repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
wkentaro/labelme | deep-learning | 1,021 | When changing the label value with label edit, the program crashes. | app.py (1171 row)
item = self.uniqLabelList.findItemsByLabel(label)[0]
there is out of range error.
add code.. good working.
def _get_rgb_by_label(self, label):
if self._config["shape_color"] == "auto":
if not self.uniqLabelList.findItemsByLabel(label):
item = QtWidgets.QListWidgetItem()
item.setData(Qt.UserRole, label)
self.uniqLabelList.addItem(item)
item = self.uniqLabelList.findItemsByLabel(label)[0] | closed | 2022-05-20T09:09:10Z | 2022-09-26T13:35:54Z | https://github.com/wkentaro/labelme/issues/1021 | [
"issue::bug",
"priority: high"
] | sangwoo89 | 1 |
feature-engine/feature_engine | scikit-learn | 493 | Ignore NaNs before using `OrdinalEncoder` | *Is your feature request related to a problem?*
`OrdinalEncoder` should accept nulls. Sometimes you don't want to impute directly but using Imputing Options of XGBoost, LightGBM or CatBoost. Because of the constraints of always Impute before this is currently not possible.
*Describe the solution you'd like*
I like that Feature Engine forces you to Impute first, but I will add some kind of default flag `ignore_nan=False` in case we want to use other imputation afterwards.
Hope you find this helpful. | closed | 2022-08-06T02:31:08Z | 2023-01-14T11:22:45Z | https://github.com/feature-engine/feature_engine/issues/493 | [
"urgent",
"priority"
] | datacubeR | 6 |
seleniumbase/SeleniumBase | pytest | 3,602 | LocalStorage variable deleted after refreshing page | Hello, Michael. Why does local storage variable, which i've set via set_local_storage_item, after refreshing page is deleted?
```
with SB(
uc=True,
browser="chrome",
locale="ru",
position="0,0",
agent=browser_config.get("user_agent"),
window_size=browser_config.get("window_size"),
ad_block=True,
devtools=True
) as driver:
```
```
def login(driver: SeleniumBase, token: str):
logger.info("Execute token to LocalStorage")
temp = json.dumps({"token": token})
driver.set_local_storage_item("wbx__tokenData", temp)
```
While we were working with raw selenium we did it like this:
```
driver.execute_script("localStorage.setItem(arguments[0], JSON.stringify({token: arguments[1]}));",
'wbx__tokenData', token)
```
And it were working fine.
_Originally posted by @leesache in https://github.com/seleniumbase/SeleniumBase/discussions/2722#discussioncomment-12462594_ | closed | 2025-03-11T14:19:29Z | 2025-03-12T06:34:54Z | https://github.com/seleniumbase/SeleniumBase/issues/3602 | [
"question",
"UC Mode / CDP Mode"
] | leesache | 1 |
microsoft/nni | data-science | 5,218 | I am very new to NNI can you please provide a MS NNI + Pytorch Lightning example for HPO | **Describe the issue**:
I am very new to NNI can you please provide a MS NNI + Pytorch Lightning example for HPO
**Environment**:
- NNI version:
- Training service (local|remote|pai|aml|etc):
- Client OS:
- Server OS (for remote mode only):
- Python version:
- PyTorch/TensorFlow version:
- Is conda/virtualenv/venv used?:
- Is running in Docker?:
**Configuration**:
- Experiment config (remember to remove secrets!):
- Search space:
**Log message**:
- nnimanager.log:
- dispatcher.log:
- nnictl stdout and stderr:
<!--
Where can you find the log files:
LOG: https://github.com/microsoft/nni/blob/master/docs/en_US/Tutorial/HowToDebug.md#experiment-root-director
STDOUT/STDERR: https://nni.readthedocs.io/en/stable/reference/nnictl.html#nnictl-log-stdout
-->
**How to reproduce it?**: | open | 2022-11-10T18:58:02Z | 2022-11-24T01:23:03Z | https://github.com/microsoft/nni/issues/5218 | [
"feature request"
] | sathishkumark27 | 0 |
OFA-Sys/Chinese-CLIP | computer-vision | 20 | import cn_clip出错UnicodeDecodeError: 'gbk' codec can't decode byte 0x81 in position 1564: illegal multibyte sequence | import cn_clip.clip as clip
发生异常: UnicodeDecodeError
Traceback (most recent call last):
File "D:\develop\anaconda3\lib\runpy.py", line 193, in _run_module_as_main
"__main__", mod_spec)
File "D:\develop\anaconda3\lib\runpy.py", line 85, in _run_code
exec(code, run_globals)
File "c:\Users\saizong\.vscode\extensions\ms-python.python-2022.4.1\pythonFiles\lib\python\debugpy\__main__.py", line 45, in <module>
cli.main()
File "c:\Users\saizong\.vscode\extensions\ms-python.python-2022.4.1\pythonFiles\lib\python\debugpy/..\debugpy\server\cli.py", line 444, in main
run()
File "c:\Users\saizong\.vscode\extensions\ms-python.python-2022.4.1\pythonFiles\lib\python\debugpy/..\debugpy\server\cli.py", line 285, in run_file
runpy.run_path(target_as_str, run_name=compat.force_str("__main__"))
File "D:\develop\anaconda3\lib\runpy.py", line 263, in run_path
pkg_name=pkg_name, script_name=fname)
File "D:\develop\anaconda3\lib\runpy.py", line 96, in _run_module_code
mod_name, mod_spec, pkg_name, script_name)
File "D:\develop\anaconda3\lib\runpy.py", line 85, in _run_code
exec(code, run_globals)
File "d:\develop\workspace\today_video\clipcn.py", line 5, in <module>
import cn_clip.clip as clip
File "D:\develop\anaconda3\Lib\site-packages\cn_clip\clip\__init__.py", line 3, in <module>
_tokenizer = FullTokenizer()
File "D:\develop\anaconda3\Lib\site-packages\cn_clip\clip\bert_tokenizer.py", line 170, in __init__
self.vocab = load_vocab(vocab_file)
File "D:\develop\anaconda3\Lib\site-packages\cn_clip\clip\bert_tokenizer.py", line 132, in load_vocab
token = convert_to_unicode(reader.readline())
UnicodeDecodeError: 'gbk' codec can't decode byte 0x81 in position 1564: illegal multibyte sequence
请问如何处理?谢谢! | closed | 2022-11-28T08:07:28Z | 2022-12-13T11:37:28Z | https://github.com/OFA-Sys/Chinese-CLIP/issues/20 | [] | bigmarten | 13 |
allenai/allennlp | data-science | 5,164 | How can i add a new subcommand? | I'm writing a transfer learning(domain adaptation) method in Allennlp, so I created a trainer by myself, and I wanted to write a new command( for example mytrain or mypredict) to begin the training and predict process.
How can I use the command: allennlp mytrain xxx.jsonnet --including-package xxx --serialization-dir xxxx ? | closed | 2021-04-28T13:34:03Z | 2021-05-06T02:58:18Z | https://github.com/allenai/allennlp/issues/5164 | [] | ding00 | 4 |
hack4impact/flask-base | sqlalchemy | 202 | Docker file for deployment | I am using this as a part of a larger project and found we need to deploy it with docker,
I'm working on this now, and will open a pull request with the patch, I'll try to make it asap. | closed | 2020-05-29T17:26:32Z | 2020-06-05T04:26:39Z | https://github.com/hack4impact/flask-base/issues/202 | [] | MohammedRashad | 1 |
openapi-generators/openapi-python-client | fastapi | 351 | Support non-file fields in Multipart requests | **Is your feature request related to a problem? Please describe.**
Given this schema for a POST request:
```
{
"requestBody": {
"content": {
"multipart/form-data": {
"schema": {
"type": "object",
"properties": {
"file": {
"type": "string",
"format": "binary"
},
"options": {
"type": "string",
"default": "{}"
}
}
}
}
}
}
}
```
the generated code will call `httpx.post(files=dict(file=file, options=options))` which causes a server-side pydantic validation error (`options` should be of type `str`).
**Describe the solution you'd like**
The correct invocation would be `httpx.post(data=dict(options=options), files=dict(file=file))` (see [relevant page from httpx docs](https://www.python-httpx.org/advanced/#multipart-file-encoding))
**Describe alternatives you've considered**
Some sort of server-side workaround, but it might be somewhat ugly.
| closed | 2021-03-19T17:08:08Z | 2021-03-22T20:25:52Z | https://github.com/openapi-generators/openapi-python-client/issues/351 | [
"✨ enhancement"
] | csymeonides-mf | 1 |
ultralytics/ultralytics | computer-vision | 18,748 | yolo11 cls train error | ### Search before asking
- [x] I have searched the Ultralytics YOLO [issues](https://github.com/ultralytics/ultralytics/issues) and found no similar bug report.
### Ultralytics YOLO Component
_No response_
### Bug
from ultralytics import YOLO
# Load a model
model = YOLO("yolo11n-cls.yaml") # build a new model from YAML
model = YOLO("yolo11n-cls.pt") # load a pretrained model (recommended for training)
model = YOLO("yolo11n-cls.yaml").load("yolo11n-cls.pt") # build from YAML and transfer weights
# Train the model
results = model.train(data="mnist160", epochs=100, imgsz=64)
error:
train: G:\Item_done\yolo\yolo5\yolov11\ultralytics-main\datasets\mnist160\train... found 80 images in 10 classes ✅
val: None...
test: G:\Item_done\yolo\yolo5\yolov11\ultralytics-main\datasets\mnist160\test... found 80 images in 10 classes ✅
Overriding model.yaml nc=80 with nc=10
from n params module arguments
0 -1 1 464 ultralytics.nn.modules.conv.Conv [3, 16, 3, 2]
1 -1 1 4672 ultralytics.nn.modules.conv.Conv [16, 32, 3, 2]
2 -1 1 6640 ultralytics.nn.modules.block.C3k2 [32, 64, 1, False, 0.25]
3 -1 1 36992 ultralytics.nn.modules.conv.Conv [64, 64, 3, 2]
4 -1 1 26080 ultralytics.nn.modules.block.C3k2 [64, 128, 1, False, 0.25]
5 -1 1 147712 ultralytics.nn.modules.conv.Conv [128, 128, 3, 2]
6 -1 1 87040 ultralytics.nn.modules.block.C3k2 [128, 128, 1, True]
7 -1 1 295424 ultralytics.nn.modules.conv.Conv [128, 256, 3, 2]
8 -1 1 346112 ultralytics.nn.modules.block.C3k2 [256, 256, 1, True]
9 -1 1 249728 ultralytics.nn.modules.block.C2PSA [256, 256, 1]
10 -1 1 343050 ultralytics.nn.modules.head.Classify [256, 10]
YOLO11n-cls summary: 151 layers, 1,543,914 parameters, 1,543,914 gradients, 3.3 GFLOPs
Transferred 234/236 items from pretrained weights
TensorBoard: Start with 'tensorboard --logdir runs\classify\train10', view at http://localhost:6006/
AMP: running Automatic Mixed Precision (AMP) checks...
AMP: checks passed ✅
train: Scanning G:\Item_done\yolo\yolo5\yolov11\ultralytics-main\datasets\mnist160\train... 80 images, 0 corrupt: 100%|██████████| 80/80 [00:00<?, ?it/s]
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\multiprocessing\spawn.py", line 116, in spawn_main
exitcode = _main(fd, parent_sentinel)
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\multiprocessing\spawn.py", line 125, in _main
prepare(preparation_data)
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\multiprocessing\spawn.py", line 236, in prepare
_fixup_main_from_path(data['init_main_from_path'])
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\multiprocessing\spawn.py", line 287, in _fixup_main_from_path
main_content = runpy.run_path(main_path,
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\runpy.py", line 288, in run_path
return _run_module_code(code, init_globals, run_name,
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\runpy.py", line 97, in _run_module_code
_run_code(code, mod_globals, init_globals,
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\runpy.py", line 87, in _run_code
exec(code, run_globals)
File "G:\Item_done\yolo\yolo5\yolov11\ultralytics-train\cls_train.py", line 9, in <module>
results = model.train(data="mnist160", epochs=100, imgsz=64)
File "G:\Item_done\yolo\yolo5\yolov11\ultralytics-train\ultralytics\engine\model.py", line 806, in train
self.trainer.train()
File "G:\Item_done\yolo\yolo5\yolov11\ultralytics-train\ultralytics\engine\trainer.py", line 207, in train
self._do_train(world_size)
File "G:\Item_done\yolo\yolo5\yolov11\ultralytics-train\ultralytics\engine\trainer.py", line 323, in _do_train
self._setup_train(world_size)
File "G:\Item_done\yolo\yolo5\yolov11\ultralytics-train\ultralytics\engine\trainer.py", line 287, in _setup_train
self.train_loader = self.get_dataloader(self.trainset, batch_size=batch_size, rank=LOCAL_RANK, mode="train")
File "G:\Item_done\yolo\yolo5\yolov11\ultralytics-train\ultralytics\models\yolo\classify\train.py", line 84, in get_dataloader
loader = build_dataloader(dataset, batch_size, self.args.workers, rank=rank)
File "G:\Item_done\yolo\yolo5\yolov11\ultralytics-train\ultralytics\data\build.py", line 143, in build_dataloader
return InfiniteDataLoader(
File "G:\Item_done\yolo\yolo5\yolov11\ultralytics-train\ultralytics\data\build.py", line 39, in __init__
self.iterator = super().__iter__()
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\site-packages\torch\utils\data\dataloader.py", line 441, in __iter__
return self._get_iterator()
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\site-packages\torch\utils\data\dataloader.py", line 388, in _get_iterator
return _MultiProcessingDataLoaderIter(self)
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\site-packages\torch\utils\data\dataloader.py", line 1042, in __init__
w.start()
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\multiprocessing\process.py", line 121, in start
self._popen = self._Popen(self)
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\multiprocessing\context.py", line 224, in _Popen
return _default_context.get_context().Process._Popen(process_obj)
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\multiprocessing\context.py", line 327, in _Popen
return Popen(process_obj)
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\multiprocessing\popen_spawn_win32.py", line 45, in __init__
prep_data = spawn.get_preparation_data(process_obj._name)
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\multiprocessing\spawn.py", line 154, in get_preparation_data
_check_not_importing_main()
File "C:\Users\lllstandout\.conda\envs\yolov8\lib\multiprocessing\spawn.py", line 134, in _check_not_importing_main
raise RuntimeError('''
RuntimeError:
An attempt has been made to start a new process before the
current process has finished its bootstrapping phase.
This probably means that you are not using fork to start your
child processes and you have forgotten to use the proper idiom
in the main module:
if __name__ == '__main__':
freeze_support()
...
The "freeze_support()" line can be omitted if the program
is not going to be frozen to produce an executable.
Exception ignored in: <function InfiniteDataLoader.__del__ at 0x000001F137A2FE50>
Traceback (most recent call last):
File "G:\Item_done\yolo\yolo5\yolov11\ultralytics-train\ultralytics\data\build.py", line 52, in __del__
if hasattr(self.iterator, "_workers"):
AttributeError: 'InfiniteDataLoader' object has no attribute 'iterator'
### Environment
windows10
python 3.8
conda
### Minimal Reproducible Example
from ultralytics import YOLO
# Load a model
model = YOLO("yolo11n-cls.yaml") # build a new model from YAML
model = YOLO("yolo11n-cls.pt") # load a pretrained model (recommended for training)
model = YOLO("yolo11n-cls.yaml").load("yolo11n-cls.pt") # build from YAML and transfer weights
# Train the model
results = model.train(data="mnist160", epochs=100, imgsz=64)
### Additional
_No response_
### Are you willing to submit a PR?
- [ ] Yes I'd like to help by submitting a PR! | closed | 2025-01-18T08:18:08Z | 2025-02-23T05:04:40Z | https://github.com/ultralytics/ultralytics/issues/18748 | [
"classify"
] | xinsuinizhuan | 4 |
adamerose/PandasGUI | pandas | 3 | MultiIndex names are not shown | When showing a DataFrame with a MultiIndex the index columns in the gui are all labeled None even though the index labels have names. | closed | 2019-06-16T02:14:46Z | 2019-06-17T18:50:07Z | https://github.com/adamerose/PandasGUI/issues/3 | [
"bug"
] | selasley | 3 |
deepspeedai/DeepSpeed | deep-learning | 6,634 | [BUG] Deepspeed MoE Multi-GPU Expert-Parallel Training/Inference freezes when a process finishes generation first. | **Describe the bug**
I am using expert-parallel MoE implementation of Deepspeed-MoE for decoder model inference using Huggingface Trainer.
During inference, if the generation lengths of the evaluation dataset distributed among the GPUs do not match,
one of the GPUs will finish the decoding inference for its batch, while the other GPU keeps running.
Because one of the GPU has finished the batch, it will not run.
The remaining running GPU will keep waiting for communication with the finished process.
Is there a way to deal with this other than feeding dummy inputs to the GPU that finishes early?
Forcing some generation lenghts could be a quick solution, too.
Has anyone dealt with this type of problem before?
**To Reproduce**
Run multi-gpu expert parallel inference without forcing any generation lengths
**Expected behavior**
Inference to run smoothly.
**ds_report output**
Don't think it's an issue with my environment, but here goes.
async_io ............... [NO] ....... [NO]
fused_adam ............. [NO] ....... [OKAY]
cpu_adam ............... [NO] ....... [OKAY]
cpu_adagrad ............ [NO] ....... [OKAY]
cpu_lion ............... [NO] ....... [OKAY]
evoformer_attn ......... [NO] ....... [OKAY]
fp_quantizer ........... [NO] ....... [OKAY]
fused_lamb ............. [NO] ....... [OKAY]
fused_lion ............. [NO] ....... [OKAY]
gds .................... [NO] ....... [NO]
transformer_inference .. [NO] ....... [OKAY]
inference_core_ops ..... [NO] ....... [OKAY]
cutlass_ops ............ [NO] ....... [OKAY]
quantizer .............. [NO] ....... [OKAY]
ragged_device_ops ...... [NO] ....... [OKAY]
ragged_ops ............. [NO] ....... [OKAY]
random_ltd ............. [NO] ....... [OKAY]
[WARNING] sparse_attn requires a torch version >= 1.5 and < 2.0 but detected 2.4
[WARNING] using untested triton version (3.0.0), only 1.0.0 is known to be compatible
sparse_attn ............ [NO] ....... [NO]
spatial_inference ...... [NO] ....... [OKAY]
transformer ............ [NO] ....... [OKAY]
stochastic_transformer . [NO] ....... [OKAY]
**Screenshots**

 | closed | 2024-10-17T05:53:47Z | 2024-11-01T21:18:16Z | https://github.com/deepspeedai/DeepSpeed/issues/6634 | [
"bug",
"training"
] | taehyunzzz | 1 |
mirumee/ariadne | api | 359 | Cannot import name 'GraphQLNamedType' | I am using version 0.11.0 of ariadne with FastAPI (0.54.1).
