code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
# Formatação de gráficos com *matplotlib*
Vamos começar refazendo os gráficos que fizemos anteriormente com o método **plot** dos *DataFrames* e *Series* utilizando as funções do **matplotlib.pyplot**.
O **matplotlib** transforma os dados em gráficos através de duas componentes: **figuras** (por exemplo janelas) e **eixos** (uma região onde os pontos podem ser determinados por meio de coordenadas). Se temos uma figura bidimensional, tipicamente os eixos são *x*-*y*, mas podemos ter coordenadas polares também. Se temos uma figura tridimensional, os eixos tipicamente são *x*-*y*-*z*, mas também podemos ter coordenadas esféricas, cilíndricas, etc.
Como as figuras são determinadas pelas posições no plano ou no espaço, utilizamos com mais frequência os **eixos** de um objeto do **matplotlib**.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt # Aqui utilizaremos a biblioteca matplotlib
serie_Idade = pd.Series({'Ana':20, 'João': 19, 'Maria': 21, 'Pedro': 22, 'Túlio': 20}, name="Idade")
serie_Peso = pd.Series({'Ana':55, 'João': 80, 'Maria': 62, 'Pedro': 67, 'Túlio': 73}, name="Peso")
serie_Altura = pd.Series({'Ana':162, 'João': 178, 'Maria': 162, 'Pedro': 165, 'Túlio': 171}, name="Altura")
dicionario_series_exemplo = {'Idade': serie_Idade, 'Peso': serie_Peso, 'Altura': serie_Altura}
df_dict_series = pd.DataFrame(dicionario_series_exemplo);df_dict_series
df_exemplo = pd.read_csv('06b-exemplo_data.csv', index_col=0)
df_exemplo['coluna_3'] = pd.Series([1,2,3,4,5,6,7,8,np.nan,np.nan],index=df_exemplo.index)
df_exemplo
covid_PB = pd.read_csv('https://superset.plataformatarget.com.br/superset/explore_json/?form_data=%7B%22slice_id%22%3A1550%7D&csv=true',
sep=',', index_col=0)
covid_PB.head()
covid_BR = pd.read_excel("06b-HIST_PAINEL_COVIDBR_18jul2020.xlsx")
covid_BR.head()
```
## Gráfico de Linhas
```
fig, ax = plt.subplots() # Este comando cria uma figura com um eixo
ax.plot(df_exemplo.index, df_exemplo['coluna_1'], label = 'Primeira Coluna') # Inserimos a linha relativa à coluna 1
ax.plot(df_exemplo.index, df_exemplo['coluna_2'], label = 'Segunda Coluna') # Inserimos a linha relativa à coluna 2
ax.plot(df_exemplo.index, df_exemplo['coluna_3'], label = 'Terceira Coluna') # Inserimos a linha relativa à coluna 3
ax.set_xlabel('Data') # Rótulo do eixo x
ax.set_ylabel('Valor') # Rótulo do eixo y
ax.set_title("Gráfico do df_exemplo")
ax.legend()
fig, ax = plt.subplots() # Este comando cria uma figura com um eixo
ax.plot(df_exemplo.index, df_exemplo['coluna_1'], label = 'Primeira Coluna',
color = 'red') # Inserimos a linha relativa à coluna 1, definimos a cor vermelha
ax.plot(df_exemplo.index, df_exemplo['coluna_2'],
label = 'Segunda Coluna', linewidth=6.0) # Inserimos a linha relativa à coluna 2 e aumentamos a grossura da linha
ax.plot(df_exemplo.index, df_exemplo['coluna_3'], label = 'Terceira Coluna') # Inserimos a linha relativa à coluna 3
ax.set_xlabel('Data') # Rótulo do eixo x
ax.set_ylabel('Valor') # Rótulo do eixo y
ax.set_title("Gráfico do df_exemplo")
ax.legend()
fig.autofmt_xdate()
covid_PB.index = pd.to_datetime(covid_PB.index)
covid_PB_casos_obitos = covid_PB[['casosAcumulados', 'obitosAcumulados']].sort_index()
fig, ax = plt.subplots()
ax.plot(covid_PB_casos_obitos.index, covid_PB_casos_obitos['casosAcumulados'], label = 'Total de Casos',
color = 'red')
ax.plot(covid_PB_casos_obitos.index, covid_PB_casos_obitos['obitosAcumulados'],
label = 'Total de Óbitos', color = 'black')
ax.set_xlabel('Data') # Rótulo do eixo x
ax.set_ylabel('Total') # Rótulo do eixo y
ax.set_title("Casos e Óbitos de COVID-19 na Paraíba")
ax.legend()
fig.autofmt_xdate()
```
Podemos alterar a apresentação das datas utilizando o subpacote *dates* do *matplotlib*.
```
import matplotlib.dates as mdates
fig, ax = plt.subplots()
ax.plot(covid_PB_casos_obitos.index, covid_PB_casos_obitos['casosAcumulados'], label = 'Total de Casos', color = 'red')
ax.plot(covid_PB_casos_obitos.index, covid_PB_casos_obitos['obitosAcumulados'], label = 'Total de Óbitos', color = 'black')
ax.set_xlabel('Data') # Rótulo do eixo x
ax.set_ylabel('Total') # Rótulo do eixo y
ax.set_title("Casos e Óbitos de COVID-19 na Paraíba")
ax.legend()
ax.xaxis.set_minor_locator(mdates.DayLocator(interval=7)) #Intervalo entre os tracinhos
ax.xaxis.set_major_locator(mdates.DayLocator(interval=21)) #Intervalo entre as datas
ax.xaxis.set_major_formatter(mdates.DateFormatter('%d/%m/%Y')) #Formato da data
fig.autofmt_xdate()
```
Vamos agora alterar o formato dos números do eixo *y*. Para tanto iremos definir uma função para realizar a formatação e utilizaremos a função *FuncFormatter* do subpacote *matplotlib.ticker*.
```
from matplotlib.ticker import FuncFormatter
def inserir_mil(x, pos):
return '%1i mil' % (x*1e-3) if x != 0 else 0
fig, ax = plt.subplots()
ax.plot(covid_PB_casos_obitos.index, covid_PB_casos_obitos['casosAcumulados'], label = 'Total de Casos', color = 'red')
ax.plot(covid_PB_casos_obitos.index, covid_PB_casos_obitos['obitosAcumulados'], label = 'Total de Óbitos', color = 'black')
ax.set_xlabel('Data') # Rótulo do eixo x
ax.set_ylabel('Total') # Rótulo do eixo y
ax.set_title("Casos e Óbitos de COVID-19 na Paraíba")
ax.legend()
ax.xaxis.set_minor_locator(mdates.DayLocator(interval=7)) #Intervalo entre os tracinhos
ax.xaxis.set_major_locator(mdates.DayLocator(interval=21)) #Intervalo entre as datas
ax.xaxis.set_major_formatter(mdates.DateFormatter('%d/%m/%Y')) #Formato da data
fig.autofmt_xdate()
ax.yaxis.set_major_formatter(FuncFormatter(inserir_mil))
covid_regioes = pd.DataFrame()
regioes = covid_BR.query('regiao != "Brasil"')['regiao'].drop_duplicates().array
for regiao in regioes:
temp_series = covid_BR.set_index('data').query('regiao == @regiao')['obitosAcumulado'].groupby('data').sum()/2
#Obs.: Utilizamos @ na frente do nome da variável para utilizar o valor da variável no query.
#Obs.: Dividimos por 2, pois os óbitos estão sendo contados duas vezes,
#uma para quando codmun == nan e outra quando não é nulo
temp_series.name = 'obitos_' + regiao
covid_regioes = pd.concat([covid_regioes, temp_series], axis=1)
covid_regioes.index = pd.to_datetime(covid_regioes.index)
covid_regioes
fig, ax = plt.subplots()
ax.plot(covid_regioes.index, covid_regioes['obitos_Norte'], label = 'Norte')
ax.plot(covid_regioes.index, covid_regioes['obitos_Nordeste'], label = 'Nordeste')
ax.plot(covid_regioes.index, covid_regioes['obitos_Sudeste'], label = 'Sudeste')
ax.plot(covid_regioes.index, covid_regioes['obitos_Sul'], label = 'Sul')
ax.plot(covid_regioes.index, covid_regioes['obitos_Centro-Oeste'], label = 'Centro-Oeste')
ax.set_xlabel('Data') # Rótulo do eixo x
ax.set_ylabel('Total de Óbitos') # Rótulo do eixo y
ax.set_title("Óbitos de COVID-19 nas regiões do Brasil")
ax.legend()
ax.xaxis.set_minor_locator(mdates.DayLocator(interval=7)) #Intervalo entre os tracinhos
ax.xaxis.set_major_locator(mdates.DayLocator(interval=21)) #Intervalo entre as datas
ax.xaxis.set_major_formatter(mdates.DateFormatter('%d/%m/%Y')) #Formato da data
fig.autofmt_xdate()
ax.yaxis.set_major_formatter(FuncFormatter(inserir_mil))
```
## Gráfico de Colunas e de Linhas
```
covid_Regioes = covid_BR[['regiao','obitosNovos']].groupby('regiao').sum().query('regiao != "Brasil"')/2
fig, ax = plt.subplots()
ax.bar(covid_Regioes.index, covid_Regioes['obitosNovos'])
ax.yaxis.set_major_formatter(FuncFormatter(inserir_mil))
ax.set_ylabel('Total de Óbitos') # Rótulo do eixo y
ax.set_title("Óbitos de COVID-19 nas regiões do Brasil até o dia 18/07/2020")
```
Podemos inserir o total de cada região em cima do retângulo correspondente. Para tanto, utilizaremos a seguinte função disponível na página do *matplotlib*:
```
def autolabel(rects):
"""Attach a text label above each bar in *rects*, displaying its height."""
for rect in rects:
height = rect.get_height()
#ax.annotate('{}'.format(height), #antigo
ax.annotate('{:.0f}'.format(height), #Modificamos para apresentar o número inteiro
xy=(rect.get_x() + rect.get_width() / 2, height),
xytext=(0, 3), # 3 points vertical offset
textcoords="offset points",
ha='center', va='bottom')
covid_Regioes = covid_BR[['regiao','obitosNovos']].groupby('regiao').sum().query('regiao != "Brasil"')/2
fig, ax = plt.subplots()
plt.ylim(0, 40000) # aumentamos o limite da coordenada y
retangulos = ax.bar(covid_Regioes.index, covid_Regioes['obitosNovos'])
ax.yaxis.set_major_formatter(FuncFormatter(inserir_mil))
ax.set_ylabel('Total de Óbitos') # Rótulo do eixo y
ax.set_title("Óbitos de COVID-19 nas regiões do Brasil até o dia 18/07/2020")
autolabel(retangulos)
```
* Para realizarmos os "plots" agrupados das barras devemos realizar 5 "plots" distintos, um para cada barra.
* Cada plot sofrerá uma translação (exceto o do meio).
* Iremos reduzir a largura de cada barra.
```
covid_Regioes = covid_BR[['regiao','obitosNovos']].groupby('regiao').sum().query('regiao != "Brasil"')/2
largura = 0.3
fig, ax = plt.subplots()
retangulo1 = ax.bar([-2*largura], covid_Regioes.loc[['Norte'],['obitosNovos']].to_numpy()[0], largura, label='Norte')
retangulo2 = ax.bar([-largura], covid_Regioes.loc[['Nordeste'],['obitosNovos']].to_numpy()[0], largura, label='Nordeste')
retangulo3 = ax.bar([0], covid_Regioes.loc[['Centro-Oeste'],['obitosNovos']].to_numpy()[0], largura, label='Centro-Oeste')
retangulo4 = ax.bar([largura], covid_Regioes.loc[['Sudeste'],['obitosNovos']].to_numpy()[0], largura, label='Sudeste')
retangulo5 = ax.bar([2*largura], covid_Regioes.loc[['Sul'],['obitosNovos']].to_numpy()[0], largura, label='Sul')
ax.yaxis.set_major_formatter(FuncFormatter(inserir_mil))
ax.set_ylabel('Total de Óbitos') # Rótulo do eixo y
ax.set_title("Óbitos de COVID-19 nas regiões do Brasil até o dia 18/07/2020")
autolabel(retangulo1); autolabel(retangulo2); autolabel(retangulo3); autolabel(retangulo4); autolabel(retangulo5)
plt.ylim(0, 40000) # aumentamos o limite da coordenada y
plt.xlim(-1, 1.3) # Limites que iremos utilizar na coordenada y
plt.xticks([], []) # Remover os "ticks" no eixo x
#plt.xticks([0], ['Região']) # Se quisermos incluir o rótulo "Região" na posição 0 do eixo x
ax.legend(title="Região")
```
Para empilharmos as barras manualmente devemos utilizar o argumento **bottom**:
```
largura = 0.25
obitos_norte = covid_Regioes.loc[['Norte'],['obitosNovos']].to_numpy()[0]
obitos_nordeste = covid_Regioes.loc[['Nordeste'],['obitosNovos']].to_numpy()[0]
obitos_centro_oeste = covid_Regioes.loc[['Centro-Oeste'],['obitosNovos']].to_numpy()[0]
obitos_sudeste = covid_Regioes.loc[['Sudeste'],['obitosNovos']].to_numpy()[0]
obitos_sul = covid_Regioes.loc[['Sul'],['obitosNovos']].to_numpy()[0]
fig, ax = plt.subplots()
retangulo1 = ax.bar([0.5], obitos_norte, largura, label='Norte')
retangulo2 = ax.bar([0.5], obitos_nordeste, largura, label='Nordeste', bottom = obitos_norte)
retangulo3 = ax.bar([0.5], obitos_centro_oeste, largura, label='Centro-Oeste', bottom = obitos_norte + obitos_nordeste)
retangulo4 = ax.bar([0.5], obitos_sudeste, largura, label='Sudeste', bottom = obitos_norte +
obitos_nordeste + obitos_centro_oeste)
retangulo5 = ax.bar([0.5], obitos_sul, largura, label='Sul', bottom = obitos_norte +
obitos_nordeste + obitos_centro_oeste + obitos_sudeste)
ax.yaxis.set_major_formatter(FuncFormatter(inserir_mil))
ax.set_ylabel('Total de Óbitos') # Rótulo do eixo y
ax.set_title("Óbitos de COVID-19 nas regiões do Brasil até o dia 18/07/2020")
plt.xticks([], [])
#plt.xticks([0], ['Região']) # Se quisermos incluir o rótulo "Região" na posição 0 do eixo x
plt.xlim(0,1)
ax.legend(title="Região")
x = np.arange(len(df_dict_series.index))
largura = 0.25
fig, ax = plt.subplots()
retangulo1 = ax.bar(x - largura, df_dict_series.Idade, largura, label='Idade')
retangulo2 = ax.bar(x, df_dict_series.Peso, largura, label='Peso')
retangulo3 = ax.bar(x + largura, df_dict_series.Altura, largura, label='Altura')
autolabel(retangulo1); autolabel(retangulo2); autolabel(retangulo3)
plt.ylim(0,200)
plt.xlim(-0.5,6)
ax.set_ylabel('Valor')
ax.set_title('Características')
ax.set_xticks(x)
ax.set_xticklabels(df_dict_series.index)
ax.legend()
x = np.arange(len(df_dict_series.index))
largura = 0.25
fig, ax = plt.subplots()
retangulo1 = ax.bar(x, df_dict_series.Idade, largura, label='Idade')
retangulo2 = ax.bar(x, df_dict_series.Peso, largura, label='Peso', bottom = df_dict_series.Idade)
retangulo3 = ax.bar(x, df_dict_series.Altura, largura, label='Altura', bottom = df_dict_series.Idade + df_dict_series.Peso)
plt.xlim(-0.5,6)
ax.set_ylabel('Valor')
ax.set_title('Características')
ax.set_xticks(x)
ax.set_xticklabels(df_dict_series.index)
ax.legend()
```
* Para construir os gráficos de barras procedemos de maneira análoga ao que foi feito acima.
* Substituímos o método **bar** por **barh**
* Caso haja interesse deve modificar a função autolabel, alterando a altura, *height*, pela largura, *width*.
## Gráfico de Setores
Neste caso devemos modificar o *DataFrame* para conter percentuais (ou pesos).
* Vamos usar os parâmetros:
* **autopct** que adiciona o percentual de cada "fatia".
* **shadow** que adiciona sombra
* **explode** que separa fatias selecionadas
```
df_dict_series_pct = df_dict_series.copy()
df_dict_series_pct.Idade = df_dict_series_pct.Idade/df_dict_series_pct.Idade.sum()
df_dict_series_pct.Peso = df_dict_series_pct.Peso/df_dict_series_pct.Peso.sum()
df_dict_series_pct.Altura = df_dict_series_pct.Altura/df_dict_series_pct.Altura.sum()
df_dict_series_pct
figs, axs = plt.subplots(1,3, figsize=(22,7)) #1 linha3 e 3 colunas de "plots"
axs[0].pie(df_dict_series_pct.Idade, labels=df_dict_series_pct.index, autopct='%1.1f%%', shadow=True)
axs[0].axis('equal') # Igualando os eixos para garantir que obteremos um círculo
axs[0].legend(loc = 'upper left')
axs[0].set_title('Idade')
axs[1].pie(df_dict_series_pct.Peso, labels=df_dict_series_pct.index, autopct='%1.1f%%', shadow=True)
axs[1].axis('equal')
axs[1].legend(loc = 'upper left')
axs[1].set_title('Peso')
axs[2].pie(df_dict_series_pct.Altura, labels=df_dict_series_pct.index, autopct='%1.1f%%', shadow=True)
axs[2].axis('equal')
axs[2].legend(loc = 'upper left')
_ = axs[2].set_title('Altura') #Atribuímos a uma variável para não termos saída
covid_Regioes_pct = covid_Regioes/covid_Regioes.sum()
covid_Regioes_pct['explodir'] = covid_Regioes_pct.index.map(lambda regiao: 0.1 if regiao == 'Nordeste' else 0)
covid_Regioes_pct
fig, ax = plt.subplots(figsize = (10,10))
ax.pie(covid_Regioes_pct.obitosNovos, explode=covid_Regioes_pct.explodir,
labels=covid_Regioes_pct.index, autopct='%1.1f%%', shadow=True)
ax.set_title('Percentual de Óbitos de COVID-19 nas Regiões do Brasil até o Dia 18/07/2020')
_ = ax.axis('equal')
```
## Gráfico de Dispersão
Para gráficos de dispersão vários argumentos são os mesmos que já vimos no método **plot** do *pandas*.
```
fig, ax = plt.subplots()
ax.scatter(df_exemplo.index, df_exemplo['coluna_1'])
fig.autofmt_xdate()
ax.set_xlabel('Data')
ax.set_ylabel('Valores da Coluna 1')
ax.set_title('Gráfico do df_exemplo')
fig, ax = plt.subplots()
ax.scatter(df_exemplo.index, df_exemplo['coluna_1'], s = np.abs(df_exemplo['coluna_2'])*100)
fig.autofmt_xdate()
ax.set_xlabel('Data')
ax.set_ylabel('Valores da Coluna 1')
ax.set_title('Gráfico do df_exemplo')
covid_PB_casos_obitos = covid_PB[['obitosNovos', 'casosNovos']].sort_index()
fig, ax = plt.subplots()
grafico = ax.scatter(covid_PB_casos_obitos.index, covid_PB_casos_obitos.casosNovos, c = covid_PB_casos_obitos.obitosNovos)
fig.autofmt_xdate()
ax.set_xlabel('Data')
ax.set_ylabel('Casos COVID-19 em PB')
ax.set_title('Casos e Óbitos de COVID-19 na Paraíba')
color_map=ax.get_children()[4]
plt.colorbar(grafico, label = 'Óbitos')
covid_PB_casos_obitos = covid_PB[['obitosNovos', 'casosNovos']].sort_index()
fig, ax = plt.subplots()
grafico = ax.scatter(covid_PB_casos_obitos.index, covid_PB_casos_obitos.casosNovos, c = covid_PB_casos_obitos.obitosNovos,
cmap='cool')
fig.autofmt_xdate()
ax.set_xlabel('Data')
ax.set_ylabel('Casos COVID-19 em PB')
ax.set_title('Casos e Óbitos de COVID-19 na Paraíba')
color_map=ax.get_children()[4]
plt.colorbar(grafico, label = 'Óbitos')
fig, ax = plt.subplots()
ax.scatter(df_exemplo.index, df_exemplo['coluna_1'], label = 'Coluna 1', color = 'black')
ax.scatter(df_exemplo.index, df_exemplo['coluna_2'], label = 'Coluna 2', color = 'red')
ax.scatter(df_exemplo.index, df_exemplo['coluna_3'], label = 'Coluna 3', color = 'green')
fig.autofmt_xdate()
ax.legend()
ax.set_ylabel("Valor")
ax.set_xlabel("Data")
```
## Gráficos Lado a Lado
```
#Vamos modificar esta função para podermos utilizá-la quando temos mais de um gráfico ao mesmo tempo
def autolabel(rects, ax):
"""Attach a text label above each bar in *rects*, displaying its height."""
for rect in rects:
height = rect.get_height()
#ax.annotate('{}'.format(height), #antigo
ax.annotate('{:.0f}'.format(height), #Modificamos para apresentar o número inteiro
xy=(rect.get_x() + rect.get_width() / 2, height),
xytext=(0, 3), # 3 points vertical offset
textcoords="offset points",
ha='center', va='bottom')
covid_Regioes = covid_BR[['regiao','obitosNovos']].groupby('regiao').sum().query('regiao != "Brasil"')/2
figs, axs = plt.subplots(1,2, figsize=(22,7))
axs[0].set_ylim(0, 40000) # aumentamos o limite da coordenada y
retangulos = axs[0].bar(covid_Regioes.index, covid_Regioes['obitosNovos'])
axs[0].yaxis.set_major_formatter(FuncFormatter(inserir_mil))
axs[0].set_ylabel('Total de Óbitos') # Rótulo do eixo y
axs[0].set_title("Óbitos de COVID-19 nas regiões do Brasil até o dia 18/07/2020")
autolabel(retangulos, axs[0])
axs[1].pie(covid_Regioes_pct.obitosNovos, explode=covid_Regioes_pct.explodir,
labels=covid_Regioes_pct.index, autopct='%1.1f%%', shadow=True)
axs[1].set_title('Percentual de Óbitos de COVID-19 nas Regiões do Brasil até o Dia 18/07/2020')
_ = axs[1].axis('equal')
```
## Histograma
```
fig, ax = plt.subplots()
ax.hist(covid_regioes.obitos_Nordeste, bins=30, color='lime')
ax.set_ylabel('Frequência')
ax.set_xlabel('Óbitos Diários por COVID-19')
ax.set_title('Nordeste')
fig, axs = plt.subplots(1,2, sharey=True, figsize = (15,7)) #sharey=True indica que o eixo y será o mesmo para todos os gráficos
axs[0].hist(covid_regioes.obitos_Nordeste, bins=30, histtype='step', color='red')
axs[0].set_ylabel('Frequência')
axs[0].set_xlabel('Óbitos Diários por COVID-19')
axs[0].set_title('Nordeste')
axs[1].hist(covid_regioes.obitos_Nordeste, bins=30, fill=False, edgecolor='red')
axs[1].set_ylabel('Frequência')
axs[1].set_xlabel('Óbitos Diários por COVID-19')
axs[1].set_title('Nordeste')
fig, ax = plt.subplots(figsize=(10,10))
ax.hist([covid_regioes.obitos_Nordeste, covid_regioes.obitos_Sudeste], bins=30, histtype='step', label=['Nordeste', 'Sudeste'])
ax.set_ylabel('Frequência')
ax.set_xlabel('Óbitos Diários por COVID-19')
_ = ax.legend()
```
## BoxPlot
O método para criar o *boxplot* utilizando o **matplotlib** se resume a fornecer uma lista (ou similar) de valores para os quais queremos os *boxplots* e uma lista (ou similar) contendo as posições nas quais queremos que os *boxplots* apareçam.
```
fig, ax = plt.subplots()
dados = [df_exemplo['coluna_1'], df_exemplo['coluna_2'], df_exemplo['coluna_3'].dropna()]
posicoes = np.array(range(len(dados))) + 1
ax.boxplot(dados, positions=posicoes)
_ = ax.set_xticklabels(['Coluna 1', 'Coluna 2', 'Coluna 3'])
covid_norte = covid_regioes.obitos_Norte
covid_nordeste = covid_regioes.obitos_Nordeste
covid_sudeste = covid_regioes.obitos_Sudeste
covid_sul = covid_regioes.obitos_Sul
covid_centro_oeste = covid_regioes['obitos_Centro-Oeste']
covid_box = [covid_norte, covid_nordeste, covid_sudeste, covid_sul, covid_centro_oeste]
fig, ax = plt.subplots()
posicoes = np.array(range(len(covid_box))) + 1
ax.boxplot(covid_box, 1, positions=posicoes, sym='+')
_ = ax.set_xticklabels(['Norte', 'Nordeste', 'Sudeste', 'Sul', 'Centro-Oeste'])
covid_box_2 = [covid_sul, covid_centro_oeste]
fig, ax = plt.subplots()
posicoes = np.array(range(len(covid_box_2))) + 1
ax.boxplot(covid_box_2, 1, positions=posicoes, sym='r+') #r indica 'red', vermelho, + é o símbolo para o outlier
_ = ax.set_xticklabels(['Sul', 'Centro-Oeste'])
covid_box_2 = [covid_sul, covid_centro_oeste]
fig, ax = plt.subplots()
posicoes = np.array(range(len(covid_box_2))) + 1
ax.boxplot(covid_box_2, 1, positions=posicoes, sym='g.') #r indica 'red', vermelho, + é o símbolo para o outlier
_ = ax.set_xticklabels(['Sul', 'Centro-Oeste'])
```
**Obs.:** Muitos dos argumentos utilizados nos métodos acima também funcionam no método **plot** do *pandas*, vale a pena testar!
| github_jupyter |
# # Duramat Webinar: US NREL Electric Futures 2021
This journal simulates the Reference and High Electrification scenarios from Electrification Futures, and comparing to a glass baseline with High bifacial future projection.
Installed Capacity considerations from bifacial installations are not considered here.
Results from this journal were presented during Duramat's webinar April 2021 – “The Impacts of Module Reliability and Lifetime on PV in the Circular Economy" presented by Teresa Barnes, Silvana Ayala, and Heather Mirletz, NREL.
```
import os
from pathlib import Path
testfolder = str(Path().resolve().parent.parent / 'PV_ICE' / 'TEMP' / 'DURAMAT')
# Another option using relative address; for some operative systems you might need '/' instead of '\'
# testfolder = os.path.abspath(r'..\..\PV_DEMICE\TEMP')
print ("Your simulation will be stored in %s" % testfolder)
MATERIALS = ['glass','silver','silicon', 'copper','aluminium_frames']
MATERIAL = MATERIALS[0]
MODULEBASELINE = r'..\..\baselines\ElectrificationFutures_2021\baseline_modules_US_NREL_Electrification_Futures_2021_basecase.csv'
MODULEBASELINE_High = r'..\..\baselines\ElectrificationFutures_2021\baseline_modules_US_NREL_Electrification_Futures_2021_LowREHighElec.csv'
import PV_ICE
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
PV_ICE.__version__
plt.rcParams.update({'font.size': 22})
plt.rcParams['figure.figsize'] = (12, 5)
pwd
r1 = PV_ICE.Simulation(name='Simulation1', path=testfolder)
r1.createScenario(name='base', file=MODULEBASELINE)
r1.scenario['base'].addMaterials(MATERIALS, r'..\..\baselines')
r1.createScenario(name='high', file=MODULEBASELINE_High)
r1.scenario['high'].addMaterials(MATERIALS, r'..\..\baselines')
r2 = PV_ICE.Simulation(name='bifacialTrend', path=testfolder)
r2.createScenario(name='base', file=MODULEBASELINE)
r2.scenario['base'].addMaterials(MATERIALS, r'..\..\baselines')
MATERIALBASELINE = r'..\..\baselines\PVSC_2021\baseline_material_glass_bifacialTrend.csv'
r2.scenario['base'].addMaterial('glass', file=MATERIALBASELINE)
r2.createScenario(name='high', file=MODULEBASELINE_High)
r2.scenario['high'].addMaterials(MATERIALS, r'..\..\baselines')
MATERIALBASELINE = r'..\..\baselines\PVSC_2021\baseline_material_glass_bifacialTrend.csv'
r2.scenario['high'].addMaterial('glass', file=MATERIALBASELINE)
IRENA= False
ELorRL = 'EL'
if IRENA:
r1.scenMod_IRENIFY(scenarios=['base', 'high'], ELorRL = ELorRL )
r2.scenMod_IRENIFY(scenarios=['base', 'high'], ELorRL = ELorRL )
title_Method = 'Irena_'+ELorRL
else:
title_Method = 'PVICE'
r1.calculateMassFlow()
r2.calculateMassFlow()
objects = [r1, r2]
scenarios = ['base', 'high']
pvice_Usyearly1, pvice_Uscum1 = r1.aggregateResults()
pvice_Usyearly2, pvice_Uscum2 = r2.aggregateResults()
UScum = pd.concat([pvice_Uscum1, pvice_Uscum2], axis=1)
USyearly = pd.concat([pvice_Usyearly1, pvice_Usyearly2], axis=1)
UScum.to_csv('pvice_USCum.csv')
USyearly.to_csv('pvice_USYearly.csv')
# OLD METHOD
'''
USyearly=pd.DataFrame()
keyword='mat_Total_Landfilled'
materials = ['glass', 'silicon', 'silver', 'copper', 'aluminium_frames']
# Loop over objects
for kk in range(0, len(objects)):
obj = objects[kk]
# Loop over Scenarios
for jj in range(0, len(scenarios)):
case = scenarios[jj]
for ii in range (0, len(materials)):
material = materials[ii]
foo = obj.scenario[case].material[material].materialdata[keyword].copy()
foo = foo.to_frame(name=material)
USyearly["Waste_"+material+'_'+obj.name+'_'+case] = foo[material]
filter_col = [col for col in USyearly if (col.startswith('Waste_') and col.endswith(obj.name+'_'+case)) ]
USyearly['Waste_Module_'+obj.name+'_'+case] = USyearly[filter_col].sum(axis=1)
# Converting to grams to Tons.
USyearly.head(20)
keyword='mat_Total_EOL_Landfilled'
materials = ['glass', 'silicon', 'silver', 'copper', 'aluminium_frames']
# Loop over objects
for kk in range(0, len(objects)):
obj = objects[kk]
# Loop over Scenarios
for jj in range(0, len(scenarios)):
case = scenarios[jj]
for ii in range (0, len(materials)):
material = materials[ii]
foo = obj.scenario[case].material[material].materialdata[keyword].copy()
foo = foo.to_frame(name=material)
USyearly["Waste_EOL_"+material+'_'+obj.name+'_'+case] = foo[material]
filter_col = [col for col in USyearly if (col.startswith('Waste_EOL_') and col.endswith(obj.name+'_'+case)) ]
USyearly['Waste_EOL_Module_'+obj.name+'_'+case] = USyearly[filter_col].sum(axis=1)
# Converting to grams to Tons.
USyearly.head(20)
keyword='mat_Virgin_Stock'
materials = ['glass', 'silicon', 'silver', 'copper', 'aluminium_frames']
# Loop over objects
for kk in range(0, len(objects)):
obj = objects[kk]
# Loop over Scenarios
for jj in range(0, len(scenarios)):
case = scenarios[jj]
for ii in range (0, len(materials)):
material = materials[ii]
foo = obj.scenario[case].material[material].materialdata[keyword].copy()
foo = foo.to_frame(name=material)
USyearly["VirginStock_"+material+'_'+obj.name+'_'+case] = foo[material]
filter_col = [col for col in USyearly if (col.startswith('VirginStock_') and col.endswith(obj.name+'_'+case)) ]
USyearly['VirginStock_Module_'+obj.name+'_'+case] = USyearly[filter_col].sum(axis=1)
# ### Converting to grams to METRIC Tons.
USyearly = USyearly/1000000 # This is the ratio for Metric tonnes
#907185 -- this is for US tons
UScum = USyearly.copy()
UScum = UScum.cumsum()
keyword='Installed_Capacity_[W]'
materials = ['glass', 'silicon', 'silver', 'copper', 'aluminium_frames']
# Loop over SF Scenarios
for kk in range(0, len(objects)):
obj = objects[kk]
# Loop over Scenarios
for jj in range(0, len(scenarios)):
case = scenarios[jj]
foo = obj.scenario[case].data[keyword]
foo = foo.to_frame(name=keyword)
UScum["Capacity_"+obj.name+'_'+case] = foo[keyword]
USyearly.index = r1.scenario['base'].data['year']
UScum.index = r1.scenario['base'].data['year']
USyearly.to_csv('USyearly_Oldmethod.csv')
UScum.to_csv('UScum_Oldmethod.csv')
''';
```
# ## Mining Capacity
```
mining2020_aluminum = 65267000
mining2020_silver = 22260
mining2020_copper = 20000000
mining2020_silicon = 8000000
objects = [r1, r2]
scenarios = ['base', 'high']
plt.rcParams.update({'font.size': 10})
plt.rcParams['figure.figsize'] = (12, 8)
keyw='VirginStock_'
materials = ['glass', 'silicon', 'silver', 'copper', 'aluminium_frames']
fig, axs = plt.subplots(1,1, figsize=(4, 6), facecolor='w', edgecolor='k')
fig.subplots_adjust(hspace = .3, wspace=.2)
# Loop over CASES
name2 = 'Simulation1_high_[Tonnes]'
name0 = 'Simulation1_base_[Tonnes]'
# ROW 2, Aluminum and Silicon: g- 4 aluminum k - 1 silicon orange - 3 copper gray - 2 silver
axs.plot(USyearly[keyw+materials[2]+'_'+name2]*100/mining2020_silver,
color = 'gray', linewidth=2.0, label='Silver')
axs.fill_between(USyearly.index, USyearly[keyw+materials[2]+'_'+name0]*100/mining2020_silver, USyearly[keyw+materials[2]+'_'+name2]*100/mining2020_silver,
color='gray', lw=3, alpha=.3)
axs.plot(USyearly[keyw+materials[1]+'_'+name2]*100/mining2020_silicon,
color = 'k', linewidth=2.0, label='Silicon')
axs.fill_between(USyearly.index, USyearly[keyw+materials[1]+'_'+name0]*100/mining2020_silicon,
USyearly[keyw+materials[1]+'_'+name2]*100/mining2020_silicon,
color='k', lw=3, alpha=.5)
axs.plot(USyearly[keyw+materials[4]+'_'+name2]*100/mining2020_aluminum,
color = 'g', linewidth=2.0, label='Aluminum')
axs.fill_between(USyearly.index, USyearly[keyw+materials[4]+'_'+name0]*100/mining2020_aluminum,
USyearly[keyw+materials[4]+'_'+name2]*100/mining2020_aluminum,
color='g', lw=3, alpha=.3)
axs.plot(USyearly[keyw+materials[3]+'_'+name2]*100/mining2020_copper,
color = 'orange', linewidth=2.0, label='Copper')
axs.fill_between(USyearly.index, USyearly[keyw+materials[3]+'_'+name0]*100/mining2020_copper,
USyearly[keyw+materials[3]+'_'+name2]*100/mining2020_copper,
color='orange', lw=3, alpha=.3)
axs.set_xlim([2020,2050])
axs.legend()
#axs.set_yscale('log')
#axs.set_ylabel('Virgin material needs as a percentage of 2020 global mining production capacity [%]')
fig.savefig(title_Method+' Fig_1x1_MaterialNeeds Ratio to Production_NREL2018.png', dpi=600)
plt.rcParams.update({'font.size': 15})
plt.rcParams['figure.figsize'] = (15, 8)
keyw='VirginStock_'
materials = ['glass', 'silicon', 'silver', 'copper', 'aluminium_frames']
f, (a0, a1) = plt.subplots(1, 2, gridspec_kw={'width_ratios': [3, 1]})
########################
# SUBPLOT 1
########################
#######################
# loop plotting over scenarios
name2 = 'Simulation1_high_[Tonnes]'
name0 = 'Simulation1_base_[Tonnes]'
# SCENARIO 1 ***************
modulemat = (USyearly[keyw+materials[0]+'_'+name0]+USyearly[keyw+materials[1]+'_'+name0]+
USyearly[keyw+materials[2]+'_'+name0]+USyearly[keyw+materials[3]+'_'+name0]+
USyearly[keyw+materials[4]+'_'+name0])
glassmat = (USyearly[keyw+materials[0]+'_'+name0])
modulemat = modulemat/1000000
glassmat = glassmat/1000000
a0.plot(USyearly.index, modulemat, 'k.', linewidth=5, label='S1: '+name0+' module mass')
a0.plot(USyearly.index, glassmat, 'k', linewidth=5, label='S1: '+name0+' glass mass only')
a0.fill_between(USyearly.index, glassmat, modulemat, color='k', alpha=0.3,
interpolate=True)
# SCENARIO 2 ***************
modulemat = (USyearly[keyw+materials[0]+'_'+name2]+USyearly[keyw+materials[1]+'_'+name2]+
USyearly[keyw+materials[2]+'_'+name2]+USyearly[keyw+materials[3]+'_'+name2]+
USyearly[keyw+materials[4]+'_'+name2])
glassmat = (USyearly[keyw+materials[0]+'_'+name2])
modulemat = modulemat/1000000
glassmat = glassmat/1000000
a0.plot(USyearly.index, modulemat, 'c.', linewidth=5, label='S2: '+name2+' module mass')
a0.plot(USyearly.index, glassmat, 'c', linewidth=5, label='S2: '+name2+' glass mass only')
a0.fill_between(USyearly.index, glassmat, modulemat, color='c', alpha=0.3,
interpolate=True)
a0.legend()
a0.set_title('Yearly Virgin Material Needs by Scenario')
a0.set_ylabel('Mass [Million Tonnes]')
a0.set_xlim([2020, 2050])
a0.set_xlabel('Years')
########################
# SUBPLOT 2
########################
#######################
# Calculate
cumulations2050 = {}
for ii in range(0, len(materials)):
matcum = []
matcum.append(UScum[keyw+materials[ii]+'_'+name0].loc[2050])
matcum.append(UScum[keyw+materials[ii]+'_'+name2].loc[2050])
cumulations2050[materials[ii]] = matcum
dfcumulations2050 = pd.DataFrame.from_dict(cumulations2050)
dfcumulations2050 = dfcumulations2050/1000000 # in Million Tonnes
dfcumulations2050['bottom1'] = dfcumulations2050['glass']
dfcumulations2050['bottom2'] = dfcumulations2050['bottom1']+dfcumulations2050['aluminium_frames']
dfcumulations2050['bottom3'] = dfcumulations2050['bottom2']+dfcumulations2050['silicon']
dfcumulations2050['bottom4'] = dfcumulations2050['bottom3']+dfcumulations2050['copper']
## Plot BARS Stuff
ind=np.arange(2)
width=0.35 # width of the bars.
p0 = a1.bar(ind, dfcumulations2050['glass'], width, color='c')
p1 = a1.bar(ind, dfcumulations2050['aluminium_frames'], width,
bottom=dfcumulations2050['bottom1'])
p2 = a1.bar(ind, dfcumulations2050['silicon'], width,
bottom=dfcumulations2050['bottom2'])
p3 = a1.bar(ind, dfcumulations2050['copper'], width,
bottom=dfcumulations2050['bottom3'])
p4 = a1.bar(ind, dfcumulations2050['silver'], width,
bottom=dfcumulations2050['bottom4'])
a1.yaxis.set_label_position("right")
a1.yaxis.tick_right()
a1.set_ylabel('Virgin Material Cumulative Needs 2020-2050 [Million Tonnes]')
a1.set_xlabel('Scenario')
a1.set_xticks(ind, ('S1', 'S2'))
#plt.yticks(np.arange(0, 81, 10))
a1.legend((p0[0], p1[0], p2[0], p3[0], p4[0] ), ('Glass', 'aluminium_frames', 'Silicon','Copper','Silver'))
f.tight_layout()
f.savefig(title_Method+' Fig_2x1_Yearly Virgin Material Needs by Scenario and Cumulatives_NREL2018.png', dpi=600)
print("Cumulative Virgin Needs by 2050 Million Tones by Scenario")
dfcumulations2050[['glass','silicon','silver','copper','aluminium_frames']].sum(axis=1)
```
# ### Bonus: Bifacial Trend Cumulative Virgin Needs (not plotted, just values)
```
name2 = 'bifacialTrend_high_[Tonnes]'
name0 = 'bifacialTrend_base_[Tonnes]'
cumulations2050 = {}
for ii in range(0, len(materials)):
matcum = []
matcum.append(UScum[keyw+materials[ii]+'_'+name0].loc[2050])
matcum.append(UScum[keyw+materials[ii]+'_'+name2].loc[2050])
cumulations2050[materials[ii]] = matcum
dfcumulations2050 = pd.DataFrame.from_dict(cumulations2050)
dfcumulations2050 = dfcumulations2050/1000000 # in Million Tonnes
print("Cumulative Virgin Needs by 2050 Million Tones by Scenario for Bifacial Trend")
dfcumulations2050[['glass','silicon','silver','copper','aluminium_frames']].sum(axis=1)
```
# ### Waste by year
```
plt.rcParams.update({'font.size': 15})
plt.rcParams['figure.figsize'] = (15, 8)
keyw='WasteAll_'
materials = ['glass', 'silicon', 'silver', 'copper', 'aluminium_frames']
f, (a0, a1) = plt.subplots(1, 2, gridspec_kw={'width_ratios': [3, 1]})
########################
# SUBPLOT 1
########################
#######################
# loop plotting over scenarios
name2 = 'Simulation1_high_[Tonnes]'
name0 = 'Simulation1_base_[Tonnes]'
# SCENARIO 1 ***************
modulemat = (USyearly[keyw+materials[0]+'_'+name0]+USyearly[keyw+materials[1]+'_'+name0]+
USyearly[keyw+materials[2]+'_'+name0]+USyearly[keyw+materials[3]+'_'+name0]+
USyearly[keyw+materials[4]+'_'+name0])
glassmat = (USyearly[keyw+materials[0]+'_'+name0])
modulemat = modulemat/1000000
glassmat = glassmat/1000000
a0.plot(USyearly.index, modulemat, 'k.', linewidth=5, label='S1: '+name0+' module mass')
a0.plot(USyearly.index, glassmat, 'k', linewidth=5, label='S1: '+name0+' glass mass only')
a0.fill_between(USyearly.index, glassmat, modulemat, color='k', alpha=0.3,
interpolate=True)
# SCENARIO 2 ***************
modulemat = (USyearly[keyw+materials[0]+'_'+name2]+USyearly[keyw+materials[1]+'_'+name2]+
USyearly[keyw+materials[2]+'_'+name2]+USyearly[keyw+materials[3]+'_'+name2]+
USyearly[keyw+materials[4]+'_'+name2])
glassmat = (USyearly[keyw+materials[0]+'_'+name2])
modulemat = modulemat/1000000
glassmat = glassmat/1000000
a0.plot(USyearly.index, modulemat, 'c.', linewidth=5, label='S2: '+name2+' module mass')
a0.plot(USyearly.index, glassmat, 'c', linewidth=5, label='S2: '+name2+' glass mass only')
a0.fill_between(USyearly.index, glassmat, modulemat, color='c', alpha=0.3,
interpolate=True)
a0.legend()
a0.set_title('Yearly Material Waste by Scenario')
a0.set_ylabel('Mass [Million Tonnes]')
a0.set_xlim([2020, 2050])
a0.set_xlabel('Years')
########################
# SUBPLOT 2
########################
#######################
# Calculate
cumulations2050 = {}
for ii in range(0, len(materials)):
matcum = []
matcum.append(UScum[keyw+materials[ii]+'_'+name0].loc[2050])
matcum.append(UScum[keyw+materials[ii]+'_'+name2].loc[2050])
cumulations2050[materials[ii]] = matcum
dfcumulations2050 = pd.DataFrame.from_dict(cumulations2050)
dfcumulations2050 = dfcumulations2050/1000000 # in Million Tonnes
dfcumulations2050['bottom1'] = dfcumulations2050['glass']
dfcumulations2050['bottom2'] = dfcumulations2050['bottom1']+dfcumulations2050['aluminium_frames']
dfcumulations2050['bottom3'] = dfcumulations2050['bottom2']+dfcumulations2050['silicon']
dfcumulations2050['bottom4'] = dfcumulations2050['bottom3']+dfcumulations2050['copper']
## Plot BARS Stuff
ind=np.arange(2)
width=0.35 # width of the bars.
p0 = a1.bar(ind, dfcumulations2050['glass'], width, color='c')
p1 = a1.bar(ind, dfcumulations2050['aluminium_frames'], width,
bottom=dfcumulations2050['bottom1'])
p2 = a1.bar(ind, dfcumulations2050['silicon'], width,
bottom=dfcumulations2050['bottom2'])
p3 = a1.bar(ind, dfcumulations2050['copper'], width,
bottom=dfcumulations2050['bottom3'])
p4 = a1.bar(ind, dfcumulations2050['silver'], width,
bottom=dfcumulations2050['bottom4'])
a1.yaxis.set_label_position("right")
a1.yaxis.tick_right()
a1.set_ylabel('Cumulative Waste by 2050 [Million Tonnes]')
a1.set_xlabel('Scenario')
a1.set_xticks(ind, ('S1', 'S2'))
#plt.yticks(np.arange(0, 81, 10))
a1.legend((p0[0], p1[0], p2[0], p3[0], p4[0] ), ('Glass', 'aluminium_frames', 'Silicon','Copper','Silver'))
f.tight_layout()
f.savefig(title_Method+' Fig_2x1_Yearly WASTE by Scenario and Cumulatives_NREL2018.png', dpi=600)
print("Cumulative Waste by 2050 Million Tones by case")
dfcumulations2050[['glass','silicon','silver','copper','aluminium_frames']].sum(axis=1)
plt.rcParams.update({'font.size': 15})
plt.rcParams['figure.figsize'] = (15, 8)
keyw='WasteEOL_'
materials = ['glass', 'silicon', 'silver', 'copper', 'aluminium_frames']
f, (a0, a1) = plt.subplots(1, 2, gridspec_kw={'width_ratios': [3, 1]})
########################
# SUBPLOT 1
########################
#######################
# loop plotting over scenarios
name2 = 'Simulation1_high_[Tonnes]'
name0 = 'Simulation1_base_[Tonnes]'
# SCENARIO 1 ***************
modulemat = (USyearly[keyw+materials[0]+'_'+name0]+USyearly[keyw+materials[1]+'_'+name0]+
USyearly[keyw+materials[2]+'_'+name0]+USyearly[keyw+materials[3]+'_'+name0]+
USyearly[keyw+materials[4]+'_'+name0])
glassmat = (USyearly[keyw+materials[0]+'_'+name0])
modulemat = modulemat/1000000
glassmat = glassmat/1000000
a0.plot(USyearly.index, modulemat, 'k.', linewidth=5, label='S1: '+name0+' module mass')
a0.plot(USyearly.index, glassmat, 'k', linewidth=5, label='S1: '+name0+' glass mass only')
a0.fill_between(USyearly.index, glassmat, modulemat, color='k', alpha=0.3,
interpolate=True)
# SCENARIO 2 ***************
modulemat = (USyearly[keyw+materials[0]+'_'+name2]+USyearly[keyw+materials[1]+'_'+name2]+
USyearly[keyw+materials[2]+'_'+name2]+USyearly[keyw+materials[3]+'_'+name2]+
USyearly[keyw+materials[4]+'_'+name2])
glassmat = (USyearly[keyw+materials[0]+'_'+name2])
modulemat = modulemat/1000000
glassmat = glassmat/1000000
a0.plot(USyearly.index, modulemat, 'c.', linewidth=5, label='S2: '+name2+' module mass')
a0.plot(USyearly.index, glassmat, 'c', linewidth=5, label='S2: '+name2+' glass mass only')
a0.fill_between(USyearly.index, glassmat, modulemat, color='c', alpha=0.3,
interpolate=True)
a0.legend()
a0.set_title('Yearly Material Waste by Scenario')
a0.set_ylabel('Mass [Million Tonnes]')
a0.set_xlim([2020, 2050])
a0.set_xlabel('Years')
########################
# SUBPLOT 2
########################
#######################
# Calculate
cumulations2050 = {}
for ii in range(0, len(materials)):
matcum = []
matcum.append(UScum[keyw+materials[ii]+'_'+name0].loc[2050])
matcum.append(UScum[keyw+materials[ii]+'_'+name2].loc[2050])
cumulations2050[materials[ii]] = matcum
dfcumulations2050 = pd.DataFrame.from_dict(cumulations2050)
dfcumulations2050 = dfcumulations2050/1000000 # in Million Tonnes
dfcumulations2050['bottom1'] = dfcumulations2050['glass']
dfcumulations2050['bottom2'] = dfcumulations2050['bottom1']+dfcumulations2050['aluminium_frames']
dfcumulations2050['bottom3'] = dfcumulations2050['bottom2']+dfcumulations2050['silicon']
dfcumulations2050['bottom4'] = dfcumulations2050['bottom3']+dfcumulations2050['copper']
## Plot BARS Stuff
ind=np.arange(2)
width=0.35 # width of the bars.
p0 = a1.bar(ind, dfcumulations2050['glass'], width, color='c')
p1 = a1.bar(ind, dfcumulations2050['aluminium_frames'], width,
bottom=dfcumulations2050['bottom1'])
p2 = a1.bar(ind, dfcumulations2050['silicon'], width,
bottom=dfcumulations2050['bottom2'])
p3 = a1.bar(ind, dfcumulations2050['copper'], width,
bottom=dfcumulations2050['bottom3'])
p4 = a1.bar(ind, dfcumulations2050['silver'], width,
bottom=dfcumulations2050['bottom4'])
a1.yaxis.set_label_position("right")
a1.yaxis.tick_right()
a1.set_ylabel('Cumulative EOL Only Waste by 2050 [Million Tonnes]')
a1.set_xlabel('Scenario')
a1.set_xticks(ind, ('S1', 'S2'))
#plt.yticks(np.arange(0, 81, 10))
a1.legend((p0[0], p1[0], p2[0], p3[0], p4[0] ), ('Glass', 'aluminium_frames', 'Silicon','Copper','Silver'))
f.tight_layout()
f.savefig(title_Method+' Fig_2x1_Yearly EOL Only WASTE by Scenario and Cumulatives_NREL2018.png', dpi=600)
print("Cumulative Eol Only Waste by 2050 Million Tones by case")
dfcumulations2050[['glass','silicon','silver','copper','aluminium_frames']].sum(axis=1)
```
| github_jupyter |
# Fun with FFT and sound files
Based on: https://realpython.com/python-scipy-fft/
Define a function for generating pure sine wave tones
```
import numpy as np
import matplotlib.pyplot as plt
SAMPLE_RATE = 44100 # Hertz
DURATION = 5 # Seconds
def generate_sine_wave(freq, sample_rate, duration):
x = np.linspace(0, duration, sample_rate * duration, endpoint=False)
frequencies = x * freq
# 2pi because np.sin takes radians
y = np.sin((2 * np.pi) * frequencies)
return x, y
# Generate a 2 hertz sine wave that lasts for 5 seconds
x, y = generate_sine_wave(2, SAMPLE_RATE, DURATION)
plt.plot(x, y)
plt.show()
```
Produce two tones, e.g. 400 Hz signal and a 4 kHz high-pitch noise
```
_, nice_tone = generate_sine_wave(400, SAMPLE_RATE, DURATION)
_, noise_tone = generate_sine_wave(4000, SAMPLE_RATE, DURATION)
noise_tone = noise_tone * 0.3
mixed_tone = nice_tone + noise_tone
#mixed_tone = noise_tone
```
For the purposes of storing the tones in an audio file, the amplitude needs to be normalized to the range of 16-bit integer
```
normalized_tone = np.int16((mixed_tone / mixed_tone.max()) * 32767)
plt.plot(normalized_tone[:1000])
plt.show()
```
Store the sound for playback
```
from scipy.io import wavfile as wf
# Remember SAMPLE_RATE = 44100 Hz is our playback rate
wf.write("mysinewave.wav", SAMPLE_RATE, normalized_tone)
```
Can also try to record the sound (NB: won't work on datahub !)
```
# import required libraries
%pip install sounddevice
import sounddevice as sd
print("Recording...")
# Start recorder with the given values
# of duration and sample frequency
recording = sd.rec(int(DURATION * SAMPLE_RATE), samplerate=SAMPLE_RATE, channels=1)
# Record audio for the given number of seconds
sd.wait()
print("Done")
# This will convert the NumPy array to an audio
# file with the given sampling frequency
wf.write("recording0.wav", 400, recording)
```
### Fourier transforms
Now try to transform the time stream into frequency space using FFT
```
from scipy.fft import fft, fftfreq
# Number of samples in normalized_tone
N = SAMPLE_RATE * DURATION
yf = fft(normalized_tone)
xf = fftfreq(N, 1 / SAMPLE_RATE)
print('Type of the output array: ',type(yf[0]))
print('Size of the input array: ',N)
print('Size of the Fourier transform: ',len(xf))
df = xf[1]-xf[0]
print(f'Width of the frequency bins: {df} Hz')
plt.plot(xf, np.abs(yf))
plt.xlabel('Frequency (Hz)')
plt.ylabel('FFT magnitude (a.u.)')
plt.show()
plt.figure()
plt.yscale('log')
plt.plot(xf, np.abs(yf))
plt.xlabel('Frequency (Hz)')
plt.ylabel('FFT magnitude (a.u.)')
plt.xlim(350,4050)
plt.show()
```
You notice that fft returns data for both positive and negative frequencies, produces the output array of the same size as input, and the output is a set of *complex* numbers. However, the information is reduntant: only half of the output values are unique. The magnitudes of the Fourier coefficients at negative frequencies are the same as at the corresponding positive frequencies. This is the property of the *real* Fourier transform, i.e. the transform applied to real-value signals. More precisely, $\mathrm{fft}(f)=\mathrm{fft}^*(-f)$
```
print(xf[1],xf[-1])
print(yf[1],yf[-1])
```
We can use this fact to save computational time and storage by computing only half of the Fourier coefficients:
```
from scipy.fft import rfft, rfftfreq
# Note the extra 'r' at the front
yf = rfft(normalized_tone)
xf = rfftfreq(N, 1 / SAMPLE_RATE)
print('Type of the output array: ',type(yf[0]))
print('Size of the input array: ',N)
print('Size of the Fourier transform: ',len(xf))
df = xf[1]-xf[0]
print(f'Width of the frequency bins: {df} Hz')
plt.plot(xf, np.abs(yf))
plt.xlabel('Frequency (Hz)')
plt.ylabel('FFT magnitude (a.u.)')
plt.show()
```
Now let's look at the Fourier transorm of a sound of a guitar string:
```
rate, data = wf.read("recording0.wav")
N=len(data)
print(rate, N)
time=np.arange(0, N)/rate
plt.plot(time, data)
plt.xlabel('time (sec)')
plt.ylabel('Sound a.u.)')
plt.show()
yf = rfft(data)
xf = rfftfreq(len(data), 1 / rate)
print('Type of the output array: ',type(yf[0]))
print('Size of the input array: ',len(data))
print('Size of the Fourier transform: ',len(xf))
df = xf[1]-xf[0]
print(f'Width of the frequency bins: {df} Hz')
plt.figure()
plt.loglog(xf, np.abs(yf))
plt.xlabel('Frequency (Hz)')
plt.ylabel('FFT magnitude (a.u.)')
plt.show()
plt.figure()
plt.plot(xf, np.abs(yf))
plt.yscale('log')
plt.xlim(100,2000)
plt.xlabel('Frequency (Hz)')
plt.ylabel('FFT magnitude (a.u.)')
plt.show()
```
| github_jupyter |
## PSO - Particle Swarm Optimisation
**About PSO -**
PSO is an biologically inspired meta-heuristic optimisation algorithm. It takes its inspiration from bird flocking or fish schooling. It works pretty good in practice. So let us code it up and optimise a function.
```
#dependencies
import random
import math
import copy # for array copying
import sys
```
### COST Function
So basically the function we are trying to optimise will become our cost function.
What cost functions we will see:
1. Sum of squares
2. Rastrigin's function
### Rastrigins function:
Rastrgins equation:

3-D Rendering

As you can see its a non-convex function with lot of local minimas (i.e multi-modal : lot of optimal solutions). It is a fairly diffucult problem for testing and we will test this out.
```
# lets code the rastrigins function
def error(position):
err = 0.0
for i in range(len(position)):
xi = position[i]
err += (xi * xi) - (10 * math.cos(2 * math.pi * xi))
err = 10*len(position) + err
return err
```
### Particle
A particle basically maintains the following params:
1. particle position
2. particle velocity
3. best position individual
4. best error individual
5. error individual
The action it can take when traversing over its search space looks like -
```
Update velocity -
w1*towards_current_direction(intertia) + w2*towards_self_best + w3*towards_swarm_best
Update position -
Add current_velocity to previous_postion to obtain new_velocity
```
Now suppose the particle finds some minima/maxima which is better than the global best it has to update the global value. So we have its fitness evaluation function -
```
evaluate fitness -
plug in current_position into test function to get where exactly you are that will give you the minima/maxima value
check against the global minima/maxima whether yours is better
assign value to global accordingly
```
```
# let us construct the class Particle
class Particle:
def __init__(self,x0):
self.position_i=[] # particle position
self.velocity_i=[] # particle velocity
self.pos_best_i=[] # best position individual
self.err_best_i=-1 # best error individual
self.err_i=-1 # error individual
for i in range(0,num_dimensions):
self.velocity_i.append(random.uniform(-1,1))
self.position_i.append(x0[i])
# evaluate current fitness
def evaluate(self,costFunc):
self.err_i=costFunc(self.position_i)
# check to see if the current position is an individual best
if self.err_i < self.err_best_i or self.err_best_i==-1:
self.pos_best_i=self.position_i
self.err_best_i=self.err_i
# update new particle velocity
def update_velocity(self,pos_best_g):
w=0.5 # constant inertia weight (how much to weigh the previous velocity)
c1=1 # cognative constant
c2=2 # social constant
for i in range(0,num_dimensions):
r1=random.uniform(-1,1)
r2=random.uniform(-1,1)
vel_cognitive=c1*r1*(self.pos_best_i[i]-self.position_i[i])
vel_social=c2*r2*(pos_best_g[i]-self.position_i[i])
self.velocity_i[i]=w*self.velocity_i[i]+vel_cognitive+vel_social
# update the particle position based off new velocity updates
def update_position(self,bounds):
for i in range(0,num_dimensions):
self.position_i[i]=self.position_i[i]+self.velocity_i[i]
# adjust maximum position if necessary
if self.position_i[i]>bounds[i][1]:
self.position_i[i]=bounds[i][1]
# adjust minimum position if neseccary
if self.position_i[i] < bounds[i][0]:
self.position_i[i]=bounds[i][0]
```
### __PSO__ (Particle Swarm Optimisation)
In particle swarm optimisation we
1. Initialise a swarm of particles to go on random exploration
2. for each particle we find whether the have discovered any new minima/maxima
3. The overall groups orientation or their velocities is guided to the global minimas
```
# Now let us define a class PSO
class PSO():
def __init__(self,costFunc,x0,bounds,num_particles,maxiter):
global num_dimensions
num_dimensions=len(x0)
err_best_g=-1 # best error for group
pos_best_g=[] # best position for group
# establish the swarm
swarm=[]
for i in range(0,num_particles):
swarm.append(Particle(x0))
# begin optimization loop
i=0
while i < maxiter:
#print i,err_best_g
# cycle through particles in swarm and evaluate fitness
for j in range(0,num_particles):
swarm[j].evaluate(costFunc)
# determine if current particle is the best (globally)
if swarm[j].err_i < err_best_g or err_best_g == -1:
pos_best_g=list(swarm[j].position_i)
err_best_g=float(swarm[j].err_i)
# cycle through swarm and update velocities and position
for j in range(0,num_particles):
swarm[j].update_velocity(pos_best_g)
swarm[j].update_position(bounds)
i+=1
# print final results
print ('\nFINAL:')
print (pos_best_g)
print (err_best_g)
%time
initial=[5,5] # initial starting location [x1,x2...]
bounds=[(-10,10),(-10,10)] # input bounds [(x1_min,x1_max),(x2_min,x2_max)...]
