repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
matplotlib/mplfinance | matplotlib | 167 | Changing spines color | I'm trying to change the color of the frame around the chart, but i'm having an hard time finding the right `rc` parameter. Until now, i managed to remove them with `'axes.spines.bottom':False` but not to edit the appearance of the spines, how can i do that? Thanks in advance! | closed | 2020-06-11T12:33:44Z | 2021-08-26T21:50:11Z | https://github.com/matplotlib/mplfinance/issues/167 | [
"question"
] | Sile25 | 4 |
s3rius/FastAPI-template | asyncio | 191 | Loguru startup error | After initializing blank project with loguru as logger `poetry run python -m project_name` it gives an error:
```shell
Traceback (most recent call last):
File "<frozen runpy>", line 198, in _run_module_as_main
File "<frozen runpy>", line 88, in _run_code
File "C:\Users\User\Documents\Projects\project_name\project_name\__main__.py", line 4, in <module>
import uvicorn
File "C:\Users\User\AppData\Local\pypoetry\Cache\virtualenvs\project-name-uwC7DCiD-py3.11\Lib\site-packages\uvicorn\__init__.py", line 1, in <module>
from uvicorn.config import Config
File "C:\Users\User\AppData\Local\pypoetry\Cache\virtualenvs\project-name-uwC7DCiD-py3.11\Lib\site-packages\uvicorn\config.py", line 1, in <module>
import asyncio
File "C:\Users\User\AppData\Local\Programs\Python\Python311\Lib\asyncio\__init__.py", line 8, in <module>
from .base_events import *
File "C:\Users\User\AppData\Local\Programs\Python\Python311\Lib\asyncio\base_events.py", line 18, in <module>
import concurrent.futures
File "C:\Users\User\AppData\Local\Programs\Python\Python311\Lib\concurrent\futures\__init__.py", line 8, in <module>
from concurrent.futures._base import (FIRST_COMPLETED,
File "C:\Users\User\AppData\Local\Programs\Python\Python311\Lib\concurrent\futures\_base.py", line 7, in <module>
import logging
File "C:\Users\User\Documents\Projects\project_name\project_name\logging.py", line 5, in <module>
from loguru import logger
File "C:\Users\User\AppData\Local\pypoetry\Cache\virtualenvs\project-name-uwC7DCiD-py3.11\Lib\site-packages\loguru\__init__.py", line 10, in <module>
from ._logger import Core as _Core
File "C:\Users\User\AppData\Local\pypoetry\Cache\virtualenvs\project-name-uwC7DCiD-py3.11\Lib\site-packages\loguru\_logger.py", line 99, in <module>
from . import _asyncio_loop, _colorama, _defaults, _filters
File "C:\Users\User\AppData\Local\pypoetry\Cache\virtualenvs\project-name-uwC7DCiD-py3.11\Lib\site-packages\loguru\_asyncio_loop.py", line 27, in <module>
get_task_loop, get_running_loop = load_loop_functions()
^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\User\AppData\Local\pypoetry\Cache\virtualenvs\project-name-uwC7DCiD-py3.11\Lib\site-packages\loguru\_asyncio_loop.py", line 11, in load_loop_functions
get_running_loop = asyncio.get_running_loop
^^^^^^^^^^^^^^^^^^^^^^^^
AttributeError: partially initialized module 'asyncio' has no attribute 'get_running_loop' (most likely due to a circular import)
```
After renaming file `project_name/logging.py` to `project_name/log.py` it works | open | 2023-10-02T12:35:53Z | 2024-07-12T08:16:43Z | https://github.com/s3rius/FastAPI-template/issues/191 | [] | RoyalGoose | 13 |
scikit-image/scikit-image | computer-vision | 7,364 | Feature smaller, focused gallery examples with a lower priority | ### Description:
> Maybe we can move examples whose focus is on a single function to the end of the gallery. I imagine they are mostly reached from the function itself rather than browsing the gallery thumbnails.
as suggested in this [comment on our SP forum](https://discuss.scientific-python.org/t/featuring-artistic-examples-in-the-gallery/904/9).
With the same idea, we could try to select gallery examples that we want to feature more prominently because the have a broad appeal. E.g. according to https://views.scientific-python.org/scikit-image.org (not public) our [regionprops example](https://scikit-image.org/docs/stable/auto_examples/segmentation/plot_regionprops.html) is our most visited one. | open | 2024-03-30T12:03:54Z | 2024-09-27T02:39:50Z | https://github.com/scikit-image/scikit-image/issues/7364 | [
":page_facing_up: type: Documentation",
":pray: Feature request",
":sleeping: Dormant"
] | lagru | 1 |
apachecn/ailearning | python | 648 | 第四章(朴素贝叶斯)中 rss 订阅失效 | ### 问题描述
在第四章-朴素贝叶斯算法的第三个小实验中,使用了 feedparser 模块来解析两个 rss 源以获取文本数据。验证发现连接已经失效,所获取的文本列表为空。
点击网站连接,会看到如下内容
> Your request has been blocked.
>
> If you have questions, please [contact us](https://www.craigslist.org/contact?step=form&reqType=help_blocks&blockID=500832).
### 问题资源地址
[第四章-朴素贝叶斯算法](https://github.com/apachecn/ailearning/blob/master/docs/ml/4.md)
### 问题位置截图

### 自测代码
```python
def localWords(feed1, feed0):
docList = []
classList = []
fullText = []
minLen = min(len(feed1["entries"]), len(feed0["entries"]))
# 1. 文本获取与统计
for i in range(minLen):
# 类别 1:每次访问一条 RSS 源
wordList = textParse(feed1["entries"][i]["summary"])
docList.append(wordList)
fullText.extend(wordList)
classList.append(1)
# 类别 0:每次访问一条 RSS 源
wordList = textParse(feed0["entries"][i]["summary"])
docList.append(wordList)
fullText.extend(wordList)
classList.append(0)
vocabList = bayes.createVocabList(docList)
top30Words = calMostFreq(vocabList, fullText)
print(f"打印获取的文本:\n{docList}")
print(f"打印单词列表:\n{vocabList}")
if __name__ == "__main__":
import feedparser as fp # type: ignore
ny = fp.parse('http://newyork.craigslist.org/stp/index.rss')
sf = fp.parse('http://sfbay.craigslist.org/stp/index.rss')
localWords(ny, sf)
```
### 输出结果
```powershell
(py38) D:\PROJECT\ml>C:/tools/Anaconda3/envs/py38/python.exe d:/PROJECT/ml/4_bayes/rss.py
打印获取的文本:
[]
打印单词列表:
[]
```
### 建议
1. 更换新的可用源
2. 或者仅展示实验结果,让大家自己找源来测试算法
| closed | 2024-01-03T01:45:17Z | 2024-01-03T02:15:31Z | https://github.com/apachecn/ailearning/issues/648 | [] | AIkikaze | 2 |
flairNLP/flair | nlp | 2,679 | Multi-label or overlapping annotations predictions | Is it possible to train a Flair NER-sequence-tagger on overlapping annotations?
I found that you introduced multi-label predictions in 2021 but I am not sure whether that fits my problem. Unfortunately, I didn't find any documentation pointing to that use case.
What I'm thinking of is to train a tagger to predict multiple labels for certain tokens, like:
`Span [1,2]: "George Washington" [− Labels: PER (0.9968), PRES (0.9734)]`
`Span [5]: "Washington" [− Labels: LOC (0.9994)]`
Is that possible with a single Flair without the use of multiple taggers? | closed | 2022-03-16T14:49:47Z | 2023-05-25T13:26:01Z | https://github.com/flairNLP/flair/issues/2679 | [
"question"
] | agademic | 5 |
assafelovic/gpt-researcher | automation | 266 | SyntaxError: invalid syntax when running uvicorn main:app --reload | INFO: Uvicorn running on http://127.0.0.1:8000 (Press CTRL+C to quit)
INFO: Started reloader process [62124] using StatReload
Process SpawnProcess-1:
Traceback (most recent call last):
File "c:\users\xx\appdata\local\programs\python\python37\lib\multiprocessing\process.py", line 297, in _bootstrap
self.run()
File "c:\users\xx\appdata\local\programs\python\python37\lib\multiprocessing\process.py", line 99, in run
self._target(*self._args, **self._kwargs)
File "c:\users\xx\appdata\local\programs\python\python37\lib\site-packages\uvicorn\_subprocess.py", line 76, in subprocess_started
target(sockets=sockets)
File "c:\users\xx\appdata\local\programs\python\python37\lib\site-packages\uvicorn\server.py", line 59, in run
return asyncio.run(self.serve(sockets=sockets))
File "c:\users\xx\appdata\local\programs\python\python37\lib\asyncio\runners.py", line 43, in run
return loop.run_until_complete(main)
File "c:\users\xx\appdata\local\programs\python\python37\lib\asyncio\base_events.py", line 587, in run_until_complete
return future.result()
File "c:\users\xx\appdata\local\programs\python\python37\lib\site-packages\uvicorn\server.py", line 66, in serve
config.load()
File "c:\users\xx\appdata\local\programs\python\python37\lib\site-packages\uvicorn\config.py", line 471, in load
self.loaded_app = import_from_string(self.app)
File "c:\users\xx\appdata\local\programs\python\python37\lib\site-packages\uvicorn\importer.py", line 21, in import_from_string
module = importlib.import_module(module_str)
File "c:\users\xx\appdata\local\programs\python\python37\lib\importlib\__init__.py", line 127, in import_module
return _bootstrap._gcd_import(name[level:], package, level)
File "<frozen importlib._bootstrap>", line 1006, in _gcd_import
File "<frozen importlib._bootstrap>", line 983, in _find_and_load
File "<frozen importlib._bootstrap>", line 967, in _find_and_load_unlocked
File "<frozen importlib._bootstrap>", line 677, in _load_unlocked
File "<frozen importlib._bootstrap_external>", line 728, in exec_module
File "<frozen importlib._bootstrap>", line 219, in _call_with_frames_removed
File "C:\Users\xx\Desktop\gpt-researcher-master\main.py", line 1, in <module>
from backend.server import app
File "C:\Users\xx\Desktop\gpt-researcher-master\backend\server.py", line 7, in <module>
from gpt_researcher.utils.websocket_manager import WebSocketManager
File "C:\Users\xx\Desktop\gpt-researcher-master\gpt_researcher\__init__.py", line 1, in <module>
from .master import GPTResearcher
File "C:\Users\x\Desktop\gpt-researcher-master\gpt_researcher\master\__init__.py", line 1, in <module>
from .agent import GPTResearcher
File "C:\Users\xx\Desktop\gpt-researcher-master\gpt_researcher\master\agent.py", line 3, in <module>
from gpt_researcher.master.functions import *
File "C:\Users\xx\Desktop\gpt-researcher-master\gpt_researcher\master\functions.py", line 18
match retriever:
^
SyntaxError: invalid syntax
| closed | 2023-11-23T16:21:26Z | 2024-06-12T06:24:29Z | https://github.com/assafelovic/gpt-researcher/issues/266 | [] | glejdis | 3 |
gradio-app/gradio | deep-learning | 10,281 | Dragging in an image a second time will not replace the original image, it will open in a new tab | ### Describe the bug
Dragging in an image a second time will not replace the original image, it will open in a new tab
### Have you searched existing issues? 🔎
- [X] I have searched and found no existing issues
### Reproduction
```python
import gradio as gr
with gr.Blocks() as demo:
with gr.Row():
input_image = gr.Image(
label="输入图像",
type="pil",
height=600,
width=400,
interactive=True
)
if __name__ == "__main__":
demo.launch(
server_name="0.0.0.0",
server_port=37865
)
```
### Screenshot
_No response_
### Logs
_No response_
### System Info
```shell
gradio 5.6.0
gradio_client 1.4.3
```
### Severity
I can work around it | open | 2025-01-03T02:42:32Z | 2025-01-05T06:32:07Z | https://github.com/gradio-app/gradio/issues/10281 | [
"bug"
] | Dazidingo | 2 |
streamlit/streamlit | streamlit | 10,353 | Makes @st.cache_resource compatible with Cython | ### Checklist
- [x] I have searched the [existing issues](https://github.com/streamlit/streamlit/issues) for similar feature requests.
- [x] I added a descriptive title and summary to this issue.
### Summary
For a user program that uses `@st.cache_resource`, currently it is not possible to use Cython to compile the user program.
As in `cache_utils`, streamlit uses `inspect.getsource(func)` to attempt to read the source, Cython compiled code will cause an TypeError exception but this is not handled by the fallback to bytecode.
### Why?
_No response_
### How?
Instead of handling `OSError`, fallback to bytecode upon any `Exception` would fix the issue.
### Additional Context
```
File "<redacted>/site-packages/streamlit/runtime/caching/cache_resource_api.py", line 238, in __call__
return self._decorator(
File "<redacted>/site-packages/streamlit/runtime/metrics_util.py", line 409, in wrapped_func
result = non_optional_func(*args, **kwargs)
File "<redacted>/site-packages/streamlit/runtime/caching/cache_resource_api.py", line 431, in _decorator
return make_cached_func_wrapper(
File "<redacted>/site-packages/streamlit/runtime/caching/cache_utils.py", line 161, in make_cached_func_wrapper
cached_func = CachedFunc(info)
File "<redacted>/site-packages/streamlit/runtime/caching/cache_utils.py", line 193, in __init__
self._function_key = _make_function_key(info.cache_type, info.func)
File "<redacted>/site-packages/streamlit/runtime/caching/cache_utils.py", line 488, in _make_function_key
source_code = inspect.getsource(func)
File "/usr/lib/python3.10/inspect.py", line 1139, in getsource
lines, lnum = getsourcelines(object)
File "/usr/lib/python3.10/inspect.py", line 1121, in getsourcelines
lines, lnum = findsource(object)
File "/usr/lib/python3.10/inspect.py", line 940, in findsource
file = getsourcefile(object)
File "/usr/lib/python3.10/inspect.py", line 817, in getsourcefile
filename = getfile(object)
File "/usr/lib/python3.10/inspect.py", line 797, in getfile
raise TypeError('module, class, method, function, traceback, frame, or '
TypeError: module, class, method, function, traceback, frame, or code object was expected, got cython_function_or_method
```
This trace was collected with `streamlit==1.41.0` but the issue persists with the current branch. | closed | 2025-02-06T11:09:52Z | 2025-02-07T11:49:35Z | https://github.com/streamlit/streamlit/issues/10353 | [
"type:enhancement",
"feature:cache"
] | tutu-sol | 1 |
pyeve/eve | flask | 1,399 | Cannot get list of elements with query contain positive timezone (with +) | Looks python eve is not able to handle query contains positive timezone (with +).
Query with is not working:
https://xxxxx/ExampleTable?where={"date":{"$gte": "2020-06-26T00:00:00+00:00", "$lte": "2020-06-27T12:12:55+00:00" }}
and working one:
https://xxxx/ExampleTable?where={"date":{"$gte": "2020-06-26T00:00:00-00:00", "$lte": "2020-06-27T12:12:55-00:00" }}
As a workaround we can do":
query=query.replace("+" , "-")
On MongoDB ExampleTable has a date: 2020-06-26T12:00:00+00:00
Can you please investigate and fix it up?
thank you
Dariusz | closed | 2020-06-29T07:50:59Z | 2022-04-16T07:39:33Z | https://github.com/pyeve/eve/issues/1399 | [
"stale"
] | dariuszsq | 2 |
jupyter-widgets-contrib/ipycanvas | jupyter | 85 | Example for interactively controlling an animation/games | I have managed to create smooth physics animations within jupyter using ipcanvas. I have also managed to successfully use ipyevents on ipycanvas to trigger events.
However I am struggling to combine events within animation loop. This would be required to run a game on ipycanvas, for example when pressing keys to change the direction of a spaceship flying across the canvas.
When the animation loop is running, it appears to block the events from being processed.
I can run my animation like this:
```python
def run_game():
for i in range(5000):
with hold_canvas(space):
space.clear()
space.fill_style = 'black'
space.fill_rect(0,0, width, height)
ship.update()
space.fill_style = 'white'
space.fill_arc(ship.position.x, ship.position.y, ship.size, 0, math.pi * 2)
```
And I can specify an event changing the ship's velocity like this:
```python
from ipyevents import Event
d = Event(source=space, watched_events=['keydown'])
d.on_dom_event(ship.thrusting)
```
Each one works on their own, but the event does not fire while the run_game() is running because it is blocking.
Is there a way to run this asynchronously?
Could you perhaps provide an example, which shows how one would write a game for ipycanvas? | closed | 2020-04-10T22:39:35Z | 2020-04-14T06:42:54Z | https://github.com/jupyter-widgets-contrib/ipycanvas/issues/85 | [] | tomanizer | 7 |
vitalik/django-ninja | django | 550 | [BUG] dead link in Proposal section | **Describe the bug**
The Enhancement Proposals intro page of the documentation currently has a dead link titled "Schemas from Django models"
Unclear whether this is a future way of getting Schemas out of Models or an old link now that SchemaModels exist
**Versions (please complete the following information):**
- Django-Ninja version: 0.19.1
| open | 2022-09-01T17:49:05Z | 2022-09-01T17:49:05Z | https://github.com/vitalik/django-ninja/issues/550 | [] | cltrudeau | 0 |
deepinsight/insightface | pytorch | 2,422 | Is the procedure of making rec file same for cosface and triplet training using arcface_mxnet ? and if not what is the procedure to make list, idx and rec file for triplet finetuning ? | open | 2023-09-05T10:01:54Z | 2023-09-05T10:17:57Z | https://github.com/deepinsight/insightface/issues/2422 | [] | Daishinkan002 | 0 | |
huggingface/pytorch-image-models | pytorch | 1,373 | Cannot create TensorRT inference engine for mobilevit | **Describe the bug**
Mobilevit onnx to TensorRT engine fails
**To Reproduce**
Steps to reproduce the behavior:
1.Export mobilevit_s model to onnx
2. Use trtexec to try and create TensorRT engine
```Shell
/usr/src/tensorrt/bin/trtexec --onnx=mobilevit.onnx --fp16 --workspace=2000 --saveEngine=mobilevit.engine
```
**Expected behavior**
Exported TensorRT engine
**Screenshots**

**Desktop (please complete the following information):**
- OS: Ubuntu 18.04 (Jetson Nano)
- This repository version: 6f103a442bb055b1fcdcf350aa816970e95ed125
- PyTorch version w/ CUDA/cuDNN PyTorch 1.10, CUDA 10.2
**Additional context**
Add any other context about the problem here.
| closed | 2022-07-27T16:19:14Z | 2022-08-01T17:06:22Z | https://github.com/huggingface/pytorch-image-models/issues/1373 | [
"bug"
] | dataplayer12 | 2 |
plotly/dash-table | dash | 503 | persisting column names across edits | Similar to #314 - if a user changes the column name in a table, being able to repopulate that column name automatically from localstorage when the table is re-rendered.
This would be a flag that the dash developer would set and the behaviour would be turned on or off. The end-user would not be able to turn this behaviour on or off. | closed | 2019-07-15T21:20:10Z | 2019-09-16T14:07:08Z | https://github.com/plotly/dash-table/issues/503 | [
"dash-type-enhancement",
"dash-meta-sponsored",
"size: 3"
] | chriddyp | 1 |
chatanywhere/GPT_API_free | api | 291 | https://api.chatanywhere.tech/audio/speech免费接口能用嘛 | **Is your feature request related to a problem? Please describe.**
A clear and concise description of what the problem is. Ex. I'm always frustrated when [...]
**Describe the solution you'd like**
A clear and concise description of what you want to happen.
**Describe alternatives you've considered**
A clear and concise description of any alternative solutions or features you've considered.
**Additional context**
Add any other context or screenshots about the feature request here.
| closed | 2024-09-10T19:59:49Z | 2024-09-11T09:23:29Z | https://github.com/chatanywhere/GPT_API_free/issues/291 | [] | lilian-lilifox | 2 |
miguelgrinberg/python-socketio | asyncio | 79 | Callback Documentation | I was having a hard time figuring out how to get client callbacks working (RPC style). After a short struggle I figured out how to respond to an individual request:
```
@sio.on('reverse', namespace='/test/')
async def reverse(sid, message):
return {'data': message[::-1]}
```
and the javascript (sorry, it's ES6 style):
```
onMyEvent = () => {
this.socket.emit(
'reverse',
{'data': 'howdy'},
response => alert(`Responded with: ${response.data}`)
);
}
```
I don't know how the inverse is supposed to work, though. I imagine it has something to do with the `callback` argument in the [`AsyncManager`](http://python-socketio.readthedocs.io/en/latest/#socketio.AsyncManager.emit). Can you explain? | closed | 2017-03-10T17:41:58Z | 2017-03-10T20:34:10Z | https://github.com/miguelgrinberg/python-socketio/issues/79 | [
"question"
] | dfee | 3 |
huggingface/transformers | pytorch | 35,957 | Cannot import 'GenerationOutput' in 4.48.1 | ### System Info
- `transformers` version: 4.48.1
- Platform: Linux-5.15.167.4-microsoft-standard-WSL2-x86_64-with-glibc2.31
- Python version: 3.9.5
- Huggingface_hub version: 0.28.0
- Safetensors version: 0.5.2
- Accelerate version: not installed
- Accelerate config: not found
- PyTorch version (GPU?): 2.5.1+cu124 (True)
- Tensorflow version (GPU?): not installed (NA)
- Flax version (CPU?/GPU?/TPU?): not installed (NA)
- Jax version: not installed
- JaxLib version: not installed
- Using distributed or parallel set-up in script?: <fill in>
- Using GPU in script?: <fill in>
- GPU type: NVIDIA GeForce MX450
### Who can help?
@gante
### Information
- [ ] The official example scripts
- [x] My own modified scripts
### Tasks
- [ ] An officially supported task in the `examples` folder (such as GLUE/SQuAD, ...)
- [x] My own task or dataset (give details below)
### Reproduction
```py
from transformers import AutoModelForCausalLM, AutoTokenizer, GenerationOutput
import torch
# Load model and tokenizer
model_name = "gpt2"
model = AutoModelForCausalLM.from_pretrained(model_name)
tokenizer = AutoTokenizer.from_pretrained(model_name)
# Encode input text
input_text = "Hello, how are you?"
input_ids = tokenizer(input_text, return_tensors="pt").input_ids
# Generate text (Return logits)
output = model.generate(
input_ids,
return_dict_in_generate=True,
output_scores=True,
return_logits=True
)
# Check if the output type is GenerationOutput
print(isinstance(output, GenerationOutput)) # True
```
### Expected behavior
The code above should run without any errors. | closed | 2025-01-29T13:22:00Z | 2025-03-13T08:03:51Z | https://github.com/huggingface/transformers/issues/35957 | [
"bug"
] | inthree3 | 4 |
raphaelvallat/pingouin | pandas | 356 | How to check n-way ANOVA assumptions using pingouin? | Suppose I'm doing this experimental design with 3 factors, 2 levels each and 3 repetitions:
```
import pingouin as pg
import numpy as np
import pandas as pd
from itertools import product
y_measures = np.array([28,25,27,18,19,23,36,32,32,31,30,29,28,25,22,18,19,23,12,32,40,31,30,29])
factors = ["Facotr A" ,"Factor B", "Factor C"]
levels_list = [['low','high'],['low','high'],['low','high']]
replicates = 3
def generate_dataframe(measures, factors, levels_list, replicates):
lines = []
for factor_combination in product(*levels_list):
line = {}
for idx, factor in enumerate(factors):
line[factor] = factor_combination[idx]
for k in range(replicates):
lines.append(line)
df = pd.DataFrame(lines,columns=factors)
df['y'] = measures
return df
df = generate_dataframe(y_measures, factors, levels_list, replicates)
```
1) What is the correct function to run ANOVA with repeatead measures using 3 or more factors? I tryed `rm_anova`, but it raises an error for more than 2 factors.
I'm trying this, but not sure if it is correct:
```
model1 = pg.anova(dv='y', between=factors, data=df, detailed=True)
```
2) I saw that there is a function `pg.power_anova`. What does exactly it measure?
3) What is the correct way to test assumptions of ANOVA for factorial design like this above? Should I test normality of measures (y) for each Factor in my experiment? Or should I test y grouping by all factors in my dataframe?
And about variance?
I wrote the code below, but not sure if I'm doing it right:
```
measures = []
for name, group in self.df.groupby(self.factors):
group_measures = group['y'].values
k2, p = stats.normaltest(group_measures)
print('Normality test for group', name, p >= 0.05)
print('Variance for group', name, np.var(group_measures, ddof=1)) # type: ignore
measures.append(group_measures)
k2, p = stats.levene(*measures) # type: ignore
print('Variance teste between groups', p, 'p >= 0.05', p >= 0.05)
```
| closed | 2023-04-19T23:07:56Z | 2023-06-04T17:37:21Z | https://github.com/raphaelvallat/pingouin/issues/356 | [
"question :raising_hand:"
] | vabatista | 1 |
babysor/MockingBird | deep-learning | 395 | 前所未有的问题……预处理时显示到100%,但是什么都没生成,什么情况啊 | aidatatang_200zh: 100%|█████████████████████████████████████████████████████| 2247/2247 [00:08<00:00, 276.10speakers/s]
The dataset consists of 0 utterances, 0 mel frames, 0 audio timesteps (0.00 hours).
Traceback (most recent call last):
File "C:\WorkSpace\Project\MockingBird-main\pre.py", line 74, in <module>
preprocess_dataset(**vars(args))
File "C:\WorkSpace\Project\MockingBird-main\synthesizer\preprocess.py", line 88, in preprocess_dataset
print("Max input length (text chars): %d" % max(len(m[5]) for m in metadata))
ValueError: max() arg is an empty sequence
这是什么报错OTZ问了一圈好像没人遇到这个…… | closed | 2022-02-20T10:14:55Z | 2023-07-01T08:13:40Z | https://github.com/babysor/MockingBird/issues/395 | [] | yy35959199 | 5 |
qwj/python-proxy | asyncio | 19 | how to set the ssr config of protocol? | closed | 2018-12-21T13:25:03Z | 2018-12-23T15:42:40Z | https://github.com/qwj/python-proxy/issues/19 | [] | fatfatson | 2 | |
NVIDIA/pix2pixHD | computer-vision | 301 | RuntimeError: DataLoader worker (pid 3752395) is killed by signal: Bus error. It is possible that dataloader's workers are out of shared memory | closed | 2022-05-12T22:47:36Z | 2022-06-21T05:31:44Z | https://github.com/NVIDIA/pix2pixHD/issues/301 | [] | Ghaleb-alnakhlani | 0 | |
plotly/dash-core-components | dash | 822 | [BUG] dcc.dropdown does not dropup when at bottom of screen/parent/viewport | With `dash==1.12`, the Dropdown component does not dropup (like `html <Select>`) when there is no space below the component. In my setup the component resides within a `dbc.ModalFooter`.
