例.矩形で対象物体を囲い,そのクラスを答えるタスク.元画像はPascal VOCデータセットより.これにChainerCVによるFaster R-CNN(両者とも後述)を適用した結果.)
#
# **画像セグメンテーション**には2種類あります.1つは,個別の物体を区別するInstance-aware Segmentationです.もう一つは,同一クラスの物体であれば個を区別しないSemantic Segmentationです.今回は,後者を扱います.
#
# 
# (上図:Semantic Segmentationの例.ピクセル単位でクラス分類を行うタスク.画像を,予め決められた数の色で塗り絵をするようなイメージ.図はCityscapesデータセットを用いて学習したあるセグメンテーションモデルの出力結果例.)
#
# 画像セグメンテーションは,4章で扱った画像全体に対して一つのクラスを割り当てる分類問題とは異なり,画像内の全ピクセルを,ピクセルごとに分類していきます.そのため,Pixel labeling タスクとも呼ばれます.これは,対象物体の**「種類」と「位置」と「形」を認識する技術**であるといえるでしょう.
#
# 今回は,深層学習フレームワークChainerを用いて,このSemantic Segmentationタスクに取り組んでみましょう.
# + [markdown] colab_type="text" id="hvW0-s0mSpq-"
# ## 環境構築
#
# ここで用いるライブラリは,
#
# - Chainer
# - CuPy
# - ChainerCV
# - matplotlib
#
# です.Google Colab上では,以下のようにしてインストールすることができます.以下のセルを実行してください.
# + colab_type="code" id="QbU-na_HWMpR" colab={}
# !curl https://colab.chainer.org/install | sh - # ChainerとCuPyのインストール
# !pip install chainercv matplotlib # ChainerCVとmatplotlibのインストール
# + [markdown] colab_type="text" id="xQivMLFwXta7"
# インストールが完了したら,以下のセルを実行して,各ライブラリのバージョンなどを確認します.
# + colab_type="code" id="4dJCchkmSpq-" outputId="b7f8ba22-d434-4f4c-a0e0-e3edbb4458c3" colab={"base_uri": "https://localhost:8080/", "height": 287}
import chainer
import cupy
import chainercv
import matplotlib
chainer.print_runtime_info()
print('ChainerCV:', chainercv.__version__)
print('matplotlib:', matplotlib.__version__)
# + [markdown] colab_type="text" id="N6-p0PhOSprA"
# ## Semantic Segmentationについて
#
# Semantic Segmentationは,Computer Visionの分野で現在も活発に研究が行われているタスクの一つで,入力画像の画素ひとつひとつに対して,なんらかのクラスを与えていくという問題です.しかし,**人間ですら,あるピクセルひとつだけを見てそれが何かを推測するのは不可能です**.そのため,いかにして**周囲のピクセルの情報を加味**しながら,ひとつひとつのピクセルの分類を行うか,が重要となります.
#
# ニューラルネットワークを用いてこの問題を解く場合は,**「画像を入力して,画像を出力するネットワーク」**を作って学習することになります.そのため,入力画像とペアになる正解ラベル画像は,同じ大きさを持つ,各ピクセルの所属クラス番号が入ったシングルチャンネルの画像とすることが一般的です.
#
# ネットワークの出力は,$C$クラス分類をする場合は$C$チャンネルの画像になります.それを各ピクセルごとにチャンネル方向にSoftmax関数を適用して確率ベクトルにし,正解のクラスの値が大きくなるよう(高い確信をもって正解クラスを予測できるよう)にすることで学習を行います.画像分類(Classification)の際の目的関数の計算を,**ピクセルごとに行っている**と考えることもできます.そして,ピクセルごとの分類誤差を,画像サイズ分だけ足し合わせたものが最小化の対象となります.
#
# ここで,$C=2$の場合だけは,ネットワークの出力を$1$チャンネルにし,損失関数をSigmoid Cross Entropyとすることもあります.
# + [markdown] colab_type="text" id="brsImW2ySprB"
# ## 使用するデータセット
#
# これから使用するデータセットは,心臓MRI画像(短軸像)と,それに専門家がラベルを付けたものです.データについて詳しくはこちらをご参照ください[1, 2, 3].
#
# [1] Sunnybrook cardiac images from earlier competition http://smial.sri.utoronto.ca/LV_Challenge/Data.html
#
# [2] 「This "Sunnybrook Cardiac MR Database" is made available under the CC0 1.0 Universal license described above, and with more detail here: http://creativecommons.org/publicdomain/zero/1.0/」
#
# [3] Attribution:
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>. "Evaluation Framework for Algorithms Segmenting Short Axis Cardiac MRI." The MIDAS Journal -Cardiac MR Left Ventricle Segmentation Challenge, http://hdl.handle.net/10380/3070
#
# まずは,データをダウンロードします.これは配布元のデータセットを今回用いやすいように加工し終えたものです.
# + colab_type="code" id="BMtGEZWIY0EW" colab={}
# !if [ ! -d train ]; then curl -L -O https://github.com/mitmul/chainer-handson/releases/download/SegmentationDataset/train.zip && unzip train.zip && rm -rf train.zip; fi
# !if [ ! -d val ]; then curl -L -O https://github.com/mitmul/chainer-handson/releases/download/SegmentationDataset/val.zip && unzip val.zip && rm -rf val.zip; fi
# + [markdown] colab_type="text" id="xS_kqoC1m2KW"
# 次に,このデータセットから抜き出した画像ペアの例を示します.下のセルを実行してみてください.
# + colab_type="code" id="fYrsnyUNSprC" outputId="971e2a59-8c26-4b02-e323-a0d89a927a1d" colab={"base_uri": "https://localhost:8080/", "height": 204}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from PIL import Image
# PILライブラリで画像を読み込む
img = np.asarray(Image.open('train/image/000.png'))
label = np.asarray(Image.open('train/label/000.png'))
# matplotlibライブラリを使って2つの画像を並べて表示
fig, axes = plt.subplots(1, 2)
axes[0].set_axis_off()
axes[0].imshow(img, cmap='gray')
axes[1].set_axis_off()
axes[1].imshow(label, cmap='gray')
plt.show()
# + [markdown] colab_type="text" id="GGUIUVdmSprE"
# 左側がMRI画像,右側がそれに対し専門家が作成した左心室の部分をマスクした画像となっています.右側のマスク画像のうち,**白く塗りつぶされている領域が,今回見つけ出したい左心室の領域となっています**.左心室の大きさは,画像ごとに異なっており,形もまた様々です.ただし,**画像全体に対して左心室が占める領域は比較的小さい**ということは共通しています.
#
# 今回は,MRI画像データを,提供元が配布している形式(DICOM形式)から扱いやすいよう一般的な画像フォーマット(PNG)に変換して用いますが,そのための作業については説明しません.もし今回用いるMRI画像群のデータ整形の方法について興味をお持ちの方は,以前行われたKaggleのコンペティションに関連して提供されているこちらのチュートリアルをご参照ください:[Kaggle competition: Second Annual Data Science Bowl](https://www.kaggle.com/c/second-annual-data-science-bowl/details/deep-learning-tutorial) [7])
#
# 今回用いるデータセットの元となったデータは,医療画像では一般的な画像フォーマットである [DICOM](https://en.wikipedia.org/wiki/DICOM) 形式で配布されており,画像サイズは 256 x 256 のグレースケール画像になっています.今回は,これをあらかじめPNG画像に変換してあります.ラベル画像は,同じ大きさの二値画像となっており,**左心室の領域内部のピクセルは画素値として1を持ち,それ以外のピクセルは0で埋められています**.今回用いる学習用データセットは234枚の画像ペア(グレースケールのMRI画像と,対応する二値のラベル画像のペア)からなり.検証用データは,26枚の画像からなります.検証用データは学習用データとは別に用意されたものです.
#
# [7] https://www.kaggle.com/c/second-annual-data-science-bowl/details/deep-learning-tutorial
# + [markdown] colab_type="text" id="rvpjNb9uSprG"
# ### Chainerを用いた学習の流れ
#
# 今回は,4章で扱ったChainerを使ってSemantic Segmentationに取り組みます.画像から画像を出力するネットワークを記述します.
# 4章でも述べたように,Chainerには,学習ループ抽象化のためのクラスである`Trainer`が用意されています.これを用いて,左心房であるかそれ以外かの2クラスにすべてのピクセルを分類するSemantic Segmentationタスクに取り組みます.`Trainer`を使って学習を行う際にユーザがする必要がある準備について再度復習しましょう.
#
# 1. Datasetオブジェクトの準備(学習に使うデータを一つ一つ返す)
# 2. DatasetオブジェクトをIteratorにくるむ(Dataset内のデータをバッチサイズ分束ねて返す)
# 3. モデルの定義(学習対象になるニューラルネットワーク.`chainer.Chain`クラスを継承して書く)
# 4. 最適化手法の選択(`chainer.optimizers`以下にある最適化手法から選ぶ)
# 5. `Updater`オブジェクトの準備(`Iterator`と`Optimizer`をとり,実際の学習部分(パラメータアップデート)を行うもの)
# 6. `Trainer`オブジェクトの作成(学習ループの管理)
#
# `Trainer`に含まれるコンポーネントは,以下のような関係になっています.
#
# 
# + [markdown] colab_type="text" id="8MwXJDwvSprG"
# - `Updater`は,`Iterator`から`Dataset`にあるデータを指定したバッチサイズ数だけ取り出し,`Model`に与えて目的関数の値を計算し,`Optimizer`によってパラメータを更新する,という一連の作業(これが1 iterationになります)を隠蔽しています.
# - `Trainer`は`Extension`という拡張機能を使うことができ,指定したタイミング(毎iterationや,毎epoch)でログを取る,目的関数の値や精度のプロットを描画して保存,などを自動的に行うことができます.
# + [markdown] colab_type="text" id="v_Fv_HaoSprH"
# Chainerを用いてネットワークの学習を記述する場合は,上の図の**内側から順に定義していき**,最後にすべてを持った`Trainer`オブジェクトを作成し,`trainer.run()`のようにして学習を開始することになります.
#
# (`Trainer`を使わず,自分で学習ループを記述することもできますが,今回は`Trainer`を使用することを前提とします.自分で学習ループを記述する方法を知りたい場合は4章を参照してください)
# + [markdown] colab_type="text" id="-5yuQ_VBSprI"
# ## 全結合型ニューラルネットワークによるセグメンテーション
#
# まずは単純なモデルから学習を開始します.全結合層3つからなるニューラルネットワークを使って,MRI画像を入力にとり,左心室らしさのグレースケール画像を出力するモデルを学習しましょう.
# + [markdown] colab_type="text" id="FScZCHDrSprJ"
# ### データセットの準備
#
# まずはデータセットの準備をします.Chainerにはいくつかの便利なデータセットまわりのクラスが用意されています.`ImageDataset`は,画像ファイルへのファイルパスのリストを渡して初期化してやると,そのパスにある画像を**学習時に**ディスクから読み込み,それを返してくれるようなデータセットクラスです.`TupleDataset`は,複数のデータセットオブジェクトを渡して初期化すると,それらから同じインデックスを持つデータをタプルに束ねて返してくれるようなデータセットオブジェクトを作成するクラスです.(Pythonの`zip`と同様です.)
#
# 今回はSemantic Segmentationなので,入力も出力も画像です.なので,2つの`ImageDataset`オブジェクトを作成します.以下のセルを実行してください.
# + colab_type="code" id="wL7MAXtNSprK" colab={}
import glob
from chainer import datasets
def create_dataset(img_filenames, label_filenames):
img = datasets.ImageDataset(img_filenames)
img = datasets.TransformDataset(img, lambda x: x / 255.) # 0-1に正規化
label = datasets.ImageDataset(label_filenames, dtype=np.int32)
dataset = datasets.TupleDataset(img, label)
return dataset
# + [markdown] colab_type="text" id="Xucn9Z1LSprN"
# 上の関数は,入力画像のファイルパスのリスト`img_filenames`と,正解ラベル画像(0 or 1の画素値を持つ二値画像)のファイルパスのリスト`label_filenames`を与えて,2つのデータセットオブジェクトを`TupleDataset`で束ねて返すものになっています.
#
# `img`は入力画像のデータセットですが,まるで入力画像が入ったリストのように振る舞い,`img[i]`は`i`番目の画像を返します(`[i]`でアクセスしたときに初めてディスクから画像が読み込まれます).
#
# `label`も同様に,ラベル画像のリストのように振る舞います.これらを`TupleDataset`で束ねて作った`dataset`は,`dataset[i]`でアクセスすると`(img[i], label[i])`というタプル(値の2つ以上の集まり)を返すものになります.(これは`img`と`label`が同じ長さのリストの場合,`zip(img, label)`の結果と同じです.)
#
# 次に,この関数内の2行目において,`ImageDataset`で作った入力データセットを元に`TransformDataset`という新しいデータセットを作っています.`TransformDataset`は,第1引数に与えられたデータセットにアクセスする際に**第2引数に与えた関数を適用してから返す**ようにできるクラスで,任意の関数を与えてデータを変換させる処理をはさむことができます.ここでは,変換を行う関数を`lambda`関数を使って与え,単純に値域を$[0, 1]$に変換するだけの処理を行っています.この他,例えば内部で乱数によって様々な変換(画像の場合,ランダムに左右反転を行ったり,ランダムな角度で回転をしたり,などがよく行われます)を施す関数を引数として渡すことでData augmentationを簡単に実装することができます.
#
# この`create_dataset`関数を使って学習用・検証用それぞれのデータセットオブジェクトを作成しましょう.下のセルを実行してください.
# + colab_type="code" id="gLwVX846SprN" colab={}
def create_datasets():
# Python標準のglobを使ってMRI画像ファイル名/ラベル画像ファイル名の一覧を取得
train_img_filenames = sorted(glob.glob('train/image/*.png'))
train_label_filenames = sorted(glob.glob('train/label/*.png'))
# リストを渡して,データセットオブジェクト train を作成
train = create_dataset(train_img_filenames, train_label_filenames)
# 同様のことをvalidationデータに対しても行う
val_img_filenames = sorted(glob.glob('val/image/*.png'))
val_label_filenames = sorted(glob.glob('val/label/*.png'))
val = create_dataset(val_img_filenames, val_label_filenames)
return train, val
# + [markdown] colab_type="text" id="7fMFbheJSprR"
# この関数`create_dataset()`では,まずPython標準に備わっている`glob`を使って,`.png`の拡張子を持つ画像ファイルを指定したディレクトリ以下から探してきて,ファイルパスが格納されたリストを作ります.次に,入力画像とラベル画像のファイルリストが同じインデックスで対応したデータをそれぞれ指すように,`sorted`を使ってファイル名をソートしています(`glob`関数で列挙されるファイルリストは必ずしもソートされているとは限りません).そのあと,それらのファイル名リストを先程の`create_dataset`関数に渡して,データセットオブジェクトを作成しています.同様のことを検証用の画像ファイルに対しても行い,`train`と`val`2つのデータセットオブジェクトを作成して返します.
#
# ではこの関数を呼んでみましょう.下のセルを実行してください.
# + colab_type="code" id="L44uqOHXSprR" outputId="4e5a6957-775d-408b-9874-60e13d75f465" colab={"base_uri": "https://localhost:8080/", "height": 71}
train, val = create_datasets()
print('Dataset size:\n\ttrain:\t{}\n\tvalid:\t{}'.format(len(train), len(val)))
# + [markdown] colab_type="text" id="p8d5DHntSprU"
# この関数を呼べば,訓練用データセットオブジェクトと検証用データセットオブジェクトを作成できます.データセットオブジェクトは基本的にはリストとして扱うことができるます.例えば組み込み関数の`len()`を使っていくつのデータが含まれているかを知ることができます.
# + [markdown] colab_type="text" id="EawmNqfnSprU"
# ### モデルの定義
#
# 次に,訓練するモデルの定義です.ここでは4章でも扱った全結合型ニューラルネットワークを使います.
# + colab_type="code" id="2tO-E0NGSprW" colab={}
import chainer
import chainer.functions as F
import chainer.links as L
class MultiLayerPerceptron(chainer.Chain):
def __init__(self, out_h, out_w):
super().__init__()
with self.init_scope():
self.l1 = L.Linear(None, 100)
self.l2 = L.Linear(100, 100)
self.l3 = L.Linear(100, out_h * out_w)
self.out_h = out_h
self.out_w = out_w
def forward(self, x):
h = F.relu(self.l1(x))
h = F.relu(self.l2(h))
h = self.l3(h)
n = x.shape[0]
return h.reshape((n, 1, self.out_h, self.out_w))
# + [markdown] colab_type="text" id="aYdKpLRHSprY"
# ここでは3つの全結合層を使い,活性化関数にReLUを用いる形で繋げています.最後に,正解のマスク画像とそのまま比較しやすいように,画像の形にreshapeして返しています.つまり,1次元配列を2次元配列に変形しています.
#
# ここで,出力のチャンネル数は1で,各ピクセルが左心室である確率を表します.
# + [markdown] colab_type="text" id="usGCNgiRSprY"
# ### Trainerの定義
#
# 次にTrainerを定義しましょう.`Trainer`オブジェクトを作成して返してくれる`create_trainer`関数を定義しましょう.各引数の定義は以下の通りです‥
#
# - ミニバッチサイズ(batchsize)
# - 学習用データセット(train)
# - 検証用データセット(val)
# - 学習を停止するタイミング(stop)
# - 使用するデバイス(device)← `-1`にするとCPU,`>=0`の場合はそのIDを持つGPU
#
# 以下のセルを実行してください.
# + colab_type="code" id="tnzh7wZySpra" colab={}
from chainer import iterators
from chainer import training
from chainer import optimizers
from chainer.training import extensions
def create_trainer(batchsize, train, val, stop, device=-1):
# 先程定義したモデルを使用
model = MultiLayerPerceptron(out_h=256, out_w=256)
# ピクセルごとの二値分類なので,目的関数にSigmoid cross entropyを,
# 精度をはかる関数としてBinary accuracyを指定しています
train_model = L.Classifier(
model, lossfun=F.sigmoid_cross_entropy, accfun=F.binary_accuracy)
# 最適化手法にAdamを使います
optimizer = optimizers.Adam()
optimizer.setup(train_model)
# データセットから,指定したバッチサイズ数のデータ点をまとめて取り出して返すイテレータを定義します
train_iter = iterators.MultiprocessIterator(train, batchsize)
val_iter = iterators.MultiprocessIterator(val, batchsize, repeat=False, shuffle=False)
# イテレータからデータを引き出し,モデルに渡して,目的関数の値を計算し,backwardしてパラメータを更新,
# までの一連の処理を行う updater を定義します
updater = training.StandardUpdater(train_iter, optimizer, device=device)
# 様々な付加機能をExtensionとして与えられるTrainerを使います
trainer = training.trainer.Trainer(updater, stop)
logging_attributes = [
'epoch', 'main/loss', 'main/accuracy', 'val/main/loss', 'val/main/accuracy']
trainer.extend(extensions.LogReport(logging_attributes))
trainer.extend(extensions.PrintReport(logging_attributes))
trainer.extend(extensions.PlotReport(['main/loss', 'val/main/loss'], 'epoch', file_name='loss.png'))
trainer.extend(extensions.PlotReport(['main/accuracy', 'val/main/accuracy'], 'epoch', file_name='accuracy.png'))
trainer.extend(extensions.Evaluator(val_iter, optimizer.target, device=device), name='val')
return trainer
# + [markdown] colab_type="text" id="Zw7vdytqSpre"
# この関数定義の中の最後の方では,**複数の Extension を追加しています**.これらはログのファイルへの自動保存(`LogReport`)やその標準出力への表示(`PrintReport`),目的関数の値や精度のプロットの自動作成(`PlotReport`),指定したタイミングおきにvalidationデータで評価(`Evaluator`),などをしてくれる拡張機能です.
#
# この他にも様々な拡張機能が使える様になっています.こちらにある`Extension`の一覧から,使い方やできることを調べることができます: [Trainer extensions](https://docs.chainer.org/en/v2.0.2/reference/extensions.html)
# + [markdown] colab_type="text" id="5nyc3T7jSprf"
# ### 学習
#
# これで学習の準備ができました.
# あとは作成した`trainer`からrun()関数を呼び出すだけです.
#
# 下のセルを実行してください.
# + colab_type="code" id="fHzNmtxXSprg" outputId="1ca1b45f-82ab-445f-d286-b540014d4f56" colab={"base_uri": "https://localhost:8080/", "height": 431}
# %%time
trainer = create_trainer(64, train, val, (20, 'epoch'), device=0)
trainer.run()
# + [markdown] colab_type="text" id="BQY_JoQsSprj"
# 大体,学習に40秒程度かると思います.この時表示されたのは`PrintReport`というExtensionが出力したログの情報です.現在のエポック数,目的関数の値,精度(学習データセットに対してのものは`main/loss`, `main/accuracy`,検証データセットに対してのものは`val/main/loss`, `val/main/accuracy`)が表示されています.
#
# それでは次に,`PlotReport`拡張が出力したグラフを見てみましょう.学習が終了したら,以下の2つのセルを実行してみてください.
# + colab_type="code" id="gSe9PO7DSprk" outputId="5f373419-c99d-42bd-ce94-44e92f0f10ff" colab={"base_uri": "https://localhost:8080/", "height": 279}
from IPython.display import Image
Image('result/loss.png')
# + colab_type="code" id="lvnccpd5Sprl" outputId="3a458e9e-08d1-45ea-a3bb-014f730cd9ac" colab={"base_uri": "https://localhost:8080/", "height": 279}
Image('result/accuracy.png')
# + [markdown] colab_type="text" id="2RPUAqyqSpro"
# うまく学習が進んでいるようです.Training loss, Validation lossともにほぼ0近くまで下がっており,また両者のデータセットに対するAccuracyも最大の1に近づいていっています.
#
# これらのプロットは,Trainerの初期化の際に渡す`out`という引数で指定された場所に画像として保存されています.これは逐次更新されているので,実際には学習の途中でもその時点でのプロットを確認することができます.学習の進み具合を視覚的に確認するのに便利です.
# + [markdown] colab_type="text" id="W5xg70FaSpro"
# ### 評価
#
# さて,ここまでの結果をみると学習や検証データに対する性能は一見良好のようにみえます.特にAccuracyは最大値の1に近い値となっていました.しかし,この指標はどういう指標なのでしょうか?何をもって「精度(Accuracy)」と言っていたのでしょうか.
#
# 一般的にSemantic Segmentationの結果は上で「accuracy」と表示されていた**Pixel accuracy**や,それとは異なる指標である**Mean Intersection over Union (mIoU)**といった値で評価が行われます.それぞれの定義は以下のようになっています.
#
# 正解クラスが$i$であるピクセルをモデルがクラス$j$に分類した数を$N_{ij}$とすると,クラス数が $k$ のとき
#
# $$
# {\rm Pixel\ Accuracy} = \frac{\sum_{i=1}^k N_{ii}}{\sum_{i=1}^k \sum_{j=1}^k N_{ij}}
# $$
#
# $$
# {\rm mIoU} = \frac{1}{k} \sum_{i=1}^k \frac{N_{ii}}{\sum_{j=1}^k N_{ij} + \sum_{j=1}^k N_{ji} - N_{ii}}
# $$
#
# です.では,改めてこの2つの値をValidationデータセットに対して,**今学習したモデルを使って計算してみましょう.**
#
# 今回は,これらの値を計算するために,[ChainerCV](https://github.com/chainer/chainercv) [11]を用います.ChainerCVはコンピュータビジョンタスクで頻出する計算やモデル・データ等の扱いを統一的に行えるChainerの追加パッケージです.上の2つの指標をあらためて計算するために,ChainerCVが提供するSemantic Segmentationタスク用の評価指標計算のための関数を用いてみましょう.
#
# 以下のセルを実行してください.
#
# [11] <NAME>, <NAME>, <NAME>, <NAME>, "ChainerCV: a Library for Deep Learning in Computer Vision", ACM Multimedia (ACMMM), Open Source Software Competition, 2017
# + colab_type="code" id="2weVbMguSpro" outputId="c8a6f0f6-fdab-45f7-9d41-1cc2f46f1eab" colab={"base_uri": "https://localhost:8080/", "height": 53}
from chainer import cuda
from chainercv import evaluations
def evaluate(trainer, val, device=-1):
# Trainerオブジェクトから学習済みモデルを取り出す
model = trainer.updater.get_optimizer('main').target.predictor
# validationデータ全部に対して予測を行う
preds = []
for img, label in val:
img = cuda.to_gpu(img[np.newaxis], device)
pred = model(img)
pred = cuda.to_cpu(pred.data[0, 0] > 0)
preds.append((pred, label[0]))
pred_labels, gt_labels = zip(*preds)
# 評価をして結果を表示
evals = evaluations.eval_semantic_segmentation(pred_labels, gt_labels)
print('Pixel Accuracy:', evals['pixel_accuracy'])
print('mIoU:', evals['miou'])
evaluate(trainer, val, device=0)
# + [markdown] colab_type="text" id="ca5XvESYSprq"
# 2つの数字が表示されました.
#
# Pixel Accuracyの値は`PrintReport`が表示した val/main/accuracy と同じ値になっています.学習中に"accuracy"として表示していたものは,Pixel Accuracyと同じものでした.こちらは,とても高い値を示しています.最大値が1であるので0.98というのは高い数値です.
#
# 一方で,同じ最大値1の指標であるmIoU(`miou`)が思ったより低いことが分かります.なぜでしょうか.
#
# Pixel Accuracyは画像全体の画素数に対して,true positive + true negative(つまり,黒を黒,白を白と当てられた合計数) の割合を見るため,画像全体に対して negative (
# 黒)が多い場合は true positive (白を当てられた数)が小さくてもtrue negativeが大きければ結果としてPixel Accuracyは高い値になります.つまり,**class imbalance(白と黒の数が大きく違う)が起きている際に,少ないクラスへの予測誤差の影響が相対的に小さくなる**ということです.
#
# 一方,mIoU の場合は,予測と正解の両画像における「positiveとtrueの和領域」(白と予測した部分と,白が正解である領域の和)に対する「true positive」(白という予測が正解していた領域)の割合を見るので,画像全体の大きさに影響されません.わかりやすく図にすると,以下のようになります.
#
# 
#
# この図の言葉で書くと,IoUは,
#
# $$IoU = \frac{\rm true\_positive}{{\rm positive} + {\rm true} - {\rm true\_positive}}$$
#
# となります.true_positiveはTrue Positiveのピクセル数,positiveは予測画像中で1の値をとるピクセル数,trueは正解画像中で1の値をとるピクセル数です.
#
# では,実際に得られたモデルを使って validation データに予測を行った結果を可視化して,**「Pixel Accuracy は高いが mIoU が低い」ことの問題を確認してみましょう**.以下のセルを実行してください.
# + colab_type="code" id="atA-a3U2Sprr" outputId="4668e4b0-31f6-454e-8fb1-372bd16c07c6" colab={"base_uri": "https://localhost:8080/", "height": 578}
def show_predicts(trainer, val, device=-1, n_sample=3):
# Trainerオブジェクトから学習済みモデルを取り出す
model = trainer.updater.get_optimizer('main').target.predictor
for i in range(n_sample):
img, label = val[i]
img = cuda.to_gpu(img, device)
pred = model(img[np.newaxis])
pred = cuda.to_cpu(pred.data[0, 0] > 0)
fig, axes = plt.subplots(1, 2)
axes[0].set_axis_off()
axes[0].imshow(pred, cmap='gray')
axes[1].set_axis_off()
axes[1].imshow(label[0], cmap='gray')
plt.show()
show_predicts(trainer, val, device=0)
# + [markdown] colab_type="text" id="uIPy6N-ISprs"
# 左の列が予測ラベルで,右の列が正解ラベルです.3行目に顕著なように,予測のpositive領域(白い領域)は正解の領域に対して小さくなっています.Pixel Accuracyは大部分を占めている黒い部分も含めての正解率ですので,Pixel Accuracyは評価指標として今回のようなデータセットにはあまり合っていない可能性があります.それに対し`mIoU`は今回のような画像中の予測対象領域の割合が少ない場合に有効な指標となります.
#
# 以降は,どうやって`mIoU`を改善するかに取り組んでみましょう.
# + [markdown] colab_type="text" id="QwYIGRnmSprv"
# ## 畳み込みネットワークを用いたセグメンテーション
#
# mIoU改善のため,モデルを全結合層のみから構成されるものから,画像関連のタスクで多く用いられる,畳み込み層を用います.それに加えてより深い(層数の多い)モデルに変えてみましょう.今回用いるLinkは,`Convolution2D`と`Deconvolution2D`の2つだけです.それぞれ,カーネルサイズ(`ksize`),ストライド(`stride`),パディング(`pad`)を指定することができます.これらがどのように出力を変化させるかを,まずはまとめてみましょう.
# + [markdown] colab_type="text" id="7gK3nG-cSprv"
# ### Convolutionレイヤ
#
# `Convolution2D`というLinkは,一般的な畳込みレイヤの実装です.Convolutionがどのようなレイヤかは前章で説明しました.畳み込み層のパラメータを設定する際には,以下の点を知っておくと便利です.
#
# - paddingを使って計算後の出力サイズを維持しやすくするために,奇数のカーネルサイズにする($\lfloor {\rm ksize} / 2 \rfloor$をpadに指定すると,stride=1の際に画像サイズが変わらなくなる)
# - 出力feature mapを縮小したい場合は,>1の値をstrideに与える(stride=nだと変換後の画像の縦横はそれぞれ元の1/nになる)
# - 出力サイズは,$({\rm input\_size} + {\rm pad} \times 2) / {\rm stride} + 1$になる.つまり,strideを大きくすると出力特徴マップは小さくなる.
# + [markdown] colab_type="text" id="D5pnY7NrSprw"
# ### Deconvolutionレイヤ
#
# `Deconvolution2D`は,歴史的な経緯からその名とは異なり数学的な意味でのdeconvolutionではありません.実際に適用している操作からTransposed convolutionや,Backward convolutionとよばれることもあります.Deconvolution2Dはフィルタの適用の仕方はConvolutionと同じですが入力特徴マップの値を飛び飛びに配置するなどの処理が入る部分が異なる処理のことです.`Deconvolution2D`レイヤのパラメータを設定する際には,以下の点を知っておくと便利です.
#
# - カーネルサイズをstrideで割り切れる数にする(checker board artifactを防ぐため.こちらを参考のこと:[Deconvolution and Checkerboard Artifacts](https://distill.pub/2016/deconv-checkerboard/))
# - 出力サイズは,${\rm stride} \times ({\rm input\_size} - 1) + {\rm ksize} - 2 \times {\rm pad}$となるので,目的の拡大後サイズになるようパラメータを調整する
#
# Deconvolution2Dにおいては,padが意味するものが少し直感的でないため,実際に行われる操作を説明した図を以下に用意しています.
#
# 
#
# 
#
# 気をつける点は,ksizeとstrideに従って配置・拡張したfeature mapの周囲を「削る量」がpadになっている点です.そのあと行われる演算自体はstride=1, pad=0のConvolutionと同じになります.
#
# こちらに,非常にわかりやすく各種Convolution/Deconvolutionの計算を表したGIFアニメがあるので,参考にしてください:[Convolution arithmetic](https://github.com/vdumoulin/conv_arithmetic)
# + [markdown] colab_type="text" id="7toHJo-ySprx"
# ### 全畳込みネットワーク
#
# それではConvolution層とDeconvolution層からなるネットワークをChainerで書いてみます.以下のモデルは,Fully Convolutional Networkと呼ばれるネットワークに類似したものです.詳しくはこちらの文献を参照してください [4], [5], [6].
#
# 以下のFullyConvolutionalNetworkというモデルの定義には,FIXME_1 ~ FIXME_5まで,5つの定数が含まれていますが,値が与えられていません.それぞれは,Convolutionの出力側のチャンネル数になります.試しにこれを,
#
# - FIXME_1 = 64
# - FIXME_2 = 128
# - FIXME_3 = 128
# - FIXME_4 = 128
# - FIXME_5 = 128
#
# と書き換えて,下のセルを実行してみましょう.入力チャンネル数は,`None`を与えておくと,実行時に自動的に決定してくれます.
#
# [4] http://fcn.berkeleyvision.org/
#
# [5] <NAME>; "Fully Convoutional Networks for Semantic Segmentation", CVPR 2015.
#
# [6] Zeiler, Krishnan, <NAME>; "Deconvolutional Networks", CVPR 2010.
# + colab_type="code" id="B7WGsingSprx" outputId="664598d5-71c6-473b-b05b-aaaa3cc6da39" colab={"base_uri": "https://localhost:8080/", "height": 35}
from chainer import reporter
from chainer import cuda
from chainercv import evaluations
class FullyConvolutionalNetwork(chainer.Chain):
def __init__(self, out_h, out_w, n_class=1):
super().__init__()
with self.init_scope():
# L.Convolution2D(in_ch, out_ch, ksize, stride, pad)
# in_chは省略することができるので,
# L.Convolution2D(out_ch, ksize, stride, pad)
# とかくこともできます.
self.conv1 = L.Convolution2D(None, FIXME_1, ksize=5, stride=2, pad=2)
self.conv2 = L.Convolution2D(None, FIXME_2, ksize=5, stride=2, pad=2)
self.conv3 = L.Convolution2D(None, FIXME_3, ksize=3, stride=1, pad=1)
self.conv4 = L.Convolution2D(None, FIXME_4, ksize=3, stride=1, pad=1)
self.conv5 = L.Convolution2D(None, FIXME_5, ksize=1, stride=1, pad=0)
# L.Deconvolution2D(in_ch, out_ch, ksize, stride, pad)
# in_chは省略することができるので,
# L.Deconvolution2D(out_ch, ksize, stride, pad)
# と書くこともできます.
self.deconv6 = L.Deconvolution2D(None, n_class, ksize=32, stride=16, pad=8)
self.out_h = out_h
self.out_w = out_w
def forward(self, x):
h = F.relu(self.conv1(x))
h = F.max_pooling_2d(h, 2, 2)
h = F.relu(self.conv2(h))
h = F.max_pooling_2d(h, 2, 2)
h = F.relu(self.conv3(h))
h = F.relu(self.conv4(h))
h = self.conv5(h)
h = self.deconv6(h)
return h.reshape(x.shape[0], 1, h.shape[2], h.shape[3])
print(FullyConvolutionalNetwork(256, 256)(np.zeros((1, 1, 256, 256), dtype=np.float32)).shape[2:])
# + [markdown] colab_type="text" id="j7-WbsG8Spry"
# FIXME_1 ~ FIXME_5を定数に書き換えた上で上のセルを実行すると,ネットワークの出力サイズが表示されます.今回の入力画像は(256, 256)サイズの画像ですから,出力が256 x 256という同じ大きさになっていれば正しく動作しています.
# + [markdown] colab_type="text" id="6Ptqh2ghSpry"
# ### Classifierクラスの改良
#
# 次に,学習中にチェックするものとして,Pixel AccuracyだけでなくmIOUも追加するために,目的関数を計算するClassifierクラスを,自分でカスタマイズしたものに置き換えます.それは,以下のように定義されます.下記のセルを実行してみましょう.
# + colab_type="code" id="ycDjvqQtSprz" colab={}
class PixelwiseSigmoidClassifier(chainer.Chain):
def __init__(self, predictor):
super().__init__()
with self.init_scope():
# 学習対象のモデルをpredictorとして保持しておく
self.predictor = predictor
def __call__(self, x, t):
# 学習対象のモデルでまず推論を行う
y = self.predictor(x)
# 2クラス分類の誤差を計算
loss = F.sigmoid_cross_entropy(y, t)
# 予測結果(0~1の連続値を持つグレースケール画像)を二値化し,
# ChainerCVのeval_semantic_segmentation関数に正解ラベルと
# 共に渡して各種スコアを計算
y, t = cuda.to_cpu(F.sigmoid(y).data), cuda.to_cpu(t)
y = np.asarray(y > 0.5, dtype=np.int32)
y, t = y[:, 0, ...], t[:, 0, ...]
evals = evaluations.eval_semantic_segmentation(y, t)
# 学習中のログに出力
reporter.report({'loss': loss,
'miou': evals['miou'],
'pa': evals['pixel_accuracy']}, self)
return loss
# + [markdown] colab_type="text" id="xc_ZaQbnSpr1"
# Trainerは,Optimizerの引数として渡されたモデルが「目的関数の値を返す」関数であると考えます.最初のモデルではモデルは出力結果を返しましたが,それを`L.Classifier`というオブジェクトに渡した上でOptimizerに渡していました.Chainerが用意しているこの`L.Classifier`は,内部で目的関数の値だけでなくAccuracyも計算し,`reporter.report`に辞書を渡す形で`LogReport`などのExtensionが補足できるように値の報告を行います.
# しかし,`L.Classifier`はmIoUの計算をしてくれません.
#
# そこで,今回は`L.Classifier`を自前の`PixelwiseSigmoidClassifier`に置き換え,自分で実際の目的関数となる`F.sigmoid_cross_entropy`の計算を書きつつ,予測(上記コード中の`y`)に対してPixel AccuracyとmIoUの両方を計算して,報告するようにします.`__call__`自体は目的関数の値(スカラ)を返すことが期待されているので,`F.sigmoid_cross_entropy`の返り値である`loss`だけを`return`しています.
# + [markdown] colab_type="text" id="4o7iJhNZSpr1"
# ### 新しいモデルを使った学習
#
# では,これらのモデルとカスタムClassifierを使って,Trainerによる学習を行ってみましょう.以下のセルを実行してください.
# + colab_type="code" id="VOwxiL8fSpr2" colab={}
def create_trainer(batchsize, train, val, stop, device=-1, log_trigger=(1, 'epoch')):
model = FullyConvolutionalNetwork(out_h=256, out_w=256)
train_model = PixelwiseSigmoidClassifier(model)
optimizer = optimizers.Adam(eps=1e-05)
optimizer.setup(train_model)
train_iter = iterators.MultiprocessIterator(train, batchsize)
val_iter = iterators.MultiprocessIterator(val, batchsize, repeat=False, shuffle=False)
updater = training.StandardUpdater(train_iter, optimizer, device=device)
trainer = training.trainer.Trainer(updater, stop, out='result_fcn')
logging_attributes = [
'epoch', 'main/loss', 'main/miou', 'main/pa',
'val/main/loss', 'val/main/miou', 'val/main/pa']
trainer.extend(extensions.LogReport(logging_attributes), trigger=log_trigger)
trainer.extend(extensions.PrintReport(logging_attributes), trigger=log_trigger)
trainer.extend(extensions.PlotReport(['main/loss', 'val/main/loss'], 'epoch', file_name='loss.png'))
trainer.extend(extensions.PlotReport(['main/miou', 'val/main/miou'], 'epoch', file_name='miou.png'))
trainer.extend(extensions.PlotReport(['main/pa', 'val/main/pa'], 'epoch', file_name='pa.png'))
trainer.extend(extensions.Evaluator(val_iter, train_model, device=device), name='val')
trainer.extend(extensions.dump_graph('main/loss'))
return trainer
# + [markdown] colab_type="text" id="cy11SO2WSpr4"
# これが今回用いるTrainerオブジェクトを作成する関数です.最初のケースと違うところは,ログをファイルに記録する`LogReport`や標準出力にログを指定項目を出力する`PrintReport`,またグラフを出力する`PlotReport`拡張で`loss`と`accuracy`(ここでは`pa`=Pixel Accuracy)だけでなく`miou`も出力しているところです.
#
# それでは学習を開始します.最初のモデルではmiouが0.68強までしかいかなかったことを思い出しつつ,経過を見てみましょう.今回はモデルが大きくなりパラメータ数も増えているため,少し学習に時間がかかります(6分強かかります)
#
# 下記のセルを実行してください.
# + colab_type="code" id="7cGbGTEWSpr4" outputId="7fc208a8-5341-44c8-aced-9228e0a6e89c" colab={"base_uri": "https://localhost:8080/", "height": 431}
# %%time
trainer = create_trainer(128, train, val, (200, 'epoch'), device=0, log_trigger=(10, 'epoch'))
trainer.run()
# + [markdown] colab_type="text" id="nwL-7kLzSpr5"
# 学習が終了しました.`PrintReport`が出力した経過の値を見る限り,mIoUが少なくとも0.90近くまで到達していることがわかります.
# + [markdown] colab_type="text" id="q7d7QqRqSpr6"
# ### 学習結果を見てみよう
#
# では,今回の学習で`PlotReport`拡張が出力したグラフを見てみましょう.下記の3つのセルを実行してください.
# + colab_type="code" id="d4QamN-YSpr6" outputId="7087c5e8-ca01-4174-dca9-832a517c69c5" colab={"base_uri": "https://localhost:8080/", "height": 297}
from IPython.display import Image
print('Loss')
Image('result_fcn/loss.png')
# + colab_type="code" id="swNzUUkqSpr8" outputId="740d9ecb-00a2-42b8-c373-0401063d62ec" colab={"base_uri": "https://localhost:8080/", "height": 297}
print('mean IoU')
Image('result_fcn/miou.png')
# + colab_type="code" id="Vo6EPBfdSpr9" outputId="0a807603-bd6b-436a-dee3-f28d002d0506" colab={"base_uri": "https://localhost:8080/", "height": 297}
print('Pixel Accuracy')
Image('result_fcn/pa.png')
# + [markdown] colab_type="text" id="CnVA6KwQSpr_"
# Pixel Accuracyが0.99以上であるだけでなく,mIoUも0.90近くまで上がっています.mIoUに注目すると,最初のモデル(0.68程度)と比べて随分精度が上がっていることがわかると思います.実際にvalidationデータに対して推論を行った際の予測ラベル画像を見て,結果を確認しましょう.以下のセルを実行してください.
# + colab_type="code" id="pkfCHRZOSpr_" outputId="5ddf6a70-ca4a-41b3-dc2b-e0e3ba6ecca3" colab={"base_uri": "https://localhost:8080/", "height": 614}
evaluate(trainer, val, device=0)
show_predicts(trainer, val, device=0, )
# + [markdown] colab_type="text" id="NKs1onagSpsB"
# 一つ目のモデルの結果を確認した際と同じ画像が3つ並べられています.一つ目の結果よりも,特に3行目に注目すると,だいぶ正解ラベルに近い形のマスクを推定できていることがわかります.
#
# 畳み込み層だけからなるより深いモデルを学習に用いることで,大きく結果を改善することができました.
#
# + [markdown] colab_type="text" id="YpnDxZ49SpsB"
# ## さらなる精度向上へのヒント
#
# 今回のモデルでも十分うまくいっているようにみえますがまだ性能改善の余地があります.Semantic Segmentationでは,どうやって入力画像における広い範囲の情報を1つのピクセルの予測に役立てるか,どうやって複数の解像度における予測結果を考慮するか,などが重要な問題意識となります.また,ニューラルネットワークでは一般に,レイヤを重ねれば重ねるほど,特徴量の抽象度が上がっていくとされています.しかし,Semantic Segmentationでは,正確に対象物体の輪郭を表すマスク画像を出力したいので,low levelな情報(エッジ・局所的な画素値勾配のような情報,色の一貫性など)も考慮して最終的な予測結果を作りたくなります.そのために,ネットワークの出力に近いレイヤでどうやって入力に近いレイヤで取り出された特徴を活用すればよいか,が重要になってきます.
#
# これらの視点からいくつもの新しいモデルが提案されています.代表的なものを挙げると,例えば以下のようなものがあります.
#
# ### SegNet [8]
#
# 各層でMax Poolingを適用した際に「どのピクセルが最大値だったか(pooling indices)
# 」の情報をとっておき,後で画像を拡大していく時に記録しておいたpooling indicesを使ってUpsamplingする手法です.[ChainerCV](https://github.com/chainer/chainercv)にてChainerで実装されたモデル及び完全な再現実験を含むコードが公開されています.
#
# 
#
# ### U-Net [9]
#
# 下層の出力特徴マップを,上層の入力に結合することで活用する構造.全体がアルファベットの "U" のような形をしていることから「U-Net」とよばれます.セグメンテーションタスクで広く使われています.
#
# 
#
# ### PSPNet [10]
#
# 異なる大きさのsub-regionごとの特徴を大域的なコンテキストを考慮するために活用することで,ImageNet 2017 Scene Parsing Challengeで優勝したモデルです.
#
# 
#
# この他に様々な手法が提案されています.例えばクラス間のサンプル数の大小だけでなく,難しいクラスと簡単なクラスがある場合にそれらを考慮した損失関数を使うことで性能をあげることができます.
#
# また,今回は簡単のためtraining splitとvalidation splitのみを持つデータセットを使いましたが,本来はハイパーパラメータをvalidation splitでの検証結果を用いて調整したあと,最終的な性能を評価する段階でtest splitを使うべきです.
