code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
<h1><center>Report 12</center></h1>
<h3><center>Jiachen Tian</center><h3>
# Introduction
This week's task is primarily glint detection. Right now there are three potential ways: KCF, Hough Transform, and Ratio Comparison. Even though Ratio Comparison works well under certain conditions within a certain displacement of the glitn as illustrated in the last report, the noise sensitive nature guarantees its failure under uncertain inputs(thousands of different frams with different noise patterns). KCF tracker works a little better comparing to ratio comparison but it tends to fail with too many inputs. In comparion, Hough transform might still work the best with proper parameter settings. All the above algorithm will be illustrated.
# Setup
Setup path to include our files. import them. use `autoreload` to get changes in as they are made.
```
import os
import sys
import cv2
from matplotlib import pyplot as plt
import statistics
import numpy as np
# load our code
sys.path.insert(0, os.path.abspath('../'))
from plotting import auto_draw
# specific to jupyter notebook
from jupyter_help import cvplt, cvplt_sub
#Import image processing function from optimization
# load any changes as we make them
%load_ext autoreload
%autoreload 2
```
# Ratio comparison
The previous report illustrates how to map the rectangle back to the original place without taking into account two crucial parameters: ratio differences between each glint caused by noise(present even with noise filtering), and variation in terms of displacements. To diminish errors caused by ratio difference, one could find the stsndard deviation between ratio for each part(Top left, top right, bottom left, bottom right) and get the smallest std. However, using standard deviation would leave the program more vulnerable to errors caused by displacement changes.
```
#Read in the original image
image = cv2.imread("../input/chosen_pic.png")
keep1 = image.copy()
keep2 = image.copy()
#Run auto threshold on the original image
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
#Don't blur it
blurred = cv2.blur(gray,(1, 1))
_,proc = cv2.threshold(blurred,153,153,cv2.THRESH_BINARY)
#Draw the rendered program
cvplt(proc)
#Set the color
color = (255, 0, 0)
#When calculating standard deviation, the individual ratio is based upon number of non-zero pixels.
sample1 = np.array(proc[158:170, 125:137]) #User chosen coordinates
show1 = cv2.rectangle(keep1, (125, 158), (137, 170), color, 2)
number1 = np.count_nonzero(sample1)
cvplt(show1)
print(number1)
#Move it right 10 unit(Standard displacement I set on the previous report)
sample2 = np.array(proc[158:170, 135:147])
show2 = cv2.rectangle(keep2, (135, 158), (147, 170), color, 2)
number2 = np.count_nonzero(sample2)
cvplt(show2)
print(number2)
#Even when it moves to the border, we could still get roughly 27 false pixel that interferes the results.
```
# Hogh Transform
Previously thought that this method is infeasible on glint due to the small size of glint. However, with the help of OTSU's method and results from the pupil detection, it might just work.
```
#Read in the original image
image = cv2.imread("../input/chosen_pic.png")
result = image.copy()
#BGR to grey to eliminate extra layers
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
#First parameter: blur(set to 10)
blur = (10, 10)
image = cv2.blur(image,blur)
#second parameter: canny(set to 40 to 50)
#Any reasonable parameter would suffice. OTSU would handle the neutralization.
#Third parameter: threshold(determined by OTSU)
#We want the threshold to be exclusively on glint. So first crop the frame
cropped = image[158:170, 125:137]
#Run OTSU
thre,proc = cv2.threshold(cropped,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
#Threshold we need
print(thre)
#Affect the threshold on the big image
_, image =cv2.threshold(image,thre,thre,cv2.THRESH_BINARY)
#Canny the image
edges = cv2.Canny(image, 40, 50)
cvplt(edges)
#If we just run the hough transform like this
circles = cv2.HoughCircles(edges, cv2.HOUGH_GRADIENT, 1,150,\
param1 = 200, param2 = 20,\
minRadius = 0, maxRadius = 0)
circles = circles[0][0]
#Draw the circle
circle = cv2.circle(result, (circles[0], circles[2]), circles[2], color, 2)
#Obviously, the algorithm thinks the circle is on the top
cvplt(circle)
#Way out: search in the cropped area cropped out by user in the first place
small_edges = edges[157:171, 125:139]
cvplt(small_edges)
circles = cv2.HoughCircles(small_edges, cv2.HOUGH_GRADIENT, 1,150,\
param1 = 200, param2 = 10,\
minRadius = 0, maxRadius = 0)
color_true = (0, 255, 0)
circles = circles[0][0]
print(circles)
#Map the circle back to the big picture
y = 157 + int(circles[1])
x = 125 + int(circles[0])
#Plot the fixed circle
circle = cv2.circle(result, (x, y), circles[2], color_true, 2)
cvplt(circle)
#As shown in the green circle, it correctly maps to the correct position.
```
# Optimization
What if the glint moves beyong the cell?
- Make the original cell bigger
- Update the cell position based on pupil
# Analysis
As shown above, for glit detection, I will use hough transform as the main algorithm and KCF as well as ratio differences as complementary analysis to get a more precise result.
# Conclusion
Both Pupil tracking and glint tracking is about to finish. The next step would be to further improve precision and find the angle of staring based upon values from both glint and pupil.
| github_jupyter |
## Seminar and homework (10 points total)
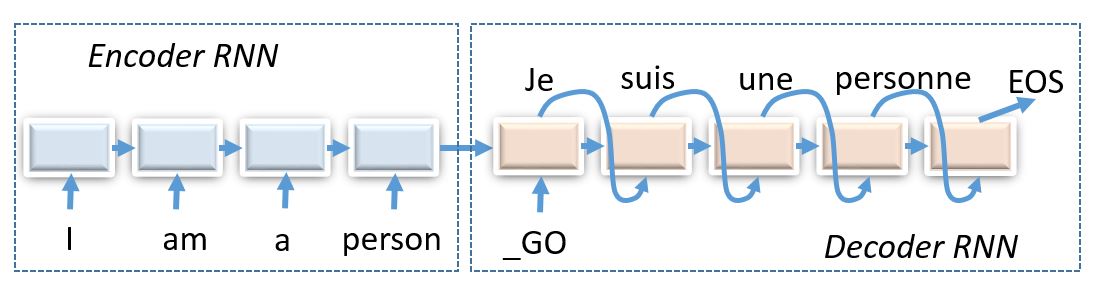
Today we shall compose encoder-decoder neural networks and apply them to the task of machine translation.
_(img: esciencegroup.files.wordpress.com)_
Encoder-decoder architectures are about converting anything to anything, including
* Machine translation and spoken dialogue systems
* [Image captioning](http://mscoco.org/dataset/#captions-challenge2015) and [image2latex](https://openai.com/requests-for-research/#im2latex) (convolutional encoder, recurrent decoder)
* Generating [images by captions](https://arxiv.org/abs/1511.02793) (recurrent encoder, convolutional decoder)
* Grapheme2phoneme - convert words to transcripts
## Our task: machine translation
We gonna try our encoder-decoder models on russian to english machine translation problem. More specifically, we'll translate hotel and hostel descriptions. This task shows the scale of machine translation while not requiring you to train your model for weeks if you don't use GPU.
Before we get to the architecture, there's some preprocessing to be done. ~~Go tokenize~~ Alright, this time we've done preprocessing for you. As usual, the data will be tokenized with WordPunctTokenizer.
However, there's one more thing to do. Our data lines contain unique rare words. If we operate on a word level, we will have to deal with large vocabulary size. If instead we use character-level models, it would take lots of iterations to process a sequence. This time we're gonna pick something inbetween.
One popular approach is called [Byte Pair Encoding](https://github.com/rsennrich/subword-nmt) aka __BPE__. The algorithm starts with a character-level tokenization and then iteratively merges most frequent pairs for N iterations. This results in frequent words being merged into a single token and rare words split into syllables or even characters.
```
# Uncomment selected lines if necessary
# !pip3 install tensorflow-gpu==2.0.0
# !pip3 install subword-nmt &> log
# !wget https://www.dropbox.com/s/yy2zqh34dyhv07i/data.txt?dl=1 -O data.txt
# !wget https://www.dropbox.com/s/fj9w01embfxvtw1/dummy_checkpoint.npz?dl=1 -O dummy_checkpoint.npz
# !wget https://raw.githubusercontent.com/yandexdataschool/nlp_course/2019/week04_seq2seq/utils.py -O utils.py
# # thanks to tilda and deephack teams for the data, Dmitry Emelyanenko for the code :)
from nltk.tokenize import WordPunctTokenizer
from subword_nmt.learn_bpe import learn_bpe
from subword_nmt.apply_bpe import BPE
tokenizer = WordPunctTokenizer()
def tokenize(x):
return ' '.join(tokenizer.tokenize(x.lower()))
# split and tokenize the data
with open('train.en', 'w') as f_src, open('train.ru', 'w') as f_dst:
for line in open('data.txt'):
src_line, dst_line = line.strip().split('\t')
f_src.write(tokenize(src_line) + '\n')
f_dst.write(tokenize(dst_line) + '\n')
# build and apply bpe vocs
bpe = {}
for lang in ['en', 'ru']:
learn_bpe(open('./train.' + lang), open('bpe_rules.' + lang, 'w'), num_symbols=8000)
bpe[lang] = BPE(open('./bpe_rules.' + lang))
with open('train.bpe.' + lang, 'w') as f_out:
for line in open('train.' + lang):
f_out.write(bpe[lang].process_line(line.strip()) + '\n')
```
### Building vocabularies
We now need to build vocabularies that map strings to token ids and vice versa. We're gonna need these fellas when we feed training data into model or convert output matrices into words.
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
data_inp = np.array(open('./train.bpe.ru').read().split('\n'))
data_out = np.array(open('./train.bpe.en').read().split('\n'))
from sklearn.model_selection import train_test_split
train_inp, dev_inp, train_out, dev_out = train_test_split(data_inp, data_out, test_size=3000,
random_state=42)
for i in range(3):
print('inp:', train_inp[i])
print('out:', train_out[i], end='\n\n')
from utils import Vocab
inp_voc = Vocab.from_lines(train_inp)
out_voc = Vocab.from_lines(train_out)
# Here's how you cast lines into ids and backwards.
batch_lines = sorted(train_inp, key=len)[5:10]
batch_ids = inp_voc.to_matrix(batch_lines)
batch_lines_restored = [' '.join(x) for x in inp_voc.to_lines(batch_ids)]
print("lines")
print(batch_lines)
print("\nwords to ids (0 = bos, 1 = eos):")
print(batch_ids)
print("\nback to words")
print(batch_lines_restored)
```
Draw source and translation length distributions to estimate the scope of the task.
```
plt.figure(figsize=[8, 4])
plt.subplot(1, 2, 1)
plt.title("source length")
plt.hist(list(map(len, map(str.split, train_inp))), bins=20);
plt.subplot(1, 2, 2)
plt.title("translation length")
plt.hist(list(map(len, map(str.split, train_out))), bins=20);
```
### Encoder-decoder model
The code below contas a template for a simple encoder-decoder model: single GRU encoder/decoder, no attention or anything. This model is implemented for you as a reference and a baseline for your homework assignment.
```
import tensorflow as tf
assert tf.__version__.startswith('2'), "Current tf version: {}; required: 2.0.*".format(tf.__version__)
L = tf.keras.layers
keras = tf.keras
from utils import infer_length, infer_mask
class BasicModel(L.Layer):
def __init__(self, inp_voc, out_voc, emb_size=64, hid_size=128):
"""
A simple encoder-decoder model
"""
super().__init__() # initialize base class to track sub-layers, trainable variables, etc.
self.inp_voc, self.out_voc = inp_voc, out_voc
self.hid_size = hid_size
self.emb_inp = L.Embedding(len(inp_voc), emb_size)
self.emb_out = L.Embedding(len(out_voc), emb_size)
self.enc0 = L.GRUCell(hid_size)
self.dec_start = L.Dense(hid_size)
self.dec0 = L.GRUCell(hid_size)
self.logits = L.Dense(len(out_voc))
def call(self, inp, out):
initial_state = self.encode(inp)
return self.decode(initial_state, out)
def encode(self, inp, **flags):
"""
Takes symbolic input sequence, computes initial state
:param inp: matrix of input tokens [batch, time]
:returns: initial decoder state tensors, one or many
"""
inp_emb = self.emb_inp(inp)
batch_size = inp.shape[0]
mask = infer_mask(inp, self.inp_voc.eos_ix, dtype=tf.bool)
state = [tf.zeros((batch_size, self.hid_size), tf.float32)]
for i in tf.range(inp_emb.shape[1]):
output, next_state = self.enc0(inp_emb[:, i], state)
state = [
tf.where(
tf.tile(mask[:, i, None],[1,next_tensor.shape[1]]),
next_tensor, tensor
) for tensor, next_tensor in zip(state, next_state)
]
dec_start = self.dec_start(state[0])
return [dec_start]
def decode_step(self, prev_state, prev_tokens, **flags):
"""
Takes previous decoder state and tokens, returns new state and logits for next tokens
:param prev_state: a list of previous decoder state tensors
:param prev_tokens: previous output tokens, an int vector of [batch_size]
:return: a list of next decoder state tensors, a tensor of logits [batch, n_tokens]
"""
<YOUR CODE HERE>
return new_dec_state, output_logits
def decode(self, initial_state, out_tokens, **flags):
state = initial_state
batch_size = out_tokens.shape[0]
# initial logits: always predict BOS
first_logits = tf.math.log(
tf.one_hot(tf.fill([batch_size], self.out_voc.bos_ix), len(self.out_voc)) + 1e-30)
outputs = [first_logits]
for i in tf.range(out_tokens.shape[1] - 1):
#on each step update the state and store new logits into output
<YOUR CODE HERE>
return tf.stack(outputs, axis=1)
def decode_inference(self, initial_state, max_len=100, **flags):
state = initial_state
outputs = [tf.ones(initial_state[0].shape[0], tf.int32) * self.out_voc.bos_ix]
all_states = initial_state
for i in tf.range(max_len):
state, logits = self.decode_step(state, outputs[-1])
outputs.append(tf.argmax(logits, axis=-1, output_type=tf.int32))
all_states.append(state[0])
return tf.stack(outputs, axis=1), tf.stack(all_states, axis=1)
def translate_lines(self, inp_lines):
inp = tf.convert_to_tensor(inp_voc.to_matrix(inp_lines))
initial_state = self.encode(inp)
out_ids, states = self.decode_inference(initial_state)
return out_voc.to_lines(out_ids.numpy()), states
model = BasicModel(inp_voc, out_voc)
```
### Training loss (2 points)
Our training objetive is almost the same as it was for neural language models:
$$ L = {\frac1{|D|}} \sum_{X, Y \in D} \sum_{y_t \in Y} - \log p(y_t \mid y_1, \dots, y_{t-1}, X, \theta) $$
where $|D|$ is the __total length of all sequences__, including BOS and first EOS, but excluding PAD.
```
dummy_inp = tf.convert_to_tensor(inp_voc.to_matrix(train_inp[:3]))
dummy_out = tf.convert_to_tensor(out_voc.to_matrix(train_out[:3]))
dummy_logits = model(dummy_inp, dummy_out)
ref_shape = (dummy_out.shape[0], dummy_out.shape[1], len(out_voc))
assert dummy_logits.shape == ref_shape, "Your logits shape should be {} but got {}".format(dummy_logits.shape, ref_shape)
assert all(dummy_logits[:, 0].numpy().argmax(-1) == out_voc.bos_ix), "first step must always be BOS"
def compute_loss(model, inp, out, **flags):
"""
Compute loss (float32 scalar) as in the formula above
:param inp: input tokens matrix, int32[batch, time]
:param out: reference tokens matrix, int32[batch, time]
In order to pass the tests, your function should
* include loss at first EOS but not the subsequent ones
* divide sum of losses by a sum of input lengths (use infer_length or infer_mask)
"""
inp, out = map(tf.convert_to_tensor, [inp, out])
#outputs of the model
logits_seq = <YOUR CODE HERE>
#log-probabilities of all tokens at all steps
logprobs_seq = <YOUR CODE HERE>
#select the correct
logp_out = logprobs_seq * tf.one_hot(out, len(model.out_voc), dtype=tf.float32)
mask = infer_mask(out, out_voc.eos_ix)
#return cross-entropy
return <YOUR CODE HERE>
dummy_loss = compute_loss(model, dummy_inp, dummy_out)
print("Loss:", dummy_loss)
assert np.allclose(dummy_loss, 8.425, rtol=0.1, atol=0.1), "We're sorry for your loss"
```
### Evaluation: BLEU
Machine translation is commonly evaluated with [BLEU](https://en.wikipedia.org/wiki/BLEU) score. This metric simply computes which fraction of predicted n-grams is actually present in the reference translation. It does so for n=1,2,3 and 4 and computes the geometric average with penalty if translation is shorter than reference.
While BLEU [has many drawbacks](http://www.cs.jhu.edu/~ccb/publications/re-evaluating-the-role-of-bleu-in-mt-research.pdf), it still remains the most commonly used metric and one of the simplest to compute.
__Note:__ in this assignment we measure token-level bleu with bpe tokens. Most scientific papers report word-level bleu. You can measure it by undoing BPE encoding before computing BLEU. Please stay with the token-level bleu for this assignment, however.
```
from nltk.translate.bleu_score import corpus_bleu
def compute_bleu(model, inp_lines, out_lines, bpe_sep='@@ ', **flags):
""" Estimates corpora-level BLEU score of model's translations given inp and reference out """
translations, _ = model.translate_lines(inp_lines, **flags)#TODO batches!
# to compute token-level BLEU we need to split the original data into list of tokens
# translation is already transformed in that form
out_lines = [x.split(' ') for x in out_lines]
return corpus_bleu([[ref] for ref in out_lines], translations) * 100
```
__Note: do not forget splitting strings into list of tokens. Otherwise nltk will split the strings into chars and char-level BLEU will be computed.__
```
compute_bleu(model, dev_inp, dev_out)
```
### Training loop
Training encoder-decoder models isn't that different from any other models: sample batches, compute loss, backprop and update
```
from IPython.display import clear_output
from tqdm import tqdm, trange
metrics = {'train_loss': [], 'dev_bleu': [] }
opt = keras.optimizers.Adam(1e-3)
batch_size = 32
for _ in trange(25000):
step = len(metrics['train_loss']) + 1
batch_ix = np.random.randint(len(train_inp), size=batch_size)
batch_inp = inp_voc.to_matrix(train_inp[batch_ix])
batch_out = out_voc.to_matrix(train_out[batch_ix])
with tf.GradientTape() as tape:
loss_t = compute_loss(model, batch_inp, batch_out)
grads = tape.gradient(loss_t, model.trainable_variables)
opt.apply_gradients(zip(grads, model.trainable_variables))
metrics['train_loss'].append((step, loss_t.numpy()))
if step % 100 == 0:
metrics['dev_bleu'].append((step, compute_bleu(model, dev_inp, dev_out)))
clear_output(True)
plt.figure(figsize=(12,4))
for i, (name, history) in enumerate(sorted(metrics.items())):
plt.subplot(1, len(metrics), i + 1)
plt.title(name)
plt.plot(*zip(*history))
plt.grid()
plt.show()
print("Mean loss=%.3f" % np.mean(metrics['train_loss'][-10:], axis=0)[1], flush=True)
# Note: it's okay if bleu oscillates up and down as long as it gets better on average over long term (e.g. 5k batches)
assert np.mean(metrics['dev_bleu'][-10:], axis=0)[1] > 35, "We kind of need a higher bleu BLEU from you. Kind of right now."
for inp_line, trans_line in zip(dev_inp[::500], model.translate_lines(dev_inp[::500])[0]):
print(inp_line)
print(trans_line)
print()
```
# Homework code templates will appear here soon!
### Your Attention Required (4 points)
In this section we want you to improve over the basic model by implementing a simple attention mechanism.
This is gonna be a two-parter: building the __attention layer__ and using it for an __attentive seq2seq model__.
```
COMMING SOON!
```
### Attention layer
Here you will have to implement a layer that computes a simple additive attention:
Given encoder sequence $ h^e_0, h^e_1, h^e_2, ..., h^e_T$ and a single decoder state $h^d$,
* Compute logits with a 2-layer neural network
$$a_t = linear_{out}(tanh(linear_{e}(h^e_t) + linear_{d}(h_d)))$$
* Get probabilities from logits,
$$ p_t = {{e ^ {a_t}} \over { \sum_\tau e^{a_\tau} }} $$
* Add up encoder states with probabilities to get __attention response__
$$ attn = \sum_t p_t \cdot h^e_t $$
You can learn more about attention layers in the leture slides or [from this post](https://distill.pub/2016/augmented-rnns/).
### Seq2seq model with attention
You can now use the attention layer to build a network. The simplest way to implement attention is to use it in decoder phase:

_image from distill.pub [article](https://distill.pub/2016/augmented-rnns/)_
On every step, use __previous__ decoder state to obtain attention response. Then feed concat this response to the inputs of next attetion layer.
The key implementation detail here is __model state__. Put simply, you can add any tensor into the list of `encode` outputs. You will then have access to them at each `decode` step. This may include:
* Last RNN hidden states (as in basic model)
* The whole sequence of encoder outputs (to attend to) and mask
* Attention probabilities (to visualize)
_There are, of course, alternative ways to wire attention into your network and different kinds of attention. Take a look at [this](https://arxiv.org/abs/1609.08144), [this](https://arxiv.org/abs/1706.03762) and [this](https://arxiv.org/abs/1808.03867) for ideas. And for image captioning/im2latex there's [visual attention](https://arxiv.org/abs/1502.03044)_
```
COMMING SOON!
```
### Training attentive model
We'll reuse the infrastructure you've built for the regular model. I hope you didn't hard-code anything :)
```
COMMING SOON!
```
## Grand Finale (4+ points)
We want you to find the best model for the task. Use everything you know.
* different recurrent units: rnn/gru/lstm; deeper architectures
* bidirectional encoder, different attention methods for decoder
* word dropout, training schedules, anything you can imagine
For a full grade we want you to conduct at least __two__ experiments from two different bullet-points or your alternative ideas (2 points each). Extra work will be rewarded with bonus points :)
As usual, we want you to describe what you tried and what results you obtained.
`[your report/log here or anywhere you please]`
| github_jupyter |
```
from sklearn.base import BaseEstimator, ClassifierMixin, clone
from sklearn.utils import check_X_y, check_random_state, check_array
from sklearn.metrics import get_scorer
from sklearn.utils.validation import column_or_1d, check_is_fitted
from sklearn.multiclass import check_classification_targets
from sklearn.utils.metaestimators import if_delegate_has_method
from sklearn.neighbors import KNeighborsClassifier
from sklearn.preprocessing import MinMaxScaler
from sklearn.feature_selection import mutual_info_classif
import numpy as np
class Knn_Forest(BaseEstimator, ClassifierMixin):
"""
Random feature selection for ensemble of knn classifiers.
Each knn model will view the samples from different perspectives.
Aggregating their views will result in a good ensemble result.
Can also bootstrap the features, optimize the knn params per each
different model and also sample the features based on their initial importance.
"""
def __init__(self,
base_estimator=KNeighborsClassifier(),
n_estimators=10,
random_state=42,
optim=False,
parameters=None,
max_features = 'auto',
bootstrap_feats = False,
feat_importance = [],
metric='accuracy'):
self.base_estimator = base_estimator
self.n_estimators = n_estimators
self.max_features = max_features
self.random_state = check_random_state(random_state)
self.bootstrap_feats = bootstrap_feats
self.optim = optim
self.feat_importance = feat_importance
if self.optim:
self.parameters = parameters
else:
self.parameters = None
self.scoring = get_scorer(metric)
self.ensemble = []
self.selected_feat_indices= []
def fit(self, X, y):
return self._fit(X, y)
def _validate_y(self, y):
y = column_or_1d(y, warn=True)
check_classification_targets(y)
self.classes_, y = np.unique(y, return_inverse=True)
self.n_classes_ = len(self.classes_)
return y
def _fit(self,X,y):
X, y = check_X_y(
X, y, ['csr', 'csc'], dtype=None, force_all_finite=False,
multi_output=True)
y = self._validate_y(y)
n_samples, self.n_features_ = X.shape
if self.max_features == 'auto':
self.N_feats_per_knn = int(np.sqrt(self.n_features_))
elif self.max_features == 'log2':
self.N_feats_per_knn = int(np.log2(self.n_features_))
elif type(self.max_features) == float:
self.N_feats_per_knn = int(self.max_features*self.n_features_)
elif type(self.max_features) == int:
self.N_feats_per_knn = int(self.max_features)
if self.feat_importance == []:
self.feat_probas = [1/float(self.n_features_) for i in xrange(self.n_features_)]
else:
self.feat_probas = self.feat_importance#MinMaxScaler().fit_transform(mutual_info_classif(X, y).reshape(1, -1),y)
print(X.shape[1], self.n_features_, len(self.feat_importance), len(self.feat_probas))
print(len(self.feat_probas), self.n_features_, len(self.feat_importance))
for i_est in xrange(self.n_estimators):
self.selected_feat_indices.append(np.random.choice(np.arange(self.n_features_),
self.N_feats_per_knn,
replace=self.bootstrap_feats,
p=self.feat_probas))
cur_X, cur_y = X[:, self.selected_feat_indices[i_est]], y
cur_mod = clone(self.base_estimator)
if self.optim:
grid_search = GridSearchCV(cur_mod, self.parameters, n_jobs=-1, verbose=0, refit=True)
grid_search.fit(cur_X, cur_y)
cur_mod = grid_search.best_estimator_
else:
cur_mod.fit(cur_X, cur_y)
self.ensemble.append(cur_mod)
#print(cur_X.shape, cur_y.shape)
print("%d ESTIMATORS -- %0.3f" % (len(self.ensemble), 100*accuracy_score(y, self.predict(X), normalize=True)))
return self
def _validate_y(self, y):
y = column_or_1d(y, warn=True)
check_classification_targets(y)
self.classes_, y = np.unique(y, return_inverse=True)
self.n_classes_ = len(self.classes_)
return y
def predict(self, X):
"""Predict class for X.
The predicted class of an input sample is computed as the class with
the highest mean predicted probability. If base estimators do not
implement a ``predict_proba`` method, then it resorts to voting.
Parameters
----------
X : {array-like, sparse matrix} of shape = [n_samples, n_features]
The training input samples. Sparse matrices are accepted only if
they are supported by the base estimator.
Returns
-------
y : array of shape = [n_samples]
The predicted classes.
"""
if hasattr(self.base_estimator, "predict_proba"):
predicted_probability = self.predict_proba(X)
return self.classes_.take((np.argmax(predicted_probability, axis=1)),
axis=0)
else:
predicted_probability = np.zeros((X.shape[0],1), dtype=int)
for i, ens in enumerate(self.ensemble):
predicted_probability = np.hstack((predicted_probability,
ens.predict(X[:, self.selected_feat_indices[i]]).reshape(-1,1)))
predicted_probability = np.delete(predicted_probability,0,axis=1)
final_pred = []
for sample in xrange(X.shape[0]):
final_pred.append(most_common(predicted_probability[sample,:]))
return np.array(final_pred)
def predict_proba(self, X):
"""Predict class probabilities for X.
The predicted class probabilities of an input sample is computed as
the mean predicted class probabilities of the base estimators in the
ensemble. If base estimators do not implement a ``predict_proba``
method, then it resorts to voting and the predicted class probabilities
of an input sample represents the proportion of estimators predicting
each class.
Parameters
----------
X : {array-like, sparse matrix} of shape = [n_samples, n_features]
The training input samples. Sparse matrices are accepted only if
they are supported by the base estimator.
Returns
-------
p : array of shape = [n_samples, n_classes]
The class probabilities of the input samples. The order of the
classes corresponds to that in the attribute `classes_`.
"""
check_is_fitted(self, "classes_")
# Check data
X = check_array(
X, accept_sparse=['csr', 'csc'], dtype=None,
force_all_finite=False
)
if self.n_features_ != X.shape[1]:
raise ValueError("Number of features of the model must "
"match the input. Model n_features is {0} and "
"input n_features is {1}."
"".format(self.n_features_, X.shape[1]))
all_proba = np.zeros((X.shape[0], self.n_classes_))
for i, ens in enumerate(self.ensemble):
all_proba += ens.predict_proba(X[:, self.selected_feat_indices[i]])
all_proba /= self.n_estimators
return all_proba
@if_delegate_has_method(delegate='base_estimator')
def decision_function(self, X):
"""Average of the decision functions of the base classifiers.
Parameters
----------
X : {array-like, sparse matrix} of shape = [n_samples, n_features]
The training input samples. Sparse matrices are accepted only if
they are supported by the base estimator.
Returns
-------
score : array, shape = [n_samples, k]
The decision function of the input samples. The columns correspond
to the classes in sorted order, as they appear in the attribute
``classes_``. Regression and binary classification are special
cases with ``k == 1``, otherwise ``k==n_classes``.
"""
check_is_fitted(self, "classes_")
# Check data
X = check_array(
X, accept_sparse=['csr', 'csc'], dtype=None,
force_all_finite=False
)
if self.n_features_ != X.shape[1]:
raise ValueError("Number of features of the model must "
"match the input. Model n_features is {0} and "
"input n_features is {1} "
"".format(self.n_features_, X.shape[1]))
all_decisions = np.zeros((X.shape[0], self.n_classes_))
for i, ens in enumerate(self.ensemble):
all_decisions += ens.predict_proba(X)
decisions = sum(all_decisions) / self.n_estimators
return decisions
from __future__ import print_function
from pprint import pprint
from time import time
import logging
from sklearn.datasets import fetch_20newsgroups
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.linear_model import SGDClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.metrics import classification_report, confusion_matrix
print(__doc__)
# Display progress logs on stdout
logging.basicConfig(level=logging.INFO,
format='%(asctime)s %(levelname)s %(message)s')
# #############################################################################
# Load some categories from the training set
categories = [
'alt.atheism',
'talk.religion.misc',
]
# Uncomment the following to do the analysis on all the categories
#categories = None
print("Loading 20 newsgroups dataset for categories:")
print(categories)
data = fetch_20newsgroups(subset='train', categories=categories)
print("%d documents" % len(data.filenames))
print("%d categories" % len(data.target_names))
print()
#############################################################################
# Define a pipeline combining a text feature extractor with a simple
# classifier
pipeline = Pipeline([
('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', SGDClassifier()),
])
# uncommenting more parameters will give better exploring power but will
# increase processing time in a combinatorial way
parameters = {
'clf__alpha': (0.00001, 0.000001),
'clf__penalty': ('l2', 'elasticnet'),
'clf__max_iter': (10, 50, 80, 150),
}
X = data.data
y = data.target
if __name__ == "__main__":
# multiprocessing requires the fork to happen in a __main__ protected
# block
# find the best parameters for both the feature extraction and the
# classifier
grid_search = GridSearchCV(pipeline, parameters, n_jobs=-1, verbose=1)
# print("Performing grid search...")
# print("pipeline:", [name for name, _ in pipeline.steps])
# print("parameters:")
# pprint(parameters)
# t0 = time()
# # grid_search.fit(data.data, data.target)
# grid_search.fit(X, y)
# print("done in %0.3fs" % (time() - t0))
# print()
# print("Best score: %0.3f" % grid_search.best_score_)
# print("Best parameters set:")
# best_parameters = grid_search.best_estimator_.get_params()
# for param_name in sorted(parameters.keys()):
# print("\t%s: %r" % (param_name, best_parameters[param_name]))
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
X_train, X_test, y_train, y_test= train_test_split(X, y)
grid_search.fit(X_train, y_train)
cur_mod = grid_search.best_estimator_
pred = cur_mod.predict(X_test)
print(accuracy_score(y_test, pred))
from sklearn.feature_selection import mutual_info_classif
from sklearn.feature_extraction.text import TfidfVectorizer
# pip = Pipeline([
# ('vect', CountVectorizer()),
# ('tfidf', TfidfTransformer())])
# tr = pip.fit_transform(X_train, y_train)
# mi = mutual_info_classif(tr, y_train)
# print(len(mi),tr.shape[1])
mi = mi/sum(mi)
# uncommenting more parameters will give better exploring power but will
# increase processing time in a combinatorial way
parameters = {
'n_neighbors': [2,4,6,8,10],
'metric':['euclidean', 'manhattan', 'cosine', 'l2']
}
parameters2 = {
'clf__max_features': [50, 0.2, 0.3, 0.4,0.8, 'auto', 'log2'],
'clf__bootstrap_feats': [True, False],
'clf__n_estimators': [100,250,500],
'clf__feat_importance':[mi, []]
}
# Define a pipeline combining a text feature extractor with a simple
# classifier
pipeline = Pipeline([
('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', Knn_Forest(n_estimators=500,
max_features=0.2,
bootstrap_feats=False,
optim=False,
parameters=parameters,
feat_importance=mi
)),
])
pipeline.fit(X_train, y_train)
pred = pipeline.predict(X_test)
# grid_search = GridSearchCV(pipeline, parameters2, n_jobs=1, verbose=2)
# grid_search.fit(X_train, y_train)
# cur_mod = grid_search.best_estimator_
# pred = cur_mod.predict(X_test)
print(accuracy_score(y_test, pred))
np.array(mi).shape
if mi == []:
feat_probas = [1/float(15285) for i in xrange(15285)]
else:
feat_probas = mi/float(sum(mi))
print(feat_probas)
from sklearn.feature_selection import mutual_info_classif
from sklearn.feature_extraction.text import TfidfVectorizer
tr = TfidfVectorizer().fit_transform(X_train, y_train)
mi = mutual_info_classif(tr, y_train)
```
| github_jupyter |
This demo provides examples of `ImageReader` class from `niftynet.io.image_reader` module.
What is `ImageReader`?
The main functionality of `ImageReader` is to search a set of folders, return a list of image files, and load the images into memory in an iterative manner.
A `tf.data.Dataset` instance can be initialised from an `ImageReader`, this makes the module readily usable as an input op to many tensorflow-based applications.
Why `ImageReader`?
- designed for medical imaging formats and applications
- works well with multi-modal input volumes
- works well with `tf.data.Dataset`
## Before the demo...
First make sure the source code is available, and import the module.
For NiftyNet installation, please checkout:
http://niftynet.readthedocs.io/en/dev/installation.html
```
import sys
niftynet_path = '/Users/bar/Documents/Niftynet/'
sys.path.append(niftynet_path)
from niftynet.io.image_reader import ImageReader
```
For demonstration purpose we download some demo data to `~/niftynet/data/`:
```
from niftynet.utilities.download import download
download('anisotropic_nets_brats_challenge_model_zoo_data')
```
## Use case: loading 3D volumes
```
from niftynet.io.image_reader import ImageReader
data_param = {'MR': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG'}}
reader = ImageReader().initialise(data_param)
reader.shapes, reader.tf_dtypes
# read data using the initialised reader
idx, image_data, interp_order = reader(idx=0)
image_data['MR'].shape, image_data['MR'].dtype
# randomly sample the list of images
for _ in range(3):
idx, image_data, _ = reader()
print('{} image: {}'.format(idx, image_data['MR'].shape))
```
The images are always read into a 5D-array, representing:
`[height, width, depth, time, channels]`
## User case: loading pairs of image and label by matching filenames
(In this case the loaded arrays are not concatenated.)
```
from niftynet.io.image_reader import ImageReader
data_param = {'image': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'T2'},
'label': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'Label'}}
reader = ImageReader().initialise(data_param)
# image file information (without loading the volumes)
reader.get_subject(0)
idx, image_data, interp_order = reader(idx=0)
image_data['image'].shape, image_data['label'].shape
```
## User case: loading multiple modalities of image and label by matching filenames
The following code initialises a reader with four modalities, and the `'image'` output is a concatenation of arrays loaded from these files. (The files are concatenated at the fifth dimension)
```
from niftynet.io.image_reader import ImageReader
data_param = {'T1': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'T1', 'filename_not_contains': 'T1c'},
'T1c': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'T1c'},
'T2': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'T2'},
'Flair': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'Flair'},
'label': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'Label'}}
grouping_param = {'image': ('T1', 'T1c', 'T2', 'Flair'), 'label':('label',)}
reader = ImageReader().initialise(data_param, grouping_param)
_, image_data, _ = reader(idx=0)
image_data['image'].shape, image_data['label'].shape
```
## More properties
The input specification supports additional properties include
```python
{'csv_file', 'path_to_search',
'filename_contains', 'filename_not_contains',
'interp_order', 'pixdim', 'axcodes', 'spatial_window_size',
'loader'}
```
see also: http://niftynet.readthedocs.io/en/dev/config_spec.html#input-data-source-section
## Using ImageReader with image-level data augmentation layers
```
from niftynet.io.image_reader import ImageReader
from niftynet.layer.rand_rotation import RandomRotationLayer as Rotate
data_param = {'MR': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG'}}
reader = ImageReader().initialise(data_param)
rotation_layer = Rotate()
rotation_layer.init_uniform_angle([-10.0, 10.0])
reader.add_preprocessing_layers([rotation_layer])
_, image_data, _ = reader(idx=0)
image_data['MR'].shape
# import matplotlib.pyplot as plt
# plt.imshow(image_data['MR'][:, :, 50, 0, 0])
# plt.show()
```
## Using ImageReader with `tf.data.Dataset`
```
import tensorflow as tf
from niftynet.io.image_reader import ImageReader
# initialise multi-modal image and label reader
data_param = {'T1': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'T1', 'filename_not_contains': 'T1c'},
'T1c': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'T1c'},
'T2': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'T2'},
'Flair': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'Flair'},
'label': {'path_to_search': '~/niftynet/data/BRATS_examples/HGG',
'filename_contains': 'Label'}}
grouping_param = {'image': ('T1', 'T1c', 'T2', 'Flair'), 'label':('label',)}
reader = ImageReader().initialise(data_param, grouping_param)
# reader as a generator
def image_label_pair_generator():
"""
A generator wrapper of an initialised reader.
:yield: a dictionary of images (numpy arrays).
"""
while True:
_, image_data, _ = reader()
yield image_data
# tensorflow dataset
dataset = tf.data.Dataset.from_generator(
image_label_pair_generator,
output_types=reader.tf_dtypes)
#output_shapes=reader.shapes)
dataset = dataset.batch(1)
iterator = dataset.make_initializable_iterator()
# run the tensorlfow graph
with tf.Session() as sess:
sess.run(iterator.initializer)
for _ in range(3):
data_dict = sess.run(iterator.get_next())
print(data_dict.keys())
print('image: {}, label: {}'.format(
data_dict['image'].shape,
data_dict['label'].shape))
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#MSVO-3,-70" data-toc-modified-id="MSVO-3,-70-1"><span class="toc-item-num">1 </span>MSVO 3, 70</a></span></li><li><span><a href="#Text-Fabric" data-toc-modified-id="Text-Fabric-2"><span class="toc-item-num">2 </span>Text-Fabric</a></span></li><li><span><a href="#Installing-Text-Fabric" data-toc-modified-id="Installing-Text-Fabric-3"><span class="toc-item-num">3 </span>Installing Text-Fabric</a></span><ul class="toc-item"><li><span><a href="#Prerequisites" data-toc-modified-id="Prerequisites-3.1"><span class="toc-item-num">3.1 </span>Prerequisites</a></span></li><li><span><a href="#TF-itself" data-toc-modified-id="TF-itself-3.2"><span class="toc-item-num">3.2 </span>TF itself</a></span></li></ul></li><li><span><a href="#Pulling-up-a-tablet-and-its-transliteration-using-a-p-number" data-toc-modified-id="Pulling-up-a-tablet-and-its-transliteration-using-a-p-number-4"><span class="toc-item-num">4 </span>Pulling up a tablet and its transliteration using a p-number</a></span></li><li><span><a href="#Non-numerical-quads" data-toc-modified-id="Non-numerical-quads-5"><span class="toc-item-num">5 </span>Non-numerical quads</a></span></li><li><span><a href="#Generating-a-list-of-sign-frequency-and-saving-it-as-a-separate-file" data-toc-modified-id="Generating-a-list-of-sign-frequency-and-saving-it-as-a-separate-file-6"><span class="toc-item-num">6 </span>Generating a list of sign frequency and saving it as a separate file</a></span></li></ul></div>
# Primer 1
This notebook is meant for those with little or no familiarity with
[Text-Fabric](https://github.com/annotation/text-fabric) and will focus on several basic tasks, including calling up an individual proto-cuneiform tablet using a p-number, the coding of complex proto-cuneiform signs using what we will call "quads" and the identification of one of the numeral systems, and a quick look at the frequency of a few sign clusters. Each primer, including this one, will focus on a single tablet and explore three or four analytical possibilities. In this primer we look at MSVO 3, 70, which has the p-number P005381 at CDLI.
## MSVO 3, 70
The proto-cuneiform tablet known as MSVO 3, 70, is held in the British Museum, where it has the museum number BM 140852. The tablet dates to the Uruk III period, ca. 3200-3000 BCE, and is slated for publication in the third volume of Materialien zu den frühen Schriftzeugnissen des Vorderen Orients (MSVO). Up to now it has only appeared as a photo in Frühe Schrift (Nissen, Damerow and Englund 1990), p. 38.
We'll show the lineart for this tablet and its ATF transcription in a moment, including a link to this tablet on CDLI.
## Text-Fabric
Text-Fabric (TF) is a model for textual data with annotations that is optimized for efficient data analysis. As we will begin to see at the end of this primer, when we check the totals on the reverse of our primer tablet, Text-Fabric also facilitates the creation of new, derived data, which can be added to the original data.
Working with TF is a bit like buying from IKEA. You get all the bits and pieces in a box, and then you assemble it yourself. TF decomposes any dataset into its components, nicely stacked, with every component uniquely labeled. And then we use short reusable bits of code to do specific things. TF is based on a model proposed by [Doedens](http://books.google.nl/books?id=9ggOBRz1dO4C) that focuses on the essential properties of texts such sequence and embedding. For a description of how Text-Fabric has been used for work on the Hebrew Bible, see Dirk Roorda's article [The Hebrew Bible as Data: Laboratory - Sharing - Experiences](https://doi.org/10.5334/bbi.18).
Once data is transformed into Text-Fabric, it can also be used to build rich online interfaces for specific groups of ancient texts. For the Hebrew Bible, have a look at [SHEBANQ](https://shebanq.ancient-data.org/hebrew/text).
The best environment for using Text-Fabric is in a [Jupyter Notebook](http://jupyter.readthedocs.io/en/latest/). This primer is in a Jupyter Notebook: the snippets of code can only be executed if you have installed Python 3, Jupyter Notebook, and Text-Fabric on your own computer.
## Installing Text-Fabric
### Prerequisites
You need to have Python on your system. Most systems have it out of the box,
but alas, that is python2 and we need at least python 3.6.
Install it from [python.org]() or from [Anaconda]().
If you got it from python.org, you also have to install [Jupyter]().
### TF itself
```
pip install text-fabric
```
if you have installed Python with the help of Anaconda, or
```
sudo -H pip3 install text-fabric
```
if you have installed Python from [python.org](https://www.python.org).
###### Execute: If all this is done, the following cells can be executed.
```
import os, sys, collections
from IPython.display import display
from tf.extra.cunei import Cunei
import sys, os
LOC = ("~/github", "Nino-cunei/uruk", "primer1")
A = Cunei(*LOC)
A.api.makeAvailableIn(globals())
```
## Pulling up a tablet and its transliteration using a p-number
Each cuneiform tablet has a unique "p-number" and we can use this p-number in Text-Fabric to bring up any images and the transliteration of a tablet, here P005381.
There is a "node" in Text-Fabric for this tablet. How do we find it and display the transliteration?
* We *search* for the tablet by means of a template;
* we use functions `A.lineart()` and `A.getSource()` to bring up the lineart and transliterations of tablets.
```
pNum = "P005381"
query = f"""
tablet catalogId={pNum}
"""
results = A.search(query)
```
The `results` is a list of "records".
Here we have only one result: `results[0]`.
Each result record is a tuple of nodes mentioned in the template.
Here we only mentioned a single thing: `tablet`.
So we find the node of the matched tablets as the firt member of the result records.
Hence the result tablet node is `results[0][0]`.
```
tablet = results[0][0]
A.lineart(tablet, width=300)
A.getSource(tablet)
```
Now we want to view the numerals on the tablet.
```
query = f"""
tablet catalogId={pNum}
sign type=numeral
"""
results = A.search(query)
```
It is easy to show them at a glance:
```
A.show(results)
```
Or we can show them in a table.
```
A.table(results)
```
There are a few different types of numerals here, but we are just going to look at the numbers belonging to the "shin prime prime" system, abbreviated here as "shinPP," which regularly adds two narrow horizatonal wedges to each number. N04, which is the basic unit in this system, is the fourth, fith and ninth of the preceding numerals: in the fourth occurrence repeated twice, in the fifth, three times and, unsurprisingly, in the ninth, which is the total on the reverse, five times. (N19, which is the next bundling unit in the same system, also occurs in the text.)
```
shinPP = dict(
N41=0.2,
N04=1,
N19=6,
N46=60,
N36=180,
N49=1800,
)
```
First, let's see if we can locate one of the occurrences of shinPP numerals, namely the set of 3(N04) in the first case of the second column on the obverse, using Text-Fabric.
```
query = f"""
tablet catalogId={pNum}
face type=obverse
column number=2
line number=1
=: sign
"""
results = A.search(query)
A.table(results)
```
Note the `:=` in `=: sign`. This is a device to require that the sign starts at the same position
as the `line` above it. Effectively, we ask for the first sign of the line.
Now the result records are tuples `(tablet, face, column, line, sign)`, so if we want
the sign part of the first result, we have to say `results[0][4]` (Python counts from 0).
```
num = results[0][4]
A.pretty(num, withNodes=True)
```
This number is the "node" in Text-Fabric that corresponds to the first sign in the first case of column 2. It is like a bar-code for that position in the entire corpus. Now let's make sure that this node, viz. 106602, is actually a numeral. To do this we check the feature "numeral" of the node 106602. And then we can use A.atfFromSign to extract the transliteration.
```
print(F.type.v(num) == "numeral")
print(A.atfFromSign(num))
```
Let's get the name of the numeral, viz. N04, and the number of times that it occurs. This amounts to splitting apart "3" and "(N04)" but since we are calling features in Text-Fabric rather than trying to pull elements out of the transliteration, we do not need to tweak the information.
```
grapheme = F.grapheme.v(num)
print(grapheme)
iteration = F.repeat.v(num)
print(iteration)
```
Now we can replace "N04" with its value, using the shinPP dictionary above, and then multiple this value by the number of iterations to arrive at the value of the numeral as a whole. Since each occurrence of the numeral N04 has a value of 1, three occurrences of it should have a value of 3.
```
valueFromDict = shinPP.get(grapheme)
value = iteration * valueFromDict
print(value)
```
Just to make sure that we are calculating these values correctly, let's try it again with a numeral whose value is not 1. There is a nice example in case 1b in column 1 on the obverse, where we have 3 occurrences of N19, each of which has a value of 6, so we expect the total value of 3(N19 to be 18.
```
query = f"""
tablet catalogId={pNum}
face type=obverse
column number=1
case number=1b
=: sign
"""
results = A.search(query)
A.table(results)
sign = results[0][4]
grapheme = F.grapheme.v(sign)
iteration = F.repeat.v(sign)
valueFromDict = shinPP.get(grapheme, 0)
value = iteration * valueFromDict
print(value)
```
The next step is to walk through the nodes on the obverse, add up the total of the shinPP system on the obverse, and then do the same for the reverse and see if the obverse and the total on the reverse add up. We expect the 3(N19) and 5(N04) on the obverse to add up to 23, viz. 18 + 5 = 23.
```
shinPPpat = "|".join(shinPP)
query = f"""
tablet catalogId={pNum}
face
sign grapheme={shinPPpat}
"""
results = A.search(query)
A.show(results)
sums = collections.Counter()
for (tablet, face, num) in results:
grapheme = F.grapheme.v(num)
iteration = F.repeat.v(num)
valueFromDict = shinPP[grapheme]
value = iteration * valueFromDict
sums[F.type.v(face)] += value
for faceType in sums:
print(f"{faceType}: {sums[faceType]}")
```
It adds up!
## Non-numerical quads
Now that we have identified the numeral system in the first case of column 2 on the obverse, let's also see what we can find out about the non-numeral signs in the same case.
We use the term "quad" to refer to all orthographic elements that occupy the space of a single proto-cuneiform sign on the surface of the tablet. This includes both an individual proto-cuneiform sign operating on its own as well as combinations of signs that occupy the same space. One of the most elaborate quads in the proto-cuneiform corpus is the following:
```
|SZU2.((HI+1(N57))+(HI+1(N57)))|
```
This quad has two sub-quads `SZU2`, `(HI+1(N57))+(HI+1(N57))`, and the second sub-quad also consists of two sub-quads `HI+1(N57)` and `HI+1(N57)`; both of these sub-quads are, in turn, composed of two further sub-quads `HI` and `1(N57)`.
First we need to pick this super-quad out of the rest of the line: this is how we get the transliteration of the entire line:
```
query = f"""
tablet catalogId={pNum}
face type=obverse
column number=2
line number=1
"""
results = A.search(query)
line = results[0][3]
A.pretty(line, withNodes=True)
```
We can just read off the node of the biggest quad.
```
bigQuad = 143015
```
Now that we have identified the "bigQuad," we can also ask Text-Fabric to show us what it looks like.
```
A.lineart(bigQuad)
```
This extremely complex quad, viz. |SZU2.((HI+1(N57))+(HI+1(N57)))|, is a hapax legomenon, meaning that it only occurs here, but there are three other non-numeral quads in this line besides |SZU2.((HI+1(N57))+(HI+1(N57)))|, namely |GISZ.TE|, GAR and GI4~a, so let's see how frequent these four non-numeral signs are in the proto-cuneiform corpus. We can do this sign by sign using the function "F.grapheme.s()".
```
GISZTEs = F.grapheme.s("|GISZ.TE|")
print(f"|GISZ.TE| {len(GISZTEs)} times")
GARs = F.grapheme.s("GAR")
print(f"GAR = {len(GARs)} times")
GI4s = F.grapheme.s("GI4")
print(f"GI4 = {len(GI4s)} times")
```
There are two problems here that we need to resolve in order to get good numbers: we have to get Text-Fabric to count |GISZ.TE| as a single unit, even though it is composed of two distinct graphemes, and we have to ask it to recognize and count the "a" variant of "GI4". In order to count the number of quads that consist of GISZ and TE, namely |GISZ.TE|, it is convenient to make a frequency index for all quads.
We walk through all the quads, pick up its ATF, and count the frequencies of ATF representations.
```
quadFreqs = collections.Counter()
for q in F.otype.s("quad"):
quadFreqs[A.atfFromQuad(q)] += 1
```
With this in hand, we can quickly count how many quads there are that have both signs `GISZ` and `TE` in them.
Added bonus: we shall also see whether there are quads with both of these signs but composed with other operators and signs as well.
```
for qAtf in quadFreqs:
if "GISZ" in qAtf and "TE" in qAtf:
print(f"{qAtf} ={quadFreqs[qAtf]:>4} times")
```
And we can also look at the set of quads in which GISZ co-occurs with another sign, and likewise, the set of quads in which TE co-occurs with another sign.
```
for qAtf in quadFreqs:
if "GISZ" in qAtf:
print(f"{quadFreqs[qAtf]:>4} x {qAtf}")
for qAtf in quadFreqs:
if "TE" in qAtf:
print(f"{quadFreqs[qAtf]:>4} x {qAtf}")
```
Most of the time, however, when we are interested in particular sign frequencies, we want to cast a wide net and get the frequency of any possibly related sign or quad. The best way to do this is to check the ATF of any sign or quad that might be relevant and add up the number of its occurrences in the corpus. This following script checks both signs and quads and casts the net widely. It looks for the frequency of our same three signs/quads, namely GAR, GI4~a and |GISZ.TE|.
```
quadSignFreqs = collections.Counter()
quadSignTypes = {"quad", "sign"}
for n in N():
nType = F.otype.v(n)
if nType not in quadSignTypes:
continue
atf = A.atfFromQuad(n) if nType == "quad" else A.atfFromSign(n)
quadSignFreqs[atf] += 1
```
We have now an frequency index for all signs and quads in their ATF representation.
Note that if a sign is part of a bigger quad, its occurrence there will be counted as an occurrence of the sign.
```
selectedAtfs = []
for qsAtf in quadSignFreqs:
if "GAR" in qsAtf or "GI4~a" in qsAtf or "|GISZ.TE|" in qsAtf:
selectedAtfs.append(qsAtf)
print(f"{quadSignFreqs[qsAtf]:>4} x {qsAtf}")
```
Let's draw all these quads.
```
for sAtf in selectedAtfs:
A.lineart(sAtf, width="5em", height="5em", withCaption="right")
```
Besides our three targets, 34 occurrences of GI4~a, 401 of GAR and 26 of |GISZ.TE|:
34 x GI4~a
401 x GAR
26 x |GISZ.TE|
it has also pulled in a number of quads that include either GAR or GI4~a, among others:
20 x |ZATU651xGAR|
3 x |NINDA2xGAR|
6 x |4(N57).GAR|
1 x |GI4~a&GI4~a|
1 x |GI4~axA|
There are also other signs tas well as signs that only resemble GAR in transliteration such as LAGAR or GARA2, but as long as we know what we are looking for this type of broader frequency count can be quite useful.
## Generating a list of sign frequency and saving it as a separate file
First, we are going to count the number of distinct signs in the corpus, look at the top hits in the list and finally save the full list to a separate file. Then we will do the same for the quads, and then lastly we are going to combine these two lists and save them as a single frequency list for both signs and quads.
```
fullGraphemes = collections.Counter()
for n in F.otype.s("sign"):
grapheme = F.grapheme.v(n)
if grapheme == "" or grapheme == "…":
continue
fullGrapheme = A.atfFromSign(n)
fullGraphemes[fullGrapheme] += 1
len(fullGraphemes)
```
So there are 1477 distinct proto-cuneiform signs in the corpus. The following snippet of code will show us the first 20 signs on that list.
```
for (value, frequency) in sorted(
fullGraphemes.items(),
key=lambda x: (-x[1], x[0]),
)[0:20]:
print(f"{frequency:>5} x {value}")
```
Now we are going to write the full set of sign frequency results to two files in your `_temp` directory, within this repo. The two files are called:
* `grapheme-alpha.txt`, an alphabetic list of graphemes, along with the frequency of each sign, and
* `grapheme-freq.txt`, which runs from the most frequent to the least.
```
def writeFreqs(fileName, data, dataName):
print(f"There are {len(data)} {dataName}s")
for (sortName, sortKey) in (
("alpha", lambda x: (x[0], -x[1])),
("freq", lambda x: (-x[1], x[0])),
):
with open(f"{A.tempDir}/{fileName}-{sortName}.txt", "w") as fh:
for (item, freq) in sorted(data, key=sortKey):
if item != "":
fh.write(f"{freq:>5} x {item}\n")
```
Now let's go through some of the same steps for quads rather than individual signs, and then export a single frequency list for both signs and quads.
```
quadFreqs = collections.Counter()
for q in F.otype.s("quad"):
quadFreqs[A.atfFromQuad(q)] += 1
print(len(quadFreqs))
```
So there are 740 quads in the corpus, and now we ask for the twenty most frequently attested quads.
```
for (value, frequency) in sorted(
quadFreqs.items(),
key=lambda x: (-x[1], x[0]),
)[0:20]:
print(f"{frequency:>5} x {value}")
```
And for the final task in this primer, we ask Text-Fabric to export a frequency list of both signs and quads in a separate file.
```
reportDir = "reports"
os.makedirs(reportDir, exist_ok=True)
def writeFreqs(fileName, data, dataName):
print(f"There are {len(data)} {dataName}s")
for (sortName, sortKey) in (
("alpha", lambda x: (x[0], -x[1])),
("freq", lambda x: (-x[1], x[0])),
):
with open(f"{reportDir}/{fileName}-{sortName}.txt", "w") as fh:
for (item, freq) in sorted(data.items(), key=sortKey):
if item != "":
fh.write(f"{freq:>5} x {item}\n")
```
This shows up as a pair of files named "quad-signs-alpha.txt" and "quad-signs-freq.txt" and if we copy a few pieces of the quad-signs-freq.txt file here, they look something like this:
29413 x ...
12983 x 1(N01)
6870 x X
3080 x 2(N01)
2584 x 1(N14)
1830 x EN~a
1598 x 3(N01)
1357 x 2(N14)
1294 x 5(N01)
1294 x SZE~a
1164 x GAL~a
Only much farther down the list do we see signs and quads interspersed; here are the signs/quads around 88 occurrences:
88 x NIMGIR
88 x NIM~a
88 x SUG5
86 x EN~b
86 x NAMESZDA
86 x |GI&GI|
85 x GU
85 x |GA~a.ZATU753|
84 x BAD~a
84 x NA2~a
84 x ZATU651
84 x |1(N58).BAD~a|
83 x ZATU759
| github_jupyter |
# Math Kernel Library (oneMKL) and SYCL Basic Parallel Kernel
In this next set of modules, we will explore how we can utilize oneAPI and SYCL to implement Matrix Multiplication using Intel® oneAPI Math Kernel Library (oneMKL) and also implement Matrix Multiplication in the most basic parallel form and then improve performance by tuning the kernel code while trying to maintain performance portability. All code improvements will be measured in terms of relative performance to oneMKL.
### Learning Objectives
- Gain familiarity with oneMKL and able to use it for a two dimensional GEMM.
- Use a basic GEMM application for basis of enhancements.
- Interpret roofline and VTune™ analyzer results as a method to measure the GEMM applications.
## Intel oneAPI Math Kernel Library (oneMKL)
One of the best ways to achieve performance portable code is to take advantage of a library. In this case oneMKL offers a compelling GEMM implementation that we will use as our baseline. If there is a library, it should always be ones first attempt at achieving performant portable code. All other implementations will be measured against oneMKL.
Intel oneMKL is included in the Intel oneAPI toolkits and there is extensive documentation at [Get Started with Intel oneAPI Math Kernel Library.](https://software.intel.com/content/www/us/en/develop/documentation/get-started-with-mkl-for-dpcpp/top.html)
The Intel® oneAPI Math Kernel Library (oneMKL) helps you achieve maximum performance with a math computing library of highly optimized, extensively parallelized routines for CPU and GPU. The library has C and Fortran interfaces for most routines on CPU, and SYCL interfaces for some routines on both CPU and GPU. You can find comprehensive support for several math operations in various interfaces including:
SYCL on CPU and GPU
(Refer to the Intel® oneAPI Math Kernel Library—Data Parallel C++ Developer Reference for more details.)
- Linear algebra
- BLAS
- Selected Sparse BLAS functionality
- Selected LAPACK functionality
- Fast Fourier Transforms (FFT)
- 1D r2c FFT
- 1D c2c FFT
- Random number generators
- Single precision Uniform, Gaussian, and Lognormal distributions
- Selected Vector Math functionality
The example below uses the GEMM function from the oneMKL BLAS routine, which computes a scalar-matrix-matrix product and add the result to a scalar-matrix product, with general matrices. The operation is defined as:
```cpp
void gemm(queue &exec_queue, transpose transa, transpose transb, std::int64_t m, std::int64_t n, std::int64_t k, T alpha, buffer<T, 1> &a, std::int64_t lda, buffer<T, 1> &b, std::int64_t ldb, T beta, buffer<T, 1> &c, std::int64_t ldc)
```
This one line of oneMKL function does all of the necessary optimizations for CPU/GPU offload compute and as you go through the exercises you will discover that it did indeed deliver the best results with the least amount of code across all of the platforms.
## Matrix Multiplication with Math Kernel Library (oneMKL)
The following SYCL code below uses a oneMKL kernel: Inspect code, there are no modifications necessary:
1. __Run__ ▶the cell following __Select offload device__, in Jupyter everything is linear, a subsequent run will need to choose a new target and then the following cell will need to be executed to get the updated results.
2. Inspect the following code cell and click __Run__ ▶ to save the code to a file.
3. Next run -- the cell in the __Build and Run__ section below the code to compile and execute the code.
#### Select Offload Device
```
run accelerator.py
%%writefile lab/mm_dpcpp_mkl.cpp
//==============================================================
// Matrix Multiplication: SYCL oneMKL
//==============================================================
// Copyright © 2021 Intel Corporation
//
// SPDX-License-Identifier: MIT
// =============================================================
#include <CL/sycl.hpp>
#include "oneapi/mkl/blas.hpp" //# oneMKL DPC++ interface for BLAS functions
using namespace sycl;
void mm_kernel(queue &q, std::vector<float> &matrix_a, std::vector<float> &matrix_b, std::vector<float> &matrix_c, size_t N, size_t M) {
std::cout << "Configuration : MATRIX_SIZE= " << N << "x" << N << "\n";
//# Create buffers for matrices
buffer a(matrix_a);
buffer b(matrix_b);
buffer c(matrix_c);
//# scalar multipliers for oneMKL
float alpha = 1.f, beta = 1.f;
//# transpose status of matrices for oneMKL
oneapi::mkl::transpose transA = oneapi::mkl::transpose::nontrans;
oneapi::mkl::transpose transB = oneapi::mkl::transpose::nontrans;
//# Submit MKL library call to execute on device
oneapi::mkl::blas::gemm(q, transA, transB, N, N, N, alpha, b, N, a, N, beta, c, N);
c.get_access<access::mode::read>();
}
```
#### Build and Run
Select the cell below and click __Run__ ▶ to compile and execute the code on selected device:
```
! chmod 755 q; chmod 755 run_mm_mkl.sh; if [ -x "$(command -v qsub)" ]; then ./q run_mm_mkl.sh "{device.value}"; else ./run_mm_mkl.sh; fi
```
### Roofline Report
Execute the following line to display the roofline results
```
run display_data/mm_mkl_roofline.py
```
### VTune™ Profiler Summary
Execute the following line to display the VTune results.
```
run display_data/mm_mkl_vtune.py
```
## Basic Parallel Kernel Implementation
In this section we will look at how matrix multiplication can be implemented using a SYCL basic parallel kernel. This is the most simplest implementation using SYCL without any optimizations. In the next few modules we will add optimization on top of this implementation to improve the performance.
<img src="Assets/naive.PNG">
We can define the kernel with `parallel_for` with a 2-dimentional range for the matrix and perform matrix multiplication as shown below:
```cpp
h.parallel_for(range<2>{N,N}, [=](item<2> item){
const int i = item.get_id(0);
const int j = item.get_id(1);
for (int k = 0; k < N; k++) {
C[i*N+j] += A[i*N+k] * B[k*N+j];
}
});
```
## Matrix Multiplication with basic parallel kernel
The following SYCL code shows the basic parallel kernel implementation of matrix multiplication. Inspect code; there are no modifications necessary:
1. Run the cell in the __Select Offload Device__ section to choose a target device to run the code on.
2. Inspect the following code cell and click __Run__ ▶ to save the code to a file.
3. Next, run the cell in the __Build and Run__ section to compile and execute the code.
#### Select Offload Device
```
run accelerator.py
%%writefile lab/mm_dpcpp_basic.cpp
//==============================================================
// Matrix Multiplication: SYCL Basic Parallel Kernel
//==============================================================
// Copyright © 2021 Intel Corporation
//
// SPDX-License-Identifier: MIT
// =============================================================
#include <CL/sycl.hpp>
using namespace sycl;
void mm_kernel(queue &q, std::vector<float> &matrix_a, std::vector<float> &matrix_b, std::vector<float> &matrix_c, size_t N, size_t M) {
std::cout << "Configuration : MATRIX_SIZE= " << N << "x" << N << "\n";
//# Create buffers for matrices
buffer a(matrix_a);
buffer b(matrix_b);
buffer c(matrix_c);
//# Submit command groups to execute on device
auto e = q.submit([&](handler &h){
//# Create accessors to copy buffers to the device
auto A = a.get_access<access::mode::read>(h);
auto B = b.get_access<access::mode::read>(h);
auto C = c.get_access<access::mode::write>(h);
//# Parallel Compute Matrix Multiplication
h.parallel_for(range<2>{N,N}, [=](item<2> item){
const int i = item.get_id(0);
const int j = item.get_id(1);
for (int k = 0; k < N; k++) {
C[i*N+j] += A[i*N+k] * B[k*N+j];
}
});
});
c.get_access<access::mode::read>();
//# print kernel compute duration from event profiling
auto kernel_duration = (e.get_profiling_info<info::event_profiling::command_end>() - e.get_profiling_info<info::event_profiling::command_start>());
std::cout << "Kernel Execution Time : " << kernel_duration / 1e+9 << " seconds\n";
}
```
#### Build and Run
Select the cell below and click __Run__ ▶ to compile and execute the code on selected device:
```
! chmod 755 q; chmod 755 run_mm_basic.sh;if [ -x "$(command -v qsub)" ]; then ./q run_mm_basic.sh "{device.value}"; else ./run_mm_basic.sh; fi
```
### Roofline Report
Execute the following line to display the roofline results
```
run display_data/mm_basic_roofline.py
```
### VTune™ Profiler Summary
Execute the following line to display the VTune results.
```
run display_data/mm_basic_vtune.py
```
### Analysis
Comparing the execution times for Basic SYCL implementation and Math Kernel Library implementation for various matrix sizes, we can see that for small matrix size of 1024x1024, Basic SYCL implementation performs better than MKL implementation. When matrix size is large, MKL implementation out performs Basic SYCL implementation significantly. The graph below shows execution times on various hardware for matrix sizes 1024x1024, 5120x5120 and 10240x10240.
<img src=Assets/ppp_basic_mkl_graph.PNG>
### Summary
In this module we looked at oneAPI Math Kernel Library (oneMKL) and implemented matrix multiplication using oneMKL function. We also implemented matrix multiplication using SYCL basic parallel kernel. We compared performance numbers for the two implementations and can see benefits of using a library link oneMKL to implement computation rather than basic implementation using SYCL.
## Resources
Check out these related resources
#### Intel® oneAPI Toolkit documentation
* [Intel Advisor Roofline](https://software.intel.com/content/www/us/en/develop/articles/intel-advisor-roofline.html)
* [Intel VTune](https://software.intel.com/content/www/us/en/develop/documentation/vtune-help/top/introduction.html)
* [Intel® oneAPI main page](https://software.intel.com/oneapi "oneAPI main page")
* [Intel® oneAPI programming guide](https://software.intel.com/sites/default/files/oneAPIProgrammingGuide_3.pdf "oneAPI programming guide")
* [Intel® DevCloud Signup](https://software.intel.com/en-us/devcloud/oneapi "Intel DevCloud") Sign up here if you do not have an account.
* [Get Started with oneAPI for Linux*](https://software.intel.com/en-us/get-started-with-intel-oneapi-linux)
* [Get Started with oneAPI for Windows*](https://software.intel.com/en-us/get-started-with-intel-oneapi-windows)
* [Intel® oneAPI Code Samples](https://software.intel.com/en-us/articles/code-samples-for-intel-oneapibeta-toolkits)
* [oneAPI Specification elements](https://www.oneapi.com/spec/)
#### SYCL
* [SYCL* 2020 Specification](https://www.khronos.org/registry/SYCL/specs/sycl-2020/pdf/sycl-2020.pdf)
#### Modern C++
* [CPPReference](https://en.cppreference.com/w/)
* [CPlusPlus](http://www.cplusplus.com/)
***
| github_jupyter |
```
from resources.workspace import *
%matplotlib inline
```
## Dynamical systems
are systems (sets of equations) whose variables evolve in time (the equations contains time derivatives). As a branch of mathematics, its theory is mainly concerned with understanding the behaviour of solutions (trajectories) of the systems.
## Chaos
is also known as the butterfly effect: "a buttefly that flaps its wings in Brazil can 'cause' a hurricane in Texas".
As opposed to the opinions of Descartes/Newton/Laplace, chaos effectively means that even in a deterministic (non-stochastic) universe, we can only predict "so far" into the future. This will be illustrated below using two toy-model dynamical systems made by Edward Lorenz.
---
## The Lorenz (1963) attractor
The [Lorenz-63 dynamical system](https://en.wikipedia.org/wiki/Lorenz_system) can be derived as an extreme simplification of *Rayleigh-Bénard convection*: fluid circulation in a shallow layer of fluid uniformly heated (cooled) from below (above).
This produces the following 3 *coupled* ordinary differential equations (ODE):
$$
\begin{aligned}
\dot{x} & = \sigma(y-x) \\
\dot{y} & = \rho x - y - xz \\
\dot{z} & = -\beta z + xy
\end{aligned}
$$
where the "dot" represents the time derivative, $\frac{d}{dt}$. The state vector is $\mathbf{x} = (x,y,z)$, and the parameters are typically set to
```
SIGMA = 10.0
BETA = 8/3
RHO = 28.0
```
The ODEs can be coded as follows
```
def dxdt(xyz, t0, sigma, beta, rho):
"""Compute the time-derivative of the Lorenz-63 system."""
x, y, z = xyz
return [
sigma * (y - x),
x * (rho - z) - y,
x * y - beta * z
]
```
#### Numerical integration to compute the trajectories
Below is a function to numerically **integrate** the ODEs and **plot** the solutions.
<!--
This function also takes arguments to control ($\sigma$, $\beta$, $\rho$) and of the numerical integration (`N`, `T`).
-->
```
from scipy.integrate import odeint
output_63 = [None]
@interact( sigma=(0.,50), beta=(0.,5), rho=(0.,50), N=(0,50), eps=(0.01,1), T=(0.,40))
def animate_lorenz(sigma=SIGMA, beta=BETA, rho=RHO , N=2, eps=0.01, T=1.0):
# Initial conditions: perturbations around some "proto" state
seed(1)
x0_proto = array([-6.1, 1.2, 32.5])
x0 = x0_proto + eps*randn((N, 3))
# Compute trajectories
tt = linspace(0, T, int(100*T)+1) # Time sequence for trajectory
dd = lambda x,t: dxdt(x,t, sigma,beta,rho) # Define dxdt(x,t) with fixed params.
xx = array([odeint(dd, xn, tt) for xn in x0]) # Integrate
output_63[0] = xx
# PLOTTING
ax = plt.figure(figsize=(10,5)).add_subplot(111, projection='3d')
ax.axis('off')
colors = plt.cm.jet(linspace(0,1,N))
for n in range(N):
ax.plot(*(xx[n,:,:].T),'-' ,c=colors[n])
ax.scatter3D(*xx[n,-1,:],s=40,c=colors[n])
```
**Exc 2**:
* Move `T` (use your arrow keys). What does it control?
* Set `T` to something small; move the sliders for `N` and `eps`. What do they control?
* Visually investigate the system's (i.e. the trajectories') sensititivy to initial conditions by moving `T`, `N` and `eps`. How long do you think it takes (on average) for two trajectories (or the estimation error) to grow twice as far apart as they started (alternatives: 0.03, 0.3, 3, 30)?
### Averages
Slide `N` and `T` to their upper bounds. Execute the code cell below.
```
# Compute the average location of the $m$-th component of the state in TWO ways.
m = 0 # state component index (must be 0,1,2)
nB = 20
xx = output_63[0][:,:,m]
plt.hist(xx[:,-1] ,normed=1,bins=nB, label="ensemble dist.",alpha=1.0) # -1: last time
plt.hist(xx[-1,:] ,normed=1,bins=nB, label="temporal dist.",alpha=0.5) # -1: last ensemble member
#plt.hist(xx.ravel(),normed=1,bins=nB, label="total distribution",alpha=0.5)
plt.legend();
```
**Exc 6*:** Answer the questions below.
* (a) Do you think the samples behind the histograms are drawn from the same distribution?
* (b) The answer to the above question means that this dynamical system is [ergodic](https://en.wikipedia.org/wiki/Ergodic_theory#Ergodic_theorems).
Now, suppose we want to investigate which (DA) method is better at estimating the true state (trajectory) for this system, on average. Should we run several short experiments or one long one?
```
#show_answer("Ergodicity a")
#show_answer("Ergodicity b")
```
---
## The "Lorenz-95" model
The Lorenz-96 system
is a "1D" model, designed to resemble atmospheric convection. Each state variable $\mathbf{x}_m$ can be considered some atmospheric quantity at grid point at a fixed lattitude of the earth. The system
is given by the coupled set of ODEs,
$$
\frac{d \mathbf{x}_m}{dt} = (\mathbf{x}_{m+1} − \mathbf{x}_{m-2}) \mathbf{x}_{m-1} − \mathbf{x}_m + F
\, ,
\quad \quad m \in \{1,\ldots,M\}
\, ,
$$
where the subscript indices apply periodically.
This model is not derived from physics but has similar characterisics, such as
<ul>
<li> there is external forcing, determined by a parameter $F$;</li>
<li> there is internal dissipation, emulated by the linear term;</li>
<li> there is energy-conserving advection, emulated by quadratic terms.</li>
</ul>
[Further description in the very readable original article](http://eaps4.mit.edu/research/Lorenz/Predicability_a_Problem_2006.pdf).
**Exc 10:** Show that the "total energy" $\sum_{m=1}^{M} \mathbf{x}_m^2$ is preserved by the quadratic terms in the ODE.
```
#show_answer("Hint: Lorenz energy")
#show_answer("Lorenz energy")
```
The model is animated below.
```
# For all m, any n: s(x,n) := x[m+n], circularly.
def s(x,n):
return np.roll(x,-n)
output_95 = [None]
@interact( M=(5,60,1), Force=(0,40,1), eps=(0.01,3,0.1), T=(0.05,40,0.05))
def animate_lorenz_95(M=40, Force=8.0, eps=0.01,T=0):
# Initial conditions: perturbations
x0 = zeros(M)
x0[0] = eps
def dxdt(x,t):
return (s(x,1)-s(x,-2))*s(x,-1) - x + Force
tt = linspace(0, T, int(40*T)+1)
xx = odeint(lambda x,t: dxdt(x,t), x0, tt)
output_95[0] = xx
plt.figure(figsize=(7,4))
# Plot last only
#plt.plot(xx[-1],'b')
# Plot multiple
Lag = 8
colors = plt.cm.cubehelix(0.1+0.6*linspace(0,1,Lag))
for k in range(Lag,0,-1):
plt.plot(xx[max(0,len(xx)-k)],c=colors[Lag-k])
plt.ylim(-10,20)
```
**Exc 12:** Investigate by moving the sliders: Under which settings of the force `F` is the system chaotic (is the predictability horizon finite)?
---
## Error/perturbation dynamics
**Exc 14*:** Suppose $x(t)$ and $z(t)$ are "twins": they evolve according to the same law $f$:
$$
\begin{align}
\frac{dx}{dt} &= f(x) \\
\frac{dz}{dt} &= f(z) \, .
\end{align}
$$
Define the "error": $\varepsilon(t) = x(t) - z(t)$.
Suppose $z(0)$ is close to $x(0)$.
Let $F = \frac{df}{dx}(x(t))$.
* a) Show that the error evolves according to the ordinary differential equation (ODE)
$$\frac{d \varepsilon}{dt} \approx F \varepsilon \, .$$
```
#show_answer("error evolution")
```
* b) Suppose $F$ is constant. Show that the error grows exponentially: $\varepsilon(t) = \varepsilon(0) e^{F t} $.
```
#show_answer("anti-deriv")
```
* c)
* 1) Suppose $F<1$.
What happens to the error?
What does this mean for predictability?
* 2) Now suppose $F>1$.
Given that all observations are uncertain (i.e. $R_t>0$, if only ever so slightly),
can we ever hope to estimate $x(t)$ with 0 uncertainty?
```
#show_answer("predictability cases")
```
* d) Consider the ODE derived above.
How might we change it in order to model (i.e. emulate) a saturation of the error at some level?
Can you solve this equation?
```
#show_answer("saturation term")
```
* e) Now suppose $z(t)$ evolves according to $\frac{dz}{dt} = g(z)$, with $g \neq f$.
What is now the differential equation governing the evolution of the error, $\varepsilon$?
```
#show_answer("liner growth")
```
**Exc 16*:** Recall the Lorenz-63 system. What is its doubling time (i.e. estimate how long does it take for two trajectories to grow twice as far apart as they were to start with) ?
*Hint: Set `N=50, eps=0.01, T=1,` and compute the spread of the particles now as compared to how they started*
```
xx = output_63[0][:,-1] # Ensemble of particles at the end of integration
### compute your answer here ###
#show_answer("doubling time")
```
The answer actually depends on where in "phase space" the particles started.
To get a universal answer one must average these experiments for many different initial conditions.
## In summary:
Prediction (forecasting) with these systems is challenging because they are chaotic: small errors grow exponentially.
In other words, chaos means that there is a limit to how far into the future we can make predictions (skillfully).
It is therefore crucial to minimize the intial error as much as possible. This is a task for DA.
### Next: [Ensemble [Monte-Carlo] approach](T7 - Ensemble [Monte-Carlo] approach.ipynb)
| github_jupyter |
## 102 - Training Regression Algorithms with the L-BFGS Solver
In this example, we run a linear regression on the *Flight Delay* dataset to predict the delay times.
We demonstrate how to use the `TrainRegressor` and the `ComputePerInstanceStatistics` APIs.
First, import the packages.
```
import numpy as np
import pandas as pd
import mmlspark
```
Next, import the CSV dataset.
```
# load raw data from small-sized 30 MB CSV file (trimmed to contain just what we use)
dataFilePath = "On_Time_Performance_2012_9.csv"
import os, urllib
if not os.path.isfile(dataFilePath):
urllib.request.urlretrieve("https://mmlspark.azureedge.net/datasets/" + dataFilePath,
dataFilePath)
flightDelay = spark.createDataFrame(
pd.read_csv(dataFilePath,
dtype={"Month": np.float64, "Quarter": np.float64,
"DayofMonth": np.float64, "DayOfWeek": np.float64,
"OriginAirportID": np.float64, "DestAirportID": np.float64,
"CRSDepTime": np.float64, "CRSArrTime": np.float64}))
# Print information on the dataset we loaded
print("Records read: " + str(flightDelay.count()))
print("Schema:")
flightDelay.printSchema()
flightDelay.limit(10).toPandas()
```
Split the dataset into train and test sets.
```
train,test = flightDelay.randomSplit([0.75, 0.25])
```
Train a regressor on dataset with `l-bfgs`.
```
from mmlspark import TrainRegressor, TrainedRegressorModel
from pyspark.ml.regression import LinearRegression
from pyspark.ml.feature import StringIndexer
# Convert columns to categorical
catCols = ["Carrier", "DepTimeBlk", "ArrTimeBlk"]
trainCat = train
testCat = test
for catCol in catCols:
simodel = StringIndexer(inputCol=catCol, outputCol=catCol + "Tmp").fit(train)
trainCat = simodel.transform(trainCat).drop(catCol).withColumnRenamed(catCol + "Tmp", catCol)
testCat = simodel.transform(testCat).drop(catCol).withColumnRenamed(catCol + "Tmp", catCol)
lr = LinearRegression().setRegParam(0.1).setElasticNetParam(0.3)
model = TrainRegressor(model=lr, labelCol="ArrDelay").fit(trainCat)
model.write().overwrite().save("flightDelayModel.mml")
```
Score the regressor on the test data.
```
flightDelayModel = TrainedRegressorModel.load("flightDelayModel.mml")
scoredData = flightDelayModel.transform(testCat)
scoredData.limit(10).toPandas()
```
Compute model metrics against the entire scored dataset
```
from mmlspark import ComputeModelStatistics
metrics = ComputeModelStatistics().transform(scoredData)
metrics.toPandas()
```
Finally, compute and show per-instance statistics, demonstrating the usage
of `ComputePerInstanceStatistics`.
```
from mmlspark import ComputePerInstanceStatistics
evalPerInstance = ComputePerInstanceStatistics().transform(scoredData)
evalPerInstance.select("ArrDelay", "Scores", "L1_loss", "L2_loss").limit(10).toPandas()
```
| github_jupyter |
### Grade: 6 / 8 -- take a look at TA-COMMENTS (you can Command + F to search for "TA-COMMENT")
I don't have the spotify portion of your HW5 -- is it somewhere else in your repo? Let me know because that part of the homework is another 8 points.
# 1) With "Lil Wayne" and "Lil Kim" there are a lot of "Lil" musicians. Do a search and print a list of 50 that are playable in the USA (or the country of your choice), along with their popularity score.
```
# !pip3 install requests
import requests
response = requests.get('https://api.spotify.com/v1/search?query=Lil+&offset=0&limit=50&type=artist&market=US')
data = response.json()
data.keys()
artist_data = data['artists']['items']
for artist in artist_data:
print(artist['name'], artist['popularity'], artist['genres'])
```
# 2) What genres are most represented in the search results? Edit your previous printout to also display a list of their genres in the format "GENRE_1, GENRE_2, GENRE_3". If there are no genres, print "No genres listed".
```
genre_list = []
for artist in artist_data:
for genre in artist['genres']:
genre_list.append(genre)
print(genre_list)
genre_count = 0
# TA-COMMENT: (-0.5) This is actually calling your temporary variable "artist" from your for loop above.
# You want to call 'artist['genres']'
for genre in artist['genres']:
# TA-COMMENT: This is actually always true and therefore, it's not doing what you'd want it to do!
if True:
genre_count = genre_count + 1
print(artist['genres'])
else:
print("No genres listed")
# TA-COMMENT: see example
if True:
print("hello")
import requests # TA-COMMENT: No need to import requests everytime -- you just once to do it once within a notebook
response = requests.get('https://api.spotify.com/v1/search?query=Lil+&offset=0&limit=50&type=artist&market=US')
data = response.json()
type(data)
data.keys()
type(data['artists'])
data['artists'].keys()
artists = data['artists']['items']
artists
# TA-COMMENT: Excellent for loop!!
for artist in artists:
print(artist['name'], artist['popularity'])
if len(artist['genres']) > 0:
genres = ", ".join(artist['genres'])
print("Genre list: ", genres)
else:
print("No genres listed")
response = requests.get('https://api.spotify.com/v1/search?query=Lil+&offset=0&limit=50&type=artist&market=US')
data = response.json()
# genres = data['genres']
# genre_count = 0
# for genre in genres:
# print(genres['name'])
# g = genres + 1
# genre_count.append(g)
data.keys()
# to figure out what data['artists'] is, we have a couple of options. My favorite!! Printing!!
print(data['artists']) #its a dictionary so look inside!
data['artists'].keys() #calling out what's inside of the dictionary
print(data['artists']['items']) #calling out a dictionary inside of a dicitonary...and look below its a list '[]'
data['artists']['items'][0] #set the list at zero.
artists = data['artists']['items'] #declare the dictionary and list as a variable to make it easier
for artist in artists:
# print(artist.keys())
print(artist['genres'])
from collections import Counter
artists = data['artists']['items']
genre_list = []
for artist in artists:
# print(artist.keys())
# print(artist['genres'])
genre_list = genre_list + artist['genres']
print(genre_list)
Counter(genre_list) #counter function - it rocks
# TA-COMMENT: Yassss
unique_genres = set(genre_list)
unique_genres
genre_count_dict = {}
for unique_item in unique_genres:
count_variable = 0
for item in genre_list:
if item == unique_item:
count_variable = count_variable + 1
genre_count_dict[unique_item] = count_variable
# TA-COMMENT: Beautiful!
genre_count_dict
```
# 3) Use a for loop to determine who BESIDES Lil Wayne has the highest popularity rating. Is it the same artist who has the largest number of followers?
```
most_popular_name = ""
most_popular_score = 0
for artist in artists:
print("Looking at", artist['name'], "who has a popularity score of", artist['popularity'])
#THE CONDITIONAL - WHAT YOU'RE TESTING
print("Comparing", artist['popularity'], "to", most_popular_score, "of")
if artist['popularity'] > most_popular_score and artist ['name'] != "Lil Wayne":
#THE CHANGE - WHAT YOU'RE KEEPING TRACK OF
most_popular_name = artist['name']
most_popular_score = artist['popularity']
print(most_popular_name, most_popular_score)
# TA-COMMENT: Excellent!
```
# 4) Print a list of Lil's that are more popular than Lil' Kim.
```
target_score = 72
#PART ONE: INITIAL CONDITON
second_best_artists = []
#AGGREGATION PROBLEM - when you're looping through a series of objects and you someitmes you want to add some
#of those objects to a DIFFERENT list
for artists in artists:
print("Looking at", artist['name'], "who has a popularity of", artist['popularity'])
#PART TWO: CONDITONAL - when we want to add someone to our list
if artist['popularity'] == 72:
#PART THREE: THE CHANGE - add artist to our list
second_best_artists.append(artist['name'])
print("OUR SECOND BEST ARTISTS ARE:")
for artist in second_best_artists:
print(artist)
# TA-COMMENT: This code doesn't work as you'd want because your temporary variable name "artists" is the same
# as your list name!
for artists in artists:
#print("Looking at", artist['name'])
if artist['name'] == "Lil' Kim":
print("Found Lil' Kim")
print(artist['popularity'])
else:
pass
#print("Not Lil' Kim")
import requests
response = requests.get('https://api.spotify.com/v1/search?query=Lil+&offset=0&limit=50&type=artist&market=US')
data = response.json()
data.keys()
artist_data = data['artists']['items']
for artist in artist_data:
print(artist['name'], artist['popularity'], artist['genres'])
```
# 5) Pick two of your favorite Lils to fight it out, and use their IDs to print out their top tracks.
```
import requests
response = requests.get('https://api.spotify.com/v1/search?query=Lil+&offset=0&limit=50&type=artist&market=US')
data = response.json()
data.keys()
artist_data = data['artists']['items']
for artist in artist_data:
print(artist['name'], "ID is:", artist['id'])
import requests
response = requests.get('https://api.spotify.com/v1/artists/55Aa2cqylxrFIXC767Z865/top-tracks?country=US')
lil_wayne = response.json()
print(lil_wayne)
type(lil_wayne)
print(type(lil_wayne))
print(lil_wayne.keys())
print(lil_wayne['tracks'])
# TA-COMMENT: (-1) Remember what our for loop structures should look like! Below, there is nothing indented below your
# for loop!
# and you cannot call ['tracks'] without specifying which dictionary to find ['tracks']
for lil_wayne in ['tracks']
print("Lil Wayne's top tracks are:")
import requests
response = requests.get('https://api.spotify.com/v1/artists/7sfl4Xt5KmfyDs2T3SVSMK/top-tracks?country=US')
lil_jon = response.json()
print(lil_jon)
type(lil_jon)
print(type(lil_jon))
print(lil_jon.keys())
print(lil_jon['tracks'])
tracks = ['tracks']
for lil_jon in tracks:
print("Lil Jon's top tracks are:")
```
# 6) Will the world explode if a musicians swears? Get an average popularity for their explicit songs vs. their non-explicit songs. How many minutes of explicit songs do they have? Non-explicit?
```
response = requests.get("https://api.spotify.com/v1/artists/55Aa2cqylxrFIXC767Z865/top-tracks?country=US")
data = response.json()
tracks = data['tracks']
explicit_count = 0
clean_count = 0
for track in tracks:
print(track['name'], track['explicit'])
# TA-COMMENT: (-0.5) if True is always True! This happens to work out because all the tracks were explicit.
if True:
explicit_count = explicit_count + 1
if not track['explicit']:
clean_count = clean_count + 1
print("We have found", explicit_count, "explicit tracks, and", clean_count, "tracks are clean")
if track_count > 0:
print("Overall, We discovered", explicit_count, "explicit tracks")
print("And", clean_count, "were non-explicit")
print("Which means", 100 * clean_count / explicit_count, " percent of tracks were clean")
else:
print("No top tracks found")
# TA-COMMENT: It's a good idea to comment out code so if you read it back later, you know what's happening at each stage
import requests
response = requests.get('https://api.spotify.com/v1/artists/55Aa2cqylxrFIXC767Z865/top-tracks?country=US')
data = response.json()
tracks = data['tracks']
#print(tracks)
explicit_count = 0
clean_count = 0
popularity_explicit = 0
popularity_nonexplicit = 0
minutes_explicit = 0
minutes_not_explicit = 0
for track in tracks:
print(track['name'], track['explicit'])
if track['explicit'] == True:
explicit_count = explicit_count + 1
popularity_explicit = popularity_explicit + track['popularity']
minutes_explicit = minutes_explicit + track['duration_ms']
print("The number of explicit songs are", explicit_count, "with a popularity of", popularity_explicit)
print( "Lil Wayne has", minutes_explicit/10000, "minutes of explicit songs" )
elif track['explicit'] == False:
clean_count = clean_count + 1
popularity_nonexplicit = popularity_nonexplicit + track['popularity']
minutes_not_explicit = minutes_not_explicit + track['duration_ms']
print("Lil Wayne has", explicit_count, "explicit tracks, and", clean_count, "clean tracks")
print( "Lil Wayne has", minutes_not_explicit/10000, "minutes of clean songs")
print("The average popularity of Lil Wayne's explicit tracks is", popularity_explicit/explicit_count)
import requests
response = requests.get('https://api.spotify.com/v1/artists/7sfl4Xt5KmfyDs2T3SVSMK/top-tracks?country=US')
data = response.json()
tracks = data['tracks']
#print(tracks)
explicit_count = 0
clean_count = 0
for track in tracks:
print(track['name'], track['explicit'])
if True:
explicit_count = explicit_count + 1
if not track['explicit']:
clean_count = clean_count + 1
print("Lil Jon has", explicit_count, "explicit tracks, and", clean_count, "clean tracks")
response = requests.get("https://api.spotify.com/v1/artists/7sfl4Xt5KmfyDs2T3SVSMK/top-tracks?country=US")
data = response.json()
tracks = data['tracks']
explicit_count = 0
clean_count = 0
for track in tracks:
print(track['name'], track['explicit'])
if True:
explicit_count = explicit_count + 1
if not track['explicit']:
clean_count = clean_count + 1
print("We have found", explicit_count, "explicit tracks, and", clean_count, "tracks are clean")
if track_count > 0:
print("Overall, We discovered", explicit_count, "explicit tracks")
print("And", clean_count, "were non-explicit")
print("Which means", 100 * clean_count / explicit_count, " percent of tracks were clean")
else:
print("No top tracks found")
import requests
response = requests.get('https://api.spotify.com/v1/artists/7sfl4Xt5KmfyDs2T3SVSMK/top-tracks?country=US')
data = response.json()
tracks = data['tracks']
#print(tracks)
explicit_count = 0
clean_count = 0
popularity_explicit = 0
popularity_nonexplicit = 0
minutes_explicit = 0
minutes_not_explicit = 0
for track in tracks:
print(track['name'], track['explicit'])
if track['explicit'] == True:
explicit_count = explicit_count + 1
popularity_explicit = popularity_explicit + track['popularity']
minutes_explicit = minutes_explicit + track['duration_ms']
print("The number of explicit songs are", explicit_count, "with a popularity of", popularity_explicit)
print( "Lil Jon has", minutes_explicit/1000, "minutes of explicit songs" )
elif track['explicit'] == False:
clean_count = clean_count + 1
popularity_nonexplicit = popularity_nonexplicit + track['popularity']
minutes_not_explicit = minutes_not_explicit + track['duration_ms']
print("Lil Jon has", explicit_count, "explicit tracks, and", clean_count, "clean tracks")
print( "Lil Jon has", minutes_not_explicit/1000, "minutes of clean songs")
print("The average popularity of Lil Jon's explicit tracks is", popularity_explicit/explicit_count)
```
| github_jupyter |
# Object-Oriented Python
During this session, we will be exploring the Oriented-Object paradigm in Python using all what we did with Pandas in previous sessions. We will be working with the same data of aircraft supervising latest Tour de France.
```
import pandas as pd
df = pd.read_json("../data/tour_de_france.json.gz")
```
There are three main principles around OOP:
- **encapsulation**: objects embed properties (attributes, methods);
- **interface**: objects expose and document services, they hide all about their inner behaviour;
- **factorisation**: objects/classes with similar behaviour are grouped together.
A common way of working with Python is to implement **protocols**. Protocols are informal interfaces defined by a set of methods allowing an object to play a particular role in the system. For instance, for an object to behave as an iterable you don't need to subclass an abstract class Iterable or implement explicitely an interface Iterable: it is enough to implement the special methods `__iter__` method or even just the `__getitem__` (we will go through these concepts hereunder).
Let's have a look at the special method `sorted`: it expects an **iterable** structure of **comparable** objects to return a sorted list of these objects. Let's have a look:
```
sorted([-2, 4, 0])
```
However it fails when object are not comparable:
```
sorted([-1, 1+1j, 1-2j])
```
Then we can write our own ComparableComplex class and implement a comparison based on modules. The **comparable** protocol expects the `<` operator to be defined (special keyword: `__lt__`)
```
class ComparableComplex(complex):
def __lt__(a, b):
return abs(a) < abs(b)
# Now this works: note the input is not a list but a generator.
sorted(ComparableComplex(i) for i in [-1, 1 + 1j, 1 - 2j])
```
We will be working with different views of pandas DataFrame for trajectories and collection of trajectories. Before we start any further, let's remember two ways to factorise behaviours in Object-Oriented Programming: **inheritance** and **composition**.
The best way to do is not always obvious and it often takes experience to find the good and bad sides of both paradigms.
In our previous examples, our ComparableComplex *offered not much more* than complex numbers. As long as we don't need to compare them, we could have *put them in a list together* with regular complex numbers *without loss of generality*: after all a ComparableComplex **is** a complex. That's a good smell for **inheritance**.
If we think about our trajectories, we will build them around pandas DataFrames. Trajectories will probably have a single attribute: the dataframe. It could be tempting to inherit from `pd.DataFrame`; it will probably work fine in the beginning but problems will occur sooner than expected (most likely with inconsistent interfaces). We **model** trajectories and collections of trajectories with dataframes, but a trajectory **is not** a dataframe. Be reasonable and go for **composition**.
So now we can start.
- The `__init__` special method defines a constructor. `self` is necessary: it represents the current object.
Note that **the constructor does not return anything**.
```
class FlightCollection:
def __init__(self, data):
self.data = data
class Flight:
def __init__(self, data):
self.data = data
FlightCollection(df)
```
## Special methods
There is nothing much we did at this point: just two classes holding a dataframe as an attribute. Even the output representation is the default one based on the class name and the object's address in memory.
- we can **override** the special `__repr__` method (which **returns** a string—**do NOT** `print`!) in order to display a more relevant output. You may use the number of lines in the underlying dataframe for instance.
<div class='alert alert-warning'>
<b>Exercice:</b> Write a relevant <code>__repr__</code> method.
</div>
```
# %load ../solutions/pandas_oo/flight_repr.py
"{0!r}".format(FlightCollection(df))
```
Note that we passed the dataframe in the constructor. We want to keep it that way (we will see later why). However we may want to create a different type of constructor to read directly from the JSON file. There is a special kind of keyword for that.
- `@classmethod` is a decorator to put before a method. It makes it an **class method**, i.e. you call it on the class and not on the object. The first parameter is no longer `self` (the instance) but by convention `cls` (the class).
<div class='alert alert-warning'>
<b>Exercice:</b> Write a relevant <code>read_json</code> class method.
</div>
```
# %load ../solutions/pandas_oo/flight_json.py
collection = FlightCollection.read_json("../data/tour_de_france.json.gz")
```
Now we want to make this `FlightCollection` iterable.
- The special method to implement is `__iter__`. This method takes no argument and **yields** elements one after the other.
<div class='alert alert-warning'>
<b>Exercice:</b> Write a relevant <code>__iter__</code> method which yields Flight instances.
</div>
Of course, you should reuse the code of last session about iteration.
```
# %load ../solutions/pandas_oo/flight_iter.py
collection = FlightCollection.read_json("../data/tour_de_france.json.gz")
for flight in collection:
print(flight)
```
<div class='alert alert-warning'>
<b>Exercice:</b> Write a relevant <code>__repr__</code> method for Flight including callsign, aircraft icao24 code and day of the flight.
</div>
```
# %load ../solutions/pandas_oo/flight_nice_repr.py
for flight in collection:
print(flight)
```
<div class='alert alert-success'>
<b>Note:</b> Since our FlightCollection is iterable, we can pass it to any method accepting iterable structures.
</div>
```
list(collection)
```
<div class='alert alert-warning'>
<b>Warning:</b> However, it won't work here, because Flight instances cannot be compared, unless we specify on which criterion we want to compare.
</div>
```
sorted(collection)
sorted(collection, key=lambda x: x.min("timestamp"))
```
<div class='alert alert-warning'>
<b>Exercice:</b> Implement the proper missing method so that a FlightCollection can be sorted.
</div>
```
# %load ../solutions/pandas_oo/flight_sort.py
sorted(collection)
```
## Data visualisation
See the following snippet of code for plotting trajectories on a map.
```
import matplotlib.pyplot as plt
from cartopy.crs import EuroPP, PlateCarree
fig, ax = plt.subplots(figsize=(10, 10), subplot_kw=dict(projection=EuroPP()))
ax.coastlines("50m")
for flight in collection:
flight.data.plot(
ax=ax,
x="longitude",
y="latitude",
legend=False,
transform=PlateCarree(),
color="steelblue",
)
ax.set_extent((-5, 10, 42, 52))
ax.set_yticks([])
```
<div class='alert alert-warning'>
<b>Exercice:</b> Implement a plot method to make the job even more simple.
</div>
```
# %load ../solutions/pandas_oo/flight_plot.py
fig, ax = plt.subplots(figsize=(10, 10), subplot_kw=dict(projection=EuroPP()))
ax.coastlines("50m")
for flight in collection:
flight.plot(ax, color="steelblue")
ax.set_extent((-5, 10, 42, 52))
ax.set_yticks([])
```
## Indexation
Until now, we implemented all what is necessary to iterate on structures.
This means we have all we need to yield elements one after the other.
Note that:
- Python does not assume your structure has a length.
(There are some infinite iterators, like the one yielding natural integers one after the other.)
- Python cannot guess for you how you want to index your flights.
```
len(collection)
collection['ASR172B']
```
There are many ways to proceed with indexing. We may want to select flights with a specific callsign, or a specific icao24 code. Also, if only one Flight is returned, we want a Flight object. If two or more segments are contained in the underlying dataframe, we want to stick to a FlightCollection.
<div class="alert alert-warning">
<b>Exercice:</b> Implement a <code>__len__</code> special method, then a <code>__getitem__</code> special method that will return a Flight or a FlightCollection (depending on the selection) wrapping data corresponding to the given callsign or icao24 code.
</div>
```
# %load ../solutions/pandas_oo/flight_index.py
collection = FlightCollection.read_json("../data/tour_de_france.json.gz")
collection
collection["3924a0"]
collection["ASR172B"]
from collections import defaultdict
count = defaultdict(int)
for flight in collection["ASR172B"]:
count[flight.icao24] += 1
count
```
As we can see here, this method for indexing is not convenient enough. We could select the only flight `collection["ASR172B"]["3924a0"]` but with current implementation, there is no way to separate the 18 other flights.
<div class='alert alert-warning'>
<b>Exercice:</b> Implement a different <code>__getitem__</code> method that checks the type of the index: filter on callsign/icao24 if the key is a <code>str</code>, filter on the day of the flight if the key is a <code>pd.Timestamp</code>.
</div>
```
# %load ../solutions/pandas_oo/flight_index_time.py
collection = FlightCollection.read_json("../data/tour_de_france.json.gz")
collection["ASR172B"][pd.Timestamp("2019-07-18")]
```
<div class='alert alert-warning'>
<b>Exercice:</b> Plot all trajectories flying on July 18th. How can they be sure to not collide with each other?
</div>
```
# %load ../solutions/pandas_oo/flight_plot_july18.py
```
| github_jupyter |
```
import re
import pandas as pd
import seaborn as sns
from jcopml.plot import plot_missing_value
import matplotlib.pyplot as plt
pd.options.display.max_columns = 100
```
# Import Data
```
df = pd.read_csv(
"./data/investments_VC.csv",
encoding= "ISO-8859-1",
parse_dates= ["founded_at", "first_funding_at", "last_funding_at"],
)
df.head()
```
# EDA Basic Info
```
df.info(verbose=True, memory_usage="deep")
```
# Missing Value Info
```
plot_missing_value(df)
(df.isna().sum() / len(df)) * 100
```
# Fixed Weird Rows / Cols name
```
def convert_str2float(fund: str) -> float:
try:
return float("".join(re.findall("\d+", fund)))
except:
return np.nan
df["market"] = df[" market "]
df["funding_total_usd"] = df[" funding_total_usd "]
df["funding_total_usd"] = df["funding_total_usd"].apply(lambda x: convert_str2float(x))
df.dropna(subset= [" funding_total_usd "], inplace=True)
df.drop(columns=[" market ", " funding_total_usd "], inplace=True)
```
# Popular Market & Country
```
# Prepare Data
df_raw = df.copy()
df_raw = df_raw[df_raw["market"].isin(dict(df_raw.market.value_counts()[:11]).keys())]
df_raw = df_raw[['funding_total_usd', 'market']].groupby('market').apply(lambda x: x.mean())
df_raw.sort_values('funding_total_usd', inplace=True)
df_raw.reset_index(inplace=True)
min_fund = df_raw.funding_total_usd.min()
max_fund = df_raw.funding_total_usd.max()
# # Draw plot
fig, ax = plt.subplots(figsize=(10,8), dpi= 80)
ax.hlines(y=df_raw.index, xmin=min_fund - 1000000, xmax=max_fund + 1500000, color='gray', alpha=0.7, linewidth=1, linestyles='dashdot')
ax.scatter(
y=df_raw.index,
x=df_raw.funding_total_usd,
s=75,
color='firebrick',
alpha=0.7
)
# # Title, Label, Ticks and Ylim
ax.set_title('Average Funding per Market Start Up', fontdict={'size':22})
ax.set_xlabel('USD')
ax.set_yticks(df_raw.index)
ax.set_yticklabels(df_raw.market.str.title(), fontdict={'horizontalalignment': 'right'})
ax.set_xlim(min_fund - 2000000, max_fund + 2500000);
market_count = df['market'].value_counts()
market_count = market_count[:10,]
market_plot = sns.barplot(
market_count.index,
market_count.values,
alpha=0.8,
)
for item in market_plot.get_xticklabels():
item.set_rotation(-90)
plt.title('Top 10 Market in Start Up Investments')
plt.ylabel('Total Companies', fontsize=12)
plt.xlabel('Market Type', fontsize=12)
plt.show()
country_count = df['country_code'].value_counts()
country_count = country_count[:10,]
country_plot = sns.barplot(
country_count.index,
country_count.values,
alpha=0.8,
)
plt.title('Top 10 Countries in Start Up Investments')
plt.ylabel('Total Companies', fontsize=12)
plt.xlabel('Countries', fontsize=12)
plt.show()
status_count = df['status'].value_counts()
status_count = status_count[:10,]
status_plot = sns.barplot(
status_count.index,
status_count.values,
alpha=0.8,
)
plt.title('Status in Start Up Investments')
plt.ylabel('Total Companies', fontsize=12)
plt.xlabel('Status', fontsize=12)
plt.show()
backup = df.copy()
df_raw = backup.copy()
df_raw.founded_year.max()
df_raw = df.copy()
top_categories = list(dict(df_raw.market.value_counts()[:5]).keys())
df_raw = df_raw[df_raw["market"].isin(top_categories)]
def get_founded_year_cat(year):
if year < 2000:
return "1900s"
elif year < 2010:
return "2000s"
elif year >= 2010:
return "2010s"
df_raw["founded_year_cat"] = df_raw.founded_year.apply(lambda x: get_founded_year_cat(x))
df_raw.dropna(subset=["founded_year_cat"], inplace=True)
df_raw = df_raw[['founded_year_cat', "market"]].groupby(['market', 'founded_year_cat']).size()
df_raw = df_raw.unstack()
df_raw
a
import matplotlib.lines as mlines
# Import Data
klass1 = ['red' if (y1-y2) < 0 else 'green' for y1, y2 in zip(df_raw['1900s'], df_raw['2000s'])]
klass2 = ['red' if (y1-y2) < 0 else 'green' for y1, y2 in zip(df_raw['2000s'], df_raw['2010s'])]
# draw line
# https://stackoverflow.com/questions/36470343/how-to-draw-a-line-with-matplotlib/36479941
def newline(p1, p2, color='black'):
ax = plt.gca()
l = mlines.Line2D([p1[0],p2[0]], [p1[1],p2[1]], color='red' if p1[1]-p2[1] > 0 else 'green', marker='o', markersize=6)
ax.add_line(l)
return l
fig, ax = plt.subplots(1,1,figsize=(14,14), dpi= 80)
# Vertical Lines
ax.vlines(x=1, ymin=50, ymax=1900, color='black', alpha=0.7, linewidth=1, linestyles='dotted')
ax.vlines(x=3, ymin=50, ymax=1900, color='black', alpha=0.7, linewidth=1, linestyles='dotted')
ax.vlines(x=5, ymin=50, ymax=1900, color='black', alpha=0.7, linewidth=1, linestyles='dotted')
# # Points
ax.scatter(y=df_raw['1900s'], x=np.repeat(1, df_raw.shape[0]), s=10, color='black', alpha=0.7)
ax.scatter(y=df_raw['2000s'], x=np.repeat(3, df_raw.shape[0]), s=10, color='black', alpha=0.7)
ax.scatter(y=df_raw['2010s'], x=np.repeat(5, df_raw.shape[0]), s=10, color='black', alpha=0.7)
# # Line Segmentsand Annotation
for p1, p2, p3, c in zip(df_raw['1900s'], df_raw['2000s'], df_raw['2010s'], df_raw.index):
newline([1, p1], [3, p2])
newline([3, p2], [5, p3])
ax.text(1+0.05, p1, c , horizontalalignment='right', verticalalignment='center', fontdict={'size':14})
ax.text(3+0.05, p2, c , horizontalalignment='right', verticalalignment='center', fontdict={'size':14})
ax.text(5+0.05, p3, c , horizontalalignment='left', verticalalignment='center', fontdict={'size':14})
# # 'Before' and 'After' Annotations
ax.text(1+0.05, 1900, '1900s', horizontalalignment='left', verticalalignment='center', fontdict={'size':18, 'weight':700})
ax.text(3+0.05, 1900, '2000s', horizontalalignment='left', verticalalignment='center', fontdict={'size':18, 'weight':700})
ax.text(5+0.05, 1900, '2010s', horizontalalignment='left', verticalalignment='center', fontdict={'size':18, 'weight':700})
# # Decoration
ax.set_title("Comparing Total Investment Start Up between 1900s, 2000s, and 2010s", fontdict={'size':22})
ax.set(xlim=(0, 6), ylim=(0, 2000), ylabel='Total Companies')
ax.set_ylabel('Total Companies', fontdict={'size':22})
ax.set_xticks([1, 3, 5])
ax.set_xticklabels(["1900s", "2000s", "2010s"])
plt.yticks(np.arange(0, 2000, 200), fontsize=12)
# # Lighten borders
plt.gca().spines["top"].set_alpha(.0)
plt.gca().spines["bottom"].set_alpha(.0)
plt.gca().spines["right"].set_alpha(.0)
plt.gca().spines["left"].set_alpha(.0)
```
| github_jupyter |
# SEC405: Scalable, Automated Anomaly Detection with Amazon GuardDuty and SageMaker
## Using IP Insights to score security findings
-------
[Return to the workshop repository](https://github.com/aws-samples/aws-security-workshops/edit/master/detection-ml-wksp/)
Amazon SageMaker IP Insights is an unsupervised anomaly detection algorithm for susicipous IP addresses that uses statistical modeling and neural networks to capture associations between online resources (such as account IDs or hostnames) and IPv4 addresses. Under the hood, it learns vector representations for online resources and IP addresses. This essentially means that if the vector representing an IP address and an online resource are close together, then it is likey for that IP address to access that online resource, even if it has never accessed it before.
In this notebook, we use the Amazon SageMaker IP Insights algorithm to train a model using the `<principal ID, IP address`> tuples we generated from the CloudTrail log data, and then use the model to perform inference on the same type of tuples generated from GuardDuty findings to determine how unusual it is to see a particular IP address for a given principal involved with a finding.
After running this notebook, you should be able to:
- obtain, transform, and store data for use in Amazon SageMaker,
- create an AWS SageMaker training job to produce an IP Insights model,
- use the model to perform inference with an Amazon SageMaker endpoint.
If you would like to know more, please check out the [SageMaker IP Inisghts Documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/ip-insights.html).
## Setup
------
*This notebook was created and tested on a ml.m4.xlarge notebook instance. We recommend using the same, but other instance types should still work.*
The following is a cell that contains Python code and can be executed by clicking the button above labelled "Run". When a cell is running, you will see a star in the parentheses to the left (e.g., `In [*]`), and when it has completed you will see a number in the parentheses. Each click of "Run" will execute the next cell in the notebook.
Go ahead and click **Run** now. You should see the text in the `print` statement get printed just beneath the cell.
All of these cells share the same interpreter, so if a cell imports modules, like this one does, those modules will be available to every subsequent cell.
```
import boto3
import botocore
import os
import sagemaker
print("Welcome to IP Insights!")
```
### Configure Amazon S3 Bucket
Before going further, we to specify the S3 bucket that SageMaker will use for input and output data for the model, which will be the bucket where our training and inference tuples from CloudTrail logs and GuardDuty findings, respectively, are located. Edit the following cell to specify the name of the bucket and then run it; you do not need to change the prefix.
```
# Specify the full name of your "sec405-tuplesbucket" here
bucket = 'sec405-tuplesbucket-########'
prefix = ''
```
Finally, run the next cell to complete the setup.
```
execution_role = sagemaker.get_execution_role()
# Check if the bucket exists
try:
boto3.Session().client('s3').head_bucket(Bucket=bucket)
except botocore.exceptions.ParamValidationError as e:
print('Hey! You either forgot to specify your S3 bucket'
' or you gave your bucket an invalid name!')
except botocore.exceptions.ClientError as e:
if e.response['Error']['Code'] == '403':
print("Hey! You don't have permission to access the bucket, {}.".format(bucket))
elif e.response['Error']['Code'] == '404':
print("Hey! Your bucket, {}, doesn't exist!".format(bucket))
else:
raise
else:
print('Training input/output will be stored in: s3://{}/{}'.format(bucket, prefix))
```
## Training
Execute the two cells below to start training. Training should take several minutes to complete. You can look at various training metrics in the log as the model trains. These logs are also available in CloudWatch.
```
from sagemaker.amazon.amazon_estimator import get_image_uri
image = get_image_uri(boto3.Session().region_name, 'ipinsights')
# Configure SageMaker IP Insights input channels
train_key = os.path.join(prefix, 'train', 'cloudtrail_tuples.csv')
s3_train_data = 's3://{}/{}'.format(bucket, train_key)
input_data = {
'train': sagemaker.session.s3_input(s3_train_data, distribution='FullyReplicated', content_type='text/csv')
}
# Set up the estimator with training job configuration
ip_insights = sagemaker.estimator.Estimator(
image,
execution_role,
train_instance_count=1,
train_instance_type='ml.m4.xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
sagemaker_session=sagemaker.Session())
# Configure algorithm-specific hyperparameters
ip_insights.set_hyperparameters(
num_entity_vectors='20000',
random_negative_sampling_rate='5',
vector_dim='128',
mini_batch_size='1000',
epochs='5',
learning_rate='0.01',
)
# Start the training job (should take 3-4 minutes to complete)
ip_insights.fit(input_data)
print('Training job name: {}'.format(ip_insights.latest_training_job.job_name))
```
## Deployment
Execute the cell below to deploy the trained model on an endpoint for inference. It should take 5-7 minutes to spin up the instance and deploy the model (the horizontal dashed line represents progress, and it will print an exclamation point \[!\] when it is complete).
```
predictor = ip_insights.deploy(
initial_instance_count=1,
instance_type='ml.m4.xlarge'
)
print('Endpoint name: {}'.format(predictor.endpoint))
```
## Inference
We can pass data in a variety of formats to our inference endpoint. In this example, we will pass CSV-formmated data.
```
from sagemaker.predictor import csv_serializer, json_deserializer
predictor.content_type = 'text/csv'
predictor.serializer = csv_serializer
predictor.accept = 'application/json'
predictor.deserializer = json_deserializer
```
When queried by a principal and an IPAddress, the model returns a score (called 'dot_product') which indicates how expected that event is. In other words, the higher the dot_product, the more normal the event is. First let's run inference on the training (normal) data for sanity check.
```
import pandas as pd
# Run inference on training (normal) data for sanity check
s3_infer_data = 's3://{}/{}'.format(bucket, train_key)
inference_data = pd.read_csv(s3_infer_data, header=None)
inference_data.head()
predictor.predict(inference_data.values)
```
Now let's run inference on the GuardDuty findings. Notice that the scores are much lower than the normal scores.
```
# Run inference on GuardDuty findings
infer_key = os.path.join(prefix, 'infer', 'guardduty_tuples.csv')
s3_infer_data = 's3://{}/{}'.format(bucket, infer_key)
inference_data = pd.read_csv(s3_infer_data, header=None)
inference_data.head()
predictor.predict(inference_data.values)
```
## Clean-up
To clean up resources created during the workshop, please see the [Cleaning up](https://github.com/aws-samples/aws-security-workshops/blob/cff322dab7cc0b9d71c4f1575c7016389b9dbe64/detection-ml-wksp/README.md) section in the workshop README guide.
| github_jupyter |
**Source of the materials**: Biopython cookbook (adapted)
<font color='red'>Status: Draft</font>
Swiss-Prot and ExPASy {#chapter:swiss_prot}
=====================
Parsing Swiss-Prot files
------------------------
Swiss-Prot (<http://www.expasy.org/sprot>) is a hand-curated database of
protein sequences. Biopython can parse the “plain text” Swiss-Prot file
format, which is still used for the UniProt Knowledgebase which combined
Swiss-Prot, TrEMBL and PIR-PSD. We do not (yet) support the UniProtKB
XML file format.
### Parsing Swiss-Prot records
In Section \[sec:SeqIO\_ExPASy\_and\_SwissProt\], we described how to
extract the sequence of a Swiss-Prot record as a `SeqRecord` object.
Alternatively, you can store the Swiss-Prot record in a
`Bio.SwissProt.Record` object, which in fact stores the complete
information contained in the Swiss-Prot record. In this section, we
describe how to extract `Bio.SwissProt.Record` objects from a Swiss-Prot
file.
To parse a Swiss-Prot record, we first get a handle to a Swiss-Prot
record. There are several ways to do so, depending on where and how the
Swiss-Prot record is stored:
- Open a Swiss-Prot file locally:
`>>> handle = open("myswissprotfile.dat")`
- Open a gzipped Swiss-Prot file:
```
import gzip
handle = gzip.open("myswissprotfile.dat.gz")
```
- Open a Swiss-Prot file over the internet:
```
import urllib.request
handle = urllib.request.urlopen("http://www.somelocation.org/data/someswissprotfile.dat")
```
- Open a Swiss-Prot file over the internet from the ExPASy database
(see section \[subsec:expasy\_swissprot\]):
```
from Bio import ExPASy
handle = ExPASy.get_sprot_raw(myaccessionnumber)
```
The key point is that for the parser, it doesn’t matter how the handle
was created, as long as it points to data in the Swiss-Prot format.
We can use `Bio.SeqIO` as described in
Section \[sec:SeqIO\_ExPASy\_and\_SwissProt\] to get file format
agnostic `SeqRecord` objects. Alternatively, we can use `Bio.SwissProt`
get `Bio.SwissProt.Record` objects, which are a much closer match to the
underlying file format.
To read one Swiss-Prot record from the handle, we use the function
`read()`:
```
from Bio import SwissProt
record = SwissProt.read(handle)
```
This function should be used if the handle points to exactly one
Swiss-Prot record. It raises a `ValueError` if no Swiss-Prot record was
found, and also if more than one record was found.
We can now print out some information about this record:
```
print(record.description)
for ref in record.references:
print("authors:", ref.authors)
print("title:", ref.title)
print(record.organism_classification)
```
To parse a file that contains more than one Swiss-Prot record, we use
the `parse` function instead. This function allows us to iterate over
the records in the file.
For example, let’s parse the full Swiss-Prot database and collect all
the descriptions. You can download this from the [ExPAYs FTP
site](ftp://ftp.expasy.org/databases/uniprot/current_release/knowledgebase/complete/uniprot_sprot.dat.gz)
as a single gzipped-file `uniprot_sprot.dat.gz` (about 300MB). This is a
compressed file containing a single file, `uniprot_sprot.dat` (over
1.5GB).
As described at the start of this section, you can use the Python
library `gzip` to open and uncompress a `.gz` file, like this:
```
import gzip
handle = gzip.open("data/uniprot_sprot.dat.gz")
```
However, uncompressing a large file takes time, and each time you open
the file for reading in this way, it has to be decompressed on the fly.
So, if you can spare the disk space you’ll save time in the long run if
you first decompress the file to disk, to get the `uniprot_sprot.dat`
file inside. Then you can open the file for reading as usual:
```
handle = open("data/uniprot_sprot.dat")
```
As of June 2009, the full Swiss-Prot database downloaded from ExPASy
contained 468851 Swiss-Prot records. One concise way to build up a list
of the record descriptions is with a list comprehension:
```
from Bio import SwissProt
handle = open("data/uniprot_sprot.dat")
descriptions = [record.description for record in SwissProt.parse(handle)]
len(descriptions)
descriptions[:5]
```
Or, using a for loop over the record iterator:
```
from Bio import SwissProt
descriptions = []
handle = open("data/uniprot_sprot.dat")
for record in SwissProt.parse(handle):
descriptions.append(record.description)
len(descriptions)
```
Because this is such a large input file, either way takes about eleven
minutes on my new desktop computer (using the uncompressed
`uniprot_sprot.dat` file as input).
It is equally easy to extract any kind of information you’d like from
Swiss-Prot records. To see the members of a Swiss-Prot record, use
```
dir(record)
```
### Parsing the Swiss-Prot keyword and category list
Swiss-Prot also distributes a file `keywlist.txt`, which lists the
keywords and categories used in Swiss-Prot. The file contains entries in
the following form:
```
ID 2Fe-2S.
AC KW-0001
DE Protein which contains at least one 2Fe-2S iron-sulfur cluster: 2 iron
DE atoms complexed to 2 inorganic sulfides and 4 sulfur atoms of
DE cysteines from the protein.
SY Fe2S2; [2Fe-2S] cluster; [Fe2S2] cluster; Fe2/S2 (inorganic) cluster;
SY Di-mu-sulfido-diiron; 2 iron, 2 sulfur cluster binding.
GO GO:0051537; 2 iron, 2 sulfur cluster binding
HI Ligand: Iron; Iron-sulfur; 2Fe-2S.
HI Ligand: Metal-binding; 2Fe-2S.
CA Ligand.
//
ID 3D-structure.
AC KW-0002
DE Protein, or part of a protein, whose three-dimensional structure has
DE been resolved experimentally (for example by X-ray crystallography or
DE NMR spectroscopy) and whose coordinates are available in the PDB
DE database. Can also be used for theoretical models.
HI Technical term: 3D-structure.
CA Technical term.
//
ID 3Fe-4S.
...
```
The entries in this file can be parsed by the `parse` function in the
`Bio.SwissProt.KeyWList` module. Each entry is then stored as a
`Bio.SwissProt.KeyWList.Record`, which is a Python dictionary.
```
from Bio.SwissProt import KeyWList
handle = open("data/keywlist.txt")
records = KeyWList.parse(handle)
for record in records:
print(record['ID'])
print(record['DE'])
```
This prints
```
2Fe-2S.
Protein which contains at least one 2Fe-2S iron-sulfur cluster: 2 iron atoms
complexed to 2 inorganic sulfides and 4 sulfur atoms of cysteines from the
protein.
...
```
Parsing Prosite records
-----------------------
Prosite is a database containing protein domains, protein families,
functional sites, as well as the patterns and profiles to recognize
them. Prosite was developed in parallel with Swiss-Prot. In Biopython, a
Prosite record is represented by the `Bio.ExPASy.Prosite.Record` class,
whose members correspond to the different fields in a Prosite record.
In general, a Prosite file can contain more than one Prosite records.
For example, the full set of Prosite records, which can be downloaded as
a single file (`prosite.dat`) from the [ExPASy FTP
site](ftp://ftp.expasy.org/databases/prosite/prosite.dat), contains 2073
records (version 20.24 released on 4 December 2007). To parse such a
file, we again make use of an iterator:
```
from Bio.ExPASy import Prosite
handle = open("myprositefile.dat")
records = Prosite.parse(handle)
```
We can now take the records one at a time and print out some
information. For example, using the file containing the complete Prosite
database, we’d find
```
from Bio.ExPASy import Prosite
handle = open("prosite.dat")
records = Prosite.parse(handle)
record = next(records)
record.accession
record.name
record.pdoc
record = next(records)
record.accession
record.name
record.pdoc
record = next(records)
record.accession
record.name
record.pdoc
```
and so on. If you’re interested in how many Prosite records there are,
you could use
```
from Bio.ExPASy import Prosite
handle = open("prosite.dat")
records = Prosite.parse(handle)
n = 0
for record in records:
n += 1
n
```
To read exactly one Prosite from the handle, you can use the `read`
function:
```
from Bio.ExPASy import Prosite
handle = open("mysingleprositerecord.dat")
record = Prosite.read(handle)
```
This function raises a ValueError if no Prosite record is found, and
also if more than one Prosite record is found.
Parsing Prosite documentation records
-------------------------------------
In the Prosite example above, the `record.pdoc` accession numbers
`'PDOC00001'`, `'PDOC00004'`, `'PDOC00005'` and so on refer to Prosite
documentation. The Prosite documentation records are available from
ExPASy as individual files, and as one file (`prosite.doc`) containing
all Prosite documentation records.
We use the parser in `Bio.ExPASy.Prodoc` to parse Prosite documentation
records. For example, to create a list of all accession numbers of
Prosite documentation record, you can use
```
from Bio.ExPASy import Prodoc
handle = open("prosite.doc")
records = Prodoc.parse(handle)
accessions = [record.accession for record in records]
```
Again a `read()` function is provided to read exactly one Prosite
documentation record from the handle.
Parsing Enzyme records
----------------------
ExPASy’s Enzyme database is a repository of information on enzyme
nomenclature. A typical Enzyme record looks as follows:
```
ID 3.1.1.34
DE Lipoprotein lipase.
AN Clearing factor lipase.
AN Diacylglycerol lipase.
AN Diglyceride lipase.
CA Triacylglycerol + H(2)O = diacylglycerol + a carboxylate.
CC -!- Hydrolyzes triacylglycerols in chylomicrons and very low-density
CC lipoproteins (VLDL).
CC -!- Also hydrolyzes diacylglycerol.
PR PROSITE; PDOC00110;
DR P11151, LIPL_BOVIN ; P11153, LIPL_CAVPO ; P11602, LIPL_CHICK ;
DR P55031, LIPL_FELCA ; P06858, LIPL_HUMAN ; P11152, LIPL_MOUSE ;
DR O46647, LIPL_MUSVI ; P49060, LIPL_PAPAN ; P49923, LIPL_PIG ;
DR Q06000, LIPL_RAT ; Q29524, LIPL_SHEEP ;
//
```
In this example, the first line shows the EC (Enzyme Commission) number
of lipoprotein lipase (second line). Alternative names of lipoprotein
lipase are “clearing factor lipase”, “diacylglycerol lipase”, and
“diglyceride lipase” (lines 3 through 5). The line starting with “CA”
shows the catalytic activity of this enzyme. Comment lines start with
“CC”. The “PR” line shows references to the Prosite Documentation
records, and the “DR” lines show references to Swiss-Prot records. Not
of these entries are necessarily present in an Enzyme record.
In Biopython, an Enzyme record is represented by the
`Bio.ExPASy.Enzyme.Record` class. This record derives from a Python
dictionary and has keys corresponding to the two-letter codes used in
Enzyme files. To read an Enzyme file containing one Enzyme record, use
the `read` function in `Bio.ExPASy.Enzyme`:
```
from Bio.ExPASy import Enzyme
with open("data/lipoprotein.txt") as handle:
record = Enzyme.read(handle)
record["ID"]
record["DE"]
record["AN"]
record["CA"]
record["PR"]
```
```
record["CC"]
record["DR"]
```
The `read` function raises a ValueError if no Enzyme record is found,
and also if more than one Enzyme record is found.
The full set of Enzyme records can be downloaded as a single file
(`enzyme.dat`) from the [ExPASy FTP
site](ftp://ftp.expasy.org/databases/enzyme/enzyme.dat), containing 4877
records (release of 3 March 2009). To parse such a file containing
multiple Enzyme records, use the `parse` function in `Bio.ExPASy.Enzyme`
to obtain an iterator:
```
from Bio.ExPASy import Enzyme
handle = open("enzyme.dat")
records = Enzyme.parse(handle)
```
We can now iterate over the records one at a time. For example, we can
make a list of all EC numbers for which an Enzyme record is available:
```
ecnumbers = [record["ID"] for record in records]
```
Accessing the ExPASy server
---------------------------
Swiss-Prot, Prosite, and Prosite documentation records can be downloaded
from the ExPASy web server at <http://www.expasy.org>. Six kinds of
queries are available from ExPASy:
get\_prodoc\_entry
: To download a Prosite documentation record in HTML format
get\_prosite\_entry
: To download a Prosite record in HTML format
get\_prosite\_raw
: To download a Prosite or Prosite documentation record in raw format
get\_sprot\_raw
: To download a Swiss-Prot record in raw format
sprot\_search\_ful
: To search for a Swiss-Prot record
sprot\_search\_de
: To search for a Swiss-Prot record
To access this web server from a Python script, we use the `Bio.ExPASy`
module.
### Retrieving a Swiss-Prot record {#subsec:expasy_swissprot}
Let’s say we are looking at chalcone synthases for Orchids (see
section \[sec:orchids\] for some justification for looking for
interesting things about orchids). Chalcone synthase is involved in
flavanoid biosynthesis in plants, and flavanoids make lots of cool
things like pigment colors and UV protectants.
If you do a search on Swiss-Prot, you can find three orchid proteins for
Chalcone Synthase, id numbers O23729, O23730, O23731. Now, let’s write a
script which grabs these, and parses out some interesting information.
First, we grab the records, using the `get_sprot_raw()` function of
`Bio.ExPASy`. This function is very nice since you can feed it an id and
get back a handle to a raw text record (no HTML to mess with!). We can
the use `Bio.SwissProt.read` to pull out the Swiss-Prot record, or
`Bio.SeqIO.read` to get a SeqRecord. The following code accomplishes
what I just wrote:
```
from Bio import ExPASy
from Bio import SwissProt
accessions = ["O23729", "O23730", "O23731"]
records = []
for accession in accessions:
handle = ExPASy.get_sprot_raw(accession)
record = SwissProt.read(handle)
records.append(record)
```
If the accession number you provided to `ExPASy.get_sprot_raw` does not
exist, then `SwissProt.read(handle)` will raise a `ValueError`. You can
catch `ValueException` exceptions to detect invalid accession numbers:
```
for accession in accessions:
handle = ExPASy.get_sprot_raw(accession)
try:
record = SwissProt.read(handle)
except ValueException:
print("WARNING: Accession %s not found" % accession)
records.append(record)
```
### Searching Swiss-Prot
Now, you may remark that I knew the records’ accession numbers
beforehand. Indeed, `get_sprot_raw()` needs either the entry name or an
accession number. When you don’t have them handy, you can use one of the
`sprot_search_de()` or `sprot_search_ful()` functions.
`sprot_search_de()` searches in the ID, DE, GN, OS and OG lines;
`sprot_search_ful()` searches in (nearly) all the fields. They are
detailed on <http://www.expasy.org/cgi-bin/sprot-search-de> and
<http://www.expasy.org/cgi-bin/sprot-search-ful> respectively. Note that
they don’t search in TrEMBL by default (argument `trembl`). Note also
that they return HTML pages; however, accession numbers are quite easily
extractable:
```
from Bio import ExPASy
import re
handle = ExPASy.sprot_search_de("Orchid Chalcone Synthase")
# or:
# handle = ExPASy.sprot_search_ful("Orchid and {Chalcone Synthase}")
html_results = handle.read()
if "Number of sequences found" in html_results:
ids = re.findall(r'HREF="/uniprot/(\w+)"', html_results)
else:
ids = re.findall(r'href="/cgi-bin/niceprot\.pl\?(\w+)"', html_results)
```
### Retrieving Prosite and Prosite documentation records
Prosite and Prosite documentation records can be retrieved either in
HTML format, or in raw format. To parse Prosite and Prosite
documentation records with Biopython, you should retrieve the records in
raw format. For other purposes, however, you may be interested in these
records in HTML format.
To retrieve a Prosite or Prosite documentation record in raw format, use
`get_prosite_raw()`. For example, to download a Prosite record and print
it out in raw text format, use
```
from Bio import ExPASy
handle = ExPASy.get_prosite_raw('PS00001')
text = handle.read()
print(text)
```
To retrieve a Prosite record and parse it into a `Bio.Prosite.Record`
object, use
```
from Bio import ExPASy
from Bio import Prosite
handle = ExPASy.get_prosite_raw('PS00001')
record = Prosite.read(handle)
```
The same function can be used to retrieve a Prosite documentation record
and parse it into a `Bio.ExPASy.Prodoc.Record` object:
```
from Bio import ExPASy
from Bio.ExPASy import Prodoc
handle = ExPASy.get_prosite_raw('PDOC00001')
record = Prodoc.read(handle)
```
For non-existing accession numbers, `ExPASy.get_prosite_raw` returns a
handle to an emptry string. When faced with an empty string,
`Prosite.read` and `Prodoc.read` will raise a ValueError. You can catch
these exceptions to detect invalid accession numbers.
The functions `get_prosite_entry()` and `get_prodoc_entry()` are used to
download Prosite and Prosite documentation records in HTML format. To
create a web page showing one Prosite record, you can use
```
from Bio import ExPASy
handle = ExPASy.get_prosite_entry('PS00001')
html = handle.read()
output = open("myprositerecord.html", "w")
output.write(html)
output.close()
```
and similarly for a Prosite documentation record:
```
from Bio import ExPASy
handle = ExPASy.get_prodoc_entry('PDOC00001')
html = handle.read()
output = open("myprodocrecord.html", "w")
output.write(html)
output.close()
```
For these functions, an invalid accession number returns an error
message in HTML format.
Scanning the Prosite database
-----------------------------
[ScanProsite](http://www.expasy.org/tools/scanprosite/) allows you to
scan protein sequences online against the Prosite database by providing
a UniProt or PDB sequence identifier or the sequence itself. For more
information about ScanProsite, please see the [ScanProsite
documentation](http://www.expasy.org/tools/scanprosite/scanprosite-doc.html)
as well as the [documentation for programmatic access of
ScanProsite](http://www.expasy.org/tools/scanprosite/ScanPrositeREST.html).
You can use Biopython’s `Bio.ExPASy.ScanProsite` module to scan the
Prosite database from Python. This module both helps you to access
ScanProsite programmatically, and to parse the results returned by
ScanProsite. To scan for Prosite patterns in the following protein
sequence:
```
MEHKEVVLLLLLFLKSGQGEPLDDYVNTQGASLFSVTKKQLGAGSIEECAAKCEEDEEFT
CRAFQYHSKEQQCVIMAENRKSSIIIRMRDVVLFEKKVYLSECKTGNGKNYRGTMSKTKN
```
you can use the following code:
```
sequence = "MEHKEVVLLLLLFLKSGQGEPLDDYVNTQGASLFSVTKKQLGAGSIEECAAKCEEDEEFT
from Bio.ExPASy import ScanProsite
handle = ScanProsite.scan(seq=sequence)
```
By executing `handle.read()`, you can obtain the search results in raw
XML format. Instead, let’s use `Bio.ExPASy.ScanProsite.read` to parse
the raw XML into a Python object:
```
result = ScanProsite.read(handle)
type(result)
```
A `Bio.ExPASy.ScanProsite.Record` object is derived from a list, with
each element in the list storing one ScanProsite hit. This object also
stores the number of hits, as well as the number of search sequences, as
returned by ScanProsite. This ScanProsite search resulted in six hits:
```
result.n_seq
result.n_match
len(result)
result[0]
result[1]
result[2]
result[3]
result[4]
result[5]
```
Other ScanProsite parameters can be passed as keyword arguments; see the
[documentation for programmatic access of
ScanProsite](http://www.expasy.org/tools/scanprosite/ScanPrositeREST.html)
for more information. As an example, passing `lowscore=1` to include
matches with low level scores lets use find one additional hit:
```
handle = ScanProsite.scan(seq=sequence, lowscore=1)
result = ScanProsite.read(handle)
result.n_match
```
| github_jupyter |
# Hierarchical Risk Parity - usage examples
## Abstract
Hierarchical Risk Parity is a novel portfolio optimisation method developed by Marcos Lopez de Prado. The working of the
algorithm can be broken down into 3 steps:
1. Assets are segregated into clusters via hierarchical tree clustering.
2. Based on these clusters, the covariance matrix of the returns is reorganized such that assets
within the same cluster are regrouped together.
3. Finally, the weights are assigned to each cluster in a recursive manner. At each node, the weights are broken down
into the sub-cluster until all the individual assets are assigned a unique weight.
Although, it is a simple algorithm, HRP has been found to be very stable as compared to its older counterparts.
This is because, HRP does not involve taking inverse of the covariance matrix matrix which makes it robust to small changes
in the covariances of the asset returns.
For a detailed explanation of how HRP works, we have written an excellent `blog post <https://hudsonthames.org/an-introduction-to-the-hierarchical-risk-parity-algorithm/>`_ about it.
The objective of this notebook is to provide examples on how to use HRP in practice
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from mlfinlab.portfolio_optimization.hrp import HierarchicalRiskParity
import warnings
warnings.filterwarnings('ignore')
def plot_weights(weights):
weights = weights.sort_values(by=0, ascending=False, axis=1)
plt.figure(figsize=(10, 5))
plt.bar(weights.columns, weights.values[0])
plt.xlabel('Tickers', fontsize=12)
plt.ylabel('Weights', fontsize=12)
plt.title('Portfolio Weights', fontsize=12)
plt.show()
data = pd.read_csv('../Sample-Data/stock_prices.csv')
data.set_index('Date', inplace=True)
data.index = pd.to_datetime(data.index)
data.head()
```
In order to work, the Algo needs 2 important matrices:
* A covariance matrix for estimating the risk associated with each cluster
* A distance_matrix often derived directly from the covariance allowing to build our clusters
## 1. Basic examples
#### First let's give the algo a dataframe or numpy array of raw prices with the corresponding names
```
hrp = HierarchicalRiskParity()
hrp.allocate(asset_prices=data, asset_names=data.columns)
plot_weights(hrp.weights)
```
By default, the algo is asked to build a minimum variance portfolio being long on each asset
The weight number k is just the proportion of your investment allocated to asset number k.
Providing asset prices means the algo will compute the returns, a covariance matrix and the distance matrix.
#### What if I want to give returns instead of price ?
```
hrp = HierarchicalRiskParity()
returns = data.pct_change()[1:]
hrp.allocate(asset_returns=returns, asset_names=data.columns)
plot_weights(hrp.weights)
```
As expected, we have found the exact same weights.
Providing asset returns means the algo only need to compute a covariance matrix and the distance matrix
#### But I have my own covariance matrix, already shrinked and proven to perform well out of sample, can I use it?
Of course Voilà
```
hrp = HierarchicalRiskParity()
my_fav_cov_matrix = returns.cov()
hrp.allocate(covariance_matrix=my_fav_cov_matrix, asset_names=data.columns)
plot_weights(hrp.weights)
```
Here the algo only has to compute the distance matrix from the covariance matrix
## 2. More advanced examples
#### What if I want to use my prefered linkage method for the tree clustering part?
```
hrp = HierarchicalRiskParity()
my_fav_cov_matrix = returns.cov()
hrp.allocate(covariance_matrix=my_fav_cov_matrix, asset_names=data.columns, linkage_method='ward')
plot_weights(hrp.weights)
```
By default, HRP uses single linkage but any other technique implemented in scipy could do the job.
The user can test various clustering method and see which one is the most robust for his particular task.
https://docs.scipy.org/doc/scipy/reference/generated/scipy.cluster.hierarchy.linkage.html
Note: The weights have changed a little bit compared to the single linkage method used in the Basic example section.
#### How to provide my own distance matrix ?
```
def cov2corr(covariance):
"""
Calculate the correlations from asset returns covariance matrix.
:param covariance: (pd.Dataframe) asset returns covariances
:return: (pd.Dataframe) correlations between asset returns
"""
d_matrix = np.zeros_like(covariance)
diagnoal_sqrt = np.sqrt(np.diag(covariance))
np.fill_diagonal(d_matrix, diagnoal_sqrt)
d_inv = np.linalg.inv(d_matrix)
corr = np.dot(np.dot(d_inv, covariance), d_inv)
corr = pd.DataFrame(corr, index=covariance.columns, columns=covariance.columns)
return corr
hrp = HierarchicalRiskParity()
cov = returns.cov()
corr = cov2corr(cov)
distance = np.sqrt((1 - corr).round(5) / 2)
hrp.allocate(distance_matrix=distance, covariance_matrix=cov, asset_names=data.columns)
plot_weights(hrp.weights)
```
Yes that's exactly the same weights than the ones given by the Basic examples section because that's how HRP computes the distance matrix. Nonetheless, using the same parameter, you can provide your custom distance matrix.
Remember to provide a covariance matrix (or prices, or returns) because the algo needs it to compute the intra cluster variance and then behave accordingly.
#### Is it possible to build a Long/Short portfolio ?
By using the side_weights parameter (pd.Series) with 1 for Buy and -1 for Sell and asset names as index.
Note: The portfolio is dollar neutral 50% is assigned to buys and 50% for sells.
```
hrp = HierarchicalRiskParity()
side_weights = pd.Series([1]*data.shape[1], index=data.columns)
#Short the first 4 stocks
side_weights.loc[data.columns[:4]] = -1
hrp.allocate(asset_prices=data, asset_names=data.columns, side_weights=side_weights)
side_weights
side_weights.value_counts()
plot_weights(hrp.weights)
short_ptf = side_weights[side_weights==-1].index
buy_ptf = side_weights[side_weights==1].index
weights = hrp.weights.T
print('Proportion $ allocated to short portf:', weights.loc[short_ptf].sum().item())
print('Proportion $ allocated to buy portf:', weights.loc[buy_ptf].sum().item())
```
## Conclusion
HRP is a robust algo for minimum variance portfolio construction and for which the user will find a lot of freedom regarding the input parameters.
You can also look at the HRP performance comparison with other portfolio construction techniques on the following notebook: https://github.com/hudson-and-thames/research/blob/master/Chapter16/Chapter16.ipynb
| github_jupyter |
# Predicting Interest Rates for Bank Loans
In this project, I use a dataset of loans from a bank to predict the interest rate an applicant can expect
- first, I wrangle the data by cleansing it of missing values, useless variables, and leaky information
- then, I feature engineer a few columns to hopefully improve my model's performance
- finally, I iterate through various models, optimizing hyperparameters for each, to find the best model
- I obtain a model which can account for 90% of the variance in the data and predict interest rate with a mean
error of 0.9%
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import shap
import seaborn as sns
import time
from pandas_profiling import ProfileReport
from sklearn.pipeline import make_pipeline
from category_encoders import OneHotEncoder
from sklearn.model_selection import train_test_split
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import cross_val_score, validation_curve # k-fold CV
from sklearn.linear_model import LinearRegression, Ridge
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from xgboost import XGBRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import r2_score
```
# Data Wrangling and Feature Engineering
Here, I clean my data into a format that will be a suitable input for modeling. I get rid of columns with high cardinality or a cardinality of 1, excessive NaN values, as well as all the columns containing leaky information. The leaky variables were variables in the dataset available after the loan had been approved - had I not gotten rid of them, my model would have had much greater predictive power (I tested this for myself but running the same analyses without a fully wrangled dataframe - that analysis is not included here) but it would have been a *useless* model since the majority of the predictive power would have come from columns like the 'grade' and 'subgrade' columns describing how likely the bank thought the applicant was to pay back the loan, which is very tightly correlated with the interest rate. The more risky a loan is for the bank, the higher interest they will charge on it.
Although it is not obvious from the code, I found the documentation to this dataset and meticulously examined each column to determine if it should be kept, removed, or changed to a more suitable format.
I engineer a few features, several of which proved to have strong predictive power.
```
# my dataframe is quite large, starting at 150 columns and over 100,000 rows.
# I will do some cleaning and feature engineering, but first I need to see all my columns
pd.set_option('display.max_columns', 150)
pd.set_option('display.max_rows', 50)
df = pd.read_csv('/Users/nicholashagemann/Lambda/Datasets/loan_data.csv')
df.head()
```
Note: The following code block is the final result of much time and computing power spent analyzing the dataframe - I have removed the code it took to create the wrangle function as its length would only serve to clutter my notebook.
```
# each row in the data is a specific loan that was issued by a bank
# I choose the issue date of the loan to be the index
df = pd.read_csv('/Users/nicholashagemann/Lambda/Datasets/loan_data.csv',
parse_dates = ['issue_d', 'earliest_cr_line'],
index_col = 'issue_d').sort_values('issue_d')
# we will need to wrangle this data quite a bit before it's suitable for predicting interest rate
def wrangle(df):
# Begin Data Cleaning
# first I get rid of all columns with more than 20% NaN values
# note that the necessary columns will be dropped at the same time at the end of wrangle()
nan_cols = []
for col in df.columns:
if df[col].isna().sum()/len(df) > 0.2:
nan_cols.append(col)
# each of the following columns had data that was only available after the loan had been approved
# we want to predict the interest rate for a loan before the bank decides to approve the loan
# thus, these columns contain leakage and should be dropped
leaky_cols = ['out_prncp', 'out_prncp_inv', 'total_pymnt', 'total_pymnt_inv', 'sub_grade',
'total_rec_int', 'total_rec_prncp', 'total_rec_late_fee', 'recoveries',
'collection_recovery_fee', 'last_pymnt_d', 'last_pymnt_amnt', 'initial_list_status',
'last_fico_range_high', 'last_fico_range_low', 'collections_12_mths_ex_med',
'funded_amnt', 'funded_amnt_inv', 'debt_settlement_flag', 'grade', 'loan_status']
# the following columns did not contain any useful data or were duplicates of other columns
cols_to_drop = ['title', 'zip_code', 'addr_state', 'fico_range_low',
'Unnamed: 0', 'emp_title', 'url', 'id', 'last_credit_pull_d',
'earliest_cr_line']
# if a column only has one value it has no predictive power
for col in df.columns:
if df[col].nunique() == 1:
cols_to_drop.append(col)
# Begin Feature Engineering
# first I create a feature which identifies how old the borrower's credit is in days
df['credit_age'] = df.index - df['earliest_cr_line']
df['credit_age'].astype('timedelta64[D]').astype(int)
df['credit_age'] = df['credit_age'].dt.days.astype(int)
# we want our interest rate to be a float, not a string
# do the same for the amount of credit the borrower is using
df['int_rate'] = df['int_rate'].str.replace('%', '').astype(float)
df['revol_util'] = df['revol_util'].str.replace('%', '').astype(float)
# we want to represent length of the loan as an integer
df.loc[df['term'] == ' 36 months', 'term'] = 36
df.loc[df['term'] == ' 60 months', 'term'] = 60
df['term'] = df['term'].astype(int)
# we want to represent employment length as an integer
df.loc[df['emp_length'] == '< 1 year', 'emp_length'] = 0
df['emp_length'] = df['emp_length'].str.replace(' ', '').str.replace('year', '').str.replace('s', '').str.replace('+', '')
df['emp_length'] = pd.to_numeric(df['emp_length'], downcast = 'integer', errors = 'ignore')
# we want to represent whether the application was individual or joint as binary
df.loc[df['application_type'] == 'Individual', 'application_type'] = 1
df.loc[df['application_type'] == 'Joint App', 'application_type'] = 0
df['application_type'] = df['application_type'].astype(int)
# drop columns from the data cleaning
df.drop(columns = nan_cols + leaky_cols + cols_to_drop, inplace = True)
return df
df = wrangle(df)
# check how many unique values we have for each of our object columns
# looks like it's small enough to use OneHotEncoder
df.select_dtypes('object').nunique()
# check shape and head of dataframe
print('Shape is: ', df.shape)
df.head()
df.describe()
```
# Creating Training and Validation Data
Here, I create the X (predictor) and y (target) matrices which will then be split into training and testing sets for my models.
```
# we want to have ~80% of our data for training and ~20% for validation
# we see that the 80th percentile is December 2016
# we have a very large dataset, so we choose to start in January 2017
display(df.iloc[0:1], df.iloc[round(df.shape[0]*0.8):round(df.shape[0]*0.8)+1])
# our prediction target is the interest rate, our predictive matrix will be everything except the target
target = 'int_rate'
y = df[target]
X = df.drop(columns = target)
# separate our training and validation data by our cutoff of January 2017
cutoff = '2017-01-01'
mask = X.index < cutoff
X_train, y_train = X[mask], y[mask]
X_val, y_val = X[~mask], y[~mask]
# check shapes to make sure dimensions are correct
print(f'The shape of the training data is {X_train.shape} for X and {y_train.shape} for y.')
print(f'The shape of the validation data is {X_val.shape} for X and {y_val.shape} for y.')
# make sure no data was lost in the splice
assert len(X_train) + len(X_val) == len(X)
# our baseline mean absolute error will be the mean interest rate
# baseline R^2 score will of course be 0 or approximately 0
baseline_reg = [y_train.mean()]*len(y_val)
print('Baseline MAE:', mean_absolute_error(y_val, baseline_reg))
print('Baseline R2:', r2_score(y_val, baseline_reg))
```
# Best Model - XGBoost
For the sake of the reader's time, I include my final model here so that it is not necessary to scroll through the various models and tuning of hyperparameters.
My final model predicts interest rate with a mean error below 0.9% and a cross-validated R^2 of 0.91.
```
%%time
model_final = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy='mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bytree = 1,
reg_alpha = 0,
reg_lambda = 1,
gamma = 0))
model_final.fit(X_train, y_train)
print('Training MAE:', mean_absolute_error(y_train, model_final.predict(X_train)))
print('Validation MAE:', mean_absolute_error(y_val, model_final.predict(X_val)))
print('Validation R2:', model_boost.score(X_val, y_val))
cross_vals = cross_val_score(model_final, X, y)
print(f'Scores: {cross_vals}\nMean: {np.mean(cross_vals)}')
```
# On Optimizing Hyperparameters
For each model, I will optimize whatever hyperparameters are available for that model.
In testing the hyperparameters, I will look at the Mean Absolute Error and R^2 (what percentage of the variance in the data my model accounts for) for the training and validation data, as well as the time it took - the time is necessary since when testing each hyperparameter I do not necessarily want the previously tested hyperparameters to be optimal since the model may take too much time.
By having the MAE, R^2, and Time for each hyperparameter I can know which hyperparameter I will use in my final model while also learning which hyperparameter will be suitable for tuning other hyperparameters - i.e. for my final model I will use the hyperparameter with the lowest MAE and highest R^2, but when tuning the other hyperparameters I will use a value which has a MAE and R^2 *similar* to the best but not as time-expensive.
I choose not to use a GridSearch for the hyperparameters since I am attempting to optimize every hyperparameter and the exponential increase in the time it would take to train my model is not worth the small amount of additional certainty I would gain from knowing I've found the absolute best combination - also, I am sure from past experience with exactly this problem that the increase in accuracy from a GridSearch would be very small compared to the overall increase in accuracy gained through my method (which itself is still small compared to the accuracy of the model itself).
In going through each model, I also choose the order in which I optimize hyperparameters very deliberately. I start with hyperparameters that will have the greatest effect on the accuracy and then move to hyperparameters with smaller effects - in essence, ordering them so that the later hyperparameters do not affect the earlier hyperparameters.
# Decision Tree Regressor
Here, I will test the decision tree model and optimize hyperparameters. It is very unlikely that a decision tree will be the best model, as we naturally expect any random forest to outcompete a single tree.
We find, after optimizing hyperparameters, the MAE is 2.28% and the R^2 is 0.4
The only significant hyperparameter to optimize for a decision tree is the depth of the tree - since we have a large dataset, we can rely on the algorithm and not have to choose the minimum samples for a split or a leaf, the maximum features or leaves, or the minimum decrease in impurity.
We see the depth of the tree significantly affects the accuracy and R^2 but the split criteria does not, and also that it is wise to stick with the default "best" splitter and not the "random" splitter.
```
# do a broad search first then a narrow search next to save time
depths = [1] + [i for i in range(4, 21, 4)]
train_acc_dt = []
val_acc_dt = []
train_r2_dt = []
val_r2_dt = []
times = []
# for each depth, train the model on the training data then add the accuracies, r^2, and time
for depth in depths:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
DecisionTreeRegressor(random_state = 42,
max_depth = depth))
model.fit(X_train, y_train)
train_acc_dt.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_dt.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_dt.append(model.score(X_train, y_train))
val_r2_dt.append(model.score(X_val, y_val))
times.append(time.time() - start)
# display stored values to determine best parameter
pd.DataFrame(list(zip(train_acc_dt, val_acc_dt, train_r2_dt, val_r2_dt, times)), index = depths,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2', 'Time'])
# time for the narrow search
depths = range(8, 17)
train_acc_dt = []
val_acc_dt = []
train_r2_dt = []
val_r2_dt = []
times = []
# for each depth, train the model on the training data then add the accuracies, r^2, and time
for depth in depths:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
DecisionTreeRegressor(random_state = 42,
max_depth = depth))
model.fit(X_train, y_train)
train_acc_dt.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_dt.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_dt.append(model.score(X_train, y_train))
val_r2_dt.append(model.score(X_val, y_val))
times.append(time.time() - start)
# display stored values to determine best parameter
pd.DataFrame(list(zip(train_acc_dt, val_acc_dt, train_r2_dt, val_r2_dt, times)), index = depths,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2', 'Time'])
# Here we decide if we should be using MSE or Friedman_MSE to split the cells in the tree - I ignore MAE
# and reducing poisson deviance because the models take significantly longer to train than the MSE variants
# and in attempting the model, my kernel crashes and must be restarted from the beginning :(
criteria = ["mse", "friedman_mse"]
train_acc_dt = []
val_acc_dt = []
train_r2_dt = []
val_r2_dt = []
for crit in criteria:
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
DecisionTreeRegressor(random_state = 42,
criterion = crit,
max_depth = 12))
model.fit(X_train, y_train)
train_acc_dt.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_dt.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_dt.append(model.score(X_train, y_train))
val_r2_dt.append(model.score(X_val, y_val))
pd.DataFrame(list(zip(train_acc_dt, val_acc_dt, train_r2_dt, val_r2_dt)), index = criteria,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2'])
# was curious how the 'random' splitter would compare to the 'best' - as expected, 'best' is much better
splits = ['best', 'random']
train_acc_dt = []
val_acc_dt = []
train_r2_dt = []
val_r2_dt = []
times = []
for split in splits:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
DecisionTreeRegressor(random_state = 42,
splitter = split,
max_depth = 12))
model.fit(X_train, y_train)
train_acc_dt.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_dt.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_dt.append(model.score(X_train, y_train))
val_r2_dt.append(model.score(X_val, y_val))
times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_dt, val_acc_dt, train_r2_dt, val_r2_dt, times)), index = splits,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2', 'Time'])
# make our decision tree model here
model_dt = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
DecisionTreeRegressor(random_state = 42,
max_depth = 12))
%%time
model_dt.fit(X_train, y_train)
print('Training MAE:', mean_absolute_error(y_train, model_dt.predict(X_train)))
print('Validation MAE:', mean_absolute_error(y_val, model_dt.predict(X_val)))
print('Validation R2:', model_dt.score(X_val,y_val))
# our MAE isn't bad for a weak model, but we can do better. R^2 is very low.
# here I graph the most important features in our model
importances = model_dt.named_steps['decisiontreeregressor'].feature_importances_
features = model_dt.named_steps['onehotencoder'].get_feature_names()
feat_imp_dt = pd.Series(importances, index = features,
name = 'tree').abs().sort_values(ascending=False)
feat_imp_dt.head(10).plot(kind = 'barh', title = 'Most Important Features')
```
# Making Tuning Dataset
For the random forest model, my kernel kept crashing when optimizing hyperparameters by using the full dataset so I decide to tune the hyperparameters using a 'tuning' dataset which is 1/10 of the original dataset
```
# make tune data which is 1/10 of the train data
X_tune_train = X_train[:len(X_train)//10]
y_tune_train = y_train[:len(y_train)//10]
X_tune_val = X_val[:len(X_val)//10]
y_tune_val = y_val[:len(y_val)//10]
# check to make sure our tuning datasets are indeed 1/10 of our previous training sets
print(f'{X_tune_train.shape}, {X_train.shape}\n{y_tune_train.shape}, {y_train.shape}')
print(f'{X_tune_val.shape}, {X_val.shape}\n{y_tune_val.shape}, {y_val.shape}')
```
# Random Forest Regressor
```
# first hyperparameter to optimize in a random forest is the number of trees
# obviously more trees will only increase the accuracy so we will instead examine when the accuracy approaches
# an asymptote - we will also learn how many estimators we should use for further tuning of hyperparameters, as
# we want enough so that the accuracy is similar to the best accuracy but also not too many for time constraints
estimators = range(10, 201, 10)
train_acc_rf = []
val_acc_rf = []
train_r2_rf = []
val_r2_rf = []
seconds = []
# we will use the previously determined max_depth hyperparameter
for estimator in estimators:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
RandomForestRegressor(random_state = 42,
n_jobs = -1,
max_depth = 12,
n_estimators = estimator))
model.fit(X_tune_train, y_tune_train)
train_acc_rf.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_rf.append(mean_absolute_error(y_train, model.predict(X_train)))
train_r2_rf.append(model.score(X_train, y_train))
val_r2_rf.append(model.score(X_val, y_val))
seconds.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_rf, val_acc_rf, train_r2_rf, val_r2_rf, seconds)), index = estimators,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2', 'Time'])
# determine what percentage of the data to sample for each decision tree
samples = np.arange(0.05, 1, 0.05)
train_acc_rf = []
val_acc_rf = []
train_r2_rf = []
val_r2_rf = []
seconds = []
for sample in samples:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
RandomForestRegressor(random_state = 42,
n_jobs = -1,
max_depth = 12,
n_estimators = 30,
max_samples = sample))
model.fit(X_tune_train, y_tune_train)
train_acc_rf.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_rf.append(mean_absolute_error(y_train, model.predict(X_train)))
train_r2_rf.append(model.score(X_train, y_train))
val_r2_rf.append(model.score(X_val, y_val))
seconds.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_rf, val_acc_rf, train_r2_rf, val_r2_rf, seconds)), index = samples,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2', 'Time'])
model_rf = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
RandomForestRegressor(random_state = 42,
n_jobs = -1,
max_depth = 12,
n_estimators = 180,
max_samples = 0.4))
%%time
model_rf.fit(X_train, y_train)
print('Training MAE:', mean_absolute_error(y_train, model_rf.predict(X_train)))
print('Validation MAE:', mean_absolute_error(y_val, model_rf.predict(X_val)))
print('Validation R2:', model_rf.score(X_val,y_val))
# interestingly, our random forest model is not significantly better than our decision tree
# the top 3 importances are the same as the decision tree
importances = model_rf.named_steps['randomforestregressor'].feature_importances_
features = model_rf.named_steps['onehotencoder'].get_feature_names()
feat_imp_rf = pd.Series(importances, index = features,
name = 'forest').abs().sort_values(ascending=False)
feat_imp_rf.head(10).plot(kind = 'barh')
```
# Boost Model
After testing out decision trees, random forests, boosted random forests, linear regression, and ridge regression, I found that the boosted model gave the lowest MAE and highest R^2 by a significant amount. Given this information, I decided to retrain every hyperparameter instead of assuming that overlapping parameters with other models like max_depth would have the same optimized value.
```
# finding the optimal depth
depths = range(1, 20, 3)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
times = []
for depth in depths:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = depth))
model.fit(X_train, y_train)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, times)), index = depths,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2', 'Time'])
# still finding the optimal depth
depths = range(7, 13)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
times = []
for depth in depths:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = depth))
model.fit(X_train, y_train)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, times)), index = depths,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2', 'Time'])
# optimal depth is 9
# seeing how number of trees affects the model accuracy
estimators = [10] + [i for i in range(50, 300, 50)]
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
times = []
for estimator in estimators:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = estimator))
model.fit(X_train, y_train)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, times)), index = estimators,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2', 'Time'])
# 100 trees should be enough for optimizing hyperparameters, but use >200 for final model
# optimizing learning rate
learns = np.arange(0.01, 0.31, 0.05)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for learn in learns:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = learn))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = learns,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# need to expand learning rate parameter as accuracy kept increasing when learning rate hit 0.26
learns = np.arange(0.25, 0.51, 0.05)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for learn in learns:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = learn))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = learns,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# finding the true optimal learning rate
learns = np.arange(0.25, 0.35, 0.01)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for learn in learns:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = learn))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = learns,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# optimizing colsample_by tree hyperparameter, i.e. the subsample ratio for the columns when constructing
# the dataset for each tree
samples = np.arange(0.01, 1.00, 0.05)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for sample in samples:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bytree = sample))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = samples,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# optimizing colsample_bylevel hyperparameter, i.e. the subsample ratio of the levels in each column
# when constructing each tree
samples = np.arange(0.01, 1.00, 0.05)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for sample in samples:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bylevel = sample))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = samples,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# optimizing colsample_bynode hyperparameter, i.e. the subsample ratio for each column when splitting
# a branch to make two new branches
samples = np.arange(0.01, 1.00, 0.05)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for sample in samples:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bynode = sample))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = samples,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# optimizing subsamples
subsamples = np.arange(0.01, 1.00, 0.05)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for sample in subsamples:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bytree = 1,
subsample = sample))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = subsamples,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# we find that the best subsample value is the default value of 1
# learning if a different boosting method would provide greater accuracy
boosters = ['gbtree', 'gblinear', 'dart']
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for boost in boosters:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bytree = 1,
booster = boost))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = boosters,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# optimizing alphas hyperparameter
alphas = range(0, 100, 5)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for alpha in alphas:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bytree = 1,
reg_alpha = alpha))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = alphas,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# finish optimizing alphas hyperparameter
alphas = range(0, 5)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for alpha in alphas:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bytree = 1,
reg_alpha = alpha))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_tune_train, model.predict(X_tune_train)))
val_acc_boost.append(mean_absolute_error(y_tune_val, model.predict(X_tune_val)))
train_r2_boost.append(model.score(X_tune_train, y_tune_train))
val_r2_boost.append(model.score(X_tune_val, y_tune_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = alphas,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# optimizing lambda hyperparameter
lambdas = range(1, 6)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for l in lambdas:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bytree = 1,
reg_lambda = l))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = lambdas,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# optimizing gamma hyperparameter
gammas = range(0, 6)
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for g in gammas:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bytree = 1,
