repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
Gozargah/Marzban | api | 1,364 | Users migration strategy when node gets blocked by censor. | First of all, I would like to thank you for the work you’ve done on this project. It’s truly an important and valuable contribution to the fight against censorship.
Is there any recommended strategy for quickly and seamlessly migrating users if the node they are on gets blocked by a censor based on its IP address?
If there is no automated solution for this task at the moment, have you considered adding such functionality?
| closed | 2024-10-14T10:42:12Z | 2024-10-14T14:39:30Z | https://github.com/Gozargah/Marzban/issues/1364 | [] | lk-geimfari | 1 |
mars-project/mars | scikit-learn | 3,268 | [BUG] Ray executor raises ValueError: WRITEBACKIFCOPY base is read-only | <!--
Thank you for your contribution!
Please review https://github.com/mars-project/mars/blob/master/CONTRIBUTING.rst before opening an issue.
-->
**Describe the bug**
A clear and concise description of what the bug is.
```python
_____________________ test_predict_sparse_callable_kernel ______________________
setup = <mars.deploy.oscar.session.SyncSession object at 0x33564eee0>
def test_predict_sparse_callable_kernel(setup):
# This is a non-regression test for #15866
# Custom sparse kernel (top-K RBF)
def topk_rbf(X, Y=None, n_neighbors=10, gamma=1e-5):
nn = NearestNeighbors(n_neighbors=10, metric="euclidean", n_jobs=-1)
nn.fit(X)
W = -1 * mt.power(nn.kneighbors_graph(Y, mode="distance"), 2) * gamma
W = mt.exp(W)
assert W.issparse()
return W.T
n_classes = 4
n_samples = 500
n_test = 10
X, y = make_classification(
n_classes=n_classes,
n_samples=n_samples,
n_features=20,
n_informative=20,
n_redundant=0,
n_repeated=0,
random_state=0,
)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=n_test, random_state=0
)
model = LabelPropagation(kernel=topk_rbf)
> model.fit(X_train, y_train)
mars/learn/semi_supervised/tests/test_label_propagation.py:143:
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
mars/learn/semi_supervised/_label_propagation.py:369: in fit
return super().fit(X, y, session=session, run_kwargs=run_kwargs)
mars/learn/semi_supervised/_label_propagation.py:231: in fit
ExecutableTuple(to_run).execute(session=session, **(run_kwargs or dict()))
mars/core/entity/executable.py:267: in execute
ret = execute(*self, session=session, **kw)
mars/deploy/oscar/session.py:1888: in execute
return session.execute(
mars/deploy/oscar/session.py:1682: in execute
execution_info: ExecutionInfo = fut.result(
../../.pyenv/versions/3.8.13/lib/python3.8/concurrent/futures/_base.py:444: in result
return self.__get_result()
../../.pyenv/versions/3.8.13/lib/python3.8/concurrent/futures/_base.py:389: in __get_result
raise self._exception
mars/deploy/oscar/session.py:1868: in _execute
await execution_info
../../.pyenv/versions/3.8.13/lib/python3.8/asyncio/tasks.py:695: in _wrap_awaitable
return (yield from awaitable.__await__())
mars/deploy/oscar/session.py:105: in wait
return await self._aio_task
mars/deploy/oscar/session.py:953: in _run_in_background
raise task_result.error.with_traceback(task_result.traceback)
mars/services/task/supervisor/processor.py:372: in run
await self._process_stage_chunk_graph(*stage_args)
mars/services/task/supervisor/processor.py:250: in _process_stage_chunk_graph
chunk_to_result = await self._executor.execute_subtask_graph(
mars/services/task/execution/ray/executor.py:551: in execute_subtask_graph
meta_list = await asyncio.gather(*output_meta_object_refs)
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
awaitable = ObjectRef(c3f6db450a565c05ffffffffffffffffffffffff0100000001000000)
@types.coroutine
def _wrap_awaitable(awaitable):
"""Helper for asyncio.ensure_future().
Wraps awaitable (an object with __await__) into a coroutine
that will later be wrapped in a Task by ensure_future().
"""
> return (yield from awaitable.__await__())
E ray.exceptions.RayTaskError(ValueError): ray::execute_subtask() (pid=15135, ip=127.0.0.1)
E At least one of the input arguments for this task could not be computed:
E ray.exceptions.RayTaskError: ray::execute_subtask() (pid=15135, ip=127.0.0.1)
E At least one of the input arguments for this task could not be computed:
E ray.exceptions.RayTaskError: ray::execute_subtask() (pid=15135, ip=127.0.0.1)
E File "/home/admin/mars/mars/services/task/execution/ray/executor.py", line 185, in execute_subtask
E execute(context, chunk.op)
E File "/home/admin/mars/mars/core/operand/core.py", line 491, in execute
E result = executor(results, op)
E File "/home/admin/mars/mars/tensor/arithmetic/core.py", line 165, in execute
E ret = cls._execute_cpu(op, xp, lhs, rhs, **kw)
E File "/home/admin/mars/mars/tensor/arithmetic/core.py", line 142, in _execute_cpu
E return cls._get_func(xp)(lhs, rhs, **kw)
E File "/home/admin/mars/mars/lib/sparse/__init__.py", line 93, in power
E return a**b
E File "/home/admin/mars/mars/lib/sparse/array.py", line 503, in __pow__
E x = self.spmatrix.power(naked_other)
E File "/home/admin/.pyenv/versions/3.8.13/lib/python3.8/site-packages/scipy/sparse/_data.py", line 114, in power
E data = self._deduped_data()
E File "/home/admin/.pyenv/versions/3.8.13/lib/python3.8/site-packages/scipy/sparse/_data.py", line 32, in _deduped_data
E self.sum_duplicates()
E File "/home/admin/.pyenv/versions/3.8.13/lib/python3.8/site-packages/scipy/sparse/_compressed.py", line 1118, in sum_duplicates
E self.sort_indices()
E File "/home/admin/.pyenv/versions/3.8.13/lib/python3.8/site-packages/scipy/sparse/_compressed.py", line 1164, in sort_indices
E _sparsetools.csr_sort_indices(len(self.indptr) - 1, self.indptr,
E ValueError: WRITEBACKIFCOPY base is read-only
../../.pyenv/versions/3.8.13/lib/python3.8/asyncio/tasks.py:695: RayTaskError(ValueError)
```
```python
________________________ test_label_binarize_multilabel ________________________
setup = <mars.deploy.oscar.session.SyncSession object at 0x332666190>
def test_label_binarize_multilabel(setup):
y_ind = np.array([[0, 1, 0], [1, 1, 1], [0, 0, 0]])
classes = [0, 1, 2]
pos_label = 2
neg_label = 0
expected = pos_label * y_ind
y_sparse = [sp.csr_matrix(y_ind)]
for y in [y_ind] + y_sparse:
> check_binarized_results(y, classes, pos_label, neg_label, expected)
mars/learn/preprocessing/tests/test_label.py:250:
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
mars/learn/preprocessing/tests/test_label.py:186: in check_binarized_results
inversed = _inverse_binarize_thresholding(
../../.pyenv/versions/3.8.13/lib/python3.8/site-packages/sklearn/preprocessing/_label.py:649: in _inverse_binarize_thresholding
y.eliminate_zeros()
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
self = <3x3 sparse matrix of type '<class 'numpy.int64'>'
with 4 stored elements in Compressed Sparse Row format>
def eliminate_zeros(self):
"""Remove zero entries from the matrix
This is an *in place* operation.
"""
M, N = self._swap(self.shape)
> _sparsetools.csr_eliminate_zeros(M, N, self.indptr, self.indices,
self.data)
E ValueError: WRITEBACKIFCOPY base is read-only
../../.pyenv/versions/3.8.13/lib/python3.8/site-packages/scipy/sparse/_compressed.py:1077: ValueError
```
related issue: https://github.com/scipy/scipy/issues/8678
**To Reproduce**
To help us reproducing this bug, please provide information below:
1. Your Python version
2. The version of Mars you use
3. Versions of crucial packages, such as numpy, scipy and pandas
4. Full stack of the error.
5. Minimized code to reproduce the error.
**Expected behavior**
A clear and concise description of what you expected to happen.
**Additional context**
Add any other context about the problem here.
| closed | 2022-09-21T09:56:43Z | 2022-10-13T03:43:28Z | https://github.com/mars-project/mars/issues/3268 | [
"type: bug",
"mod: learn"
] | fyrestone | 0 |
supabase/supabase-py | fastapi | 516 | Cannot set options when instantiating Supabase client | **Describe the bug**
Cannot set options (such as schema, timeout etc.) for Supabase client in terminal or Jupyter notebook.
**To Reproduce**
Steps to reproduce the behavior:
1. Create a Supabase client in a new .py file using your database URL and service role key, and attempt to set an option:
```python
supabase: Client = create_client(url, key, {"schema": "some_other_schema"})
```
``
2. Attempt to run your .py file in the terminal, VSCode debug mode or a Jupyter notebook.
3. It will throw the error: `AttributeError: 'dict' object has no attribute 'headers'`
**Expected behavior**
Successful declaration of a new Supabase client, allowing the user to fetch or insert new data into, for example, a table on a different schema to 'public'.
**Screenshots**
<img width="916" alt="image" src="https://github.com/supabase-community/supabase-py/assets/101295184/7fbbce52-4b21-4190-b270-92d763535f65">
**Desktop (please complete the following information):**
- OS: macOS
| closed | 2023-08-08T04:41:12Z | 2023-08-08T05:31:18Z | https://github.com/supabase/supabase-py/issues/516 | [] | d-c-turner | 1 |
scikit-image/scikit-image | computer-vision | 6,906 | regionprops and regionprops_table crash when spacing != 1 | ### Description:
The `skimage.measure.regionprops` and `skimage.measure.regionprops_table` will crash when particular properties are passed and the `spacing` parameter is not 1 (or unspecified).
I think the `spacing` parameter is a new feature in v0.20.0 so this is probably a new bug.
I've got the code below to reproduce it.
When I pass the properties `label`, `area`, and `equivalent_diameter_area` then everything works fine with a custom `spacing`. Everything else seems to be trying to index a `float` value.
```props_dict_passing 1 ... PASSED
props_dict_passing 2 ... PASSED
Output exceeds the [size limit](command:workbench.action.openSettings?%5B%22notebook.output.textLineLimit%22%5D). Open the full output data [in a text editor](command:workbench.action.openLargeOutput?e9d046cb-8f53-4a89-b9dd-bc7dd3482e9b)---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
Cell In[14], line 42
39 print("props_dict_passing 2 ... PASSED")
41 # Fails now that spacing != 1 and eccentricity is passed
---> 42 props_dict_failing = regionprops_table(
43 label_image=label_image,
44 intensity_image=test_img,
45 spacing=0.5,
46 properties=bad_properties,
47 )
48 print("props_dict_failing ... PASSED")
File [~/mambaforge/envs/test-robusta-package/lib/python3.9/site-packages/skimage/measure/_regionprops.py:1038](https://vscode-remote+ssh-002dremote-002bbasic-002dst-002dm6ixlarge-002d1.vscode-resource.vscode-cdn.net/users/tony_reina/resilience_projects/starbux-image-viewer/notebooks/~/mambaforge/envs/test-robusta-package/lib/python3.9/site-packages/skimage/measure/_regionprops.py:1038), in regionprops_table(label_image, intensity_image, properties, cache, separator, extra_properties, spacing)
1031 intensity_image = np.zeros(
1032 label_image.shape + intensity_image.shape[ndim:],
1033 dtype=intensity_image.dtype
1034 )
1035 regions = regionprops(label_image, intensity_image=intensity_image,
1036 cache=cache, extra_properties=extra_properties, spacing=spacing)
-> 1038 out_d = _props_to_dict(regions, properties=properties,
1039 separator=separator)
1040 return {k: v[:0] for k, v in out_d.items()}
1042 return _props_to_dict(
1043 regions, properties=properties, separator=separator
...
264 delta[:, np.newaxis] ** np.arange(order + 1, dtype=float_dtype)
265 )
266 calc = np.rollaxis(calc, dim, image.ndim)
TypeError: 'float' object is not subscriptable```
### Way to reproduce:
```python
from skimage.measure import regionprops_table
from skimage.filters import threshold_otsu
from skimage.segmentation import clear_border
from skimage.measure import label, regionprops_table
from skimage.morphology import closing, square
import numpy as np
# Create random image 640x480
test_img = np.random.randint(0, 255, (640, 480))
# Detect objects
thresh = threshold_otsu(test_img)
bw = closing(test_img > thresh, square(3))
cleared = clear_border(bw)
label_image = label(cleared)
# Bugs start
good_properties = [
"label",
"equivalent_diameter_area",
"area",
]
# Works
props_dict_passing = regionprops_table(
label_image=label_image,
intensity_image=test_img,
spacing=0.5,
properties=good_properties,
)
print("props_dict_passing 1 ... PASSED")
# Add eccentricity (or major/minor axis and other float properties)
bad_properties = good_properties + ["eccentricity"]
# Works because spacing == 1
props_dict_passing2 = regionprops_table(
label_image=label_image,
intensity_image=test_img,
spacing=1,
properties=good_properties,
)
print("props_dict_passing 2 ... PASSED")
# Fails now that spacing != 1 and eccentricity is passed
props_dict_failing = regionprops_table(
label_image=label_image,
intensity_image=test_img,
spacing=0.5,
properties=bad_properties,
)
print("props_dict_failing ... PASSED")
```
### Version information:
```Shell
3.9.16 | packaged by conda-forge | (main, Feb 1 2023, 21:39:03)
[GCC 11.3.0]
Linux-5.10.147-133.644.amzn2.x86_64-x86_64-with-glibc2.26
scikit-image version: 0.21.0rc0
numpy version: 1.24.2
```
| open | 2023-04-21T18:31:04Z | 2023-09-16T14:09:05Z | https://github.com/scikit-image/scikit-image/issues/6906 | [
":bug: Bug"
] | tony-res | 6 |
onnx/onnx | machine-learning | 6,011 | [Feature request] Shape Inference for Einsum instead of Rank Inference | ### System information
v1.15.0
### What is the problem that this feature solves?
In the development of ONNX Runtime, we need know the output shape of each Op node for static graph compilation. However, we found that we could use onnx shape inference to achieve almost all output shapes except the output shape of Einsum. In `onnx/defs/math/defs.cc`, we found that there was only Rank Inference function for Einsum instead of Shape Inference. In a nutshell, shape inference for Einsum will be helpful for static graph compilations.
### Alternatives considered
_No response_
### Describe the feature
Just like the shape inference for all other ops, shape inference for Einsum should infer the output shape instead of rank according to the input shapes and the equation attribute.
We have developed a prototype version, which can be found in PR https://github.com/onnx/onnx/pull/6010. We would be delighted if this feature request is accepted. Alternatively, we are more than willing to provide assistance in incorporating this feature.
### Will this influence the current api (Y/N)?
No
### Feature Area
shape_inference
### Are you willing to contribute it (Y/N)
Yes
### Notes
_No response_ | closed | 2024-03-11T06:06:49Z | 2024-03-26T23:52:17Z | https://github.com/onnx/onnx/issues/6011 | [
"topic: enhancement",
"module: shape inference"
] | peishenyan | 1 |
huggingface/diffusers | deep-learning | 10,987 | Spatio-temporal diffusion models | **Is your feature request related to a problem? Please describe.**
Including https://github.com/yyysjz1997/Awesome-TimeSeries-SpatioTemporal-Diffusion-Model/blob/main/README.md models
| open | 2025-03-06T14:39:11Z | 2025-03-06T14:39:11Z | https://github.com/huggingface/diffusers/issues/10987 | [] | moghadas76 | 0 |
JaidedAI/EasyOCR | pytorch | 681 | Accelerate reader.readtext() with OpenMP | Hello all, this is more a question than an issue. I know `reader.readtext()` can be accelerated if I have a GPU with CUDA available; I was wondering if there was a flag to accelerate it with multi-threading (OpenMP).
Regards,
Victor | open | 2022-03-14T01:31:44Z | 2022-03-14T01:31:44Z | https://github.com/JaidedAI/EasyOCR/issues/681 | [] | vkrGitHub | 0 |
open-mmlab/mmdetection | pytorch | 11,753 | RuntimeError: handle_0 INTERNAL ASSERT FAILED at "../c10/cuda/driver_api.cpp":15, please report a bug to PyTorch. ```none | Thanks for your error report and we appreciate it a lot.
**Checklist**
1. I have searched related issues but cannot get the expected help.
2. I have read the [FAQ documentation](https://mmdetection.readthedocs.io/en/latest/faq.html) but cannot get the expected help.
3. The bug has not been fixed in the latest version.
**Describe the bug**
When I used the mask2former for instance segmentation, an error came out.
mask_pred = mask_pred[is_thing]
RuntimeError: handle_0 INTERNAL ASSERT FAILED at "../c10/cuda/driver_api.cpp":15, please report a bug to PyTorch.
**Reproduction**
1. What command or script did you run?
```none
A placeholder for the command.
```
2. Did you make any modifications on the code or config? Did you understand what you have modified?
3. What dataset did you use?
A segmentation dataset
**Environment**
1. Please run `python mmdet/utils/collect_env.py` to collect necessary environment information and paste it here.
2. You may add addition that may be helpful for locating the problem, such as
- How you installed PyTorch \[e.g., pip, conda, source\]
- Other environment variables that may be related (such as `$PATH`, `$LD_LIBRARY_PATH`, `$PYTHONPATH`, etc.)
**Error traceback**
If applicable, paste the error trackback here.
return _VF.meshgrid(tensors, **kwargs) # type: ignore[attr-defined]
05/29 16:12:42 - mmengine - INFO - Saving checkpoint at 44 iterations
Traceback (most recent call last):
File "tools/train.py", line 121, in <module>
main()
File "tools/train.py", line 117, in main
runner.train()
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/mmengine/runner/runner.py", line 1777, in train
model = self.train_loop.run() # type: ignore
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/mmengine/runner/loops.py", line 294, in run
self.runner.val_loop.run()
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/mmengine/runner/loops.py", line 373, in run
self.run_iter(idx, data_batch)
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/torch/utils/_contextlib.py", line 115, in decorate_context
return func(*args, **kwargs)
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/mmengine/runner/loops.py", line 393, in run_iter
outputs = self.runner.model.val_step(data_batch)
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/mmengine/model/base_model/base_model.py", line 133, in val_step
return self._run_forward(data, mode='predict') # type: ignore
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/mmengine/model/base_model/base_model.py", line 361, in _run_forward
results = self(**data, mode=mode)
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/torch/nn/modules/module.py", line 1518, in _wrapped_call_impl
return self._call_impl(*args, **kwargs)
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/torch/nn/modules/module.py", line 1527, in _call_impl
return forward_call(*args, **kwargs)
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/mmdet/models/detectors/base.py", line 94, in forward
return self.predict(inputs, data_samples)
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/mmdet/models/detectors/maskformer.py", line 103, in predict
results_list = self.panoptic_fusion_head.predict(
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/mmdet/models/seg_heads/panoptic_fusion_heads/maskformer_fusion_head.py", line 255, in predict
ins_results = self.instance_postprocess(
File "/home/xuym/miniconda3/envs/openmmlab/lib/python3.8/site-packages/mmdet/models/seg_heads/panoptic_fusion_heads/maskformer_fusion_head.py", line 167, in instance_postprocess
mask_pred = mask_pred[is_thing]
RuntimeError: handle_0 INTERNAL ASSERT FAILED at "../c10/cuda/driver_api.cpp":15, please report a bug to PyTorch.
```none
A placeholder for trackback.
```
**Bug fix**
If you have already identified the reason, you can provide the information here. If you are willing to create a PR to fix it, please also leave a comment here and that would be much appreciated!
| open | 2024-05-29T08:23:07Z | 2024-05-29T08:23:22Z | https://github.com/open-mmlab/mmdetection/issues/11753 | [] | AIzealotwu | 0 |
cookiecutter/cookiecutter-django | django | 4,872 | You probably don't need `get_user_model` | ## Description
Import `User` directly, rather than using `get_user_model`.