#
# [8] <NAME>, <NAME> and <NAME> "SegNet: A Deep Convolutional Encoder-Decoder Architecture for Image Segmentation." PAMI, 2017
#
# [9] <NAME>, <NAME>, <NAME>, "U-Net: Convolutional Networks for Biomedical Image Segmentation", MICCAI 2015
#
# [10] <NAME>, <NAME>, <NAME>, <NAME> and <NAME>, "Pyramid Scene Parsing Network", CVPR 2017
#
# + [markdown] colab_type="text" id="DCWZoBoCSpsC"
# ## その他の参考資料
#
# 最後に,本資料作成者によるいくつかのセグメンテーションに関する資料をここに載せます.
#
# - [最近のセグメンテーション手法の簡単な紹介](https://www.slideshare.net/mitmul/a-brief-introduction-to-recent-segmentation-methods)
# - [Pyramid Scene Parsing Network (CVPR 2017)の紹介](https://www.slideshare.net/mitmul/unofficial-pyramid-scene-parsing-network-cvpr-2017)
#
# また,以下のレビュー論文も昨今のDeep learningを活用したセグメンテーション手法についてよくまとまっており,参考になります.
#
# - [A Review on Deep Learning Techniques Applied to Semantic Segmentation](https://arxiv.org/abs/1704.06857)
|
source/source/notebooks/Image_Segmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/skojaku/Practical-Guide-to-Sentence-Transformers/blob/main/notebook/Practical_Guide_to_Sentence_Transformers.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="G7Vl68CX6mwh"
# # **Tutorial: Practical Guide to Sentence Transformers**
# _Monday, 27th Sep 2021_
#
# By <NAME>.
#
# ## References
# - *Paper*:
# Reimers, Nils, and <NAME>. 2019. “Sentence-BERT: Sentence Embeddings
# Using Siamese BERT-Networks.” EMNLP. arXiv [cs.CL]. arXiv. http://arxiv.org/abs/1908.10084.
#
#
# - *Library*:
# https://www.sbert.net/
#
#
# - *Video*:
# https://www.youtube.com/watch?v=Ey81KfQ3PQU
# + [markdown] id="K7nHP_Uv7Ugy"
# # **1. How to use sentence-transformer models**
#
# ## **1.1. Setup**
#
# First, we need the following libraries to use the sentence BERT.
#
# - [transformers](https://huggingface.co/transformers/) provides a variety of pre-trained transformer-based models.
# - [sentence-transformer](https://www.sbert.net/index.html) provides a lightweight wrapper for transformers and training procedures.
# + id="neBL0sqqceM9"
# %%capture
# !pip install -U sentence-transformers datasets==1.9.0 transformers==4.9.0
# + [markdown] id="ANVzo3FL9gJd"
# After installing the libraries, we import the modules for loading sentence transformers.
# + id="ZOfeGp2Kcn7A"
from sentence_transformers import SentenceTransformer
# + [markdown] id="GioGzK4t8EUo"
# ## **1.2. Model**
#
# Next, we need to select the transformer-based model for embedding. There are more than 15,000 models, and what model to use is a critical modeling decision.
#
# The key feature of sentence-transformers is fine-tuning, i.e., they are trained such that ***sentence embeddings*** are useful, whereas pre-trained models are trained such that ***word embeddings*** are useful. The fine-tuned models can be downloaded from [sentence-transformers library](https://www.sbert.net/docs/pretrained_models.html). You can also use pre-trained models in [Hugging Face model hub](https://huggingface.co/models).
#
# As a demonstration of sentence-transformers, we use a fine-tuned model. The model is trained on various sentence pairs in Wikipedia, scientific papers, reviews, and Q&A websites.
# + id="_UthTu4Mc9dp"
MODEL_NAME = 'paraphrase-MiniLM-L6-v2'
# + [markdown] id="Ph6xR526_HUD"
# The model can be downloaded by
# + id="FJ6dZp_-dIT_"
model = SentenceTransformer(MODEL_NAME)
#model = SentenceTransformer(MODEL_NAME, device = 0) # device = -1 == CPU, device = 0 == GPU
# + [markdown] id="n0PR6IBd_WvH"
# This method takes a list of sentences and produces an array of embedding (`numpy.ndarray`). Each row in the array is the embedding vector for a given sentence.
# + id="xqsSK_hbdK2a"
sentences = [
"This framework generates embeddings for each input sentence",
"Sentences are passed as a list of string.",
"The quick brown fox jumps over the lazy dog.",
]
# Sentences are encoded by calling model.encode()
embeddings = model.encode(sentences)
# Print the embeddings
for i, sentence in enumerate(sentences):
print("Sentence:", sentence)
print("Embedding[:10]:", embeddings[i, :10])
print("")
# + [markdown] id="n9LUjzydahwW"
# ## **1.3. Semantic search**
#
# A key feature of BERT is its ability to capture semantics. To demonstrate this, let us consider a basic NLP task:
# - You are given pairs of sentences, e.g., "He likes eating noodles", and "His favorite food is noodles".
# - You are asked to provide semantic relatedness of the given sentences.
#
# To calculate the semantic relatedness, we'll embed the given sentences and calculate the similarity.
#
# Let us consider the following sentence pairs in order of semantic relatedness.
# + id="HI5HDrbwdG-V"
sentence_pairs = [
[
"The little bird is bathing in the sink.",
"Birdie is washing itself in the water basin.",
],
[
"Two boys on a couch are playing video games.",
"Two boys are playing a video game.",
],
[
"John said he is considered a witness but not suspect.",
"'He is not a suspect anymore', John said.",
],
[
"They flew out of the nest in groups.",
"They flew into the nest together."
],
[
"The woman is playing the violin.",
"The young lady enjoys listening to the guitar.",
],
[
"The black dog is running through the snow.",
"A race car driver is driving his car through the mud.",
],
]
# + [markdown] id="8_wXgzSHdb9I"
# The first sentence is semantically equivalent although no word except 'is' and 'in' are in common (and thus a very challenging example). The second sentence pair is also semantically very similar but some details are different. The last sentence pair is semantically different.
#
# Can the sentence-transformers really capture the semantic relatedness?
# + id="OaAdkPWrdN_K"
import numpy as np
MODEL_NAME = 'paraphrase-MiniLM-L6-v2'
model = SentenceTransformer(MODEL_NAME)
def cosine_similarity_matrix(emb):
emb = np.einsum("ij,i->ij", emb, 1 / np.linalg.norm(emb, axis = 1))
return emb @ emb.T
for sentence_pair in sentence_pairs:
emb = model.encode(sentence_pair)
sim = cosine_similarity_matrix(emb)[0, 1]
print("sim = {sim:.2f}: '{sent1}' '{sent2}'".format(sent1 = sentence_pair[0], sent2 = sentence_pair[1], sim = sim))
# + [markdown] id="YbQUkoYuemNF"
# The similarity for the first sentence is relatively high even though only two general words ('in' and 'is') are in common. The second to the fourth sentences have clearly higher similarity than those of semantically less related sentence pairs (the fifth and sixth).
# + [markdown] id="PphSc5Mc6kB1"
# ## **1.4. Semantic search with pre-trained models**
#
# The `sentence-transformers` makes it easy for you to generate sentence embeddings with pre-trained models.
# Although pre-trained models are not trained for sentence embeddings, they would capture some aspects of semantic relatedness of words. With pre-trained models, the embedding for a sentence is calculated by the average of the embeddings of words in the sentence.
#
# Pre-trained models are sometimes useful because there are more than 15,000 pre-trained models trained for various tasks, whereas there are only less than 50 sentence-transformer models trained for some specific tasks.
#
# An example is sentiment analysis: given a sentence, decide whether or not the sentiment is positive or negative. As of 09/23/2021, there is no sentence-transformers model but numerous pre-trained models for sentiment analysis.
#
# Here, we use a model in [hugging models hub](https://huggingface.co/) for sentiment analysis.
# + id="yacobsF7mnMI"
PRE_TRAINED_MODEL_NAME = 'cardiffnlp/twitter-roberta-base-sentiment'
# + id="DtsCWzHO7J0X"
model = SentenceTransformer(PRE_TRAINED_MODEL_NAME)
# + [markdown] id="16imGSSUmqMz"
# Suppose that we have a list of sentences with different sentiments:
# + id="dcTyZ2uAnPs6"
sentences = [
"I love you",
"I don't like you",
"I know you",
"I like you before and although you did something good to me, I hate you",
]
# + [markdown] id="y2fh2B21nQGL"
# Our task is that, given a query sentence, rank the sentences based on sentiment similarity:
# + id="ZjFnO66gfojq"
query = "I like you" # Query sentence
emb = model.encode([query] + sentences)
sim = cosine_similarity_matrix(emb)[0, 1:]
hits = np.argsort(-sim)
for i in hits:
print("sim = {sim:.2f}: '{sent}'".format(sent=sentences[i], sim=sim[i]))
# + [markdown] id="qZSvI2Kfn9H1"
# ## **1.5. Interactive Hands-On**
# - With fine-tuned models
# 1. Go to the [sentence-transformers library](https://www.sbert.net/docs/pretrained_models.html) and find "Model Overview" section
# 2. Copy a model name into `MODEL_NAME`.
# 3. Adapt the text and make your first semantic search.
# - With pre-trained models
# 1. Go to the [Hugging Face model hub](https://huggingface.co/models) and click a model for text classification.
# 2. In the model card of a model, copy the model name at the top left and past it into `MODEL_NAME`.
# 3. Adapt the text and make your first semantic search.
# + id="RCQNWPzOpNUw"
MODEL_NAME = ""
# + id="qC1RKHQFmora"
sentences = []
# + id="CaeBiXxppJiw"
query = "" # Query sentence
model = SentenceTransformer(MODEL_NAME)
emb = model.encode([query] + sentences)
sim = cosine_similarity_matrix(emb)[0, 1:]
hits = np.argsort(-sim)
for i in hits:
print("sim = {sim:.2f}: '{sent}'".format(sent=sentences[i], sim=sim[i]))
# + [markdown] id="CgHuFFmGqOHY"
# # **2. Fine-tuning models with your data**
#
# So far, we have used existing fine-tuned or pre-trained models trained with generic texts. However, we all have different problems, and thus often want to tailor the models with our data. In the following, we will walk through how to fine-tune transformer-based models using sentence-transformer architecture.
# + [markdown] id="onMUSzGJr4uH"
# ## **2.1 Setup**
#
# To start, we need to import some libraries:
# + id="qkGaqtHsr7lG"
from sentence_transformers import (
InputExample,
SentencesDataset,
SentenceTransformer,
evaluation,
losses,
models,
)
from torch.utils.data import DataLoader
# + [markdown] id="ByIkIAgusRkG"
# ## **2.2 Define a model**
#
# `sentence-transformers` library provides building blocks to define a model for sentence-transformers. Here, we construct a sentence-transformer model with a pre-trained model, `distilroberta-base`, and average pooling layer.
# + id="_xB78JbpsRDi"
# Define the base model for word embeddings
word_embedding_model = models.Transformer("distilroberta-base", max_seq_length=512)
# Define the pooling layer that aggregates word embeddings into a sentence embedding
pooling_model = models.Pooling(
word_embedding_model.get_word_embedding_dimension(), pooling_mode="mean"
)
# Construct a sentence transformer
model = SentenceTransformer(
modules=[word_embedding_model, pooling_model],
# device=0, # Set GPU device id if GPU is available
)
# + [markdown] id="zx8bEGBCuBNE"
# ##### **The followings are alternative model designs:**
#
# Model that produces unit-norm sentence embeddings:
# ```python
# word_embedding_model = models.Transformer("distilroberta-base", max_seq_length=512)
# pooling_model = models.Pooling(
# word_embedding_model.get_word_embedding_dimension(), pooling_mode="mean"
# )
#
# # This ensures the unit norm of the sentence embedding.
# normalize_model = models.Normalize()
#
# model = SentenceTransformer(
# modules=[word_embedding_model, pooling_model, normalize_model],
# device=-1
# )
# ```
#
# Model that produces more compact sentence embeddings:
# ```python
# word_embedding_model = models.Transformer("distilroberta-base", max_seq_length=512)
#
# pooling_model = models.Pooling(
# word_embedding_model.get_word_embedding_dimension(), pooling_mode="mean"
# )
#
# # This reduces 768 dimensional embeddings to 256 dimensional embeddings.
# dense_model = models.Dense(
# in_features=pooling_model.get_sentence_embedding_dimension(),
# out_features=256,
# activation_function=nn.Tanh(),
# )
#
# model = SentenceTransformer(
# modules=[word_embedding_model, pooling_model, dense_model], device=-1
# )
# ```
# + [markdown] id="ZY2OixvTvoRg"
# ## **2.3 Training Data**
#
#
# To train the sentence-transformer, we need pairs of sentences that are semantically similar. The sentence pair should be wrapped with `InputExample`, and all pairs should be stored in `DataLoader`. For example,
#
# ```python
# train_examples = [
# InputExample(texts=["My first sentence", "My second sentence"]),
# InputExample(texts=["Another pair", "Related sentence"]),
# ]
# train_dataloader = DataLoader(train_examples, shuffle=True, batch_size=16)
# ```
#
# As a toy example, here we use the title of Physics papers. This dataset consists of 5000 pairs of papers published from the American Physical Society journals. A paper `i` is paired with another paper `j` when `i` cites `j`.
# + id="Kpurmm2KuL9u"
import pandas as pd
data_table = pd.read_csv("https://raw.githubusercontent.com/skojaku/Practical-Guide-to-Sentence-Transformers/main/data/training-data.csv")
data_table.head()
# + id="TLhMWDgS1w0i"
train_examples=[
InputExample(texts=[row["src"], row["trg"]]) for _, row in data_table.iterrows()
]
train_dataloader = DataLoader(train_examples, shuffle=True, batch_size=16)
# + [markdown] id="BEpL-LWNwKyP"
# ## **2.4 Loss function**
#
# The loss function is by far the most critical for performance. There are several loss functions available in [sentence-transformers library](https://www.sbert.net/docs/package_reference/losses.html?highlight=loss%20functions). A common choice is *triplet loss*. See [this paper](https://arxiv.org/abs/1703.07737) for details.
#
# Another important variable is the type of similarity for embeddings. Euclidean, dot-product, and cosine similarity are the commonly used metric for similarity. Here we use cosine similarity as a metric for similarity, which can be specified through `distance_metric` argument of the loss function.
# + id="f80ff0KIxIrj"
train_loss = losses.BatchSemiHardTripletLoss(
model=model,
distance_metric=losses.BatchHardTripletLossDistanceFunction.cosine_distance,
)
# + [markdown] id="uUnJgUEEyD_O"
# ## **2.5 Evaluator**
#
# The training usually takes some time, and we might want to monitor the learning progress. `evaluation` module contains various evaluators that measure the performance improvements during the training phase. See [here](https://www.sbert.net/docs/package_reference/evaluation.html?highlight=evaluators) for the available evaluators.
#
#
# + id="CzD_ZKzpyDg0"
# We will make two groups of sentece pairs, `pos_pairs` and `neg_pairs`.
# `pos_pairs` is composed of sentences paired by citatons.
pos_pairs = data_table.sample(frac = 0.1)
pos_pairs["score"] = 1 # group label
# `neg_pairs` is composed of the pairs of randomly selected sentences.
neg_pairs = data_table.copy()
neg_pairs["trg"] = neg_pairs["trg"].sample(frac = 1).values
neg_pairs = neg_pairs.sample(frac = 0.1)
neg_pairs["score"] = 0 # group label
# Concatenate the pairs
eval_data_table = pd.concat([pos_pairs, neg_pairs])
# Set up the evaluator
evaluator = evaluation.EmbeddingSimilarityEvaluator(
eval_data_table["src"].values.tolist(), # sentence
eval_data_table["trg"].values.tolist(), # sentence
scores=eval_data_table["score"].values.tolist(), # similarity
show_progress_bar=True,
)
# + [markdown] id="wjvEgdi3zkO0"
# ## **2.6. Training**
#
# Set some parameters for training:
# + id="NRPi89-Jzj0_"
num_epochs = 4
warmup_steps = 100
evaluation_steps = 1000
model_save_path = "model"
# + [markdown] id="4an59GHszYVo"
# All set. We can train the model by
# + id="H4oCQr-Pzgvu"
model.fit(
train_objectives=[(train_dataloader, train_loss)],
evaluator=evaluator,
epochs=num_epochs,
evaluation_steps=evaluation_steps,
warmup_steps=warmup_steps,
output_path=model_save_path,
)
|
notebook/Practical_Guide_to_Sentence_Transformers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="wH_vQSTZ0SsW"
import pandas as pd
import numpy as np
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
import seaborn as sns
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="FkIBeYEm1nuU" outputId="35dbca2c-208b-4179-e505-a7ca6b966607"
df = pd.read_csv('churn_data.csv')
df.head()
# + id="TbJuwXhV1ucx"
df.drop('customerID', axis = 1, inplace=True)
df['TotalCharges'] = pd.to_numeric(df['TotalCharges'],errors='coerce')
# + id="LVs08-0iY5Yg"
df = df.dropna()
# + colab={"base_uri": "https://localhost:8080/"} id="hu9l7Xwa1zRY" outputId="4e5d341e-778f-4b57-ce35-a7f83a030f99"
cat_cols = []
num_cols = []
for col in df:
if df[col].dtype == 'object':
print(f'{col}, values : {df[col].unique()}')
cat_cols.append(col)
else:
num_cols.append(col)
# + colab={"base_uri": "https://localhost:8080/"} id="VLsChAZAWOk7" outputId="eb15c4d7-59db-4368-bc53-e1cd68e66f31"
print(num_cols)
# + id="B4C18BLi8VEm"
duplicates_col = ['MultipleLines','StreamingMovies','StreamingTV', 'TechSupport','OnlineSecurity','DeviceProtection','OnlineBackup']
df[duplicates_col] = df[duplicates_col].replace('No internet service', 'No')
df['InternetService'] = df['InternetService'].replace('No internet service', 'No')
# + colab={"base_uri": "https://localhost:8080/"} id="mx1zyCTCAqhW" outputId="c9e031ee-f2bd-4751-b859-6b3d92ff305b"
for col in df:
if df[col].dtype == 'object':
print(f'{col}, values : {df[col].unique()}')
cat_cols.append(col)
else:
num_cols.append(col)
# + colab={"base_uri": "https://localhost:8080/"} id="fkkBYKR44R0g" outputId="c933fee0-0529-4bcf-9813-b3ca3d24dd5d"
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
df[cat_cols] = df[cat_cols].apply(encoder.fit_transform)
print(df[cat_cols])
# + colab={"base_uri": "https://localhost:8080/"} id="CPHeHkhj-JK5" outputId="cd6567ee-c138-46ed-9dbb-72ea06a9f035"
for col in df:
print(f'{col}, values : {df[col].unique()}')
# + id="3U6U7rc_A_j_"
from sklearn.preprocessing import MinMaxScaler
df[num_cols] = MinMaxScaler().fit_transform(df[num_cols])
# + id="oxEfJQ8icviZ"
X = df.drop('Churn', axis = 1)
y = df['Churn']
# + id="66NwmtSBbWBd"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
# + colab={"base_uri": "https://localhost:8080/"} id="_-qarvIQaYOH" outputId="b42b4002-3883-4500-c420-4c863f7d0667"
model_1 = keras.Sequential([
keras.layers.Dense(30, input_shape = (19,), activation = 'relu'),
keras.layers.Dense(15, activation = 'relu'),
keras.layers.Dense(1, activation = 'sigmoid')
])
model_1.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
model_1.fit(X_train, y_train, epochs = 150)
# + colab={"base_uri": "https://localhost:8080/"} id="wHZ3WKBGe6ZS" outputId="f9c05e10-dd8f-49f8-e538-3cfdf836705e"
model_1_params = model_1.evaluate(X_test, y_test)
y_model_1 = model_1.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="dg03wsdSfG3R" outputId="52ea9938-413c-4426-b20d-3bd30f7790c9"
model_2 = keras.Sequential([
keras.layers.Dense(30, input_shape = (19,), activation = 'relu'),
keras.layers.Dense(15, activation = 'relu'),
keras.layers.Dense(1, activation = 'sigmoid')
])
model_2.compile(optimizer='SGD', loss='binary_crossentropy', metrics=['accuracy'])
model_2.fit(X_train, y_train, epochs = 150)
# + colab={"base_uri": "https://localhost:8080/"} id="FCbWf1cifRPW" outputId="8e5e2da3-ffb9-4f9e-c39d-6b34e074543d"
model_2_params = model_2.evaluate(X_test, y_test)
y_model_2 = model_2.predict(X_test)
# + id="Cpzt_rEJfejS"
y_preds_model_1 = []
for y in y_model_1:
if y > 0.5:
y_preds_model_1.append(1)
else:
y_preds_model_1.append(0)
# + id="ozEKuyc2jEG1"
y_preds_model_2 = []
for y in y_model_2:
if y > 0.5:
y_preds_model_2.append(1)
else:
y_preds_model_2.append(0)
# + colab={"base_uri": "https://localhost:8080/"} id="JKsimB-ZjI8u" outputId="acb36806-f282-4d00-ca7c-65cc3e9d2ac9"
from sklearn.metrics import classification_report
#Resulting in a low f1 score for the class with label 1 since that the dataset used in imbalanded/need more data.
print(classification_report(y_test,y_preds_model_1))
# + colab={"base_uri": "https://localhost:8080/"} id="jUxlof54jaSe" outputId="cd4e69c2-2b5a-4f2b-d900-73acdf9942e5"
print(classification_report(y_test,y_preds_model_2))
# + colab={"base_uri": "https://localhost:8080/", "height": 459} id="F9g0n2jfjddl" outputId="fd388af5-7db0-4738-f3ea-95d329f8c60d"
#Confusion matrix for model 1
confusion_matrix = tf.math.confusion_matrix(labels=y_test,predictions=y_preds_model_1)
plt.figure(figsize = (10,7))
sns.heatmap(confusion_matrix, annot=True, fmt='d')
plt.xlabel('Predicted')
plt.ylabel('Truth')
# + colab={"base_uri": "https://localhost:8080/", "height": 459} id="JrwdXPtmk6xO" outputId="024f6723-249e-4d56-8af0-60e5216606e0"
#Confusion matrix for model 2
confusion_matrix = tf.math.confusion_matrix(labels=y_test,predictions=y_preds_model_2)
plt.figure(figsize = (10,7))
sns.heatmap(confusion_matrix, annot=True, fmt='d')
plt.xlabel('Predicted')
plt.ylabel('Truth')
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="uR4U8yTxk-EG" outputId="5d6cd325-4449-427c-e2ae-94ab1cc1f125"
#Bar plot graph showing the accuracy of each model on the test set.
plt.figure(figsize= (12, 6))
plt.bar(['Model 1', 'Model 2'], [model_1_params[1] * 100, model_2_params[1] * 100])
plt.xlabel('Model #')
plt.ylabel('Accuracy (%)')
plt.show()
|
ANN_churn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn as sns # Why sns? It's a reference to The West Wing
import matplotlib.pyplot as plt # seaborn is based on matplotlib
sns.set(color_codes=True) # adds a nice background to the graphs
# %matplotlib inline
# tells python to actually display the graphs
auto = pd.read_csv('Automobile (1).csv')
auto.head()
# ## Plotting univariate distributions
#
# The most convenient way to take a quick look at a univariate distribution in seaborn is the distplot() function. By default, this will draw a histogram and fit a kernel density estimate (KDE).
#
#
sns.distplot(auto.highway_mpg);
# we can turn the kde off and put a tic mark along the x-axis for every data point with rug
sns.rugplot(data = auto, x = 'city_mpg');
sns.histplot(auto.highway_mpg)
# ## Plotting bivariate distributions
#
# It can also be useful to visualize a relationship between two variables. The easiest way to do this in seaborn is to use the jointplot() function, which creates a scatterplot of the two variables along with the histograms of each next to the appropriate axes.
#
#
sns.jointplot(data = auto, x = 'engine_size', y = 'horsepower');
# ### Hex Bin Plots
#
# We can make a hex bin plot that breaks the 2D area into hexagons and the number of points in each hexagon determines the color
sns.jointplot(data = auto, x = 'engine_size', y = 'horsepower', kind="hex");
# ### Kernel Density Estimation
#
# We can make a 2D estimation of the density also
sns.jointplot(data = auto, x= 'engine_size', y = 'horsepower', kind="kde");
# ## Visualizing pairwise relationships in a dataset
#
# To plot multiple pairwise scatterplots in a dataset, you can use the pairplot() function. This creates a matrix of axes and shows the relationship for each pair of columns in a DataFrame, it also draws the histogram of each variable on the diagonal Axes:
#
# Be careful about toggle scrolling in the cell menu!!!
sns.pairplot(auto[['normalized_losses', 'engine_size', 'horsepower']]);
# ## Plotting with categorical data
#
#
# In a strip plot, the scatterplot points will usually overlap. This makes it difficult to see the full distribution of data. One easy solution is to adjust the positions (only along the categorical axis) using “jitter"
sns.stripplot(data = auto, x = 'fuel_type', y='horsepower', jitter=True);
# A different approach would be to use the function swarmplot(), which positions each scatterplot point on the categorical axis and avoids overlapping points:
sns.swarmplot(data = auto, x='fuel_type', y ='horsepower');
# ## Boxplots
#
# Another common graph is a boxplot(). This kind of plot shows the three quartile values of the distribution along with extreme values. The “whiskers” extend to points that lie within 1.5 IQRs of the lower and upper quartile, and then observations that fall outside this range are displayed independently.
sns.boxplot(data = auto, x ='number_of_doors', y ='horsepower');
sns.boxplot(data = auto,x='number_of_doors',y='horsepower', hue=auto['fuel_type']);
# ## Bar plots
#
# We can plot the mean of a a dataset, separated in categories using the barplot() function. When there are multiple observations in each category, it uses bootstrapping to compute a confidence interval around the estimate and plots that using error bars:
#
# Bar plots start at 0, which can sometimes be practical if zero is a number you want to compare to
sns.barplot(data = auto, x='body_style',y='horsepower', hue=auto['fuel_type']);
# A special case for the bar plot is when you want to show the number of observations in each category rather than computing the mean of a second variable. This is similar to a histogram over a categorical, rather than quantitative, variable. In seaborn, it’s easy to do so with the countplot() function:
sns.countplot(data = auto, x='body_style',hue='fuel_type');
# ## Point plots
#
# An alternative style for visualizing the same information is offered by the pointplot() function. This function also encodes the value of the estimate with height on the other axis, but rather than show a full bar it just plots the point estimate and confidence interval. Additionally, pointplot connects points from the same hue category. This makes it easy to see how the main relationship is changing as a function of a second variable, because your eyes are quite good at picking up on differences of slopes:
sns.pointplot(data = auto,x='body_style',y= 'horsepower', hue='number_of_doors');
# ## Drawing multi-panel categorical plots
sns.catplot(x="fuel_type",
y = "horsepower",
hue="number_of_doors",
col="drive_wheels",
data=auto,
kind="box");
# Various types of kind input : {``point``, ``bar``, ``count``, ``box``, ``violin``, ``strip``}
# ## Function to draw linear regression models
#
# lmplot() is one of the most widely used function to quickly plot the Linear Relationship between 2 variables
sns.lmplot(y="horsepower", x="engine_size", data=auto);
sns.lmplot(y="horsepower", x="engine_size",hue="fuel_type", data=auto);
|
notebooks/Visualization-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# # Automated Machine Learning
#
# #### Forecasting away from training data
#
#
# ## Contents
# 1. [Introduction](#Introduction)
# 2. [Setup](#Setup)
# 3. [Data](#Data)
# 4. [Prepare remote compute and data.](#prepare_remote)
# 4. [Create the configuration and train a forecaster](#train)
# 5. [Forecasting from the trained model](#forecasting)
# 6. [Forecasting away from training data](#forecasting_away)
# ## Introduction
# This notebook demonstrates the full interface of the `forecast()` function.
#
# The best known and most frequent usage of `forecast` enables forecasting on test sets that immediately follows training data.
#
# However, in many use cases it is necessary to continue using the model for some time before retraining it. This happens especially in **high frequency forecasting** when forecasts need to be made more frequently than the model can be retrained. Examples are in Internet of Things and predictive cloud resource scaling.
#
# Here we show how to use the `forecast()` function when a time gap exists between training data and prediction period.
#
# Terminology:
# * forecast origin: the last period when the target value is known
# * forecast periods(s): the period(s) for which the value of the target is desired.
# * lookback: how many past periods (before forecast origin) the model function depends on. The larger of number of lags and length of rolling window.
# * prediction context: `lookback` periods immediately preceding the forecast origin
#
# 
# ## Setup
# Please make sure you have followed the `configuration.ipynb` notebook so that your ML workspace information is saved in the config file.
# +
import os
import pandas as pd
import numpy as np
import logging
import warnings
import azureml.core
from azureml.core.dataset import Dataset
from pandas.tseries.frequencies import to_offset
from azureml.core.compute import AmlCompute
from azureml.core.compute import ComputeTarget
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
# Squash warning messages for cleaner output in the notebook
warnings.showwarning = lambda *args, **kwargs: None
np.set_printoptions(precision=4, suppress=True, linewidth=120)
# -
# This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
print("This notebook was created using version 1.21.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
# +
from azureml.core.workspace import Workspace
from azureml.core.experiment import Experiment
from azureml.train.automl import AutoMLConfig
ws = Workspace.from_config()
# choose a name for the run history container in the workspace
experiment_name = 'automl-forecast-function-demo'
experiment = Experiment(ws, experiment_name)
output = {}
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['SKU'] = ws.sku
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Run History Name'] = experiment_name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
# -
# ## Data
# For the demonstration purposes we will generate the data artificially and use them for the forecasting.
# +
TIME_COLUMN_NAME = 'date'
TIME_SERIES_ID_COLUMN_NAME = 'time_series_id'
TARGET_COLUMN_NAME = 'y'
def get_timeseries(train_len: int,
test_len: int,
time_column_name: str,
target_column_name: str,
time_series_id_column_name: str,
time_series_number: int = 1,
freq: str = 'H'):
"""
Return the time series of designed length.
:param train_len: The length of training data (one series).
:type train_len: int
:param test_len: The length of testing data (one series).
:type test_len: int
:param time_column_name: The desired name of a time column.
:type time_column_name: str
:param time_series_number: The number of time series in the data set.
:type time_series_number: int
:param freq: The frequency string representing pandas offset.
see https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html
:type freq: str
:returns: the tuple of train and test data sets.
:rtype: tuple
"""
data_train = [] # type: List[pd.DataFrame]
data_test = [] # type: List[pd.DataFrame]
data_length = train_len + test_len
for i in range(time_series_number):
X = pd.DataFrame({
time_column_name: pd.date_range(start='2000-01-01',
periods=data_length,
freq=freq),
target_column_name: np.arange(data_length).astype(float) + np.random.rand(data_length) + i*5,
'ext_predictor': np.asarray(range(42, 42 + data_length)),
time_series_id_column_name: np.repeat('ts{}'.format(i), data_length)
})
data_train.append(X[:train_len])
data_test.append(X[train_len:])
X_train = pd.concat(data_train)
y_train = X_train.pop(target_column_name).values
X_test = pd.concat(data_test)
y_test = X_test.pop(target_column_name).values
return X_train, y_train, X_test, y_test
n_test_periods = 6
n_train_periods = 30
X_train, y_train, X_test, y_test = get_timeseries(train_len=n_train_periods,
test_len=n_test_periods,
time_column_name=TIME_COLUMN_NAME,
target_column_name=TARGET_COLUMN_NAME,
time_series_id_column_name=TIME_SERIES_ID_COLUMN_NAME,
time_series_number=2)
# -
# Let's see what the training data looks like.
X_train.tail()
# plot the example time series
import matplotlib.pyplot as plt
whole_data = X_train.copy()
target_label = 'y'
whole_data[target_label] = y_train
for g in whole_data.groupby('time_series_id'):
plt.plot(g[1]['date'].values, g[1]['y'].values, label=g[0])
plt.legend()
plt.show()
# ### Prepare remote compute and data. <a id="prepare_remote"></a>
# The [Machine Learning service workspace](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-workspace), is paired with the storage account, which contains the default data store. We will use it to upload the artificial data and create [tabular dataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.tabulardataset?view=azure-ml-py) for training. A tabular dataset defines a series of lazily-evaluated, immutable operations to load data from the data source into tabular representation.
# We need to save thw artificial data and then upload them to default workspace datastore.
DATA_PATH = "fc_fn_data"
DATA_PATH_X = "{}/data_train.csv".format(DATA_PATH)
if not os.path.isdir('data'):
os.mkdir('data')
pd.DataFrame(whole_data).to_csv("data/data_train.csv", index=False)
# Upload saved data to the default data store.
ds = ws.get_default_datastore()
ds.upload(src_dir='./data', target_path=DATA_PATH, overwrite=True, show_progress=True)
train_data = Dataset.Tabular.from_delimited_files(path=ds.path(DATA_PATH_X))
# You will need to create a [compute target](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute) for your AutoML run. In this tutorial, you create AmlCompute as your training compute resource.
# +
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
amlcompute_cluster_name = "fcfn-cluster"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=amlcompute_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',
max_nodes=6)
compute_target = ComputeTarget.create(ws, amlcompute_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# -
# ## Create the configuration and train a forecaster <a id="train"></a>
# First generate the configuration, in which we:
# * Set metadata columns: target, time column and time-series id column names.
# * Validate our data using cross validation with rolling window method.
# * Set normalized root mean squared error as a metric to select the best model.
# * Set early termination to True, so the iterations through the models will stop when no improvements in accuracy score will be made.
# * Set limitations on the length of experiment run to 15 minutes.
# * Finally, we set the task to be forecasting.
# * We apply the lag lead operator to the target value i.e. we use the previous values as a predictor for the future ones.
# * [Optional] Forecast frequency parameter (freq) represents the period with which the forecast is desired, for example, daily, weekly, yearly, etc. Use this parameter for the correction of time series containing irregular data points or for padding of short time series. The frequency needs to be a pandas offset alias. Please refer to [pandas documentation](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#dateoffset-objects) for more information.
from azureml.automl.core.forecasting_parameters import ForecastingParameters
lags = [1,2,3]
forecast_horizon = n_test_periods
forecasting_parameters = ForecastingParameters(
time_column_name=TIME_COLUMN_NAME,
forecast_horizon=forecast_horizon,
time_series_id_column_names=[ TIME_SERIES_ID_COLUMN_NAME ],
target_lags=lags
)
# Run the model selection and training process. Validation errors and current status will be shown when setting `show_output=True` and the execution will be synchronous.
# +
from azureml.core.workspace import Workspace
from azureml.core.experiment import Experiment
from azureml.train.automl import AutoMLConfig
automl_config = AutoMLConfig(task='forecasting',
debug_log='automl_forecasting_function.log',
primary_metric='normalized_root_mean_squared_error',
experiment_timeout_hours=0.25,
enable_early_stopping=True,
training_data=train_data,
compute_target=compute_target,
n_cross_validations=3,
verbosity = logging.INFO,
max_concurrent_iterations=4,
max_cores_per_iteration=-1,
label_column_name=target_label,
forecasting_parameters=forecasting_parameters)
remote_run = experiment.submit(automl_config, show_output=False)
# -
remote_run.wait_for_completion()
# Retrieve the best model to use it further.
_, fitted_model = remote_run.get_output()
# ## Forecasting from the trained model <a id="forecasting"></a>
# In this section we will review the `forecast` interface for two main scenarios: forecasting right after the training data, and the more complex interface for forecasting when there is a gap (in the time sense) between training and testing data.
# ### X_train is directly followed by the X_test
#
# Let's first consider the case when the prediction period immediately follows the training data. This is typical in scenarios where we have the time to retrain the model every time we wish to forecast. Forecasts that are made on daily and slower cadence typically fall into this category. Retraining the model every time benefits the accuracy because the most recent data is often the most informative.
#
# 
#
# We use `X_test` as a **forecast request** to generate the predictions.
# #### Typical path: X_test is known, forecast all upcoming periods
# +
# The data set contains hourly data, the training set ends at 01/02/2000 at 05:00
# These are predictions we are asking the model to make (does not contain thet target column y),
# for 6 periods beginning with 2000-01-02 06:00, which immediately follows the training data
X_test
# +
y_pred_no_gap, xy_nogap = fitted_model.forecast(X_test)
# xy_nogap contains the predictions in the _automl_target_col column.
# Those same numbers are output in y_pred_no_gap
xy_nogap
# -
# #### Confidence intervals
# Forecasting model may be used for the prediction of forecasting intervals by running ```forecast_quantiles()```.
# This method accepts the same parameters as forecast().
quantiles = fitted_model.forecast_quantiles(X_test)
quantiles
# #### Distribution forecasts
#
# Often the figure of interest is not just the point prediction, but the prediction at some quantile of the distribution.
# This arises when the forecast is used to control some kind of inventory, for example of grocery items or virtual machines for a cloud service. In such case, the control point is usually something like "we want the item to be in stock and not run out 99% of the time". This is called a "service level". Here is how you get quantile forecasts.
# +
# specify which quantiles you would like
fitted_model.quantiles = [0.01, 0.5, 0.95]
# use forecast_quantiles function, not the forecast() one
y_pred_quantiles = fitted_model.forecast_quantiles(X_test)
# quantile forecasts returned in a Dataframe along with the time and time series id columns
y_pred_quantiles
# -
# #### Destination-date forecast: "just do something"
#
# In some scenarios, the X_test is not known. The forecast is likely to be weak, because it is missing contemporaneous predictors, which we will need to impute. If you still wish to predict forward under the assumption that the last known values will be carried forward, you can forecast out to "destination date". The destination date still needs to fit within the forecast horizon from training.
# +
# We will take the destination date as a last date in the test set.
dest = max(X_test[TIME_COLUMN_NAME])
y_pred_dest, xy_dest = fitted_model.forecast(forecast_destination=dest)
# This form also shows how we imputed the predictors which were not given. (Not so well! Use with caution!)
xy_dest
# -
# ## Forecasting away from training data <a id="forecasting_away"></a>
#
# Suppose we trained a model, some time passed, and now we want to apply the model without re-training. If the model "looks back" -- uses previous values of the target -- then we somehow need to provide those values to the model.
#
# 
#
# The notion of forecast origin comes into play: the forecast origin is **the last period for which we have seen the target value**. This applies per time-series, so each time-series can have a different forecast origin.
#
# The part of data before the forecast origin is the **prediction context**. To provide the context values the model needs when it looks back, we pass definite values in `y_test` (aligned with corresponding times in `X_test`).
# +
# generate the same kind of test data we trained on,
# but now make the train set much longer, so that the test set will be in the future
X_context, y_context, X_away, y_away = get_timeseries(train_len=42, # train data was 30 steps long
test_len=4,
time_column_name=TIME_COLUMN_NAME,
target_column_name=TARGET_COLUMN_NAME,
time_series_id_column_name=TIME_SERIES_ID_COLUMN_NAME,
time_series_number=2)
# end of the data we trained on
print(X_train.groupby(TIME_SERIES_ID_COLUMN_NAME)[TIME_COLUMN_NAME].max())
# start of the data we want to predict on
print(X_away.groupby(TIME_SERIES_ID_COLUMN_NAME)[TIME_COLUMN_NAME].min())
# -
# There is a gap of 12 hours between end of training and beginning of `X_away`. (It looks like 13 because all timestamps point to the start of the one hour periods.) Using only `X_away` will fail without adding context data for the model to consume.
try:
y_pred_away, xy_away = fitted_model.forecast(X_away)
xy_away
except Exception as e:
print(e)
# How should we read that eror message? The forecast origin is at the last time the model saw an actual value of `y` (the target). That was at the end of the training data! The model is attempting to forecast from the end of training data. But the requested forecast periods are past the forecast horizon. We need to provide a define `y` value to establish the forecast origin.
#
# We will use this helper function to take the required amount of context from the data preceding the testing data. It's definition is intentionally simplified to keep the idea in the clear.
def make_forecasting_query(fulldata, time_column_name, target_column_name, forecast_origin, horizon, lookback):
"""
This function will take the full dataset, and create the query
to predict all values of the time series from the `forecast_origin`
forward for the next `horizon` horizons. Context from previous
`lookback` periods will be included.
fulldata: pandas.DataFrame a time series dataset. Needs to contain X and y.
time_column_name: string which column (must be in fulldata) is the time axis
target_column_name: string which column (must be in fulldata) is to be forecast
forecast_origin: datetime type the last time we (pretend to) have target values
horizon: timedelta how far forward, in time units (not periods)
lookback: timedelta how far back does the model look?
Example:
```
forecast_origin = pd.to_datetime('2012-09-01') + pd.DateOffset(days=5) # forecast 5 days after end of training
print(forecast_origin)
X_query, y_query = make_forecasting_query(data,
forecast_origin = forecast_origin,
horizon = pd.DateOffset(days=7), # 7 days into the future
lookback = pd.DateOffset(days=1), # model has lag 1 period (day)
)
```
"""
X_past = fulldata[ (fulldata[ time_column_name ] > forecast_origin - lookback) &
(fulldata[ time_column_name ] <= forecast_origin)
]
X_future = fulldata[ (fulldata[ time_column_name ] > forecast_origin) &
(fulldata[ time_column_name ] <= forecast_origin + horizon)
]
y_past = X_past.pop(target_column_name).values.astype(np.float)
y_future = X_future.pop(target_column_name).values.astype(np.float)
# Now take y_future and turn it into question marks
y_query = y_future.copy().astype(np.float) # because sometimes life hands you an int
y_query.fill(np.NaN)
print("X_past is " + str(X_past.shape) + " - shaped")
print("X_future is " + str(X_future.shape) + " - shaped")
print("y_past is " + str(y_past.shape) + " - shaped")
print("y_query is " + str(y_query.shape) + " - shaped")
X_pred = pd.concat([X_past, X_future])
y_pred = np.concatenate([y_past, y_query])
return X_pred, y_pred
# Let's see where the context data ends - it ends, by construction, just before the testing data starts.
print(X_context.groupby(TIME_SERIES_ID_COLUMN_NAME)[TIME_COLUMN_NAME].agg(['min','max','count']))
print(X_away.groupby(TIME_SERIES_ID_COLUMN_NAME)[TIME_COLUMN_NAME].agg(['min','max','count']))
X_context.tail(5)
# +
# Since the length of the lookback is 3,
# we need to add 3 periods from the context to the request
# so that the model has the data it needs
# Put the X and y back together for a while.
# They like each other and it makes them happy.
X_context[TARGET_COLUMN_NAME] = y_context
X_away[TARGET_COLUMN_NAME] = y_away
fulldata = pd.concat([X_context, X_away])
# forecast origin is the last point of data, which is one 1-hr period before test
forecast_origin = X_away[TIME_COLUMN_NAME].min() - pd.DateOffset(hours=1)
# it is indeed the last point of the context
assert forecast_origin == X_context[TIME_COLUMN_NAME].max()
print("Forecast origin: " + str(forecast_origin))
# the model uses lags and rolling windows to look back in time
n_lookback_periods = max(lags)
lookback = pd.DateOffset(hours=n_lookback_periods)
horizon = pd.DateOffset(hours=forecast_horizon)
# now make the forecast query from context (refer to figure)
X_pred, y_pred = make_forecasting_query(fulldata, TIME_COLUMN_NAME, TARGET_COLUMN_NAME,
forecast_origin, horizon, lookback)
# show the forecast request aligned
X_show = X_pred.copy()
X_show[TARGET_COLUMN_NAME] = y_pred
X_show
# -
# Note that the forecast origin is at 17:00 for both time-series, and periods from 18:00 are to be forecast.
# +
# Now everything works
y_pred_away, xy_away = fitted_model.forecast(X_pred, y_pred)
# show the forecast aligned
X_show = xy_away.reset_index()
# without the generated features
X_show[['date', 'time_series_id', 'ext_predictor', '_automl_target_col']]
# prediction is in _automl_target_col
# -
# ## Forecasting farther than the forecast horizon <a id="recursive forecasting"></a>
# When the forecast destination, or the latest date in the prediction data frame, is farther into the future than the specified forecast horizon, the `forecast()` function will still make point predictions out to the later date using a recursive operation mode. Internally, the method recursively applies the regular forecaster to generate context so that we can forecast further into the future.
#
# To illustrate the use-case and operation of recursive forecasting, we'll consider an example with a single time-series where the forecasting period directly follows the training period and is twice as long as the forecasting horizon given at training time.
#
# 
#
# Internally, we apply the forecaster in an iterative manner and finish the forecast task in two interations. In the first iteration, we apply the forecaster and get the prediction for the first forecast-horizon periods (y_pred1). In the second iteraction, y_pred1 is used as the context to produce the prediction for the next forecast-horizon periods (y_pred2). The combination of (y_pred1 and y_pred2) gives the results for the total forecast periods.
#
# A caveat: forecast accuracy will likely be worse the farther we predict into the future since errors are compounded with recursive application of the forecaster.
