repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
katanaml/sparrow | computer-vision | 29 | Random prediction and wrong prediction in repeated characters | Hello,
I have trained a donut base model on our custom dataset, which consists of a total of 12,480 images. I then fine-tuned this base model with default parameters.
During the analysis of predictions, I observed certain patterns in the JSON output. Specifically, when similar keys appear almost simultaneously, the model tends to make the following types of errors:
It predicts extra characters (e.g., "Paneer cheese paratha with butter" is predicted as "Paneer Paneer cheese paratha with butter").
It misses some characters (e.g., "199.00" is predicted as "19.00").
It predicts incorrect characters (e.g., "119.00" is predicted as "159.00").
Additionally, I noticed that the model often predicts characters such as "5," "7," and "1," even though these characters are not present in the images.
**Ground Truth:**
{
"table": [
{
"key": "Paneer paratha with butter",
"value": "199.00"
},
{
"key": "Paneer cheese paratha with butter",
"value": "119.00"
}
]
}
**Prediction:**
{
"table": [
{
"key": "Paneer paratha with butter",
"value": "19.00"
},
{
"key": "Paneer Paneer cheese paratha with butter",
"value": "159.00"
}
]
}
In the below json, model misses in between characters, predicts something else other than ground truth or gives extra characters in prediction which are not there in image/json. The image is clean enough for a model to get proper predictions still it gets wrong predictions as mentioned above.
As per analysis, the model makes more mistakes in values(Numeric) than keys(Alphabetic), maybe the reason is data imbalancing.
**Ground Truth:**
{
"table": [
{
"key": "Accessible Amount",
"value": "9123.23"
},
{
"key": "Car parts due :",
"value": "2,09,233.19"
},
{
"key": "Paint brushes :",
"value": "200.00"
}
]
}
**Predicted:**
{
"table": [
{
"key": "Accesible Amount",
"value": "9123.33"
},
{
"key": "Car parts due :",
"value": "9,1,233.19"
},
{
"key": "Paint brushes :",
"value": "200.000"
}
]
}
In the JSON provided below, despite the clarity of the image, the model consistently exhibits several issues:
Missing Characters: The model frequently fails to recognize certain characters.
Duplicate Keys: It tends to predict the same type of key multiple times, resulting in an extra key, such as "Oil fluid," which is a combination of two adjacent keys.
Missing Colon (:) at the End of Keys: The model omits the colon character at the end of keys.
Missing Plus Sign (+) in Values: It also overlooks the plus sign in values.
**Ground Truth :**
{
"table": [
{
"key": "Delivery charges :",
"value": "(+)470.00"
},
{
"key": "Oil charge:",
"value": "3,120.00"
},
{
"key": "Washer fluid :",
"value": "3,120.00"
}
]
}
**Predicted:**
{
"table": [
{
"key": "Delivery charges",
"value": "( )470.00"
},
{
"key": "Oil charge:",
"value": "3,120.00"
},
{
"key": "Oil fluid :",
"value": "157.00"
},
{
"key": "Washer fluid :",
"value": "3,120.00"
}
]
}
In the below json, I have found the same pattern that sometimes model predict a character only one time even after that character there two times in the image. like; (‘@ @’, ‘: :’) then the model will predict it only once. Also predicts the same keys twice.
**Ground Truth:**
{
"table": [
{
"key": "Transport charges::",
"value": "144.00"
},
{
"key": "Freight charges",
"value": ""
},
{
"key": "Washer fluid @ @ 18 %",
"value": "3,120.00"
}
]
}
**Prediction:**
{
"table": [
{
"key": "Transport charges:",
"value": "144.00"
},
{
"key": "Freight charges:",
"value": ""
},
{
"key": "Freight charges:",
"value": ""
},
{
"key": "Freight charges:",
"value": ""
},
{
"key": "Washer fluid @ 18 %",
"value": "3,120.00"
}
]
}
| closed | 2023-11-07T18:51:32Z | 2023-11-07T18:59:18Z | https://github.com/katanaml/sparrow/issues/29 | [] | Asha-12502 | 1 |
graphql-python/graphene | graphql | 870 | Resolver Not Receiving Arguments when Nested | I have a list of classes stored in memory that I am trying to parse through various types. It is referenced through the method `get_inventory()`.
When I call the classes individually, they resolve as I would expect.
But when I try to nest one in the other, the value is returning null.
The code, followed by some examples:
class Account(graphene.ObjectType):
account_name = graphene.String()
account_id = graphene.String()
def resolve_account(
self, info,
account_id=None,
account_name=None
):
inventory = get_inventory()
result = [Account(
account_id=i.account_id,
account_name=i.account_name
) for i in inventory if (
(i.account_id == account_id) or
(i.account_name == account_name)
)]
if len(result):
return result[0]
else:
return Account()
account = graphene.Field(
Account,
resolver=Account.resolve_account,
account_name=graphene.String(default_value=None),
account_id=graphene.String(default_value=None)
)
class Item(graphene.ObjectType):
item_name = graphene.String()
region = graphene.String()
account = account
def resolve_item(
self, info,
item_name=None
):
inventory = get_inventory()
result = [Item(
item_name=i.item_name,
region=i.region,
account=Account(
account_id=i.account_id
)
) for i in inventory if (
(i.item_name == item_name)
)]
if len(result):
return result[0]
else:
return Item()
item = graphene.Field(
Item,
resolver=Item.resolve_item,
item_name=graphene.String(default_value=None)
)
class Query(graphene.ObjectType):
account = account
item = item
schema = graphene.Schema(query=Query)
Let's assume I have an account `foo` that has an item `bar`. The below queries return the fields correctly.
{
account(accountName:"foo") {
accountName
accountId
}
}
{
item(itemName: "bar") {
itemName
region
}
}
So if I wanted to find the account that has the item `bar`, I would think I could query `bar` and get `foo`. But it returns the `account` fields as `null`.
{
item(itemName: "bar") {
itemName
region
account {
accountId
accountName
}
}
}
Recall that as part of `resolve_item`, I am doing `account=Account(account_id=i.account_id)` - I would expect this to work.
If I alter the last return statement of `resolve_account` to the below, `accountId` always returns `yo`.
...
else:
return Account(
account_id='yo'
)
So this tells me that my resolver is firing, but the invocation in `resolve_item` is not passing `account_id` properly.
Not sure if this is a bug or user error. | closed | 2018-11-26T18:56:26Z | 2019-08-05T21:18:01Z | https://github.com/graphql-python/graphene/issues/870 | [
"wontfix",
"👀 more info needed"
] | getglad | 5 |
mwouts/itables | jupyter | 328 | 2 Problems with SearchPanes | In this code , there are 2 problems :
1. 'Header of the column is not shown without a reset_index()
2. Only the first column appears in the searchPanes although 3 are asked.
Can someone help me please ? Thanks
I tried the most recent version and version 2.0 of itables
import itables
from itables.sample_dfs import get_countries
df = get_countries(html=False)
# df.reset_index(inplace=True) with reset_index header 'region' is shown , without reset header is not shown
show(
df,
layout={"top1": "searchPanes"},
searchPanes={"layout": "columns-3", "cascadePanes": True, "columns": [1, 2, 3]})
#only the first column is shown in the Panes | closed | 2024-10-21T11:56:10Z | 2024-11-02T22:38:50Z | https://github.com/mwouts/itables/issues/328 | [] | tomnobelsAM | 2 |
sgl-project/sglang | pytorch | 3,802 | [Feature] SGlang_router start with healthy given worker_urls | ### Checklist
- [x] 1. If the issue you raised is not a feature but a question, please raise a discussion at https://github.com/sgl-project/sglang/discussions/new/choose Otherwise, it will be closed.
- [x] 2. Please use English, otherwise it will be closed.
### Motivation
the router start fails if not all the given worker_url are healthy, this may cause some problems when one worker is broken. Besides, if the router can start with no worker_url, this problem can also be solved by adding the heathy workers only.
### Related resources
_No response_ | open | 2025-02-24T03:36:18Z | 2025-03-20T09:31:30Z | https://github.com/sgl-project/sglang/issues/3802 | [] | slr1997 | 2 |
seleniumbase/SeleniumBase | web-scraping | 3,284 | dublicate url when using sb.open in cycle | https://www.sofascore.comhttps//www.sofascore.com/ru/football/match/pharco-fc-enppi/owrsdJWb#id:13015449 getting like this when using
def get_games_info(self, links):
games_info = []
for link in links:
self.sb.open(link)
time.sleep(1) | closed | 2024-11-22T17:09:30Z | 2024-11-22T17:55:42Z | https://github.com/seleniumbase/SeleniumBase/issues/3284 | [
"invalid"
] | SenseiSol | 1 |
matplotlib/cheatsheets | matplotlib | 16 | Image for "Basic plots - plot" should hint at markers | Currently the image is just sine line, which could trick users in thinking that `plot` is for lines and `scatter` is for markers.

I propose to additionally show another set of values with markers, e.g. something like:

| open | 2020-07-06T20:42:10Z | 2020-07-07T20:26:42Z | https://github.com/matplotlib/cheatsheets/issues/16 | [] | timhoffm | 9 |
sqlalchemy/sqlalchemy | sqlalchemy | 11,282 | Unable to close a connection | ### Describe the bug
when using the method .close() in my code (on **sqlalchemy.engine.base.Connection** object), and then checking the connections in the container, I still see the connection.
<img width="641" alt="image" src="https://github.com/sqlalchemy/sqlalchemy/assets/102469772/ecd2d3e7-0e8d-4a21-b3a3-092198720d0b">
Is there a problem with this method?
### Optional link from https://docs.sqlalchemy.org which documents the behavior that is expected
_No response_
### SQLAlchemy Version in Use
20.0.29
### DBAPI (i.e. the database driver)
pyodbc
### Database Vendor and Major Version
Teradata
### Python Version
3.10
### Operating system
Linux
### To Reproduce
```python
# sqlalchemy.engine.base.Connection - Object
client.connection.close()
# after closing still see the connection in the container
```
### Error
There is no error
### Additional context
_No response_ | closed | 2024-04-17T13:09:27Z | 2024-04-17T17:16:22Z | https://github.com/sqlalchemy/sqlalchemy/issues/11282 | [] | rshunim | 0 |
Neoteroi/BlackSheep | asyncio | 218 | docs: Error in `dispose_http_client` | **Describe the bug**
A description of what the bug is, possibly including how to reproduce it.
How it should be:
```diff
async def dispose_http_client(app):
- http_client = app.services.get(ClientSession) # obtain the http client
+ http_client = app.service_provider.get(ClientSession) # obtain the http client
await http_client.close()
``` | closed | 2021-12-15T14:45:11Z | 2021-12-15T18:50:58Z | https://github.com/Neoteroi/BlackSheep/issues/218 | [] | q0w | 2 |
mckinsey/vizro | plotly | 566 | How to make an editable table/Ag Grid be the source for a chart/figure | Hello all 👋 ,
based on a recent user question I wanted to show how one can make an editable table power a graph. The original request was:
> I would like to create a Table in my dashboard (in default blank) that allows the user to input values (the index are dates). Once clicking on a "Save" button, the data is used to create a line-chart on the left.
> How should I realize this with the AgGrid model and custimizable actions?
Here is the solution (I made the update automatic - but of course another "Save" button could be added!)

The code for this can be found here, where you can also run the dashboard live: https://py.cafe/maxi.schulz/vizro-user-question-editable-table
#### Notes
- this example showcases how Vizro is a framework on top of dash - and you can easily revert to using pure Dash for custom functionality
- some parts done here with `@callback` could be done with `vm.Action`, but not all
#### Important
When creating dashboards that allow for user input that the dashboard creator is always responsible for the security of the dashboard. Never evaluate untrusted data.
#### Sources
This solution was heavily inspired by https://www.youtube.com/watch?v=LNQhY8NZmCY (thanks @Coding-with-Adam )
#### Discussion
@petar-qb @antonymilne Maybe take note of this issue in case you have any comments, I find this especially interesting because it highlights the need for:
- specifying triggers
- ideally not returning entire figs in callbacks/actions, but rather modifying data in the DM (which we do not do in this example)
| open | 2024-07-04T12:44:10Z | 2024-07-08T15:03:27Z | https://github.com/mckinsey/vizro/issues/566 | [
"General Question :question:"
] | maxschulz-COL | 0 |
dgtlmoon/changedetection.io | web-scraping | 2,935 | Resetting of watch group tab selection when interacting with UI. | **Describe the bug**
Resetting of watch group tab selection when interacting with UI.
**Version**
0.49.0
**How did you install?**
pip
**To Reproduce**
You already fixed couple of UI actions reseting tab selection here https://github.com/dgtlmoon/changedetection.io/issues/2785 (big thank you for that!!!)
There are couple more:
-, watch edit > "clear history"
-, click on "CHANGEDETECTION.io in logo area" (i use it often when i want to go (from watch edit) to home, aka "watch list" view page)
-, watch checkbox > ''unpause"
...
= in general in my opinion it's better to make tab selection "persistent" / changeable only by clicking watch group "tab", cause if user focused his attention on particular group (narrowed it down) resetting his choice is not friendly.
but all this of course if you agree and if your time permits.
| open | 2025-01-29T01:01:13Z | 2025-01-30T14:54:53Z | https://github.com/dgtlmoon/changedetection.io/issues/2935 | [
"user-interface",
"triage"
] | gety9 | 3 |
littlecodersh/ItChat | api | 973 | 微信机器人 | open | 2022-12-10T17:19:48Z | 2022-12-10T17:19:48Z | https://github.com/littlecodersh/ItChat/issues/973 | [] | oucos | 0 | |
xonsh/xonsh | data-science | 5,623 | [docs] Cli command examples in guides should be fully copy&pasteable, i.e., without @ | ## Current Behavior
You can't copy&paste a line from a guide and run it, but need to cleanup `@` symbol
For example
> We suggest using the branchname::
`@ cp TEMPLATE.rst branch.rst`
When triple-clicking it on https://xon.sh/devguide.html you get `@`
Interestingly enough, the front page https://xon.sh seems to be better and doesn't include `@`
## Expected Behavior
No `@` on copying a line
## For community
⬇️ **Please click the 👍 reaction instead of leaving a `+1` or 👍 comment**
| open | 2024-07-23T04:57:22Z | 2024-07-23T11:25:46Z | https://github.com/xonsh/xonsh/issues/5623 | [
"docs"
] | eugenesvk | 0 |
microsoft/nni | tensorflow | 5,720 | NNI is starting, it's time to run an epoch but there's no value in the page? | **Describe the issue**:
it's time to run an epoch but there's no value in the page?
**Environment**:
- NNI version:2.5
- Training service (local|remote|pai|aml|etc):local
- Client OS:Win10
- Server OS (for remote mode only):
- Python version: 3.7
- PyTorch/TensorFlow version:PyTorch
- Is conda/virtualenv/venv used?:conda
- Is running in Docker?: no
**Configuration**:
searchSpaceFile: search_space.json
trialCommand: python train_nni.py
trialGpuNumber: 0
trialConcurrency: 1
tuner:
name: TPE
classArgs:
optimize_mode: maximize
trainingService:
platform: local

