repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
davidteather/TikTok-Api | api | 463 | [FEATURE_REQUEST] - Account Names of Commenters | **Requested Feature**
Hi! I'm wondering if it's possible to get a feature that scrapes the account names of commenters for an inputted Tiktok.
(or if there is already a function that does this). Would be incredibly helpful!
**Additional context**
One of the use cases is after finding a Tiktok related to recruiting for a right wing extremist group, with hundreds of commenters, would like to be able to scrape the account handles of the commenters for Open Source Investigation / OSINT purposes. | closed | 2021-01-07T21:24:55Z | 2022-02-14T03:08:06Z | https://github.com/davidteather/TikTok-Api/issues/463 | [
"feature_request"
] | ETedward | 2 |
comfyanonymous/ComfyUI | pytorch | 7,078 | load image No components choose image | ### Your question
After updating to the new version.0.3.19 load image No components choose image
### Logs
```powershell
```
### Other
_No response_ | closed | 2025-03-05T03:33:23Z | 2025-03-05T14:40:44Z | https://github.com/comfyanonymous/ComfyUI/issues/7078 | [
"User Support"
] | wangnima007 | 5 |
dbfixtures/pytest-postgresql | pytest | 918 | modulenotfounderror while using load in factory | ### What action do you want to perform
When I try to load an sql file through the load argument of the factory :
`postgresql_in_docker = factories.postgresql_noproc(user = DBConfig().user, password=DBConfig().password)
postgresql_mok = factories.postgresql("postgresql_in_docker",
dbname="mydb", load=["test_base.sql", ])
`
I get this error :
> ERROR tests/test_decorators.py::test_decorator_fk_table_not_exists - ModuleNotFoundError: No module named 'test_base'
However if I write :
`postgresql_in_docker = factories.postgresql_noproc(user = DBConfig().user, password=DBConfig().password)
postgresql_mok = factories.postgresql("postgresql_in_docker",
dbname="mydb", load=["./test_base.sql", ])
`
Note the **./** in front of the filename, in that case everything works well
I just upgraded my version to 5.1.1 and the issue is still there.
G.
| closed | 2024-03-07T13:10:50Z | 2024-03-11T14:49:58Z | https://github.com/dbfixtures/pytest-postgresql/issues/918 | [] | GFuhr | 2 |
ibis-project/ibis | pandas | 10,135 | bug: polars flatten does not work as expected | ### What happened?
Following up on https://github.com/ibis-project/ibis/issues/9995
Thanks @gforsyth and @cpcloud for the quick fix of the above issue. However I don't think it totally fixed the issue.
I am trying to get an `array<array<int64>>` to become an `array<int64>`
Using Polars:
```python
import ibis
ibis.set_backend("polars")
t = ibis.memtable([
{
"arr": [[1, 5, 7], [3,4]]
},
])
t.arr.flatten()
```
```
┏━━━━━━━━━━━━━━━━━━━┓
┃ ArrayFlatten(arr) ┃
┡━━━━━━━━━━━━━━━━━━━┩
│ array<int64> │
├───────────────────┤
│ [1, 5, ... +1] │
│ [3, 4] │
└───────────────────┘
```
Using pandas:
```python
ibis.set_backend("pandas")
t = ibis.memtable([
{
"arr": [[1, 5, 7], [3,4]]
},
])
t.arr.flatten()
```
```
┏━━━━━━━━━━━━━━━━━━━┓
┃ ArrayFlatten(arr) ┃
┡━━━━━━━━━━━━━━━━━━━┩
│ array<int64> │
├───────────────────┤
│ [1, 5, ... +3] │
└───────────────────┘
```
Notice how using `pandas` I get a single line back with a single "flattened" array, while using `polars`, I get 2 rows back. Hence I think something is still missing somewhere, unless there is another way to achieve this using ibis or I misunderstand the flatten function. Anyways, I believe the result should be the same by using pandas or polars.
I expect the result to be `[1, 5, 7, 3, 4]` in both cases.
### What version of ibis are you using?
9.5.0
### What backend(s) are you using, if any?
polars 1.5.0
### Relevant log output
_No response_
### Code of Conduct
- [X] I agree to follow this project's Code of Conduct | closed | 2024-09-16T06:48:57Z | 2024-09-23T13:27:50Z | https://github.com/ibis-project/ibis/issues/10135 | [
"bug"
] | GLeurquin | 9 |
ScottfreeLLC/AlphaPy | scikit-learn | 27 | Model.yml encoding error | Running this via Windows WSL (winbash).
Python 3.7
Run mflow and get this error:
Traceback (most recent call last):
File "/home/d/.local/bin/mflow", line 8, in <module>
sys.exit(main())
File "/home/d/.local/lib/python3.7/site-packages/alphapy/market_flow.py", line 412, in main
model_specs = get_model_config()
File "/home/d/.local/lib/python3.7/site-packages/alphapy/model.py", line 274, in get_model_config
raise ValueError("model.yml features:encoding:type %s unrecognized" % encoder)
ValueError: model.yml features:encoding:type factorize unrecognized
This is from the config entry in console:
[03/01/20 01:54:19] INFO ********************************************************************************
[03/01/20 01:54:19] INFO MarketFlow Start
[03/01/20 01:54:19] INFO ********************************************************************************
[03/01/20 01:54:19] INFO Training Date: 1900-01-01
[03/01/20 01:54:19] INFO Prediction Date: 2020-03-01
[03/01/20 01:54:19] INFO MarketFlow Configuration
[03/01/20 01:54:19] INFO Getting Features
[03/01/20 01:54:19] INFO No Features Found
[03/01/20 01:54:19] INFO Defining Groups
[03/01/20 01:54:19] INFO Added: {'googl', 'fb', 'aapl', 'amzn', 'nflx'}
[03/01/20 01:54:19] INFO Defining Aliases
[03/01/20 01:54:19] INFO Getting System Parameters
[03/01/20 01:54:19] INFO Defining AlphaPy Variables [phigh, plow]
[03/01/20 01:54:19] INFO Defining User Variables
[03/01/20 01:54:19] INFO No Variables Found
[03/01/20 01:54:19] INFO Getting Variable Functions
[03/01/20 01:54:19] INFO No Variable Functions Found
[03/01/20 01:54:19] INFO MARKET PARAMETERS:
[03/01/20 01:54:19] INFO api_key = None
[03/01/20 01:54:19] INFO api_key_name = None
[03/01/20 01:54:19] INFO create_model = False
[03/01/20 01:54:19] INFO data_fractal = 1d
[03/01/20 01:54:19] INFO data_history = 500
[03/01/20 01:54:19] INFO features = {}
[03/01/20 01:54:19] INFO forecast_period = 1
[03/01/20 01:54:19] INFO fractal = 1d
[03/01/20 01:54:19] INFO lag_period = 1
[03/01/20 01:54:19] INFO leaders = []
[03/01/20 01:54:19] INFO predict_history = 50
[03/01/20 01:54:19] INFO schema = yahoo
[03/01/20 01:54:19] INFO subject = stock
[03/01/20 01:54:19] INFO subschema = None
[03/01/20 01:54:19] INFO system = {'name': 'closer', 'holdperiod': 0, 'longentry': 'hc', 'longexit': None, 'shortentry': 'lc', 'shortexit': None, 'scale': False}
| closed | 2020-03-01T02:16:40Z | 2020-03-03T01:57:46Z | https://github.com/ScottfreeLLC/AlphaPy/issues/27 | [] | thegamecat | 3 |
thunlp/OpenPrompt | nlp | 203 | Two placeholders are two restrictive. | Hi there,
I am working on a template that has three placeholders but InputExample only supports "text_a" and "text_b". I wonder if you can adapt it to more placeholders and let us define the names of the placehoders, instead of just "text_a" and "text_b". This will make it way more flexible. Thanks! | open | 2022-10-19T22:29:30Z | 2022-11-11T06:20:34Z | https://github.com/thunlp/OpenPrompt/issues/203 | [] | zihaohe123 | 1 |
dbfixtures/pytest-postgresql | pytest | 449 | documentation / new user issue: process fixture does not take kwarg 'load' | I'm reading the [intro documentation on pypi](https://pypi.org/project/pytest-postgresql/):
> The process fixture performs the load once per test session, and loads the data into the template database. Client fixture then creates test database out of the template database each test, which significantly speeds up the tests.
```
postgresql_my_proc = factories.postgresql_proc(
load=["schemafile.sql", "otherschema.sql", "import.path.to.function", "import.path.to:otherfunction", load_this]
)
```
but when trying to give the `load` keyword to `postgresql_proc`:
```
TypeError: postgresql_proc() got an unexpected keyword argument 'load'
```
it looks like `load` is not a kwarg to the `postgresql_proc()` process-based fixture, only to the `postgresql()` client-based fixture:
```
help(factories.postgresql_proc)
Help on function postgresql_proc in module pytest_postgresql.factories:
postgresql_proc(executable: str = None, host: str = None, port: Union[str, int, Iterable] = -1, user: str = None, password: str = None, options: str = '', startparams: str = None, unixsocketdir: str = None, logs_prefix: str = '') -> Callable[[_pytest.fix
tures.FixtureRequest, _pytest.tmpdir.TempdirFactory], pytest_postgresql.executor.PostgreSQLExecutor]
Postgresql process factory.
:param str executable: path to postgresql_ctl
:param str host: hostname
:param str|int|tuple|set|list port:
exact port (e.g. '8000', 8000)
randomly selected port (None) - any random available port
-1 - command line or pytest.ini configured port
[(2000,3000)] or (2000,3000) - random available port from a given range
[{4002,4003}] or {4002,4003} - random of 4002 or 4003 ports
[(2000,3000), {4002,4003}] - random of given range and set
:param str user: postgresql username
:param str options: Postgresql connection options
:param str startparams: postgresql starting parameters
:param str unixsocketdir: directory to create postgresql's unixsockets
:param str logs_prefix: prefix for log filename
:rtype: func
:returns: function which makes a postgresql process
```
So the documentation seems to be out of date or something right now.
Also, if you could let me know the best way to have a pytest-postgresql db running throughout the pytest, that loads my schema file when first created, that would be helpful to me. Should I be using startparams maybe? | closed | 2021-06-17T16:03:29Z | 2021-06-18T08:26:06Z | https://github.com/dbfixtures/pytest-postgresql/issues/449 | [] | sinback | 2 |
dadadel/pyment | numpy | 46 | list, tuple, dict default param values are not parsed correctly | When a function has parameters with default values that are list, dictionary or tuple, Pyment will just consider several parameters splitting on coma.
The following python code:
```python
def func1(param1=[1, None, "hehe"]):
pass
def func2(param1=(1, None, "hehe")):
pass
def func3(param1={0: 1, "a": None}):
pass
```
Will produce the patch:
```diff
# Patch generated by Pyment v0.3.2-dev4
--- a/issue46.py
+++ b/issue46.py
@@ -1,9 +1,29 @@
def func1(param1=[1, None, "hehe"]):
+ """
+
+ :param param1: (Default value = [1)
+ :param None:
+ :param "hehe"]:
+
+ """
pass
def func2(param1=(1, None, "hehe")):
+ """
+
+ :param param1: (Default value = (1)
+ :param None:
+ :param "hehe":
+
+ """
pass
def func3(param1={0: 1, "a": None}):
+ """
+
+ :param param1: (Default value = {0: 1)
+ :param "a": None}:
+
+ """
pass
``` | closed | 2017-10-01T13:26:57Z | 2021-03-08T13:50:12Z | https://github.com/dadadel/pyment/issues/46 | [] | dadadel | 1 |
voila-dashboards/voila | jupyter | 812 | Unpin xtl in the tests | In #808, we pinned to `xtl=0.6.23` to fix the tests (`xeus-cling` depends on `xtl`):
https://github.com/voila-dashboards/voila/blob/3209c1f5c2f6588645e6f046f62d893f8ec5d18a/.github/workflows/main.yml#L34
From https://github.com/voila-dashboards/voila/pull/808#issuecomment-764650478
> The xeus-cling stack must be rebuilt so it uses the same standard lib as the last `xtl`
This issue is a reminder to unpin at some point in the future. | closed | 2021-01-21T14:23:57Z | 2021-01-27T07:32:13Z | https://github.com/voila-dashboards/voila/issues/812 | [] | jtpio | 0 |
huggingface/datasets | machine-learning | 6,950 | `Dataset.with_format` behaves inconsistently with documentation | ### Describe the bug
The actual behavior of the interface `Dataset.with_format` is inconsistent with the documentation.
https://huggingface.co/docs/datasets/use_with_pytorch#n-dimensional-arrays
https://huggingface.co/docs/datasets/v2.19.0/en/use_with_tensorflow#n-dimensional-arrays
> If your dataset consists of N-dimensional arrays, you will see that by default they are considered as nested lists.
> In particular, a PyTorch formatted dataset outputs nested lists instead of a single tensor.
> A TensorFlow formatted dataset outputs a RaggedTensor instead of a single tensor.
But I get a single tensor by default, which is inconsistent with the description.
Actually the current behavior seems more reasonable to me. Therefore, the document needs to be modified.
### Steps to reproduce the bug
```python
>>> from datasets import Dataset
>>> data = [[[1, 2],[3, 4]],[[5, 6],[7, 8]]]
>>> ds = Dataset.from_dict({"data": data})
>>> ds = ds.with_format("torch")
>>> ds[0]
{'data': tensor([[1, 2],
[3, 4]])}
>>> ds = ds.with_format("tf")
>>> ds[0]
{'data': <tf.Tensor: shape=(2, 2), dtype=int64, numpy=
array([[1, 2],
[3, 4]])>}
```
### Expected behavior
```python
>>> from datasets import Dataset
>>> data = [[[1, 2],[3, 4]],[[5, 6],[7, 8]]]
>>> ds = Dataset.from_dict({"data": data})
>>> ds = ds.with_format("torch")
>>> ds[0]
{'data': [tensor([1, 2]), tensor([3, 4])]}
>>> ds = ds.with_format("tf")
>>> ds[0]
{'data': <tf.RaggedTensor [[1, 2], [3, 4]]>}
```
### Environment info
datasets==2.19.1
torch==2.1.0
tensorflow==2.13.1 | closed | 2024-06-04T09:18:32Z | 2024-06-25T08:05:49Z | https://github.com/huggingface/datasets/issues/6950 | [
"documentation"
] | iansheng | 2 |
albumentations-team/albumentations | deep-learning | 1,471 | Dynamic dependency on OpenCV is brittle | ## 🐛 Bug
The [dynamic dependency](https://github.com/albumentations-team/albumentations/blob/e3b47b3a127f92541cfeb16abbb44a6f8bf79cc8/setup.py#L10-L16) of `albumentations` on OpenCV means that downstream users who want to install the package at the same time as a pinned version of `opencv-python` may end up with simultaneous installations of `opencv-python` and `opencv-python-headless` which may not be compatible with each other. As a concrete example, today's release of OpenCV 4.8.0.76 broke the build of a product I maintain because the new version is not API-compatible with the project's pinned version of `opencv-python` (which is `4.5.5.64`, released 9 Mar 2022).
## To Reproduce
Steps to reproduce the behavior:
1) Attempt to install `albumentations` and a pinned version of `opencv-python` before `4.6.0.66` in the same `pip` invocation
2) `import cv2`
I've included in this report a reproduction script that illustrates the buggy behavior, example output below.
```
$ PIP_QUIET=1 ./repro.sh # omit PIP_QUIET=1 if you want to see all of pip's output
Installing albumentations 1.3.1 and opencv-python 4.5.5.64 at the same time
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "/tmp/bad_repro_venv/lib/python3.8/site-packages/cv2/__init__.py", line 190, in <module>
bootstrap()
File "/tmp/bad_repro_venv/lib/python3.8/site-packages/cv2/__init__.py", line 184, in bootstrap
if __load_extra_py_code_for_module("cv2", submodule, DEBUG):
File "/tmp/bad_repro_venv/lib/python3.8/site-packages/cv2/__init__.py", line 37, in __load_extra_py_code_for_module
py_module = importlib.import_module(module_name)
File "/usr/lib/python3.8/importlib/__init__.py", line 127, in import_module
return _bootstrap._gcd_import(name[level:], package, level)
File "/tmp/bad_repro_venv/lib/python3.8/site-packages/cv2/typing/__init__.py", line 169, in <module>
LayerId = cv2.dnn.DictValue
AttributeError: module 'cv2.dnn' has no attribute 'DictValue'
OpenCV installation is broken
---
Installing opencv-python 4.5.5.64 before installing albumentations 1.3.1
OpenCV installation is okay
---
Installing opencv-python 4.5.5.64 *and* opencv-python-headless 4.5.5.64 along with albumentations 1.3.1
OpenCV installation is okay
---
```
<details><summary>click for `repro.sh`</summary>
```sh
#!/bin/sh
set -o errexit
rm -fr /tmp/bad_repro_venv /tmp/good_repro_venv
ALBUMENTATIONS_VERSION=${ALBUMENTATIONS_VERSION:-1.3.1}
OPENCV_VERSION=${OPENCV_VERSION:-4.5.5.64}
echo "Installing albumentations ${ALBUMENTATIONS_VERSION} and opencv-python ${OPENCV_VERSION} at the same time"
python3 -m venv /tmp/bad_repro_venv
/tmp/bad_repro_venv/bin/python3 -m pip install "albumentations==${ALBUMENTATIONS_VERSION}" "opencv-python==${OPENCV_VERSION}"
/tmp/bad_repro_venv/bin/python3 -c "import cv2; print('OpenCV installation is okay')" || echo "OpenCV installation is broken"
echo "---"
echo "Installing opencv-python ${OPENCV_VERSION} before installing albumentations ${ALBUMENTATIONS_VERSION}"
python3 -m venv /tmp/good_repro_venv
/tmp/good_repro_venv/bin/python3 -m pip install "opencv-python==${OPENCV_VERSION}"
/tmp/good_repro_venv/bin/python3 -m pip install "albumentations==${ALBUMENTATIONS_VERSION}"
/tmp/good_repro_venv/bin/python3 -c "import cv2; print('OpenCV installation is okay')" || echo "OpenCV installation is broken"
echo "---"
echo "Installing opencv-python ${OPENCV_VERSION} *and* opencv-python-headless ${OPENCV_VERSION} along with albumentations ${ALBUMENTATIONS_VERSION}"
python3 -m venv /tmp/good_repro_venv
/tmp/good_repro_venv/bin/python3 -m pip install "albumentations==${ALBUMENTATIONS_VERSION}" "opencv-python==${OPENCV_VERSION}"
/tmp/good_repro_venv/bin/python3 -c "import cv2; print('OpenCV installation is okay')" || echo "OpenCV installation is broken"
echo "---"
```
</details>
<!-- If you have a code sample, error messages, stack traces, please provide it here as well -->
## Expected behavior
My expectation is that `albumentations` would not require careful handling when installed alongside a pinned version of OpenCV. This dynamic dependency makes it harder for me to maintain a project that includes `albumentations`.
I would much rather have a runtime error from the library telling me to install OpenCV if one of the compatible packages is not installed than the current behavior, which requires a workaround. That is, I suggest that `albumentations` drops OpenCV from its `install_requires` and replaces it with an import-time error if the OpenCV dependency is not satisfied. It may still be appropriate to issue a warning at installation time if OpenCV does not seem to be installed, although such a warning would be a false positive in the case described by this report.
If the maintainers are open to this approach, I would be happy to send a PR.
## Environment
- Albumentations version (e.g., 0.1.8): 1.3.1
- Python version (e.g., 3.7): 3.8.10 (I have also observed this behavior with Python 3.9.14)
- OS (e.g., Linux): Ubuntu 20.04
- How you installed albumentations (`conda`, `pip`, source): `pip`
- Any other relevant information: N/A
## Additional context
Note that the `qudida` library suffers from the same inflexible behavior, so if the relaxed `install_requires` solution I propose were to be adopted, this library would need to be dealt with as well. Since that library is relatively small, has not had a release in 2 years, and uses the permissive MIT License, I would suggest that its contents be folded directly into `albumentations`.
### Related issues
#1100 (possible)
#1139
#1202 (possible)
#1293
https://github.com/aleju/imgaug/issues/737
| closed | 2023-08-09T18:40:47Z | 2024-05-28T22:44:16Z | https://github.com/albumentations-team/albumentations/issues/1471 | [] | jgerityneurala | 7 |
huggingface/transformers | nlp | 36,267 | ci/v4.49-release: tests collection fail with "No module named 'transformers.models.marian.convert_marian_to_pytorch'" on v4.48/v4.49 release branches | Tests collection fails with the following error in https://github.com/huggingface/transformers/tree/v4.49-release branch (and for v4.48-release branch too).
```
python3 -m pytest tests/
===================================================== test session starts =====================================================
platform linux -- Python 3.10.12, pytest-7.4.4, pluggy-1.5.0
rootdir: /home/dvrogozh/git/huggingface/transformers
configfile: pyproject.toml
plugins: anyio-4.8.0, rich-0.2.0, subtests-0.14.1, xdist-3.6.1, asyncio-0.23.8, timeout-2.3.1, hypothesis-6.122.3, reportlog-0.4.0, dash-2.18.2, cov-6.0.0, typeguard-4.3.0
asyncio: mode=strict
collected 84774 items / 1 error
=========================================================== ERRORS ============================================================
________________________________ ERROR collecting tests/models/marian/test_modeling_marian.py _________________________________
ImportError while importing test module '/home/dvrogozh/git/huggingface/transformers/tests/models/marian/test_modeling_marian.py'.
Hint: make sure your test modules/packages have valid Python names.