#
# 
# 
# +
# generate the same kind of test data we trained on, but with a single time-series and test period twice as long
# as the forecast_horizon.
_, _, X_test_long, y_test_long = get_timeseries(train_len=n_train_periods,
test_len=forecast_horizon*2,
time_column_name=TIME_COLUMN_NAME,
target_column_name=TARGET_COLUMN_NAME,
time_series_id_column_name=TIME_SERIES_ID_COLUMN_NAME,
time_series_number=1)
print(X_test_long.groupby(TIME_SERIES_ID_COLUMN_NAME)[TIME_COLUMN_NAME].min())
print(X_test_long.groupby(TIME_SERIES_ID_COLUMN_NAME)[TIME_COLUMN_NAME].max())
# -
# forecast() function will invoke the recursive forecast method internally.
y_pred_long, X_trans_long = fitted_model.forecast(X_test_long)
y_pred_long
# What forecast() function does in this case is equivalent to iterating it twice over the test set as the following.
y_pred1, _ = fitted_model.forecast(X_test_long[:forecast_horizon])
y_pred_all, _ = fitted_model.forecast(X_test_long, np.concatenate((y_pred1, np.full(forecast_horizon, np.nan))))
np.array_equal(y_pred_all, y_pred_long)
# #### Confidence interval and distributional forecasts
# AutoML cannot currently estimate forecast errors beyond the forecast horizon set during training, so the `forecast_quantiles()` function will return missing values for quantiles not equal to 0.5 beyond the forecast horizon.
fitted_model.forecast_quantiles(X_test_long)
# Similarly with the simple senarios illustrated above, forecasting farther than the forecast horizon in other senarios like 'multiple time-series', 'Destination-date forecast', and 'forecast away from the training data' are also automatically handled by the `forecast()` function.
|
how-to-use-azureml/automated-machine-learning/forecasting-forecast-function/auto-ml-forecasting-function.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 🖼️ Diagrams
# ## PlantUML (iPython)
import iplantuml
# +
# %%plantuml --jar --name sample_database
@startuml
' hide the spot
hide circle
' avoid problems with angled crows feet
skinparam linetype ortho
skinparam packageStyle rectangle
package Tables {
entity "Feed" as feed {
*id : number <<generated>>
--
*name : text
*type : number <<FK>>
*uri : text
update : datetime
}
entity "Feed Type" as feedtype {
*id : number <<generated>>
--
*name : text
*specification : text
*schema : text
update : datetime
}
entity "User Feed" as userfeeds {
*id : number <<generated>>
--
*feed_id : number <<FK>>
*user_id : number <<FK>>
*order_seq : number
update : datetime
}
entity "User" as users {
*id : number <<generated>>
--
*name : text
*email : text
update : datetime
}
entity "User Schedule" as usersched {
*id : number <<generated>>
--
*user_id : number <<FK>>
*active : boolean
*cronsched : text
*timezone : text
update : datetime
}
}
package "Config Flatfile" as flatfile{
entity "Ingest File" as ingest #lightgreen {
*filename : text
--
*user name : text
*user email : text
*user cronsched : text
*user timezone : text
*user feeds : obj
}
}
feedtype ||..o{ feed
userfeeds }o..o{ feed
userfeeds }o..|| users
usersched ||..|| users
@enduml
# +
# %%plantuml --jar --name sample_deploy
@startuml
file "Config File" as configfile
interface "FastAPI" as fastapi
rectangle "NiFi" as nifi
rectangle "NiFi Registry" as nifireg
rectangle "HWX Schema Registry" as schemareg
queue "Kafka" as kafka
database "MySQL" as mysql
database "MS SQL" as mssql
cloud "Internet" as internet
configfile --> fastapi
fastapi <--> nifi
fastapi <--> internet
nifi <--> nifireg
nifi <--> kafka
nifi <--> internet
schemareg --> nifi
schemareg --> kafka
mssql --> kafka
mysql <--> nifi
@enduml
# -
# ## Graphviz
# +
# referece: https://github.com/xflr6/graphviz/blob/master/examples/graphviz-notebook.ipynb
# http://www.graphviz.org/pdf/dotguide.pdf Figure 20
from graphviz import Digraph
g = Digraph("G")
g.attr(compound="true")
with g.subgraph(name="cluster0") as c:
c.edges(["ab", "ac", "bd", "cd"])
with g.subgraph(name="cluster1") as c:
c.edges(["eg", "ef"])
g.edge("b", "f", lhead="cluster1")
g.edge("d", "e")
g.edge("c", "g", ltail="cluster0", lhead="cluster1")
g.edge("c", "e", ltail="cluster0")
g.edge("d", "h")
g
# -
# ## mingrammer/diagrams
# +
# reference: https://diagrams.mingrammer.com/docs/getting-started/examples
from diagrams import Cluster, Diagram
from diagrams.aws.compute import ECS, EKS, Lambda
from diagrams.aws.database import Redshift
from diagrams.aws.integration import SQS
from diagrams.aws.storage import S3
with Diagram("Event Processing", show=False) as diag:
source = EKS("k8s source")
with Cluster("Event Flows"):
with Cluster("Event Workers"):
workers = [ECS("worker1"), ECS("worker2"), ECS("worker3")]
queue = SQS("event queue")
with Cluster("Processing"):
handlers = [Lambda("proc1"), Lambda("proc2"), Lambda("proc3")]
store = S3("events store")
dw = Redshift("analytics")
source >> workers >> queue >> handlers
handlers >> store
handlers >> dw
diag
# -
# ## Kroki (Remote)
# +
import base64
import sys
import zlib
import requests
from IPython.display import SVG
kroki_host = "https://kroki.io"
# -
puml = b"""
start
:Hello world;
:This is defined on
several **lines**;
stop
"""
encoded_diag = base64.urlsafe_b64encode(zlib.compress(puml, 9))
encoded_diag.decode("utf-8")
result = requests.get(
"{}/plantuml/svg/{}".format(kroki_host, encoded_diag.decode("utf-8"))
)
result
SVG(result.content)
erd = """[Person] {bgcolor: "#ececfc"}
*name
height
weight
+birth_location_id
[Location] {bgcolor: "#d0e0d0"}
*id
city
state
country
Person *--1 Location
"""
data = {
"diagram_source": erd,
"diagram_type": "erd",
"output_format": "svg",
}
result = requests.post(kroki_host, json=data)
SVG(result.content)
# ## Kroki (Local JAR)
# +
import base64
import os
import subprocess
import sys
import zlib
import psutil
import requests
from IPython.display import SVG
kroki_jar = (
"https://github.com/yuzutech/kroki/releases/download/v0.7.1/kroki-server-0.7.1.jar"
)
kroki_host = "http://localhost:8000"
# -
# download the jar if not already available
kroki_jar_filepath = "{}/{}".format(os.getcwd(), os.path.basename(kroki_jar))
if not os.path.isfile(kroki_jar_filepath):
result = requests.get(kroki_jar)
with open(kroki_jar_filepath, "wb") as f:
f.write(result.content)
# run the jar
kroki_proc = subprocess.Popen(["java", "-jar", kroki_jar_filepath])
print("kroki host process started with pid: {}".format(kroki_proc.pid))
print("kroki pid exists: {}".format(psutil.pid_exists(kroki_proc.pid)))
data = {
"diagram_source": "Bob -> Alice : hello",
"diagram_type": "plantuml",
"output_format": "svg",
}
result = requests.post(kroki_host, json=data)
SVG(result.content)
# kill the kroki subprocess
proc_killed = kroki_proc.kill()
print("kroki pid exists: {}".format(psutil.pid_exists(kroki_proc.pid)))
|
diagrams.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# + id="E934hrzgTSEB"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import datetime
import matplotlib as mpl
import matplotlib.pyplot as plt
# + [markdown] id="hIcYQojwTSEE"
# # Ayiti Analytics Probability and Statistics Project
# + [markdown] id="4q2hjm_ZTSEH"
# Use all the files to have a final datasets to have the following columns
# <ul>
# <li>questid</li>
# <li>gender</li>
# <li>age (age of applicant)</li>
# <li>communeName (use data prep in commune file)</li>
# <li>application_date (created at in quest file)</li>
# <li>enroll_date (created_at in enroll file)</li>
# <li>is_enroll (Yes/No) (use data prep in enroll file)</li>
# <li>Education Level</li>
# <li>Communication channels(hear_AA1)</li>
# <li>Bootcamp Insterest (after_AA)</li>
# <li>Payement Date (use ord and transaction files)</li>
# <li>Payed (Yes/No)</li>
# <li>list Technologies as columns based (use get_dummies)</li>
# <li>list Study domains (use get_dummies)</li>
# <li>Job is formal</li>
# <li>Have computer at home</li>
# <li>Have internet at home</li>
#
# </ul>
#
# + id="-njqkLFsgImS"
commune=pd.read_excel(r"commune.xlsx")
quest= pd.read_csv(r"quest.csv")
enroll= pd.read_csv(r"enroll.csv")
industry=pd.read_csv(r"industry.csv")
ord= pd.read_csv(r"ord.csv")
study_domain=pd.read_csv(r"study_domain.csv")
technology=pd.read_csv(r"technology.csv")
transaction= pd.read_csv(r"transaction.csv")
# + id="C0ik6NSgT5ju"
# def read_file(r="", file=""):
# if file.type()==excel:
# df= pd.read_excel(file)
# else:
# df=pd.read_csv(file)
# def read_file(file_name=""):
# read= pd.read_csv(r"file_name")
# return read
# + colab={"base_uri": "https://localhost:8080/"} id="IWa5f-giX1_H" outputId="8b8a189d-85e4-4c75-b3cc-8f2e62dd5f52"
# read_file(file_name="quest.csv")
len(quest)
# + id="oMLQ3wjttIbt"
def merging (y,x):
df= pd.merge(left=x, right=y, how="left", on="quest_id" )
display(df.head(2))
return df
# -
def dummies (data):
dum= pd.get_dummies(data[["key", "quest_id","values"]], columns=["values"], prefix="", prefix_sep="")
groupe= dum.groupby("quest_id").sum()
return groupe
techno= dummies(technology)
study= dummies(study_domain)
# + id="igfstI-pcCem"
enroll["is_enroll"]="yes"
# -
df4=pd.concat([ord,transaction],axis=0)
df4["payed"]="yes"
df4.head(2)
df4
df4.rename(columns={"created_at":"Payment_Date"}, inplace=True)
df4.head(2)
df4["Payment_Date"]
df4=df4.loc[:,["Payment_Date","quest_id","payed"]]
df4.head(2)
# + colab={"base_uri": "https://localhost:8080/", "height": 440} id="CUTs-fHewyTk" outputId="fa60a972-2cae-4cd9-c753-5849c80bb869"
df1= merging(x=quest,y=enroll)
# -
df1.head(2)
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="im4Q9OOLKbbN" outputId="d68b1105-f47b-41ec-b2d0-229c1f392cf7"
df1.rename(columns={"created_at_x":"application_date","created_at_y":"enroll_date"},inplace=True)
df1.head(2)
# + id="XOeq_jYkeo5k"
df1["is_enroll"] = df1["is_enroll"].fillna("No")
# + colab={"base_uri": "https://localhost:8080/"} id="EhXs4ea0SC2i" outputId="ddcc08cf-898f-4625-d91b-1875d136e95d"
df1.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 222} id="Lg_Iu589yJ51" outputId="125dfbab-5ea6-4d53-e2ac-60300ef0ad40"
df1=df1.loc[:,["gender","dob","application_date","education_level","formal_sector_job","have_computer_home","internet_at_home","hear_AA_1","after_AA","quest_id","enroll_date","commune","is_enroll"]]
df1.head(2)
# + colab={"base_uri": "https://localhost:8080/", "height": 105} id="TOm3LkEx5qyD" outputId="76defcef-6c33-45c2-95ca-51f34d8aceab"
commune["Commune_Id"] = commune["Commune_Id"].str.lower()
commune.rename (columns={"Commune_Id":"commune"},inplace=True)
#commune["ADM1_PCODE"] = commune["ADM1_PCODE"].str.lower()
commune.head(2)
#commune = commune.loc[:,["Commune_FR"]]
# + colab={"base_uri": "https://localhost:8080/"} id="8DgBY3nQ59Fn" outputId="40e5fae9-0b35-4cc3-d23d-39ccbbab16cd"
df2= pd.merge(left=df1, right=commune[["commune","Commune_FR"]], how="left", on="commune")
df2.shape
# -
techno
df3=merging(x=df2, y= techno)
df3.shape
df5= merging(x=df3 , y=study )
# + id="Guj8aFmMpCmi"
df5.shape
# -
df4.shape
# + id="hUFKkyOmoeGK"
df_final=merging(x=df5,y=df4)
# -
df_final["payed"]
df_final["payed"]=df_final["payed"].fillna("No")
df_final["payed"]
df_final=df_final.rename(columns={"hear_AA_1":"Communication_channels","after_AA":"Bootcamp_Insterest","dob":"Birth_Date"})
# +
# times=pd.Timestamp(year=2021, month=3, day=20)
# +
# df_final["Birth_Date"]= df_final["Birth_Date"].fillna(times)
# -
df_final["Birth_Date"].replace({'3 aout 1977': '03/08/1977'}, inplace=True)
df_final["Birth_Date"].isna().value_counts()
df_final["application_date"] = df_final["application_date"].apply(lambda x: str(x)[:10])
df_final["Birth_Date"] = pd.to_datetime(df_final["Birth_Date"])
df_final["application_date"] = pd.to_datetime(df_final["application_date"])
df_final["Age"]=df_final["application_date"]-df_final["Birth_Date"]
df_final["Age"].value_counts()
df_final["Age"] = df_final["Age"]// np.timedelta64(1,"Y")
df_final["Age"].mean()
# +
# df_final["Age"]=df_final["Age"]//365
# +
# df_final["Age"] = df_final["Age"].apply(lambda x: str(x)[:2])
# -
df_final.columns
df_final.Age.value_counts()
df_final.Age=df_final.Age.replace({-1.00000:np.nan,1.00000:np.nan,0.00000:np.nan})
df_final.Age= df_final.Age.fillna(df_final.Age.mean())
df_final["Age"] = df_final["Age"].apply(lambda x: int(x))
df_final.Age
df_final.Age.value_counts()
# + [markdown] id="ehC_9a1Z2Zc_"
#
# + [markdown] id="zJ<KEY>"
#
# + [markdown] id="hlpCdKUSTSEK"
# ### 1. How many observations and variables are there in the dataset
#
# + id="7lZAWdkITSEM"
# here your codes
a=df_final.shape
print(f"The number of observations are: {a[0]} and the number of variables are {a[1]}")
# + [markdown] id="-FP7KXrNTSEO"
# ### 2.A Calculate the average age of the observations
# * On the whole dataset
# * On the whole male dataset
# * On all the female dataset
# ### 2.B Calculate
# * variance of the whole datatset,male and female
# * kurtosis of the whole datatset,male and female
# * skewness of the whole datatset,male and female
# ### 2.C Does the age distribution follow a normal distribution
#
# -
#2A Average age of the observations
# on the whole dataset
df_final["Age"].mean()
# on the whole male dataset
male_avg= df_final[df_final["gender"] == "male"]
male_avg.Age.mean()
female_avg= df_final[df_final["gender"] == "female"]
female_avg.Age.mean()
# 2.B variance on the whole dataset
df_final["Age"].var()
# Variance on the female dataset age
female_avg.Age.var()
#variance on the male dataset age
male_avg.Age.var()
# kurtosis on the male age
a=(male_avg.Age)
a.kurtosis()
# kurtosis on the female
female_avg.Age.kurtosis()
# Kurtosis on the whole dataset
df_final["Age"].kurtosis()
# !pip install scipy
import scipy as stats
from scipy.stats import skew
# +
#skewness on the whole age dataset
skew(df_final.Age)
# -
# skweness on the male dataset
skew(male_avg.Age)
# skewness on the female dataset
skew(female_avg.Age)
figure=df_final.groupby('gender').Age.plot(kind='kde')
# +
# #generate_barchart(data=gender_result,title="Total et Percent By Sex",abs_value="Total",rel_value="Percent")
# figure_1 = df_final.Age.plot(kind="bar")
# # Get a Matplotlib figure from the axes object for formatting purposes
# fig = figure_1.get_figure()
# # Change the plot dimensions (width, height)
# fig.set_size_inches(6,6)
# # Change the axes labels
# # figure_1.set_xlabel("hear_AA_1",fontsize=12)
# # figure_1.set_ylabel("gender",fontsize=12)
# # figure_1.set_title("Distribution of women according to the channel used",fontsize=12)
# # to put a font size to the graph
# plt.rcParams.update({'font.size': 10})
# #add legend
# figure_1.legend()
# plt.tight_layout()
# #plt.show()
# figure_1
# -
bins = np.linspace(df_final.Age.min(),df_final.Age.max(),15,endpoint=True)
plt.figure(figsize=(10,6))
sns.histplot(data =df_final,x=df_final["Age"],kde=True,bins=bins,hue="gender")
plt.title("Density")
plt.show()
# Does the age distribution follow a normal distribution
import scipy.stats as stats
# normal test on the whole dataset
stats.jarque_bera(df_final.Age)
# +
# normal test on the female dataset
female_avg= df_final[df_final["gender"] == "female"]
stats.jarque_bera(female_avg.Age)
# -
# normal test on the male dataset
stats.jarque_bera(male_avg.Age)
# + [markdown] id="LKBJSmbATSES"
# ### 3.Display the age frequency disbrution
# * On the whole dataset
# * On the whole male dataset
# * On all the female dataset
# -
# Age frequency distribution
# on the whole dataset
display(df_final.Age.value_counts(normalize=True).to_frame())
# +
# display((df_final["gender"]=="female").Age.value_counts(normalize=True).to_frame())
# -
def pivo(data="y"):
gender_pivot=pd.pivot_table(df_final, values='quest_id', index=['Age'], columns=data,aggfunc='count')
return gender_pivot
display((df_final[df_final["gender"]=="male"]).Age.value_counts(normalize=True).to_frame())
# on the female dataset
display((df_final[df_final["gender"]=="female"]).Age.value_counts(normalize=True).to_frame())
# + [markdown] id="qtzZrZ6MTSEW"
# ### 4. Can we say that the average age (24 years old) of the observations can be considered as the average age of the population likely to participate in this bootcamp. Justify your answer
# -
# + [markdown] id="ROlYpGTBTSEa"
# ### 5. Calculate the average age of participants for each communication channel
# -
age_channel = pd.pivot_table(df_final, index= "Communication_channels", values='Age', aggfunc='mean')
age_channel
# # pivo=pd.pivot_table(df_final, values='quest_id', index=['Communication_channels'], columns="Age",aggfunc='mean')
# # pivo
# age_channel =df_final.groupby("Communication_channels")["Age"].nunique().to_frame()
# age_channel
# +
# df_final.pivot_table(index="Communication_channels", columns="Age", values=["quest_id"], aggfunc=np.mean, fill_value=0)
# + [markdown] id="N5iUnz4kTSEe"
# ### 6. Display an age boxplot for each communication channel
# -
fig = df_final.boxplot(column=['Age'], by=['Communication_channels'], figsize=(18,10), fontsize=10)
plt.figure(figsize=(10,6))
sns.boxplot(data=df_final,x="Communication_channels",y="Age")
# + [markdown] id="CqffjuvuTSEh"
# ### 7 .Is there a significant age difference between these groups
# -
age_channel =df_final.groupby("Communication_channels")["Age"].count().to_frame()
age_channel.reset_index()
# + id="mquCtkRATSEi"
#here your codes
#anova test
# fvalue, pvalue = stats.f_oneway(age_channel["Communication_channels"], age_channel["Age"] )
# print(fvalue, pvalue)
# + [markdown] id="Zf2Z2T1dTSEk"
# ### 8.Plan a BA strategy for each communication channel regarding the age group
# + id="0Irx-WL7TSEl"
#here your codes
# + [markdown] id="VRiNTp0OTSEn"
# ### 9.According to the observations what is the probability of being in each channel of communication knowing your are a woman
# -
def prob_all_comchannels(donne="x"):
probability_1= df_final[df_final.Communication_channels==donne].shape[0]/df_final.shape[0]
return probability_1
def prob_knowing_female(y,z):
prob_boot= (y*z)/z
return prob_boot
# + id="KfbbCD5ETSEo"
#here your codes
prob_friend= prob_all_comchannels(donne="Friend")
prob_whatssap= prob_all_comchannels(donne="WhatsApp")
prob_facebook= prob_all_comchannels(donne="Facebook")
prob_esih= prob_all_comchannels(donne="ESIH")
prob_ayiti_analitics= prob_all_comchannels(donne="Ayiti Analytics Website")
prob_linkedin= prob_all_comchannels(donne="LinkedIn")
prob_alumni= prob_all_comchannels(donne="Bootcamp Alumni")
prob_other_com= prob_all_comchannels(donne="other")
prob_instagram= prob_all_comchannels(donne="Instagram")
prob_unknown_com= prob_all_comchannels(donne="unknown")
prob_female= df_final[df_final.gender=='female'].shape[0]/df_final.shape[0]
prob_male= df_final[df_final.gender=='male'].shape[0]/df_final.shape[0]
# -
prob_friend_female=prob_knowing_female(y=prob_friend,z=prob_female)
prob_whatssap_female=prob_knowing_female(y=prob_whatssap,z=prob_female)
prob_facebook_female =prob_knowing_female(y=prob_facebook,z=prob_female)
prob_esih_female =prob_knowing_female(y=prob_esih,z=prob_female)
prob_ayiti_female =prob_knowing_female(y=prob_ayiti_analitics,z=prob_female)
prob_linkedin_female =prob_knowing_female(y=prob_linkedin,z=prob_female)
prob_alumni_female =prob_knowing_female(y=prob_alumni,z=prob_female)
prob_other_female =prob_knowing_female(y=prob_other_com,z=prob_female)
prob_instagram_female =prob_knowing_female(y=prob_instagram,z=prob_female)
prob_unknown_female =prob_knowing_female(y=prob_unknown_com,z=prob_female)
# + [markdown] id="wChpmw9gTSEp"
# ### 10.According to the observations what is the probability of being in each channel of communication knowing your are a man
# + id="1O7UFuMCTSEq"
#here your codes
prob_friend_male=prob_knowing_female(y=prob_friend,z=prob_male)
prob_friend_male
# -
prob_whatssap_male=prob_knowing_female(y=prob_whatssap,z=prob_male)
prob_facebook_male =prob_knowing_female(y=prob_facebook,z=prob_male)
prob_esih_male =prob_knowing_female(y=prob_esih,z=prob_male)
prob_ayiti_male =prob_knowing_female(y=prob_ayiti_analitics,z=prob_male)
prob_linkedin_male =prob_knowing_female(y=prob_linkedin,z=prob_male)
prob_alumni_male =prob_knowing_female(y=prob_alumni,z=prob_male)
prob_other_male =prob_knowing_female(y=prob_other_com,z=prob_male)
prob_instagram_male =prob_knowing_female(y=prob_instagram,z=prob_male)
prob_unknown_male =prob_knowing_female(y=prob_unknown_com,z=prob_male)
# + [markdown] id="5y0RxWx3TSEr"
# ### 11. Deduce the probability of being a man knowing each communication channel
# + id="Ao4HbLVjTSEs"
#here your codes
prob_male_unknown=prob_knowing_female(y=prob_male,z=prob_unknown_com)
# -
prob_male_instagram=prob_knowing_female(y=prob_male,z=prob_instagram)
prob_male_other=prob_knowing_female(y=prob_male,z=prob_other_com)
prob_male_alumni=prob_knowing_female(y=prob_male,z=prob_alumni)
prob_male_linkedin =prob_knowing_female(y=prob_male,z=prob_linkedin)
prob_male_ayiti =prob_knowing_female(y=prob_male,z=prob_ayiti_analitics)
prob_male_esih =prob_knowing_female(y=prob_male,z=prob_esih)
prob_male_facebook =prob_knowing_female(y=prob_male,z=prob_facebook)
prob_male_whatsapp =prob_knowing_female(y=prob_male,z=prob_whatssap)
# + [markdown] id="wGkJ_GrkTSEt"
# ### 13. Deduce the probability of being a woman knowing each communication channel
# + id="BWP4fk0LTSEu"
# here your codes
prob_female_whatsapp =prob_knowing_female(y=prob_female,z=prob_whatssap)
# -
prob_female_facebook =prob_knowing_female(y=prob_female,z=prob_facebook)
prob_female_esih =prob_knowing_female(y=prob_female,z=prob_esih)
prob_female_ayiti =prob_knowing_female(y=prob_female,z=prob_ayiti_analitics)
prob_female_linkedin =prob_knowing_female(y=prob_female,z=prob_linkedin)
prob_female_alumni =prob_knowing_female(y=prob_female,z=prob_alumni)
prob_female_other =prob_knowing_female(y=prob_female,z=prob_other_com)
prob_female_instagram =prob_knowing_female(y=prob_female,z=prob_instagram)
prob_female_unknown =prob_knowing_female(y=prob_female,z=prob_unknown_com)
# + [markdown] id="NalU9FkTTSEy"
# ### 15 Display a plot to see Gender vs Communication Channels .Is there any dependency between communication channels and gender?
# -
import seaborn as sns
cross1=pd.crosstab(index=df_final.gender,columns=df_final.Communication_channels)
cross1.plot.bar(figsize=(12,10), rot=0)
from scipy.stats import chi2_contingency
chi2_contingency(cross1.values)
stat, p, dof, expected = chi2_contingency(cross1.values)
alpha = 0.05
print("p value is " + str(p))
if p <= alpha:
print('The two variables are dependent (reject Null Hypothesis H0)')
else:
print('The two variables are independent (Null Hypothesis H0 holds true)')
# + [markdown] id="9f5e18m_TSE0"
# ### 16 Use the same method to display plot and know if is there any dependency between communication channels and Bootcamp Insterest?
# + id="0nP4EPNXTSE1"
# here you
cross2=pd.crosstab(index=df_final.Communication_channels,columns=df_final.Bootcamp_Insterest)
cross2.plot.bar(figsize=(12,10), rot=0)
# -
from scipy.stats import chi2_contingency
chi2_contingency(cross2.values)
stat, p, dof, expected = chi2_contingency(cross2.values)
alpha = 0.05
print("p value is " + str(p))
if p <= alpha:
print('The two variables are dependent (reject Null Hypothesis H0)')
else:
print('The two variables are independent (Null Hypothesis H0 holds true)')
# + [markdown] id="iVjj8TXfTSE2"
# ### 17.Plan a BA strategy for each communication channel, Bootcamp Insterest regarding the gender
# + id="ImytFuNcTSE3"
#here your codes
# + [markdown] id="0NdV7YBXTSE3"
# ### 18.Plan a BA strategy for each communication channel, Bootcamp Insterest regarding the gender
# + id="PM2izr9nTSE4"
# here your codes
# + [markdown] id="F9QXFvT4TSE5"
# ### 19. Calculate
# * P(Bootcamp Insterest,Communication channels,Gender/Payed=yes)
# * P(Bootcamp Insterest,Communication channels,Gender/Payed=no)
# -
df_final["Bootcamp_Insterest"].value_counts()
# calcul of all the probability in the bootcamp interest columns
def prob_all_bootcamp_interest(data="x"):
probability_0= df_final[df_final.Bootcamp_Insterest==data].shape[0]/df_final.shape[0]
return probability_0
prob_improving_data_skills= prob_all_bootcamp_interest(data="Improving my data analysis skills")
# Finding a job probability
prob_finding_job = prob_all_bootcamp_interest(data="Finding a job/internship")
prob_start_company=prob_all_bootcamp_interest(data="Training to start my own company")
prob_switch_carrer=prob_all_bootcamp_interest(data="Switching careers")
prob_Mentorship=prob_all_bootcamp_interest(data="Mentorship")
prob_other=prob_all_bootcamp_interest(data="other")
prob_unknow=prob_all_bootcamp_interest(data="unknown")
df_final.Communication_channels.value_counts()
#calcul of the communication channels probability
def prob_all_comchannels(donne="x"):
probability_1= df_final[df_final.Communication_channels==donne].shape[0]/df_final.shape[0]
return probability_1
prob_friend= prob_all_comchannels(donne="Friend")
prob_whatssap= prob_all_comchannels(donne="WhatsApp")
prob_facebook= prob_all_comchannels(donne="Facebook")
prob_esih= prob_all_comchannels(donne="ESIH")
prob_ayiti_analitics= prob_all_comchannels(donne="Ayiti Analytics Website")
prob_linkedin= prob_all_comchannels(donne="LinkedIn")
prob_alumni= prob_all_comchannels(donne="Bootcamp Alumni")
prob_other_com= prob_all_comchannels(donne="other")
prob_instagram= prob_all_comchannels(donne="Instagram")
prob_unknown_com= prob_all_comchannels(donne="unknown")
# +
# calcul of the probability for gender
# -
prob_male= df_final[df_final.gender=='male'].shape[0]/df_final.shape[0]
prob_male
prob_female= df_final[df_final.gender=='female'].shape[0]/df_final.shape[0]
prob_female
# +
# calcul of all the probability for the payed columns
# -
prob_payed_yes= df_final[df_final.payed=="yes"].shape[0]/df_final.shape[0]
prob_payed_yes
prob_payed_no= df_final[df_final.payed=="No"].shape[0]/df_final.shape[0]
prob_payed_no
# +
#* P(Bootcamp Insterest,Communication channels,Gender/Payed=yes)
# -
def prob_knowing_payed_yes(y,z):
prob_boot= (y*z)/z
return prob_boot
prob_improve_yes=prob_knowing_payed_yes(y=prob_improving_data_skills,z=prob_payed_yes)
prob_job_yes=prob_knowing_payed_yes(y=prob_finding_job,z=prob_payed_yes)
prob_company_yes=prob_knowing_payed_yes(y=prob_start_company,z=prob_payed_yes)
prob_carrer_yes=prob_knowing_payed_yes(y=prob_switch_carrer,z=prob_payed_yes)
prob_mentor_yes=prob_knowing_payed_yes(y=prob_Mentorship,z=prob_payed_yes)
prob_other_yes=prob_knowing_payed_yes(y=prob_other,z=prob_payed_yes)
prob_unknow_yes=prob_knowing_payed_yes(y=prob_unknow,z=prob_payed_yes)
# +
# * P(Communication channels/Payed=no)"""
# -
prob_friend_yes=prob_knowing_payed_yes(y=prob_friend,z=prob_payed_yes)
prob_whatsapp_yes=prob_knowing_payed_yes(y=prob_whatssap,z=prob_payed_yes)
prob_facebook_yes=prob_knowing_payed_yes(y=prob_facebook,z=prob_payed_yes)
prob_ayiti_yes=prob_knowing_payed_yes(y=prob_ayiti_analitics,z=prob_payed_yes)
prob_linkedin_yes=prob_knowing_payed_yes(y=prob_linkedin,z=prob_payed_yes)
prob_alumni_yes=prob_knowing_payed_yes(y=prob_alumni,z=prob_payed_yes)
prob_other_com_yes=prob_knowing_payed_yes(y=prob_other_com,z=prob_payed_yes)
prob_instagram_yes=prob_knowing_payed_yes(y=prob_instagram,z=prob_payed_yes)
prob_unknown_com_yes=prob_knowing_payed_yes(y=prob_unknown_com,z=prob_payed_yes)
# +
#* P(Gender/Payed=yes)
# -
prob_female_yes=prob_knowing_payed_yes(y=prob_female,z=prob_payed_yes)
prob_male_yes=prob_knowing_payed_yes(y=prob_male,z=prob_payed_yes)
# + [markdown] id="Zzz2oWknTSE6"
# ### 20 reduce
# * P(Payed="yes"/Bootcamp Insterest="Increase Skill",Communication channels="Friend",Gender="male")
# * P(Payed="no"/Bootcamp Insterest="Increase Skill",Communication channels="Friend",Gender="male")
# -
# + id="u9BsceOETSE7"
# here your codes
# + [markdown] id="QpMBm7IBTSE8"
# ### Based on these findings, propose strategies to increase our sales?
# + id="KWx7jdBtTSE9"
#here your codes
# + [markdown] id="9lvWxgLhTSE-"
#
|
Statistics_project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import recombination as rb
import timeit
# Please, to run the experiment donwload the following dataset and put it in the /Datasets folder:
# - 3D_spatial_network.txt -
# https://archive.ics.uci.edu/ml/machine-learning-databases/00246/3D_spatial_network.txt
filename = "Datasets/3D_spatial_network.txt"
df = pd.read_csv(filename, sep=',', usecols=[1,2,3])
df = df.dropna()
print(list(df.columns.values) )
# +
df.columns = ['Longitude', 'Latitude', 'Altitude']
print(list(df.columns.values))
df['Longitude'] = pd.to_numeric(df['Longitude'], errors='coerce')
df['Latitude'] = pd.to_numeric(df['Latitude'], errors='coerce')
df['Altitude'] = pd.to_numeric(df['Altitude'], errors='coerce')
df = df.dropna()
print(df.shape)
print(df.dtypes)
df.head()
# -
x = df[['Longitude','Latitude']]
x = x.to_numpy()
y = df['Altitude']
y = y.to_numpy()
n = x.shape[1]
# +
print(x.shape)
X = np.append(x,y[np.newaxis].T,1)
xy_sq = rb.tens_sq(X)
print(xy_sq.shape)
print(xy_sq[:,n+1:].shape)
N, d = xy_sq[:,n+1:].shape
# +
mean_t = 0.
time_rand = []
iterations_rand = []
min_t = np.inf
max_t = 0.
sample = 1000
COV = np.matmul(x.T,x)/N
for i in range(sample):
tic = timeit.default_timer()
w_star, idx_star, _, _, _, iterations, eliminated_points = rb.recomb_Mor_reset(
xy_sq[:,n+1:]-np.mean(xy_sq[:,n+1:],0), 400)
time_rand.append((timeit.default_timer()-tic)*1000)
iterations_rand.append(iterations)
################ CHECK THE BARYCENTER IS THE SAME
COV_recomb = np.zeros(COV.shape)
jj = 0
for j in idx_star:
tmp = np.matmul(x[j,:][np.newaxis].T,x[j,:][np.newaxis])
COV_recomb += tmp * w_star[jj]
jj += 1
assert np.allclose(COV_recomb,COV), "ERROR COV != COV_RECOMB"
################ CHECK FINISHED
mean_t += time_rand[-1]
print("sample = ", i)
print("time = ", time_rand[-1], "ms")
print("mean time = ", mean_t/(i+1), "ms")
min_t = min(time_rand)
max_t = max(time_rand)
print("---------------------------------------")
print("max t = ", max_t, "ms")
print("min t = ", min_t, "ms")
print("mean = ", mean_t/sample, "ms")
print("std = ", np.std(time_rand))
print("---------------------------------------")
# +
mean_t = 0.
sample = 100
time_MT = []
min_t = np.inf
max_t = 0.
COV = np.matmul(x.T,x)/N
for i in range(sample):
x_cp = np.copy(xy_sq[:,n+1:])
tic = timeit.default_timer()
w_star, idx_star, _, _, _, iterations, eliminated_points = rb.Tchernychova_Lyons(
x_cp)
time_MT.append((timeit.default_timer()-tic)*1000)
################ CHECK THE BARYCENTER IS THE SAME
COV_recomb = np.zeros(COV.shape)
jj = 0
for j in idx_star:
tmp = np.matmul(x[j,:][np.newaxis].T,x[j,:][np.newaxis])
COV_recomb += tmp * w_star[jj]
jj += 1
assert np.allclose(COV_recomb,COV), "ERROR COV != COV_RECOMB"
################ CHECK FINISHED
mean_t += time_MT[-1]
print("sample = ", i)
print("time = ", time_MT[-1], "ms")
print("mean time = ", mean_t/(i+1), "ms")
min_t = min(time_MT)
max_t = max(time_MT)
print("---------------------------------------")
print("max t = ", max_t, "ms")
print("min t = ", min_t, "ms")
print("mean = ", mean_t/sample, "ms")
print("std FC = ", np.std(time_MT))
print("---------------------------------------")
# +
from Maalouf_Jubran_Feldman import Fast_Caratheodory
time_FC = []
mean_t = 0.
for i in range(100):
tic = timeit.default_timer()
Fast_Caratheodory(xy_sq[:,n+1:],np.ones(N),d+1)
time_FC.append((timeit.default_timer()-tic)*1000)
mean_t += time_FC[-1]
print("sample = ", i)
print("time = ", time_FC[-1], "ms")
print("mean time = ", mean_t/(i+1), "ms")
print("---------------------------------------")
print("max FC = ", np.max(time_FC), " ms")
print("min FC = ", np.min(time_FC), " ms")
print("mean FC = ", np.mean(time_FC), " ms")
print("std FC = ", np.std(time_FC))
print("---------------------------------------")
# +
mean_t = 0.
sample = 1000
time_log = np.zeros(sample)
min_t = 0.
max_t = 0.
COV = np.matmul(x[:,:].T,x[:,:])/N
for i in range(sample):
x_cp = np.copy(xy_sq[:,n+1:])
tic = timeit.default_timer()
w_star, idx_star, _, _, _, _, _ = rb.recomb_log(x_cp)
time_log[i] = (timeit.default_timer()-tic)*1000
################ CHECK THE BARYCENTER IS THE SAME
COV_recomb = np.zeros(COV.shape)
jj = 0
for j in idx_star:
tmp = np.matmul(x[j,:][np.newaxis].T,x[j,:][np.newaxis])
COV_recomb += tmp * w_star[jj]
jj += 1
assert np.allclose(COV_recomb,COV), "ERROR COV != COV_RECOMB"
################ CHECK FINISHED
mean_t += time_log[i]
print("sample = ", i)
print("time = ", time_log[i], "ms")
print("mean time = ", mean_t/(i+1), "ms")
mean_t = np.mean(time_log)
min_t = np.min(time_log)
max_t = np.max(time_log)
print("---------------------------------------")
print("max t = ", max_t, "ms")
print("min t = ", min_t, "ms")
print("mean = ", mean_t, "ms")
print("---------------------------------------")
# +
time_rand = np.array(time_rand)
iterations_rand = np.array(iterations_rand)
time_FC = np.array(time_FC)
time_log = np.array(time_log)
time_MT = np.array(time_MT)
np.set_printoptions(precision=1)
print("Probability to be faster = ",
np.sum(np.array(time_rand)<np.mean(time_FC))/sample*100, "%")
print("Probability to be 4x faster = ",
np.sum(np.array(time_rand)<np.mean(time_FC)/4)/sample*100, "%")
print("Standard deviation = ", np.std(time_rand))
print("The expected time of the log-random is ", np.mean(time_log), "ms")
print("Standard deviation of the log-random is = ", np.std(time_log))
np.set_printoptions(precision=1)
print('''Some statistics for the randomized algorithm are:
average running time = ''', np.round(np.mean(time_rand),1),
"ms, min = " , np.round(np.min(time_rand),1), "ms, max = ", np.round(np.max(time_rand),1),
"ms, std ", np.round(np.std(time_rand),1),
"ms. Using the log-random strategy they are: average running time = ", np.round(np.mean(time_log),1),
"ms, min = ", np.round(np.min(time_log),1), "ms, max = ", np.round(np.max(time_log),1),
", std = ", np.round(np.std(time_log),1), "ms.",
" Average runnig times of determinsitic: TL = ", np.round(np.mean(time_MT),1),
"ms, MJF = ", np.round(np.mean(time_FC),1),"ms.")
# +
import matplotlib.pyplot as plt
plt.rcParams.update({'font.size': 16})
fig, axs = plt.subplots(5,1,figsize=(10,4.5*5))
################################################
plt.subplot(5, 1, 1)
plt.hist(time_rand, bins=int(90))
plt.axvline(np.mean(time_rand), 0, max(time_rand), linestyle='-', color="blue", label="mean randomized algo")
plt.axvline(np.mean(time_MT), 0, max(time_MT), linestyle='-.', color="orange", label="mean det3")
plt.axvline(np.mean(time_FC), 0, max(time_rand), linestyle=':', color="red", label="mean det4")
plt.xlim((0, max(time_rand)))
plt.legend()
plt.title('Distribution of the running time - 3D Roads')
plt.xlabel('time (ms)')
################################################
plt.subplot(5, 1, 2)
plt.hist(iterations_rand, bins=int(90))
plt.title('Distribution of the iterations')
plt.xlabel('number of iterations')
plt.xscale('linear')
################################################
plt.subplot(5, 1, 3)
plt.plot(iterations_rand,time_rand, '.')
plt.xlabel('iterations')
plt.ylabel('time (ms)')
plt.title('Iterations vs time')
################################################
plt.subplot(5, 1, 4)
plt.hist(time_log, bins=int(40),color='limegreen')
plt.axvline(np.mean(time_rand), linestyle='-', color="blue", label="mean randomized algo")
plt.axvline(np.mean(time_log), linestyle='--', color="green", label="mean log-random algo")
plt.axvline(np.mean(time_MT), linestyle='-.', color="orange", label="mean det3")
plt.axvline(np.mean(time_FC), linestyle=':', color="red", label="mean det4")
plt.xlim((0, max(time_rand)))
plt.legend()
plt.title('Distribution of the running time of the log-random algorithm')
################################################
plt.subplot(5, 1, 5)
plt.hist(time_rand, bins=int(250))
plt.hist(time_log, bins=int(55),color='limegreen')
plt.axvline(np.mean(time_rand), 0, max(time_rand), linestyle='-', color="blue", label="mean randomized algo")
plt.axvline(np.mean(time_log), 0, max(time_log), linestyle='--', color="green", label="mean log-random algo")
plt.axvline(np.mean(time_MT), 0, max(time_MT), linestyle='-.', color="orange", label="mean det3")
plt.axvline(np.mean(time_FC), 0, max(time_rand), linestyle=':', color="red", label="mean det4")
plt.xlim((min(time_rand), max(time_rand)/2.5))
plt.legend()
plt.title('Distribution of the running time of the log-random algorithm')
plt.xlabel('time (ms)')
fig.tight_layout()
# plt.savefig('Distrib_running_time_3DRoads.pdf')#, bbox_inches='tight')
plt.show()
# +
mean_t = 0.
time_combined = []
min_t = np.inf
max_t = 0.
sample = 1000
COV = np.matmul(x.T,x)/N
for i in range(sample):
tic = timeit.default_timer()
w_star, idx_star, _, _, _, iterations, eliminated_points = rb.recomb_combined(
xy_sq[:,n+1:], 400)
time_combined.append((timeit.default_timer()-tic)*1000)
################ CHECK THE BARYCENTER IS THE SAME
COV_recomb = np.zeros(COV.shape)
jj = 0
for j in idx_star:
tmp = np.matmul(x[j,:][np.newaxis].T,x[j,:][np.newaxis])
COV_recomb += tmp * w_star[jj]
jj += 1
assert np.allclose(COV_recomb,COV), "ERROR COV != COV_RECOMB"
################ CHECK FINISHED
mean_t += time_combined[-1]
print("sample = ", i)
print("time = ", time_combined[-1], "ms")
print("mean time = ", mean_t/(i+1), "ms")
min_t = min(time_combined)
max_t = max(time_combined)
print("---------------------------------------")
print("max t = ", max_t, "ms")
print("min t = ", min_t, "ms")
print("mean = ", mean_t/sample, "ms")
print("std = ", np.std(time_combined))
print("---------------------------------------")
# +
maximum = max(np.mean(time_rand),np.mean(time_log),np.mean(time_MT),np.mean(time_rand),np.mean(time_combined))*2
plt.rcParams.update({'font.size': 16})
fig, axs = plt.subplots(figsize=(10,4.5))
plt.hist(time_combined,bins=int(100),color='grey')
plt.axvline(np.mean(time_combined), linestyle='--', color="grey", label="mean combined algo")
plt.axvline(np.mean(time_rand), linestyle='-', color="blue", label="mean randomized algo")
plt.axvline(np.mean(time_log), linestyle='--', color="green", label="mean log-random algo")
plt.axvline(np.mean(time_MT), linestyle='-.', color="orange", label="mean det3")
plt.axvline(np.mean(time_FC), linestyle=':', color="red", label="mean det4")
plt.xlim((0, maximum/1.3))
plt.legend()
plt.title('Distr. running time Combined Algorithm - 3D Roads')
plt.xlabel('time (ms)')
fig.tight_layout()
# plt.savefig('Distrib_running_time_3DRoads_combined.pdf')#, bbox_inches='tight')
plt.show()
# -
|
Comparison_3D_spatial_network.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # %load train_cropped.py
import numpy as np
from tensorflow.python.keras import backend as K
from tensorflow.python.keras.models import Model
from tensorflow.python.keras.layers import Flatten, Dense, Dropout
from tensorflow.python.keras.applications.resnet50 import ResNet50, preprocess_input
from tensorflow.python.keras.optimizers import Adam
from tensorflow.python.keras.preprocessing.image import ImageDataGenerator
DATASET_PATH = './catsdogs/sample'
IMAGE_SIZE = (256, 256)
CROP_LENGTH = 224
NUM_CLASSES = 2
BATCH_SIZE = 8 # try reducing batch size or freeze more layers if your GPU runs out of memory
FREEZE_LAYERS = 2 # freeze the first this many layers for training
NUM_EPOCHS = 20
WEIGHTS_FINAL = 'model-cropped-final.h5'
def random_crop(img, random_crop_size):
# Note: image_data_format is 'channel_last'
assert img.shape[2] == 3
height, width = img.shape[0], img.shape[1]
dy, dx = random_crop_size
x = np.random.randint(0, width - dx + 1)
y = np.random.randint(0, height - dy + 1)
return img[y:(y+dy), x:(x+dx), :]
def crop_generator(batches, crop_length):
"""Take as input a Keras ImageGen (Iterator) and generate random
crops from the image batches generated by the original iterator.