gamma = g))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = gammas,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# should we choose median instead of mean for SimpleImputer?
strats = ['mean', 'median']
train_acc_boost = []
val_acc_boost = []
train_r2_boost = []
val_r2_boost = []
train_times = []
score_times = []
for strat in strats:
start = time.time()
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = strat),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bytree = 1,
reg_alpha = 0,
reg_lambda = 1,
gamma = 0))
model.fit(X_train, y_train)
train_times.append(time.time() - start)
train_acc_boost.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_boost.append(mean_absolute_error(y_val, model.predict(X_val)))
train_r2_boost.append(model.score(X_train, y_train))
val_r2_boost.append(model.score(X_val, y_val))
score_times.append(time.time() - start)
pd.DataFrame(list(zip(train_acc_boost, val_acc_boost, train_r2_boost, val_r2_boost, train_times, score_times)),
index = strats,
columns = ['Training Accuracy', 'Validation Accuracy', 'Training R^2', 'Validation R^2',
'Train Time', 'Score Time'])
# mean works best, as expected - but median works surprisingly well
# time to build our final model with all our hyperparameters
model_boost = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy='mean'),
XGBRegressor(random_state = 42,
n_jobs = -1,
max_depth = 9,
n_estimators = 100,
learning_rate = 0.29,
colsample_bytree = 1,
reg_alpha = 0,
reg_lambda = 1,
gamma = 0))
%%time
model_boost.fit(X_train, y_train)
%%time
print('Training MAE:', mean_absolute_error(y_train, model_boost.predict(X_train)))
print('Validation MAE:', mean_absolute_error(y_val, model_boost.predict(X_val)))
print('Validation R2:', model_boost.score(X_val, y_val))
# check cross validation score
cross_vals = cross_val_score(model_boost, X, y)
print(np.mean(cross_vals))
# it's actually higher than our previous R^2
importances = model_boost.named_steps['xgbregressor'].feature_importances_
features = model_boost.named_steps['onehotencoder'].get_feature_names()
feat_imp_boost = pd.Series(importances, index = features,
name = 'boost').abs().sort_values(ascending=False)
feat_imp_boost.head(10).plot(kind = 'barh')
```
# Ridge Regression
```
alphas = np.arange(0, 6)
train_acc_ridge = []
val_acc_ridge = []
for a in alphas:
model = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
Ridge(alpha = a))
model.fit(X_train, y_train)
train_acc_ridge.append(mean_absolute_error(y_train, model.predict(X_train)))
val_acc_ridge.append(mean_absolute_error(y_val, model.predict(X_val)))
pd.DataFrame(list(zip(train_acc_ridge, val_acc_ridge)), index = alphas,
columns = ['Training Accuracy', 'Validation Accuracy'])
model_ridge = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
Ridge(alpha = 1))
model_ridge.fit(X_train, y_train)
print('Training MAE:', mean_absolute_error(y_train, model_ridge.predict(X_train)))
print('Validation MAE:', mean_absolute_error(y_val, model_ridge.predict(X_val)))
print('Validation R2:', model_ridge.score(X_val,y_val))
importances = model_ridge.named_steps['ridge'].coef_
features = model_ridge.named_steps['onehotencoder'].get_feature_names()
feat_imp_ridge = pd.Series(importances, index = features,
name = 'ridge').abs().sort_values(ascending=False)
feat_imp_ridge.head(10).plot(kind = 'barh')
```
# Linear Regression
```
model_linear = make_pipeline(OneHotEncoder(use_cat_names = True),
StandardScaler(),
SimpleImputer(strategy = 'mean'),
LinearRegression())
model_linear.fit(X_train, y_train)
print('Training MAE:', mean_absolute_error(y_train, model_linear.predict(X_train)))
print('Validation MAE:', mean_absolute_error(y_val, model_linear.predict(X_val)))
print('Validation R2:', model_linear.score(X_val,y_val))
importances = model_linear.named_steps['linearregression'].coef_
features = model_linear.named_steps['onehotencoder'].get_feature_names()
feat_imp_linear = pd.Series(importances, index = features,
name = 'linear').abs().sort_values(ascending=False)
feat_imp_linear.head(10).plot(kind = 'barh')
```
| github_jupyter |
## For running the T5 vanilla baseline
Here we provide code to
1. Setup datafiles for running t5 (i.e. produce json files)
1. Run the seq2seq model to produce outputs, saving in a directory that matches k_t5_outputs below
- Train model (produces saved checkpoints)
- Eval top performing model (load from top checkpoint, produces json outputs)
1. run eval code using the json output from model eval
```
%load_ext autoreload
%autoreload 2
from decrypt.scrape_parse import (
load_guardian_splits,
load_guardian_splits_disjoint,
load_guardian_splits_disjoint_hash
)
import os
from decrypt import config
from decrypt.common import validation_tools as vt
from decrypt.common.util_data import clue_list_tuple_to_train_split_json
import logging
logging.getLogger(__name__)
k_json_folder = config.DataDirs.Guardian.json_folder
```
## 1 Produce datasets
```
def make_dataset(split_type: str, overwrite=False):
assert split_type in ['naive_random', 'naive_disjoint', 'word_init_disjoint']
if split_type == 'naive_random':
load_fn = load_guardian_splits
tgt_dir = config.DataDirs.DataExport.guardian_naive_random_split
elif split_type == 'naive_disjoint':
load_fn = load_guardian_splits_disjoint
tgt_dir = config.DataDirs.DataExport.guardian_naive_disjoint_split
else:
load_fn = load_guardian_splits_disjoint_hash
tgt_dir = config.DataDirs.DataExport.guardian_word_init_disjoint_split
_, _, (train, val, test) = load_fn(k_json_folder)
os.makedirs(tgt_dir, exist_ok=True)
# write the output as json
try:
clue_list_tuple_to_train_split_json((train, val, test),
comment=f'Guardian data. Split: {split_type}',
export_dir=tgt_dir,
overwrite=overwrite)
except FileExistsError:
logging.warning(f'You have already generated the {split_type} dataset.\n'
f'It is located at {tgt_dir}\n'
f'To regenerate, pass overwrite=True or delete it\n')
make_dataset('naive_random')
make_dataset('word_init_disjoint')
# you can also make_dataset('naive_disjoint')
```
## 2 Running (training) the model
1. Setup environment
1. You should setup wandb for logging (that's where metrics will show up).
If you try to run without wandb, then wandb will tell you what you need to do to initialize
1. The relevant libraries used for our runs are
- transformers==4.4.2
- wandb==0.10.13 # this can probably be updated
- torch==1.7.1+cu110
- torchvision==0.8.2+cu110
- Choose place for your wandb dir, e.g., `'./wandb' `
1. Train the model
1. Note that the default arguments are given in args_cryptic. See `--default_train` and `--default_val`
- Note that, when looking at logging messages or wandb, it will appear that epochs start at 11.
This is done so that we have "space" for 10 "warmup" epochs for curricular training.
This space causes all plots in wandb to line up.
1. from directory seq2seq, run the commands in the box below.
This will produce model checkpoints that can then be used for evaluation.
Baseline naive
```python
python train_clues.py --default_train=base --name=baseline_naive --project=baseline --wandb_dir='./wandb' --data_dir='../data/clue_json/guardian/naive_random'
```
Baseline (naive split), without lengths
```python
python train_clues.py --default_train=base --name=baseline_naive_nolens --project=baseline --wandb_dir='./wandb' --data_dir='../data/clue_json/guardian/word_initial_disjoint' --special=no_lens
```
Baseline disjoint (word initial disjoint)
```python
python train_clues.py --default_train=base --name=baseline_disj --project=baseline --wandb_dir='./wandb' --data_dir='../data/clue_json/guardian/word_initial_disjoint'
```
Baseline disjoint (word initial disjoint), without lengths
```python
python train_clues.py --default_train=base --name=baseline_disj --project=baseline --wandb_dir='./wandb' --data_dir='../data/clue_json/guardian/word_initial_disjoint' --special=no_lens
```
## 3 Evaluating the model
During training we generate only 5 beams for efficiency. For eval we generate 100.
1. Select the best model based on num_match_top_sampled. There should be
a logging statement at the end of the run that prints the location
of the best model checkpoint.
You can also find it by matching the peak in the wandb.ai metrics graph
to the appropriate model save.
2. Run eval using that model (see commands below), which will
produce a file in a (new, different) wandb directory that looks like `epoch_11.pth.tar.preds.json` (i.e only a single epoch)
For example,
Baseline naive, if epoch 20 is best (you'll need to set the run_name)
This produces generations for the validation set
```python
python train_clues.py --default_val=base --name=baseline_naive_val --project=baseline --data_dir='../data/clue_json/guardian/naive_random' --ckpt_path='./wandb/run_name/files/epoch_20.pth.tar
```
To produce generations for the test set,
```python
python train_clues.py --default_val=base --name=baseline_naive_val --project=baseline --data_dir='../data/clue_json/guardian/naive_random' --ckpt_path='./wandb/run_name/files/epoch_10.pth.tar --test
```
This should also be run for the no-lengths versions if you want to replicate those results.
Now we produce metrics by evaluating the json that was produced
1. Change the k_t5_outputs_dir value to the location where you have saved the json files.
- Recommend copying all of the preds.json files into a common directory and working from that.
- Alternatively you could modify the code below and pass in a full path name to each of the json outputs (using the wandb directory path)
1. For each t5 model eval (above) that you ran (each of which produced some `..preds.json` file, run `load_and_run()` to get metrics for those outputs
1. The resulting outputs are the values we report in the tables. See `decrypt/common/validation_tools.ModelEval` for more details about the numbers that are produced. Percentages are prefixed by agg_
Note that for the Main Results Table 2, the metrics we include in the table correspond to
- `agg_top_match`
- `agg_top_10_after_filter`
More details of these metric calculations can be found in `decrypt.common.validation_tools`
```
# for example, if your output files are in
# 'decrypt/t5_outputs/'
# and you will run the below, e.g., if you have named the files
# baseline_naive_e12_test.json
# (.json will be appended for you by the load_and_run_t5 function)
# a better name for load_and_run is load_and_eval
# for example
### primary - test
vt.load_and_run_t5('decrypt/t5_outputs/baseline_naive_e12_test')
vt.load_and_run_t5('decrypt/t5_outputs/baseline_naive_nolens_e15_test') # test set
## primary val
vt.load_and_run_t5('decrypt/t5_outputs/baseline_naive_e12_val')
vt.load_and_run_t5('decrypt/t5_outputs/baseline_naive_nolens_e15_val')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/shivammehta007/Information-Retrieval/blob/master/MLinRanking.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Application of ML in Ranking
# Application of ML in Ranking of Search Engines
## Downloading Data From Kaggle
#### Might Take a while
```
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# Download Data
!pip install kaggle
import json
import os
with open('/content/drive/My Drive/kaggle.json') as kc:
kaggle_config = json.load(kc)
os.environ['KAGGLE_USERNAME'] = kaggle_config['username']
os.environ['KAGGLE_KEY'] = kaggle_config['key']
!kaggle competitions download -c text-relevance-competition-ir-2-itmo-fall-2019
!unzip docs.tsv.zip
!rm docs.tsv.zip
```
## Code
```
!pip install -U tqdm
!pip install rank_bm25
!pip install keyw
import os
import gzip
import nltk
import random
import pickle
import pandas as pd
import keyw
import logging
import itertools
import re
import time
import numpy as np
import pandas as pd
from rank_bm25 import BM25Okapi
from nltk.corpus import stopwords
nltk.download("stopwords")
from string import punctuation
from collections import defaultdict
from collections import OrderedDict
from tqdm.notebook import tqdm
from sklearn.metrics.pairwise import cosine_similarity
from sklearn.feature_extraction.text import TfidfVectorizer
logging.basicConfig(level=logging.DEBUG)
logging.debug('Test Logger')
indexing_data_location = '/content/docs.tsv'
queries_list = '/content/queries.numerate.txt'
```
### Building Inverted Index
#### Inverted Indexer Class
```
class InvertedIndexer:
"""
This class makes inverted index
"""
def __init__(self, filename=False):
self.filename = filename
self.stemmer_ru = nltk.SnowballStemmer("russian")
self.stopwords = set(stopwords.words("russian")) | set(stopwords.words("english"))
self.punctuation = punctuation # from string import punctuation
if filename:
self.inverted_index = self._build_index(self.filename)
else:
self.inverted_index = defaultdict(set)
def preprocess(self, sentence):
"""
Method to remove stop words and punctuations return tokens
"""
NONTEXT = re.compile('[^0-9 a-z#+_а-яё]')
sentence = sentence.lower()
sentence = sentence.translate(str.maketrans('', '', punctuation))
sentence = re.sub(NONTEXT,'',sentence)
tokens = [token for token in sentence.split() if token not in self.stopwords]
return tokens
def stem_keys(self, inverted_index):
"""
Called after index is built to stem all the keys and normalize them
"""
logging.debug('Indexing Complete will not Stem keys and remap indexes')
temp_dict = defaultdict(set)
i = 0
for word in tqdm(inverted_index):
stemmed_key = self.stemmer_ru.stem(word)
temp_dict[stemmed_key].update(inverted_index[word])
inverted_index[word] = None
inverted_index = temp_dict
logging.debug('Done Stemmping Indexes')
return inverted_index
def _build_index(self, indexing_data_location):
"""
This method builds the inverted index and returns the invrted index dictionary
"""
inverted_index = defaultdict(set)
with open(indexing_data_location, "rb") as f:
for line in tqdm(f):
line = line.decode().split('\t')
file_number = line[0]
subject = line[1]
text = line[2]
line = subject + ' ' + text
for word in self.preprocess(line):
inverted_index[word].add(int(file_number))
inverted_index = self.stem_keys(inverted_index)
return inverted_index
def save(self, filename_to_save):
"""
Save method to save the inverted indexes
"""
with open(filename_to_save, mode='wb') as f:
pickle.dump(self.inverted_index, f)
def load(self, filelocation_to_load):
"""
Load method to load the inverted indexes
"""
with open(filelocation_to_load, mode='rb') as f:
self.inverted_index = pickle.load(f)
```
#### SolutionPredictor Class
```
class SolutionPredictor:
"""
This classes uses object of Hw1SolutionIndexer
to make boolean search
"""
def __init__(self, indexer):
"""
indexer : object of class Hw1SolutionIndexer
"""
self.indexer = indexer
def find_docs(self, query):
"""
This method provides booleaen search
query : string with text of query
Returns Python set with documents which contain query words
Should return maximum 100 docs
"""
query = keyw.engrus(query)
tokens = self.indexer.preprocess(query)
tokens = [self.indexer.stemmer_ru.stem(word) for word in tokens]
docs_list = set()
for word in tokens:
if word in self.indexer.inverted_index:
if len(docs_list) > 0:
docs_list.intersection_update(self.indexer.inverted_index[word]) # changed from intersection_update
else:
docs_list.update(self.indexer.inverted_index[word])
return docs_list
```
#### Generate Index
Run Only when Index is not generated Otherwise use the Loading code block
```
logging.debug('Index is creating...')
start = time.time()
new_index = InvertedIndexer(indexing_data_location)
end = time.time()
logging.debug('Index has been created and saved as inverted_index.pickle in {:.4f}s'.format(end-start))
len(new_index.inverted_index)
new_index.save('inverted_index.pickle')
!cp inverted_index.pickle '/content/drive/My Drive/Homeworks/InformationRetrieval/inverted_index_4.pickle'
```
#### Load Index
Run when Index is not generated
```
index_location = '/content/drive/My Drive/Homeworks/InformationRetrieval/inverted_index_all_rus.pickle'
logging.debug('Loading Index...')
start = time.time()
new_index = InvertedIndexer()
new_index.load(index_location)
end = time.time()
logging.debug('Index has been loaded from inverted_index.pickle in {:.4f}s'.format(end-start))
len(new_index.inverted_index)
```
#### Convert keys to all Russian only text
```
temp_dict = defaultdict(set)
i = 0
for word in new_index.inverted_index:
i += 1
if i % 100000 == 0:
print(i)
rus_key = keyw.engrus(word)
temp_dict[rus_key].update(new_index.inverted_index[word])
new_index.inverted_index[word] = None
new_index.inverted_index = temp_dict
temp_dict = None
del temp_dict
len(new_index.inverted_index)
new_index.save('inverted_index_all_rus_no_stem.pickle')
!cp inverted_index_all_rus_no_stem.pickle '/content/drive/My Drive/Homeworks/InformationRetrieval/inverted_index_all_rus_no_stem.pickle'
```
##### Testing Block
```
test_sentence = 'бандиты боятся ли ментов'
boolean_model = SolutionPredictor(new_index)
start = time.time()
value_present = boolean_model.find_docs(test_sentence)
print(value_present)
logging.debug('Search Time: {}'.format(time.time() - start))
```
### Loading File to DataFrame
We are doing this for faster search based on index numbers
```
%%time
database = pd.read_csv(indexing_data_location, sep='\t', header=None,
names=['id', 'subject', 'content'],
dtype = {'id': int,'subject': str,
'content': str})
database.head()
```
### Implementing TFIDF Cosine Distances for text relevancy
```
# boolean_model = SolutionPredictor(new_index)
output=open('submission.csv', 'w')
with open(queries_list) as queries:
output.write('QueryId,DocumentId\n')
for line in tqdm(queries.readlines()):
line = line.split('\t')
id = int(line[0])
query = line[1]
value_present = boolean_model.find_docs(query)
data = database[database['id'].isin(list(value_present))].copy()
data.loc[:, 'content'] = data[['subject', 'content']].astype(str).apply(' '.join, axis=1)
del data['subject']
data = OrderedDict(data.set_index('id')['content'].to_dict())
for key in data:
data[key] = ' '.join(new_index.preprocess(data[key]))
if data:
tfidf = TfidfVectorizer()
ids = data.keys()
data = tfidf.fit_transform(data.values())
query = tfidf.transform([query])[0]
distances = cosine_similarity(query, data)
rankings = sorted(list(zip(ids, distances[0])), key=lambda x: x[1], reverse=True)[:10]
for rank in rankings:
output.write('{},{}\n'.format(id, rank[0]))
data = None
query = None
distances = None
rankings = None
output.close()
!cp submission.csv '/content/drive/My Drive/Homeworks/InformationRetrieval/submission_tfidf.csv'
```
# Crossed Baseline with this submission
## Implementing BM25 for text relevancy
(could not run for memery issues with all boolean searches check different doc for its working)
```
# boolean_model = SolutionPredictor(new_index)
output=open('submission.csv', 'w')
with open(queries_list) as queries:
output.write('QueryId,DocumentId\n')
for line in tqdm(queries.readlines()):
line = line.split('\t')
id = int(line[0])
query = line[1]
value_present = boolean_model.find_docs(query)
data = database[database['id'].isin(list(value_present))].copy()
data.loc[:, 'content'] = data[['subject', 'content']].astype(str).apply(' '.join, axis=1)
del data['subject']
data = OrderedDict(data.set_index('id')['content'].to_dict())
for key in data:
data[key] = new_index.preprocess(data[key])
if data:
bm25 = BM25Okapi(data.values())
rankings = sorted(list(zip(data.keys(), bm25.get_scores(new_index.preprocess(query)))), key=lambda x: x[1], reverse=True)[:10]
for rank in rankings:
output.write('{},{}\n'.format(id, rank[0]))
if (not data) or len(rankings) < 10:
rankings = []
logging.debug('Found Some values with no query results or less than 10 {} ranking'.format(id, len(rankings)))
print(id, query)
# for i in range(10 - len(rankings)):
# output.write('{},{}\n'.format(id, random.randrange(50000)))
output.close()
```
| github_jupyter |
# Import packages and functions
```
import sys
# force the notebook to look for files in the upper level directory
sys.path.insert(1, '../')
import pandas as pd
from glob import glob
import pymatgen as mg
from data.compound_featurizer import read_new_struct, \
get_struct, get_elem_info, get_elem_distances, \
calc_mm_dists, calc_mx_dists, calc_xx_dists, calc_elem_max_potential
```
# Read in the initial dataframe
```
# initialize an empty list of dataframes
df_lst = []
# iterate over all the cif files
for struct_file_path in glob("./user_defined_structures/featurizer_sub_function_demo/*.cif"):
# add the newly read in dataframe to the list
df_lst.append(read_new_struct(struct_file_path))
# concatenate all the dataframes in the list
df = pd.concat(df_lst, ignore_index=True)
# assign oxidation states to BaTiO3 and Mg2AlFeO5
df.at[df[df.Compound == "BaTiO3"].index[0], "structure"].add_oxidation_state_by_element({"Ba": 2, "Ti": 4, "O": -2})
df.at[df[df.Compound == "Mg2AlFeO5"].index[0], "structure"].add_oxidation_state_by_element({"Mg": 2, "Al": 3, "Fe": 3, "O": -2})
# here is a print out of the dataframe
df
```
# Demo usage of relevant sub-functions
## 1. get_struct("compound_formula", input_df) -> Pymatgen Structure
Since we've already read in all the structures in dataframe, we can access the individual Pymatgen structure using the compound formula.
_Tip_: when you have questions about a specific function, you can always go to the original .py file or you can press <kbd>⇧ Shift</kbd> + <kbd>⇥ Tab</kbd> for its docstring
```
test_struct = get_struct("BaTiO3", df)
test_struct
```
If you happen to type in a formula that doesn't have an exact match, the function will return an error message along with several possible suggestions
```
get_struct("BaTiO", df)
```
_BaTiO3_ will be used consistently as the demo test structure from now on.
## 2. get_elem_distances(Pymatgen_Structure, Pymatgen_Element_1, Pymatgen_Element_2) -> Array of distances (Å)
Now that we have the structure, we can use **get_elem_distances()** to calculate the distance between any two elements in the structure
But before doing that, we first need to know which site(s) each element occupies through the **get_elem_info()** function
```
elem_indices, _, modified_struct = get_elem_info(test_struct)
print(elem_indices, "\n")
print(modified_struct)
```
If you compare this to the printout from the original, you will find that the modified structure have double the amount of sites
```
print(test_struct)
```
This is because if we keep the original function, _Ba_ and _Ti_ will only occupy one site
```
elem_indices_orig, *_ = get_elem_info(test_struct, makesupercell=False)
elem_indices_orig
```
The reason for returning a supercell of the original structure is related to the inner workings of **get_elem_distances()** function. It basically works by getting the site indices of the two elements (they can be the same) and using the built-in method of **pymatgen.Structure.get_distance(i, j)** to calculate the distance between site i and site j. There is one scenario where only using the original structure can cause a problem:
1. If we have a structure where an element only occupies one site and we want to know the distance between the same elements, e.g. _Ba_-_Ba_ or _Ti_-_Ti_ in _BaTiO3_, we would have **pymatgen.Structure.get_distance(i, j)** where i = j and we would only get 0 for that distance.
By making a supercell (in this case a'=2a, b'=b, c'=c), we would be able to get a non-zero distance betweem the original site and the newly translated site along the a-axis. That being said, if all elements in the original structure all occupy more than one site, the structure will not be modified.
Let's first try to calculate the _Ba_-_Ba_ distance using the supercell structure
```
get_elem_distances(test_struct,
elem_1=mg.Element("Ba"),
elem_indices=elem_indices, only_unique=True)
```
**Note**: when the `only_unique` parameter is set to be `True`, the function will only return the unique values of distance since in a structure the same distance can occur multiple times due to symmetry.
Let's see what happens when we use the original reduced structure
```
get_elem_distances(test_struct,
elem_1=mg.Element("Ba"),
elem_indices=elem_indices_orig, only_unique=True)
```
As expected, we get 0 Å. We can also calculate the distance between different elements. Let's see the distance between _Ti_ and _O_
```
get_elem_distances(test_struct,
elem_1=mg.Element("O"), elem_2=mg.Element("Ti"),
elem_indices=elem_indices_orig, only_unique=True)
```
This function can also handle structures where multiple elements can occupy the same site (La$_{2.8}$Mg$_{1.2}$Mn$_4$O$_{12}$ is a made-up structure generated for the purpose of this demo)
```
special_struct = get_struct("La2.8Mg1.2Mn4O12", df)
print(special_struct)
elem_indices, *_ = get_elem_info(special_struct)
distances = get_elem_distances(special_struct,
elem_1=mg.Element("La"), elem_2=mg.Element("Mn"),
elem_indices=elem_indices, only_unique=True)
distances
```
It may seem that there are some distances that are equal to each other, but since the values displayed do not have all the decimal places shown, there are still slight differences among them.
```
distances[0] - distances[1]
```
## 3. Wrapper functions around get_elem_distances() to calculate distances between different types of elements
### 3.1 calc_mm_dists() to calculate distances between metal-metal elements
```
calc_mm_dists(test_struct, return_unique=True)
```
### 3.2 calc_mx_dists() to calculate distances between metal-non_metal elements
```
calc_mx_dists(test_struct, return_unique=True)
```
### 3.3 calc_xx_dists() to calculate distances between non_metal-non_metal elements
```
calc_xx_dists(test_struct, return_unique=True)
```
This functionality is realized again through the **get_elem_info()** function where all the elements in the structure is classified as either a metal or a non_metal.
```
_, elem_groups, _ = get_elem_info(test_struct)
elem_groups
```
Once we know which elements are metal and which ones are non_metal, we can then use the elem_indices to find where they are (i.e. the site indices) and compute the distances using the generic element distance finder **get_elem_distances()**.
## 4. calc_elem_max_potential() to calculate Madelung Site Potentials
The **calc_elem_max_potential()** utilizes the EwaldSummation() module from Pymatgen to calculate site energy for all the sites in a structure and convert the site energy to site potential using the relation as follows. ($U_{E_\text{tot}}$: the total potential energy of the structure, $U_{E_i}$: the site energy at site i, $N$: the total number of sites, $q_i$: the charge at site i, $\Phi(r_i)$: the site potential at site i)
$$
\begin{align*}
U_{E_\text{tot}}&=\sum_{i=1}^{N}U_{E_i}=\frac{1}{2}\sum_{i=1}^{N}q_i\Phi(r_i)\\
U_{E_i}&=\frac{1}{2}q_i\Phi(r_i)\\
\Phi(r_i)&=\frac{2U_{E_i}}{q_i}
\end{align*}
$$
The default output unit for the Madelung site potential is in $V$
```
calc_elem_max_potential(test_struct, full_list=True)
```
But the unit can be converted from $V$ to $e/Å$ for easier comparison with the results from VESTA
```
calc_elem_max_potential(test_struct, full_list=True, check_vesta=True)
```
If we don't specify the `full_list` parameter, it will be set to `False` and the function only return the maximum site potential for each element.
```
calc_elem_max_potential(test_struct)
```
Just like before, this function can also work with structures where multiple elements occupy the same site. We can try a compound with non-integer stoichiometry this time. (again, Mg$_2$AlFeO$_5$ is a made-up structure)
```
non_stoich_struct = get_struct("Mg2AlFeO5", df)
print(non_stoich_struct)
calc_elem_max_potential(non_stoich_struct, check_vesta=True)
```
# Now it's your turn
If you want to test the functions with structures that are not in the loaded dataframe, you can also upload your own .cif file to the `user_defined` folder located at this path
_./user_defined_structures/_
```
USER_DEFINED_FOLDER_PATH = "./user_defined_structures/"
example_new_struct = mg.Structure.from_file(USER_DEFINED_FOLDER_PATH + "CuNiO2_mp-1178372_primitive.cif")
example_new_struct
```
## Define a wrapper function around get_elem_distances()
```
def get_elem_distances_wrapper(structure: mg.Structure, **kwargs):
"""A wrapper function around get_elem_distances() such that there is no need to get elem_indices manually"""
elem_indices, _, structure = get_elem_info(structure)
return get_elem_distances(structure, elem_indices=elem_indices, only_unique=True, **kwargs)
```
Check the _Cu_-_Ni_ distance
```
get_elem_distances_wrapper(example_new_struct, elem_1=mg.Element("Cu"), elem_2=mg.Element("Ni"))
```
Check the _Ni_-_O_ distance
```
get_elem_distances_wrapper(example_new_struct, elem_1=mg.Element("O"), elem_2=mg.Element("Ni"))
```
Check the _Cu_-_Cu_ distance
```
get_elem_distances_wrapper(example_new_struct, elem_1=mg.Element("Cu"))
```
## Get distances of all three types of element pairs
```
calc_mm_dists(example_new_struct)
calc_mx_dists(example_new_struct)
calc_xx_dists(example_new_struct)
```
## A note for site potential calculation
To use the EwaldSummation technique, the input structure has to have oxidation states (that's where the charge value comes from) associated with all the sites. A structure without oxidation states will raise an error in the function.
```
calc_elem_max_potential(example_new_struct)
```
To overcome this problem, we can add oxidation states to the structure using the add_oxidation_state_by_guess() method from Pymatgen
```
example_new_struct.add_oxidation_state_by_guess()
example_new_struct
```
Now that we should be able to obtain proper results from the function.
```
calc_elem_max_potential(example_new_struct, check_vesta=True)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import bokeh
import bokeh.plotting
from bokeh.plotting import ColumnDataSource
from bokeh.models import LabelSet
from bokeh.models import FuncTickFormatter
import bokeh.io
bokeh.io.output_notebook()
df = pd.read_csv('191031_Deuterium_Transfer_Peak_Areas.csv', comment='#')
df.tail()
#function to normalize the peak are of each unique peak using the area of the C18 peak for each sample
def normalize_area(data):
#check to make sure that the dataframe hasn't already been normalized
if not {'Normalized Area', 'Hydrocarbon amount'}.issubset(data.columns):
#Create array of c18 values repeated in groups of three, matching the number of unique peaks for each sample
c18_Areas = np.repeat(data['Area'].loc[data['Peak ID'] == 'C18'],3).reset_index().drop("index", axis=1)
#Divide the area of each peak in the dataframe by the corresponding c18 peak area
data['Normalized Area'] = np.divide(data['Area'],c18_Areas['Area'])
#Calculate the hydrocarbon amount, in ng, by multiplying the normalized peak area by 25.
#25 ng of c18 was injected in each sample (1 microliter of a 25 ng/microliter solution of c18 in hexane)
data['Hydrocarbon amount'] = data['Normalized Area']*25
return data
df = normalize_area(df)
#function to create an identifier for each sample in the order (Liometopum or not : Beetle: Type)
def add_identifier(data):
#check if identifier column already exists
if not {'Identifier'}.issubset(data.columns):
#create identifier column, populating it with the beetle species used in each well
data = data.merge(data[['WellNumber','Species']].loc[data['Species'] != 'L'].rename(columns={"Species": "Identifier"}).drop_duplicates())
#modify the identifier column, adding a leading 'L' for Liometopum runs and adding a suffix from the 'Type' column
data['Identifier'] = ((data['Species']=='L')*pd.Series('L',index=data.index)) + data['Identifier'] + data['Type']
return data
df = add_identifier(df)
def box_and_whisker(data,plt,x_vals,y_vals):
groups = data.groupby(x_vals)
q1 = groups.quantile(q=0.25)
q2 = groups.quantile(q=0.5)
q3 = groups.quantile(q=0.75)
iqr = q3 - q1
upper = q3 + 1.5*iqr
lower = q1 - 1.5*iqr
# find the outliers for each category
def outliers(group):
cat = group.name
return group[(group[y_vals] > upper.loc[cat][y_vals]) | (group[y_vals] < lower.loc[cat][y_vals])][y_vals]
out = groups.apply(outliers).dropna()
# prepare outlier data for plotting, we need coordinates for every outlier.
if not out.empty:
outx = []
outy = []
for keys in out.index:
outx.append(keys[0])
outy.append(out.loc[keys[0]].loc[keys[1]])
# if no outliers, shrink lengths of stems to be no longer than the minimums or maximums
qmin = groups.quantile(q=0.00)
qmax = groups.quantile(q=1.00)
upper[y_vals] = [min([x,y]) for (x,y) in zip(list(qmax.loc[:,y_vals]),upper[y_vals])]
lower[y_vals] = [max([x,y]) for (x,y) in zip(list(qmin.loc[:,y_vals]),lower[y_vals])]
# stems
plt.segment(q3.reset_index()[x_vals], upper[y_vals], q3.reset_index()[x_vals], q3[y_vals], line_color="black")
plt.segment(q3.reset_index()[x_vals], lower[y_vals], q3.reset_index()[x_vals], q1[y_vals], line_color="black")
#boxes
plt.vbar(q3.reset_index()[x_vals], 0.7, q2[y_vals], q3[y_vals], fill_color=None, line_color="black")
plt.vbar(q3.reset_index()[x_vals], 0.7, q1[y_vals], q2[y_vals], fill_color=None, line_color="black")
# whiskers (almost-0 height rects simpler than segments)
plt.vbar(q3.reset_index()[x_vals], top=lower[y_vals],bottom=lower[y_vals], width=0.2, line_color="black")
plt.vbar(q3.reset_index()[x_vals], top=upper[y_vals],bottom=upper[y_vals], width=0.2, line_color="black")
return plt
df.head()
plotting_df=df.loc[df['Peak ID']=='D50C24']
Identifiers = ['LSU','SU','LSL','SL','LPU','PU','LPL','PL','LDU','DU','LDL','DL',]
Labels={'LSU': '','SU': 'Sceptobius Control','LSL': '','SL': 'Sceptobius Treated',
'LPU': '','PU': 'Platyusa Control','LPL': '','PL': 'Platyusa Treated',
'LDU': '','DU': 'Dalotia Control','LDL': '','DL': 'Dalotia Treated'}
np.random.seed(666)
p = bokeh.plotting.figure(plot_width=800,
plot_height=600,
title='C24',
x_range=Identifiers,
y_axis_label='ng D50C24',
y_axis_type='log')
colors=['#494949','#5F56FF','#494949','#5F56FF','#494949','#C42F2F','#494949','#C42F2F','#494949','#832161','#494949','#832161']
p = box_and_whisker(plotting_df,p,'Identifier','Hydrocarbon amount')
for _,i in enumerate(np.unique(plotting_df['WellNumber'])):
data = {'identifier': plotting_df['Identifier'].loc[(plotting_df['WellNumber']==i)].values,
'hydrocarbon amount': plotting_df['Hydrocarbon amount'].loc[(plotting_df['WellNumber']==i)].values}
source = ColumnDataSource(data=data)
outlier=np.unique(plotting_df['Outlier'].loc[(plotting_df['WellNumber']==i)])
offsetVal=(np.random.rand(1)[0]-0.5)*0.5
p.line(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='black',
alpha=0.3)
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color=bokeh.transform.factor_cmap('identifier',colors,Identifiers),
alpha=0.6,
size=7)
if outlier=='AI':
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='lime',
alpha=1,
size=7,
legend='Ant Injured')
if outlier=='A':
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='purple',
alpha=1,
size=7,
legend='Ant Chomped')
if outlier=='BI':
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='orange',
alpha=1,
size=7,
legend='Beetle Injured')
if outlier=='B':
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='pink',
alpha=1,
size=7,
legend='Beetle Chomped')
if outlier=='Y':
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='cyan',
alpha=1,
size=7,
legend='Ant Crushed or Not Labeled')
p.xgrid.visible = False
p.ygrid.visible = False
# Add custom axis
p.xaxis.formatter = FuncTickFormatter(code="""
var labels = %s;
return labels[tick];
""" %Labels)
bokeh.io.show(p)
plotting_df=df.loc[df['Peak ID']=='D62C30']
Identifiers = ['LSU','SU','LSL','SL','LPU','PU','LPL','PL','LDU','DU','LDL','DL',]
Labels={'LSU': '','SU': 'Sceptobius Control','LSL': '','SL': 'Sceptobius Treated',
'LPU': '','PU': 'Platyusa Control','LPL': '','PL': 'Platyusa Treated',
'LDU': '','DU': 'Dalotia Control','LDL': '','DL': 'Dalotia Treated'}
np.random.seed(666)
p = bokeh.plotting.figure(plot_width=800,
plot_height=600,
title='C30',
x_range=Identifiers,
y_axis_label='ng D62C30',
y_axis_type='log')
colors=['#494949','#5F56FF','#494949','#5F56FF','#494949','#C42F2F','#494949','#C42F2F','#494949','#832161','#494949','#832161']
p = box_and_whisker(plotting_df,p,'Identifier','Hydrocarbon amount')
for _,i in enumerate(np.unique(plotting_df['WellNumber'])):
data = {'identifier': plotting_df['Identifier'].loc[(plotting_df['WellNumber']==i)].values,
'hydrocarbon amount': plotting_df['Hydrocarbon amount'].loc[(plotting_df['WellNumber']==i)].values}
source = ColumnDataSource(data=data)
outlier=np.unique(plotting_df['Outlier'].loc[(plotting_df['WellNumber']==i)])
offsetVal=(np.random.rand(1)[0]-0.5)*0.5
p.line(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='black',
alpha=0.3)
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color=bokeh.transform.factor_cmap('identifier',colors,Identifiers),
alpha=0.6,
size=7)
if outlier=='AI':
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='lime',
alpha=1,
size=7,
legend='Ant Injured')
if outlier=='A':
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='purple',
alpha=1,
size=7,
legend='Ant Chomped')
if outlier=='BI':
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='orange',
alpha=1,
size=7,
legend='Beetle Injured')
if outlier=='B':
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='pink',
alpha=1,
size=7,
legend='Beetle Chomped')
if outlier=='Y':
p.circle(bokeh.transform.dodge('identifier', offsetVal, range=p.x_range),
'hydrocarbon amount',
source=source,
color='cyan',
alpha=1,
size=7,
legend='Ant Crushed or Not Labeled')
p.xgrid.visible = False
p.ygrid.visible = False
# Add custom axis
p.xaxis.formatter = FuncTickFormatter(code="""
var labels = %s;
return labels[tick];
""" %Labels)
bokeh.io.show(p)
```
| github_jupyter |
```
import sys
import numpy as np
import scipy.io as sio
import keras
import numpy as np
import os
import matplotlib
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from keras.optimizers import SGD
from keras.optimizers import Adam, Adadelta
from keras.callbacks import ModelCheckpoint, ReduceLROnPlateau
from keras import backend as K
from keras.datasets import cifar10
from keras.models import load_model
from data_input.data_input import getDataGenerator
from model.DenseNet import createSTNDenseNet
def flip(data):
y_4 = np.zeros_like(data)
y_1 = y_4
y_2 = y_4
first = np.concatenate((y_1, y_2, y_1), axis=1)
second = np.concatenate((y_4, data, y_4), axis=1)
third = first
Data = np.concatenate((first, second, third), axis=0)
return Data
```
# 数据集1
```
matfn1 = 'data/dsm/dsm.mat'
data1 = sio.loadmat(matfn1)
dsm = data1['dsm']
matfn2 = 'data/dsm/truemap.mat'
data2 = sio.loadmat(matfn2)
groundtruth = data2['groundtruth']
```
# 数据集2
```
matfn1 = 'data/recology/Recology_data_DSM_cube.mat'
data1 = sio.loadmat(matfn1)
dsm = data1['Recology_data_DSM_cube']
matfn2 = 'data/recology/Recology_truthmap.mat'
data2 = sio.loadmat(matfn2)
groundtruth = data2['groundtruth']
dsm = 1 * ((dsm - np.min(dsm)) / (np.max(dsm) - np.min(dsm)) - 0.5)
[nRow, nCol] = dsm.shape
nTrain = 3000
nTest = 2000
num_class = int(np.max(groundtruth))
dsm=flip(dsm)
groundtruth = flip(groundtruth)
HalfWidth = 16
Wid = 2 * HalfWidth
G = groundtruth[nRow-HalfWidth : 2 * nRow + HalfWidth, nCol - HalfWidth : 2 * nCol + HalfWidth]
data = dsm[nRow-HalfWidth : 2 * nRow + HalfWidth, nCol - HalfWidth : 2 * nCol + HalfWidth]
[row, col] = G.shape
NotZeroMask = np.zeros([row, col])
Wid = 2 * HalfWidth
NotZeroMask[HalfWidth + 1: -1 - HalfWidth + 1, HalfWidth + 1: -1 - HalfWidth + 1] = 1
G = G * NotZeroMask
[Row, Col] = np.nonzero(G)
nSample = np.size(Row)
imdb = {}
imdb['data'] = np.zeros([nTrain + nTest, 2 * HalfWidth, 2 * HalfWidth],dtype=np.float64)
imdb['Labels'] = np.zeros([nTrain + nTest], dtype=np.int64)
imdb['set'] = np.zeros([nTrain + nTest], dtype=np.int64)
RandPerm = np.random.permutation(nSample)
for iSample in range(nTrain + nTest):
imdb['data'][iSample,:, :] = data[Row[RandPerm[iSample]] - HalfWidth: Row[RandPerm[iSample]] + HalfWidth, \
Col[RandPerm[iSample]] - HalfWidth: Col[RandPerm[iSample]] + HalfWidth].astype(np.float64)
imdb['Labels'][iSample] = G[Row[RandPerm[iSample]],
Col[RandPerm[iSample]]].astype(np.int64)
print('Data is OK.')
imdb['data'].shape
num_class
#define DenseNet parms
ROWS = 32
COLS = 32
CHANNELS = 1
nb_classes = num_class
batch_size = 4
nb_epoch = 100
img_dim = (ROWS,COLS,CHANNELS)
densenet_depth = 40
densenet_growth_rate = 12
#define filepath parms
check_point_file = r"./densenet_check_point.h5"
loss_trend_graph_path = r"./loss.jpg"
acc_trend_graph_path = r"./acc.jpg"
resume = False
print('Now,we start compiling DenseNet model...')
model = createSTNDenseNet(nb_classes=nb_classes,img_dim=img_dim,depth=densenet_depth,
growth_rate = densenet_growth_rate)
if resume == True:
try:
model.load_weights(check_point_file)
except:
pass
# optimizer = Adam()
optimizer = SGD(lr=0.001)
#optimizer = Adadelta()
model.compile(loss='categorical_crossentropy',optimizer=optimizer,metrics=['accuracy'])
print('Now,we start loading data...')
data_x = imdb['data']
data_x = np.expand_dims(data_x, axis=3)
data_y = imdb['Labels']-1
x_train, x_test, y_train, y_test = train_test_split(data_x, data_y, test_size=0.1, random_state=42)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
y_train = keras.utils.to_categorical(y_train, nb_classes)
y_test= keras.utils.to_categorical(y_test, nb_classes)
print('Now,we start defining callback functions...')
"""
lr_reducer = ReduceLROnPlateau(monitor='val_acc', factor=np.sqrt(0.1),
cooldown=0, patience=3, min_lr=1e-6)
"""
model_checkpoint = ModelCheckpoint(check_point_file, monitor="val_acc", save_best_only=True,
save_weights_only=True, verbose=1)
#callbacks=[lr_reducer,model_checkpoint]
callbacks=[model_checkpoint]
print("Now,we start training...")
history = model.fit(x_train, y_train,
epochs=nb_epoch,
batch_size = batch_size,
callbacks=callbacks,
validation_data=(x_test,y_test),
verbose=1)
print("Now,we start drawing the loss and acc trends graph...")
#summarize history for accuracy
fig = plt.figure(1)
plt.plot(history.history["acc"])
plt.plot(history.history["val_acc"])
plt.title("Model accuracy")
plt.ylabel("accuracy")
plt.xlabel("epoch")
plt.legend(["train","test"],loc="upper left")
plt.savefig(acc_trend_graph_path)
plt.close(1)
#summarize history for loss
fig = plt.figure(2)
plt.plot(history.history["loss"])
plt.plot(history.history["val_loss"])
plt.title("Model loss")
plt.ylabel("loss")
plt.xlabel("epoch")
plt.legend(["train","test"],loc="upper left")
plt.savefig(loss_trend_graph_path)
plt.close(2)
print("We are done, everything seems OK...")
data_x.shape
data_y.shape
```
| github_jupyter |
# Navigation
---
You are welcome to use this coding environment to train your agent for the project. Follow the instructions below to get started!
### 1. Start the Environment
Run the next code cell to install a few packages. This line will take a few minutes to run!
```
import os
if os.path.isdir('./python'):
!pip -q install ./python
else:
from unityagents import UnityEnvironment
import numpy as np
```
The environment is already saved in the Workspace and can be accessed at the file path provided below. Please run the next code cell without making any changes.
```
from unityagents import UnityEnvironment
import numpy as np
# TODO: MODIFY THE FILENAME TO FIT YOUR ENVIRONMENT
BANANA_FNAME = "/data/Banana_Linux_NoVis/Banana.x86_64"
env = UnityEnvironment(file_name=BANANA_FNAME)
```
Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
```
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
```
### 2. Examine the State and Action Spaces
Run the code cell below to print some information about the environment.
```
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents in the environment
print('Number of agents:', len(env_info.agents))
# number of actions
action_size = brain.vector_action_space_size
print('Number of actions:', action_size)
# examine the state space
state = env_info.vector_observations[0]
print('States look like:', state)
state_size = len(state)
print('States have length:', state_size)
```
### 3. Take Random Actions in the Environment
In the next code cell, you will learn how to use the Python API to control the agent and receive feedback from the environment.
Note that **in this coding environment, you will not be able to watch the agent while it is training**, and you should set `train_mode=True` to restart the environment.
```
# TODO: UNCOMMMENT BELOW IF YOU WANT TO WATCH RANDOM ACTIONS
# env_info = env.reset(train_mode=True)[brain_name] # reset the environment
# state = env_info.vector_observations[0] # get the current state
# score = 0 # initialize the score
# while True:
# action = np.random.randint(action_size) # select an action
# env_info = env.step(action)[brain_name] # send the action to the environment
# next_state = env_info.vector_observations[0] # get the next state
# reward = env_info.rewards[0] # get the reward
# done = env_info.local_done[0] # see if episode has finished
# score += reward # update the score
# state = next_state # roll over the state to next time step
# if done: # exit loop if episode finished
# break
# print("Score: {}".format(score))
```
### 4. It's Your Turn!
Now it's your turn to train your own agent to solve the environment! A few **important notes**:
- When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
```python
env_info = env.reset(train_mode=True)[brain_name]
```
- To structure your work, you're welcome to work directly in this Jupyter notebook, or you might like to start over with a new file! You can see the list of files in the workspace by clicking on **_Jupyter_** in the top left corner of the notebook.
- In this coding environment, you will not be able to watch the agent while it is training. However, **_after training the agent_**, you can download the saved model weights to watch the agent on your own machine!
```
from dqn_agent import Agent
from model import QNetwork
from collections import deque
import torch
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents in the environment
print('Number of agents:', len(env_info.agents))
# number of actions
action_size = brain.vector_action_space_size
print('Number of actions:', action_size)
# examine the state space
state = env_info.vector_observations[0]
print('States look like:', state)
state_size = len(state)
print('States have length:', state_size)
# initialize agent
agent = Agent(state_size=state_size, action_size=action_size, seed=12)
# enable double DQN
agent.enable_double_DQN()
def get_state(env_info):
return env_info.vector_observations[0]
def get_reward(env_info):
return env_info.rewards[0]
def get_done(env_info):
return env_info.local_done[0]
def extract_full_state(env_info):
return get_state(env_info), get_reward(env_info), get_done(env_info)
# run dqn training
def dqn(n_episodes=3000, max_t=1000, eps_start=1.0, eps_end=0.01, eps_decay=0.995):
"""Deep Q-Learning.
Params
======
n_episodes (int): maximum number of training episodes
max_t (int): maximum number of timesteps per episode
eps_start (float): starting value of epsilon, for epsilon-greedy action selection
eps_end (float): minimum value of epsilon
eps_decay (float): multiplicative factor (per episode) for decreasing epsilon
"""
scores = [] # list containing scores from each episode
scores_window = deque(maxlen=100) # last 100 scores
eps = eps_start # initialize epsilon
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=True)[brain_name]
score = 0
for t in range(max_t):
state = get_state(env_info)
action = agent.act(state, eps)
next_env = env.step(action)[brain_name]
next_state, reward, done = extract_full_state(next_env)
agent.step(state, action, reward, next_state, done)
env_info = next_env
score += reward
if done:
break
scores_window.append(score) # save most recent score
scores.append(score) # save most recent score
eps = max(eps_end, eps_decay*eps) # decrease epsilon
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)), end="")
if i_episode % 100 == 0:
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)))
if np.mean(scores_window) >= 14.0:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, np.mean(scores_window)))
torch.save(agent.qnetwork_local.state_dict(), 'checkpoint.pth')
break
return scores
scores = dqn()
# plot the scores graph
import matplotlib.pyplot as plt
%matplotlib inline
def plot_scores(scores):
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
plot_scores(scores)
# load the training checkpoint
agent.qnetwork_local.load_state_dict(torch.load('checkpoint.pth'))
# watch it dance by running 1 episode
NUM_TESTS = 10
for _ in range(NUM_TESTS):
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
state = get_state(env_info) # get the current state
score = 0 # initialize the score
while True:
action = agent.act(state) # select an action
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = get_state(env_info) # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
score += reward # update the score
state = next_state # roll over the state to next time step
if done: # exit loop if episode finished
break
print("Score: {}".format(score))
env.close()
```
| github_jupyter |
# CycleGAN, Image-to-Image Translation
In this notebook, we're going to define and train a CycleGAN to read in an image from a set $X$ and transform it so that it looks as if it belongs in set $Y$. Specifically, we'll look at a set of images of [Yosemite national park](https://en.wikipedia.org/wiki/Yosemite_National_Park) taken either during the summer of winter. The seasons are our two domains!
>The objective will be to train generators that learn to transform an image from domain $X$ into an image that looks like it came from domain $Y$ (and vice versa).
Some examples of image data in both sets are pictured below.
<img src='notebook_images/XY_season_images.png' width=50% />
### Unpaired Training Data
These images do not come with labels, but CycleGANs give us a way to learn the mapping between one image domain and another using an **unsupervised** approach. A CycleGAN is designed for image-to-image translation and it learns from unpaired training data. This means that in order to train a generator to translate images from domain $X$ to domain $Y$, we do not have to have exact correspondences between individual images in those domains. For example, in [the paper that introduced CycleGANs](https://arxiv.org/abs/1703.10593), the authors are able to translate between images of horses and zebras, even though there are no images of a zebra in exactly the same position as a horse or with exactly the same background, etc. Thus, CycleGANs enable learning a mapping from one domain $X$ to another domain $Y$ without having to find perfectly-matched, training pairs!
<img src='notebook_images/horse2zebra.jpg' width=50% />
### CycleGAN and Notebook Structure
A CycleGAN is made of two types of networks: **discriminators, and generators**. In this example, the discriminators are responsible for classifying images as real or fake (for both $X$ and $Y$ kinds of images). The generators are responsible for generating convincing, fake images for both kinds of images.
This notebook will detail the steps you should take to define and train such a CycleGAN.
>1. You'll load in the image data using PyTorch's DataLoader class to efficiently read in images from a specified directory.
2. Then, you'll be tasked with defining the CycleGAN architecture according to provided specifications. You'll define the discriminator and the generator models.
3. You'll complete the training cycle by calculating the adversarial and cycle consistency losses for the generator and discriminator network and completing a number of training epochs. *It's suggested that you enable GPU usage for training.*
4. Finally, you'll evaluate your model by looking at the loss over time and looking at sample, generated images.
---
## Load and Visualize the Data
We'll first load in and visualize the training data, importing the necessary libraries to do so.
> If you are working locally, you'll need to download the data as a zip file by [clicking here](https://s3.amazonaws.com/video.udacity-data.com/topher/2018/November/5be66e78_summer2winter-yosemite/summer2winter-yosemite.zip).
It may be named `summer2winter-yosemite/` with a dash or an underscore, so take note, extract the data to your home directory and make sure the below `image_dir` matches. Then you can proceed with the following loading code.
```
# loading in and transforming data
import os
import torch
from torch.utils.data import DataLoader
import torchvision
import torchvision.datasets as datasets
import torchvision.transforms as transforms
# visualizing data
import matplotlib.pyplot as plt
import numpy as np
import warnings
%matplotlib inline
```
### DataLoaders
The `get_data_loader` function returns training and test DataLoaders that can load data efficiently and in specified batches. The function has the following parameters:
* `image_type`: `summer` or `winter`, the names of the directories where the X and Y images are stored
* `image_dir`: name of the main image directory, which holds all training and test images
* `image_size`: resized, square image dimension (all images will be resized to this dim)
* `batch_size`: number of images in one batch of data
The test data is strictly for feeding to our generators, later on, so we can visualize some generated samples on fixed, test data.
You can see that this function is also responsible for making sure our images are of the right, square size (128x128x3) and converted into Tensor image types.
**It's suggested that you use the default values of these parameters.**
Note: If you are trying this code on a different set of data, you may get better results with larger `image_size` and `batch_size` parameters. If you change the `batch_size`, make sure that you create complete batches in the training loop otherwise you may get an error when trying to save sample data.
```
def get_data_loader(image_type, image_dir='summer2winter-yosemite',
image_size=128, batch_size=16, num_workers=0):
"""Returns training and test data loaders for a given image type, either 'summer' or 'winter'.
These images will be resized to 128x128x3, by default, converted into Tensors, and normalized.
"""
# resize and normalize the images
transform = transforms.Compose([transforms.Resize(image_size), # resize to 128x128
transforms.ToTensor()])
# get training and test directories
image_path = './' + image_dir
train_path = os.path.join(image_path, image_type)
test_path = os.path.join(image_path, 'test_{}'.format(image_type))
# define datasets using ImageFolder
train_dataset = datasets.ImageFolder(train_path, transform)
test_dataset = datasets.ImageFolder(test_path, transform)
# create and return DataLoaders
train_loader = DataLoader(dataset=train_dataset, batch_size=batch_size, shuffle=True, num_workers=num_workers)
test_loader = DataLoader(dataset=test_dataset, batch_size=batch_size, shuffle=False, num_workers=num_workers)
return train_loader, test_loader
# Create train and test dataloaders for images from the two domains X and Y
# image_type = directory names for our data
dataloader_X, test_dataloader_X = get_data_loader(image_type='summer')
dataloader_Y, test_dataloader_Y = get_data_loader(image_type='winter')
```
## Display some Training Images
Below we provide a function `imshow` that reshape some given images and converts them to NumPy images so that they can be displayed by `plt`. This cell should display a grid that contains a batch of image data from set $X$.
```
# helper imshow function
def imshow(img):
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
# get some images from X
dataiter = iter(dataloader_X)
# the "_" is a placeholder for no labels
images, _ = dataiter.next()
# show images
fig = plt.figure(figsize=(12, 8))
imshow(torchvision.utils.make_grid(images))
```
Next, let's visualize a batch of images from set $Y$.
```
# get some images from Y
dataiter = iter(dataloader_Y)
images, _ = dataiter.next()
# show images
fig = plt.figure(figsize=(12,8))
imshow(torchvision.utils.make_grid(images))
```
### Pre-processing: scaling from -1 to 1
We need to do a bit of pre-processing; we know that the output of our `tanh` activated generator will contain pixel values in a range from -1 to 1, and so, we need to rescale our training images to a range of -1 to 1. (Right now, they are in a range from 0-1.)
```
# current range
img = images[0]
print('Min: ', img.min())
print('Max: ', img.max())
# helper scale function
def scale(x, feature_range=(-1, 1)):
''' Scale takes in an image x and returns that image, scaled
with a feature_range of pixel values from -1 to 1.
This function assumes that the input x is already scaled from 0-255.'''
# scale from 0-1 to feature_range
min, max = feature_range
x = x * (max - min) + min
return x
# scaled range
scaled_img = scale(img)
print('Scaled min: ', scaled_img.min())
print('Scaled max: ', scaled_img.max())
```
---
## Define the Model
A CycleGAN is made of two discriminator and two generator networks.
## Discriminators
The discriminators, $D_X$ and $D_Y$, in this CycleGAN are convolutional neural networks that see an image and attempt to classify it as real or fake. In this case, real is indicated by an output close to 1 and fake as close to 0. The discriminators have the following architecture:
<img src='notebook_images/discriminator_layers.png' width=80% />
This network sees a 128x128x3 image, and passes it through 5 convolutional layers that downsample the image by a factor of 2. The first four convolutional layers have a BatchNorm and ReLu activation function applied to their output, and the last acts as a classification layer that outputs one value.
### Convolutional Helper Function
To define the discriminators, you're expected to use the provided `conv` function, which creates a convolutional layer + an optional batch norm layer.
```
import torch.nn as nn
import torch.nn.functional as F
# helper conv function
def conv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a convolutional layer, with optional batch normalization.
"""
layers = []
conv_layer = nn.Conv2d(in_channels=in_channels, out_channels=out_channels,
kernel_size=kernel_size, stride=stride, padding=padding, bias=False)
layers.append(conv_layer)
if batch_norm:
layers.append(nn.BatchNorm2d(out_channels))
return nn.Sequential(*layers)
```
### Define the Discriminator Architecture
Your task is to fill in the `__init__` function with the specified 5 layer conv net architecture. Both $D_X$ and $D_Y$ have the same architecture, so we only need to define one class, and later instantiate two discriminators.
> It's recommended that you use a **kernel size of 4x4** and use that to determine the correct stride and padding size for each layer. [This Stanford resource](http://cs231n.github.io/convolutional-networks/#conv) may also help in determining stride and padding sizes.
* Define your convolutional layers in `__init__`
* Then fill in the forward behavior of the network
The `forward` function defines how an input image moves through the discriminator, and the most important thing is to pass it through your convolutional layers in order, with a **ReLu** activation function applied to all but the last layer.
You should **not** apply a sigmoid activation function to the output, here, and that is because we are planning on using a squared error loss for training. And you can read more about this loss function, later in the notebook.
```
class Discriminator(nn.Module):
def __init__(self, conv_dim=64):
super(Discriminator, self).__init__()
# Define all convolutional layers
# Should accept an RGB image as input and output a single value
def forward(self, x):
# define feedforward behavior
return x
```
## Generators
The generators, `G_XtoY` and `G_YtoX` (sometimes called F), are made of an **encoder**, a conv net that is responsible for turning an image into a smaller feature representation, and a **decoder**, a *transpose_conv* net that is responsible for turning that representation into an transformed image. These generators, one from XtoY and one from YtoX, have the following architecture:
<img src='notebook_images/cyclegan_generator_ex.png' width=90% />
This network sees a 128x128x3 image, compresses it into a feature representation as it goes through three convolutional layers and reaches a series of residual blocks. It goes through a few (typically 6 or more) of these residual blocks, then it goes through three transpose convolutional layers (sometimes called *de-conv* layers) which upsample the output of the resnet blocks and create a new image!
Note that most of the convolutional and transpose-convolutional layers have BatchNorm and ReLu functions applied to their outputs with the exception of the final transpose convolutional layer, which has a `tanh` activation function applied to the output. Also, the residual blocks are made of convolutional and batch normalization layers, which we'll go over in more detail, next.
---
### Residual Block Class
To define the generators, you're expected to define a `ResidualBlock` class which will help you connect the encoder and decoder portions of the generators. You might be wondering, what exactly is a Resnet block? It may sound familiar from something like ResNet50 for image classification, pictured below.
<img src='notebook_images/resnet_50.png' width=90%/>
ResNet blocks rely on connecting the output of one layer with the input of an earlier layer. The motivation for this structure is as follows: very deep neural networks can be difficult to train. Deeper networks are more likely to have vanishing or exploding gradients and, therefore, have trouble reaching convergence; batch normalization helps with this a bit. However, during training, we often see that deep networks respond with a kind of training degradation. Essentially, the training accuracy stops improving and gets saturated at some point during training. In the worst cases, deep models would see their training accuracy actually worsen over time!
One solution to this problem is to use **Resnet blocks** that allow us to learn so-called *residual functions* as they are applied to layer inputs. You can read more about this proposed architecture in the paper, [Deep Residual Learning for Image Recognition](https://arxiv.org/pdf/1512.03385.pdf) by Kaiming He et. al, and the below image is from that paper.
<img src='notebook_images/resnet_block.png' width=40%/>
### Residual Functions
Usually, when we create a deep learning model, the model (several layers with activations applied) is responsible for learning a mapping, `M`, from an input `x` to an output `y`.
>`M(x) = y` (Equation 1)
Instead of learning a direct mapping from `x` to `y`, we can instead define a **residual function**
> `F(x) = M(x) - x`
This looks at the difference between a mapping applied to x and the original input, x. `F(x)` is, typically, two convolutional layers + normalization layer and a ReLu in between. These convolutional layers should have the same number of inputs as outputs. This mapping can then be written as the following; a function of the residual function and the input x. The addition step creates a kind of loop that connects the input x to the output, y:
>`M(x) = F(x) + x` (Equation 2) or
>`y = F(x) + x` (Equation 3)
#### Optimizing a Residual Function
The idea is that it is easier to optimize this residual function `F(x)` than it is to optimize the original mapping `M(x)`. Consider an example; what if we want `y = x`?
From our first, direct mapping equation, **Equation 1**, we could set `M(x) = x` but it is easier to solve the residual equation `F(x) = 0`, which, when plugged in to **Equation 3**, yields `y = x`.
### Defining the `ResidualBlock` Class
To define the `ResidualBlock` class, we'll define residual functions (a series of layers), apply them to an input x and add them to that same input. This is defined just like any other neural network, with an `__init__` function and the addition step in the `forward` function.
In our case, you'll want to define the residual block as:
* Two convolutional layers with the same size input and output
* Batch normalization applied to the outputs of the convolutional layers
* A ReLu function on the output of the *first* convolutional layer
Then, in the `forward` function, add the input x to this residual block. Feel free to use the helper `conv` function from above to create this block.
```
# residual block class
class ResidualBlock(nn.Module):
"""Defines a residual block.
This adds an input x to a convolutional layer (applied to x) with the same size input and output.
These blocks allow a model to learn an effective transformation from one domain to another.
"""
def __init__(self, conv_dim):
super(ResidualBlock, self).__init__()
# conv_dim = number of inputs
# define two convolutional layers + batch normalization that will act as our residual function, F(x)
# layers should have the same shape input as output; I suggest a kernel_size of 3
def forward(self, x):
# apply a ReLu activation the outputs of the first layer
# return a summed output, x + resnet_block(x)
return x
```
### Transpose Convolutional Helper Function
To define the generators, you're expected to use the above `conv` function, `ResidualBlock` class, and the below `deconv` helper function, which creates a transpose convolutional layer + an optional batchnorm layer.
```
# helper deconv function
def deconv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a transpose convolutional layer, with optional batch normalization.
"""
layers = []
# append transpose conv layer
layers.append(nn.ConvTranspose2d(in_channels, out_channels, kernel_size, stride, padding, bias=False))
# optional batch norm layer
if batch_norm:
layers.append(nn.BatchNorm2d(out_channels))
return nn.Sequential(*layers)
```
---
## Define the Generator Architecture
* Complete the `__init__` function with the specified 3 layer **encoder** convolutional net, a series of residual blocks (the number of which is given by `n_res_blocks`), and then a 3 layer **decoder** transpose convolutional net.
* Then complete the `forward` function to define the forward behavior of the generators. Recall that the last layer has a `tanh` activation function.
Both $G_{XtoY}$ and $G_{YtoX}$ have the same architecture, so we only need to define one class, and later instantiate two generators.
```
class CycleGenerator(nn.Module):
def __init__(self, conv_dim=64, n_res_blocks=6):
super(CycleGenerator, self).__init__()
# 1. Define the encoder part of the generator
# 2. Define the resnet part of the generator
# 3. Define the decoder part of the generator
def forward(self, x):
"""Given an image x, returns a transformed image."""
# define feedforward behavior, applying activations as necessary
return x
```
---
## Create the complete network
Using the classes you defined earlier, you can define the discriminators and generators necessary to create a complete CycleGAN. The given parameters should work for training.
First, create two discriminators, one for checking if $X$ sample images are real, and one for checking if $Y$ sample images are real. Then the generators. Instantiate two of them, one for transforming a painting into a realistic photo and one for transforming a photo into into a painting.
```
def create_model(g_conv_dim=64, d_conv_dim=64, n_res_blocks=6):
"""Builds the generators and discriminators."""
# Instantiate generators
G_XtoY =
G_YtoX =
# Instantiate discriminators
D_X =
D_Y =
# move models to GPU, if available
if torch.cuda.is_available():
device = torch.device("cuda:0")
G_XtoY.to(device)
G_YtoX.to(device)
D_X.to(device)
D_Y.to(device)
print('Models moved to GPU.')
else:
print('Only CPU available.')
return G_XtoY, G_YtoX, D_X, D_Y
# call the function to get models
G_XtoY, G_YtoX, D_X, D_Y = create_model()
```
## Check that you've implemented this correctly
The function `create_model` should return the two generator and two discriminator networks. After you've defined these discriminator and generator components, it's good practice to check your work. The easiest way to do this is to print out your model architecture and read through it to make sure the parameters are what you expected. The next cell will print out their architectures.
```
# helper function for printing the model architecture
def print_models(G_XtoY, G_YtoX, D_X, D_Y):
"""Prints model information for the generators and discriminators.
"""
print(" G_XtoY ")
print("-----------------------------------------------")
print(G_XtoY)
print()
print(" G_YtoX ")
print("-----------------------------------------------")
print(G_YtoX)
print()
print(" D_X ")
print("-----------------------------------------------")
print(D_X)
print()
print(" D_Y ")
print("-----------------------------------------------")
print(D_Y)
print()
# print all of the models
print_models(G_XtoY, G_YtoX, D_X, D_Y)
```
## Discriminator and Generator Losses
Computing the discriminator and the generator losses are key to getting a CycleGAN to train.
<img src='notebook_images/CycleGAN_loss.png' width=90% height=90% />
**Image from [original paper](https://arxiv.org/abs/1703.10593) by Jun-Yan Zhu et. al.**
* The CycleGAN contains two mapping functions $G: X \rightarrow Y$ and $F: Y \rightarrow X$, and associated adversarial discriminators $D_Y$ and $D_X$. **(a)** $D_Y$ encourages $G$ to translate $X$ into outputs indistinguishable from domain $Y$, and vice versa for $D_X$ and $F$.
* To further regularize the mappings, we introduce two cycle consistency losses that capture the intuition that if
we translate from one domain to the other and back again we should arrive at where we started. **(b)** Forward cycle-consistency loss and **(c)** backward cycle-consistency loss.
## Least Squares GANs
We've seen that regular GANs treat the discriminator as a classifier with the sigmoid cross entropy loss function. However, this loss function may lead to the vanishing gradients problem during the learning process. To overcome such a problem, we'll use a least squares loss function for the discriminator. This structure is also referred to as a least squares GAN or LSGAN, and you can [read the original paper on LSGANs, here](https://arxiv.org/pdf/1611.04076.pdf). The authors show that LSGANs are able to generate higher quality images than regular GANs and that this loss type is a bit more stable during training!
### Discriminator Losses
The discriminator losses will be mean squared errors between the output of the discriminator, given an image, and the target value, 0 or 1, depending on whether it should classify that image as fake or real. For example, for a *real* image, `x`, we can train $D_X$ by looking at how close it is to recognizing and image `x` as real using the mean squared error:
```
out_x = D_X(x)
real_err = torch.mean((out_x-1)**2)
```
### Generator Losses
Calculating the generator losses will look somewhat similar to calculating the discriminator loss; there will still be steps in which you generate fake images that look like they belong to the set of $X$ images but are based on real images in set $Y$, and vice versa. You'll compute the "real loss" on those generated images by looking at the output of the discriminator as it's applied to these _fake_ images; this time, your generator aims to make the discriminator classify these fake images as *real* images.
#### Cycle Consistency Loss
In addition to the adversarial losses, the generator loss terms will also include the **cycle consistency loss**. This loss is a measure of how good a reconstructed image is, when compared to an original image.
Say you have a fake, generated image, `x_hat`, and a real image, `y`. You can get a reconstructed `y_hat` by applying `G_XtoY(x_hat) = y_hat` and then check to see if this reconstruction `y_hat` and the orginal image `y` match. For this, we recommed calculating the L1 loss, which is an absolute difference, between reconstructed and real images. You may also choose to multiply this loss by some weight value `lambda_weight` to convey its importance.
<img src='notebook_images/reconstruction_error.png' width=40% height=40% />
The total generator loss will be the sum of the generator losses and the forward and backward cycle consistency losses.
---
### Define Loss Functions
To help us calculate the discriminator and gnerator losses during training, let's define some helpful loss functions. Here, we'll define three.
1. `real_mse_loss` that looks at the output of a discriminator and returns the error based on how close that output is to being classified as real. This should be a mean squared error.
2. `fake_mse_loss` that looks at the output of a discriminator and returns the error based on how close that output is to being classified as fake. This should be a mean squared error.
3. `cycle_consistency_loss` that looks at a set of real image and a set of reconstructed/generated images, and returns the mean absolute error between them. This has a `lambda_weight` parameter that will weight the mean absolute error in a batch.
It's recommended that you take a [look at the original, CycleGAN paper](https://arxiv.org/pdf/1703.10593.pdf) to get a starting value for `lambda_weight`.
```
def real_mse_loss(D_out):
# how close is the produced output from being "real"?
def fake_mse_loss(D_out):
# how close is the produced output from being "false"?
def cycle_consistency_loss(real_im, reconstructed_im, lambda_weight):
# calculate reconstruction loss
# return weighted loss
```
### Define the Optimizers
Next, let's define how this model will update its weights. This, like the GANs you may have seen before, uses [Adam](https://pytorch.org/docs/stable/optim.html#algorithms) optimizers for the discriminator and generator. It's again recommended that you take a [look at the original, CycleGAN paper](https://arxiv.org/pdf/1703.10593.pdf) to get starting hyperparameter values.
```
import torch.optim as optim
# hyperparams for Adam optimizers
lr=
beta1=
beta2=
g_params = list(G_XtoY.parameters()) + list(G_YtoX.parameters()) # Get generator parameters
# Create optimizers for the generators and discriminators
g_optimizer = optim.Adam(g_params, lr, [beta1, beta2])
d_x_optimizer = optim.Adam(D_X.parameters(), lr, [beta1, beta2])
d_y_optimizer = optim.Adam(D_Y.parameters(), lr, [beta1, beta2])
```
---
## Training a CycleGAN
When a CycleGAN trains, and sees one batch of real images from set $X$ and $Y$, it trains by performing the following steps:
**Training the Discriminators**
1. Compute the discriminator $D_X$ loss on real images
2. Generate fake images that look like domain $X$ based on real images in domain $Y$
3. Compute the fake loss for $D_X$
4. Compute the total loss and perform backpropagation and $D_X$ optimization
5. Repeat steps 1-4 only with $D_Y$ and your domains switched!
**Training the Generators**
1. Generate fake images that look like domain $X$ based on real images in domain $Y$
2. Compute the generator loss based on how $D_X$ responds to fake $X$
3. Generate *reconstructed* $\hat{Y}$ images based on the fake $X$ images generated in step 1
4. Compute the cycle consistency loss by comparing the reconstructions with real $Y$ images
5. Repeat steps 1-4 only swapping domains
6. Add up all the generator and reconstruction losses and perform backpropagation + optimization
<img src='notebook_images/cycle_consistency_ex.png' width=70% />
### Saving Your Progress
A CycleGAN repeats its training process, alternating between training the discriminators and the generators, for a specified number of training iterations. You've been given code that will save some example generated images that the CycleGAN has learned to generate after a certain number of training iterations. Along with looking at the losses, these example generations should give you an idea of how well your network has trained.
Below, you may choose to keep all default parameters; your only task is to calculate the appropriate losses and complete the training cycle.
```
# import save code
from helpers import save_samples, checkpoint
# train the network
def training_loop(dataloader_X, dataloader_Y, test_dataloader_X, test_dataloader_Y,
n_epochs=1000):
print_every=10
# keep track of losses over time
losses = []
test_iter_X = iter(test_dataloader_X)
test_iter_Y = iter(test_dataloader_Y)
# Get some fixed data from domains X and Y for sampling. These are images that are held
# constant throughout training, that allow us to inspect the model's performance.
fixed_X = test_iter_X.next()[0]
fixed_Y = test_iter_Y.next()[0]
fixed_X = scale(fixed_X) # make sure to scale to a range -1 to 1
fixed_Y = scale(fixed_Y)
# batches per epoch
iter_X = iter(dataloader_X)
iter_Y = iter(dataloader_Y)
batches_per_epoch = min(len(iter_X), len(iter_Y))
for epoch in range(1, n_epochs+1):
# Reset iterators for each epoch
if epoch % batches_per_epoch == 0:
iter_X = iter(dataloader_X)
iter_Y = iter(dataloader_Y)
images_X, _ = iter_X.next()
images_X = scale(images_X) # make sure to scale to a range -1 to 1
images_Y, _ = iter_Y.next()
images_Y = scale(images_Y)
# move images to GPU if available (otherwise stay on CPU)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
images_X = images_X.to(device)
images_Y = images_Y.to(device)
# ============================================
# TRAIN THE DISCRIMINATORS
# ============================================
## First: D_X, real and fake loss components ##
# 1. Compute the discriminator losses on real images
# 2. Generate fake images that look like domain X based on real images in domain Y
# 3. Compute the fake loss for D_X
# 4. Compute the total loss and perform backprop
d_x_loss =
## Second: D_Y, real and fake loss components ##
d_y_loss =
# =========================================
# TRAIN THE GENERATORS
# =========================================
## First: generate fake X images and reconstructed Y images ##
# 1. Generate fake images that look like domain X based on real images in domain Y
# 2. Compute the generator loss based on domain X
# 3. Create a reconstructed y
# 4. Compute the cycle consistency loss (the reconstruction loss)
## Second: generate fake Y images and reconstructed X images ##
# 5. Add up all generator and reconstructed losses and perform backprop
g_total_loss =
# Print the log info
if epoch % print_every == 0:
# append real and fake discriminator losses and the generator loss
losses.append((d_x_loss.item(), d_y_loss.item(), g_total_loss.item()))
print('Epoch [{:5d}/{:5d}] | d_X_loss: {:6.4f} | d_Y_loss: {:6.4f} | g_total_loss: {:6.4f}'.format(
epoch, n_epochs, d_x_loss.item(), d_y_loss.item(), g_total_loss.item()))
sample_every=100
# Save the generated samples
if epoch % sample_every == 0:
G_YtoX.eval() # set generators to eval mode for sample generation
G_XtoY.eval()
save_samples(epoch, fixed_Y, fixed_X, G_YtoX, G_XtoY, batch_size=16)
G_YtoX.train()
G_XtoY.train()
# uncomment these lines, if you want to save your model
# checkpoint_every=1000
# # Save the model parameters
# if epoch % checkpoint_every == 0:
# checkpoint(epoch, G_XtoY, G_YtoX, D_X, D_Y)
return losses
n_epochs = 1000 # keep this small when testing if a model first works, then increase it to >=1000
losses = training_loop(dataloader_X, dataloader_Y, test_dataloader_X, test_dataloader_Y, n_epochs=n_epochs)
```
## Tips on Training and Loss Patterns
A lot of experimentation goes into finding the best hyperparameters such that the generators and discriminators don't overpower each other. It's often a good starting point to look at existing papers to find what has worked in previous experiments, I'd recommend this [DCGAN paper](https://arxiv.org/pdf/1511.06434.pdf) in addition to the original [CycleGAN paper](https://arxiv.org/pdf/1703.10593.pdf) to see what worked for them. Then, you can try your own experiments based off of a good foundation.
#### Discriminator Losses
When you display the generator and discriminator losses you should see that there is always some discriminator loss; recall that we are trying to design a model that can generate good "fake" images. So, the ideal discriminator will not be able to tell the difference between real and fake images and, as such, will always have some loss. You should also see that $D_X$ and $D_Y$ are roughly at the same loss levels; if they are not, this indicates that your training is favoring one type of discriminator over the and you may need to look at biases in your models or data.
#### Generator Loss
The generator's loss should start significantly higher than the discriminator losses because it is accounting for the loss of both generators *and* weighted reconstruction errors. You should see this loss decrease a lot at the start of training because initial, generated images are often far-off from being good fakes. After some time it may level off; this is normal since the generator and discriminator are both improving as they train. If you see that the loss is jumping around a lot, over time, you may want to try decreasing your learning rates or changing your cycle consistency loss to be a little more/less weighted.
```
fig, ax = plt.subplots(figsize=(12,8))
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator, X', alpha=0.5)
plt.plot(losses.T[1], label='Discriminator, Y', alpha=0.5)
plt.plot(losses.T[2], label='Generators', alpha=0.5)
plt.title("Training Losses")
plt.legend()
```
---
## Evaluate the Result!
As you trained this model, you may have chosen to sample and save the results of your generated images after a certain number of training iterations. This gives you a way to see whether or not your Generators are creating *good* fake images. For example, the image below depicts real images in the $Y$ set, and the corresponding generated images during different points in the training process. You can see that the generator starts out creating very noisy, fake images, but begins to converge to better representations as it trains (though, not perfect).
<img src='notebook_images/sample-004000-summer2winter.png' width=50% />
Below, you've been given a helper function for displaying generated samples based on the passed in training iteration.
```
import matplotlib.image as mpimg
# helper visualization code
def view_samples(iteration, sample_dir='samples_cyclegan'):
# samples are named by iteration
path_XtoY = os.path.join(sample_dir, 'sample-{:06d}-X-Y.png'.format(iteration))
path_YtoX = os.path.join(sample_dir, 'sample-{:06d}-Y-X.png'.format(iteration))
# read in those samples
try:
x2y = mpimg.imread(path_XtoY)
y2x = mpimg.imread(path_YtoX)
except:
print('Invalid number of iterations.')
fig, (ax1, ax2) = plt.subplots(figsize=(18,20), nrows=2, ncols=1, sharey=True, sharex=True)
ax1.imshow(x2y)
ax1.set_title('X to Y')
ax2.imshow(y2x)
ax2.set_title('Y to X')
# view samples at iteration 100
view_samples(100, 'samples_cyclegan')
# view samples at iteration 1000
view_samples(1000, 'samples_cyclegan')
```
---
## Further Challenges and Directions
* One shortcoming of this model is that it produces fairly low-resolution images; this is an ongoing area of research; you can read about a higher-resolution formulation that uses a multi-scale generator model, in [this paper](https://arxiv.org/abs/1711.11585).
* Relatedly, we may want to process these as larger (say 256x256) images at first, to take advantage of high-res data.
* It may help your model to converge faster, if you initialize the weights in your network.
* This model struggles with matching colors exactly. This is because, if $G_{YtoX}$ and $G_{XtoY}$ may change the tint of an image; the cycle consistency loss may not be affected and can still be small. You could choose to introduce a new, color-based loss term that compares $G_{YtoX}(y)$ and $y$, and $G_{XtoY}(x)$ and $x$, but then this becomes a supervised learning approach.
* This unsupervised approach also struggles with geometric changes, like changing the apparent size of individual object in an image, so it is best suited for stylistic transformations.
* For creating different kinds of models or trying out the Pix2Pix Architecture, [this Github repository](https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/) which implements CycleGAN *and* Pix2Pix in PyTorch is a great resource.
**Once you are satified with your model, you are ancouraged to test it on a different dataset to see if it can find different types of mappings!**
---
### Different datasets for download
You can download a variety of datasets used in the Pix2Pix and CycleGAN papers, by following instructions in the [associated Github repository](https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/blob/master/README.md). You'll just need to make sure that the data directories are named and organized correctly to load in that data.
| github_jupyter |
## Problem 1: Visualize a static map
```
import glob
import numpy as np
import pysal as ps
import pandas as pd
import geopandas as gpd
import contextily as ctx
from geopandas.tools import geocode
import matplotlib.pyplot as plt
%matplotlib inline
# reading in the addresses of the six shopping centers and geocoding them
addresses = pd.read_csv('data/addresses.txt', sep=';')
addresses = geocode(addresses['address'], provider = 'nominatim', user_agent = 'autogis_student_er')
# reading in the TravelTime filepaths and MetropAccess shape file
filepaths = glob.glob('data/Travel*.txt')
grid = gpd.read_file('data/MetropAccess_YKR_grid_EurefFIN.shp')
# adding the car and pt data in the TravelTime files to the grid dataframe
for idx, fp in enumerate(filepaths):
data = pd.read_csv(fp, sep=';')
data = data[['from_id', 'pt_r_t', 'car_r_t']]
data.rename(columns={'pt_r_t': ('pt_r_t_' + str(idx))}, inplace=True)
data.rename(columns={'car_r_t': ('car_r_t_' + str(idx))}, inplace=True)
grid = grid.merge(data, left_on='YKR_ID', right_on='from_id')
# reporojecting the two dataframes
addresses = addresses.to_crs(epsg=3857)
grid = grid.to_crs(epsg=3857)
# calculating the travel time by pt to the closest shopping center for each grid cell
cols_pt = ['pt_r_t_'+str(i) for i in range(0,7)]
grid['pt_min'] = grid[cols_pt].min(1)
# calculating the travel time by car to the closest shopping center for each grid cell
cols_car = ['car_r_t_'+str(i) for i in range(0,7)]
grid['car_min'] = grid[cols_car].min(1)
# creating a new dataframe that only has data for the car_min, pt_min, and geometry columns,
# ignoring all columns where car_min or pt_min is -1 (no data)
cols_subset = ['car_min', 'pt_min', 'geometry']
travel_times = grid[cols_subset].loc[np.where((grid['pt_min'] != -1) & (grid['car_min'] != -1))]
# creating a new column in the travel_times dataframe that shows the difference between travel
# times by car and pt to each gridcell's closest shopping center
travel_times['car_fast'] = travel_times['pt_min'] - travel_times['car_min']
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(10, 10));
# plotting the minutes saved from travelling by car (in comparison to travelling by pt) to
# the nearest shopping center
travel_times.plot(column="car_fast", legend=True, cmap='Reds', linewidth=0, alpha=0.75,
scheme='quantiles', k=8, ax=ax);
# adding a basemap
ctx.add_basemap(ax, url=ctx.tile_providers.ST_TONER_LITE);
# plotting the locations of the shopping centers
addresses['geometry'].plot(ax=ax, markersize=60);
# adding a title to the map
ax.set_title('Travel Times to Nearest Shopping Center\nMinutes Saved When Travelling by Car, in Comparison to Public Transport');
# setting the bounds for the map
ax.set_xbound(2725000, 2815000);
ax.set_ybound(8420000, 8490000);
fig.tight_layout()
fig.savefig('docs/Shopping_centers_accessibility_car_pt_comparisons.png')
```
It seems that as long as one lives close to Helsinki's main trasport axes, one's nearest shopping center is about as far away by car as by public transport. In a few cases public transport is even faster. However, in the not so well connected parts of Helsinki (especially the north-west) travelling by car is much faster than by public transport.
| github_jupyter |
# Gaussian elimination and LU decomposition
Say we want to compute the solution of
$$Ax = b$$
for the vector $x$. We learn how to do this by transforming it to the problem of solving
$$U x = y$$
where $U$ is an upper-triangular matrix obtained by performing Gaussian elimiantion on $A$ and $y$ is obtained by performing the same operations on $b$. We can then use back substitution to solve $Ux=y$ more easily than solving $Ax=b$ directly.
This approach is directly related to the LU decomposition of a matrix, where we wish to factor a matrix $A$ into a product of a lower triangular matrix $L$ and an upper triangular matrix $U$ to give $A = LU$. To understand how to compute the LU decomposition of a matrix, let us start by reminding ourselves of how to do Gaussian elimination.
## Gaussian elimination by hand
To start, consider the following 3x3 matrix
$$ A = \begin{bmatrix} 1 & 2 & 3 \\ 4 & 5 & 6 \\ 7 & 8 & 10 \end{bmatrix}$$
1. Use Gaussian elimination to transform this by hand to an upper triangular matrix $U$ (in row echelon form). Record each elementary row operation you perform along the way.
2. Apply the same sequence of row operations to the vector
$$b = \begin{bmatrix} 1 \\ 2 \\ 3 \end{bmatrix}$$
to obtain the transformed vector $y$.
3. Use back substitution to solve $U x = y$.
### Solution
Using the standard Gaussian elimination algorithm, we would perform the following steps:
1. Subtract 4x(row 1) from row 2. This leaves us with $$\begin{bmatrix} 1 & 2 & 3 \\ 0 & -3 & -6 \\ 7 & 8 & 10 \end{bmatrix}$$
2. Subtract 7x(row 1) from row 3. This leaves us with $$\begin{bmatrix} 1 & 2 & 3 \\ 0 & -3 & -6 \\ 0 & -6 & -11 \end{bmatrix}$$
3. Subtract 2x(row 2) from row 3. This leaves us with $$\begin{bmatrix} 1 & 2 & 3 \\ 0 & -3 & -6 \\ 0 & 0 & 1 \end{bmatrix}$$
We now have an upper-triangular matrix $U$. Applying the same sequence of operations to $b$:
1. Subtract 4x(row 1) from row 2. This leaves us with $$\begin{bmatrix} 1 \\ -2 \\ 3\end{bmatrix}$$
2. Subtract 7x(row 1) from row 3. This leaves us with $$\begin{bmatrix} 1 \\ -2 \\ -4 \end{bmatrix}$$
3. Subtract 2x(row 2) from row 3. This leaves us with $$\begin{bmatrix} 1 \\ -2 \\ 0 \end{bmatrix}$$
Finally, we use backsubstitution to solve $Ux = y$ for x. Starting with the last entry
$$ x_n = 0 / 1 = 0$$
$$ x_{n-1} = \frac{-2 - (-6)(0)}{-3} = \frac23$$
$$ x_{n-2} = \frac{1 - (2)(\frac23) - (3)(0)}{1} = -\frac13$$
so we have the solution
$$x = \begin{bmatrix} -\frac13 \\ \frac23 \\ 0\end{bmatrix}$$
## Gaussian elimination in Python
We will now transform the previous algorithm into Python code. First of all we define the matrix $A$ and the vector $b$.
```
import numpy as np
A = np.array([[1,2,3],[4,5,6],[7,8,10]])
b = np.array([1,2,3])
n = 3
```
Now perform Gaussian elimination and store the result in a matrix $U$ and a vector $y$. We keep track of the multiplication factors for each step in a matrix $L$.
```
U = np.array(A)
y = np.array(b)
L = np.identity(n)
# loop over columns and transform to get zeros below the pivot
for k in range(0,n):
# loop over all rows below the pivot
for i in range(k + 1, n):
# Store the multiplication factors in the matrix L
L[i,k] = U[i,k] / U[k,k]
# Subtract a multiple of row k from row i
# for j in range(k, n):
# U[i,j] = U[i,j] - L[i,k] * U[k,j]
U[i,:] = U[i,:] - L[i,k] * U[k,:]
y[i] = y[i] - L[i,k] * y[k]
U
y
```
If we consider how many operations this took, there are: ($n$ iterations of the outer loop) x ($n-(k+1)$ iterations of the inner loop) x (n multiplications for the subtraction). This means we require $\mathcal{O}(n^3)$ operations for the Gaussian elimination step.
Let us now solve for $x$ using back substitution on $U x = y$.
```
x = np.zeros(U.shape[1])
# Start with the last entry in x
x[n-1] = y[n-1] / U[n-1,n-1]
# Iterate upwards from the second last entry to the first entry
for i in range(n-2,-1,-1):
# Subtract all of the previously computed values from y, then divide by U[i,i]
#sum = 0
#for j in range(i+1,n):
# sum += U[i,j] * x[j]
x[i] = (y[i] - U[i,i+1:n]@x[i+1:n])/U[i,i]
x
```
We can check that our original matrix is given by $A=LU$:
```
L@U
```
## Gaussian elimination by matrix multiplication
We could consider each of the steps in Gaussian elimination in terms of multiplication on the left by a sequence of *elementary elimination matrices*. These come in three forms:
1. Multiplying row $i$ by a scalar $c$: $\mathbf{r}_i \to c\, \mathbf{r}_i$. This is equivalent to pre-multiplying by a matrix with $1$'s along the diagonal and c in the $i$-th diagonal,$$E_1(i, c) = \begin{bmatrix}
1 & & & & & & \\
& \ddots & & & & & \\
& & 1 & & & & \\
& & & c & & & \\
& & & & 1 & & \\
& & & & & \ddots & \\
& & & & & & 1
\end{bmatrix}$$
Note that the inverse is given by $E_1(c)^{-1} = E_1(c^{-1})$.
2. Add a multiple $c$ of row $j$ to row $i$: $\mathbf{r}_i \to \mathbf{r}_i + c\, \mathbf{r}_j$. This is equivalent to premultiplying by a matrix with $1$'s along the diagonal and $c$ in $(i, j)$-th entry:
$$E_2(i,j,c) = \begin{bmatrix}
1 & & & & & & \\
& \ddots & & & & & \\
& & 1 & & & & \\
& & & \ddots & & & \\
& & c & & 1 & & \\
& & & & & \ddots & \\
& & & & & & 1
\end{bmatrix}$$
In this case the inverse is given by $E_2(c)^{-1} = E_2(-c)$.
3. Interchanging rows $i$ and $j$: $\mathbf{r}_i \leftrightarrow \mathbf{r}_j$. This is equivalent to pre-multiplying by a matrix which is the identity with rows $i$ and $j$ swapped: $$E_3(i,j) = \begin{bmatrix}
1 & & & & & & \\
& \ddots & & & & & \\
& & 0 & & 1 & & \\
& & & \ddots & & & \\
& & 1 & & 0 & & \\
& & & & & \ddots & \\
& & & & & & 1
\end{bmatrix}$$
In this case the $E_3$ is a permutation matrix and it is its own inverse $E_3^{-1} = E_3$.
Let's work out the sequence of elimination matrices we need to perform the Gaussian elimination from the previous example. First, we define Python functions produce each of the three types of elimination matrix:
```
def E1(i, c):
e1 = np.identity(n)
e1[i, i] = c
return e1
def E2(i, j, c):
e2 = np.identity(n)
e2[i, j] = c
return e2
def E3(i, j):
e3 = np.identity(n)
e3[i, i] = 0
e3[j, j] = 0
e3[i, j] = 1
e3[j, i] = 1
return e3
```
Now, we can see that the Gaussian elimination steps correspond to
$$ U = E_2(2,1,-2) E_2(2,0,-7) E_2(1,0,-4) A$$
```
E2(1,0,-4)@A
E2(2,0,-7)@E2(1,0,-4)@A
E2(2,1,-2)@E2(2,0,-7)@E2(1,0,-4)@A
```
We therefore have
$$
\begin{aligned}
A &= [E_2(2,1,-2) E_2(2,0,-7) E_2(1,0,-4)]^{-1} U \\
&= E_2(1,0,-4)^{-1} E_2(2,0,-7)^{-1} E_2(2,1,-2)^{-1} U \\
&= E_2(1,0,4) E_2(2,0,7) E_2(2,1,2) U \\
&= L U
\end{aligned}
$$
so we have $L$ in terms of elementry elimination matrices.
```
E2(1,0,4)@E2(2,0,7)@E2(2,1,2)
L
```
## LU decomposition and rank-1 matrices
In the lecture videos we emphasized the idea of matrix multiplication in terms of columns-times-rows and the related idea of breaking a matrix into a sum of rank-1 matrices. Now, let's see how this gives a different way of looking at the LU decomposition.
The idea is that we would like to split $A$ into a rank-1 piece that picks out the first row and first column, plus a rank-1 piece that picks out the next row and column, and so on:
$$
\begin{aligned}
A = \begin{bmatrix} 1 & 2 & 3 \\ 4 & 5 & 6 \\ 7 & 8 & 10 \end{bmatrix}
&= \begin{bmatrix} 1 & 2 & 3 \\ 4 & \_ & \_ \\ 7 & \_ & \_ \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & \_ & \_ \\ 0 & \_ & \_ \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & 0 & 0 \\ 0 & 0 & \_ \end{bmatrix}
\end{aligned}
$$
We can fill in all the blanks here by insisting that each term is rank-1 and that we recover $A$.
Doing so, we get
$$
\begin{aligned}
A = \begin{bmatrix} 1 & 2 & 3 \\ 4 & 5 & 6 \\ 7 & 8 & 10 \end{bmatrix}
&= \begin{bmatrix} 1 & 2 & 3 \\ 4 & \_ & \_ \\ 7 & \_ & \_ \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & \_ & \_ \\ 0 & \_ & \_ \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & 0 & 0 \\ 0 & 0 & \_ \end{bmatrix}\\
&= \begin{bmatrix} 1 & 2 & 3 \\ 4 & 8 & 12 \\ 7 & 14 & 21 \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & \_ & \_ \\ 0 & \_ & \_ \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & 0 & 0 \\ 0 & 0 & \_ \end{bmatrix} \quad \text{(rank-1)}\\
&= \begin{bmatrix} 1 & 2 & 3 \\ 4 & 8 & 12 \\ 7 & 14 & 21 \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & -3 & -6 \\ 0 & -6 & \_ \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & 0 & 0 \\ 0 & 0 & \_ \end{bmatrix} \quad \text{(=$A$)}\\
&= \begin{bmatrix} 1 & 2 & 3 \\ 4 & 8 & 12 \\ 7 & 14 & 21 \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & -3 & -6 \\ 0 & -6 & -12 \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & 0 & 0 \\ 0 & 0 & \_ \end{bmatrix} \quad \text{(rank-1)}\\
&= \begin{bmatrix} 1 & 2 & 3 \\ 4 & 8 & 12 \\ 7 & 14 & 21 \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & -3 & -6 \\ 0 & -6 & -12 \end{bmatrix}
+ \begin{bmatrix} 0 & 0 & 0 \\ 0 & 0 & 0 \\ 0 & 0 & 1 \end{bmatrix} \quad \text{(=$A$)} \\
&= \begin{bmatrix} 1 \\ 4 \\ 7 \end{bmatrix} \begin{bmatrix} 1 & 2 & 3 \end{bmatrix}
+ \begin{bmatrix} 0 \\ 1 \\ 2 \end{bmatrix} \begin{bmatrix} 0 & -3 & -6 \end{bmatrix}
+ \begin{bmatrix} 0 \\ 0 \\ 1 \end{bmatrix} \begin{bmatrix} 0 & 0 & 1 \end{bmatrix} \\
&= l_1 u_1{}^T + l_2 u_2{}^T + l_3 u_3{}^T = LU
\end{aligned}
$$
```
l1 = L[:,0:1]
u1T = U[0:1]
l2 = L[:,1:2]
u2T = U[1:2]
l3 = L[:,2:3]
u3T = U[2:3]
l1@u1T
l2@u2T
l3@u3T
l1@u1T + l2@u2T + l3@u3T
```
| github_jupyter |
First, let's load all the necessary libraries:
```
library(openxlsx)
library(ggplot2)
library(ggrepel)
library(foreach)
```
Loading the functions:
```
source('functions.R')
source("visualizations.R")
```
Next we'll read the peak information and intensities from the provided Excel file. The file contains 538 features from a LC-MS experiment with HILIC column and negative ionization.
```
X <- read.xlsx("HILIC_NEG_TEST.xlsx", sheet = 1)
P <- read.xlsx("HILIC_NEG_TEST.xlsx", sheet = 2)
# compute the median peak area for each feature, used in visualizations
P$MPA <- sapply(X[P$Name], median, na.rm = TRUE)
```
This function finds connections between features according to set conditions: correlation threshhold and retention time window. If two features have a Pearson correlation coefficient above the set threshold and their retention times differ less than the set window, they are deemed connected.
The function returns a data frame with one row per connection, with the following columns:
- x & y: the names of the features
- cor: the pearson correlation coefficient
- mz_diff: the mass difference between the features
- rt_diff: the retention time difference between the features
```
conn <- find_connections(data = X, features = P, corr_thresh = 0.85, rt_window = 1/60, name_col = "Name", mz_col = "mz", rt_col = "rt")
```
The next part is where the actual ~~magic~~ science happens. The algorithm first constructs a network of the connected features, then starts stripping down the connected components of the network until we are left with densely connected clusters. The threshold for the relative degree can be set by the user. For example, using a degree threshold of 0.8 means that all the features in a cluster need to be connected with 80% of the other features of the cluster.
After each iteration of the function, all the complete components are removed from the graph. The number of components left is reported in the beginning of each iteration.
```
clusters <- find_clusters(conn)
```
The resulting list has one element per cluster, each containing a character vector of feature names and an igraph object of the cluster graph. The graph object is useful for plotting
Now that we know which features are clustered together we can create modified versions of the original intensity and feature information data frames.
```
pulled <- pull_features(clusters, data= X, features = P, name_col = "Name")
cdata <- pulled$cdata
cpeaks <- pulled$cfeatures
```
The following function will then save the modified data frames as separate sheet to the given Excel spreadsheet file. (no need to run this on Binder)
```
#multisheet_xlsx(dfs = pulled, filename = "HILIC_NEG_output2.xlsx")
```
#### Visualizations
Let's pick a cluster and take a closer look at it:
```
cluster <- clusters[[2]]
cluster$features
```
The cluster seems to have X features. We can also visualize the relationship between the fetures:
```
plot_graph(features = P, cluster, name_col = "Name",
mz_col = "mz", rt_col = "rt")
plot_features(P, cluster, name_col = "Name", mz_col = "mz", rt_col = "rt", rt_window = 1/60)
plot_heatmaps(data = X, features = P, cluster = cluster, name_col = "Name", mz_col = "mz", rt_col = "rt")
```
| github_jupyter |
# TensorFlow
In this notebook, we'll learn the basics of [TensorFlow + Keras](https://tensorflow.org), which is a machine learning library used to build dynamic neural networks. We'll learn about the basics, like creating and using Tensors.
# Set seeds
```
%tensorflow_version 2.x
import numpy as np
import tensorflow as tf
SEED = 1234
# Set seed for reproducibility
np.random.seed(seed=SEED)
tf.random.set_seed(SEED)
```
# Basics
```
# Constants
x = tf.constant(1)
print (x)
# Creating a random tensor
x = tf.random.uniform((2,3))
print(f"Type: {x.dtype}")
print(f"Size: {x.shape}")
print(f"Values: \n{x}")
# Zero and Ones tensor
x = tf.zeros((2, 3))
print (x)
x = tf.ones((2, 3))
print (x)
# List → Tensor
x = tf.convert_to_tensor([[1, 2, 3],[4, 5, 6]], dtype='int32')
print(f"Size: {x.shape}")
print(f"Values: \n{x}")
# NumPy array → Tensor
x = tf.convert_to_tensor(np.random.rand(2, 3), dtype='float32')
print(f"Size: {x.shape}")
print(f"Values: \n{x}")
# Changing tensor type
x = tf.random.uniform((2,3))
print(f"Type: {x.dtype}")
x = tf.random.uniform((2,3), dtype='float64')
print(f"Type: {x.dtype}")
```
# Operations
```
# Addition
x = tf.random.uniform((2,3))
y = tf.random.uniform((2,3))
z = x + y
print(f"Size: {z.shape}")
print(f"Values: \n{z}")
# Dot product
x = tf.random.uniform((2,3))
y = tf.random.uniform((3,2))
z = tf.matmul(x, y)
print(f"Size: {z.shape}")
print(f"Values: \n{z}")
# Transpose
x = tf.random.uniform((2,3))
print(f"Size: {x.shape}")
print(f"Values: \n{x}")
y = tf.transpose(x)
print(f"Size: {y.shape}")
print(f"Values: \n{y}")
# Reshape
x = tf.random.uniform((2,3))
z = tf.reshape(x, (1, 6))
print(f"Size: {z.shape}")
print(f"Values: \n{z}")
# Dimensional operations
x = tf.random.uniform((2,3))
print(f"Values: \n{x}")
y = tf.math.reduce_sum(x, axis=0) # add each row's value for every column
print(f"Values: \n{y}")
z = tf.math.reduce_sum(x, axis=1) # add each columns's value for every row
print(f"Values: \n{z}")
```
# Indexing, Splicing and Joining
```
x = tf.random.uniform((2,3))
print (f"x: \n{x}")
print (f"x[:1]: \n{x[:1]}")
print (f"x[:1, 1:3]: \n{x[:1, 1:3]}")
# Select with dimensional indicies
x = tf.random.uniform((2,3))
print(f"Values: \n{x}")
col_indices = tf.convert_to_tensor([0, 2])
chosen = tf.gather(x, indices=col_indices, axis=1) # values from column 0 & 2
print(f"Values: \n{chosen}")
# Concatenation
x = tf.random.uniform((2,3))
print(f"Values: \n{x}")
y = tf.concat([x, x], axis=0) # stack by rows (axis=1 to stack by columns)
print(f"Values: \n{y}")
```
# Gradients
```
# Tensors with gradient bookkeeping
x = tf.constant(3.0)
with tf.GradientTape() as g:
g.watch(x)
y = 3 * x + 2
g.gradient(y, x).numpy()
```
* $ y = 3x + 2 $
* $ \frac{\partial(y)}{\partial(x)} = 3 $
# GPUs
```
# Is CUDA available?
print (tf.config.list_physical_devices('GPU'))
```
If False (CUDA is not available), let's change that by following these steps: Go to *Runtime* > *Change runtime type* > Change *Hardware accelertor* to *GPU* > Click *Save*
```
%tensorflow_version 2.x
import tensorflow as tf
# Is CUDA available now?
print (tf.config.list_physical_devices('GPU'))
```
| github_jupyter |
```
%load_ext watermark
import numpy as np
import pandas as pd
```
## Load the JupyterHub logs
```
columns = ['user', 'machine', 'session_start', 'session_end', 'session_length', 'log_file']
df_all = pd.read_csv("../data/jhub_logs.csv.gz", parse_dates=["session_start", "session_end"])
df_all["session_length"] = (df_all.session_end - df_all.session_start).dt.total_seconds().div(60)
df_all = df_all[columns]
df_all.head()
df_all["machine"].unique()
dfs = {'all': df_all,
'Cheyenne': df_all[df_all["machine"] == 'cheyenne'],
'Casper': df_all[df_all["machine"] == 'casper']}
```
## Basic Statistics
Some basic statistics from the logs.
```
print(f'Total number of sessions:')
for name in dfs:
print(f'{name.rjust(10)}: {len(dfs[name])}')
def print_span(name):
print(f'Total span of {name} logs: {dfs[name].session_end.max() - dfs[name].session_start.min()}')
print(f' From: {dfs[name].session_start.min()}')
print(f' To: {dfs[name].session_end.max()}')
for name in dfs:
print_span(name)
print()
def print_lengths(name):
print(f'Session Lengths on {name} [minutes]:')
print(f' Minimum: {dfs[name].session_length.min()}')
print(f' Maximum: {dfs[name].session_length.max()}')
print(f' Mean: {dfs[name].session_length.mean()}')
print(f' Median: {dfs[name].session_length.median()}')
for name in dfs:
print_lengths(name)
print()
print(f'Total Number of Unique Users:')
for name in dfs:
print(f'{name.rjust(10)}: {dfs[name].user.nunique()}')
```
## Tutorials
Three tutorials have been given during the duration of the logs. We note the dates for these tutorials so that we can reference their times in the plots below.
```
tutorials = {'jun': ["2019-06-03", "2019-06-05"],
'sep': ["2019-09-18", "2019-09-20"],
'oct': ["2019-10-16", "2019-10-18"]}
```
## Sessions by Day, Week & Month
First, we take a look at the number of sessions per day, week, and month over the span of the logs.
```
days = pd.date_range(dfs['all'].session_start.min(),
dfs['all'].session_end.max() + pd.DateOffset(days=1),
freq='D').normalize()
wks = pd.date_range(dfs['all'].session_start.min() - pd.DateOffset(months=1),
dfs['all'].session_end.max() + pd.DateOffset(months=1),
freq='W').normalize()
mons = pd.date_range(dfs['all'].session_start.min() - pd.DateOffset(months=1),
dfs['all'].session_end.max() + pd.DateOffset(months=1),
freq='MS').normalize()
sess_per_day = pd.DataFrame()
sess_per_wk = pd.DataFrame()
sess_per_mon = pd.DataFrame()
for name in dfs:
sess_per_day[name] = 0.5 * (dfs[name].groupby(pd.cut(dfs[name].session_start, days)).size() +
dfs[name].groupby(pd.cut(dfs[name].session_end, days)).size()).rename(f'Sessions per Day ({name})')
sess_per_wk[name] = 0.5 * (dfs[name].groupby(pd.cut(dfs[name].session_start, wks)).size() +
dfs[name].groupby(pd.cut(dfs[name].session_end, wks)).size()).rename(f'Sessions per Week ({name})')
sess_per_mon[name] = 0.5 * (dfs[name].groupby(pd.cut(dfs[name].session_start, mons)).size() +
dfs[name].groupby(pd.cut(dfs[name].session_end, mons)).size()).rename(f'Sessions per Month ({name})')
ax = sess_per_day.plot.area(figsize=(14,5), stacked=False)
xticks = np.linspace(0, len(days)-1, 10, dtype=int)
ax.set_xticks(xticks)
ax.set_xticklabels([days[i].strftime('%Y-%m-%d') for i in xticks])
ax.set_title('Number of JupyterHub Sessions per Day', fontsize=16)
for tutorial in tutorials:
t1, t2 = tutorials[tutorial]
ax.axvspan(days.get_loc(t1)-1, days.get_loc(t2)+1, facecolor='r', alpha=.3, edgecolor='none')
ax = sess_per_wk.plot.area(figsize=(14,5))
xticks = np.linspace(0, len(wks)-1, 10, dtype=int)
ax.set_xticks(xticks)
ax.set_xticklabels([wks[i].strftime('%Y W%U') for i in xticks])
ax.set_title('Number of JupyterHub Sessions per Week', fontsize=16);
ax = sess_per_mon.plot.bar(figsize=(14,5))
xticks = np.linspace(0, len(mons)-1, 10, dtype=int)
ax.set_xticks(xticks)
ax.set_xticklabels([mons[i].strftime('%Y-%m') for i in xticks])
ax.set_title('Number of JupyterHub Sessions per Month', fontsize=16);
```
**NOTE:** You can see a definite up-tick in the number of sessions following the September and October tutorials. During this period of time, from August through October, the average number of sessions per day and month more than doubles.
## Unique Users by Day, Week & Month
Next, we look at the number of unique users per day, week and month.
```
users_per_day = pd.DataFrame()
users_per_wk = pd.DataFrame()
users_per_mon = pd.DataFrame()
for name in dfs:
users_per_day[name] = 0.5 * (dfs[name].user.groupby(pd.cut(dfs[name].session_start, days)).nunique() +
dfs[name].user.groupby(pd.cut(dfs[name].session_end, days)).nunique()).rename(f'Unique Users per Day ({name})')
users_per_wk[name] = 0.5 * (dfs[name].user.groupby(pd.cut(dfs[name].session_start, wks)).nunique() +
dfs[name].user.groupby(pd.cut(dfs[name].session_end, wks)).nunique()).rename(f'Unique Users per Week ({name})')
users_per_mon[name] = 0.5 * (dfs[name].user.groupby(pd.cut(dfs[name].session_start, mons)).nunique() +
dfs[name].user.groupby(pd.cut(dfs[name].session_end, mons)).nunique()).rename(f'Unique Users per Month ({name})')
ax = users_per_day.plot.area(figsize=(14,5), stacked=False)
xticks = np.linspace(0, len(days)-1, 10, dtype=int)
ax.set_xticks(xticks)
ax.set_xticklabels([days[i].strftime('%Y-%m-%d') for i in xticks])
ax.set_title('Number of Unqiue JupyterHub Users per Day', fontsize=16)
for tutorial in tutorials:
t1, t2 = tutorials[tutorial]
ax.axvspan(days.get_loc(t1)-1, days.get_loc(t2)+1, facecolor='r', alpha=.3, edgecolor='none')
ax = users_per_wk.plot.area(figsize=(14,5))
xticks = np.linspace(0, len(wks)-1, 10, dtype=int)
ax.set_xticks(xticks)
ax.set_xticklabels([wks[i].strftime('%Y W%U') for i in xticks])
ax.set_title('Number of Unique JupyterHub Users per Week', fontsize=16);
ax = users_per_mon.plot.bar(figsize=(14,5))
xticks = np.linspace(0, len(mons)-1, 10, dtype=int)
ax.set_xticks(xticks)
ax.set_xticklabels([mons[i].strftime('%Y-%m') for i in xticks])
ax.set_title('Number of Unique JupyterHub Users per Month', fontsize=16);
```
**NOTE:** Again, you can note a growth in users following each tutorial/hackathon (September and October).
```
%watermark -iv -d -u
```
| github_jupyter |
### Script
- **Input:** Real-time in-DB recommendation using Neo4j Graph database.
- **Output:** Sample recommendations.
### Import libraries
```
!pip3 install py2neo
from pprint import pprint
from py2neo import Graph, NodeMatcher
g = Graph("bolt://localhost:7687/neo4j", password = "test")
query="""
MATCH (a:Amenity) RETURN a.name LIMIT 3
"""
g.run(query).to_data_frame()
query="""
MATCH (a:Listing) WHERE a.price='$150.00' RETURN a.name,a.price,a.url LIMIT 3
"""
g.run(query).to_data_frame()
```
### Content-based filtering
```
query = """
MATCH (u:User {name:$cid})-[:RATED]->(s:Listing)-[:HAS_AMENITY]->(c:Amenity)<-[:HAS_AMENITY]-(z:Listing)
WHERE NOT EXISTS ((u)-[:RATED]->(z))
WITH s, z, COUNT(c) AS intersection
MATCH (s)-[:HAS_AMENITY]->(sc:Amenity)
WITH s, z, intersection, COLLECT(sc.name) AS s1
MATCH (z)-[:HAS_AMENITY]->(zc:Amenity)
WITH s, z, s1, intersection, COLLECT(zc.name) AS s2
WITH s, z, intersection, s1+[x IN s2 WHERE NOT x IN s1] AS union, s1, s2
RETURN s.name as UserListing, z.name as Recommendate, s1 as UserListingAmenities, s2 as RecommendateListingAmenities, ((1.0*intersection)/SIZE(union)) AS jaccard ORDER BY jaccard DESC LIMIT $k;
"""
recos=[]
recoAmenity=[]
for row in g.run(query, cid = "8726758", k = 5).data():
recos.append('https://www.airbnb.com/rooms/'+row['Recommendate'])
recoAmenity.append(row['UserListingAmenities'])
recos
```
### Collaborative Filtering
- https://www.datatheque.com/posts/collaborative-filtering/
- https://www.kernix.com/article/an-efficient-recommender-system-based-on-graph-database/
```
from pprint import pprint
from py2neo import Graph, NodeMatcher
g = Graph("bolt://localhost:7687/neo4j", password = "test")
user_id="27130534"
query = """
// Get count of all distinct listings that user 4789 has rated and find other users who have rated them
MATCH (u1:User {name:$uid})-[x:RATED]->(m:Listing)<-[y:RATED]-(u2:User)
WHERE u1 <> u2
WITH u1, u2, COUNT(DISTINCT m) as intersection_count
// Get count of all the distinct products that are unique to each user
MATCH (u:User)-[:RATED]->(m:Listing)
WHERE u in [u1, u2]
WITH u1, u2, intersection_count, COUNT(DISTINCT m) as union_count
// Compute Jaccard index
WITH u1, u2, intersection_count, union_count, (intersection_count*1.0/union_count) as jaccard_index
// Get top k neighbours based on Jaccard index
ORDER BY jaccard_index DESC, u2.id
WITH u1, COLLECT([u2.name, jaccard_index, intersection_count, union_count])[0..$k] as neighbours
RETURN u1.name as user, neighbours
"""
neighbours = {}
for row in g.run(query, uid=user_id, k=5):
neighbours[row[0]] = row[1]
neighbours_ids = [x[0] for x in neighbours[row[0]]]
query = """
// Get top n recommendations for user from the selected neighbours
MATCH (u1:User),
(neighbour:User)-[:RATED]->(p:Listing) // get all listings rated by neighbour
WHERE u1.name = $uid
AND neighbour.id in $neighbours
AND not (u1)-[:RATED]->(p) // which u1 has not already bought
WITH u1, p, COUNT(DISTINCT neighbour) as cnt // count times rated by neighbours
ORDER BY u1.name, cnt DESC // and sort by count desc
RETURN u1.name as user, COLLECT([p.name,cnt])[0..$k] as recos
"""
recos = {}
for row in g.run(query, uid=user_id, neighbours=neighbours_ids, k=5):
recos[row[0]] = row[1]
recommended_ids = [x[0] for x in recos[row[0]]]
query="""
MATCH (a:Listing) WHERE a.name in $name RETURN a.name,a.beds,a.bedrooms,a.bathrooms,a.picture_url,a.accomodates,a.review_scores_rating,a.price,a.url;
"""
g.run(query, name=recommended_ids).to_data_frame()
```
| github_jupyter |
```
%serialconnect
# below is the esp8266 version
# RST | GPIO1 TX
# A0 | GPIO3 RX
# D0 GPIO16 | GPIO5 D1 SCL
# SCK D5 GPIO14 | GPIO4 D2 SDA
# MISO D6 GPIO12 | GPIO0 D3
# MOSI D7 GPIO13 | GPIO2 D4 LED
# SS D8 GPIO15 | GND
# 3V3 | 5V
# You need the esp8266 code built with mqtt_as libraries
# See https://github.com/goatchurchprime/jupyter_micropython_developer_notebooks/blob/master/mqtt_async_projects/esp8266_commissioning_mqtt_as.ipynb
#HW-166
# 5V VM | PWMA D8
# 3V VCC | AIN2 D0
# GND | AIN1 D3
# MotL A01 | STBY 3V
# MotL A02 | BIN1 D5
# MotR B02 | BIN2 D6
# MotR B01 | PWMB D7
# G GND | GND G
%sendtofile --source stdmqttas.py
%sendtofile --quiet --source utils.py
%sendtofile config.txt
wifiname DoESLiverpool
wifipassword decafbad00
mqttbroker sensorcity.io
pinled 2
boardname miniwheels1
mAen 15
mAfore 0
mAback 16
mBen 13
mBfore 14
mBback 12
%serialconnect
%sendtofile main.py
import os, time, sys, machine, itertools, time
from machine import Pin, PWM
import uasyncio as asyncio
from stdmqttas import fconfig, config, mqttconnecttask, callbackcmdtask
from mqtt_as import MQTTClient
from stdmqttas import pinled, flashpinled, shortmac
#flashpinled(5, 300, 300)
import network
network.WLAN().active(0) # disable the connection at startup
from machine import Pin, PWM
mApwm = PWM(Pin(int(fconfig["mAen"])))
mAfore = Pin(int(fconfig["mAfore"]), Pin.OUT)
mAback = Pin(int(fconfig["mAback"]), Pin.OUT)
mApwm.freq(10)
mApwm.duty(0)
mBpwm = PWM(Pin(int(fconfig["mBen"])))
mBfore = Pin(int(fconfig["mBfore"]), Pin.OUT)
mBback = Pin(int(fconfig["mBback"]), Pin.OUT)
mBpwm.freq(10)
mBpwm.duty(0)
timeZeroBy = 1
async def zeroingtask():
global timeZeroBy
while True:
if timeZeroBy != 0:
if time.ticks_ms() > timeZeroBy:
timeZeroBy = 0
mAfore.value(0)
mAback.value(0)
mBfore.value(0)
mBback.value(0)
pinled.value(1)
await asyncio.sleep_ms(50)
def updatewheels(msg):
global timeZeroBy
try:
s = msg.split()
timeZeroBy = time.ticks_ms() + int(s[6])
mApwm.duty(int(s[0]))
mAfore.value(int(s[1]))
mAback.value(int(s[2]))
mBpwm.duty(int(s[3]))
mBfore.value(int(s[4]))
mBback.value(int(s[5]))
pinled.value(0)
except Exception as e:
print("Bad wheels", msg, e)
topicstem = fconfig["boardname"].encode()
topicboardcmd = topicstem + b"/cmd"
topicboardwheels = topicstem + b"/wheels"
topicstatus = topicstem+b"/status"
print("topicstem", topicstem)
import network
network.WLAN().active(0) # disable the connection at startup
for l in ["500 1 0 500 1 0 1", "500 0 1 500 0 1 1",
"500 1 0 500 0 1 1", "500 0 1 500 1 0 1", "500 0 0 500 0 0 1"]:
updatewheels(l)
time.sleep_ms(500)
def callbackcmd(topic, msg, retained):
print(topic, msg)
if topic == topicboardcmd:
aloop.create_task(callbackcmdtask(client, topicreply, msg))
else:
updatewheels(msg)
async def onconnecttask(client):
ipnumber = client._sta_if.ifconfig()[0]
await client.publish(topicstatus, ipnumber, retain=True)
await client.subscribe(topiccmd, 1)
async def onconnecttask(client):
print("subscribing")
ipnumber = client._sta_if.ifconfig()[0]
print("ipnumber", ipnumber)
await client.publish(topicstatus, ipnumber, retain=True)
print("subscribing to", topicboardcmd, topicboardwheels)
await client.subscribe(topicboardcmd, 1)
await client.subscribe(topicboardwheels, 1)
config['subs_cb'] = callbackcmd
config['connect_coro'] = onconnecttask
config['will'] = (topicstatus, "offline", True)
client = MQTTClient(config)
client.DEBUG = True
aloop = asyncio.get_event_loop()
aloop.create_task(zeroingtask())
aloop.create_task(mqttconnecttask(client))
aloop.run_forever()
# leftover attempts at light controls
from machine import Pin, I2C
i2c = I2C(scl=Pin(15), sda=Pin(4), freq=450000)
print(list(map(hex, i2c.scan())))
# TCS3472
i2c.writeto_mem(0x29, 0x80, b'\x03')
print(hex(i2c.readfrom_mem(0x29, 0x92, 1)[0]))
x = i2c.readfrom_mem(0x29, 0x8F, 1) # gain of 1x
print(x)
x = i2c.readfrom_mem(0x29, 0x81, 1)
print(x)
print(hex(i2c.readfrom_mem(0x29, 0x92, 1)[0]))
i2c.writeto_mem(0x29, 0x8F, b'\x00') # gain of 1x
i2c.writeto_mem(0x29, 0x81, b'\x00') # integration time 700ms
i2c.writeto_mem(0x29, 0x8F, b'\x01') # gain of 2x
i2c.writeto_mem(0x29, 0x81, b'\xC0') # integration time 154ms
i2c.writeto_mem(0x29, 0x8F, b'\x02') # gain of 4x
i2c.writeto_mem(0x29, 0x81, b'\xEB') # integration time 50ms
import struct, time
t0 = time.ticks_ms()
for i in range(20):
print(time.ticks_ms() - t0, struct.unpack("<HHHH", i2c.readfrom_mem(0x29, 0x94, 8)))
time.sleep(0.01)
```
| github_jupyter |
```
from pulp import *
import numpy as np
import networkx as nx
import osmnx as ox
import pandas as pd
from ast import literal_eval
import re
from time import time
np.random.seed(seed=42)
# columns = ['Periods','Retailers', 'Plants', 'Products', 'Vehicles', '# variables', '# constraints', 'Optimal solution (s)']
# experiment_results_df = pd.DataFrame(columns=columns)
# %store experiment_results_df
# %store -r experiment_results_df
# for col in experiment_results_df.columns[:-1]:
# experiment_results_df[col] = experiment_results_df[col].map(int)
# experiment_results_df[experiment_results_df.columns[-1]] = experiment_results_df[experiment_results_df.columns[-1]].apply(lambda x: round(x, 2))
```
# Set hyperparameters
```
NUM_CAFES = 4
NUM_PERIODS = 2
NUM_PRODUCTS = 2
NUM_VEHICLES = 2
NUM_WAR = 1
```
# Denver data
```
# place = {'city' : 'Denver',
# 'state' : 'Colorado',
# 'country': 'USA'}
# G = ox.graph_from_place(place, network_type='drive', truncate_by_edge=True)
# %store G
%store -r G
fig, ax = ox.plot_graph(G, figsize=(10, 10), node_size=0, edge_color='y', edge_linewidth=0.2)
# subway data pre-proccessed earlier
subway = pd.read_csv('https://raw.githubusercontent.com/olesyat/dataset_test/master/denver_subway.csv')
def get_city(address):
return address['city']
subway['address'] = subway['address'].map(literal_eval)
subway['city'] = subway['address'].map(get_city)
subway['geo'] = subway['geo'].map(literal_eval)
subway['lat'] = subway['geo'].apply(lambda x: x['lat'])
subway['lon'] = subway['geo'].apply(lambda x: x['lon'])
subway_stores = [ox.get_nearest_node(G, (x[1].lat, x[1].lon)) for x in subway.iterrows()]
np.random.shuffle(subway_stores)
subway_stores = subway_stores[:NUM_CAFES]
nodes = list(set(G.nodes.keys())-set(subway_stores))
manufacturers = [nodes[np.random.randint(len(nodes))] for i in range(NUM_WAR)]
colors = ['r' if node in manufacturers else 'g' if node in subway_stores else 'w' for node in G.nodes.keys()]
sizes = [100 if node in manufacturers else 30 if node in subway_stores else 0 for node in G.nodes.keys()]
ox.plot_graph(G, node_color=colors, node_size=sizes)
stores = subway_stores.copy()
for manuf in manufacturers[:NUM_WAR]:
stores.insert(0, manuf) # add warehouse as 0
distance_matrix = []
# time_matrix = []
path_matrix = []
for i in range(len(stores)):
new_path = []
new_distance = []
new_time = []
for j in range(len(stores)):
try:
length = nx.shortest_path_length(
G, stores[i], stores[j], weight='length')
# t = nx.shortest_path_length(
# G, stores[i], stores[j], weight='time')
path = nx.shortest_path(
G, stores[i], stores[j], weight='length')
new_distance.append(length)
new_path.append(path)
# new_time.append(t)
except:
new_distance.append(np.inf)
new_path.append(np.inf)
# new_time.append(np.inf)
path_matrix.append(new_path)
distance_matrix.append(new_distance)
# time_matrix.append(new_time)
cost = 0.62 # per liter NUMBEO
km_per_liter = 0.4
cost_per_km = cost*km_per_liter
routing_cost = cost_per_km * pd.DataFrame(distance_matrix)/1000
```
# Notations/Prerequisites
```
products = [f'product{i}' for i in range(NUM_PRODUCTS)] # n
locations = [f'cafe{i}' for i in range(NUM_CAFES)] # i
periods = [f'period{i}' for i in range(NUM_PERIODS)] # t
warehouses = [f'W{i}' for i in range(NUM_WAR)] # w
vehicles = [f'vehicle{i}' for i in range(NUM_VEHICLES)] # v
routing_cost = routing_cost.to_numpy()
routing_cost = np.vstack([routing_cost]*(len(periods))).reshape(len(periods), len(locations+warehouses), len(locations+warehouses))
```
# Parameters
## Inventory
```
D_i_n_t = np.random.randint(30, 100, size=( len(periods), len(locations), len(products))) # demand
max_D = np.max(D_i_n_t)
mean_D = np.mean(D_i_n_t)
D_i_n_t = makeDict([periods, locations, products], D_i_n_t, 0)
N_n = np.random.randint(1, 3, size=(len(products))) # Product volume per 1 unit
max_N = np.max(N_n)
N_n = makeDict([products], N_n, 0)
S_i_n = abs(np.random.rand(len(locations), len(products))) # unit storage cost
S_i_n = makeDict([locations, products],S_i_n,0)
L_i_n = np.random.randint(mean_D//2, max_D, size=(len(locations), len(products))) # storage capacity at locations: maybe do same as with vehicles?
L_i_n = makeDict([locations, products], L_i_n, 0)
# perish periods for product
PP_n = np.random.randint(2, 5, size=(len(products))) # how many periods can a product go without going bad
PP_n = makeDict([products], PP_n, 0)
```
## Production
# TODO read numbeo
```
numbeo_df = pd.read_csv('https://raw.githubusercontent.com/olesyat/dataset_test/master/numbeo.csv')
# %store -r numbeo_df
numbeo_df['price'] = numbeo_df['price'].apply(lambda x: float(x.split()[0].replace(',', '')))
product_list = ['Domestic Beer (0.5 liter draught) ',
'Imported Beer (0.33 liter bottle) ', 'Cappuccino (regular) ',
'Coke/Pepsi (0.33 liter bottle) ', 'Water (0.33 liter bottle) ',
'Milk (regular), (1 liter) ', 'Loaf of Fresh White Bread (500g) ',
'Rice (white), (1kg) ', 'Eggs (regular) (12) ',
'Local Cheese (1kg) ', 'Chicken Fillets (1kg) ',
'Beef Round (1kg) (or Equivalent Back Leg Red Meat) ',
'Apples (1kg) ', 'Banana (1kg) ', 'Oranges (1kg) ',
'Tomato (1kg) ', 'Potato (1kg) ', 'Onion (1kg) ',
'Lettuce (1 head) ', 'Water (1.5 liter bottle) ',
'Bottle of Wine (Mid-Range) ', 'Domestic Beer (0.5 liter bottle) ',
'Imported Beer (0.33 liter bottle) ']
mean_price_onethrird = (numbeo_df[numbeo_df['product'].isin(product_list)]['price']/3).mean()
std_price_onethrird = (numbeo_df[numbeo_df['product'].isin(product_list)]['price']/3).std()
C_w_n_t = np.random.randint(1.5*NUM_CAFES*max_D, 3*NUM_CAFES*max_D, size=(len(periods), len(warehouses), len(products))) # production capacity per period (maybe consider hours, but can do without it, just estimate considering hours)
C_w_n_t = makeDict([periods, warehouses, products], C_w_n_t, 0)
Y_w_n = np.abs(mean_price_onethrird + std_price_onethrird*np.random.randn(len(warehouses), len(products))) # unit production cost
Y_w_n = makeDict([warehouses, products],Y_w_n,0)
# F_w_n = np.random.randint(500, 600, size=(len(warehouses), len(products))) # production fixed cost
# F_w_n = makeDict([warehouses, products], F_w_n, 0)
```
## Routing
```
T_i_j_t = makeDict([periods,warehouses+locations,warehouses+locations],routing_cost,0)
R_v = [NUM_PRODUCTS*max_N*NUM_CAFES*max_D//NUM_VEHICLES for i in range(len(vehicles))] # consider homogeneous case: each vehicle has the same capacity
R_v = makeDict([vehicles], R_v, 0)
```
# Variables
## Production
```
P_w_n_t = LpVariable.dicts("Production",(periods, warehouses,products),0,None,LpInteger) # production at w in period t of n
```
## Inventory
```
Perished_i_n_t_dummy = LpVariable.dicts("Dummy_Perished",(periods, locations,products),None,None,LpInteger)# amount perished
Perished_i_n_t = LpVariable.dicts("Perished",(periods, locations,products),0,None,LpInteger)# amount perished
B_i_n_t = LpVariable.dicts("b_dummy",(periods, locations,products),None,None,LpBinary) # dummy for max constraint
I_i_n_t = LpVariable.dicts("In.storage",(periods, locations,products),0,None,LpInteger)# amount left at location
```
## Routing
```
Z_t_i_v = LpVariable.dicts("Location.visited",(periods, locations+warehouses, vehicles),0,None,LpBinary) # 1/0 if location/warehouse i visited by vehicle v in period t
X_i_j_t_v = LpVariable.dicts("Arc.traversed",(periods, locations+warehouses, locations+warehouses, vehicles),0,None,LpBinary) # 1/0 if route between traversed at time t for vehicle v
Q_i_n_t_v = LpVariable.dicts("Quantity.delivered",(periods, locations,vehicles, products),0,None,LpInteger) # quantity of n delivered to i at t
W_v_n_t = LpVariable.dicts("Load.vehicle",(periods, vehicles, products),0,None,LpInteger) # load of vehicle ?????????
U_t_i = LpVariable.dicts("Subtour.eliminated",(periods, locations),0,None,LpInteger) # dummy for elimination of subtours
```
# Objective function
```
model = LpProblem('Minimize_costs', LpMinimize)
tin = [(t, i, n) for t in periods for i in locations for n in products]
tijv = [(t, i, j, v) for t in periods for i in locations+warehouses for j in locations+warehouses for v in vehicles]
twn = [(t, w, n) for t in periods for w in warehouses for n in products]
model += lpSum([T_i_j_t[t][i][j]*X_i_j_t_v[t][i][j][v] for (t, i, j, v) in tijv]) + \
lpSum([S_i_n[i][n]*I_i_n_t[t][i][n] for (t, i, n) in tin]) + \
lpSum([ Y_w_n[w][n]*P_w_n_t[t][w][n]for t, w, n in twn]), 'Costs'
```
# Constratints
## Production
### 1: the production capacity is not exceeded
```
for t, w, n in twn:
model += P_w_n_t[t][w][n] <= C_w_n_t[t][w][n]
```
### 2: Quantity delivered doesn't exceed production amount
```
iv = [(i, v) for i in locations for v in vehicles]
for t, n in [(t, n) for t in periods for n in products]:
model += lpSum([Q_i_n_t_v[t][i][v][n] for i, v in iv]) <= lpSum([P_w_n_t[t][w][n] for w in warehouses])
```
## Inventory
### 3: Amount in storage does not exceed the storage capacity
```
for t, i, n in tin:
model += I_i_n_t[t][i][n] <= L_i_n[i][n]
```
### 4: Amount in storage (t) = (t-1) delivered - demnad + stored
```
for t_i, t in enumerate(periods):
for i in locations:
for n in products:
if t == 'period0':
model += I_i_n_t[t][i][n] == 0
else:
t_1 = periods[t_i-1]
model += I_i_n_t[t][i][n] == lpSum([Q_i_n_t_v[t_1][i][v][n] for v in vehicles]) - D_i_n_t[t_1][i][n] + I_i_n_t[t_1][i][n] - Perished_i_n_t[t][i][n]
```
### 5: Product saved for N periods is thrown away. (without penalty)
```
for t_i, t in enumerate(periods):
for i in locations:
for n in products:
going_bad = PP_n[n]
if t_i < going_bad: # no product can be perished before its perish date
model += Perished_i_n_t[t][i][n] == 0
else:
temp = (lpSum(Q_i_n_t_v[periods[t_i-going_bad]][i][v][n] for v in vehicles) - lpSum( D_i_n_t[periods[t_i-gb]][i][n] for gb in range(t_i,0, -1)))
model += Perished_i_n_t_dummy[t][i][n] == temp
model += Perished_i_n_t[t][i][n] >= Perished_i_n_t_dummy[t][i][n]
model += Perished_i_n_t[t][i][n] >= 0
M = 1000
model += Perished_i_n_t[t][i][n] <= Perished_i_n_t_dummy[t][i][n] + M*B_i_n_t[t][i][n]
model += Perished_i_n_t[t][i][n] <= 0 + M*(1-B_i_n_t[t][i][n])
```
### 6: (storage period) amout in storage + delivered exceeds the demand
```
for t, i, n in tin:
model += I_i_n_t[t][i][n] + \
lpSum([Q_i_n_t_v[t][i][v][n] for v in vehicles]) >= D_i_n_t[t][i][n]
```
## Routing
### 7: Load of vehicle is not exceeded
```
# capacity of vehicle is not exceeded
tv = [(t, v) for t in periods for v in vehicles]
for t, v in tv:
model += lpSum([N_n[n]*W_v_n_t[t][v][n] for n in products]) <= R_v[v]
```
### 8: load of vehicle does not exceed the produced amount
```
tn = [(t, n) for t in periods for n in products]
for t, n in tn:
model += lpSum([W_v_n_t[t][v][n] for v in vehicles]) <= lpSum([P_w_n_t[t][w][n] for w in warehouses])
```
### 9: quantity delivered does not exceed the vehicle load
```
# load of vehicle constraints
# quantity delivered does not exceed the vehicle load
tvn = [(t, v, n) for t in periods for v in vehicles for n in products]
for t, v, n in tvn:
model += ([Q_i_n_t_v[t][i][v][n] for i in locations]) <= W_v_n_t[t][v][n]
```
### 10: quantity delivered is not 0 only if LOCATION VISITED (route exists)
```
# quantity delivered is not 0 only if LOCATION VISITED (route exists)
# quantity delivered is less than the availible space
tinv = [(t, i, n, v) for t in periods for i in locations for n in products for v in vehicles]
for t, i, n, v in tinv:
model += Q_i_n_t_v[t][i][v][n] <= Z_t_i_v[t][i][v] * (L_i_n[i][n] + D_i_n_t[t][i][n])
```
### 11: quantity loaded only if warehouse visited
```
for t, v, n in tvn:
model += W_v_n_t[t][v][n] <= lpSum([Z_t_i_v[t][w][v] * C_w_n_t[t][w][n] for w in warehouses])
```
### 12: no route between same locations
```
tv = [(t, v) for t in periods for v in vehicles]
for i in locations+warehouses:
for j in locations+warehouses:
if i == j:
model += lpSum([X_i_j_t_v[t][i][j][v] for (t, v) in tv]) == 0
```
### 13: Flow constraints
```
tvi = [(t, v, i) for t in periods for v in vehicles for i in locations+warehouses]
for t, v, i in tvi:
model += lpSum([X_i_j_t_v[t][i][j][v] for j in locations+warehouses]) == Z_t_i_v[t][i][v]
```
### 14: if vehicle arrived at i it must leave from it
```
for t, v, i in tvi:
model += lpSum([X_i_j_t_v[t][j][i][v] for j in locations+warehouses]) == lpSum([X_i_j_t_v[t][i][l][v] for l in locations+warehouses])
```
### 15 start and end at warehouse
```
twv = [(t, w, v) for t in periods for w in warehouses for v in vehicles]
for t, w, v in twv:
model += lpSum([X_i_j_t_v[t][w][j][v] for j in locations+warehouses]) == lpSum([X_i_j_t_v[t][i][w][v] for i in locations])
```
### 16: Subtour elimination
```
n = len(locations)
m = len(warehouses)
tij = [(t, i, j) for t in periods for i in locations for j in locations]
for t, i, j in tij:
model += U_t_i[t][j] >= U_t_i[t][i] + 1 - (n+m)*(1 - lpSum([X_i_j_t_v[t][i][j][v] for v in vehicles]))
```
# Solve
```
start_time = time()
model.solve()
end_time = time() - start_time
# experiment_results_df.loc[len(experiment_results_df)] = [NUM_PERIODS, NUM_CAFES, NUM_WAR, NUM_PRODUCTS, NUM_VEHICLES, model.numVariables(), model.numConstraints(), end_time]
print("Status:", LpStatus[model.status])
print('Solver:', model.solver)
print("Total Cost = ", value(model.objective))
model.numConstraints(), model.numVariables(), model.isMIP()
for v in model.variables():
print(v.name, "=", v.varValue)
# %store experiment_results_df
# experiment_results_df
```
# Check
```
In_storage = pd.DataFrame(columns=['period', 'location', 'product', 'value'])
Load_vehicle = pd.DataFrame(columns=['period', 'vehicle', 'product', 'value'])
Production = pd.DataFrame(columns=['period', 'warehouse', 'product', 'value'])
Quantity_delivered = pd.DataFrame(columns=['period', 'location', 'vehicle', 'product', 'value'])
Arc_traversed = pd.DataFrame(columns=['period', 'location1', 'location2', 'vehicle', 'value'])
Location_visited = pd.DataFrame(columns=['period', 'location', 'vehicle', 'value'])
Perished = pd.DataFrame(columns=['period', 'location', 'product', 'value'])
Subtour_eliminated = pd.DataFrame()
check_dict = {'In.storage': In_storage,
'Load.vehicle': Load_vehicle,
'Production': Production,
'Quantity.delivered': Quantity_delivered,
'Arc.traversed': Arc_traversed,
'Location.visited': Location_visited,
'Perished': Perished,
'Subtour.eliminated': Subtour_eliminated}
frames = []
for v in model.variables():
splited_name = v.name.split('_')
if splited_name[0] in ['b', 'Dummy']:
continue
temp_df = check_dict[splited_name[0]]
if splited_name[0] in ['Arc.traversed', 'Quantity.delivered']:
t1 = splited_name[1]
t2 = splited_name[2]
t3 = splited_name[3]
t4 = splited_name[4]
temp_df.loc[len(temp_df)] = [t1, t2, t3, t4, v.varValue]
elif splited_name[0] == 'Subtour.eliminated':
pass
else:
t1 = splited_name[1]
t2 = splited_name[2]
t3 = splited_name[3]
temp_df.loc[len(temp_df)] = [t1, t2, t3, v.varValue]
```
# Validation
```
D_i_n_t
Production
Quantity_delivered[Quantity_delivered.period == 'period0']
# check_dict['Arc.traversed'][(check_dict['Arc.traversed'].period == 'period0')
# & (check_dict['Arc.traversed'].vehicle == 'vehicle0')].pivot_table(index='location1',
# columns='location2',
# values='value',
# aggfunc = sum)
```
| github_jupyter |
# MAPEM de Pierro algorithm for the Bowsher prior
One of the more popular methods for guiding a reconstruction based on a high quality image was suggested by Bowsher. This notebook explores this prior.
We highly recommend you look at the [PET/MAPEM](../PET/MAPEM.ipynb) notebook first. This example extends upon the quadratic prior used in that notebook to use an anatomical prior.
Authors: Kris Thielemans, Sam Ellis, Richard Brown, Casper da Costa-Luis
First version: 22nd of October 2019
Second version: 27th of October 2019
Third version: June 2021
CCP SyneRBI Synergistic Image Reconstruction Framework (SIRF)
Copyright 2019,20201 University College London
Copyright 2019 King's College London
This is software developed for the Collaborative Computational
Project in Synergistic Reconstruction for Biomedical Imaging. (http://www.synerbi.ac.uk/).
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
# Brief description of the Bowsher prior
The "usual" quadratic prior penalises differences between neighbouring voxels (using the square of the difference). This tends to oversmooth parts of the image where you know there should be an edge. To overcome this, it is natural to not penalise the difference between those "edge" voxels. This can be done after segmentation of the anatomical image for instance.
Bowsher suggested a segmentation-free approach to use an anatomical (or any "side" image) as follows:
- compute edge information on the anatomical image.
- for each voxel, consider only the $N_B$ neighbours which have the lowest difference in the anatomical image.
The paper is
Bowsher, J. E., Hong Yuan, L. W. Hedlund, T. G. Turkington, G. Akabani, A. Badea, W. C. Kurylo, et al. ‘Utilizing MRI Information to Estimate F18-FDG Distributions in Rat Flank Tumors’. In IEEE Symposium Conference Record Nuclear Science 2004., 4:2488-2492 Vol. 4, 2004. https://doi.org/10.1109/NSSMIC.2004.1462760.
# All the normal imports and handy functions
```
%matplotlib notebook
# Setup the working directory for the notebook
import notebook_setup
from sirf_exercises import cd_to_working_dir
cd_to_working_dir('Synergistic', 'MAPEM_Bowsher')
#%% Initial imports etc
import numpy
import matplotlib.pyplot as plt
import os
import sys
import shutil
from tqdm.auto import tqdm, trange
import time
from scipy.ndimage.filters import gaussian_filter
import sirf.STIR as pet
from numba import jit
from sirf_exercises import exercises_data_path
brainweb_sim_data_path = exercises_data_path('working_folder', 'Synergistic', 'BrainWeb')
# set-up redirection of STIR messages to files
msg_red = pet.MessageRedirector('info.txt', 'warnings.txt', 'errors.txt')
# plotting settings
plt.ion() # interactive 'on' such that plots appear during loops
#%% some handy function definitions
def imshow(image, limits=None, title=''):
"""Usage: imshow(image, [min,max], title)"""
plt.title(title)
bitmap = plt.imshow(image)
if limits is None:
limits = [image.min(), image.max()]
plt.clim(limits[0], limits[1])
plt.colorbar(shrink=.6)
plt.axis('off')
return bitmap
def make_cylindrical_FOV(image):
"""truncate to cylindrical FOV"""
filt = pet.TruncateToCylinderProcessor()
filt.apply(image)
#%% define a function for plotting images and the updates
# This is the same function as in `ML_reconstruction`
def plot_progress(all_images, title, subiterations, cmax):
if len(subiterations)==0:
num_subiters = all_images[0].shape[0]-1
subiterations = range(1, num_subiters+1)
num_rows = len(all_images);
slice_show = 60
for it in subiterations:
plt.figure()
for r in range(num_rows):
plt.subplot(num_rows,2,2*r+1)
imshow(all_images[r][it,slice_show,:,:], [0,cmax], '%s at %d' % (title[r], it))
plt.subplot(num_rows,2,2*r+2)
imshow(all_images[r][it,slice_show,:,:]-all_images[r][it-1,slice_show,:,:],[-cmax*.1,cmax*.1], 'update')
plt.show();
def subplot_(idx,vol,title,clims=None,cmap="viridis"):
plt.subplot(*idx)
plt.imshow(vol,cmap=cmap)
if not clims is None:
plt.clim(clims)
plt.colorbar()
plt.title(title)
plt.axis("off")
```
# Load the data
To generate the data needed for this notebook, run the [BrainWeb](./BrainWeb.ipynb) notebook first.
```
full_acquired_data = pet.AcquisitionData(os.path.join(brainweb_sim_data_path, 'FDG_sino_noisy.hs'))
atten = pet.ImageData(os.path.join(brainweb_sim_data_path, 'uMap_small.hv'))
# Anatomical image
anatomical = pet.ImageData(os.path.join(brainweb_sim_data_path, 'T1_small.hv')) # could be T2_small.hv
anatomical_arr = anatomical.as_array()
# create initial image
init_image=atten.get_uniform_copy(atten.as_array().max()*.1)
make_cylindrical_FOV(init_image)
plt.figure()
imshow(anatomical.as_array()[64, :, :])
plt.show()
plt.figure()
imshow(full_acquired_data.as_array()[0, 64, :, :])
plt.show()
```
# Code from first MAPEM notebook
The following chunk of code is copied and pasted more-or-less directly from the other notebook as a starting point.
First, run the code chunk to get the objective functions etc
### construction of Likelihood objective functions and OSEM
```
def get_obj_fun(acquired_data, atten):
print('\n------------- Setting up objective function')
# #%% create objective function
#%% create acquisition model
am = pet.AcquisitionModelUsingRayTracingMatrix()
am.set_num_tangential_LORs(5)
# Set up sensitivity due to attenuation
asm_attn = pet.AcquisitionSensitivityModel(atten, am)
asm_attn.set_up(acquired_data)
bin_eff = pet.AcquisitionData(acquired_data)
bin_eff.fill(1.0)
asm_attn.unnormalise(bin_eff)
asm_attn = pet.AcquisitionSensitivityModel(bin_eff)
# Set sensitivity of the model and set up
am.set_acquisition_sensitivity(asm_attn)
am.set_up(acquired_data,atten);
#%% create objective function
obj_fun = pet.make_Poisson_loglikelihood(acquired_data)
obj_fun.set_acquisition_model(am)
print('\n------------- Finished setting up objective function')
return obj_fun
def get_reconstructor(num_subsets, num_subiters, obj_fun, init_image):
print('\n------------- Setting up reconstructor')
#%% create OSEM reconstructor
OSEM_reconstructor = pet.OSMAPOSLReconstructor()
OSEM_reconstructor.set_objective_function(obj_fun)
OSEM_reconstructor.set_num_subsets(num_subsets)
OSEM_reconstructor.set_num_subiterations(num_subiters)
#%% initialise
OSEM_reconstructor.set_up(init_image)
print('\n------------- Finished setting up reconstructor')
return OSEM_reconstructor
# Use rebin to create a smaller sinogram to speed up calculations
acquired_data = full_acquired_data.clone()
acquired_data = acquired_data.rebin(3)
# Get the objective function
obj_fun = get_obj_fun(acquired_data, atten)
```
# Implement de Pierro MAP-EM for a quadratic prior with arbitrary weights
The following code is almost a copy-paste of the implementation by A. Mehranian and S. Ellis [contributed during one of our hackathons](https://github.com/SyneRBI/SIRF-Contribs/tree/master/src/Python/sirf/contrib/kcl). It is copied here for you to have an easier look.
Note that the code avoids the `for` loops in our simplistic version above and hence should be faster (however, the construction of the neighbourhood is still slow, but you should have to do this only once). Also, this is a Python reimplementation of MATLAB code (hence the use of "Fortran order" below).
```
def dePierroReg(image,weights,nhoodIndVec):
"""Get the de Pierro regularisation image (xreg)"""
imSize = image.shape
# vectorise image for indexing
imageVec = image.reshape(-1,order='F')
# retrieve voxel intensities for neighbourhoods
resultVec = imageVec[nhoodIndVec]
result = resultVec.reshape(weights.shape,order='F')
# compute xreg
imageReg = 0.5*numpy.sum(weights*(result + image.reshape(-1,1,order='F')),axis=1)/numpy.sum(weights,axis=1)
imageReg = imageReg.reshape(imSize,order='F')
return imageReg
def compute_nhoodIndVec(imageSize,weightsSize):
"""Get the neigbourhoods of each voxel"""
w = int(round(weightsSize[1]**(1.0/3))) # side length of neighbourhood
nhoodInd = neighbourExtract(imageSize,w)
return nhoodInd.reshape(-1,order='F')
def neighbourExtract(imageSize,w):
"""Adapted from kcl.Prior class"""
n = imageSize[0]
m = imageSize[1]
h = imageSize[2]
wlen = 2*numpy.floor(w/2)
widx = xidx = yidx = numpy.arange(-wlen/2,wlen/2+1)
if h==1:
zidx = [0]
nN = w*w
else:
zidx = widx
nN = w*w*w
Y,X,Z = numpy.meshgrid(numpy.arange(0,m), numpy.arange(0,n), numpy.arange(0,h))
N = numpy.zeros([n*m*h, nN],dtype='int32')
l = 0
for x in xidx:
Xnew = setBoundary(X + x,n)
for y in yidx:
Ynew = setBoundary(Y + y,m)
for z in zidx:
Znew = setBoundary(Z + z,h)
N[:,l] = ((Xnew + (Ynew)*n + (Znew)*n*m)).reshape(-1,1).flatten('F')
l += 1
return N
def setBoundary(X,n):
"""Boundary conditions for neighbourExtract.
Adapted from kcl.Prior class"""
idx = X<0
X[idx] = X[idx] + n
idx = X>n-1
X[idx] = X[idx] - n
return X.flatten('F')
@jit
def dePierroUpdate(xEM, imageReg, beta):
"""Update the image based on the de Pierro regularisation image"""
return (2*xEM)/(((1 - beta*imageReg)**2 + 4*beta*xEM)**0.5 + (1 - beta*imageReg) + 0.00001)
def MAPEM_iteration(OSEM_reconstructor,current_image,weights,nhoodIndVec,beta):
image_reg = dePierroReg(current_image.as_array(),weights,nhoodIndVec) # compute xreg
OSEM_reconstructor.update(current_image); # compute EM update
image_EM=current_image.as_array() # get xEM as a numpy array
updated = dePierroUpdate(image_EM, image_reg, beta) # compute new uxpdate
current_image.fill(updated) # store for next iteration
return current_image
```
## Create uniform and Bowsher weights
We will use the `kcl.Prior` class here to construct the Bowsher weights given an anatomical image. The `kcl.Prior` class (and the above code) assumes that the `weights` are returned an $N_v \times N_n$ array, with $N_v$ the number of voxels and $N_n$ the number of neighbours (here 27 as the implementation is in 3D).
```
import sirf.contrib.kcl.Prior as pr
def update_bowsher_weights(prior,side_image,num_bowsher_neighbours):
return prior.BowshserWeights\
(side_image.as_array(),num_bowsher_neighbours)
```
For illustration, we will keep only a few neighbours in the Bowsher prior. This makes the contrast with "uniform" weights higher of course.
```
num_bowsher_neighbours = 3
myPrior = pr.Prior(anatomical_arr.shape)
BowsherWeights = update_bowsher_weights(myPrior,anatomical,num_bowsher_neighbours)
```
Ignore the warning about `divide by zero`, it is actually handled in the `kcl.Prior` class.
```
# compute indices of the neighbourhood for each voxel
nhoodIndVec=compute_nhoodIndVec(anatomical_arr.shape,BowsherWeights.shape)
# illustrate that only a few of the weights in the neighbourhood are kept
# (taking an arbitrary voxel)
print(BowsherWeights[500,:])
```
You could try to understand the neighbourhood structure using the following, but it is quite complicated due to the Fortran order and linear indices.
```
toLinearIndices=nhoodIndVec.reshape(BowsherWeights.shape,order='F')
print(toLinearIndices[500,:])
```
We will also use uniform weights where every neighbour is counted the same (often people will use 1/distance between voxels as weighting, but this isn't implemented here).
```
uniformWeights=BowsherWeights.copy()
uniformWeights[:,:]=1
# set "self-weight" of the voxel to zero
uniformWeights[:,27//2]=0
print(uniformWeights[500,:])
```
# Run some experiments
```
num_subsets = 21
num_subiters = 42
```
## Do a normal OSEM (for comparison and initialisation)
```
# Do initial OSEM recon
OSEM_reconstructor = get_reconstructor(num_subsets, num_subiters, obj_fun, init_image)
osem_image = init_image.clone()
OSEM_reconstructor.reconstruct(osem_image)
plt.figure()
imshow(osem_image.as_array()[60,:,:])
plt.show();
```
## Run MAP-EM with the 2 different sets of weights
To save some time, we will initialise the algorithms with the OSEM image. This makes sense of course as in the initial iterations, the penalty will just slow everything down (as it smooths an already too smooth image even more!).
```
# arbitrary value for the weight of the penalty. You might have to tune it
beta=1
```
Compute with Bowsher penalty
```
current_image=osem_image.clone()
for it in trange(1, num_subiters+1):
current_image = MAPEM_iteration(OSEM_reconstructor,current_image,BowsherWeights,nhoodIndVec,beta)
Bowsher=current_image.clone()
```
Compute with uniform weights (we'll call the result UQP for "uniform quadratic penalty")
```
current_image=osem_image.clone()
for it in trange(1, num_subiters+1):
current_image = MAPEM_iteration(OSEM_reconstructor,current_image,uniformWeights,nhoodIndVec,beta)
UQP=current_image.clone()
# Plot the anatomical, OSEM, and two MAPEM images
plt.figure()
cmax=osem_image.max()*.6
clim=[0,cmax]
subplot_([1,2,1],anatomical.as_array()[60,:,:],"anatomical")
subplot_([1,2,2],osem_image.as_array()[60,:,:],"OSEM",clim)
plt.figure()
subplot_([1,2,1],UQP.as_array()[60,:,:],"Uniform Quadratic prior",clim)
subplot_([1,2,2],Bowsher.as_array()[60,:,:],"Bowsher Quadratic prior",clim)
plt.figure()
y_idx=osem_image.dimensions()[1]//2
plt.plot(osem_image.as_array()[60,y_idx,:],label="OSEM")
plt.plot(UQP.as_array()[60,y_idx,:],label="Uniform Quadratic prior")
plt.plot(Bowsher.as_array()[60,y_idx,:],label="Bowsher Quadratic prior")
plt.legend()
```
You will probably see that the MAP-EM are quite smooth, and that there is very little difference between the "uniform" and "Bowsher" weights after this number of updates. The difference will get larger with higher number of updates (try it!).
Also, with the Bowsher weights you should be able to increase `beta` more than for the uniform weights without oversmoothing the image too much.
# Misalignment between anatomical and emission images
What happens if you want to use an anatomical prior but the image isn't aligned with the image you're trying to reconstruct?
You'll have to register them of course! Have a look at the [registration notebook](../Reg/sirf_registration.ipynb) if you haven't already.
The idea here would be to run an initial reconstruction (say, OSEM), and then register the anatomical image to the resulting reconstruction...
Once we've got the anatomical image in the correct space, we can calculate the Bowsher weights.
```
import sirf.Reg as Reg
registration = Reg.NiftyAladinSym()
registration.set_reference_image
registration.set_reference_image(osem_image)
registration.set_floating_image(anatomical)
registration.set_parameter('SetPerformRigid','1')
registration.set_parameter('SetPerformAffine','0')
registration.process()
anatomical_in_emission_space = registration.get_output()
Bweights = update_bowsher_weights(myPrior,anatomical_in_emission_space,num_bowsher_neighbours)
```
| github_jupyter |
```
%matplotlib inline
```
*************************
Text rendering With LaTeX
*************************
Rendering text with LaTeX in Matplotlib.
Matplotlib has the option to use LaTeX to manage all text layout. This
option is available with the following backends:
* Agg
* PS
* PDF
The LaTeX option is activated by setting ``text.usetex : True`` in your rc
settings. Text handling with matplotlib's LaTeX support is slower than
matplotlib's very capable :doc:`mathtext </tutorials/text/mathtext>`, but is
more flexible, since different LaTeX packages (font packages, math packages,
etc.) can be used. The results can be striking, especially when you take care
to use the same fonts in your figures as in the main document.
Matplotlib's LaTeX support requires a working LaTeX_ installation, dvipng_
(which may be included with your LaTeX installation), and Ghostscript_
(GPL Ghostscript 9.0 or later is required). The executables for these
external dependencies must all be located on your :envvar:`PATH`.
There are a couple of options to mention, which can be changed using
:doc:`rc settings </tutorials/introductory/customizing>`. Here is an example
matplotlibrc file::
font.family : serif
font.serif : Times, Palatino, New Century Schoolbook, Bookman, Computer Modern Roman
font.sans-serif : Helvetica, Avant Garde, Computer Modern Sans serif
font.cursive : Zapf Chancery
font.monospace : Courier, Computer Modern Typewriter
text.usetex : true
The first valid font in each family is the one that will be loaded. If the
fonts are not specified, the Computer Modern fonts are used by default. All of
the other fonts are Adobe fonts. Times and Palatino each have their own
accompanying math fonts, while the other Adobe serif fonts make use of the
Computer Modern math fonts. See the PSNFSS_ documentation for more details.
To use LaTeX and select Helvetica as the default font, without editing
matplotlibrc use::
from matplotlib import rc
rc('font',**{'family':'sans-serif','sans-serif':['Helvetica']})
## for Palatino and other serif fonts use:
#rc('font',**{'family':'serif','serif':['Palatino']})
rc('text', usetex=True)
Here is the standard example,
:file:`/gallery/text_labels_and_annotations/tex_demo`:
.. figure:: ../../gallery/text_labels_and_annotations/images/sphx_glr_tex_demo_001.png
:target: ../../gallery/text_labels_and_annotations/tex_demo.html
:align: center
:scale: 50
TeX Demo
Note that display math mode (``$$ e=mc^2 $$``) is not supported, but adding the
command ``\displaystyle``, as in the above demo, will produce the same results.
<div class="alert alert-info"><h4>Note</h4><p>Certain characters require special escaping in TeX, such as::
# $ % & ~ _ ^ \ { } \( \) \[ \]
Therefore, these characters will behave differently depending on
the rcParam ``text.usetex`` flag.</p></div>
usetex with unicode
===================
It is also possible to use unicode strings with the LaTeX text manager, here is
an example taken from :file:`/gallery/text_labels_and_annotations/tex_demo`.
The axis labels include Unicode text:
.. figure:: ../../gallery/text_labels_and_annotations/images/sphx_glr_tex_demo_001.png
:target: ../../gallery/text_labels_and_annotations/tex_demo.html
:align: center
:scale: 50
TeX Unicode Demo
Postscript options
==================
In order to produce encapsulated postscript files that can be embedded in a new
LaTeX document, the default behavior of matplotlib is to distill the output,
which removes some postscript operators used by LaTeX that are illegal in an
eps file. This step produces results which may be unacceptable to some users,
because the text is coarsely rasterized and converted to bitmaps, which are not
scalable like standard postscript, and the text is not searchable. One
workaround is to set ``ps.distiller.res`` to a higher value (perhaps 6000)
in your rc settings, which will produce larger files but may look better and
scale reasonably. A better workaround, which requires Poppler_ or Xpdf_, can be
activated by changing the ``ps.usedistiller`` rc setting to ``xpdf``. This
alternative produces postscript without rasterizing text, so it scales
properly, can be edited in Adobe Illustrator, and searched text in pdf
documents.
Possible hangups
================
* On Windows, the :envvar:`PATH` environment variable may need to be modified
to include the directories containing the latex, dvipng and ghostscript
executables. See `environment-variables` and
`setting-windows-environment-variables` for details.
* Using MiKTeX with Computer Modern fonts, if you get odd \*Agg and PNG
results, go to MiKTeX/Options and update your format files
* On Ubuntu and Gentoo, the base texlive install does not ship with
the type1cm package. You may need to install some of the extra
packages to get all the goodies that come bundled with other latex
distributions.
* Some progress has been made so matplotlib uses the dvi files
directly for text layout. This allows latex to be used for text
layout with the pdf and svg backends, as well as the \*Agg and PS
backends. In the future, a latex installation may be the only
external dependency.
Troubleshooting
===============
* Try deleting your :file:`.matplotlib/tex.cache` directory. If you don't know
where to find :file:`.matplotlib`, see `locating-matplotlib-config-dir`.
* Make sure LaTeX, dvipng and ghostscript are each working and on your
:envvar:`PATH`.
* Make sure what you are trying to do is possible in a LaTeX document,
that your LaTeX syntax is valid and that you are using raw strings
if necessary to avoid unintended escape sequences.
* Most problems reported on the mailing list have been cleared up by
upgrading Ghostscript_. If possible, please try upgrading to the
latest release before reporting problems to the list.
* The ``text.latex.preamble`` rc setting is not officially supported. This
option provides lots of flexibility, and lots of ways to cause
problems. Please disable this option before reporting problems to
the mailing list.
* If you still need help, please see `reporting-problems`
| github_jupyter |
```
#all_slow
```
# Tutorial - Migrating from Lightning
> Incrementally adding fastai goodness to your Lightning training
We're going to use the MNIST training code from Lightning's 'Quick Start' (as at August 2020), converted to a module. See `migrating_lightning.py` for the Lightning code we are importing here.
```
from migrating_lightning import *
from fastai.vision.all import *
```
## Using fastai's training loop
We can use the Lightning module directly:
```
model = LitModel()
```
To use it in fastai, we first pull the DataLoaders from the module into a `DataLoaders` object:
```
data = DataLoaders(model.train_dataloader(), model.val_dataloader()).cuda()
```
We can now create a `Learner` and fit:
```
learn = Learner(data, model, loss_func=F.cross_entropy, opt_func=Adam, metrics=accuracy)
learn.fit_one_cycle(1, 0.001)
```
As you can see, migrating from Lightning allowed us to reduce the amount of code, and doesn't require you to change any of your existing data pipelines, optimizers, loss functions, models, etc. Once you've made this change, you can then benefit from fastai's rich set of callbacks, transforms, visualizations, and so forth.
For instance, in the Lightning example, Tensorboard support was defined a special-case "logger". In fastai, Tensorboard is just another `Callback` that you can add, with the parameter `cbs=Tensorboard`, when you create your `Learner`. The callbacks all work together, so you can add an remove any schedulers, loggers, visualizers, and so forth. You don't have to learn about special types of functionality for each - they are all just plain callbacks.
Note that fastai is very different from Lightning, in that it is much more than just a training loop (although we're only using the training loop in this example) - it is a complete framework including GPU-accelerated transformations, end-to-end inference, integrated applications for vision, text, tabular, and collaborative filtering, and so forth. You can use any part of the framework on its own, or combine them together, as described in the [fastai paper](https://arxiv.org/abs/2002.04688).
### Taking advantage of fastai Data Blocks
One problem in the Lightning example is that it doesn't actually use a validation set - it's just using the training set a second time as a validation set.
You might prefer to use fastai's Data Block API, which makes it really easy to create, visualize, and test your input data processing. Here's how you can create input data for MNIST, for instance:
```
mnist = DataBlock(blocks=(ImageBlock(cls=PILImageBW), CategoryBlock),
get_items=get_image_files,
splitter=GrandparentSplitter(),
get_y=parent_label)
```
Here, we're telling `DataBlock` that we have a B&W image input, and a category output, our input items are file names of images, the images are labeled based on the name of the parent folder, and they are split by training vs validation based on the grandparent folder name. It's important to actually look at your data, so fastai also makes it easy to visualize your inputs and outputs, for instance:
```
dls = mnist.dataloaders(untar_data(URLs.MNIST_TINY))
dls.show_batch(max_n=9, figsize=(4,4))
```
| github_jupyter |
<img src="images/dask_horizontal.svg" align="right" width="30%">
# Data Storage
<img src="images/hdd.jpg" width="20%" align="right">
Efficient storage can dramatically improve performance, particularly when operating repeatedly from disk.
Decompressing text and parsing CSV files is expensive. One of the most effective strategies with medium data is to use a binary storage format like HDF5. Often the performance gains from doing this is sufficient so that you can switch back to using Pandas again instead of using `dask.dataframe`.
In this section we'll learn how to efficiently arrange and store your datasets in on-disk binary formats. We'll use the following:
1. [Pandas `HDFStore`](http://pandas.pydata.org/pandas-docs/stable/io.html#io-hdf5) format on top of `HDF5`
2. Categoricals for storing text data numerically
**Main Take-aways**
1. Storage formats affect performance by an order of magnitude
2. Text data will keep even a fast format like HDF5 slow
3. A combination of binary formats, column storage, and partitioned data turns one second wait times into 80ms wait times.
## Create data
```
%run prep.py -d accounts
```
## Read CSV
First we read our csv data as before.
CSV and other text-based file formats are the most common storage for data from many sources, because they require minimal pre-processing, can be written line-by-line and are human-readable. Since Pandas' `read_csv` is well-optimized, CSVs are a reasonable input, but far from optimized, since reading required extensive text parsing.
```
import os
filename = os.path.join('data', 'accounts.*.csv')
filename
import dask.dataframe as dd
df_csv = dd.read_csv(filename)
df_csv.head()
```
### Write to HDF5
HDF5 and netCDF are binary array formats very commonly used in the scientific realm.
Pandas contains a specialized HDF5 format, `HDFStore`. The ``dd.DataFrame.to_hdf`` method works exactly like the ``pd.DataFrame.to_hdf`` method.
```
target = os.path.join('data', 'accounts.h5')
target
# convert to binary format, takes some time up-front
%time df_csv.to_hdf(target, '/data')
# same data as before
df_hdf = dd.read_hdf(target, '/data')
df_hdf.head()
```
### Compare CSV to HDF5 speeds
We do a simple computation that requires reading a column of our dataset and compare performance between CSV files and our newly created HDF5 file. Which do you expect to be faster?
```
%time df_csv.amount.sum().compute()
%time df_hdf.amount.sum().compute()
```
Sadly they are about the same, or perhaps even slower.
The culprit here is `names` column, which is of `object` dtype and thus hard to store efficiently. There are two problems here:
1. How do we store text data like `names` efficiently on disk?
2. Why did we have to read the `names` column when all we wanted was `amount`
### 1. Store text efficiently with categoricals
We can use Pandas categoricals to replace our object dtypes with a numerical representation. This takes a bit more time up front, but results in better performance.
More on categoricals at the [pandas docs](http://pandas.pydata.org/pandas-docs/stable/categorical.html) and [this blogpost](http://matthewrocklin.com/blog/work/2015/06/18/Categoricals).
```
# Categorize data, then store in HDFStore
%time df_hdf.categorize(columns=['names']).to_hdf(target, '/data2')
# It looks the same
df_hdf = dd.read_hdf(target, '/data2')
df_hdf.head()
# But loads more quickly
%time df_hdf.amount.sum().compute()
```
This is now definitely faster than before. This tells us that it's not only the file type that we use but also how we represent our variables that influences storage performance.
How does the performance of reading depend on the scheduler we use? You can try this with threaded, processes and distributed.
However this can still be better. We had to read all of the columns (`names` and `amount`) in order to compute the sum of one (`amount`). We'll improve further on this with `parquet`, an on-disk column-store. First though we learn about how to set an index in a dask.dataframe.
### Exercise
`fastparquet` is a library for interacting with parquet-format files, which are a very common format in the Big Data ecosystem, and used by tools such as Hadoop, Spark and Impala.
```
target = os.path.join('data', 'accounts.parquet')
df_csv.categorize(columns=['names']).to_parquet(target, storage_options={"has_nulls": True}, engine="fastparquet")
```
Investigate the file structure in the resultant new directory - what do you suppose those files are for?
`to_parquet` comes with many options, such as compression, whether to explicitly write NULLs information (not necessary in this case), and how to encode strings. You can experiment with these, to see what effect they have on the file size and the processing times, below.
```
ls -l data/accounts.parquet/
df_p = dd.read_parquet(target)
# note that column names shows the type of the values - we could
# choose to load as a categorical column or not.
df_p.dtypes
```
Rerun the sum computation above for this version of the data, and time how long it takes. You may want to try this more than once - it is common for many libraries to do various setup work when called for the first time.
```
%time df_p.amount.sum().compute()
```
When archiving data, it is common to sort and partition by a column with unique identifiers, to facilitate fast look-ups later. For this data, that column is `id`. Time how long it takes to retrieve the rows corresponding to `id==100` from the raw CSV, from HDF5 and parquet versions, and finally from a new parquet version written after applying `set_index('id')`.
```
df_p.set_index('id').to_parquet(target, storage_options={"has_nulls": True}, engine="fastparquet")
%time df_csv[df_csv.id == 100].compute()
df_p = dd.read_parquet(target)
%time df_p.loc[100].compute()
```
## Remote files
Dask can access various cloud- and cluster-oriented data storage services such as Amazon S3 or HDFS
Advantages:
* scalable, secure storage
Disadvantages:
* network speed becomes bottleneck
The way to set up dataframes (and other collections) remains very similar to before. Note that the data here is available anonymously, but in general an extra parameter `storage_options=` can be passed with further details about how to interact with the remote storage.
```python
taxi = dd.read_csv('s3://nyc-tlc/trip data/yellow_tripdata_2015-*.csv',
storage_options={'anon': True})
```
**Warning**: operations over the Internet can take a long time to run. Such operations work really well in a cloud clustered set-up, e.g., amazon EC2 machines reading from S3 or Google compute machines reading from GCS.
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
```
# LDLS Demo
This notebook demonstrates how to use LDLS to perform instance segmentation of a LiDAR point cloud. This demo uses Frame 571 from the KITTI object detection dataset.
## Setup
Import LiDAR segmentation modules:
```
import numpy as np
from pathlib import Path
import skimage
from lidar_segmentation.detections import MaskRCNNDetections
from lidar_segmentation.segmentation import LidarSegmentation
from lidar_segmentation.kitti_utils import load_kitti_lidar_data, load_kitti_object_calib
from lidar_segmentation.utils import load_image
from mask_rcnn.mask_rcnn import MaskRCNNDetector
```
# Load input data
Load the following files:
- Calibration data (relates the LiDAR and camera sensor coordinate frames)
- Image
- LiDAR point cloud
```
# Define file paths
calib_path = Path("data/") / "kitti_demo" / "calib" / "000571.txt"
image_path = Path("data/") / "kitti_demo" / "image_2" / "000571.png"
lidar_path = Path("data/") / "kitti_demo" / "velodyne" / "000571.bin"
# Load calibration data
projection = load_kitti_object_calib(calib_path)
# Load image
image = load_image(image_path)
skimage.io.imshow(image)
# Load lidar
lidar = load_kitti_lidar_data(lidar_path, load_reflectance=False)
print("Loaded LiDAR point cloud with %d points" % lidar.shape[0])
```