If there were at least an option for defining drop direction, I would be happy. | open | 2020-06-16T09:20:11Z | 2022-07-01T07:10:19Z | https://github.com/plotly/dash-core-components/issues/822 | [] | MM-Lehmann | 2 |
sanic-org/sanic | asyncio | 2,616 | How to change worker amount online | ### Is there an existing issue for this?
- [X] I have searched the existing issues
### Is your feature request related to a problem? Please describe.
Condition: must keep server online, then:
1. Is there a way to change (increase) the amount of running workers?
2. How can I add a new worker, by multiplexer, or manager?
The "online" means never lose any request all the time
### Describe the solution you'd like
1. like gunicorn, can modify the "workers" in setting, then send a HUP signal to main process.
2. like gunicorn, send a TTIN signal to main process.
### Additional context
thanks. | closed | 2022-12-07T14:08:13Z | 2022-12-08T01:14:43Z | https://github.com/sanic-org/sanic/issues/2616 | [
"feature request"
] | yangbo1024 | 3 |
ultralytics/yolov5 | deep-learning | 12,806 | An error in YOLOv5s summary | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
After my YOLO v5 model trained, an error was apear in "YOLOv5s summary" process, and I have no idea how to solve it. If anyone know that, please tell me, thank you!

### Additional
_No response_ | closed | 2024-03-11T03:01:30Z | 2024-10-20T19:41:15Z | https://github.com/ultralytics/yolov5/issues/12806 | [
"question",
"Stale"
] | qruiwu | 6 |
psf/requests | python | 6,164 | How to write python script setting system-wide http proxy using requests | Hello,
I've written a Python script exporting http proxy:
```
import os
proxy = "http://proxy:port"
os.environ['http_proxy'] = proxy
os.environ['HTTP_PROXY'] = proxy
```
But the thing is that I want to set the http proxy on system wide without exporting the environment variables using the requests library.
How to do that? I am Python newbie.. Please help. | closed | 2022-06-17T11:06:52Z | 2023-06-18T00:03:26Z | https://github.com/psf/requests/issues/6164 | [] | 1nrho12 | 4 |
HumanSignal/labelImg | deep-learning | 20 | how to create and save rotation bounding box? | I want to create and save the rotation bounding box by record the rotation angle,
| open | 2016-09-29T09:22:00Z | 2023-02-03T18:29:18Z | https://github.com/HumanSignal/labelImg/issues/20 | [
"enhancement"
] | taopanpan | 16 |
jina-ai/serve | machine-learning | 5,653 | CUDA_VISIBLE_DEVICES=RR & env={"CUDA_VISIBLE_DEVICES":"RR"} can not work | I tried to deploy multiple replica wtih multiple GPUs, but `CUDA_VISIBLE_DEVICES=RR` and `env={"CUDA_VISIBLE_DEVICES":"RR"}` do not work [as document said](https://docs.jina.ai/concepts/flow/scale-out/#replicate-on-multiple-gpus).
### Code
```python
# CUDA_VISIBLE_DEVICES=RR JINA_MP_START_METHOD=spawn python test_flow.py
from diffusers import DiffusionPipeline,EulerAncestralDiscreteScheduler
from diffusers import DPMSolverMultistepScheduler
from jina import Executor, requests,Flow
import torch
import time
class ZRExecutor(Executor):
def __init__(self,**kwargs):
super().__init__(**kwargs)
print('torch.cuda.device_count()',torch.cuda.device_count())
print('torch.cuda.current_device()',torch.cuda.current_device())
before_load=(torch.cuda.memory_allocated())/1024/1024
print('before load model:',before_load)
model_path = "#######"
lms = EulerAncestralDiscreteScheduler(
beta_start=0.00085,
beta_end=0.012,
beta_schedule="scaled_linear"
)
pipe = DiffusionPipeline.from_pretrained(
model_path,
cache_dir="./huggingface",
resume_download=True,
custom_pipeline="lpw_stable_diffusion",
torch_dtype=torch.float16,
scheduler=lms,
use_auth_token="#######",
safety_checker=None
)
pipe.to("cuda")
print('after load model:',(torch.cuda.memory_allocated())/1024/1024)
print('used memory:',(torch.cuda.memory_allocated()-before_load)/1024/1024)
def main():
f = Flow().add(uses=ZRExecutor,name='testens',replicas=3,env={"CUDA_VISIBLE_DEVICES":"RR"})
with f:
f.block()
if __name__ == '__main__':
main()
```
it will raise error
```python
ERROR testens/rep-0@778717 RuntimeError('No CUDA GPUs are available') during <class 'jina.serve.runtimes.worker.WorkerRuntime'> initialization [02/03/23 17:26:04]
add "--quiet-error" to suppress the exception details
Traceback (most recent call last):
File "/root/envs/(***)/lib/python3.8/site-packages/jina/orchestrate/pods/__init__.py", line 76, in run
runtime = runtime_cls(
File "/root/envs/(***)/lib/python3.8/site-packages/jina/serve/runtimes/worker/__init__.py", line 36, in __init__
super().__init__(args, **kwargs)
File "/root/envs/(***)/lib/python3.8/site-packages/jina/serve/runtimes/asyncio.py", line 88, in __init__
self._loop.run_until_complete(self.async_setup())
File "/root/envs/(***)/lib/python3.8/asyncio/base_events.py", line 616, in run_until_complete
return future.result()
File "/root/envs/(***)/lib/python3.8/site-packages/jina/serve/runtimes/worker/__init__.py", line 101, in async_setup
self._request_handler = WorkerRequestHandler(
File "/root/envs/(***)/lib/python3.8/site-packages/jina/serve/runtimes/worker/request_handling.py", line 49, in __init__
self._load_executor(
File "/root/envs/(***)/lib/python3.8/site-packages/jina/serve/runtimes/worker/request_handling.py", line 140, in _load_executor
self._executor: BaseExecutor = BaseExecutor.load_config(
File "/root/envs/(***)/lib/python3.8/site-packages/jina/jaml/__init__.py", line 760, in load_config
obj = JAML.load(tag_yml, substitute=False, runtime_args=runtime_args)
File "/root/envs/(***)/lib/python3.8/site-packages/jina/jaml/__init__.py", line 174, in load
r = yaml.load(stream, Loader=get_jina_loader_with_runtime(runtime_args))
File "/root/envs/(***)/lib/python3.8/site-packages/yaml/__init__.py", line 81, in load
return loader.get_single_data()
File "/root/envs/(***)/lib/python3.8/site-packages/yaml/constructor.py", line 51, in get_single_data
return self.construct_document(node)
File "/root/envs/(***)/lib/python3.8/site-packages/yaml/constructor.py", line 55, in construct_document
data = self.construct_object(node)
File "/root/envs/(***)/lib/python3.8/site-packages/yaml/constructor.py", line 100, in construct_object
data = constructor(self, node)
File "/root/envs/(***)/lib/python3.8/site-packages/jina/jaml/__init__.py", line 582, in _from_yaml
return get_parser(cls, version=data.get('version', None)).parse(
File "/root/envs/(***)/lib/python3.8/site-packages/jina/jaml/parsers/executor/legacy.py", line 45, in parse
obj = cls(
File "/root/envs/(***)/lib/python3.8/site-packages/jina/serve/executors/decorators.py", line 60, in arg_wrapper
f = func(self, *args, **kwargs)
File "/root/envs/(***)/lib/python3.8/site-packages/jina/serve/helper.py", line 71, in arg_wrapper
f = func(self, *args, **kwargs)
File "/root/autodl-nas/zrr/jina_test/test_flow.py", line 30, in __init__
pipe.to("cuda")
File "/root/envs/(***)/lib/python3.8/site-packages/diffusers/pipelines/pipeline_utils.py", line 272, in to
module.to(torch_device)
File "/root/envs/(***)/lib/python3.8/site-packages/transformers/modeling_utils.py", line 1682, in to
return super().to(*args, **kwargs)
File "/root/envs/(***)/lib/python3.8/site-packages/torch/nn/modules/module.py", line 987, in to
return self._apply(convert)
File "/root/envs/(***)/lib/python3.8/site-packages/torch/nn/modules/module.py", line 639, in _apply
module._apply(fn)
File "/root/envs/(***)/lib/python3.8/site-packages/torch/nn/modules/module.py", line 639, in _apply
module._apply(fn)
File "/root/envs/(***)/lib/python3.8/site-packages/torch/nn/modules/module.py", line 639, in _apply
module._apply(fn)
File "/root/envs/(***)/lib/python3.8/site-packages/torch/nn/modules/module.py", line 662, in _apply
param_applied = fn(param)
File "/root/envs/(***)/lib/python3.8/site-packages/torch/nn/modules/module.py", line 985, in convert
return t.to(device, dtype if t.is_floating_point() or t.is_complex() else None, non_blocking)
File "/root/envs/(***)/lib/python3.8/site-packages/torch/cuda/__init__.py", line 229, in _lazy_init
torch._C._cuda_init()
RuntimeError: No CUDA GPUs are available
```
If I remove `CUDA_VISIBLE_DEVICES=RR` and run the code, it will run successfully. However I checked the GPU usage, it seems that all models are running on GPU 0. And the script output `torch.cuda.current_device() 0` three times.
```bash
nvidia-smi
Fri Feb 3 17:47:23 2023
+-----------------------------------------------------------------------------+
| NVIDIA-SMI 510.60.02 Driver Version: 510.60.02 CUDA Version: 11.6 |
|-------------------------------+----------------------+----------------------+
| GPU Name Persistence-M| Bus-Id Disp.A | Volatile Uncorr. ECC |
| Fan Temp Perf Pwr:Usage/Cap| Memory-Usage | GPU-Util Compute M. |
| | | MIG M. |
|===============================+======================+======================|
| 0 NVIDIA RTX A5000 On | 00000000:01:00.0 Off | Off |
| 30% 29C P2 58W / 230W | 8491MiB / 24564MiB | 0% Default |
| | | N/A |
+-------------------------------+----------------------+----------------------+
| 1 NVIDIA RTX A5000 On | 00000000:25:00.0 Off | Off |
| 30% 23C P8 14W / 230W | 2MiB / 24564MiB | 0% Default |
| | | N/A |
+-------------------------------+----------------------+----------------------+
| 2 NVIDIA RTX A5000 On | 00000000:41:00.0 Off | Off |
| 30% 22C P8 14W / 230W | 2MiB / 24564MiB | 0% Default |
| | | N/A |
+-------------------------------+----------------------+----------------------+
+-----------------------------------------------------------------------------+
| Processes: |
| GPU GI CI PID Type Process name GPU Memory |
| ID ID Usage |
|=============================================================================|
| 0 N/A N/A 997681 C 2811MiB |
| 0 N/A N/A 997682 C 2839MiB |
| 0 N/A N/A 997683 C 2839MiB |
+-----------------------------------------------------------------------------+
```
### Jina version
```bash
jina --version-full
- jina 3.13.2
- docarray 0.21.0
- jcloud 0.2.1
- jina-hubble-sdk 0.32.0
- jina-proto 0.1.13
- protobuf 4.21.12
- proto-backend upb
- grpcio 1.47.2
- pyyaml 6.0
- python 3.8.10
- platform Linux
- platform-release 5.4.0-91-generic
- platform-version #102-Ubuntu SMP Fri Nov 5 16:31:28 UTC 2021
- architecture x86_64
- processor x86_64
- uid 2485377892355
- session-id 87650b9c-a3b0-11ed-8d5d-0242ac110003
- uptime 2023-02-03T18:50:09.712441
- ci-vendor (unset)
- internal False
* JINA_DEFAULT_HOST (unset)
* JINA_DEFAULT_TIMEOUT_CTRL (unset)
* JINA_DEPLOYMENT_NAME (unset)
* JINA_DISABLE_UVLOOP (unset)
* JINA_EARLY_STOP (unset)
* JINA_FULL_CLI (unset)
* JINA_GATEWAY_IMAGE (unset)
* JINA_GRPC_RECV_BYTES (unset)
* JINA_GRPC_SEND_BYTES (unset)
* JINA_HUB_NO_IMAGE_REBUILD (unset)
* JINA_LOG_CONFIG (unset)
* JINA_LOG_LEVEL (unset)
* JINA_LOG_NO_COLOR (unset)
* JINA_MP_START_METHOD (unset)
* JINA_OPTOUT_TELEMETRY (unset)
* JINA_RANDOM_PORT_MAX (unset)
* JINA_RANDOM_PORT_MIN (unset)
* JINA_LOCKS_ROOT (unset)
* JINA_K8S_ACCESS_MODES (unset)
* JINA_K8S_STORAGE_CLASS_NAME (unset)
* JINA_K8S_STORAGE_CAPACITY (unset)
* JINA_STREAMER_ARGS (unset)
``` | closed | 2023-02-03T11:52:50Z | 2023-02-22T09:06:52Z | https://github.com/jina-ai/serve/issues/5653 | [] | ruanrz | 9 |
statsmodels/statsmodels | data-science | 8,688 | ENH: model equivalence testing | reversing the null hypothesis in model comparisons and gof tests.
not a very popular topic, but should be more used
Main practical problem is how to specify insignificant "effect size" margin for model comparisons and gof.
example where it would work is FTestRegressionPower, i.e. Cohen's f2 effect size that can be related to (partial) R-square. In these cases we have a interpretable scaled noncentrality nc/nobs.
Independently of how to define equivalence margin, we could just add the functions.
The target would be to extend equivalence testing from one and two sample functions to models, model comparisons and diagnostic and specification tests.
These equivalence tests would be targeted to specific statistics, e.g. gof in general, directional misspecification, params, predictive test (?), ... depending on the test statistic that is used.
e.g.
- show that interaction effect is close to zero (equivalent models with and without interaction effect)
- show that some statistics are equivalent for models with different link functions.
- ...
| open | 2023-02-19T19:09:34Z | 2023-02-19T19:12:43Z | https://github.com/statsmodels/statsmodels/issues/8688 | [
"type-enh",
"comp-stats",
"topic-diagnostic"
] | josef-pkt | 1 |
tensorflow/tensor2tensor | machine-learning | 957 | t2t-trainer eval_early_stopping crashed at GetAccumulator() with KeyError 'run0' | ### Description
With t2t 1.6.6, tensorflow 1.8.0, I ran cifar100 with eval early stopping. The cmd failed quickly with crash at tensorboard/backend/event_processing/event_multiplexer.py, GetAccumulator() with KeyError 'run0'
### Environment information
```
OS: CentOS 7.4 x64
$ pip freeze | grep tensor
tensor2tensor==1.6.6
tensorboard==1.8.0
tensorflow==1.8.0
$ python -V
Python 2.7.5
```
### For bugs: reproduction and error logs
```
t2t-trainer --generate_data --tmp_dir=./tmp --data_dir=./cifar-100-python --output_dir=./cifar-out --problem=image_cifar100 --model=resnet --hparams_set=resnet_18 --hparams=learning_rate=0.001 --worker_gpu=1 --eval_early_stopping_steps=10 --schedule=train --train_steps=3000 --eval_steps=100
```
```
# Error logs:
INFO:tensorflow:Generating data for image_cifar100
INFO:tensorflow:Not downloading, file already found: ./tmp/cifar-100-python.tar.gz
INFO:tensorflow:Not downloading, file already found: ./tmp/cifar-100-python.tar.gz
2018-07-26 13:48:31.403947: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1356] Found device 0 with properties:
name: Tesla P100-PCIE-16GB major: 6 minor: 0 memoryClockRate(GHz): 1.3285
pciBusID: 0000:04:00.0
totalMemory: 15.90GiB freeMemory: 15.61GiB
2018-07-26 13:48:31.590744: I tensorflow/stream_executor/cuda/cuda_gpu_executor.cc:898] successful NUMA node read from SysFS had negative value (-1), but there must be at least one NUMA node, so returning NUMA node zero
2018-07-26 13:48:31.591518: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1356] Found device 1 with properties:
name: Tesla P100-PCIE-16GB major: 6 minor: 0 memoryClockRate(GHz): 1.3285
pciBusID: 0000:84:00.0
totalMemory: 15.90GiB freeMemory: 15.61GiB
2018-07-26 13:48:31.591578: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1435] Adding visible gpu devices: 0, 1
2018-07-26 13:48:32.201392: I tensorflow/core/common_runtime/gpu/gpu_device.cc:923] Device interconnect StreamExecutor with strength 1 edge matrix:
2018-07-26 13:48:32.201453: I tensorflow/core/common_runtime/gpu/gpu_device.cc:929] 0 1
2018-07-26 13:48:32.201463: I tensorflow/core/common_runtime/gpu/gpu_device.cc:942] 0: N N
2018-07-26 13:48:32.201469: I tensorflow/core/common_runtime/gpu/gpu_device.cc:942] 1: N N
2018-07-26 13:48:32.202163: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1053] Created TensorFlow device (/job:localhost/replica:0/task:0/device:GPU:0 with 15137 MB memory) -> physical GPU (device: 0, name: Tesla P100-PCIE-16GB, pci bus id: 0000:04:00.0, compute capability: 6.0)
2018-07-26 13:48:32.380463: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1053] Created TensorFlow device (/job:localhost/replica:0/task:0/device:GPU:1 with 15137 MB memory) -> physical GPU (device: 1, name: Tesla P100-PCIE-16GB, pci bus id: 0000:84:00.0, compute capability: 6.0)
INFO:tensorflow:Generating case 0.
INFO:tensorflow:Generated 50000 Examples
2018-07-26 13:48:58.058549: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1435] Adding visible gpu devices: 0, 1
2018-07-26 13:48:58.058660: I tensorflow/core/common_runtime/gpu/gpu_device.cc:923] Device interconnect StreamExecutor with strength 1 edge matrix:
2018-07-26 13:48:58.058672: I tensorflow/core/common_runtime/gpu/gpu_device.cc:929] 0 1
2018-07-26 13:48:58.058679: I tensorflow/core/common_runtime/gpu/gpu_device.cc:942] 0: N N
2018-07-26 13:48:58.058685: I tensorflow/core/common_runtime/gpu/gpu_device.cc:942] 1: N N
2018-07-26 13:48:58.058947: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1053] Created TensorFlow device (/job:localhost/replica:0/task:0/device:GPU:0 with 15137 MB memory) -> physical GPU (device: 0, name: Tesla P100-PCIE-16GB, pci bus id: 0000:04:00.0, compute capability: 6.0)
2018-07-26 13:48:58.059052: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1053] Created TensorFlow device (/job:localhost/replica:0/task:0/device:GPU:1 with 15137 MB memory) -> physical GPU (device: 1, name: Tesla P100-PCIE-16GB, pci bus id: 0000:84:00.0, compute capability: 6.0)
INFO:tensorflow:Generating case 0.
INFO:tensorflow:Generated 10000 Examples
INFO:tensorflow:Shuffling data...
INFO:tensorflow:Data shuffled.
INFO:tensorflow:Overriding hparams in resnet_18 with learning_rate=0.001
WARNING:tensorflow:From /usr/lib/python2.7/site-packages/tensor2tensor/utils/trainer_lib.py:165: __init__ (from tensorflow.contrib.learn.python.learn.estimators.run_config) is deprecated and will be removed in a future version.
Instructions for updating:
When switching to tf.estimator.Estimator, use tf.estimator.RunConfig instead.
INFO:tensorflow:schedule=train
INFO:tensorflow:worker_gpu=1
INFO:tensorflow:sync=False
INFO:tensorflow:datashard_devices: ['/job:localhost']
INFO:tensorflow:caching_devices: None
INFO:tensorflow:ps_devices: ['gpu:0']
INFO:tensorflow:Using config: {'_save_checkpoints_secs': None, '_keep_checkpoint_max': 20, '_task_type': None, '_cluster_spec': <tensorflow.python.training.server_lib.ClusterSpec object at 0xb2d5190>, '_keep_checkpoint_every_n_hours': 10000, '_session_config': gpu_options {
per_process_gpu_memory_fraction: 0.95
}
allow_soft_placement: true
graph_options {
optimizer_options {
}
}
, 'use_tpu': False, '_tf_random_seed': None, '_num_worker_replicas': 0, '_task_id': 0, 't2t_device_info': {'num_async_replicas': 1}, '_evaluation_master': '', '_log_step_count_steps': 100, '_num_ps_replicas': 0, '_train_distribute': None, '_is_chief': True, '_tf_config': gpu_options {
per_process_gpu_memory_fraction: 1.0
}
, '_save_checkpoints_steps': 1000, '_environment': 'local', '_master': '', '_model_dir': './cifar-out', 'data_parallelism': <tensor2tensor.utils.expert_utils.Parallelism object at 0xb2d51d0>, '_save_summary_steps': 100}
WARNING:tensorflow:Estimator's model_fn (<function wrapping_model_fn at 0xaaecf50>) includes params argument, but params are not passed to Estimator.
INFO:tensorflow:Using EarlyStoppingHook
INFO:tensorflow:Event Multiplexer initializing.
INFO:tensorflow:Event Multplexer doing initialization load for {'run0': './cifar-out/eval_continuous/'}
INFO:tensorflow:Constructing EventAccumulator for ./cifar-out/eval_continuous/
INFO:tensorflow:Event Multiplexer done initializing
INFO:tensorflow:Reading data files from ./cifar-100-python/image_cifar100-train*
INFO:tensorflow:partition: 0 num_data_files: 10
INFO:tensorflow:Calling model_fn.
INFO:tensorflow:Setting T2TModel mode to 'train'
INFO:tensorflow:Using variable initializer: normal_unit_scaling
INFO:tensorflow:Transforming feature 'inputs' with image_modality.bottom
INFO:tensorflow:Transforming 'targets' with class_label_modality_100_64.targets_bottom
INFO:tensorflow:Building model body
INFO:tensorflow:Transforming body output with class_label_modality_100_64.top
WARNING:tensorflow:From /usr/lib/python2.7/site-packages/tensor2tensor/layers/modalities.py:703: calling reduce_mean (from tensorflow.python.ops.math_ops) with keep_dims is deprecated and will be removed in a future version.
Instructions for updating:
keep_dims is deprecated, use keepdims instead
INFO:tensorflow:Applying exp learning rate warmup for 100 steps
INFO:tensorflow:Applying learning rate decay: cosine.
INFO:tensorflow:Base learning rate: 0.001000
INFO:tensorflow:Applying weight decay, decay_rate: 0.00010
INFO:tensorflow:Trainable Variables Total size: 11231140
INFO:tensorflow:Using optimizer Momentum
INFO:tensorflow:Done calling model_fn.
INFO:tensorflow:Create CheckpointSaverHook.
INFO:tensorflow:Graph was finalized.
2018-07-26 13:49:08.290671: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1435] Adding visible gpu devices: 0, 1
2018-07-26 13:49:08.290789: I tensorflow/core/common_runtime/gpu/gpu_device.cc:923] Device interconnect StreamExecutor with strength 1 edge matrix:
2018-07-26 13:49:08.290802: I tensorflow/core/common_runtime/gpu/gpu_device.cc:929] 0 1
2018-07-26 13:49:08.290809: I tensorflow/core/common_runtime/gpu/gpu_device.cc:942] 0: N N
2018-07-26 13:49:08.290815: I tensorflow/core/common_runtime/gpu/gpu_device.cc:942] 1: N N
2018-07-26 13:49:08.291089: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1053] Created TensorFlow device (/job:localhost/replica:0/task:0/device:GPU:0 with 15137 MB memory) -> physical GPU (device: 0, name: Tesla P100-PCIE-16GB, pci bus id: 0000:04:00.0, compute capability: 6.0)
2018-07-26 13:49:08.291227: I tensorflow/core/common_runtime/gpu/gpu_device.cc:1053] Created TensorFlow device (/job:localhost/replica:0/task:0/device:GPU:1 with 15137 MB memory) -> physical GPU (device: 1, name: Tesla P100-PCIE-16GB, pci bus id: 0000:84:00.0, compute capability: 6.0)
INFO:tensorflow:Running local_init_op.
INFO:tensorflow:Done running local_init_op.
INFO:tensorflow:Saving checkpoints for 1 into ./cifar-out/model.ckpt.