## Rationale
`get_user_model` is meant for *reusable* apps, while it is my understanding this project is targeted more towards creating websites than packages. Especially within the `users` app it doesn't make any sense to use it (are we expecting users to create a new user model in a custom app but then keep the `users` app in their project?) , and switching can prevent new users from assuming they have to use it in all their custom apps (which may or may not be what happened to me). For a more in-depth explanation of why it's an anti-pattern, read this [blog post](https://adamj.eu/tech/2022/03/27/you-probably-dont-need-djangos-get-user-model/). | closed | 2024-02-18T02:15:15Z | 2024-02-21T10:01:58Z | https://github.com/cookiecutter/cookiecutter-django/issues/4872 | [
"enhancement"
] | mfosterw | 1 |
apache/airflow | data-science | 47,630 | AIP-38 Turn dag run breadcrumb into a dropdown | ### Body
Make it easier to switch between dag runs in the graph view by using the breadcrumb as a dropdown like we had in the designs when the graph view was in its own modal.
### Committer
- [x] I acknowledge that I am a maintainer/committer of the Apache Airflow project. | closed | 2025-03-11T15:37:11Z | 2025-03-17T14:06:34Z | https://github.com/apache/airflow/issues/47630 | [
"kind:feature",
"area:UI",
"AIP-38"
] | bbovenzi | 2 |
Miserlou/Zappa | django | 1,525 | Support for generating slimmer packages | Currently zappa packaging will include all pip packages installed in the virtualenv. Installing zappa in the venv brings in a ton of dependencies. Depending on the app's actual needs, most/all of these don't actually need to be packaged and shipped to lambda. This unnecessarily increases the size of the package which makes zappa deploy/update much slower than it would otherwise.
As an example, for a simple hello world app, the package is over 8MB. The vast majority of this data is unneeded.
A possible approach here is to have an option to:
- don't package up anything from venv
- use requirements.txt in a way that doesn't slow deploy down
I see #525 and #542 but they don't seem to be resolved yet. Let me know if I'm missing anything! | open | 2018-06-08T20:15:28Z | 2019-04-04T14:08:19Z | https://github.com/Miserlou/Zappa/issues/1525 | [
"feature-request"
] | figelwump | 15 |
ultralytics/yolov5 | machine-learning | 12,514 | a questions when improve YOLOv5 | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and found no similar bug report.
### YOLOv5 Component
Training, Detection
### Bug
I want to improve ECA-attention, but there have same bug, which i cant not solve, i want your help@glenn-jocher . When i run yolo.py it work, but run train.py, there have been some issues.
`class EfficientChannelAttention(nn.Module): # Efficient Channel Attention module
def __init__(self, c, b=1, gamma=2):
super(EfficientChannelAttention, self).__init__()
t = int(abs((math.log(c, 2) + b) / gamma))
k = t if t % 2 else t + 1
self.avg_pool = nn.AdaptiveAvgPool2d(1)
self.conv1 = nn.Conv1d(1, 1, kernel_size=k, padding=int(k/2), bias=False)
self.sigmoid = nn.Sigmoid()
def forward(self, x):
# print('x是:{}'.format(x.size))
out = self.avg_pool(x)
# print('out是:{}'.format(out))
out_flat = out.view(-1)
orig_shape = out.size()
print('out_flat:{}'.format(out_flat))
sorted_indices = torch.argsort(out_flat,descending=True)
print('sorted_indices为:{}'.format(sorted_indices))
reshape_indices = sorted_indices.view(*orig_shape)
# print('reshape_indices:{}'.format(reshape_indices.shape))
soted_out = out.flatten()[sorted_indices].reshape(*orig_shape)
# print('soted_out为:{}'.format(soted_out))
# sorted_x = x.view(x.size()[0],-1,x.size()[-2],x.size()[-1])[reshape_indices]
sorted_x = torch.index_select(x, dim = 1, index =sorted_indices)
# print('sorted_x的形状:{}'.format(sorted_x.shape))
# print('排序后的x:{}'.format(sorted_x))
out2 = self.avg_pool(sorted_x)
# print('avgpool验证排序:{}'.format(out2))
soted_out = self.conv1(soted_out.squeeze(-1).transpose(-1, -2)).transpose(-1, -2).unsqueeze(-1)
soted_out = self.sigmoid(soted_out)
# print('out的形状:{}'.format(out.shape))
# print(out * sorted_x)
return soted_out * sorted_x`
`# parameters
nc: 80 # number of classes
depth_multiple: 0.33 # model depth multiple
width_multiple: 0.50 # layer channel multiple
# anchors
anchors:
- [10,13, 16,30, 33,23] # P3/8
- [30,61, 62,45, 59,119] # P4/16
- [116,90, 156,198, 373,326] # P5/32
# YOLOv5 backbone
backbone:
# [from, number, module, args]
[[-1, 1, Focus, [64, 3]], # 0-P1/2
[-1, 1, Conv, [128, 3, 2]], # 1-P2/4
[-1, 3, C3, [128]],
[-1, 1, Conv, [256, 3, 2]], # 3-P3/8
[-1, 9, C3, [256]],
[-1, 1, Conv, [512, 3, 2]], # 5-P4/16
[-1, 9, C3, [512]],
[-1, 1, Conv, [1024, 3, 2]], # 7-P5/32
[-1, 1, SPP, [1024, [5, 9, 13]]],
[-1, 3, C3, [1024, False]], # 9
]
# YOLOv5 head
head:
[[-1, 1, Conv, [512, 1, 1]],
[-1, 1, nn.Upsample, [None, 2, 'nearest']],
[[-1, 6], 1, Concat, [1]], # cat backbone P4
[-1, 3, C3, [512, False]], # 13
[-1, 1, Conv, [256, 1, 1]],
[-1, 1, nn.Upsample, [None, 2, 'nearest']],
[[-1, 4], 1, Concat, [1]], # cat backbone P3
[-1, 3, C3, [256, False]], # 17 (P3/8-small)
[-1, 1, Conv, [256, 3, 2]],
[[-1, 14], 1, Concat, [1]], # cat head P4
[-1, 3, C3, [512, False]], # 20 (P4/16-medium)
[-1, 1, Conv, [512, 3, 2]],
[-1, 1, EfficientChannelAttention, [512]],
[[-1, 10], 1, Concat, [1]], # cat head P5
[-1, 3, C3, [1024, False]], # 23 (P5/32-large)
[[17, 20, 24], 1, Detect, [nc, anchors]], # Detect(P3, P4, P5)
]`
When i use cpu the follow problem appear:
` Epoch gpu_mem box obj cls total labels img_size
0%| | 0/11049 [00:00<?, ?it/s]
out_flat:tensor([ 0.13989, 0.01097, 0.67497, ..., 0.14956, 0.13888, -0.00238], grad_fn=<ViewBackward0>)
sorted_indices为:tensor([ 27, 84, 107, ..., 539, 596, 706])
Traceback (most recent call last):
File "/home/wjh/learning/1/yolov5-5.0/train.py", line 543, in <module>
train(hyp, opt, device, tb_writer)
File "/home/wjh/learning/1/yolov5-5.0/train.py", line 303, in train
pred = model(imgs) # forward
File "/home/wjh/.conda/envs/Yolov5/lib/python3.9/site-packages/torch/nn/modules/module.py", line 1194, in _call_impl
return forward_call(*input, **kwargs)
File "/home/wjh/learning/1/yolov5-5.0/models/yolo.py", line 123, in forward
return self.forward_once(x, profile) # single-scale inference, train
File "/home/wjh/learning/1/yolov5-5.0/models/yolo.py", line 139, in forward_once
x = m(x) # run
File "/home/wjh/.conda/envs/Yolov5/lib/python3.9/site-packages/torch/nn/modules/module.py", line 1194, in _call_impl
return forward_call(*input, **kwargs)
File "/home/wjh/learning/1/yolov5-5.0/models/common.py", line 411, in forward
sorted_x = torch.index_select(x, dim = 1, index =sorted_indices)
RuntimeError: INDICES element is out of DATA bounds, id=918 axis_dim=256
进程已结束,退出代码`
The display exceeds the index, but I have checked the index during yolo. py runtime and everything is fine,
### Environment
_No response_
### Minimal Reproducible Example
_No response_
### Additional
_No response_
### Are you willing to submit a PR?
- [ ] Yes I'd like to help by submitting a PR! | closed | 2023-12-16T12:44:17Z | 2024-10-20T19:34:34Z | https://github.com/ultralytics/yolov5/issues/12514 | [
"bug",
"Stale"
] | haoaZ | 5 |
praw-dev/praw | api | 1,404 | Smarter MoreComments Algorithim | If we basically wait on a bunch of MoreComments, why could we not add up all of the MoreComments into one big MoreComment and then replace as needed? It would make the replace_more algorithm much faster.
PR #1403 implements a queue, so we could theoretically combine and get a lot at once. If it's a matter of linking the new comments to match with their parent comments, we could use a dict of some sort, and have the new objects go to the Comment key. | closed | 2020-04-22T03:59:07Z | 2021-05-20T17:46:48Z | https://github.com/praw-dev/praw/issues/1404 | [
"Feature",
"Discussion"
] | PythonCoderAS | 3 |
BeastByteAI/scikit-llm | scikit-learn | 86 | can you share link to Agent Dingo | can you share link to Agent Dingo | closed | 2024-03-03T19:33:56Z | 2024-03-04T21:57:06Z | https://github.com/BeastByteAI/scikit-llm/issues/86 | [] | Sandy4321 | 1 |
AutoGPTQ/AutoGPTQ | nlp | 363 | Why inference gets slower by going down to lower bits?(in comparison with ggml) | Hi Team,
Thanks for the great work.
I had a few doubts about quantized inference. I was doing the benchmark test and found that inference gets slower by going down to lower bits(4->3->2). Below are the inference details on the A6000 GPU:
4 bit(3.7G): 48 tokens/s
3 bit(2.9G): 38 tokens/s
2 bit(2.2G): 39 tokens/s
What is the reason behind the inference getting slower by going to lower bits? but this is not the case for ggml, where the inference speed gets better.
Thanks | closed | 2023-10-06T21:26:56Z | 2023-10-25T12:54:20Z | https://github.com/AutoGPTQ/AutoGPTQ/issues/363 | [
"bug"
] | Darshvino | 1 |
mwaskom/seaborn | data-science | 2,986 | swarmplot change point maximum displacement from center | Hi,
I am trying to plot a `violinplot` + `swarmplot` combination for with multiple hues and many points and am struggling to get the optimal clarity with as few points as possible overlapping. I tried both `swarmplot` and `stripplot`, with and without `dodge`.
Since i have multiple categories on the y-axis , I have also played around with the figure size, setting it to large height values. It helps to improve the clarity of the violin plots but the swarm/strip plots remain unchanged and crowded with massive overlap. I know that there will always be overlap with many points sharing the same/similar x-values, but i would like to maximize the use of space available between y-values for the swarms. Is there a way i can increase the maximum displacement from center for the swarm plots? With the `stripplot` `jitter` i can disperese the point, but they tend to overlap randomly quite a bit still and also start to move over into other violin plots.
Tried with Seaborn versions: `0.11.2` and `0.12.0rc0 `
I attached a partial plot, as the original is quite large:
```
...
sns.set_theme()
sns.set(rc={"figure.figsize": (6, 18)})
...
PROPS = {'boxprops': {'edgecolor': 'black'},
'medianprops': {'color': 'black'},
'whiskerprops': {'color': 'black'},
'capprops': {'color': 'black'}}
ax = sns.violinplot(x=stat2show, y=y_cat, data=data_df, width=1.7, fliersize=0,
linewidth=0.75, order=y_order, palette=qual_colors,
scale="count", inner="quartile", **PROPS)
sns.swarmplot(x=stat2show, y=y_cat, data=data_df, size=5.2, color='white',
linewidth=0.5, hue="Data Set", edgecolor='black',
palette=data_set_palette, order=y_order, dodge=False,
hue_order=data_set_hue_order)
...
```

Thanks for any help! | closed | 2022-08-30T10:05:12Z | 2022-08-30T11:44:51Z | https://github.com/mwaskom/seaborn/issues/2986 | [] | ohickl | 4 |
plotly/dash | dash | 2,302 | [BUG] Error when passing list of components in dash component properties other than children. | ```
dash 2.7.0
dash-core-components 2.0.0
dash-html-components 2.0.0
dash-table 5.0.0
dash-mantine-components 0.11.0a0
```
**Describe the bug**
When passing components in dash component properties other than `children`, an error is thrown if a list of components is passed.
```python
from dash import Dash
from dash_iconify import DashIconify
import dash_mantine_components as dmc
app = Dash(__name__)
app.layout = dmc.Divider(label=["GitHub", DashIconify(icon="fa:github")])
if __name__ == "__main__":
app.run_server(debug=True)
```
Error:
<img width="1614" alt="Screenshot 2022-11-06 at 12 10 52 AM" src="https://user-images.githubusercontent.com/91216500/200135808-1b1ea37d-4b7b-4871-9b02-e02412340600.png">
This behaviour is observed even if a single component is passed in the list:
```python
app.layout = dmc.Divider(label=[DashIconify(icon="fa:github")])
```
**Expected behavior**
No error should be displayed even when multiple components are passed.
**Screenshots**
NA | closed | 2022-11-05T18:48:04Z | 2022-12-05T16:24:34Z | https://github.com/plotly/dash/issues/2302 | [] | snehilvj | 0 |
junyanz/pytorch-CycleGAN-and-pix2pix | deep-learning | 730 | Looking for performance metric for cyclegan | Hi, we often apply cycleGAN for unpaired data. So, some of the performance metric will be not applied
- SSIM
- PSNR
For my dataset, I would like to use cyclegan to mapping an image from winter session to spring session and they have no pair data for each image. Could you tell me how can I evaluate the cyclegan performance (i.e how to know the output is close to a realistic image...)
| closed | 2019-08-14T21:55:25Z | 2020-04-25T18:18:55Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/730 | [] | John1231983 | 6 |
Evil0ctal/Douyin_TikTok_Download_API | api | 439 | 抖音视频可以解析,但无法下载 | ***发生错误的平台?***
抖音
***发生错误的端点?***
Web APP
***提交的输入值?***
如:短视频链接
***是否有再次尝试?***
如:是,发生错误后X时间后错误依旧存在。
***你有查看本项目的自述文件或接口文档吗?***
如:有,并且很确定该问题是程序导致的。
{
"code": 400,
"message": "Client error '403 Forbidden' for url 'http://v3-web.douyinvod.com/045365e0ddece3cd7bb6ee83a1f2207c/6687bd6a/video/tos/cn/tos-cn-ve-15c001-alinc2/oEheBigbIOzQoZA3EBy2VNiBQOkaRAfR30AEpT/?a=6383&ch=26&cr=3&dr=0&lr=all&cd=0%7C0%7C0%7C3&cv=1&br=1508&bt=1508&cs=0&ds=4&ft=pEaFx4hZffPdhb~NI1VNvAq-antLjrKaM9V.RkaFmfTeejVhWL6&mime_type=video_mp4&qs=0&rc=MztoNTVkODxoZGc8ZDg7M0BpM29oN3E5cjw2czMzNGkzM0A0NTZeLjYwNmMxNi42YGAyYSNmZDA0MmRzNm1gLS1kLWFzcw%3D%3D&btag=c0000e00008000&cquery=100B_100x_100z_100o_100w&dy_q=1720164672&feature_id=46a7bb47b4fd1280f3d3825bf2b29388&l=20240705153112480A6FC15A49E08AB062'\nFor more information check: https://developer.mozilla.org/en-US/docs/Web/HTTP/Status/403",
"support": "Please contact us on Github: https://github.com/Evil0ctal/Douyin_TikTok_Download_API",
"time": "2024-07-05 07:28:20",
"router": "/api/download",
"params": {
"url": "https://v.douyin.com/i6CgdQHY/",
"prefix": "true",
"with_watermark": "false"
}
}
| closed | 2024-07-05T07:34:35Z | 2024-07-10T02:50:07Z | https://github.com/Evil0ctal/Douyin_TikTok_Download_API/issues/439 | [
"BUG"
] | zttlovedouzi | 3 |
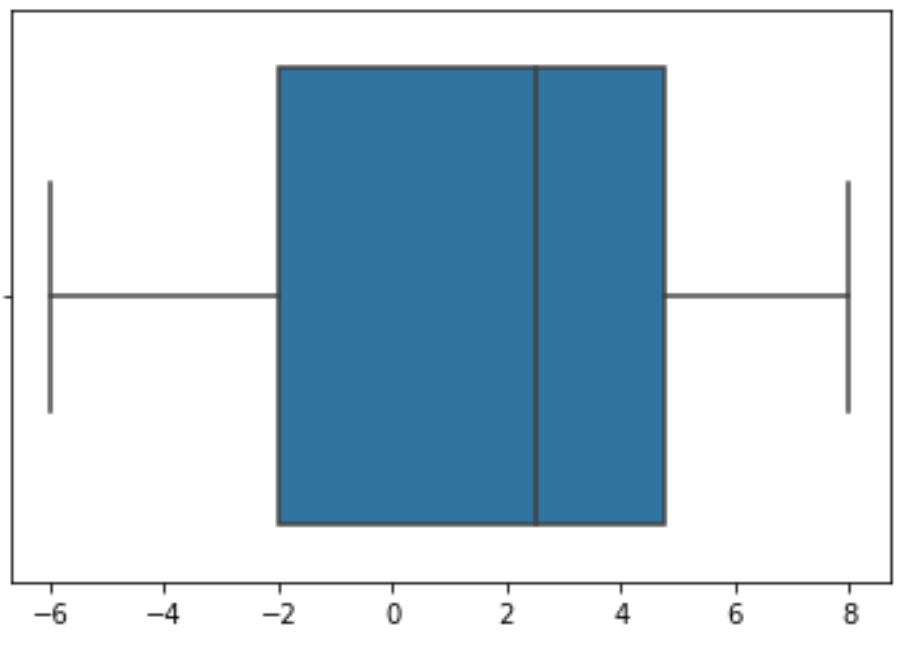
mwaskom/seaborn | pandas | 3,025 | Boxplot Bug | Hi
The issue happens when use sns.boxplot for quartiles. In terms of the quartile Wikipedia with links (https://en.wikipedia.org/wiki/Quartile), Q1, Q2(median), and Q3 are all mean but in the different subsets. If that is the case, in the list containing elements - -6,-3,1,4,5,8, Q1 is -3, median for 2.5, and Q3 for 5. However, when running the method, sns.boxplot, I find Q1 shows at the edge of the blue box is -2. Hence, I think this is bug.
In order to facilitate your inquiry, I paste both source code and plot, which are shown below.

| closed | 2022-09-14T18:06:19Z | 2022-09-15T00:58:12Z | https://github.com/mwaskom/seaborn/issues/3025 | [] | tac628 | 1 |
davidteather/TikTok-Api | api | 855 | time out error and new connection error | when i run the code it shows time out error and new connection error,but i can get access to https://www.tiktok.com/@laurenalaina
the code is :
```
from TikTokApi import TikTokApi
verify_fp = " "
api = TikTokApi(custom_verify_fp=verify_fp)
user = api.user(username="laurenalaina")
for video in user.videos():
print(video.id)
```
it shows
```
TimeoutError Traceback (most recent call last)
~\anaconda3\lib\site-packages\urllib3\connection.py in _new_conn(self)
158 try:
--> 159 conn = connection.create_connection(
160 (self._dns_host, self.port), self.timeout, **extra_kw
~\anaconda3\lib\site-packages\urllib3\util\connection.py in create_connection(address, timeout, source_address, socket_options)
83 if err is not None:
---> 84 raise err
85
~\anaconda3\lib\site-packages\urllib3\util\connection.py in create_connection(address, timeout, source_address, socket_options)
73 sock.bind(source_address)
---> 74 sock.connect(sa)
75 return sock
TimeoutError: [WinError 10060] 由于连接方在一段时间后没有正确答复或连接的主机没有反应,连接尝试失败。
During handling of the above exception, another exception occurred:
NewConnectionError Traceback (most recent call last)
~\anaconda3\lib\site-packages\urllib3\connectionpool.py in urlopen(self, method, url, body, headers, retries, redirect, assert_same_host, timeout, pool_timeout, release_conn, chunked, body_pos, **response_kw)
669 # Make the request on the httplib connection object.