**How to reproduce it?**: | open | 2023-12-10T11:22:42Z | 2023-12-10T11:22:42Z | https://github.com/microsoft/nni/issues/5720 | [] | yao-ao | 0 |
jmcnamara/XlsxWriter | pandas | 313 | Minor gridlines in scatter chart do not support log_base | I am trying to set Y in log base 10 with minor gridlines. If I add minor gridlines the log_base category do not work anymore:
```
chart.set_y_axis({
'minor_gridlines': {
'visible': True,
'line': {'width': 1.25, 'dash_type': 'solid'},
'log_base' : 10
},
})
chart.set_y_axis({'log_base': 10})
```
Using second line removes minor gridline, opposite sequence removed log scale.
| closed | 2015-11-10T07:52:36Z | 2015-11-17T08:04:44Z | https://github.com/jmcnamara/XlsxWriter/issues/313 | [
"question",
"ready to close"
] | MichalMisiaszek | 2 |
igorbenav/fastcrud | pydantic | 209 | [Bug / Skill Issue] many to many relationship | **Describe the bug or question**
I would like to have a class for the Assosciate table that references the ids, from 2 other tables as Foreign Keys and make the records in there to be deleted when one of the referred objects gets deleted (cascade).
**To Reproduce**
Please provide a self-contained, minimal, and reproducible example of your use case
```python
from sqlalchemy.orm import Mapped, mapped_column
from sqlalchemy import String, ForeignKey, UniqueConstraint
from src.schemas.level import LanguageEnum
from src.setup.database import Base
class Tag(Base):
__tablename__ = "tag"
id: Mapped[int] = mapped_column(autoincrement=True, primary_key=True)
name: Mapped[str]
language: Mapped[LanguageEnum] = mapped_column(String)
user_id: Mapped[int] = mapped_column(ForeignKey("user.id"), index=True)
__table_args__ = (UniqueConstraint("name", "language", name="uq_tag_language"),)
class PhraseTag(Base):
__tablename__ = "phrase_tag"
tag_id: Mapped[int] = mapped_column(ForeignKey("tag.id", ondelete="CASCADE"), primary_key=True)
phrase_id: Mapped[int] = mapped_column(ForeignKey("phrase.id", ondelete="CASCADE"), primary_key=True)
```
```python
from sqlalchemy import String, ForeignKey
from sqlalchemy.orm import Mapped, mapped_column
from src.setup.database import Base
from src.models.level import LanguageEnum
class Phrase(Base):
__tablename__ = "phrase"
id: Mapped[int] = mapped_column(autoincrement=True, primary_key=True)
language: Mapped[LanguageEnum] = mapped_column(String)
phrase: Mapped[str]
user_id: Mapped[int] = mapped_column(ForeignKey("user.id"))
```
**Description**
Expected: I expect that whenever the phrase or tag gets deleted then all the objects that references the id of one of them gets deleted as well.
Actual: The PhraseTag table still has all the records present when I delete either Phrase or Tag
**Screenshots**
N/A
**Additional context**
N/A
| closed | 2025-03-18T20:34:45Z | 2025-03-18T22:45:46Z | https://github.com/igorbenav/fastcrud/issues/209 | [] | maktowon | 1 |
dynaconf/dynaconf | flask | 1,075 | [CI] Update codecov configuration file | I've closed #990 but forget to create an issue about the codecov configuration issue.
Apparently, we should have a `coverage.yml` file for the Codecov app/github-action, but I'm not sure how this will go with local coverage reports, which seems to use `.coveragerc`. This require a little investigation.
The goal here is to have:
- a single config file (local reports and CI)
- an up-to-date configuration file (as recommended in codecov docs):
- as a consequence, we can more easily customize the codecov config (if we discuss we need to) | open | 2024-03-06T12:42:47Z | 2024-03-06T12:42:48Z | https://github.com/dynaconf/dynaconf/issues/1075 | [
"enhancement",
"CI"
] | pedro-psb | 0 |
noirbizarre/flask-restplus | flask | 54 | No way to configure .../swagger.json to go over HTTPS? | Hello,
I have an application deployed using Flask-RESTPlus. Everything works nicely when it goes over http, but soon as I switch to https, it can't load the swagger.json because it is making requests to http://host/swagger.json and not https://host/swagger.json
The endpoints are registered properly with blueprints, because if I manually go to https://host/swagger.json, I get the correct JSON. However, the generated swagger page tries to load the JSON from a http endpoint, instead of the correct https endpoint.
Is there any way to fix this?
Thanks
| closed | 2015-06-25T17:54:24Z | 2022-01-05T01:52:28Z | https://github.com/noirbizarre/flask-restplus/issues/54 | [] | Iulian7 | 11 |
zappa/Zappa | django | 526 | [Migrated] Deployed flask endpoint not working: [run_wsgi_app TypeError: 'NoneType' object is not callable] App function file missing in Lamda zip package | Originally from: https://github.com/Miserlou/Zappa/issues/1396 by [NakedKoala](https://github.com/NakedKoala)
## Context
I am trying to deploy my deep learning model to lambda. The entire deployment package is over 1 GB. I am using slim-handler.
The app function file looks like this:
```
import logging
from flask import Flask
from flask import request
from flask import json
from model import TacoModel
from utils import image
from keras.preprocessing.image import img_to_array
import numpy as np
app = Flask(__name__)
logging.basicConfig()
logger = logging.getLogger(__name__)
logger.setLevel(logging.DEBUG)
taco_model = TacoModel()
@app.route('/', methods=['GET'])
def index():
path = "data/"
img = image.load_img(path+"test/unknown/hotdog679.JPEG",target_size=(224, 224))
img = np.array([img_to_array(img)])
res = taco_model.predict(img)
return json.dumps(res)
if __name__ == '__main__':
app.run()
```
Everything runs fine locally. When I curl my local server, I am able to get back the expected response.
The zappa deployment was successful and without error. But get request produces
```
{u'message': u'An uncaught exception happened while servicing this request. You can investigate this with the `zappa tail` command.', u'traceback': ['Traceback (most recent call last):\\n', ' File \"/var/task/handler.py\", line 452, in handler\\n response = Response.from_app(self.wsgi_app, environ)\\n', ' File \"/private/var/folders/kl/35lz1q3x2gs0y0r_1njxfd0m0000gn/T/pip-build-R7GWCg/Werkzeug/werkzeug/wrappers.py\", line 903, in from_app\\n', ' File \"/private/var/folders/kl/35lz1q3x2gs0y0r_1njxfd0m0000gn/T/pip-build-R7GWCg/Werkzeug/werkzeug/wrappers.py\", line 57, in _run_wsgi_app\\n', ' File \"/private/var/folders/kl/35lz1q3x2gs0y0r_1njxfd0m0000gn/T/pip-build-R7GWCg/Werkzeug/werkzeug/test.py\", line 884, in run_wsgi_app\\n', \"TypeError: 'NoneType' object is not callable\\n\"]}
```
## Expected Behavior
Endpoint should work and return a dummy prediction on get request
## Actual Behavior
Get request failed. Produce the above error
## Possible Fix
I have tried clean up my virtualenv & recreated virtualenv and re-deploy. Error still persists
zappa tail reveals this error
```
[1518497133157] No module named my_app: ImportError
Traceback (most recent call last):
File "/var/task/handler.py", line 509, in lambda_handler
return LambdaHandler.lambda_handler(event, context)
File "/var/task/handler.py", line 237, in lambda_handler
handler = cls()
File "/var/task/handler.py", line 129, in __init__
self.app_module = importlib.import_module(self.settings.APP_MODULE)
File "/usr/lib64/python2.7/importlib/__init__.py", line 37, in import_module
__import__(name)
ImportError: No module named my_app
```
I tried exporting my lambda zip package. Upon inspection , my flask app function file is missing in the deployment package. I am new to Zappa, but this looks wrong to me. So I manually added the app function file and re-zip & upload. But the get request still broke and zappa tail returns some different error.
Is something wrong during the packaging step ?
## Steps to Reproduce
live link
https://knbxmrhkt0.execute-api.us-west-2.amazonaws.com/dev
## Your Environment
* Zappa version used: 0.45.1
* Operating System and Python version: MacOS python 2.7
* The output of `pip freeze`:
```
absl-py==0.1.10
appnope==0.1.0
argcomplete==1.9.2
backports-abc==0.5
backports.functools-lru-cache==1.5
backports.shutil-get-terminal-size==1.0.0
backports.weakref==1.0.post1
base58==0.2.4
bcolz==1.1.2
bleach==2.1.2
boto3==1.5.27
botocore==1.8.41
certifi==2018.1.18
cfn-flip==1.0.0
chardet==3.0.4
click==6.7
configparser==3.5.0
cycler==0.10.0
decorator==4.2.1
docutils==0.14
durationpy==0.5
entrypoints==0.2.3
enum34==1.1.6
Flask==0.12.2
funcsigs==1.0.2
functools32==3.2.3.post2
future==0.16.0
futures==3.1.1
h5py==2.7.1
hjson==3.0.1
html5lib==0.9999999
idna==2.6
ipykernel==4.8.1
ipython==5.5.0
ipython-genutils==0.2.0
ipywidgets==7.1.1
itsdangerous==0.24
Jinja2==2.10
jmespath==0.9.3
jsonschema==2.6.0
jupyter==1.0.0
jupyter-client==5.2.2
jupyter-console==5.2.0
jupyter-core==4.4.0
kappa==0.6.0
Keras==1.2.2
lambda-packages==0.19.0
Markdown==2.6.11
MarkupSafe==1.0
matplotlib==2.1.2
mistune==0.8.3
mock==2.0.0
mpmath==1.0.0
nbconvert==5.3.1
nbformat==4.4.0
notebook==5.4.0
numpy==1.14.0
pandas==0.22.0
pandocfilters==1.4.2
pathlib2==2.3.0
pbr==3.1.1
pexpect==4.4.0
pickleshare==0.7.4
Pillow==5.0.0
placebo==0.8.1
prompt-toolkit==1.0.15
protobuf==3.5.1
ptyprocess==0.5.2
Pygments==2.2.0
pyparsing==2.2.0
python-dateutil==2.6.1
python-slugify==1.2.4
pytz==2018.3
PyYAML==3.12
pyzmq==17.0.0
qtconsole==4.3.1
requests==2.18.4
s3transfer==0.1.12
scandir==1.7
scikit-learn==0.19.1
scipy==1.0.0
Send2Trash==1.4.2
simplegeneric==0.8.1
singledispatch==3.4.0.3
six==1.11.0
subprocess32==3.2.7
sympy==1.1.1
tensorflow==1.5.0
tensorflow-tensorboard==1.5.1
terminado==0.8.1
testpath==0.3.1
Theano==1.0.1
toml==0.9.4
tornado==4.5.3
tqdm==4.19.1
traitlets==4.3.2
troposphere==2.2.0
Unidecode==1.0.22
urllib3==1.22
wcwidth==0.1.7
Werkzeug==0.12
widgetsnbextension==3.1.3
wsgi-request-logger==0.4.6
zappa==0.45.1
```
* Link to your project (optional):
* Your `zappa_settings.py`:
```
{
"dev": {
"app_function": "my_app.app",
"aws_region": "us-west-2",
"profile_name": "default",
"project_name": "freshstart3",
"runtime": "python2.7",
"s3_bucket": "[redacted]",
"slim_handler": true
}
}
``` | closed | 2021-02-20T09:43:57Z | 2022-07-16T07:15:31Z | https://github.com/zappa/Zappa/issues/526 | [] | jneves | 1 |
coleifer/sqlite-web | flask | 24 | UnicodeDecodeError: 'ascii' codec can't decode byte 0xff in position 0: ordinal not in range(128) | classic unicode issue, please fix, thanks
| closed | 2016-09-13T09:58:41Z | 2016-09-17T17:45:05Z | https://github.com/coleifer/sqlite-web/issues/24 | [] | hugowan | 7 |
vllm-project/vllm | pytorch | 15,217 | [Bug]: RuntimeError: please ensure that world_size (2) is less than than max local gpu count (1) | ### Your current environment
```text
Collecting environment information...
PyTorch version: 2.7.0a0+git6c0e746
Is debug build: False
CUDA used to build PyTorch: N/A
ROCM used to build PyTorch: 6.3.42133-1b9c17779
OS: Ubuntu 22.04.5 LTS (x86_64)
GCC version: (Ubuntu 11.4.0-1ubuntu1~22.04) 11.4.0
Clang version: 18.0.0git (https://github.com/RadeonOpenCompute/llvm-project roc-6.3.1 24491 1e0fda770a2079fbd71e4b70974d74f62fd3af10)
CMake version: version 3.31.4
Libc version: glibc-2.35
Python version: 3.12.9 (main, Feb 5 2025, 08:49:00) [GCC 11.4.0] (64-bit runtime)
Python platform: Linux-5.14.21-150500.55.83_13.0.62-cray_shasta_c-x86_64-with-glibc2.35
Is CUDA available: True
CUDA runtime version: Could not collect
CUDA_MODULE_LOADING set to: LAZY
GPU models and configuration: AMD Instinct MI250X (gfx90a:sramecc+:xnack-)
Nvidia driver version: Could not collect
cuDNN version: Could not collect
HIP runtime version: 6.3.42133
MIOpen runtime version: 3.3.0
Is XNNPACK available: True
CPU:
Architecture: x86_64
CPU op-mode(s): 32-bit, 64-bit
Address sizes: 48 bits physical, 48 bits virtual
Byte Order: Little Endian
CPU(s): 128
On-line CPU(s) list: 0-127
Vendor ID: AuthenticAMD
Model name: AMD EPYC 7A53 64-Core Processor
CPU family: 25
Model: 48
Thread(s) per core: 2
Core(s) per socket: 64
Socket(s): 1
Stepping: 1
Frequency boost: enabled
CPU max MHz: 3541.0149
CPU min MHz: 1500.0000
BogoMIPS: 3992.57
Flags: fpu vme de pse tsc msr pae mce cx8 apic sep mtrr pge mca cmov pat pse36 clflush mmx fxsr sse sse2 ht syscall nx mmxext fxsr_opt pdpe1gb rdtscp lm constant_tsc rep_good nopl nonstop_tsc cpuid extd_apicid aperfmperf rapl pni pclmulqdq monitor ssse3 fma cx16 pcid sse4_1 sse4_2 x2apic movbe popcnt aes xsave avx f16c rdrand lahf_lm cmp_legacy svm extapic cr8_legacy abm sse4a misalignsse 3dnowprefetch osvw ibs wdt tce topoext perfctr_core perfctr_nb bpext perfctr_llc mwaitx cpb cat_l3 cdp_l3 invpcid_single hw_pstate ssbd mba ibrs ibpb stibp vmmcall fsgsbase bmi1 avx2 smep bmi2 erms invpcid cqm rdt_a rdseed adx smap clflushopt clwb sha_ni xsaveopt xsavec xgetbv1 xsaves cqm_llc cqm_occup_llc cqm_mbm_total cqm_mbm_local clzero irperf xsaveerptr rdpru wbnoinvd amd_ppin arat npt lbrv svm_lock nrip_save tsc_scale vmcb_clean flushbyasid decodeassists pausefilter pfthreshold avic v_vmsave_vmload vgif v_spec_ctrl umip pku ospke vaes vpclmulqdq rdpid overflow_recov succor smca fsrm
Virtualization: AMD-V
L1d cache: 2 MiB (64 instances)
L1i cache: 2 MiB (64 instances)
L2 cache: 32 MiB (64 instances)
L3 cache: 256 MiB (8 instances)
NUMA node(s): 4
NUMA node0 CPU(s): 0-15,64-79
NUMA node1 CPU(s): 16-31,80-95
NUMA node2 CPU(s): 32-47,96-111
NUMA node3 CPU(s): 48-63,112-127
Vulnerability Gather data sampling: Not affected
Vulnerability Itlb multihit: Not affected
Vulnerability L1tf: Not affected
Vulnerability Mds: Not affected
Vulnerability Meltdown: Not affected
Vulnerability Mmio stale data: Not affected
Vulnerability Reg file data sampling: Not affected
Vulnerability Retbleed: Not affected
Vulnerability Spec rstack overflow: Mitigation; Safe RET
Vulnerability Spec store bypass: Mitigation; Speculative Store Bypass disabled via prctl and seccomp
Vulnerability Spectre v1: Mitigation; usercopy/swapgs barriers and __user pointer sanitization
Vulnerability Spectre v2: Mitigation; Retpolines; IBPB conditional; IBRS_FW; STIBP always-on; RSB filling; PBRSB-eIBRS Not affected, BHI Not affected
Vulnerability Srbds: Not affected
Vulnerability Tsx async abort: Not affected
Versions of relevant libraries:
[pip3] numpy==1.26.4
[pip3] pyzmq==26.2.1
[pip3] torch==2.7.0a0+git6c0e746
[pip3] torchvision==0.21.0+7af6987
[pip3] transformers==4.49.0
[pip3] triton==3.2.0+gite5be006a
[conda] Could not collect
ROCM Version: 6.3.42133-1b9c17779
Neuron SDK Version: N/A
vLLM Version: 0.7.4.dev49+gc0dd5adf6
vLLM Build Flags:
CUDA Archs: Not Set; ROCm: Disabled; Neuron: Disabled
GPU Topology:
============================ ROCm System Management Interface ============================
================================ Weight between two GPUs =================================
GPU0
GPU0 0
================================= Hops between two GPUs ==================================
GPU0
GPU0 0
=============================== Link Type between two GPUs ===============================
GPU0
GPU0 0
======================================= Numa Nodes =======================================
GPU[0] : (Topology) Numa Node: 0
GPU[0] : (Topology) Numa Affinity: 0
================================== End of ROCm SMI Log ===================================
PYTORCH_ROCM_ARCH=gfx90a;gfx942
LD_LIBRARY_PATH=/usr/local/lib/python3.12/dist-packages/cv2/../../lib64::/opt/cray/pe/mpich/default/ofi/gnu/9.1/lib-abi-mpich:/opt/cray/pe/mpich/default/gtl/lib:/opt/cray/xpmem/default/lib64:/opt/cray/pe/pmi/default/lib:/opt/cray/pe/pals/default/lib:/opt/cray/pe/gcc-libs:/opt/rocm/lib:/usr/local/lib::/.singularity.d/libs:/.singularity.d/libs
NCCL_CUMEM_ENABLE=0
TORCHINDUCTOR_COMPILE_THREADS=1
CUDA_MODULE_LOADING=LAZY
```
### 🐛 Describe the bug
I want to run vllm on 2 nodes. I already setup the ray, and it showed 2 GPUs:
```
Singularity> ray status
======== Autoscaler status: 2025-03-20 20:40:48.252496 ========
Node status
---------------------------------------------------------------
Active:
1 node_5373538893976cc08e332acdf25b3f043ca49757d98d5f6ab048d1ed
1 node_1689389e001b234e7d470b2da54b6c17b4441403358cf341f778e2ea
Pending:
(no pending nodes)
Recent failures:
(no failures)
Resources
---------------------------------------------------------------
Usage:
0.0/256.0 CPU
0.0/2.0 GPU
0B/311.15GiB memory
0B/137.34GiB object_store_memory
Demands:
(no resource demands)
```
Then I run `vllm serve ./DeepSeek-V3 --trust-remote-code --tensor-parallel-size 1 --pipeline-parallel-size 2` on the head node, it showed:
```
INFO 03-20 20:38:08 [__init__.py:207] Automatically detected platform rocm.
INFO 03-20 20:38:09 [api_server.py:911] vLLM API server version 0.7.4.dev49+gc0dd5adf6
INFO 03-20 20:38:09 [api_server.py:912] args: Namespace(subparser='serve', model_tag='./DeepSeek-V3', config='', host=None, port=8000, uvicorn_log_level='info', allow_credentials=False, allowed_origins=['*'], allowed_methods=['*'], allowed_headers=['*'], api_key=None, lora_modules=None, prompt_adapters=None, chat_template=None, chat_template_content_format='auto', response_role='assistant', ssl_keyfile=None, ssl_certfile=None, ssl_ca_certs=None, enable_ssl_refresh=False, ssl_cert_reqs=0, root_path=None, middleware=[], return_tokens_as_token_ids=False, disable_frontend_multiprocessing=False, enable_request_id_headers=False, enable_auto_tool_choice=False, enable_reasoning=False, reasoning_parser=None, tool_call_parser=None, tool_parser_plugin='', model='./DeepSeek-V3', task='auto', tokenizer=None, skip_tokenizer_init=False, revision=None, code_revision=None, tokenizer_revision=None, tokenizer_mode='auto', trust_remote_code=True, allowed_local_media_path=None, download_dir=None, load_format='auto', config_format=<ConfigFormat.AUTO: 'auto'>, dtype='auto', kv_cache_dtype='auto', max_model_len=None, guided_decoding_backend='xgrammar', logits_processor_pattern=None, model_impl='auto', distributed_executor_backend=None, pipeline_parallel_size=2, tensor_parallel_size=1, max_parallel_loading_workers=None, ray_workers_use_nsight=False, block_size=None, enable_prefix_caching=None, disable_sliding_window=False, use_v2_block_manager=True, num_lookahead_slots=0, seed=0, swap_space=4, cpu_offload_gb=0, gpu_memory_utilization=0.9, num_gpu_blocks_override=None, max_num_batched_tokens=None, max_num_partial_prefills=1, max_long_partial_prefills=1, long_prefill_token_threshold=0, max_num_seqs=None, max_logprobs=20, disable_log_stats=False, quantization=None, rope_scaling=None, rope_theta=None, hf_overrides=None, enforce_eager=False, max_seq_len_to_capture=8192, disable_custom_all_reduce=False, tokenizer_pool_size=0, tokenizer_pool_type='ray', tokenizer_pool_extra_config=None, limit_mm_per_prompt=None, mm_processor_kwargs=None, disable_mm_preprocessor_cache=False, enable_lora=False, enable_lora_bias=False, max_loras=1, max_lora_rank=16, lora_extra_vocab_size=256, lora_dtype='auto', long_lora_scaling_factors=None, max_cpu_loras=None, fully_sharded_loras=False, enable_prompt_adapter=False, max_prompt_adapters=1, max_prompt_adapter_token=0, device='auto', num_scheduler_steps=1, multi_step_stream_outputs=True, scheduler_delay_factor=0.0, enable_chunked_prefill=None, speculative_model=None, speculative_model_quantization=None, num_speculative_tokens=None, speculative_disable_mqa_scorer=False, speculative_draft_tensor_parallel_size=None, speculative_max_model_len=None, speculative_disable_by_batch_size=None, ngram_prompt_lookup_max=None, ngram_prompt_lookup_min=None, spec_decoding_acceptance_method='rejection_sampler', typical_acceptance_sampler_posterior_threshold=None, typical_acceptance_sampler_posterior_alpha=None, disable_logprobs_during_spec_decoding=None, model_loader_extra_config=None, ignore_patterns=[], preemption_mode=None, served_model_name=None, qlora_adapter_name_or_path=None, show_hidden_metrics_for_version=None, otlp_traces_endpoint=None, collect_detailed_traces=None, disable_async_output_proc=False, scheduling_policy='fcfs', scheduler_cls='vllm.core.scheduler.Scheduler', override_neuron_config=None, override_pooler_config=None, compilation_config=None, kv_transfer_config=None, worker_cls='auto', generation_config=None, override_generation_config=None, enable_sleep_mode=False, calculate_kv_scales=False, additional_config=None, disable_log_requests=False, max_log_len=None, disable_fastapi_docs=False, enable_prompt_tokens_details=False, dispatch_function=<function ServeSubcommand.cmd at 0x14a39b61c180>)
INFO 03-20 20:38:09 [config.py:209] Replacing legacy 'type' key with 'rope_type'
INFO 03-20 20:38:24 [config.py:570] This model supports multiple tasks: {'score', 'generate', 'reward', 'embed', 'classify'}. Defaulting to 'generate'.
INFO 03-20 20:38:31 [config.py:1479] Defaulting to use mp for distributed inference
INFO 03-20 20:38:31 [config.py:1536] Disabled the custom all-reduce kernel because it is not supported with pipeline parallelism.
WARNING 03-20 20:38:31 [arg_utils.py:1218] The model has a long context length (163840). This may cause OOM errors during the initial memory profiling phase, or result in low performance due to small KV cache space. Consider setting --max-model-len to a smaller value.
WARNING 03-20 20:38:31 [fp8.py:59] Detected fp8 checkpoint. Please note that the format is experimental and subject to change.
INFO 03-20 20:38:31 [config.py:3454] MLA is enabled on a non-cuda platform; forcing chunked prefill and prefix caching to be disabled.
INFO 03-20 20:38:31 [rocm.py:228] Aiter main switch (VLLM_USE_AITER) is not set. Disabling individual Aiter components
INFO 03-20 20:38:31 [async_llm_engine.py:267] Initializing a V0 LLM engine (v0.7.4.dev49+gc0dd5adf6) with config: model='./DeepSeek-V3', speculative_config=None, tokenizer='./DeepSeek-V3', skip_tokenizer_init=False, tokenizer_mode=auto, revision=None, override_neuron_config=None, tokenizer_revision=None, trust_remote_code=True, dtype=torch.bfloat16, max_seq_len=163840, download_dir=None, load_format=LoadFormat.AUTO, tensor_parallel_size=1, pipeline_parallel_size=2, disable_custom_all_reduce=True, quantization=fp8, enforce_eager=False, kv_cache_dtype=auto, device_config=cuda, decoding_config=DecodingConfig(guided_decoding_backend='xgrammar'), observability_config=ObservabilityConfig(show_hidden_metrics=False, otlp_traces_endpoint=None, collect_model_forward_time=False, collect_model_execute_time=False), seed=0, served_model_name=./DeepSeek-V3, num_scheduler_steps=1, multi_step_stream_outputs=True, enable_prefix_caching=False, chunked_prefill_enabled=False, use_async_output_proc=False, disable_mm_preprocessor_cache=False, mm_processor_kwargs=None, pooler_config=None, compilation_config={"splitting_ops":[],"compile_sizes":[],"cudagraph_capture_sizes":[256,248,240,232,224,216,208,200,192,184,176,168,160,152,144,136,128,120,112,104,96,88,80,72,64,56,48,40,32,24,16,8,4,2,1],"max_capture_size":256}, use_cached_outputs=False,
INFO 03-20 20:38:31 [config.py:209] Replacing legacy 'type' key with 'rope_type'
Traceback (most recent call last):
File "/usr/local/bin/vllm", line 8, in <module>
sys.exit(main())
^^^^^^
File "/usr/local/lib/python3.12/dist-packages/vllm/entrypoints/cli/main.py", line 73, in main
args.dispatch_function(args)
File "/usr/local/lib/python3.12/dist-packages/vllm/entrypoints/cli/serve.py", line 34, in cmd
uvloop.run(run_server(args))
File "/usr/local/lib/python3.12/dist-packages/uvloop/__init__.py", line 109, in run
return __asyncio.run(
^^^^^^^^^^^^^^
File "/usr/lib/python3.12/asyncio/runners.py", line 195, in run
return runner.run(main)
^^^^^^^^^^^^^^^^
File "/usr/lib/python3.12/asyncio/runners.py", line 118, in run
return self._loop.run_until_complete(task)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "uvloop/loop.pyx", line 1518, in uvloop.loop.Loop.run_until_complete
File "/usr/local/lib/python3.12/dist-packages/uvloop/__init__.py", line 61, in wrapper
return await main
^^^^^^^^^^
File "/usr/local/lib/python3.12/dist-packages/vllm/entrypoints/openai/api_server.py", line 946, in run_server
async with build_async_engine_client(args) as engine_client:
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/lib/python3.12/contextlib.py", line 210, in __aenter__
return await anext(self.gen)
^^^^^^^^^^^^^^^^^^^^^
File "/usr/local/lib/python3.12/dist-packages/vllm/entrypoints/openai/api_server.py", line 138, in build_async_engine_client
async with build_async_engine_client_from_engine_args(
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/lib/python3.12/contextlib.py", line 210, in __aenter__
return await anext(self.gen)
^^^^^^^^^^^^^^^^^^^^^
File "/usr/local/lib/python3.12/dist-packages/vllm/entrypoints/openai/api_server.py", line 162, in build_async_engine_client_from_engine_args
engine_client = AsyncLLMEngine.from_engine_args(
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/local/lib/python3.12/dist-packages/vllm/engine/async_llm_engine.py", line 644, in from_engine_args
engine = cls(
^^^^
File "/usr/local/lib/python3.12/dist-packages/vllm/engine/async_llm_engine.py", line 594, in __init__
self.engine = self._engine_class(*args, **kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/local/lib/python3.12/dist-packages/vllm/engine/async_llm_engine.py", line 267, in __init__
super().__init__(*args, **kwargs)
File "vllm/engine/llm_engine.py", line 273, in vllm.engine.llm_engine.LLMEngine.__init__
File "/usr/local/lib/python3.12/dist-packages/vllm/executor/executor_base.py", line 271, in __init__
super().__init__(*args, **kwargs)
File "/usr/local/lib/python3.12/dist-packages/vllm/executor/executor_base.py", line 52, in __init__
self._init_executor()
File "/usr/local/lib/python3.12/dist-packages/vllm/executor/mp_distributed_executor.py", line 60, in _init_executor
self._check_cuda()
File "/usr/local/lib/python3.12/dist-packages/vllm/executor/mp_distributed_executor.py", line 46, in _check_cuda
raise RuntimeError(
RuntimeError: please ensure that world_size (2) is less than than max local gpu count (1)
```
It seemed didn't recognize gpu on other nodes.
### Before submitting a new issue...
- [x] Make sure you already searched for relevant issues, and asked the chatbot living at the bottom right corner of the [documentation page](https://docs.vllm.ai/en/latest/), which can answer lots of frequently asked questions. | open | 2025-03-20T12:44:35Z | 2025-03-20T12:44:35Z | https://github.com/vllm-project/vllm/issues/15217 | [
"bug"
] | chn-lee-yumi | 0 |
plotly/dash | plotly | 3,201 | Typescript Components which have props with hyphens generate a syntax error in Python | When converting components which have props containing a hyphen, e.g. "aria-expanded", the generated Python class has "area-expander" in its parameters, which throws an invalid syntax error.
```
def __init__(self, children=None, value=Component.REQUIRED, aria-expanded=Component.UNDEFINED, **kwargs):
self._prop_names = ['children', 'id', 'about', 'accessKey', 'aria-expanded', ]
```
`SyntaxError: invalid syntax
` | open | 2024-04-29T16:16:12Z | 2025-03-07T14:15:49Z | https://github.com/plotly/dash/issues/3201 | [] | tsveti22 | 0 |
globaleaks/globaleaks-whistleblowing-software | sqlalchemy | 3,342 | Wizard cannot be completed | When reinstalling, the wizard starts as usual.
Unfortunately, it does not continue at step 5, the confirmation of the license. When pressing the "continue" button nothing happens and no error is displayed. We have tested it on different systems and with different browsers - unfortunately without any result.
We tried to call the wizard via subdomain or IP address. Same result.
This is the output in the console:
The Cross-Origin-Opener-Policy header has been ignored, because the URL's origin was untrustworthy. It was defined either in the final response or a redirect. Please deliver the response using the HTTPS protocol. You can also use the 'localhost' origin instead. See https://www.w3.org/TR/powerful-features/#potentially-trustworthy-origin and https://html.spec.whatwg.org/#the-cross-origin-opener-policy-header.
How can we complete the wizard? | closed | 2023-02-03T13:58:36Z | 2023-02-06T08:37:48Z | https://github.com/globaleaks/globaleaks-whistleblowing-software/issues/3342 | [] | stefan-h3 | 2 |
serengil/deepface | machine-learning | 1,323 | [FEATURE]: support pipx install on Ubuntu 24.04, fails with "ValueError: You have tensorflow 2.17.0 and this requires tf-keras package. Please run `pip install tf-keras` or downgrade your tensorflow" | ### Description
It would be good if `pipx install` worked for Ubuntu 24.04:
```
pipx install deepface==0.0.93
```
but then:
```
deepface help
```
blows up with:
```
2024-08-27_21-42-31@ciro@ciro-p14s$ deepface help
2024-08-27 21:42:38.810014: I tensorflow/core/util/port.cc:153] oneDNN custom operations are on. You may see slightly different numerical results due to fl
oating-point round-off errors from different computation orders. To turn them off, set the environment variable `TF_ENABLE_ONEDNN_OPTS=0`.
2024-08-27 21:42:38.810549: I external/local_xla/xla/tsl/cuda/cudart_stub.cc:32] Could not find cuda drivers on your machine, GPU will not be used.
2024-08-27 21:42:38.814904: I external/local_xla/xla/tsl/cuda/cudart_stub.cc:32] Could not find cuda drivers on your machine, GPU will not be used.
2024-08-27 21:42:38.822156: E external/local_xla/xla/stream_executor/cuda/cuda_fft.cc:485] Unable to register cuFFT factory: Attempting to register factory
for plugin cuFFT when one has already been registered
2024-08-27 21:42:38.834452: E external/local_xla/xla/stream_executor/cuda/cuda_dnn.cc:8454] Unable to register cuDNN factory: Attempting to register factor
y for plugin cuDNN when one has already been registered
2024-08-27 21:42:38.837693: E external/local_xla/xla/stream_executor/cuda/cuda_blas.cc:1452] Unable to register cuBLAS factory: Attempting to register fact
ory for plugin cuBLAS when one has already been registered
2024-08-27 21:42:38.846987: I tensorflow/core/platform/cpu_feature_guard.cc:210] This TensorFlow binary is optimized to use available CPU instructions in p
erformance-critical operations.
To enable the following instructions: AVX2 AVX512F AVX512_VNNI AVX512_BF16 FMA, in other operations, rebuild TensorFlow with the appropriate compiler flags
.
2024-08-27 21:42:39.511588: W tensorflow/compiler/tf2tensorrt/utils/py_utils.cc:38] TF-TRT Warning: Could not find TensorRT
Traceback (most recent call last):
File "/home/ciro/.local/pipx/venvs/deepface/lib/python3.12/site-packages/retinaface/commons/package_utils.py", line 19, in validate_for_keras3
import tf_keras
ModuleNotFoundError: No module named 'tf_keras'
The above exception was the direct cause of the following exception:
Traceback (most recent call last):
File "/home/ciro/.local/bin/deepface", line 5, in <module>
from deepface.DeepFace import cli
File "/home/ciro/.local/pipx/venvs/deepface/lib/python3.12/site-packages/deepface/DeepFace.py", line 20, in <module>
from deepface.modules import (
File "/home/ciro/.local/pipx/venvs/deepface/lib/python3.12/site-packages/deepface/modules/modeling.py", line 16, in <module>
from deepface.models.face_detection import (
File "/home/ciro/.local/pipx/venvs/deepface/lib/python3.12/site-packages/deepface/models/face_detection/RetinaFace.py", line 3, in <module>
from retinaface import RetinaFace as rf
File "/home/ciro/.local/pipx/venvs/deepface/lib/python3.12/site-packages/retinaface/RetinaFace.py", line 20, in <module>
package_utils.validate_for_keras3()
File "/home/ciro/.local/pipx/venvs/deepface/lib/python3.12/site-packages/retinaface/commons/package_utils.py", line 24, in validate_for_keras3
raise ValueError(
ValueError: You have tensorflow 2.17.0 and this requires tf-keras package. Please run `pip install tf-keras` or downgrade your tensorflow.
```
And you can't just:
```
pip install tf-keras
```
because it blows up with:
```
error: externally-managed-environment
× This environment is externally managed
╰─> To install Python packages system-wide, try apt install
python3-xyz, where xyz is the package you are trying to
install.
If you wish to install a non-Debian-packaged Python package,
create a virtual environment using python3 -m venv path/to/venv.
Then use path/to/venv/bin/python and path/to/venv/bin/pip. Make
sure you have python3-full installed.
If you wish to install a non-Debian packaged Python application,
it may be easiest to use pipx install xyz, which will manage a
virtual environment for you. Make sure you have pipx installed.
See /usr/share/doc/python3.12/README.venv for more information.
note: If you believe this is a mistake, please contact your Python installation or OS distribution provider. You can override this, at the risk of breaking your Python installation or OS, by passing --break-system-packages.
hint: See PEP 668 for the detailed specification.
``` | closed | 2024-08-27T20:46:13Z | 2024-10-22T17:04:49Z | https://github.com/serengil/deepface/issues/1323 | [
"enhancement",
"dependencies"
] | cirosantilli | 2 |
flairNLP/flair | nlp | 3,081 | [Question]: Combining BERT & Flair | ### Question
Hey,
I have some questions regarding the [following tutorial](https://github.com/flairNLP/flair/blob/master/resources/docs/TUTORIAL_4_ELMO_BERT_FLAIR_EMBEDDING.md) for creating a multi-lingual Flair Stacked embedding model that combines Flair Embeddings & BERT.
https://github.com/flairNLP/flair/blob/23618cd8e072ec2a3f325985c18bfa14315c9554/resources/docs/TUTORIAL_4_ELMO_BERT_FLAIR_EMBEDDING.md?plain=1#L33-L42
By default, the parameter fine_tune is set to True. My question is **should you fine-tune the TransformerWordEmbeddings when including them in a Flair Stacked Embedding?**
I have noticed that training this model can be incredibly slow, even on a big GPU (I have an A100 with 80GB available).
With 2400 training sentences takes about 40 minutes per epoch with a mini_batch_size=4 and a mini_chunk_size=1 | closed | 2023-02-06T07:25:35Z | 2023-02-07T17:31:38Z | https://github.com/flairNLP/flair/issues/3081 | [
"question"
] | Guust-Franssens | 1 |
xlwings/xlwings | automation | 1,777 | Unable to include Image comment in xlwings | #### OS (e.g. Windows 10 or macOS Sierra)
Windows 10
#### Versions of xlwings, Excel and Python (e.g. 0.11.8, Office 365, Python 3.7)
0.24.9, Office 2019 and Python 3.9.7
#### Describe your issue (incl. Traceback!)
I unable to include image in XL comment. In Excel, if we follow the following steps we would able to include pictures in the comment. I wanted to make it through python
1. Right-click the cell which contains the comment.
2. Choose Show/Hide Comments, and clear any text from the comment.
3. Click on the border of the comment, to select it.
4. Choose Format|Comment
5. On the Colors and Lines tab, click the drop-down arrow for Color.
6. Click Fill Effects
7. On the picture tab, click Select Picture
8. Locate and select the picture
9. To keep the picture in proportion, add a check mark to Lock Picture Aspect Ratio
10. Click Insert, click OK, click OK
Sample Output expecting from Python is attached above.
Could you please suggest how to achieve this in xlwings? | open | 2021-12-02T06:04:26Z | 2022-02-01T13:26:50Z | https://github.com/xlwings/xlwings/issues/1777 | [] | agrangaraj | 1 |
pydantic/pydantic-ai | pydantic | 1,143 | Validation errors for _GeminiResponse | ### Initial Checks
- [x] I confirm that I'm using the latest version of Pydantic AI
### Description
I get the below error when trying to run simple agent with Gemini provider:
```
lib/python3.11/site-packages/pydantic/type_adapter.py", line 468, in validate_json
return self.validator.validate_json(
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
pydantic_core._pydantic_core.ValidationError: 4 validation errors for _GeminiResponse
candidates.0.avgLogProbs
Field required [type=missing, input_value={'content': {'parts': [{'...': -0.05566399544477463}, input_type=dict]
For further information visit https://errors.pydantic.dev/2.11/v/missing
candidates.0.index
Field required [type=missing, input_value={'content': {'parts': [{'...': -0.05566399544477463}, input_type=dict]
For further information visit https://errors.pydantic.dev/2.11/v/missing
candidates.0.safetyRatings
Field required [type=missing, input_value={'content': {'parts': [{'...': -0.05566399544477463}, input_type=dict]
For further information visit https://errors.pydantic.dev/2.11/v/missing
promptFeedback
Field required [type=missing, input_value={'candidates': [{'content...on': 'gemini-2.0-flash'}, input_type=dict]
For further information visit https://errors.pydantic.dev/2.11/v/missing
```
### Example Code
```Python
from pydantic_ai import Agent
agent = Agent(model="google-gla:gemini-2.0-flash")
result = agent.run_sync("Hello, how are you?")
print(result.data)
```
### Python, Pydantic AI & LLM client version
```Text
Python: 3.11.11
Pydantic: 2.11.0b1
Pydantic AI: 0.0.40
``` | closed | 2025-03-16T23:08:50Z | 2025-03-17T09:49:05Z | https://github.com/pydantic/pydantic-ai/issues/1143 | [
"bug"
] | torayeff | 2 |
Lightning-AI/pytorch-lightning | data-science | 20,110 | CSV Logger acts weirdly in Callbacks | ### Bug description
I use CSV logger inside callbacks as the following
```
pl_module.log('epoch_throughput', throughput, on_epoch=True, logger=True, sync_dist=True, reduce_fx="sum")
pl_module.log('epoch_time', epoch_time, logger=True, on_epoch=True, sync_dist=True)
```
When I use two callbacks to log data at the same time point (i.e. on_train_epoch_end), it logs the data into two rows

### What version are you seeing the problem on?
master
### How to reproduce the bug
```python
class CallbackX(Callback):
def on_train_epoch_start(self, trainer, pl_module):
self.epoch_start_time = time.time()
def on_train_epoch_end(self, trainer, pl_module):
epoch_time = time.time() - self.epoch_start_time
pl_module.log('epoch_time', epoch_time, logger=True, on_epoch=True, sync_dist=True)
class CallbackY(Callback):
def on_train_epoch_start(self, trainer, pl_module):
self.epoch_start_time = time.time()
def on_train_epoch_end(self, trainer, pl_module):
epoch_time = time.time() - self.epoch_start_time
pl_module.log('epoch_time2', epoch_time, logger=True, on_epoch=True, sync_dist=True)
```
### Error messages and logs
```
# Error messages and logs here please
```
### Environment
<details>
<summary>Current environment</summary>
* CUDA:
- GPU: Tesla T4
- available: False
- version: 12.1
* Lightning:
- lightning: 2.3.0
- lightning-utilities: 0.11.2
- pytorch-lightning: 2.3.0
- torch: 2.3.1
- torch-tb-profiler: 0.4.3
- torchmetrics: 1.4.0.post0
- torchvision: 0.18.1
* Packages:
- absl-py: 2.1.0
- aiohttp: 3.9.5
- aiosignal: 1.3.1
- annotated-types: 0.7.0
- attrs: 23.2.0
- certifi: 2024.6.2
- cffi: 1.16.0
- charset-normalizer: 3.3.2
- cloudpickle: 3.0.0
- deepspeed: 0.14.4
- dool: 1.3.2
- filelock: 3.15.4
- frozenlist: 1.4.1
- fsspec: 2024.6.0
- graphviz: 0.8.4
- grpcio: 1.64.1
- gviz-api: 1.10.0
- hjson: 3.1.0
- idna: 3.7
- jinja2: 3.1.4
- lightning: 2.3.0
- lightning-utilities: 0.11.2
- markdown: 3.6
- markupsafe: 2.1.5
- mpmath: 1.3.0
- multidict: 6.0.5
- mxnet: 1.9.1
- networkx: 3.3
- ninja: 1.11.1.1
- numpy: 1.25.0
- nvidia-cublas-cu12: 12.1.3.1
- nvidia-cuda-cupti-cu12: 12.1.105
- nvidia-cuda-nvrtc-cu12: 12.1.105
- nvidia-cuda-runtime-cu12: 12.1.105
- nvidia-cudnn-cu12: 8.9.2.26
- nvidia-cufft-cu12: 11.0.2.54
- nvidia-curand-cu12: 10.3.2.106
- nvidia-cusolver-cu12: 11.4.5.107
- nvidia-cusparse-cu12: 12.1.0.106
- nvidia-ml-py: 12.555.43
- nvidia-nccl-cu12: 2.20.5
- nvidia-nvjitlink-cu12: 12.5.40
- nvidia-nvtx-cu12: 12.1.105
- packaging: 24.1
- pandas: 2.2.2
- pillow: 10.3.0
- pip: 24.1
- protobuf: 4.25.3
- psutil: 6.0.0
- py-cpuinfo: 9.0.0
- pycparser: 2.22
- pydantic: 2.7.4
- pydantic-core: 2.18.4
- python-dateutil: 2.9.0.post0
- pytorch-lightning: 2.3.0
- pytz: 2024.1
- pyyaml: 6.0.1
- requests: 2.32.3
- setuptools: 65.5.0
- six: 1.16.0
- sympy: 1.12.1
- tensorboard: 2.17.0
- tensorboard-data-server: 0.7.2
- tensorboard-plugin-profile: 2.15.1
- tensorboardx: 2.6.2.2
- torch: 2.3.1
- torch-tb-profiler: 0.4.3
- torchmetrics: 1.4.0.post0
- torchvision: 0.18.1
- tqdm: 4.66.4
- triton: 2.3.1
- typing-extensions: 4.12.2
- tzdata: 2024.1
- urllib3: 2.2.2
- werkzeug: 3.0.3
- wheel: 0.43.0
- yarl: 1.9.4
* System:
- OS: Linux
- architecture:
- 64bit
- ELF
- processor: x86_64
- python: 3.11.7
- release: 3.10.0-1127.19.1.el7.x86_64
- version: #1 SMP Tue Aug 25 17:23:54 UTC 2020
</details>
### More info
_No response_ | open | 2024-07-20T19:17:17Z | 2024-07-20T19:17:31Z | https://github.com/Lightning-AI/pytorch-lightning/issues/20110 | [
"bug",
"needs triage",
"ver: 2.2.x"
] | oabuhamdan | 0 |
globaleaks/globaleaks-whistleblowing-software | sqlalchemy | 4,154 | Translation issue in "Revoke access" | ### What version of GlobaLeaks are you using?
4.15.8
### What browser(s) are you seeing the problem on?
Microsoft Edge
### What operating system(s) are you seeing the problem on?
Windows
### Describe the issue
The "close" button in the Revoke Access window is forced to be in English in all language versions. It is not possible to set a translation.