INFO:tensorflow:Beginning EventMultiplexer.Reload()
WARNING:tensorflow:Deleting accumulator 'run0'
INFO:tensorflow:Finished with EventMultiplexer.Reload()
Traceback (most recent call last):
File "/usr/bin/t2t-trainer", line 32, in <module>
tf.app.run()
File "/usr/lib/python2.7/site-packages/tensorflow/python/platform/app.py", line 126, in run
_sys.exit(main(argv))
File "/usr/bin/t2t-trainer", line 28, in main
t2t_trainer.main(argv)
File "/usr/lib/python2.7/site-packages/tensor2tensor/bin/t2t_trainer.py", line 359, in main
execute_schedule(exp)
File "/usr/lib/python2.7/site-packages/tensor2tensor/bin/t2t_trainer.py", line 306, in execute_schedule
getattr(exp, FLAGS.schedule)()
File "/usr/lib/python2.7/site-packages/tensor2tensor/utils/trainer_lib.py", line 303, in train
max_steps=self._train_spec.max_steps)
File "/usr/lib/python2.7/site-packages/tensorflow/python/estimator/estimator.py", line 363, in train
loss = self._train_model(input_fn, hooks, saving_listeners)
File "/usr/lib/python2.7/site-packages/tensorflow/python/estimator/estimator.py", line 843, in _train_model
return self._train_model_default(input_fn, hooks, saving_listeners)
File "/usr/lib/python2.7/site-packages/tensorflow/python/estimator/estimator.py", line 859, in _train_model_default
saving_listeners)
File "/usr/lib/python2.7/site-packages/tensorflow/python/estimator/estimator.py", line 1059, in _train_with_estimator_spec
_, loss = mon_sess.run([estimator_spec.train_op, estimator_spec.loss])
File "/usr/lib/python2.7/site-packages/tensorflow/python/training/monitored_session.py", line 567, in run
run_metadata=run_metadata)
File "/usr/lib/python2.7/site-packages/tensorflow/python/training/monitored_session.py", line 1043, in run
run_metadata=run_metadata)
File "/usr/lib/python2.7/site-packages/tensorflow/python/training/monitored_session.py", line 1134, in run
raise six.reraise(*original_exc_info)
File "/usr/lib/python2.7/site-packages/tensorflow/python/training/monitored_session.py", line 1119, in run
return self._sess.run(*args, **kwargs)
File "/usr/lib/python2.7/site-packages/tensorflow/python/training/monitored_session.py", line 1199, in run
run_metadata=run_metadata))
File "/usr/lib/python2.7/site-packages/tensor2tensor/utils/metrics_hook.py", line 81, in after_run
metrics = self._collect_metrics()
File "/usr/lib/python2.7/site-packages/tensor2tensor/utils/metrics_hook.py", line 95, in _collect_metrics
accum = self._event_multiplexer.GetAccumulator(self._RUN_NAME % i)
File "/usr/lib/python2.7/site-packages/tensorboard/backend/event_processing/event_multiplexer.py", line 482, in GetAccumulator
return self._accumulators[run]
KeyError: 'run0'
```
| open | 2018-07-25T22:18:25Z | 2018-09-10T02:04:24Z | https://github.com/tensorflow/tensor2tensor/issues/957 | [] | LiweiPeng | 1 |
microsoft/qlib | machine-learning | 939 | fail to generate analysis report graphs | ## ❓ Questions and Help
I run the successfully the model training following ( examples/workflow_by_code.ipynb),pass the step "prediction, backtest & analysis", but fail to generate analyze graphs. not sure what happened. | closed | 2022-02-28T09:31:37Z | 2022-06-03T15:01:56Z | https://github.com/microsoft/qlib/issues/939 | [
"question",
"stale"
] | cssaudrey | 2 |
axnsan12/drf-yasg | django | 497 | Import ruamel.yaml issue | ``` from drf_yasg import openapi, views
File "/usr/local/lib/python2.7/site-packages/drf_yasg/views.py", line 13, in <module>
from .renderers import (
File "/usr/local/lib/python2.7/site-packages/drf_yasg/renderers.py", line 11, in <module>
from .codecs import VALIDATORS, OpenAPICodecJson, OpenAPICodecYaml
File "/usr/local/lib/python2.7/site-packages/drf_yasg/codecs.py", line 9, in <module>
from ruamel import yaml
ImportError: No module named ruamel
```
```from rest_framework import permissions
from drf_yasg.views import get_schema_view
from drf_yasg import openapi
schema_view = get_schema_view(
openapi.Info(
title="FPAAS APIS",
default_version='v1',
description="Food Personalisation Platform APIS",
terms_of_service="https://spoonshot.com/terms/"
),
public=False,
permission_classes=(permissions.IsAdminUser,),
)
urlpatterns = [
url(r'^swagger(?P<format>\.json|\.yaml)$',
schema_view.without_ui(cache_timeout=0),
name='schema-json'),
url(r'^swagger/$',
schema_view.with_ui('swagger', cache_timeout=0),
name='schema-swagger-ui'),
url(r'^redoc/$',
schema_view.with_ui('redoc', cache_timeout=0),
name='schema-redoc'),
]
```
This is the code i use
I am able to do `from ruamel import yaml` in python manage.py shell but when i run using uwsgi-nginx image by tianglo - python2.7-alpine3,9 it gives import error | closed | 2019-11-20T13:49:26Z | 2020-10-26T01:02:44Z | https://github.com/axnsan12/drf-yasg/issues/497 | [] | appunni-m | 2 |
plotly/dash-core-components | dash | 736 | Fix test instability | Some tests randomly fail on CI runs and make it hard to get (a) an accurate picture of the impact of changes, (b) a final approval for merge in GH, resulting in wasted time & effort.
Here's a sample taken from runs in the last few days. In all cases the test failed for some `test-pyXX` but passed for another `test-pyYY` run. Items with `***` are the most frequent offenders.
- test_stcp100_clear_data_on_all_types ***
- test_stdl001_data_lifecycle_with_different_condition ***
- test_graph_extend_trace[False]
- test_location_link
- test_grbs002_wrapped_graph_has_no_infinite_loop[False]
I haven't checked if the tests are unstable locally or if this only happens in CI. | open | 2020-01-20T14:03:13Z | 2020-01-30T20:50:56Z | https://github.com/plotly/dash-core-components/issues/736 | [
"dash-type-maintenance"
] | Marc-Andre-Rivet | 0 |
ydataai/ydata-profiling | jupyter | 1,331 | html.inline = False gets the src for javascript files wrong | ### Current Behaviour
setting `profile.config.html.inline = False`
and then `profile.to_file("all_data/longi_report.html")'`
assets are stored in `longi_report_assets/`
however the html file in several places has `src=_assets`
Loading the HTML file gives a broken page
### Expected Behaviour
Correct prefix is used.
### Data Description
N/A
### Code that reproduces the bug
```Python
profile = ProfileReport(data, title="Longitudinal profiling", minimal=True)
profile.config.html.inline = False
profile.to_file("all_data/longi_report.html")
```
### pandas-profiling version
v4.1.2
### Dependencies
```Text
# packages in environment at /gpfs/fs1/home/m/mchakrav/gdevenyi/mambaforge:
#
# Name Version Build Channel
_libgcc_mutex 0.1 conda_forge conda-forge
_openmp_mutex 4.5 2_gnu conda-forge
aiohttp 3.8.4 py310h1fa729e_0 conda-forge
aiosignal 1.3.1 pyhd8ed1ab_0 conda-forge
alsa-lib 1.2.8 h166bdaf_0 conda-forge
aom 3.5.0 h27087fc_0 conda-forge
argcomplete 3.0.5 pyhd8ed1ab_0 conda-forge
arrow-cpp 11.0.0 ha770c72_13_cpu conda-forge
asttokens 2.2.1 pyhd8ed1ab_0 conda-forge
async-timeout 4.0.2 pyhd8ed1ab_0 conda-forge
attr 2.5.1 h166bdaf_1 conda-forge
attrs 22.2.0 pyh71513ae_0 conda-forge
aws-c-auth 0.6.26 hf365957_1 conda-forge
aws-c-cal 0.5.21 h48707d8_2 conda-forge
aws-c-common 0.8.14 h0b41bf4_0 conda-forge
aws-c-compression 0.2.16 h03acc5a_5 conda-forge
aws-c-event-stream 0.2.20 h00877a2_4 conda-forge
aws-c-http 0.7.6 hf342b9f_0 conda-forge
aws-c-io 0.13.19 h5b20300_3 conda-forge
aws-c-mqtt 0.8.6 hc4349f7_12 conda-forge
aws-c-s3 0.2.7 h909e904_1 conda-forge
aws-c-sdkutils 0.1.8 h03acc5a_0 conda-forge
aws-checksums 0.1.14 h03acc5a_5 conda-forge
aws-crt-cpp 0.19.8 hf7fbfca_12 conda-forge
aws-sdk-cpp 1.10.57 h17c43bd_8 conda-forge
backcall 0.2.0 pyh9f0ad1d_0 conda-forge
backports 1.0 pyhd8ed1ab_3 conda-forge
backports.functools_lru_cache 1.6.4 pyhd8ed1ab_0 conda-forge
backports.zoneinfo 0.2.1 py310hff52083_7 conda-forge
boltons 23.0.0 pyhd8ed1ab_0 conda-forge
brotli 1.0.9 h166bdaf_8 conda-forge
brotli-bin 1.0.9 h166bdaf_8 conda-forge
brotlipy 0.7.0 py310h5764c6d_1005 conda-forge
bzip2 1.0.8 h7f98852_4 conda-forge
c-ares 1.18.1 h7f98852_0 conda-forge
ca-certificates 2023.5.7 hbcca054_0 conda-forge
cairo 1.16.0 ha61ee94_1014 conda-forge
certifi 2023.5.7 pyhd8ed1ab_0 conda-forge
cffi 1.15.1 py310h255011f_3 conda-forge
charset-normalizer 2.1.1 pyhd8ed1ab_0 conda-forge
click 8.1.3 unix_pyhd8ed1ab_2 conda-forge
colorama 0.4.6 pyhd8ed1ab_0 conda-forge
comm 0.1.3 pyhd8ed1ab_0 conda-forge
conda 23.3.1 py310hff52083_0 conda-forge
conda-package-handling 2.0.2 pyh38be061_0 conda-forge
conda-package-streaming 0.7.0 pyhd8ed1ab_1 conda-forge
contourpy 1.0.7 py310hdf3cbec_0 conda-forge
cryptography 40.0.1 py310h34c0648_0 conda-forge
curl 7.88.1 hdc1c0ab_1 conda-forge
cycler 0.11.0 pyhd8ed1ab_0 conda-forge
dbus 1.13.6 h5008d03_3 conda-forge
debugpy 1.6.7 py310heca2aa9_0 conda-forge
decorator 5.1.1 pyhd8ed1ab_0 conda-forge
double-conversion 3.2.0 h27087fc_1 conda-forge
eigen 3.4.0 h4bd325d_0 conda-forge
executing 1.2.0 pyhd8ed1ab_0 conda-forge
expat 2.5.0 hcb278e6_1 conda-forge
ffmpeg 5.1.2 gpl_h8dda1f0_106 conda-forge
fftw 3.3.10 nompi_hf0379b8_106 conda-forge
fmt 9.1.0 h924138e_0 conda-forge
font-ttf-dejavu-sans-mono 2.37 hab24e00_0 conda-forge
font-ttf-inconsolata 3.000 h77eed37_0 conda-forge
font-ttf-source-code-pro 2.038 h77eed37_0 conda-forge
font-ttf-ubuntu 0.83 hab24e00_0 conda-forge
fontconfig 2.14.2 h14ed4e7_0 conda-forge
fonts-conda-ecosystem 1 0 conda-forge
fonts-conda-forge 1 0 conda-forge
fonttools 4.39.3 py310h1fa729e_0 conda-forge
freetype 2.12.1 hca18f0e_1 conda-forge
frozenlist 1.3.3 py310h5764c6d_0 conda-forge
gettext 0.21.1 h27087fc_0 conda-forge
gflags 2.2.2 he1b5a44_1004 conda-forge
gl2ps 1.4.2 h0708190_0 conda-forge
glew 2.1.0 h9c3ff4c_2 conda-forge
glib 2.74.1 h6239696_1 conda-forge
glib-tools 2.74.1 h6239696_1 conda-forge
glog 0.6.0 h6f12383_0 conda-forge
gmp 6.2.1 h58526e2_0 conda-forge
gnutls 3.7.8 hf3e180e_0 conda-forge
graphite2 1.3.13 h58526e2_1001 conda-forge
gst-plugins-base 1.22.0 h4243ec0_2 conda-forge
gstreamer 1.22.0 h25f0c4b_2 conda-forge
gstreamer-orc 0.4.33 h166bdaf_0 conda-forge
harfbuzz 6.0.0 h8e241bc_0 conda-forge
hdf4 4.2.15 h9772cbc_5 conda-forge
hdf5 1.12.2 nompi_h4df4325_101 conda-forge
htmlmin 0.1.12 py_1 conda-forge
icu 70.1 h27087fc_0 conda-forge
idna 3.4 pyhd8ed1ab_0 conda-forge
imagehash 4.3.1 pyhd8ed1ab_0 conda-forge
importlib-metadata 6.1.0 pyha770c72_0 conda-forge
importlib_metadata 6.1.0 hd8ed1ab_0 conda-forge
importlib_resources 5.12.0 pyhd8ed1ab_0 conda-forge
ipykernel 6.23.1 pyh210e3f2_0 conda-forge
ipython 8.12.0 pyh41d4057_0 conda-forge
ipywidgets 8.0.6 pyhd8ed1ab_0 conda-forge
itk 5.3.0 py310hfdc917e_0 conda-forge
itk-core 5.3.0 pypi_0 pypi
itk-filtering 5.3.0 pypi_0 pypi
itk-numerics 5.3.0 pypi_0 pypi
itk-registration 5.3.0 pypi_0 pypi
itk-segmentation 5.3.0 pypi_0 pypi
jack 1.9.22 h11f4161_0 conda-forge
jedi 0.18.2 pyhd8ed1ab_0 conda-forge
jinja2 3.1.2 pyhd8ed1ab_1 conda-forge
joblib 1.2.0 pyhd8ed1ab_0 conda-forge
jpeg 9e h0b41bf4_3 conda-forge
jsoncpp 1.9.5 h4bd325d_1 conda-forge
jsonpatch 1.32 pyhd8ed1ab_0 conda-forge
jsonpointer 2.0 py_0 conda-forge
jupyter_client 8.2.0 pyhd8ed1ab_0 conda-forge
jupyter_core 5.3.0 py310hff52083_0 conda-forge
jupyterlab_widgets 3.0.7 pyhd8ed1ab_1 conda-forge
keyutils 1.6.1 h166bdaf_0 conda-forge
kiwisolver 1.4.4 py310hbf28c38_1 conda-forge
krb5 1.20.1 h81ceb04_0 conda-forge
lame 3.100 h166bdaf_1003 conda-forge
lcms2 2.14 h6ed2654_0 conda-forge
ld_impl_linux-64 2.40 h41732ed_0 conda-forge
lerc 4.0.0 h27087fc_0 conda-forge
libabseil 20230125.0 cxx17_hcb278e6_1 conda-forge
libaec 1.0.6 hcb278e6_1 conda-forge
libarchive 3.6.2 h3d51595_0 conda-forge
libarrow 11.0.0 h93537a5_13_cpu conda-forge
libblas 3.9.0 16_linux64_openblas conda-forge
libbrotlicommon 1.0.9 h166bdaf_8 conda-forge
libbrotlidec 1.0.9 h166bdaf_8 conda-forge
libbrotlienc 1.0.9 h166bdaf_8 conda-forge
libcap 2.67 he9d0100_0 conda-forge
libcblas 3.9.0 16_linux64_openblas conda-forge
libclang 15.0.7 default_had23c3d_1 conda-forge
libclang13 15.0.7 default_h3e3d535_1 conda-forge
libcrc32c 1.1.2 h9c3ff4c_0 conda-forge
libcups 2.3.3 h36d4200_3 conda-forge
libcurl 7.88.1 hdc1c0ab_1 conda-forge
libdb 6.2.32 h9c3ff4c_0 conda-forge
libdeflate 1.14 h166bdaf_0 conda-forge
libdrm 2.4.114 h166bdaf_0 conda-forge
libedit 3.1.20191231 he28a2e2_2 conda-forge
libev 4.33 h516909a_1 conda-forge
libevent 2.1.10 h28343ad_4 conda-forge
libexpat 2.5.0 hcb278e6_1 conda-forge
libffi 3.4.2 h7f98852_5 conda-forge
libflac 1.4.2 h27087fc_0 conda-forge
libgcc-ng 12.2.0 h65d4601_19 conda-forge
libgcrypt 1.10.1 h166bdaf_0 conda-forge
libgfortran-ng 12.2.0 h69a702a_19 conda-forge
libgfortran5 12.2.0 h337968e_19 conda-forge
libglib 2.74.1 h606061b_1 conda-forge
libglu 9.0.0 he1b5a44_1001 conda-forge
libgomp 12.2.0 h65d4601_19 conda-forge
libgoogle-cloud 2.8.0 h0bc5f78_1 conda-forge
libgpg-error 1.46 h620e276_0 conda-forge
libgrpc 1.52.1 hcf146ea_1 conda-forge
libhwloc 2.9.0 hd6dc26d_0 conda-forge
libiconv 1.17 h166bdaf_0 conda-forge
libidn2 2.3.4 h166bdaf_0 conda-forge
libitk 5.3.0 hcedbc38_0 conda-forge
liblapack 3.9.0 16_linux64_openblas conda-forge
libllvm15 15.0.7 hadd5161_1 conda-forge
libmamba 1.4.1 hcea66bb_0 conda-forge
libmambapy 1.4.1 py310h1428755_0 conda-forge
libnetcdf 4.8.1 nompi_h261ec11_106 conda-forge
libnghttp2 1.52.0 h61bc06f_0 conda-forge
libnsl 2.0.0 h7f98852_0 conda-forge
libnuma 2.0.16 h0b41bf4_1 conda-forge
libogg 1.3.4 h7f98852_1 conda-forge
libopenblas 0.3.21 pthreads_h78a6416_3 conda-forge
libopus 1.3.1 h7f98852_1 conda-forge
libpciaccess 0.17 h166bdaf_0 conda-forge
libpng 1.6.39 h753d276_0 conda-forge
libpq 15.2 hb675445_0 conda-forge
libprotobuf 3.21.12 h3eb15da_0 conda-forge
libsndfile 1.2.0 hb75c966_0 conda-forge
libsodium 1.0.18 h36c2ea0_1 conda-forge
libsolv 0.7.23 h3eb15da_0 conda-forge
libsqlite 3.40.0 h753d276_0 conda-forge
libssh2 1.10.0 hf14f497_3 conda-forge
libstdcxx-ng 12.2.0 h46fd767_19 conda-forge
libsystemd0 253 h8c4010b_1 conda-forge
libtasn1 4.19.0 h166bdaf_0 conda-forge
libtheora 1.1.1 h7f98852_1005 conda-forge
libthrift 0.18.1 h5e4af38_0 conda-forge
libtiff 4.4.0 h82bc61c_5 conda-forge
libtool 2.4.7 h27087fc_0 conda-forge
libudev1 253 h0b41bf4_1 conda-forge
libunistring 0.9.10 h7f98852_0 conda-forge
libutf8proc 2.8.0 h166bdaf_0 conda-forge
libuuid 2.38.1 h0b41bf4_0 conda-forge
libva 2.18.0 h0b41bf4_0 conda-forge
libvorbis 1.3.7 h9c3ff4c_0 conda-forge
libvpx 1.11.0 h9c3ff4c_3 conda-forge
libwebp-base 1.3.0 h0b41bf4_0 conda-forge
libxcb 1.13 h7f98852_1004 conda-forge
libxkbcommon 1.5.0 h79f4944_1 conda-forge
libxml2 2.10.3 hca2bb57_4 conda-forge
libzip 1.9.2 hc929e4a_1 conda-forge
libzlib 1.2.13 h166bdaf_4 conda-forge
loguru 0.6.0 py310hff52083_2 conda-forge
lz4-c 1.9.4 hcb278e6_0 conda-forge
lzo 2.10 h516909a_1000 conda-forge
mamba 1.4.1 py310h51d5547_0 conda-forge
markupsafe 2.1.2 py310h1fa729e_0 conda-forge
matplotlib-base 3.6.3 py310he60537e_0 conda-forge
matplotlib-inline 0.1.6 pyhd8ed1ab_0 conda-forge
mizani 0.8.1 pyhd8ed1ab_1 conda-forge
mpg123 1.31.3 hcb278e6_0 conda-forge
multidict 6.0.4 py310h1fa729e_0 conda-forge
multimethod 1.4 py_0 conda-forge
munkres 1.1.4 pyh9f0ad1d_0 conda-forge
mysql-common 8.0.32 ha901b37_1 conda-forge
mysql-libs 8.0.32 hd7da12d_1 conda-forge
ncurses 6.3 h27087fc_1 conda-forge
nest-asyncio 1.5.6 pyhd8ed1ab_0 conda-forge
nettle 3.8.1 hc379101_1 conda-forge
networkx 3.1 pyhd8ed1ab_0 conda-forge
nlohmann_json 3.11.2 h27087fc_0 conda-forge
nspr 4.35 h27087fc_0 conda-forge
nss 3.89 he45b914_0 conda-forge
numpy 1.23.5 py310h53a5b5f_0 conda-forge
openh264 2.3.1 hcb278e6_2 conda-forge
openjpeg 2.5.0 h7d73246_1 conda-forge
openssl 3.1.0 hd590300_3 conda-forge
orc 1.8.3 hfdbbad2_0 conda-forge
p11-kit 0.24.1 hc5aa10d_0 conda-forge
packaging 23.0 pyhd8ed1ab_0 conda-forge
palettable 3.3.0 py_0 conda-forge
pandas 1.5.3 py310h9b08913_1 conda-forge
parquet-cpp 1.5.1 2 conda-forge
parso 0.8.3 pyhd8ed1ab_0 conda-forge
patsy 0.5.3 pyhd8ed1ab_0 conda-forge
pcre2 10.40 hc3806b6_0 conda-forge
pexpect 4.8.0 pyh1a96a4e_2 conda-forge
phik 0.12.3 py310h7270e96_0 conda-forge
pickleshare 0.7.5 py_1003 conda-forge
pillow 9.2.0 py310h454ad03_3 conda-forge
pip 23.0.1 pyhd8ed1ab_0 conda-forge
pipx 1.2.0 pyhd8ed1ab_0 conda-forge
pixman 0.40.0 h36c2ea0_0 conda-forge
platformdirs 3.2.0 pyhd8ed1ab_0 conda-forge
plotnine 0.10.1 pyhd8ed1ab_2 conda-forge
pluggy 1.0.0 pyhd8ed1ab_5 conda-forge
polars 0.17.13 py310hcb5633a_0 conda-forge
pooch 1.7.0 pyha770c72_3 conda-forge
proj 9.1.0 h93bde94_0 conda-forge
prompt-toolkit 3.0.38 pyha770c72_0 conda-forge
prompt_toolkit 3.0.38 hd8ed1ab_0 conda-forge
psutil 5.9.5 py310h1fa729e_0 conda-forge
pthread-stubs 0.4 h36c2ea0_1001 conda-forge
ptyprocess 0.7.0 pyhd3deb0d_0 conda-forge
pugixml 1.11.4 h9c3ff4c_0 conda-forge
pulseaudio 16.1 hcb278e6_3 conda-forge
pulseaudio-client 16.1 h5195f5e_3 conda-forge
pulseaudio-daemon 16.1 ha8d29e2_3 conda-forge
pure_eval 0.2.2 pyhd8ed1ab_0 conda-forge
pyarrow 11.0.0 py310h633f555_13_cpu conda-forge
pybind11-abi 4 hd8ed1ab_3 conda-forge
pycosat 0.6.4 py310h5764c6d_1 conda-forge
pycparser 2.21 pyhd8ed1ab_0 conda-forge
pydantic 1.10.7 py310h1fa729e_0 conda-forge
pygments 2.14.0 pyhd8ed1ab_0 conda-forge
pyopenssl 23.1.1 pyhd8ed1ab_0 conda-forge
pyparsing 3.0.9 pyhd8ed1ab_0 conda-forge
pysocks 1.7.1 pyha2e5f31_6 conda-forge
python 3.10.10 he550d4f_0_cpython conda-forge
python-dateutil 2.8.2 pyhd8ed1ab_0 conda-forge
python-tzdata 2023.3 pyhd8ed1ab_0 conda-forge
python_abi 3.10 3_cp310 conda-forge
pytz 2023.3 pyhd8ed1ab_0 conda-forge
pywavelets 1.4.1 py310h0a54255_0 conda-forge
pyyaml 6.0 py310h5764c6d_5 conda-forge
pyzmq 25.0.2 py310h059b190_0 conda-forge
qt-main 5.15.8 h5d23da1_6 conda-forge
re2 2023.02.02 hcb278e6_0 conda-forge
readline 8.2 h8228510_1 conda-forge
reproc 14.2.4 h0b41bf4_0 conda-forge
reproc-cpp 14.2.4 hcb278e6_0 conda-forge
requests 2.28.2 pyhd8ed1ab_1 conda-forge
ruamel.yaml 0.17.21 py310h1fa729e_3 conda-forge
ruamel.yaml.clib 0.2.7 py310h1fa729e_1 conda-forge
s2n 1.3.41 h3358134_0 conda-forge
scikit-learn 1.2.2 py310h41b6a48_1 conda-forge
scipy 1.9.3 py310hdfbd76f_2 conda-forge
seaborn-base 0.12.2 pyhd8ed1ab_0 conda-forge
setuptools 67.6.1 pyhd8ed1ab_0 conda-forge
simpleitk 2.2.1 py310h2b9ea3a_1 conda-forge
six 1.16.0 pyh6c4a22f_0 conda-forge
snappy 1.1.10 h9fff704_0 conda-forge
sqlite 3.40.0 h4ff8645_0 conda-forge
stack_data 0.6.2 pyhd8ed1ab_0 conda-forge
statadict 1.1.0 pypi_0 pypi
statsmodels 0.13.5 py310hde88566_2 conda-forge
svt-av1 1.4.1 hcb278e6_0 conda-forge
sweetviz 2.1.4 pyhd8ed1ab_0 conda-forge
tangled-up-in-unicode 0.2.0 pyhd8ed1ab_0 conda-forge
tbb 2021.8.0 hf52228f_0 conda-forge
tbb-devel 2021.8.0 hf52228f_0 conda-forge
threadpoolctl 3.1.0 pyh8a188c0_0 conda-forge
tk 8.6.12 h27826a3_0 conda-forge
toolz 0.12.0 pyhd8ed1ab_0 conda-forge
tornado 6.3.2 py310h2372a71_0 conda-forge
tqdm 4.64.1 pyhd8ed1ab_0 conda-forge
traitlets 5.9.0 pyhd8ed1ab_0 conda-forge
typeguard 2.13.3 pyhd8ed1ab_0 conda-forge
typing-extensions 4.5.0 hd8ed1ab_0 conda-forge
typing_extensions 4.5.0 pyha770c72_0 conda-forge
tzdata 2023c h71feb2d_0 conda-forge
ucx 1.14.0 ha0ee010_0 conda-forge
unicodedata2 15.0.0 py310h5764c6d_0 conda-forge
urllib3 1.26.15 pyhd8ed1ab_0 conda-forge
userpath 1.7.0 pyhd8ed1ab_0 conda-forge
utfcpp 3.2.3 ha770c72_0 conda-forge
visions 0.7.5 pyhd8ed1ab_0 conda-forge
vtk 9.2.5 qt_py310hc895abb_200 conda-forge
wcwidth 0.2.6 pyhd8ed1ab_0 conda-forge
wheel 0.40.0 pyhd8ed1ab_0 conda-forge
widgetsnbextension 4.0.7 pyhd8ed1ab_0 conda-forge
wslink 1.10.1 pyhd8ed1ab_0 conda-forge
x264 1!164.3095 h166bdaf_2 conda-forge
x265 3.5 h924138e_3 conda-forge
xcb-util 0.4.0 h166bdaf_0 conda-forge
xcb-util-image 0.4.0 h166bdaf_0 conda-forge
xcb-util-keysyms 0.4.0 h166bdaf_0 conda-forge
xcb-util-renderutil 0.3.9 h166bdaf_0 conda-forge
xcb-util-wm 0.4.1 h166bdaf_0 conda-forge
xkeyboard-config 2.38 h0b41bf4_0 conda-forge
xorg-fixesproto 5.0 h7f98852_1002 conda-forge
xorg-kbproto 1.0.7 h7f98852_1002 conda-forge
xorg-libice 1.0.10 h7f98852_0 conda-forge
xorg-libsm 1.2.3 hd9c2040_1000 conda-forge
xorg-libx11 1.8.4 h0b41bf4_0 conda-forge
xorg-libxau 1.0.9 h7f98852_0 conda-forge
xorg-libxdmcp 1.1.3 h7f98852_0 conda-forge
xorg-libxext 1.3.4 h0b41bf4_2 conda-forge
xorg-libxfixes 5.0.3 h7f98852_1004 conda-forge
xorg-libxrender 0.9.10 h7f98852_1003 conda-forge
xorg-libxt 1.2.1 h7f98852_2 conda-forge
xorg-renderproto 0.11.1 h7f98852_1002 conda-forge
xorg-xextproto 7.3.0 h0b41bf4_1003 conda-forge
xorg-xproto 7.0.31 h7f98852_1007 conda-forge
xz 5.2.6 h166bdaf_0 conda-forge
yaml 0.2.5 h7f98852_2 conda-forge
yaml-cpp 0.7.0 h27087fc_2 conda-forge
yarl 1.8.2 py310h5764c6d_0 conda-forge
ydata-profiling 4.1.2 pyhd8ed1ab_0 conda-forge
zeromq 4.3.4 h9c3ff4c_1 conda-forge
zipp 3.15.0 pyhd8ed1ab_0 conda-forge
zlib 1.2.13 h166bdaf_4 conda-forge
zstandard 0.19.0 py310hdeb6495_1 conda-forge
zstd 1.5.2 h3eb15da_6 conda-forge
```
### OS
Ubuntu 22.04
### Checklist
- [X] There is not yet another bug report for this issue in the [issue tracker](https://github.com/ydataai/pandas-profiling/issues)
- [X] The problem is reproducible from this bug report. [This guide](http://matthewrocklin.com/blog/work/2018/02/28/minimal-bug-reports) can help to craft a minimal bug report.