PSO(error,initial,bounds,num_particles=15,maxiter=30)
```
Now further on we will try to parallelize PSO algorithm
| github_jupyter |
```
import matplotlib
%matplotlib inline
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
import matplotlib.patches as mpatches
from sklearn.decomposition import PCA
sns.set_context('poster')
sns.set_style('white')
pd.options.mode.chained_assignment = None # default='warn'
import hdbscan
from collections import Counter
from collections import defaultdict
from numpy import random
def normalize(x, r):
M = np.divide(x, r)
M_norm = np.full_like(M, 0)
for i in range(np.shape(M)[0]):
rev = 1 - M[i, :]
if np.dot(M[i, :], M[i, :]) > np.dot(rev, rev):
M_norm[i, :] = rev
else:
M_norm[i, :] = M[i, :]
return M_norm
```
(Вспомогательная процедура, которая рисует легенду с обозначением цветов.)
```
def draw_legend(class_colours, classes, right=False):
recs = []
for i in range(0, len(classes)):
recs.append(mpatches.Rectangle((0,0), 1, 1, fc=class_colours[i]))
if right:
plt.legend(recs, classes, bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
else:
plt.legend(recs, classes)
```
SNP, встречающиеся в комбинации стрейнов
```
def plot_shared_snps(f_pca, f_0_pca, mask, names, draw_all=False):
combs = []
combs_nums = []
combinations = []
for m in mask:
if not draw_all:
if not (np.sum(m) > 1):
combinations.append(-1)
continue
cur = ""
for i in range(len(m)):
if m[i] == 1:
if cur != "":
cur += " + "
cur += names[i]
if cur == "":
cur = "none"
if cur not in combs:
combs.append(cur)
combs_nums.append(0)
combs_nums[combs.index(cur)] += 1
combinations.append(combs.index(cur))
df = pd.DataFrame({'pc1':f_pca[:, 0], 'pc2':f_pca[:, 1], 'combination':combinations})
df_valid = df.loc[df['combination'] != -1]
# reoder combinations by sizes of groups
order = sorted(zip(combs_nums, combs, range(12)), reverse=True)
new_comb_order = [0] * (2 ** len(mask[0]))
new_comb_names = []
for i in range(len(order)):
old_order = order[i][2]
new_comb_order[old_order] = i
new_comb_names.append('{:5d}'.format(order[i][0]) + ' ' + order[i][1])
#new_comb_names.append(order[i][1])
for i in df_valid.index:
df_valid.loc[i, "combination"] = new_comb_order[df_valid.loc[i, "combination"]]
# Kelly’s 20 (except the first 2) Colours of Maximum Contrast
colors = ['yellow', 'purple', 'orange', '#96cde6', 'red', '#c0bd7f', '#5fa641', '#d485b2',
'#4277b6', '#df8461', '#463397', '#e1a11a', '#91218c', '#e8e948', '#7e1510',
'#92ae31', '#6f340d', '#d32b1e', '#2b3514']
color_palette = sns.color_palette(colors)
cluster_colors = [color_palette[x] for x in df_valid["combination"]]
plt.figure(figsize=(15, 8))
ax = plt.gca()
ax.set_aspect('equal')
plt.xlabel("PC 1")
plt.ylabel("PC 2")
plt.scatter(f_0_pca[:, 0], f_0_pca[:, 1], s=40, linewidth=0, c="grey", alpha=0.2);
plt.scatter(df_valid["pc1"], df_valid["pc2"], s=40, linewidth=0, c=cluster_colors);
#plt.title("[Sharon et al, 2013]")
draw_legend(color_palette, new_comb_names, right=True)
def clusterization(f, pca=True, num_of_comp=2):
if pca:
f_pca = PCA(n_components = num_of_comp).fit(f).transform(f)
cur_f = f_pca
else:
cur_f = f
f_pca = PCA(n_components = 2).fit(f).transform(f)
#N = (nt) (len(f) * 0.005)
#print(N)
N = 100
clusterer = hdbscan.HDBSCAN(min_cluster_size=N, min_samples=1).fit(cur_f)
plt.figure(figsize=(15, 8))
ax = plt.gca()
ax.set_aspect('equal')
plt.xlabel("PC 1")
plt.ylabel("PC 2")
if pca:
plt.title("Clustering %s primary components" % num_of_comp)
else:
plt.title("Clustering initial frequencies")
color_palette = sns.color_palette("Set2", 20)
cluster_colors = [color_palette[x] if x >= 0
else (0.5, 0.5, 0.5)
for x in clusterer.labels_]
cluster_member_colors = [sns.desaturate(x, p) for x, p in
zip(cluster_colors, clusterer.probabilities_)]
plt.scatter(f_pca[:, 0], f_pca[:, 1], s=40, linewidth=0, c=cluster_member_colors, alpha=0.3);
sizes_of_classes = Counter(clusterer.labels_)
print(sizes_of_classes.get(-1, 0), "outliers\n")
labels = [str(x) + ' - ' + str(sizes_of_classes[x]) for x in range(max(clusterer.labels_)+1)]
draw_legend(color_palette, labels, right=True)
print("Medians in clusters:")
for i in range(max(clusterer.labels_)+1):
f_with_labels = f.copy()
f_with_labels = np.hstack([f_with_labels, clusterer.labels_.reshape(len(f_with_labels),1)])
col = f_with_labels[:, -1]
idx = (col == i)
print(i, np.round(np.median(f_with_labels[idx,:-1], axis=0), 2))
```
# Infant Gut, выровненный на Strain 1
(Преобразование не делаем, так как референс есть в данных)
##### Частоты стрейнов в Infant Gut:
strain1 0.73 0.74 0.04 0.13 0.17 0.04 0.32 0.75 0.30 0.20 0.0
strain3 0.24 0.20 0.95 0.80 0.80 0.93 0.52 0.19 0.64 0.65 1.0
strain4 0.03 0.06 0.02 0.07 0.03 0.02 0.16 0.06 0.06 0.15 0.0
```
def filter_by_coverage(cur_r, bad_percent, bad_samples):
def filter_row(row):
num_of_samples = len(row)
valid = np.sum(np.array(([(min_coverage < row) & (row < max_coverage)])))
return num_of_samples - valid <= bad_samples
min_coverage = np.percentile(cur_r, bad_percent, axis=0)
max_coverage = np.percentile(cur_r, 100-bad_percent, axis=0)
good_coverage = np.array([filter_row(row) for row in cur_r])
return good_coverage
r_0 = np.genfromtxt("infant_gut_pure_STRAIN1/matrices/R_all", dtype=int, delimiter=' ')
x_0 = np.genfromtxt("infant_gut_pure_STRAIN1/matrices/X_all", dtype=int, delimiter=' ')
print(len(r_0))
names = ["strain 1", "strain 3", "strain 4"]
r_0 = np.delete(r_0, [i for i in range(len(names))], axis=1)
x_0 = np.delete(x_0, [i for i in range(len(names))], axis=1)
Ncut = 6
print("Delete zero and almost zero profiles:")
good_ind = [i for i in range(np.shape(x_0)[0])
if not ((np.abs(r_0[i, :] - x_0[i, :]) <= Ncut).all() or (x_0[i, :] <= Ncut).all())]
print(len(good_ind), "remained")
x_0 = x_0[good_ind, :]
r_0 = r_0[good_ind, :]
good_coverage = filter_by_coverage(r_0, 15, 2)
r_0 = r_0[good_coverage, :]
x_0 = x_0[good_coverage, :]
print(len(r_0))
r = np.genfromtxt("infant_gut_pure_STRAIN1/matrices/R_filtered", dtype=int, delimiter=' ')
x = np.genfromtxt("infant_gut_pure_STRAIN1/matrices/X_filtered", dtype=int, delimiter=' ')
print("%s sites" % len(r))
mask = np.genfromtxt("infant_gut_pure_STRAIN1/clomial_results/genotypes_3.txt",
dtype=float, delimiter=' ', skip_header=1)
mask = np.delete(mask, [0], axis=1)
mask = np.rint(mask)
names = ["C1", "C2", "C3"]
```
Рисуем получившиеся фичи на главных компонентах.
```
f_0 = np.divide(x_0, r_0)
f_0_pca = PCA(n_components = 2).fit(f_0).transform(f_0)
f = np.divide(x, r)
f_pca = PCA(n_components = 2).fit(f_0).transform(f)
plot_shared_snps(f_pca, f_0_pca, mask, names, draw_all=True)
```
# Infant Gut, выровненный на референс NCBI + подмешали референс
(Преобразование делаем)
```
r_0 = np.genfromtxt("infant_gut/infant_gut_pure_without_ref/matrices/R_all", dtype=int, delimiter=' ')
x_0 = np.genfromtxt("infant_gut/infant_gut_pure_without_ref/matrices/X_all", dtype=int, delimiter=' ')
print(len(r_0))
names = ["strain 1", "strain 3", "strain 4"]
r_0 = np.delete(r_0, [i for i in range(len(names))], axis=1)
x_0 = np.delete(x_0, [i for i in range(len(names))], axis=1)
Ncut = 6
print("Delete zero and almost zero profiles:")
good_ind = [i for i in range(np.shape(x_0)[0])
if not ((np.abs(r_0[i, :] - x_0[i, :]) <= Ncut).all() or (x_0[i, :] <= Ncut).all())]
print(len(good_ind), "remained")
x_0 = x_0[good_ind, :]
r_0 = r_0[good_ind, :]
good_coverage = filter_by_coverage(r_0, 15, 2)
r_0 = r_0[good_coverage, :]
x_0 = x_0[good_coverage, :]
print(len(r_0))
r = np.genfromtxt("infant_gut/infant_gut_pure_without_ref/matrices/R_filtered", dtype=int, delimiter=' ')
x = np.genfromtxt("infant_gut/infant_gut_pure_without_ref/matrices/X_filtered", dtype=int, delimiter=' ')
r = np.delete(r, [0], axis=1)
r = r / 1.1
r = np.rint(r)
r = r.astype(int)
x = np.delete(x, [0], axis=1)
print("%s sites" % len(r))
mask = np.genfromtxt("infant_gut/infant_gut_pure_without_ref/clomial_results/genotypes_4.txt",
dtype=float, delimiter=' ', skip_header=1)
mask = np.delete(mask, [0, 1], axis=1)
mask = np.rint(mask)
names = ["C2", "C3", "C4"]
```
Рисуем получившиеся фичи на главных компонентах.
```
f_0 = np.divide(x_0, r_0)
f_0_pca = PCA(n_components = 2).fit(f_0).transform(f_0)
f = np.divide(x, r)
f_pca = PCA(n_components = 2).fit(f_0).transform(f)
plot_shared_snps(f_pca, f_0_pca, mask, names, draw_all=True)
f_0 = normalize(x_0, r_0)
f_0_pca = PCA(n_components = 2).fit(f_0).transform(f_0)
f = normalize(x, r)
f_pca = PCA(n_components = 2).fit(f_0).transform(f)
plot_shared_snps(f_pca, f_0_pca, mask, names, draw_all=True)
```
| github_jupyter |
```
import sys, os
if 'google.colab' in sys.modules:
# https://github.com/yandexdataschool/Practical_RL/issues/256
!pip install tensorflow-gpu==1.13.1
if not os.path.exists('.setup_complete'):
!wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/spring20/setup_colab.sh -O- | bash
!wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/spring20/week07_seq2seq/basic_model_tf.py
!wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/spring20/week07_seq2seq/he-pron-wiktionary.txt
!wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/spring20/week07_seq2seq/main_dataset.txt
!wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/spring20/week07_seq2seq/voc.py
!touch .setup_complete
# This code creates a virtual display to draw game images on.
# It will have no effect if your machine has a monitor.
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY")) == 0:
!bash ../xvfb start
os.environ['DISPLAY'] = ':1'
```
## Reinforcement Learning for seq2seq
This time we'll solve a problem of transribing hebrew words in english, also known as g2p (grapheme2phoneme)
* word (sequence of letters in source language) -> translation (sequence of letters in target language)
Unlike what most deep learning practitioners do, we won't only train it to maximize likelihood of correct translation, but also employ reinforcement learning to actually teach it to translate with as few errors as possible.
### About the task
One notable property of Hebrew is that it's consonant language. That is, there are no wovels in the written language. One could represent wovels with diacritics above consonants, but you don't expect people to do that in everyay life.
Therefore, some hebrew characters will correspond to several english letters and others - to none, so we should use encoder-decoder architecture to figure that out.
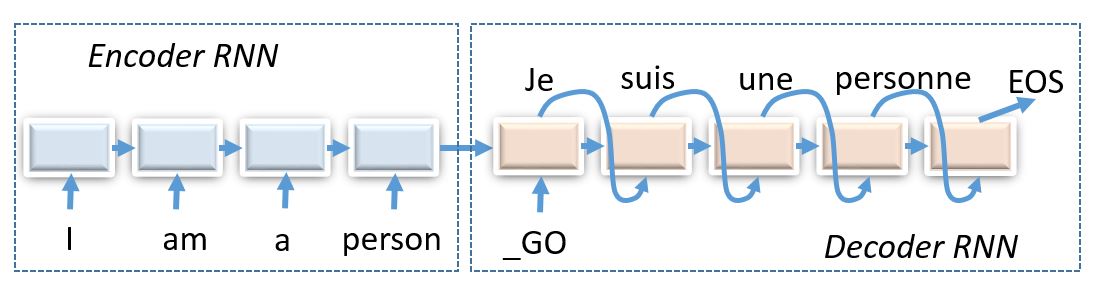
_(img: esciencegroup.files.wordpress.com)_
Encoder-decoder architectures are about converting anything to anything, including
* Machine translation and spoken dialogue systems
* [Image captioning](http://mscoco.org/dataset/#captions-challenge2015) and [image2latex](https://openai.com/requests-for-research/#im2latex) (convolutional encoder, recurrent decoder)
* Generating [images by captions](https://arxiv.org/abs/1511.02793) (recurrent encoder, convolutional decoder)
* Grapheme2phoneme - convert words to transcripts
We chose simplified __Hebrew->English__ machine translation for words and short phrases (character-level), as it is relatively quick to train even without a gpu cluster.
```
# If True, only translates phrases shorter than 20 characters (way easier).
EASY_MODE = True
# Useful for initial coding.
# If false, works with all phrases (please switch to this mode for homework assignment)
MODE = "he-to-en" # way we translate. Either "he-to-en" or "en-to-he"
# maximal length of _generated_ output, does not affect training
MAX_OUTPUT_LENGTH = 50 if not EASY_MODE else 20
REPORT_FREQ = 100 # how often to evaluate validation score
```
### Step 1: preprocessing
We shall store dataset as a dictionary
`{ word1:[translation1,translation2,...], word2:[...],...}`.
This is mostly due to the fact that many words have several correct translations.
We have implemented this thing for you so that you can focus on more interesting parts.
__Attention python2 users!__ You may want to cast everything to unicode later during homework phase, just make sure you do it _everywhere_.
```
import numpy as np
from collections import defaultdict
word_to_translation = defaultdict(list) # our dictionary
bos = '_'
eos = ';'
with open("main_dataset.txt") as fin:
for line in fin:
en, he = line[:-1].lower().replace(bos, ' ').replace(eos,
' ').split('\t')
word, trans = (he, en) if MODE == 'he-to-en' else (en, he)
if len(word) < 3:
continue
if EASY_MODE:
if max(len(word), len(trans)) > 20:
continue
word_to_translation[word].append(trans)
print("size = ", len(word_to_translation))
# get all unique lines in source language
all_words = np.array(list(word_to_translation.keys()))
# get all unique lines in translation language
all_translations = np.array(
[ts for all_ts in word_to_translation.values() for ts in all_ts])
```
### split the dataset
We hold out 10% of all words to be used for validation.
```
from sklearn.model_selection import train_test_split
train_words, test_words = train_test_split(
all_words, test_size=0.1, random_state=42)
```
### Building vocabularies
We now need to build vocabularies that map strings to token ids and vice versa. We're gonna need these fellas when we feed training data into model or convert output matrices into english words.
```
from voc import Vocab
inp_voc = Vocab.from_lines(''.join(all_words), bos=bos, eos=eos, sep='')
out_voc = Vocab.from_lines(''.join(all_translations), bos=bos, eos=eos, sep='')
# Here's how you cast lines into ids and backwards.
batch_lines = all_words[:5]
batch_ids = inp_voc.to_matrix(batch_lines)
batch_lines_restored = inp_voc.to_lines(batch_ids)
print("lines")
print(batch_lines)
print("\nwords to ids (0 = bos, 1 = eos):")
print(batch_ids)
print("\nback to words")
print(batch_lines_restored)
```
Draw word/translation length distributions to estimate the scope of the task.
```
import matplotlib.pyplot as plt
%matplotlib inline
plt.figure(figsize=[8, 4])
plt.subplot(1, 2, 1)
plt.title("words")
plt.hist(list(map(len, all_words)), bins=20)
plt.subplot(1, 2, 2)
plt.title('translations')
plt.hist(list(map(len, all_translations)), bins=20)
```
### Step 3: deploy encoder-decoder (1 point)
__assignment starts here__
Our architecture consists of two main blocks:
* Encoder reads words character by character and outputs code vector (usually a function of last RNN state)
* Decoder takes that code vector and produces translations character by character
Than it gets fed into a model that follows this simple interface:
* __`model.symbolic_translate(inp, **flags) -> out, logp`__ - takes symbolic int32 matrix of hebrew words, produces output tokens sampled from the model and output log-probabilities for all possible tokens at each tick.
* if given flag __`greedy=True`__, takes most likely next token at each iteration. Otherwise samples with next token probabilities predicted by model.
* __`model.symbolic_score(inp, out, **flags) -> logp`__ - takes symbolic int32 matrices of hebrew words and their english translations. Computes the log-probabilities of all possible english characters given english prefices and hebrew word.
* __`model.weights`__ - weights from all model layers [a list of variables]
That's all! It's as hard as it gets. With those two methods alone you can implement all kinds of prediction and training.
```
import tensorflow as tf
tf.reset_default_graph()
s = tf.InteractiveSession()
# ^^^ if you get "variable *** already exists": re-run this cell again
from basic_model_tf import BasicTranslationModel
model = BasicTranslationModel('model', inp_voc, out_voc,
emb_size=64, hid_size=128)
s.run(tf.global_variables_initializer())
# Play around with symbolic_translate and symbolic_score
inp = tf.placeholder_with_default(np.random.randint(
0, 10, [3, 5], dtype='int32'), [None, None])
out = tf.placeholder_with_default(np.random.randint(
0, 10, [3, 5], dtype='int32'), [None, None])
# translate inp (with untrained model)
sampled_out, logp = model.symbolic_translate(inp, greedy=False)
print("\nSymbolic_translate output:\n", sampled_out, logp)
print("\nSample translations:\n", s.run(sampled_out))
# score logp(out | inp) with untrained input
logp = model.symbolic_score(inp, out)
print("\nSymbolic_score output:\n", logp)
print("\nLog-probabilities (clipped):\n", s.run(logp)[:, :2, :5])
# Prepare any operations you want here
input_sequence = tf.placeholder('int32', [None, None])
greedy_translations, logp = <YOUR CODE: build symbolic translations with greedy = True>
def translate(lines):
"""
You are given a list of input lines.
Make your neural network translate them.
:return: a list of output lines
"""
# Convert lines to a matrix of indices
lines_ix = <YOUR CODE>
# Compute translations in form of indices
trans_ix = s.run(greedy_translations, { <YOUR CODE: feed_dict> })
# Convert translations back into strings
return out_voc.to_lines(trans_ix)
print("Sample inputs:", all_words[:3])
print("Dummy translations:", translate(all_words[:3]))
assert isinstance(greedy_translations,
tf.Tensor) and greedy_translations.dtype.is_integer, "trans must be a tensor of integers (token ids)"
assert translate(all_words[:3]) == translate(
all_words[:3]), "make sure translation is deterministic (use greedy=True and disable any noise layers)"
assert type(translate(all_words[:3])) is list and (type(translate(all_words[:1])[0]) is str or type(
translate(all_words[:1])[0]) is unicode), "translate(lines) must return a sequence of strings!"
print("Tests passed!")
```
### Scoring function
LogLikelihood is a poor estimator of model performance.
* If we predict zero probability once, it shouldn't ruin entire model.
* It is enough to learn just one translation if there are several correct ones.
* What matters is how many mistakes model's gonna make when it translates!
Therefore, we will use minimal Levenshtein distance. It measures how many characters do we need to add/remove/replace from model translation to make it perfect. Alternatively, one could use character-level BLEU/RougeL or other similar metrics.
The catch here is that Levenshtein distance is not differentiable: it isn't even continuous. We can't train our neural network to maximize it by gradient descent.
```
import editdistance # !pip install editdistance
def get_distance(word, trans):
"""
A function that takes word and predicted translation
and evaluates (Levenshtein's) edit distance to closest correct translation
"""
references = word_to_translation[word]
assert len(references) != 0, "wrong/unknown word"
return min(editdistance.eval(trans, ref) for ref in references)
def score(words, bsize=100):
"""a function that computes levenshtein distance for bsize random samples"""
assert isinstance(words, np.ndarray)
batch_words = np.random.choice(words, size=bsize, replace=False)
batch_trans = translate(batch_words)
distances = list(map(get_distance, batch_words, batch_trans))
return np.array(distances, dtype='float32')
# should be around 5-50 and decrease rapidly after training :)
[score(test_words, 10).mean() for _ in range(5)]
```
## Step 2: Supervised pre-training
Here we define a function that trains our model through maximizing log-likelihood a.k.a. minimizing crossentropy.
```
# import utility functions
from basic_model_tf import initialize_uninitialized, infer_length, infer_mask, select_values_over_last_axis
class supervised_training:
# variable for inputs and correct answers
input_sequence = tf.placeholder('int32', [None, None])
reference_answers = tf.placeholder('int32', [None, None])
# Compute log-probabilities of all possible tokens at each step. Use model interface.
logprobs_seq = <YOUR CODE>
# compute mean crossentropy
crossentropy = - select_values_over_last_axis(logprobs_seq, reference_answers)
mask = infer_mask(reference_answers, out_voc.eos_ix)
loss = tf.reduce_sum(crossentropy * mask)/tf.reduce_sum(mask)
# Build weights optimizer. Use model.weights to get all trainable params.
train_step = <YOUR CODE>
# intialize optimizer params while keeping model intact
initialize_uninitialized(s)
```
Actually run training on minibatches
```
import random
def sample_batch(words, word_to_translation, batch_size):
"""
sample random batch of words and random correct translation for each word
example usage:
batch_x,batch_y = sample_batch(train_words, word_to_translations,10)
"""
# choose words
batch_words = np.random.choice(words, size=batch_size)
# choose translations
batch_trans_candidates = list(map(word_to_translation.get, batch_words))
batch_trans = list(map(random.choice, batch_trans_candidates))
return inp_voc.to_matrix(batch_words), out_voc.to_matrix(batch_trans)
bx, by = sample_batch(train_words, word_to_translation, batch_size=3)
print("Source:")
print(bx)
print("Target:")
print(by)
from IPython.display import clear_output
from tqdm import tqdm, trange # or use tqdm_notebook,tnrange
loss_history = []
editdist_history = []
for i in trange(25000):
bx, by = sample_batch(train_words, word_to_translation, 32)
feed_dict = {
supervised_training.input_sequence: bx,
supervised_training.reference_answers: by
}
loss, _ = s.run([supervised_training.loss,
supervised_training.train_step], feed_dict)
loss_history.append(loss)
if (i+1) % REPORT_FREQ == 0:
clear_output(True)
current_scores = score(test_words)
editdist_history.append(current_scores.mean())
plt.figure(figsize=(12, 4))
plt.subplot(131)
plt.title('train loss / traning time')
plt.plot(loss_history)
plt.grid()
plt.subplot(132)
plt.title('val score distribution')
plt.hist(current_scores, bins=20)
plt.subplot(133)
plt.title('val score / traning time')
plt.plot(editdist_history)
plt.grid()
plt.show()
print("llh=%.3f, mean score=%.3f" %
(np.mean(loss_history[-10:]), np.mean(editdist_history[-10:])))
# Note: it's okay if loss oscillates up and down as long as it gets better on average over long term (e.g. 5k batches)
for word in train_words[:10]:
print("%s -> %s" % (word, translate([word])[0]))
test_scores = []
for start_i in trange(0, len(test_words), 32):
batch_words = test_words[start_i:start_i+32]
batch_trans = translate(batch_words)
distances = list(map(get_distance, batch_words, batch_trans))
test_scores.extend(distances)
print("Supervised test score:", np.mean(test_scores))
```
## Preparing for reinforcement learning (2 points)
First we need to define loss function as a custom tf operation.
The simple way to do so is through `tensorflow.py_func` wrapper.
```
def my_func(x):
# x will be a numpy array with the contents of the placeholder below
return np.sinh(x)
inp = tf.placeholder(tf.float32)
y = tf.py_func(my_func, [inp], tf.float32)
```
__Your task__ is to implement `_compute_levenshtein` function that takes matrices of words and translations, along with input masks, then converts those to actual words and phonemes and computes min-levenshtein via __get_distance__ function above.
```
def _compute_levenshtein(words_ix, trans_ix):
"""
A custom tensorflow operation that computes levenshtein loss for predicted trans.
Params:
- words_ix - a matrix of letter indices, shape=[batch_size,word_length]
- words_mask - a matrix of zeros/ones,
1 means "word is still not finished"
0 means "word has already finished and this is padding"
- trans_mask - a matrix of output letter indices, shape=[batch_size,translation_length]
- trans_mask - a matrix of zeros/ones, similar to words_mask but for trans_ix
Please implement the function and make sure it passes tests from the next cell.
"""
# convert words to strings
words = <YOUR CODE: restore words (a list of strings) from words_ix. Use vocab>
assert type(words) is list and type(
words[0]) is str and len(words) == len(words_ix)
# convert translations to lists
translations = <YOUR CODE: restore trans (a list of lists of phonemes) from trans_ix
assert type(translations) is list and type(
translations[0]) is str and len(translations) == len(trans_ix)
# computes levenstein distances. can be arbitrary python code.
distances = <YOUR CODE: apply get_distance to each pair of [words, translations]>
assert type(distances) in (list, tuple, np.ndarray) and len(
distances) == len(words_ix)
distances = np.array(list(distances), dtype='float32')
return distances
def compute_levenshtein(words_ix, trans_ix):
out = tf.py_func(_compute_levenshtein, [words_ix, trans_ix, ], tf.float32)
out.set_shape([None])
return tf.stop_gradient(out)
```
Simple test suite to make sure your implementation is correct. Hint: if you run into any bugs, feel free to use print from inside _compute_levenshtein.
```
# test suite
# sample random batch of (words, correct trans, wrong trans)
batch_words = np.random.choice(train_words, size=100)
batch_trans = list(map(random.choice, map(
word_to_translation.get, batch_words)))
batch_trans_wrong = np.random.choice(all_translations, size=100)
batch_words_ix = tf.constant(inp_voc.to_matrix(batch_words))
batch_trans_ix = tf.constant(out_voc.to_matrix(batch_trans))
batch_trans_wrong_ix = tf.constant(out_voc.to_matrix(batch_trans_wrong))
# assert compute_levenshtein is zero for ideal translations
correct_answers_score = compute_levenshtein(
batch_words_ix, batch_trans_ix).eval()
assert np.all(correct_answers_score ==
0), "a perfect translation got nonzero levenshtein score!"
print("Everything seems alright!")
# assert compute_levenshtein matches actual scoring function
wrong_answers_score = compute_levenshtein(
batch_words_ix, batch_trans_wrong_ix).eval()
true_wrong_answers_score = np.array(
list(map(get_distance, batch_words, batch_trans_wrong)))
assert np.all(wrong_answers_score ==
true_wrong_answers_score), "for some word symbolic levenshtein is different from actual levenshtein distance"
print("Everything seems alright!")
```
Once you got it working...
* You may now want to __remove/comment asserts__ from function code for a slight speed-up.
* There's a more detailed tutorial on custom tensorflow ops: [`py_func`](https://www.tensorflow.org/api_docs/python/tf/py_func), [`low-level`](https://www.tensorflow.org/api_docs/python/tf/py_func).
## 3. Self-critical policy gradient (2 points)
In this section you'll implement algorithm called self-critical sequence training (here's an [article](https://arxiv.org/abs/1612.00563)).
The algorithm is a vanilla policy gradient with a special baseline.
$$ \nabla J = E_{x \sim p(s)} E_{y \sim \pi(y|x)} \nabla log \pi(y|x) \cdot (R(x,y) - b(x)) $$
Here reward R(x,y) is a __negative levenshtein distance__ (since we minimize it). The baseline __b(x)__ represents how well model fares on word __x__.
In practice, this means that we compute baseline as a score of greedy translation, $b(x) = R(x,y_{greedy}(x)) $.

Luckily, we already obtained the required outputs: `model.greedy_translations, model.greedy_mask` and we only need to compute levenshtein using `compute_levenshtein` function.
```
class trainer:
input_sequence = tf.placeholder('int32', [None, None])
# use model to __sample__ symbolic translations given input_sequence
sample_translations, sample_logp = <YOUR CODE>
# use model to __greedy__ symbolic translations given input_sequence
greedy_translations, greedy_logp = <YOUR CODE>
rewards = - compute_levenshtein(input_sequence, sample_translations)
# compute __negative__ levenshtein for greedy mode
baseline = <YOUR CODE>
# compute advantage using rewards and baseline
advantage = <YOUR CODE: compute advantage>
assert advantage.shape.ndims == 1, "advantage must be of shape [batch_size]"
# compute log_pi(a_t|s_t), shape = [batch, seq_length]
logprobs_phoneme = <YOUR CODE>
# ^-- hint: look at how crossentropy is implemented in supervised learning loss above
# mind the sign - this one should not be multiplied by -1 :)
# Compute policy gradient
# or rather surrogate function who's gradient is policy gradient
J = logprobs_phoneme*advantage[:, None]
mask = infer_mask(sample_translations, out_voc.eos_ix)
loss = - tf.reduce_sum(J*mask) / tf.reduce_sum(mask)
# regularize with negative entropy. Don't forget the sign!
# note: for entropy you need probabilities for all tokens (sample_logp), not just phoneme_logprobs
entropy = <compute entropy matrix of shape[batch, seq_length], H = -sum(p*log_p), don't forget the sign!>
# hint: you can get sample probabilities from sample_logp using math :)
assert entropy.shape.ndims == 2, "please make sure elementwise entropy is of shape [batch,time]"
loss -= 0.01*tf.reduce_sum(entropy*mask) / tf.reduce_sum(mask)
# compute weight updates, clip by norm
grads = tf.gradients(loss, model.weights)
grads = tf.clip_by_global_norm(grads, 50)[0]
train_step = tf.train.AdamOptimizer(
learning_rate=1e-5).apply_gradients(zip(grads, model.weights,))
initialize_uninitialized()
```
# Policy gradient training
```
for i in trange(100000):
bx = sample_batch(train_words, word_to_translation, 32)[0]
pseudo_loss, _ = s.run([trainer.loss, trainer.train_step], {
trainer.input_sequence: bx})
loss_history.append(
pseudo_loss
)
if (i+1) % REPORT_FREQ == 0:
clear_output(True)
current_scores = score(test_words)
editdist_history.append(current_scores.mean())
plt.figure(figsize=(8, 4))
plt.subplot(121)
plt.title('val score distribution')
plt.hist(current_scores, bins=20)
plt.subplot(122)
plt.title('val score / traning time')
plt.plot(editdist_history)
plt.grid()
plt.show()
print("J=%.3f, mean score=%.3f" %
(np.mean(loss_history[-10:]), np.mean(editdist_history[-10:])))
```
### Results
```
for word in train_words[:10]:
print("%s -> %s" % (word, translate([word])[0]))
test_scores = []
for start_i in trange(0, len(test_words), 32):
batch_words = test_words[start_i:start_i+32]
batch_trans = translate(batch_words)
distances = list(map(get_distance, batch_words, batch_trans))
test_scores.extend(distances)
print("Supervised test score:", np.mean(test_scores))
# ^^ If you get Out Of Memory, please replace this with batched computation
```
## Step 6: Make it actually work (5++ pts)
<img src=https://github.com/yandexdataschool/Practical_RL/raw/master/yet_another_week/_resource/do_something_scst.png width=400>
In this section we want you to finally __restart with EASY_MODE=False__ and experiment to find a good model/curriculum for that task.
We recommend you to start with the following architecture
```
encoder---decoder
P(y|h)
^
LSTM -> LSTM
^ ^
biLSTM -> LSTM
^ ^
input y_prev
```
__Note:__ you can fit all 4 state tensors of both LSTMs into a in a single state - just assume that it contains, for example, [h0, c0, h1, c1] - pack it in encode and update in decode.
Here are some cool ideas on what you can do then.
__General tips & tricks:__
* In some tensorflow versions and for some layers, it is required that each rnn/gru/lstm cell gets it's own `tf.variable_scope(unique_name, reuse=False)`.
* Otherwise it will complain about wrong tensor sizes because it tries to reuse weights from one rnn to the other.
* You will likely need to adjust pre-training time for such a network.
* Supervised pre-training may benefit from clipping gradients somehow.
* SCST may indulge a higher learning rate in some cases and changing entropy regularizer over time.
* It's often useful to save pre-trained model parameters to not re-train it every time you want new policy gradient parameters.
* When leaving training for nighttime, try setting REPORT_FREQ to a larger value (e.g. 500) not to waste time on it.
__Formal criteria:__
To get 5 points we want you to build an architecture that:
* _doesn't consist of single GRU_
* _works better_ than single GRU baseline.
* We also want you to provide either learning curve or trained model, preferably both
* ... and write a brief report or experiment log describing what you did and how it fared.
### Attention
There's more than one way to connect decoder to encoder
* __Vanilla:__ layer_i of encoder last state goes to layer_i of decoder initial state
* __Every tick:__ feed encoder last state _on every iteration_ of decoder.
* __Attention:__ allow decoder to "peek" at one (or several) positions of encoded sequence on every tick.
The most effective (and cool) of those is, of course, attention.
You can read more about attention [in this nice blog post](https://distill.pub/2016/augmented-rnns/). The easiest way to begin is to use "soft" attention with "additive" or "dot-product" intermediate layers.
__Tips__
* Model usually generalizes better if you no longer allow decoder to see final encoder state
* Once your model made it through several epochs, it is a good idea to visualize attention maps to understand what your model has actually learned
* There's more stuff [here](https://github.com/yandexdataschool/Practical_RL/blob/master/week8_scst/bonus.ipynb)
* If you opted for hard attention, we recommend [gumbel-softmax](https://blog.evjang.com/2016/11/tutorial-categorical-variational.html) instead of sampling. Also please make sure soft attention works fine before you switch to hard.
### UREX
* This is a way to improve exploration in policy-based settings. The main idea is that you find and upweight under-appreciated actions.
* Here's [video](https://www.youtube.com/watch?v=fZNyHoXgV7M&feature=youtu.be&t=3444)
and an [article](https://arxiv.org/abs/1611.09321).
* You may want to reduce batch size 'cuz UREX requires you to sample multiple times per source sentence.
* Once you got it working, try using experience replay with importance sampling instead of (in addition to) basic UREX.
### Some additional ideas:
* (advanced deep learning) It may be a good idea to first train on small phrases and then adapt to larger ones (a.k.a. training curriculum).
* (advanced nlp) You may want to switch from raw utf8 to something like unicode or even syllables to make task easier.
* (advanced nlp) Since hebrew words are written __with vowels omitted__, you may want to use a small Hebrew vowel markup dataset at `he-pron-wiktionary.txt`.
### Bonus hints: [here](https://github.com/yandexdataschool/Practical_RL/blob/master/week8_scst/bonus.ipynb)
```
assert not EASY_MODE, "make sure you set EASY_MODE = False at the top of the notebook."
```
`[your report/log here or anywhere you please]`
__Contributions:__ This notebook is brought to you by
* Yandex [MT team](https://tech.yandex.com/translate/)
* Denis Mazur ([DeniskaMazur](https://github.com/DeniskaMazur)), Oleg Vasilev ([Omrigan](https://github.com/Omrigan/)), Dmitry Emelyanenko ([TixFeniks](https://github.com/tixfeniks)) and Fedor Ratnikov ([justheuristic](https://github.com/justheuristic/))
* Dataset is parsed from [Wiktionary](https://en.wiktionary.org), which is under CC-BY-SA and GFDL licenses.
| github_jupyter |
```
# Load libraries
import pandas as pd
from pandas.tools.plotting import scatter_matrix
import matplotlib.pyplot as plt
from sklearn import cross_validation
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
# Load dataset
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data"
names = ['sepal-length', 'sepal-width', 'petal-length', 'petal-width', 'class']
dataset = pd.read_csv(url, names=names)
print(dataset.shape)
print(dataset.head(20))
# box and whisker plot for each attribute
dataset.plot(kind='box', subplots=True, layout=(2,2), sharex=False, sharey=False)
plt.show()
dataset.hist()
plt.show()
scatter_matrix(dataset)
plt.show()
# Split-out validation dataset
array = dataset.values
X = array[:,0:4]
Y = array[:,4]
validation_size = 0.20
seed = 7
X_train, X_validation, Y_train, Y_validation = cross_validation.train_test_split(X, Y, test_size=validation_size, random_state=seed)
# Test options and evaluation metric
seed = 7
scoring = 'accuracy'
# Spot Check Algorithms
models = []
models.append(('LR', LogisticRegression()))
models.append(('LDA', LinearDiscriminantAnalysis()))
models.append(('KNN', KNeighborsClassifier()))
models.append(('CART', DecisionTreeClassifier()))
models.append(('NB', GaussianNB()))
models.append(('SVM', SVC()))
# evaluate each model in turn
results = []
names = []
for name, model in models:
cv_results=cross_validation.cross_val_score(model, X_train, Y_train, cv=10, scoring=scoring)
results.append(cv_results)
names.append(name)
msg = "%s: %f (%f)" % (name, cv_results.mean(), cv_results.std())
print(msg)
# Compare Algorithms
fig = plt.figure()
fig.suptitle('Algorithm Comparsion')
ax = fig.add_subplot(111)
plt.boxplot(results)
ax.set_xticklabels(names)
plt.show()
knn = KNeighborsClassifier()
knn.fit(X_train, Y_train)
predictions = knn.predict(X_validation)
print("Accuracy: %.2f" % accuracy_score(Y_validation, predictions))
indexes = ['Iris-setosa', 'Iris-versicolor', 'Iris-virginica']
#accuracy_score(Y_validation,predictions)[0:,:]
confusion_matrix_df = pd.DataFrame(
data = confusion_matrix(Y_validation, predictions),
index = indexes,
columns = indexes
)
print()
print(confusion_matrix_df)
print()
print(classification_report(Y_validation, predictions))
```
| github_jupyter |
```
#@title Copyright 2022 The Cirq Developers
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Gate Zoo
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://quantumai.google/cirq/gatezoo.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/quantumai_logo_1x.png" />View on QuantumAI</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/quantumlib/Cirq/blob/master/docs/gatezoo.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/colab_logo_1x.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/quantumlib/Cirq/blob/master/docs/gatezoo.ipynbb"><img src="https://quantumai.google/site-assets/images/buttons/github_logo_1x.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/Cirq/docs/gatezoo.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/download_icon_1x.png" />Download notebook</a>
</td>
</table>
## Setup
Note: this notebook relies on unreleased Cirq features. If you want to try these features, make sure you install cirq via `pip install cirq --pre`
```
try:
import cirq
except ImportError:
print("installing cirq...")
!pip install --quiet --pre cirq
print("installed cirq.")
import IPython.display as ipd
import cirq
import inspect
def display_gates(*gates):
for gate_name in gates:
ipd.display(ipd.Markdown("---"))
gate = getattr(cirq, gate_name)
ipd.display(ipd.Markdown(f"#### cirq.{gate_name}"))
ipd.display(ipd.Markdown(inspect.cleandoc(gate.__doc__ or "")))
else:
ipd.display(ipd.Markdown("---"))
```
Cirq comes with many gates that are standard across quantum computing. This notebook serves as a reference sheet for these gates.
## Single Qubit Gates
### Gate Constants
Cirq defines constants which are gate instances for particular important single qubit gates.
```
display_gates("X", "Y", "Z", "H", "S", "T")
```
### Traditional Pauli Rotation Gates
Cirq defines traditional single qubit rotations that are rotations in radiants abougt different Pauli directions.
```
display_gates("Rx", "Ry", "Rz")
```
### Pauli PowGates
If you think of the `cirq.Z` gate as phasing the state $|1\rangle$ by $-1$, then you might think that the square root of this gate phases the state $|1\rangle$ by $i=\sqrt{-1}$. The `XPowGate`, `YPowGate` and `ZPowGate`s all act in this manner, phasing the state corresponding to their $-1$ eigenvalue by a prescribed amount. This ends up being the same as the `Rx`, `Ry`, and `Rz` up to a global phase.
```
display_gates("XPowGate", "YPowGate", "ZPowGate")
```
### More Single Qubit Gate
Many quantum computing implementations use qubits whose energy eigenstate are the computational basis states. In these cases it is often useful to move `cirq.ZPowGate`'s through other single qubit gates, "phasing" the other gates. For these scenarios, the following phased gates are useful.
```
display_gates("PhasedXPowGate", "PhasedXZGate", "HPowGate")
```
## Two Qubit Gates
### Gate Constants
Cirq defines convenient constants for common two qubit gates.
```
display_gates("CX", "CZ", "SWAP", "ISWAP", "SQRT_ISWAP", "SQRT_ISWAP_INV")
```
### Parity Gates
If $P$ is a non-identity Pauli matrix, then it has eigenvalues $\pm 1$. $P \otimes P$ similarly has eigenvalues $\pm 1$ which are the product of the eigenvalues of the single $P$ eigenvalues. In this sense, $P \otimes P$ has an eigenvalue which encodes the parity of the eigenvalues of the two qubits. If you think of $P \otimes P$ as phasing its $-1$ eigenvectors by $-1$, then you could consider $(P \otimes P)^{\frac{1}{2}}$ as the gate that phases the $-1$ eigenvectors by $\sqrt{-1} =i$. The Parity gates are exactly these gates for the three different non-identity Paulis.
```
display_gates("XXPowGate", "YYPowGate", "ZZPowGate")
```
There are also constants that one can use to define the parity gates via exponentiating them.
```
display_gates("XX", "YY", "ZZ")
```
### Fermionic Gates
If we think of $|1\rangle$ as an excitation, then the gates that preserve the number of excitations are the fermionic gates. There are two implementations, with differing phase conventions.
```
display_gates("FSimGate", "PhasedFSimGate")
```
### Two qubit PowGates
Just as `cirq.XPowGate` represents a powering of `cirq.X`, our two qubit gate constants also have corresponding "Pow" versions.
```
display_gates("SwapPowGate", "ISwapPowGate", "CZPowGate", "CXPowGate", "PhasedISwapPowGate")
```
## Three Qubit Gates
### Gate Constants
Cirq provides constants for common three qubit gates.
```
display_gates("CCX", "CCZ", "CSWAP")
```
### Three Qubit Pow Gates
Corresponding to some of the above gate constants are the corresponding PowGates.
```
display_gates("CCXPowGate", "CCZPowGate")
```
## N Qubit Gates
### Do Nothing Gates
Sometimes you just want a gate to represent doing nothing.
```
display_gates("IdentityGate", "WaitGate")
```
### Measurement Gates
Measurement gates are gates that represent a measurement and can operate on any number of qubits.
```
display_gates("MeasurementGate")
```
### Matrix Gates
If one has a specific unitary matrix in mind, then one can construct it using matrix gates, or, if the unitary is diagonal, the diagonal gates.
```
display_gates("MatrixGate", "DiagonalGate", "TwoQubitDiagonalGate", "ThreeQubitDiagonalGate")
```
### Pauli String Gates
Pauli strings are expressions like "XXZ" representing the Pauli operator X operating on the first two qubits, and Z on the last qubit, along with a numeric (or symbolic) coefficient. When the coefficient is a unit complex number, then this is a valid unitary gate. Similarly one can construct gates which phases the $\pm 1$ eigenvalues of such a Pauli string.
```
display_gates("DensePauliString", "MutableDensePauliString", "PauliStringPhasorGate")
```
### Algorithm Based Gates
It is useful to define composite gates which correspond to algorithmic primitives, i.e. one can think of the fourier transform as a single unitary gate.
```
display_gates("BooleanHamiltonianGate", "QuantumFourierTransformGate", "PhaseGradientGate")
```
### Classiscal Permutation Gates
Sometimes you want to represent shuffling of qubits.
```
display_gates("QubitPermutationGate")
```
| github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/19.Financial_Contract_NER.ipynb)
## 19.Finance Contract NER with Chunk Merger
### Colab Setup
```
import json, os
from google.colab import files
license_keys = files.upload()
with open(list(license_keys.keys())[0]) as f:
license_keys = json.load(f)
# Defining license key-value pairs as local variables
locals().update(license_keys)
# Adding license key-value pairs to environment variables
os.environ.update(license_keys)
# Installing pyspark and spark-nlp
! pip install --upgrade -q pyspark==3.1.2 spark-nlp==$PUBLIC_VERSION
# Installing Spark NLP Healthcare
! pip install --upgrade -q spark-nlp-jsl==$JSL_VERSION --extra-index-url https://pypi.johnsnowlabs.com/$SECRET
# Installing Spark NLP Display Library for visualization
! pip install -q spark-nlp-display
import json
import os
from pyspark.ml import Pipeline,PipelineModel
from pyspark.sql import SparkSession
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
from sparknlp.base import *
from sparknlp.util import *
import sparknlp_jsl
import sparknlp
from pyspark.sql import functions as F
params = {"spark.driver.memory":"16G",
"spark.kryoserializer.buffer.max":"2000M",
"spark.driver.maxResultSize":"2000M"}
spark = sparknlp_jsl.start(license_keys['SECRET'],params=params)
print (sparknlp.version())
print (sparknlp_jsl.version())
spark
```
## Prediction Pipeline
```
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
# Sentence Detector annotator, processes various sentences per line
sentenceDetector = SentenceDetector()\
.setInputCols(["document"])\
.setOutputCol("sentence")\
# Tokenizer splits words in a relevant format for NLP
tokenizer = Tokenizer()\
.setInputCols(["sentence"])\
.setOutputCol("token")
word_embeddings = WordEmbeddingsModel.pretrained("glove_6B_300",'xx')\
.setInputCols(["sentence", 'token'])\
.setOutputCol("word_embeddings")\
.setCaseSensitive(False)
financial_ner_model =MedicalNerModel.pretrained('ner_financial_contract', 'en', 'clinical/models')\
.setInputCols(["sentence", "token", "word_embeddings"])\
.setOutputCol("ner")
ner_converter_1 = NerConverter() \
.setInputCols(["sentence", "token", "ner"]) \
.setOutputCol("ner_chunk_fin")
onto_embeddings = WordEmbeddingsModel.pretrained("glove_100d",'en')\
.setInputCols(["sentence", 'token'])\
.setOutputCol("onto_embeddings")\
.setCaseSensitive(False)
ner_onto = NerDLModel.pretrained(name='onto_100', lang='en')\
.setInputCols(["sentence", "token", "onto_embeddings"])\
.setOutputCol("ner_onto")
ner_converter_2 = NerConverter() \
.setInputCols(["sentence", "token", "ner_onto"]) \
.setOutputCol("ner_chunk_onto")
chunk_merger = ChunkMergeApproach()\
.setInputCols('ner_chunk_onto', "ner_chunk_fin")\
.setOutputCol('ner_chunk')
text_pipeline = Pipeline(stages = [
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings,
onto_embeddings,
financial_ner_model,
ner_onto,
ner_converter_1,ner_converter_2,
chunk_merger
])
empty_df = spark.createDataFrame([['']]).toDF("text")
model_for_text = text_pipeline.fit(empty_df)
ner_onto.getStorageRef()
financial_ner_model.getStorageRef()
financial_ner_model.getClasses()
text = '''6 AFFIRMATIVE COVENANTS |
Borrower has good title to the Collateral , free from liens on 29 November 2018 in Michogan. |
6 . 6 FURTHER ASSURANCES . |
During the additional time , the failure to cure the default is not an Event of Default ( but no Credit Extensions will be made during the cure period ); |
( d ) Apply to the Obligations any ( i ) balances and deposits of Borrower it holds , or ( ii ) any amount held by Bank owing to or for the credit or the account of Borrower ; |
Bank ' s appointment as Borrower ' s attorney in fact , and all of Bank of Michigan ' s rights and powers , coupled |
If Bank complies with reasonable banking practices it is not liable for ( a ) the safekeeping of the Collateral ; ( b ) any loss or damage to the Collateral ; ( c ) any diminution in the value of the Collateral ; or ( d ) any act or default of any carrier , warehouseman , bailee , or other person . |
If there is a default in any agreement between Borrower and a third party that gives the third party the right to accelerate any Indebtedness exceeding $ 100,000 or that could cause a Material Adverse Change ; |
13 CONTRACT CLAIMS , TORT CLAIMS , BREACH OF DUTY CLAIMS , AND ALL OTHER COMMON LAW OR STATUTORY CLAIMS . |
Borrower waives demand, notice of default or dishonor, notice of payment and nonpayment, notice of any default, nonpayment at maturity, release, compromise, settlement, extension, or renewal of accounts, documents, instruments, chattel paper, and guarantees held by Bank on , which Borrower is liable .'''
light_model = LightPipeline(model_for_text)
ann_text = light_model.fullAnnotate(text)
result = light_model.annotate(text)
list(zip(result['token'], result['ner']))
import pandas as pd
result = light_model.fullAnnotate(text)
ner_df= pd.DataFrame([(int(x.metadata['sentence']), x.result, x.begin, x.end, y.result) for x,y in zip(result[0]["token"], result[0]["ner"])],
columns=['sent_id','token','start','end','ner'])
print('Number of Detected NERs in the given Text is :', ner_df.ner[ner_df.ner!='O'].count())
ner_df
chunks = []
entities = []
for n in result[0]['ner_chunk']:
chunks.append(n.result)
entities.append(n.metadata['entity'])
df = pd.DataFrame({'ner_chunk':chunks, 'entities':entities})
df.sample(15)
```
## Highlighting NERs in the Text
```
from sparknlp_display import NerVisualizer
visualiser = NerVisualizer()
visualiser.display(result[0], label_col='ner_chunk', document_col='document')
```
| github_jupyter |
## Comparing results on various datasets using PyDP Naive Bayes vs Scikit-Learn's Naive Bayes
This notebook has implemented both the Naive Bayes algorithm on various datasets such as "adult", "mushroom", "nursery", "digit" and "skin".
Different Training and Testing Accuracies along with the plots aids in understanding the difference in results.
Sources:
- https://github.com/OpenMined/PyDP/blob/dev/src/pydp/ml/naive_bayes.py
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
sns.set_theme(style="white")
np.random.seed(0)
from sklearn.model_selection import KFold
```
## Datasets
Load all the datasets i.e. "adult", "mushroom", "nursery", "digit", "skin"
```
import data_utils
datasets = {
"adult": data_utils.load_adult_dataset(),
"mushroom": data_utils.load_mushroom_dataset(),
"nursery": data_utils.load_nursery_dataset(),
"digit": data_utils.load_digit_dataset(),
"skin": data_utils.load_skin_dataset(),
}
```
## Finding average training and testing accuracies of Scikit-learn's Gaussian Naive Bayes
```
# Train GaussianNB from Sklearn for basline 10-fold average training and testing accuracies
from sklearn.naive_bayes import GaussianNB
clf = GaussianNB()
kfold = KFold(n_splits=10, random_state=42)
```
kfold: The data is split using scikit-learn's K-Folds cross validator
```
def get_fit_result(clf, kfold, df, X, y):
training_accuracies = []
testing_accuracies = []
for train, test in kfold.split(df.values):
X_train = X.values[train]
y_train = y.values[train]
X_test = X.values[test]
y_test = y.values[test]
clf.fit(X_train, y_train)
training_accuracy = clf.score(X_train, y_train)
testing_accuracy = clf.score(X_test, y_test)
training_accuracies.append(training_acucracy)
testing_accuracies.append(testing_acucracy)
mean_train_acc = np.mean(training_accuracies)
mean_test_acc = np.mean(testing_accuracies)
print("Average Training Accuracy")
print(mean_train_acc)
print("Average Testing Accuracy")
print(mean_test_acc)
return mean_train_acc, mean_test_acc
baselines = {}
for name, dataset in datasets.items():
baseline_result = get_fit_result(clf, kfold, dataset[0], dataset[1], dataset[2])
baselines[name] = baseline_result
```
clf: Gaussian Naive Bayes from scikit-learn is used as a classifier on five different datasets
training_accuracies: A dict containing all accuracies of a k-fold cross validator on five different datasets
testing_accuracies: A dict containing all accuracies of a k-fold cross validator on five different datasets
mean_train_acc: Average of training accuracy of a particular dataset
mean_test_acc: Average of testing accuracy of a particular dataset
baseline_result: Stores a tuple containing average training and testing accuracies
baselines: A dict containing names of five different datasets and their corresponding training and testing accuracies as a tuple
```
from pydp.ml.naive_bayes import GaussianNB
epsilons = [10.0, 1.0, 0.1, 0.05, 0.01, 0.005, 0.001]
dp_results = {}
for name, dataset in datasets.items():
mean_train_accs = []
mean_test_accs = []
for epsilon in epsilons:
clf = GaussianNB(epsilon=epsilon)
training_accuracy, test_accuracy = get_fit_result(
clf, kfold, dataset[0], dataset[1], dataset[2]
)
mean_train_accs.append(training_accuracy * 100)
mean_test_accs.append(test_accuracy * 100)
dp_results[name] = mean_train_accs, mean_test_accs
```
clf: Gaussian Naive Bayes from PyDP is used as a classifier on five different datasets
mean_train_accs: Average of training accuracy of a particular dataset
mean_test_accs: Average of testing accuracy of a particular dataset
dp_results: A dict containing names of five different datasets and their corresponding training and testing accuracies as a tuple
```
def plot_result(
ax, dataset_name, baseline_train, baseline_test, dp_train, dp_test, epsilons
):
ax.plot(epsilons, dp_train, marker="o", label="Train (DP)")
ax.plot(epsilons, dp_test, marker="x", label="Test (DP)")
# Set limit on y and axis
ax.set_xlim(0.1, -3.0)
ax.set_ylim(0, 100)
# Set title
ax.set_title(
f"{dataset_name.capitalize()} Dataset:Average Accuracy over 10-fold Cross Validation"
)
# Set label on y and axis
ax.set_ylabel("Train Accuracy (linear scale)")
ax.set_xlabel("Epsilons (log scale 10^)")
# Turn off grid background
ax.grid(False)
# Adjust x axis to display only specific numbers
ax.set_xticks(np.arange(ax.get_xticks().min(), ax.get_xticks().max(), 1.0))
# Draw the read line
ax.axhline(baseline_train * 100, color="r", linewidth=2, label="Baseline Train")
ax.axhline(baseline_test * 100, color="c", linewidth=2, label="Baseline Test")
```
## Plotting corresponding train and test accuracies over 10 folds cross validation using scikit-learn and PyDP's Naive Bayes algorithm
```
fig, axs = plt.subplots(3, 2, figsize=(15, 15), constrained_layout=True)
fig.delaxes(axs[2, 1])
epsilons = np.log10(epsilons)
dataset_name = "adult"
plot_result(
axs[0, 0],
dataset_name,
baselines[dataset_name][0],
baselines[dataset_name][1],
dp_results[dataset_name][0],
dp_results[dataset_name][1],
epsilons,
)
dataset_name = "mushroom"
plot_result(
axs[0, 1],
dataset_name,
baselines[dataset_name][0],
baselines[dataset_name][1],
dp_results[dataset_name][0],
dp_results[dataset_name][1],
epsilons,
)
dataset_name = "nursery"
plot_result(
axs[1, 0],
dataset_name,
baselines[dataset_name][0],
baselines[dataset_name][1],
dp_results[dataset_name][0],
dp_results[dataset_name][1],
epsilons,
)
dataset_name = "digit"
plot_result(
axs[1, 1],
dataset_name,
baselines[dataset_name][0],
baselines[dataset_name][1],
dp_results[dataset_name][0],
dp_results[dataset_name][1],
epsilons,
)
dataset_name = "skin"
plot_result(
axs[2, 0],
dataset_name,
baselines[dataset_name][0],
baselines[dataset_name][1],
dp_results[dataset_name][0],
dp_results[dataset_name][1],
epsilons,
)
handles, labels = axs[2, 0].get_legend_handles_labels()
fig.legend(handles, labels, loc="lower right")
plt.savefig("./result.png")
```
| github_jupyter |
# Breakpoint analysis for damaging winds or rain
Here, we explore the idea that wind/rain damage occurs above some threshold of wind speed, rain rate or rain accumulation.
The damage survey results are classified into damaged/not damaged, and the rate of damaged buildings for a given wind speed/rain rate/rain accumulation is determined by binning the number of damaged buildings per wind speed interval.