"""
while True:
batch_x, batch_y = next(batches)
batch_crops = np.zeros((batch_x.shape[0], crop_length, crop_length, 3))
for i in range(batch_x.shape[0]):
batch_crops[i] = random_crop(batch_x[i], (crop_length, crop_length))
yield (batch_crops, batch_y)
train_datagen = ImageDataGenerator(preprocessing_function=preprocess_input,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
channel_shift_range=10,
horizontal_flip=True,
fill_mode='nearest')
train_batches = train_datagen.flow_from_directory(DATASET_PATH + '/train',
target_size=IMAGE_SIZE,
interpolation='bicubic',
class_mode='categorical',
shuffle=True,
batch_size=BATCH_SIZE)
valid_datagen = ImageDataGenerator(preprocessing_function=preprocess_input)
valid_batches = valid_datagen.flow_from_directory(DATASET_PATH + '/valid',
target_size=IMAGE_SIZE,
interpolation='bicubic',
class_mode='categorical',
shuffle=False,
batch_size=BATCH_SIZE)
train_crops = crop_generator(train_batches, CROP_LENGTH)
valid_crops = crop_generator(valid_batches, CROP_LENGTH)
# show class indices
print('****************')
for cls, idx in train_batches.class_indices.items():
print('Class #{} = {}'.format(idx, cls))
print('****************')
# build our classifier model based on pre-trained ResNet50:
# 1. we don't include the top (fully connected) layers of ResNet50
# 2. we add a DropOut layer followed by a Dense (fully connected)
# layer which generates softmax class score for each class
# 3. we compile the final model using an Adam optimizer, with a
# low learning rate (since we are 'fine-tuning')
net = ResNet50(include_top=False, weights='imagenet', input_tensor=None,
input_shape=(CROP_LENGTH,CROP_LENGTH,3))
x = net.output
x = Flatten()(x)
x = Dropout(0.5)(x)
output_layer = Dense(NUM_CLASSES, activation='softmax', name='softmax')(x)
net_final = Model(inputs=net.input, outputs=output_layer)
for layer in net_final.layers[:FREEZE_LAYERS]:
layer.trainable = False
for layer in net_final.layers[FREEZE_LAYERS:]:
layer.trainable = True
net_final.compile(optimizer=Adam(lr=1e-5),
loss='categorical_crossentropy', metrics=['accuracy'])
print(net_final.summary())
# train the model
net_final.fit_generator(train_crops,
steps_per_epoch = train_batches.samples // BATCH_SIZE,
validation_data = valid_crops,
validation_steps = valid_batches.samples // BATCH_SIZE,
epochs = NUM_EPOCHS)
# save trained weights
net_final.save(WEIGHTS_FINAL)
# -
|
train_cropped.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Ref: https://github.com/Lancelotl/Gale-Shapley/blob/master/stable_matching.py
import random
from typing import List, Dict, Tuple
# Declaring types
Person = str
People = List[Person]
Preferences = List[Person]
Side = Dict[Person, Preferences]
Participants = Dict[str, Side]
Pair = Tuple[Person, Person]
Matching = Dict[Person, Person]
Stable_Matching = List[Pair]
class MissingPreferences(Exception):
pass
def other_side(current_side: str, all_sides: list) -> str:
"""Given a side and all possible sides, returns the opposite side"""
for s in all_sides:
if s != current_side:
return s
def all_preferences(participants: Participants) -> bool:
"""Checks whether all participants have all participants of the other side in their own preference"""
sides = list(participants.keys())
for side, people in participants.items():
other_side_participants = participants[other_side(side, sides)]
for name, preferences in people.items():
for o in other_side_participants:
if o not in preferences:
return False
else:
return True
def is_free(person: Person, engaged: Matching) -> bool:
"""Is the person missing from all current pairs?"""
return person not in engaged
def current_match(person: Person, engaged: Matching) -> bool:
"""Returns the current match for that person"""
return engaged.get(person)
def free_participants(people: People, engaged: Matching) -> list:
"""Returns all participants are that are still currently free"""
return filter(lambda x: x not in engaged, people)
def preferred(a: Person, b: Person, preferences: Preferences) -> Person:
"""Is a preferred over b according to the preferences ordering?"""
for preference in preferences:
if preference == a:
return True
if preference == b:
return False
def stable_matching(participants: Participants) -> Stable_Matching:
"""For a group of participants and their respective preferences of the other group, returns a list of stable matches according to the Gale–Shapley algorithm"""
# The algorithm requires each participant expresses a preference that includes all other participants
if not all_preferences(participants):
raise MissingPreferences
sides = list(participants.keys())
proposing = sides[0] # Taking the 1st side
receiving = sides[1] # Taking the 2nd side
proposers = participants[proposing]
receivers = participants[receiving]
free_proposers = proposers
proposal_history = {k: {} for k in proposers}
engagements = {}
while free_proposers:
for proposer in free_proposers:
preferences = proposers[proposer]
for target in preferences:
# Has proposed yet?
if target not in proposal_history[proposer]:
# Record proposal
proposal_history[proposer][target] = ""
# Is receiver free?
if is_free(target, engagements):
# Engagement
engagements[proposer] = target
engagements[target] = proposer
else:
# Pair already exists
current = current_match(target, engagements)
target_preferences = receivers[target]
if preferred(proposer, current, target_preferences):
# Proposer replaces the current individual
engagements[target] = proposer
engagements[proposer] = target
# Freeing the incumbent
del engagements[current]
# Done proposing this round
break
# Updating the list of proposers that are free
# Must be a list since a generator always evaluates to True
free_proposers = list(free_participants(free_proposers, engagements))
# Composing the stable matchings
stable_matchings = set()
for a, b in engagements.items():
# Checking the reverse isn't already in
if (b, a) not in stable_matchings:
stable_matchings.add((a, b))
return list(stable_matchings)
if __name__ == "__main__":
listA = ["A1", "A2", "A3", "A4", "A5", "A6", "A7", "A8"]
listB = ["B1", "B2", "B3", "B4", "B5", "B6", "B7", "B8"]
results = []
count = 0
for i in range(1000):
sample_participants = {
"Super_Group_A": {
"A1": random.sample(listB,8),
"A2": random.sample(listB,8),
"A3": random.sample(listB,8),
"A4": random.sample(listB,8),
"A5": random.sample(listB,8),
"A6": random.sample(listB,8),
"A7": random.sample(listB,8),
"A8": random.sample(listB,8)
},
"Super_Group_B": {
"B1": random.sample(listA,8),
"B2": random.sample(listA,8),
"B3": random.sample(listA,8),
"B4": random.sample(listA,8),
"B5": random.sample(listA,8),
"B6": random.sample(listA,8),
"B7": random.sample(listA,8),
"B8": random.sample(listA,8)
}
}
result = stable_matching(sample_participants)
results.append(result)
for result in results:
if result is not results[-1]:
count += 1
p = count/1000
print(p)
|
Assignment1-Q9-B.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from fbprophet import *
# +
hadoop(자바 기반 오픈소스 프레임워크)
MR-MapReduce-HDFS에서 데이터를 mp로 코딩했으나 지금은 spark을 사용
YARN-리소스 메니저
HDFS-대용량 파일을 분산된 서버에 저장하고, 그 저장된 데이터를 빠르게 처리할 수 있게 하는 파일시스템
MapReduce란? : 대용량 데이터를 처리를 위한 분산 프로그래밍 모델 타고난 병행성(병렬 처리 지원)을 내포,
누구든지 임의로 활용할 수 있는 충분한 서버를 이용하여 대규모 데이터 분석 가능 - 흩어져 있는 데이터를 수직화하여,
그 데이터를 각각의 종류 별로 모으고(Map), Filtering과 Sorting을 거쳐 데이터를 뽑아내는(Reduce) 하는 분산처리 기술과 관련 프레임워크를 의미
# -
배울것
batch-일괄처리
stream-지속적인 데이터
- HDFS 설치방식 : Mode 3가지
-------------------------------------------------------------------------------
- Stand alone (독립실행모드) : 기본 실행모드.... 분산저장 안함.... 코딩은 가능....
- Pseudo-distributed (가상분산모드) : 하나의 컴퓨터에 설치해 여러 클러스터가 하나의 서버에서 돌아가는 것
- Fully distributed (완전분산모드) : 여러 대의 컴퓨터에 설치....
# +
hadoop-env.sh(환경)설정
vi /kikang/hadoop3/etc/hadoop/hadoop-env.sh
master namenode = 각 분산한 데이터들의 디렉토리를 관리하는 마스터 네임노드
resource manager = 네임노드의 디렉토리를 관리
slave(worker) datanode = 각 분산된 데이터들이 저장되어 있는 곳
nodemanager= 각 데이터 노드의 데이터만 관리
seonderdary namen node= 백업용
# +
설정파일들
hadoop: core-site.xml(설정파일) ha
hadoop:env.sh(환경설정,실행파일): 실행을 하면 env에 설정되어 있는 설정이 구성되고 hadoop 실행됨
hdfs(store)
hdfs-site.xml
namenode, datanode, secondary namenode
mr(mapreduce)=>spark
narored-site
:
# -
리눅스 : bin 명령어들 모임
|
Spark.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venti
# language: python
# name: venti
# ---
import torch
import numpy as np
torch.ones(5,4)
torch.tensor(torch.finfo(torch.float32).min)
tmp_actions = torch.rand(5,4,3)
tmp_inv = torch.rand(5,3,1)
out = torch.matmul(tmp_actions, tmp_inv)
out.shape
tmp = torch.rand(5,4)
torch.cat([tmp, tmp], dim=0)
tmp[0:0+1,:]
torch.rand(5,4).numpy()
torch.rand(2,).unsqueeze(dim=1)
torch.rand(5,4)[0:2, 0:2]
torch.randint(low=0, high=9, size=(5,4)).numpy().dtype
torch.randint(low=0, high=9, size=(9,))
torch.cat([
torch.randint(low=0, high=9, size=(9,)).unsqueeze(0),
torch.randint(low=0, high=9, size=(9,)).unsqueeze(0)
], dim=0)
torch.randint(low=0, high=9, size=(9,)).unsqueeze(0)
import torch
torch.embedding.__file__
|
tests/deprecated/test-tensors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BitPanda
# ## Imports and Params
# +
from Ionomy import BitPanda
from decouple import config
MARKET = 'btc-hive'
CURRENCY = 'hive'
BASE = 'btc'
TIME = 'day'
# -
# ## Instantiation
bp = BitPanda(config('TREX_KEY'), config('TREX_SECRET'))
# ## Public Endpoint Methods
# #### Markets
markets_pd = bp.markets()
markets_pd.head()
# #### Currencies
currencies_pd = bp.currencies()
currencies_pd.head()
# ### Ticker
bp.ticker(MARKET)
# #### Market Summaries
market_summaries_pd = bp.market_summaries()
market_summaries_pd.head()
# #### Market Summary
bp.market_summary(MARKET)
# #### Order Book
order_book_pd = bt.order_book(MARKET)
order_book_pd.head()
# #### Market History
market_history_pd = bp.market_history(MARKET)
market_history_pd.head()
# ## Market Endpoint Methods
# #### Limit Buy/Sell Order
order_uuid = bp.buy_limit(MARKET, QUANTITY, RATE, TIMEINFORCE)
order_uuid = bp.sell_limit(MARKET, QUANTITY, RATE, TIMEINFORCE)
# #### Cancel Order
order_uuid = bp.cancel(UUID)
# #### Order Status
order = bp.get_order(UUID)
# ## Account Endpoint Methods
# #### Balances
balances_pd = bp.balances()
balances_pd.head()
# #### Balance
bp.balance(CURRENCY)
# #### Order History
order_history_pd = bp.order_history()
order_history_pd.head()
# #### Deposit History
deposit_history_pd = bp.deposit_history(CURRENCY)
deposit_history_pd.head()
# #### Deposit History
bp.deposit_address(CURRENCY)
# #### Withdrawal History
withdrawal_history_pd = bp.withdrawal_history(CURRENCY)
withdrawal_history_pd.head()
# #### Withdraw
uuid = bt.withdraw(CURRENCY, QUANTITY, ADDRESS, PAYMENTID_OPTIONAL)
|
docs/source/notebooks/bitpanda.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Text processing from the command line
#
# It's often the case that we can do a huge amount of cleanup on unstructured text before using Python to process it more formally. We can delete unwanted characters, squeeze repeated characters, reformat, etc... In this section you will do a number of exercises that get you used to processing files from the command line. If you'd like to dig further, you can see [this link](http://www.tldp.org/LDP/abs/html/textproc.html).
#
# The operating system launches all commands in a pipeline sequence as separate processes, which means they can run on multiple processors simultaneously. This gives us parallel processing without having to write complicated code. As data is completed by one stage, it passes it to the next stage of the pipeline, and continues to work on its input. The next stage consumes that input in parallel. Consequently, processing text from the command line can be extremely efficient, much more so than doing it in Python.
#
# Let's get the Tesla text again:
# ! curl https://www.tesla.com/sites/default/files/tesla-model-s.pdf > /tmp/tsla.pdf
# That command downloads the file and because of the redirection operator, `>`, the output gets written to `tsla.pdf` up in `/tmp` directory.
#
# Once we have the data, we can pass the filename to `pdftotext` to extract the text:
# ! pdftotext /tmp/tsla.pdf # saves into /tmp/tsla.txt
# ### Exercise
#
# Using the `tr` (translate) command from the terminal, strip all of the new lines from the text file you created above (`/tmp/tsla.txt`). Look at the manual page with this command:
#
# ```python
# $ man tr
# ```
#
# You can pipe the output of `tr` to `head -c 200` to only print out the first 200 characters of the output.
# ! tr -s '\n' ' ' < /tmp/tsla.txt | head -c 200
# ### Exercise
#
# Reformat the text using `tr` and `fold`. The `fold` command wraps lines at 80 characters; use its `-s` option to making break lines at spaces between words.
# ! tr -s '\n' ' ' < /tmp/tsla.txt | fold -s | head -10
# ### Exercise
#
# It is sometimes useful to put a line number at the left edge of all lines. For example, you might want to create a unique ID number for each row of a CSV file. Pipe the output of the previous command to `nl` so that you get the line number on the left edge.
# ! tr -s '\n' ' ' < /tmp/tsla.txt | fold -s | nl | head -10
# ### Exercise
#
# Convert the text to all lowercase using `tr`. Hint: `a-z` and `A-Z` are [regular expressions](http://www.rexegg.com/regex-quickstart.html) that describe English characters and uppercase English characters.
# ! tr 'A-Z' 'a-z' < /tmp/tsla.txt | head -c 150
# ### Exercise
#
# Do the same thing but on the text that has the new lines removed.
# ! tr -s '\n' ' ' < /tmp/tsla.txt | tr 'A-Z' 'a-z' | head -c 150
# ### Exercise
#
# Get a histogram of all words in the document. Start by converting all spaces to newlines so that all words are unaligned by themselves. Then pipe that output to `tr` to make everything lowercase as we did before. Next `sort` that output so you see a list of all the words in sorted order. Once they are sorted, we can use `uniq -c` to count unique instances. Finally, we can do a reverse sort numerically on the word counts that are in the first column with `sort -r -n`.
# ! tr -s ' ' '\n' < /tmp/tsla.txt | tr 'A-Z' 'a-z' | sort | head -10
# ! tr -s ' ' '\n' < /tmp/tsla.txt | tr 'A-Z' 'a-z' | \
# sort | uniq -c | head -10
# ! tr -s ' ' '\n' < /tmp/tsla.txt | tr 'A-Z' 'a-z' | \
# sort | uniq -c | sort -r -n | head -20
|
notes/text-cmd-line.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %matplotlib nbagg
import datetime
from collections import defaultdict
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# +
dp_end_dates = [datetime.datetime(2022, 6, 1), # DP0.1
datetime.datetime(2023, 6, 1), # DP0.2
datetime.datetime(2024, 3, 1), # DP1
datetime.datetime(2025, 3, 1), # DP2
datetime.datetime(2025, 9, 1), # DR1
]
one_year = datetime.timedelta(365)
# -
ops_allocations = {'Computing Environment': 0.6,
'Data Verification & Validation': 0.3,
'Databases and data access': 0.4,
'Image processing pipeline development': 0.75,
'Image processing pipeline management': 0.2,
'Image simulation development': 1.4,
'Image simulation pipeline development': 0,
'Image simulation pipeline management': 0,
'Science Release management': 0.35,
'Software Release management': 0.15
}
def get_date(date):
tokens = date.split('/')
year = (int(tokens[-1]) % 2000) + 2000
month = int(tokens[0])
day = int(tokens[1]) if len(tokens) == 3 else 1
return datetime.datetime(year, month, day)
def read_sheet(csv_file):
data = defaultdict(list)
with open(csv_file) as fobj:
for line in fobj:
tokens = line.split(',')
if (len(tokens[0].split('.')) != 3 #check for a deliverable entry
or tokens[5] == '' # skip rows without baseline completion date
or tokens[14] == '' # skip rows without Operations role
):
continue
data['deliverable'].append(tokens[2])
start = get_date(tokens[4])
end = get_date(tokens[5])
data['start'].append(start)
data['end'].append(end)
# Assume each PS/CI effort level entry is in FTE-years.
data['FTE-years'].append(float(tokens[11]))
# Convert to FTE (as a rate) by dividing by the fraction of a year.
frac = (end - start)/one_year
data['FTE'].append(float(tokens[11])/frac)
data['ops_role'].append(tokens[14])
return pd.DataFrame(data=data)
df0 = read_sheet('WBS_Dictionary_Computing.csv')
ops_roles = set(df0['ops_role'])
plt.figure(figsize=(8, 12))
for i, ops_role in enumerate(ops_roles, 1):
if ops_role == '':
continue
plt.subplot(4, 2, i)
df = df0.query(f'ops_role == "{ops_role}"')
edges = np.array(sorted(list(set(df['start']).union(df['end']))))
mid_points = edges[:-1] + (edges[1:] - edges[:-1])/2.
bin_values = np.zeros(len(mid_points))
for start, end, fte, fte_years in zip(df['start'], df['end'], df['FTE'], df['FTE-years']):
for i, mid_point in enumerate(mid_points):
if start < mid_point < end:
bin_values[i] += fte
plt.stairs(bin_values, edges, label=ops_role)
for dp_end_date in dp_end_dates:
plt.axvline(dp_end_date, linestyle='--', color='green', alpha=0.3)
plt.axhline(ops_allocations[ops_role], linestyle=':', color='green', alpha=0.3)
plt.xticks([datetime.datetime(year, 1, 1) for year in range(2022, 2026, 1)], fontsize='small')
plt.yticks(fontsize='small')
plt.title(ops_role)
plt.ylabel('FTE')
plt.xlabel('date')
plt.tight_layout()
|
notebooks/WBS_Ops_FTE_plots.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import all the necessary libraries
import numpy as np
import pandas as pd
import pickle
from rdkit.Chem import AllChem
from rdkit.Chem.rdMolDescriptors import GetMACCSKeysFingerprint
from rdkit import DataStructs
# # Processing the data to be predicted
# Below codes demonstrate how to process the smiles strings in an xlsx file. You can download the file above.
## Load the file
df = pd.read_excel("Aropha_AB_regression_predict_example.xlsx", sheet_name='Sheet1')
df.head()
## Convert smiles to MACCS molecular fingerprint (the model we will be using was built based on MACCS fingerprints)
df['mol'] = [AllChem.MolFromSmiles(smiles) for smiles in df['SMILES']]
df['fp'] = [GetMACCSKeysFingerprint(mol) for mol in df['mol']]
df = pd.concat([df['fp'], df['Guideline'], df['Principle'], df['Reliability']], axis=1)
df.head()
## Mannually encode the categorical data
cat_dict_guideline = {'EU Method C.4-C': 0, 'EU Method C.4-D': 1, 'EU Method C.4-E': 2,
'OECD 301B': 3, 'OECD 301C': 4, 'OECD 301D': 5, 'OECD 301F': 6,
'OECD 310': 7}
cat_dict_principle = {'CO2 evolution': 0, 'Closed respirometer': 1, 'Closed bottle test': 2}
df = df.replace({'Guideline': cat_dict_guideline, 'Principle': cat_dict_principle})
df.head()
## Obtain the final X_input for the model
X = []
X_fp = np.array(df.iloc[:, 0])
X_other = np.array(df.iloc[:, 1:4])
for i in range(len(df)):
record_fp = np.array(X_fp[i]).tolist()
other = np.array(X_other[i]).tolist()
for item in other:
record_fp.append(item) ## Append each categorical data into fp
X.append(record_fp)
X = np.array(X)
X
# # Load the model and perform the prediction
## Load the model (you can download this model use the link above)
model = pickle.load(open('Aropha_AB_XGBRegressor_model.pkl', 'rb'))
## Perform the prediction and save the results to a column named "Prediction" in the orginal dataframe
prediction = model.predict(X)
# # Calculate the prediction performance
# The prediction performance is based on the similarity between the query compound and the dataset used to build the model.
## Load the data that was used to build the model. It can be downloaded in the "Dataset" tab
model_data = pd.read_excel('Aropha_AB_regression_model_data.xlsx', sheet_name='Sheet1')
model_mols = [AllChem.MolFromSmiles(smiles) for smiles in model_data['Smiles']]
model_fp = [GetMACCSKeysFingerprint(mol) for mol in model_mols]
'''The prediction performance is based on the similarity score.
For example, during the model development, chemicals with a similarity score of >=0.9 with each other
demonstrated an R2 or 0.79 and RMSE of 0.14 between the predicted and true values.'''
def prediction_acc(similarity):
if similarity >= 0.9:
R2 = 0.79
RMSE = 0.14
elif 0.8 <= similarity <= 0.9:
R2 = 0.66
RMSE = 0.21
elif 0.7 <= similarity <= 0.8:
R2 = 0.59
RMSE = 0.23
elif 0.6 <= similarity <= 0.7:
R2 = 0.44
RMSE = 0.26
else:
R2 = 'Out of AD'
RMSE = 'Out of AD'
return R2, RMSE
similarity_list = []
R2_list = []
RMSE_list = []
for fp in df['fp']:
similarities = DataStructs.BulkTanimotoSimilarity(fp, model_fp) ## Compare the query compound with all the model data
similarities.sort()
similarity = round(similarities[-1], 2)
R2, RMSE = prediction_acc(similarity)
similarity_list.append(similarity)
R2_list.append(R2)
RMSE_list.append(RMSE)
## Add the similarity and accuracy scores to the dataframe
df_0 = pd.read_excel("Aropha_AB_regression_predict_example.xlsx", sheet_name='Sheet1')
df_0['Prediction'] = ['{:.1%}'.format(i) for i in prediction]
df_0['Similarity'] = similarity_list
df_0['Expected prediction R2'] = R2_list
df_0['Expected prediction RMSE'] = RMSE_list
df_0.head()
# # Save the results to a csv file
df_0.to_csv("prediction_result.csv")
|
example/Aropha_AB_regression_jupyter_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A2: Bias in the Data
# <NAME> <br>
# In this notebook,
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# ## Step 0: Background and EDA
# In this session, I will walk you through some visualizations and distributions for the demographic information of crowdflower workers who labeled toxicity and aggression. Those simple EDA helped me to think about the bias in data problem that I want to solve.
# Load the data related to toxicity and aggression
# +
# data for toxicity
demographic_worker = pd.read_csv('toxicity_data/toxicity_worker_demographics.tsv', sep = '\t')
annotated_comments = pd.read_csv('toxicity_data/toxicity_annotated_comments.tsv', sep = '\t')
annotations = pd.read_csv('toxicity_data/toxicity_annotations.tsv', sep = '\t')
# data for aggression
demographic_worker_agg = pd.read_csv('~/Desktop/aggression_data/aggression_worker_demographics.tsv', sep = '\t')
annotated_comments_agg = pd.read_csv('~/Desktop/aggression_data/aggression_annotated_comments.tsv', sep = '\t')
annotations_agg = pd.read_csv('~/Desktop/aggression_data/aggression_annotations.tsv', sep = '\t')
# -
# Show percentage of the gender distribution of the workers who labeled toxicity
demographic_worker.gender.value_counts('female')
# Show percentage of the gender distribution of the workers who labeled aggression
demographic_worker_agg.gender.value_counts('female')
# Show the count of workers in each gender for toxicity
sns.set(rc={'figure.figsize':(10,6)})
sns.countplot(x="gender", hue="gender", data=demographic_worker)
# Show percentage of the age group distribution of the workers for toxicity through visualization
df= demographic_worker.age_group.value_counts('age_group')*100
worker_age_df = pd.DataFrame({'age_group':df.index, 'proportion':df.values})
df
sns.set(rc={'figure.figsize':(10,6)})
sns.barplot(x="age_group", y = "proportion", hue="age_group", data=worker_age_df,
hue_order = ['Under 18','18-30', '30-45', '45-60', 'Over 60'],
order = ['Under 18','18-30', '30-45', '45-60', 'Over 60'])
# Show percentage of the age-bucket distribution of the workers who labeled aggression
agg_df= demographic_worker_agg.age_group.value_counts('age_group')*100
worker_age_df_agg = pd.DataFrame({'age_group':agg_df.index, 'proportion':agg_df.values})
agg_df
# ## Step 1: Analysis
# There are two anlysis that I would like to do. First, I want to furthur analyze the demographic information about the Crowdflower workers that is available in the dataset and answering the questions as the follwoing: How well does the demographic profile of the crowdworkers match that of the general population? What are some potential consequences of skewed labeller demographic distribution on the behavior of a model trained on this dataset?
#
# The second analysis I want to do is to explore relationships between worker demographics and labeling behavior. I would like to answer the questions as the following: Are female-identified labelers more or less likely to label comments as aggressive than male-identified labelers? If the labelling behaviors are different, what are some possible causes and consequences of this difference?
#
# I will be using both the toxicity data and the aggression data for my analysis here.
# ### Analysis 1
# In the first analysis, I would like to compare the demographic information about the crowflower workers who labeled the toxicity data and aggression data with the demographic information of the general population. Here are the overview of the steps I took:
#
# - Find the gender distribution data and the age distribution information of the general population from UN's data
# - Choose the year 2015 since the age and gender distribution stays fairly constant in a range of years
# - Perform data manipulation to compare the demographic information of the general public with the crowflowers'
# Load the age distribution data download from UN's website
xls = pd.ExcelFile('PopulationAgeSex-20201019065126.xlsx')
population_age_df = pd.read_excel(xls, 'Data')
# Perform data manipulation to be able to match with the age-bucket in our crowdflower workers' data.
# +
population_age_df = population_age_df.iloc[[13]]
population_age_df = population_age_df.drop(columns=['ISO 3166-1 numeric code', 'Location', 'Time', 'Sex'])
population_age_df = pd.melt(population_age_df, var_name='age_group', value_name='population')
# define new age group for the UN data
age_group_population = ['Under 18', 'Under 18', 'Under 18', 'Under 18',
'18-30', '18-30', '30-45', '30-45', '30-45',
'45-60', '45-60','45-60',
'Over 60', 'Over 60', 'Over 60', 'Over 60', 'Over 60', 'Over 60', 'Over 60', 'Over 60', 'Over 60']
population_age_df['age_group_population'] = age_group_population
population_age_df
# -
# drop the unused column
new_population_age_df = population_age_df.drop(columns = ['age_group'])
# calculate the distribution of propotion of each age group
world_demographic = new_population_age_df.groupby('age_group_population').agg('sum')['population']/new_population_age_df.sum()['population']*100
world_demographic
# reformat the poportion into a dataframe
world_demographic = pd.DataFrame({'age_group':world_demographic.index, 'world_proportion':world_demographic.values})
world_demographic
# We can now compare the distribution of age group of workers who labeled toxicity vs the general population
worker_vs_world_age_df = world_demographic.merge(worker_age_df, on = 'age_group')
worker_vs_world_age_df
# We now repeat the same procedure for comparing the distribution of age group of workers who labeled aggression vs the general population
agg_worker_vs_world_age_df = world_demographic.merge(worker_age_df_agg, on = 'age_group')
agg_worker_vs_world_age_df
# ### Analysis 1 findings
# From UN's data, I find the demographic distribution for age-group and gender for the year 2015. I did some data manipulation for the age-group to match up with the age-group in the dataset. For gender information (https://population.un.org/wpp/DataQuery/), male vs female ratio is 101.7 : 100 which is distributed evenly. Compared to the demographic info for the crowdflower workers, we see a very uneven distribution across gender and age-group population distribution. Workers who labeled for toxicity data and aggression data shows the same skewed demographic information. The demographic profile of the crowdflower workers do not match that of the general population well at all. We see that there are twice as much male crowdflower workers than females. We also see that there are a lot more younger people over 18-year-old among workers than the proportion of this age group in the general population. Such bias in the demographic data of the workers is very obvious.
# We should pay attention to such bias in the data since it is possible that male vs female or young people vs old people have different level of sentiment when they label the wiki comments on a scale of -2 to 2 for both the toxicity and aggression datasets. Thus in our second analysis, we go furthur to analysis suh bias in the demographic profile of the crodflower workers.
# ## Analysis 2
# In this analysis, I will furthur explore the bias in the demographic profile of workers. I want to answer the question: are female-identified labelers more or less likely to label comments as aggressive than male-identified labelers? In order to answer this question, I followed the steps below:
#
# - Merge annotations for toxicity with the corresponding worker demographic data and do the same thing for aggression data
# - Calculate the conditional probability using Bayes' theorem for P(aggression level given female workers) and P(aggression level given male workers)
# - Do the same thing for the toxicity data
#
# #### Aggression
# Merge annotations for aggression with the corresponding worker demographic data
annotations_agg_demo = annotations_agg.merge(demographic_worker_agg, on = 'worker_id')
annotations_agg_demo
# Find the porprotion of female and male who give aggression score < 0 for aggression data
annotations_agg_new = annotations_agg_demo[annotations_agg_demo['aggression_score'] < 0]
annotations_agg_new.gender.value_counts('female')
# Calculate the joint probability for female/male and aggression score < 0
print('P(aggression < 0 and gender = female): {}'.format(len(annotations_agg_new) * 0.386576 / len(annotations_agg_demo)))
print('P(aggression < 0 and gender = male): {}'.format(len(annotations_agg_new) * 0.613188 / len(annotations_agg_demo)))
# Find the porprotion of female and male who give extreme aggression score = -2
annotations_agg_neg2 = annotations_agg_demo[annotations_agg_demo['aggression_score'] == -2]
annotations_agg_neg2.gender.value_counts('female')
print('P(aggression = -2 and gender = female): {}'.format(len(annotations_agg_neg2) * 0.364790 / len(annotations_agg_demo)))
print('P(aggression = -2 and gender = male): {}'.format(len(annotations_agg_neg2) * 0.634996 / len(annotations_agg_demo)))
# Below is what we calculated before for p(female) worker in the aggression dataset.
annotations_agg_demo.gender.value_counts('female')
# We use the Bayes' theorem to figure how the condition probability: P(aggression_score < 0 | gender = female) and P(aggression_score < 0 | gender = male). <br>
#
#
# \begin{equation*}
# P(aggression < 0 | gender = female/male) =
# \frac{P(aggression < 0 \cap gender = female/male)}{P(gender = female/male)}
# \end{equation*} <br>
# \begin{equation*}
# P(aggression < 0 | gender = female) =
# \frac{0.0708369}{0.360138} = 0.19667
# \end{equation*}
# \begin{equation*}
# P(aggression < 0 | gender = male) =
# \frac{0.1123617}{0.639765} = 0.1756
# \end{equation*}
# We calculate the conditional probability also for the extreme aggressive scores
# \begin{equation*}
# P(aggression = -2 | gender = female) =
# \frac{0.016}{0.360138} = 0.044
# \end{equation*}
# \begin{equation*}
# P(aggression = -2 | gender = male) =
# \frac{0.028}{0.639765} = 0.044
# \end{equation*}
# #### Toxicity
# We now do the same thing for the toxicity data
annotations_demo = annotations.merge(demographic_worker, on = 'worker_id')
# Find the porprotion of female and male who give toxicty score < 0
annotations_new = annotations_demo[annotations_demo['toxicity_score'] < 0]
annotations_new.gender.value_counts('female')
# Calculate the joint probability for female/male and toxicity score < 0
print('P(toxicity < 0 and gender = female): {}'.format(len(annotations_new) * 0.366003 / len(annotations_demo)))
print('P(toxicity < 0 and gender = male): {}'.format(len(annotations_new) * 0.633697 / len(annotations_demo)))
# Find the porprotion of female and male who give extreme toxicity score = -2
annotations_neg2 = annotations_demo[annotations_demo['toxicity_score'] == -2]
annotations_neg2.gender.value_counts('female')
print('P(toxicity = -2 and gender = female): {}'.format(len(annotations_neg2) * 0.391500 / len(annotations_demo)))
print('P(toxicity = -2 and gender = male): {}'.format(len(annotations_neg2) * 0.608083 / len(annotations_demo)))
# Below is what we calculated before for p(female) worker in the toxicity dataset.
annotations_demo.gender.value_counts('female')
# We use the Bayes' theorem to figure how the condition probability: P(toxicity_score < 0 | gender = female) and P(toxicity_score < 0 | gender = male). <br>
#
#
# \begin{equation*}
# P(toxicity < 0 | gender = female/male) =
# \frac{P(toxicity < 0 \cap gender = female/male)}{P(gender = female/male)}
# \end{equation*} <br>
# \begin{equation*}
# P(toxicity < 0 | gender = female) =
# \frac{0.053}{0.339541} = 0.156
# \end{equation*}
# \begin{equation*}
# P(toxicity < 0 | gender = male) =
# \frac{0.092}{0.660194} = 0.139
# \end{equation*}
# \begin{equation*}
# P(toxicity = -2 | gender = female) =
# \frac{0.01045}{0.339541} = 0.031
# \end{equation*}
# \begin{equation*}
# P(toxicity = -2 | gender = male) =
# \frac{0.016}{0.660194} = 0.024
# \end{equation*}
# ### Analysis 2 findings
# Under the assumption that workers are randomly selected to do the annotations for toxicity data and the aggression data, we see that suprising, female-identified labelers are equally likely to label comments as aggressive than male-identified labelers. Based on the results of the conditional probability we have, we see P(toxicity_score/aggression_score < 0 | gender = female) and P(toxicity_score/aggression < 0 | gender = male) are very similar to each other. When I compared the extreme comments with aggression_score = -2 and toxicty_score = -2, we still see no difference between female-identified labelers and male identified labelers. P(toxicity_score/aggression = -2 | gender = female) and P(toxicity_score/aggression_score = -2 | gender = male) are very similar.
# I conclude that female-identified labelers are equally likely to label comments as aggressive than male-identified labelers. Thus gender disproportion does not contribute to the bias in the data. We do not need to worry about that. However, I believe the age-group demographic profile of workers would make a difference. If more time is granted, furthur analysis on the age-group should be done.
# ## Step 2: Future implications
# 1. Which, if any, of these demo applications would you expect the Perspective API to perform poorly in? Why?
# We might not forsee the bias in the demographic data beside gender of the Crowdflower workers. I suspect that there is bias towards the proportion of young people in the worker population dominates the likelyhood to label comments as aggressive. Thus the API such as Comment Filter might have high false positive errors.
# 2. Imagine you are one of the Google data scientists who maintains the Perspective API. If you wanted to improve the model or the API itself to make it work better for any of these purposes, how should they go about doing that?
# We should take a deeper investigation towards the potential bias in the demographic of workers. There are a lot more to explore besides age-group. Since all the comments were in English and only 17% of the annotators said they speak English as their first language there might be some error in interpretation of the comments. This data also grouped annotators by age into ranges of 15, grouping by a smaller range might produce different results. We would want to weight the aggression/toxicity scores of the comments axcording to out furthur analysis.
# 3. What are some potential unintended, negative consequences of using the Perspective API for any of these purposes? In your opinion, are these consequences likely or serious enough that you would recommend that the Perspective API not be used in these applications? Why or why not?
# some potential unintended, negative consequences of using the Perspective API could be both the type I and the type II error. The API falsely accusing a person of being a negative user on a specific platform or missing a toxic user thus allowing them to stay on the platform.
#
|
data_512_a2/data_512_a2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Univariate Distribution Similarity
#
# [](https://colab.research.google.com/github/ing-bank/probatus/blob/master/docs/tutorials/nb_distribution_statistics.ipynb)
# There are many situations when you want to perform univariate distribution comparison of a given feature, e.g. stability of the feature over different months.
#
# In order to do that, you can use statistical tests. In this tutorial we present how to easily do this using the `DistributionStatistics` class, and with the statistical tests directly.
#
# The available statistical tests in `probatus.stat_tests` are:
# - Epps-Singleton ('ES')
# - Kolmogorov-Smirnov statistic ('KS')
# - Population Stability Index ('PSI')
# - Shapiro-Wilk based difference statistic ('SW')
# - Anderson-Darling TS ('AD')
#
# You can perform all these tests using a convenient wrapper class called `DistributionStatistics`.
#
# In this tutorial we will focus on how to perform two useful tests: Population Stability Index (widely applied in banking industry) and Kolmogorov-Smirnov.
# ## Setup
# +
# %load_ext autoreload
# %autoreload 2
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from probatus.binning import AgglomerativeBucketer, SimpleBucketer, QuantileBucketer
from probatus.stat_tests import DistributionStatistics, psi, ks
# -
# Let's define some test distributions and visualize them. For these examples, we will use a normal distribution and a shifted version of the same distribution.
counts = 1000
np.random.seed(0)
d1 = pd.Series(np.random.normal(size=counts), name='feature_1')
d2 = pd.Series(np.random.normal(loc=0.5, size=counts), name='feature_1')
# +
from probatus.utils.plots import plot_distributions_of_feature
feature_distributions = [d1, d2]
sample_names = ['expected', 'actual']
plot_distributions_of_feature(feature_distributions, sample_names=sample_names, plot_perc_outliers_removed=0.01)
# -
# ### Binning - QuantileBucketer
#
# To visualize the data, we will bin the data using a quantile bucketer, available in the `probatus.binning` module.
#
# Binning is used by all the `stats_tests` in order to group observations.
# +
bins = 10
myBucketer = QuantileBucketer(bins)
d1_bincounts = myBucketer.fit_compute(d1)
d2_bincounts = myBucketer.compute(d2)
print("Bincounts for d1 and d2:")
print(d1_bincounts)
print(d2_bincounts)
# -
# Let's plot the distribution for which we will calculate the statistics.
plt.figure(figsize=(20,5))
plt.bar(range(0, len(d1_bincounts)), d1_bincounts, label='d1: expected')
plt.bar(range(0, len(d2_bincounts)), d2_bincounts, label='d2: actual', alpha=0.5)
plt.title('PSI (bucketed)', fontsize=16, fontweight='bold')
plt.legend(fontsize=15)
plt.show()
# By visualizing the bins, we can already notice that the distributions are different.
#
# Let's use the statistical test to prove that.
# ## PSI - Population Stability Index
# The population stability index ([Karakoulas, 2004](https://cms.rmau.org/uploadedFiles/Credit_Risk/Library/RMA_Journal/Other_Topics_(1998_to_present)/Empirical%20Validation%20of%20Retail%20Credit-Scoring%20Models.pdf)) has long been used to evaluate distribution similarity in banking industry, while developing credit decision models.
#
# In probatus we have implemented the PSI according to [Yurdakul 2018](https://scholarworks.wmich.edu/cgi/viewcontent.cgi?article=4249&context=dissertations), which derives a p-value, based on the hard to interpret PSI statistic. Using the p-value is a more reliable choice, because the banking industry-standard PSI critical values of 0.1 and 0.25 are unreliable heuristics because there is a strong dependency on sample sizes and number of bins. Aside from these heuristics, the PSI value is not easily interpretable in the context of common statistical frameworks (like a p-value or confidence levels).
psi_value, p_value = psi(d1_bincounts, d2_bincounts, verbose=True)
# Based on the above test, the distribution between the two samples significantly differs.
# Not only the PSI statistic is above the commonly used critical value, but also the p-value shows a very high confidence.
# ## PSI with DistributionStatistics
# Using `DistributionStatistics` class one can apply the above test, without the need to manually perform the binning. We initialize a `DistributionStatistics` instance with the desired test, binning_strategy (or choose `"default"` to choose the test's most appropriate binning strategy) and the number of bins. Then we start the test with the unbinned values as input.
distribution_test = DistributionStatistics("psi", binning_strategy="default", bin_count=10)
psi_value, p_value = distribution_test.compute(d1, d2, verbose=True)
# ## KS: Kolmogorov-Smirnov with DistributionStatistics
# The Kolmogorov-Smirnov test compares two distributions by calculating the maximum difference of the two samples' distribution functions, as illustrated by the black arrow in the following figure. The KS test is available in `probatus.stat_tests.ks`.
#
# <img align="middle" src="../img/KS2_Example.png" alt="Example of the Kolmogorov-Smirnov value" width="400" height="400">
#
# The main advantage of this method is its sensitivity to differences in both location and shape of the empirical cumulative distribution functions of the two samples.
#
# The main disadwantages are that: it works for continuous distributions (unless modified, e.g. see ([Jeng 2006](https://bmcmedresmethodol.biomedcentral.com/track/pdf/10.1186/1471-2288-6-45)), in large samples, small and unimportant differences can be statistically significant ([Taplin & Hunt 2019](https://www.mdpi.com/2227-9091/7/2/53/pdf)), and finally in small samples, large and important differences can be statistically insignificant ([Taplin & Hunt 2019](https://www.mdpi.com/2227-9091/7/2/53/pdf)).
# As before, using the test requires you to perform the binning beforehand
k_value, p_value = ks(d1, d2, verbose=True)
# Again, we can also choose to combine the binning and the statistical test using the `DistributionStatistics` class.
distribution_test = DistributionStatistics("ks", binning_strategy=None)
ks_value, p_value = distribution_test.compute(d1, d2, verbose=True)
# ## AutoDist
from probatus.stat_tests import AutoDist
# Multiple statistics can automatically be calculated using `AutoDist`. To show this, let's create two new dataframes with two features each.