### Proposed solution
_No response_ | closed | 2024-08-14T13:34:01Z | 2024-09-06T16:37:48Z | https://github.com/globaleaks/globaleaks-whistleblowing-software/issues/4154 | [
"T: Bug",
"C: Client"
] | evariitta | 1 |
open-mmlab/mmdetection | pytorch | 11,777 | Is it possible to calculate a validation loss? | I want to conduct an experiment with an object detection model that I trained. My experiment is as follows: I want to understand a little more about the images in my test set. For this, I would like to obtain some individual metrics per image from the test dataset, in addition to getting the loss (validation) for each image. My current code is as follows:
```
config_file = 'swin/custom_mask_rcnn_swin-s-p4-w7_fpn_fp16_ms-crop-3x_coco.py'
checkpoint_file = 'mmdet/swin/epoch_40.pth'
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
cfg = Config.fromfile(config_file)
dataset = build_dataset(cfg.data.test)
data_loader = build_dataloader(
dataset,
samples_per_gpu=cfg.data.samples_per_gpu,
workers_per_gpu=cfg.data.workers_per_gpu,
dist=False,
shuffle=False
)
model = init_detector(config_file, checkpoint_file, device=device)
dataset = build_dataset(cfg.data.test)
model.eval()
results = []
prog_bar = mmcv.ProgressBar(len(dataset))
for i, data in enumerate(data_loader):
with torch.no_grad():
data = scatter(data, [device])[0]
result = model(return_loss=True, **data)
prog_bar.update()
for elem in result:
print(elem)
```
I am getting a tuple of tensors as the output, like the example below:
```
([array([], shape=(0, 5), dtype=float32), array([[1.3915492e+02, 2.6474759e+02, 1.5779597e+02, 2.9583871e+02,
1.4035654e-01],
[1.3974554e+02, 2.6289932e+02, 1.6024533e+02, 3.1977335e+02,
9.8360136e-02]], dtype=float32), array([[8.7633228e+02, 2.1958812e+02, 8.8837036e+02, 2.4044472e+02,
7.2545749e-01]], dtype=float32), array([], shape=(0, 5), dtype=float32), array([[8.7622699e+02, 2.1944591e+02, 8.8825470e+02, 2.4135008e+02,
2.4266671e-01]], dtype=float32)], [[], [array([[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
...,
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False]]), array([[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
...,
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False]])], [array([[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
...,
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False]])], [], [array([[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
...,
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False]])]])
([array([], shape=(0, 5), dtype=float32), array([], shape=(0, 5), dtype=float32), array([], shape=(0, 5), dtype=float32), array([], shape=(0, 5), dtype=float32), array([[2.9585770e+02, 3.7026846e+02, 3.1038290e+02, 3.8506241e+02,
5.9989877e-02]], dtype=float32)], [[], [], [], [], [array([[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
...,
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False],
[False, False, False, ..., False, False, False]])]])
```
My intuition tells me that this output is something like bounding box positions (x, y, w, h), confidence score per class, and also binary mask information due to the boolean values. Additionally, I added the argument "return_loss=True", and I imagine that some of this information must also be related to the loss that I want to obtain. How can I parse this output? That is, identify what each of the pieces of information in these results is to be able to find the desired loss.
I'm using MMDetection v2.28.2. | open | 2024-06-07T22:40:25Z | 2024-06-07T22:43:20Z | https://github.com/open-mmlab/mmdetection/issues/11777 | [] | psantiago-lsbd | 0 |
tensorpack/tensorpack | tensorflow | 876 | Save Model | Hi, i have a question about TrainConfig. How to save model in specific steps rather than every epoch. | closed | 2018-08-28T12:15:38Z | 2018-09-06T20:06:40Z | https://github.com/tensorpack/tensorpack/issues/876 | [
"usage"
] | lizaigaoge550 | 1 |
microsoft/unilm | nlp | 724 | Question about learning performance when finetuning LayoutLM V3 on PubLayNet-like dataset | I'm trying to finetune _LayoutLM V3_ [Base model](https://huggingface.co/microsoft/layoutlmv3-base) using the provided `dit/train_net.py` script on my own custom dataset that is similar to _PubLayNet_. The learning starts well but after reaching the first checkpoint (2000 iterations) and doing the first evaluation the loss starts going up and the accuracy keeps going down.
Someone can point me to some critical factors that may lead to this behavior ? Thanks in advance !
| closed | 2022-05-19T21:16:13Z | 2022-05-23T03:18:37Z | https://github.com/microsoft/unilm/issues/724 | [] | naourass | 3 |
TencentARC/GFPGAN | deep-learning | 290 | Enhanced | open | 2022-10-13T18:49:37Z | 2022-10-13T18:49:37Z | https://github.com/TencentARC/GFPGAN/issues/290 | [] | abhaydasah | 0 | |
vimalloc/flask-jwt-extended | flask | 481 | Refresh with cookies | I have a pretty standard refresh endpoint with the intent to create a new access token and set the new access token as a cookie.
```python
@identity_bp.get('/refresh')
@jwt_required(refresh=True)
def refresh_token():
identity = get_jwt_identity()
access_token = create_access_token(identity=identity)
res = jsonify({'refresh': True})
set_access_cookies(res, access_token)
return jsonify(access_token=access_token), HTTPStatus.OK
```
A new access token is created, but the access token cookie remains the same as the access token set upon login. So, despite refreshing, the token stored in the cookie is old.
For reference, this is our login endpoint, which does successfully create access/refresh tokens and stores them into cookies:
```python
@identity.post('/login')
def login():
email = request.json.get('email', None)
password = request.json.get('password', None)
# ... some logic
access_token = create_access_token(identity=identity)
refresh_token = create_refresh_token(identity=identity)
res = jsonify({'msg': 'Successful login.',
'access_token': access_token,
'refresh_token': refresh_token})
set_access_cookies(res, access_token)
set_refresh_cookies(res, refresh_token)
return res, HTTPStatus.OK
```
In our React web app, we handle the logic necessary logic to refresh the tokens. I can confirm that when we hit the refresh endpoint there, a new token is created, but the cookie just doesn't change. I've also tried to unset the cookies with `unset_access_cookies`, but it also doesn't remove the cookie.
Perhaps this is just a misunderstanding on my end with how overriding the cookie works, or a bug. I'm not too sure. | closed | 2022-06-02T03:01:40Z | 2022-06-02T03:16:55Z | https://github.com/vimalloc/flask-jwt-extended/issues/481 | [] | leelerm | 2 |
piskvorky/gensim | machine-learning | 3,368 | CoherenceModel does not finish with computing | #### Problem description
When computing coherence scores, it newer finishes with computing on a bit bigger dataset. Run the code below (with the provided dataset) to reproduce.
#### Steps/code/corpus to reproduce
```python
with open("coherence-bug.pkl", "rb") as f:
model, tokens = pickle.load(f)
print("conherence")
print(datetime.now())
t = time.time()
cm = CoherenceModel(model=model, texts=tokens, coherence="c_v")
coherence = cm.get_coherence()
print(time.time() - t)
```
[coherence-bug.pkl.zip](https://github.com/RaRe-Technologies/gensim/files/9178910/coherence-bug.pkl.zip)
#### Versions
The bug appears on Gensim version 4.2, but it does not happen on 4.1.2
macOS-10.16-x86_64-i386-64bit
Python 3.8.12 (default, Oct 12 2021, 06:23:56)
[Clang 10.0.0 ]
Bits 64
NumPy 1.22.3
SciPy 1.8.1
gensim 4.2.1.dev0
FAST_VERSION 0
| closed | 2022-07-25T07:23:04Z | 2022-12-06T07:38:18Z | https://github.com/piskvorky/gensim/issues/3368 | [
"bug",
"difficulty easy",
"impact HIGH",
"reach LOW"
] | PrimozGodec | 5 |
modoboa/modoboa | django | 2,569 | LDAP: password sync is broken | # Impacted versions
* OS Type: Debian
* OS Version: bullseye 11.4
* Database Type: MySQL / MariaDB
* Database version: 10.5.15
* Modoboa: 2.0.1
* installer used: yes
* Webserver: nginx
# Steps to reproduce
1. have openldap / slapd installation
(i can certainly get more info on this, but i was only tasked with fixing the issue and have not yet much fiddled with slapd config)
the important point is: slapd must automatically encrypt new userPasswords if it thinks the hash type is unknown
2. install modoboa
3. configure ldap connection
4. use modoboa to change users password
# Current behavior
there are two issues we were able to identify:
## 1. modoboa does not send password hashing scheme to ldap-server
TL;DR: modoboa sends `$6$rounds=70000$...` to ldap server instead of `{SHA512-CRYPT}$6$rounds=70000$...`
we were able to capture this with tcpdump. when modoboa sends password to ldap https://github.com/modoboa/modoboa/blob/c26379478445da5888bf05be0ba4cf98e20ea046/modoboa/ldapsync/lib.py#L119 we always found it only sends the actual hash starting with `$6$rounds=70000$...`
## 2. ldap does only understand "{CRYPT}"
TL;DR: modoboa sends `{SHA512-CRYPT}$6$rounds=70000$...` to ldap server instead of `{CRYPT}$6$rounds=70000$...`
the second issue is with slapd only supporting {CRYPT} as a scheme. it can understand, operate and generate multiple different hash types (like `$1$`, `$5$`, and `$6$`) but this is controlled only by the actual hash, not the scheme prefix.
these do not work: {SHA256-CRYPT} {SHA512-CRYPT} {BLF-CRYPT}
but their hashes work if stored in userPassword field in LDAP with {CRYPT} as prefix.
# Expected behavior
included in "Current behavior" section
# Possible fixes:
## 1. modoboa does not send password hashing scheme to ldap-server
the update_ldap_account function uses get_user_password from that same file https://github.com/modoboa/modoboa/blob/c26379478445da5888bf05be0ba4cf98e20ea046/modoboa/ldapsync/lib.py#L50. we identified an issue in line https://github.com/modoboa/modoboa/blob/c26379478445da5888bf05be0ba4cf98e20ea046/modoboa/ldapsync/lib.py#L56 which prevents the scheme from being sent if the accounts is not disabled.
i fixed it by adding parentheses around the disabled check (see: https://github.com/modoboa/modoboa/commit/53dd6c7502d8f8aeb81c8e4caec13a065e92f172)
afterwards tcpdump showed the correct full hash with scheme prepended (i.e. `{SHA512-CRYPT}$6$rounds=70000$...`)
## 2. ldap does only understand "{CRYPT}"
to fix this issue, i added ~~"LDAP_DROP_SCHEME_PREFIX"~~"LDAP_DROP_CRYPT_PREFIX" to settings.py and a check in get_user_password which sets scheme to "{CRYPT" when this option is set. (see: https://github.com/modoboa/modoboa/commit/7432877c3429a0f8bc3d8084b3e00eee7887a0f5)
we verified it working with tcpdump which now showed correct updates to userPassword with full hash like `{CRYPT}$6$rounds=70000$...`
sadly i am not very good with python and was unable to find where to "declare" that new option for the generated settings.py so this needs to be added by s/o else.
| closed | 2022-07-21T18:00:13Z | 2023-03-11T13:49:10Z | https://github.com/modoboa/modoboa/issues/2569 | [
"feedback-needed",
"stale"
] | elgarfo | 7 |
davidsandberg/facenet | computer-vision | 852 | error when running Validate_on_lfw | I am new to the machine learning @davidsandberg
I am trying to run the (Validate_on_lfw .py) in windows but i get this error:
Model directory: pretrained_model
Metagraph file: model-20180402-114759.meta
Checkpoint file: model-20180402-114759.ckpt-275
Traceback (most recent call last):
File "C:\Users\user\Desktop\noor2\project2\facenet-master\facenet-master\src\validate_on_lfw.py", line 166, in <module>
main(parse_arguments(sys.argv[1:]))
File "C:\Users\user\Desktop\noor2\project2\facenet-master\facenet-master\src\validate_on_lfw.py", line 75, in main
facenet.load_model("pretrained_model", input_map=input_map)
File "C:\Users\user\Desktop\noor2\project2\facenet-master\facenet-master\src\facenet.py", line 381, in load_model
saver = tf.train.import_meta_graph(os.path.join(model_exp, meta_file), input_map=input_map)
File "C:\Users\user\AppData\Local\Programs\Python\Python36\lib\site-packages\tensorflow\python\training\saver.py", line 1686, in import_meta_graph
**kwargs)
File "C:\Users\user\AppData\Local\Programs\Python\Python36\lib\site-packages\tensorflow\python\framework\meta_graph.py", line 504, in import_scoped_meta_graph
producer_op_list=producer_op_list)
File "C:\Users\user\AppData\Local\Programs\Python\Python36\lib\site-packages\tensorflow\python\framework\importer.py", line 283, in import_graph_def
raise ValueError('No op named %s in defined operations.' % node.op)
ValueError: No op named DecodeBmp in defined operations.
any help ?
| closed | 2018-08-22T09:28:34Z | 2021-02-15T16:33:18Z | https://github.com/davidsandberg/facenet/issues/852 | [] | nonameever95 | 17 |
httpie/cli | rest-api | 527 | httpie hangs in MobaXterm | I am attempting to use httpie in [mobaXterm](http://mobaxterm.mobatek.net/) (version 8.6) on Windows 7 Ultimate, but httpie hangs and never returns a result.


I configured mobaXterm to use my Windows PATH, and both python and http are on my path:


Although httpie does not work in mobaXterm, it does work in cmd.exe:


I think this issue might be related to another similar issue that I found on StackOverflow:
http://stackoverflow.com/questions/3250749/using-windows-python-from-cygwin
| closed | 2016-10-13T11:32:23Z | 2020-06-18T08:26:13Z | https://github.com/httpie/cli/issues/527 | [] | knilling | 4 |
pyeve/eve | flask | 879 | XML Parsing Error: not well-formed | Hi, I am doing some codes with **python-eve**. I got an issue.
In a **blog editor** page, I have a **textarea** to store **markdown style** string and the other **textarea** to store the **html tags style** string. Then post this two big long strings to the **mongodb** by **eve**. Then I find the blog list by API _http://127.0.0.1:8000/api/v1/blog_, it shows with error:**XML Parsing Error: not well-formed.**