We then attempt to determine the threshold at which the damage rate increases significantly, using a Bayesian approach.
```
%matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import seaborn as sns
sns.set_context("poster")
sns.set_style("whitegrid")
sns.set_palette("hls")
```
Read in the damage dataset
```
#filename = "//nas/gemd/georisk/HaRIA_B_Wind/projects/impact_forecasting/data/exposure/NSW/April_2015_Impact_Assessment/Property_Damage_cleaned.csv"
filename = "C:/Workspace/data/derived/exposure/NSW/Property_Damage_cleaned.csv"
df = pd.read_csv(filename)
```
There are a number of blank fields throughout the data where a value was not entered into the dataset by the assessor. We need to keep track of the missing data, as well as the entered data, so we will find all 'NaN' values in the dataset, and change these to 'Not given' so we can include them in subsequent analyses.
```
df = df.fillna('Not given')
```
Now we add a column that indicates whether the building was damaged or not. Any building which is flagged as 'Minor', 'Major', 'Severe' or 'Destroyed' is tagged as damaged
```
damaged = np.zeros(len(df))
damaged[df['EICU_Degdamage'].isin(['Destroyed - 76-100%',
'Severe Impact - 51-75%',
'Major Impact - 26-50%',
'Minor Impact - 1-25%'])] = 1
df['Damaged'] = damaged
```
Determine the maximum wind speed for all data points, and set up bins to determine the rate of damage.
```
vmax = df[df['Damaged']>0]["combined_alltimes_maxwind_stage4_ens12"].max()
bins = np.arange(0, vmax, 0.5)
hist, edges = np.histogram(df[df['Damaged']==1]["combined_alltimes_maxwind_stage4_ens12"].values,
bins=len(bins),
density=False)
plt.bar(bins, hist, width=0.5)
_ = plt.xlabel("Model forecast wind speed (m/s)")
_ = plt.ylabel("Number of damaged buildings")
```
Now we will explore the onset of damage as a function of wind speed.
```
import pymc
switchpoint = pymc.DiscreteUniform('switchpoint',lower=0, upper=vmax)
early_mean = pymc.Exponential('early_mean', beta=1)
late_mean = pymc.Exponential('late_mean', beta=1)
@pymc.deterministic(plot=False)
def rate(s=switchpoint, e=early_mean, l=late_mean):
out = np.empty(len(bins))
out[:s] = e
out[s:] = l
return out
damage = pymc.Poisson('damage', mu=rate, value=hist, observed=True)
model = pymc.Model([switchpoint, early_mean, late_mean, rate, damage])
mcmc = pymc.MCMC(model)
mcmc.sample(iter=10000, burn=1000, thin=10)
plt.figure(figsize=(12,12))
plt.subplot(311);
plt.plot(mcmc.trace('switchpoint')[:]);
plt.ylabel("Switch point");
plt.subplot(312);
plt.plot(mcmc.trace('early_mean')[:]);
plt.ylabel("Early mean");
plt.subplot(313);
plt.plot(mcmc.trace('late_mean')[:]);
plt.xlabel("Iteration");
plt.ylabel("Late mean");
plt.tight_layout()
plt.figure(figsize=(14,3))
plt.subplot(131);
plt.hist(mcmc.trace('switchpoint')[:], 15,);
plt.xlabel("Switch point")
plt.ylabel("Distribution")
plt.subplot(132);
plt.hist(mcmc.trace('early_mean')[:], 15);
plt.xlabel("Early mean");
plt.subplot(133);
plt.hist(mcmc.trace('late_mean')[:], 15);
plt.xlabel("Late mean");
plt.tight_layout()
yp = np.round(mcmc.trace('switchpoint')[:].mean(), 0)
em = mcmc.trace('early_mean')[:].mean()
es = mcmc.trace('early_mean')[:].std()
lm = mcmc.trace('late_mean')[:].mean()
ls = mcmc.trace('late_mean')[:].std()
print((bins[int(yp)], em, es, lm, ls))
plt.figure(figsize=(12,6));
plt.bar(bins, hist, width=0.5);
plt.axvline(bins[int(yp)], color='k', ls='--', label="Mean breakpoint");
plt.plot([0, bins[int(yp)]], [em, em], '-b', lw=3, label="Average damage count below threshold");
plt.plot([bins[int(yp)], len(bins)], [lm, lm], '-r', lw=3, label="Average damage count above threshold");
plt.legend(loc=10, bbox_to_anchor=(0.5, -0.2), ncol=3)
plt.xlim(0, vmax);
plt.xlabel("Model forecast wind speed (m/s)");
plt.ylabel("Number damaged buildings");
```
Repeat this process, using rainfall rate as the predictor.
```
rmax = df[df['Damaged']>0]["combined_alltimes_accum_ls_rainrate_stage4_ens00"].max()
bins = np.linspace(0, rmax, 100)
hist, edges = np.histogram(df[df['Damaged']==1]["combined_alltimes_accum_ls_rainrate_stage4_ens00"].values,
bins=len(bins),
density=False)
plt.bar(bins, hist,width=(bins[1]-bins[0]))
_ = plt.xlabel("Modelled precipitation rate (kg/m^2/s)")
_ = plt.ylabel("Number of damaged buildings")
switchpoint = pymc.DiscreteUniform('switchpoint',lower=0, upper=rmax)
early_mean = pymc.Exponential('early_mean', beta=1)
late_mean = pymc.Exponential('late_mean', beta=1)
@pymc.deterministic(plot=False)
def rate(s=switchpoint, e=early_mean, l=late_mean):
out = np.empty(len(bins))
out[:s] = e
out[s:] = l
return out
damage = pymc.Poisson('damage', mu=rate, value=hist, observed=True)
model = pymc.Model([switchpoint, early_mean, late_mean, rate, damage])
mcmc = pymc.MCMC(model)
mcmc.sample(iter=10000, burn=1000, thin=10)
plt.figure(figsize=(12,12))
plt.subplot(311);
plt.plot(mcmc.trace('switchpoint')[:]);
plt.ylabel("Switch point");
plt.subplot(312);
plt.plot(mcmc.trace('early_mean')[:]);
plt.ylabel("Early mean");
plt.subplot(313);
plt.plot(mcmc.trace('late_mean')[:]);
plt.xlabel("Iteration");
plt.ylabel("Late mean");
plt.tight_layout()
plt.figure(figsize=(14,3))
plt.subplot(131);
plt.hist(mcmc.trace('switchpoint')[:], 15,);
plt.xlabel("Switch point")
plt.ylabel("Distribution")
plt.subplot(132);
plt.hist(mcmc.trace('early_mean')[:], 15);
plt.xlabel("Early mean");
plt.subplot(133);
plt.hist(mcmc.trace('late_mean')[:], 15);
plt.xlabel("Late mean");
plt.tight_layout()
yp = np.round(mcmc.trace('switchpoint')[:].mean(), 0)
em = mcmc.trace('early_mean')[:].mean()
es = mcmc.trace('early_mean')[:].std()
lm = mcmc.trace('late_mean')[:].mean()
ls = mcmc.trace('late_mean')[:].std()
print((bins[int(yp)], em, es, lm, ls))
plt.figure(figsize=(12,6));
plt.bar(bins, hist, width=bins[1]-bins[0]);
plt.axvline(bins[int(yp)], color='k', ls='--', label="Mean breakpoint");
plt.plot([0, bins[int(yp)]], [em, em], '-b', lw=3, label="Average damage count below threshold");
plt.plot([bins[int(yp)], len(bins)], [lm, lm], '-r', lw=3, label="Average damage count above threshold");
plt.legend(loc=10, bbox_to_anchor=(0.5, -0.2), ncol=3)
plt.xlim(0, rmax);
plt.xlabel("Rainfall rate (kg/m^2/s)");
plt.ylabel("Number damaged buildings");
```
TODO:
* Compare to NEXIS building points per bin (wind speed/rainfall rate) for the region
| github_jupyter |
## Module 2.2: Working with CNNs in Keras (A Review)
We turn to implementing a CNN in the Keras functional API. In this module we will pay attention to:
1. Using the Keras functional API for defining models.
2. Implementing dropout regularization.
Those students who are comfortable with all these matters might consider skipping ahead.
Note that we will not spend time tuning hyper-parameters: The purpose is to show how different techniques can be implemented in Keras, not to solve particular data science problems as optimally as possible. Obviously, most techniques include hyper-parameters that need to be tuned for optimal performance.
We start by importing required libraries.
```
import numpy as np
from sklearn.metrics import confusion_matrix,classification_report
from keras.datasets import cifar10
from keras.models import Sequential
from keras import Model
from keras.layers import Dense,Dropout,Flatten,Activation,Input
from keras.optimizers import Adam
from keras.layers.convolutional import Conv2D,MaxPooling2D
from keras.callbacks import EarlyStopping
from keras.utils import np_utils
import matplotlib.pyplot as plt
```
We will use the CIFAR10 dataset. This consists of small (32 x 32 pixel) color images of 10 different types of objects. It is included in the keras.datasets library.
We load the images. These are already split into training and test cases. We need to normalize the pixel values to be between 0 and 1, and turn our integer labels into one-hot vectors - these are 1d-arrays of length the same as the number of classes, with zeros everywhere except the label specified, which is a 1. They are the probability that the image is of different classes.
We also make a vector of class/label names for display purposes, as the label arrays contain only integers.
```
# Load images
(train_images, train_labels), (test_images, test_labels) = cifar10.load_data()
# Normalize pixel values to be between 0 and 1
train_images, test_images = train_images / 255.0, test_images / 255.0
# Make versions of the labels that are one-hot vectors
train_labels_array=np_utils.to_categorical(train_labels, 10)
test_labels_array=np_utils.to_categorical(test_labels, 10)
# Make vector of classnames
class_names = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
train_labels_array.shape
```
Let's make a function to have a look at the images.
```
def show_images(images,labels,class_names,random=True):
plt.figure(figsize=(10,10))
if random:
indices=np.random.randint(0,images.shape[0],25)
else:
indices=np.array([i for i in range(25)])
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(images[indices[i]], cmap=plt.cm.binary)
# The CIFAR labels happen to be arrays,
# which is why we need the extra index
plt.xlabel(class_names[labels[indices[i]][0]])
plt.show()
```
Now we run it. We will see 25 random images from the dataset that we pass. If you set random=False you will see the first 25 images, the variety of which reassures us that the data is in a random order. (If this was a real world problem, such re-assurances would be insufficient, and we would shuffle the data.)
```
show_images(train_images,train_labels,class_names,False)
```
Now we create a function that will define the network architecture. Note that we introduce dropout layers for regularization purposes. We discussed these in the last module.
For comparison, the code to specify the same network using the sequential approach is provided in a second function.
```
def get_model():
inputs = Input(shape=(32, 32, 3),name="Input")
conv1 = Conv2D(32, (3, 3), activation='relu', input_shape=(32, 32, 3))(inputs)
pool1 = MaxPooling2D((2, 2))(conv1)
drop1 = Dropout(0.5)(pool1)
conv2 = Conv2D(64, (3, 3), activation='relu')(drop1)
pool2 = MaxPooling2D((2, 2))(conv2)
drop2 = Dropout(0.5)(pool2)
conv3 = Conv2D(64, (3, 3), activation='relu')(drop2)
flat = Flatten()(conv3)
dense1 = Dense(64, activation='relu')(flat)
outputs = Dense(10, activation='softmax')(dense1)
model = Model(inputs=inputs,outputs=outputs)
return model
# For comparison, this is how we would use the sequential process
def get_model_seqential():
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', input_shape=(32, 32, 3)))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.5))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.5))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(Flatten())
model.add(Dense(64, activation='relu'))
model.add(Dense(10, activation='softmax'))
return model
```
We will get our model.
```
model=get_model()
```
Now we will define an optimizer and compile it. If you are unfamiliar with the different types of optimizers available in keras, I suggest you read the keras documentation [here](https://keras.io/optimizers/) and play around training the model with different alternatives.
```
opt=Adam()
```
And we compile our model with the optimizer ready for training. We use categorical crossentropy as our loss function as this is a good default choice for working with a multi-class categorical target variable (i.e. the image labels).
```
model.compile(optimizer=opt,
loss='categorical_crossentropy',
metrics=['accuracy'])
```
Now we fit (train) the model. We will set the training to continue for 100 epochs, but use an early stopping callback which means it should terminate much quicker than this.
```
# Before calling fit, we create the Early Stopping callback.
# We set it up to stop if improvement in the validation loss
# does not occur over 10 epochs. When stopping occurs, the
# weights associated with the best validation loss are restored.
earlyStopping = EarlyStopping(monitor="val_loss",
patience=10,
verbose=1,
restore_best_weights=True)
# We need to use the one-hot vector version of the labels
# This shouldn't go through all 100 epoches, because of the
# early stopping, but can take some time.
history = model.fit(train_images,
train_labels_array,
epochs=100,
shuffle=True,
callbacks=[earlyStopping],
validation_split=.2)
```
We will plot the training history to see a graphical representation of the training.
```
def plot_training_history(history):
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model accuracy and loss')
plt.xlabel('Epoch')
plt.legend(['Accuracy','Validation Accuracy', 'Loss',
'Validation Loss'], loc='upper right')
plt.show()
plot_training_history(history)
```
Finally, for fun lets see how our improved model performs on our test data. But remember that we have not spent any time or effort optimizing this model - for a real problem we would determine good values for the dropout regularization, as well as tune the architecture and optimizer.
We make a function that will show the confusion matrix, and then run it.
```
def test_model(model,x,y):
y_pred = model.predict(x)
y_pred = np.argmax(y_pred,axis=1)
cm = confusion_matrix(y, y_pred)
print("Confusion Matrix:")
print(cm)
print("Classification report:")
print(classification_report(y, y_pred))
test_model(model,test_images,test_labels)
```
| github_jupyter |
## Dependencies
```
import os
import sys
import cv2
import shutil
import random
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import multiprocessing as mp
import matplotlib.pyplot as plt
from tensorflow import set_random_seed
from sklearn.utils import class_weight
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, cohen_kappa_score
from keras import backend as K
from keras.models import Model
from keras.utils import to_categorical
from keras import optimizers, applications
from keras.preprocessing.image import ImageDataGenerator
from keras.layers import Dense, Dropout, GlobalAveragePooling2D, Input
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, Callback, LearningRateScheduler
def seed_everything(seed=0):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
set_random_seed(0)
seed = 0
seed_everything(seed)
%matplotlib inline
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
sys.path.append(os.path.abspath('../input/efficientnet/efficientnet-master/efficientnet-master/'))
from efficientnet import *
```
## Load data
```
hold_out_set = pd.read_csv('../input/aptos-split-oldnew/hold-out_5.csv')
X_train = hold_out_set[hold_out_set['set'] == 'train']
X_val = hold_out_set[hold_out_set['set'] == 'validation']
test = pd.read_csv('../input/aptos2019-blindness-detection/test.csv')
test["id_code"] = test["id_code"].apply(lambda x: x + ".png")
print('Number of train samples: ', X_train.shape[0])
print('Number of validation samples: ', X_val.shape[0])
print('Number of test samples: ', test.shape[0])
display(X_train.head())
```
# Model parameters
```
# Model parameters
FACTOR = 4
BATCH_SIZE = 8 * FACTOR
EPOCHS = 20
WARMUP_EPOCHS = 5
LEARNING_RATE = 1e-4 * FACTOR
WARMUP_LEARNING_RATE = 1e-3 * FACTOR
HEIGHT = 224
WIDTH = 224
CHANNELS = 3
TTA_STEPS = 5
ES_PATIENCE = 5
RLROP_PATIENCE = 3
DECAY_DROP = 0.5
LR_WARMUP_EPOCHS_1st = 2
LR_WARMUP_EPOCHS_2nd = 5
STEP_SIZE = len(X_train) // BATCH_SIZE
TOTAL_STEPS_1st = WARMUP_EPOCHS * STEP_SIZE
TOTAL_STEPS_2nd = EPOCHS * STEP_SIZE
WARMUP_STEPS_1st = LR_WARMUP_EPOCHS_1st * STEP_SIZE
WARMUP_STEPS_2nd = LR_WARMUP_EPOCHS_2nd * STEP_SIZE
```
# Pre-procecess images
```
old_data_base_path = '../input/diabetic-retinopathy-resized/resized_train/resized_train/'
new_data_base_path = '../input/aptos2019-blindness-detection/train_images/'
test_base_path = '../input/aptos2019-blindness-detection/test_images/'
train_dest_path = 'base_dir/train_images/'
validation_dest_path = 'base_dir/validation_images/'
test_dest_path = 'base_dir/test_images/'
# Making sure directories don't exist
if os.path.exists(train_dest_path):
shutil.rmtree(train_dest_path)
if os.path.exists(validation_dest_path):
shutil.rmtree(validation_dest_path)
if os.path.exists(test_dest_path):
shutil.rmtree(test_dest_path)
# Creating train, validation and test directories
os.makedirs(train_dest_path)
os.makedirs(validation_dest_path)
os.makedirs(test_dest_path)
def crop_image(img, tol=7):
if img.ndim ==2:
mask = img>tol
return img[np.ix_(mask.any(1),mask.any(0))]
elif img.ndim==3:
gray_img = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
mask = gray_img>tol
check_shape = img[:,:,0][np.ix_(mask.any(1),mask.any(0))].shape[0]
if (check_shape == 0): # image is too dark so that we crop out everything,
return img # return original image
else:
img1=img[:,:,0][np.ix_(mask.any(1),mask.any(0))]
img2=img[:,:,1][np.ix_(mask.any(1),mask.any(0))]
img3=img[:,:,2][np.ix_(mask.any(1),mask.any(0))]
img = np.stack([img1,img2,img3],axis=-1)
return img
def circle_crop(img):
img = crop_image(img)
height, width, depth = img.shape
largest_side = np.max((height, width))
img = cv2.resize(img, (largest_side, largest_side))
height, width, depth = img.shape
x = width//2
y = height//2
r = np.amin((x, y))
circle_img = np.zeros((height, width), np.uint8)
cv2.circle(circle_img, (x, y), int(r), 1, thickness=-1)
img = cv2.bitwise_and(img, img, mask=circle_img)
img = crop_image(img)
return img
def preprocess_image(image_id, base_path, save_path, HEIGHT=HEIGHT, WIDTH=WIDTH, sigmaX=10):
image = cv2.imread(base_path + image_id)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = circle_crop(image)
image = cv2.resize(image, (HEIGHT, WIDTH))
# image = cv2.addWeighted(image, 4, cv2.GaussianBlur(image, (0,0), sigmaX), -4 , 128)
cv2.imwrite(save_path + image_id, image)
def preprocess_data(df, HEIGHT=HEIGHT, WIDTH=WIDTH, sigmaX=10):
df = df.reset_index()
for i in range(df.shape[0]):
item = df.iloc[i]
image_id = item['id_code']
item_set = item['set']
item_data = item['data']
if item_set == 'train':
if item_data == 'new':
preprocess_image(image_id, new_data_base_path, train_dest_path)
if item_data == 'old':
preprocess_image(image_id, old_data_base_path, train_dest_path)
if item_set == 'validation':
if item_data == 'new':
preprocess_image(image_id, new_data_base_path, validation_dest_path)
if item_data == 'old':
preprocess_image(image_id, old_data_base_path, validation_dest_path)
def preprocess_test(df, base_path=test_base_path, save_path=test_dest_path, HEIGHT=HEIGHT, WIDTH=WIDTH, sigmaX=10):
df = df.reset_index()
for i in range(df.shape[0]):
image_id = df.iloc[i]['id_code']
preprocess_image(image_id, base_path, save_path)
n_cpu = mp.cpu_count()
train_n_cnt = X_train.shape[0] // n_cpu
val_n_cnt = X_val.shape[0] // n_cpu
test_n_cnt = test.shape[0] // n_cpu
# Pre-procecss old data train set
pool = mp.Pool(n_cpu)
dfs = [X_train.iloc[train_n_cnt*i:train_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = X_train.iloc[train_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_data, [x_df for x_df in dfs])
pool.close()
# Pre-procecss validation set
pool = mp.Pool(n_cpu)
dfs = [X_val.iloc[val_n_cnt*i:val_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = X_val.iloc[val_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_data, [x_df for x_df in dfs])
pool.close()
# Pre-procecss test set
pool = mp.Pool(n_cpu)
dfs = [test.iloc[test_n_cnt*i:test_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = test.iloc[test_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_test, [x_df for x_df in dfs])
pool.close()
```
# Data generator
```
datagen=ImageDataGenerator(rescale=1./255,
rotation_range=360,
horizontal_flip=True,
vertical_flip=True)
train_generator=datagen.flow_from_dataframe(
dataframe=X_train,
directory=train_dest_path,
x_col="id_code",
y_col="diagnosis",
class_mode="raw",
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
seed=seed)
valid_generator=datagen.flow_from_dataframe(
dataframe=X_val,
directory=validation_dest_path,
x_col="id_code",
y_col="diagnosis",
class_mode="raw",
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
seed=seed)
test_generator=datagen.flow_from_dataframe(
dataframe=test,
directory=test_dest_path,
x_col="id_code",
batch_size=1,
class_mode=None,
shuffle=False,
target_size=(HEIGHT, WIDTH),
seed=seed)
def cosine_decay_with_warmup(global_step,
learning_rate_base,
total_steps,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0):
"""
Cosine decay schedule with warm up period.
In this schedule, the learning rate grows linearly from warmup_learning_rate
to learning_rate_base for warmup_steps, then transitions to a cosine decay
schedule.
:param global_step {int}: global step.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param global_step {int}: global step.
:Returns : a float representing learning rate.
:Raises ValueError: if warmup_learning_rate is larger than learning_rate_base, or if warmup_steps is larger than total_steps.
"""
if total_steps < warmup_steps:
raise ValueError('total_steps must be larger or equal to warmup_steps.')
learning_rate = 0.5 * learning_rate_base * (1 + np.cos(
np.pi *
(global_step - warmup_steps - hold_base_rate_steps
) / float(total_steps - warmup_steps - hold_base_rate_steps)))
if hold_base_rate_steps > 0:
learning_rate = np.where(global_step > warmup_steps + hold_base_rate_steps,
learning_rate, learning_rate_base)
if warmup_steps > 0:
if learning_rate_base < warmup_learning_rate:
raise ValueError('learning_rate_base must be larger or equal to warmup_learning_rate.')
slope = (learning_rate_base - warmup_learning_rate) / warmup_steps
warmup_rate = slope * global_step + warmup_learning_rate
learning_rate = np.where(global_step < warmup_steps, warmup_rate,
learning_rate)
return np.where(global_step > total_steps, 0.0, learning_rate)
class WarmUpCosineDecayScheduler(Callback):
"""Cosine decay with warmup learning rate scheduler"""
def __init__(self,
learning_rate_base,
total_steps,
global_step_init=0,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0,
verbose=0):
"""
Constructor for cosine decay with warmup learning rate scheduler.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param global_step_init {int}: initial global step, e.g. from previous checkpoint.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param verbose {int}: quiet, 1: update messages. (default: {0}).
"""
super(WarmUpCosineDecayScheduler, self).__init__()
self.learning_rate_base = learning_rate_base
self.total_steps = total_steps
self.global_step = global_step_init
self.warmup_learning_rate = warmup_learning_rate
self.warmup_steps = warmup_steps
self.hold_base_rate_steps = hold_base_rate_steps
self.verbose = verbose
self.learning_rates = []
def on_batch_end(self, batch, logs=None):
self.global_step = self.global_step + 1
lr = K.get_value(self.model.optimizer.lr)
self.learning_rates.append(lr)
def on_batch_begin(self, batch, logs=None):
lr = cosine_decay_with_warmup(global_step=self.global_step,
learning_rate_base=self.learning_rate_base,
total_steps=self.total_steps,
warmup_learning_rate=self.warmup_learning_rate,
warmup_steps=self.warmup_steps,
hold_base_rate_steps=self.hold_base_rate_steps)
K.set_value(self.model.optimizer.lr, lr)
if self.verbose > 0:
print('\nBatch %02d: setting learning rate to %s.' % (self.global_step + 1, lr))
```
# Model
```
def create_model(input_shape):
input_tensor = Input(shape=input_shape)
base_model = EfficientNetB3(weights=None,
include_top=False,
input_tensor=input_tensor)
base_model.load_weights('../input/efficientnet-keras-weights-b0b5/efficientnet-b3_imagenet_1000_notop.h5')
x = GlobalAveragePooling2D()(base_model.output)
final_output = Dense(1, activation='linear', name='final_output')(x)
model = Model(input_tensor, final_output)
return model
```
# Train top layers
```
model = create_model(input_shape=(HEIGHT, WIDTH, CHANNELS))
for layer in model.layers:
layer.trainable = False
for i in range(-2, 0):
model.layers[i].trainable = True
cosine_lr_1st = WarmUpCosineDecayScheduler(learning_rate_base=WARMUP_LEARNING_RATE,
total_steps=TOTAL_STEPS_1st,
warmup_learning_rate=0.0,
warmup_steps=WARMUP_STEPS_1st,
hold_base_rate_steps=(2 * STEP_SIZE))
metric_list = ["accuracy"]
callback_list = [cosine_lr_1st]
optimizer = optimizers.Adam(lr=WARMUP_LEARNING_RATE)
model.compile(optimizer=optimizer, loss='mean_squared_error', metrics=metric_list)
model.summary()
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history_warmup = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=WARMUP_EPOCHS,
callbacks=callback_list,
verbose=2).history
```
# Fine-tune the complete model
```
for layer in model.layers:
layer.trainable = True
es = EarlyStopping(monitor='val_loss', mode='min', patience=ES_PATIENCE, restore_best_weights=True, verbose=1)
cosine_lr_2nd = WarmUpCosineDecayScheduler(learning_rate_base=LEARNING_RATE,
total_steps=TOTAL_STEPS_2nd,
warmup_learning_rate=0.0,
warmup_steps=WARMUP_STEPS_2nd,
hold_base_rate_steps=(3 * STEP_SIZE))
callback_list = [es, cosine_lr_2nd]
optimizer = optimizers.Adam(lr=LEARNING_RATE)
model.compile(optimizer=optimizer, loss='mean_squared_error', metrics=metric_list)
model.summary()
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=EPOCHS,
callbacks=callback_list,
verbose=2).history
fig, (ax1, ax2) = plt.subplots(2, 1, sharex='col', figsize=(20, 6))
ax1.plot(cosine_lr_1st.learning_rates)
ax1.set_title('Warm up learning rates')
ax2.plot(cosine_lr_2nd.learning_rates)
ax2.set_title('Fine-tune learning rates')
plt.xlabel('Steps')
plt.ylabel('Learning rate')
sns.despine()
plt.show()
```
# Model loss graph
```
fig, (ax1, ax2) = plt.subplots(2, 1, sharex='col', figsize=(20, 14))
ax1.plot(history['loss'], label='Train loss')
ax1.plot(history['val_loss'], label='Validation loss')
ax1.legend(loc='best')
ax1.set_title('Loss')
ax2.plot(history['acc'], label='Train accuracy')
ax2.plot(history['val_acc'], label='Validation accuracy')
ax2.legend(loc='best')
ax2.set_title('Accuracy')
plt.xlabel('Epochs')
sns.despine()
plt.show()
# Create empty arays to keep the predictions and labels
df_preds = pd.DataFrame(columns=['label', 'pred', 'set'])
train_generator.reset()
valid_generator.reset()
# Add train predictions and labels
for i in range(STEP_SIZE_TRAIN + 1):
im, lbl = next(train_generator)
preds = model.predict(im, batch_size=train_generator.batch_size)
for index in range(len(preds)):
df_preds.loc[len(df_preds)] = [lbl[index], preds[index][0], 'train']
# Add validation predictions and labels
for i in range(STEP_SIZE_VALID + 1):
im, lbl = next(valid_generator)
preds = model.predict(im, batch_size=valid_generator.batch_size)
for index in range(len(preds)):
df_preds.loc[len(df_preds)] = [lbl[index], preds[index][0], 'validation']
df_preds['label'] = df_preds['label'].astype('int')
def classify(x):
if x < 0.5:
return 0
elif x < 1.5:
return 1
elif x < 2.5:
return 2
elif x < 3.5:
return 3
return 4
# Classify predictions
df_preds['predictions'] = df_preds['pred'].apply(lambda x: classify(x))
train_preds = df_preds[df_preds['set'] == 'train']
validation_preds = df_preds[df_preds['set'] == 'validation']
```
# Model Evaluation
## Confusion Matrix
### Original thresholds
```
labels = ['0 - No DR', '1 - Mild', '2 - Moderate', '3 - Severe', '4 - Proliferative DR']
def plot_confusion_matrix(train, validation, labels=labels):
train_labels, train_preds = train
validation_labels, validation_preds = validation
fig, (ax1, ax2) = plt.subplots(1, 2, sharex='col', figsize=(24, 7))
train_cnf_matrix = confusion_matrix(train_labels, train_preds)
validation_cnf_matrix = confusion_matrix(validation_labels, validation_preds)
train_cnf_matrix_norm = train_cnf_matrix.astype('float') / train_cnf_matrix.sum(axis=1)[:, np.newaxis]
validation_cnf_matrix_norm = validation_cnf_matrix.astype('float') / validation_cnf_matrix.sum(axis=1)[:, np.newaxis]
train_df_cm = pd.DataFrame(train_cnf_matrix_norm, index=labels, columns=labels)
validation_df_cm = pd.DataFrame(validation_cnf_matrix_norm, index=labels, columns=labels)
sns.heatmap(train_df_cm, annot=True, fmt='.2f', cmap="Blues",ax=ax1).set_title('Train')
sns.heatmap(validation_df_cm, annot=True, fmt='.2f', cmap=sns.cubehelix_palette(8),ax=ax2).set_title('Validation')
plt.show()
plot_confusion_matrix((train_preds['label'], train_preds['predictions']), (validation_preds['label'], validation_preds['predictions']))
```
## Quadratic Weighted Kappa
```
def evaluate_model(train, validation):
train_labels, train_preds = train
validation_labels, validation_preds = validation
print("Train Cohen Kappa score: %.3f" % cohen_kappa_score(train_preds, train_labels, weights='quadratic'))
print("Validation Cohen Kappa score: %.3f" % cohen_kappa_score(validation_preds, validation_labels, weights='quadratic'))
print("Complete set Cohen Kappa score: %.3f" % cohen_kappa_score(np.append(train_preds, validation_preds), np.append(train_labels, validation_labels), weights='quadratic'))
evaluate_model((train_preds['label'], train_preds['predictions']), (validation_preds['label'], validation_preds['predictions']))
```
## Apply model to test set and output predictions
```
def apply_tta(model, generator, steps=10):
step_size = generator.n//generator.batch_size
preds_tta = []
for i in range(steps):
generator.reset()
preds = model.predict_generator(generator, steps=step_size)
preds_tta.append(preds)
return np.mean(preds_tta, axis=0)
preds = apply_tta(model, test_generator, TTA_STEPS)
predictions = [classify(x) for x in preds]
results = pd.DataFrame({'id_code':test['id_code'], 'diagnosis':predictions})
results['id_code'] = results['id_code'].map(lambda x: str(x)[:-4])
# Cleaning created directories
if os.path.exists(train_dest_path):
shutil.rmtree(train_dest_path)
if os.path.exists(validation_dest_path):
shutil.rmtree(validation_dest_path)
if os.path.exists(test_dest_path):
shutil.rmtree(test_dest_path)
```
# Predictions class distribution
```
fig = plt.subplots(sharex='col', figsize=(24, 8.7))
sns.countplot(x="diagnosis", data=results, palette="GnBu_d").set_title('Test')
sns.despine()
plt.show()
results.to_csv('submission.csv', index=False)
display(results.head())
```
## Save model
```
model.save_weights('../working/effNetB3_img224.h5')
```
| github_jupyter |
```
import collections
from collections import defaultdict
import sys
import json
import random
from jsmin import jsmin
from io import StringIO
import numpy as np
import copy
import importlib
from functools import partial
import math
import os
# script_n = os.path.basename(__file__).split('.')[0]
script_n = 'pf_synapse_area_plot_210829'
# from lib_weight_correlation import hist_to_mpd
sys.path.insert(0, '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc')
import my_plot
importlib.reload(my_plot)
from my_plot import MyPlotData
from weight_database import WeightDatabase
weightdb = WeightDatabase()
def weight_fn(syn):
props = syn['props']['area_erode0']
if 'mesh_area' in props:
area = props['mesh_area']
else:
area = 0
if area < .01125:
area = 0
return area
diameter = math.sqrt(area/math.pi)*2
return diameter*1000
gzdb = '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/gen_db/pfs/' \
'gen_210429_setup01_syndb_threshold_10_coalesced.gz'
weightdb.load_syn_db(gzdb,
weight_fn=weight_fn)
pf_mpd = MyPlotData()
pf_hist = defaultdict(int)
weights_db = weightdb.get_weights()
n_pairs = 0
syn_weights = []
avg_data = []
# mpd_data = MyPlotData()
# hist = defaultdict(int)
for neuron, pc_weights in weights_db.items():
# print(n)
for pc, weights in pc_weights.items():
n_pairs += 1
# if len(weights) != 1:
# continue
for avg in weights:
# avg = weights[0]
syn_weights.append(avg)
pf_mpd.add_data_point(
w=avg,
x='Data',
)
pf_hist[avg] += 1
avg_data.append(avg)
print(f'n = {len(syn_weights)} synapses')
print(f'n = {n_pairs} connections')
importlib.reload(my_plot); my_plot.my_displot(
pf_mpd,
x="w",
# y='count_cdf',
# hue='model',
# ci='sd',
# save_filename=f'{script_n}_kde.svg',
show=True,
)
grc_pc_db_f = '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/gen_db/grc_axons/' \
'gen_210429_setup01_syndb_threshold_10_coalesced.gz'
local_weightdb = WeightDatabase()
local_weightdb.load_syn_db(grc_pc_db_f,
weight_fn=weight_fn)
local_mpd = MyPlotData()
hist_local = defaultdict(int)
local_weights_db = local_weightdb.get_weights()
local_syn_weights = []
# avg_data = []
# mpd_data = MyPlotData()
for neuron, pc_weights in local_weights_db.items():
# print(n)
for pc, weights in pc_weights.items():
n_pairs += 1
# if len(weights) != 1:
# continue
for avg in weights:
local_syn_weights.append(avg)
local_mpd.add_data_point(
w=avg,
x='Data',
)
hist_local[avg] += 1
avg_data.append(avg)
print(f'n = {len(syn_weights)} synapses')
print(f'n = {n_pairs} connections')
importlib.reload(my_plot); my_plot.my_displot(
local_mpd,
x="w",
# y='count_cdf',
# hue='model',
# ci='sd',
# save_filename=f'{script_n}_kde.svg',
show=True,
)
mpd_all = MyPlotData()
mpd_all.append(local_mpd)
mpd_all.append(pf_mpd)
importlib.reload(my_plot); my_plot.my_displot(
mpd_all,
x="w",
stat='probability',
bins=40,
# kde=True,
# stat='frequency',
# y='count_cdf',
# hue='model',
# ci='sd',
context='paper',
height=3,
width=5,
xlim=(None, .5),
y_axis_label='Probability',
x_axis_label='Cleft area (µm$^2$)',
show=True,
save_filename=f'{script_n}.svg',
)
print(f'n = {len(mpd_all.data)}')
cv = lambda x: np.std(x, ddof=1) / np.mean(x)
print(f'CV = {cv(avg_data)}')
print(f'n = {len(avg_data)}')
```
| github_jupyter |
```
import csv, random, math
import statistics as st
def loadCsv(filename):
lines = csv.reader(open(filename, "r"));
dataset = list(lines)
for i in range(len(dataset)):
dataset[i] = [float(x) for x in dataset[i]]
return dataset
def splitDataset(dataset, splitRatio):
testSize = int(len(dataset) * splitRatio);
trainSet = list(dataset);
testSet = []
while len(testSet) < testSize:
index = random.randrange(len(trainSet));
testSet.append(trainSet.pop(index))
return [trainSet, testSet]
def separateByClass(dataset):
separated = {}
for i in range(len(dataset)):
x = dataset[i] # current row
if (x[-1] not in separated):
separated[x[-1]] = []
separated[x[-1]].append(x)
return separated
def compute_mean_std(dataset):
mean_std = [ (st.mean(attribute), st.stdev(attribute))
for attribute in zip(*dataset)]; #zip(*res) transposes a matrix (2-d array/list)
del mean_std[-1] # Exclude label, i.e., target
return mean_std
def summarizeByClass(dataset): # summary is the mean and STD of class values
separated = separateByClass(dataset);
summary = {} # to store mean and std of +ve and -ve instances
for classValue, instances in separated.items():
#summaries is a dictionary of tuples(mean,std) for each class value
summary[classValue] = compute_mean_std(instances)
return summary
#For continuous attributes, p is estimated using Gaussian distribution
def estimateProbability(x, mean, stdev):
exponent = math.exp(-(math.pow(x-mean,2)/(2*math.pow(stdev,2))))
return (1 / (math.sqrt(2*math.pi) * stdev)) * exponent
# calculate class probabilities of that entire row (testVector)
def calculateClassProbabilities(summaries, testVector):
p = {}
#class and attribute information as mean and sd
for classValue, classSummaries in summaries.items():
p[classValue] = 1
for i in range(len(classSummaries)):
mean, stdev = classSummaries[i]
x = testVector[i] #testvector's i-th attribute
#use normal distribution
p[classValue] *= estimateProbability(x, mean, stdev)
return p
# calculate best out of all class probabilities of that entire row (testVector)
def predict(summaries, testVector):
all_p = calculateClassProbabilities(summaries, testVector)
bestLabel, bestProb = None, -1
for lbl, p in all_p.items():#assigns that class which has he highest prob
if bestLabel is None or p > bestProb:
bestProb = p
bestLabel = lbl
return bestLabel
# find predicted class for each row in testSet
def perform_classification(summaries, testSet):
predictions = []
for i in range(len(testSet)):
result = predict(summaries, testSet[i])
predictions.append(result)
return predictions
def getAccuracy(testSet, predictions):
correct = 0
for i in range(len(testSet)):
if testSet[i][-1] == predictions[i]:
correct += 1
return (correct/float(len(testSet))) * 100.0
# dataset = loadCsv('pima-indians-diabetes.csv');
dataset = loadCsv('prog5_dataset.csv');
#print dataset
print('Pima Indian Diabetes Dataset loaded...')
print('Total instances available :',len(dataset))
print('Total attributes present :',len(dataset[0])-1)
print("First Five instances of dataset:")
for i in range(5):
print(i+1 , ':' , dataset[i])
splitRatio = 0.2
trainingSet, testSet = splitDataset(dataset, splitRatio)
print('\nDataset is split into training and testing set.')
print('Training examples = {0} \nTesting examples = {1}'.format(len(trainingSet), len(testSet)))
summaries = summarizeByClass(trainingSet);
predictions = perform_classification(summaries, testSet)
accuracy = getAccuracy(testSet, predictions)
print('\nAccuracy of the Naive Baysian Classifier is :', accuracy)
```
| github_jupyter |
# Dependent density regression
In another [example](dp_mix.ipynb), we showed how to use Dirichlet processes to perform Bayesian nonparametric density estimation. This example expands on the previous one, illustrating dependent density regression.
Just as Dirichlet process mixtures can be thought of as infinite mixture models that select the number of active components as part of inference, dependent density regression can be thought of as infinite [mixtures of experts](https://en.wikipedia.org/wiki/Committee_machine) that select the active experts as part of inference. Their flexibility and modularity make them powerful tools for performing nonparametric Bayesian Data analysis.
```
import arviz as az
import numpy as np
import pandas as pd
import pymc3 as pm
import seaborn as sns
from IPython.display import HTML
from matplotlib import animation as ani
from matplotlib import pyplot as plt
from theano import tensor as tt
print(f"Running on PyMC3 v{pm.__version__}")
%config InlineBackend.figure_format = 'retina'
plt.rc("animation", writer="ffmpeg")
blue, *_ = sns.color_palette()
az.style.use("arviz-darkgrid")
SEED = 972915 # from random.org; for reproducibility
np.random.seed(SEED)
```
We will use the LIDAR data set from Larry Wasserman's excellent book, [_All of Nonparametric Statistics_](http://www.stat.cmu.edu/~larry/all-of-nonpar/). We standardize the data set to improve the rate of convergence of our samples.
```
DATA_URI = "http://www.stat.cmu.edu/~larry/all-of-nonpar/=data/lidar.dat"
def standardize(x):
return (x - x.mean()) / x.std()
df = pd.read_csv(DATA_URI, sep=r"\s{1,3}", engine="python").assign(
std_range=lambda df: standardize(df.range), std_logratio=lambda df: standardize(df.logratio)
)
df.head()
```
We plot the LIDAR data below.
```
fig, ax = plt.subplots(figsize=(8, 6))
ax.scatter(df.std_range, df.std_logratio, color=blue)
ax.set_xticklabels([])
ax.set_xlabel("Standardized range")
ax.set_yticklabels([])
ax.set_ylabel("Standardized log ratio");
```
This data set has a two interesting properties that make it useful for illustrating dependent density regression.
1. The relationship between range and log ratio is nonlinear, but has locally linear components.
2. The observation noise is [heteroskedastic](https://en.wikipedia.org/wiki/Heteroscedasticity); that is, the magnitude of the variance varies with the range.
The intuitive idea behind dependent density regression is to reduce the problem to many (related) density estimates, conditioned on fixed values of the predictors. The following animation illustrates this intuition.
```
fig, (scatter_ax, hist_ax) = plt.subplots(ncols=2, figsize=(16, 6))
scatter_ax.scatter(df.std_range, df.std_logratio, color=blue, zorder=2)
scatter_ax.set_xticklabels([])
scatter_ax.set_xlabel("Standardized range")
scatter_ax.set_yticklabels([])
scatter_ax.set_ylabel("Standardized log ratio")
bins = np.linspace(df.std_range.min(), df.std_range.max(), 25)
hist_ax.hist(df.std_logratio, bins=bins, color="k", lw=0, alpha=0.25, label="All data")
hist_ax.set_xticklabels([])
hist_ax.set_xlabel("Standardized log ratio")
hist_ax.set_yticklabels([])
hist_ax.set_ylabel("Frequency")
hist_ax.legend(loc=2)
endpoints = np.linspace(1.05 * df.std_range.min(), 1.05 * df.std_range.max(), 15)
frame_artists = []
for low, high in zip(endpoints[:-1], endpoints[2:]):
interval = scatter_ax.axvspan(low, high, color="k", alpha=0.5, lw=0, zorder=1)
*_, bars = hist_ax.hist(
df[df.std_range.between(low, high)].std_logratio, bins=bins, color="k", lw=0, alpha=0.5
)
frame_artists.append((interval,) + tuple(bars))
animation = ani.ArtistAnimation(fig, frame_artists, interval=500, repeat_delay=3000, blit=True)
plt.close()
# prevent the intermediate figure from showing
HTML(animation.to_html5_video())
```
As we slice the data with a window sliding along the x-axis in the left plot, the empirical distribution of the y-values of the points in the window varies in the right plot. An important aspect of this approach is that the density estimates that correspond to close values of the predictor are similar.
In the previous example, we saw that a Dirichlet process estimates a probability density as a mixture model with infinitely many components. In the case of normal component distributions,
$$y \sim \sum_{i = 1}^{\infty} w_i \cdot N(\mu_i, \tau_i^{-1}),$$
where the mixture weights, $w_1, w_2, \ldots$, are generated by a [stick-breaking process](https://en.wikipedia.org/wiki/Dirichlet_process#The_stick-breaking_process).
Dependent density regression generalizes this representation of the Dirichlet process mixture model by allowing the mixture weights and component means to vary conditioned on the value of the predictor, $x$. That is,
$$y\ |\ x \sim \sum_{i = 1}^{\infty} w_i\ |\ x \cdot N(\mu_i\ |\ x, \tau_i^{-1}).$$
In this example, we will follow Chapter 23 of [_Bayesian Data Analysis_](http://www.stat.columbia.edu/~gelman/book/) and use a probit stick-breaking process to determine the conditional mixture weights, $w_i\ |\ x$. The probit stick-breaking process starts by defining
$$v_i\ |\ x = \Phi(\alpha_i + \beta_i x),$$
where $\Phi$ is the cumulative distribution function of the standard normal distribution. We then obtain $w_i\ |\ x$ by applying the stick breaking process to $v_i\ |\ x$. That is,
$$w_i\ |\ x = v_i\ |\ x \cdot \prod_{j = 1}^{i - 1} (1 - v_j\ |\ x).$$
For the LIDAR data set, we use independent normal priors $\alpha_i \sim N(0, 5^2)$ and $\beta_i \sim N(0, 5^2)$. We now express this this model for the conditional mixture weights using `PyMC3`.
```
def norm_cdf(z):
return 0.5 * (1 + tt.erf(z / np.sqrt(2)))
def stick_breaking(v):
return v * tt.concatenate(
[tt.ones_like(v[:, :1]), tt.extra_ops.cumprod(1 - v, axis=1)[:, :-1]], axis=1
)
N = len(df)
K = 20
std_range = df.std_range.values[:, np.newaxis]
std_logratio = df.std_logratio.values
with pm.Model(coords={"N": np.arange(N), "K": np.arange(K) + 1, "one": [1]}) as model:
alpha = pm.Normal("alpha", 0.0, 5.0, dims="K")
beta = pm.Normal("beta", 0.0, 5.0, dims=("one", "K"))
x = pm.Data("x", std_range)
v = norm_cdf(alpha + pm.math.dot(x, beta))
w = pm.Deterministic("w", stick_breaking(v), dims=["N", "K"])
```
We have defined `x` as a `pm.Data` container in order to use `PyMC3`'s posterior prediction capabilities later.
While the dependent density regression model theoretically has infinitely many components, we must truncate the model to finitely many components (in this case, twenty) in order to express it using `PyMC3`. After sampling from the model, we will verify that truncation did not unduly influence our results.
Since the LIDAR data seems to have several linear components, we use the linear models
$$
\begin{align*}
\mu_i\ |\ x
& \sim \gamma_i + \delta_i x \\
\gamma_i
& \sim N(0, 10^2) \\
\delta_i
& \sim N(0, 10^2)
\end{align*}
$$
for the conditional component means.
```
with model:
gamma = pm.Normal("gamma", 0.0, 10.0, dims="K")
delta = pm.Normal("delta", 0.0, 10.0, dims=("one", "K"))
mu = pm.Deterministic("mu", gamma + pm.math.dot(x, delta))
```
Finally, we place the prior $\tau_i \sim \textrm{Gamma}(1, 1)$ on the component precisions.
```
with model:
tau = pm.Gamma("tau", 1.0, 1.0, dims="K")
y = pm.Data("y", std_logratio)
obs = pm.NormalMixture("obs", w, mu, tau=tau, observed=y)
pm.model_to_graphviz(model)
```
We now sample from the dependent density regression model.
```
SAMPLES = 20000
BURN = 10000
with model:
step = pm.Metropolis()
trace = pm.sample(SAMPLES, tune=BURN, step=step, random_seed=SEED, return_inferencedata=True)
```
To verify that truncation did not unduly influence our results, we plot the largest posterior expected mixture weight for each component. (In this model, each point has a mixture weight for each component, so we plot the maximum mixture weight for each component across all data points in order to judge if the component exerts any influence on the posterior.)
```
fig, ax = plt.subplots(figsize=(8, 6))
max_mixture_weights = trace.posterior["w"].mean(("chain", "draw")).max("N")
ax.bar(max_mixture_weights.coords.to_index(), max_mixture_weights)
ax.set_xlim(1 - 0.5, K + 0.5)
ax.set_xticks(np.arange(0, K, 2) + 1)
ax.set_xlabel("Mixture component")
ax.set_ylabel("Largest posterior expected\nmixture weight");
```
Since only three mixture components have appreciable posterior expected weight for any data point, we can be fairly certain that truncation did not unduly influence our results. (If most components had appreciable posterior expected weight, truncation may have influenced the results, and we would have increased the number of components and sampled again.)
Visually, it is reasonable that the LIDAR data has three linear components, so these posterior expected weights seem to have identified the structure of the data well. We now sample from the posterior predictive distribution to get a better understand the model's performance.
```
PP_SAMPLES = 5000
lidar_pp_x = np.linspace(std_range.min() - 0.05, std_range.max() + 0.05, 100)
with model:
pm.set_data({"x": lidar_pp_x[:, np.newaxis]})
pp_trace = pm.sample_posterior_predictive(trace, PP_SAMPLES, random_seed=SEED)
```
Below we plot the posterior expected value and the 95% posterior credible interval.
```
fig, ax = plt.subplots(figsize=(8, 6))
ax.scatter(df.std_range, df.std_logratio, color=blue, zorder=10, label=None)
low, high = np.percentile(pp_trace["obs"], [2.5, 97.5], axis=0)
ax.fill_between(
lidar_pp_x, low, high, color="k", alpha=0.35, zorder=5, label="95% posterior credible interval"
)
ax.plot(lidar_pp_x, pp_trace["obs"].mean(axis=0), c="k", zorder=6, label="Posterior expected value")
ax.set_xticklabels([])
ax.set_xlabel("Standardized range")
ax.set_yticklabels([])
ax.set_ylabel("Standardized log ratio")
ax.legend(loc=1)
ax.set_title("LIDAR Data");
```
The model has fit the linear components of the data well, and also accomodated its heteroskedasticity. This flexibility, along with the ability to modularly specify the conditional mixture weights and conditional component densities, makes dependent density regression an extremely useful nonparametric Bayesian model.
To learn more about depdendent density regression and related models, consult [_Bayesian Data Analysis_](http://www.stat.columbia.edu/~gelman/book/), [_Bayesian Nonparametric Data Analysis_](http://www.springer.com/us/book/9783319189673), or [_Bayesian Nonparametrics_](https://www.google.com/webhp?sourceid=chrome-instant&ion=1&espv=2&ie=UTF-8#q=bayesian+nonparametrics+book).
This example first appeared [here](http://austinrochford.com/posts/2017-01-18-ddp-pymc3.html).
Author: [Austin Rochford](https://github.com/AustinRochford/)
```
%load_ext watermark
%watermark -n -u -v -iv -w
```
| github_jupyter |
```
# run this code to login to https://okpy.org/ and setup the assignment for submission
from ist256 import okclient
ok = okclient.Lab()
```
# Class Coding Lab: Variables And Types
The goals of this lab are to help you to understand:
1. Python data types
1. Getting input as different types
1. Formatting output as different types
1. Basic arithmetic operators
1. How to create a program from an idea.
## Variable Types
Every Python variable has a **type**. The Type determines how the data is stored in the computer's memory:
```
a = "4"
type(a) # should be str
a = 4
type(a) # should be int
```
### Types Matter
Python's built in functions and operators work differently depending on the type of the variable.:
```
a = 4
b = 5
a + b # this plus in this case means add so 9
a = "4"
b = "5"
a + b # the plus + in this case means concatenation, so '45'
```
### Switching Types
there are built-in Python functions for switching types. For example:
```
x = "45" # x is a str
y = int(x) # y is now an int
z = float(x) # z is a float
print(x,y,z)
```
### Inputs type str
When you use the `input()` function the result is of type `str`:
```
age = input("Enter your age: ")
type(age)
```
We can use a built in Python function to convert the type from `str` to our desired type:
```
age = input("Enter your age: ")
age = int(age)
type(age)
```
We typically combine the first two lines into one expression like this:
```
age = int(input("Enter your age: "))
type(age)
```
## Now Try This:
Write a program to:
- input your age, convert it to an int and store it in a variable
- add one to your age, store it in another variable
- print out your current age and your age next year.
For example:
```
Enter your age: 45
Today you are 45 next year you will be 46
```
```
# TODO: Write your code here
age=int(input("enter your age: "))
ageb=int(1)
print(age+ageb)
```
## Format Codes
Python has some string format codes which allow us to control the output of our variables.
- %s = format variable as str
- %d = format variable as int
- %f = format variable as float
You can also include the number of spaces to use for example `%5.2f` prints a float with 5 spaces 2 to the right of the decimal point.
```
name = "Mike"
age = 45
gpa = 3.4
print("%s is %d years old. His gpa is %.3f" % (name, age,gpa))
```
## Formatting with F-Strings
The other method of formatting data in Python is F-strings. As we saw in the last lab, F-strings use interpolation to specify the variables we would like to print in-line with the print string.
You can format an f-string
- `{var:d}` formats `var` as integer
- `{var:f}` formats `var` as float
- `{var:.3f}` formats `var` as float to `3` decimal places.
Example:
```
name ="Mike"
wage = 15
print(f"{name} makes ${wage:.2f} per hour")
```
## Now Try This:
Write a print statement using F-strings. print GPA to 3 decimal places, rest of the output should appear just like the example above
```
name = "Mike"
age = 45
gpa = 3.4
print("%s is %d years old. His gpa is %.3f" % (name, age,gpa))
## TODO: Rewrite the above line to use an F-string instead
```
## Now Try This:
Print the PI variable out 3 times. Once as a string, once as an int and once as a float to 4 decimal places. Use either F-strings for Format Codes.
```
PI = 3.1415927
#TODO: Write Code Here
```
## Putting it all together: Fred's Fence Estimator
Fred's Fence has hired you to write a program to estimate the cost of their fencing projects. For a given length and width you will calculate the number of 6 foot fence sections, and the total cost of the project. Each fence section costs $23.95. Assume the posts and labor are free.
Program Inputs:
- Length of yard in feet
- Width of yard in feet
Program Outputs:
- Perimeter of yard ( 2 x (Length + Width))
- Number of fence sections required (Permiemer divided by 6 )
- Total cost for fence ( fence sections multiplied by $23.95 )
NOTE: All outputs should be formatted to 2 decimal places: e.g. 123.05
```
#TODO:
# 1. Input length of yard as float, assign to a variable
# 2. Input Width of yard as float, assign to a variable
# 3. Calculate perimeter of yard, assign to a variable
# 4. calculate number of fence sections, assign to a variable
# 5. calculate total cost, assign to variable
# 6. print perimeter of yard
# 7. print number of fence sections
# 8. print total cost for fence.
length=float(input("Input length of yard: "))
width=float(input("Input width of yard: "))
perimeter=(2*(length+width))
fencesec=perimeter//6
cost=fencesec*float(23.95)
print(perimeter)
print(fencesec)
print(cost)
```
## Now Try This
Based on the provided TODO, write the program in python in the cell below. Your solution should have 8 lines of code, one for each TODO.
**HINT**: Don't try to write the program in one sitting. Instead write a line of code, run it, verify it works and fix any issues with it before writing the next line of code.
```
# TODO: Write your code here
print("oops i think i did it above")
```
## Metacognition
Please answer the following questions. This should be a personal narrative, in your own voice. Answer the questions by double clicking on the question and placing your answer next to the Answer: prompt.
1. Record any questions you have about this lab that you would like to ask in recitation. It is expected you will have questions if you did not complete the code sections correctly. Learning how to articulate what you do not understand is an important skill of critical thinking.
Answer: I don't really understand most things that involve use with the percent sign (%)
2. What was the most difficult aspect of completing this lab? Least difficult?
Answer: The difficult part of completing this lab was figuring out how to make the code spit out values that end in two decimal places
3. What aspects of this lab do you find most valuable? Least valuable?
Answer: i like learning about the different values, i find intigers to be the least valuble since im working with real numbers most of the time
4. Rate your comfort level with this week's material so far.
1 ==> I can do this on my own and explain how to do it.
2 ==> I can do this on my own without any help.
3 ==> I can do this with help or guidance from others. If you choose this level please list those who helped you.
4 ==> I don't understand this at all yet and need extra help. If you choose this please try to articulate that which you do not understand.
Answer: 2
```
# to save and turn in your work, execute this cell. Your latest submission will be graded.
ok.submit()
```
| github_jupyter |
# MLflow end-to-end example
In this example we are going to build a model using `mlflow`, pack and deploy locally using `tempo` (in docker and local kubernetes cluster).
We are are going to use follow the MNIST pytorch example from `mlflow`, check this [link](https://github.com/mlflow/mlflow/tree/master/examples/pytorch/MNIST) for more information.
In this example we will:
* [Train MNIST Model using mlflow and pytorch](#Train-model)
* [Create tempo artifacts](#Save-model-environment)
* [Deploy Locally to Docker](#Deploy-to-Docker)
* [Deploy Locally to Kubernetes](#Deploy-to-Kubernetes)
## Prerequisites
This notebooks needs to be run in the `tempo-examples` conda environment defined below. Create from project root folder:
```bash
conda env create --name tempo-examples --file conda/tempo-examples.yaml
```
## Train model
We train MNIST model below:
### Install prerequisites
```
!pip install mlflow 'torchvision>=0.9.1' torch==1.9.0 pytorch-lightning==1.4.0
!rm -fr /tmp/mlflow
%cd /tmp
!git clone https://github.com/mlflow/mlflow.git
```
### Train model using `mlflow`
```
%cd mlflow/examples/pytorch/MNIST
!mlflow run . --no-conda
!tree -L 1 mlruns/0
```
### Choose test image
```
from torchvision import datasets
mnist_test = datasets.MNIST('/tmp/data', train=False, download=True)
# change the index below to get a different image for testing
mnist_test = list(mnist_test)[0]
img, category = mnist_test
display(img)
print(category)
```
### Tranform test image to numpy
```
import numpy as np
img_np = np.asarray(img).reshape((1, 28*28)).astype(np.float32)
```
## Save model environment
```
import glob
import os
files = glob.glob("mlruns/0/*/")
files.sort(key=os.path.getmtime)
ARTIFACTS_FOLDER = os.path.join(
os.getcwd(),
files[-1],
"artifacts",
"model"
)
assert os.path.exists(ARTIFACTS_FOLDER)
print(ARTIFACTS_FOLDER)
```
### Define `tempo` model
```
from tempo.serve.metadata import ModelFramework
from tempo.serve.model import Model
mlflow_tag = "mlflow"
pytorch_mnist_model = Model(
name="test-pytorch-mnist",
platform=ModelFramework.MLFlow,
local_folder=ARTIFACTS_FOLDER,
# if we deploy to kube, this defines where the model artifacts are stored
uri="s3://tempo/basic/mnist",
description="A pytorch MNIST model",
)
```
### Save model (environment) using `tempo`
Tempo hides many details required to save the model environment for `mlserver`:
- Add required runtime dependencies
- Create a conda pack `environment.tar.gz`
```
from tempo.serve.loader import save
save(pytorch_mnist_model)
```
## Deploy to Docker
```
from tempo import deploy_local
local_deployed_model = deploy_local(pytorch_mnist_model)
local_prediction = local_deployed_model.predict(img_np)
print(np.nonzero(local_prediction.flatten() == 0))
local_deployed_model.undeploy()
```
## Deploy to Kubernetes
### Prerequisites
Create a Kind Kubernetes cluster with Minio and Seldon Core installed using Ansible as described [here](https://tempo.readthedocs.io/en/latest/overview/quickstart.html#kubernetes-cluster-with-seldon-core).