Traceback:
/usr/lib/python3.10/importlib/__init__.py:126: in import_module
return _bootstrap._gcd_import(name[level:], package, level)
tests/models/marian/test_modeling_marian.py:50: in <module>
from transformers.models.marian.convert_marian_to_pytorch import (
E ModuleNotFoundError: No module named 'transformers.models.marian.convert_marian_to_pytorch'
====================================================== warnings summary =======================================================
src/transformers/optimization.py:640
/home/dvrogozh/git/huggingface/transformers/src/transformers/optimization.py:640: FutureWarning: This implementation of AdamW is deprecated and will be removed in a future version. Use the PyTorch implementation torch.optim.AdamW instead, or set `no_deprecation_warning=True` to disable this warning
warnings.warn(
-- Docs: https://docs.pytest.org/en/stable/how-to/capture-warnings.html
=================================================== short test summary info ===================================================
ERROR tests/models/marian/test_modeling_marian.py
!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!! Interrupted: 1 error during collection !!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!
================================================ 1 warning, 1 error in 17.54s =================================================
```
This happens due to some files are being pruned on release creation commit, see https://github.com/huggingface/transformers/commit/a22a4378d97d06b7a1d9abad6e0086d30fdea199. **Can we resolve this issue to be able to trigger all the tests on release branches?**
CC: @ArthurZucker @ydshieh
| closed | 2025-02-18T23:09:36Z | 2025-02-20T12:22:11Z | https://github.com/huggingface/transformers/issues/36267 | [] | dvrogozh | 1 |
kizniche/Mycodo | automation | 683 | Daemon Stops Running When SPI Enabled | ## Mycodo Issue Report:
- Specific Mycodo Version: 7.6.3
#### Problem Description
I am trying to add a Waveshare High Precision AD/DA (ADS1256) board for adding some analog sensors. After adding it in the output tab, an unmet dependencies warning popped up and I was prompted to to install the missing dependencies. I then ran "raspi-config" to enable SPI. When I then activated that input on the data, the daemon immediately stopped running. I tried rebooting and the daemon was still not running. I ran "raspi-config" again to disable SPI and the daemon started running again. When I repeated the process again, activating the ADS1256 on the data page caused the daemon to stop running.
### Errors
"Error: Error: invalid message type: None"
See Traceback below:
Traceback (most recent call last):
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/flask/app.py", line 2446, in wsgi_app
response = self.full_dispatch_request()
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/flask/app.py", line 1951, in full_dispatch_request
rv = self.handle_user_exception(e)
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/flask/app.py", line 1820, in handle_user_exception
reraise(exc_type, exc_value, tb)
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/flask/_compat.py", line 39, in reraise
raise value
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/flask/app.py", line 1949, in full_dispatch_request
rv = self.dispatch_request()
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/flask/app.py", line 1935, in dispatch_request
return self.view_functions[rule.endpoint](**req.view_args)
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/flask_login/utils.py", line 261, in decorated_view
return func(*args, **kwargs)
File "/home/pi/Mycodo/mycodo/mycodo_flask/routes_page.py", line 911, in page_info
ram_use_daemon = control.ram_use()
File "/home/pi/Mycodo/mycodo/mycodo_client.py", line 150, in ram_use
return self.rpyc_client.root.ram_use()
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/rpyc/core/protocol.py", line 485, in root
self._remote_root = self.sync_request(consts.HANDLE_GETROOT)
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/rpyc/core/protocol.py", line 455, in sync_request
return self.async_request(handler, *args, timeout=timeout).value
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/rpyc/core/async_.py", line 100, in value
self.wait()
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/rpyc/core/async_.py", line 47, in wait
self._conn.serve(self._ttl)
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/rpyc/core/protocol.py", line 384, in serve
self._dispatch(data)
File "/home/pi/Mycodo/env/lib/python3.7/site-packages/rpyc/core/protocol.py", line 358, in _dispatch
raise ValueError("invalid message type: %r" % (msg,))
ValueError: invalid message type: None
### Steps to Reproduce the issue:
1. Connect the WaveShare board (https://www.waveshare.com/High-Precision-AD-DA-Board.htm)
2. Go to the data tab and add an input, selecting the "Texas Instruments: ADS1256: Voltage (WaveShare, Analog to Digital Converter)(UART)" option.
3. Get dependencies flag
4. Click "Install Dependencies" button.
5. Wait for installation to complete.
6. Go back to the "Data" page.
7. Try to activate input source.
8. Get an error and realize that SPI is not enabled.
9. Run "raspi-config" and enable SPI.
10. Notice the daemon is not running.
11. Reboot the Pi.
12. See
### Additional Notes
TBD | closed | 2019-09-03T21:24:53Z | 2019-09-11T21:31:59Z | https://github.com/kizniche/Mycodo/issues/683 | [] | smatthews95 | 9 |
tqdm/tqdm | jupyter | 1,415 | resizing spams the terminal??? | Category: visual output "bug" (not sure i'd call it a bug, but it's definitely different behavior than i would expect).
I have read the known issues
I have searched github open issues
I am using Python 3.10.7, tqdm 4.64.0, on ubuntu 22.10, with tqdm installed via apt-get (python3-tqdm version 4.64.0-2)
When i resize a window (with "dynamic_ncols" defaulting to False), i would expect to simply be left with an inappropriately-sized progress-bar line.
instead, it repeatedly prints to screen (printing new lines, not overwriting the previous line like i would expect), and blows up the terminal output.
functionally, this effectively prevents you from resizing a terminal window, unless i had the foresight to use the extra argument (and now it's too late because my thing takes eons to run, ha!)
here are a couple screenshots; this comes from unmaximizing my full-screen terminal window, then remaximizing. https://imgur.com/a/HBZON55 | open | 2023-01-11T17:39:44Z | 2023-01-11T17:40:21Z | https://github.com/tqdm/tqdm/issues/1415 | [] | tpchuckles | 0 |
yihong0618/running_page | data-visualization | 331 | riding data | If I want to download riding data from Keep, which parameters should be modified? | closed | 2022-10-27T02:42:39Z | 2022-10-31T05:59:23Z | https://github.com/yihong0618/running_page/issues/331 | [] | Dabao55 | 1 |
man-group/arctic | pandas | 211 | inconsistent behavior between update and append in chunkstore | ```
df = pd.DataFrame(data={'data': [1]}, index=pd.MultiIndex.from_tuples([(dt(2016,1,1), 1)], names=['date', 'id']))
df2 = pd.DataFrame(data={'data': [2]}, index=pd.MultiIndex.from_tuples([(dt(2016,1,2), 2)], names=['date', 'id']))
l.write('test', df, 'D')
l.append('test', df2)
---------------------------------------------------------------------------
Exception Traceback (most recent call last)
<ipython-input-28-790fa6766bb4> in <module>()
----> 1 l.append('test', df2)
/users/is/ahlpypi/egg_cache/a/arctic-1.25.0-py2.7-linux-x86_64.egg/arctic/chunkstore/chunkstore.pyc in append(self, symbol, item)
278
279 if str(dtype) != sym['dtype']:
--> 280 raise Exception("Dtype mismatch - cannot append")
281
282 data = item.tostring()
Exception: Dtype mismatch - cannot append
l.update('test', df2)
l.read('test')
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
<ipython-input-30-8bc9f991dd86> in <module>()
----> 1 l.read('test')
/users/is/ahlpypi/egg_cache/a/arctic-1.25.0-py2.7-linux-x86_64.egg/arctic/chunkstore/chunkstore.pyc in read(self, symbol, chunk_range)
133
134 dtype = PandasSerializer()._dtype(sym['dtype'], sym.get('dtype_metadata', {}))
--> 135 records = np.fromstring(data, dtype=dtype).reshape(sym.get('shape', (-1)))
136
137 data = deserialize(records, sym['type'])
ValueError: string size must be a multiple of element size
```
Update and Append should behave similarly. Also, should probably allow addition of new column within reason, and rely on pandas' ability to combine separate dataframes with missing columns:
```
pd.concat([df, df2])
data price
date id
2016-01-01 1 1 NaN
2016-01-02 2 2 50
```
| closed | 2016-09-07T18:22:38Z | 2016-09-07T18:41:24Z | https://github.com/man-group/arctic/issues/211 | [
"bug"
] | bmoscon | 1 |
ymcui/Chinese-LLaMA-Alpaca | nlp | 769 | plus和pro模型的区别在什么地方呢? | ### 提交前必须检查以下项目
- [X] 请确保使用的是仓库最新代码(git pull),一些问题已被解决和修复。
- [X] 由于相关依赖频繁更新,请确保按照[Wiki](https://github.com/ymcui/Chinese-LLaMA-Alpaca/wiki)中的相关步骤执行
- [X] 我已阅读[FAQ章节](https://github.com/ymcui/Chinese-LLaMA-Alpaca/wiki/常见问题)并且已在Issue中对问题进行了搜索,没有找到相似问题和解决方案
- [X] 第三方插件问题:例如[llama.cpp](https://github.com/ggerganov/llama.cpp)、[text-generation-webui](https://github.com/oobabooga/text-generation-webui)、[LlamaChat](https://github.com/alexrozanski/LlamaChat)等,同时建议到对应的项目中查找解决方案
- [X] 模型正确性检查:务必检查模型的[SHA256.md](https://github.com/ymcui/Chinese-LLaMA-Alpaca/blob/main/SHA256.md),模型不对的情况下无法保证效果和正常运行
### 问题类型
效果问题
### 基础模型
None
### 操作系统
None
### 详细描述问题
根据项目wiki描述,plus模型的回复相对较短,pro模型的回复相对较长。
请问下是通过SFT数据的input/output长度调整的吗?还是有其他实现思路?后续如果要在现有模型的基础上,继续进行SFT的话,是推荐plus模型,还是pro模型呢?二者的区别大不大?
感谢🙏
| closed | 2023-07-19T16:46:20Z | 2023-08-14T22:02:30Z | https://github.com/ymcui/Chinese-LLaMA-Alpaca/issues/769 | [
"stale"
] | minlik | 9 |
xonsh/xonsh | data-science | 5,798 | pytest: The (fspath: py.path.local) argument to XshFile is deprecated. Please use the (path: pathlib.Path) argument instead. | ## Current Behavior
Currently if I try to add tests for functions in a xsh script (foo.xsh), say this script:
```xsh
def bar():
return 42
```
and I then use this test file (test_foo.xsh):
```xsh
from foo import bar
def test_foo():
assert 42 == bar()
```
Then if I run the test using pytest like so `pytest test_foo.xsh` then I get this output
```console
============================================================ test session starts ============================================================
platform linux -- Python 3.12.3, pytest-7.4.4, pluggy-1.4.0
rootdir: /tmp/foo
plugins: xonsh-0.14.4
collected 1 item
test_foo.xsh . [100%]
============================================================= warnings summary ==============================================================
../../usr/lib/python3/dist-packages/xonsh/pytest/plugin.py:61
/usr/lib/python3/dist-packages/xonsh/pytest/plugin.py:61: PytestRemovedIn8Warning: The (fspath: py.path.local) argument to XshFile is deprecated. Please use the (path: pathlib.Path) argument instead.
See https://docs.pytest.org/en/latest/deprecations.html#fspath-argument-for-node-constructors-replaced-with-pathlib-path
return XshFile.from_parent(parent, fspath=path)
-- Docs: https://docs.pytest.org/en/stable/how-to/capture-warnings.html
======================================================= 1 passed, 1 warning in 0.01s ========================================================
```
So it all works and the test pass, but I get the warning: "PytestRemovedIn8Warning: The (fspath: py.path.local) argument to XshFile is deprecated. Please use the (path: pathlib.Path) argument instead."
## Expected Behavior
I would expect no warning, i.e. this output:
```console
============================================================ test session starts ============================================================
platform linux -- Python 3.12.3, pytest-7.4.4, pluggy-1.4.0
rootdir: /tmp/foo
plugins: xonsh-0.14.4
collected 1 item
test_foo.xsh . [100%]
============================================================= 1 passed in 0.01s =============================================================
```
| closed | 2025-02-24T12:42:29Z | 2025-02-25T11:56:39Z | https://github.com/xonsh/xonsh/issues/5798 | [
"to-close-in-the-future",
"pytest"
] | svenskmand | 4 |
microsoft/nni | deep-learning | 5,600 | How to specify the target of distilling when there are multiple outputs? | **Describe the issue**:
I am using the transcript of [pruning BERT on MNLI](https://nni.readthedocs.io/en/v3.0rc1/tutorials/new_pruning_bert_glue.html). I find a problem of distillation when I change the output format of the encoder, which is as follows.
``` python
layer_outputs = layer_module(hidden_states, ...) # original BERT encoder
layer_outputs, second_output = layer_module(hidden_states, ...) # modified BERT encoder
```
I changed the configuration of the distiller
``` python
def dynamic_distiller(student_model: BertForSequenceClassification, teacher_model: BertForSequenceClassification,
student_trainer: Trainer):
layer_num = len(student_model.bert.encoder.layer)
config_list = [{
'op_names': [f'bert.encoder.layer.{i}'],
'link': [f'bert.encoder.layer.{j}' for j in range(i, layer_num)],
'lambda': 0.9,
'apply_method': 'mse',
'target_names': ['_output_0'] # this line is new added to specify which output is used to distil.
} for i in range(layer_num)]
config_list.append({
'op_names': ['classifier'],
'link': ['classifier'],
'lambda': 0.9,
'apply_method': 'kl',
})
evaluator = TransformersEvaluator(student_trainer)
def teacher_predict(batch, teacher_model):
return teacher_model(**batch)
return DynamicLayerwiseDistiller(student_model, config_list, evaluator, teacher_model, teacher_predict, origin_loss_lambda=0.1)
```
But the errors I put in the Log message occurs. Could you help me with this problem?
**Environment**:
- NNI version: 999.dev0 (installed from the source, I guess it is the newest).
**Log message**:
```
Traceback (most recent call last):
File "/home/work/project/prune_bert_glue/pruning_bert_glue.py", line 178, in <module>
finetuned_model=post_distillation(task_name, finetuned_model, teacher_model, output_dir=stage_softmax_dir)
File "/home/work/project/prune_bert_glue/prune_modules.py", line 107, in post_distillation
dynamic_distillation(task_name, model, copy.deepcopy(teacher_model), output_dir, None, 3)
File "/home/work/project/prune_bert_glue/distiller.py", line 52, in dynamic_distillation
distiller.compress(max_steps, max_epochs)
File "/home/work/project/nni/nni/contrib/compression/base/compressor.py", line 190, in compress
self._single_compress(max_steps, max_epochs)
File "/home/work/project/nni/nni/contrib/compression/distillation/basic_distiller.py", line 144, in _single_compress
self._fusion_compress(max_steps, max_epochs)
File "/home/work/project/nni/nni/contrib/compression/base/compressor.py", line 183, in _fusion_compress
self.evaluator.train(max_steps, max_epochs)
File "/home/work/project/nni/nni/contrib/compression/utils/evaluator.py", line 1084, in train
self.trainer.train()
File "/home/work/anaconda3/envs/nni/lib/python3.10/site-packages/transformers/trainer.py", line 1664, in train
return inner_training_loop(
File "/home/work/anaconda3/envs/nni/lib/python3.10/site-packages/transformers/trainer.py", line 1940, in _inner_training_loop
tr_loss_step = self.training_step(model, inputs)
File "/home/work/anaconda3/envs/nni/lib/python3.10/site-packages/transformers/trainer.py", line 2735, in training_step
loss = self.compute_loss(model, inputs)
File "/home/work/project/nni/nni/contrib/compression/utils/evaluator.py", line 1044, in patched_compute_loss
result = old_compute_loss(model, inputs, return_outputs)
File "/home/work/anaconda3/envs/nni/lib/python3.10/site-packages/transformers/trainer.py", line 2767, in compute_loss
outputs = model(**inputs)
File "/home/work/anaconda3/envs/nni/lib/python3.10/site-packages/torch/nn/modules/module.py", line 1130, in _call_impl
return forward_call(*input, **kwargs)
File "/home/work/project/prune_bert_glue/models/modeling_bert.py", line 1592, in forward
outputs = self.bert(
File "/home/work/anaconda3/envs/nni/lib/python3.10/site-packages/torch/nn/modules/module.py", line 1130, in _call_impl
return forward_call(*input, **kwargs)
File "/home/work/project/prune_bert_glue/models/modeling_bert.py", line 1050, in forward
encoder_outputs = self.encoder(
File "/home/work/anaconda3/envs/nni/lib/python3.10/site-packages/torch/nn/modules/module.py", line 1130, in _call_impl
return forward_call(*input, **kwargs)
File "/home/work/project/prune_bert_glue/models/modeling_bert.py", line 627, in forward
layer_outputs, sparse_mask = layer_module(
File "/home/work/anaconda3/envs/nni/lib/python3.10/site-packages/torch/nn/modules/module.py", line 1130, in _call_impl
return forward_call(*input, **kwargs)
File "/home/work/project/nni/nni/contrib/compression/base/wrapper.py", line 407, in forward
outputs = self.patch_outputs(outputs)
File "/home/work/project/nni/nni/contrib/compression/base/wrapper.py", line 369, in patch_outputs
new_outputs.append(self.patch_helper(target_name, target))
File "/home/work/project/nni/nni/contrib/compression/base/wrapper.py", line 331, in patch_helper
target = self._distil_observe_helper(target, self.distillation_target_spaces[target_name])
File "/home/work/project/nni/nni/contrib/compression/base/wrapper.py", line 303, in _distil_observe_helper
target_space.hidden_state = target
File "/home/work/project/nni/nni/contrib/compression/base/target_space.py", line 387, in hidden_state
raise TypeError('Only support saving tensor as distillation hidden_state.')
TypeError: Only support saving tensor as distillation hidden_state.
``` | closed | 2023-06-07T09:02:02Z | 2023-06-09T12:40:13Z | https://github.com/microsoft/nni/issues/5600 | [] | hobbitlzy | 3 |
pytorch/vision | machine-learning | 8,625 | ImageReadMode should support strings | It's pretty inconvenient to have to import ImageReadMode just to ask `decode_*` for an RGB image. We should just allow strings as well.
Also, `RGBA` should be a valid option (like in PIL). `RGB_ALPHA` is... long. | closed | 2024-09-03T14:46:03Z | 2024-09-04T10:38:52Z | https://github.com/pytorch/vision/issues/8625 | [] | NicolasHug | 0 |
sigmavirus24/github3.py | rest-api | 497 | Add issue import beta endpoint(s) | https://gist.github.com/jonmagic/5282384165e0f86ef105
| closed | 2015-12-08T21:33:25Z | 2018-07-20T14:11:49Z | https://github.com/sigmavirus24/github3.py/issues/497 | [
"in progress"
] | sigmavirus24 | 9 |
vanna-ai/vanna | data-visualization | 702 | Error: The server returned an error. See the server logs for more details. If you are running in Colab, this function is probably not supported. Please try running in a local environment. | **Describe the bug**
I have vanna running in docker, but I'm getting the following error when submitting my training
> Error: The server returned an error. See the server logs for more details. If you are running in Colab, this function is probably not
> supported. Please try running in a local environment.