- [X] The issue has not been resolved by the entries listed under [Common Issues](https://pandas-profiling.ydata.ai/docs/master/pages/support_contrib/common_issues.html). | open | 2023-05-18T14:10:43Z | 2024-10-09T05:47:44Z | https://github.com/ydataai/ydata-profiling/issues/1331 | [
"bug 🐛",
"getting started ☝"
] | gdevenyi | 1 |
vchaptsev/cookiecutter-django-vue | graphql | 60 | Update to instructions | Just a couple helpful updates to the instructions:
1. Add `pip install autopep8` if you don't have it installed.
1. run `npm i && npm run lint --fix` from the **frontend** directory. | open | 2020-09-25T20:57:06Z | 2020-11-11T14:27:35Z | https://github.com/vchaptsev/cookiecutter-django-vue/issues/60 | [] | ndunn219 | 2 |
modoboa/modoboa | django | 2,200 | error 451 4.3.5 | hi, i have new error without any reasson .. maybe restart or update any package .. but now have this :/
debian 10, maria 10.5 , nginx mainline, --
Mar 18 16:56:16 mail postfix/postscreen[20786]: CONNECT from [127.0.0.1]:57479 to [127.0.0.1]:25
Mar 18 16:56:16 mail postfix/postscreen[20786]: WHITELISTED [127.0.0.1]:57479
Mar 18 16:56:16 mail postfix/smtpd[20787]: connect from localhost[127.0.0.1]
Mar 18 16:56:17 mail postfix/smtpd[20787]: warning: problem talking to server 127.0.0.1:9999: Success
Mar 18 16:56:17 mail postfix/smtpd[20787]: NOQUEUE: reject: RCPT from localhost[127.0.0.1]: 451 4.3.5 <user@domain2.eu>: Recipient address rejected: Server configuration problem; from=<user@domain1.net> to=<user@domain2.eu> proto=ESMTP helo=<email.domain1.net>
Mar 18 16:56:17 mail postfix/smtpd[20787]: disconnect from localhost[127.0.0.1] ehlo=1 auth=1 mail=1 rcpt=0/1 rset=1 quit=1 commands=5/6
thanks a lot ! | closed | 2021-03-18T16:09:19Z | 2022-05-07T07:12:16Z | https://github.com/modoboa/modoboa/issues/2200 | [] | CHazz | 24 |
huggingface/diffusers | pytorch | 10,680 | stabilityai/stable-diffusion-2-1-base is missing diffusion_pytorch_model.fp16.bin | Got this warning on my console
```
stabilityai/stable-diffusion-2-1-base is missing diffusion_pytorch_model.fp16.bin
```
Was asked to raise this issue, can you please upload the necessary checkpoints in the hugging face repo? | closed | 2025-01-29T14:35:31Z | 2025-01-30T20:52:19Z | https://github.com/huggingface/diffusers/issues/10680 | [] | rohit901 | 5 |
gradio-app/gradio | machine-learning | 10,821 | Provide the CSS style names for components. | Gradio is an excellent and convenient project for deploying model frontends. I really like it and have been using it for a long time. However, there is one issue that has troubled me for a while. I want to make this project look better, but finding the style names of each component to modify many parameters is extremely difficult and troublesome. Additionally, there are many inline styles for which I can't even find the style names (although this might be due to my oversight). Therefore, I sincerely hope that in the documentation for each component, at the bottom or somewhere, the CSS section for that component could be provided. Even just a small portion would be very helpful. Thank you very much. | open | 2025-03-17T19:01:43Z | 2025-03-17T19:34:34Z | https://github.com/gradio-app/gradio/issues/10821 | [
"enhancement",
"docs/website"
] | MeliodasZHAO | 4 |
QuivrHQ/quivr | api | 3,421 | Better logging | * log all error and exception level in parseable
* log request body
* log response body | closed | 2024-10-23T09:15:29Z | 2025-02-02T00:26:38Z | https://github.com/QuivrHQ/quivr/issues/3421 | [
"Stale"
] | linear[bot] | 2 |
flairNLP/flair | pytorch | 2,751 | Iterating over the cuda-devices fails with a Type-Error | **Describe the bug**
While iterating over the cuda-devices, flair fails to load the device properly. This method of iteration is analogous to the method described in this issue: https://github.com/flairNLP/flair/issues/464
**To Reproduce**
```
import flair
import torch
for i in range(0,torch.cuda.device_count()):
current_device = torch.cuda.device(i)
print(current_device)
flair.device = torch.cuda.device(i)
# Actually use flair, this is cut from the example for brevity
```
**Expected behavior**
That the code snipped changes the flair device properly. Instead, the programm crashes with the following error:
```
<torch.cuda.device object at 0x7f8fd0eee880>
2022-05-07 09:24:41,338 loading file /vol/fob-vol4/mi17/weyaaron/.flair/models/ner-german-large/6b8de9edd73722050be2547acf64c037b2df833c6e8f0e88934de08385e26c1e.4b0797effcc6ebb1889d5d29784b97f0a099c1569b319d87d7c387e44e2bba48
Traceback (most recent call last):
File "/vol/fob-vol4/mi17/weyaaron/Zitatsuchmaschine/main.py", line 54, in <module>
main()
File "/vol/fob-vol4/mi17/weyaaron/Zitatsuchmaschine/main.py", line 50, in main
active_mode.execute_full_cycle()
File "/vol/fob-vol4/mi17/weyaaron/Zitatsuchmaschine/src/modes/nlp_mode.py", line 14, in execute_full_cycle
self.setup()
File "/vol/fob-vol4/mi17/weyaaron/Zitatsuchmaschine/src/modes/impl/extraction_mode.py", line 14, in setup
self.quote_extractor = QuoteExtractor(
File "<string>", line 16, in __init__
File "/vol/fob-vol4/mi17/weyaaron/Zitatsuchmaschine/src/nlp/quote_extraction/quote_extraction.py", line 94, in __post_init__
self.__tagger = MultiTagger.load([self.__ner_model, self.__quote_model, self.__quotee_model])
File "/glusterfs/dfs-gfs-dist/qse-common/miniconda-common/envs/qse-dev-system/lib/python3.9/site-packages/flair/models/sequence_tagger_model.py", line 1330, in load
model = SequenceTagger.load(model_name)
File "/glusterfs/dfs-gfs-dist/qse-common/miniconda-common/envs/qse-dev-system/lib/python3.9/site-packages/flair/nn.py", line 88, in load
state = torch.load(f, map_location='cpu')
File "/glusterfs/dfs-gfs-dist/qse-common/miniconda-common/envs/qse-dev-system/lib/python3.9/site-packages/torch/serialization.py", line 607, in load
return _load(opened_zipfile, map_location, pickle_module, **pickle_load_args)
File "/glusterfs/dfs-gfs-dist/qse-common/miniconda-common/envs/qse-dev-system/lib/python3.9/site-packages/torch/serialization.py", line 882, in _load
result = unpickler.load()
File "/glusterfs/dfs-gfs-dist/qse-common/miniconda-common/envs/qse-dev-system/lib/python3.9/site-packages/flair/embeddings/token.py", line 1284, in __setstate__
embedding = TransformerWordEmbeddings(
File "/glusterfs/dfs-gfs-dist/qse-common/miniconda-common/envs/qse-dev-system/lib/python3.9/site-packages/flair/embeddings/token.py", line 856, in __init__
self.model.to(flair.device)
File "/glusterfs/dfs-gfs-dist/qse-common/miniconda-common/envs/qse-dev-system/lib/python3.9/site-packages/torch/nn/modules/module.py", line 880, in to
device, dtype, non_blocking, convert_to_format = torch._C._nn._parse_to(*args, **kwargs)
TypeError: to() received an invalid combination of arguments - got (device), but expected one of:
* (torch.device device, torch.dtype dtype, bool non_blocking, bool copy, *, torch.memory_format memory_format)
* (torch.dtype dtype, bool non_blocking, bool copy, *, torch.memory_format memory_format)
* (Tensor tensor, bool non_blocking, bool copy, *, torch.memory_format memory_format)
```
**Environment (please complete the following information):**
- Linux
- Version 0.8.0.post1
**Additional context**
This is my naive attempt at iterating over all cuda devices. If there is another approach, let me know.
| closed | 2022-05-07T07:42:31Z | 2022-05-08T06:12:48Z | https://github.com/flairNLP/flair/issues/2751 | [
"bug"
] | Weyaaron | 3 |
SciTools/cartopy | matplotlib | 1,527 | [proposal] adding pm/prime_meridian as an option for Globe? | The PROJ parameter `pm` can support a string or a number.
The prime meridian is used when defining a datum or for handing the `-180/180` issues.
- https://pyproj4.github.io/pyproj/stable/build_crs.html
- https://lists.osgeo.org/pipermail/proj/2020-April/009540.html
So, I propose adding `prime_meridian` as an input parameter to the `Globe` object that maps to the `pm` PROJ parameter.
This would enable:
```python
from cartopy.crs import CRS as cCRS, Globe
from pyproj.crs import CRS
proj_crs = CRS.from_epsg(4326)
globe = Globe(
ellipse=None,
semimajor_axis=proj_crs.ellipsoid.semi_major_metre,
semiminor_axis=proj_crs.ellipsoid.semi_minor_metre,
inverse_flattening=proj_crs.ellipsoid.inverse_flattening,
prime_meridian=proj_crs.prime_meridian.longitude,
)
proj_dict = proj_crs.to_dict()
cart_crs = cCRS(proj_dict, globe=globe)
``` | closed | 2020-04-15T13:48:12Z | 2021-08-31T20:35:20Z | https://github.com/SciTools/cartopy/issues/1527 | [] | snowman2 | 1 |
flaskbb/flaskbb | flask | 368 | Plugin interaction with forms | While writing a plugin to modify the registration/update details forms (see also #367) - I ran into the following problem:
The following (simplified) code works as expected as the change details form works using `form.populate_obj()`
@impl
def flaskbb_additional_setup():
from flaskbb.user.forms import ChangeDetailsForm
from flaskbb.user.models import User
ChangeDetailsForm.plugin_extrafield = TextAreaField("Plugin Stuff")
User.plugin_extrafield = db.Column(db.String(200))
@impl
def flaskbb_tpl_after_user_details_form():
#code to render extra field
however trying to do the same thing with the registration form is made more difficult as the plugin needs to monkeypatch the save method. Not sure what the best way to address this is - a hook just before the user.save() method? | closed | 2017-11-30T16:09:18Z | 2018-04-15T07:47:49Z | https://github.com/flaskbb/flaskbb/issues/368 | [] | djsilcock | 9 |
gevent/gevent | asyncio | 1,356 | Monkey patch raises an error under PyPy2.7-7.0.0 | * gevent version: 1.4.0
* Python version: PyPy2.7 v7.0.0
* Operating System: Ubuntu 18.04
### Description:
PyPy2 now uses a backported version of Python3's CLock, which can't be patched. The following raises an attribute error under PyPy2.7-7:
```python
from gevent import monkey
monkey.patch_all()
```
See https://bitbucket.org/pypy/pypy/issues/2962/gevent-cannot-patch-rlock-under-pypy-27-7 for the underlying PyPy change and a minimal dockerfile reproducing the error. | closed | 2019-02-21T01:37:56Z | 2019-03-27T20:15:00Z | https://github.com/gevent/gevent/issues/1356 | [] | olliemath | 0 |
google-research/bert | nlp | 1,057 | is it necessary to drop stop words before training? | When I use the model for text classification,is it necessary to drop stop words before training?If I did so,will it improve the accuracy?And could you please tell me the reasons?Thanks a lot. | open | 2020-04-14T04:17:08Z | 2020-06-13T08:21:26Z | https://github.com/google-research/bert/issues/1057 | [] | haozheshz | 2 |
ploomber/ploomber | jupyter | 232 | Show appropriate error messages for Python-only features | Like debugging | closed | 2020-08-13T17:56:34Z | 2020-10-03T19:57:36Z | https://github.com/ploomber/ploomber/issues/232 | [] | edublancas | 0 |
python-gino/gino | sqlalchemy | 426 | Retry options in init_app for Sanic | * GINO version: 0.8.1
* Python version: 3.7
* asyncpg version: 0.18.2
* aiocontextvars version:
* PostgreSQL version: 11
### Description
Today I was trying to write a simple app using docker-compose. One service is the database service and another is the web-app written with Sanic and GINO.
When starting both of these, since Postgres takes it's time starting up, I'll get an error from Sanic `ConnectionRefused` etc...
Is there any way or is there any plan to add options for retrying the connection?
### What I Did
Started a Sanic app with Gino using Gino.ext.sanic configured as follow
```
app = Sanic()
app.config.DB_HOST = 'db-service'
app.config.DB_DATABASE = 'vpn_service'
app.config.DB_USER = 'postgres'
app.config.DB_PASSWORD = 'postgres'
db.init_app(app)
```
Get a traceback with a `ConnectionRefused` starting from the `set_bind` inside the `Gino.ext.sanic` module.
| closed | 2019-01-25T23:21:44Z | 2019-02-02T17:19:11Z | https://github.com/python-gino/gino/issues/426 | [
"question"
] | choco | 3 |
plotly/jupyter-dash | dash | 90 | No module named 'jupyter-dash' | I encountered this problem when I was trying to run `from jupyter_dash import JupyterDash` in both jupyter notebook and jupyter lab.
OS: Windows 10
python version: 3.8.13
Installed packages:
Package Version
ansi2html 0.0.0
anyio 3.6.1
appnope 0.1.3
argon2-cffi 21.3.0
argon2-cffi-bindings 21.2.0
asttokens 2.0.5
async-generator 1.10
attrs 21.4.0
Babel 2.10.1
backcall 0.2.0
backports.functools-lru-cache 1.6.4
beautifulsoup4 4.11.1
bleach 5.0.0
Bottleneck 1.3.4
Brotli 1.0.9
brotlipy 0.7.0
certifi 2022.5.18.1
cffi 1.15.0
chardet 4.0.0
charset-normalizer 2.0.12
click 8.1.3
colorama 0.4.4
cryptography 37.0.1
dash 2.4.1
dash-bootstrap-components 1.1.0
dash-core-components 2.0.0
dash-html-components 2.0.0
dash-renderer 1.9.1
dash-table 5.0.0
debugpy 1.6.0
decorator 5.1.1
defusedxml 0.7.1
deprecation 2.1.0
entrypoints 0.4
executing 0.8.3
fastjsonschema 2.15.3
Flask 2.1.2
Flask-Compress 0.0.0
flit_core 3.7.1
future 0.18.2
gunicorn 20.1.0
idna 3.3
importlib-metadata 4.11.4
importlib-resources 5.7.1
ipykernel 6.13.1
ipython 8.4.0
ipython-genutils 0.2.0
itsdangerous 2.1.2
jedi 0.18.1
Jinja2 3.1.2
joblib 1.1.0
json5 0.9.6
jsonschema 4.6.0
jupyter-client 7.3.3
jupyter-core 4.10.0
jupyter-dash 0.4.2
jupyter-packaging 0.12.0
jupyter-server 1.17.0
jupyterlab 3.4.2
jupyterlab-pygments 0.2.2
jupyterlab-server 2.14.0
MarkupSafe 2.1.1
matplotlib-inline 0.1.3
mistune 0.8.4
mkl-fft 1.3.1
mkl-random 1.2.2
mkl-service 2.4.0
nbclassic 0.3.7
nbclient 0.6.4
nbconvert 6.5.0
nbformat 5.4.0
nest-asyncio 1.5.5
notebook 6.4.11
notebook-shim 0.1.0
numexpr 2.8.1
numpy 1.22.3
packaging 21.3
pandas 1.4.2
pandocfilters 1.5.0
parso 0.8.3
pexpect 4.8.0
pickleshare 0.7.5
pip 21.2.2
plotly 5.8.0
prometheus-client 0.14.1
prompt-toolkit 3.0.29
psutil 5.9.1
ptyprocess 0.7.0
pure-eval 0.2.2
pycparser 2.21
Pygments 2.12.0
pyOpenSSL 22.0.0
pyparsing 3.0.9
pyrsistent 0.18.1
PySocks 1.7.1
python-dateutil 2.8.2
pytz 2022.1
pywin32 303
pywinpty 2.0.2
pyzmq 23.1.0
requests 2.27.1
retrying 1.3.3
scikit-learn 1.0.2
scipy 1.7.3
Send2Trash 1.8.0
setuptools 61.2.0
six 1.16.0
sniffio 1.2.0
soupsieve 2.3.1
stack-data 0.2.0
tenacity 8.0.1
terminado 0.15.0
testpath 0.6.0
threadpoolctl 2.2.0
tinycss2 1.1.1
tomlkit 0.11.0
tornado 6.1
traitlets 5.2.2.post1
urllib3 1.26.9
wcwidth 0.2.5
webencodings 0.5.1
websocket-client 1.3.2
Werkzeug 2.1.2
wheel 0.37.1
win-inet-pton 1.1.0
wincertstore 0.2
zipp 3.8.0 | closed | 2022-06-08T21:49:48Z | 2022-06-09T22:17:02Z | https://github.com/plotly/jupyter-dash/issues/90 | [] | Dwingkak | 6 |
allenai/allennlp | nlp | 5,211 | Models loaded using the `from_archive` method need to be saved with original config | When `allennlp train` is used to fine-tune a pretrained model (`model A`) using `from_archive(path_to_A)`, the finetuned model (`model B`) is saved with the config that contains `from_archive`. This means that if you try to now finetune the `model B`, it needs the original `model A` at the exact `path_to_A`, as well as `model B`. In the normal usecase, this will fail if the user does not have access to the original `model A`. On beaker, depending on how the code is setup, if the path to the pretrained model remains the same in `experiment A -> B` and `experiment B -> C`, it will cause a `maximum recursion depth` error.
Potential solution is to store the original configuration when saving a fine-tuned model (i.e., the `from_archive` case). | open | 2021-05-18T19:28:40Z | 2021-05-28T16:33:03Z | https://github.com/allenai/allennlp/issues/5211 | [
"bug"
] | AkshitaB | 1 |
tiangolo/uvicorn-gunicorn-fastapi-docker | pydantic | 26 | Unable to start container | I have the following Dockerfile:
```
FROM tiangolo/uvicorn-gunicorn-fastapi:python3.7
COPY . /app
#COPY ./requirements.txt /app
#
#run pip install --upgrade pip && \
# pip install -r /app/requirements.txt
ENV APP_NAME "control.controller:app"
```
I have a folder `control` and a `controller.py` in it.
But my control app is not started. It starts the simple `main.py` from this repo instead.
Any ideas? | closed | 2020-02-05T22:01:14Z | 2020-04-13T13:36:04Z | https://github.com/tiangolo/uvicorn-gunicorn-fastapi-docker/issues/26 | [] | heitorPB | 3 |
matplotlib/matplotlib | data-visualization | 29,736 | [ENH]: ConnectionPatch's connection line arrow size and position issue | ### Problem
I want to mimic a graph to create a global traffic flow map using matplotlib
The graph has a feature where the arrows of the connecting lines are close around the circle.

In order to reproduce this image, I use matplotlib to create scatters and sizes of countries and use ConnectionPatch to create connection lines.
But I can't control the size of the arrows to stay the same and not be covered by the circle, and I can't do it even after consulting all the parameters.
```
connection = ConnectionPatch(
xyA=(x1, y1),
xyB=(x2, y2),
coordsA='data',
coordsB='data',
connectionstyle='arc3,rad=0.3',
linewidth=linewidth,
arrowstyle='->',
color='#2A5079',
zorder=2,
transform=ccrs.PlateCarree()
)
ax.add_artist(connection)
```

### Proposed solution
I'm wondering if it's possible to set the parameter to control the arrow to be a consistent size, with the arrow fitting immediately outside the circle of the scatter. If this is not possible, consider whether the position of the arrow can be set in the middle of the connecting line.
 | open | 2025-03-12T02:16:37Z | 2025-03-20T03:18:24Z | https://github.com/matplotlib/matplotlib/issues/29736 | [
"New feature",
"Community support"
] | Curallin | 9 |
mars-project/mars | scikit-learn | 2,907 | [BUG] Mars serialization took too much time | <!--
Thank you for your contribution!
Please review https://github.com/mars-project/mars/blob/master/CONTRIBUTING.rst before opening an issue.
-->
**Describe the bug**
When executing following task with 3000 parallelism, mars took 2.5h to finished using 55 workers each has 9 subpools. But the serialization of supervisor took 2.2h in it:
```python
df = md.DataFrame(
mt.random.RandomState(0).rand(3000_000, 5, chunk_size=1000),
columns=list('abcde'))
df.groupby(['a', 'b']).apply(lambda df: [1,2]).reset_index().execute()
```
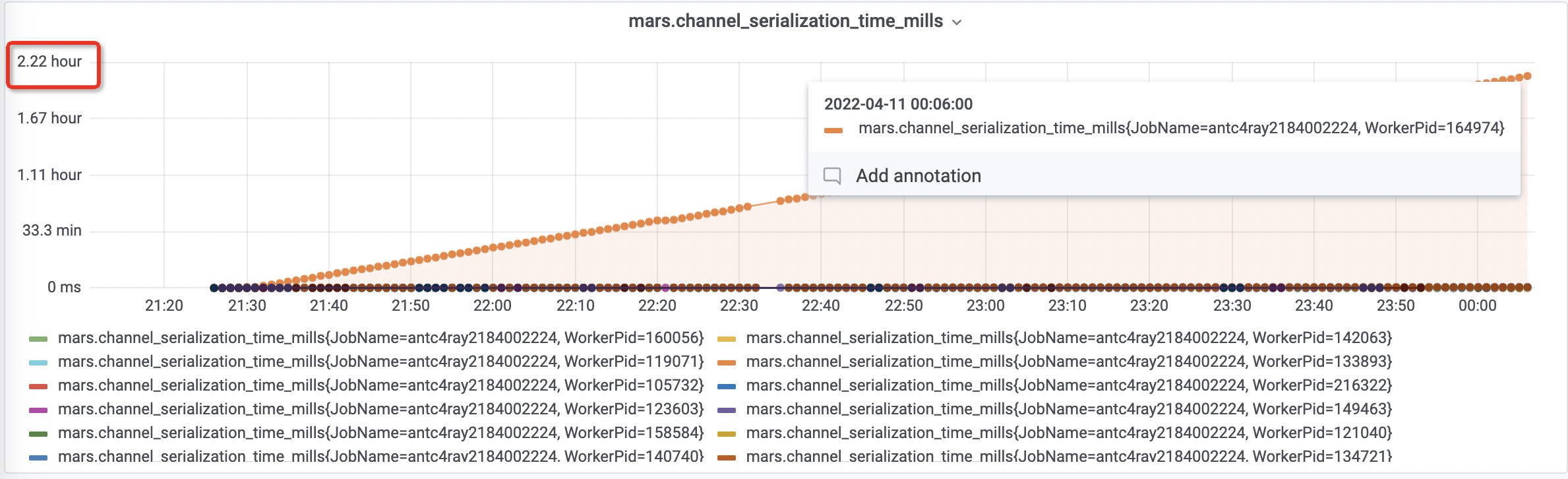

**To Reproduce**
To help us reproducing this bug, please provide information below:
1. Your Python version: python 3.7.9
2. The version of Mars you use: master
3. Versions of crucial packages, such as numpy, scipy and pandas
4. Full stack of the error.
5. Minimized code to reproduce the error.
```python
df = md.DataFrame(
mt.random.RandomState(0).rand(3000_000, 5, chunk_size=1000),
columns=list('abcde'))
df.groupby(['a', 'b']).apply(lambda df: [1, 2]).reset_index().execute()
```
**Expected behavior**
We need to:
* Find out what is being sent to the supervisor
* reduce the data sent to mars supervisor
* Speed up mars serialization performance.