# Run Mask-RCNN detector on image
The first step in the LDLS pipeline is to run Mask-RCNN on the input image to generate 2D segmentation masks. The following code block runs Mask-RCNN and visualizes results on the input image.
```
detector = MaskRCNNDetector()
detections = detector.detect(image)
detections.visualize(image)
```
# Perform LiDAR segmentation
Next, perform 3D segmentation using a LidarSegmentation object. The LidarSegmentation.run() method takes as inputs a LiDAR point cloud, Mask-RCNN detections, and a maximum number of iterations parameter.
```
lidarseg = LidarSegmentation(projection)
# Be sure to set save_all=False when running segmentation
# If set to true, returns label diffusion results at each iteration in the results
# This is useful for analysis or visualizing the diffusion, but slow.
results = lidarseg.run(lidar, detections, max_iters=50, save_all=False)
%timeit lidarseg.run(lidar, detections, max_iters=50, save_all=False)
```
# Visualize results using Plotly
Plot the resulting labeled pointcloud using [Plotly](https://plot.ly/). You can visualize the results with points colored according to class labels (Person, Car, ...), or instance labels (Person 1, Person 2, Car 1, ...).
```
from lidar_segmentation.plotting import plot_segmentation_result
# Show points colored by class label
plot_segmentation_result(results, label_type='class')
# Show points colored by instance label
plot_segmentation_result(results, label_type='instance')
```
You can also visualize the label diffusion over time. This requires running the lidar segmentation with the `save_all` parameter set to `true` (note that this is significantly slower due to saving the full diffusion results in an array).
Run the following code block to visualize this. You can use the slide bar on the bottom to see results at different iterations.
```
from lidar_segmentation.plotting import plot_diffusion
results_all = lidarseg.run(lidar, detections, max_iters=50, save_all=True)
plot_diffusion(results_all)
```
| github_jupyter |
<a id='step_1'></a>
# Step 1
Read `candidates.txt` and `contributors.txt` and insert their values into the respective tables.
<a id='step_2'></a>
# Step 2: Various Queries
### Do the following queries:
* Display the contributors where the state is "PA"
* Display the contributors where the amount contributed is greater than $\$1000.00$.
* Display the contributors from "UT" where the amount contributed is greater than $\$1000.00$.
* Display the contributors who didn't list their state
- **Hint**: Match `state` to the empty string
* Display the contributors from "WA" and "PA"
- **Hint**: You will need to use `IN ("WA", "PA")` in your `SELECT` statement.
* Display the contributors who contributed between $\$100.00$ and $\$200.00$.
- **Hint**: You can use the `BETWEEN 100.00 and 200.00` clause.
<a id='step_3'></a>
# Step 3: Sorting
### Do the following sorts on the `contributors` table:
* Sort the `candidates` table by `last_name`.
* Sort by the `amount` in decending order where `amount` is restricted to be between $\$1000.00$ and $\$5000.00$.
* Sort the contributors who donted between $\$1000.00$ and $\$5000.00$ by `candidate_id` and then by `amount` in descending order.
- **Hint**: Multiple orderings can be accomplished by separating requests after `ORDER BY` with commas.
- e.g. `ORDER BY amount ASC, last_name DESC`
<a id='step_4'></a>
# Step 4: Selecting Columns
### Do the following:
* Get the first and last name of contributors. Make sure each row has distinct values.
<a id='step_5'></a>
# Step 5: Altering Tables
### Do the following:
* Add a new column to the contributors table called `full_name`. The value in that column should be in the form `last_name, first_name`.
* Change the value in the `full_name` column to the string `"Too Much"` if someone donated more than $\$1000.00$
<a id='step_6'></a>
# Step 6: Aggregation
### Do the following:
* Count how many donations there were above $\$1000.00$.
* Calculate the average donation.
* Calculate the average contribution from each state and display in a table.
- **Hint**: Use code that looks like:
```python
"SELECT state,SUM(amount) FROM contributors GROUP BY state"
```
<a id='step_7'></a>
# Step 7: DELETE
### Do the following:
* Delete rows in the `contributors` table with last name "Ahrens".
<a id='step_8'></a>
# Step 8: LIMIT
### Do the following:
* Query and display the ten most generous donors.
* Query and display the ten least generous donors who donated a positive amount of money (since the data we have has some negative numbers in it...).
# Save
Don't forget to save all of these changes to your database using `db.commit()`. Before closing shop, be sure to close the database connection with `db.close()`.
```
import sqlite3
import pandas as pd
from IPython.core.display import display
pd.set_option('display.width', 500)
pd.set_option('display.max_columns', 100)
pd.set_option('display.notebook_repr_html', True)
# Create and connect to database
db = sqlite3.connect('L18DB.sqlite')
cursor = db.cursor()
cursor.execute("DROP TABLE IF EXISTS candidates")
cursor.execute("DROP TABLE IF EXISTS contributors")
cursor.execute("PRAGMA foreign_keys=1")
# Create the candidates Table
cursor.execute('''CREATE TABLE candidates (
id INTEGER PRIMARY KEY NOT NULL,
first_name TEXT,
last_name TEXT,
middle_init TEXT,
party TEXT NOT NULL)''')
db.commit()
# Create the contributors Table
cursor.execute('''CREATE TABLE contributors (
id INTEGER PRIMARY KEY AUTOINCREMENT NOT NULL,
last_name TEXT,
first_name TEXT,
middle_name TEXT,
street_1 TEXT,
street_2 TEXT,
city TEXT,
state TEXT,
zip TEXT,
amount REAL,
date DATETIME,
candidate_id INTEGER NOT NULL,
FOREIGN KEY(candidate_id) REFERENCES candidates(id))''')
db.commit()
# The given helper function to visualize tables
def viz_tables(cols, query):
q = cursor.execute(query).fetchall()
framelist = []
for i, col_name in enumerate(cols):
framelist.append((col_name, [col[i] for col in q]))
return pd.DataFrame.from_items(framelist)
# ==========================================
# Step 1
# Populate the candidates table
with open ("candidates.txt") as candidates:
next(candidates) # jump over the header
for line in candidates.readlines():
cid, first_name, last_name, middle_name, party = line.strip().split('|')
vals_to_insert = (int(cid), first_name, last_name, middle_name, party)
cursor.execute('''INSERT INTO candidates
(id, first_name, last_name, middle_init, party)
VALUES (?, ?, ?, ?, ?)''', vals_to_insert)
# Populate the contributors table
with open ("contributors.txt") as contributors:
next(contributors)
for line in contributors.readlines():
cid, last_name, first_name, middle_name, street_1, street_2, \
city, state, zip_code, amount, date, candidate_id = line.strip().split('|')
vals_to_insert = (last_name, first_name, middle_name, street_1, street_2,
city, state, int(zip_code), amount, date, candidate_id)
cursor.execute('''INSERT INTO contributors (last_name, first_name, middle_name,
street_1, street_2, city, state, zip, amount, date, candidate_id)
VALUES (?, ?, ?, ?, ?, ?, ?, ?, ?, ?, ?)''', vals_to_insert)
# ==========================================
# Step 2
# get all columns from the contributors table
contributor_cols = [col[1] for col in cursor.execute("PRAGMA table_info(contributors)")]
# the list of queries
queries = []
# select rows where states = PA
queries.append('''SELECT * FROM contributors WHERE state = "PA" ''')
# select rows where amount > 1000
queries.append('''SELECT * FROM contributors WHERE amount > 1000 ''')
# select rows where state = UT and amount > 1000
queries.append('''SELECT * FROM contributors WHERE state = "UT" and amount > 1000 ''')
# select rows rows where state is ""
queries.append('''SELECT * FROM contributors WHERE state = "" ''')
# select rows where state is IN ("WA", "PA")
queries.append('''SELECT * FROM contributors WHERE state IN ("WA", "PA") ''')
# select rows where amount within range (100, 200)
queries.append('''SELECT * FROM contributors WHERE amount BETWEEN 100 and 200 ''')
# ==========================================
# Step 3
# sort by last_name
queries.append('''SELECT * FROM contributors ORDER BY last_name ''')
# select amount within (1000, 5000) and sort by amount in decending order
queries.append('''SELECT * FROM contributors WHERE amount BETWEEN 1000 and 5000 ORDER BY amount DESC ''')
# select amount within (1000, 5000)
# sort rows firstly by candidate_id and then by amount in decending order
queries.append('''SELECT * FROM contributors WHERE amount BETWEEN 1000 and 5000 ORDER BY candidate_id, amount DESC ''')
# visualize the queries results
for q in queries:
display(viz_tables(contributor_cols, q))
# ==========================================
# Step 4
query = '''SELECT DISTINCT first_name, last_name FROM contributors'''
display(viz_tables(['first_name', 'last_name'], query))
# ==========================================
# Step 5
# create full_name column
cursor.execute('''ALTER TABLE contributors ADD COLUMN full_name TEXT''')
# Select last_name, first_name, id
query = '''SELECT last_name, first_name, id FROM contributors'''
full_name_and_id = [(attr[0] + ", " + attr[1], attr[2]) for attr in cursor.execute(query).fetchall()]
# Update the full_name at rows specified by id
update = '''UPDATE contributors SET full_name = ? WHERE id = ?'''
for rows in full_name_and_id:
cursor.execute(update, rows)
# Set full_name to 'Too Much' where amount > 1000
update = '''UPDATE contributors SET full_name = "Too Much" WHERE amount > 1000'''
cursor.execute(update)
# get all columns from the contributors table after adding full_name
contributor_cols = [col[1] for col in cursor.execute("PRAGMA table_info(contributors)")]
# ==========================================
# Step 6
# Count # of amounts > 1000
count = '''SELECT COUNT(id) FROM contributors WHERE amount > 1000'''
display(viz_tables(['count'], count))
# Calculate the average donation
avg_don = '''SELECT AVG(amount) FROM contributors'''
display(viz_tables(['avg_don'], avg_don))
# Calculate the average donation from each state
state_avg = '''SELECT state,AVG(amount) FROM contributors GROUP BY state'''
display(viz_tables(['state', 'avg_don'], state_avg))
# ==========================================
# Step 7
# Delete rows in the contributors table where last_name = "Ahrens".
cursor.execute('''DELETE FROM contributors WHERE last_name = "Ahrens" ''')
display(viz_tables(contributor_cols, '''SELECT * FROM contributors'''))
# ==========================================
# Step 8
# Select the first ten rows order by amount descendingly
query = '''SELECT * FROM contributors ORDER BY amount DESC LIMIT 10'''
display(viz_tables(contributor_cols, query))
# Select the first ten rows where amount > 0 (least generous donors) order by amount ascendingly
query = '''SELECT * FROM contributors WHERE amount > 0 ORDER BY amount LIMIT 10'''
display(viz_tables(contributor_cols, query))
# ==========================================
# Save
db.commit()
db.close()
```
| github_jupyter |
```
# ========================================
# library
# ========================================
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import StratifiedKFold, KFold,GroupKFold
from sklearn.metrics import mean_squared_error
%matplotlib inline
import torch
import torch.nn as nn
from torch.utils.data import Dataset, DataLoader, Subset
import transformers
from transformers import LongformerTokenizer, LongformerModel,AutoTokenizer,RobertaModel,FunnelModel
from transformers import AdamW, get_linear_schedule_with_warmup
from torch.cuda.amp import autocast, GradScaler
import logging
from ast import literal_eval
import sys
from contextlib import contextmanager
import time
import random
from tqdm import tqdm
import os
# ==================
# Constant
# ==================
TRAIN_PATH = "../data/train.csv"
DATA_DIR = "../data/funnel-large/"
if not os.path.exists(DATA_DIR):
os.makedirs(DATA_DIR)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# ===============
# Configs
# ===============
max_len = 512
MODEL_PATH = 'funnel-transformer/large'
tokenizer = AutoTokenizer.from_pretrained(MODEL_PATH)
# ===============
# Functions
# ===============
def set_seed(seed: int = 42):
random.seed(seed)
np.random.seed(seed)
os.environ["PYTHONHASHSEED"] = str(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
target_map_rev = {0:'Lead', 1:'Position', 2:'Evidence', 3:'Claim', 4:'Concluding Statement',
5:'Counterclaim', 6:'Rebuttal', 7:'blank'}
def get_preds_collatte(dataset, verbose,text_ids, preds, preds_len):
all_predictions = []
for id_num in tqdm(range(len(preds))):
# GET ID
#if (id_num%100==0)&(verbose):
# print(id_num,', ',end='')
n = text_ids[id_num]
max_len = int(preds_len[id_num])
# GET TOKEN POSITIONS IN CHARS
name = f'../data/{dataset}/{n}.txt'
txt = open(name, 'r').read()
tokens = tokenizer.encode_plus(txt, max_length=max_len, padding='max_length',
truncation=True, return_offsets_mapping=True)
off = tokens['offset_mapping']
# GET WORD POSITIONS IN CHARS
w = []
blank = True
for i in range(len(txt)):
if (txt[i]!=' ')&(txt[i]!='\n')&(txt[i]!='\xa0')&(txt[i]!='\x85')&(blank==True):
w.append(i)
blank=False
elif (txt[i]==' ')|(txt[i]=='\n')|(txt[i]=='\xa0')|(txt[i]=='\x85'):
blank=True
w.append(1e6)
# MAPPING FROM TOKENS TO WORDS
word_map = -1 * np.ones(max_len,dtype='int32')
w_i = 0
for i in range(len(off)):
if off[i][1]==0: continue
while off[i][0]>=w[w_i+1]: w_i += 1
word_map[i] = int(w_i)
# CONVERT TOKEN PREDICTIONS INTO WORD LABELS
### KEY: ###
# 0: LEAD_B, 1: LEAD_I
# 2: POSITION_B, 3: POSITION_I
# 4: EVIDENCE_B, 5: EVIDENCE_I
# 6: CLAIM_B, 7: CLAIM_I
# 8: CONCLUSION_B, 9: CONCLUSION_I
# 10: COUNTERCLAIM_B, 11: COUNTERCLAIM_I
# 12: REBUTTAL_B, 13: REBUTTAL_I
# 14: NOTHING i.e. O
### NOTE THESE VALUES ARE DIVIDED BY 2 IN NEXT CODE LINE
pred = preds[id_num,]/2.0
i = 0
while i<max_len:
prediction = []
start = pred[i]
if start in [0,1,2,3,4,5,6,7]:
prediction.append(word_map[i])
i += 1
if i>=max_len: break
while pred[i]==start+0.5:
if not word_map[i] in prediction:
prediction.append(word_map[i])
i += 1
if i>=max_len: break
else:
i += 1
prediction = [x for x in prediction if x!=-1]
if len(prediction)>4:
all_predictions.append( (n, target_map_rev[int(start)],
' '.join([str(x) for x in prediction]) ) )
# MAKE DATAFRAME
df = pd.DataFrame(all_predictions)
df.columns = ['id','class','predictionstring']
return df
def calc_overlap(row):
"""
Calculates the overlap between prediction and
ground truth and overlap percentages used for determining
true positives.
"""
set_pred = set(row.predictionstring_pred.split(' '))
set_gt = set(row.predictionstring_gt.split(' '))
# Length of each and intersection
len_gt = len(set_gt)
len_pred = len(set_pred)
inter = len(set_gt.intersection(set_pred))
overlap_1 = inter / len_gt
overlap_2 = inter/ len_pred
return [overlap_1, overlap_2]
def score_feedback_comp(pred_df, gt_df):
"""
A function that scores for the kaggle
Student Writing Competition
Uses the steps in the evaluation page here:
https://www.kaggle.com/c/feedback-prize-2021/overview/evaluation
"""
gt_df = gt_df[['id','discourse_type','predictionstring']] \
.reset_index(drop=True).copy()
pred_df = pred_df[['id','class','predictionstring']] \
.reset_index(drop=True).copy()
pred_df['pred_id'] = pred_df.index
gt_df['gt_id'] = gt_df.index
# Step 1. all ground truths and predictions for a given class are compared.
joined = pred_df.merge(gt_df,
left_on=['id','class'],
right_on=['id','discourse_type'],
how='outer',
suffixes=('_pred','_gt')
)
joined['predictionstring_gt'] = joined['predictionstring_gt'].fillna(' ')
joined['predictionstring_pred'] = joined['predictionstring_pred'].fillna(' ')
joined['overlaps'] = joined.apply(calc_overlap, axis=1)
# 2. If the overlap between the ground truth and prediction is >= 0.5,
# and the overlap between the prediction and the ground truth >= 0.5,
# the prediction is a match and considered a true positive.
# If multiple matches exist, the match with the highest pair of overlaps is taken.
joined['overlap1'] = joined['overlaps'].apply(lambda x: eval(str(x))[0])
joined['overlap2'] = joined['overlaps'].apply(lambda x: eval(str(x))[1])
joined['potential_TP'] = (joined['overlap1'] >= 0.5) & (joined['overlap2'] >= 0.5)
joined['max_overlap'] = joined[['overlap1','overlap2']].max(axis=1)
tp_pred_ids = joined.query('potential_TP') \
.sort_values('max_overlap', ascending=False) \
.groupby(['id','predictionstring_gt']).first()['pred_id'].values
# 3. Any unmatched ground truths are false negatives
# and any unmatched predictions are false positives.
fp_pred_ids = [p for p in joined['pred_id'].unique() if p not in tp_pred_ids]
matched_gt_ids = joined.query('potential_TP')['gt_id'].unique()
unmatched_gt_ids = [c for c in joined['gt_id'].unique() if c not in matched_gt_ids]
# Get numbers of each type
TP = len(tp_pred_ids)
FP = len(fp_pred_ids)
FN = len(unmatched_gt_ids)
#calc microf1
my_f1_score = TP / (TP + 0.5*(FP+FN))
return my_f1_score
def collatte(d,train=True):
mask_len = int(d["mask"].sum(axis=1).max())
if train:
return {"token" : d['token'][:,:mask_len],
"mask" : d['mask'][:,:mask_len],
"y" : d['y'][:,:mask_len],
"max_len" : mask_len}
else:
return {"token" : d['token'][:,:mask_len],
"mask" : d['mask'][:,:mask_len],
"max_len" : mask_len}
# ================================
# Main
# ================================
train = pd.read_csv(TRAIN_PATH)
IDS = train.id.unique()
id_array = np.array(IDS)
MAX_LEN = max_len
# # THE TOKENS AND ATTENTION ARRAYS
train_tokens = np.zeros((len(IDS),MAX_LEN), dtype='int32')
train_attention = np.zeros((len(IDS),MAX_LEN), dtype='int32')
# THE 14 CLASSES FOR NER
lead_b = np.zeros((len(IDS),MAX_LEN))
lead_i = np.zeros((len(IDS),MAX_LEN))
position_b = np.zeros((len(IDS),MAX_LEN))
position_i = np.zeros((len(IDS),MAX_LEN))
evidence_b = np.zeros((len(IDS),MAX_LEN))
evidence_i = np.zeros((len(IDS),MAX_LEN))
claim_b = np.zeros((len(IDS),MAX_LEN))
claim_i = np.zeros((len(IDS),MAX_LEN))
conclusion_b = np.zeros((len(IDS),MAX_LEN))
conclusion_i = np.zeros((len(IDS),MAX_LEN))
counterclaim_b = np.zeros((len(IDS),MAX_LEN))
counterclaim_i = np.zeros((len(IDS),MAX_LEN))
rebuttal_b = np.zeros((len(IDS),MAX_LEN))
rebuttal_i = np.zeros((len(IDS),MAX_LEN))
# HELPER VARIABLES
train_lens = []
targets_b = [lead_b, position_b, evidence_b, claim_b, conclusion_b, counterclaim_b, rebuttal_b]
targets_i = [lead_i, position_i, evidence_i, claim_i, conclusion_i, counterclaim_i, rebuttal_i]
target_map = {'Lead':0, 'Position':1, 'Evidence':2, 'Claim':3, 'Concluding Statement':4,
'Counterclaim':5, 'Rebuttal':6}
# # WE ASSUME DATAFRAME IS ASCENDING WHICH IT IS
# assert( np.sum(train.groupby('id')['discourse_start'].diff()<=0)==0 )
# # FOR LOOP THROUGH EACH TRAIN TEXT
for id_num in tqdm(range(len(IDS))):
# READ TRAIN TEXT, TOKENIZE, AND SAVE IN TOKEN ARRAYS
n = IDS[id_num]
name = f'../data/train/{n}.txt'
txt = open(name, 'r').read()
train_lens.append( len(txt.split()))
tokens = tokenizer.encode_plus(txt, max_length=MAX_LEN, padding='max_length',
truncation=True, return_offsets_mapping=True)
train_tokens[id_num,] = tokens['input_ids']
train_attention[id_num,] = tokens['attention_mask']
# FIND TARGETS IN TEXT AND SAVE IN TARGET ARRAYS
offsets = tokens['offset_mapping']
offset_index = 1
df = train.loc[train.id==n]
for index,row in df.iterrows():
a = row.discourse_start
b = row.discourse_end
if offset_index>len(offsets)-1:
break
c = offsets[offset_index][0]
d = offsets[offset_index][1]
beginning = True
while b>c:
if (c>=a)&(b>=d):
k = target_map[row.discourse_type]
if beginning:
targets_b[k][id_num][offset_index] = 1
beginning = False
else:
targets_i[k][id_num][offset_index] = 1
offset_index += 1
if offset_index>len(offsets)-1:
break
c = offsets[offset_index][0]
d = offsets[offset_index][1]
targets = np.zeros((len(IDS),MAX_LEN,15), dtype='int32')
for k in range(7):
targets[:,:,2*k] = targets_b[k]
targets[:,:,2*k+1] = targets_i[k]
targets[:,:,14] = 1-np.max(targets,axis=-1)
np.save(DATA_DIR + f"targets_{max_len}.npy",targets)
np.save(DATA_DIR + f"tokens_{max_len}.npy",train_tokens)
np.save(DATA_DIR + f"attention_{max_len}.npy",train_attention)
```
| github_jupyter |
Model of bouting as published in the paper
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
from matplotlib import cm
matplotlib.rcParams['figure.figsize'] = [12, 7]
matplotlib.rcParams['text.usetex'] = False
```
Parameters:
```
h = 0.005
tau1 = tau2 = tau = 0.06
sigma = 10.0
rho1 = -1.0
rho2 = 1.0
T = 50000
```
Simulation using Euler's method
```
# Map
f = lambda t, x: np.array([-tau*x[0] + (1.0 - x[0])*np.exp(-sigma*(x[2] - rho1)**2),
-tau*x[1] + (1.0 - x[1])*np.exp(-sigma*(x[2] - rho2)**2),
-(x[2] - rho1)*(x[2] - rho2)*(x[2] - (rho1+rho2)/2.0) -
0.5*(1-x[0])*(x[2] - rho1) - 0.5*(1-x[1])*(x[2] - rho2)])
def integrate( f, T, x0 ):
X = np.zeros((len(x0), T))
time = np.zeros(T)
X[:,0] = x0
for i in range(T-1):
k1 = f(time[i], X[:,i])
k2 = f(time[i] + h/2.0, X[:,i] + h*k1/2.0)
k3 = f(time[i] + h/2.0, X[:,i] + h*k2/2.0)
k4 = f(time[i] + h, X[:,i] + h*k3)
X[:, i+1] = X[:,i] + h*(k1 + 2*k2 + 2*k3 + k4)/6.0
time[i+1] = time[i] + h
return time, X
t, X = integrate( f, T, [1.0, 0.0, 0.5] )
fig, ax = plt.subplots(3, 1)
ax[0].plot(t, X[0,:])
ax[0].set_xlabel('time')
ax[0].set_ylabel('alpha_1')
ax[0].set_ylim((0, 1))
ax[1].plot(t, X[1,:])
ax[1].set_xlabel('time')
ax[1].set_ylabel('alpha_2')
ax[1].set_ylim((0, 1))
ax[2].plot(X[0,:],X[1,:])
# ax[2].plot(t, X[2,:])
ax[2].set_xlabel('time')
ax[2].set_ylabel('rho')
# ax[2].set_ylim((-1, 1))
print np.max(X[0,int(2*T/3):])-np.min(X[0,int(2*T/3):])
```
Now adding the gradient:
```
tau1 = 0.01
tau2 = 0.03
rho1 = 0.0
rho2 = 1.0
x1 = 1.0
x2 = -1.0
gs = 1.0
G = lambda x: np.exp(-gs*(x - x1)**2)
F = lambda x: np.exp(-gs*(x - x2)**2)
DG = lambda x: -2.0*gs*(x - x1)*np.exp(-gs*(x - x1)**2)
DF = lambda x: -2.0*gs*(x - x2)*np.exp(-gs*(x - x2)**2)
g = lambda t, y: np.array([-tau1*y[0] + (1.0 - y[0])*G(y[3]),
-tau2*y[1] + (1.0 - y[1])*F(y[3]),
-(y[2] - rho1)*(y[2] - rho2)*(y[2] - (rho1+rho2)/2.0) - 0.5*(1-y[0])*(y[2] - rho1) - 0.5*(1-y[1])*(y[2] - rho2),
1.0*(y[2]*DF(y[3]) + (1.0 - y[2])*DG(y[3])) - 0.0*( y[2]*DG(y[3]) - (1.0 - y[2])*DF(y[3])) ])
# g = lambda t, y: np.array([0,0,DF(y[2]),DG(y[3]) ])
t, X = integrate( g, T, [0.0, 0.0, 0.3, 0.0] )
x = np.linspace(-1.5, 1.5, 100)
plt.plot(x, DF(x))
plt.plot(x, F(x))
plt.plot(x, DG(x))
plt.plot(x, G(x))
plt.show()
fig, ax = plt.subplots(2, 2)
ax[0,0].plot(t, X[0,:])
ax[0,0].set_xlabel('time')
ax[0,0].set_ylabel('alpha_1')
ax[0,0].set_ylim((0, 1))
ax[0,1].plot(t, X[1,:])
ax[0,1].set_xlabel('time')
ax[0,1].set_ylabel('alpha_2')
ax[0,1].set_ylim((0, 1))
ax[1,0].plot(t, X[2,:])
ax[1,0].set_xlabel('time')
ax[1,0].set_ylabel('rho')
ax[1,0].set_ylim((0, 1))
ax[1,1].plot(t, X[3,:])
ax[1,1].set_xlabel('time')
ax[1,1].set_ylabel('x')
ax[1,1].set_ylim((-1.1, 1.1))
```
No simulation of the potential in a two dimensional domain and the proof that the corresponding coordinates stay on a line (Are affine)
```
a = 0.0
b = 0.0
r1 = np.array([1, 0])
r2 = np.array([0, 1])
nr = lambda X, Y, r: (X - r[0])**2 + (Y - r[1])**2
phi = lambda X,Y, a, b:nr(X,Y,r1)*nr(X,Y,r2) + a*nr(X,Y,r1)**2 + b*nr(X,Y,r2)
dp = lambda rho, a, b: -0.1*np.array([2*(rho[0]-1)*(rho[0]**2 + (rho[1] - 1)**2) + 2*rho[0]*((rho[0] - 1)**2 + rho[1]**2) + 2*a*(rho[0]-1) + 2*b*rho[0],
2*(rho[1]-1)*(rho[1]**2 + (rho[0] - 1)**2) + 2*rho[1]*((rho[1] - 1)**2 + rho[0]**2) + 2*b*(rho[1]-1) + 2*a*rho[1]])
T = 10000
rho = np.zeros((2, T))
rho0 = [0.2, 0.51]
rho[:,0] = rho0
x = y = np.arange(-0.7, 1.9, 0.02)
X, Y = np.meshgrid(x, y)
Z = phi(X,Y, a, b)
fig, ax = plt.subplots(1,2)
ax[0].contourf(X, Y, Z, levels = np.arange(0, 1.5, 0.01), cmap = plt.cm.jet)
for i in range(T-1):
rho[:,i + 1] = rho[:,i] + h*dp(rho[:,i], a , b)
if i > 4000:
a += 0.01*(1 - a)
ax[0].plot(rho[0,:], rho[1,:], 'w--', linewidth = 3.0)
ax[0].plot(rho[0,0], rho[1,0], 'w.', markersize = 15.0)
ax[0].set_xlabel('motivation1')
ax[0].set_ylabel('motivation2')
ax[0].set_title('Motivational phase space')
ax[1].plot(rho[0,:] + rho[1,:], linewidth = 2.0, label = 'rho1 + rho2')
ax[1].plot(rho[0,:], linewidth = 2.0, label = 'rho1')
ax[1].plot(rho[1,:], linewidth = 2.0, label = 'rho1')
ax[1].legend()
ax[1].set_xlabel('step')
ax[1].set_title('Conservation law')
```
Now simulation in the two dimensional plane
```
T = 80000
mu1 = 0.01
mu2 = 0.01
gamma1 = 1.0
gamma2 = 1.0
e = 1.0
rho1 = np.array([1.0, 0.0])
rho2 = np.array([0.0, 1.0])
L = 1.0
x1 = np.array([1.0, 1.0])
x2 = np.array([-1.0, -1.0])
gs = 0.5
G = lambda x,y: np.exp(-gs*((x - x1[0])**2 + (y - x1[1])**2))
F = lambda x,y: np.exp(-gs*((x - x2[0])**2 + (y - x2[1])**2))
# G = lambda x,y: (gs/np.pi)*1.0/((x - x1[0])**2 + (y - x1[1])**2 + gs**2)
# F = lambda x,y: (gs/np.pi)*1.0/((x - x2[0])**2 + (y - x2[1])**2 + gs**2)
DG = lambda x: np.array([-2.0*gs*(x[0] - x1[0])*np.exp(-gs*np.linalg.norm(x - x1)**2),
-2.0*gs*(x[1] - x1[1])*np.exp(-gs*np.linalg.norm(x - x1)**2)])
DF = lambda x: np.array([-2.0*gs*(x[0] - x2[0])*np.exp(-gs*np.linalg.norm(x - x2)**2),
-2.0*gs*(x[1] - x2[1])*np.exp(-gs*np.linalg.norm(x - x2)**2)])
# DG = lambda x: np.array([-2.0*(gs/np.pi)*(x[0] - x1[0])*1.0/(np.linalg.norm(x - x1)**2 + gs**2),
# -2.0*(gs/np.pi)*(x[1] - x1[1])*1.0/(np.linalg.norm(x - x1)**2 + gs**2)])
# DF = lambda x: np.array([-2.0*(gs/np.pi)*(x[0] - x2[0])*1.0/(np.linalg.norm(x - x2)**2 + gs**2),
# -2.0*(gs/np.pi)*(x[1] - x2[1])*1.0/(np.linalg.norm(x - x2)**2 + gs**2)])
c = 1.0
def g(t, theta):
a1 = theta[0]
a2 = theta[1]
rhox = theta[2]
rhoy = theta[3]
x = theta[4]
y = theta[5]
psy = np.array([e*(-mu1*a1 + (gamma1)*(1-a1)*(np.exp(-sigma*((rhoy)**2))*(0.01 + F(x, y)))),
e*(-mu2*a2 + (gamma2)*(1-a2)*(np.exp(-sigma*((rhox)**2))*(0.01 + G(x, y)))),
-2.0*L*((rhox-c)*(rhox**2 + (rhoy - c)**2) + rhox*((rhox - c)**2 + rhoy**2) + (1-a1)*(rhox-c) + (1-a2)*rhox),
-2.0*L*((rhoy-c)*(rhoy**2 + (rhox - c)**2) + rhoy*((rhoy - c)**2 + rhox**2) + (1-a2)*(rhoy-c) + (1-a1)*rhoy)])
motor = 1.0*(rhox*DF(np.array([x, y])) + rhoy*DG(np.array([x, y])))
return np.concatenate([psy, motor])
#- (1-y[3])*DG(np.array([y[4], y[5]])) - (1.0-y[2])*DF(np.array([y[4], y[5]]))
# g = lambda t, y: np.array([0,0,DF(y[2]),DG(y[3]) ])
t, X = integrate( g, T, [0.0, 1.0, 0.3, 1.0, 1.0, -0.5] )
print(X)
fig, ax = plt.subplots(2,2)
x = y = np.arange(-2.5, 2.9, 0.02)
xx, yy = np.meshgrid(x, y)
Z = G(xx,yy)
# ax[0,0].contourf(xx, yy, Z, levels = np.arange(0, 1.5, 0.01), cmap = plt.cm.jet)
Z = np.maximum(F(xx,yy), G(xx, yy))
ax[0,0].contourf(xx, yy, Z, 10, cmap = plt.cm.jet)
ax[0,0].plot(X[4,:], X[5,:], color = 'white')
ax[0,0].set_xlabel('x')
ax[0,0].set_ylabel('y')
ax[0,0].set_title('World')
ax[0,1].plot(t, X[2,:])
ax[0,1].plot(t, X[3,:])
ax[0,1].plot(t, X[3,:]+X[2,:], 'k--', linewidth = 3.0 )
ax[0,1].set_xlabel('rhox')
ax[0,1].set_ylabel('rhoy')
ax[0,1].set_title('Motivation')
ax[1,0].plot(t, X[0,:])
ax[1,0].axis([0, T*h, 0, 1])
ax[1,0].set_xlabel('time')
ax[1,0].set_ylabel('a')
ax[1,0].set_title('Drive 1')
ax[1,1].plot(t, X[1,:])
ax[1,1].axis([0, T*h, 0, 1])
ax[1,1].set_xlabel('time')
ax[1,1].set_ylabel('b')
ax[1,1].set_title('Drive 2')
# fig = figure()
fig, ax = plt.subplots(1,2)
rhox = 0.4
rhoy = 1 - rhox
x1 = np.array([1.0, 1.0])
x2 = np.array([-1.0, -1.0])
DGx = lambda x,y: -2.0*gs*(x - x1[0])*np.exp(-gs*((x - x1[0])**2 + (y - x1[1])**2))
DGy = lambda x,y: -2.0*gs*(y - x1[1])*np.exp(-gs*((x - x1[0])**2 + (y - x1[1])**2))
DFx = lambda x,y: -2.0*gs*(x - x2[0])*np.exp(-gs*((x - x2[0])**2 + (y - x2[1])**2))
DFy = lambda x,y: -2.0*gs*(y - x2[1])*np.exp(-gs*((x - x2[0])**2 + (y - x2[1])**2))
dHx = lambda x, y: rhox*DGx(x, y) + rhoy*DFx(x, y)
dHy = lambda x, y: rhox*DGy(x, y) + rhoy*DFy(x, y)
x = y = np.arange(-2.5, 2.9, 0.02)
xx, yy = np.meshgrid(x, y)
Zx = dHx(xx, yy)
Zy = dHy(xx, yy)
b1 = lambda x: -np.log(-(rhox/rhoy)*(x - x1[0])/(x - x2[0]))/(2.0*gs) + (x1[0]**2 - x2[0]**2 + x1[1]**2 - x2[1]**2)/2.0
yb1 = lambda x: -((x1[0] - x2[0])/(x1[1] - x2[1]))*x + b1(x)/(x1[1] - x2[1])
b2 = lambda y: -np.log(-(rhox/rhoy)*(y - x1[1])/(y - x2[1]))/(2.0*gs) + (x1[0]**2 - x2[0]**2 + x1[1]**2 - x2[1]**2)/2.0
yb2 = lambda y: -((x1[1] - x2[1])/(x1[0] - x2[0]))*y + b1(y)/(x1[0] - x2[0])
ax[0].contourf(xx, yy, Zx, 10, cmap = plt.cm.jet)
ax[0].plot(x, yb1(x))
# plt.contourf(xx, yy, Zy, levels =[-2,-1,0, 1], cmap = plt.cm.jet)
# ax[0].colorbar()
ax[0].axis([-1.5, 2.0, -1.5, 2.0])
ax[1].contourf(xx, yy, Zy, 10, cmap = plt.cm.jet)
ax[1].plot(yb2(y), y)
# plt.contourf(xx, yy, Zy, levels =[-2,-1,0, 1], cmap = plt.cm.jet)
# ax[0].colorbar()
ax[1].axis([-1.5, 2.0, -1.5, 2.0])
plt.show()
```
Now dynamics of a simple agent
```
T = 10000
g = lambda t, theta: np.array([dHx(theta[0], theta[1]), dHy(theta[0], theta[1])])
s = 0.3
print(yb2(s))
t, X = integrate(g, T, [1.3, -0.8])
x = y = np.arange(-2.5, 2.9, 0.02)
xx, yy = np.meshgrid(x, y)
# ax[0,0].contourf(xx, yy, Z, levels = np.arange(0, 1.5, 0.01), cmap = plt.cm.jet)
Z = np.maximum(F(xx,yy), G(xx, yy))
plt.contourf(xx, yy, Z, 10, cmap = plt.cm.hot)
plt.plot(x, yb1(x), '--', color = (0,0,0), linewidth = 2.0)
plt.plot(yb2(y), y, '--', color = (0,0,0), linewidth = 2.0)
plt.plot(X[0,:], X[1,:], color = (0,0,1), linewidth = 3.0)
plt.plot(X[0,0], X[1,0], '.', color = (0,0,1), markersize = 15)
plt.axis([-2.5, 2.9, -2.5, 2.9])
plt.xlabel('x')
plt.ylabel('y')
plt.title('Physical space')
plt.show()
```
The condition for existence of equilibrium
```
# x = np.linspace(-0.9, 1, 100)
from scipy.optimize import fsolve
a = (x1[0]**2 - x2[0]**2)/2.0 + (x1[1]**2 - x2[1]**2)/2.0
g1 = lambda x, y: -(rhox/rhoy)*((y - x1[1])/(y - x2[1]))*np.exp(2.0*gs*(x1[1]-x2[1])*y) - np.exp(-2*gs*((x1[0] - x2[0])*x - a))
g2 = lambda x, y: -(rhox/rhoy)*((x - x1[0])/(x - x2[0]))*np.exp(2.0*gs*(x1[0]-x2[0])*x) - np.exp(-2*gs*((x1[1] - x2[1])*y - a))
def equations(p):
x, y = p
return (g1(x, y), g2(x, y))
res = fsolve(equations, (-0.9, -0.9))
print (res)
f1 = lambda x: np.exp(-2*gs*((x1[0] - x2[0])*x - a)) + (rhox/rhoy)*((x - x1[0])/(x - x2[0]))*np.exp(2.0*gs*(x1[0]-x2[0])*x)
f2 = lambda y: -(rhox/rhoy)*((y - x1[1])/(y - x2[1]))*np.exp(2.0*gs*(x1[1]-x2[1])*y) - np.exp(-2*gs*((x1[1] - x2[1])*y - a))
plt.plot(x, f1(x))
plt.plot(x, f2(x))
plt.plot(res[0], 0, 'k.', markersize = 15)
plt.axis([-3.0, 3.0, -1.0, 1.0])
plt.show()
```
A simplified phase space
```
gs =0.1
x1 = -1.0
x2 = 1.0
a = 0.1
b = 0.9
rho1 = -1.0
rho2 = 1.0
DG = lambda x: -2.0*gs*(x - x1)*np.exp(-gs*((x - x1)**2))
DF = lambda x: -2.0*gs*(x - x2)*np.exp(-gs*((x - x2)**2))
DH = lambda rho, x: rho*DF(x) - (1 - rho)*DG(x)
frho = lambda rho, x: -(rho - rho1)*(rho - rho2)*(rho - (rho1+rho2)/2.0) - \
0.5*(1-a)*(rho - rho1) - 0.5*(1-b)*(rho - rho2)
xr = np.linspace(-2.0, 2.0, 100)
rhor = np.linspace(-1, 1, 100)
Rg,Xg = np.meshgrid(rhor, xr)
plt.contour( Rg, Xg, DH(Rg, Xg), [0] )
plt.contour( Rg, Xg, frho(Rg, Xg), [0] )
plt.xlabel('rho')
plt.ylabel('x')
plt.show()
```
Now the drives
```
mu = 0.1
gamma = 0.7
b = 0.4
rho1 = -1.0
rho2 = 1.0
a1 = lambda rho: -gamma*rho/(mu -gamma*rho)
Jp = lambda rho, a: -(rho - rho1)*(rho - rho2)*(rho - (rho1+rho2)/2.0) - 0.5*(1-a)*(rho - rho1) - 0.5*(1-b)*(rho - rho2)
ar = np.linspace(0.0, 1.0, 100)
rhor = np.linspace(-1, 1, 100)
Rg,Ag = np.meshgrid(rhor, ar)
plt.plot(rhor, a1(rhor))
plt.contour( Rg, Ag, Jp(Rg, Ag), [0])
plt.legend(("a1", "Potential"))
plt.axis([-1, 1, 0, 1])
plt.show()
```
Comparing the two systems' evolution
```
T = 30000
tau1 = tau2 = tau = 0.01
sigma = 20.0
rho1 = 0.0
rho2 = 1.0
f1 = lambda t, x: np.array([-tau*x[0] + (1.0 - x[0])*np.exp(-sigma*(x[2] - rho1)**2),
-tau*x[1] + (1.0 - x[1])*np.exp(-sigma*(x[2] - rho2)**2),
-8.0*((x[2] - rho1)*(x[2] - rho2)*(x[2] - (rho1+rho2)/2.0) +
(1.0/4.0)*(1-x[0])*(x[2] - rho1) + (1.0/4.0)*(1-x[1])*(x[2] - rho2))])
L = 1.0
def f2(t, theta):
a1 = theta[0]
a2 = theta[1]
rhox = theta[2]
rhoy = theta[3]
psy = np.array([(-tau*a1 + 1.0*(1-a1)*np.exp(-sigma*((rhoy)**2))),
(-tau*a2 + 1.0*(1-a2)*np.exp(-sigma*((rhox)**2 ))),
-2.0*L*((rhox-c)*(rhox**2 + (rhoy - c)**2) + rhox*((rhox - c)**2 + rhoy**2) + (1-a1)*(rhox-c) + (1-a2)*rhox),
-2.0*L*((rhoy-c)*(rhoy**2 + (rhox - c)**2) + rhoy*((rhoy - c)**2 + rhox**2) + (1-a2)*(rhoy-c) + (1-a1)*rhoy)])
return psy
t, X1 = integrate( f2, T, [1.0, 0.0, 0.5, 0.5] )
t, X2 = integrate( f1, T, [1.0, 0.0, 0.5] )
fig, ax = plt.subplots(3,1, figsize = (8, 10))
# ax[0].plot(t, X1[2,:] , 'k')
ax[0].plot(t, X1[3,:], 'k', label = 'Full system')
ax[0].plot(t, X2[2,:], 'r--', label = 'Reduced system')
ax[0].set_title('Motivational dynamics')
ax[0].legend()
ax[1].plot(t,X1[0,:], 'k', label = 'Full system')
ax[1].plot(t, X2[0,:], 'r--', label = 'Reduced system')
ax[1].set_ylabel('a1')
ax[1].axis([0, T*h, 0, 1.3])
ax[1].set_title('Drive 1')
ax[1].legend()
ax[2].plot(t,X1[1,:], 'k', label = 'Full system')
ax[2].plot(t,X2[1,:], 'r--', label = 'Reduced system')
ax[2].set_ylabel('a2')
ax[2].axis([0, T*h, 0, 1])
ax[2].set_xlabel('time')
ax[2].set_title('Drive 2')
ax[2].legend()
plt.show()
fig = plt.figure()
T = 100000
tau = 4.0
gamma1 = 30.0
sigma = 10.0
a2 = 0.4#0.74357
k = -0.5
rho1 = 0.0
rho2 = 1.0
epsilon = 0.01
G = lambda rho: np.exp(-sigma*((rho - rho1)**2))
a1_f = lambda rho: (gamma1*G(rho) )/(tau + gamma1*G(rho))
a_p = lambda rho, a2: 4*((rho - rho1)*(rho- rho2)*(rho - (rho1+rho2)/2.0) + (1.0/4.0)*(1-a2)*(rho - rho2))/(rho - rho1) + 1
rhor = np.linspace(0.01, 2, 100)
L = 1.0
# f1 = lambda t, x: np.array([-tau*(x[0]) + gamma1*(1.0 - (x[0]))*(G(x[2])),
# 0.0,
# -8.0*L*((x[2] - rho1)*(x[2] - rho2)*(x[2] - (rho1+rho2)/2.0) +
# (1.0/4.0)*(1-x[0])*(x[2] - rho1) + (1.0/4.0)*(1-x[1])*(x[2] - rho2))])
f1 = lambda t, x: np.array([epsilon*(-tau*(x[0]) + gamma1*(1 - x[0])*(G(x[2]))),
0.0,
-8.0*L*((x[2] - rho1)*(x[2] - rho2)*(x[2] - (rho1+rho2)/2.0) +
(1.0/4.0)*(1-x[0])*(x[2] - rho1) + (1.0/4.0)*(1-x[1])*(x[2] - rho2))])
t,X = integrate( f1, T, [0.7, a2, 0.3] )
# plt.contour( Rg, Ag, Jp(Rg, Ag, 0.9), [0])
plt.plot(rhor, a1_f(rhor), linewidth = 2.0)
plt.plot(rhor, a_p(rhor, a2), linewidth = 2.0)
rho_c = np.sqrt(-(1.0/(2.0*sigma))*np.log(tau**2/gamma1**2))
print(rho_c)
plt.plot([rho_c, rho_c],[0, 1], 'k--')
plt.plot(X[2,:], X[0, :], 'r')
plt.plot(X[2,0], X[0, 0], 'r*')
plt.axis([0, 1, -0.1, 1.8])
plt.show()
plt.figure()
plt.plot(t, X[0,:])
plt.plot(t, X[2,:])
plt.show()
```
Trying to get a fast slow equivalent
```
T = 50000
tau1 = tau2 = tau = 4.0
sigma = 10.0
rho1 = 0.0
rho2 = 1.0
epsilon = 0.001
gamma = 30.0
f1 = lambda t, x: np.array([epsilon*(-tau*x[0] + gamma*(1.0 - x[0])*G(x[2])),
epsilon*(-tau*x[1] + gamma*(1.0 - x[1])*G(1-x[2])),
-8.0*((x[2] - rho1)*(x[2] - rho2)*(x[2] - (rho1+rho2)/2.0) +
(1.0/4.0)*(1-x[0])*(x[2] - rho1) + (1.0/4.0)*(1-x[1])*(x[2] - rho2))])
t, X1 = integrate( f1, T, [0.0, 1.0, 0.5] )
fig, ax = plt.subplots(3,1, figsize = (8, 10))
a = tau
b = gamma
ss = lambda t: (b/(a + b))*(1 - np.exp(-epsilon*(a + b)*t))
sf = lambda t: np.exp(-epsilon*a*t)
# ax[0].plot(t, X1[2,:] , 'k')
ax[0].plot(t, X1[2,:], 'k')
ax[0].set_title('Motivational dynamics')
ax[1].plot(t,X1[0,:], 'k')
ax[1].plot(t, ss(t))
ax[1].plot(t, sf(t))
ax[1].plot(t, (sf(t) + ss(t))/2.0, 'r--')
vm = 0.5*((1.0/a + b/(a + b) - b/((a + b)**2)))
ax[1].plot([0, T*h], [vm, vm], 'k--')
ax[1].set_ylabel('a1')
ax[1].axis([0, T*h, 0, 1.3])
ax[1].set_title('Drive 1')
ax[2].plot(t,X1[1,:], 'k')
ax[2].set_ylabel('a2')
ax[2].axis([0, T*h, 0, 1])
ax[2].set_xlabel('time')
ax[2].set_title('Drive 2')
plt.show()
```
Exploring the nullcline parameters
```
a = 4.0
b = 30.0
a2 = 0.73
G = lambda rho: np.exp(-sigma*(rho**2))
a1_f = lambda rho, a, b: b*G(rho)/(a + b*G(rho))
a_p = lambda rho, a2: 2*(rho*(rho-1.0)*(2*rho - 1.0) + (1.0/2.0)*(1-a2)*(rho - 1))/(rho) + 1
ass = np.linspace(0, 10, 30)
bs = np.linspace(0, 50, 30)
ps = np.linspace(0, 1, 30)
rho = np.linspace(0.01, 1, 100)
fig, ax = plt.subplots(3, 1)
ax[0].plot(rho, a_p(rho, a2))
for i in range(len(ps)):
c = float(i)/float(len(ps))
ac = b*(1 - ps[i])/ps[i]
ax[0].plot(rho, a1_f(rho, ac, b), color = (c, c, 0))
ax[0].axis([0, 1, 0, 1.5])
ax[1].plot(rho, a_p(rho, a2))
for i in range(len(ps)):
bc = a*ps[i]/(1 - ps[i])
c = float(i)/float(len(ps))
ax[1].plot(rho, a1_f(rho, a, bc), color = (c, c, 0))
ax[1].axis([0, 1, 0, 1.5])
plt.show()
```
| github_jupyter |
```
# Import Splinter and BeautifulSoup
from splinter import Browser
from bs4 import BeautifulSoup as soup
from webdriver_manager.chrome import ChromeDriverManager
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
# Visit the mars nasa news site
url = 'https://redplanetscience.com'
browser.visit(url)
# Optional delay for loading the page
browser.is_element_present_by_css('div.list_text', wait_time=1)
# Convert the browser html to soup and quit the browser
html = browser.html
news_soup = soup(html, 'html.parser')
slide_elem = news_soup.select_one('div.list_text')
slide_elem.find("div",class_="content_title")
# Use the parent element to find the first `a` tag and save it as `news_title`
news_title = slide_elem.find('div', class_='content_title').get_text()
news_title
# Use the parent element to find the paragraph text
news_p = slide_elem.find('div', class_='article_teaser_body').get_text()
news_p
# Import Splinter and BeautifulSoup
from splinter import Browser
from bs4 import BeautifulSoup as soup
from webdriver_manager.chrome import ChromeDriverManager
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
### Featured Image
# Visit URL
# Visit URL
url = 'https://spaceimages-mars.com'
browser.visit(url)
# Import Splinter and BeautifulSoup
from splinter import Browser
from bs4 import BeautifulSoup as soup
from webdriver_manager.chrome import ChromeDriverManager
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
### Featured Image
# Visit URL
# Visit URL
url = 'https://spaceimages-mars.com'
browser.visit(url)
# Find and click the full image button
full_image_elem = browser.find_by_tag('button')[1]
full_image_elem.click()
html = browser.html
img_soup = soup(html, 'html.parser')
# Find the relative image url
img_url_rel = img_soup.find('img', class_='fancybox-image').get('src')
img_url_rel
# Use the base URL to create an absolute URL
img_url = f'https://spaceimages-mars.com/{img_url_rel}'
print(img_url)
```
# Mars Facts
```
import pandas as pd
import os
df = pd.read_html('https://galaxyfacts-mars.com')[0]
df.columns=['description', 'Mars', 'Earth']
df.set_index('description', inplace=True)
df
df.to_html()
# Import Splinter and BeautifulSoup
from splinter import Browser
from bs4 import BeautifulSoup as soup
from webdriver_manager.chrome import ChromeDriverManager
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
### Featured Image
# Visit URL
# Visit URL
url = 'https://spaceimages-mars.com'
browser.visit(url)
# Find and click the full image button
full_image_elem = browser.find_by_tag('button')[1]
full_image_elem.click()
html = browser.html
img_soup = soup(html, 'html.parser')
# Find the relative image url
img_url_rel = img_soup.find('img', class_='fancybox-image').get('src')
img_url_rel
img_url = f'https://spaceimages-mars.com/{img_url_rel}'
print(img_url)
df = pd.read_html('https://galaxyfacts-mars.com')[0]
df.columns=['description', 'Mars', 'Earth']
df.set_index('description', inplace=True)
print(df.to_html())
browser.quit()
browser.quit()
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Transfer learning with TensorFlow Hub
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/images/transfer_learning_with_hub"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/images/transfer_learning_with_hub.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/images/transfer_learning_with_hub.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/images/transfer_learning_with_hub.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
[TensorFlow Hub](http://tensorflow.org/hub) is a way to share pretrained model components. See the [TensorFlow Module Hub](https://tfhub.dev/) for a searchable listing of pre-trained models. This tutorial demonstrates:
1. How to use TensorFlow Hub with `tf.keras`.
1. How to do image classification using TensorFlow Hub.
1. How to do simple transfer learning.
## Setup
```
import matplotlib.pylab as plt
import tensorflow as tf
!pip install -U tf-hub-nightly
!pip install tfds-nightly
import tensorflow_hub as hub
from tensorflow.keras import layers
```
## An ImageNet classifier
### Download the classifier
Use `hub.module` to load a mobilenet, and `tf.keras.layers.Lambda` to wrap it up as a keras layer. Any [TensorFlow 2 compatible image classifier URL](https://tfhub.dev/s?q=tf2&module-type=image-classification) from tfhub.dev will work here.
```
classifier_url ="https://tfhub.dev/google/tf2-preview/mobilenet_v2/classification/2" #@param {type:"string"}
IMAGE_SHAPE = (224, 224)
classifier = tf.keras.Sequential([
hub.KerasLayer(classifier_url, input_shape=IMAGE_SHAPE+(3,))
])
```
### Run it on a single image
Download a single image to try the model on.
```
import numpy as np
import PIL.Image as Image
grace_hopper = tf.keras.utils.get_file('image.jpg','https://storage.googleapis.com/download.tensorflow.org/example_images/grace_hopper.jpg')
grace_hopper = Image.open(grace_hopper).resize(IMAGE_SHAPE)
grace_hopper
grace_hopper = np.array(grace_hopper)/255.0
grace_hopper.shape
```
Add a batch dimension, and pass the image to the model.
```
result = classifier.predict(grace_hopper[np.newaxis, ...])
result.shape
```
The result is a 1001 element vector of logits, rating the probability of each class for the image.
So the top class ID can be found with argmax:
```
predicted_class = np.argmax(result[0], axis=-1)
predicted_class
```
### Decode the predictions
We have the predicted class ID,
Fetch the `ImageNet` labels, and decode the predictions
```
labels_path = tf.keras.utils.get_file('ImageNetLabels.txt','https://storage.googleapis.com/download.tensorflow.org/data/ImageNetLabels.txt')
imagenet_labels = np.array(open(labels_path).read().splitlines())
plt.imshow(grace_hopper)
plt.axis('off')
predicted_class_name = imagenet_labels[predicted_class]
_ = plt.title("Prediction: " + predicted_class_name.title())
```
## Simple transfer learning
Using TF Hub it is simple to retrain the top layer of the model to recognize the classes in our dataset.
### Dataset
For this example you will use the TensorFlow flowers dataset:
```
data_root = tf.keras.utils.get_file(
'flower_photos','https://storage.googleapis.com/download.tensorflow.org/example_images/flower_photos.tgz',
untar=True)
```
The simplest way to load this data into our model is using `tf.keras.preprocessing.image.ImageDataGenerator`,
All of TensorFlow Hub's image modules expect float inputs in the `[0, 1]` range. Use the `ImageDataGenerator`'s `rescale` parameter to achieve this.
The image size will be handled later.
```
image_generator = tf.keras.preprocessing.image.ImageDataGenerator(rescale=1/255)
image_data = image_generator.flow_from_directory(str(data_root), target_size=IMAGE_SHAPE)
```
The resulting object is an iterator that returns `image_batch, label_batch` pairs.
```
for image_batch, label_batch in image_data:
print("Image batch shape: ", image_batch.shape)
print("Label batch shape: ", label_batch.shape)
break
```
### Run the classifier on a batch of images
Now run the classifier on the image batch.
```
result_batch = classifier.predict(image_batch)
result_batch.shape
predicted_class_names = imagenet_labels[np.argmax(result_batch, axis=-1)]
predicted_class_names
```
Now check how these predictions line up with the images:
```
plt.figure(figsize=(10,9))
plt.subplots_adjust(hspace=0.5)
for n in range(30):
plt.subplot(6,5,n+1)
plt.imshow(image_batch[n])
plt.title(predicted_class_names[n])
plt.axis('off')
_ = plt.suptitle("ImageNet predictions")
```
See the `LICENSE.txt` file for image attributions.
The results are far from perfect, but reasonable considering that these are not the classes the model was trained for (except "daisy").
### Download the headless model
TensorFlow Hub also distributes models without the top classification layer. These can be used to easily do transfer learning.
Any [Tensorflow 2 compatible image feature vector URL](https://tfhub.dev/s?module-type=image-feature-vector&q=tf2) from tfhub.dev will work here.
```
feature_extractor_url = "https://tfhub.dev/google/tf2-preview/mobilenet_v2/feature_vector/2" #@param {type:"string"}
```
Create the feature extractor.
```
feature_extractor_layer = hub.KerasLayer(feature_extractor_url,
input_shape=(224,224,3))
```
It returns a 1280-length vector for each image:
```
feature_batch = feature_extractor_layer(image_batch)
print(feature_batch.shape)
```
Freeze the variables in the feature extractor layer, so that the training only modifies the new classifier layer.
```
feature_extractor_layer.trainable = False
```
### Attach a classification head
Now wrap the hub layer in a `tf.keras.Sequential` model, and add a new classification layer.
```
model = tf.keras.Sequential([
feature_extractor_layer,
layers.Dense(image_data.num_classes)
])
model.summary()
predictions = model(image_batch)
predictions.shape
```
### Train the model
Use compile to configure the training process:
```
model.compile(
optimizer=tf.keras.optimizers.Adam(),
loss=tf.keras.losses.CategoricalCrossentropy(from_logits=True),
metrics=['acc'])
```
Now use the `.fit` method to train the model.
To keep this example short train just 2 epochs. To visualize the training progress, use a custom callback to log the loss and accuracy of each batch individually, instead of the epoch average.
```
class CollectBatchStats(tf.keras.callbacks.Callback):
def __init__(self):
self.batch_losses = []
self.batch_acc = []
def on_train_batch_end(self, batch, logs=None):
self.batch_losses.append(logs['loss'])
self.batch_acc.append(logs['acc'])
self.model.reset_metrics()
steps_per_epoch = np.ceil(image_data.samples/image_data.batch_size)
batch_stats_callback = CollectBatchStats()
history = model.fit(image_data, epochs=2,
steps_per_epoch=steps_per_epoch,
callbacks=[batch_stats_callback])
```
Now after, even just a few training iterations, we can already see that the model is making progress on the task.
```
plt.figure()
plt.ylabel("Loss")
plt.xlabel("Training Steps")
plt.ylim([0,2])
plt.plot(batch_stats_callback.batch_losses)
plt.figure()
plt.ylabel("Accuracy")
plt.xlabel("Training Steps")
plt.ylim([0,1])
plt.plot(batch_stats_callback.batch_acc)
```
### Check the predictions
To redo the plot from before, first get the ordered list of class names:
```
class_names = sorted(image_data.class_indices.items(), key=lambda pair:pair[1])
class_names = np.array([key.title() for key, value in class_names])
class_names
```
Run the image batch through the model and convert the indices to class names.
```
predicted_batch = model.predict(image_batch)
predicted_id = np.argmax(predicted_batch, axis=-1)
predicted_label_batch = class_names[predicted_id]
```
Plot the result
```
label_id = np.argmax(label_batch, axis=-1)
plt.figure(figsize=(10,9))
plt.subplots_adjust(hspace=0.5)
for n in range(30):
plt.subplot(6,5,n+1)
plt.imshow(image_batch[n])
color = "green" if predicted_id[n] == label_id[n] else "red"
plt.title(predicted_label_batch[n].title(), color=color)
plt.axis('off')
_ = plt.suptitle("Model predictions (green: correct, red: incorrect)")
```
## Export your model
Now that you've trained the model, export it as a saved model:
```
import time
t = time.time()
export_path = "/tmp/saved_models/{}".format(int(t))
model.save(export_path, save_format='tf')
export_path
```
Now confirm that we can reload it, and it still gives the same results:
```
reloaded = tf.keras.models.load_model(export_path)
result_batch = model.predict(image_batch)
reloaded_result_batch = reloaded.predict(image_batch)
abs(reloaded_result_batch - result_batch).max()
```
This saved model can be loaded for inference later, or converted to [TFLite](https://www.tensorflow.org/lite/convert/) or [TFjs](https://github.com/tensorflow/tfjs-converter).
| github_jupyter |
<a href="https://colab.research.google.com/github/JeromeMberia/KNN_and_Naive_Bayes/blob/main/Independent_Project_Week_9(Titanic__Data_Set).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **Titanic Data Set**
## Defining the Question
### Specifying the Data Analytic Question
> Predicting wheather person can survive or not based on it's features
### Defining the Metric for Success
> Getting an accuracy of 80 or above it.
### Understanding the context
> This is titanic data frame does not contain information from the crew, but it
does contain actual ages of half of the passengers. The principal source for data about. Titanic passengers is the Encyclopedia Titanica. The datasets used here were begun by a variety of researchers.
### Recording the Experimental Design
1. Load , read and explore the dataset.
2. Clean the dataset.
3. Perform Exploratory Data Analysis.
4. Create k-nearest neighbors(KNN) modal.
## The Dataset
### Importing the libraries
```
import pandas as pd
import numpy as np
# Visualization
import seaborn as sns
import matplotlib.pyplot as plt
# Machine Learning Models
from sklearn.neighbors import KNeighborsClassifier
# others
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn import preprocessing
from sklearn.preprocessing import Normalizer
# constant
RANDOM_STATE = 42
cv = 10
```
### Reading the dataset
```
Train = pd.read_csv('train.csv')
Test = pd.read_csv('test.csv')
```
### Checking the dataset
#### Train
```
Train.head()
Train.tail()
Train.columns.values.tolist()
Train.nunique()
rows = Train.shape[0]
columns = Train.shape[1]
print(f'This dataset has {columns} columns and {rows} rows')
Train.describe()
Train.info()
Train.dtypes
col = list(Train.columns)
for i in col:
print(f'{i}')
print('')
print(Train[i].unique())
print('****************************************************************************')
print('')
```
#### Test
```
Test.head()
Test.tail()
Test.columns.values.tolist()
Test.nunique()
rows = Test.shape[0]
columns = Test.shape[1]
print(f'This dataset has {columns} columns and {rows} rows')
Test.describe()
Test.info()
Test.dtypes
col = list(Test.columns)
for i in col:
print(f'{i}')
print('')
print(Test[i].unique())
print('****************************************************************************')
print('')
```
## Data Wrangling / Data Cleaning
### Duplicate Records
#### Train
```
Train.duplicated().sum()
```
#### Test
```
Test.duplicated().sum()
```
### Irrelevant Data
```
Train.columns.values.tolist()
Train.head()
dropped = ['PassengerId','Name','Ticket', 'Cabin', 'Fare']
Train = Train.drop(dropped,axis=1)
Test = Test.drop(dropped,axis=1)
```
### Missing Data
```
Train.isnull().sum()
Test.isnull().sum()
Test.loc[(Test['Pclass'] == 1) & (Test['Age'].isnull()), 'Age'] = Test[(Test['Pclass']==1)]['Age'].mean()
Test.loc[(Test['Pclass'] == 2) & (Test['Age'].isnull()), 'Age'] = Test[(Test['Pclass']==2)]['Age'].mean()
Test.loc[(Test['Pclass'] == 3) & (Test['Age'].isnull()), 'Age'] = Test[(Test['Pclass']==3)]['Age'].mean()
Train.loc[(Train['Pclass'] == 1) & (Train['Age'].isnull()), 'Age'] = Train[(Train['Pclass']==1)]['Age'].mean()
Train.loc[(Train['Pclass'] == 2) & (Train['Age'].isnull()), 'Age'] = Train[(Train['Pclass']==2)]['Age'].mean()
Train.loc[(Train['Pclass'] == 3) & (Train['Age'].isnull()), 'Age'] = Train[(Train['Pclass']==3)]['Age'].mean()
Train.loc[(Train['Embarked'].isnull()), 'Embarked'] = Train['Embarked'].mode()
Train['Embarked'] = Train['Embarked'].fillna(Train['Embarked'].mode()[0])
Test.isnull().sum()
Train.isnull().sum()
```
### Outliers
```
df = Train
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
plo = plt.figure(figsize=(20,4))
plo.add_subplot(1,2,1)
sns.distplot(df[column])
plo.add_subplot(1,2,2)
sns.boxplot(df[column])
df = Test
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
plo = plt.figure(figsize=(20,4))
plo.add_subplot(1,2,1)
sns.distplot(df[column])
plo.add_subplot(1,2,2)
sns.boxplot(df[column])
```
## Exploratory Data Analysis(EDA)
### Univariate Analysis
#### Categorical Variable
```
Train.columns
sns.countplot(Train['Sex'])
sns.countplot(Test['Sex'])
sns.countplot(Train['Embarked'])
sns.countplot(Test['Embarked'])
sns.countplot(Train['Pclass'])
sns.countplot(Test['Pclass'])
sns.countplot(Train['Survived'])
```
#### Numerical Variable
**1. Measures of Central Tendency**
```
# 1.1 Mean
numeric = Train[['Age','SibSp','Parch']]
columns = None
columns = list(numeric.columns)
for column in columns:
mean = Train[column].mean()
print(f'This is the mean of the {column} column: ')
print(f'{mean}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = None
columns = list(numeric.columns)
for column in columns:
mean = Test[column].mean()
print(f'This is the mean of the {column} column: ')
print(f'{mean}')
print(' ')
# 1.2 Median
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
median = Train[column].median()
print(f'This is the median of the {column} column: ')
print(f'{median}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
median = Test[column].median()
print(f'This is the median of the {column} column: ')
print(f'{median}')
print(' ')
# 1.3 Mode
df = Train[['Survived', 'Age', 'SibSp', 'Parch']]
columns = list(df.columns)
for column in columns:
mode = Train[column].mode()[0]
print(f'This is the mode of the {column} column: ')
print(f'{mode}')
print(' ')
df = Test[['Age', 'SibSp', 'Parch']]
columns = list(df.columns)
for column in columns:
mode = Test[column].mode()[0]
print(f'This is the mode of the {column} column: ')
print(f'{mode}')
print(' ')
```
**2. Measures of Dispersion**
```
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
## 2.1 Standard Deviation
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
standard_deviation = Train[column].std()
print(f'This is the Standard Deviation of the {column} column: ')
print(f'{standard_deviation}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
standard_deviation = Test[column].std()
print(f'This is the Standard Deviation of the {column} column: ')
print(f'{standard_deviation}')
print(' ')
## 2.2 Variance
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
variance = Train[column].var()
print(f'This is the variance of the {column} column: ')
print(f'{variance}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
variance = Test[column].var()
print(f'This is the variance of the {column} column: ')
print(f'{variance}')
print(' ')
## 2.3 Maximum
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
max = Train[column].max()
print(f'This is the largest value in the {column} column: ')
print(f'{max}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
max = Test[column].max()
print(f'This is the largest value in the {column} column: ')
print(f'{max}')
print(' ')
## 2.4 Minimum
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
min = Train[column].min()
print(f'This is the small value in the {column} column: ')
print(f'{min}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
min = Test[column].min()
print(f'This is the small value in the {column} column: ')
print(f'{min}')
print(' ')
## 2.5 Quantile
### 2.5.1 First Quantile
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
first_quantile = Train[column].quantile(0.25)
print(f'This is the first quantile of the {column} column: ')
print(f'{first_quantile}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
first_quantile = Test[column].quantile(0.25)
print(f'This is the first quantile of the {column} column: ')
print(f'{first_quantile}')
print(' ')
### 2.5.2 Second Quantile / Mean
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
second_quantile = Train[column].quantile(0.5)
print(f'This is the second quantile of the {column} column: ')
print(f'{second_quantile}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
second_quantile = Test[column].quantile(0.5)
print(f'This is the second quantile of the {column} column: ')
print(f'{second_quantile}')
print(' ')
### 2.5.3 Third Quantile
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
third_quantile = Train[column].quantile(0.75)
print(f'This is the third quantile of the {column} column: ')
print(f'{third_quantile}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
third_quantile = Test[column].quantile(0.75)
print(f'This is the third quantile of the {column} column: ')
print(f'{third_quantile}')
print(' ')
```
**3. Descriptions of the distribution**
```
## 3.1 Skewness
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
skewness = Train[column].skew()
print(f'This is the skewness of the {column} column: ')
print(f'{skewness}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
skewness = Test[column].skew()
print(f'This is the skewness of the {column} column: ')
print(f'{skewness}')
print(' ')
## 3.2 Kurtosis
numeric = Train[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
kurtosis = Train[column].kurt()
print(f'This is the kurtosis of the {column} column: ')
print(f'{kurtosis}')
print(' ')
numeric = Test[['Age','SibSp','Parch']]
columns = list(numeric.columns)
for column in columns:
kurtosis = Test[column].kurt()
print(f'This is the kurtosis of the {column} column: ')
print(f'{kurtosis}')
print(' ')
```
### Bivariate Analysis
```
sns.heatmap(Train.corr(),annot=True, linewidths=2, linecolor='white')
plt.show()
sns.heatmap(Test.corr(),annot=True, linewidths=2, linecolor='white')
plt.show()
sns.countplot('Pclass',hue='Survived',data=Train)
plt.title('Pclass:Survived vs Dead')
plt.show()
```
> Most of People that survived come from Pclass 3 .
```
sns.countplot('Embarked',hue='Survived',data=Train)
plt.title('Embarked:Survived vs Dead')
plt.show()
```
> Most of the passangers come form Embarked S
```
sns.countplot('Sex',hue='Survived',data=Train)
plt.title('Sex:Survived vs Dead')
plt.show()
```
>* Most of the people how dead were female
>
>* Most of the people how survived were male
## Implementing the Solution
```
Train['Sex'] = preprocessing.LabelEncoder().fit_transform(Train['Sex'])
Test['Sex'] = preprocessing.LabelEncoder().fit_transform(Test['Sex'])
Train['Embarked'] = preprocessing.LabelEncoder().fit_transform(Train['Embarked'])
Test['Embarked'] = preprocessing.LabelEncoder().fit_transform(Test['Embarked'])
Train.head()
Test.head()
X = Train.drop(['Survived'], axis=1).copy()
y = Train['Survived'].copy()
from sklearn.preprocessing import Normalizer
# Create normalizer
normalizer = Normalizer()
# Transform feature matrix
normalizer.transform(X)
```
### 80% training dataset - 20% testing dataset
```
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state=42)
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
lda = LDA(n_components=1)
x_train = lda.fit_transform(x_train, y_train)
x_test = lda.transform(x_test)
parameters = {'n_neighbors': list(range(1,30)),
'leaf_size': list(range(1,50)),
'metric':['euclidean', 'manhattan', 'chebyshev', 'minkowski', 'wminkowski', 'seuclidean', 'mahalanobis'],
}
knn = KNeighborsClassifier()
grid_search = GridSearchCV(estimator=knn,
param_grid=parameters)
print(grid_search.fit(x_train, y_train))
print('')
print('')
print('The parameters:')
print(grid_search.best_params_)
grid_search.best_params_
par = grid_search.best_params_
leaf_size = par['leaf_size']
n_neighbors = par['n_neighbors']
metric = par['metric']
classifier = KNeighborsClassifier(n_neighbors=n_neighbors, leaf_size=leaf_size, metric=metric)
classifier.fit(x_train, y_train)
y_pred = classifier.predict(x_test)
y_pred
from sklearn.metrics import accuracy_score
print(f'Accuracy score of {accuracy_score(y_test, y_pred)}')
#0.7877094972067039 lda
# 0.7988826815642458
# 0.8044692737430168 sd
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(y_test, y_pred)
print('The confusion matrix:')
confusion_matrix
```
### 70% training dataset - 30% testing dataset
```
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state=42)
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
lda = LDA(n_components=1)
x_train = lda.fit_transform(x_train, y_train)
x_test = lda.transform(x_test)
parameters = {'n_neighbors': list(range(1,30)),
'leaf_size': list(range(1,50)),
'metric':['euclidean', 'manhattan', 'chebyshev', 'minkowski', 'wminkowski', 'seuclidean', 'mahalanobis'],
}
knn = KNeighborsClassifier()
grid_search = GridSearchCV(estimator=knn,
param_grid=parameters)
print(grid_search.fit(x_train, y_train))
print('')
print('')
print('The parameters:')
print(grid_search.best_params_)
par = grid_search.best_params_
leaf_size = par['leaf_size']
n_neighbors = par['n_neighbors']
metric = par['metric']
classifier = KNeighborsClassifier(n_neighbors=n_neighbors, leaf_size=leaf_size)
classifier.fit(x_train, y_train)
y_pred = classifier.predict(x_test)
from sklearn.metrics import accuracy_score
print(f'Accuracy score of {accuracy_score(y_test, y_pred)}')
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(y_test, y_pred)
print('The confusion matrix:')
confusion_matrix
```
### 60% training dataset - 40% testing dataset
```
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size = 0.4, random_state=42)
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
lda = LDA(n_components=1)
x_train = lda.fit_transform(x_train, y_train)
x_test = lda.transform(x_test)
parameters = {'n_neighbors': list(range(1,30)),
'leaf_size': list(range(1,50)),
'metric':['euclidean', 'manhattan', 'chebyshev', 'minkowski', 'wminkowski', 'seuclidean', 'mahalanobis'],
}
knn = KNeighborsClassifier()
grid_search = GridSearchCV(estimator=knn,
param_grid=parameters)
print(grid_search.fit(x_train, y_train))
print('')
print('')
print('The parameters:')
print(grid_search.best_params_)
grid_search.best_params_
par = grid_search.best_params_
leaf_size = par['leaf_size']
n_neighbors = par['n_neighbors']
metric = par['metric']
classifier = KNeighborsClassifier(n_neighbors=n_neighbors, leaf_size=leaf_size)
classifier.fit(x_train, y_train)
y_pred = classifier.predict(x_test)
from sklearn.metrics import accuracy_score
print(f'Accuracy score of {accuracy_score(y_test, y_pred)}')
# 0.803921568627451
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(y_test, y_pred)
print('The confusion matrix:')
confusion_matrix
```
## Conlusion
> the 80 - 20 modal performed best with an accurancy score 81
| github_jupyter |
# Background Computation with "ofilter"
This notebook illustrate background calculations using `ofilter` algorithm adapted from IRAF.
```
import matplotlib.pyplot as plt
import numpy as np
import scipy
import ofiltsky
%matplotlib inline
```
### Generate Data
```
# Set the seed for reproducibility
np.random.seed(0)
# Random Poisson data
data = np.random.poisson(lam=1, size=(50, 50))
plt.imshow(data, cmap='viridis', interpolation='none')
plt.colorbar()
h = plt.hist(data.flatten(), bins=5, histtype='stepfilled')
```
### Quick IRAF Detour
To write this data out for IRAF, uncomment and run the following:
```
# from astropy.io import fits
#
# hdu0 = fits.PrimaryHDU()
# hdu1 = fits.ImageHDU(data.astype(np.float32))
# hdu = fits.HDUList([hdu0, hdu1])
# hdu.writeto('im_poisson_0.fits', clobber=True)
```
In addition to the FITS file, you also have to create an one-liner `im_poisson_0.coo` file with the following contents:
25 25
Inside your IRAF session, set the following parameters (more or less) using `epar`:
datapars.sigma = 1.
fitskypars.salgo = "ofilter"
fitskypars.annulus = 1.
fitskypars.dannulus = 20.
fitskypars.smaxiter = 10
fitskypars.snreject = 10
Then, run the following command:
fitsky im_poisson_0.fits im_poisson_0.coo
The result would be in `im_poisson_0.fits1.sky.1`:
MSKY = 0.7501966
STDEV = 0.7515768
SSKEW = 0.5060266
### Back to Python
For this dataset, Python version of the `ofilter` algorithm gives slightly lower sky and skew values, but comparable sigma. The Python version uses third-party libraries like Numpy, Scipy, and Astropy. Thus, it is not shocking that we are not getting complete agreement here.
Some questions that could be pursued:
1. Is it good enough? (Also see the next sub-section.)
2. Do we even care about the skew? Maybe not? In Python, it is calculated using `scipy.stats.skew()`.
```
np.ceil(data.std())
# NOTE: Sigma clipping does not matter much for this dataset.
ofil_results = ofiltsky.fitsky_ofilter(data, binsize=data.std(), sigclip_sigma=None)
print('MSKY =', ofil_results[0])
print('STDEV =', ofil_results[1])
print('SSKEW =', ofil_results[2])
```
We can also compare with some other available statistics:
```
sky_mean = data.mean()
sky_med = np.median(data)
sky_mode = scipy.stats.mode(data, axis=None).mode[0]
print('MEAN =', sky_mean)
print('MEDIAN =', sky_med)
print('MODE =', sky_mode)
```
### Comparing Results
This sub-section attempts to generate a plot not unlike what was published in WFPC2 ISR 1996-03 (Ferguson 1996). Perhaps the plot here can answer, "Is it good enough?"
```
# Populate desired background values
vals = np.arange(0, 4.5, 0.1)
# Initialize arrays to store results
sky_vals = []
sky_ofil = []
sky_med = []
sky_mean = []
# Generate results
for i, val in enumerate(vals):
np.random.seed(i)
data = np.random.poisson(lam=val, size=(50, 50))
try:
msky = ofiltsky.fitsky_ofilter(data, sigclip_sigma=None)[0]
except ValueError as e:
print('i={0}, val={1:.1f}, errmsg={2}'.format(i, val, str(e)))
continue
sky_vals.append(val)
sky_ofil.append(msky)
sky_med.append(np.median(data))
sky_mean.append(data.mean())
# Convert result to Numpy arrays
sky_vals = np.asarray(sky_vals)
sky_ofil = np.asarray(sky_ofil)
sky_med = np.asarray(sky_med)
sky_mean = np.asarray(sky_mean)
print()
print('Number of data points for plotting:', sky_ofil.size)
plt.scatter(sky_mean, sky_med - sky_mean, color='b', marker='x', label='median')
plt.scatter(sky_mean, sky_ofil - sky_mean, color='r', marker='o', label='ofilter (Python)')
plt.scatter([0.9868], [0.7501699 - 0.9868], color='k', marker='o', label='ofilter (IRAF)')
plt.xlabel('mean')
plt.ylabel('X - mean')
plt.axhline(0, color='k', linestyle='--')
plt.legend(bbox_to_anchor=(1.01, 1), loc='upper left', scatterpoints=1)
plt.scatter(sky_med, sky_mean - sky_med, color='b', marker='x', label='mean')
plt.scatter(sky_med, sky_ofil - sky_med, color='r', marker='o', label='ofilter (Python)')
plt.scatter([1], [0.7501699 - 1], color='k', marker='o', label='ofilter (IRAF)')
plt.xlabel('median')
plt.ylabel('X - median')
plt.axhline(0, color='k', linestyle='--')
plt.legend(bbox_to_anchor=(1.01, 1), loc='upper left', scatterpoints=1)
plt.scatter(sky_vals, sky_mean - sky_vals, color='b', marker='x', label='mean')
plt.scatter(sky_vals, sky_med - sky_vals, color='g', marker='*', label='median')
plt.scatter(sky_vals, sky_ofil - sky_vals, color='r', marker='o', label='ofilter (Python)')
plt.scatter([1], [0.7501699 - 1], color='k', marker='o', label='ofilter (IRAF)')
plt.xlabel('vals')
plt.ylabel('X - vals')
plt.axhline(0, color='k', linestyle='--')
plt.legend(bbox_to_anchor=(1.01, 1), loc='upper left', scatterpoints=1)
```
### Try with Other Skewed Distribution
```
from scipy.stats import gumbel_r
# Populate desired background values
vals = np.arange(0.1, 4.6, 0.1)
# Initialize arrays to store results
sky_vals = []
sky_ofil = []
sky_med = []
sky_mean = []
# Generate results
for i, val in enumerate(vals):
np.random.seed(i) # Does this control Scipy?
data = gumbel_r.rvs(loc=val, size=(50, 50))
try:
msky = ofiltsky.fitsky_ofilter(data, sigclip_sigma=None)[0]
except ValueError as e:
print('i={0}, val={1:.1f}, errmsg={2}'.format(i, val, str(e)))
continue
sky_vals.append(val)
sky_ofil.append(msky)
sky_med.append(np.median(data))
sky_mean.append(data.mean())
# Convert result to Numpy arrays
sky_vals = np.asarray(sky_vals)
sky_ofil = np.asarray(sky_ofil)
sky_med = np.asarray(sky_med)
sky_mean = np.asarray(sky_mean)
print()
print('Number of data points for plotting:', sky_ofil.size)
plt.scatter(sky_vals, sky_mean - sky_vals, color='b', marker='x', label='mean')
plt.scatter(sky_vals, sky_med - sky_vals, color='g', marker='*', label='median')
plt.scatter(sky_vals, sky_ofil - sky_vals, color='r', marker='o', label='ofilter (Python)')
plt.xlabel('vals')
plt.ylabel('X - vals')
plt.axhline(0, color='k', linestyle='--')
plt.legend(bbox_to_anchor=(1.01, 1), loc='upper left', scatterpoints=1)
```
Display histogram of the data from the last iteration above:
```
h = plt.hist(data.flatten(), bins=20, histtype='stepfilled')
plt.axvline(msky, color='r')
plt.axvline(np.median(data), color='g')
plt.axvline(data.mean(), color='b')
plt.axvline(val, color='k')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D2_DynamicNetworks/student/W3D2_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Neuromatch Academy: Week 3, Day 2, Tutorial 1
# Neuronal Network Dynamics: Neural Rate Models
__Content creators:__ Qinglong Gu, Songtin Li, Arvind Kumar, John Murray, Julijana Gjorgjieva
__Content reviewers:__ Maryam Vaziri-Pashkam, Ella Batty, Lorenzo Fontolan, Richard Gao, Spiros Chavlis, Michael Waskom
---
# Tutorial Objectives
The brain is a complex system, not because it is composed of a large number of diverse types of neurons, but mainly because of how neurons are connected to each other. The brain is indeed a network of highly specialized neuronal networks.
The activity of a neural network constantly evolves in time. For this reason, neurons can be modeled as dynamical systems. The dynamical system approach is only one of the many modeling approaches that computational neuroscientists have developed (other points of view include information processing, statistical models, etc.).
How the dynamics of neuronal networks affect the representation and processing of information in the brain is an open question. However, signatures of altered brain dynamics present in many brain diseases (e.g., in epilepsy or Parkinson's disease) tell us that it is crucial to study network activity dynamics if we want to understand the brain.
In this tutorial, we will simulate and study one of the simplest models of biological neuronal networks. Instead of modeling and simulating individual excitatory neurons (e.g., LIF models that you implemented yesterday), we will treat them as a single homogeneous population and approximate their dynamics using a single one-dimensional equation describing the evolution of their average spiking rate in time.
In this tutorial, we will learn how to build a firing rate model of a single population of excitatory neurons.
**Steps:**
- Write the equation for the firing rate dynamics of a 1D excitatory population.
- Visualize the response of the population as a function of parameters such as threshold level and gain, using the frequency-current (F-I) curve.
- Numerically simulate the dynamics of the excitatory population and find the fixed points of the system.
- Investigate the stability of the fixed points by linearizing the dynamics around them.
---
# Setup
```
# Imports
import matplotlib.pyplot as plt
import numpy as np
import scipy.optimize as opt # root-finding algorithm
# @title Figure Settings
import ipywidgets as widgets # interactive display
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# @title Helper functions
def plot_fI(x, f):
plt.figure(figsize=(6, 4)) # plot the figure
plt.plot(x, f, 'k')
plt.xlabel('x (a.u.)', fontsize=14)
plt.ylabel('F(x)', fontsize=14)
plt.show()
def plot_dr_r(r, drdt, x_fps=None):
plt.figure()
plt.plot(r, drdt, 'k')
plt.plot(r, 0. * r, 'k--')
if x_fps is not None:
plt.plot(x_fps, np.zeros_like(x_fps), "ko", ms=12)
plt.xlabel(r'$r$')
plt.ylabel(r'$\frac{dr}{dt}$', fontsize=20)
plt.ylim(-0.1, 0.1)
def plot_dFdt(x, dFdt):
plt.figure()
plt.plot(x, dFdt, 'r')
plt.xlabel('x (a.u.)', fontsize=14)
plt.ylabel('dF(x)', fontsize=14)
plt.show()
```
---
# Section 1: Neuronal network dynamics
```
# @title Video 1: Dynamic networks
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="p848349hPyw", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
## Section 1.1: Dynamics of a single excitatory population
Individual neurons respond by spiking. When we average the spikes of neurons in a population, we can define the average firing activity of the population. In this model, we are interested in how the population-averaged firing varies as a function of time and network parameters. Mathematically, we can describe the firing rate dynamic as:
\begin{align}
\tau \frac{dr}{dt} &= -r + F(w\cdot r + I_{\text{ext}}) \quad\qquad (1)
\end{align}
$r(t)$ represents the average firing rate of the excitatory population at time $t$, $\tau$ controls the timescale of the evolution of the average firing rate, $w$ denotes the strength (synaptic weight) of the recurrent input to the population, $I_{\text{ext}}$ represents the external input, and the transfer function $F(\cdot)$ (which can be related to f-I curve of individual neurons described in the next sections) represents the population activation function in response to all received inputs.
To start building the model, please execute the cell below to initialize the simulation parameters.
```
# @markdown *Execute this cell to set default parameters for a single excitatory population model*
def default_pars_single(**kwargs):
pars = {}
# Excitatory parameters
pars['tau'] = 1. # Timescale of the E population [ms]
pars['a'] = 1.2 # Gain of the E population
pars['theta'] = 2.8 # Threshold of the E population
# Connection strength
pars['w'] = 0. # E to E, we first set it to 0
# External input
pars['I_ext'] = 0.
# simulation parameters
pars['T'] = 20. # Total duration of simulation [ms]
pars['dt'] = .1 # Simulation time step [ms]
pars['r_init'] = 0.2 # Initial value of E
# External parameters if any
pars.update(kwargs)
# Vector of discretized time points [ms]
pars['range_t'] = np.arange(0, pars['T'], pars['dt'])
return pars
```
You can now use:
- `pars = default_pars_single()` to get all the parameters, and then you can execute `print(pars)` to check these parameters.
- `pars = default_pars_single(T=T_sim, dt=time_step)` to set new simulation time and time step
- To update an existing parameter dictionary, use `pars['New_para'] = value`
Because `pars` is a dictionary, it can be passed to a function that requires individual parameters as arguments using `my_func(**pars)` syntax.
## Section 1.2: F-I curves
In electrophysiology, a neuron is often characterized by its spike rate output in response to input currents. This is often called the **F-I** curve, denoting the output spike frequency (**F**) in response to different injected currents (**I**). We estimated this for an LIF neuron in yesterday's tutorial.
The transfer function $F(\cdot)$ in Equation $1$ represents the gain of the population as a function of the total input. The gain is often modeled as a sigmoidal function, i.e., more input drive leads to a nonlinear increase in the population firing rate. The output firing rate will eventually saturate for high input values.
A sigmoidal $F(\cdot)$ is parameterized by its gain $a$ and threshold $\theta$.
$$ F(x;a,\theta) = \frac{1}{1+\text{e}^{-a(x-\theta)}} - \frac{1}{1+\text{e}^{a\theta}} \quad(2)$$
The argument $x$ represents the input to the population. Note that the second term is chosen so that $F(0;a,\theta)=0$.
Many other transfer functions (generally monotonic) can be also used. Examples are the rectified linear function $ReLU(x)$ or the hyperbolic tangent $tanh(x)$.
### Exercise 1: Implement F-I curve
Let's first investigate the activation functions before simulating the dynamics of the entire population.
In this exercise, you will implement a sigmoidal **F-I** curve or transfer function $F(x)$, with gain $a$ and threshold level $\theta$ as parameters.
```
def F(x, a, theta):
"""
Population activation function.
Args:
x (float): the population input
a (float): the gain of the function
theta (float): the threshold of the function
Returns:
float: the population activation response F(x) for input x
"""
#################################################
## TODO for students: compute f = F(x) ##
# Fill out function and remove
raise NotImplementedError("Student excercise: implement the f-I function")
#################################################
# Define the sigmoidal transfer function f = F(x)
f = ...
return f
pars = default_pars_single() # get default parameters
x = np.arange(0, 10, .1) # set the range of input
# Uncomment below to test your function
# f = F(x, pars['a'], pars['theta'])
# plot_fI(x, f)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_45ddc05f.py)
*Example output:*
<img alt='Solution hint' align='left' width=416 height=272 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D2_DynamicNetworks/static/W3D2_Tutorial1_Solution_45ddc05f_0.png>
### Interactive Demo: Parameter exploration of F-I curve
Here's an interactive demo that shows how the F-I curve changes for different values of the gain and threshold parameters. How do the gain and threshold parameters affect the F-I curve?
```
# @title
# @markdown Make sure you execute this cell to enable the widget!
def interactive_plot_FI(a, theta):
"""
Population activation function.
Expecxts:
a : the gain of the function
theta : the threshold of the function
Returns:
plot the F-I curve with give parameters
"""
# set the range of input
x = np.arange(0, 10, .1)
plt.figure()
plt.plot(x, F(x, a, theta), 'k')
plt.xlabel('x (a.u.)', fontsize=14)
plt.ylabel('F(x)', fontsize=14)
plt.show()
_ = widgets.interact(interactive_plot_FI, a=(0.3, 3, 0.3), theta=(2, 4, 0.2))
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_1c0165d7.py)
## Section 1.3: Simulation scheme of E dynamics
Because $F(\cdot)$ is a nonlinear function, the exact solution of Equation $1$ can not be determined via analytical methods. Therefore, numerical methods must be used to find the solution. In practice, the derivative on the left-hand side of Equation $1$ can be approximated using the Euler method on a time-grid of stepsize $\Delta t$:
\begin{align}
&\frac{dr}{dt} \approx \frac{r[k+1]-r[k]}{\Delta t}
\end{align}
where $r[k] = r(k\Delta t)$.
Thus,
$$\Delta r[k] = \frac{\Delta t}{\tau}[-r[k] + F(w\cdot r[k] + I_{\text{ext}}(k;a,\theta))]$$
Hence, Equation (1) is updated at each time step by:
$$r[k+1] = r[k] + \Delta r[k]$$
```
# @markdown *Execute this cell to enable the single population rate model simulator: `simulate_single`*
def simulate_single(pars):
"""
Simulate an excitatory population of neurons
Args:
pars : Parameter dictionary
Returns:
rE : Activity of excitatory population (array)
Example:
pars = default_pars_single()
r = simulate_single(pars)
"""
# Set parameters
tau, a, theta = pars['tau'], pars['a'], pars['theta']
w = pars['w']
I_ext = pars['I_ext']
r_init = pars['r_init']
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# Initialize activity
r = np.zeros(Lt)
r[0] = r_init
I_ext = I_ext * np.ones(Lt)
# Update the E activity
for k in range(Lt - 1):
dr = dt / tau * (-r[k] + F(w * r[k] + I_ext[k], a, theta))
r[k+1] = r[k] + dr
return r
help(simulate_single)
```
### Interactive Demo: Parameter Exploration of single population dynamics
Note that $w=0$, as in the default setting, means no recurrent input to the neuron population in Equation (1). Hence, the dynamics are entirely determined by the external input $I_{\text{ext}}$. Explore these dynamics in this interactive demo.
How does $r_{\text{sim}}(t)$ change with different $I_{\text{ext}}$ values? How does it change with different $\tau$ values? Investigate the relationship between $F(I_{\text{ext}}; a, \theta)$ and the steady value of $r(t)$.
Note that, $r_{\rm ana}(t)$ denotes the analytical solution - you will learn how this is computed in the next section.
```
# @title
# @markdown Make sure you execute this cell to enable the widget!
# get default parameters
pars = default_pars_single(T=20.)
def Myplot_E_diffI_difftau(I_ext, tau):
# set external input and time constant
pars['I_ext'] = I_ext
pars['tau'] = tau
# simulation
r = simulate_single(pars)
# Analytical Solution
r_ana = (pars['r_init']
+ (F(I_ext, pars['a'], pars['theta'])
- pars['r_init']) * (1. - np.exp(-pars['range_t'] / pars['tau'])))
# plot
plt.figure()
plt.plot(pars['range_t'], r, 'b', label=r'$r_{\mathrm{sim}}$(t)', alpha=0.5,
zorder=1)
plt.plot(pars['range_t'], r_ana, 'b--', lw=5, dashes=(2, 2),
label=r'$r_{\mathrm{ana}}$(t)', zorder=2)
plt.plot(pars['range_t'],
F(I_ext, pars['a'], pars['theta']) * np.ones(pars['range_t'].size),
'k--', label=r'$F(I_{\mathrm{ext}})$')
plt.xlabel('t (ms)', fontsize=16.)
plt.ylabel('Activity r(t)', fontsize=16.)
plt.legend(loc='best', fontsize=14.)
plt.show()
_ = widgets.interact(Myplot_E_diffI_difftau, I_ext=(0.0, 10., 1.),
tau=(1., 5., 0.2))
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_65dee3e7.py)
## Think!
Above, we have numerically solved a system driven by a positive input and that, if $w_{EE} \neq 0$, receives an excitatory recurrent input (**extra challenge: try changing the value of $w_{EE}$ to a positive number and plotting the results of simulate_single**). Yet, $r_E(t)$ either decays to zero or reaches a fixed non-zero value.
- Why doesn't the solution of the system "explode" in a finite time? In other words, what guarantees that $r_E$(t) stays finite?
- Which parameter would you change in order to increase the maximum value of the response?
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_5a95a98e.py)
---
# Section 2: Fixed points of the single population system
```
# @title Video 2: Fixed point
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="Ox3ELd1UFyo", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
As you varied the two parameters in the last Interactive Demo, you noticed that, while at first the system output quickly changes, with time, it reaches its maximum/minimum value and does not change anymore. The value eventually reached by the system is called the **steady state** of the system, or the **fixed point**. Essentially, in the steady states the derivative with respect to time of the activity ($r$) is zero, i.e. $\displaystyle \frac{dr}{dt}=0$.
We can find that the steady state of the Equation. (1) by setting $\displaystyle{\frac{dr}{dt}=0}$ and solve for $r$:
$$-r_{\text{steady}} + F(w\cdot r_{\text{steady}} + I_{\text{ext}};a,\theta) = 0, \qquad (3)$$
When it exists, the solution of Equation. (3) defines a **fixed point** of the dynamical system in Equation (1). Note that if $F(x)$ is nonlinear, it is not always possible to find an analytical solution, but the solution can be found via numerical simulations, as we will do later.
From the Interactive Demo, one could also notice that the value of $\tau$ influences how quickly the activity will converge to the steady state from its initial value.
In the specific case of $w=0$, we can also analytically compute the solution of Equation (1) (i.e., the thick blue dashed line) and deduce the role of $\tau$ in determining the convergence to the fixed point:
$$\displaystyle{r(t) = \big{[}F(I_{\text{ext}};a,\theta) -r(t=0)\big{]} (1-\text{e}^{-\frac{t}{\tau}})} + r(t=0)$$ \\
We can now numerically calculate the fixed point with a root finding algorithm.
## Exercise 2: Visualization of the fixed points
When it is not possible to find the solution for Equation (3) analytically, a graphical approach can be taken. To that end, it is useful to plot $\displaystyle{\frac{dr}{dt}}$ as a function of $r$. The values of $r$ for which the plotted function crosses zero on the y axis correspond to fixed points.
Here, let us, for example, set $w=5.0$ and $I^{\text{ext}}=0.5$. From Equation (1), you can obtain
$$\frac{dr}{dt} = [-r + F(w\cdot r + I^{\text{ext}})]\,/\,\tau $$
Then, plot the $dr/dt$ as a function of $r$, and check for the presence of fixed points.
```
def compute_drdt(r, I_ext, w, a, theta, tau, **other_pars):
"""Given parameters, compute dr/dt as a function of r.
Args:
r (1D array) : Average firing rate of the excitatory population
I_ext, w, a, theta, tau (numbers): Simulation parameters to use
other_pars : Other simulation parameters are unused by this function
Returns
drdt function for each value of r
"""
#########################################################################
# TODO compute drdt and disable the error
raise NotImplementedError("Finish the compute_drdt function")
#########################################################################
# Calculate drdt
drdt = ...
return drdt
# Define a vector of r values and the simulation parameters
r = np.linspace(0, 1, 1000)
pars = default_pars_single(I_ext=0.5, w=5)
# Uncomment to test your function
# drdt = compute_drdt(r, **pars)
# plot_dr_r(r, drdt)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_c5280901.py)
*Example output:*
<img alt='Solution hint' align='left' width=558 height=413 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D2_DynamicNetworks/static/W3D2_Tutorial1_Solution_c5280901_0.png>
## Exercise 3: Fixed point calculation
We will now find the fixed points numerically. To do so, we need to specif initial values ($r_{\text{guess}}$) for the root-finding algorithm to start from. From the line $\displaystyle{\frac{dr}{dt}}$ plotted above in Exercise 2, initial values can be chosen as a set of values close to where the line crosses zero on the y axis (real fixed point).
The next cell defines three helper functions that we will use:
- `my_fp_single(r_guess, **pars)` uses a root-finding algorithm to locate a fixed point near a given initial value
- `check_fp_single(x_fp, **pars)`, verifies that the values of $r_{\rm fp}$ for which $\displaystyle{\frac{dr}{dt}} = 0$ are the true fixed points
- `my_fp_finder(r_guess_vector, **pars)` accepts an array of initial values and finds the same number of fixed points, using the above two functions
```
# @markdown *Execute this cell to enable the fixed point functions*
def my_fp_single(r_guess, a, theta, w, I_ext, **other_pars):
"""
Calculate the fixed point through drE/dt=0
Args:
r_guess : Initial value used for scipy.optimize function
a, theta, w, I_ext : simulation parameters
Returns:
x_fp : value of fixed point
"""
# define the right hand of E dynamics
def my_WCr(x):
r = x
drdt = (-r + F(w * r + I_ext, a, theta))
y = np.array(drdt)
return y
x0 = np.array(r_guess)
x_fp = opt.root(my_WCr, x0).x.item()
return x_fp
def check_fp_single(x_fp, a, theta, w, I_ext, mytol=1e-4, **other_pars):
"""
Verify |dr/dt| < mytol
Args:
fp : value of fixed point
a, theta, w, I_ext: simulation parameters
mytol : tolerance, default as 10^{-4}
Returns :
Whether it is a correct fixed point: True/False
"""
# calculate Equation(3)
y = x_fp - F(w * x_fp + I_ext, a, theta)
# Here we set tolerance as 10^{-4}
return np.abs(y) < mytol
def my_fp_finder(pars, r_guess_vector, mytol=1e-4):
"""
Calculate the fixed point(s) through drE/dt=0
Args:
pars : Parameter dictionary
r_guess_vector : Initial values used for scipy.optimize function
mytol : tolerance for checking fixed point, default as 10^{-4}
Returns:
x_fps : values of fixed points
"""
x_fps = []
correct_fps = []
for r_guess in r_guess_vector:
x_fp = my_fp_single(r_guess, **pars)
if check_fp_single(x_fp, **pars, mytol=mytol):
x_fps.append(x_fp)
return x_fps
help(my_fp_finder)
r = np.linspace(0, 1, 1000)
pars = default_pars_single(I_ext=0.5, w=5)
drdt = compute_drdt(r, **pars)
#############################################################################
# TODO for students:
# Define initial values close to the intersections of drdt and y=0
# (How many initial values? Hint: How many times do the two lines intersect?)
# Calculate the fixed point with these initial values and plot them
#############################################################################
r_guess_vector = [...]
# Uncomment to test your values
# x_fps = my_fp_finder(pars, r_guess_vector)
# plot_dr_r(r, drdt, x_fps)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_0637b6bf.py)
*Example output:*
<img alt='Solution hint' align='left' width=558 height=413 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D2_DynamicNetworks/static/W3D2_Tutorial1_Solution_0637b6bf_0.png>
## Interactive Demo: fixed points as a function of recurrent and external inputs.
You can now explore how the previous plot changes when the recurrent coupling $w$ and the external input $I_{\text{ext}}$ take different values. How does the number of fixed points change?
```
# @title
# @markdown Make sure you execute this cell to enable the widget!
def plot_intersection_single(w, I_ext):
# set your parameters
pars = default_pars_single(w=w, I_ext=I_ext)
# find fixed points
r_init_vector = [0, .4, .9]
x_fps = my_fp_finder(pars, r_init_vector)
# plot
r = np.linspace(0, 1., 1000)
drdt = (-r + F(w * r + I_ext, pars['a'], pars['theta'])) / pars['tau']
plot_dr_r(r, drdt, x_fps)
_ = widgets.interact(plot_intersection_single, w=(1, 7, 0.2),
I_ext=(0, 3, 0.1))
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_20486792.py)
---
# Summary
In this tutorial, we have investigated the dynamics of a rate-based single population of neurons.
We learned about:
- The effect of the input parameters and the time constant of the network on the dynamics of the population.
- How to find the fixed point(s) of the system.
Next, we have two Bonus, but important concepts in dynamical system analysis and simulation. If you have time left, watch the next video and proceed to solve the exercises. You will learn:
- How to determine the stability of a fixed point by linearizing the system.
- How to add realistic inputs to our model.
---
# Bonus 1: Stability of a fixed point
```
# @title Video 3: Stability of fixed points
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="KKMlWWU83Jg", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
#### Initial values and trajectories
Here, let us first set $w=5.0$ and $I_{\text{ext}}=0.5$, and investigate the dynamics of $r(t)$ starting with different initial values $r(0) \equiv r_{\text{init}}$. We will plot the trajectories of $r(t)$ with $r_{\text{init}} = 0.0, 0.1, 0.2,..., 0.9$.
```
# @markdown Execute this cell to see the trajectories!
pars = default_pars_single()
pars['w'] = 5.0
pars['I_ext'] = 0.5
plt.figure(figsize=(8, 5))
for ie in range(10):
pars['r_init'] = 0.1 * ie # set the initial value
r = simulate_single(pars) # run the simulation
# plot the activity with given initial
plt.plot(pars['range_t'], r, 'b', alpha=0.1 + 0.1 * ie,
label=r'r$_{\mathrm{init}}$=%.1f' % (0.1 * ie))
plt.xlabel('t (ms)')
plt.title('Two steady states?')
plt.ylabel(r'$r$(t)')
plt.legend(loc=[1.01, -0.06], fontsize=14)
plt.show()
```
## Interactive Demo: dynamics as a function of the initial value
Let's now set $r_{\rm init}$ to a value of your choice in this demo. How does the solution change? What do you observe?
```
# @title
# @markdown Make sure you execute this cell to enable the widget!
pars = default_pars_single(w=5.0, I_ext=0.5)
def plot_single_diffEinit(r_init):
pars['r_init'] = r_init
r = simulate_single(pars)
plt.figure()
plt.plot(pars['range_t'], r, 'b', zorder=1)
plt.plot(0, r[0], 'bo', alpha=0.7, zorder=2)
plt.xlabel('t (ms)', fontsize=16)
plt.ylabel(r'$r(t)$', fontsize=16)
plt.ylim(0, 1.0)
plt.show()
_ = widgets.interact(plot_single_diffEinit, r_init=(0, 1, 0.02))
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_4d2de6a0.py)
### Stability analysis via linearization of the dynamics
Just like Equation $1$ in the case ($w=0$) discussed above, a generic linear system
$$\frac{dx}{dt} = \lambda (x - b),$$
has a fixed point for $x=b$. The analytical solution of such a system can be found to be:
$$x(t) = b + \big{(} x(0) - b \big{)} \text{e}^{\lambda t}.$$
Now consider a small perturbation of the activity around the fixed point: $x(0) = b+ \epsilon$, where $|\epsilon| \ll 1$. Will the perturbation $\epsilon(t)$ grow with time or will it decay to the fixed point? The evolution of the perturbation with time can be written, using the analytical solution for $x(t)$, as:
$$\epsilon (t) = x(t) - b = \epsilon \text{e}^{\lambda t}$$
- if $\lambda < 0$, $\epsilon(t)$ decays to zero, $x(t)$ will still converge to $b$ and the fixed point is "**stable**".
- if $\lambda > 0$, $\epsilon(t)$ grows with time, $x(t)$ will leave the fixed point $b$ exponentially, and the fixed point is, therefore, "**unstable**" .
### Compute the stability of Equation $1$
Similar to what we did in the linear system above, in order to determine the stability of a fixed point $r^{*}$ of the excitatory population dynamics, we perturb Equation (1) around $r^{*}$ by $\epsilon$, i.e. $r = r^{*} + \epsilon$. We can plug in Equation (1) and obtain the equation determining the time evolution of the perturbation $\epsilon(t)$:
\begin{align}
\tau \frac{d\epsilon}{dt} \approx -\epsilon + w F'(w\cdot r^{*} + I_{\text{ext}};a,\theta) \epsilon
\end{align}
where $F'(\cdot)$ is the derivative of the transfer function $F(\cdot)$. We can rewrite the above equation as:
\begin{align}
\frac{d\epsilon}{dt} \approx \frac{\epsilon}{\tau }[-1 + w F'(w\cdot r^* + I_{\text{ext}};a,\theta)]
\end{align}
That is, as in the linear system above, the value of
$$\lambda = [-1+ wF'(w\cdot r^* + I_{\text{ext}};a,\theta)]/\tau \qquad (4)$$
determines whether the perturbation will grow or decay to zero, i.e., $\lambda$ defines the stability of the fixed point. This value is called the **eigenvalue** of the dynamical system.
## Exercise 4: Compute $dF$
The derivative of the sigmoid transfer function is:
\begin{align}
\frac{dF}{dx} & = \frac{d}{dx} (1+\exp\{-a(x-\theta)\})^{-1} \\
& = a\exp\{-a(x-\theta)\} (1+\exp\{-a(x-\theta)\})^{-2}. \qquad (5)
\end{align}
Let's now find the expression for the derivative $\displaystyle{\frac{dF}{dx}}$ in the following cell and plot it.
```
def dF(x, a, theta):
"""
Population activation function.
Args:
x : the population input
a : the gain of the function
theta : the threshold of the function
Returns:
dFdx : the population activation response F(x) for input x
"""
###########################################################################
# TODO for students: compute dFdx ##
raise NotImplementedError("Student excercise: compute the deravitive of F")
###########################################################################
# Calculate the population activation
dFdx = ...
return dFdx
pars = default_pars_single() # get default parameters
x = np.arange(0, 10, .1) # set the range of input
# Uncomment below to test your function
# df = dF(x, pars['a'], pars['theta'])
# plot_dFdt(x, df)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_ce2e3bc5.py)
*Example output:*
<img alt='Solution hint' align='left' width=560 height=416 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D2_DynamicNetworks/static/W3D2_Tutorial1_Solution_ce2e3bc5_0.png>
## Exercise 5: Compute eigenvalues
As discussed above, for the case with $w=5.0$ and $I_{\text{ext}}=0.5$, the system displays **three** fixed points. However, when we simulated the dynamics and varied the initial conditions $r_{\rm init}$, we could only obtain **two** steady states. In this exercise, we will now check the stability of each of the three fixed points by calculating the corresponding eigenvalues with the function `eig_single`. Check the sign of each eigenvalue (i.e., stability of each fixed point). How many of the fixed points are stable?
Note that the expression of the eigenvalue at fixed point $r^*$
$$\lambda = [-1+ wF'(w\cdot r^* + I_{\text{ext}};a,\theta)]/\tau$$
```
def eig_single(fp, tau, a, theta, w, I_ext, **other_pars):
"""
Args:
fp : fixed point r_fp
tau, a, theta, w, I_ext : Simulation parameters
Returns:
eig : eigevalue of the linearized system
"""
#####################################################################
## TODO for students: compute eigenvalue and disable the error
raise NotImplementedError("Student excercise: compute the eigenvalue")
######################################################################
# Compute the eigenvalue
eig = ...
return eig
# Find the eigenvalues for all fixed points of Exercise 2
pars = default_pars_single(w=5, I_ext=.5)
r_guess_vector = [0, .4, .9]
x_fp = my_fp_finder(pars, r_guess_vector)
# Uncomment below lines after completing the eig_single function.
# for fp in x_fp:
# eig_fp = eig_single(fp, **pars)
# print(f'Fixed point1 at {fp:.3f} with Eigenvalue={eig_fp:.3f}')
```
**SAMPLE OUTPUT**
```
Fixed point1 at 0.042 with Eigenvalue=-0.583
Fixed point2 at 0.447 with Eigenvalue=0.498
Fixed point3 at 0.900 with Eigenvalue=-0.626
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_e285f60d.py)
## Think!
Throughout the tutorial, we have assumed $w> 0 $, i.e., we considered a single population of **excitatory** neurons. What do you think will be the behavior of a population of inhibitory neurons, i.e., where $w> 0$ is replaced by $w< 0$?
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial1_Solution_579bc9c9.py)
---
# Bonus 2: Noisy input drives the transition between two stable states
## Ornstein-Uhlenbeck (OU) process
As discussed in several previous tutorials, the OU process is usually used to generate a noisy input into the neuron. The OU input $\eta(t)$ follows:
$$\tau_\eta \frac{d}{dt}\eta(t) = -\eta (t) + \sigma_\eta\sqrt{2\tau_\eta}\xi(t)$$
Execute the following function `my_OU(pars, sig, myseed=False)` to generate an OU process.
```
# @title OU process `my_OU(pars, sig, myseed=False)`
# @markdown Make sure you execute this cell to visualize the noise!
def my_OU(pars, sig, myseed=False):
"""
A functions that generates Ornstein-Uhlenback process
Args:
pars : parameter dictionary
sig : noise amplitute
myseed : random seed. int or boolean
Returns:
I : Ornstein-Uhlenbeck input current
"""
# Retrieve simulation parameters
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
tau_ou = pars['tau_ou'] # [ms]
# set random seed
if myseed:
np.random.seed(seed=myseed)
else:
np.random.seed()
# Initialize
noise = np.random.randn(Lt)
I_ou = np.zeros(Lt)
I_ou[0] = noise[0] * sig
# generate OU
for it in range(Lt - 1):
I_ou[it + 1] = (I_ou[it]
+ dt / tau_ou * (0. - I_ou[it])
+ np.sqrt(2 * dt / tau_ou) * sig * noise[it + 1])
return I_ou
pars = default_pars_single(T=100)
pars['tau_ou'] = 1. # [ms]
sig_ou = 0.1
I_ou = my_OU(pars, sig=sig_ou, myseed=2020)
plt.figure(figsize=(10, 4))
plt.plot(pars['range_t'], I_ou, 'r')
plt.xlabel('t (ms)')
plt.ylabel(r'$I_{\mathrm{OU}}$')
plt.show()
```
## Example: Up-Down transition
In the presence of two or more fixed points, noisy inputs can drive a transition between the fixed points! Here, we stimulate an E population for 1,000 ms applying OU inputs.
```
# @title Simulation of an E population with OU inputs
# @markdown Make sure you execute this cell to spot the Up-Down states!
pars = default_pars_single(T=1000)
pars['w'] = 5.0
sig_ou = 0.7
pars['tau_ou'] = 1. # [ms]
pars['I_ext'] = 0.56 + my_OU(pars, sig=sig_ou, myseed=2020)
r = simulate_single(pars)
plt.figure(figsize=(10, 4))
plt.plot(pars['range_t'], r, 'b', alpha=0.8)
plt.xlabel('t (ms)')
plt.ylabel(r'$r(t)$')
plt.show()
```
| github_jupyter |
# Machine Learning with Python
## Setup
```
# Package imports
import matplotlib.pyplot as plt # data visualisation
import numpy as np # mathematics
import pandas as pd # data manipulation
from sklearn.linear_model import LinearRegression # regression
# Graphical properties
plt.rcParams['figure.facecolor'] = 'white'
# Parameters
BASE_URL = 'https://raw.githubusercontent.com/warwickdatascience/' + \
'helping-hack-heart-of-england/main/resources/'
```
## Exploratory Analysis
Data available from the [hackathon website](https://warwickdatascience.github.io/helping-hack-heart-of-england/).
```
# Data import
imd = pd.read_csv(BASE_URL + 'imd.csv')
ref = pd.read_csv(BASE_URL + 'ref.csv')
# Sample datsets
display(imd.sample(10))
display(ref.sample(10))
# Change in IMD
imd_train = imd.loc[imd['imd_2019'].notnull()].copy()
imd_train['imd_change'] = imd_train['imd_2019'] - imd_train['imd_2015']
fig, ax = plt.subplots()
ax.hist(imd_train['imd_change'], bins=25,
edgecolor='black', color='lightblue')
ax.set_xlabel("Change in IMD")
ax.set_ylabel("Count")
plt.show()
# Data aggregation
group_cols = ['lad_code', 'lad_name', 'class', 'category']
ref_agg = ref.groupby(group_cols)['expenditure'].mean().reset_index()
display(ref_agg.sample(10))
```
A massive flaw is already obvious: by not including county/London boroughs/other authorities, we are not accurately reflecting spending. We will not address this issue here as this is one way you can approve your leaderboard score.
```
# Expenditure by category
categories = ref_agg['category'].unique()
fig, axs = plt.subplots((len(categories) + 1) // 2, 2,
figsize=(8, 16))
for i, cat in enumerate(categories):
ax = axs[i // 2][i % 2]
ref_sub = ref_agg.loc[(ref_agg['category'] == cat) &
(ref_agg['expenditure'] > 0)]
ax.hist(np.log10(ref_sub['expenditure']), bins=25,
edgecolor='black', color='lightblue')
ax.set_title(cat)
fig.text(0.5, 0.1, 'Expenditure', ha='center', va='center')
fig.text(0.06, 0.5, 'Count', ha='center', va='center', rotation='vertical')
plt.subplots_adjust(hspace=0.4)
plt.show()
# Expenditure by category and class
ref_agg['expenditure_sign'] = 'Negative'
ref_agg.loc[ref_agg['expenditure'] > 0, 'expenditure_sign'] = 'Positive'
ref_agg.loc[np.isclose(ref_agg['expenditure'], 0), 'expenditure_sign'] = 'Zero'
ref_agg['class_type'] = ref_agg['class'].map({
'L': 'Individual', 'UA': 'Individual', 'SD': 'Individual',
'MD': 'Combined', 'SC': 'Combined',
'O': 'Other'
})
fig, axs = plt.subplots(1, 3, figsize=(12, 6))
for i, ct in enumerate(('Combined', 'Individual', 'Other')):
ax = axs[i]
ref_sub = ref_agg.loc[ref_agg['class_type'] == ct]
prev_height = 0
for es, c in zip(('Negative', 'Positive', 'Zero'), ('r', 'g', 'b')):
ref_sub2 = ref_sub.loc[ref_sub['expenditure_sign'] == es]
curr_height = ref_sub2.groupby('category')['expenditure'].count()
curr_height = curr_height.reindex(categories).fillna(0)
ax.barh(categories, curr_height, left=prev_height, color=c,
label=es if i == 0 else None)
ax.set_title(ct)
prev_height = curr_height + prev_height
if i: ax.set_yticklabels([])
ax.set_xticklabels([])
fig.text(0.5, 0.07, 'Proportion', ha='center', va='center')
fig.text(-0.13, 0.5, 'Expenditure Category',
ha='center', va='center', rotation='vertical')
fig.legend(loc='upper left', bbox_to_anchor=(0.05, 0, 1, 0.87))
plt.subplots_adjust(hspace=0.4)
plt.show()
# Correlations
combi_sub = pd.merge(
imd_train[['lad_code', 'imd_change']],
ref_agg[['lad_code', 'category', 'expenditure']].loc[
ref_agg['expenditure'] > 0
], how='inner'
)
fig, axs = plt.subplots((len(categories) + 1) // 2, 2,
figsize=(8, 16))
for i, cat in enumerate(categories):
ax = axs[i // 2][i % 2]
combi_sub2 = combi_sub.loc[(combi_sub['category'] == cat)]
exp = np.log10(combi_sub2['expenditure'])
imd_chng = combi_sub2['imd_change']
ax.plot(exp, imd_chng, 'ko', exp,
np.poly1d(np.polyfit(exp, imd_chng, 1))(exp), 'b')
ax.set_title(cat)
fig.text(0.5, 0.1, 'Expenditure', ha='center', va='center')
fig.text(0.06, 0.5, 'Count', ha='center', va='center', rotation='vertical')
plt.subplots_adjust(hspace=0.4)
plt.show()
```
## Modelling
```
# Transform data
ref_agg['clean_cat'] = ref_agg['category'].str.replace(' ', '_')
ref_agg['clean_cat'] = ref_agg['clean_cat'].str.replace(r'[^\w]+', '')
ref_wide = ref_agg.pivot_table(index=['lad_code', 'lad_name', 'class'],
columns='clean_cat',
values='expenditure').reset_index()
ref_wide.columns.name = None
display(ref_wide.sample(10))
# Combine datasets
combi_train = pd.merge(
imd_train[['lad_code', 'imd_change']],
ref_wide.drop(['lad_name', 'class'], axis=1),
how='left'
)
# Build model
reg = LinearRegression()
reg.fit(combi_train.drop(['lad_code', 'imd_change'], axis=1),
combi_train['imd_change']); # baseline model on leaderboard
```
_For null model checks and ANOVA, see the R version of this notebook._
## Output
```
# Make predictions
imd_test = imd.loc[imd['imd_2019'].isnull()].copy()[['lad_code', 'imd_2015']]
combi_test = pd.merge(
imd_test, ref_wide.drop(['lad_name', 'class'], axis=1),
how='left'
)
pred = reg.predict(combi_test.drop(['lad_code', 'imd_2015'], axis=1))
# Output predictions
imd_test['imd_2019'] = imd_test['imd_2015'] + pred
imd_test.to_csv('tutorial.csv')
```
## Closing Notes
### Comments
Possible improvements:
- Use lookup tables to bring in all data sources
- Consider using multi-level modeling techniques
- Implement cross-validation to ensure generalisation
- Use regularisation to reduce generalisation gap
- Consider more complex models
- Model using a transformed expenditure
- Use time series modelling
### Learning Resources
- [Python Data Science Handbook](https://jakevdp.github.io/PythonDataScienceHandbook/)
- [Python Cookbook](https://d.cxcore.net/Python/Python_Cookbook_3rd_Edition.pdf)
- [DataCamp Trial](https://www.datacamp.com/github-students)
| github_jupyter |
# Creating city models and objects
In this tutorial we explore how to create new city models with using `cjio`'s
API.
```
from pathlib import Path
from cjio import cityjson
from cjio.models import CityObject, Geometry
```
Set up paths for the tutorial.
```
package_dir = Path(__name__).resolve().parent.parent.parent
schema_dir = package_dir / 'cjio' / 'schemas'/ '1.0.0'
data_dir = package_dir / 'tests' / 'data'
```
## Creating a single CityObject
We are building a single CityObject of type *Building*. This building has an
LoD2 geometry, thus it has Semantic Surfaces. The geometric shape of the
building is a simple cube (size 10x10x10), which is sufficient for this
demonstration.
The idea is that we create empty containers for the CityModel, CityObjects and
Geometries, then fill those up and add to the CityModel.
We create an empty CityModel
```
cm = cityjson.CityJSON()
print(cm)
```
An empty CityObject. Note that the ID is required.
```
co = CityObject(
id='1'
)
```
We can also add attributes
```
co_attrs = {
'some_attribute': 42,
'other_attribute': 'bla bla'
}
co.attributes = co_attrs
```
Let's see what do we have
```
print(co)
```
Instantiate a Geometry without boundaries and semantics
```
geom = Geometry(type='Solid', lod=2)
```
We build the boundary Solid of the cube
The surfaces are in this order: WallSurface, WallSurface, WallSurface, WallSurface, GroundSurface, RoofSurface
```
bdry = [
[[(0.0, 0.0, 0.0), (10.0, 0.0, 0.0), (10.0, 0.0, 10.0), (0.0, 0.0, 10.0)]],
[[(10.0, 0.0, 0.0), (10.0, 10.0, 0.0), (10.0, 10.0, 10.0), (10.0, 0.0, 10.0)]],
[[(10.0, 10.0, 0.0), (0.0, 10.0, 0.0), (0.0, 10.0, 10.0), (10.0, 10.0, 10.0)]],
[[(0.0, 10.0, 0.0), (0.0, 0.0, 0.0), (0.0, 0.0, 10.0), (0.0, 10.0, 10.0)]],
[[(0.0, 0.0, 0.0), (0.0, 10.0, 0.0), (10.0, 10.0, 0.0), (10.0, 0.0, 0.0)]],
[[(10.0, 0.0, 10.0), (10.0, 10.0, 10.0), (0.0, 10.0, 10.0), (0.0, 0.0, 10.0)]]
]
```
Add the boundary to the Geometry
```
geom.boundaries.append(bdry)
```
We build the SemanticSurfaces for the boundary. The `surfaces` attribute must
contain at least the `surface_idx` and `type` keys, optionally `attributes`.
We have three semantic surface types, WallSurface, GroundSurface, RoofSurface.
```
srf = {
0: {'surface_idx': [], 'type': 'WallSurface'},
1: {'surface_idx': [], 'type': 'GroundSurface'},
2: {'surface_idx': [], 'type': 'RoofSurface'}
}
```
We use the `surface_idx` to point to the surfaces of the boundary. Thus the
index to a single boundary surface is composed as [Solid index, Shell index, Surface index].
Consequently, in case of a CompositeSolid which first Solid, outer Shell,
second Surface is a WallSurface, one element in the `surface_idx` would be
`[0, 0, 1]`. Then assuming that there is only a single WallSurface in the
mentioned CompositeSolid, the index to the WallSurfaces is composed as
`{'surface_idx': [ [0, 0, 1] ], 'type': 'WallSurface'}`.
In case of a Solid boundary type the *Solid index* is omitted from the elements
of `surface_idx`. In case of a MultiSurface boundary type both the *Solid index*
and *Shell index* are omitted from the elements of `surface_idx`.
We create the surface index accordingly and assign them to the geometry.
```
geom.surfaces[0] = {'surface_idx': [[0,0], [0,1], [0,2], [0,3]], 'type': 'WallSurface'}
geom.surfaces[1] = {'surface_idx': [[0,4]], 'type': 'GroundSurface'}
geom.surfaces[2] = {'surface_idx': [[0,5]], 'type': 'RoofSurface'}
```
Then we test if it works.
```
ground = geom.get_surfaces('groundsurface')
ground_boundaries = []
for g in ground.values():
ground_boundaries.append(geom.get_surface_boundaries(g))
```
We have a list of generators
```
res = list(ground_boundaries[0])
```
The generator creates a list of surfaces --> a MultiSurface
```
assert res[0] == bdry[4]
# %%
wall = geom.get_surfaces('wallsurface')
wall_boundaries = []
for w in wall.values():
wall_boundaries.append(geom.get_surface_boundaries(w))
```
We put everything together, first filling up the CityObject
```
co.geometry.append(geom)
co.type = 'Building'
```
Then adding the CityObject to the CityModel.
```
cm.cityobjects[co.id] = co
```
Let's validate the citymodel before writing it to a file. However, first we
need to index the geometry boundaries and create the vertex list, second we
need to add the cityobject and vertices to the internal json-store of the
citymodel so the `validate()` method can validate them.
Note: CityJSON version 1.0.0 only accepts the Geometry `lod` as a numeric
value and not a string.
```
cityobjects, vertex_lookup = cm.reference_geometry()
cm.add_to_j(cityobjects,vertex_lookup)
cm.update_bbox()
#cm.validate(folder_schemas=schema_dir)
cm
```
Finally, we write the citymodel to a CityJSON file.
```
outfile = data_dir / 'test_create.json'
cityjson.save(cm, outfile)
```
| github_jupyter |
# Image classification of simulated AT-TPC events
Welcome to this project in applied machine learning. In this project we will tackle a simple classification problem of two different classes. The classes are simulated reaction types for the Ar(p, p') experiment conducted at MSU, in this task we'll focus on the classification task and simply treat the experiment as a black box.
### This is a completed notebook with solution examples, for your implementation we suggest you implement your own solution in the `project.ipynb` notebook
This project has three tasks with a recommendation for the time to spend on each task:
- Preparation, Data exploration and standardization: 0.5hr
- Model construction: 1hr
- Hyperparameter tuning and performance validation: 1hr
There is a notebook `project_solution.ipynb` included with suggestions to solutions for each task included, for reference or to easily move on to a part of the project more appealing to your interests.
## Preparation:
This project uses python and the machine learning library `keras`. As well as some functionality from `numpy` and `scikit-learn`. We recommend a `Python` verson of `>3.4`. These libraries should be installed to your specific system by using the command `pip3 install --user LIBRARY_NAME`
## Task 1: Data exploration and standardization
In machine learning, as in many other fields, the task of preparing data for analysis is as vital as it can be troublesome and tedious. In data-analysis the researcher can expect to spend the majority of their time merely processing data to prepare for analysis. In this projcet we will focus more on the entire pipeline of analysis, and so the data at hand has already been shaped to an image format suitable for our analysis.
## Task 1a: Loading the data
The data is stored in the `.npy` format using vecotrized code to speed up the read process. The files pointed to in this task are downsampled images with dimensions $64 x 64$ (if the images are to big for your laptop to handle, the script included in `../scripts/downsample_images.py` can further reduce the dimension).
```
import numpy as np # we'll be using this shorthand for the NumPy library throughout
dataset = np.load("../data/images/project_data.npy")
n_samples = dataset.shape[0]
print("Data shape: ", dataset.shape)
```
## Task 1b: Inspecting the data
The data is stored as xy projections of the real events, who take place in a 3d volume. This allows a simple exploratiuon of the data as images. In this task you should plot a few different events in a grid using `matplotlib.pyplot`
```
import matplotlib.pyplot as plt
rows = 2
cols = 2
n_plots = rows*cols
fig, axs = plt.subplots(nrows=rows, ncols=cols, figsize=(10, 10 ))
for row in axs:
for ax in row:
"""
one of pythons most wonderful attributes is that if an object is iterable it can be
directly iterated over, like above.
ax is an axis object from the 2d array of axis objects
"""
which = np.random.randint(0, n_samples)
ax.imshow(dataset[which].reshape(64, 64))
ax.axis("off")
```
## Task 1c: Standardizing the data
An important part of the preprocessing of data is the standardization of the input. The intuition here is simply that the model should expect similar values in the input to mean the same.
You should implement a standardization of the input. Perhaps the most common standardization is the centering of the mean of the distribution, and scaling by the standard deviation:
$X_s = \frac{X - \mu}{\sigma}$
Note that for our data we only want to standardize the signal part of our image, we know the rest is zero and we don't want the standardization to be unduly effected. This also means we don't necessarily want a zero mean for our signal distribution. So for this example we stick with the scaling:
$X_s = \frac{X}{\sigma}$
Another important fact is that at already at this point is it recommended to separate the data in train and test sets. The partion of the data to test on should be roughly 10-20%. And to remember to compute the standardization variables only from the training set.
### MORE OPEN ENDED !!!
```
from sklearn.model_selection import train_test_split
targets = np.load("../data/targets/project_targets.npy")
train_X, test_X, train_y, test_y = train_test_split(dataset, targets, test_size=0.15)
nonzero_indices = np.nonzero(train_X)
nonzero_elements = train_X[nonzero_indices]
print("Train Mean: ", nonzero_elements.mean())
print("Train Std.: ", nonzero_elements.std())
print("-------------")
print("Test Mean: ", test_X[np.nonzero(test_X)].mean())
print("Test Std.: ", test_X[np.nonzero(test_X)].std())
print("############")
nonzero_scaled = nonzero_elements/nonzero_elements.std()
train_X[nonzero_indices] = nonzero_scaled
test_X[np.nonzero(test_X)] /= nonzero_elements.std()
print("Train Mean: ", nonzero_scaled.mean())
print("Train Std.: ", nonzero_scaled.std())
print("-------------")
print("Test Mean: ", test_X[np.nonzero(test_X)].mean())
print("Test Std.: ", test_X[np.nonzero(test_X)].std())
```
#### We also want to plot up the data again to confirm that our scaling is sensible, you should reuse your code from above for this.
```
import matplotlib.pyplot as plt
rows = 2
cols = 2
n_plots = rows*cols
fig, axs = plt.subplots(nrows=rows, ncols=cols, figsize=(10, 10 ))
for row in axs:
for ax in row:
"""
one of pythons most wonderful attributes is that if an object is iterable it can be
directly iterated over, like above.
ax is an axis object from the 2d array of axis objects
"""
which = np.random.randint(0, train_X.shape[0])
ax.imshow(train_X[which].reshape(64, 64))
ax.text(5, 5, "{}".format(int(train_y[which])), bbox={'facecolor': 'white', 'pad': 10})
ax.axis("off")
```
## 1d: Encoding the targets:
For classification one ordinarily encodes the target as a n-element zero vector with one element valued at 1 indicating the target class. This is simply called one-hot encoding.
You should inspect the values of the target vectors and use the imported `OneHotEncoder` to convert the targets.
Note that this is not necessary for the two class case, but we do it to demonstrate a general approach.
```
from sklearn.preprocessing import OneHotEncoder
onehot_train_y = OneHotEncoder(sparse=False, categories="auto").fit_transform(train_y.reshape(-1, 1))
onehot_test_y = OneHotEncoder(sparse=False, categories="auto").fit_transform(test_y.reshape(-1, 1))
print("Onehot train targets:", onehot_train_y.shape)
print("Onehot test targets:",onehot_test_y.shape)
```
## Prelude to task 2:
In this task we'll be constructing a CNN model for the two class data. Before that it is useful to characterize the performance of a less complex model, for example a logistic regression model. For this task then you should construct a logistic regression model to classify the two class problem.
```
from keras.models import Sequential, Model
from keras.layers import Dense
from keras.regularizers import l2
from keras.optimizers import SGD, adam
flat_train_X = np.reshape(train_X, (train_X.shape[0], train_X.shape[1]*train_X.shape[2]*train_X.shape[3]))
flat_test_X = np.reshape(test_X, (test_X.shape[0], train_X.shape[1]*train_X.shape[2]*train_X.shape[3]))
logreg = Sequential()
logreg.add(Dense(2, kernel_regularizer=l2(0.01), activation="softmax"))
eta = 0.001
optimizer = SGD(eta)
logreg.compile(optimizer, loss="binary_crossentropy", metrics=["accuracy",])
history = logreg.fit(
x=flat_train_X,
y=onehot_train_y,
batch_size=100,
epochs=200,
validation_split=0.15,
verbose=2
)
```
The logistic regression model doesn't work, clearly. What about the data prohobits it from doing so?
## 2a: Creating a model
In this task we will create a CNN with fully connected bottom-layers for classification. You should base your code on Morten's code for a model. We suggest you complete one of the following for this task:
1. Implement a class or function `cnn` that returns a compiled Keras model with an arbitrary number of convolutional and fully connected layers with optional configuration of regularization terms or layers.
2. Implement a simple hard-coded function `cnn` that returns a Keras model object. The architecture should be specified in the function.
Both implementations should include multiple convolutional layers and ending with a couple fully connected layers. The output of the network should be a softmax or log-softmax layer of logits.
You should experiment with where in the network you place the non-linearities and whether to use striding or pooling to reduce the input. As well as the use of padding, would you need one for the first layer?
```
model_config = {
"n_conv":2,
"receptive_fields":[3, 3],
"strides":[1, 1,],
"n_filters":[2, 2],
"conv_activation":[1, 1],
"max_pool":[1, 1],
"n_dense":1,
"neurons":[10,],
"dense_activation":[1,]
}
from keras.models import Sequential, Model
from keras.layers import Dense, Conv2D, Flatten, MaxPooling2D, ReLU, Input, Softmax
from keras.regularizers import l2
def create_convolutional_neural_network_keras(input_shape, config, n_classes=2):
"""
Modified from MH Jensen's course on machine learning in physics:
https://github.com/CompPhysics/MachineLearningMSU/blob/master/doc/pub/CNN/ipynb/CNN.ipynb
"""
model=Sequential()
for i in range(config["n_conv"]):
receptive_field = config["receptive_fields"][i]
strides = config["strides"][i]
n_filters = config["n_filters"][i]
pad = "same" if i == 0 else "same"
input_shape = input_shape if i==0 else None
if i == 0:
conv = Conv2D(
n_filters,
(receptive_field, receptive_field),
input_shape=input_shape,
padding=pad,
strides=strides,
kernel_regularizer=l2(0.01)
)
else:
conv = Conv2D(
n_filters,
(receptive_field, receptive_field),
padding=pad,
strides=strides,
kernel_regularizer=l2(0.01)
)
model.add(conv)
pool = config["max_pool"][i]
activation = config["conv_activation"][i]
if activation:
model.add(ReLU())
if pool:
model.add(MaxPooling2D(2))
model.add(Flatten())
for i in range(config["n_dense"]):
n_neurons = config["neurons"][i]
model.add(
Dense(
n_neurons,
kernel_regularizer=l2(0.01)
))
activation = config["dense_activation"][i]
if activation:
model.add(ReLU())
model.add(
Dense(
n_classes,
activation='softmax',
kernel_regularizer=l2(0.01))
)
return model
model_o = create_convolutional_neural_network_keras(train_X.shape[1:], model_config, n_classes=2)
#model_o = mhj(train_X.shape[1:], 3, 2, 10, 2, 0.01)
print(model_o.summary())
```
## 2b: Plot your model
`Keras` provides a convenient class for plotting your model architecture. You should both do this and inspect the model summary to see how many trainable parameters you have as well as to confirm that your model is reasonably put together with no dangling edges in the graph etc.
```
from keras.utils import plot_model
plot_model(model_o, to_file="convnet.png")
```

## 2b: Compiling your model
With the constructed model ready it can now be compiled. Compiling entails unrolling the computational graph underlying the model and attaching losses at the layers you specify. For more complex models one can attach loss functions at arbitrary layers or one could define a specific loss for your particular problem.
For our case we will simply use a categorical cross-entropy, which means our network parametrizes an output of logits which we softmax to produce probabilities. In this task you should simply compile the above model with an optimizer of your choice and a categorical cross-entropy loss.
```
eta = 0.01
sgd = SGD(lr=eta, )
adam = adam(lr=eta, beta_1=0.5, )
model_o.compile(loss='binary_crossentropy', optimizer=adam, metrics=['accuracy'])
```
## 2c: Running your model
Here you should simply use the `.fit` method of the model to train on the training set. Select a suitable subset of
train to use as validation, this should not be the test-set. Take care to note how many trainable parameters your model has. A model with $10^5$ parameters takes about a minute per epoch to run on a 7th gen i9 intel processor. If your laptop has a nvidia GPU training should be considerably faster.
Hint: this model is quite easy to over-fit, you should build your network with a relatively low complexity ($10^3$ parameters).
The `.fit` method returns a `history` object that you can use to plot the progress of your training.
```
%matplotlib notebook
import matplotlib.pyplot as plt
history = model_o.fit(
x=train_X,
y=onehot_train_y,
batch_size=50,
epochs=40,
validation_split=0.15,
verbose=2
)
# copied from https://keras.io/visualization/
# Plot training & validation accuracy values
fig, axs= plt.subplots(figsize=(10, 8), nrows=2)
fig.suptitle('Model performance')
axs[0].plot(history.history['acc'], "x-",alpha=0.8)
axs[0].plot(history.history['val_acc'], "x-", alpha=0.8)
axs[0].set_ylabel('Accuracy')
axs[0].set_xlabel('Epoch')
axs[0].legend(['Train', 'Test'], loc='upper left')
# Plot training & validation loss values
axs[1].plot(history.history['loss'], "o-",alpha=0.8)
axs[1].plot(history.history['val_loss'], "o-", alpha=0.8)
axs[1].set_ylabel('Loss')
axs[1].legend(['Train', 'Test'], loc='upper left')
```
## 3a: Performance validation
As mentioned in the lectures machine learning models suffer from the problem of overfitting as scaling to an almost arbitrary complexity is simple with todays hardware. The challenge then is often that of finding the correct architecture and type of model suitable for your problem. In this task you will be familiarized with some tools to monitor and estimate the degree of overfitting.
In task 1c you separated your data in training and test sets. The test set is used to estimate generalization performance. Typically we use this to fine-tune the hyperparameters. Another trick is to use a validation set during training, which we implemented in the last task for the training.
In this task we will start with attaching callbacks to the fitting process. They are listed in the documentation for `Keras`here: https://keras.io/callbacks/
Pick ones you think are suitable for our problem and re-run the training from above.
```
from keras.callbacks import EarlyStopping, ModelCheckpoint
callbacks = [EarlyStopping(min_delta=0.0001, patience=4), ModelCheckpoint("../checkpoints/ckpt")
history = model_o.fit(
x=train_X,
y=onehot_train_y,
batch_size=50,
epochs=150,
validation_split=0.15,
verbose=2,
callbacks=callbacks
)
```
## 3b: Hyperparameter tuning
Hyperparameters, like the number of layers, learning rate or others can have a very big impact on the model quality. The model performance should also be statistically quantified using cross validation, bootstrapped confidence intervals for your performance metrics or other tools depending on model.
In this task you should then implement a function or for loop doing either random search or a grid search over parameters and finally you should plot those results in a suitable way.
| github_jupyter |
### Import Package
```
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
import numpy as np
import traceback
import contextlib
import pathlib
```
### Load Dataset
```
mnist = tf.keras.datasets.fashion_mnist
(X_train, y_train), (X_test, y_test) = mnist.load_data()
print("Train Image shape:", X_train.shape, "Test Image shape:", X_test.shape)
# Normalize the images
X_train = X_train / 255.0
X_test = X_test / 255.0
```
### Conv2D Model - Base Model
```
model = keras.Sequential([
keras.layers.InputLayer(input_shape=(28, 28)),
keras.layers.Reshape(target_shape=(28, 28, 1)),
keras.layers.Conv2D(filters=12, kernel_size=(3, 3), activation='relu'),
keras.layers.MaxPooling2D(pool_size=(2, 2)),
keras.layers.Flatten(),
keras.layers.Dense(10)
])
# Model summary
model.summary()
```
### Train Conv2D
```
model.compile(optimizer='adam',
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
model.fit(X_train,
y_train,
batch_size=64,
epochs=10,
validation_data=(X_test, y_test))
# Saving Model
model.save('1_fashion_mnist_model.h5')
# Evaluate the model on test set
score = model.evaluate(X_test, y_test, verbose=0)
# Print test accuracy
print('\n', 'Test accuracy:', score[1])
```
### Train model with pruning
```
! pip install -q tensorflow-model-optimization
import tensorflow_model_optimization as tfmot
prune_low_magnitude = tfmot.sparsity.keras.prune_low_magnitude
# Compute end step to finish pruning after 2 epochs.
batch_size = 128
epochs = 40
validation_split = 0.1 # 10% of training set will be used for validation set.
num_images = X_train.shape[0] * (1 - validation_split)
end_step = np.ceil(num_images / batch_size).astype(np.int32) * epochs
# Define model for pruning.
pruning_params = {
'pruning_schedule': tfmot.sparsity.keras.PolynomialDecay(initial_sparsity=0.50,
final_sparsity=0.80,
begin_step=0,
end_step=end_step)
}
model_for_pruning = prune_low_magnitude(model, **pruning_params)
# `prune_low_magnitude` requires a recompile.
model_for_pruning.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
model_for_pruning.summary()
callbacks = [
tfmot.sparsity.keras.UpdatePruningStep(),
tfmot.sparsity.keras.PruningSummaries(log_dir='log'),
]
model_for_pruning.fit(X_train, y_train,
batch_size=batch_size, epochs=epochs, validation_split=validation_split,
callbacks=callbacks)
_, model_for_pruning_accuracy = model_for_pruning.evaluate(
X_train, y_train, verbose=0)
print('Pruned test accuracy:', model_for_pruning_accuracy)
model_for_export = tfmot.sparsity.keras.strip_pruning(model_for_pruning)
tf.keras.models.save_model(model_for_export, '2_fashion_mnist_model_pruning.h5', include_optimizer=False)
```
### Q-aware Training
```
import tensorflow_model_optimization as tfmot
quantize_model = tfmot.quantization.keras.quantize_model
# q_aware stands for for quantization aware.
q_aware_model = quantize_model(model)
# `quantize_model` requires a recompile.
q_aware_model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
q_aware_model.summary()
# Train and evaluate the model against baseline
train_images_subset = X_train[0:1000] # out of 60000
train_labels_subset = y_train[0:1000]
q_aware_model.fit(train_images_subset, train_labels_subset,
batch_size=10, epochs=50, validation_split=0.1)
# Evaluate the model on test set
import time
start_time_infer = time.time()
score = q_aware_model.evaluate(X_test, y_test, verbose=0)
# Print test accuracy
result = {'Time to full set infer': (time.time() - start_time_infer),
'Score' : score[1]}
print(result)
start_time_infer = time.time()
#model = tf.keras.models.load_model('fashion_mnist_model_qaware.h5', compile = True)
data = X_test[0]
data = data.reshape((1, 28, 28))
data_y = y_test[0:1]
score = q_aware_model.evaluate(data, data_y, verbose=0)
result = {'Time to single unit infer': (time.time() - start_time_infer),
'Score' : score[1]}
print(result)
q_aware_model.save('3_fashion_mnist_model_qaware.h5')
```
### Convert Model to TFLite
```
def ConvertTFLite(model_path, filename):
try:
# Loading Model
model = tf.keras.models.load_model(model_path)
# Converter
converter = tf.lite.TFLiteConverter.from_keras_model(model)
tflite_model = converter.convert()
#Specify path
tflite_models_dir = pathlib.Path("tflite_models/")
tflite_models_dir.mkdir(exist_ok=True, parents=True)
filename = filename+".tflite"
tflite_model_file = tflite_models_dir/filename
# Save Model
tflite_model_file.write_bytes(tflite_model)
return f'Converted to TFLite, path {tflite_model_file}'
except Exception as e:
return str(e)
ConvertTFLite('./1_fashion_mnist_model.h5','4_fashion_mnist_model')
ConvertTFLite('./2_fashion_mnist_model_pruning.h5','5_fashion_mnist_pruning_model')
converter = tf.lite.TFLiteConverter.from_keras_model(q_aware_model)
quantized_tflite_model = converter.convert()
quantized_aware_tflite_file = '6_fashion_mnist_model_qaware.tflite'
with open(quantized_aware_tflite_file, 'wb') as f:
f.write(quantized_tflite_model)
print('Saved quvantaised aware TFLite model to:', quantized_aware_tflite_file)
```
### Integer with Float fallback quantaization
```
def Quant_int_with_float(model_name, filename):
try:
model = tf.keras.models.load_model(model_name)
converter = tf.lite.TFLiteConverter.from_keras_model(model)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
tflite_model_quant = converter.convert()
filename = filename+'.tflite'
tflite_models_dir = pathlib.Path("tflite_models/")
tflite_models_dir.mkdir(exist_ok=True, parents=True)
tflite_model_quant_file = tflite_models_dir/filename
tflite_model_quant_file.write_bytes(tflite_model_quant)
return f'Converted - path {tflite_model_quant_file}'
except Exception as e:
return str(e)
Quant_int_with_float('./1_fashion_mnist_model.h5', '7_fashion_mnist_Integer_float_model')
Quant_int_with_float('./2_fashion_mnist_model_pruning.h5','8_fashion_mnist_pruning_Integer_float_model')
converter = tf.lite.TFLiteConverter.from_keras_model(q_aware_model)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
mnist_train, _ = tf.keras.datasets.fashion_mnist.load_data()
images = tf.cast(mnist_train[0], tf.float32) / 255.0
mnist_ds = tf.data.Dataset.from_tensor_slices((images)).batch(1)
def representative_data_gen():
for input_value in mnist_ds.take(100):
yield [input_value]
converter.representative_dataset = representative_data_gen
quantized_tflite_model = converter.convert()
tflite_models_dir = pathlib.Path("tflite_models/")
tflite_models_dir.mkdir(exist_ok=True, parents=True)
tflite_model_quant_file = tflite_models_dir/"9_fashion_mnist_Qaware_Integer_float_model.tflite"
tflite_model_quant_file.write_bytes(quantized_tflite_model)
```
### Float 16 Quantization
```
def Quant_float(model_name, filename):
try:
model = tf.keras.models.load_model(model_name)
converter = tf.lite.TFLiteConverter.from_keras_model(model)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter.target_spec.supported_types = [tf.float16]
tflite_fp16_model = converter.convert()
filename = filename+'.tflite'
tflite_models_fp16_dir = pathlib.Path("tflite_models/")
tflite_models_fp16_dir.mkdir(exist_ok=True, parents=True)
tflite_model_fp16_file = tflite_models_fp16_dir/filename
tflite_model_fp16_file.write_bytes(tflite_fp16_model)
return f'Converted - path {tflite_model_fp16_file}'
except Exception as e:
return str(e)
Quant_float('./1_fashion_mnist_model.h5', '10_fashion_mnist_float16_model')
Quant_float('./2_fashion_mnist_model_pruning.h5', '11_fashion_mnist_float_pruning_model')
converter = tf.lite.TFLiteConverter.from_keras_model(q_aware_model)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter.target_spec.supported_types = [tf.float16]
tflite_fp16_model = converter.convert()
tflite_model_fp16_file = tflite_models_dir/"12_fashion_mnist_Qaware_float16_model.tflite"
tflite_model_fp16_file.write_bytes(tflite_fp16_model)
```
### Integer Only
```
def Quant_integer(model_name, filename):
try:
model = tf.keras.models.load_model(model_name)
converter = tf.lite.TFLiteConverter.from_keras_model(model)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
mnist_train, _ = tf.keras.datasets.fashion_mnist.load_data()
images = tf.cast(mnist_train[0], tf.float32) / 255.0
mnist_ds = tf.data.Dataset.from_tensor_slices((images)).batch(1)
def representative_data_gen():
for input_value in mnist_ds.take(100):
yield [input_value]
converter.representative_dataset = representative_data_gen
converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS_INT8]
converter.inference_input_type = tf.int8 # or tf.uint8
converter.inference_output_type = tf.int8 # or tf.uint8
tflite_int_quant_model = converter.convert()
filename = filename+'.tflite'
tflite_models_dir = pathlib.Path("tflite_models/")
tflite_models_dir.mkdir(exist_ok=True, parents=True)
tflite_model_integeronly_file = tflite_models_dir/filename
tflite_model_integeronly_file.write_bytes(tflite_int_quant_model)
return f'Converted - path {tflite_model_integeronly_file}'
except Exception as e:
return str(e)
Quant_integer('./1_fashion_mnist_model.h5', '13_fashion_mnist_integeronly_model')
Quant_integer('./2_fashion_mnist_model_pruning.h5', '14_fashion_mnist_Integeronly_pruning_model')
# Quant_integer('3_fashion_mnist_model_qaware.h5','15_fashion_mnist_qaware_integer_model')
```
### Evalvate Model
```
import time
```
### Keras model Evaluation
```
def evaluate_keras_model_single_unit(model_path):
start_time_infer = time.time()
model = tf.keras.models.load_model(model_path, compile = True)
model.compile(optimizer='adam',
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
data = X_test[0]
data = data.reshape((1, 28, 28))
data_y = y_test[0:1]
score = model.evaluate(data, data_y, verbose=0)
result = {'Time to single unit infer': (time.time() - start_time_infer),
'Score' : score[1]}
return result
evaluate_keras_model_single_unit('./1_fashion_mnist_model.h5')
evaluate_keras_model_single_unit('./2_fashion_mnist_model_pruning.h5')
def evaluate_keras_model_test_set(model_path):
start_time_infer = time.time()
model = tf.keras.models.load_model(model_path, compile = True)
model.compile(optimizer='adam',
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
score = score = model.evaluate(X_test, y_test, verbose =0)
result = {'Time to single unit infer': (time.time() - start_time_infer),
'Score' : score[1]}
return result
evaluate_keras_model_test_set('./1_fashion_mnist_model.h5')
evaluate_keras_model_test_set('./2_fashion_mnist_model_pruning.h5')
```
### TF Lite Model Evaluvation
```
# Evaluate the mode
def evaluate_tflite_model_test_set(interpreter):
start_time = time.time()
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
# Run predictions on every image in the "test" dataset.
prediction_digits = []
for test_image in X_test:
# Pre-processing: add batch dimension and convert to float32 to match with
# the model's input data format.
test_image = np.expand_dims(test_image, axis=0).astype(np.float32)
interpreter.set_tensor(input_index, test_image)
# Run inference.
interpreter.invoke()
# Post-processing: remove batch dimension and find the digit with highest
# probability.
output = interpreter.tensor(output_index)
digit = np.argmax(output()[0])
prediction_digits.append(digit)
# Compare prediction results with ground truth labels to calculate accuracy.
accurate_count = 0
for index in range(len(prediction_digits)):
if prediction_digits[index] == y_test[index]:
accurate_count += 1
accuracy = accurate_count * 1.0 / len(prediction_digits)
results = {'time': (time.time() - start_time),
'accuracy': accuracy}
return results
```
### TF Lite Models
```
# TF Lite
tflite_model_file = 'tflite_models/4_fashion_mnist_model.tflite'
interpreter = tf.lite.Interpreter(model_path=str(tflite_model_file))
interpreter.allocate_tensors()
evaluate_tflite_model_test_set(interpreter)
# Purning TF Lite
tflite_pruning_model_file = 'tflite_models/5_fashion_mnist_pruning_model.tflite'
interpreter_pruning = tf.lite.Interpreter(model_path=str(tflite_pruning_model_file))
interpreter_pruning.allocate_tensors()
evaluate_tflite_model_test_set(interpreter_pruning)
# Qaware Model
tflite_model_file = '6_fashion_mnist_model_qaware.tflite'
interpreter_qaware = tf.lite.Interpreter(model_path=str(tflite_model_file))
interpreter_qaware.allocate_tensors()
evaluate_tflite_model_test_set(interpreter_qaware)
```
### Integer Float TF Lite models
```
# TF Lite
tflite_model_file = 'tflite_models/7_fashion_mnist_Integer_float_model.tflite'
interpreter_int_float = tf.lite.Interpreter(model_path=str(tflite_model_file))
interpreter_int_float.allocate_tensors()
evaluate_tflite_model_test_set(interpreter_int_float)
# Purning TF Lite
tflite_pruning_model_file = 'tflite_models/8_fashion_mnist_pruning_Integer_float_model.tflite'
interpreter_int_float_pruning = tf.lite.Interpreter(model_path=str(tflite_pruning_model_file))
interpreter_int_float_pruning.allocate_tensors()
evaluate_tflite_model_test_set(interpreter_int_float_pruning)
# Q-aware TF Lite
tflite_qaware_model_file = 'tflite_models/9_fashion_mnist_Qaware_Integer_float_model.tflite'
interpreter_tflite_qaware = tf.lite.Interpreter(model_path=str(tflite_qaware_model_file))
interpreter_tflite_qaware.allocate_tensors()
evaluate_tflite_model_test_set(interpreter_tflite_qaware)
```
### Float Tflite
```
# TF Lite
tflite_model_file = 'tflite_models/10_fashion_mnist_float16_model.tflite'
interpreter_float = tf.lite.Interpreter(model_path=str(tflite_model_file))
interpreter_float.allocate_tensors()
evaluate_tflite_model_test_set(interpreter_float)
# Purning TF Lite
tflite_pruning_model_file = 'tflite_models/11_fashion_mnist_float_pruning_model.tflite'
interpreter_float_pruning = tf.lite.Interpreter(model_path=str(tflite_pruning_model_file))
interpreter_float_pruning.allocate_tensors()
evaluate_tflite_model_test_set(interpreter_float_pruning)
tflite_qaware_model_file = 'tflite_models/12_fashion_mnist_Qaware_float16_model.tflite'
interpreter_tflite_qaware = tf.lite.Interpreter(model_path=str(tflite_qaware_model_file))
interpreter_tflite_qaware.allocate_tensors()
evaluate_tflite_model_test_set(interpreter_tflite_qaware)
```
### Integer Only TFlite
```
# TF Lite
tflite_model_file = 'tflite_models/13_fashion_mnist_integeronly_model.tflite'
interpreter_int = tf.lite.Interpreter(model_path=str(tflite_model_file))
interpreter_int.allocate_tensors()
evaluate_tflite_model_test_set(interpreter_int)
# Purning TF Lite
tflite_pruning_model_file = 'tflite_models/14_fashion_mnist_Integeronly_pruning_model.tflite'
interpreter_int_pruning = tf.lite.Interpreter(model_path=str(tflite_pruning_model_file))
interpreter_int_pruning.allocate_tensors()
evaluate_tflite_model_test_set(interpreter_int_pruning)
# tflite_qaware_model_file = 'tflite_models/15_fashion_mnist_Qaware_integer_model.tflite'
# interpreter_tflite_qaware = tf.lite.Interpreter(model_path=str(tflite_qaware_model_file))
# interpreter_tflite_qaware.allocate_tensors()
# evaluate_tflite_model_test_set(interpreter_tflite_qaware)
```
### Find unit inference time
```
# Evaluate the mode
def evaluate_tflite_model_single_unit(interpreter):
start_time = time.time()
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
test_image = np.expand_dims(X_test[0], axis=0).astype(np.float32)
interpreter.set_tensor(input_index, test_image)
# Run inference.
interpreter.invoke()
# Post-processing: remove batch dimension and find the digit with highest
# probability.
output = interpreter.tensor(output_index)
results = {'time': (time.time() - start_time)}
return results
# TF Lite
evaluate_tflite_model_single_unit(interpreter)
evaluate_tflite_model_single_unit(interpreter_pruning)
evaluate_tflite_model_single_unit(interpreter_int_float)
evaluate_tflite_model_single_unit(interpreter_qaware)
evaluate_tflite_model_single_unit(interpreter_int_float_pruning)
evaluate_tflite_model_single_unit(interpreter_float)
evaluate_tflite_model_single_unit(interpreter_float_pruning)
evaluate_tflite_model_single_unit(interpreter_int)
evaluate_tflite_model_single_unit(interpreter_float_pruning)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/main/notebooks/bnn_hmc_gaussian.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# (SG)HMC for inferring params of a 2d Gaussian
Based on
https://github.com/google-research/google-research/blob/master/bnn_hmc/notebooks/mcmc_gaussian_test.ipynb
```
import jax
print(jax.devices())
!git clone https://github.com/google-research/google-research.git
%cd /content/google-research
!ls bnn_hmc
!pip install optax
```
# Setup
```
from jax.config import config
import jax
from jax import numpy as jnp
import numpy as onp
import numpy as np
import os
import sys
import time
import tqdm
import optax
import functools
from matplotlib import pyplot as plt
from bnn_hmc.utils import losses
from bnn_hmc.utils import train_utils
from bnn_hmc.utils import tree_utils
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
# Data and model
```
mu = jnp.zeros([2,])
# sigma = jnp.array([[1., .5], [.5, 1.]])
sigma = jnp.array([[1.e-4, 0], [0., 1.]])
sigma_l = jnp.linalg.cholesky(sigma)
sigma_inv = jnp.linalg.inv(sigma)
sigma_det = jnp.linalg.det(sigma)
onp.random.seed(0)
samples = onp.random.multivariate_normal(onp.asarray(mu), onp.asarray(sigma), size=1000)
plt.scatter(samples[:, 0], samples[:, 1], alpha=0.3)
plt.grid()
def log_density_fn(params):
assert params.shape == mu.shape, "Shape error"
diff = params - mu
k = mu.size
log_density = -jnp.log(2 * jnp.pi) * k / 2
log_density -= jnp.log(sigma_det) / 2
log_density -= diff.T @ sigma_inv @ diff / 2
return log_density
def log_likelihood_fn(_, params, *args, **kwargs):
return log_density_fn(params), jnp.array(jnp.nan)
def log_prior_fn(_):
return 0.
def log_prior_diff_fn(*args):
return 0.
fake_net_apply = None
fake_data = jnp.array([[jnp.nan,],]), jnp.array([[jnp.nan,],])
fake_net_state = jnp.array([jnp.nan,])
```
# HMC
```
step_size = 1e-1
trajectory_len = jnp.pi / 2
max_num_leapfrog_steps = int(trajectory_len // step_size + 1)
print("Leapfrog steps per iteration:", max_num_leapfrog_steps)
update, get_log_prob_and_grad = train_utils.make_hmc_update(
fake_net_apply, log_likelihood_fn, log_prior_fn, log_prior_diff_fn,
max_num_leapfrog_steps, 1., 0.)
# Initial log-prob and grad values
# params = jnp.ones_like(mu)[None, :]
params = jnp.ones_like(mu)
log_prob, state_grad, log_likelihood, net_state = (
get_log_prob_and_grad(fake_data, params, fake_net_state))
%%time
num_iterations = 500
all_samples = []
key = jax.random.PRNGKey(0)
for iteration in tqdm.tqdm(range(num_iterations)):
(params, net_state, log_likelihood, state_grad, step_size, key,
accept_prob, accepted) = (
update(fake_data, params, net_state, log_likelihood, state_grad,
key, step_size, trajectory_len, True))
if accepted:
all_samples.append(onp.asarray(params).copy())
# print("It: {} \t Accept P: {} \t Accepted {} \t Log-likelihood: {}".format(
# iteration, accept_prob, accepted, log_likelihood))
len(all_samples)
log_prob, state_grad, log_likelihood, net_state
all_samples_cat = onp.stack(all_samples)
plt.scatter(all_samples_cat[:, 0], all_samples_cat[:, 1], alpha=0.3)
plt.grid()
```
# Blackjax
```
!pip install blackjax
import jax
import jax.numpy as jnp
import jax.scipy.stats as stats
import matplotlib.pyplot as plt
import numpy as np
import blackjax.hmc as hmc
import blackjax.nuts as nuts
import blackjax.stan_warmup as stan_warmup
print(jax.devices())
potential = lambda x: -log_density_fn(**x)
num_integration_steps = 30
kernel_generator = lambda step_size, inverse_mass_matrix: hmc.kernel(
potential, step_size, inverse_mass_matrix, num_integration_steps
)
rng_key = jax.random.PRNGKey(0)
initial_position = {"params": np.zeros(2)}
initial_state = hmc.new_state(initial_position, potential)
print(initial_state)
%%time
nsteps = 500
final_state, (step_size, inverse_mass_matrix), info = stan_warmup.run(
rng_key,
kernel_generator,
initial_state,
nsteps,
)
%%time
kernel = nuts.kernel(potential, step_size, inverse_mass_matrix)
kernel = jax.jit(kernel)
def inference_loop(rng_key, kernel, initial_state, num_samples):
def one_step(state, rng_key):
state, _ = kernel(rng_key, state)
return state, state
keys = jax.random.split(rng_key, num_samples)
_, states = jax.lax.scan(one_step, initial_state, keys)
return states
%%time
nsamples = 500
states = inference_loop(rng_key, kernel, initial_state, nsamples)
samples = states.position["params"].block_until_ready()
print(samples.shape)
plt.scatter(samples[:, 0], samples[:, 1], alpha=0.3)
plt.grid()
```
| github_jupyter |
# Deep Learning & Art: Neural Style Transfer
In this assignment, you will learn about Neural Style Transfer. This algorithm was created by [Gatys et al. (2015).](https://arxiv.org/abs/1508.06576)
**In this assignment, you will:**
- Implement the neural style transfer algorithm
- Generate novel artistic images using your algorithm
Most of the algorithms you've studied optimize a cost function to get a set of parameter values. In Neural Style Transfer, you'll optimize a cost function to get pixel values!
## <font color='darkblue'>Updates</font>
#### If you were working on the notebook before this update...
* The current notebook is version "3a".
* You can find your original work saved in the notebook with the previous version name ("v2")
* To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#### List of updates
* Use `pprint.PrettyPrinter` to format printing of the vgg model.
* computing content cost: clarified and reformatted instructions, fixed broken links, added additional hints for unrolling.
* style matrix: clarify two uses of variable "G" by using different notation for gram matrix.
* style cost: use distinct notation for gram matrix, added additional hints.
* Grammar and wording updates for clarity.
* `model_nn`: added hints.
```
import os
import sys
import scipy.io
import scipy.misc
import matplotlib.pyplot as plt
from matplotlib.pyplot import imshow
from PIL import Image
from nst_utils import *
import numpy as np
import tensorflow as tf
import pprint
%matplotlib inline
```
## 1 - Problem Statement
Neural Style Transfer (NST) is one of the most fun techniques in deep learning. As seen below, it merges two images, namely: a **"content" image (C) and a "style" image (S), to create a "generated" image (G**).
The generated image G combines the "content" of the image C with the "style" of image S.
In this example, you are going to generate an image of the Louvre museum in Paris (content image C), mixed with a painting by Claude Monet, a leader of the impressionist movement (style image S).
<img src="images/louvre_generated.png" style="width:750px;height:200px;">
Let's see how you can do this.
## 2 - Transfer Learning
Neural Style Transfer (NST) uses a previously trained convolutional network, and builds on top of that. The idea of using a network trained on a different task and applying it to a new task is called transfer learning.
Following the [original NST paper](https://arxiv.org/abs/1508.06576), we will use the VGG network. Specifically, we'll use VGG-19, a 19-layer version of the VGG network. This model has already been trained on the very large ImageNet database, and thus has learned to recognize a variety of low level features (at the shallower layers) and high level features (at the deeper layers).
Run the following code to load parameters from the VGG model. This may take a few seconds.
```
pp = pprint.PrettyPrinter(indent=4)
model = load_vgg_model("pretrained-model/imagenet-vgg-verydeep-19.mat")
pp.pprint(model)
```
* The model is stored in a python dictionary.
* The python dictionary contains key-value pairs for each layer.
* The 'key' is the variable name and the 'value' is a tensor for that layer.
#### Assign input image to the model's input layer
To run an image through this network, you just have to feed the image to the model. In TensorFlow, you can do so using the [tf.assign](https://www.tensorflow.org/api_docs/python/tf/assign) function. In particular, you will use the assign function like this:
```python
model["input"].assign(image)
```
This assigns the image as an input to the model.
#### Activate a layer
After this, if you want to access the activations of a particular layer, say layer `4_2` when the network is run on this image, you would run a TensorFlow session on the correct tensor `conv4_2`, as follows:
```python
sess.run(model["conv4_2"])
```
## 3 - Neural Style Transfer (NST)
We will build the Neural Style Transfer (NST) algorithm in three steps:
- Build the content cost function $J_{content}(C,G)$
- Build the style cost function $J_{style}(S,G)$
- Put it together to get $J(G) = \alpha J_{content}(C,G) + \beta J_{style}(S,G)$.
### 3.1 - Computing the content cost
In our running example, the content image C will be the picture of the Louvre Museum in Paris. Run the code below to see a picture of the Louvre.
```
content_image = scipy.misc.imread("images/louvre.jpg")
imshow(content_image);
```
The content image (C) shows the Louvre museum's pyramid surrounded by old Paris buildings, against a sunny sky with a few clouds.
** 3.1.1 - Make generated image G match the content of image C**
#### Shallower versus deeper layers
* The shallower layers of a ConvNet tend to detect lower-level features such as edges and simple textures.
* The deeper layers tend to detect higher-level features such as more complex textures as well as object classes.
#### Choose a "middle" activation layer $a^{[l]}$
We would like the "generated" image G to have similar content as the input image C. Suppose you have chosen some layer's activations to represent the content of an image.
* In practice, you'll get the most visually pleasing results if you choose a layer in the **middle** of the network--neither too shallow nor too deep.
* (After you have finished this exercise, feel free to come back and experiment with using different layers, to see how the results vary.)
#### Forward propagate image "C"
* Set the image C as the input to the pretrained VGG network, and run forward propagation.
* Let $a^{(C)}$ be the hidden layer activations in the layer you had chosen. (In lecture, we had written this as $a^{[l](C)}$, but here we'll drop the superscript $[l]$ to simplify the notation.) This will be an $n_H \times n_W \times n_C$ tensor.
#### Forward propagate image "G"
* Repeat this process with the image G: Set G as the input, and run forward progation.
* Let $a^{(G)}$ be the corresponding hidden layer activation.
#### Content Cost Function $J_{content}(C,G)$
We will define the content cost function as:
$$J_{content}(C,G) = \frac{1}{4 \times n_H \times n_W \times n_C}\sum _{ \text{all entries}} (a^{(C)} - a^{(G)})^2\tag{1} $$
* Here, $n_H, n_W$ and $n_C$ are the height, width and number of channels of the hidden layer you have chosen, and appear in a normalization term in the cost.
* For clarity, note that $a^{(C)}$ and $a^{(G)}$ are the 3D volumes corresponding to a hidden layer's activations.
* In order to compute the cost $J_{content}(C,G)$, it might also be convenient to unroll these 3D volumes into a 2D matrix, as shown below.
* Technically this unrolling step isn't needed to compute $J_{content}$, but it will be good practice for when you do need to carry out a similar operation later for computing the style cost $J_{style}$.
<img src="images/NST_LOSS.png" style="width:800px;height:400px;">
**Exercise:** Compute the "content cost" using TensorFlow.
**Instructions**: The 3 steps to implement this function are:
1. Retrieve dimensions from `a_G`:
- To retrieve dimensions from a tensor `X`, use: `X.get_shape().as_list()`
2. Unroll `a_C` and `a_G` as explained in the picture above
- You'll likey want to use these functions: [tf.transpose](https://www.tensorflow.org/versions/r1.15/api_docs/python/tf/transpose) and [tf.reshape](https://www.tensorflow.org/versions/r1.15/api_docs/python/tf/reshape).
3. Compute the content cost:
- You'll likely want to use these functions: [tf.reduce_sum](https://www.tensorflow.org/api_docs/python/tf/reduce_sum), [tf.square](https://www.tensorflow.org/api_docs/python/tf/square) and [tf.subtract](https://www.tensorflow.org/api_docs/python/tf/subtract).
#### Additional Hints for "Unrolling"
* To unroll the tensor, we want the shape to change from $(m,n_H,n_W,n_C)$ to $(m, n_H \times n_W, n_C)$.
* `tf.reshape(tensor, shape)` takes a list of integers that represent the desired output shape.
* For the `shape` parameter, a `-1` tells the function to choose the correct dimension size so that the output tensor still contains all the values of the original tensor.
* So tf.reshape(a_C, shape=[m, n_H * n_W, n_C]) gives the same result as tf.reshape(a_C, shape=[m, -1, n_C]).
* If you prefer to re-order the dimensions, you can use `tf.transpose(tensor, perm)`, where `perm` is a list of integers containing the original index of the dimensions.
* For example, `tf.transpose(a_C, perm=[0,3,1,2])` changes the dimensions from $(m, n_H, n_W, n_C)$ to $(m, n_C, n_H, n_W)$.
* There is more than one way to unroll the tensors.
* Notice that it's not necessary to use tf.transpose to 'unroll' the tensors in this case but this is a useful function to practice and understand for other situations that you'll encounter.
```
# GRADED FUNCTION: compute_content_cost
def compute_content_cost(a_C, a_G):
"""
Computes the content cost
Arguments:
a_C -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing content of the image C
a_G -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing content of the image G
Returns:
J_content -- scalar that you compute using equation 1 above.
"""
### START CODE HERE ###
# Retrieve dimensions from a_G (≈1 line)
m, n_H, n_W, n_C = a_G.get_shape().as_list()
# Reshape a_C and a_G (≈2 lines)
a_C_unrolled = None
a_G_unrolled = None
# compute the cost with tensorflow (≈1 line)
J_content = 1 / (4 * n_H * n_W * n_C) * tf.reduce_sum((a_C - a_G)**2)
### END CODE HERE ###
return J_content
tf.reset_default_graph()
with tf.Session() as test:
tf.set_random_seed(1)
a_C = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
a_G = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
J_content = compute_content_cost(a_C, a_G)
print("J_content = " + str(J_content.eval()))
```
**Expected Output**:
<table>
<tr>
<td>
**J_content**
</td>
<td>
6.76559
</td>
</tr>
</table>
#### What you should remember
- The content cost takes a hidden layer activation of the neural network, and measures how different $a^{(C)}$ and $a^{(G)}$ are.
- When we minimize the content cost later, this will help make sure $G$ has similar content as $C$.
### 3.2 - Computing the style cost
For our running example, we will use the following style image:
```
style_image = scipy.misc.imread("images/monet_800600.jpg")
imshow(style_image);
```
This was painted in the style of *[impressionism](https://en.wikipedia.org/wiki/Impressionism)*.
Lets see how you can now define a "style" cost function $J_{style}(S,G)$.
### 3.2.1 - Style matrix
#### Gram matrix
* The style matrix is also called a "Gram matrix."
* In linear algebra, the Gram matrix G of a set of vectors $(v_{1},\dots ,v_{n})$ is the matrix of dot products, whose entries are ${\displaystyle G_{ij} = v_{i}^T v_{j} = np.dot(v_{i}, v_{j}) }$.
* In other words, $G_{ij}$ compares how similar $v_i$ is to $v_j$: If they are highly similar, you would expect them to have a large dot product, and thus for $G_{ij}$ to be large.
#### Two meanings of the variable $G$
* Note that there is an unfortunate collision in the variable names used here. We are following common terminology used in the literature.
* $G$ is used to denote the Style matrix (or Gram matrix)
* $G$ also denotes the generated image.
* For this assignment, we will use $G_{gram}$ to refer to the Gram matrix, and $G$ to denote the generated image.
#### Compute $G_{gram}$
In Neural Style Transfer (NST), you can compute the Style matrix by multiplying the "unrolled" filter matrix with its transpose:
<img src="images/NST_GM.png" style="width:900px;height:300px;">
$$\mathbf{G}_{gram} = \mathbf{A}_{unrolled} \mathbf{A}_{unrolled}^T$$
#### $G_{(gram)i,j}$: correlation
The result is a matrix of dimension $(n_C,n_C)$ where $n_C$ is the number of filters (channels). The value $G_{(gram)i,j}$ measures how similar the activations of filter $i$ are to the activations of filter $j$.
#### $G_{(gram),i,i}$: prevalence of patterns or textures
* The diagonal elements $G_{(gram)ii}$ measure how "active" a filter $i$ is.
* For example, suppose filter $i$ is detecting vertical textures in the image. Then $G_{(gram)ii}$ measures how common vertical textures are in the image as a whole.
* If $G_{(gram)ii}$ is large, this means that the image has a lot of vertical texture.
By capturing the prevalence of different types of features ($G_{(gram)ii}$), as well as how much different features occur together ($G_{(gram)ij}$), the Style matrix $G_{gram}$ measures the style of an image.
**Exercise**:
* Using TensorFlow, implement a function that computes the Gram matrix of a matrix A.
* The formula is: The gram matrix of A is $G_A = AA^T$.
* You may use these functions: [matmul](https://www.tensorflow.org/api_docs/python/tf/matmul) and [transpose](https://www.tensorflow.org/api_docs/python/tf/transpose).
```
# GRADED FUNCTION: gram_matrix
def gram_matrix(A):
"""
Argument:
A -- matrix of shape (n_C, n_H*n_W)
Returns:
GA -- Gram matrix of A, of shape (n_C, n_C)
"""
### START CODE HERE ### (≈1 line)
GA = tf.matmul(A, A, transpose_b=True)
### END CODE HERE ###
return GA
tf.reset_default_graph()
with tf.Session() as test:
tf.set_random_seed(1)
A = tf.random_normal([3, 2*1], mean=1, stddev=4)
GA = gram_matrix(A)
print("GA = \n" + str(GA.eval()))
```
**Expected Output**:
<table>
<tr>
<td>
**GA**
</td>
<td>
[[ 6.42230511 -4.42912197 -2.09668207] <br>
[ -4.42912197 19.46583748 19.56387138] <br>
[ -2.09668207 19.56387138 20.6864624 ]]
</td>
</tr>
</table>
### 3.2.2 - Style cost
Your goal will be to minimize the distance between the Gram matrix of the "style" image S and the gram matrix of the "generated" image G.
* For now, we are using only a single hidden layer $a^{[l]}$.
* The corresponding style cost for this layer is defined as:
$$J_{style}^{[l]}(S,G) = \frac{1}{4 \times {n_C}^2 \times (n_H \times n_W)^2} \sum _{i=1}^{n_C}\sum_{j=1}^{n_C}(G^{(S)}_{(gram)i,j} - G^{(G)}_{(gram)i,j})^2\tag{2} $$
* $G_{gram}^{(S)}$ Gram matrix of the "style" image.
* $G_{gram}^{(G)}$ Gram matrix of the "generated" image.
* Remember, this cost is computed using the hidden layer activations for a particular hidden layer in the network $a^{[l]}$
**Exercise**: Compute the style cost for a single layer.
**Instructions**: The 3 steps to implement this function are:
1. Retrieve dimensions from the hidden layer activations a_G:
- To retrieve dimensions from a tensor X, use: `X.get_shape().as_list()`
2. Unroll the hidden layer activations a_S and a_G into 2D matrices, as explained in the picture above (see the images in the sections "computing the content cost" and "style matrix").
- You may use [tf.transpose](https://www.tensorflow.org/versions/r1.15/api_docs/python/tf/transpose) and [tf.reshape](https://www.tensorflow.org/versions/r1.15/api_docs/python/tf/reshape).
3. Compute the Style matrix of the images S and G. (Use the function you had previously written.)
4. Compute the Style cost:
- You may find [tf.reduce_sum](https://www.tensorflow.org/api_docs/python/tf/reduce_sum), [tf.square](https://www.tensorflow.org/api_docs/python/tf/square) and [tf.subtract](https://www.tensorflow.org/api_docs/python/tf/subtract) useful.
#### Additional Hints
* Since the activation dimensions are $(m, n_H, n_W, n_C)$ whereas the desired unrolled matrix shape is $(n_C, n_H*n_W)$, the order of the filter dimension $n_C$ is changed. So `tf.transpose` can be used to change the order of the filter dimension.
* for the product $\mathbf{G}_{gram} = \mathbf{A}_{} \mathbf{A}_{}^T$, you will also need to specify the `perm` parameter for the `tf.transpose` function.
```
# GRADED FUNCTION: compute_layer_style_cost
def compute_layer_style_cost(a_S, a_G):
"""
Arguments:
a_S -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing style of the image S
a_G -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing style of the image G
Returns:
J_style_layer -- tensor representing a scalar value, style cost defined above by equation (2)
"""
### START CODE HERE ###
# Retrieve dimensions from a_G (≈1 line)
m, n_H, n_W, n_C = a_G.get_shape().as_list()
# Reshape the images to have them of shape (n_C, n_H*n_W) (≈2 lines)
a_S = tf.transpose(tf.reshape(a_S, shape=[-1, n_C]))
a_G = tf.transpose(tf.reshape(a_G, shape=[-1, n_C]))
# Computing gram_matrices for both images S and G (≈2 lines)
GS = gram_matrix(a_S)
GG = gram_matrix(a_G)
# Computing the loss (≈1 line)
J_style_layer = 1 / (2*n_C*n_H*n_W)**2 * tf.reduce_sum((GS-GG)**2)
### END CODE HERE ###
return J_style_layer
tf.reset_default_graph()
with tf.Session() as test:
tf.set_random_seed(1)
a_S = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
a_G = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
J_style_layer = compute_layer_style_cost(a_S, a_G)
print("J_style_layer = " + str(J_style_layer.eval()))
```
**Expected Output**:
<table>
<tr>
<td>
**J_style_layer**
</td>
<td>
9.19028
</td>
</tr>
</table>
### 3.2.3 Style Weights
* So far you have captured the style from only one layer.
* We'll get better results if we "merge" style costs from several different layers.
* Each layer will be given weights ($\lambda^{[l]}$) that reflect how much each layer will contribute to the style.
* After completing this exercise, feel free to come back and experiment with different weights to see how it changes the generated image $G$.
* By default, we'll give each layer equal weight, and the weights add up to 1. ($\sum_{l}^L\lambda^{[l]} = 1$)
```
STYLE_LAYERS = [
('conv1_1', 0.2),
('conv2_1', 0.2),
('conv3_1', 0.2),
('conv4_1', 0.2),
('conv5_1', 0.2)]
```
You can combine the style costs for different layers as follows:
$$J_{style}(S,G) = \sum_{l} \lambda^{[l]} J^{[l]}_{style}(S,G)$$
where the values for $\lambda^{[l]}$ are given in `STYLE_LAYERS`.
### Exercise: compute style cost
* We've implemented a compute_style_cost(...) function.
* It calls your `compute_layer_style_cost(...)` several times, and weights their results using the values in `STYLE_LAYERS`.
* Please read over it to make sure you understand what it's doing.
#### Description of `compute_style_cost`
For each layer:
* Select the activation (the output tensor) of the current layer.
* Get the style of the style image "S" from the current layer.
* Get the style of the generated image "G" from the current layer.
* Compute the "style cost" for the current layer
* Add the weighted style cost to the overall style cost (J_style)
Once you're done with the loop:
* Return the overall style cost.
```
def compute_style_cost(model, STYLE_LAYERS):
"""
Computes the overall style cost from several chosen layers
Arguments:
model -- our tensorflow model
STYLE_LAYERS -- A python list containing:
- the names of the layers we would like to extract style from
- a coefficient for each of them
Returns:
J_style -- tensor representing a scalar value, style cost defined above by equation (2)
"""
# initialize the overall style cost
J_style = 0
for layer_name, coeff in STYLE_LAYERS:
# Select the output tensor of the currently selected layer
out = model[layer_name]
# Set a_S to be the hidden layer activation from the layer we have selected, by running the session on out
a_S = sess.run(out)
# Set a_G to be the hidden layer activation from same layer. Here, a_G references model[layer_name]
# and isn't evaluated yet. Later in the code, we'll assign the image G as the model input, so that
# when we run the session, this will be the activations drawn from the appropriate layer, with G as input.
a_G = out
# Compute style_cost for the current layer
J_style_layer = compute_layer_style_cost(a_S, a_G)
# Add coeff * J_style_layer of this layer to overall style cost
J_style += coeff * J_style_layer
return J_style
```
**Note**: In the inner-loop of the for-loop above, `a_G` is a tensor and hasn't been evaluated yet. It will be evaluated and updated at each iteration when we run the TensorFlow graph in model_nn() below.
<!--
How do you choose the coefficients for each layer? The deeper layers capture higher-level concepts, and the features in the deeper layers are less localized in the image relative to each other. So if you want the generated image to softly follow the style image, try choosing larger weights for deeper layers and smaller weights for the first layers. In contrast, if you want the generated image to strongly follow the style image, try choosing smaller weights for deeper layers and larger weights for the first layers