--> 670 httplib_response = self._make_request(
671 conn,
~\anaconda3\lib\site-packages\urllib3\connectionpool.py in _make_request(self, conn, method, url, timeout, chunked, **httplib_request_kw)
380 try:
--> 381 self._validate_conn(conn)
382 except (SocketTimeout, BaseSSLError) as e:
~\anaconda3\lib\site-packages\urllib3\connectionpool.py in _validate_conn(self, conn)
975 if not getattr(conn, "sock", None): # AppEngine might not have `.sock`
--> 976 conn.connect()
977
~\anaconda3\lib\site-packages\urllib3\connection.py in connect(self)
307 # Add certificate verification
--> 308 conn = self._new_conn()
309 hostname = self.host
~\anaconda3\lib\site-packages\urllib3\connection.py in _new_conn(self)
170 except SocketError as e:
--> 171 raise NewConnectionError(
172 self, "Failed to establish a new connection: %s" % e
NewConnectionError: <urllib3.connection.HTTPSConnection object at 0x00000251A9F6D670>: Failed to establish a new connection: [WinError 10060] 由于连接方在一段时间后没有正确答复或连接的主机没有反应,连接尝试失败。
During handling of the above exception, another exception occurred:
MaxRetryError Traceback (most recent call last)
~\anaconda3\lib\site-packages\requests\adapters.py in send(self, request, stream, timeout, verify, cert, proxies)
438 if not chunked:
--> 439 resp = conn.urlopen(
440 method=request.method,
~\anaconda3\lib\site-packages\urllib3\connectionpool.py in urlopen(self, method, url, body, headers, retries, redirect, assert_same_host, timeout, pool_timeout, release_conn, chunked, body_pos, **response_kw)
723
--> 724 retries = retries.increment(
725 method, url, error=e, _pool=self, _stacktrace=sys.exc_info()[2]
~\anaconda3\lib\site-packages\urllib3\util\retry.py in increment(self, method, url, response, error, _pool, _stacktrace)
438 if new_retry.is_exhausted():
--> 439 raise MaxRetryError(_pool, url, error or ResponseError(cause))
440
MaxRetryError: HTTPSConnectionPool(host='www.tiktok.com', port=443): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPSConnection object at 0x00000251A9F6D670>: Failed to establish a new connection: [WinError 10060] 由于连接方在一段时间后没有正确答复或连接的主机没有反应,连接尝试失败。'))
During handling of the above exception, another exception occurred:
ConnectionError Traceback (most recent call last)
<ipython-input-10-c2db16034fb7> in <module>
6 user = api.user(username="laurenalaina")
7
----> 8 for video in user.videos():
9 print(video.id)
~\anaconda3\lib\site-packages\TikTokApi\api\user.py in videos(self, count, cursor, **kwargs)
131
132 if not self.user_id and not self.sec_uid:
--> 133 self.__find_attributes()
134
135 first = True
~\anaconda3\lib\site-packages\TikTokApi\api\user.py in __find_attributes(self)
261 # It is more efficient to check search first, since self.user_object() makes HTML request.
262 found = False
--> 263 for u in self.parent.search.users(self.username):
264 if u.username == self.username:
265 found = True
~\anaconda3\lib\site-packages\TikTokApi\api\search.py in search_type(search_term, obj_type, count, offset, **kwargs)
78 cursor = offset
79
---> 80 spawn = requests.head(
81 "https://www.tiktok.com",
82 proxies=Search.parent._format_proxy(processed.proxy),
~\anaconda3\lib\site-packages\requests\api.py in head(url, **kwargs)
102
103 kwargs.setdefault('allow_redirects', False)
--> 104 return request('head', url, **kwargs)
105
106
~\anaconda3\lib\site-packages\requests\api.py in request(method, url, **kwargs)
59 # cases, and look like a memory leak in others.
60 with sessions.Session() as session:
---> 61 return session.request(method=method, url=url, **kwargs)
62
63
~\anaconda3\lib\site-packages\requests\sessions.py in request(self, method, url, params, data, headers, cookies, files, auth, timeout, allow_redirects, proxies, hooks, stream, verify, cert, json)
528 }
529 send_kwargs.update(settings)
--> 530 resp = self.send(prep, **send_kwargs)
531
532 return resp
~\anaconda3\lib\site-packages\requests\sessions.py in send(self, request, **kwargs)
641
642 # Send the request
--> 643 r = adapter.send(request, **kwargs)
644
645 # Total elapsed time of the request (approximately)
~\anaconda3\lib\site-packages\requests\adapters.py in send(self, request, stream, timeout, verify, cert, proxies)
514 raise SSLError(e, request=request)
515
--> 516 raise ConnectionError(e, request=request)
517
518 except ClosedPoolError as e:
ConnectionError: HTTPSConnectionPool(host='www.tiktok.com', port=443): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPSConnection object at 0x00000251A9F6D670>: Failed to establish a new connection: [WinError 10060] 由于连接方在一段时间后没有正确答复或连接的主机没有反应,连接尝试失败。'))
```
**Desktop (please complete the following information):**
- OS: [Windows 10][jupyter notebook]
- TikTokApi Version [e.g. 5.0.0] - if out of date upgrade before posting an issue
**Additional context**
Add any other context about the problem here.
| closed | 2022-03-13T04:00:57Z | 2023-08-08T22:18:06Z | https://github.com/davidteather/TikTok-Api/issues/855 | [
"bug"
] | sxy-dawnwind | 2 |
litestar-org/litestar | pydantic | 3,814 | Enhancement: consider adding mypy plugin for type checking `data.create_instance(id=1, address__id=2)` | ### Summary
Right now [`create_instance`](https://docs.litestar.dev/latest/reference/dto/data_structures.html#litestar.dto.data_structures.DTOData.create_instance) can take any `**kwargs`.
But, mypy has no way of actually checking that `id=1, address__id=2` are valid keywords for this call.
It can be caught when executed, sure. But typechecking is much faster than writing code + writing tests + running them.
In Django we have a similar pattern of passing keywords like this to filters. Like: `User.objects.filter(id=1, settings__profile="public")`. For this we use a custom mypy plugin: https://github.com/typeddjango/django-stubs/blob/c9c729073417d0936cb944ab8585ad236ab30321/mypy_django_plugin/transformers/orm_lookups.py#L10
What it does?
- It checks that simple keyword arguments are indeed the correct ones
- It checks that nested `__` ones also exist on the nested model
- It still allows `**custom_data` unpacking
- It generates an error that can be silenced with `type: ignore[custom-code]`
- All checks like this can be turned off when `custom-code` is disabled in mypy checks
- It does not affect anything else
- It slows down type-checking a bit for users who added this plugin
- For users without a plugin - nothing happens
- Pyright and PyCharm are unaffected
- It is better to bundle this plugin, but it can be a 3rd party (really hard to maintain)
Plus, in the future more goodies can be added, included DI proper checking, URL params, etc.
It will require its own set of tests via `typing.assert_type` and maintaince time. Mypy sometimes break plugin-facing APIs.
I can write this plugin if others are interested :)
### Basic Example
_No response_
### Drawbacks and Impact
_No response_
### Unresolved questions
_No response_ | open | 2024-10-16T09:31:03Z | 2025-03-20T15:55:00Z | https://github.com/litestar-org/litestar/issues/3814 | [
"Enhancement",
"Typing",
"DTOs"
] | sobolevn | 0 |
graphdeco-inria/gaussian-splatting | computer-vision | 720 | how to construct our own dataset as input for 3d-GS from images taken by a phone | Hi,
Thanks for your great work. I'd like to try your pipeline on my own dataset. I took a few images and wanted to use colmap to obtain the necessary files as in your dataset. When I run your python file "convert.py", the output files were totally different from yours. The images taken are stored in this format: ./360_v2/bottles/input . Is there anything wrong with this data format?
thx for your time
best,
| open | 2024-03-21T10:02:58Z | 2024-03-21T10:02:58Z | https://github.com/graphdeco-inria/gaussian-splatting/issues/720 | [] | Ericgone | 0 |
django-cms/django-cms | django | 7,482 | [BUG] Wizard create page doesnt work | ## Description
When i start new. I get the wizard with 'new page'. I get the message in red "Please choose an option from below to proceed to the next step.".
## Steps to reproduce
I used this docs: https://django-cms-docs.readthedocs.io/en/latest/how_to/01-install.html
To setup django-cms v4.
## Expected behaviour
That when you choose a option in the wizard the form comes in front.
## Actual behaviour
When choosing an option i get in red "Please choose an option from below to proceed to the next step."
## Screenshots
<img width="1375" alt="Schermafbeelding 2023-01-22 om 12 01 49" src="https://user-images.githubusercontent.com/34129243/213912639-352a1761-08a6-4c2a-92cb-31aad98bd552.png">
## Additional information (CMS/Python/Django versions)
Python 3.9
Django 4.1.5
Django CMS 4.1.0rc1
## Do you want to help fix this issue?
* [ + ] Yes, I want to help fix this issue and I will join #workgroup-pr-review on [Slack](https://www.django-cms.org/slack) to confirm with the community that a PR is welcome.
* [ ] No, I only want to report the issue.
| closed | 2023-01-22T11:09:12Z | 2023-01-28T13:36:56Z | https://github.com/django-cms/django-cms/issues/7482 | [] | svandeneertwegh | 1 |
microsoft/MMdnn | tensorflow | 608 | pytorch to IR error?? | Platform (like ubuntu 16.04/win10):
Ubuntu 16.04.6 LTS
Python version:
2.7
Source framework with version (like Tensorflow 1.4.1 with GPU):
1.13.1with GPU
Destination framework with version (like CNTK 2.3 with GPU):
pytorch verson: 1.0.1.post2
Pre-trained model path (webpath or webdisk path):
mmdownload -f pytorch -n resnet101 -o ./
Running scripts:
mmtoir -f pytorch -d resnet101 --inputShape 3,224,224 -n imagenet_resnet101.pth
mmdnn setup: pip install -U git+https://github.com/Microsoft/MMdnn.git@master
mmtoir -f pytorch -d resnet101 --inputShape 3,224,224 -n imagenet_resnet101.pth
Traceback (most recent call last):
File "/home/luna/.local/bin/mmtoir", line 10, in <module>
sys.exit(_main())
File "/home/luna/.local/lib/python2.7/site-packages/mmdnn/conversion/_script/convertToIR.py", line 192, in _main
ret = _convert(args)
File "/home/luna/.local/lib/python2.7/site-packages/mmdnn/conversion/_script/convertToIR.py", line 92, in _convert
parser = PytorchParser(model, inputshape[0])
File "/home/luna/.local/lib/python2.7/site-packages/mmdnn/conversion/pytorch/pytorch_parser.py", line 85, in __init__
self.pytorch_graph.build(self.input_shape)
File "/home/luna/.local/lib/python2.7/site-packages/mmdnn/conversion/pytorch/pytorch_graph.py", line 124, in build
trace.set_graph(PytorchGraph._optimize_graph(trace.graph(), False))
File "/home/luna/.local/lib/python2.7/site-packages/mmdnn/conversion/pytorch/pytorch_graph.py", line 74, in _optimize_graph
graph = torch._C._jit_pass_onnx(graph, aten)
TypeError: _jit_pass_onnx(): incompatible function arguments. The following argument types are supported:
1. (arg0: torch::jit::Graph, arg1: torch._C._onnx.OperatorExportTypes) -> torch::jit::Graph
| open | 2019-03-07T06:31:41Z | 2019-06-26T15:19:47Z | https://github.com/microsoft/MMdnn/issues/608 | [] | lunalulu | 16 |
flasgger/flasgger | api | 517 | swag_from did not update components.schemas - OpenAPI 3.0 | Hi
I am converting a project OpenAPI from v2.0 to v3.0
the problem is : `swag_from` will not update `components.schemas` when I load an additional yaml file (previously with v2.0, where we had `definitions`, everything worked fine and `definitions` will updated by `swag_from` from extra yml file)
here is a simple code to reproduce this issue
`swagger_all.yml`
```yml
openapi: 3.0.1
info:
title: Test
description: Testing
version: 1.0.0
paths: {}
# paths:
# /get_cost:
# post:
# summary: a test with cascading $refs
# requestBody:
# description: request
# content:
# application/json:
# schema:
# $ref: '#/components/schemas/GetCostRequest'
# required: true
# responses:
# 200:
# description: OK
# content:
# application/json:
# schema:
# type: array
# items:
# $ref: '#/components/schemas/GetCostResponse'
# 201:
# description: Created
# content: {}
# 401:
# description: Unauthorized
# content: {}
components:
schemas:
Cost:
title: Cost
type: object
properties:
currency:
type: string
description: cost currency (3-letters code)
value:
type: number
description: cost value
GeoPosition:
title: GeoPosition
type: object
properties:
latitude:
type: number
description: latitude in float
format: double
longitude:
type: number
description: longitude in float
format: double
# GetCostRequest:
# title: GetCost Request
# type: object
# properties:
# level:
# type: integer
# location:
# $ref: '#/components/schemas/Location'
# GetCostResponse:
# title: GetCost response
# type: object
# properties:
# cost:
# $ref: '#/components/schemas/Cost'
# description:
# type: string
Location:
title: Location
type: object
properties:
name:
type: string
description: name of the location
position:
$ref: '#/components/schemas/GeoPosition'
```
and `extra.yml`
```yml
summary: a test
requestBody:
description: request
content:
application/vnd.api+json:
schema:
$ref: '#/components/schemas/GetCostRequest'
required: true
responses:
200:
description: OK
content:
application/json:
schema:
type: array
items:
$ref: '#/components/schemas/GetCostResponse'
201:
description: Created
content: {}
401:
description: Unauthorized
content: {}
components:
schemas:
GetCostRequest:
title: GetCost Request
type: object
properties:
level:
type: integer
location:
$ref: '#/components/schemas/Location'
GetCostResponse:
title: GetCost response
type: object
properties:
cost:
$ref: '#/components/schemas/Cost'
description:
type: string
```
and the python code :
```py
from flask import Flask, jsonify, Blueprint
from flasgger import swag_from, Swagger
try:
import simplejson as json
except ImportError:
import json
app = Flask(__name__)
app.config['SWAGGER'] = {'openapi': '3.0.1','uiversion': 3}
swagger = Swagger(app, template_file='swagger_all.yml')
api = Blueprint('api', __name__, url_prefix='/api')
@api.route('/get_cost', methods=['POST'])
@swag_from(specs='extra.yml',validation=True)
def get_cost():
result = dict(description='The best place',
cost=dict(currency='EUR', value=123456))
return jsonify([result])
app.register_blueprint(api)
if __name__ == '__main__':
app.run(debug=True)
```
when I look at to the generated `apispec_1.json` file, the input entry `components.schemas` did not updated by `swag_from` (only `paths` updated), and contains only inputs from file passed by `template_file`
here is the error I am getting
```bash
Errors
Resolver error at paths./api/get_cost.post.requestBody.content.application/vnd.api+json.schema.$ref
Could not resolve reference: Could not resolve pointer: /components/schemas/GetCostRequest does not exist in document
Resolver error at paths./api/get_cost.post.responses.200.content.application/json.schema.items.$ref
Could not resolve reference: Could not resolve pointer: /components/schemas/GetCostResponse does not exist in document
```
any suggestion ?
| open | 2022-01-14T14:58:46Z | 2022-03-10T05:43:20Z | https://github.com/flasgger/flasgger/issues/517 | [] | arabnejad | 2 |
pyjanitor-devs/pyjanitor | pandas | 959 | Extend select_columns to groupby objects | # Brief Description
Allow column selection on pandas dataframe groupby objects with `select_columns`
# Example API
``mtcars.groupby('cyl').select_columns('*p')``
| closed | 2021-11-27T03:12:29Z | 2021-12-11T10:27:08Z | https://github.com/pyjanitor-devs/pyjanitor/issues/959 | [] | samukweku | 2 |
MilesCranmer/PySR | scikit-learn | 424 | [BUG]: PySR runs well once and then stops after error | ### What happened?
Hello,
I was trying to use PySR and I ran into a problem: I ran it once and the model was able to identify the equation correctly. However, after trying to run my code on other data, nothing happens but the code stops at the following error (see below)
I am not sure if I am causing this problem or what the problem could be. I am running the code in Python 3.11.0 and Julia 1.8.5. If there is already an issue that would help, then sorry for posting the same question twice. I hope that you can help me in resolving this problem.
Best wishes,
Bartosz
### Version
0.16.3
### Operating System
Windows
### Package Manager
pip
### Interface
Jupyter Notebook
### Relevant log output
```shell
UserWarning Traceback (most recent call last)
Cell In[45], line 19
1 from pysr import PySRRegressor
3 model = PySRRegressor(
4 niterations=40, # < Increase me for better results
5 binary_operators=["+", "*", "-"],
(...)
17 progress=False
18 )
---> 19 model.fit(x_train_ic,x_dot)
File ~\Anaconda3\envs\tristan\Lib\site-packages\pysr\sr.py:1904, in PySRRegressor.fit(self, X, y, Xresampled, weights, variable_names, X_units, y_units)
1900 seed = random_state.get_state()[1][0] # For julia random
1902 self._setup_equation_file()
-> 1904 mutated_params = self._validate_and_set_init_params()
1906 (
1907 X,
1908 y,
(...)
1915 X, y, Xresampled, weights, variable_names, X_units, y_units
1916 )
1918 if X.shape[0] > 10000 and not self.batching:
File ~\Anaconda3\envs\tristan\Lib\site-packages\pysr\sr.py:1346, in PySRRegressor._validate_and_set_init_params(self)
1344 parameter_value = 1
1345 elif parameter == "progress" and not buffer_available:
-> 1346 warnings.warn(
1347 "Note: it looks like you are running in Jupyter. "
1348 "The progress bar will be turned off."
1349 )
1350 parameter_value = False
1351 packed_modified_params[parameter] = parameter_value
UserWarning: Note: it looks like you are running in Jupyter. The progress bar will be turned off.
```
### Extra Info
This the minimal example, the x_train_ic is just a time series and x_dot the derivatives of it.
```
from pysr import PySRRegressor
model = PySRRegressor(
niterations=40, # < Increase me for better results
binary_operators=["+", "*", "-"],
#unary_operators=[
# "cos",
# "exp",
# "sin",
# "inv(x) = 1/x",
# ^ Custom operator (julia syntax)
#],
#extra_sympy_mappings={"inv": lambda x: 1 / x},
# ^ Define operator for SymPy as well
loss="loss(prediction, target) = (prediction - target)^2",
# ^ Custom loss function (julia syntax)
progress=False
)
model.fit(x_train_ic,x_dot)
``` | open | 2023-09-13T13:51:57Z | 2023-09-13T18:34:50Z | https://github.com/MilesCranmer/PySR/issues/424 | [
"bug"
] | BMP-TUD | 1 |
PaddlePaddle/PaddleNLP | nlp | 9,482 | [Docs]:预测demo中加载了两次模型参数,不符合逻辑 | ### 软件环境
```Markdown
- paddlepaddle:
- paddlepaddle-gpu:
- paddlenlp:
```
### 详细描述
```Markdown
这个文档里,predict时加载了两次模型参数,第一次是原始模型,第二次是训练后的参数,按理说,只需要加载训练后的参数即可,是不是可以再完善一下
```
| closed | 2024-11-22T09:03:43Z | 2025-02-05T00:20:47Z | https://github.com/PaddlePaddle/PaddleNLP/issues/9482 | [
"documentation",
"stale"
] | williamPENG1 | 6 |
jupyter/nbviewer | jupyter | 703 | Markdown rendering issue for ipython notebooks in Github but not nbviewer | (I wasn't certain where to raise this issue but the nbviewer blog recommended this repo.).
**Issue**: For ipython notebooks viewed in Github (but not nbviewer), if there is any Markdown-formatted text nested between inline LaTeX _within the same paragraph block_, the Markdown formatting does not render correctly.
For example, take a look at this [ipython notebook](https://github.com/redwanhuq/machine-learning/blob/master/sms_spam_filter.ipynb). If you search for the word "subsets", you'll notice that the 1st example displays as "\<em>subsets\</em>", instead of italicized Markdown rendering. (FYI in the ipython notebook editor, I'm using the proper Markdown syntax, i.e., \*subsets*) Whereas, the same notebook in nbviewer [doesn't exhibit this issue](http://nbviewer.jupyter.org/github/redwanhuq/machine-learning/blob/master/sms_spam_filter.ipynb).