**Additional context**
Add any other context about the problem here.
| closed | 2022-04-11T03:02:14Z | 2022-04-15T10:55:32Z | https://github.com/mars-project/mars/issues/2907 | [] | chaokunyang | 11 |
GibbsConsulting/django-plotly-dash | plotly | 431 | Question may I integrate djangodash with dash-leaflet framework or assing function from from dash_extensions.javascript import assign | Thank You for DjangoDash , i will use it, and i like it.
Today i need integrate openstreet map , for this i try use dash-leaflet from documentation https://dash-leaflet-docs.onrender.com/
i will try start example with geojson and change icon markers
============================== Example from site https://dash-leaflet-docs.onrender.com/ =======================
import dash_leaflet as dl
import dash_leaflet.express as dlx
from dash import Dash, html
from dash_extensions.javascript import assign
# A few countries.
countries = [dict(name="Denmark", iso2="dk", lat=56.26392, lon=9.501785),
dict(name="Sweden", iso2="se", lat=59.334591, lon=18.063240),
dict(name="Norway", iso2="no", lat=59.911491, lon=9.501785)]
# Generate geojson with a marker for each country and name as tooltip.
geojson = dlx.dicts_to_geojson([{**c, **dict(tooltip=c['name'])} for c in countries])
# Create javascript function that draws a marker with a custom icon, in this case a flag hosted by flagcdn.
draw_flag = assign("""function(feature, latlng){
const flag = L.icon({iconUrl: `https://flagcdn.com/64x48/${feature.properties.iso2}.png`, iconSize: [64, 48]});
return L.marker(latlng, {icon: flag});
}""")
# Create example app.
app = Dash()
app.layout = html.Div([
dl.Map(children=[
dl.TileLayer(), dl.GeoJSON(data=geojson, options=dict(pointToLayer=draw_flag), zoomToBounds=True)
], style={'width': '100%', 'height': '50vh', 'margin': "auto", "display": "block"}, id="map"),
])
if __name__ == '__main__':
app.run_server()
================================= End Example ==============================================
instead of app = Dash() i use app = DjangoDash('Graph', external_stylesheets=[dbc.themes.BOOTSTRAP])
as a result i see in browser map without markers and i get Error
No match for [dashExtensions.default.function0] in the global window object ( dash_renderer.min.js:2)
If the way to make friends djangodash with use javascript from assing operator ?

Thank for the answer | open | 2022-12-04T17:52:21Z | 2022-12-05T19:52:40Z | https://github.com/GibbsConsulting/django-plotly-dash/issues/431 | [] | Nizhurin | 1 |
kymatio/kymatio | numpy | 789 | `sigma0` should be `0.13` | AFAIK 0.1 is a heuristic which aims to allow subsampling by `T` by setting `sigma=0.1/T`. It is, however, overly conservative, according to `criterion_amplitude=1e-3`:
```python
from kymatio.scattering1d.filter_bank import gauss_1d, compute_temporal_support
T = 128
phi = gauss_1d(16384, 0.1 / T)
f_at_subsampling = len(phi)//2//T
f_halfsupport = compute_temporal_support(ifft(phi), criterion_amplitude=1e-3)
print(f_halfsupport, f_at_subsampling)
```
```
49 64
```
vs `0.13 / T`:
```
63 64
```
`criterion_amplitude` conditions the extent of decay upon which boundary effects, i.e. tail contributions, are negligible. Hence, if contributions past `1e-3` are deemed negligible, then we have ~lossless subsampling - and `0.1` obtaining `49` means we subsample much less (by 30%) than we safely could.
It's a simple yet significant change by which the wavelets would also be affected (since `J` is also defined per `sigma0`) and will require changing some tests and maybe docs, so I leave the idea here for now. | closed | 2022-01-01T23:21:29Z | 2022-05-30T15:16:03Z | https://github.com/kymatio/kymatio/issues/789 | [] | OverLordGoldDragon | 12 |
flasgger/flasgger | api | 176 | How to fetch next result "idStatus=1&idStatus=3" | I have next yml file
```
Return list of foo
---
tags:
- "Foo"
summary: "Return list of foo"
description: ""
produces:
- "application/json"
parameters:
- in: "query"
name: "limit"
type: "integer"
description: ""
default: 10
- in: "query"
name: "page"
type: "integer"
description: ""
default: 1
- in: "query"
name: "idStatus"
description: ""
type: "array"
items:
type: "integer"
style: "form"
explode: true
``` | closed | 2018-01-22T12:37:15Z | 2018-01-24T13:37:55Z | https://github.com/flasgger/flasgger/issues/176 | [
"question"
] | Kvasela | 1 |
serengil/deepface | deep-learning | 502 | "AttributeError: 'NoneType' object has no attribute 'copy'" | I get this error whenever I try to use DeepFace.find on a folder full of pictures.
Relevant folder structure:
F:\Pictures\jpg\ (images with the names "db (1).jpg" through "db (13713).jpg")
D:\Work\known\ (find.jpg)
Here's my code:
"""
from deepface import DeepFace
import os
#The path containing the face(s) I want too search
known_path = "D:\Work\known"
#Making a list containing the path to all pictures in a certain folder to search through
pictures = []
for (root, dirs, known) in os.walk(known_path): pass
for i in known:
pictures.append("D://Work//known//" + i)
#The actual searching
df = DeepFace.find(img_path = pictures, db_path = "F:\Pictures\jpg", enforce_detection=(False), model_name = "Facenet512", detector_backend = "ssd")
"""
Here's the full error:
"""
Traceback (most recent call last):
File "C:\Users\----\Desktop\Programming\DeepFace.py", line 14, in <module>
df = DeepFace.find(img_path = pictures, db_path = "F:\Pictures\jpg", enforce_detection=(False), model_name = "Facenet512", detector_backend = "ssd")
File "C:\Users\----\anaconda3\envs\face_recognition\lib\site-packages\deepface\DeepFace.py", line 581, in find
representation = represent(img_path = employee
File "C:\Users\----\anaconda3\envs\face_recognition\lib\site-packages\deepface\DeepFace.py", line 754, in represent
img = functions.preprocess_face(img = img_path
File "C:\Users\----\anaconda3\envs\face_recognition\lib\site-packages\deepface\commons\functions.py", line 176, in preprocess_face
base_img = img.copy()
AttributeError: 'NoneType' object has no attribute 'copy'
"""
Python version 3.10.5, using an Anaconda environment. The only packages installed are DeepFace (and all dependencies) and face_recognition (and all dependencies).
My issue gives the same error as issue #13 and issue #206. However, I don't think either solution is relevant to me, as there are no non-English characters in my path (like issue #13) and the solution to issue #206 should already be implemented(?). Also, both other issue's occurred when using DeepFace.stream, which I am not doing
Also, thanks for creating this program and making it available for free to everyone. I've had some other (successful) tests with this program, and it worked perfectly every time, with good performance and clear syntax. | closed | 2022-07-04T13:33:08Z | 2022-07-07T07:58:53Z | https://github.com/serengil/deepface/issues/502 | [
"question"
] | maxyvisser | 5 |
horovod/horovod | machine-learning | 3,952 | Horovod stack trace from Signal 7 | **Environment:**
1. Framework: TensorFlow
2. Framework version: 2.12.0
3. Horovod version: 0.28.1
4. MPI version: 4.1.4-3 (openmpi40-aws)
5. CUDA version: 11.8
6. NCCL version: 2.16.5-1+cuda11.8
7. Python version: 3.10
8. Spark / PySpark version:
10. Ray version:
11. OS and version: Ubuntu 20.04
12. GCC version: 9.4.0
13. CMake version: 3.26.0
**Checklist:**
1. Did you search issues to find if somebody asked this question before? Yes, i searched issues
2. If your question is about hang, did you read [this doc](https://github.com/horovod/horovod/blob/master/docs/running.rst)? It is not about a hang.
3. If your question is about docker, did you read [this doc](https://github.com/horovod/horovod/blob/master/docs/docker.rst)? Not it is not about docker.
4. Did you check if you question is answered in the [troubleshooting guide] (https://github.com/horovod/horovod/blob/master/docs/troubleshooting.rst)? Yes
**Bug report:**
```
Tue Jun 27 22:49:58 2023[1,3]<stderr>:*** Received signal 7 ***
Tue Jun 27 22:49:58 2023[1,3]<stderr>:*** BEGIN MANGLED STACK TRACE ***
Tue Jun 27 22:49:58 2023[1,4]<stderr>:/usr/local/lib/python3.10/site-packages/tensorflow/python/platform/../../libtensorflow_framework.so.2(+0x1793d31)[0x7fbf52976d31]
Tue Jun 27 22:49:58 2023[1,3]<stderr>:/usr/local/lib/python3.10/site-packages/tensorflow/python/platform/../../libtensorflow_framework.so.2(+0x1793d31)[0x7fbe6c106d31]
Tue Jun 27 22:49:58 2023[1,3]<stderr>:/usr/lib/x86_64-linux-gnu/libc.so.6(+0x43090)[0x7fbf20ce4090]
Tue Jun 27 22:49:58 2023[1,4]<stderr>:/usr/lib/x86_64-linux-gnu/libc.so.6(+0x43090)[0x7fc007554090]
Tue Jun 27 22:49:58 2023[1,4]<stderr>:/usr/lib/x86_64-linux-gnu/libc.so.6(+0x18bb41)[0x7fc00769cb41]
Tue Jun 27 22:49:58 2023[1,3]<stderr>:/usr/lib/x86_64-linux-gnu/libc.so.6(+0x18bb41)[0x7fbf20e2cb41]
Tue Jun 27 22:49:58 2023[1,3]<stderr>:/usr/local/lib/python3.10/site-packages/horovod/tensorflow/mpi_lib.cpython-310-x86_64-linux-gnu.so(+0x224e35)[0x7fbd556c4e35]
Tue Jun 27 22:49:58 2023[1,3]<stderr>:/usr/local/lib/python3.10/site-packages/horovod/tensorflow/mpi_lib.cpython-310-x86_64-linux-gnu.so(+0x21b6ab)[0x7fbd556bb6ab]
Tue Jun 27 22:49:58 2023[1,3]<stderr>:/usr/lib/x86_64-linux-gnu/libpthread.so.0(+0x8609)[0x7fbf20c86609]
Tue Jun 27 22:49:58 2023[1,4]<stderr>:/usr/local/lib/python3.10/site-packages/horovod/tensorflow/mpi_lib.cpython-310-x86_64-linux-gnu.so(+0x224e35)[0x7fbe3bf34e35]
Tue Jun 27 22:49:58 2023[1,4]<stderr>:/usr/local/lib/python3.10/site-packages/horovod/tensorflow/mpi_lib.cpython-310-x86_64-linux-gnu.so(+0x21b6ab)[0x7fbe3bf2b6ab]
Tue Jun 27 22:49:58 2023[1,4]<stderr>:/usr/lib/x86_64-linux-gnu/libpthread.so.0(+0x8609)[0x7fc0074f6609]
Tue Jun 27 22:49:58 2023[1,3]<stderr>:/usr/lib/x86_64-linux-gnu/libc.so.6(clone+0x43)[0x7fbf20dc0133]
Tue Jun 27 22:49:58 2023[1,3]<stderr>:*** END MANGLED STACK TRACE ***
```
Please describe erroneous behavior you're observing and steps to reproduce it.
The behavior is unpredictable. Training can run for multiple epochs before stack trace shown above. This bug does not show up when training on a single EC2 `p3dn.24xlarge`. It only shows up when training on multiple nodes.
| open | 2023-06-27T23:26:38Z | 2023-06-27T23:26:38Z | https://github.com/horovod/horovod/issues/3952 | [
"bug"
] | ajayvohra2005 | 0 |
koxudaxi/fastapi-code-generator | pydantic | 297 | Array of array loses its constraints | An array of arrays gets translated into List[List[T]], even when the second array has contraints, losing these constraints. Minimal working example :
```json
{
"components": {
"schemas": {
"Mod": {
"type": "object",
"properties": {
"prop": {
"type": "array",
"items": {
"type": "array",
"items": {
"type": "number",
},
"minItems": 3,
"maxItems": 6
}
}
},
}
}
}
}
```
I see 2 solutions to that :
1 - Use a conlist to keep the contraints. Albeit compact and easy (only a few lines to add around datamodel-code-generator/types.py L87,L254 and datamodel-code-generator/parser/jsonschema.py L772), it would stand out quite a bit, compared to the rest (the conX objects being alone in their own Item model).
2 - Handle constrained array the same way object is handled: by giving it its own Model, which would allow to store the constraints in Field, as it is done for a single array. | open | 2022-11-23T20:29:55Z | 2022-11-23T20:29:55Z | https://github.com/koxudaxi/fastapi-code-generator/issues/297 | [] | Aedial | 0 |
zappa/Zappa | flask | 906 | [Migrated] cannot be assumed by principal 'events.amazonaws.com', when using events | Originally from: https://github.com/Miserlou/Zappa/issues/2168 by [saeedesmaili](https://github.com/saeedesmaili)
I have a flask project and it is deployed to the AWS Lambda using the Zappa, and it works fine. I'm trying to add an event in the `zappa_settings.json` to run some function regularly. The settings config that was working (without events) was:
```
{
"dev": {
"app_function": "app.app",
"profile_name": "default",
"project_name": "contactclipper2",
"runtime": "python3.8",
"s3_bucket": "zappa-i4hsr8rya",
"aws_region": "us-west-2",
"keep_warm": false,
"use_precompiled_packages": false,
"memory_size": 3008
}
}
```
and I added these two lines, so the settings changed to:
```
{
"dev": {
"app_function": "app.app",
"profile_name": "default",
"project_name": "contactclipper2",
"runtime": "python3.8",
"s3_bucket": "zappa-i4hsr8rya",
"aws_region": "us-west-2",
"keep_warm": false,
"use_precompiled_packages": false,
"memory_size": 3008,
"events": [{
"function": "alerts.test_alert",
"expression": "rate(1 minute)"
}]
}
}
```
But now I can't update or schedule the project and I get this error:
```
botocore.exceptions.ClientError: An error occurred (ValidationException) when calling the PutRule operation: Provided role 'arn:aws:iam::199151782709:role/contactclipper2-dev-ZappaLambdaExecutionRole' cannot be assumed by principal 'events.amazonaws.com'.
```
This is the role's trust entities:

What should I do to fix this and have a working event (cron job)? | closed | 2021-02-20T13:03:36Z | 2024-04-13T19:36:29Z | https://github.com/zappa/Zappa/issues/906 | [
"no-activity",
"auto-closed"
] | jneves | 2 |
databricks/spark-sklearn | scikit-learn | 81 | ImportError: Module not found with Azure Spark Cluster | I'm trying to run spark-sklearns GridSearch on an HDInsight Cluster from Azure. Here is a Code Snippet:
```
model = KerasRegressor(build_fn=build_model, verbose=0)
kf = KFold(n_splits=self.cv_split, shuffle=True) # Cross validation with k=5
sc = SparkContext.getOrCreate()
grid = GridSearchCV(sc=sc, estimator=model, param_grid=self.params,
cv=kf, return_train_score=True, verbose=2,
fit_params={'epochs': nb_epoch, 'batch_size': 32})
hist = grid.fit(x_train, y_train)
```
It works fine until I call the grid.fit method, which returns the following exception:
```
File "/mnt/resource/hadoop/yarn/local/usercache/nsusshuser/appcache/application_1526397916826_0020/container_e01_1526397916826_0020_01_000001/py4j-0.10.4-src.zip/py4j/protocol.py", line 319, in get_return_value
format(target_id, ".", name), value)
py4j.protocol.Py4JJavaError: An error occurred while calling z:org.apache.spark.api.python.PythonRDD.collectAndServe.
: org.apache.spark.SparkException: Job aborted due to stage failure: Task 2 in stage 0.0 failed 4 times, most recent failure: Lost task 2.3 in stage 0.0 (TID 13, wn0-bt-nsu.kkatsjzvwzuephdjshji40kxae.ax.internal.cloudapp.net, executor 1): org.apache.spark.api.python.PythonException: Traceback (most recent call last):
File "/mnt/resource/hadoop/yarn/local/usercache/nsusshuser/appcache/application_1526397916826_0020/container_e01_1526397916826_0020_01_000002/pyspark.zip/pyspark/worker.py", line 166, in main
func, profiler, deserializer, serializer = read_command(pickleSer, infile)
File "/mnt/resource/hadoop/yarn/local/usercache/nsusshuser/appcache/application_1526397916826_0020/container_e01_1526397916826_0020_01_000002/pyspark.zip/pyspark/worker.py", line 55, in read_command
command = serializer._read_with_length(file)
File "/mnt/resource/hadoop/yarn/local/usercache/nsusshuser/appcache/application_1526397916826_0020/container_e01_1526397916826_0020_01_000002/pyspark.zip/pyspark/serializers.py", line 169, in _read_with_length
return self.loads(obj)
File "/mnt/resource/hadoop/yarn/local/usercache/nsusshuser/appcache/application_1526397916826_0020/container_e01_1526397916826_0020_01_000002/pyspark.zip/pyspark/serializers.py", line 455, in loads
return pickle.loads(obj, encoding=encoding)
ImportError: No module named 'ml'
...
Caused by: org.apache.spark.api.python.PythonException: Traceback (most recent call last):
File "/mnt/resource/hadoop/yarn/local/usercache/nsusshuser/appcache/application_1526397916826_0020/container_e01_1526397916826_0020_01_000002/pyspark.zip/pyspark/worker.py", line 166, in main
func, profiler, deserializer, serializer = read_command(pickleSer, infile)
File "/mnt/resource/hadoop/yarn/local/usercache/nsusshuser/appcache/application_1526397916826_0020/container_e01_1526397916826_0020_01_000002/pyspark.zip/pyspark/worker.py", line 55, in read_command
command = serializer._read_with_length(file)
File "/mnt/resource/hadoop/yarn/local/usercache/nsusshuser/appcache/application_1526397916826_0020/container_e01_1526397916826_0020_01_000002/pyspark.zip/pyspark/serializers.py", line 169, in _read_with_length
return self.loads(obj)
File "/mnt/resource/hadoop/yarn/local/usercache/nsusshuser/appcache/application_1526397916826_0020/container_e01_1526397916826_0020_01_000002/pyspark.zip/pyspark/serializers.py", line 455, in loads
return pickle.loads(obj, encoding=encoding)
ImportError: No module named 'ml'
```
The ml module is part of our project. I checked sys.modules and it is in there. Don't really understand the error message. Can somebody help me out? | closed | 2018-05-25T13:34:37Z | 2018-12-08T19:59:41Z | https://github.com/databricks/spark-sklearn/issues/81 | [] | Nimi42 | 1 |
WZMIAOMIAO/deep-learning-for-image-processing | pytorch | 847 | faster-RCNN进行批量推理 | 需要在predict.py文件里将predictions = model(img.to(device))[0]中的img替换成list[torch.tensor]吗,但是这样就没有.to方法了。按照作者在其他问题下的解决方案,我测试了
``image_tensors = [img, img] ``
``batch = torch.stack(image_tensors)``
``predictions = model(batch .to(device))[0]``
有报错transform.py", line 244, in forward
raise ValueError("images is expected to be a list of 3d tensors "
ValueError: images is expected to be a list of 3d tensors of shape [C, H, W], got torch.Size([1, 3, 1080, 1920])
请问如何修改predict.py文件可以支持多个img同时推理呢? | closed | 2024-12-23T08:38:17Z | 2024-12-23T08:51:49Z | https://github.com/WZMIAOMIAO/deep-learning-for-image-processing/issues/847 | [] | Smartog | 2 |
supabase/supabase-py | fastapi | 405 | rpc() - value is not a valid list | **Describe the bug**
`rpc()` crashes (calling a db function) with following error:
> ./tests/test_claims.py::test_get_claims Failed: [undefined]postgrest.exceptions.APIError: {'provider': 'email', 'providers': ['email'], 'claims_admin': True}
> self = <postgrest._sync.request_builder.SyncFilterRequestBuilder object at 0x103d5a680>
>
> def execute(self) -> APIResponse:
> """Execute the query.
>
> .. tip::
> This is the last method called, after the query is built.
>
> Returns:
> :class:`APIResponse`
>
> Raises:
> :class:`APIError` If the API raised an error.
> """
> r = self.session.request(
> self.http_method,
> self.path,
> json=self.json,
> params=self.params,
> headers=self.headers,
> )
> try:
> if (
> 200 <= r.status_code <= 299
> ): # Response.ok from JS (https://developer.mozilla.org/en-US/docs/Web/API/Response/ok)
> > return APIResponse.from_http_request_response(r)
>
> env/lib/python3.10/site-packages/postgrest/_sync/request_builder.py:66:
> _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
>
> cls = <class 'postgrest.base_request_builder.APIResponse'>
> request_response = <Response [200 OK]>
>
> @classmethod
> def from_http_request_response(
> cls: Type[APIResponse], request_response: RequestResponse
> ) -> APIResponse:
> try:
> data = request_response.json()
> except JSONDecodeError as e:
> return cls(data=[], count=0)
> count = cls._get_count_from_http_request_response(request_response)
> > return cls(data=data, count=count)
>
> env/lib/python3.10/site-packages/postgrest/base_request_builder.py:162:
> _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
>
> > ???
> E pydantic.error_wrappers.ValidationError: 1 validation error for APIResponse
> E data
> E value is not a valid list (type=type_error.list)
>
> pydantic/main.py:342: ValidationError
>
> The above exception was the direct cause of the following exception:
>
> def test_get_claims():
> client = get_client()
> > claims = get_claims(client, USER_ID)
>
> tests/test_claims.py:11:
> _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
> middlewares/claims.py:27: in get_claims
> res = client.rpc("get_claims", {"uid": uid}).execute()
Using `get_claims` function from:
https://github.com/wantpinow/supabase-custom-claims/blob/patch-1/install.sql
Is it because the function returns a `jsonb` ?
**To Reproduce**
Steps to reproduce the behavior:
1. Install the function `get_claims` from this PR https://github.com/wantpinow/supabase-custom-claims/blob/patch-1/install.sql
2. Call it from latest version of supabase-py
**Expected behavior**
Returns the function output
**Desktop (please complete the following information):**
- OS: MacOS 13.3
- Version `supabase==1.0.2`
| closed | 2023-04-02T14:25:32Z | 2024-02-10T15:13:45Z | https://github.com/supabase/supabase-py/issues/405 | [] | louis030195 | 16 |
dynaconf/dynaconf | django | 482 | [bug] Attribute error when accessing formatted value in layered config | **Describe the bug**
Attribute error when accessing settings value in layered configuration with multiple settings files.
<details>
<summary>Exception</summary>
```
Traceback (most recent call last):
File "app.py", line 11, in <module>
assert settings['s3_url'] == expected_value # fails
File "/home/user/dev/venv/lib/python3.7/site-packages/dynaconf/utils/functional.py", line 17, in inner
return func(self._wrapped, *args)
File "/home/user/dev/venv/lib/python3.7/site-packages/dynaconf/base.py", line 285, in __getitem__
value = self.get(item, default=empty)
File "/home/user/dev/venv/lib/python3.7/site-packages/dynaconf/base.py", line 419, in get
data = (parent or self.store).get(key, default)
File "/home/user/dev/venv/lib/python3.7/site-packages/dynaconf/utils/boxing.py", line 15, in evaluate
value = f(dynabox, item, *args, **kwargs)
File "/home/user/dev/venv/lib/python3.7/site-packages/dynaconf/utils/boxing.py", line 64, in get
return super(DynaBox, self).get(item, default, *args, **kwargs)
File "/home/user/dev/venv/lib/python3.7/site-packages/dynaconf/vendor/box/box.py", line 109, in get
return B[C]
File "/home/user/dev/venv/lib/python3.7/site-packages/dynaconf/utils/boxing.py", line 23, in evaluate
return recursively_evaluate_lazy_format(value, settings)
File "/home/user/dev/venv/lib/python3.7/site-packages/dynaconf/utils/__init__.py", line 355, in recursively_evaluate_lazy_format
value = value(settings)
File "/home/user/dev/venv/lib/python3.7/site-packages/dynaconf/utils/parse_conf.py", line 176, in __call__
return self.formatter(self.value, **self.context)
File "/home/user/dev/venv/lib/python3.7/site-packages/dynaconf/utils/parse_conf.py", line 138, in __call__
return self.function(value, **context)
AttributeError: 'Settings' object has no attribute 's3_protocol'
```
</details>
**To Reproduce**
Steps to reproduce the behavior:
1. Having the following folder structure
<details>
<summary> Project structure </summary>
```bash
$ ls
. .. app.py conf-test-default.toml conf-test-layer.toml
```
</details>
2. Having the following config files:
<details>
<summary> Config files </summary>
**conf-test-default.toml**
```toml
s3_protocol = 's3a'
s3_url = '@format {this.s3_protocol}://{this.s3_bucket}'
```
and
**conf-test-layer.toml**
```toml
s3_bucket = 'kewl_bucket'
```
</details>
3. Having the following app code:
<details>
<summary> Code </summary>
**app.py**
```python
from dynaconf import Dynaconf
settings_files = ['conf-test-default.toml', 'conf-test-layer.toml']
settings = Dynaconf(settings_files=settings_files)
expected_value = 's3a://kewl_bucket'
assert settings.s3_url == expected_value # succeeds
assert settings['s3_url'] == expected_value # fails
assert settings.get('s3_url', default='s3://default') == expected_values # fails
assert settings('s3_url', cast=str) == expected_value # fails
```
</details>
4. Executing under the following environment
<details>
<summary> Execution </summary>
```bash
$ python app.py
```
</details>
**Expected behavior**
I would expect all documented settings access methods to function properly.