```
%cd -0
!kubectl apply -f k8s/rbac -n production
```
### Upload artifacts to minio
```
from tempo.examples.minio import create_minio_rclone
import os
create_minio_rclone(os.getcwd()+"/rclone.conf")
from tempo.serve.loader import upload
upload(pytorch_mnist_model)
```
### Deploy to `kind`
```
from tempo.serve.metadata import SeldonCoreOptions
runtime_options = SeldonCoreOptions(**{
"remote_options": {
"namespace": "production",
"authSecretName": "minio-secret"
}
})
from tempo import deploy_remote
remote_deployed_model = deploy_remote(pytorch_mnist_model, options=runtime_options)
remote_prediction = remote_deployed_model.predict(img_np)
print(np.nonzero(remote_prediction.flatten() == 0))
remote_deployed_model.undeploy()
```
| github_jupyter |
####**... Meanwhile on 15 Dec 2020.**
---
###<div align="left">I have been quite familiar how **softmax** and **negative log likelihood** function works in general. But why below Crossentropy function from *pytorch.org* gave me headache? </div>
> ##### <div align="right">*author : alvinwatner*</div>

https://pytorch.org/docs/stable/generated/torch.nn.CrossEntropyLoss.html
### If you feel the same way, then take a look how i tried to boil down above equation.
----
#### Based on my **perspective**, that above equation is not being consistent, since it seems like they try to combine the math style with the programming style. But well, in terms of code documentation it's not necessary to stick on which particular style as long as it's still readable and maintain the actual computation.
<br />
<br />
#### Anyway moving on, so actually that just softmax & negative natural log likelihood. How is that?

<br />
###...Now let's try to describe it more mathy.
---
###### <div align="center">---- *Note : I use the bold letter to represent a **vector** and non-bold as **scalar**. ----*</div>
---
<br />
#### First of all, let's have a look how softmax function looks like in general,
> $$S(\mathbf{x}) = \mathbf{b}$$
#### The above function $S$ take an input vector $\mathbf{x}$ and output another vector $\mathbf{b}$.
> $$\mathbf{b} = [ b_{1}, b_{2}, \dots, b_{n}]$$
### Below is the calculation for each $b_i$
> $$b_i = \displaystyle \frac{e^x_i}{\sum_{i=1}^{n} e^x_j}$$
##### Since we make use of the magical euler number $e$, which is a mathematical constant approximately equal to 2.71828, it turns out that the sum of $$b_{1}, b_{2}, \dots, b_{n} = 1.0$$
### Let the target class vector denoted as $\mathbf{c}$.
> $$\mathbf{c} = [ c_{1}, c_{2}, \dots, c_{n}]$$
### Let the *$b_i$ that correspond to the target class $c_i$ equal to $b_c$* .
#### Then we attempt to maximize the $b_c$ by minimizing the negative natural log of $b_c$ shown as below
> ### $$loss( \mathbf{b} , \mathbf{c}) = - ln(b_{c})$$
### Allright... Let's see if it is true...
```
_input_vector_x = [0,2,0,1]
_target_class_c = [2]
import torch
import torch.nn as nn
from torch.autograd import Variable
x_torchTensor = Variable(torch.FloatTensor(_input_vector_x)).view(1, -1)
t_torchTensor = Variable(torch.LongTensor(_target_class_c))
print(f"x_torchTensor = {x_torchTensor}")
print(f"t_torchTensor = {t_torchTensor}")
# |Pytorch| Cross Entropy Loss (Version)
criterion = nn.CrossEntropyLoss()
torchC_Loss = criterion(x_torchTensor, t_torchTensor)
print(f"Pytorch Cross Entropy Loss = {torchC_Loss}")
# |Pytorch| Negative Log Likelihood and Softmax (Version)
fsoftmax_ = nn.Softmax(dim=1)
b_ = fsoftmax_(x_torchTensor)
bt_ = b_[0, _target_class_c] # Pull out the value of the index of _target_class_ in b_
print(f"Softmax of x_torchTensor = {b_}")
print(f"The index of _target_class_ from the softmax probability b_ = {bt_}")
torchNegLog = -1 * torch.log(bt_)
print("")
print(f"Pytorch Negative log of bt_ = {torchNegLog}")
# No Torch, Let's use Numpy
# |Numpy| Negative Log Likelihood and Softmax (Version)
import numpy as np
x_npTensor = np.array(_input_vector_x)
t_npTensor = np.array(_target_class_c)
def softmax_(x):
b_ = np.array([])
denominator = np.array([])
''' ∑ e^x_j '''
for x_j in x:
denominator = np.append(denominator, np.exp(x_j))
denominator = np.sum(denominator)
''' softmax_vector (denoted as b_) = [ e^x_i / ∑ e^x_j ]'''
for numerator in x:
b_ = np.append(b_, np.exp(numerator)/denominator)
return b_
def neglogLoss(b_, target_class):
negLog_vector = np.array([])
for t in target_class:
negLog = -1 * np.log(b_[t])
negLog_vector = np.append(negLog_vector, negLog)
return negLog_vector
b_ = softmax_(x_npTensor)
bt_ = neglogLoss(b_, t_npTensor)
print(f"Numpy Negative log of bt_ = {bt_}")
# No Torch, Let's use Numpy
# |Pytorch| CrossEntropy Loss (Version)
def crossEntropyLoss(x, target_class):
b_ = softmax_(x)
bt_ = neglogLoss(b_, target_class)
return bt_
npC_Loss = crossEntropyLoss(x_npTensor, t_npTensor)
print(f"Numpy Cross Entropy Loss = {npC_Loss}")
```
| github_jupyter |
# Language Translation
In this project, you’re going to take a peek into the realm of neural network machine translation. You’ll be training a sequence to sequence model on a dataset of English and French sentences that can translate new sentences from English to French.
## Get the Data
Since translating the whole language of English to French will take lots of time to train, we have provided you with a small portion of the English corpus.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import helper
import problem_unittests as tests
source_path = 'data/small_vocab_en'
target_path = 'data/small_vocab_fr'
source_text = helper.load_data(source_path)
target_text = helper.load_data(target_path)
```
## Explore the Data
Play around with view_sentence_range to view different parts of the data.
```
view_sentence_range = (0, 10)
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import numpy as np
print('Dataset Stats')
print('Roughly the number of unique words: {}'.format(len({word: None for word in source_text.split()})))
sentences = source_text.split('\n')
word_counts = [len(sentence.split()) for sentence in sentences]
print('Number of sentences: {}'.format(len(sentences)))
print('Average number of words in a sentence: {}'.format(np.average(word_counts)))
print()
print('English sentences {} to {}:'.format(*view_sentence_range))
print('\n'.join(source_text.split('\n')[view_sentence_range[0]:view_sentence_range[1]]))
print()
print('French sentences {} to {}:'.format(*view_sentence_range))
print('\n'.join(target_text.split('\n')[view_sentence_range[0]:view_sentence_range[1]]))
```
## Implement Preprocessing Function
### Text to Word Ids
As you did with other RNNs, you must turn the text into a number so the computer can understand it. In the function `text_to_ids()`, you'll turn `source_text` and `target_text` from words to ids. However, you need to add the `<EOS>` word id at the end of each sentence from `target_text`. This will help the neural network predict when the sentence should end.
You can get the `<EOS>` word id by doing:
```python
target_vocab_to_int['<EOS>']
```
You can get other word ids using `source_vocab_to_int` and `target_vocab_to_int`.
```
def text_to_ids(source_text, target_text, source_vocab_to_int, target_vocab_to_int):
"""
Convert source and target text to proper word ids
:param source_text: String that contains all the source text.
:param target_text: String that contains all the target text.
:param source_vocab_to_int: Dictionary to go from the source words to an id
:param target_vocab_to_int: Dictionary to go from the target words to an id
:return: A tuple of lists (source_id_text, target_id_text)
"""
# TODO: Implement Function
source_id_text = [[source_vocab_to_int.get(word, 0) for word in line.split()] for line in source_text.split('\n')]
target_id_text = [[target_vocab_to_int[word] for word in line.split()] + [target_vocab_to_int[ '<EOS>']] for line in target_text.split('\n')]
return source_id_text, target_id_text
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_text_to_ids(text_to_ids)
```
### Preprocess all the data and save it
Running the code cell below will preprocess all the data and save it to file.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
helper.preprocess_and_save_data(source_path, target_path, text_to_ids)
```
# Check Point
This is your first checkpoint. If you ever decide to come back to this notebook or have to restart the notebook, you can start from here. The preprocessed data has been saved to disk.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import numpy as np
import helper
(source_int_text, target_int_text), (source_vocab_to_int, target_vocab_to_int), _ = helper.load_preprocess()
```
### Check the Version of TensorFlow and Access to GPU
This will check to make sure you have the correct version of TensorFlow and access to a GPU
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
from distutils.version import LooseVersion
import warnings
import tensorflow as tf
# Check TensorFlow Version
assert LooseVersion(tf.__version__) >= LooseVersion('1.0'), 'Please use TensorFlow version 1.0 or newer'
print('TensorFlow Version: {}'.format(tf.__version__))
# Check for a GPU
if not tf.test.gpu_device_name():
warnings.warn('No GPU found. Please use a GPU to train your neural network.')
else:
print('Default GPU Device: {}'.format(tf.test.gpu_device_name()))
```
## Build the Neural Network
You'll build the components necessary to build a Sequence-to-Sequence model by implementing the following functions below:
- `model_inputs`
- `process_decoding_input`
- `encoding_layer`
- `decoding_layer_train`
- `decoding_layer_infer`
- `decoding_layer`
- `seq2seq_model`
### Input
Implement the `model_inputs()` function to create TF Placeholders for the Neural Network. It should create the following placeholders:
- Input text placeholder named "input" using the TF Placeholder name parameter with rank 2.
- Targets placeholder with rank 2.
- Learning rate placeholder with rank 0.
- Keep probability placeholder named "keep_prob" using the TF Placeholder name parameter with rank 0.
Return the placeholders in the following the tuple (Input, Targets, Learing Rate, Keep Probability)
```
def model_inputs():
"""
Create TF Placeholders for input, targets, and learning rate.
:return: Tuple (input, targets, learning rate, keep probability)
"""
# TODO: Implement Function\
inputs = tf.placeholder(tf.int32,[None, None], name='input')
target = tf.placeholder(tf.int32,[None, None])
learning_rate = tf.placeholder(tf.float32)
keep_prob = tf.placeholder(tf.float32, name='keep_prob')
return inputs, target, learning_rate, keep_prob
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_model_inputs(model_inputs)
```
### Process Decoding Input
Implement `process_decoding_input` using TensorFlow to remove the last word id from each batch in `target_data` and concat the GO ID to the begining of each batch.
```
def process_decoding_input(target_data, target_vocab_to_int, batch_size):
"""
Preprocess target data for dencoding
:param target_data: Target Placehoder
:param target_vocab_to_int: Dictionary to go from the target words to an id
:param batch_size: Batch Size
:return: Preprocessed target data
"""
# TODO: Implement Function
ending = tf.strided_slice(target_data, [0, 0], [batch_size, -1], [1, 1])
dec_input = tf.concat([tf.fill([batch_size, 1], target_vocab_to_int['<GO>']), ending], 1)
return dec_input
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_process_decoding_input(process_decoding_input)
```
### Encoding
Implement `encoding_layer()` to create a Encoder RNN layer using [`tf.nn.dynamic_rnn()`](https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn).
```
def encoding_layer(rnn_inputs, rnn_size, num_layers, keep_prob):
"""
Create encoding layer
:param rnn_inputs: Inputs for the RNN
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param keep_prob: Dropout keep probability
:return: RNN state
"""
# TODO: Implement Function
enc_cell = tf.contrib.rnn.MultiRNNCell([tf.contrib.rnn.BasicLSTMCell(rnn_size)] * num_layers)
dropout = tf.contrib.rnn.DropoutWrapper(enc_cell, output_keep_prob=keep_prob)
_, enc_state = tf.nn.dynamic_rnn(dropout, rnn_inputs, dtype=tf.float32)
return enc_state
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_encoding_layer(encoding_layer)
```
### Decoding - Training
Create training logits using [`tf.contrib.seq2seq.simple_decoder_fn_train()`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/simple_decoder_fn_train) and [`tf.contrib.seq2seq.dynamic_rnn_decoder()`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/dynamic_rnn_decoder). Apply the `output_fn` to the [`tf.contrib.seq2seq.dynamic_rnn_decoder()`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/dynamic_rnn_decoder) outputs.
```
def decoding_layer_train(encoder_state, dec_cell, dec_embed_input, sequence_length, decoding_scope,
output_fn, keep_prob):
"""
Create a decoding layer for training
:param encoder_state: Encoder State
:param dec_cell: Decoder RNN Cell
:param dec_embed_input: Decoder embedded input
:param sequence_length: Sequence Length
:param decoding_scope: TenorFlow Variable Scope for decoding
:param output_fn: Function to apply the output layer
:param keep_prob: Dropout keep probability
:return: Train Logits
"""
# TODO: Implement Function
train_decoder_fn = tf.contrib.seq2seq.simple_decoder_fn_train(encoder_state)
train_pred, _, _ = tf.contrib.seq2seq.dynamic_rnn_decoder(dec_cell, train_decoder_fn, dec_embed_input, sequence_length, scope=decoding_scope)
train_logits = output_fn(train_pred)
return train_logits
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer_train(decoding_layer_train)
```
### Decoding - Inference
Create inference logits using [`tf.contrib.seq2seq.simple_decoder_fn_inference()`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/simple_decoder_fn_inference) and [`tf.contrib.seq2seq.dynamic_rnn_decoder()`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/dynamic_rnn_decoder).
```
def decoding_layer_infer(encoder_state, dec_cell, dec_embeddings, start_of_sequence_id, end_of_sequence_id,
maximum_length, vocab_size, decoding_scope, output_fn, keep_prob):
"""
Create a decoding layer for inference
:param encoder_state: Encoder state
:param dec_cell: Decoder RNN Cell
:param dec_embeddings: Decoder embeddings
:param start_of_sequence_id: GO ID
:param end_of_sequence_id: EOS Id
:param maximum_length: Maximum length of
:param vocab_size: Size of vocabulary
:param decoding_scope: TensorFlow Variable Scope for decoding
:param output_fn: Function to apply the output layer
:param keep_prob: Dropout keep probability
:return: Inference Logits
"""
# TODO: Implement Function
infer_decoder_fn = tf.contrib.seq2seq.simple_decoder_fn_inference(output_fn, encoder_state, dec_embeddings,
start_of_sequence_id, end_of_sequence_id,
maximum_length, vocab_size, dtype=tf.int32)
inference_logits, _, _ = tf.contrib.seq2seq.dynamic_rnn_decoder(dec_cell, infer_decoder_fn, scope=decoding_scope)
inference_logits = tf.contrib.layers.dropout(inference_logits, keep_prob)
return inference_logits
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer_infer(decoding_layer_infer)
```
### Build the Decoding Layer
Implement `decoding_layer()` to create a Decoder RNN layer.
- Create RNN cell for decoding using `rnn_size` and `num_layers`.
- Create the output fuction using [`lambda`](https://docs.python.org/3/tutorial/controlflow.html#lambda-expressions) to transform it's input, logits, to class logits.
- Use the your `decoding_layer_train(encoder_state, dec_cell, dec_embed_input, sequence_length, decoding_scope, output_fn, keep_prob)` function to get the training logits.
- Use your `decoding_layer_infer(encoder_state, dec_cell, dec_embeddings, start_of_sequence_id, end_of_sequence_id, maximum_length, vocab_size, decoding_scope, output_fn, keep_prob)` function to get the inference logits.
Note: You'll need to use [tf.variable_scope](https://www.tensorflow.org/api_docs/python/tf/variable_scope) to share variables between training and inference.
```
def decoding_layer(dec_embed_input, dec_embeddings, encoder_state, vocab_size, sequence_length, rnn_size,
num_layers, target_vocab_to_int, keep_prob):
"""
Create decoding layer
:param dec_embed_input: Decoder embedded input
:param dec_embeddings: Decoder embeddings
:param encoder_state: The encoded state
:param vocab_size: Size of vocabulary
:param sequence_length: Sequence Length
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param target_vocab_to_int: Dictionary to go from the target words to an id
:param keep_prob: Dropout keep probability
:return: Tuple of (Training Logits, Inference Logits)
"""
# TODO: Implement Function
start_of_sequence_id = target_vocab_to_int['<GO>']
end_of_sequence_id = target_vocab_to_int['<EOS>']
lstm = tf.contrib.rnn.BasicLSTMCell(rnn_size)
dec_cell = tf.contrib.rnn.MultiRNNCell([lstm]*num_layers)
dec_cell = tf.contrib.rnn.DropoutWrapper(dec_cell, output_keep_prob=keep_prob)
# Output Layer
output_fn = lambda x: tf.contrib.layers.fully_connected(x, vocab_size, None, scope=decoding_scope)
with tf.variable_scope('decoding') as decoding_scope:
# Training Decoder
training_logits = decoding_layer_train(encoder_state, dec_cell, dec_embed_input, sequence_length,
decoding_scope, output_fn, keep_prob)
with tf.variable_scope("decoding", reuse=True) as decoding_scope:
# Inference Decoder
inference_logits = decoding_layer_infer(encoder_state, dec_cell, dec_embeddings, start_of_sequence_id,
end_of_sequence_id, sequence_length, vocab_size,
decoding_scope, output_fn, keep_prob)
return training_logits, inference_logits
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer(decoding_layer)
```
### Build the Neural Network
Apply the functions you implemented above to:
- Apply embedding to the input data for the encoder.
- Encode the input using your `encoding_layer(rnn_inputs, rnn_size, num_layers, keep_prob)`.
- Process target data using your `process_decoding_input(target_data, target_vocab_to_int, batch_size)` function.
- Apply embedding to the target data for the decoder.
- Decode the encoded input using your `decoding_layer(dec_embed_input, dec_embeddings, encoder_state, vocab_size, sequence_length, rnn_size, num_layers, target_vocab_to_int, keep_prob)`.
```
def seq2seq_model(input_data, target_data, keep_prob, batch_size, sequence_length, source_vocab_size, target_vocab_size,
enc_embedding_size, dec_embedding_size, rnn_size, num_layers, target_vocab_to_int):
"""
Build the Sequence-to-Sequence part of the neural network
:param input_data: Input placeholder
:param target_data: Target placeholder
:param keep_prob: Dropout keep probability placeholder
:param batch_size: Batch Size
:param sequence_length: Sequence Length
:param source_vocab_size: Source vocabulary size
:param target_vocab_size: Target vocabulary size
:param enc_embedding_size: Decoder embedding size
:param dec_embedding_size: Encoder embedding size
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param target_vocab_to_int: Dictionary to go from the target words to an id
:return: Tuple of (Training Logits, Inference Logits)
"""
# TODO: Implement Function
rnn_inputs = tf.contrib.layers.embed_sequence(input_data, source_vocab_size, enc_embedding_size)
encoder_state = encoding_layer(rnn_inputs, rnn_size, num_layers, keep_prob)
dec_input = process_decoding_input(target_data, target_vocab_to_int, batch_size)
dec_embeddings = tf.Variable(tf.random_uniform([target_vocab_size, dec_embedding_size]))
dec_embed_input = tf.nn.embedding_lookup(dec_embeddings, dec_input)
training_logits, inference_logits = decoding_layer(dec_embed_input, dec_embeddings,
encoder_state, target_vocab_size, sequence_length,
rnn_size, num_layers, target_vocab_to_int, keep_prob)
return training_logits, inference_logits
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_seq2seq_model(seq2seq_model)
```
## Neural Network Training
### Hyperparameters
Tune the following parameters:
- Set `epochs` to the number of epochs.
- Set `batch_size` to the batch size.
- Set `rnn_size` to the size of the RNNs.
- Set `num_layers` to the number of layers.
- Set `encoding_embedding_size` to the size of the embedding for the encoder.
- Set `decoding_embedding_size` to the size of the embedding for the decoder.
- Set `learning_rate` to the learning rate.
- Set `keep_probability` to the Dropout keep probability
```
# Number of Epochs
epochs = 20
# Batch Size
batch_size = 512
# RNN Size
rnn_size = 256
# Number of Layers
num_layers = 2
# Embedding Size
encoding_embedding_size = 200
decoding_embedding_size = 200
# Learning Rate
learning_rate = 0.001
# Dropout Keep Probability
keep_probability = 0.5
```
### Build the Graph
Build the graph using the neural network you implemented.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
save_path = 'checkpoints/dev'
(source_int_text, target_int_text), (source_vocab_to_int, target_vocab_to_int), _ = helper.load_preprocess()
max_target_sentence_length = max([len(sentence) for sentence in source_int_text])
train_graph = tf.Graph()
with train_graph.as_default():
input_data, targets, lr, keep_prob = model_inputs()
sequence_length = tf.placeholder_with_default(max_target_sentence_length, None, name='sequence_length')
input_shape = tf.shape(input_data)
train_logits, inference_logits = seq2seq_model(
tf.reverse(input_data, [-1]), targets, keep_prob, batch_size, sequence_length, len(source_vocab_to_int), len(target_vocab_to_int),
encoding_embedding_size, decoding_embedding_size, rnn_size, num_layers, target_vocab_to_int)
tf.identity(inference_logits, 'logits')
with tf.name_scope("optimization"):
# Loss function
cost = tf.contrib.seq2seq.sequence_loss(
train_logits,
targets,
tf.ones([input_shape[0], sequence_length]))
# Optimizer
optimizer = tf.train.AdamOptimizer(lr)
# Gradient Clipping
gradients = optimizer.compute_gradients(cost)
capped_gradients = [(tf.clip_by_value(grad, -1., 1.), var) for grad, var in gradients if grad is not None]
train_op = optimizer.apply_gradients(capped_gradients)
```
### Train
Train the neural network on the preprocessed data. If you have a hard time getting a good loss, check the forms to see if anyone is having the same problem.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import time
def get_accuracy(target, logits):
"""
Calculate accuracy
"""
max_seq = max(target.shape[1], logits.shape[1])
if max_seq - target.shape[1]:
target = np.pad(
target_batch,
[(0,0),(0,max_seq - target_batch.shape[1]), (0,0)],
'constant')
if max_seq - batch_train_logits.shape[1]:
logits = np.pad(
logits,
[(0,0),(0,max_seq - logits.shape[1]), (0,0)],
'constant')
return np.mean(np.equal(target, np.argmax(logits, 2)))
train_source = source_int_text[batch_size:]
train_target = target_int_text[batch_size:]
valid_source = helper.pad_sentence_batch(source_int_text[:batch_size])
valid_target = helper.pad_sentence_batch(target_int_text[:batch_size])
with tf.Session(graph=train_graph) as sess:
sess.run(tf.global_variables_initializer())
for epoch_i in range(epochs):
for batch_i, (source_batch, target_batch) in enumerate(
helper.batch_data(train_source, train_target, batch_size)):
start_time = time.time()
_, loss = sess.run(
[train_op, cost],
{input_data: source_batch,
targets: target_batch,
lr: learning_rate,
sequence_length: target_batch.shape[1],
keep_prob: keep_probability})
batch_train_logits = sess.run(
inference_logits,
{input_data: source_batch, keep_prob: 1.0})
batch_valid_logits = sess.run(
inference_logits,
{input_data: valid_source, keep_prob: 1.0})
train_acc = get_accuracy(target_batch, batch_train_logits)
valid_acc = get_accuracy(np.array(valid_target), batch_valid_logits)
end_time = time.time()
print('Epoch {:>3} Batch {:>4}/{} - Train Accuracy: {:>6.3f}, Validation Accuracy: {:>6.3f}, Loss: {:>6.3f}'
.format(epoch_i, batch_i, len(source_int_text) // batch_size, train_acc, valid_acc, loss))
# Save Model
saver = tf.train.Saver()
saver.save(sess, save_path)
print('Model Trained and Saved')
```
### Save Parameters
Save the `batch_size` and `save_path` parameters for inference.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# Save parameters for checkpoint
helper.save_params(save_path)
```
# Checkpoint
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import tensorflow as tf
import numpy as np
import helper
import problem_unittests as tests
_, (source_vocab_to_int, target_vocab_to_int), (source_int_to_vocab, target_int_to_vocab) = helper.load_preprocess()
load_path = helper.load_params()
```
## Sentence to Sequence
To feed a sentence into the model for translation, you first need to preprocess it. Implement the function `sentence_to_seq()` to preprocess new sentences.
- Convert the sentence to lowercase
- Convert words into ids using `vocab_to_int`
- Convert words not in the vocabulary, to the `<UNK>` word id.
```
def sentence_to_seq(sentence, vocab_to_int):
"""
Convert a sentence to a sequence of ids
:param sentence: String
:param vocab_to_int: Dictionary to go from the words to an id
:return: List of word ids
"""
# TODO: Implement Function
input_sentence = sentence.split()
input_sentence = [vocab_to_int.get(word.lower(), vocab_to_int['<UNK>']) for word in input_sentence]
return input_sentence
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_sentence_to_seq(sentence_to_seq)
```
## Translate
This will translate `translate_sentence` from English to French.
```
translate_sentence = 'he saw a yellow truck .'
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
translate_sentence = sentence_to_seq(translate_sentence, source_vocab_to_int)
loaded_graph = tf.Graph()
with tf.Session(graph=loaded_graph) as sess:
# Load saved model
loader = tf.train.import_meta_graph(load_path + '.meta')
loader.restore(sess, load_path)
input_data = loaded_graph.get_tensor_by_name('input:0')
logits = loaded_graph.get_tensor_by_name('logits:0')
keep_prob = loaded_graph.get_tensor_by_name('keep_prob:0')
translate_logits = sess.run(logits, {input_data: [translate_sentence], keep_prob: 1.0})[0]
print('Input')
print(' Word Ids: {}'.format([i for i in translate_sentence]))
print(' English Words: {}'.format([source_int_to_vocab[i] for i in translate_sentence]))
print('\nPrediction')
print(' Word Ids: {}'.format([i for i in np.argmax(translate_logits, 1)]))
print(' French Words: {}'.format([target_int_to_vocab[i] for i in np.argmax(translate_logits, 1)]))
```
## Imperfect Translation
You might notice that some sentences translate better than others. Since the dataset you're using only has a vocabulary of 227 English words of the thousands that you use, you're only going to see good results using these words. For this project, you don't need a perfect translation. However, if you want to create a better translation model, you'll need better data.
You can train on the [WMT10 French-English corpus](http://www.statmt.org/wmt10/training-giga-fren.tar). This dataset has more vocabulary and richer in topics discussed. However, this will take you days to train, so make sure you've a GPU and the neural network is performing well on dataset we provided. Just make sure you play with the WMT10 corpus after you've submitted this project.
## Submitting This Project
When submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as "dlnd_language_translation.ipynb" and save it as a HTML file under "File" -> "Download as". Include the "helper.py" and "problem_unittests.py" files in your submission.
| github_jupyter |
# Hierarchical radon model
(c) 2019 Thomas Wiecki, Junpeng Lao
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pymc4 as pm
import pandas as pd
import tensorflow as tf
import arviz as az
data = pd.read_csv('/home/junpenglao/Documents/pymc3/docs/source/data/radon.csv')
county_names = data.county.unique()
county_idx = data['county_code'].values.astype(np.int32)
@pm.model
def hierarchical_model(data, county_idx):
# Hyperpriors
mu_a = yield pm.Normal('mu_alpha', mu=0., sigma=1)
sigma_a = yield pm.HalfCauchy('sigma_alpha', beta=1)
mu_b = yield pm.Normal('mu_beta', mu=0., sigma=1)
sigma_b = yield pm.HalfCauchy('sigma_beta', beta=1)
# Intercept for each county, distributed around group mean mu_a
a = yield pm.Normal('alpha', mu=mu_a, sigma=sigma_a, plate=len(data.county.unique()))
# Intercept for each county, distributed around group mean mu_a
b = yield pm.Normal('beta', mu=mu_b, sigma=sigma_b, plate=len(data.county.unique()))
# Model error
eps = yield pm.HalfCauchy('eps', beta=1)
# Expected value
#radon_est = a[county_idx] + b[county_idx] * data.floor.values
radon_est = tf.gather(a, county_idx) + tf.gather(
b, county_idx) * data.floor.values
# Data likelihood
y_like = yield pm.Normal('y_like', mu=radon_est, sigma=eps, observed=data.log_radon)
%%time
init_num_chains = 50
model = hierarchical_model(data, county_idx)
pm4_trace, _ = pm.inference.sampling.sample(
model, num_chains=init_num_chains, num_samples=10, burn_in=10, step_size=1., xla=False)
```
### Mass matrix adaptation
Essentially, we are doing a 2 window adaptation similar to Stan below: dual averaging for burn_in samples, and then normal sample for diagonal mass matrix estimation.
```
%%time
for i in range(3):
step_size_ = []
for _, x in pm4_trace.items():
std = tf.math.reduce_std(x, axis=[0, 1])
step_size_.append(
std[tf.newaxis, ...] * tf.ones([init_num_chains] + std.shape, dtype=std.dtype))
pm4_trace, _ = pm.inference.sampling.sample(
model, num_chains=init_num_chains, num_samples=10 + 10*i, burn_in=10 + 10*i,
step_size=step_size_, xla=False)
%%time
num_chains = 5
step_size_ = []
for _, x in pm4_trace.items():
std = tf.math.reduce_std(x, axis=[0, 1])
step_size_.append(
std[tf.newaxis, ...] * tf.ones([num_chains]+std.shape, dtype=std.dtype))
pm4_trace, sample_stat = pm.inference.sampling.sample(
model, num_chains=num_chains, num_samples=500, burn_in=500,
step_size=step_size_, xla=False)
az_trace = pm.inference.utils.trace_to_arviz(pm4_trace, sample_stat)
np.mean(np.exp(az_trace.sample_stats['mean_tree_accept']), axis=1)
plt.hist(np.ravel(az_trace.sample_stats['tree_size']), 50);
az.summary(az_trace)
az.plot_energy(az_trace);
az.plot_trace(az_trace, compact=True, combined=True);
```
| github_jupyter |
```
from scipy.sparse import csr_matrix
import implicit
import pandas as pd
from tqdm.notebook import tqdm
import matplotlib.pyplot as plt
from metrics import mean_average_presision_k, hitrate_k
# from sacred.observers import
# Load data
path = '/data/pet_ML/groupLe_recsys/gl/'
df_prefix = '30k/'
test_df = pd.read_csv(path + df_prefix + 'test/views.csv')
train_df = pd.read_csv(path + df_prefix + 'train/views.csv')
train_df.head(2)
train_df.item_id.drop_duplicates().hist(bins=100, figsize=(20, 3))
test_df.item_id.drop_duplicates().hist(bins=100, figsize=(20, 3))
# delete negative samples
train_df.shape
d = train_df.groupby('user_id').sum()['rate']
dislikers = d[d <= 0].index.to_list()
test_items = test_df.item_id.drop_duplicates()
train_df = train_df[train_df.item_id.isin(test_items) & (~train_df.user_id.isin(dislikers))]
test_df = test_df[(test_df.user_id.isin(train_df.user_id.drop_duplicates()))]
train_df.shape
calc_preference = lambda v: 1 if v>0 else -10
train_df['metric']= train_df.rate.apply(calc_preference)
alpha = 10
train_df.metric = train_df.metric * alpha * train_df.rate
train_df.rate = train_df.metric
train_df = train_df.drop('metric', axis=1)
df_users = pd.read_csv(path + df_prefix + 'users.csv')
df_users.head(2)
df_manga = pd.read_csv(path+ df_prefix + 'manga.csv')
df_manga.head(2)
```
# Filter data
```
train_df.groupby('user_id').count()['rate'].hist(bins=100)
```
# indexing data sequentially
```
users = sorted(train_df.user_id.drop_duplicates().to_list())
items = sorted(train_df.item_id.drop_duplicates().to_list())
print(len(users), len(items))
users_pivot = pd.DataFrame.from_records([(uid, i) for i, uid in enumerate(users)], columns='user_id user_index'.split())
items_pivot = pd.DataFrame.from_records([(iid, i) for i, iid in enumerate(items)], columns='item_id item_index'.split())
users_pivot.head()
train_df = pd.merge(train_df, users_pivot, on='user_id')
train_df = pd.merge(train_df, items_pivot, on='item_id')
train_df.head()
indexed_df = train_df.drop('user_id item_id'.split(), axis=1)
indexed_df.head()
users = indexed_df.user_index.to_list()
items = indexed_df.item_index.to_list()
rate = indexed_df.rate.to_list()
shape = (len(set(users)), len(set(items)))
ui_mat = csr_matrix((rate, (users, items)), shape=shape)
ui_mat.shape
# initialize a model
model = implicit.als.AlternatingLeastSquares(factors=10, calculate_training_loss=True, iterations=20)
# train the model on a sparse matrix of item/user/confidence weights
model.fit(ui_mat)
```
# Calc metrics
```
ui_mat.shape
recs = []
k = 100
for uid, uix in tqdm(list(users_pivot.to_records(index=False))):
recs.append(model.recommend(uix, ui_mat, N=k, filter_already_liked_items=False))
gt = test_df.groupby('user_id')['item_id'].apply(list).tolist()
items_pivot_dict = {rec_id : item_id for item_id, rec_id in list(items_pivot.to_records(index=False))}
# select only items ids and map it to real ids. Without mapping cant calculating metrics with gt dataframe
rec_items = list(map(lambda rec_list: [items_pivot_dict[x[0]] for x in rec_list], recs))
mean_average_presision_k(rec_items, gt, k=k)
hitrate_k(rec_items, gt, k=k)
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Distributed training with TensorFlow
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/distributed_training"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/distributed_training.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/distributed_training.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/distributed_training.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
## Overview
`tf.distribute.Strategy` is a TensorFlow API to distribute training
across multiple GPUs, multiple machines or TPUs. Using this API, you can distribute your existing models and training code with minimal code changes.
`tf.distribute.Strategy` has been designed with these key goals in mind:
* Easy to use and support multiple user segments, including researchers, ML engineers, etc.
* Provide good performance out of the box.
* Easy switching between strategies.
`tf.distribute.Strategy` can be used with a high-level API like [Keras](https://www.tensorflow.org/guide/keras), and can also be used to distribute custom training loops (and, in general, any computation using TensorFlow).
In TensorFlow 2.0, you can execute your programs eagerly, or in a graph using [`tf.function`](function.ipynb). `tf.distribute.Strategy` intends to support both these modes of execution. Although we discuss training most of the time in this guide, this API can also be used for distributing evaluation and prediction on different platforms.
You can use `tf.distribute.Strategy` with very few changes to your code, because we have changed the underlying components of TensorFlow to become strategy-aware. This includes variables, layers, models, optimizers, metrics, summaries, and checkpoints.
In this guide, we explain various types of strategies and how you can use them in different situations.
Note: For a deeper understanding of the concepts, please watch [this deep-dive presentation](https://youtu.be/jKV53r9-H14). This is especially recommended if you plan to write your own training loop.
```
# Import TensorFlow
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
```
## Types of strategies
`tf.distribute.Strategy` intends to cover a number of use cases along different axes. Some of these combinations are currently supported and others will be added in the future. Some of these axes are:
* *Synchronous vs asynchronous training:* These are two common ways of distributing training with data parallelism. In sync training, all workers train over different slices of input data in sync, and aggregating gradients at each step. In async training, all workers are independently training over the input data and updating variables asynchronously. Typically sync training is supported via all-reduce and async through parameter server architecture.
* *Hardware platform:* You may want to scale your training onto multiple GPUs on one machine, or multiple machines in a network (with 0 or more GPUs each), or on Cloud TPUs.
In order to support these use cases, there are six strategies available. In the next section we explain which of these are supported in which scenarios in TF 2.0 at this time. Here is a quick overview:
| Training API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy | OneDeviceStrategy |
|:----------------------- |:------------------- |:--------------------- |:--------------------------------- |:--------------------------------- |:-------------------------- | :-------------------------------------- |
| **Keras API** | Supported | Experimental support | Experimental support | Experimental support | Supported planned post 2.0 | Supported |
| **Custom training loop** | Experimental support | Experimental support | Support planned post 2.0 | Support planned post 2.0 | No support yet | Supported |
| **Estimator API** | Limited Support | Not supported | Limited Support | Limited Support | Limited Support | Limited Support |
Note: Estimator support is limited. Basic training and evaluation are experimental, and advanced features—such as scaffold—are not implemented. We recommend using Keras or custom training loops if a use case is not covered.
### MirroredStrategy
`tf.distribute.MirroredStrategy` supports synchronous distributed training on multiple GPUs on one machine. It creates one replica per GPU device. Each variable in the model is mirrored across all the replicas. Together, these variables form a single conceptual variable called `MirroredVariable`. These variables are kept in sync with each other by applying identical updates.
Efficient all-reduce algorithms are used to communicate the variable updates across the devices.
All-reduce aggregates tensors across all the devices by adding them up, and makes them available on each device.
It’s a fused algorithm that is very efficient and can reduce the overhead of synchronization significantly. There are many all-reduce algorithms and implementations available, depending on the type of communication available between devices. By default, it uses NVIDIA NCCL as the all-reduce implementation. You can choose from a few other options we provide, or write your own.
Here is the simplest way of creating `MirroredStrategy`:
```
mirrored_strategy = tf.distribute.MirroredStrategy()
```
This will create a `MirroredStrategy` instance which will use all the GPUs that are visible to TensorFlow, and use NCCL as the cross device communication.
If you wish to use only some of the GPUs on your machine, you can do so like this:
```
mirrored_strategy = tf.distribute.MirroredStrategy(devices=["/gpu:0", "/gpu:1"])
```
If you wish to override the cross device communication, you can do so using the `cross_device_ops` argument by supplying an instance of `tf.distribute.CrossDeviceOps`. Currently, `tf.distribute.HierarchicalCopyAllReduce` and `tf.distribute.ReductionToOneDevice` are two options other than `tf.distribute.NcclAllReduce` which is the default.
```
mirrored_strategy = tf.distribute.MirroredStrategy(
cross_device_ops=tf.distribute.HierarchicalCopyAllReduce())
```
### CentralStorageStrategy
`tf.distribute.experimental.CentralStorageStrategy` does synchronous training as well. Variables are not mirrored, instead they are placed on the CPU and operations are replicated across all local GPUs. If there is only one GPU, all variables and operations will be placed on that GPU.
Create an instance of `CentralStorageStrategy` by:
```
central_storage_strategy = tf.distribute.experimental.CentralStorageStrategy()
```
This will create a `CentralStorageStrategy` instance which will use all visible GPUs and CPU. Update to variables on replicas will be aggregated before being applied to variables.
Note: This strategy is [`experimental`](https://www.tensorflow.org/guide/versions#what_is_not_covered) as we are currently improving it and making it work for more scenarios. As part of this, please expect the APIs to change in the future.
### MultiWorkerMirroredStrategy
`tf.distribute.experimental.MultiWorkerMirroredStrategy` is very similar to `MirroredStrategy`. It implements synchronous distributed training across multiple workers, each with potentially multiple GPUs. Similar to `MirroredStrategy`, it creates copies of all variables in the model on each device across all workers.
It uses [CollectiveOps](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/ops/collective_ops.py) as the multi-worker all-reduce communication method used to keep variables in sync. A collective op is a single op in the TensorFlow graph which can automatically choose an all-reduce algorithm in the TensorFlow runtime according to hardware, network topology and tensor sizes.
It also implements additional performance optimizations. For example, it includes a static optimization that converts multiple all-reductions on small tensors into fewer all-reductions on larger tensors. In addition, we are designing it to have a plugin architecture - so that in the future, you will be able to plugin algorithms that are better tuned for your hardware. Note that collective ops also implement other collective operations such as broadcast and all-gather.
Here is the simplest way of creating `MultiWorkerMirroredStrategy`:
```
multiworker_strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
```
`MultiWorkerMirroredStrategy` currently allows you to choose between two different implementations of collective ops. `CollectiveCommunication.RING` implements ring-based collectives using gRPC as the communication layer. `CollectiveCommunication.NCCL` uses [Nvidia's NCCL](https://developer.nvidia.com/nccl) to implement collectives. `CollectiveCommunication.AUTO` defers the choice to the runtime. The best choice of collective implementation depends upon the number and kind of GPUs, and the network interconnect in the cluster. You can specify them in the following way:
```
multiworker_strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy(
tf.distribute.experimental.CollectiveCommunication.NCCL)
```
One of the key differences to get multi worker training going, as compared to multi-GPU training, is the multi-worker setup. The `TF_CONFIG` environment variable is the standard way in TensorFlow to specify the cluster configuration to each worker that is part of the cluster. Learn more about [setting up TF_CONFIG](#TF_CONFIG).
Note: This strategy is [`experimental`](https://www.tensorflow.org/guide/versions#what_is_not_covered) as we are currently improving it and making it work for more scenarios. As part of this, please expect the APIs to change in the future.
### TPUStrategy
`tf.distribute.experimental.TPUStrategy` lets you run your TensorFlow training on Tensor Processing Units (TPUs). TPUs are Google's specialized ASICs designed to dramatically accelerate machine learning workloads. They are available on Google Colab, the [TensorFlow Research Cloud](https://www.tensorflow.org/tfrc) and [Cloud TPU](https://cloud.google.com/tpu).
In terms of distributed training architecture, `TPUStrategy` is the same `MirroredStrategy` - it implements synchronous distributed training. TPUs provide their own implementation of efficient all-reduce and other collective operations across multiple TPU cores, which are used in `TPUStrategy`.
Here is how you would instantiate `TPUStrategy`:
Note: To run this code in Colab, you should select TPU as the Colab runtime. We will have a tutorial soon that will demonstrate how you can use TPUStrategy.
```
cluster_resolver = tf.distribute.cluster_resolver.TPUClusterResolver(
tpu=tpu_address)
tf.config.experimental_connect_to_cluster(cluster_resolver)
tf.tpu.experimental.initialize_tpu_system(cluster_resolver)
tpu_strategy = tf.distribute.experimental.TPUStrategy(cluster_resolver)
```
The `TPUClusterResolver` instance helps locate the TPUs. In Colab, you don't need to specify any arguments to it.
If you want to use this for Cloud TPUs:
- You must specify the name of your TPU resource in the `tpu` argument.
- You must initialize the tpu system explicitly at the *start* of the program. This is required before TPUs can be used for computation. Initializing the tpu system also wipes out the TPU memory, so it's important to complete this step first in order to avoid losing state.
Note: This strategy is [`experimental`](https://www.tensorflow.org/guide/versions#what_is_not_covered) as we are currently improving it and making it work for more scenarios. As part of this, please expect the APIs to change in the future.
### ParameterServerStrategy
`tf.distribute.experimental.ParameterServerStrategy` supports parameter servers training on multiple machines. In this setup, some machines are designated as workers and some as parameter servers. Each variable of the model is placed on one parameter server. Computation is replicated across all GPUs of all the workers.
In terms of code, it looks similar to other strategies:
```
ps_strategy = tf.distribute.experimental.ParameterServerStrategy()
```
For multi worker training, `TF_CONFIG` needs to specify the configuration of parameter servers and workers in your cluster, which you can read more about in [TF_CONFIG below](#TF_CONFIG) below.
### OneDeviceStrategy
`tf.distribute.OneDeviceStrategy` runs on a single device. This strategy will place any variables created in its scope on the specified device. Input distributed through this strategy will be prefetched to the specified device. Moreover, any functions called via `strategy.experimental_run_v2` will also be placed on the specified device.
You can use this strategy to test your code before switching to other strategies which actually distributes to multiple devices/machines.
```
strategy = tf.distribute.OneDeviceStrategy(device="/gpu:0")
```
So far we've talked about what are the different strategies available and how you can instantiate them. In the next few sections, we will talk about the different ways in which you can use them to distribute your training. We will show short code snippets in this guide and link off to full tutorials which you can run end to end.
## Using `tf.distribute.Strategy` with Keras
We've integrated `tf.distribute.Strategy` into `tf.keras` which is TensorFlow's implementation of the
[Keras API specification](https://keras.io). `tf.keras` is a high-level API to build and train models. By integrating into `tf.keras` backend, we've made it seamless for you to distribute your training written in the Keras training framework.
Here's what you need to change in your code:
1. Create an instance of the appropriate `tf.distribute.Strategy`
2. Move the creation and compiling of Keras model inside `strategy.scope`.
We support all types of Keras models - sequential, functional and subclassed.
Here is a snippet of code to do this for a very simple Keras model with one dense layer:
```
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = tf.keras.Sequential([tf.keras.layers.Dense(1, input_shape=(1,))])
model.compile(loss='mse', optimizer='sgd')
```
In this example we used `MirroredStrategy` so we can run this on a machine with multiple GPUs. `strategy.scope()` indicated which parts of the code to run distributed. Creating a model inside this scope allows us to create mirrored variables instead of regular variables. Compiling under the scope allows us to know that the user intends to train this model using this strategy. Once this is set up, you can fit your model like you would normally. `MirroredStrategy` takes care of replicating the model's training on the available GPUs, aggregating gradients, and more.
```
dataset = tf.data.Dataset.from_tensors(([1.], [1.])).repeat(100).batch(10)
model.fit(dataset, epochs=2)
model.evaluate(dataset)
```
Here we used a `tf.data.Dataset` to provide the training and eval input. You can also use numpy arrays:
```
import numpy as np
inputs, targets = np.ones((100, 1)), np.ones((100, 1))
model.fit(inputs, targets, epochs=2, batch_size=10)
```
In both cases (dataset or numpy), each batch of the given input is divided equally among the multiple replicas. For instance, if using `MirroredStrategy` with 2 GPUs, each batch of size 10 will get divided among the 2 GPUs, with each receiving 5 input examples in each step. Each epoch will then train faster as you add more GPUs. Typically, you would want to increase your batch size as you add more accelerators so as to make effective use of the extra computing power. You will also need to re-tune your learning rate, depending on the model. You can use `strategy.num_replicas_in_sync` to get the number of replicas.
```
# Compute global batch size using number of replicas.
BATCH_SIZE_PER_REPLICA = 5
global_batch_size = (BATCH_SIZE_PER_REPLICA *
mirrored_strategy.num_replicas_in_sync)
dataset = tf.data.Dataset.from_tensors(([1.], [1.])).repeat(100)
dataset = dataset.batch(global_batch_size)
LEARNING_RATES_BY_BATCH_SIZE = {5: 0.1, 10: 0.15}
learning_rate = LEARNING_RATES_BY_BATCH_SIZE[global_batch_size]
```
### What's supported now?
In TF 2.0 release, `MirroredStrategy`, `TPUStrategy`, `CentralStorageStrategy` and `MultiWorkerMirroredStrategy` are supported in Keras. Except `MirroredStrategy`, others are currently experimental and are subject to change.
Support for other strategies will be coming soon. The API and how to use will be exactly the same as above.
| Training API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy | OneDeviceStrategy |
|---------------- |--------------------- |----------------------- |----------------------------------- |----------------------------------- |--------------------------- | --------------------------------------- |
| Keras APIs | Supported | Experimental support | Experimental support | Experimental support | Support planned post 2.0 | Supported |
### Examples and Tutorials
Here is a list of tutorials and examples that illustrate the above integration end to end with Keras:
1. Tutorial to train [MNIST](../tutorials/distribute/keras.ipynb) with `MirroredStrategy`.
2. Official [ResNet50](https://github.com/tensorflow/models/blob/master/official/vision/image_classification/resnet_imagenet_main.py) training with ImageNet data using `MirroredStrategy`.
3. [ResNet50](https://github.com/tensorflow/tpu/blob/master/models/experimental/resnet50_keras/resnet50_tf2.py) trained with Imagenet data on Cloud TPUs with `TPUStrategy`.
4. [Tutorial](../tutorials/distribute/multi_worker_with_keras.ipynb) to train MNIST using `MultiWorkerMirroredStrategy`.
5. [NCF](https://github.com/tensorflow/models/blob/master/official/recommendation/ncf_keras_main.py) trained using `MirroredStrategy`.
2. [Transformer]( https://github.com/tensorflow/models/blob/master/official/transformer/v2/transformer_main.py) trained using `MirroredStrategy`.
## Using `tf.distribute.Strategy` with custom training loops
As you've seen, using `tf.distribute.Strategy` with high-level APIs (Estimator and Keras) requires changing only a couple lines of your code. With a little more effort, you can also use `tf.distribute.Strategy` with custom training loops.
If you need more flexibility and control over your training loops than is possible with Estimator or Keras, you can write custom training loops. For instance, when using a GAN, you may want to take a different number of generator or discriminator steps each round. Similarly, the high level frameworks are not very suitable for Reinforcement Learning training.
To support custom training loops, we provide a core set of methods through the `tf.distribute.Strategy` classes. Using these may require minor restructuring of the code initially, but once that is done, you should be able to switch between GPUs, TPUs, and multiple machines simply by changing the strategy instance.
Here we will show a brief snippet illustrating this use case for a simple training example using the same Keras model as before.
First, we create the model and optimizer inside the strategy's scope. This ensures that any variables created with the model and optimizer are mirrored variables.
```
with mirrored_strategy.scope():
model = tf.keras.Sequential([tf.keras.layers.Dense(1, input_shape=(1,))])
optimizer = tf.keras.optimizers.SGD()
```
Next, we create the input dataset and call `tf.distribute.Strategy.experimental_distribute_dataset` to distribute the dataset based on the strategy.
```
dataset = tf.data.Dataset.from_tensors(([1.], [1.])).repeat(1000).batch(
global_batch_size)
dist_dataset = mirrored_strategy.experimental_distribute_dataset(dataset)
```
Then, we define one step of the training. We will use `tf.GradientTape` to compute gradients and optimizer to apply those gradients to update our model's variables. To distribute this training step, we put in a function `step_fn` and pass it to `tf.distrbute.Strategy.experimental_run_v2` along with the dataset inputs that we get from `dist_dataset` created before:
```
@tf.function
def train_step(dist_inputs):
def step_fn(inputs):
features, labels = inputs
with tf.GradientTape() as tape:
logits = model(features)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(
logits=logits, labels=labels)
loss = tf.reduce_sum(cross_entropy) * (1.0 / global_batch_size)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(list(zip(grads, model.trainable_variables)))
return cross_entropy
per_example_losses = mirrored_strategy.experimental_run_v2(
step_fn, args=(dist_inputs,))
mean_loss = mirrored_strategy.reduce(
tf.distribute.ReduceOp.MEAN, per_example_losses, axis=0)
return mean_loss
```
A few other things to note in the code above:
1. We used `tf.nn.softmax_cross_entropy_with_logits` to compute the loss. And then we scaled the total loss by the global batch size. This is important because all the replicas are training in sync and number of examples in each step of training is the global batch. So the loss needs to be divided by the global batch size and not by the replica (local) batch size.
2. We used the `tf.distribute.Strategy.reduce` API to aggregate the results returned by `tf.distribute.Strategy.experimental_run_v2`. `tf.distribute.Strategy.experimental_run_v2` returns results from each local replica in the strategy, and there are multiple ways to consume this result. You can `reduce` them to get an aggregated value. You can also do `tf.distribute.Strategy.experimental_local_results` to get the list of values contained in the result, one per local replica.
3. When `apply_gradients` is called within a distribution strategy scope, its behavior is modified. Specifically, before applying gradients on each parallel instance during synchronous training, it performs a sum-over-all-replicas of the gradients.
Finally, once we have defined the training step, we can iterate over `dist_dataset` and run the training in a loop:
```
with mirrored_strategy.scope():
for inputs in dist_dataset:
print(train_step(inputs))
```
In the example above, we iterated over the `dist_dataset` to provide input to your training. We also provide the `tf.distribute.Strategy.make_experimental_numpy_dataset` to support numpy inputs. You can use this API to create a dataset before calling `tf.distribute.Strategy.experimental_distribute_dataset`.
Another way of iterating over your data is to explicitly use iterators. You may want to do this when you want to run for a given number of steps as opposed to iterating over the entire dataset.
The above iteration would now be modified to first create an iterator and then explicitly call `next` on it to get the input data.
```
with mirrored_strategy.scope():
iterator = iter(dist_dataset)
for _ in range(10):
print(train_step(next(iterator)))
```
This covers the simplest case of using `tf.distribute.Strategy` API to distribute custom training loops. We are in the process of improving these APIs. Since this use case requires more work to adapt your code, we will be publishing a separate detailed guide in the future.
### What's supported now?
In TF 2.0 release, training with custom training loops is supported using `MirroredStrategy` as shown above and `TPUStrategy`.
`MultiWorkerMirorredStrategy` support will be coming in the future.
| Training API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy | OneDeviceStrategy |
|:----------------------- |:------------------- |:------------------- |:----------------------------- |:------------------------ |:------------------------- | :-------------------------- |
| Custom Training Loop | Experimental support | Experimental support | Support planned post 2.0 | Support planned post 2.0 | No support yet | Supported |
### Examples and Tutorials
Here are some examples for using distribution strategy with custom training loops:
1. [Tutorial](../tutorials/distribute/custom_training.ipynb) to train MNIST using `MirroredStrategy`.
2. [DenseNet](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/densenet/distributed_train.py) example using `MirroredStrategy`.
1. [BERT](https://github.com/tensorflow/models/blob/master/official/nlp/bert/run_classifier.py) example trained using `MirroredStrategy` and `TPUStrategy`.
This example is particularly helpful for understanding how to load from a checkpoint and generate periodic checkpoints during distributed training etc.
2. [NCF](https://github.com/tensorflow/models/blob/master/official/recommendation/ncf_keras_main.py) example trained using `MirroredStrategy` and `TPUStrategy` that can be enabled using the `keras_use_ctl` flag.
3. [NMT](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/nmt_with_attention/distributed_train.py) example trained using `MirroredStrategy`.
## Using `tf.distribute.Strategy` with Estimator (Limited support)
`tf.estimator` is a distributed training TensorFlow API that originally supported the async parameter server approach. Like with Keras, we've integrated `tf.distribute.Strategy` into `tf.Estimator`. If you're using Estimator for your training, you can easily change to distributed training with very few changes to your code. With this, Estimator users can now do synchronous distributed training on multiple GPUs and multiple workers, as well as use TPUs. This support in Estimator is, however, limited. See [What's supported now](#estimator_support) section below for more details.
The usage of `tf.distribute.Strategy` with Estimator is slightly different than the Keras case. Instead of using `strategy.scope`, now we pass the strategy object into the [`RunConfig`](https://www.tensorflow.org/api_docs/python/tf/estimator/RunConfig) for the Estimator.
Here is a snippet of code that shows this with a premade Estimator `LinearRegressor` and `MirroredStrategy`:
```
mirrored_strategy = tf.distribute.MirroredStrategy()
config = tf.estimator.RunConfig(
train_distribute=mirrored_strategy, eval_distribute=mirrored_strategy)
regressor = tf.estimator.LinearRegressor(
feature_columns=[tf.feature_column.numeric_column('feats')],
optimizer='SGD',
config=config)
```
We use a premade Estimator here, but the same code works with a custom Estimator as well. `train_distribute` determines how training will be distributed, and `eval_distribute` determines how evaluation will be distributed. This is another difference from Keras where we use the same strategy for both training and eval.
Now we can train and evaluate this Estimator with an input function:
```
def input_fn():
dataset = tf.data.Dataset.from_tensors(({"feats":[1.]}, [1.]))
return dataset.repeat(1000).batch(10)
regressor.train(input_fn=input_fn, steps=10)
regressor.evaluate(input_fn=input_fn, steps=10)
```
Another difference to highlight here between Estimator and Keras is the input handling. In Keras, we mentioned that each batch of the dataset is split automatically across the multiple replicas. In Estimator, however, we do not do automatic splitting of batch, nor automatically shard the data across different workers. You have full control over how you want your data to be distributed across workers and devices, and you must provide an `input_fn` to specify how to distribute your data.
Your `input_fn` is called once per worker, thus giving one dataset per worker. Then one batch from that dataset is fed to one replica on that worker, thereby consuming N batches for N replicas on 1 worker. In other words, the dataset returned by the `input_fn` should provide batches of size `PER_REPLICA_BATCH_SIZE`. And the global batch size for a step can be obtained as `PER_REPLICA_BATCH_SIZE * strategy.num_replicas_in_sync`.
When doing multi worker training, you should either split your data across the workers, or shuffle with a random seed on each. You can see an example of how to do this in the [Multi-worker Training with Estimator](../tutorials/distribute/multi_worker_with_estimator.ipynb).
We showed an example of using `MirroredStrategy` with Estimator. You can also use `TPUStrategy` with Estimator as well, in the exact same way:
```
config = tf.estimator.RunConfig(
train_distribute=tpu_strategy, eval_distribute=tpu_strategy)
```
And similarly, you can use multi worker and parameter server strategies as well. The code remains the same, but you need to use `tf.estimator.train_and_evaluate`, and set `TF_CONFIG` environment variables for each binary running in your cluster.
<a name="estimator_support"></a>
### What's supported now?
In TF 2.0 release, there is limited support for training with Estimator using all strategies except `TPUStrategy`. Basic training and evaluation should work, but a number of advanced features such as scaffold do not yet work. There may also be a number of bugs in this integration. At this time, we do not plan to actively improve this support, and instead are focused on Keras and custom training loop support. If at all possible, you should prefer to use `tf.distribute` with those APIs instead.
| Training API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy | OneDeviceStrategy |
|:--------------- |:------------------ |:------------- |:----------------------------- |:------------------------ |:------------------------- | :-------------------------- |
| Estimator API | Limited Support | Not supported | Limited Support | Limited Support | Limited Support | Limited Support |
### Examples and Tutorials
Here are some examples that show end to end usage of various strategies with Estimator:
1. [Multi-worker Training with Estimator](../tutorials/distribute/multi_worker_with_estimator.ipynb) to train MNIST with multiple workers using `MultiWorkerMirroredStrategy`.
2. [End to end example](https://github.com/tensorflow/ecosystem/tree/master/distribution_strategy) for multi worker training in tensorflow/ecosystem using Kubernetes templates. This example starts with a Keras model and converts it to an Estimator using the `tf.keras.estimator.model_to_estimator` API.
3. Official [ResNet50](https://github.com/tensorflow/models/blob/master/official/vision/image_classification/resnet_imagenet_main.py) model, which can be trained using either `MirroredStrategy` or `MultiWorkerMirroredStrategy`.
## Other topics
In this section, we will cover some topics that are relevant to multiple use cases.