I also use **Django** to find the blog list and it's fine to display on the html page.
I have tested for some codes and found below:
If I post the **markdown style** string, and with code, such as Linux vi code,
```
#vi /root/.vnc/xstartup
```
then I got the error.
| closed | 2016-06-27T17:04:01Z | 2016-06-28T12:25:12Z | https://github.com/pyeve/eve/issues/879 | [] | Penguin7z | 2 |
pydantic/pydantic | pydantic | 11,508 | Add Unicode string normalization tests | ### Initial Checks
- [x] I have searched Google & GitHub for similar requests and couldn't find anything
- [x] I have read and followed [the docs](https://docs.pydantic.dev) and still think this feature is missing
## Description
I'd like to add a test coverage to test_edge_cases.py for Unicode string normalization.
### Proposed Test Coverage
- Emoji with zero-width joiners (👨👩👧👦)
- Variation selectors (🏳️🌈)
- Skin tone modifiers (👋🏾)
- Combining characters vs precomposed characters (e\u0301 vs é)
- Surrogate pairs (🌍)
- Right-to-left text (سلام)
- Bidirectional text mixing (Hello سلام)
- Korean Hangul decomposition (한)
- Replacement characters (�)
- Zero-width spaces (a\u200Bb)
- Characters with different NFKC/NFKD normalizations (ffi)
Proposed Code:
```
def test_unicode_string_normalization():
class UnicodeModel(BaseModel):
text: str
test_cases = {
"zwj_sequence": "👨👩👧👦",
"variation_selector": "🏳️🌈",
"skin_modifier": "👋🏾",
"decomposed_e": "e\u0301",
"precomposed_e": "é",
"surrogate_pair": "🌍",
"rtl_chars": "سلام", # right to left text
"mixed_direction": "Hello سلام", # Mixed RTL and LTR
"decomposed_hangul": "한", # Korean character that can be decomposed
"invalid_sequence": "�", # Replacement character
"zero_width": "a\u200Bb", # Zero-width space
"different_normalized": "ffi", # Character that differs in NFKC/NFKD
}
for case_name, test_str in test_cases.items():
model = UnicodeModel(text=test_str)
assert model.text == test_str
assert len(model.text) == len(test_str) # Checking Length
json_str = model.model_dump_json()
decoded = UnicodeModel.model_validate_json(json_str) # Checking if it survived json roundtrip
assert decoded.text == test_str
for form in ['NFC', 'NFD', 'NFKC', 'NFKD']:
normalized = unicodedata.normalize(form, test_str)
normalized_model = UnicodeModel(text=normalized)
assert normalized_model.text == normalized
renormalized = unicodedata.normalize(form, normalized_model.text) # checking if it is stable under normalization
assert renormalized == normalized
```
### Affected Components
- [ ] [Compatibility between releases](https://docs.pydantic.dev/changelog/)
- [x] [Data validation/parsing](https://docs.pydantic.dev/concepts/models/#basic-model-usage)
- [ ] [Data serialization](https://docs.pydantic.dev/concepts/serialization/) - `.model_dump()` and `.model_dump_json()`
- [ ] [JSON Schema](https://docs.pydantic.dev/concepts/json_schema/)
- [ ] [Dataclasses](https://docs.pydantic.dev/concepts/dataclasses/)
- [ ] [Model Config](https://docs.pydantic.dev/concepts/config/)
- [ ] [Field Types](https://docs.pydantic.dev/api/types/) - adding or changing a particular data type
- [ ] [Function validation decorator](https://docs.pydantic.dev/concepts/validation_decorator/)
- [ ] [Generic Models](https://docs.pydantic.dev/concepts/models/#generic-models)
- [ ] [Other Model behaviour](https://docs.pydantic.dev/concepts/models/) - `model_construct()`, pickling, private attributes, ORM mode
- [ ] [Plugins](https://docs.pydantic.dev/) and integration with other tools - mypy, FastAPI, python-devtools, Hypothesis, VS Code, PyCharm, etc. | closed | 2025-02-28T19:01:53Z | 2025-03-03T17:51:50Z | https://github.com/pydantic/pydantic/issues/11508 | [
"feature request"
] | saturnines | 2 |
encode/apistar | api | 8 | Request Data | This is a bit of a missing component ATM.
Parse the request body, raising 400 or 415 if required.
We'll want to automatically created TypedRequestData components for type arguments on POST requests.
| closed | 2017-03-30T09:41:05Z | 2017-08-17T11:38:55Z | https://github.com/encode/apistar/issues/8 | [
"Component"
] | tomchristie | 1 |
feder-cr/Jobs_Applier_AI_Agent_AIHawk | automation | 99 | Huge delay in filling the forms | It takes around 30 seconds to fill one single input field. Needed to be optimized. | closed | 2024-08-28T04:04:52Z | 2024-08-29T12:55:37Z | https://github.com/feder-cr/Jobs_Applier_AI_Agent_AIHawk/issues/99 | [] | sanjeethboddi | 1 |
Yorko/mlcourse.ai | numpy | 369 | Docker Image | The Docker image needs to be updated to the latest packages versioning, maybe... | closed | 2018-10-10T12:09:24Z | 2018-10-11T13:47:20Z | https://github.com/Yorko/mlcourse.ai/issues/369 | [
"enhancement"
] | AdityaSoni19031997 | 1 |
anselal/antminer-monitor | dash | 64 | Fan speed color | For the S9 we have to two fans. One front and one rear. Antminer-monitor display both of them.
We should think of a color code. If the fan is less than 5000 = green, 5000 or more = yellow and more than 6000 = red.
More important, if the rear fan speed is faster than the front, then the rear **fan speed should blink**.
This would bring the situation to our eye. We want maximum air pressure inside the antminer so that there is more air particules in contact with the heat sink to exchange the heat. If the rear one is faster, this could mean that the air pressure is too low inside the antminer, or we have a problem on the antminer or that the output tube or output windows (depending on our setups) is too small.
| open | 2018-01-28T15:12:23Z | 2018-02-22T09:33:28Z | https://github.com/anselal/antminer-monitor/issues/64 | [
":sparkles: enhancement :sparkles:"
] | carlcbilodeau | 7 |
WZMIAOMIAO/deep-learning-for-image-processing | pytorch | 269 | 关于faster rcnn的评价指标 | 您好,关于关于faster rcnn的评价指标有点问题向您请教下。就是您代码中给出的评价指标是map,是按照coco数据集的格式给出的,但是现在我想指导具体的每一类的ap指,不知道应该从哪里去修改代码呢? | closed | 2021-05-23T08:08:52Z | 2022-07-27T08:32:42Z | https://github.com/WZMIAOMIAO/deep-learning-for-image-processing/issues/269 | [] | hhhuuuh | 2 |
polakowo/vectorbt | data-visualization | 157 | Change line color in plot | I'm trying to change the line color of the RSI plot that I make from red to green. I looked through the documentation and tried to mimic an example. Here is the code:
```python
import vectorbt as vbt
btc_prices = vbt.BinanceData.download('BTCUSDT', interval='1d')
rsi = vbt.RSI.run(btc_prices.get('Close'))
long_entries = rsi.rsi_above(50.0, crossover=True)
long_exits = rsi.rsi_above(70.0)
portfolio = vbt.Portfolio.from_signals(btc_prices.get('Close'), long_entries, long_exits, init_cash=1000)
def plot_rsi(portfolio, rsi=None, add_trace_kwargs=None, fig=None):
rsi.rsi.vbt.plot(add_trace_kwargs=add_trace_kwargs, fig=fig)
rsi_subplots = ('rsi', dict(
title='RSI',
yaxis_title='rsi_val',
can_plot_groups=False,
plot_func=plot_rsi
))
subplots =['orders', rsi_subplots, 'trade_returns', 'cum_returns']
portfolio.plot(subplots=subplots, rsi_kwargs=dict(rsi=rsi, add_trace_kwargs=dict(line_color='green'))).show()
```
but this produces the following error:
```python
Traceback (most recent call last):
File "/home/aclifton/quant/backtests/test_vectorbt.py", line 36, in <module>
portfolio.plot(subplots=subplots, rsi_kwargs=dict(rsi=rsi, add_trace_kwargs=dict(line_color='green'))).show()
File "/home/aclifton/venvs/quant/lib/python3.7/site-packages/vectorbt/portfolio/base.py", line 2979, in plot
plot_func(self_col, **custom_kwargs, fig=fig)
File "/home/aclifton/quant/backtests/test_vectorbt.py", line 25, in plot_rsi
rsi.rsi.vbt.plot(add_trace_kwargs=add_trace_kwargs, fig=fig)
File "/home/aclifton/venvs/quant/lib/python3.7/site-packages/vectorbt/generic/accessors.py", line 1213, in plot
**kwargs
File "/home/aclifton/venvs/quant/lib/python3.7/site-packages/vectorbt/generic/plotting.py", line 344, in __init__
fig.add_trace(scatter, **add_trace_kwargs)
TypeError: add_trace() got an unexpected keyword argument 'line_color'
```
For reference, I was looking at the following example found [here](https://polakowo.io/vectorbt/docs/portfolio/base.html#vectorbt.portfolio.base.Portfolio):
```python
>>> from vectorbt.utils.colors import adjust_opacity
>>> portfolio.plot(
... subplots=['drawdowns', 'underwater'],
... drawdowns_kwargs=dict(top_n=3),
... underwater_kwargs=dict(
... trace_kwargs=dict(
... line_color='#FF6F00',
... fillcolor=adjust_opacity('#FF6F00', 0.3)
... )
... )
... )
``` | closed | 2021-06-04T14:06:34Z | 2021-06-04T16:30:57Z | https://github.com/polakowo/vectorbt/issues/157 | [] | aclifton314 | 12 |
biolab/orange3 | scikit-learn | 6,312 | Row number as a variable |
**What's your use case?**
When using Select Rows, it should also be possible to select by row number. For example, in some datasets from connected products, the first x rows originate from prototypes or they are otherwise not representative, for instance due to start-up problems. In such cases it is useful to be able to use "row number is greater than" as a row selection criterion.
I have also once encountered a situation where I would have liked to use the row number as a variable in Feature Constructor, but I cannot remember what the exact use case was ...
**What's your proposed solution?**
Make the row number available as a variable in Select Rows and Feature Constructor. Even better, allow use such as
`newvar := existingvar (row - 2) * othervar`
in Feature Constructor to refer to the value of a variable 2 rows back (which will of course not work for the first two rows
**Are there any alternative solutions?**
Yes:
- use a Python Script as suggested [here](https://discord.com/channels/633376992607076354/822470786346516501/940962165001695242) or
- abuse Melt and Group By as suggested [here](https://discord.com/channels/633376992607076354/822470786346516501/1016643062480511056)
| closed | 2023-01-24T14:22:43Z | 2023-01-25T21:11:59Z | https://github.com/biolab/orange3/issues/6312 | [] | wvdvegte | 5 |
iperov/DeepFaceLab | deep-learning | 5,609 | NEW to DFL wanna try it with the DFL, it works fine with 3090 but cant run SAEHD with 4080 HELP! Can run with QUICK tho | Running trainer.
[new] No saved models found. Enter a name of a new model :
new
Model first run.
Choose one or several GPU idxs (separated by comma).
[CPU] : CPU
[0] : NVIDIA GeForce RTX 4080
[0] Which GPU indexes to choose? :
0
[0] Autobackup every N hour ( 0..24 ?:help ) :
0
[n] Write preview history ( y/n ?:help ) :
n
[0] Target iteration :
0
[n] Flip SRC faces randomly ( y/n ?:help ) :
n
[y] Flip DST faces randomly ( y/n ?:help ) :
y
[8] Batch_size ( ?:help ) : 12
12
[128] Resolution ( 64-640 ?:help ) : 320
320
[f] Face type ( h/mf/f/wf/head ?:help ) : wf
wf
[liae-ud] AE architecture ( ?:help ) :
liae-ud
[256] AutoEncoder dimensions ( 32-1024 ?:help ) :
256
[64] Encoder dimensions ( 16-256 ?:help ) :
64
[64] Decoder dimensions ( 16-256 ?:help ) :
64
[22] Decoder mask dimensions ( 16-256 ?:help ) :
22
[y] Masked training ( y/n ?:help ) :
y
[n] Eyes and mouth priority ( y/n ?:help ) :
n
[n] Uniform yaw distribution of samples ( y/n ?:help ) :
n
[n] Blur out mask ( y/n ?:help ) :
n
[y] Place models and optimizer on GPU ( y/n ?:help ) :
y
[y] Use AdaBelief optimizer? ( y/n ?:help ) :
y
[n] Use learning rate dropout ( n/y/cpu ?:help ) :
n
[y] Enable random warp of samples ( y/n ?:help ) :
y
[0.0] Random hue/saturation/light intensity ( 0.0 .. 0.3 ?:help ) :
0.0
[0.0] GAN power ( 0.0 .. 5.0 ?:help ) :
0.0
[0.0] Face style power ( 0.0..100.0 ?:help ) :
0.0
[0.0] Background style power ( 0.0..100.0 ?:help ) :
0.0
[none] Color transfer for src faceset ( none/rct/lct/mkl/idt/sot ?:help ) :
none
[n] Enable gradient clipping ( y/n ?:help ) :
n
[n] Enable pretraining mode ( y/n ?:help ) :
n
Initializing models: 100%|###############################################################| 5/5 [00:01<00:00, 3.10it/s]
Loading samples: 100%|############################################################| 1053/1053 [00:04<00:00, 231.28it/s]
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "multiprocessing\spawn.py", line 105, in spawn_main
File "multiprocessing\spawn.py", line 115, in _main
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\__init__.py", line 1, in <module>
ImportError: numpy.core.multiarray failed to import
Traceback (most recent call last):
Traceback (most recent call last):
File "<string>", line 1, in <module>
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "multiprocessing\spawn.py", line 105, in spawn_main
Traceback (most recent call last):
File "multiprocessing\spawn.py", line 115, in _main
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "multiprocessing\spawn.py", line 105, in spawn_main
File "multiprocessing\spawn.py", line 115, in _main
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\__init__.py", line 1, in <module>
from .Sample import Sample
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\Sample.py", line 4, in <module>
ImportError: numpy.core.multiarray failed to import
Traceback (most recent call last):
File "<string>", line 1, in <module>
ImportError: numpy.core.multiarray failed to import
File "multiprocessing\spawn.py", line 105, in spawn_main
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "multiprocessing\spawn.py", line 115, in _main
File "multiprocessing\spawn.py", line 105, in spawn_main
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\__init__.py", line 1, in <module>
ImportError: numpy.core.multiarray failed to import
File "multiprocessing\spawn.py", line 115, in _main
from .Sample import Sample
Traceback (most recent call last):
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\Sample.py", line 4, in <module>
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\__init__.py", line 1, in <module>
File "<string>", line 1, in <module>
from .Sample import Sample
File "multiprocessing\spawn.py", line 105, in spawn_main
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\Sample.py", line 4, in <module>
File "multiprocessing\spawn.py", line 115, in _main
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\__init__.py", line 1, in <module>
from .Sample import Sample
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\Sample.py", line 4, in <module>
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "multiprocessing\spawn.py", line 105, in spawn_main
File "multiprocessing\spawn.py", line 115, in _main
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\__init__.py", line 1, in <module>
from .Sample import Sample
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\Sample.py", line 7, in <module>
from core.cv2ex import *
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\core\cv2ex.py", line 5, in <module>
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "multiprocessing\spawn.py", line 105, in spawn_main
File "multiprocessing\spawn.py", line 115, in _main
File "multiprocessing\heap.py", line 55, in __setstate__
OSError: [WinError 1455] The paging file is too small for this operation to complete
from core import imagelib
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\core\imagelib\__init__.py", line 9, in <module>
Process Process-18:
Process Process-16:
Traceback (most recent call last):
Traceback (most recent call last):
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\SampleGeneratorFace.py", line 134, in batch_func
x, = SampleProcessor.process ([sample], self.sample_process_options, self.output_sample_types, self.debug, ct_sample=ct_sample)
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\SampleGeneratorFace.py", line 134, in batch_func
x, = SampleProcessor.process ([sample], self.sample_process_options, self.output_sample_types, self.debug, ct_sample=ct_sample)
from .morph import morph_by_points File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\SampleProcessor.py", line 113, in process
warp_rnd_state=warp_rnd_state,
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\SampleProcessor.py", line 56, in process
sample_bgr = sample.load_bgr()
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\core\imagelib\warp.py", line 133, in gen_warp_params
mapy = cv2.resize(mapy, (w+cell_size,)*2 )[half_cell_size:-half_cell_size,half_cell_size:-half_cell_size].astype(np.float32)
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\Sample.py", line 112, in load_bgr
img = cv2_imread (self.filename, loader_func=self.read_raw_file).astype(np.float32) / 255.0
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\core\imagelib\morph.py", line 3, in <module>
MemoryError: Unable to allocate 400. KiB for an array with shape (320, 320) and data type float32
MemoryError: Unable to allocate 3.00 MiB for an array with shape (512, 512, 3) and data type float32
During handling of the above exception, another exception occurred:
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
Traceback (most recent call last):
File "multiprocessing\process.py", line 258, in _bootstrap
File "multiprocessing\process.py", line 258, in _bootstrap
File "multiprocessing\process.py", line 93, in run
File "multiprocessing\process.py", line 93, in run
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\core\joblib\SubprocessGenerator.py", line 54, in process_func
gen_data = next (self.generator_func)
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\core\joblib\SubprocessGenerator.py", line 54, in process_func
gen_data = next (self.generator_func)
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\SampleGeneratorFace.py", line 136, in batch_func
raise Exception ("Exception occured in sample %s. Error: %s" % (sample.filename, traceback.format_exc() ) )
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\SampleGeneratorFace.py", line 136, in batch_func
raise Exception ("Exception occured in sample %s. Error: %s" % (sample.filename, traceback.format_exc() ) )
Exception: Exception occured in sample F:\DeepFaceLab_NVIDIA_RTX3000_series\workspace\data_src\aligned\00731_0.jpg. Error: Traceback (most recent call last):
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\SampleGeneratorFace.py", line 134, in batch_func
x, = SampleProcessor.process ([sample], self.sample_process_options, self.output_sample_types, self.debug, ct_sample=ct_sample)
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\SampleProcessor.py", line 113, in process
warp_rnd_state=warp_rnd_state,
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\core\imagelib\warp.py", line 133, in gen_warp_params
mapy = cv2.resize(mapy, (w+cell_size,)*2 )[half_cell_size:-half_cell_size,half_cell_size:-half_cell_size].astype(np.float32)
MemoryError: Unable to allocate 400. KiB for an array with shape (320, 320) and data type float32
Exception: Exception occured in sample F:\DeepFaceLab_NVIDIA_RTX3000_series\workspace\data_src\aligned\01047_0.jpg. Error: Traceback (most recent call last):
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\SampleGeneratorFace.py", line 134, in batch_func
x, = SampleProcessor.process ([sample], self.sample_process_options, self.output_sample_types, self.debug, ct_sample=ct_sample)
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\SampleProcessor.py", line 56, in process
sample_bgr = sample.load_bgr()
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\DeepFaceLab\samplelib\Sample.py", line 112, in load_bgr
img = cv2_imread (self.filename, loader_func=self.read_raw_file).astype(np.float32) / 255.0
MemoryError: Unable to allocate 3.00 MiB for an array with shape (512, 512, 3) and data type float32
from scipy.spatial import Delaunay
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\python-3.6.8\lib\site-packages\scipy\spatial\__init__.py", line 97, in <module>
from .kdtree import *
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\python-3.6.8\lib\site-packages\scipy\spatial\kdtree.py", line 8, in <module>
import scipy.sparse
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\python-3.6.8\lib\site-packages\scipy\sparse\__init__.py", line 232, in <module>
from .lil import *
File "F:\DeepFaceLab_NVIDIA_RTX3000_series\_internal\python-3.6.8\lib\site-packages\scipy\sparse\lil.py", line 20, in <module>
from . import _csparsetools
ImportError: DLL load failed: The paging file is too small for this operation to complete.
| open | 2023-01-09T13:54:26Z | 2023-06-08T23:08:08Z | https://github.com/iperov/DeepFaceLab/issues/5609 | [] | kknpnfrom | 4 |
ultralytics/yolov5 | machine-learning | 12,828 | Batch Inference with Fine-tuned YOLOv5x6 Model on Custom Data | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
How can I perform batch inference to speed up image processing with a YOLOv5x6 model that has been fine-tuned on custom data?
### Additional
_No response_ | closed | 2024-03-19T08:12:10Z | 2024-10-20T19:41:45Z | https://github.com/ultralytics/yolov5/issues/12828 | [
"question"
] | Bycqg | 8 |
mckinsey/vizro | plotly | 571 | Connecting Backend Paging with a Graph | ### Question
Hey team,
I have a question regarding the use of Connecting Backend Paging with a Graph.
Based on the filtering of the table graph needs to be change, How we can achieve the same by using vizro.
Thank you so much for the amazing work on this tool, and for the help in advance!
```
from dash import Dash, dash_table, dcc, html, Input, Output, callback
import pandas as pd
app = Dash(__name__)
df = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/gapminder2007.csv')
PAGE_SIZE = 5
app.layout = html.Div(
className="row",
children=[
html.Div(
dash_table.DataTable(
id='table-paging-with-graph',
columns=[
{"name": i, "id": i} for i in sorted(df.columns)
],
page_current=0,
page_size=20,
page_action='custom',
filter_action='custom',
filter_query='',
sort_action='custom',
sort_mode='multi',
sort_by=[]
),
style={'height': 750, 'overflowY': 'scroll'},
className='six columns'
),
html.Div(
id='table-paging-with-graph-container',
className="five columns"
)
]
)
operators = [['ge ', '>='],
['le ', '<='],
['lt ', '<'],
['gt ', '>'],
['ne ', '!='],
['eq ', '='],
['contains '],
['datestartswith ']]
def split_filter_part(filter_part):
for operator_type in operators:
for operator in operator_type:
if operator in filter_part:
name_part, value_part = filter_part.split(operator, 1)
name = name_part[name_part.find('{') + 1: name_part.rfind('}')]
value_part = value_part.strip()
v0 = value_part[0]
if (v0 == value_part[-1] and v0 in ("'", '"', '`')):
value = value_part[1: -1].replace('\\' + v0, v0)
else:
try:
value = float(value_part)
except ValueError:
value = value_part
# word operators need spaces after them in the filter string,
# but we don't want these later
return name, operator_type[0].strip(), value
return [None] * 3
@callback(
Output('table-paging-with-graph', "data"),
Input('table-paging-with-graph', "page_current"),
Input('table-paging-with-graph', "page_size"),
Input('table-paging-with-graph', "sort_by"),
Input('table-paging-with-graph', "filter_query"))
def update_table(page_current, page_size, sort_by, filter):
filtering_expressions = filter.split(' && ')
dff = df
for filter_part in filtering_expressions:
col_name, operator, filter_value = split_filter_part(filter_part)
if operator in ('eq', 'ne', 'lt', 'le', 'gt', 'ge'):
# these operators match pandas series operator method names
dff = dff.loc[getattr(dff[col_name], operator)(filter_value)]
elif operator == 'contains':
dff = dff.loc[dff[col_name].str.contains(filter_value)]
elif operator == 'datestartswith':
# this is a simplification of the front-end filtering logic,
# only works with complete fields in standard format
dff = dff.loc[dff[col_name].str.startswith(filter_value)]
if len(sort_by):
dff = dff.sort_values(
[col['column_id'] for col in sort_by],
ascending=[
col['direction'] == 'asc'
for col in sort_by
],
inplace=False
)
return dff.iloc[
page_current*page_size: (page_current + 1)*page_size
].to_dict('records')
@callback(
Output('table-paging-with-graph-container', "children"),
Input('table-paging-with-graph', "data"))
def update_graph(rows):
dff = pd.DataFrame(rows)
return html.Div(
[
dcc.Graph(
id=column,
figure={
"data": [
{
"x": dff["country"],
"y": dff[column] if column in dff else [],
"type": "bar",
"marker": {"color": "#0074D9"},
}
],
"layout": {
"xaxis": {"automargin": True},
"yaxis": {"automargin": True},
"height": 250,
"margin": {"t": 10, "l": 10, "r": 10},
},
},
)
for column in ["pop", "lifeExp", "gdpPercap"]
]
)
if __name__ == '__main__':
app.run(debug=True)
```
[https://dash.plotly.com/datatable/callbacks](url)
### Code/Examples
_No response_
### Other information
_No response_
### Which package?
None
### Package version
_No response_
### Python version
_No response_
### OS
_No response_
### Code of Conduct
- [X] I agree to follow the [Code of Conduct](https://github.com/mckinsey/vizro/blob/main/CODE_OF_CONDUCT.md). | open | 2024-07-08T05:53:08Z | 2024-07-10T16:33:56Z | https://github.com/mckinsey/vizro/issues/571 | [
"General Question :question:"
] | BalaNagendraReddy | 2 |
Evil0ctal/Douyin_TikTok_Download_API | fastapi | 23 | 国际Tiktok 下载的是720p, 可以下1080p 吗? | 国际Tiktok 下载的是720p, 可以下1080p 吗? | closed | 2022-05-05T16:50:45Z | 2022-11-09T21:10:24Z | https://github.com/Evil0ctal/Douyin_TikTok_Download_API/issues/23 | [
"Fixed"
] | EddyN8 | 13 |
litestar-org/polyfactory | pydantic | 77 | Bug: pytest plugin causing runtime error | Looks like introduced in #74
```
app_1 | File "/app/./app/main.py", line 14, in <module>
app_1 | from starlite import Provide, Starlite
app_1 | File "/usr/local/lib/python3.10/site-packages/starlite/__init__.py", line 1, in <module>
app_1 | from starlite.app import Starlite
app_1 | File "/usr/local/lib/python3.10/site-packages/starlite/app.py", line 11, in <module>
app_1 | from starlite.asgi import (
app_1 | File "/usr/local/lib/python3.10/site-packages/starlite/asgi.py", line 33, in <module>
app_1 | from starlite.utils import AsyncCallable
app_1 | File "/usr/local/lib/python3.10/site-packages/starlite/utils/__init__.py", line 8, in <module>
app_1 | from .model import convert_dataclass_to_model, create_parsed_model_field
app_1 | File "/usr/local/lib/python3.10/site-packages/starlite/utils/model.py", line 5, in <module>
app_1 | from pydantic_factories.utils import create_model_from_dataclass
app_1 | File "/usr/local/lib/python3.10/site-packages/pydantic_factories/__init__.py", line 11, in <module>
app_1 | from .plugins import register_fixture
app_1 | File "/usr/local/lib/python3.10/site-packages/pydantic_factories/plugins/__init__.py", line 1, in <module>
app_1 | from .pytest_plugin import register_fixture
app_1 | File "/usr/local/lib/python3.10/site-packages/pydantic_factories/plugins/pytest_plugin.py", line 6, in <module>
app_1 | import pytest
app_1 | ModuleNotFoundError: No module named 'pytest'
``` | closed | 2022-10-08T12:31:24Z | 2022-10-08T16:45:23Z | https://github.com/litestar-org/polyfactory/issues/77 | [] | peterschutt | 0 |
huggingface/datasets | deep-learning | 7,440 | IterableDataset raises FileNotFoundError instead of retrying | ### Describe the bug
In https://github.com/huggingface/datasets/issues/6843 it was noted that the streaming feature of `datasets` is highly susceptible to outages and doesn't back off for long (or even *at all*).
I was training a model while streaming SlimPajama and training crashed with a `FileNotFoundError`. I can only assume that this was due to a momentary outage considering the file in question, `train/chunk9/example_train_3889.jsonl.zst`, [exists like all other files in SlimPajama](https://huggingface.co/datasets/cerebras/SlimPajama-627B/blob/main/train/chunk9/example_train_3889.jsonl.zst).
```python
...
File "/miniconda3/envs/draft/lib/python3.11/site-packages/datasets/iterable_dataset.py", line 2226, in __iter__
for key, example in ex_iterable:
File "/miniconda3/envs/draft/lib/python3.11/site-packages/datasets/iterable_dataset.py", line 1499, in __iter__
for x in self.ex_iterable:
File "/miniconda3/envs/draft/lib/python3.11/site-packages/datasets/iterable_dataset.py", line 1067, in __iter__
yield from self._iter()
File "/miniconda3/envs/draft/lib/python3.11/site-packages/datasets/iterable_dataset.py", line 1231, in _iter
for key, transformed_example in iter_outputs():
File "/miniconda3/envs/draft/lib/python3.11/site-packages/datasets/iterable_dataset.py", line 1207, in iter_outputs
for i, key_example in inputs_iterator:
File "/miniconda3/envs/draft/lib/python3.11/site-packages/datasets/iterable_dataset.py", line 1111, in iter_inputs
for key, example in iterator:
File "/miniconda3/envs/draft/lib/python3.11/site-packages/datasets/iterable_dataset.py", line 371, in __iter__
for key, pa_table in self.generate_tables_fn(**gen_kwags):
File "/miniconda3/envs/draft/lib/python3.11/site-packages/datasets/packaged_modules/json/json.py", line 99, in _generate_tables
for file_idx, file in enumerate(itertools.chain.from_iterable(files)):
File "/miniconda3/envs/draft/lib/python3.11/site-packages/datasets/utils/track.py", line 50, in __iter__
for x in self.generator(*self.args):
File "/miniconda3/envs/draft/lib/python3.11/site-packages/datasets/utils/file_utils.py", line 1378, in _iter_from_urlpaths
raise FileNotFoundError(urlpath)
FileNotFoundError: zstd://example_train_3889.jsonl::hf://datasets/cerebras/SlimPajama-627B@2d0accdd58c5d5511943ca1f5ff0e3eb5e293543/train/chunk9/example_train_3889.jsonl.zst
```
That final `raise` is at the bottom of the following snippet:
https://github.com/huggingface/datasets/blob/f693f4e93aabafa878470c80fd42ddb10ec550d6/src/datasets/utils/file_utils.py#L1354-L1379
So clearly, something choked up in `xisfile`.
### Steps to reproduce the bug
This happens when streaming a dataset and iterating over it. In my case, that iteration is done in Trainer's `inner_training_loop`, but this is not relevant to the iterator.
```python
File "/miniconda3/envs/draft/lib/python3.11/site-packages/accelerate/data_loader.py", line 835, in __iter__
next_batch, next_batch_info = self._fetch_batches(main_iterator)
```
### Expected behavior
This bug and the linked issue have one thing in common: *when streaming fails to retrieve an example, the entire program gives up and crashes*. As users, we cannot even protect ourselves from this: when we are iterating over a dataset, we can't make `datasets` skip over a bad example or wait a little longer to retry the iteration, because when a Python generator/iterator raises an error, it loses all its context.
In other words: if you have something that looks like `for b in a: for c in b: for d in c:`, errors in the innermost loop can only be caught by a `try ... except` in `c.__iter__()`. There should be such exception handling in `datasets` and it should have a **configurable exponential back-off**: first wait and retry after 1 minute, then 2 minutes, then 4 minutes, then 8 minutes, ... and after a given amount of retries, **skip the bad example**, and **only after** skipping a given amount of examples, give up and crash. This was requested in https://github.com/huggingface/datasets/issues/6843 too, since currently there is only linear backoff *and* it is clearly not applied to `xisfile`.
### Environment info
- `datasets` version: 3.3.2 *(the latest version)*
- Platform: Linux-4.18.0-513.24.1.el8_9.x86_64-x86_64-with-glibc2.28
- Python version: 3.11.7
- `huggingface_hub` version: 0.26.5
- PyArrow version: 15.0.0
- Pandas version: 2.2.0
- `fsspec` version: 2024.10.0 | open | 2025-03-07T19:14:18Z | 2025-03-22T21:48:02Z | https://github.com/huggingface/datasets/issues/7440 | [] | bauwenst | 5 |
pydata/xarray | numpy | 10,169 | Grouping first and last on numpy datetime data with missing groups fails on flox | ### What happened?
I have a data array of datetime values and I want to get the first value for each group. If there are any missing groups, the operation fails as numpy can't promote datetime data to float.
This is new in xarray 2025.3.
### What did you expect to happen?
I expected to receive the first value for each group and NaT for missing groups.
### Minimal Complete Verifiable Example
```Python
import xarray as xr
# A datetime array
time = xr.DataArray(xr.date_range('2000-01-01', periods=400, freq='D'), dims=('time',))
# Remove december, so there is a missing group
time = time.sel(time=time.dt.month != 12)
time.resample(time='MS').first()
```
### MVCE confirmation
- [x] Minimal example — the example is as focused as reasonably possible to demonstrate the underlying issue in xarray.
- [x] Complete example — the example is self-contained, including all data and the text of any traceback.
- [x] Verifiable example — the example copy & pastes into an IPython prompt or [Binder notebook](https://mybinder.org/v2/gh/pydata/xarray/main?urlpath=lab/tree/doc/examples/blank_template.ipynb), returning the result.
- [x] New issue — a search of GitHub Issues suggests this is not a duplicate.
- [x] Recent environment — the issue occurs with the latest version of xarray and its dependencies.
### Relevant log output
```Python
---------------------------------------------------------------------------
DTypePromotionError Traceback (most recent call last)
Cell In[41], line 1
----> 1 time.resample(time='MS').first()
File ~/Projets/xarray/xarray/core/groupby.py:1420, in GroupBy.first(self, skipna, keep_attrs)
1401 def first(
1402 self, skipna: bool | None = None, keep_attrs: bool | None = None
1403 ) -> T_Xarray:
1404 """
1405 Return the first element of each group along the group dimension
1406
(...) 1418
1419 """
-> 1420 return self._first_or_last("first", skipna, keep_attrs)
File ~/Projets/xarray/xarray/core/resample.py:114, in Resample._first_or_last(self, op, skipna, keep_attrs)
109 def _first_or_last(
110 self, op: Literal["first", "last"], skipna: bool | None, keep_attrs: bool | None
111 ) -> T_Xarray:
112 from xarray.core.dataset import Dataset
--> 114 result = super()._first_or_last(op=op, skipna=skipna, keep_attrs=keep_attrs)
115 if isinstance(result, Dataset):
116 # Can't do this in the base class because group_dim is RESAMPLE_DIM
117 # which is not present in the original object
118 for var in result.data_vars:
File ~/Projets/xarray/xarray/core/groupby.py:1389, in GroupBy._first_or_last(self, op, skipna, keep_attrs)
1383 keep_attrs = _get_keep_attrs(default=True)
1384 if (
1385 module_available("flox", minversion="0.10.0")
1386 and OPTIONS["use_flox"]
1387 and contains_only_chunked_or_numpy(self._obj)
1388 ):
-> 1389 result = self._flox_reduce(
1390 dim=None, func=op, skipna=skipna, keep_attrs=keep_attrs
1391 )
1392 else:
1393 result = self.reduce(
1394 getattr(duck_array_ops, op),
1395 dim=[self._group_dim],
1396 skipna=skipna,
1397 keep_attrs=keep_attrs,
1398 )
File ~/Projets/xarray/xarray/core/resample.py:59, in Resample._flox_reduce(self, dim, keep_attrs, **kwargs)
52 def _flox_reduce(
53 self,
54 dim: Dims,
55 keep_attrs: bool | None = None,
56 **kwargs,
57 ) -> T_Xarray:
58 result: T_Xarray = (
---> 59 super()
60 ._flox_reduce(dim=dim, keep_attrs=keep_attrs, **kwargs)
61 .rename({RESAMPLE_DIM: self._group_dim}) # type: ignore[assignment]
62 )
63 return result
File ~/Projets/xarray/xarray/core/groupby.py:1099, in GroupBy._flox_reduce(self, dim, keep_attrs, **kwargs)
1097 from IPython import embed
1098 embed()
-> 1099 result = xarray_reduce(
1100 obj.drop_vars(non_numeric.keys()),
1101 *codes,
1102 dim=parsed_dim,
1103 expected_groups=expected_groups,
1104 isbin=False,
1105 keep_attrs=keep_attrs,
1106 **kwargs,
1107 )
1109 # we did end up reducing over dimension(s) that are
1110 # in the grouped variable
1111 group_dims = set(grouper.group.dims)
File ~/miniforge3/envs/xclim-dev/lib/python3.13/site-packages/flox/xarray.py:410, in xarray_reduce(obj, func, expected_groups, isbin, sort, dim, fill_value, dtype, method, engine, keep_attrs, skipna, min_count, reindex, *by, **finalize_kwargs)
407 output_sizes = group_sizes
408 output_sizes.update({dim.name: dim.size for dim in newdims if dim.size != 0})
--> 410 actual = xr.apply_ufunc(
411 wrapper,
412 ds_broad.drop_vars(tuple(missing_dim)).transpose(..., *grouper_dims),
413 *by_da,
414 input_core_dims=input_core_dims,
415 # for xarray's test_groupby_duplicate_coordinate_labels
416 exclude_dims=set(dim_tuple),
417 output_core_dims=[output_core_dims],
418 dask="allowed",
419 dask_gufunc_kwargs=dict(
420 output_sizes=output_sizes,
421 output_dtypes=[dtype] if dtype is not None else None,
422 ),
423 keep_attrs=keep_attrs,
424 kwargs={
425 "func": func,
426 "axis": axis,
427 "sort": sort,
428 "fill_value": fill_value,
429 "method": method,
430 "min_count": min_count,
431 "skipna": skipna,
432 "engine": engine,
433 "reindex": reindex,
434 "expected_groups": tuple(expected_groups_valid_list),
435 "isbin": isbins,
436 "finalize_kwargs": finalize_kwargs,
437 "dtype": dtype,
438 "core_dims": input_core_dims,
439 },
440 )
442 # restore non-dim coord variables without the core dimension
443 # TODO: shouldn't apply_ufunc handle this?
444 for var in set(ds_broad._coord_names) - set(ds_broad._indexes) - set(ds_broad.dims):
File ~/Projets/xarray/xarray/computation/apply_ufunc.py:1255, in apply_ufunc(func, input_core_dims, output_core_dims, exclude_dims, vectorize, join, dataset_join, dataset_fill_value, keep_attrs, kwargs, dask, output_dtypes, output_sizes, meta, dask_gufunc_kwargs, on_missing_core_dim, *args)
1253 # feed datasets apply_variable_ufunc through apply_dataset_vfunc
1254 elif any(is_dict_like(a) for a in args):
-> 1255 return apply_dataset_vfunc(
1256 variables_vfunc,
1257 *args,
1258 signature=signature,
1259 join=join,
1260 exclude_dims=exclude_dims,
1261 dataset_join=dataset_join,
1262 fill_value=dataset_fill_value,
1263 keep_attrs=keep_attrs,
1264 on_missing_core_dim=on_missing_core_dim,
1265 )
1266 # feed DataArray apply_variable_ufunc through apply_dataarray_vfunc
1267 elif any(isinstance(a, DataArray) for a in args):
File ~/Projets/xarray/xarray/computation/apply_ufunc.py:526, in apply_dataset_vfunc(func, signature, join, dataset_join, fill_value, exclude_dims, keep_attrs, on_missing_core_dim, *args)
521 list_of_coords, list_of_indexes = build_output_coords_and_indexes(
522 args, signature, exclude_dims, combine_attrs=keep_attrs
523 )
524 args = tuple(getattr(arg, "data_vars", arg) for arg in args)
--> 526 result_vars = apply_dict_of_variables_vfunc(
527 func,
528 *args,
529 signature=signature,
530 join=dataset_join,
531 fill_value=fill_value,
532 on_missing_core_dim=on_missing_core_dim,
533 )
535 out: Dataset | tuple[Dataset, ...]
536 if signature.num_outputs > 1:
File ~/Projets/xarray/xarray/computation/apply_ufunc.py:450, in apply_dict_of_variables_vfunc(func, signature, join, fill_value, on_missing_core_dim, *args)
448 core_dim_present = _check_core_dims(signature, variable_args, name)
449 if core_dim_present is True:
--> 450 result_vars[name] = func(*variable_args)
451 else:
452 if on_missing_core_dim == "raise":
File ~/Projets/xarray/xarray/computation/apply_ufunc.py:821, in apply_variable_ufunc(func, signature, exclude_dims, dask, output_dtypes, vectorize, keep_attrs, dask_gufunc_kwargs, *args)
816 if vectorize:
817 func = _vectorize(
818 func, signature, output_dtypes=output_dtypes, exclude_dims=exclude_dims
819 )
--> 821 result_data = func(*input_data)
823 if signature.num_outputs == 1:
824 result_data = (result_data,)
File ~/miniforge3/envs/xclim-dev/lib/python3.13/site-packages/flox/xarray.py:367, in xarray_reduce.<locals>.wrapper(array, func, skipna, core_dims, *by, **kwargs)
364 if "nan" not in func and func not in ["all", "any", "count"]:
365 func = f"nan{func}"
--> 367 result, *groups = groupby_reduce(array, *by, func=func, **kwargs)
369 # Transpose the new quantile dimension to the end. This is ugly.
370 # but new core dimensions are expected at the end :/
371 # but groupby_reduce inserts them at the beginning
372 if func in ["quantile", "nanquantile"]:
File ~/miniforge3/envs/xclim-dev/lib/python3.13/site-packages/flox/core.py:2559, in groupby_reduce(array, func, expected_groups, sort, isbin, axis, fill_value, dtype, min_count, method, engine, reindex, finalize_kwargs, *by)
2556 fill_value = np.nan
2558 kwargs = dict(axis=axis_, fill_value=fill_value)
-> 2559 agg = _initialize_aggregation(func, dtype, array.dtype, fill_value, min_count_, finalize_kwargs)
2561 # Need to set this early using `agg`
2562 # It cannot be done in the core loop of chunk_reduce
2563 # since we "prepare" the data for flox.
2564 kwargs["engine"] = _choose_engine(by_, agg) if engine is None else engine
File ~/miniforge3/envs/xclim-dev/lib/python3.13/site-packages/flox/aggregations.py:809, in _initialize_aggregation(func, dtype, array_dtype, fill_value, min_count, finalize_kwargs)
804 # np.dtype(None) == np.dtype("float64")!!!
805 # so check for not None
806 dtype_: np.dtype | None = (
807 np.dtype(dtype) if dtype is not None and not isinstance(dtype, np.dtype) else dtype
808 )
--> 809 final_dtype = dtypes._normalize_dtype(
810 dtype_ or agg.dtype_init["final"], array_dtype, agg.preserves_dtype, fill_value
811 )
812 agg.dtype = {
813 "user": dtype, # Save to automatically choose an engine
814 "final": final_dtype,
(...) 823 ),
824 }
826 # Replace sentinel fill values according to dtype
File ~/miniforge3/envs/xclim-dev/lib/python3.13/site-packages/flox/xrdtypes.py:171, in _normalize_dtype(dtype, array_dtype, preserves_dtype, fill_value)
169 dtype = np.dtype(dtype)
170 if fill_value not in [None, INF, NINF, NA]:
--> 171 dtype = np.result_type(dtype, fill_value)
172 return dtype
DTypePromotionError: The DType <class 'numpy.dtypes.DateTime64DType'> could not be promoted by <class 'numpy.dtypes._PyFloatDType'>. This means that no common DType exists for the given inputs. For example they cannot be stored in a single array unless the dtype is `object`. The full list of DTypes is: (<class 'numpy.dtypes.DateTime64DType'>, <class 'numpy.dtypes._PyFloatDType'>)
```
### Anything else we need to know?
This is was introduced by #10148 I believe, but the reason is that the `_flox_reduce` method assumes `np.nan` as a fill value when groups are missing:
https://github.com/pydata/xarray/blob/4174aa1d6104bc853bdf4de08019194c9eececc0/xarray/core/groupby.py#L1105
It also fails for cftime data, but I think this is another issue internal to flox.
Deactivating flox fixes both issues ( `xr.set_options(use_flox=False)`).
### Environment
<details>
INSTALLED VERSIONS
------------------
commit: fd7c76562ff70e07d63ff03808b4f87a62955bcd
python: 3.13.2 | packaged by conda-forge | (main, Feb 17 2025, 14:10:22) [GCC 13.3.0]
python-bits: 64
OS: Linux
OS-release: 6.12.12-200.fc41.x86_64
machine: x86_64
processor:
byteorder: little
LC_ALL: None
LANG: fr_CA.UTF-8
LOCALE: ('fr_CA', 'UTF-8')
libhdf5: 1.14.4
libnetcdf: 4.9.2
xarray: 2025.3.1.dev5+gfd7c7656.d20250324
pandas: 2.2.3
numpy: 2.1.3
scipy: 1.15.2
netCDF4: 1.7.2
pydap: None
h5netcdf: 1.6.1
h5py: 3.12.1
zarr: None
cftime: 1.6.4
nc_time_axis: 1.4.1
iris: None
bottleneck: 1.4.2
dask: 2025.3.0
distributed: 2025.3.0
matplotlib: 3.10.1
cartopy: None
seaborn: None
numbagg: None
fsspec: 2025.3.0
cupy: None
pint: 0.24.4
sparse: None
flox: 0.10.0
numpy_groupies: 0.11.2
setuptools: 75.8.2
pip: 25.0.1
conda: None
pytest: 8.3.5
mypy: 1.15.0
IPython: 9.0.2
sphinx: 8.1.
</details>
| open | 2025-03-24T16:10:37Z | 2025-03-24T17:15:12Z | https://github.com/pydata/xarray/issues/10169 | [
"bug",
"topic-groupby"
] | aulemahal | 0 |
Avaiga/taipy | data-visualization | 1,922 | Make a navbar to be sticky | ### What went wrong? 🤔
The navbar goes away after scrolling down