When I launch the server, I have a systematic error of which here is the stack.
```
uvicorn app.main:app --reload
INFO: Uvicorn running on http://127.0.0.1:8000 (Press CTRL+C to quit)
INFO: Started reloader process [7818]
Process SpawnProcess-1:
Traceback (most recent call last):
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/multiprocessing/process.py", line 315, in _bootstrap
self.run()
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/multiprocessing/process.py", line 108, in run
self._target(*self._args, **self._kwargs)
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/site-packages/uvicorn/subprocess.py", line 61, in subprocess_started
target(sockets=sockets)
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/site-packages/uvicorn/main.py", line 382, in run
loop.run_until_complete(self.serve(sockets=sockets))
File "uvloop/loop.pyx", line 1456, in uvloop.loop.Loop.run_until_complete
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/site-packages/uvicorn/main.py", line 389, in serve
config.load()
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/site-packages/uvicorn/config.py", line 288, in load
self.loaded_app = import_from_string(self.app)
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/site-packages/uvicorn/importer.py", line 23, in import_from_string
raise exc from None
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/site-packages/uvicorn/importer.py", line 20, in import_from_string
module = importlib.import_module(module_str)
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/importlib/__init__.py", line 127, in import_module
return _bootstrap._gcd_import(name[level:], package, level)
File "<frozen importlib._bootstrap>", line 1014, in _gcd_import
File "<frozen importlib._bootstrap>", line 991, in _find_and_load
File "<frozen importlib._bootstrap>", line 975, in _find_and_load_unlocked
File "<frozen importlib._bootstrap>", line 671, in _load_unlocked
File "<frozen importlib._bootstrap_external>", line 783, in exec_module
File "<frozen importlib._bootstrap>", line 219, in _call_with_frames_removed
File "./app/main.py", line 21, in <module>
from ariadne.asgi import GraphQL
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/site-packages/ariadne/__init__.py", line 1, in <module>
from .enums import EnumType
File "/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/site-packages/ariadne/enums.py", line 5, in <module>
from graphql.type import GraphQLEnumType, GraphQLNamedType, GraphQLSchema
ImportError: cannot import name 'GraphQLNamedType' from 'graphql.type' (/Library/Frameworks/Python.framework/Versions/3.8/lib/python3.8/site-packages/graphql/type/__init__.py)
```
| closed | 2020-04-18T07:39:17Z | 2020-04-18T08:06:39Z | https://github.com/mirumee/ariadne/issues/359 | [] | wkup | 1 |
coqui-ai/TTS | python | 2,905 | [Bug] Documentation for Simple Usage Not Working | ### Describe the bug
Hi, I was attempting to setup and run TTS locally on my Macbook this morning, and I ran into what seems like a documentation issue. The README gives the following example for a single speaker ([link](https://github.com/coqui-ai/TTS#running-a-single-speaker-model)):
```python
# Init TTS with the target model name
tts = TTS(model_name="tts_models/de/thorsten/tacotron2-DDC", progress_bar=False).to(device)
...
```
When I run this locally with everything installed, and `device` set to `cpu`, I get the following error:
```
AttributeError: 'TTS' object has no attribute 'to'
```
Note the previous setup I've already run in the same file / session:
```python
import torch
from TTS.api import TTS
# Get device
device = "cuda" if torch.cuda.is_available() else "cpu"
```
Is this an issue with the documentation? Or is this something to do with my setup or an error in the library? This seems like SUPER simple usage.
### To Reproduce
1. Install using `pip install TTS`, MacOS Ventura 13.4
2. Install `espeak` with `brew install espeak`
3. Run `python`
4. Run the following code:
```python
from TTS.api import TTS
device = 'cpu'
tts = TTS(model_name="tts_models/de/thorsten/tacotron2-DDC", progress_bar=False).to(device)
```
5. Observe the error:
```
AttributeError: 'TTS' object has no attribute 'to'
```
### Expected behavior
I'd expect this to load the model.
### Logs
_No response_
### Environment
```shell
{
"CUDA": {
"GPU": [],
"available": false,
"version": null
},
"Packages": {
"PyTorch_debug": false,
"PyTorch_version": "2.0.1",
"TTS": "0.14.3",
"numpy": "1.21.6"
},
"System": {
"OS": "Darwin",
"architecture": [
"64bit",
""
],
"processor": "i386",
"python": "3.8.8",
"version": "Darwin Kernel Version 22.5.0: Mon Apr 24 20:51:50 PDT 2023; root:xnu-8796.121.2~5/RELEASE_X86_64"
}
}
```
### Additional context
_No response_ | closed | 2023-08-30T13:55:47Z | 2023-09-04T10:55:06Z | https://github.com/coqui-ai/TTS/issues/2905 | [
"bug"
] | jeremyblalock | 2 |
Evil0ctal/Douyin_TikTok_Download_API | web-scraping | 102 | 视频ID获取失败 | 
__最初由 @kk20180123 在 https://github.com/Evil0ctal/Douyin_TikTok_Download_API/issues/101#issuecomment-1312716925 发布__ | closed | 2022-11-13T20:00:09Z | 2022-11-14T09:53:46Z | https://github.com/Evil0ctal/Douyin_TikTok_Download_API/issues/102 | [
"BUG",
"Fixed"
] | Evil0ctal | 6 |
microsoft/qlib | machine-learning | 1,058 | How can I do incremental training? | ## ❓ Questions and Help
I found a solution how to save and load models.
But I can't find a solution how to do incremental training with it.
Is there any example or documents to refer?
I think online_srv example is close to the solution.
But, there is only first_train in the example | closed | 2022-04-15T15:45:24Z | 2022-05-16T06:11:35Z | https://github.com/microsoft/qlib/issues/1058 | [
"question"
] | heury | 2 |
plotly/dash | jupyter | 3,236 | Dash 3 not properly rendering initially hidden plotly figures | **Describe the bug**
If more than 1 plots are created into a `Div` (which is initially hidden) and subsequently updated and shown, then the width of some of the plots is not correctly set. If using `responsive=True`, resizing the browser window fixes the behaviour. This appeared in Dash 3 and it was not present in Dash 2.
Consider the following MWE
<details>
```python
import dash
from dash import dcc, html, Input, Output
import plotly.graph_objects as go
app = dash.Dash(__name__)
app.layout = html.Div(
[
html.Button("Generate Figures", id="generate-btn", n_clicks=0),
html.Div(
id="graph-container",
children=[
dcc.Graph(
id="prec-climate-daily",
style={"height": "45vh"},
config={"responsive": True}
),
dcc.Graph(
id="temp-climate-daily",
style={"height": "45vh"},
config={"responsive": True}
),
],
style={"display": "none"}
),
]
)
@app.callback(
[
Output("prec-climate-daily", "figure"),
Output("temp-climate-daily", "figure"),
Output("graph-container", "style"),
],
[Input("generate-btn", "n_clicks")],
prevent_initial_call=True,
)
def update_figures(n_clicks):
fig_acc = go.Figure(data=[go.Scatter(x=[0, 1, 2], y=[0, 1, 0], mode="lines")])
fig_daily = go.Figure(data=[go.Scatter(x=[0, 1, 2], y=[1, 0, 1], mode="lines")])
return fig_acc, fig_daily, {"display": "block"}
if __name__ == "__main__":
app.run(debug=True)
```
</details>
When running you'll notice that one of the figures is created with default width and height values instead of filling the whole container.

There are different workarounds (which however are not compatible with different logics).
1) create `Graph` components directly in the callback (but this means they're not available in the initial app layout and thus not exposed to other callbacks)
<details>
```python
import dash
from dash import dcc, html, Input, Output
import plotly.graph_objects as go
app = dash.Dash(__name__)
app.layout = html.Div(
[
html.Button("Generate Figures", id="generate-btn", n_clicks=0),
html.Div(
id="graph-container",
children=[], # Initially empty
style={"display": "none"} # Initially hidden
),
]
)
@app.callback(
[
Output("graph-container", "children"),
Output("graph-container", "style"),
],
[Input("generate-btn", "n_clicks")],
prevent_initial_call=True,
)
def update_figures(n_clicks):
fig_acc = go.Figure(data=[go.Scatter(x=[0, 1, 2], y=[0, 1, 0], mode="lines")])
fig_daily = go.Figure(data=[go.Scatter(x=[0, 1, 2], y=[1, 0, 1], mode="lines")])
children = [
dcc.Graph(
id="prec-climate-daily",
style={"height": "45vh"},
config={"responsive": True},
figure=fig_acc
),
dcc.Graph(
id="temp-climate-daily",
style={"height": "45vh"},
config={"responsive": True},
figure=fig_daily
),
]
return children, {"display": "block"}
if __name__ == "__main__":
app.run(debug=True)
```
</details>

2) Removing the `hidden` property, thus showing empty plots on app startup
3) Reverting back to Dash 2.* where this issue is not present
4) Only having 1 plot into the app (I tried to make 1 `Div` per plot with different callbacks but this still trigger the issue...)
I don't have any error nor warning in the console so I'm kind of confused on what is triggering this issue in Dash 3 | open | 2025-03-24T12:28:36Z | 2025-03-24T15:01:26Z | https://github.com/plotly/dash/issues/3236 | [
"regression",
"bug",
"P1"
] | guidocioni | 0 |
kochlisGit/ProphitBet-Soccer-Bets-Predictor | seaborn | 83 | NSButton: 0x7fe573a48080 | Hello, I have successfully completed the installation, but I'm getting the following error and encountering the issue shown in the screenshot. How can I fix this?
024-04-15 23:28:53.160 python[10142:131011] Warning: Expected min height of view: (<NSButton: 0x7fe573a48080>) to be less than or equal to 30 but got a height of 32.000000. This error will be logged once per view in violation.
![Uploading Ekran Resmi 2024-04-15 23.41.34.png…]()
| open | 2024-04-15T20:42:06Z | 2024-04-16T07:01:08Z | https://github.com/kochlisGit/ProphitBet-Soccer-Bets-Predictor/issues/83 | [] | omerkymcu | 3 |
labmlai/annotated_deep_learning_paper_implementations | deep-learning | 283 | How to run model on multiple GPUs? | Hi, I am trying diffusion model.
SOLVED | closed | 2025-02-14T05:17:25Z | 2025-02-21T08:08:45Z | https://github.com/labmlai/annotated_deep_learning_paper_implementations/issues/283 | [] | yuffon | 0 |
axnsan12/drf-yasg | rest-api | 314 | How to wrapping with envelope? | I want to wrapping response value with `data`
In my class view code like this.
```
class UserList(BaseAPIView):
@swagger_auto_schema(responses={200: UserSerializer(many=True)})
def get(self, request: Request) -> Union[Response, NoReturn]:
```
And response is here -> `{'user_id': 'test', 'user_pw':'123'}`
But I want to wrap it like this -> `{'data': {'user_id': 'test', 'user_pw':'123'}}`
Is there any solution here? | closed | 2019-02-15T08:41:54Z | 2021-03-25T23:31:53Z | https://github.com/axnsan12/drf-yasg/issues/314 | [] | teamhide | 5 |
keras-rl/keras-rl | tensorflow | 287 | DDPG with MultiInputProcessor not working | Hi,
I have used MultiInputProcessor with DQN and it works fine. I am trying to use that feature to train an agent using DDPG.
I have 3 inputs from my environment: Image, two 1D vectors of size (1,3) (called vec_1 and vec_2)
My Actor-network is defined as:
```
actor = Model ( inputs = [image, vec_1, vec_2] ,
outputs = actions
)
```
My Critic-network is defined as:
```
critic = Model (inputs =[actions, image,vec_1, vec_2],
outputs = action_1
)
```
"action_1" is of the same dimension as "action".
`processor = MultiInputProcessor(nb_inputs=3)`
`agent = DDPGAgent(processor=processor,....,target_model_update=1e-3)`
When I run the code, I am getting an AssertionError from processor.py. Basically the len(observation) is not same as nb_inputs.
When I check the observation variable that is passed to the processor.py, I see that my "vec_1" and "vec_2" variables are not passed.
Since Critic network takes action as an input along with observation, the number of input to the critic model is one more than Actor-network. So how does MultiInputProcessor work with DDPG? Are there any examples of using DDPG with MultiInputProcessor?
Thanks
| open | 2019-01-17T00:10:19Z | 2019-01-18T17:15:57Z | https://github.com/keras-rl/keras-rl/issues/287 | [] | srivatsankrishnan | 6 |
ClimbsRocks/auto_ml | scikit-learn | 148 | ImportError: cannot import name Predictor | When importing using the example from the readme get the error in the title :-/ | closed | 2016-12-13T01:00:55Z | 2016-12-14T23:47:05Z | https://github.com/ClimbsRocks/auto_ml/issues/148 | [] | jeregrine | 13 |
autogluon/autogluon | computer-vision | 3,846 | Documentation for test set size minus train set size > prediction length | ### Describe the issue linked to the documentation
Perhaps this can be included in the documentation because it is unclear how to have settings for timeseries forecasting where I want to predict 1 step ahead.
The docs state to make the train_set and then the test_set = train_set + prediction_length.
If the prediction_length ==1 then the output plots for the test set has just a single point being predicted.
So here's an example.
Suppose I have 50 time series, all time-varying unknown. Suppose I have 100 rows of data total per time series.
Make train set == 90 rows
Test set would be all 100 rows.
prediction_length == 1
context_length (if applicable) is 10 rows
Walkforward validation is desired.
To my understanding from the docs, this would be something like:
```
predictor = TimeSeriesPredictor(
prediction_length=1,
target="target",
eval_metric="MSE",
)
predictor.fit(
train_data,
presets="best_quality",
random_seed=42,
num_val_windows=80, # this is 90 rows in train set minus 10 for the rolling context window.
val_step_size=1, # walkforward the 10 rows in the context_window by 1 timestep
"TemporalFusionTransformer": [ {"context_length": 10}],
....<other models>
)
```
After fitting the model, doing:
```
predictions = predictor.predict(train_data, random_seed=42)
```
I see that there is only a 1 mean value + the quantiles. Same results when passing in `test_data` as well.
Is the only option to iteratively pass in the test data in a for loop?
rows (80,90) -> 91
rows (81,91) -> 92
rows (82, 92) -> 93
...
rows (89, 99) -> 100
### Suggest a potential alternative/fix
Keep same API from `.fit`. i.e. do:
```
predictions = predictor.predict(train_data, random_seed=42, test_step_size=1)
```
To do autoregressive forecasting.
I'd also have `context_length` apply to all models globally. | open | 2024-01-04T03:49:48Z | 2024-06-27T17:36:12Z | https://github.com/autogluon/autogluon/issues/3846 | [
"API & Doc",
"module: timeseries"
] | esnvidia | 0 |
widgetti/solara | flask | 989 | Feature Request: accept async functions in `on_kernel_start` | ## Feature Request
- [ ] The requested feature would break current behaviour
- [x] I would be interested in opening a PR for this feature
### What problem would this feature solve? Please describe.
For certain functionality that you might want to execute on kernel start, you need to call asynchronous functions. It would be useful to accept these for `solara.lab.on_kernel_start`
### Describe the solution you'd like
The cleanest would be simply allowing
```python
@solara.lab.on_kernel_start
async def init():
await database.connect()
async def cleanup():
await database.disconnect()
return cleanup
```
| open | 2025-01-28T14:18:35Z | 2025-01-28T14:38:43Z | https://github.com/widgetti/solara/issues/989 | [
"enhancement"
] | iisakkirotko | 0 |
gunthercox/ChatterBot | machine-learning | 2,216 | Maintainer Needed? | What's the state of maintenance for Chatterbot? It seems a number of important things are unresolved without comment, and there's been only a single doc fixing commit this year.
It can often be quite reasonable to find new maintainers for projects by reaching out to PR contributors and fork owners. There are also some looking-for-maintainer hubs out there that try to bring people in.
@gunthercox are you still on this project or do you need help? Is anyone available to step in? | closed | 2021-11-05T18:31:36Z | 2025-02-25T22:47:50Z | https://github.com/gunthercox/ChatterBot/issues/2216 | [] | xloem | 8 |
vitalik/django-ninja | django | 499 | serialization without a view? | Please describe what you are trying to achieve
I have some places in my code where I need to get at a JSON version of an object that aren't within the API. Is there some way to take advantage of the Schemas I've defined to convert?
Please include code examples (like models code, schemes code, view function) to help understand the issue
```
from ninja.hope_this_exists import serialization_helper
class Job(models.Model):
...
class JobOut(Schema):
...
job = Job.objects.get(id=1)
serialized = serialization_helper(JobOut, job)
```
| closed | 2022-07-06T18:22:25Z | 2023-04-02T11:39:51Z | https://github.com/vitalik/django-ninja/issues/499 | [] | cltrudeau | 7 |
httpie/http-prompt | api | 70 | Feature proposal: saving current request setup under named alias | This is great package, thanks for sharing your work!
As I've been using it couple of days I noticed that it's quite inconvenient, having to repeatedly change request settings every time I want to send different request.
The option of saving current state of request would solve the problem.
Given this output of `httpie post`:
``` bash
http://127.0.0.1:3000/api/oauth/token> http --form --style native POST http://127.0.0.1:3000/api/oauth/token user_id==value client_id=value client_secret=value grant_type=password password=test username=testuser1 Content-Type:application/x-www-form-urlencoded
```
We could use eg.: `http://127.0.0.1:3000/api/oauth/token> alias getToken`
which would save the request under `getToken` keyword. Later, we could call getToken request and optionally overwrite any req options like so:
``` bash
http://127.0.0.1:3000/api/oauth/token> getToken username=testuser2 password=test2
```
| closed | 2016-07-25T10:08:26Z | 2016-09-18T06:44:03Z | https://github.com/httpie/http-prompt/issues/70 | [
"enhancement"
] | fogine | 17 |
WZMIAOMIAO/deep-learning-for-image-processing | pytorch | 41 | About pytorch version | ssd_model.py
line4:from torch.jit.annotations import Optional, List, Dict, Tuple, Module
I just deleted “Optional” and run it directly under pytorch1.0. It seems like module Optional is useless......
| closed | 2020-07-31T08:38:35Z | 2020-08-21T03:08:52Z | https://github.com/WZMIAOMIAO/deep-learning-for-image-processing/issues/41 | [] | Venquieu | 1 |
babysor/MockingBird | deep-learning | 598 | protoc 跑不起来训练,_pb2.py in <module>到底哪个版本可以啊? | protoc 哪个版本能跑起来synthesizer_traina啊?