**To Reproduce**
Core content of the app.py file
```python
index_html_path = os.path.abspath('./static/index.html')
logo_path = os.path.abspath('./static/assets/vanna.svg')
assets_folder = os.path.abspath('./static/assets')
app = VannaFlaskApp(
vn,
auth=SimplePassword(users=accounts.users, tokens=accounts.tokens),
allow_llm_to_see_data=True,
debug=True,
logo=logo_path,
index_html_path=index_html_path,
assets_folder=assets_folder
)
```
Full content of the index.html file
```html
<!doctype html>
<html lang="en">
<head>
<meta charset="UTF-8" />
<link rel="icon" type="image/svg+xml" href="/assets/vanna.svg" />
<meta name="viewport" content="width=device-width, initial-scale=1.0" />
<link href="https://fonts.googleapis.com/css2?family=Roboto+Slab:wght@350&display=swap" rel="stylesheet">
<script src="https://cdn.plot.ly/plotly-latest.min.js" type="text/javascript"></script>
<title>Vanna.AI</title>
<script type="module" crossorigin src="/assets/index-2dc047a4.js"></script>
<link rel="stylesheet" href="/assets/index-a3ae634d.css">
</head>
<body class="bg-white dark:bg-slate-900">
<div id="app"></div>
</body>
</html>
```
**Error logs**
```shell
2024-11-14T05:43:04.707037285Z Traceback (most recent call last):
2024-11-14T05:43:04.707040046Z File "/usr/local/lib/python3.10/site-packages/requests/adapters.py", line 667, in send
2024-11-14T05:43:04.707042991Z resp = conn.urlopen(
2024-11-14T05:43:04.707045775Z File "/usr/local/lib/python3.10/site-packages/urllib3/connectionpool.py", line 843, in urlopen
2024-11-14T05:43:04.707048730Z retries = retries.increment(
2024-11-14T05:43:04.707051493Z File "/usr/local/lib/python3.10/site-packages/urllib3/util/retry.py", line 519, in increment
2024-11-14T05:43:04.707055770Z raise MaxRetryError(_pool, url, reason) from reason # type: ignore[arg-type]
2024-11-14T05:43:04.707060043Z urllib3.exceptions.MaxRetryError: HTTPSConnectionPool(host='vanna.ai', port=443): Max retries exceeded with url: /img/vanna.svg (Caused by ConnectTimeoutError(<urllib3.connection.HTTPSConnection object at 0x7f10cb9a0ee0>, 'Connection to vanna.ai timed out. (connect timeout=None)'))
2024-11-14T05:43:04.707063493Z
2024-11-14T05:43:04.707066201Z During handling of the above exception, another exception occurred:
2024-11-14T05:43:04.707072641Z
2024-11-14T05:43:04.707075489Z Traceback (most recent call last):
2024-11-14T05:43:04.707078291Z File "/usr/local/lib/python3.10/site-packages/flask/app.py", line 1463, in wsgi_app
2024-11-14T05:43:04.707081175Z response = self.full_dispatch_request()
2024-11-14T05:43:04.707083977Z File "/usr/local/lib/python3.10/site-packages/flask/app.py", line 872, in full_dispatch_request
2024-11-14T05:43:04.707086930Z rv = self.handle_user_exception(e)
2024-11-14T05:43:04.707089720Z File "/usr/local/lib/python3.10/site-packages/flask/app.py", line 870, in full_dispatch_request
2024-11-14T05:43:04.707092703Z rv = self.dispatch_request()
2024-11-14T05:43:04.707095481Z File "/usr/local/lib/python3.10/site-packages/flask/app.py", line 855, in dispatch_request
2024-11-14T05:43:04.707098407Z return self.ensure_sync(self.view_functions[rule.endpoint])(**view_args) # type: ignore[no-any-return]
2024-11-14T05:43:04.707101400Z File "/usr/local/lib/python3.10/site-packages/vanna/flask/__init__.py", line 1276, in proxy_vanna_svg
2024-11-14T05:43:04.707104372Z response = requests.get(remote_url, stream=True)
2024-11-14T05:43:04.707107183Z File "/usr/local/lib/python3.10/site-packages/requests/api.py", line 73, in get
2024-11-14T05:43:04.707110071Z return request("get", url, params=params, **kwargs)
2024-11-14T05:43:04.707113028Z File "/usr/local/lib/python3.10/site-packages/requests/api.py", line 59, in request
2024-11-14T05:43:04.707115928Z return session.request(method=method, url=url, **kwargs)
2024-11-14T05:43:04.707118773Z File "/usr/local/lib/python3.10/site-packages/requests/sessions.py", line 589, in request
2024-11-14T05:43:04.707121739Z resp = self.send(prep, **send_kwargs)
2024-11-14T05:43:04.707124543Z File "/usr/local/lib/python3.10/site-packages/requests/sessions.py", line 703, in send
2024-11-14T05:43:04.707127499Z r = adapter.send(request, **kwargs)
2024-11-14T05:43:04.707130250Z File "/usr/local/lib/python3.10/site-packages/requests/adapters.py", line 688, in send
2024-11-14T05:43:04.707133616Z raise ConnectTimeout(e, request=request)
2024-11-14T05:43:04.707136878Z requests.exceptions.ConnectTimeout: HTTPSConnectionPool(host='vanna.ai', port=443): Max retries exceeded with url: /img/vanna.svg (Caused by ConnectTimeoutError(<urllib3.connection.HTTPSConnection object at 0x7f10cb9a0ee0>, 'Connection to vanna.ai timed out. (connect timeout=None)'))
2024-11-14T05:43:09.954900441Z [2024-11-14 13:43:09,954] ERROR in app: Exception on /vanna.svg [GET]
2024-11-14T05:43:09.954932822Z Traceback (most recent call last):
2024-11-14T05:43:09.954937482Z File "/usr/local/lib/python3.10/site-packages/urllib3/connection.py", line 199, in _new_conn
2024-11-14T05:43:09.954942589Z sock = connection.create_connection(
2024-11-14T05:43:09.954946462Z File "/usr/local/lib/python3.10/site-packages/urllib3/util/connection.py", line 85, in create_connection
2024-11-14T05:43:09.954959922Z raise err
2024-11-14T05:43:09.954963003Z File "/usr/local/lib/python3.10/site-packages/urllib3/util/connection.py", line 73, in create_connection
2024-11-14T05:43:09.954966439Z sock.connect(sa)
2024-11-14T05:43:09.954969394Z TimeoutError: [Errno 110] Connection timed out
2024-11-14T05:43:09.954972418Z
2024-11-14T05:43:09.954975202Z The above exception was the direct cause of the following exception:
2024-11-14T05:43:09.954978288Z
2024-11-14T05:43:09.954982785Z Traceback (most recent call last):
2024-11-14T05:43:09.954985905Z File "/usr/local/lib/python3.10/site-packages/urllib3/connectionpool.py", line 789, in urlopen
2024-11-14T05:43:09.954988847Z response = self._make_request(
2024-11-14T05:43:09.954991620Z File "/usr/local/lib/python3.10/site-packages/urllib3/connectionpool.py", line 490, in _make_request
2024-11-14T05:43:09.954994632Z raise new_e
2024-11-14T05:43:09.954997546Z File "/usr/local/lib/python3.10/site-packages/urllib3/connectionpool.py", line 466, in _make_request
2024-11-14T05:43:09.955000511Z self._validate_conn(conn)
2024-11-14T05:43:09.955003326Z File "/usr/local/lib/python3.10/site-packages/urllib3/connectionpool.py", line 1095, in _validate_conn
2024-11-14T05:43:09.955006317Z conn.connect()
2024-11-14T05:43:09.955009180Z File "/usr/local/lib/python3.10/site-packages/urllib3/connection.py", line 693, in connect
2024-11-14T05:43:09.955012152Z self.sock = sock = self._new_conn()
2024-11-14T05:43:09.955014956Z File "/usr/local/lib/python3.10/site-packages/urllib3/connection.py", line 208, in _new_conn
2024-11-14T05:43:09.955017975Z raise ConnectTimeoutError(
2024-11-14T05:43:09.955020902Z urllib3.exceptions.ConnectTimeoutError: (<urllib3.connection.HTTPSConnection object at 0x7f10c8bc6260>, 'Connection to vanna.ai timed out. (connect timeout=None)')
2024-11-14T05:43:09.955024500Z
2024-11-14T05:43:09.955027325Z The above exception was the direct cause of the following exception:
2024-11-14T05:43:09.955030204Z
2024-11-14T05:43:09.955032918Z Traceback (most recent call last):
2024-11-14T05:43:09.955035696Z File "/usr/local/lib/python3.10/site-packages/requests/adapters.py", line 667, in send
2024-11-14T05:43:09.955038612Z resp = conn.urlopen(
2024-11-14T05:43:09.955041416Z File "/usr/local/lib/python3.10/site-packages/urllib3/connectionpool.py", line 843, in urlopen
2024-11-14T05:43:09.955044358Z retries = retries.increment(
2024-11-14T05:43:09.955047177Z File "/usr/local/lib/python3.10/site-packages/urllib3/util/retry.py", line 519, in increment
2024-11-14T05:43:09.955051971Z raise MaxRetryError(_pool, url, reason) from reason # type: ignore[arg-type]
2024-11-14T05:43:09.955056251Z urllib3.exceptions.MaxRetryError: HTTPSConnectionPool(host='vanna.ai', port=443): Max retries exceeded with url: /img/vanna.svg (Caused by ConnectTimeoutError(<urllib3.connection.HTTPSConnection object at 0x7f10c8bc6260>, 'Connection to vanna.ai timed out. (connect timeout=None)'))
2024-11-14T05:43:09.955062996Z
2024-11-14T05:43:09.955065825Z During handling of the above exception, another exception occurred:
2024-11-14T05:43:09.955068775Z
2024-11-14T05:43:09.955071509Z Traceback (most recent call last):
2024-11-14T05:43:09.955074330Z File "/usr/local/lib/python3.10/site-packages/flask/app.py", line 1463, in wsgi_app
2024-11-14T05:43:09.955077279Z response = self.full_dispatch_request()
2024-11-14T05:43:09.955080163Z File "/usr/local/lib/python3.10/site-packages/flask/app.py", line 872, in full_dispatch_request
2024-11-14T05:43:09.955083108Z rv = self.handle_user_exception(e)
2024-11-14T05:43:09.955085947Z File "/usr/local/lib/python3.10/site-packages/flask/app.py", line 870, in full_dispatch_request
2024-11-14T05:43:09.955089072Z rv = self.dispatch_request()
2024-11-14T05:43:09.955091860Z File "/usr/local/lib/python3.10/site-packages/flask/app.py", line 855, in dispatch_request
2024-11-14T05:43:09.955094981Z return self.ensure_sync(self.view_functions[rule.endpoint])(**view_args) # type: ignore[no-any-return]
2024-11-14T05:43:09.955098039Z File "/usr/local/lib/python3.10/site-packages/vanna/flask/__init__.py", line 1276, in proxy_vanna_svg
2024-11-14T05:43:09.955101013Z response = requests.get(remote_url, stream=True)
2024-11-14T05:43:09.955103859Z File "/usr/local/lib/python3.10/site-packages/requests/api.py", line 73, in get
2024-11-14T05:43:09.955106822Z return request("get", url, params=params, **kwargs)
2024-11-14T05:43:09.955109829Z File "/usr/local/lib/python3.10/site-packages/requests/api.py", line 59, in request
2024-11-14T05:43:09.955112766Z return session.request(method=method, url=url, **kwargs)
2024-11-14T05:43:09.955115639Z File "/usr/local/lib/python3.10/site-packages/requests/sessions.py", line 589, in request
2024-11-14T05:43:09.955119003Z resp = self.send(prep, **send_kwargs)
2024-11-14T05:43:09.955121995Z File "/usr/local/lib/python3.10/site-packages/requests/sessions.py", line 703, in send
2024-11-14T05:43:09.955124886Z r = adapter.send(request, **kwargs)
2024-11-14T05:43:09.955127674Z File "/usr/local/lib/python3.10/site-packages/requests/adapters.py", line 688, in send
2024-11-14T05:43:09.955130571Z raise ConnectTimeout(e, request=request)
2024-11-14T05:43:09.955134066Z requests.exceptions.ConnectTimeout: HTTPSConnectionPool(host='vanna.ai', port=443): Max retries exceeded with url: /img/vanna.svg (Caused by ConnectTimeoutError(<urllib3.connection.HTTPSConnection object at 0x7f10c8bc6260>, 'Connection to vanna.ai timed out. (connect timeout=None)'))
```
**Desktop (please complete the following information where):**
- OS: [e.g. Ubuntu]
- Version: [e.g. 20.04]
- Python: [3.10]
- Vanna: [0.7.5]
**Additional context**
I tested it and the result seems to be that all routes return the content of index.html by this method. `vanna/flask/__init__.py`
```python
@self.flask_app.route("/index", defaults={"path": ""})
@self.flask_app.route("/<path:path>")
def hello(path: str):
if self.index_html_path:
directory = os.path.dirname(self.index_html_path)
filename = os.path.basename(self.index_html_path)
return send_from_directory(directory=directory, path=filename)
return html_content
```
| open | 2024-11-14T06:34:36Z | 2025-01-21T13:18:45Z | https://github.com/vanna-ai/vanna/issues/702 | [
"bug"
] | SharkSyl | 2 |
newpanjing/simpleui | django | 331 | 多标签标题更新错乱 | **bug描述**
* *Bug description * *
* 打开多个model标签(`首页`、`A`、`B`)后,在`A`标签刷新时马上切换到`B`标签,之后`A`标签刷新完成更新标签标题时,新标题名称被更新到`B`标签,`A`标签的标题未变;
* 打开多个model标签(`首页`、`A`、`B`)后,`A`标签页面点击任意一个条目进入详情页面,加载完成后,切换到`首页`标签,快捷操作中`A`标签对应model的文本标题变更为`A`标签的标题。
**环境**
** environment**
1.Operating System:Centos 6
2.Python Version:3.7.4
3.Django Version:2.2.17
4.SimpleUI Version:2021.1.1
| closed | 2020-12-31T03:14:22Z | 2021-01-26T07:28:35Z | https://github.com/newpanjing/simpleui/issues/331 | [
"bug"
] | eshxcmhk | 1 |
FactoryBoy/factory_boy | django | 340 | DESCRIPTION.rst and METADATA still refer to fake-factory | `factory_boy` no longer relies on `fake-factory`, but in `METADATA` and `DESCRIPTION.rst` there's still:
> For this, factory_boy relies on the excellent `fake-factory <https://pypi.python.org/pypi/fake-factory>`
Super minor and I'm happy to PR it, I just haven't set up dev for `factory_boy` before | closed | 2017-01-09T15:43:09Z | 2017-01-10T20:30:00Z | https://github.com/FactoryBoy/factory_boy/issues/340 | [] | jisantuc | 2 |
sczhou/CodeFormer | pytorch | 43 | metrics | In your article, you wrote:
For the evaluation on real-world datasets without ground truth, we employ the widely-used non-reference perceptual metrics: FID and NIQE.
Is FID a non-reference indicator?
I have some trouble understanding this sentence. Thanks! | open | 2022-10-05T08:20:22Z | 2022-11-01T01:22:04Z | https://github.com/sczhou/CodeFormer/issues/43 | [] | 123456789-qwer | 4 |
strawberry-graphql/strawberry | asyncio | 2,845 | relay: NodeID does not support auto | <!-- Provide a general summary of the bug in the title above. -->
Something like relay.NodeID[auto] is not possible. The evaled type is there auto not Annotated | open | 2023-06-12T22:49:54Z | 2025-03-20T15:56:13Z | https://github.com/strawberry-graphql/strawberry/issues/2845 | [
"bug"
] | devkral | 0 |
coleifer/sqlite-web | flask | 135 | Support multiple DB's | Hi there
I would like to use sqlite_web to display the gui for multiple db's, which almost works since I can use multiple calls to initialize_app. However, the globals in sqlite_web.py are a deal breaker. Would you accept a PR that simply turns the globals to a dict with the file_name as key or something the like? | closed | 2023-10-20T12:27:18Z | 2025-02-23T16:04:01Z | https://github.com/coleifer/sqlite-web/issues/135 | [] | aersam | 5 |
apache/airflow | machine-learning | 47,413 | Scheduler HA mode, DagFileProcessor Race Condition | ### Apache Airflow version
Other Airflow 2 version (please specify below)
### If "Other Airflow 2 version" selected, which one?
2.10.1
### What happened?
We use dynamic dag generation to generate dags in our Airflow environment. We have one base dag definition file, we will call `big_dag.py`, generating >1500 dags. Recently, after the introduction of a handful more dags generated from `big_dag.py`, all the `big_dag.py` generated dags have disappeared from UI and reappear randomly in a loop.
We noticed that if we restart our env a couple times, we could randomly achieve stability. We started to believe some timing issue was at play.
### What you think should happen instead?
Goal State: Dags that generate >1500 dags should not cause any disruptions to environment, given appropriate timeouts.
After checking the dag_process_manager log stream we noticed a prevalence of this error:
`psycopg2.errors.UniqueViolation) duplicate key value violates unique constraint "serialized_dag_pkey" DETAIL: Key (dag_id)=(<dag_name>)`
I believe the issue is on this line of the `write_dag` function of the `SerializedDagModel`:
**This code is from the main branch, I believe the issue is still present in main**
https://github.com/apache/airflow/blob/7bfe283cf4fa28453c857e659f4c1d5917f9e11c/airflow/models/serialized_dag.py#L197
The check for if a serialized dag should be updated or not is NOT ATOMIC, which leads to the issue where more than 1 scheduler runs into a race condition while trying to update serialization.
I believe a "check-then-update" atomic action should be used here through a mechanism like the row level `SELECT ... FOR UPDATE`.
### How to reproduce
You can reproduce this by having an environment with multiple schedulers/standalone_dag_file_processors and dag files that dynamically generate > 1500 dags. Time for a full processing of a >1500 dag file should be ~200 seconds (make sure timeout accommodates this).
To increase the likelihood the duplicate serialized pkey issue happens, reduce min_file_process_interval to like 30 seconds.
### Operating System
Amazon Linux 2023
### Versions of Apache Airflow Providers
_No response_
### Deployment
Amazon (AWS) MWAA
### Deployment details
2.10.1
2 Schedulers
xL Environment Size:

min_file_process_interval= 600
standalone_dag_processor = True (we believe MWAA creates one per scheduler)
dag_file_processor_timeout = 900
dagbag_import_timeout = 900
### Anything else?
I am not sure why the timing works out when dag definitio files are generating <<1500 dags, but could just be the speed of the environment is finishing all work before a race condition can occur.
### Are you willing to submit PR?
- [x] Yes I am willing to submit a PR!
### Code of Conduct
- [x] I agree to follow this project's [Code of Conduct](https://github.com/apache/airflow/blob/main/CODE_OF_CONDUCT.md)
| open | 2025-03-05T19:43:20Z | 2025-03-11T16:09:10Z | https://github.com/apache/airflow/issues/47413 | [
"kind:bug",
"area:Scheduler",
"area:MetaDB",
"area:core",
"needs-triage"
] | robertchinezon | 4 |
lyhue1991/eat_tensorflow2_in_30_days | tensorflow | 46 | tf serving预测会有错误 | tf serving预测会有错误帮忙看下
{ "error": "Malformed request: POST /v1/models/linear_model" }{ "error": "In[0] is not a matrix. Instead it has shape [3]\n\t [[{{node model/outputs/BiasAdd}}]]" }% | closed | 2020-05-29T07:01:30Z | 2020-05-29T07:03:51Z | https://github.com/lyhue1991/eat_tensorflow2_in_30_days/issues/46 | [] | binzhouchn | 0 |
huggingface/transformers | python | 36,227 | <spam> | spam | closed | 2025-02-17T09:39:21Z | 2025-02-17T16:17:43Z | https://github.com/huggingface/transformers/issues/36227 | [
"bug"
] | j3801996 | 0 |
plotly/dash-table | plotly | 294 | Defining copy and paste behaviour on editable and non-editable columns | Related to https://github.com/plotly/dash-table/pull/293#issuecomment-445967446 creating a test for checking behaviour of copying and pasting onto a table with both editable and non-editable cells:
what should the behaviour be when copying & pasting 2 cols onto another table with an editable col and non-editable col? | open | 2018-12-10T21:18:00Z | 2019-07-06T12:24:49Z | https://github.com/plotly/dash-table/issues/294 | [
"dash-type-enhancement",
"Status: Discussion Needed"
] | cldougl | 0 |
horovod/horovod | tensorflow | 2,983 | can not run "horovodrun -np 4 -H localhost:4 python keras_mnist_advanced.py" inside horovod container | System: 8 A100 PCIe NVIDIA GPU
CUDA: 11.2
Driver: 460.27.04
OS: Ubuntu 18.04.4 LTS
Run following cmd as horovod instructions:
```
docker pull horovod/horovod
docker run -it imageID
#example/keras horovodrun -np 4 -H localhost:4 python keras_mnist_advanced.py
```
getting following Errors, please help!
```
2021-06-15 00:52:36.479715: I tensorflow/stream_executor/platform/default/dso_loader.cc:53] Successfully opened dynamic library libcudart.so.11.0
[1,0]<stderr>:2021-06-15 00:52:38.200811: I tensorflow/stream_executor/platform/default/dso_loader.cc:53] Successfully opened dynamic library libcudart.so.11.0
[1,1]<stderr>:2021-06-15 00:52:38.223588: I tensorflow/stream_executor/platform/default/dso_loader.cc:53] Successfully opened dynamic library libcudart.so.11.0
[1,2]<stderr>:2021-06-15 00:52:38.223589: I tensorflow/stream_executor/platform/default/dso_loader.cc:53] Successfully opened dynamic library libcudart.so.11.0
[1,3]<stderr>:2021-06-15 00:52:38.223610: I tensorflow/stream_executor/platform/default/dso_loader.cc:53] Successfully opened dynamic library libcudart.so.11.0
[1,0]<stderr>:Traceback (most recent call last):
[1,0]<stderr>: File "./keras/keras_mnist_advanced.py", line 3, in <module>
[1,0]<stderr>: import keras
[1,0]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/__init__.py", line 20, in <module>
[1,0]<stderr>: from . import initializers
[1,0]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/initializers/__init__.py", line 124, in <module>
[1,0]<stderr>: populate_deserializable_objects()
[1,0]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/initializers/__init__.py", line 82, in populate_deserializable_objects
[1,0]<stderr>: generic_utils.populate_dict_with_module_objects(
[1,0]<stderr>:AttributeError: module 'keras.utils.generic_utils' has no attribute 'populate_dict_with_module_objects'
[1,2]<stderr>:Traceback (most recent call last):
[1,2]<stderr>: File "./keras/keras_mnist_advanced.py", line 3, in <module>
[1,2]<stderr>: import keras
[1,2]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/__init__.py", line 20, in <module>
[1,2]<stderr>: from . import initializers
[1,2]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/initializers/__init__.py", line 124, in <module>
[1,2]<stderr>: populate_deserializable_objects()
[1,2]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/initializers/__init__.py", line 82, in populate_deserializable_objects
[1,2]<stderr>: generic_utils.populate_dict_with_module_objects(
[1,2]<stderr>:AttributeError: module 'keras.utils.generic_utils' has no attribute 'populate_dict_with_module_objects'
[1,1]<stderr>:Traceback (most recent call last):
[1,1]<stderr>: File "./keras/keras_mnist_advanced.py", line 3, in <module>
[1,1]<stderr>: import keras
[1,1]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/__init__.py", line 20, in <module>
[1,1]<stderr>: from . import initializers
[1,1]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/initializers/__init__.py", line 124, in <module>
[1,1]<stderr>: populate_deserializable_objects()
[1,1]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/initializers/__init__.py", line 82, in populate_deserializable_objects
[1,1]<stderr>: generic_utils.populate_dict_with_module_objects(
[1,1]<stderr>:AttributeError: module 'keras.utils.generic_utils' has no attribute 'populate_dict_with_module_objects'
[1,3]<stderr>:Traceback (most recent call last):
[1,3]<stderr>: File "./keras/keras_mnist_advanced.py", line 3, in <module>
[1,3]<stderr>: import keras
[1,3]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/__init__.py", line 20, in <module>
[1,3]<stderr>: from . import initializers
[1,3]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/initializers/__init__.py", line 124, in <module>
[1,3]<stderr>: populate_deserializable_objects()
[1,3]<stderr>: File "/usr/local/lib/python3.7/dist-packages/keras/initializers/__init__.py", line 82, in populate_deserializable_objects
[1,3]<stderr>: generic_utils.populate_dict_with_module_objects(
[1,3]<stderr>:AttributeError: module 'keras.utils.generic_utils' has no attribute 'populate_dict_with_module_objects'
-------------------------------------------------------
Primary job terminated normally, but 1 process returned
a non-zero exit code. Per user-direction, the job has been aborted.