<a name="TF_CONFIG"></a>
### Setting up TF\_CONFIG environment variable
For multi-worker training, as mentioned before, you need to set `TF_CONFIG` environment variable for each
binary running in your cluster. The `TF_CONFIG` environment variable is a JSON string which specifies what
tasks constitute a cluster, their addresses and each task's role in the cluster. We provide a Kubernetes template in the
[tensorflow/ecosystem](https://github.com/tensorflow/ecosystem) repo which sets
`TF_CONFIG` for your training tasks.
One example of `TF_CONFIG` is:
```
os.environ["TF_CONFIG"] = json.dumps({
"cluster": {
"worker": ["host1:port", "host2:port", "host3:port"],
"ps": ["host4:port", "host5:port"]
},
"task": {"type": "worker", "index": 1}
})
```
This `TF_CONFIG` specifies that there are three workers and two ps tasks in the
cluster along with their hosts and ports. The "task" part specifies that the
role of the current task in the cluster, worker 1 (the second worker). Valid roles in a cluster is
"chief", "worker", "ps" and "evaluator". There should be no "ps" job except when using `tf.distribute.experimental.ParameterServerStrategy`.
## What's next?
`tf.distribute.Strategy` is actively under development. We welcome you to try it out and provide and your feedback using [GitHub issues](https://github.com/tensorflow/tensorflow/issues/new).
| github_jupyter |
# SUMMER ANALYTICS 2021
# Week-1 Assignment
# **Gotta catch 'em all !**

**Welcome to your first assignment of Summer Analytics 2021! We hope you are excited to implement and test everything you have learnt up until now. The dataset which you'll use includes information about Pokemons.**
**We've got an interesting set of questions for you to get a basic understanding of pandas and data visualization libraries. GOOD LUCK!**
***Let's get started with importing numpy, pandas, seaborn and matplotlib!***
Note - matplotlib should be imported with the command :
`import matplotlib.pyplot as plt`
### 1) Start by importing all important libraries
For eg, "import numpy as np"
```
#your code here
import pandas as pd
import numpy as np
```
### 2) Read the csv file and assign it to a variable .
```
#your code here
data = pd.read_csv('SA2021_W1_Pokemon.csv')
data
data.describe()
data['Name']
```
### 3) Display shape of dataframe
Expected Output - (800, 13)
```
#your code here
data.shape
```
### 4) Print all columns of dataframe
Return an array containing names of all the columns.
```
#your code here
data.columns.values
```
### 5) Remove the column '#' and update the dataframe.
```
#your code here
data = data.drop(['#'], axis=1)
data.columns.values
```
### 6) Set the 'Name' column as the index of dataframe
```
#your code here
data.set_index(['Name'])
```
### 7) Print a list of all the unique Type-1 powers
```
#your code here
data['Type 1'].unique()
```
### 8) Create a column which contains the Type 1 and Type 2 abilities of pokemons, seperated with a '+'' sign. Also, display the no. of pokemons that have type-1 power as 'Psychic' and type 2 power as 'Flying' using this new column.
```
#your code here
data['combined'] = data[['Type 1', 'Type 2']].fillna('').agg(' + '.join, axis=1)
data.head()
data[data['Type 1'].str.contains('Psychic') & data['Type 2'].str.contains('Flying')]
```
## GRADED Questions (To be answered in the quiz)
### Try to retrieve some information from the data and answer the questions below . BEST OF LUCK !!
```
s = pd.Series(np.random.randn(4))
print(s.ndim)
import matplotlib.pyplot as plt
plt.plot([1,2, 3],[4,5, 1])
df = pd.DataFrame(['ff', 'gg', 'hh', 'yy'], [24, 12, 48, 30], columns = ['Name', 'Age'])
pd.date_range("2021-01-01", periods=3, freq="H")
array_1 = np.array([1, 2])
array_2 = np.array([4, 6])
array_3 = np.array(np.meshgrid(array_1, array_2)).T.reshape(-1,2)
print(array_3)
```
### 1. How many pokemons have 'Mega' in their name?
```
data['Name']
#your code here
Mega = data[data['Name'].str.contains('Mega')]
Mega.head()
len(Mega)
```
### 2. What is the standard deviation of Sp. Def. in the dataset ?
```
#your code here
data['Sp. Def'].std()
```
### 3. What percentage (upto 3 decimal places) of pokemons are legendary ?
```
#your code here
data.Legendary.value_counts(normalize=True)
```
### 4. Name the pokemon(s) with Maximum Defense.
```
#your code here
data.sort_values(by ='Defense', ascending=False).head()
```
### 5. Which poison pokemon has the strongest attack ?
```
#your code here
data[(data['Type 1'] == 'Poison') | (data['Type 2'] == 'Poison')].sort_values(by ='Attack', ascending=False).head()
```
### 6. Using seaborn make different types of plots, observe the trend and answer the questions given in the form.
```
#your code here
#your code here
```
### 7. Which is the second fastest non-legendary 'Ghost' type pokemon from 4th generation ?
```
#your code here
data[((data['Type 1'] == 'Ghost') | (data['Type 2'] == 'Ghost')) & (data['Legendary'] == False) & (data['Generation'] == 4)].sort_values(by = 'Speed').head()
```
### 8. How many non-legendary pokemons have stronger defence but weaker attack than Charizard?
```
data[data['Name'] == 'Charizard']
#your code here
len(data[(data['Legendary'] == False) & (data['Defense'] > 78) & (data['Attack'] < 84)])
```
## *Congratulations on coming this far! Since we were having so much fun playing with this dataset, here are some bonus questions that you can try to further deepen your understanding of the topic!*
### **Note:** These questions are UNGRADED, and are given as an extra exercise.
### Which pokemon has the highest combined value of Attack and Sp. Atk ?
```
#your code here
```
### Which type of legendary pokemons are the most common?
```
#your code here
```
### How many generation-3, non-legendary pokemons have higher HP than the weakest generation-6, legendary pokemon?
```
#your code here
```
### Print out the third slowest pokemon(s) in the dataset.
```
#your code here
```
### Which pokemon type has the highest average HP?
```
#your code here
```
| github_jupyter |
---
# Predicting Credit Card Default
### Preliminary Analysis and Preparation of Data
---
# Preliminary Analysis
## Loading data
Let's load our data and take a peak at the first 10 data objects and their corresponding features
```
import pandas as pd
import numpy as np
import imblearn #libary for imbalanced functions i.e. K-means SMOTE
from sklearn import preprocessing
#from google.colab import drive
#drive.mount('/content/drive')
# filename = "drive/Shareddrives/DS-project/default.xls"
filename = "default.xls"
data = pd.read_excel(filename)
data.head(10)
```
Based on the image above, there are a couple of issues that need to be addressed:
1. Our dataset feature names appear as the first object in our dataset
2. The first feature is an unique ID corresponding to a data object. We can assume that this feature will is irrelevant to the task at hand and can thus drop it in our feature selection stage.
We will address the first issue now:
```
feature_names = data.iloc[0, :] #acquire the names of the features, located on object with index = 0
feature_names = feature_names.values.tolist() #convert pandas frame to list
data.columns = feature_names #rename columns
data.rename(columns={'default payment next month': 'DEFAULT'}, inplace=True) #rename target variable name
data = data.drop([0]) #drop the first object
data.head(10) #inspect data
```
## Exploratory Data Analysis
We will start our analysis by checking the data types of each column as well as if any samples contain mising values.
By checking `data.dtypes`, we notice that all column's data types were imported as objects. In order to analyze their statistics (mean, std, etc..) we will need to convert the data types as follows:
1. The `'AGE'` attribute needs to be converted to an integer
2. The `'SEX'`, `'MARRIAGE'`, `'EDUCATION'`, `'DEFAULT'`, `'PAY_0'`, `'PAY_2'`,`'PAY_3'`, `'PAY_4'`, `'PAY_5'`, `'PAY_6'` will be converted to categorical values
3. The remaining columns will be converted to float values
Note that there are no missing values in our data
```
data = data.astype(str).astype(float) #coverting to float
int_cols=['AGE']
data[int_cols] = data[int_cols].astype('int')
categ_cols = ['SEX', 'MARRIAGE', 'EDUCATION', 'DEFAULT', 'PAY_0', 'PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6']
data[categ_cols] = data[categ_cols].astype('category')
data.info()
```
### Distribution of our target variable
Let's investigate the distribution of our target variable.
The Default variable is a binary number $\{0, 1\}$
```
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
#function to display barplot of a given categorical feature and print it's value distribution
def plot_barplot(feature):
count_values = feature.value_counts()
sns.barplot(count_values.index, count_values.values)
plt.title("Class Distribution")
plt.show()
print("Value Distribution:")
print(count_values)
index,counts = np.unique(data['DEFAULT'], return_counts=True) #
sns.set(rc={'figure.figsize':(4.0,4.0)})
sns.barplot(index, counts)
plt.title("Class Distribution")
plt.show()
print("Value Distribution:")
print(counts)
data[categ_cols].describe()
```
There are seven unique cateogries for the education feature, but only 4 of them are **known** `(1,2,3,4)`.
Therefore, we will bin together the values of `0, 5, 6` under one category called **unknown**.
```
plot_barplot(data['EDUCATION'])
```
### Numerical Statistics
```
data.describe()
```
### Feature Importance
Feature importance gives you a score for each feature of your data, the higher the score more important or relevant is the feature towards your output variable.
Feature importance is an inbuilt class that comes with Tree Based Classifiers we will be using Extra Tree Classifier for extracting the ranking the features in the dataset.
```
from sklearn.ensemble import ExtraTreesClassifier
import matplotlib.pyplot as plt
features = (data.iloc[:, 1:-1]).reset_index(drop=True)
features = features.astype('int64')
y = data.iloc[:,-1].astype('int8')
#use inbuilt class feature_importances of tree based classifiers
model = ExtraTreesClassifier()
model.fit(features,y)
feature_importances = model.feature_importances_
#print(feature_importances)
#plot graph of feature importances for better visualization
feat_importances = pd.Series(feature_importances, index=features.columns)
plt.figure(figsize=(6,6))
feat_importances.nlargest(22).plot(kind='barh')
plt.show()
```
### Correlation Matrix with Heatmap
Correlation states how the features are related to each other or the target variable.
**Positive Correlation** - Increase in one value of feature increases the value of the target variable.
**Negative Correlation** - increase in one value of feature decreases the value of the target variable.
Heatmap helps to identify which features are most related to the target variable.
```
### Correlation Matrix with Heatmap
features = data[features.columns]
features= features.astype('int64')
corrmat = features.corr()
#plot heat map
plt.figure(figsize=(22,22))
g=sns.heatmap(corrmat,
#norm=divnorm,
annot=True,
cmap="RdYlGn",
#center=0,
fmt ='.2f')
```
### Bivariate Analysis
```
sns.displot(data = data, x = 'LIMIT_BAL', kind="kde", label='Default Type', hue='DEFAULT', common_norm=False)
plt.title("Density Plot of Limit Balance per Default Type\n")
sns.displot(data = data, x = 'AGE', kind="kde", label='Default Type', hue='DEFAULT', common_norm=False)
plt.title("Density Plot of Age per Default Type\n")
data_tmp = data
data_tmp['PAY_0'] = data['PAY_0'].astype('int')
sns.displot(data = data_tmp, x = 'PAY_0', kind="kde", label='Default Type', hue='DEFAULT', common_norm=False)
plt.title("Density Plot of PAY_0 per Default Type\n")
sns.displot(data = data_tmp, x = 'BILL_AMT1', kind="kde", label='Default Type', hue='DEFAULT', common_norm=False)
plt.title("Density Plot of BIL_AMT1 per Default Type\n")
```
# Data Preparation
## Standarizing and Feature Selection
```
#drop first feature
data = data.drop(['ID'], axis='columns')
data.head(10)
```
## Binning unkwown values for Education feature
```
education_feature = data['EDUCATION']
data['EDUCATION'] = education_feature.replace(to_replace=[0.0, 5.0, 6.0], value=5.0)
#data2 = data.copy()
#data2['EDUCATION'] = data2['EDUCATION'].astype("int16")
sns.histplot(data = data2, x='EDUCATION', label='Default Type', hue='DEFAULT', stat='count', multiple='stack', palette='pastel')
plt.title("Stacked Histogram of Education per Default Type\n")
data.isnull().sum(axis=0)
```
**Writing the data to a comma-separated values (csv) file.**
```
data.to_csv('default_processed.csv', index=False)
```
| github_jupyter |
# Inheritance Exercise Clothing
The following code contains a Clothing parent class and two children classes: Shirt and Pants.
Your job is to code a class called Blouse. Read through the code and fill out the TODOs. Then check your work with the unit tests at the bottom of the code.
```
class Clothing:
def __init__(self, color, size, style, price):
self.color = color
self.size = size
self.style = style
self.price = price
def change_price(self, price):
self.price = price
def calculate_discount(self, discount):
return self.price * (1 - discount)
def calculate_shipping(self, weight, rate):
return weight*rate
class Shirt(Clothing):
def __init__(self, color, size, style, price, long_or_short):
Clothing.__init__(self, color, size, style, price)
self.long_or_short = long_or_short
def double_price(self):
self.price = 2*self.price
class Pants(Clothing):
def __init__(self, color, size, style, price, waist):
Clothing.__init__(self, color, size, style, price)
self.waist = waist
def calculate_discount(self, discount):
return self.price * (1 - discount / 2)
# TODO: Write a class called Blouse, that inherits from the Clothing class
# and has the the following attributes and methods:
# attributes: color, size, style, price, country_of_origin
# where country_of_origin is a string that holds the name of a
# country
#
# methods: triple_price, which has no inputs and returns three times
# the price of the blouse
#
#
class Blouse(Clothing):
def __init__(self, color, size, style, price, country_of_origin):
Clothing.__init__(self, color, size, style, price)
self.country_of_origin = country_of_origin
def triple_price(self):
#self.price = 3* self.price
return 3* self.price
# TODO: Add a method to the clothing class called calculate_shipping.
# The method has two inputs: weight and rate. Weight is a float
# representing the weight of the article of clothing. Rate is a float
# representing the shipping weight. The method returns weight * rate
# Unit tests to check your solution
import unittest
class TestClothingClass(unittest.TestCase):
def setUp(self):
self.clothing = Clothing('orange', 'M', 'stripes', 35)
self.blouse = Blouse('blue', 'M', 'luxury', 40, 'Brazil')
self.pants = Pants('black', 32, 'baggy', 60, 30)
def test_initialization(self):
self.assertEqual(self.clothing.color, 'orange', 'color should be orange')
self.assertEqual(self.clothing.price, 35, 'incorrect price')
self.assertEqual(self.blouse.color, 'blue', 'color should be blue')
self.assertEqual(self.blouse.size, 'M', 'incorrect size')
self.assertEqual(self.blouse.style, 'luxury', 'incorrect style')
self.assertEqual(self.blouse.price, 40, 'incorrect price')
self.assertEqual(self.blouse.country_of_origin, 'Brazil', 'incorrect country of origin')
def test_calculateshipping(self):
self.assertEqual(self.clothing.calculate_shipping(.5, 3), .5 * 3,\
'Clothing shipping calculation not as expected')
self.assertEqual(self.blouse.calculate_shipping(.5, 3), .5 * 3,\
'Clothing shipping calculation not as expected')
tests = TestClothingClass()
tests_loaded = unittest.TestLoader().loadTestsFromModule(tests)
unittest.TextTestRunner().run(tests_loaded)
shirt_one = Shirt('orange', 'M', 'stripes', 35, 'long')
shirt_one
shirt_one.calculate_discount(0.1)
blouse_one = Blouse('orange', 'M', 'stripes', 35, 'China')
blouse_one.price
blouse_one.triple_price()
blouse_one.price
```
| github_jupyter |
<br>
# Einblick in die Arbeit mit <i>zufall</i>
von Holger Böttcher - hbomat@posteo.de
<br><br>
Diese Arbeit steht unter der freien Lizenz [CC BY-SA 4.0](https://creativecommons.org/licenses/by-sa/4.0/deed.de)
<br><br>
### Abitur Bayern 2019
### Stochastik Teil B, Aufgabengruppe 1
<br>
Quelle: [serlo.org](https://de.serlo.org/mathe/deutschland/bayern/gymnasium/abiturpr%C3%BCfungen-l%C3%B6sung/mathematik-abitur-bayern-2019/stochastik,-teil-b,-aufgabengruppe-1)
<br>
<b>1.</b> Ein Unternehmen organisiert Fahrten mit einem Ausflugsschiff, das Platz für 60 <br>
Personen bietet.
Betrachtet wird eine Fahrt, bei der das Schiff voll besetzt ist. Unter den Fahrgästen
<br>befinden sich Erwachsene, Jugendliche und Kinder. Die Hälfte der Fahrgäste isst <br>
während der Fahrt ein Eis, von den Erwachsenen nur jeder Dritte, von den <br>
Jugendlichen und Kindern 75 %. Berechnen Sie, wie viele Erwachsene an der <br>
Fahrt teilnehmen.
<b>2.</b> Möchte man an einer Fahrt mit einem Ausflugsschiff, das Platz für 60 Fahrgäste<br>
bietet, teilnehmen, so muss man dafür im Voraus eine Reservierung vornehmen, ohne
<br>dabei schon den Fahrpreis bezahlen zu müssen. Erfahrungsgemäß erscheinen von<br>
den Personen mit Reservierung einige nicht zur Fahrt. Für die 60 zur Verfügung <br>
stehenden Plätze lässt das Unternehmen deshalb bis zu 64 Reservierungen zu. Es soll <br>
davon ausgegangen werden, dass für jede Fahrt tatsächlich 64 Reservierungen<br>
vorgenommen werden. Erscheinen mehr als 60 Personen mit Reservierung zur Fahrt, <br>
so können nur 60 von ihnen daran teilnehmen; die übrigen müssen abgewiesen <br>
werden.
Die Zufallsgröße $X$ beschreibt die Anzahl der Personen mit Reservierung, die nicht zur <br>
Fahrt erscheinen. Vereinfachend soll angenommen werden, dass $X$ binomialverteilt ist, <br>
wobei die Wahrscheinlichkeit dafür, dass eine zufällig ausgewählte Person mit <br>
Reservierung nicht zur Fahrt erscheint, 10% beträgt.
<b>a)</b> Geben Sie einen Grund dafür an, dass es sich bei der Annahme, die <br>
Zufallsgröße $X$ ist binomialverteilt, im Sachzusammenhang um eine Vereinfachung <br>handelt.
<b>b)</b> Bestimmen Sie die Wahrscheinlichkeit dafür, dass keine Person mit <br>
Reservierung abgewiesen werden muss.
<b>c)</b> Für das Unternehmen wäre es hilfreich, wenn die Wahrscheinlichkeit dafür, <br>
mindestens eine Person mit Reservierung abweisen zu müssen, höchstens ein <br>
Prozent wäre. Dazu müsste die Wahrscheinlichkeit dafür, dass eine zufällig <br>
ausgewählte Person mit Reservierung nicht zur Fahrt erscheint, mindestens einen <br>bestimmenten Wert haben. Ermitteln Sie diesen Wert auf ganze Prozent genau.
Das Unternhmen richtet ein Online-Portal zur Reservierung ein und vermutet, dass <br>
dadurch der Anteil der Personen mit Reservierung, die zur jeweiligen Fahrt nicht <br>
erscheinen, zunehmen könnte. Als Grundlage für die Entscheidung darüber, ob pro <br>
Fahrt künftig mehr als 64 Reservierungen zugelassen werden, soll die Nullhypothese <br>
"Die Wahrscheinlichkeit dafür, dass eine zufällig ausgewählte Person mit <br>
Reservierung nicht zur Fahrt erscheint, beträgt höchstens 10%." mithilfe einer <br>
Stichprobe von 200 Personen mit Reservierung auf einem Signifikanzniveau von 5% <br>
getestet werden. Vor der Durchführung des Tests wird festgelegt, die Anzahl der für <br>
eine Fahrt möglichen Reservierungen nur dann zu erhöhen, wenn die Nullhypothese <br>aufgrund des Testergebnisses abgelehnt werden müsste.
<b>d)</b> Ermitteln Sie die zughörige Entscheidungsregel.
<b>e)</b> Entscheiden Sie, ob bei der Wahl der Nullhypothese eher das Interesse, dass <br>
weniger Platz frei bleiben sollen, oder das Interesse, dass nicht mehr Personen mit <br>
Reservierung abgewiesen werden müssen, im Vordergrund stand. Begründen Sie <br>
Ihre Entscheidung.
<b>f)</b> Beschreiben Sie den zugehörigen Fehler zweiter Art sowie die daraus resultierende <br>Konsequenz im Sachzusammenhang.
<br><br>
```
%run zufall/start
```
<br>
### Zu 1.
$e$ - Erwachsene, $k$ - Kinder + Jugendliche$\quad E$ ist ungünstig, da mit $2.71828...$ belegt
Gleichungen
$\quad$$e+k = 60\quad\quad\,$ */ alle Personen zusammen*
<br><br>
$\quad$$\dfrac{e}{3} + \dfrac{3}{4} k = 30\quad$ */ eisessende Personen*
<br>
Die rechten Seiten werden zu Null gemacht und dann die linken Seiten als <br>
lineares Gleichungssystem gelöst
<br>
```
löse([e + k - 60, 1/3*e + 3/4*k - 30])
```
<br>
### Zu 2.
### a)
Bei einer zufälligen Reisegruppe kann nicht davon ausgegangen werden, daß <br>die Trefferwahrscheinlichkeit für alle Personen gleich ist<br><br>
Außerdem kann nicht vorausgesetzt werden, dass die Ereignisse unabhängig <br>
voneinander sind, da z.B. Ehepaare die Reise normalerweise zusammenen <br>
antreten oder absagen
<br>
### b)
Die zugrundeliegennde Binomialverteilung ist
```
bv = BV(64, 0.1) # BV - BinomialVerteilung
```
Mindestens 4 der Personen mit Reservierung dürfen nicht erscheinen
```
bv.P(X >= 4) # der exakte Wert
bv.P(X >= 4, p=ja) # der (gerundete) Prozentwert
```
### c)
$X$ soll eine Bernoullikette mit der Länge 64 und der unbekannten <br>
Trefferwahrscheinlichkeit $p$ sein
Das Ereignis "Mindestens eine Person mit Reservierung wird abgewiesen" tritt dann <br>
ein, wenn "Höchstens 3 Personen mit Reservierung erscheinen nicht" eintritt
$p$ muss also so gewählt werden, dass $P(X \le 3) \le 0.01$ ist
Für $p=0.1 = 10 \%$ und $p=0.2=20\%$ ergeben sich die Werte
```
p1 = 0.1
p2 = 0.2
BV(64, p1).P(X <= 3, p=ja) # in Prozent
BV(64, p2).P(X <= 3, p=ja) # ebenso
```
Das zeigt, dass der gesuchte Wert im Intervall $\;(0.1, \,0.2)\,$ liegt
Man könnte jetzt durch schrittweise (manuelle) Verfeinerung dieses Intervalls <br>
den Bereich so lange einengen, bis ein Wert mit der geforderten Genauigkeit <br>
gefunden ist (das geht recht schnell)
Hier wird eine Wertetabelle für den intereressierenden Bereich in ausreichend <br>
feiner Untergliederung erzeugt (sie entspricht der in der Aufgabenstellung <br>
angegebenen Tabelle)
```
ber = [0.1 + i*0.01 for i in range(11)] # p-Werte
ber
```
Die Ungenauigkeiten sind duch die Gleitkommarechnung verursacht (und nicht <br>
zu vermeiden), schöner ist
```
ber = [N(0.1 + i*0.01, 3) for i in range(11)]
bv = [(p, BV(64, p).P(X <= 3, p=ja)) for p in ber]
bv
```
Es kann abgelesen werden, dass der Spung unter die $1\%$ - Grenze bei $15\%$ liegt
### d)
Es ist ein statistischer Test für die unbekannte Wahrscheinlichkeit einer <br>Binomialverteilung durchzuführen
(Einzelheiten für die Argumente siehe unten)
```
test = STP(0.1, 0.05, 'rechts', 200, 'B')
test
test.regel
test.schema
test.K
test.an_ber # Annahmebereich
ab = list(test.ab_ber) # Mit etwas Tricksen kommt man zu einer
# passablen Ausgabe des großen Ablehnungsbereiches
ab[:5] + [Symbol('...')] + ab[-5:]
```
### e)
Bei dieser Wahl der Nullhypothese soll die Wahrscheinlichkeit, den Fehler irrtümlich <br>
mehr als 64 Reservierungen zuzulassen gering, höchstens 5% , gehalten werden.
Damit steht das Interesse, dass nicht mehr Personen mit Reservierung abgewiesen <br>
werden müssen, im Vordergrund.
### f)
Der zum Test gehörende Fehler 2. Art liegt vor, wenn die Nullhypothese, d.h. die <br>
Wahrscheinlichkeit, dass eine Person die reserviert hat, aber nicht erscheint höchstens <br>
10% ist, falsch ist und dennoch angenommen wird. D.h. es bleibt bei 64 möglichen <br>
Reservierungen, obwohl der Anteil nicht erscheinender Personen zugenommen hat.
Im Sachzusammenhang bedeutet dies, dass der Unternehmer mit mehr nicht besetzten <br>
Plätzen zu rechnen hat und mit größerer Wahrscheinlichkeit einen finanziellen Verlust <br>hinnehmen muss.
| github_jupyter |
## How-to guide for Real-Time Forecasting use-case on Abacus.AI platform
This notebook provides you with a hands on environment to build a real-time forecasting model using the Abacus.AI Python Client Library.
We'll be using the [Household Electricity Usage Dataset](https://s3.amazonaws.com/realityengines.exampledatasets/rtforecasting/household_electricity_usage.csv), which contains data about electricity usage in a specified household.
1. Install the Abacus.AI library.
```
!pip install abacusai
```
We'll also import pandas and pprint tools for neat visualization in this notebook.
```
import pandas as pd # A tool we'll use to download and preview CSV files
import pprint # A tool to pretty print dictionary outputs
pp = pprint.PrettyPrinter(indent=2)
```
2. Add your Abacus.AI [API Key](https://abacus.ai/app/profile/apikey) generated using the API dashboard as follows:
```
#@title Abacus.AI API Key
api_key = '' #@param {type: "string"}
```
3. Import the Abacus.AI library and instantiate a client.
```
from abacusai import ApiClient
client = ApiClient(api_key)
```
## 1. Create a Project
Abacus.AI projects are containers that have datasets and trained models. By specifying a business **Use Case**, Abacus.AI tailors the deep learning algorithms to produce the best performing model possible for your data.
We'll call the `list_use_cases` method to retrieve a list of the available Use Cases currently available on the Abacus.AI platform.
```
client.list_use_cases()
```
In this notebook, we're going to create a real-time forecasting model using the Household Electricity Usage dataset. The 'ENERGY' use case is best tailored for this situation.
```
#@title Abacus.AI Use Case
use_case = 'ENERGY' #@param {type: "string"}
```
By calling the `describe_use_case_requirements` method we can view what datasets are required for this use_case.
```
for requirement in client.describe_use_case_requirements(use_case):
pp.pprint(requirement.to_dict())
```
Finally, let's create the project.
```
real_time_project = client.create_project(name='Electricity Usage Forecasting', use_case=use_case)
real_time_project.to_dict()
```
**Note: When feature_groups_enabled is True then the use case supports feature groups (collection of ML features). Feature groups are created at the organization level and can be tied to a project to further use it for training ML models**
## 2. Add Datasets to your Project
Abacus.AI can read datasets directly from `AWS S3` or `Google Cloud Storage` buckets, otherwise you can also directly upload and store your datasets with Abacus.AI. For this notebook, we will have Abacus.AI read the datasets directly from a public S3 bucket's location.
We are using one dataset for this notebook. We'll tell Abacus.AI how the dataset should be used when creating it by tagging the dataset with a special Abacus.AI **Dataset Type**.
- [Household Electricity Usage Dataset](https://s3.amazonaws.com/realityengines.exampledatasets/rtforecasting/household_electricity_usage.csv) (**TIMESERIES**):
This dataset contains information about electricity usage in specified households over a period of time.
### Add the dataset to Abacus.AI
First we'll use Pandas to preview the file, then add it to Abacus.AI.
```
pd.read_csv('https://s3.amazonaws.com/realityengines.exampledatasets/rtforecasting/household_electricity_usage.csv')
```
Using the Create Dataset API, we can tell Abacus.AI the public S3 URI of where to find the datasets. We will also give each dataset a Refresh Schedule, which tells Abacus.AI when it should refresh the dataset (take an updated/latest copy of the dataset).
If you're unfamiliar with Cron Syntax, Crontab Guru can help translate the syntax back into natural language: [https://crontab.guru/#0_12_\*_\*_\*](https://crontab.guru/#0_12_*_*_*)
**Note: This cron string will be evaluated in UTC time zone**
```
real_time_dataset = client.create_dataset_from_file_connector(name='Household Electricity Usage',table_name='Household_Electricity_Usage',
location='s3://realityengines.exampledatasets/rtforecasting/household_electricity_usage.csv',
refresh_schedule='0 12 * * *')
datasets = [real_time_dataset]
for dataset in datasets:
dataset.wait_for_inspection()
```
## 3. Create Feature Groups and add them to your Project
Datasets are created at the organization level and can be used to create feature groups as follows:
```
feature_group = client.create_feature_group(table_name='real_time_forecasting',sql='SELECT * FROM Household_Electricity_Usage')
```
Adding Feature Group to the project:
```
client.add_feature_group_to_project(feature_group_id=feature_group.feature_group_id,project_id = real_time_project.project_id)
```
Setting the Feature Group type according to the use case requirements:
```
client.set_feature_group_type(feature_group_id=feature_group.feature_group_id, project_id = real_time_project.project_id, feature_group_type= "TIMESERIES")
```
Check current Feature Group schema:
```
client.get_feature_group_schema(feature_group_id=feature_group.feature_group_id)
```
#### For each **Use Case**, there are special **Column Mappings** that must be applied to a column to fulfill use case requirements. We can find the list of available **Column Mappings** by calling the *Describe Use Case Requirements* API:
```
client.describe_use_case_requirements(use_case)[0].allowed_feature_mappings
client.set_feature_mapping(project_id = real_time_project.project_id,feature_group_id= feature_group.feature_group_id, feature_name='value',feature_mapping='TARGET')
client.set_feature_mapping(project_id = real_time_project.project_id,feature_group_id= feature_group.feature_group_id, feature_name='time',feature_mapping='DATE')
client.set_feature_mapping(project_id = real_time_project.project_id,feature_group_id= feature_group.feature_group_id, feature_name='id',feature_mapping='ITEM_ID')
```
For each required Feature Group Type within the use case, you must assign the Feature group to be used for training the model:
```
client.use_feature_group_for_training(project_id=real_time_project.project_id, feature_group_id=feature_group.feature_group_id)
```
Now that we've our feature groups assigned, we're almost ready to train a model!
To be sure that our project is ready to go, let's call project.validate to confirm that all the project requirements have been met:
```
real_time_project.validate()
```
## 4. Train a Model
For each **Use Case**, Abacus.AI has a bunch of options for training. We can call the *Get Training Config Options* API to see the available options.
```
real_time_project.get_training_config_options()
```
In this notebook, we'll just train with the default options, but definitely feel free to experiment, especially if you have familiarity with Machine Learning.
```
real_time_model = real_time_project.train_model(training_config={})
real_time_model.to_dict()
```
After we start training the model, we can call this blocking call that routinely checks the status of the model until it is trained and evaluated.
```
real_time_model.wait_for_evaluation()
```
**Note that model training might take some minutes to some hours depending upon the size of datasets, complexity of the models being trained and a variety of other factors**
## **Checkpoint** [Optional]
As model training can take an hours to complete, your page could time out or you might end up hitting the refresh button, this section helps you restore your progress:
```
!pip install abacusai
import pandas as pd
import pprint
pp = pprint.PrettyPrinter(indent=2)
api_key = '' #@param {type: "string"}
from abacusai import ApiClient
client = ApiClient(api_key)
real_time_project = next(project for project in client.list_projects() if project.name == 'Electricity Usage Forecasting')
real_time_model = real_time_project.list_models()[-1]
real_time_model.wait_for_evaluation()
```
## Evaluate your Model Metrics
After your model is done training you can inspect the model's quality by reviewing the model's metrics
```
pp.pprint(real_time_model.get_metrics().to_dict())
```
To get a better understanding on what these metrics mean, visit our [documentation](https://abacus.ai/app/help/useCases/ENERGY/training) page.
## 5. Deploy Model
After the model has been trained, we need to deploy the model to be able to start making predictions. Deploying a model will reserve cloud resources to host the model for Realtime and/or batch predictions.
```
real_time_deployment = client.create_deployment(name='Electricity Usage Deployment',description='Electricity Usage Deployment',model_id=real_time_model.model_id)
real_time_deployment.wait_for_deployment()
```
After the model is deployed, we need to create a deployment token for authenticating prediction requests. This token is only authorized to predict on deployments in this project, so it's safe to embed this token inside of a user-facing application or website.
```
deployment_token = real_time_project.create_deployment_token().deployment_token
deployment_token
```
## 6. Predict
Now that you have an active deployment and a deployment token to authenticate requests, you can make the `get_forecast` API call below.
This command will return a forecast under each percentile for the specified ITEM_ID. The forecast will be performed based on attributes specified in the dataset.
```
ApiClient().get_forecast(deployment_token=deployment_token,
deployment_id=real_time_deployment.deployment_id,
query_data={"id":"MT_001"})
```
| github_jupyter |
# Imports
```
import pandas as pd
from bs4 import BeautifulSoup
import string
import re
from datetime import datetime
```
# Import Data
```
process = pd.read_csv('https://github.com/ftmnl/asr/raw/main/data/allExport.csv', sep='|')
process
#remove noise
process = process.dropna()
process = process.rename(columns = {"file_name_sort": "title", "content": "abstract"})
remove = ['.DS_Store', 'NaN', 'Readme.md']
process = process[~process.title.isin(remove)]
```
# Transform Content
```
#abstract html to string
translate_table = dict((ord(char), None) for char in string.punctuation)
def prettify(text):
text = BeautifulSoup(text, 'html.parser').get_text()
text = text.replace("\r", "")
#text = text.replace('\n', '')
#text = text.translate(translate_table)
return str(text)
process.abstract = process.abstract.apply(prettify)
```
# Search formats
```
#extract id, type and date from title
import re
numberlist = []
typelist = []
datelist = []
for title in process.title:
id = re.search('^[0-9\.]+', title)
if id == None:
numberlist.append(None)
else:
numberlist.append(id.group(0))
type = re.search('(?<=[0-9\_]\_)[a-z A-Z]+(?=\_)', title)
if type != None:
typelist.append(type.group(0))
else:
typelist.append("Onbekend")
date = re.search("[0-9-?]+(?=.pdf$)", title)
if date == None:
datelist.append(None)
else:
try: datelist.append(datetime.strptime(date.group(0), '%d-%m-%Y'))
except:
try: datelist.append(datetime.strptime(date.group(0), '%-d-%-m-%Y'))
except: datelist.append(date.group(0))
process["id"] = numberlist
process["type"] = typelist
process["date"] = datelist
#Improve some Dating
betterDate = []
for index, row in process.iterrows():
regexDate = re.search("(?<=Date : )[0-9-]{3,5}-20[0-9]{2} [0-9:]{8}", row.abstract)
if regexDate != None:
date = datetime.strptime(regexDate.group(0), '%d-%m-%Y %H:%M:%S')
betterDate.append(date)
else:
betterDate.append(None)
process["betterDate"] = betterDate
```
# Clean Data
```
def cleanTitle(title):
title = re.sub('^[0-9\.]+_+[a-z A-Z]+_', '', title)
title = re.sub('[0-9\-]+.pdf$', '', title)
title = re.sub('.msg_', ' ', title)
return title
process.title = process.title.apply(cleanTitle)
```
# Export processing
```
process[['id','type','date','betterDate','title','abstract']].to_excel(r'..\data\preprocessed.xlsx')
```
| github_jupyter |
```
import os
# set the current working directory to the deployed package folder. This is required by isaac.
# This cell should only run once.
os.chdir("/home/davis/deploy/davis/simple_joint_control-pkg")
os.getcwd()
from IPython.display import display
import json
import numpy as np
import time
import threading
from engine.pyalice import Application, Codelet
from engine.pyalice.gui.composite_widget import CompositeWidget
np.set_printoptions(precision=3)
# A Python codelet for joint control through widget
class JointPositionControl(Codelet):
def start(self):
self.rx = self.isaac_proto_rx("CompositeProto", "state")
self.tx = self.isaac_proto_tx("CompositeProto", "command")
joints = self.config.joints
limits = self.config.limits
self._widget = CompositeWidget(joints, "position", limits)
if self._widget is None:
report_failure("Cannot create valid widget")
return
display(self._widget.panel)
self.tick_periodically(0.1)
def tick(self):
state_msg = self.rx.message
if state_msg is None:
return
print(state_msg)
self._widget.composite = state_msg
self.tx._msg = self._widget.composite
if self.tx._msg is not None:
self.tx.publish()
```
UR10 and Smarthand in Omniverse Isaac Sim
======
```
# set kinematic file and get list of joints
kinematic_file = "/home/davis/deploy/davis/rm_isaac_bridge-pkg/apps/assets/kinematic_trees/rm_ur10.kinematic.json"
joints = []
with open(kinematic_file,'r') as fd:
kt = json.load(fd)
for link in kt['links']:
if 'motor' in link and link['motor']['type'] != 'constant':
joints.append(link['name'])
print(joints)
app = Application(name="simple_joint_control_sim")
# load subgraphcs
app.load(filename="packages/planner/apps/multi_joint_lqr_control.subgraph.json", prefix="lqr")
app.load(filename="packages/navsim/apps/navsim_tcp.subgraph.json", prefix="simulation")
# edges
simulation_node = app.nodes["simulation.interface"]
lqr_interface = app.nodes["lqr.subgraph"]["interface"]
app.connect(simulation_node["output"], "joint_state", lqr_interface, "joint_state")
app.connect(lqr_interface, "joint_command", simulation_node["input"], "joint_position")
# configs
app.nodes["lqr.kinematic_tree"]["KinematicTree"].config.kinematic_file = kinematic_file
lqr_planner = app.nodes["lqr.local_plan"]["MultiJointLqrPlanner"]
lqr_planner.config.speed_min = [-50.0] * len(joints)
lqr_planner.config.speed_max = [50.0] * len(joints)
lqr_planner.config.acceleration_min = [-50.0] * len(joints)
lqr_planner.config.acceleration_max = [50.0] * len(joints)
# add pycodelet JointPositionControl
widget_node = app.add("command_generator")
joint_commander = widget_node.add(JointPositionControl)
joint_commander.config.joints = joints
joint_commander.config.limits = [[-7, 7]] * len(joints)
app.connect(joint_commander, "command", lqr_interface, "joint_target")
app.connect(simulation_node["output"], "joint_state", joint_commander, "state")
app.start()
# stop Isaac app
app.stop()
```
Kinova Jaco (gen2, 7 joints) Hardware
======
Install the KinoveJaco SDK in /opt/JACO2SDK (tested with v1.4.2) and connect to workstation via USB. Make sure the USB port has write permission
```
kinematic_file = "apps/assets/kinematic_trees/kinova_j2n7.kinematic.json"
joints = []
with open(kinematic_file,'r') as fd:
kt = json.load(fd)
for link in kt['links']:
if 'motor' in link and link['motor']['type'] != 'constant':
joints.append(link['name'])
print(joints)
app = Application(name="simple_joint_control_kinova_real")
# load lqr subgraphcs
app.load(filename="packages/planner/apps/multi_joint_lqr_control.subgraph.json", prefix="lqr")
lqr_interface = app.nodes["lqr.subgraph"]["interface"]
# add kinova driver codelet
app.load_module("kinova_jaco")
driver = app.add("driver").add(app.registry.isaac.kinova_jaco.KinovaJaco)
# edges
app.connect(driver, "arm_state", lqr_interface, "joint_state")
app.connect(lqr_interface, "joint_command", driver, "arm_command")
# configs
app.nodes["lqr.kinematic_tree"]["KinematicTree"].config.kinematic_file = kinematic_file
lqr_planner = app.nodes["lqr.local_plan"]["MultiJointLqrPlanner"]
lqr_planner.config.speed_min = [-0.5] * len(joints)
lqr_planner.config.speed_max = [0.5] * len(joints)
lqr_planner.config.acceleration_min = [-0.5] * len(joints)
lqr_planner.config.acceleration_max = [0.5] * len(joints)
driver.config.kinematic_tree = "lqr.kinematic_tree"
driver.config.kinova_jaco_sdk_path = "/opt/JACO2SDK/API/"
driver.config.tick_period = "50ms"
# add pycodelet JointPositionControl
widget_node = app.add("command_generator")
joint_commander = widget_node.add(JointPositionControl)
joint_commander.config.joints = joints
joint_commander.config.limits = [[-2*np.pi, 2*np.pi]] * len(joints)
app.connect(joint_commander, "command", lqr_interface, "joint_target")
app.connect(driver, "arm_state", joint_commander, "state")
app.start()
app.stop()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import torch
from tqdm.notebook import tqdm
from torch.utils.data import TensorDataset
from transformers import (ElectraForSequenceClassification, ElectraTokenizerFast)
from matplotlib import pyplot as plt
import seaborn as sns
import sklearn
from sklearn.metrics import classification_report, confusion_matrix
```
# Get data
```
df = pd.read_csv('./../../../labeledTweets/allLabeledTweets.csv')
df = df[['id', 'message', 'label']]
df = df.drop_duplicates()
print(df.shape[0])
df.head()
df['label'].value_counts()
newLine ="\\n|\\r"
urls = '(https?:\/\/(?:www\.|(?!www))[a-zA-Z0-9][a-zA-Z0-9-]+[a-zA-Z0-9]\.[^\s]{2,}|www\.[a-zA-Z0-9][a-zA-Z0-9-]+[a-zA-Z0-9]\.[^\s]{2,}|https?:\/\/(?:www\.|(?!www))[a-zA-Z0-9]+\.[^\s]{2,}|www\.[a-zA-Z0-9]+\.[^\s]{2,})'
numbers = '\d+((\.|\-)\d+)?'
mentions = '\B\@([\w\-]+)'
hashtag = '#'
whitespaces = '\s+'
leadTrailWhitespace = '^\s+|\s+?$'
df['clean_message'] = df['message']
df['clean_message'] = df['clean_message'].str.replace(newLine,' ',regex=True)
df['clean_message'] = df['clean_message'].str.replace(urls,' URL ',regex=True)
df['clean_message'] = df['clean_message'].str.replace(mentions,' MENTION ',regex=True)
df['clean_message'] = df['clean_message'].str.replace(numbers,' NMBR ',regex=True)
df['clean_message'] = df['clean_message'].str.replace(hashtag,' ',regex=True)
df['clean_message'] = df['clean_message'].str.replace(whitespaces,' ',regex=True)
df['clean_message'] = df['clean_message'].str.replace(leadTrailWhitespace,'',regex=True)
df.head()
```
# Train, validate split (balanced)
```
df_0 = df[df['label']==0]
df_1 = df[df['label']==1]
df_2 = df[df['label']==2]
trainLabelSize = round(df_1.shape[0]*0.85)
trainLabelSize
df_0 = df_0.sample(trainLabelSize, random_state=42)
df_1 = df_1.sample(trainLabelSize, random_state=42)
df_2 = df_2.sample(trainLabelSize, random_state=42)
df_train = pd.concat([df_0, df_1, df_2])
# Shuffle rows
df_train = sklearn.utils.shuffle(df_train, random_state=42)
df_train['label'].value_counts()
df_val = df.merge(df_train, on=['id', 'message', 'label', 'clean_message'], how='left', indicator=True)
df_val = df_val[df_val['_merge']=='left_only']
df_val = df_val[['id', 'message', 'label', 'clean_message']]
df_val['label'].value_counts()
```
# Tokenizer "google/electra-base-discriminator"
```
tokenizer = ElectraTokenizerFast.from_pretrained('google/electra-base-discriminator', do_lower_case=True)
```
### Find max length for tokenizer
```
token_lens = []
for txt in list(df.clean_message.values):
tokens = tokenizer.encode(txt, max_length=512, truncation=True)
token_lens.append(len(tokens))
max_length = max(token_lens)
max_length
```
### Encode messages
```
encoded_data_train = tokenizer.batch_encode_plus(
df_train["clean_message"].values.tolist(),
add_special_tokens=True,
return_attention_mask=True,
padding='max_length',
truncation=True,
max_length=max_length,
return_tensors='pt'
)
encoded_data_val = tokenizer.batch_encode_plus(
df_val["clean_message"].values.tolist(),
add_special_tokens=True,
return_attention_mask=True,
padding='max_length',
truncation=True,
max_length=max_length,
return_tensors='pt'
)
input_ids_train = encoded_data_train['input_ids']
attention_masks_train = encoded_data_train['attention_mask']
labels_train = torch.tensor(df_train.label.values)
input_ids_val = encoded_data_val['input_ids']
attention_masks_val = encoded_data_val['attention_mask']
labels_val = torch.tensor(df_val.label.values)
dataset_train = TensorDataset(input_ids_train, attention_masks_train, labels_train)
dataset_val = TensorDataset(input_ids_val, attention_masks_val, labels_val)
len(dataset_train), len(dataset_val)
```
# Model "google/electra-base-discriminator"
```
model = ElectraForSequenceClassification.from_pretrained("google/electra-base-discriminator",
num_labels=3,
output_attentions=False,
output_hidden_states=False)
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
batch_size = 32
dataloader_train = DataLoader(dataset_train, sampler=RandomSampler(dataset_train), batch_size=batch_size)
dataloader_validation = DataLoader(dataset_val, sampler=SequentialSampler(dataset_val), batch_size=batch_size)
from transformers import get_linear_schedule_with_warmup
optimizer = torch.optim.AdamW(model.parameters(), lr=1e-5, eps=1e-8)
# optimizer = torch.optim.SGD(model.parameters(), lr=0.0001)
epochs = 5
scheduler = get_linear_schedule_with_warmup(optimizer, num_warmup_steps=0, num_training_steps=len(dataloader_train)*epochs)
# Function to measure weighted F1
from sklearn.metrics import f1_score
def f1_score_func(preds, labels):
preds_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return f1_score(labels_flat, preds_flat, average='weighted')
import random
seed_val = 17
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
device = torch.device('cpu')
model.to(device)
print(device)
# Function to evaluate model. Returns average validation loss, predictions, true values
def evaluate(dataloader_val):
model.eval()
loss_val_total = 0
predictions, true_vals = [], []
progress_bar = tqdm(dataloader_val, desc='Validating:', leave=False, disable=False)
for batch in progress_bar:
batch = tuple(b.to(device) for b in batch)
inputs = {'input_ids': batch[0], 'attention_mask': batch[1], 'labels': batch[2]}
with torch.no_grad():
outputs = model(**inputs)
loss = outputs[0]
logits = outputs[1]
loss_val_total += loss.item()
logits = logits.detach().cpu().numpy()
label_ids = inputs['labels'].cpu().numpy()
predictions.append(logits)
true_vals.append(label_ids)
loss_val_avg = loss_val_total/len(dataloader_val)
predictions = np.concatenate(predictions, axis=0)
true_vals = np.concatenate(true_vals, axis=0)
return loss_val_avg, predictions, true_vals
```
# Evaluate untrained model
```
_, predictions, true_vals = evaluate(dataloader_validation)
from sklearn.metrics import classification_report, confusion_matrix
preds_flat = np.argmax(predictions, axis=1).flatten()
print(classification_report(true_vals, preds_flat))
print(f1_score_func(predictions, true_vals))
pd.DataFrame(confusion_matrix(true_vals, preds_flat),
index = [['actual', 'actual', 'actual'], ['neutral', 'positive', 'negative']],
columns = [['predicted', 'predicted', 'predicted'], ['neutral', 'positive', 'negative']])
```
# Train
```
for epoch in tqdm(range(1, epochs+1)):
model.train()
loss_train_total = 0
progress_bar = tqdm(dataloader_train, desc='Epoch {:1d}'.format(epoch), leave=False, disable=False)
for batch in progress_bar:
model.zero_grad()
batch = tuple(b.to(device) for b in batch)
inputs = {'input_ids': batch[0], 'attention_mask': batch[1], 'labels': batch[2]}
outputs = model(**inputs)
loss = outputs[0]
loss_train_total += loss.item()
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
optimizer.step()
scheduler.step()
progress_bar.set_postfix({'training_loss': '{:.3f}'.format(loss.item()/len(batch))})
torch.save(model.state_dict(), f'modelsCleaned/finetuned_ELECTRAbase_epoch_{epoch}.model')
tqdm.write(f'\nEpoch {epoch}')
loss_train_avg = loss_train_total/len(dataloader_train)
tqdm.write(f'Training loss: {loss_train_avg}')
val_loss, predictions, true_vals = evaluate(dataloader_validation)
val_f1 = f1_score_func(predictions, true_vals)
tqdm.write(f'Validation loss: {val_loss}')
tqdm.write(f'F1 Score (Weighted): {val_f1}')
preds_flat = np.argmax(predictions, axis=1).flatten()
print('Classification report:')
print(classification_report(true_vals, preds_flat))
print('Confusion matrix:')
print(pd.DataFrame(confusion_matrix(true_vals, preds_flat),
index = [['actual', 'actual', 'actual'], ['neutral', 'positive', 'negative']],
columns = [['predicted', 'predicted', 'predicted'], ['neutral', 'positive', 'negative']]))
```
# Evaluate best model
```
model.load_state_dict(torch.load('modelsBase/finetuned_ELECTRAbase_epoch_X.model', map_location=torch.device('cpu')))
_, predictions, true_vals = evaluate(dataloader_validation)
preds_flat = np.argmax(predictions, axis=1).flatten()
print(f1_score_func(predictions, true_vals))
print(classification_report(true_vals, preds_flat))
pd.DataFrame(confusion_matrix(true_vals, preds_flat),
index = [['actual', 'actual', 'actual'], ['neutral', 'positive', 'negative']],
columns = [['predicted', 'predicted', 'predicted'], ['neutral', 'positive', 'negative']])
```
| github_jupyter |
```
import os
import requests
import calendar
data = {'formQuery:menuAldId':1,
'formQuery:selectRad':'incidentLevel',
'dateFromCrime':'01/01/2005',
'dateToCrime':'07/10/2017',
'dateFromAcci':'MM/DD/YYYY',
'dateToAcci':'MM/DD/YYYY',
'formQuery:radioFormat':'excel',
'formQuery:buttonQueryId':'Submit',
'formQuery:buttonResetId':'Clear',
'formQuery_SUBMIT':1,
'javax.faces.ViewState':'j_id6:j_id7'}
headers = {'Accept':'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8',
'Accept-Encoding':'gzip, deflate',
'Accept-Language':'en-US,en;q=0.8',
'Cache-Control':'max-age=0',
'Connection':'keep-alive',
'Content-Length':'289',
'Content-Type':'application/x-www-form-urlencoded',
'Cookie':'__utma=116865050.46345168.1496938709.1498690358.1498763206.4; __utmz=116865050.1498763206.4.4.utmccn=(referral)|utmcsr=city.milwaukee.gov|utmcct=/DownloadMapData3497.htm|utmcmd=referral; JSESSIONID=0001kTUljano4n8x8_sv0IhnhPS:166pun9c5; _gat_gacity=1; _gat_gaitmd=1; _ga=GA1.2.1203834187.1496847807; _gid=GA1.2.1853685424.1499888152',
'Host':'itmdapps.milwaukee.gov',
'Origin':'http://itmdapps.milwaukee.gov',
'Referer':'http://itmdapps.milwaukee.gov/publicApplication_QD/queryDownload/aldermanicDistfm.faces',
'Upgrade-Insecure-Requests':'1',
'User-Agent':'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/59.0.3071.115 Safari/537.36'
}
url = 'http://itmdapps.milwaukee.gov/publicApplication_QD/queryDownload/aldermanicDistfm.faces'
month_list = ['jan','feb','mar','april','may','june','july','aug','sep','oct',
'nov','dec']
for x in range(2005,2017):
if(x==2015):
continue
try:
os.mkdir(f'data_{x}')
except:
pass
for index,month in enumerate(month_list):
m_num = 1 + index
month_end = calendar.monthrange(x,m_num)[1]
try:
os.mkdir(f'./data_{x}/{month}')
except:
pass
for ald in range(1,16):
data['formQuery:menuAldId'] = ald
if(len(str(m_num))== 1):
data['dateFromCrime'] = f'0{m_num}/01/{x}'
data['dateToCrime'] = f'0{m_num}/{month_end}/{x}'
else:
data['dateFromCrime'] = f'{m_num}/01/{x}'
data['dateToCrime'] = f'{m_num}/{month_end}/{x}'
r = requests.post(url,data=data, headers=headers)
if(r.status_code == 200):
print(f'Okay:./data_{x}/{month}/ald{ald}.xls')
else:
print(f'Not Okay: ./data_{x}/{month}/ald{ald}.xls')
with open(f'./data_{x}/{month}/ald{ald}.xls','wb') as f:
f.write(r.content)
calendar.monthrange(2002,1)
r = requests.post(url,data=data, headers=headers)
with open('test.xls','wb') as f:
f.write(r.content)
```
| github_jupyter |
# Tic Tac Toe
By Devesh Gupta
Func Display Board
```
from IPython.display import clear_output
def display_board(board):
clear_output()
print(' '+board[7]+'|'+board[8]+'|'+board[9])
print('-------')
print(' '+board[4]+'|'+board[5]+'|'+board[6])
print('-------')
print(' '+board[1]+'|'+board[2]+'|'+board[3])
```
Func Player input
```
def player_input():
marker = ''
while not (marker == 'X' or marker == 'O'):
marker = input('Player 1: Do you want to be X or O? ').upper()
if marker == 'X':
return ('X', 'O')
else:
return ('O', 'X')
```
Func Place marker
```
def place_marker(board,marker,position):
board[position] = marker
```
Func Win check
```
def win_check(board,mark):
return ((board[7] == mark and board[8] == mark and board[9] == mark) or # across the top
(board[4] == mark and board[5] == mark and board[6] == mark) or # across the middle
(board[1] == mark and board[2] == mark and board[3] == mark) or # across the bottom
(board[7] == mark and board[4] == mark and board[1] == mark) or # down the middle
(board[8] == mark and board[5] == mark and board[2] == mark) or # down the middle
(board[9] == mark and board[6] == mark and board[3] == mark) or # down the right side
(board[7] == mark and board[5] == mark and board[3] == mark) or # diagonal
(board[9] == mark and board[5] == mark and board[1] == mark)) # diagonal
```
Func Choose first randomly
```
import random
def choose_first():
flip = random.randint(0,1)
if flip == 0:
return 'Player1'
else:
return 'Player2'
```
Func Space check
```
def space_check(board,position):
return (board[position]==' ')
```
Func Full board check
```
def full_board_check(board):
for i in range(1,10):
if space_check(board,i):
return False
#Return True if the board is Full
return True
```
Func player choice
```
def player_choice(board):
position = 0
while position not in [1,2,3,4,5,6,7,8,9] or not space_check(board,position):
position = int(input("Choose a position (1-9) : "))
return position
```
Func to replay
```
def replay():
choice = input("Play again? Enter 'Yes' or 'No' - ")
return choice == 'Yes'
```
# Driver Func
```
print('===== Welcome to Tic Tac Toe! =====')
while True:
# Reset the board
theBoard = [' '] * 10
player1_marker, player2_marker = player_input()
turn = choose_first()
print(turn + ' will go first.')
play_game = input('Are you ready to play? Enter Y(for Yes) or N(for No).')
if play_game.lower()[0] == 'y':
game_on = True
else:
game_on = False
while game_on:
if turn == 'Player 1':
# Player1's turn.
print('Player 1\'s turn :-')
display_board(theBoard)
position = player_choice(theBoard)
place_marker(theBoard, player1_marker, position)
if win_check(theBoard, player1_marker):
display_board(theBoard)
print('Congratulations! Player 1 has won!')
game_on = False
else:
if full_board_check(theBoard):
display_board(theBoard)
print('Its a TIE!!')
break
else:
turn = 'Player 2'
else:
# Player2's turn.
print('Player 2\'s turn :-')
display_board(theBoard)
position = player_choice(theBoard)
place_marker(theBoard, player2_marker, position)
if win_check(theBoard, player2_marker):
display_board(theBoard)
print('Congratulations! Player 2 has won!')
game_on = False
else:
if full_board_check(theBoard):
display_board(theBoard)
print('Its a TIE!!')
break
else:
turn = 'Player 1'
if not replay():
break
```
| github_jupyter |
# Data analysis for Almentor Facebook Data
### read required libraries
```
import pandas as pd
import seaborn as sns
import regex as re
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import datetime
```
## Read and optemize files
#### reading text data file
```
## open the file and read it
with open("C:\\Users\\20812018100700\\working\\work\\rd_part\\result_full.txt", encoding='utf-8')as f:
result = f.read()
## split .txt file
result = re.split("done [0-9$,.%]+\d*\n", result)
```
#### reading orignal data frame
```
## read DataFrame
df = pd.read_csv("C:\\Users\\20812018100700\\working\\work\\rd_part\\test2_data.csv")
## drop null values
df = df[df["message"].notnull()]
## reset the index of DF
df.reset_index(drop=True, inplace=True)
## slice the DF to the len of our data
df = df.iloc[:len(result), :]
df.info()
```
### extract values from text file
```
## store values in dict
data = {
"Translate": [],
"is_human" : [],
"sentment" : [],
"positive" : [],
"neutral" : [],
"negative" : []
}
## loop for each value in result list extract is_human
## if not human: will extract the Translation, Sentment overall rank, sentment score
## else the sentment is nutral/no translation
for i in range(len(result)):
iz = re.search('(?<=is human: )(.*)', result[i]).groups()[0]
data["is_human"].append(iz[0])
if (iz == "True"):
data["sentment"].append("neutral")
data["Translate"].append(result[i].split(" \n\n")[0])
h = ['positive=0.00', 'neutral=1.00', 'negative=0.00 ']
for j in h:
s = j.split("=")
data[s[0]].append(s[1])
else:
m = re.search('(?<=Document Sentiment: )(.*)', result[i]).groups()
data["sentment"].append(m[0])
n = re.search('(?<=Translate: )(.*)', result[i]).groups()
data["Translate"].append(n[0])
h = str(re.search('(?<=Overall scores: )(.*)', result[i]).groups()[0]).split("; ")
for j in h:
s = j.split("=")
data[s[0]].append(s[1])
## convert dict to dataFrame
df1= pd.DataFrame(data)
df1.sample(10)
```
### merge the 2 dataframes to 1 df
```
## merge 2 dataFrames into the index
ndf = pd.merge(df, df1, left_index=True, right_index=True)
ndf.head(25)
```
#### save dataframe to .csv file
```
ndf.to_csv("concatnated_DF", index=False)
```
# read the data again
```
ndf = pd.read_csv("C:\\Users\\20812018100700\\working\\work\\rd_part\\concatnated_DF.csv")
```
#### explore some data
```
## extract date from created_time
ndf["created_time"] = ndf["created_time"].str.slice(0, 7)
ndf["created_time"]
ndf.is_human.unique()
ndf.query('is_human =="N" and sentment == "negative"').sample(10)
```
## explore and fix the data
```
## drop unused col
ndf.drop("from", axis=1, inplace = True)
ndf.info()
ndf["created_time"] = ndf["created_time"].str.slice(0, 7)
ndf["created_time"] = pd.to_datetime(ndf["created_time"])
ndf["date"] = ndf["created_time"].dt.date
ndf.info()
ndf.query('date == datetime.date(2021, 9, 1)')
ndf.drop(index=17, inplace = True)
ndf.reset_index(drop=True, inplace=True)
ndf.sentment.unique()
## convert rate ti float
ndf["positive"] = ndf["positive"].astype('float')
ndf["neutral"] = ndf["neutral"].astype('float')
ndf["negative"] = ndf["negative"].astype('float')
## checking distripution of the rate
ndf.positive.describe()
ndf.negative.describe()
plot = plt.subplots(figsize = (16,10))
plot = sns.countplot(data = ndf, x ="date",hue = "sentment", palette = 'Paired')
_ = plt.xticks(rotation=45)
```
## select non-human comments
```
## select not human data to explore
info_df = ndf.query('is_human != "T"')
print(info_df.info())
info_df.head()
## select non natural comments
info_df = ndf.query('neutral <0.5')
info_df.reset_index(drop=True, inplace=True)
print(info_df.info())
info_df.head()
info_df.positive.describe()
info_df.negative.describe()
fig, plot = plt.subplots(figsize = (16,10))
plot = sns.lineplot(data=info_df, x="created_time", y="negative", color = "r", alpha = 0.5)
plot = sns.lineplot(data=info_df, x="created_time", y="positive", color = "g", alpha = 0.5)
plt.title('rate ber month')
plt.ylabel("value")
fig, plot = plt.subplots(figsize = (16,10))
plt.scatter(x = info_df["negative"], y =info_df["positive"] )
```
##### the corelation between positivity and negativity is linear :O
### lets check what happen in 5/2021
```
negative_df = info_df.query('date == datetime.date(2021, 5, 1) and sentment == "negative"')
negative_df.sample(10)
```
#### more than 80% of comment about "Gaza" and considerd as negative
## extract "Gaza" commetns and fix the rate
```
negative_df = negative_df[negative_df["message"].str.find("#GazaUnderAttack")!= -1]
negative_df
```
### replace old and new values
```
n_negative = negative_df["positive"].copy()
n_positive = negative_df["negative"].copy()
negative_df["negative"] = n_negative
negative_df["positive"] = n_positive
negative_df["sentment"] = "positive"
negative_df
```
#### readd data to orignal DataFrame
```
info_df.loc[negative_df.index, :] = negative_df.copy()
info_df.loc[negative_df.index, :]
```
#### replot the graph
```
fig, plot = plt.subplots(figsize = (16,10))
plot = sns.lineplot(data=info_df, x="created_time", y="negative", color = "r", alpha = 0.5)
plot = sns.lineplot(data=info_df, x="created_time", y="positive", color = "g", alpha = 0.5)
plt.title('rate ber month')
plt.ylabel("value")
```
## check distripution of rate
```
fig, plot = plt.subplots(figsize = (11,10))
plot = sns.distplot(x=info_df["positive"] )
plt.title('positive rate distripution')
fig, plot = plt.subplots(figsize = (11,10))
plot = sns.distplot(x=info_df["negative"])
info_df.is_human.unique()
plt.pie(ndf.is_human.value_counts(), labels = ndf.is_human.unique(),
startangle = 90, counterclock = False, wedgeprops = {'width' : 0.4});
plt.pie(info_df.sentment.value_counts(), labels = info_df.sentment.unique(),
startangle = 90, counterclock = False, wedgeprops = {'width' : 0.4});
colors = sns.color_palette()
plot = plt.subplots(figsize = (13,8))
plot = sns.barplot(data = info_df, x ="date", y="positive", color = colors[-1], alpha = 0.5)
plot = sns.barplot(data = info_df, x ="date", y="negative", color = colors[1], alpha = 0.5)
plt.title("positive/negative rate comparison")
plt.ylabel("Rate")
plt.xlabel("Date")
_ = plt.xticks(rotation=45)
plot = plt.subplots(figsize = (16,10))
plot = sns.countplot(data = info_df, x ="date",hue = "sentment", palette = 'Paired')
plt.title("rate comparison per month")
plt.xlabel("Data")
plt.ylabel("Count")
_ = plt.xticks(rotation=45)
plot = plt.subplots(figsize = (16,10))
plot = sns.countplot(data = ndf,y = "date", color = colors[0])
plt.title("Page interaction per month")
_ = plt.yticks(rotation=45)
ndf.query('sentment == "mixed"').sample()
ndf.query('sentment == "neutral"').sample()
```
| github_jupyter |
```
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
<img src="http://developer.download.nvidia.com/compute/machine-learning/frameworks/nvidia_logo.png" style="width: 90px; float: right;">
## Overview
In this notebook, we want to provide an overview what HugeCTR framework is, its features and benefits. We will use HugeCTR to train a basic neural network architecture and deploy the saved model to Triton Inference Server.