3.19.4
21.0.rc
21.1
都不行
还是一大堆_pb2.py file,in<module>
traceback
File "D:\MockingBird-main\synthesizer_train.py", line 2, in <module>
from synthesizer.train import train
File "D:\MockingBird-main\synthesizer\train.py", line 5, in <module>
from torch.utils.tensorboard import SummaryWriter
File "C:\Users\H410MH\AppData\Local\Programs\Python\Python39\lib\site-packages\torch\utils\tensorboard\__init__.py", line 10, in <module>
from .writer import FileWriter, SummaryWriter # noqa: F401
File "C:\Users\H410MH\AppData\Local\Programs\Python\Python39\lib\site-packages\torch\utils\tensorboard\writer.py", line 9, in <module>
from tensorboard.compat.proto.event_pb2 import SessionLog
File "C:\Users\H410MH\AppData\Local\Programs\Python\Python39\lib\site-packages\tensorboard\compat\proto\event_pb2.py", line 17, in <module>
from tensorboard.compat.proto import summary_pb2 as tensorboard_dot_compat_dot_proto_dot_summary__pb2
File "C:\Users\H410MH\AppData\Local\Programs\Python\Python39\lib\site-packages\tensorboard\compat\proto\summary_pb2.py", line 17, in <module>
from tensorboard.compat.proto import tensor_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__pb2
File "C:\Users\H410MH\AppData\Local\Programs\Python\Python39\lib\site-packages\tensorboard\compat\proto\tensor_pb2.py", line 16, in <module>
from tensorboard.compat.proto import resource_handle_pb2 as tensorboard_dot_compat_dot_proto_dot_resource__handle__pb2
File "C:\Users\H410MH\AppData\Local\Programs\Python\Python39\lib\site-packages\tensorboard\compat\proto\resource_handle_pb2.py", line 16, in <module>
from tensorboard.compat.proto import tensor_shape_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__shape__pb2
File "C:\Users\H410MH\AppData\Local\Programs\Python\Python39\lib\site-packages\tensorboard\compat\proto\tensor_shape_pb2.py", line 36, in <module>
_descriptor.FieldDescriptor(
File "C:\Users\H410MH\AppData\Local\Programs\Python\Python39\lib\site-packages\google\protobuf\descriptor.py", line 560, in __new__
_message.Message._CheckCalledFromGeneratedFile()
类型
TypeError: Descriptors cannot not be created directly.
If this call came from a _pb2.py file, your generated code is out of date and must be regenerated with protoc >= 3.19.0.
If you cannot immediately regenerate your protos, some other possible workarounds are:
1. Downgrade the protobuf package to 3.20.x or lower.
| closed | 2022-06-02T02:27:13Z | 2022-06-11T07:22:47Z | https://github.com/babysor/MockingBird/issues/598 | [] | kidwanttolearn | 2 |
graphql-python/graphene | graphql | 973 | Graphene Django and Classes instead of models | I am using graphene on a django project. It's nice to have graphene-django since you can add the Meta class and set the model from django models. But what I want to do is return my own custom classes.
What I have is a Profile class, I am using project without db, so this profile class has properties like name, email, age, etc
What I need is to return this Profile class instead of a django model class or the graphene Object Types, how is that posible
I woul like to do something like
```
import Profile
class ProfileType(Profile)
pass
```
in that case profile has all the attributes but like a python object and not as a django model.
or maybe add this class as a Meta property | closed | 2019-05-23T02:02:25Z | 2019-05-23T03:12:03Z | https://github.com/graphql-python/graphene/issues/973 | [] | pmcarlos | 0 |
RomelTorres/alpha_vantage | pandas | 237 | Alpha Vantage API return empty results [bug?] | Dear,
The following query does not work anymore (returns an empty dictionary).
Do you introduce some restrictions for Italian stocks?
https://www.alphavantage.co/query?function=TIME_SERIES_DAILY&outputsize=full&symbol=ENI.MI&apikey=
Best
Lorenzo | closed | 2020-06-26T06:33:59Z | 2020-06-26T16:14:20Z | https://github.com/RomelTorres/alpha_vantage/issues/237 | [] | lorenzo-gabrielli | 6 |
horovod/horovod | tensorflow | 3,251 | How to load a pre-trained model using horovod | **Environment:**
1. Framework: PyTorch 1.4
2. python version: horovod with ppytorch
3. Horovod version: latest version
4. MPI version: 4.0.3
5. CUDA version: cuda10.1
6. NCCL version: 2.5.6
7. Python version: python3.6
**Question**
I can successfully train my model using horovod in a multi-machine-multi-card fashion, and save the checkpoint models only on worker 0 to prevent other workers from corrupting them. My question is how to load such pre-trained models using horovod in a multi-machine-multi-card fashion, Do I need to load the pre-trained models only on worker 0 ?
| closed | 2021-11-01T06:52:27Z | 2021-11-02T12:34:25Z | https://github.com/horovod/horovod/issues/3251 | [
"bug"
] | ForawardStar | 0 |
vaexio/vaex | data-science | 2,230 | [BUG-REPORT] median_approx and percentile_approx return nan instead of actual value | **Description**
When using `median_approx` or `percentile_approx` to calculate the median or a percentile of a (Vaex) data frame or one of its columns, they return `nan` instead of the actual value of the statistic. I'm not sure if this might be related to #1304.
See for instance the behavior of the median approximation function illustrated below with the Vaex example dataset.
```
import vaex
vdf = vaex.example()
v_med_x = vdf.median_approx('x')
v_med_xy = vdf.median_approx(['x', 'y'])
v_med_df = vdf.median_approx(vdf.get_column_names())
pdf = vdf.to_pandas_df()
p_med_x = pdf['x'].median()
p_med_xy = pdf[['x', 'y']].median()
p_med_df = pdf.median()
print(f'Vaex results:\n\tMedian of \'x\':\t\t{v_med_x}\n\tMedian of \'x\' and \'y\':\t{v_med_xy}\n\tMedian of all df:\t{v_med_df}\n\nPandas results:\n\tMedian of \'x\':\t\t{p_med_x}\n\tMedian of \'x\' and \'y\':\t{list(p_med_xy)}\n\tMedian of all df:\t{list(p_med_df)}')
```
This little program returns the following output on the screen, where one can see that **the Vaex medians are all nan's and differ from those reported by Pandas (note that they exist and are finite)**. May there be an issue with the interpolation algorithm used by Vaex to estimate these metrics?
_Is there a way to get the median/percentile with Vaex at the moment?_
```
Vaex results:
Median of 'x': nan
Median of 'x' and 'y': [nan nan]
Median of all df: [nan nan nan nan nan nan nan nan nan nan nan]
Pandas results:
Median of 'x': -0.05056469142436981
Median of 'x' and 'y': [-0.05056469142436981, -0.032549407333135605]
Median of all df: [16.0, -0.05056469142436981, -0.032549407333135605, -0.007299995049834251, 0.3338821530342102, 2.5303449630737305, 0.5708029270172119, -105428.9765625, 752.062744140625, -164.71417236328125, -1.6478899717330933]
```
**Software information**
- Vaex version: {'vaex-core': '4.14.0', 'vaex-viz': '0.5.4', 'vaex-hdf5': '0.13.0', 'vaex-ml': '0.18.0', 'vaex-arrow': '0.4.2'}
- Vaex was installed via: conda-forge (conda install -c conda-forge vaex-core vaex-viz vaex-ml vaex-hdf5 vaex-arrow)
- OS: Windows 10 Pro version 21H2
- Python version: Python 3.9.13 | packaged by conda-forge
- NumPy version: 1.23.3
- Pandas version: 1.5.0
| open | 2022-10-14T16:24:15Z | 2022-10-19T03:40:49Z | https://github.com/vaexio/vaex/issues/2230 | [] | abianco88 | 17 |
AUTOMATIC1111/stable-diffusion-webui | pytorch | 15,483 | [Bug]: Loading an fp16 SDXL model fails on MPS when model loading RAM optimization is enabled | ### Checklist
- [ ] The issue exists after disabling all extensions
- [ ] The issue exists on a clean installation of webui
- [ ] The issue is caused by an extension, but I believe it is caused by a bug in the webui
- [X] The issue exists in the current version of the webui
- [ ] The issue has not been reported before recently
- [ ] The issue has been reported before but has not been fixed yet
### What happened?
Loading an fp16 model on MPS fails when `--disable-model-loading-ram-optimization` is not enabled. I suspect this is because fp32 is natively supported by MPS, but something is causing an attempted cast to fp64 from fp16, which is not supported.
### Steps to reproduce the problem
1. Try to load an fp16 model on MPS (Apple Silicon).
### What should have happened?
The model loading RAM optimization hack shouldn't interfere with this.
### What browsers do you use to access the UI ?
_No response_
### Sysinfo
```
{
"Platform": "macOS-14.2.1-arm64-arm-64bit",
"Python": "3.10.13",
"Version": "v1.9.0-RC-12-gac8ffb34",
"Commit": "ac8ffb34e3d1196319c8dcfcf0f5d3eded713176",
"Commandline": [
"webui.py",
"--skip-torch-cuda-test",
"--upcast-sampling",
"--no-half-vae",
"--use-cpu",
"interrogate",
"--skip-load-model-at-start"
],
}
```
### Console logs
```Shell
Traceback (most recent call last):
File "stable-diffusion-webui/modules/options.py", line 165, in set
option.onchange()
File "stable-diffusion-webui/modules/call_queue.py", line 13, in f
res = func(*args, **kwargs)
File "stable-diffusion-webui/modules/initialize_util.py", line 181, in <lambda>
shared.opts.onchange("sd_model_checkpoint", wrap_queued_call(lambda: sd_models.reload_model_weights()), call=False)
File "stable-diffusion-webui/modules/sd_models.py", line 860, in reload_model_weights
sd_model = reuse_model_from_already_loaded(sd_model, checkpoint_info, timer)
File "stable-diffusion-webui/modules/sd_models.py", line 826, in reuse_model_from_already_loaded
load_model(checkpoint_info)
File "stable-diffusion-webui/modules/sd_models.py", line 748, in load_model
load_model_weights(sd_model, checkpoint_info, state_dict, timer)
File "stable-diffusion-webui/modules/sd_models.py", line 393, in load_model_weights
model.load_state_dict(state_dict, strict=False)
File "stable-diffusion-webui/modules/sd_disable_initialization.py", line 223, in <lambda>
module_load_state_dict = self.replace(torch.nn.Module, 'load_state_dict', lambda *args, **kwargs: load_state_dict(module_load_state_dict, *args, **kwargs))
File "stable-diffusion-webui/modules/sd_disable_initialization.py", line 221, in load_state_dict
original(module, state_dict, strict=strict)
File "stable-diffusion-webui/venv/lib/python3.10/site-packages/torch/nn/modules/module.py", line 2139, in load_state_dict
load(self, state_dict)
File "stable-diffusion-webui/venv/lib/python3.10/site-packages/torch/nn/modules/module.py", line 2127, in load
load(child, child_state_dict, child_prefix)
File "stable-diffusion-webui/venv/lib/python3.10/site-packages/torch/nn/modules/module.py", line 2127, in load
load(child, child_state_dict, child_prefix)
File "stable-diffusion-webui/venv/lib/python3.10/site-packages/torch/nn/modules/module.py", line 2127, in load
load(child, child_state_dict, child_prefix)
[Previous line repeated 3 more times]
File "stable-diffusion-webui/venv/lib/python3.10/site-packages/torch/nn/modules/module.py", line 2121, in load
module._load_from_state_dict(
File "stable-diffusion-webui/modules/sd_disable_initialization.py", line 226, in <lambda>
conv2d_load_from_state_dict = self.replace(torch.nn.Conv2d, '_load_from_state_dict', lambda *args, **kwargs: load_from_state_dict(conv2d_load_from_state_dict, *args, **kwargs))
File "stable-diffusion-webui/modules/sd_disable_initialization.py", line 191, in load_from_state_dict
module._parameters[name] = torch.nn.parameter.Parameter(torch.zeros_like(param, device=device, dtype=dtype), requires_grad=param.requires_grad)
File "stable-diffusion-webui/venv/lib/python3.10/site-packages/torch/_meta_registrations.py", line 4806, in zeros_like
res = aten.empty_like.default(
File "stable-diffusion-webui/venv/lib/python3.10/site-packages/torch/_ops.py", line 513, in __call__
return self._op(*args, **(kwargs or {}))
File "stable-diffusion-webui/venv/lib/python3.10/site-packages/torch/_prims_common/wrappers.py", line 250, in _fn
result = fn(*args, **kwargs)
File "stable-diffusion-webui/venv/lib/python3.10/site-packages/torch/_refs/__init__.py", line 4755, in empty_like
return torch.empty_permuted(
TypeError: Cannot convert a MPS Tensor to float64 dtype as the MPS framework doesn't support float64. Please use float32 instead
```
### Additional information
_No response_ | open | 2024-04-11T05:57:42Z | 2024-04-11T05:58:36Z | https://github.com/AUTOMATIC1111/stable-diffusion-webui/issues/15483 | [
"bug-report"
] | akx | 0 |
coleifer/sqlite-web | flask | 37 | Some safety features for remote DB access | I just needed a sqlite db browser for testing of my program on remote Linux server.
Sqlite-web is the only I found. It is simple and useful. Thank you!
What I'm missing it is **simple password protection** to access the database remotely.
For example specifying password as parameter in the start command line would be fine. Now each time I have to stop the server and it is unsafe if I forget it. There might be ways to handle it but I'm not good at web server administration.
Another useful option would be **server auto stop** after some time of inactivity | closed | 2018-02-14T23:57:43Z | 2018-02-23T17:42:09Z | https://github.com/coleifer/sqlite-web/issues/37 | [] | lunfardo314 | 6 |
JoeanAmier/TikTokDownloader | api | 316 | 无法下载内容 | 版本
TikTokDownloader_V5.4_WIN

| open | 2024-10-18T16:18:00Z | 2024-10-29T03:17:08Z | https://github.com/JoeanAmier/TikTokDownloader/issues/316 | [] | ZX828 | 2 |
nerfstudio-project/nerfstudio | computer-vision | 3,332 | COLMAP crashed causing the machine to restart | **Describe the bug**
When using nerfstudio to process image data, the computer will restart directly during the bundle adjustment stage. It doesn't happen when the number of pictures is small, but it does happen with a large number of pictures (around 400 pics, 4000*6000).
**Hardware**
GPU: 4090, 24G
Mem: 126G
Swap: 2G
**To Reproduce**
Steps to reproduce the behavior:
1. Run ns-process-data
2. In BA stage, the machine reboots.
**Expected behavior**
After completing BA, we get the sparse reconstruction result.
**Additional context**
- In the same environment, if the number of images is small, such as 100, nerfstudio can call colmap to process it normally, and subsequent training is also good.
- Even with large amounts of data, openSfM can still be used for sparse reconstruction. I wonder if there are plans to support the openSfM format in the future. Or if the processed colmap format data can be used directly without nerfstudio calling it again. | open | 2024-07-25T09:18:33Z | 2024-07-29T13:11:22Z | https://github.com/nerfstudio-project/nerfstudio/issues/3332 | [] | YuhsiHu | 2 |
biolab/orange3 | pandas | 6,881 | Switch the language of orange3 interface to Chinese. | <!--
Thanks for taking the time to submit a feature request!
For the best chance at our team considering your request, please answer the following questions to the best of your ability.
-->
**What's your use case?**
<!-- In other words, what's your pain point? -->
<!-- Is your request related to a problem, or perhaps a frustration? -->
<!-- Tell us the story that led you to write this request. -->
Because there is no Chinese version of orange3 software, a group of people in China have translated it and sold it at a high price. As a student and a learning enthusiast, its price is unbearable. I hope I can add an option to switch the software language to Chinese in the settings, which can be downloaded and used as a plug-in
**What's your proposed solution?**
<!-- Be specific, clear, and concise. -->
1. Chinese language packs are downloaded as plug-ins.
2. Add options about Chinese language packs in the settings.
**Are there any alternative solutions?**
Use the default language of the computer system.
| closed | 2024-08-23T03:20:41Z | 2024-11-17T01:47:50Z | https://github.com/biolab/orange3/issues/6881 | [] | TonyEinstein | 5 |
widgetti/solara | jupyter | 619 | "Page not found by Solara router" with the boilerplate package from `solara create portal ` | I am considering using solara for a prototype/showcase app. Starting from the boilerplate from `solara create portal` seems not to work out of the box with the current version 1.32
I'll see if I can figure out a PR, but no promise - not my field.