Oddly, if the Markdown-formatted text is flanked by inline LaTeX either before or after (but not both) within the same paragraph block, then the issue doesn't appear at all in Github viewer. | closed | 2017-06-15T14:12:38Z | 2017-06-23T23:39:00Z | https://github.com/jupyter/nbviewer/issues/703 | [] | redwanhuq | 3 |
sigmavirus24/github3.py | rest-api | 403 | Bug in Feeds API | In line 361-362 of `github.py`
```
for d in links.values():
d['href'] = URITemplate(d['href'])
```
there is a bug. When user have no `current_user_organization_url` or `current_user_organization_urls` or something like this ,the d will be blank array `[]`. So this would throw an error of `TypeError: list indices must be integers, not unicode`
So I add a line between them like this `if d:` . It will work well.
| closed | 2015-06-30T13:57:50Z | 2015-11-08T00:25:37Z | https://github.com/sigmavirus24/github3.py/issues/403 | [] | jzau | 1 |
sammchardy/python-binance | api | 773 | Failed to parse error [SOLVED] | My code works fine If I'm running it using VScode with the Python virtualenv
Python version: [3.8.5] 64-bit
however if I run the same code in the terminal using Python [3.8.5] 64-bit
I get this error here
nick-pc@nickpc-HP-xw4300-Workstation:~/Documents/pythonprojects/DCAbot$ /usr/bin/python3 /home/nick-pc/Documents/pythonprojects/DCAbot/DCAETHbot.py
Traceback (most recent call last):
File "/home/nick-pc/.local/lib/python3.8/site-packages/requests/models.py", line 382, in prepare_url
scheme, auth, host, port, path, query, fragment = parse_url(url)
File "/usr/lib/python3/dist-packages/urllib3/util/url.py", line 392, in parse_url
return six.raise_from(LocationParseError(source_url), None)
File "<string>", line 3, in raise_from
urllib3.exceptions.LocationParseError: Failed to parse: https://api.binance.com/api/v3/ping
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "/home/nick-pc/Documents/pythonprojects/DCAbot/DCAETHbot.py", line 8, in <module>
client = Client(keys.api_key, keys.api_secret)
File "/home/nick-pc/.local/lib/python3.8/site-packages/binance/client.py", line 105, in __init__
self.ping()
File "/home/nick-pc/.local/lib/python3.8/site-packages/binance/client.py", line 392, in ping
return self._get('ping', version=self.PRIVATE_API_VERSION)
File "/home/nick-pc/.local/lib/python3.8/site-packages/binance/client.py", line 237, in _get
return self._request_api('get', path, signed, version, **kwargs)
File "/home/nick-pc/.local/lib/python3.8/site-packages/binance/client.py", line 202, in _request_api
return self._request(method, uri, signed, **kwargs)
File "/home/nick-pc/.local/lib/python3.8/site-packages/binance/client.py", line 196, in _request
self.response = getattr(self.session, method)(uri, **kwargs)
File "/home/nick-pc/.local/lib/python3.8/site-packages/requests/sessions.py", line 555, in get
return self.request('GET', url, **kwargs)
File "/home/nick-pc/.local/lib/python3.8/site-packages/requests/sessions.py", line 528, in request
prep = self.prepare_request(req)
File "/home/nick-pc/.local/lib/python3.8/site-packages/requests/sessions.py", line 456, in prepare_request
p.prepare(
File "/home/nick-pc/.local/lib/python3.8/site-packages/requests/models.py", line 316, in prepare
self.prepare_url(url, params)
File "/home/nick-pc/.local/lib/python3.8/site-packages/requests/models.py", line 384, in prepare_url
raise InvalidURL(*e.args)
requests.exceptions.InvalidURL: Failed to parse: https://api.binance.com/api/v3/ping
- Python version: [3.8.5] 64-bit
- Virtual Env: [virtualenv]
- OS: [Ubuntu]
- python-binance version [0.7.9]
| closed | 2021-04-18T04:27:02Z | 2021-05-11T12:43:30Z | https://github.com/sammchardy/python-binance/issues/773 | [] | fuzzybannana | 1 |
mars-project/mars | pandas | 2,584 | [BUG] mars.dataframe.DataFrame.loc[i:j] semantics is different with pandas | # Reporting a bug
```
import pandas as pd
import mars
import numpy as np
df = pd.DataFrame(np.random.rand(5,3))
sliced_df = df.loc[0:1]
# Out[6]: sliced_df
# 0 1 2
# 0 0.362741 0.466188 0.750695
# 1 0.775940 0.544655 0.711621
mars.new_session()
md = mars.dataframe.DataFrame(np.random.rand(5, 3))
sliced_md = md.loc[0:1]
sliced_md.execute()
# Out[14]: sliced_md
# 0 1 2
# 0 0.851917 0.508231 0.908007
```
As you can see, mars choose `right open`, while pandas choose `right closed`. They contain different behaviours.
Python 3.7.9 & Pandas 1.2.0 & mars 0.7.5 | closed | 2021-11-24T06:33:29Z | 2022-09-05T03:26:57Z | https://github.com/mars-project/mars/issues/2584 | [
"type: bug",
"mod: dataframe"
] | dlee992 | 0 |
coqui-ai/TTS | pytorch | 4,043 | [Feature request] Support for Quantized ONNX Model Conversion for Stream Inference | <!-- Welcome to the 🐸TTS project!
We are excited to see your interest, and appreciate your support! --->
**🚀 Feature Description**
Is there support in Coqui TTS for converting models to a quantized ONNX format for stream inference? This feature would enhance model performance and reduce inference time for real-time applications.
**Solution**
Implement a workflow or tool within Coqui TTS for easy conversion of TTS models to quantized ONNX format.
**Alternative Solutions**
Currently, external tools like ONNX Runtime or TensorRT can be used for post-conversion quantization, but having this feature natively would streamline the process.
**Additional context**
Any existing documentation or insights on this topic would be appreciated. Thank you!
| closed | 2024-11-02T04:01:41Z | 2024-12-28T11:58:22Z | https://github.com/coqui-ai/TTS/issues/4043 | [
"wontfix",
"feature request"
] | TranDacKhoa | 1 |
modin-project/modin | data-science | 7,405 | BUG: incorrect iloc behavior in modin when assigning index values based on row indices | ### Modin version checks
- [X] I have checked that this issue has not already been reported.
- [X] I have confirmed this bug exists on the latest released version of Modin.
- [ ] I have confirmed this bug exists on the main branch of Modin. (In order to do this you can follow [this guide](https://modin.readthedocs.io/en/stable/getting_started/installation.html#installing-from-the-github-main-branch).)
### Reproducible Example
```python
import pandas as pd
import modin.pandas as mpd
dict1 = {
'index_test': [-1, -1, -1]
}
df1 = pd.DataFrame(dict1)
mdf1 = mpd.DataFrame(dict1)
row_indices = [2, 0]
df1.iloc[row_indices, 0] = df1.iloc[row_indices].index
mdf1.iloc[row_indices, 0] = mdf1.iloc[row_indices].index
print(df1) # as expected: 0, -1, 2
print('-------------')
print(mdf1) # NOT as expected: 2, -1, 0
# index_test
# 0 0
# 1 -1
# 2 2
# -------------
# index_test
# 0 2
# 1 -1
# 2 0
```
### Issue Description
When assigning values using iloc in modin, the behavior deviates from the expected behavior seen with pandas. Specifically, assigning index values to a subset of rows works correctly in pandas, but modin assigns values in wrong order.
### Expected Behavior
This issue occurs consistently when trying to assign values based on row indices using iloc in modin. The expected behavior is for modin to mirror pandas behavior, but instead, the values are assigned in a different order.
expected output produced with pandas:
```
index_test
0 0
1 -1
2 2
```
actual output produced with modin:
```
index_test
0 2
1 -1
2 0
```
### Error Logs
_No response_
### Installed Versions
<details>
PyDev console: using IPython 8.23.0
INSTALLED VERSIONS
------------------
commit : 3e951a63084a9cbfd5e73f6f36653ee12d2a2bfa
python : 3.11.8
python-bits : 64
OS : Windows
OS-release : 10
Version : 10.0.22631
machine : AMD64
processor : Intel64 Family 6 Model 186 Stepping 2, GenuineIntel
byteorder : little
LC_ALL : None
LANG : None
LOCALE : English_Austria.1252
Modin dependencies
------------------
modin : 0.32.0
ray : 2.20.0
dask : 2024.5.2
distributed : 2024.5.2
pandas dependencies
-------------------
pandas : 2.2.3
numpy : 1.26.4
pytz : 2024.1
dateutil : 2.9.0.post0
pip : 24.2
Cython : 3.0.10
sphinx : None
IPython : 8.23.0
adbc-driver-postgresql: None
adbc-driver-sqlite : None
bs4 : 4.12.3
blosc : None
bottleneck : 1.3.7
dataframe-api-compat : None
fastparquet : None
fsspec : 2023.10.0
html5lib : None
hypothesis : None
gcsfs : None
jinja2 : 3.1.3
lxml.etree : None
matplotlib : 3.8.2
numba : None
numexpr : 2.8.7
odfpy : None
openpyxl : None
pandas_gbq : None
psycopg2 : None
pymysql : None
pyarrow : 14.0.2
pyreadstat : None
pytest : 8.1.1
python-calamine : None
pyxlsb : None
s3fs : None
scipy : 1.14.0
sqlalchemy : 2.0.25
tables : 3.9.2
tabulate : 0.9.0
xarray : None
xlrd : None
xlsxwriter : None
zstandard : 0.23.0
tzdata : 2024.1
qtpy : 2.4.1
pyqt5 : None
</details>
| closed | 2024-10-14T07:13:20Z | 2025-02-27T19:59:55Z | https://github.com/modin-project/modin/issues/7405 | [
"bug 🦗",
"P1"
] | SchwurbeI | 3 |
fastapi/sqlmodel | pydantic | 75 | Add sessionmaker | ### First Check
- [X] I added a very descriptive title to this issue.
- [X] I used the GitHub search to find a similar issue and didn't find it.
- [X] I searched the SQLModel documentation, with the integrated search.
- [X] I already searched in Google "How to X in SQLModel" and didn't find any information.
- [X] I already read and followed all the tutorial in the docs and didn't find an answer.
- [X] I already checked if it is not related to SQLModel but to [Pydantic](https://github.com/samuelcolvin/pydantic).
- [X] I already checked if it is not related to SQLModel but to [SQLAlchemy](https://github.com/sqlalchemy/sqlalchemy).
### Commit to Help
- [X] I commit to help with one of those options 👆
### Example Code
```python
Session = sessionmaker(engine)
```
### Description
Add an sqlalchemy compatible sessionmaker that generates SqlModel sessions
### Wanted Solution
I would like to have a working sessionmaker
### Wanted Code
```python
from sqlmodel import sessionmaker
```
### Alternatives
_No response_
### Operating System
macOS
### Operating System Details
_No response_
### SQLModel Version
0.0.4
### Python Version
3.9.6
### Additional Context
_No response_ | open | 2021-09-02T21:00:03Z | 2024-05-14T11:03:00Z | https://github.com/fastapi/sqlmodel/issues/75 | [
"feature"
] | hitman-gdg | 6 |
junyanz/pytorch-CycleGAN-and-pix2pix | deep-learning | 1,119 | Always out of memory when testing | Hi,
My testing set has about 7,000 images in all. Some images in the testing set are very large, like 2,000*3,000 pixels. The memory is always overflow. The testing program can only run on one gpu instead of multi-gpu. How can I fix this problem? Many thanks! | closed | 2020-08-06T13:47:36Z | 2020-08-07T04:12:09Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/1119 | [] | GuoLanqing | 2 |
gradio-app/gradio | python | 10,564 | Misplaced Chat Avatar While Thinking | ### Describe the bug
When the chatbot is thinking, the Avatar icon is misplaced. When it is actually inferencing or done inferencing, the avatar is fine.
Similar to https://github.com/gradio-app/gradio/issues/9655 I believe, but a special edge case. Also, I mostly notice the issue with rectangular images.
### Have you searched existing issues? 🔎
- [x] I have searched and found no existing issues
### Reproduction
```python
import gradio as gr
from time import sleep
AVATAR = "./car.png"
# Define a simple chatbot function
def chatbot_response(message, hist):
sleep(10)
return f"Gradio is pretty cool!"
# Create a chat interface using gr.ChatInterface
chatbot = gr.ChatInterface(fn=chatbot_response,
chatbot=gr.Chatbot(
label="LLM",
elem_id="chatbot",
avatar_images=(
None,
AVATAR
),
)
)
# Launch the chatbot
chatbot.launch()
```
### Screenshot



### Logs
```shell
```
### System Info
```shell
(base) carter.yancey@Yancy-XPS:~$ gradio environment
Gradio Environment Information:
------------------------------
Operating System: Linux
gradio version: 5.13.1
gradio_client version: 1.6.0
------------------------------------------------
gradio dependencies in your environment:
aiofiles: 23.2.1
anyio: 3.7.1
audioop-lts is not installed.
fastapi: 0.115.7
ffmpy: 0.3.2
gradio-client==1.6.0 is not installed.
httpx: 0.25.1
huggingface-hub: 0.27.1
jinja2: 3.1.2
markupsafe: 2.1.3
numpy: 1.26.2
orjson: 3.9.10
packaging: 23.2
pandas: 1.5.3
pillow: 10.0.0
pydantic: 2.5.1
pydub: 0.25.1
python-multipart: 0.0.20
pyyaml: 6.0.1
ruff: 0.2.2
safehttpx: 0.1.6
semantic-version: 2.10.0
starlette: 0.45.3
tomlkit: 0.12.0
typer: 0.15.1
typing-extensions: 4.8.0
urllib3: 2.3.0
uvicorn: 0.24.0.post1
authlib; extra == 'oauth' is not installed.
itsdangerous; extra == 'oauth' is not installed.
gradio_client dependencies in your environment:
fsspec: 2023.10.0
httpx: 0.25.1
huggingface-hub: 0.27.1
packaging: 23.2
typing-extensions: 4.8.0
websockets: 11.0.3
```
### Severity
I can work around it | closed | 2025-02-11T18:31:28Z | 2025-03-04T21:23:07Z | https://github.com/gradio-app/gradio/issues/10564 | [
"bug",
"💬 Chatbot"
] | CarterYancey | 0 |
ultralytics/yolov5 | machine-learning | 13,447 | training stuck | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
I used your framework to modify yolo and define a model by myself, but when I was training, why did I get stuck at the beginning, in the position shown below

### Additional
_No response_ | closed | 2024-12-06T06:35:00Z | 2024-12-06T10:43:26Z | https://github.com/ultralytics/yolov5/issues/13447 | [
"question"
] | passingdragon | 3 |
Kanaries/pygwalker | matplotlib | 670 | [BUG] Installation from conda-forge yields No module named 'lib2to3' | **Describe the bug**
When installing pygwalker via conda, the actual installation works, but subsequent import yields:
```sh
----> 1 import pygwalker as pyg
File ~/miniforge3/envs/scratchpad/lib/python3.13/site-packages/pygwalker/__init__.py:16
13 __version__ = "0.3.17"
14 __hash__ = __rand_str()
---> 16 from pygwalker.api.walker import walk
17 from pygwalker.api.gwalker import GWalker
18 from pygwalker.api.html import to_html
File ~/miniforge3/envs/scratchpad/lib/python3.13/site-packages/pygwalker/api/walker.py:10
8 from pygwalker.data_parsers.database_parser import Connector
9 from pygwalker._typing import DataFrame
---> 10 from pygwalker.services.format_invoke_walk_code import get_formated_spec_params_code_from_frame
11 from pygwalker.services.kaggle import auto_set_kanaries_api_key_on_kaggle, adjust_kaggle_default_font_size
12 from pygwalker.utils.execute_env_check import check_convert, get_kaggle_run_type, check_kaggle
File ~/miniforge3/envs/scratchpad/lib/python3.13/site-packages/pygwalker/services/format_invoke_walk_code.py:3
1 from typing import Optional, List, Any
2 from types import FrameType
----> 3 from lib2to3 import fixer_base, refactor
4 import logging
5 import inspect
ModuleNotFoundError: No module named 'lib2to3'
```
**To Reproduce**
```
$ conda install pygwalker
... # open REPL
>>> import pygwalker
```
**Expected behavior**
Import should work
**Versions**
- pygwalker version: 0.3.17
- python version: 3.13.1
- browser
**Additional context**
A pip install seems to be successful, and installs a newer version (0.4.9.13) so I'm guessing the conda-forge recipe is just outdated?
| open | 2024-12-19T16:37:40Z | 2025-02-17T02:16:55Z | https://github.com/Kanaries/pygwalker/issues/670 | [
"bug"
] | WillAyd | 1 |
onnx/onnx | deep-learning | 6,364 | Sonarcloud for static code analysis? | ### System information
_No response_
### What is the problem that this feature solves?
Introduction of sonarcloud
### Alternatives considered
Focus on codeql ?
### Describe the feature
Thanks to the improvements made by @cyyever I wonder if we want to officially set up a tool like Sonarcloud, for example. ( I could do that)
For a fork of mine, for example, it looks like this:
https://sonarcloud.io/project/issues?rules=python%3AS6711&issueStatuses=OPEN%2CCONFIRMED&id=andife_onnx&open=AZHq5D8n6JXh0XXyfRwb&tab=code
(My general experience with sonarcloud/sonarqube has been very positive)
Is the codeql integrated in github systematically used so far?
I know different static linkers produce different results and blindly following the suggestions does not necessarily lead to better code quality.
A comparison can be found at https://medium.com/@suthakarparamathma/sonarqube-vs-codeql-code-quality-tool-comparison-32395f2a77b3
### Will this influence the current api (Y/N)?
no
### Feature Area
best practices, code quality
### Are you willing to contribute it (Y/N)
Yes
### Notes
I could create it for our regulate onnx/onnx branch. It is free for open source projects
https://www.sonarsource.com/plans-and-pricing/ | open | 2024-09-14T16:01:12Z | 2024-09-25T04:41:40Z | https://github.com/onnx/onnx/issues/6364 | [
"topic: enhancement"
] | andife | 4 |
Nemo2011/bilibili-api | api | 165 | 将获取 HTML 内 JSON 信息的操作单独提出成函数 | 指的是
```python
try:
resp = await session.get(
f"https://www.bilibili.com/bangumi/play/ep{epid}",
cookies=credential.get_cookies(),
headers={"User-Agent": "Mozilla/5.0"},
)
except Exception as e:
raise ResponseException(str(e))
else:
content = resp.text
pattern = re.compile(r"window.__INITIAL_STATE__=(\{.*?\});")
match = re.search(pattern, content)
if match is None:
raise ApiException("未找到番剧信息")
try:
content = json.loads(match.group(1))
except json.JSONDecodeError:
raise ApiException("信息解析错误")
return content
```
很显然这个重复的操作已经出现了很多次了...为什么没做成函数啊
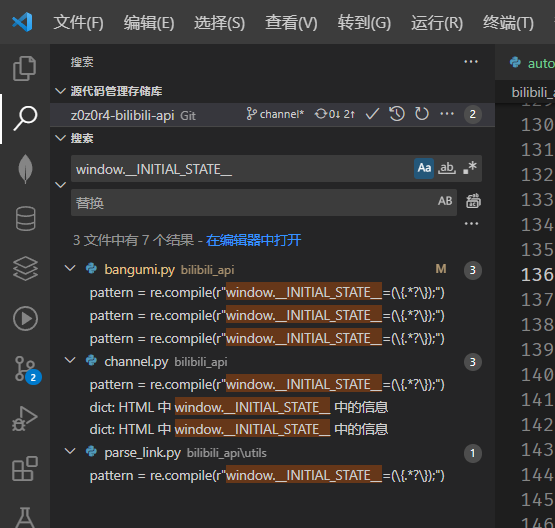
| closed | 2023-01-27T07:47:10Z | 2023-01-27T08:37:22Z | https://github.com/Nemo2011/bilibili-api/issues/165 | [
"need"
] | z0z0r4 | 4 |
piskvorky/gensim | nlp | 3,043 | gensim.scripts.word2vec2tensor results in UnicodeDecodeError | #### Problem description
- I created a word2vec model from the tokens read from 1.4L files using the following call
model.wv.save_word2vec_format(f"{folder}/wvmodel.wv", binary=True)
- Ran the following command to convert word-vectors from word2vec format into Tensorflow 2D tensor format
python -m gensim.scripts.word2vec2tensor -i model/wvmodel.wv -o model/ -b
The above command works for tokens read from 5000 files. But it fails when I read the tokens from 6000 files. Looks like there is some content in the one of the files (5000 to 6000) that the **word2vec2tensor** script has problems with.