**Environment (please complete the following information):**
- OS: Linux 5.9.10-arch1-1 #1 SMP PREEMPT Sun, 22 Nov 2020 14:16:59 +0000 x86_64 GNU/Linux
- Dynaconf Version 3.1.2
| closed | 2020-12-04T22:36:30Z | 2021-03-01T14:06:41Z | https://github.com/dynaconf/dynaconf/issues/482 | [
"bug"
] | billcrook | 3 |
WeblateOrg/weblate | django | 13,437 | Trailing format string in Android is broken | ### Describe the issue
In our Android app, we have the following string in our `values/strings.xml`:
```
<string name="cmd_camera_response_success">"The device is taking a picture and will send it to FMD Server. You can view it here soon: %s</string>
```
Note the trailing `%s`.
Recently, Weblate started to ignore the trailing "s", and suggest to users that they remove it from the translations.
This has resulted in some translators removing the "s", even though it should be there.
This results in the linter (rightfully!) complaining when compiling our Android app:
```
Error: Format string 'cmd_camera_response_success' is not a valid format string so it should not be passed to String.format [StringFormatInvalid]
transport.send(context, context.getString(R.string.cmd_camera_response_success, serverUrl))
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
/builds/Nulide/findmydevice/app/src/main/res/values-ta/strings.xml:21: This definition does not require arguments
```
This recently (in the last few weeks) broke. The string in question has not been touched on our main branch since June 2024, and up to now Weblate has not had this issue.
### I already tried
- [X] I've read and searched [the documentation](https://docs.weblate.org/).
- [X] I've searched for similar filed issues in this repository.
### Steps to reproduce the behavior
1. Define a string with a trailing "%s" in an Android app
2. Pull it into Weblate
### Expected behavior
The %s should be part of the string.
### Screenshots

### Exception traceback
```pytb
n/a
```
### How do you run Weblate?
weblate.org service
### Weblate versions
_No response_
### Weblate deploy checks
_No response_
### Additional context
Weblate project: https://hosted.weblate.org/projects/findmydevice/fmd-android/
Offending string in Gitlab: https://gitlab.com/Nulide/findmydevice/-/blame/1d0dbe75677f4a66bf8c6b2cb3fc5b3edf0013e0/app/src/main/res/values/strings.xml#L192
Offending string in Weblate: https://hosted.weblate.org/translate/findmydevice/fmd-android/en/?checksum=d7d28e73f83c973f | closed | 2025-01-05T21:29:29Z | 2025-01-10T14:16:55Z | https://github.com/WeblateOrg/weblate/issues/13437 | [
"bug",
"translate-toolkit"
] | thgoebel | 5 |
jupyter/nbgrader | jupyter | 1,153 | Randomising Values in questions and tests as part of assignment process | Is there a way / would it be useful to provide a way of incorporating randomised elements into a question.
For example, I might set a simple task:
*Load the file X into a pandas dataframe and preview the first {{import random;N=random.choice(random.randint(6,10)+random.randint(11,16)}} rows of it.*
and then in the next cell test on:
```python
assert_equal(_, pd.read_csv(X).head({{N}})
```
When the assignment is created, execute the `{{...}}` code as part of generating the assigned notebook....hmmm... once. per. student. That could get expensive, couldn't it?!
Okay, so maybe not for each student. But if you're in the habit of recycling questions one year to the next, with the occasional parameter change, then that could work?! | open | 2019-06-11T15:47:26Z | 2021-01-27T13:12:21Z | https://github.com/jupyter/nbgrader/issues/1153 | [
"enhancement"
] | psychemedia | 13 |
JoeanAmier/XHS-Downloader | api | 81 | 可否对视频笔记选择下载其封面图呢 | open | 2024-04-26T05:17:17Z | 2024-06-27T02:32:58Z | https://github.com/JoeanAmier/XHS-Downloader/issues/81 | [] | hzllllllll | 2 | |
igorbenav/fastcrud | pydantic | 91 | Nested Join Should Return List When Necessary | This was mentioned in #90
```python
async def get_card(self, card_id: uuid.UUID):
async with async_session_maker() as db:
return await card_crud.get_joined(
db=db,
id=card_id,
nest_joins=True,
joins_config=[
JoinConfig(
model=Article,
join_on=Article.card_id == Card.id,
join_type="left",
join_prefix="articles_",
schema_to_select=Article_schema,
)
]
)
```
Assuming a `card` has multiple `articles`, `articles` in joined response should be a list of articles. | closed | 2024-05-21T04:21:04Z | 2024-05-27T07:31:33Z | https://github.com/igorbenav/fastcrud/issues/91 | [
"bug",
"FastCRUD Methods"
] | igorbenav | 0 |
mars-project/mars | scikit-learn | 2,523 | [BUG] df.loc failed when df is empty: RuntimeError: generator raised StopIteration | <!--
Thank you for your contribution!
Please review https://github.com/mars-project/mars/blob/master/CONTRIBUTING.rst before opening an issue.
-->
**Describe the bug**
df.loc failed when df is empty: RuntimeError: generator raised StopIteration.
**To Reproduce**
To help us reproducing this bug, please provide information below:
1. Your Python version
2. The version of Mars you use
3. Versions of crucial packages, such as numpy, scipy and pandas
4. Full stack of the error.
5. Minimized code to reproduce the error.
```
In [8]: from mars.dataframe import DataFrame
...: df = DataFrame(pd.DataFrame({'a': [], 'b': []}))
...: df.set_index('a')
...: print(df.loc[df.index.to_tensor().to_numpy()].execute())
100%|█████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████| 100.0/100 [00:00<00:00, 1274.28it/s]
0%| | 0/100 [00:00<?, ?it/s]Unexpected error happens in <function TaskProcessor.get_next_stage_processor at 0x7fd82e1b1050>
Traceback (most recent call last):
File "/Users/qinxuye/Workspace/mars/mars/tensor/indexing/index_lib.py", line 937, in _process
context.out_nsplits = calc_nsplits(index_to_shape)
File "/Users/qinxuye/Workspace/mars/mars/utils.py", line 569, in calc_nsplits
ndim = len(next(iter(chunk_idx_to_shape)))
StopIteration
The above exception was the direct cause of the following exception:
Traceback (most recent call last):
File "/Users/qinxuye/Workspace/mars/mars/services/task/supervisor/processor.py", line 52, in inner
return await func(processor, *args, **kwargs)
File "/Users/qinxuye/Workspace/mars/mars/services/task/supervisor/processor.py", line 300, in get_next_stage_processor
chunk_graph = await self._get_next_chunk_graph(self._chunk_graph_iter)
File "/Users/qinxuye/Workspace/mars/mars/services/task/supervisor/processor.py", line 235, in _get_next_chunk_graph
chunk_graph = await fut
File "/Users/qinxuye/Workspace/mars/mars/lib/aio/_threads.py", line 36, in to_thread
return await loop.run_in_executor(None, func_call)
File "/Users/qinxuye/miniconda3/lib/python3.7/concurrent/futures/thread.py", line 57, in run
result = self.fn(*self.args, **self.kwargs)
File "/Users/qinxuye/Workspace/mars/mars/services/task/supervisor/processor.py", line 230, in next_chunk_graph
return next(chunk_graph_iter)
File "/Users/qinxuye/Workspace/mars/mars/services/task/supervisor/preprocessor.py", line 141, in tile
for chunk_graph in chunk_graph_builder.build():
File "/Users/qinxuye/Workspace/mars/mars/core/graph/builder/chunk.py", line 237, in build
yield from self._build()
File "/Users/qinxuye/Workspace/mars/mars/core/graph/builder/chunk.py", line 233, in _build
yield from self.tiler
File "/Users/qinxuye/Workspace/mars/mars/services/task/supervisor/preprocessor.py", line 69, in __iter__
to_update_tileables = self._iter()
File "/Users/qinxuye/Workspace/mars/mars/core/graph/builder/chunk.py", line 179, in _iter
next_tileable_handlers, to_update_tileables, visited)
File "/Users/qinxuye/Workspace/mars/mars/core/graph/builder/chunk.py", line 94, in _tile
need_process = next(tile_handler)
File "/Users/qinxuye/Workspace/mars/mars/core/graph/builder/chunk.py", line 70, in _tile_handler
tiled_tileables = yield from handler.tile(tiled_tileables)
File "/Users/qinxuye/Workspace/mars/mars/core/entity/tileables.py", line 77, in tile
tiled_result = yield from tile_handler(op)
File "/Users/qinxuye/Workspace/mars/mars/dataframe/indexing/loc.py", line 333, in tile
return [(yield from handler.handle(op))]
File "/Users/qinxuye/Workspace/mars/mars/tensor/indexing/index_lib.py", line 906, in handle
yield from self._process(context, index_infos)
RuntimeError: generator raised StopIteration
0%|
```
| closed | 2021-10-14T10:21:01Z | 2021-10-15T02:15:03Z | https://github.com/mars-project/mars/issues/2523 | [
"type: bug",
"mod: dataframe",
"prio: high"
] | qinxuye | 0 |
aws/aws-sdk-pandas | pandas | 2,453 | Error with typ='series' for read_json | ### Describe the bug
When using `s3.read_json` with the additional pandas argument `typ='series'` I get the following error: `AttributeError: 'Series' object has no attribute 'select_dtypes'`.
Using pandas.read_json with the same json file and `typ='series'` works without problems.
### How to Reproduce
```
d = {'a': 'dd', 'b': 'yy'}
j = json.dumps(d)
pd.read_json(j, typ='series')
```
This works without problems.
Now save as a json file somewhere on s3
```
s3 = boto3.resource('s3')
s3object = s3.Object(bucket, key)
s3object.put(Body=(bytes(json.dumps(d, indent=4).encode('UTF-8'))))
```
and try to load the json file
```
wr.s3.read_json(s3_path, typ='series')
```
Gives this error:
`AttributeError: 'Series' object has no attribute 'select_dtypes'`
### Expected behavior
No error, same as using pd.read_json()
### Your project
_No response_
### Screenshots
_No response_
### OS
Linux
### Python version
3.8
### AWS SDK for pandas version
3.3.0
### Additional context
_No response_ | closed | 2023-09-04T14:28:06Z | 2023-09-25T15:02:56Z | https://github.com/aws/aws-sdk-pandas/issues/2453 | [
"bug"
] | ClaudiaSchulz | 3 |
flairNLP/flair | pytorch | 3,540 | [Question]: Installing `pyab3p` | ### Question
When trying to run `species_linker = EntityMentionLinker.load("species-linker")` I am getting `'pyab3p' is not found, switching to a model without abbreviation resolution. This might impact the model performance. To reach full performance, please install pyab3p by running: pip install pyab3p`.
When I try to run `pip install pyab3p` I get `ERROR: Failed building wheel for pyab3p` due to `ab3p_source/Ab3P.cpp:1:10: fatal error: 'Ab3P.h' file not found`. Is this a known issue and is there a known fix? | closed | 2024-08-27T12:37:32Z | 2024-08-30T09:19:43Z | https://github.com/flairNLP/flair/issues/3540 | [
"question"
] | jessicapetrochuk | 9 |
allenai/allennlp | nlp | 4,837 | model_save_interval and restoring checkpoints | Say I have a very long running training process with a lot of data, where an epoch could take an hour.
So if i use model_save_interval to save the model every 30 minutes, can I restore the model to these intermediate training states? (if they get interrupted before a complete epoch)
If yes, then these intermediate states, do they restore the position of the batch as well (i.e do they start training from the exact batch that was interrupted) how is this handled ? | closed | 2020-12-04T05:16:22Z | 2020-12-18T16:46:12Z | https://github.com/allenai/allennlp/issues/4837 | [
"question"
] | vikigenius | 3 |
sinaptik-ai/pandas-ai | pandas | 1,294 | Skill name is not defined | ### System Info
OS version: macOS 14.5
Python version: Python 3.10.7
The current version of pandasai being used: 2.2.12
### 🐛 Describe the bug
# Bug: Skill Calculations Fail in PandasAI
## Issue Description
Skills that perform calculations are failing with a `NameError: name '<skill>' is not defined` error. This occurs because the `_extract_fix_dataframe_redeclarations` method executes code in an environment that lacks skill definitions.
## Root Cause
The `_extract_fix_dataframe_redeclarations` method uses an environment created by `get_environment()`, which does not include skill definitions:
```python
def _extract_fix_dataframe_redeclarations(
self, node: ast.AST, code_lines: list[str]
) -> ast.AST:
# ...
code = "\n".join(code_lines)
env = get_environment(self._additional_dependencies)
env["dfs"] = copy.deepcopy(self._get_originals(self._dfs))
exec(code, env)
# ...
```
The `get_environment()` function returns a dictionary with pandas, matplotlib, numpy, and some whitelisted builtins, but no skills:
```python
def get_environment(additional_deps: List[dict]) -> dict:
return {
"pd": pd,
"plt": plt,
"np": np,
# Additional dependencies and whitelisted builtins...
}
```
## Contrast with Correct Implementation
In contrast, the `execute_code` method in the `CodeExecution` class correctly adds skills to the environment:
```python
def execute_code(self, code: str, context: ExecutionContext):
# ...
if context.skills_manager.used_skills:
for skill_func_name in context.skills_manager.used_skills:
skill = context.skills_manager.get_skill_by_func_name(skill_func_name)
environment[skill_func_name] = skill
# ...
```
## Proposed Solution
To fix this issue, the `_extract_fix_dataframe_redeclarations` method should be updated to include skill definitions in its execution environment, similar to the `execute_code` method.
## Example
```
import os
import pandas as pd
from pandasai import Agent
from pandasai.skills import skill
from pandasai.llm import OpenAI
employees_data = {
"EmployeeID": [1, 2, 3, 4, 5],
"Name": ["John", "Emma", "Liam", "Olivia", "William"],
"Department": ["HR", "Sales", "IT", "Marketing", "Finance"],
}
salaries_data = {
"EmployeeID": [1, 2, 3, 4, 5],
"Salary": [5000, 6000, 4500, 7000, 5500],
}
employees_df = pd.DataFrame(employees_data)
salaries_df = pd.DataFrame(salaries_data)
# Add function docstring to give more context to model
@skill
def plot_salaries(names: list[str], salaries: list[int]):
"""
Displays the bar chart having name on x-axis and salaries on y-axis using matplotlib
Args:
names (list[str]): Employees' names
salaries (list[int]): Salaries
"""
import matplotlib.pyplot as plt
plt.bar(names, salaries)
plt.xlabel("Employee Name")
plt.ylabel("Salary")
plt.title("Employee Salaries")
plt.xticks(rotation=45)
@skill
def calculate_salary_betas(salaries: list[int]) -> list[float]:
"""
Calculates the betas (25th, 50th and 75th percentiles) of salaries.
Args:
salaries (list[int]): List of employee salaries
Returns:
list[float]: A list containing the 25th, 50th, and 75th percentiles
"""
import numpy as np
percentiles = np.percentile(salaries, [25, 50, 75])
return percentiles.tolist()
# By default, unless you choose a different LLM, it will use BambooLLM.
# You can get your free API key signing up at https://pandabi.ai (you can also configure it in your .env file)
llm = OpenAI(
api_token=os.getenv("OPENAI_API_KEY"), temperature=0, seed=26, model="gpt-4o"
)
agent = Agent(
[employees_df, salaries_df],
config={"llm": llm, "enforce_privacy": True},
memory_size=10,
)
agent.add_skills(plot_salaries, calculate_salary_betas)
# Chat with the agent
response = agent.chat("Create a table with salary betas")
```
Error:
```
Traceback (most recent call last):
File "pandas-ai/pandasai/pipelines/chat/code_cleaning.py", line 95, in execute
code_to_run = self.get_code_to_run(input, code_context)
File "pandas-ai/pandasai/pipelines/chat/code_cleaning.py", line 152, in get_code_to_run
code_to_run = self._clean_code(code, context)
File "pandas-ai/pandasai/pipelines/chat/code_cleaning.py", line 515, in _clean_code
self._extract_fix_dataframe_redeclarations(node, clean_code_lines)
File "pandas-ai/pandasai/pipelines/chat/code_cleaning.py", line 420, in _extract_fix_dataframe_redeclarations
exec(code, env)
File "<string>", line 5, in <module>
NameError: name 'calculate_salary_betas' is not defined
``` | closed | 2024-07-26T10:16:50Z | 2024-08-31T11:04:56Z | https://github.com/sinaptik-ai/pandas-ai/issues/1294 | [
"bug"
] | WojtAcht | 1 |
pywinauto/pywinauto | automation | 797 | Pywinauto Installation of Add-ons | ## Expected Behavior
I have to automate IBM Rhapsody8.3.1 setup.
There is a window where you need to select which features to be installed. Features are hold into a TreeView object and each has a combobox for selecting whether or not to install this feature.
I am using inspect.exe from Windows in order to find out the related settings.
## Actual Behavior
I Can select a treeview item but I cannot find the dropdownlist object/ option to click on it (to expand the dropdownlist),
Actually I need only the first TreeView Item(parent) which will select to install everything in list.
## Steps to Reproduce the Problem
See below picture
## Short Example of Code to Demonstrate the Problem
This is what `print_control_identifiers()` returns for that window:
```
Dialog - 'IBM Rational Rhapsody 8.3.1 64bit - InstallShield Wizard' (L708, T328, R1212, B711)
['IBM Rational Rhapsody 8.3.1 64bit - InstallShield WizardDialog', 'IBM Rational Rhapsody 8.3.1 64bit - InstallShield Wizard', 'Dialog']
child_window(title="IBM Rational Rhapsody 8.3.1 64bit - InstallShield Wizard", control_type="Window")
|
| TreeView - '' (L721, T427, R1034, B649)
| ['TreeView', 'Rhapsody Add-ons provide enhanced utilities for your Rhapsody environmentTreeView']
| child_window(auto_id="76538", control_type="Tree")
| |
| | ScrollBar - 'Vertical' (L1014, T430, R1031, B629)
| | ['Vertical', 'ScrollBar1', 'ScrollBar0', 'ScrollBar', 'VerticalScrollBar']
| | child_window(title="Vertical", auto_id="NonClientVerticalScrollBar", control_type="ScrollBar")
| | |
| | | Button - 'Line up' (L1014, T430, R1031, B447)
| | | ['Button1', 'Line upButton', 'Button0', 'Line up', 'Button']
| | | child_window(title="Line up", auto_id="UpButton", control_type="Button")
| | |
| | | Thumb - 'Position' (L1014, T447, R1031, B588)
| | | ['Thumb', 'PositionThumb1', 'PositionThumb', 'Position1', 'Position', 'Thumb1', 'Thumb0', 'Position0', 'PositionThumb0']
| | | child_window(title="Position", auto_id="ScrollbarThumb", control_type="Thumb")
| | |
| | | Button - 'Page down' (L1014, T588, R1031, B612)
| | | ['Page down', 'Button2', 'Page downButton']
| | | child_window(title="Page down", auto_id="DownPageButton", control_type="Button")
| | |
| | | Button - 'Line down' (L1014, T612, R1031, B629)
| | | ['Line down', 'Line downButton', 'Button3']
| | | child_window(title="Line down", auto_id="DownButton", control_type="Button")
| |
| | ScrollBar - 'Horizontal' (L724, T629, R1014, B646)
| | ['ScrollBar2', 'Horizontal', 'HorizontalScrollBar']
| | child_window(title="Horizontal", auto_id="NonClientHorizontalScrollBar", control_type="ScrollBar")
| | |
| | | Button - 'Column left' (L724, T629, R741, B646)
| | | ['Column left', 'Button4', 'Column leftButton']
| | | child_window(title="Column left", auto_id="UpButton", control_type="Button")
| | |
| | | Thumb - 'Position' (L741, T629, R846, B646)
| | | ['Position2', 'Thumb2', 'PositionThumb2']
| | | child_window(title="Position", auto_id="ScrollbarThumb", control_type="Thumb")
| | |
| | | Button - 'Page right' (L846, T629, R997, B646)
| | | ['Button5', 'Page right', 'Page rightButton']
| | | child_window(title="Page right", auto_id="DownPageButton", control_type="Button")
| | |
| | | Button - 'Column right' (L997, T629, R1014, B646)
| | | ['Column right', 'Column rightButton', 'Button6']
| | | child_window(title="Column right", auto_id="DownButton", control_type="Button")
| |
| | Thumb - '' (L1014, T629, R1031, B646)
| | ['Rhapsody Add-ons provide enhanced utilities for your Rhapsody environmentThumb', 'Thumb3']
| |
| | TreeItem - 'Rhapsody Add Ons - This feature will be installed on local hard drive.' (L797, T430, R1014, B446)
| | ['Rhapsody Add Ons - This feature will be installed on local hard drive.TreeItem', 'TreeItem0', 'TreeItem1', 'Rhapsody Add Ons - This feature will be installed on local hard drive.', 'TreeItem']
| | child_window(title="Rhapsody Add Ons - This feature will be installed on local hard drive.", control_type="TreeItem")
| | |
| | | TreeItem - 'Rational Rhapsody Gateway Add On - Requirements Traceability - This feature will not be available.' (L832, T446, R1014, B462)
| | | ['Rational Rhapsody Gateway Add On - Requirements Traceability - This feature will not be available.', 'Rational Rhapsody Gateway Add On - Requirements Traceability - This feature will not be available.TreeItem', 'TreeItem2']
| | | child_window(title="Rational Rhapsody Gateway Add On - Requirements Traceability - This feature will not be available.", control_type="TreeItem")
| | |
| | | TreeItem - 'Rational Rhapsody XMI Toolkit - XML Metadata Interchange - This feature will not be available.' (L832, T462, R1014, B478)
| | | ['Rational Rhapsody XMI Toolkit - XML Metadata Interchange - This feature will not be available.TreeItem', 'Rational Rhapsody XMI Toolkit - XML Metadata Interchange - This feature will not be available.', 'TreeItem3']
| | | child_window(title="Rational Rhapsody XMI Toolkit - XML Metadata Interchange - This feature will not be available.", control_type="TreeItem")
| | |
| | | TreeItem - 'Rational Rhapsody TestConductor Add On - This feature will not be available.' (L832, T478, R1014, B494)
| | | ['Rational Rhapsody TestConductor Add On - This feature will not be available.TreeItem', 'TreeItem4', 'Rational Rhapsody TestConductor Add On - This feature will not be available.']
| | | child_window(title="Rational Rhapsody TestConductor Add On - This feature will not be available.", control_type="TreeItem")
| | |
| | | TreeItem - 'Rational Rhapsody Automatic Test Generation Add On - This feature will not be available.' (L832, T494, R1014, B510)
| | | ['TreeItem5', 'Rational Rhapsody Automatic Test Generation Add On - This feature will not be available.TreeItem', 'Rational Rhapsody Automatic Test Generation Add On - This feature will not be available.']
| | | child_window(title="Rational Rhapsody Automatic Test Generation Add On - This feature will not be available.", control_type="TreeItem")
| | |
| | | TreeItem - 'Rational Rhapsody Rules Composer Add On - This feature will not be available.' (L832, T510, R1014, B526)
| | | ['Rational Rhapsody Rules Composer Add On - This feature will not be available.TreeItem', 'TreeItem6', 'Rational Rhapsody Rules Composer Add On - This feature will not be available.']
| | | child_window(title="Rational Rhapsody Rules Composer Add On - This feature will not be available.", control_type="TreeItem")
| | |
| | | TreeItem - 'Automotive, AUTOSAR system authoring and behavioral design and AutomotiveC profile - This feature will not be available.' (L832, T526, R1014, B542)
| | | ['Automotive, AUTOSAR system authoring and behavioral design and AutomotiveC profile - This feature will not be available.TreeItem', 'Automotive, AUTOSAR system authoring and behavioral design and AutomotiveC profile - This feature will not be available.', 'TreeItem7']
| | | child_window(title="Automotive, AUTOSAR system authoring and behavioral design and AutomotiveC profile - This feature will not be available.", control_type="TreeItem")
| | |
| | | TreeItem - 'Microsoft Visual Studio Workflow Integration - This feature, and all subfeatures, will be installed on local hard drive.' (L832, T542, R1014, B558)
| | | ['Microsoft Visual Studio Workflow Integration - This feature, and all subfeatures, will be installed on local hard drive.TreeItem', 'TreeItem8', 'Microsoft Visual Studio Workflow Integration - This feature, and all subfeatures, will be installed on local hard drive.']
| | | child_window(title="Microsoft Visual Studio Workflow Integration - This feature, and all subfeatures, will be installed on local hard drive.", control_type="TreeItem")
| | |
| | | TreeItem - 'Systems Engineering Add On - This feature, and all subfeatures, will be installed on local hard drive.' (L832, T558, R1014, B574)
| | | ['Systems Engineering Add On - This feature, and all subfeatures, will be installed on local hard drive.', 'TreeItem9', 'Systems Engineering Add On - This feature, and all subfeatures, will be installed on local hard drive.TreeItem']
| | | child_window(title="Systems Engineering Add On - This feature, and all subfeatures, will be installed on local hard drive.", control_type="TreeItem")
| | |
| | | TreeItem - 'Spell Checker - This feature will not be available.' (L832, T574, R1014, B590)
| | | ['TreeItem10', 'Spell Checker - This feature will not be available.TreeItem', 'Spell Checker - This feature will not be available.']