!-->
## What you should remember
- The style of an image can be represented using the Gram matrix of a hidden layer's activations.
- We get even better results by combining this representation from multiple different layers.
- This is in contrast to the content representation, where usually using just a single hidden layer is sufficient.
- Minimizing the style cost will cause the image $G$ to follow the style of the image $S$.
### 3.3 - Defining the total cost to optimize
Finally, let's create a cost function that minimizes both the style and the content cost. The formula is:
$$J(G) = \alpha J_{content}(C,G) + \beta J_{style}(S,G)$$
**Exercise**: Implement the total cost function which includes both the content cost and the style cost.
```
# GRADED FUNCTION: total_cost
def total_cost(J_content, J_style, alpha = 10, beta = 40):
"""
Computes the total cost function
Arguments:
J_content -- content cost coded above
J_style -- style cost coded above
alpha -- hyperparameter weighting the importance of the content cost
beta -- hyperparameter weighting the importance of the style cost
Returns:
J -- total cost as defined by the formula above.
"""
### START CODE HERE ### (≈1 line)
J = alpha * J_content + beta * J_style
### END CODE HERE ###
return J
tf.reset_default_graph()
with tf.Session() as test:
np.random.seed(3)
J_content = np.random.randn()
J_style = np.random.randn()
J = total_cost(J_content, J_style)
print("J = " + str(J))
```
**Expected Output**:
<table>
<tr>
<td>
**J**
</td>
<td>
35.34667875478276
</td>
</tr>
</table>
## What you should remember
- The total cost is a linear combination of the content cost $J_{content}(C,G)$ and the style cost $J_{style}(S,G)$.
- $\alpha$ and $\beta$ are hyperparameters that control the relative weighting between content and style.
## 4 - Solving the optimization problem
Finally, let's put everything together to implement Neural Style Transfer!
Here's what the program will have to do:
1. Create an Interactive Session
2. Load the content image
3. Load the style image
4. Randomly initialize the image to be generated
5. Load the VGG19 model
7. Build the TensorFlow graph:
- Run the content image through the VGG19 model and compute the content cost
- Run the style image through the VGG19 model and compute the style cost
- Compute the total cost
- Define the optimizer and the learning rate
8. Initialize the TensorFlow graph and run it for a large number of iterations, updating the generated image at every step.
Lets go through the individual steps in detail.
#### Interactive Sessions
You've previously implemented the overall cost $J(G)$. We'll now set up TensorFlow to optimize this with respect to $G$.
* To do so, your program has to reset the graph and use an "[Interactive Session](https://www.tensorflow.org/api_docs/python/tf/InteractiveSession)".
* Unlike a regular session, the "Interactive Session" installs itself as the default session to build a graph.
* This allows you to run variables without constantly needing to refer to the session object (calling "sess.run()"), which simplifies the code.
#### Start the interactive session.
```
# Reset the graph
tf.reset_default_graph()
# Start interactive session
sess = tf.InteractiveSession()
```
#### Content image
Let's load, reshape, and normalize our "content" image (the Louvre museum picture):
```
content_image = scipy.misc.imread("images/louvre_small.jpg")
content_image = reshape_and_normalize_image(content_image)
```
#### Style image
Let's load, reshape and normalize our "style" image (Claude Monet's painting):
```
style_image = scipy.misc.imread("images/monet.jpg")
style_image = reshape_and_normalize_image(style_image)
```
#### Generated image correlated with content image
Now, we initialize the "generated" image as a noisy image created from the content_image.
* The generated image is slightly correlated with the content image.
* By initializing the pixels of the generated image to be mostly noise but slightly correlated with the content image, this will help the content of the "generated" image more rapidly match the content of the "content" image.
* Feel free to look in `nst_utils.py` to see the details of `generate_noise_image(...)`; to do so, click "File-->Open..." at the upper-left corner of this Jupyter notebook.
```
generated_image = generate_noise_image(content_image)
imshow(generated_image[0]);
```
#### Load pre-trained VGG19 model
Next, as explained in part (2), let's load the VGG19 model.
```
model = load_vgg_model("pretrained-model/imagenet-vgg-verydeep-19.mat")
```
#### Content Cost
To get the program to compute the content cost, we will now assign `a_C` and `a_G` to be the appropriate hidden layer activations. We will use layer `conv4_2` to compute the content cost. The code below does the following:
1. Assign the content image to be the input to the VGG model.
2. Set a_C to be the tensor giving the hidden layer activation for layer "conv4_2".
3. Set a_G to be the tensor giving the hidden layer activation for the same layer.
4. Compute the content cost using a_C and a_G.
**Note**: At this point, a_G is a tensor and hasn't been evaluated. It will be evaluated and updated at each iteration when we run the Tensorflow graph in model_nn() below.
```
# Assign the content image to be the input of the VGG model.
sess.run(model['input'].assign(content_image))
# Select the output tensor of layer conv4_2
out = model['conv4_2']
# Set a_C to be the hidden layer activation from the layer we have selected
a_C = sess.run(out)
# Set a_G to be the hidden layer activation from same layer. Here, a_G references model['conv4_2']
# and isn't evaluated yet. Later in the code, we'll assign the image G as the model input, so that
# when we run the session, this will be the activations drawn from the appropriate layer, with G as input.
a_G = out
# Compute the content cost
J_content = compute_content_cost(a_C, a_G)
```
#### Style cost
```
# Assign the input of the model to be the "style" image
sess.run(model['input'].assign(style_image))
# Compute the style cost
J_style = compute_style_cost(model, STYLE_LAYERS)
```
### Exercise: total cost
* Now that you have J_content and J_style, compute the total cost J by calling `total_cost()`.
* Use `alpha = 10` and `beta = 40`.
```
### START CODE HERE ### (1 line)
J = total_cost(J_content, J_style, alpha=10, beta=40)
### END CODE HERE ###
```
### Optimizer
* Use the Adam optimizer to minimize the total cost `J`.
* Use a learning rate of 2.0.
* [Adam Optimizer documentation](https://www.tensorflow.org/api_docs/python/tf/train/AdamOptimizer)
```
# define optimizer (1 line)
optimizer = tf.train.AdamOptimizer(2.0)
# define train_step (1 line)
train_step = optimizer.minimize(J)
```
### Exercise: implement the model
* Implement the model_nn() function.
* The function **initializes** the variables of the tensorflow graph,
* **assigns** the input image (initial generated image) as the input of the VGG19 model
* and **runs** the `train_step` tensor (it was created in the code above this function) for a large number of steps.
#### Hints
* To initialize global variables, use this:
```Python
sess.run(tf.global_variables_initializer())
```
* Run `sess.run()` to evaluate a variable.
* [assign](https://www.tensorflow.org/versions/r1.14/api_docs/python/tf/assign) can be used like this:
```python
model["input"].assign(image)
```
```
def model_nn(sess, input_image, num_iterations = 200):
# Initialize global variables (you need to run the session on the initializer)
### START CODE HERE ### (1 line)
sess.run(tf.global_variables_initializer())
### END CODE HERE ###
# Run the noisy input image (initial generated image) through the model. Use assign().
### START CODE HERE ### (1 line)
sess.run(model['input'].assign(input_image))
### END CODE HERE ###
for i in range(num_iterations):
# Run the session on the train_step to minimize the total cost
### START CODE HERE ### (1 line)
sess.run(train_step)
### END CODE HERE ###
# Compute the generated image by running the session on the current model['input']
### START CODE HERE ### (1 line)
generated_image = sess.run(model['input'])
### END CODE HERE ###
# Print every 20 iteration.
if i%20 == 0:
Jt, Jc, Js = sess.run([J, J_content, J_style])
print("Iteration " + str(i) + " :")
print("total cost = " + str(Jt))
print("content cost = " + str(Jc))
print("style cost = " + str(Js))
# save current generated image in the "/output" directory
save_image("output/" + str(i) + ".png", generated_image)
# save last generated image
save_image('output/generated_image.jpg', generated_image)
return generated_image
```
Run the following cell to generate an artistic image. It should take about 3min on CPU for every 20 iterations but you start observing attractive results after ≈140 iterations. Neural Style Transfer is generally trained using GPUs.
```
model_nn(sess, generated_image)
```
**Expected Output**:
<table>
<tr>
<td>
**Iteration 0 : **
</td>
<td>
total cost = 5.05035e+09 <br>
content cost = 7877.67 <br>
style cost = 1.26257e+08
</td>
</tr>
</table>
You're done! After running this, in the upper bar of the notebook click on "File" and then "Open". Go to the "/output" directory to see all the saved images. Open "generated_image" to see the generated image! :)
You should see something the image presented below on the right:
<img src="images/louvre_generated.png" style="width:800px;height:300px;">
We didn't want you to wait too long to see an initial result, and so had set the hyperparameters accordingly. To get the best looking results, running the optimization algorithm longer (and perhaps with a smaller learning rate) might work better. After completing and submitting this assignment, we encourage you to come back and play more with this notebook, and see if you can generate even better looking images.
Here are few other examples:
- The beautiful ruins of the ancient city of Persepolis (Iran) with the style of Van Gogh (The Starry Night)
<img src="images/perspolis_vangogh.png" style="width:750px;height:300px;">
- The tomb of Cyrus the great in Pasargadae with the style of a Ceramic Kashi from Ispahan.
<img src="images/pasargad_kashi.png" style="width:750px;height:300px;">
- A scientific study of a turbulent fluid with the style of a abstract blue fluid painting.
<img src="images/circle_abstract.png" style="width:750px;height:300px;">
## 5 - Test with your own image (Optional/Ungraded)
Finally, you can also rerun the algorithm on your own images!
To do so, go back to part 4 and change the content image and style image with your own pictures. In detail, here's what you should do:
1. Click on "File -> Open" in the upper tab of the notebook
2. Go to "/images" and upload your images (requirement: (WIDTH = 300, HEIGHT = 225)), rename them "my_content.png" and "my_style.png" for example.
3. Change the code in part (3.4) from :
```python
content_image = scipy.misc.imread("images/louvre.jpg")
style_image = scipy.misc.imread("images/claude-monet.jpg")
```
to:
```python
content_image = scipy.misc.imread("images/my_content.jpg")
style_image = scipy.misc.imread("images/my_style.jpg")
```
4. Rerun the cells (you may need to restart the Kernel in the upper tab of the notebook).
You can share your generated images with us on social media with the hashtag #deeplearniNgAI or by direct tagging!
You can also tune your hyperparameters:
- Which layers are responsible for representing the style? STYLE_LAYERS
- How many iterations do you want to run the algorithm? num_iterations
- What is the relative weighting between content and style? alpha/beta
## 6 - Conclusion
Great job on completing this assignment! You are now able to use Neural Style Transfer to generate artistic images. This is also your first time building a model in which the optimization algorithm updates the pixel values rather than the neural network's parameters. Deep learning has many different types of models and this is only one of them!
## What you should remember
- Neural Style Transfer is an algorithm that given a content image C and a style image S can generate an artistic image
- It uses representations (hidden layer activations) based on a pretrained ConvNet.
- The content cost function is computed using one hidden layer's activations.
- The style cost function for one layer is computed using the Gram matrix of that layer's activations. The overall style cost function is obtained using several hidden layers.
- Optimizing the total cost function results in synthesizing new images.
# Congratulations on finishing the course!
This was the final programming exercise of this course. Congratulations--you've finished all the programming exercises of this course on Convolutional Networks! We hope to also see you in Course 5, on Sequence models!
### References:
The Neural Style Transfer algorithm was due to Gatys et al. (2015). Harish Narayanan and Github user "log0" also have highly readable write-ups from which we drew inspiration. The pre-trained network used in this implementation is a VGG network, which is due to Simonyan and Zisserman (2015). Pre-trained weights were from the work of the MathConvNet team.
- Leon A. Gatys, Alexander S. Ecker, Matthias Bethge, (2015). [A Neural Algorithm of Artistic Style](https://arxiv.org/abs/1508.06576)
- Harish Narayanan, [Convolutional neural networks for artistic style transfer.](https://harishnarayanan.org/writing/artistic-style-transfer/)
- Log0, [TensorFlow Implementation of "A Neural Algorithm of Artistic Style".](http://www.chioka.in/tensorflow-implementation-neural-algorithm-of-artistic-style)
- Karen Simonyan and Andrew Zisserman (2015). [Very deep convolutional networks for large-scale image recognition](https://arxiv.org/pdf/1409.1556.pdf)
- [MatConvNet.](http://www.vlfeat.org/matconvnet/pretrained/)
| github_jupyter |
```
import numpy as np
np.random.seed(1337)
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Convolution2D, MaxPooling2D
from keras.utils import np_utils
from keras import backend as K
batch_size = 128
nb_classes = 10
nb_epoch = 5
# 输入图像的维度,此处是mnist图像,因此是28*28
img_rows, img_cols = 28, 28
# 卷积层中使用的卷积核的个数
nb_filters = 32
# 池化层操作的范围
pool_size = (2,2)
# 卷积核的大小
kernel_size = (3,3)
# keras中的mnist数据集已经被划分成了60,000个训练集,10,000个测试集的形式,按以下格式调用即可
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# 后端使用tensorflow时,即tf模式下,
# 会将100张RGB三通道的16*32彩色图表示为(100,16,32,3),
# 第一个维度是样本维,表示样本的数目,
# 第二和第三个维度是高和宽,
# 最后一个维度是通道维,表示颜色通道数
X_train = X_train.reshape(X_train.shape[0], img_rows, img_cols, 1)
X_test = X_test.reshape(X_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
# 将X_train, X_test的数据格式转为float32
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
# 归一化
X_train /= 255
X_test /= 255
# 打印出相关信息
print('X_train shape:', X_train.shape)
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
# 将类别向量(从0到nb_classes的整数向量)映射为二值类别矩阵,
# 相当于将向量用one-hot重新编码
Y_train = np_utils.to_categorical(y_train, nb_classes)
Y_test = np_utils.to_categorical(y_test, nb_classes)
# 输入图像的维度,此处是mnist图像,因此是28*28
img_rows, img_cols = 28, 28
# 卷积层中使用的卷积核的个数
nb_filters = 32
# 池化层操作的范围
pool_size = (2,2)
# 卷积核的大小
kernel_size = (3,3)
input_shape = (img_rows, img_cols,1)
# 建立序贯模型
model = Sequential()
# 卷积层,对二维输入进行滑动窗卷积
# 当使用该层为第一层时,应提供input_shape参数,在tf模式中,通道维位于第三个位置
# border_mode:边界模式,为"valid","same"或"full",即图像外的边缘点是补0
# 还是补成相同像素,或者是补1
cnn2d = Convolution2D(nb_filters, (kernel_size[0] ,kernel_size[1]),
padding='valid',
input_shape=input_shape)
model.add(cnn2d)
model.add(Activation('relu'))
# 卷积层,激活函数是ReLu
model.add(Convolution2D(nb_filters, (kernel_size[0], kernel_size[1])))
model.add(Activation('relu'))
# 池化层,选用Maxpooling,给定pool_size,dropout比例为0.25
model.add(MaxPooling2D(pool_size=pool_size))
model.add(Dropout(0.25))
# Flatten层,把多维输入进行一维化,常用在卷积层到全连接层的过渡
model.add(Flatten())
# 包含128个神经元的全连接层,激活函数为ReLu,dropout比例为0.5
model.add(Dense(256))
model.add(Activation('relu'))
model.add(Dropout(0.5))
# 包含10个神经元的输出层,激活函数为Softmax
model.add(Dense(nb_classes))
model.add(Activation('softmax'))
# 输出模型的参数信息
model.summary()
# 配置模型的学习过程
model.compile(loss='categorical_crossentropy',
optimizer='adadelta',
metrics=['accuracy'])
# 训练模型
history = model.fit(X_train, Y_train, batch_size=batch_size, epochs=nb_epoch,
verbose=1, validation_data=(X_test, Y_test))
# 按batch计算在某些输入数据上模型的误差
score = model.evaluate(X_test, Y_test, verbose=0)
# 输出训练好的模型在测试集上的表现
print('Test score:', score[0])
print('Test accuracy:', score[1])
score = model.evaluate(X_train, Y_train, verbose=0)
# 输出训练好的模型在测试集上的表现
print('X_train score:', score[0])
print('Test accuracy:', score[1])
import matplotlib.pyplot as plt
%matplotlib inline
print(cnn2d.get_weights()[0].shape)
from keras import models
import imageio
from keras.preprocessing import image
layer_outputs = [layer.output for layer in model.layers[:12]] # Extracts the outputs of the top 12 layers
activation_model = models.Model(inputs=model.input, outputs=layer_outputs) # Creates a model that will return these outputs, given the model input
# img_path = '../2019-02-21/my_own_images/9.png'
# img = image.load_img(img_path, target_size=(28, 28),color_mode="grayscale")
img = x_test[3].reshape(28,28)
plt.imshow(img)
# img_tensor = image.img_to_array(img)
img_tensor = img
img_tensor = img_tensor[:,:,np.newaxis]
img_tensor = np.expand_dims(img_tensor, axis=0)
# img_tensor = img_tensor[:,:,:,np.newaxis]
img_tensor /= 255.
print(img_tensor.shape)
# plt.imshow(img_tensor)
# plt.show()
activations = activation_model.predict(img_tensor) # Returns a list of five Numpy arrays: one array per layer activation
print(activations[1].shape)
first_layer_activation = activations[0]
print(first_layer_activation.shape)
plt.matshow(first_layer_activation[0, :, :, 4], cmap='viridis')
layer_names = []
for layer in model.layers[:12]:
layer_names.append(layer.name) # Names of the layers, so you can have them as part of your plot
images_per_row = 6
for layer_name, layer_activation in zip(layer_names, activations): # Displays the feature maps
n_features = layer_activation.shape[-1] # Number of features in the feature map
size = layer_activation.shape[1] #The feature map has shape (1, size, size, n_features).
n_cols = n_features // images_per_row # Tiles the activation channels in this matrix
display_grid = np.zeros((size * n_cols, images_per_row * size))
for col in range(n_cols): # Tiles each filter into a big horizontal grid
for row in range(images_per_row):
a = layer_activation[0,:, :, col * images_per_row + row]
# plt.imshow(layer_activation[0,:, :, col * images_per_row + row])
channel_image = a
channel_image -= channel_image.mean() # Post-processes the feature to make it visually palatable
channel_image /= channel_image.std()
channel_image *= 64
channel_image += 128
channel_image = np.clip(channel_image, 0, 255).astype('uint8')
display_grid[col * size : (col + 1) * size, # Displays the grid
row * size : (row + 1) * size] = channel_image
scale = 1. / size
plt.figure(figsize=(scale * display_grid.shape[1],
scale * display_grid.shape[0]))
plt.title(layer_name)
plt.grid(False)
plt.imshow(display_grid, aspect='auto', cmap='viridis')
m_test = []
m_label = []
for i in range(10):
img_path = '../2019-02-21/my_own_images/%s.png' % (i)
img = image.load_img(img_path, target_size=(28, 28),color_mode="grayscale")
img_tensor = image.img_to_array(img)
plt.imshow(img)
img_tensor = np.expand_dims(img_tensor, axis=0)
img_tensor /= 255.
# print(img_tensor.shape)
m_test.append(img_tensor)
label_arr = np.zeros(10)
label_arr[i] = 1
m_label.append(label_arr)
# plt.imshow(img_tensor.reshape((28,28)))
# plt.show()
# print(img_tensor.reshape((28,28)))
r = model.predict(img_tensor)
# print(r)
# plt.matshow(r)
plt.show()
print('实际是%s\t神经网络认为是 %s\t概率 %s' % (i,np.argmax(r),np.max(r)))
# r = np.expand_dims(r, axis=0)
print(r.shape)
plt.xticks(np.arange(0,10,1))
plt.yticks(np.arange(0,1,0.1))
# plt.matshow(r)
plt.bar(np.arange(0,10,1),r[0])
plt.show()
m_test = np.asarray(m_test)
m_label = np.asarray(m_label)
# print(Y_test[0])
# print(m_label[0])
# print(X_test.shape)
# print(m_test.shape)
# plt.imshow(X_test[1000].reshape(28,28))
score = model.evaluate(m_test, m_label, verbose=0)
print('m_test accuracy:', score[1],score[0])
```
| github_jupyter |
# CIFAR10 전이학습 기반 분류기
이 노트북은 사전 훈련된 심층-CNN 중에서 VGG16으로 전이학습의 개념을 확용한 분류기를 구축하는 단계를 개략적으로 설명한다.
```
%matplotlib inline
# Pandas and Numpy for data structures and util fucntions
import scipy as sp
import numpy as np
import pandas as pd
from numpy.random import rand
pd.options.display.max_colwidth = 600
# Scikit 임포트
from sklearn import preprocessing
from sklearn.metrics import roc_curve, auc, precision_recall_curve
from sklearn.model_selection import train_test_split
import cnn_utils as utils
# Matplot 임포트
import matplotlib.pyplot as plt
params = {'legend.fontsize': 'x-large',
'figure.figsize': (15, 5),
'axes.labelsize': 'x-large',
'axes.titlesize':'x-large',
'xtick.labelsize':'x-large',
'ytick.labelsize':'x-large'}
plt.rcParams.update(params)
# 판다스는 데이터 프레임을 테이블로 보여준다.
from IPython.display import display, HTML
import warnings
warnings.filterwarnings('ignore')
import tensorflow as tf
from tensorflow.keras import callbacks
from tensorflow.keras import optimizers
from tensorflow.keras.datasets import cifar10
from tensorflow.keras import Model
from tensorflow.keras.applications import vgg16 as vgg
from tensorflow.keras.layers import Dropout, Flatten, Dense, GlobalAveragePooling2D,BatchNormalization
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.python.keras.utils import np_utils
```
## 데이터 세트 로딩과 준비
```
BATCH_SIZE = 32
EPOCHS = 40
NUM_CLASSES = 10
LEARNING_RATE = 1e-4
MOMENTUM = 0.9
(X_train, y_train), (X_test, y_test) = cifar10.load_data()
```
Split training dataset in train and validation sets
```
X_train, X_val, y_train, y_val = train_test_split(X_train,
y_train,
test_size=0.15,
stratify=np.array(y_train),
random_state=42)
```
Transform target variable/labels into one hot encoded form
```
Y_train = np_utils.to_categorical(y_train, NUM_CLASSES)
Y_val = np_utils.to_categorical(y_val, NUM_CLASSES)
Y_test = np_utils.to_categorical(y_test, NUM_CLASSES)
```
### 전처리
VGG16을 특성 추출기로 사용할 것이기 때문에, 이미지의 최소 크기는 48x48이어야 한다. ```scipy```로 이미지 크기를 필요한 차원으로 재조정 한다.
```
X_train = np.array([sp.misc.imresize(x,
(48, 48)) for x in X_train])
X_val = np.array([sp.misc.imresize(x,
(48, 48)) for x in X_val])
X_test = np.array([sp.misc.imresize(x,
(48, 48)) for x in X_test])
```
## 모델 준비
* 최상위층 없이 VGG16 로딩
* 커스텀 분류기 준비
* 모델의 맨 위에 새로운 층 쌓기
```
base_model = vgg.VGG16(weights='imagenet',
include_top=False,
input_shape=(48, 48, 3))
```
목표는 분류층만 훈련시키는 것이기 때문에 훈련할 수 있는 파라미터 세팅을 False로 해서 나머지 층을 동결했다. 이렇게 하면 덜 강력한 기반 구조에서도 기존 아키텍처를 활용할 수 있고 학습된 가중치를 한 도메인에서 다른 도메인으로 전이할 수 있다.
```
# VGG16 모델의 세 번째 블록에서 마지막 층 추출
last = base_model.get_layer('block3_pool').output
# 상위 층에 분류층 추가
x = GlobalAveragePooling2D()(last)
x= BatchNormalization()(x)
x = Dense(256, activation='relu')(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.6)(x)
pred = Dense(NUM_CLASSES, activation='softmax')(x)
model = Model(base_model.input, pred)
```
우리의 목표는 커스컴 분류기를 훈련시키는 것이기 때문에 VGG16 층은 동결한다.
```
for layer in base_model.layers:
layer.trainable = False
model.compile(loss='binary_crossentropy',
optimizer=optimizers.Adam(lr=LEARNING_RATE),
metrics=['accuracy'])
model.summary()
```
## 데이터 늘리기
소규모 데이터 세트의 한계를 극복하고 모델을 일반화할 수 있도록 ```케라스``` 유틸리티로 데이터 세트를 늘려준다.
```
# 데이터 늘리기 구성의 준비
train_datagen = ImageDataGenerator(
rescale=1. / 255,
horizontal_flip=False)
train_datagen.fit(X_train)
train_generator = train_datagen.flow(X_train,
Y_train,
batch_size=BATCH_SIZE)
val_datagen = ImageDataGenerator(rescale=1. / 255,
horizontal_flip=False)
val_datagen.fit(X_val)
val_generator = val_datagen.flow(X_val,
Y_val,
batch_size=BATCH_SIZE)
```
## 모델 훈련
이제 모델을 몇 번의 에포크로 훈련시키고 그 성능을 측정해 보자. 다음 코드로 모델에 새로 추가된 층을 훈련시키기 위한 fit_generator() 함수를 호출한다.
```
train_steps_per_epoch = X_train.shape[0] // BATCH_SIZE
val_steps_per_epoch = X_val.shape[0] // BATCH_SIZE
history = model.fit_generator(train_generator,
steps_per_epoch=train_steps_per_epoch,
validation_data=val_generator,
validation_steps=val_steps_per_epoch,
epochs=EPOCHS,
verbose=1)
```
## 모델 성능 분석
```
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(15, 5))
t = f.suptitle('Deep Neural Net Performance', fontsize=12)
f.subplots_adjust(top=0.85, wspace=0.3)
epochs = list(range(1,EPOCHS+1))
ax1.plot(epochs, history.history['acc'], label='Train Accuracy')
ax1.plot(epochs, history.history['val_acc'], label='Validation Accuracy')
ax1.set_xticks(epochs)
ax1.set_ylabel('Accuracy Value')
ax1.set_xlabel('Epoch')
ax1.set_title('Accuracy')
l1 = ax1.legend(loc="best")
ax2.plot(epochs, history.history['loss'], label='Train Loss')
ax2.plot(epochs, history.history['val_loss'], label='Validation Loss')
ax2.set_xticks(epochs)
ax2.set_ylabel('Loss Value')
ax2.set_xlabel('Epoch')
ax2.set_title('Loss')
l2 = ax2.legend(loc="best")
predictions = model.predict(X_test/255.)
test_labels = list(y_test.squeeze())
predictions = list(predictions.argmax(axis=1))
get_metrics(true_labels=y_test,
predicted_labels=predictions)
```
## 예측 시각화
```
label_dict = {0:'airplane',
1:'automobile',
2:'bird',
3:'cat',
4:'deer',
5:'dog',
6:'frog',
7:'horse',
8:'ship',
9:'truck'}
utils.plot_predictions(model=model,dataset=X_test/255.,
dataset_labels=Y_test,
label_dict=label_dict,
batch_size=16,
grid_height=4,
grid_width=4)
```
| github_jupyter |
# Table of Contents
<p><div class="lev1 toc-item"><a href="#Setting-up-your-machine-Learning-Application" data-toc-modified-id="Setting-up-your-machine-Learning-Application-1"><span class="toc-item-num">1 </span>Setting up your machine Learning Application</a></div><div class="lev2 toc-item"><a href="#Train-/-Dev-/-Test-sets" data-toc-modified-id="Train-/-Dev-/-Test-sets-11"><span class="toc-item-num">1.1 </span>Train / Dev / Test sets</a></div><div class="lev2 toc-item"><a href="#Bias-/-Variance" data-toc-modified-id="Bias-/-Variance-12"><span class="toc-item-num">1.2 </span>Bias / Variance</a></div><div class="lev2 toc-item"><a href="#Basic-Recipe-for-Machine-Learning" data-toc-modified-id="Basic-Recipe-for-Machine-Learning-13"><span class="toc-item-num">1.3 </span>Basic Recipe for Machine Learning</a></div><div class="lev1 toc-item"><a href="#Regularizing-your-neural-network" data-toc-modified-id="Regularizing-your-neural-network-2"><span class="toc-item-num">2 </span>Regularizing your neural network</a></div><div class="lev2 toc-item"><a href="#Regularization" data-toc-modified-id="Regularization-21"><span class="toc-item-num">2.1 </span>Regularization</a></div><div class="lev2 toc-item"><a href="#Why-regularization-reduces-overfitting?" data-toc-modified-id="Why-regularization-reduces-overfitting?-22"><span class="toc-item-num">2.2 </span>Why regularization reduces overfitting?</a></div><div class="lev2 toc-item"><a href="#Dropout-Regularization" data-toc-modified-id="Dropout-Regularization-23"><span class="toc-item-num">2.3 </span>Dropout Regularization</a></div><div class="lev2 toc-item"><a href="#Understanding-Dropout" data-toc-modified-id="Understanding-Dropout-24"><span class="toc-item-num">2.4 </span>Understanding Dropout</a></div><div class="lev2 toc-item"><a href="#Other-regularization-methods" data-toc-modified-id="Other-regularization-methods-25"><span class="toc-item-num">2.5 </span>Other regularization methods</a></div><div class="lev1 toc-item"><a href="#Setting-up-your-optimization-problem" data-toc-modified-id="Setting-up-your-optimization-problem-3"><span class="toc-item-num">3 </span>Setting up your optimization problem</a></div><div class="lev2 toc-item"><a href="#Normalizing-inputs" data-toc-modified-id="Normalizing-inputs-31"><span class="toc-item-num">3.1 </span>Normalizing inputs</a></div><div class="lev2 toc-item"><a href="#Vanishing-/-Exploding-gradients" data-toc-modified-id="Vanishing-/-Exploding-gradients-32"><span class="toc-item-num">3.2 </span>Vanishing / Exploding gradients</a></div><div class="lev2 toc-item"><a href="#Weight-Initialization-for-Deep-Networks" data-toc-modified-id="Weight-Initialization-for-Deep-Networks-33"><span class="toc-item-num">3.3 </span>Weight Initialization for Deep Networks</a></div><div class="lev2 toc-item"><a href="#Numerical-approximation-of-gradients" data-toc-modified-id="Numerical-approximation-of-gradients-34"><span class="toc-item-num">3.4 </span>Numerical approximation of gradients</a></div><div class="lev2 toc-item"><a href="#Gradient-Checking" data-toc-modified-id="Gradient-Checking-35"><span class="toc-item-num">3.5 </span>Gradient Checking</a></div><div class="lev2 toc-item"><a href="#Gradient-Checking-Implementation-Notes" data-toc-modified-id="Gradient-Checking-Implementation-Notes-36"><span class="toc-item-num">3.6 </span>Gradient Checking Implementation Notes</a></div>
# Setting up your machine Learning Application
## Train / Dev / Test sets



## Bias / Variance

## Basic Recipe for Machine Learning

# Regularizing your neural network
## Regularization


## Why regularization reduces overfitting?


## Dropout Regularization



## Understanding Dropout

## Other regularization methods


# Setting up your optimization problem
## Normalizing inputs


## Vanishing / Exploding gradients

## Weight Initialization for Deep Networks

## Numerical approximation of gradients

## Gradient Checking


## Gradient Checking Implementation Notes

| github_jupyter |
# Self-Driving Car Engineer Nanodegree
## Deep Learning
## Project: Build a Traffic Sign Recognition Classifier
In this notebook, a template is provided for you to implement your functionality in stages, which is required to successfully complete this project. If additional code is required that cannot be included in the notebook, be sure that the Python code is successfully imported and included in your submission if necessary.
> **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the iPython Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to \n",
"**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
In addition to implementing code, there is a writeup to complete. The writeup should be completed in a separate file, which can be either a markdown file or a pdf document. There is a [write up template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) that can be used to guide the writing process. Completing the code template and writeup template will cover all of the [rubric points](https://review.udacity.com/#!/rubrics/481/view) for this project.
The [rubric](https://review.udacity.com/#!/rubrics/481/view) contains "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. The stand out suggestions are optional. If you decide to pursue the "stand out suggestions", you can include the code in this Ipython notebook and also discuss the results in the writeup file.
>**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
---
## Step 0: Load The Data
```
# Load pickled data
import pickle
# TODO: Fill this in based on where you saved the training and testing data
training_file = r'../data/train.p'
validation_file = r'../data/valid.p'
testing_file = r'../data/test.p'
with open(training_file, mode='rb') as f:
train = pickle.load(f)
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
```
---
## Step 1: Dataset Summary & Exploration
The pickled data is a dictionary with 4 key/value pairs:
- `'features'` is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
- `'labels'` is a 1D array containing the label/class id of the traffic sign. The file `signnames.csv` contains id -> name mappings for each id.
- `'sizes'` is a list containing tuples, (width, height) representing the original width and height the image.
- `'coords'` is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image. **THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES**
Complete the basic data summary below. Use python, numpy and/or pandas methods to calculate the data summary rather than hard coding the results. For example, the [pandas shape method](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.shape.html) might be useful for calculating some of the summary results.
### Provide a Basic Summary of the Data Set Using Python, Numpy and/or Pandas
```
### Replace each question mark with the appropriate value.
### Use python, pandas or numpy methods rather than hard coding the results
import numpy as np
import pandas as pd
# TODO: Number of training examples
n_train = train['features'].shape[0]
# TODO: Number of validation examples
n_validation = valid['features'].shape[0]
# TODO: Number of testing examples.
n_test = test['features'].shape[0]
# TODO: What's the shape of an traffic sign image?
image_shape = train['features'].shape[1:3]
# TODO: How many unique classes/labels there are in the dataset.
classes = pd.read_csv('signnames.csv', index_col=0)
n_classes = classes.shape[0]
print("Number of training examples =", n_train)
print("Number of validation examples =", n_validation)
print("Number of testing examples =", n_test)
print("Image data shape =", image_shape)
print("Number of classes =", n_classes)
```
### Include an exploratory visualization of the dataset
Visualize the German Traffic Signs Dataset using the pickled file(s). This is open ended, suggestions include: plotting traffic sign images, plotting the count of each sign, etc.
The [Matplotlib](http://matplotlib.org/) [examples](http://matplotlib.org/examples/index.html) and [gallery](http://matplotlib.org/gallery.html) pages are a great resource for doing visualizations in Python.
**NOTE:** It's recommended you start with something simple first. If you wish to do more, come back to it after you've completed the rest of the sections. It can be interesting to look at the distribution of classes in the training, validation and test set. Is the distribution the same? Are there more examples of some classes than others?
```
### Data exploration visualization code goes here.
### Feel free to use as many code cells as needed.
import matplotlib.pyplot as plt
# Visualizations will be shown in the notebook.
%matplotlib inline
# Choose an example from the training set
classid = classes.loc[classes['SignName'] == 'Roundabout mandatory'].index.values[0]
example = X_train[y_train == classid][0]
# Print a summary of the image
print('Class of the training example = ', classid)
plt.imshow(example)
def visualize_dataset(dataset, name, color):
"""
Helper function to visualize datasets
"""
# Create a figure
plt.figure(figsize=(18, 6))
# Plot histogram of the dataset
counts, _, _ = plt.hist(dataset, bins=n_classes, range=(0, n_classes), color=color, edgecolor='white')
# Show histogram information
plt.title('{} Set: {} Examples'.format(name, int(counts.sum())))
plt.xlabel('Traffic Sign Class')
plt.ylabel('Number of Examples')
visualize_dataset(y_train, 'Training', 'xkcd:salmon')
visualize_dataset(y_valid, 'Validation', 'xkcd:turquoise')
visualize_dataset(y_test, 'Testing', 'xkcd:lavender')
```
----
## Step 2: Design and Test a Model Architecture
Design and implement a deep learning model that learns to recognize traffic signs. Train and test your model on the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset).
The LeNet-5 implementation shown in the [classroom](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) at the end of the CNN lesson is a solid starting point. You'll have to change the number of classes and possibly the preprocessing, but aside from that it's plug and play!
With the LeNet-5 solution from the lecture, you should expect a validation set accuracy of about 0.89. To meet specifications, the validation set accuracy will need to be at least 0.93. It is possible to get an even higher accuracy, but 0.93 is the minimum for a successful project submission.
There are various aspects to consider when thinking about this problem:
- Neural network architecture (is the network over or underfitting?)
- Play around preprocessing techniques (normalization, rgb to grayscale, etc)
- Number of examples per label (some have more than others).
- Generate fake data.
Here is an example of a [published baseline model on this problem](http://yann.lecun.com/exdb/publis/pdf/sermanet-ijcnn-11.pdf). It's not required to be familiar with the approach used in the paper but, it's good practice to try to read papers like these.
### Pre-process the Data Set (normalization, grayscale, etc.)
Minimally, the image data should be normalized so that the data has mean zero and equal variance. For image data, `(pixel - 128)/ 128` is a quick way to approximately normalize the data and can be used in this project.
Other pre-processing steps are optional. You can try different techniques to see if it improves performance.
Use the code cell (or multiple code cells, if necessary) to implement the first step of your project.
```
### Preprocess the data here. It is required to normalize the data. Other preprocessing steps could include
### converting to grayscale, etc.
### Feel free to use as many code cells as needed.
import cv2
def normalize(data):
return (data.astype(float) - 128) / 128
def preprocess(data):
dim = list(data.shape)
dim[-1] = 1
X = np.zeros(tuple(dim))
for i in range(data.shape[0]):
grayscale = cv2.cvtColor(data[i], cv2.COLOR_RGB2GRAY)
equalized = cv2.equalizeHist(grayscale)
X[i] = np.expand_dims(equalized, axis=2)
return normalize(X)
# Apply pre-processing to the datasets
X_train = preprocess(X_train)
X_valid = preprocess(X_valid)
X_test = preprocess(X_test)
# Save the number of channels after pre-processing
channels = X_train.shape[-1]
```
### Setup TensorFlow
```
import tensorflow as tf
EPOCHS = 10
BATCH_SIZE = 128
```
### Model Architecture
```
### Define your architecture here.
### Feel free to use as many code cells as needed.
from tensorflow.contrib.layers import flatten
def LeNet(x):
# Arguments used for tf.truncated_normal, randomly defines variables for the weights and biases for each layer
mu = 0
sigma = 0.1
# SOLUTION: Layer 1: Convolutional. Input = 32x32xC. Output = 28x28x6.
conv1_W = tf.Variable(tf.truncated_normal(shape=(5, 5, channels, 6), mean = mu, stddev = sigma))
conv1_b = tf.Variable(tf.zeros(6))
conv1 = tf.nn.conv2d(x, conv1_W, strides=[1, 1, 1, 1], padding='VALID') + conv1_b
# SOLUTION: Activation.
conv1 = tf.nn.relu(conv1)
# SOLUTION: Pooling. Input = 28x28x6. Output = 14x14x6.
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# Drop-out.
drop1 = tf.nn.dropout(conv1, keep_prob=keep_prob)
# SOLUTION: Layer 2: Convolutional. Output = 10x10x16.
conv2_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 16), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(16))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides=[1, 1, 1, 1], padding='VALID') + conv2_b
# SOLUTION: Activation.
conv2 = tf.nn.relu(conv2)
# SOLUTION: Pooling. Input = 10x10x16. Output = 5x5x16.
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# Drop-out.
drop2 = tf.nn.dropout(conv2, keep_prob=keep_prob)
# SOLUTION: Flatten. Input = 5x5x16. Output = 400.
fc0 = flatten(drop2)
# SOLUTION: Layer 3: Fully Connected. Input = 400. Output = 120.
fc1_W = tf.Variable(tf.truncated_normal(shape=(400, 120), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(120))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# SOLUTION: Activation.
fc1 = tf.nn.relu(fc1)
# SOLUTION: Layer 4: Fully Connected. Input = 120. Output = 84.
fc2_W = tf.Variable(tf.truncated_normal(shape=(120, 84), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(84))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# SOLUTION: Activation.
fc2 = tf.nn.relu(fc2)
# SOLUTION: Layer 5: Fully Connected. Input = 84. Output = n_classes.
fc3_W = tf.Variable(tf.truncated_normal(shape=(84, n_classes), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(n_classes))
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits
```
### Features and Labels
```
x = tf.placeholder(tf.float32, (None, 32, 32, channels))
y = tf.placeholder(tf.int32, (None))
keep_prob = tf.placeholder(tf.float32)
one_hot_y = tf.one_hot(y, n_classes)
```
### Train, Validate and Test the Model
A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation
sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
### Training Pipeline
```
rate = 0.001
logits = LeNet(x)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=one_hot_y, logits=logits)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = optimizer.minimize(loss_operation)
```
### Model Evaluation
```
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
saver = tf.train.Saver()
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE]
accuracy = sess.run(accuracy_operation, feed_dict={x: batch_x, y: batch_y, keep_prob: 1.0})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
```
### Train the Model
```
### Train your model here.
### Calculate and report the accuracy on the training and validation set.
### Once a final model architecture is selected,
### the accuracy on the test set should be calculated and reported as well.
### Feel free to use as many code cells as needed.
from sklearn.utils import shuffle
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print("Training...")
print()
for i in range(EPOCHS):
X_train, y_train = shuffle(X_train, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y, keep_prob: 0.5})
training_accuracy = evaluate(X_train, y_train)
validation_accuracy = evaluate(X_valid, y_valid)
print("EPOCH {} ...".format(i+1))
print("Training Accuracy = {:.3f} - Validation Accuracy = {:.3f}".format(training_accuracy, validation_accuracy))
print()
saver.save(sess, './lenet')
print("Model saved")
```
### Evaluate the Model
```
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy = evaluate(X_test, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy))
```
---
## Step 3: Test a Model on New Images
To give yourself more insight into how your model is working, download at least five pictures of German traffic signs from the web and use your model to predict the traffic sign type.
You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
### Load and Output the Images
```
### Load the images and plot them here.
### Feel free to use as many code cells as needed.
import os
def load(path):
# Create a list of file names in the current path
fnames = os.listdir(path)
# Create variables to store the results
m = len(fnames)
X = np.zeros((m, 32, 32, 3), dtype=np.uint8)
y = np.zeros(m, dtype=int)
for i, file in enumerate(fnames):
X[i] = plt.imread(os.path.join(path, file))
y[i] = int(os.path.splitext(file)[0])
return X, y
# Load new images
X_new, y_new = load('images')
# Plot a new image
plt.imshow(X_new[0])
```
### Predict the Sign Type for Each Image
```
### Run the predictions here and use the model to output the prediction for each image.
### Make sure to pre-process the images with the same pre-processing pipeline used earlier.
### Feel free to use as many code cells as needed.
X_new = preprocess(X_new)
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
prediction = np.argmax(sess.run(logits, feed_dict={x: X_new, keep_prob: 1.0}), axis=1)
for i, p in enumerate(prediction):
print('Image {} - Target = {:02d}, Predicted = {:02d}'.format(i, y_new[i], p))
print('> Model accuracy: {:02f}'.format(np.sum(y_new == prediction)/y_new.shape[0]))
prediction
```
### Analyze Performance
```
### Calculate the accuracy for these 5 new images.
### For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate on these new images.
```
### Output Top 5 Softmax Probabilities For Each Image Found on the Web
For each of the new images, print out the model's softmax probabilities to show the **certainty** of the model's predictions (limit the output to the top 5 probabilities for each image). [`tf.nn.top_k`](https://www.tensorflow.org/versions/r0.12/api_docs/python/nn.html#top_k) could prove helpful here.
The example below demonstrates how tf.nn.top_k can be used to find the top k predictions for each image.
`tf.nn.top_k` will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
Take this numpy array as an example. The values in the array represent predictions. The array contains softmax probabilities for five candidate images with six possible classes. `tf.nn.top_k` is used to choose the three classes with the highest probability:
```
# (5, 6) array
a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
0.12789202],
[ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
0.15899337],
[ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
0.23892179],
[ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
0.16505091],
[ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
0.09155967]])
```
Running it through `sess.run(tf.nn.top_k(tf.constant(a), k=3))` produces:
```
TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
[ 0.28086119, 0.27569815, 0.18063401],
[ 0.26076848, 0.23892179, 0.23664738],
[ 0.29198961, 0.26234032, 0.16505091],
[ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
[0, 1, 4],
[0, 5, 1],
[1, 3, 5],
[1, 4, 3]], dtype=int32))
```
Looking just at the first row we get `[ 0.34763842, 0.24879643, 0.12789202]`, you can confirm these are the 3 largest probabilities in `a`. You'll also notice `[3, 0, 5]` are the corresponding indices.
```
### Print out the top five softmax probabilities for the predictions on the German traffic sign images found on the web.
### Feel free to use as many code cells as needed.
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
top = sess.run(tf.nn.top_k(tf.nn.softmax(logits), k=5), feed_dict={x: X_new, keep_prob: 1.0})
top
def format(predictions):
n = predictions.values.shape[0]
probs = np.zeros((n, n_classes))
for i in range(n):
for j in range(predictions.values.shape[1]):
probs[i, predictions.indices[i, j]] = predictions.values[i, j]
return probs
def batch_compare(probs):
n = probs.shape[0]
f, ax = plt.subplots(n, 2, figsize=(18, 6*n))
images, _ = load('images')
for i in range(n):
ax[i][0].imshow(images[i])
ax[i][1].bar(np.arange(n_classes), probs[i], width=2.5, color='xkcd:sky blue', edgecolor='white')
batch_compare(format(top))
```
### Project Writeup
Once you have completed the code implementation, document your results in a project writeup using this [template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) as a guide. The writeup can be in a markdown or pdf file.
> **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
"**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
---
## Step 4 (Optional): Visualize the Neural Network's State with Test Images
This Section is not required to complete but acts as an additional excersise for understaning the output of a neural network's weights. While neural networks can be a great learning device they are often referred to as a black box. We can understand what the weights of a neural network look like better by plotting their feature maps. After successfully training your neural network you can see what it's feature maps look like by plotting the output of the network's weight layers in response to a test stimuli image. From these plotted feature maps, it's possible to see what characteristics of an image the network finds interesting. For a sign, maybe the inner network feature maps react with high activation to the sign's boundary outline or to the contrast in the sign's painted symbol.
Provided for you below is the function code that allows you to get the visualization output of any tensorflow weight layer you want. The inputs to the function should be a stimuli image, one used during training or a new one you provided, and then the tensorflow variable name that represents the layer's state during the training process, for instance if you wanted to see what the [LeNet lab's](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) feature maps looked like for it's second convolutional layer you could enter conv2 as the tf_activation variable.
For an example of what feature map outputs look like, check out NVIDIA's results in their paper [End-to-End Deep Learning for Self-Driving Cars](https://devblogs.nvidia.com/parallelforall/deep-learning-self-driving-cars/) in the section Visualization of internal CNN State. NVIDIA was able to show that their network's inner weights had high activations to road boundary lines by comparing feature maps from an image with a clear path to one without. Try experimenting with a similar test to show that your trained network's weights are looking for interesting features, whether it's looking at differences in feature maps from images with or without a sign, or even what feature maps look like in a trained network vs a completely untrained one on the same sign image.
<figure>
<img src="visualize_cnn.png" width="380" alt="Combined Image" />
<figcaption>
<p></p>
<p style="text-align: center;"> Your output should look something like this (above)</p>
</figcaption>
</figure>
<p></p>
```
### Visualize your network's feature maps here.
### Feel free to use as many code cells as needed.
# image_input: the test image being fed into the network to produce the feature maps
# tf_activation: should be a tf variable name used during your training procedure that represents the calculated state of a specific weight layer
# activation_min/max: can be used to view the activation contrast in more detail, by default matplot sets min and max to the actual min and max values of the output
# plt_num: used to plot out multiple different weight feature map sets on the same block, just extend the plt number for each new feature map entry
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it may be having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
```
| github_jupyter |
[source](../api/alibi_detect.ad.model_distillation.rst)
# Model distillation
## Overview
[Model distillation](https://arxiv.org/abs/1503.02531) is a technique that is used to transfer knowledge from a large network to a smaller network. Typically, it consists of training a second model with a simplified architecture on soft targets (the output distributions or the logits) obtained from the original model.
Here, we apply model distillation to obtain harmfulness scores, by comparing the output distributions of the original model with the output distributions
of the distilled model, in order to detect adversarial data, malicious data drift or data corruption.
We use the following definition of harmful and harmless data points:
* Harmful data points are defined as inputs for which the model's predictions on the uncorrupted data are correct while the model's predictions on the corrupted data are wrong.
* Harmless data points are defined as inputs for which the model's predictions on the uncorrupted data are correct and the model's predictions on the corrupted data remain correct.
Analogously to the [adversarial AE detector](https://arxiv.org/abs/2002.09364), which is also part of the library, the model distillation detector picks up drift that reduces the performance of the classification model.
The detector can be used as follows:
* Given an input $x,$ an adversarial score $S(x)$ is computed. $S(x)$ equals the value loss function employed for distillation calculated between the original model's output and the distilled model's output on $x$.
* If $S(x)$ is above a threshold (explicitly defined or inferred from training data), the instance is flagged as adversarial.
## Usage
### Initialize
Parameters:
* `threshold`: threshold value above which the instance is flagged as an adversarial instance.
* `distilled_model`: `tf.keras.Sequential` instance containing the model used for distillation. Example:
```python
distilled_model = tf.keras.Sequential(
[
tf.keras.InputLayer(input_shape=(input_dim,)),
tf.keras.layers.Dense(output_dim, activation=tf.nn.softmax)
]
)
```
* `model`: the classifier as a `tf.keras.Model`. Example:
```python
inputs = tf.keras.Input(shape=(input_dim,))
hidden = tf.keras.layers.Dense(hidden_dim)(inputs)
outputs = tf.keras.layers.Dense(output_dim, activation=tf.nn.softmax)(hidden)
model = tf.keras.Model(inputs=inputs, outputs=outputs)
```
* `loss_type`: type of loss used for distillation. Supported losses: 'kld', 'xent'.
* `temperature`: Temperature used for model prediction scaling. Temperature <1 sharpens the prediction probability distribution which can be beneficial for prediction distributions with high entropy.
* `data_type`: can specify data type added to metadata. E.g. *'tabular'* or *'image'*.
Initialized detector example:
```python
from alibi_detect.ad import ModelDistillation
ad = ModelDistillation(
distilled_model=distilled_model,
model=model,
temperature=0.5
)
```
### Fit
We then need to train the detector. The following parameters can be specified:
* `X`: training batch as a numpy array.
* `loss_fn`: loss function used for training. Defaults to the custom model distillation loss.
* `optimizer`: optimizer used for training. Defaults to [Adam](https://arxiv.org/abs/1412.6980) with learning rate 1e-3.
* `epochs`: number of training epochs.
* `batch_size`: batch size used during training.
* `verbose`: boolean whether to print training progress.
* `log_metric`: additional metrics whose progress will be displayed if verbose equals True.
* `preprocess_fn`: optional data preprocessing function applied per batch during training.
```python
ad.fit(X_train, epochs=50)
```
The threshold for the adversarial / harmfulness score can be set via ```infer_threshold```. We need to pass a batch of instances $X$ and specify what percentage of those we consider to be normal via `threshold_perc`. Even if we only have normal instances in the batch, it might be best to set the threshold value a bit lower (e.g. $95$%) since the model could have misclassified training instances.
```python
ad.infer_threshold(X_train, threshold_perc=95, batch_size=64)
```
### Detect
We detect adversarial / harmful instances by simply calling `predict` on a batch of instances `X`. We can also return the instance level score by setting `return_instance_score` to True.
The prediction takes the form of a dictionary with `meta` and `data` keys. `meta` contains the detector's metadata while `data` is also a dictionary which contains the actual predictions stored in the following keys:
* `is_adversarial`: boolean whether instances are above the threshold and therefore adversarial instances. The array is of shape *(batch size,)*.
* `instance_score`: contains instance level scores if `return_instance_score` equals True.
```python
preds_detect = ad.predict(X, batch_size=64, return_instance_score=True)
```
## Examples
### Image
[Harmful drift detection through model distillation on CIFAR10](../examples/cd_distillation_cifar10.nblink)
| github_jupyter |
## 각 형태소 분석기 비교 (KKMA, KOMORAN, MECAB, TWITTER)
##### 긍정리뷰 3개 vs. 부정리뷰 3개를 기준으로
------------
#### 긍정리뷰
"3달정도 사용해오고 있는데 가성비부터 최고예요. 운동도 하고 교통비도 아끼고 대만족입니다."
"QR코드 이용해서 대여하니 빠르고 편해요. 처음 이용해봤는데 좋았네요^^"
"너무너무 좋아요. 결제도 쉽고 대여소 찾는 것도 쉽고 정기권 끊어서 타고있어요. 이거 덕분에 자전거 많이 타고있어요!! 감사합니다~"
#### 부정리뷰
"소셜로그인은 자동로그인이 안되나요? 너무 불편해요ㅠ"
"회원가입도 안되고 로그인도 잘 안되요."
"제발 앱좀 고쳐주세요..매번 카드정보입력하게할거면 뭐하러 회원정보수집하고 뭐하러 로그인시키나요.. 탈때마다 불편해죽겠어요. "
"매번 카드정보입력하게할거면 뭐하러 회원정보수집하고 뭐하러 로그인시키나요.. 탈때마다 불편해죽겠어요. "
```
# 각 형태소 분석기 import
from konlpy.tag import *
twitter = Okt()
mecab = Mecab()
han = Hannanum()
kkma = Kkma()
komo = Komoran()
b_text1 = '소셜로그인은 자동로그인이 안되나요? 너무 불편해요ㅠ'
b_text2 = "회원가입도 안되고 로그인도 잘 안되요."
b_text3 = "제발 앱좀 고쳐주세요..매번 카드정보입력하게할거면 뭐하러 회원정보수집하고 뭐하러 로그인시키나요.. 탈때마다 불편해죽겠어요. "
b_text_extra = "매번 카드정보입력하게할거면 뭐하러 회원정보수집하고 뭐하러 로그인시키나요.. 탈때마다 불편해죽겠어요. "
g_text1 = "3달정도 사용해오고 있는데 가성비부터 최고예요. 운동도 하고 교통비도 아끼고 대만족입니다."
g_text2 = "QR코드이용해서 대여하니빠르고 편해요 ㅎㅎㅎ. 처음 이용해봤는데 좋았네요^^"
g_text3 = "너무너무 좋아요. 결제도 쉽고 대여소 찾는 것도 쉽고 정기권 끊어서 타고있어요. 이거 덕분에 자전거 많이 타고있어요!! 감사합니다~"
def posComparsion(text):
kkma_text = kkma.pos(text)
han_text = han.pos(text)
komo_text = komo.pos(text)
mecab_text = mecab.pos(text)
twitter_text =twitter.pos(text)
return print('KKMA POS\n', kkma_text, '\n\nHannanum POS\n', han_text, '\n\nKomoran POS\n', komo_text, '\n\nMecab POS\n', mecab_text,
'\n\nTwitter POS\n' ,twitter_text)
def makeSlash(pos_text):
pos_slash = []
for compo in pos_text:
word =compo[0]+'/'
pos = compo[1]
words = (word, pos)
pos_slash.append(words)
return pos_slash
def posComparsionText(text):
kkma_text = kkma.pos(text)
han_text = han.pos(text)
komo_text = komo.pos(text)
mecab_text = mecab.pos(text)
twitter_text =twitter.pos(text, stem=True)
kkma_slash = makeSlash(kkma_text)
han_slash =makeSlash(han_text)
komo_slash = makeSlash(komo_text)
mecab_slash = makeSlash(mecab_text)
twitter_slash =makeSlash(twitter_text)
return print('KKMA POS\n', kkma_slash, '\n\nHannanum POS\n', han_slash, '\n\nKomoran POS\n', komo_slash, '\n\nMecab POS\n', mecab_slash,
'\n\nTwitter POS\n' ,twitter_slash)
```
### 긍정리뷰
#### "3달정도 사용해오고 있는데 가성비부터 최고예요. 운동도 하고 교통비도 아끼고 대만족입니다."
```
posComparsionText(g_text1)
```
</br>
#### "QR코드이용해서 대여하니빠르고 편해요 ㅎㅎㅎ. 처음 이용해봤는데 좋았네요^^"
```
posComparsionText(g_text2)
```
</br>
#### "너무너무 좋아요. 결제도 쉽고 대여소 찾는 것도 쉽고 정기권 끊어서 타고있어요. 이거 덕분에 자전거 많이 타고있어요!! 감사합니다~"
```
posComparsion(g_text3)
```
</br>
### 부정리뷰
#### "소셜로그인은 자동로그인이 안되나요? 너무 불편해요ㅠ"
```
posComparsion(b_text1)
```
</br>
#### "회원가입도 안되고 로그인도 잘 안되요."
```
posComparsion(b_text2)
```
</br>
#### "제발 앱좀 고쳐주세요..매번 카드정보입력하게할거면 뭐하러 회원정보수집하고 뭐하러 로그인시키나요.. 탈때마다 불편해죽겠어요. "
```
posComparsion(b_text3)
posComparsionText(b_text_extra)
twitter.pos("3달정도 사용해오고 있는데 가성비부터 최고예요. 운동도 하고 교통비도 아끼고 대만족입니다." , stem=True)
```
| github_jupyter |
# Trying to determine Customer Satisfaction based on numeric variables
**_By Isaiah Torain_**
## Summary/Introduction
Modern day businesses now rely on data to help their business. They need data help them market, data to help them run their business more efficiently, and data to help improve their products. The last topic is where I am going to focus. How to improve a product or service using data? One way is to gather customer feedback to determine if they are satisfied with your product/service and which factors best determine whether a customer is satisfied. That way you can determine which areas of the product and service to improve upon. That is the heart of the data set I am working with. Given certain factors, can I determine if a customer is satisfied with a service? A competition on Kaggle provides a dataset to try and determine if a customer is satisfied with a product. The data is presented in a csv format where there are 371 numerical variables representing different customer factors (these actual field variable names are hidden due to legal reasons). The target is simply category: 0 for customer sastification and 1 for a customer being unsatisfied. One of the difficulties with this data is that the data provided is skewed. About 96% of the samples are satisfied customers and the rest are unsatisfied instead of being a 50/50 split. I determined an optimal model for predicting customer satisfaction by comparing LDA, QDA, logistic, and Non-logistic regression. Feature engineering was also used to select only the variables that mattered most in predicting customer satisfaction. The result is that
## Methods
Let me start by visualizing the initial data to see whats going on.
```
import pandas
import numpy as np
import time
import matplotlib.pyplot as plt
import mlutils as ml
import neuralnetworks as nn
import qdalda as ql
import itertools
import imp
from sklearn.metrics import roc_auc_score, roc_curve, auc
from sklearn.feature_selection import SelectFromModel
imp.reload(nn)
imp.reload(ml)
%matplotlib inline
def trainLDA(X, T, parameters):
lda = ql.LDA()
lda.train(X,T)
return lda
def evaluateLDA(model, X, T):
pclass, probabilities, discriminants = model.use(X)
return np.sum(pclass.ravel()==T.ravel()) / float(len(T)) * 100
def trainQDA(X, T, parameters):
qda = ql.QDA()
qda.train(X, T)
return qda
def evaluateQDA(model, X, T):
pclass, probabilities, discriminants = model.use(X)
return np.sum(pclass.ravel()==T.ravel()) / float(len(T)) * 100
def trainNN(X, T, parameters):
classes = np.unique(T)
if parameters == 0 or parameters == [0] or parameters is None or parameters == [None]:
nnet = nn.NeuralNetworkClassifier(X.shape[1], None, classes.shape[0])
nnet.train(X, T)
elif type(parameters) is int:
nnet = nn.NeuralNetworkClassifier(X.shape[1], parameters, classes.shape[0])
nnet.train(X, T)
elif type(parameters) is list and len(parameters) == 1:
nnet = nn.NeuralNetworkClassifier(X.shape[1], parameters[0], classes.shape[0])
nnet.train(X, T)
else:
nnet = nn.NeuralNetworkClassifier(X.shape[1], parameters[0], classes.shape[0])
nnet.train(X, T, nIterations = parameters[1])
return nnet
def evaluateNN(model, X, T):
results = model.use(X)
return np.sum(results.ravel()==T.ravel()) / float(len(T)) * 100
def showResults(pclass, Targets, whichAlgo, probabilities=None):
Tclasses = np.array([0,1])
classes,counts = np.unique(pclass,return_counts=True)
print('classes',classes)
print('counts',counts)
ml.confusionMatrix(Targets,pclass,Tclasses)
print("ROC AUC: ", roc_auc_score(Targets, pclass))
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(2):
fpr[i], tpr[i], _ = roc_curve(Targets, pclass)
roc_auc[i] = auc(fpr[i], tpr[i])
fpr["micro"], tpr["micro"], _ = roc_curve(Targets.ravel(), pclass.ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
plt.figure()
plt.plot(fpr[1], tpr[1], label='ROC curve (area = %0.2f)' % roc_auc[1])
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic for %s' % whichAlgo)
plt.legend(loc="lower right")
plt.show()
def printResults(label,results):
print('{:4s} {:>20s}{:>8s}{:>8s}{:>8s}'.format('Algo','Parameters','TrnAcc','ValAcc','TesAcc'))
print('-------------------------------------------------')
for row in results:
# 20 is expected maximum number of characters in printed parameter value list
print('{:>4s} {:>20s} {:7.2f} {:7.2f} {:7.2f}'.format(label,str(row[0]),*row[1:]))
data = pandas.read_csv("train.csv")
data.head()
targets = pandas.DataFrame(data.TARGET.value_counts())
targets['Percentage'] = 100*targets['TARGET']/data.shape[0]
targets
```
It's clear that the unsatisfied customer are the outliers in this case but which factors are the best predictor of these outliers is the question. There are many ways to do classification, one of the first is removing categorical variables.
I want to make sure the data is cleaned before I proceed with analyzing it. To do this I will find out which variables are duplicates and which rows are duplicates.
```
def removeDuplicates(data):
data.drop_duplicates()
remove = []
columns = data.columns
for i in range(len(columns) - 1):
values = data[columns[i]].values
for j in range(i+1,len(columns)):
if np.array_equal(values, data[columns[j]].values):
remove.append(columns[j])
data.drop(remove, axis=1, inplace=True)
```
I also need to remove columns with the same constant values
```
def removeConstantColumns(data):
remove = []
for col in data.columns:
if data[col].std() == 0:
remove.append(col)
data.drop(remove, axis=1, inplace=True)
removeDuplicates(data)
removeConstantColumns(data)
data.describe()
```
According to this link () column var3 is supposed to be the nationality of the customer. -999999 represents the nationality or country of origin not being known. Thus it is a categorical variable. I will take -9999999 as being 2. The ID column also needs to be dropped.
```
data.var3.replace(-999999, 2)
data.drop(["ID"], axis =1)
```
Now I will try out my various models to see how they do.
```
#test_data = pandas.read_csv("test.csv")
#removeDuplicates(test_data)
#removeConstantColumns(test_data)
#test_data.var3.replace(-999999, 2)
#test_data.drop(["ID"], axis = 1)
#test_data = test_data.as_matrix()
train_data = data.as_matrix()
Targets = train_data[:,-1]
Targets = Targets.reshape((-1,1))
train_data = np.delete(train_data, -1, 1)
print (train_data.shape)
#print (test_data.shape)
lda = ql.LDA()
lda.train(train_data, Targets)
pclass, probabilities, discriminants = lda.use(train_data)
showResults(pclass,Targets, "LDA")
qda = ql.QDA()
qda.train(train_data, Targets)
pc, prob, d = qda.use(train_data)
showResults(pc, Targets, "QDA")
nnet = nn.NeuralNetworkClassifier(train_data.shape[1], None, Tclasses.shape[0])
nnet.train(train_data, Targets)
results = nnet.use(train_data)
print (np.sum(results.ravel()==Targets.ravel()) / float(len(Targets)) * 100)
showResults(results, Targets, "Logistic Regresssion")
nnet = nn.NeuralNetworkClassifier(train_data.shape[1], [10, 30], Tclasses.shape[0])
nnet.train(train_data, Targets)
results = nnet.use(train_data)
print (np.sum(results.ravel()==Targets.ravel()) / float(len(Targets)) * 100)
showResults(results, Targets, "NL Logistic Regresssion")
resultsLDA = ml.trainValidateTestKFoldsClassification(trainLDA, evaluateLDA, train_data, Targets, [None], nFolds=6, shuffle=True, verbose=False)
printResults('LDA: ', resultsLDA)
resultsQDA = ml.trainValidateTestKFoldsClassification(trainQDA, evaluateQDA, train_data, Targets, [None], nFolds=6, shuffle=True, verbose=False)
printResults('QDA: ', resultsQDA)
#parameters for neural net is [[0], 10] where the first input is the number of hidden units and the second is iterations
resultsNN = ml.trainValidateTestKFoldsClassification(trainNN, evaluateNN, train_data, Targets, [None], nFolds=6, shuffle=True, verbose=False)
printResults('NN: ', resultsNN)
resultsNN = ml.trainValidateTestKFoldsClassification(trainNN, evaluateNN, train_data, Targets, [[[5], 10]], nFolds=10, shuffle=True, verbose=False)
printResults('NN: ', resultsNN)
resultsNN = ml.trainValidateTestKFoldsClassification(trainNN, evaluateNN, train_data, Targets, [[[50, 50], 5]], nFolds=10, shuffle=True, verbose=False)
printResults('NN: ', resultsNN)
```
Seems as though the neural network could do better but that could also be the result of the data being skewed and it doing really well on the targets of 0. Next I will remove features to try to get a better model.
```
from sklearn.ensemble import ExtraTreesClassifier
clf = ExtraTreesClassifier(random_state=1729)
data = data.drop(["TARGET"], axis =1)
selector = clf.fit(data, Targets)
# plot most important features
feat_imp = pandas.Series(clf.feature_importances_, index = data.columns.values).sort_values(ascending=False)
feat_imp[:40].plot(kind='bar', title='Feature Importances according to ExtraTreesClassifier', figsize=(12, 8))
plt.ylabel('Feature Importance Score')
plt.subplots_adjust(bottom=0.3)
plt.savefig('1.png')
plt.show()
# clf.feature_importances_
fs = SelectFromModel(selector, prefit=True)
newTrain = fs.transform(data)
print (newTrain.shape)
lda = ql.LDA()
lda.train(newTrain, Targets)
pclass, probabilities, discriminants = lda.use(newTrain)
showResults(pclass,Targets, "LDA")
qda = ql.QDA()
qda.train(newTrain, Targets)
pclass, probabilities, discriminants = qda.use(newTrain)
showResults(pclass,Targets, "QDA")
nnet = nn.NeuralNetworkClassifier(newTrain.shape[1], None, Tclasses.shape[0])
nnet.train(newTrain, Targets)
results = nnet.use(newTrain)
print (np.sum(results.ravel()==Targets.ravel()) / float(len(Targets)) * 100)
showResults(results, Targets, "Logistic Regresssion")
nnet = nn.NeuralNetworkClassifier(newTrain.shape[1], [5, 10, 15, 20, 15, 10, 5], Tclasses.shape[0])
nnet.train(newTrain, Targets, nIterations=100)
results = nnet.use(newTrain)
print (np.sum(results.ravel()==Targets.ravel()) / float(len(Targets)) * 100)
showResults(results, Targets, "NL Logistic Regresssion")
import xgboost
boosted = xgboost.XGBClassifier(n_estimators=110, nthread=-1, max_depth=4, seed=1729)
boosted.fit(newTrain, Targets, eval_metric="auc", verbose=False)
print ("AUC: ", roc_auc_score(Targets, boosted.predict_proba(newTrain), average='macro'))
```
## Results
It seems I could achieve no better than an AUC of about .70 in all tests. That is not bad however some of the Kaggle competitors were able to achieve about .84 using a tool called xgboost (Extreme Gradient Boosting).
## Conclusion
In conclusion, using several different models it seems the best way to predict an unbalanced dataset with a lot of attributes is using Linear discriminant analysis. One difficulty I ran into was calculating the ROC but once I found out about the sklearn toolkit it was easy.
| github_jupyter |
<a href="https://colab.research.google.com/github/liscolme/EscapeEarth/blob/main/Interns/Elise/BLS_Function_Test.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from google.colab import drive
drive.mount('/content/gdrive')
###################################
!pip install lightkurve
import lightkurve as lk
import numpy as np
import pandas as pd
import astropy.units as u
from astropy.timeseries import BoxLeastSquares
!ls /content/gdrive/MyDrive/EscapeEarthData
#/content/gdrive/My \Drive/EscapeEarthData/Sec14_cleaned/{ticid}/lc.fits
#/content/gdrive/My \Drive/EscapeEarthData/Sec15_cleaned/{ticid}/lc.fits
#/content/gdrive/My \Drive/EscapeEarthData/all_targets_S014_v1.csv
#timestamps
t = np.random.uniform(0, 20, 2000)
#observations
y = np.ones_like(t) - 0.1*((t%3)<0.2) + 0.01*np.random.randn(len(t))
model = BoxLeastSquares(t * u.day, y, dy=0.01)
periodogram = model.autopower(0.2)
import matplotlib.pyplot as plt
plt.plot(periodogram.period, periodogram.power)
model = BoxLeastSquares(t * u.day, y, dy=0.01)
periodogram = model.autopower(0.2, objective="snr")
plt.plot(periodogram.period, periodogram.power)
# This objective will generally produce a periodogram that is qualitatively similar to the log likelihood spectrum
# but it has been used to improve the reliability of transit search in the presence of correlated noise.
```
**Period Grid**
```
# It is possible to provide a specific period grid as follows:
model = BoxLeastSquares(t * u.day, y, dy=0.01)
periods = np.linspace(2.5, 3.5, 1000) * u.day
periodogram = model.power(periods, 0.2)
plt.plot(periodogram.period, periodogram.power)
# However, if the period grid is too coarse, the correct period might be missed.
model = BoxLeastSquares(t * u.day, y, dy=0.01)
periods = np.linspace(0.5, 10.5, 15) * u.day
periodogram = model.power(periods, 0.2)
plt.plot(periodogram.period, periodogram.power)
```
**Peak Statistics**
```
# The compute_stats method can be used to calculate several statistics of a candidate transit.
model = BoxLeastSquares(t * u.day, y, dy=0.01)
periodogram = model.autopower(0.2)
max_power = np.argmax(periodogram.power)
stats = model.compute_stats(periodogram.period[max_power],
periodogram.duration[max_power],
periodogram.transit_time[max_power])
```
**Function**
```
def BLS(periodgrid,durationgrid,lightcurve):
'''
Purppose
------------------
A Box Least Squares function to print out the compute stats of the periodogram.
Parameters
-------------------
Period grid - describes how often a transit is happening
Duration Grid - describes the width of a transit
Lightcurve - lightkurve class object
Return
------------------
Calculate the peak using the method compute_stats.
'''
# assinging parameters to variables.
period = periodgrid
lc = lightcurve
# t = timestamps
t = durationgrid * u.day
# y = observations
y = lc * u.dimensionless_unscaled # indicate quantities that do not have a physical dimension
# assigned to a BLS object
model = BoxLeastSquares(t , y , dy=lc.flux_err) # dy is the uncertainty
periodogram = model.autopower(period,durationgrid)
#calculates the maximum statstics within the transit for period, duration, and transit time
max_power = np.argmax(periodogram.power)
stats = model.compute_stats(periodogram.period[max_power],
periodogram.duration[max_power],
periodogram.transit_time[max_power])
# stats is the one peak, periodgram is the arrays
return stats, periodogram
def periods(N=1000):
period=np.logspace(-0.523, 1.43, N, endpoint=True)
return period
def duration_grid(N=10):
duration=np.linspace(.01, 0.298, N)
return duration
pd = periods(N=1000)
dg = duration_grid(N=10)
```
| github_jupyter |
```
# from google.colab import drive
# drive.mount('/content/gdrive')
#model3_ifelif_blend_S8_2847_time_124791_public_8_2847_private_8_3057_local_7_592444_lgb
# 1 nov-------------------------
# на ЛБ ещё чуть лучше, опять же, поиграл с периодами:
# PERIODS = ["5H", "4H", "6H"]
# скор на ЛБ: 8.3581
# 17 nov--------------------------
# change "num_leaves": to 34 and min_gain_to_split to 3
# [150] training's mape: 0.358244 valid_1's mape: 0.358244
# y2019_11m_16d_22h_14min_49s_397741_ 7.592444 7.592444 30 5 3
# LB -> 8.2847
# 18 nov------------------------------
# S8_2847: ifelif 8.3057 8.2847 8.3057 Нет y2019_11m_16d_22h_07min_31s_124791_lgb_sub.csv 17.11.2019 01:13
#import utils
import pathlib
import time
import tqdm
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os,sys,inspect,pickle,json,time,datetime,re
import random as rn
import lightgbm as lgb
%matplotlib inline
#DATA_DIR = pathlib.Path("./")
# DATA_DIR = "../input/activity-atactic/"
# train_data = pd.read_csv(DATA_DIR + "activity_train.csv.zip", parse_dates=["date"], index_col="date", compression="zip")
# test_data = pd.read_csv(DATA_DIR + "activity_test.csv.zip", parse_dates=["date"], index_col="date", compression="zip")
# activity_test_target = pd.read_csv(DATA_DIR + "activity_test_timestamps.csv", index_col="date", parse_dates=["date"])
# print('train',train_data.shape, 'test',test_data.shape, 'activity_test_target',activity_test_target.shape)#, 'atactic_test_target',atactic_test_target.shape)
DATA_DIR = "../input/sibur-csv/"#"../baseline/"
# activity_tags = pd.read_csv(DATA_DIR.joinpath('gdrive/My Drive/activity_tags.csv'),
# index_col='feature', sep=';')
# activity_tags['description'] = activity_tags['description'] + ' (' + activity_tags['units'] + ')'
# activity_tags_dict = activity_tags['description'].to_dict()
train_data = pd.read_csv(DATA_DIR+"activity_train.csv.zip",
parse_dates=["date"], index_col="date",
compression="zip")
test_data = pd.read_csv(DATA_DIR+"activity_test.csv.zip",
parse_dates=["date"], index_col="date",
compression="zip")
# train_data.rename(activity_tags_dict, axis=1, inplace=True)
# test_data.rename(activity_tags_dict, axis=1, inplace=True)
activity_test_target = pd.read_csv(DATA_DIR+"activity_test_timestamps.csv",
index_col="date",
parse_dates=["date"])
atactic_test_target = pd.read_csv(DATA_DIR+"atactic_test_timestamps.csv",
index_col="date",
parse_dates=["date"])
# prepare data for feature engineering
data = pd.concat([train_data[test_data.columns], test_data]) ;print('data',data.shape) # use all columns except 4 targets
train_targets = train_data[["activity", "atactic_1", "atactic_2", "atactic_3"]].copy() ;print('train_targets',train_targets.shape)
#series = train_targets[['activity']].notnull().all(axis=1) ;print(len(series[series==True]), 'rows without any null in whole row') # 300759
# generate 825 new cols from joineddata
# data.drop('ТЭА (л/c)', axis=1, inplace=True) # 'f28' ## try without inplace!
data.drop("f28", axis=1, inplace=True)
#ACOLS = ["atactic_1", "atactic_2", "atactic_3"]
#not_null_atactic = train_targets.loc[train_targets[ACOLS].notnull().all(axis=1), ACOLS] ;print(not_null_atactic.shape)
PERIODS = ["4H", "5H", "6H"]
AGGREGATES = ["mean", "median", "std", "max", "min"]#, "sum"]
#PERIODS = ["3H"]
#AGGREGATES = ["mean"]
all_features = []
for period in tqdm.tqdm_notebook(PERIODS):
for agg in AGGREGATES:
print(period,agg,end=',')
rolling_features = data.rolling(period).aggregate(agg)
rolling_features.rename(lambda x: "_".join([x, period, agg]), axis=1, inplace=True)
all_features.append(rolling_features)
all_features = pd.concat(all_features, axis=1)
print('all_features',all_features.shape) #825/5/3 = 55 #15 new cols for each of 55 features
# join all new features
full_data = data.join(all_features) ;print('full_data',full_data.shape) # (566709, 880)
activity_train_with_nulls = train_targets[["activity"]].join(full_data.shift(6, freq="H")) ;print('activity_train_with_nulls',activity_train_with_nulls.shape) # add 6 hours empty rows to start, delete last 6 hours rows
#series = activity_train_with_nulls.notnull().all(axis=1) ;print(len(series[series==True]), 'rows without any null in whole row') # 300759
activity_train = activity_train_with_nulls[activity_train_with_nulls.notnull().all(axis=1)] ;print('activity_train',activity_train.shape)
#activity_test = full_data[full_data.index >= test_data.index[0]] ;print('activity_test',activity_test.shape)
test_activity_data = activity_test_target.join(full_data.shift(6, freq="H")).ffill() ; print('test_activity_data',test_activity_data.shape) # test_activity_data (85891, 880)
# full_data.tail(3)
# rename data
train_data = activity_train
#test_data_useless = activity_test
test_data = test_activity_data
extra_data1 = activity_test_target
print('train_data',train_data.shape, 'test_data',test_data.shape, 'extra_data1',extra_data1.shape)
# config for models tuning
config_lgb = """{
"models":"lgb",
"datasize":"max",
"data":"v02_add55x5x3roll",
"rs":17,
"lgb": {
"_comment":"lambda_l2,lambda_l1,min_gain_to_split 30 5 8",
"name":"lgb",
"tune":"",
"rs":17,
"metric":"mape",
"num_threads": -1,
"objective": "regression",
"verbosity": 1,
"rounds":150,
"verbose_eval":100,
"early_stopping_rounds":20000,
"lambda_l2":30,
"lambda_l1":5,
"min_gain_to_split":3,
"num_leaves":34
}
}"""
conf = json.loads(config_lgb.replace("'", "\""))
# lgb wrapper
class TrainLGB():
model = None
conf = None
history_callback = None
def __init__(self, conf):
self.conf = conf
#print('lgb init done')
return
def compile(self, x_train_shape):
#print('lgb define done')
return
def fit(self, x_train, y_train, x_val, y_val):
lgb_train = lgb.Dataset(x_train, label=y_train.reshape(-1))
lgb_val = lgb.Dataset(x_val, label=y_val.reshape(-1))
evals_result = {}
c=self.conf
#params = {'metric':'mape', 'num_threads': -1, 'objective': 'regression', 'verbosity': 1} # rmse
params = {
'metric':c['metric'],
'num_threads':c['num_threads'],
'objective': c['objective'],
'verbosity': c['verbosity'],
'is_training_metric': True,
'lambda_l2':c['lambda_l2'],
'lambda_l1':c['lambda_l1'],
'min_gain_to_split':c['min_gain_to_split'],
'num_leaves':c['num_leaves'],
} # rmse
rounds = c['rounds']
# https://lightgbm.readthedocs.io/en/latest/pythonapi/lightgbm.train.html
self.model = lgb.train(
params = params,
#lgb_train,
train_set = lgb_train,
num_boost_round = rounds,
valid_sets = [lgb_train,lgb_val],
verbose_eval=c['verbose_eval'],
early_stopping_rounds=c['early_stopping_rounds'],
callbacks=[lgb.record_evaluation(evals_result)] #same -> evals_result=evals_result,
#callbacks=[lgb.print_evaluation(period=1, show_stdv=True)] #same -> evals_result=evals_result,
)
steps = self.model.best_iteration #print(f"Best: {steps}") #print('lgb fit done')
self.history_callback = evals_result
return
def predict(self, x):
#print('lgb predict done')
return self.model.predict(x)
def get_costs(self):
oy_train_list = self.history_callback['training']['mape']
oy_train_list = list(np.array(oy_train_list, dtype=np.float32))
oy_val_list = self.history_callback['valid_1']['mape']
oy_val_list = list(np.array(oy_val_list, dtype=np.float32))
#ox_list = [x for x in range(1,self.conf['rounds']+1)] # early stopping can cause error
ox_list = [x for x in range(1,len(oy_train_list)+1)]
title = self.conf['name'] + '_'
for t in self.conf['tune'].split(','):
if t!='': title += t+'_'+str(self.conf[t])+'_'
return {'ox_list':ox_list, 'oy_train_list':oy_train_list, 'oy_val_list':oy_val_list, 'title':title}
TARGET = 'activity'
rn.seed(conf['rs'])
np.random.seed(conf['rs'])
# split rows
train = train_data
val = train_data
test = test_data
# split cols to numpy and norm
def split_cols_np(train, val, test, TARGET):
x_train = train.drop(TARGET, axis=1, inplace=False).values # faster if target first col: train.iloc[:, 1:].values
x_val = val.drop(TARGET, axis=1, inplace=False).values # val.iloc[:, 1:].values
x_test = test.values
y_train = train[[TARGET]].values
y_val = val[[TARGET]].values
return x_train, x_val, x_test, y_train, y_val
x_train, x_val, x_test, y_train, y_val = split_cols_np(train, val, test, TARGET) ;print('x_train',x_train.shape, 'y_train',y_train.shape, 'x_val',x_val.shape, 'y_val',y_val.shape, 'x_test',x_test.shape)
# normalize
def normalize_by_train(x_train, x_val, x_test, y_train, y_val, train, TARGET):
#center, scale = train.iloc[:, 1:].mean().values, train.iloc[:, 1:].std().values
center_x, scale_x = train.drop(TARGET, axis=1, inplace=False).mean().values, train.drop(TARGET, axis=1, inplace=False).std().values
center_y, scale_y = train[TARGET].mean(), train[TARGET].std()
x_train_n = (x_train - center_x)/scale_x
x_val_n = (x_val - center_x)/scale_x
x_test_n = (x_test - center_x)/scale_x
y_train_n = (y_train - center_y)/scale_y
y_val_n = (y_val - center_y)/scale_y
#print('x_train_n',x_train_n.shape, 'y_train_n',y_train_n.shape, 'x_val_n',x_val_n.shape, 'y_val_n', y_val_n.shape)
return x_train_n, x_val_n, x_test_n, y_train_n, y_val_n
x_train_n, x_val_n, x_test_n, y_train_n, y_val_n = normalize_by_train(x_train, x_val, x_test, y_train, y_val, train, TARGET) #;print('x_train_n',x_train_n.shape, 'y_train_n',y_train_n.shape, 'x_val_n',x_val_n.shape, 'y_val_n', y_val_n.shape)
def denormalize_by_train(y_train_pred_n, y_val_pred_n, y_test_pred_n, train, TARGET):
y_train_pred = y_train_pred_n * train[TARGET].std() + train[TARGET].mean()
y_val_pred = y_val_pred_n * train[TARGET].std() + train[TARGET].mean() # cause center/scale made from train only
y_test_pred = y_test_pred_n * train[TARGET].std() + train[TARGET].mean() # cause center/scale made from train only
return y_train_pred, y_val_pred, y_test_pred
def evaluate(y_train_pred_n, y_val_pred_n, y_test_pred_n, conf_i, submit=False):
# denormalize
y_train_pred, y_val_pred, y_test_pred = denormalize_by_train(y_train_pred_n, y_val_pred_n, y_test_pred_n, train, TARGET)
# convert to pandas
y_train_pred_p = pd.Series(y_train_pred.flatten(), index=train.index, name=TARGET+"_pred").sort_index()
y_val_pred_p = pd.Series(y_val_pred.flatten(), index=val.index, name=TARGET+"_pred").sort_index()
y_test_pred_p = pd.DataFrame(y_test_pred, columns=[TARGET], index=extra_data1.index) # activity_test_target
# evaluate
def mape(y_true, y_pred): return 100 * np.mean(np.abs((y_pred-y_true)/y_true))
conf_i['score_train']= mape(train[TARGET].values, y_train_pred_p.values)
conf_i['score_val'] = mape(val[TARGET].values, y_val_pred_p.values)
# submit
curr_time = re.sub(r'^(.*)-(.*)-(.*) (.*):(.*):(.*)\.(.*)$',r'y\1_\2m_\3d_\4h_\5min_\6s_\7_',str(datetime.datetime.now()))
#if submit: y_test_pred_p.loc[test.index].to_csv(os.path.join('output',curr_time+mtype+'_sub.csv'))
#if submit: y_test_pred_p.loc[test.index].to_csv(curr_time+mtype+'_sub.csv')
if submit: y_test_pred_p.loc[test.index].to_csv('submit_model3.csv')
# add log to log_model.csv file
#log_file = 'log_'+mtype+'.csv'
log_file = 'log\\log_model3_'+mtype+'.csv'
log_df = pd.DataFrame([conf_i], index=[curr_time])
if log_file in os.listdir(): log_df = pd.read_csv(log_file, index_col=0).append(log_df)
log_df.to_csv(log_file)
#if log_file in os.listdir('output'): log_df = pd.read_csv(os.path.join('output', log_file), index_col=0).append(log_df)
#log_df.to_csv(os.path.join('output', log_file))
# add log to separate file
#pd.DataFrame([conf_i], index=[curr_time]).to_csv(os.path.join('output',curr_time+mtype+'_log.csv'))
return
def split_model_config(conf_mtype): # in: model params with lists, out: list of params-combination
if conf_mtype['tune']=='': return [conf_mtype]
import itertools; model_configs = []; list_of_lists = []
def tunelists_to_floats(conf_mtype, comb):
conf_i = conf_mtype.copy()
for i,t in enumerate(conf_mtype['tune'].split(',')): conf_i[t] = comb[i]
return conf_i
# 1 build list of lists
for t in conf_mtype['tune'].split(','): list_of_lists.append(conf_mtype[t])
# 2 get all combinations of a list of lists
combs = list(itertools.product(*list_of_lists)) # [(2734, 10), (2734, 20), (2734, 30), (1, 10), (1, 20), (1, 30), (2, 10), (2, 20), (2, 30)]
# 3 fill separate model_config from each combination
for comb in combs:model_configs.append(tunelists_to_floats(conf_mtype, comb))
return model_configs
#print(split_model_config(conf['lgb']))
for mtype in conf['models'].split(','):
for conf_i in split_model_config(conf[mtype]):
if mtype=='lgb':
#from train_lgb import TrainLGB
model = TrainLGB(conf_i)
model.compile(x_train.shape)
model.fit(x_train_n, y_train_n, x_val_n, y_val_n)
y_train_pred_n = model.predict(x_train_n)
y_val_pred_n = model.predict(x_val_n)
y_test_pred_n = model.predict(x_test_n)
conf_i['history'] = model.get_costs() #model.history_callback
conf_i['datasize'] = conf['datasize']
conf_i['data'] = conf['data']
conf_i['rs_global'] = conf['rs']
evaluate(y_train_pred_n, y_val_pred_n, y_test_pred_n, conf_i, submit=True)
#plot_costs(*model.get_costs())
# change "num_leaves": to 34 and min_gain_to_split to 3
# [150] training's mape: 0.358244 valid_1's mape: 0.358244
# y2019_11m_16d_22h_14min_49s_397741_ 7.592444 7.592444 30 5 3
# LB -> 8.2847
mtype = conf['models'].split(',')[0]
#pd.read_csv('output/log_'+mtype+'.csv', index_col=0)[['score_train','score_val','lambda_l2','lambda_l1','min_gain_to_split']].sort_values(by=['score_val'])
#pd.read_csv('log_'+mtype+'.csv', index_col=0)[['score_train','score_val','lambda_l2','lambda_l1','min_gain_to_split']]
pd.read_csv('log\\log_model3_'+mtype+'.csv', index_col=0)[['score_train','score_val','lambda_l2','lambda_l1','min_gain_to_split']]
y_train_pred, y_val_pred, y_test_pred = denormalize_by_train(y_train_pred_n, y_val_pred_n, y_test_pred_n, train, TARGET)
y_train_pred_p = pd.Series(y_train_pred.flatten(), index=train.index, name=TARGET+"_pred").sort_index()
train["activity"].plot(c="steelblue")
y_train_pred_p.plot(c="forestgreen")
plt.legend(loc=0)
plt.tight_layout()
plt.title("Activity", fontsize=12)
```
| github_jupyter |
```
import math
import numpy as np
from sympy import *
t, s, k = symbols('t, s, k')
# Defino los métodos de suma de rectángulos - izquierda
def _Riemman_izq(Func, limA, limB, numI):
""" Método de la Bisección para encontrar raíces en un intervalo.
## Parámetros:
Func (function) : función que depende de una variable.
limA (function) : derivada de la función.
limB (int) : semilla de la solución.
numI (int) : número máximo de iteraciones.
## Devoluciones:
Area (float) : valor de area encontrado.
"""
Δx = (limB - limA) / numI
Alturas = 0
for i in range(numI):
Alturas += Func(limA + i * Δx)
Area = Δx * Alturas
return Area
# Defino los métodos de suma de rectángulos - izquierda
def _Riemman_der(Func, limA, limB, numI):
""" Método de la Bisección para encontrar raíces en un intervalo.
## Parámetros:
Func (function) : función que depende de una variable.
limA (function) : derivada de la función.
limB (int) : semilla de la solución.
numI (int) : número máximo de iteraciones.
## Devoluciones:
Area (float) : valor de area encontrado.
"""
Δx = (limB - limA) / numI
Alturas = 0
for i in range(numI):
Alturas += Func(limB - i * Δx)
Area = Δx * Alturas
return Area
# Defino los métodos de suma de rectángulos - izquierda
def _Riemman_med(Func, limA, limB, numI):
""" Método de la Bisección para encontrar raíces en un intervalo.
## Parámetros:
Func (function) : función que depende de una variable.
limA (function) : derivada de la función.
limB (int) : semilla de la solución.
numI (int) : número máximo de iteraciones.
## Devoluciones:
Area (float) : valor de area encontrado.
"""
Δx = (limB - limA) / numI
Alturas = 0
for i in range(numI):
Alturas += Func(limA + (2*i + 1)/2 * Δx)
Area = Δx * Alturas
return Area
# 1
A = [(1, 3), (0, 3)]
b = [14, 21]
x = np.matmul(np.linalg.inv(A), np.transpose(b))
x1 = x[0]
x2 = x[1]
ft = 2*t
Ft = Integral(ft, t)
Ft_ab = Integral(ft, (t, x1, x2))
# Evaluar la integral
Rt = Ft.doit()
Rt_ab = Ft_ab.doit()
display(Eq(Ft, Rt))
display(Ft_ab, Rt_ab)
# 2
A = [(1, 2), (-3, 4)]
b = [14, 18]
M11 = 4
detA = np.linalg.det(A)
Zt = Integral(s, (s, t, M11))
funtion_Zt = Zt.doit()
Zt1 = detA * funtion_Zt.subs(t, 1)
display(Zt, funtion_Zt, Zt1)
# math.floor(Z11)
# 3
Vr = math.e
Va = 2.72
er = 100 * abs(Va - Vr)/Vr
print(f"{er:7.1e}")
# 5
F1 = lambda x: x**2
_Riemman_izq(F1, 0, 2, 3)
# 6
F2 = lambda x: (x**3)**2
Va = _Riemman_der(F2, 0, 4, 5)
print(4250 - Va)
print(4500 - Va)
# 7
F3 = lambda k: math.e**k
vr = Integral(math.e**k, (k, 0, 3)).doit()
va = _Riemman_med(F3, 0, 3, 2)
er = 100 * abs(vr-va)/va
print(vr, va, er)
```
| github_jupyter |
```
file_1 = """Stock Close Beta Cap
Apple 188.72 0.2 895.667B
Tesla 278.62 0.5 48.338B"""
file_2 = """Employee Wage Hired Promotion
Linda 3000 2017 Yes
Bob 2000 2016 No
Joshua 800 2019 Yes"""
```
### My solution
Other approaches are possible
```
def parser(stringa):
"""
Parse string and returns dict of lists: keys are first line, lists are columns.
"""
# lines will be a list of lists
# each sub list contains the words of a single line
lines = list()
for line in stringa.splitlines():
lines.append(line.split())
keys = lines[0] # the first line is the key
lines = lines[1:] # now lines does not include the first line
result = dict()
count = 0
for key in keys:
values = [line[count] for line in lines]
result[key] = values
count += 1
return result
parser(file_1)
parser(file_2)
```
### Test
We want to verify carefully that everything works as intended
```
def feel_bored_1_test(function):
"""
Verify that function returns result1 and result2.
"""
result_1 = {'Stock': ['Apple', 'Tesla'], 'Close': ['188.72', '278.62'], 'Beta': ['0.2', '0.5'],
'Cap': ['895.667B', '48.338B']}
result_2 = {'Employee': ['Linda', 'Bob', 'Joshua'], 'Wage': ['3000', '2000', '800'], 'Hired': ['2017', '2016', '2019'],
'Promotion': ['Yes', 'No', 'Yes']}
results = list()
if function(file1) == result1:
print("Test 1 passed")
results.append(True)
else:
print("Test 1 not passed")
results.append(False)
if function(file2) == result2:
print("Test 2 passed")
results.append(True)
else:
print("Test 2 not passed")
results.append(False)
return results
```
We can follow DRY (Don't Repeat Yourself) with a for loop to improve the testing function
```
def feel_bored_1_test(function):
result_1 = {'Stock': ['Apple', 'Tesla'], 'Close': ['188.72', '278.62'], 'Beta': ['0.2', '0.5'],
'Cap': ['895.667B', '48.338B']}
result_2 = {'Employee': ['Linda', 'Bob', 'Joshua'], 'Wage': ['3000', '2000', '800'], 'Hired': ['2017', '2016', '2019'],
'Promotion': ['Yes', 'No', 'Yes']}
input_to_output = {file_1: result_1, file_2: result_2}
results = list()
count = 1
for key, value in input_to_output.items():
if function(key) == value:
results.append(True)
print(f"Test {count} passed")
else:
results.append(False)
print(f"Test {count} not passed")
count += 1
return results
feel_bored_1_test(parser)
```
### Improve code
```
def fast_parser(stringa):
"""
Parse string and returns dict of lists: keys are first line, lists are columns.
"""
lines = [line.split() for line in stringa.splitlines()] # list of lists
keys = lines.pop(0) # remove first line and assign to keys
result = {
key: [line[index] for line in lines] for index, key in enumerate(keys)
}
return result
```
### Everything appears to work as intended
```
feel_bored_1_test(fast_parser)
```
### Efficiency does not matter, but it's still interesting to measure
We can see that the difference is insignificant for small inputs
```
%%timeit
parser(file_1)
%%timeit
fast_parser(file_1)
```
### With bigger inputs, parsing efficiency becomes relevant
<br>
**Key Takeaway:**
<br>
Do not waste time on optimizing code if you don't need it
<br>
<br>
**Premature optimization is the root of all evil**
```
big_input = (file_1 + '\n') * 100
print(big_input)
%%timeit
parser(big_input)
%%timeit
fast_parser(big_input)
```
| github_jupyter |
# TV Script Generation
In this project, you'll generate your own [Seinfeld](https://en.wikipedia.org/wiki/Seinfeld) TV scripts using RNNs. You'll be using part of the [Seinfeld dataset](https://www.kaggle.com/thec03u5/seinfeld-chronicles#scripts.csv) of scripts from 9 seasons. The Neural Network you'll build will generate a new ,"fake" TV script, based on patterns it recognizes in this training data.
## Get the Data
The data is already provided for you in `./data/Seinfeld_Scripts.txt` and you're encouraged to open that file and look at the text.
>* As a first step, we'll load in this data and look at some samples.
* Then, you'll be tasked with defining and training an RNN to generate a new script!
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# load in data
import helper
data_dir = './data/Seinfeld_Scripts.txt'
text = helper.load_data(data_dir)
```
## Explore the Data
Play around with `view_line_range` to view different parts of the data. This will give you a sense of the data you'll be working with. You can see, for example, that it is all lowercase text, and each new line of dialogue is separated by a newline character `\n`.
```
view_line_range = (0, 10)
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import numpy as np
print('Dataset Stats')
print('Roughly the number of unique words: {}'.format(len({word: None for word in text.split()})))
lines = text.split('\n')
print('Number of lines: {}'.format(len(lines)))
word_count_line = [len(line.split()) for line in lines]
print('Average number of words in each line: {}'.format(np.average(word_count_line)))
print()
print('The lines {} to {}:'.format(*view_line_range))
print('\n'.join(text.split('\n')[view_line_range[0]:view_line_range[1]]))
```
---
## Implement Pre-processing Functions
The first thing to do to any dataset is pre-processing. Implement the following pre-processing functions below:
- Lookup Table
- Tokenize Punctuation
### Lookup Table
To create a word embedding, you first need to transform the words to ids. In this function, create two dictionaries:
- Dictionary to go from the words to an id, we'll call `vocab_to_int`
- Dictionary to go from the id to word, we'll call `int_to_vocab`
Return these dictionaries in the following **tuple** `(vocab_to_int, int_to_vocab)`
```
import problem_unittests as tests
from collections import Counter
def create_lookup_tables(text):
"""
Create lookup tables for vocabulary
:param text: The text of tv scripts split into words
:return: A tuple of dicts (vocab_to_int, int_to_vocab)
"""
# TODO: Implement Function
counts = Counter(text)
vocab = sorted(counts, key=counts.get, reverse=True)
vocab_to_int = {text: ii for ii, text in enumerate(vocab, 0)}
int_to_vocab = {ii: text for ii, text in enumerate(vocab, 0)}
# return tuple
return (vocab_to_int, int_to_vocab)
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_create_lookup_tables(create_lookup_tables)
```
### Tokenize Punctuation
We'll be splitting the script into a word array using spaces as delimiters. However, punctuations like periods and exclamation marks can create multiple ids for the same word. For example, "bye" and "bye!" would generate two different word ids.
Implement the function `token_lookup` to return a dict that will be used to tokenize symbols like "!" into "||Exclamation_Mark||". Create a dictionary for the following symbols where the symbol is the key and value is the token:
- Period ( **.** )
- Comma ( **,** )
- Quotation Mark ( **"** )
- Semicolon ( **;** )
- Exclamation mark ( **!** )
- Question mark ( **?** )
- Left Parentheses ( **(** )
- Right Parentheses ( **)** )
- Dash ( **-** )
- Return ( **\n** )
This dictionary will be used to tokenize the symbols and add the delimiter (space) around it. This separates each symbols as its own word, making it easier for the neural network to predict the next word. Make sure you don't use a value that could be confused as a word; for example, instead of using the value "dash", try using something like "||dash||".
```
def token_lookup():
"""
Generate a dict to turn punctuation into a token.
:return: Tokenized dictionary where the key is the punctuation and the value is the token
"""
# TODO: Implement Function
keys = ['.', ',', '"', ';', '!', '?', '(', ')', '-', '\n']
values = ['||Period||', '||Comma||', '||Quotation_Mark||', '||Semicolon||', '||Exclamation_mark||', '||Question_mark||',
'||Left_Parentheses||', '||Right_Parentheses||', '||Dash||', '||Return||']
tokens = {keys[i] : values[i] for i in range(len(keys))}
return tokens
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_tokenize(token_lookup)
```
## Pre-process all the data and save it
Running the code cell below will pre-process all the data and save it to file. You're encouraged to lok at the code for `preprocess_and_save_data` in the `helpers.py` file to see what it's doing in detail, but you do not need to change this code.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# pre-process training data
helper.preprocess_and_save_data(data_dir, token_lookup, create_lookup_tables)
```
# Check Point
This is your first checkpoint. If you ever decide to come back to this notebook or have to restart the notebook, you can start from here. The preprocessed data has been saved to disk.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import helper
import problem_unittests as tests
int_text, vocab_to_int, int_to_vocab, token_dict = helper.load_preprocess()
```
## Build the Neural Network
In this section, you'll build the components necessary to build an RNN by implementing the RNN Module and forward and backpropagation functions.
### Check Access to GPU
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import torch
# Check for a GPU
train_on_gpu = torch.cuda.is_available()
if not train_on_gpu:
print('No GPU found. Please use a GPU to train your neural network.')
```
## Input
Let's start with the preprocessed input data. We'll use [TensorDataset](http://pytorch.org/docs/master/data.html#torch.utils.data.TensorDataset) to provide a known format to our dataset; in combination with [DataLoader](http://pytorch.org/docs/master/data.html#torch.utils.data.DataLoader), it will handle batching, shuffling, and other dataset iteration functions.
You can create data with TensorDataset by passing in feature and target tensors. Then create a DataLoader as usual.
```
data = TensorDataset(feature_tensors, target_tensors)
data_loader = torch.utils.data.DataLoader(data,
batch_size=batch_size)
```
### Batching
Implement the `batch_data` function to batch `words` data into chunks of size `batch_size` using the `TensorDataset` and `DataLoader` classes.
>You can batch words using the DataLoader, but it will be up to you to create `feature_tensors` and `target_tensors` of the correct size and content for a given `sequence_length`.
For example, say we have these as input:
```
words = [1, 2, 3, 4, 5, 6, 7]
sequence_length = 4
```
Your first `feature_tensor` should contain the values:
```
[1, 2, 3, 4]
```
And the corresponding `target_tensor` should just be the next "word"/tokenized word value:
```
5
```
This should continue with the second `feature_tensor`, `target_tensor` being:
```
[2, 3, 4, 5] # features
6 # target
```
```
from torch.utils.data import TensorDataset, DataLoader
def batch_data(words, sequence_length, batch_size):
"""
Batch the neural network data using DataLoader
:param words: The word ids of the TV scripts
:param sequence_length: The sequence length of each batch
:param batch_size: The size of each batch; the number of sequences in a batch
:return: DataLoader with batched data
"""
# TODO: Implement function
n_batches = len(words) - sequence_length
features, target = [], []
for i in range(n_batches):
f = words[i: i+sequence_length]
t = words[i+sequence_length]
features.append(f)
target.append(t)
feature_tensors, target_tensors = torch.from_numpy(np.array(features)), torch.from_numpy(np.array(target))
data = TensorDataset(feature_tensors, target_tensors)
data_loader = torch.utils.data.DataLoader(data, batch_size=batch_size, shuffle=True)
# return a dataloader
return data_loader
# there is no test for this function, but you are encouraged to create
# print statements and tests of your own
```
### Test your dataloader
You'll have to modify this code to test a batching function, but it should look fairly similar.
Below, we're generating some test text data and defining a dataloader using the function you defined, above. Then, we are getting some sample batch of inputs `sample_x` and targets `sample_y` from our dataloader.
Your code should return something like the following (likely in a different order, if you shuffled your data):
```
torch.Size([10, 5])
tensor([[ 28, 29, 30, 31, 32],
[ 21, 22, 23, 24, 25],
[ 17, 18, 19, 20, 21],
[ 34, 35, 36, 37, 38],
[ 11, 12, 13, 14, 15],
[ 23, 24, 25, 26, 27],
[ 6, 7, 8, 9, 10],
[ 38, 39, 40, 41, 42],
[ 25, 26, 27, 28, 29],
[ 7, 8, 9, 10, 11]])
torch.Size([10])
tensor([ 33, 26, 22, 39, 16, 28, 11, 43, 30, 12])
```
### Sizes
Your sample_x should be of size `(batch_size, sequence_length)` or (10, 5) in this case and sample_y should just have one dimension: batch_size (10).
### Values
You should also notice that the targets, sample_y, are the *next* value in the ordered test_text data. So, for an input sequence `[ 28, 29, 30, 31, 32]` that ends with the value `32`, the corresponding output should be `33`.
```
# test dataloader
test_text = range(50)
t_loader = batch_data(test_text, sequence_length=5, batch_size=10)
data_iter = iter(t_loader)
sample_x, sample_y = data_iter.next()
print(sample_x.shape)
print(sample_x)
print()
print(sample_y.shape)
print(sample_y)
```
---
## Build the Neural Network
Implement an RNN using PyTorch's [Module class](http://pytorch.org/docs/master/nn.html#torch.nn.Module). You may choose to use a GRU or an LSTM. To complete the RNN, you'll have to implement the following functions for the class:
- `__init__` - The initialize function.
- `init_hidden` - The initialization function for an LSTM/GRU hidden state
- `forward` - Forward propagation function.
The initialize function should create the layers of the neural network and save them to the class. The forward propagation function will use these layers to run forward propagation and generate an output and a hidden state.
**The output of this model should be the *last* batch of word scores** after a complete sequence has been processed. That is, for each input sequence of words, we only want to output the word scores for a single, most likely, next word.
### Hints
1. Make sure to stack the outputs of the lstm to pass to your fully-connected layer, you can do this with `lstm_output = lstm_output.contiguous().view(-1, self.hidden_dim)`
2. You can get the last batch of word scores by shaping the output of the final, fully-connected layer like so:
```
# reshape into (batch_size, seq_length, output_size)
output = output.view(batch_size, -1, self.output_size)
# get last batch
out = output[:, -1]
```
```
import torch.nn as nn
class RNN(nn.Module):
def __init__(self, vocab_size, output_size, embedding_dim, hidden_dim, n_layers, dropout=0.5):
"""
Initialize the PyTorch RNN Module
:param vocab_size: The number of input dimensions of the neural network (the size of the vocabulary)
:param output_size: The number of output dimensions of the neural network
:param embedding_dim: The size of embeddings, should you choose to use them
:param hidden_dim: The size of the hidden layer outputs
:param dropout: dropout to add in between LSTM/GRU layers
"""
super(RNN, self).__init__()
# TODO: Implement function
self.output_size = output_size
self.n_layers = n_layers
self.hidden_dim = hidden_dim
# set class variables
self.embedding = nn.Embedding()
self.lstm = nn.LSTM()
# define model layers
def forward(self, nn_input, hidden):
"""
Forward propagation of the neural network
:param nn_input: The input to the neural network
:param hidden: The hidden state
:return: Two Tensors, the output of the neural network and the latest hidden state
"""
# TODO: Implement function
# return one batch of output word scores and the hidden state
return None, None
def init_hidden(self, batch_size):
'''
Initialize the hidden state of an LSTM/GRU
:param batch_size: The batch_size of the hidden state
:return: hidden state of dims (n_layers, batch_size, hidden_dim)
'''
# Implement function
# initialize hidden state with zero weights, and move to GPU if available
return None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_rnn(RNN, train_on_gpu)
```
### Define forward and backpropagation
Use the RNN class you implemented to apply forward and back propagation. This function will be called, iteratively, in the training loop as follows:
```
loss = forward_back_prop(decoder, decoder_optimizer, criterion, inp, target)
```
And it should return the average loss over a batch and the hidden state returned by a call to `RNN(inp, hidden)`. Recall that you can get this loss by computing it, as usual, and calling `loss.item()`.
**If a GPU is available, you should move your data to that GPU device, here.**
```
def forward_back_prop(rnn, optimizer, criterion, inp, target, hidden):
"""
Forward and backward propagation on the neural network
:param decoder: The PyTorch Module that holds the neural network
:param decoder_optimizer: The PyTorch optimizer for the neural network
:param criterion: The PyTorch loss function
:param inp: A batch of input to the neural network
:param target: The target output for the batch of input
:return: The loss and the latest hidden state Tensor
"""
# TODO: Implement Function
# move data to GPU, if available
# perform backpropagation and optimization
# return the loss over a batch and the hidden state produced by our model
return None, None
# Note that these tests aren't completely extensive.
# they are here to act as general checks on the expected outputs of your functions
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_forward_back_prop(RNN, forward_back_prop, train_on_gpu)
```
## Neural Network Training
With the structure of the network complete and data ready to be fed in the neural network, it's time to train it.
### Train Loop
The training loop is implemented for you in the `train_decoder` function. This function will train the network over all the batches for the number of epochs given. The model progress will be shown every number of batches. This number is set with the `show_every_n_batches` parameter. You'll set this parameter along with other parameters in the next section.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
def train_rnn(rnn, batch_size, optimizer, criterion, n_epochs, show_every_n_batches=100):
batch_losses = []
rnn.train()
print("Training for %d epoch(s)..." % n_epochs)
for epoch_i in range(1, n_epochs + 1):
# initialize hidden state
hidden = rnn.init_hidden(batch_size)
for batch_i, (inputs, labels) in enumerate(train_loader, 1):
# make sure you iterate over completely full batches, only
n_batches = len(train_loader.dataset)//batch_size
if(batch_i > n_batches):
break
# forward, back prop
loss, hidden = forward_back_prop(rnn, optimizer, criterion, inputs, labels, hidden)
# record loss
batch_losses.append(loss)
# printing loss stats
if batch_i % show_every_n_batches == 0:
print('Epoch: {:>4}/{:<4} Loss: {}\n'.format(
epoch_i, n_epochs, np.average(batch_losses)))
batch_losses = []
# returns a trained rnn
return rnn
```
### Hyperparameters
Set and train the neural network with the following parameters:
- Set `sequence_length` to the length of a sequence.
- Set `batch_size` to the batch size.
- Set `num_epochs` to the number of epochs to train for.
- Set `learning_rate` to the learning rate for an Adam optimizer.
- Set `vocab_size` to the number of unique tokens in our vocabulary.
- Set `output_size` to the desired size of the output.
- Set `embedding_dim` to the embedding dimension; smaller than the vocab_size.
- Set `hidden_dim` to the hidden dimension of your RNN.
- Set `n_layers` to the number of layers/cells in your RNN.
- Set `show_every_n_batches` to the number of batches at which the neural network should print progress.
If the network isn't getting the desired results, tweak these parameters and/or the layers in the `RNN` class.
```
# Data params
# Sequence Length
sequence_length = # of words in a sequence
# Batch Size
batch_size =
# data loader - do not change
train_loader = batch_data(int_text, sequence_length, batch_size)
# Training parameters
# Number of Epochs
num_epochs =
# Learning Rate
learning_rate =
# Model parameters
# Vocab size
vocab_size =
# Output size
output_size =
# Embedding Dimension
embedding_dim =
# Hidden Dimension
hidden_dim =
# Number of RNN Layers
n_layers =
# Show stats for every n number of batches
show_every_n_batches = 500
```
### Train
In the next cell, you'll train the neural network on the pre-processed data. If you have a hard time getting a good loss, you may consider changing your hyperparameters. In general, you may get better results with larger hidden and n_layer dimensions, but larger models take a longer time to train.
> **You should aim for a loss less than 3.5.**
You should also experiment with different sequence lengths, which determine the size of the long range dependencies that a model can learn.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# create model and move to gpu if available
rnn = RNN(vocab_size, output_size, embedding_dim, hidden_dim, n_layers, dropout=0.5)
if train_on_gpu:
rnn.cuda()
# defining loss and optimization functions for training
optimizer = torch.optim.Adam(rnn.parameters(), lr=learning_rate)
criterion = nn.CrossEntropyLoss()
# training the model
trained_rnn = train_rnn(rnn, batch_size, optimizer, criterion, num_epochs, show_every_n_batches)
# saving the trained model
helper.save_model('./save/trained_rnn', trained_rnn)
print('Model Trained and Saved')
```
### Question: How did you decide on your model hyperparameters?
For example, did you try different sequence_lengths and find that one size made the model converge faster? What about your hidden_dim and n_layers; how did you decide on those?
**Answer:** (Write answer, here)
---
# Checkpoint
After running the above training cell, your model will be saved by name, `trained_rnn`, and if you save your notebook progress, **you can pause here and come back to this code at another time**. You can resume your progress by running the next cell, which will load in our word:id dictionaries _and_ load in your saved model by name!
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import torch
import helper
import problem_unittests as tests
_, vocab_to_int, int_to_vocab, token_dict = helper.load_preprocess()
trained_rnn = helper.load_model('./save/trained_rnn')
```
## Generate TV Script
With the network trained and saved, you'll use it to generate a new, "fake" Seinfeld TV script in this section.
### Generate Text
To generate the text, the network needs to start with a single word and repeat its predictions until it reaches a set length. You'll be using the `generate` function to do this. It takes a word id to start with, `prime_id`, and generates a set length of text, `predict_len`. Also note that it uses topk sampling to introduce some randomness in choosing the most likely next word, given an output set of word scores!
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import torch.nn.functional as F
def generate(rnn, prime_id, int_to_vocab, token_dict, pad_value, predict_len=100):
"""
Generate text using the neural network
:param decoder: The PyTorch Module that holds the trained neural network
:param prime_id: The word id to start the first prediction
:param int_to_vocab: Dict of word id keys to word values
:param token_dict: Dict of puncuation tokens keys to puncuation values
:param pad_value: The value used to pad a sequence
:param predict_len: The length of text to generate
:return: The generated text
"""
rnn.eval()
# create a sequence (batch_size=1) with the prime_id
current_seq = np.full((1, sequence_length), pad_value)
current_seq[-1][-1] = prime_id
predicted = [int_to_vocab[prime_id]]
for _ in range(predict_len):
if train_on_gpu:
current_seq = torch.LongTensor(current_seq).cuda()
else:
current_seq = torch.LongTensor(current_seq)
# initialize the hidden state
hidden = rnn.init_hidden(current_seq.size(0))
# get the output of the rnn
output, _ = rnn(current_seq, hidden)
# get the next word probabilities
p = F.softmax(output, dim=1).data
if(train_on_gpu):
p = p.cpu() # move to cpu
# use top_k sampling to get the index of the next word
top_k = 5
p, top_i = p.topk(top_k)
top_i = top_i.numpy().squeeze()
# select the likely next word index with some element of randomness
p = p.numpy().squeeze()
word_i = np.random.choice(top_i, p=p/p.sum())
# retrieve that word from the dictionary
word = int_to_vocab[word_i]
predicted.append(word)
# the generated word becomes the next "current sequence" and the cycle can continue
current_seq = np.roll(current_seq, -1, 1)
current_seq[-1][-1] = word_i
gen_sentences = ' '.join(predicted)
# Replace punctuation tokens
for key, token in token_dict.items():
ending = ' ' if key in ['\n', '(', '"'] else ''
gen_sentences = gen_sentences.replace(' ' + token.lower(), key)
gen_sentences = gen_sentences.replace('\n ', '\n')
gen_sentences = gen_sentences.replace('( ', '(')
# return all the sentences
return gen_sentences
```
### Generate a New Script
It's time to generate the text. Set `gen_length` to the length of TV script you want to generate and set `prime_word` to one of the following to start the prediction:
- "jerry"
- "elaine"
- "george"
- "kramer"
You can set the prime word to _any word_ in our dictionary, but it's best to start with a name for generating a TV script. (You can also start with any other names you find in the original text file!)
```
# run the cell multiple times to get different results!
gen_length = 400 # modify the length to your preference
prime_word = 'jerry' # name for starting the script
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
pad_word = helper.SPECIAL_WORDS['PADDING']
generated_script = generate(trained_rnn, vocab_to_int[prime_word + ':'], int_to_vocab, token_dict, vocab_to_int[pad_word], gen_length)
print(generated_script)
```
#### Save your favorite scripts
Once you have a script that you like (or find interesting), save it to a text file!
```
# save script to a text file
f = open("generated_script_1.txt","w")
f.write(generated_script)
f.close()
```
# The TV Script is Not Perfect
It's ok if the TV script doesn't make perfect sense. It should look like alternating lines of dialogue, here is one such example of a few generated lines.
### Example generated script
>jerry: what about me?
>
>jerry: i don't have to wait.
>
>kramer:(to the sales table)
>
>elaine:(to jerry) hey, look at this, i'm a good doctor.
>
>newman:(to elaine) you think i have no idea of this...
>
>elaine: oh, you better take the phone, and he was a little nervous.
>
>kramer:(to the phone) hey, hey, jerry, i don't want to be a little bit.(to kramer and jerry) you can't.
>
>jerry: oh, yeah. i don't even know, i know.
>
>jerry:(to the phone) oh, i know.
>
>kramer:(laughing) you know...(to jerry) you don't know.
You can see that there are multiple characters that say (somewhat) complete sentences, but it doesn't have to be perfect! It takes quite a while to get good results, and often, you'll have to use a smaller vocabulary (and discard uncommon words), or get more data. The Seinfeld dataset is about 3.4 MB, which is big enough for our purposes; for script generation you'll want more than 1 MB of text, generally.
# Submitting This Project
When submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as "dlnd_tv_script_generation.ipynb" and save another copy as an HTML file by clicking "File" -> "Download as.."->"html". Include the "helper.py" and "problem_unittests.py" files in your submission. Once you download these files, compress them into one zip file for submission.
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import sys
from math import sqrt
```
# Bar Graph
```
sets = ["SG.txt", "SA.txt", "FC.txt", "NF.txt"]
# n query steps, SG/SA/FC/NF, nsamples, word/letter
data = np.empty((4,4,5,2))
# n query steps, SG/SA/FC/NF, word/letter
means = np.empty((4,4,2))
for i, nq in enumerate(["", "1_", "5_", "10_"]):
for j, file in enumerate(sets):
data[i,j,:,:] = np.loadtxt("t_e2e_enc_query_" + nq + file, dtype=object).reshape(5,2,2)[:,:,1].astype(np.float32)
means[i,j,0] = np.mean(data[i,j,:,0]) # mean of word accuracy
means[i,j,1] = np.mean(data[i,j,:,1]) # mean of letter accuracy
# shape: nquery steps, SG/SA/FC/NF, word/letter
ci = np.empty((4,4,2))
for i in range(4):
for j in range(4):
for k in range(2):
ci[i,j,k] = (np.std(data[i,j,:,k])/sqrt(5)) * 1.96
labels = ["Standard Generalization", "Spurious Anticorrelation", "Full Combinatorial", "Novel Filler"]
x = np.arange(len(labels))
width = .10
fig, ax = plt.subplots(figsize=(20,6))
#ax.bar(x-width*3.5, means[0,:,0], width, yerr=ci[0,:,0], capsize=5, label="Inccorect Q - Word Level")
ax.bar(x-width*2.5, means[0,:,1], width, yerr=ci[0,:,1], capsize=5, label="Incorrect Q - Letter Level")
#ax.bar(x-width*1.5, means[1,:,0], width, yerr=ci[1,:,0], capsize=5, label="E2E 1Q - Word Level")
ax.bar(x-width/2, means[1,:,1], width, yerr=ci[1,:,1], capsize=5, label="E2E 1Q - Letter Level")
#ax.bar(x+width/2, means[2,:,0], width, yerr=ci[2,:,0], capsize=5, label="E2E 5Q - Word Level")
ax.bar(x+width*1.5, means[2,:,1], width, yerr=ci[2,:,1], capsize=5, label="E2E 5Q - Letter Level")
#ax.bar(x+width*2.5, means[3,:,0], width, yerr=ci[3,:,0], capsize=5, label="E2E 10Q - Word Level")
ax.bar(x+width*3.5, means[3,:,1], width, yerr=ci[3,:,1], capsize=5, label="E2E 10Q - Letter Level")
ax.set_ylabel('Mean Accuracy (%)',fontsize=14)
ax.set_xticks(x)
ax.set_xticklabels(labels, fontsize=14)
ax.legend(loc="lower left",fontsize=12)
fig.tight_layout()
plt.savefig("Mean_Accuracy_Bargraph.png")
plt.show()
labels = ["Standard Generalization", "Spurious Anticorrelation", "Full Combinatorial", "Novel Filler"]
x = np.arange(len(labels))
width = .10
fig, ax = plt.subplots(figsize=(20,6))
ax.bar(x-width*1.5, wm_letters_mean, width, yerr=wm_letters_ci, capsize=5, label="E2E - Letter Level", color="#3d5941")
ax.bar(x-width/2, wm_words_mean, width, yerr=wm_words_ci, capsize=5, label="E2E - Word Level", color = "#778868")
ax.bar(x+width/2, nested_letters_mean, width, yerr=nested_letters_ci, capsize=5, label="E2E + Q - Letter Level", color="#ca562c")
ax.bar(x+width*1.5, nested_words_mean, width, yerr=nested_words_ci, capsize=5, label="E2E + Q - Word Level", color="#de8a5a")
ax.set_ylabel('Mean Accuracy (%)',fontsize=14)
ax.set_xticks(x)
ax.set_xticklabels(labels, fontsize=14)
ax.legend(loc="lower left",fontsize=12)
fig.tight_layout()
plt.savefig("Mean_Accuracy_Bargraph.png")
plt.show()
```
# Box Plot
```
fig, (ax1, ax2, ax3, ax4) = plt.subplots(nrows=1, ncols=4, figsize=(16,8))
labels = ["Indirection - Letters", "Indirection - Words", "Enc/Dec - Letters", "Enc/Dec - Words"]
# rectangular box plot
bplot1 = ax1.boxplot([wm_letters[0], wm_words[0], nested_letters[0], nested_words[0]],
vert=True, # vertical box alignment
notch=True,
patch_artist=True, # fill with color
labels=labels) # will be used to label x-ticks
ax1.set_ylim(0,105)
ax1.tick_params(labelrotation=45)
ax1.set_title('Standard Generalization',fontsize=16)
ax1.set_ylabel('Mean Accuracy (%)',fontsize=16)
bplot2 = ax2.boxplot([wm_letters[1], wm_words[1], nested_letters[1], nested_words[1]],
vert=True, # vertical box alignment
notch=True,
patch_artist=True, # fill with color
labels=labels) # will be used to label x-ticks
ax2.set_ylim(0,105)
ax2.set_yticks([])
ax2.tick_params(labelrotation=45)
ax2.set_title('Spurious Anticorrelation',fontsize=16)
bplot3 = ax3.boxplot([wm_letters[2], wm_words[2], nested_letters[2], nested_words[2]],
vert=True, # vertical box alignment
notch=True,
patch_artist=True, # fill with color
labels=labels) # will be used to label x-ticks
ax3.set_ylim(0,105)
ax3.set_yticks([])
ax3.tick_params(labelrotation=45)
ax3.set_title('Full Combinatorial',fontsize=16)
bplot4 = ax4.boxplot([wm_letters[3], wm_words[3], nested_letters[3], nested_words[3]],
vert=True, # vertical box alignment
notch=True,
patch_artist=True, # fill with color
labels=labels) # will be used to label x-ticks
ax4.set_ylim(0,105)
ax4.set_yticks([])
ax4.tick_params(labelrotation=45)
ax4.set_title('Novel Filler',fontsize=16)
# fill with colors
colors = ['#3d5941', '#778868', '#ca562c', '#de8a5a']
for bplot in (bplot1, bplot2, bplot3, bplot4):
for patch, color in zip(bplot['boxes'], colors):
patch.set_facecolor(color)
fig.tight_layout()
plt.savefig("Mean_Accuracy_Boxplot.png")
plt.show()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import xarray as xr
from glob import glob
import pymongo
import pdb
from datetime import datetime, timedelta
from sqlalchemy import create_engine
import time
import psycopg2
import os
from io import StringIO
from scipy import sparse
from scipy.sparse.linalg import svds
import dask.dataframe as dd
engine = create_engine('postgresql://postgres:postgres@localhost:5432/atmos')
verdziFileNames = '/home/tyler/slicer/**/*.csv'
df = dd.read_csv(verdziFileNames)
filenames = glob(verdziFileNames)
filenames[0]
def create_idx(lat, lon, layer, delta):
row = (180 - lat) / delta
col = (360 - lon) / delta
idx = (1 + row)*col*layer
return idx
def row_idx(lat, delta=1):
return (90 - lat) / delta
def col_idx(lon, delta=1):
return (180 - lon) / delta
def vector_idx(row):
return (row['rowIdx'] + 1) * (row['colIdx'] + 1) * row['pres']
def get_layer(row, delta=1):
delta = 1
nrow = (180 - row['lat']) / delta
ncol = (360 - row['lon']) / delta
layer = int(row['index'] / ((1 + nrow)*ncol))
return layer
vdf = pd.DataFrame()
for idx, file in enumerate(filenames):
df = pd.read_csv(file, index_col=0)
time = file.split('_')[-1].strip('.csv')
value = 'T' + time
df = df.rename(index=str, columns={value:'temp'})
df['time'] = int(time)
df = df.reset_index()
df['index'] = df['index'].astype(int)
df['pres'] = df.apply(get_layer, axis=1)
df = df[['time', 'temp', 'lat', 'lon', 'pres', 'index']]
vdf = pd.concat([vdf, df], axis=0, sort=False)
#get depth from index
if idx % 100 == 0:
print('on idx: {}'.format(idx))
print('finished')
vdf = vdf.dropna()
vdf.to_csv('verdzi.csv')
vdf = pd.read_csv('verdzi.csv', index_col=0)
vdf = vdf.sort_values(['time', 'lat', 'lon'], ascending=True)
vdf['rowIdx'] = vdf['lat'].apply(lambda x: row_idx(x))
vdf['colIdx'] = vdf['lon'].apply(lambda x: col_idx(x))
for time, colDf in vdf.groupby('time'):
colDf['idx'] = colDf.apply(lambda row: vector_idx(row), axis=1)
data = colDf['temp'].values * np.cos(np.deg2rad(colDf['lat']))
row = colDf['idx'].astype(int).values
col = np.zeros(len(row))
colM = sparse.csc_matrix((data, (row, col)), shape=(180*360*25, 1))
if time == 0:
M = colM
else:
M = sparse.hstack([M, colM])
if time % 100 == 0:
print('on time: {}'.format(time))
print('finished')
sparse.save_npz('verdzi.npz', M)
u, s, vt = svds(M, k=30)
u.sum()
colDf['temp'].shape
vdf.to_csv('verdzi.csv')
M = sparse.csc_matrix([0], shape=(180*360*25, 1))
nCol = M.shape[1]
u.shape
```
| github_jupyter |
## SICP 习题 (2.79)解题总结: 为通用算术计算增加操作
SICP 习题 2.79 , 2.80 都是让我们给上面题目的通用算术计算增加操作,分别是 `equ?` 操作和`=zero?`操作,`equ?`用于判断两个数是否相等,`=zero?`用于判断一个数是否等于零。
这里没有太多难理解的,就是工程实现,给所有的类型添加操作。
代码如下:
```
(import 'math)
(python-exec "
def cos(input_number):
return math.cos(input_number)
")
(python-exec "
def sin(input_number):
return math.sin(input_number)
")
(python-exec "
def atan(input_number):
return math.atan(input_number)
")
(define (gcd a b)
(if (= b 0)
a
(gcd b (remainder a b))))
(define (assoc key records)
(cond ((null? records) #f)
((equal? key (caar records)) (car records))
(else (assoc key (cdr records)))))
(define (make-table)
(let ((local-table (list '*table*)))
(define (lookup key-1 key-2)
(let ((subtable (assoc key-1 (cdr local-table))))
(if subtable
(let ((record (assoc key-2 (cdr subtable))))
(if record
(cdr record)
(begin
(display "record not found: ") (display key-2) (newline)
#f
)))
(begin
(display "subtable not found: ") (display key-1)(newline)
#f))))
(define (insert! key-1 key-2 value)
(let ((subtable (assoc key-1 (cdr local-table))))
(if subtable
(let ((record (assoc key-2 (cdr subtable))))
(if record
(set-cdr! record value)
(set-cdr! subtable
(cons (cons key-2 value)
(cdr subtable)))))
(set-cdr! local-table
(cons (list key-1
(cons key-2 value))
(cdr local-table)))))
'ok)
(define (dispatch m)
(cond ((eq? m 'lookup-proc) lookup)
((eq? m 'insert-proc!) insert!)
(else (error "Unknown operation -- TABLE" m))))
dispatch))
(define operation-table (make-table))
(define get (operation-table 'lookup-proc))
(define put (operation-table 'insert-proc!))
(define (attach-tag type-tag contents)
(cond ((number? contents) contents)
(else (cons type-tag contents))))
(define (type-tag datum)
(if (pair? datum)
(car datum)
(if (number? datum)
'scheme-number
(error "Bad tagged type -- TYPE " datum))))
(define (contents datum)
(if (pair? datum)
(cdr datum)
(if (number? datum)
datum
(error "Bad tagged datum -- CONTENTS" datum))))
(define (add x y) (apply-generic 'add x y))
(define (sub x y) (apply-generic 'sub x y))
(define (mul x y) (apply-generic 'mul x y))
(define (div x y) (apply-generic 'div x y))
(define (d-equ? x y) (apply-generic 'equ? x y))
(define (d=zero? x) (apply-generic '=zero? x))
(define (install-scheme-number-package)
(define (tag x)
(attach-tag 'scheme-number x))
(put 'add '(scheme-number scheme-number)
(lambda (x y) (tag (+ x y))))
(put 'sub '(scheme-number scheme-number)
(lambda (x y) (tag (- x y))))
(put 'mul '(scheme-number scheme-number)
(lambda (x y) (tag (* x y))))
(put 'div '(scheme-number scheme-number)
(lambda (x y) (tag (/ x y))))
(put 'make 'scheme-number
(lambda (x) (tag x)))
(put 'equ? '(scheme-number scheme-number)
(lambda (x y) (tag (eq? x y))))
(put '=zero? '(scheme-number)
(lambda (x) (tag (zero? x))))
'done)
(define (make-scheme-number n)
((get 'make 'scheme-number) n))
(define (install-rational-package)
(define (numer x) (car x))
(define (denom x) (cdr x))
(define (make-rat n d)
(let ((g (gcd n d)))
(cons (/ n g) (/ d g))))
(define (add-rat x y)
(make-rat (+ (* (numer x) (denom y))
(* (numer y) (denom x)))
(* (denom x) (denom y))))
(define (sub-rat x y)
(make-rat (- (* (numer x) (denom y))
(* (numer y) (denom x)))
(* (denom x) (denom y))))
(define (mul-rat x y)
(make-rat (* (numer x) (numer y))
(* (denom x) (denom y))))
(define (div-rat x y)
(make-rat (* (numer x) (denom y))
(* (denom x) (numer y))))
(define (tag x) (attach-tag 'rational x))
(put 'add '(rational rational)
(lambda (x y) (tag (add-rat x y))))
(put 'sub '(rational rational)
(lambda (x y) (tag (sub-rat x y))))
(put 'mul '(rational rational)
(lambda (x y) (tag (mul-rat x y))))
(put 'div '(rational rational)
(lambda (x y) (tag (div-rat x y))))
(put 'equ? '(rational rational)
(lambda (x y) (tag (and (eq? (numer x) (numer y)) (eq? (denom x) (denom y))))))
(put '=zero? '(rational)
(lambda (x) (tag (zero? (numer x)))))
(put 'make 'rational
(lambda (n d) (tag (make-rat n d))))
'done)
(define (make-rational n d)
((get 'make 'rational) n d))
(define (apply-generic op . args)
(let ((type-tags (map type-tag args)))
(let ((proc (get op type-tags)))
(if proc
(apply proc (map contents args))
(error
"No method for these types -- APPLY-GENERIC"
(list op type-tags))))))
(define (real-part z) (apply-generic 'real-part z))
(define (imag-part z) (apply-generic 'imag-part z))
(define (magnitude z) (apply-generic 'magnitude z))
(define (angle z) (apply-generic 'angle z))
(define (install-rectangular-package)
(define (real-part z) (car z))
(define (imag-part z) (cdr z))
(define (make-from-real-imag x y) (cons x y))
(define (magnitude z)
(sqrt (+ (square (real-part z))
(square (imag-part z)))))
(define (angle z)
(atan (/ (imag-part z) (real-part z))))
(define (make-from-mag-ang r a)
(cons (* r (cos a)) (* r (sin a))))
(define (tag x) (attach-tag 'rectangular x))
(put 'real-part '(rectangular) real-part)
(put 'imag-part '(rectangular) imag-part)
(put 'magnitude '(rectangular) magnitude)
(put 'angle '(rectangular) angle)
(put 'make-from-real-imag 'rectangular
(lambda (x y) (tag (make-from-real-imag x y))))
(put 'make-from-mag-ang 'rectangular
(lambda (r a) (tag (make-from-mag-ang r a))))
'done)
(define (install-polar-package)
(define (magnitude z) (car z))
(define (angle z) (cdr z))
(define (make-from-mag-ang r a) (cons r a))
(define (real-part z)
(* (magnitude z) (cos (angle z))))
(define (imag-part z)
(* (magnitude z) (sing (angle z))))
(define (make-from-real-imag x y)
(cons (sqrt (+ (square x) (square y)))
(atan (/ y x))))
(define (tag x) (attach-tag 'polar x))
(put 'real-part '(polar) real-part)
(put 'imag-part '(polar) imag-part)
(put 'magnitude '(polar) magnitude)
(put 'angle '(polar) angle)
(put 'make-from-real-imag 'polar
(lambda (x y) (tag (make-from-real-imag x y))))
(put 'make-from-mag-ang 'polar
(lambda (r a) (tag (make-from-mag-ang r a))))
'done)
(define (install-complex-package)
(define (make-from-real-imag x y)
((get 'make-from-real-imag 'rectangular) x y))
(define (make-from-mag-ang r a)
((get 'make-from-mag-ang 'polar) r a))
(define (add-complex z1 z2)
(make-from-real-imag (+ (real-part z1) (real-part z2))
(+ (imag-part z1) (imag-part z2))))
(define (sub-complex z1 z2)
(make-from-real-imag (- (real-part z1) (real-part z2))
(- (imag-part z1) (imag-part z2))))
(define (mul-complex z1 z2)
(make-from-mag-ang (* (magnitude z1) (magnitude z2))
(+ (angle z1) (angle z2))))
(define (div-complex z1 z2)
(make-from-mag-ang (/ (magnitude z1) (magnitude z2))
(- (angle z1 ) (angle z2))))
(define (tag z) (attach-tag 'complex z))
; the following lines are added for this execise:
(put 'real-part '(complex) real-part)
(put 'imag-part '(complex) imag-part)
(put 'magnitude '(complex) magnitude)
(put 'angle '(complex) angle)
;end of the execise
(put 'add '(complex complex)
(lambda (z1 z2) (tag (add-complex z1 z2))))
(put 'sub '(complex complex)
(lambda (z1 z2) (tag (sub-complex z1 z2))))
(put 'mul '(complex complex)
(lambda (z1 z2) (tag (mul-complex z1 z2))))
(put 'div '(complex complex)
(lambda (z1 z2) (tag (div-complex z1 z2))))
(put 'make-from-real-imag 'complex
(lambda (x y) (tag (make-from-real-imag x y))))
(put 'make-from-mag-ang 'complex
(lambda (r a) (tag (make-from-mag-ang r a))))
'done)
(define (make-complex-from-real-imag x y)
((get 'make-from-real-imag 'complex) x y))
(define (make-complex-from-mag-ang r a)
((get 'make-from-mag-ang 'complex) r a))
(define (display-complex complex)
(display "--------------------") (newline)
(display "real part:") (display (real-part complex)) (newline)
(display "imag part:") (display (imag-part complex)) (newline)
(display "magnitude:") (display (magnitude complex)) (newline)
(display "angle :") (display (angle complex))(newline)
(display "--------------------") (newline)
)
(define (start-test-2-79)
(install-scheme-number-package)
(install-rational-package)
(display (d-equ? 2 2))(newline)
(display (d-equ? 2 3))(newline)
(display (d=zero? 2))(newline)
(display (d=zero? 0))(newline)
(display (d-equ? (make-scheme-number 2) (make-scheme-number 2)))(newline)
(display (d-equ? (make-scheme-number 2) (make-scheme-number 3)))(newline)
(display (d=zero? (make-scheme-number 2)))(newline)
(display (d=zero? (make-scheme-number 0)))(newline)
(display (d-equ? (make-rational 2 3) (make-rational 2 3)))(newline)
(display (d-equ? (make-rational 2 3) (make-rational 3 3)))(newline)
(display (d-equ? (make-rational 4 6) (make-rational 2 3)))(newline)
(display (d=zero? (make-rational 2 3)))(newline)
(display (d=zero? (make-rational 0 3)))(newline)
)
(start-test-2-79)
```
| github_jupyter |
```
%matplotlib inline
```
# TP1: The Interaction-driven Metal-Insulator Transition
## GOAL:
1) To understand the difference between the electronic spectral functions of a metal and of a Mott insulator.
2) To understand the interaction-driven metal-insulator transition in the Hubbard Model (the reference model for this problem), known as the **Mott Metal-Insulator Transition** (MIT).
To achieve these goals you will have to understand how to read Green's function and connect them to physical properties. We will use the (exact!) Dynamical-Mean-Field-Theory (DMFT) solution of the Hubbard Model on the infinite-dimensional Bethe lattice. The DMFT is numerically implemented using the Iterative Perturbation Theory (IPT), which provides (approximate but good) Green's functions. To this purpose you will have to run a PYTHON code, manipulating the input/output in order to simulate different physical situations and interpret the outputs.
It would be interesting to study also the details of the DMFT implementation and the IPT impurity solver, but it will not be possible on this occasion. Use it then as a black box to perform virtual theoretical experiments.
You will have to provide us at the end of the Tutorial a complete report, answering to the questions of the section EXERCISES. Include graphs and commentaries whenever asked (or possible) at your best convenience.
Real frequency IPT solver single band Hubbard model
===================================================
Here it is the IPT code.
```
# Author: Óscar Nájera
# License: 3-clause BSD
from __future__ import division, absolute_import, print_function
import scipy.signal as signal
import numpy as np
import matplotlib.pyplot as plt
plt.matplotlib.rcParams.update({'axes.labelsize': 22,
'axes.titlesize': 22, })
def fermi_dist(energy, beta):
""" Fermi Dirac distribution"""
exponent = np.asarray(beta * energy).clip(-600, 600)
return 1. / (np.exp(exponent) + 1)
def semi_circle_hiltrans(zeta, D):
"""Calculate the Hilbert transform with a semicircular DOS """
sqr = np.sqrt(zeta**2 - D**2)
sqr = np.sign(sqr.imag) * sqr
return 2 * (zeta - sqr) / D**2
def pertth(Aw, nf, U):
"""Imaginary part of the second order diagram"""
# because of ph and half-fill in the Single band one can work with
# A^+ only
Ap = Aw * nf
# convolution A^+ * A^+
App = signal.fftconvolve(Ap, Ap, mode='same')
# convolution A^-(-w) * App
Appp = signal.fftconvolve(Ap, App, mode='same')
return -np.pi * U**2 * (Appp + Appp[::-1])
def dmft_loop(gloc, w, U, beta, D, loops):
"""DMFT Loop for the single band Hubbard Model at Half-Filling
Parameters
----------
gloc : complex 1D ndarray
local Green's function to use as seed
w : real 1D ndarray
real frequency points
U : float
On site interaction, Hubbard U
beta : float
Inverse temperature
D : float
Half-bandwidth of the noninteracting system
loops : int
Amount of DMFT loops to perform
Returns
-------
gloc : complex 1D ndarray
DMFT iterated local Green's function
sigma : complex 1D ndarray
DMFT iterated self-energy
"""
dw = w[1] - w[0]
eta = 2j * dw
nf = fermi_dist(w, beta)
for i in range(loops):
# Self-consistency
g0 = 1 / (w + eta - .25 * D**2 * gloc)
# Spectral-function of Weiss field
A0 = -g0.imag / np.pi
# Clean for PH and Half-fill
A0 = 0.5 * (A0 + A0[::-1])
# Second order diagram
isi = pertth(A0, nf, U) * dw * dw
# Kramers-Kronig relation, uses Fourier Transform to speed convolution
hsi = -signal.hilbert(isi, len(isi) * 4)[:len(isi)].imag
sigma = hsi + 1j * isi
# Semi-circle Hilbert Transform
gloc = semi_circle_hiltrans(w - sigma, D)
return gloc, sigma
```
## Example Simulation
```
U = 2.0
D = 2.0
beta = 10**6
w = np.linspace(-6*D, 6*D, 2**12)
gloc = semi_circle_hiltrans(w + 1e-3j,D)
#gloc= 0.0
gloc, sigma_loc = dmft_loop(gloc, w, U, beta, D, 100)
fig,ax = plt.subplots(2,1)
ax[0].plot(w, -gloc.imag, label=r'$\pi A(\omega)$')
ax[0].legend(loc=0)
ax[1].plot(w, sigma_loc.real, '-', label=r'$Re \Sigma(\omega)$')
ax[1].plot(w, sigma_loc.imag, '-', label=r'$Im \Sigma(\omega)$')
ax[1].set_xlabel(r'$\omega$')
ax[1].legend(loc=0)
#plt.ylim([0, 0.1])
#plt.xlim([-1, 1])
```
## The effect of the self-consistency condition
1. The DMFT implementation is an iterative process which requires a certain number of loops before convergence is reached. But one needs a starting point, i.e. one needs to input an initial guess of the spectral function. Can you say which is the starting guess in the DMFT implementation considered above?
2. For $U < Uc_2$, run the code with *just one dmft iteration*, using the same values of $U$ that you have employed above. This one-loop run corresponds to the solution of the Single Impurity Anderson Model SIAM (or **Kondo Model**, see Marc's lecture notes) for a semi-circular conduction band. Compare the spectral function $A(\omega)$ of the one-iteration loop with the fully converged one. Do you see the MIT?
3. Repeat the same excercise of point 2., this time inputing no bath (i.e. the intial guess is zero), and always one iteration loop. This means that you are essentially solving the isolated atom problem with on-site interaction $U$. Draw a conclusions about the effect of the self-conistency in solving the MIT problem.
# Let's start now
## Excercises
We shall first work at low temperature, choose $\beta=100$. The imaginary part of the Green's function $A(\omega)= -\Im G(\omega)/\pi$ can be directly connected to physical observables (e.g. spectral functions) or transport (e.g. specific heat).
## The Metal-to-Insulator Transition
1. Plot first $A(\omega)$ in the case $U=0$. This is the semicircular density of states used in the DMFT code, i.e. $-\Im G(\omega)/\pi= D(\omega)$. The system is in this case "halffilled", i.e. there is one elctron per atomic orbital. Can you indicate then where the occupied states are on the plot of the density of states $A(\omega)$? Why the system is a metal according to the band theory of solids?
2. Run the code for several values of $U$ (the half-bandwidth is set $D=1$) to check out the metal-insulator transition. Display $A(\omega)$ for some rapresentative values of $U$, in the metallic and insulating sides.
3. Approach the MIT at $Uc_2$ (metal to insulator) from below ($U< Uc_2$) and describe how the different contributions to the $A(\omega)$ (quasiparticle peak and Hubbard bands) evolve as you get close to the critical value. Why can we state that for $U> Uc_2$ the system is finally insulating? Give your extimation of $Uc_2$.
4. At the MIT point, the quaiparticle peak has completely disappered. But where is the spectral weight of the peak gone? Explain why we expect that the total spectral weight [i.e. the area under the curve $A(\omega)$] is conserved.
## Selfenergy
1. Plot now the real part and the imaginary part of the self-energy $\Sigma(\omega)$, first for $U< Uc_2$, then for $U \ge Uc_2$. Use the same values of $U$ that you have used above.
2. We are going now to concentrate on the low energy part $\omega\to 0$, where a quasiparticle peak is observable in the metallic phase, even for values of $U$ very close to the MIT. The Fermi liquid theory of metals states that the selfenergy is a regular function of $\omega$, which can be Taylor expanded: $$
\Re\Sigma \simeq \mu_0 + \alpha \omega+ O(\omega^2) \quad \quad
\Im\Sigma \sim \omega^2+ O(\omega^3)
$$
For which values of $U$ can we state that the system is in a Fermi liquid state?
3. We shall now see how the Fermi liquid theory may be useful to understand the beahvior of spectral functions. The local Green's function (entering the DMFT self-consistency condition) is the Hilbert transform of the lattice Green's function $G(\omega,\varepsilon)$
$$
G_{loc}(\omega)= \,\int D(\varepsilon) \, G(\omega, \varepsilon)\, d\varepsilon=
\,\int D(\varepsilon)/\left[ \omega-\varepsilon- \Sigma(\omega) \right] \, d\varepsilon $$
where $D(\varepsilon)= -(1/\pi) \Im G_{loc}(\varepsilon,U=0)$ and in our case (infinite dimension) $\Sigma(\omega)$ is independent of $\varepsilon$. Show that the lattice Green's function can be written within the Fermi liquid theory ($\omega\to0$) as a almost free-particle Green's function $$
G(\omega, \xi)\sim \frac{Z}{\omega-\xi} $$
where the main difference is the factor $Z$, known as *quasi-particle renormalization factor* .
* Evaluate $Z$ for different values of $U$ (use the real part of the self-energy) and plot it as a function of $U$. Explain why $Z$ is useful to describe the MIT. *Hint: in order to extract a part of the Green's function array (let's call it e.g. $gx$) between frequencies, e.g. $[-2,2]$, define a logical array of frequencies $w_r= (w< 2 )*(w> -2)$, then the desired reduced Green's function array is simply given by $gx[w_r]$.*
* Calculate (for a couple of $U$ values used above) the area under the quasiparticle peaks and plot them as a function of $U$. Relate this area with the quasiparticle residue $Z$ (you should plot them as a function of $U$).
* Determine the exact relation between the quasiparticle peak and $Z$ also analytically. Explain finally why it is said that *a Mott transition is a breaking of the Fermi Liquid Theory*.
# First order transition
The MIT is a first order transiton, i.e. there is a region of coexistence between the insulator and the metallic phase.
1. Show that there is an interval of $U$ where DMFT self-consistency provides two solutions. Determine the $Uc_1$, i.e. the critical value of $U$ where the insulating state changes into a metal by reducing $U$. *(Hint: you have to use an appropriate starting guess to obtain the insulating solution, start your investigations with values of $U$ in the insulating side).*
2. Is $Z$ the right parameter to describe the insulator-to-metal transition too? Argument your answer.
# $T-U \,$ phase diagram and the $T$-driven transition
1. Run now the code for higher temperatures $T$, i.e. lower inverse temperatures $\beta= 75, 50, 25$ and determine for each temperature $T$ the $Uc_1$ and the $Uc_2$. Sketch then the coexistence region in the $T-U$ plane.
2. Observe the shape of the coexistence region in the $T-U$ space. Fix the interaction $U$ inside this co-existence region (e.g. $U=2.9$) and systematically increase the temperature. Compare the different density of states. Can you now explain the phase-diagram of VO$_2$, in particular the unusual not-metallic behavior at high temperature?
3. Can you give an interpretation of this physical behaviour with temperature in terms of the associated SIAM model, i.e. a spin impurity fluctuating with temperature in a bath of free electrons?
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style(style='whitegrid')
```
# Importing Data
Data was scraped from the Amazon website for Charge 2
```
df = pd.read_csv('charge2.csv', parse_dates =['date'])
df.head()
df = df.drop(columns=['web-scraper-order','web-scraper-start-url'])
df['stars'] = df.stars.apply(lambda x: x.split(' ')[0])
df.head()
df['length'] = df.review.apply(lambda x: len(x))
df['year'] = df.date.apply(lambda x: x.year)
df['stars'] = pd.to_numeric(df.stars)
df.head()
fig, ax = plt.subplots(figsize = (14,8))
sns.boxplot(df.stars, df.length)
plt.ylim(-20, 2000)
fig, ax = plt.subplots(figsize = (14,8))
df.groupby(['date'])['stars'].mean().plot(marker='.',linestyle='none')
fig, ax = plt.subplots(figsize = (14,8))
plt.plot(df.date, df.stars, alpha=0.02, marker='.', linestyle='none', markersize=40)
df.shape
fig, ax = plt.subplots(figsize = (14,8))
df.groupby('stars').size().plot(kind='bar', color='orange', alpha=0.7)
plt.xlabel('Star Rating')
plt.ylabel('Number of Ratings')
plt.title('Amazon Reviews for Fitbit Charge 2', fontsize=14)
```
# NLP
```
import gensim
import spacy
from nltk import FreqDist
from gensim.models import LdaModel, phrases
from gensim.models.wrappers import LdaMallet
from gensim.corpora import Dictionary
import pyLDAvis.gensim
import os, re, operator, warnings
warnings.filterwarnings('ignore') # Let's not pay heed to them right now
```
# SpaCy
```
terms = [u"heart rate", u"gps", u"sync", u"syncing", u"design",
u"charging", u"charger", u"battery", u"quality",
u"screen", u"service", u"sleep", u"alarm", u"reminder"]
#$ python -m spacy download en_core_web_md (run in shell if not installed)
nlp = spacy.load('en_core_web_lg')
text = df.review[12]
print(text)
# Process the text
doc = nlp(text)
for token in doc:
if token.lemma_ in terms:
print("{0}/{1} <--{2}-- {3}/{4}".format(
token.text, token.tag_, token.dep_, token.head.text, token.head.tag_))
for sentence in doc.sents:
chunk = []
for token in sentence:
if token.lemma_ in terms:
print(sentence)
print('---')
print("{0}/{1} <--{2}-- {3}/{4}".format(
token.text, token.tag_, token.dep_, token.head.text, token.head.tag_))
print('---')
for child in sentence.root.children:
chunk.append(child.text)
print(f'child: {child}')
from spacy import displacy
# In[2]:
doc = nlp(u"i am very disapointed especially about the heart rate tracker")
displacy.render(doc, style="dep", jupyter=True, options={'distance':90})
for sentence in doc.sents:
print(sentence)
print('--------------------')
'''#stemming
from nltk.stem.snowball import SnowballStemmer
stemmer = SnowballStemmer(language='english')'''
#A very slow way of extracting search terms
'''for term in terms:
for i in doc:
sims = nlp(stemmer.stem(term))[0].similarity(nlp(stemmer.stem(str(i))))
if sims > 0.7:
print(sims, i)'''
```
## Phrase Matcher
```
from spacy.matcher import PhraseMatcher
terms = [u"heart rate", u"gps", u"sync", u"syncing", u"design",
u"charging", u"charger", u"battery", u"quality",
u"screen", u"service", u"sleep", u"alarm", u"reminder"]
def phrase_matcher(review):
matched_list = []
matcher = PhraseMatcher(nlp.vocab)
# Only run nlp.make_doc to speed things up
patterns = [nlp.make_doc(text) for text in terms]
matcher.add("TerminologyList", None, *patterns)
matches = matcher(review)
for match_id, start, end in matches:
span = review[start:end]
matched_list.append(span.text)
return list(set(matched_list))
```
# Sentiment Analysis - TextBlob, Afinn, NLTK+Vader
```
#TEXTBLOB
from textblob import TextBlob
#AFINN
from afinn import Afinn
af = Afinn()
#NLTK
from nltk.sentiment.vader import SentimentIntensityAnalyzer
import nltk
nltk.download('vader_lexicon')
sid = SentimentIntensityAnalyzer()
```
## Phrase count in new DataFrame
```
#create a dataframe with our phrases as columns, to keep count of sentiment
phrase_df = pd.DataFrame(columns=terms)
def get_sentiment(input_string):
'''
Takes a string (sentence), and returns an averaged, three-model
sentiment score, from -1 to 1. Where -1 is negative sentiment, 0 is neutral,
and +1 is positive sentiment.
'''
#calculate sentiment score with TextBlob, Afinn, NLTK+Vader
txt_blob = TextBlob(input_string.text).sentiment[0]
a_fin = af.score(input_string.text)
nltk_vader = sid.polarity_scores(input_string.text).get('compound')
#TextBlob
if txt_blob > 0.1:
blob = 1
elif txt_blob == 0 and txt_blob <= 0.1:
blob = 0
else: blob = -1
#afinn
if a_fin > 1:
fin = 1
elif a_fin == 0 and a_fin <= 1:
fin = 0
else: fin = -1
#NLTK-Vader
if nltk_vader > 0.1:
vader = 1
elif nltk_vader == 0 and nltk_vader <= 0.1:
vader = 0
else: vader = -1
#Averaged return
avg_score = (blob + fin + vader) / 3
return avg_score
#need SpaCy lg dictionary and TextBlob
def review_sent(review):
'''
for a given review, returns dictionary of matched phrases : sentiment
'''
sent_dict = {}
count_df = pd.DataFrame(columns=['phrase','sentiment'])
doc = nlp(review)
for sentence in doc.sents:
matched_phrases = phrase_matcher(nlp(str(sentence)))
phrases = ', '.join(matched_phrases)
#Get Sentiment from models
sentence_sentiment = get_sentiment(sentence)
#check if more than one phrase found by phrase matcher
if len(matched_phrases) > 1:
#add each phrase of the given sentence to the dataframe
for item in matched_phrases:
count_df.loc[-1] = [item, sentence_sentiment]
count_df.index = count_df.index + 1
count_df = count_df.sort_index()
else:
count_df.loc[-1] = [phrases, sentence_sentiment]
count_df.index = count_df.index + 1
count_df = count_df.sort_index()
grouped_df = count_df.groupby('phrase').sum()
grouped_dict = grouped_df.to_dict()
sentimen_dict = grouped_dict['sentiment']
return sentimen_dict
df['sentiment'] = df.review.apply(lambda x: review_sent(x))
df_concat = pd.concat([df, phrase_df], axis=1)
for i in phrase_df.columns:
df_concat[i] = df_concat.apply(lambda x: x.sentiment.get(i), axis=1)
new_df = df_concat.iloc[:, 12:]
df_concat.groupby('stars').sum().iloc[:, 2:]
new_df.head()
fig, ax = plt.subplots(figsize = (14,8))
new_df.count().sort_values().plot(kind='barh', color='orange', alpha=0.7)
plt.title('Total Count of Topics Mentioned in Reviews', fontsize=16)
plt
fig, ax = plt.subplots(figsize = (14,8))
new_df.sum().sort_values().plot(kind='barh', color='orange', alpha=0.7)
plt.title('Feature Sentiment from Charge2 Amazon Reviews', fontsize=16)
plt.xlabel('Count of Negative and Positive Comments', fontsize=12)
sns.set(font_scale=1.05)
fig, ax = plt.subplots(figsize = (20,6))
sns.heatmap(df_concat.groupby('stars').sum().iloc[:, 2:], vmin=-120, vmax=120,
annot=True, linewidths=0.3, cmap='YlGnBu',
cbar_kws={'label': '<--- negative sentiment --- positive sentiment --->'})
ax.invert_yaxis()
plt.title('Net Sentiment Score of Topic by Start Rating', fontsize=16)
plt.xlabel('Topic', fontsize=14)
plt.ylabel('Amazon Star Rating', fontsize=14)
fig.savefig('Net_Sentiment_Scores.png')
```
## Phrase count in new DataFrame (TextBlob only)
```
#create a dataframe with our phrases as columns, to keep count of sentiment
phrase_df = pd.DataFrame(columns=terms)
#need SpaCy lg dictionary and TextBlob
def review_sent(review):
'''
for a given review, returns dictionary of matched phrases : sentiment
'''
sent_dict = {}
count_df = pd.DataFrame(columns=['phrase','sentiment'])
#create textblob object
rev = TextBlob(review)
for sentence in rev.sentences:
matched_phrases = phrase_matcher(nlp(str(sentence)))
phrases = ', '.join(matched_phrases)
if sentence.sentiment[0] > 0.1:
sentence_sentiment = 1
elif sentence.sentiment[0] == 0 and sentence.sentiment[0] <= 0.1:
sentence_sentiment = 0
else:
sentence_sentiment = -1
#check if more than one phrase found by phrase matcher
if len(matched_phrases) > 1:
#add each phrase of the given sentence to the dataframe
for item in matched_phrases:
count_df.loc[-1] = [item, sentence_sentiment]
count_df.index = count_df.index + 1
count_df = count_df.sort_index()
else:
count_df.loc[-1] = [phrases, sentence_sentiment]
count_df.index = count_df.index + 1
count_df = count_df.sort_index()
grouped_df = count_df.groupby('phrase').sum()
grouped_dict = grouped_df.to_dict()
sentimen_dict = grouped_dict['sentiment']
return sentimen_dict
df['sentiment'] = df.review.apply(lambda x: review_sent(x))
df_concat = pd.concat([df, phrase_df], axis=1)
for i in phrase_df.columns:
df_concat[i] = df_concat.apply(lambda x: x.sentiment.get(i), axis=1)
new_df = df_concat.iloc[:, 12:]
new_df.head()
fig, ax = plt.subplots(figsize = (14,8))
new_df.count().sort_values().plot(kind='barh', color='orange', alpha=0.7)
fig, ax = plt.subplots(figsize = (14,8))
new_df.sum().sort_values().plot(kind='barh', color='orange', alpha=0.7)
plt.title('Feature Sentiment from Charge2 Amazon Reviews', fontsize=16)
plt.xlabel('Count of Negative and Positive Comments', fontsize=12)
```
## Phrase count in new DataFrame (AFFIN only)
```
from afinn import Afinn
af = Afinn()
#need SpaCy lg dictionary and AFFIN
def review_sent(review):
'''
for a given review, returns dictionary of matched phrases : sentiment
'''
sent_dict = {}
count_df = pd.DataFrame(columns=['phrase','sentiment'])
#create textblob object
rev = TextBlob(review)
for sentence in rev.sentences:
matched_phrases = phrase_matcher(nlp(str(sentence)))
phrases = ', '.join(matched_phrases)
if af.score(str(sentence)) > 1:
sentence_sentiment = 1
elif af.score(str(sentence)) == 0 and af.score(str(sentence)) <= 1:
sentence_sentiment = 0
else:
sentence_sentiment = -1
#check if more than one phrase found by phrase matcher
if len(matched_phrases) > 1:
#add each phrase of the given sentence to the dataframe
for item in matched_phrases:
count_df.loc[-1] = [item, sentence_sentiment]
count_df.index = count_df.index + 1
count_df = count_df.sort_index()
else:
count_df.loc[-1] = [phrases, sentence_sentiment]
count_df.index = count_df.index + 1
count_df = count_df.sort_index()
grouped_df = count_df.groupby('phrase').sum()
grouped_dict = grouped_df.to_dict()
sentimen_dict = grouped_dict['sentiment']
return sentimen_dict
df['sentiment'] = df.review.apply(lambda x: review_sent(x))
df_concat = pd.concat([df, phrase_df], axis=1)
for i in phrase_df.columns:
df_concat[i] = df_concat.apply(lambda x: x.sentiment.get(i), axis=1)
new_df_afinn = df_concat.iloc[:, 12:]
new_df_afinn.head()
fig, ax = plt.subplots(figsize = (14,8))
new_df_afinn.count().sort_values().plot(kind='barh', color='orange', alpha=0.7)
fig, ax = plt.subplots(figsize = (14,8))
new_df_afinn.sum().sort_values().plot(kind='barh', color='orange', alpha=0.7)
plt.title('Feature Sentiment from Charge2 Amazon Reviews', fontsize=16)
plt.xlabel('Count of Negative and Positive Comments', fontsize=12)
```
## Phrase count in new DataFrame (NLTK only)
```
from nltk.sentiment.vader import SentimentIntensityAnalyzer
import nltk
nltk.download('vader_lexicon')
#need SpaCy lg dictionary
def review_sent(review):
'''
for a given review, returns dictionary of matched phrases : sentiment
'''
sent_dict = {}
count_df = pd.DataFrame(columns=['phrase','sentiment'])
#create textblob object
rev = TextBlob(review)
for sentence in rev.sentences:
matched_phrases = phrase_matcher(nlp(str(sentence)))
phrases = ', '.join(matched_phrases)
if sid.polarity_scores(str(sentence)).get('compound') > 0.1:
sentence_sentiment = 1
elif sid.polarity_scores(str(sentence)).get('compound') == 0 \
and sid.polarity_scores(str(sentence)).get('compound') <= 0.1:
sentence_sentiment = 0
else:
sentence_sentiment = -1
#check if more than one phrase found by phrase matcher
if len(matched_phrases) > 1:
#add each phrase of the given sentence to the dataframe
for item in matched_phrases:
count_df.loc[-1] = [item, sentence_sentiment]
count_df.index = count_df.index + 1
count_df = count_df.sort_index()
else:
count_df.loc[-1] = [phrases, sentence_sentiment]
count_df.index = count_df.index + 1
count_df = count_df.sort_index()
grouped_df = count_df.groupby('phrase').sum()
grouped_dict = grouped_df.to_dict()
sentimen_dict = grouped_dict['sentiment']
return sentimen_dict
df['sentiment'] = df.review.apply(lambda x: review_sent(x))
df_concat = pd.concat([df, phrase_df], axis=1)
for i in phrase_df.columns:
df_concat[i] = df_concat.apply(lambda x: x.sentiment.get(i), axis=1)
new_df_nltk = df_concat.iloc[:, 12:]
new_df_nltk.head()
fig, ax = plt.subplots(figsize = (14,8))
new_df_nltk.count().sort_values().plot(kind='barh', color='orange', alpha=0.7)
fig, ax = plt.subplots(figsize = (14,8))
new_df_nltk.sum().sort_values().plot(kind='barh', color='orange', alpha=0.7)
plt.title('Feature Sentiment from Charge2 Amazon Reviews', fontsize=16)
plt.xlabel('Count of Negative and Positive Comments', fontsize=12)
```
# Sentiment Analyis with StanfordCoreNLP
```
from pycorenlp import StanfordCoreNLP
stan_nlp = StanfordCoreNLP('http://localhost:9000')
text = df.review[19]
print(text)
reviews = text
result = stan_nlp.annotate(text,
properties={
'annotators': 'sentiment, ner, pos',
'outputFormat': 'json',
'timeout': 5000,
})
for s in result["sentences"]:
print("{}: '{}': {} (Sentiment Value) {} (Sentiment)".format(
s["index"],
" ".join([t["word"] for t in s["tokens"]]),
s["sentimentValue"], s["sentiment"]))
sid.polarity_scores('The sleep tracker breaks down sleep by stages which is great .').get('compound')
```
# Training Keras Deep Sentiment Model
```
df['aspect_category'] = df.review.apply(lambda x: sorted(phrase_matcher(nlp(x))))
df.head()
df['aspect_cat_string'] = df.aspect_category.apply(lambda x: ', '.join(x))
df['review'] = df.review.str.lower()
aspect_terms = []
for review in nlp.pipe(df.review):
chunks = [(chunk.root.text) for chunk in review.noun_chunks if chunk.root.pos_ == 'NOUN']
aspect_terms.append(' '.join(chunks))
df['aspect_terms'] = aspect_terms
df.head(5)
from keras.models import load_model
from keras.models import Sequential
from keras.layers import Dense, Activation
aspect_categories_model = Sequential()
aspect_categories_model.add(Dense(512, input_shape=(6000,), activation='relu'))
aspect_categories_model.add(Dense(205, activation='softmax'))
aspect_categories_model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
from keras.preprocessing.text import Tokenizer
vocab_size = 6000 # We set a maximum size for the vocabulary
tokenizer = Tokenizer(num_words=vocab_size)
tokenizer.fit_on_texts(df.review)
aspect_tokenized = pd.DataFrame(tokenizer.texts_to_matrix(df.aspect_terms))
from sklearn.preprocessing import LabelEncoder
from keras.utils import to_categorical
label_encoder = LabelEncoder()
integer_category = label_encoder.fit_transform(df.aspect_cat_string)
dummy_category = to_categorical(integer_category)
aspect_categories_model.fit(aspect_tokenized, dummy_category, epochs=5, verbose=1)
new_review = text
chunks = [(chunk.root.text) for chunk in nlp(new_review).noun_chunks if chunk.root.pos_ == 'NOUN']
new_review_aspect_terms = ' '.join(chunks)
new_review_aspect_tokenized = tokenizer.texts_to_matrix([new_review_aspect_terms])
new_review_category = label_encoder.inverse_transform(aspect_categories_model.predict_classes(new_review_aspect_tokenized))
print(new_review_category)
sentiment_terms = []
for review in nlp.pipe(df['review']):
if review.is_parsed:
sentiment_terms.append(' '.join([token.lemma_ for token in review if (not token.is_stop and not token.is_punct and (token.pos_ == "ADJ" or token.pos_ == "VERB"))]))
else:
sentiment_terms.append('')
df['sentiment_terms'] = sentiment_terms
df.head()
sentiment_model = Sequential()
sentiment_model.add(Dense(512, input_shape=(6000,), activation='relu'))
sentiment_model.add(Dense(3, activation='softmax'))
sentiment_model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
sentiment_tokenized = pd.DataFrame(tokenizer.texts_to_matrix(df.sentiment_terms))
def sentiment_return(rating):
sentiment = 'positive'
if rating == 3:
sentiment = 'neutral'
elif rating < 3:
sentiment = 'negative'
return sentiment
df['sentiment'] = df.stars.apply(lambda x: sentiment_return(x))
df.head()
label_encoder_2 = LabelEncoder()
integer_sentiment = label_encoder_2.fit_transform(df.sentiment)
dummy_sentiment = to_categorical(integer_sentiment)
sentiment_model.fit(sentiment_tokenized, dummy_sentiment, epochs=10, verbose=1)
new_review = 'hate everything, and the watch does not work, sync is bad, sreen is dead'
chunks = [(chunk.root.text) for chunk in nlp(new_review).noun_chunks if chunk.root.pos_ == 'NOUN']
new_review_aspect_terms = ' '.join(chunks)
new_review_aspect_tokenized = tokenizer.texts_to_matrix([new_review_aspect_terms])
new_review_category = label_encoder_2.inverse_transform(sentiment_model.predict_classes(new_review_aspect_tokenized))
print(new_review_category)
test_reviews = [
"customer support they tell me to restart the watch. The problem has not been fixed, i'm returing it. Also, the watch does not sync!",
"Sreen went blank after 3 days. Fitbit stopped syncing after 2 days",
"Customer service just kept telling me to restart the watch, but could not fix my sync problem. I can't return it. awful support."
]
# Aspect preprocessing
test_reviews = [review.lower() for review in test_reviews]
test_aspect_terms = []
for review in nlp.pipe(test_reviews):
chunks = [(chunk.root.text) for chunk in review.noun_chunks if chunk.root.pos_ == 'NOUN']
test_aspect_terms.append(' '.join(chunks))
test_aspect_terms = pd.DataFrame(tokenizer.texts_to_matrix(test_aspect_terms))
# Sentiment preprocessing
test_sentiment_terms = []
for review in nlp.pipe(test_reviews):
if review.is_parsed:
test_sentiment_terms.append(' '.join([token.lemma_ for token in review if (not token.is_stop and not token.is_punct and (token.pos_ == "ADJ" or token.pos_ == "VERB"))]))
else:
test_sentiment_terms.append('')
test_sentiment_terms = pd.DataFrame(tokenizer.texts_to_matrix(test_sentiment_terms))
# Models output
test_aspect_categories = label_encoder.inverse_transform(aspect_categories_model.predict_classes(test_aspect_terms))
test_sentiment = label_encoder_2.inverse_transform(sentiment_model.predict_classes(test_sentiment_terms))
for i in range(3):
print("Review " + str(i+1) + " is expressing a " + test_sentiment[i] + " opinion about " + test_aspect_categories[i])
```
| github_jupyter |
<a href="https://colab.research.google.com/github/satyajitghana/PadhAI-Course/blob/master/11_VectorizedGDAlgorithms.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.colors
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, mean_squared_error, log_loss
from tqdm import tqdm_notebook
import seaborn as sns
import imageio
import time
from IPython.display import HTML
sns.set()
from sklearn.preprocessing import OneHotEncoder
from sklearn.datasets import make_blobs
my_cmap = 'inferno'
np.random.seed(0)
```
# Generate Data
```
data, labels = make_blobs(n_samples=1000, centers=4, n_features=2, random_state=0)
print(data.shape, labels.shape)
plt.scatter(data[:,0], data[:,1], c=labels, cmap=my_cmap)
plt.show()
labels_orig = labels
labels = np.mod(labels_orig, 2)
plt.scatter(data[:,0], data[:,1], c=labels, cmap=my_cmap)
plt.show()
```
# MultiClass Classification
```
X_train, X_val, Y_train, Y_val = train_test_split(data, labels_orig, stratify=labels_orig, random_state=0)
print(X_train.shape, X_val.shape, labels_orig.shape)
enc = OneHotEncoder()
# 0 -> (1, 0, 0, 0), 1 -> (0, 1, 0, 0), 2 -> (0, 0, 1, 0), 3 -> (0, 0, 0, 1)
y_OH_train = enc.fit_transform(np.expand_dims(Y_train,1)).toarray()
y_OH_val = enc.fit_transform(np.expand_dims(Y_val,1)).toarray()
print(y_OH_train.shape, y_OH_val.shape)
W1 = np.random.randn(2,2)
W2 = np.random.randn(2,4)
print(W1)
print(W2)
```
# FF Class
```
class FFNetwork:
def __init__(self, W1, W2):
self.params={}
self.params["W1"]=W1.copy()
self.params["W2"]=W2.copy()
self.params["B1"]=np.zeros((1,2))
self.params["B2"]=np.zeros((1,4))
self.num_layers=2
self.gradients={}
self.update_params={}
self.prev_update_params={}
for i in range(1,self.num_layers+1):
self.update_params["v_w"+str(i)]=0
self.update_params["v_b"+str(i)]=0
self.update_params["m_b"+str(i)]=0
self.update_params["m_w"+str(i)]=0
self.prev_update_params["v_w"+str(i)]=0
self.prev_update_params["v_b"+str(i)]=0
def forward_activation(self, X):
return 1.0/(1.0 + np.exp(-X))
def grad_activation(self, X):
return X*(1-X)
def softmax(self, X):
exps = np.exp(X)
return exps / np.sum(exps, axis=1).reshape(-1,1)
def forward_pass(self, X, params = None):
if params is None:
params = self.params
self.A1 = np.matmul(X, params["W1"]) + params["B1"] # (N, 2) * (2, 2) -> (N, 2)
self.H1 = self.forward_activation(self.A1) # (N, 2)
self.A2 = np.matmul(self.H1, params["W2"]) + params["B2"] # (N, 2) * (2, 4) -> (N, 4)
self.H2 = self.softmax(self.A2) # (N, 4)
return self.H2
def grad(self, X, Y, params = None):
if params is None:
params = self.params
self.forward_pass(X, params)
m = X.shape[0]
self.gradients["dA2"] = self.H2 - Y # (N, 4) - (N, 4) -> (N, 4)
self.gradients["dW2"] = np.matmul(self.H1.T, self.gradients["dA2"]) # (2, N) * (N, 4) -> (2, 4)
self.gradients["dB2"] = np.sum(self.gradients["dA2"], axis=0).reshape(1, -1) # (N, 4) -> (1, 4)
self.gradients["dH1"] = np.matmul(self.gradients["dA2"], params["W2"].T) # (N, 4) * (4, 2) -> (N, 2)
self.gradients["dA1"] = np.multiply(self.gradients["dH1"], self.grad_activation(self.H1)) # (N, 2) .* (N, 2) -> (N, 2)
self.gradients["dW1"] = np.matmul(X.T, self.gradients["dA1"]) # (2, N) * (N, 2) -> (2, 2)
self.gradients["dB1"] = np.sum(self.gradients["dA1"], axis=0).reshape(1, -1) # (N, 2) -> (1, 2)
def fit(self, X, Y, epochs=1, algo= "GD", display_loss=False,
eta=1, mini_batch_size=100, eps=1e-8,
beta=0.9, beta1=0.9, beta2=0.9, gamma=0.9 ):
if display_loss:
loss = {}
for num_epoch in tqdm_notebook(range(epochs), total=epochs, unit="epoch"):
m = X.shape[0]
if algo == "GD":
self.grad(X, Y)
for i in range(1,self.num_layers+1):
self.params["W"+str(i)] -= eta * (self.gradients["dW"+str(i)]/m)
self.params["B"+str(i)] -= eta * (self.gradients["dB"+str(i)]/m)
elif algo == "MiniBatch":
for k in range(0,m,mini_batch_size):
self.grad(X[k:k+mini_batch_size], Y[k:k+mini_batch_size])
for i in range(1,self.num_layers+1):
self.params["W"+str(i)] -= eta * (self.gradients["dW"+str(i)]/mini_batch_size)
self.params["B"+str(i)] -= eta * (self.gradients["dB"+str(i)]/mini_batch_size)
elif algo == "Momentum":
self.grad(X, Y)
for i in range(1,self.num_layers+1):
self.update_params["v_w"+str(i)] = gamma *self.update_params["v_w"+str(i)] + eta * (self.gradients["dW"+str(i)]/m)
self.update_params["v_b"+str(i)] = gamma *self.update_params["v_b"+str(i)] + eta * (self.gradients["dB"+str(i)]/m)
self.params["W"+str(i)] -= self.update_params["v_w"+str(i)]
self.params["B"+str(i)] -= self.update_params["v_b"+str(i)]
elif algo == "NAG":
temp_params = {}
for i in range(1,self.num_layers+1):
self.update_params["v_w"+str(i)]=gamma*self.prev_update_params["v_w"+str(i)]
self.update_params["v_b"+str(i)]=gamma*self.prev_update_params["v_b"+str(i)]
temp_params["W"+str(i)]=self.params["W"+str(i)]-self.update_params["v_w"+str(i)]
temp_params["B"+str(i)]=self.params["B"+str(i)]-self.update_params["v_b"+str(i)]