## Repro info
```sh
python --version # 3.9.13; debian linux
cd $HOME/.venvs/
python -m venv solara-env
source ./solara-env/bin/activate
pip install solara # installs 1.32.0
~/.venvs/solara-env/bin/python -m pip install --upgrade pip
pip install matplotlib pandas vaex plotly
mkdir -p $HOME/$BLAH/showcase/
cd $HOME/$BLAH/showcase/
solara create --help
solara create portal stream-temperature
cd stream-temperature/
pip install --no-deps -e .
cd ..
solara run stream_temperature
```
| closed | 2024-04-29T07:59:59Z | 2024-04-29T08:06:47Z | https://github.com/widgetti/solara/issues/619 | [] | jmp75 | 1 |
gunthercox/ChatterBot | machine-learning | 1,827 | Converting chatterbot used py to exe | After converting a simple chatbot program (using chatterbot module) to exe i encountered thousands of module not found runtime error when i run the exe file.... I simply keep adding hidden imports while compiling to exe using pyinstaller but i am tired now as there are tons of hidden modules require i guess...... So is there any easy way to solve the error??? | closed | 2019-10-06T10:39:16Z | 2020-06-20T17:32:17Z | https://github.com/gunthercox/ChatterBot/issues/1827 | [
"answered"
] | ProPython007 | 2 |
marcomusy/vedo | numpy | 1,085 | Bug: Thumbnail slices part of mesh | ```
import vedo
import numpy as np
mesh = vedo.Mesh("snare.stl")
image_size=224
num_views = 16
sqrt_num_views = 4
images = []
for i in range(sqrt_num_views):
for j in range(sqrt_num_views):
image = mesh.thumbnail(
size=(image_size, image_size),
azimuth=360 * i / sqrt_num_views - 180,
elevation=180 * j / sqrt_num_views - 90,
zoom=1.25,
axes=1,
)
images.append(image)
images = np.array(images)
ncols = np.floor(np.sqrt(num_views))
nrows = np.ceil(np.sqrt(num_views))
# Create a figure and axes
fig, axs = plt.subplots(4, 4, figsize=(10, 10))
# Flatten axes for easier indexing
axs = axs.flatten()
# Loop through each image and plot
for i in range(16):
axs[i].imshow(images[i])
# axs[i].axis("off") # Turn off axis
# Adjust layout
plt.tight_layout()
# Show plot
plt.show()
```
[snare.zip](https://github.com/marcomusy/vedo/files/14878751/snare.zip)
| closed | 2024-04-05T01:26:17Z | 2024-04-05T17:59:57Z | https://github.com/marcomusy/vedo/issues/1085 | [
"bug",
"fixed"
] | JeffreyWardman | 1 |
ets-labs/python-dependency-injector | asyncio | 262 | Does the `from_ini` method of the Configuration provider always return a str even when the value is an int? | I just started working with the newest version of the dependency injector and I'm curious about the Configuration provider and its return types. Using `from_dict` seems to return the correct values and correct types of those values, e.g.:
```python
config.from_dict(
{
'section1': {
'option1': 1, # returns int as value type
'option2': 2, # returns int as value type
},
},
)
```
Using the `from_ini` method has a different result from the same structure, e.g.:
```ini
[section1]
option1 = 1 # returns str as value type
option2 = 2 # returns str as value type
```
Is there something I'm overlooking in the documentation or is there something that can be done to make `from_ini` return its values in the same way as the `from_dict` method?
As an intermediate solution, I'm casting all the values from the dict to their appropriate (int or float) value in a new dict because the original dict is unassignable.
For each section I currently have the following snippet:
```python
sec1 = {k: float(v) if '.' in v else int(v)
for (k, v) in Core.config.section1().items()}
```
Also, how does the Configuration provider handle changes in the source file? Let's say I already used `from_ini` once and then updated the file with new configurations, will the `from_ini` method load the file again or maybe does the `provider.Configuration.section1` auto-magically have updated values? | closed | 2020-07-08T08:59:22Z | 2020-08-24T17:42:33Z | https://github.com/ets-labs/python-dependency-injector/issues/262 | [
"question"
] | zborowa | 2 |
itamarst/eliot | numpy | 185 | Test against newer version of Twisted | Probably 15.2.
| closed | 2015-07-24T20:26:50Z | 2018-09-22T20:59:18Z | https://github.com/itamarst/eliot/issues/185 | [] | itamarst | 0 |
Netflix/metaflow | data-science | 1,712 | add __repr__ methods to Parameter | I think it would be useful to add __str__ and __repr__ methods to the Parameter class (https://github.com/Netflix/metaflow/blob/master/metaflow/parameters.py#L212) to improve understanding and debugging | closed | 2024-02-02T17:02:10Z | 2024-02-07T10:52:10Z | https://github.com/Netflix/metaflow/issues/1712 | [] | raybellwaves | 0 |
hack4impact/flask-base | sqlalchemy | 53 | Add auto-formatting to flask-base | From working on Go and working at Google (where most code is auto-formatted) I've noticed that there are immense benefits to auto-formatting code.
1. Code style and conformation to standards, like pep8, are ensured.
2. Writing code is faster.
3. Less cognitive load, it's one less thing to worry about.
4. No arguments about style.
5. If all of our python code is formatted the same, getting up to speed with code you didn't write will take less effort.
6. Diffs show actual code, not 'diff noise' where someone adds a space or changes formatting.
| closed | 2016-08-21T05:02:41Z | 2016-08-28T22:44:07Z | https://github.com/hack4impact/flask-base/issues/53 | [] | sandlerben | 3 |
ansible/awx | django | 15,581 | awx.awx.role module failed to assign role to teams when using "lookup_organization" parameter | ### Please confirm the following
- [X] I agree to follow this project's [code of conduct](https://docs.ansible.com/ansible/latest/community/code_of_conduct.html).
- [X] I have checked the [current issues](https://github.com/ansible/awx/issues) for duplicates.
- [X] I understand that AWX is open source software provided for free and that I might not receive a timely response.
- [X] I am **NOT** reporting a (potential) security vulnerability. (These should be emailed to `security@ansible.com` instead.)
### Bug Summary
awx.awx.role module failed when calling with "lookup_organization" pointing to a different organization where the teams are located.
### AWX version
any
### Select the relevant components
- [ ] UI
- [ ] UI (tech preview)
- [ ] API
- [ ] Docs
- [X] Collection
- [ ] CLI
- [ ] Other
### Installation method
N/A
### Modifications
no
### Ansible version
_No response_
### Operating system
_No response_
### Web browser
_No response_
### Steps to reproduce
For example. We want to assign use for project ABC-project under organization ABC to team XYZ under organization XYZ.
tasks:
- name: Assign Project ABC-project use permission to Team XYZ across organizations
ansible.controller.role:
teams: XYZ
projects: ABC-project
role: use
lookup_organization: ABC
### Expected results
Should be able to assign role permission.
### Actual results
The task return failed message
fatal: [localhost]: FAILED! => {"changed": false, "msg": "There were 1 missing items, missing items: ['XYZ']"}
### Additional information
It seems to be a bug in role.py
https://github.com/ansible/awx/pull/15580 | closed | 2024-10-11T23:23:38Z | 2025-03-12T17:33:45Z | https://github.com/ansible/awx/issues/15581 | [
"type:bug",
"component:awx_collection",
"needs_triage"
] | ecchong | 1 |
davidsandberg/facenet | computer-vision | 626 | align_dataset_mtcnn.py too slow | I followed the instructions in https://github.com/davidsandberg/facenet/wiki/Classifier-training-of-inception-resnet-v1 and ran python src/align/align_dataset_mtcnn.py ~/datasets/casia/CASIA-maxpy-clean/ ~/datasets/casia/casia_maxpy_mtcnnpy_182 --image_size 182 --margin 44.
It's working but seems to be very slow. The gpu utilization is also low ( 3% of gpu0 and %0 of gpu1). I don't think I can wait that long. Could anyone give me some help? Thanks! | closed | 2018-01-23T08:26:23Z | 2018-01-23T09:55:22Z | https://github.com/davidsandberg/facenet/issues/626 | [] | eurus-yu | 1 |
stanfordnlp/stanza | nlp | 479 | Connecting to remote CoreNLP server | closed | 2020-10-06T04:53:40Z | 2020-10-10T17:28:20Z | https://github.com/stanfordnlp/stanza/issues/479 | [
"question"
] | ronkow | 8 | |
pytorch/vision | computer-vision | 8,297 | Allow empty class folders in ImageFolder | Follow up to https://github.com/pytorch/vision/issues/4925 and as summarized in https://github.com/pytorch/vision/issues/4925#issuecomment-1976785097, we need to allow empty class folders in ImageFolder i.e. we need a way for users to bypass this error:
https://github.com/pytorch/vision/blob/c8c3839cb9b0fc52885cafa431460f686c46f5ae/torchvision/datasets/folder.py#L98-L99
The most obvious way to enable that is by adding a new `allow_empty` parameter to `make_dataset()` and `ImageFolder()` / `DatasetFolder()`. (name TBD). | closed | 2024-03-05T10:30:55Z | 2024-03-13T13:29:44Z | https://github.com/pytorch/vision/issues/8297 | [
"enhancement",
"module: datasets"
] | NicolasHug | 0 |
tfranzel/drf-spectacular | rest-api | 473 | How can I disable schema for patch requests across all APIs | Currently I'm using this on all my viewsets:
```
@extend_schema_view(
partial_update=extend_schema(exclude=True),
)
class MyViewSet(...):
...
```
Any better suggestions? | closed | 2021-07-31T05:12:42Z | 2021-08-01T08:40:09Z | https://github.com/tfranzel/drf-spectacular/issues/473 | [] | tiholic | 2 |
pallets/quart | asyncio | 37 | http2_push example hangs when using nghttp | I was trying to verify the http2_push example by using nghttp. The example does seem to work fine from a browser (Chrome or Firefox).
Repro:
> $ QUART_APP=http2_push:app quart run
> Running on https://127.0.0.1:5000 (CTRL + C to quit)
> [2018-12-19 14:44:13,507] ASGI Framework Lifespan error, continuing without Lifespan support
>
> $ nghttp -nvy https://localhost:5000/push
>
| closed | 2018-12-19T22:49:49Z | 2022-07-06T00:24:01Z | https://github.com/pallets/quart/issues/37 | [] | jonozzz | 4 |
pallets/flask | flask | 4,425 | flask app timeout urgent issue | Dears,
Hope all is well.
I have an application, my app build based on the flask, my issue is as follows,
after sending 1000 to 3000 requests per minute to my flask app the flask going down (no response ) kindly keep in your mind that my issue is not a resource issue because my memory percentage is less than 20% and CPU 30 to40 %
so how can I fix this issue in flask
PS: I did the same testing in fastapi and there is no issue with them
| closed | 2022-01-20T12:29:16Z | 2022-01-20T12:30:35Z | https://github.com/pallets/flask/issues/4425 | [] | themeswordpress | 1 |
polakowo/vectorbt | data-visualization | 165 | Is there a way to change the start date in portfolio.stats() ? | Hi guys,
I have a doubt, is there a way to change the start date in portfolio.stats() ? I'm backtesting a portfolio optimization model using a rolling window to rebalance the portfolio monthly, but when I use portfolio.stats(), the days consider the initial period (about 5 years) where the portfolio is only cash. I want to calculate the stats starting the first date when I rebalance the portfolio. I try the solution:
`a = portfolio.value().iloc[252*5:].pct_change().dropna()` # I extract data from the vectorbt portfolio,
`a.vbt.returns(freq='D').stats(0)` # I calculate stats from portfolio returns
The Total Return obtained using this method is different from that obtained using portfolio.stats(), however they very are close.
Best
Dany | closed | 2021-06-12T07:05:41Z | 2021-06-13T05:25:28Z | https://github.com/polakowo/vectorbt/issues/165 | [] | dcajasn | 2 |
encode/apistar | api | 68 | API Mocking | Once we've got the schemas fully in place we'd like to be able to run a mock API purely based on the function annotations, so that users can start up a mock API even before they've implemented any functionality.
We'd want to use randomised output that fits the given schema constraints, as well as validating any input, and returning those values accordingly.
eg.
**schemas.py**:
```python
class KittenName(schema.String):
max_length = 100
class KittenColor(schema.Enum):
enum = [
'black',
'brown',
'white',
'grey',
'tabby'
]
class Kitten(schema.Object):
properties = {
'name': KittenName,
'color': KittenColor,
'cuteness': schema.Number(
minimum=0.0,
maximum=10.0,
multiple_of=0.1
)
}
```
**views.py:**
```python
def list_favorite_kittens(color: KittenColor=None) -> List[Kitten]:
"""
List your favorite kittens, optionally filtered by color.
"""
pass
def add_favorite_kitten(name: KittenName) -> Kitten:
"""
Add a kitten to your favorites list.
"""
pass
```
We should be able to do this sort of thing...
```bash
$ apistar mock
mock api running on 127.0.0.1:5000
$ curl http://127.0.0.1:5000/kittens/fav/
[
{
"name": "congue",
"color": "tabby",
"cuteness": 5.9
},
{
"name": "aenean",
"color": "white",
"cuteness": 9.3
},
{
"name": "etiam",
"color": "tabby",
"cuteness": 8.8
}
]
``` | open | 2017-04-20T13:50:35Z | 2017-08-17T11:38:46Z | https://github.com/encode/apistar/issues/68 | [
"Baseline feature"
] | tomchristie | 2 |
pyeve/eve | flask | 769 | GeoJson types limited to 2 fields | Quoting from the [GeoJSON specification](http://geojson.org/geojson-spec.html#geojson-objects):
> The GeoJSON object may have any number of members (name/value pairs)
Using one of the GeoJson types in Eve schemas I cannot add additional fields besides "types" and "coordinates".
For istance the following GeoJSON is valid according to http://geojsonlint.com/, but is rejected by Eve, having defined the field as type "point"
```
{
"type": "Point",
"coordinates": [
-105.01621,
39.57422
],
"customField": "blabla"
}
```
| closed | 2015-11-24T16:10:28Z | 2018-05-18T18:19:37Z | https://github.com/pyeve/eve/issues/769 | [
"stale"
] | mion00 | 1 |
slackapi/python-slack-sdk | asyncio | 903 | ActionsBlock elements are not parsed | ### Description
In ActionsBlock __init__, elements are simplie copied.
[Exact bug place](https://github.com/slackapi/python-slack-sdk/blob/4e4524230ed41ef7cd9d637c52f4e86b1ffedad9/slack/web/classes/blocks.py#L241)
For all other Blocks, similar elements are parsed into BlockElement
[How it parsed for SectionBlock](https://github.com/slackapi/python-slack-sdk/blob/4e4524230ed41ef7cd9d637c52f4e86b1ffedad9/slack/web/classes/blocks.py#L143)
[How it is parsed for ContextBlock](https://github.com/slackapi/python-slack-sdk/blob/4e4524230ed41ef7cd9d637c52f4e86b1ffedad9/slack/web/classes/blocks.py#L269)
To fix this we can deal with it the same way as in ContextBlock -> replace [this line of code](https://github.com/slackapi/python-slack-sdk/blob/4e4524230ed41ef7cd9d637c52f4e86b1ffedad9/slack/web/classes/blocks.py#L241) by
`self.elements = BlockElement.parse_all(elements)`
Also the same issue present in current sdk version(3.1.0)
[Link for bug place in 3.1.0](https://github.com/slackapi/python-slack-sdk/blob/5340ee337a2364e84c38d696c107f19c341dd6eb/slack_sdk/models/blocks/blocks.py#L246)
### Reproducible in:
#### The Slack SDK version
slackclient==2.9.3
#### Python runtime version
Python 3.7.0
#### OS info
Ubuntu 20.04.1 LTS
#### Steps to reproduce:
1.Copy example json from slack API docs for ActionsBlock (https://api.slack.com/reference/block-kit/blocks#actions_examples)
2.Create an ActionsBlock from parsed (to dict) json
3. Check the type of elements attribute from created ActionsBlock object
### Expected result:
Elements should be instances of BlockElement, the same as it is for SectionBlock.accessory
### Actual result:
Elements of ActionsBlock is not parsed
| closed | 2020-12-23T12:07:14Z | 2020-12-24T06:31:24Z | https://github.com/slackapi/python-slack-sdk/issues/903 | [
"bug",
"Version: 3x",
"good first issue"
] | KharchenkoDmitriy | 3 |
pyg-team/pytorch_geometric | deep-learning | 9,310 | ModuleNotFoundError: 'CuGraphSAGEConv' requires 'pylibcugraphops>=23.02' | ### 😵 Describe the installation problem
I am trying to run this code block:
```
import torch.nn.functional as F
from torch_geometric.nn import CuGraphSAGEConv
class CuGraphSAGE(torch.nn.Module):
def __init__(self, in_channels, hidden_channels, out_channels, num_layers):
super().__init__()
self.convs = torch.nn.ModuleList()
self.convs.append(CuGraphSAGEConv(in_channels, hidden_channels))
for _ in range(num_layers - 1):
conv = CuGraphSAGEConv(hidden_channels, hidden_channels)
self.convs.append(conv)
self.lin = torch.nn.Linear(hidden_channels, out_channels)
def forward(self, x, edge, size):
edge_csc = CuGraphSAGEConv.to_csc(edge, (size[0], size[0]))
for conv in self.convs:
x = conv(x, edge_csc)[: size[1]]
x = F.relu(x)
x = F.dropout(x, p=0.5)
return self.lin(x)
model = CuGraphSAGE(128, 64, 349, 3).to(torch.float32).to("cuda")
optimizer = torch.optim.Adam(model.parameters(), lr=0.01)
```
but I get this:
```
---------------------------------------------------------------------------
ModuleNotFoundError Traceback (most recent call last)
Cell In[26], line 25
21 x = F.dropout(x, p=0.5)
23 return self.lin(x)
---> 25 model = CuGraphSAGE(128, 64, 349, 3).to(torch.float32).to("cuda")
26 optimizer = torch.optim.Adam(model.parameters(), lr=0.01)
Cell In[26], line 9, in CuGraphSAGE.__init__(self, in_channels, hidden_channels, out_channels, num_layers)
6 super().__init__()
8 self.convs = torch.nn.ModuleList()
----> 9 self.convs.append(CuGraphSAGEConv(in_channels, hidden_channels))
10 for _ in range(num_layers - 1):
11 conv = CuGraphSAGEConv(hidden_channels, hidden_channels)
File ~/.conda/envs/rapids-24.04/lib/python3.11/site-packages/torch_geometric/nn/conv/cugraph/sage_conv.py:40, in CuGraphSAGEConv.__init__(self, in_channels, out_channels, aggr, normalize, root_weight, project, bias)
30 def __init__(
31 self,
32 in_channels: int,
(...)
38 bias: bool = True,
39 ):
---> 40 super().__init__()
42 if aggr not in ['mean', 'sum', 'min', 'max']:
43 raise ValueError(f"Aggregation function must be either 'mean', "
44 f"'sum', 'min' or 'max' (got '{aggr}')")
File ~/.conda/envs/rapids-24.04/lib/python3.11/site-packages/torch_geometric/nn/conv/cugraph/base.py:41, in CuGraphModule.__init__(self)
38 super().__init__()
40 if not HAS_PYLIBCUGRAPHOPS and not LEGACY_MODE:
---> 41 raise ModuleNotFoundError(f"'{self.__class__.__name__}' requires "
42 f"'pylibcugraphops>=23.02'")
ModuleNotFoundError: 'CuGraphSAGEConv' requires 'pylibcugraphops>=23.02'
```
### Environment
* PyG version:2.5.3
* PyTorch version: 2.1.2.post303
* OS: Windows 11
* Python version: 3.11.9
* CUDA/cuDNN version: 12.2, V12.2.140
* How you installed PyTorch and PyG (`conda`, `pip`, source): conda
* Any other relevant information (*e.g.*, version of `torch-scatter`):
| closed | 2024-05-10T21:36:59Z | 2024-07-22T00:36:21Z | https://github.com/pyg-team/pytorch_geometric/issues/9310 | [
"installation"
] | d3netxer | 2 |
microsoft/nni | data-science | 4,756 | Current support of Retiarii for TensorFlow | I am interesting in evaluating Retiarii for my particular use-case. Looking at the examples, and issues, it seems like currently only Pytorch is supported. However, I do see some mention of TensorFlow support to be introduced in the roadmap [V2.4](https://github.com/microsoft/nni/discussions/3744)
My question is then: Can retiarii be used with TensorFlow at the moment ?