# +
size, n_features = 100, 2
df1 = pd.DataFrame(np.random.normal(size=(size, n_features)), columns=[f'feat_{x}' for x in range(n_features)])
df2 = pd.DataFrame(np.random.normal(size=(size, n_features)), columns=[f'feat_{x}' for x in range(n_features)])
# -
# We can now specify the statistical tests we want to perform and the binning strategies to perform. We can also set both of these variables to `'all'` or binning strategies to `'default'` to use the default binning strategy for every chosen statistical test.
statistical_tests = ["KS", "PSI"]
binning_strategies = "default"
# Let's compute the statistics and their p_values:
myAutoDist = AutoDist(statistical_tests=statistical_tests, binning_strategies=binning_strategies, bin_count=10)
myAutoDist.compute(df1, df2)
|
docs/tutorials/nb_distribution_statistics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import powerlaw
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
import scipy.stats
# +
a, xmin = 2.5 , 1.0
N = 10000
# generates random variates of power law distribution
xx = list(powerlaw.Power_Law(xmin=xmin, parameters=[a]).generate_random(N))
# -
max(xx)
min(xx)
# # 1-Plotting linear histogram
plt.hist(xx,bins=100)
plt.xlabel('x')
plt.ylabel('P(x)')
plt.show()
# +
ybinLin,xbinLin=np.histogram(xx,bins=100)
ynorm=[float(i)/sum(ybinLin) for i in ybinLin]
plt.plot(xbinLin[:-1],ynorm,'o-')
plt.xlabel('x')
plt.ylabel('P(x)')
plt.show()
# -
# # 2-Plotting log-log histogram
# +
plt.loglog(xbinLin[:-1],ynorm,'o')
plt.xlabel('x')
plt.ylabel('P(x)')
plt.show()
# -
# # 3- Plotting cumulative distribution
ycum=[sum(ynorm[i:]) for i in range(len(ynorm))]
plt.loglog(xbinLin[:-1],ycum,'o')
plt.title('cumulative')
plt.xlabel('x')
plt.ylabel('$P^C(x)$')
plt.show()
# # 4- Plotting binned distribution
#
def logBinning(degreeList,nbin):
kmin=min(degreeList)
kmax=max(degreeList)
logBins = np.logspace(np.log10(kmin), np.log10(kmax),num=nbin)
logBinDensity, binedges = np.histogram(degreeList, bins=logBins, density=True)
logBins = np.delete(logBins, -1)
return logBinDensity, logBins
y,x=logBinning(np.array(xx),50)
plt.loglog(x,y,'o')
plt.xlabel('x')
plt.ylabel('P(x)')
plt.show()
# # 5-calculating power-law fit
def powerLaw(x, a, b):
return b*x**(a)
# +
fitx=[i for (i,j) in zip(x,y) if i<100]
fity=[j for (i,j) in zip(x,y) if i<100]
popt, pcov = curve_fit(powerLaw, fitx, fity)
print (popt[0])
plt.loglog(sorted(x), powerLaw(sorted(x), *popt), '--',c='k',linewidth=3)
plt.loglog(x,y,'o')
plt.xlabel('x')
plt.ylabel('P(x)')
plt.show()
# -
# # 6-Plot ranking
# +
plt.loglog(range(len(xx)),sorted(xx,reverse=True),'o')
plt.xlabel('rank')
plt.ylabel('x')
plt.show()
# -
# # BIVARIATE STATISTICS
# xx is distributed according to a power-law distribution. yy is derived from a uniform distribution. I have a set of bivariate variables: (x,y)
yy=[random.uniform(0,1) for i in xx]
plt.hist(yy)
plt.show()
# simple plot of the (x,y) couples
plt.loglog(xx,yy,'o',alpha=0.1)
plt.show()
logBins=np.logspace(np.log2(np.min(xx)),np.log2(np.max(xx)),base=2,num=10)
ybin,xbin,binnumber=scipy.stats.binned_statistic(xx,yy,statistic='mean',bins=logBins)
plt.semilogx(xx,yy,'o',alpha=0.1)
plt.semilogx(xbin[:-1],ybin,'o',markersize=10)
plt.show()
bin_stdevs, _, _ = scipy.stats.binned_statistic(xx, yy,
statistic='std',
bins=logBins)
# +
logBins=np.logspace(np.log2(np.min(xx)),np.log2(np.max(xx)),base=2,num=10)
ybin,xbin,binnumber=scipy.stats.binned_statistic(xx,yy,statistic='mean',bins=logBins)
plt.semilogx(xx,yy,'o',alpha=0.1)
plt.semilogx(xbin[:-1],ybin,'o',markersize=10)
plt.errorbar(xbin[:-1], ybin, bin_stdevs, fmt='o',color="k")
plt.show()
# -
|
CC4/exClasse/Seance4_ExerciseClasse1.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
import java.nio.file.Paths
import org.apache.spark.sql.functions._
val dir_ot = Paths.get(System.getProperty("user.home"), "data", "ot", "extract")
var df = spark.read.json(dir_ot.resolve("evidence.json").toString)
df.select("scores").printSchema()
// +
// ES Query:
// GET master_evidence-data/_search
// {
// "query": {
// "bool": {
// "must": [
// { "match": { "disease.id": "EFO_0001645" }},
// { "match": { "target.id": "ENSG00000226777"}},
// { "match": { "sourceID": "sysbio"}}
// ]
// }
// }
// }
df.filter($"id" === "abb040109d9414a1f5e764f031ffc3b4")
.select($"scores", $"target.id", $"disease.id", $"sourceID").show()
// -
df.groupBy("sourceID").count().show()
df.filter($"sourceID" === "sysbio").show(3)
// +
var dfe = df
.filter($"disease.id" === "EFO_0001645")
.filter($"target.id" === "ENSG00000226777")
.filter($"sourceID" === "sysbio")
dfe.show(3)
// -
// Results from scoring-poc after explosion:
//
// ```
// +--------------------+---------+---------------+---------------+-----+-----------+------------+
// | id|source_id|orig_disease_id| target_id|score| disease_id|is_direct_id|
// +--------------------+---------+---------------+---------------+-----+-----------+------------+
// |abb040109d9414a1f...| sysbio| EFO_0001645|ENSG00000226777|0.859|EFO_0003777| false|
// |abb040109d9414a1f...| sysbio| EFO_0001645|ENSG00000226777|0.859|EFO_0000408| false|
// |abb040109d9414a1f...| sysbio| EFO_0001645|ENSG00000226777|0.859|EFO_0000319| false|
// |abb040109d9414a1f...| sysbio| EFO_0001645|ENSG00000226777|0.859|EFO_0001645| true|
// +--------------------+---------+---------------+---------------+-----+-----------+------------+
// ```
dfe.select($"scores", $"target", $"disease", $"disease.id").show(1, 1000)
df.printSchema
|
notebooks/dev/sysbio-debug.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="c10QRmlWYxnn"
# # 1d CNNs for sentiment classification
#
# We use 1d CNNs for IMDB movie review classification.
# Based on sec 15.3 of http://d2l.ai/chapter_natural-language-processing-applications/sentiment-analysis-cnn.html
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="NjPuDYiqft1B" outputId="8e2410be-0a9c-4f8a-b1d0-947545fdbf56"
# !pip install flax
# + id="o6VNlv_FYTbS"
import numpy as np
import matplotlib.pyplot as plt
import math
from IPython import display
import jax
import jax.numpy as jnp
from flax import linen as nn
from flax.training import train_state
import optax
import collections
import re
import random
import os
import requests
import zipfile
import tarfile
import hashlib
import time
import functools
from typing import Sequence
rng = jax.random.PRNGKey(0)
# !mkdir figures # for saving plots
# + [markdown] id="eAqzyDB2ZI1E"
# # Data
#
# We use IMDB dataset. Details in [this colab](https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/rnn_sentiment_torch.ipynb).
# + id="Cy4JfSNBgnHS"
# Required functions for downloading data
def download(name, cache_dir=os.path.join('..', 'data')):
"""Download a file inserted into DATA_HUB, return the local filename."""
assert name in DATA_HUB, f"{name} does not exist in {DATA_HUB}."
url, sha1_hash = DATA_HUB[name]
os.makedirs(cache_dir, exist_ok=True)
fname = os.path.join(cache_dir, url.split('/')[-1])
if os.path.exists(fname):
sha1 = hashlib.sha1()
with open(fname, 'rb') as f:
while True:
data = f.read(1048576)
if not data:
break
sha1.update(data)
if sha1.hexdigest() == sha1_hash:
return fname # Hit cache
print(f'Downloading {fname} from {url}...')
r = requests.get(url, stream=True, verify=True)
with open(fname, 'wb') as f:
f.write(r.content)
return fname
def download_extract(name, folder=None):
"""Download and extract a zip/tar file."""
fname = download(name)
base_dir = os.path.dirname(fname)
data_dir, ext = os.path.splitext(fname)
if ext == '.zip':
fp = zipfile.ZipFile(fname, 'r')
elif ext in ('.tar', '.gz'):
fp = tarfile.open(fname, 'r')
else:
assert False, 'Only zip/tar files can be extracted.'
fp.extractall(base_dir)
return os.path.join(base_dir, folder) if folder else data_dir
# + id="1mJDy6S3QFic"
def read_imdb(data_dir, is_train):
data, labels = [], []
for label in ('pos', 'neg'):
folder_name = os.path.join(data_dir, 'train' if is_train else 'test',
label)
for file in os.listdir(folder_name):
with open(os.path.join(folder_name, file), 'rb') as f:
review = f.read().decode('utf-8').replace('\n', '')
data.append(review)
labels.append(1 if label == 'pos' else 0)
return data, labels
# + [markdown] id="nx6X6FoeQNVX"
# We tokenize using words, and drop words which occur less than 5 times in training set when creating the vocab.
# + id="nJmK1jtkg5wC"
def tokenize(lines, token='word'):
"""Split text lines into word or character tokens."""
if token == 'word':
return [line.split() for line in lines]
elif token == 'char':
return [list(line) for line in lines]
else:
print('ERROR: unknown token type: ' + token)
class Vocab:
"""Vocabulary for text."""
def __init__(self, tokens=None, min_freq=0, reserved_tokens=None):
if tokens is None:
tokens = []
if reserved_tokens is None:
reserved_tokens = []
# Sort according to frequencies
counter = count_corpus(tokens)
self.token_freqs = sorted(counter.items(), key=lambda x: x[1],
reverse=True)
# The index for the unknown token is 0
self.unk, uniq_tokens = 0, ['<unk>'] + reserved_tokens
uniq_tokens += [
token for token, freq in self.token_freqs
if freq >= min_freq and token not in uniq_tokens]
self.idx_to_token, self.token_to_idx = [], dict()
for token in uniq_tokens:
self.idx_to_token.append(token)
self.token_to_idx[token] = len(self.idx_to_token) - 1
def __len__(self):
return len(self.idx_to_token)
def __getitem__(self, tokens):
if not isinstance(tokens, (list, tuple)):
return self.token_to_idx.get(tokens, self.unk)
return [self.__getitem__(token) for token in tokens]
def to_tokens(self, indices):
if not isinstance(indices, (list, tuple)):
return self.idx_to_token[indices]
return [self.idx_to_token[index] for index in indices]
def count_corpus(tokens):
"""Count token frequencies."""
# Here `tokens` is a 1D list or 2D list
if len(tokens) == 0 or isinstance(tokens[0], list):
# Flatten a list of token lists into a list of tokens
tokens = [token for line in tokens for token in line]
return collections.Counter(tokens)
def set_figsize(figsize=(3.5, 2.5)):
"""Set the figure size for matplotlib."""
display.set_matplotlib_formats('svg')
plt.rcParams['figure.figsize'] = figsize
# + [markdown] id="P0UGjVDmQYr2"
# We pad all sequences to length 500, for efficient minibatching.
# + id="7KjC8-syi-id"
def truncate_pad(line, num_steps, padding_token):
"""Truncate or pad sequences."""
if len(line) > num_steps:
return line[:num_steps] # Truncate
return line + [padding_token] * (num_steps - len(line))
# + [markdown] id="0HHnFY2UQiRE"
# Putting it altogether.
# + id="J4C0tLI1QgN5"
def load_data_imdb(num_steps=500):
data_dir = download_extract('aclImdb', 'aclImdb')
train_data = read_imdb(data_dir, True)
test_data = read_imdb(data_dir, False)
train_tokens = tokenize(train_data[0], token='word')
test_tokens = tokenize(test_data[0], token='word')
vocab = Vocab(train_tokens, min_freq=5)
train_features = jnp.asarray([
truncate_pad(vocab[line], num_steps, vocab['<pad>'])
for line in train_tokens])
test_features = jnp.asarray([
truncate_pad(vocab[line], num_steps, vocab['<pad>'])
for line in test_tokens])
train_labels = jnp.asarray(train_data[1])
test_labels = jnp.asarray(test_data[1])
train_ds = {
'text': train_features,
'label': train_labels
}
test_ds = {
'text': test_features,
'label': test_labels
}
return train_ds, test_ds, vocab
# + id="5ADmjrfwPnTw" colab={"base_uri": "https://localhost:8080/"} outputId="f45fd488-1d68-4796-c742-8c0d2bb8e8e2"
DATA_HUB = dict()
DATA_HUB['aclImdb'] = (
'http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz',
'01ada507287d82875905620988597833ad4e0903')
data_dir = download_extract('aclImdb', 'aclImdb')
# + id="KVS7n4hNQmQR"
train_ds, test_ds, vocab = load_data_imdb()
# + id="1_QNbMw0OZK3"
assert train_ds['text'].shape == (25000, 500)
assert train_ds['label'].shape == (25000,)
assert test_ds['text'].shape == (25000, 500)
assert test_ds['label'].shape == (25000,)
# + [markdown] id="-tf9T5YRZh3z"
# # Model
#
# We load pretrained Glove vectors. We use these to initialize the embedding layers, one of which is frozen.
#
# + id="JnEDTL4Fj--D"
class TokenEmbedding:
"""Token Embedding."""
def __init__(self, embedding_name):
self.idx_to_token, self.idx_to_vec = self._load_embedding(
embedding_name)
self.unknown_idx = 0
self.token_to_idx = {
token: idx for idx, token in enumerate(self.idx_to_token)}
def _load_embedding(self, embedding_name):
idx_to_token, idx_to_vec = ['<unk>'], []
data_dir = download_extract(embedding_name)
# GloVe website: https://nlp.stanford.edu/projects/glove/
# fastText website: https://fasttext.cc/
with open(os.path.join(data_dir, 'vec.txt'), 'r') as f:
for line in f:
elems = line.rstrip().split(' ')
token, elems = elems[0], [float(elem) for elem in elems[1:]]
# Skip header information, such as the top row in fastText
if len(elems) > 1:
idx_to_token.append(token)
idx_to_vec.append(elems)
idx_to_vec = [[0] * len(idx_to_vec[0])] + idx_to_vec
return idx_to_token, np.array(idx_to_vec)
def __getitem__(self, tokens):
indices = [
self.token_to_idx.get(token, self.unknown_idx)
for token in tokens]
vecs = self.idx_to_vec[np.array(indices)]
return vecs
def __len__(self):
return len(self.idx_to_token)
# + id="QjKHAIjdZ9vt" colab={"base_uri": "https://localhost:8080/"} outputId="af4aff6f-f07b-466f-a93a-4dd778b5ff98"
DATA_URL = 'http://d2l-data.s3-accelerate.amazonaws.com/glove.6B.100d.zip'
DATA_HUB['glove.6b.100d'] = (DATA_URL, 'cd43bfb07e44e6f27cbcc7bc9ae3d80284fdaf5a')
glove_embedding = TokenEmbedding('glove.6b.100d')
embeds = jnp.array(glove_embedding[vocab.idx_to_token])
assert embeds.shape == (49346, 100)
# + [markdown] id="NX98u24BaAkP"
# We use 2 embedding layers, one with frozen weights, and one with learnable weights. We feed their concatenation to the 1d CNN. We then do average pooling over time before passing into the final MLP to map to the 2 output logits.
# + id="slGCo3OBZN3-"
class TextCNN(nn.Module):
vocab_size: int
embed_size: int
kernel_sizes: Sequence[int]
num_channels: Sequence[int]
embeds: jnp.array
def setup(self):
self.embedding = nn.Embed(self.vocab_size, self.embed_size,
embedding_init = lambda *_: self.embeds)
# The embedding layer does not participate in training
self.constant_embedding = nn.Embed(self.vocab_size, self.embed_size,
embedding_init = lambda *_: self.embeds)
self.dropout = nn.Dropout(0.5)
self.decoder = nn.Dense(2)
self.convs = [nn.Conv(features=c, kernel_size=(k,), padding='VALID')
for c, k in zip(self.num_channels, self.kernel_sizes)]
def __call__(self, x, *, train):
batch_size, num_words = x.shape
# Concatenate the output of two embedding layers with shape of
# (batch size, no. of words, word vector dimension) by word vector
embeddings = jnp.concatenate(
[self.embedding(x), self.constant_embedding(x)], axis=2)
assert embeddings.shape == (batch_size, num_words, 2*self.embed_size)
# For each one-dimensional convolutional layer, after max-over-time
# pooling, a tensor with the shape of (batch size, 1, channel size)
# can be obtained
for conv, c, k in zip(self.convs, self.num_channels, self.kernel_sizes):
assert conv(embeddings).shape == (batch_size, num_words+1-k, c)
assert nn.max_pool(conv(embeddings), window_shape=(num_words+1-k,)).shape == (batch_size, 1, c)
# Use the flatten function to remove the penultimate dimension and
# then concatenate on the channel dimension
encoding = jnp.concatenate([
jnp.squeeze(nn.relu(nn.max_pool(conv(embeddings), window_shape=(num_words+1-k,))), axis=1)
for conv, k in zip(self.convs, self.kernel_sizes)], axis=1)
assert encoding.shape == (batch_size, sum(self.num_channels))
# After applying the dropout method, use a fully connected layer to
# obtain the output
outputs = self.decoder(self.dropout(encoding, deterministic=not train))
assert outputs.shape == (batch_size, 2)
return outputs
# + id="K3Yb77I95MIG"
embed_size, kernel_sizes, num_channels = 100, [3, 4, 5], [100, 100, 100]
TextCNN = functools.partial(TextCNN, len(vocab), embed_size, kernel_sizes, num_channels, embeds)
# + [markdown] id="6MW7I7lbTH82"
# # Training
# + id="tA_QeZH5rp1o"
class Animator:
"""For plotting data in animation."""
def __init__(self, xlabel=None, ylabel=None, legend=None, xlim=None,
ylim=None, xscale='linear', yscale='linear',
fmts=('-', 'm--', 'g-.', 'r:'), nrows=1, ncols=1,
figsize=(3.5, 2.5)):
# Incrementally plot multiple lines
if legend is None:
legend = []
display.set_matplotlib_formats('svg')
self.fig, self.axes = plt.subplots(nrows, ncols, figsize=figsize)
if nrows * ncols == 1:
self.axes = [self.axes,]
# Use a lambda function to capture arguments
self.config_axes = lambda: set_axes(self.axes[
0], xlabel, ylabel, xlim, ylim, xscale, yscale, legend)
self.X, self.Y, self.fmts = None, None, fmts
def add(self, x, y):
# Add multiple data points into the figure
if not hasattr(y, "__len__"):
y = [y]
n = len(y)
if not hasattr(x, "__len__"):
x = [x] * n
if not self.X:
self.X = [[] for _ in range(n)]
if not self.Y:
self.Y = [[] for _ in range(n)]
for i, (a, b) in enumerate(zip(x, y)):
if a is not None and b is not None:
self.X[i].append(a)
self.Y[i].append(b)
self.axes[0].cla()
for x, y, fmt in zip(self.X, self.Y, self.fmts):
self.axes[0].plot(x, y, fmt)
self.config_axes()
display.display(self.fig)
display.clear_output(wait=True)
class Timer:
"""Record multiple running times."""
def __init__(self):
self.times = []
self.start()
def start(self):
"""Start the timer."""
self.tik = time.time()
def stop(self):
"""Stop the timer and record the time in a list."""
self.times.append(time.time() - self.tik)
return self.times[-1]
def avg(self):
"""Return the average time."""
return sum(self.times) / len(self.times)
def sum(self):
"""Return the sum of time."""
return sum(self.times)
def cumsum(self):
"""Return the accumulated time."""
return np.array(self.times).cumsum().tolist()
class Accumulator:
"""For accumulating sums over `n` variables."""
def __init__(self, n):
self.data = [0.0] * n
def add(self, *args):
self.data = [a + float(b) for a, b in zip(self.data, args)]
def reset(self):
self.data = [0.0] * len(self.data)
def __getitem__(self, idx):
return self.data[idx]
# + id="FqnIuSGwxG0L"
def set_axes(axes, xlabel, ylabel, xlim, ylim, xscale, yscale, legend):
"""Set the axes for matplotlib."""
axes.set_xlabel(xlabel)
axes.set_ylabel(ylabel)
axes.set_xscale(xscale)
axes.set_yscale(yscale)
axes.set_xlim(xlim)
axes.set_ylim(ylim)
if legend:
axes.legend(legend)
axes.grid()
# + id="vBqC89zOr_Si"
def compute_metrics(*, logits, labels):
one_hot = jax.nn.one_hot(labels, num_classes=2)
loss = jnp.mean(optax.softmax_cross_entropy(logits=logits, labels=one_hot))
accuracy = jnp.mean(jnp.argmax(logits, -1) == labels)
metrics = {
'loss': loss,
'accuracy': accuracy,
}
return metrics
def create_train_state(rng, learning_rate):
"""Creates initial `TrainState`."""
cnn = TextCNN()
params = cnn.init(rng, jnp.ones([1, 500], dtype=int), train=False)['params']
tx = optax.adam(learning_rate, )
return train_state.TrainState.create(
apply_fn=cnn.apply, params=params, tx=tx)
# + id="wQGd4KeCrMzT"
@jax.jit
def train_step(state, batch, dropout_rng):
"""Train for a single step."""
def loss_fn(params):
logits = TextCNN().apply({'params': params}, batch['text'], train=True,
rngs={'dropout': dropout_rng})
one_hot = jax.nn.one_hot(batch['label'], num_classes=2)
loss = jnp.sum(optax.softmax_cross_entropy(logits=logits, labels=one_hot))
return loss, logits
grad_fn = jax.value_and_grad(loss_fn, has_aux=True)
(_, logits), grads = grad_fn(state.params)
grads = grads.copy({'constant_embedding': {'embedding': 0}}) # don't update this layer
state = state.apply_gradients(grads=grads)
metrics = compute_metrics(logits=logits, labels=batch['label'])
return state, metrics
@jax.jit
def eval_step(params, batch):
logits = TextCNN().apply({'params': params}, batch['text'], train=False)
return compute_metrics(logits=logits, labels=batch['label'])
def train_epoch(state, train_ds, batch_size, epoch, rng, dropout_rng, animator):
"""Train for a single epoch."""
train_ds_size = len(train_ds['text'])
steps_per_epoch = train_ds_size // batch_size
perms = jax.random.permutation(rng, train_ds_size)
perms = perms[:steps_per_epoch * batch_size] # skip incomplete batch
perms = perms.reshape((steps_per_epoch, batch_size))
batch_metrics = []
for perm in perms:
batch = {k: v[perm, ...] for k, v in train_ds.items()}
state, metrics = train_step(state, batch, dropout_rng)
batch_metrics.append(metrics)
# compute mean of metrics across each batch in epoch.
batch_metrics_np = jax.device_get(batch_metrics)
epoch_metrics_np = {
k: np.mean([metrics[k] for metrics in batch_metrics_np])
for k in batch_metrics_np[0]}
animator.add(epoch, (epoch_metrics_np['loss'], epoch_metrics_np['accuracy'], None))
print('train epoch: %d, loss: %.4f, accuracy: %.2f' % (
epoch, epoch_metrics_np['loss'], epoch_metrics_np['accuracy'] * 100))
return state
def eval_model(params, test_ds, batch_size):
test_ds_size = len(test_ds['text'])
steps_per_epoch = test_ds_size // batch_size
perms = jax.random.permutation(rng, test_ds_size)
perms = perms[:steps_per_epoch * batch_size] # skip incomplete batch
perms = perms.reshape((steps_per_epoch, batch_size))
batch_metrics = []
for perm in perms:
batch = {k: v[perm, ...] for k, v in test_ds.items()}
metrics = eval_step(params, batch)
batch_metrics.append(metrics)
# compute mean of metrics across each batch in epoch.
batch_metrics_np = jax.device_get(batch_metrics)
epoch_metrics_np = {
k: np.mean([metrics[k] for metrics in batch_metrics_np])
for k in batch_metrics_np[0]}
return epoch_metrics_np['loss'], epoch_metrics_np['accuracy']
# + [markdown] id="mUty6y23Lh7C"
# # Learning curve
# + id="bEnxoEyZGSF5"
rng, init_rng = jax.random.split(rng)
learning_rate = 0.001
state = create_train_state(init_rng, learning_rate)
del init_rng # Must not be used anymore.
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="VaspjLDbaP4Y" outputId="8c053ec2-c8fe-47f5-9c7b-0f24e364953f"
num_epochs = 5
batch_size = 64
animator = Animator(xlabel='epoch', xlim=[1, num_epochs], legend=['train loss', 'train acc', 'test acc'])
for epoch in range(1, num_epochs + 1):
# Use a separate PRNG key to permute image data during shuffling
rng, input_rng = jax.random.split(rng)
# dropout needs PRNG for "dropout"
rng, dropout_rng = jax.random.split(rng)
# Run an optimization step over a training batch
state = train_epoch(state, train_ds, batch_size, epoch, input_rng, dropout_rng, animator)
# Evaluate on the test set after each training epoch
test_loss, test_accuracy = eval_model(state.params, test_ds, batch_size)
animator.add(epoch, (None, None, test_accuracy))
print('test epoch: %d, test_loss: %.2f, accuracy: %.2f' % (epoch, test_loss, test_accuracy * 100))
# + [markdown] id="rPcdmigTaT8X"
# # Testing
# + id="aNrZA-eGTIt6"
def predict_sentiment(params, vocab, sentence):
sentence = jnp.asarray(vocab[sentence.split()])
logits = TextCNN().apply({'params': params}, sentence.reshape(1, -1), train=False)
label = jnp.argmax(logits, axis=1)
return 'positive' if label == 1 else 'negative'
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="wZhdLe6IaUpq" outputId="e0bc0a81-1500-4286-890b-45d78b2937b7"
predict_sentiment(state.params, vocab, 'this movie is so great')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="uNdbHZmYaXRc" outputId="e48ffde7-db7e-418d-a960-7c87df8bc639"
predict_sentiment(state.params, vocab, 'this movie is so bad')
|
notebooks-d2l/cnn1d_sentiment_jax.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="g3EbbVLwIJ74"
# # Horizontal DAS + vibroseis data from Brady Hot Springs geothermal site
#
#
# + [markdown] id="YKMXiJEWyDhY"
# Notebook created by <NAME> (Virginia Tech) and edited by <NAME> (Tel Aviv University). Assistance with editing provided by <NAME> (Stanford), <NAME> (Caltech), <NAME> (Colorado School of Mines).
# + [markdown] id="7uuXh507_EhX"
# In this notebook, we'll take a look at how to do some basic visualization of DAS data with a trenched surface array at Brady Hot Springs in Nevada. A brief overview of this project can be found at https://www.energy.gov/eere/articles/nevada-site-home-geothermal-community-focused-expediting-research-and-development
#
# 
#
# *Image of Fumeroles at Brady Hot Springs, photo by <NAME> (originally from https://www.energy.gov/eere/articles/nevada-site-home-geothermal-community-focused-expediting-research-and-development)*
#
#
#
# These data were acquired by the PoroTomo team: http://geoscience.wisc.edu/geoscience/people/faculty/feigl/porotomo/
#
# **References:**
# * <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, PoroTomoTeam, 2017, "[High-resolution Shallow Structure Revealed with Ambient Noise Tomography on a Dense Array](https://www.researchgate.net/profile/Dante_Fratta/publication/316092404_High-resolution_Shallow_Structure_Revealed_with_Ambient_Noise_Tomography_on_a_Dense_Array/links/58eff4a0a6fdcc11e569dc8d/High-resolution-Shallow-Structure-Revealed-with-Ambient-Noise-Tomography-on-a-Dense-Array.pdf)," Proceedings, 42nd Workshop on Geothermal Reservoir Engineering, Stanford University, Stanford, CA, Feb. 13-15.
# * <NAME> and the PoroTomo Team, 2017, "[Overview and Preliminary Results from the PoroTomo project at Brady Hot Springs, Nevada: Poroelastic Tomography by Adjoint Inverse Modeling of Data from Seismology, Geodesy, and Hydrology](https://www.researchgate.net/profile/Dante_Fratta/publication/316092238_Overview_and_Preliminary_Results_from_the_PoroTomo_project_at_Brady_Hot_Springs_Nevada_Poroelastic_Tomography_by_Adjoint_Inverse_Modeling_of_Data_from_Seismology_Geodesy_and_Hydrology/links/58eff6e1458515ff23a88b48/Overview-and-Preliminary-Results-from-the-PoroTomo-project-at-Brady-Hot-Springs-Nevada-Poroelastic-Tomography-by-Adjoint-Inverse-Modeling-of-Data-from-Seismology-Geodesy-and-Hydrology.pdf)," Proceedings, 42nd Workshop on Geothermal Reservoir Engineering, Stanford University, Stanford, CA, Feb. 13-15.
# * <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, 2017, "[Geothermal Reservoir Characterization Using Distributed Temperature Sensing at Brady Geothermal Field, Nevada](https://library.seg.org/doi/am-pdf/10.1190/tle36121024a1.1)," The Leading Edge, 36(12), 1024a1-1024a7.
# * <NAME>, 2017, "[Active Souce 3D Seismic Tomography of Brady Hot Springs Geothermal Field, Nevada](https://gdr.openei.org/files/1070/Parker_MS_Thesis.pdf)," M.S. Thesis, University of Wisconsin-Madison.
# * <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, 2018, "[Ground motion response to an ML 4.3 earthquake using co-located distributed acoustic sensing and seismometer arrays](https://academic.oup.com/gji/article/213/3/2020/4942237)," Geophysical Journal International, 213(3), 2020-2036.
# * <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, 2018, "[Active-Source Seismic Tomography at the Brady Geothermal Field, Nevada with Dense Nodal and Fiber-Optic Seismic Arrays](https://pubs.geoscienceworld.org/ssa/srl/article-abstract/545060/active-source-seismic-tomography-at-the-brady)," Seismological Research Letters, 89(5), 1629-1640.
# * <NAME>, <NAME>, <NAME>, 2018, "[Characterizing volumetric strain at Brady Hot Springs, Nevada, USA using geodetic data, numerical models and prior information](https://academic.oup.com/gji/article/215/2/1501/5078355)," Geophysical Journal International, 215(2), 1501-1513.
# * <NAME> and <NAME>, 2018, "[Pushing the limit of earthquake detection with distributed acoustic sensing and template matching: a case study at the Brady geothermal field](https://academic.oup.com/gji/article/215/3/1583/5090146)," Geophysical Journal International, 215(3), 1583-1593.
# * <NAME>., <NAME>., <NAME>., <NAME>., and <NAME>., 2019, "3D Imaging of Geothermal Faults from a Vertical DAS Fiber at Brady Hot Spring, NV USA," Energies, 12(1401). https://doi.org/10.3390/en12071401
# * <NAME>., <NAME>, and PoroTomo_Team, 2019, "PoroTomo Final Technical Report: Poroelastic Tomography by Adjoint Inverse Modeling of Data from Seismology, Geodesy, and Hydrology," https://www.osti.gov/servlets/purl/1499141
# + [markdown] id="oq4wML6FJUBa"
# The Python packages we'll use are:
#
# * numpy, already available in Colab: https://numpy.org/
# * matplotlib.pyplot, already available in Colab: https://matplotlib.org/
# * obspy, needs to be installed: https://docs.obspy.org/
# * scipy.fft, already available in Colab: https://docs.scipy.org/doc/scipy/reference/fft.html#module-scipy.fft
# * csv, already available in Colab: https://docs.python.org/3/library/csv.html
# * ipywidgets, already available in Colab: https://ipywidgets.readthedocs.io/en/stable/
# * gdown to grab large files from Google Drive folders (since we haven't stress-tested the GDR for 100s of users at same time on same file): https://pypi.org/project/gdown/
#
# The only packages we'll need to install are Obspy and gdown. To do this, we would normally type something like "pip install obspy" at the command line (if running this on our local laptops). Here we'll need to run a bash (command-line) command in our Jupyter notebook on Colab, so we'll use a ! before the command. Below is the command to install obspy. As done in the FORGE notebook, you can also use a -q flag here if you don't want to see the output from the Obspy installation or gdown installation.
#
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 37027, "status": "ok", "timestamp": 1628862626119, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="5pkYNYW5IFPn" outputId="c4132a50-06d8-4dde-ce39-9fd6d30bc0e5"
# !pip install obspy gdown
# + [markdown] id="DWO7IR99KBxW"
# All packages need to be imported before they can be used. Now that we've installed Obspy, let's load all relevant packages.
#
# Note: If you decide to run this on your own local computer instead of Google Colab, you are likely to need to make some changes.
# + executionInfo={"elapsed": 1140, "status": "ok", "timestamp": 1628862627249, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="p4L9IsZ_KA5c"
# %matplotlib inline
import numpy as np
import scipy.fft as ft
import matplotlib.pyplot as plt
import obspy
import csv
import ipywidgets as widgets
# + [markdown] id="0Rbir7lFMMbY"
# # Reading Array Geometry
# + [markdown] id="1BdDeUSWMcqd"
# Grab the sensor array geometry information from the Geothermal Data Repository, which is stored in a csv file (comma separated value). File is at url: https://gdr.openei.org/files/829/Surface_DAS_DTS_UTM_coordinates.csv
#
# and more info is at https://openei.org/doe-opendata/dataset/brady-s-geothermal-field-das-and-dts-surface-and-borehole-array-metadata/resource/f0000003-58cc-4372-a567-000000000829
#
# Note that is a small Excel file (just has 3 numbers per channel stored), so we don't need to use gdown. You could download this csv file to your laptop and ready it like an Excel file should you wish.
# + executionInfo={"elapsed": 958, "status": "ok", "timestamp": 1628862628196, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="9vysLszqMTsA"
# Normally you would use the command below
# # !wget https://gdr.openei.org/files/829/Surface_DAS_DTS_UTM_coordinates.csv
# but we don't want to crash the GDR with 300+ people requesting the same file at once
# so here's a backup location and command on a public Google Drive folder:
# !wget -q --no-check-certificate 'https://docs.google.com/uc?export=download&id=1xl3diOA8eFHkPENqLEXgnynFpjcyORCn' -O Surface_DAS_DTS_UTM_coordinates.csv
# + [markdown] id="mTHJDcnjGabF"
# Read the geometry data from the CSV file with geometry information. Note that some parameters (noted below) were noted on the website.
# + executionInfo={"elapsed": 11, "status": "ok", "timestamp": 1628862628197, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="Y8o63bqkGVwP"
geomData = 'Surface_DAS_DTS_UTM_coordinates.csv'
# For this experiment, some of the geometry is in this csv file.
# if you look in the CSV file, these are the first and last rows with channels that have a location assigned
minRow = 53
maxRow = 8673
# read the geometry of all channels in the CSV file
with open(geomData) as myFile:
myReader = csv.reader(myFile, delimiter=',')
nRows = maxRow - minRow + 1
ch = np.zeros(nRows,dtype=np.int) # array to hold channel indices
x = np.zeros(nRows,dtype=np.float32) # array to hold x location
y = np.zeros(nRows,dtype=np.float32) # array to hold y location
myFile.seek(0)
rowCtr = 1
for row in myReader:
if((rowCtr >= minRow) and (rowCtr <= maxRow)):
ch[rowCtr-minRow] = int(row[0])
x[rowCtr-minRow] = float(row[1])
y[rowCtr-minRow] = float(row[2])
rowCtr = rowCtr + 1
# + [markdown] id="8bvc1fzQGu-K"
# In a later cell, we're going to use a data file with all sensors recording data while a single vibroseis seismic source is set off. The GDR website lists the location of this source. Let's calculate and plot the distance of each sensor from the source:
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 299} executionInfo={"elapsed": 1035, "status": "ok", "timestamp": 1628862629222, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="ujh_zq_mGk_4" outputId="782fbf7c-a52c-4e3f-abc2-6c50e72411bb"
# location of source from https://gdr.openei.org/submissions/849
# converted to UTM by https://www.latlong.net/lat-long-utm.html
srcX = 328552.49 # example source's x location in UTM meters
srcY = 4408006.65 # example source's y location in UTM meters
# calculate distances in meters of each sensor from the source location
distances = np.sqrt((x-srcX)**2 + (y-srcY)**2)
plt.plot(ch,distances)
plt.xlabel("channel index",fontsize=12)
plt.ylabel("distance (m)",fontsize=12)
plt.title("Distance of each sensor from example vibroseis source",fontsize=14)
plt.show()
# + [markdown] id="UftYpGL81esO"
# Plot the geometry of all channels. An easy way to do this is to do a scatter plot of all sensor locations, then color-code their markers by their corresponding channel index.
# + colab={"base_uri": "https://localhost:8080/", "height": 299} executionInfo={"elapsed": 31, "status": "ok", "timestamp": 1628862629224, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="BiZAn12kHZUY" outputId="42a80b90-60d8-4317-c6ff-26b04cb7d8eb"
plt.scatter(x,y,c=ch,linewidth=0,s=2,cmap=plt.get_cmap('jet'))
plt.colorbar()
plt.xlabel('x UTM (m)', fontsize=12)
plt.locator_params(axis='x', nbins=5) # reduce number of tick marks so labels aren't so crowded
plt.ylabel('y UTM (m)', fontsize=12)
plt.title('array geometry', fontsize=14)
plt.scatter(srcX,srcY,c='k',linewidth=0)
plt.show()
# + [markdown] id="IbluscYqI_AT"
# Grab the data acquired during a single vibroseis sweep from the Geothermal Data Repository (much larger file than geometry). Again, this will need to be done as a bash command with an !. This time we'll use the wget program to grab the data by its URL.
#
# More info at http://search.geothermaldata.org/dataset/98046653-40ef-4274-96e5-56bb8ae4dbde
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 22103, "status": "ok", "timestamp": 1628862651299, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="MCpqPJXxH53X" outputId="34063d91-7a40-4b33-e914-89fe0e7eb80f"
# Normally you would get data using the command below:
# # !wget https://gdr.openei.org/files/849/PoroTomo_iDAS16043_160325140048.sgy
# but we don't want to crash the GDR with a few hundred people requesting the
# same large file all at once, so may also instead use this command to access the data
# from a backup location in a public Google Drive folder:
# !gdown https://drive.google.com/uc?id=1hr-tMIyG7fXEi77XVBBa08GIAz6BxcjX -O PoroTomo_iDAS16043_160325140048.sgy
# + [markdown] id="xUGkIhDYMH-x"
# These data are SEG-Y formatted, which is a common format for active source exploration seismic data. Check the [SEG Wiki page](https://wiki.seg.org/wiki/SEG-Y#:~:text=The%20SEG%2DY%20(sometimes%20SEG,%2C%20a%20non%2Dprofit%20organization.) for more details.
# + executionInfo={"elapsed": 8968, "status": "ok", "timestamp": 1628862660264, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="VHzrcoHJIDWz"
chNumberFor0thTrace = -20 # just as a note- had to look in the csv file to know this (specific to this dataset's channel/geometry mapping info)
startTrNumber = minRow # for full array
endTrNumber = maxRow # for full array
nTr = endTrNumber-startTrNumber+1
# open up the SEGY file using Obspy
datafile = "PoroTomo_iDAS16043_160325140048.sgy"
# Read the file:
import obspy.io.segy.core
st = obspy.io.segy.core._read_segy(datafile, format='segy', unpack_trace_headers=True)
# Below is an alternative way to read:
#st = obspy.read(datafile,format='SEGY') # st is a stream and it has thousands of traces in it
# get header info from first channel (trace) of interest
stats = st[startTrNumber].stats
samplesPerSecond = stats.sampling_rate
secondsPerRecord = stats.npts*stats.delta
nSamples = stats.npts
# To try yourself: print out all of these stats and see what other information
# you can get about the dataset.
# create an empty 2D numpy array to store data
# (rows correspond to channel, columns correspond to time)
data = np.zeros((nTr,nSamples))
# read data from all channels and store in the numpy array
for tr in range(startTrNumber,endTrNumber+1):
data[tr-startTrNumber,:] = st[tr].data
# + [markdown] id="ilyu48IKBfSM"
# Just to be clear about dimensions (since we needed to do some offsets), let's print those out:
#
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 20, "status": "ok", "timestamp": 1628862660264, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="dXDE2MpFBewa" outputId="2243afd8-8a1e-44d6-f2cb-79ef1139fcda"
print("x has "+str(x.size)+" entries")
print("y has "+str(y.size)+" entries")
print("ch has "+str(ch.size)+" entries")
print("data array dimensions are: "+str(data.shape))
# + [markdown] id="VItFYYq4Yh_4"
# Let's visualize one of these channels (traces) with a wiggle plot:
# + colab={"base_uri": "https://localhost:8080/", "height": 254} executionInfo={"elapsed": 498, "status": "ok", "timestamp": 1628862660746, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="yTlvRJ46Yqc6" outputId="5c38a1c3-4867-4d98-ac85-fcfaecbc05c8"
st[2000].plot();
# + [markdown] id="BzihaAU3KgvM"
# However, visualizing wiggles from a stream with thousands of traces would be difficult: crowded looking, and taking a long time to generate. Often in array seismology, we use raster plots rather than wiggles to visualize many traces together. Define a function to plot an array of data from minCh to maxCh, and from minSec seconds to maxSec seconds with a title on the plot.
#
# Here we'll use the seismic colormap (red - white - blue), with one pixel colored to indicate the strain rate at each time sample and channel. Other colormaps can be used for a variety of visualizations: https://matplotlib.org/3.1.0/tutorials/colors/colormaps.html
#
#
# PURPOSE:
# This function plots a subset of someData from channel minCh to maxCh and minSec to maxSec seconds of time.
#
# INPUTS:
# * someData: 2D numpy array, storing seismic data such that each row represents a channel, and columns represent time (i.e. 0th axis is channel, 1st axis is time)
# * minSec: float, minimum time of interest (expressed in seconds beyond the start of someData)
# * maxSec: float, maximum time of interest (expressed in seconds beyond the start of someData)
# * minCh: int, minimum channel index of interest (don't forget offset of 0th channel if needed, as in BHS data)
# * maxCh: int, maximum channel index of interest (don't forget offset of 0th channel if needed, as in BHS data)
# * title: string, will be the title of your plot
# * sampleRate: a float indicating number of samples per second (of each sensor)
# + executionInfo={"elapsed": 15, "status": "ok", "timestamp": 1628862660747, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="QBipOtgaJGjX"
def plotSpaceTime(someData,minSec,maxSec,minCh,maxCh,title,sampleRate):
# Basic error checking
if (minSec >= maxSec) or (minSec < 0) or (maxSec*sampleRate > someData.shape[1]):
print("ERROR in plotSpaceTime inputs minSec: "+str(minSec)+" or maxSec: "+str(maxSec))
return
if (minCh >= maxCh) or (minCh < 0) or (maxCh > someData.shape[0]):
print("Error in plotSpaceTime inputs minCh: "+str(minCh)+" or maxCh: "+str(maxCh)+" referring to array with "+str(someData.shape[0])+" channels.")
return
# turn time range (in seconds) to indices
minSecID = int(minSec*sampleRate)
maxSecID = int(maxSec*sampleRate)
# make the plot
plt.figure()
plt.imshow(someData[minCh:maxCh,minSecID:maxSecID],aspect='auto',interpolation='none',cmap='seismic',extent=(minSec,maxSec,maxCh,minCh))
plt.xlabel('time (s)',fontsize=12)
plt.ylabel('channel',fontsize=12)
plt.title(title,fontsize=14)
plt.colorbar()
# + [markdown] id="wCJFaryshvR-"
# Now call this for the vibroseis sweep record.
# + colab={"base_uri": "https://localhost:8080/", "height": 299} executionInfo={"elapsed": 8971, "status": "ok", "timestamp": 1628862669703, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="8H37RKIbhLq_" outputId="5ff161eb-e475-4e29-bed7-4ca568dd9d2c"
title='response to first vibroseis sweep'
beginningCh = 0
endingCh = nTr
plotSpaceTime(data, 0, secondsPerRecord, beginningCh, endingCh, title, samplesPerSecond) # call the actual plotting function
# + [markdown] id="UVriBvtuLBeO"
# Write a function to plot a snapshot of the wavefield in time with a colored marker at each channel location. Each marker color should indicate the strain rate at the particular time and location.
#
# INPUTS:
# * xLocs: 1D numpy array of floats, specifying x locations of channels in meters
# * yLocs: 1D numpy array of floats, specifying y locations of channels in meters, should have same number of entries as xLocs
# * dataChT: 2D numpy array of floats containing data organized so each row is a channel and columns indicate time
# * timeInSeconds: float, indicating number of seconds beyond the start time of dataChT that you want to visualize the snapshot of data at
# * sampleRate: float, number of samples per second acquired by each channel
# * subsample: a positive integer, indicates the stride across channels if only a subset of channels will be visualized. For instance subsample=2 would take every other channel/row starting with the 0th index in dataChT. If = 1 (which is the default) no spatial subsampling is done.
# * clip: float, a nonegative number that sets the maximum value in the colorbar. By default this is 0 which would then be a flag to set the max positive value in the colorscale to the maximum absolute value throughout dataChT.
#
# No outputs, just generates a figure.
# + executionInfo={"elapsed": 29, "status": "ok", "timestamp": 1628862669704, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="Sk3AdkrzLJdR"
def plotTimeSnapshot(xLocs, yLocs, dataChT, timeInSeconds, sampleRate, subsample=1, clip=0):
# Basic error checking
if (xLocs.size != yLocs.size) or (xLocs.size != dataChT.shape[0]):
print("ERROR in plotTimeSnapshot inputs. Dimensions of xLocs, yLocs and rows of dataChT must match.")
return
maxTimeAvailable = float(dataChT.shape[1])/sampleRate # maximum time represented in dataChT (seconds)
if (timeInSeconds < 0) or (timeInSeconds > maxTimeAvailable):
print("ERROR in plotTimeSnapshot inputs. timeInSeconds "+str(timeInSeconds)+" is not represented in dataChT.")
return
# time sample of interest
timeID = int(timeInSeconds*sampleRate)
# default color clipping behavior is to just set the max color value to the max absolute value in this snapshot
if(clip == 0):
clip = np.max(np.absolute(dataChT[:,timeID]))
plt.figure()
plt.scatter(xLocs[::subsample],yLocs[::subsample],c=dataChT[::subsample,timeID],linewidth=0,s=2,cmap='seismic',vmin=-clip,vmax=clip)
plt.colorbar()
plt.xlabel('x UTM (m)')
plt.locator_params(axis='x', nbins=5) # reduce number of tick marks so labels aren't so crowded
plt.ylabel('y UTM (m)')
plt.title('signal at '+str(timeInSeconds)+' seconds',fontsize=14)
# + [markdown] id="qr4l5PGbqU4p"
# Plot a snapshot of the wavefield at a particular time, 3.5 seconds for example.
# + colab={"base_uri": "https://localhost:8080/", "height": 296} executionInfo={"elapsed": 479, "status": "ok", "timestamp": 1628862670155, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="GPYawOxhnf54" outputId="32d271ca-9eca-4e95-a9bf-4c769e780aa7"
time = 3.5 # time of intereset (in seconds)
plotTimeSnapshot(x, y, data, time, samplesPerSecond)
# + [markdown] id="qaL5uNc6qdfs"
# We can use a widget to easily change the time of the snapshot. As you move the widget to different times, look back at the raster plot above showing all channels. Does this make sense with what you're seeing there?