-------------------------------------------------------
--------------------------------------------------------------------------
mpirun detected that one or more processes exited with non-zero status, thus causing
the job to be terminated. The first process to do so was:
Process name: [[5508,1],2]
Exit code: 1
``` | open | 2021-06-15T00:55:56Z | 2022-08-11T15:38:47Z | https://github.com/horovod/horovod/issues/2983 | [
"bug"
] | jeff-yajun-liu | 3 |
recommenders-team/recommenders | machine-learning | 1,965 | [BUG] Error in lgithgbm quickstart due to change in API for early stopping | ### Description
<!--- Describe your issue/bug/request in detail -->
See https://github.com/microsoft/recommenders/actions/runs/5853247386/job/15866899086
```
tests/smoke/examples/test_notebooks_python.py F. [ 96%]
tests/integration/examples/test_notebooks_python.py . [100%]
=================================== FAILURES ===================================
________________________ test_lightgbm_quickstart_smoke ________________________
notebooks = ***'als_deep_dive': '/mnt/azureml/cr/j/25a5baf22b8c4b9db7a6c6abea64c9a5/exe/wd/examples/02_model_collaborative_filtering...rk_movielens': '/mnt/azureml/cr/j/25a5baf22b8c4b9db7a6c6abea64c9a5/exe/wd/examples/06_benchmarks/movielens.ipynb', ...***
output_notebook = 'output.ipynb', kernel_name = 'python3'
@pytest.mark.smoke
@pytest.mark.notebooks
def test_lightgbm_quickstart_smoke(notebooks, output_notebook, kernel_name):
notebook_path = notebooks["lightgbm_quickstart"]
pm.execute_notebook(
notebook_path,
output_notebook,
kernel_name=kernel_name,
parameters=dict(
MAX_LEAF=64,
MIN_DATA=20,
NUM_OF_TREES=100,
TREE_LEARNING_RATE=0.15,
EARLY_STOPPING_ROUNDS=20,
> METRIC="auc",
),
)
tests/smoke/examples/test_notebooks_python.py:124:
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
/azureml-envs/azureml_42c7166d644ccdc54af662a7cb4b4218/lib/python3.7/site-packages/papermill/execute.py:128: in execute_notebook
raise_for_execution_errors(nb, output_path)
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
nb = ***'cells': [***'id': '18084673', 'cell_type': 'markdown', 'source': '<span style="color:red; font-family:Helvetica Neue, ...end_time': '2023-08-14T08:35:23.189125', 'duration': 16.39677, 'exception': True***, 'nbformat': 4, 'nbformat_minor': 5***
output_path = 'output.ipynb'
def raise_for_execution_errors(nb, output_path):
"""Assigned parameters into the appropriate place in the input notebook
Parameters
----------
nb : NotebookNode
Executable notebook object
output_path : str
Path to write executed notebook
"""
error = None
for index, cell in enumerate(nb.cells):
if cell.get("outputs") is None:
continue
for output in cell.outputs:
if output.output_type == "error":
if output.ename == "SystemExit" and (output.evalue == "" or output.evalue == "0"):
continue
error = PapermillExecutionError(
cell_index=index,
exec_count=cell.execution_count,
source=cell.source,
ename=output.ename,
evalue=output.evalue,
traceback=output.traceback,
)
break
if error:
# Write notebook back out with the Error Message at the top of the Notebook, and a link to
# the relevant cell (by adding a note just before the failure with an HTML anchor)
error_msg = ERROR_MESSAGE_TEMPLATE % str(error.exec_count)
error_msg_cell = nbformat.v4.new_markdown_cell(error_msg)
error_msg_cell.metadata['tags'] = [ERROR_MARKER_TAG]
error_anchor_cell = nbformat.v4.new_markdown_cell(ERROR_ANCHOR_MSG)
error_anchor_cell.metadata['tags'] = [ERROR_MARKER_TAG]
# put the anchor before the cell with the error, before all the indices change due to the
# heading-prepending
nb.cells.insert(error.cell_index, error_anchor_cell)
nb.cells.insert(0, error_msg_cell)
write_ipynb(nb, output_path)
> raise error
E papermill.exceptions.PapermillExecutionError:
E ---------------------------------------------------------------------------
E Exception encountered at "In [8]":
E ---------------------------------------------------------------------------
E TypeError Traceback (most recent call last)
E /tmp/ipykernel_137/2864762[359](https://github.com/microsoft/recommenders/actions/runs/5853247386/job/15866899086#step:3:367).py in <module>
E 7 early_stopping_rounds=EARLY_STOPPING_ROUNDS,
E 8 valid_sets=lgb_valid,
E ----> 9 categorical_feature=cate_cols)
E
E TypeError: train() got an unexpected keyword argument 'early_stopping_rounds'
/azureml-envs/azureml_42c7166d644ccdc54af662a7cb4b[421](https://github.com/microsoft/recommenders/actions/runs/5853247386/job/15866899086#step:3:429)8/lib/python3.7/site-packages/papermill/execute.py:232: PapermillExecutionError
```
### In which platform does it happen?
<!--- Describe the platform where the issue is happening (use a list if needed) -->
<!--- For example: -->
<!--- * Azure Data Science Virtual Machine. -->
<!--- * Azure Databricks. -->
<!--- * Other platforms. -->
### How do we replicate the issue?
<!--- Please be specific as possible (use a list if needed). -->
<!--- For example: -->
<!--- * Create a conda environment for pyspark -->
<!--- * Run unit test `test_sar_pyspark.py` with `pytest -m 'spark'` -->
<!--- * ... -->
### Expected behavior (i.e. solution)
<!--- For example: -->
<!--- * The tests for SAR PySpark should pass successfully. -->
### Other Comments
| closed | 2023-08-14T08:59:51Z | 2023-08-18T22:45:30Z | https://github.com/recommenders-team/recommenders/issues/1965 | [
"bug"
] | miguelgfierro | 1 |
Yorko/mlcourse.ai | matplotlib | 360 | Add demo gif to README | __Disclaimer: This is a bot__
It looks like your repo is trending. The [github_trending_videos](https://www.instagram.com/github_trending_videos/) Instgram account automatically shows the demo gifs of trending repos in Github.
Your README doesn't seem to have any demo gifs. Add one and the next time the parser runs it will pick it up and post it on its Instagram feed. If you don't want to just close this issue we won't bother you again. | closed | 2018-10-02T07:09:26Z | 2018-10-09T15:47:16Z | https://github.com/Yorko/mlcourse.ai/issues/360 | [
"invalid"
] | va3093 | 0 |
autokey/autokey | automation | 995 | Update or remove new_features.rst | ### AutoKey is a Xorg application and will not function in a Wayland session. Do you use Xorg (X11) or Wayland?
Xorg
### Has this issue already been reported?
- [X] I have searched through the existing issues.
### Is this a question rather than an issue?
- [X] This is not a question.
### What type of issue is this?
Documentation
### Choose one or more terms that describe this issue:
- [ ] autokey triggers
- [ ] autokey-gtk
- [ ] autokey-qt
- [ ] beta
- [ ] bug
- [ ] critical
- [ ] development
- [X] documentation
- [ ] enhancement
- [ ] installation/configuration
- [ ] phrase expansion
- [ ] scripting
- [X] technical debt
- [ ] user interface
### Other terms that describe this issue if not provided above:
_No response_
### Which Linux distribution did you use?
N/A
### Which AutoKey GUI did you use?
None
### Which AutoKey version did you use?
0.96.0
### How did you install AutoKey?
Debs from AutoKey GitHub
### Can you briefly describe the issue?
new_features.rst was inherited from 0.90.4 and was never updated. It is a fossil. It contains a few useful lines about the high_level API that should be moved elsewhere (untill tatt API is upgraded or elimiated).
### Can the issue be reproduced?
None
### What are the steps to reproduce the issue?
_No response_
### What should have happened?
Anything useful should be extracted, probably to our wiki.
### What actually happened?
_No response_
### Do you have screenshots?
_No response_
### Can you provide the output of the AutoKey command?
_No response_
### Anything else?
_No response_
<br/>
<hr/>
<details><summary>This repo is using Opire - what does it mean? 👇</summary><br/>💵 Everyone can add rewards for this issue commenting <code>/reward 100</code> (replace <code>100</code> with the amount).<br/>🕵️♂️ If someone starts working on this issue to earn the rewards, they can comment <code>/try</code> to let everyone know!<br/>🙌 And when they open the PR, they can comment <code>/claim #995</code> either in the PR description or in a PR's comment.<br/><br/>🪙 Also, everyone can tip any user commenting <code>/tip 20 @josephj11</code> (replace <code>20</code> with the amount, and <code>@josephj11</code> with the user to tip).<br/><br/>📖 If you want to learn more, check out our <a href="https://docs.opire.dev">documentation</a>.</details> | open | 2024-12-21T04:04:08Z | 2024-12-26T13:22:30Z | https://github.com/autokey/autokey/issues/995 | [
"help-wanted",
"documentation",
"low-priority",
"technical debt"
] | josephj11 | 4 |
wandb/wandb | data-science | 9,203 | [Bug-App]: graphql: panic occurred: runtime error: invalid memory address or nil pointer dereference | ### Describe the bug
<!--- Describe your issue here --->
When I log into my account using Github, every wandb page shows this error
> graphql: panic occurred: runtime error: invalid memory address or nil pointer dereference
An application error occurred. | closed | 2025-01-07T20:19:48Z | 2025-01-08T15:52:50Z | https://github.com/wandb/wandb/issues/9203 | [
"ty:bug",
"a:app"
] | lhy0807 | 3 |
davidsandberg/facenet | computer-vision | 1,244 | Unable to use .pb in tensorflow's java api | I'm trying to use this pre-trained model in Java. I'm using Intellij Idea and I've added library dependency of TensorFlow and Added OpenCV via project structure.
`libraryDependency += "org.tensorflow" % "tensorflow" % "1.15.0"`
I've downloaded the VGGFace2 pre-trained model, and trying to use its .pb file and find face embeddings.
**Code:**
```
import org.opencv.core.Core;
import org.opencv.core.Mat;
import org.opencv.imgcodecs.Imgcodecs;
import org.tensorflow.*;
import java.nio.ByteBuffer;
import java.nio.FloatBuffer;
import java.nio.file.Paths;
import java.nio.file.Files;
import java.io.IOException;
public class DirectTensorflowTest {
public static void main(String[] args) {
// Load OpenCV library
System.loadLibrary(Core.NATIVE_LIBRARY_NAME);
// Path to the FaceNet model
String modelDir = "/home/zaryab/Downloads/20170512-110547";
try {
// Read the FaceNet model graph
byte[] graphDef = readAllBytesOrExit(Paths.get(modelDir, "20170512-110547.pb"));
// Import the graph definition into TensorFlow Graph
Graph g = new Graph();
g.importGraphDef(graphDef);
// Create a TensorFlow session with the imported graph
try (Session s = new Session(g)) {
// Load an image using OpenCV (replace this with your image loading logic)
String imagePath = "/home/zaryab/Desktop/IMG_20231124_171435.jpg";
Mat openCVMat = Imgcodecs.imread(imagePath);
// Convert OpenCV Mat to float array
float[] floatArray = convertMatToFloatArray(openCVMat);
// Byte array to TensorFlow Tensor
ByteBuffer byteBuffer = ByteBuffer.allocate(floatArray.length * Float.BYTES);
byteBuffer.asFloatBuffer().put(floatArray);
byte[] imageByte = byteBuffer.array();
FloatBuffer fb = ByteBuffer.wrap(imageByte).asFloatBuffer();
Tensor<Float> imageF = Tensor.create(new long[]{1, openCVMat.rows(), openCVMat.cols(), 1}, fb);
Tensor<Boolean> falseTensor = Tensors.create(false);
// Run the session to get embeddings
Tensor<Float> result = s.runner()
.feed("input", imageF)
.feed("phase_train", falseTensor)
.fetch("embeddings")
.run()
.get(0)
.expect(Float.class);
// Access the embeddings
float[][] embeddings = new float[1][(int) result.shape()[1]]; // Assuming shape[1] gives the embedding size
result.copyTo(embeddings);
System.out.println(embeddings);
result.close();
imageF.close();
}
} catch (IOException e) {
e.printStackTrace();
}
}
// Function to convert OpenCV Mat to float array
public static float[] convertMatToFloatArray(Mat mat) {
int rows = mat.rows();
int cols = mat.cols();
float[] floatArray = new float[rows * cols];
for (int i = 0; i < rows; i++) {
for (int j = 0; j < cols; j++) {
floatArray[i * cols + j] = (float) (mat.get(i, j)[0] / 255.0); // Normalize pixel values between 0 and 1
}
}
return floatArray;
}
// Function to read all bytes from a file
public static byte[] readAllBytesOrExit(java.nio.file.Path path) throws IOException {
return Files.readAllBytes(path);
}
}
```
I've taken some help from [https://github.com/davidsandberg/facenet/issues/659](url) but still getting error:
```
2023-12-01 11:19:51.689467: W tensorflow/core/framework/op_kernel.cc:1651] OP_REQUIRES failed at conv_ops.cc:491 : Invalid argument: input depth must be evenly divisible by filter depth: 1 vs 3
Exception in thread "main" java.lang.IllegalArgumentException: input depth must be evenly divisible by filter depth: 1 vs 3
[[{{node InceptionResnetV1/Conv2d_1a_3x3/convolution}}]]
at org.tensorflow.Session.run(Native Method)
at org.tensorflow.Session.access$100(Session.java:48)
at org.tensorflow.Session$Runner.runHelper(Session.java:326)
at org.tensorflow.Session$Runner.run(Session.java:276)
at DirectTensorflowTest.main(DirectTensorflowTest.java:111)
```
**Someone Guide me how can I successfully use this model in java using Tensorflow?** | open | 2023-12-01T06:28:13Z | 2023-12-01T06:28:13Z | https://github.com/davidsandberg/facenet/issues/1244 | [] | zaryabRiasat | 0 |
clovaai/donut | nlp | 238 | How to estimate the required video memory? | when i adjust processor.image_processor.size from {'height': 960, 'width': 720} to higher, GPU said CUDA out of memory..
my Video Card 8GB video, i wanna know, if i wanna use {'height': 2560, 'width': 1920}, how much VRAM will be used? does 16GB ok ? | closed | 2023-08-11T08:19:48Z | 2023-10-17T00:34:17Z | https://github.com/clovaai/donut/issues/238 | [] | chopin1998 | 1 |
babysor/MockingBird | deep-learning | 691 | 关于语音转换Voice Conversion(PPG based)中的ppg2mel.yaml修改里面的地址指向预训练好的文件夹 | **Summary[问题简述(一句话)]**
原文件似乎是为linux环境创建的,没有说明如何在windows下修改地址,请教一下如何在windows下修改ppg2mel.yaml文件里的地址,有好多文件我并没有在预处理后文件夹中找到
**Env & To Reproduce[复现与环境]**
环境:windows11,anaconda:python3.9.12
数据集文件夹:C:\test\test8\aidatatang_200zh
预处理生成文件夹:C:\test\test8\PPGVC\ppg2mel
我只能在预训练生成文件夹里找到原文件4,5,6行的同名的文件,这么修改对不对?剩下的7-14行怎么修改?
**Screenshots[截图(如有)]**
直接运行报错
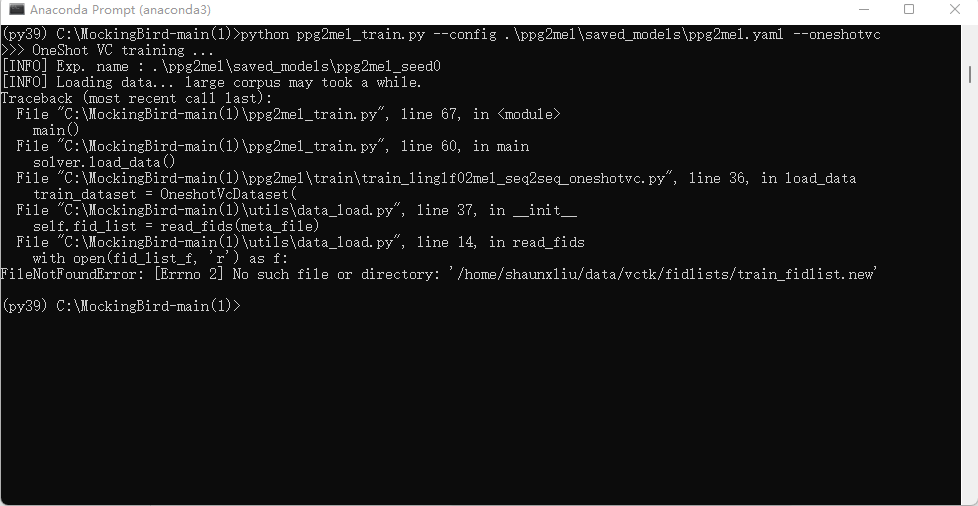
| open | 2022-07-31T07:03:28Z | 2022-07-31T07:19:04Z | https://github.com/babysor/MockingBird/issues/691 | [] | ms903x1 | 0 |
joouha/euporie | jupyter | 50 | Incorrect Description of Changing cell type | [Changing Cell's Type](https://euporie.readthedocs.io/en/latest/apps/notebook.html#changing-a-cell-s-type) has incorrect description .
It is explaining `how to close notebook` instead of `how to change cell's type`.
Corresponding [file](https://github.com/joouha/euporie/blob/main/docs/apps/notebook.rst#changing-a-cells-type) in repo.
Maybe we can add good first issue | closed | 2022-12-06T03:03:02Z | 2023-01-23T19:30:58Z | https://github.com/joouha/euporie/issues/50 | [
"good first issue"
] | DivyanshuBist | 3 |
cleanlab/cleanlab | data-science | 1,019 | exact issue name should be listed for each issue type in the Datalab Issue Type Guide | Otherwise it's hard to know how to run an audit for this issue type (eg. data valuation say). | closed | 2024-02-20T09:15:40Z | 2024-02-24T02:09:33Z | https://github.com/cleanlab/cleanlab/issues/1019 | [
"next release"
] | jwmueller | 1 |
matplotlib/matplotlib | data-visualization | 29,711 | [ENH]: directional antialiasing filter | ### Problem
Currently, the antialiasing filter used by imshow() is applied both in the x and the y direction. There are cases where "image-like" data is highly sampled in one direction (requiring antialiasing) but not in the other, and also has some nans; an example is a kymograph like subfigure 1c at https://www.nature.com/articles/s41467-017-01462-y/figures/1
For such data, the current default interpolation settings (antialiased + rgba (because *one* of the directions is oversampled) can yield not nice artefacts, in particular at the boundary between "data" and "nan":
```python
from pylab import *
vals = np.full((50, 500), np.nan)
for i, row in enumerate(vals):
n = 200 + np.random.randint(300)
row[:n] = np.sin(np.arange(n) / 10 + i / 5) # make some fake data
axs = figure(layout="constrained", figsize=(10, 10)).subplots(
3, 3, subplot_kw={"xticks": [], "yticks": []})
for i, interpolation in enumerate([None, "none", "antialiased"]):
for j, stage in enumerate([None, "data", "rgba"]):
axs[i, j].text(0, 0, f"{interpolation=}, {stage=}", bbox={"fc": "w"})
axs[i, j].imshow(vals.T, aspect="auto", origin="lower",
interpolation=interpolation, interpolation_stage=stage)
show()
```

For this specific dataset, the best settings currently available is probably to go back to interpolation="data". But ideally, it would be nice if it was possible to have a separable antialiasing filter (I don't actually know if the current one is separable), so that one can apply it (here) only in the vertical direction (ideally in rgba space).
### Proposed solution
_No response_ | open | 2025-03-06T11:49:33Z | 2025-03-17T14:31:07Z | https://github.com/matplotlib/matplotlib/issues/29711 | [
"New feature",
"topic: images"
] | anntzer | 17 |
waditu/tushare | pandas | 1,043 | 具体赞助多少才能获取2000积分? | 现在需要抓取tushare_pro的数据 需要赞助多少才能拥有2000积分,5000积分又需要赞助多少呢? | closed | 2019-05-14T01:54:38Z | 2019-05-20T06:14:05Z | https://github.com/waditu/tushare/issues/1043 | [] | Anthony0722 | 1 |
chiphuyen/stanford-tensorflow-tutorials | nlp | 4 | lack of definition of CONTENT_WEIGHT, STYLE_WEIGHT(in style_transfer_sols.py), prev_layer_name (in vgg_model_sols.py) | HI, Tks for the post, very helpful.
As title, I found several variable undefined,
for prev_layer_name, I think, it should be prev_name.name, however ':' is not accepted as scope name. So I changed ':' to '_', and it works
For CONTENT_WEIGHT and STYLE_WEIGHT, how to define it ?
(of course, omitted the weight could let the program keep running)
Tks
Larry | closed | 2017-03-01T06:27:02Z | 2017-03-01T22:58:01Z | https://github.com/chiphuyen/stanford-tensorflow-tutorials/issues/4 | [] | tcglarry | 1 |
jackmpcollins/magentic | pydantic | 194 | Are there some models in ollama can support function calling or object return? | closed | 2024-04-27T13:49:57Z | 2024-05-12T23:30:08Z | https://github.com/jackmpcollins/magentic/issues/194 | [] | chaos369 | 6 | |
pydantic/pydantic | pydantic | 10,525 | Unable to import JsonSchemaHandler | ### Initial Checks
- [X] I confirm that I'm using Pydantic V2
### Description

Hey, this piece of code has not been working since today morning (GMT +5:30). Checked all latest commits but still not sure why this error is popping up. This ends up messing up my llama-index imports as well.