I am particularly interested in using ProxylessNAS and FBNet | closed | 2022-04-12T13:34:33Z | 2022-04-13T08:00:09Z | https://github.com/microsoft/nni/issues/4756 | [] | Hrayo712 | 2 |
open-mmlab/mmdetection | pytorch | 11,747 | TypeError: __init__() got an unexpected keyword argument 'pretrained' |
**Describe the bug**
i try to run the following code:
from mmseg.apis import init_model, inference_model, show_result_pyplot
import mmcv
from mmcv import imread
import mmengine
from mmengine.registry import init_default_scope
import numpy as np
from mmpose.apis import inference_topdown
from mmpose.apis import init_model as init_pose_estimator
from mmpose.evaluation.functional import nms
from mmpose.registry import VISUALIZERS
from mmpose.structures import merge_data_samples
try:
from mmdet.apis import inference_detector, init_detector
has_mmdet = True
except (ImportError, ModuleNotFoundError):
has_mmdet = False
local_runtime = True
img = 'img.png'
rear_pose_config = 'models/v1/rear_pose/res152_animalpose_256x256.py'
rear_pose_checkpoint = 'models/v1/rear_pose/epoch_210.pth'
side_pose_config = 'models/v1/side_pose/res152_animalpose_256x256.py'
side_pose_checkpoint = 'models/v1/side_pose/epoch_210.pth'
det_config = 'models/v1/det/faster_rcnn_r50_fpn_coco.py'
det_checkpoint = 'models/v1/det/faster_rcnn_r50_fpn_1x_coco_20200130-047c8118.pth'
device = 'cpu'
cfg_options = dict(model=dict(test_cfg=dict(output_heatmaps=True)))
detector = init_detector(
det_config,
det_checkpoint,
device=device
)
side_pose_estimator = init_pose_estimator(
side_pose_config,
side_pose_checkpoint,
device=device,
cfg_options=cfg_options
)
side_pose_estimator.cfg.visualizer.radius = 3
side_pose_estimator.cfg.visualizer.line_width = 1
visualizer = VISUALIZERS.build(side_pose_estimator.cfg.visualizer)
def visualize_img(img_path, detector, pose_estimator, visualizer,
show_interval, out_file):
"""Visualize predicted keypoints (and heatmaps) of one image."""
# predict bbox
scope = detector.cfg.get('default_scope', 'mmdet')
if scope is not None:
init_default_scope(scope)
detect_result = inference_detector(detector, img_path)
pred_instance = detect_result.pred_instances.cpu().numpy()
bboxes = np.concatenate(
(pred_instance.bboxes, pred_instance.scores[:, None]), axis=1)
bboxes = bboxes[np.logical_and(pred_instance.labels == 0,
pred_instance.scores > 0.3)]
bboxes = bboxes[nms(bboxes, 0.3)][:, :4]
# predict keypoints
pose_results = inference_topdown(pose_estimator, img_path, bboxes)
data_samples = merge_data_samples(pose_results)
# show the results
img = mmcv.imread(img_path, channel_order='rgb')
visualizer.add_datasample(
'result',
img,
data_sample=data_samples,
draw_gt=False,
draw_heatmap=True,
draw_bbox=True,
show=False,
wait_time=show_interval,
out_file=out_file,
kpt_thr=0.3)
visualize_img(
'img.png',
detector,
side_pose_estimator,
visualizer,
show_interval=0,
out_file=None)
vis_result = visualizer.get_image()
and get the following error:
Traceback (most recent call last):
File "/Users/aftaabhussain/Work/Implementation cattle/predict.py", line 33, in <module>
detector = init_detector(
File "/Users/aftaabhussain/Work/Implementation cattle/.venv/lib/python3.9/site-packages/mmdet/apis/inference.py", line 66, in init_detector
model = MODELS.build(config.model)
File "/Users/aftaabhussain/Work/Implementation cattle/.venv/lib/python3.9/site-packages/mmengine/registry/registry.py", line 570, in build
return self.build_func(cfg, *args, **kwargs, registry=self)
File "/Users/aftaabhussain/Work/Implementation cattle/.venv/lib/python3.9/site-packages/mmengine/registry/build_functions.py", line 232, in build_model_from_cfg
return build_from_cfg(cfg, registry, default_args)
File "/Users/aftaabhussain/Work/Implementation cattle/.venv/lib/python3.9/site-packages/mmengine/registry/build_functions.py", line 121, in build_from_cfg
obj = obj_cls(**args) # type: ignore
TypeError: __init__() got an unexpected keyword argument 'pretrained'
i checked the init() file but there is no keyword argument called pretrained present over there,
my environment:
pytorch version - 2.3.0 cuda available = False
mmseg version 1.2.2
mmpose version 1.1.0
mmdet version 3.3.0
| open | 2024-05-28T15:37:41Z | 2025-02-04T15:20:52Z | https://github.com/open-mmlab/mmdetection/issues/11747 | [] | theaftaab | 1 |
jupyterlab/jupyter-ai | jupyter | 852 | /export command uses name "Agent" for Jupyternaut's messages | When I use the `/export` command (see #658), the name used for Jupyternaut's messages is "Agent", not "Jupyternaut". If we let users change the default character's name, or create additional characters for AI assistance, this will be ambiguous. We should use the same character name in the chat panel and in exported files.
```
**Agent**: Hi there! I'm Jupyternaut, your programming assistant.
You can ask me a question using the text box below. You can also use these commands:
* `/ask` — Ask a question about your learned data
* `/clear` — Clear the chat window
* `/generate` — Generate a Jupyter notebook from a text prompt
* `/learn` — Teach Jupyternaut about files on your system
* `/export` — Export chat history to a Markdown file
* `/fix` — Fix an error cell selected in your notebook
* `/help` — Display this help message
Jupyter AI includes [magic commands](https://jupyter-ai.readthedocs.io/en/latest/users/index.html#the-ai-and-ai-magic-commands) that you can use in your notebooks.
For more information, see the [documentation](https://jupyter-ai.readthedocs.io).
**jweill**: /help
``` | closed | 2024-06-21T16:40:00Z | 2024-06-21T23:42:39Z | https://github.com/jupyterlab/jupyter-ai/issues/852 | [
"bug",
"scope:chat-ux"
] | JasonWeill | 0 |
pyjanitor-devs/pyjanitor | pandas | 731 | Add an `also` method akin to Kotlins `also` | # Brief Description
I would like to propose a new function `also` that takes a function with a single dataframe for input, runs the function on the chained dataframe, and returns the input dataframe.
I would love to take a whack at implementing this if you're interested incorporating it into the library.
As I've been trying to adopt chaining as much as possible, I've routinely bumped into scenarios where I need to break the chain for one reason or another. This API would eliminate most reasons for breaking the chain
This function is inspired by Kotlin's `also` function. If we were to mimic Kotlin's `also`, the signature would look something like
`also: (f: T -> None) -> T`
It would behave very similar to `pipe` or `then`, but can be used on functions that do not return anything. This can be very useful for any sort of side-effect operations within a long chain, such as periodic logging or debugging.
# Example API
<!-- One of the selling points of pyjanitor is the API. Hence, we guard the API very carefully, and want to
make sure that it is accessible and understandable to many people. Please provide a few examples of what the API
of the new function you're proposing will look like. We have provided an example that you should modify. -->
Below are some examples of how this API would simplify code I've written in the past.
### print the df while processing.
```python
# instead of defining an extra function
def print_and(df):
print(df)
return df
df.groupby(["column1"]).agg(sum).then(print_and).then(my_transform)
# or breaking the chain up
df_tmp = df.groupby(["column1"]).agg(sum)
print(df_tmp)
df_tmp.then(my_transform)
# you could just inline the print statement
df.groupby(["column1"]).agg(sum).also(print).then(my_transform)
```
### Insert a breakpoint while debugging, but keep going when you're done
Similar to above, to add a breakpoint for debugging, you would either have to create a new `pipe` compatible function or break the chain into smaller steps.
Instead, you could chain a breakpoint like so
```python
df.groupby(["column1"]).agg(sum).also(lambda x: breakpoint()).then(my_transform)
```
### Save a midpoint of a processing chain and then keep going
Another possibility would be running IO in the middle of the chain. Often, a midpoint in the chain is just as important as the end result. `also` would allow me to save the results and keep going without breaking a chain.
```
df.groupby(["column1"]).agg(sum).also(lambda x: x.to_csv("filename.csv)).then(my_transform)
```
### pipe-dream: Save a midpoint as a variable
I haven't figured out how to get this to work given Python's scoping rules, maybe the new `:=` operator will make this possible.
This could be really helpful in scenarios where you're running groupbys or other aggregations and want to join those back into an ungrouped frame.
```
df.groupby(["column1"]).agg(sum).also(lambda x: df_mid = x).then(my_transform).assign(original_var = df_mid)
```
| closed | 2020-09-03T20:27:08Z | 2020-09-10T23:51:56Z | https://github.com/pyjanitor-devs/pyjanitor/issues/731 | [] | sauln | 9 |
openapi-generators/openapi-python-client | fastapi | 656 | x-www-form-urlencoded in requestBody: no parameters generated | I am trying to generate a client for the following spec:
```yaml
openapi: 3.0.3
info:
title: Test
version: 1.2.3
paths:
/bar:
post:
summary: Do something
requestBody:
required: true
content:
application/x-www-form-urlencoded:
schema:
type: object
required: [ form_param ]
properties:
form_param:
type: string
responses:
200:
description: ""
```
The output however doesn't contain any parameters:
```python
def sync_detailed(
*,
client: Client,
) -> Response[Any]:
"""Do something
Returns:
Response[Any]
"""
...
```
I am using the latest version:
```
❯ openapi-python-client --version
openapi-python-client version: 0.11.5
```
Any hints? Is this a missing feature? From the closed issues, I got the impression that form data should be supported, in general?
The parameter is rendered correctly in Swagger UI / editor, so I am pretty sure that the spec is correct. | closed | 2022-08-19T09:24:29Z | 2022-08-27T21:25:59Z | https://github.com/openapi-generators/openapi-python-client/issues/656 | [] | supermihi | 1 |
alyssaq/face_morpher | numpy | 24 | ImportError: No module named 'locator' | I tried using the project as a library.
Python 3.5, OpenCV 3.3, Ubuntu 16.04
I have images in the `imgs` folder.
Code-
```
(cv) ubuntu@ip:~/face_morpher_aslib$ cat fm.py
import facemorpher
facemorpher.averager(['imgs/trump.jpg', 'imgs/madara.jpg'], plot=True)
```
But, when I try to run the program, I end up with the following error-
```
(cv) ubuntu@ip:~/face_morpher_aslib$ python fm.py
Traceback (most recent call last):
File "fm.py", line 1, in <module>
import facemorpher
File "/home/ubuntu/.virtualenvs/cv/lib/python3.5/site-packages/facemorpher/__init__.py", line 5, in <module>
from .morpher import morpher, list_imgpaths
File "/home/ubuntu/.virtualenvs/cv/lib/python3.5/site-packages/facemorpher/morpher.py", line 34, in <module>
import locator
ImportError: No module named 'locator'
```
I also did a pip search, couldn't find the module.
```
(cv) ubuntu@ip:~/face_morpher_aslib$ pip search locator
resource-allocator (0.1.0) - Python Resource Allocation API
bareon-allocator (1.0.0.dev) -
allocator (0.1.7) - Optimally Allocate Geographically Distributed Tasks
celery_geolocator (0.0.11) - Celery Geolocator
django-cms-storelocator (1.10.0) - A store locator extension for Django CMS
consul-locator (0.1.6) - python consul discovery locator for http
robotframework-selenium2library-divfor (1.8.0.1) - Web testing library for Robot Framework with locator wrapper feature
django-storelocator (0.1) - Django Storelocator is a Django App for locating stores near a geographical location.
django-locator (1.0.0) - An easy to integrate store locator plugin for Django.
djangocms-store-locator (0.1.3) - A simple store locator django CMS plugin that uses Google Maps.
EditorConfig (0.12.1) - EditorConfig File Locator and Interpreter for Python
geolocator (0.1.1) - geolocator library: locate places and calculate distances between them
inmatelocator (0.0.8) - Library for locating people incarcerated in the US by querying official prison search tools
service-locator (0.1.3) - Dead simple python service locator
selenium-smart-locator (0.2.0) - A (somewhat) smart locator class for Selenium.
luogo (0.1.0) - A service locator for python.
maidenhead (1.1.1) - Maidenhead Locator
mlocs (1.0.5) - Effective Location Storage via a Maidenhead Locator System For Python
modulocator (0.1) - Want to use a python module you're developing in you Jupyter notebook? Use modulocator to do that
osxrelocator (1.0.1) - Utility to relocate OSX libraries
pylisp (0.4.2) - Locator/ID Separation Protocol (LISP) library
pylocator (1.0.beta.1.dev) - Program for the localization of EEG-electrodes.
relocator (0.1) - change Location field in responses using WSGI middleware
solu (0.1) - Self-service Office resource Locator and Updater
Task_allocator (1.0.2) - This program takes commandline input for the datafile from where configuration will be fetched.
ulp (1.1.1) - ULP is a Locator Picker - a PathPicker clone for URLs
```
Please can you help me fix this? | closed | 2017-08-23T10:47:29Z | 2017-08-23T11:16:09Z | https://github.com/alyssaq/face_morpher/issues/24 | [] | bozzmob | 1 |
explosion/spaCy | data-science | 13,710 | Unable to finetune transformer based ner model after initial tuning | ### Discussed in https://github.com/explosion/spaCy/discussions/13394
<div type='discussions-op-text'>
<sup>Originally posted by **jlustgarten** March 23, 2024</sup>
<!-- NOTE: For questions or install related issues, please open a Discussion instead. -->
## How to reproduce the behaviour
Create a transformer ner model
Train it on data using the cfg and cli which auto-saves it
Create a new cfg file that points to your existing model
Try triggering the training using the CLI
You will get a missing config.json error
## Your Environment
<!-- Include details of your environment. You can also type `python -m spacy info --markdown` and copy-paste the result here.-->
- **spaCy version:** 3.7.2
- **Platform:** Linux-5.15.146.1-microsoft-standard-WSL2-x86_64-with-glibc2.35
- **Python version:** 3.10.13
</div>
This still is occurring with the same text:
Config:
[paths]
train = null
dev = null
vectors = null
init_tok2vec = null
[system]
gpu_allocator = "pytorch"
seed = 0
[nlp]
lang = "en"
pipeline = ["transformer","ner"]
batch_size = 128
disabled = []
before_creation = null
after_creation = null
after_pipeline_creation = null
tokenizer = {"@Tokenizers":"spacy.Tokenizer.v1"}
vectors = {"@vectors":"spacy.Vectors.v1"}
[components]
[components.ner]
factory = "ner"
incorrect_spans_key = null
moves = null
scorer = {"https://github.com/scorers":"spacy.ner_scorer.v1"}
update_with_oracle_cut_size = 100
[components.ner.model]
@architectures = "spacy.TransitionBasedParser.v2"
state_type = "ner"
extra_state_tokens = false
hidden_width = 64
maxout_pieces = 2
use_upper = false
nO = null
[components.ner.model.tok2vec]
@architectures = "spacy-transformers.TransformerListener.v1"
grad_factor = 1.0
pooling = {"https://github.com/layers":"reduce_mean.v1"}
upstream = "*"
[components.transformer]
factory = "transformer"
max_batch_items = 4096
set_extra_annotations = {"@annotation_setters":"spacy-transformers.null_annotation_setter.v1"}
[components.transformer.model]
@architectures = "spacy-transformers.TransformerModel.v3"
name = "/home/user/Coding/PatientHistory/original_pt_hist_ner"
mixed_precision = false
[components.transformer.model.get_spans]
@span_getters = "spacy-transformers.strided_spans.v1"
window = 128
stride = 96
[components.transformer.model.grad_scaler_config]
[components.transformer.model.tokenizer_config]
use_fast = true
[components.transformer.model.transformer_config]
[corpora]
[corpora.dev]
@readers = "spacy.Corpus.v1"
path = ${paths.dev}
max_length = 0
gold_preproc = false
limit = 0
augmenter = null
[corpora.train]
@readers = "spacy.Corpus.v1"
path = ${paths.train}
max_length = 0
gold_preproc = false
limit = 0
augmenter = null
[training]
accumulate_gradient = 4
dev_corpus = "corpora.dev"
train_corpus = "corpora.train"
seed = ${system.seed}
gpu_allocator = ${system.gpu_allocator}
dropout = 0.1
patience = 2000
max_epochs = 0
max_steps = 80000
eval_frequency = 200
frozen_components = []
annotating_components = []
before_to_disk = null
before_update = null
[training.batcher]
@batchers = "spacy.batch_by_padded.v1"
discard_oversize = false
size = 2000
buffer = 256
get_length = null
[training.logger]
@Loggers = "spacy.ConsoleLogger.v1"
progress_bar = false
[training.optimizer]
https://github.com/optimizers = "Adam.v1"
beta1 = 0.9
beta2 = 0.999
L2_is_weight_decay = true
L2 = 0.01
grad_clip = 1.0
use_averages = false
eps = 0.00000001
[training.optimizer.learn_rate]
https://github.com/schedules = "warmup_linear.v1"
warmup_steps = 250
total_steps = 200000
initial_rate = 0.00005
[training.score_weights]
ents_f = 1.0
ents_p = 0.0
ents_r = 0.0
ents_per_type = null
[pretraining]
[initialize]
vectors = ${paths.vectors}
init_tok2vec = ${paths.init_tok2vec}
vocab_data = null
lookups = null
before_init = null
after_init = null
[initialize.components]
[initialize.tokenizer]
Here's the CLI:
python -m spacy train '/home/user/Coding/PatientHistory/refine_pt_hist_ner.cfg' --output '/home/user/Coding/PatientHistory/improved_pt_hist_3_22_2024' --paths.train '/home/user/Coding/PatientHistory/train.spacy' --paths.dev '/home/user/Coding/PatientHistory/test.spacy' --gpu-id 0
Here's the output:
ℹ Saving to output directory:
/home/user/Coding/PatientHistory/improved_pt_hist_3_22_2024
ℹ Using GPU: 0
=========================== Initializing pipeline ===========================
/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/transformers/utils/generic.py:441: UserWarning: torch.utils._pytree._register_pytree_node is deprecated. Please use torch.utils._pytree.register_pytree_node instead.