Is there anyway I can fix this issue? Or atleast identify the offending file and remove it?
#### Steps/code/corpus to reproduce
Unfortunately I cannot share the dataset as it is huge.
2021-02-11 05:28:33,305 - utils_any2vec - INFO - loading projection weights from model/wvmodel.wv
**Traceback** (most recent call last):
File "/usr/local/lib/python3.9/runpy.py", line 197, in _run_module_as_main
return _run_code(code, main_globals, None,
File "/usr/local/lib/python3.9/runpy.py", line 87, in _run_code
exec(code, run_globals)
File "/usr/local/lib/python3.9/site-packages/gensim/scripts/word2vec2tensor.py", line 94, in <module>
word2vec2tensor(args.input, args.output, args.binary)
File "/usr/local/lib/python3.9/site-packages/gensim/scripts/word2vec2tensor.py", line 68, in word2vec2tensor
model = gensim.models.KeyedVectors.load_word2vec_format(word2vec_model_path, binary=binary)
File "/usr/local/lib/python3.9/site-packages/gensim/models/keyedvectors.py", line 1547, in load_word2vec_format
return _load_word2vec_format(
File "/usr/local/lib/python3.9/site-packages/gensim/models/utils_any2vec.py", line 285, in _load_word2vec_format
_word2vec_read_binary(fin, result, counts,
File "/usr/local/lib/python3.9/site-packages/gensim/models/utils_any2vec.py", line 204, in _word2vec_read_binary
processed_words, chunk = _add_bytes_to_result(
File "/usr/local/lib/python3.9/site-packages/gensim/models/utils_any2vec.py", line 186, in _add_bytes_to_result
word = chunk[start:i_space].decode("utf-8", errors=unicode_errors)
UnicodeDecodeError: 'utf-8' codec can't decode byte 0xbf in position 0: invalid start byte
#### Versions
Linux-4.14.214-160.339.amzn2.x86_64-x86_64-with-debian-10.6
Python 3.6.12 (default, Nov 18 2020, 14:46:32)
[GCC 8.3.0]
Bits 64
NumPy 1.19.5
SciPy 1.5.4
gensim 3.8.3
FAST_VERSION 1 | closed | 2021-02-11T07:32:37Z | 2021-02-12T09:34:08Z | https://github.com/piskvorky/gensim/issues/3043 | [] | sreedevigattu | 6 |
plotly/dash-table | dash | 762 | Let nully work for all data types | Currently, `Format(nully='N/A')` only works if the column type is explicitly set to `numeric` | open | 2020-04-23T00:32:37Z | 2020-04-23T00:32:37Z | https://github.com/plotly/dash-table/issues/762 | [] | chriddyp | 0 |
coqui-ai/TTS | pytorch | 3,591 | buyongle | 不用了
| closed | 2024-02-18T00:51:13Z | 2024-03-10T14:13:27Z | https://github.com/coqui-ai/TTS/issues/3591 | [
"feature request"
] | fanghaiquan1 | 1 |
scikit-tda/kepler-mapper | data-visualization | 97 | User Defined Cover | Hi there
I can see that there is a TODO to implement a cover defining API. I was wondering what is the best way of creating a user-defined cover at the moment (if it is possible at all). From what I can tell, we are currently restricted to a `(n_bins, overlap_perc)` method. Is it possible to define a cover explicity (for one or more dimensions in the lens), using cutoff values or similar (like, setting the maximum and minimum values of the covering space in each dimension)? I ask because in its current implementation I think the [non-]presence of an outlier can skew the covering space quite drastically
Let me know what my options are for the covering space. I would also be interested to know the status of the above TODO. More information as to how the cover class currently works might also be useful if I was going to write my own.
Thanks!
Edit: I've modified the code such that you can pass `kmapper.map` a `CoverBounds` variable.
`if CoverBounds == None:` Normal behavior
However, CoverBounds can also be a `(ndim_lens, 2)` array, with `min, max` for every dimension of your lens. If the default behavior is fine for a particular dimension, pass it `np.float('inf'), np.float('inf')`.
For example, if I have a lens in **R**2 and want to set the maximum and minimum of the second dimension to be 0 and 1, I can pass:
`mapper.map(CoverBounds = np.array([[np.float('inf'), np.float('inf')],[0,1]]))` and that should have the desired behavior.
Edit 2: Might change it so rather than `inf` detection, works off `None` detection in the `CoverBounds` array.
I think a system designed like this should produce exactly the same cover, independent of input data limits.
Devs - let me hear your thoughts on this - I can clean up and submit a pull request.
| closed | 2018-06-05T08:41:28Z | 2018-07-12T22:38:38Z | https://github.com/scikit-tda/kepler-mapper/issues/97 | [] | leesteinberg | 3 |
cupy/cupy | numpy | 8,281 | Discover cuTENSOR wheels when building CuPy | CuPy can utilize `cutensor-cuXX` packages at runtime but not at build time. It is better to support building CuPy using headers and libraries from these packages. | open | 2024-04-11T10:34:39Z | 2024-04-12T04:38:47Z | https://github.com/cupy/cupy/issues/8281 | [
"cat:enhancement",
"prio:medium"
] | kmaehashi | 0 |
xonsh/xonsh | data-science | 5,241 | Error `Bad file descriptor` in `prompt_toolkit` > 3.0.40 | Testing environment: macOS Sonoma 14.1.2
After a git checkout, or directory change I very randomly get the following error:
```xsh
Traceback (most recent call last):
File "/opt/homebrew/lib/python3.11/site-packages/xonsh/main.py", line 469, in main
sys.exit(main_xonsh(args))
^^^^^^^^^^^^^^^^
File "/opt/homebrew/lib/python3.11/site-packages/xonsh/main.py", line 513, in main_xonsh
shell.shell.cmdloop()
File "/opt/homebrew/lib/python3.11/site-packages/xonsh/ptk_shell/shell.py", line 401, in cmdloop
line = self.singleline(auto_suggest=auto_suggest)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/homebrew/lib/python3.11/site-packages/xonsh/ptk_shell/shell.py", line 369, in singleline
line = self.prompter.prompt(**prompt_args)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/homebrew/lib/python3.11/site-packages/prompt_toolkit/shortcuts/prompt.py", line 1026, in prompt
return self.app.run(
^^^^^^^^^^^^^
File "/opt/homebrew/lib/python3.11/site-packages/prompt_toolkit/application/application.py", line 998, in run
return asyncio.run(coro)
^^^^^^^^^^^^^^^^^
File "/opt/homebrew/Cellar/python@3.11/3.11.6_1/Frameworks/Python.framework/Versions/3.11/lib/python3.11/asyncio/runners.py", line 189, in run
with Runner(debug=debug) as runner:
File "/opt/homebrew/Cellar/python@3.11/3.11.6_1/Frameworks/Python.framework/Versions/3.11/lib/python3.11/asyncio/runners.py", line 63, in __exit__
self.close()
File "/opt/homebrew/Cellar/python@3.11/3.11.6_1/Frameworks/Python.framework/Versions/3.11/lib/python3.11/asyncio/runners.py", line 77, in close
loop.close()
File "/opt/homebrew/Cellar/python@3.11/3.11.6_1/Frameworks/Python.framework/Versions/3.11/lib/python3.11/asyncio/unix_events.py", line 68, in close
super().close()
File "/opt/homebrew/Cellar/python@3.11/3.11.6_1/Frameworks/Python.framework/Versions/3.11/lib/python3.11/asyncio/selector_events.py", line 91, in close
self._close_self_pipe()
File "/opt/homebrew/Cellar/python@3.11/3.11.6_1/Frameworks/Python.framework/Versions/3.11/lib/python3.11/asyncio/selector_events.py", line 99, in _close_self_pipe
self._ssock.close()
File "/opt/homebrew/Cellar/python@3.11/3.11.6_1/Frameworks/Python.framework/Versions/3.11/lib/python3.11/socket.py", line 503, in close
self._real_close()
File "/opt/homebrew/Cellar/python@3.11/3.11.6_1/Frameworks/Python.framework/Versions/3.11/lib/python3.11/socket.py", line 497, in _real_close
_ss.close(self)
OSError: [Errno 9] Bad file descriptor
Xonsh encountered an issue during launch
Failback to /bin/bash
The default interactive shell is now zsh.
To update your account to use zsh, please run `chsh -s /bin/zsh`.
For more details, please visit https://support.apple.com/kb/HT208050.
bash-3.2$
```
Don't know how to force reproduction of this bug since it occurs very randomly at different times (but pretty often) | closed | 2023-12-03T07:27:23Z | 2024-05-09T21:30:45Z | https://github.com/xonsh/xonsh/issues/5241 | [
"prompt-toolkit",
"upstream",
"threading"
] | doronz88 | 22 |
ipython/ipython | data-science | 14,303 | Unexpected exception formatting exception in Python 3.13.0a3 | I appreciate that Python 3.13 is still in alpha, but some incompatibility seems to have been introduced with the way that exception data is produced that causes `ipython`'s pretty execution formatting to fail, cause the raising of a separate "Unexpected exception formatting exception".
## Steps to reproduce
1) Build Python 3.13.0a3 from source and install it somewhere.
2) Create a venv using the new Python 3.13 interpreter.
3) Build the latest master branch of [`parso`](https://github.com/davidhalter/parso) from source and install it into the venv.
4) Install `ipython` using `pip`.
5) Run `ipython` in a way that triggers an exception (such as `ipython -c 'print(1/0)'`)
## Expected result
`ipython` should print a nicely formatted exception. For instance, on Python 3.12 the result is:
```
(venv_3.12) nicko@testvm ~ % ipython -c 'print(1/0)'
---------------------------------------------------------------------------
ZeroDivisionError Traceback (most recent call last)
Cell In[1], line 1
----> 1 print(1/0)
ZeroDivisionError: division by zero
```
## Actual result
It appears that `ipython`, or possibly the `executing` library, is choking on the stack data and generates an `Unexpected exception formatting exception` message:
```
(venv_3.13) nicko@testvm ~ % ipython -c 'print(1/0)'
Unexpected exception formatting exception. Falling back to standard exception
Traceback (most recent call last):
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/IPython/core/interactiveshell.py", line 3553, in run_code
exec(code_obj, self.user_global_ns, self.user_ns)
~~~~^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "<ipython-input-1-2fc232d1511a>", line 1, in <module>
print(1/0)
~^~
ZeroDivisionError: division by zero
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/IPython/core/interactiveshell.py", line 2144, in showtraceback
stb = self.InteractiveTB.structured_traceback(
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~^
etype, value, tb, tb_offset=tb_offset
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
)
^
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/IPython/core/ultratb.py", line 1435, in structured_traceback
return FormattedTB.structured_traceback(
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~^
self, etype, evalue, etb, tb_offset, number_of_lines_of_context
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
)
^
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/IPython/core/ultratb.py", line 1326, in structured_traceback
return VerboseTB.structured_traceback(
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~^
self, etype, value, tb, tb_offset, number_of_lines_of_context
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
)
^
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/IPython/core/ultratb.py", line 1173, in structured_traceback
formatted_exception = self.format_exception_as_a_whole(etype, evalue, etb, number_of_lines_of_context,
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
tb_offset)
^^^^^^^^^^
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/IPython/core/ultratb.py", line 1063, in format_exception_as_a_whole
self.get_records(etb, number_of_lines_of_context, tb_offset) if etb else []
~~~~~~~~~~~~~~~~^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/IPython/core/ultratb.py", line 1160, in get_records
res = list(stack_data.FrameInfo.stack_data(etb, options=options))[tb_offset:]
~~~~^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/stack_data/core.py", line 597, in stack_data
yield from collapse_repeated(
...<4 lines>...
)
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/stack_data/utils.py", line 83, in collapse_repeated
yield from map(mapper, original_group)
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/stack_data/core.py", line 587, in mapper
return cls(f, options)
~~~^^^^^^^^^^^^
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/stack_data/core.py", line 551, in __init__
self.executing = Source.executing(frame_or_tb)
~~~~~~~~~~~~~~~~^^^^^^^^^^^^^
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/executing/executing.py", line 283, in executing
assert_(new_stmts <= stmts)
~~~~~~~^^^^^^^^^^^^^^^^^^^^
File "/Users/nvansomeren/python_tests/venv_3.13/lib/python3.13/site-packages/executing/executing.py", line 80, in assert_
raise AssertionError(str(message))
AssertionError
```
| open | 2024-01-24T04:52:46Z | 2024-02-03T22:33:33Z | https://github.com/ipython/ipython/issues/14303 | [] | nickovs | 3 |
yeongpin/cursor-free-vip | automation | 60 | Where can I get the user's password? | I need the user's password to log in to the cursor, but I can't find where the user's password is stored | closed | 2025-02-12T11:07:57Z | 2025-02-13T11:44:28Z | https://github.com/yeongpin/cursor-free-vip/issues/60 | [] | mnguyen081002 | 16 |
deepset-ai/haystack | pytorch | 8,540 | Add a ranker component that uses an LLM to rerank documents | **Describe the solution you'd like**
I’d like to add a new ranker component that leverages a LLM to rerank retrieved documents based on their relevance to the query. This would better assess the quality of the top-ranked documents, helping ensure that only relevant results are given to the LLM to answer the question.
Additionally, having an ability for the LLM to choose how many documents to keep would also be nice. A sort of dynamic top-k if you will.
**Additional context**
We have started to employ this for some clients especially in situations where we need to provide extensive references. Basically for a given answer we need to provide all relevant documents that support the answer text. Having one reference in these situations is not enough. As a result in these situations we are willing to pay the extra cost to use an LLM to rerank and only keep the most relevant documents.
| open | 2024-11-12T14:59:54Z | 2025-01-23T09:48:44Z | https://github.com/deepset-ai/haystack/issues/8540 | [
"P3"
] | sjrl | 6 |
tflearn/tflearn | tensorflow | 1,166 | Xception Example model | I wish you will add more examples at tflearn/examples. It would be really cool if you add the Xception model as well. Instead of Keras, tflearn is much more convenient for me, I am not able to write Xception from scratch so i would grateful if you add it👍 💯💯 :) | open | 2021-05-24T13:53:29Z | 2021-05-24T13:53:29Z | https://github.com/tflearn/tflearn/issues/1166 | [] | KfurkK | 0 |
supabase/supabase-py | flask | 673 | Even though the row is deleted, it still appears as if it exists (Python) | I'm not sure if this is a bug or not, but I'll try to explain it as best I can with screenshots.
With the API I developed with FastAPI, I first pull reviews from Tripadvisor, analyze them, and then send them to two interconnected tables called reviews and analysis on Supabase.
<img width="615" alt="Screenshot 2024-01-22 at 23 52 54" src="https://github.com/supabase-community/supabase-py/assets/68559468/7ecc8b50-a22a-4753-861b-5eff65d1706e">
When the tables are both empty, I can insert comments into the tables when I first post them.
<img width="1377" alt="Screenshot 2024-01-22 at 23 54 22" src="https://github.com/supabase-community/supabase-py/assets/68559468/fb027f58-944b-4adf-95cb-600204869244">
But then, when I delete the comments in both tables via supabase UI and try to insert the same comments again via the API, I encounter the following error:
<img width="886" alt="Screenshot 2024-01-22 at 23 56 31" src="https://github.com/supabase-community/supabase-py/assets/68559468/ec61026a-1f92-4933-a704-ee0e4a8d17c1">
Even though both tables are empty, when I want to insert the same comment again, it says that there is already a row with the same ID. What could be the reason for this?
| closed | 2024-01-22T21:00:11Z | 2024-03-12T23:40:34Z | https://github.com/supabase/supabase-py/issues/673 | [] | cenkerozkan | 3 |
liangliangyy/DjangoBlog | django | 405 | 什么时候能够支持markdown呢? | <!--
如果你不认真勾选下面的内容,我可能会直接关闭你的 Issue。
提问之前,建议先阅读 https://github.com/ruby-china/How-To-Ask-Questions-The-Smart-Way
-->
**我确定我已经查看了** (标注`[ ]`为`[x]`)
- [ ] [DjangoBlog的readme](https://github.com/liangliangyy/DjangoBlog/blob/master/README.md)
- [ ] [配置说明](https://github.com/liangliangyy/DjangoBlog/blob/master/bin/config.md)
- [ ] [其他 Issues](https://github.com/liangliangyy/DjangoBlog/issues)
----
**我要申请请求技术支持** (标注`[ ]`为`[x]`)
- [ ] BUG 反馈
- [x] 添加新的特性或者功能
- [ ] 请求技术支持
| closed | 2020-06-01T12:48:09Z | 2020-06-02T14:50:25Z | https://github.com/liangliangyy/DjangoBlog/issues/405 | [] | a532233648 | 0 |
wagtail/wagtail | django | 12,627 | Ordering documents in search causes error | <!--
Found a bug? Please fill out the sections below. 👍
-->
### Issue Summary
Issue was discovered by editors when searching through uploaded documents with similar names. Attempt to order them by date has failed.
### Steps to Reproduce
1. Login to wagtail admin
2. Search for an existing document
3. In search results click Documents tab
4. Click `Created` to sort the documents
5. Error - Cannot sort search results
### Technical details
- Python version: 3.10.15
- Django version: 5.0 - 5.1
- Wagtail version: 6.0.6 - 6.3.1
- Browser version: Chrome 131
### Working on this
Error message suggests adding index.FilterField('created_at') to AbstractDocument model. Adding this line to a local instance in virtual environment has fixed the issue
| closed | 2024-11-25T15:32:37Z | 2025-01-13T12:13:11Z | https://github.com/wagtail/wagtail/issues/12627 | [
"type:Bug"
] | JuraZakarija | 2 |
laughingman7743/PyAthena | sqlalchemy | 27 | Ctrl-C while running query kills python session | Signal handling should be improved if possible, because both:
1. Being unable to abort at all, and
2. Abort at the cost of quitting a running REPL
are barely acceptable for interactive usage. | closed | 2018-03-16T15:15:48Z | 2018-03-16T21:45:00Z | https://github.com/laughingman7743/PyAthena/issues/27 | [] | memeplex | 3 |
plotly/dash-table | plotly | 249 | Select all rows | I don't think its possible to select all rows in the table / filtered view.
Is this something that can be added?
Thanks! And thanks for all your work on the project - excited to see how it develops | open | 2018-11-20T19:31:24Z | 2022-07-11T13:15:06Z | https://github.com/plotly/dash-table/issues/249 | [
"dash-type-enhancement",
"size: 2"
] | pmajmudar | 13 |
marcomusy/vedo | numpy | 576 | Snapping multiple meshes together and extract transformation matrices | Hi @marcomusy,
I have the following problem that I am trying to address and I am trying to figure out how possibly I could automate the whole procedure. Imagine that I have multiple pieces of a complete object which are randomly given as input (different orientation, position, etc) and then I would like to find a way to automatize (not perfectly) how to assemble them all together to the final object and extract the transformation matrices. So imagine that I have the following 5 pieces:

which if you put them together in the correct order you should get the following complete object:

Currently someone could do that manually by using a corresponding 3d analysis tool, e.g. Meshlab, CloudCompare, Blender, Meshmixer, etc... and as I did. However, this takes a lot of time especially if you plan to do it for multiple objects and moreover the result still might not be the best. Thus, I wanted to ask you from your experience if you know any tool that could help me on that or if you believe I could do something with vedo.
My idea would be to extract some kind of boundary/shape contours and try to apply some kind of shape fitting metric or something similar but I am not sure what that could be. I've found your discussion here about [shape decomposition](https://github.com/marcomusy/vedo/issues/39) but I am not sure whether this could be related or not. I've tried to apply and test with different aligning algorithms but these are not working properly since they look for similar features that overlay each other while in this case I am looking for features that complement each other instead.