<b>Learning Objectives</b>:
* Adopt NVTabular workflow to provide input files to HugeCTR
* Define HugeCTR neural network architecture
* Train a deep learning model with HugeCTR
* Deploy HugeCTR to Triton Inference Server
### Why using HugeCTR?
HugeCTR is a GPU-accelerated recommender framework designed to distribute training across multiple GPUs and nodes and estimate Click-Through Rates (CTRs).<br>
HugeCTR offers multiple advantages to train deep learning recommender systems:
1. **Speed**: HugeCTR is a highly efficient framework written C++. We experienced up to 10x speed up. HugeCTR on a NVIDIA DGX A100 system proved to be the fastest commercially available solution for training the architecture Deep Learning Recommender Model (DLRM) developed by Facebook.
2. **Scale**: HugeCTR supports model parallel scaling. It distributes the large embedding tables over multiple GPUs or multiple nodes.
3. **Easy-to-use**: Easy-to-use Python API similar to Keras. Examples for popular deep learning recommender systems architectures (Wide&Deep, DLRM, DCN, DeepFM) are available.
### Other Features of HugeCTR
HugeCTR is designed to scale deep learning models for recommender systems. It provides a list of other important features:
* Proficiency in oversubscribing models to train embedding tables with single nodes that don’t fit within the GPU or CPU memory (only required embeddings are prefetched from a parameter server per batch)
* Asynchronous and multithreaded data pipelines
* A highly optimized data loader.
* Supported data formats such as parquet and binary
* Integration with Triton Inference Server for deployment to production
### Getting Started
In this example, we will train a neural network with HugeCTR. We will use NVTabular for preprocessing.
#### Preprocessing and Feature Engineering with NVTabular
We use NVTabular to `Categorify` our categorical input columns.
```
# External dependencies
import os
import shutil
import gc
import nvtabular as nvt
import cudf
import numpy as np
from os import path
from sklearn.model_selection import train_test_split
from nvtabular.utils import download_file
```
We define our base directory, containing the data.
```
# path to store raw and preprocessed data
BASE_DIR = "/model/data/"
```
If the data is not available in the base directory, we will download and unzip the data.
```
download_file(
"http://files.grouplens.org/datasets/movielens/ml-25m.zip", os.path.join(BASE_DIR, "ml-25m.zip")
)
```
## Preparing the dataset with NVTabular
First, we take a look at the movie metadata.
Let's load the movie ratings.
```
ratings = cudf.read_csv(os.path.join(BASE_DIR, "ml-25m", "ratings.csv"))
ratings.head()
```
We drop the timestamp column and split the ratings into training and test dataset. We use a simple random split.
```
ratings = ratings.drop("timestamp", axis=1)
train, valid = train_test_split(ratings, test_size=0.2, random_state=42)
train.head()
```
We save our train and valid datasets as parquet files on disk, and below we will read them in while initializing the Dataset objects.
```
train.to_parquet(BASE_DIR + "train.parquet")
valid.to_parquet(BASE_DIR + "valid.parquet")
del train
del valid
gc.collect()
```
Let's define our categorical and label columns. Note that in that example we do not have numerical columns.
```
CATEGORICAL_COLUMNS = ["userId", "movieId"]
LABEL_COLUMNS = ["rating"]
```
Let's add Categorify op for our categorical features, userId, movieId.
```
cat_features = CATEGORICAL_COLUMNS >> nvt.ops.Categorify(cat_cache="device")
```
The ratings are on a scale between 1-5. We want to predict a binary target with 1 are all ratings >=4 and 0 are all ratings <=3. We use the LambdaOp for it.
```
ratings = nvt.ColumnGroup(["rating"]) >> (lambda col: (col > 3).astype("int8"))
```
We can visualize our calculation graph.
```
output = cat_features + ratings
(output).graph
```
We initialize our NVTabular workflow.
```
workflow = nvt.Workflow(output)
```
We initialize NVTabular Datasets, and use the part_size parameter, which defines the size read into GPU-memory at once, in nvt.Dataset.
```
train_dataset = nvt.Dataset(BASE_DIR + "train.parquet", part_size="100MB")
valid_dataset = nvt.Dataset(BASE_DIR + "valid.parquet", part_size="100MB")
```
First, we collect the training dataset statistics.
```
%%time
workflow.fit(train_dataset)
```
This step is slightly different for HugeCTR. HugeCTR expect the categorical input columns as `int64` and continuous/label columns as `float32` We can define output datatypes for our NVTabular workflow.
```
dict_dtypes = {}
for col in CATEGORICAL_COLUMNS:
dict_dtypes[col] = np.int64
for col in LABEL_COLUMNS:
dict_dtypes[col] = np.float32
```
Note: We do not have numerical output columns
```
train_dir = os.path.join(BASE_DIR, "train")
valid_dir = os.path.join(BASE_DIR, "valid")
if path.exists(train_dir):
shutil.rmtree(train_dir)
if path.exists(valid_dir):
shutil.rmtree(valid_dir)
```
In addition, we need to provide the data schema to the output calls. We need to define which output columns are `categorical`, `continuous` and which is the `label` columns. NVTabular will write metadata files, which HugeCTR requires to load the data and optimize training.
```
workflow.transform(train_dataset).to_parquet(
output_path=BASE_DIR + "train/",
shuffle=nvt.io.Shuffle.PER_PARTITION,
cats=CATEGORICAL_COLUMNS,
labels=LABEL_COLUMNS,
dtypes=dict_dtypes,
)
workflow.transform(valid_dataset).to_parquet(
output_path=BASE_DIR + "valid/",
shuffle=False,
cats=CATEGORICAL_COLUMNS,
labels=LABEL_COLUMNS,
dtypes=dict_dtypes,
)
```
## Scaling Accelerated training with HugeCTR
HugeCTR is a deep learning framework dedicated to recommendation systems. It is written in CUDA C++. As HugeCTR optimizes the training in CUDA++, we need to define the training pipeline and model architecture and execute it via the commandline. We will use the Python API, which is similar to Keras models.
HugeCTR has three main components:
* Solver: Specifies various details such as active GPU list, batchsize, and model_file
* Optimizer: Specifies the type of optimizer and its hyperparameters
* Model: Specifies training/evaluation data (and their paths), embeddings, and dense layers. Note that embeddings must precede the dense layers
**Solver**
Let's take a look on the parameter for the `Solver`. We should be familiar from other frameworks for the hyperparameter.
```
solver = hugectr.solver_parser_helper(
- vvgpu: GPU indices used in the training process, which has two levels. For example: [[0,1],[1,2]] indicates that two nodes are used in the first node. GPUs 0 and 1 are used while GPUs 1 and 2 are used for the second node. It is also possible to specify non-continuous GPU indices such as [0, 2, 4, 7]
- max_iter: Total number of training iterations
- batchsize: Minibatch size used in training
- display: Intervals to print loss on the screen
- eval_interval: Evaluation interval in the unit of training iteration
- max_eval_batches: Maximum number of batches used in evaluation. It is recommended that the number is equal to or bigger than the actual number of bathces in the evaluation dataset.
If max_iter is used, the evaluation happens for max_eval_batches by repeating the evaluation dataset infinitely.
On the other hand, with num_epochs, HugeCTR stops the evaluation if all the evaluation data is consumed
- batchsize_eval: Maximum number of batches used in evaluation. It is recommended that the number is equal to or
bigger than the actual number of bathces in the evaluation dataset
- mixed_precision: Enables mixed precision training with the scaler specified here. Only 128,256, 512, and 1024 scalers are supported
)
```
**Optimizer**
The optimizer is the algorithm to update the model parameters. HugeCTR supports the common algorithms.
```
optimizer = CreateOptimizer(
- optimizer_type: Optimizer algorithm - Adam, MomentumSGD, Nesterov, and SGD
- learning_rate: Learning Rate for optimizer
)
```
**Model**
We initialize the model with the solver and optimizer:
```
model = hugectr.Model(solver, optimizer)
```
We can add multiple layers to the model with `model.add` function. We will focus on:
- `Input` defines the input data
- `SparseEmbedding` defines the embedding layer
- `DenseLayer` defines dense layers, such as fully connected, ReLU, BatchNorm, etc.
**HugeCTR organizes the layers by names. For each layer, we define the input and output names.**
Input layer:
This layer is required to define the input data.
```
hugectr.Input(
data_reader_type: Data format to read
source: The training dataset file list.
eval_source: The evaluation dataset file list.
check_type: The data error detection machanism (Sum: Checksum, None: no detection).
label_dim: Number of label columns
label_name: Name of label columns in network architecture
dense_dim: Number of continous columns
dense_name: Name of contiunous columns in network architecture
slot_size_array: The list of categorical feature cardinalities
data_reader_sparse_param_array: Configuration how to read sparse data
sparse_names: Name of sparse/categorical columns in network architecture
)
```
SparseEmbedding:
This layer defines embedding table
```
hugectr.SparseEmbedding(
embedding_type: Different embedding options to distribute embedding tables
max_vocabulary_size_per_gpu: Maximum vocabulary size or cardinality across all the input features
embedding_vec_size: Embedding vector size
combiner: Intra-slot reduction op (0=sum, 1=average)
sparse_embedding_name: Layer name
bottom_name: Input layer names
)
```
DenseLayer:
This layer is copied to each GPU and is normally used for the MLP tower.
```
hugectr.DenseLayer(
layer_type: Layer type, such as FullyConnected, Reshape, Concat, Loss, BatchNorm, etc.
bottom_names: Input layer names
top_names: Layer name
...: Depending on the layer type additional parameter can be defined
)
```
## Let's define our model
We walked through the documentation, but it is useful to understand the API. Finally, we can define our model. We will write the model to `./model.py` and execute it afterwards.
We need the cardinalities of each categorical feature to assign as `slot_size_array` in the model below.
```
from nvtabular.ops import get_embedding_sizes
embeddings = get_embedding_sizes(workflow)
print(embeddings)
```
In addition, we need the total cardinalities to be assigned as `max_vocabulary_size_per_gpu` parameter.
```
total_cardinality = embeddings["userId"][0] + embeddings["movieId"][0]
total_cardinality
%%writefile './model.py'
import hugectr
from mpi4py import MPI # noqa
solver = hugectr.solver_parser_helper(
vvgpu=[[0]],
max_iter=2000,
batchsize=2048,
display=100,
eval_interval=200,
batchsize_eval=2048,
max_eval_batches=160,
i64_input_key=True,
use_mixed_precision=False,
repeat_dataset=True,
snapshot=1900,
)
optimizer = hugectr.optimizer.CreateOptimizer(
optimizer_type=hugectr.Optimizer_t.Adam, use_mixed_precision=False
)
model = hugectr.Model(solver, optimizer)
model.add(
hugectr.Input(
data_reader_type=hugectr.DataReaderType_t.Parquet,
source="/model/data/train/_file_list.txt",
eval_source="/model/data/valid/_file_list.txt",
check_type=hugectr.Check_t.Non,
label_dim=1,
label_name="label",
dense_dim=0,
dense_name="dense",
slot_size_array=[162542, 56586],
data_reader_sparse_param_array=[
hugectr.DataReaderSparseParam(hugectr.DataReaderSparse_t.Distributed, 3, 1, 2)
],
sparse_names=["data1"],
)
)
model.add(
hugectr.SparseEmbedding(
embedding_type=hugectr.Embedding_t.DistributedSlotSparseEmbeddingHash,
max_vocabulary_size_per_gpu=219128,
embedding_vec_size=16,
combiner=0,
sparse_embedding_name="sparse_embedding1",
bottom_name="data1",
)
)
model.add(
hugectr.DenseLayer(
layer_type=hugectr.Layer_t.Reshape,
bottom_names=["sparse_embedding1"],
top_names=["reshape1"],
leading_dim=32,
)
)
model.add(
hugectr.DenseLayer(
layer_type=hugectr.Layer_t.InnerProduct,
bottom_names=["reshape1"],
top_names=["fc1"],
num_output=128,
)
)
model.add(
hugectr.DenseLayer(
layer_type=hugectr.Layer_t.ReLU,
bottom_names=["fc1"],
top_names=["relu1"],
)
)
model.add(
hugectr.DenseLayer(
layer_type=hugectr.Layer_t.InnerProduct,
bottom_names=["relu1"],
top_names=["fc2"],
num_output=128,
)
)
model.add(
hugectr.DenseLayer(
layer_type=hugectr.Layer_t.ReLU,
bottom_names=["fc2"],
top_names=["relu2"],
)
)
model.add(
hugectr.DenseLayer(
layer_type=hugectr.Layer_t.InnerProduct,
bottom_names=["relu2"],
top_names=["fc3"],
num_output=1,
)
)
model.add(
hugectr.DenseLayer(
layer_type=hugectr.Layer_t.BinaryCrossEntropyLoss,
bottom_names=["fc3", "label"],
top_names=["loss"],
)
)
model.compile()
model.summary()
model.fit()
!python model.py
```
We trained our model.
After training terminates, we can see that two `.model` files are generated. We need to move them inside `1` folder under the `movielens_hugectr` folder. Let's create these folders first.
```
!mkdir -p /model/movielens_hugectr/1
```
Now we move our saved `.model` files inside `1` folder.
```
!mv *.model /model/movielens_hugectr/1/
```
Note that these stored `.model` files will be used in the inference. Now we have to create a JSON file for inference which has a similar configuration as our training file. We should remove the solver and optimizer clauses and add the inference clause in the JSON file. The paths of the stored dense model and sparse model(s) should be specified at dense_model_file and sparse_model_file within the inference clause. We need to make some modifications to data in the layers clause. Besides, we need to change the last layer from BinaryCrossEntropyLoss to Sigmoid. The rest of "layers" should be exactly the same as that in the training model.py file.
Now let's create a `movielens.json` file inside the `movielens/1` folder. We have already retrieved the cardinality of each categorical column using `get_embedding_sizes` function above. We will use these cardinalities below in the `movielens.json` file as well.
```
%%writefile '/model/movielens_hugectr/1/movielens.json'
{
"inference": {
"max_batchsize": 64,
"hit_rate_threshold": 0.6,
"dense_model_file": "/model/models/movielens/1/_dense_1900.model",
"sparse_model_file": "/model/models/movielens/1/0_sparse_1900.model",
"label": 1,
"input_key_type": "I64"
},
"layers": [
{
"name": "data",
"type": "Data",
"format": "Parquet",
"slot_size_array": [162542, 56586],
"source": "/model/data/train/_file_list.txt",
"eval_source": "/model/data/valid/_file_list.txt",
"check": "Sum",
"label": {"top": "label", "label_dim": 1},
"dense": {"top": "dense", "dense_dim": 0},
"sparse": [
{
"top": "data1",
"type": "DistributedSlot",
"max_feature_num_per_sample": 3,
"slot_num": 2
}
]
},
{
"name": "sparse_embedding1",
"type": "DistributedSlotSparseEmbeddingHash",
"bottom": "data1",
"top": "sparse_embedding1",
"sparse_embedding_hparam": {
"max_vocabulary_size_per_gpu": 219128,
"embedding_vec_size": 16,
"combiner": 0
}
},
{
"name": "reshape1",
"type": "Reshape",
"bottom": "sparse_embedding1",
"top": "reshape1",
"leading_dim": 32
},
{
"name": "fc1",
"type": "InnerProduct",
"bottom": "reshape1",
"top": "fc1",
"fc_param": {"num_output": 128}
},
{"name": "relu1", "type": "ReLU", "bottom": "fc1", "top": "relu1"},
{
"name": "fc2",
"type": "InnerProduct",
"bottom": "relu1",
"top": "fc2",
"fc_param": {"num_output": 128}
},
{"name": "relu2", "type": "ReLU", "bottom": "fc2", "top": "relu2"},
{
"name": "fc3",
"type": "InnerProduct",
"bottom": "relu2",
"top": "fc3",
"fc_param": {"num_output": 1}
},
{"name": "sigmoid", "type": "Sigmoid", "bottom": "fc3", "top": "sigmoid"}
]
}
```
Now we can save our models to be deployed at the inference stage. To do so we will use `export_hugectr_ensemble` method below. With this method, we can generate the `config.pbtxt` files automatically for each model. In doing so, we should also create a `hugectr_params` dictionary, and define the parameters like where the `movielens.json` file will be read, `slots` which corresponds to number of categorical features, `embedding_vector_size`, `max_nnz`, and `n_outputs` which is number of outputs.
The script below creates an ensemble triton server model where
- `workflow` is the the nvtabular workflow used in preprocessing,
- `hugectr_model_path` is the HugeCTR model that should be served. This path includes the `.model` files.
- `name` is the base name of the various triton models
- `output_path` is the path where is model will be saved to.
```
from nvtabular.inference.triton import export_hugectr_ensemble
hugectr_params = dict()
hugectr_params["config"] = "/model/models/movielens/1/movielens.json"
hugectr_params["slots"] = 2
hugectr_params["max_nnz"] = 2
hugectr_params["embedding_vector_size"] = 16
hugectr_params["n_outputs"] = 1
export_hugectr_ensemble(
workflow=workflow,
hugectr_model_path="/model/movielens_hugectr/1/",
hugectr_params=hugectr_params,
name="movielens",
output_path="/model/models/",
label_columns=["rating"],
cats=CATEGORICAL_COLUMNS,
max_batch_size=64,
)
```
After we run the script above, we will have three model folders saved as `movielens_nvt`, `movielens` and `movielens_ens`. Now we can move to the next notebook, `movielens-HugeCTR-inference`, to send request to the Triton Inference Server using the saved ensemble model.
| github_jupyter |

<div class="alert alert-block alert-info">
<b>Important:</b> This notebook uses <code>ipywidgets</code> that take advantage of the javascript interface in a web browser. The downside is the functionality does not render well on saved notebooks. Run this notebook locally to see the widgets in action.
</div>
# Qiskit Jupyter Tools
Qiskit was designed to be used inside of the Jupyter notebook interface. As such it includes many useful routines that take advantage of this platform, and make performing tasks like exploring devices and tracking job progress effortless.
Loading all the qiskit Jupyter goodness is done via:
```
from qiskit import *
import qiskit.providers.ibmq.jupyter # This is the where the magic happens (literally).
```
## Table of contents
1) [IQX Dashboard](#dashboard)
2) [Backend Details](#details)
To start, load your IQX account information and select a provider:
```
IBMQ.load_account();
provider = IBMQ.get_provider(group='open')
```
## IQX Dashboard <a name="dashboard"></a>
Perhaps the most useful Jupyter tool is the `iqx_dashboard`. This widget consists of a `Devices` tab and a `Jobs` tab. The `Devices` tab provides an overview of all the devices you have access to. The `Jobs` tab automatically tracks and displays information for the jobs submitted in this session.
To start the dashboard you run the Jupyter magic:
```
%iqx_dashboard
```
You should now see a small window titled "IQX Dashboard" in the upper left corner of the notebook. Click on the drop down symbol to see the two tabs. The `Devices` tab may take a few seconds to load as it needs to communicate with the server for device information. The `Jobs` tab should contain no job information as none has been submitted yet.
### Getting an Overview of Backends
The `Devices` tab provides an overview of all the backends you have access to. You can use it to compare, for example, the average CNOT error rates. In addition, the number of pending jobs on the devices is continuously being updated along with the operational status.
### Automatic Job Tracking
The `Jobs` tab automatically tracks and displays information for the jobs submitted in this session.
Now, let's submit a job to a device to see this in action:
```
backend = provider.get_backend('ibmq_essex')
qc = QuantumCircuit(2, 2)
qc.h(0)
qc.cx(0, 1)
qc.measure([0,1], [0,1])
job = execute(qc, backend)
```
Click on the `Jobs` tab and you will see that the job has been added to the list of jobs. Its status, queue position (if any), and estimated start time are being automatically tracked and updated. If the job is running, the scheduling mode for the job is also displayed. For example, if the job status is `RUNNING[F]`, that means the job is actively running and was scheduled using a fair-share algorithm. The button to the left of a job ID allows you to cancel the job.
If you want to kill the dashboard you can do so by calling:
```
%disable_ibmq_dashboard
```
Although the dashboard itself is killed, the underlying framework is still tracking jobs for you and will show this information if loaded once again.
## Viewing Backend Details <a name="details"></a>
The IBM Quantum devices contain a large amount of configuration data and properties. This information can be retrieved by calling:
```
config = backend.configuration()
params = backend.properties()
```
However, parsing through this information quickly becomes tedious. Instead, all the information for a single backend can be displayed graphically by just calling the backend instance itself:
```
backend
```
This widget displays all the information about a backend in a single tabbed-window.
```
import qiskit.tools.jupyter
%qiskit_version_table
%qiskit_copyright
```
| github_jupyter |
# Millikan Oil Drop
___**Meansurement of the electron charge**
```
rho=886 # kg/m^3
dV = .5 #volts
dd = .000005 # meters
dP = 5 # pascals
g=9.8 # m/s^2
eta= 1.8330*10**(-5) # N*s/m^2
b=8.20*10**(-3) # Pa*m
p=101325 #Pa
V=500 #V
e=1.6*10**(-19)
d_array=10**(-3)*np.array([7.55,7.59,7.60,7.60,7.60,7.61]) # unit: m
d=d_array.mean()
d_std=d_array.std()
print("d_mean: ",d_mean)
print("d_std: ",d_std)
def reject_outliers(data, m=2):
'''
remove anomalous data points that outside 2 standard deviation in the array
'''
return data[abs(data - np.mean(data)) < m * np.std(data)]
```
**Load data from files**
```
data_path = "/Users/Angel/Documents/MilikanData/"
statistics=[]
for file_name in os.listdir(data_path):
name=file_name[:3]
obj_drop=pd.read_csv(data_path+file_name).dropna()
# seperate rising and falling velocities, remove anomalous velocities at switching field direction
v_y=obj_drop["v_{y}"].values
y = obj_drop["y"] #y values
n_points=len(v_y)
v_r=reject_outliers(v_y[v_y>0])
v_f=reject_outliers(v_y[v_y<0])
# calculate mean and deviation
(v_r_mean,v_r_std)=(v_r.mean(),v_r.std())
(v_f_mean,v_f_std)=(np.abs(v_f.mean()),v_f.std())
# calculate other properties
a=np.sqrt((b/2/p)**2+9*eta*v_f_mean/2/rho/g)-b/(2*p) #droplet radius
m=4*np.pi/3*a**3*rho # droplet mass
q=m*g*d_mean*(v_f_mean+v_r_mean)/V/v_f_mean #droplet charge
# error propagation
dely = np.roll(y, -2)-y
delt = .4
error_y = 2e-6
error_t = .1
error_v = np.sqrt((2*error_y/dely)**2+(2*error_t/delt)**2)
error_v.pop(n_points-1)
error_v.pop(n_points-2)
error_v = np.append([0.5],error_v)
error_v = np.append(error_v, [0.5])
error_v = np.abs(v_y)*error_v
meanerror_v = error_v[~np.isinf(error_v)].mean()
dqdvf = 2*np.pi*(((b/(2*p))**2+9*eta*v_f_mean/(2*rho*g))**(-.5))*((np.sqrt(9*eta*v_f_mean/(2*rho*g)+(b/(2*p))**2)-b/(2*p))**2)*9*eta/(2*rho*g)*rho*g*d*(v_f_mean+v_r_mean)/(V*v_f_mean) + 4*np.pi/3*((np.sqrt((b/(2*p))**2+9*eta*v_f_mean/(2*rho*g))-b/(2*p))**3)*(V*v_f_mean*rho*g*d*v_r_mean-rho*g*d*(v_f_mean+v_r_mean)*V)/((V*v_f_mean)**2)
dqdvr = 4*np.pi/3*((np.sqrt((b/(2*p))**2+9*eta*v_f_mean/(2*rho*g))-b/(2*p))**3)*(rho*g*d/V)
dqdV = -4*np.pi/3*((np.sqrt((b/(2*p))**2+9*eta*v_f_mean/(2*rho*g))-b/(2*p))**3)*(v_f_mean*rho*g*d*(v_f_mean+v_r_mean)/((V*v_f_mean)**2))
dqdd = 4*np.pi/3*((np.sqrt((b/(2*p))**2+9*eta*v_f_mean/(2*rho*g))-b/(2*p))**3)*rho*g*(v_f_mean+v_r_mean)/(V*v_f_mean)
dqdP1 = 2*np.pi*((np.sqrt((b/(2*p))**2+9*eta*v_f_mean/(2*rho*g))-b/(2*p))**2)*rho*g*d*(v_f_mean+v_r_mean)/(V*v_f_mean)
dqdP2 = -(((b/(2*p))**2+9*eta*v_f_mean/(2*rho*g))**(-.5))*(b**2)/(2*p**3)+b/(4*p**2)
error_func = np.sqrt(((dqdvf)*(meanerror_v))**2+((dqdvr)*(meanerror_v))**2+((dqdV)*(dV))**2+((dqdd)*(dd))**2+((dqdP1*dqdP2)*(dP))**2)
statistics.append(np.array((name,n_points,v_r_mean,v_r_std,v_f_mean,v_f_std, meanerror_v, a,m,q, error_func)))
```
Calculation of the attached charge
```
labels = ["name","n_points","v_r_mean","v_r_std","v_f_mean","v_f_std","meanerror_v","a","m","q","q_error"]
overall = pd.DataFrame(statistics,columns=labels,dtype="float64")
overall
import matplotlib.pylab as plt
plt.figure().dpi=100
plt.xlabel("Charge attached")
plt.ylabel("Number of droplets")
plt.title("Histogram of charge carried by droplets")
(overall.q/e).hist(bins=21)
def clustering(arr,x):
arr = list(arr/x)
num = int(max(arr))
clusters= []
for i in range(num+1):
clusters.append(list(filter(lambda x:i<x<i+1,arr)))
return clusters
from scipy.optimize import minimize
def obj_error(x):
test = list(map(np.mean,clustering(overall.q,x)))
estimate_delta_q = np.array(test[:-1])-np.array(test[1:])
estimate_e = estimate_delta_q[~np.isnan(estimate_delta_q)]
estimate_e = estimate_e*e
return abs(estimate_e.mean())
obj_error(e)
#valuee = minimize(obj_error,.8e-19)
#print(valuee.x)
```
| github_jupyter |
```
import json
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.collections import PatchCollection
from matplotlib.patches import Rectangle
from matplotlib.lines import Line2D
from tqdm import tqdm
import mc_bfm_2d
from importlib import reload
reload(mc_bfm_2d)
%matplotlib inline
polymers = []
for j in range(48):
monomers= [mc_bfm_2d.Monomer([1 + 2*j, 1 + 2*i]) for i in range(8)]
for i in range(len(monomers)-1):
mc_bfm_2d.Bond(monomers[i], monomers[i+1])
polymers.append(mc_bfm_2d.Polymer(monomers))
occlusion = mc_bfm_2d.Occlusion(
sorted(set(
[(x, 0) for x in range(100)] +
[(x, 99) for x in range(100)] +
[(0, y) for y in range(100)] +
[(99, y) for y in range(100)] +
[(x, 50) for x in range(45)] +
[(x, 50) for x in range(55, 100)]
))
)
config = mc_bfm_2d.Configuration(100, 100, polymers, occlusions=[occlusion])
def plot_config(config):
monomer_boxes = [Rectangle(monomer.lower_right_occupancy, 2, 2)
for polymer in config.polymers
for monomer in polymer.monomers]
pc = PatchCollection(monomer_boxes, facecolor='blue', edgecolor='black')
plt.gca().add_collection(pc)
occlusion_boxes = [Rectangle(site, 1, 1)
for occlusion in config.occlusions
for site in occlusion.sites]
pc = PatchCollection(occlusion_boxes, facecolor='grey', edgecolor='grey')
plt.gca().add_collection(pc)
bonds = {bond
for polymer in config.polymers
for monomer in polymer.monomers
for bond in monomer.bonds}
for bond in bonds:
a = bond.a.lower_right_occupancy + np.array([1,1])
b = bond.b.lower_right_occupancy + np.array([1,1])
x, y = np.array([a,b]).T
line = Line2D(x, y, color='red', linewidth=2)
plt.gca().add_line(line)
plt.xlim(0, config.width)
plt.ylim(0, config.height)
plt.xticks([])
plt.yticks([])
plt.figure(figsize=(6,6))
plot_config(config)
plt.savefig('initial_config.png')
with open('config0.json', 'wt') as fp:
fp.write(json.dumps(config.to_json()))
plt.figure(figsize=(6,6))
with tqdm() as t:
with open('configs_long.json', 'rt') as fp:
for i,line in enumerate(fp):
json_config = json.loads(line.strip())
config = mc_bfm_2d.Configuration.from_json(json_config)
t.update()
plt.clf()
plot_config(config)
plt.xticks([])
plt.yticks([])
plt.savefig('data/movie0/%05d.png' % (i))
with open('configs_long.json', 'rt') as fp:
for line in fp:
pass
json_config = json.loads(line.strip())
config = mc_bfm_2d.Configuration.from_json(json_config)
plt.figure(figsize=(6,6))
plot_config(config)
plt.savefig('final_config.png')
plt.hist(np.array([p.get_center_of_mass() for p in config.polymers])[::, 1])
acc_center_of_mass = []
with open('configs_long.json', 'rt') as fp:
for line in fp:
json_config = json.loads(line.strip())
config = mc_bfm_2d.Configuration.from_json(json_config)
acc_center_of_mass.append(
np.array([p.get_center_of_mass() for p in config.polymers])[::, 1].mean()
)
plt.plot(acc_center_of_mass)
acc_above_threshold = []
with open('configs_long.json', 'rt') as fp:
for line in fp:
json_config = json.loads(line.strip())
config = mc_bfm_2d.Configuration.from_json(json_config)
acc_above_threshold.append(
(np.array([p.get_center_of_mass() for p in config.polymers])[::, 1] > 50).sum()
)
plt.plot(acc_above_threshold)
plt.xlabel('Simulation Cycles')
plt.ylabel('Number of Polymer Chains\nDiffused through Pore')
plt.savefig('polymer_diffusion.png')
```
| github_jupyter |
# 3.6 Refinements with federated learning
## Data loading and preprocessing
```
# read more: https://www.tensorflow.org/federated/tutorials/federated_learning_for_text_generation
import nest_asyncio # pip install nest_asyncio
import tensorflow_federated as tff # pip install tensorflow_federated
import collections
import functools
import os
import time
import numpy as np
import tensorflow as tf
#nest_asyncio.apply()
tf.compat.v1.enable_v2_behavior()
np.random.seed(0)
# Test the TFF is working:
tff.federated_computation(lambda: 'Hello, World!')()
import numpy as np
# A fixed vocabularly of ASCII chars that occur in the works of Shakespeare and Dickens:
vocab = list('dhlptx@DHLPTX $(,048cgkoswCGKOSW[_#\'/37;?bfjnrvzBFJNRVZ"&*.26:\naeimquyAEIMQUY]!%)-159\r')
# Creating a mapping from unique characters to indices
char2idx = {u:i for i, u in enumerate(vocab)}
idx2char = np.array(vocab)
```
## Data
```
train_data, test_data = tff.simulation.datasets.shakespeare.load_data()
# Here the play is "The Tragedy of King Lear" and the character is "King".
raw_example_dataset = train_data.create_tf_dataset_for_client(
'THE_TRAGEDY_OF_KING_LEAR_KING')
# To allow for future extensions, each entry x
# is an OrderedDict with a single key 'snippets' which contains the text.
for x in raw_example_dataset.take(2):
print(x['snippets'])
# Input pre-processing parameters
SEQ_LENGTH = 100
BATCH_SIZE = 8
BUFFER_SIZE = 10000 # For dataset shuffling
```
## Text generation
```
import tensorflow as tf
# Construct a lookup table to map string chars to indexes,
# using the vocab loaded above:
table = tf.lookup.StaticHashTable(
tf.lookup.KeyValueTensorInitializer(
keys=vocab, values=tf.constant(list(range(len(vocab))),
dtype=tf.int64)),
default_value=0)
def to_ids(x):
s = tf.reshape(x['snippets'], shape=[1])
chars = tf.strings.bytes_split(s).values
ids = table.lookup(chars)
return ids
def split_input_target(chunk):
input_text = tf.map_fn(lambda x: x[:-1], chunk)
target_text = tf.map_fn(lambda x: x[1:], chunk)
return (input_text, target_text)
def preprocess(dataset):
return (
# Map ASCII chars to int64 indexes using the vocab
dataset.map(to_ids)
# Split into individual chars
.unbatch()
# Form example sequences of SEQ_LENGTH +1
.batch(SEQ_LENGTH + 1, drop_remainder=True)
# Shuffle and form minibatches
.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
# And finally split into (input, target) tuples,
# each of length SEQ_LENGTH.
.map(split_input_target))
example_dataset = preprocess(raw_example_dataset)
print(example_dataset.element_spec)
import os
def load_model(batch_size):
urls = {
1: 'https://storage.googleapis.com/tff-models-public/dickens_rnn.batch1.kerasmodel',
8: 'https://storage.googleapis.com/tff-models-public/dickens_rnn.batch8.kerasmodel'}
assert batch_size in urls, 'batch_size must be in ' + str(urls.keys())
url = urls[batch_size]
local_file = tf.keras.utils.get_file(os.path.basename(url), origin=url)
return tf.keras.models.load_model(local_file, compile=False)
def generate_text(model, start_string):
# Evaluation step (generating text using the learned model)
# Number of characters to generate
num_generate = 1000
# Converting our start string to numbers (vectorizing)
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, 0)
# Empty string to store our results
text_generated = []
# Low temperatures results in more predictable text.
# Higher temperatures results in more surprising text.
# Experiment to find the best setting.
temperature = 1.0
# Here batch size == 1
model.reset_states()
for i in range(num_generate):
predictions = model(input_eval)
# remove the batch dimension
predictions = tf.squeeze(predictions, 0)
# using a categorical distribution to predict the character returned by the model
predictions = predictions / temperature
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1,0].numpy()
# We pass the predicted character as the next input to the model
# along with the previous hidden state
input_eval = tf.expand_dims([predicted_id], 0)
text_generated.append(idx2char[predicted_id])
return (start_string + ''.join(text_generated))
# Text generation requires a batch_size=1 model.
keras_model_batch1 = load_model(batch_size=1)
print(generate_text(keras_model_batch1, 'What of TensorFlow Federated, you ask? '))
BATCH_SIZE = 8 # The training and eval batch size for the rest of this tutorial.
keras_model = load_model(batch_size=BATCH_SIZE)
keras_model.compile(loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True))
```
## Federated learning
```
import collections
# Clone the keras_model inside `create_tff_model()`, which TFF will
# call to produce a new copy of the model inside the graph that it will
# serialize. Note: we want to construct all the necessary objects we'll need
# _inside_ this method.
def create_tff_model():
# TFF uses a `dummy_batch` so it knows the types and shapes
# that your model expects.
x = np.random.randint(1, len(vocab), size=[BATCH_SIZE, SEQ_LENGTH])
dummy_batch = collections.OrderedDict(x=x, y=x)
keras_model_clone = tf.keras.models.clone_model(keras_model)
return tff.learning.from_keras_model(
keras_model_clone,
dummy_batch=dummy_batch,
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True))
fed_avg = tff.learning.build_federated_averaging_process(
model_fn=create_tff_model,
client_optimizer_fn=lambda: tf.keras.optimizers.SGD(lr=0.5))
nest_asyncio.apply()
NUM_ROUNDS = 5
state = fed_avg.initialize()
for _ in range(NUM_ROUNDS):
state, metrics = fed_avg.next(state, [example_dataset.take(5)])
print(f'loss={metrics.loss}')
```
| github_jupyter |
# 0.前言
这个文档主要是用来入门下XGBOOST,主要就是参考的https://blog.csdn.net/qq_24519677/article/details/81869196
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import xgboost as xgb
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn import cross_validation
from sklearn.preprocessing import LabelEncoder
import sklearn
import warnings
warnings.filterwarnings('ignore')
```
# 1.数据特征处理
```
train = pd.read_csv('data/train.csv')
test = pd.read_csv('data/test.csv')
train.info() # 打印训练数据的信息
test.info()
```
对数据的缺失值进行处理,这里采用的方法是对连续值用该列的平均值进行填充,非连续值用该列的众数进行填充,还可以使用机器学习的模型对缺失值进行预测,用预测的值来填充缺失值,该方法这里不做介绍:
```
def handle_na(train, test): # 将Cabin特征删除
fare_mean = train['Fare'].mean() # 测试集的fare特征有缺失值,用训练数据的均值填充
test.loc[pd.isnull(test.Fare), 'Fare'] = fare_mean
embarked_mode = train['Embarked'].mode() # 用众数填充
train.loc[pd.isnull(train.Embarked), 'Embarked'] = embarked_mode[0]
train.loc[pd.isnull(train.Age), 'Age'] = train['Age'].mean() # 用均值填充年龄
test.loc[pd.isnull(test.Age), 'Age'] = train['Age'].mean()
return train, test
new_train, new_test = handle_na(train, test) # 填充缺失值
```
由于Embarked,Sex,Pclass特征是离散特征,所以对其进行one-hot/get_dummies编码
```
# 对Embarked和male特征进行one-hot/get_dummies编码
new_train = pd.get_dummies(new_train, columns=['Embarked', 'Sex', 'Pclass'])
new_test = pd.get_dummies(new_test, columns=['Embarked', 'Sex', 'Pclass'])
```
然后再去除掉PassengerId,Name,Ticket,Cabin, Survived列,这里不使用这些特征做预测
```
target = new_train['Survived'].values
# 删除PassengerId,Name,Ticket,Cabin, Survived列
df_train = new_train.drop(['PassengerId','Name','Ticket','Cabin','Survived'], axis=1).values
df_test = new_test.drop(['PassengerId','Name','Ticket','Cabin'], axis=1).values
```
# 2.XGBoost模型
## 2.1使用XGBoost原生版本模型
```
X_train,X_test,y_train,y_test = train_test_split(df_train,target,test_size = 0.3,random_state = 1) # 将数据划分为训练集和测试集
data_train = xgb.DMatrix(X_train, y_train) # 使用XGBoost的原生版本需要对数据进行转化
data_test = xgb.DMatrix(X_test, y_test)
param = {'max_depth': 5, 'eta': 1, 'objective': 'binary:logistic'}
watchlist = [(data_test, 'test'), (data_train, 'train')]
n_round = 3 # 迭代训练3轮
booster = xgb.train(param, data_train, num_boost_round=n_round, evals=watchlist)
# 计算错误率
y_predicted = booster.predict(data_test)
y = data_test.get_label()
accuracy = sum(y == (y_predicted > 0.5))
accuracy_rate = float(accuracy) / len(y_predicted)
print ('样本总数:{0}'.format(len(y_predicted)))
print ('正确数目:{0}'.format(accuracy) )
print ('正确率:{0:.3f}'.format((accuracy_rate)))
```
## 2.2XGBoost的sklearn接口版本
```
X_train,X_test,y_train,y_test = train_test_split(df_train,target,test_size = 0.3,random_state = 1)
model = xgb.XGBClassifier(max_depth=3, n_estimators=200, learn_rate=0.01)
model.fit(X_train, y_train)
test_score = model.score(X_test, y_test)
print('test_score: {0}'.format(test_score))
```
利用xgboost做一次预测。
```
try_pred = X_test[[0,1],:]
try_pred
try_pred_y = y_test[0:2]
try_pred_y
pred = model.predict(try_pred)
pred
```
# 3.使用其他模型于XGBoost进行对比
```
# 应用模型进行预测
model_lr = LogisticRegression()
model_rf = RandomForestClassifier(n_estimators=200)
model_xgb = xgb.XGBClassifier(max_depth=5, n_estimators=200, learn_rate=0.01)
models = [model_lr, model_rf, model_xgb]
model_name = ['LogisticRegression', '随机森林', 'XGBoost']
cv =cross_validation.ShuffleSplit(len(df_train), n_iter=3, test_size=0.3, random_state=1)
for i in range(3):
print(model_name[i] + ":")
model = models[i]
for train, test in cv:
model.fit(df_train[train], target[train])
train_score = model.score(df_train[train], target[train])
test_score = model.score(df_train[test], target[test])
print('train score: {0:.5f} \t test score: {0:.5f}'.format(train_score, test_score))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content-dl/blob/main/projects/ComputerVision/Image_Alignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Image Alignment
**By Neuromatch Academy**
__Content creators:__ Kaleb Vinehout
**Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
<p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
---
# Intro to Image Alignment
This notebook will give you starting points to perform Spatial Transformers. These can be used for registraion of images. This is useful when comparing multiple datasets together. Check out https://arxiv.org/abs/1506.02025 for more details.These can also be used to plug into any CNN architecture to deal with dataset rotations and scale invariance in a given dataset
* Spatial transformers contain three main parts.The first is a localizaion net the second is grid generator and the last is a sampler.
## Image Alignment Applications
* To answer many biological questions, it is necessary to align sets of images together
* Use Spatial Transfomers as a preprocessing step for any CNN achitecutre. This could be done before facial recognition in order to crop and align images before spatial recognition.
#Acknowledgments:
This Notebook was developed by Kaleb Vinehout. It borrows from material by Ghassen Hamrouni, Asror Wali, and Erwin Russel.
```
%matplotlib inline
# Import dependencies
from __future__ import print_function
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torchvision
from torchvision import datasets, transforms
import matplotlib.pyplot as plt
import numpy as np
import glob
import torch
import numpy as np
import sklearn.decomposition
import matplotlib.pyplot as plt
from PIL import Image
from torchvision import transforms
from torchvision.utils import make_grid
from torchvision.datasets import ImageFolder
from skimage.util import random_noise
#from facenet_pytorch import MTCNN, InceptionResnetV1
plt.ion() # interactive mode
```
# Data loading
## loader for classic MNIST as an example
```
from six.moves import urllib
opener = urllib.request.build_opener()
opener.addheaders = [('User-agent', 'Mozilla/5.0')]
urllib.request.install_opener(opener)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# Training dataset
train_loader = torch.utils.data.DataLoader(
datasets.MNIST(root='.', train=True, download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])), batch_size=64, shuffle=True, num_workers=4)
# Test dataset
test_loader = torch.utils.data.DataLoader(
datasets.MNIST(root='.', train=False, transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])), batch_size=64, shuffle=True, num_workers=4)
```
Define functions to convert between Tensor and numpy image
```
def convert_image_np(inp):
"""Convert a Tensor to numpy image."""
inp = inp.numpy().transpose((1, 2, 0))
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
inp = std * inp + mean
inp = np.clip(inp, 0, 1)
return inp
def convert2tensor(self, args):
data = np.asarray([e[0] for e in self.binary_train_dataset])
target = np.asarray([e[1] for e in self.binary_train_dataset])
tensor_data = torch.from_numpy(data)
tensor_data = tensor_data.float()
tensor_target = torch.from_numpy(target)
train = data_utils.TensorDataset(tensor_data, tensor_target)
train_loader = data_utils.DataLoader(train, batch_size=args.batch_size, shuffle = True)
return train_loader
```
Plot the data
```
## Display Images
# Get a batch of training data
data = next(iter(test_loader))[0].to(device)
input_tensor = data.cpu()
in_grid = convert_image_np(torchvision.utils.make_grid(input_tensor))
# Plot the images
plt.imshow(in_grid)
#plot ONE image
plt.figure()
plt.imshow(torchvision.utils.make_grid(input_tensor).numpy().transpose((1, 2, 0)))
```
## loader for data from github --> modify for your data
```
#what needs to be done here to load data from github? (Hint: How did you do it durring the tutorials?)
# Training dataset
#train_loader=
# Test dataset
#test_loader =
```
# Spatial Transformer on images
## Spatial Transformer Network
```
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 10, kernel_size=5)
self.conv2 = nn.Conv2d(10, 20, kernel_size=5)
self.conv2_drop = nn.Dropout2d()
self.fc1 = nn.Linear(320, 50)
self.fc2 = nn.Linear(50, 10)
# Spatial transformer localization-network
self.localization = nn.Sequential(
nn.Conv2d(1, 8, kernel_size=7),
nn.MaxPool2d(2, stride=2),
nn.ReLU(True),
nn.Conv2d(8, 10, kernel_size=5),
nn.MaxPool2d(2, stride=2),
nn.ReLU(True)
)
# Regressor for the 3 * 2 affine matrix
self.fc_loc = nn.Sequential(
nn.Linear(10 * 3 * 3, 32),
nn.ReLU(True),
nn.Linear(32, 3 * 2)
)
# Initialize the weights/bias with identity transformation
self.fc_loc[2].weight.data.zero_()
self.fc_loc[2].bias.data.copy_(torch.tensor([1, 0, 0, 0, 1, 0], dtype=torch.float))
# Spatial transformer network forward function
def stn(self, x):
xs = self.localization(x)
xs = xs.view(-1, 10 * 3 * 3)
theta = self.fc_loc(xs)
theta = theta.view(-1, 2, 3)
grid = F.affine_grid(theta, x.size())
x = F.grid_sample(x, grid)
return x
def forward(self, x):
# transform the input
x = self.stn(x)
# Perform the usual forward pass
x = F.relu(F.max_pool2d(self.conv1(x), 2))
x = F.relu(F.max_pool2d(self.conv2_drop(self.conv2(x)), 2))
x = x.view(-1, 320)
x = F.relu(self.fc1(x))
x = F.dropout(x, training=self.training)
x = self.fc2(x)
return F.log_softmax(x, dim=1)
model = Net().to(device)
```
## Train and Test functions for the STN
Train function
```
optimizer = optim.SGD(model.parameters(), lr=0.01)
def train(epoch):
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
data, target = data.to(device), target.to(device)
optimizer.zero_grad()
output = model(data)
loss = F.nll_loss(output, target)
loss.backward()
optimizer.step()
if batch_idx % 500 == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(train_loader.dataset),
100. * batch_idx / len(train_loader), loss.item()))
```
Test function
```
def test():
with torch.no_grad():
model.eval()
test_loss = 0
correct = 0
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
# sum up batch loss
test_loss += F.nll_loss(output, target, size_average=False).item()
# get the index of the max log-probability
pred = output.max(1, keepdim=True)[1]
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader.dataset)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'
.format(test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
```
## Run Train and test the data
```
for epoch in range(1, 20 + 1):
train(epoch)
test()
```
Visualize the results
```
def visualize_stn():
with torch.no_grad():
# Get a batch of training data
data = next(iter(test_loader))[0].to(device)
input_tensor = data.cpu()
transformed_input_tensor = model.stn(data).cpu()
in_grid = convert_image_np(
torchvision.utils.make_grid(input_tensor))
out_grid = convert_image_np(
torchvision.utils.make_grid(transformed_input_tensor))
# Plot the results side-by-side
f, axarr = plt.subplots(1, 2)
axarr[0].imshow(in_grid)
axarr[0].set_title('Dataset Images')
axarr[1].imshow(out_grid)
axarr[1].set_title('Transformed Images')
return in_grid, out_grid
# Visualize the STN transformation on some input batch
[in_grid, out_grid] = visualize_stn()
plt.ioff()
plt.show()
```
# Use Sørensen–Dice coefficient (DSC) to calculate the simularity of images --> modify for your data
Function to Compare similarity of images
Compare the similarity of two images with the the Sørensen–Dice coefficient. See details here: https://en.wikipedia.org/wiki/S%C3%B8rensen%E2%80%93Dice_coefficient
```
def calc_dice(im1,im2):
"""
This calculates the DICE between two images. The maximum DICE is 1, the minimum is Zero.
Args:
- im1, im2: one of the imges to calcualte DICE coeffeicnet. Note image1.shape has to equal image2.shape
Returns:
- dice: the dice coeffeicent
"""
im1 = np.asarray(im1).astype(np.bool)
im2 = np.asarray(im2).astype(np.bool)
if im1.shape != im2.shape:
raise ValueError("Shape mismatch: im1 and im2 must have the same shape.")