| | | child_window(title="Spell Checker - This feature will not be available.", control_type="TreeItem")
| | |
| | | TreeItem - 'Rhapsody Apps - This feature will not be available.' (L832, T590, R1014, B606)
| | | ['Rhapsody Apps - This feature will not be available.', 'Rhapsody Apps - This feature will not be available.TreeItem', 'TreeItem11']
| | | child_window(title="Rhapsody Apps - This feature will not be available.", control_type="TreeItem")
| | |
| | | TreeItem - 'Design Manager Client Extension 6.0.6 - This feature will not be available.' (L832, T606, R1014, B622)
| | | ['TreeItem12', 'Design Manager Client Extension 6.0.6 - This feature will not be available.', 'Design Manager Client Extension 6.0.6 - This feature will not be available.TreeItem']
| | | child_window(title="Design Manager Client Extension 6.0.6 - This feature will not be available.", control_type="TreeItem")
| | |
| | | TreeItem - 'Rhapsody Model Manager - This feature will be installed on local hard drive.' (L832, T622, R1014, B629)
| | | ['Rhapsody Model Manager - This feature will be installed on local hard drive.TreeItem', 'Rhapsody Model Manager - This feature will be installed on local hard drive.', 'TreeItem13']
| | | child_window(title="Rhapsody Model Manager - This feature will be installed on local hard drive.", control_type="TreeItem")
| | | |
| | | | TreeItem - 'Design Manager Importer - This feature will not be available.' (L0, T0, R0, B0)
| | | | ['Design Manager Importer - This feature will not be available.', 'Design Manager Importer - This feature will not be available.TreeItem', 'TreeItem14']
| | | | child_window(title="Design Manager Importer - This feature will not be available.", control_type="TreeItem")
|
| Button - 'Help' (L740, T678, R828, B700)
| ['HelpButton', 'Button7', 'Help']
| child_window(title="Help", auto_id="164", control_type="Button")
|
| Button - 'Space' (L835, T678, R923, B700)
| ['Space', 'SpaceButton', 'Button8']
| child_window(title="Space", auto_id="74817", control_type="Button")
|
| Button - '< Back' (L929, T678, R1017, B700)
| ['Button9', '< Back', '< BackButton']
| child_window(title="< Back", auto_id="74777", control_type="Button")
|
| Button - 'Next >' (L1017, T678, R1105, B700)
| ['Next >', 'Next >Button', 'Button10']
| child_window(title="Next >", auto_id="74847", control_type="Button")
|
| Button - 'Cancel' (L1112, T678, R1200, B700)
| ['Button11', 'Cancel', 'CancelButton']
| child_window(title="Cancel", auto_id="74774", control_type="Button")
|
| GroupBox - 'Feature Description' (L1044, T422, R1204, B650)
| ['Feature DescriptionGroupBox', 'Feature Description', 'GroupBox']
| child_window(title="Feature Description", auto_id="76541", control_type="Group")
|
| Static - '' (L1052, T444, R1197, B641)
| ['Static', 'Rhapsody Add-ons provide enhanced utilities for your Rhapsody environmentStatic1', 'Rhapsody Add-ons provide enhanced utilities for your Rhapsody environmentStatic', 'Static1', 'Rhapsody Add-ons provide enhanced utilities for your Rhapsody environmentStatic0', 'Static0']
| child_window(auto_id="75734", control_type="Text")
|
| Static - 'Rhapsody Add-ons provide enhanced utilities for your Rhapsody environment' (L733, T384, R1122, B417)
| ['Rhapsody Add-ons provide enhanced utilities for your Rhapsody environment', 'Rhapsody Add-ons provide enhanced utilities for your Rhapsody environmentStatic2', 'Static2']
| child_window(title="Rhapsody Add-ons provide enhanced utilities for your Rhapsody environment", auto_id="74807", control_type="Text")
|
| Static - 'Add-on Installation' (L723, T362, R1112, B395)
| ['Add-on Installation', 'Add-on Installation0', 'Add-on Installation1', 'Static3', 'Add-on InstallationStatic']
| child_window(title="Add-on Installation", auto_id="74809", control_type="Text")
|
| Image - 'Add-on Installation' (L711, T354, R1209, B412)
| ['Image', 'Add-on InstallationImage', 'Image0', 'Image1', 'Add-on Installation2']
| child_window(title="Add-on Installation", auto_id="76494", control_type="Image")
|
| Image - 'NewBinary20' (L711, T412, R1207, B414)
| ['NewBinary20', 'NewBinary20Image', 'Image2']
| child_window(title="NewBinary20", auto_id="76496", control_type="Image")
|
| Static - 'InstallShield' (L716, T659, R782, B676)
| ['Static4', 'InstallShield', 'InstallShield0', 'InstallShieldStatic0', 'InstallShieldStatic1', 'InstallShield1', 'InstallShieldStatic']
| child_window(title="InstallShield", auto_id="76498", control_type="Text")
|
| Static - 'InstallShield' (L715, T658, R781, B675)
| ['InstallShieldStatic2', 'InstallShield2', 'Static5']
| child_window(title="InstallShield", auto_id="76500", control_type="Text")
|
| Image - 'InstallShield' (L775, T666, R1207, B668)
| ['InstallShield3', 'Image3', 'InstallShieldImage']
| child_window(title="InstallShield", auto_id="76509", control_type="Image")
|
| TitleBar - '' (L727, T331, R1209, B354)
| ['', 'TitleBar']
| |
| | Menu - 'System' (L716, T336, R738, B358)
| | ['Menu', 'System', 'SystemMenu', 'System0', 'System1']
| | child_window(title="System", auto_id="MenuBar", control_type="MenuBar")
| | |
| | | MenuItem - 'System' (L716, T336, R738, B358)
| | | ['System2', 'MenuItem', 'SystemMenuItem']
| | | child_window(title="System", control_type="MenuItem")
| |
| | Button - 'Close' (L0, T0, R0, B0)
| | ['Close', 'Button12', 'CloseButton']
| | child_window(title="Close", control_type="Button")
```
## Specifications
- Pywinauto version: 0.6.7
- Python version and bitness: 3.5.4 x64
- Platform and OS: win10, x64

| open | 2019-08-27T13:40:11Z | 2019-09-29T16:43:17Z | https://github.com/pywinauto/pywinauto/issues/797 | [
"question"
] | Bujy | 2 |
CTFd/CTFd | flask | 2,129 | Add a healthcheck endpoint | Add a simple healthcheck endpoint. Likely something like `/healthcheck`. It should likely so a simple `SELECT 1` on the database and do a simple `get_config()` call to validate that everything is working and then return a 200 with "OK". On any failure it should return 500. | closed | 2022-05-25T18:58:46Z | 2022-06-16T18:39:47Z | https://github.com/CTFd/CTFd/issues/2129 | [
"easy"
] | ColdHeat | 0 |
minimaxir/textgenrnn | tensorflow | 13 | Word-level enhancements | Word level addition was a last-min change, so need to work on it a bit:
* Make sure the `vocab` abides by `max_length`.
* Add a feature to collapse punctuation. | closed | 2018-04-21T00:08:11Z | 2018-04-30T03:52:48Z | https://github.com/minimaxir/textgenrnn/issues/13 | [
"enhancement"
] | minimaxir | 3 |
hankcs/HanLP | nlp | 1,402 | hanlp 2.0.0-alpha.25 加载 hanlp.pretrained.pos.CTB5_POS_RNN_FASTTEXT_ZH 出错 | <!--
Please carefully fill out this form to bypass our spam filter. Please make sure that this is a bug. We only address bugs and feature requests issues on GitHub. Other questions should be posted on stackoverflow or https://bbs.hankcs.com/
以下必填,否则直接关闭。
-->
**Describe the bug**
hanlp 2.0.0-alpha.25 加载 hanlp.pretrained.pos.CTB5_POS_RNN_FASTTEXT_ZH 出错
**Code to reproduce the issue**
Provide a reproducible test case that is the bare minimum necessary to generate the problem.
```
tagger = hanlp.load(hanlp.pretrained.pos.CTB5_POS_RNN_FASTTEXT_ZH)
tagger(['我', '的', '希望', '是', '希望', '和平'])
```
**Describe the current behavior**
加载 hanlp.pretrained.pos.CTB5_POS_RNN_FASTTEXT_ZH 报错
**Expected behavior**
能够运行测试代码
**不会是内存不够吧?**
**System information**
- Ubuntu 18.04
- Python version: 3.6
- HanLP version: 2.0.0-alpha.25
**Other info / logs**
Downloading https://file.hankcs.com/hanlp/pos/ctb5_pos_rnn_fasttext_20191230_202639.zip to /root/.hanlp/pos/ctb5_pos_rnn_fasttext_20191230_202639.zip
100.00%, 1.4 MB/1.4 MB, 677 KB/s, ETA 0 s
Extracting /root/.hanlp/pos/ctb5_pos_rnn_fasttext_20191230_202639.zip to /root/.hanlp/pos
Downloading https://dl.fbaipublicfiles.com/fasttext/vectors-wiki/wiki.zh.zip#wiki.zh.bin to /root/.hanlp/thirdparty/dl.fbaipublicfiles.com/fasttext/vectors-wiki/wiki.zh.zip
1.68%, 53.6 MB/3.1 GB, 9.6 MB/s, ETA 5 m 27 s
100.00%, 3.1 GB/3.1 GB, 8.0 MB/s, ETA 0 s
Extracting /root/.hanlp/thirdparty/dl.fbaipublicfiles.com/fasttext/vectors-wiki/wiki.zh.zip to /root/.hanlp/thirdparty/dl.fbaipublicfiles.com/fasttext/vectors-wiki/wiki.zh
Failed to load https://file.hankcs.com/hanlp/pos/ctb5_pos_rnn_fasttext_20191230_202639.zip. See stack trace below
Traceback (most recent call last):
File "/usr/local/lib/python3.6/dist-packages/hanlp/utils/component_util.py", line 43, in load_from_meta_file
obj.load(save_dir, **load_kwargs)
File "/usr/local/lib/python3.6/dist-packages/hanlp/common/component.py", line 244, in load
self.build(**merge_dict(self.config, training=False, logger=logger, **kwargs, overwrite=True, inplace=True))
File "/usr/local/lib/python3.6/dist-packages/hanlp/common/component.py", line 255, in build
loss=kwargs.get('loss', None)))
File "/usr/local/lib/python3.6/dist-packages/hanlp/components/taggers/rnn_tagger.py", line 34, in build_model
embeddings = build_embedding(embeddings, self.transform.word_vocab, self.transform)
File "/usr/local/lib/python3.6/dist-packages/hanlp/layers/embeddings/__init__.py", line 33, in build_embedding
layer: tf.keras.layers.Embedding = tf.keras.utils.deserialize_keras_object(embeddings)
File "/usr/local/lib/python3.6/dist-packages/tensorflow_core/python/keras/utils/generic_utils.py", line 305, in deserialize_keras_object
return cls.from_config(cls_config)
File "/usr/local/lib/python3.6/dist-packages/tensorflow_core/python/keras/engine/base_layer.py", line 519, in from_config
return cls(**config)
File "/usr/local/lib/python3.6/dist-packages/hanlp/layers/embeddings/fast_text.py", line 35, in __init__
self.model = fasttext.load_model(filepath)
File "/usr/local/lib/python3.6/dist-packages/fasttext/FastText.py", line 350, in load_model
return _FastText(model_path=path)
File "/usr/local/lib/python3.6/dist-packages/fasttext/FastText.py", line 43, in __init__
self.f.loadModel(model_path)
**MemoryError: std::bad_alloc**
**https://file.hankcs.com/hanlp/pos/ctb5_pos_rnn_fasttext_20191230_202639.zip** was created with **hanlp-2.0.0, while you are running 2.0.0-alpha.25.** Try to upgrade hanlp with
pip install --upgrade hanlp
* [x] I've completed this form and searched the web for solutions.
| closed | 2020-01-14T04:14:01Z | 2020-01-14T05:58:08Z | https://github.com/hankcs/HanLP/issues/1402 | [
"question"
] | SoaringTiger | 1 |
flasgger/flasgger | flask | 130 | decorators requires_basic_auth not works |
```python
def requires_basic_auth(f):
"""Decorator to require HTTP Basic Auth for your endpoint."""
def check_auth(username, password):
logger.info('2'*100)
user = User.query.filter_by(username=username).first()
logger.info('username:%s', user.username)
if not user or not user.verify_password(password):
return False
g.user = user
return True
def authenticate():
return Response(
"Authentication required.", 401,
{"WWW-Authenticate": "Basic realm='Login Required'"},
)
@wraps(f)
def decorated(*args, **kwargs):
# NOTE: This example will require Basic Auth only when you run the
# app directly. For unit tests, we can't block it from getting the
# Swagger specs so we just allow it to go thru without auth.
# The following two lines of code wouldn't be needed in a normal
# production environment.
if __name__ != "__main__":
return f(*args, **kwargs)
auth = request.authorization
# if not auth or not check_auth(auth.username, auth.password):
if not check_auth(auth.username, auth.password):
return authenticate()
return f(*args, **kwargs)
return decorated
swagger = Swagger(
decorators=[requires_basic_auth],
template={
"swagger": "2.0",
"info": {
"title": "SimpleWay Core api",
"version": "1.0",
},
"consumes": [
"application/json",
],
"produces": [
"application/json",
],
}, config={
"headers": [
],
"specs": [
{
"endpoint": 'apispec_1',
"route": '/apispec_1.json',
"rule_filter": lambda rule: True, # all in
"model_filter": lambda tag: True, # all in
}
],
"static_url_path": "/flasgger_static",
# "static_folder": "static", # must be set by user
"swagger_ui": True,
"specs_route": "/apis/"
}
)
| closed | 2017-07-04T12:24:17Z | 2017-07-05T11:13:16Z | https://github.com/flasgger/flasgger/issues/130 | [
"bug"
] | CptJason | 3 |
microsoft/qlib | deep-learning | 1,630 | MemoryError triggered when importing 'TopkDropoutStrategy' from qlib.contrib.strategy |
```python
from typing import Tuple
import pandas as pd
import qlib
from qlib.contrib.strategy import TopkDropoutStrategy
from qlib.data import D
from qlib.utils import hash_args, init_instance_by_config
if __name__ == "__main__":
qlib.init(
provider_uri=r"D:\qlib_data",
region="cn",
)
TRAIN_PERIODS: Tuple = ("2013-01-01", "2017-12-31")
VALID_PERIODS: Tuple = ("2018-01-01", "2019-12-31")
TEST_PERIODS: Tuple = ("2020-01-01", "2023-05-31")
dataset_config = {
"class": "DatasetH",
"module_path": "qlib.data.dataset",
"kwargs": {
"handler": {
"class": "Alpha158",
"module_path": "qlib.contrib.data.handler",
"kwargs": {
"start_time": TRAIN_PERIODS[0],
"end_time": TEST_PERIODS[1],
"fit_start_time": TRAIN_PERIODS[0],
"fit_end_time": TRAIN_PERIODS[1],
"instruments": "csi300",
},
},
"segments": {
"train": TRAIN_PERIODS,
"valid": VALID_PERIODS,
"test": TEST_PERIODS,
},
},
}
dataset = init_instance_by_config(dataset_config)
```
此时会报错


*qlib版本为0.9.3*
在注释掉**from qlib.contrib.strategy import TopkDropoutStrategy**后便不会有问题 | open | 2023-08-22T02:47:51Z | 2023-11-21T10:35:19Z | https://github.com/microsoft/qlib/issues/1630 | [
"bug"
] | hugo2046 | 2 |
ibis-project/ibis | pandas | 10,213 | feat: error at construction time for illegal casts | ### Is your feature request related to a problem?
Consider `ibis.literal(1).cast("array<int64>")`. This currently doesn't error. It only errors once you try to execute the result. I don't think there is any backend where this cast would succeed. It would be great if I got this error as early as possible.
We DON'T want to be overly-sensitive, and disallow a cast that is actually implemented by a backend, but I think there are a subset of casts that we could be sure aren't illegal., and we should error earlier. Initial first thoughts:
- non-string to struct
- non-string to array
- non-string to map
- struct to non-string
- array to non-string
- map to non-string
- non-binary and non-string to geom
- geom to non-binary and non-string
### What is the motivation behind your request?
I'm getting errors, but it was tricky to debug the code at fault, since it happened so much earlier.
### Describe the solution you'd like
Use the already-implemented `castable()` function?
### What version of ibis are you running?
main
### What backend(s) are you using, if any?
_No response_
### Code of Conduct
- [X] I agree to follow this project's Code of Conduct | closed | 2024-09-24T20:29:36Z | 2024-11-02T12:10:36Z | https://github.com/ibis-project/ibis/issues/10213 | [
"feature"
] | NickCrews | 1 |
globaleaks/globaleaks-whistleblowing-software | sqlalchemy | 4,349 | v5.0.32 issue - Recipients unable to upload attachments to submissions | ### What version of GlobaLeaks are you using?
v5.0.32
### What browser(s) are you seeing the problem on?
All
### What operating system(s) are you seeing the problem on?
Windows, N/A
### Describe the issue
Hi @evilaliv3
With the newest release, recipients are not able to upload attachments to submissions.
The "Upload" button is grayed out for all recipients.
### Proposed solution
_No response_ | closed | 2024-12-06T12:27:41Z | 2024-12-06T16:10:14Z | https://github.com/globaleaks/globaleaks-whistleblowing-software/issues/4349 | [] | aetdr | 3 |
Lightning-AI/pytorch-lightning | data-science | 19,745 | When calling trainer.test() train_dataloader is also validated, which makes no sense | ### Bug description
In the current logic of pytorch-lightning everytime I call a` trainer.test() `it is also checked if the `train_dataloader()` function makes sense. This is problematic.
For example, I use a `WeightedRandomSampler` only in the` train_dataloader` for obvious reasons. In order for this to work I calculate
the `weights` and `num_samples` parameters in the `setup() stage="fit"` section of my code.
Of course when I trigger` trainer.test()` this code is not executed and thus weights and num_samples are never calculated, which
leads to an error when lightning validates the` train_dataloader` function.
I dont see any best practices to avoid this and no reason to validate code which is never executed.
### What version are you seeing the problem on?
v2.2
### How to reproduce the bug
_No response_
### Error messages and logs
```
# Error messages and logs here please
```
### Environment
<details>
<summary>Current environment</summary>
```
#- Lightning Component (e.g. Trainer, LightningModule, LightningApp, LightningWork, LightningFlow):
#- PyTorch Lightning Version (e.g., 1.5.0):
#- Lightning App Version (e.g., 0.5.2):
#- PyTorch Version (e.g., 2.0):
#- Python version (e.g., 3.9):
#- OS (e.g., Linux):
#- CUDA/cuDNN version:
#- GPU models and configuration:
#- How you installed Lightning(`conda`, `pip`, source):
#- Running environment of LightningApp (e.g. local, cloud):
```
</details>
### More info
_No response_
cc @justusschock @awaelchli | open | 2024-04-08T13:51:43Z | 2024-04-11T15:48:32Z | https://github.com/Lightning-AI/pytorch-lightning/issues/19745 | [
"bug",
"strategy: deepspeed"
] | asusdisciple | 2 |
LibreTranslate/LibreTranslate | api | 290 | Polish lang not working | Hi guys,
I wanted to test the website libretranslate.com but it looks like the polish language is not working. Looks like a bug.
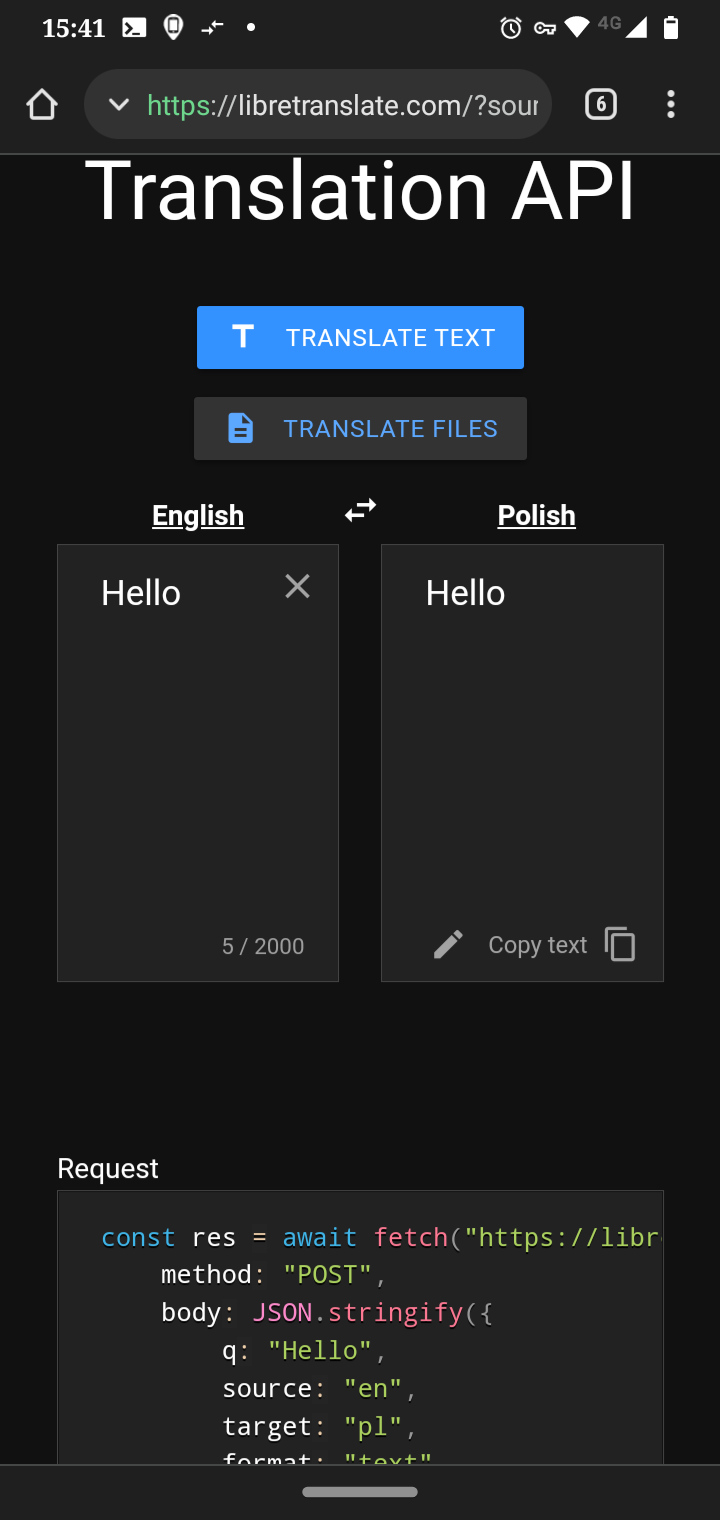
Also the dark mode on desktop does not show a list of languages properly in the dropdown/switch. | closed | 2022-07-31T14:43:51Z | 2022-09-24T12:54:16Z | https://github.com/LibreTranslate/LibreTranslate/issues/290 | [
"possible bug"
] | fairking | 5 |
comfyanonymous/ComfyUI | pytorch | 6,854 | Real-time sampling previews broken? | ### Your question
I'm still learning the terminology so please forgive me if I'm not using it right.
I'm running ComfyUI via StabilityMatrix. Up until about a week ago, both the ComfyUI web interface and the inference page within StabilityMatrix would show the sampler rendering the image in realtime. Now, some recent update in the past week has caused sampler previews to break; StabilityMatrix's inference preview won't show until an image is completed, and none of the built-in or add-on samplers in the web interface show anything at all, so I need to have a preview node (which again doesn't show an image until it's completed). Is there a workaround for this? Or a specific branch or commit I can check out that was prior to whatever broke sampling previews?
I apologize if there's already an issue addressing this but my cursory search of the repo didn't find one, but that might just be because I'm looking for the wrong terms.
### Logs
```powershell
```
### Other
_No response_ | closed | 2025-02-18T03:58:03Z | 2025-02-18T17:51:01Z | https://github.com/comfyanonymous/ComfyUI/issues/6854 | [
"User Support"
] | HowellBP | 4 |
sigmavirus24/github3.py | rest-api | 614 | Add repository attribute to Pull Destination object | [Here](https://github.com/sigmavirus24/github3.py/blob/develop/github3/pulls.py#L46) we check to see if there's a `'repo'` key in the decoded JSON. We should add `self.repository = Repository(...)` which uses that data.
##
<bountysource-plugin>
---
Want to back this issue? **[Post a bounty on it!](https://www.bountysource.com/issues/34820541-add-repository-attribute-to-pull-destination-object?utm_campaign=plugin&utm_content=tracker%2F183477&utm_medium=issues&utm_source=github)** We accept bounties via [Bountysource](https://www.bountysource.com/?utm_campaign=plugin&utm_content=tracker%2F183477&utm_medium=issues&utm_source=github).
</bountysource-plugin> | closed | 2016-06-03T01:12:58Z | 2018-03-22T02:20:56Z | https://github.com/sigmavirus24/github3.py/issues/614 | [
"help wanted",
"Mentored/Pair available"
] | sigmavirus24 | 2 |
docarray/docarray | fastapi | 1,599 | DocArray as a Retriever in Langchain | Follow up of https://github.com/docarray/docarray/issues/1580
Implement DocArray as a Retriever inside Langchain, supporting all doc index backends. Should look like this:
```python
from langchain.retrievers import DocArrayRetriever
from docarray import BaseDoc
from docarray.index import HnswDocumentIndex
from docarray.typing import NdArray
class MyDoc(BaseDoc):
title: str
title_embedding: NdArray[768]
# initialize docarray index (in this case hnsw, but will work for any backend)
db = HnswDocumentIndex[MyDoc](work_dir='./path/to/db')
# index data
db.index(
[
MyDoc(
title=f"My document {i}",
title_embedding=np.random.random(768),
)
for i in range(100)
]
# initialize retriever
# **search_params - search parameters, such as search_field, filters, etc.
retriever = DocArrayRetriever(index=db, **search_params)
```
And the usage:
```python
from langchain.chat_models import ChatOpenAI
from langchain.chains import ConversationalRetrievalChain
# use docarray retriever
model = ChatOpenAI(model_name='gpt-3.5-turbo')
qa = ConversationalRetrievalChain.from_llm(model, retriever=retriever)
``` | closed | 2023-05-31T12:18:09Z | 2023-06-19T06:41:05Z | https://github.com/docarray/docarray/issues/1599 | [] | jupyterjazz | 1 |
plotly/dash | jupyter | 2,794 | title option in options seems to not work | **Describe your context**
I would like to display information on hover an element of a checklist, but, it seems that does not work with the option title of options.