#
# Note: it does take a little time to regenerate each one, so if you're impatient you may want to set subsample to an integer > 1 (so for instance is subsample = 10, then only every 10th channel will be plotted).
# + colab={"base_uri": "https://localhost:8080/", "height": 328, "referenced_widgets": ["3b0387dc0bb94d4cb9511cbc7ff75ad5", "<KEY>", "9ec930c61e564cbab452519fbfe7017e", "191b0331451a43fcb936b381fee0f519", "0b0afe59223540ca9e13a47cc5d195d7", "cbbb6ea81a1343779c561d571d83a248", "95e10ec9ad0d4dd383d0f2bc596ffec6"]} executionInfo={"elapsed": 5805, "status": "ok", "timestamp": 1628862675948, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="IrZA9KAkLZm9" outputId="156a6312-4428-4eb2-95f4-ba41f7c09b13"
# set a single color clip for all time snapshots (so they're all on the same colorscale)
overallClip = np.percentile(np.absolute(data),99) # take the 99th %tile of absolute values of whole data window as max
def interactiveSnapshot(timeInSec):
# This is a wrapper around plotTimeSnapshot so that the widget just depends on the
# time of interest in seconds. This has no subsampling (so 2nd to last arg is 1).
plotTimeSnapshot(x, y, data, timeInSec, samplesPerSecond, 1, overallClip)
# has a widget that lets you check the time snapshot at 1, 1.5, 2, 2.5, ..., 24.5, 25 seconds
interactive_plot = widgets.interactive(interactiveSnapshot, timeInSec=(1,25,0.5))
interactive_plot
# + [markdown] id="55mJ9Sp56jKH"
# ## Effect of sensor orientation on response
# + [markdown] id="erMyFIhP6DTp"
# Let's zoom in on just channels 1500-2500 during 7-8 seconds while the vibroseis sweep is happening. These channels make a few tight turns, so they'll let us see how response varies with geometry. First check out this subset of the geometry.
# + colab={"base_uri": "https://localhost:8080/", "height": 295} executionInfo={"elapsed": 674, "status": "ok", "timestamp": 1628862676619, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="1jSnv4ilAgbL" outputId="bbe6f384-ee2c-4930-cd3d-f31c7c414670"
startCh = 1500
endCh = 2500
startSec = 7
endSec=8
plt.scatter(x[startCh:endCh],y[startCh:endCh],c=ch[startCh:endCh],linewidth=0,s=2,cmap=plt.get_cmap('jet'))
plt.colorbar()
plt.xlabel('x UTM (m)')
plt.locator_params(axis='x', nbins=5)
plt.ylabel('y UTM (m)')
plt.title('array geometry')
plt.scatter(srcX,srcY,c='k',linewidth=0) # plot the source location
plt.axis('equal')
plt.show()
# + [markdown] id="MQx5TNQtA7VM"
# Now actually plot the data. Investigate where the corners of the array line up with the data, and the direction of the waves moving outwards from the source. What do you notice?
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 299} executionInfo={"elapsed": 709, "status": "ok", "timestamp": 1628862677314, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="-_oI2YQd6qAw" outputId="ad27e327-3ec6-4343-f64c-5296be7abf54"
title='vibroseis sweep - zoom in'
plotSpaceTime(data, startSec, endSec, startCh, endCh, title, samplesPerSecond) # call the actual plotting function
# + [markdown] id="ihVnp6utCQq5"
# **Array geometry questions for small group investigation:**
#
# 1. Look at the last segment. How would waves from that source location be hitting it? What do you notice about the amplitudes?
# 2. Change your visualization codes in the cell above and below so you can zoom in on much smaller subsets of channels. Can you figure out where corners are based on the waveforms?
# 3. Why is the amplitude from ~1750 to ~1900 so much lower than 1500 to ~1650?
# 4. Try zooming in on different subsets of the array. What do you see about their response?
# + [markdown] id="2q_E3IXoR-nR"
# ## Spectral Analysis
# + [markdown] id="OmIDQMwICURG"
# Often we can understand some of our sensor response by looking at space-time domain data, but we can learn more by also looking at the spectrum of our data in the frequency domain.
#
# If you aren't familiar with the Fourier Transform, this video by <NAME> is a good starting place: https://www.youtube.com/watch?v=1JnayXHhjlg
#
# Here, we'll use the Scipy Fourier Transform package: https://docs.scipy.org/doc/scipy/reference/fft.html#module-scipy.fft
# which has multiple implementations based on the Fast Fourier Transform (FFT) algorithm.
#
# For visualizing how much wave energy is being detected at each frequency, we often look at the amplitude spectrum (magnitude of the complex-valued Fourier transform at each frequency), and sometimes at the power spectrum (squared magnitude of the complex-valued Fourier transform at each frequency).
#
# In the cell below, we define a function to calculate the power spectrum of a single channel's recording.
# + executionInfo={"elapsed": 14, "status": "ok", "timestamp": 1628862677315, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="AFHSy4IuDP6c"
def plotSpecOneCh(chOfInterest,dataArray,sampleRate,title):
# Basic error checking on inputs
if(chOfInterest >= dataArray.shape[0]):
print("ERROR in plotSpecOneCh inputs. Channel requested "+str(chOfInterest)+" not available in data with "+str(dataArray.shape[0])+" channels ")
return
spectrum = ft.fft(dataArray[chOfInterest,:])
frqBins = int(spectrum.size/2)
# frequencies of interest
NyquistFrq = sampleRate/2.0 # the Nyquist frequency
frqs = np.linspace(0,NyquistFrq,num=frqBins)
plt.figure()
plt.plot(frqs,np.absolute(spectrum[:frqBins]))
plt.xlabel('Frequency (Hz)',fontsize=12)
plt.ylabel('Amplitude',fontsize=12)
plt.title(title)
# + [markdown] id="oFkitgMXAi4F"
# Let's plot this for one particular channel of interest, 2300.
# + colab={"base_uri": "https://localhost:8080/", "height": 298} executionInfo={"elapsed": 13, "status": "ok", "timestamp": 1628862677315, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="4TJHnmURFZ1N" outputId="2b8e9525-66a3-44ea-e40b-efd3f09de379"
channel = 2300
plotSpecOneCh(channel,data,samplesPerSecond,'Spectrum channel '+str(channel))
# + [markdown] id="WNDU6VLYBDhp"
# Define a function that will plot the amplitude spectrum of each channel (amplitude indicated by color) within a certain time window and frequency range.
#
#
# INPUTS:
# * dataArray: 2D numpy array of space-time domain seismic data, rows represent channels, columns represent time samples
# * minSec: float, minimum time (seconds) from the start of the dataArray of interest
# * maxSec: float, maximum time (seconds) from the start of the dataArray of interest
# * minCh: int, minimum channel index of interest (don't forget offset of 0th channel if needed, as in BHS data)
# * maxCh: int, maximum channel index of interest (don't forget offset of 0th channel if needed, as in BHS data)
# * minFrq: float, minimum frequency of interest (Hz)
# * maxFrq: float, maximum frequency of interest (Hz)
# * sampleRate: float, number of samples per second acquired by each sensor
# * title: string, what you'd like as your plot title
# * scale: string, keyword of either 'linear' or 'log10' specifying whether the plot is colored based on the amplitudes or the log_10 of the amplitudes.
# + executionInfo={"elapsed": 13, "status": "ok", "timestamp": 1628862677316, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="p772gD0hCaR0"
def plotArraySpec(dataArray, minSec, maxSec, minCh, maxCh, minFrq, maxFrq, sampleRate, title, scale='linear'):
# check for valid inputs (note- these checks aren't exhaustive)
if(maxFrq <= minFrq):
print("ERROR in plotArraySpec inputs: minFrq "+str(minFrq)+" >= maxFrq "+str(maxFrq))
return
if maxSec <= minSec:
print("ERROR in plotArraySpec inputs: minSec "+str(minSec)+" >= maxSec "+str(maxSec))
return
if (scale != 'linear') and (scale != 'log10'): # only 2 scale keywords are recognized
print("ERROR in plotArraySpec inputs: scale keyword "+scale+" is not recognized")
# figure out sample indices for time window of interest
startTimeIdx = int(minSec*sampleRate)
endTimeIdx = int(maxSec*sampleRate)
if endTimeIdx > dataArray.shape[1]: # another opportunity for error checking: don't request a time bigger than what's available.
print("ERROR in plotArraySpec inputs: maxSec "+str(maxSec)+" exceeds last time in dataArray")
# calculate the amplitude spectrum (not amplitude symmetry for +/- frequencies)
spectrum = ft.fft(dataArray[minCh:maxCh+1,startTimeIdx:endTimeIdx+1],axis=-1)
nFrqBins = int(spectrum.shape[1]/2) # number of frequency bins
amplitudeSpec =np.absolute(spectrum[:,:nFrqBins])
# calculate indices corresponding to the frequencies of interest
NyquistFrq = sampleRate/2.0 # the Nyquist frequency
# make sure maxFrq doesn't exceed Nyquist frequency
if(maxFrq > NyquistFrq):
print("ERROR in plotArraySpec inputs: maxFrq "+str(maxFrq)+" >= Nyquist frequency "+str(NyquistFrq)+" indicated by sampleRate "+str(sampleRate))
return
# convert frequencies to an index in the array
HzPerBin = NyquistFrq/float(nFrqBins)
minFrqIdx = int(minFrq/HzPerBin)
maxFrqIdx = int(maxFrq/HzPerBin)
frqs = np.linspace(minFrqIdx*HzPerBin,maxFrqIdx*HzPerBin,num=(maxFrqIdx-minFrqIdx+1))
# actually do the plot
plt.figure()
if scale == 'linear':
plt.imshow(amplitudeSpec[:,minFrqIdx:maxFrqIdx],aspect='auto',interpolation='none',cmap='inferno',extent=(minFrq,maxFrq,maxCh,minCh))
elif scale == 'log10':
plt.imshow(np.log10(amplitudeSpec[:,minFrqIdx:maxFrqIdx]),aspect='auto',interpolation='none',cmap='inferno',extent=(minFrq,maxFrq,maxCh,minCh))
plt.xlabel('Frequency (Hz)')
plt.ylabel('Channel')
plt.colorbar()
plt.title(title)
# + [markdown] id="WxX709qmCdmr"
# Let's first look at all channels' amplitude spectra up to a few hundred Hz for the entire sweep (say 0 to 25 seconds):
# + colab={"base_uri": "https://localhost:8080/", "height": 295} executionInfo={"elapsed": 28026, "status": "ok", "timestamp": 1628862705329, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="W0-XRhv-BAq-" outputId="0fd5c014-f41c-4232-9c7b-85d7bd9d8e4f"
# visualize same channels as you did for whole time domain plot above
# choose frequency range of interest
minFrq = 0.5
maxFrq = 200
# choose which time window to investigate
startSec = 0
endSec = 25
# call the function to plot
plotArraySpec(data, startSec, endSec, beginningCh, endingCh, minFrq, maxFrq, samplesPerSecond, 'Array-wide log amplitude spectrum','log10')
# + [markdown] id="AsN4llXHLJbe"
# ***Spectral analysis questions for small group investigation:***
#
# Add some cells to look at different subsets of the frequency content, subsets in time, and in space. Investigate some of the following questions:
# 1. During this vibroseis sweep how does the frequency content change over time?
# 2. How does the frequency content differ on channels that are closer to the source? Farther from the source? Would there be way to normalize energy between near and far channels to bring out these changes in frequency content?
# 3. Does the geometry matter in the frequency content acquired on a channel?
# 4. How much does using the linear or log plot matter in finding spectral variability in space/time? Does the colormap matter in your ability to visualize these changes? Investigate this by changing the cmap parameter, and changing log or linear (maybe even add it as a parameter in the function call).
# 5. Within a single channel, we often use a "waterfall plot" that shows the spectral amplitude after calculating the short time Fourier transform in many shorter successive time windows. Try to code this up and use it to investigate changes over time on different channels.
# + executionInfo={"elapsed": 25, "status": "ok", "timestamp": 1628862705329, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="YWpYTUSwR7ab"
# + [markdown] id="wDSPXKXshR0Q"
# ## Removing source signature
# + [markdown] id="-YrTjRqUn-s-"
# In seismic exploration, a common method of removing the signature of the vibroseis sweep is to cross-correlate the known sweep with the recorded data. The resulting data is, ideally, the autocorrelation of a sweep, which is known as a Klauder wavelet. While it is not fully deconvolved yet, it is much easier to understand.
# + executionInfo={"elapsed": 25680, "status": "ok", "timestamp": 1628862730985, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="USPX-ep-hWde"
import scipy
startCh = 1500
endCh = 2500
nch,nt = data.shape
swp_time = 20.0
swp_f1 = 5.0
swp_f2 = 80.0
sweep = scipy.signal.chirp(np.linspace(0.0,swp_time,int(swp_time*samplesPerSecond)), swp_f1, 20, swp_f2, method='linear')
sweep = np.flip(sweep)
corr_data = np.zeros(shape=data.shape)
for chind in range(nch):
corr_data[chind,:] = scipy.signal.convolve(data[chind,:],sweep,mode='same')
# + colab={"base_uri": "https://localhost:8080/", "height": 299} executionInfo={"elapsed": 820, "status": "ok", "timestamp": 1628862731792, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="zDG6rz6Gn2Ms" outputId="f918e27a-a31a-4cc9-ec21-f6d9714ba3fa"
startCh = 1500
endCh = 2500
title='vibroseis sweep - zoom in, before correlation'
plotSpaceTime(data, 11.7, 13, startCh, endCh, title, samplesPerSecond) # call the actual plotting function
# + colab={"base_uri": "https://localhost:8080/", "height": 299} executionInfo={"elapsed": 750, "status": "ok", "timestamp": 1628862732528, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12798709312997120137"}, "user_tz": -180} id="6cCt-XDouVDu" outputId="7f05dd49-fb9d-40c0-9ed8-5856d53855f5"
title='vibroseis sweep - zoom in, after correlation'
plotSpaceTime(corr_data, 11.7, 13, startCh, endCh, title, samplesPerSecond) # call the actual plotting function
|
.ipynb_checkpoints/BradyHotSprings-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <h2> Assignment 2</h2>
#
# <p>This time, you're going to attempt to load your first csv dataset! Open up the starter code located in Module2/assignment2.py. Read through it and follow the directions to: </p>
#
#
# <ol>
# <li> Load up Module2/Datasets/tutorial.csv </li>
# <li> Print the entire dataframe, using print df </li>
# <li> Use the .describe() method on the dataset </li>
# <li> Slice the dataset using [2:4, 'col3'] </li>
# </ol>
#
# +
# # %load 'assignment2.py'
import pandas as pd
# TODO: Load up the 'tutorial.csv' dataset
#
# .. your code here ..
# TODO: Print the results of the .describe() method
#
# .. your code here ..
# TODO: Figure out which indexing method you need to
# use in order to index your dataframe with: [2:4,'col3']
# And print the results
#
# .. your code here ..
# -
import pandas as pd
data = pd.read_csv('Datasets/tutorial.csv')
# <h4> Question 1 </h4>
# <p> When you print the results of calling .describe() on your dataframe, what is the value displayed in the bottom right corner (col3 max)? </p>
data.describe()
# <h4> Question 2 </h4>
#
# <p> Which of the many indexing methods did you use to get [2:4,'col3'] working?</p>
#
# <h4> Question 3 </h4>
#
# <p> How many values are returned when you print the results of the [2:4,'col3'] indexing operation? </p>
data.loc[2:4,'col3']
# <h2> Assignment 3</h2>
#
# <p> MIT's Karl Ulrich donated a dataset titled Servo Data Set to the UCI Machine Learning Repository in the 1980's. The dataset has been described as "an interesting collection of data that covers an extremely non-linear phenomenon - predicting the rise time of a servomechanism in terms of two (continuous) gain settings and two (discrete) choices of mechanical linkages."</p>
#
# <p> As noted on the dataset website above, the column names are defined in order as:</p>
#
# <p> ['motor', 'screw', 'pgain', 'vgain', 'class'] </p>
#
# <p> Your mission, should you choose to accept, is to figure out a few stats about this dataset, which has been conveniently copied to your Module2/Datasets/servo.data. You can get started by opening up the assignment starter code, saved to Module2/assignment3.py.</p>
# +
# # %load 'assignment3.py'
import pandas as pd
# TODO: Load up the dataset
# Ensuring you set the appropriate header column names
#
# .. your code here ..
# TODO: Create a slice that contains all entries
# having a vgain equal to 5. Then print the
# length of (# of samples in) that slice:
#
# .. your code here ..
# TODO: Create a slice that contains all entries
# having a motor equal to E and screw equal
# to E. Then print the length of (# of
# samples in) that slice:
#
# .. your code here ..
# TODO: Create a slice that contains all entries
# having a pgain equal to 4. Use one of the
# various methods of finding the mean vgain
# value for the samples in that slice. Once
# you've found it, print it:
#
# .. your code here ..
# TODO: (Bonus) See what happens when you run
# the .dtypes method on your dataframe!
# -
import pandas as pd
# +
data = pd.read_csv('Datasets/servo.data', header = None)
# +
headers = ['motor', 'screw', 'pgain', 'vgain', 'class']
data.columns = headers
data.head()
# -
# <h4> Question 1a </h4>
#
# <p> How many samples in this dataset have a vgain feature value equal to 5?</p>
t = data[data["vgain"] == 5]
len(t)
# <h4> Question 1b </h4>
#
# <p> How many samples in this dataset contain the value E for both motor and screw features? Be sure to validate you've correctly loaded your data before answering!</p>
t = data[(data["motor"] == "E") & (data["screw"] == "E")]
t.head()
len(t)
# <h4> Question 1c </h4>
#
# <p> What is the mean vgain value of those samples that have a pgain feature value equal to 4?</p>
t = data[data["pgain"] == 4]
t.describe()
t.head()
# <h2> Assignment 4 </h2>
#
# <p> Navigate over to ESPN's website for NHL Historic Player Points Statistics, for the years 2014-2015. This page has a table on it with a few stats we're interested in obtaining. But it's a bit messy! Clean it up for us, using the appropriate commands to: </p>
#
# <ol>
#
# <li> Load up the table on just this page into a Pandas dataframe. No need to worry about the other pages! </li>
# <li> Rename the columns so that they match the column definitions on the website. </li>
# <li> Get rid of (drop) any erroneous rows that has at least 4 NANs in them. </li>
# <li> Get rid of the RK column. </li>
# <li> Ensure there are no nan "holes" in your index. </li>
# <li> Check the dtypes of all columns, and ensure those that should be numeric are numeric. </li>
#
# </ol>
#
# +
# # %load 'assignment4.py'
import pandas as pd
# TODO: Load up the table, and extract the dataset
# out of it. If you're having issues with this, look
# carefully at the sample code provided in the reading
#
# .. your code here ..
# TODO: Rename the columns so that they are similar to the
# column definitions provided to you on the website.
# Be careful and don't accidentially use any names twice.
#
# .. your code here ..
# TODO: Get rid of any row that has at least 4 NANs in it,
# e.g. that do not contain player points statistics
#
# .. your code here ..
# TODO: At this point, look through your dataset by printing
# it. There probably still are some erroneous rows in there.
# What indexing command(s) can you use to select all rows
# EXCEPT those rows?
#
# .. your code here ..
# TODO: Get rid of the 'RK' column
#
# .. your code here ..
# TODO: Ensure there are no holes in your index by resetting
# it. By the way, don't store the original index
#
# .. your code here ..
# TODO: Check the data type of all columns, and ensure those
# that should be numeric are numeric
#
# .. your code here ..
# TODO: Your dataframe is now ready! Use the appropriate
# commands to answer the questions on the course lab page.
#
# .. your code here ..
# -
import pandas as pd
# <h4> Question 1 </h4>
#
# <p> After completing the 6 steps above, how many rows remain in this dataset? (Not to be confused with the index!) </p>
data = pd.read_html?
data = pd.read_html
url = "http://www.espn.com/nhl/statistics/player/_/stat/points/sort/points/year/2015/seasontype/2"
dataTables = pd.read_html(url, skiprows = 2)
len(dataTables)
# +
data = dataTables[0]
data.head()
# +
data.columns = ["x_x", "player", "team", "gp", "g", "a", "pts", "plusminus", "pim", "pts_g", "sog", "pct", "gwg",
"pp_g", "pp_a", "sh_g", "sh_a"]
data = data.drop(labels = ["x_x"], axis = 1)
# -
data.head()
print len(data)
data = data.dropna(axis = 0, thresh = 4)
print len(data)
data = data.reset_index(drop= True)
# +
print data.dtypes
for i in list(data.columns)[2:]:
data[i] = pd.to_numeric(data[i], errors="coerce")
print data.dtypes
# +
print len(data)
data = data.dropna(axis = 0, thresh = 4)
print len(data)
data = data.reset_index(drop= True)
# -
# <h4> Question 2 </h4>
#
# <p> How many unique PCT values exist in the table? </p>
len(set(data["pct"]))
# <h4> Question 3 </h4>
#
# <p> What is the value you get by adding the GP values at indices 15 and 16 of this table?</p>
print type(data.loc[15:16, 'gp'])
data.loc[15:16, 'gp']
sum(data.loc[15:16, 'gp'])
# <h2> Assignment 5 </h2>
#
# <p> <NAME> extracted a reasonably clean subset of the 1994, U.S. Census database, with a goal of running predictions to determine whether a person makes over 50K a year. The dataset is hosted on the University of California, Irvine's Machine Learning Repository and includes features such as the person's age, occupation, and hours worked per week, etc.</p>
#
# <p> As clean as the data is, it still isn't quite ready for analysis by SciKit-Learn! Using what you've learned in this chapter, clean up the various columns by encode them properly using the best practices so that they're ready to be examined. We've included a modifies subset of the dataset at Module2/Datasets/census.data and also have some started code to get you going located at Module2/assignment5.py.</p>
#
#
# <ol>
# <li> Load up the dataset and set header label names to:
#
# ['education', 'age', 'capital-gain', 'race', 'capital-loss', 'hours-per-week', 'sex', 'classification'] </li>
#
#
# <li> Ensure you use the right command to do this, as there is more than one command! To verify you used the correct one, open the dataset in a text editor like SublimeText or Notepad, and double check your df.head() to ensure the first values match up. </li>
# <li> Make sure any value that needs to be replaced with a NAN is set as such. There are at least three ways to do this. One is much easier than the other two. </li>
# <li> Look through the dataset and ensure all of your columns have appropriate data types. Numeric columns should be float64 or int64, and textual columns should be object. </li>
# <li> Properly encode any ordinal features using the method discussed in the chapter. </li>
# <li> Properly encode any nominal features by exploding them out into new, separate, boolean features. </li>
#
# </ol>
#
# +
# # %load 'assignment5.py'
import pandas as pd
import numpy as np
#
# TODO:
# Load up the dataset, setting correct header labels.
#
# .. your code here ..
#
# TODO:
# Use basic pandas commands to look through the dataset... get a
# feel for it before proceeding! Do the data-types of each column
# reflect the values you see when you look through the data using
# a text editor / spread sheet program? If you see 'object' where
# you expect to see 'int32' / 'float64', that is a good indicator
# that there is probably a string or missing value in a column.
# use `your_data_frame['your_column'].unique()` to see the unique
# values of each column and identify the rogue values. If these
# should be represented as nans, you can convert them using
# na_values when loading the dataframe.
#
# .. your code here ..
#
# TODO:
# Look through your data and identify any potential categorical
# features. Ensure you properly encode any ordinal and nominal
# types using the methods discussed in the chapter.
#
# Be careful! Some features can be represented as either categorical
# or continuous (numerical). If you ever get confused, think to yourself
# what makes more sense generally---to represent such features with a
# continuous numeric type... or a series of categories?
#
# .. your code here ..
#
# TODO:
# Print out your dataframe
#
# .. your code here ..
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# %matplotlib inline
# -
data = pd.read_csv('Datasets/census.data', header = None, usecols = list(range(1,9)))
data.head()
len(data)
# +
cols = ['education', 'age', 'capital-gain', 'race', 'capital-loss', 'hours-per-week', 'sex', 'classification']
data.columns = cols
data.head()
# -
data.dtypes
# +
#for i in ['education', 'race', 'sex', 'classification']:
#data[i].value_counts().plot(kind = 'bar')
data['education'].value_counts().plot(kind = 'bar')
# +
data['capital-gain'] = pd.to_numeric(data['capital-gain'], errors = 'coerce')
data.dtypes
# -
data = pd.get_dummies(data, columns = ['race', 'sex'])
data.head()
for i in ['education', 'hours-per-week']:
for x in list(set(data[i])):
print x
print ''
# +
orderedEducation = ["Preschool", "1st-4th", "5th-6th", "7th-8th", "9th", "10th", "11th", "12th", "HS-grad",
"Some-college", "Bachelors", "Masters", "Doctorate"]
data['education'] = data['education'].astype("category",
ordered = True, categories = orderedEducation).cat.codes
data.head()
# -
|
Module2/02b - Data and Features Lab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import os.path as osp
from tqdm.autonotebook import tqdm
import torch
import torch.nn.functional as F
from torch.nn import Sequential, Linear, ReLU, GRU
import torch_geometric.transforms as T
from torch_geometric.data import DataLoader
# -
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split, GroupShuffleSplit
from torch.utils.data import Subset
from kaggle_champs import constants
# # Load and preprocessing data
# ## Load data
script_dir = os.path.abspath(os.path.dirname(__file__))
train = pd.read_csv(script_dir + '/../../data/train.csv')
test = pd.read_csv(script_dir + '/../../data/test.csv')
struct_dir = script_dir + "/../../data/xyz/"
y_mean = train.scalar_coupling_constant.mean()
y_std = train.scalar_coupling_constant.std()
np.log((train.scalar_coupling_constant - train.type.map(train.groupby('type').scalar_coupling_constant.mean())).abs().groupby(train.type).mean())
# ## Split train valid
molecules = train.molecule_name.drop_duplicates().sort_values()
train_ind, valid_ind = train_test_split(np.arange(len(molecules)),
test_size=5000,
random_state=1234)
assert not set(train_ind).intersection(valid_ind)
len(train_ind), len(valid_ind)
# ## Create train valid subet
# Check reproducibility
rs = np.random.RandomState(seed=1234)
print(rs.choice(train_ind, 10))
print(rs.choice(valid_ind, 10))
train_data = train.loc[train.molecule_name.isin(molecules.iloc[train_ind])]
val_data = train.loc[train.molecule_name.isin(molecules.iloc[valid_ind])]
# ## Create dataset
from kaggle_champs.dataset import ChampsDataset
# +
import os
import numpy as np
import openbabel
import torch
from torch_geometric.data import Data
from torch.utils.data import Dataset
from tqdm.autonotebook import tqdm
from kaggle_champs.dataset import mol_to_data_v2
# -
class MoleculeDataset(Dataset):
def __init__(self, metadata=None, base_dir=None, transform=None):
self.molecules = metadata.molecule_name.unique()
self.metadata = dict([
(ind, df) for ind, df in tqdm(metadata.groupby('molecule_name'))
])
self.base_dir = base_dir
self.transform = transform
self.conversion = openbabel.OBConversion()
self.conversion.SetInAndOutFormats("xyz", "mdl")
def __len__(self):
return len(self.metadata)
def __getitem__(self, index):
mol = openbabel.OBMol()
mol_name = self.molecules[index]
xyz_file = os.path.join(self.base_dir, f'{mol_name}.xyz')
if not os.path.exists(xyz_file):
raise FileNotFoundError(f'Expecting file {xyz_file} not found')
self.conversion.ReadFile(mol, xyz_file)
data = mol_to_data_v2(mol)
data.mol_ind = torch.tensor([[index]], dtype=torch.long)
data = self._add_targets(data, metadata=self.metadata[mol_name])
data.graph = nx.Graph()
data.graph.add_edges_from(data.edge_index.transpose(1,0).cpu().numpy())
if self.transform:
data = self.transform(data)
if hasattr(data, 'graph'):
del data.graph
return data
def _add_inverse_couple(self, couples):
inverse_direction = couples.rename(
{'atom_index_1': 'atom_index_0',
'atom_index_0': 'atom_index_1'},
axis=1)
couples = couples.append(
inverse_direction,
sort=False
)
couples = couples.sort_values(['atom_index_0',
'atom_index_1'])
return couples
def _add_y(self, data, couples):
if 'scalar_coupling_constant' in couples.columns:
data.y = torch.tensor(
couples['scalar_coupling_constant'].values,
dtype=torch.float).view(-1,1)
else:
data.y = torch.zeros((len(couples), 1), dtype=torch.float)
return data
def _add_targets(self, data, metadata):
couples = metadata.copy()
couples = self._add_inverse_couple(couples)
data.couples_ind = torch.tensor(
couples[['atom_index_0',
'atom_index_1']].values,
dtype=torch.long)
data = self._add_y(data, couples)
data.type = torch.tensor(
couples['type'].map(constants.TYPES_DICT).values,
dtype=torch.long)
data.sample_weight = torch.tensor(
couples['type'].map(constants.TYPES_WEIGHTS).values,
dtype=torch.float)
return data
from kaggle_champs.preprocessing import RandomRotation, AddVirtualEdges, AddEdgeDistanceAndDirection, SortTarget
import networkx as nx
class AddEdgeDistanceAndDirection:
def __init__(self, dist_noise=0., gauss_base_max=4, gauss_base_steps=20, keep=True):
self.dist_noise = dist_noise
self.gauss_base_max = gauss_base_max
self.gauss_base_steps = gauss_base_steps
self.keep = True
def __call__(self, data):
(row, col), pos, edge_attr = data.edge_index, data.pos, data.edge_attr
dist = torch.norm(pos[col] - pos[row], p=2, dim=-1).view(-1, 1)
if self.dist_noise > 0:
noise = 1 + torch.randn_like(dist, dtype=dist.dtype) * self.dist_noise
dist = dist * noise
direction = (pos[col] - pos[row]) / dist
if self.keep:
data.dist = dist
data.direction = direction
base = torch.linspace(self.gauss_base_max/self.gauss_base_steps,
self.gauss_base_max,
self.gauss_base_steps,
dtype=torch.float).view(1, -1) # shape 1xn for broadcasting
dist = torch.exp(-(dist - base) ** 2 / 0.5 ** 2)
edge_attr = edge_attr.view(-1, 1) if edge_attr.dim() == 1 else edge_attr
data.edge_attr = torch.cat(
[edge_attr,
dist.type_as(edge_attr),
direction.type_as(edge_attr)],
dim=-1)
return data
class AddBondLinks:
def __call__(self, data):
bonds_ind = data.bonds_edge_ind
bonds_from = bonds_ind.view(-1, 1).repeat(1, (len(bonds_ind))).view(-1)
bonds_to = bonds_ind.view(-1).repeat(1, len(data.bonds_edge_ind)).view(-1)
bonds_links = torch.stack([bonds_from, bonds_to], dim=1) # all couples, will filter
filter_correct_common_node = (data.edge_index[:, bonds_from][1] == data.edge_index[:, bonds_to][0])
filter_remove_self_loop = (data.edge_index[:, bonds_from][0] != data.edge_index[:, bonds_to][1])
data.bonds_links_edge_ind = bonds_links[filter_correct_common_node * filter_remove_self_loop]
return data
class AddCounts:
def __call__(self, data):
data.count_nodes = torch.tensor([[data.num_nodes]], dtype=torch.long)
data.count_edges = torch.tensor([[data.num_edges]], dtype=torch.long)
data.count_couples = torch.tensor([[data.couples_ind.size(0)]], dtype=torch.long)
return data
class AddGlobalAttr:
def __init__(self):
pass
def __call__(self, data):
data.global_attr = torch.zeros((1, 1), dtype=torch.float)
return data
class SortTarget:
def _get_index(self, data, row, col):
idx = row * (data.num_nodes-1) + col
idx[row < col] = idx[row < col] - 1
return idx
def __call__(self, data):
target = torch.zeros((data.num_edges, data.y.size()[1]), dtype=torch.float)
weights = torch.zeros((data.num_edges), dtype=torch.float)
mask = torch.zeros((data.num_edges), dtype=torch.bool)
types = torch.zeros((data.num_edges), dtype=torch.long)
row, col = data.couples_ind.transpose(1,0)
indexes = self._get_index(data, row, col)
mask[indexes] = True
weights[indexes] = data.sample_weight
target[indexes] = data.y
types[indexes] = data.type
#data.mask = mask
data.y = target[mask]
data.sample_weight = weights[mask]
data.type = types[mask]
assert torch.equal(data.couples_ind, data.edge_index[:, mask].transpose(1,0))
data.couples_edge_ind = torch.arange(data.num_edges, dtype=torch.long)[mask].view(-1,1)
return data
class AddBondPath:
def __call__(self, data):
# suffix _index to get node index adjustment
data.paths_index = self.find_paths(data).transpose(1,0)
data.paths_edge_ind = torch.cat(
[self._nodes_to_edge_ind(data, data.paths_index[i], data.paths_index[i+1]) for i in range(3)],
dim=1)
return data
def _nodes_to_edge_ind(self, data, node_from, node_to):
edge_ind = node_from * (data.num_nodes-1) + node_to
edge_ind[node_from < node_to] = edge_ind[node_from < node_to] - 1
return edge_ind.view(-1, 1)
def find_paths(self, data):
assert hasattr(data, 'couples_ind')
assert hasattr(data, 'graph')
all_paths = nx.shortest_path(data.graph)
paths = []
for (from_, to_) in data.couples_ind.numpy():
path = torch.tensor(all_paths[from_][to_], dtype=torch.long).view(-1,1)
paths.append(path)
paths = torch.nn.utils.rnn.pad_sequence(paths, batch_first=True).squeeze()
if paths.size(1) < 4:
paths = torch.nn.functional.pad(paths, (0, 4 - paths.size(1)))
return paths
class AddInverseCouples:
def find_inverse_couple_position(self, node_from, node_to):
df = pd.DataFrame({
'from': node_from,
'to': node_to,
}).reset_index()
inverse = df.rename({
'from': 'to',
'to': 'from',
},axis=1)
merged = pd.merge(df, inverse, on=['from', 'to'], suffixes=('', '_inverse'))
assert merged.shape[0] == df.shape[0]
return merged.sort_values('index').index_inverse.values
def __call__(self, data):
assert hasattr(data, 'couples_ind')
node_from, node_to = data.couples_ind[:,0].numpy(), data.couples_ind[:,1].numpy()
data.inverse_couple_ind = torch.tensor(self.find_inverse_couple_position(node_from, node_to), dtype=torch.long)
return data
def find_inverse_couple_position(self, node_from, node_to):
df = pd.DataFrame({
'from': node_from,
'to': node_to,
}).reset_index()
inverse = df.rename({
'from': 'to',
'to': 'from',
},axis=1)
merged = pd.merge(df, inverse, left_on=['from', 'to'], right_on=['to', 'from'], suffixes=('', '_inverse'))
assert merged.shape[0] == df.shape[0]
return merged.sort_values('index').index_inverse
def correct_batch_edge_ind(batch):
offset_edge_ind = torch.zeros_like(batch.count_edges)
offset_edge_ind[1:] = batch.count_edges[:-1].cumsum(dim=0)
for k in ['bonds_edge_ind', 'bonds_links_edge_ind', 'paths_edge_ind', 'couples_edge_ind']:
if hasattr(batch, k):
batch[k] = batch[k] + offset_edge_ind[batch[k+'_batch']]
return batch
def correct_inverse_couples_ind(batch):
offset = torch.zeros_like(batch.count_couples)
offset[1:] = batch.count_couples[:-1].cumsum(dim=0)
batch.inverse_couple_ind = batch.inverse_couple_ind + offset[batch.inverse_couple_ind_batch].view(-1)
assert torch.equal(batch.couples_edge_ind[batch.inverse_couple_ind][batch.inverse_couple_ind],
batch.couples_edge_ind)
return batch
# +
train_dataset = MoleculeDataset(metadata=train_data,
base_dir=struct_dir,
transform=T.Compose([
AddBondPath(),
AddVirtualEdges(),
RandomRotation(),
AddEdgeDistanceAndDirection(dist_noise=0.),
AddGlobalAttr(),
SortTarget(),
AddBondLinks(),
AddCounts(),
AddInverseCouples(),
]))
val_dataset = MoleculeDataset(metadata=val_data,
base_dir=struct_dir,
transform=T.Compose([
AddBondPath(),
AddVirtualEdges(),
AddEdgeDistanceAndDirection(dist_noise=0.),
AddGlobalAttr(),
SortTarget(),
AddBondLinks(),
AddCounts(),
AddInverseCouples(),
]))
# -
data = train_dataset[10]
data
torch.equal(data.inverse_couple_ind[data.inverse_couple_ind], torch.arange(86))
# # Model
from kaggle_champs.modelling import MegNetBlock, create_mlp_v2, MegNetBlock_v2, MegNetBlock_v3
from torch import nn
from torch_scatter import scatter_add
def gather_embedding(data, x_out, edge_out, u_out, couple_type):
n_bonds = int(couple_type[0])
couple_filter = (data.type == constants.TYPES_DICT[couple_type])
couples_edge_ind = data.couples_edge_ind.view(-1)
merged = [
u_out[data.batch[data.edge_index[0][couples_edge_ind][couple_filter]]],
]
if n_bonds > 1:
merged.append(edge_out[couples_edge_ind][couple_filter])
node_ind = data.paths_index.transpose(1,0)[:, :n_bonds+1][couple_filter] # convert_node_ind(data, 'paths')[:, :n_bonds+1]
for i in range(n_bonds+1):
merged.append(x_out[node_ind[:,i]])
for i in range(n_bonds):
edge_ind = data.paths_edge_ind[:,i] # convert_couple_to_edge_ind(data, data.paths_index[i], data.paths_index[i+1], data.paths_edge_ind_batch)
merged.append(edge_out[edge_ind[couple_filter]])
return torch.cat(merged, dim=1)
class OutputLayer_new(torch.nn.Module):
def __init__(self, rep_dim, dim, y_mean, y_std, couple_type):
super(OutputLayer_new, self).__init__()
self.scaling = torch.nn.Linear(1, 1)
self.scaling.bias = torch.nn.Parameter(torch.tensor(y_mean,
dtype=torch.float),
requires_grad=False)
self.scaling.weight = torch.nn.Parameter(torch.tensor(
[[y_std]], dtype=torch.float),
requires_grad=False)
self.couple_type = couple_type
n_bonds = int(couple_type[0])
if n_bonds == 1:
input_dim = dim * (n_bonds + (n_bonds + 1) + 1) # edges + nodes + u
else:
input_dim = dim * (n_bonds + (n_bonds + 1) + 2) # edges + nodes + u + direct edge
self.mlp = create_mlp_v2(
input_dim=input_dim,
output_dim=1,
hidden_dims=[input_dim//2, input_dim//2, input_dim//2],
normalization_cls=torch.nn.LayerNorm,
activation_cls=torch.nn.ELU,
dropout_cls=torch.nn.Dropout,
dropout_prob=0.