Any idea is welcome.
p.s. actually even an easier interactive mode, where I can select whether two edges should snap together would be helpful in regards to the current solution where I am trying to bring two pieces close together manually.
[tombstone.zip](https://github.com/marcomusy/vedo/files/7847455/tombstone.zip) | closed | 2022-01-11T15:00:02Z | 2022-07-25T12:01:19Z | https://github.com/marcomusy/vedo/issues/576 | [] | ttsesm | 31 |
healthchecks/healthchecks | django | 314 | error during ./manage.py migrate | Hi,
in the end of install process when i run ./manage.py migrate
i get this:
```
(hc-venv) check@healthcheck:~/webapps/healthchecks$ ./manage.py migrate
Traceback (most recent call last):
File "./manage.py", line 10, in <module>
execute_from_command_line(sys.argv)
File "/home/check/webapps/hc-venv/lib/python3.6/site-packages/django/core/management/__init__.py", line 401, in execute_from_command_line
utility.execute()
File "/home/check/webapps/hc-venv/lib/python3.6/site-packages/django/core/management/__init__.py", line 345, in execute
settings.INSTALLED_APPS
File "/home/check/webapps/hc-venv/lib/python3.6/site-packages/django/conf/__init__.py", line 76, in __getattr__
self._setup(name)
File "/home/check/webapps/hc-venv/lib/python3.6/site-packages/django/conf/__init__.py", line 63, in _setup
self._wrapped = Settings(settings_module)
File "/home/check/webapps/hc-venv/lib/python3.6/site-packages/django/conf/__init__.py", line 142, in __init__
mod = importlib.import_module(self.SETTINGS_MODULE)
File "/usr/lib/python3.6/importlib/__init__.py", line 126, in import_module
return _bootstrap._gcd_import(name[level:], package, level)
File "<frozen importlib._bootstrap>", line 994, in _gcd_import
File "<frozen importlib._bootstrap>", line 971, in _find_and_load
File "<frozen importlib._bootstrap>", line 955, in _find_and_load_unlocked
File "<frozen importlib._bootstrap>", line 665, in _load_unlocked
File "<frozen importlib._bootstrap_external>", line 678, in exec_module
File "<frozen importlib._bootstrap>", line 219, in _call_with_frames_removed
File "/home/check/webapps/healthchecks/hc/settings.py", line 39, in <module>
for line in f.readlines():
File "/usr/lib/python3.6/encodings/ascii.py", line 26, in decode
return codecs.ascii_decode(input, self.errors)[0]
UnicodeDecodeError: 'ascii' codec can't decode byte 0xe2 in position 1970: ordinal not in range(128)
```
what can i do?
cc: @ilanmimoun
| closed | 2019-12-18T15:21:05Z | 2019-12-21T19:29:23Z | https://github.com/healthchecks/healthchecks/issues/314 | [] | jonathanparsy | 8 |
Asabeneh/30-Days-Of-Python | numpy | 641 | Muito Bom | Excelente repositório sobre python para quem está começando!!! | open | 2025-01-17T00:16:43Z | 2025-01-17T00:16:43Z | https://github.com/Asabeneh/30-Days-Of-Python/issues/641 | [] | lucasmpeg | 0 |
pyg-team/pytorch_geometric | pytorch | 9,602 | outdated conda build | ### 😵 Describe the installation problem
as shown in https://anaconda.org/pyg/pyg/files, the latest pyg build for conda is 2.5.2 for pytorch 2.2, while the latest releases are 2.5.3 and 2.4, respectively. are there plans for publishing newer conda builds for newer pytorch (and potentially newer cuda)?
### Environment
_No response_ | open | 2024-08-17T15:34:48Z | 2024-08-17T15:34:48Z | https://github.com/pyg-team/pytorch_geometric/issues/9602 | [
"installation"
] | moetayuko | 0 |
jeffknupp/sandman2 | rest-api | 75 | Distribute as docker images | Last year I started using sandman2 at my company for building a quick admin console. Since we use docker to deploy things, and since the project doesn't provide official docker images, I dockerized it myself and published the image on [docker hub](https://hub.docker.com/r/mondora/sandman2-mssql/).
The image (which only targets mssql) has since been downloaded 100k+ times, probably because it's the first result when [searching for **sandman**](https://hub.docker.com/search/?isAutomated=0&isOfficial=0&page=1&pullCount=0&q=sandman&starCount=0) on docker hub.
A couple of days ago I received [a PR on the docker image repo](https://github.com/mondora/docker-sandman2-mssql/pull/1) to update some dependencies, but I'm hesitant to merge it to master because the update could potentially break existing users (back when I created the image I didn't think about setting up a versioning scheme for it, so everything that goes into the master branch of the repo gets published as the `latest` tag of the docker image).
So to get to the point, since there appears to be some demand for a dockerized version of sandman2, you might want to consider directly publishing images for it. | closed | 2018-10-22T06:05:08Z | 2018-10-29T19:53:15Z | https://github.com/jeffknupp/sandman2/issues/75 | [] | pscanf | 3 |
flaskbb/flaskbb | flask | 17 | Searching | I want to use Whoosh for searching, but I still need to look into it how to do it.
| closed | 2014-02-27T13:20:09Z | 2018-04-15T07:47:30Z | https://github.com/flaskbb/flaskbb/issues/17 | [
"enhancement"
] | sh4nks | 2 |
microsoft/unilm | nlp | 921 | [unimim] mismatched positional_embed about vit-large/14 for input resolution with 196 | hello, for CLIP knowledge distilation paper, i.e.,A Unified View of Masked Image Modeling:
when the teacher is CLIP vit-large/14 for 196's input resolution, and the student is vit-base/16 for 224's input resolution, vit-large/14's positional embed (i.e.,257) for CLIP mismatch with the positional embed of our teacher (i.e., 197). How should I fix this to align with the paper.
Thanks very much!
| open | 2022-11-17T06:56:28Z | 2022-11-18T12:51:32Z | https://github.com/microsoft/unilm/issues/921 | [] | futureisatyourhand | 1 |
mirumee/ariadne-codegen | graphql | 178 | Incorrect import with top level fragment with ShorterResultsPlugin | Let's take example schema and query:
```gql
type Query {
hello: TypeA!
}
type TypeA {
valueB: TypeB!
}
type TypeB {
id: ID!
}
```
```gql
query testQuery {
...fragmentHello
}
fragment fragmentHello on Query {
hello {
valueB {
id
}
}
}
```
From these we generate `test_query.py`
```py
class TestQuery(FragmentHello):
pass
```
and `fragments.py`
```py
class FragmentHello(BaseModel):
hello: "FragmentHelloHello"
class FragmentHelloHello(BaseModel):
value_b: "FragmentHelloHelloValueB" = Field(alias="valueB")
```
Without `ShorterResultsPlugin` we generate client which looks like this:
```py
from .test_query import TestQuery
...
async def test_query(self) -> TestQuery:
...
return TestQuery.parse_obj(data)
```
With `ShorterResultsPlugin`:
```py
from .test_query import FragmentHelloHello, TestQuery
...
async def test_query(self) -> FragmentHelloHello:
...
return TestQuery.parse_obj(data).hello
```
Problem with client generated with `ShorterResultsPlugin` is that it imports `FragmentHelloHello` from `test_query.py`, but instead it should be imported from `fragments.py`. | closed | 2023-06-22T14:32:13Z | 2023-07-07T07:37:04Z | https://github.com/mirumee/ariadne-codegen/issues/178 | [
"bug"
] | mat-sop | 1 |
piskvorky/gensim | machine-learning | 2,665 | `train()` doc-comments don't explain `corpus_file` requires both `total_words` and `total_examples` | As using the `corpus_file` option requires **both** `total_words` and `total_examples` to be specified (unlike how the iteratable-corpus needed just one or the other), the doc-comments for `train()` in `Word2Vec`, `Doc2Vec`, & `FastText` are out-of-date about the 'optional' status of these parameters & description of when they're needed. | open | 2019-10-31T22:03:52Z | 2019-11-01T01:10:56Z | https://github.com/piskvorky/gensim/issues/2665 | [
"documentation"
] | gojomo | 0 |
keras-team/keras | deep-learning | 20,210 | Embedding Projector using TensorBoard callback | # Environment
- Python 3.12.4
- Tensorflow v2.16.1-19-g810f233968c 2.16.2
- Keras 3.5.0
- TensorBoard 2.16.2
# How to reproduce it?
I tried to visualizing data using [the embedding Projector in TensorBoard](https://github.com/tensorflow/tensorboard/blob/2.16.2/docs/tensorboard_projector_plugin.ipynb). So I added the following args to TensorBoard callback:
```python
metadata_filename = "metadata.tsv"
os.makedirs(logs_path, exist_ok=True)
# Save Labels separately on a line-by-line manner.
with open(os.path.join(logs_path, metadata_filename), "w") as f:
for token in vectorizer.get_vocabulary():
f.write("{}\n".format(token))
keras.callbacks.TensorBoard(
log_dir=logs_path,
embeddings_freq=1,
embeddings_metadata=metadata_filename
)
```
Anyway TensorBoard embedding tab only shows [this HTML page](https://github.com/tensorflow/tensorboard/blob/4c004d4bddb5040de138815b3bec3cb2829d2878/tensorboard/plugins/projector/vz_projector/vz-projector-dashboard.ts#L23-L65).
# Issues
The above HTML page is returned because [`dataNotFound` is true](https://github.com/tensorflow/tensorboard/blob/4c004d4bddb5040de138815b3bec3cb2829d2878/tensorboard/plugins/projector/vz_projector/vz-projector-dashboard.ts#L22). This happens because [this route](https://github.com/tensorflow/tensorboard/blob/4c004d4bddb5040de138815b3bec3cb2829d2878/tensorboard/plugins/projector/vz_projector/vz-projector-dashboard.ts#L97) (`http://localhost:6006/data/plugin/projector/runs`) returns an [empty JSON](https://github.com/tensorflow/tensorboard/blob/4c004d4bddb5040de138815b3bec3cb2829d2878/tensorboard/plugins/projector/projector_plugin_test.py#L71-L72). In particular, this route is addressed by [this Python function](https://github.com/tensorflow/tensorboard/blob/4c004d4bddb5040de138815b3bec3cb2829d2878/tensorboard/plugins/projector/projector_plugin.py#L545-L549). Under the hood this function tries to [find the latest checkpoint](https://github.com/tensorflow/tensorboard/blob/4c004d4bddb5040de138815b3bec3cb2829d2878/tensorboard/plugins/projector/projector_plugin.py#L458). In particular, it gets the path of the latest checkpoint using [`tf.train.latest_checkpoint`](https://github.com/tensorflow/tensorflow/blob/810f233968cec850915324948bbbc338c97cf57f/tensorflow/python/checkpoint/checkpoint_management.py#L328-L365). Like doc string states, this TF function finds a **TensorFlow (2 or 1.x) checkpoint**. Now, TensorBoard callback [saves a checkpoint](https://github.com/keras-team/keras/blob/fa834a767bfab5d8e4180ada03fd0b7a597d6d55/keras/src/callbacks/tensorboard.py#L591-L596), at the end of the epoch, but it is a **Keras checkpoint**.
Furthermore, `projector_config.pbtxt` is written in the [wrong place](https://github.com/keras-team/keras/blob/fa834a767bfab5d8e4180ada03fd0b7a597d6d55/keras/src/callbacks/tensorboard.py#L304): TensorBoard [expects this file](https://github.com/tensorflow/tensorboard/blob/4c004d4bddb5040de138815b3bec3cb2829d2878/tensorboard/plugins/projector/projector_plugin.py#L441) in the same place where checkpoints are saved.
Finally, choosing [a fixed name](https://github.com/keras-team/keras/blob/fa834a767bfab5d8e4180ada03fd0b7a597d6d55/keras/src/callbacks/tensorboard.py#L278-L283) is a strong assumption. In my model, tensor associated to Embedding layer had a different name (obviously).
## Notes
IMO this feature stopped working when the callback updated to TF 2.0. Indeed, callback for TF 1.x should work. For example, it [saves checkpoint](https://github.com/keras-team/tf-keras/blob/c5f97730b2e495f5f56fc2267d22504075e46337/tf_keras/callbacks_v1.py#L493-L497) using TF format. But when callback was updated to be compatible with TF 2.0 it was used `tf.keras.Model.save_weights` and not `tf.train.Checkpoint`: perfectly legit like reported [here](https://github.com/tensorflow/tensorflow/blob/810f233968cec850915324948bbbc338c97cf57f/tensorflow/python/training/saver.py#L646-L650).
# Possible solution
Saving only weights from Embedding layer. [Here](https://github.com/tensorflow/tensorboard/blob/4c004d4bddb5040de138815b3bec3cb2829d2878/docs/tensorboard_projector_plugin.ipynb#L242-L249), you can find an example. To get model, you can use [`self._model`](https://github.com/keras-team/keras/blob/fa834a767bfab5d8e4180ada03fd0b7a597d6d55/keras/src/callbacks/tensorboard.py#L203). Plus it is not necessary to specify tensor name because there is only one tensor to save. The only drawback is: how to handle two or more embeddings? | open | 2024-09-04T16:58:15Z | 2024-09-19T16:17:55Z | https://github.com/keras-team/keras/issues/20210 | [
"stat:awaiting keras-eng",
"type:Bug"
] | miticollo | 4 |
Miserlou/Zappa | flask | 1,524 | there's a bug to delete lambda versions | self.lambda_client.delete_function(FunctionNmae=function_name,Qualifier=version)
FunctionNmae should be FunctionName。
| open | 2018-06-08T09:47:35Z | 2018-06-10T21:48:16Z | https://github.com/Miserlou/Zappa/issues/1524 | [] | bjmayor | 1 |
matterport/Mask_RCNN | tensorflow | 2,714 | How to plot loss curves in Tensorboard | Can someone guide how to use tensorboard to look at learning curves, I really tried few things available but no graphs coming up.
| open | 2021-10-26T10:06:38Z | 2021-11-27T23:09:03Z | https://github.com/matterport/Mask_RCNN/issues/2714 | [] | chhigansharma | 8 |
mckinsey/vizro | data-visualization | 719 | Apply code formatting to code examples in our docs | Currently our code examples are not formatted using `black` or linted in any way.
* Investigate what mkdocs extensions there are to do this and what they would do (e.g. they might run `ruff` or `black`)
* Find a good solution and apply it! | open | 2024-09-18T17:01:13Z | 2024-12-03T10:42:17Z | https://github.com/mckinsey/vizro/issues/719 | [
"Docs :spiral_notepad:",
"Good first issue :baby_chick:",
"hacktoberfest"
] | antonymilne | 14 |
deezer/spleeter | tensorflow | 616 | [Discussion] Why are separate U-Nets used for each instrument? | Hello! I have a more general question about the model architecture used – Spleeter appears to train a separate U-Net for each instrument track, effectively training separate models for each instrument. What motivated this architecture, as opposed to using a single encoder-decoder that predicts masks for everything all at once? (which is more common in analogous image segmentation models)
I'm exploring source separation for music which doesn't fit the vocals/drums/piano/bass format and it doesn't seem like there's a straightforward way to fine-tune these models for different instruments or more than 5 stems. It also seems to imply that you can train these separators separately (i.e. a piano extractor, a voice extractor, etc.) which is potentially interesting.
Apologies if this has already been discussed elsewhere, I couldn't find anything in the issues/wiki/paper about it.
Thanks! | open | 2021-04-28T00:58:27Z | 2021-04-28T00:58:27Z | https://github.com/deezer/spleeter/issues/616 | [
"question"
] | somewacko | 0 |
rthalley/dnspython | asyncio | 927 | resolve's "Answer" is incorrectly typed (pyright) | **Describe the bug**
Pyright isn't able to get correct types:
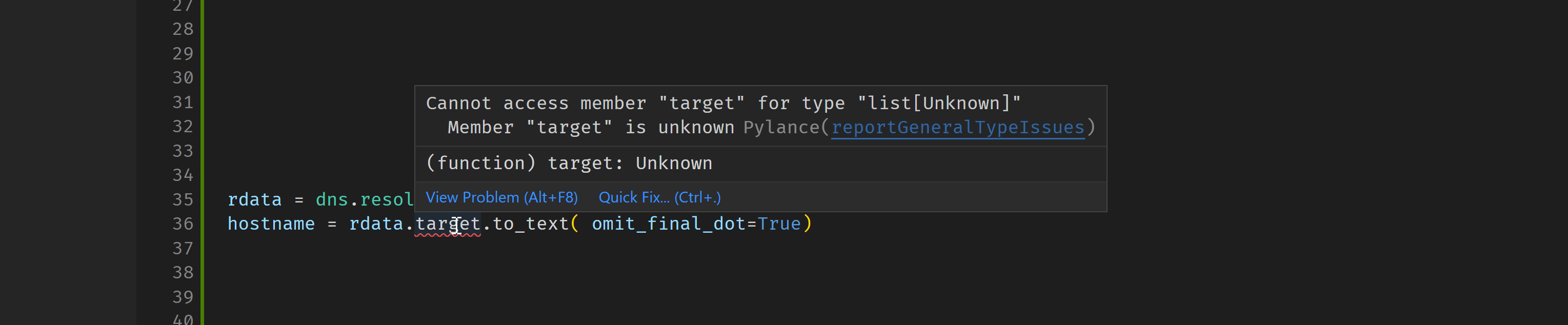
**To Reproduce**
```python
import dns.resolver
rdata = dns.resolver.resolve(cname, "CNAME")[0]
hostname = rdata.target.to_text( omit_final_dot=True)
```
You can use pyright or pylance (which uses pyright).
**Context (please complete the following information):**
- dnspython version: 2.3.0
- Python version: 3.9.16
- OS: Ubuntu
| closed | 2023-04-23T23:04:01Z | 2023-04-30T21:02:55Z | https://github.com/rthalley/dnspython/issues/927 | [] | karolzlot | 2 |
ultralytics/yolov5 | deep-learning | 12,854 | Get Scalar Validation Metrics | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
How can I get the metrics on `val.py ` scalar (numbers) instead of the graphs?
i want something like:
```
mAP@.5: 0.976
mAP@.5:.95: 0.612
Precision: 0.841
Recall: 0.973
```
### Additional
_No response_ | closed | 2024-03-26T18:50:46Z | 2024-10-20T19:42:21Z | https://github.com/ultralytics/yolov5/issues/12854 | [
"question"
] | ArtBreguez | 2 |
slackapi/bolt-python | fastapi | 292 | Is Slack App required to be listed on App directory to be used for sign in with slack | I have developed a web app using Strapi+React. I want a button for Sign in with Slack. Is it necessary to list my Slack app on the App directory to be used for Sign in with Slack integration?
I am getting error `Method not allowed`
#### The `slack_bolt` version
1.1.2
#### Python runtime version
3.7
#### OS info
Microsoft Windows [Version 10.0.19042.906]
| closed | 2021-04-12T11:44:12Z | 2021-04-19T03:43:15Z | https://github.com/slackapi/bolt-python/issues/292 | [
"question"
] | sudhir512kj | 3 |
arogozhnikov/einops | numpy | 67 | Requirements Text | Can we use this library only for numpy operations when we do not have tensorflow/torch/etc?
I was looking for the `requirements.txt` file and it was missing in the Github repo.
It would be helpful for starters if there is info about library requirements. | closed | 2020-09-05T14:10:05Z | 2020-09-09T15:50:48Z | https://github.com/arogozhnikov/einops/issues/67 | [
"question"
] | bhishanpdl | 2 |
hack4impact/flask-base | flask | 160 | Documentation on http://hack4impact.github.io/flask-base outdated. Doesn't match with README | Hi,
It seems that part of the documentation on https://hack4impact.github.io/flask-base/ are outdated.
For example, the **setup section** of the documentation mentions
```
$ pip install -r requirements/common.txt
$ pip install -r requirements/dev.txt
```
But there is no **requirements** folder.