self.grad(X,Y,temp_params)
for i in range(1,self.num_layers+1):
self.update_params["v_w"+str(i)] = gamma *self.update_params["v_w"+str(i)] + eta * (self.gradients["dW"+str(i)]/m)
self.update_params["v_b"+str(i)] = gamma *self.update_params["v_b"+str(i)] + eta * (self.gradients["dB"+str(i)]/m)
self.params["W"+str(i)] -= eta * (self.update_params["v_w"+str(i)])
self.params["B"+str(i)] -= eta * (self.update_params["v_b"+str(i)])
self.prev_update_params=self.update_params
elif algo == "AdaGrad":
self.grad(X, Y)
for i in range(1,self.num_layers+1):
self.update_params["v_w"+str(i)] += (self.gradients["dW"+str(i)]/m)**2
self.update_params["v_b"+str(i)] += (self.gradients["dB"+str(i)]/m)**2
self.params["W"+str(i)] -= (eta/(np.sqrt(self.update_params["v_w"+str(i)])+eps)) * (self.gradients["dW"+str(i)]/m)
self.params["B"+str(i)] -= (eta/(np.sqrt(self.update_params["v_b"+str(i)])+eps)) * (self.gradients["dB"+str(i)]/m)
elif algo == "RMSProp":
self.grad(X, Y)
for i in range(1,self.num_layers+1):
self.update_params["v_w"+str(i)] = beta*self.update_params["v_w"+str(i)] +(1-beta)*((self.gradients["dW"+str(i)]/m)**2)
self.update_params["v_b"+str(i)] = beta*self.update_params["v_b"+str(i)] +(1-beta)*((self.gradients["dB"+str(i)]/m)**2)
self.params["W"+str(i)] -= (eta/(np.sqrt(self.update_params["v_w"+str(i)]+eps)))*(self.gradients["dW"+str(i)]/m)
self.params["B"+str(i)] -= (eta/(np.sqrt(self.update_params["v_b"+str(i)]+eps)))*(self.gradients["dB"+str(i)]/m)
elif algo == "Adam":
self.grad(X, Y)
num_updates=0
for i in range(1,self.num_layers+1):
num_updates+=1
self.update_params["m_w"+str(i)]=beta1*self.update_params["m_w"+str(i)]+(1-beta1)*(self.gradients["dW"+str(i)]/m)
self.update_params["v_w"+str(i)]=beta2*self.update_params["v_w"+str(i)]+(1-beta2)*((self.gradients["dW"+str(i)]/m)**2)
m_w_hat=self.update_params["m_w"+str(i)]/(1-np.power(beta1,num_updates))
v_w_hat=self.update_params["v_w"+str(i)]/(1-np.power(beta2,num_updates))
self.params["W"+str(i)] -=(eta/np.sqrt(v_w_hat+eps))*m_w_hat
self.update_params["m_b"+str(i)]=beta1*self.update_params["m_b"+str(i)]+(1-beta1)*(self.gradients["dB"+str(i)]/m)
self.update_params["v_b"+str(i)]=beta2*self.update_params["v_b"+str(i)]+(1-beta2)*((self.gradients["dB"+str(i)]/m)**2)
m_b_hat=self.update_params["m_b"+str(i)]/(1-np.power(beta1,num_updates))
v_b_hat=self.update_params["v_b"+str(i)]/(1-np.power(beta2,num_updates))
self.params["B"+str(i)] -=(eta/np.sqrt(v_b_hat+eps))*m_b_hat
if display_loss:
Y_pred = self.predict(X)
loss[num_epoch] = log_loss(np.argmax(Y, axis=1), Y_pred)
if display_loss:
plt.plot(list(loss.values()), '-o', markersize=5)
plt.xlabel('Epochs')
plt.ylabel('Log Loss')
plt.show()
def predict(self, X):
Y_pred = self.forward_pass(X)
return np.array(Y_pred).squeeze()
def print_accuracy():
Y_pred_train = model.predict(X_train)
Y_pred_train = np.argmax(Y_pred_train,1)
Y_pred_val = model.predict(X_val)
Y_pred_val = np.argmax(Y_pred_val,1)
accuracy_train = accuracy_score(Y_pred_train, Y_train)
accuracy_val = accuracy_score(Y_pred_val, Y_val)
print("Training accuracy", round(accuracy_train, 4))
print("Validation accuracy", round(accuracy_val, 4))
if False:
plt.scatter(X_train[:,0], X_train[:,1], c=Y_pred_train, cmap=my_cmap, s=15*(np.abs(np.sign(Y_pred_train-Y_train))+.1))
plt.show()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=1, algo="GD", display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=1, algo="MiniBatch", mini_batch_size=128, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=1, algo="MiniBatch", mini_batch_size=8, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=1, algo="Momentum", gamma=0.5, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=1, algo="Momentum", gamma=0.9, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=1, algo="Momentum", gamma=0.99, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=1, algo="NAG", gamma=0.99, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=1, algo="NAG", gamma=0.5, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=1, algo="NAG", gamma=0.9, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=1, algo="AdaGrad", display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=.1, algo="AdaGrad", display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=.1, algo="RMSProp", beta=0.9, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=.9, algo="RMSProp", beta=0.9, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=.9, algo="Adam", beta=0.9, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=100, eta=.1, algo="Adam", beta=0.9, display_loss=True)
print_accuracy()
```
# Good Configuration for each Algorithm
```
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=10000, eta=0.5, algo="GD", display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=1000, eta=0.5, algo="Momentum", gamma=0.9, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=1000, eta=0.5, algo="NAG", gamma=0.9, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=500, eta=1, algo="AdaGrad", display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=2000, eta=.01, algo="RMSProp", beta=0.9, display_loss=True)
print_accuracy()
%%time
model = FFNetwork(W1, W2)
model.fit(X_train, y_OH_train, epochs=200, eta=.1, algo="Adam", beta=0.9, display_loss=True)
print_accuracy()
```
| github_jupyter |
**Example**
```
(define (sum term a next b)
(if (> a b)
0
(+ (term a)
(sum term (next a) next b))))
(define (inc n) (+ n 1))
(define (cube n) (* n n n))
(define (sum-cube a b)
(sum cube a inc b))
(sum-cube 1 10)
(define (identify x) x)
(define (sum-integers a b)
(sum identify a inc b))
(sum-integers 1 10)
(define (pi-sum a b)
(define (pi-term x)
(/ 1.0 (* x (+ x 2))))
(define (pi-next x)
(+ x 4))
(sum pi-term a pi-next b))
(* 8 (pi-sum 1 10000))
(define (integral f a b dx)
(define (add-dx x) (+ x dx))
(* (sum f (+ a (/ dx 2.0)) add-dx b) dx ))
(integral cube 0 1 0.001)
```
**Exericse 1.29**
```
(define (round-to-next-even x)
(+ x (remainder x 2)))
(define (simpson f a b n)
(define fixed-n (round-to-next-even n))
(define (even? x) (= (remainder x 2) 0))
(define (inc x)(+ x 1))
(define h (/ (- b a) fixed-n))
(define (simpson-term k)
(define y (f (+ a (* k h))))
(if (or (= k 0) (= k fixed-n))
(* 1 y)
(if (even? k)
(* 2 y)
(* 4 y))))
(* (/ h 3) (sum simpson-term 0 inc fixed-n)))
(simpson cube 0 1 10)
```
**Exercise 1.30**
```
(define (sum term a next b)
(define (iter a result)
(if (> a b)
result
(iter (next a) (+ (term a) result))))
(iter a 0))
```
**Exercise 1.31(a)**
```
(define (product term a next b)
(if (> a b)
1
(* (term a) (product term (next a) next b))))
(define (pi-term x)
(/ (* (- x 1) (+ x 1) 1.0) (* x x)))
(define (pi-next n) (+ n 2))
(define (product-pi n)
(* 4 (product pi-term 3 pi-next (+ (* 2 n) 1))))
(product-pi 100)
```
**Exercise 1.31(b)**
```
(define (product term a next b)
(define (iter a result)
(if (> a b)
result
(iter (next a) (* (term a) result))))
(iter a 1))
(define (pi-term x)
(/ (* (- x 1) (+ x 1) 1.0) (* x x)))
(define (pi-next n) (+ n 2))
(define (product-pi n)
(* 4 (product pi-term 3 pi-next (+ (* 2 n) 1))))
(product-pi 100)
```
**Exercise 1.32(a)**
```
(define (accumulate combiner null-value term a next b)
(if (> a b)
null-value
(combiner (term a) (accumulate combiner null-value term (next a) next b))))
(define (sum term a next b)
(define (add a b)(+ a b))
(accumulate add 0 term a next b))
(define (product term a next b)
(define (multiply a b)(* a b))
(accumulate multiply 1 term a next b))
```
**Exercise 1.32(b)**
```
(define (accumulate combiner null-value term a next b)
(define (iter a result)
(if (> a b)
result
(iter (next a) (combiner (term a) result))))
(iter a null-value))
```
**Exercise 1.33(a)**
```
(define (filtered-accumulate filter combiner null-value term a next b)
(define (predict x)
(if (filter x)
x
null-value))
(if (> a b)
null-value
(combiner (filter (term a)) (filtered-accumulate filter combiner null-value term (next a) next b ))))
(define (even? x)(= (remainder x 2) 0))
(define (add a b)(+ a b))
(define (identify x) x )
(define (inc x)(+ x 1))
(filtered-accumulate even? add 0 identify 1 inc 8)
(define (prime-sum a b)
(define (prime? n)
(define (smallest-divisor n)
(find-divisor n 2))
(define (find-divisor n test-divisor)
(cond ((> (square test-divisor) n) n)
((divides? test-divisor n) test-divisor)
(else (find-divisor n (+ test-divisor 1)))))
(define (divides? a b)
(= (remainder b a) 0))
(define (square n)(* n n))
(= n (smallest-divisor n)))
(define (add a b)(+ a b))
(define (identify x) x)
(define (inc x)(+ x 1))
(filtered-accumulate prime? add 0 identify a inc b))
(prime-sum 0 10)
```
**Exercise 1.33(b)**
```
(define (gcd a b)
(if (= b 0)
a
(gcd b (remainder a b))))
(define (gcd-product n)
(define (relative-prime? x)
(= (gcd n x) 1))
(define (product a b)(* a b))
(define (identify x) x)
(define (inc x)(+ x 1))
(filtered-accumulate relative-prime? product 1 identify 1 inc n))
(gcd-product 10)
```
**Example**
```
(lambda x (+ x 4))
(define (pi-sum a b)
(sum (lambda (x) (/ 1.0 (* x (+ x 2))))
a
(lambda (x) (+ x 4))
b))
(pi-sum 1 10)
```
**Example**
$$f(x,y)=x(1+xy)^{2}+y(1-y)+(1+xy)(1-y)$$
$a=1+xy$
$b=1- y$
$$f(x, y)=xa^2+yb+ab$$
```
(define (f x y)
(define (square n)(* n n))
(let ((a (+ 1 (* x y)))
(b (- 1 y)))
(+ (* x (square a))
(* y b)
(* a b))))
(f 2 4)
```
**Exercise 1.34**
```
(define (f g)
(g 2))
(define (square x)(* x x))
(f square)
(f (lambda (z)(* z (+ z 1))))
```
**Example**
```
(define (search f neg-point pos-point)
(let ((midpoint (average neg-point pos-point)))
(if (close-enough? neg-point pos-point)
midpoint
(let ((test-value (f midpoint)))
(cond ((positive? test-value)
(search f neg-point midpoint))
((negative? test-value)
(search f midpoint pos-point))
(else midpoint))))))
(define (average a b)(/ (+ a b) 2.0))
(define (close-enough? x y)
(< (abs(- x y)) 0.001))
(define (positive? a)(> a 0))
(define (negative? a)(< a 0))
(search (lambda (x)(* x x x)) -2 1)
(define (half-interval-method f a b)
(let ((a-value (f a))
(b-value (f b)))
(cond ((and (negative? a-value)(positive? b-value))
(search f a b))
((and (negative? b-value)(positive? a-value))
(search f b a))
(else
(error "Values are not of opposite sign" a b)))))
```
**Example**
```
(define tolerance 0.00001)
(define (fixed-point f first-guess)
(define (close-enough? v1 v2)
(< (abs(- v1 v2)) tolerance))
(define (try guess)
(let ((next (f guess)))
(if (close-enough? guess next)
next
(try next))))
(try first-guess))
(define (sqrt x)
(define (average a b)(/ (+ a b) 2.0))
(fixed-point (lambda (y) (average y (/ x y)))
2.0))
```
**Exercise 1.35**
```
(define gold-ratio
(fixed-point (lambda (y) (+ 1 (/ 1.0 y))) 0.5))
```
**Exercise 1.37 (a)**
```
(define (cont-frac fn fd k)
(define (cont-frac-iter fn fd k result)
(if (= k 0)
result
(cont-frac-iter fn fd (- k 1) (/ (fn (- k 1)) (+ (fd (- k 1)) result)))))
(cont-frac-iter fn fd k 0.0))
(cont-frac (lambda (i) 1.0) (lambda (i) 1.0) 100)
```
**Exercise 1.37(b)**
```
(define (cont-frac fn fd k)
(define (cont-frac-rec fn fd i)
(if (= i k)
(/ (fn i)(fd i))
(/ (fn i)(+ (fd i) (cont-frac-rec fn fd (+ i 1))))))
(cont-frac-rec fn fd 1))
(cont-frac (lambda (i) 1.0) (lambda (i) 1.0) 100)
```
**Exercise 1.38**
```
(define (fn i) 1)
(define (fd i)
(cond ((= i 1) 1)
((= i 2) 2)
((or (= (remainder (- i 2) 3) 1) (= (remainder (- i 2) 3) 2)) 1)
((+ (* (/ 2.0 3)(- i 2)) 2))))
(define (natural n)
(+ (cont-frac fn fd n) 2))
(natural 1000)
```
**Exercise 1.39**
```
(define (tan-cf x k)
(define (tan-fc-iter x i result)
(if (= i 1)
(/ x (- 1 result))
(tan-fc-iter x (- i 1) (/ (* x x)(- (* i 2) 1)))))
(tan-fc-iter x k 0.0))
(tan-cf 1.0 300)
```
**Example**
```
(define (average-damp f)
(define (average a b)(/ (+ a b) 2))
(lambda (x) (average x (f x))))
((average-damp (lambda (i) (* i i))) 10)
```
**Example**
```
(define (deriv g)
(lambda (x)(/ (- (g (+ x dx)) (g x)) dx)))
(define dx 0.0001)
(define (cube x)(* x x x))
((deriv cube) 5)
```
**Exercise1.41**
```
(define (double f)
(lambda (x) (f (f x))))
(define (inc x) (+ x 1))
(((double (double double)) inc) 5)
```
**Exercise 1.42**
```
(define (compose f g)
(lambda (x) (f (g x))))
(define (square x)(* x x))
(define (inc x)(+ x 1))
((compose square inc) 6)
```
**Exercise 1.43**
```
(define (repeated f n)
(if (= n 1)
(lambda (x) (f x))
(lambda (x) (f ((repeated f (- n 1)) x)))))
(define (square x)(* x x))
((repeated square 2) 5)
```
**Exericse 1.44**
```
(define dx 0.00001)
(define (smooth f)
(lambda (x)
(/ (+ (f (- x dx))
(f x)
(f (+ x dx)))
3)))
(define (n-fold-smooth f n)
((repeated smooth n) f))
```
| github_jupyter |
```
"""
Set up yagmail: https://towardsdatascience.com/automate-sending-emails-with-gmail-in-python-449cc0c3c317
Set up auto schedule: https://towardsdatascience.com/10-step-guide-to-schedule-your-script-using-cloud-services-7f4a8e517d81
Enjoy!
"""
!pip3 install yagmail
from urllib.request import urlopen
import yagmail
user = 'your_id'
app_password = 'generate_your_app_passwrd'
to = 'sender\'s_address'
try:
def jobNotiSSC(i):
dateList = []
contentList = []
urlList = []
data = ""
j = 1
url = "https://ssc.nic.in/"
page = urlopen(url)
html_bytes = page.read()
html = html_bytes.decode("utf-8")
firstIndex = html.find("<div class=\"leftsideNotifications_New\">")
firstIndex = firstIndex + \
len("<div class=\"leftsideNotifications_New\">")
while i > 0:
firstIndex = html.find(
"<div class=\"eachNotification \">", firstIndex)
firstIndex = firstIndex+len("<div class=\"eachNotification \">")
firstIndex = html.find("<span>", firstIndex)
firstIndex = firstIndex+len("<span>")
lastIndex = html.find("</span>", firstIndex)
# Date is fetched here
date = html[firstIndex:lastIndex]
date = date.split("<i>")
date = " ".join(date)
date = date.split("</i>")
date = " ".join(date)
# Linked document URL is fetched here.
firstIndex = html.find("<a href=", lastIndex)
firstIndex = firstIndex+len("<a href=\"")
lastIndex = html.find("target=\"_blank\">", firstIndex)
urlDoc = html[firstIndex:lastIndex].strip()
urlDoc = urlDoc.replace(" ", "%20")
urlDoc = urlDoc.replace("\"", "")
# Content will be fetched here.
firstIndex = lastIndex+len("target=\"_blank\">")
lastIndex = html.find("</a>", firstIndex)
content = html[firstIndex:lastIndex].strip()
dateList.append(date)
contentList.append(content)
urlList.append(urlDoc)
j = j+1
i = i-1
return dateList, urlList, contentList
def html_builder():
header = """\
<!DOCTYPE html PUBLIC "-//W3C//DTD XHTML 1.0 Transitional //EN" "http://www.w3.org/TR/xhtml1/DTD/xhtml1-transitional.dtd">
<html>
<head>
<meta http-equiv="Content-Type" content="text/html; charset=UTF-8" />
<meta name="viewport" content="width=device-width, initial-scale=1.0" />
<meta name="x-apple-disable-message-reformatting" />
<!--[if !mso]><!-->
<meta http-equiv="X-UA-Compatible" content="IE=edge" />
<!--<![endif]-->
</head>
<body>
<table>
<tr>
<th>Sr. No.</th>
<th>Date</th>
<th>Link</th>
<th>Content</th>
</tr>"""
body = '''<tr>
<td style="padding:20px"><?a?></td>
<td style="padding:20px"><?b?></td>
<td style="padding:20px"><a href="<?c?>" target="_blank">Link</a></td>
<td style="padding:20px"><b><?d?></b></td>
</tr>'''
footer = """</table>
<p><a href="https://ssc.nic.in/">SSC Online</a></p>
</body>
</html>"""
new_str = ''
for i in range(no_of_data):
temp = body
str1 = "dateList["+str(i)+"]"
str2 = "urlList["+str(i)+"]"
str3 = "contentList["+str(i)+"]"
temp = temp.replace("<?a?>", str(i+1))
temp = temp.replace("<?b?>", str1)
temp = temp.replace("<?c?>", str2)
temp = temp.replace("<?d?>", str3)
new_str += temp
content = header + new_str + footer
for i in range(no_of_data):
str1 = "dateList["+str(i)+"]"
str2 = "urlList["+str(i)+"]"
str3 = "contentList["+str(i)+"]"
content = content.replace(str1, str(dateList[i]))
content = content.replace(str2, str(urlList[i]))
content = content.replace(str3, str(contentList[i]))
return content
"""main"""
no_of_data = 3
dateList, urlList, contentList = jobNotiSSC(no_of_data)
subject = 'Job Alert'
content = html_builder()
# print(content)
with yagmail.SMTP(user, app_password) as yag:
yag.send(to, subject, content)
print('Sent email successfully')
except:
print("Error occured")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mottaquikarim/PYTH2/blob/master/src/Topics/nb/basic_data_types.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Basic Data Types
Let's discuss data types, variables, and naming.
A data type is a unit of information that can be stored and retrieved using a program language. **Variables** store different types of data. You can display the value stored in a variable on the screen using the `print()` statement. Because variables are so ubiquitous, there are some **rules for naming variables** that help avoid confusion.
- **Rule:** snake_case with the underscore character
- **Rule:** Can’t use keywords as var names, e.g. True, None, function, class, object
- **Rule:** Can’t start with a numeric value, e.g. can't name something "1var", but you CAN do function1
- **Best Practice:** You COULD use CamelCase to name a variable, but this is typically reserved for naming a different Python object called a "class".
#### Creating & Reading a Variable
```
first_prime = 2
print(first_prime) # expect to see 2
```
#### Comments
Real quick, let's see how we create comments so that you can write notes to yourself in your code - **very** useful!
```
# I'm a comment!
# Here is a multi-line comment:
"""
'Cause if you liked it, then you should have put a ring on it
If you liked it, then you should have put a ring on it
Don't be mad once you see that he want it
If you liked it, then you should have put a ring on it
"""
```
These are the most basic data types, which compose the more complex data types we'll learn about later:
* Strings: alphanumeric characters read as text
* **Important takeaway!** If you use a `'`, say in a contraction, the string might misinterpret it as the ending `'` for the string. To avoid this, add `\` right before that `'` like this: 'I\'ll`.
* Integers: whole numbers
* Floats: decimals
* Booleans: represents True or False values; used for evaluating conditional statements
* Nonetype: represents variables not yet defined or essentially blank values (shown as None, used as placeholder to avoid error)
```
# Strings
cat = 'Layla' # or "Layla" because single & double quotes are both valid
cat_description = 'Layla\'s a mischievous, but very cute kitty!' # Note the use of \' here.
# Integers
age = 8
# Floats
weight = 10.5
# Booleans
vaccinated = True
good_with_other_cats = False
# Nonetype
good_with_other_dogs = None
```
### String Formatting
Strings have some special qualities that make dealing with them easier. For instance, you can store several separate snippets of text within a single string. One way to do this is adding `\n` to signify that you want to create a new line. However, there's another way to do this that makes it much more readable! To do this, you can use triple quotes i.e. `'''` or `"""`. You can use single and double quotes within the string freely, so no need to worry about that detail!
Let's say you're storing song lyrics, so you want to have a line break between each line of the song.
```
'Cause if you liked it, then you should have put a ring on it\nIf you liked it, then you should have put a ring on it\nDon\'t be mad once you see that he want it\nIf you liked it, then you should have put a ring on it'
'''
'Cause if you liked it, then you should have put a ring on it
If you liked it, then you should have put a ring on it
Don't be mad once you see that he want it
If you liked it, then you should have put a ring on it
'''
```
If you want to insert variables within a string, you can do this:
```
cat = 'Layla'
print(cat, 'is a mischievous, but very cute kitty!')
# Note that the ',' automatically adds a space character.
```
But that can be annoying if you're inserting multiple variables. To simplify this, you dynamically add variables directly within a string like this:
```
cat = 'Layla' # or "Layla" because single & double quotes are both valid
age = 8
weight = 10.5
print(f'{cat} is {age} years old.\n{cat} weighs {weight} pounds.')
print(f'''
{cat} is {age} years old.
{cat} weighs {weight} pounds.
''')
"""
Both of these print...
Layla is 8 years old.
Layla weighs 10.5 pounds.
"""
```
**Pro Tip**: You can use several versions of the the `.strip()` function to remove leading and trailing spaces from strings.
* `lstrip()`: remove all leading spaces
* `rstrip()`: remove all trailing spaces
* `strip()`: remove all leading AND trailing spaces
```
a = ' hi mom!'
print(f'LEADING SPACES: \n{a} \n{a.strip()}\n\n')
b = 'hi mom! '
print(f'TRAILING SPACES: \n{b} \n{b.strip()}\n\n')
c = ' hi mom! '
# print(c, '\n', c.strip(), '\n\n')
print(f'LEADING & TRAILING SPACES: \n{c} \n{c.strip()}')
```
## Typecasting
**Typecasting** is a way to convert an instance of one data type to another. Numbers are the most flexible for converting, while strings are very inflexible. Booleans get a bit more complicated, so we'll look at those last!
First you need to know what data type your variable is in the first place! Check what data type is stored in a variable using the `type()` statement or the `isinstance()` statement.
```
# Example 1 - type()
a = 1
print(type(a)) # <class 'int'>
b = '2.5'
print(type(b)) # <class 'str'>
# Example 2 - isinstance()
c = -1
print(f'Is {c} a boolean?', isinstance(c, bool)) # False
d = False
print(f'Is {d} a boolean?', isinstance(d, bool)) # True
```
### Convert Data Types
```
"""INTEGERS"""
int_to_float = float(10) # 10.0, <class 'float'>
int_to_string = str(10) # 10, <class 'str'>
"""FLOATS"""
float_to_int = int(2.5) # 2, <class 'float'>
print(float_to_int)
# Notice it does NOT ROUND!
float_to_string = str(2.5) # '2.5', <class 'str'>
```
#### Converting Strings
```
string_to_int = int('mango') # ERROR!
string_to_float = str('strawberry') # ERROR!
```
#### Converting Booleans
As you'll see below, you can convert any number to a True boolean, BUT a True boolean will only ever become 1 or 1.0.
```
"""EVALS TO TRUE"""
int_to_boolean = bool(10) # True, <class 'bool'>
int_to_boolean = bool(0) # False
float_to_boolean = print(bool(-1)) # True, <class 'bool'>
string_to_boolean = bool('peach') # True, <class 'bool'>
"""EVALS TO FALSE"""
int_to_boolean = bool(0) # False, <class 'bool'>
float_to_boolean = bool(0.0) # False, <class 'bool'>
string_to_boolean = print(bool(' ')) # False, <class 'bool'>
```
Notice that the **ONLY** way a string converted into a boolean will be False is if it's empty. Spaces count as characters even though they wouldn't display anything if printed.
## Simple Integer, Float, & String Operators
**Operators** are shortcuts for manipulating values stored in variables.
### Integer/Float Operators
We can operate on integers/floats in the following ways:
* Addition
* Subtraction
* Multiplication
* Division
* Modulus (This one divides and returns **only the remainder**.)
```
orig_num = 10
# Addition
num1 = orig_num + 5 # 15
# Subtraction
num2 = orig_num - 5 # 5
# Multiplication
num3 = orig_num * 5 # 50
# Division
num4 = orig_num / 5 # 2
# Modulus
num5 = orig_num % 5 # 0
num6 = orig_num % 3 # 1
# if num1 % 3 == 0, then it's even
#
```
### String Operators
* We can "add" strings
* We CANNOT add strings to non strings
```
a = 'this string'
b = 'that string'
print(a + b) # 'this stringthat string'
print(a + ' and ' + b) # 'this string and that string'
print(a, 'and', b) # 'this string and that string'
"""ERROR!!!
print('this will not work' + 4) doesn't work
because you can't add a number to a string"""
# quant = 4
# product = 'bananas'
# how_much = quant + product # '4 bananas'
```
## Class Practice PSETs
[Basic Data Types](https://colab.research.google.com/github/mottaquikarim/PYTH2/blob/master/src/PSETS/nb/basic_data_inclass_psets.ipynb)
## Additional Resources
* [A Repl.it Summarizing Print Statements](https://repl.it/@brandiw/Python-01-Variables-4?lite=true)
* [Python For Beginners](http://www.pythonforbeginners.com/basics/python-variables)
* [Python Programming Tutorial: Variables](https://www.youtube.com/watch?v=vKqVnr0BEJQ)
* [Variables in Python](https://www.guru99.com/variables-in-python.html)
* [Operators Cheatsheet](http://python-reference.readthedocs.io/en/latest/docs/operators/)
* [Python Style Guide: Naming](https://www.python.org/dev/peps/pep-0008/#descriptive-naming-styles)
* [Python-Strings](https://www.tutorialspoint.com/python/python_strings.htm)
* [String Concatenation and Formatting](http://www.pythonforbeginners.com/concatenation/string-concatenation-and-formatting-in-python)
* [String Concatenation and Formatting - Video](https://www.youtube.com/watch?v=jA5LW3bR0Us)
| github_jupyter |
# Generate a Noise Model using Calibration Data
We will use pairs of noisy calibration observations $x_i$ and clean signal $s_i$ (created by averaging these noisy, calibration images) to estimate the conditional distribution $p(x_i|s_i)$. Histogram-based and Gaussian Mixture Model-based noise models are generated and saved.
__Note:__ Noise model can also be generated if calibration data is not available. In such a case, we use an approach called ```Bootstrapping```. Take a look at the notebook ```0b-CreateNoiseModel (With Bootstrapping)``` on how to do so. To understand more about the ```Bootstrapping``` procedure, take a look at the readme [here](https://github.com/juglab/PPN2V).
```
import warnings
warnings.filterwarnings('ignore')
# import torch
import os
import urllib
import zipfile
from torch.distributions import normal
import matplotlib.pyplot as plt, numpy as np, pickle
from scipy.stats import norm
from tifffile import imread
import sys
sys.path.append('../../')
from divnoising.gaussianMixtureNoiseModel import GaussianMixtureNoiseModel
from divnoising import histNoiseModel
from divnoising.utils import plotProbabilityDistribution
dtype = torch.float
device = torch.device("cuda:0")
```
### Download data
Download the data from https://zenodo.org/record/5156913/files/Convallaria_diaphragm.zip?download=1. Here we show the pipeline for Convallaria dataset. Save the dataset in an appropriate path. For us, the path is the data folder which exists at `./data`.
```
# Download data
if not os.path.isdir('./data'):
os.mkdir('./data')
zipPath="./data/Convallaria_diaphragm.zip"
if not os.path.exists(zipPath):
data = urllib.request.urlretrieve('https://zenodo.org/record/5156913/files/Convallaria_diaphragm.zip?download=1', zipPath)
with zipfile.ZipFile(zipPath, 'r') as zip_ref:
zip_ref.extractall("./data")
```
The noise model is a characteristic of your camera. The downloaded data folder contains a set of calibration images (For the Convallaria dataset, it is ```20190726_tl_50um_500msec_wf_130EM_FD.tif``` showing microscope diaphragm and the data to be denoised is named ```20190520_tl_25um_50msec_05pc_488_130EM_Conv.tif```). We can either bin the noisy - GT pairs (obtained from noisy calibration images) as a 2-D histogram or fit a GMM distribution to obtain a smooth, parametric description of the noise model.
Specify ```path``` where the noisy calibration data will be loaded from. It is the same path where noise model will be stored when created later, ```dataName``` is the name you wish to have for the noise model, ```n_gaussian``` to indicate how many Gaussians willbe used for learning a GMM based noise model, ```n_coeff``` for indicating number of polynomial coefficients will be used to patrametrize the mean, standard deviation and weight of GMM noise model. The default settings for ```n_gaussian``` and ```n_coeff``` generally work well for most datasets.
```
path="./data/Convallaria_diaphragm/"
observation= imread(path+'20190726_tl_50um_500msec_wf_130EM_FD.tif') # Load the appropriate calibration data
dataName = 'convallaria' # Name of the noise model
n_gaussian = 3 # Number of gaussians to use for Gaussian Mixture Model
n_coeff = 2 # No. of polynomial coefficients for parameterizing the mean, standard deviation and weight of Gaussian components.
nameHistNoiseModel ='HistNoiseModel_'+dataName+'_'+'calibration'
nameGMMNoiseModel = 'GMMNoiseModel_'+dataName+'_'+str(n_gaussian)+'_'+str(n_coeff)+'_'+'calibration'
# The data contains 100 images of a static sample (microscope diaphragm).
# We estimate the clean signal by averaging all images.
signal=np.mean(observation[:, ...],axis=0)[np.newaxis,...]
# Let's look the raw data and our pseudo ground truth signal
print(signal.shape)
plt.figure(figsize=(12, 12))
plt.subplot(1, 2, 2)
plt.title(label='average (ground truth)')
plt.imshow(signal[0],cmap='gray')
plt.subplot(1, 2, 1)
plt.title(label='single raw image')
plt.imshow(observation[0],cmap='gray')
plt.show()
```
### Creating the Histogram Noise Model
Using the raw pixels $x_i$, and our averaged GT $s_i$, we are now learning a histogram based noise model. It describes the distribution $p(x_i|s_i)$ for each $s_i$.
```
# We set the range of values we want to cover with our model.
# The pixel intensities in the images you want to denoise have to lie within this range.
minVal, maxVal = 234, 7402
bins = 256
# We are creating the histogram.
# This can take a minute.
histogram = histNoiseModel.createHistogram(bins, minVal, maxVal, observation,signal)
# Saving histogram to disc.
np.save(path+nameHistNoiseModel+'.npy', histogram)
histogramFD=histogram[0]
# Let's look at the histogram-based noise model.
plt.xlabel('Observation Bin')
plt.ylabel('Signal Bin')
plt.imshow(histogramFD**0.25, cmap='gray')
plt.show()
```
### Creating the GMM noise model
Using the raw pixels $x_i$, and our averaged GT $s_i$, we are now learning a GMM based noise model. It describes the distribution $p(x_i|s_i)$ for each $s_i$.
```
min_signal=np.min(signal)
max_signal=np.max(signal)
print("Minimum Signal Intensity is", min_signal)
print("Maximum Signal Intensity is", max_signal)
```
Iterating the noise model training for `n_epoch=2000` and `batchSize=250000` works the best for `Convallaria` dataset.
```
gaussianMixtureNoiseModel = GaussianMixtureNoiseModel(min_signal = min_signal,
max_signal =max_signal,
path=path, weight = None,
n_gaussian = n_gaussian,
n_coeff = n_coeff,
min_sigma = 50,
device = device)
gaussianMixtureNoiseModel.train(signal, observation, batchSize = 250000, n_epochs = 2000,
learning_rate=0.1, name = nameGMMNoiseModel)
```
### Visualizing the Histogram-based and GMM-based noise models
```
plotProbabilityDistribution(signalBinIndex=25, histogram=histogramFD,
gaussianMixtureNoiseModel=gaussianMixtureNoiseModel, min_signal=minVal,
max_signal=maxVal, n_bin= bins, device=device)
```
| github_jupyter |
# CT-LTI: Multiple Sample Performance Evaluation Table
This table is found in the appendix section A.4. and summarizes the performance comparison between NODEC and OC in relative terms of error and energy. Without extensive hyperparameter optimization we see that NODEC is competitive to OC for all graphs and intial-target state settings.
Furthermore, please make sure that the required data folder is available at the paths used by the script.
You may generate the required data by running the python script
```nodec_experiments/ct_lti/gen_parameters.py```.
Please also make sure that a trainingproceedure has produced results in the corresponding paths used below.
Running ```nodec_experiments/ct_lti/multi_sample/train.ipynb``` with default paths is expected to generate at the requiered location.
As neural network intialization is stochastic, please make sure that appropriate seeds are used or expect some variance to paper results.
```
%load_ext autoreload
%autoreload 2
import numpy as np
import pandas as pd
```
## Gather data from files
Below we gather the data from files generated by the ```train_and_eval.ipynb``` file. Please run this first if the data files are not present!
```
data_folder = '../../../../data/results/ct_lti/multi_sample/'
graphs = ['lattice', 'ba', 'tree']
graph_name = {'lattice' : 'Square Lattice', 'ba' : 'Barabasi Albert', 'tree' : 'Random Tree'}
resulting_rows = []
for graph in graphs:
graph_folder = data_folder + graph + '/'
interactions = [50, 500, 5000]
for interaction in interactions:
mse_diffs = []
energy_diffs = []
for i in range(100):
nnres = pd.read_csv(graph_folder+'nn_sample_'+str(i)+'_train_'+str(interaction)+'/epoch_metadata.csv')
ocres = pd.read_csv(graph_folder+'oc_sample'+str(i)+'_ninter_'+str(interaction)+'/epoch_metadata.csv')
nn_en = nnres['total_energy'].item()
oc_en = ocres['total_energy'].item()
nn_fl = nnres['final_loss'].item()
oc_fl = ocres['final_loss'].item()
mse_diffs.append((nn_fl-oc_fl)/oc_fl)
energy_diffs.append((nn_en-oc_en)/oc_en)
row = {'Graph' : graph_name[graph], 'Interaction Interval': 5.0/interaction,
'Median Energy' : round(np.quantile(energy_diffs, 0.5), 2),
'IQR Energy' : round(np.quantile(energy_diffs, 0.75)-np.quantile(energy_diffs,0.25), 2),
'Median MSE' : round(np.quantile(mse_diffs, 0.5), 2),
'IQR MSE' : round(np.quantile(mse_diffs, 0.75)-np.quantile(mse_diffs, 0.25), 2),
'Numerical Instabilities' : round((np.array(mse_diffs) > 10).mean(), 2)
}
resulting_rows.append(row)
```
## Resulting Table
```
df = pd.DataFrame(resulting_rows).groupby(['Graph', 'Interaction Interval']).first()
styler = df.style.apply(lambda x: ["background: lightblue" if v <= 0.1 and i in [0,2] else "" for i,v in enumerate(x)], axis = 1)
styler
```
| github_jupyter |
___
<a href='http://www.pieriandata.com'><img src='../Pierian_Data_Logo.png'/></a>
___
<center><em>Copyright Pierian Data</em></center>
<center><em>For more information, visit us at <a href='http://www.pieriandata.com'>www.pieriandata.com</a></em></center>
# Pandas Exercises
Time to test your new pandas skills! Use the <tt>population_by_county.csv</tt> file in the Data folder to complete the tasks in bold below!
<div class="alert alert-info" style="margin: 10px"><strong>NOTE:</strong> ALL TASKS CAN BE DONE IN ONE LINE OF PANDAS CODE. GET STUCK? NO PROBLEM! CHECK OUT THE SOLUTIONS LECTURE!</div>
<div class="alert alert-danger" style="margin: 10px"><strong>IMPORTANT NOTE!</strong> Make sure you don't run the cells directly above the example output shown, <br>otherwise you will end up writing over the example output!</div>
#### 1. Import pandas and read in the <tt>population_by_county.csv</tt> file into a dataframe called <tt>pop</tt>.
```
import pandas as pd
pop = pd.read_csv("population_by_county.csv")
```
#### 2. Show the head of the dataframe
```
# CODE HERE
pop.head()
# DON'T WRITE HERE
```
#### 3. What are the column names?
```
pop.columns
# DON'T WRITE HERE
```
<strong>4. How many States are represented in this data set?</strong> <em>Note: the data includes the District of Columbia</em>
```
pop["State"].nunique()
# DON'T WRITE HERE
```
#### 5. Get a list or array of all the states in the data set.
```
pop.State.unique()
# DON'T WRITE HERE
```
#### 6. What are the five most common County names in the U.S.?
```
pop['County'].value_counts()[:5]
# DON'T WRITE HERE
```
#### 7. What are the top 5 most populated Counties according to the 2010 Census?
```
pop.sort_values(["2010Census"], ascending=False)[:5]
# DON'T WRITE HERE
```
#### 8. What are the top 5 most populated States according to the 2010 Census?
```
pop.groupby("State").sum().sort_values(["2010Census"], ascending=False)[:5]
# DON'T WRITE HERE
```
#### 9. How many Counties have 2010 populations greater than 1 million?
```
pop["County"][pop["2010Census"] > 1000000].nunique()
# DON'T WRITE HERE
```
#### 10. How many Counties don't have the word "County" in their name?
```
len(set([x for x in pop["County"] if "County" not in x]))
# DON'T WRITE HERE
```
#### 11. Add a column that calculates the percent change between the 2010 Census and the 2017 Population Estimate
```
# CODE HERE
pop["PercentChange"] = (pop["2017PopEstimate"] - pop["2010Census"]) / pop["2010Census"] * 100
# USE THIS TO SHOW THE RESULT
pop.head()
# DON'T WRITE HERE
```
<strong>Bonus: What States have the highest estimated percent change between the 2010 Census and the 2017 Population Estimate?</strong><br>This will take several lines of code, as it requires a recalculation of PercentChange.
```
# CODE HERE
new_pop = pop.groupby("State").sum()
new_pop["PercentChange"] = (new_pop["2017PopEstimate"] - new_pop["2010Census"]) / new_pop["2010Census"] * 100
new_pop.sort_values(["PercentChange"], ascending=False)[:5]
# DON'T WRITE HERE
```
# GREAT JOB!
| github_jupyter |
## PLINK GWAS Regression Tutorial
These commands walk through running the GWAS regressions from Marees et al. 2018 using PLINK.
As in all PLINK tutorials, the comments and code from the original tutorial are included with R steps commented out (and replaced by python where necessary) and to disambiguate between comments from the original authors and me, the ```#*#``` character is used.
```
import os.path as osp
import pandas as pd
import numpy as np
import plotnine as pn
from IPython.display import display, Image
%run ../init/benchmark.py
register_timeop_magic(get_ipython(), 'plink')
prev_dir = osp.expanduser('~/data/gwas/tutorial/2_PS_GWAS')
data_dir = osp.expanduser('~/data/gwas/tutorial/3_AA_GWAS')
data_dir
```
Copy necessary data from directory for second step in tutorial (population stratification) to directory for this step:
```
%%timeop -o ps0
%%bash -s "$prev_dir" "$data_dir"
set -e
echo "$1 $2"
cp $1/HapMap_3_r3_13.* $2/
cp $1/covar_mds.txt $2/
```
### Step 1: Association Analysis via Hypothesis Testing
Run chi-square tests for each variant testing whether or not independence between the frequency of cases and controls is unlikely for dominant and recessive alleles (as well as several others, all as separate tests). See [PLINK#assoc](https://www.cog-genomics.org/plink/1.9/assoc) for more details.
```
%%timeop -o aa1
%%bash -s "$data_dir"
set -e; cd $1
# assoc
plink --bfile HapMap_3_r3_13 --assoc --out assoc_results
# Note, the --assoc option does not allow to correct covariates such as principal components (PC's)/ MDS components, which makes it
# less suited for association analyses.
printf '=%.0s' {1..80}; echo # HR
echo "head assoc_results.assoc"
head assoc_results.assoc
```
### Step 2: Association Analysis va Regression
Use the PLINK [--logistic](https://www.cog-genomics.org/plink/1.9/assoc#linear) command to run regressions adjusted for population stratification covariates.
```
%%timeop -o aa2
%%bash -s "$data_dir"
set -e; cd $1
# logistic
# We will be using 10 principal components as covariates in this logistic analysis. We use the MDS components calculated from the previous tutorial: covar_mds.txt.
plink --bfile HapMap_3_r3_13 --covar covar_mds.txt --logistic --hide-covar --out logistic_results
# Note, we use the option -ñhide-covar to only show the additive results of the SNPs in the output file.
# Remove NA values, those might give problems generating plots in later steps.
awk '!/'NA'/' logistic_results.assoc.logistic > logistic_results.assoc_2.logistic
printf '=%.0s' {1..80}; echo # HR
echo "head logistic_results.assoc_2.logistic"
head logistic_results.assoc_2.logistic
```
### Step 3: Visualization
```
def get_data(path):
return (
pd.read_csv(path, sep='\s+')
.sort_values(['CHR', 'BP'])
.reset_index(drop=True)
.rename_axis('POS', axis='index')
.reset_index()
)
get_data(osp.join(data_dir, 'assoc_results.assoc')).head()
get_data(osp.join(data_dir, 'logistic_results.assoc_2.logistic')).head()
def manhattan_plot(df, limit=20000):
return (
pn.ggplot(
df
.sort_values('P')
.reset_index(drop=True)
.head(limit)
.assign(LOGP=lambda df: -np.log10(df['P']))
.assign(CHR=lambda df: df['CHR'].astype(str))
,
pn.aes(x='POS', y='LOGP', fill='CHR', color='CHR')
) +
pn.geom_point() +
pn.geom_hline(yintercept=5) +
pn.theme_bw() +
pn.theme(figure_size=(16, 4))
)
def qq_plot(df, limit=20000):
return (
pn.ggplot(
df
.sort_values('P')
.assign(OBS=lambda df: -np.log10(df['P']))
.assign(EXP=lambda df: -np.log10(np.arange(1, len(df) + 1) / float(len(df))))
.head(limit),
pn.aes(x='EXP', y='OBS')
) +
pn.geom_point() +
pn.geom_abline() +
pn.theme_bw()
)
display(manhattan_plot(get_data(osp.join(data_dir, 'logistic_results.assoc_2.logistic'))) +
pn.ggtitle('Manhattan Plot (Logistic Regression)'))
display(manhattan_plot(get_data(osp.join(data_dir, 'assoc_results.assoc'))) +
pn.ggtitle('Manhattan Plot (Assocation Tests)'))
display(Image('figures/manhattan-logistic.jpeg'))
display(Image('figures/manhattan-assoc.jpeg'))
display(qq_plot(get_data(osp.join(data_dir, 'logistic_results.assoc_2.logistic'))) +
pn.ggtitle('QQ Plot (Logistic Regression)'))
display(qq_plot(get_data(osp.join(data_dir, 'assoc_results.assoc'))) +
pn.ggtitle('QQ Plot (Assocation Tests)'))
display(Image('figures/QQ-Plot_logistic.jpeg'))
display(Image('figures/QQ-Plot_assoc.jpeg'))
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Convert Your Existing Code to TensorFlow 2.0
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/beta/guide/migration_guide">
<img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/guide/migration_guide.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/guide/migration_guide.ipynb">
<img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
View source on GitHub</a>
</td>
</table>
It is still possible to run 1.X code, unmodified (except for contrib), in TensorFlow 2.0:
```
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
```
However, this does not let you take advantage of many of the improvements made in TensorFlow 2.0. This guide will help you upgrade your code, making it simpler, more performant, and easier to maintain.
## Automatic conversion script
The first step is to try running the [upgrade script](./upgrade.md).
This will do an initial pass at upgrading your code to TensorFlow 2.0. But it can't make your code idiomatic to TensorFlowF 2.0. Your code may still make use of `tf.compat.v1` endpoints to access placeholders, sessions, collections, and other 1.x-style functionality.
## Make the code 2.0-native
This guide will walk through several examples of converting TensorFlow 1.x code to TensorFlow 2.0. These changes will let your code take advantage of performance optimizations and simplified API calls.
In each case, the pattern is:
### 1. Replace `tf.Session.run` calls
Every `tf.Session.run` call should be replaced by a Python function.
* The `feed_dict` and `tf.placeholder`s become function arguments.
* The `fetches` become the function's return value.
You can step-through and debug the function using standard Python tools like `pdb`.
When you're satisfied that it works, add a `tf.function` decorator to make it run efficiently in graph mode. See the [Autograph Guide](autograph.ipynb) for more on how this works.
### 2. Use python objects to track variables and losses
Use `tf.Variable` instead of `tf.get_variable`.
Every `variable_scope` can be converted to a Python object. Typically this will be one of:
* `tf.keras.layers.Layer`
* `tf.keras.Model`
* `tf.Module`
If you need to aggregate lists of variables (like `tf.Graph.get_collection(tf.GraphKeys.VARIABLES)`), use the `.variables` and `.trainable_variables` attributes of the `Layer` and `Model` objects.
These `Layer` and `Model` classes implement several other properties that remove the need for global collections. Their `.losses` property can be a replacement for using the `tf.GraphKeys.LOSSES` collection.
See the [keras guides](keras.ipynb) for details.
Warning: Many `tf.compat.v1` symbols use the global collections implicitly.
### 3. Upgrade your training loops
Use the highest level API that works for your use case. Prefer `tf.keras.Model.fit` over building your own training loops.
These high level functions manage a lot of the low-level details that might be easy to miss if you write your own training loop. For example, they automatically collect the regularization losses, and set the `training=True` argument when calling the model.
### 4. Upgrade your data input pipelines
Use `tf.data` datasets for data input. Thse objects are efficient, expressive, and integrate well with tensorflow.
They can be passed directly to the `tf.keras.Model.fit` method.
```
model.fit(dataset, epochs=5)
```
They can be iterated over directly standard Python:
```
for example_batch, label_batch in dataset:
break
```
## Converting models
### Setup
```
from __future__ import absolute_import, division, print_function, unicode_literals
!pip install tensorflow==2.0.0-beta0
import tensorflow as tf
import tensorflow_datasets as tfds
```
### Low-level variables & operator execution
Examples of low-level API use include:
* using variable scopes to control reuse
* creating variables with `tf.get_variable`.
* accessing collections explicitly
* accessing collections implicitly with methods like :
* `tf.global_variables`
* `tf.losses.get_regularization_loss`
* using `tf.placeholder` to set up graph inputs
* executing graphs with `session.run`
* initializing variables manually
#### Before converting
Here is what these patterns may look like in code using TensorFlow 1.x.
```python
in_a = tf.placeholder(dtype=tf.float32, shape=(2))
in_b = tf.placeholder(dtype=tf.float32, shape=(2))
def forward(x):
with tf.variable_scope("matmul", reuse=tf.AUTO_REUSE):
W = tf.get_variable("W", initializer=tf.ones(shape=(2,2)),
regularizer=tf.contrib.layers.l2_regularizer(0.04))
b = tf.get_variable("b", initializer=tf.zeros(shape=(2)))
return W * x + b
out_a = forward(in_a)
out_b = forward(in_b)
reg_loss = tf.losses.get_regularization_loss(scope="matmul")
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
outs = sess.run([out_a, out_b, reg_loss],
feed_dict={in_a: [1, 0], in_b: [0, 1]})
```
#### After converting
In the converted code:
* The variables are local Python objects.
* The `forward` function still defines the calculation.
* The `sess.run` call is replaced with a call to `forward`
* The optional `tf.function` decorator can be added for performance.
* The regularizations are calculated manually, without referring to any global collection.
* **No sessions or placeholders.**
```
W = tf.Variable(tf.ones(shape=(2,2)), name="W")
b = tf.Variable(tf.zeros(shape=(2)), name="b")
@tf.function
def forward(x):
return W * x + b
out_a = forward([1,0])
print(out_a)
out_b = forward([0,1])
regularizer = tf.keras.regularizers.l2(0.04)
reg_loss = regularizer(W)
```
### Models based on `tf.layers`
The `tf.layers` module is used to contain layer-functions that relied on `tf.variable_scope` to define and reuse variables.
#### Before converting
```python
def model(x, training, scope='model'):
with tf.variable_scope(scope, reuse=tf.AUTO_REUSE):
x = tf.layers.conv2d(x, 32, 3, activation=tf.nn.relu,
kernel_regularizer=tf.contrib.layers.l2_regularizer(0.04))
x = tf.layers.max_pooling2d(x, (2, 2), 1)
x = tf.layers.flatten(x)
x = tf.layers.dropout(x, 0.1, training=training)
x = tf.layers.dense(x, 64, activation=tf.nn.relu)
x = tf.layers.batch_normalization(x, training=training)
x = tf.layers.dense(x, 10, activation=tf.nn.softmax)
return x
train_out = model(train_data, training=True)
test_out = model(test_data, training=False)
```
#### After converting
* The simple stack of layers fits neatly into `tf.keras.Sequential`. (For more complex models see [custom layers and models](keras/custom_layers_and_models.ipynb), and [the functional API](keras/functional.ipynb).)
* The model tracks the variables, and regularization losses.
* The conversion was one-to-one because there is a direct mapping from `tf.layers` to `tf.keras.layers`.
Most arguments stayed the same. But notice the differences:
* The `training` argument is passed to each layer by the model when it runs.
* The first argument to the original `model` function (the input `x`) is gone. This is because object layers separate building the model from calling the model.
Also note that:
* If you were using regularizers of initializers from `tf.contrib`, these have more argument changes than others.
* The code no longer writes to collections, so functions like `tf.losses.get_regularization_loss` will no longer return these values, potentially breaking your training loops.
```
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu',
kernel_regularizer=tf.keras.regularizers.l2(0.04),
input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dropout(0.1),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(10, activation='softmax')
])
train_data = tf.ones(shape=(1, 28, 28, 1))
test_data = tf.ones(shape=(1, 28, 28, 1))
train_out = model(train_data, training=True)
print(train_out)
test_out = model(test_data, training=False)
print(test_out)
# Here are all the trainable variables.
len(model.trainable_variables)
# Here is the regularization loss.
model.losses
```
### Mixed variables & tf.layers
Existing code often mixes lower-level TF 1.x variables and operations with higher-level `tf.layers`.
#### Before converting
```python
def model(x, training, scope='model'):
with tf.variable_scope(scope, reuse=tf.AUTO_REUSE):
W = tf.get_variable(
"W", dtype=tf.float32,
initializer=tf.ones(shape=x.shape),
regularizer=tf.contrib.layers.l2_regularizer(0.04),
trainable=True)
if training:
x = x + W
else:
x = x + W * 0.5
x = tf.layers.conv2d(x, 32, 3, activation=tf.nn.relu)
x = tf.layers.max_pooling2d(x, (2, 2), 1)
x = tf.layers.flatten(x)
return x
train_out = model(train_data, training=True)
test_out = model(test_data, training=False)
```
#### After converting
To convert this code, follow the pattern of mapping layers to layers as in the previous example.
The `tf.variable_scope` is effectively a layer of its own. So rewrite it as a `tf.keras.layers.Layer`. See [the guide](keras/custom_layers_and_models.ipynb) for details.
The general pattern is:
* Collect layer parameters in `__init__`.
* Build the variables in `build`.
* Execute the calculations in `call`, and return the result.
The `tf.variable_scope` is essentially a layer of its own. So rewrite it as a `tf.keras.layers.Layer`. See [the guide](keras/custom_layers_and_models.ipynb) for details.
```
# Create a custom layer for part of the model
class CustomLayer(tf.keras.layers.Layer):
def __init__(self, *args, **kwargs):
super(CustomLayer, self).__init__(*args, **kwargs)
def build(self, input_shape):
self.w = self.add_weight(
shape=input_shape[1:],
dtype=tf.float32,
initializer=tf.keras.initializers.ones(),
regularizer=tf.keras.regularizers.l2(0.02),
trainable=True)
# Call method will sometimes get used in graph mode,
# training will get turned into a tensor
@tf.function
def call(self, inputs, training=None):
if training:
return inputs + self.w
else:
return inputs + self.w * 0.5
custom_layer = CustomLayer()
print(custom_layer([1]).numpy())
print(custom_layer([1], training=True).numpy())
train_data = tf.ones(shape=(1, 28, 28, 1))
test_data = tf.ones(shape=(1, 28, 28, 1))
# Build the model including the custom layer
model = tf.keras.Sequential([
CustomLayer(input_shape=(28, 28, 1)),
tf.keras.layers.Conv2D(32, 3, activation='relu'),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
])
train_out = model(train_data, training=True)
test_out = model(test_data, training=False)
```
Some things to note:
* Subclassed Keras models & layers need to run in both v1 graphs (no automatic control dependencies) and in eager mode
* Wrap the `call()` in a `tf.function()` to get autograph and automatic control dependencies
* Don't forget to accept a `training` argument to `call`.
* Sometimes it is a `tf.Tensor`
* Sometimes it is a Python boolean.
* Create model variables in constructor or `def build()` using `self.add_weight()`.
* In `build` you have access to the input shape, so can create weights with matching shape.
* Using `tf.keras.layers.Layer.add_weight` allows Keras to track variables and regularization losses.
* Don't keep `tf.Tensors` in your objects.
* They might get created either in a `tf.function` or in the eager context, and these tensors behave differently.
* Use `tf.Variable`s for state, they are always usable from both contexts
* `tf.Tensors` are only for intermediate values.
### A note on Slim & contrib.layers
A large amount of older TensorFlow 1.x code uses the [Slim](https://ai.googleblog.com/2016/08/tf-slim-high-level-library-to-define.html) library, which was packaged with TensorFlow 1.x as `tf.contrib.layers`. As a `contrib` module, this is no longer available in TensorFlow 2.0, even in `tf.compat.v1`. Converting code using Slim to TF 2.0 is more involved than converting repositories that use `tf.layers`. In fact, it may make sense to convert your Slim code to `tf.layers` first, then convert to Keras.
* Remove `arg_scopes`, all args need to be explicit
* If you use them, split `normalizer_fn` and `activation_fn` into their own layers
* Separable conv layers map to one or more different Keras layers (depthwise, pointwise, and separable Keras layers)
* Slim and `tf.layers` have different arg names & default values
* Some args have different scales
* If you use Slim pre-trained models, try out `tf.keras.applications` or [TFHub](https://tensorflow.orb/hub)
Some `tf.contrib` layers might not have been moved to core TensorFlow but have instead been moved to the [TF add-ons package](https://github.com/tensorflow/addons).
## Training
There are many ways to feed data to a `tf.keras` model. They will accept Python generators and Numpy arrays as input.
The recomended way to feed data to a model is to use the `tf.data` package, which contains a collection of high performance classes for manipulating data.
If you are still using `tf.queue`, these are only supported as data-structures, not as input pipelines.
### Using Datasets
The [TensorFlow Datasets](https://tensorflow.org/datasets) package (`tfds`) contains utilities for loading predefined datasets as `tf.data.Dataset` objects.
For this example, load the MNISTdataset, using `tfds`:
```
datasets, info = tfds.load(name='mnist', with_info=True, as_supervised=True)
mnist_train, mnist_test = datasets['train'], datasets['test']
```
Then prepare the data for training:
* Re-scale each image.
* Shuffle the order of the examples.
* Collect batches of images and labels.
```
BUFFER_SIZE = 10 # Use a much larger value for real code.
BATCH_SIZE = 64
NUM_EPOCHS = 5
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255
return image, label
```
To keep the example short, trim the dataset to only return 5 batches:
```
train_data = mnist_train.map(scale).shuffle(BUFFER_SIZE).batch(BATCH_SIZE).take(5)
test_data = mnist_test.map(scale).batch(BATCH_SIZE).take(5)
STEPS_PER_EPOCH = 5
train_data = train_data.take(STEPS_PER_EPOCH)
test_data = test_data.take(STEPS_PER_EPOCH)
image_batch, label_batch = next(iter(train_data))
```
### Use Keras training loops
If you don't need low level control of your training process, using Keras's built-in `fit`, `evaluate`, and `predict` methods is recomended. These methods provide a uniform interface to train the model regardless of the implementation (sequential, functional, or sub-classed).
The advantages of these methods include:
* They accept Numpy arrays, Python generators and, `tf.data.Datasets`
* They apply regularization, and activation losses automatically.
* They support `tf.distribute` [for multi-device training](distribute_strategy.ipynb).
* They support arbitrary callables as losses and metrics.
* They support callbacks like `tf.keras.callbacks.TensorBoard`, and custom callbacks.
* They are performant, automatically using TensorFlow graphs.
Here is an example of training a model using a `Dataset`. (For details on how this works see [tutorials](../tutorials).)
```
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu',
kernel_regularizer=tf.keras.regularizers.l2(0.02),
input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dropout(0.1),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(10, activation='softmax')
])
# Model is the full model w/o custom layers
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(train_data, epochs=NUM_EPOCHS)
loss, acc = model.evaluate(test_data)
print("Loss {}, Accuracy {}".format(loss, acc))
```
### Write your own loop
If the Keras model's training step works for you, but you need more control outside that step, consider using the `tf.keras.model.train_on_batch` method, in your own data-iteration loop.
Remember: Many things can be implemented as a `tf.keras.Callback`.
This method has many of the advantages of the methods mentioned in the previous section, but gives the user control of the outer loop.
You can also use `tf.keras.model.test_on_batch` or `tf.keras.Model.evaluate` to check performance during training.
Note: `train_on_batch` and `test_on_batch`, by default return the loss and metrics for the single batch. If you pass `reset_metrics=False` they return accumulated metrics and you must remember to appropriately reset the metric accumulators. Also remember that some metrics like `AUC` require `reset_metrics=False` to be calculated correctly.
To continue training the above model:
```
# Model is the full model w/o custom layers
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
metrics_names = model.metrics_names
for epoch in range(NUM_EPOCHS):
#Reset the metric accumulators
model.reset_metrics()
for image_batch, label_batch in train_data:
result = model.train_on_batch(image_batch, label_batch)
print("train: ",
"{}: {:.3f}".format(metrics_names[0], result[0]),
"{}: {:.3f}".format(metrics_names[1], result[1]))
for image_batch, label_batch in test_data:
result = model.test_on_batch(image_batch, label_batch,
# return accumulated metrics
reset_metrics=False)
print("\neval: ",
"{}: {:.3f}".format(metrics_names[0], result[0]),
"{}: {:.3f}".format(metrics_names[1], result[1]))
```
<p id="custom_loops"/>
### Customize the training step
If you need more flexibility and control, you can have it by implementing your own training loop. There are three steps:
1. Iterate over a Python generator or `tf.data.Dataset` to get batches of examples.
2. Use `tf.GradientTape` to collect gradients.
3. Use a `tf.keras.optimizer` to apply weight updates to the model's variables.
Remember:
* Always include a `training` argument on the `call` method of subclassed layers and models.
* Make sure to call the model with the `training` argument set correctly.
* Depending on usage, model variables may not exist until the model is run on a batch of data.
* You need to manually handle things like regularization losses for the model.
Note the simplifications relative to v1:
* There is no need to run variable initializers. Variables are initialized on creation.
* There is no need to add manual control dependencies. Even in `tf.function` operations act as in eager mode.
```
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu',
kernel_regularizer=tf.keras.regularizers.l2(0.02),
input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dropout(0.1),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(10, activation='softmax')
])
optimizer = tf.keras.optimizers.Adam(0.001)
loss_fn = tf.keras.losses.SparseCategoricalCrossentropy()
@tf.function
def train_step(inputs, labels):
with tf.GradientTape() as tape:
predictions = model(inputs, training=True)
regularization_loss = tf.math.add_n(model.losses)
pred_loss = loss_fn(labels, predictions)
total_loss = pred_loss + regularization_loss
gradients = tape.gradient(total_loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
for epoch in range(NUM_EPOCHS):
for inputs, labels in train_data:
train_step(inputs, labels)
print("Finished epoch", epoch)
```
### New-style metrics
In TensorFlow 2.0, metrics are objects. Metric objects work both eagerly and in `tf.function`s. A metric object has the following methods:
* `update_state()` — add new observations
* `result()` —get the current result of the metric, given the observed values
* `reset_states()` — clear all observations.
The object itself is callable. Calling updates the state with new observations, as with `update_state`, and returns the new result of the metric.
You don't have to manually initialize a metric's variables, and because TensorFlow 2.0 has automatic control dependencies, you don't need to worry about those either.
The code below uses a metric to keep track of the mean loss observed within a custom training loop.
```
# Create the metrics
loss_metric = tf.keras.metrics.Mean(name='train_loss')
accuracy_metric = tf.keras.metrics.SparseCategoricalAccuracy(name='train_accuracy')
@tf.function
def train_step(inputs, labels):
with tf.GradientTape() as tape:
predictions = model(inputs, training=True)
regularization_loss = tf.math.add_n(model.losses)
pred_loss = loss_fn(labels, predictions)
total_loss = pred_loss + regularization_loss
gradients = tape.gradient(total_loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
# Update the metrics
loss_metric.update_state(total_loss)
accuracy_metric.update_state(labels, predictions)
for epoch in range(NUM_EPOCHS):
# Reset the metrics
loss_metric.reset_states()
accuracy_metric.reset_states()
for inputs, labels in train_data:
train_step(inputs, labels)
# Get the metric results
mean_loss = loss_metric.result()
mean_accuracy = accuracy_metric.result()
print('Epoch: ', epoch)
print(' loss: {:.3f}'.format(mean_loss))
print(' accuracy: {:.3f}'.format(mean_accuracy))
```
## Saving & Loading
### Checkpoint compatibility
TensorFlow 2.0 uses [object-based checkpoints](checkpoints.ipynb).
Old-style name-based checkpoints can still be loaded, if you're careful.
The code conversion process may result in variable name changes, but there are workarounds.
The simplest approach it to line up the names of the new model with the names in the checkpoint:
* Variables still all have a `name` argument you can set.
* Keras models also take a `name` argument as which they set as the prefix for their variables.
* The `tf.name_scope` function can be used to set variable name prefixes. This is very different from `tf.variable_scope`. It only affects names, and doesn't track variables & reuse.
If that does not work for your use-case, try the `tf.compat.v1.train.init_from_checkpoint` function. It takes an `assignment_map` argument, which specifies the mapping from old names to new names.
Note: Unlike object based checkpoints, which can [defer loading](checkpoints.ipynb#loading_mechanics), name-based checkpoints require that all variables be built when the function is called. Some models defer building variables until you call `build` or run the model on a batch of data.
### Saved models compatibility
There are no significant compatibility concerns for saved models.
* TensorFlow 1.x saved_models work in TensorFlow 2.0.
* TensorFlow 2.0 saved_models even load work in TensorFlow 1.x if all the ops are supported.
## Estimators
### Training with Estimators
Estimators are supported in TensorFlow 2.0.
When you use estimators, you can use `input_fn()`, `tf.estimator.TrainSpec`, and `tf.estimator.EvalSpec` from TensorFlow 1.x.
Here is an example using `input_fn` with train and evaluate specs.
#### Creating the input_fn and train/eval specs
```
# Define the estimator's input_fn
def input_fn():
datasets, info = tfds.load(name='mnist', with_info=True, as_supervised=True)
mnist_train, mnist_test = datasets['train'], datasets['test']
BUFFER_SIZE = 10000
BATCH_SIZE = 64
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255
return image, label[..., tf.newaxis]
train_data = mnist_train.map(scale).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
return train_data.repeat()
# Define train & eval specs
train_spec = tf.estimator.TrainSpec(input_fn=input_fn,
max_steps=STEPS_PER_EPOCH * NUM_EPOCHS)
eval_spec = tf.estimator.EvalSpec(input_fn=input_fn,
steps=STEPS_PER_EPOCH)
```
### Using a Keras model definition
There are some differences in how to construct your estimators in TensorFlow 2.0.
We recommend that you define your model using Keras, then use the `tf.keras.model_to_estimator` utility to turn your model into an estimator. The code below shows how to use this utility when creating and training an estimator.
```
def make_model():
return tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu',
kernel_regularizer=tf.keras.regularizers.l2(0.02),
input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dropout(0.1),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(10, activation='softmax')
])
model = make_model()
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
estimator = tf.keras.estimator.model_to_estimator(
keras_model = model
)
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
```
### Using a custom `model_fn`
If you have an existing custom estimator `model_fn` that you need to maintain, you can convert your `model_fn` to use a Keras model.
However, for compatibility reasons, a custom `model_fn` will still run in 1.x-style graph mode. This means there is no eager execution and no automatic control dependencies.
Using a Keras models in a custom `model_fn` is similar to using it in a custom training loop:
* Set the `training` phase appropriately, based on the `mode` argument.
* Explicitly pass the model's `trainable_variables` to the optimizer.
But there are important differences, relative to a [custom loop](#custom_loop):
* Instead of using `model.losses`, extract the losses using `tf.keras.Model.get_losses_for`.
* Extract the model's updates using `tf.keras.Model.get_updates_for`
Note: "Updates" are changes that need to be applied to a model after each batch. For example, the moving averages of the mean and variance in a `tf.keras.layers.BatchNormalization` layer.
The following code creates an estimator from a custom `model_fn`, illustrating all of these concerns.
```
def my_model_fn(features, labels, mode):
model = make_model()
optimizer = tf.compat.v1.train.AdamOptimizer()
loss_fn = tf.keras.losses.SparseCategoricalCrossentropy()
training = (mode == tf.estimator.ModeKeys.TRAIN)
predictions = model(features, training=training)
reg_losses = model.get_losses_for(None) + model.get_losses_for(features)
total_loss = loss_fn(labels, predictions) + tf.math.add_n(reg_losses)
accuracy = tf.compat.v1.metrics.accuracy(labels=labels,
predictions=tf.math.argmax(predictions, axis=1),
name='acc_op')
update_ops = model.get_updates_for(None) + model.get_updates_for(features)
minimize_op = optimizer.minimize(
total_loss,
var_list=model.trainable_variables,
global_step=tf.compat.v1.train.get_or_create_global_step())
train_op = tf.group(minimize_op, update_ops)
return tf.estimator.EstimatorSpec(
mode=mode,
predictions=predictions,
loss=total_loss,
train_op=train_op, eval_metric_ops={'accuracy': accuracy})
# Create the Estimator & Train
estimator = tf.estimator.Estimator(model_fn=my_model_fn)
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
```
### Premade Estimators
[Premade Estimators](https://www.tensorflow.org/guide/premade_estimators) in the family of `tf.estimator.DNN*`, `tf.estimator.Linear*` and `tf.estimator.DNNLinearCombined*` are still supported in in the TensorFlow 2.0 API, however, some arguments have changed:
1. `input_layer_partitioner`: Removed in 2.0.
2. `loss_reduction`: Updated to `tf.keras.losses.Reduction` instead of `tf.compat.v1.losses.Reduction`. Its default value is also changed to `tf.keras.losses.Reduction.SUM_OVER_BATCH_SIZE` from `tf.compat.v1.losses.Reduction.SUM`.
3. `optimizer`, `dnn_optimizer` and `linear_optimizer`: this arg has been updated to `tf.keras.optimizers` instead of the `tf.compat.v1.train.Optimizer`.
To migrate the above changes:
1. No migration is needed for `input_layer_partitioner` since [`Distribution Strategy`](https://www.tensorflow.org/guide/distribute_strategy) will handle it automatically in TF 2.0.
2. For `loss_reduction`, check [`tf.keras.losses.Reduction`](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/losses/Reduction) for the supported options.
3. For `optimizer` args, if you do not pass in an `optimizer`, `dnn_optimizer` or `linear_optimizer` arg, or if you specify the `optimizer` arg as a `string` in your code, you don't need to change anything. `tf.keras.optimizers` is used by default. Otherwise, you need to update it from `tf.compat.v1.train.Optimizer` to its corresponding [`tf.keras.optimizers`](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/optimizers)
#### Checkpoint Converter
The migration of `optimizer` will break the checkpoints in TF 1.x, as `tf.keras.optimizers` generates a different set of variables to be saved in checkpoints. To make the checkpoint reusable after your migration to TF 2.0, please check the checkpoint converter tool for optimizers to convert the checkpoints from TF 1.x to TF 2.0. The converted checkpoints can be used to restore the pre-trained models in TF 2.0.
## TensorShape
This class was simplified to hold `int`s, instead of `tf.compat.v1.Dimension` objects. So there is no need to call `.value()` to get an `int`.
Individual `tf.compat.v1.Dimension` objects are still accessible from `tf.TensorShape.dims`.
The following demonstrate the differences between TensorFlow 1.x and TensorFlow 2.0.
```
# Create a shape and choose an index
i = 0
shape = tf.TensorShape([16, None, 256])
shape
```
If you had this in TF 1.x:
```python
value = shape[i].value
```
Then do this in TF 2.0:
```
value = shape[i]
value
```
If you had this in TF 1.x:
```python
for dim in shape:
value = dim.value
print(value)
```
Then do this in TF 2.0:
```
for value in shape:
print(value)
```
If you had this in TF 1.x (Or used any other dimension method):
```python
dim = shape[i]
dim.assert_is_compatible_with(other_dim)
```
Then do this in TF 2.0:
```
other_dim = 16
Dimension = tf.compat.v1.Dimension
if shape.rank is None:
dim = Dimension(None)
else:
dim = shape.dims[i]
dim.is_compatible_with(other_dim) # or any other dimension method
shape = tf.TensorShape(None)
if shape:
dim = shape.dims[i]
dim.is_compatible_with(other_dim) # or any other dimension method
```
The boolean value of a `tf.TensorShape` is `True` if the rank is known, `False` otherwise.
```
print(bool(tf.TensorShape([]))) # Scalar
print(bool(tf.TensorShape([0]))) # 0-length vector
print(bool(tf.TensorShape([1]))) # 1-length vector
print(bool(tf.TensorShape([None]))) # Unknown-length vector
print(bool(tf.TensorShape([1, 10, 100]))) # 3D tensor
print(bool(tf.TensorShape([None, None, None]))) # 3D tensor with no known dimensions
print()
print(bool(tf.TensorShape(None))) # A tensor with unknown rank.
```
## Other behavioral changes
There are a few other behavioral changes in TensorFlow 2.0 that you may run into.
### ResourceVariables
TensorFlow 2.0 creates `ResourceVariables` by default, not `RefVariables`.
`ResourceVariables` are locked for writing, and so provide more intuitive consistency guarantees.
* This may change behavior in edge cases.
* This may occasionally create extra copies, can have higher memory usage
* This can be disabled by passing `use_resource=False` to the `tf.Variable` constructor.
### Control Flow
The control flow op implementation has been simplified, and so produces different graphs in TensorFlow 2.0
## Conclusions
The overall process is:
1. Run the upgrade script.
2. Remove contrib symbols.
3. Switch your models to an object oriented style (Keras).
4. Use `tf.keras` or `tf.estimator` training and evaluation loops where you can.
5. Otherwise, use custom loops, but be sure to avoid sessions & collections.
It takes a little work to convert code to idiomatic TensorFlow 2.0, but every change results in:
* Fewer lines of code.
* Increased clarity and simplicity.
* Easier debugging.
| github_jupyter |
```
# Example-2-GP-BS-Derivatives
# Author: Matthew Dixon
# Version: 1.0 (28.4.2020)
# License: MIT
# Email: matthew.dixon@iit.edu
# Notes: tested on Mac OS X running Python 3.6.9 with the following packages:
# scikit-learn=0.22.1, numpy=1.18.1, matplotlib=3.1.3
# Citation: Please cite the following reference if this notebook is used for research purposes:
# Dixon M.F. Halperin I. and P. Bilokon, Machine Learning in Finance: From Theory to Practice, Springer Graduate textbook Series, 2020.
```
# Calculating the Greeks
# Overview
The purpose of this notebook is to demonstrate the derivation of the greeks in a Gaussian Process Regression model (GP), fitted to option price data.
In this notebook, European option prices are generated from the Black-Scholes model. The notebook begins by building a GP call model, where the input is the underlying price. The delta is then derived and compared with the Black-Scholes (BS)
delta. The exercise is repeated, but using the volatility as the input instead of the underlying price. The vega of the GP is then derived and compared with the BS vega.
```
from BlackScholes import bsformula
import numpy as np
import scipy as sp
from sklearn import gaussian_process
from sklearn.gaussian_process.kernels import RBF
import matplotlib.pyplot as plt
%matplotlib inline
```
## Black-Scholes Model
First, set the model parameters
```
KC = 130 # Call strike
KP = 70 # Put strike
r = 0.002 # risk-free rate
sigma = 0.4 # implied volatility
T = 2.0 # Time to maturity
S0 = 100 # Underlying spot
lb = 0 # lower bound on domain
ub = 300 # upper bound on domain
training_number = 100 # Number of training samples
testing_number = 50 # Number of testing samples
sigma_n = 1e-8 # additive noise in GP
```
Define the call and put prices using the BS model
```
call = lambda x, y: bsformula(1, lb+(ub-lb)*x, KC, r, T, y, 0)[0]
put = lambda x, y: bsformula(-1, lb+(ub-lb)*x, KP, r, T, y, 0)[0]
```
# Delta
Generate the training and testing data, where the input is the gridded underlying and the output are the option prices.
```
x_train = np.array(np.linspace(0.01, 1.2, training_number), dtype='float32').reshape(training_number, 1)
x_test = np.array(np.linspace(0.01, 1.0, testing_number), dtype='float32').reshape(testing_number, 1)
y_train = []
for idx in range(len(x_train)):
y_train.append(call(x_train[idx], sigma))
y_train = np.array(y_train)
```
Fit the GP model to the generated data
```
sk_kernel = RBF(length_scale=1.0, length_scale_bounds=(0.01, 10000.0))
gp = gaussian_process.GaussianProcessRegressor(kernel=sk_kernel, n_restarts_optimizer=20)
gp.fit(x_train, y_train)
```
Get the model's predicted outputs for each of the test inputs
```
y_pred, sigma_hat = gp.predict(x_test, return_std=True)
```
Derive the GP delta
```
l = gp.kernel_.length_scale
rbf = gaussian_process.kernels.RBF(length_scale=l)
Kernel = rbf(x_train, x_train)
K_y = Kernel + np.eye(training_number) * sigma_n
L = sp.linalg.cho_factor(K_y)
alpha_p = sp.linalg.cho_solve(np.transpose(L), y_train)
k_s = rbf(x_test, x_train)
k_s_prime = (x_train.T - x_test) * k_s / l**2
f_prime = np.dot(k_s_prime, alpha_p) / (ub - lb)
```
Calculate the BS delta
```
delta = lambda x, y: bsformula(1, lb+(ub-lb)*x, KC, r, T, y, 0)[1]
plt.plot(delta(x_test, sigma) - f_prime)
```
Compare the GP delta with the BS delta
```
plt.figure(figsize = (10,6),facecolor='white', edgecolor='black')
plt.plot(lb+(ub-lb)*x_test, delta(x_test,sigma), color = 'black', label = 'Exact')
plt.plot(lb+(ub-lb)*x_test, f_prime, color = 'red', label = 'GP')
plt.grid(True)
plt.xlabel('S')
plt.ylabel('$\Delta$')
plt.legend(loc = 'best', prop={'size':10});
```
Show the error between the GP delta and the BS delta
```
plt.figure(figsize = (10,6),facecolor='white', edgecolor='black')
plt.plot(lb+(ub-lb)*x_test, delta(x_test,sigma) - f_prime, color = 'black', label = 'GP Error')
plt.grid(True)
plt.xlabel('S')
plt.ylabel('Error in $\Delta$')
plt.legend(loc = 'best', prop={'size':10});
```
## Vega
Generate the training and testing data, where the input is the gridded underlying and the output are the option prices. The inputs are again scaled to the unit domain.
```
x_train = np.array(np.linspace(0.01, 1.2, training_number), dtype='float32').reshape(training_number, 1)
x_test = np.array(np.linspace(0.01, 1.0, testing_number), dtype='float32').reshape(testing_number, 1)
y_train = []
for idx in range(len(x_train)):
y_train.append(call((S0-lb)/(ub-lb), x_train[idx]))
y_train = np.array(y_train)
```
Fit the GP model to the generated data
```
sk_kernel = RBF(length_scale=1.0, length_scale_bounds=(0.01, 10000.0))
gp = gaussian_process.GaussianProcessRegressor(kernel=sk_kernel, n_restarts_optimizer=20)
gp.fit(x_train, y_train)
```
Get the model's predicted outputs for each of the test inputs
```
y_pred, sigma_hat = gp.predict(x_test, return_std=True)
```
Derive the GP delta
```
l = gp.kernel_.length_scale
rbf = gaussian_process.kernels.RBF(length_scale=l)
Kernel= rbf(x_train, x_train)
K_y = Kernel + np.eye(training_number) * sigma_n
L = sp.linalg.cho_factor(K_y)
alpha_p = sp.linalg.cho_solve(np.transpose(L), y_train)
k_s = rbf(x_test, x_train)
k_s_prime = np.zeros([len(x_test), len(x_train)])
for i in range(len(x_test)):
for j in range(len(x_train)):
k_s_prime[i, j] = (1.0/l**2) * (x_train[j] - x_test[i]) * k_s[i, j]
f_prime = np.dot(k_s_prime, alpha_p)
```
Calculate the BS delta
```
vega = lambda x, y: bsformula(1, lb + (ub-lb) * x, KC, r, T, y, 0)[2]
vega((S0-lb)/(ub-lb), x_test) - f_prime
```
Compare the GP vega with the BS vega
```
plt.figure(figsize = (10,6), facecolor='white', edgecolor='black')
plt.plot(x_test, vega((S0-lb)/(ub-lb), x_test), color = 'black', label = 'Exact')
plt.plot(x_test, f_prime, color = 'red', label = 'GP')
plt.grid(True)
plt.xlabel('$\\sigma$')
plt.ylabel('$\\nu$')
plt.legend(loc = 'best', prop={'size':10});
```
Plot the error between the GP vega and the BS vega
```
plt.figure(figsize = (10,6), facecolor='white', edgecolor='black')
plt.plot(x_test, vega((S0-lb)/(ub-lb), x_test)-f_prime, color = 'black', label = 'GP Error')
plt.grid(True)
plt.xlabel('$\\sigma$')
plt.ylabel('Error in $\\nu$')
plt.legend(loc = 'best', prop={'size':10});
```
# Idea: Calculate Gamma and then explain what a Gamma Squeeze is
https://www.fool.com/investing/2021/01/28/what-is-a-gamma-squeeze/
| github_jupyter |
### quero usar matplotlib para ilustrar permutações
A primeira coisa é fazer circulos numerados
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
circle1=plt.Circle((0,0),.1,color='r', alpha=0.2, clip_on=False)
plt.axes(aspect="equal")
fig = plt.gcf()
fig.gca().add_artist(circle1)
plt.axis("off")
circle1=plt.Circle((0,0),.1,color='r', alpha=0.2, clip_on=False)
circle2=plt.Circle((0,0.2),.1,color='y', alpha=0.2, clip_on=False)
circle3=plt.Circle((0,0.4),.1,color='b', alpha=0.2, clip_on=False)
circle4=plt.Circle((0,0.6),.1,color='g', alpha=0.2, clip_on=False)
circle5=plt.Circle((0,0.8),.1,color=(0.2,0.6,0.7), alpha=0.2, clip_on=False)
plt.axes(aspect="equal")
fig1 = plt.gcf()
plt.axis("off")
fig1.gca().add_artist(circle1)
fig1.gca().add_artist(circle2)
fig1.gca().add_artist(circle3)
fig1.gca().add_artist(circle4)
fig1.gca().add_artist(circle5)
circle1=plt.Circle((0,0),.1,color='r', alpha=0.2, clip_on=False)
circle2=plt.Circle((0,0.2),.1,color='y', alpha=0.2, clip_on=False)
circle3=plt.Circle((0,0.4),.1,color='b', alpha=0.2, clip_on=False)
circle4=plt.Circle((0,0.6),.1,color='g', alpha=0.2, clip_on=False)
circle5=plt.Circle((0,0.8),.1,color=(0.2,0.6,0.7), alpha=0.2, clip_on=False)
circled1=plt.Circle((1,0),.1,color='r', alpha=0.2, clip_on=False)
circled2=plt.Circle((1,0.2),.1,color='y', alpha=0.2, clip_on=False)
circled3=plt.Circle((1,0.4),.1,color='b', alpha=0.2, clip_on=False)
circled4=plt.Circle((1,0.6),.1,color='g', alpha=0.2, clip_on=False)
circled5=plt.Circle((1,0.8),.1,color=(0.2,0.6,0.7), alpha=0.2, clip_on=False)
plt.axes(aspect="equal")
fig1 = plt.gcf()
plt.axis("off")
fig1.gca().add_artist(circle1)
fig1.gca().add_artist(circle2)
fig1.gca().add_artist(circle3)
fig1.gca().add_artist(circle4)
fig1.gca().add_artist(circle5)
fig1.gca().add_artist(circled1)
fig1.gca().add_artist(circled2)
fig1.gca().add_artist(circled3)
fig1.gca().add_artist(circled4)
fig1.gca().add_artist(circled5)
circle1=plt.Circle((0,0),.1,color='r', alpha=0.2, clip_on=False)
circle2=plt.Circle((0,0.2),.1,color='y', alpha=0.2, clip_on=False)
circle3=plt.Circle((0,0.4),.1,color='b', alpha=0.2, clip_on=False)
circle4=plt.Circle((0,0.6),.1,color='g', alpha=0.2, clip_on=False)
circle5=plt.Circle((0,0.8),.1,color=(0.2,0.6,0.7), alpha=0.2, clip_on=False)
circled1=plt.Circle((1,0),.1,color='r', alpha=0.2, clip_on=False)
circled2=plt.Circle((1,0.2),.1,color='y', alpha=0.2, clip_on=False)
circled3=plt.Circle((1,0.4),.1,color='b', alpha=0.2, clip_on=False)
circled4=plt.Circle((1,0.6),.1,color='g', alpha=0.2, clip_on=False)
circled5=plt.Circle((1,0.8),.1,color=(0.2,0.6,0.7), alpha=0.2, clip_on=False)
plt.axes(aspect="equal")
fig1 = plt.gcf()
plt.axis("off")
fig1.gca().add_artist(circle1)
fig1.gca().add_artist(circle2)
fig1.gca().add_artist(circle3)
fig1.gca().add_artist(circle4)
fig1.gca().add_artist(circle5)
fig1.gca().add_artist(circled1)
fig1.gca().add_artist(circled2)
fig1.gca().add_artist(circled3)
fig1.gca().add_artist(circled4)
fig1.gca().add_artist(circled5)
# as arestas
fig1.gca().plot([0.15,0.85],[0,0.8], color="red", alpha=0.6 )
fig1.gca().text(0.,0.,r'$5$', fontsize=20,verticalalignment='center', horizontalalignment='center')
fig1.gca().text(1,0,r'$5$', fontsize=20, verticalalignment='center', horizontalalignment='center')
fig1.gca().text(1,0.8,r'$1$', fontsize=20, verticalalignment='center', horizontalalignment='center')
fig1.gca().text(0,0.8,r'$1$', fontsize=20, verticalalignment='center', horizontalalignment='center')
fig1.gca().plot([0.15,0.85],[0.8,0.4], color=(.2,.6,.7), alpha=0.6 )
# agora faremos as funções. primeiro a cor de um inteiro
def cor(n):
''' Dado um inteiro n designa uma cor'''
return (n/(n+1), 1- n/(n+1), 1-(n+2)/(n+5))
#teste
circle1=plt.Circle((0,0),.1,color=cor(1), alpha=0.2, clip_on=False)
circle2=plt.Circle((0,0.2),.1,color=cor(3), alpha=0.2, clip_on=False)
plt.axes(aspect="equal")
fig1 = plt.gcf()
plt.axis("off")
fig1.gca().add_artist(circle1)
fig1.gca().add_artist(circle2)
def circulo(x,n):
'''Define um circulo de centro (x,0.2*n) de raio 0.1 e cor n'''
return plt.Circle((x,0.2*n), .1, color=cor(n), alpha=0.3, clip_on=False )
#teste
plt.axes(aspect="equal")
fig1 = plt.gcf()
plt.axis("off")
fig1.gca().add_artist(circulo(0,3))
fig1.gca().add_artist(circulo(0,4))
# função pilha de circulos
def pilha_de_circulos(x,n):
'''Faz uma pilha de n circulos sobre a abcissa x'''
for k in range(n):
fig1.gca().add_artist(circulo(x,k))
fig1.gca().text(x,0.2*k,r'$'+str(k+1)+'$', fontsize=20,verticalalignment='center', horizontalalignment='center')
return
# teste desta função:
plt.axes(aspect="equal")
fig1 = plt.gcf()
plt.axis("off")
pilha_de_circulos(0,3)
pilha_de_circulos(1,3)
pilha_de_circulos(2,3)
# agora a função mapa_permu
def mapa_permu(x,p):
''' desenha a permutação p (uma lista) na posição x'''
l=len(p)
x1= x+.15
x2= x+.85
for y in range(l):
fig1.gca().plot([x1,x2],[0.2*y,0.2*(p[y]-1)], color=cor(y), alpha=0.6 )
return
# teste
plt.axes(aspect="equal")
fig1 = plt.gcf()
plt.axis("off")
pilha_de_circulos(0,3)
pilha_de_circulos(1,3)
pilha_de_circulos(2,3)
mapa_permu(0,[2,1,3])
mapa_permu(1.0, [3,1,2])
plt.axes(aspect="equal")
fig1 = plt.gcf()
plt.axis("off")
pilha_de_circulos(0,5)
pilha_de_circulos(1,5)
mapa_permu(0,[3,2,1,5,4])
def pgrafico(x,p):
'''Faz o grafico da permutação p começando em x'''
n=len(p)
fig1= plt.gcf()
plt.axis("off")
pilha_de_circulos(x,n)
pilha_de_circulos(x+1,n)
return mapa_permu(x,p)
#teste
plt.axes(aspect="equal")
fig1= plt.gcf()
plt.axis("off")
pgrafico(0,[3,1,2])
```
| github_jupyter |
# Fitting to existing data
```
# Base Data Science snippet
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os
import time
from tqdm import tqdm_notebook
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
Inspiration - https://www.lewuathe.com/covid-19-dynamics-with-sir-model.html
```
import sys
sys.path.append("../")
from covid.dataset import fetch_daily_case
from covid.models import SIR
from covid.models.states import CompartmentStates
```
# Fitting French data to SIR model
```
cases = fetch_daily_case(return_data=True)
cases.head()
```
## Getting cases for all France
### Fetching French data and prepare it
```
cases_fr = (
cases.query("granularite =='pays'")
.query("source_nom=='Ministère des Solidarités et de la Santé'")
[["date","cas_confirmes","deces","gueris"]]
.drop_duplicates(subset = ["date"])
.fillna(0.0)
.assign(date = lambda x : pd.to_datetime(x["date"]))
.set_index("date")
)
start,end = cases_fr.index[0],cases_fr.index[-1]
date_range = pd.date_range(start,end,freq="D")
cases_fr = cases_fr.reindex(date_range).fillna(method="ffill")
cases_fr.plot(figsize = (15,4))
plt.show()
```
### Recomputing compartments
```
cases_fr["I"] = cases_fr["cas_confirmes"] - (cases_fr["deces"] + cases_fr["gueris"])
cases_fr["R"] = (cases_fr["deces"] + cases_fr["gueris"])
pop_fr = 66.99*1e6
cases_fr["S"] = pop_fr - cases_fr["I"] - cases_fr["R"]
cases_fr[["S","I","R"]].plot(figsize = (15,4));
cases_fr[["I","R"]].plot(figsize = (15,4));
```
### Smoothing curves
```
from scipy.signal import savgol_filter
import statsmodels.api as sm
def smooth(y,p = 1600):
cycle, trend = sm.tsa.filters.hpfilter(y, p)
return trend
pd.Series(savgol_filter(cases_fr["I"], 51, 2)).plot(figsize = (15,4))
pd.Series(savgol_filter(cases_fr["R"], 51, 2)).plot()
plt.show()
pd.Series(smooth(cases_fr["I"],6.25)).plot(figsize = (15,4),label = "Is")
pd.Series(cases_fr["I"]).plot(label = "I")
pd.Series(smooth(cases_fr["R"],6.25)).plot(label = "Rs")
pd.Series(cases_fr["R"]).plot(label = "R")
plt.legend()
plt.show()
pd.Series(smooth(cases_fr["I"],1600)).plot(figsize = (15,4),label = "Is")
pd.Series(cases_fr["I"]).plot(label = "I")
pd.Series(smooth(cases_fr["R"],1600)).plot(label = "Rs")
pd.Series(cases_fr["R"]).plot(label = "R")
plt.legend()
plt.show()
```
## Preparing SIR model
```
from covid.models import SIR
# Parameters
N = pop_fr
beta = 5/4
gamma = 1/4
start_date = cases_fr.index[0]
sir = SIR(N,beta,gamma)
states = sir.solve((N-1,1,0),start_date = start_date)
states.head()
states.show(plotly = False)
```
So of course parameters are not correct, in this version of the SIR model 30+ million persons get infected.<br>
Even because of test biases, estimates are more around 5 to 12% not 40%. <br>
Moreover, starting from first cases a peak was to be expected in mid-February, and in France lockdown started on the 10th of March.
```
states["I"].plot(figsize = (15,4))
cases_fr["I"].plot()
plt.show()
states["I"].plot(figsize = (15,4),label = "I_pred")
cases_fr["I"].plot(secondary_y = True,label = "I_true")
plt.legend()
plt.show()
```
## Some intuition about parameter sensibility
```
from ipywidgets import interact
@interact(beta = 5/4,gamma = 1/4)
def show_sir(beta,gamma):
# Create SIR model
N = pop_fr
start_date = cases_fr.index[0]
sir = SIR(N,beta,gamma)
states = sir.solve((N-1,1,0),start_date = start_date)
# Plot result
states["I"].plot(figsize = (15,2),label = "I_pred")
cases_fr["I"].plot(secondary_y = True,label = "I_true")
plt.legend()
plt.show()
states["I"].plot(figsize = (15,2),label = "I_pred")
cases_fr["I"].plot(label = "I_true")
plt.legend()
plt.show()
```
## Fitting parameters with hyperopt
### First attempt
##### References
- https://towardsdatascience.com/hyperparameter-optimization-in-python-part-2-hyperopt-5f661db91324
- http://hyperopt.github.io/hyperopt/
##### Space
```
from hyperopt import hp
space = {
"beta":hp.uniform('beta',0.1,5),
"gamma":hp.uniform('gamma',1/15,1/3),
}
```
##### Loss function between prediction and true
```
def loss_pred(states,true,cols = None):
if cols is None: cols = states.columns.tolist()
loss = 0
for col in cols:
loss = np.linalg.norm(states.loc[true.index,col].values - true[col].values)
return loss
loss_pred(states,cases_fr,cols = ["I","R"])
loss_pred(cases_fr,cases_fr,cols = ["I","R"])
```
##### Final loss function
```
def objective(params):
sir = SIR(N,params["beta"],params["gamma"])
states = sir.solve((N-1,1,0),start_date = start_date)
return loss_pred(states,cases_fr,cols = ["I","R"])
```
##### Hyperopt optimization
```
from hyperopt import fmin, tpe, Trials
trials = Trials()
best = fmin(
fn=objective,
space=space,
trials=trials,
algo=tpe.suggest,
max_evals=1000)
print(best)
```
##### Visualizing results
```
sir = SIR(N,best["beta"],best["gamma"])
states = sir.solve((N-1,1,0),start_date = start_date)
states["I"].plot(figsize = (15,2),label = "pred")
cases_fr["I"].plot(label = "true")
plt.legend()
plt.show()
states["R"].plot(figsize = (15,4),label = "pred")
cases_fr["R"].plot(label = "true")
plt.legend()
plt.show()
```
| github_jupyter |
# Venture Funding with Deep Learning
You work as a risk management associate at Alphabet Soup, a venture capital firm. Alphabet Soup’s business team receives many funding applications from startups every day. This team has asked you to help them create a model that predicts whether applicants will be successful if funded by Alphabet Soup.
The business team has given you a CSV containing more than 34,000 organizations that have received funding from Alphabet Soup over the years. With your knowledge of machine learning and neural networks, you decide to use the features in the provided dataset to create a binary classifier model that will predict whether an applicant will become a successful business. The CSV file contains a variety of information about these businesses, including whether or not they ultimately became successful.
## Instructions:
The steps for this challenge are broken out into the following sections:
* Prepare the data for use on a neural network model.
* Compile and evaluate a binary classification model using a neural network.
* Optimize the neural network model.
### Prepare the Data for Use on a Neural Network Model
Using your knowledge of Pandas and scikit-learn’s `StandardScaler()`, preprocess the dataset so that you can use it to compile and evaluate the neural network model later.
Open the starter code file, and complete the following data preparation steps:
1. Read the `applicants_data.csv` file into a Pandas DataFrame. Review the DataFrame, looking for categorical variables that will need to be encoded, as well as columns that could eventually define your features and target variables.
2. Drop the “EIN” (Employer Identification Number) and “NAME” columns from the DataFrame, because they are not relevant to the binary classification model.
3. Encode the dataset’s categorical variables using `OneHotEncoder`, and then place the encoded variables into a new DataFrame.
4. Add the original DataFrame’s numerical variables to the DataFrame containing the encoded variables.
> **Note** To complete this step, you will employ the Pandas `concat()` function that was introduced earlier in this course.
5. Using the preprocessed data, create the features (`X`) and target (`y`) datasets. The target dataset should be defined by the preprocessed DataFrame column “IS_SUCCESSFUL”. The remaining columns should define the features dataset.
6. Split the features and target sets into training and testing datasets.
7. Use scikit-learn's `StandardScaler` to scale the features data.
### Compile and Evaluate a Binary Classification Model Using a Neural Network
Use your knowledge of TensorFlow to design a binary classification deep neural network model. This model should use the dataset’s features to predict whether an Alphabet Soup–funded startup will be successful based on the features in the dataset. Consider the number of inputs before determining the number of layers that your model will contain or the number of neurons on each layer. Then, compile and fit your model. Finally, evaluate your binary classification model to calculate the model’s loss and accuracy.
To do so, complete the following steps:
1. Create a deep neural network by assigning the number of input features, the number of layers, and the number of neurons on each layer using Tensorflow’s Keras.
> **Hint** You can start with a two-layer deep neural network model that uses the `relu` activation function for both layers.
2. Compile and fit the model using the `binary_crossentropy` loss function, the `adam` optimizer, and the `accuracy` evaluation metric.
> **Hint** When fitting the model, start with a small number of epochs, such as 20, 50, or 100.
3. Evaluate the model using the test data to determine the model’s loss and accuracy.
4. Save and export your model to an HDF5 file, and name the file `AlphabetSoup.h5`.
### Optimize the Neural Network Model
Using your knowledge of TensorFlow and Keras, optimize your model to improve the model's accuracy. Even if you do not successfully achieve a better accuracy, you'll need to demonstrate at least two attempts to optimize the model. You can include these attempts in your existing notebook. Or, you can make copies of the starter notebook in the same folder, rename them, and code each model optimization in a new notebook.
> **Note** You will not lose points if your model does not achieve a high accuracy, as long as you make at least two attempts to optimize the model.
To do so, complete the following steps:
1. Define at least three new deep neural network models (the original plus 2 optimization attempts). With each, try to improve on your first model’s predictive accuracy.
> **Rewind** Recall that perfect accuracy has a value of 1, so accuracy improves as its value moves closer to 1. To optimize your model for a predictive accuracy as close to 1 as possible, you can use any or all of the following techniques:
>
> * Adjust the input data by dropping different features columns to ensure that no variables or outliers confuse the model.
>
> * Add more neurons (nodes) to a hidden layer.
>
> * Add more hidden layers.
>
> * Use different activation functions for the hidden layers.
>
> * Add to or reduce the number of epochs in the training regimen.
2. After finishing your models, display the accuracy scores achieved by each model, and compare the results.
3. Save each of your models as an HDF5 file.
```
# Imports
import pandas as pd
from pathlib import Path
import tensorflow as tf
from tensorflow.keras.layers import Dense
from tensorflow.keras.models import Sequential
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler,OneHotEncoder
# Google Colab imports.
import io
from google.colab import files
```
---
## Prepare the data to be used on a neural network model
### Step 1: Read the `applicants_data.csv` file into a Pandas DataFrame. Review the DataFrame, looking for categorical variables that will need to be encoded, as well as columns that could eventually define your features and target variables.
```
# Read the applicants_data.csv file from the Resources folder into a Pandas DataFrame
uploaded = files.upload()
applicant_data_df = pd.read_csv(io.BytesIO(uploaded["applicants_data.csv"]))
# Review the DataFrame
# YOUR CODE HERE
applicant_data_df.head(-5)
# Review the data types associated with the columns
# YOUR CODE HERE
applicant_data_df.dtypes
```
### Step 2: Drop the “EIN” (Employer Identification Number) and “NAME” columns from the DataFrame, because they are not relevant to the binary classification model.
```
# Drop the 'EIN' and 'NAME' columns from the DataFrame
applicant_data_df = applicant_data_df.drop(columns = ["EIN", "NAME"])
# Review the DataFrame
# YOUR CODE HERE
applicant_data_df.head(-5)
```
### Step 3: Encode the dataset’s categorical variables using `OneHotEncoder`, and then place the encoded variables into a new DataFrame.
```
# Create a list of categorical variables
categorical_variables = list(applicant_data_df.dtypes[applicant_data_df.dtypes == "object"].index)
# Display the categorical variables list
# YOUR CODE HERE
display(categorical_variables)
# Create a OneHotEncoder instance
enc = OneHotEncoder(sparse = False)
# Encode the categorcal variables using OneHotEncoder
encoded_data = enc.fit_transform(applicant_data_df[categorical_variables])
# Create a DataFrame with the encoded variables
encoded_df = pd.DataFrame(
encoded_data,
columns = enc.get_feature_names(categorical_variables)
)
# Review the DataFrame
encoded_df.head(-5)
```
### Step 4: Add the original DataFrame’s numerical variables to the DataFrame containing the encoded variables.
> **Note** To complete this step, you will employ the Pandas `concat()` function that was introduced earlier in this course.
```
# Add the numerical variables from the original DataFrame to the one-hot encoding DataFrame
numerical_variables = list(applicant_data_df.dtypes[applicant_data_df.dtypes != "object"].index)
encoded_df = pd.concat([encoded_df, applicant_data_df[numerical_variables]], axis = 1)
# Review the Dataframe
# YOUR CODE HERE
encoded_df.head(-5)
```
### Step 5: Using the preprocessed data, create the features (`X`) and target (`y`) datasets. The target dataset should be defined by the preprocessed DataFrame column “IS_SUCCESSFUL”. The remaining columns should define the features dataset.
```
# Define the target set y using the IS_SUCCESSFUL column
y = encoded_df["IS_SUCCESSFUL"]
# Display a sample of y
y
# Define features set X by selecting all columns but IS_SUCCESSFUL
X = encoded_df.drop(columns = ["IS_SUCCESSFUL"])
# Review the features DataFrame
X.head(-5)
```
### Step 6: Split the features and target sets into training and testing datasets.
```
# Split the preprocessed data into a training and testing dataset
# Assign the function a random_state equal to 1
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state = 1)
```
### Step 7: Use scikit-learn's `StandardScaler` to scale the features data.
```
# Create a StandardScaler instance
scaler = StandardScaler()
# Fit the scaler to the features training dataset
X_scaler = scaler.fit(X_train)
# Fit the scaler to the features training dataset
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.fit_transform(X_test)
```
---
## Compile and Evaluate a Binary Classification Model Using a Neural Network
### Step 1: Create a deep neural network by assigning the number of input features, the number of layers, and the number of neurons on each layer using Tensorflow’s Keras.
> **Hint** You can start with a two-layer deep neural network model that uses the `relu` activation function for both layers.
```
# Define the the number of inputs (features) to the model
number_input_features = len(X_train.loc[0])
# Review the number of features
number_input_features
# Define the number of neurons in the output layer
number_output_neurons = 1
# Define the number of hidden nodes for the first hidden layer
hidden_nodes_layer1 = 8
# Review the number hidden nodes in the first layer
hidden_nodes_layer1
# Define the number of hidden nodes for the second hidden layer
hidden_nodes_layer2 = 4
# Review the number hidden nodes in the second layer
hidden_nodes_layer2
# Create the Sequential model instance
nn = Sequential()
# Add the first hidden layer
# YOUR CODE HERE
nn.add(Dense(units = hidden_nodes_layer1, input_dim = number_input_features, activation = "relu"))
# Add the second hidden layer
# YOUR CODE HERE
nn.add(Dense(units = hidden_nodes_layer2, activation = "relu"))
# Add the output layer to the model specifying the number of output neurons and activation function
# YOUR CODE HERE
nn.add(Dense(units = number_output_neurons, activation = "sigmoid"))
# Display the Sequential model summary
# YOUR CODE HERE
nn.summary()
```
### Step 2: Compile and fit the model using the `binary_crossentropy` loss function, the `adam` optimizer, and the `accuracy` evaluation metric.
```
# Compile the Sequential model
# YOUR CODE HERE
nn.compile(loss = "binary_crossentropy", optimizer = "adam", metrics = ["accuracy"])
# Fit the model using 50 epochs and the training data
# YOUR CODE HERE
nn.fit(X_train_scaled, y_train, epochs = 50)
```
### Step 3: Evaluate the model using the test data to determine the model’s loss and accuracy.
```
# Evaluate the model loss and accuracy metrics using the evaluate method and the test data
model_loss, model_accuracy = nn.evaluate(X_test_scaled, y_test, verbose = 2)
# Display the model loss and accuracy results
print(f"Loss: {model_loss}, Accuracy: {model_accuracy}")
```
### Step 4: Save and export your model to an HDF5 file, and name the file `AlphabetSoup.h5`.
```
# Set the model's file path
file_path = Path("./H5/AlphabetSoup.h5")
# Export your model to a HDF5 file
nn.save(file_path)
```
---
## Optimize the neural network model
### Step 1: Define at least three new deep neural network models (resulting in the original plus 3 optimization attempts). With each, try to improve on your first model’s predictive accuracy.
> **Rewind** Recall that perfect accuracy has a value of 1, so accuracy improves as its value moves closer to 1. To optimize your model for a predictive accuracy as close to 1 as possible, you can use any or all of the following techniques:
>
> * Adjust the input data by dropping different features columns to ensure that no variables or outliers confuse the model.
>
> * Add more neurons (nodes) to a hidden layer.
>
> * Add more hidden layers.
>
> * Use different activation functions for the hidden layers.
>
> * Add to or reduce the number of epochs in the training regimen.
### Alternative Model 1
```
# Define the the number of inputs (features) to the model
number_input_features = len(X_train.iloc[0])
# Review the number of features
number_input_features
# Define the number of neurons in the output layer
number_output_neurons_A1 = 1
# Define the number of hidden nodes for the first hidden layer
hidden_nodes_layer1_A1 = 8
hidden_nodes_layer2_A1 = 6
hidden_nodes_layer3_A1 = 4
# Review the number of hidden nodes in the first layer
print(f"Hidden nodes (first layer): {hidden_nodes_layer1_A1}")
print(f"Hidden nodes (second layer): {hidden_nodes_layer2_A1}")
print(f"Hidden nodes (third layer): {hidden_nodes_layer3_A1}")
# Create the Sequential model instance
nn_A1 = Sequential()
# First hidden layer
nn_A1.add(Dense(units = hidden_nodes_layer1_A1, input_dim = number_input_features, activation = "relu"))
nn_A1.add(Dense(units = hidden_nodes_layer2_A1, activation = "relu"))
nn_A1.add(Dense(units = hidden_nodes_layer3_A1, activation = "relu"))
# Output layer
nn_A1.add(Dense(units = number_output_neurons, activation = "sigmoid"))
# Check the structure of the model
nn_A1.summary()
# Compile the Sequential model
nn_A1.compile(loss = "binary_crossentropy", optimizer = "adam", metrics = ["accuracy"])
# Fit the model using 50 epochs and the training data
fit_model_A1 = nn_A1.fit(X_train_scaled, y_train, epochs = 50)
```
#### Alternative Model 2
```
# Define the the number of inputs (features) to the model
number_input_features = len(X_train.iloc[0])
# Review the number of features
number_input_features
# Define the number of neurons in the output layer
number_output_neurons_A2 = 1 # YOUR CODE HERE
# Define the number of hidden nodes for the first hidden layer
hidden_nodes_layer1_A2 = 8
hidden_nodes_layer2_A2 = 6
hidden_nodes_layer3_A2 = 4
# Review the number of hidden nodes in the first, second and third layer.
print(f"Hidden nodes (first layer): {hidden_nodes_layer1_A2}")
print(f"Hidden nodes (second layer): {hidden_nodes_layer2_A2}")
print(f"Hidden nodes (third layer): {hidden_nodes_layer3_A2}")
# Create the Sequential model instance
nn_A2 = Sequential()
# First layer
nn_A2.add(Dense(units = hidden_nodes_layer1_A2, input_dim = number_input_features, activation = "selu"))
nn_A2.add(Dense(units = hidden_nodes_layer2_A2, activation = "selu"))
nn_A2.add(Dense(units = hidden_nodes_layer3_A2, activation = "selu"))
# Output layer
nn_A2.add(Dense(units = number_output_neurons, activation = "sigmoid"))
# Check the structure of the model
nn_A2.summary()
# Compile the model
# YOUR CODE HERE
nn_A2.compile(loss = "binary_crossentropy", optimizer = "adam", metrics = ["accuracy"])
# Fit the model
# YOUR CODE HERE
nn_A2.fit(X_train_scaled, y_train, epochs = 50)
```
### Step 2: After finishing your models, display the accuracy scores achieved by each model, and compare the results.
```
print("Original Model Results")
# Evaluate the model loss and accuracy metrics using the evaluate method and the test data
model_loss, model_accuracy = nn.evaluate(X_test_scaled, y_test, verbose = 2)
# Display the model loss and accuracy results
print(f"Loss: {model_loss}, Accuracy: {model_accuracy}")
print("Alternative Model 1 Results")
# Evaluate the model loss and accuracy metrics using the evaluate method and the test data
model_loss, model_accuracy = nn_A1.evaluate(X_test_scaled, y_test, verbose = 2)
# Display the model loss and accuracy results
print(f"Loss: {model_loss}, Accuracy: {model_accuracy}")
print("Alternative Model 2 Results")
# Evaluate the model loss and accuracy metrics using the evaluate method and the test data
model_loss, model_accuracy = nn_A2.evaluate(X_test_scaled, y_test, verbose = 2)
# Display the model loss and accuracy results
print(f"Loss: {model_loss}, Accuracy: {model_accuracy}")
```
### Step 3: Save each of your alternative models as an HDF5 file.
```
# Set the file path for the first alternative model
file_path = Path("./H5/AlphabetSoup_A1.h5")
# Export your model to a HDF5 file
nn_A1.save(file_path)
# Set the file path for the second alternative model
file_path = Path("./H5/AlphabetSoup_A2.h5")
# Export your model to a HDF5 file
nn_A2.save(file_path)
```
| github_jupyter |
##### Copyright 2019 Google LLC.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Spherical Harmonics Approximation
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/graphics/blob/master/tensorflow_graphics/notebooks/spherical_harmonics_approximation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/graphics/blob/master/tensorflow_graphics/notebooks/spherical_harmonics_approximation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
This Colab covers an advanced topic and hence focuses on providing controllable
toy examples to form a high level understanding of Spherical Harmonics and their
use for lighting rather than providing step by step details. For those
interested, these details are nevertheless available in the code. A great
resource to form a good understanding of Spherical Harmonics and their use for
lighting is
[Spherical Harmonics Lighting: the Gritty Details](http://silviojemma.com/public/papers/lighting/spherical-harmonic-lighting.pdf).
This Colab demonstrates how to approximate functions defined over a sphere using
Spherical Harmonics. These can be used to approximate lighting and
[reflectance](https://colab.research.google.com/github/tensorflow/graphics/blob/master/tensorflow_graphics/notebooks/reflectance.ipynb),
leading to very efficient rendering.
In more details, the following cells demonstrate:
* Approximation of lighting environments with Spherical Harmonics (SH)
* Approximation of the Lambertian BRDF with Zonal Harmonics (ZH)
* Rotation of Zonal Harmonics
* Rendering via Spherical Harmonics convolution of the SH lighting and ZH BRDF
## Setup & Imports
If Tensorflow Graphics is not installed on your system, the following cell can install the Tensorflow Graphics package for you.
```
!pip install tensorflow_graphics
```
Now that Tensorflow Graphics is installed, let's import everything needed to run the demo contained in this notebook.
```
###########
# Imports #
###########
import math
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from tensorflow_graphics.geometry.representation import grid
from tensorflow_graphics.geometry.representation import ray
from tensorflow_graphics.geometry.representation import vector
from tensorflow_graphics.rendering.camera import orthographic
from tensorflow_graphics.math import spherical_harmonics
from tensorflow_graphics.math import math_helpers as tf_math
tf.compat.v1.enable_eager_execution()
```
## Approximation of lighting with Spherical Harmonics
```
#@title Controls { vertical-output: false, run: "auto" }
max_band = 2 #@param { type: "slider", min: 0, max: 10 , step: 1 }
#########################################################################
# This cell creates a lighting function which we approximate with an SH #
#########################################################################
def image_to_spherical_coordinates(image_width, image_height):
pixel_grid_start = np.array((0, 0), dtype=type)
pixel_grid_end = np.array((image_width - 1, image_height - 1), dtype=type)
pixel_nb = np.array((image_width, image_height))
pixels = grid.generate(pixel_grid_start, pixel_grid_end, pixel_nb)
normalized_pixels = pixels / (image_width - 1, image_height - 1)
spherical_coordinates = tf_math.square_to_spherical_coordinates(
normalized_pixels)
return spherical_coordinates
def light_function(theta, phi):
theta = tf.convert_to_tensor(theta)
phi = tf.convert_to_tensor(phi)
zero = tf.zeros_like(theta)
return tf.maximum(zero,
-4.0 * tf.sin(theta - np.pi) * tf.cos(phi - 2.5) - 3.0)
light_image_width = 30
light_image_height = 30
type = np.float64
# Builds the pixels grid and compute corresponding spherical coordinates.
spherical_coordinates = image_to_spherical_coordinates(light_image_width,
light_image_height)
theta = spherical_coordinates[:, :, 1]
phi = spherical_coordinates[:, :, 2]
# Samples the light function.
sampled_light_function = light_function(theta, phi)
ones_normal = tf.ones_like(theta)
spherical_coordinates_3d = tf.stack((ones_normal, theta, phi), axis=-1)
samples_direction_to_light = tf_math.spherical_to_cartesian_coordinates(
spherical_coordinates_3d)
# Samples the SH.
l, m = spherical_harmonics.generate_l_m_permutations(max_band)
l = tf.convert_to_tensor(l)
m = tf.convert_to_tensor(m)
l_broadcasted = tf.broadcast_to(l, [light_image_width, light_image_height] +
l.shape.as_list())
m_broadcasted = tf.broadcast_to(m, [light_image_width, light_image_height] +
l.shape.as_list())
theta = tf.expand_dims(theta, axis=-1)
theta_broadcasted = tf.broadcast_to(
theta, [light_image_width, light_image_height, 1])
phi = tf.expand_dims(phi, axis=-1)
phi_broadcasted = tf.broadcast_to(phi, [light_image_width, light_image_height, 1])
sh_coefficients = spherical_harmonics.evaluate_spherical_harmonics(
l_broadcasted, m_broadcasted, theta_broadcasted, phi_broadcasted)
sampled_light_function_broadcasted = tf.expand_dims(
sampled_light_function, axis=-1)
sampled_light_function_broadcasted = tf.broadcast_to(
sampled_light_function_broadcasted,
[light_image_width, light_image_height] + l.shape.as_list())
# Integrates the light function times SH over the sphere.
projection = sh_coefficients * sampled_light_function_broadcasted * 4.0 * math.pi / (
light_image_width * light_image_height)
light_coeffs = tf.reduce_sum(projection, (0, 1))
# Reconstructs the image.
reconstructed_light_function = tf.squeeze(
vector.dot(sh_coefficients, light_coeffs))
print(
"average l2 reconstruction error ",
np.linalg.norm(sampled_light_function - reconstructed_light_function) /
(light_image_width * light_image_height))
vmin = np.minimum(
np.amin(np.minimum(sampled_light_function, reconstructed_light_function)),
0.0)
vmax = np.maximum(
np.amax(np.maximum(sampled_light_function, reconstructed_light_function)),
1.0)
# Plots results.
plt.figure(figsize=(10, 10))
ax = plt.subplot("131")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Original lighting function")
_ = ax.imshow(sampled_light_function, vmin=vmin, vmax=vmax)
ax = plt.subplot("132")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Spherical Harmonics approximation")
_ = ax.imshow(reconstructed_light_function, vmin=vmin, vmax=vmax)
ax = plt.subplot("133")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Difference")
_ = ax.imshow(
np.abs(reconstructed_light_function - sampled_light_function),
vmin=vmin,
vmax=vmax)
```
## Approximates the Lambertian BRDF with Zonal Harmonics
```
#################################################################
# This cell creates an SH that approximates the Lambertian BRDF #
#################################################################
# The image dimensions control how many uniform samples we draw from the BRDF.
brdf_image_width = 30
brdf_image_height = 30
type = np.float64
# Builds the pixels grid and compute corresponding spherical coordinates.
spherical_coordinates = image_to_spherical_coordinates(brdf_image_width,
brdf_image_height)
# Samples the BRDF function.
cos_theta = tf.cos(spherical_coordinates[:, :, 1])
sampled_brdf = tf.maximum(tf.zeros_like(cos_theta), cos_theta / np.pi)
# Samples the zonal SH.
l, m = spherical_harmonics.generate_l_m_zonal(max_band)
l_broadcasted = tf.broadcast_to(l, [brdf_image_width, brdf_image_height] +
l.shape.as_list())
m_broadcasted = tf.broadcast_to(m, [brdf_image_width, brdf_image_height] +
l.shape.as_list())
theta = tf.expand_dims(spherical_coordinates[:, :, 1], axis=-1)
theta_broadcasted = tf.broadcast_to(
theta, [brdf_image_width, brdf_image_height, 1])
phi = tf.expand_dims(spherical_coordinates[:, :, 2], axis=-1)
phi_broadcasted = tf.broadcast_to(phi, [brdf_image_width, brdf_image_height, 1])
sh_coefficients = spherical_harmonics.evaluate_spherical_harmonics(
l_broadcasted, m_broadcasted, theta_broadcasted, phi_broadcasted)
sampled_brdf_broadcasted = tf.expand_dims(sampled_brdf, axis=-1)
sampled_brdf_broadcasted = tf.broadcast_to(
sampled_brdf_broadcasted,
[brdf_image_width, brdf_image_height] + l.shape.as_list())
# Integrates the BRDF function times SH over the sphere.
projection = sh_coefficients * sampled_brdf_broadcasted * 4.0 * math.pi / (
brdf_image_width * brdf_image_height)
brdf_coeffs = tf.reduce_sum(projection, (0, 1))
# Reconstructs the image.
reconstructed_brdf = tf.squeeze(vector.dot(sh_coefficients, brdf_coeffs))
print(
"average l2 reconstruction error ",
np.linalg.norm(sampled_brdf - reconstructed_brdf) /
(brdf_image_width * brdf_image_height))
vmin = np.minimum(np.amin(np.minimum(sampled_brdf, reconstructed_brdf)), 0.0)
vmax = np.maximum(
np.amax(np.maximum(sampled_brdf, reconstructed_brdf)), 1.0 / np.pi)
# Plots results.
plt.figure(figsize=(10, 10))
ax = plt.subplot("131")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Original reflectance function")
_ = ax.imshow(sampled_brdf, vmin=vmin, vmax=vmax)
ax = plt.subplot("132")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Zonal Harmonics approximation")
_ = ax.imshow(reconstructed_brdf, vmin=vmin, vmax=vmax)
ax = plt.subplot("133")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Difference")
_ = ax.imshow(np.abs(sampled_brdf - reconstructed_brdf), vmin=vmin, vmax=vmax)
plt.figure(figsize=(10, 5))
plt.plot(
spherical_coordinates[:, 0, 1],
sampled_brdf[:, 0],
label="max(0,cos(x) / pi)")
plt.plot(
spherical_coordinates[:, 0, 1],
reconstructed_brdf[:, 0],
label="SH approximation")
plt.title("Approximation quality")
plt.legend()
plt.show()
```
## Rotation of Zonal Harmonics
```
###############################
# Rotation of zonal harmonics #
###############################
r_theta = tf.constant(np.pi / 2, shape=(1,), dtype=brdf_coeffs.dtype)
r_phi = tf.constant(0.0, shape=(1,), dtype=brdf_coeffs.dtype)
rotated_zonal_coefficients = spherical_harmonics.rotate_zonal_harmonics(
brdf_coeffs, r_theta, r_phi)
# Builds the pixels grid and compute corresponding spherical coordinates.
pixel_grid_start = np.array((0, 0), dtype=type)
pixel_grid_end = np.array((brdf_image_width - 1, brdf_image_height - 1),
dtype=type)
pixel_nb = np.array((brdf_image_width, brdf_image_height))
pixels = grid.generate(pixel_grid_start, pixel_grid_end, pixel_nb)
normalized_pixels = pixels / (brdf_image_width - 1, brdf_image_height - 1)
spherical_coordinates = tf_math.square_to_spherical_coordinates(
normalized_pixels)
# reconstruction.
l, m = spherical_harmonics.generate_l_m_permutations(max_band)
l_broadcasted = tf.broadcast_to(
l, [light_image_width, light_image_height] + l.shape.as_list())
m_broadcasted = tf.broadcast_to(
m, [light_image_width, light_image_height] + l.shape.as_list())
theta = tf.expand_dims(spherical_coordinates[:, :, 1], axis=-1)
theta_broadcasted = tf.broadcast_to(
theta, [light_image_width, light_image_height, 1])
phi = tf.expand_dims(spherical_coordinates[:, :, 2], axis=-1)
phi_broadcasted = tf.broadcast_to(
phi, [light_image_width, light_image_height, 1])
sh_coefficients = spherical_harmonics.evaluate_spherical_harmonics(
l_broadcasted, m_broadcasted, theta_broadcasted, phi_broadcasted)
reconstructed_rotated_brdf_function = tf.squeeze(
vector.dot(sh_coefficients, rotated_zonal_coefficients))
plt.figure(figsize=(10, 10))
ax = plt.subplot("121")
ax.set_title("Zonal SH")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
_ = ax.imshow(reconstructed_brdf)
ax = plt.subplot("122")
ax.set_title("Rotated version")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
_ = ax.imshow(reconstructed_rotated_brdf_function)
```
## Reconstruction via Spherical Harmonics convolution of the SH lighting and ZH BRDF
```
############################################################################################
# Helper function allowing to estimate sphere normal and depth for each pixel in the image #
############################################################################################
def compute_intersection_normal_sphere(image_width, image_height, sphere_radius,
sphere_center, type):
pixel_grid_start = np.array((0.5, 0.5), dtype=type)
pixel_grid_end = np.array((image_width - 0.5, image_height - 0.5), dtype=type)
pixel_nb = np.array((image_width, image_height))
pixels = grid.generate(pixel_grid_start, pixel_grid_end, pixel_nb)
pixel_ray = tf.math.l2_normalize(orthographic.ray(pixels), axis=-1)
zero_depth = np.zeros([image_width, image_height, 1])
pixels_3d = orthographic.unproject(pixels, zero_depth)
intersections_points, normals = ray.intersection_ray_sphere(
sphere_center, sphere_radius, pixel_ray, pixels_3d)
return intersections_points[0, :, :, :], normals[0, :, :, :]
###############################
# Setup the image, and sphere #
###############################
# Image dimensions
image_width = 100
image_height = 80
# Sphere center and radius
sphere_radius = np.array((30.0,), dtype=type)
sphere_center = np.array((image_width / 2.0, image_height / 2.0, 100.0),
dtype=type)
# Builds the pixels grid and compute corresponding spherical coordinates.
pixel_grid_start = np.array((0, 0), dtype=type)
pixel_grid_end = np.array((image_width - 1, image_height - 1), dtype=type)
pixel_nb = np.array((image_width, image_height))
pixels = grid.generate(pixel_grid_start, pixel_grid_end, pixel_nb)
normalized_pixels = pixels / (image_width - 1, image_height - 1)
spherical_coordinates = tf_math.square_to_spherical_coordinates(
normalized_pixels)
################################################################################################
# For each pixel in the image, estimate the corresponding surface point and associated normal. #
################################################################################################
intersection_3d, surface_normal = compute_intersection_normal_sphere(
image_width, image_height, sphere_radius, sphere_center, type)
surface_normals_spherical_coordinates = tf_math.cartesian_to_spherical_coordinates(
surface_normal)
# SH
l, m = spherical_harmonics.generate_l_m_permutations(
max_band) # recomputed => optimize
l = tf.convert_to_tensor(l)
m = tf.convert_to_tensor(m)
l_broadcasted = tf.broadcast_to(l,
[image_width, image_height] + l.shape.as_list())
m_broadcasted = tf.broadcast_to(m,
[image_width, image_height] + l.shape.as_list())
#################################################
# Estimates result using SH convolution - cheap #
#################################################
sh_integration = spherical_harmonics.integration_product(
light_coeffs,
spherical_harmonics.rotate_zonal_harmonics(
brdf_coeffs,
tf.expand_dims(surface_normals_spherical_coordinates[:, :, 1], axis=-1),
tf.expand_dims(surface_normals_spherical_coordinates[:, :, 2],
axis=-1)),
keepdims=False)
# Sets pixels not belonging to the sphere to 0.
sh_integration = tf.where(
tf.greater(intersection_3d[:, :, 2], 0.0), sh_integration,
tf.zeros_like(sh_integration))
# Sets pixels with negative light to 0.
sh_integration = tf.where(
tf.greater(sh_integration, 0.0), sh_integration,
tf.zeros_like(sh_integration))
###########################################
# 'Brute force' solution - very expensive #
###########################################
factor = 4.0 * np.pi / (light_image_width * light_image_height)
gt = tf.einsum(
"hwn,uvn->hwuv", surface_normal,
samples_direction_to_light *
tf.expand_dims(sampled_light_function, axis=-1))
gt = tf.maximum(gt, 0.0) # removes negative dot products
gt = tf.reduce_sum(gt, axis=(2, 3))
# Sets pixels not belonging to the sphere to 0.
gt = tf.where(tf.greater(intersection_3d[:, :, 2], 0.0), gt, tf.zeros_like(gt))
gt *= factor
# TODO(b/124463095): gt and sh_integration differ by a factor of pi.
sh_integration = np.transpose(sh_integration, (1, 0))
gt = np.transpose(gt, (1, 0))
plt.figure(figsize=(10, 20))
ax = plt.subplot("121")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("SH light and SH BRDF")
_ = ax.imshow(sh_integration, vmin=0.0)
ax = plt.subplot("122")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("GT light and GT BRDF")
_ = ax.imshow(gt, vmin=0.0)
```
| github_jupyter |
# Sheep Coordinates and Ensembl
Ensembl stores information regarding SNPs on sheep assembly 3.1. I want to verify if such information are updated with snpchimp or not
```
import os
import itertools
import random
from pathlib import Path
from importlib import reload
from ensemblrest import EnsemblRest
import src.features.illumina
import src.features.snpchimp
random.seed = 42
```
Start by reading snpchimp and snpchip illumina data from files
```
project_dir = Path.cwd().parents[1]
illumina_chip = project_dir / "data/external/SHE/ILLUMINA/ovinesnp50-genome-assembly-oar-v3-1.csv"
snpchimp_file = project_dir / "data/external/SHE/SNPCHIMP/SNPchimp_SHE_SNP50v1_oar3.1.csv"
new_chip3 = dict()
for record in src.features.illumina.read_Manifest(illumina_chip):
new_chip3[record.name] = (record.chr, record.mapinfo)
snpchimp3 = dict()
for record in src.features.snpchimp.read_snpChimp(snpchimp_file):
snpchimp3[record.snp_name] = [(record.chromosome, record.position), record.rs]
```
Ok, first track snp ids with different positions in both files. Skip SNP without rsID
```
count = 0
rs_different = {}
for i, (key, value) in enumerate(new_chip3.items()):
if value != snpchimp3[key][0]:
count += 1
if snpchimp3[key][1] != 'NULL':
rs_different[snpchimp3[key][1]] = key
if i < 10:
print(key, value, snpchimp3[key])
elif i == 10:
print("...")
print(f"\nN of SNPs in different positions from illumina to SNPchimp: {count}")
print(f"\nN of SNPs with rsID with different positions from illumina to SNPchimp: {len(rs_different)}")
```
Select some snps to test coordinates with ensembl
```
# filter out None rs
rs_keys = [rs for rs in rs_different.keys() if rs is not None]
selected_rsID = random.sample(sorted(rs_keys), 10)
# add a custom rs
selected_rsID.append("rs408606108")
```
Ok, search in Ensembl for snp coordinates:
```
ensRest = EnsemblRest()
result = ensRest.getVariationByMultipleIds(ids=selected_rsID, species="ovis_aries", genotyping_chips=True)
for key, value in result.items():
for location in value['mappings']:
print(key, location['seq_region_name'], location['start'], snpchimp3[rs_different[key]], new_chip3[rs_different[key]])
```
The coordinates are different: sometimes they are equal to SNPchimp, sometimes with illumina chip. Sometimes there are totally different. I think that it depends on the dbSNP version used for build or the align method in general.
When both SNPchimp and Illumina chip doesn't have coordinates, the SNP has a multiple aligment
<div class="alert alert-block alert-danger">
<b>Danger:</b> <code>rs408606108</code>, for example, failed the alignment with ensembl since the <code>C/T</code> allele doesn't match the reference genome (which is <code>T</code>). In EVA this snps is deprecated in Oarv4.0. Does this applies to any others SNPs? are they BOT/reverse? need to validate all SNPs data
</div>
| github_jupyter |
<img src="https://raw.githubusercontent.com/brazil-data-cube/code-gallery/master/img/logo-bdc.png" align="right" width="64"/>
# <span style="color:#336699">Introduction to the SpatioTemporal Asset Catalog (STAC)</span>
<hr style="border:2px solid #0077b9;">
<div style="text-align: left;">
<a href="https://nbviewer.jupyter.org/github/brazil-data-cube/code-gallery/blob/master/jupyter/Python/stac/stac-introduction.ipynb"><img src="https://raw.githubusercontent.com/jupyter/design/master/logos/Badges/nbviewer_badge.svg" align="center"/></a>
</div>
<br/>
<div style="text-align: center;font-size: 90%;">
Matheus Zaglia<sup><a href="https://orcid.org/0000-0001-6181-2158"><i class="fab fa-lg fa-orcid" style="color: #a6ce39"></i></a></sup>, Rennan Marujo<sup><a href="https://orcid.org/0000-0002-0082-9498"><i class="fab fa-lg fa-orcid" style="color: #a6ce39"></i></a></sup>, Gilberto R. Queiroz<sup><a href="https://orcid.org/0000-0001-7534-0219"><i class="fab fa-lg fa-orcid" style="color: #a6ce39"></i></a></sup>
<br/><br/>
Earth Observation and Geoinformatics Division, National Institute for Space Research (INPE)
<br/>
Avenida dos Astronautas, 1758, Jardim da Granja, São José dos Campos, SP 12227-010, Brazil
<br/><br/>
Contact: <a href="mailto:brazildatacube@inpe.br">brazildatacube@inpe.br</a>
<br/><br/>
Last Update: March 12, 2021
</div>
<br/>
<div style="text-align: justify; margin-left: 25%; margin-right: 25%;">
<b>Abstract.</b> This Jupyter Notebook gives an overview on how to use the STAC service to discover and access the data products from the <em>Brazil Data Cube</em>.
</div>
<br/>
<div style="text-align: justify; margin-left: 25%; margin-right: 25%;font-size: 75%; border-style: solid; border-color: #0077b9; border-width: 1px; padding: 5px;">
<b>This Jupyter Notebook is a supplement to the following paper:</b>
<div style="margin-left: 10px; margin-right: 10px">
Zaglia, M.; Vinhas, L.; Queiroz, G. R.; Simões, R. <a href="http://urlib.net/rep/8JMKD3MGPDW34R/3UFEFD8" target="_blank">Catalogação de Metadados do Cubo de Dados do Brasil com o SpatioTemporal Asset Catalog</a>. In: Proceedings XX GEOINFO, November 11-13, 2019, São José dos Campos, SP, Brazil. p 280-285.
</div>
</div>
# Introduction
<hr style="border:1px solid #0077b9;">
The [**S**patio**T**emporal **A**sset **C**atalog (STAC)](https://stacspec.org/) is a specification created through the colaboration of several organizations intended to increase satellite image search interoperability.
The diagram depicted in the picture contains the most important concepts behind the STAC data model:
<center>
<img src="https://raw.githubusercontent.com/brazil-data-cube/code-gallery/master/img/stac/stac-model.png" width="480" />
<br/>
<b>Figure 1</b> - STAC model.
</center>
The description of the concepts below are adapted from the [STAC Specification](https://github.com/radiantearth/stac-spec):
- **Item**: a `STAC Item` is the atomic unit of metadata in STAC, providing links to the actual `assets` (including thumbnails) that they represent. It is a `GeoJSON Feature` with additional fields for things like time, links to related entities and mainly to the assets. According to the specification, this is the atomic unit that describes the data to be discovered in a `STAC Catalog` or `Collection`.
- **Asset**: a `spatiotemporal asset` is any file that represents information about the earth captured in a certain space and time.
- **Catalog**: provides a structure to link various `STAC Items` together or even to other `STAC Catalogs` or `Collections`.
- **Collection:** is a specialization of the `Catalog` that allows additional information about a spatio-temporal collection of data.
# STAC Client API
<hr style="border:1px solid #0077b9;">
For running the examples in this Jupyter Notebook you will need to install the [STAC client for Python](https://github.com/brazil-data-cube/stac.py). To install it from PyPI using `pip`, use the following command:
```
#!pip install stac.py
```
In order to access the funcionalities of the client API, you should import the `stac` package, as follows:
```
import stac
```
After that, you can check the installed `stac` package version:
```
stac.__version__
```
Then, create a `STAC` object attached to the Brazil Data Cube' STAC service:
```
service = stac.STAC('https://brazildatacube.dpi.inpe.br/stac/', access_token='change-me')
```
# Listing the Available Data Products
<hr style="border:1px solid #0077b9;">
In the Jupyter environment, the `STAC` object will list the available image and data cube collections from the service:
```
service
```
or, access the `collections` property:
```
service.collections
```
# Retrieving the Metadata of a Data Product
<hr style="border:1px solid #0077b9;">
The `collection` method returns information about a given image or data cube collection identified by its name. In this example we are retrieving information about the datacube collection `CB4_64_16D_STK-1`:
```
collection = service.collection('CB4_64_16D_STK-1')
collection
```
# Retrieving Collection Items
<hr style="border:1px solid #0077b9;">
The `get_items` method returns the items of a given collection:
```
collection.get_items()
```
The `get_items` method also supports filtering rules through the specification of a rectangle (`bbox`) or a date and time (`datatime`) criterias:
```
items = collection.get_items(
filter={
'bbox':'-46.62597656250001,-13.19716452328198,-45.03570556640626,-12.297068292853805',
'datetime':'2018-08-01/2019-07-31',
'limit':10
}
)
items
```
From the item collection retrieved with the `get_item` method, it is possible to traverse the list of items:
```
for item in items:
print(item.id)
```
or, it is possible to use the index operador (`[]`) with the ``features`` property in order to retrieve a specific item from the collection:
```
item = items.features[0]
item.id
```
# Assets
<hr style="border:1px solid #0077b9;">
The assets with the links to the images, thumbnails or specific metadata files, can be accessed through the property `assets` (from a given item):
```
assets = item.assets
```
Then, from the assets it is possible to traverse or access individual elements:
```
for k in assets.keys():
print(k)
```
The metadata related to the CBERS-4/AWFI blue band is available under the dictionary key `BAND13`:
```
blue_asset = assets['BAND13']
blue_asset
```
To iterate in the item's assets, use the following pattern:
```
for asset in assets.values():
print(asset)
```
# Using RasterIO and NumPy
<hr style="border:1px solid #0077b9;">
The `rasterio` library can be used to read image files from the Brazil Data Cube' service on-the-fly and then to create `NumPy` arrays. The `read` method of an `Item` can be used to perform the reading and array creation:
```
nir = item.read('BAND16')
```
<div style="text-align: justify; margin-left: 15%; margin-right: 15%; border-style: solid; border-color: #0077b9; border-width: 1px; padding: 5px;">
<b>Note:</b> If there are errors because of your pyproj version, you can run the code below as specified in <a href="https://rasterio.readthedocs.io/en/latest/faq.html#why-can-t-rasterio-find-proj-db-rasterio-from-pypi-versions-1-2-0" target="_blank">rasterio documentation</a> and try again:
import os
del os.environ['PROJ_LIB']
</div>
```
nir
```
The next cell code import the `Window` class from the `rasterio` library in order to retrieve a subset of an image and then create an array:
```
from rasterio.windows import Window
```
We can specify a subset of the image file (window or chunck) to be read. Let's read a range that starts on pixel (0, 0) with 500 x 500 and column 0 to column 500, for the spectral bands `red`, `green` and `blue`:
```
red = item.read('BAND15', window=Window(0, 0, 500, 500)) # Window(col_off, row_off, width, height)
green = item.read('BAND14', window=Window(0, 0, 500, 500))
blue = item.read('BAND13', window=Window(0, 0, 500, 500))
blue
```
# Using Matplotlib to Visualize Images
<hr style="border:1px solid #0077b9;">
The `Matplotlib` cab be used to plot the arrays read in the last section:
```
%matplotlib inline
from matplotlib import pyplot as plt
fig, (ax1, ax2, ax3) = plt.subplots(1,3, figsize=(12, 4))
ax1.imshow(red, cmap='gray')
ax2.imshow(green, cmap='gray')
ax3.imshow(blue, cmap='gray')
```
Using `Numpy` we can stack the previous arrays and use `Matplotlib` to plot a color image, but first we need to normalize their values:
```
import numpy
def normalize(array):
"""Normalizes numpy arrays into scale 0.0 - 1.0"""
array_min, array_max = array.min(), array.max()
return ((array - array_min)/(array_max - array_min))
rgb = numpy.dstack((normalize(red), normalize(green), normalize(blue)))
plt.imshow(rgb)
```
# Retrieving Image Files
<hr style="border:1px solid #0077b9;">
The file related to an asset can be retrieved through the `download` method. The cell code below shows ho to download the image file associated to the asset into a folder named `img`:
```
blue_asset.download('img')
```
In order to download all files related to an item, use the `Item.download` method:
```
item.download('images')
```
Note that the URL for a given asset can be retrieved by the property `href`:
```
blue_asset.href
```
# References
<hr style="border:1px solid #0077b9;">
- [Spatio Temporal Asset Catalog Specification](https://stacspec.org/)
- [Brazil Data Cube Python Client Library for STAC Service - GitHub Repository](https://github.com/brazil-data-cube/stac.py)
# See also the following Jupyter Notebooks
<hr style="border:1px solid #0077b9;">
* [NDVI calculation on images obtained through STAC](./stac-ndvi-calculation.ipynb)
* [Thresholding images obtained through STAC](./stac-image-threshold.ipynb)
* [Calculating Image Difference on images obtained through STAC](./stac-image-difference.ipynb)
| github_jupyter |
# Homework03: Topic Modeling with Latent Semantic Analysis
Latent Semantic Analysis (LSA) is a method for finding latent similarities between documents treated as a bag of words by using a low rank approximation. It is used for document classification, clustering and retrieval. For example, LSA can be used to search for prior art given a new patent application. In this homework, we will implement a small library for simple latent semantic analysis as a practical example of the application of SVD. The ideas are very similar to PCA. SVD is also used in recommender systems in an similar fashion (for an SVD-based recommender system library, see [Surpise](http://surpriselib.com).
We will implement a toy example of LSA to get familiar with the ideas. If you want to use LSA or similar methods for statistical language analysis, the most efficient Python libraries are probably [gensim](https://radimrehurek.com/gensim/) and [spaCy](https://spacy.io) - these also provide an online algorithm - i.e. the training information can be continuously updated. Other useful functions for processing natural language can be found in the [Natural Language Toolkit](http://www.nltk.org/).
**Note**: The SVD from scipy.linalg performs a full decomposition, which is inefficient since we only need to decompose until we get the first k singluar values. If the SVD from `scipy.linalg` is too slow, please use the `sparsesvd` function from the [sparsesvd](https://pypi.python.org/pypi/sparsesvd/) package to perform SVD instead. You can install in the usual way with
```
!pip install sparsesvd
```
Then import the following
```python
from sparsesvd import sparsesvd
from scipy.sparse import csc_matrix
```
and use as follows
```python
sparsesvd(csc_matrix(M), k=10)
```
**Exercise 1 (20 points)**. Calculating pairwise distance matrices.
Suppose we want to construct a distance matrix between the rows of a matrix. For example, given the matrix
```python
M = np.array([[1,2,3],[4,5,6]])
```
the distance matrix using Euclidean distance as the measure would be
```python
[[ 0.000 1.414 2.828]
[ 1.414 0.000 1.414]
[ 2.828 1.414 0.000]]
```
if $M$ was a collection of column vectors.
Write a function to calculate the pairwise-distance matrix given the matrix $M$ and some arbitrary distance function. Your functions should have the following signature:
```
def func_name(M, distance_func):
pass
```
0. Write a distance function for the Euclidean, squared Euclidean and cosine measures.
1. Write the function using looping for M as a collection of row vectors.
2. Write the function using looping for M as a collection of column vectors.
3. Wrtie the function using broadcasting for M as a collection of row vectors.
4. Write the function using broadcasting for M as a collection of column vectors.
For 3 and 4, try to avoid using transposition (but if you get stuck, there will be no penalty for using transposition). Check that all four functions give the same result when applied to the given matrix $M$.
```
import numpy as np
import scipy.linalg as la
import string
import pandas as pd
from scipy import stats
np.set_printoptions(precision=4)
def Euc(x, y):
return np.sqrt(np.sum((x - y) ** 2))
def sqEuc(x, y):
return np.sum((x - y) ** 2)
def Cos(x, y):
return np.dot(x.T, y)/(np.linalg.norm(x) * np.linalg.norm(y))
M = np.array([[1,2,3],[4,5,6]])
def loop_row(M, distance_func):
n = M.shape[0]
dist = np.zeros((n, n))
for i in range(n):
for j in range(i + 1, n):
dist[i, j] = dist[j, i] = distance_func(M[i, :], M[j, :])
return dist
def loop_col(M, distance_func):
return loop_row(M.T, distance_func)
def broadcast_row(M, distance_func):
dist = np.sum(M ** 2, axis = 1) + np.sum(M ** 2, axis = 1)[:, np.newaxis] - 2 * np.dot(M, M.T)
return dist
broadcast_row(M, Euc)
```
**Exercise 2 (20 points)**.
**Exercise 2 (20 points)**. Write 3 functions to calculate the term frequency (tf), the inverse document frequency (idf) and the product (tf-idf). Each function should take a single argument `docs`, which is a dictionary of (key=identifier, value=document text) pairs, and return an appropriately sized array. Convert '-' to ' ' (space), remove punctuation, convert text to lowercase and split on whitespace to generate a collection of terms from the document text.
- tf = the number of occurrences of term $i$ in document $j$
- idf = $\log \frac{n}{1 + \text{df}_i}$ where $n$ is the total number of documents and $\text{df}_i$ is the number of documents in which term $i$ occurs.
Print the table of tf-idf values for the following document collection
```
s1 = "The quick brown fox"
s2 = "Brown fox jumps over the jumps jumps jumps"
s3 = "The the the lazy dog elephant."
s4 = "The the the the the dog peacock lion tiger elephant"
docs = {'s1': s1, 's2': s2, 's3': s3, 's4': s4}
```
```
def tf(docs):
doc_words = [doc.strip().lower().translate(str.maketrans('-', ' ', string.punctuation)).split()
for key, doc in docs.items()]
words = [word for words in doc_words for word in words]
terms = set(words)
table = np.zeros((len(terms), len(docs)), dtype = 'int')
for i, term in enumerate(terms):
for j, doc in enumerate(doc_words):
table[i, j] = doc.count(term)
df = pd.DataFrame(table, columns = docs.keys(), index=terms)
return df
def idf(docs):
doc_words = [doc.strip().lower().translate(str.maketrans('-', ' ', string.punctuation)).split()
for key, doc in docs.items()]
words = [word for words in doc_words for word in words]
terms = set(words)
table = np.zeros((len(terms)), dtype = 'int')
for i, term in enumerate(terms):
for doc in doc_words:
table[i] += int(term in doc)
table = np.log(len(docs) / (1 + table))
df = pd.DataFrame(table, columns=['idf'], index = terms)
return df
def tfidf(docs):
tf_tbl = tf(docs)
idf_tbl = idf(docs)
tfidf_tbl = pd.DataFrame(np.array(tf_tbl) * np.array(idf_tbl),columns = docs.keys(), index = idf_tbl.index)
return tfidf_tbl
s1 = "The quick brown fox"
s2 = "Brown fox jumps over the jumps jumps jumps"
s3 = "The the the lazy dog elephant."
s4 = "The the the the the dog peacock lion tiger elephant"
docs = {'s1': s1, 's2': s2, 's3': s3, 's4': s4}
print(tf(docs))
print(idf(docs))
print(tfidf(docs))
```
**Exercise 3 (20 points)**.
1. Write a function that takes a matrix $M$ and an integer $k$ as arguments, and reconstructs a reduced matrix using only the $k$ largest singular values. Use the `scipy.linagl.svd` function to perform the decomposition. This is the least squares approximation to the matrix $M$ in $k$ dimensions.
2. Apply the function you just wrote to the following term-frequency matrix for a set of $9$ documents using $k=2$ and print the reconstructed matrix $M'$.
```
M = np.array([[1, 0, 0, 1, 0, 0, 0, 0, 0],
[1, 0, 1, 0, 0, 0, 0, 0, 0],
[1, 1, 0, 0, 0, 0, 0, 0, 0],
[0, 1, 1, 0, 1, 0, 0, 0, 0],
[0, 1, 1, 2, 0, 0, 0, 0, 0],
[0, 1, 0, 0, 1, 0, 0, 0, 0],
[0, 1, 0, 0, 1, 0, 0, 0, 0],
[0, 0, 1, 1, 0, 0, 0, 0, 0],
[0, 1, 0, 0, 0, 0, 0, 0, 1],
[0, 0, 0, 0, 0, 1, 1, 1, 0],
[0, 0, 0, 0, 0, 0, 1, 1, 1],
[0, 0, 0, 0, 0, 0, 0, 1, 1]])
```
3. Calculate the pairwise correlation matrix for the original matrix M and the reconstructed matrix using $k=2$ singular values (you may use [scipy.stats.spearmanr](http://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.spearmanr.html) to do the calculations). Consider the fist 5 sets of documents as one group $G1$ and the last 4 as another group $G2$ (i.e. first 5 and last 4 columns). What is the average within group correlation for $G1$, $G2$ and the average cross-group correlation for G1-G2 using either $M$ or $M'$. (Do not include self-correlation in the within-group calculations.).
```
def reconstruct(M, k):
U, s, Vt = la.svd(M, full_matrices = False)
M_reduced = U[:, :k] @ np.diag(s[:k]) @ Vt[:k, :]
return M_reduced
M = np.array([[1, 0, 0, 1, 0, 0, 0, 0, 0],
[1, 0, 1, 0, 0, 0, 0, 0, 0],
[1, 1, 0, 0, 0, 0, 0, 0, 0],
[0, 1, 1, 0, 1, 0, 0, 0, 0],
[0, 1, 1, 2, 0, 0, 0, 0, 0],
[0, 1, 0, 0, 1, 0, 0, 0, 0],
[0, 1, 0, 0, 1, 0, 0, 0, 0],
[0, 0, 1, 1, 0, 0, 0, 0, 0],
[0, 1, 0, 0, 0, 0, 0, 0, 1],
[0, 0, 0, 0, 0, 1, 1, 1, 0],
[0, 0, 0, 0, 0, 0, 1, 1, 1],
[0, 0, 0, 0, 0, 0, 0, 1, 1]])
k = 2
Mp = reconstruct(M, k)
Mp
M_cor = stats.spearmanr(M).correlation
Mp_cor = stats.spearmanr(Mp).correlation
print(M_cor)
print(Mp_cor)
G1 = M[:, :5]
G2 = M[:, 5:]
G1_cor = stats.spearmanr(G1).correlation
G1_cor_mean = G1_cor[0, 1:].mean()
print(G1_cor_mean)
G2_cor = stats.spearmanr(G2).correlation
G2_cor_mean = G2_cor[0, 1:].mean()
print(G2_cor_mean)
G1_G2_cor = stats.spearmanr(G1, G2).correlation
G1_G2_cor_mean = G1_G2_cor[0, :].mean()
print(G1_G2_cor_mean)
```
**Exercise 4 (40 points)**. Clustering with LSA
1. Begin by loading a PubMed database of selected article titles using 'pickle'. With the following:
```import pickle
docs = pickle.load(open('pubmed.pic', 'rb'))```
Create a tf-idf matrix for every term that appears at least once in any of the documents. What is the shape of the tf-idf matrix?
2. Perform SVD on the tf-idf matrix to obtain $U \Sigma V^T$ (often written as $T \Sigma D^T$ in this context with $T$ representing the terms and $D$ representing the documents). If we set all but the top $k$ singular values to 0, the reconstructed matrix is essentially $U_k \Sigma_k V_k^T$, where $U_k$ is $m \times k$, $\Sigma_k$ is $k \times k$ and $V_k^T$ is $k \times n$. Terms in this reduced space are represented by $U_k \Sigma_k$ and documents by $\Sigma_k V^T_k$. Reconstruct the matrix using the first $k=10$ singular values.
3. Use agglomerative hierarchical clustering with complete linkage to plot a dendrogram and comment on the likely number of document clusters with $k = 100$. Use the dendrogram function from [SciPy ](https://docs.scipy.org/doc/scipy-0.15.1/reference/generated/scipy.cluster.hierarchy.dendrogram.html).
4. Determine how similar each of the original documents is to the new document `data/mystery.txt`. Since $A = U \Sigma V^T$, we also have $V = A^T U S^{-1}$ using orthogonality and the rule for transposing matrix products. This suggests that in order to map the new document to the same concept space, first find the tf-idf vector $v$ for the new document - this must contain all (and only) the terms present in the existing tf-idx matrix. Then the query vector $q$ is given by $v^T U_k \Sigma_k^{-1}$. Find the 10 documents most similar to the new document and the 10 most dissimilar.
```
import pickle
docs = pickle.load(open('pubmed.pic', 'rb'))
tfidf_df = tfidf(docs)
tfidf_m = np.array(tfidf_df)
tfidf_m.shape
tfidf_mp = reconstruct(tfidf_m, 10)
tfidf_mp
from matplotlib import pyplot as plt
from scipy.cluster.hierarchy import dendrogram, linkage
k = 100
Z = linkage(reconstruct(tfidf_m, k), 'complete')
plt.figure(figsize=(25, 10))
plt.title('Hierarchical Clustering Dendrogram')
plt.xlabel('index')
plt.ylabel('distance')
dendrogram(
Z,
truncate_mode = 'level',
p = 15,
leaf_font_size=15., # font size for the x axis labels
)
plt.show()
U, s, Vt = la.svd(tfidf_m, full_matrices = False)
with open("mystery.txt") as f:
newtext = f.read()
idf_df = idf(docs)
newtext_words = newtext.strip().lower().translate(str.maketrans('-', ' ', string.punctuation)).split()
terms = [term for term in idf_df.index]
tf_new = np.zeros((len(terms)), dtype = 'int')
for i, term in enumerate(terms):
tf_new[i] = newtext_words.count(term)
tfidf_new = np.array(idf_df) * tf_new.reshape((-1,1))
tfidf_new.T @ U[:, :k] @ la.inv(np.diag(s[:k]))
```
**Notes on the Pubmed articles**
These were downloaded with the following script.
```python
from Bio import Entrez, Medline
Entrez.email = "YOUR EMAIL HERE"
import cPickle
try:
docs = cPickle.load(open('pubmed.pic'))
except Exception, e:
print e
docs = {}
for term in ['plasmodium', 'diabetes', 'asthma', 'cytometry']:
handle = Entrez.esearch(db="pubmed", term=term, retmax=50)
result = Entrez.read(handle)
handle.close()
idlist = result["IdList"]
handle2 = Entrez.efetch(db="pubmed", id=idlist, rettype="medline", retmode="text")
result2 = Medline.parse(handle2)
for record in result2:
title = record.get("TI", None)
abstract = record.get("AB", None)
if title is None or abstract is None:
continue
docs[title] = '\n'.join([title, abstract])
print title
handle2.close()
cPickle.dump(docs, open('pubmed.pic', 'w'))
docs.values()
```
| github_jupyter |
### Visualizzare i 6 grafici
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import scipy
import sklearn
import seaborn as sns
import xlrd
import funzioni as fn
import statsmodels.api as sm
```
Per il sesso M=0 e F=1
```
data=pd.read_excel('Data/Mini Project EFSA.xlsx')
data.rename(columns={'sex \n(0=M, 1=F)':'sex'}, inplace=True)
data
```
## Grafici maschili
```
male_data=data[data.sex==0]
male_data
```
### Endpoint 1
```
male_data_1=male_data[male_data.endpoint==1]
male_data_1.plot(x='dose',y='response',yerr='SD',kind='bar',figsize=(12,6))
```
### Endpoint 2
```
male_data_2=male_data[male_data.endpoint==2]
male_data_2.plot(x='dose',y='response',yerr='SD',kind='bar',figsize=(12,6))
```
### Endpoint 3
```
male_data_3=male_data[male_data.endpoint==3]
male_data_3.plot(x='dose',y='response',yerr='SD',kind='bar',figsize=(12,6))
```
## Grafici femminili
```
female_data=data[data.sex==1]
female_data
```
### Endpoint 1
```
female_data_1=female_data[female_data.endpoint==1]
female_data_1.plot(x='dose',y='response',yerr='SD',kind='bar',figsize=(12,6))
```
### Endpoint 2
```
female_data_2=female_data[female_data.endpoint==2]
female_data_2.plot(x='dose',y='response',yerr='SD',kind='bar',figsize=(12,6))
```
### Endpoint 3
```
female_data_3=female_data[female_data.endpoint==3]
female_data_3.plot(x='dose',y='response',yerr='SD',kind='bar',figsize=(12,6))
```
## prove plot
```
data_Endpoint1 = data[data.endpoint == 1]
data_Endpoint2 = data[data.endpoint == 2]
data_Endpoint3 = data[data.endpoint == 3]
Y = data_Endpoint1.response
weights = data.SD
X = data_Endpoint1.drop(columns=["response", "SD", "endpoint"])
group_of_models_endpoint1 = fn.mainForward(X, Y, weights)
display(group_of_models_endpoint1)
pred=pd.DataFrame([data_Endpoint1['number of animals'],group_of_models_endpoint1.Y_pred[1]],index=["noa","y_pred"])
pred1=pred.T
pred=pd.DataFrame([data_Endpoint1['number of animals'],data_Endpoint1['sex'],group_of_models_endpoint1.Y_pred[2]],index=["noa","sex","y_pred"])
pred2=pred.T
pred=pd.DataFrame([data_Endpoint1['number of animals'],data_Endpoint1['sex'],data_Endpoint1['dose'],group_of_models_endpoint1.Y_pred[3]],index=["noa","sex","dose","y_pred"])
pred3=pred.T
pred3
fig, axs = plt.subplots(figsize=(15,20),nrows=3)
data_Endpoint1.plot(x='number of animals', y='response',s=100,marker='x', ax=axs[0],kind='scatter')
pred1.plot(x='noa', y='y_pred',color='r', ax=axs[0])
data_Endpoint1.plot(x=data_Endpoint1[['number of animals','sex']], y='response',s=100,marker='x', ax=axs[1],kind='scatter')
pred1.plot(x='noa', y='y_pred',color='r', ax=axs[0])
p = model1.params()
#Plot
x = np.arange(0,40)
ax = data_Endpoint1.plot(kind='scatter', x='number of animals', y='response')
ax.plot(x, p.)
ax.set_xlim([0,30])
#Seaborn
sns.lmplot(x='Xvalue', y='Yvalue', data=data)
group_of_models_endpoint1.plot(x = 'number_of_predictors', y = 'RSS')
#print(model1.summary())
#fig, ax = plt.subplots()
#fig = sm.graphics.plot_fit(model1, 0, ax=ax)
```
# prova push
| github_jupyter |
# Handling infeasible models with Docplex
This tutorial includes everything you need to set up Decision Optimization engines, build a mathematical programming model, then use the progress listeners to monitor progress, capture intermediate solutions and stop the solve on your own criteria.
When you finish this tutorial, you'll have a foundational knowledge of _Prescriptive Analytics_.
>This notebook is part of **[Prescriptive Analytics for Python](http://ibmdecisionoptimization.github.io/docplex-doc/)**
>
>It requires either an [installation of CPLEX Optimizers](http://ibmdecisionoptimization.github.io/docplex-doc/getting_started.html) or it can be run on [IBM Cloud Pak for Data as a Service](https://www.ibm.com/products/cloud-pak-for-data/as-a-service/) (Sign up for a [free IBM Cloud account](https://dataplatform.cloud.ibm.com/registration/stepone?context=wdp&apps=all>)
and you can start using `IBM Cloud Pak for Data as a Service` right away).
>
> CPLEX is available on <i>IBM Cloud Pack for Data</i> and <i>IBM Cloud Pak for Data as a Service</i>:
> - <i>IBM Cloud Pak for Data as a Service</i>: Depends on the runtime used:
> - <i>Python 3.x</i> runtime: Community edition
> - <i>Python 3.x + DO</i> runtime: full edition
> - <i>Cloud Pack for Data</i>: Community edition is installed by default. Please install `DO` addon in `Watson Studio Premium` for the full edition
Table of contents:
* [Use decision optimization](#Use-decision-optimization)
* [Step 1: Set up a basic infeasible model](#Step-1:-Set-up-the-prescriptive-model)
* [Step 2: Monitoring CPLEX progress](#Step-2:-Monitoring-CPLEX-progress)
* [Step 3: Aborting the search with a custom progress listener](#Step-3:-Aborting-the-search-with-a-custom-progress-listener)
* [Variant: using matplotlib to plot a chart of gap vs. time](#Variant:-using-matplotlib-to-plot-a-chart-of-gap-vs.-time)
* [Summary](#Summary)
****
## How Decision Optimization can help
* Prescriptive analytics (Decision Optimization) technology recommends actions that are based on desired outcomes. It takes into account specific scenarios, resources, and knowledge of past and current events. With this insight, your organization can make better decisions and have greater control of business outcomes.
* Prescriptive analytics is the next step on the path to insight-based actions. It creates value through synergy with predictive analytics, which analyzes data to predict future outcomes.
* Prescriptive analytics takes that insight to the next level by suggesting the optimal way to handle that future situation. Organizations that can act fast in dynamic conditions and make superior decisions in uncertain environments gain a strong competitive advantage.
<br/>
<u>With prescriptive analytics, you can:</u>
* Automate the complex decisions and trade-offs to better manage your limited resources.
* Take advantage of a future opportunity or mitigate a future risk.
* Proactively update recommendations based on changing events.
* Meet operational goals, increase customer loyalty, prevent threats and fraud, and optimize business processes.
```
from docplex.mp.environment import Environment
Environment().print_information()
```
## Example 1: handling a cyclic infeasible model.
You start with a very simple infeasible model: you have three variables, each of which is greater than the previous one, in a cyclic fashion. Of course this leads to an infeasible model
```
from docplex.mp.model import Model
def build_infeasible_cyclic_model3():
m = Model(name='cyclic3')
x,y,z = m.continuous_var_list(keys=['x', 'y', 'z'], name=str)
m.add( y >= x+1, name="y_gt_x")
m.add( z >= y+1, name="z_gt_y")
m.add( x >= z+1, name="x_gt_z")
# add another constraint, should noever appear in conflicts
m.add(x + y + z <= 33)
return m
cycle3 = build_infeasible_cyclic_model3()
cycle3.print_information()
```
As expected, the model is infeasible.
```
s = cycle3.solve(log_output=True)
assert s is None
print("the model is infeasible")
```
### Using the conflict refiner on the infeasible cyclic model
First, you can use the Conflict refiner on this model. The conflict refiner computes a minimal cluster of constraints, which causes the infeasibility.
Using the conflict refiner requires the following steps:
- instantiate a `ConflictRefiner` instance
- call `refine_conflict' on the model.
The output is an object of type `ConflictRefinerResults` which holds all information about the minimal conflict.
Displaying this result object lists all modeling objects which belong to the minimal conflict.
```
from docplex.mp.conflict_refiner import ConflictRefiner
cr = ConflictRefiner()
crr = cr.refine_conflict(cycle3, display=True)
```
### Using the constraint relaxer on the infeasible cyclic model
Another way to handle infeasibilities is to use the _relaxer_ (class `docplex.mp.relaxer.Relaxer`). The relaxer tries to find a _minimal_ feasible relaxation of the model, by relaxing certain constraints.
For example, a constraint `x == 1` can be relaxed with a slack of 1 to accept a value of 2 for x.
the relaxer tries to minimize the total value of slack in finding a feasible relaxation.
```
from docplex.mp.relaxer import Relaxer
rx = Relaxer()
rs = rx.relax(cycle3)
rx.print_information()
rs.display()
```
The relaxer has relaxed one constraint ( x>= z+1) by 3, and found a solution wiht x=0, x=1, z=2, breaking the cyclic chain of constraints.
Unlike the conflict refiner, the relaxer provides a _relaxed_ solution to the initial model, with minimal slack. But there's more: the relaxer can also search for the best business objective, once it has found the minimal slack.
To illustrate, add an objective to your model, try to minimize z, and see what happens:
```
# retrieve the z variable using Model.get_var_by_name()
z = cycle3.get_var_by_name('z')
assert z
cycle3.minimize(z)
rs = rx.relax(cycle3)
rx.print_information()
rs.display()
```
Th relaxed solution has changed, finding a minimum objective of 0 for z, and the relaxations have also changed: now two constraints are relaxed, but the total absolute slack remains unchanged, equal to 3.
To summarize, the relaxer finds a relaxed solution in two steps:
- first it finds the minimal amount of slack that is necessary to find a feasible solution
- second, with this minimal slack value it searches for the best objective value, if any.
In this case, the minimal conflict contains the three cyclic constraints, but not the fourth (x+y+z <= 30).
## Example 2: Handling an infeasible production problem
The model aims at minimizing the production cost for a number of products
while satisfying customer demand.
Production is constrained by the company's resources.
The model first declares the products and the resources.
The data consists of the description of the products (the demand, the inside
and outside costs, and the resource consumption) and the capacity of the
various resources.
The variables for this problem are the quantities produced for each products.
Of course, this model is naive, un-realistic and not robust, as it fails as soon as demand is not satisfied. But this is excatly why it is well adapted to show you how to repair infeasible models.
First define the data:
```
# costs are stored in a dict from product names to cost
COSTS = {"kluski": 0.6, "capellini": 0.8, "fettucine": 0.3}
# demands are stored in a dict from product names to demands
DEMANDS = {"kluski": 10, "capellini": 20, "fettucine": 30}
# resources are stored as a dict of resource name to resource capacity
RESOURCES = {"flour": 40, "eggs": 60}
CONSUMPTIONS = {("kluski", "flour"): 0.5,
("kluski", "eggs"): 0.2,
("capellini", "flour"): 0.4,
("capellini", "eggs"): 0.4,
("fettucine", "flour"): 0.3,
("fettucine", "eggs"): 0.6}
from docplex.mp.model import Model
import six
def build_production_problem(costs, resources, consumptions, demands, **kwargs):
products = [p for p, _ in six.iteritems(costs)]
mdl = Model(name='pasta_production', **kwargs)
# --- decision variables ---
mdl.q_vars = mdl.continuous_var_dict(products, name="q")
# --- constraints ---
# demand satisfaction
mdl.add_constraints((mdl.q_vars[p] >= demands[p], 'ct_demand_%s' % p) for p in products)
# --- resource capacity ---
mdl.add_constraints((mdl.sum(mdl.q_vars[p] * consumptions[p, res] for p in products) <= cap,
'ct_res_%s' % res) for res, cap in six.iteritems(resources))
# --- objective ---
mdl.minimize(mdl.dotf(mdl.q_vars, lambda p: costs[p]))
return mdl
pasta1 = build_production_problem(COSTS, RESOURCES, CONSUMPTIONS, DEMANDS)
pasta1.print_information()
```
This default model is feasible, solve the model
```
s1 = pasta1.solve()
s1.display()
demands2 = {p: 2*d for p, d in six.iteritems(DEMANDS)}
pasta2 = build_production_problem(COSTS, RESOURCES, CONSUMPTIONS, demands2)
s2 = pasta2.solve()
if s2 is None:
print("!! Pasta production with double demand is impossible")
```
Now double the demand.
### Using the conflict refiner on the production problem
Start by running the conflict refiner on the second production model.
```
from docplex.mp.conflict_refiner import ConflictRefiner
crr = ConflictRefiner().refine_conflict(pasta2, display=True)
crr.display()
```
Not surprisingly, you can see that the conflict involves all three demands but also the flour resource constraint, but we have no idea which quantity demands cannot be satisfied.
### Using the default relaxer on the production problem
The purpose of the relaxer is to allow relaxation of some constraints by a minimal amount to provide both a _relaxed solution_ and also a measure of how the constraints were infeasible.
```
from docplex.mp.relaxer import Relaxer
# create an instance of relaxer
rx = Relaxer()
rs = rx.relax(pasta2)
rx.print_information()
rs.display()
```
The relaxer managed to satisfy all demands by _relaxing_ the flour constraint by an amount of 4. What does this mean?
To explain, first remember what this flour constraint was all about. You can use the `Model.get_constraint_by_name` method to retrieve a constraint from its name.
```
# get back the constraint from its name
ctf = pasta2.get_constraint_by_name("ct_res_flour")
assert ctf is not None
print(str(ctf))
```
Now you can see what the _left hand side_ evaluates in the relaxed solution
```
ctf.lhs.solution_value
```
This explains the relaxation of 4 for the flour resource constraint
### Managing constraint priorities
It might well happen that the relaxation found by the relaxer does not make sense in real world. For example, in our production example, resource constraints can be impossible to relax, but demands could be.
This is where priorities enter the game. By setting priorities, users can control how the relaxer chooses constraints to relax. In the following code, you can set a HIGH priority to resource constraints (you could even make them mandatory) and a LOW priority to demand constraints.
```
from docplex.mp.basic import Priority
for ctr in pasta2.find_matching_linear_constraints("ct_res"):
ctr.priority = Priority.HIGH
for ctd in pasta2.find_matching_linear_constraints("ct_dem"):
ctd.priority = Priority.LOW
rx2 = Relaxer()
rs2 = rx2.relax(pasta2)
rx2.print_information()
rs2.display()
```
In this new relaxed solution, all resource constraints are satisfied, but one demand is not: kluski demand has an unfilled quantity of 8.
Setting constraint priorities explicitly is the most basic way to control relaxation, but there are others. A _function_ can be used: the relaxer will call the function for each constraint to determine its priority. Possible values are:
- relaxable priorities: VERY_LOW, LOW, MEDIUM, HIGH, VERY_HIGH
- non-relaxable priority: MANDATORY
Constraints with higher priority are less likely to be relaxed than constraints with lower priorities. Still, relaxation of a high-priority constraint cannot be ruled out, if it is the only way to provide a relaxed solution.
### Managing priorities with functions
In this section, you can see how to use a function to compute the priority of a constraint. The function must take a constraint and return a priority (an enumerated type, see `docplex.mp.basic.Priority`
First, you reset all priorities to None (the default).
```
# reset all priorities
for c in pasta2.iter_constraints():
c.priority = None
# define the constraint -> priority function
def map_priority(ct):
ctname = ct.name
if not ctname:
return Priority.MANDATORY
elif "ct_res" in ctname:
return Priority.HIGH
elif "ct_dem" in ctname:
return Priority.LOW
else:
# will not be relaxed
return Priority.MANDATORY
# create a new instance of Relaxer with this function.
rx3 = Relaxer(prioritizer=map_priority)
# use it to relax pasta2 model
rx3.relax(pasta2)
# display relaxation.
rx3.print_information()
```
As expected, you get the same result as with the explicit priorities: an unsatisfied demand of 8 for kluski.
Note that relaxer can also accept a _dictionary_ of constraints to priorities.
### The default relaxer revisited
Now that you know about setting priorities, you can understand the default behavior of the `relaxer` class: for each constraint, you use either its explicit priority (if set) or the default`MEDIUM` priority.
If no priority has been set, all constraints are considered equally relaxable.
## Example 3: Handling variable bounds in an infeasible model
### Variable bounds in conflict refiner
The conflict refiner takes into account variable bounds. This is illustrated with a very simple model, with three integer variables with lower bound 1, the sum of which should be less than 2.
```
m4 = Model(name='m4')
ijs = m4.integer_var_list(keys=["i", "j", "k", "l"], name =str, lb = 1)
m4.add(m4.sum(ijs) <= 2)
s4 = m4.solve()
assert s4 is None
ConflictRefiner().refine_conflict(m4, display=True);
```
The resulting conflict contains the sum constraint _and_ the three lower bounds.
### Variable bounds in relaxer
The relaxer only relaxes _constraints_ , so in this case it relaxes the sum constraint.
```
r4 = Relaxer()
r4.relax(m4)
r4.print_information()
```
Changing variable lower bounds to constraints allows the relaxer to take them into account.
You can set a *LOW* priority to lower bounds, so we expect the default relaxer to relax them before the sum constraint, which will be considered with the default MEDIUM priority.
```
for v in m4.iter_variables():
v.lb = 0
clb = m4.add(v >= 1, "{0}_ge_1".format(v.name))
clb.priority = Priority.LOW
r4 = Relaxer()
r4.relax(m4)
r4.print_information()
```
As expected, the relaxer has relaxed two lower bound constraints, but not the sum constraint.
## Summary
You have learned how to use both the conflict refiner and the relaxer, and the differences between them
- The conflict refiner lists constraints which are participating in the infeasibility. Constraints not mentioned in the conflict are not a problem.
- The conflict refiner considers both constraints and variable bounds.
- The conflict refiner does not provide any relaxed solution, nor any quantitative information.
In constrast, the relaxer provides a relaxed solution, and indicates which constraints are relaxed, and with what quantities. It does not consider variables bounds. It requires a mapping from constraints to Priority objects, which can take many forms: a function, a dictionary,...
#### References
* [Decision Optimization CPLEX Modeling for Python documentation](http://ibmdecisionoptimization.github.io/docplex-doc/)
* Need help with DOcplex or to report a bug? Please go [here](https://stackoverflow.com/questions/tagged/docplex)
* Contact us at dofeedback@wwpdl.vnet.ibm.com"
Copyright © 2017-2019 IBM. Sample Materials.
| github_jupyter |
# Natural Language Processing (NLP) with SciKit Learn
## Introduction to Natural Language Processing
Natural language processing is an automated way to understand and analyze natural human languages and extract information from such data by applying machine algorithms.