)
def forward(self, data, x_out, edge_out, u_out):
in_ = gather_embedding(data, x_out, edge_out, u_out, self.couple_type)
out = self.mlp(in_)
out = self.scaling(out)
return out
from torch_scatter import scatter_mean
class EdgeAgg(torch.nn.Module):
def __init__(self, dim=32):
super(EdgeAgg, self).__init__()
self.body_mlp = nn.Sequential(
create_mlp_v2(
input_dim=dim,
output_dim=dim*2,
hidden_dims=[dim*2],
normalization_cls=torch.nn.LayerNorm,
activation_cls=torch.nn.ELU,
dropout_cls=torch.nn.Dropout,
dropout_prob=0.),
nn.LayerNorm(dim*2)
)
self.value_out = nn.Linear(dim*2, dim)
self.gating = nn.Sequential(
nn.Linear(dim*2, dim),
nn.Sigmoid()
)
def forward(self, edge_out, edges_ind):
out = self.body_mlp(edge_out)
out = self.value_out(out) * self.gating(out)
result = scatter_add(out, edges_ind, dim=0)
return result
class MegNetBlock(torch.nn.Module):
def __init__(self, edge_dim, x_dim, u_dim, dim=32, layer_norm=False,
normalization_cls=None, activation_cls=nn.ReLU,
dropout_cls=nn.Dropout, dropout_prob=0., residual=True, pooling='mean'):
super(MegNetBlock, self).__init__()
self.dim = dim
self.residual = residual
self.pooling = pooling
if layer_norm:
normalization_cls = nn.LayerNorm
kwargs = dict(
normalization_cls=normalization_cls,
activation_cls=activation_cls,
dropout_cls=dropout_cls,
dropout_prob=dropout_prob)
self.edge_dense = create_mlp_v2(
input_dim=edge_dim, output_dim=dim, hidden_dims=[dim * 2], **kwargs)
self.edge_agg = EdgeAgg(dim=dim)
self.node_dense = create_mlp_v2(
input_dim=x_dim, output_dim=dim, hidden_dims=[dim * 2], **kwargs)
self.global_dense = create_mlp_v2(
input_dim=u_dim, output_dim=dim, hidden_dims=[dim * 2], **kwargs)
self.edge_msg = create_mlp_v2(
input_dim=dim * 4, output_dim=dim, hidden_dims=[dim*2, dim*2], **kwargs)
self.node_msg = create_mlp_v2(
input_dim=dim * 3, output_dim=dim, hidden_dims=[dim*2, dim*2], **kwargs)
self.global_msg = create_mlp_v2(
input_dim=dim * 3, output_dim=dim, hidden_dims=[dim*2, dim*2], **kwargs)
def edge_model(self, src, dest, edge_attr, u, batch):
# source, target: [E, F_x], where E is the number of edges.
# edge_attr: [E, F_e]
# u: [B, F_u], where B is the number of graphs.
# batch: [E] with max entry B - 1.
out = torch.cat([src, dest, edge_attr, u[batch]], 1)
out = self.edge_msg(out)
return out
def node_model(self, x, edge_index, edge_attr, u, batch):
# x: [N, F_x], where N is the number of nodes.
# edge_index: [2, E] with max entry N - 1.
# edge_attr: [E, F_e]
# u: [B, F_u]
# batch: [N] with max entry B - 1.
row, _ = edge_index
out = self.edge_agg(edge_attr, row)
out = torch.cat([out, x, u[batch]], dim=1)
out = self.node_msg(out)
return out
def global_model(self, x, edge_index, edge_attr, u, batch):
# x: [N, F_x], where N is the number of nodes.
# edge_index: [2, E] with max entry N - 1.
# edge_attr: [E, F_e]
# u: [B, F_u]
# batch: [N] with max entry B - 1.
row, _ = edge_index
edge_mean = scatter_mean(edge_attr, batch[row], dim=0)
out = torch.cat(
[u, scatter_mean(x, batch, dim=0), edge_mean], dim=1)
out = self.global_msg(out)
return out
def forward(self, x, edge_index, edge_attr, u, batch, first_block=False):
# first block
edge_out = self.edge_dense(edge_attr)
x_out = self.node_dense(x)
u_out = self.global_dense(u)
x_res_base = x_out if first_block else x
edge_res_base = edge_out if first_block else edge_attr
u_res_base = u_out if first_block else u
row, col = edge_index
edge_out = self.edge_model(x_out[row], x_out[col], edge_out, u_out,
batch[row])
if self.residual:
edge_out = edge_res_base + edge_out
x_out = self.node_model(x_out, edge_index, edge_out, u_out, batch)
if self.residual:
x_out = x_res_base + x_out
u_out = self.global_model(x_out, edge_index, edge_out, u_out, batch)
if self.residual:
u_out = u_res_base + u_out
return x_out, edge_out, u_out
class MultiplicativeGaussianNoise(torch.nn.Module):
def __init__(self, scale=0.):
super(MultiplicativeGaussianNoise, self).__init__()
self.scale = scale
def forward(self, x):
if not self.training:
return x
noise = 1 + torch.randn_like(x) * self.scale
return x * noise
class MegNetModel_new(torch.nn.Module):
def __init__(self,
edge_dim,
x_dim,
u_dim,
dim=32,
head_dim=32,
n_megnet_blocks=3,
y_mean=0,
y_std=1,
layer_norm=False):
super(MegNetModel_new, self).__init__()
self.dim = dim
self.n_megnet_blocks = n_megnet_blocks
self.node_proj = torch.nn.Linear(x_dim, dim)
self.edge_proj = torch.nn.Linear(edge_dim, dim)
self.global_proj = torch.nn.Linear(u_dim, dim)
self.megnet_blocks = torch.nn.ModuleList([
MegNetBlock(dim,
dim,
dim,
dim,
normalization_cls=torch.nn.LayerNorm,
activation_cls=torch.nn.ELU,
dropout_cls=torch.nn.Dropout,
dropout_prob=0.,
residual=True) for i in range(n_megnet_blocks)
])
self.out_mlp = torch.nn.ModuleList([
OutputLayer_new(
dim,
head_dim,
y_mean=y_mean[i],
y_std=y_std[i],
couple_type=type_,
) for i, type_ in enumerate(constants.TYPES_LIST)
])
self.noise = MultiplicativeGaussianNoise(scale=0.05)
def forward(self, data, add_noise=False):
data = correct_batch_edge_ind(data)
data = correct_inverse_couples_ind(data)
if not hasattr(data, 'global_attr'):
data.global_attr = torch.zeros((data.num_graphs, 1),
dtype=torch.float,
device=data.x.device)
x_out, edge_out, u_out = self.node_proj(data.x), self.edge_proj(data.edge_attr), self.global_proj(data.global_attr)
for i in range(self.n_megnet_blocks):
x_out, edge_out, u_out = self.megnet_blocks[i](
x_out,
data.edge_index,
edge_out,
u_out,
data.batch,
first_block=(i==0))
if add_noise:
x_out = self.noise(x_out)
edge_out = self.noise(edge_out)
u_out = self.noise(u_out)
pred = torch.zeros_like(data.type,
dtype=torch.float,
device=x_out.device)
for type_ in range(8):
if (data.type == type_).any():
pred[data.type == type_] = self.out_mlp[type_](data, x_out,
edge_out,
u_out).view(-1)
return pred
# # Training
from kaggle_champs.metrics import MeanLogGroupMAE, AverageMetric
from kaggle_champs.training import train_epoch
def cycle(iterable):
while True:
for x in iterable:
yield x
# +
import torch
from tqdm.autonotebook import tqdm
def train_epoch(global_iteration, epoch, model, device, optimizer,
train_loader, tb_logger, gradient_accumulation_steps=1, swa=False, noise=False):
model.train()
avg_loss = AverageMetric()
log_mae = MeanLogGroupMAE()
pbar = tqdm(train_loader)
for step, data in enumerate(pbar):
data = data.to(device)
pred = model(data, add_noise=noise)
loss = torch.nn.L1Loss(reduction='mean')(pred.view(-1),
data.y.view(-1))
loss.backward()
if (step + 1) % gradient_accumulation_steps == 0:
optimizer.step()
optimizer.zero_grad()
global_iteration += 1
if swa:
optimizer.update_swa()
tb_logger.add_scalar('loss', loss.item(), global_iteration)
avg_loss.update(loss.item() * data.num_graphs, data.num_graphs)
log_mae.update(pred.view(-1), data.y.view(-1), data.type)
pbar.set_postfix_str(f'loss: {avg_loss.compute():.4f}')
return avg_loss.compute(), log_mae, global_iteration
# +
def test_model(model, loader):
model.eval()
log_mae = MeanLogGroupMAE()
avg_loss = AverageMetric()
with torch.no_grad():
for data in loader:
data = data.to(device)
pred = model(data)
loss = torch.nn.L1Loss(reduction='mean')(pred.view(-1),
data.y.view(-1))
avg_loss.update(loss.item() * data.num_graphs, data.num_graphs)
log_mae.update(pred.view(-1), data.y.view(-1), data.type.view(-1))
return avg_loss.compute(), log_mae
def make_log(epoch, lr, loss, tr_logmae, val_logmae):
results = {
'epoch': epoch,
'lr': lr,
'loss': loss,
'tr_logmae': tr_logmae.compute(),
'val_logmae': val_logmae.compute(),
}
for k, v in tr_logmae.compute_individuals().items():
results.update({'tr_' + k: v})
for k, v in val_logmae.compute_individuals().items():
results.update({'val_' + k: v})
return results
def save_checkpoint(dir_path, model, optimizer, scheduler, epoch):
torch.save(model.state_dict(), dir_path + f'model_epoch_{epoch}.pth')
torch.save(optimizer.state_dict(),
dir_path + f'optimizer_epoch_{epoch}.pth')
torch.save(scheduler.state_dict(),
dir_path + f'scheduler_epoch_{epoch}.pth')
# -
# # Run
from tensorboardX import SummaryWriter
import shutil
OUTPUT_DIR = './models/megnet_256x10_new_arch_3/'
# !mkdir -p {OUTPUT_DIR}
tb_logger = SummaryWriter(OUTPUT_DIR+'tb_log/')
global_iteration = 0
SAVE_INTERVAL = 10
MAX_EPOCH = 150
val_loader = DataLoader(val_dataset,
batch_size=64,
shuffle=False,
num_workers=8,
follow_batch=[
'bonds_edge_ind',
'bonds_links_edge_ind',
'paths_edge_ind',
'couples_edge_ind',
'inverse_couple_ind',
])
train_loader = DataLoader(train_dataset,
batch_size=32,
num_workers=8,
shuffle=True,
follow_batch=[
'bonds_edge_ind',
'bonds_links_edge_ind',
'paths_edge_ind',
'couples_edge_ind',
'inverse_couple_ind',
], drop_last=True)
batch = next(iter(val_loader))
batch = correct_batch_edge_ind(batch)
batch = correct_inverse_couples_ind(batch)
batch
((batch.y[batch.inverse_couple_ind] - batch.y) != 0).any()
y_mean = train.groupby(train.type.map(
constants.TYPES_DICT)).scalar_coupling_constant.mean().sort_index().values
y_std = train.groupby(train.type.map(
constants.TYPES_DICT)).scalar_coupling_constant.std().sort_index().values
device = torch.device('cuda')
model = MegNetModel_new(edge_dim=data.edge_attr.size()[1],
x_dim=data.x.size()[1],
u_dim=1,
dim=300,
head_dim=300,
n_megnet_blocks=10,
y_mean=y_mean,
y_std=y_std,
layer_norm=False).to(device)
from kaggle_champs.optimizer import RAdam
optimizer = RAdam(model.parameters(), lr=2e-4)
scheduler = torch.optim.lr_scheduler.StepLR(optimizer, step_size=20, gamma=1.)
# train loop
logs = []
for epoch in range(1, 145):
lr = scheduler.optimizer.param_groups[0]['lr']
tr_loss, tr_logmae, global_iteration = train_epoch(global_iteration,
epoch,
model,
device,
optimizer,
train_loader,
tb_logger,
gradient_accumulation_steps=2)
#optimizer.update_swa()
scheduler.step()
val_loss, val_logmae = test_model(model, val_loader)
epoch_log = make_log(epoch, lr, tr_loss, tr_logmae, val_logmae)
logs.append(epoch_log)
pd.DataFrame(logs).to_csv(OUTPUT_DIR + 'log.csv')
print('Epoch: {epoch:03d}, LR: {lr:7f}, Loss: {loss:.7f}, \
Train LogMAE: {tr_logmae:.7f}, Val LogMAE: {val_logmae:.7f}'.format(
**epoch_log))
#optimizer.swap_swa_sgd()
#val_loss_swa, val_logmae_swa = test_model(model, val_loader)
#optimizer.swap_swa_sgd()
#print(f'Val LogMAE SWA: {val_logmae_swa.compute():.7f}')
if epoch % SAVE_INTERVAL == 0:
save_checkpoint(OUTPUT_DIR, model, optimizer, scheduler, epoch)
tb_logger.add_scalar('lr', lr, global_iteration)
tb_logger.add_scalar('val_loss', val_loss, global_iteration)
tb_logger.add_scalars('global_logmae', {
'tr_logmae': epoch_log['tr_logmae'],
'val_logmae': epoch_log['val_logmae']
}, global_iteration)
for type_ in constants.TYPES_LIST:
tb_logger.add_scalars(
type_, {
'tr_' + type_: epoch_log['tr_' + type_],
'val_' + type_: epoch_log['val_' + type_]
}, global_iteration)
save_checkpoint(OUTPUT_DIR, model, optimizer, scheduler, epoch=145)
optimizer.param_groups[0]['lr'] = 1e-4
scheduler = torch.optim.lr_scheduler.StepLR(optimizer, step_size=3, gamma=0.5)
# train loop
logs = []
for epoch in range(146, 168):
lr = scheduler.optimizer.param_groups[0]['lr']
tr_loss, tr_logmae, global_iteration = train_epoch(global_iteration,
epoch,
model,
device,
optimizer,
train_loader,
tb_logger,
gradient_accumulation_steps=1)
#optimizer.update_swa()
scheduler.step()
val_loss, val_logmae = test_model(model, val_loader)
epoch_log = make_log(epoch, lr, tr_loss, tr_logmae, val_logmae)
logs.append(epoch_log)
pd.DataFrame(logs).to_csv(OUTPUT_DIR + 'log.csv')
print('Epoch: {epoch:03d}, LR: {lr:7f}, Loss: {loss:.7f}, \
Train LogMAE: {tr_logmae:.7f}, Val LogMAE: {val_logmae:.7f}'.format(
**epoch_log))
#optimizer.swap_swa_sgd()
#val_loss_swa, val_logmae_swa = test_model(model, val_loader)
#optimizer.swap_swa_sgd()
#print(f'Val LogMAE SWA: {val_logmae_swa.compute():.7f}')
if epoch % SAVE_INTERVAL == 0:
save_checkpoint(OUTPUT_DIR, model, optimizer, scheduler, epoch)
tb_logger.add_scalar('lr', lr, global_iteration)
tb_logger.add_scalar('val_loss', val_loss, global_iteration)
tb_logger.add_scalars('global_logmae', {
'tr_logmae': epoch_log['tr_logmae'],
'val_logmae': epoch_log['val_logmae']
}, global_iteration)
for type_ in constants.TYPES_LIST:
tb_logger.add_scalars(
type_, {
'tr_' + type_: epoch_log['tr_' + type_],
'val_' + type_: epoch_log['val_' + type_]
}, global_iteration)
save_checkpoint(OUTPUT_DIR, model, optimizer, scheduler, epoch=168)
# # Make sub
def merge_direction(df):
inverse_direction = df.rename(
{
'atom_index_1': 'atom_index_0',
'atom_index_0': 'atom_index_1'
},
axis=1)
merged = pd.merge(df,
inverse_direction,
on=['molecule_name', 'atom_index_0', 'atom_index_1'],
suffixes=('', '_bis'))
merged['scalar_coupling_constant'] = (merged['scalar_coupling_constant'] + merged['scalar_coupling_constant_bis']) / 2
return merged.drop('scalar_coupling_constant_bis', axis=1)
def predict(model, input_data, checkpoint_path):
model.load_state_dict(torch.load(checkpoint_path))
pred_dataset = MoleculeDataset(
metadata=input_data,
base_dir=constants.STRUCT_DATA_PATH,
transform=T.Compose([
AddBondPath(),
AddVirtualEdges(),
AddEdgeDistanceAndDirection(dist_noise=0.),
AddGlobalAttr(),
SortTarget(),
AddBondLinks(),
AddCounts(),
AddInverseCouples(),
]))
pred_loader = DataLoader(pred_dataset,
batch_size=64,
shuffle=False,
num_workers=8,
follow_batch=[
'bonds_edge_ind', 'bonds_links_edge_ind',
'paths_edge_ind', 'couples_edge_ind', 'inverse_couple_ind'
])
model.eval()
preds = []
inds = []
couples = []
for data in tqdm(pred_loader):
with torch.no_grad():
data = data.to(device)
pred = model(data).detach().cpu().numpy()
ind = data.mol_ind[data.couples_edge_ind_batch].detach().cpu().numpy()
couple_ind = data.couples_ind.cpu().numpy()
df = pd.DataFrame({
'molecule_name' : pred_dataset.molecules[ind].ravel(),
'molecule_ind': ind.ravel(),
'atom_index_0': couple_ind[:,0].ravel(),
'atom_index_1': couple_ind[:,1].ravel(),
})
df.sort_values(['molecule_ind', 'atom_index_0', 'atom_index_1'], ascending=True, inplace=True)
np.testing.assert_array_equal(df.molecule_ind, ind.ravel())
df['scalar_coupling_constant'] = pred
preds.append(df.drop('molecule_ind', axis=1))
pred = pd.concat(preds)
pred = merge_direction(pred)
merged = pd.merge(input_data,
pred,
on=['molecule_name', 'atom_index_0', 'atom_index_1'],
how='left', suffixes=('_truth', ''))
assert merged.dropna().shape[0] == input_data.shape[0]
return merged.loc[:, ['id', 'scalar_coupling_constant']].set_index('id'), pred
pred_val, p = predict(model, val_data, f'{OUTPUT_DIR}/model_epoch_168.pth')
pred_val.head()
def score(pred, ref_data):
merged = pd.merge(ref_data, pred, how='left', left_on='id', right_index=True, suffixes=('', '_pred'))
merged['abs_error'] = (merged['scalar_coupling_constant'] - merged['scalar_coupling_constant_pred']).abs()
result = merged.groupby('type')['abs_error'].mean()
result.iloc[:] = np.log(np.maximum(result.values, 1e-9))
return result.mean(), result.to_dict()
score(pred_val, val_data)
test = pd.read_csv(script_dir + '/../../data/test.csv')
sub, _ = predict(model, test, f'{OUTPUT_DIR}/model_epoch_168.pth')
sub.head()
# !mkdir -p subs/lam_03_v1/
sub.to_csv('./subs/lam_03_v1/sub.csv', index=True)
pred_val.to_csv('./subs/lam_03_v1/pred_val.csv', index=True)
|
solutions/5/lam/Megnet-256x10-new_arch_3-Copy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression
#
# A simple linear regression model to serve as a benchmark against which the performance of other techniques can be compared. This model will not be optimised.
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from time import time
matplotlib.rcParams['figure.figsize'] = (16, 9)
pd.options.display.max_columns = 999
# ## Load Dataset
df = pd.read_csv('../datasets/hourly-weather-wind_direction.csv', parse_dates=[0], index_col='DateTime')
print(df.shape)
df.head()
# ## Define Parameters
#
# Make predictions for 24-hour period using a training period of four weeks.
dataset_name = 'Hourly Weather Wind Direction'
dataset_abbr = 'HWD'
model_name = 'LinearRegression'
context_length = 24*7*4 # Four weeks
prediction_length = 24
# ## Define Error Metric
#
# The seasonal variant of the mean absolute scaled error (MASE) will be used to evaluate the forecasts.
def calc_sMASE(training_series, testing_series, prediction_series, seasonality=prediction_length):
a = training_series.iloc[seasonality:].values
b = training_series.iloc[:-seasonality].values
d = np.sum(np.abs(a-b)) / len(a)
errors = np.abs(testing_series - prediction_series)
return np.mean(errors) / d
# ## Example Linear Regression Model
#
# Functions will be used to readily prepare datasets for each time series.
def select_ts(ts, df=df):
dft = df.iloc[:, ts-1]
return dft
def create_features(dft, window=prediction_length):
dftf = dft.copy()
dftf = dftf.reset_index()
# One hot encode day of week
day_names = ['mon', 'tue', 'wed', 'thu', 'fri', 'sat', 'sun']
for i in range(7):
dftf[day_names[i]] = (dftf['DateTime'].dt.dayofweek == i).astype(int)
# One hot encode hour of day
hour_names = ['hour_%s' % str(i) for i in range(24)]
for i in range(24):
dftf[hour_names[i]] = (dftf['DateTime'].dt.hour == i).astype(int)
# Continuous variable for years
dftf['annual'] = (dftf['DateTime'] - dftf['DateTime'][0]).dt.days / 365
# Moving average
dftf['ma'] = dftf.iloc[:, 1].rolling(window).mean()
dftf['ma'] = dftf['ma'].shift(window)
dftf.dropna(inplace=True)
dftf.set_index('DateTime', inplace=True)
return dftf
def split_data(dftf, test_length=prediction_length):
X_train = dftf.iloc[:-test_length, 1:]
X_test = dftf.iloc[-test_length:, 1:]
y_train = dftf.iloc[:-test_length, 0]
y_test = dftf.iloc[-test_length:, 0]
return X_train, X_test, y_train, y_test
# Illustrate model for an example household. Ridge regression is used for regularisation.
# +
from sklearn.linear_model import Ridge
ts = 1
df_ex = select_ts(ts)
df_ex = create_features(df_ex)
X_train, X_test, y_train, y_test = split_data(df_ex)
lr = Ridge()
lr.fit(X_train, y_train)
y_pred = lr.predict(X_test)
y_pred[y_pred < 0] = 0
sMASE = calc_sMASE(y_train[-(context_length + prediction_length):-prediction_length], y_test, y_pred)
print("sMASE: {:.4f}".format(sMASE))
# -
# Plot example predictions.
plt.plot(y_test.values, label='Actual', c='k', linewidth=1)
plt.plot(y_pred, label='Predicted', c='b', linestyle='--', linewidth=2)
plt.legend()
plt.title('ts%s' % ts);
# Examine example linear regression model coefficients.
# +
coefs = lr.coef_
cols = df_ex.columns[1:]
params = pd.Series(coefs, index=cols)
fig, ax = plt.subplots()
ax.barh(range(len(params)), params.values, align='center', color='b')
plt.xlabel('Feature coefficient')
plt.yticks(np.arange(len(params)), params.index)
plt.ylabel('Feature')
plt.ylim(-1, len(params))
plt.title('ts%s' % ts);
# -
# ## Evaluate Linear Regresion
# +
results = df.copy()
for i in range(len(df.columns)):
ts = i + 1
dft = select_ts(ts)
dftf = create_features(dft)
X_train, X_test, y_train, y_test = split_data(dftf)
lr = Ridge()
lr.fit(X_train, y_train)
y_pred = lr.predict(X_test)
results['pred%s' % ts] = np.nan
results['pred%s' % ts].iloc[-prediction_length:] = y_pred
# -
results0 = results.copy()
results0[results0 < 0] = 0
results0.head()
sMASEs = []
for i, col in enumerate(df.columns):
sMASEs.append(calc_sMASE(results[col].iloc[-(context_length + prediction_length):-prediction_length],
results[col].iloc[-prediction_length:],
results['pred%s' % str(i+1)].iloc[-prediction_length:]))
fig, ax = plt.subplots()
ax.hist(sMASEs, bins=20)
ax.set_title('Distributions of sMASEs for {} dataset'.format(dataset_name))
ax.set_xlabel('sMASE')
ax.set_ylabel('Count');
sMASE = np.mean(sMASEs)
print("Overall sMASE: {:.4f}".format(sMASE))
# Show some example forecasts.
# +
fig, ax = plt.subplots(5, 2, sharex=True)
ax = ax.ravel()
for col in range(1, 11):
ax[col-1].plot(results.index[-prediction_length:], results['ts%s' % col].iloc[-prediction_length:],
label='Actual', c='k', linestyle='--', linewidth=1)
ax[col-1].plot(results.index[-prediction_length:], results['pred%s' % col].iloc[-prediction_length:],
label='Predicted', c='b')
fig.suptitle('{} Predictions'.format(dataset_name))
ax[0].legend();
# -
# Store the predictions and accuracy score for the Linear Regression models.
# +
import pickle
with open('{}-sMASE.pkl'.format(dataset_abbr), 'wb') as f:
pickle.dump(sMASE, f)
with open('../_results/{}/{}-results.pkl'.format(model_name, dataset_abbr), 'wb') as f:
pickle.dump(results.iloc[-prediction_length:], f)
# -
|
LinearRegression/hourly-weather-wind_direction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import json
import numpy as np
import pandas as pd
from tqdm.notebook import tqdm
# +
def clean_name(name):
name = name.replace('(c)', '')
name = name.replace('†', '')
name = name.strip()
return name
def find_wicket_c(wicket):
wicket = wicket.replace('†', '')
wicket = wicket.strip()
if wicket.startswith('c & b '):
wicket_c = wicket[len('c & b '):]
elif wicket.startswith('c '):
wicket_c = wicket[len('c '):wicket.find(' b ')]
else:
wicket_c = 'NA'
return wicket_c
def find_wicket_b(wicket):
wicket = wicket.replace('†', '')
wicket = wicket.strip()
if wicket.startswith('c & b '):
wicket_b = wicket[len('c & b '):]
elif wicket.startswith('b '):
wicket_b = wicket[len('b '):]
elif wicket.startswith('lbw b '):
wicket_b = wicket[len('lbw b '):]
elif wicket.startswith('c '):
wicket_b = wicket[wicket.find(' b ')+3:]
elif wicket.startswith('st '):
wicket_b = wicket[wicket.find(' b ')+3:]
else:
wicket_b = 'NA'
return wicket_b
def find_wicket_st(wicket):
wicket = wicket.replace('†', '')
wicket = wicket.strip()
if wicket.startswith('st '):
wicket_st = wicket[len('st '):wicket.find(' b ')]
else:
wicket_st = 'NA'
return wicket_st
def find_wicket_ro(wicket):
wicket = wicket.replace('†', '')
wicket = wicket.strip()
if wicket.startswith('run out ('):
if "/" in wicket:
wicket_ro_t, wicket_ro_c = wicket[len('run out ('):-1].split('/')[:2]
else:
wicket_ro_t, wicket_ro_c = wicket[len('run out ('):-1], wicket[len('run out ('):-1]
else:
wicket_ro_t, wicket_ro_c = 'NA', 'NA'
return wicket_ro_t, wicket_ro_c
def match_player_caught(row, players_1, players_2):
if row['wicket_c'] == 'NA':
return 'NA'
if row['inning'] == 1:
players = players_2
else:
players = players_1
for player in players:
if row['wicket_c'] in player:
return player
else:
return 'Not Matched'
def match_player_bowled(row, players_1, players_2):
if row['wicket_b'] == 'NA':
return 'NA'
if row['inning'] == 1:
players = players_2
else:
players = players_1
for player in players:
if row['wicket_b'] in player:
return player
else:
return 'Not Matched'
def match_player_stumped(row, players_1, players_2):
if row['wicket_st'] == 'NA':
return 'NA'
if row['inning'] == 1:
players = players_2
else:
players = players_1
for player in players:
if row['wicket_st'] in player:
return player
else:
return 'Not Matched'
def match_player_ro_throwed(row, players_1, players_2):
if row['wicket_ro_t'] == 'NA':
return 'NA'
if row['inning'] == 1:
players = players_2
else:
players = players_1
for player in players:
if row['wicket_ro_t'] in player:
return player
else:
return 'Not Matched'
def match_player_ro_caught(row, players_1, players_2):
if row['wicket_ro_c'] == 'NA':
return 'NA'
if row['inning'] == 1:
players = players_2
else:
players = players_1
for player in players:
if row['wicket_ro_c'] in player:
return player
else:
return 'Not Matched'
# -
def create_player_df(match_dir):
batsman_df = pd.read_csv(os.path.join(match_dir, 'batsman_df.csv'))
if len(batsman_df) == 0:
return pd.DataFrame(columns=['NA'])
bowler_df = pd.read_csv(os.path.join(match_dir, 'bowler_df.csv'))
bowler_df = bowler_df.rename(columns={
'runs': 'runs_given',
'fours': 'fours_given',
'sixes': 'sixes_given'
})
if 'inning' in bowler_df.columns:
bowler_df = bowler_df.drop(columns=['inning'])
with open(os.path.join(match_dir, 'meta_data.json'), 'r') as file:
meta_data = json.load(file)
team_map = {1:meta_data['team_1'], 2:meta_data['team_2']}
player_df = batsman_df[batsman_df['name'] != 'Extras'].copy()
player_df['is_captain'] = player_df['name'].str.contains('(c)', regex=False, case=True)
player_df['is_wicket_keeper'] = player_df['name'].str.contains('†', regex=False, case=True)
player_df['name'] = player_df['name'].apply(clean_name)
player_df['team'] = player_df['inning'].map(team_map)
player_df['wicket_c'] = player_df['wicket'].apply(find_wicket_c)
#player_df['wicket_b'] = player_df['wicket'].apply(find_wicket_b)
player_df['wicket_st'] = player_df['wicket'].apply(find_wicket_st)
player_df['wicket_ro_t'], player_df['wicket_ro_c'] = zip(*player_df['wicket'].apply(find_wicket_ro))
players_1 = player_df[player_df['inning']==1]['name']
players_2 = player_df[player_df['inning']==2]['name']
player_df['player_caught'] = player_df.apply(match_player_caught, axis=1,
players_1=players_1, players_2=players_2)
#player_df['player_bowled'] = player_df.apply(match_player_bowled, axis=1,
# players_1=players_1, players_2=players_2)
player_df['player_stumped'] = player_df.apply(match_player_stumped, axis=1,
players_1=players_1, players_2=players_2)
player_df['player_ro_throwed'] = player_df.apply(match_player_ro_throwed, axis=1,
players_1=players_1, players_2=players_2)
player_df['player_ro_caught'] = player_df.apply(match_player_ro_caught, axis=1,
players_1=players_1, players_2=players_2)
player_df['caught_count'] = player_df['name'].map(player_df['player_caught'].value_counts())
player_df['caught_count'] = player_df['caught_count'].fillna(0).astype('int')
#player_df['bowled_count'] = player_df['name'].map(player_df['player_bowled'].value_counts())
#player_df['bowled_count'] = player_df['bowled_count'].fillna(0).astype('int')
player_df['stumped_count'] = player_df['name'].map(player_df['player_stumped'].value_counts())
player_df['stumped_count'] = player_df['stumped_count'].fillna(0).astype('int')
player_df['ro_throwed_count'] = player_df['name'].map(player_df['player_ro_throwed'].value_counts())
player_df['ro_throwed_count'] = player_df['ro_throwed_count'].fillna(0).astype('int')
player_df['ro_caught_count'] = player_df['name'].map(player_df['player_ro_caught'].value_counts())
player_df['ro_caught_count'] = player_df['ro_caught_count'].fillna(0).astype('int')
player_df = pd.merge(player_df, bowler_df, on='name', how='outer').fillna('NA')
return player_df
def create_dream11_df(match_dir):
player_df = pd.read_csv(os.path.join(match_dir, 'player_df.csv'))
if len(player_df) == 0:
return pd.DataFrame(columns=['NA'])
player_df['strike_rate'] = player_df['strike_rate'].replace('-', -1).astype('float')
player_df['runs'] = player_df['runs'].replace('-', 0).astype('float')
player_df['wicket'] = player_df['wicket'].replace('absent hurt', 'Did not bat')
player_df['economy'] = pd.to_numeric(player_df['economy'].replace('-', np.nan))
dream11_df = player_df[['name', 'team']].copy()
dream11_df['starting_11_actual'] = 'Yes'
dream11_df['starting_11_points'] = 4
dream11_df['runs_actual'] = player_df['runs']
dream11_df['runs_points'] = dream11_df['runs_actual']
dream11_df['fours_actual'] = player_df['fours']
dream11_df['fours_points'] = dream11_df['fours_actual']
dream11_df['sixes_actual'] = player_df['sixes']
dream11_df['sixes_points'] = dream11_df['sixes_actual'] * 2
atleast_10_balls = player_df['balls'] >= 10
strike_rate_points = pd.to_numeric(pd.cut(player_df['strike_rate'],
bins=[0, 50, 60, 70],
right=False,
labels=[-6, -4, -2])).fillna(0)
dream11_df['atleast_10_balls'] = atleast_10_balls
dream11_df['strike_rate_actual'] = player_df['strike_rate']
dream11_df['strike_rate_points'] = dream11_df['atleast_10_balls'] * strike_rate_points
points_50 = (player_df['runs'] >= 50) * 8
points_100 = (player_df['runs'] >= 100) * 8
dream11_df['50_100_points'] = points_50 + points_100
dream11_df['duck_actual'] = (player_df['runs'] == 0) & (player_df['wicket'] != 'Did not bat')
dream11_df['duck_points'] = dream11_df['duck_actual'] * -2
dream11_df['wickets_actual'] = player_df['wickets'].replace('NA', 0).fillna(0)
dream11_df['wickets_points'] = dream11_df['wickets_actual'] * 25
dream11_df['maidens_actual'] = player_df['maidens'].replace('NA', 0).fillna(0)
dream11_df['maidens_points'] = dream11_df['maidens_actual'] * 8
atleast_2_overs = player_df['overs'] >= 2
economy_rate_points = pd.to_numeric(pd.cut(player_df['economy'],
bins=[0, 4, 5, 6.01, 9, 10.01, 11.01, 36],
right=False,
labels=[6, 4, 2, 0, -2, -4, -6])).fillna(0)
dream11_df['atleast_2_overs'] = atleast_2_overs
dream11_df['economy_rate_actual'] = player_df['economy']
dream11_df['economy_rate_points'] = dream11_df['atleast_2_overs'] * economy_rate_points
wickets_4 = (player_df['wickets'] >= 4) * 8
wickets_4 = (player_df['wickets'] >= 5) * 8
dream11_df['bonus_points'] = wickets_4 + wickets_4
dream11_df['catches_actual'] = player_df['caught_count']
dream11_df['catches_points'] = dream11_df['catches_actual'] * 8
dream11_df['run_out_actual'] = player_df['ro_throwed_count'] + player_df['ro_caught_count']
dream11_df['run_out_points'] = dream11_df['run_out_actual'] * 6
dream11_df['stumping_actual'] = player_df['stumped_count']
dream11_df['stumping_points'] = dream11_df['stumping_actual'] * 12
points_cols = [col for col in dream11_df.columns if col.endswith('_points')]
dream11_df['total_points'] = dream11_df[points_cols].sum(axis=1)
dream11_df = dream11_df.sort_values(by='total_points', ascending=False).reset_index(drop=True)
dream11_df['rank'] = dream11_df.index + 1
cols = list(dream11_df.columns)
cols.remove('total_points')
cols = ['total_points'] + cols
dream11_df = dream11_df[cols]
return dream11_df
# +
data_dir = 'Data'
seasons = range(2010, 2021)
for season in tqdm(seasons, desc='seasons'):
season_dir = os.path.join(data_dir, str(season))
matches = sorted(os.listdir(season_dir))
for match in tqdm(matches, desc='matches', leave=False):
match_dir = os.path.join(season_dir, match)
try:
player_df = create_player_df(match_dir)
except Exception as e:
print(0, match_dir)
print(e)
raise
player_df.to_csv(os.path.join(match_dir, 'player_df.csv'), index=False)
try:
dream11_df = create_dream11_df(match_dir)
except Exception as e:
print(1, match_dir)
print(e)
raise
dream11_df.to_csv(os.path.join(match_dir, 'dream11_df.csv'), index=False)
# -
# ### Merging all dream11 tables
# +
data_dir = 'Data'
seasons = range(2010, 2021)
dream11_df = pd.DataFrame()
for season in tqdm(seasons, desc='seasons'):
season_dir = os.path.join(data_dir, str(season))
matches = sorted(os.listdir(season_dir))
for match in tqdm(matches, desc='matches', leave=False):
match_dir = os.path.join(season_dir, match)
with open(os.path.join(match_dir, 'meta_data.json'), 'r') as file:
meta_data = json.load(file)
match_dream11_df = pd.read_csv(os.path.join(match_dir, 'dream11_df.csv'))
cols = list(match_dream11_df.columns)
if len(match_dream11_df):
match_dream11_df['season_id'] = season
match_dream11_df['match_id'] = meta_data['match_id']
cols = ['season_id', 'match_id'] + cols
match_dream11_df = match_dream11_df[cols]
dream11_df = pd.concat([dream11_df, match_dream11_df], ignore_index=True)
dream11_df['match_id'] = dream11_df['match_id'].astype('int')
dream11_df = dream11_df.sort_values(by=['season_id', 'match_id'])
dream11_df.to_csv(os.path.join(data_dir, 'dream11_df.csv'), index=False)
|
2-Computing-Dream11-Points.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dougscohen/DS-Unit-2-Linear-Models/blob/master/module4-logistic-regression/LS_DS_214_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="kJVfOZGu7jVO" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 4*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Logistic Regression
#
#
# ## Assignment 🌯
#
# You'll use a [**dataset of 400+ burrito reviews**](https://srcole.github.io/100burritos/). How accurately can you predict whether a burrito is rated 'Great'?
#
# > We have developed a 10-dimensional system for rating the burritos in San Diego. ... Generate models for what makes a burrito great and investigate correlations in its dimensions.
#
# - [ ] Do train/validate/test split. Train on reviews from 2016 & earlier. Validate on 2017. Test on 2018 & later.
# - [ ] Begin with baselines for classification.
# - [ ] Use scikit-learn for logistic regression.
# - [ ] Get your model's validation accuracy. (Multiple times if you try multiple iterations.)
# - [ ] Get your model's test accuracy. (One time, at the end.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
#
#
# ## Stretch Goals
#
# - [ ] Add your own stretch goal(s) !
# - [ ] Make exploratory visualizations.
# - [ ] Do one-hot encoding.
# - [ ] Do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html).
# - [ ] Get and plot your coefficients.
# - [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="AZxmpNlu7jVW" colab_type="code" colab={}
# Load data downloaded from https://srcole.github.io/100burritos/
import pandas as pd
df = pd.read_csv(DATA_PATH+'burritos/burritos.csv')
# + id="429OyKqP7jVZ" colab_type="code" colab={}
# Derive binary classification target:
# We define a 'Great' burrito as having an
# overall rating of 4 or higher, on a 5 point scale.
# Drop unrated burritos.
df = df.dropna(subset=['overall'])
df['Great'] = df['overall'] >= 4
# + id="ueaLOU_W7jVb" colab_type="code" colab={}
# Clean/combine the Burrito categories
df['Burrito'] = df['Burrito'].str.lower()
california = df['Burrito'].str.contains('california')
asada = df['Burrito'].str.contains('asada')
surf = df['Burrito'].str.contains('surf')
carnitas = df['Burrito'].str.contains('carnitas')
df.loc[california, 'Burrito'] = 'California'
df.loc[asada, 'Burrito'] = 'Asada'
df.loc[surf, 'Burrito'] = 'Surf & Turf'
df.loc[carnitas, 'Burrito'] = 'Carnitas'
df.loc[~california & ~asada & ~surf & ~carnitas, 'Burrito'] = 'Other'
# + id="i6XDWr6Y7jVe" colab_type="code" colab={}
# Drop some high cardinality categoricals
df = df.drop(columns=['Notes', 'Location', 'Reviewer', 'Address', 'URL', 'Neighborhood'])
# + id="-67KefPx7jVh" colab_type="code" colab={}
# Drop some columns to prevent "leakage"
df = df.drop(columns=['Rec', 'overall'])
# + id="oyjncWVt7jVm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 252} outputId="e6ced009-8c96-4cd6-dbdc-2d9ba36a0722"
print(df.shape)
df.head()
# + id="l39B5LamJQTG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 123} outputId="f445d738-57ec-4c60-f478-bb5a233a99af"
# Looking at datatypes of columns and specifically the 'Date' column
df.dtypes.head()
# + id="UkmFOW5_Ja0S" colab_type="code" colab={}
# Use pandas to_datetime function to turn 'Date' column into datetime format
import pandas as pd
df['Date'] = pd.to_datetime(df['Date'], infer_datetime_format=True)
# + id="GX5j_iknJz4m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="81e97e55-e042-4b1a-9a1f-3115e062c7cf"
# Look at values in 'Date' column
df['Date']
# + id="iHAjc0xjJ2Vh" colab_type="code" colab={}
# Establish cutoffs so that we can separate our dataframe by dates
cutoff_1 = pd.to_datetime('2017-01-01')
cutoff_2 = pd.to_datetime('2018-01-01')
# Train dataset will include all observations before 2017
train = df[df['Date'] < cutoff_1]
# Validate dataset will include all observations in 2017
val = df[(df['Date'] >= cutoff_1) & (df['Date'] < cutoff_2)]
# Test dataset will include all observations after in 2018 and later
test = df[df['Date'] >= cutoff_2]
# + id="4xzigCtQKv92" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 335} outputId="05af03ae-6017-44ee-9ab4-df62bb954391"
print(train.shape)
train.head()
# + id="9sCm2Um8Kw7Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 335} outputId="f55c1f08-f6a8-4ce3-f32d-6d47edcc8b4c"
print(val.shape)
val.head()
# + id="ZLwkLTdpK-Pt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 335} outputId="16b6d37a-2c20-41d1-8a57-ef4252dc9236"
print(test.shape)
test.head()
# + id="Wn1nBkp-LAS_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="d0a54f5e-ec92-4773-e723-d7904a6e4012"
# Baseline for Classification
target = 'Great'
y_train = train[target]
y_val = val[target]
y_train.value_counts(normalize=True)
# + id="wBlK3U6XLknG" colab_type="code" colab={}
# Majority Class is that a burrito is Not Great
# + id="Fx-CjBw1L12t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="a5579b83-71a9-4605-e910-5a9e3cd3a41f"
majority_class = y_train.mode()[0]
# Guessing the majority class for every prediction
y_pred_base = [majority_class] * len(y_train)
y_pred_base
# + id="UVZc_V2ZL7wR" colab_type="code" colab={}
from sklearn.metrics import accuracy_score
# + id="tRpnkHmyMrk1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a77140cb-6355-43ad-9699-8b1e3e92da0c"
# Baseline accuracy if we guessed the majority class (Not a great burrito)
# for every predicition
accuracy_score(y_train, y_pred_base)
# + id="GhbX97woO7b8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3e7f39f9-a5e3-4b71-d48b-0ac81a83a4a9"
# Looking at the categorical columns
train.describe(exclude='number').T.sort_values(by='unique')
# + id="LUkwKWgTPUrB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 324} outputId="0a0e8e76-3b61-4f33-b4ba-8a1b2a7f2257"
# Look at numerical columns
train.describe(include='number')
# + id="ZDhIteUoRDnb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="de919e94-e6e5-4fb1-8364-5ef302aef2b8"
train.describe(include='number').columns
# + id="TxtKfmfaMwoZ" colab_type="code" colab={}
# 1. Import estimator class
from sklearn.linear_model import LogisticRegression
# 2. Substantiate the class
log_reg = LogisticRegression()
# 3. Arrange X features matrix (already did y target vectors)
# Take out columns that are entirely NaNs ('Mass (g), 'Density (g/mL), and 'Queso)
features = ['Yelp', 'Google', 'Cost', 'Hunger', 'Length', 'Circum', 'Volume',
'Tortilla', 'Temp', 'Meat', 'Fillings', 'Meat:filling',
'Uniformity', 'Salsa', 'Synergy', 'Wrap', 'Burrito']
X_train = train[features]
X_val = val[features]
# One hot encode the 'Burrito' column
import category_encoders as ce
encoder = ce.OneHotEncoder(use_cat_names=True)
X_train_encoded = encoder.fit_transform(X_train)
X_val_encoded = encoder.transform(X_val)
# Impute Missing Values
from sklearn.impute import SimpleImputer
imputer = SimpleImputer()
X_train_imputed = imputer.fit_transform(X_train_encoded)
X_val_imputed = imputer.transform(X_val_encoded)
# Standardize the Data
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train_imputed)
X_val_scaled = scaler.transform(X_val_imputed)
# + id="UXxYSbuPTGMq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 194} outputId="b11b2013-e0dd-4b11-f33e-65f9fdc50a70"
# 4. Fit the model
log_reg.fit(X_train_scaled, y_train)
# 5. Apply the model to the new data
log_reg.predict(X_val_scaled)
# + id="aQ7rm-73TcGM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9bdfe2f6-22f7-4741-e098-221be3247d8f"
# Predicting the dependent variable (whethere a burrito is great)
# on the Validation set using Logistic Regression
y_pred_log = log_reg.predict(X_val_scaled)
# Accuracy of the model
accuracy_score(y_val, y_pred_log)
# + id="o7jYM0DQYttl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="189f7fde-3cec-4741-fdf4-5bfa72eb77d1"
# Coefficients
log_reg.coef_
# + id="-wmd5xjgUIea" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="acb81c67-9081-427f-e506-176d04e948c3"
from sklearn.linear_model import LogisticRegressionCV
# Fitting logistic regression CV to the data
log_reg_CV = LogisticRegressionCV()
log_reg_CV.fit(X_train_scaled, y_train)
# + id="Ra89vnPmVq5L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="24df92ba-d9c8-4f20-fb4e-17491480ea8e"
# Predicting whether a burrito is great or not based on Logistic Regression CV
y_pred_logCV = log_reg_CV.predict(X_val_scaled)
# Accuracy of the model
accuracy_score(y_val, y_pred_logCV)
# + id="4KjGINsvV7QW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1cb1d5d7-fade-457d-f72d-35428442882a"
log_reg_CV.intercept_
# + id="hGVa87sTWV8Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="e7d5b724-4504-400d-b540-15c816785b6b"
# Coefficients
log_reg_CV.coef_
# + id="3UWTw5-BWYSr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="5eb98dc3-f8ca-4af4-a58d-8ed4e55c7b94"
# Plotting the coefficients to see which independant variables are most
# strongly correlated with the dependant variable
import matplotlib.pyplot as plt
# %matplotlib inline
coefficients = pd.Series(log_reg_CV.coef_[0], X_train_encoded.columns)
coefficients.sort_values().plot.barh();
# + id="fQxgX0cPop7n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="51971485-c751-43c7-90f6-306d22eca33c"
# Predict whether or not a burrito is great on the test data
X_test = test[features]
X_test_encoded = encoder.transform(X_test)
X_test_imputed = imputer.transform(X_test_encoded)
X_test_scaled = scaler.transform(X_test_imputed)
y_pred_final = log_reg_CV.predict(X_test_scaled)
y_pred_final
# + id="GFN4DUgQq4P9" colab_type="code" colab={}
|
module4-logistic-regression/LS_DS_214_assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Algoritmos en cuerpos finitos
# +
from IPython.display import Math, display
import os
if 'examples' in os.getcwd():
os.chdir('..')
# -
# ## Inverso de un elemento
from compalg import IF
# +
p = 17
F = IF(p)
for a in range(1, p):
display(Math(r"[" + str(a) + r"]_{" + str(p) + r"}^{-1} = " + str(F.inverse(a))))
# -
try:
F.inverse(0)
except:
print("0 no tiene inverso")
from compalg import Var
# #### Funciona también con cuerpos finitos con $q = p^n : n > 1$
# +
x = Var('x')
p = 2
h = x ** 8 + x ** 4 + x ** 3 + x + 1
F = IF(p, h)
f = x ** 6 + x ** 4 + x + 1
display(Math(r"(" + f.__latex__() + r")^{-1} = " + F.inverse(f).__latex__() + " \mod " + h.__latex__()))
# -
# ## Test de irreducibilidad
# +
t = Var('t')
x = Var('x')
tests = [
(IF(2)[t], t + 1),
(IF(2)[t], t ** 7 + t ** 6 + t ** 5 + t ** 4 + t ** 2 + t + 1),
(IF(3)[t], t ** 3 + 1), # (t + 1) ** 3
(IF(2, x ** 2 + x + 1)[t], t ** 2 - t + x),
(IF(2, x ** 2 + x + 1)[t], t ** 2 - (x + 1)), # (t - x) * (t + x)
]
for F, p in tests:
result = "" if F.is_irreducible(p) else "no "
display(Math(r"\text{El polinomio }" + p.__latex__() + r"\in " + F.__latex__() + r"\text{ " + result + r"es irreducible}"))
# -
# ## Logaritmo discreto
#
# $g^x = h \implies x = \log_g h$
# +
x = Var('x')
tests = [
(IF(5), 2, 3),
(IF(59), 2, 11),
(IF(383), 2, 228),
(IF(2, x ** 2 + x + 1), x, x + 1),
(IF(3, x ** 2 + 1), x + 1, x + 2)
]
for F, g, h in tests:
r = getattr(g, '__latex__', lambda: repr(g))
s = getattr(h, '__latex__', lambda: repr(h))
display(Math(r"\log_{" + r() + r"} " + s() + r" = " + str(F.discrete_logarithm(g, h)) + r" \in " + F.__latex__()))
|
examples/finite_fields_tests.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Introduction to Data Science
# # Lecture 13 continued
# *COMP 5360 / MATH 4100, University of Utah, http://datasciencecourse.net/*
#
# We continue lecture 13 but switch from using the [statsmodels](http://statsmodels.sourceforge.net/) library to the [scikit-learn](http://scikit-learn.org/) library.