Whereas the setup section in the README mentions
```
pip install -r requirements.txt
```
I find it confusing to have two sources with different information. | closed | 2018-03-20T09:35:42Z | 2018-05-31T17:57:06Z | https://github.com/hack4impact/flask-base/issues/160 | [] | s-razaq | 0 |
sktime/sktime | data-science | 7,775 | [BUG] HierarchicalPdMultiIndex fails to recognize two-level hierarchical indexes |
**Describe the bug**
While implementing a transformer, I encountered an error raised by `BaseTransformer`'s `_convert_output`, which appears to be a bug in `HierarchicalPdMultiIndex._check`. There can be hierarchical indexes with only two levels, for example:
```
value
level_1 time
__total 2020-01-01 100
regionA 2020-01-01 40
regionB 2020-01-01 30
regionC 2020-01-01 30
```
However, the function does not consider this a valid case.
This results in an error with the following message:
*"obj must have a MultiIndex with 3 or more levels, found 2."*
The fix could be implemented in `HierarchicalPdMultiIndex` or by modifying `_check_pdmultiindex_panel`.
Alternatively, this might be a misunderstanding on my part regarding the definition of a hierarchical multi-index.
---
**To Reproduce**
```python
import pandas as pd
from sktime.datatypes._hierarchical._check import HierarchicalPdMultiIndex
# Creating the MultiIndex DataFrame
index = pd.MultiIndex.from_tuples(
[
("__total", "2020-01-01"),
("regionA", "2020-01-01"),
("regionB", "2020-01-01"),
("regionC", "2020-01-01"),
],
names=["level_1", "time"]
)
data = {"value": [100, 40, 30, 30]}
df = pd.DataFrame(data, index=index)
# Should be valid
output = HierarchicalPdMultiIndex()._check(df, return_metadata=True)
is_valid = output[0]
assert is_valid
```
---
**Expected behavior**
The DataFrame should be considered a valid hierarchical scitype.
---
**Versions**
<details>
<summary>System & Dependencies</summary>
System:
- Python: 3.11.11 (main, Dec 26 2024, 12:31:23) [Clang 16.0.0 (clang-1600.0.26.6)]
- Executable: `/Users/felipeangelim/.pyenv/versions/3.11.11/envs/sktime-3.11/bin/python`
- Machine: macOS-15.3-arm64-arm-64bit
Python dependencies:
- `pip`: 24.0
- `sktime`: 0.35.0
- `sklearn`: 1.5.2
- `skbase`: 0.11.0
- `numpy`: 2.1.3
- `scipy`: 1.15.0
- `pandas`: 2.2.3
- `matplotlib`: None
- `joblib`: 1.4.2
- `numba`: None
- `statsmodels`: 0.14.4
- `pmdarima`: 1.8.5
- `statsforecast`: None
- `tsfresh`: None
- `tslearn`: None
- `torch`: None
- `tensorflow`: None
</details> | closed | 2025-02-07T15:12:35Z | 2025-02-07T19:31:08Z | https://github.com/sktime/sktime/issues/7775 | [
"bug",
"module:datatypes"
] | felipeangelimvieira | 3 |
ray-project/ray | data-science | 51,446 | [core] Cover cpplint for `ray/core_worker/transport` | ## Description
As part of the initiative to introduce cpplint into the pre-commit hook, we are gradually cleaning up C++ folders to ensure compliance with code style requirements. This issue focuses on cleaning up /src/ray/core_worker/transport
## Goal
- Ensure all .h and .cc files in `/src/ray/core_worker/transport` comply with cpplint rules.
- Address or suppress all cpplint warnings.
## Steps to Complete
- Checkout the latest main branch and install the pre-commit hook.
- Manually modify all C++ files in `/src/ray/core_worker/transport` to trigger cpplint (e.g., by adding a newline).
- Run git commit to trigger cpplint and identify issues.
- Fix the reported issues or suppress them using clang-tidy if necessary.
This is a sub issue from https://github.com/ray-project/ray/issues/50583 | closed | 2025-03-18T08:26:35Z | 2025-03-19T14:16:08Z | https://github.com/ray-project/ray/issues/51446 | [] | nishi-t | 1 |
scikit-optimize/scikit-optimize | scikit-learn | 226 | Update installation instructions when sklearn 0.18 is released | closed | 2016-09-16T03:56:01Z | 2016-09-29T13:37:39Z | https://github.com/scikit-optimize/scikit-optimize/issues/226 | [
"Easy"
] | MechCoder | 3 | |
SALib/SALib | numpy | 172 | Wrong values in the installaltion test question_interpretation | Hey,
I installed the SALib v 1.1.0 with pip install SALib and tested the exampled as described in the docs.
http://salib.readthedocs.io/en/latest/getting-started.html#testing-installation
According to the docs the Sis should be [ 0.30644324 0.44776661 -0.00104936].
However, I get [ 0.03443819 0.09611386 0.12723021].
I use python 3.5 .2, numpy 1.13.1+mkl, scipy 0.19.1 and matplotlib 1.5.3.
Why do I have these wrong values?
Thanks a lot. | closed | 2017-11-15T09:09:05Z | 2017-11-20T00:20:16Z | https://github.com/SALib/SALib/issues/172 | [] | witteire | 1 |
sqlalchemy/alembic | sqlalchemy | 438 | if/when SQLAlchemy provides truncation for naming convention names, need to do that same truncation on the name comparison side | **Migrated issue, originally created by Danny Milosavljevic**
Hi,
postgresql automatically truncates too-long index names (for the limit see "SELECT max_identifier_length - 1 FROM pg_control_init()") but alembic does not truncate index names in this manner.
That means that if an index name is too long then alembic will always generate a spurious migration where it tries to create the index with the long name and drop the index with the short name.
The bug is not that bad because for cases where the sqlalchemy naming convention generates index names that are too long you can just override it in the model by specifying a non-autogenerated index name ("name=...").
But in the long run it would be nice if alembic would also auto-truncate index names like postgres does.
It is apparently not possible to disable autotruncation in postgresql 9.6.1, so it might be a bit difficult to find these cases.
| open | 2017-07-20T12:57:36Z | 2020-02-12T15:08:25Z | https://github.com/sqlalchemy/alembic/issues/438 | [
"bug",
"autogenerate - detection",
"low priority",
"naming convention issues"
] | sqlalchemy-bot | 5 |
microsoft/UFO | automation | 9 | Error making API request: ('Connection aborted.', RemoteDisconnected('Remote end closed connection without response')) | I followed the Getting Started steps to configure the OpenAI endpoint, but encountered an error during execution.
Error making API request: ('Connection aborted.', RemoteDisconnected('Remote end closed connection without response'))
In the config.yml, I ONLY modified the following parameters:
OPENAI_API_BASE: "https://api.openai.com/v1/chat/completions"
OPENAI_API_KEY: "###"
Could anybody tell me why and how to solve it ? | open | 2024-02-20T13:52:48Z | 2024-06-13T23:18:24Z | https://github.com/microsoft/UFO/issues/9 | [] | xdzha133733 | 5 |
ansible/ansible | python | 83,954 | Trying to create a postgresqlflexibleserver fail with an API internal server error | ### Summary
When trying to create a postgresqlflexibleserver with ansible, I end up with a fatal server error and absolutely no output to understand what's happening.
It works fine with postgresqlserver though.
### Issue Type
Bug Report
### Component Name
postgresqlflexibleserver
### Ansible Version
```console
$ ansible --version
ansible [core 2.17.4]
config file = None
configured module search path = ['/home/michel/.ansible/plugins/modules', '/usr/share/ansible/plugins/modules']
ansible python module location = /home/michel/ws/projects/mypath-sa/venv/lib/python3.12/site-packages/ansible
ansible collection location = /home/michel/.ansible/collections:/usr/share/ansible/collections
executable location = /home/michel/ws/projects/mypath-sa/venv/bin/ansible
python version = 3.12.3 (main, Sep 11 2024, 14:17:37) [GCC 13.2.0] (/home/michel/ws/projects/mypath-sa/venv/bin/python)
jinja version = 3.1.4
libyaml = True
```
### Configuration
```console
# if using a version older than ansible-core 2.12 you should omit the '-t all'
$ ansible-config dump --only-changed -t all
CONFIG_FILE() = None
EDITOR(env: EDITOR) = vim
```
### OS / Environment
linux mint (ubuntu based)
### Steps to Reproduce
<!--- Paste example playbooks or commands between quotes below -->
```yaml (paste below)
- name: make db creation fail
azure.azcollection.azure_rm_postgresqlflexibleserver:
name: dbserver
version: 16
administrator_login: admin_login
administrator_login_password: ###########
resource_group: myresourcegroup
sku:
name: Standard_B1ms
tier: Burstable
```
### Expected Results
Database creation, or at least an error saying what's going on.
### Actual Results
```console
The full traceback is:
File "/tmp/ansible_azure.azcollection.azure_rm_postgresqlflexibleserver_payload_2r8g4x2b/ansible_azure.azcollection.azure_rm_postgresqlflexibleserver_payload.zip/ansible_collections/azure/azcollection/plugins/modules/azure_rm_postgresqlflexibleserver.py", line 862, in create_postgresqlflexibleserver
response = self.get_poller_result(response)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/tmp/ansible_azure.azcollection.azure_rm_postgresqlflexibleserver_payload_2r8g4x2b/ansible_azure.azcollection.azure_rm_postgresqlflexibleserver_payload.zip/ansible_collections/azure/azcollection/plugins/module_utils/azure_rm_common.py", line 635, in get_poller_result
poller.wait(timeout=delay)
File "/home/michel/ws/projects/mypath-sa/venv/lib/python3.12/site-packages/azure/core/tracing/decorator.py", line 76, in wrapper_use_tracer
return func(*args, **kwargs)
^^^^^^^^^^^^^^^^^^^^^
File "/home/michel/ws/projects/mypath-sa/venv/lib/python3.12/site-packages/azure/core/polling/_poller.py", line 261, in wait
raise self._exception # type: ignore
^^^^^^^^^^^^^^^^^^^^^
File "/home/michel/ws/projects/mypath-sa/venv/lib/python3.12/site-packages/azure/core/polling/_poller.py", line 176, in _start
self._polling_method.run()
File "/home/michel/ws/projects/mypath-sa/venv/lib/python3.12/site-packages/azure/core/polling/base_polling.py", line 745, in run
raise HttpResponseError(response=self._pipeline_response.http_response, error=err) from err
fatal: [localhost]: FAILED! => {
"changed": false,
"invocation": {
"module_args": {
"ad_user": null,
"adfs_authority_url": null,
"administrator_login": "admin_login",
"administrator_login_password": "VALUE_SPECIFIED_IN_NO_LOG_PARAMETER",
"api_profile": "latest",
"append_tags": true,
"auth_source": "auto",
"availability_zone": null,
"backup": null,
"cert_validation_mode": null,
"client_id": null,
"cloud_environment": "AzureCloud",
"create_mode": null,
"disable_instance_discovery": false,
"fully_qualified_domain_name": null,
"high_availability": null,
"identity": null,
"is_restart": false,
"is_start": false,
"is_stop": false,
"location": null,
"log_mode": null,
"log_path": null,
"maintenance_window": null,
"name": "test",
"network": null,
"password": null,
"point_in_time_utc": null,
"profile": null,
"resource_group": "myresourcegroup",
"secret": null,
"sku": {
"name": "Standard_B2ms",
"tier": "Burstable"
},
"source_server_resource_id": null,
"state": "present",
"storage": null,
"subscription_id": null,
"tags": null,
"tenant": null,
"thumbprint": null,
"version": "16",
"x509_certificate_path": null
}
},
"msg": "Error creating the PostgreSQL Flexible Server instance: (InternalServerError) An unexpected error occured while processing the request. Tracking ID: '00d0955c-e4e1-48e3-a57b-e5868f78523d'\nCode: InternalServerError\nMessage: An unexpected error occured while processing the request. Tracking ID: '00d0955c-e4e1-48e3-a57b-e5868f78523d'"
}
```
### Code of Conduct
- [X] I agree to follow the Ansible Code of Conduct | closed | 2024-09-17T18:05:17Z | 2024-10-03T13:00:09Z | https://github.com/ansible/ansible/issues/83954 | [
"bug",
"bot_closed",
"affects_2.17"
] | mbegoc | 3 |
mars-project/mars | numpy | 3,267 | [BUG] Ray executor run inv_mapper raises ValueError: assignment destination is read-only | <!--
Thank you for your contribution!
Please review https://github.com/mars-project/mars/blob/master/CONTRIBUTING.rst before opening an issue.
-->
**Describe the bug**
A clear and concise description of what the bug is.
```python
__________________________ test_label_encoder[int64] ___________________________
setup = <mars.deploy.oscar.session.SyncSession object at 0x337a3c7f0>
values = array([2, 1, 3, 1, 3]), classes = array([1, 2, 3])
unknown = array([4])
@pytest.mark.parametrize(
"values, classes, unknown",
[
(
np.array([2, 1, 3, 1, 3], dtype="int64"),
np.array([1, 2, 3], dtype="int64"),
np.array([4], dtype="int64"),
),
(
np.array(["b", "a", "c", "a", "c"], dtype=object),
np.array(["a", "b", "c"], dtype=object),
np.array(["d"], dtype=object),
),
(
np.array(["b", "a", "c", "a", "c"]),
np.array(["a", "b", "c"]),
np.array(["d"]),
),
],
ids=["int64", "object", "str"],
)
def test_label_encoder(setup, values, classes, unknown):
# Test LabelEncoder's transform, fit_transform and
# inverse_transform methods
values_t = mt.tensor(values)
le = LabelEncoder()
le.fit(values_t)
assert_array_equal(le.classes_.fetch(), classes)
assert_array_equal(le.transform(values_t).fetch(), [1, 0, 2, 0, 2])
assert_array_equal(le.inverse_transform(mt.tensor([1, 0, 2, 0, 2])).fetch(), values)
le = LabelEncoder()
> ret = le.fit_transform(values)
mars/learn/preprocessing/tests/test_label.py:300:
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
mars/learn/preprocessing/_label.py:122: in fit_transform
self.classes_, y = execute_tileable(
mars/deploy/oscar/session.py:1888: in execute
return session.execute(
mars/deploy/oscar/session.py:1682: in execute
execution_info: ExecutionInfo = fut.result(
../../.pyenv/versions/3.8.13/lib/python3.8/concurrent/futures/_base.py:444: in result
return self.__get_result()
../../.pyenv/versions/3.8.13/lib/python3.8/concurrent/futures/_base.py:389: in __get_result
raise self._exception
mars/deploy/oscar/session.py:1868: in _execute
await execution_info
../../.pyenv/versions/3.8.13/lib/python3.8/asyncio/tasks.py:695: in _wrap_awaitable
return (yield from awaitable.__await__())
mars/deploy/oscar/session.py:105: in wait
return await self._aio_task
mars/deploy/oscar/session.py:953: in _run_in_background
raise task_result.error.with_traceback(task_result.traceback)
mars/services/task/supervisor/processor.py:372: in run
await self._process_stage_chunk_graph(*stage_args)
mars/services/task/supervisor/processor.py:250: in _process_stage_chunk_graph
chunk_to_result = await self._executor.execute_subtask_graph(
mars/services/task/execution/ray/executor.py:551: in execute_subtask_graph
meta_list = await asyncio.gather(*output_meta_object_refs)
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
awaitable = ObjectRef(b2d1bf24c5f98f84ffffffffffffffffffffffff0100000001000000)
@types.coroutine
def _wrap_awaitable(awaitable):
"""Helper for asyncio.ensure_future().
Wraps awaitable (an object with __await__) into a coroutine
that will later be wrapped in a Task by ensure_future().
"""
> return (yield from awaitable.__await__())
E ray.exceptions.RayTaskError(ValueError): ray::execute_subtask() (pid=97485, ip=127.0.0.1)
E File "/home/admin/mars/mars/services/task/execution/ray/executor.py", line 185, in execute_subtask
E execute(context, chunk.op)
E File "/home/admin/mars/mars/core/operand/core.py", line 491, in execute
E result = executor(results, op)
E File "/home/admin/mars/mars/core/custom_log.py", line 94, in wrap
E return func(cls, ctx, op)
E File "/home/admin/mars/mars/utils.py", line 1160, in wrapped
E return func(cls, ctx, op)
E File "/home/admin/mars/mars/tensor/base/map_chunk.py", line 170, in execute
E ctx[op.outputs[0].key] = op.func(in_data, *args, **kwargs)
E File "/home/admin/mars/mars/learn/utils/_encode.py", line 72, in inv_mapper
E c[c > idx] = idx
E ValueError: assignment destination is read-only
../../.pyenv/versions/3.8.13/lib/python3.8/asyncio/tasks.py:695: RayTaskError(ValueError)
```
**To Reproduce**
To help us reproducing this bug, please provide information below:
1. Your Python version
2. The version of Mars you use
3. Versions of crucial packages, such as numpy, scipy and pandas
4. Full stack of the error.
5. Minimized code to reproduce the error.
**Expected behavior**
A clear and concise description of what you expected to happen.
**Additional context**
Add any other context about the problem here.
| closed | 2022-09-20T08:54:21Z | 2022-10-13T03:43:59Z | https://github.com/mars-project/mars/issues/3267 | [
"type: bug"
] | fyrestone | 0 |
unit8co/darts | data-science | 1,778 | [BUG] TopDownReconciliator modifies Top forecasst | **Describe the bug**
I'm trying to reconcile some hierarchical forecast with Top-Down approach using `TopDownReconciliator`, but the top time series gets also modified. I'm aware there were similar issues in the past, in which something like this happened depending on the order of the time series in the object with `TimeSeries` (https://github.com/unit8co/darts/issues/1582), but in my case the behaviour is order-independent and I'm not using that version of darts.
**To Reproduce**
Here is the code where I define a `TimeSeries` object with three time series and a simple hierarchy, reconcile the forecast with `TopDownReconciliator` and the top series gets modified:
```python
# imports
import pandas as pd
from darts.dataprocessing.transformers import TopDownReconciliator
from darts.timeseries import TimeSeries
# define time series in a pd.DataFrame
df_ts_raw = pd.DataFrame(data={"date": ["2022-02-28",
"2022-03-31",
"2022-04-30",
"2022-05-31",
"2022-06-30"],
"ts_1": [671.450520,
780.584530,
695.618301,
837.410714,
616.029596],
"ts_2": [383.85076871,
412.18267353,
401.97344016,
416.19073302,
488.0563728],
"ts_top": [1228.1294012,
1100.9604,
1045.31923,
1394.89629,
928.9595]})
# transform date column to datetime
df_ts_raw.date = pd.to_datetime(df_ts_raw.date)
# transform pd.DataFrame to TimeSeries
ts_raw = TimeSeries.from_dataframe(df=df_ts_raw,
time_col="date",
value_cols=["ts_1", "ts_2", "ts_top"])
# apply hierarchy
ts_raw = ts_raw.with_hierarchy(hierarchy={"ts_1": "ts_top",
"ts_2": "ts_top"})
# initialise instance of TopDownReconciliator
reconciliator = TopDownReconciliator()
# fit and transform
reconciliator.fit(series=ts_raw)
ts_reconciled = reconciliator.transform(ts_raw)
# print differences of "ts_top" between df_ts_raw and the reconciled time series
print(f'{ts_reconciled.pd_dataframe().reset_index()["ts_top"] - df_ts_raw["ts_top"]}')
```
being the output:
> 0 1.095486
1 0.982052
2 0.932420
3 1.244242
4 0.828628
Name: ts_top, dtype: float64
I modified the order of the columns (e.g. introducing `"ts_top"` before `"ts_1"` and `"ts_2"` in the dictionary to define `df_ts_raw` and modifying accordingly `value_cols` in the definition of `ts_raw` just in case), but the result is exactly the same.