### Expected Behavior
The navbar should be fixed at the top of the page

### Steps to Reproduce Issue
_No response_
### Solution Proposed
_No response_
### Screenshots
_No response_
### Runtime Environment
_No response_
### Browsers
Chrome
### OS
Windows
### Version of Taipy
_No response_
### Additional Context
_No response_
### Acceptance Criteria
- [ ] Ensure new code is unit tested, and check code coverage is at least 90%.
- [ ] Create related issue in taipy-doc for documentation and Release Notes.
### Code of Conduct
- [X] I have checked the [existing issues](https://github.com/Avaiga/taipy/issues?q=is%3Aissue+).
- [X] I am willing to work on this issue (optional) | closed | 2024-10-04T13:40:36Z | 2024-10-05T01:24:53Z | https://github.com/Avaiga/taipy/issues/1922 | [
"💥Malfunction"
] | MOHDNEHALKHAN | 3 |
pytorch/vision | machine-learning | 8,224 | `pr-labels `job is failing on CI | Our "PR label bot" hasn't been commenting on PRs for a bit. Looks like it's broken https://github.com/pytorch/vision/actions/workflows/pr-labels.yml
```
Run mshick/add-pr-comment@v1
with:
message: Hey @NicolasHug!
You merged this PR, but no labels were added. The list of valid labels is available at https://github.com/pytorch/vision/blob/main/.github/process_commit.py
allow-repeats: false
env:
pythonLocation: /opt/hostedtoolcache/Python/3.12.1/x64
LD_LIBRARY_PATH: /opt/hostedtoolcache/Python/3.12.1/x64/lib
GITHUB_TOKEN: ***
Error: Resource not accessible by integration
```
(Note that https://github.com/pytorch/vision/pull/8221 is related but the pr-labels action was broken before this was merged) | closed | 2024-01-19T11:38:32Z | 2024-02-14T11:10:38Z | https://github.com/pytorch/vision/issues/8224 | [] | NicolasHug | 4 |
flairNLP/flair | pytorch | 2,924 | How can I assure the best model is saved? | I'm confused by the parameters given in https://github.com/flairNLP/flair/blob/493b61fc07f83d412928301571b0e3abe780348f/flair/trainers/trainer.py#L74
and the conditions shown in https://github.com/flairNLP/flair/blob/493b61fc07f83d412928301571b0e3abe780348f/flair/trainers/trainer.py#L796
` if (
(not train_with_dev or anneal_with_restarts or anneal_with_prestarts)
and not param_selection_mode
and current_epoch_has_best_model_so_far
and not use_final_model_for_eval
):
log.info("saving best model")
self.model.save(base_path / "best-model.pt", checkpoint=save_optimizer_state)`
It seems that by default a best-model.pt should be saved, however I can only find a final-model.pt in my output folder
My fine_tune arguments are:
` trainer.fine_tune(
classifier_save_path,
learning_rate=0.05,
mini_batch_size=4,
mini_batch_chunk_size=None,
max_epochs=1,
train_with_dev=False,
checkpoint=False,
embeddings_storage_mode='none',
**kwargs
)`
As I am training large models I'm currently only testing with max_epochs=1, but I don't think that should prevent the best-model.pt from being saved? | closed | 2022-08-30T10:15:20Z | 2022-09-05T13:39:13Z | https://github.com/flairNLP/flair/issues/2924 | [
"question"
] | Flowhill | 2 |
alpacahq/alpaca-trade-api-python | rest-api | 59 | Request ability to cancel order by client order id | If we are able to tag our orders with a Client Order ID, it seems reasonable for a call like `api.cancel_order_by_client_order_id('my_custom_client_order_id')` to exist. I ask because it seems rather confusing to provide developers with a nice way of tagging orders internally, then forcing them to query for the proper id with a call to `get_order_by_client_order_id` if they chose to use that feature.
| closed | 2019-03-15T07:42:43Z | 2019-05-06T21:21:23Z | https://github.com/alpacahq/alpaca-trade-api-python/issues/59 | [] | ztaylor54 | 1 |
microsoft/Bringing-Old-Photos-Back-to-Life | pytorch | 10 | Using CPU instead of CUDA | Is it possible to use CPU or even OpenCL instead? CUDA is nvidia proprietary code and I have an AMD laptop. Thanks. | closed | 2020-09-21T03:07:12Z | 2020-09-24T18:03:43Z | https://github.com/microsoft/Bringing-Old-Photos-Back-to-Life/issues/10 | [] | jersobh | 2 |
axnsan12/drf-yasg | rest-api | 852 | Update MarkupSafe version | # Update MarkupSafe package
drf_yasg uses MarkupSafe==1.1.1 and it is really old. when I use a new version in my own project. I will encounter errors.
errors are about drf_yasg. I will be grateful if you can use a newer version in drf_yasg
| open | 2023-06-11T07:04:36Z | 2025-03-07T12:09:04Z | https://github.com/axnsan12/drf-yasg/issues/852 | [
"triage"
] | RasoulRostami | 0 |
joeyespo/grip | flask | 15 | Add authentication to increase GitHub API limit | GitHub imposes a limit of 60 unauthenticated requests per hour. This is really easy to hit when using grip, especially if you're doing other things using the API at the same time. Once your reach this limit, grip no longer functions (you see an error message about rate limiting when you try to render a page).
I'd like to add options to grip to specify GitHub username and password so that grip can perform Basic auth in renderer.py. This bumps the limit to 5000 requests/hour which should be enough for anyone :)
Any thoughts on this change?
I guess I see these being new command line options.
| closed | 2013-07-06T18:55:28Z | 2014-02-04T17:44:10Z | https://github.com/joeyespo/grip/issues/15 | [
"enhancement"
] | joelittlejohn | 2 |
mlflow/mlflow | machine-learning | 14,841 | [FR] way to bookmark/share a UI view with `Search Experiments` text box populated | ### Willingness to contribute
No. I cannot contribute this feature at this time.
### Proposal Summary
When there are man experiments accumulated in the sidebar/database, it would be useful to be able to bookmark a view where the search field to filter the list of visible experiments is pre-populated.
As far as I can tell, this isn't really possible using currently available query parameters in the URL.
### Motivation
> #### What is the use case for this feature?
Automatically filter long list of experiment names to the experiments you're currently working on with out having to manually enter a search string into the filter box each time you connect to the server.
> #### Why is this use case valuable to support for MLflow users in general?
Less clutter in the UI, fewer steps to get to relevant data, etc.
> #### Why is this use case valuable to support for your project(s) or organization?
Same reasons
> #### Why is it currently difficult to achieve this use case?
You can limit the experiment list, but only by manually going to the filter box each time you open the UI and typing in a search string. It's just often inconvenient when you're working on multiple projects.
### Details
_No response_
### What component(s) does this bug affect?
- [ ] `area/artifacts`: Artifact stores and artifact logging
- [ ] `area/build`: Build and test infrastructure for MLflow
- [ ] `area/deployments`: MLflow Deployments client APIs, server, and third-party Deployments integrations
- [ ] `area/docs`: MLflow documentation pages
- [ ] `area/examples`: Example code
- [ ] `area/model-registry`: Model Registry service, APIs, and the fluent client calls for Model Registry
- [ ] `area/models`: MLmodel format, model serialization/deserialization, flavors
- [ ] `area/recipes`: Recipes, Recipe APIs, Recipe configs, Recipe Templates
- [ ] `area/projects`: MLproject format, project running backends
- [ ] `area/scoring`: MLflow Model server, model deployment tools, Spark UDFs
- [ ] `area/server-infra`: MLflow Tracking server backend
- [ ] `area/tracking`: Tracking Service, tracking client APIs, autologging
### What interface(s) does this bug affect?
- [x] `area/uiux`: Front-end, user experience, plotting, JavaScript, JavaScript dev server
- [ ] `area/docker`: Docker use across MLflow's components, such as MLflow Projects and MLflow Models
- [ ] `area/sqlalchemy`: Use of SQLAlchemy in the Tracking Service or Model Registry
- [ ] `area/windows`: Windows support
### What language(s) does this bug affect?
- [ ] `language/r`: R APIs and clients
- [ ] `language/java`: Java APIs and clients
- [ ] `language/new`: Proposals for new client languages
### What integration(s) does this bug affect?
- [ ] `integrations/azure`: Azure and Azure ML integrations
- [ ] `integrations/sagemaker`: SageMaker integrations
- [ ] `integrations/databricks`: Databricks integrations | open | 2025-03-04T19:17:59Z | 2025-03-15T14:37:57Z | https://github.com/mlflow/mlflow/issues/14841 | [
"enhancement",
"area/uiux",
"help wanted"
] | mazer-ai | 5 |
mkhorasani/Streamlit-Authenticator | streamlit | 166 | Logout takes a lot of time when there is more data in the application's session state. | I have implemented the authentication module in my application and it works great. The only thing is Logout takes an ample amount of time when there is a significant amount of session data. This is because of streamlit's top-down execution model whenever there is some user interaction. Moreover, it is better to delete all the session key data on the click of logout instead of just the session keys set by the streamlit authenticator module.
I have a solution and would like to contribute by creating the pull request.
Thank you so much for creating such a great and easy-to-use authentication module for streamlit. Keep it up.
| closed | 2024-05-21T15:08:00Z | 2024-05-21T15:20:13Z | https://github.com/mkhorasani/Streamlit-Authenticator/issues/166 | [] | harsh9898 | 1 |
strawberry-graphql/strawberry | django | 3,809 | Automatic object type resolution does not trigger in reference resolvers | <!-- Provide a general summary of the bug in the title above. -->
<!--- This template is entirely optional and can be removed, but is here to help both you and us. -->
<!--- Anything on lines wrapped in comments like these will not show up in the final text. -->
## Describe the Bug
<!-- A clear and concise description of what the bug is. -->
This is a bit of a tricky one to explain, so do let me know if any further clarity is needed.
TL;DR: Strawberry is unable to resolve federated types where the reference resolver does not explicitly return an object type, or where the type being federated defines inline field resolvers.
It's easier to explain with an MRE. This system has two services, `groups` and `users`, each pertaining to group and user queries respectively. The `Group` type is federated, and the `users` service has two queries. `Users` contains `group` as a field.
groups/app.py:
```py
from types import SimpleNamespace
from typing import Self
import strawberry
groups = {
"1": SimpleNamespace(id="1", name="Hello", altname="Hey"),
"2": SimpleNamespace(id="2", name="Strawberry"),
"3": SimpleNamespace(id="3", name="World", altname="Earth"),
}
@strawberry.federation.type(keys=["id"])
class Group:
id: strawberry.ID
name: str
altname: str = strawberry.field(
resolver=lambda root: getattr(root, "altname", root.name),
)
@classmethod
def resolve_reference(cls, id: str) -> Self:
return groups.get(id)
schema = strawberry.federation.Schema(
types=[Group],
enable_federation_2=True,
)
```
users/app.py:
```py
from types import SimpleNamespace
import strawberry
users = {
"1": SimpleNamespace(id="1", group_id="1"),
"2": SimpleNamespace(id="2", group_id="2"),
"3": SimpleNamespace(id="3", group_id="3"),
}
@strawberry.federation.type(keys=["id"])
class Group:
id: strawberry.ID
@strawberry.type
class User:
id: int
group: Group = strawberry.field(
resolver=lambda root: Group(id=root.group_id),
)
@strawberry.type
class Query:
@strawberry.field
def users(self) -> list[User]:
return list(users.values())
@strawberry.field
def user(self) -> User:
return users.get("1")
schema = strawberry.federation.Schema(
query=Query,
enable_federation_2=True,
)
```
Posting the following query (`altname` is intentionally omitted for now):
```json
{"query": "query { users { id group { id name } } }"
```
returns the following error in the `groups` service:
```
GraphQL request:1:37
1 | query($representations: [_Any!]!) { _entities(representations: $representations) { ... on Group { name } } }
| ^
Traceback (most recent call last):
File "/Users/ethanhenderson/Programs/Playground/strawberry_test/.venv/lib/python3.13/site-packages/graphql/execution/execute.py", line 728, in complete_list_value
completed_item = self.complete_value(
item_type, field_nodes, info, item_path, item
)
File "/Users/ethanhenderson/Programs/Playground/strawberry_test/.venv/lib/python3.13/site-packages/graphql/execution/execute.py", line 646, in complete_value
return self.complete_abstract_value(
~~~~~~~~~~~~~~~~~~~~~~~~~~~~^
cast(GraphQLAbstractType, return_type), field_nodes, info, path, result
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
)
^
File "/Users/ethanhenderson/Programs/Playground/strawberry_test/.venv/lib/python3.13/site-packages/graphql/execution/execute.py", line 798, in complete_abstract_value
runtime_type = resolve_type_fn(result, info, return_type)
File "/Users/ethanhenderson/Programs/Playground/strawberry_test/.venv/lib/python3.13/site-packages/strawberry/types/union.py", line 185, in _resolve_union_type
raise WrongReturnTypeForUnion(info.field_name, str(type(root)))
strawberry.exceptions.WrongReturnTypeForUnion: The type "<class 'types.SimpleNamespace'>" cannot be resolved for the field "_entities" , are you using a strawberry.field?
```
This error can be resolved by explicitly returning a `Group` object type, like so:
```py
@classmethod
def resolve_reference(cls, id: str) -> Self:
group = groups.get(id)
return Group(id=group.id, name=group.name)
```
However, when querying `altname` as well, which uses an inline resolver:
```json
{"query": "query { users { id group { id name altname } } }"
```
The `groups` service raises this error instead:
```
GraphQL request:1:104
1 | query($representations: [_Any!]!) { _entities(representations: $representations) { ... on Group { name altname } } }
| ^
Traceback (most recent call last):
File "/Users/ethanhenderson/Programs/Playground/strawberry_test/.venv/lib/python3.13/site-packages/graphql/execution/execute.py", line 542, in execute_field
completed = self.complete_value(
return_type, field_nodes, info, path, result
)
File "/Users/ethanhenderson/Programs/Playground/strawberry_test/.venv/lib/python3.13/site-packages/graphql/execution/execute.py", line 614, in complete_value
completed = self.complete_value(
cast(GraphQLNonNull, return_type).of_type,
...<3 lines>...
result,
)
File "/Users/ethanhenderson/Programs/Playground/strawberry_test/.venv/lib/python3.13/site-packages/graphql/execution/execute.py", line 641, in complete_value
return self.complete_leaf_value(cast(GraphQLLeafType, return_type), result)
~~~~~~~~~~~~~~~~~~~~~~~~^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/ethanhenderson/Programs/Playground/strawberry_test/.venv/lib/python3.13/site-packages/graphql/execution/execute.py", line 776, in complete_leaf_value
serialized_result = return_type.serialize(result)
File "/Users/ethanhenderson/Programs/Playground/strawberry_test/.venv/lib/python3.13/site-packages/graphql/type/scalars.py", line 177, in serialize_string
raise GraphQLError("String cannot represent value: " + inspect(output_value))
graphql.error.graphql_error.GraphQLError: String cannot represent value: <method>
```
`altname` can't be passed as an argument to the constructor as it has an inline resolver, and it can't be resolved when being federated from another type as Strawberry doesn't perform the necessary resolution. This makes it very difficult to use inline resolvers in federated types.
If there's another way of doing this I'm missing, do let me know.
## System Information
- Operating system: MacOS 15.3.2
- Strawberry version (if applicable): 0.262.3
## Additional Context
The MRE is based on the setup in the [Federation v2 Guide](https://strawberry.rocks/docs/guides/federation#apollo-federation-2-guide) to try and make it simpler.
| open | 2025-03-13T16:14:33Z | 2025-03-13T16:21:00Z | https://github.com/strawberry-graphql/strawberry/issues/3809 | [
"bug"
] | parafoxia | 0 |
wkentaro/labelme | deep-learning | 485 | I want to do a multi-category data set, where the same category in the data set has the same color,how can i do?thank you very much! | closed | 2019-09-16T02:52:22Z | 2019-09-19T12:44:50Z | https://github.com/wkentaro/labelme/issues/485 | [] | leedoge | 1 | |
jupyter/nbgrader | jupyter | 1,107 | Release nbgrader 0.6.0 | I am planning to make a release for 0.6.0 after we get in the changes from the hackathon, so hopefully within the next few weeks!
If there is anything you'd really like to see in 0.6.0 that is currently not marked for 0.6.0, please let me know. (However, if it's a major change it's unlikely to be something that will make it in as I'd rather not keep pushing the release off). | closed | 2019-05-30T23:11:44Z | 2022-07-13T15:17:11Z | https://github.com/jupyter/nbgrader/issues/1107 | [
"maintenance"
] | jhamrick | 2 |
OpenInterpreter/open-interpreter | python | 1,228 | ollama llama3 How to remove the first line " ` " when generating code in Windows 11 terminal | ### Describe the bug
ollama llama3 "`" symbols frequently appear in the first line when generating code in Windows 11 terminal
### Reproduce
ollama llama3 "`" symbols frequently appear in the first line when generating code in Windows 11 terminal
### Expected behavior
ollama llama3 When generating code in Windows 11 terminal, the "`" symbol should not appear in the first line
### Screenshots
![Uploading 屏幕截图 2024-04-23 212337.png…]()
### Open Interpreter version
0.2.4
### Python version
3.11.9
### Operating System name and version
windows11
### Additional context
open interpreter This error occurs when running ollama llama3 | closed | 2024-04-23T13:37:39Z | 2024-04-23T23:42:48Z | https://github.com/OpenInterpreter/open-interpreter/issues/1228 | [] | ltsyk | 3 |
mljar/mercury | data-visualization | 436 | Disable page refresh on disconnect? | Is there an option to prevent page refresh on disconnect? Currently, mercury refreshes the page when it loses connection to the server.
For my use case, I want to persist whatever is on the page if the connection gets lost.
It would also be nice to not "gray-out" the screen.
Use case: a page takes a long time to render and there's no need to refresh it afterwards. The server is flaky and loses connection sometime and I don't want to lose the contents of the webpage when the connection the server is bad | open | 2024-03-25T19:28:16Z | 2024-03-25T21:07:59Z | https://github.com/mljar/mercury/issues/436 | [] | kapily | 1 |
litestar-org/polyfactory | pydantic | 583 | Bug: Strange behavior with self-reference model | ### Description
Encountering some strange errors with self-referencing models.
In the first case, there seems to be a 50/50 chance for the `pytest` to work:
```python
from __future__ import annotations
from typing import Optional, Dict
import pydantic as pyd
from polyfactory.factories.pydantic_factory import ModelFactory
class Bar(pyd.BaseModel):
fields: Dict[str, Foo] = pyd.Field(default_factory=dict)
class Foo(pyd.BaseModel):
sometimes: Optional[Bar] = pyd.Field(default=None, exclude=True)
# never: Dict[str, Bar] = pyd.Field(default_factory=dict)
class FooFactory(ModelFactory[Foo]):
__model__ = Foo
def test_foo():
foo_instance = FooFactory.build()
assert isinstance(foo_instance, Foo)
```
In the second case, the `pytest` seems to never work:
```python
from __future__ import annotations
from typing import Optional, Dict
import pydantic as pyd
from polyfactory.factories.pydantic_factory import ModelFactory
class Bar(pyd.BaseModel):
fields: Dict[str, Foo] = pyd.Field(default_factory=dict)
class Foo(pyd.BaseModel):
# sometimes: Optional[Bar] = pyd.Field(default=None, exclude=True)
never: Dict[str, Bar] = pyd.Field(default_factory=dict)
class FooFactory(ModelFactory[Foo]):
__model__ = Foo
def test_foo():
foo_instance = FooFactory.build()
assert isinstance(foo_instance, Foo)
```
### Release Version
2.16.2
### Platform
- [X] Linux
- [ ] Mac
- [ ] Windows
- [ ] Other (Please specify in the description above) | closed | 2024-09-13T19:12:29Z | 2025-03-20T15:53:18Z | https://github.com/litestar-org/polyfactory/issues/583 | [
"bug"
] | FredrikBakken | 1 |
huggingface/datasets | tensorflow | 7,364 | API endpoints for gated dataset access requests | ### Feature request
I would like a programatic way of requesting access to gated datasets. The current solution to gain access forces me to visit a website and physically click an "agreement" button (as per the [documentation](https://huggingface.co/docs/hub/en/datasets-gated#access-gated-datasets-as-a-user)).
An ideal approach would be HF API download methods that negotiate access on my behalf based on information from my CLI login and/or token. I realise that may be naive given the various types of access semantics available to dataset authors (automatic versus manual approval, for example) and complexities it might add to existing methods, but something along those lines would be nice.
Perhaps using the `*_access_request` methods available to dataset authors can be a precedent; see [`reject_access_request`](https://huggingface.co/docs/huggingface_hub/main/en/package_reference/hf_api#huggingface_hub.HfApi.reject_access_request) for example.
### Motivation
When trying to download files from a gated dataset, I'm met with a `GatedRepoError` and instructed to visit the repository's website to gain access:
```
Cannot access gated repo for url https://huggingface.co/datasets/open-llm-leaderboard/meta-llama__Meta-Llama-3.1-70B-Instruct-details/resolve/main/meta-llama__Meta-Llama-3.1-70B-Instruct/samples_leaderboard_math_precalculus_hard_2024-07-19T18-47-29.522341.jsonl.
Access to dataset open-llm-leaderboard/meta-llama__Meta-Llama-3.1-70B-Instruct-details is restricted and you are not in the authorized list. Visit https://huggingface.co/datasets/open-llm-leaderboard/meta-llama__Meta-Llama-3.1-70B-Instruct-details to ask for access.
```
This makes task automation extremely difficult. For example, I'm interested in studying sample-level responses of models on the LLM leaderboard -- how they answered particular questions on a given evaluation framework. As I come across more and more participants that gate their data, it's becoming unwieldy to continue my work (there over 2,000 participants, so in the worst case that's the number of website visits I'd need to manually undertake).
One approach is use Selenium to react to the `GatedRepoError`, but that seems like overkill; and a potential violation HF terms of service (?).
As mentioned in the previous section, there seems to be an [API for gated dataset owners](https://huggingface.co/docs/hub/en/datasets-gated#via-the-api) to managed access requests, and thus some appetite for allowing automated management of gating. This feature request is to extend that to dataset users.
### Your contribution
Whether I can help depends on a few things; one being the complexity of the underlying gated access design. If this feature request is accepted I am open to being involved in discussions and testing, and even development under the right time-outcome tradeoff. | closed | 2025-01-09T06:21:20Z | 2025-01-09T11:17:40Z | https://github.com/huggingface/datasets/issues/7364 | [
"enhancement"
] | jerome-white | 3 |
ultralytics/ultralytics | deep-learning | 19,432 | Integrate Bifpn and Coordatt into yolov8 | i was trying to integrating both coordatt and bifpn into yolov8
when i modify yolov8 with bifpn alone it work perfect
and when i modify yolov8 with coordatt alone it work
when i try to combine them in single custom model
issue arise
```
KeyError Traceback (most recent call last)
[<ipython-input-30-32ffe6bf1a5e>](https://localhost:8080/#) in <cell line: 0>()
5
6 # Load the model with the custom architecture from your new YAML
----> 7 model = YOLO(model_path)
8
9 # Print the model architecture
4 frames
[/content/ultralytics/ultralytics/nn/tasks.py](https://localhost:8080/#) in parse_model(d, ch, verbose)
1016 C2fPSA,
1017 C2fCIB,
-> 1018 C2PSA,
1019 A2C2f,
1020
```
yaml of both
`# Ultralytics YOLO 🚀, AGPL-3.0 license
# YOLOv8 object detection model with P3-P5 outputs. For Usage examples see https://docs.ultralytics.com/tasks/detect
# Parameters
nc: 2 # number of classes
scales: # model compound scaling constants, i.e. 'model=yolov8n.yaml' will call yolov8.yaml with scale 'n'
# [depth, width, max_channels]
n: [0.33, 0.25, 1024] # YOLOv8n summary: 225 layers, 3157200 parameters, 3157184 gradients, 8.9 GFLOPs
s: [0.33, 0.50, 1024] # YOLOv8s summary: 225 layers, 11166560 parameters, 11166544 gradients, 28.8 GFLOPs
m: [0.67, 0.75, 768] # YOLOv8m summary: 295 layers, 25902640 parameters, 25902624 gradients, 79.3 GFLOPs
l: [1.00, 1.00, 512] # YOLOv8l summary: 365 layers, 43691520 parameters, 43691504 gradients, 165.7 GFLOPs
x: [1.00, 1.25, 512] # YOLOv8x summary: 365 layers, 68229648 parameters, 68229632 gradients, 258.5 GFLOPs
# YOLOv8.0n backbone
backbone:
# [from, repeats, module, args]
- [-1, 1, Conv, [64, 3, 2]] # 0-P1/2
- [-1, 1, Conv, [128, 3, 2]] # 1-P2/4
- [-1, 3, C2f, [128, True]]
- [-1, 1, Conv, [256, 3, 2]] # 3-P3/8
- [-1, 6, C2f, [256, True]]
- [-1, 1, Conv, [512, 3, 2]] # 5-P4/16
- [-1, 6, C2f, [512, True]]
- [-1, 1, Conv, [1024, 3, 2]] # 7-P5/32
- [-1, 3, C2f, [1024, True]]
- [-1, 1, SPPF, [1024, 5]] # 9
# YOLOv8.0n head
head:
- [-1, 1, nn.Upsample, [None, 2, 'nearest']]
- [[-1, 6], 1, BiFPN_Concat2, [1]] # cat backbone P4
- [-1, 3, C2f, [512]] # 12
- [-1, 1, nn.Upsample, [None, 2, 'nearest']]
- [[-1, 4], 1, BiFPN_Concat2, [1]] # cat backbone P3
- [-1, 3, C2f, [256]] # 15 (P3/8-small)
- [-1, 1, CoordAtt, [64, 64]] # 16 CA 추가
- [-1, 1, Conv, [256, 3, 2]] # 17
- [[-1, 12], 1, BiFPN_Concat3, [1]] # 18 cat head P4
- [-1, 3, C2f, [512]] # 19 (P4/16-medium)
- [-1, 1, CoordAtt, [128, 128]] # 20 CA 추가
- [-1, 1, Conv, [512, 3, 2]] # 21
- [[-1, 9], 1, BiFPN_Concat2, [1]] # 22 cat head P5
- [-1, 3, C2f, [1024]] # 23 (P5/32-large)
- [-1, 1, CoordAtt, [256, 256]] # 24 CA 추가
- [[16, 20, 24], 1, Detect, [nc]] # Detect(P3, P4, P5)`
i also added import in tasks.py
`
import torch
from ultralytics.nn.modules.bifpn import BiFPN_Concat2 ,BiFPN_Concat3
globals()['BiFPN_Concat3'] = BiFPN_Concat3
globals()['BiFPN_Concat2'] = BiFPN_Concat2
from ultralytics.nn.modules.coordatt import CoordAtt
globals()['CoordAtt'] = CoordAtt
`
i created file for bifpn and file for coordatt
class of coordatt
`
import torch
import torch.nn as nn
import torch.nn.functional as F
__all__ = ('CoordAtt')
class h_sigmoid(nn.Module):
def __init__(self, inplace=True):
super(h_sigmoid, self).__init__()
self.relu = nn.ReLU6(inplace=inplace)
def forward(self, x):
return self.relu(x + 3) / 6
class h_swish(nn.Module):
def __init__(self, inplace=True):
super(h_swish, self).__init__()
self.sigmoid = h_sigmoid(inplace=inplace)
def forward(self, x):
return x * self.sigmoid(x)
class CoordAtt(nn.Module):
def __init__(self, inp, oup, reduction=32): # inp: number of input channel, oup: number of output channel
super(CoordAtt, self).__init__()
self.pool_h = nn.AdaptiveAvgPool2d((None, 1))
self.pool_w = nn.AdaptiveAvgPool2d((1, None))
mip = max(8, inp // reduction)
self.conv1 = nn.Conv2d(inp, mip, kernel_size=1, stride=1, padding=0)
self.bn1 = nn.BatchNorm2d(mip)
self.act = h_swish()
self.conv_h = nn.Conv2d(mip, oup, kernel_size=1, stride=1, padding=0)
self.conv_w = nn.Conv2d(mip, oup, kernel_size=1, stride=1, padding=0)
def forward(self, x):
identity = x
_,_,h,w = x.size()
x_h = self.pool_h(x)
x_w = self.pool_w(x).permute(0, 1, 3, 2)
y = torch.cat([x_h, x_w], dim=2)
y = self.conv1(y)
y = self.bn1(y)
y = self.act(y)
x_h, x_w = torch.split(y, [h, w], dim=2)
x_w = x_w.permute(0, 1, 3, 2)
a_h = self.conv_h(x_h).sigmoid()
a_w = self.conv_w(x_w).sigmoid()
out = identity * a_w * a_h
return out`
and bifpn class
`import math
import numpy as np
import torch
import torch.nn as nn
# 结合BIFPM 设置的学习参数 学习不同分类的权重
# 两个分类Concat操作
class BiFPN_Concat2(nn.Module):
def __init__(self, dimension=1):
super(BiFPN_Concat2, self).__init__()
self.d = dimension
self.w = nn.Parameter(torch.ones(2, dtype=torch.float32), requires_grad=True)
self.epsilon = 0.0001
def forward(self, x):
w = self.w
weight = w / (torch.sum(w, dim=0) + self.epsilon) # 将权重进行归一化
# Fast normalized fusion
x = [weight[0] * x[0], weight[1] * x[1]]
return torch.cat(x, self.d)
# 三个分类Concat操作
class BiFPN_Concat3(nn.Module):
def __init__(self, dimension=1):
super(BiFPN_Concat3, self).__init__()
self.d = dimension
# 设置的学习参数 nn.Parameter的作用是:将一个不可调谐的类型Tensor转换成可以调谐的类型parameter
# 并且会向宿主模型注册该参数 成为唯一的一组Model_parameters()会包含这个parameter
# 从前任参数优化到目标调用后的一些优化
self.w = nn.Parameter(torch.ones(3, dtype=torch.float32), requires_grad=True)
self.epsilon = 0.0001
def forward(self, x):
w = self.w
weight = w / (torch.sum(w, dim=0) + self.epsilon) # 将权重进行归一化
# Fast normalized fusion
x = [weight[0] * x[0], weight[1] * x[1], weight[2] * x[2]]
return torch.cat(x, self.d)
`
i also imported them in init.py
i hope you can help me with the error | open | 2025-02-26T00:06:01Z | 2025-02-27T00:44:27Z | https://github.com/ultralytics/ultralytics/issues/19432 | [
"detect"
] | Youssef-Hassan-Git | 2 |
blb-ventures/strawberry-django-plus | graphql | 86 | How to work with mutations with foreignkeys | Hi Bellini,
I have some problems, as always, I'm sorry.
I don't think it should be an issue, but maybe you can show me in one sentence how to make things better/easier/properly?
I am trying set some fields + a ForeignKey in the Create Mutation.
```
mutation createMapService {
createMapService(input: {url: "https://bbbb.eu", mapSet: 5}) {
pk
}
}
```
As I see here: [https://strawberry.rocks/docs/types/object-types](url) I define the field: `map_set: "MapSet"` (Im using `""` because the MapSet type is defined in my code later).
This works in another scenario (without strawberry-django-plus), but with strawberry-django-plus the mutation fails and I can find this code in `strawberry_django_plus/mutations/resolvers.py`, line 276.
```
elif isinstance(field, models.ForeignKey) and isinstance(
value,
(ParsedObject, strawberry.ID),
):
```
This fails (`isinstance() arg 2 must be a type, a tuple of types, or a union`) which comes I believe from the comparision with `strawberry.id`. I can the code here make perfectly good for my goals - `type()` to prevent the error, `int` to allow plain id).
```
elif isinstance(field, models.ForeignKey) and isinstance(
value,
(ParsedObject, type(strawberry.ID), int),
):
```
However I am really week in typing in python, so I don't know if such change is acceptable. | closed | 2022-07-20T13:28:54Z | 2022-07-21T14:04:51Z | https://github.com/blb-ventures/strawberry-django-plus/issues/86 | [] | zvolsky | 3 |
mckinsey/vizro | pydantic | 424 | Check all links in docs and make them more accessible | Guidelines for links are as follows:
* We should comply with https://vizro.readthedocs.io/en/stable/pages/development/documentation-style-guide/#language
* Links should always work.
### Task (1) Fix all links
Before we have perfection, we need to run through all the docs pages and fix issues like this one https://github.com/mckinsey/vizro/pull/422#discussion_r1566440210 where they arise.
### Task (2) Set up style checking for future content
One way to enforce this ongoing would be to use Vale. I've asked how to do this:
https://github.com/errata-ai/vale/discussions/807
### Task (3) Add external link checking (if not already running) to CI.
We already have this for internal links in that we build with `--strict` but need to have something check links to Dash etc as Kedro does.
This ticket doesn't need technical writing skills nor does it need Vale knowledge. Good first issue for a new contributor! | closed | 2024-04-16T05:33:06Z | 2024-10-09T09:20:33Z | https://github.com/mckinsey/vizro/issues/424 | [