It is also referred to as, the field of computer science or AI to extract the linguistics information from the underlying data.
### Why Natural Language Processing
The world is now connected globally due to the advancement of technology and devices. this has resulted in high volume of data around the world and has led to a number of challenges in analysing data, such as
* Analyzing tons of data
* Identifying various languages
* Applying quantitative analysis
* Handling ambiguities
NLP can achieve full automation by using modern software libraries, modules, and packages.

### NLP Terminology
* __Word boundaries__:
* Determines where one word ends and the other begins
* __Tokenization__:
* Splits words, phrases, and idioms
* __Stemming__:
* Maps to the valid root word
* __Tf-idf_:
* Represents term frequency and inverse document frequency
* __Semantic analytics__:
* Compares words, phrases, and idioms in a set of documents to extract meaning
* __Disambiguation__:
* Determines meaning and sense of words (context vs. intent)
* __Topic models__:
* Discovers topics in a collection of documents
### NLP Approach for Text Data
Let us look at the Natural Language Processing approaches to analyze text data.
* Conduct basic text processing
* Categorize and tag words
* Classify text
* Extract information
* Analyze sentence structure
* Build feature-based structure
* Analyze the meaning
### Demo - 01: Demonstrate the installation of NLP environment
In this demo we will see how to install NLP environment.
* Open Anaconda prompt
* pip Intall sklearn
* pip install nltk
* Once installed, open python terminal in anaconda command prompt.
* Type import nlkt, and press enter.
* Type nltk.download()
* A window will prompt up, from there download required nltk subpackages like stopword etc,.
### Demo - 02: Demonstrate how to perform the sentence analysis
```
#Eliminate punctuation and stowords from the sentence
#Import required library
import string
from nltk.corpus import stopwords
#view first 10 stopwords present in English corpus
stopwords.words('english')[0:10]
#Create a test sentence
test_sentence = 'This is my first test string. wow!! we are doing hust fine.'
#eliminate the punctuation in-from of characters and print then
no_punctuation = [char for char in test_sentence if char not in string.punctuation]
no_punctuation
#now eliminate the punctuations and print them as a whole sentence.
no_punctuation = ''.join(no_punctuation)
no_punctuation
#split each words presentin the new sentence (remove stopwords)
no_punctuation.split()
#Eliminate stopwords
clean_sentence = [word for word in no_punctuation.split() if word.lower() not in stopwords.words('english')]
print(clean_sentence)
```
### Applications of NLP
1. __Machine Translation:__
Machine translation is used to translate one language into another. Google Translate is an example. It uses NLP to translate the input data from one language to another.
2. __Speech Recognition:__
The speech recognition application understands human speech and uses it as input information. It is useful for applications like Siri, Google Now, and Microsoft Cortana.
3. __Sentiment Analysis:__
Sentiment analysis is achieved by processing tons of data received from different interfaces and sources. For example, NLP uses all social media activities to find out the popular topic of discussion or importance.
### Major NLP Libraries
* NLTK
* Scikit-Learn
* TextBlob
* spaCy
## The Scikit-Learn Approach
It is a very powerful library with a set of modules to process and analyze natural language data, such as text and images, and extract information using machine learning algorithms.
* __Built-in module:__
Contains built-in modules to load the dataset’s content and categories.
* __Feature extraction:__
A way to extract information from data which can be text or images.
* __Model training:__
Analyzes the content based on particular categories and then trains them according to a specific model.
* __Pipeline building mechanism:__
A technique in Scikit-learn approach to streamline the NLP process into stages.
Various stages of pipeline learning:
1. Vectorization
2. Transformation
3. Model training and application
* __Performance optimization:__
In this stage we train the models to optimize the overall process.
* __Grid search for finding good parameters:__
It’s a powerful way to search parameters affecting the outcome for model training purposes.
### Modules to Load Content and Category
Scikit-learn has many built-in datasets. There are several methods to load these datasets with the help of a data load object.