**Expected behavior**
If I understood properly the approach, the time series `"ts_top"` shouldn't get modified after the Top-Down reconciliation, so the output of the previous code should be:
> 0 0.0
1 0.0
2 0.0
3 0.0
4 0.0
Name: ts_top, dtype: **float64**
**System:**
- OS: Windows 10
- Python version: 3.9.13
- darts version: 0.24.0
- pandas version: 1.5.3
| closed | 2023-05-17T08:05:26Z | 2023-08-08T16:05:33Z | https://github.com/unit8co/darts/issues/1778 | [
"wontfix"
] | PL-EduardoSanchez | 3 |
sigmavirus24/github3.py | rest-api | 677 | Docs show a function issues_on for GitHub object, but I am getting attribute error | ```
Traceback (most recent call last):
File "/home/phoenista/Desktop/ghubby/meet/chubby/.env/bin/chubby", line 11, in <module>
load_entry_point('chubby', 'console_scripts', 'chubby')()
File "/home/phoenista/Desktop/ghubby/meet/chubby/chubby/chubby.py", line 112, in main
for iss in gh.issues_on(username=username,
AttributeError: 'GitHub' object has no attribute 'issues_on'
``` | closed | 2017-01-29T15:15:41Z | 2017-01-29T15:35:33Z | https://github.com/sigmavirus24/github3.py/issues/677 | [] | meetmangukiya | 6 |
graphql-python/graphene-django | graphql | 1,373 | Duplicate types when using SerializerMutation with a Model having "choices" | **Note: for support questions, please use stackoverflow**. This repository's issues are reserved for feature requests and bug reports.
* **What is the current behavior?**
When defining a Mutation with the parent SerializerMutation, graphene will try to generate two types with the same name, resulting in the error `AssertionError: Found different types with the same name in the schema: xxx, xxx.`
* **If the current behavior is a bug, please provide the steps to reproduce and if possible a minimal demo of the problem** via
a github repo, https://repl.it or similar.
Define these classes
```
class Demo(models.Model):
class YesNoChoices(models.TextChoices):
YES = "y"
NO = "n"
field_with_choices = models.CharField(max_length=1, choices=YesNoChoices)
class DemoSerializer(serializers.ModelSerializer):
class Meta:
model = Demo
fields = ["field_with_choices"]
class DemoMutation(SerializerMutation):
class Meta:
serializer_class = DemoSerializer
class Mutation(object):
demo_mutation = DemoMutation.Field()
```
then run
```
python manage.py graphql_schema
```
* **What is the expected behavior?**
Types should be created in a clean manner without naming conflicts, resulting in a valid schema.
* **What is the motivation / use case for changing the behavior?**
* **Please tell us about your environment:**
- Version: graphene-django: 2.15.0, python 3.10.6
- Platform: Ubuntu 22.04
* **Other information** (e.g. detailed explanation, stacktraces, related issues, suggestions how to fix, links for us to have context, eg. stackoverflow)
Most likely relates to graphql-python/graphene#1384, where Enums are used. I cannot seem influence how the SerializerMutation handles the choices, however. | open | 2022-11-22T14:53:50Z | 2022-11-23T21:02:52Z | https://github.com/graphql-python/graphene-django/issues/1373 | [
"🐛bug"
] | ramonwenger | 2 |
ExpDev07/coronavirus-tracker-api | rest-api | 1 | The latest and all route is not working on the API server | The latest and all route is not working on the API server
https://coronavirus-tracker-api.herokuapp.com/latest
https://coronavirus-tracker-api.herokuapp.com/all
Thanks! | closed | 2020-02-11T07:06:39Z | 2020-02-11T08:09:50Z | https://github.com/ExpDev07/coronavirus-tracker-api/issues/1 | [] | altezza04 | 2 |
sloria/TextBlob | nlp | 276 | No module named 'xml.etree' | While importing textblob using `from textblob import TextBlob` I get the following error:
```
ModuleNotFoundError Traceback (most recent call last)
~/Documents/GitHub/python-test/lib/python3.7/site-packages/nltk/internals.py in <module>
23 try:
---> 24 from xml.etree import cElementTree as ElementTree
25 except ImportError:
ModuleNotFoundError: No module named 'xml.etree'
During handling of the above exception, another exception occurred:
ModuleNotFoundError Traceback (most recent call last)
<ipython-input-20-3fa94cbd0c01> in <module>
1 # Import TextBlob module
----> 2 from textblob import TextBlob
~/Documents/GitHub/python-test/lib/python3.7/site-packages/textblob/__init__.py in <module>
1 import os
----> 2 from .blob import TextBlob, Word, Sentence, Blobber, WordList
3
4 __version__ = '0.15.3'
5 __license__ = 'MIT'
~/Documents/GitHub/python-test/lib/python3.7/site-packages/textblob/blob.py in <module>
26 from collections import defaultdict
27
---> 28 import nltk
29
30 from textblob.decorators import cached_property, requires_nltk_corpus
~/Documents/GitHub/python-test/lib/python3.7/site-packages/nltk/__init__.py in <module>
97 ]
98
---> 99 from nltk.internals import config_java
100
101 # support numpy from pypy
~/Documents/GitHub/python-test/lib/python3.7/site-packages/nltk/internals.py in <module>
24 from xml.etree import cElementTree as ElementTree
25 except ImportError:
---> 26 from xml.etree import ElementTree
27
28 from six import string_types
ModuleNotFoundError: No module named 'xml.etree'
```
I am trying to import it in a virtualenv. Thanks | open | 2019-07-11T07:40:31Z | 2019-07-11T07:40:31Z | https://github.com/sloria/TextBlob/issues/276 | [] | rmrbytes | 0 |
labmlai/annotated_deep_learning_paper_implementations | machine-learning | 78 | Can you open the webside? | closed | 2021-08-12T02:12:07Z | 2021-08-14T11:45:45Z | https://github.com/labmlai/annotated_deep_learning_paper_implementations/issues/78 | [
"question"
] | JLUForever | 2 | |
lux-org/lux | jupyter | 34 | Add Pivot action to support identity case | ```
df.set_context([lux.Spec(attribute = "Horsepower"),lux.Spec(attribute = "Horsepower")])
df
```
Right now, we penalize views that have duplicate attributes with an interestingness score of -1, which is why we don't have Enhance and Filter here. This would actually be one of the few places where `Pivot` might be helpful to help users to "get unstuck".

| closed | 2020-07-17T05:39:13Z | 2021-01-11T12:38:26Z | https://github.com/lux-org/lux/issues/34 | [] | dorisjlee | 0 |
PaddlePaddle/models | nlp | 4,721 | No such file or directory: './data/vangogh2photo/trainA.txt' | 这个trainA.txt文件该去哪里找呢?下载过来的数据集只有4个图像文件
| closed | 2020-06-28T06:39:12Z | 2020-06-28T07:28:50Z | https://github.com/PaddlePaddle/models/issues/4721 | [] | shaunhurryup | 1 |
horovod/horovod | deep-learning | 4,023 | Horovod + Deepspeed : Device mismatch error | **Environment:**
Machine Info : 8xA100 (80G)
1. Framework: (TensorFlow, Keras, PyTorch, MXNet) : Pytorch
2. Framework version: 1.12.1+cu113
3. Horovod version: 0.28.1
4. MPI version: 3.1.5
5. CUDA version:
6. NCCL version:
7. Python version: 3.8.10
8. Spark / PySpark version:
9. Ray version:
10. OS and version: Ubuntu 20.04
11. GCC version:
12. CMake version:
**Checklist:**
1. Did you search issues to find if somebody asked this question before? Yes
2. If your question is about hang, did you read [this doc](https://github.com/horovod/horovod/blob/master/docs/running.rst)?
3. If your question is about docker, did you read [this doc](https://github.com/horovod/horovod/blob/master/docs/docker.rst)?
4. Did you check if you question is answered in the [troubleshooting guide](https://github.com/horovod/horovod/blob/master/docs/troubleshooting.rst)?
**Bug report:**
Please describe erroneous behavior you're observing and steps to reproduce it.
```
[1,1]<stderr>:Traceback (most recent call last):
[1,1]<stderr>: File "sc2.py", line 178, in <module>
[1,1]<stderr>: outputs = model(input_ids=d['input_ids'],attention_mask=d['attention_mask'])
[1,1]<stderr>: File "/usr/local/lib/python3.8/dist-packages/torch/nn/modules/module.py", line 1130, in _call_impl
[1,1]<stderr>: return forward_call(*input, **kwargs)
[1,1]<stderr>: File "/usr/local/lib/python3.8/dist-packages/deepspeed/utils/nvtx.py", line 15, in wrapped_fn
[1,1]<stderr>: ret_val = func(*args, **kwargs)
[1,1]<stderr>: File "/usr/local/lib/python3.8/dist-packages/deepspeed/runtime/engine.py", line 1842, in forward
[1,1]<stderr>: loss = self.module(*inputs, **kwargs)
[1,1]<stderr>: File "/usr/local/lib/python3.8/dist-packages/torch/nn/modules/module.py", line 1148, in _call_impl
[1,1]<stderr>: result = forward_call(*input, **kwargs)
[1,1]<stderr>: File "/usr/local/lib/python3.8/dist-packages/transformers/models/llama/modeling_llama.py", line 1183, in forward
[1,1]<stderr>: outputs = self.model(
[1,1]<stderr>: File "/usr/local/lib/python3.8/dist-packages/torch/nn/modules/module.py", line 1148, in _call_impl
[1,1]<stderr>: result = forward_call(*input, **kwargs)
[1,1]<stderr>: File "/usr/local/lib/python3.8/dist-packages/transformers/models/llama/modeling_llama.py", line 1027, in forward
[1,1]<stderr>: inputs_embeds = self.embed_tokens(input_ids)
[1,1]<stderr>: File "/usr/local/lib/python3.8/dist-packages/torch/nn/modules/module.py", line 1148, in _call_impl
[1,1]<stderr>: result = forward_call(*input, **kwargs)
[1,1]<stderr>: File "/usr/local/lib/python3.8/dist-packages/torch/nn/modules/sparse.py", line 158, in forward
[1,1]<stderr>: return F.embedding(
[1,1]<stderr>: File "/usr/local/lib/python3.8/dist-packages/torch/nn/functional.py", line 2199, in embedding
[1,1]<stderr>: return torch.embedding(weight, input, padding_idx, scale_grad_by_freq, sparse)
[1,1]<stderr>:RuntimeError: Expected all tensors to be on the same device, but found at least two devices, cuda:1 and cpu! (when checking argument for argument index in method wrapper__index_select)
```
Environment setup
```
Docker : horovod/horovod:latest
pip install datasets evaluate accelerate==0.25.0 transformers==4.37.0 deepspeed==0.13.1
pip install git+https://github.com/aicrumb/datasettokenizer -q
```
[Script](https://drive.google.com/file/d/1KYcMZ4Rg0oyg6pgNd6ZzR_PgZDINlPq_/view?usp=drive_link)
I think I am not sure if script is correct or not. I am still under process of making it work.
Let me know if need any additional information. | closed | 2024-02-15T04:19:26Z | 2024-02-16T01:33:07Z | https://github.com/horovod/horovod/issues/4023 | [
"bug"
] | PurvangL | 0 |
GibbsConsulting/django-plotly-dash | plotly | 414 | target param for links no longer working | html.A('google', href='google.com', target="_blank")
Works as intended in version 1.6.4
Breaks in any updated versions.
target="_self" does work | open | 2022-08-02T21:14:53Z | 2022-09-07T12:26:56Z | https://github.com/GibbsConsulting/django-plotly-dash/issues/414 | [
"question"
] | amd-pscannell | 1 |
cvat-ai/cvat | computer-vision | 8,802 | Where are the annotation txt files corresponding to the images in the project or task? | ### Actions before raising this issue
- [X] I searched the existing issues and did not find anything similar.
- [X] I read/searched [the docs](https://docs.cvat.ai/docs/)
### Is your feature request related to a problem? Please describe.
I want to be able to directly find the corresponding txt file for each image after annotating on the CVAT platform, so that I don't have to use the export dataset feature in tasks.thanks
### Describe the solution you'd like
_No response_
### Describe alternatives you've considered
_No response_
### Additional context
_No response_ | closed | 2024-12-09T13:41:44Z | 2024-12-11T07:01:25Z | https://github.com/cvat-ai/cvat/issues/8802 | [
"enhancement"
] | stephen-TT | 5 |
deepset-ai/haystack | nlp | 8,824 | Reliably check whether a component has been warmed up or not | In the current Pipeline, whenever the `Pipeline.run()` is called the `warm_up()` for every component is run. We want to avoid that an expensive operation is executed multiple times, we cannot to this from the pipeline side. We should review that every component which has a `warm_up()` makes this check.
For instance, `SentenceTransformersTextEmbedder` is [doing it properly](https://github.com/deepset-ai/haystack/blob/main/haystack/components/embedders/sentence_transformers_text_embedder.py#L178) by checking if the sentence transformers model was already initialized.
The `NamedEntityExtractor` [uses a boolean](https://github.com/deepset-ai/haystack/blob/main/haystack/components/extractors/named_entity_extractor.py#L142) to keep track of this state.
We should review all the `warm_up()` methods and make sure this is the current behaviour.
| closed | 2025-02-06T11:37:13Z | 2025-02-06T11:41:31Z | https://github.com/deepset-ai/haystack/issues/8824 | [] | davidsbatista | 1 |
ivy-llc/ivy | numpy | 28,348 | Fix Frontend Failing Test: numpy - math.tensorflow.math.argmin | To-do List: https://github.com/unifyai/ivy/issues/27497 | closed | 2024-02-20T11:40:04Z | 2024-02-20T15:36:44Z | https://github.com/ivy-llc/ivy/issues/28348 | [
"Sub Task"
] | Sai-Suraj-27 | 0 |
jupyter-incubator/sparkmagic | jupyter | 658 | [BUG] Inconsistent behavior in "spark add" | **Describe the bug**
```
%spark add -u LIVY_HOST -s "new_session" -l "python"
```
results in
```
An error was encountered:
Cannot get session kind for "python".
```
However if I do:
```
from sparkmagic.utils.configuration import get_livy_kind
get_livy_kind("python")
```
it returns
```pyspark```
**Expected behavior**
Should not return "Cannot get session kind for "python"."
**Versions:**
- SparkMagic: 0.15.0
| closed | 2020-07-10T18:22:19Z | 2020-07-10T19:00:29Z | https://github.com/jupyter-incubator/sparkmagic/issues/658 | [] | kyprifog | 1 |
joerick/pyinstrument | django | 288 | nevergrad import fails when profiler is active | To reproduce:
```
from pyinstrument import Profiler
profiler = Profiler()
profiler.start()
import nevergrad as ng
profiler.stop()
profiler.print()
```
This is under python 3.11, nevergrad 0.13.0, and pyinstrument 4.6.1
Traceback:
```
---------------------------------------------------------------------------
RuntimeError Traceback (most recent call last)
Cell In[1], line 6
3 profiler = Profiler()
4 profiler.start()
----> 6 import nevergrad as ng
8 profiler.stop()
10 profiler.print()
File ~/micromamba/envs/dev/lib/python3.11/site-packages/nevergrad/__init__.py:8
6 from .common import typing as typing
7 from .parametrization import parameter as p
----> 8 from .optimization import optimizerlib as optimizers # busy namespace, likely to be simplified
9 from .optimization import families as families
10 from .optimization import callbacks as callbacks
File ~/micromamba/envs/dev/lib/python3.11/site-packages/nevergrad/optimization/__init__.py:7
1 # Copyright (c) Meta Platforms, Inc. and affiliates.
2 #
3 # This source code is licensed under the MIT license found in the
4 # LICENSE file in the root directory of this source tree.
6 from .base import Optimizer # abstract class, for type checking
----> 7 from . import optimizerlib
8 from .optimizerlib import registry as registry
File ~/micromamba/envs/dev/lib/python3.11/site-packages/nevergrad/optimization/optimizerlib.py:26
24 from nevergrad.parametrization import _layering
25 from nevergrad.parametrization import _datalayers
---> 26 from . import oneshot
27 from . import base
28 from . import mutations
File ~/micromamba/envs/dev/lib/python3.11/site-packages/nevergrad/optimization/oneshot.py:461
455 ScrHammersleySearch = SamplingSearch(sampler="Hammersley", scrambled=True).set_name(
456 "ScrHammersleySearch", register=True
457 )
458 QOScrHammersleySearch = SamplingSearch(
459 sampler="Hammersley", scrambled=True, opposition_mode="quasi"
460 ).set_name("QOScrHammersleySearch", register=True)
--> 461 OScrHammersleySearch = SamplingSearch(
462 sampler="Hammersley", scrambled=True, opposition_mode="opposite"
463 ).set_name("OScrHammersleySearch", register=True)
464 CauchyScrHammersleySearch = SamplingSearch(cauchy=True, sampler="Hammersley", scrambled=True).set_name(
465 "CauchyScrHammersleySearch", register=True
466 )
467 LHSSearch = SamplingSearch(sampler="LHS").set_name("LHSSearch", register=True)
File ~/micromamba/envs/dev/lib/python3.11/site-packages/nevergrad/optimization/oneshot.py:407, in SamplingSearch.__init__(self, sampler, scrambled, middle_point, opposition_mode, cauchy, autorescale, scale, rescaled, recommendation_rule)
394 def __init__(
395 self,
396 *,
(...)
405 recommendation_rule: str = "pessimistic",
406 ) -> None:
--> 407 super().__init__(_SamplingSearch, locals())
File ~/micromamba/envs/dev/lib/python3.11/site-packages/nevergrad/optimization/base.py:776, in ConfiguredOptimizer.__init__(self, OptimizerClass, config, as_config)
774 self._as_config = as_config
775 self._config = config # keep all, to avoid weird behavior at mismatch between optim and configoptim
--> 776 diff = ngtools.different_from_defaults(instance=self, instance_dict=config, check_mismatches=True)
777 params = ", ".join(f"{x}={y!r}" for x, y in sorted(diff.items()))
778 self.name = f"{self.__class__.__name__}({params})"
File ~/micromamba/envs/dev/lib/python3.11/site-packages/nevergrad/common/tools.py:185, in different_from_defaults(instance, instance_dict, check_mismatches)
183 miss = set(instance_dict.keys()) - set(defaults.keys())
184 if add or miss: # this is to help during development
--> 185 raise RuntimeError(
186 f"Mismatch between attributes and arguments of {instance.__class__}:\n"
187 f"- additional: {add}\n- missing: {miss}"
188 )
189 else:
190 defaults = {x: y for x, y in defaults.items() if x in instance.__dict__}
RuntimeError: Mismatch between attributes and arguments of <class 'nevergrad.optimization.oneshot.SamplingSearch'>:
- additional: set()
- missing: {'__class__', 'self'}
``` | open | 2024-01-18T18:58:57Z | 2024-08-26T13:49:05Z | https://github.com/joerick/pyinstrument/issues/288 | [] | stephanos-stephani | 4 |
streamlit/streamlit | data-science | 10,351 | Data Editor New Row Added to Bottom is a Usability Issue | ### Checklist
- [x] I have searched the [existing issues](https://github.com/streamlit/streamlit/issues) for similar issues.
- [x] I added a very descriptive title to this issue.
- [x] I have provided sufficient information below to help reproduce this issue.
### Summary
When using the data editor component, in both Snowflake and Open Source, and adding new rows, they are appended to the bottom of the data frame, not the top. If the data set is greater than 5-6 rows. This causes usability issues because 1. the user can't tell if a row has been added, and 2. requires a tremendous amount of scrolling for larger data sets.
Is it possible to insert new rows at the top?
### Reproducible Code Example
```Python
import streamlit as st
import pandas as pd
import numpy as np
np.random.seed(42)
num_rows = 100
data = {
"ID": np.arange(1, num_rows + 1),
"Name": [f"User_{i}" for i in range(1, num_rows + 1)],
"Age": np.random.randint(18, 65, size=num_rows),
"Score": np.round(np.random.uniform(50, 100, size=num_rows), 2)
}
df = pd.DataFrame(data)
df_edited = st.data_editor(df, num_rows="dynamic")
```
### Steps To Reproduce
1. Add rows
2. Must scroll to bottom
### Expected Behavior
New rows added to top.
### Current Behavior
_No response_
### Is this a regression?
- [ ] Yes, this used to work in a previous version.
### Debug info
- Streamlit version: 1.39
- Python version: 3.11
- Operating System: Snowflake SaaS
- Browser: Prisma / Chrome / All
### Additional Information
_No response_ | closed | 2025-02-06T01:53:21Z | 2025-02-14T22:32:57Z | https://github.com/streamlit/streamlit/issues/10351 | [
"type:enhancement",
"feature:st.dataframe",
"feature:st.data_editor"
] | sfc-gh-acarson | 3 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.