"Help Wanted :pray:",
"Docs :spiral_notepad:",
"Good first issue :baby_chick:",
"hacktoberfest"
] | stichbury | 17 |
robotframework/robotframework | automation | 4,482 | WHILE and FOR loop contents not shown in log if running them fails due to errors | When debugging #4480 I noticed that if WHILE loop condition is invalid, the loop is totally empty in the log file. The same was true also if the loop had syntax errors (e.g. missing END) and also FOR loops had same problem. We in general try to show all data in log even if it's not run (#3842) and have, for example, enhanced handling FOR loops that aren't run but are nevertheless valid so like that (#4184). Let's show loop contents also in these error situations. | closed | 2022-09-27T21:18:10Z | 2022-09-29T21:09:09Z | https://github.com/robotframework/robotframework/issues/4482 | [
"bug",
"priority: medium",
"rc 1"
] | pekkaklarck | 0 |
aio-libs/aiomysql | asyncio | 224 | "UPDATE" has no effect | await aiomysql.create_pool(maxsize=db['pool'], host=connection['host'], port=connection['port'],
user=connection['user'], password=connection['password'],
db=connection['database'], autocommit=True, loop=loop, charset='utf8')
But, Run "UPDATE" has no effect. | open | 2017-10-30T12:03:18Z | 2022-06-20T11:39:45Z | https://github.com/aio-libs/aiomysql/issues/224 | [
"docs"
] | darkforest42 | 9 |
Ehco1996/django-sspanel | django | 102 | 大佬 关于面板生成的ssr无法连接问题 | 大佬 前后端成功配置连接 后端使用webapi 面板显示后端在线 用户有剩余流量 但是生成的ssr连不上外网
打开软件之后甚至百度都打不开了... 我感觉是 加密方式 混淆 或者协议的问题 请问后端的在哪改呢? | closed | 2018-04-20T02:36:19Z | 2018-04-20T02:48:45Z | https://github.com/Ehco1996/django-sspanel/issues/102 | [] | wangchencom | 1 |
scikit-multilearn/scikit-multilearn | scikit-learn | 164 | 'MLTSVM' object has no attribute 'wk_norms' | When I try to predict test set i have this error:
'MLTSVM' object has no attribute 'wk_norms'
Here is my classifier trained params:
MLTSVM(c_k=0.5, lambda_param=1.0, max_iteration=500, sor_omega=1.0,
threshold=1e-06)
| open | 2019-04-15T06:36:12Z | 2019-05-21T11:45:33Z | https://github.com/scikit-multilearn/scikit-multilearn/issues/164 | [] | veseliy | 1 |
yt-dlp/yt-dlp | python | 11,732 | Vimeo/Patreon link returns 403 even with cookies fetched. | ### DO NOT REMOVE OR SKIP THE ISSUE TEMPLATE
- [X] I understand that I will be **blocked** if I *intentionally* remove or skip any mandatory\* field
### Checklist
- [X] I'm reporting that yt-dlp is broken on a **supported** site
- [X] I've verified that I have **updated yt-dlp to nightly or master** ([update instructions](https://github.com/yt-dlp/yt-dlp#update-channels))
- [X] I've checked that all provided URLs are playable in a browser with the same IP and same login details
- [X] I've checked that all URLs and arguments with special characters are [properly quoted or escaped](https://github.com/yt-dlp/yt-dlp/wiki/FAQ#video-url-contains-an-ampersand--and-im-getting-some-strange-output-1-2839-or-v-is-not-recognized-as-an-internal-or-external-command)
- [X] I've searched [known issues](https://github.com/yt-dlp/yt-dlp/issues/3766) and the [bugtracker](https://github.com/yt-dlp/yt-dlp/issues?q=) for similar issues **including closed ones**. DO NOT post duplicates
- [X] I've read the [guidelines for opening an issue](https://github.com/yt-dlp/yt-dlp/blob/master/CONTRIBUTING.md#opening-an-issue)
- [X] I've read about [sharing account credentials](https://github.com/yt-dlp/yt-dlp/blob/master/CONTRIBUTING.md#are-you-willing-to-share-account-details-if-needed) and I'm willing to share it if required
### Region
Serbia
### Provide a description that is worded well enough to be understood
I'm trying to download the vimeo video from Patreon and nothing that I tried of worked. Inevitable I'm getting a 403 access denied error. --cookies-from-browser chrome won't work due to "Could not copy Chrome cookie database. See https://github.com/yt-dlp/yt-dlp/issues/7271", tried every possible solution. In the end I managed to extract cookies to a .txt file and put a path to that file as advised here: https://github.com/yt-dlp/yt-dlp/issues/10927. Still no luck. Maybe I'm doing something wrong, I dunno.
403 error is being returned both when browser is closed / when it's open.
First time raising an issue here, I think I went through the checklist and didn't break any rules. Sorry if I did something wrong.
### Provide verbose output that clearly demonstrates the problem
- [X] Run **your** yt-dlp command with **-vU** flag added (`yt-dlp -vU <your command line>`)
- [ ] If using API, add `'verbose': True` to `YoutubeDL` params instead
- [X] Copy the WHOLE output (starting with `[debug] Command-line config`) and insert it below
### Complete Verbose Output
```shell
[debug] Command-line config: ['-vU', '--cookies', 'D:\\ffmpeg\\bin\\cookies.txt', '--referer', 'https://www.patreon.com/posts/wintersun-sons-110460144', 'https://vod-adaptive-ak.vimeocdn.com/exp=1733335309~acl=%2F17615183-bd54-4774-ae2f-5e2f01278e32%2F%2A~hmac=4f594011d2648f87d31d9548cc106303bf4ade3009747170124534498c36ca81/17615183-bd54-4774-ae2f-5e2f01278e32/v2/playlist/av/primary/prot/cXNyPTE/playlist.m3u8']
[debug] Encodings: locale cp1251, fs utf-8, pref cp1251, out utf-8, error utf-8, screen utf-8
[debug] yt-dlp version master@2024.12.03.202240 from yt-dlp/yt-dlp-master-builds [2b67ac300] (win_exe)
[debug] Python 3.10.11 (CPython AMD64 64bit) - Windows-10-10.0.19045-SP0 (OpenSSL 1.1.1t 7 Feb 2023)
[debug] exe versions: ffmpeg 2023-12-11-git-1439784ff0-full_build-www.gyan.dev (setts), ffprobe 2023-12-11-git-1439784ff0-full_build-www.gyan.dev, phantomjs 2.1.1
[debug] Optional libraries: Cryptodome-3.21.0, brotli-1.1.0, certifi-2024.08.30, curl_cffi-0.5.10, mutagen-1.47.0, requests-2.32.3, sqlite3-3.40.1, urllib3-2.2.3, websockets-14.1
[debug] Proxy map: {}
[debug] Request Handlers: urllib, requests, websockets, curl_cffi
[debug] Loaded 1837 extractors
[debug] Fetching release info: https://api.github.com/repos/yt-dlp/yt-dlp-master-builds/releases/latest
Latest version: master@2024.12.03.202240 from yt-dlp/yt-dlp-master-builds
yt-dlp is up to date (master@2024.12.03.202240 from yt-dlp/yt-dlp-master-builds)
[generic] Extracting URL: https://vod-adaptive-ak.vimeocdn.com/exp=1733335309~acl=%2F17615183-bd54-4774-ae2f-5e2f01278e32%2F%2A~hmac=4f594011d2648f87d31d9548cc106303bf4ade3009747170124534498c36ca81/17615183-bd54-4774-ae2f-5e2f01278e32/v2/playlist/av/primary/prot/cXNyPTE/playlist.m3u8
[generic] playlist: Downloading webpage
ERROR: [generic] Unable to download webpage: HTTP Error 403: Forbidden (caused by <HTTPError 403: Forbidden>)
File "yt_dlp\extractor\common.py", line 742, in extract
File "yt_dlp\extractor\generic.py", line 2393, in _real_extract
File "yt_dlp\extractor\common.py", line 911, in _request_webpage
File "yt_dlp\extractor\common.py", line 898, in _request_webpage
File "yt_dlp\YoutubeDL.py", line 4162, in urlopen
File "yt_dlp\networking\common.py", line 117, in send
File "yt_dlp\networking\_helper.py", line 208, in wrapper
File "yt_dlp\networking\common.py", line 340, in send
File "yt_dlp\networking\_requests.py", line 365, in _send
yt_dlp.networking.exceptions.HTTPError: HTTP Error 403: Forbidden
```
| closed | 2024-12-04T17:16:26Z | 2024-12-04T17:23:36Z | https://github.com/yt-dlp/yt-dlp/issues/11732 | [
"question"
] | TrulyStucker | 2 |
mars-project/mars | scikit-learn | 3,220 | [BUG] test_numexpr_execution.py::test_unary_execution may raises NotImplementedError: couldn't find matching opcode for 'invert_dd' | <!--
Thank you for your contribution!
Please review https://github.com/mars-project/mars/blob/master/CONTRIBUTING.rst before opening an issue.
-->
**Describe the bug**
A clear and concise description of what the bug is.
```python
Traceback (most recent call last):
File "/home/vsts/miniconda/envs/test/lib/python3.9/site-packages/numexpr/necompiler.py", line 820, in evaluate
compiled_ex = _numexpr_cache[numexpr_key]
KeyError: ('(~((abs((sin((cosh(V_0))))))))', (('optimization', 'aggressive'), ('truediv', False)), (('V_0', <class 'numpy.float64'>),))
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "/home/vsts/work/1/s/mars/tensor/fuse/numexpr.py", line 49, in execute
res = ne.evaluate(expr, local_dict=local_dict, global_dict={})
File "/home/vsts/miniconda/envs/test/lib/python3.9/site-packages/numexpr/necompiler.py", line 822, in evaluate
compiled_ex = _numexpr_cache[numexpr_key] = NumExpr(ex, signature, **context)
File "/home/vsts/miniconda/envs/test/lib/python3.9/site-packages/numexpr/necompiler.py", line 621, in NumExpr
threeAddrProgram, inputsig, tempsig, constants, input_names = precompile(ex, signature, context)
File "/home/vsts/miniconda/envs/test/lib/python3.9/site-packages/numexpr/necompiler.py", line 566, in precompile
ast = typeCompileAst(ast)
File "/home/vsts/miniconda/envs/test/lib/python3.9/site-packages/numexpr/necompiler.py", line 202, in typeCompileAst
raise NotImplementedError(
NotImplementedError: couldn't find matching opcode for 'invert_dd'
The above exception was the direct cause of the following exception:
Traceback (most recent call last):
File "/home/vsts/work/1/s/mars/services/subtask/worker/processor.py", line 189, in _execute_operand
return execute(ctx, op)
File "/home/vsts/work/1/s/mars/core/operand/core.py", line 491, in execute
result = executor(results, op)
File "/home/vsts/work/1/s/mars/tensor/fuse/numexpr.py", line 51, in execute
raise RuntimeError(
RuntimeError: Failed to evaluate numexpr '(~((abs((sin((cosh(V_0))))))))' on local dict {'V_0': array([[[0.21645726, 0.16604782, 0.92275661, 0.29407666],
```
The problem is occurred in CI test, the ~ can not apply to a float array. Similar issue: https://github.com/pandas-dev/pandas/blob/main/pandas/tests/computation/test_eval.py#L368
**To Reproduce**
To help us reproducing this bug, please provide information below:
1. Your Python version
2. The version of Mars you use
3. Versions of crucial packages, such as numpy, scipy and pandas
4. Full stack of the error.
5. Minimized code to reproduce the error.
**Expected behavior**
A clear and concise description of what you expected to happen.
**Additional context**
Add any other context about the problem here.
| closed | 2022-08-11T10:01:18Z | 2022-08-19T03:30:10Z | https://github.com/mars-project/mars/issues/3220 | [
"type: bug"
] | fyrestone | 0 |
StackStorm/st2 | automation | 5,352 | please support redis + tls (rediss) | ## SUMMARY
Redis with TLS not supported (rediss)
### STACKSTORM VERSION
Paste the output of ``st2 --version``:
```
$ st2 --version
st2 3.4.1, on Python 3.6.14
```
##### OS, environment, install method
Post what OS you are running this on, along with any other relevant information/
- e.g. Docker, Vagrant, Kubernetes, etc. Describe how you installed ST2
- e.g. one-line install, custom install, etc -->
Custom/Ansible
```
uname -a
Linux xxx 5.8.0-1042-aws #44~20.04.1-Ubuntu SMP Mon Aug 2 11:25:34 UTC 2021 x86_64 x86_64 x86_64 GNU/Linux
```
## Steps to reproduce the problem
add the following to `st2.conf`:
```
[coordination]
#local redis without tls enabled works fine...
#url = redis://127.0.0.1
url = rediss://:someredispassword@myredishost:6379
```
## Expected Results
What did you expect to happen when running the steps above?
`rediss` should work (ie TLS)
## Actual Results
What happened? What output did you get?
```
2021-09-05 17:38:46,483 140159477849888 WARNING named [-] Could not load rediss
2021-09-05 17:38:46,484 140159477849888 ERROR scheduler [-] (PID=1236037) Scheduler quit due to exception.
Traceback (most recent call last):
File "/opt/stackstorm/st2/lib/python3.6/site-packages/st2actions/cmd/scheduler.py", line 126, in main
return _run_scheduler()
File "/opt/stackstorm/st2/lib/python3.6/site-packages/st2actions/cmd/scheduler.py", line 68, in _run_scheduler
handler = scheduler_handler.get_handler()
File "/opt/stackstorm/st2/lib/python3.6/site-packages/st2actions/scheduler/handler.py", line 469, in get_handler
return ActionExecutionSchedulingQueueHandler()
File "/opt/stackstorm/st2/lib/python3.6/site-packages/st2actions/scheduler/handler.py", line 66, in __init__
self._coordinator = coordination_service.get_coordinator(start_heart=True)
File "/opt/stackstorm/st2/lib/python3.6/site-packages/st2common/services/coordination.py", line 227, in get_coordinator
COORDINATOR = coordinator_setup(start_heart=start_heart)
File "/opt/stackstorm/st2/lib/python3.6/site-packages/st2common/services/coordination.py", line 192, in coordinator_setup
coordinator = coordination.get_coordinator(url, member_id, lock_timeout=lock_timeout)
File "/opt/stackstorm/st2/lib/python3.6/site-packages/tooz/coordination.py", line 802, in get_coordinator
invoke_args=(member_id, parsed_url, options)).driver
File "/opt/stackstorm/st2/lib/python3.6/site-packages/stevedore/driver.py", line 61, in __init__
warn_on_missing_entrypoint=warn_on_missing_entrypoint
File "/opt/stackstorm/st2/lib/python3.6/site-packages/stevedore/named.py", line 89, in __init__
self._init_plugins(extensions)
File "/opt/stackstorm/st2/lib/python3.6/site-packages/stevedore/driver.py", line 113, in _init_plugins
(self.namespace, name))
stevedore.exception.NoMatches: No 'tooz.backends' driver found, looking for 'rediss'
```
Making sure to follow these steps will guarantee the quickest resolution possible.
Thanks!
| closed | 2021-09-05T17:42:57Z | 2021-09-24T13:37:26Z | https://github.com/StackStorm/st2/issues/5352 | [
"documentation"
] | bkk-bcd | 2 |
sktime/pytorch-forecasting | pandas | 831 | know and unknow values | - PyTorch-Forecasting version: 0.9.2
- PyTorch version: '1.10.1+cu113'
- Python version: 3.7
- Operating System: ubuntu
### Expected behavior
I want to use the TimeSeriesDataSet function, the description says that the UNKNOW values represent the past and the KNOW values represent what we know in the past and the future.
### Actual behavior
I did some values in a random test, and I pass feature '1' and '2' as KNOW values, even '3' as UNKNOW. But I can't interpret the result shown in the output. Both "encoder_cont" and "decoder_cont" see all values of all 3 features, although my idea is that the decoder should have one less parameter.
### Code to reproduce the problem
```
data = np.array([[1,2,3,4,5,6,7,8,9,1,100,0],
[11,12,13,14,15,16,17,18,19,2,101,0],
[21,22,23,24,25,26,27,28,29,3,102,0],
[31,32,33,34,35,36,37,38,39,4,103,0],
[41,42,43,44,45,46,47,48,49,5,104,0],
[51,52,53,54,55,56,57,58,59,6,105,0],
[61,62,63,64,65,66,67,68,69,7,106,0],
[71,72,73,74,75,76,77,78,79,8,107,0],
[81,82,83,84,85,86,87,88,89,9,108,0]
])
data = pd.DataFrame(data)
data.columns = ['1','2','3','4','5','6','7','8','9','time_idx','target','NODE_ID']
data['NODE_ID'] = data['NODE_ID'].astype(str)
max_encoder_length = 4
max_prediction_length = 4
training = TimeSeriesDataSet(
data,
time_idx='time_idx',
target="target",
group_ids=['NODE_ID'],
min_encoder_length=max_encoder_length,
max_encoder_length=max_encoder_length,
min_prediction_length=max_prediction_length,
max_prediction_length=max_prediction_length,
time_varying_known_reals=['1','2'],
time_varying_unknown_reals=['3'],
scalers= {'1': None,'2': None, '3': None}
)
train_dataloader = training.to_dataloader(train=True, batch_size=2, num_workers=0)
training.get_parameters()
{'time_idx': 'time_idx',
'target': 'target',
'group_ids': ['NODE_ID'],
'weight': None,
'max_encoder_length': 4,
'min_encoder_length': 4,
'min_prediction_idx': 1,
'min_prediction_length': 4,
'max_prediction_length': 4,
'static_categoricals': [],
'static_reals': [],
'time_varying_known_categoricals': [],
'time_varying_known_reals': ['1', '2'],
'time_varying_unknown_categoricals': [],
'time_varying_unknown_reals': ['3'],
'variable_groups': {},
'constant_fill_strategy': {},
'allow_missing_timesteps': False,
'lags': {},
'add_relative_time_idx': False,
'add_target_scales': False,
'add_encoder_length': False,
'target_normalizer': NaNLabelEncoder(),
'categorical_encoders': {'__group_id__NODE_ID': NaNLabelEncoder()},
'scalers': {'1': None, '2': None, '3': None},
'randomize_length': None,
'predict_mode': False}
x, y = next(iter(train_dataloader))
x
{'encoder_cat': tensor([], size=(2, 4, 0), dtype=torch.int64),
'encoder_cont': tensor([[[11., 12., 13.],
[21., 22., 23.],
[31., 32., 33.],
[41., 42., 43.]],
[[ 1., 2., 3.],
[11., 12., 13.],
[21., 22., 23.],
[31., 32., 33.]]]),
'encoder_target': tensor([[1, 2, 3, 4],
[0, 1, 2, 3]]),
'encoder_lengths': tensor([4, 4]),
'decoder_cat': tensor([], size=(2, 4, 0), dtype=torch.int64),
'decoder_cont': tensor([[[51., 52., 53.],
[61., 62., 63.],
[71., 72., 73.],
[81., 82., 83.]],
[[41., 42., 43.],
[51., 52., 53.],
[61., 62., 63.],
[71., 72., 73.]]]),
'decoder_target': tensor([[5, 6, 7, 8],
[4, 5, 6, 7]]),
'decoder_lengths': tensor([4, 4]),
'decoder_time_idx': tensor([[6, 7, 8, 9],
[5, 6, 7, 8]]),
'groups': tensor([[0],
[0]]),
'target_scale': tensor([[0., 0.],
[0., 0.]])}
```
| open | 2022-01-17T11:29:02Z | 2022-05-12T03:04:53Z | https://github.com/sktime/pytorch-forecasting/issues/831 | [] | korosig | 5 |
microsoft/nni | pytorch | 5,126 | requests.exceptions.ConnectionError: HTTPConnectionPool(host='localhost', port=8140): Max retries exceeded with url: /api/v1/nni/check-status (Caused by NewConnectionError('<urllib3.connection.HTTPConnection object at 0x7ffa090c0d00>: Failed to establish a new connection: [Errno 111] Connection refused')) | when i run ''nnictl create --config config.yml -p 8140", i get the error:
```
requests.exceptions.ConnectionError: HTTPConnectionPool(host='localhost', port=8140): Max retries exceeded with url: /api/v1/nni/check-status (Caused by NewConnectionError('<urllib3.connection.HTTPConnection object at 0x7ffa090c0d00>: Failed to establish a new connection: [Errno 111] Connection refused'))
```
**Environment**:
- NNI version:
- v2.9
- Training service (local|remote|pai|aml|etc):
- remote
- Client OS:
- Server OS (for remote mode only):
- Python version:
- 3.8
- PyTorch/TensorFlow version:
- PyTorch 1.7.0
- Is conda/virtualenv/venv used?:
- conda
- Is running in Docker?:
no
**Configuration**:
- Experiment config (remember to remove secrets!):
```
trialConcurrency: 2 #trail的并发数,根据GPU数量设置,此值为几就有几个train在同时跑
trainingService:
platform: local
gpuIndices: [6,7] # 使用哪几个GPU
# gpuIndices: [0] # 使用哪几个GPU
useActiveGpu: True # 默认值false。是否使用已经被其他进程使用的gpu,包括graphical desktop占用的。
maxTrialNumberPerGpu: 1 #指定1个GPU上最大并发trail的数量,在确保显存达到足以容下任何两个trail时,再设置为2。
trialGpuNumber: 1 # 每个trail所需要的gpu
```
- Search space:
```
{
"epochs":{"_type":"choice","_value":[400,500]},
"lr":{"_type":"quniform","_value":[0.0001,0.0025,0.0005]},
}
```
**Log message**:
- nnimanager.log:
```
[2022-09-13 20:23:23] INFO (main) Start NNI manager
```
- dispatcher.log:
none
- nnictl stdout and stderr:
none
**How to reproduce it?**: | closed | 2022-09-13T12:33:13Z | 2022-09-18T13:31:15Z | https://github.com/microsoft/nni/issues/5126 | [
"waiting user confirm",
"support",
"NNI SDK",
"need more info"
] | xiangtaowong | 15 |
onnx/onnx | deep-learning | 6,302 | a) Feature Request: Function sample_dirichlet, b) State of probabilistic model support? | I am very interested in converting DeepLearning models, that contain the PowerSpherical function (https://github.com/andife/power_spherical) to ONNX.
Currently this fails because of the Dirichlet function (https://github.com/pytorch/pytorch/issues/116336).
After my research, I came across https://github.com/onnx/onnxmltools/issues/549, among others
and wondered whether it would be useful to have gamma, dirichlet, beta available in general?
For this reason, the question arises as to what the current state of probabilistic model support looks like?
Dirichlet is available in
pytorch: https://pytorch.org/docs/stable/distributions.html#dirichlet
tensorflow: https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/ops/distributions/dirichlet_multinomial.py
Would it be a suitable direction, e.g. to create a sample Dirichlet method as an Onnx function based on RandomUniform (https://onnx.ai/onnx/operators/onnx__RandomUniform.html#l-onnx-doc-randomuniform)?
| open | 2024-08-17T04:38:43Z | 2024-09-30T21:38:34Z | https://github.com/onnx/onnx/issues/6302 | [] | andife | 6 |
allure-framework/allure-python | pytest | 39 | Parametrized session-scoped fixtures: KeyError in logger | Reproducing:
```python
import pytest
@pytest.fixture(scope='session', params=['param a'])
def param(request):
return request.param
def test_smth(param):
with pytest.allure.step('Test {}'.format(param)):
assert param == 'param a'
```
Trace:
```
$ py.test test.py --alluredir allure-results/
============================================================================================================ test session starts =============================================================================================================
platform linux -- Python 3.4.5, pytest-3.0.6, py-1.4.32, pluggy-0.4.0
rootdir: /home/igorock/Work/backend-tests, inifile:
plugins: timeout-1.2.0, allure-adaptor-2.0.1
collected 1 items
test.py E
=================================================================================================================== ERRORS ===================================================================================================================
____________________________________________________________________________________________________ ERROR at setup of test_smth[param b] ____________________________________________________________________________________________________
self = <allure.listener.AllureListener object at 0x7fee15ef7c50>, fixturedef = <FixtureDef name='param' scope='session' baseid='test.py' >, request = <SubRequest 'param' for <Function 'test_smth[param b]'>>
@pytest.hookimpl(hookwrapper=True)
def pytest_fixture_setup(self, fixturedef, request):
uuid = uuid4()
node_id = request.node.nodeid
parent_uuid = self._cache.get(node_id) if fixturedef.scope == 'function' else self._cache.get(fixturedef)
parameters = allure_parameters(fixturedef, request)
# ToDo autouse fixtures
if fixturedef.baseid and parent_uuid:
fixture = ExecutableItem(start=now(), name=fixturedef.argname)
self.allure_logger.start_before_fixture(parent_uuid, uuid, fixture)
if parameters and parent_uuid:
parameters = Parameter(**parameters) if parameters else []
> self.allure_logger.update_test(self._cache.get(node_id), parameters=parameters)
/usr/lib/python3.4/site-packages/allure/listener.py:95:
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
/usr/lib/python3.4/site-packages/allure/logger.py:57: in update_test
self._update_item(uuid, **kwargs)
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
self = <allure.logger.AllureLogger object at 0x7fee15ef7c88>, uuid = None, kwargs = {'parameters': Parameter(name='param', value='param b')}
def _update_item(self, uuid, **kwargs):
> item = self._items[uuid]
E KeyError: None
/usr/lib/python3.4/site-packages/allure/logger.py:18: KeyError
========================================================================================================== 1 error in 0.05 seconds ===========================================================================================================
```
Environment:
```
$ sudo pip freeze | grep pytest
You are using pip version 8.1.2, however version 9.0.1 is available.
You should consider upgrading via the 'pip install --upgrade pip' command.
pytest==3.0.6
pytest-allure-adaptor==2.0.1
pytest-timeout==1.2.0
```
P.S. allure-python2 was updated 2017.02.10 at 16:24 MSK :) | closed | 2017-02-10T13:24:34Z | 2017-02-12T14:47:23Z | https://github.com/allure-framework/allure-python/issues/39 | [] | i-feofilaktov | 0 |
tqdm/tqdm | jupyter | 1,476 | Error when sleep() in trange with rich | >>> from time import sleep
>>> from tqdm.rich import trange
>>> for i in trange(10):
... sleep(1)
...
/usr/lib/python3.11/site-packages/tqdm/rich.py:145: TqdmExperimentalWarning: rich is experimental/alpha
return tqdm_rich(range(*args), **kwargs)
0% ━━━━━━ 0/10 [ 0:00… < -:--:… , ? ]
it/s Bad system call | open | 2023-06-03T23:19:50Z | 2023-06-03T23:19:50Z | https://github.com/tqdm/tqdm/issues/1476 | [] | o-murphy | 0 |
serengil/deepface | machine-learning | 770 | Value error when trying to analyze emotion | ValueError Traceback (most recent call last)
Cell In[16], line 1
----> 1 objs = DeepFace.analyze(img_path = "test_photo.jpg",
2 actions = ['age', 'gender', 'race', 'emotion']
3 )
4 display_image_with_matplotlib(image)
File ~\anaconda3\envs\tf-gpu\lib\site-packages\deepface\DeepFace.py:336, in analyze(img_path, actions, enforce_detection, detector_backend, align, silent)
333 img_gray = cv2.resize(img_gray, (48, 48))
334 img_gray = np.expand_dims(img_gray, axis=0)
--> 336 emotion_predictions = models["emotion"].predict(img_gray, verbose=0)[0, :]
338 sum_of_predictions = emotion_predictions.sum()
340 obj["emotion"] = {}
File ~\anaconda3\envs\tf-gpu\lib\site-packages\tensorflow\python\keras\engine\training.py:1749, in Model.predict(self, x, batch_size, verbose, steps, callbacks, max_queue_size, workers, use_multiprocessing)
1747 for step in data_handler.steps():
1748 callbacks.on_predict_batch_begin(step)
-> 1749 tmp_batch_outputs = self.predict_function(iterator)
1750 if data_handler.should_sync:
1751 context.async_wait()
File ~\anaconda3\envs\tf-gpu\lib\site-packages\tensorflow\python\eager\def_function.py:885, in Function.__call__(self, *args, **kwds)
882 compiler = "xla" if self._jit_compile else "nonXla"
884 with OptionalXlaContext(self._jit_compile):
--> 885 result = self._call(*args, **kwds)
887 new_tracing_count = self.experimental_get_tracing_count()
888 without_tracing = (tracing_count == new_tracing_count)
File ~\anaconda3\envs\tf-gpu\lib\site-packages\tensorflow\python\eager\def_function.py:924, in Function._call(self, *args, **kwds)
921 self._lock.release()
922 # In this case we have not created variables on the first call. So we can
923 # run the first trace but we should fail if variables are created.
--> 924 results = self._stateful_fn(*args, **kwds)
925 if self._created_variables and not ALLOW_DYNAMIC_VARIABLE_CREATION:
926 raise ValueError("Creating variables on a non-first call to a function"
927 " decorated with tf.function.")
File ~\anaconda3\envs\tf-gpu\lib\site-packages\tensorflow\python\eager\function.py:3038, in Function.__call__(self, *args, **kwargs)
3035 """Calls a graph function specialized to the inputs."""
3036 with self._lock:
3037 (graph_function,
-> 3038 filtered_flat_args) = self._maybe_define_function(args, kwargs)
3039 return graph_function._call_flat(
3040 filtered_flat_args, captured_inputs=graph_function.captured_inputs)
File ~\anaconda3\envs\tf-gpu\lib\site-packages\tensorflow\python\eager\function.py:3459, in Function._maybe_define_function(self, args, kwargs)
3449 with ag_ctx.ControlStatusCtx(
3450 status=ag_status, options=self._autograph_options):
3451
(...)
3454 # and 2. there's no provided input signature
3455 # and 3. there's been a cache miss for this calling context
3456 if (self._experimental_relax_shapes and
3457 self.input_signature is None and
3458 call_context_key in self._function_cache.missed):
-> 3459 return self._define_function_with_shape_relaxation(
3460 args, kwargs, flat_args, filtered_flat_args, cache_key_context)
3462 self._function_cache.missed.add(call_context_key)
3463 graph_function = self._create_graph_function(args, kwargs)
File ~\anaconda3\envs\tf-gpu\lib\site-packages\tensorflow\python\eager\function.py:3381, in Function._define_function_with_shape_relaxation(self, args, kwargs, flat_args, filtered_flat_args, cache_key_context)
3374 (relaxed_arg_specs, relaxed_kwarg_specs) = nest.pack_sequence_as(
3375 (args, kwargs), relaxed_arg_specs, expand_composites=False)
3376 (args, kwargs) = nest.pack_sequence_as(
3377 (relaxed_arg_specs, relaxed_kwarg_specs),
3378 flat_args,
3379 expand_composites=True)
-> 3381 graph_function = self._create_graph_function(
3382 args, kwargs, override_flat_arg_shapes=relaxed_arg_shapes)
3383 self._function_cache.arg_relaxed[rank_only_cache_key] = graph_function
3385 return (graph_function, [
3386 t for t in nest.flatten((args, kwargs), expand_composites=True)
3387 if isinstance(t, (ops.Tensor,
3388 resource_variable_ops.BaseResourceVariable))
3389 ])
File ~\anaconda3\envs\tf-gpu\lib\site-packages\tensorflow\python\eager\function.py:3298, in Function._create_graph_function(self, args, kwargs, override_flat_arg_shapes)
3293 missing_arg_names = [
3294 "%s_%d" % (arg, i) for i, arg in enumerate(missing_arg_names)
3295 ]
3296 arg_names = base_arg_names + missing_arg_names
3297 graph_function = ConcreteFunction(
-> 3298 func_graph_module.func_graph_from_py_func(
3299 self._name,
3300 self._python_function,
3301 args,
3302 kwargs,
3303 self.input_signature,
3304 autograph=self._autograph,
3305 autograph_options=self._autograph_options,
3306 arg_names=arg_names,
3307 override_flat_arg_shapes=override_flat_arg_shapes,
3308 capture_by_value=self._capture_by_value),
3309 self._function_attributes,
3310 function_spec=self.function_spec,
3311 # Tell the ConcreteFunction to clean up its graph once it goes out of
3312 # scope. This is not the default behavior since it gets used in some
3313 # places (like Keras) where the FuncGraph lives longer than the
3314 # ConcreteFunction.
3315 shared_func_graph=False)
3316 return graph_function
File ~\anaconda3\envs\tf-gpu\lib\site-packages\tensorflow\python\framework\func_graph.py:1007, in func_graph_from_py_func(name, python_func, args, kwargs, signature, func_graph, autograph, autograph_options, add_control_dependencies, arg_names, op_return_value, collections, capture_by_value, override_flat_arg_shapes, acd_record_initial_resource_uses)
1004 else:
1005 _, original_func = tf_decorator.unwrap(python_func)
-> 1007 func_outputs = python_func(*func_args, **func_kwargs)
1009 # invariant: `func_outputs` contains only Tensors, CompositeTensors,
1010 # TensorArrays and `None`s.
1011 func_outputs = nest.map_structure(convert, func_outputs,
1012 expand_composites=True)
File ~\anaconda3\envs\tf-gpu\lib\site-packages\tensorflow\python\eager\def_function.py:668, in Function._defun_with_scope.<locals>.wrapped_fn(*args, **kwds)
664 with default_graph._variable_creator_scope(scope, priority=50): # pylint: disable=protected-access
665 # __wrapped__ allows AutoGraph to swap in a converted function. We give
666 # the function a weak reference to itself to avoid a reference cycle.
667 with OptionalXlaContext(compile_with_xla):
--> 668 out = weak_wrapped_fn().__wrapped__(*args, **kwds)
669 return out
File ~\anaconda3\envs\tf-gpu\lib\site-packages\tensorflow\python\framework\func_graph.py:994, in func_graph_from_py_func.<locals>.wrapper(*args, **kwargs)
992 except Exception as e: # pylint:disable=broad-except
993 if hasattr(e, "ag_error_metadata"):
--> 994 raise e.ag_error_metadata.to_exception(e)
995 else:
996 raise
ValueError: Input 0 of layer sequential is incompatible with the layer: : expected min_ndim=4, found ndim=3. Full shape received: (None, 48, 48) | closed | 2023-06-04T20:20:52Z | 2023-06-04T20:23:40Z | https://github.com/serengil/deepface/issues/770 | [
"question"
] | ionut-girla | 1 |
netbox-community/netbox | django | 18,927 | Shortcut to make Primary when adding new MAC | ### NetBox version
v4.2.4
### Feature type
Change to existing functionality
### Proposed functionality
When you click the plus button to add a new MAC address to an interface, there should be a checkbox for "make this the primary MAC for the interface" on this page.
This would be similar to the checkbox for selecting the Primary IP when adding a new IP Address.