The text files are loaded with categories as subfolder names.


The attributes of a data load object are:

The example shows how a dataset can be loaded using Scikit-learn:
__Example on Digits dataset to view the data using built-in methods.__
```
#Import dataset
from sklearn.datasets import load_digits
#Load the dataset
#create object of the loaded dataset
digit_dataset = load_digits()
#Describe the dataset
#USe built-in DESCR method to describe dataset
digit_dataset.DESCR
#View type of dataset
type(digit_dataset)
#View the data
digit_dataset.data
#View the target
digit_dataset.target
```
## Feature Extraction
Feature extraction is a technique to convert the content into the numerical vectors to perform machine learning.
1. __Text feature extraction:__
For example: Large datasets or documents
2. __Image feature extraction:__
For example: Patch extraction, hierarchical clustering
### Bag of Words
Bag of words is used to convert text data into numerical feature vectors with a fixed size.
__Step 1 - Tokenizing:__
Assign a fixed integer id to each word
__Step 2 - Counting:__
Number of occurrences of each word
__Step 3 - Storing:__
Store as the value feature.

### CountVectorizer Class Signature

### Demo - 03: Bags of words
Demonstrate the Bag of Words technique to process document
```
#import the required library
from sklearn.feature_extraction.text import CountVectorizer
#instantiate the vectorizer
vectorizer = CountVectorizer()
#Create 3 Documents
documnet1 = 'Hi How are you'
documnet2 = 'today is very very very pleasent day and we can have some fun fun fun'
documnet3 = 'This was an amazing experience'
#put all documents together
list_of_documents = [documnet1,documnet2,documnet3]
#fit them as bag of words
bag_of_words = vectorizer.fit(list_of_documents)
#check bag of words
bag_of_words
#Apply transform method
bag_of_words = vectorizer.transform(list_of_documents)
#print bag of words
print(bag_of_words)
```
As we see, the first number is tuple, and the second number is the frequency of words.
The tuple here indicates document number and feature indices of each word which belongs to the document.
Feature indices are generated from the transform method. This is called as the feature extraction process and to extract the features we have to refer to the indexes, which is assigned to any particular feature.
```
#verifying the vocabulary for repeated words
print(vectorizer.vocabulary_.get('very'))
print(vectorizer.vocabulary_.get('fun'))
#check the type of bag of words
type(bag_of_words)
```
In this demo, we learned how to use bag of words technique and transformed documents using transform method.
### Text Feature Extraction Considerations
* __Sparse__:
This utility deals with sparse matrix while storing them in memory. Sparse data is commonly noticed when it comes to extracting feature values, especially for large document datasets.
* __Vectorizer__:
It implements tokenization and occurrence. Words with minimum two letters get tokenized. We can use the analyzer function to vectorize the text data.
* __Tf-idf__:
It is a term weighing utility for term frequency and inverse document frequency. Term frequency indicates the frequency of a particular term in the document. Inverse document frequency is a factor which diminishes the weight of terms that occur frequently.
* __Decoding__:
This utility can decode text files if their encoding is specified.
## Model Training
An important task in model training is to identify the right model for the given dataset. The choice of model completely depends on the type of dataset.
1. Supervised:
* Models predict the outcome of new observations and datasets, and classify documents based on the features and response of a given dataset.
* Example: Naïve Bayes, SVM, linear regression, K-NN neighbors
2. Unsupervised:
* Models identify patterns in the data and extract its structure. They are also used to group documents using clustering algorithms.
* Example: K-means
### Naïve Bayes Classifier
It is the most basic technique for classification of text.
__Advantages:__
* It is efficient as it uses limited CPU and memory.
* It is fast as the model training takes less time.
__Uses:__
* Naïve Bayes is used for sentiment analysis, email spam detection, categorization of documents, and language detection.
* Multinomial Naïve Bayes is used when multiple occurrences of the words matter.
Let us take a look at the signature of the multinomial Naïve Bayes classifier:

### Grid Search and Multiple Parameters
Document classifiers can have many parameters. A Grid approach helps to search the best parameters for model training and predicting the outcome accurately.


In grid search mechanism, the whole dataset can be divided into multiple grids and a search can be run on the entire grid or a combination of grids.

### Pipeline
A pipeline is a combination of vectorizers, transformers, and model training.

### Demo - 04: Pipeline and Grid Search
Demonstrate the Pipeline and Grid Search technique.
```
#import required libraries
import pandas as pd
import string
from pprint import pprint
from time import time
#import dataset
#get spam data collection with the help of the pandas library
df_spam_collection = pd.read_csv('D:\\NIPUN_SC_REC\\3_Practice_Project\\Course_5_Data Science with Python\\Lesson_recap\\SMSSpamCollection.csv', sep = ',', names=['response','message'])
#view first 5 records with head method
df_spam_collection.head()
#import text processing library from scikit-learn
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
#import SGD Classifier
from sklearn.linear_model import SGDClassifier
#import Grid Search
from sklearn.model_selection import GridSearchCV
#import pipeline
from sklearn.pipeline import Pipeline
#define pipeline
pipeline = Pipeline([
('vect',CountVectorizer()),
('tfidf',TfidfTransformer()),
('clf',SGDClassifier())
])
#parameter for grid search
parameters = {'tfidf__use_idf':(True,False)}
#perform the gridsearch with pipeline and parameters
grid_search = GridSearchCV(pipeline,parameters,n_jobs = -1,verbose = 1)
print('Performing Grid Search now..')
print('Parameters...')
pprint(parameters)
to = time()
grid_search.fit(df_spam_collection['message'],
df_spam_collection['response'])
print('done in %0.3fs' %(time() - to))
print()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
import plotly.offline as py
import plotly.graph_objs as go
import plotly.tools as tls
import seaborn as sns
import io
from google.colab import files
uploaded = files.upload()
data = uploaded['LAND TEMPERATURES FILE FROM KAGGLE IMPORT']
inputData = pd.read_csv(io.BytesIO(data))
#Process and Clean Data - from Kaggle DataSet
countriesToClear = ['CHOOSE COUNTRIES']
countriesToBeReplaced = ['CHOOSE COUNTRIES']
countriedReplacedWith = ['CHOOSE REPLACEMENTS']
cleanedCountries = inputData[~inputData['Country'].isin(countriesToClear)]
cleanedCountries = cleanedCountries.replace(countriesToBeReplaced ,countriedReplacedWith)
#Average Country Temperatures
countries = np.unique(cleanedCountries['Country'])
meanTemperatures = []
for country in countries:
meanTempsByCountry = cleanedCountries[cleanedCountries['Country'] == country]['AverageTemperature'].mean()
meanTemperatures.append(meanTempsByCountry)
meanTempBar, countryBar = (list(x) for x in zip(*sorted(zip(meanTemperatures, countries), reverse = True)))
#Plot Graphs of Hottest and Coldest
sns.set(font_scale=1)
a,b = plt.subplots(figsize=(5, 10))
sns.barplot(meanTempBar[:25], countryBar[:25], palette = sns.color_palette('coolwarm', len(countries))[::1])
b.set(xlabel='Average temperature', title='Warmest 25 Countries')
sns.set(font_scale=1)
a,b = plt.subplots(figsize=(5, 10))
sns.barplot(meanTempBar[-25:], countryBar[-25:], palette = sns.color_palette('coolwarm', len(countries))[::1])
b.set(xlabel='Average temperature', title='Coldest 25 Countries')
# Combine Data Into Zipped List
countriesToPlot = countryBar
yAxis = np.arange(len(countriesToPlot))
tempsToPlot = meanTempBar
countries_zipped = list(zip(countryBar,meanTempBar))
# Plot All Countries Mean Temperatures
plt.bar(yAxis,tempsToPlot, align='center', alpha=0.5)
plt.ylabel('Temperature / degrees celcius')
plt.xlabel('Countries / country number')
plt.title('Average Global Temperatures 1750-2013')
plt.show()
# Cycle Through Each Country
counter = 0 #
for country in countries:
if counter < 10:
temperature = meanTempsByCountry = cleanedCountries[cleanedCountries['Country'] == country]['AverageTemperature'].tail(165).iloc[::12]
date = meanTempsByCountry = cleanedCountries[cleanedCountries['Country'] == country]['dt'].tail(165).iloc[::12]
plot(date,temperature)
# Plot Temperatures of Each Country Over A Year
xlabel('Time / Date ')
ylabel('Temperature / Degrees Celcius')
title(country)
plt.xticks(rotation=90)
grid(True)
show()
```
| github_jupyter |
```
import bare
```
#### Plot detected interest points over images
```
image_file_name = 'image.tif'
ip_file_name = 'image.vwip'
ip_csv_file_name = bare.core.write_ip_to_csv(ip_file_name)
bare.plot.ip_plot(image_file_name, ip_csv_file_name)
```

#### Plot match points found between two images
```
image1_file_name = 'image1.tif'
image2_file_name = 'image2.tif'
match_file_name = 'image1__image2-clean.match'
match_csv_file_name = bare.core.write_mp_to_csv(match_file_name)
bare.plot.mp_plot(image1_file_name, image2_file_name, match_csv_file_name,out_dir_abs='qc_plots/match_points')
```

#### Plot dxdy after bundle adjustment
```
ba_dir = 'bundle_adjust_output_directory/'
bare.plot.plot_dxdy(ba_dir)
```

#### Plot tsai camera xyz positions bef|ore and after bundle adjustment
```
ba_dir = 'bundle_adjust_output_directory/'
img_dir = 'input_image_directory/'
input_cam_dir = 'input_cameras_directory/'
bare.plot.plot_tsai_camera_positions_before_and_after(ba_dir, input_cam_dir)
```


#### Plot residuals after bundle adjustment
Figure on the right shows outliers is a result of outliers.
```
ba_dir = 'bundle_adjust_output_directory/'
bare.plot.plot_residuals(ba_dir)
```

#### Plot WV3 footprint and scanner positions during aquisition.
```
image_file_name = 'wv_image.tif'
camera_file = 'wv_image.xml'
reference_dem = 'reference_dem.tif'
bare.plot.plot_footprint(image_file_name,
camera_file,
reference_dem)
```

#### Plot WV3 footprint.
```
image_file_name = 'wv_image.tif'
camera_file = 'wv_image.xml'
reference_dem = 'reference_dem.tif'
bare.plot.plot_footprint(image_file_name,
camera_file,
reference_dem,
cam_on=False)
```

#### Plot tsai camera footprint and position.
```
image_file_name = 'image.tif'
camera_file = 'image.tsai'
reference_dem = 'reference_dem.tif'
bare.plot.plot_footprint(image_file_name,
camera_file,
reference_dem)
```

#### Batch plot tsai camera footprints.
```
img_dir = 'input_image_directory/'
cam_dir = 'input_tsai_camera_directory/'
reference_dem = 'reference_dem.tif'
bare.batch.plot_footprints(cam_dir, img_dir, reference_dem)
```

#### Run all
```
ba_dir = 'bundle_adjust_output_directory/'
img_dir = 'input_image_directory/'
input_cam_dir = 'input_cameras_directory/'
bare.batch.plot_all_qc_products(ba_dir,
img_dir,
input_cam_dir,
img_extension='8.tif')
```
| github_jupyter |
<a id="introduction"></a>
## Introduction to Clustering
#### By Paul Hendricks
-------
Clustering is an important technique for helping data scientists partition data, especially when that data doesn't have labels or annotations associated with it. Since these data often don't have labels, clustering is often described as an unsupervised learning technique. While there are many different algorithms that partition data into unique clusters, we will show in this notebook how in certain cases the DBSCAN algorithm can do a better job of clustering than traditional algorithms such as K-Means or Agglomerative Clustering.
We will also show how to cluster data with K-Means and DBSCAN in NVIDIA RAPIDS – an open-source data analytics and machine learning acceleration platform that leverages GPUs to accelerate computations. We will see that porting this example from CPU to GPU is trivial and that we can experience massive performance gains by doing so.
In this notebook, we will describe K-Means and DBSCAN - several popular clustering algorithms - and show how to use the GPU accelerated implementation of these algorithms in RAPIDS.
**Table of Contents**
* [Introduction to Clustering](#introduction)
* [Setup](#setup)
* [Generating Data](#generate)
* [Overview of K-Means and Agglomerative Clustering](#overview)
* [Clustering using DBSCAN](#dbscan)
* [Accelerating Clustering with RAPIDS](#accelerating)
* [Clustering using GPU Accelerated K-Means](#gpukmeans)
* [Benchmarking: Comparing GPU and CPU](#benchmarking)
* [K-Means](#benchmarkingkmeans)
* [GPU](#benchmarkingkmeansgpu)
* [CPU](#benchmarkingkmeanscpu)
* [DBSCAN](#benchmarkingdbscan)
* [GPU](#benchmarkingdbscangpu)
* [CPU](#benchmarkingdbscancpu)
* [Conclusion](#conclusion)
Before going any further, let's make sure we have access to `matplotlib`, a popular Python library for visualizing data.
```
import os
try:
import matplotlib; print('Matplotlib Version:', matplotlib.__version__)
except ModuleNotFoundError:
os.system('conda install -y matplotlib')
```
<a id="setup"></a>
## Setup
This notebook was tested using the following Docker containers:
* `rapidsai/rapidsai-nightly:0.8-cuda10.0-devel-ubuntu18.04-gcc7-py3.7` from [DockerHub - rapidsai/rapidsai-nightly](https://hub.docker.com/r/rapidsai/rapidsai-nightly)
This notebook was run on the NVIDIA Tesla V100 GPU. Please be aware that your system may be different and you may need to modify the code or install packages to run the below examples.
If you think you have found a bug or an error, please file an issue here: https://github.com/rapidsai/notebooks/issues
Before we begin, let's check out our hardware setup by running the `nvidia-smi` command.
```
!nvidia-smi
```
Next, let's see what CUDA version we have:
```
!nvcc --version
```
Next, let's load some helper functions from `matplotlib` and configure the Jupyter Notebook for visualization.
```
from matplotlib.colors import ListedColormap
import matplotlib.pyplot as plt
%matplotlib inline
```
<a id="generate"></a>
## Generating Data
We'll generate some fake data using the `make_moons` function from the `sklearn.datasets` module. This function generates data points from two equations, each describing a half circle with a unique center. Since each data point is generated by one of these two equations, the cluster each data point belongs to is clear. The ideal clustering algorithm will identify two clusters and associate each data point with the equation that generated it.
```
import sklearn; print('Scikit-Learn Version:', sklearn.__version__)
from sklearn.datasets import make_moons
X, y = make_moons(n_samples=int(1e2), noise=0.05, random_state=0)
print(X.shape)
```
Let's visualize our data:
```
# create figure
figure = plt.figure()
axis = figure.add_subplot(111)
axis.scatter(X[y == 0, 0], X[y == 0, 1],
edgecolor='black',
c='lightblue', marker='o', s=40, label='cluster 1')
axis.scatter(X[y == 1, 0], X[y == 1, 1],
edgecolor='black',
c='red', marker='s', s=40, label='cluster 2')
axis.set_title('Empirical Data Points with Labels')
plt.legend()
plt.tight_layout()
plt.show()
```
<a id="overview"></a>
## Overview of K-Means and Agglomerative Clustering
There exist several algorithms for partitioning data into partitions, two of the more common of which are called K-Means and Agglomerative Clustering.
The K-Means algorithm approaches the clustering problem by partitioning a set of data points into disjoint clusters, where each cluster is described by the mean of the samples in the cluster. The mean of the samples in a particular cluster is called a centroid; the K-Means algorithm finds the centroids and associates data points with centroids in such a way as to minimize the within-cluster sum-of-squares.
For more information on the K-Means algorithm and its implementatin in scikit-learn, check out this resource: http://scikit-learn.org/stable/modules/clustering.html#k-means
In the code cell below, we instantiate the `KMeans` algorithm from the `sklearn.cluster` module and apply it to our data using the `fit_predict` method. We see that `KMeans` identifies two centroids; one located at about (-0.23, 0.56) and the other located at (1.17, -0.05).
```
from sklearn.cluster import KMeans
km = KMeans(n_clusters=2, random_state=0)
y_km = km.fit_predict(X)
print(km.cluster_centers_)
```
The Agglomerative Clustering algorithm behaves a little bit differently and does not identify clusters using centroids. Instead, it recursively merges the pair of clusters that minimally increases a given linkage distance. Put another way, the Agglomerative Clustering algorithm identifies the two data points that are "closest" out of all the data samples. It then takes those two data points and identifies a third data point that is "closest" to those two data points. The algorithm continues in this fashion for each data point; finding the next data point that is "closest" to the preceeding cluster of data points, where the definition of "closest" depends on the distance metric chosen.
For more information on the Agglomerative Clustering algorithm and its implementatin in scikit-learn, check out this resource: http://scikit-learn.org/stable/modules/clustering.html#hierarchical-clustering
Below, we instantiate the `AgglomerativeClustering` algorithm from the `sklearn.cluster` module and apply it to our data.
```
from sklearn.cluster import AgglomerativeClustering
ac = AgglomerativeClustering(n_clusters=2,
affinity='euclidean',
linkage='complete')
y_ac = ac.fit_predict(X)
```
We can visualize the results of both algorithms applied to the data. Visually, we see that neither algorithm ideally clusters our data. The ideal algorithm for this unique set of data would recognize that both sets of samples are generated from two different equations describing two different half circles.
```
# Create two subplots
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 3))
ax1.scatter(X[y_km == 0, 0], X[y_km == 0, 1],
edgecolor='black',
c='lightblue', marker='o', s=40, label='cluster 1')
ax1.scatter(X[y_km == 1, 0], X[y_km == 1, 1],
edgecolor='black',
c='red', marker='s', s=40, label='cluster 2')
ax1.set_title('K Means Clustering')
ax2.scatter(X[y_ac == 0, 0], X[y_ac == 0, 1], c='lightblue',
edgecolor='black',
marker='o', s=40, label='cluster 1')
ax2.scatter(X[y_ac == 1, 0], X[y_ac == 1, 1], c='red',
edgecolor='black',
marker='s', s=40, label='cluster 2')
ax2.set_title('Agglomerative Clustering')
plt.legend()
plt.tight_layout()
plt.show()
```
<a id="dbscan"></a>
## Clustering using DBSCAN
Unlike K Means or Agglomerative Clustering, DBSCAN is a density-based approach to spatial clustering. It views clusters as areas of high density separated by areas of low density. This approach has several advantages; whereas K Means focuses on finding centroids and assoicating data points with that centroid in a spherical manner, the DBSCAN algorithm can identify clusters of any convex shape. Additionally, DBSCAN is robust to areas of low density. In the above visualization, we see that Agglomerative Clustering ignores the low density space space between the interleaving circles and instead focuses on finding a clustering hierarchy that minimizes the Euclidean distance. While minimizing Euclidean distance is important for some clustering problems, it is visually apparent to a human that following the density trail of points results in the ideal clustering.
For more information on the DBSCAN algorithm and its implementation in scikit-learn, check out this resource: http://scikit-learn.org/stable/modules/clustering.html#dbscan
```
from sklearn.cluster import DBSCAN
db = DBSCAN(eps=0.2, min_samples=2)
```
Next, let's fit our model to the data and generate predictions.
```
y_db = db.fit_predict(X)
```
Lastly, let's visualize the model applied to our data. We see that the DBSCAN algorithm correctly identifies which half-circle each data point is generated from.
```
plt.scatter(X[y_db == 0, 0], X[y_db == 0, 1],
c='lightblue', marker='o', s=40,
edgecolor='black',
label='cluster 1')
plt.scatter(X[y_db == 1, 0], X[y_db == 1, 1],
c='red', marker='s', s=40,
edgecolor='black',
label='cluster 2')
plt.legend()
plt.tight_layout()
plt.show()
```
<a id="accelerating"></a>
## Accelerating Clustering with RAPIDS
So how can we accelerate our code using RAPIDS? First, we cast our data to a `pandas.DataFrame` and use that to create a `cudf.DataFrame`.
```
import cudf; print('cuDF Version:', cudf.__version__)
import pandas as pd; print('Pandas Version:', pd.__version__)
# iteratively create a column for each column of X
X_df = pd.DataFrame({'fea%d'%i: X[:, i] for i in range(X.shape[1])}) # shape = (n_row, n_col)
X_gdf = cudf.DataFrame.from_pandas(X_df)
```
<a id="gpukmeans"></a>
## Clustering using GPU Accelerated K-Means
Next, we load the `KMeans` class from the `cuml` package and instantiate it in the same way we did with the `sklearn.cluster.KMeans` class.
```
from cuml import KMeans as KMeans_GPU
km_gpu = KMeans_GPU(n_clusters=2)
```
The `KMeans` class from `cuml` implements the same API as the `sklearn` version; we can use the `fit` and `fit_predict` methods to fit our `KMeans` model to the data and generate predictions.
```
y_km_gpu = km_gpu.fit_predict(X_gdf)
```
Lastly, let's visualize our results:
```
# convert output to array
plt.scatter(X[y_km_gpu.to_array() == 0, 0], X[y_km_gpu.to_array() == 0, 1],
c='lightblue', marker='o', s=40,
edgecolor='black',
label='cluster 1')
plt.scatter(X[y_km_gpu.to_array() == 1, 0], X[y_km_gpu.to_array() == 1, 1],
c='red', marker='s', s=40,
edgecolor='black',
label='cluster 2')
plt.legend()
plt.tight_layout()
plt.show()
```
<a id="gpudbscan"></a>
## Clustering using GPU Accelerated DBSCAN
Next, we load the `DBSCAN` class from the `cuml` package and instantiate it in the same way we did with the `sklearn.cluster.DBSCAN` class.
```
import cuml; print('cuML Version:', cuml.__version__)
from cuml import DBSCAN as DBSCAN_GPU
db_gpu = DBSCAN_GPU(eps=0.2, min_samples=2)
```
The `DBSCAN` class from `cuml` implements the same API as the `sklearn` version; we can use the `fit` and `fit_predict` methods to fit our `DBSCAN` model to the data and generate predictions.
```
y_db_gpu = db_gpu.fit_predict(X_gdf)
```
Lastly, let's visualize our results:
```
plt.scatter(X[y_db_gpu.to_array() == 0, 0], X[y_db_gpu.to_array() == 0, 1],
c='lightblue', marker='o', s=40,
edgecolor='black',
label='cluster 1')
plt.scatter(X[y_db_gpu.to_array() == 1, 0], X[y_db_gpu.to_array() == 1, 1],
c='red', marker='s', s=40,
edgecolor='black',
label='cluster 2')
plt.legend()
plt.tight_layout()
plt.show()
```
<a id="benchmarking"></a>
## Benchmarking: Comparing GPU and CPU
RAPIDS uses GPUs to parallelize operations and accelerate computations. We saw porting an example from the traditional scikit-learn interface to cuML was trivial. So how much speedup do we get from using RAPIDS?
The answer to this question varies depending on the size and shape of the data. As a good rule of thumb, larger datasets will benefit from RAPIDS. There is overhead associated with using a GPU; data has to be transferred from the CPU to the GPU, computations have to take place on the GPU, and the results need to be transferred back from the GPU to the CPU. However, the transactional overhead of moving data back and forth from the CPU to the GPU can quickly become negligible due to the performance speedup from computing on a GPU instead of a CPU.
Feel free to change the number of rows and columns to see how this speedup might change depending on the size and shape of the data.
```
import numpy as np; print('NumPy Version:', np.__version__)
n_rows, n_cols = 100000, 128
X = np.random.rand(n_rows, n_cols)
print(X.shape)
X_df = pd.DataFrame({'fea%d'%i: X[:, i] for i in range(X.shape[1])})
X_gdf = cudf.DataFrame.from_pandas(X_df)
```
<a id="benchmarkingkmeans"></a>
## K-Means
<a id="benchmarkingkmeansgpu"></a>
### GPU
```
km_gpu = KMeans_GPU(n_clusters=2)
%%time
y_km_gpu = km_gpu.fit_predict(X_gdf)
```
<a id="benchmarkingkmeanscpu"></a>
### CPU
```
km = KMeans(n_clusters=2)
%%time
y_km = km.fit_predict(X)
```
<a id="benchmarkingdbscan"></a>
## DBSCAN
<a id="benchmarkingdbscangpu"></a>
### GPU
```
db_gpu = DBSCAN_GPU(eps=3, min_samples=2)
%%time
y_db_gpu = db_gpu.fit_predict(X_gdf)
```
<a id="benchmarkingdbscancpu"></a>
### CPU
```
db = DBSCAN(eps=3, min_samples=2)
%%time
print('This takes a very long time!')
y_db = db.fit_predict(X) # uncomment this line to execute - run at your own risk of boredom!
```
<a id="conclusion"></a>
## Conclusion
In this notebook, we described K-Means and DBSCAN - several popular clustering algorithms - and showed how to use the GPU accelerated implementation of these algorithms in RAPIDS.
To learn more about RAPIDS, be sure to check out:
* [Open Source Website](http://rapids.ai)
* [GitHub](https://github.com/rapidsai/)
* [Press Release](https://nvidianews.nvidia.com/news/nvidia-introduces-rapids-open-source-gpu-acceleration-platform-for-large-scale-data-analytics-and-machine-learning)
* [NVIDIA Blog](https://blogs.nvidia.com/blog/2018/10/10/rapids-data-science-open-source-community/)
* [Developer Blog](https://devblogs.nvidia.com/gpu-accelerated-analytics-rapids/)
* [NVIDIA Data Science Webpage](https://www.nvidia.com/en-us/deep-learning-ai/solutions/data-science/)
| github_jupyter |
```
%matplotlib inline
import numpy as np
import pandas as pd
import scipy
import sklearn
import spacy
import matplotlib.pyplot as plt
import seaborn as sns
import re
from nltk.corpus import state_union, stopwords
from collections import Counter
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings("ignore")
import nltk
nltk.download()
state_union.fileids()
#Let's analyze Eisenhower and Kennedy
eisenhower = state_union.raw('1953-Eisenhower.txt')
kennedy = state_union.raw('1962-Kennedy.txt')
# Utility function for standard text cleaning.
def text_cleaner(text):
# Visual inspection identifies a form of punctuation spaCy does not
# recognize: the double dash '--'. Better get rid of it now!
text = re.sub(r'--',' ',text)
text = re.sub("[\[].*?[\]]", "", text)
text = ' '.join(text.split())
return text
eisenhower = text_cleaner(eisenhower)
kennedy = text_cleaner(kennedy)
#SpaCy
nlp = spacy.load('en')
eisenhower_doc = nlp(eisenhower)
kennedy_doc = nlp(kennedy)
# Group into sentences.
eisenhower_sents = [[sent, 'Eisenhower'] for sent in eisenhower_doc.sents]
kennedy_sents = [[sent, "Kennedy"] for sent in kennedy_doc.sents]
# Combine the sentences from the two novels into one data frame.
sentences = pd.DataFrame(eisenhower_sents + kennedy_sents)
sentences.head()
# how long are their speeches?
print('Eisenhower speech length:', len(eisenhower_doc))
print('Kennedy speech length:', len(kennedy_doc))
# check excerpts for any other cleaning needed
print(eisenhower_doc[:100])
print(kennedy_doc[:100])
```
## Bag of Words
```
# Utility function to create a list of the 2000 most common words.
def bag_of_words(text):
# Filter out punctuation and stop words.
allwords = [token.lemma_
for token in text
if not token.is_punct
and not token.is_stop]
# Return the most common words.
return [item[0] for item in Counter(allwords).most_common(2000)]
# Creates a data frame with features for each word in our common word set.
# Each value is the count of the times the word appears in each sentence.
def bow_features(sentences, common_words):
# Scaffold the data frame and initialize counts to zero.
df = pd.DataFrame(columns=common_words)
df['text_sentence'] = sentences[0]
df['text_source'] = sentences[1]
df.loc[:, common_words] = 0
# Process each row, counting the occurrence of words in each sentence.
for i, sentence in enumerate(df['text_sentence']):
# Convert the sentence to lemmas, then filter out punctuation,
# stop words, and uncommon words.
words = [token.lemma_
for token in sentence
if (
not token.is_punct
and not token.is_stop
and token.lemma_ in common_words
)]
# Populate the row with word counts.
for word in words:
df.loc[i, word] += 1
# This counter is just to make sure the kernel didn't hang.
if i % 500 == 0:
print("Processing row {}".format(i))
return df
# Set up the bags.
eisenhowerwords = bag_of_words(eisenhower_doc)
kennedywords = bag_of_words(kennedy_doc)
# Combine bags to create a set of unique words.
common_words = set(eisenhowerwords + kennedywords)
# Create bow features
bow = bow_features(sentences, common_words)
bow.head()
```
## TF-IDF
```
#sentences
eisenhower = state_union.sents('1953-Eisenhower.txt')
kennedy = state_union.sents('1962-Kennedy.txt')
# lists
eisenhower_list = [" ".join(sent) for sent in eisenhower]
kennedy_list = [" ".join(sent) for sent in kennedy]
together = eisenhower_list + kennedy_list
from sklearn.feature_extraction.text import TfidfVectorizer
#X_train, X_test = train_test_split(together, test_size=0.4, random_state=0)
vectorizer = TfidfVectorizer(max_df=0.5, # drop words that occur in more than half the paragraphs
min_df=2, # only use words that appear at least twice
stop_words='english',
lowercase=True, #convert everything to lower case (since Alice in Wonderland has the HABIT of CAPITALIZING WORDS for EMPHASIS)
use_idf=True,#we definitely want to use inverse document frequencies in our weighting
norm=u'l2', #Applies a correction factor so that longer paragraphs and shorter paragraphs get treated equally
smooth_idf=True #Adds 1 to all document frequencies, as if an extra document existed that used every word once. Prevents divide-by-zero errors
)
#Applying the vectorizer
together_tfidf=vectorizer.fit_transform(together)
print("Number of features: %d" % together_tfidf.get_shape()[1])
tfidf = vectorizer.fit_transform(together).tocsr()
```
### These two texts, even though just a few years apart, are not highly correlated. There could be many reasons for this, but perhaps it's a shift in party in the White House? Or, different events at the time.
## Supervised Learning Models
#### Logistic Regression
```
#Imports
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score, cross_val_predict
from sklearn import metrics
# Set X, y and train, test, split
y = bow['text_source']
X = np.array(bow.drop(['text_sentence','text_source'], 1))
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size=0.4,
random_state=0)
# Logistic Regression Model with BoW
lrb = LogisticRegression()
model = lrb.fit(X_train, y_train)
pred = lrb.predict(X_test)
print(X_train.shape, y_train.shape)
print('BoW Training set score:', lrb.score(X_train, y_train))
print('BoW Test set score:', lrb.score(X_test, y_test))
print('BoW Predictions:', pred[0:5])
#5 fold Cross Validation
scores = cross_val_score(model, X, y, cv=5)
print('Cross-validated scores:', scores)
print('Avg. Score ', np.mean(cross_val_score(lrb, X, y, cv=5)))
# Tfidf
X_tfidf = tfidf
y_tfidf = ['Eisenhower']*len(eisenhower_list) + ['Kennedy']*len(kennedy_list)
X2_train, X2_test, y2_train, y2_test = train_test_split(X_tfidf,
y_tfidf,
test_size=0.4,
random_state=0)
# Logistic Regression Model with TFIDF
lrt = LogisticRegression()
model = lrt.fit(X2_train, y2_train)
pred = lrt.predict(X2_test)
print('\nTFIDF Training set score:', lrt.score(X2_train, y2_train))
print('TFIDF Test set score:', lrt.score(X2_test, y2_test))
print('Predictions:', pred[0:5])
#5 fold Cross Validation
scores = cross_val_score(model, X_tfidf, y_tfidf, cv=5)
print('Cross-validated scores:', scores)
print('Avg. Score ', np.mean(cross_val_score(lrt, X_tfidf, y_tfidf, cv=5)))
```
#### Random Forest
```
#Import
from sklearn.ensemble import RandomForestClassifier
#Random Forest Model with BoW
rfcb = RandomForestClassifier()
model = rfcb.fit(X_train, y_train)
pred = rfcb.predict(X_test)
print('Training set score:', rfcb.score(X_train, y_train))
print('Test set score:', rfcb.score(X_test, y_test))
print('Predictions:', pred[0:5])
#5 fold cross validation
scores = cross_val_score(model, X, y, cv=5)
print('Cross-validated scores:', scores)
print('Avg. Score ', np.mean(cross_val_score(rfcb, X, y, cv=5)))
# Random Forest Model with TFIDF
rfct = RandomForestClassifier()
model = rfct.fit(X2_train, y2_train)
pred = rfct.predict(X2_test)
print('\nTFIDF Training set score:', rfct.score(X2_train, y2_train))
print('TFIDF Test set score:', rfct.score(X2_test, y2_test))
print('Predictions:', pred[0:5])
#5 fold Cross Validation
scores = cross_val_score(model, X_tfidf, y_tfidf, cv=5)
print('Cross-validated scores:', scores)
print('Avg. Score ', np.mean(cross_val_score(rfct, X_tfidf, y_tfidf, cv=5)))
```
#### XGBoost Classifier
```
#import
from xgboost import XGBClassifier
#Our XGBoost Classifier
clfb = XGBClassifier()
model= clfb.fit(X_train, y_train)
print('Training set score:', clfb.score(X_train, y_train))
print('Test set score:', clfb.score(X_test, y_test))
#5 fold cross validation
scores = cross_val_score(model, X, y, cv=5)
print('Cross-validated scores:', scores)
print('Avg. Score ', np.mean(cross_val_score(clfb, X, y, cv=5)))
# Random Forest Model with TFIDF
clft = XGBClassifier()
model = clft.fit(X2_train, y2_train)
pred = clft.predict(X2_test)
print('\nTFIDF Training set score:', clft.score(X2_train, y2_train))
print('TFIDF Test set score:', clft.score(X2_test, y2_test))
print('Predictions:', pred[0:5])
#5 fold Cross Validation
scores = cross_val_score(model, X_tfidf, y_tfidf, cv=5)
print('Cross-validated scores:', scores)
print('Avg. Score ', np.mean(cross_val_score(clft, X_tfidf, y_tfidf, cv=5)))
# We'll make 500 iterations, use 2-deep trees, and set our loss function.
params = {'n_estimators': 500,
'max_depth': 2,
'loss': 'deviance'}
# Initialize and fit the model.
clfb = ensemble.GradientBoostingClassifier(**params)
model= clfb.fit(X_train, y_train)
print('Training set score:', clfb.score(X_train, y_train))
print('Test set score:', clfb.score(X_test, y_test))
#5 fold cross validation
scores = cross_val_score(model, X, y, cv=5)
print('Cross-validated scores:', scores)
print('Avg. Score ', np.mean(cross_val_score(clfb, X, y, cv=5)))
# Random Forest Model with TFIDF
clft = ensemble.GradientBoostingClassifier(**params)
model = clft.fit(X2_train, y2_train)
pred = clft.predict(X2_test)
print('\nTFIDF Training set score:', clft.score(X2_train, y2_train))
print('TFIDF Test set score:', clft.score(X2_test, y2_test))
print('Predictions:', pred[0:5])
#5 fold Cross Validation
scores = cross_val_score(model, X_tfidf, y_tfidf, cv=5)
print('Cross-validated scores:', scores)
print('Avg. Score ', np.mean(scores))
```
### Increase Accuracy by 5% on Random Forest
```
# Utility function to create a list of the 3000 most common words and add in punctuation.
def bag_of_words(text):
# Filter out punctuation and stop words.
allwords = [token.lemma_
for token in text
if not token.is_stop]
# Return the most common words.
return [item[0] for item in Counter(allwords).most_common(4000)]
# Creates a data frame with features for each word in our common word set.
# Each value is the count of the times the word appears in each sentence.
def bow_features(sentences, common_words):
# Scaffold the data frame and initialize counts to zero.
df = pd.DataFrame(columns=common_words)
df['text_sentence'] = sentences[0]
df['text_source'] = sentences[1]
df.loc[:, common_words] = 0
# Process each row, counting the occurrence of words in each sentence.
for i, sentence in enumerate(df['text_sentence']):
# Convert the sentence to lemmas, then filter out punctuation,
# stop words, and uncommon words.
words = [token.lemma_
for token in sentence
if (
not token.is_punct
and not token.is_stop
and token.lemma_ in common_words
)]
# Populate the row with word counts.
for word in words:
df.loc[i, word] += 1
# This counter is just to make sure the kernel didn't hang.
if i % 500 == 0:
print("Processing row {}".format(i))
return df
# Set up the bags.
eisenhowerwords = bag_of_words(eisenhower_doc)
kennedywords = bag_of_words(kennedy_doc)
# Combine bags to create a set of unique words.
common_words = set(eisenhowerwords + kennedywords)
# Create bow features
bow_inc = bow_features(sentences, common_words)
bow.head()
from sklearn.model_selection import GridSearchCV
# Set X, y and train, test, split
y2 = bow_inc['text_source']
X2 = np.array(bow_inc.drop(['text_sentence','text_source'], 1))
X2_train, X2_test, y2_train, y2_test = train_test_split(X2,
y2,
test_size=0.4,
random_state=0)
# Logistic Regression Model with GridSearchCV on BoW
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10, 100, 1000] }
lrb2 = GridSearchCV(LogisticRegression(penalty='l2',
random_state=42,
dual=True,
class_weight=None), param_grid)
model2 = lrb2.fit(X2_train, y2_train)
pred = lrb2.predict(X2_test)
print(X2_train.shape, y2_train.shape)
print('BoW Training set score:', lrb2.score(X2_train, y2_train))
print('BoW Test set score:', lrb2.score(X2_test, y2_test))
print('BoW Predictions:', pred[0:5])
#10 fold Cross Validation
scores = cross_val_score(model2, X2, y2, cv=10)
print('Cross-validated scores:', scores)
print('Avg. Score ', np.mean(scores))
```
### Adding Grid Search, tuning parameters and Cross Validating with 10 folds was the best solution to increase by 5%.
| github_jupyter |
```
import pandas as pd
import numpy as np
from pathlib import Path
import os
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import GridSearchCV
from sklearn.decomposition import PCA
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import confusion_matrix
from sklearn.metrics import plot_confusion_matrix
from sklearn.metrics import plot_roc_curve
from sklearn.metrics import classification_report
from sklearn.pipeline import Pipeline
from scipy import stats
from joblib import dump
from joblib import load
import xgboost as xgb
import matplotlib.pyplot as plt
from typing import Dict
from kaggle.api.kaggle_api_extended import KaggleApi
from dotenv import find_dotenv, load_dotenv
from collections import Counter
from src.data import make_dataset
from src.visualization.visualize import classification_reports, classification_reports_2
from src.visualization.visualize import plot_pie, create_feature_importance_df, plot_feature_importances
load_dotenv(find_dotenv())
api = KaggleApi()
api.authenticate()
competition = os.environ['COMPETITION']
```
# Set up directories
```
project_dir = Path.cwd().parent
data_dir = project_dir / 'data'
raw_data_dir = data_dir / 'raw'
interim_data_dir = data_dir / 'interim'
processed_data_dir = data_dir / 'processed'
models_dir = project_dir / 'models'
```
# Load data
```
df_train = pd.read_csv(raw_data_dir / 'train.csv')
df_test = pd.read_csv(raw_data_dir / 'test.csv')
X_train = np.load(interim_data_dir / 'X_train.npy')
X_val = np.load(interim_data_dir / 'X_val.npy')
y_train = np.load(interim_data_dir / 'y_train.npy')
y_val = np.load(interim_data_dir / 'y_val.npy')
X_test = np.load(interim_data_dir / 'X_test.npy')
test_id = pd.read_csv(interim_data_dir / 'test_id.csv')
```
# Baseline
The base line prediction is simply to make them all negative.
```
labels = 'Positive', 'Negative'
pos_count = (y_train == 1).sum()
neg_count = (y_train == 0).sum()
sizes = [pos_count, neg_count]
explode = (0, 0.1) # only "explode" the 2nd slice (i.e. 'Hogs')
fig1, ax1 = plt.subplots()
ax1.pie(sizes, explode=explode, labels=labels, autopct='%1.1f%%',
shadow=True, startangle=90)
ax1.axis('equal') # Equal aspect ratio ensures that pie is drawn as a circle.
plt.show()
pd.Series(y_train).value_counts(normalize=True)
labels = 'Positive', 'Negative'
pos_count = (y_val == 1).sum()
neg_count = (y_val == 0).sum()
sizes = [pos_count, neg_count]
explode = (0, 0.1) # only "explode" the 2nd slice (i.e. 'Hogs')
fig1, ax1 = plt.subplots()
ax1.pie(sizes, explode=explode, labels=labels, autopct='%1.1f%%',
shadow=True, startangle=90)
ax1.axis('equal') # Equal aspect ratio ensures that pie is drawn as a circle.
plt.show()
```
## Baseline Score
The baseline score is to merely predict the larger class for all instances of the training set.
```
preds = [0] * len(y_val)
roc_auc_score(y_val, preds)
```
# Naive XGB
```
clf = xgb.XGBClassifier()
clf.fit(X_train, y_train)
preds = clf.predict(X_val)
probs = clf.predict_proba(X_val)
auc = roc_auc_score(y_val, probs[:, 1])
auc
plot_roc_curve(clf, X_val, y_val)
plot_confusion_matrix(clf, X_val, y_val)
print(classification_report(y_val, preds))
```
# RandomizedSearchCV XGB
```
pipe = Pipeline([
('scaler', StandardScaler()),
('pca', PCA()),
('classifier', xgb.XGBClassifier(
objective='binary:logistic',
use_label_encoder=False,
eval_metric='logloss'
))
])
param_dist = {
'pca__n_components': stats.randint(1, X_train.shape[1]),
'classifier__n_estimators': stats.randint(150, 1000),
'classifier__learning_rate': stats.uniform(0.01, 0.6),
'classifier__subsample': stats.uniform(0.3, 0.9),
'classifier__max_depth': [3, 4, 5, 6, 7, 8, 9],
'classifier__colsample_bytree': stats.uniform(0.5, 0.9),
'classifier__min_child_weight': [1, 2, 3, 4]
}
cv = RandomizedSearchCV(
estimator=pipe,
param_distributions=param_dist,
random_state=42,
n_iter=100,
cv=5,
n_jobs=7,
verbose=10,
scoring='roc_auc'
)
cv.fit(X_train, y_train)
dump(cv, models_dir / 'randomised_xgb.joblib')
```
## Training set results
```
preds = cv.predict(X_train)
probs = cv.predict_proba(X_train)[:, 1]
classification_reports_2(y_train, preds, probs)
pd.DataFrame(cv.cv_results_).query('mean_test_score.notnull()').sort_values(by=['mean_test_score'], ascending=False)
pd.DataFrame(cv.cv_results_).query('mean_test_score.isnull()').sort_values(by=['mean_test_score'], ascending=False)
pd.DataFrame(cv.cv_results_).query('mean_test_score.notnull()').sort_values(by=['mean_test_score'], ascending=False).iloc[0]
```
## Predictions with the best model
```
preds = cv.predict(X_val)
probs = cv.predict_proba(X_val)
roc_auc_score(y_val, probs[:, 1])
plot_roc_curve(cv, X_val, y_val)
plot_confusion_matrix(cv, X_val, y_val)
print(classification_report(y_val, preds))
```
The model is misclassifying the negative classes.
```
# fpr
260/(260+3)
```
## Predict on test set
```
preds = cv.predict(X_test)
probs = cv.predict_proba(X_test)[:, 1]
```
## Save predictions
```
pred_name = 'TARGET_5Yrs'
pred_path = processed_data_dir / 'preds_randomised_xgb.csv'
make_dataset.save_predictions(probs, pred_name, test_id, pred_path)
pred_path.stem
```
## Submit predictions
| github_jupyter |
<a href="https://colab.research.google.com/github/GivanTsai/Bert-cookbook/blob/main/BERT_Fine_Tuning_Sentence_Classification_v4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# BERT Fine-Tuning Tutorial with PyTorch
By Chris McCormick and Nick Ryan
*Revised on March 20, 2020 - Switched to `tokenizer.encode_plus` and added validation loss. See [Revision History](https://colab.research.google.com/drive/1pTuQhug6Dhl9XalKB0zUGf4FIdYFlpcX#scrollTo=IKzLS9ohzGVu) at the end for details.*
In this tutorial I'll show you how to use BERT with the huggingface PyTorch library to quickly and efficiently fine-tune a model to get near state of the art performance in sentence classification. More broadly, I describe the practical application of transfer learning in NLP to create high performance models with minimal effort on a range of NLP tasks.
This post is presented in two forms--as a blog post [here](http://mccormickml.com/2019/07/22/BERT-fine-tuning/) and as a Colab Notebook [here](https://colab.research.google.com/drive/1pTuQhug6Dhl9XalKB0zUGf4FIdYFlpcX).
The content is identical in both, but:
* The blog post includes a comments section for discussion.
* The Colab Notebook will allow you to run the code and inspect it as you read through.
I've also published a video walkthrough of this post on my YouTube channel! [Part 1](https://youtu.be/x66kkDnbzi4) and [Part 2](https://youtu.be/Hnvb9b7a_Ps).
# Contents
See "Table of contents" in the sidebar to the left.
# Introduction
## History
2018 was a breakthrough year in NLP. Transfer learning, particularly models like Allen AI's ELMO, OpenAI's Open-GPT, and Google's BERT allowed researchers to smash multiple benchmarks with minimal task-specific fine-tuning and provided the rest of the NLP community with pretrained models that could easily (with less data and less compute time) be fine-tuned and implemented to produce state of the art results. Unfortunately, for many starting out in NLP and even for some experienced practicioners, the theory and practical application of these powerful models is still not well understood.
## What is BERT?
BERT (Bidirectional Encoder Representations from Transformers), released in late 2018, is the model we will use in this tutorial to provide readers with a better understanding of and practical guidance for using transfer learning models in NLP. BERT is a method of pretraining language representations that was used to create models that NLP practicioners can then download and use for free. You can either use these models to extract high quality language features from your text data, or you can fine-tune these models on a specific task (classification, entity recognition, question answering, etc.) with your own data to produce state of the art predictions.
This post will explain how you can modify and fine-tune BERT to create a powerful NLP model that quickly gives you state of the art results.
## Advantages of Fine-Tuning
In this tutorial, we will use BERT to train a text classifier. Specifically, we will take the pre-trained BERT model, add an untrained layer of neurons on the end, and train the new model for our classification task. Why do this rather than train a train a specific deep learning model (a CNN, BiLSTM, etc.) that is well suited for the specific NLP task you need?
1. **Quicker Development**
* First, the pre-trained BERT model weights already encode a lot of information about our language. As a result, it takes much less time to train our fine-tuned model - it is as if we have already trained the bottom layers of our network extensively and only need to gently tune them while using their output as features for our classification task. In fact, the authors recommend only 2-4 epochs of training for fine-tuning BERT on a specific NLP task (compared to the hundreds of GPU hours needed to train the original BERT model or a LSTM from scratch!).
2. **Less Data**
* In addition and perhaps just as important, because of the pre-trained weights this method allows us to fine-tune our task on a much smaller dataset than would be required in a model that is built from scratch. A major drawback of NLP models built from scratch is that we often need a prohibitively large dataset in order to train our network to reasonable accuracy, meaning a lot of time and energy had to be put into dataset creation. By fine-tuning BERT, we are now able to get away with training a model to good performance on a much smaller amount of training data.
3. **Better Results**
* Finally, this simple fine-tuning procedure (typically adding one fully-connected layer on top of BERT and training for a few epochs) was shown to achieve state of the art results with minimal task-specific adjustments for a wide variety of tasks: classification, language inference, semantic similarity, question answering, etc. Rather than implementing custom and sometimes-obscure architetures shown to work well on a specific task, simply fine-tuning BERT is shown to be a better (or at least equal) alternative.
### A Shift in NLP
This shift to transfer learning parallels the same shift that took place in computer vision a few years ago. Creating a good deep learning network for computer vision tasks can take millions of parameters and be very expensive to train. Researchers discovered that deep networks learn hierarchical feature representations (simple features like edges at the lowest layers with gradually more complex features at higher layers). Rather than training a new network from scratch each time, the lower layers of a trained network with generalized image features could be copied and transfered for use in another network with a different task. It soon became common practice to download a pre-trained deep network and quickly retrain it for the new task or add additional layers on top - vastly preferable to the expensive process of training a network from scratch. For many, the introduction of deep pre-trained language models in 2018 (ELMO, BERT, ULMFIT, Open-GPT, etc.) signals the same shift to transfer learning in NLP that computer vision saw.
Let's get started!
[](https://bit.ly/30JzuBH)
# 1. Setup
## 1.1. Using Colab GPU for Training
Google Colab offers free GPUs and TPUs! Since we'll be training a large neural network it's best to take advantage of this (in this case we'll attach a GPU), otherwise training will take a very long time.
A GPU can be added by going to the menu and selecting:
`Edit 🡒 Notebook Settings 🡒 Hardware accelerator 🡒 (GPU)`
Then run the following cell to confirm that the GPU is detected.
```
import tensorflow as tf
# Get the GPU device name.
device_name = tf.test.gpu_device_name()
# The device name should look like the following:
if device_name == '/device:GPU:0':
print('Found GPU at: {}'.format(device_name))
else:
raise SystemError('GPU device not found')
```
In order for torch to use the GPU, we need to identify and specify the GPU as the device. Later, in our training loop, we will load data onto the device.
```
import torch
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
```
## 1.2. Installing the Hugging Face Library
Next, let's install the [transformers](https://github.com/huggingface/transformers) package from Hugging Face which will give us a pytorch interface for working with BERT. (This library contains interfaces for other pretrained language models like OpenAI's GPT and GPT-2.) We've selected the pytorch interface because it strikes a nice balance between the high-level APIs (which are easy to use but don't provide insight into how things work) and tensorflow code (which contains lots of details but often sidetracks us into lessons about tensorflow, when the purpose here is BERT!).
At the moment, the Hugging Face library seems to be the most widely accepted and powerful pytorch interface for working with BERT. In addition to supporting a variety of different pre-trained transformer models, the library also includes pre-built modifications of these models suited to your specific task. For example, in this tutorial we will use `BertForSequenceClassification`.
The library also includes task-specific classes for token classification, question answering, next sentence prediciton, etc. Using these pre-built classes simplifies the process of modifying BERT for your purposes.
```
!pip install transformers
```
The code in this notebook is actually a simplified version of the [run_glue.py](https://github.com/huggingface/transformers/blob/master/examples/run_glue.py) example script from huggingface.
`run_glue.py` is a helpful utility which allows you to pick which GLUE benchmark task you want to run on, and which pre-trained model you want to use (you can see the list of possible models [here](https://github.com/huggingface/transformers/blob/e6cff60b4cbc1158fbd6e4a1c3afda8dc224f566/examples/run_glue.py#L69)). It also supports using either the CPU, a single GPU, or multiple GPUs. It even supports using 16-bit precision if you want further speed up.
Unfortunately, all of this configurability comes at the cost of *readability*. In this Notebook, we've simplified the code greatly and added plenty of comments to make it clear what's going on.
# 2. Loading CoLA Dataset
We'll use [The Corpus of Linguistic Acceptability (CoLA)](https://nyu-mll.github.io/CoLA/) dataset for single sentence classification. It's a set of sentences labeled as grammatically correct or incorrect. It was first published in May of 2018, and is one of the tests included in the "GLUE Benchmark" on which models like BERT are competing.
## 2.1. Download & Extract
We'll use the `wget` package to download the dataset to the Colab instance's file system.
```
!pip install wget
```
The dataset is hosted on GitHub in this repo: https://nyu-mll.github.io/CoLA/
```
import wget
import os
print('Downloading dataset...')
# The URL for the dataset zip file.
url = 'https://nyu-mll.github.io/CoLA/cola_public_1.1.zip'
# Download the file (if we haven't already)
if not os.path.exists('./cola_public_1.1.zip'):
wget.download(url, './cola_public_1.1.zip')
```
Unzip the dataset to the file system. You can browse the file system of the Colab instance in the sidebar on the left.
```
# Unzip the dataset (if we haven't already)
if not os.path.exists('./cola_public/'):
!unzip cola_public_1.1.zip
```
## 2.2. Parse
We can see from the file names that both `tokenized` and `raw` versions of the data are available.
We can't use the pre-tokenized version because, in order to apply the pre-trained BERT, we *must* use the tokenizer provided by the model. This is because (1) the model has a specific, fixed vocabulary and (2) the BERT tokenizer has a particular way of handling out-of-vocabulary words.
We'll use pandas to parse the "in-domain" training set and look at a few of its properties and data points.
```
import pandas as pd
# Load the dataset into a pandas dataframe.
df = pd.read_csv("./cola_public/raw/in_domain_train.tsv", delimiter='\t', header=None, names=['sentence_source', 'label', 'label_notes', 'sentence'])
# Report the number of sentences.
print('Number of training sentences: {:,}\n'.format(df.shape[0]))
# Display 10 random rows from the data.
df.sample(10)
```
The two properties we actually care about are the the `sentence` and its `label`, which is referred to as the "acceptibility judgment" (0=unacceptable, 1=acceptable).
Here are five sentences which are labeled as not grammatically acceptible. Note how much more difficult this task is than something like sentiment analysis!
```
df.loc[df.label == 0].sample(5)[['sentence', 'label']]
```
Let's extract the sentences and labels of our training set as numpy ndarrays.
```
# Get the lists of sentences and their labels.
sentences = df.sentence.values
labels = df.label.values
```
# 3. Tokenization & Input Formatting
In this section, we'll transform our dataset into the format that BERT can be trained on.
## 3.1. BERT Tokenizer
To feed our text to BERT, it must be split into tokens, and then these tokens must be mapped to their index in the tokenizer vocabulary.
The tokenization must be performed by the tokenizer included with BERT--the below cell will download this for us. We'll be using the "uncased" version here.
```
from transformers import BertTokenizer
# Load the BERT tokenizer.
print('Loading BERT tokenizer...')
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased', do_lower_case=True)
```
Let's apply the tokenizer to one sentence just to see the output.
```
# Print the original sentence.
print(' Original: ', sentences[0])
# Print the sentence split into tokens.
print('Tokenized: ', tokenizer.tokenize(sentences[0]))
# Print the sentence mapped to token ids.
print('Token IDs: ', tokenizer.convert_tokens_to_ids(tokenizer.tokenize(sentences[0])))
```
When we actually convert all of our sentences, we'll use the `tokenize.encode` function to handle both steps, rather than calling `tokenize` and `convert_tokens_to_ids` separately.
Before we can do that, though, we need to talk about some of BERT's formatting requirements.
## 3.2. Required Formatting
The above code left out a few required formatting steps that we'll look at here.
*Side Note: The input format to BERT seems "over-specified" to me... We are required to give it a number of pieces of information which seem redundant, or like they could easily be inferred from the data without us explicity providing it. But it is what it is, and I suspect it will make more sense once I have a deeper understanding of the BERT internals.*
We are required to:
1. Add special tokens to the start and end of each sentence.
2. Pad & truncate all sentences to a single constant length.
3. Explicitly differentiate real tokens from padding tokens with the "attention mask".
### Special Tokens
**`[SEP]`**
At the end of every sentence, we need to append the special `[SEP]` token.
This token is an artifact of two-sentence tasks, where BERT is given two separate sentences and asked to determine something (e.g., can the answer to the question in sentence A be found in sentence B?).
I am not certain yet why the token is still required when we have only single-sentence input, but it is!
**`[CLS]`**
For classification tasks, we must prepend the special `[CLS]` token to the beginning of every sentence.
This token has special significance. BERT consists of 12 Transformer layers. Each transformer takes in a list of token embeddings, and produces the same number of embeddings on the output (but with the feature values changed, of course!).