# + slideshow={"slide_type": "slide"}
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn import metrics
from sklearn.datasets import make_moons, load_iris
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10, 6)
plt.style.use('ggplot')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Example: Classifying two moons
#
# Let's consider a synthetic dataset in the shape of "two moons". Here, each sample has two pieces of information:
# * the *features*, denoted by $x_i$, which are just a two-dimensional coordinate and
# * a *class*, denoted by $y_i$, which is either 0 and 1.
# + slideshow={"slide_type": "-"}
# there are two features contained in X and the labels are contained in y
X,y = make_moons(n_samples=500,random_state=1,noise=0.3)
# X is a 500x2 numpy.ndarray containing the coordinates for each sample
# y is a 500x1 numpy.ndarray containing the class for each sample
print(type(X), type(y))
print(np.shape(X), np.shape(y))
print(np.concatenate((X,y[:, np.newaxis]),axis=1)[:10,:])
# Plot the data, color by class
plt.scatter(X[y == 1, 0], X[y == 1, 1], color="DarkBlue", marker="s",label="class 1")
plt.scatter(X[y == 0, 0], X[y == 0, 1], color="DarkRed", marker="o",label="class 2")
plt.legend(scatterpoints=1)
x_min, x_max = X[:,0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.title('Two Moons Dataset')
plt.xlabel('feature 1')
plt.ylabel('feature 2')
plt.show()
# + [markdown] slideshow={"slide_type": "-"}
# Recall that the goal in **classification** is to develop a rule for classifying the points.
#
# Let's see how to use [scikit-learn](http://scikit-learn.org/) for logistic regression.
# + slideshow={"slide_type": "-"}
# set up the model
# we could specify adittinoal parameters here, but we'll just use the default ones
model = LogisticRegression()
# use the model to fit the data
model.fit(X, y)
# Plot the data, color by class
plt.scatter(X[y == 1, 0], X[y == 1, 1], color="DarkBlue", marker="s",label="class 1")
plt.scatter(X[y == 0, 0], X[y == 0, 1], color="DarkRed", marker="o",label="class 2")
plt.legend(scatterpoints=1)
# Plot the predictions made by Logistic Regression
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 200),np.linspace(y_min, y_max, 200))
zz = model.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.contourf(xx, yy, zz, cmap=ListedColormap(['DarkRed', 'DarkBlue']), alpha=.2)
plt.contour(xx, yy, zz, colors="black", alpha=1, linewidths=0.2)
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.title('Classification of Two Moons using Logistic Regression')
plt.xlabel('feature 1')
plt.ylabel('feature 2')
plt.show()
# + slideshow={"slide_type": "slide"}
print('Confusion Matrix:')
y_pred = model.predict(X)
print(metrics.confusion_matrix(y_true = y, y_pred = y_pred))
print('Precision = ', metrics.precision_score(y_true = y, y_pred = y_pred))
print('Recall = ', metrics.recall_score(y_true = y, y_pred = y_pred))
print('F-score = ', metrics.f1_score(y_true = y, y_pred = y_pred))
print('Jaccard similarity score', metrics.jaccard_similarity_score(y_true = y, y_pred = y_pred))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Another method for classification: k Nearest Neighbors (k-NN)
#
# **Idea:** To decide the class of a given point, find the k nearest neighbors of that point, and let them "vote" on the class. That is, we assign the class to the sample that is most common among its k nearest neighbors.
#
# **Considerations:**
# 1. We must pick k, the number of voting neighbors (typically a small number, say k=10)
# # + 'Nearest' means closest in distance, so there is some flexibility in defining the distance
# # + There are different ways to vote. For example, of the k nearest neighbors, I might give the closest ones more weight than farther ones.
# # + We have to decide how to break ties in the vote.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Dataset: The Iris dataset
#
# This dataset was introduced in 1936 by the statistician [<NAME>](https://en.wikipedia.org/wiki/Ronald_Fisher).
#
# The dataset contains 4 features (attributes) of 50 samples containing 3 different species of iris plants. The goal is to classify the species of iris plant given the attributes.
#
# **Classes:**
# 1. Iris Setosa
# # + Iris Versicolour
# # + Iris Virginica
#
# **Features (attributes):**
# 1. sepal length (cm)
# # + sepal width (cm)
# # + petal length (cm)
# # + petal width (cm)
#
# <img src="iris.png" title="http://mirlab.org/jang/books/dcpr/dataSetIris.asp?title=2-2%20Iris%20Dataset" width="20%">
#
# + [markdown] slideshow={"slide_type": "-"}
# The [seaborn library](https://stanford.edu/~mwaskom/software/seaborn/) is a visualization library. It can be installed from the terminal using
# ```
# conda install seaborn
# ```
# + slideshow={"slide_type": "-"}
import seaborn as sns
sns.set()
df = sns.load_dataset("iris") # built-in dataset in seaborn
print(df.describe())
sns.pairplot(df, hue="species");
# + slideshow={"slide_type": "-"}
# import data, scikit-learn also has this dataset built-in
iris = load_iris()
# For easy plotting and interpretation, we only use first 2 features here.
# We're throwing away useful information - don't do this at home!
X = iris.data[:,:2]
y = iris.target
# Create color maps
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
# plot data
plt.scatter(X[:, 0], X[:, 1], c=y, marker="o", cmap=cmap_bold, s=30)
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.title('Iris dataset')
plt.xlabel('feature 1')
plt.ylabel('feature 2')
plt.show()
# + [markdown] slideshow={"slide_type": "-"}
# We see that it would be fairly easy to separate the "red" irises from the two classes. However, separating the "green" and "blue" ones would be a challenge.
#
# There are three classes, so we can't apply logistic regression. (This isn't completely true; there are extensions of logistic regression to handle more classes, but these are not very popular.) Instead we'll use k-NN.
# + slideshow={"slide_type": "-"}
# set up the model, k-NN classification with k = ?
k = 1
clf = KNeighborsClassifier(n_neighbors=k)
clf.fit(X, y)
# plot classification
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 400),np.linspace(y_min, y_max, 400))
zz = clf.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, zz, cmap=cmap_light)
# plot data
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=cmap_bold,s=30)
plt.title('Classification of Iris dataset using k-NN with k = '+ str(k))
plt.xlabel('feature 1')
plt.ylabel('feature 2')
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.show()
# + slideshow={"slide_type": "-"}
print('Confusion Matrix:')
y_pred = clf.predict(X)
print(metrics.confusion_matrix(y_true = y, y_pred = y_pred))
print('Jaccard similarity score', metrics.jaccard_similarity_score(y_true = y, y_pred = y_pred))
# + [markdown] slideshow={"slide_type": "-"}
# **Some preliminary comments on the parameter, $k$:**
#
# - For k large (say $k=100$), the *decision boundary* (boundary between classes) is smooth. The model is not very complex - it could basically be described by a few lines. The model has low variance in the sense that if the data were to change slightly, the model wouldn't change much. (There are many voters.) Since the model doesn't depend on the data very much, we might expect that it would *generalize* to new data points.
#
# - For k small (say $k=1$), the decision boundary is very wiggly. The model is very complex - it definitely can't be described by a few lines. The model has high variance in the sense that if the data were to change slightly, the model would change quite a bit. Since the model is very dependent on the dataset, we would say that it wouldn't generalize to new data points well. In this case, we would say that the model has overfit the data. (We saw a similar phenomena in regression using high degree polynomials.)
#
#
# **Questions:**
# 1. How to choose k? (more on this below)
# # + Which does a better job on the two moons dataset: k-NN or logistic regression?
#
# To the moons!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Example: k-NN on the moons dataset
# + slideshow={"slide_type": "-"}
# moons
X,y = make_moons(n_samples=500,random_state=1,noise=0.3)
# + slideshow={"slide_type": "-"}
model = KNeighborsClassifier(n_neighbors = 100)
model.fit(X, y)
# Plot the data, color by class
plt.scatter(X[y == 1, 0], X[y == 1, 1], color="darkblue", marker="s",label="class 1")
plt.scatter(X[y == 0, 0], X[y == 0, 1], color="darkred", marker="o",label="class 2")
plt.legend(scatterpoints=1)
# Plot the predictions made by Logistic Regression
x_min, x_max = X[:,0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 200),np.linspace(y_min, y_max, 200))
zz = model.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.contourf(xx, yy, zz, cmap=ListedColormap(['DarkRed', 'DarkBlue']), alpha=.2)
plt.contour(xx, yy, zz, colors="black", alpha=1, linewidths=0.2)
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.xlabel('feature 1')
plt.ylabel('feature 2')
plt.title('Classification of Two Moons using k-NN')
plt.show()
# + slideshow={"slide_type": "-"}
print('Confusion Matrix:')
y_pred = model.predict(X)
print(metrics.confusion_matrix(y_true = y, y_pred = y_pred))
print('Precision = ', metrics.precision_score(y_true = y, y_pred = y_pred))
print('Recall = ', metrics.recall_score(y_true = y, y_pred = y_pred))
print('F-score = ', metrics.f1_score(y_true = y, y_pred = y_pred))
print('Jaccard similarity score', metrics.jaccard_similarity_score(y_true = y, y_pred = y_pred))
# + [markdown] slideshow={"slide_type": "-"}
# For *good* choices of the parameter k, k-NN has better performance than logistic regression. Logistic regression suffers because the decision boundary isn't curved. For this reason, it is called a *linear classifier*. (However there are extensions to logistic regression that allow the decision boundary to curve).
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Model generalizability and cross-validation
#
# In classification, and other prediction problems, we would like to develop a model on a dataset, the *training dataset*, that will not only perform well on that dataset but on similar data that the model hasn't yet seen, the *testing dataset*. If a model satisfies this criterion, we say that it is *generalizable*.
#
# If a model has 100% accuracy on the training dataset ($k=1$ in k-NN) but doesn't generalize to new data, then it isn't a very good model. We say that this model has *overfit* the data. On the other hand, it isn't difficult to see that we could also *underfit* the data (taking $k$ large in k-NN). In this case, the model isn't complex enough to have good accuracy on the training dataset.
#
# **Cross-validation** is a general method for assessing how the results of a predictive model (classification, regression,...) will *generalize* to an independent data set. In classification, cross-validation is a method for assessing how well the classification model will predict the class of points that weren't used to *train* the model.
#
# The idea of the method is simple:
# 1. Split the dataset into two groups: the training dataset and the test dataset.
# # + Train the model on the training dataset.
# # + Check the accuracy of the model on the test dataset.
#
# In practice, you have to decide how to split the data into groups (i.e. how large the groups should be). You might also want to repeat the experiment so that the assessment doesn't depend on the way in which you split the data into groups. We'll worry about this in a later lecture.
#
# For now, I just want you to conceptually understand how generalizable k-NN is as we vary the parameter, k.
#
# <img src="BiasVarianceTradeoff.png" width="500">
#
# $\qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad$
# source: [this blog](https://blog.cambridgecoding.com/2016/03/24/misleading-modelling-overfitting-cross-validation-and-the-bias-variance-trade-off/)
#
# As the model becomes more complex (k decreases), the accuracy always increases for the training dataset. But, at some point, it starts to overfit the data and the accuracy decreases for the test dataset! Cross validation techniques will allow us to find the sweet-spot for the parameter k! (Think: Goldilocks and the Three Bears.)
#
#
# Let's see this concept for the two moons dataset. You can use the *train_test_split* function in scikit-learn to split the dataset into a training dataset and a test dataset.
# + slideshow={"slide_type": "-"}
def detect_plot_dimension(X, h=0.02, b=0.05):
x_min, x_max = X[:, 0].min() - b, X[:, 0].max() + b
y_min, y_max = X[:, 1].min() - b, X[:, 1].max() + b
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
dimension = xx, yy
return dimension
def detect_decision_boundary(dimension, model):
xx, yy = dimension # unpack the dimensions
boundary = model.predict(np.c_[xx.ravel(), yy.ravel()])
boundary = boundary.reshape(xx.shape) # Put the result into a color plot
return boundary
def plot_decision_boundary(panel, dimension, boundary, colors=['#DADDED', '#FBD8D8']):
xx, yy = dimension # unpack the dimensions
panel.contourf(xx, yy, boundary, cmap=ListedColormap(colors), alpha=1)
panel.contour(xx, yy, boundary, colors="g", alpha=1, linewidths=0.5) # the decision boundary in green
def plot_dataset(panel, X, y, colors=["#EE3D34", "#4458A7"], markers=["x", "o"]):
panel.scatter(X[y == 1, 0], X[y == 1, 1], color=colors[0], marker=markers[0])
panel.scatter(X[y == 0, 0], X[y == 0, 1], color=colors[1], marker=markers[1])
def calculate_prediction_error(model, X, y):
yPred = model.predict(X)
score = round(metrics.accuracy_score(y, yPred), 2)
return score
def plot_prediction_error(panel, dimension, score, b=.3):
xx, yy = dimension # unpack the dimensions
panel.text(xx.max() - b, yy.min() + b, ('%.2f' % score).lstrip('0'), size=15, horizontalalignment='right')
def explore_fitting_boundaries(model, n_neighbors, datasets, width):
# determine the height of the plot given the aspect ration of each panel should be equal
height = float(width)/len(n_neighbors) * len(datasets.keys())
nrows = len(datasets.keys())
ncols = len(n_neighbors)
# set up the plot
figure, axes = plt.subplots(
nrows,
ncols,
figsize=(width, height),
sharex=True,
sharey=True
)
dimension = detect_plot_dimension(X, h=0.02) # the dimension each subplot based on the data
# Plotting the dataset and decision boundaries
i = 0
for n in n_neighbors:
model.n_neighbors = n
model.fit(datasets["Training Set"][0], datasets["Training Set"][1])
boundary = detect_decision_boundary(dimension, model)
j = 0
for d in datasets.keys():
try:
panel = axes[j, i]
except (TypeError, IndexError):
if (nrows * ncols) == 1:
panel = axes
elif nrows == 1: # if you only have one dataset
panel = axes[i]
elif ncols == 1: # if you only try one number of neighbors
panel = axes[j]
plot_decision_boundary(panel, dimension, boundary) # plot the decision boundary
plot_dataset(panel, X=datasets[d][0], y=datasets[d][1]) # plot the observations
score = calculate_prediction_error(model, X=datasets[d][0], y=datasets[d][1])
plot_prediction_error(panel, dimension, score, b=0.2) # plot the score
# make compacted layout
panel.set_frame_on(False)
panel.set_xticks([])
panel.set_yticks([])
# format the axis labels
if i == 0:
panel.set_ylabel(d)
if j == 0:
panel.set_title('k={}'.format(n))
j += 1
i += 1
plt.subplots_adjust(hspace=0, wspace=0) # make compacted layout
# + slideshow={"slide_type": "-"}
# Split into training and test sets
XTrain, XTest, yTrain, yTest = train_test_split(X, y, random_state=1, test_size=0.5)
# specify the model and settings
model = KNeighborsClassifier()
n_neighbors = [200, 99, 50, 23, 11, 1]
datasets = {
"Training Set": [XTrain, yTrain],
"Test Set": [XTest, yTest]
}
width = 20
explore_fitting_boundaries(model=model, n_neighbors=n_neighbors, datasets=datasets, width=width)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Conclusions and take-away
#
# 1. k-NN is a very simple method that can be used for classification. (It can be used for regression too! How?)
#
# # + Model accuracy (measured on the training dataset) and generalizability (measured on the testing dataset) are both important and often in contention with one another. Model accuracy can be measured using the confusion matrix, precision, recall, F-measure, or the Jaccard similarity score. Generalizability can be measured via cross validation.
#
# # + Picking parameters in models (such as k in k-NN) is non-trivial, but can be done via cross validation.
#
#
# ### Classification method preview
# For a quick preview of other classification methods, see the comparison [here](http://scikit-learn.org/stable/auto_examples/classification/plot_classifier_comparison.html).
#
# 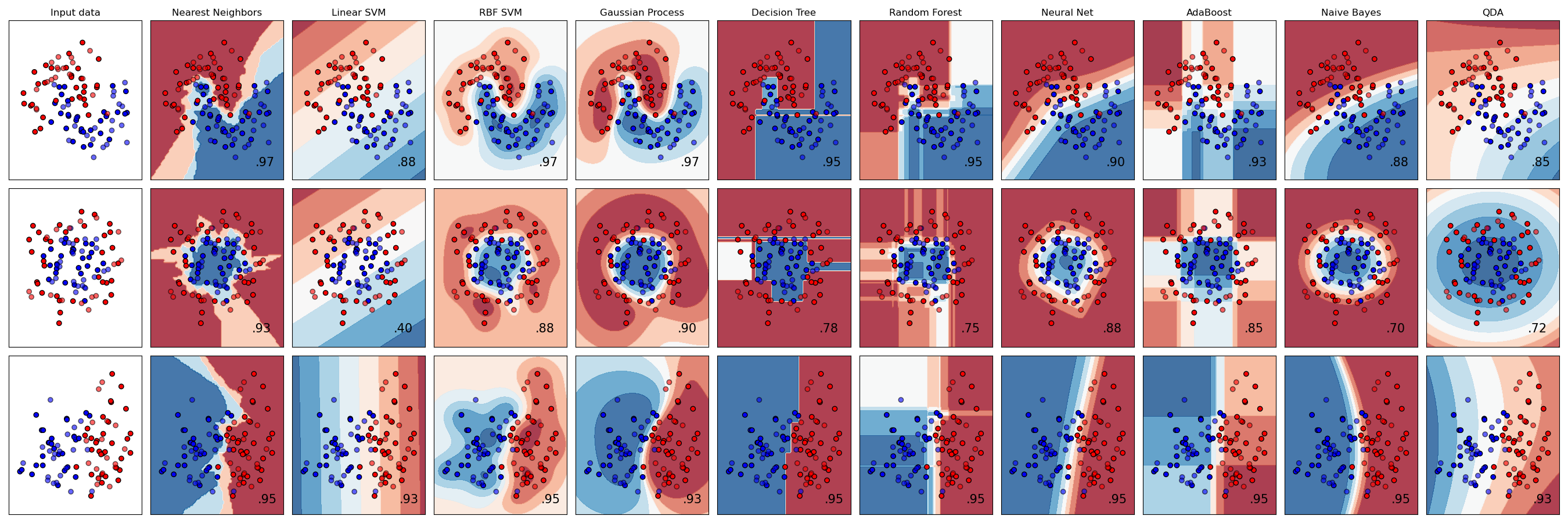
#
# -
|
13-Classification1/13b-Classification1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/inhauser/DSND_Term2/blob/master/Titanic.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="NCgaq2zDS1Wu"
# # Nova seção
# + colab={"base_uri": "https://localhost:8080/", "height": 175} id="TPoC3q7-TefO" outputId="4d355a8f-3330-491e-d1bf-d6756a8b4c56"
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
dados = pd.read_csv('/content/tested.csv')
dados.head()
dados = dados.drop(['Name','Ticket','Cabin','Embarked'], axis = 1)
dados.head()
dados = dados.set_index(['PassengerId'])
dados = dados.rename(columns = {'Survived' : 'target'}, inplace = False)
dados.head()
dados.describe()
dados.describe(include=['O'])
# + [markdown] id="gsHqQfBIXfg9"
# # Nova seção
# + colab={"base_uri": "https://localhost:8080/", "height": 571} id="zz3w8sViXhZ7" outputId="25692eb2-08dd-43e7-df14-8f378025eac8"
dados['Sex_F'] = np.where(dados['Sex'] == 'female', 1, 0)
dados['Pclass_1'] = np.where(dados['Pclass'] == 1,1,0)
dados['Pclass_2'] = np.where(dados['Pclass'] == 2,1,0)
dados['Pclass_3'] = np.where(dados['Pclass'] == 3,1,0)
# + id="TiD5O72IYKtm"
dados = dados.drop(['Pclass','Sex'], axis = 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="xp73Kr5gZNgD" outputId="96f3e6c6-eab9-454b-d674-a217579a1438"
dados.head()
# + colab={"base_uri": "https://localhost:8080/"} id="4FQYnYYMZT9F" outputId="e77322f1-a6de-4f02-f685-caff196f6cff"
dados.isnull().sum()
# + id="5_8Q8uuHZbfk"
dados.fillna(0, inplace = True)
# + colab={"base_uri": "https://localhost:8080/"} id="hm8E60asZumc" outputId="51e92661-7d50-4125-8ce0-43e454ed0594"
dados.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="8gSJw0-yZ2D2" outputId="97dba6cc-c240-4347-9e35-e7e7e45e36dc"
x_train, x_test, y_train, y_test = train_test_split(dados.drop(['target'], axis = 1),
dados["target"],
test_size = 0.3,
random_state = 1234)
[{'treino': x_train.shape},
{'teste': x_test.shape}]
# + colab={"base_uri": "https://localhost:8080/"} id="55jD7dvQazbX" outputId="bf7805a2-eedb-4fd8-9e78-25003dbf9017"
from sklearn.ensemble import RandomForestClassifier
rndforest = RandomForestClassifier(n_estimators = 1000,
criterion='gini',
max_depth = 5)
rndforest.fit(x_train, y_train)
# + id="pLjrDO8MdDfi"
probabilidade = rndforest.predict_proba(dados.drop('target', axis = 1))[:,1]
classificacao = rndforest.predict(dados.drop('target',axis = 1))
# + id="zfzz5s1WdzBm"
dados['probabilidade'] = probabilidade
dados['classificacao'] = classificacao
# + colab={"base_uri": "https://localhost:8080/", "height": 455} id="uwq6h_NOd9gK" outputId="db0ec772-9864-432e-d611-9b0c25abe33f"
dados
|
Titanic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 5GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# -
import sklearn
import matplotlib.pyplot as plt
# %matplotlib inline
import pandas
from sklearn.model_selection import train_test_split
import numpy
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
Tweet= pd.read_csv("/kaggle/input/twitter-airline-sentiment/Tweets.csv")
Tweet.head()
# +
import re
import nltk
from nltk.corpus import stopwords
# 数据清洗
def tweet_to_words(raw_tweet):
letters_only = re.sub("[^a-zA-Z]", " ",raw_tweet)
words = letters_only.lower().split()
stops = set(stopwords.words("english"))
meaningful_words = [w for w in words if not w in stops]
return( " ".join( meaningful_words ))
def clean_tweet_length(raw_tweet):
letters_only = re.sub("[^a-zA-Z]", " ",raw_tweet)
words = letters_only.lower().split()
stops = set(stopwords.words("english"))
meaningful_words = [w for w in words if not w in stops]
return(len(meaningful_words))
# 将标签转换成数字
Tweet['sentiment']=Tweet['airline_sentiment'].apply(lambda x: 0 if x=='negative' else 1)
Tweet.fillna('-1', inplace=True)
#用的小数据集
Tweet['clean_tweet'] = Tweet[['negativereason', 'name', 'text']].apply(lambda x: ' '.join(x), axis=1)
Tweet['clean_tweet']=Tweet['clean_tweet'].apply(lambda x: tweet_to_words(x))
Tweet['Tweet_length']=Tweet['text'].apply(lambda x: clean_tweet_length(x))
# -
Tweet.head()
train,test = train_test_split(Tweet,test_size=0.2,random_state=42)
# 转换成list,方便特征提取
train_clean_tweet=[]
for tweet in train['clean_tweet']:
train_clean_tweet.append(tweet)
test_clean_tweet=[]
for tweet in test['clean_tweet']:
test_clean_tweet.append(tweet)
from sklearn.feature_extraction.text import CountVectorizer
v = CountVectorizer(analyzer = "word")
train_features= v.fit_transform(train_clean_tweet)
test_features=v.transform(test_clean_tweet)
# +
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC, LinearSVC, NuSVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier, GradientBoostingClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.metrics import accuracy_score
from xgboost import XGBClassifier
from sklearn.metrics import accuracy_score
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
import matplotlib.pyplot as plt
from itertools import cycle
from sklearn import svm, datasets
from sklearn.metrics import roc_curve, auc
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import label_binarize
from sklearn.multiclass import OneVsRestClassifier
from scipy import interp
from sklearn.metrics import roc_auc_score
from sklearn.metrics import classification_report
# +
Classifiers = [
LogisticRegression(C=0.000000001,solver='liblinear',max_iter=200),
KNeighborsClassifier(3),
SVC(kernel="rbf", C=0.025, probability=True),
DecisionTreeClassifier(),
RandomForestClassifier(n_estimators=200),
AdaBoostClassifier(),
GaussianNB(),
XGBClassifier(),
MLPClassifier(solver='sgd', alpha=1e-5,
hidden_layer_sizes=(5, 2), random_state=1, max_iter=500),
GradientBoostingClassifier(random_state=0)
]#gpu不能一次性加载这么多模型兵训练
# -
dense_features=train_features.toarray()
dense_test= test_features.toarray()
Accuracy=[]
Model=[]
for classifier in Classifiers:
try:
fit = classifier.fit(train_features,train['sentiment'])
pred = fit.predict(test_features)
except Exception:
fit = classifier.fit(dense_features,train['sentiment'])
pred = fit.predict(dense_test)
predictions = [round(value) for value in pred]
accuracy = accuracy_score(test['sentiment'],predictions)
print(classification_report(test.sentiment, predictions, labels=[0,2,4]))
Accuracy.append(accuracy)
Model.append(classifier.__class__.__name__)
print('Accuracy of '+classifier.__class__.__name__+ ' is: '+str(accuracy))
|
others/sentiment-analysis-kernel-airline-baseline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import zarr
import sys
import allel
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
def build_sample_zarr(input, output):
allel.vcf_to_zarr(
input=input,
output=output,
compressor=zarr.Blosc(cname='zstd', clevel=5, shuffle=0),
overwrite=True,
fields=['*'],
alt_number=3,
chunk_length=2**18,
log=sys.stdout,
)
input_vcf = 'merged.vcf'
output_path = 'merged.zarr'
build_sample_zarr(input_vcf, output_path)
callset = zarr.open_group('merged.zarr', mode='r')
callset
list(callset['calldata'].keys())
samples = callset['samples'][:]
samples
callset['calldata/DP'][0].shape
callset['calldata/GT'][:].shape
callset['calldata/AD'][0][0]
chrom = callset['variants/CHROM'][:]
chromunq = np.unique(chrom)
len(chromunq)
chromunq
gt = allel.GenotypeArray(callset['calldata/GT'])
gt
ismiss = gt.is_missing()
ismiss
persnp_miss = np.count_nonzero(ismiss, axis=1)/121
np.count_nonzero(persnp_miss <= 0.15)
plt.hist(persnp_miss, bins=20);
|
notebooks/20180607_44_uniref_vcf_to_zarr_plus_initial_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## DELETE THIS SESSION AFTER EDITING
# + language="javascript"
# IPython.notebook.kernel.execute('nb_name = "' + IPython.notebook.notebook_name + '"')
# //DELETE THIS CELL AFTER EDITING THE NOTEBOOK
# -
raise Exception(f'{nb_name} template has been left inaltered. Delete notebook or alter file and delete first notebook session')
#DELETE THIS CELL AFTER EDITING THE NOTEBOOK
# ## Nbdev and Docs Imports
# +
#default_exp fit
# +
#hide
from nbdev.showdoc import *
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append('..') #appends project root to path in order to import project packages since `noteboks_dev` is not on the root
#DO NOT EDIT
# +
#Internal Imports
#imports that are going to be used only during development and are not intended to be loaded inside the generated modules.
#for example: use imported modules to generate graphs for documentation, but lib is unused in actual package
#import ...
# -
# # fit
# > Module containing feature modeling/model fitting functionalities
#
# + [markdown] heading_collapsed=true
# ## Dev comments
# + [markdown] hidden=true
# ### TODOs
# + [markdown] hidden=true
# - [X] TODO: do something
# - [ ] TODO: do something else
# + [markdown] hidden=true
# ### <comment section 2>
# -
# ## External imports
# > imports that are intended to be loaded in the actual modules e.g.: module dependencies
# +
#export
#import ...
# -
# ## Code session
# ### <func 1> -
#export
def func1():
pass
# `func1` comments and usage examples for documentation:
# ### <func 2> -
#export
def func2():
pass
# `func2` comments and usage examples for documentation
# ### <class 1> -
#export
class Class1:
pass
# `Class1` comments and usage examples for documentation
# ## Export -
#hide
from nbdev.export import notebook2script
notebook2script()
|
notebooks_dev/03_fit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="HnU0fNSuG2aD"
# # Lab: Feature Analysis Using TensorFlow Data Validation and Facets
# + [markdown] colab_type="text" id="iVkPBosnIFlu"
# **Learning Objectives:**
# 1. Use TFRecords to load record-oriented binary format data
# 2. Use TFDV to generate statistics and Facets to visualize the data
# 3. Use the TFDV widget to answer questions
# 4. Analyze label distribution for subset groups
#
#
# ## Introduction
#
# Bias can manifest in any part of a typical machine learning pipeline, from an unrepresentative dataset, to learned model representations, to the way in which the results are presented to the user. Errors that result from this bias can disproportionately impact some users more than others.
#
# [TensorFlow Data Validation](https://www.tensorflow.org/tfx/data_validation/get_started) (TFDV) is one tool you can use to analyze your data to find potential problems in your data, such as missing values and data imbalances - that can lead to Fairness disparities. The TFDV tool analyzes training and serving data to compute descriptive statistics, infer a schema, and detect data anomalies. [Facets Overview](https://pair-code.github.io/facets/) provides a succinct visualization of these statistics for easy browsing. Both the TFDV and Facets are tools that are part of the [Fairness Indicators](https://www.tensorflow.org/tfx/fairness_indicators).
#
# In this notebook, we use TFDV to compute descriptive statistics that provide a quick overview of the data in terms of the features that are present and the shapes of their value distributions. We use Facets Overview to visualize these statistics using the Civil Comments dataset.
#
# Each learning objective will correspond to a __#TODO__ in this student lab notebook -- try to complete this notebook first and then review the [solution notebook](../solutions/adv_tfdv_facets.ipynb).
#
# -
# ## Set up environment variables and load necessary libraries
# We will start by importing the necessary dependencies for the libraries we'll be using in this exercise. First, run the cell below to install Fairness Indicators.
#
# **NOTE:** You can ignore the "pip" being invoked by an old script wrapper, as it will not affect the lab's functionality.
#
# !pip3 install fairness-indicators --user
# <strong>Restart the kernel</strong> after you do a pip3 install (click on the <strong>Restart the kernel</strong> button above).
# + [markdown] colab_type="text" id="mdLlKWbIlxYH"
# Next, import all the dependencies we'll use in this exercise, which include Fairness Indicators, TensorFlow Data Validation (tfdv), and the What-If tool (WIT) Facets Overview.
# + colab={} colab_type="code" id="6E__x2XkJDFW"
# # %tensorflow_version 2.x
import sys, os
import warnings
warnings.filterwarnings('ignore')
#os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3' # Ignore deprecation warnings
import tempfile
import apache_beam as beam
import numpy as np
import pandas as pd
from datetime import datetime
import tensorflow_hub as hub
import tensorflow as tf
import tensorflow_model_analysis as tfma
import tensorflow_data_validation as tfdv
from tensorflow_model_analysis.addons.fairness.post_export_metrics import fairness_indicators
from tensorflow_model_analysis.addons.fairness.view import widget_view
from fairness_indicators.examples import util
import warnings
warnings.filterwarnings("ignore")
from witwidget.notebook.visualization import WitConfigBuilder
from witwidget.notebook.visualization import WitWidget
print(tf.version.VERSION)
print(tf) # This statement shows us what version of Python we are currently running.
# + [markdown] colab_type="text" id="J3R2QWkru1WN"
# ### About the Civil Comments dataset
#
# Click below to learn more about the Civil Comments dataset, and how we've preprocessed it for this exercise.
# + [markdown] colab_type="text" id="ZZswcJJMCDjU"
# The Civil Comments dataset comprises approximately 2 million public comments that were submitted to the Civil Comments platform. [Jigsaw](https://jigsaw.google.com/) sponsored the effort to compile and annotate these comments for ongoing [research](https://arxiv.org/abs/1903.04561); they've also hosted competitions on [Kaggle](https://www.kaggle.com/c/jigsaw-unintended-bias-in-toxicity-classification) to help classify toxic comments as well as minimize unintended model bias.
#
# #### Features
#
# Within the Civil Comments data, a subset of comments are tagged with a variety of identity attributes pertaining to gender, sexual orientation, religion, race, and ethnicity. Each identity annotation column contains a value that represents the percentage of annotators who categorized a comment as containing references to that identity. Multiple identities may be present in a comment.
#
# **NOTE:** These identity attributes are intended *for evaluation purposes only*, to assess how well a classifier trained solely on the comment text performs on different tag sets.
#
# To collect these identity labels, each comment was reviewed by up to 10 annotators, who were asked to indicate all identities that were mentioned in the comment. For example, annotators were posed the question: "What genders are mentioned in the comment?", and asked to choose all of the following categories that were applicable.
#
# * Male
# * Female
# * Transgender
# * Other gender
# * No gender mentioned
#
# **NOTE:** *We recognize the limitations of the categories used in the original dataset, and acknowledge that these terms do not encompass the full range of vocabulary used in describing gender.*
#
# Jigsaw used these ratings to generate an aggregate score for each identity attribute representing the percentage of raters who said the identity was mentioned in the comment. For example, if 10 annotators reviewed a comment, and 6 said that the comment mentioned the identity "female" and 0 said that the comment mentioned the identity "male," the comment would receive a `female` score of `0.6` and a `male` score of `0.0`.
#
# **NOTE:** For the purposes of annotation, a comment was considered to "mention" gender if it contained a comment about gender issues (e.g., a discussion about feminism, wage gap between men and women, transgender rights, etc.), gendered language, or gendered insults. Use of "he," "she," or gendered names (e.g., Donald, Margaret) did not require a gender label.
#
# #### Label
#
# Each comment was rated by up to 10 annotators for toxicity, who each classified it with one of the following ratings.
#
# * Very Toxic
# * Toxic
# * Hard to Say
# * Not Toxic
#
# Again, Jigsaw used these ratings to generate an aggregate toxicity "score" for each comment (ranging from `0.0` to `1.0`) to serve as the [label](https://developers.google.com/machine-learning/glossary?utm_source=Colab&utm_medium=fi-colab&utm_campaign=fi-practicum&utm_content=glossary&utm_term=label#label), representing the fraction of annotators who labeled the comment either "Very Toxic" or "Toxic." For example, if 10 annotators rated a comment, and 3 of them labeled it "Very Toxic" and 5 of them labeled it "Toxic", the comment would receive a toxicity score of `0.8`.
#
# **NOTE:** For more information on the Civil Comments labeling schema, see the [Data](https://www.kaggle.com/c/jigsaw-unintended-bias-in-toxicity-classification/data) section of the Jigsaw Untended Bias in Toxicity Classification Kaggle competition.
#
# ### Preprocessing the data
# For the purposes of this exercise, we converted toxicity and identity columns to booleans in order to work with our neural net and metrics calculations. In the preprocessed dataset, we considered any value ≥ 0.5 as True (i.e., a comment is considered toxic if 50% or more crowd raters labeled it as toxic).
#
# For identity labels, the threshold 0.5 was chosen and the identities were grouped together by their categories. For example, if one comment has `{ male: 0.3, female: 1.0, transgender: 0.0, heterosexual: 0.8, homosexual_gay_or_lesbian: 1.0 }`, after processing, the data will be `{ gender: [female], sexual_orientation: [heterosexual, homosexual_gay_or_lesbian] }`.
#
# **NOTE:** Missing identity fields were converted to False.
#
# + [markdown] colab_type="text" id="0YNqAJW5JjZD"
# ### Use TFRecords to load record-oriented binary format data
#
#
#
#
# -------------------------------------------------------------------------------------------------------
#
# The [TFRecord format](https://www.tensorflow.org/tutorials/load_data/tfrecord) is a simple [Protobuf](https://developers.google.com/protocol-buffers)-based format for storing a sequence of binary records. It gives you and your machine learning models to handle arbitrarily large datasets over the network because it:
# 1. Splits up large files into 100-200MB chunks
# 2. Stores the results as serialized binary messages for faster ingestion
#
# If you already have a dataset in TFRecord format, you can use the tf.keras.utils functions for accessing the data (as you will below!). If you want to practice creating your own TFRecord datasets you can do so outside of this lab by [viewing the documentation here](https://www.tensorflow.org/tutorials/load_data/tfrecord).
#
# #### TODO 1: Use the utility functions tf.keras to download and import our datasets
# Run the following cell to download and import the training and validation preprocessed datasets.
# + colab={} colab_type="code" id="duPWGTQAvYKK"
download_original_data = False #@param {type:"boolean"}
# TODO 1
# TODO: Your code goes here
# The identity terms list will be grouped together by their categories
# (see 'IDENTITY_COLUMNS') on threshould 0.5. Only the identity term column,
# text column and label column will be kept after processing.
# TODO: Your code goes here
# TODO 1a
# TODO: Your code goes here
# + [markdown] colab_type="text" id="aLup7wY0_Q3K"
# ### Use TFDV to generate statistics and Facets to visualize the data
#
#
# TensorFlow Data Validation supports data stored in a TFRecord file, a CSV input format, with extensibility for other common formats. You can find the available data decoders [here](https://github.com/tensorflow/data-validation/tree/master/tensorflow_data_validation/coders). In addition, TFDV provides the [tfdv.generate_statistics_from_dataframe](https://www.tensorflow.org/tfx/data_validation/api_docs/python/tfdv/generate_statistics_from_dataframe) utility function for users with in-memory data represented as a pandas DataFrame.
#
# In addition to computing a default set of data statistics, TFDV can also compute statistics for semantic domains (e.g., images, text). To enable computation of semantic domain statistics, pass a tfdv.StatsOptions object with enable_semantic_domain_stats set to True to tfdv.generate_statistics_from_tfrecord.Before we train the model, let's do a quick audit of our training data using [TensorFlow Data Validation](https://www.tensorflow.org/tfx/data_validation/get_started), so we can better understand our data distribution.
#
# #### TODO 2: Use TFDV to get quick statistics on your dataset
#
# The following cell may take 2–3 minutes to run. **NOTE:** Please ignore the deprecation warnings.
# + colab={} colab_type="code" id="vkzcE_g8_m_h"
# TODO 2
# The computation of statistics using TFDV. The returned value is a DatasetFeatureStatisticsList protocol buffer.
# TODO: Your code goes here
# TODO 2a
# A visualization of the statistics using Facets Overview.
# TODO: Your code goes here
# + [markdown] colab_type="text" id="wZU1Djze6E-s"
# ### TODO 3: Use the TensorFlow Data Validation widget above to answer the following questions.
# + [markdown] colab_type="text" id="ne2_vKAb-XGD"
# #### **1. How many total examples are in the training dataset?**
# + [markdown] colab_type="text" id="UFBqqnRD-Zkj"
# #### Solution
#
# See below solution.
#
# + [markdown] colab_type="text" id="XSkOfchI-arC"
# **There are 1.08 million total examples in the training dataset.**
#
# The count column tells us how many examples there are for a given feature. Each feature (`sexual_orientation`, `comment_text`, `gender`, etc.) has 1.08 million examples. The missing column tells us what percentage of examples are missing that feature.
#
# 
#
# Each feature is missing from 0% of examples, so we know that the per-feature example count of 1.08 million is also the total number of examples in the dataset.
# + [markdown] colab_type="text" id="_PgFNm6sAZB2"
# #### **2. How many unique values are there for the `gender` feature? What are they, and what are the frequencies of each of these values?**
#
# **NOTE #1:** `gender` and the other identity features (`sexual_orientation`, `religion`, `disability`, and `race`) are included in this dataset for evaluation purposes only, so we can assess model performance on different identity slices. The only feature we will use for model training is `comment_text`.
#
# **NOTE #2:** *We recognize the limitations of the categories used in the original dataset, and acknowledge that these terms do not encompass the full range of vocabulary used in describing gender.*
# + [markdown] colab_type="text" id="6KmrCS-uAz0s"
# #### Solution
#
# See below solution.
# + [markdown] colab_type="text" id="wkc7P1nvA4cw"
# The **unique** column of the **Categorical Features** table tells us that there are 4 unique values for the `gender` feature.
#
# To view the 4 values and their frequencies, we can click on the **SHOW RAW DATA** button:
#
# 
#
# The raw data table shows that there are 32,208 examples with a gender value of `female`, 26,758 examples with a value of `male`, 1,551 examples with a value of `transgender`, and 4 examples with a value of `other gender`.
#
#
#
#
# + [markdown] colab_type="text" id="NDUO57bdNUQR"
# **NOTE:** As described [earlier](#scrollTo=J3R2QWkru1WN), a `gender` feature can contain zero or more of these 4 values, depending on the content of the comment. For example, a comment containing the text "I am a transgender man" will have both `transgender` and `male` as `gender` values, whereas a comment that does not reference gender at all will have an empty/false `gender` value.
# + [markdown] colab_type="text" id="wX62Ktwp-qoF"
# #### **3. What percentage of total examples are labeled toxic? Overall, is this a class-balanced dataset (relatively even split of examples between positive and negative classes) or a class-imbalanced dataset (majority of examples are in one class)?**
#
# **NOTE:** In this dataset, a `toxicity` value of `0` signifies "not toxic," and a `toxicity` value of `1` signifies "toxic."
# + [markdown] colab_type="text" id="IvvxNMgM-6A2"
# #### Solution
#
# See below solution.
# + [markdown] colab_type="text" id="QmCtkzZqOvC2"
# **7.98 percent of examples are toxic.**
#
# Under **Numeric Features**, we can see the distribution of values for the `toxicity` feature. 92.02% of examples have a value of 0 (which signifies "non-toxic"), so 7.98% of examples are toxic.
#
# 
#
# This is a [**class-imbalanced dataset**](https://developers.google.com/machine-learning/glossary?utm_source=Colab&utm_medium=fi-colab&utm_campaign=fi-practicum&utm_content=glossary&utm_term=class-imbalanced-dataset#class-imbalanced-dataset), as the overwhelming majority of examples (over 90%) are classified as nontoxic.
# -
# Notice that there is one numeric feature (count of toxic comments) and six categorical features.
# ### TODO 4: Analyze label distribution for subset groups
# + [markdown] colab_type="text" id="9MGLCsVhGWz0"
# Run the following code to analyze label distribution for the subset of examples that contain a `gender` value**
#
#
# **NOTE:** *The cell run should for just a few minutes*
# + cellView="form" colab={} colab_type="code" id="f5pEWIkgLTKz"
#@title Calculate label distribution for gender-related examples
raw_dataset = tf.data.TFRecordDataset(train_tf_file)
toxic_gender_examples = 0
nontoxic_gender_examples = 0
# TODO 4
# There are 1,082,924 examples in the dataset
# TODO: Your code goes here
# TODO 4a
# TODO: Your code goes here
# + [markdown] colab_type="text" id="WJag4cEKNINy"
# #### **What percentage of `gender` examples are labeled toxic? Compare this percentage to the percentage of total examples that are labeled toxic from #3 above. What, if any, fairness concerns can you identify based on this comparison?**
# + [markdown] colab_type="text" id="-J4hbOhgHZid"
# #### Solution
#
# Click below for one possible solution.
# + [markdown] colab_type="text" id="2KK3VWzkHmJ7"
# There are 7,189 gender-related examples that are labeled toxic, which represent 14.7% of all gender-related examples.
#
# The percentage of gender-related examples that are toxic (14.7%) is nearly double the percentage of toxic examples overall (7.98%). In other words, in our dataset, gender-related comments are almost two times more likely than comments overall to be labeled as toxic.
#
# This skew suggests that a model trained on this dataset might learn a correlation between gender-related content and toxicity. This raises fairness considerations, as the model might be more likely to classify nontoxic comments as toxic if they contain gender terminology, which could lead to [disparate impact](https://developers.google.com/machine-learning/glossary?utm_source=Colab&utm_medium=fi-colab&utm_campaign=fi-practicum&utm_content=glossary&utm_term=disparate-impact#disparate-impact) for gender subgroups.
# -
# Copyright 2020 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
|
courses/business/managingmlprojects/labs/bias_tfdv_facets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pathlib import Path as path
import os
# +
location = path.cwd()
print(f'Location: {location}')
user_home_dir = path.home()
print(f'Home: {user_home_dir}')
join_path = location.joinpath('testing', 'python.py')
print(f'Join Path (example, such as ): {join_path}')
join_path.name, join_path.stem, join_path.suffix, join_path.parent, join_path.anchor,join_path.parent.parent, join_path.drive
# -
join_path.with_suffix('.md'), join_path.with_name('python.md'), list(join_path.parents), join_path.exists()
list(location.glob('*.ipynb'))
list(location.rglob('*'))
# ## Directories and files can be deleted using .rmdir() and .unlink() respectively. (Again, be careful!)
list(location.iterdir())
# cheking whether it is a file or directory
location.is_dir()
location.is_file()
# +
import collections as collect
file_types = collect.Counter(f.suffix for f in location.iterdir())
file_types
# -
# # Last modified file
# +
from datetime import datetime
time, file_path = max((f.stat().st_mtime, f) for f in location.iterdir())
time = datetime.fromtimestamp(time)
print(f"{time}: {file_path}")
# -
file = location / 'pathlib_module_tutorials.ipynb'
print(file.stat())
file.is_file(), file.exists()
file.stat().st_size # in bytes
file.stat().st_size/1024 # in KB
# # Create a Unique File Name
# +
def unique_path(directory, name_pattern):
counter = 0
while True:
counter += 1
path = directory / name_pattern.format(counter)
if not path.exists():
return path
new_file = unique_path(location, 'test{:03d}.txt')
# If the directory already contains the files test001.txt and test002.txt, the above code will set path to test003.txt.
new_file
# -
location/'pathlib_module_tutorials.ipynb'/'test'
|
Simulations/THz/testing/.ipynb_checkpoints/pathlib_module_tutorials-checkpoint.ipynb
|