_torch_pytree._register_pytree_node(
/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/transformers/utils/generic.py:309: UserWarning: torch.utils._pytree._register_pytree_node is deprecated. Please use torch.utils._pytree.register_pytree_node instead.
_torch_pytree._register_pytree_node(
Traceback (most recent call last):
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/runpy.py", line 196, in _run_module_as_main
return _run_code(code, main_globals, None,
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/runpy.py", line 86, in _run_code
exec(code, run_globals)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/spacy/main.py", line 4, in
setup_cli()
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/spacy/cli/_util.py", line 87, in setup_cli
command(prog_name=COMMAND)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/click/core.py", line 1157, in call
return self.main(*args, **kwargs)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/typer/core.py", line 778, in main
return _main(
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/typer/core.py", line 216, in _main
rv = self.invoke(ctx)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/click/core.py", line 1688, in invoke
return _process_result(sub_ctx.command.invoke(sub_ctx))
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/click/core.py", line 1434, in invoke
return ctx.invoke(self.callback, **ctx.params)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/click/core.py", line 783, in invoke
return __callback(*args, **kwargs)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/typer/main.py", line 683, in wrapper
return callback(**use_params) # type: ignore
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/spacy/cli/train.py", line 54, in train_cli
train(config_path, output_path, use_gpu=use_gpu, overrides=overrides)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/spacy/cli/train.py", line 81, in train
nlp = init_nlp(config, use_gpu=use_gpu)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/spacy/training/initialize.py", line 95, in init_nlp
nlp.initialize(lambda: train_corpus(nlp), sgd=optimizer)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/spacy/language.py", line 1349, in initialize
proc.initialize(get_examples, nlp=self, **p_settings)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/spacy_transformers/pipeline_component.py", line 351, in initialize
self.model.initialize(X=docs)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/thinc/model.py", line 318, in initialize
self.init(self, X=X, Y=Y)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/spacy_transformers/layers/transformer_model.py", line 131, in init
hf_model = huggingface_from_pretrained(name, tok_cfg, trf_cfg)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/spacy_transformers/layers/transformer_model.py", line 267, in huggingface_from_pretrained
tokenizer = tokenizer_cls.from_pretrained(str_path, **tok_config)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/transformers/models/auto/tokenization_auto.py", line 752, in from_pretrained
config = AutoConfig.from_pretrained(
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/transformers/models/auto/configuration_auto.py", line 1082, in from_pretrained
config_dict, unused_kwargs = PretrainedConfig.get_config_dict(pretrained_model_name_or_path, **kwargs)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/transformers/configuration_utils.py", line 644, in get_config_dict
config_dict, kwargs = cls._get_config_dict(pretrained_model_name_or_path, **kwargs)
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/transformers/configuration_utils.py", line 699, in _get_config_dict
resolved_config_file = cached_file(
File "/home/user/miniconda3/envs/patienthistoryclassifier/lib/python3.10/site-packages/transformers/utils/hub.py", line 360, in cached_file
raise EnvironmentError(
OSError: /home/user/Coding/PatientHistory/original_pt_hist_ner does not appear to have a file named config.json. Checkout 'https://huggingface.co//home/user/Coding/PatientHistory/original_pt_hist_ner/None' for available files. | open | 2024-12-06T04:55:54Z | 2024-12-06T04:55:54Z | https://github.com/explosion/spaCy/issues/13710 | [] | jlustgarten | 0 |
mljar/mercury | jupyter | 210 | YAML cell is not detected | I have setup a yamls cell (the first one) as RAW but the widgets on it are not shown.
> title: PSFB dashboard
>
> author: xxx
> description: yyyyyyyy
> show-code: False
> params:
> primary_switch:
> label: This is select label
> input: select
> value: Cześć
> choices: [Cześć, Hi, Hello]
> multi: False
> Fsw:
> input: numeric
> label: Switching frequency [Hz]
> value: 50000
> min: 0
> max: 300000
> step: 1000
the second cell has the variables assigned a default value.
When running the notebook, no single widget appears. Am I missing anything?
thanks!!
| closed | 2023-02-15T14:56:50Z | 2023-02-16T14:43:28Z | https://github.com/mljar/mercury/issues/210 | [] | mizamae | 1 |
hbldh/bleak | asyncio | 1,299 | Add support for adapter selection in Windows | * bleak version: master
* Python version: all
* Operating System: Windows
### Description
bluezdbus supports passing the adapter to use as parameter (e.g. https://github.com/hbldh/bleak/pull/524). Winrt doesn't. Would it be possible to add this support for Windows as well?
Thanks!
| closed | 2023-05-04T13:54:29Z | 2023-05-04T14:42:34Z | https://github.com/hbldh/bleak/issues/1299 | [] | eranzim | 1 |
vitalik/django-ninja | rest-api | 884 | [Feature request] simple way to generate error schema from validation error for models | **Is your feature request related to a problem? Please describe.**
Generating schemas from model fields is super easy and a huge boon to productivity. However, on the other side of the happy path I have not had good luck finding a good pattern on how to map model errors into a schema. The best I've been able to do would be something like this...
```python
class Widget(Model):
name = models.TextField()
amount = models.IntegerField()
class WidgetErrorSchema(Schema):
name = Optional[List[str]]
amount = Optional[List[str]]
@api.post("/widgets", response={200: WidgetModelSchema, 400: WidgetErrorSchema})
def create_widget(request, payload: WidgetCreateSchema):
widget = Widget(**payload.dict())
try:
widget.full_clean()
except ValidationError as e:
print(e.message_dict)
return 400, e.messages_dict
widget.save()
return widget
```
But this approach has at least one gotcha to it that I have run into. If your validation puts an error on the '__all__' field of the model error dictionary that will not come across in this schema. My immediate reaction was to try...
```python
class WidgetErrorSchema(Schema):
__all__: Optional[List[str]]
name = Optional[List[str]]
amount = Optional[List[str]]
```
But that did not appear to work either. What I actually had to do was...
```python
class WidgetErrorSchema(Schema):
all: Optional[List[str]]
name = Optional[List[str]]
amount = Optional[List[str]]
# ... abbreviated code
def create_widget(request, payload: WidgetCreateSchema):
widget = Widget(**payload.dict())
try:
widget.full_clean()
except ValidationError as e:
print(e.message_dict)
return 400, {"all": e.message_dict.get("__all__", []), **e.message_dict}
# ...
```
And that 'works' but feels very hacky and error-prone to duplicate that pattern across multiple model CRUD endpoints.
**Describe the solution you'd like**
It would be nice if Ninja had a built-in way to generate a schema for a model's validation errors similar to how you can generate a model fields schema.
| closed | 2023-10-21T22:54:00Z | 2023-10-22T20:24:16Z | https://github.com/vitalik/django-ninja/issues/884 | [] | RileyMathews | 2 |
scrapy/scrapy | python | 6,118 | Infinite request: producer-consumer scrapy | I am using redis achieve producer-consumer scrapy, as we can see in the code, if code get empty urls, It will fall into a loop, and if there are still running request tasks, then those request tasks will not be process.
```python
class ExampleSpider(Spider):
name = "ExampleSpider"
@property
def start_urls(self):
while True:
urls = self.redis.keys()
if len(urls) == 0:
time.sleep(0.001)
continue
for url in urls:
yield url
```
I want to know if there is a way to release the main process control loop like this?
```python
import asyncio
class ExampleSpider(Spider):
name = "ExampleSpider"
@property
def start_urls(self):
while True:
urls = self.redis.keys()
if len(urls) == 0:
await asyncio.sleep(0.001)
continue
for url in urls:
yield url
```
I use this way to implement now, but I think it is not very good, It is achieved by set Request params ```dont_filter=Flase```:
```python
class ExampleSpider(Spider):
name = "ExampleSpider"
@property
def start_urls(self):
while True:
urls = self.redis.keys()
if len(urls) == 0:
time.sleep(0.001)
flag = 0
yield flag, "https://www.google.com"
continue
for url in urls:
flag = 1
yield flag, url
def start_requests(self):
for flag, url in self.start_urls:
if flag:
yield Request(url , lambda x: None, dont_filter=True)
else:
yield Request(url , lambda x: None)
```
Please give me some advice, thanks! | closed | 2023-10-19T09:47:00Z | 2023-10-19T10:25:36Z | https://github.com/scrapy/scrapy/issues/6118 | [] | Yakuho | 1 |
huggingface/transformers | nlp | 36,384 | Set non_blocking=True When moving data from the CPU to the GPU | ### System Info
No Need
### Who can help?
_No response_
### Information
- [ ] The official example scripts
- [x] My own modified scripts
### Tasks
- [ ] An officially supported task in the `examples` folder (such as GLUE/SQuAD, ...)
- [x] My own task or dataset (give details below)
### Reproduction
I used the transformer's Trainer to train the model, but used my own Dataloader.
Then I used pytorch profile to check my training performance and found that the CPU execution time accounted for a high proportion

After a period of investigation, it was found that the non_blocking was not set when the data was transferred from the CPU to the GPU.
https://github.com/huggingface/transformers/blob/v4.49.0/src/transformers/trainer.py#L3625-L3631
The modified code is:
```python
kwargs = {"device": self.args.device,“non_blocking”: True}
```
Then I re-profiled my code and the results were as follows:

You can see that the performance has been greatly improved.
I'm not sure if this is a bug in the code or a problem with the way I'm using it.
But there is no doubt that setting non_blocking=True has brought a great performance improvement to my training.
Looking forward to your reply
### Expected behavior
No Need | open | 2025-02-25T03:31:37Z | 2025-02-26T03:11:34Z | https://github.com/huggingface/transformers/issues/36384 | [
"bug"
] | Hukongtao | 3 |
oegedijk/explainerdashboard | dash | 94 | Can not handel stack models | I am trying to train and deploy a stack model but I get the following error!
````
ValueError: Parameter shap='gues'', but failed to to guess the type of shap explainer to use. Please explicitly pass a `shap` parameter to the explainer, e.g. shap='tree', shap='linear', etc.
````
Here is the code I used.
````
from explainerdashboard.datasets import titanic_fare, titanic_names
from explainerdashboard.datasets import feature_descriptions
from explainerdashboard import RegressionExplainer
from explainerdashboard import ClassifierExplainer, ExplainerDashboard
from sklearn.datasets import load_diabetes
from sklearn.linear_model import RidgeCV
from sklearn.svm import SVR
from sklearn.ensemble import RandomForestRegressor
from sklearn.ensemble import StackingRegressor
estimators = [
('lr', RidgeCV()),
('svr', SVR())
]
model = StackingRegressor(
estimators=estimators,
final_estimator=RandomForestRegressor(random_state=42))
# model.fit(X_train, y_train)
# model.score(X_test, y_test)
explainer = RegressionExplainer(model, X_test, y_test,
cats=['Sex', 'Deck', 'Embarked'],
idxs=test_names,
descriptions = feature_descriptions,
target='Fare',
units="$")
ExplainerDashboard(explainer, mode='inline').run()
```` | closed | 2021-03-03T01:55:42Z | 2021-03-03T19:30:17Z | https://github.com/oegedijk/explainerdashboard/issues/94 | [] | jkiani64 | 2 |
ymcui/Chinese-LLaMA-Alpaca | nlp | 147 | Resume损失与断点处差异较大 | 请问大家,断点续训时损失值的大小能接上吗,resume发现无论是load checkpoint还是最后保存的adapters_model.bin 损失值都接不上 | closed | 2023-04-13T03:04:37Z | 2023-04-26T12:00:40Z | https://github.com/ymcui/Chinese-LLaMA-Alpaca/issues/147 | [
"stale"
] | xxr11 | 1 |
proplot-dev/proplot | data-visualization | 145 | Midpoint normalizer doesn't work well for data with large difference | ### Description
Midpoint normalizer doesn't work well for data with large difference.
Zero value doesn't divide the colormap.
### Steps to reproduce
```python
import proplot as plot
import numpy as np
state = np.random.RandomState(51423)
data1 = (state.rand(20, 20) - 0.43).cumsum(axis=0)
data2 = (state.rand(20, 20) - 0.57).cumsum(axis=0)
data1[data1>2] *= 5
data2[data2<-3] *= 5
f, axs = plot.subplots(ncols=2, axwidth=2.5, aspect=1.5)
cmap = plot.Colormap('DryWet', cut=0.1)
axs.format(suptitle='Midpoint normalizer demo')
for ax, data, mode in zip(axs, (data1, data2), ('positive', 'negative')):
m = ax.contourf(data, norm='midpoint', cmap='Div')
ax.colorbar(m, loc='b')
ax.format(title=f'Skewed {mode} data')
```
**Expected behavior**: [What you expected to happen]
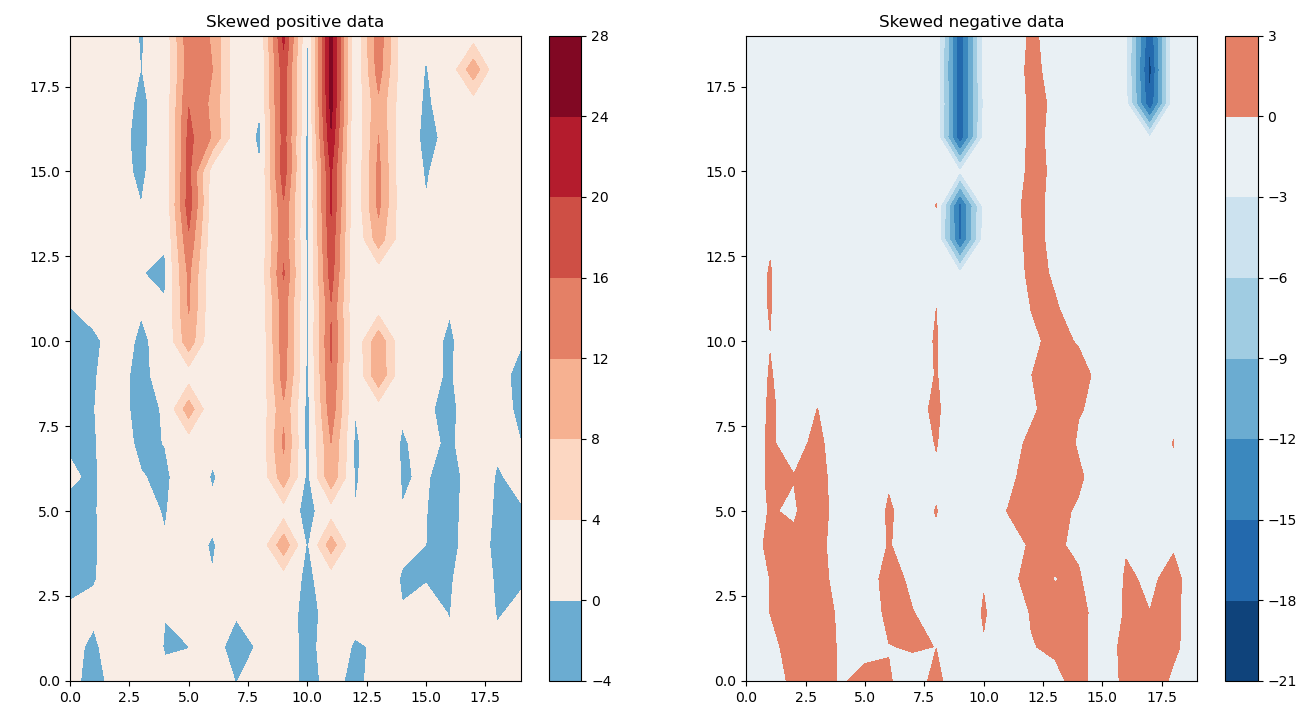
**Actual behavior**: [What actually happened]
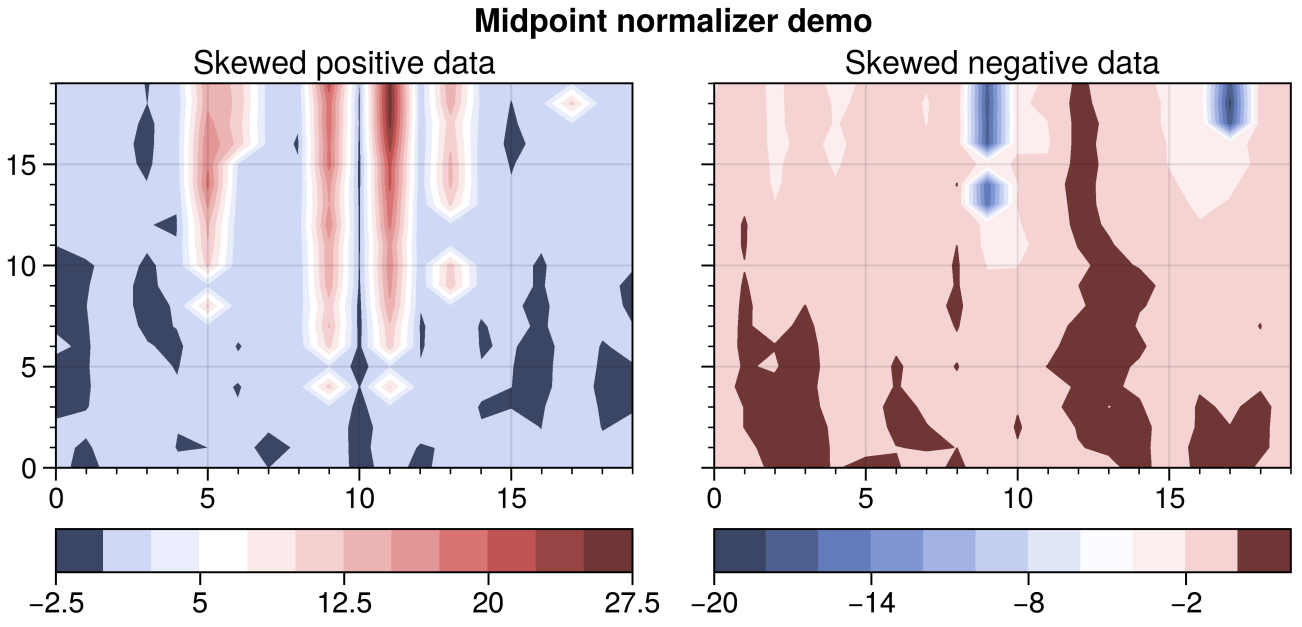
### Equivalent steps in matplotlib
```python
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
class MidpointNormalize(matplotlib.colors.Normalize):
def __init__(self, vmin=None, vmax=None, midpoint=None, clip=False):
self.midpoint = midpoint
matplotlib.colors.Normalize.__init__(self, vmin, vmax, clip)
def __call__(self, value, clip=None):
# Note that I'm ignoring clipping and other edge cases here.
result, is_scalar = self.process_value(value)
x, y = [self.vmin, self.midpoint, self.vmax], [0, 0.5, 1]
return np.ma.array(np.interp(value, x, y), mask=result.mask, copy=False)
state = np.random.RandomState(51423)
data1 = (state.rand(20, 20) - 0.43).cumsum(axis=0)
data2 = (state.rand(20, 20) - 0.57).cumsum(axis=0)
data1[data1>2] *= 5
data2[data2<-3] *= 5
f, axs = plt.subplots(1, 2)
for ax, data, mode in zip(axs, (data1, data2), ('positive', 'negative')):
m = ax.contourf(data, norm=MidpointNormalize(midpoint=0), cmap='RdBu_r')
plt.colorbar(m, ax=ax)
ax.set_title(f'Skewed {mode} data')
```
### Proplot version
0.5.0 | closed | 2020-04-26T08:31:03Z | 2020-05-10T08:25:55Z | https://github.com/proplot-dev/proplot/issues/145 | [
"bug"
] | zxdawn | 3 |
strawberry-graphql/strawberry | graphql | 3,074 | Strawberry trivia | <!--- Provide a general summary of the changes you want in the title above. -->
<!--- Anything on lines wrapped in comments like these will not show up in the final text. -->
I've been using strawberry at work for a few months now and can't get these silly things out of my mind.