### Example Code
```Python
from pydantic import GetJsonSchemaHandler
#error -> ImportError: cannot import name 'GetJsonSchemaHandler' from 'pydantic' (/home/jovyan/DBLLM/smebot2/lib/python3.11/site-packages/pydantic/__init__.cpython-311-x86_64-linux-gnu.so)
```
### Python, Pydantic & OS Version
```Text
pydantic version: 2.9.2
pydantic-core version: 2.23.4
pydantic-core build: profile=release pgo=false
install path: /home/jovyan/DBLLM/smebot2/lib/python3.11/site-packages/pydantic
python version: 3.11.9 | packaged by conda-forge | (main, Apr 19 2024, 18:36:13) [GCC 12.3.0]
platform: Linux-5.15.0-121-generic-x86_64-with-glibc2.35
related packages: fastapi-0.112.2 typing_extensions-4.12.2
commit: unknown
```
| closed | 2024-10-01T06:04:04Z | 2024-10-01T15:00:14Z | https://github.com/pydantic/pydantic/issues/10525 | [
"bug V2",
"pending"
] | ShivamGoswami-TCL | 2 |
erdewit/ib_insync | asyncio | 468 | How to cancel all open orders and close all positions? | Currently I have orders that have OCA trailing stop and take profit for an order attached and at EOD I like to close all my positions. As such below code doesn't exit all positions, they tend to keep the opposite orders open. If I keep them open, in the falling market they r getting triggered and become a working market order. I have to manually cancel those each day. Can you please advice how we close all positions.
```
def cancel_order(ib, order):
try:
ib.cancelOrder(order)
except Exception as ex:
print("Unable to cancel the order Reason: {}".format(ex))
return
exit_orders = ib.oneCancelsAll([tp_order, sl_order], "OCA_{}".format(str(datetime.now())), 1)
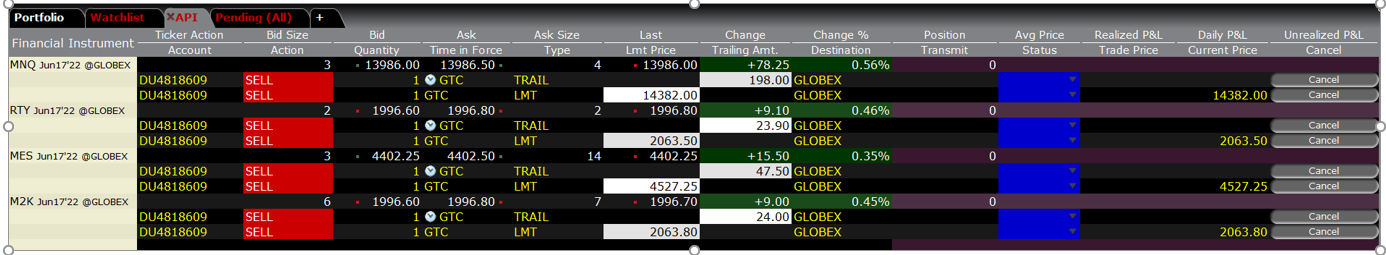
``` | closed | 2022-04-25T20:57:03Z | 2022-05-07T15:16:09Z | https://github.com/erdewit/ib_insync/issues/468 | [] | msacs09 | 3 |
remsky/Kokoro-FastAPI | fastapi | 30 | 🔄 Automatic master to develop merge failed | Automatic merge from master to develop failed.
Please resolve this manually
Workflow run: https://github.com/remsky/Kokoro-FastAPI/actions/runs/12737419308 | closed | 2025-01-12T22:00:10Z | 2025-01-12T22:01:12Z | https://github.com/remsky/Kokoro-FastAPI/issues/30 | [
"merge-failed",
"automation"
] | github-actions[bot] | 0 |
explosion/spaCy | machine-learning | 12,976 | span_ruler is not working on ENT patterns | <!-- NOTE: For questions or install related issues, please open a Discussion instead. -->
Text : "A 50-year-old male patient lodges chief complaint pain in lower right side of chest for three days associated with burning in epigastrium. History is cuffed with sputum."
Get Ent in Doc :
0 | 50-year | 2 | 5 | Age
0 | male | 7 | 8 | Sex
0 | pain in lower right side of chest | 12 | 19 | chief complaint
0 | three days | 20 | 22 | time_unit
0 | burning in epigastrium | 24 | 27 | chief complaint

Now ,
Add span_ruler in Pipe.
`patterns = srsly.read_jsonl("./drugs_patterns_3.jsonl")
span_ruler = nlp.add_pipe("span_ruler" , name= "span_ruler_3" , after="entity_ruler_2" , config= { } )
span_ruler.add_patterns(patterns)`
drugs_patterns_3.jsonl File have 1 patterns ,
`{"label":"cc_time_unit","pattern":[ {"ENT_TYPE": "cc"} , {"LOWER": "for"} , {"ENT_TYPE": "time_unit"} ]}`
Output came :

But Output come as per logic ,
0 | pain in lower right side of chest | 12 | 19 | chief complaint
0 | three days | 20 | 22 | time_unit
## How to reproduce the behaviour
<!-- Include a code example or the steps that led to the problem. Please try to be as specific as possible. -->
Above Side Explain All Details. Still if any need , Please update to me.
## Your Environment
<!-- Include details of your environment. You can also type `python -m spacy info --markdown` and copy-paste the result here.-->
* Operating System: Windows 11
* Python Version Used: 3.7
* spaCy Version Used: 3.6
* Environment Information:
| closed | 2023-09-12T12:50:18Z | 2023-09-13T06:21:35Z | https://github.com/explosion/spaCy/issues/12976 | [
"feat / spanruler"
] | kamlesh0606 | 1 |
junyanz/pytorch-CycleGAN-and-pix2pix | deep-learning | 974 | Cityscapes label2photo evaluation issue | Hello!
I am trying to reproduce the results in Table 1 of the pix2pix paper by evaluating the results generated from pix2pix on the Cityscapes (label2photo) dataset using your scripts.
I resized all leftImg8bit images and gtFine images (incl. *_color.png, *_labels.png and *_instances.png, although I probably only need labels) of the original dataset to (256, 256). I also renamed the generated images to the original images' names from leftImg8bit/val folder.
Although the generated images seemed quite realistic, the evaluation results were bad and pretty far from the reported ones in the paper. Suspecting that something was wrong with the evaluation, I ran the evaluation on the original resized images (created as *_real_B.png in the results and renamed) to generate the ground truth evaluation results. Results showed that something is wrong with my evaluation since I am getting similar results to what I got from the pix2pix generated images (which are bad).
I am trying to understand what I'm doing wrong here. Which images exactly do I need to resize in the original dataset in order to correctly run the script? As stated above, I resized all leftImg8bit images and gtFine images. Did I only need to resize the left8bitImg images?
A little help would be appreciated.
Thank you in advance | open | 2020-04-01T12:07:29Z | 2020-04-01T18:36:54Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/974 | [] | parvanitis15 | 1 |
albumentations-team/albumentations | machine-learning | 2,293 | [Speed up] GaussianBlur | Benchmark shows that `imgaug` has faster GaussianBlur implementation => need to learn from it and fix. | closed | 2025-01-24T15:56:46Z | 2025-02-18T18:26:41Z | https://github.com/albumentations-team/albumentations/issues/2293 | [
"Speed Improvements"
] | ternaus | 3 |
microsoft/qlib | deep-learning | 1,859 | error: subprocess-exited-with-error note: This error originates from a subprocess, and is likely not a problem with pip. error: metadata-generation-failed. | ## ❓ Questions and Help
I try to install pyqlib in anaconda prompt cmd of windows 11, suffering from this when in subprogress collecting scs:
error: subprocess-exited-with-error
note: This error originates from a subprocess, and is likely not a problem with pip.
error: metadata-generation-failed. | closed | 2024-11-11T11:13:06Z | 2024-11-13T02:45:23Z | https://github.com/microsoft/qlib/issues/1859 | [
"question"
] | Agvensome | 1 |
keras-team/autokeras | tensorflow | 1,719 | Would it be possible to add jax? | In according to Google's [JAX](https://jax.readthedocs.io/en/latest/notebooks/Common_Gotchas_in_JAX.html), would it be possible to include JAX into `AutoKeras`, please?
I guess, if no array-slicing would be performed, JAX should be fully working?
https://github.com/keras-team/autokeras/blob/c51da2dd87b195ab3bd0941ea70862c3cf66e9a9/autokeras/adapters/input_adapters.py#L25-L29 | open | 2022-05-02T20:01:43Z | 2022-05-02T20:01:43Z | https://github.com/keras-team/autokeras/issues/1719 | [] | Anselmoo | 0 |
kizniche/Mycodo | automation | 963 | upstream variable name change breaks input: SCD30 CircuitPython (fix also enclosed here) | Please DO NOT OPEN AN ISSUE:
- If your Mycodo version is not the latest release version, please update your device before submitting your issue (unless your issue is related to not being able to upgrade). Your problem might already be solved.
- If your issue has been addressed before (i.e., duplicated issue), please ask in the original issue.
Please complete as many of the sections below, if applicable, to provide the most information that may help with investigating your issue. The details requested potentially affect which options to pursue. The small amount of time you spend completing the template will also help those providing assistance by reducing the time required to help you.
### Describe the problem/bug
The CircuitPython SCD30 sensor input currently errors at every reading, saying there is no parameter called "eCO2". This is because Adafruit [updated](https://github.com/adafruit/Adafruit_CircuitPython_SCD30/commit/500c4b9d704dbc60bbb064c4dc1b5b444e5cf0e0#diff-eaa2c02e845bd776b0ca8e9fc462498b19e161a63b11fa3cf27db0d6d3f1164a) their library a few months ago, changing that variable name.
### Versions:
- Mycodo Version: current version, 8.9.2
- Raspberry Pi Version: 3B+
- Raspbian OS Version: Buster
### Reproducibility
Please list specific setup details that are involved and the steps to reproduce the behavior:
1. Connect SCD30 i2c sensor
2. Install CircuitPython Dependency for SCD30
3. Add sensor and activate input
4. Check Daemon logs and see error:
`2021-03-26 10:24:15,292 - ERROR - mycodo.inputs.scd30_circuitpython_d830dda1 - InputModule raised an exception when taking a reading: 'SCD30' object has no attribute 'eCO2'
Traceback (most recent call last):
File "/var/mycodo-root/mycodo/inputs/base_input.py", line 128, in read
self._measurements = self.get_measurement()
File "/home/pi/Mycodo/mycodo/inputs/scd30_circuitpython.py", line 95, in get_measurement
co2 = self.scd.eCO2
AttributeError: 'SCD30' object has no attribute 'eCO2'`
### Expected behavior
Sensor takes readings per normal
### Additional context
I fixed this myself by editing one line of code, replacing "eCO2" with "CO2"
Here's the line:
https://github.com/kizniche/Mycodo/blob/master/mycodo/inputs/scd30_circuitpython.py#L95
Thanks for making Mycodo, it's awesome!
AKA
| closed | 2021-03-26T14:34:29Z | 2021-04-24T18:22:34Z | https://github.com/kizniche/Mycodo/issues/963 | [
"bug",
"Fixed and Committed"
] | AKAMEDIASYSTEM | 1 |
hzwer/ECCV2022-RIFE | computer-vision | 96 | 训练最后会crash | 感谢楼主的辛苦付出,测了几段视频,大运动效果和纹理很稳定,重影很少。
所以尝试复现一下效果,用了你给的100个数据做了一下测试,跑20个epoch,跑完最后会crash
机器是P40,双卡。
脚本是:
`python3 -u -m torch.distributed.launch --nproc_per_node=1 train.py --local_rank=0`
所有输出的日志:
nohup: ignoring input
457012eb-9ba5-4d55-9bbe-a3f39dfe66c5:82576:82576 [0] NCCL INFO Bootstrap : Using [0]eth1:9.73.145.207<0>
457012eb-9ba5-4d55-9bbe-a3f39dfe66c5:82576:82576 [0] NCCL INFO NET/Plugin : No plugin found (libnccl-net.so).
457012eb-9ba5-4d55-9bbe-a3f39dfe66c5:82576:82576 [0] misc/ibvwrap.cc:63 NCCL WARN Failed to open libibverbs.so[.1]
457012eb-9ba5-4d55-9bbe-a3f39dfe66c5:82576:82576 [0] NCCL INFO NET/Socket : Using [0]eth1:9.73.145.207<0>
NCCL version 2.4.8+cuda10.1
457012eb-9ba5-4d55-9bbe-a3f39dfe66c5:82576:82608 [0] NCCL INFO Setting affinity for GPU 0 to 01f000,0000001f
457012eb-9ba5-4d55-9bbe-a3f39dfe66c5:82576:82608 [0] NCCL INFO Using 256 threads, Min Comp Cap 6, Trees enabled up to size -2
457012eb-9ba5-4d55-9bbe-a3f39dfe66c5:82576:82608 [0] NCCL INFO comm 0x7f8880001a40 rank 0 nranks 1 cudaDev 0 nvmlDev 0 - Init COMPLETE
------------this evaluating is start....--------
/tools/conda/envs/rife/lib/python3.9/site-packages/torch/nn/functional.py:3000: UserWarning: The default behavior for interpolate/upsample with float scale_factor changed in 1.6.0 to align with other frameworks/libraries, and uses scale_factor directly, instead of relying on the computed output size. If you wish to keep the old behavior, please set recompute_scale_factor=True. See the documentation of nn.Upsample for details.
warnings.warn("The default behavior for interpolate/upsample with float scale_factor changed "
eval time: 4.768563508987427
------------this evaluating is end--------
training...
running epoch is 0
epoch:0 0/3 time:0.31+2.41 loss_l1:6.6461e-02
epoch:0 4/3 time:0.00+0.49 loss_l1:6.6527e-02
epoch:0 2/3 time:0.88+0.39 loss_l1:6.1271e-02
------------last barrier is end....--------
running epoch is 1
epoch:1 0/3 time:0.47+0.38 loss_l1:6.6090e-02
epoch:1 1/3 time:0.00+0.38 loss_l1:6.4821e-02
epoch:1 2/3 time:0.00+0.38 loss_l1:5.8615e-02
------------last barrier is end....--------
running epoch is 2
epoch:2 0/3 time:0.47+0.38 loss_l1:6.2020e-02
epoch:2 1/3 time:0.00+0.37 loss_l1:6.0823e-02
epoch:2 2/3 time:0.00+0.37 loss_l1:6.3036e-02
------------last barrier is end....--------
running epoch is 3
epoch:3 0/3 time:0.48+0.37 loss_l1:6.6802e-02
epoch:3 1/3 time:0.00+0.37 loss_l1:6.0749e-02
epoch:3 2/3 time:0.00+0.37 loss_l1:5.8643e-02
------------last barrier is end....--------
running epoch is 4
epoch:4 0/3 time:0.45+0.38 loss_l1:6.1430e-02
epoch:4 1/3 time:0.00+0.37 loss_l1:5.8594e-02
epoch:4 2/3 time:0.00+0.37 loss_l1:5.8876e-02
------------this evaluating is start....--------
eval time: 1.697829008102417
------------this evaluating is end--------
------------last barrier is end....--------
running epoch is 5
epoch:5 0/3 time:2.19+0.39 loss_l1:6.0860e-02
epoch:5 1/3 time:0.00+0.39 loss_l1:5.6333e-02
epoch:5 2/3 time:0.00+0.39 loss_l1:5.6805e-02
------------last barrier is end....--------
running epoch is 6
epoch:6 0/3 time:0.49+0.39 loss_l1:5.8091e-02
epoch:6 1/3 time:0.00+0.39 loss_l1:5.8843e-02
epoch:6 2/3 time:0.00+0.38 loss_l1:5.5708e-02
------------last barrier is end....--------
running epoch is 7
epoch:7 0/3 time:0.45+0.37 loss_l1:5.5774e-02
epoch:7 1/3 time:0.00+0.37 loss_l1:5.7322e-02
epoch:7 2/3 time:0.00+0.37 loss_l1:5.1374e-02
------------last barrier is end....--------
running epoch is 8
epoch:8 0/3 time:0.49+0.39 loss_l1:5.5491e-02
epoch:8 1/3 time:0.00+0.39 loss_l1:5.4187e-02
epoch:8 2/3 time:0.00+0.39 loss_l1:4.9916e-02
------------last barrier is end....--------
running epoch is 9
epoch:9 0/3 time:0.52+0.37 loss_l1:5.7686e-02
epoch:9 1/3 time:0.00+0.37 loss_l1:5.2956e-02
epoch:9 2/3 time:0.00+0.37 loss_l1:4.9873e-02
------------this evaluating is start....--------
eval time: 1.6385705471038818
------------this evaluating is end--------
------------last barrier is end....--------
running epoch is 10
epoch:10 0/3 time:2.12+0.37 loss_l1:5.2009e-02
epoch:10 1/3 time:0.00+0.37 loss_l1:4.7775e-02
epoch:10 2/3 time:0.00+0.37 loss_l1:5.0613e-02
------------last barrier is end....--------
running epoch is 11
epoch:11 0/3 time:0.51+0.37 loss_l1:4.8291e-02
epoch:11 1/3 time:0.00+0.37 loss_l1:4.5139e-02
epoch:11 2/3 time:0.00+0.37 loss_l1:4.7583e-02
------------last barrier is end....--------
running epoch is 12
epoch:12 0/3 time:0.48+0.37 loss_l1:4.8362e-02
epoch:12 1/3 time:0.00+0.37 loss_l1:4.9962e-02
epoch:12 2/3 time:0.00+0.37 loss_l1:4.7941e-02
------------last barrier is end....--------
running epoch is 13
epoch:13 0/3 time:0.49+0.38 loss_l1:4.6211e-02
epoch:13 1/3 time:0.00+0.37 loss_l1:4.7942e-02
epoch:13 2/3 time:0.00+0.37 loss_l1:4.4998e-02
------------last barrier is end....--------
running epoch is 14
epoch:14 0/3 time:0.48+0.38 loss_l1:4.4784e-02
epoch:14 1/3 time:0.00+0.37 loss_l1:4.7616e-02
epoch:14 2/3 time:0.00+0.37 loss_l1:4.3033e-02
------------this evaluating is start....--------
eval time: 1.6448850631713867
------------this evaluating is end--------
------------last barrier is end....--------
running epoch is 15
epoch:15 0/3 time:2.13+0.38 loss_l1:4.2588e-02
epoch:15 1/3 time:0.00+0.37 loss_l1:4.5357e-02
epoch:15 2/3 time:0.00+0.38 loss_l1:4.2175e-02
------------last barrier is end....--------
running epoch is 16
epoch:16 0/3 time:0.61+0.39 loss_l1:4.3911e-02
epoch:16 1/3 time:0.00+0.37 loss_l1:4.1421e-02
epoch:16 2/3 time:0.00+0.37 loss_l1:4.3421e-02
------------last barrier is end....--------
running epoch is 17
epoch:17 0/3 time:0.47+0.38 loss_l1:4.3027e-02
epoch:17 1/3 time:0.00+0.37 loss_l1:4.2315e-02
epoch:17 2/3 time:0.00+0.37 loss_l1:4.3238e-02
------------last barrier is end....--------
running epoch is 18
epoch:18 0/3 time:0.49+0.38 loss_l1:4.0845e-02
epoch:18 1/3 time:0.00+0.37 loss_l1:3.9075e-02
epoch:18 2/3 time:0.00+0.37 loss_l1:4.0612e-02
------------last barrier is end....--------
running epoch is 19
epoch:19 0/3 time:0.47+0.38 loss_l1:4.0664e-02
epoch:19 1/3 time:0.00+0.37 loss_l1:4.0039e-02
epoch:19 2/3 time:0.00+0.37 loss_l1:4.1695e-02
------------this evaluating is start....--------
eval time: 1.6566226482391357
------------this evaluating is end--------
------------last barrier is end....--------
terminate called without an active exception
terminate called without an active exception
Traceback (most recent call last):
File "/tools/conda/envs/rife/lib/python3.9/runpy.py", line 197, in _run_module_as_main
return _run_code(code, main_globals, None,
File "/tools/conda/envs/rife/lib/python3.9/runpy.py", line 87, in _run_code
exec(code, run_globals)
File "/tools/conda/envs/rife/lib/python3.9/site-packages/torch/distributed/launch.py", line 261, in <module>
main()
File "/tools/conda/envs/rife/lib/python3.9/site-packages/torch/distributed/launch.py", line 256, in main
raise subprocess.CalledProcessError(returncode=process.returncode,
subprocess.CalledProcessError: Command '['/tools/conda/envs/rife/bin/python3', '-u', 'train.py', '--local_rank=0', '--local_rank=0']' died with <Signals.SIGABRT: 6>.
跑完最后一个epoch,保存了模型,处理完dist.barrier()后按道理应该就结束了,但不知道为什么会crash,看起来和分布式有关,不知道楼主有什么经验吗?感谢感谢 | closed | 2021-01-20T07:20:51Z | 2021-03-19T10:00:05Z | https://github.com/hzwer/ECCV2022-RIFE/issues/96 | [] | xiazhenyz | 8 |
docarray/docarray | pydantic | 939 | Figure out handling of Union types | Many features of our current implementation rely on the type hints under the hood to infer behaviour.
This break whenever the type hint is of form `Union` or `Optional`, since those cannot be used in instance and subclass checks.
This is especially problematic, since our type `Tensor` is a Union under the hood, and it is the type used by our pre buit documents. For example, `find()` and `stack()` don't currently work with `Image`.