On the output of the final (12th) transformer, *only the first embedding (corresponding to the [CLS] token) is used by the classifier*.
> "The first token of every sequence is always a special classification token (`[CLS]`). The final hidden state
corresponding to this token is used as the aggregate sequence representation for classification
tasks." (from the [BERT paper](https://arxiv.org/pdf/1810.04805.pdf))
You might think to try some pooling strategy over the final embeddings, but this isn't necessary. Because BERT is trained to only use this [CLS] token for classification, we know that the model has been motivated to encode everything it needs for the classification step into that single 768-value embedding vector. It's already done the pooling for us!
### Sentence Length & Attention Mask
The sentences in our dataset obviously have varying lengths, so how does BERT handle this?
BERT has two constraints:
1. All sentences must be padded or truncated to a single, fixed length.
2. The maximum sentence length is 512 tokens.
Padding is done with a special `[PAD]` token, which is at index 0 in the BERT vocabulary. The below illustration demonstrates padding out to a "MAX_LEN" of 8 tokens.
<img src="https://drive.google.com/uc?export=view&id=1cb5xeqLu_5vPOgs3eRnail2Y00Fl2pCo" width="600">
The "Attention Mask" is simply an array of 1s and 0s indicating which tokens are padding and which aren't (seems kind of redundant, doesn't it?!). This mask tells the "Self-Attention" mechanism in BERT not to incorporate these PAD tokens into its interpretation of the sentence.
The maximum length does impact training and evaluation speed, however.
For example, with a Tesla K80:
`MAX_LEN = 128 --> Training epochs take ~5:28 each`
`MAX_LEN = 64 --> Training epochs take ~2:57 each`
## 3.3. Tokenize Dataset
The transformers library provides a helpful `encode` function which will handle most of the parsing and data prep steps for us.
Before we are ready to encode our text, though, we need to decide on a **maximum sentence length** for padding / truncating to.
The below cell will perform one tokenization pass of the dataset in order to measure the maximum sentence length.
```
max_len = 0
# For every sentence...
for sent in sentences:
# Tokenize the text and add `[CLS]` and `[SEP]` tokens.
input_ids = tokenizer.encode(sent, add_special_tokens=True)
# Update the maximum sentence length.
max_len = max(max_len, len(input_ids))
print('Max sentence length: ', max_len)
```
Just in case there are some longer test sentences, I'll set the maximum length to 64.
Now we're ready to perform the real tokenization.
The `tokenizer.encode_plus` function combines multiple steps for us:
1. Split the sentence into tokens.
2. Add the special `[CLS]` and `[SEP]` tokens.
3. Map the tokens to their IDs.
4. Pad or truncate all sentences to the same length.
5. Create the attention masks which explicitly differentiate real tokens from `[PAD]` tokens.
The first four features are in `tokenizer.encode`, but I'm using `tokenizer.encode_plus` to get the fifth item (attention masks). Documentation is [here](https://huggingface.co/transformers/main_classes/tokenizer.html?highlight=encode_plus#transformers.PreTrainedTokenizer.encode_plus).
```
# Tokenize all of the sentences and map the tokens to thier word IDs.
input_ids = []
attention_masks = []
# For every sentence...
for sent in sentences:
# `encode_plus` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
# (5) Pad or truncate the sentence to `max_length`
# (6) Create attention masks for [PAD] tokens.
encoded_dict = tokenizer.encode_plus(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = 64, # Pad & truncate all sentences.
pad_to_max_length = True,
return_attention_mask = True, # Construct attn. masks.
return_tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding).
attention_masks.append(encoded_dict['attention_mask'])
# Convert the lists into tensors.
input_ids = torch.cat(input_ids, dim=0)
attention_masks = torch.cat(attention_masks, dim=0)
labels = torch.tensor(labels)
# Print sentence 0, now as a list of IDs.
print('Original: ', sentences[0])
print('Token IDs:', input_ids[0])
```
## 3.4. Training & Validation Split
Divide up our training set to use 90% for training and 10% for validation.
```
from torch.utils.data import TensorDataset, random_split
# Combine the training inputs into a TensorDataset.
dataset = TensorDataset(input_ids, attention_masks, labels)
# Create a 90-10 train-validation split.
# Calculate the number of samples to include in each set.
train_size = int(0.9 * len(dataset))
val_size = len(dataset) - train_size
# Divide the dataset by randomly selecting samples.
train_dataset, val_dataset = random_split(dataset, [train_size, val_size])
print('{:>5,} training samples'.format(train_size))
print('{:>5,} validation samples'.format(val_size))
```
We'll also create an iterator for our dataset using the torch DataLoader class. This helps save on memory during training because, unlike a for loop, with an iterator the entire dataset does not need to be loaded into memory.
```
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
# The DataLoader needs to know our batch size for training, so we specify it
# here. For fine-tuning BERT on a specific task, the authors recommend a batch
# size of 16 or 32.
batch_size = 32
# Create the DataLoaders for our training and validation sets.
# We'll take training samples in random order.
train_dataloader = DataLoader(
train_dataset, # The training samples.
sampler = RandomSampler(train_dataset), # Select batches randomly
batch_size = batch_size # Trains with this batch size.
)
# For validation the order doesn't matter, so we'll just read them sequentially.
validation_dataloader = DataLoader(
val_dataset, # The validation samples.
sampler = SequentialSampler(val_dataset), # Pull out batches sequentially.
batch_size = batch_size # Evaluate with this batch size.
)
```
# 4. Train Our Classification Model
Now that our input data is properly formatted, it's time to fine tune the BERT model.
## 4.1. BertForSequenceClassification
For this task, we first want to modify the pre-trained BERT model to give outputs for classification, and then we want to continue training the model on our dataset until that the entire model, end-to-end, is well-suited for our task.
Thankfully, the huggingface pytorch implementation includes a set of interfaces designed for a variety of NLP tasks. Though these interfaces are all built on top of a trained BERT model, each has different top layers and output types designed to accomodate their specific NLP task.
Here is the current list of classes provided for fine-tuning:
* BertModel
* BertForPreTraining
* BertForMaskedLM
* BertForNextSentencePrediction
* **BertForSequenceClassification** - The one we'll use.
* BertForTokenClassification
* BertForQuestionAnswering
The documentation for these can be found under [here](https://huggingface.co/transformers/v2.2.0/model_doc/bert.html).
We'll be using [BertForSequenceClassification](https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#bertforsequenceclassification). This is the normal BERT model with an added single linear layer on top for classification that we will use as a sentence classifier. As we feed input data, the entire pre-trained BERT model and the additional untrained classification layer is trained on our specific task.
OK, let's load BERT! There are a few different pre-trained BERT models available. "bert-base-uncased" means the version that has only lowercase letters ("uncased") and is the smaller version of the two ("base" vs "large").
The documentation for `from_pretrained` can be found [here](https://huggingface.co/transformers/v2.2.0/main_classes/model.html#transformers.PreTrainedModel.from_pretrained), with the additional parameters defined [here](https://huggingface.co/transformers/v2.2.0/main_classes/configuration.html#transformers.PretrainedConfig).
```
from transformers import BertForSequenceClassification, AdamW, BertConfig
# Load BertForSequenceClassification, the pretrained BERT model with a single
# linear classification layer on top.
model = BertForSequenceClassification.from_pretrained(
"bert-base-uncased", # Use the 12-layer BERT model, with an uncased vocab.
num_labels = 2, # The number of output labels--2 for binary classification.
# You can increase this for multi-class tasks.
output_attentions = False, # Whether the model returns attentions weights.
output_hidden_states = False, # Whether the model returns all hidden-states.
)
# Tell pytorch to run this model on the GPU.
model.cuda()
```
Just for curiosity's sake, we can browse all of the model's parameters by name here.
In the below cell, I've printed out the names and dimensions of the weights for:
1. The embedding layer.
2. The first of the twelve transformers.
3. The output layer.
```
# Get all of the model's parameters as a list of tuples.
params = list(model.named_parameters())
print('The BERT model has {:} different named parameters.\n'.format(len(params)))
print('==== Embedding Layer ====\n')
for p in params[0:5]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== First Transformer ====\n')
for p in params[5:21]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== Output Layer ====\n')
for p in params[-4:]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
```
## 4.2. Optimizer & Learning Rate Scheduler
Now that we have our model loaded we need to grab the training hyperparameters from within the stored model.
For the purposes of fine-tuning, the authors recommend choosing from the following values (from Appendix A.3 of the [BERT paper](https://arxiv.org/pdf/1810.04805.pdf)):
>- **Batch size:** 16, 32
- **Learning rate (Adam):** 5e-5, 3e-5, 2e-5
- **Number of epochs:** 2, 3, 4
We chose:
* Batch size: 32 (set when creating our DataLoaders)
* Learning rate: 2e-5
* Epochs: 4 (we'll see that this is probably too many...)
The epsilon parameter `eps = 1e-8` is "a very small number to prevent any division by zero in the implementation" (from [here](https://machinelearningmastery.com/adam-optimization-algorithm-for-deep-learning/)).
You can find the creation of the AdamW optimizer in `run_glue.py` [here](https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L109).
```
# Note: AdamW is a class from the huggingface library (as opposed to pytorch)
# I believe the 'W' stands for 'Weight Decay fix"
optimizer = AdamW(model.parameters(),
lr = 2e-5, # args.learning_rate - default is 5e-5, our notebook had 2e-5
eps = 1e-8 # args.adam_epsilon - default is 1e-8.
)
from transformers import get_linear_schedule_with_warmup
# Number of training epochs. The BERT authors recommend between 2 and 4.
# We chose to run for 4, but we'll see later that this may be over-fitting the
# training data.
epochs = 4
# Total number of training steps is [number of batches] x [number of epochs].
# (Note that this is not the same as the number of training samples).
total_steps = len(train_dataloader) * epochs
# Create the learning rate scheduler.
scheduler = get_linear_schedule_with_warmup(optimizer,
num_warmup_steps = 0, # Default value in run_glue.py
num_training_steps = total_steps)
```
## 4.3. Training Loop
Below is our training loop. There's a lot going on, but fundamentally for each pass in our loop we have a trianing phase and a validation phase.
> *Thank you to [Stas Bekman](https://ca.linkedin.com/in/stasbekman) for contributing the insights and code for using validation loss to detect over-fitting!*
**Training:**
- Unpack our data inputs and labels
- Load data onto the GPU for acceleration
- Clear out the gradients calculated in the previous pass.
- In pytorch the gradients accumulate by default (useful for things like RNNs) unless you explicitly clear them out.
- Forward pass (feed input data through the network)
- Backward pass (backpropagation)
- Tell the network to update parameters with optimizer.step()
- Track variables for monitoring progress
**Evalution:**
- Unpack our data inputs and labels
- Load data onto the GPU for acceleration
- Forward pass (feed input data through the network)
- Compute loss on our validation data and track variables for monitoring progress
Pytorch hides all of the detailed calculations from us, but we've commented the code to point out which of the above steps are happening on each line.
> *PyTorch also has some [beginner tutorials](https://pytorch.org/tutorials/beginner/blitz/cifar10_tutorial.html#sphx-glr-beginner-blitz-cifar10-tutorial-py) which you may also find helpful.*
Define a helper function for calculating accuracy.
```
import numpy as np
# Function to calculate the accuracy of our predictions vs labels
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
```
Helper function for formatting elapsed times as `hh:mm:ss`
```
import time
import datetime
def format_time(elapsed):
'''
Takes a time in seconds and returns a string hh:mm:ss
'''
# Round to the nearest second.
elapsed_rounded = int(round((elapsed)))
# Format as hh:mm:ss
return str(datetime.timedelta(seconds=elapsed_rounded))
```
We're ready to kick off the training!
```
import random
import numpy as np
# This training code is based on the `run_glue.py` script here:
# https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L128
# Set the seed value all over the place to make this reproducible.
seed_val = 42
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# We'll store a number of quantities such as training and validation loss,
# validation accuracy, and timings.
training_stats = []
# Measure the total training time for the whole run.
total_t0 = time.time()
# For each epoch...
for epoch_i in range(0, epochs):
# ========================================
# Training
# ========================================
# Perform one full pass over the training set.
print("")
print('======== Epoch {:} / {:} ========'.format(epoch_i + 1, epochs))
print('Training...')
# Measure how long the training epoch takes.
t0 = time.time()
# Reset the total loss for this epoch.
total_train_loss = 0
# Put the model into training mode. Don't be mislead--the call to
# `train` just changes the *mode*, it doesn't *perform* the training.
# `dropout` and `batchnorm` layers behave differently during training
# vs. test (source: https://stackoverflow.com/questions/51433378/what-does-model-train-do-in-pytorch)
model.train()
# For each batch of training data...
for step, batch in enumerate(train_dataloader):
# Progress update every 40 batches.
if step % 40 == 0 and not step == 0:
# Calculate elapsed time in minutes.
elapsed = format_time(time.time() - t0)
# Report progress.
print(' Batch {:>5,} of {:>5,}. Elapsed: {:}.'.format(step, len(train_dataloader), elapsed))
# Unpack this training batch from our dataloader.
#
# As we unpack the batch, we'll also copy each tensor to the GPU using the
# `to` method.
#
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1]: attention masks
# [2]: labels
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
# Always clear any previously calculated gradients before performing a
# backward pass. PyTorch doesn't do this automatically because
# accumulating the gradients is "convenient while training RNNs".
# (source: https://stackoverflow.com/questions/48001598/why-do-we-need-to-call-zero-grad-in-pytorch)
model.zero_grad()
# Perform a forward pass (evaluate the model on this training batch).
# In PyTorch, calling `model` will in turn call the model's `forward`
# function and pass down the arguments. The `forward` function is
# documented here:
# https://huggingface.co/transformers/model_doc/bert.html#bertforsequenceclassification
# The results are returned in a results object, documented here:
# https://huggingface.co/transformers/main_classes/output.html#transformers.modeling_outputs.SequenceClassifierOutput
# Specifically, we'll get the loss (because we provided labels) and the
# "logits"--the model outputs prior to activation.
result = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels,
return_dict=True)
loss = result.loss
logits = result.logits
# Accumulate the training loss over all of the batches so that we can
# calculate the average loss at the end. `loss` is a Tensor containing a
# single value; the `.item()` function just returns the Python value
# from the tensor.
total_train_loss += loss.item()
# Perform a backward pass to calculate the gradients.
loss.backward()
# Clip the norm of the gradients to 1.0.
# This is to help prevent the "exploding gradients" problem.
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
# Update parameters and take a step using the computed gradient.
# The optimizer dictates the "update rule"--how the parameters are
# modified based on their gradients, the learning rate, etc.
optimizer.step()
# Update the learning rate.
scheduler.step()
# Calculate the average loss over all of the batches.
avg_train_loss = total_train_loss / len(train_dataloader)
# Measure how long this epoch took.
training_time = format_time(time.time() - t0)
print("")
print(" Average training loss: {0:.2f}".format(avg_train_loss))
print(" Training epcoh took: {:}".format(training_time))
# ========================================
# Validation
# ========================================
# After the completion of each training epoch, measure our performance on
# our validation set.
print("")
print("Running Validation...")
t0 = time.time()
# Put the model in evaluation mode--the dropout layers behave differently
# during evaluation.
model.eval()
# Tracking variables
total_eval_accuracy = 0
total_eval_loss = 0
nb_eval_steps = 0
# Evaluate data for one epoch
for batch in validation_dataloader:
# Unpack this training batch from our dataloader.
#
# As we unpack the batch, we'll also copy each tensor to the GPU using
# the `to` method.
#
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1]: attention masks
# [2]: labels
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
# Tell pytorch not to bother with constructing the compute graph during
# the forward pass, since this is only needed for backprop (training).
with torch.no_grad():
# Forward pass, calculate logit predictions.
# token_type_ids is the same as the "segment ids", which
# differentiates sentence 1 and 2 in 2-sentence tasks.
result = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels,
return_dict=True)
# Get the loss and "logits" output by the model. The "logits" are the
# output values prior to applying an activation function like the
# softmax.
loss = result.loss
logits = result.logits
# Accumulate the validation loss.
total_eval_loss += loss.item()
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Calculate the accuracy for this batch of test sentences, and
# accumulate it over all batches.
total_eval_accuracy += flat_accuracy(logits, label_ids)
# Report the final accuracy for this validation run.
avg_val_accuracy = total_eval_accuracy / len(validation_dataloader)
print(" Accuracy: {0:.2f}".format(avg_val_accuracy))
# Calculate the average loss over all of the batches.
avg_val_loss = total_eval_loss / len(validation_dataloader)
# Measure how long the validation run took.
validation_time = format_time(time.time() - t0)
print(" Validation Loss: {0:.2f}".format(avg_val_loss))
print(" Validation took: {:}".format(validation_time))
# Record all statistics from this epoch.
training_stats.append(
{
'epoch': epoch_i + 1,
'Training Loss': avg_train_loss,
'Valid. Loss': avg_val_loss,
'Valid. Accur.': avg_val_accuracy,
'Training Time': training_time,
'Validation Time': validation_time
}
)
print("")
print("Training complete!")
print("Total training took {:} (h:mm:ss)".format(format_time(time.time()-total_t0)))
```
Let's view the summary of the training process.
```
import pandas as pd
# Display floats with two decimal places.
pd.set_option('precision', 2)
# Create a DataFrame from our training statistics.
df_stats = pd.DataFrame(data=training_stats)
# Use the 'epoch' as the row index.
df_stats = df_stats.set_index('epoch')
# A hack to force the column headers to wrap.
#df = df.style.set_table_styles([dict(selector="th",props=[('max-width', '70px')])])
# Display the table.
df_stats
```
Notice that, while the the training loss is going down with each epoch, the validation loss is increasing! This suggests that we are training our model too long, and it's over-fitting on the training data.
(For reference, we are using 7,695 training samples and 856 validation samples).
Validation Loss is a more precise measure than accuracy, because with accuracy we don't care about the exact output value, but just which side of a threshold it falls on.
If we are predicting the correct answer, but with less confidence, then validation loss will catch this, while accuracy will not.
```
import matplotlib.pyplot as plt
% matplotlib inline
import seaborn as sns
# Use plot styling from seaborn.
sns.set(style='darkgrid')
# Increase the plot size and font size.
sns.set(font_scale=1.5)
plt.rcParams["figure.figsize"] = (12,6)
# Plot the learning curve.
plt.plot(df_stats['Training Loss'], 'b-o', label="Training")
plt.plot(df_stats['Valid. Loss'], 'g-o', label="Validation")
# Label the plot.
plt.title("Training & Validation Loss")
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.legend()
plt.xticks([1, 2, 3, 4])
plt.show()
```
# 5. Performance On Test Set
Now we'll load the holdout dataset and prepare inputs just as we did with the training set. Then we'll evaluate predictions using [Matthew's correlation coefficient](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.matthews_corrcoef.html) because this is the metric used by the wider NLP community to evaluate performance on CoLA. With this metric, +1 is the best score, and -1 is the worst score. This way, we can see how well we perform against the state of the art models for this specific task.
### 5.1. Data Preparation
We'll need to apply all of the same steps that we did for the training data to prepare our test data set.
```
import pandas as pd
# Load the dataset into a pandas dataframe.
df = pd.read_csv("./cola_public/raw/out_of_domain_dev.tsv", delimiter='\t', header=None, names=['sentence_source', 'label', 'label_notes', 'sentence'])
# Report the number of sentences.
print('Number of test sentences: {:,}\n'.format(df.shape[0]))
# Create sentence and label lists
sentences = df.sentence.values
labels = df.label.values
# Tokenize all of the sentences and map the tokens to thier word IDs.
input_ids = []
attention_masks = []
# For every sentence...
for sent in sentences:
# `encode_plus` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
# (5) Pad or truncate the sentence to `max_length`
# (6) Create attention masks for [PAD] tokens.
encoded_dict = tokenizer.encode_plus(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = 64, # Pad & truncate all sentences.
pad_to_max_length = True,
return_attention_mask = True, # Construct attn. masks.
return_tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding).
attention_masks.append(encoded_dict['attention_mask'])
# Convert the lists into tensors.
input_ids = torch.cat(input_ids, dim=0)
attention_masks = torch.cat(attention_masks, dim=0)
labels = torch.tensor(labels)
# Set the batch size.
batch_size = 32
# Create the DataLoader.
prediction_data = TensorDataset(input_ids, attention_masks, labels)
prediction_sampler = SequentialSampler(prediction_data)
prediction_dataloader = DataLoader(prediction_data, sampler=prediction_sampler, batch_size=batch_size)
```
## 5.2. Evaluate on Test Set
With the test set prepared, we can apply our fine-tuned model to generate predictions on the test set.
```
# Prediction on test set
print('Predicting labels for {:,} test sentences...'.format(len(input_ids)))
# Put model in evaluation mode
model.eval()
# Tracking variables
predictions , true_labels = [], []
# Predict
for batch in prediction_dataloader:
# Add batch to GPU
batch = tuple(t.to(device) for t in batch)
# Unpack the inputs from our dataloader
b_input_ids, b_input_mask, b_labels = batch
# Telling the model not to compute or store gradients, saving memory and
# speeding up prediction
with torch.no_grad():
# Forward pass, calculate logit predictions.
result = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
return_dict=True)
logits = result.logits
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Store predictions and true labels
predictions.append(logits)
true_labels.append(label_ids)
print(' DONE.')
```
Accuracy on the CoLA benchmark is measured using the "[Matthews correlation coefficient](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.matthews_corrcoef.html)" (MCC).
We use MCC here because the classes are imbalanced:
```
print('Positive samples: %d of %d (%.2f%%)' % (df.label.sum(), len(df.label), (df.label.sum() / len(df.label) * 100.0)))
from sklearn.metrics import matthews_corrcoef
matthews_set = []
# Evaluate each test batch using Matthew's correlation coefficient
print('Calculating Matthews Corr. Coef. for each batch...')
# For each input batch...
for i in range(len(true_labels)):
# The predictions for this batch are a 2-column ndarray (one column for "0"
# and one column for "1"). Pick the label with the highest value and turn this
# in to a list of 0s and 1s.
pred_labels_i = np.argmax(predictions[i], axis=1).flatten()
# Calculate and store the coef for this batch.
matthews = matthews_corrcoef(true_labels[i], pred_labels_i)
matthews_set.append(matthews)
```
The final score will be based on the entire test set, but let's take a look at the scores on the individual batches to get a sense of the variability in the metric between batches.
Each batch has 32 sentences in it, except the last batch which has only (516 % 32) = 4 test sentences in it.
```
# Create a barplot showing the MCC score for each batch of test samples.
ax = sns.barplot(x=list(range(len(matthews_set))), y=matthews_set, ci=None)
plt.title('MCC Score per Batch')
plt.ylabel('MCC Score (-1 to +1)')
plt.xlabel('Batch #')
plt.show()
```
Now we'll combine the results for all of the batches and calculate our final MCC score.
```
# Combine the results across all batches.
flat_predictions = np.concatenate(predictions, axis=0)
# For each sample, pick the label (0 or 1) with the higher score.
flat_predictions = np.argmax(flat_predictions, axis=1).flatten()
# Combine the correct labels for each batch into a single list.
flat_true_labels = np.concatenate(true_labels, axis=0)
# Calculate the MCC
mcc = matthews_corrcoef(flat_true_labels, flat_predictions)
print('Total MCC: %.3f' % mcc)
```
Cool! In about half an hour and without doing any hyperparameter tuning (adjusting the learning rate, epochs, batch size, ADAM properties, etc.) we are able to get a good score.
> *Note: To maximize the score, we should remove the "validation set" (which we used to help determine how many epochs to train for) and train on the entire training set.*
The library documents the expected accuracy for this benchmark [here](https://huggingface.co/transformers/examples.html#glue) as `49.23`.
You can also look at the official leaderboard [here](https://gluebenchmark.com/leaderboard/submission/zlssuBTm5XRs0aSKbFYGVIVdvbj1/-LhijX9VVmvJcvzKymxy).
Note that (due to the small dataset size?) the accuracy can vary significantly between runs.
# Conclusion
This post demonstrates that with a pre-trained BERT model you can quickly and effectively create a high quality model with minimal effort and training time using the pytorch interface, regardless of the specific NLP task you are interested in.
# Appendix
## A1. Saving & Loading Fine-Tuned Model
This first cell (taken from `run_glue.py` [here](https://github.com/huggingface/transformers/blob/35ff345fc9df9e777b27903f11fa213e4052595b/examples/run_glue.py#L495)) writes the model and tokenizer out to disk.
```
import os
# Saving best-practices: if you use defaults names for the model, you can reload it using from_pretrained()
output_dir = './model_save/'
# Create output directory if needed
if not os.path.exists(output_dir):
os.makedirs(output_dir)
print("Saving model to %s" % output_dir)
# Save a trained model, configuration and tokenizer using `save_pretrained()`.
# They can then be reloaded using `from_pretrained()`
model_to_save = model.module if hasattr(model, 'module') else model # Take care of distributed/parallel training
model_to_save.save_pretrained(output_dir)
tokenizer.save_pretrained(output_dir)
# Good practice: save your training arguments together with the trained model
# torch.save(args, os.path.join(output_dir, 'training_args.bin'))
```
Let's check out the file sizes, out of curiosity.
```
!ls -l --block-size=K ./model_save/
```
The largest file is the model weights, at around 418 megabytes.
```
!ls -l --block-size=M ./model_save/pytorch_model.bin
```
To save your model across Colab Notebook sessions, download it to your local machine, or ideally copy it to your Google Drive.
```
# Mount Google Drive to this Notebook instance.
from google.colab import drive
drive.mount('/content/drive')
# Copy the model files to a directory in your Google Drive.
!cp -r ./model_save/ "./drive/Shared drives/ChrisMcCormick.AI/Blog Posts/BERT Fine-Tuning/"
```
The following functions will load the model back from disk.
```
# Load a trained model and vocabulary that you have fine-tuned
model = model_class.from_pretrained(output_dir)
tokenizer = tokenizer_class.from_pretrained(output_dir)
# Copy the model to the GPU.
model.to(device)
```
## A.2. Weight Decay
The huggingface example includes the following code block for enabling weight decay, but the default decay rate is "0.0", so I moved this to the appendix.
This block essentially tells the optimizer to not apply weight decay to the bias terms (e.g., $ b $ in the equation $ y = Wx + b $ ). Weight decay is a form of regularization--after calculating the gradients, we multiply them by, e.g., 0.99.
```
# This code is taken from:
# https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L102
# Don't apply weight decay to any parameters whose names include these tokens.
# (Here, the BERT doesn't have `gamma` or `beta` parameters, only `bias` terms)
no_decay = ['bias', 'LayerNorm.weight']
# Separate the `weight` parameters from the `bias` parameters.
# - For the `weight` parameters, this specifies a 'weight_decay_rate' of 0.01.
# - For the `bias` parameters, the 'weight_decay_rate' is 0.0.
optimizer_grouped_parameters = [
# Filter for all parameters which *don't* include 'bias', 'gamma', 'beta'.
{'params': [p for n, p in param_optimizer if not any(nd in n for nd in no_decay)],
'weight_decay_rate': 0.1},
# Filter for parameters which *do* include those.
{'params': [p for n, p in param_optimizer if any(nd in n for nd in no_decay)],
'weight_decay_rate': 0.0}
]
# Note - `optimizer_grouped_parameters` only includes the parameter values, not
# the names.
```
# Revision History
**Version 4** - *Feb 2nd, 2020* - (current)
* Updated all calls to `model` (fine-tuning and evaluation) to use the [`SequenceClassifierOutput`](https://huggingface.co/transformers/main_classes/output.html#transformers.modeling_outputs.SequenceClassifierOutput) class.
* Moved illustration images to Google Drive--Colab appears to no longer support images at external URLs.
**Version 3** - *Mar 18th, 2020*
* Simplified the tokenization and input formatting (for both training and test) by leveraging the `tokenizer.encode_plus` function.
`encode_plus` handles padding *and* creates the attention masks for us.
* Improved explanation of attention masks.
* Switched to using `torch.utils.data.random_split` for creating the training-validation split.
* Added a summary table of the training statistics (validation loss, time per epoch, etc.).
* Added validation loss to the learning curve plot, so we can see if we're overfitting.
* Thank you to [Stas Bekman](https://ca.linkedin.com/in/stasbekman) for contributing this!
* Displayed the per-batch MCC as a bar plot.
**Version 2** - *Dec 20th, 2019* - [link](https://colab.research.google.com/drive/1Y4o3jh3ZH70tl6mCd76vz_IxX23biCPP)
* huggingface renamed their library to `transformers`.
* Updated the notebook to use the `transformers` library.
**Version 1** - *July 22nd, 2019*
* Initial version.
## Further Work
* It might make more sense to use the MCC score for “validation accuracy”, but I’ve left it out so as not to have to explain it earlier in the Notebook.
* Seeding -- I’m not convinced that setting the seed values at the beginning of the training loop is actually creating reproducible results…
* The MCC score seems to vary substantially across different runs. It would be interesting to run this example a number of times and show the variance.
| github_jupyter |
# Example: Polynomial Cureve Fitting
Observse a real-valued input variable $x$ $\rightarrow$ predict a real-valued target variable $t$
* $\textbf{x} \equiv (x_1, \cdots, x_i, \cdots, x_N)^T, \quad x_i \in [0, 1]$
* $\textbf{t} \equiv (t_1, \cdots, t_i, \cdots, t_N)^T, \quad t_i = \sin(2\pi x_i) + N(\mu, \sigma^2)$
```
import numpy as np
import matplotlib.pylab as plt
# making data
seed = 62
np.random.seed(seed)
N = 10
x = np.random.rand(N)
t = np.sin(2*np.pi*x) + np.random.randn(N) * 0.1
x_sin = np.linspace(0, 1)
t_sin = np.sin(2*np.pi*x_sin)
plt.plot(x_sin, t_sin, c='green')
plt.scatter(x, t)
plt.xlabel('x', fontsize=16)
plt.ylabel('t', rotation=0, fontsize=16)
plt.show()
```
* Goal: exploit this training set in order to make predictions of the value $\hat{t}$ of the target variable for some new value $\hat{x}$ of the input variable.
* Use some theories:
* Probability theory: provides a framework for expressing such uncertainty in a precise and quantitative manner
* Decision theory: allows us to exploit this probabilistic representation in order to make predictions that are optimal according to appropriate criteria
* For the moment, let's use polynomial function, where $M$ is the order of polynomial. $y(x, \mathbf{w})$ is a linear function of coefficients ($\mathbf{w}$)
$$y(x, \mathbf{w}) = w_0 + w_1 x + w_2 x^2 + \cdots + w_M x^M = \sum_{j=0}^{M} w_j x^j$$
```
def vandermonde_matrix(x, m):
"""we will introduce vandermonde_matrix, when we find solution of polynomial regression"""
return np.array([x**i for i in range(m+1)]).T
def polynomial_function(x, w, m):
assert w.size == m+1, "coefficients number must same as M+1"
V = vandermonde_matrix(x, m) # shape (x.size, M+1)
return np.dot(V, w)
np.random.seed(seed)
M = 3
w = np.random.randn(M+1)
t_hat = polynomial_function(x, w, M)
print(t_hat.round(3))
```
* The values of the coefficients will be determined by fitting the polynomial to the training data, this can be done by minimizing an error function, which measure the misfit between the function $y(x, \mathbf{w})$ and training data points.
$$E(\mathbf{w}) = \dfrac{1}{2} \sum_{n=1}^{N} (y(x_n, \mathbf{w}) - t_n)^2$$
```
def error_function(pred, target):
return (1/2)*((pred-target)**2).sum()
error_value = error_function(t_hat, t)
error_value
```
* Because error function is quadratic function of $\mathbf{w}$, its derivatives with respect to the coefficients will be linear in the elements of $\mathbf{w}$, so the minimization of the error function has a unique solution.
* The remain problem is choosing the order $M$, this is called **model comparison or model selection**.
* Then how to choose optimal $M$?
* use test data with 100 data points
* evaluate the residual value of error
```
np.random.seed(seed)
N_test = 100
x_test = np.random.rand(N_test)
t_test = np.sin(2*np.pi*x_test) + np.random.randn(N_test) * 0.1
plt.plot(x_sin, t_sin, c='green')
plt.scatter(x_test, t_test, c='red')
plt.xlabel('x', fontsize=16)
plt.ylabel('t', rotation=0, fontsize=16)
plt.show()
def root_mean_square_error(error, n_samples):
return np.sqrt(2*error/n_samples)
# M=3
error = error_function(polynomial_function(x_test, w, M), t_test)
rms = root_mean_square_error(error, N_test)
rms
```
### using normal equation to find soulution
First define $V$(size is $(N, M+1)$) matrix named **Vandermode matrix** which is looks like below. $M$ is degree of polynomial function.
$$V = \begin{bmatrix}
1 & x_1 & x_1^2 & \cdots & x_1^M \\
1 & x_2 & x_2^2 & \cdots & x_2^M \\
\vdots & \vdots & \vdots & \ddots & \vdots \\
1 & x_N & x_N^2 & \cdots & x_N^M
\end{bmatrix}$$
```
def vandermonde_matrix(x, m):
"""vandermonde matrix"""
return np.array([x**i for i in range(m+1)]).T
M = 3
V = vandermonde_matrix(x, M)
print(V.round(3))
```
So, we can define polynomial as $y=V\cdot w$. Where $w$ is a column vector called **coefficients** , $w = [w_0, w_1, \cdots , w_M]^T$
$$y = \begin{bmatrix} y_1 \\ y_2 \\ \vdots \\ y_N \end{bmatrix} =
\begin{bmatrix} w_0 + w_1x_1 + w_2x_1^2 + \cdots + w_Mx_1^M \\ w_0 + w_1x_2 + w_2x_2^2 + \cdots + w_Mx_2^M \\ \vdots \\
w_0 + w_1x_N + w_2x_N^2 + \cdots + w_Mx_N^M \end{bmatrix}$$
We already defined error function, $E(\mathbf{w}) = \dfrac{1}{2} \sum_{n=1}^{N} (y(x_n, \mathbf{w}) - t_n)^2 = \dfrac{1}{2} \Vert y - V \cdot w \Vert^2$. which is can solved by minimization, $\hat{w} = \underset{w}{\arg \min} E(w)$.
Define residual $r = y - V \cdot w$ then error function becomes $E(\mathbf{w}) = \dfrac{1}{2} r^2 $ Because error function is quadratic function, the minimization of the error function has a unique solution.
Then we can get derivatives, and when it becomes to $0$, error function has minimum value.
$$\begin{aligned} \dfrac{\partial E}{\partial w} &= \begin{bmatrix} \dfrac{\partial E}{\partial w_0} \\ \dfrac{\partial E}{\partial w_1} \\ \vdots \\ \dfrac{\partial E}{\partial w_M} \end{bmatrix} \\
&= \begin{bmatrix}
\dfrac{\partial E}{\partial r_1}\dfrac{\partial r_1}{\partial w_0} + \dfrac{\partial E}{\partial r_2}\dfrac{\partial r_2}{\partial w_0} + \cdots +\dfrac{\partial E}{\partial r_N}\dfrac{\partial r_N}{\partial w_0} \\
\dfrac{\partial E}{\partial r_1}\dfrac{\partial r_1}{\partial w_1} + \dfrac{\partial E}{\partial r_2}\dfrac{\partial r_2}{\partial w_1} + \cdots +\dfrac{\partial E}{\partial r_N}\dfrac{\partial r_N}{\partial w_1} \\
\vdots \\
\dfrac{\partial E}{\partial r_1}\dfrac{\partial r_1}{\partial w_M} + \dfrac{\partial E}{\partial r_2}\dfrac{\partial r_2}{\partial w_M} + \cdots +\dfrac{\partial E}{\partial r_N}\dfrac{\partial r_N}{\partial w_M}
\end{bmatrix} \\
&= \begin{bmatrix}
\dfrac{\partial r_1}{\partial w_0} & \dfrac{\partial r_2}{\partial w_0} & \cdots & \dfrac{\partial r_N}{\partial w_0} \\
\dfrac{\partial r_1}{\partial w_1} & \dfrac{\partial r_2}{\partial w_1} & \cdots & \dfrac{\partial r_N}{\partial w_1} \\
\vdots & \vdots & \ddots & \vdots \\
\dfrac{\partial r_1}{\partial w_M} & \dfrac{\partial r_2}{\partial w_M} & \cdots & \dfrac{\partial r_N}{\partial w_M}
\end{bmatrix} \cdot
\begin{bmatrix} \dfrac{\partial E}{\partial r_1} \\ \dfrac{\partial E}{\partial r_2} \\ \vdots \\ \dfrac{\partial E}{\partial r_N} \end{bmatrix} \\
&= \dfrac{\partial r}{\partial w} \cdot \dfrac{\partial E}{\partial r} \\
&= V^T \cdot (y - V\cdot w) = 0
\end{aligned}$$
So, we can find solution of coefficient $w$.
$$w = (V^TV)^{-1}V^Ty$$
```
def poly_solution(x, t, m):
V = vandermonde_matrix(x, m)
return np.linalg.inv(np.dot(V.T, V)).dot(V.T).dot(t)
print(f"Solution of coefficients are {poly_solution(x, t, M).round(3)}")
# confirm we are right
from numpy.polynomial import polynomial as P
print(P.polyfit(x, t, M).round(3))
```
Let's find optimal degree of polynomial now!
```
def get_rms_error(t_hat, t, n_sample, m):
error = error_function(t_hat, t)
rms = root_mean_square_error(error, n_sample)
return rms
all_w = []
all_rms_train = []
all_rms_test = []
for m in range(10):
optimal_w = poly_solution(x, t, m)
t_hat = polynomial_function(x, optimal_w, m)
t_hat_test = polynomial_function(x_test, optimal_w, m)
rms_train = get_rms_error(t_hat, t, N, m) # N=10
rms_test = get_rms_error(t_hat_test, t_test, N_test, m) # N_test = 100
print(f"M={m} | rms_train: {rms_train:.4f} rms_test: {rms_test:.4f}")
# Plot predicted line
plt.plot(x_sin, t_sin, c="green", label="sin function")
plt.plot(x_sin, polynomial_function(x_sin, optimal_w, m), c="red", label=f"model M={m}")
plt.scatter(x, t)
plt.xlim((0, 1))
plt.ylim((-1.25, 1.25))
plt.xlabel('x', fontsize=16)
plt.ylabel('t', rotation=0, fontsize=16)
plt.legend()
plt.show()
all_w.append(optimal_w)
all_rms_train.append(rms_train)
all_rms_test.append(rms_test)
plt.scatter(np.arange(10), all_rms_train, facecolors='none', edgecolors='b')
plt.plot(np.arange(10), all_rms_train, c='b', label='Training')
plt.scatter(np.arange(len(all_rms_test)), all_rms_test, facecolors='none', edgecolors='r')
plt.plot(np.arange(len(all_rms_test)), all_rms_test, c='r', label='Test')
plt.legend()
plt.xlim((-0.1, 10))
plt.ylim((-0.1, 1.2))
plt.ylabel("root-mean-squared Error", fontsize=16)
plt.xlabel("M", fontsize=16)
plt.show()
np.set_printoptions(precision=3)
for i in [0, 1, 3, 9]:
print(f"coefficients at M={i} is {all_w[i].round(3)}")
```
### Test for Different size of datas
```
np.random.seed(seed)
N1 = 15
N2 = 100
x1, x2 = np.random.rand(N1), np.random.rand(N2)
t1 = np.sin(2*np.pi*x1) + np.random.randn(N1) * 0.1
t2 = np.sin(2*np.pi*x2) + np.random.randn(N2) * 0.1
optimal_w1 = poly_solution(x1, t1, m=9)
optimal_w2 = poly_solution(x2, t2, m=9)
# Plot predicted line
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(16, 6))
def plot(x, t, x_sin, t_sin, optimal_w, m, ax):
ax.plot(x_sin, t_sin, c="green", label="sin function")
ax.plot(x_sin, polynomial_function(x_sin, optimal_w, m), c="red", label=f"model N={len(x)}")
ax.scatter(x, t)
ax.set_xlim((0, 1))
ax.set_ylim((-1.25, 1.25))
ax.set_xlabel('x', fontsize=16)
ax.set_ylabel('t', rotation=0, fontsize=16)
ax.legend()
plot(x1, t1, x_sin, t_sin, optimal_w1, m=9, ax=ax1)
plot(x2, t2, x_sin, t_sin, optimal_w2, m=9, ax=ax2)
plt.show()
```
## Regularization
$$
E(\mathbf{w}) = \dfrac{1}{2} \Vert y - V \cdot w \Vert^2 + \frac{\lambda}{2} \Vert w \Vert^2 \qquad \cdots (4)
$$
where, $\Vert \mathbf{w} \Vert^2 \equiv \mathbf{w}^T\mathbf{w}=w_0^2 + w_1^2 + \cdots w_M^2$
easy to get solution for this
$$
\begin{aligned}
\frac{\partial E(w)}{\partial w} &= V^Ty-V^TV\cdot w+\lambda w = 0 \\
w &= (V^TV- \lambda I_{(M+1)})^{-1}V^Ty
\end{aligned}
$$
when regularizer $\lambda \uparrow$, means that more regularization
```
def ridge_solution(x, t, m, alpha=0):
V = vandermonde_matrix(x, m)
return np.linalg.inv(np.dot(V.T, V) - alpha * np.eye(m+1)).dot(V.T).dot(t)
M=9
optimal_w1 = ridge_solution(x, t, m=M, alpha=1e-8)
optimal_w2 = ridge_solution(x, t, m=M, alpha=1.0)
# Plot predicted line
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(16, 6))
def plot_ridge(x, t, x_sin, t_sin, optimal_w, m, text, ax):
ax.plot(x_sin, t_sin, c="green", label="sin function")
ax.plot(x_sin, polynomial_function(x_sin, optimal_w, m), c="red", label=f"model M={m}")
ax.scatter(x, t)
ax.set_xlim((0, 1))
ax.set_ylim((-1.25, 1.25))
ax.set_xlabel('x', fontsize=16)
ax.set_ylabel('t', rotation=0, fontsize=16)
ax.legend()
ax.annotate(text, (0.6, 0.5), fontsize=14)
plot_ridge(x, t, x_sin, t_sin, optimal_w1, m=M, text='lambda = 1e-8', ax=ax1)
plot_ridge(x, t, x_sin, t_sin, optimal_w2, m=M, text='lambda = 1.0', ax=ax2)
plt.show()
print(f"coefficients at lambda=1e-8 is {optimal_w1.round(3)}")
print(f"coefficients at lambda=1.0 is {optimal_w2.round(3)}")
```
## see ridge effect
```
all_w = []
all_rms_train = []
all_rms_test = []
M = 9
for alpha in np.exp(np.arange(-28, -15)):
optimal_w = ridge_solution(x, t, m=M, alpha=alpha)
t_hat = polynomial_function(x, optimal_w, m=M)
t_hat_test = polynomial_function(x_test, optimal_w, m=M)
rms_train = get_rms_error(t_hat, t, N, m=M)
rms_test = get_rms_error(t_hat_test, t_test, N_test, m=M) # N_test = 100
print(f"lambda={alpha} | rms_train: {rms_train:.4f} rms_test: {rms_test:.4f}")
# Plot predicted line
# plt.plot(x_sin, t_sin, c="green", label="sin function")
# plt.plot(x_sin, polynomial_function(x_sin, optimal_w, m), c="red", label=f"model M={m}")
# plt.scatter(x, t)
# plt.xlim((0, 1))
# plt.ylim((-1.25, 1.25))
# plt.xlabel('x', fontsize=16)
# plt.ylabel('t', rotation=0, fontsize=16)
# plt.legend()
# plt.show()
all_w.append(optimal_w)
all_rms_train.append(rms_train)
all_rms_test.append(rms_test)
plt.scatter(np.arange(len(all_rms_train)), all_rms_train, facecolors='none', edgecolors='b')
plt.plot(np.arange(len(all_rms_train)), all_rms_train, c='b', label='Training')
plt.scatter(np.arange(len(all_rms_test)), all_rms_test, facecolors='none', edgecolors='r')
plt.plot(np.arange(len(all_rms_test)), all_rms_test, c='r', label='Test')
plt.legend()
plt.xticks(np.arange(len(all_rms_test)), np.arange(-28, -15))
plt.ylabel("root-mean-squared Error", fontsize=16)
plt.xlabel("np.log(lambda)", fontsize=16)
plt.show()
```
| github_jupyter |
## Initialization
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import math
import scipy.io
from scipy.special import expit
from math import *
from scipy import optimize
sns.set_style('whitegrid')
%matplotlib inline
```
## Loading Data
```
mat = scipy.io.loadmat('ex4data1.mat')
X = mat['X']
y = mat['y']
X = np.insert(X,0,1,axis= 1)
m,n = X.shape
input_layer_size = 400
hidden_layer_size = 25
num_labels = 10
_lambda = 1
```
## Function Section
```
#functions Sections
def magic_display(matrix = None):
if matrix is None:
# selecting 100 random rows of the X
rand_indces = np.random.permutation(m)[0:100]
X_dis = X[rand_indces]
else:
X_dis = matrix
if( len(X_dis.shape) > 1 ):
m_test,n_test = X_dis.shape
axis_bound = 1
else:
m_test = 1
n_test = X_dis.shape[0]
axis_bound = 0
# each number width , height in plot
example_width = int(round(sqrt(n_test)))
example_height = int(round( n_test / example_width ))
# number of numbers to show in plot
display_rows = floor(sqrt(m_test))
display_cols = ceil(m_test / display_rows )
# padding between numbers
pad = 2
# intilazation array for holding previos 100 random numbers
display_array = np.ones((
pad + display_rows * ( example_height + pad ),
pad + display_cols * ( example_width + pad )
))
count = 0;
for i in range(display_rows):
for j in range(display_cols):
if( count >= m_test ):
break
# max_val of each row in X_dis
max_val = np.max( X_dis[count : count+1], axis= axis_bound)
# Starting x,y point of numbers shape in array
ex_x_range = pad + ( i ) * ( example_height + pad )
ex_y_range = pad + ( j ) * ( example_width + pad )
if(m_test > 1):
ex_arr = X_dis[ count : count + 1 , 1:].reshape(example_height , example_width)
else:
ex_arr = X_dis[1:].reshape(example_height , example_width)
# Setting values
display_array[ ex_x_range : ex_x_range + example_height,
ex_y_range : ex_y_range + example_width ] = np.divide(ex_arr , max_val)
count += 1
# Plotting 100 random data
plt.figure(figsize=(12,8))
# Get rod of grid
plt.grid(False)
plt.imshow(display_array)
def hyp(matrix):
return expit(matrix)
def neural_cost_function(nn_params, input_layer_size, hidden_layer_size, num_labels, X, y, _lam):
# initialization some varibles
if(len(X.shape) > 1):
axis_bound = 1
else:
axis_bound = 0
# reshaping from one dimensional to 2d dimensional parameter vector
end_indx_theta1 = hidden_layer_size * ( input_layer_size + 1 )
Theta1 = np.reshape( nn_params[0 : end_indx_theta1 ],
( hidden_layer_size, input_layer_size + 1 ))
# reshaping from one dimensional to 2d dimensional parameter vector
Theta2 = np.reshape( nn_params[end_indx_theta1 : ],
( num_labels, hidden_layer_size + 1 ))
# Copmuting hidden level activation
z_2 = np.dot(X, Theta1.T )
hidden_activation = hyp( z_2 )
hidden_activation = np.insert( hidden_activation, 0, 1, axis=axis_bound )
# Copmuting output level activation
z_3 = np.dot(hidden_activation, Theta2.T)
out_activation = hyp(z_3)
# finding hypotesis matrix
h = out_activation
# Computing Log(sigmoid(x)) for all of the hypotesis elements
h1 = np.log(h)
# Computing Log( 1 - simgoid(x)) for all of the hypotesis elements
h2 = np.log(1 - h)
# Creating new matrix for y
new_y0 = ( y - 1 ).copy()
new_y1 = np.zeros(out_activation.shape)
new_y1[np.arange(0,out_activation.shape[0]),new_y0.T] = 1
# Computing Regularization Part Varibles
Theta1_pow2 = Theta1 * Theta1
Theta2_pow2 = Theta2 * Theta2
#Computing Cost of the hypotesis
J = ( -1 / m ) * sum(sum( new_y1 * h1 + (1 - new_y1) * h2)) + \
( _lam / ( 2 * m )) * ( sum(sum( Theta1_pow2 )) + sum(sum( Theta2_pow2 )) )
return J
def neural_gradient_function(nn_params, input_layer_size, hidden_layer_size, num_labels, X, y, _lam):
# initialization some varibles
if(len(X.shape) > 1):
axis_bound = 1
else:
axis_bound = 0
# Number of training examples
m = X.shape[0]
# reshaping from one dimensional to 2d dimensional parameter vector
end_indx_theta1 = hidden_layer_size * ( input_layer_size + 1 )
Theta1 = np.reshape( nn_params[0 : end_indx_theta1 ],
( hidden_layer_size, input_layer_size + 1 ))
# reshaping from one dimensional to 2d dimensional parameter vector
Theta2 = np.reshape( nn_params[end_indx_theta1 : ],
( num_labels, hidden_layer_size + 1 ))
# Defining Delta's
Delta1 = np.zeros(Theta1.shape)
Delta2 = np.zeros(Theta2.shape)
# Defining Theta_grad Matrixs
Theta1_grad = np.zeros(Theta1.shape)
Theta2_grad = np.zeros(Theta2.shape)
for i in range(m):
X_input = X[i : i + 1,:]
# Copmuting hidden level activation
z_2 = np.dot( X_input, Theta1.T )
hidden_activation = hyp( z_2 )
hidden_activation = np.insert( hidden_activation, 0, 1, axis=axis_bound )
# Copmuting output level activation
z_3 = np.dot( hidden_activation, Theta2.T )
out_activation = hyp( z_3 )
# finding hypotesis matrix
h = out_activation
# Creating new matrix for y
new_y0 = ( y - 1 ).copy()
new_y1 = np.zeros(out_activation.shape[1])
new_y1[new_y0[i]] = 1
# Computing erros
out_error = h - new_y1
z_2 = np.insert(z_2, 0, 1, axis=1)
hidden_error = np.dot( out_error , Theta2 ).T * sigmoid_gradient(z_2).T
hidden_error = hidden_error[ 1: ]
# Computing Delta
Delta1 = Delta1 + hidden_error * X_input
Delta2 = Delta2 + out_error.T * hidden_activation
Theta1_grad[:, 0:1 ] = ( 1 / m ) * ( Delta1[:, 0:1 ] )
Theta1_grad[:, 1: ] = ( 1 / m ) * ( Delta1[:, 1: ] ) + ( _lam / m ) * Theta1[:, 1: ]
Theta2_grad[:, 0:1 ] = ( 1 / m ) * ( Delta2[:, 0:1 ] )
Theta2_grad[:, 1: ] = ( 1 / m ) * ( Delta2[:, 1: ] ) + ( _lam / m ) * Theta2[:, 1: ]
# Converting Weigths to 1 Dimensional Matrix's
Theta1_grad_flat = np.array(Theta1_grad.flat)
Theta2_grad_flat = np.array(Theta2_grad.flat)
return np.concatenate((Theta1_grad_flat, Theta2_grad_flat)) * 1e-3
def sigmoid_gradient(matrix):
return hyp(matrix) * ( 1 - hyp(matrix) )
def checking_gradient(_lambda):
if(_lambda == None):
_lambda = 0
input_layer_size = 3
hidden_layer_size = 5
num_labels = 3
m = 5
Theta1 = debug_initialaize_weights(hidden_layer_size, input_layer_size)
Theta2 = debug_initialaize_weights(num_labels, hidden_layer_size)
X = debug_initialaize_weights(m, input_layer_size - 1)
y = 1 + np.mod(np.arange(0,m), num_labels)
# initialization some varibles
if(len(X.shape) > 1):
axis_bound = 1
else:
axis_bound = 0
# Inserting 1's column to matrix
X = np.insert( X, 0, 1, axis= axis_bound)
Theta1_flat = np.array(Theta1.flat)
Theta2_flat = np.array(Theta2.flat)
Theta = np.concatenate((Theta1_flat, Theta2_flat))
grad = neural_gradient_function(Theta, input_layer_size, hidden_layer_size, num_labels, X, y, _lambda)
numerical_grad = numerical_gradinet_function(Theta, input_layer_size, hidden_layer_size, num_labels, X, y, _lambda)
print(np.linalg.norm(numerical_grad - grad) / np.linalg.norm(numerical_grad + grad))
def numerical_gradinet_function(Theta, input_layer_size, hidden_layer_size, num_labels, X, y, _lambda):
new_grad = np.zeros(Theta.size)
p = np.zeros(Theta.size)
e = 1e-4
for i in range(Theta.size):
p[i] = e
j1 = neural_cost_function(Theta + p, input_layer_size, hidden_layer_size, num_labels, X, y, _lambda)
j2 = neural_cost_function(Theta - p, input_layer_size, hidden_layer_size, num_labels, X, y, _lambda)
new_grad[i] = (j1 - j2) / ( 2 * e )
p[i] = 0
return new_grad
def debug_initialaize_weights(output_layer, input_layer):
matrix = np.zeros((output_layer, input_layer + 1))
return np.sin(np.arange(1,matrix.size + 1)).reshape(matrix.shape) / 10
checking_gradient(3)
```
## Visualizing Data
```
magic_display()
```
## Feedforward Propagation Algorithm
```
# Loading Weights
weights = scipy.io.loadmat('ex4weights.mat')
Theta1 = weights['Theta1']
Theta2 = weights['Theta2']
Theta1.shape
Theta2.shape
# Converting Weigths to 1 Dimensional Matrix's
Theta1_flat = np.array(Theta1.flat)
Theta2_flat = np.array(Theta2.flat)
# Creating New 1d Matrix for holding all of the weights
Theta = np.concatenate((Theta1_flat, Theta2_flat))
neural_cost_function(Theta, input_layer_size, hidden_layer_size, num_labels, X, y, 3)
```
| github_jupyter |
# X-TEC-s (label smoothing) analysis of Sr$_3$Rh$_4$Sn$_{13}$ XRD data
In this tutorial, we analyze the XRD data of Sr<sub>3</sub>Rh<sub>4</sub>Sn<sub>13</sub>, a quasi-skutterudite family which shows charge density wave ordering below a quantum critical point.
This tutorial takes single-crystal x-ray diffraction data in the data file `srn0_XTEC.nxs` available for download at https://dx.doi.org/10.18126/iidy-30e7
## System requirements
The tutorial should run on any python version. We recommend: python 3.8 and Jupyter Notebook 6.3.0.
## Installing XTEC
Released versions of *XTEC* can be installed using either
```
$ pip install xtec
```
or by cloning the Git repository and installing from source:
```
$ git clone https://github.com/KimGroup/XTEC
```
## XTEC codes
XTEC codes are available in GitHub repository: https://github.com/KimGroup/XTEC
## Required python libraries
```
import glob
import os
import time
import matplotlib.pyplot as plt
import numpy as np
from nexusformat.nexus import (nxgetmemory, nxload, nxsetmemory, NXdata,
NXentry, NXfield)
```
## Required XTEC modules
The following modules are to be imported for the XTEC analysis:
```
from xtec.Preprocessing import Mask_Zeros # to remove zero intensity points from data
from xtec.Preprocessing import Threshold_Background # for thresholding background intensity
from xtec.GMM import GMM # for XTEC GMM clustering
from xtec.GMM import GMM_kernels
```
# X-TEC flowchart
The flowchart below gives the steps we follow in this tutorial.

# Step A. Import the XRD Data
The data loading can be customized to the type of file and structure of data stored.
Ultimately, one requires a temperature series of intensity at (Qh, Qk, Ql) points of the 3D reciprocal lattice (or equivalently in 2D reciprocal lattice) as an array with
```
Data. shape = (num_Temperature, num_Ql, num_Qk, num_Qh) for 3D data (equivalently for 2D data).
```
#### From https://dx.doi.org/10.18126/iidy-30e7, download the data file ```srn0_XTEC.nxs```
```
data_filename = 'srn0_XTEC.nxs' # path to your datafile: srn0_XTEC.nxs
data_file = nxload(data_filename, 'r')
data = data_file['entry/data'][:, 0.0:0.2, -10.0:10.0, -15.0:15.0]
I = data.nxsignal.nxvalue # intensities, shape= (num_Temperature, num_Ql, num_Qk, num_Qh)
Qh = data['Qh'].nxvalue # Qh in r.l.u
Qk = data['Qk'].nxvalue # Qk in r.l.u
Ql = data['Ql'].nxvalue # Ql in r.l.u
Temp = data['Te'].nxvalue # Temperatures
```
# Step B. Threshold background intensity
The data is first put through a preprocessing to remove low intensity background noise. Assumes that the peaks/relevant x-ray features are sparse so that most momenta have predominantely background intensity contributions.
Cutoff automatically estimated by minimizing the Kullback-Leibler (KL) distance between the distribution of $\log[\overline{I_q(T)}]$ and a Gaussian distribution with (high intensity) sliding cutoff. See supplementary materials of Venderley et al., https://arxiv.org/abs/2008.03275.
See preprocessing.py for comments on the code
```
start_time = time.time()
masked = Mask_Zeros(I) # removes data with zero intensity.
threshold = Threshold_Background(masked) # remove low intensity background
print("Thresholding Run Time:",time.time() - start_time)
```
The results from thresholding are stored as
```
1. threshold.data_thresholded : contains thresholded data.
Shape=(num_T, num_thresholded_data)
2. threshold.ind_thresholded : contains hkl indices of the thresholded data.
Shape=(num_data_thresholded, 3) for 3D or (num_data_thresholded, 2) for 2D.
3. threshold.thresholded : stores whether the corresponding pixel is removed (=0) or not (=1).
Shape=(num_l, num_k, num_h) or shape=(num_k, num_h)
```
```
Data_thresh = threshold.data_thresholded # shape=(num_temperatures, num_thresholded data)
Data_ind = threshold.ind_thresholded # contains hkl indices of the thresholded data.
print('num of original data pixels=', np.prod(I.shape[1:]))
print('num of thresholded data pixels=', Data_thresh.shape[1])
```
Let us see the thresholding results. Below is the distribution of intensities $(\log[\overline{I_q(T)}])$ and the truncation point (red dot) determined by thresholding. Only intensities above the truncation are retained.
```
figsize_=(10,5)
threshold.plot_cutoff(figsize_)
```
Note the non Gaussian features in the distribution just above the cutoff intensity (red dot). Those are the useful data we are after.
All data points that have $\log[\overline{I_q(T)}]$ above estimated cutoff are retained. The plot below shows those remaining points in the hk plane as grey clusters
```
figsize_=(8,8)
axis_ind = 0 # Select 0 for plotting l plane, 1 for k plane, 2 for h plane
slice_value = 0.0 # Value of the axis orthogonal to the selected plane
slice_ind = data.nxaxes[axis_ind+1].index(slice_value) # index of the selected axis (h/k/l plane) to plot
threshold.plot_thresholding_2D_slice(figsize_, slice_ind, axis_ind)
# threshold.plot_thresholding_2D_slice(figsize_, slice_ind_, axis_) : plots the 2D image
# with points whose intensity distribution is above the cutoff marked as grey.
# if data is 3D, then set the slice ind and axis_ (default=None) to select the index of the 2D slice along axis_ =0,1 or 2.
ax = plt.gca()
ax.get_images()[0].set_extent((Qh[0], Qh[-1], Qk[0], Qk[-1]))
plt.xlabel('H')
plt.ylabel('K')
plt.title(f'L={slice_value}')
```
# Step. C Rescale intensity
To reduce the large range in the scale of intensities and for efficient feature selection of the temperature trajectory, we provide two options to rescale intensities $I_q(T)$:
#### Rescaling by mean:
subtracting and dividing their mean over temperature $\mu_{\vec{q}_i}=d_T^{-1}\sum_j I_{\vec{q}_i}(T_j)$ for each $q$,
\begin{eqnarray}
\tilde{I}_{\vec{q}_i}(T_j)=\dfrac{I_{\vec{q}_i}(T_j)}{\mu_{\vec{q}_i}}-1,\label{eq:mean_rescaling}
\end{eqnarray}
#### Z-scoring the data:
subtracting the mean (over T) and then dividing by standard deviation (over T)
\begin{eqnarray}
\tilde{I}_{\vec{q}_i}(T_j)=\dfrac{I_{\vec{q}_i}(T_j)-\mu_{\vec q_i}}{\sigma_{\vec{q}_i}},
\end{eqnarray}
where $\sigma_{\vec{q}_i}=\sqrt{d_T^{-1}\sum_j \left(I_{\vec{q}_i}(T_j)-\mu_{\vec q_i}\right)^2}$ is the standard deviation.
```
Rescaled_data=threshold.Rescale_mean(Data_thresh) # rescale by mean
#Rescaled_data=threshold.Rescale_zscore(data) # rescale by z-score
print('num of thresholded (and rescaled) data = ', Rescaled_data.shape[1] )
print('num T = ', Rescaled_data.shape[0])
```
## Optional step: remove data with low variance in temperature (for faster results)
To see faster results, and when large number of data post thresholding is made of noise, this step removes the noisy data by eliminating low variance trajectories. We have to however set the cutoff manually
```
#cutoff=0.3
#threshold.Get_High_Variance(Rescaled_data,cutoff) # get (rescaled) data with std_dev (in T) > cutoff
#Rescaled_data=threshold.data_high_std_dev # The high variance data, shape=(num_T, num_high_var_data)
#Data_ind = threshold.ind_high_std_dev # (hkl) indices of high var data, shape=(num_high_var_data,3)
```
The downside is the setting of manual cutoff for this step.
###### Recommended course of action: Cluster without any cutoff on smaller datasets. Apply cutoff on larger datasets for faster results
# Step D: set number of clusters
Start with an initial guess (say 3 or 4), and run till step F (visualization). Revisit this step to change the num of clusters to arrive at the optimal choice.
```
num_clusters = 4 # change this to 3, 4, 5 etc and see.
```
# Step E.2: XTEC-s (label smoothing)
We now apply clustering with label smoothing. Here temperature series of intensity at each $q$ point is clustered independently.
### Parameters for label smoothing (Build Markov matrix)
```
# Smoothing through peak averaging is better for larger data sets
L_scale = .05 # change L_scale to control the length scale across which the labels are smoothed
# for periodic kernel (enforce label smoothing in neighbouring pixels in and across Brillouin zone)
kernel_type = 'periodic'
unit_cell_shape = np.array([20,20,20]) # The size of unit cell (integer lengths in unit of number of pixels)
Markov_matrix = GMM_kernels.Build_Markov_Matrix(Data_ind, L_scale,kernel_type,unit_cell_shape)
Data_for_GMM = Rescaled_data.transpose() # data should be arranged as (num_data,num_T) for GMM
start_time = time.time()
label_smoothing_flag=True
clusterGMM = GMM(Data_for_GMM,num_clusters) # set the data and number of clusters for GMM
clusterGMM.RunEM(label_smoothing_flag,Markov_matrix) # RunEM with Markov matrix
print("GMM Run Time:",time.time() - start_time)
cluster_assignments=clusterGMM.cluster_assignments
print('num of trajectories in each cluster = ', clusterGMM.num_per_cluster)
```
Clustering results are stored in clusterGMM as
```
clusterGMM.cluster_assignments : cluster assignment k in range(num_clusters), of each sample. Shape=(num_data)
clusterGMM.num_per_cluster : number of trajectories in each cluster.
clusterGMM.cluster[k].mean: cluster mean trajectory (dim=num_Temperature) of the k-th cluster
clusterGMM.cluster[k].cov : cluster covariance [dim=num_Temperature] of the k-th cluster
```
# Step F. Visualization of clustered results.
Let us plot the cluster mean of the rescaled data (lines), with 1 standard deviation shaded, for the clusters color coded as red blue green violet etc.
```
clusterGMM.Plot_Cluster_Results_traj(Temp) # Plotting for only less than 7 clusters currently enabled.
plt.xlabel('T(K)', size=18)
plt.ylabel('$\widetilde{I}_q(T)$', size=18)
plt.title('Cluster mean and variance \n (rescaled) intensity trajectory ');
```
The order parameter like behavior of one of the cluster is apparent from here, with a transition temperature $T_c\approx 130 K$ This cluster corresponds to the CDW peak. To see that, let us plot and see which pixels in the $(h,k)$ plane belong to the red/blue/green/violet cluster.
```
figsize_=(8,8)
axis_ind=0 # Select 0 for plotting l plane, 1 for k plane, 2 for h plane
slice_value = 0.0 # Value of the axis orthogonal to the selected plane
slice_ind = data.nxaxes[axis_ind+1].index(slice_value) # index of the selected axis (h/k/l plane) to plot
Data_for_GMM_ind= Data_ind #(hkl) indices of the data that was clustered
clusterGMM.Plot_Cluster_kspace_2D_slice(threshold,figsize_,Data_for_GMM_ind,slice_ind,axis_ind)
ax = plt.gca()
ax.get_images()[0].set_extent((Qh[0], Qh[-1], Qk[0], Qk[-1]))
plt.xlabel("H", size=22)
plt.ylabel("K", size=22)
plt.title(f'L={slice_value}')
```
```
clusterGMM.Plot_Cluster_kspace_2D_slice(threshold,figsize_,data_ind,slice_ind,axis_ind)
```
Plots the 2D image slice, with each (thresholded) pixel color coded by the clustering label, or colored grey if not clustered.
The input parameters are:
```
threshold : class Threshold_Background from Preprocessing
figsize_ : size of image. If None, will not plot the image
data_ind : (h,k,l) or (h,k) indices of the data. shape=(num_data,2) or (num_data,3)
slice_ind : if 3D, the h/k/l index of the selected axis along which to take the data slice.
axis_ind : selected axis_ of slice_ind. 0 for l, 1 for k and 2 for h axis
```
Output attributes for ploting with imshow(),
```
clusterGMM.plot_image : 2D image to be plotted
clusterGMM.plot_cmap : color map for the clustering, matching the same color scheme of the cluster trajectories
clusterGMM.plot_norm : the norm to be used in imshow() so that cmap matches the cluster assignments
```
```
# a zoom in image of the above figure..
plt.figure(figsize=(10,10))
plt.imshow(clusterGMM.plot_image[200:300,0:100],origin='lower',cmap=clusterGMM.plot_cmap,norm=clusterGMM.plot_norm,extent=[Qh[0], Qh[100], Qk[200],Qk[300]])
plt.xlabel("Q$_h$",size=22)
plt.ylabel("Q$_k$",size=22)
plt.title(str('Ql='+str(Ql[slice_ind])),size=22)
```
Notice that CDW peaks at $q_{CDW} = (H+0.5,K+0.5,0)$ have overwhelmingly the same color matching the color of the order parameter like trajectory.
### Remove the dominating diffuse cluster to focus on the peaks .
# Step G: Identify the distinct cluster
There are other features such as the thermal diffuse scattering surrounding the Bragg peaks (trajectories increasing with $T$), and the noise at the centre of the reciprocal space. To focus on the more interesting features (the CDW and Bragg peaks), let us remove these noise like features and cluster the remaining trajectories again.
### Select the clusters you want to remove
```
Bad_Clusters=[0] # Select the list clusters to be removed: 0 for Red, 1 for blue, 2 for green, 3 for violet, 4 for yellow, 5 for orange, 6 for pink clusters...
# Note that the color assignments can change with each run.
# This block removes the data belonging to the bad clusters from the first clustering
cluster_assignments = clusterGMM.cluster_assignments
BadC_mask = ~np.isin(cluster_assignments, Bad_Clusters)
Good_data = Data_thresh[:,BadC_mask]
Good_rescaled_data = Rescaled_data[:,BadC_mask] # the data belonging to the good clusters
Good_ind = Data_ind[BadC_mask] # (h,k,l) indices of the good data
```
### Redo the GMM clustering with only the interesting clusters (steps E and F again)
```
unit_cell_shape = np.array([20,20,20])
L_scale = .05 # change L_scale to control the length scale for smoothing the labels
kernel_type = 'periodic'
data_inds=Good_ind
Markov_matrix = GMM_kernels.Build_Markov_Matrix(data_inds, L_scale,kernel_type,unit_cell_shape)
num_clusters_2 = 2 # set the number of new clusters for the second step clustering
Data_for_GMM_2= Good_rescaled_data.transpose()
clusterGMM_2 = GMM(Data_for_GMM_2,num_clusters_2)
smoothing_iterations=1 # number of times Markov matrix is introduced between E and M step
clusterGMM_2.RunEM(True, Markov_matrix,smoothing_iterations) # RunEM with Markov matrix
print('num of trajectories in each cluster:', clusterGMM_2.num_per_cluster)
```
Let us plot the cluster trajectories
```
clusterGMM_2.Plot_Cluster_Results_traj(Temp)
plt.xlabel('T(K)', size=18)
plt.ylabel('$\widetilde{I}_q(T)$', size=18)
plt.title(' ')
```
Now we clearly see the order parameter like cluster. Let us plot and see which pixels in the (ℎ,𝑘) plane belong to the new clustering assignments.
```
plt.figure(figsize=(10,10))
axis_ind = 0 # Select 0 for plotting l plane, 1 for k plane, 2 for h plane
slice_value = 0.0 # Value of the axis orthogonal to the selected plane
slice_ind = data.nxaxes[axis_ind+1].index(slice_value) # index of the selected axis (h/k/l plane) to plot
Data_for_GMM_ind=Good_ind
clusterGMM_2.Plot_Cluster_kspace_2D_slice(threshold,None,Data_for_GMM_ind,slice_ind,axis_ind)
plt.imshow(clusterGMM_2.plot_image[200:300,0:100],origin='lower',cmap=clusterGMM_2.plot_cmap,norm=clusterGMM_2.plot_norm,extent=[Qh[0], Qh[100], Qk[200],Qk[300]])
plt.xlabel("H",size=22)
plt.ylabel("K",size=22)
plt.title(f'L={slice_value}',size=22)
```
The bad clusters not selected for the second round clustering are now labeled as grey pixels.
#### The order parameter trajectory is reflected clearly at the CDW peaks (no cross talk in the cluster colors between neighboring pixels)
# Final step: Physics!
Store the clustered results and identify the physics in them
### Plot the cluster averaged intensities to see the order parameter
```
plt.figure()
color_list = ['red', 'blue', 'green', 'purple', 'yellow', 'orange', 'pink', 'black', 'grey', 'cyan']
yc_plot={}
for i in range(clusterGMM_2.cluster_num):
cluster_mask_i = (clusterGMM_2.cluster_assignments == i)
yc_plot[i]=np.mean(Good_data[:,cluster_mask_i],axis=1).flatten()
plt.plot(Temp,yc_plot[i],color=color_list[i], lw=2);
plt.yscale("log")
plt.xlabel('T')
plt.ylabel('I (cluster avged)')
```
### Store the final clustering results
```
import pickle
obj = {}
obj['Temp'] = Temp # 1 sigma var of the cluster trajectory, shape=(num_cluster,num_T)
obj['Data'] = Good_data.transpose() # shape=(num_data, num_T)
obj['Data_ind'] = Good_ind # (hkl) indices of data, shape= (num_data,3)
obj['cluster_assignments'] = clusterGMM_2.cluster_assignments # cluster number = 0/1/2 for each data, shape=(num_data)
obj['Num_clusters'] = clusterGMM_2.cluster_num
obj['cluster_mean'] = clusterGMM_2.means # mean trajectory of clusters, shape=(num_cluster, num_T)
obj['cluster_var'] = clusterGMM_2.covs # 1 sigma var of the cluster trajectory, shape=(num_cluster,num_T)
pickle.dump( obj, open( "csrs_0x0_XTEC_s.p", "wb" ) )
```
# https://github.com/KimGroup/X-TEC
## Harnessing Interpretable and Unsupervised Machine Learning to Address Big Data from Modern X-ray Diffraction
### https://arxiv.org/abs/2008.03275
###### Jordan Venderley, Michael Matty, Krishnanand Mallaya, Matthew Krogstad, Jacob Ruff, Geoff Pleiss, Varsha Kishore, David Mandrus, Daniel Phelan, Lekh Poudel, Andrew Gordon Wilson, Kilian Weinberger, Puspa Upreti, M. R. Norman, Stephan Rosenkranz, Raymond Osborn, Eun-Ah Kim
### Tutorial by Krishnanand Mallayya.
### For questions about X-TEC code and tutorial contact kmm537@cornell.edu (Krishnanand)
| github_jupyter |
```
%matplotlib inline
sample_submission = pd.read_csv('C:\\dss\\team_project2\\walmart\\sample_submission.csv')
sample_submission
test = pd.read_csv('C:\\dss\\team_project2\\walmart\\test.csv')
test.tail()
train = pd.read_csv('C:\\dss\\team_project2\\walmart\\train.csv')
train.tail()
df = train.copy()
```
# Delete column
```
del df["Upc"]
del df["FinelineNumber"]
```
# Get dummy
```
weekday = pd.get_dummies(df['Weekday'])
weekday = weekday.sort_index()
weekday
DepartmentDescription = pd.get_dummies(df['DepartmentDescription'])
DepartmentDescription = DepartmentDescription.sort_index()
DepartmentDescription
del df["Weekday"]
del df["DepartmentDescription"]
df = pd.concat([df, weekday, DepartmentDescription],axis=1)
df
```
# Make a data
```
gdf1 = df[['TripType', "VisitNumber", "Friday", "Monday", "Saturday", "Thursday", "Tuesday", "Wednesday"]].groupby("VisitNumber").mean()
gdf2 = df[['VisitNumber', 'ScanCount', 'SHOES', 'DAIRY', 'GROCERY DRY GOODS', 'DSD GROCERY', 'BAKERY',
'IMPULSE MERCHANDISE', 'PERSONAL CARE', 'FABRICS AND CRAFTS',
'BOYS WEAR', 'PRE PACKED DELI', 'MENS WEAR', 'CELEBRATION',
'HOUSEHOLD CHEMICALS/SUPP', 'FROZEN FOODS', 'PRODUCE',
'HOME MANAGEMENT', 'PHARMACY OTC', 'CANDY, TOBACCO, COOKIES',
'HOUSEHOLD PAPER GOODS', 'SPORTING GOODS', 'HARDWARE',
'COMM BREAD', 'HOME DECOR', 'COOK AND DINE', 'SERVICE DELI',
'PETS AND SUPPLIES', 'MEAT - FRESH & FROZEN', 'FINANCIAL SERVICES',
'SEAFOOD', 'WIRELESS', 'MEDIA AND GAMING', 'PAINT AND ACCESSORIES',
'LADIESWEAR', 'SLEEPWEAR/FOUNDATIONS', 'BEAUTY',
'BRAS & SHAPEWEAR', 'FURNITURE', 'LAWN AND GARDEN', 'AUTOMOTIVE',
'BATH AND SHOWER', 'ELECTRONICS', 'OFFICE SUPPLIES',
'INFANT CONSUMABLE HARDLINES', 'GIRLS WEAR, 4-6X AND 7-14',
'INFANT APPAREL', 'LIQUOR,WINE,BEER', 'BOOKS AND MAGAZINES',
'HORTICULTURE AND ACCESS', 'JEWELRY AND SUNGLASSES', 'ACCESSORIES',
'LADIES SOCKS', 'BEDDING', 'TOYS', 'OPTICAL - FRAMES',
'PHARMACY RX', 'CAMERAS AND SUPPLIES', 'MENSWEAR',
'PLAYERS AND ELECTRONICS', 'PLUS AND MATERNITY',
'SWIMWEAR/OUTERWEAR', 'CONCEPT STORES', 'SHEER HOSIERY',
'LARGE HOUSEHOLD GOODS', 'OPTICAL - LENSES', '1-HR PHOTO',
'OTHER DEPARTMENTS', 'SEASONAL']].groupby('VisitNumber').sum()
gdf = pd.concat([gdf1, gdf2], axis=1)
X = gdf.copy()
X.drop(columns = 'TripType', inplace=True)
X.tail()
y = gdf1['TripType'].copy()
y
```
# modeling
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2)
import xgboost
model_xgb = xgboost.XGBClassifier(n_estimators=100, max_depth=11)
model_xgb.fit(X_train, y_train)
pred = model_xgb.predict(X_test)
pred
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
print(classification_report(y_test, pred))
# plot the important features #
fig, ax = plt.subplots(figsize=(12,25))
xgboost.plot_importance(model_xgb, ax=ax)
plt.show()
# test[test["DepartmentDescription"] == "HEALTH AND BEAUTY AIDS"]
train[train["DepartmentDescription"] == "HEALTH AND BEAUTY AIDS"]
```
# submission
```
sub = test.copy()
del sub["Upc"]
del sub["FinelineNumber"]
test_weekday = pd.get_dummies(sub['Weekday'])
test_weekday = test_weekday.sort_index()
test_DepartmentDescription = pd.get_dummies(sub['DepartmentDescription'])
test_DepartmentDescription = test_DepartmentDescription.sort_index()
del sub["Weekday"]
del sub["DepartmentDescription"]
test_submission = pd.concat([sub, test_weekday, test_DepartmentDescription],axis=1)
test_submission
test_submission1 = test_submission[["VisitNumber", "Friday", "Monday", "Saturday", "Thursday", "Tuesday", "Wednesday"]].groupby("VisitNumber").mean()
test_submission1
test_submission2 = test_submission[['VisitNumber', 'ScanCount', 'SHOES', 'DAIRY', 'GROCERY DRY GOODS', 'DSD GROCERY', 'BAKERY',
'IMPULSE MERCHANDISE', 'PERSONAL CARE', 'FABRICS AND CRAFTS',
'BOYS WEAR', 'PRE PACKED DELI', 'MENS WEAR', 'CELEBRATION',
'HOUSEHOLD CHEMICALS/SUPP', 'FROZEN FOODS', 'PRODUCE',
'HOME MANAGEMENT', 'PHARMACY OTC', 'CANDY, TOBACCO, COOKIES',
'HOUSEHOLD PAPER GOODS', 'SPORTING GOODS', 'HARDWARE',
'COMM BREAD', 'HOME DECOR', 'COOK AND DINE', 'SERVICE DELI',
'PETS AND SUPPLIES', 'MEAT - FRESH & FROZEN', 'FINANCIAL SERVICES',
'SEAFOOD', 'WIRELESS', 'MEDIA AND GAMING', 'PAINT AND ACCESSORIES',
'LADIESWEAR', 'SLEEPWEAR/FOUNDATIONS', 'BEAUTY',
'BRAS & SHAPEWEAR', 'FURNITURE', 'LAWN AND GARDEN', 'AUTOMOTIVE',
'BATH AND SHOWER', 'ELECTRONICS', 'OFFICE SUPPLIES',
'INFANT CONSUMABLE HARDLINES', 'GIRLS WEAR, 4-6X AND 7-14',
'INFANT APPAREL', 'LIQUOR,WINE,BEER', 'BOOKS AND MAGAZINES',
'HORTICULTURE AND ACCESS', 'JEWELRY AND SUNGLASSES', 'ACCESSORIES',
'LADIES SOCKS', 'BEDDING', 'TOYS', 'OPTICAL - FRAMES',
'PHARMACY RX', 'CAMERAS AND SUPPLIES', 'MENSWEAR',
'PLAYERS AND ELECTRONICS', 'PLUS AND MATERNITY',
'SWIMWEAR/OUTERWEAR', 'CONCEPT STORES', 'SHEER HOSIERY',
'LARGE HOUSEHOLD GOODS', 'OPTICAL - LENSES', '1-HR PHOTO',
'OTHER DEPARTMENTS', 'SEASONAL']].groupby('VisitNumber').sum()
test_submission2
testt = pd.concat([test_submission1, test_submission2], axis=1)
testt
sub_pred = model_xgb.predict(testt)
sub_pred
submi = pd.DataFrame(pd.get_dummies(sub_pred))
submi
submi.columns
a = pd.DataFrame(submi, columns=[ 3, 4, 5, 6, 7, 8, 9, 12, 14, 15, 18, 19, 20, 21,
22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34,
35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 999])
a1 = a.fillna(0).astype(int)
VisitNumber = pd.DataFrame(testt.index)
my_submission = pd.concat([VisitNumber, a1], axis=1,)
my_submission.columns=['VisitNumber', 'TripType_3', 'TripType_4', 'TripType_5', 'TripType_6',
'TripType_7', 'TripType_8', 'TripType_9', 'TripType_12', 'TripType_14',
'TripType_15', 'TripType_18', 'TripType_19', 'TripType_20',
'TripType_21', 'TripType_22', 'TripType_23', 'TripType_24',
'TripType_25', 'TripType_26', 'TripType_27', 'TripType_28',
'TripType_29', 'TripType_30', 'TripType_31', 'TripType_32',
'TripType_33', 'TripType_34', 'TripType_35', 'TripType_36',
'TripType_37', 'TripType_38', 'TripType_39', 'TripType_40',
'TripType_41', 'TripType_42', 'TripType_43', 'TripType_44',
'TripType_999']
my_submission
my_submission.to_csv("C:\\dss\\team_project2\\walmart\\my_submission",index=False)
sample_submission
```
| github_jupyter |
```
import nsepython
from portfoliotools.screener.stock_detail import StockDetail
import pandas as pd
from pandas.plotting import register_matplotlib_converters
import warnings
import seaborn as sns
import matplotlib.pyplot as plt
from plotly.subplots import make_subplots
import plotly.graph_objects as go
import numpy as np
from datetime import datetime
from statsmodels.tsa.arima_model import ARIMA
from sklearn.metrics import mean_squared_error
import scipy.stats as st
warnings.filterwarnings("ignore")
register_matplotlib_converters()
%matplotlib inline
sns.set()
pd.options.display.max_columns = None
pd.options.display.max_rows = None
ticker = 'RELIANCE'
focus_period = 1000
lookback_period = 30
return_period = 1 # Daily returns
stockObj = StockDetail(ticker, period = focus_period)
prices = stockObj.calculate_return(return_period) # return_period Returns
prices['Risk'] = prices['Return'].rolling( window = lookback_period).std()*np.sqrt(360/return_period) # return_period Risk
prices['Return'] = prices['Return']*(360/return_period)
fig = make_subplots(rows = 2, cols = 1, shared_xaxes= True, vertical_spacing = 0.08,
column_widths = [15], row_heights = [2,2])
# Add Returns
fig.add_trace(
go.Scatter(x=prices.index, y=prices['Return']/360, name="Return", line = {'color':'purple'}), row = 1, col =1,
)
fig.add_trace(
go.Scatter(x=prices.index, y=[prices['Return'].mean()/360]*len(prices), name="Avg Return", line = {'color':'skyblue'}), row = 1, col =1,
)
# Add Risk
fig.add_trace(
go.Scatter(x=prices.index, y=prices['Risk'], name="Risk", line = {'color':'red'}), row = 2, col =1,
)
fig.add_trace(
go.Scatter(x=prices.index, y=[prices['Risk'].mean()]*len(prices), name="Avg Risk", line = {'color':'pink'}), row = 2, col =1,
)
fig['layout']['yaxis1'].update(title='Return')
fig['layout']['yaxis2'].update(title='Risk')
fig.show()
print("Avg Return : " + str(round(prices['Return'].mean(),3)))
print("Avg Risk : " + str(round(prices['Risk'].mean(),3)))
print("Latest Risk : " + str(round(prices['Risk'].values[-1],3)))
fnoList = nsepython.fnolist()
details = nsepython.nse_fno(ticker)
option_chain = nsepython.option_chain(ticker)
fnoPrice = [z['metadata'] for z in details['stocks']]
fnoPrice = pd.DataFrame(fnoPrice)
optionPrice = fnoPrice[fnoPrice['instrumentType'] == 'Stock Options']
futurePrice = fnoPrice[fnoPrice['instrumentType'] == 'Stock Futures']
strikePrices = optionPrice['strikePrice'].unique()
info = details['info']
info['underlying'] = details['underlyingValue']
result = []
for data in option_chain['records']['data']:
pe = data.get('PE', None)
ce = data.get('CE', None)
if pe is not None:
result.append({
'strikePrice': data.get('strikePrice',0),
'expiryDate': data.get('expiryDate', ''),
'optionType': 'Put',
'closePrice': pe.get('lastPrice', 0),
'totalBuyQuantity': pe.get('totalBuyQuantity', 0),
'totalSellQuantity' : pe.get('totalSellQuantity', 0),
'openInterest' : pe.get('openInterest', 0),
'pchangeinOpenInterest' : pe.get('pchangeinOpenInterest', 0),
'identifier' : pe.get('identifier', ''),
'numberOfContractsTraded' : pe.get('totalTradedVolume', 0),
'impliedVolatility' : pe.get('impliedVolatility', 0),
'pChange' : pe.get('pChange', 0),
'underlyingValue' : pe.get('underlyingValue', 0),
})
if ce is not None:
result.append({
'strikePrice': data.get('strikePrice',0),
'expiryDate': data.get('expiryDate', ''),
'optionType': 'Call',
'closePrice': ce.get('lastPrice', 0),
'totalBuyQuantity': ce.get('totalBuyQuantity', 0),
'totalSellQuantity' : ce.get('totalSellQuantity', 0),
'openInterest' : ce.get('openInterest', 0),
'pchangeinOpenInterest' : ce.get('pchangeinOpenInterest', 0),
'identifier' : ce.get('identifier', ''),
'numberOfContractsTraded' : ce.get('totalTradedVolume', 0),
'impliedVolatility' : ce.get('impliedVolatility', 0),
'pChange' : ce.get('pChange', 0),
'underlyingValue' : ce.get('underlyingValue', 0),
})
option_chain = pd.DataFrame(result)
option_chain['expiryDate'] = option_chain['expiryDate'].apply(lambda x: datetime.strptime(x, '%d-%b-%Y').strftime('%Y-%m-%d'))
expiryDates = option_chain['expiryDate'].unique()
option_chain
def smape_kun(y_true, y_pred):
return np.mean((np.abs(y_pred - y_true) * 200/ (np.abs(y_pred) + np.abs(y_true))))
def predict(df):
result = {
'MSE' : np.nan,
'SMAPE KUN' : np.nan,
'Pred Value' : np.nan,
'SD' : np.nan,
'Pred Low 50%' : np.nan,
'Pred High 50%' :np.nan,
'Model':None
}
train_data, test_data = df[0:int(len(df)*0.8)], df[int(len(df)*0.8):]
train, test = train_data['data'].values, test_data['data'].values
history = [x for x in train]
predictions = list()
p = 5
d = 0
q = 1
for t in range(len(test)):
model = ARIMA(history, order=(p,q,d))
model_fit = model.fit(disp=0)
output = model_fit.forecast()
yhat = output[0]
predictions.append(yhat)
obs = test[t]
history.append(obs)
error = mean_squared_error(test, predictions)
result['MSE'] = np.round(error, 3)
error2 = smape_kun(test, predictions)
result['SMAPE KUN'] = np.round(error2, 3)
model = ARIMA(history, order=(p,q,d))
model_fit = model.fit(disp=0)
result['Model'] = model_fit
output = model_fit.forecast(alpha =0.5)
result['Pred Value'] = np.round(output[0][0],2)
result['SD'] = np.round(output[1][0],2)
result['Pred Low 50%'] = np.round(output[2][0][0],2)
result['Pred High 50%']= np.round(output[2][0][1],2)
return result
# Predict Price Range
data = prices[['Adj Close']]
data.rename(columns={'Adj Close' : 'Close'}, inplace=True)
# Expiry Dates
daysToExpiry = [(datetime.strptime(d, '%Y-%m-%d') - datetime.now()).days for d in expiryDates]
daysToExpiry = [z - round(z/7)*2 for z in daysToExpiry]
forecast = {}
i=0
for days in daysToExpiry:
data['Low_'+ str(days)] = data['Close'].rolling( window = days).min()
data['High_'+ str(days)] = data['Close'].rolling( window = days).max()
#data['Return_'+ str(days)] = (data['Close']/data['Close'].shift(days)-1)*100
data['High_'+ str(days)] = ((data['High_'+ str(days)]/data['Close'])-1)*100
data['Low_'+ str(days)] = ((data['Low_'+ str(days)]/data['Close'])-1)*100
df_High = pd.DataFrame(data = data['High_'+ str(days)].values, columns = ['data'])
df_Low = pd.DataFrame(data = data['Low_'+ str(days)].values, columns = ['data'])
df_High.dropna(inplace=True)
df_Low.dropna(inplace=True)
temp = {}
temp['High'] = predict(df_High)
temp['Low'] = predict(df_Low)
temp['DaysToExpiry'] = days
forecast[expiryDates[i]] = temp
i+=1
# Straddle Details
def straddleCost(data):
try:
callPrice = list(data[data['optionType'] == 'Call']['closePrice'].values)[0]
putPrice = list(data[data['optionType'] == 'Put']['closePrice'].values)[0]
return callPrice + putPrice
except:
return 0
def callPrice(data):
try:
callPrice = list(data[data['optionType'] == 'Call']['closePrice'].values)[0]
return callPrice
except:
return 0
def putPrice(data):
try:
putPrice = list(data[data['optionType'] == 'Put']['closePrice'].values)[0]
return putPrice
except:
return 0
def straddleBreakEven(data, direction = 'up', displayPercent = False):
try:
cost = straddleCost(data)
strike = list(data['strikePrice'].values)[0]
spot = list(data['underlyingValue'].values)[0]
if direction == 'up':
price = strike + cost
else:
price = strike - cost
if displayPercent:
if spot != 0:
return ((price - spot)*100 / spot)
else:
np.nan
else:
return price
except:
return 0
def groupImpliedVolatility(data, optionType = 'Call'):
try:
return list(data[data['optionType'] == optionType]['impliedVolatility'].values)[0]
except:
return 0
# Append price ranges
option_chain['predHighMean'] = option_chain['expiryDate'].apply(lambda x: forecast[x]['High']['Pred Value'])
option_chain['predLowMean'] = option_chain['expiryDate'].apply(lambda x: forecast[x]['Low']['Pred Value'])
option_chain['predHighSD'] = option_chain['expiryDate'].apply(lambda x: forecast[x]['High']['SD'])
option_chain['predLowSD'] = option_chain['expiryDate'].apply(lambda x: forecast[x]['Low']['SD'])
option_chain['daysToExpiry'] = option_chain['expiryDate'].apply(lambda x: forecast[x]['DaysToExpiry'])
straddleDetails = option_chain.groupby(['expiryDate', 'strikePrice']).agg({'numberOfContractsTraded' : sum,
'underlyingValue': max,
'predHighMean': max,
'predLowMean':max,
'predHighSD':max,
'predLowSD':max,
'daysToExpiry':max})
straddleDetails['call_price'] = option_chain.groupby(['expiryDate', 'strikePrice']).apply(callPrice)
straddleDetails['put_price'] = option_chain.groupby(['expiryDate', 'strikePrice']).apply(putPrice)
straddleDetails['cost'] = option_chain.groupby(['expiryDate', 'strikePrice']).apply(straddleCost)
straddleDetails['breakeven_up'] = option_chain.groupby(['expiryDate', 'strikePrice']).apply(straddleBreakEven,'up')
straddleDetails['breakeven_down'] = option_chain.groupby(['expiryDate', 'strikePrice']).apply(straddleBreakEven,'down')
straddleDetails['breakeven_up_per'] = option_chain.groupby(['expiryDate', 'strikePrice']).apply(straddleBreakEven,'up', True)
straddleDetails['breakeven_down_per'] = option_chain.groupby(['expiryDate', 'strikePrice']).apply(straddleBreakEven,'down', True)
straddleDetails['iv_pe'] = option_chain.groupby(['expiryDate', 'strikePrice']).apply(groupImpliedVolatility,'Put')
straddleDetails['iv_ce'] = option_chain.groupby(['expiryDate', 'strikePrice']).apply(groupImpliedVolatility,'Call')
straddleDetails = straddleDetails[straddleDetails['numberOfContractsTraded'] > 0]
straddleDetails = straddleDetails[straddleDetails['iv_ce'] > 0]
straddleDetails = straddleDetails[straddleDetails['iv_pe'] > 0]
straddleDetails['strikePrice'] = straddleDetails.index.get_level_values(1)
#straddleDetails['probUpStd'] = straddleDetails[['strikePrice', 'call_price', 'underlyingValue', 'iv_ce', 'daysToExpiry']].apply(lambda x: round(100-100*st.norm.cdf(np.log((x['strikePrice'] + x['call_price'])/x['underlyingValue'])/(x['iv_ce']*.01 * np.sqrt(x['daysToExpiry']/250))),2), axis=1)
straddleDetails['probUpStd'] = straddleDetails[['breakeven_up', 'underlyingValue', 'iv_ce', 'daysToExpiry']].apply(lambda x: round(100-100*st.norm.cdf(np.log(x['breakeven_up']/x['underlyingValue'])/(x['iv_ce']*.01 * np.sqrt(x['daysToExpiry']/250))),2), axis=1)
straddleDetails['probUpPredict'] = straddleDetails[['predHighMean', 'predHighSD','breakeven_up_per']].apply(lambda x: round(100-st.norm.cdf((x['breakeven_up_per'] - x['predHighMean'])/x['predHighSD'])*100,2), axis=1)
#straddleDetails['probDownStd'] = straddleDetails[['strikePrice', 'put_price', 'underlyingValue', 'iv_pe', 'daysToExpiry']].apply(lambda x: round(100*st.norm.cdf(np.log((x['strikePrice'] - x['put_price'])/x['underlyingValue'])/(x['iv_pe']*.01 * np.sqrt(x['daysToExpiry']/250))),2), axis=1)
straddleDetails['probDownStd'] = straddleDetails[['breakeven_down', 'underlyingValue', 'iv_pe', 'daysToExpiry']].apply(lambda x: round(100*st.norm.cdf(np.log(x['breakeven_down']/x['underlyingValue'])/(x['iv_pe']*.01 * np.sqrt(x['daysToExpiry']/250))),2), axis=1)
straddleDetails['probDownPredict'] = straddleDetails[['predLowMean', 'predLowSD','breakeven_down_per']].apply(lambda x: round(st.norm.cdf((x['breakeven_down_per'] - x['predLowMean'])/x['predLowSD'])*100,2), axis=1)
straddleDetails['probStraddle'] = (straddleDetails['probUpPredict'] + straddleDetails['probDownPredict'])/2
straddleDetails['probStraddleStd'] = straddleDetails['probUpStd'] + straddleDetails['probDownStd']
straddleDetails = straddleDetails[straddleDetails.columns.drop(['predHighMean', 'predHighSD','predLowMean', 'predLowSD', 'strikePrice'])]
straddleDetails
expiryDates.sort()
rows = round(len(expiryDates)/2)
fig = make_subplots(rows = rows, cols = 2, shared_xaxes= True, vertical_spacing = 0.08,
column_widths = [15,15], row_heights = [2]*rows)
i=1
j=1
for date in expiryDates:
data = straddleDetails[straddleDetails.index.isin([date], level=0)]
# Add iv_ce
fig.add_trace(
go.Scatter(x=data.index.get_level_values(1), y=data['iv_ce'], name="IV CE", line = {'color':'green'}), row = i, col =j,
)
# Add iv_pe
fig.add_trace(
go.Scatter(x=data.index.get_level_values(1), y=data['iv_pe'], name="IV PE", line = {'color':'red'}), row = i, col =j,
)
fig['layout']['yaxis' + str(2*(i-1)+j)].update(title=date)
if j == 2:
i+=1
j = 1 if j==2 else 2
fig.show()
straddleDetails.to_clipboard()
straddleDetails.reset_index()
```
| github_jupyter |
# Dataframes Handling
**Inhalt:** Ein bessere Gefühl dafür erhalten, wie Dataframes aufgebaut sind und wie man damit umgeht
**Nötige Skills:** Erste Schritte mit Pandas
**Lernziele:**
- Verschiedene Konstruktur-Methoden für Dataframes kennenlernen
- Wie Series und Dataframes zusammenspielen
- Der Index und verschiedene Arten, ihn zu referenzieren
- Spalten und Zeilen hinzufügen und löschen
- Aufmerksamkeit schärfen für Dinge wie `inplace=True`
## Vorbereitung
```
import pandas as pd
```
## Dataframes konstruieren
Man kann Daten nicht nur aus Dateien in ein Dataframe laden, sondern auch aus anderen Datenstrukturen. Es gibt gefühlt 1000 Arten, ein Dataframe zu konstruieren.
Einige Beispiele dazu finden sich hier: https://pandas.pydata.org/pandas-docs/stable/dsintro.html
Eine Referenz zur Funktion `DataFrame()` findet sich hier: https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html. An diesem Ort sind übrigens auch alle Methoden verzeichnet, die auf DataFrame-Objekte angewandt werden können.
Wir schauen uns hier eine kleine Auswahl davon an.
### Dataframe aus Dictionaries erstellen
Daten können aus verschiedenen Quellen kommen. Aus Dateien, oder zB auch von gescrapten Webseiten.
Nehmen wir mal an, wir haben Daten von einer Social Media Seite zusammengestellt, und sie liegen nun Form von Dictionaries vor.
Wie können wir damit in Pandas arbeiten?
```
person1 = {'Name': 'Peter', 'Groesse': 180, 'Alter': 30}
person2 = {'Name': 'Paul', 'Groesse': 176, 'Alter': 44}
person3 = {'Name': 'Maria', 'Groesse': 165, 'Alter': 25}
person1
```
Schritt 1: Wir machen daraus eine "Liste von Dictionaries":
```
l = [person1, person2, person3]
l
```
Schritt 2: Wir machen aus der Liste ein Dataframe:
```
df = pd.DataFrame(l)
```
Die einzelnen Einträge in der Liste sind nun die *Zeilen* in der Tabelle:
```
df
```
### Dataframe aus Listen erstellen
Vielleicht sind wir aber auch auf anderem Weg an diese Daten gekommen.
Und wir haben nun eine Liste von Personen sowie von deren Attributen, statt ein Dictionary.
```
namen = ['Peter', 'Paul', 'Maria']
groesse = [180, 176, 165]
alter = [30, 44, 25]
```
Schritt 1: Wir machen ein "Dictionary von Listen" daraus.
```
d = {'Name': namen, 'Groesse': groesse, 'Alter': alter}
d
```
Schritt 2: Wir machen aus dem Dictionary ein Dataframe
```
df = pd.DataFrame(d)
```
Die einzelnen Einträge im Dictionary sind nun die *Spalten* in unserer Tabelle:
```
df
```
**Note:** Wichtig ist nicht unbedingt, diese Techniken auswendig zu können. Sondern einfach zu wissen: `DataFrame()` ist eine sehr vielseitige Funktion, man kann fast alles darin reinfüttern.
Falls das Resultat nicht so herauskommt, wie gewünscht, ist die Chance gross, dass sich eine andere Variante findet, wie man aus irgendwelchen Datenstrukturen ein Dataframe erhält. Es gibt zB auch:
- pd.DataFrame.from_dict()
- pd.DataFrame.from_records()
- pd.DataFrame.from_items()
Im Zweifelsfall: Dokumentation konsultieren, Beispiele anschauen, ausprobieren!
Dataframes kann man auch aus Serien generieren.
### Serien
Was sind schon wieder Serien? Ach ja: Es ist das, was herauskommt, wenn man einzelne Spalten eines Dataframes herauszieht.
```
df['Name']
```
Serien sind ganz einfach gesagt eine Art von indexierten Listen.
Wir kann sie ähnlich wie Dataframes auch from Scratch konstruieren – mittels `pd.Series()`
```
s = pd.Series(['Peter', 'Paul', 'Maria'])
s
```
Der Index muss übrigens nicht zwingend aus einer Zahlenreihe bestehen.
```
s = pd.Series(['Peter', 'Paul', 'Maria'], index=['a', 'b', 'c'])
s
```
Series sind ähnlich wie Listen, aber auch ähnlich wie Dictionaries. Man kann basierend auf dem Index zB einzelne Elemente aufrufen
```
s['a']
```
... oder auch andere, lustige Auswahloperationen machen
```
s['b':]
```
Eine Reihe von Serien zu einem Dataframe zusammenzufügen, funktioniert sehr ähnlich wie mit Listen.
### Dataframe aus Series erstellen
Nehmen wir also an, unsere Daten liegen in Form von Listen vor:
```
s_name = pd.Series(['Peter', 'Paul', 'Maria'])
s_groesse= pd.Series([180, 176, 165])
s_alter= pd.Series([30, 44, 25])
s_name
s_groesse
s_alter
```
Um die drei Serien zu einem Dataframe zu machen, gehen wir ähnlich vor wie zuvor.
Schirtt 1: Wir basteln einen Dictionary aus den Serien:
```
d = {"Name": s_name, "Groesse": s_groesse, "Alter": s_alter}
d
```
Schritt 2: Wir kontstruieren aus dem Dictionary ein Dataframe
```
pd.DataFrame(d)
```
### Dataframes aus mühsamen Strukturen erstellen
Nehmen wir mal an, unsere Daten liegen in einer etwas merkwürdigen Form vor (merkwürdig zB deshalb, weil man normalerweise den Inhalt eines Datenfeldes ("Peter") nicht als Variablennamen verwenden würde):
```
Peter = {'Groesse': 180, 'Alter': 30}
Paul = {'Groesse': 176, 'Alter': 44}
Maria = {'Groesse': 165, 'Alter': 25}
Peter
```
Eine Variante, um ein Dataframe zu erstellen, könnte so funktionieren:
```
df = pd.DataFrame([Peter, Paul, Maria], index=['Peter', 'Paul', 'Maria'])
df
```
Wir haben nun eine etwas anders aufgebaute Tabelle. Die Index-Spalte besteht nicht mehr aus Zahlen, sondern aus den Werten in einer bestimmten Spalte.
Wir schauen uns nun die Index-Spalte etwas genauer an.
### Mit dem Index arbeiten
Wir haben bereits kennengelernt, wie man die Werte einer einzelnen Spalte herausziehen kann:
```
df['Alter']
```
Mit einzelnen Zeilen funktioniert es ähnlich, allerdings brauchen wir dazu noch eine spezielle Syntax: `df.loc[]`
```
df.loc['Peter']
```
`df.loc['Peter']` funktioniert, weil es eine Zeile gibt, die den Indexwert "Peter" hat.
Falls nötig, können wir eine bestimmte Zeile aber auch jederzeit mit einem Integer-Zahlenwert referenzieren.
Dafür gibt es die Syntax `df.iloc[]` - die Zählung startet bei null, nicht bei eins.
```
df.iloc[0]
```
Zwei Spalten anzeigen
```
df.iloc[0:2]
```
Wollen wir nur bestimmte Spaltenwerte anzeigen, funktioniert das so:
```
df.loc['Peter', 'Alter']
```
Diese Syntax dient übrigens auch dazu, Werte zu überschreiben:
```
df.loc['Peter', 'Alter'] = 60
```
Peter ist nun nicht mehr 30, sondern 60 Jahre alt.
```
df
```
**Achtung:** Das hier funktioniert übrigens nicht: (Warum nicht?)
```
df[df['Alter'] == 60]['Alter'] = 30
df
```
Im allgemeinen ist es allerdings unschön, bestimmte Datenfelder als Index zu verwenden - könnte ja sein, dass zwei Leute Peter heissen, das gäbe an irgendeinem Punkt ziemlich sicher einen Fehler.
Um das zu ändern, gibt es eine einfache Funktion: `reset_index()`
Nun haben wir einen neuen Index (**fett**), und die Namen wurden in eine separate Spalte kopiert.
```
df = df.reset_index()
df
```
Das Problem ist, die Spalte mit den Namen heisst jetzt "index"...
Um das zu ändern, benutzen wir `df.rename()` und geben einen Dictionary mit auf den Weg, der die zu ändernden Spaltennamen beinhaltet.
```
df.rename(columns={'index': 'Name'})
```
**Achtung:** Wie die meisten Funktionen bei Pandas, ändert `df.rename()` nicht das Dataframe selbst, sondern spuckt ein neues Dataframe aus.
Um die Änderung zu speichern, haben wir zwei Optionen:
a) Wir speichern das retournierte Dataframe unter demselben oder unter einem neuen Namen (`df2`) ab:
```
df_2 = df.rename(columns={'index': 'Name'})
df_2
```
b) Wir verwenden das Argument `inplace=True`, das so viel bedeutet wie: "mach das gleich an Ort und Stelle".
```
df.rename(columns={'index': 'Name'}, inplace=True)
df
```
`inplace=True` funktioniert übrigens auch mit einigen anderen Funktionen, z.B. auch mit `df.reset_index()`
```
df.reset_index(inplace=True)
df
```
Oops! Nun haben wir bereits eine Spalte zu viel drin, die wir gar nicht wollen.
Macht nix, wir werfen sie einfach wieder raus. Und zwar mit `pop()`
```
df.pop('index')
df
```
**Note:** Anders als zuvor hat `df.pop()` gleich an Ort und Stelle die Spalte rausgeworfen, statt ein neues Dataframe auszuspucken. Das Default-Verhalten ist über Pandas hinweg nicht ganz konsistent: Manchmal gilt `inplace=True` automatisch, manchmal nicht.
Wir können übrigens auch einzelne Zeilen rauswerfen, basierend auf dem Index der Zeile:
```
df.drop([2], inplace=True)
df
```
Falls wir den Index nicht kennen, suchen wir ihn, und zwar mit dem Attribut `.index`
```
df.drop(df[df['Name'] == 'Paul'].index, inplace=True)
df
```
Zu viel gelöscht? Egal, fügen wir die Zeile einfach wieder hinzu, und zwar mit `df.append()`
```
Paul = pd.Series({'Alter': 44, 'Groesse': 176, 'Name': 'Paul'})
Paul
df = df.append(Paul, ignore_index=True)
df
```
Und zu guter letzt, falls wir doch wieder die Namen als Index wollen: `df.set_index()`
```
df.set_index('Name', inplace=True)
df
```
**Achtung:** Was nicht funktioniert, ist diese Zeile hier: (warum?)
```
#df2 = df.set_index('Alter', inplace=True)
```
**Note:** Die Index-Spalte hat jetzt einen ganz bestimmten Namen: "Name". Wir können diesen Namen auch löschen, und zwar mit `df.rename_axis()`:
```
df.rename_axis(None, inplace=True)
df
```
## Schluss
Es ist nicht wichtig, dass ihr jede dieser Funktionen auswendig könnt und genau wisst, wie man sie anwenden muss, ob inplace=True oder nicht.
Wichtig ist aber:
- Seid euch bewusst, dass es für fast alles, was man mit einem Dataframe anstellen will, in Pandas eine Funktion gibt
- Falls ihr den Namen der gesuchten Funktion nicht kennt: Googelt einfach. Die Chance ist gross, dass ihr schnell fündig werdet
- In der Dokumentation (zB über Google auffindbar: "Pandas set index") steht immer, welche Argumente eine Funktion nimmt
- Sonst einfach ausprobieren. Falls irgendwas komplett falsch läuft, einfach den Kernel nochmals starten und alles nochmal ausführen
| github_jupyter |
# Shopee-Product-Matching

1. If you want to learn more about this amazing competition hosted by [Shopee](https://www.kaggle.com/c/shopee-product-matching), Please visit following [Shopee EDA Image AutoEncoder](https://www.kaggle.com/code/chiragtagadiya/shopee-basic-autoencoder).
2. This Notebook contains EDA and Image AutoEncoder solution.
```
%config Completer.use_jedi = False
```
# Import Packages
```
import sys
sys.path.append('../input/timmmaster')
import timm
import math
import os
import numpy as np
import cv2
import pandas as pd
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import StratifiedKFold
import timm
import torch
from torch import nn
from torch.utils.data import Dataset, DataLoader
import torch.nn.functional as F
import albumentations
from albumentations.pytorch.transforms import ToTensorV2
from torch.optim import lr_scheduler
from tqdm import tqdm
import matplotlib.pyplot as plt
from sklearn import metrics
from datetime import date
from sklearn.metrics import f1_score, accuracy_score
```
# Configuration Options
```
TRAIN_DIR = '../input/shopee-product-matching/train_images'
TEST_DIR = '../input/shopee-product-matching/test_images'
TRAIN_CSV = '../input/crossvalidationfolds/folds.csv'
MODEL_PATH = './'
class CFG:
seed = 123
img_size = 512
classes = 11014
fc_dim = 512
epochs = 15
batch_size = 32
num_workers = 3
model_name = 'tf_efficientnet_b4'
device = 'cuda' if torch.cuda.is_available() else 'cpu'
scheduler_params = {
"lr_start": 1e-3,
"lr_max": 1e-5 * batch_size,
"lr_min": 1e-6,
"lr_ramp_ep": 5,
"lr_sus_ep": 0,
"lr_decay": 0.8,
}
model_path='../input/21-mar-lr-large/2022-03-20_softmax_512x512_tf_efficientnet_b4.pt'
isTraining=False
```
# Solution Approach
* In this competition it is given that,if two or more images have **same label group** then they are **similar products.**
* Basically we can use this information to transfer the business problem into **multi class classification** problem.
* From Image EDA, I found out that we have **11014** different classes, and dataset is **not balanced dataset**
* If you see below plot, we can clearly see that there are **hardly 1000 data points having more than 10 products per label.*
* In this notebook I used **Weighted Sampler technique used in pytorch for handling imbalanced classification problem**
```
train_df=pd.read_csv('../input/shopee-product-matching/train.csv')
labelGroups = train_df.label_group.value_counts()
# print(labelGroups)
plt.figure(figsize=(15,5))
plt.plot(np.arange(len(labelGroups)), labelGroups.values)
plt.xlabel("Index for unique label_group_item", size=12)
plt.ylabel("Number of product data for label ", size=12)
plt.title("label vs label frequency", size=15)
plt.show()
```
# Create Custom DataSet
```
class ShopeeDataset(Dataset):
def __init__(self, df,root_dir, isTraining=False, transform=None):
self.df = df
self.transform = transform
self.root_dir = root_dir
def __len__(self):
return len(self.df)
def __getitem__(self, idx):
# get row at index idx
# print("idx",idx)
row = self.df.iloc[idx]
# print(row)
label = row.label_group
image_path = os.path.join(self.root_dir, row.image)
# read image convert to RGB and apply augmentation
image = cv2.imread(image_path)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
if self.transform:
aug = self.transform(image=image)
image = aug['image']
return image, torch.tensor(label).long()
```
# Create Data Augmentation For training and validation Data
```
def getAugmentation(IMG_SIZE, isTraining=False):
if isTraining:
return albumentations.Compose([
albumentations.Resize(IMG_SIZE, IMG_SIZE, always_apply=True),
albumentations.HorizontalFlip(p=0.5),
albumentations.VerticalFlip(p=0.5),
albumentations.Rotate(limit=120, p=0.75),
albumentations.RandomBrightness(limit=(0.09, 0.6), p=0.5),
albumentations.Normalize(
mean = [0.485, 0.456, 0.406],
std = [0.229, 0.224, 0.225]
),
ToTensorV2(p=1.0)
])
else:
return albumentations.Compose([
albumentations.Resize(IMG_SIZE, IMG_SIZE, always_apply=True),
albumentations.Normalize(
mean = [0.485, 0.456, 0.406],
std = [0.229, 0.224, 0.225]
),
ToTensorV2(p=1.0)
])
```
# Build Model
```
class ShopeeLabelGroupClassfier(nn.Module):
def __init__(self,
model_name='tf_efficientnet_b0',
loss_fn='softmax',
classes = CFG.classes,
fc_dim = CFG.fc_dim,
pretrained=True,
use_fc=True,
isTraining=False
):
super(ShopeeLabelGroupClassfier,self).__init__()
# create bottlenack backbone network from pretrained model
self.backbone = timm.create_model(model_name, pretrained=pretrained)
in_features = self.backbone.classifier.in_features
# we will put FC layers over backbone to classfy images based on label groups
self.backbone.classifier = nn.Identity()
self.backbone.global_pool = nn.Identity()
self.pooling = nn.AdaptiveAvgPool2d(1)
self.use_fc = use_fc
self.loss_fn =loss_fn
# build top fc layers
if self.use_fc:
self.dropout = nn.Dropout(0.2)
self.fc = nn.Linear(in_features,fc_dim )
self.bn = nn.BatchNorm1d(fc_dim)
in_features = fc_dim
self.loss_fn = loss_fn
if self.loss_fn=='softmax':
self.final = nn.Linear(in_features, CFG.classes)
def forward(self, image, label):
features = self.get_features(image)
if self.loss_fn=='softmax':
logits = self.final(features)
return logits
def get_features(self,inp):
batch_dim = inp.shape[0]
inp = self.backbone(inp)
inp = self.pooling(inp).view(batch_dim, -1)
if self.use_fc:
inp = self.dropout(inp)
inp = self.fc(inp)
inp = self.bn(inp)
return inp
# shoppe_label_classfier = ShopeeLabelGroupClassfier()
```
# Training Single Epoch
```
def training_one_epoch(epoch_num,model, dataloader,optimizer, scheduler, device, loss_criteria):
avgloss = 0.0
# put model in traning model
model.train()
tq = tqdm(enumerate(dataloader), total=len(dataloader))
for idx, data in tq:
batch_size = data[0].shape[0]
images = data[0]
targets = data[1]
# zero out gradient
optimizer.zero_grad()
# put input and target to device
images = images.to(device)
targets = targets.to(device)
# pass input to the model
output = model(images,targets)
# get loss
loss = loss_criteria(output,targets)
# backpropogation
loss.backward()
# update learning rate step
optimizer.step()
# avg loss
avgloss += loss.item()
tq.set_postfix({'loss' : '%.6f' %float(avgloss/(idx+1)), 'LR' : optimizer.param_groups[0]['lr']})
# lr scheduler step after each epoch
scheduler.step()
return avgloss / len(dataloader)
```
# Validating Single Epoch
```
def validation_one_epoch(model, dataloader, epoch, device, loss_criteria):
avgloss = 0.0
# put model in traning model
model.eval()
tq = tqdm(enumerate(dataloader), desc = "Training Epoch { }" + str(epoch+1))
y_true=[]
y_pred=[]
with torch.no_grad():
for idx, data in tq:
batch_size = data[0].shape[0]
images = data[0]
targets = data[1]
images = images.to(device)
targets = targets.to(device)
output = model(images,targets)
predicted_label=torch.argmax(output,1)
y_true.extend(targets.detach().cpu().numpy())
y_pred.extend(predicted_label.detach().cpu().numpy())
loss = loss_criteria(output,targets)
avgloss += loss.item()
tq.set_postfix({'validation loss' : '%.6f' %float(avgloss/(idx+1))})
f1_score_metric = f1_score(y_true, y_pred, average='micro')
tq.set_postfix({'validation f1 score' : '%.6f' %float(f1_score_metric)})
return avgloss / len(dataloader),f1_score_metric
```
## Helper Function for Handling class imbalanced data
```
import numpy as np
def get_class_weights(data):
weight_dict=dict()
# Format of row : PostingId, Image, ImageHash, Title, LabelGroup
# LabelGroup index is 4 and it is representating class information
for row in data.values:
weight_dict[row[4]]=0
# Word dictionary keys will be label and value will be frequency of label in dataset
for row in data.values:
weight_dict[row[4]]+=1
# for each data point get label count data
class_sample_count= np.array([weight_dict[row[4]] for row in data.values])
# each data point weight will be inverse of frequency
weight = 1. / class_sample_count
weight=torch.from_numpy(weight)
return weight
```
# Training Loop
```
def run_training():
data = pd.read_csv('../input/crossvalidationfolds/folds.csv')
# label encoding
labelencoder= LabelEncoder()
data['label_group_original']=data['label_group']
data['label_group'] = labelencoder.fit_transform(data['label_group'])
#data['weights'] = data['label_group'].map(1/data['label_group'].value_counts())
# create training_data and validation data initially not using k fold
train_data = data[data['fold']!=0]
# get weights for classes
samples_weight=get_class_weights(train_data)
print("samples_weight", len(samples_weight))
validation_data = data[data['fold']==0]
# training augmentation
train_aug = getAugmentation(CFG.img_size,isTraining=True )
validation_aug = getAugmentation(CFG.img_size, isTraining=False)
# create custom train and validation dataset
trainset = ShopeeDataset(train_data, TRAIN_DIR, isTraining=True, transform = train_aug)
validset = ShopeeDataset(validation_data, TRAIN_DIR, isTraining=False, transform = validation_aug)
print(len(data), len(samples_weight))
print(len(trainset))
# create data sampler
sampler = torch.utils.data.sampler.WeightedRandomSampler(samples_weight, num_samples=len(samples_weight))
# create custom training and validation data loader num_workers=CFG.num_workers,
train_dataloader = DataLoader(trainset, batch_size=CFG.batch_size,
drop_last=True,pin_memory=True, sampler=sampler)
validation_dataloader = DataLoader(validset, batch_size=CFG.batch_size,
drop_last=True,pin_memory=True)
# define loss function
loss_criteria = nn.CrossEntropyLoss()
loss_criteria.to(CFG.device)
# define model
model = ShopeeLabelGroupClassfier()
model.to(CFG.device)
# define optimzer
optimizer = torch.optim.Adam(model.parameters(),lr= CFG.scheduler_params['lr_start'])
# learning rate scheudler
scheduler = lr_scheduler.CosineAnnealingWarmRestarts(optimizer, T_0=7, T_mult=1, eta_min=1e-6, last_epoch=-1)
history = {'train_loss':[],'validation_loss':[]}
for epoch in range(CFG.epochs):
# get current epoch training loss
avg_train_loss = training_one_epoch(epoch_num = epoch,
model = model,
dataloader = train_dataloader,
optimizer = optimizer,
scheduler = scheduler,
device = CFG.device,
loss_criteria = loss_criteria)
# get current epoch validation loss
avg_validation_loss = validation_one_epoch(model = model,
dataloader = validation_dataloader,
epoch = epoch,
device = CFG.device,
loss_criteria = loss_criteria)
history['train_loss'].append(avg_train_loss)
history['validation_loss'].append(avg_validation_loss)
# save model
torch.save(model.state_dict(), MODEL_PATH + str(date.today()) +'_softmax_512x512_{}.pt'.format(CFG.model_name))
torch.save({
'epoch': epoch,
'model_state_dict': model.state_dict(),
'optimizer': optimizer.state_dict(),
# 'scheduler': lr_scheduler.state_dict()
},
MODEL_PATH + str(date.today()) +'_softmax_512x512_{}_checkpoints.pt'.format(CFG.model_name)
)
return model, history
history=None
if CFG.isTraining:
model, history = run_training()
```
# Plot Training and Validation Loss and Accuracy
```
if CFG.isTraining:
epoch_lst = [ i+1 for i in range(15)]
plt.plot(epoch_lst,history['train_loss'])
plt.xlabel("Epoch number")
plt.ylabel('Training Loss')
plt.title('Training Loss SoftMax Loss Function')
plt.show()
if CFG.isTraining:
plt.plot(epoch_lst,history['validation_loss'])
plt.xlabel("Epoch number")
plt.ylabel('Validation Loss')
plt.title('Validation Loss SoftMax Loss Function')
plt.show()
```
# Prediction
```
def prediction(model):
data = pd.read_csv('../input/crossvalidationfolds/folds.csv')
# label encoding
labelencoder= LabelEncoder()
data['label_group'] = labelencoder.fit_transform(data['label_group'])
# Prepare Validation data
validation_data = data[data['fold']==0]
validation_aug = getAugmentation(CFG.img_size,isTraining=False)
validset = ShopeeDataset(validation_data, TRAIN_DIR, isTraining=False, transform = validation_aug)
test_data_loader = torch.utils.data.DataLoader(validset,batch_size=CFG.batch_size)
# put model in evalution mode
model.eval()
tq = tqdm(enumerate(test_data_loader))
y_true=[]
y_pred=[]
with torch.no_grad():
for idx, data in tq:
images = data[0]
targets = data[1]
images = images.to(CFG.device)
targets = targets.to(CFG.device)
y_true.extend(targets.detach().cpu().numpy())
output = model(images,targets)
outputs=torch.argmax(output,1)
y_pred.extend(outputs.detach().cpu().numpy())
f1_score_metric = f1_score(y_true, y_pred, average='micro')
return f1_score_metric
if not CFG.isTraining:
model = ShopeeLabelGroupClassfier(pretrained=False).to(CFG.device)
model.load_state_dict(torch.load(CFG.model_path))
f1=prediction(model)
print("F1 score {}".format(f1))
```
| github_jupyter |
# Convolutional Layer
In this notebook, we visualize four filtered outputs (a.k.a. activation maps) of a convolutional layer.
In this example, *we* are defining four filters that are applied to an input image by initializing the **weights** of a convolutional layer, but a trained CNN will learn the values of these weights.
<img src='notebook_ims/conv_layer.gif' height=60% width=60% />
### Import the image
```
import cv2
import matplotlib.pyplot as plt
%matplotlib inline
# TODO: Feel free to try out your own images here by changing img_path
# to a file path to another image on your computer!
img_path = 'data/udacity_sdc.png'
# load color image
bgr_img = cv2.imread(img_path)
# convert to grayscale
gray_img = cv2.cvtColor(bgr_img, cv2.COLOR_BGR2GRAY)
# normalize, rescale entries to lie in [0,1]
gray_img = gray_img.astype("float32")/255
# plot image
plt.imshow(gray_img, cmap='gray')
plt.show()
```
### Define and visualize the filters
```
import numpy as np
## TODO: Feel free to modify the numbers here, to try out another filter!
filter_vals = np.array([[-1, -1, 1, 1], [-1, -1, 1, 1], [-1, -1, 1, 1], [-1, -1, 1, 1]])
print('Filter shape: ', filter_vals.shape)
# Defining four different filters,
# all of which are linear combinations of the `filter_vals` defined above
# define four filters
filter_1 = filter_vals
filter_2 = -filter_1
filter_3 = filter_1.T
filter_4 = -filter_3
filters = np.array([filter_1, filter_2, filter_3, filter_4])
# For an example, print out the values of filter 1
print('Filter 1: \n', filter_1)
# visualize all four filters
fig = plt.figure(figsize=(10, 5))
for i in range(4):
ax = fig.add_subplot(1, 4, i+1, xticks=[], yticks=[])
ax.imshow(filters[i], cmap='gray')
ax.set_title('Filter %s' % str(i+1))
width, height = filters[i].shape
for x in range(width):
for y in range(height):
ax.annotate(str(filters[i][x][y]), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if filters[i][x][y]<0 else 'black')
```
## Define a convolutional layer
The various layers that make up any neural network are documented, [here](http://pytorch.org/docs/stable/nn.html). For a convolutional neural network, we'll start by defining a:
* Convolutional layer
Initialize a single convolutional layer so that it contains all your created filters. Note that you are not training this network; you are initializing the weights in a convolutional layer so that you can visualize what happens after a forward pass through this network!
#### `__init__` and `forward`
To define a neural network in PyTorch, you define the layers of a model in the function `__init__` and define the forward behavior of a network that applyies those initialized layers to an input (`x`) in the function `forward`. In PyTorch we convert all inputs into the Tensor datatype, which is similar to a list data type in Python.
Below, I define the structure of a class called `Net` that has a convolutional layer that can contain four 3x3 grayscale filters.
```
import torch
import torch.nn as nn
import torch.nn.functional as F
# define a neural network with a single convolutional layer with four filters
class Net(nn.Module):
def __init__(self, weight):
super(Net, self).__init__()
# initializes the weights of the convolutional layer to be the weights of the 4 defined filters
k_height, k_width = weight.shape[2:]
# assumes there are 4 grayscale filters
self.conv = nn.Conv2d(1, 4, kernel_size=(k_height, k_width), bias=False)
self.conv.weight = torch.nn.Parameter(weight)
def forward(self, x):
# calculates the output of a convolutional layer
# pre- and post-activation
conv_x = self.conv(x)
activated_x = F.relu(conv_x)
# returns both layers
return conv_x, activated_x
# instantiate the model and set the weights
weight = torch.from_numpy(filters).unsqueeze(1).type(torch.FloatTensor)
model = Net(weight)
# print out the layer in the network
print(model)
```
### Visualize the output of each filter
First, we'll define a helper function, `viz_layer` that takes in a specific layer and number of filters (optional argument), and displays the output of that layer once an image has been passed through.
```
# helper function for visualizing the output of a given layer
# default number of filters is 4
def viz_layer(layer, n_filters= 4):
fig = plt.figure(figsize=(20, 20))
for i in range(n_filters):
ax = fig.add_subplot(1, n_filters, i+1, xticks=[], yticks=[])
# grab layer outputs
ax.imshow(np.squeeze(layer[0,i].data.numpy()), cmap='gray')
ax.set_title('Output %s' % str(i+1))
```
Let's look at the output of a convolutional layer, before and after a ReLu activation function is applied.
```
# plot original image
plt.imshow(gray_img, cmap='gray')
# visualize all filters
fig = plt.figure(figsize=(12, 6))
fig.subplots_adjust(left=0, right=1.5, bottom=0.8, top=1, hspace=0.05, wspace=0.05)
for i in range(4):
ax = fig.add_subplot(1, 4, i+1, xticks=[], yticks=[])
ax.imshow(filters[i], cmap='gray')
ax.set_title('Filter %s' % str(i+1))
# convert the image into an input Tensor
gray_img_tensor = torch.from_numpy(gray_img).unsqueeze(0).unsqueeze(1)
# get the convolutional layer (pre and post activation)
conv_layer, activated_layer = model(gray_img_tensor)
# visualize the output of a conv layer
viz_layer(conv_layer)
```
#### ReLu activation
In this model, we've used an activation function that scales the output of the convolutional layer. We've chose a ReLu function to do this, and this function simply turns all negative pixel values in 0's (black). See the equation pictured below for input pixel values, `x`.
<img src='notebook_ims/relu_ex.png' height=50% width=50% />
```
# after a ReLu is applied
# visualize the output of an activated conv layer
viz_layer(activated_layer)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.