- Why is it called strawberry?
- Who made that logo? It deserves a credit.
- Am I the only one who thinks of the Beatles song when I type `strawberry.field`? Should `strawberry.fields.forever` be some kind of utility or easter egg?
I'd like answers. And those things should be in the documentation for weird people like me 🤪 | open | 2023-09-05T06:08:53Z | 2025-03-20T15:56:21Z | https://github.com/strawberry-graphql/strawberry/issues/3074 | [] | brunodantas | 2 |
ultralytics/ultralytics | deep-learning | 18,815 | TensorBoard does not work properly in Docker | ### Search before asking
- [x] I have searched the Ultralytics YOLO [issues](https://github.com/ultralytics/ultralytics/issues) and found no similar bug report.
### Ultralytics YOLO Component
Train
### Bug
Training with docker images:
```
docker run -it --rm \
--ipc=host \
--gpus all \
-p 6006:6006 \
ultralytics/ultralytics:8.3.40 \
yolo detect train data=coco8.yaml model=yolo11n.pt
```
see the following message:
TensorBoard: Start with 'tensorboard --logdir /ultralytics/runs/detect/train', view at http://localhost:6006/
But the browser shows it as follows:

### Environment
```
Ultralytics 8.3.40 🚀 Python-3.11.10 torch-2.5.0+cu124 CUDA:0 (NVIDIA GeForce RTX 2070, 8192MiB)
Setup complete ✅ (8 CPUs, 5.8 GB RAM, 267.6/1006.9 GB disk)
OS Linux-5.15.167.4-microsoft-standard-WSL2-x86_64-with-glibc2.35
Environment Docker
Python 3.11.10
Install git
RAM 5.79 GB
Disk 267.6/1006.9 GB
CPU Intel Core(TM) i7-9700K 3.60GHz
CPU count 8
GPU NVIDIA GeForce RTX 2070, 8192MiB
GPU count 1
CUDA 12.4
numpy ✅ 1.23.5>=1.23.0
numpy ✅ 1.23.5<2.0.0; sys_platform == "darwin"
matplotlib ✅ 3.9.3>=3.3.0
opencv-python ✅ 4.10.0.84>=4.6.0
pillow ✅ 10.2.0>=7.1.2
pyyaml ✅ 6.0.2>=5.3.1
requests ✅ 2.32.3>=2.23.0
scipy ✅ 1.14.1>=1.4.1
torch ✅ 2.5.0+cu124>=1.8.0
torch ✅ 2.5.0+cu124!=2.4.0,>=1.8.0; sys_platform == "win32"
torchvision ✅ 0.20.0+cu124>=0.9.0
tqdm ✅ 4.66.5>=4.64.0
psutil ✅ 6.0.0
py-cpuinfo ✅ 9.0.0
pandas ✅ 2.2.3>=1.1.4
seaborn ✅ 0.13.2>=0.11.0
ultralytics-thop ✅ 2.0.12>=2.0.0
```
### Minimal Reproducible Example
```
docker run -it --rm \
--ipc=host \
--gpus all \
-p 6006:6006 \
ultralytics/ultralytics:8.3.40 \
yolo detect train data=coco8.yaml model=yolo11n.pt
```
### Additional
_No response_
### Are you willing to submit a PR?
- [ ] Yes I'd like to help by submitting a PR! | closed | 2025-01-22T07:07:09Z | 2025-01-22T08:44:41Z | https://github.com/ultralytics/ultralytics/issues/18815 | [
"bug",
"detect",
"devops"
] | hexchip | 6 |
autogluon/autogluon | scikit-learn | 4,991 | [Feature Request] Support Past Covariates in AutoGluon Time Series Models | ## Description
I would like to request support for incorporating past covariates in the training process of time series models within AutoGluon. This feature would enhance the flexibility and predictive power of time series models by allowing them to leverage additional historical information.
- This proposal refers to the time-series module.
## Requested Enhancements:
- Enable `PatchTSTModel` and `DeepAR` to support the inclusion of past covariates during training.
- Allow fine-tuned `Chronos-Bolt` models to accept past covariates for improved forecasting capabilities.
## Motivation:
- Many real-world time series problems require contextual historical information beyond the target variable itself.
- This enhancement would enable more accurate and robust forecasting, especially for datasets with external influencing factors.
If there are any current workarounds or ongoing developments related to this, I would appreciate any insights. Thank you for considering this feature request. I appreciate the efforts of the AutoGluon team and look forward to any discussions about feasibility and potential implementation!
| open | 2025-03-21T08:19:11Z | 2025-03-21T08:25:33Z | https://github.com/autogluon/autogluon/issues/4991 | [
"enhancement",
"module: timeseries"
] | LiPingYen | 0 |
netbox-community/netbox | django | 18,572 | Populate CORS_ORIGIN_WHITELIST documentation | ### Change Type
Addition
### Area
Configuration
### Proposed Changes
The [page](https://demo.netbox.dev/static/docs/configuration/security/) which describes CORS_ORIGIN_WHITELIST doesn’t define the value. | closed | 2025-02-05T11:49:58Z | 2025-02-05T13:11:17Z | https://github.com/netbox-community/netbox/issues/18572 | [
"type: documentation"
] | mr1716 | 1 |
rthalley/dnspython | asyncio | 754 | resolve_address has wrong type | The type definition of resolve_address address is `def resolve_address(self, ipaddr: str, *args: Any, **kwargs: Optional[Dict]):`, but the real method has the signature `def resolve_address(ipaddr, *args, **kwargs):` --> the `self` parameter is wrong.
Can you please have a look? I'm happy to open a PR
| closed | 2022-01-18T16:47:52Z | 2022-01-18T17:18:32Z | https://github.com/rthalley/dnspython/issues/754 | [
"Bug",
"Fixed"
] | kasium | 5 |
AntonOsika/gpt-engineer | python | 547 | Unable to use gpt-engineer with API from a free account with OpenAI | Hi, I am using a MacOS and a free OpenAI account.
After following the readme and setting the API key, when I run the "gpt-engineer first_auto" command, I get the following info:
```openai:error_code=insufficient_quota error_message='You exceeded your current quota, please check your plan and billing details.' error_param=None error_type=insufficient_quota message='OpenAI API error received' stream_error=False```
The execution then fails with the following error:
```openai.error.RateLimitError: You exceeded your current quota, please check your plan and billing details.```
I am new to this, could you please help me understand what is happening? | closed | 2023-07-20T06:26:29Z | 2023-08-16T19:34:44Z | https://github.com/AntonOsika/gpt-engineer/issues/547 | [] | AnkiBhatia | 15 |
JaidedAI/EasyOCR | deep-learning | 717 | Help, a issues! | D:\ProgramData\Anaconda3\envs\pytorch\lib\site-packages\torchvision\io\image.py:13: UserWarning: Failed to load image Python extension: Could not find module 'D:\ProgramData\Anaconda3\envs\pytorch\Lib\site-packages\torchvision\image.pyd' (or one of its dependencies). Try using the full path with constructor syntax.
warn(f"Failed to load image Python extension: {e}")

| open | 2022-05-03T13:02:58Z | 2022-05-03T13:02:58Z | https://github.com/JaidedAI/EasyOCR/issues/717 | [] | cobition | 0 |
yeongpin/cursor-free-vip | automation | 324 | [讨论]: 使用自定义邮箱注册时出错 | ### Issue 检查清单
- [x] 我理解 Issue 是用于反馈和解决问题的,而非吐槽评论区,将尽可能提供更多信息帮助问题解决。
- [x] 我确认自己需要的是提出问题并且讨论问题,而不是 Bug 反馈或需求建议。
- [x] 我已阅读 [Github Issues](https://github.com/yeongpin/cursor-free-vip/issues) 并搜索了现有的 [开放 Issue](https://github.com/yeongpin/cursor-free-vip/issues) 和 [已关闭 Issue](https://github.com/yeongpin/cursor-free-vip/issues?q=is%3Aissue%20state%3Aclosed%20),没有找到类似的问题。
### 平台
Windows x64
### 版本
1.7.12
### 您的问题

### 补充信息
```shell
使用自定义邮箱注册时出错,提示如图。
```
### 优先级
中 (希望尽快得到答复) | closed | 2025-03-20T01:59:08Z | 2025-03-20T03:46:20Z | https://github.com/yeongpin/cursor-free-vip/issues/324 | [
"question"
] | mygithub424525 | 1 |
davidteather/TikTok-Api | api | 841 | [INSTALLATION] - requests.exceptions.ConnectionError | Describe the error
I need to use VPN to login www.tiktok.com, but when i finished coding , displayed the below errors:
requests.exceptions.ConnectionError: HTTPSConnectionPool(host='www.tiktok.com', port=443): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPSConnection object at 0x0000023FE34B5640>: Failed to establish a new connection: [WinError 10060] A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond'))
I thought maybe was VPN problem, but i am not sure, have anyone can help me to solve this issue, thank you very much.
**The buggy code**
```
from TikTokApi import TikTokApi
api = TikTokApi(custom_verify_fp="******")
for video in api.trending.videos(count=50):
print(video.as_dict)
```
**Error Trace (if any)**
Put the error trace below if there's any error thrown.
```
Traceback (most recent call last):
File "D:\software\Python\lib\site-packages\urllib3\connection.py", line 169, in _new_conn
conn = connection.create_connection(
File "D:\software\Python\lib\site-packages\urllib3\util\connection.py", line 96, in create_connection
raise err
File "D:\software\Python\lib\site-packages\urllib3\util\connection.py", line 86, in create_connection
sock.connect(sa)
TimeoutError: [WinError 10060] A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "D:\software\Python\lib\site-packages\urllib3\connectionpool.py", line 699, in urlopen
httplib_response = self._make_request(
File "D:\software\Python\lib\site-packages\urllib3\connectionpool.py", line 382, in _make_request
self._validate_conn(conn)
File "D:\software\Python\lib\site-packages\urllib3\connectionpool.py", line 1010, in _validate_conn
conn.connect()
File "D:\software\Python\lib\site-packages\urllib3\connection.py", line 353, in connect
conn = self._new_conn()
File "D:\software\Python\lib\site-packages\urllib3\connection.py", line 181, in _new_conn
raise NewConnectionError(
urllib3.exceptions.NewConnectionError: <urllib3.connection.HTTPSConnection object at 0x0000023FE34B5640>: Failed to establish a new connection: [WinError 10060] A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "D:\software\Python\lib\site-packages\requests\adapters.py", line 439, in send
resp = conn.urlopen(
File "D:\software\Python\lib\site-packages\urllib3\connectionpool.py", line 755, in urlopen
retries = retries.increment(
File "D:\software\Python\lib\site-packages\urllib3\util\retry.py", line 574, in increment
raise MaxRetryError(_pool, url, error or ResponseError(cause))
urllib3.exceptions.MaxRetryError: HTTPSConnectionPool(host='www.tiktok.com', port=443): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPSConnection object at 0x0000023FE34B5640>: Failed to establish a new connection: [WinError 10060] A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond'))
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "D:\software\Python\Python projects\Test1\testing32.py", line 9, in <module>
for video in api.trending.videos(count=50):
File "D:\software\Python\lib\site-packages\TikTokApi\api\trending.py", line 35, in videos
spawn = requests.head(
File "D:\software\Python\lib\site-packages\requests\api.py", line 104, in head
return request('head', url, **kwargs)
File "D:\software\Python\lib\site-packages\requests\api.py", line 61, in request
return session.request(method=method, url=url, **kwargs)
File "D:\software\Python\lib\site-packages\requests\sessions.py", line 542, in request
resp = self.send(prep, **send_kwargs)
File "D:\software\Python\lib\site-packages\requests\sessions.py", line 655, in send
r = adapter.send(request, **kwargs)
File "D:\software\Python\lib\site-packages\requests\adapters.py", line 516, in send
raise ConnectionError(e, request=request)
requests.exceptions.ConnectionError: HTTPSConnectionPool(host='www.tiktok.com', port=443): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPSConnection object at 0x0000023FE34B5640>: Failed to establish a new connection: [WinError 10060] A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond'))
Process finished with exit code 1
```
**Desktop (please complete the following information):**
- OS: [e.g. Windows [10]
- TikTokApi Version [5.0]
- Python Version [3.95]
| closed | 2022-02-26T15:45:14Z | 2023-08-08T22:20:22Z | https://github.com/davidteather/TikTok-Api/issues/841 | [
"installation_help"
] | Jerrywangmax | 2 |
ndleah/python-mini-project | data-visualization | 262 | Modularization for Caterpillar_Game | # Description
With this implementation the actual game can be more easier to use it, diving the part of the code into two main files, the game mechanism and the user interface, doing this the code will be more flexibility and reusability, also with this we can have a better testing for the function of the game.
## Type of issue
- [x] Feature (New Script)
- [ ] Bug
- [ ] Documentation
## Checklist:
- [x] I have read the project guidelines.
- [x] I have checked previous issues to avoid duplicates.
- [x] This issue will be meaningful for the project.
| open | 2024-05-15T07:41:29Z | 2024-06-11T08:35:39Z | https://github.com/ndleah/python-mini-project/issues/262 | [] | Gabriela20103967 | 0 |
ultralytics/ultralytics | pytorch | 18,939 | ONNX inference with GPU not working on NVIDIA ORIN | ### Search before asking
- [x] I have searched the Ultralytics YOLO [issues](https://github.com/ultralytics/ultralytics/issues) and found no similar bug report.
### Ultralytics YOLO Component
_No response_
### Bug
Dear ultralytics team and community,
I'm trying to run very simple inference on one image of yolov8n.onnx (yolov8n.pt exported to onnx format) and It's not working with the NVIDIA Jetson GPU. I need to run it on GPU because it has to be real time.
Thank you very much in advance for your help.
Best regards,
Xuban
### Environment
Ultralytics 8.3.28 🚀 Python-3.10.12 torch-2.1.0 CUDA:0 (Orin, 30697MiB)
Setup complete ✅ (12 CPUs, 30.0 GB RAM, 194.7/455.9 GB disk)
OS Linux-5.15.122-tegra-aarch64-with-glibc2.35
Environment Linux
Python 3.10.12
Install pip
RAM 29.98 GB
Disk 194.7/455.9 GB
CPU Cortex-A78AE
CPU count 12
GPU Orin, 30697MiB
GPU count 1
CUDA 12.2
numpy ✅ 1.23.5>=1.23.0
matplotlib ✅ 3.5.1>=3.3.0
opencv-python ✅ 4.10.0.84>=4.6.0
pillow ✅ 11.0.0>=7.1.2
pyyaml ✅ 5.4.1>=5.3.1
requests ✅ 2.32.3>=2.23.0
scipy ✅ 1.12.0>=1.4.1
torch ✅ 2.1.0>=1.8.0
torchvision ✅ 0.16.0+fbb4cc5>=0.9.0
tqdm ✅ 4.67.0>=4.64.0
psutil ✅ 5.9.0
py-cpuinfo ✅ 9.0.0
pandas ✅ 1.3.5>=1.1.4
seaborn ✅ 0.13.1>=0.11.0
ultralytics-thop ✅ 2.0.10>=2.0.0
numpy ✅ 1.23.5<2.0.0; sys_platform == "darwin"
torch ✅ 2.1.0!=2.4.0,>=1.8.0; sys_platform == "win32"
### Minimal Reproducible Example
Here is the python code:
`from ultralytics import YOLO
def run_inference(onnx_model_path, image_path, output_image_path):
model = YOLO(onnx_model_path)
results = model(image_path)
results[0].save(output_image_path)
print(f"Inference completed. Output saved to {output_image_path}")
if __name__ == "__main__":
run_inference("yolov8n.onnx", "input.jpg", "output.jpg")`
### Additional
`WARNING ⚠️ Unable to automatically guess model task, assuming 'task=detect'. Explicitly define task for your model, i.e. 'task=detect', 'segment', 'classify','pose' or 'obb'.
Loading yolov8n.onnx for ONNX Runtime inference...
requirements: Ultralytics requirement ['onnxruntime-gpu'] not found, attempting AutoUpdate...