The simplest workaround is probably to check if a type is union or optional, extract the types inside them, and perform subclass/instance checks on those | closed | 2022-12-14T09:10:44Z | 2023-01-03T08:51:56Z | https://github.com/docarray/docarray/issues/939 | [
"DocArray v2"
] | JohannesMessner | 0 |
Hironsan/BossSensor | computer-vision | 20 | run boss_train.py error | I user tensorflow 0.12, and I put some pictures in boss and other directories, but when I run boss_train.py ,it report error:
Traceback (most recent call last):
File "/home/zxx/PycharmProjects/BossSensor/boss_train.py", line 176, in <module>
dataset.read()
File "/home/zxx/PycharmProjects/BossSensor/boss_train.py", line 35, in read
X_train, X_test, y_train, y_test = train_test_split(images, labels, test_size=0.3, random_state=random.randint(0, 100))
File "/usr/lib/python2.7/dist-packages/sklearn/cross_validation.py", line 1556, in train_test_split
arrays = check_arrays(*arrays, **options)
File "/usr/lib/python2.7/dist-packages/sklearn/utils/validation.py", line 287, in check_arrays
array.ndim)
ValueError: Found array with dim 4. Expected <= 2
Could you give me some help, thanks.
| closed | 2017-03-12T15:49:31Z | 2019-03-22T10:32:04Z | https://github.com/Hironsan/BossSensor/issues/20 | [] | asd51731 | 1 |
raphaelvallat/pingouin | pandas | 19 | Epsilon and Mauchly interaction in rm_anova2 differ from JASP | open | 2019-04-26T17:16:24Z | 2019-08-06T15:14:55Z | https://github.com/raphaelvallat/pingouin/issues/19 | [
"bug :boom:",
"help wanted :bell:",
"invalid :triangular_flag_on_post:"
] | raphaelvallat | 3 | |
MycroftAI/mycroft-core | nlp | 3,156 | ModuleNotFoundError: No module named 'xdg.BaseDirectory' | **Describe the bug**
Hi, all. I cloned mycroft-core on my computer and installed it. But I get the following error when running.
**To Reproduce**
```sh
~$ source .venv/bin/activate
(.venv) blink@blink-eq:~/mycroft-core$ mycroft-skill-testrunner
Traceback (most recent call last):
File "<frozen runpy>", line 198, in _run_module_as_main
File "<frozen runpy>", line 88, in _run_code
File "/home/blink/mycroft-core/test/integrationtests/skills/runner.py", line 32, in <module>
from test.integrationtests.skills.skill_tester import MockSkillsLoader
File "/home/blink/mycroft-core/test/integrationtests/skills/skill_tester.py", line 44, in <module>
from mycroft.messagebus.message import Message
File "/home/blink/mycroft-core/mycroft/__init__.py", line 17, in <module>
from mycroft.api import Api
File "/home/blink/mycroft-core/mycroft/api/__init__.py", line 22, in <module>
from mycroft.configuration import Configuration
File "/home/blink/mycroft-core/mycroft/configuration/__init__.py", line 15, in <module>
from .config import Configuration, LocalConf, RemoteConf
File "/home/blink/mycroft-core/mycroft/configuration/config.py", line 22, in <module>
import xdg.BaseDirectory
ModuleNotFoundError: No module named 'xdg.BaseDirectory'
(.venv) blink@blink-eq:~/mycroft-core$ mycroft-pip install xdg
Requirement already satisfied: xdg in ./.venv/lib/python3.11/site-packages (6.0.0)
```
**Expected behavior**
A clear and concise description of what you expected to happen.
**Log files**
If possible, add log files from `/var/log/mycroft/` to help explain your problem.
You may also include screenshots, however screenshots of log files are often difficult to read and parse.
If you are running Mycroft, the [Support Skill](https://github.com/MycroftAI/skill-support) helps to automate gathering this information. Simply say "Create a support ticket" and the Skill will put together a support package and email it to you.
**Environment (please complete the following information):**
- Device type: [ desktop x86_64]
- OS: [debian bookworm]
- Mycroft-core version: [ git main ]
- Other versions: [e.g. Adapt v0.3.7]
| closed | 2023-09-13T05:24:25Z | 2024-09-08T08:15:27Z | https://github.com/MycroftAI/mycroft-core/issues/3156 | [
"bug"
] | yjdwbj | 1 |
graphql-python/graphene-django | django | 751 | How do I make the pagination like using django rest framework? | I didn't see any docs about pagination, will that be coming in future? | closed | 2019-08-18T02:58:49Z | 2019-10-25T11:15:06Z | https://github.com/graphql-python/graphene-django/issues/751 | [] | tinc0709 | 3 |
aiogram/aiogram | asyncio | 1,388 | Pretty work with state | ### aiogram version
3.x
### Problem
I've been writing telegram bots on aiogram for quite some time.
I like everything, but I don’t really like working with states. You have to write a lot of typical code.
You must describe the state model, and then write handlers for each item.
I believe that this can be implemented a little more convenient for developers
### Possible solution
I believe that it is possible to write a wrapper over the existing state system that can process a state in one line
### Alternatives
I know about state scenes in 3.x, but I want something a little different
### Code example
```python3
# For example:
# We init model with data types and other additional information(custom validators for users answers and other)
class Form(StatesGroup):
name: str = State(custom_validator) # custom validator is a link to a function that returns a validator of the user's response
age: int = State(custom_validator)
@form_router.message(CommandStart())
async def command_start(message: Message, state: FSMContext) -> None:
name = await Form.name.process_state()
await message.answer(f"Your name is {name}")
# After we work with user input value without typical code and boring work with states.
```
### Additional information
If I'm wrong on some issues, please correct me, this is my first micro contribution to open source, sorry if something is wrong | closed | 2024-01-02T12:17:32Z | 2024-08-17T13:56:59Z | https://github.com/aiogram/aiogram/issues/1388 | [
"enhancement"
] | dop3file | 4 |
junyanz/pytorch-CycleGAN-and-pix2pix | deep-learning | 707 | I reduced the image size . but result is poor | Hello, I appreciate to your great research and implementations.
I would like to thank you for doing such a great job.
Anyway, I have a Question.
When I learned with 3x256x256 images , it was fine
but I learn with 1x50x50 image, it is poor
I want to learn with 1(channel) x 50 x 50 image from this model.
So I modify some options
images = A 4000 : B 4000
input_nc = 1
output_nc = 1
load size = 50
crop size = 50 (or preprocessing = none)
batch size = 64 ( when I learn with 3x256x256 image , it was 3)
epoch = 200
(Discriminator loss is very poor. this is got 0.00xx when 10~20 epoch.)
I want know that what I modified options is influence to poor generate
and if it is influenced my learn, I want to know how tune the options
Thank you.
| closed | 2019-07-17T08:17:48Z | 2019-07-19T03:52:11Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/707 | [] | Realdr4g0n | 2 |
xlwings/xlwings | automation | 1,865 | Access to chart sheets Excel | #### OS (Windows 10)
#### Versions of xlwings, Excel and Python (xlwings 0.27.2, Microsoft Excel 2010, Python 3.6)
#### Describe your issue (incl. Traceback!)
Hallo,
I have a chart sheet in my Excel book.
If I try to find the sheets of my book the once which contains the chart is not listed out.
Also, if i manually activate the chart sheet and ask xlwings to list the charts, I obtain an empty list.
How can I access a chart sheet form xlwings API and interact with the chart?
Thank you in advance.
Alberto
``` | closed | 2022-03-16T10:49:04Z | 2022-03-23T13:14:48Z | https://github.com/xlwings/xlwings/issues/1865 | [] | AlbertoCasetta | 1 |
anselal/antminer-monitor | dash | 54 | Support for avalon 741 and 821 | Anyone have an example of the JSON RPC data that cgminer returns?
I have some that are arriving and wanted to add support for this before they are here.
| open | 2018-01-17T14:37:19Z | 2018-02-12T08:51:26Z | https://github.com/anselal/antminer-monitor/issues/54 | [
":pick: miner_support"
] | sergioclemente | 7 |
ets-labs/python-dependency-injector | asyncio | 431 | service is not being injected, instead Provide object is present | ```
@auth_blp.route('/')
class Login(MethodView):
@auth_blp.arguments(schema=AuthLoginFormSchema, location="form", as_kwargs=True)
@auth_blp.response(status_code=200, schema=AuthLoginRespSchema)
def post(self, **kwargs):
"""
POST to login
"""
user = User.get_by_user_name(
user_name=kwargs["username"]
)
if not verify_password(
kwargs["password"],
user.password
):
abort(401, message="User credentials don't match.")
encoded_jwt, expire = create_access_token(
data={
"blacklist_id": str(uuid4()),
"id": user.id,
"group": user.group,
}
)
return {
"access_token": encoded_jwt.decode("utf-8")
}
@auth_blp.response(status_code=200, schema=AuthLogoutRespSchema)
@auth_blp.doc(security=[{"Oauth2PasswordBearer": []}])
@inject
def delete(
self,
token_user_service: TokenUserService = Provide[TokenUserContainer.token_user_service]
):
"""
DELETE to logout
"""
raise Exception(token_user_service.get_token())
```
[Public Repo of the code](https://github.com/zero-shubham/flask_tm/blob/master/tm/tm/auth/routes.py)
Getting error - **AttributeError: 'Provide' object has no attribute 'get_token'**
I'm wiring the contain with the whole package. (check main.py) | closed | 2021-03-22T01:54:48Z | 2021-03-22T17:11:58Z | https://github.com/ets-labs/python-dependency-injector/issues/431 | [
"question"
] | zero-shubham | 2 |
keras-team/keras | python | 20,912 | What is the python package google used for? | The package was added to requirements.txt in https://github.com/keras-team/keras/commit/699e4c3174976d9b308a60458259a0e8544e0777 and is still present in requirements-common.txt as of https://github.com/keras-team/keras/commit/e045b6abe42ed9335a85b553cc9c609b76a8a54c. The package `google` on [pypi](https://pypi.org/project/google/) refers to a package providing search engine bindings via the `googlesearch` module. However there is no match for the string `googlesearch` in either the keras or tensorboard projects. Was google added to requirements.txt in addition to protobuf by mistake because of the `from google.protobuf import text_format` import? | closed | 2025-02-16T20:25:12Z | 2025-02-20T18:38:46Z | https://github.com/keras-team/keras/issues/20912 | [
"type:support",
"keras-team-review-pending"
] | loqs | 2 |
WZMIAOMIAO/deep-learning-for-image-processing | deep-learning | 512 | SSD error | **System information**
* Have I written custom code:
* OS Platform(e.g., window10 or Linux Ubuntu 16.04):
* Python version:
* Deep learning framework and version(e.g., Tensorflow2.1 or Pytorch1.3):
* Use GPU or not:
* CUDA/cuDNN version(if you use GPU):
* The network you trained(e.g., Resnet34 network):
**Describe the current behavior**
**Error info / logs**
| closed | 2022-04-10T13:17:39Z | 2022-04-11T00:49:01Z | https://github.com/WZMIAOMIAO/deep-learning-for-image-processing/issues/512 | [] | shangshanghuiliyi | 2 |
deepset-ai/haystack | nlp | 9,016 | Add run_async for `AzureOpenAIDocumentEmbedder` | We should be able to reuse the implementation once it is made for the `OpenAIDocumentEmbedder` | open | 2025-03-11T11:07:12Z | 2025-03-23T07:08:53Z | https://github.com/deepset-ai/haystack/issues/9016 | [
"Contributions wanted!",
"P2"
] | sjrl | 0 |
e2b-dev/code-interpreter | jupyter | 23 | 401 Unauthorized when pushing new template | Would appreciate any help here. I tried logging out and logging back in, still seeing the same error.
```
gonzalonunez@Gonzalos-MacBook-Pro e2b % npx e2b template build -c "/root/.jupyter/start-up.sh"
Found sandbox template or4llhf3739qmlizq7u1 fluent-production-sandbox <-> ./e2b.toml
Found ./Dockerfile that will be used to build the sandbox template.
Requested build for the sandbox template or4llhf3739qmlizq7u1 fluent-production-sandbox
Login Succeeded
Building docker image...
[+] Building 1.1s (14/14) FINISHED docker:desktop-linux
=> [internal] load build definition from Dockerfile 0.0s
=> => transferring dockerfile: 1.37kB 0.0s
=> [internal] load metadata for docker.io/e2bdev/code-interpreter:latest 1.0s
=> [auth] e2bdev/code-interpreter:pull token for registry-1.docker.io 0.0s
=> [internal] load .dockerignore 0.0s
=> => transferring context: 2B 0.0s
=> [1/8] FROM docker.io/e2bdev/code-interpreter:latest@sha256:ea769ea3ca9793958ba47167affd6420f0563f43c2a262960ffde2cab7ad9f2 0.0s
=> => resolve docker.io/e2bdev/code-interpreter:latest@sha256:ea769ea3ca9793958ba47167affd6420f0563f43c2a262960ffde2cab7ad9f2 0.0s
=> [internal] load build context 0.0s
=> => transferring context: 3.27kB 0.0s
=> CACHED [2/8] RUN apt update && apt install -y curl gnupg odbc-postgresql 0.0s
=> CACHED [3/8] RUN curl https://packages.microsoft.com/keys/microsoft.asc | tee /etc/apt/trusted.gpg.d/microsoft.asc 0.0s
=> CACHED [4/8] RUN curl https://packages.microsoft.com/config/debian/12/prod.list | sed 's/ signed-by=[^ ]*] /] /' | tee /et 0.0s
=> CACHED [5/8] RUN apt-get update 0.0s
=> CACHED [6/8] RUN ACCEPT_EULA=Y apt-get install -y msodbcsql18 0.0s
=> CACHED [7/8] COPY requirements.txt . 0.0s
=> CACHED [8/8] RUN pip install -r requirements.txt 0.0s
=> exporting to image 0.0s
=> => exporting layers 0.0s
=> => exporting manifest sha256:cea8f2b46a3b44e483f33fe8c6120242bef7f925a0b59ef510fceaf280ea5f95 0.0s
=> => exporting config sha256:81c63da58b71993260eef8d1dee103e60f19d9942c10e4edf5a54ba6ee40c06d 0.0s
=> => exporting attestation manifest sha256:51e5968a8895dc40ef3518c3e1fc8a4aac8c16393c231b5dbbb9a9d3c11c8ada 0.0s
=> => exporting manifest list sha256:f7ed59fa12fef26ba246736d7c44264dc22970c30cbe87284c2c70289fa2275e 0.0s
=> => naming to docker.e2b.dev/e2b/custom-envs/or4llhf3739qmlizq7u1:2b4b2493-38cc-4583-a260-68a46596bb31 0.0s
Docker image built.
Pushing docker image...
The push refers to repository [docker.e2b.dev/e2b/custom-envs/or4llhf3739qmlizq7u1]
896d5b1b5dfc: Waiting
891494355808: Waiting
f6d9e20bee33: Waiting
57b6589e7635: Waiting
f26285f1a25b: Waiting
a99509a32390: Waiting
5d4c3b3f6734: Waiting
e56e34e87760: Waiting
752667ab907a: Waiting
d76e704b1be9: Waiting
73a707b4cb54: Waiting
240b0335931b: Waiting
a091fb61d64f: Waiting
8e804c790eec: Waiting
b2ab5d29389b: Waiting
58bdfb10d51a: Waiting
bf2c3e352f3d: Waiting
d518c3c1be5f: Waiting
795d8e6db33d: Waiting
bc0ba82972c2: Waiting
6582c62583ef: Waiting
c6cf28de8a06: Waiting
946285778af4: Waiting
580d0969862c: Waiting
unexpected status from HEAD request to https://docker.e2b.dev/v2/e2b/custom-envs/or4llhf3739qmlizq7u1/blobs/sha256:d518c3c1be5f702c70fe1b70f04246b9a02b5ccfd4d9f5887ac83f648f7863cc: 401 Unauthorized
Error: Command failed: docker push docker.e2b.dev/e2b/custom-envs/or4llhf3739qmlizq7u1:2b4b2493-38cc-4583-a260-68a46596bb31
at __node_internal_genericNodeError (node:internal/errors:932:15)
at checkExecSyncError (node:child_process:890:11)
at Object.execSync (node:child_process:962:15)
at e.<anonymous> (/Users/gonzalonunez/fluent/node_modules/.pnpm/@e2b+cli@0.5.5/node_modules/@e2b/cli/src/commands/template/build.ts:298:23) {
status: 1,
signal: null,
output: [ null, null, null ],
pid: 99393,
stdout: null,
stderr: null
}
``` | closed | 2024-06-21T16:55:20Z | 2024-06-23T19:08:03Z | https://github.com/e2b-dev/code-interpreter/issues/23 | [] | gonzalonunez | 11 |
ShishirPatil/gorilla | api | 371 | Cannot load hugging face dataset for function calling | To repro, try the following:
```
from datasets import load_dataset
dataset = load_dataset("gorilla-llm/Berkeley-Function-Calling-Leaderboard")
```
You will get the following error
```
File "pyarrow/_json.pyx", line 308, in pyarrow._json.read_json
File "pyarrow/error.pxi", line 154, in pyarrow.lib.pyarrow_internal_check_status
File "pyarrow/error.pxi", line 91, in pyarrow.lib.check_status
pyarrow.lib.ArrowInvalid: JSON parse error: Column(/execution_result/[]) changed from number to string in row 13
``` | closed | 2024-04-19T06:51:17Z | 2024-04-23T04:49:59Z | https://github.com/ShishirPatil/gorilla/issues/371 | [] | sebastiangonsal | 1 |
python-restx/flask-restx | flask | 83 | Upgrade packages in light of deprecation warnings | Currently the unit tests are always throwing the following DeprecationWarnings:
> app/lib/python3.8/site-packages/flask_restx/model.py:12
> /private/var/www/html/api/app/lib/python3.8/site-packages/flask_restx/model.py:12: DeprecationWarning: Using or importing the ABCs from 'collections' instead of from 'collections.abc' is deprecated since Python 3.3, and in 3.9 it will stop working
> from collections import OrderedDict, MutableMapping
>
> app/lib/python3.8/site-packages/flask_restx/api.py:28
> /private/var/www/html/api/app/lib/python3.8/site-packages/flask_restx/api.py:28: DeprecationWarning: The import 'werkzeug.cached_property' is deprecated and will be removed in Werkzeug 1.0. Use 'from werkzeug.utils import cached_property' instead.
> from werkzeug import cached_property
>
> app/lib/python3.8/site-packages/flask_restx/swagger.py:12
> /private/var/www/html/api/app/lib/python3.8/site-packages/flask_restx/swagger.py:12: DeprecationWarning: Using or importing the ABCs from 'collections' instead of from 'collections.abc' is deprecated since Python 3.3, and in 3.9 it will stop working
> from collections import OrderedDict, Hashable
It would be a good idea to upgrade them to avoid broken releases in the near future | closed | 2020-03-12T23:48:35Z | 2020-03-22T23:35:16Z | https://github.com/python-restx/flask-restx/issues/83 | [
"enhancement"
] | andreixk | 2 |
pyro-ppl/numpyro | numpy | 1,045 | FR: Fail with informative error message when same plate name used with inconsistent dimensions | Currently reusing the same plate name with different arguments to e.g. dim will cause problems. Models like this could be rejected with an informative error message quickly, before confusing problems which can include arrays changing shape happen. See below for the current behaviour.
```py
import sys
import numpyro
import numpyro.distributions as dist
from jax import numpy as jnp
from jax import random
from numpyro.infer import SVI, Trace_ELBO
from numpyro.infer.autoguide import AutoDelta
from numpyro.infer.initialization import init_to_median
from numpyro.optim import Adam
def irt2pl(ncls, resp, word_subsample_size=None):
nstud, nitems = resp.shape
difficulty_offsets = numpyro.sample(
"difficulty_offsets",
dist.TransformedDistribution(
dist.Normal(0, 1).expand([ncls - 1]), dist.transforms.OrderedTransform()
),
)
with numpyro.plate("nstud", nstud, dim=-1):
abilities = numpyro.sample("abilities", dist.Normal())
with numpyro.plate("nitems", nitems, dim=-1):
difficulties = numpyro.sample("difficulties", dist.Normal())
discriminations = numpyro.sample("discriminations", dist.HalfNormal())
offset_difficulties = jnp.expand_dims(difficulties, 1) + jnp.expand_dims(
difficulty_offsets, 0
)
print("abilities.shape", abilities.shape)
print("discriminations.shape", discriminations.shape)
predictor = jnp.expand_dims(abilities, 1) * jnp.expand_dims(discriminations, 0)
cutpoints = jnp.expand_dims(
offset_difficulties * jnp.expand_dims(discriminations, 0), 0
)
with numpyro.plate("nstud", nstud, dim=-2), numpyro.plate("nitems", nitems, dim=-1):
numpyro.sample("resp", dist.OrderedLogistic(predictor, cutpoints), obs=resp)
resp = jnp.array(
[[0, 1, 4, 3], [0, 1, 4, 3], [3, 4, 4, 4], [2, 2, 4, 4], [2, 2, 4, 4], [1, 2, 3, 3]]
)
optim = Adam(0.1, 0.8, 0.99)
elbo = Trace_ELBO()
guide = AutoDelta(irt2pl, init_loc_fn=init_to_median())
rng_key = random.PRNGKey(42)
svi = SVI(irt2pl, guide, optim, loss=elbo)
svi.run(rng_key, 200, 5, resp)
print(guide(5, resp))
```
Running gets an error like:
```
abilities.shape (6,)
discriminations.shape (4,)
abilities.shape (6,)
discriminations.shape (4,)
abilities.shape (6,)
discriminations.shape (4,)
abilities.shape (6,)
discriminations.shape (4,)
abilities.shape (6,)
discriminations.shape (4,)
abilities.shape (6, 6)
discriminations.shape (4,)
Traceback (most recent call last):
File "broken.py", line 55, in <module>
svi.run(rng_key, 200, 5, resp)
...