# Compute Dice coefficient
intersection = np.logical_and(im1, im2)
dice = 2. * intersection.sum() / (im1.sum() + im2.sum())
print("The Dice is {}".format(dice))
return dice
```
run DICE on INDIVUAL images set
```
#what do you need to do to calculate dice for your images?
dice = calc_dice(yourdata_im1,yourdata_im2)
```
# Optional preprocessing steps --> modify for your data
## Add salt/pepper noise to the dataset
add noise class
```
#are there other types of noise you can add? What effect do different types of noise have? EX: gasusain?
def salt_pepper_noise(trainloader):
for data in trainloader:
img, _ = data[0], data[1]
s_and_p = torch.tensor(random_noise(img, mode='s&p', salt_vs_pepper=0.5, clip=True))
return s_and_p
```
add noise to both train and test datasets
plot clean and noisy data
## remove nosie with noise2void
Check out: https://aswali.github.io/WNet/ and this paper:https://arxiv.org/abs/1711.08506
import noise2void dependances
```
from n2v.models import N2VConfig, N2V
from csbdeep.utils import plot_history
from n2v.utils.n2v_utils import manipulate_val_data
from n2v.internals.N2V_DataGenerator import N2V_DataGenerator
```
make denoising fucntion
```
def noise2void(data, model_name, patch_size=64):
"""
Removes noise in 3d image using the noise 2 void method. Based on https://arxiv.org/abs/1811.10980 w/ this implementation: https://github.com/juglab/n2v
Args:
- data: Numpy array 2d to be deionised
- model: name of model to load, if provided this model is used to denoise instead of model made from data otherwise this is name given to model (ex:#model_name = 'n2v_3D_blk')
-patch_size: this is the size of patches in X and Y, default is 64
Returns:
- data_denoise: Numpy array noise removed
"""
# We create our DataGenerator-object.
datagen = N2V_DataGenerator()
# In the 'dims' parameter we specify the order of dimensions in the image files we are reading.
if data.ndim == 2:
print('2D image found to denosie')
dataZYX = data
data_exp = np.expand_dims(dataZYX, axis=(0, 1, 4)) # expand dimensions One at the front is used to hold a potential stack of images such as a movie, One at the end could hold color channels such as RGB. #expand dimensions One at the front is used to hold a potential stack of images such as a movie, One at the end could hold color channels such as RGB.
patch_shape = (patch_size, patch_size)
model_axis = 'YX'
print('arrary with extra dimensions is size of {}'.format(data_exp.shape))
print('patches are {}'.format(patch_shape))
# the base directory in which our model will live
basedir = 'models'
path = basedir + '/' + model_name
if not os.path.exists(path):
print(path)
# create model
patches = datagen.generate_patches_from_list(data_exp, shape=patch_shape)
print('patches shape {}'.format(patches.shape))
# Patches are created so they do not overlap.
# (Note: this is not the case if you specify a number of patches. See the docstring for details!)
# Non-overlapping patches enable us to split them into a training and validation set.
# modify split so set as a %
perc_95 = int(patches.shape[0] * 0.95)
X = patches[:perc_95] # this is 600/640
X_val = patches[perc_95:] # this is 40/640
# train model
# You can increase "train_steps_per_epoch" to get even better results at the price of longer computation.
fast = 128 # default
slow = 50 # to get better results? --> apply same model to Z plane
speed = fast
config = N2VConfig(X, unet_kern_size=3, train_steps_per_epoch=int(X.shape[0] / speed), train_epochs=20,
train_loss='mse', batch_norm=True, train_batch_size=4, n2v_perc_pix=0.198,
n2v_patch_shape=patch_shape, n2v_manipulator='uniform_withCP', n2v_neighborhood_radius=5)
# Let's look at the parameters stored in the config-object.
vars(config)
# We are now creating our network model.
model = N2V(config=config, name=model_name, basedir=basedir)
history = model.train(X, X_val)
print(sorted(list(history.history.keys())))
model.export_TF(name='Noise2Void - data',
description='This is the 3D Noise2Void for data.',
authors=["Kaleb Vinehout"],
test_img=X_val[0, ..., 0], axes=model_axis,
patch_shape=patch_shape)
# run prediction model on rest of data in 3D image
# A previously trained model is loaded by creating a new N2V-object without providing a 'config'.
model = N2V(config=None, name=model_name, basedir=basedir)
# Here we process the data.
# The 'n_tiles' parameter can be used if images are too big for the GPU memory.
# If we do not provide the 'n_tiles' parameter the system will automatically try to find an appropriate tiling.
data_denoise = model.predict(dataZYX, axes=model_axis) # , n_tiles=(2, 4, 4))
return data_denoise
```
run the denoise on our data
```
data_denoise = noise2void(data, model_name='test_model', patch_size=64)
```
Make figures of raw data and denoised data
```
plt.figure()
plt.imshow(data, cmap='magma', vmin=np.percentile(data, 0.1), vmax=np.percentile(data, 99.9))
plt.figure()
plt.imshow(data_denoise, cmap='magma', vmin=np.percentile(data_denoise, 0.1), vmax=np.percentile(data_denoise, 99.9))
```
## Segment image with W-net
Wnet Class
```
# Wnet
class Block(nn.Module):
def __init__(self, in_filters, out_filters, seperable=True):
super(Block, self).__init__()
if seperable:
self.spatial1=nn.Conv2d(in_filters, in_filters, kernel_size=3, groups=in_filters, padding=1)
self.depth1=nn.Conv2d(in_filters, out_filters, kernel_size=1)
self.conv1=lambda x: self.depth1(self.spatial1(x))
self.spatial2=nn.Conv2d(out_filters, out_filters, kernel_size=3, padding=1, groups=out_filters)
self.depth2=nn.Conv2d(out_filters, out_filters, kernel_size=1)
self.conv2=lambda x: self.depth2(self.spatial2(x))
else:
self.conv1=nn.Conv2d(in_filters, out_filters, kernel_size=3, padding=1)
self.conv2=nn.Conv2d(out_filters, out_filters, kernel_size=3, padding=1)
self.relu1 = nn.ReLU()
self.dropout1 = nn.Dropout(0.65)
self.batchnorm1=nn.BatchNorm2d(out_filters)
self.relu2 = nn.ReLU()
self.dropout2 = nn.Dropout(0.65)
self.batchnorm2=nn.BatchNorm2d(out_filters)
def forward(self, x):
x = self.batchnorm1(self.conv1(x)).clamp(0)
x = self.relu1(x)
x = self.dropout1(x)
x = self.batchnorm2(self.conv2(x)).clamp(0)
x = self.relu2(x)
x = self.dropout2(x)
return x
class UEnc(nn.Module):
def __init__(self, squeeze, ch_mul=64, in_chans=3):
super(UEnc, self).__init__()
self.enc1=Block(in_chans, ch_mul, seperable=False)
self.enc2=Block(ch_mul, 2*ch_mul)
self.enc3=Block(2*ch_mul, 4*ch_mul)
self.enc4=Block(4*ch_mul, 8*ch_mul)
self.middle=Block(8*ch_mul, 16*ch_mul)
self.up1=nn.ConvTranspose2d(16*ch_mul, 8*ch_mul, kernel_size=3, stride=2, padding=1, output_padding=1)
self.dec1=Block(16*ch_mul, 8*ch_mul)
self.up2=nn.ConvTranspose2d(8*ch_mul, 4*ch_mul, kernel_size=3, stride=2, padding=1, output_padding=1)
self.dec2=Block(8*ch_mul, 4*ch_mul)
self.up3=nn.ConvTranspose2d(4*ch_mul, 2*ch_mul, kernel_size=3, stride=2, padding=1, output_padding=1)
self.dec3=Block(4*ch_mul, 2*ch_mul)
self.up4=nn.ConvTranspose2d(2*ch_mul, ch_mul, kernel_size=3, stride=2, padding=1, output_padding=1)
self.dec4=Block(2*ch_mul, ch_mul, seperable=False)
self.final=nn.Conv2d(ch_mul, squeeze, kernel_size=(1, 1))
self.softmax = nn.Softmax2d()
def forward(self, x):
enc1=self.enc1(x)
enc2=self.enc2(F.max_pool2d(enc1, (2, 2)))
enc3=self.enc3(F.max_pool2d(enc2, (2,2)))
enc4=self.enc4(F.max_pool2d(enc3, (2,2)))
middle=self.middle(F.max_pool2d(enc4, (2,2)))
up1=torch.cat([enc4, self.up1(middle)], 1)
dec1=self.dec1(up1)
up2=torch.cat([enc3, self.up2(dec1)], 1)
dec2=self.dec2(up2)
up3=torch.cat([enc2, self.up3(dec2)], 1)
dec3=self.dec3(up3)
up4=torch.cat([enc1, self.up4(dec3)], 1)
dec4=self.dec4(up4)
final=self.final(dec4)
return final
class UDec(nn.Module):
def __init__(self, squeeze, ch_mul=64, in_chans=3):
super(UDec, self).__init__()
self.enc1=Block(squeeze, ch_mul, seperable=False)
self.enc2=Block(ch_mul, 2*ch_mul)
self.enc3=Block(2*ch_mul, 4*ch_mul)
self.enc4=Block(4*ch_mul, 8*ch_mul)
self.middle=Block(8*ch_mul, 16*ch_mul)
self.up1=nn.ConvTranspose2d(16*ch_mul, 8*ch_mul, kernel_size=3, stride=2, padding=1, output_padding=1)
self.dec1=Block(16*ch_mul, 8*ch_mul)
self.up2=nn.ConvTranspose2d(8*ch_mul, 4*ch_mul, kernel_size=3, stride=2, padding=1, output_padding=1)
self.dec2=Block(8*ch_mul, 4*ch_mul)
self.up3=nn.ConvTranspose2d(4*ch_mul, 2*ch_mul, kernel_size=3, stride=2, padding=1, output_padding=1)
self.dec3=Block(4*ch_mul, 2*ch_mul)
self.up4=nn.ConvTranspose2d(2*ch_mul, ch_mul, kernel_size=3, stride=2, padding=1, output_padding=1)
self.dec4=Block(2*ch_mul, ch_mul, seperable=False)
self.final=nn.Conv2d(ch_mul, in_chans, kernel_size=(1, 1))
def forward(self, x):
enc1 = self.enc1(x)
enc2 = self.enc2(F.max_pool2d(enc1, (2, 2)))
enc3 = self.enc3(F.max_pool2d(enc2, (2,2)))
enc4 = self.enc4(F.max_pool2d(enc3, (2,2)))
middle = self.middle(F.max_pool2d(enc4, (2,2)))
up1 = torch.cat([enc4, self.up1(middle)], 1)
dec1 = self.dec1(up1)
up2 = torch.cat([enc3, self.up2(dec1)], 1)
dec2 = self.dec2(up2)
up3 = torch.cat([enc2, self.up3(dec2)], 1)
dec3 =self.dec3(up3)
up4 = torch.cat([enc1, self.up4(dec3)], 1)
dec4 = self.dec4(up4)
final=self.final(dec4)
return final
class WNet(nn.Module):
def __init__(self, squeeze, ch_mul=64, in_chans=3, out_chans=1000):
super(WNet, self).__init__()
if out_chans==1000:
out_chans=in_chans
self.UEnc=UEnc(squeeze, ch_mul, in_chans)
self.UDec=UDec(squeeze, ch_mul, out_chans)
def forward(self, x, returns='both'):
enc = self.UEnc(x)
if returns=='enc':
return enc
dec=self.UDec(F.softmax(enc, 1))
if returns=='dec':
return dec
if returns=='both':
return enc, dec
else:
raise ValueError('Invalid returns, returns must be in [enc dec both]')
```
Wnet train/test/loss
```
softmax = nn.Softmax2d()
criterionIdt = torch.nn.MSELoss()
def train_op(model, optimizer, input, k, img_size, psi=0.5):
enc = model(input, returns='enc')
d = enc.clone().detach()
n_cut_loss=soft_n_cut_loss(input, softmax(enc), img_size)
n_cut_loss.backward()
optimizer.step()
optimizer.zero_grad()
dec = model(input, returns='dec')
rec_loss=reconstruction_loss(input, dec)
rec_loss.backward()
optimizer.step()
optimizer.zero_grad()
return (model, n_cut_loss, rec_loss)
def reconstruction_loss(x, x_prime):
rec_loss = criterionIdt(x_prime, x)
return rec_loss
def test():
wnet=WNet.WNet(4)
synthetic_data=torch.rand((1, 3, 128, 128))
optimizer=torch.optim.SGD(wnet.parameters(), 0.001) #.cuda()
train_op(wnet, optimizer, synthetic_data)
def show_image(image):
img = image.numpy().transpose((1, 2, 0))
plt.imshow(img)
plt.show()
```
WNET MAIN
```
# Check if CUDA is available
CUDA = torch.cuda.is_available()
epochsall=100
# Create empty lists for average N_cut losses and reconstruction losses
n_cut_losses_avg = []
rec_losses_avg = []
# Squeeze k
k = 4
img_size = #define image size from test_loader
wnet = WNet(k)
learning_rate = 0.003
optimizer = torch.optim.SGD(wnet.parameters(), lr=learning_rate)
transform = transforms.Compose([transforms.Resize(img_size),
transforms.ToTensor()])
# Train 1 image set batch size=1 and set shuffle to False
#Here we can train new model for each image, or batch it
dataloader = train_loader # or test_loader
# Run for every epoch
for epoch in range(epochsall):
# At 1000 epochs divide SGD learning rate by 10
if (epoch > 0 and epoch % 1000 == 0):
learning_rate = learning_rate/10
optimizer = torch.optim.SGD(wnet.parameters(), lr=learning_rate)
# Print out every epoch:
print("Epoch = " + str(epoch))
# Create empty lists for N_cut losses and reconstruction losses
n_cut_losses = []
rec_losses = []
start_time = time.time()
for (idx, batch) in enumerate(dataloader):
# Train 1 image idx > 1
if(idx > 1): break
# Train Wnet with CUDA if available
if CUDA:
batch[0] = batch[0].cuda()
wnet, n_cut_loss, rec_loss = train_op(wnet, optimizer, batch[0], k, img_size)
n_cut_losses.append(n_cut_loss.detach())
rec_losses.append(rec_loss.detach())
n_cut_losses_avg.append(torch.mean(torch.FloatTensor(n_cut_losses)))
rec_losses_avg.append(torch.mean(torch.FloatTensor(rec_losses)))
print("--- %s seconds ---" % (time.time() - start_time))
images, labels = next(iter(dataloader))
# Run wnet with cuda if enabled
if CUDA:
images = images.cuda()
enc, dec = wnet(images)
torch.save(wnet.state_dict(), "model_" + args.name)
wnet_model = wnet
np.save("n_cut_losses_" + args.name, n_cut_losses_avg)
np.save("rec_losses_" + args.name, rec_losses_avg)
print("Done")
```
Apply Wnet to set of images
```
model = WNet.WNet(4) #squezze layers
model.load_state_dict(torch.load(wnet_model))
model.eval()
transform = transforms.Compose([transforms.Resize((64, 64)),
transforms.ToTensor()])
image = Image.open(args.image).convert('RGB')
x = transform(image)[None, :, :, :]
enc, dec = model(x)
show_image(x[0])
show_image(enc[0, :1, :, :].detach())
show_image(dec[0, :, :, :].detach())
```
| github_jupyter |
###### Background
- As you know that, the non-zero value is very few in target in this competition. It is like imbalance of target in classfication problem.
- For solving the imbalance in classfication problem, we commonly use the "stratifed sampling".
- For this cometition, we can simply apply the stratified sampling to get more well-distributed sampling for continuous target.
- To compare the effect of this strategy, I forked the good kernel(https://www.kaggle.com/prashantkikani/rstudio-lgb-single-model-lb1-6607) and used same parameters, same random seeds.
- I just change the sampling strategy. Ok, Let's see whether it works.
```
import os
import json
import numpy as np
import pandas as pd
from pandas.io.json import json_normalize
import time
from datetime import datetime
import gc
import psutil
from sklearn.preprocessing import LabelEncoder
PATH="../input/"
NUM_ROUNDS = 20000
VERBOSE_EVAL = 500
STOP_ROUNDS = 100
N_SPLITS = 10
#the columns that will be parsed to extract the fields from the jsons
cols_to_parse = ['device', 'geoNetwork', 'totals', 'trafficSource']
def read_parse_dataframe(file_name):
#full path for the data file
path = PATH + file_name
#read the data file, convert the columns in the list of columns to parse using json loader,
#convert the `fullVisitorId` field as a string
data_df = pd.read_csv(path,
converters={column: json.loads for column in cols_to_parse},
dtype={'fullVisitorId': 'str'})
#parse the json-type columns
for col in cols_to_parse:
#each column became a dataset, with the columns the fields of the Json type object
json_col_df = json_normalize(data_df[col])
json_col_df.columns = [f"{col}_{sub_col}" for sub_col in json_col_df.columns]
#we drop the object column processed and we add the columns created from the json fields
data_df = data_df.drop(col, axis=1).merge(json_col_df, right_index=True, left_index=True)
return data_df
def process_date_time(data_df):
print("process date time ...")
data_df['date'] = data_df['date'].astype(str)
data_df["date"] = data_df["date"].apply(lambda x : x[:4] + "-" + x[4:6] + "-" + x[6:])
data_df["date"] = pd.to_datetime(data_df["date"])
data_df["year"] = data_df['date'].dt.year
data_df["month"] = data_df['date'].dt.month
data_df["day"] = data_df['date'].dt.day
data_df["weekday"] = data_df['date'].dt.weekday
data_df['weekofyear'] = data_df['date'].dt.weekofyear
data_df['month_unique_user_count'] = data_df.groupby('month')['fullVisitorId'].transform('nunique')
data_df['day_unique_user_count'] = data_df.groupby('day')['fullVisitorId'].transform('nunique')
data_df['weekday_unique_user_count'] = data_df.groupby('weekday')['fullVisitorId'].transform('nunique')
return data_df
def process_format(data_df):
print("process format ...")
for col in ['visitNumber', 'totals_hits', 'totals_pageviews']:
data_df[col] = data_df[col].astype(float)
data_df['trafficSource_adwordsClickInfo.isVideoAd'].fillna(True, inplace=True)
data_df['trafficSource_isTrueDirect'].fillna(False, inplace=True)
return data_df
def process_device(data_df):
print("process device ...")
data_df['browser_category'] = data_df['device_browser'] + '_' + data_df['device_deviceCategory']
data_df['browser_os'] = data_df['device_browser'] + '_' + data_df['device_operatingSystem']
return data_df
def process_totals(data_df):
print("process totals ...")
data_df['visitNumber'] = np.log1p(data_df['visitNumber'])
data_df['totals_hits'] = np.log1p(data_df['totals_hits'])
data_df['totals_pageviews'] = np.log1p(data_df['totals_pageviews'].fillna(0))
data_df['mean_hits_per_day'] = data_df.groupby(['day'])['totals_hits'].transform('mean')
data_df['sum_hits_per_day'] = data_df.groupby(['day'])['totals_hits'].transform('sum')
data_df['max_hits_per_day'] = data_df.groupby(['day'])['totals_hits'].transform('max')
data_df['min_hits_per_day'] = data_df.groupby(['day'])['totals_hits'].transform('min')
data_df['var_hits_per_day'] = data_df.groupby(['day'])['totals_hits'].transform('var')
data_df['mean_pageviews_per_day'] = data_df.groupby(['day'])['totals_pageviews'].transform('mean')
data_df['sum_pageviews_per_day'] = data_df.groupby(['day'])['totals_pageviews'].transform('sum')
data_df['max_pageviews_per_day'] = data_df.groupby(['day'])['totals_pageviews'].transform('max')
data_df['min_pageviews_per_day'] = data_df.groupby(['day'])['totals_pageviews'].transform('min')
return data_df
def process_geo_network(data_df):
print("process geo network ...")
data_df['sum_pageviews_per_network_domain'] = data_df.groupby('geoNetwork_networkDomain')['totals_pageviews'].transform('sum')
data_df['count_pageviews_per_network_domain'] = data_df.groupby('geoNetwork_networkDomain')['totals_pageviews'].transform('count')
data_df['mean_pageviews_per_network_domain'] = data_df.groupby('geoNetwork_networkDomain')['totals_pageviews'].transform('mean')
data_df['sum_hits_per_network_domain'] = data_df.groupby('geoNetwork_networkDomain')['totals_hits'].transform('sum')
data_df['count_hits_per_network_domain'] = data_df.groupby('geoNetwork_networkDomain')['totals_hits'].transform('count')
data_df['mean_hits_per_network_domain'] = data_df.groupby('geoNetwork_networkDomain')['totals_hits'].transform('mean')
return data_df
def process_traffic_source(data_df):
print("process traffic source ...")
data_df['source_country'] = data_df['trafficSource_source'] + '_' + data_df['geoNetwork_country']
data_df['campaign_medium'] = data_df['trafficSource_campaign'] + '_' + data_df['trafficSource_medium']
data_df['medium_hits_mean'] = data_df.groupby(['trafficSource_medium'])['totals_hits'].transform('mean')
data_df['medium_hits_max'] = data_df.groupby(['trafficSource_medium'])['totals_hits'].transform('max')
data_df['medium_hits_min'] = data_df.groupby(['trafficSource_medium'])['totals_hits'].transform('min')
data_df['medium_hits_sum'] = data_df.groupby(['trafficSource_medium'])['totals_hits'].transform('sum')
return data_df
#Feature processing
## Load data
print('reading train')
train_df = read_parse_dataframe('train.csv')
trn_len = train_df.shape[0]
train_df = process_date_time(train_df)
print('reading test')
test_df = read_parse_dataframe('test.csv')
test_df = process_date_time(test_df)
## Drop columns
cols_to_drop = [col for col in train_df.columns if train_df[col].nunique(dropna=False) == 1]
train_df.drop(cols_to_drop, axis=1, inplace=True)
test_df.drop([col for col in cols_to_drop if col in test_df.columns], axis=1, inplace=True)
###only one not null value
train_df.drop(['trafficSource_campaignCode'], axis=1, inplace=True)
###converting columns format
train_df['totals_transactionRevenue'] = train_df['totals_transactionRevenue'].astype(float)
train_df['totals_transactionRevenue'] = train_df['totals_transactionRevenue'].fillna(0)
# train_df['totals_transactionRevenue'] = np.log1p(train_df['totals_transactionRevenue'])
## Features engineering
train_df = process_format(train_df)
train_df = process_device(train_df)
train_df = process_totals(train_df)
train_df = process_geo_network(train_df)
train_df = process_traffic_source(train_df)
test_df = process_format(test_df)
test_df = process_device(test_df)
test_df = process_totals(test_df)
test_df = process_geo_network(test_df)
test_df = process_traffic_source(test_df)
## Categorical columns
print("process categorical columns ...")
num_cols = ['month_unique_user_count', 'day_unique_user_count', 'weekday_unique_user_count',
'visitNumber', 'totals_hits', 'totals_pageviews',
'mean_hits_per_day', 'sum_hits_per_day', 'min_hits_per_day', 'max_hits_per_day', 'var_hits_per_day',
'mean_pageviews_per_day', 'sum_pageviews_per_day', 'min_pageviews_per_day', 'max_pageviews_per_day',
'sum_pageviews_per_network_domain', 'count_pageviews_per_network_domain', 'mean_pageviews_per_network_domain',
'sum_hits_per_network_domain', 'count_hits_per_network_domain', 'mean_hits_per_network_domain',
'medium_hits_mean','medium_hits_min','medium_hits_max','medium_hits_sum']
not_used_cols = ["visitNumber", "date", "fullVisitorId", "sessionId",
"visitId", "visitStartTime", 'totals_transactionRevenue', 'trafficSource_referralPath']
cat_cols = [col for col in train_df.columns if col not in num_cols and col not in not_used_cols]
merged_df = pd.concat([train_df, test_df])
print('Cat columns : ', len(cat_cols))
ohe_cols = []
for i in cat_cols:
if len(set(merged_df[i].values)) < 100:
ohe_cols.append(i)
print('ohe_cols : ', ohe_cols)
print(len(ohe_cols))
merged_df = pd.get_dummies(merged_df, columns = ohe_cols)
train_df = merged_df[:trn_len]
test_df = merged_df[trn_len:]
del merged_df
gc.collect()
for col in cat_cols:
if col in ohe_cols:
continue
#print(col)
lbl = LabelEncoder()
lbl.fit(list(train_df[col].values.astype('str')) + list(test_df[col].values.astype('str')))
train_df[col] = lbl.transform(list(train_df[col].values.astype('str')))
test_df[col] = lbl.transform(list(test_df[col].values.astype('str')))
print('FINAL train shape : ', train_df.shape, ' test shape : ', test_df.shape)
#print(train_df.columns)
train_df = train_df.sort_values('date')
X = train_df.drop(not_used_cols, axis=1)
y = train_df['totals_transactionRevenue']
X_test = test_df.drop([col for col in not_used_cols if col in test_df.columns], axis=1)
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import KFold, StratifiedKFold
from sklearn import model_selection, preprocessing, metrics
import matplotlib.pyplot as plt
import seaborn as sns
lgb_params1 = {"objective" : "regression", "metric" : "rmse",
"max_depth": 8, "min_child_samples": 20,
"reg_alpha": 1, "reg_lambda": 1,
"num_leaves" : 257, "learning_rate" : 0.01,
"subsample" : 0.8, "colsample_bytree" : 0.8,
"verbosity": -1}
```
# Stratified sampling
- Before stratified samling, we need to pseudo-label for continous target.
- In this case, I categorize the continous target into 12 class using range of 2.
```
# def categorize_target(x):
# if x < 2:
# return 0
# elif x < 4:
# return 1
# elif x < 6:
# return 2
# elif x < 8:
# return 3
# elif x < 10:
# return 4
# elif x < 12:
# return 5
# elif x < 14:
# return 6
# elif x < 16:
# return 7
# elif x < 18:
# return 8
# elif x < 20:
# return 9
# elif x < 22:
# return 10
# else:
# return 11
from sklearn.model_selection import StratifiedKFold
import lightgbm as lgb
```
## Target, prediction process
- 1st log1p to target
- 2nd exmp1 predictions
- 3rd sum predictions
- 4th log1p to sum
```
# y_categorized = y.apply(categorize_target)
y_log = np.log1p(y)
y_categorized= pd.cut(y_log, bins=range(0,25,3), include_lowest=True,right=False, labels=range(0,24,3)) # Thanks to Vitaly Portnoy
FOLDs = StratifiedKFold(n_splits=5, shuffle=True, random_state=7)
oof_lgb = np.zeros(len(train_df))
predictions_lgb = np.zeros(len(test_df))
features_lgb = list(X.columns)
feature_importance_df_lgb = pd.DataFrame()
for fold_, (trn_idx, val_idx) in enumerate(FOLDs.split(X, y_categorized)):
trn_data = lgb.Dataset(X.iloc[trn_idx], label=y_log.iloc[trn_idx])
val_data = lgb.Dataset(X.iloc[val_idx], label=y_log.iloc[val_idx])
print("LGB " + str(fold_) + "-" * 50)
num_round = 20000
clf = lgb.train(lgb_params1, trn_data, num_round, valid_sets = [trn_data, val_data], verbose_eval=1000, early_stopping_rounds = 100)
oof_lgb[val_idx] = clf.predict(X.iloc[val_idx], num_iteration=clf.best_iteration)
fold_importance_df_lgb = pd.DataFrame()
fold_importance_df_lgb["feature"] = features_lgb
fold_importance_df_lgb["importance"] = clf.feature_importance()
fold_importance_df_lgb["fold"] = fold_ + 1
feature_importance_df_lgb = pd.concat([feature_importance_df_lgb, fold_importance_df_lgb], axis=0)
predictions_lgb += clf.predict(X_test, num_iteration=clf.best_iteration) / FOLDs.n_splits
#lgb.plot_importance(clf, max_num_features=30)
cols = feature_importance_df_lgb[["feature", "importance"]].groupby("feature").mean().sort_values(by="importance", ascending=False)[:50].index
best_features_lgb = feature_importance_df_lgb.loc[feature_importance_df_lgb.feature.isin(cols)]
plt.figure(figsize=(14,10))
sns.barplot(x="importance", y="feature", data=best_features_lgb.sort_values(by="importance", ascending=False))
plt.title('LightGBM Features (avg over folds)')
plt.tight_layout()
plt.savefig('lgbm_importances.png')
x = []
for i in oof_lgb:
if i < 0:
x.append(0.0)
else:
x.append(i)
cv_lgb = mean_squared_error(x, y_log)**0.5
cv_lgb = str(cv_lgb)
cv_lgb = cv_lgb[:10]
pd.DataFrame({'preds': x}).to_csv('lgb_oof_' + cv_lgb + '.csv', index = False)
print("CV_LGB : ", cv_lgb)
sub_df = test_df[['fullVisitorId']].copy()
predictions_lgb[predictions_lgb<0] = 0
sub_df["PredictedLogRevenue"] = np.expm1(predictions_lgb)
sub_df = sub_df.groupby("fullVisitorId")["PredictedLogRevenue"].sum().reset_index()
sub_df.columns = ["fullVisitorId", "PredictedLogRevenue"]
sub_df["PredictedLogRevenue"] = np.log1p(sub_df["PredictedLogRevenue"])
sub_df.to_csv("submission.csv", index=False)
```
* - My result is LB : 1.4627
# Conclusion
- The improvement seems to be small, but you know that the small result can change the medal winner.
- This strategy would be improved using more category, etc.
- How about using it?
| github_jupyter |
# FIDO - the unified downloader tool for SunPy
### NOTE: Internet access is required in order to use this tutorial
FIDO is a new feature as part of the 0.8 SunPy release. It provides a unified interface to search for and download data from multiple sources and clients. For example, it can be used to search for images via the VSO, or timeseries data via an instrument-specific client, or both simultaneously.
In this tutorial, we will show some examples of how FIDO may be used to both search for and download data of various types.
### Importing modules - getting started
Python is modular, so first we have to import some modules we will need.
```
import sunpy # import sunpy itself
from sunpy.net import Fido #import the Fido module from sunpy
from sunpy.net import attrs as a #these are the attributes that are used to construct searches with the FIDO client
import matplotlib.pyplot as plt # for plotting
import astropy.units as u # Much of SunPy uses astropy units and quantities
```
### Prelude - a quick note on AstroPy Quantities
Some Fido search queries require the use of AstroPy quantities. We are not going to cover these in detail here, but a brief explanation is needed.
In short, an Astropy Quantity is just a value attached so some units or dimensions. In the first cell, you will see that we already imported astropy.units and gave it the name u.
#### Simple quantity example
Here is a very quick example showing how quantities can be used. Here we construct a velocity in km/s and a distance in km. When we calculate the time to travel the distance at the given velocity, the result is unit aware. More details of Units and Quantities can be found in other notebooks.
```
velocity = 100*u.km/u.s
distance = 150000*u.km
time_to_travel = distance/velocity
print(time_to_travel)
print(time_to_travel.unit)
```
### Example 1 - A simple search for AIA data
First, let's construct a simple search for AIA images using the Fido client. To do this, we have to construct a search query using valid *attributes*. Above, we imported the attributes module and gave it the short name *a*.
Let's search for all AIA files between 06:20 and 06:30 on 2011 June 7, during which there was an M-class solar flare.
```
example1_search = Fido.search(a.Time('2011-06-07 06:20','2011-06-07 06:30'), a.Instrument('AIA'))
print(example1_search)
```
From this summary of the search we can understand a few things. First, the search returned 402 files. We can see the properties of these files, such as the wavelength and time interval. Secondly, we can see that these entries were sourced from the Virtual Solar Observatory (VSO) client.
Let's refine this search to return only files with the 171A filter.
```
example1_search = Fido.search(a.Time('2011-06-07 06:20','2011-06-07 06:30'),
a.Instrument('AIA'), a.Wavelength(171*u.angstrom))
print(example1_search)
```
Now we can see that only 51 results were returned, all for files with the 171A filter. Notice also that we had to specify the wavelength using the astropy unit u.angstroms). Many functions in SunPy use these Units, which remove ambiguity in functions. *HINT*: see what happens if you try to carry out the search using just '171' as the wavelength.
#### Example 1 - downloading the result
The Fido module allows us to easily download the search results using the Fido.fetch function.
Let's download just the first file in our search results:
```
Fido.fetch(example1_search[:,0], path='.')
```
Now check that the file was correctly downloaded
```
ls *.fits
```
### Example 2 - querying multiple instruments simultaneously
We often want to retrieve data from different instruments of missions simultaneously, for example we may want SDO/AIA and STEREO/EUV images together. FIDO allows us to easily construct such searches.
Let's search for AIA and STEREO/EUVI data for the same time interval as before, between 06:20 - 06:30 on 2011 June 7.
```
example2_search = Fido.search(a.Time('2011-06-07 06:20','2011-06-07 06:30'),
a.Instrument('AIA') | a.Instrument('EUVI'))
```
Using the '|' symbol we can construct queries with multiple options for a given attribute, in this case 'Instrument'.
```
print(example2_search)
```
Looking at these results, we can see that we have search result in two blocks: the first block contains all the SDO/AIA search results, and the second block contains the STEREO/EUVI search results. As before, there were 402 AIA files found, and 22 STEREO/EUVI files.
These blocks can be indexed and retrieved separately, for example:
```
print(example2_search[1])
```
...returns only the STEREO search results block.
As before, we can download these files using the Fido.fetch command. Let's download just the first result from the STEREO results block.
```
Fido.fetch(example2_search[1,0],path='.')
ls
```
### Example 3 - querying multiple clients for different data types simultaneously
In both examples above, Fido returned results exclusively from the VSO client. Crucially though, Fido supports the search for data files from multiple different clients simulaneously.
As an example, let's construct a query where we want to obtain the AIA data, STEREO/EUVI data, and the GOES/XRS time series data from the 2011 June 7 event simultaneously.
```
example3_search = Fido.search(a.Time('2011-06-07 06:20','2011-06-07 06:30'),
a.Instrument('AIA') | a.Instrument('EUVI') | a.Instrument('GOES'))
print(example3_search)
```
We can see that this time, we have three blocks of responses. The third block contains results from another client, the GOESClient, which is for GOES/XRS time series data. FIDO automatically communicates with all of the different SunPy clients as required to find the requested data.
```
example3_search[2]
```
Once again, we can download just the GOES/XRS timeseries data using Fido.fetch.
```
Fido.fetch(example3_search[2,:],path='.')
ls
```
### Summary
By the end of this notebook you should have a basic understanding of how to use FIDO, the unified downloader client in SunPy. Here are some key things to remember:
- The function Fido.search() allows you to search online for data files from a large suite of solar missions.
- The Fido.fetch() command allows you to download the reults of a search.
- Searches are constructed using *attributes*. Sometimes these attributes need to be in the form of an *AstroPy Quantity*.
- Complex queries can be constructed via combinations of attributes - multiple options for an attribute may be used simultaneously.
- Fido can search for different data types from different sources simultaneously, e.g. timeseries, images and spectra.
| github_jupyter |
```
import os
import sys
#sys.path.append('D:\ProgramData\Anaconda3\envs\KKeras\Lib\site-packages')
import csv
import cv2
import matplotlib.pyplot as plt
import numpy as np
import sklearn
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split
```
## 处理数据文件名称列表
```
print(os.path.abspath('__file__'))
lines = []
with open('./data/driving_log.csv') as csvfile: # 相对路径,用./
csvsheet = csv.reader(csvfile)
for line in csvsheet:
lines.append(line)
lines = lines[1:-1] # Remove the title line
print(lines[0]) # show the first effective data, the structure should be
# [center pic], [left pic], [right pic], [steering],[throttle],[brake],[speed]
```
## 数据集生成
```
images = []
images_rz = []
measurements = []
for line in lines:
source_path = line[0] # 看CSV,表里面第一项是地址
#print(source_path)
pic_name = source_path.split('/')[-1] #被‘/’分割后的最后一项
#print(pic_name)
current_path ='./data/IMG/'+pic_name
image = cv2.imread(current_path)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)# 因为cv2读取的是BGR因此要转换成RGB显示才正常,包括drive.py使用也是
# image_rz = cv2.resize(image,(160,160),cv2.INTER_AREA) #取中间有效部分然后缩放
# images_rz.append(image_rz)
images.append(image)
measurement = float(line[3])
measurements.append(measurement) # get all the steering angle
plt.imshow(images[1])
plt.title('Example of input image')
plt.show()
#plt.imshow(images_rz[1])
#plt.title('Example after Resize')
#plt.show()
X_train = np.array(images)
#X_train = np.array(images)
Y_train = np.array(measurements)
train_samples, validation_samples = train_test_split(lines,test_size =0.2)
```
## Generator分割数据
```
def generator(samples, batch_size=32):
num_samples = len(samples)
while 1:
shuffle(samples)
for offset in range(0,num_samples,batch_size):
batch_samples = samples[offset:offset+batch_size]
images =[]
angles =[]
for batch_sample in batch_samples:
for ii in range(2):
name ='./data/IMG/'+batch_sample[ii].split('/')[-1] # the name of the pic
center_image = cv2.imread(name)
center_image = cv2.cvtColor(center_image, cv2.COLOR_BGR2RGB)
center_angle = float(batch_sample[3])
images.append(center_image)
angles.append(center_angle)
X_train = np.array(images)
Y_train = np.array(angles)
yield sklearn.utils.shuffle(X_train, Y_train)
train_generator = generator(train_samples,batch_size = 32)
validation_generator = generator(validation_samples, batch_size = 32)
```
## 数据准备完毕,开始构建网络
```
from keras.models import Sequential
from keras.layers import Flatten, Dense, Lambda, Dropout
from keras.layers.convolutional import Convolution2D, Conv2D
from keras.layers.pooling import MaxPooling2D
from keras.layers import Cropping2D
#Build Model
model = Sequential()
#Pre-processing
model.add(Lambda(lambda x: x / 255 - 0.5, input_shape=(160,320,3)))
model.add(Cropping2D(cropping=((50, 20), (0, 0))))
# NIVIDIA End to End Learning for Self-Driving Cars, Figure(4)
# First Conv, 24*5*5
model.add(Conv2D(24, 5, strides=(2, 2), activation='relu'))
model.add(Dropout(0.7))
# Second Conv, 36*5*5
model.add(Conv2D(36, 5, strides=(2, 2), activation='relu'))
# Thrid Conv, 48*5*5
model.add(Conv2D(48, 5, strides=(2, 2), activation='relu'))
# Fourth Conv, 64*5*5
model.add(Conv2D(64, 3, activation='relu'))
model.add(Conv2D(64, 3, activation='relu'))
model.add(Flatten())
# Four Fully-Connected layers
model.add(Dense(100))
model.add(Dense(50))
model.add(Dense(10))
model.add(Dense(1))
#Run the Model
model.compile(loss = 'mse', optimizer = 'adam') #Regression问题,而不是Classification问
#history_object = model.fit(X_train, Y_train,validation_split = 0.2, shuffle = True, epochs = 5, verbose = 1)
history_object = model.fit_generator(train_generator,
steps_per_epoch = len(train_samples),
validation_data = validation_generator,
validation_steps = len(validation_samples),
epochs = 5)
#history_object = model.fit_generator(train_generator, steps_per_epoch = len(train_samples),validation_data = validation_generator,nb_val_samples = len(validation_samples), nb_epoch = 5)
print(history_object.history.keys())
### plot the training and validation loss for each epoch
plt.plot(history_object.history['loss'])
plt.plot(history_object.history['val_loss'])
plt.title('model mean squared error loss')
plt.ylabel('mean squared error loss')
plt.xlabel('epoch')
plt.legend(['training set', 'validation set'], loc='upper right')
plt.show()
model.save('model.h5')
```
| github_jupyter |
```
#an interestin link https://github.com/jasperdebie/VisInfo/blob/master/us-state-capitals.csv
#we want to create a hexbin of the food insecurity level
#for this task, we want a national coordinate system for the us namely USNG (united states national grid)
#we want to have the coordinates for the capital of each state to plot our hexbin correctly
import pandas as pd
df6 = pd.read_csv('/content/Book1.csv')
df6.head()
import plotly.express as px
fig = px.choropleth(df6, #the dataframe from which we will use the values
locations="Code", #the column with the USPS code
color="Percentage of the population who is food insecure in %", #the variable defining the range of colors
locationmode = 'USA-states') #a plot with all the us states
fig.update_layout(title_text = 'Level of food insecurity in each state', #the title of our plot
geo_scope='usa', #we only want the USA and not the whole world
)
fig.show() #to display the plot
#We used this link: https://wellsr.com/python/creating-python-choropleth-maps-with-plotly/
#We also used this: https://plotly.github.io/plotly.py-docs/generated/plotly.express.choropleth.html
from google.colab import files
files.upload()
import plotly.graph_objects as go
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/2011_us_ag_exports.csv')
df1 = pd.read_csv('data_project.csv')
fig = go.Figure(data=go.Choropleth(
locations=df['code'], # Spatial coordinates
z = df1['Percentage of the population who is food insecure in %'], # Data to be color-coded
locationmode = 'USA-states', # set of locations match entries in `locations`
colorscale = 'Reds',
colorbar_title = "Percentage",
))
fig.update_layout(
title_text = 'Percentage of the population which is food insecure in each state',
geo_scope='usa', # limite map scope to USA
)
fig.show()
#we want to create a timeline graph to display the evolution of food insecurity in the USA over time
#we will focus on the 21st century mostly
import matplotlib.pyplot as plt
import pandas as pd
from google.colab import files
files.upload()
timeline_df = pd.read_csv('FI percentages data.csv')
timeline_df
plt.figure(figsize=(10,5))
plt.plot(timeline_df['Year'], timeline_df['Percentage FI individuals'], color='c', marker='o')
plt.grid(True)
plt.title('Evolution of the food insecurity rate in the US between 1998 and 2019')
plt.xlabel('Year')
plt.ylabel('Percentage of individuals who are food insecure')
```
| github_jupyter |
## Day Agenda
- Reading the diffent format of data sets
- statistical information about data sets
- Concatination of data frames
- grouping of dataframes
- merging data frames
- filtering data from data frames
-
## Reading the diffent format of data sets
- csv(comma separated values)
- json
- xlsx
- tsv(tab separated values)
```
import pandas as pd
dir(pd)
# Loading the dataset from browser
df=pd.read_json("https://raw.githubusercontent.com/AP-State-Skill-Development-Corporation/Data-Science-Using-Python-Internship-EB1/main/Notebooks/Day23-Pandas%20continue/cafes.json")
df
pd.read_json("cafes.json")# loading dataset from local system
# convert json format into csv data set
df.to_csv("cafes.csv")
df1=pd.read_csv("https://raw.githubusercontent.com/AP-State-Skill-Development-Corporation/Data-Science-Using-Python-Internship-EB1/main/Notebooks/Day23-Pandas%20continue/chipotle.tsv.txt",
sep='\t')
df1.head()
df2=pd.read_csv("https://raw.githubusercontent.com/AP-State-Skill-Development-Corporation/Data-Science-Using-Python-Internship-EB1/main/Notebooks/Day23-Pandas%20continue/weather.csv")
df2.head()
```
## statistical information about data sets
- describe()
- info()
- corr()
- varies from +1(highely corrletaed) to -1(not correlated)
```
df1.head()
df1.describe()
df1.info()
df1.corr()
```
## Concatination of data frames
- pandas.concat(df1,df2)
```
d1=pd.DataFrame({'clg':['spmvv','Gist','aec','Alts'],
'temp':[35,47,68,33],
'hum':[45,23,78,89]})
d1
d2=pd.DataFrame({'clg':['vvit','kits','nec','Anu'],
'temp':[36,50,60,30],
'hum':[47,77,88,90]})
d2
import pandas as pd
d3=pd.concat((d1,d2))
d3
import pandas as pd
d3=pd.concat((d1,d2),ignore_index=True)
d3
import pandas as pd
d3=pd.concat((d1,d2),keys=['k1','k2'])
d3
```
## Grouping of dataframes
- df.groupby
```
help(df1.groupby)
d1
g1=d1.groupby(by='temp')# spliting a specific object
g1
d1=g1.max()# applying a function to groupby object and dispay a result
d1
d1
g2=d1.groupby(by='clg')
d1=g2
d1
```
## merging data frames
df.merge()
```
a=pd.DataFrame({'x':[11,22,33,44],'y':[33,44,35,16]})
b=pd.DataFrame({'x':[11,21,33,44],'y':[3,4,5,6]})
a
b
df1 = pd.DataFrame({'lkey': ['foo', 'bar', 'baz', 'foo'],
'value': [1, 2, 3, 5]})
df2 = pd.DataFrame({'rkey': ['foo', 'bar', 'baz', 'foo'],
'value': [5, 6, 7, 8]})
df1
df2
df3=pd.merge(df1,df2,indicator=True)
df3
df3=pd.merge(df1,df2,how='right',indicator=True)
df3
m1=pd.DataFrame({"city":['x','y','z'],
'temp':[11,22,33]})
m1
m2=pd.DataFrame({"city":['x','y','a'],
'hum':[44,55,66]})
m2
m3=pd.merge(m1,m2)
m3
m3=pd.merge(m1,m2,on='city',how='left',indicator=True)
m3
```
| github_jupyter |
# Potentiostats and Galvanostats
## References
---
Adams, Scott D., et al. "MiniStat: Development and evaluation of a mini-potentiostat for electrochemical measurements." Ieee Access 7 (2019): 31903-31912. https://ieeexplore.ieee.org/stamp/stamp.jsp?arnumber=8657694
---
Ainla, Alar, et al. "Open-source potentiostat for wireless electrochemical detection with smartphones." Analytical chemistry 90.10 (2018): 6240-6246. https://gmwgroup.harvard.edu/files/gmwgroup/files/1308.pdf
---
Bianchi, Valentina, et al. "A Wi-Fi cloud-based portable potentiostat for electrochemical biosensors." IEEE Transactions on Instrumentation and Measurement 69.6 (2019): 3232-3240.
---
Dobbelaere, Thomas, Philippe M. Vereecken, and Christophe Detavernier. "A USB-controlled potentiostat/galvanostat for thin-film battery characterization." HardwareX 2 (2017): 34-49. https://doi.org/10.1016/j.ohx.2017.08.001
---
Hoilett, Orlando S., et al. "KickStat: A coin-sized potentiostat for high-resolution electrochemical analysis." Sensors 20.8 (2020): 2407. https://www.mdpi.com/1424-8220/20/8/2407/htm
---
Irving, P., R. Cecil, and M. Z. Yates. "MYSTAT: A compact potentiostat/galvanostat for general electrochemistry measurements." HardwareX 9 (2021): e00163. https://www.sciencedirect.com/science/article/pii/S2468067220300729
> 2, 3, and 4 wire cell configurations with +/- 12 volts at 200ma.
---
Lopin, Prattana, and Kyle V. Lopin. "PSoC-Stat: A single chip open source potentiostat based on a Programmable System on a Chip." PloS one 13.7 (2018): e0201353. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0201353
---
Matsubara, Yasuo. "A Small yet Complete Framework for a Potentiostat, Galvanostat, and Electrochemical Impedance Spectrometer." (2021): 3362-3370. https://pubs.acs.org/doi/full/10.1021/acs.jchemed.1c00228
> Elegant 2 omp amp current source for a galvanostat.
---
## Application to Electrical Impedence Spectroscopy
---
Wang, Shangshang, et al. "Electrochemical impedance spectroscopy." Nature Reviews Methods Primers 1.1 (2021): 1-21. https://www.nature.com/articles/s43586-021-00039-w.pdf
> Tutorial presentation of EIS, including instrumentation and data analysis.
---
Magar, Hend S., Rabeay YA Hassan, and Ashok Mulchandani. "Electrochemical Impedance Spectroscopy (EIS): Principles, Construction, and Biosensing Applications." Sensors 21.19 (2021): 6578. https://www.mdpi.com/1424-8220/21/19/6578/pdf
> Tutorial introduction with descriptions of application to solutions and reactions at surfaces.
---
Instruments, Gamry. "Basics of electrochemical impedance spectroscopy." G. Instruments, Complex impedance in Corrosion (2007): 1-30. https://www.c3-analysentechnik.eu/downloads/applikationsberichte/gamry/5657-Application-Note-EIS.pdf
> Tutorial introduction to EIS with extensive modeling discussion.
---
| github_jupyter |
# Class 9: Functions
## A painful analogy
What do you do when you wake up in the morning?
I don't know about you, but I **get ready.**
"Obviously," you say, a little too snidely for my liking. You're particular, very detail-oriented, and need more information out of me.
Fine, then. Since you're going to be nitpicky, I might be able to break it down a little bit more for you...
1. I get out of bed
2. I take a shower
3. I get dressed
4. **I eat breakfast**
Unfortunately that's not good enough for you. "But how do you eat breakfast?" Well, maybe I...
1. Get a bowl out of a cabinet
2. Get some cereal out of the pantry
3. Get some milk out of the fridge
4. Pour some cereal into a bowl
5. Pour some milk into the bowl
6. Sit down at the table and start eating
"Are you eating with a spoon?" you interrupt. "When did you get the spoon out? Was that after the milk, or before the bowl?"
It's annoying people like this that make us have **functions.**
> **FUN FACT:** The joke's on you, because **I don't even actually eat cereal.** Maybe I don't even get ready in the morning, either.
## What is a function?
Functions are chunks of code that do something. They're different than the code we've written so far because **they have names**.
Instead of detailing each and every step involved in eating breakfast, I just use "I eat breakfast" as a shorthand for many, many detailed steps. Functions are the same - they allow us to take complicated parts of code, give it a name, and type **`just_eat_breakfast()`** every morning instead of twenty-five lines of code.
## What are some examples of functions?
We've used a lot of functions in our time with Python. You remember our good buddy `len`? It's a **function** that gives back the length of whatever you send its way, e.g. `len("ghost")` is `5` and `len("cartography")` is 11.
```
len
```
**Almost everything useful is a function.** Python has [a ton of other built-in functions](https://docs.python.org/2/library/functions.html)!
Along with `len`, a couple you might have seen are:
* `abs(...)` takes a number and returns the absolute value of the number
* `int(...)` takes a string or float and returns it as an integer
* `round(...)` takes a float and returns a rounded version of it
* `sum(...)` takes a list and returns the sum of all of its elements
* `max(...)` takes a list and returns the largest of all of its selements
* `print(...)` takes whatever you want to give it and displays it on the screen
Functions can also come from packages and libraries. The `.get` part of `requests.get` is a function, too!
And here, to prove it to you?
```
max
print
import requests
requests.get
# And if we just wanted to use them, for some reason
n = -34
print(n, "in absolute value is", abs(n))
print("We can add after casting to int:", 55 + int("55"))
n = 4.4847
print(n, "can be rounded to", round(n))
print(n, "can also be rounded to 2 decimal points", round(n, 2))
numbers = [4, 22, 40, 54]
print("The total of the list is", sum(numbers))
```
**See? Functions make the world run.**
One useful role they play is **functions hide code that you wouldn't want to type a thousand times.** For example, you might have used `urlretrieve` from `urllib` to download files from around the internet. If you *didn't* use `urlretrieve` you'd have to type all of this:
```
def urlretrieve(url, filename=None, reporthook=None, data=None):
url_type, path = splittype(url)
with contextlib.closing(urlopen(url, data)) as fp:
headers = fp.info()
# Just return the local path and the "headers" for file://
# URLs. No sense in performing a copy unless requested.
if url_type == "file" and not filename:
return os.path.normpath(path), headers
# Handle temporary file setup.
if filename:
tfp = open(filename, 'wb')
else:
tfp = tempfile.NamedTemporaryFile(delete=False)
filename = tfp.name
_url_tempfiles.append(filename)
with tfp:
result = filename, headers
bs = 1024*8
size = -1
read = 0
blocknum = 0
if "content-length" in headers:
size = int(headers["Content-Length"])
if reporthook:
reporthook(blocknum, bs, size)
while True:
block = fp.read(bs)
if not block:
break
read += len(block)
tfp.write(block)
blocknum += 1
if reporthook:
reporthook(blocknum, bs, size)
if size >= 0 and read < size:
raise ContentTooShortError(
"retrieval incomplete: got only %i out of %i bytes"
% (read, size), result)
return result
```
Horrifying, right? Thank goodness for functions.
## Writing your own functions
I've always been kind of jealous of `len(...)` and its crowd. It seemed unfair that Python made a list of cool, important functions, and neither me nor you had any say in the matter. What if I want a function that turns all of the periods in a sentence into exclamation points, or prints out a word a hundred million times?
Well, turns out **that isn't a problem**. We can do that. Easily! *And we will*. If you can type `def` and use a colon, you can write a function.
A function that you write yourself looks like this:
```
# A function to multiply a number by two
```
It has a handful of parts:
1. **`def`** - tells Python "hey buddy, we're about to define a function! Get ready." And Python appropriately prepares itself.
2. **`double`** - is the **name** of the function, and it's how you'll refer to the function later on. For example, `len`'s function name is (obviously) `len`.
3. **`(number)`** - defines the **parameters** that the function "takes." You can see that this function is called `double`, and you send it one parameter that will be called `number`.
4. **`return bigger`** - is called the **return statement**. If the function is a factory, this is the shipping department - **return** tells you what to send back to the main program.
You'll see it doesn't *do* anything, though. That's because we haven't **called** the function, which is a programmer's way of saying **use** the function. Let's use it!
```
print("2 times two is", double(2))
print("10 times two is", double(10))
print("56 times two is", double(56))
age = 76
print("Double your age is", double(age))
```
## Function Naming
Your function name has to be **unique**, otherwise Python will get confused. No other functions or variabels can share its name!
For example, if you call it `len` it'll forget about the built-in `len` function, and if you give one of your variables the name `print` suddenly Python won't understand how `print(...)` works anymore.
If you end up doing this, you'll get errors like the one below
```
def greet(name):
return "Hello " + name
# This one works
print(greet("Soma"))
# Overwrite the function greet with a string
greet = "blah"
# Trying the function again breaks
print(greet("Soma"))
```
## Parameters
In our function `double`, we have a parameter called `number`.
````py
def double(number):
bigger = number * 2
return bigger
````
Notice in the last example up above, though, we called `double(age)`. Those don't match!!!
The thing is, **your function doesn't care what the variable you send it is called**. Whatever you send it, it will rename. It's like if someone adopted my cat *Smushface*, they might think calling her *Petunia* would be a little bit nicer (it wouldn't be, but I wouldn't do anything about it).
Here's an example with my favorite variable name `potato_soup`
```
def exclaim(potato_soup):
return potato_soup + "!!!!!!!!!!"
invitation = "I hope you can come to my wedding"
print(exclaim(invitation))
line = "I am sorry to hear you have the flu"
print(exclaim(line))
```
`invitation` and `line` both get renamed to `potato_soup` inside of the function, so you can reuse the function with **any** variable of **any** name.
Let's say I have a function that does some **intense calculations**:
````py
def sum_times_two(a, b):
added = a + b
return added * 2
````
To reiterate: **`a` and `b` have nothing to do with the values outside of the function**. You don't have to make variables called `a` and `b` and then send them to the function, the function takes care of that by itself. For example, the below examples are perfectly fine.
````py
sum_times_two(2, 3)
r = 4
y = 7
sum_times_two(r, y)
````
When you're outside of the function, you almost **never have to think about what's inside the function.** You don't care about what variabels are called or *anything*. It's a magic box. Think about how you don't know what `len` looks like inside, or `print`, but you use them all of the time!
## Why functions?
Two reasons to use functions, since maybe you'll ask:
**Don't Repeat Yourself** - If you find yourself writing the same code again and again, it's a good time to put that code into a function. `len(...)` is a function because Python people decided that you shouldn't have to write length-calculating code every time you wanted to see how many characters were in a string.
**Code Modularity** - sometimes it's just nice to *organize* your code. All of your parts that deal with counting dog names can go over here, and all of the stuff that has to do with boroughs goes over there. In the end it can make for more readable and maintanable code. (Maintainable code = code you can edit in the future without thinking real hard)
Those reasons probably don't mean much to you right now, and I sure don't blame you. Abstract programming concepts are just dumb abstract things until you actually start using them.
Let's say I wanted to greet someone and then tell them how long their name is, because I'm pedantic.
```
name = "Nancy"
name_length = len(name)
print("Hello", name, "your name is", name_length, "letters long")
name = "Brick"
name_length = len(name)
print("Hello", name, "your name is", name_length, "letters long")
name = "Saint Augustine"
name_length = len(name)
print("Hello", name, "your name is", name_length, "letters long")
```
**Do you know how exhausted I got typing all of that out?** And how it makes no sense at all? Luckily, functions save us: all of our code goes into one place so we don't have to repeat ourselves, *and* we can give it a descriptive name.
```
def weird_greeting(name):
name_length = len(name)
print("Hello", name, "your name is", name_length, "letters long")
weird_greeting("Nancy")
weird_greeting("Brick")
weird_greeting("Saint Augustine")
```
# `return`
The role of a function is generally **to do something and then send the result back to us**. `len` sends us back the length of the string, `requests.get` sends us back the web page we requested.
````py
def double(a):
return a * 2
````
**This is called the `return` statement.** You don't *have* to send something back (`print` doesn't) but you usually want to.
# Writing a custom function
Let's say we have some code that compares the number of boats you have to the number of cars you have.
````python
if boat_count > car_count:
print "Larger"
else:
print "Smaller"
````
Simple, right? But unfortunately we're at a rich people convention where they're always comparing the number of boats to the number of cars to the number of planes etc etc etc. If we have to check *again and again and again and again* for all of those people and always print *Larger* or *Smaller* I'm sure we'd get bored of typing all that. So let's convert it to a function!
Let's give our function a **name** of `size_comparison`. Remember: We can name our functions whatever we want, *as long as it's unique*.
Our function will take **two parameters**. they're `boat_coat` and `car_count` above, but we want generic, re-usable names, so maybe like, uh, `a` and `b`?
For our function's **return value**, let's have it send back `"Larger"` or `"Smaller"`.
```
# Our cool function
def size_comparison(a, b):
if a > b:
return "Larger"
else:
return "Smaller"
print(size_comparison(4, 5.5))
print(size_comparison(65, 2))
print(size_comparison(34.2, 33))
```
# Your Turn
This is a do-now even though it's not the beginning of class!
### 1a. Driving Speed
With the code below, it tells you how fast you're driving. I figure that a lot of people are more familiar with kilometers an hour, though, so let's write a function that does the conversion. I wrote a skeleton, now you can fill in the conversion.
Make it display a whole number.
```
def to_kmh(speeed):
"YOUR CODE HERE"
mph = 40
print("You are driving", mph, "in mph")
print("You are driving", to_kmh(mph), "in kmh")
```
### 1b. Driving Speed Part II
Now write a function called `to_mpm` that, when given miles per hour, computes the meters per minute.
### 1c. Driving Speed Part III
Rewrite `to_mpm` to use the `to_kmh` function. **D.R.Y.**!
### 2. Broken Function
The code below won't work. Why not?
```
# You have to wash ten cars on every street, along with the cars in your driveway.
# With the following list of streets, how many cars do we have?
def total(n):
return n * 10
# Here are the streets
streets = ['10th Ave', '11th Street', '45th Ave']
# Let's count them up
total = len(streets)
# And add one
count = total + 1
# And see how many we have
print(total(count))
```
### 3. Data converter
We have a bunch of data in different formats, and we need to normalize it! The data looks like this:
````python
var first = { 'measurement': 3.4, 'scale': 'kilometer' }
var second = { 'measurement': 9.1, 'scale': 'mile' }
var third = { 'measurement': 2.0, 'scale': 'meter' }
var fourth = { 'measurement': 9.0, 'scale': 'inches' }
````
Write a function called `to_meters(...)`. When you send it a dictionary, have it examine the `measurement` and `scale` and return the adjusted value. For the values above, 3.4 kilometers should be 3400.0 meters, 9.1 miles should be around 14600, and 9 inches should be apprxoimately 0.23.
| github_jupyter |
# <font color="blue">Failure Cases in Tesseract OCR</font>
In this notebook, we will see some instances where Tesseract does not work as expected and provide logical reasons for them. We will see some examples from scanned documents as well as natural camera images.
We will discuss how to improve the OCR output in the next notebook.
# <font color="blue">Install Tesseract</font>
```
!apt install libtesseract-dev tesseract-ocr > /dev/null
!pip install pytesseract > /dev/null
```
# <font color="blue">Import Libraries </font>
```
import pytesseract
import cv2
import glob
import matplotlib.pyplot as plt
%matplotlib inline
from IPython.display import Image
```
# <font color="blue">Failure Cases Categories </font>
Even though failure cannot be always attributed to one specific reasons, We have listed a few major reasons for failure of OCR using Tesseract and in general. They are:
1. **Cluttered Background** - The text might not be visibly clear or it might appear camouflaged with the background.
1. **Small Text** - The text might be too small to be detected.
1. **Rotation or Perspective Distortion** - The text might be rotated in the image or the image itself might be distorted.
# <font color="blue">Test Image 1: Image of a Book</font>
```
!wget https://www.dropbox.com/s/uwrdek4jjac4ysz/book2.jpg?dl=1 -O book2.jpg --quiet
```
### <font color="green">Downloaded Image</font>
<img src="https://www.dropbox.com/s/uwrdek4jjac4ysz/book2.jpg?dl=1" width=500>
### <font color="green">Output</font>
```
book2_text = pytesseract.image_to_string('book2.jpg')
print(book2_text)
```
### <font color="green">Observation </font>
We saw this example in the last notebook. The major reason is the relatively cluttered background and low contrast of the white text on orange background. Another issue with these kind of images is the variability of text size. **"Black"** is written in extra-large size whereas **"The impact of"** is normal.
# <font color="blue">Test Image 2: Driving License </font>
```
!wget https://www.dropbox.com/s/rdaha84n8jo3bmw/dl.jpg?dl=0 -O dl.jpg --quiet
```
### <font color="green">Downloaded Image</font>

### <font color="green">Output</font>
```
dl_text = pytesseract.image_to_string('dl.jpg')
print(dl_text)
Image("dl.jpg")
```
### <font color="green">Observation </font>
- It is unable to detect small text( Date of Expiry, Address, etc.)
- Same issue with cluttered background ( New York on Top Left )
# <font color="blue">Test Image 3: License Plate</font>
```
!wget "https://www.dropbox.com/s/xz24vxrp4uvvnri/license_plate.jpg?dl=0" -O lp1.jpg --quiet
```
### <font color="green">Downloaded Image</font>
<img src="https://www.dropbox.com/s/xz24vxrp4uvvnri/license_plate.jpg?dl=1" width=500>
### <font color="green">Output</font>
```
lp_text = pytesseract.image_to_string('lp1.jpg')
print(lp_text)
```
### <font color="green">Observation </font>
Even though the text is very clear to us, Tesseract finds it difficult to recognize. The major issue is with contrast.
# <font color="blue">Street Signs</font>
Street signs are one of the most difficult ones to recognize. Let us see some examples.
# <font color="blue">Test Image 4</font>
```
!wget https://www.dropbox.com/s/uwlnxiihqgni57o/streetsign1.jpg?dl=0 -O streetsign1.jpg --quiet
```
### <font color="green">Downloaded Image</font>
<img src="https://www.dropbox.com/s/uwlnxiihqgni57o/streetsign1.jpg?dl=1" width=500>
### <font color="green">Output</font>
```
ss1_text = pytesseract.image_to_string('streetsign1.jpg')
print(ss1_text)
```
# <font color="blue">Test Image 5</font>
```
!wget https://www.dropbox.com/s/dbkag5gsicxqoqg/streetsign2.jpg?dl=0 -O streetsign2.jpg --quiet
```
### <font color="green">Downloaded Image</font>
<img src="https://www.dropbox.com/s/dbkag5gsicxqoqg/streetsign2.jpg?dl=1" width=500>
### <font color="green">Output</font>
```
ss2_text = pytesseract.image_to_string('streetsign2.jpg')
print(ss2_text)
```
# <font color="blue">Test Image 6</font>
```
!wget https://www.dropbox.com/s/cgni28zl1k9sesk/streetsign3.jpg?dl=0 -O streetsign3.jpg --quiet
```
### <font color="green">Downloaded Image</font>
<img src="https://www.dropbox.com/s/cgni28zl1k9sesk/streetsign3.jpg?dl=1" width=500>
### <font color="green">Output</font>
```
ss3_text = pytesseract.image_to_string('streetsign3.jpg')
print(ss3_text)
```
### <font color="green">Observation</font>
In all the above 3 images, there are multiple issues:
- text is not aligned. It is either rotated within the plane or into the plane.