ERROR: Could not find a version that satisfies the requirement onnxruntime-gpu (from versions: none)
[notice] A new release of pip is available: 24.3.1 -> 25.0
[notice] To update, run: python3 -m pip install --upgrade pip
ERROR: No matching distribution found for onnxruntime-gpu
Retry 1/2 failed: Command 'pip install --no-cache-dir "onnxruntime-gpu" ' returned non-zero exit status 1.
ERROR: Could not find a version that satisfies the requirement onnxruntime-gpu (from versions: none)
[notice] A new release of pip is available: 24.3.1 -> 25.0
[notice] To update, run: python3 -m pip install --upgrade pip
ERROR: No matching distribution found for onnxruntime-gpu
Retry 2/2 failed: Command 'pip install --no-cache-dir "onnxruntime-gpu" ' returned non-zero exit status 1.
requirements: ❌ Command 'pip install --no-cache-dir "onnxruntime-gpu" ' returned non-zero exit status 1.
WARNING ⚠️ Failed to start ONNX Runtime session with CUDA. Falling back to CPU...
Preferring ONNX Runtime AzureExecutionProvider
image 1/1 /home/ikerlan/Documents/xubanceccon/paper/input.jpg: 640x640 4 persons, 2 ties, 131.7ms
Speed: 9.3ms preprocess, 131.7ms inference, 3.7ms postprocess per image at shape (1, 3, 640, 640)
Inference completed. Output saved to output.jpg`
### Are you willing to submit a PR?
- [ ] Yes I'd like to help by submitting a PR! | open | 2025-01-29T15:54:15Z | 2025-02-03T09:30:54Z | https://github.com/ultralytics/ultralytics/issues/18939 | [
"bug",
"detect",
"embedded",
"exports"
] | xceccon | 5 |
eamigo86/graphene-django-extras | graphql | 177 | Tried installing graphene-django-extras, but get errors about incompatibility | Hi, just tried to installed your library, but then I get this error :
ERROR: pip's dependency resolver does not currently take into account all the packages that are installed. This behaviour is the source of the following dependency conflicts.
django-graphql-jwt 0.3.0 requires graphql-core<3,>=2.1, but you have graphql-core 3.1.5 which is incompatible.
Is it incompatible to my graphene version, is that the issue?
What should I do in order to use the library? | open | 2021-07-11T19:19:05Z | 2021-08-26T06:48:17Z | https://github.com/eamigo86/graphene-django-extras/issues/177 | [] | Instrumedley | 3 |
gradio-app/gradio | data-science | 10,141 | Python version is not 3.10+ in devcontainer | - [x] I have searched to see if a similar issue already exists.
**Is your feature request related to a problem? Please describe.**
Since the CONTRIBUTING guide (https://github.com/gradio-app/gradio/blob/main/CONTRIBUTING.md) says that Python version 3.10+ is highly recommended, it would be nice to have the devcontainer configured for that. Currently, it uses 3.9 on Debian 11 (see below)
https://github.com/gradio-app/gradio/blob/9a6ce6f6b089d94c06da0b8620f28967f39f8383/.devcontainer/devcontainer.json#L4
**Describe the solution you'd like**
I expected that changing L4 of `devcontainer.json` to `"image": "mcr.microsoft.com/devcontainers/python:3.10",` would take care of the problem but it does not have any effect. After doing that, I still have `/usr/bin/python3` aliased to `/usr/bin/python3.9` and there is no 3.10.
**Additional context**
Additional improvements, perhaps?
- Running `apt-get update && apt-get upgrade` as part of the `postCreateCommand` as suggested by https://hub.docker.com/r/microsoft/devcontainers-python.
| closed | 2024-12-06T01:39:23Z | 2024-12-08T04:52:58Z | https://github.com/gradio-app/gradio/issues/10141 | [
"bug"
] | anirbanbasu | 15 |
databricks/spark-sklearn | scikit-learn | 32 | Link pyspark docs in generated docs | Need to configure intersphinx
| closed | 2016-06-28T00:01:38Z | 2018-12-08T20:01:33Z | https://github.com/databricks/spark-sklearn/issues/32 | [] | vlad17 | 1 |
piskvorky/gensim | data-science | 3,376 | word2vec doesn't scale linearly with multi-cpu configuration ? | #### Problem description
I've tried to use script from:
https://github.com/RaRe-Technologies/gensim/releases/3.6.0
with varying number of cores (num_cores) and obtained following times: 8 -> 26 sec, 16 -> 17 sec, 24 -> 14.4 sec, 32 -> 15.9 sec, 48 -> 16 sec. So it doesn't scale linearly with number of cores and peak seems to be at 24 cores.
My machine reports 48 cores by cpu_count(), by lscpu: CPUs: 48, Threads per core: 2, Cores per socket:12, Socket: 2, Numa nodes: 2, Model name: Intel Xeon E5-2650 v4 2.2 Ghz. Note, the same behaviour occurs for Doc2Vec and FastText.
Is it possible that only one socket is used, or I miss something?
#### Steps/code/corpus to reproduce
```
import gensim.downloader as api
from multiprocessing import cpu_count
from gensim.utils import save_as_line_sentence
from gensim.test.utils import get_tmpfile
from gensim.models import Word2Vec, Doc2Vec, FastText
from linetimer import CodeTimer #pip install linetimer
# Convert any corpus to the needed format: 1 document per line, words delimited by " "
corpus = api.load("text8")
corpus_fname = get_tmpfile("text8-file-sentence.txt")
save_as_line_sentence(corpus, corpus_fname)
# Choose num of cores that you want to use (let's use all, models scale linearly now!)
num_cores = 8 # 16, 24, 32, 48 cpu_count()
# Train models using all cores
with CodeTimer(unit="s"):
w2v_model = Word2Vec(corpus_file=corpus_fname, workers=num_cores)
#d2v_model = Doc2Vec(corpus_file=corpus_fname, workers=num_cores)
#ft_model = FastText(corpus_file=corpus_fname, workers=num_cores)
```
#### Versions
Linux-4.15...generic_x86_64_with_debian_buster_sid
64
numpy: 1.21.4
scipy: 1.7.3
gensim : 4.2.0
FAST_VERSION: the same behavior with 0 and 1 | open | 2022-08-09T06:20:56Z | 2022-08-23T05:21:50Z | https://github.com/piskvorky/gensim/issues/3376 | [] | mglowacki100 | 7 |
pallets/flask | flask | 5,408 | send_file doesn't work with objects | I had a file in an object of type io.BytesIO(), but when I put this object in a send_file made me the error that the parameter is incorrect.
In the version 2 this work well but in the version 3.0.2 doesn't.
Environment:
- Python version: 3.10.12
- Flask version: 3.0.2
| closed | 2024-02-08T16:41:20Z | 2024-02-23T00:05:36Z | https://github.com/pallets/flask/issues/5408 | [] | AxelGarciaTello | 1 |
waditu/tushare | pandas | 1,494 | LPR利率数据缺少 | shibor_lpr接口获取的LPR数据只有截止2019年11月的
ID:265422 | open | 2021-01-17T12:23:31Z | 2021-01-17T12:23:31Z | https://github.com/waditu/tushare/issues/1494 | [] | EthanHsiung | 0 |
gunthercox/ChatterBot | machine-learning | 1,893 | ListTrainer Import issue | I have already installed chatterbot and chatterbot corpus twice. Initially it was showing Import error for ChatBot which somehow resolved, but it started showing Import error for ListTrainer. Can someone please help me out here.
I am using Linux
This is the error being displayed
Traceback (most recent call last):
File "/home/anant/Documents/ss.py", line 2, in <module>
from chatterbot import ListTrainer
ImportError: cannot import name 'ListTrainer'
I have already tried ListTrainers fix, didn't work
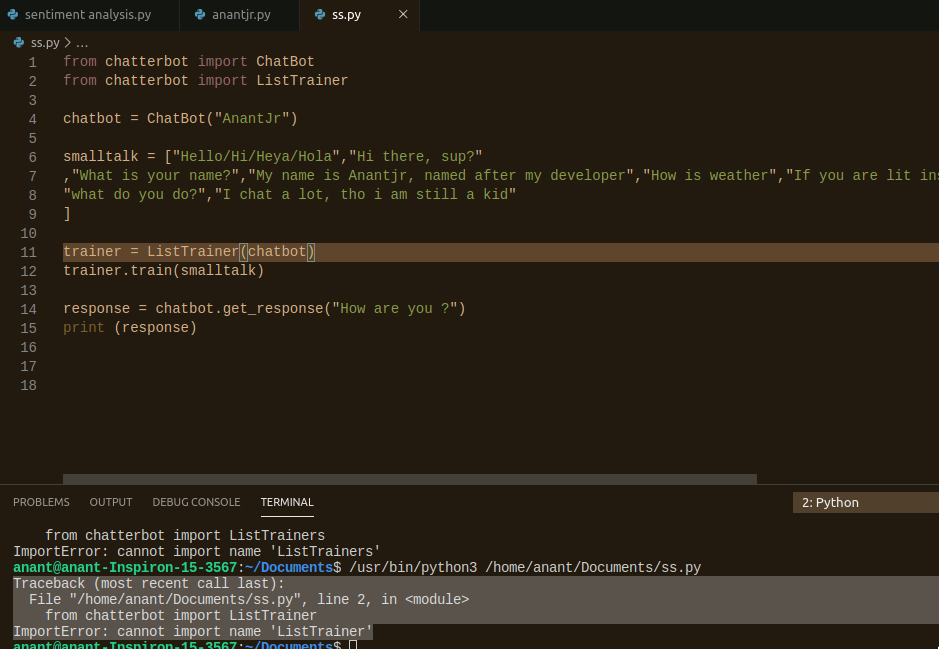
| closed | 2020-01-08T04:50:26Z | 2025-03-24T12:12:47Z | https://github.com/gunthercox/ChatterBot/issues/1893 | [] | anantshahi | 1 |
httpie/cli | api | 681 | Fix Travis / tests | https://travis-ci.org/jakubroztocil/httpie/jobs/385658131#L598-L616 | closed | 2018-05-30T12:15:43Z | 2018-06-09T10:13:59Z | https://github.com/httpie/cli/issues/681 | [
"help wanted"
] | jkbrzt | 3 |
ResidentMario/geoplot | matplotlib | 237 | webmap() + polyplot() does not seem to work | Hi there,
first of all congrats on a great product. I'm trying to get `webmap()` to work in conjunction with `polyplot()` but can't seem to get the polys to show. Without `webmap()`, this works no problem:
```
ax = gplt.polyplot(gdf1, extent=extent, projection=gcrs.Mercator(), **kwargs)
gplt.polyplot(gdf2, ax=ax, **kwargs)
```
If I try this with `webmap()`, however, the basemap shows (with the right zoom level and extent), but the polys don't.
```
ax = gplt.webmap(gdf1, extent=extent, projection=gcrs.WebMercator(), **kwargs)
gplt.polyplot(gdf2, ax=ax, **kwargs)
```
All examples in the documentation use `webmap()` in conjunction with `pointplot()` only. Is that the issue? All my GDFs are in lon lat as required by geoplot, by the way.
Cheers! | closed | 2021-06-28T13:34:17Z | 2021-07-03T21:50:29Z | https://github.com/ResidentMario/geoplot/issues/237 | [] | gregorhd | 3 |
iperov/DeepFaceLab | machine-learning | 850 | Error when I click on graph icon on analysis page. | ### Problem
It is solved. | closed | 2020-08-04T21:12:35Z | 2020-08-04T21:48:39Z | https://github.com/iperov/DeepFaceLab/issues/850 | [] | Xtendera | 0 |
waditu/tushare | pandas | 1,271 | pro.dividend读出的002050.SZ股票的分红出现重复,还有其他的股票 没有都列出 [tushare id=13936436049] | ts_code end_date ann_date div_proc stk_div stk_bo_rate stk_co_rate cash_div cash_div_tax record_date ex_date pay_date div_listdate imp_ann_date
0 002050.SZ 20190630 20190830 预案 0 0 0
1 002050.SZ 20181231 20190403 实施 0.3 0.3 0.25 0.25 20190509 20190510 20190510 20190510 20190430
2 002050.SZ 20181231 20190403 实施 0.3 0.3 0.25 0.25 20190509 20190510 20190510 20190510 20190430
| open | 2020-01-31T03:48:37Z | 2020-01-31T03:49:31Z | https://github.com/waditu/tushare/issues/1271 | [] | smqhrb | 0 |
django-cms/django-cms | django | 7,985 | [BUG] cms middleware defeats ASGI adaptation | ## Description
Django async support adapts synchronous middleware to an asynchronous middleware stack automagically. It does that by assuming it's synchronous and testing for async. Subclassing MiddlewareMixin makes that test positive. However, some cms middleware then overrides \_\_call\_\_, which django does not adapt, causing it to fail.
Of the five middlewares provided, LanguageCookieMiddleware and CurrentUserMiddleware both suffer from this.
## Steps to reproduce
Run django-cms on an ASGI server (uvicorn in my case).
## Expected behaviour
Django async support allows non-async django-cms to run alongside async-aware apps.
## Actual behaviour
Failure is immediate if LanguageCookieMiddleware is installed:
```
AttributeError at /
'coroutine' object has no attribute 'set_cookie'
Request Method: GET
Request URL: https://test-server.example.com/
Django Version: 4.2.15
Exception Type: AttributeError
Exception Value:
'coroutine' object has no attribute 'set_cookie'
Exception Location: /usr/local/share/sites/csweb/lib/python3.11/site-packages/cms/middleware/language.py, line 54, in __call__
Raised during: cms.views.details
Python Executable: /usr/local/share/sites/csweb/bin/python
Python Version: 3.11.2
```
## Additional information (CMS/Python/Django versions)
python 3.11.2
django 4.2.14
django-cms 3.11.6
## Do you want to help fix this issue?
* [ X] Yes, I want to help fix this issue and I will join the channel #pr-reviews on [the Discord Server](https://discord-pr-review-channel.django-cms.org) to confirm with the community that a PR is welcome.
* [ ] No, I only want to report the issue.
| closed | 2024-09-04T16:21:21Z | 2024-10-06T16:27:11Z | https://github.com/django-cms/django-cms/issues/7985 | [
"3.11"
] | jbazik | 4 |
wkentaro/labelme | computer-vision | 995 | multiple labels output in different colors each time json file is converted to png | As you can see that after labeling the 3 classes they come out in different colors after extracting from the json file. i want each of them to be of distinct colors.
Can you help?

| closed | 2022-02-26T06:39:53Z | 2022-03-06T09:54:47Z | https://github.com/wkentaro/labelme/issues/995 | [] | rexxar0105 | 1 |
PeterL1n/RobustVideoMatting | computer-vision | 62 | Questions on reproducing the results | Hi, Thanks for sharing this amazing work on video matting!
I'm trying to reproduce the numbers in Table 1 of the paper and have some questions here:
1. In table 1, all the results are under training stage 1,2,3 and 4, right? I trained the model for stage 1,2,3 and got the results 12.16 / 3.08 (MAD/MSE) on VM while in the paper it is 6.08/1.47 (MAD/MSE) on VM.
2. How important is the 8k image backgrounds for reproducing the numbers in Table 1? I used the 200 image backgrounds in test set that you released for training stage 1,2,3.
3. The overlap about training / test set on video backgrounds. In paper, you mentioned 3118 clips (while in dvm_background_train_clips.txt there are 3117 lines) are selected for training, while in dvm_background_test_clips.txt there are some overlap clips with training set (like 0245/0246), does it mean that we need to manually remove them during training? By the way, in generate_videomatte_with_background_video.py, 0245/0246 are also selected for compositing test set.
Could you help elaborate on them? Thanks. | closed | 2021-10-01T22:34:48Z | 2021-10-01T23:37:05Z | https://github.com/PeterL1n/RobustVideoMatting/issues/62 | [] | chrisjuniorli | 2 |
jumpserver/jumpserver | django | 15,112 | [Bug] 通过jumpserver登录dell idrac后无法打开服务器里的系统 | ### Product Version
4.8.0
### Product Edition
- [x] Community Edition
- [ ] Enterprise Edition
- [ ] Enterprise Trial Edition
### Installation Method
- [x] Online Installation (One-click command installation)
- [ ] Offline Package Installation
- [ ] All-in-One
- [ ] 1Panel
- [ ] Kubernetes
- [ ] Source Code
### Environment Information
ubuntu22.04 jumpserver4.8.0
### 🐛 Bug Description
通过jumpserver登录dell idrac后无法打开服务器里的系统,不通过jumpserve登录dell idrac后可以打开服务器里的系统
### Recurrence Steps
1、用jumpserver web登录dell idrac管理页面,然后点击登录系统


2、不通过jumpserver登录dell idrac管理界面,登录系统正常

### Expected Behavior
_No response_
### Additional Information
_No response_
### Attempted Solutions
_No response_ | closed | 2025-03-24T08:49:10Z | 2025-03-24T09:02:33Z | https://github.com/jumpserver/jumpserver/issues/15112 | [
"🐛 Bug"
] | ryswork1993 | 1 |
PaddlePaddle/ERNIE | nlp | 361 | ernie是基于字训练还是基于词训练的啊 | closed | 2019-11-03T06:50:14Z | 2020-05-28T09:53:04Z | https://github.com/PaddlePaddle/ERNIE/issues/361 | [
"wontfix"
] | wxlduter | 5 | |
pykaldi/pykaldi | numpy | 73 | 【Question】Get cuArray from pytorch or cupy | Can I convert `kaldi::cuArray` <-> any python CUDA-library such as `pytorch` or `cupy` directly without transferring to CPU? I'm interesting to glue `chain` with such framework. | closed | 2019-01-09T02:12:08Z | 2019-01-11T06:46:27Z | https://github.com/pykaldi/pykaldi/issues/73 | [] | kamo-naoyuki | 2 |
miguelgrinberg/Flask-SocketIO | flask | 947 | How many namespaces? | In my app i have a live table with 2 parts:
the live table and also the system overload variable. Lets say i want to update the live table using the user room and the system overload making a broadcast to all users in namespace. Is it bad to have a namespace for every client ? lets say i plan to scale up to 200 300 clients.
Also, given i want a user admin with his table to get all user tables updates and append in one table, can i get all the emits from the room of a single name space ? or would i have to join to every user room to that admin ? | closed | 2019-04-14T07:33:05Z | 2019-04-21T21:38:40Z | https://github.com/miguelgrinberg/Flask-SocketIO/issues/947 | [
"question"
] | valentin-ballester | 7 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.