File "broken.py", line 31, in irt2pl
predictor = jnp.expand_dims(abilities, 1) * jnp.expand_dims(discriminations, 0)
File "/path/to/python3.8/site-packages/jax/_src/numpy/lax_numpy.py", line 5256, in deferring_binary_op
return binary_op(self, other)
File "/home/frankier/edu/doc/vocabmodel/.direnv/python-3.8.6/lib/python3.8/site-packages/jax/_src/numpy/lax_numpy.py", line 396, in fn
x1, x2 = _promote_args(numpy_fn.__name__, x1, x2)
File "/path/to/python3.8/site-packages/jax/_src/numpy/lax_numpy.py", line 333, in _promote_args
return _promote_shapes(fun_name, *_promote_dtypes(*args))
File "/path/to/python3.8/site-packages/jax/_src/numpy/lax_numpy.py", line 251, in _promote_shapes
result_rank = len(lax.broadcast_shapes(*shapes))
File "/path/to/python3.8/site-packages/jax/_src/util.py", line 198, in wrapper
return cached(bool(config.x64_enabled), *args, **kwargs)
File "/path/to/python3.8/site-packages/jax/_src/util.py", line 191, in cached
return f(*args, **kwargs)
File "/path/to/python3.8/site-packages/jax/_src/lax/lax.py", line 97, in broadcast_shapes
raise ValueError("Incompatible shapes for broadcasting: {}"
ValueError: Incompatible shapes for broadcasting: ((6, 1, 6), (1, 1, 4))
``` | closed | 2021-05-28T06:24:00Z | 2021-06-01T00:03:23Z | https://github.com/pyro-ppl/numpyro/issues/1045 | [
"warnings & errors"
] | frankier | 0 |
Farama-Foundation/PettingZoo | api | 844 | [Proposal] Use GitHub Issue Forms | ### Proposal
I propose to use [GitHub Issue Forms](https://docs.github.com/en/communities/using-templates-to-encourage-useful-issues-and-pull-requests/configuring-issue-templates-for-your-repository#creating-issue-forms) when an issue is created in this repo
### Motivation
This facilitates the proper filling in of information when an issue is created. Information can be marked as required. I created a PR in the [Gymnasium](https://github.com/Farama-Foundation/Gymnasium) repo and was asked to create one here as well :)
### Alternatives
Stick with the current solution
### Additional context
Creating an issue could look like this (screenshot taken from [stable-baselines3 repo](https://github.com/DLR-RM/stable-baselines3):
<img width="945" alt="image" src="https://user-images.githubusercontent.com/17867382/198526648-296e879b-829a-4da2-8140-16a4065d2f19.png">
See [this repo](https://github.com/tobirohrer/dummy-repo-git-issue-template) to see implementation details.
### Checklist
- [x] I have checked that there is no similar [issue](https://github.com/Farama-Foundation/Gymnasium/issues) in the repo (**required**)
| closed | 2022-10-31T09:03:12Z | 2022-10-31T16:29:10Z | https://github.com/Farama-Foundation/PettingZoo/issues/844 | [] | tobirohrer | 1 |
plotly/dash | data-visualization | 3,226 | add/change type annotations to satisfy mypy and other tools | With the release of dash 3.0 our CI/CD fails for stuff that used to work.
Here's a minimum working example:
```python
from dash import Dash, dcc, html
from dash.dependencies import Input, Output
from typing import Callable
app = Dash(__name__)
def create_layout() -> html.Div:
return html.Div([
dcc.Input(id='input-text', type='text', value='', placeholder='Enter text'),
html.Div(id='output-text')
])
app.layout = create_layout
@app.callback(Output('output-text', 'children'), Input('input-text', 'value'))
def update_output(value: str) -> str:
return f'You entered: {value}'
if __name__ == '__main__':
app.run(debug=True, port=9000)
```
running mypy on this file results in:
```bash
$ mypy t.py --strict
t.py:1: error: Skipping analyzing "dash": module is installed, but missing library stubs or py.typed marker [import-untyped]
t.py:2: error: Skipping analyzing "dash.dependencies": module is installed, but missing library stubs or py.typed marker [import-untyped]
t.py:2: note: See https://mypy.readthedocs.io/en/stable/running_mypy.html#missing-imports
t.py:15: error: Untyped decorator makes function "update_output" untyped [misc]
Found 3 errors in 1 file (checked 1 source file)
```
which we used to solve by adding `# type: ignore[misc]` to every `callback` call and by adding
```toml
[[tool.mypy.overrides]]
module = [
"dash.*",
"dash_ag_grid",
"dash_bootstrap_components.*",
"plotly.*",
]
ignore_missing_imports = true
```
to our `pyproject.toml`.
However, when updating to version 3, we get:
```bash
$ mypy --strict t.py
t.py:8: error: Returning Any from function declared to return "Div" [no-any-return]
t.py:13: error: Property "layout" defined in "Dash" is read-only [misc]
t.py:15: error: Call to untyped function "Input" in typed context [no-untyped-call]
t.py:15: error: Call to untyped function "Output" in typed context [no-untyped-call]
t.py:15: error: Call to untyped function "callback" in typed context [no-untyped-call]
t.py:15: note: Error code "no-untyped-call" not covered by "type: ignore" comment
Found 5 errors in 1 file (checked 1 source file)
```
without changing anything else.
This can't be intended behavior, right? How to fix that?
community post: https://community.plotly.com/t/dash-3-0-fails-mypy/91308 | open | 2025-03-18T10:25:57Z | 2025-03-20T16:30:13Z | https://github.com/plotly/dash/issues/3226 | [
"bug",
"P2"
] | gothicVI | 6 |
Miserlou/Zappa | django | 1,295 | PermissionError trying to make first deployment | ## Context
I'm trying to do my first Zappa deployment and hit an exception:
Calling deploy for stage dev..
Creating project-dev-ZappaLambdaExecutionRole IAM Role..
Creating zappa-permissions policy on project-dev-ZappaLambdaExecutionRole IAM Role.
Oh no! An error occurred! :(
==============
Traceback (most recent call last):
File "/.../.direnv/python-3.6.3/lib/python3.6/site-packages/zappa/cli.py", line 2610, in handle
sys.exit(cli.handle())
File "/.../.direnv/python-3.6.3/lib/python3.6/site-packages/zappa/cli.py", line 505, in handle
self.dispatch_command(self.command, stage)
File "/.../.direnv/python-3.6.3/lib/python3.6/site-packages/zappa/cli.py", line 539, in dispatch_command
self.deploy(self.vargs['zip'])
File "/.../.direnv/python-3.6.3/lib/python3.6/site-packages/zappa/cli.py", line 709, in deploy
self.create_package()
File "/.../.direnv/python-3.6.3/lib/python3.6/site-packages/zappa/cli.py", line 2171, in create_package
disable_progress=self.disable_progress
File "/.../.direnv/python-3.6.3/lib/python3.6/site-packages/zappa/core.py", line 504, in create_lambda_zip
copytree(cwd, temp_project_path, symlinks=False, ignore=shutil.ignore_patterns(*excludes))
File "/.../.direnv/python-3.6.3/lib/python3.6/site-packages/zappa/utilities.py", line 54, in copytree
copytree(s, d, symlinks, ignore)
File "/.../.direnv/python-3.6.3/lib/python3.6/site-packages/zappa/utilities.py", line 54, in copytree
copytree(s, d, symlinks, ignore)
File "/.../.direnv/python-3.6.3/lib/python3.6/site-packages/zappa/utilities.py", line 54, in copytree
copytree(s, d, symlinks, ignore)
File "/.../.direnv/python-3.6.3/lib/python3.6/site-packages/zappa/utilities.py", line 32, in copytree
shutil.copystat(src, dst)
File "/.../.direnv/python-3.6.3/lib/python3.6/shutil.py", line 218, in copystat
lookup("chflags")(dst, st.st_flags, follow_symlinks=follow)
PermissionError: [Errno 1] Operation not permitted: '/var/folders/dy/0lgxvt4n0nb4c6h0sgq66bl80000gn/T/1513418744/.venv/include/python2.7'
## Your Environment
* Python environment managed by `direnv`
* macOS 10.13.2
* Zappa version used: 0.45.1
* Operating System and Python version: 3.6.3
* The output of `pip freeze`:
```
argcomplete==1.9.2
base58==0.2.4
boto3==1.5.1
botocore==1.8.15
certifi==2017.11.5
cfn-flip==1.0.0
chardet==3.0.4
click==6.7
defusedxml==0.5.0
dj-database-url==0.4.2
Django==2.0
docutils==0.14
durationpy==0.5
future==0.16.0
gunicorn==19.7.1
hjson==3.0.1
idna==2.6
jmespath==0.9.3
kappa==0.6.0
lambda-packages==0.19.0
oauthlib==2.0.6
placebo==0.8.1
psycopg2==2.7.3.1
PyJWT==1.5.3
python-dateutil==2.6.1
python-slugify==1.2.4
python3-openid==3.1.0
pytz==2017.3
PyYAML==3.12
requests==2.18.4
requests-oauthlib==0.8.0
s3transfer==0.1.12
six==1.11.0
social-auth-app-django==2.0.0
social-auth-core==1.5.0
toml==0.9.3
tqdm==4.19.1
troposphere==2.1.2
Unidecode==0.4.21
urllib3==1.22
Werkzeug==0.12
whitenoise==3.3.1
wsgi-request-logger==0.4.6
zappa==0.45.1
```
* Your `zappa_settings.py`:
```
{
"dev": {
"aws_region": null,
"django_settings": "mysite.settings",
"profile_name": "default",
"project_name": "project",
"runtime": "python3.6",
"s3_bucket": "xxxxx"
}
}
``` | closed | 2017-12-16T10:10:51Z | 2018-02-25T16:57:13Z | https://github.com/Miserlou/Zappa/issues/1295 | [] | rgov | 7 |
JoeanAmier/TikTokDownloader | api | 408 | 下载视频提示[配置文件cookie参数未登陆,数据获取已提前结束] |
使用chrome浏览器能正常获取cookie,下载视频提示[配置文件cookie参数未登陆,数据获取已提前结束],实际浏览器的账号是正常登陆状态,账号发布有53个视频,只能获取到18个视频,其他获取不到也无法下载.
 | open | 2025-02-26T03:00:44Z | 2025-02-28T02:07:53Z | https://github.com/JoeanAmier/TikTokDownloader/issues/408 | [] | yzy64 | 10 |
aidlearning/AidLearning-FrameWork | jupyter | 54 | Cannot use sklearn? | I install the sklearn lib using "pip3 install sklearn",then I import sklearn in the python3,there were many errors like this:
ImportError: cannot import name 'SemLock'
......
ImportError: This platform lacks a functioning sem_open implementation, therefore, the required synchronization primitives needed will not function, see issue 3770.
I found that maybe we must mount /dev/shm first then install python.
https://blog.csdn.net/u010454261/article/details/80216581 (This web is Chinese)
Could you please solve this problem? Or dose the Aid Learning not support sklearn? Thank you! | closed | 2019-09-21T05:26:17Z | 2020-08-02T01:43:43Z | https://github.com/aidlearning/AidLearning-FrameWork/issues/54 | [] | zwdnet | 1 |
clovaai/donut | computer-vision | 39 | Input size parameter clarification | I'm trying to run my own fine-tunning for document parsing. When building the train configuration I wondered: Is the input_size parameter related to the the size of the images in the dataset or is it only used for the Swin transfomer to create the embedding windows?
In case it's the second. When should it be customized and what constraints apply to the values provided?
Thank you! | closed | 2022-08-30T19:39:45Z | 2024-08-01T23:25:52Z | https://github.com/clovaai/donut/issues/39 | [] | leitouran | 3 |
kizniche/Mycodo | automation | 1,126 | Atlas Pump I2C connection doesn't understand command |
### Describe the problem/bug
We are using Mycodo with Raspberry Pi4 to automate pH monitoring of water. Everything seems to be working fine except for the pump. When given the command the pump changes the light color from blue to green to red and then back to blue. Troubleshooting with Atlas, we found that pump can not understand the command
### Versions:
- Mycodo Version: 8.12.9
- Raspberry Pi Version: 4+
- Raspbian OS Version: Bulls Eye
### Reproducibility
Please list specific setup details that are involved and the steps to reproduce the behavior:
1. Connect the circuit as represented in the [blog ](https://kylegabriel.com/projects/2020/06/automated-hydroponic-system-build.html)
2. Calibrate pH
3. While calibrating pump, press "Dispense Volume", the pump displays blue->green->red->blue
4. Check the logs below
`2021-12-21 14:29:36,699 - DEBUG - mycodo.outputs.pump_atlas_ezo_pmp_bfb22612 - Calibration command: D,10.0
2021-12-21 14:29:36,701 - INFO - mycodo.outputs.pump_atlas_ezo_pmp_bfb22612 - Command returned: None
2021-12-21 14:29:36,702 - INFO - mycodo.outputs.pump_atlas_ezo_pmp_bfb22612 - Device Calibrated?: None
2021-12-21 14:30:06,252 - DEBUG - mycodo.controllers.controller_conditional_da513008 - Conditional check. pH: 8.316
2021-12-21 14:30:06,257 - DEBUG - mycodo.controllers.controller_conditional_da513008 - pH is dangerously high: 8.316. Should be < 7.0. Dispensing 1 ml acid
2021-12-21 14:30:06,334 - DEBUG - mycodo.outputs.pump_atlas_ezo_pmp_bfb22612 - output_on_off(on, 0, sec, 10.0, 0.0, True)
2021-12-21 14:30:06,373 - DEBUG - mycodo.outputs.pump_atlas_ezo_pmp_bfb22612 - Output bfb22612-153b-4603-b907-112630869018 CH0 (pump) on for 10.0 seconds. Output returned: None
2021-12-21 14:30:06,376 - DEBUG - mycodo.outputs.pump_atlas_ezo_pmp_bfb22612 - EZO-PMP command: D,*
2021-12-21 14:30:16,384 - DEBUG - mycodo.outputs.pump_atlas_ezo_pmp_bfb22612 - output_on_off(off, 0, None, 0.0, 0.0, True)
2021-12-21 14:30:16,385 - DEBUG - mycodo.outputs.pump_atlas_ezo_pmp_bfb22612 - EZO-PMP command: X
2021-12-21 14:30:16,386 - DEBUG - mycodo.outputs.pump_atlas_ezo_pmp_bfb22612 - Output bfb22612-153b-4603-b907-112630869018 CH0 (pump) OFF at 2021-12-21 14:30:16. Output returned: None
`
Is there an error somewhere in Mycodo or is probably the pump faulty?
| closed | 2021-12-21T14:33:26Z | 2022-03-28T22:40:25Z | https://github.com/kizniche/Mycodo/issues/1126 | [] | vipul-khatana | 1 |
ContextLab/hypertools | data-visualization | 163 | add soundtrack | include an animation flag that will (optionally) add a dramatic soundtrack (or a user-selected audio file) to animations.
eventually we could automatically generate an audio soundtrack using the data. 🎶 | open | 2017-10-24T17:16:31Z | 2017-10-24T17:16:31Z | https://github.com/ContextLab/hypertools/issues/163 | [] | jeremymanning | 0 |
frappe/frappe | rest-api | 31,815 | Dark Theme Color Issue | Like Button not showing clearly

Frappe Framework: v15.59.0 (version-15) | open | 2025-03-20T03:46:32Z | 2025-03-20T03:46:49Z | https://github.com/frappe/frappe/issues/31815 | [
"bug"
] | nilpatel42 | 0 |
ludwig-ai/ludwig | data-science | 3,462 | Image Classification: Config | The Config.

It starts a syntax error, it should actually be:
config = {
'input_features': [
{
'name': 'image_path',
'type': 'image',
'preprocessing': {'num_processes': 4},
'encoder': 'stacked_cnn',
'conv_layers': [
{'num_filters': 32, 'filter_size': 3, 'pool_size': 2, 'pool_stride': 2},
{'num_filters': 64, 'filter_size': 3, 'pool_size': 2, 'pool_stride': 2, 'dropout': 0.4}
],
'fc_layers': [{'output_size': 128, 'dropout': 0.4}]
}
],
'output_features': [{'name': 'label', 'type': 'binary'}],
'trainer': {'epochs': 5}
}
| closed | 2023-07-13T11:01:32Z | 2024-10-18T13:48:23Z | https://github.com/ludwig-ai/ludwig/issues/3462 | [] | AnaMiguelRodrigues1 | 4 |
twopirllc/pandas-ta | pandas | 176 | Pandas TA strategy method goes into infinite loop: Windows Freeze Support Error | @twopirllc I was trying to run the below code on a dataframe
```python
df.ta.strategy("Momentum")
print(df)
```
It goes into infinite loop and consumes all CPU on my windows 10 laptop.
Then I need to kill all VSCodium and python exes.
Could you please help?
_Originally posted by @rahulmr in https://github.com/twopirllc/pandas-ta/issues/138#issuecomment-751864255_ | closed | 2020-12-28T21:16:15Z | 2023-03-17T19:53:02Z | https://github.com/twopirllc/pandas-ta/issues/176 | [
"question",
"info"
] | twopirllc | 20 |
huggingface/transformers | tensorflow | 36,348 | None | closed | 2025-02-22T18:12:03Z | 2025-02-22T18:25:42Z | https://github.com/huggingface/transformers/issues/36348 | [] | Hashmapw | 0 | |
dmlc/gluon-cv | computer-vision | 844 | Failed loading Parameter | I have trained yolov3 on my own dataset and got params,and i have read some issues like #229,but still failed loading Parameter
In my training script train_yolo3.py, i added the following lines:
class VOCLike(VOCDetection):
CLASSES=['number']
def __init__(self,root,splits,transform=None,index_map=None,preload_label=True):
super(VOCLike,self).__init__(root,splits,transform,index_map,preload_label)
and changes as follows:
def get_dataset(dataset, args):
if dataset.lower() == 'voc':
train_dataset = VOCLike(root='/VOCtemplate', splits=((2018, 'train'),))
val_dataset = VOCLike(root='/VOCtemplate', splits=((2018, 'val'),))
val_metric = VOC07MApMetric(iou_thresh=0.5, class_names=val_dataset.classes)
and my inference code as follow:
from gluoncv import model_zoo,data,utils
net=model_zoo.get_model('yolo3_darknet53_custom',classes=['number'],pretrained=False)
net.load_parameters('yolo3_darknet53_voc_best.params')
the error log:
Traceback (most recent call last):
File "inference_yolo_v3.py", line 13, in <module>
net.load_parameters('yolo3_darknet53_voc_best.params')
File "/home/user/anaconda3/envs/gluoncv/lib/python3.6/site-packages/mxnet/gluon/block.py", line 405, in load_parameters
params[name]._load_init(loaded[name], ctx, cast_dtype=cast_dtype)
File "/home/user/anaconda3/envs/gluoncv/lib/python3.6/site-packages/mxnet/gluon/parameter.py", line 251, in _load_init
self.name, str(self.shape), str(data.shape))
AssertionError: Failed loading Parameter 'yolov30_yolooutputv30_conv0_weight' from saved params: shape incompatible expected (18, 0, 1, 1) vs saved (75, 1024, 1, 1)
Thanks for any help.
| closed | 2019-06-29T14:39:52Z | 2021-05-26T04:04:10Z | https://github.com/dmlc/gluon-cv/issues/844 | [] | narutobns | 3 |
streamlit/streamlit | streamlit | 10,162 | Redirect to last viewed page in multipage app after `st.login` | ### Checklist
- [X] I have searched the [existing issues](https://github.com/streamlit/streamlit/issues) for similar feature requests.
- [X] I added a descriptive title and summary to this issue.
### Summary
We're soon launching native authentication in Streamlit (see https://github.com/streamlit/streamlit/issues/8518). If you initiate the auth flow in a multipage app, it will afterwards redirect to the main page of the app. It would be nicer to redirect to the page you last viewed.
### Why?
We're losing track of the current page because auth makes the app initiate a new session: To show the OAuth flow, we redirect to the identity provider. After completing the flow, they redirect back to the app again. This means Streamlit initiates a completely new session and doesn't have the state from before the auth flow anymore. This can by itself be annoying but it might be especially confusing in a multipage app, where you land somewhere completely differently from what you viewed before.
### How?
_No response_
### Additional Context
_No response_ | open | 2025-01-10T23:48:40Z | 2025-01-10T23:48:54Z | https://github.com/streamlit/streamlit/issues/10162 | [
"type:enhancement",
"feature:st.login"
] | jrieke | 1 |
onnx/onnx | deep-learning | 5,885 | [Feature request] Provide a means to convert to numpy array without byteswapping | ### System information
ONNX 1.15
### What is the problem that this feature solves?
Issue onnx/tensorflow-onnx#1902 in tf2onnx occurs on big endian systems, and it is my observation that attributes which end up converting to integers are incorrectly byteswapped because the original data resided within a tensor. If `numpy_helper.to_array()` could be updated to optionally not perform byteswapping, then that could help solve this issue.
### Alternatives considered
As an alternative, additional logic could be added in tf2onnx to perform byteswapping on the data again, but this seems excessive.
### Describe the feature
I believe this feature is necessary to improve support for big endian systems.
### Will this influence the current api (Y/N)?
_No response_
### Feature Area
converters
### Are you willing to contribute it (Y/N)
Yes
### Notes
_No response_ | closed | 2024-01-31T20:58:56Z | 2024-02-02T16:52:35Z | https://github.com/onnx/onnx/issues/5885 | [
"topic: enhancement"
] | tehbone | 4 |
desec-io/desec-stack | rest-api | 389 | Extend API for easy access to TLSA, SSHFP, OPENPGPKEY, ... records | Setting these "DNSSEC-only" record types often requires complicated computations, let's do them on the API side wherever safely possible. | open | 2020-05-18T09:27:23Z | 2024-10-07T17:21:00Z | https://github.com/desec-io/desec-stack/issues/389 | [
"enhancement",
"api"
] | nils-wisiol | 0 |
mage-ai/mage-ai | data-science | 5,411 | Specify scheduler name on k8s executor | Is your feature request related to a problem? Please describe.
Currently, the scheduler pod name is not configurable in the k8sconfig file for Mage. This limitation forces users to rely on the default Kubernetes scheduler, which can be restrictive in scenarios where alternative schedulers are required, such as those optimized for specific workloads, custom scheduling algorithms, or multi-tenant environments. The lack of flexibility can be frustrating for users who need more control over scheduling policies to optimize resource usage or meet specific performance requirements.
Describe the solution you’d like
I would like the ability to specify the scheduler pod name directly in the k8sconfig file for Mage. This addition would allow users to easily select and switch between different schedulers without needing to modify other parts of the configuration. The change would involve adding a field where users can define the desired scheduler's name, enabling seamless integration of custom or third-party schedulers with Mage workflows.
Describe alternatives you’ve considered
Alternatives considered include manually configuring the scheduler settings at the pod level, which can be cumbersome and error-prone, especially when managing large or dynamic environments. Another alternative is to fork and modify the Mage codebase to support this feature, but this approach is not scalable, introduces maintenance overhead, and creates potential compatibility issues with future updates.