### Use case
Currently, this requires too many clicks. You have to first create the MAC, then edit the interface and assign the Primary MAC. It was easier before the 4.2 update.
### Database changes
_No response_
### External dependencies
_No response_ | closed | 2025-03-17T17:51:49Z | 2025-03-18T12:47:55Z | https://github.com/netbox-community/netbox/issues/18927 | [
"status: duplicate",
"type: feature"
] | llamafilm | 2 |
hpcaitech/ColossalAI | deep-learning | 5,948 | [FEATURE]: Request updates for pretraining roberta | ### Describe the feature
I encountered issues while trying to run the RoBERTa pretraining using the provided ColossalAI repository code. The code has not been maintained and is currently not functional.


| open | 2024-07-29T09:24:02Z | 2024-07-29T09:24:02Z | https://github.com/hpcaitech/ColossalAI/issues/5948 | [
"enhancement"
] | jiahuanluo | 0 |
python-restx/flask-restx | flask | 549 | SwaggerUIBundle is not defined | I am using `flask-restx==1.1.0`
My Python is `3.8.10`
Sometime I am seeing this issue in my swagger dashboard
`GET https://{host}/api/swaggerui/swagger-ui-standalone-preset.js net::ERR_ABORTED 404 (NOT FOUND)
{host}/:71 GET https://{host}/api/swaggerui/swagger-ui-bundle.js net::ERR_ABORTED 404 (NOT FOUND)
{host}/:7 GET https://{host}/api/swaggerui/swagger-ui.css net::ERR_ABORTED 404 (NOT FOUND)
(index):75 Uncaught ReferenceError: SwaggerUIBundle is not defined
at window.onload ((index):75:40)`
And my dashboard is not getting load because of this issue
Can someone please help me with this ? I am not able to find much about this issue in internet already | open | 2023-06-29T10:29:34Z | 2023-07-07T03:26:03Z | https://github.com/python-restx/flask-restx/issues/549 | [
"bug"
] | viveksahu56722 | 5 |
viewflow/viewflow | django | 361 | OrderBy for JSONField support | Seems there is no automatic way to disable sortable_by in the Django admin on the field level. So probably, its simple to recearch a way to sort by virtual column | open | 2022-12-21T07:09:58Z | 2022-12-21T07:09:58Z | https://github.com/viewflow/viewflow/issues/361 | [
"request/enhancement",
"dev/flow"
] | kmmbvnr | 0 |
kizniche/Mycodo | automation | 1,027 | anyleaf __init__.py has no reference to EcSensor | ### Mycodo > mycodo > inputs > anyleaf_ec.py tries to import EcSensor from anyleaf. A closed examination of anyleaf indicates that there is no EcSensor section.
### Versions:
- Mycodo Version: [8.11.0]
- Raspberry Pi Version: [4]
- Raspbian OS Version: [Raspberry OS, latest and updated]
### Reproducibility
**Mycodo > mycodo > inputs > anyleaf_ec.py**
line 73 tries to import EcSensor from anyleaf, which is not referenced in the __init__ file.
58 class InputModule(AbstractInput):
59 """A sensor support class that monitors AnyLeaf sensor conductivity (EC)"""
60
61 def __init__(self, input_dev, testing=False):
62 super(InputModule, self).__init__(input_dev, testing=testing, name=__name__)
63
64 self.sensor = None
65 self.constant_k = None
66
67 if not testing:
68 self.setup_custom_options(
69 INPUT_INFORMATION['custom_options'], input_dev)
70 self.initialize_input()
71
72 def initialize_input(self):
73 from anyleaf import EcSensor
74
75 self.sensor = EcSensor(K=self.constant_k)
**Mycodo > env > lib > python3.7 > site-packages > anyleaf > __init_**
it appears that the whole EcSensor section has been removed from this _init__py version
# Driver for the Anyleaf pH module
from dataclasses import dataclass
from enum import Enum, auto
from typing import Optional, List, Union
import adafruit_ads1x15.ads1115 as ADS
from adafruit_ads1x15.analog_in import AnalogIn
from adafruit_max31865 import MAX31865
from filterpy.kalman import KalmanFilter
from filterpy.common import Q_discrete_white_noise
from . import filter
I found on github the __ini__.py file that does reference EcSensor.
**github > AnyLeaf >anyleaf-python > anyleaf > __init__.py**
This __init__ has the EcSensor starting on line 387
# Driver for the Anyleaf pH module
from dataclasses import dataclass
from enum import Enum, auto
from typing import Optional, List, Union
import struct
import adafruit_ads1x15.ads1115 as ADS
from adafruit_ads1x15.analog_in import AnalogIn
from adafruit_max31865 import MAX31865
from filterpy.kalman import KalmanFilter
from filterpy.common import Q_discrete_white_noise
import serial
from serial.tools import list_ports
from . import filter
…
386 @dataclass
387 class EcSensor:
388 """An interface for our EC module, which communicates a serial message over UART."""
389 ser: serial.Serial
390 K: CellConstant
391 cal: Optional[CalPtEc]
392 excitation_mode: ExcMode
393
394 def __init__(self, K: float=1.0, cal: Optional[CalPtEc]=None, 395 exc_mode=ExcMode.READING_ONLY,
395 uart_location='/dev/serial0'):
396 # Same baud as set in firmware: 9,600.
397 self.ser = serial.Serial(uart_location, 9_600, timeout=10)
398 # self.ser = serial.Serial('/dev/ttyS0', 9_600, timeout=10)
399 # self.ser = serial.Serial('/dev/ttyAMA0', 9_600, timeout=10)
### Expected behavior
### Additional context
It appears that the anyleaf > __init__.py from github just needs to be uploaded to Mycodo.
| closed | 2021-06-17T19:03:07Z | 2021-08-30T02:47:45Z | https://github.com/kizniche/Mycodo/issues/1027 | [] | keefer223 | 1 |
jmcnamara/XlsxWriter | pandas | 810 | Comment size stretched when intervening rows are enlarged by word wrap cells | I am using Python version 3.9.2 and XlsxWriter 1.4.3 and Excel for Mac 2016.
This is very similar to problem #403, but instead can be reproduced by cells stretched in height by word wrap. In this case, the stretching will happen regardless of when the text wrap formatting is applied.
```python
from xlsxwriter import Workbook
comment_text = 'These comments should be identically sized.'
comment_options = {'width': 200, 'height': 30, 'visible': True}
cell_text = 'Fillertext, Fillertext, Fillertext, Fillertext'
with Workbook('comment_test.xlsx') as wb:
ws = wb.add_worksheet()
wrap_text = wb.add_format()
wrap_text.set_text_wrap()
ws.write_string('G5', cell_text, wrap_text)
ws.write_comment('B3', comment_text, comment_options)
ws.write_comment('B6', comment_text, comment_options)
ws.write_comment('G3', comment_text, comment_options)
ws.write_comment('G6', comment_text, comment_options)
```
The resulting worksheet is shown in the image below:
<img width="608" alt="Screen Shot 2021-06-04 at 4 59 29 PM" src="https://user-images.githubusercontent.com/5733291/120862046-475bf400-c556-11eb-9bab-1ed5ce57655b.png">
| closed | 2021-06-04T21:02:28Z | 2021-06-04T23:20:24Z | https://github.com/jmcnamara/XlsxWriter/issues/810 | [] | rlad78 | 1 |
Kav-K/GPTDiscord | asyncio | 181 | /index query results are cut off for larger prompts | If the prompt is large or multiple sentences, the results for /index query run the risk of being cut off:

| closed | 2023-02-27T01:25:07Z | 2023-03-12T03:12:35Z | https://github.com/Kav-K/GPTDiscord/issues/181 | [
"bug",
"help wanted",
"good first issue",
"help-wanted-important"
] | Kav-K | 0 |
jupyter/nbviewer | jupyter | 450 | ipythonblocks not rendering on github.com | Copied from ipython/nbconvert#204.
With GitHub's [new notebook rendering](https://github.com/blog/1995-github-jupyter-notebooks-3) [ipythonblocks](https://github.com/jiffyclub/ipythonblocks) tables aren't rendered correctly as they are in nbviewer. Examples:
GitHub: https://github.com/jiffyclub/ipythonblocks/blob/master/demos/starry_night_to_text.ipynb
nbviewer: http://nbviewer.ipython.org/github/jiffyclub/ipythonblocks/blob/master/demos/starry_night_to_text.ipynb
Looking at the HTML it looks like GitHub is stripping out `style` tags and attributes, and `id` attributes:
GitHub:

nbviewer:

| open | 2015-05-07T21:22:11Z | 2018-07-16T02:34:53Z | https://github.com/jupyter/nbviewer/issues/450 | [
"type:Bug",
"tag:GitHub"
] | jiffyclub | 7 |
InstaPy/InstaPy | automation | 6,104 | Could not pass the login A/B test. Trying last string... | <!-- Did you know that we have a Discord channel ? Join us: https://discord.gg/FDETsht -->
<!-- Is this a Feature Request ? Please, check out our Wiki first https://github.com/timgrossmann/InstaPy/wiki -->
## Expected Behavior
After running the quickstart.py with valid credentials a successful login should happen.
## Current Behavior
INFO [2021-03-04 12:03:08] [...] Cookie file not found, creating cookie...
WARNING [2021-03-04 12:03:13] [...] Login A/B test detected! Trying another string...
WARNING [2021-03-04 12:03:18] [...] Could not pass the login A/B test. Trying last string...
...........................................................................................................................
CRITICAL [2021-03-04 12:03:57] [...] Unable to login to Instagram! You will find more information in the logs above.
## Possible Solution (optional)
Something seems to Pop Up up in the browser window "Save login", "activate notifications" etc. I tried clicking manually.
## InstaPy configuration
Only credentials.
Edit: running on linux with pip install on python 3.9.1 | closed | 2021-03-04T13:43:28Z | 2021-07-21T06:18:36Z | https://github.com/InstaPy/InstaPy/issues/6104 | [
"wontfix"
] | SeironWP | 3 |
explosion/spaCy | deep-learning | 13,228 | displacy.js for NER | https://github.com/explosion/spaCy/blob/e2a3952de51abb2620b4ff799ac461c87fec7bb4/website/docs/usage/visualizers.mdx#L417
I might fail to see where but [displacy.js](https://github.com/explosion/displacy/blob/master/assets/js/displacy.js) doesn't seem to render the NER tags but only dependency arcs. | closed | 2024-01-09T15:45:18Z | 2024-01-16T13:56:05Z | https://github.com/explosion/spaCy/issues/13228 | [
"feat / visualizers"
] | ch-sander | 1 |
nschloe/tikzplotlib | matplotlib | 189 | Manual legend | Hello,
I implemented a modification of the rendering of legends that I hope you will find interesting.
See the diff [here](https://github.com/nschloe/matplotlib2tikz/compare/master...haji-ali:manual-legend)
When passing `manual_legend=True`, the legend is rendered using `\matrix` instead of `legend`. This is much more flexible and allows for easy customization of the legend (for example changing the order to match the order from `matplotlib`).
Also in this commit is what I believe is a fix to color.py that returns `none` when the alpha value is exactly zero.
| closed | 2017-06-29T09:07:22Z | 2019-03-21T10:53:31Z | https://github.com/nschloe/tikzplotlib/issues/189 | [] | haji-ali | 6 |
svc-develop-team/so-vits-svc | pytorch | 163 | [Help]: 推理时音频文件报错 | ### 请勾选下方的确认框。
- [X] 我已仔细阅读[README.md](https://github.com/svc-develop-team/so-vits-svc/blob/4.0/README_zh_CN.md)和[wiki中的Quick solution](https://github.com/svc-develop-team/so-vits-svc/wiki/Quick-solution)。
- [X] 我已通过各种搜索引擎排查问题,我要提出的问题并不常见。
- [X] 我未在使用由第三方用户提供的一键包/环境包。
### 系统平台版本号
colab平台
### GPU 型号
Tesla T4
### Python版本
python3.8.9
### PyTorch版本
1.13.1+cu117
### sovits分支
4.0(默认)
### 数据集来源(用于判断数据集质量)
纯人声语音
### 出现问题的环节或执行的命令
Start inference (and download)
### 问题描述
下载时无法找到文件,应该是上半部分报错的问题,是speaker设置错了吗?
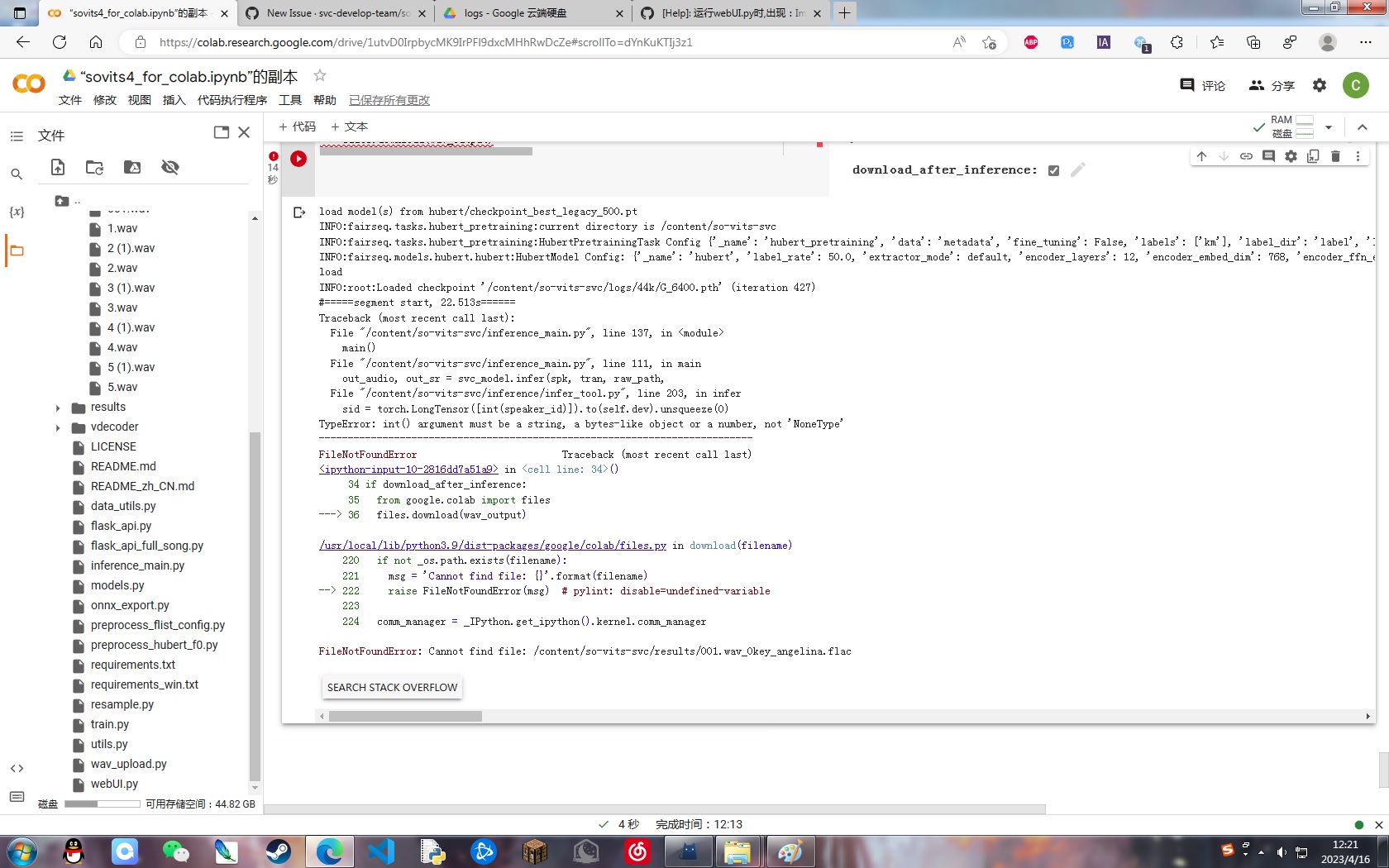
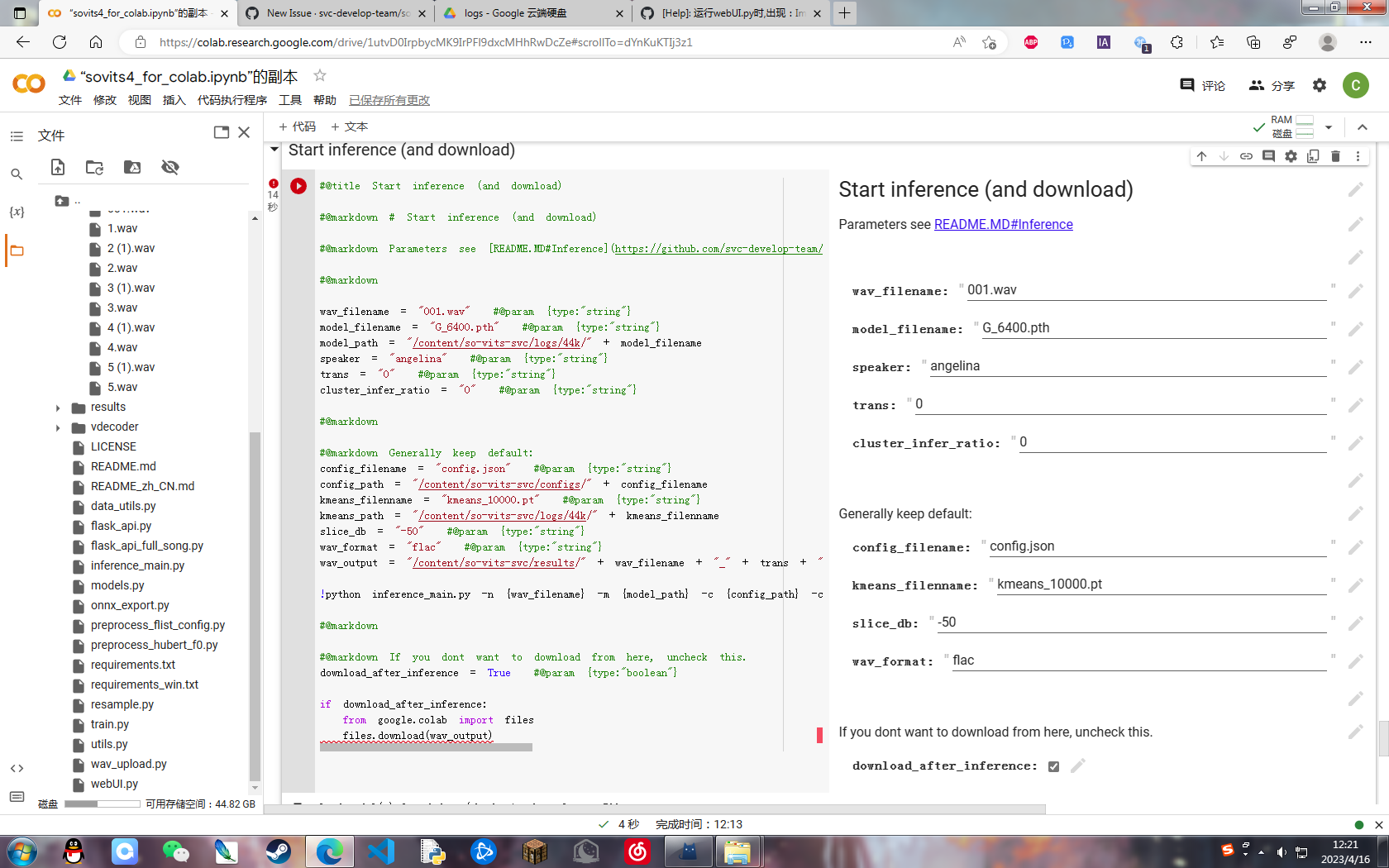
### 日志
```python
load model(s) from hubert/checkpoint_best_legacy_500.pt
INFO:fairseq.tasks.hubert_pretraining:current directory is /content/so-vits-svc
INFO:fairseq.tasks.hubert_pretraining:HubertPretrainingTask Config {'_name': 'hubert_pretraining', 'data': 'metadata', 'fine_tuning': False, 'labels': ['km'], 'label_dir': 'label', 'label_rate': 50.0, 'sample_rate': 16000, 'normalize': False, 'enable_padding': False, 'max_keep_size': None, 'max_sample_size': 250000, 'min_sample_size': 32000, 'single_target': False, 'random_crop': True, 'pad_audio': False}
INFO:fairseq.models.hubert.hubert:HubertModel Config: {'_name': 'hubert', 'label_rate': 50.0, 'extractor_mode': default, 'encoder_layers': 12, 'encoder_embed_dim': 768, 'encoder_ffn_embed_dim': 3072, 'encoder_attention_heads': 12, 'activation_fn': gelu, 'layer_type': transformer, 'dropout': 0.1, 'attention_dropout': 0.1, 'activation_dropout': 0.0, 'encoder_layerdrop': 0.05, 'dropout_input': 0.1, 'dropout_features': 0.1, 'final_dim': 256, 'untie_final_proj': True, 'layer_norm_first': False, 'conv_feature_layers': '[(512,10,5)] + [(512,3,2)] * 4 + [(512,2,2)] * 2', 'conv_bias': False, 'logit_temp': 0.1, 'target_glu': False, 'feature_grad_mult': 0.1, 'mask_length': 10, 'mask_prob': 0.8, 'mask_selection': static, 'mask_other': 0.0, 'no_mask_overlap': False, 'mask_min_space': 1, 'mask_channel_length': 10, 'mask_channel_prob': 0.0, 'mask_channel_selection': static, 'mask_channel_other': 0.0, 'no_mask_channel_overlap': False, 'mask_channel_min_space': 1, 'conv_pos': 128, 'conv_pos_groups': 16, 'latent_temp': [2.0, 0.5, 0.999995], 'skip_masked': False, 'skip_nomask': False, 'checkpoint_activations': False, 'required_seq_len_multiple': 2, 'depthwise_conv_kernel_size': 31, 'attn_type': '', 'pos_enc_type': 'abs', 'fp16': False}
load
INFO:root:Loaded checkpoint '/content/so-vits-svc/logs/44k/G_6400.pth' (iteration 427)
#=====segment start, 22.513s======
Traceback (most recent call last):
File "/content/so-vits-svc/inference_main.py", line 137, in <module>
main()
File "/content/so-vits-svc/inference_main.py", line 111, in main
out_audio, out_sr = svc_model.infer(spk, tran, raw_path,
File "/content/so-vits-svc/inference/infer_tool.py", line 203, in infer
sid = torch.LongTensor([int(speaker_id)]).to(self.dev).unsqueeze(0)
TypeError: int() argument must be a string, a bytes-like object or a number, not 'NoneType'
---------------------------------------------------------------------------
FileNotFoundError Traceback (most recent call last)
<ipython-input-10-2816dd7a51a9> in <cell line: 34>()
34 if download_after_inference:
35 from google.colab import files
---> 36 files.download(wav_output)
/usr/local/lib/python3.9/dist-packages/google/colab/files.py in download(filename)
220 if not _os.path.exists(filename):
221 msg = 'Cannot find file: {}'.format(filename)
--> 222 raise FileNotFoundError(msg) # pylint: disable=undefined-variable
223
224 comm_manager = _IPython.get_ipython().kernel.comm_manager
FileNotFoundError: Cannot find file: /content/so-vits-svc/results/001.wav_0key_angelina.flac
```
### 截图`so-vits-svc`、`logs/44k`文件夹并粘贴到此处
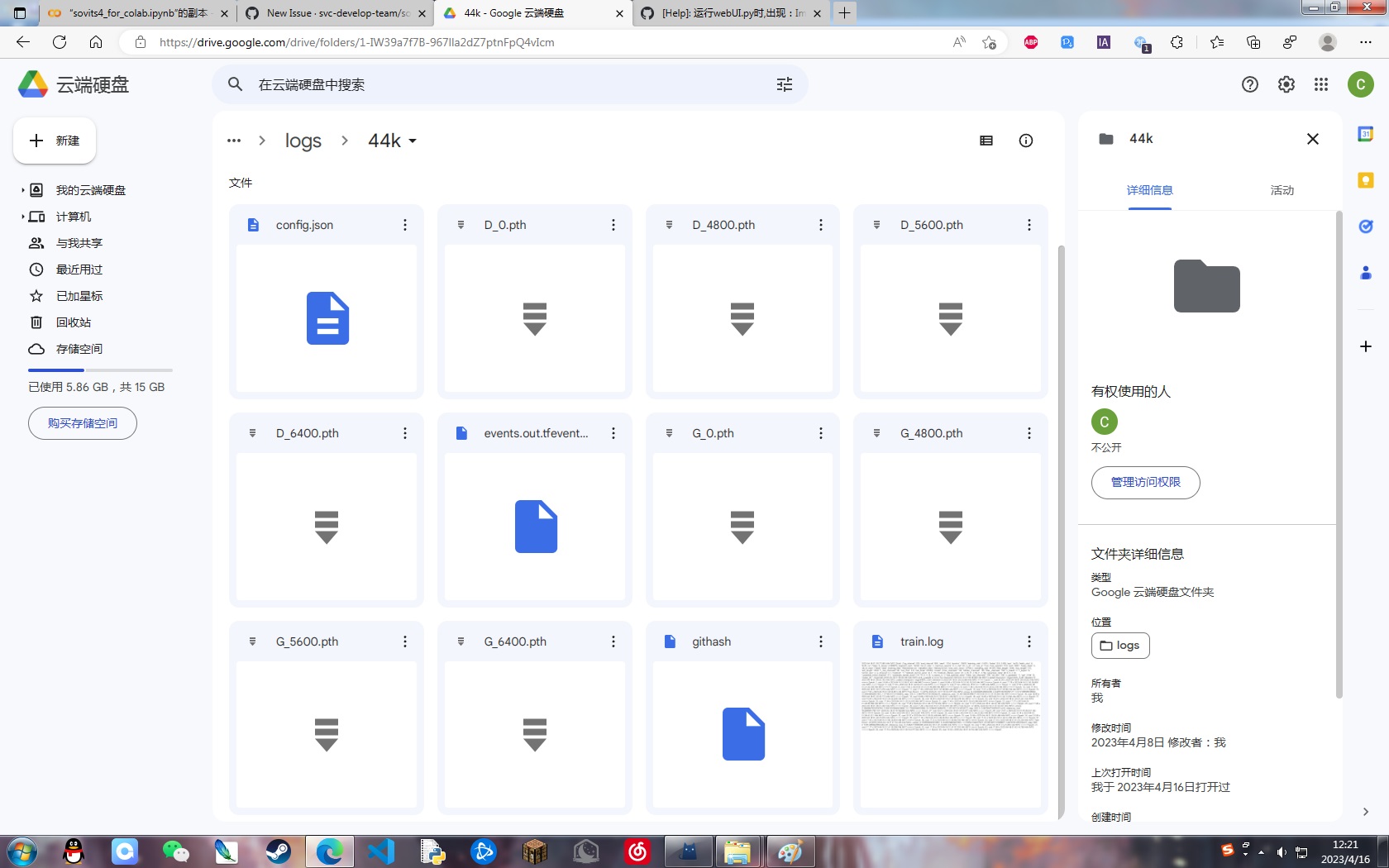
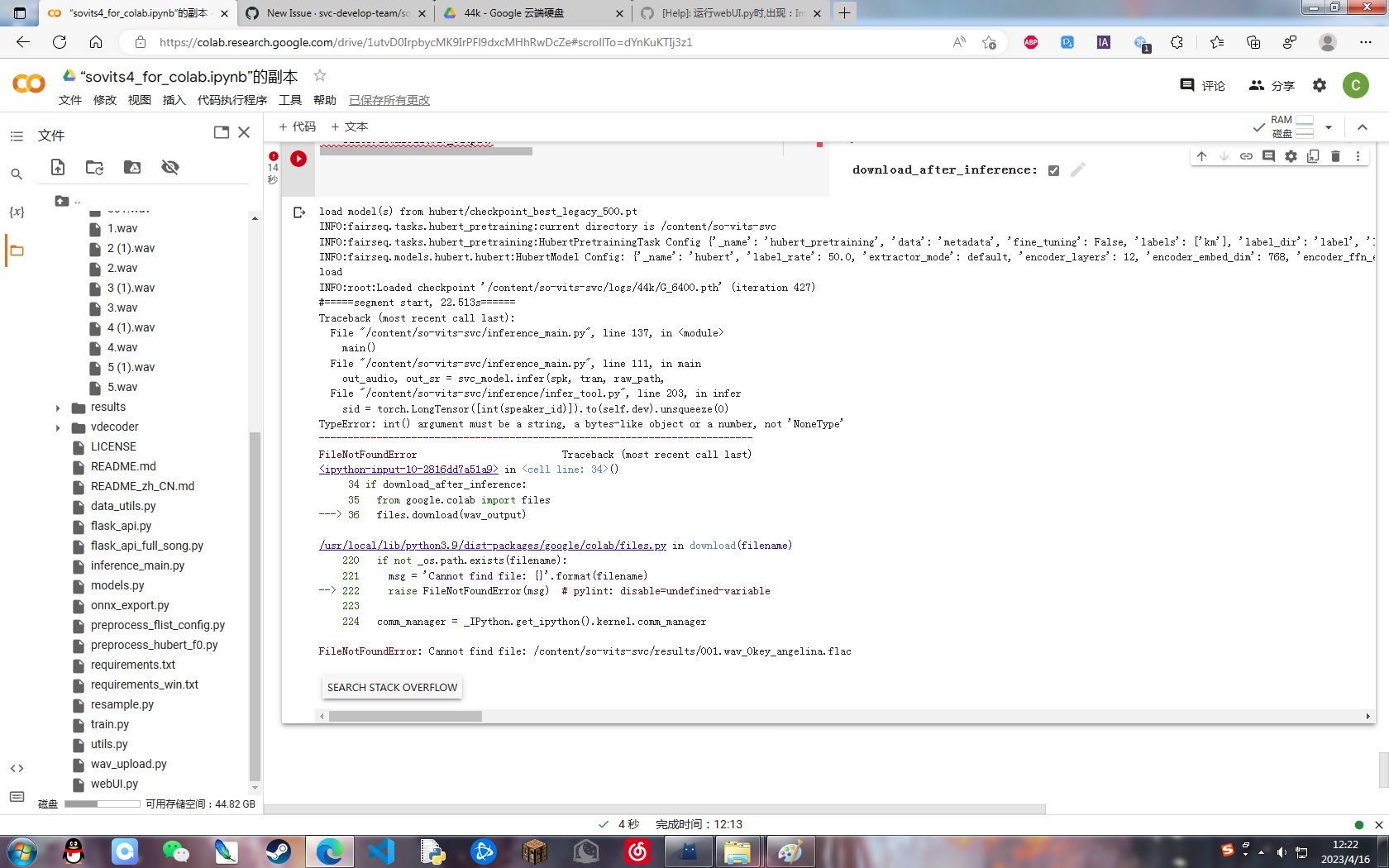
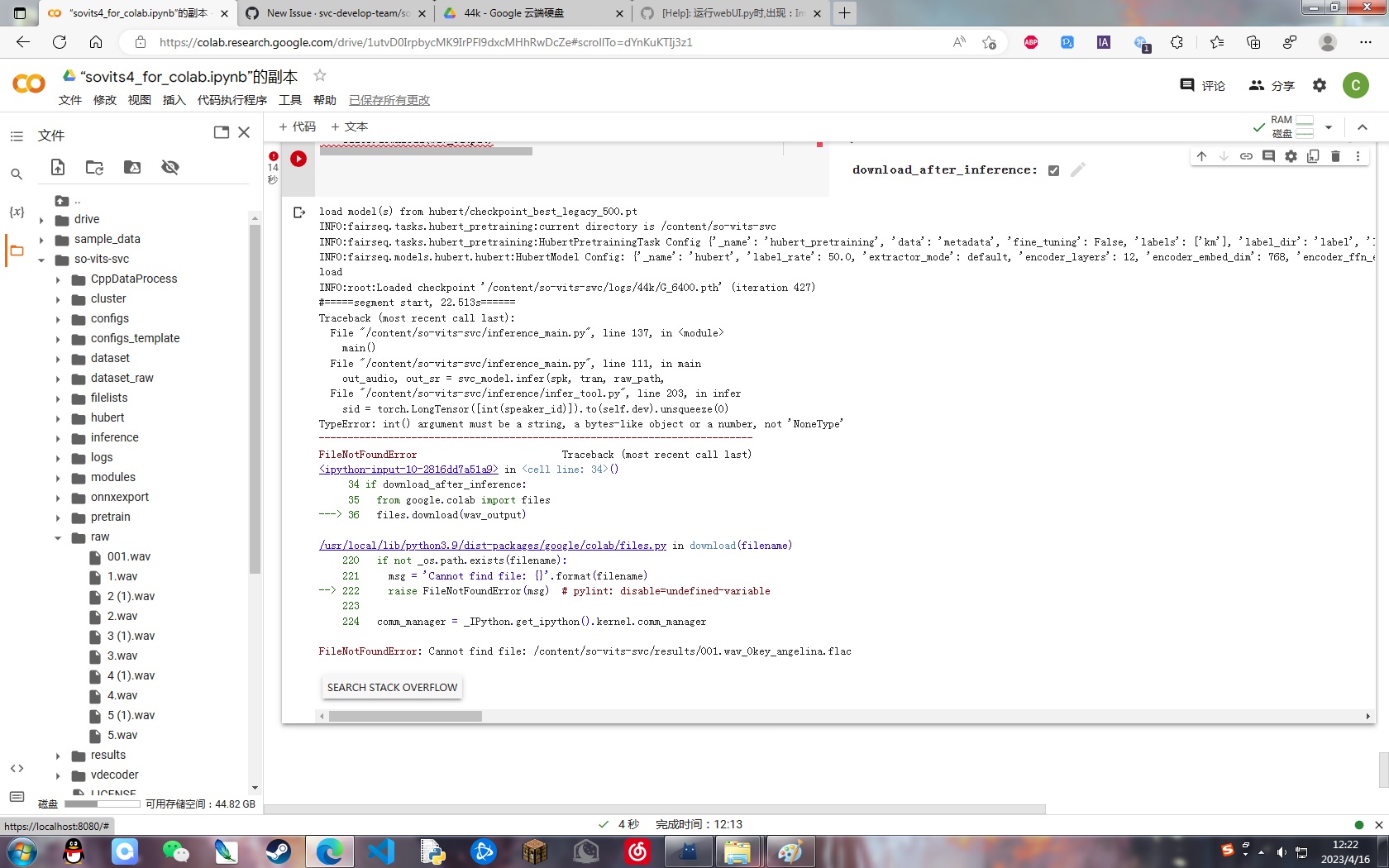
### 补充说明
_No response_ | closed | 2023-04-16T04:23:17Z | 2023-04-17T01:12:32Z | https://github.com/svc-develop-team/so-vits-svc/issues/163 | [
"help wanted"
] | bilbillm | 1 |
sktime/pytorch-forecasting | pandas | 1,283 | info on validation methods for TFT | Hello everyone! I'm new to the library and I'm trying to understand how to approach the validation part.
Let's say I want a forward chaining cross validation. To be clear, here an example. There is an initial amount of data to train on, let's say five folds in this example, and then you evaluate on the sixth fold and save that performance metric. You re-train now on the first six folds and evaluate on the seventh. You repeat until all folds are exhausted and again take the average of your performance metric. The folds using this technique would look like this:
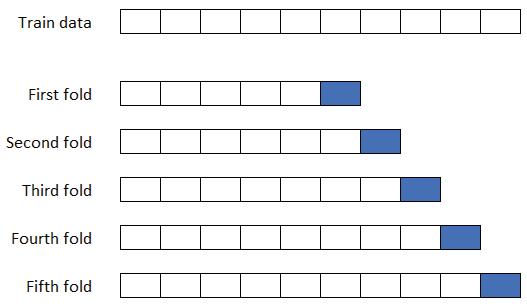
To run something like this, I assume I need a code like this, correect?
```
training = TimeSeriesDataSet(
data[lambda x: x.time_idx <= training_cutoff],
min_prediction_idx = training_cutoff_idx
.............
)
# create validation set (predict=True) which means to predict the last max_prediction_length points in time
# for each series
validation = TimeSeriesDataSet.from_dataset(training, data, predict=False, stop_randomization=True)
# create dataloaders for model
batch_size = 128 # set this between 32 to 128
train_dataloader = training.to_dataloader(train=True, batch_size=batch_size, num_workers=0)
val_dataloader = validation.to_dataloader(train=False, batch_size=batch_size * 10, num_workers=0)
```
Is my approach correct or am I missing something? **Basically what I need to do is set the ''predict'' from true to false and set the right min_prediction_idx, or is it more than that**?
**Or am I supposed to do that by myself with a loop?**
| open | 2023-03-31T19:37:19Z | 2023-04-18T07:28:36Z | https://github.com/sktime/pytorch-forecasting/issues/1283 | [] | ianux22 | 1 |
langmanus/langmanus | automation | 80 | Browser agent error | Query: Search OCR, object detection, and instance segmentation models and based on scenarios such as training, fine-tuning, and inference, provide a summary report on GPU consumption required by common small models. Note that the training cost is measured by multiplying the number of NVIDIA V100 32 GB graphics cards by the training or inference time.
env.
```
# Reasoning LLM (for complex reasoning tasks)
REASONING_API_KEY=EMPTY
REASONING_BASE_URL=http://localhost:8000/v1
REASONING_MODEL=qwq32b
# Non-reasoning LLM (for straightforward tasks)
BASIC_API_KEY=EMPTY
BASIC_BASE_URL=http://localhost:8001/v1
BASIC_MODEL=qwen2.57b
# Vision-language LLM (for tasks requiring visual understanding)
VL_API_KEY=EMPTY
VL_BASE_URL=http://localhost:8002/v1
VL_MODEL=qwen2.5vl7b
```
```
DEBUG [src.graph.nodes] Supervisor response: {'next': 'browser'}
INFO [src.graph.nodes] Supervisor delegating to: browser
INFO [src.graph.nodes] Browser agent starting task
DEBUG [src.tools.decorators] Tool BrowserTool._run called with parameters: instruction=Go to https://epoch.ai/data/all_ai_models.csv
INFO [agent] 🚀 Starting task: Go to https://epoch.ai/data/all_ai_models.csv
INFO [agent]
📍 Step 1
ERROR [browser] Failed to initialize Playwright browser: BrowserType.launch: Target page, context or browser has been closed
Browser logs:
╔════════════════════════════════════════════════════════════════════════════════════════════════╗
║ Looks like you launched a headed browser without having a XServer running. ║
║ Set either 'headless: true' or use 'xvfb-run <your-playwright-app>' before running Playwright. ║
║ ║
║ <3 Playwright Team ║
╚════════════════════════════════════════════════════════════════════════════════════════════════╝
Call log:
- <launching> /root/.cache/ms-playwright/chromium-1155/chrome-linux/chrome --disable-field-trial-config --disable-background-networking --disable-background-timer-throttling --disable-backgrounding-occluded-windows --disable-back-forward-cache --disable-breakpad --d
isable-client-side-phishing-detection --disable-component-extensions-with-background-pages --disable-component-update --no-default-browser-check --disable-default-apps --disable-dev-shm-usage --disable-extensions --disable-features=ImprovedCookieControls,LazyFrameLoad
ing,GlobalMediaControls,DestroyProfileOnBrowserClose,MediaRouter,DialMediaRouteProvider,AcceptCHFrame,AutoExpandDetailsElement,CertificateTransparencyComponentUpdater,AvoidUnnecessaryBeforeUnloadCheckSync,Translate,HttpsUpgrades,PaintHolding,ThirdPartyStoragePartition
ing,LensOverlay,PlzDedicatedWorker --allow-pre-commit-input --disable-hang-monitor --disable-ipc-flooding-protection --disable-popup-blocking --disable-prompt-on-repost --disable-renderer-backgrounding --force-color-profile=srgb --metrics-recording-only --no-first-run
--enable-automation --password-store=basic --use-mock-keychain --no-service-autorun --export-tagged-pdf --disable-search-engine-choice-screen --unsafely-disable-devtools-self-xss-warnings --no-sandbox --no-sandbox --disable-blink-features=AutomationControlled --disab
le-infobars --disable-background-timer-throttling --disable-popup-blocking --disable-backgrounding-occluded-windows --disable-renderer-backgrounding --disable-window-activation --disable-focus-on-load --no-first-run --no-default-browser-check --no-startup-window --win
dow-position=0,0 --disable-web-security --disable-site-isolation-trials --disable-features=IsolateOrigins,site-per-process --user-data-dir=/tmp/playwright_chromiumdev_profile-AyLoZM --remote-debugging-pipe --no-startup-window
- - <launched> pid=4044115
- - [pid=4044115][err] [4044115:4044115:0320/195144.093934:ERROR:ozone_platform_x11.cc(245)] Missing X server or $DISPLAY
- - [pid=4044115][err] [4044115:4044115:0320/195144.093972:ERROR:env.cc(257)] The platform failed to initialize. Exiting.
WARNING [browser] Page load failed, continuing...
ERROR [browser] Failed to initialize Playwright browser: BrowserType.launch: Target page, context or browser has been closed
Browser logs:
╔════════════════════════════════════════════════════════════════════════════════════════════════╗
║ Looks like you launched a headed browser without having a XServer running. ║
║ Set either 'headless: true' or use 'xvfb-run <your-playwright-app>' before running Playwright. ║
║ ║
║ <3 Playwright Team ║
╚════════════════════════════════════════════════════════════════════════════════════════════════╝
Call log:
- <launching> /root/.cache/ms-playwright/chromium-1155/chrome-linux/chrome --disable-field-trial-config --disable-background-networking --disable-background-timer-throttling --disable-backgrounding-occluded-windows --disable-back-forward-cache --disable-breakpad --d
isable-client-side-phishing-detection --disable-component-extensions-with-background-pages --disable-component-update --no-default-browser-check --disable-default-apps --disable-dev-shm-usage --disable-extensions --disable-features=ImprovedCookieControls,LazyFrameLoad
ing,GlobalMediaControls,DestroyProfileOnBrowserClose,MediaRouter,DialMediaRouteProvider,AcceptCHFrame,AutoExpandDetailsElement,CertificateTransparencyComponentUpdater,AvoidUnnecessaryBeforeUnloadCheckSync,Translate,HttpsUpgrades,PaintHolding,ThirdPartyStoragePartition
ing,LensOverlay,PlzDedicatedWorker --allow-pre-commit-input --disable-hang-monitor --disable-ipc-flooding-protection --disable-popup-blocking --disable-prompt-on-repost --disable-renderer-backgrounding --force-color-profile=srgb --metrics-recording-only --no-first-run
--enable-automation --password-store=basic --use-mock-keychain --no-service-autorun --export-tagged-pdf --disable-search-engine-choice-screen --unsafely-disable-devtools-self-xss-warnings --no-sandbox --no-sandbox --disable-blink-features=AutomationControlled --disab
le-infobars --disable-background-timer-throttling --disable-popup-blocking --disable-backgrounding-occluded-windows --disable-renderer-backgrounding --disable-window-activation --disable-focus-on-load --no-first-run --no-default-browser-check --no-startup-window --win
dow-position=0,0 --disable-web-security --disable-site-isolation-trials --disable-features=IsolateOrigins,site-per-process --user-data-dir=/tmp/playwright_chromiumdev_profile-YnbrnR --remote-debugging-pipe --no-startup-window
- - <launched> pid=4044156
- - [pid=4044156][err] [4044156:4044156:0320/195144.562060:ERROR:ozone_platform_x11.cc(245)] Missing X server or $DISPLAY
- - [pid=4044156][err] [4044156:4044156:0320/195144.562094:ERROR:env.cc(257)] The platform failed to initialize. Exiting.
ERROR [agent] ❌ Result failed 1/3 times:
BrowserType.launch: Target page, context or browser has been closed
```
But my XServe is running. Or not ?
```
+-----------------------------------------------------------------------------------------+
| Processes: |
| GPU GI CI PID Type Process name GPU Memory |
| ID ID Usage |
|=========================================================================================|
| 0 N/A N/A 46318 C ...aconda3/envs/openblas/bin/python3.9 320MiB |
| 0 N/A N/A 3891547 C ...t/anaconda3/envs/vllm080/bin/python 88770MiB |
| 0 N/A N/A 4039561 G /usr/lib/xorg/Xorg 106MiB |
| 1 N/A N/A 3892292 C ...t/anaconda3/envs/vllm080/bin/python 49512MiB |
| 1 N/A N/A 4039561 G /usr/lib/xorg/Xorg 4MiB |
| 2 N/A N/A 3893033 C ...t/anaconda3/envs/vllm080/bin/python 48948MiB |
| 2 N/A N/A 4039561 G /usr/lib/xorg/Xorg 4MiB |
| 3 N/A N/A 3913871 C ...naconda3/envs/yuanhao310/bin/python 92716MiB |
| 3 N/A N/A 4039561 G /usr/lib/xorg/Xorg 4MiB |
| 4 N/A N/A 3913872 C ...naconda3/envs/yuanhao310/bin/python 92730MiB |
| 4 N/A N/A 4039561 G /usr/lib/xorg/Xorg 4MiB |
| 5 N/A N/A 4039561 G /usr/lib/xorg/Xorg 4MiB |
| 6 N/A N/A 46318 C ...aconda3/envs/openblas/bin/python3.9 2070MiB |
| 6 N/A N/A 4039561 G /usr/lib/xorg/Xorg 4MiB |
| 7 N/A N/A 4039561 G /usr/lib/xorg/Xorg 4MiB |
+-----------------------------------------------------------------------------------------+
```
| closed | 2025-03-20T12:09:57Z | 2025-03-21T06:33:37Z | https://github.com/langmanus/langmanus/issues/80 | [] | liuruijin17 | 0 |
ijl/orjson | numpy | 333 | Datetime serialization precision | Hi Folks,
Due to compatibility with other languages, we only need milliseconds instead of microseconds in the serialized datetime.
What do you think if we added a few options for specifications of time accuracy? Something like OPT_DT_PRECISION_S, OPT_DT_PRECISION_MS, OPT_DT_PRECISION_US instead of OPT_OMIT_MICROSECONDS?
@M0dEx is keen to implement such functionality, | closed | 2023-01-12T15:22:37Z | 2023-05-07T17:57:17Z | https://github.com/ijl/orjson/issues/333 | [] | lejmr | 6 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.