**options (list of dicts; optional): An array of options.
title (string; optional): The HTML ‘title’ attribute for the option. Allows for information on hover. For more information on this attribute, see [title - HTML: HyperText Markup Language | MDN](https://developer.mozilla.org/en-US/docs/Web/HTML/Global_attributes/title).**
Below, this is the version of dash package that I use:
dash 2.14.2
dash-ag-grid 31.0.1
dash-bootstrap-components 1.5.0
dash-bootstrap-templates 1.1.2
dash-core-components 2.0.0
dash-html-components 2.0.0
dash-table 5.0.0
How I use this option :
{
"label": html.Div(['London'], style={'color': 'LightGreen', 'font-size': 20}),
"value": "London",
**"title": "test",**
},
| open | 2024-03-13T09:12:07Z | 2024-08-13T19:47:13Z | https://github.com/plotly/dash/issues/2794 | [
"bug",
"P3"
] | PatSev | 2 |
autogluon/autogluon | data-science | 3,935 | [BUG] refit_full does not expand memory allowance when `use_bag_holdout=True` (`good_quality` preset) | - refit_full does not expand memory allowance when `use_bag_holdout=True` (`good_quality` preset)
- This can cause exceptions when the system is low on memory that should otherwise not occur.
- Further, sometimes memory avail can shrink dramatically between initial fit and refit (ex: 130GB avail at train, 72GB avail at refit). This can make things very challenging. Might require memory-safe sub-fits to avoid.
Example Logs:
```
Fitting model: XGBoost_BAG_L1 ... Training model for up to 7219.57s of the 62860.07s of remaining time.
Memory not enough to fit 8 folds in parallel. Will train 1 folds in parallel instead (Estimated 61.62% memory usage per fold, 61.62%/80.00% total).
Fitting 8 child models (S1F1 - S1F8) | Fitting with ParallelLocalFoldFittingStrategy (1 workers, per: cpus=24, gpus=0, memory=61.62%)
Switching to pseudo sequential ParallelFoldFittingStrategy to avoid Python memory leakage.
Overrule this behavior by setting fold_fitting_strategy to 'sequential_local' in ag_args_ensemble when when calling `predictor.fit`
-0.185 = Validation score (-mean_squared_error)
10665.07s = Training runtime
141.46s = Validation runtime
```
During Refit:
```
Fitting model: XGBoost_BAG_L1_FULL ...
Warning: Not enough memory to safely train model. Estimated to require 87.167 GB out of 72.005 GB available memory (121.057%)... (100.000% of avail memory is the max safe size)
To force training the model, specify the model hyperparameter "ag.max_memory_usage_ratio" to a larger value (currently 1.0, set to >=1.26 to avoid the error)
To set the same value for all models, do the following when calling predictor.fit: `predictor.fit(..., ag_args_fit={"ag.max_memory_usage_ratio": VALUE})`
Setting "ag.max_memory_usage_ratio" to values above 1 may result in out-of-memory errors. You may consider using a machine with more memory as a safer alternative.
Not enough memory to train XGBoost_BAG_L1_FULL... Skipping this model.
```
Because refit failed, this leads to an exception downstream.
| open | 2024-02-20T00:55:36Z | 2024-11-02T02:13:19Z | https://github.com/autogluon/autogluon/issues/3935 | [
"bug",
"module: tabular",
"Needs Triage",
"priority: 0"
] | Innixma | 2 |
matplotlib/mplfinance | matplotlib | 557 | Add table as a panel | Is there a way to add a custom panel, in my case a table of params?
Or a way to add an `mpf` figure to a `plt` grid? That way, I could put the table directly under the plt figure, combined into a single figure.
Thanks | closed | 2022-10-08T03:14:42Z | 2023-01-16T08:21:28Z | https://github.com/matplotlib/mplfinance/issues/557 | [
"question"
] | GAEfan | 3 |
plotly/dash | jupyter | 3,057 | Serious performance issues related to React context | When using components associated with the `XxxProvider`, severe performance issues can arise when there is a large amount of page content. Here are some examples related to well-known component libraries in the Dash ecosystem:
- with `dmc`
In `dmc`, it is required that the application be wrapped inside the `MantineProvider`. With the React Developer Tools, you can see that any interaction with an internal component will trigger a **re-render** of all components on the current page.

```python
import dash_mantine_components as dmc
from dash import Dash, _dash_renderer
_dash_renderer._set_react_version("18.2.0")
app = Dash(external_stylesheets=dmc.styles.ALL)
app.layout = dmc.MantineProvider([dmc.Button("test", style={"margin": 5})] * 200)
if __name__ == "__main__":
app.run(debug=True)
```
Even placing components from `dcc` under the `MantineProvider` will cause the same issue:

```python
import dash_mantine_components as dmc
from dash import Dash, _dash_renderer, dcc
_dash_renderer._set_react_version("18.2.0")
app = Dash(external_stylesheets=dmc.styles.ALL)
app.layout = dmc.MantineProvider([dcc.Input(style={"margin": 5})] * 200)
if __name__ == "__main__":
app.run(debug=True)
```
- with `fac`
In [fac](https://github.com/CNFeffery/feffery-antd-components), the similar component `AntdConfigProvider` is not a must-use, but the same issue will also occur:

```python
import dash
from dash import html
import feffery_antd_components as fac
app = dash.Dash(__name__)
app.layout = html.Div(
fac.AntdConfigProvider(
[fac.AntdButton("test", type="primary", style={"margin": 5})] * 100
)
)
if __name__ == "__main__":
app.run(debug=True)
```
---
However, the issue of global re-rendering does not occur with components within `html`, such as for `html.Div` (which has the functionality to update the click event to the component's `n_clicks` property):
- with `dmc`
```python
import dash_mantine_components as dmc
from dash import Dash, _dash_renderer, html
_dash_renderer._set_react_version("18.2.0")
app = Dash(external_stylesheets=dmc.styles.ALL)
app.layout = dmc.MantineProvider(
[html.Div(style={"height": 25, "border": "1px solid black", "marginBottom": 5})]
* 100
)
if __name__ == "__main__":
app.run(debug=True)
```
- with `fac`
```python
import dash
from dash import html
import feffery_antd_components as fac
app = dash.Dash(__name__)
app.layout = html.Div(
fac.AntdConfigProvider(
[html.Div(style={"height": 25, "border": "1px solid black", "marginBottom": 5})]
* 100
)
)
if __name__ == "__main__":
app.run(debug=True)
```
I hope to receive more help on this issue, to explore the deeper reasons and possible solutions.
| closed | 2024-11-02T03:28:25Z | 2025-02-06T14:21:32Z | https://github.com/plotly/dash/issues/3057 | [
"performance",
"P1"
] | CNFeffery | 4 |
ymcui/Chinese-BERT-wwm | tensorflow | 78 | 你在google tpu v3-8上训练roberta large的时候, batch size大小是多少 | closed | 2019-12-03T02:11:04Z | 2019-12-03T06:08:37Z | https://github.com/ymcui/Chinese-BERT-wwm/issues/78 | [] | xiongma | 3 | |
Kanaries/pygwalker | pandas | 638 | Support for Pygwalker Data Visualizations in `marimo` | **Is your feature request related to a problem? Please describe.**
When attempting to use pygwalker within marimo (a Python notebook framework), I encountered an issue where marimo was unable to display the pygwalker visualization. Specifically, I received the error message:
```
Unsupported mimetype: application/vnd.jupyter.widget-view+json
```

This prevents users from utilizing pygwalker's data visualization capabilities within marimo notebooks.
**Describe the solution you'd like**
I would like pygwalker to implement support for marimo by adding either a `__repr_html__` or `__mime__` method to the `pygwalker.api.pygwalker.PygWalker` class. This would allow marimo to properly render pygwalker visualizations, as described in the [marimo documentation for displaying objects](https://docs.marimo.io/guides/integrating_with_marimo/displaying_objects.html).
**Describe alternatives you've considered**
I initially tried using pygwalker with marimo following the standard instructions provided in the pygwalker repository, similar to how it's used in Jupyter notebooks. However, this approach resulted in the aforementioned error.
**Additional context**
This feature request originated from an attempt to integrate pygwalker with marimo, as documented in [marimo issue #2486](https://github.com/marimo-team/marimo/issues/2486). I got suggested filing this feature request with pygwalker to implement the necessary methods for compatibility.
Implementing this feature would greatly enhance the usability of pygwalker across different Python notebook environments, particularly benefiting users of marimo who wish to use pygwalker's data visualization capabilities. | closed | 2024-10-03T14:42:42Z | 2024-10-31T02:30:20Z | https://github.com/Kanaries/pygwalker/issues/638 | [
"enhancement",
"P1"
] | Haleshot | 18 |
scikit-learn/scikit-learn | data-science | 30,934 | DOC Missing doc string in tests present in sklearn/linear_model/_glm/tests/test_glm.py | ### Describe the issue related to documentation
The file `sklearn/linear_model/_glm/tests/test_glm.py` has the following tests without any doc string to describe what these functions aim to test.
- test_glm_wrong_y_range
- test_warm_start
- test_tags
- test_linalg_warning_with_newton_solver
### Suggested fix/improvement
Add doc strings to these tests similar to ones present in other tests with doc strings in the same file.
for example:
```
def test_linalg_warning_with_newton_solver(global_random_seed):
"""Test PoissonRegressor's behavior with the Newton solver under collinearity."""
```
### Additional Comments
I would like to work on this for my first documentation related work on this project. | closed | 2025-03-03T13:44:51Z | 2025-03-18T08:48:42Z | https://github.com/scikit-learn/scikit-learn/issues/30934 | [
"Documentation"
] | Rishab260 | 3 |
explosion/spaCy | nlp | 13,151 | No such command 'fill-curated-transformer' | I get the error `No such command 'fill-curated-transformer'.` when I try to run [`spacy init fill-curated-transformer`](https://spacy.io/api/cli#init-fill-curated-transformer).
Thank you for a great product and for your assistance in advance.
## How to reproduce the behaviour
1. Create a new python environment
2. `python -m pip install spacy spacy-curated-transformers`
3. `python -m spacy init fill-curated-transformer config.cfg - --model-name prajjwal1/bert-tiny`
```
Usage: python -m spacy init [OPTIONS] COMMAND [ARGS]...
Try 'python -m spacy init --help' for help.
Error: No such command 'fill-curated-transformer'.
```
## Your Environment
- **spaCy version:** 3.7.2
- **Platform:** macOS-14.1.1-x86_64-i386-64bit
- **Python version:** 3.10.11
| closed | 2023-11-24T20:16:29Z | 2023-12-29T00:02:02Z | https://github.com/explosion/spaCy/issues/13151 | [
"bug"
] | DanShatford | 4 |
LAION-AI/Open-Assistant | python | 3,723 | Chat doesn't open | dear ladies and gentelmen,
I try to open a new chat and I click on the button "Create a new chat" in the following link: https://open-assistant.io/chat
But it doesn't work and it doesn't open any new chat for me.
Please help me to fix this. Thank you very much.
Best Regards
Ehsan Pazooki | closed | 2023-11-04T17:50:50Z | 2023-11-28T07:16:10Z | https://github.com/LAION-AI/Open-Assistant/issues/3723 | [] | epz1371 | 1 |
davidsandberg/facenet | computer-vision | 899 | mtcnn+facenet bad performance on the new person not trained by knn or svm | hi,@davidsandberg,I used the facenet framework to train my own images ,then I used the mtcnn+facenet framework to real-time-recognition video stream, for the new and unknown person face. At present,I have a face data set that I give the 128-d embedding data as some one face data,which I call the face data set.And the data set has 20 people face (which was not trained by knn or svm)data now.The goal is that when someone go to in the front of the camera,the system will identify the face whether included by the data face set,if not included,then put in the data face.
But,when I run the system I give the very bad performance,can you help me explain the reason?
thanks | open | 2018-10-23T03:28:30Z | 2019-11-15T20:41:40Z | https://github.com/davidsandberg/facenet/issues/899 | [] | yuqj1991 | 6 |
flavors/django-graphql-jwt | graphql | 257 | JWT_REUSE_REFRESH_TOKENS documentation is wrong | Current [documentation](https://django-graphql-jwt.domake.io/en/latest/settings.html#jwt-reuse-refresh-tokens):
> Reuse the long running refreshed token instead of generating a new one
> Default: `False`
The correct description would be:
> A new long running refresh token is being generated but replaces the existing database record and thus invalidates the previous long running refresh token.
See this [test](https://github.com/flavors/django-graphql-jwt/blob/28e4f9749bac839d327914cfdda2ea3bb77bd775/tests/refresh_token/test_models.py#L56).
See the [code](https://github.com/flavors/django-graphql-jwt/blob/master/graphql_jwt/refresh_token/models.py#L37).
The [changelog](https://github.com/flavors/django-graphql-jwt/blob/28e4f9749bac839d327914cfdda2ea3bb77bd775/CHANGES.rst) says:
> Add JWT_REUSE_REFRESH_TOKENS setting in order to reuse the refresh token instances
| open | 2021-03-01T17:03:37Z | 2021-03-01T17:04:24Z | https://github.com/flavors/django-graphql-jwt/issues/257 | [] | googol7 | 0 |
pytorch/pytorch | machine-learning | 149,509 | `torch.compile` has a graph break when one of the `out_dims` of `torch.vmap` is set to `None` | ### 🐛 Describe the bug
I want to `torch.compile` a vmapped function (`torch.vmap(..., in_dims=(None, 0), out_dims=(None, 0))`) with the default "inductor" backend and `fullgraph=True`; however, it failed due to a graph break caused by the `torch._C._functorch.is_batchedtensor` function, which was invoked by `torch.vmap`.
This problem seems to be caused by setting an `out_dim` to `None` since the `is_batchedtensor` function will not be invoked otherwise.
I have searched for [the existing and past issues](https://github.com/pytorch/pytorch/issues); however, I failed to find issues related to `torch.compile` and `is_batchedtensor`/`out_dims`.
## Minimal reproducer
```
import torch
def test(x: torch.Tensor, y: torch.Tensor):
return x, y * 2
vmap_test = torch.vmap(test, in_dims=(None, 0), out_dims=(None, 0))
compiled_vmap_test = torch.compile(vmap_test, fullgraph=True)
print(compiled_vmap_test(torch.rand(3), torch.rand(3, 4)))
```
## Ablation
I have tried all of the ablations in https://pytorch.org/docs/main/torch.compiler_troubleshooting.html#reporting-issues. However, I got the same error as long as `fullgraph=True`.
### Error logs
```
Traceback (most recent call last):
File "c:\Users\admin\Documents\python_tests\unit_test\problems\test.py", line 8, in <module>
print(compiled_vmap_test(torch.rand(3), torch.rand(3, 4)))
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\eval_frame.py", line 574, in _fn
return fn(*args, **kwargs)
^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\convert_frame.py", line 1380, in __call__
return self._torchdynamo_orig_callable(
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\convert_frame.py", line 547, in __call__
return _compile(
^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\convert_frame.py", line 986, in _compile
guarded_code = compile_inner(code, one_graph, hooks, transform)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\convert_frame.py", line 715, in compile_inner
return _compile_inner(code, one_graph, hooks, transform)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_utils_internal.py", line 95, in wrapper_function
return function(*args, **kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\convert_frame.py", line 750, in _compile_inner
out_code = transform_code_object(code, transform)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\bytecode_transformation.py", line 1361, in transform_code_object
transformations(instructions, code_options)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\convert_frame.py", line 231, in _fn
return fn(*args, **kwargs)
^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\convert_frame.py", line 662, in transform
tracer.run()
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 2868, in run
super().run()
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 1052, in run
while self.step():
^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 962, in step
self.dispatch_table[inst.opcode](self, inst)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 659, in wrapper
return inner_fn(self, inst)
^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 1736, in CALL_FUNCTION_EX
self.call_function(fn, argsvars.items, kwargsvars)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 897, in call_function
self.push(fn.call_function(self, args, kwargs)) # type: ignore[arg-type]
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\higher_order_ops.py", line 1598, in call_function
return super().call_function(tx, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\functions.py", line 317, in call_function
return super().call_function(tx, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\functions.py", line 118, in call_function
return tx.inline_user_function_return(self, [*self.self_args(), *args], kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 903, in inline_user_function_return
return InliningInstructionTranslator.inline_call(self, fn, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 3072, in inline_call
return cls.inline_call_(parent, func, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 3198, in inline_call_
tracer.run()
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 1052, in run
while self.step():
^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 962, in step
self.dispatch_table[inst.opcode](self, inst)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 659, in wrapper
return inner_fn(self, inst)
^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 1736, in CALL_FUNCTION_EX
self.call_function(fn, argsvars.items, kwargsvars)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 897, in call_function
self.push(fn.call_function(self, args, kwargs)) # type: ignore[arg-type]
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\functions.py", line 317, in call_function
return super().call_function(tx, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\functions.py", line 118, in call_function
return tx.inline_user_function_return(self, [*self.self_args(), *args], kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 903, in inline_user_function_return
return InliningInstructionTranslator.inline_call(self, fn, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 3072, in inline_call
return cls.inline_call_(parent, func, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 3198, in inline_call_
tracer.run()
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 1052, in run
while self.step():
^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 962, in step
self.dispatch_table[inst.opcode](self, inst)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 659, in wrapper
return inner_fn(self, inst)
^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 2341, in CALL
self._call(inst)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 2335, in _call
self.call_function(fn, args, kwargs)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 897, in call_function
self.push(fn.call_function(self, args, kwargs)) # type: ignore[arg-type]
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\functions.py", line 317, in call_function
return super().call_function(tx, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\functions.py", line 118, in call_function
return tx.inline_user_function_return(self, [*self.self_args(), *args], kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 903, in inline_user_function_return
return InliningInstructionTranslator.inline_call(self, fn, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 3072, in inline_call
return cls.inline_call_(parent, func, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 3198, in inline_call_
tracer.run()
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 1052, in run
while self.step():
^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 962, in step
self.dispatch_table[inst.opcode](self, inst)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 659, in wrapper
return inner_fn(self, inst)
^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 2341, in CALL
self._call(inst)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 2335, in _call
self.call_function(fn, args, kwargs)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 897, in call_function
self.push(fn.call_function(self, args, kwargs)) # type: ignore[arg-type]
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\functions.py", line 317, in call_function
return super().call_function(tx, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\functions.py", line 118, in call_function
return tx.inline_user_function_return(self, [*self.self_args(), *args], kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 903, in inline_user_function_return
return InliningInstructionTranslator.inline_call(self, fn, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 3072, in inline_call
return cls.inline_call_(parent, func, args, kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 3198, in inline_call_
tracer.run()
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 1052, in run
while self.step():
^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 962, in step
self.dispatch_table[inst.opcode](self, inst)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 659, in wrapper
return inner_fn(self, inst)
^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 2341, in CALL
self._call(inst)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 2335, in _call
self.call_function(fn, args, kwargs)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\symbolic_convert.py", line 897, in call_function
self.push(fn.call_function(self, args, kwargs)) # type: ignore[arg-type]
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\torch.py", line 953, in call_function
tensor_variable = wrap_fx_proxy(
^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\builder.py", line 2153, in wrap_fx_proxy
return wrap_fx_proxy_cls(target_cls=TensorVariable, **kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\builder.py", line 2219, in wrap_fx_proxy_cls
return _wrap_fx_proxy(
^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\builder.py", line 2317, in _wrap_fx_proxy
return handle_traced_output(
^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\variables\builder.py", line 2517, in handle_traced_output
unimplemented(
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_dynamo\exc.py", line 317, in unimplemented
raise Unsupported(msg, case_name=case_name)
torch._dynamo.exc.Unsupported: torch.* op returned non-Tensor bool call_function <built-in method is_batchedtensor of PyCapsule object at 0x000001AAFF3C9470>
from user code:
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_functorch\apis.py", line 203, in wrapped
return vmap_impl(
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_functorch\vmap.py", line 331, in vmap_impl
return _flat_vmap(
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_functorch\vmap.py", line 480, in _flat_vmap
return _unwrap_batched(batched_outputs, out_dims, vmap_level, batch_size, func)
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_functorch\vmap.py", line 222, in _unwrap_batched
_maybe_remove_batch_dim(
File "C:\Users\admin\Documents\python_tests\.venv\Lib\site-packages\torch\_functorch\vmap.py", line 167, in _maybe_remove_batch_dim
if isinstance(batched_output, torch.Tensor) and is_batchedtensor(
Set TORCH_LOGS="+dynamo" and TORCHDYNAMO_VERBOSE=1 for more information
You can suppress this exception and fall back to eager by setting:
import torch._dynamo
torch._dynamo.config.suppress_errors = True
```
### Versions
PyTorch version: 2.6.0+cu126
Is debug build: False
CUDA used to build PyTorch: 12.6
ROCM used to build PyTorch: N/A
OS: Microsoft Windows 11 家庭中文版 (10.0.26100 64 位)
GCC version: Could not collect
Clang version: Could not collect
CMake version: Could not collect
Libc version: N/A
Python version: 3.12.7 (tags/v3.12.7:0b05ead, Oct 1 2024, 03:06:41) [MSC v.1941 64 bit (AMD64)] (64-bit runtime)
Python platform: Windows-11-10.0.26100-SP0
Is CUDA available: True
CUDA runtime version: 12.6.68
CUDA_MODULE_LOADING set to: LAZY
GPU models and configuration: GPU 0: NVIDIA GeForce RTX 4090
Nvidia driver version: 566.36
cuDNN version: Could not collect
HIP runtime version: N/A
MIOpen runtime version: N/A
Is XNNPACK available: True
CPU:
Name: 13th Gen Intel(R) Core(TM) i9-13900K
Manufacturer: GenuineIntel
Family: 207
Architecture: 9
ProcessorType: 3
DeviceID: CPU0
CurrentClockSpeed: 3000
MaxClockSpeed: 3000
L2CacheSize: 32768
L2CacheSpeed: None
Revision: None
Versions of relevant libraries:
[pip3] numpy==2.1.2
[pip3] torch==2.6.0+cu126
[pip3] torchaudio==2.6.0+cu126
[pip3] torchvision==0.21.0+cu126
[pip3] triton==3.2.0
[conda] Could not collect
cc @chauhang @penguinwu @voznesenskym @EikanWang @jgong5 @Guobing-Chen @XiaobingSuper @zhuhaozhe @blzheng @wenzhe-nrv @jiayisunx @chenyang78 @kadeng @amjames | open | 2025-03-19T13:09:55Z | 2025-03-20T16:05:01Z | https://github.com/pytorch/pytorch/issues/149509 | [
"triaged",
"oncall: pt2",
"module: dynamo",
"dynamo-triage-jan2025"
] | sses7757 | 2 |
mljar/mljar-supervised | scikit-learn | 378 | Setting to More Fully Explore Golden Features? | Hi,
For a project involving predicting several hierarchal compositional data analysis (CODA) chemical composition labels MLJAR-supervised works very well. Interestingly the golden features identified mostly have physical interpretation - essentially we are discovering feature combinations that are used in a real life laboratory to distinguish among samples.
So, we have a wish list:
1) can we have a setting to expand the compute budget and number of golden features discovered so we can see if additional known physical properties 'pop up'?
2) Somewhat more complex (and perhapsunreasonable to ask) would be to expand the operations considered in golden feature search, perhaps with this library or other symbolic regression package (https://github.com/MilesCranmer/PySR). In the current field of study there are many very large public datasets to which this technique could be applied with the aim to discover previously unknown physical relationships among laboratory measures that would provide better real world labeling of samples.
PM me if interested in collaboration
thanks!
| closed | 2021-04-17T13:40:34Z | 2021-04-26T14:31:26Z | https://github.com/mljar/mljar-supervised/issues/378 | [
"enhancement"
] | strelzoff-erdc | 2 |
ading2210/poe-api | graphql | 108 | Tokens not working (again) 😒 | # Pull request at #109
Same as #105 but it started happening again a few minutes ago | closed | 2023-06-09T19:13:00Z | 2023-06-09T20:13:07Z | https://github.com/ading2210/poe-api/issues/108 | [
"bug"
] | mak448a | 5 |
stanfordnlp/stanza | nlp | 1,383 | [QUESTION]Semantic Sentence Tokenization | I'm working with a corpus that primarily consists of longer documents. I'm seeking recommendations for the most effective approach to semantically tokenize them.
Examples:
```
Original Text: "I like the ambiance but the food was terrible."
Desired Output: ["I like the ambiance"] ["but the food was terrible."]
Original Text: "I don't know. I like the restaurant but not the food."
Desired Output: ["I don't know."] ["I like the restaurant"] ["but not the food."]
```
Any suggestions or advice on how to achieve this would be greatly appreciated! | closed | 2024-04-18T14:09:09Z | 2025-01-31T22:03:26Z | https://github.com/stanfordnlp/stanza/issues/1383 | [
"question",
"stale"
] | TheAIMagics | 3 |
2noise/ChatTTS | python | 909 | 音频token,如何获取 | 您好:
我想请问下,标签的音频token如何获取。微调的数据格式是什么样的,有示例么?损失函数是啥样的呢? | open | 2025-03-03T02:18:44Z | 2025-03-03T02:37:47Z | https://github.com/2noise/ChatTTS/issues/909 | [] | panhu | 0 |
modin-project/modin | data-science | 7,429 | BUG: Should check individual storage format and engine instead of global ones | This bug follows up on #7427.
There are places in the code where we check the global `Engine` or `StorageFormat` but to be precise we should check the configuration of the individual frame. I'll fix some of these in the PR For #7427, but others are more difficult to fix.
Places I've found so far:
- https://github.com/modin-project/modin/blob/1c4d173d3b2c44a1c1b5d5516552c7717b26de32/modin/core/execution/modin_aqp.py#L94
- https://github.com/sfc-gh-mvashishtha/modin/blob/a0d05698ebced75d539a0eb6bb0dd66dbb66f539/modin/core/execution/utils.py#L44
- https://github.com/sfc-gh-mvashishtha/modin/blob/a0d05698ebced75d539a0eb6bb0dd66dbb66f539/modin/error_message.py#L61
- Input methods, like read_pickle_glob, should continue using get_current_execution
- Output methods, like _to_pickle_glob, should not check get_current_execution() and should instead check the current dataframe’s execution
- There are some uses of get_current_execution() in the experimental batch mode. Can check dataframe’s engine instead.
| open | 2025-01-27T23:55:44Z | 2025-01-27T23:55:44Z | https://github.com/modin-project/modin/issues/7429 | [
"bug 🦗",
"P3"
] | sfc-gh-mvashishtha | 0 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.