Additional context
Allowing users to specify the scheduler pod name enhances flexibility, performance, and resource optimization. It also aligns with Kubernetes' philosophy of providing configurable components, giving users the ability to fine-tune their infrastructure based on their unique needs. This feature would be particularly beneficial in environments that use advanced schedulers like Volcano for machine learning workloads, Coscheduling for multi-step workflows, or custom schedulers designed for specific business logic. | closed | 2024-09-12T15:07:15Z | 2024-09-18T20:54:50Z | https://github.com/mage-ai/mage-ai/issues/5411 | [] | messerzen | 1 |
pyro-ppl/numpyro | numpy | 1,989 | Check transformation domain in `TransformedDistribution` constructor? | ### Feature Summary
Distributions know their support. Transformations know their domain and codomain. As far as I can tell, it should therefore be possible to confirm that a transformation used in constructing a `TransformedDistribution` from a base distribution has a domain equal to the support of the base distribution, and error if not.
### Why is this needed?
The lack of an error here can lead to unexpected and somewhat opaque behavior. See e.g. #1756
### Questions
- Is there reason to allow the creation of `TransformedDistributions` with a transformation whose domain is _not_ equal to the base distribution's support?
- Are there barriers to implementation that I'm not seeing?
Happy to take this on if it would be valuable. | open | 2025-02-27T17:35:16Z | 2025-03-01T19:23:20Z | https://github.com/pyro-ppl/numpyro/issues/1989 | [
"enhancement"
] | dylanhmorris | 2 |
pytorch/pytorch | python | 148,938 | [triton 3.3] `AOTInductorTestABICompatibleGpu.test_triton_kernel_tma_descriptor_1d_dynamic_False_cuda` | ### 🐛 Describe the bug
1. Update triton to `release/3.3.x` https://github.com/triton-lang/triton/tree/release/3.3.x
2. run `python test/inductor/test_aot_inductor.py -vvv -k test_triton_kernel_tma_descriptor_1d_dynamic_False_cuda`
Possibly an easier repro is
```
TORCHINDUCTOR_CPP_WRAPPER=1 python test/inductor/test_triton_kernels.py -k test_tma_descriptor_1d_dynamic_False_backend_inductor
```
errors:
<details>
```
/home/dberard/local/triton-env2/pytorch/torch/backends/cudnn/__init__.py:108: UserWarning: PyTorch was compiled without cuDNN/MIOpen support. To use cuDNN/MIOpen, rebuild PyTorch making sure the library is visible to the build system.
warnings.warn(
/home/dberard/local/triton-env2/pytorch/torch/backends/mkldnn/__init__.py:78: UserWarning: TF32 acceleration on top of oneDNN is available for Intel GPUs. The current Torch version does not have Intel GPU Support. (Triggered internally at /home/dberard/local/triton-env2/pytorch/aten/src/ATen/Context.cpp:148.)
torch._C._set_onednn_allow_tf32(_allow_tf32)
W0310 18:58:17.091000 2102274 torch/_export/__init__.py:67] +============================+
W0310 18:58:17.091000 2102274 torch/_export/__init__.py:68] | !!! WARNING !!! |
W0310 18:58:17.092000 2102274 torch/_export/__init__.py:69] +============================+
W0310 18:58:17.092000 2102274 torch/_export/__init__.py:70] torch._export.aot_compile()/torch._export.aot_load() is being deprecated, please switch to directly calling torch._inductor.aoti_compile_and_package(torch.export.export())/torch._inductor.aoti_load_package() instead.
ETEST SUITE EARLY TERMINATION due to torch.cuda.synchronize() failure
CUDA error: an illegal memory access was encountered
CUDA kernel errors might be asynchronously reported at some other API call, so the stacktrace below might be incorrect.
For debugging consider passing CUDA_LAUNCH_BLOCKING=1
Compile with `TORCH_USE_CUDA_DSA` to enable device-side assertions.
======================================================================
ERROR: test_triton_kernel_tma_descriptor_1d_dynamic_False_cuda (__main__.AOTInductorTestABICompatibleGpu)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/home/dberard/local/triton-env2/pytorch/torch/testing/_comparison.py", line 1221, in not_close_error_metas
pair.compare()
File "/home/dberard/local/triton-env2/pytorch/torch/testing/_comparison.py", line 700, in compare
self._compare_values(actual, expected)
File "/home/dberard/local/triton-env2/pytorch/torch/testing/_comparison.py", line 830, in _compare_values
compare_fn(
File "/home/dberard/local/triton-env2/pytorch/torch/testing/_comparison.py", line 1009, in _compare_regular_values_close
matches = torch.isclose(
RuntimeError: CUDA error: an illegal memory access was encountered
CUDA kernel errors might be asynchronously reported at some other API call, so the stacktrace below might be incorrect.
For debugging consider passing CUDA_LAUNCH_BLOCKING=1
Compile with `TORCH_USE_CUDA_DSA` to enable device-side assertions.
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "/home/dberard/local/triton-env2/pytorch/torch/testing/_internal/common_utils.py", line 3150, in wrapper
method(*args, **kwargs)
File "/home/dberard/local/triton-env2/pytorch/test/inductor/test_torchinductor.py", line 12836, in new_test
return value(self)
File "/home/dberard/local/triton-env2/pytorch/torch/testing/_internal/common_utils.py", line 552, in instantiated_test
test(self, **param_kwargs)
File "/home/dberard/local/triton-env2/pytorch/test/inductor/test_aot_inductor.py", line 2568, in test_triton_kernel_tma_descriptor_1d
self.check_model(
File "/home/dberard/local/triton-env2/pytorch/test/inductor/test_aot_inductor_utils.py", line 207, in check_model
self.assertEqual(actual, expected, atol=atol, rtol=rtol)
File "/home/dberard/local/triton-env2/pytorch/torch/testing/_internal/common_utils.py", line 4052, in assertEqual
error_metas = not_close_error_metas(
File "/home/dberard/local/triton-env2/pytorch/torch/testing/_comparison.py", line 1228, in not_close_error_metas
f"Comparing\n\n"
File "/home/dberard/local/triton-env2/pytorch/torch/testing/_comparison.py", line 367, in __repr__
body = [
File "/home/dberard/local/triton-env2/pytorch/torch/testing/_comparison.py", line 368, in <listcomp>
f" {name}={value!s},"
File "/home/dberard/local/triton-env2/pytorch/torch/_tensor.py", line 590, in __repr__
return torch._tensor_str._str(self, tensor_contents=tensor_contents)
File "/home/dberard/local/triton-env2/pytorch/torch/_tensor_str.py", line 710, in _str
return _str_intern(self, tensor_contents=tensor_contents)
File "/home/dberard/local/triton-env2/pytorch/torch/_tensor_str.py", line 631, in _str_intern
tensor_str = _tensor_str(self, indent)
File "/home/dberard/local/triton-env2/pytorch/torch/_tensor_str.py", line 363, in _tensor_str
formatter = _Formatter(get_summarized_data(self) if summarize else self)
File "/home/dberard/local/triton-env2/pytorch/torch/_tensor_str.py", line 146, in __init__
tensor_view, torch.isfinite(tensor_view) & tensor_view.ne(0)
RuntimeError: CUDA error: an illegal memory access was encountered
CUDA kernel errors might be asynchronously reported at some other API call, so the stacktrace below might be incorrect.
For debugging consider passing CUDA_LAUNCH_BLOCKING=1
Compile with `TORCH_USE_CUDA_DSA` to enable device-side assertions.
To execute this test, run the following from the base repo dir:
python test/inductor/test_aot_inductor.py AOTInductorTestABICompatibleGpu.test_triton_kernel_tma_descriptor_1d_dynamic_False_cuda
This message can be suppressed by setting PYTORCH_PRINT_REPRO_ON_FAILURE=0
----------------------------------------------------------------------
Ran 1 test in 5.612s
FAILED (errors=1)
inline_call []
unimplemented []
stats [('calls_captured', 2), ('unique_graphs', 1)]
inductor [('extern_calls', 4), ('async_compile_cache_miss', 2), ('benchmarking.InductorBenchmarker.benchmark_gpu', 2), ('pattern_matcher_count', 1), ('pattern_matcher_nodes', 1), ('async_compile_cache_hit', 1)]
graph_break []
aten_mm_info []
```
</details>
errors w/ compute-sanitizer:
https://gist.github.com/davidberard98/ecd9fefff91393b3a3fa0725dea96e22
### Versions
triton: release/3.3.x
pytorch: viable/strict from mar 10
cc @chauhang @penguinwu @voznesenskym @EikanWang @jgong5 @Guobing-Chen @XiaobingSuper @zhuhaozhe @blzheng @wenzhe-nrv @jiayisunx @ipiszy @chenyang78 @kadeng @muchulee8 @amjames @aakhundov @bertmaher @int3 @nmacchioni @embg @peterbell10 @avikchaudhuri @gmagogsfm @zhxchen17 @tugsbayasgalan @angelayi @suo @ydwu4 @desertfire @yushangdi @oulgen | open | 2025-03-11T02:00:45Z | 2025-03-13T16:58:07Z | https://github.com/pytorch/pytorch/issues/148938 | [
"oncall: pt2",
"module: inductor",
"upstream triton",
"oncall: export",
"module: aotinductor",
"module: user triton"
] | davidberard98 | 1 |
opengeos/streamlit-geospatial | streamlit | 54 | Page not working | I am trying to access to this page to create a timelapse but it seems that is not working. | closed | 2022-07-04T09:54:57Z | 2022-07-06T18:13:03Z | https://github.com/opengeos/streamlit-geospatial/issues/54 | [] | marparven1 | 1 |
widgetti/solara | fastapi | 246 | Cached variables? | I have a rather large app now, so I don't have any good isolated examples, but on multiple occasions I've navigated the app, perform changes that are saved to a database, then refreshed and seen old states of variables appearing. Then, I restart the development with `solara run main.py` and I get the actual (correct) state back again. Thus, I believe there is some caching mechanism that is causing problems. | open | 2023-08-18T18:11:58Z | 2023-09-16T08:34:54Z | https://github.com/widgetti/solara/issues/246 | [] | FerusAndBeyond | 3 |
lanpa/tensorboardX | numpy | 146 | histogramm error | pytorch 0.3.1.post2
TensorBoard 1.7.0
Tensorbord distributions consist of one value 3e+18.
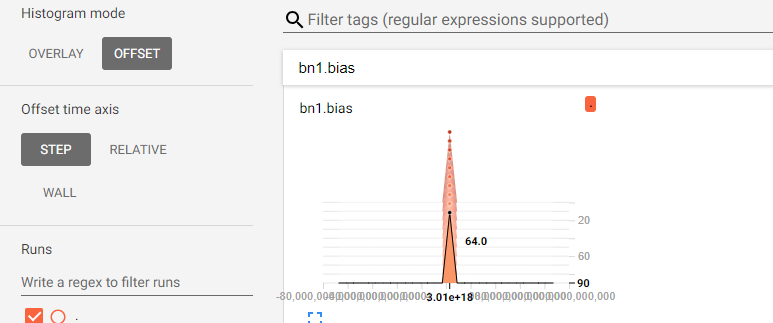
How can I fix it? | closed | 2018-05-24T15:34:19Z | 2018-05-24T17:48:01Z | https://github.com/lanpa/tensorboardX/issues/146 | [] | E1eMenta | 1 |
coqui-ai/TTS | python | 3,315 | [Feature request] Using local whisper transcription with word time stamps to remove tts hallucinations | **🚀 Feature Description**
Currently all the transformer based tts models I've run into deal with issues of hallucinations, especially at the end for instance even with XTTS V2, I was wondering if there was any planned way to remove at least the hallucinations that appear at the end of the generated audio, for instance the text could be "hey, bob" and the output audio will be "hey, bob. Other!"
**Solution**
You could use a solution like this where you have a cleanup method using this whisper repo: https://github.com/linto-ai/whisper-timestamped: to generate a transcription with the time stamps for each word in the generated output audio, to then compare that transcription-with-time-stamps to the words used in the input for the tts models
**Additional context**
I'm planning on trying to make and use a method like that for my own project, and was just wondering if anything like that was in the works. Thanks! | closed | 2023-11-27T02:46:22Z | 2024-01-28T22:40:19Z | https://github.com/coqui-ai/TTS/issues/3315 | [
"wontfix",
"feature request"
] | DrewThomasson | 2 |
keras-team/autokeras | tensorflow | 1,163 | Example code not working - MPG example | ### Bug Description
Trying to get started using AutoKeras and finding that most of the example code does not work.
### Bug Reproduction
Running the example here: https://autokeras.com/tutorial/structured_data_regression/
### Setup Details
Include the details about the versions of:
- OS type and version: MacOS Catalina 10.15.4
- Python: 3.8
- autokeras: master (pulled 20.06.01)
- keras-tuner: 1.0.1
- scikit-learn: 0.23.1
- numpy: 1.18.4
- pandas: 1.0.4
- tensorflow: 2.2.0
### Error
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
<ipython-input-12-3dda062a9e40> in <module>
5 # Evaluate the accuracy of the found model.
6 print('Accuracy: {accuracy}'.format(
----> 7 accuracy=regressor.evaluate(x=test_dataset.drop(columns=['MPG']), y=test_dataset['MPG'])))
~/opt/miniconda2/envs/AutoML/lib/python3.8/site-packages/autokeras-1.0.3-py3.8.egg/autokeras/tasks/structured_data.py in evaluate(self, x, y, batch_size, **kwargs)
133 if isinstance(x, str):
134 x, y = self._read_from_csv(x, y)
--> 135 return super().evaluate(x=x,
136 y=y,
137 batch_size=batch_size,
~/opt/miniconda2/envs/AutoML/lib/python3.8/site-packages/autokeras-1.0.3-py3.8.egg/autokeras/auto_model.py in evaluate(self, x, y, **kwargs)
443 """
444 dataset = self._process_xy(x, y, False)
--> 445 return self.tuner.get_best_model().evaluate(x=dataset, **kwargs)
446
447 def export_model(self):
~/opt/miniconda2/envs/AutoML/lib/python3.8/site-packages/autokeras-1.0.3-py3.8.egg/autokeras/engine/tuner.py in get_best_model(self)
43
44 def get_best_model(self):
---> 45 model = super().get_best_models()[0]
46 model.load_weights(self.best_model_path)
47 return model
~/opt/miniconda2/envs/AutoML/lib/python3.8/site-packages/kerastuner/engine/tuner.py in get_best_models(self, num_models)
229 """
230 # Method only exists in this class for the docstring override.
--> 231 return super(Tuner, self).get_best_models(num_models)
232
233 def _deepcopy_callbacks(self, callbacks):
~/opt/miniconda2/envs/AutoML/lib/python3.8/site-packages/kerastuner/engine/base_tuner.py in get_best_models(self, num_models)
236 """
237 best_trials = self.oracle.get_best_trials(num_models)
--> 238 models = [self.load_model(trial) for trial in best_trials]
239 return models
240
~/opt/miniconda2/envs/AutoML/lib/python3.8/site-packages/kerastuner/engine/base_tuner.py in <listcomp>(.0)
236 """
237 best_trials = self.oracle.get_best_trials(num_models)
--> 238 models = [self.load_model(trial) for trial in best_trials]
239 return models
240
~/opt/miniconda2/envs/AutoML/lib/python3.8/site-packages/kerastuner/engine/tuner.py in load_model(self, trial)
154 best_epoch = trial.best_step
155 with hm_module.maybe_distribute(self.distribution_strategy):
--> 156 model.load_weights(self._get_checkpoint_fname(
157 trial.trial_id, best_epoch))
158 return model
~/opt/miniconda2/envs/AutoML/lib/python3.8/site-packages/tensorflow/python/keras/engine/training.py in load_weights(self, filepath, by_name, skip_mismatch)
248 raise ValueError('Load weights is not yet supported with TPUStrategy '
249 'with steps_per_run greater than 1.')
--> 250 return super(Model, self).load_weights(filepath, by_name, skip_mismatch)
251
252 def compile(self,
~/opt/miniconda2/envs/AutoML/lib/python3.8/site-packages/tensorflow/python/keras/engine/network.py in load_weights(self, filepath, by_name, skip_mismatch)
1229 else:
1230 try:
-> 1231 py_checkpoint_reader.NewCheckpointReader(filepath)
1232 save_format = 'tf'
1233 except errors_impl.DataLossError:
~/opt/miniconda2/envs/AutoML/lib/python3.8/site-packages/tensorflow/python/training/py_checkpoint_reader.py in NewCheckpointReader(filepattern)
93 """
94 try:
---> 95 return CheckpointReader(compat.as_bytes(filepattern))
96 # TODO(b/143319754): Remove the RuntimeError casting logic once we resolve the
97 # issue with throwing python exceptions from C++.
ValueError: Unsuccessful TensorSliceReader constructor: Failed to get matching files on ./structured_data_regressor/trial_a2b0718070dcd1d815fe093a8ebb90ab/checkpoints/epoch_52/checkpoint: Not found: ./structured_data_regressor/trial_a2b0718070dcd1d815fe093a8ebb90ab/checkpoints/epoch_52; No such file or directory
| closed | 2020-06-02T15:50:26Z | 2023-03-27T09:19:30Z | https://github.com/keras-team/autokeras/issues/1163 | [
"bug report",
"pinned"
] | KirkDCO | 27 |
ScrapeGraphAI/Scrapegraph-ai | machine-learning | 415 | ImportError: cannot import name 'Required' from 'typing' (/usr/local/lib/python3.11/typing.py) | When I run this code
`from scrapegraphai.graphs import SmartScraperGraph
import nest_asyncio
graph_config = {
"llm": {
"model": "ollama/mistral",
"temperature": 0,
"format": "json", # Ollama needs the format to be specified explicitly
"base_url": "http://localhost:11434", # set Ollama URL
},
"embeddings": {
"model": "ollama/nomic-embed-text",
"base_url": "http://localhost:11434", # set Ollama URL
}
}
smart_scraper_graph = SmartScraperGraph(
prompt="List me all items",
# also accepts a string with the already downloaded HTML code
source="https://www.url.com",
config=graph_config
)
nest_asyncio.apply()
result = smart_scraper_graph.run()
print(result)
`
I got the following error

**VERSIONS**
Ubuntu - 20.04
Scrapegraphai - 1.7.4
| closed | 2024-06-28T07:23:39Z | 2024-08-18T08:43:15Z | https://github.com/ScrapeGraphAI/Scrapegraph-ai/issues/415 | [] | sunejay | 2 |
polarsource/polar | fastapi | 4,729 | Cannot mark issue is not solved but closed/reject funding | ### Description
<!-- A brief description with a link to the page on the site where you found the issue. -->
An issue was not solved but closed.
How do I mark that?
### Current Behavior
<!-- A brief description of the current behavior of the issue. -->
I can only mark the issue as done and pay it out.

### Expected Behavior
<!-- A brief description of what you expected to happen. -->
Nobody worked on this. How can I reject the funding?

### Screenshots
<!-- Add screenshots, if applicable, to help explain your problem. -->
### Environment:
- Operating System: [e.g., Windows, macOS, Linux] Linux
- Browser (if applicable): [e.g., Chrome, Firefox, Safari] Firefox
---
<!-- Thank you for contributing to Polar! We appreciate your help in improving it. -->
<!-- Questions: [Discord Server](https://discord.com/invite/Pnhfz3UThd). --> | closed | 2024-12-23T14:37:44Z | 2024-12-23T19:36:12Z | https://github.com/polarsource/polar/issues/4729 | [
"bug"
] | niccokunzmann | 1 |
pytorch/pytorch | machine-learning | 149,370 | UNSTABLE pull / cuda12.4-py3.10-gcc9-sm75 / test (pr_time_benchmarks) | See https://hud.pytorch.org/hud/pytorch/pytorch/main/1?per_page=50&name_filter=pr_time&mergeLF=true <- job passes and fails intermittently with no apparent commit that could have started it
cc @chauhang @penguinwu @seemethere @pytorch/pytorch-dev-infra | open | 2025-03-18T01:24:52Z | 2025-03-18T13:11:18Z | https://github.com/pytorch/pytorch/issues/149370 | [
"module: ci",
"triaged",
"oncall: pt2",
"unstable"
] | malfet | 2 |
twopirllc/pandas-ta | pandas | 699 | Schaff trend cycle (STC) giving negatively biased results: bug? | **Which version are you running? The lastest version is on Github. Pip is for major releases.**
pandas_ta: 0.3.14b0
**Do you have _TA Lib_ also installed in your environment?**
Yes: TA-Lib: 0.4.19
**Have you tried the _development_ version? Did it resolve the issue?**
No
**Describe the bug**
The values are always negatively biased, using ta.stc(x).iloc[:, 0].mean(). The indicator ranges from 0-100. I would expect a mean of 50. However, what I observe is always around 20. I tested ta.stc with different datasets and default parameters (or any other meaningful combination: I also tested changing each single parameter at a time).
For comparison, using the same data, the finta TA.STC(x).mean() consistently results in a mean of around 50, as would be expected.
**To Reproduce**
```python
import yfinance as yf
import pandas_ta as ta
from finta import TA
x = yf.download('ABB.NS', start = "2021-12-08", end = "2023-01-01", progress = False)
ta.stc(x['Close']).iloc[:, 0].mean()
TA.STC(x[['Open', 'Low', 'High', 'Close']]).mean()
```
```sh
pandas_ta result: 17.77649708406015
finta result: 47.476128567356724
```
**Expected behavior**
Value should be equal (or at least comparable) to finta result and around 50.
(also, stc calculation in finta seems a lot faster)
Thanks for developing Pandas TA! 😄 👍
| open | 2023-06-26T17:23:45Z | 2023-10-12T21:15:09Z | https://github.com/twopirllc/pandas-ta/issues/699 | [
"bug",
"help wanted"
] | halterc | 1 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.