- Large variation in Text size
- Background clutter
Next, We will discuss how to overcome some of the limitations of Tesseract by using different techniques.
| github_jupyter |
**Note that the name of the callback `AccumulateStepper` has been changed into `AccumulateScheduler`**
https://forums.fast.ai/t/accumulating-gradients/33219/90?u=hwasiti
https://github.com/fastai/fastai/blob/fbbc6f91e8e8e91ba0e3cc98ac148f6b26b9e041/fastai/train.py#L99-L134
```
import fastai
from fastai.vision import *
gpu_device = 0
defaults.device = torch.device(f'cuda:{gpu_device}')
torch.cuda.set_device(gpu_device)
def seed_everything(seed):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.deterministic = True
seed_everything(42)
path = untar_data(URLs.PETS)
path_anno = path/'annotations'
path_img = path/'images'
fnames = get_image_files(path_img)
pat = re.compile(r'/([^/]+)_\d+.jpg$')
# Simplified RunningBatchNorm
# 07_batchnorm.ipynb (fastai course v3 part2 2019)
class RunningBatchNorm2d(nn.Module):
def __init__(self, nf, mom=0.1, eps=1e-5):
super().__init__()
# I have added self.nf so that it can be represented when
# printing the model in the extra_repr method below
self.nf = nf
self.mom, self.eps = mom, eps
self.mults = nn.Parameter(torch.ones (nf,1,1))
self.adds = nn.Parameter(torch.zeros(nf,1,1))
self.register_buffer('sums', torch.zeros(1,nf,1,1))
self.register_buffer('sqrs', torch.zeros(1,nf,1,1))
self.register_buffer('count', tensor(0.))
self.register_buffer('factor', tensor(0.))
self.register_buffer('offset', tensor(0.))
self.batch = 0
def update_stats(self, x):
bs,nc,*_ = x.shape
self.sums.detach_()
self.sqrs.detach_()
dims = (0,2,3)
s = x .sum(dims, keepdim=True)
ss = (x*x).sum(dims, keepdim=True)
c = s.new_tensor(x.numel()/nc)
mom1 = s.new_tensor(1 - (1-self.mom)/math.sqrt(bs-1))
self.sums .lerp_(s , mom1)
self.sqrs .lerp_(ss, mom1)
self.count.lerp_(c , mom1)
self.batch += bs
means = self.sums/self.count
varns = (self.sqrs/self.count).sub_(means*means)
if bool(self.batch < 20): varns.clamp_min_(0.01)
self.factor = self.mults / (varns+self.eps).sqrt()
self.offset = self.adds - means*self.factor
def forward(self, x):
if self.training: self.update_stats(x)
return x*self.factor + self.offset
def extra_repr(self):
return '{nf}, mom={mom}, eps={eps}'.format(**self.__dict__)
class RunningBatchNorm1d(nn.Module):
def __init__(self, nf, mom=0.1, eps=1e-5):
super().__init__()
# I have added self.nf so that it can be represented when
# printing the model in the extra_repr method below
self.nf = nf
self.mom, self.eps = mom, eps
self.mults = nn.Parameter(torch.ones (nf,1,1))
self.adds = nn.Parameter(torch.zeros(nf,1,1))
self.register_buffer('sums', torch.zeros(1,nf,1,1))
self.register_buffer('sqrs', torch.zeros(1,nf,1,1))
self.register_buffer('count', tensor(0.))
self.register_buffer('factor', tensor(0.))
self.register_buffer('offset', tensor(0.))
self.batch = 0
def update_stats(self, x):
bs,nc,*_ = x.shape
self.sums.detach_()
self.sqrs.detach_()
dims = (0,2)
s = x .sum(dims, keepdim=True)
ss = (x*x).sum(dims, keepdim=True)
c = s.new_tensor(x.numel()/nc)
mom1 = s.new_tensor(1 - (1-self.mom)/math.sqrt(bs-1))
self.sums .lerp_(s , mom1)
self.sqrs .lerp_(ss, mom1)
self.count.lerp_(c , mom1)
self.batch += bs
means = self.sums/self.count
varns = (self.sqrs/self.count).sub_(means*means)
if bool(self.batch < 20): varns.clamp_min_(0.01)
self.factor = self.mults / (varns+self.eps).sqrt()
self.offset = self.adds - means*self.factor
def forward(self, x):
if self.training: self.update_stats(x)
return x*self.factor + self.offset
def extra_repr(self):
return '{nf}, mom={mom}, eps={eps}'.format(**self.__dict__)
```
### No Grad Acc (BS 64), No running BN
```
seed_everything(2)
data = ImageDataBunch.from_name_re(path_img, fnames, pat, ds_tfms=get_transforms(), size=224, bs=64
).normalize(imagenet_stats)
learn = cnn_learner(data, models.resnet18, metrics=accuracy)
learn.fit(1)
data.batch_size
```
### No Grad Acc (BS 2), No running BN
```
seed_everything(2)
data = ImageDataBunch.from_name_re(path_img, fnames, pat, ds_tfms=get_transforms(), size=224, bs=2
).normalize(imagenet_stats)
learn = cnn_learner(data, models.resnet18, metrics=accuracy)
learn.fit(1)
```
### Naive Grad Acc (BS 2) x 32 steps, No running BN
```
seed_everything(2)
data = ImageDataBunch.from_name_re(path_img, fnames, pat, ds_tfms=get_transforms(), size=224, bs=2
).normalize(imagenet_stats)
learn = cnn_learner(data, models.resnet18, metrics=accuracy,
callback_fns=[partial(AccumulateScheduler, n_step=32)])
learn.loss_func = CrossEntropyFlat(reduction='sum')
learn.fit(1)
```
### No Grad Acc (BS 2), Running BN
```
def bn2rbn(bn):
if isinstance(bn, nn.BatchNorm1d): rbn = RunningBatchNorm1d(bn.num_features, eps=bn.eps, mom=bn.momentum)
elif isinstance(bn, nn.BatchNorm2d): rbn = RunningBatchNorm2d(bn.num_features, eps=bn.eps, mom=bn.momentum)
rbn.weight = bn.weight
rbn.bias = bn.bias
return (rbn).to(bn.weight.device)
def convert_bn(list_mods, func=bn2rbn):
for i in range(len(list_mods)):
if isinstance(list_mods[i], bn_types):
list_mods[i] = func(list_mods[i])
elif list_mods[i].__class__.__name__ in ("Sequential", "BasicBlock"):
list_mods[i] = nn.Sequential(*convert_bn(list(list_mods[i].children()), func))
return list_mods
seed_everything(2)
data = ImageDataBunch.from_name_re(path_img, fnames, pat, ds_tfms=get_transforms(), size=224, bs=2
).normalize(imagenet_stats)
learn = cnn_learner(data, models.resnet18, metrics=accuracy)
# learn.loss_func = CrossEntropyFlat(reduction='sum')
learn.model
learn.summary()
learn.model = nn.Sequential(*convert_bn(list(learn.model.children()), bn2rbn))
learn.model
learn.summary()
%debug
learn.fit(1)
```
### GroupNorm
```
seed_everything(2)
data = ImageDataBunch.from_name_re(path_img, fnames, pat, ds_tfms=get_transforms(), size=224, bs=2
).normalize(imagenet_stats)
learn = create_cnn(data, models.resnet18, metrics=accuracy,
callback_fns=[partial(AccumulateScheduler, n_step=32)])
# learn.loss_func = CrossEntropyFlat(reduction='sum')
groups = 64
def bn2group(bn):
groupnorm = nn.GroupNorm(groups, bn.num_features, affine=True)
groupnorm.weight = bn.weight
groupnorm.bias = bn.bias
groupnorm.eps = bn.eps
return (groupnorm).to(bn.weight.device)
def convert_bn(list_mods, func=bn2group):
for i in range(len(list_mods)):
if isinstance(list_mods[i], bn_types):
list_mods[i] = func(list_mods[i])
elif list_mods[i].__class__.__name__ in ("Sequential", "BasicBlock"):
list_mods[i] = nn.Sequential(*convert_bn(list(list_mods[i].children()), func))
return list_mods
seed_everything(2)
data = ImageDataBunch.from_name_re(path_img, fnames, pat, ds_tfms=get_transforms(), size=224, bs=2
).normalize(imagenet_stats)
learn = create_cnn(data, models.vgg16_bn, metrics=accuracy,
callback_fns=[partial(AccumulateStepper, n_step=32)])
learn.loss_func = CrossEntropyFlat(reduction='sum')
learn.model = nn.Sequential(*convert_bn(list(learn.model.children()), bn2group))
learn.freeze()
learn.fit(1)
```
### Resnet + GroupNorm
```
seed_everything(2)
data = ImageDataBunch.from_name_re(path_img, fnames, pat, ds_tfms=get_transforms(), size=224, bs=2
).normalize(imagenet_stats)
learn = create_cnn(data, models.resnet18, metrics=accuracy,
callback_fns=[partial(AccumulateStepper, n_step=32)])
learn.loss_func = CrossEntropyFlat(reduction='sum')
def change_all_BN(module):
for i in range(5):
atr = 'bn'+str(i)
if hasattr(module, atr):
setattr(module, atr, bn2group(getattr(module,atr)))
def wrap_BN(model):
for i in range(len(model)):
for j in range(len(model[i])):
if isinstance(model[i][j], bn_types):
model[i][j] = bn2group(model[i][j])
elif model[i][j].__class__.__name__ == "Sequential":
for k in range(len(model[i][j])):
if isinstance(model[i][j][k], bn_types):
model[i][j][k] = bn2group(model[i][j][k])
elif model[i][j][k].__class__.__name__ == "BasicBlock":
change_all_BN(model[i][j][k])
if hasattr(model[i][j][k],'downsample'):
if model[i][j][k].downsample is not None:
for l in range(len(model[i][j][k].downsample)):
if isinstance(model[i][j][k].downsample[l], bn_types):
model[i][j][k].downsample[l] = bn2group(model[i][j][k].downsample[l])
wrap_BN(learn.model)
learn.freeze()
learn.fit(1)
```
### Resnet + GroupNorm (No Acc)
```
seed_everything(2)
data = ImageDataBunch.from_name_re(path_img, fnames, pat, ds_tfms=get_transforms(), size=224, bs=2
).normalize(imagenet_stats)
learn = create_cnn(data, models.resnet18, metrics=accuracy)
wrap_BN(learn.model)
learn.freeze()
learn.fit(1)
```
### Resnet + GroupNorm (No Acc) bs = 1
```
seed_everything(2)
data = ImageDataBunch.from_name_re(path_img, fnames, pat, ds_tfms=get_transforms(), size=224, bs=1
).normalize(imagenet_stats)
learn = create_cnn(data, models.resnet18, metrics=accuracy)
wrap_BN(learn.model)
learn.freeze()
learn.fit(1)
```
| github_jupyter |
```
import torch, pickle, time, os, random
from sklearn.model_selection import train_test_split
import numpy as np
import os.path as osp
import matplotlib.pyplot as plt
import torch_geometric as tg
from torch_geometric.loader import DataLoader
# accelerate huggingface to GPU
if torch.cuda.is_available():
from accelerate import Accelerator
accelerator = Accelerator()
device = accelerator.device
torch.manual_seed(42)
random.seed(42)
os.listdir(osp.expanduser('~/../../../scratch/gpfs/cj1223/GraphStorage/'))
case='vlarge_all_smass_z0.0_quantile_quant'
case='vlarge_all_smass'
case='vlarge_all_4t_z0.0_quantile_raw'
# case='vlarge_all_4t_z0.0_standard_raw'
datat=pickle.load(open(osp.expanduser(f'~/../../../scratch/gpfs/cj1223/GraphStorage/{case}/data.pkl'), 'rb'))
from torch_geometric.data import Data
data=[]
for d in datat:
data.append(Data(x=d.x, edge_index=d.edge_index, edge_attr=d.edge_attr, y=d.y[[0]]))
try:
n_targ=len(data[0].y)
except:
n_targ=1
n_feat=len(data[0].x[0])
n_feat
from torch import nn
class MLP(nn.Module):
def __init__(self, n_in, n_out, hidden=64, nlayers=2, layer_norm=True):
super().__init__()
layers = [nn.Linear(n_in, hidden), nn.ReLU()]
for i in range(nlayers):
layers.append(nn.Linear(hidden, hidden))
layers.append(nn.ReLU())
if layer_norm:
layers.append(nn.LayerNorm(hidden))
layers.append(nn.Linear(hidden, n_out))
self.mlp = nn.Sequential(*layers)
def forward(self, x):
return self.mlp(x)
import torch.nn.functional as F
from torch.nn import Linear, BatchNorm1d, LayerNorm
from torch_geometric.nn import SAGEConv, global_mean_pool, norm, global_max_pool, global_add_pool
class GCN(torch.nn.Module):
def __init__(self, hidden_channels, nlin=3):
super(GCN, self).__init__()
self.node_enc = MLP(n_feat, hidden_channels, layer_norm=True)
ag='add'
self.conv1 = SAGEConv(hidden_channels, hidden_channels, aggr=ag)
self.conv2 = SAGEConv(hidden_channels, hidden_channels, aggr=ag)
self.conv3 = SAGEConv(hidden_channels, hidden_channels, aggr=ag)
self.conv4 = SAGEConv(hidden_channels, hidden_channels, aggr=ag)
self.conv5 = SAGEConv(hidden_channels, hidden_channels, aggr=ag)
self.lin = Linear(hidden_channels, hidden_channels)
self.norm = LayerNorm(normalized_shape=hidden_channels) # layer_norm instead
self.lin_f = Linear(hidden_channels, n_targ)
self.lin_var = Linear(hidden_channels, hidden_channels)
self.norm_var = LayerNorm(normalized_shape=hidden_channels) # layer_norm instead
self.lin_f_var = Linear(hidden_channels, n_targ)
def forward(self, x, edge_index, batch):
# 1. Obtain node embeddings
x = self.node_enc(x)
x = self.conv1(x, edge_index)
x = x.relu()
x = self.conv2(x, edge_index)
x = x.relu()
x = self.conv3(x, edge_index)
x = x.relu()
x = self.conv4(x, edge_index)
x = x.relu()
x = self.conv5(x, edge_index)
x = x.relu()
x1 = global_add_pool(x, batch)
# x1 = torch.cat([global_max_pool(x, batch),global_add_pool(x, batch)], 1) ## Miles says use sumpool
x = self.lin(x1)
x = self.lin_f(self.norm(x))
sig = self.lin_var(x1)
sig = self.lin_f_var(self.norm(sig))
return x, torch.abs(sig)
model = GCN(hidden_channels=64)
next(model.parameters()).is_cuda ##check number one
criterion = torch.nn.GaussianNLLLoss()
n_epochs=250
n_trials=1
batch_size=128
split=0.8
test_data=data[int(len(data)*split):]
train_data=data[:int(len(data)*split)]
# train_data, test_data=train_test_split(data, test_size=0.2)
l1_lambda = 1e-5
l2_lambda = 1e-5
def l_func(pred, ys, sig):
global z, sigloss
z=(pred-ys)/sig
sigloss=torch.sum(torch.log(sig))
err_loss = torch.sum(z**2)/2
return err_loss+sigloss, err_loss, sigloss
trains, tests, scatter = [], [], []
yss, preds=[],[]
model = GCN(hidden_channels=128)
train_loader=DataLoader(train_data, batch_size=batch_size, shuffle=1, num_workers=4)
test_loader=DataLoader(test_data, batch_size=batch_size, shuffle=0,num_workers=4)
optimizer = torch.optim.Adam(model.parameters(), lr=0.003)
_, _, test_loader = accelerator.prepare(model, optimizer, test_loader)
model, optimizer, train_loader = accelerator.prepare(model, optimizer, train_loader)
print('GPU ', next(model.parameters()).is_cuda)
# Initialize our train function
def train():
model.train()
for data in train_loader:
out, var = model(data.x, data.edge_index, data.batch)
# out, var1=outs[:,0],torch.abs(outs[:,1])
# var=torch.ones_like(var)/100
# loss = criterion(out, data.y.view(-1,n_targ), var)
loss, _,_ = l_func(out, data.y.view(-1,n_targ), var)
l1_norm = sum(p.abs().sum() for p in model.parameters())
l2_norm = sum(p.pow(2.0).sum() for p in model.parameters())
loss = loss + l1_lambda * l1_norm + l2_lambda * l2_norm
# loss.backward()
accelerator.backward(loss)
optimizer.step()
optimizer.zero_grad()
# print(loss, l1_norm*l1_lambda, l2_norm*l2_lambda)
# test function
def test(loader):
model.eval()
outss = []
ys = []
varss = []
with torch.no_grad(): ##this solves it!!!
for dat in loader:
out, var = model(dat.x, dat.edge_index, dat.batch)
# outs = model(dat.x, dat.edge_index, dat.batch)
# out, var=outs[:,0],torch.abs(outs[:,1])
ys.append(dat.y.view(-1,n_targ))
outss.append(out)
varss.append(var)
outss=torch.vstack(outss)
yss=torch.vstack(ys)
varss=torch.vstack(varss)
return torch.std(outss - yss, axis=0), outss, yss, varss
# return outss, yss, varss
s, outs, ys, varss=test(train_loader)
#this uses about 1 GB of memory on the GPU
tr_acc, te_acc = [], []
start=time.time()
for epoch in range(n_epochs):
train()
if (epoch+1)%2==0:
train_acc, _ , _, _ = test(train_loader)
test_acc, _ , _ , _= test(test_loader)
tr_acc.append(train_acc.cpu().numpy())
te_acc.append(test_acc.cpu().numpy())
print(f'Epoch: {epoch+1:03d}, Train scatter: {np.round(train_acc.cpu().numpy(), 4)} \n \
Test scatter: {np.round(test_acc.cpu().numpy(), 4)}')
stop=time.time()
spent=stop-start
print(f"{spent:.2f} seconds spent training, {spent/n_epochs:.3f} seconds per epoch. Processed {len(data)*split*n_epochs/spent:.0f} trees per second")
plt.plot(tr_acc)
plt.plot(te_acc)
teststd, outtest, ytest, var = test(test_loader)
pred=outtest.cpu().numpy()
ys=ytest.cpu().numpy()
var=var.cpu().numpy()
teststd
tot=np.vstack([ys,pred])
fig , ax = plt.subplots(1, figsize=(10,6))
l=0
for k in range(n_targ):
# ax[k].hist(outtest.cpu().numpy()[:,k]-ytest.cpu().numpy()[:,k], bins=1000, histtype='step', label='res')
ax.hist(pred, bins=50, range=list(np.percentile(tot, [l,100-l])), histtype='step', label='pred' )
ax.hist(ys, bins=50, range=list(np.percentile(tot, [l,100-l])),histtype='step', label='true')
ax.legend()
plt.hist((ys-pred)/var, bins=50);
Mh=[]
for d in test_data:
Mh.append(d.x.numpy()[0,3])
plt.hist(Mh, bins=50);
np.corrcoef(Mh, var.flatten())
fig , ax = plt.subplots(1,1, figsize=(10,10))
yte=ys
predte=pred
ax.plot(yte, predte, 'ro', alpha=0.2)
ax.plot([min(yte),max(yte)],[min(yte),max(yte)], 'k--', label='Perfect correspondance')
ax.set(title=[f'Stellar mass after {epoch} epochs', np.round(np.std(yte-predte),3), np.round(np.mean(yte-predte),2)], xlabel='True [dex]', ylabel='Predicted [dex]')
ax.legend()
transform='quantile'
transform_path=osp.expanduser(f"~/../../../scratch/gpfs/cj1223/GraphStorage/transformers/{transform}_alltarg_1.pkl")
target_scaler=pickle.load(open(transform_path, 'rb'))
transform_path=osp.expanduser(f"~/../../../scratch/gpfs/cj1223/GraphStorage/transformers/{transform}_allfeat_1.pkl")
feat_scaler=pickle.load(open(transform_path, 'rb'))
fig , ax = plt.subplots(1,1, figsize=(10,10))
yte=np.hstack(target_scaler[8].inverse_transform(ys.reshape(-1,1)))
predte=np.hstack(target_scaler[8].inverse_transform(pred.reshape(-1,1)))
ax.plot(yte, predte, 'ro', alpha=0.2)
ax.plot([min(yte),max(yte)],[min(yte),max(yte)], 'k--', label='Perfect correspondance')
ax.set(title=[f'Stellar mass after {epoch} epochs', np.round(np.std(yte-predte),2), np.round(np.mean(yte-predte),2)], xlabel='True [dex]', ylabel='Predicted [dex]')
ax.legend()
```
| github_jupyter |
# SpaceNet Rio Chip Classification Data Prep
This notebook prepares data for training a chip classification model on the Rio SpaceNet dataset.
* Set `raw_uri` to the local or S3 directory containing the raw dataset.
* Set `processed_uri` to a local or S3 directory (you can write to), which will store the processed data generated by this notebook.
This is all you will need to do in order to run this notebook.
```
raw_uri = 's3://spacenet-dataset/'
# processed_uri = 's3://raster-vision-lf-dev/examples/spacenet/rio/processed-data/'
processed_uri = '/opt/data/examples/spacenet/rio/processed-data'
```
The steps we'll take to make the data are as follows:
- Get the building labels and AOI (most likely from the SpaceNet AWS public dataset bucket)
- Use the AOI and the image bounds to determine which images can be used for training and validation
- Split the building labels by image, save a label GeoJSON file per image
- Split the labeled images into a training and validation set, using the percentage of the AOI each covers, aiming at an 80%/20% split.
This process will save the split label files, and `train_scenes.csv` and `val_scenes.csv` files that are used by the experiment at `experiments.spacenet.chip_classification`
```
import os
from os.path import join
import json
import tempfile
import boto3
import botocore
import rasterio
from shapely.geometry import (Polygon, shape)
import rastervision as rv
from rastervision.utils.files import (
download_if_needed, list_paths, file_to_json, json_to_file,
get_local_path, make_dir, sync_to_dir, str_to_file)
```
## Get the label and AOI data from AWS's public dataset of Space Net
```
label_uri = join(raw_uri, 'AOI_1_Rio/srcData/buildingLabels/Rio_Buildings_Public_AOI_v2.geojson')
aoi_uri = join(raw_uri, 'AOI_1_Rio/srcData/buildingLabels/Rio_OUTLINE_Public_AOI.geojson')
label_json = file_to_json(label_uri)
aoi_json = file_to_json(aoi_uri)
```
## Use the AOI to determine what images are inside the training set
Here we compare the AOI to the image extents to deteremine which images we can use for training and validation. We're using `rasterio`'s ability to read the metadata from raster data on S3 without downloading the whole image
```
aoi = shape(aoi_json['features'][0]['geometry'])
aoi
images_uri = join(raw_uri, 'AOI_1_Rio/srcData/mosaic_3band')
image_paths = list_paths(images_uri)
def bounds_to_shape(bounds):
return Polygon([[bounds.left, bounds.bottom],
[bounds.left, bounds.top],
[bounds.right, bounds.top],
[bounds.right, bounds.bottom],
[bounds.left, bounds.bottom]])
image_to_extents = {}
for img in image_paths:
with rasterio.open(img, 'r') as ds:
image_to_extents[img] = bounds_to_shape(ds.bounds)
intersecting_images = []
for img in image_to_extents:
if image_to_extents[img].intersects(aoi):
intersecting_images.append(img)
intersecting_images
```
## Match labels to images
Find the labels that intersect with the image's bounding box, which will be saved into a labels geojson that matches the image name.
```
# Add a class_id and class_name to the properties of each feature
for feature in label_json['features']:
feature['properties']['class_id'] = 1
feature['properties']['class_name'] = 'building'
image_to_features = {}
for img in intersecting_images:
image_to_features[img] = []
bbox = image_to_extents[img]
for feature in label_json['features']:
if shape(feature['geometry']).intersects(bbox):
image_to_features[img].append(feature)
processed_labels_uri = join(processed_uri, 'labels')
for img in image_to_features:
fc = {}
fc['type'] = 'FeatureCollection'
fc['crs'] = label_json['crs']
fc['features'] = image_to_features[img]
img_id = os.path.splitext(os.path.basename(img))[0]
label_path = join(processed_labels_uri, '{}.geojson'.format(img_id))
json_to_file(fc, label_path)
```
## Split into train and validation
Split up training and validation data. There's an odd shaped AOI and not that many images, so we'll split the train and validation roughly based on how much area each scene covers of the AOI.
Create a CSV that our experiments will use to load the training and validation data.
```
# Split training and validation
ratio = 0.8
aoi_area = aoi.area
images_to_area = {}
for img in intersecting_images:
area = image_to_extents[img].intersection(aoi).area
images_to_area[img] = area / aoi_area
train_imgs = []
val_imgs = []
train_area_covered = 0
for img in sorted(intersecting_images, reverse=True, key=lambda img: images_to_area[img]):
if train_area_covered < ratio:
train_imgs.append(img)
train_area_covered += images_to_area[img]
else:
val_imgs.append(img)
print("{} training images with {}% area.".format(len(train_imgs), train_area_covered))
print("{} validation images with {} area.".format(len(val_imgs), 1 - train_area_covered))
def save_split_csv(imgs, path):
csv_rows = []
for img in imgs:
img_id = os.path.splitext(os.path.basename(img))[0]
img_path = join('AOI_1_Rio', 'srcData/mosaic_3band', '{}.tif'.format(img_id))
label_path = join('labels','{}.geojson'.format(img_id))
csv_rows.append('"{}","{}"'.format(img_path, label_path))
str_to_file('\n'.join(csv_rows), path)
save_split_csv(train_imgs, join(processed_uri, 'train-scenes.csv'))
save_split_csv(val_imgs, join(processed_uri, 'val-scenes.csv'))
```
| github_jupyter |
```
import os, re
import pandas as pd
import missingno as msno
import numpy as np
import matplotlib.pyplot as plt
# CURRENT DIRECTORY (PLACE SCRIPT IN /home/mydirectory)
cd = os.getcwd()
fileNames=[]
reviewRegex = re.compile(r'newest_rev(ie|ei)ws')
i = 0
columns = range(1,100)
dfList = []
for root, dirs, files in os.walk(cd):
for fname in files:
if reviewRegex.search(fname):
frame = pd.read_csv(os.path.join(root, fname), names=columns,skiprows=1)
if fname.find("EDUCATION") != -1:
frame['CATEGORY'] = "EDUCATION"
elif fname.find("FINANCE") !=-1:
frame['CATEGORY'] = "FINANCE"
elif fname.find("ENTERTAINMENT") != -1:
frame['CATEGORY'] = "ENTERTAINMENT"
elif fname.find("FAMILY") != -1:
frame['CATEGORY'] = "FAMILY"
elif fname.find("GAME") != -1:
frame['CATEGORY'] = "GAME-ACTION"
elif fname.find("HEALTH") != -1:
frame['CATEGORY'] = "HEALTH AND FITNESS"
elif fname.find("LIFESTYLE") != -1:
frame['CATEGORY'] = "LIFESTYLE"
elif fname.find("MUSIC") != -1:
frame['CATEGORY'] = "MUSIC AND AUDIO"
#frame['key'] = "file{}".format(i)
dfList.append(frame)
fileNames.append(fname)
i += 1
detailsRegex = re.compile(r'_all_detailed')
i = 0
columns = range(1,100)
detailsFiles=[]
detailsList=[]
for root, dirs, files in os.walk(cd):
for fname in files:
if detailsRegex.search(fname):
frame = pd.read_csv(os.path.join(root, fname), names=columns,skiprows=0)
if fname.find("EDUCATION") != -1:
frame['CATEGORY'] = "EDUCATION"
elif fname.find("FINANCE") !=-1:
frame['CATEGORY'] = "FINANCE"
elif fname.find("ENTERTAINMENT") != -1:
frame['CATEGORY'] = "ENTERTAINMENT"
elif fname.find("FAMILY") != -1:
frame['CATEGORY'] = "FAMILY"
elif fname.find("GAME") != -1:
frame['CATEGORY'] = "GAME-ACTION"
elif fname.find("HEALTH") != -1:
frame['CATEGORY'] = "HEALTH AND FITNESS"
elif fname.find("LIFESTYLE") != -1:
frame['CATEGORY'] = "LIFESTYLE"
elif fname.find("MUSIC") != -1:
frame['CATEGORY'] = "MUSIC AND AUDIO"
#frame['key'] = "file{}".format(i)
detailsList.append(frame)
detailsFiles.append(fname)
i += 1
```
##### ALL detailed dataset analysis
```
len(detailsList)
detailsDf = pd.concat(detailsList)
detailsDf
detailsDf.columns = detailsDf.iloc[0]
pd.set_option('display.max_columns', 30)
detailsDf.dropna(axis=1,how="all",inplace=True)
#msno.matrix(detailsDf)
detailsDf.tail()
detailsDf = detailsDf.loc[:, detailsDf.columns.notnull()]
detailsDf.reset_index(drop=True,inplace=True)
detailsDf.drop(index=0,inplace=True)
detailsDf.appTitle = detailsDf.appTitle.str.lower()
detailsDf.drop_duplicates(subset=['appTitle','summary','installs','minInstalls','score','ratings','reviews','free','offersIAP','size','androidVersionText','developerId','genreId','familyGenreId','contentRating'],inplace=True)
detailsDf.drop(index=527,inplace=True)
detailsDf.rename(columns={"EDUCATION":"CATEGORY"},inplace=True)
detailsDf['reviews'] = pd.to_numeric(detailsDf['reviews'], errors='coerce')
detailsDf.contentRating.unique()
detailsDf
detailsDf
```
###### Content Rating groups in each of the eight categories and their number
```
detailsDf.contentRating.unique()
pd.DataFrame(detailsDf.groupby("CATEGORY")["contentRating"].unique())
detailsDf.groupby(["CATEGORY","contentRating"])["contentRating"].count()
```
*******
#### Reviews Analysis
```
len(fileNames)
df = pd.concat(dfList)
df.reset_index(drop=True,inplace=True)
df.rename(columns={1:"appTitle",2:"userName",3:"date",4:"score",5:"text"},inplace=True)
df.dropna(axis=1,how="all",inplace=True)
df['appTitle'] = df['appTitle'].str.lower()
```
#### Reviews Data set after reading each CSV and attaching category as additional column
```
df
df.tail()
```
##### Convert all the app title to lower to avoid any duplicates with other columns as a result of having same name but are different as a result of case sensitivity
```
df['appTitle'] = df['appTitle'].str.lower()
```
##### Drop all the possible duplicates
```
df.drop_duplicates(subset=['appTitle','userName','date','score','text'],inplace=True)
```
##### Drop any duplicates that have the same app Title and text to clear out even more misleading data
```
df.dropna(subset=['appTitle','text'],inplace=True)
df
```
##### Total number of unique reviews
```
len(df.text.unique())
```
##### Unique number of Apps in the dataset
```
len(df.appTitle.unique())
```
##### Number of apps in each of the specified 8 categories
```
df.groupby("CATEGORY")['appTitle'].nunique()
```
#### Number of reviews per category
```
df.groupby("CATEGORY")['text'].count()
JoinedDf = pd.merge(df,detailsDf[['appTitle','contentRating']],how="left",on="appTitle",)
JoinedDf.drop_duplicates(subset=['appTitle','userName','date','score','text','CATEGORY'],inplace=True)
JoinedDf.reset_index(drop=True,inplace=True)
JoinedDf
JoinedDf.groupby(["CATEGORY","contentRating"])["contentRating"].count()
```
##### Remove non english reviews using NLTK
```
import nltk
nltk.download('words')
words = set(nltk.corpus.words.words())
def review(text):
return (" ".join(w for w in nltk.wordpunct_tokenize(text) if w.lower() in words or not w.isalpha()))
JoinedDf["processedText"] = JoinedDf["text"]
JoinedDf["processedText"] = JoinedDf["processedText"].astype(str)
JoinedDf["processedText"] = JoinedDf["processedText"].apply(review)
JoinedDf
```
#### Remove non ascii characters
```
JoinedDf.processedText.replace({r'[^\x00-\x7F]+':''}, regex=True, inplace=True)
```
#### Remove punctuations
```
JoinedDf["processedText"] = JoinedDf['processedText'].str.replace('[^\w\s]','',)
```
##### Remove multiple characters if they occur more than two times continuously
```
def removeMultiple(text):
return re.sub(r'([a-z])\1{2,}', r'\1\1', text)
JoinedDf['processedText']= JoinedDf['processedText'].map(removeMultiple)
JoinedDf
```
###### 7. Should we remove the reviews that contain two or less number of words?
A) We should not remove reviews that contain two more words. In case sentiment analysis needs to be carried , a two word review such as, "Great APP" is meaningful about the app and gives the true sentiment of that review. Hence taking away such reviews would just loose make us loose out on meaningful data that could be used for analysis.
B)Removing reviews just based on the score would not be recommended. This would cause the the eventual model to be biased.
###### Remove reviews that have less than two words
```
def moreThanTwoWords(text):
if len(text.split(" ")) > 2:
return text
return np.nan
JoinedDf["processedText"] = JoinedDf["processedText"].apply(moreThanTwoWords)
JoinedDf
```
###### 8. How many reviews exist in each of the eight app-categories? Compare with question 3.
```
print("The number of Unique reviews after processing the text is ",len(JoinedDf["processedText"].unique()),
"and before text processing is",len(JoinedDf["text"].unique()))
len(JoinedDf["text"].unique())
JoinedDf.groupby("CATEGORY")["processedText"].count()
pd.concat([df.groupby("CATEGORY")['text'].count(),JoinedDf.groupby("CATEGORY")["processedText"].count()],axis=1)
```
###### 9. How many reviews in each contentRating-sub-groups in each app-category? Compare with question 6.
```
pd.concat([JoinedDf.groupby(["CATEGORY","contentRating"])["text"].count(),JoinedDf.groupby(["CATEGORY","contentRating"])["processedText"].count()],axis=1)
```
##### Analyzing each category
##### 10. What is the number of reviews for each score (score column)? For example, 35000 reviews have a score of 1, etc.
###### Before text processing
```
JoinedDf.groupby("score")["text"].count()
```
##### After text processing
```
JoinedDf.groupby("score")["processedText"].count()
```
###### 11. How many apps exist in each score-sub-group?
```
JoinedDf.groupby("score")["appTitle"].nunique()
```
###### 12. Compare the number of reviews for each score in a plot (Remember to normalize the numbers when you are comparing them).
```
JoinedDf
Normalize = pd.DataFrame(JoinedDf.groupby("score")["processedText"].count())
Normalize.columns = ["Number of Reviews for processed text"]
Normalize
## Normalize using mix max data
Normalize["Normalizereviews"]= (Normalize["Number of Reviews for processed text"]-Normalize["Number of Reviews for processed text"].min())/(Normalize["Number of Reviews for processed text"].max()-Normalize["Number of Reviews for processed text"].min())
Normalize
Normalize["score"] = Normalize.index
Normalize.plot.bar(x="score",y="Normalizereviews")
```
###### 13. What is the average length of the reviews in each score-sub-group?
```
AvgLength = JoinedDf.copy()
AvgLength ["AverageLen"]= JoinedDf["text"].str.len()
AvgLengthTable = pd.DataFrame(AvgLength.groupby("score")["AverageLen"].mean())
AvgLengthTable
```
##### 14. Compare the average length of reviews in each score-sub-group in the 8 app categories (draw a plot).
```
AvgLength
AvgPlot = pd.DataFrame(AvgLength.groupby(["CATEGORY",'score'])['AverageLen'].mean())
AvgPlot.unstack().plot(kind="bar",title="Average Length of reviews for each category with each score")
plt.legend(loc="center left",bbox_to_anchor=(1,0.4))
```
###### 15. Compare the number of reviews in each score-sub-group in the 8 app categories (draw a plot).
```
AvgReview = pd.DataFrame(AvgLength.groupby(["CATEGORY","score"])["text"].count())
AvgReview.unstack().plot(kind="bar",title="Average Number of reviews for each category with each score")
plt.legend(loc="center left",bbox_to_anchor=(1,0.4))
```
###### 16. Compare the number of apps in each score-sub-group in the 8 app categories (draw a plot).
```
AverageApp = pd.DataFrame(AvgLength.groupby(['CATEGORY',"score"])['appTitle'].nunique())
AverageApp.unstack().plot(kind="bar",title="Average Number of Apps in each subgroup in 8 categories")
plt.legend(loc="center left",bbox_to_anchor=(1,0.4))
```
###### 17. Is there any correlation between the length of the reviews and the score in each app-category?
```
RevScore = AvgLength.groupby("CATEGORY")[["score","AverageLen"]].apply(lambda x:x.corr()["AverageLen"])
RevScore.iloc[:,[0]]
```
###### There is negative correlation between the average Length and the score
###### 18. Find the evolution/changes of the star rating and length of reviews during time for each app category. Draw plots.
###### a. Can we use the date column for this question?
###### b. If not, what is the solution based on the data you have?
```
detailsRegex = re.compile(r'newest')
i = 0
columns = range(1,100)
df2files=[]
df2List=[]
for root, dirs, files in os.walk(cd):
for fname in files:
if detailsRegex.search(fname):
frame = pd.read_csv(os.path.join(root, fname), names=columns,skiprows=1)
if fname.find("EDUCATION") != -1:
frame['CATEGORY'] = "EDUCATION"
elif fname.find("FINANCE") !=-1:
frame['CATEGORY'] = "FINANCE"
elif fname.find("ENTERTAINMENT") != -1:
frame['CATEGORY'] = "ENTERTAINMENT"
elif fname.find("FAMILY") != -1:
frame['CATEGORY'] = "FAMILY"
elif fname.find("GAME") != -1:
frame['CATEGORY'] = "GAME-ACTION"
elif fname.find("HEALTH") != -1:
frame['CATEGORY'] = "HEALTH AND FITNESS"
elif fname.find("LIFESTYLE") != -1:
frame['CATEGORY'] = "LIFESTYLE"
elif fname.find("MUSIC") != -1:
frame['CATEGORY'] = "MUSIC AND AUDIO"
frame['month'] = fname[5:7]
#frame['key'] = "file{}".format(i)
df2List.append(frame)
df2files.append(fname)
i += 1
df2 = pd.concat(df2List)
df2.reset_index(drop=True,inplace=True)
df2.rename(columns={1:"appTitle",2:"userName",3:"date",4:"score",5:"text"},inplace=True)
df2.dropna(axis=1,how="all",inplace=True)
df2['appTitle'] = df2['appTitle'].str.lower()
df2.drop_duplicates(subset=['appTitle','userName','date','score','text'],inplace=True)
df2.dropna(subset=['appTitle','text'],inplace=True)
df2
```
##### 1)Drop duplicates and record how many unique reviews are collected?
###### Recording of unique number of reviews is done after dropping duplicates which have similar app Title,UserName,score,text and Category and then again dropping duplicates that have the same apptitle and review
```
len(df.text.unique())
```
Total number of unique reviews is 673592
###### 2. How many unique apps are in the dataset? How many apps in each of the 8 specified app categories?
```
print("Number of unique apps are",len(df['appTitle'].unique()))
print("Apps in each of the 8 specified app categories are")
df.groupby("CATEGORY")['appTitle'].nunique()
```
##### 3. How many reviews exist in each of the eight app-categories?
```
df.groupby("CATEGORY")['text'].count()
```
###### 4. Based on the contentRating column in the details files for each app, what are the different contentRating groups in each app-category?
###### 5. How many apps exist in each of these contentRating-sub-groups in each app-category?
```
pd.DataFrame(detailsDf.groupby("CATEGORY")["contentRating"].unique())
detailsDf.groupby(["CATEGORY","contentRating"])["contentRating"].count()
JoinedDf.groupby(["CATEGORY","contentRating"])["contentRating"].count()
```
##### 6. How many reviews in each contentRating-sub-groups in each app-category?
```
JoinedDf.groupby(["CATEGORY","contentRating"])["text"].count()
import nltk
nltk.download('words')
words = set(nltk.corpus.words.words())
def review(text):
return (" ".join(w for w in nltk.wordpunct_tokenize(text) if w.lower() in words or not w.isalpha()))
JoinedDf["processedText"] = JoinedDf["text"]
JoinedDf["processedText"] = JoinedDf["processedText"].astype(str)
JoinedDf["processedText"] = JoinedDf["processedText"].apply(review)
JoinedDf
```
#### Remove non ascii characters
```
JoinedDf.processedText.replace({r'[^\x00-\x7F]+':''}, regex=True, inplace=True)
```
#### Remove punctuations
```
JoinedDf["processedText"] = JoinedDf['processedText'].str.replace('[^\w\s]','',)
```
##### Remove multiple characters if they occur more than two times continuously
```
def removeMultiple(text):
return re.sub(r'([a-z])\1{2,}', r'\1\1', text)
JoinedDf['processedText']= JoinedDf['processedText'].map(removeMultiple)
JoinedDf
```
###### 7. Should we remove the reviews that contain two or less number of words?
A) We should not remove reviews that contain two more words. In case sentiment analysis needs to be carried , a two word review such as, "Great APP" is meaningful about the app and gives the true sentiment of that review. Hence taking away such reviews would just loose make us loose out on meaningful data that could be used for analysis.
B)Removing reviews just based on the score would not be recommended. This would cause the the eventual model to be biased.
###### Remove reviews that have less than two words
```
def moreThanTwoWords(text):
if len(text.split(" ")) > 2:
return text
return np.nan
JoinedDf["processedText"] = JoinedDf["processedText"].apply(moreThanTwoWords)
JoinedDf
```
###### 8. How many reviews exist in each of the eight app-categories? Compare with question 3.
```
print("The number of Unique reviews after processing the text is ",len(JoinedDf["processedText"].unique()),
"and before text processing is",len(JoinedDf["text"].unique()))
len(JoinedDf["text"].unique())
JoinedDf.groupby("CATEGORY")["processedText"].count()
pd.concat([df.groupby("CATEGORY")['text'].count(),JoinedDf.groupby("CATEGORY")["processedText"].count()],axis=1)
```
###### 9. How many reviews in each contentRating-sub-groups in each app-category? Compare with question 6.
```
pd.concat([JoinedDf.groupby(["CATEGORY","contentRating"])["text"].count(),JoinedDf.groupby(["CATEGORY","contentRating"])["processedText"].count()],axis=1)
```
##### Analyzing each category
##### 10. What is the number of reviews for each score (score column)? For example, 35000 reviews have a score of 1, etc.
###### Before text processing
```
JoinedDf.groupby("score")["text"].count()
```
##### After text processing
```
JoinedDf.groupby("score")["processedText"].count()
```
###### 11. How many apps exist in each score-sub-group?
```
JoinedDf.groupby("score")["appTitle"].nunique()
```
###### 12. Compare the number of reviews for each score in a plot (Remember to normalize the numbers when you are comparing them).
```
JoinedDf
Normalize = pd.DataFrame(JoinedDf.groupby("score")["processedText"].count())
Normalize.columns = ["Number of Reviews for processed text"]
Normalize
## Normalize using mix max data
Normalize["Normalizereviews"]= (Normalize["Number of Reviews for processed text"]-Normalize["Number of Reviews for processed text"].min())/(Normalize["Number of Reviews for processed text"].max()-Normalize["Number of Reviews for processed text"].min())
Normalize
Normalize["score"] = Normalize.index
Normalize.plot.bar(x="score",y="Normalizereviews")
```
###### 13. What is the average length of the reviews in each score-sub-group?
```
AvgLength = JoinedDf.copy()
AvgLength ["AverageLen"]= JoinedDf["text"].str.len()
AvgLengthTable = pd.DataFrame(AvgLength.groupby("score")["AverageLen"].mean())
AvgLengthTable
```
##### 14. Compare the average length of reviews in each score-sub-group in the 8 app categories (draw a plot).
```
AvgLength
AvgPlot = pd.DataFrame(AvgLength.groupby(["CATEGORY",'score'])['AverageLen'].mean())
AvgPlot.unstack().plot(kind="bar",title="Average Length of reviews for each category with each score")
plt.legend(loc="center left",bbox_to_anchor=(1,0.4))
```
###### 15. Compare the number of reviews in each score-sub-group in the 8 app categories (draw a plot).
```
AvgReview = pd.DataFrame(AvgLength.groupby(["CATEGORY","score"])["text"].count())
AvgReview.unstack().plot(kind="bar",title="Average Number of reviews for each category with each score")
plt.legend(loc="center left",bbox_to_anchor=(1,0.4))
```
###### 16. Compare the number of apps in each score-sub-group in the 8 app categories (draw a plot).
```
AverageApp = pd.DataFrame(AvgLength.groupby(['CATEGORY',"score"])['appTitle'].nunique())
AverageApp.unstack().plot(kind="bar",title="Average Number of Apps in each subgroup in 8 categories")
plt.legend(loc="center left",bbox_to_anchor=(1,0.4))
```
###### 17. Is there any correlation between the length of the reviews and the score in each app-category?
```
RevScore = AvgLength.groupby("CATEGORY")[["score","AverageLen"]].apply(lambda x:x.corr()["AverageLen"])
RevScore.iloc[:,[0]]
```
###### There is negative correlation between the average Length and the score
###### 18. Find the evolution/changes of the star rating and length of reviews during time for each app category. Draw plots.
###### a. Can we use the date column for this question?
###### b. If not, what is the solution based on the data you have?
```
detailsRegex = re.compile(r'newest')
i = 0
columns = range(1,100)
df2files=[]
df2List=[]
for root, dirs, files in os.walk(cd):
for fname in files:
if detailsRegex.search(fname):
frame = pd.read_csv(os.path.join(root, fname), names=columns,skiprows=1)
if fname.find("EDUCATION") != -1:
frame['CATEGORY'] = "EDUCATION"
elif fname.find("FINANCE") !=-1:
frame['CATEGORY'] = "FINANCE"
elif fname.find("ENTERTAINMENT") != -1:
frame['CATEGORY'] = "ENTERTAINMENT"
elif fname.find("FAMILY") != -1:
frame['CATEGORY'] = "FAMILY"
elif fname.find("GAME") != -1:
frame['CATEGORY'] = "GAME-ACTION"
elif fname.find("HEALTH") != -1:
frame['CATEGORY'] = "HEALTH AND FITNESS"
elif fname.find("LIFESTYLE") != -1:
frame['CATEGORY'] = "LIFESTYLE"
elif fname.find("MUSIC") != -1:
frame['CATEGORY'] = "MUSIC AND AUDIO"
frame['month'] = fname[5:7]
#frame['key'] = "file{}".format(i)
df2List.append(frame)
df2files.append(fname)
i += 1
df2 = pd.concat(df2List)
df2.reset_index(drop=True,inplace=True)
df2.rename(columns={1:"appTitle",2:"userName",3:"date",4:"score",5:"text"},inplace=True)
df2.dropna(axis=1,how="all",inplace=True)
df2['appTitle'] = df2['appTitle'].str.lower()
df2.drop_duplicates(subset=['appTitle','userName','date','score','text'],inplace=True)
df2.dropna(subset=['appTitle','text'],inplace=True)
df2
```
| github_jupyter |
```
# imports
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from sklearn.metrics import r2_score
import matplotlib.pyplot as plt
import numpy
from keras.optimizers import Adam
import keras
from matplotlib import pyplot
from keras.callbacks import EarlyStopping
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
import sklearn.metrics as sm
# we will read the dataset that we created in the previous exercise
df = pd.read_csv("DSDPartners_Data.csv", encoding='ISO-8859-1')
df.head()
#also try Operator Adjustments as potential target
target = 'PropOrderQty'
y = np.asarray(df[target])
#y = np.reshape(y,(y.shape[0],1))
X = df.drop(['CustStorItemTriadID','BaseorderID','Createdate','ModelUsed','RecDeliveryDate',
'ConversionFactor','Previous2DelDate','MaxScanDate','MaxShipDate','Reviewed','IncInAnom'],axis = 1).drop(target, axis=1).fillna(0)
#df.isna().sum()
#Establish training and testing data sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=314)
len(X_train), len(X_test), len(y_test), len(y_train)
# Create model
model = Sequential()
model.add(Dense(X.shape[1], activation="relu", input_dim=X.shape[1]))
model.add(Dense(X.shape[1]*.75, activation="relu"))
model.add(Dense(1))
# Compile model: The model is initialized with the Adam optimizer and then it is compiled.
model.compile(loss='mean_squared_error', optimizer=Adam(lr=1e-3, decay=1e-3 / 200))
# Patient early stopping
es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=200)
# Fit the model
history = model.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=100, batch_size=10, verbose=2, callbacks=[es])
# Calculate predictions
PredTestSet = model.predict(X_train)
PredValSet = model.predict(X_test)
print("Mean absolute error =", round(sm.mean_absolute_error(y_test, PredValSet), 2))
print("Mean squared error =", round(sm.mean_squared_error(y_test, PredValSet), 2))
print("Median absolute error =", round(sm.median_absolute_error(y_test, PredValSet), 2))
print("Explain variance score =", round(sm.explained_variance_score(y_test, PredValSet), 2))
print("R2 score =", round(sm.r2_score(y_test, PredValSet), 3))
pyplot.plot(history.history['loss'], label='train')
pyplot.plot(history.history['val_loss'], label='test')
pyplot.legend()
pyplot.show()
#Neural Network Model Accuracy
r_squared = r2_score(y_test,PredValSet)
#add RMSE,MSE, MAE
adjusted_r_squared = 1 - (1-r_squared)*(len(y)-1)/(len(y)-X.shape[1]-1)
r_squared,adjusted_r_squared
#Use the below code to see what percent where our predictions fall within 3, 4 or 5 off from actual
#As of Sunday night 98.6, 98.0 and 96.8 percent of predictions are +- 3,4 or 5 of actual.
#Biggest improvement is that the model gets it exactly right 83% of the time with new features, compared to 70% prior.
y_test_vals =np.reshape(y_test,(y_test.shape[0],))
Preds = np.reshape(PredValSet,(PredValSet.shape[0],))
compare = pd.DataFrame(np.array([y_test_vals, Preds]))
compare = np.transpose(compare)
compare.to_csv(r'compare.csv', index = False)
```
| github_jupyter |
# Introduction
## Why the FLUX pipeline?
The aim of the FLUX pipeline is to provide a standard implemented as a tutorial for using some of most common toolboxes to analyze full MEG datasets. The pipeline will be used for education and as well as provide a common framework for MEG data analysis.
This section will focus on [MNE-Python](https://mne.tools/stable/index.html). The core arguments for using MNE-Python are:
- The MNE-Python toolbox is supported by an active and dynamic group of developers ensuring that the latest analysis tools are available.
- The toolbox and pipeline are Open Source and henceforth compatible with Open Science approaches.
- The toolbox is based on Python which is free (as opposed to Matlab, for instance).
- Python is becoming the preferred programming language for novel data analyses approaches including a wealth of machine learning tools. Python modules are under constant development by a huge open sources community ensuring the availability of state-of-the-art data science tools.
## Target users
The target users are researchers new to MNE-Python as well as more advanced users seeking to standardize their analyses. The FLUX pipeline is particular geared towards cognitive neuroscientist with an interest in task-based paradigms. The developers of the pipeline are cognitve neuroscientists with a strong interest on brain oscillations and multivariate approaches.
## Using the pipeline in self-studies and education
The MNE Python toolbox has an excellent selection of [Tutorials](https://mne.tools/stable/auto_tutorials/index.html)
and [Examples](https://mne.tools/dev/auto_examples/index.html). Nevertheless, the options are so many that the learning curve for new users is very steep. The FLUX pipeline provides a set of procedures for what we consider best practice at this moment in time. Consistent with an Open Science approach, the FLUX pipeline provides a validated and documented approach for MEG data analysis. Each analysis step comes with explanations and illustrations. Furthermore, questions are embedded in the tutorials which will be useful for self-study or they can be used in educational settings.
We will not link back to sections on the MNE Python webpage as they will change over time; nevertheless, users are are strongly encouraged to consult the website as they develop their skills and insight.
To use the FLUX toolbox the following installations and downloads are required:
1. Install the Python environment; we recommend Anaconda3 which provide Jupyter notebooks and Spyder; see [anaconda.com](https://www.anaconda.com/)
2. Instal MNE Python: [instructions](https://mne.tools/dev/install/index.html). To make 3D rendering possible we recommend
> conda create --name=mne --channel=conda-forge mne
> conda install --name base nb_conda_kernels
3. Download the example datasets (see [FLUX website](https://neuosc.com/flux/))
4. Get the Jupyther notebooks from Github (see [FLUX website](https://neuosc.com/flux/); alternatively copy/paste the code from Github an execute using e.g. Spyder.
To create forward models based on T1 MRIs for source modeling [Freesurfer](https://surfer.nmr.mgh.harvard.edu/) is required.
## Preregistration and publication
Publication, example:
"The data were analyzed using the open source toolbox MNE Python v0.24.0 (Gramfort et al., 2013) following the standards defined in the FLUX Pipeline (Ferrrante et al., 2022)"
## References
Alexandre Gramfort, Martin Luessi, Eric Larson, Denis A. Engemann, Daniel Strohmeier, Christian Brodbeck, Roman Goj, Mainak Jas, Teon Brooks, Lauri Parkkonen, and Matti S. Hämäläinen. MEG and EEG data analysis with MNE-Python. Frontiers in Neuroscience, 7(267):1–13, 2013. doi:10.3389/fnins.2013.00267.
| github_jupyter |
```
# default_exp modeling.question_answering
#hide
%reload_ext autoreload
%autoreload 2
%matplotlib inline
import os
os.environ["TOKENIZERS_PARALLELISM"] = "false"
```
# modeling.question_answering
> This module contains custom models, loss functions, custom splitters, etc... for question answering tasks
```
#export
import ast
import torch
from transformers import *
from fastai.text.all import *
from blurr.utils import *
from blurr.data.core import *
from blurr.data.question_answering import *
from blurr.modeling.core import *
logging.set_verbosity_error()
#hide
import pdb
from nbdev.showdoc import *
from fastcore.test import *
from fastai import __version__ as fa_version
from torch import __version__ as pt_version
from transformers import __version__ as hft_version
print(f'Using pytorch {pt_version}')
print(f'Using fastai {fa_version}')
print(f'Using transformers {hft_version}')
#cuda
torch.cuda.set_device(1)
print(f'Using GPU #{torch.cuda.current_device()}: {torch.cuda.get_device_name()}')
```
## Question Answer
Given a document (context) and a question, the objective of these models is to predict the start and end token of the correct answer as it exists in the context.
Again, we'll use a subset of pre-processed SQUAD v2 for our purposes below.
```
# full
# squad_df = pd.read_csv('./data/task-question-answering/squad_cleaned.csv'); len(squad_df)
# sample
squad_df = pd.read_csv('./squad_sample.csv'); len(squad_df)
squad_df.head(2)
pretrained_model_name = 'bert-large-uncased-whole-word-masking-finetuned-squad'
hf_model_cls = BertForQuestionAnswering
hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(pretrained_model_name, model_cls=hf_model_cls)
# # here's a pre-trained roberta model for squad you can try too
# pretrained_model_name = "ahotrod/roberta_large_squad2"
# hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(pretrained_model_name,
# model_cls=AutoModelForQuestionAnswering)
# # here's a pre-trained xlm model for squad you can try too
# pretrained_model_name = 'xlm-mlm-ende-1024'
# hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(pretrained_model_name,
# model_cls=AutoModelForQuestionAnswering)
squad_df = squad_df.apply(partial(pre_process_squad, hf_arch=hf_arch, hf_tokenizer=hf_tokenizer), axis=1)
max_seq_len= 128
squad_df = squad_df[(squad_df.tokenized_input_len < max_seq_len) & (squad_df.is_impossible == False)]
#hide
squad_df.head(2)
vocab = list(range(max_seq_len))
# vocab = dict(enumerate(range(max_seq_len)));
# account for tokenizers that pad on right or left side
trunc_strat = 'only_second' if (hf_tokenizer.padding_side == 'right') else 'only_first'
before_batch_tfm = HF_QABeforeBatchTransform(hf_arch, hf_config, hf_tokenizer, hf_model,
max_length=max_seq_len,
truncation=trunc_strat,
tok_kwargs={ 'return_special_tokens_mask': True })
blocks = (
HF_TextBlock(before_batch_tfm=before_batch_tfm, input_return_type=HF_QuestionAnswerInput),
CategoryBlock(vocab=vocab),
CategoryBlock(vocab=vocab)
)
def get_x(x):
return (x.question, x.context) if (hf_tokenizer.padding_side == 'right') else (x.context, x.question)
dblock = DataBlock(blocks=blocks,
get_x=get_x,
get_y=[ColReader('tok_answer_start'), ColReader('tok_answer_end')],
splitter=RandomSplitter(),
n_inp=1)
dls = dblock.dataloaders(squad_df, bs=4)
len(dls.vocab), dls.vocab[0], dls.vocab[1]
dls.show_batch(dataloaders=dls, max_n=2)
```
### Training
Here we create a question/answer specific subclass of `HF_BaseModelCallback` in order to get all the start and end prediction. We also add here a new loss function that can handle multiple targets
```
#export
class HF_QstAndAnsModelCallback(HF_BaseModelCallback):
"""The prediction is a combination start/end logits"""
def after_pred(self):
super().after_pred()
self.learn.pred = (self.pred.start_logits, self.pred.end_logits)
```
And here we provide a custom loss function our question answer task, expanding on some techniques learned from here and here.
In fact, this new loss function can be used in many other multi-modal architectures, with any mix of loss functions. For example, this can be ammended to include the `is_impossible` task, as well as the start/end token tasks in the SQUAD v2 dataset.
```
#export
class MultiTargetLoss(Module):
"""Provides the ability to apply different loss functions to multi-modal targets/predictions"""
def __init__(self, loss_classes=[CrossEntropyLossFlat, CrossEntropyLossFlat], loss_classes_kwargs=[{}, {}],
weights=[1, 1], reduction='mean'):
loss_funcs = [ cls(reduction=reduction, **kwargs) for cls, kwargs in zip(loss_classes, loss_classes_kwargs) ]
store_attr(self=self, names='loss_funcs, weights')
self._reduction = reduction
# custom loss function must have either a reduction attribute or a reduction argument (like all fastai and
# PyTorch loss functions) so that the framework can change this as needed (e.g., when doing lear.get_preds
# it will set = 'none'). see this forum topic for more info: https://bit.ly/3br2Syz
@property
def reduction(self): return self._reduction
@reduction.setter
def reduction(self, v):
self._reduction = v
for lf in self.loss_funcs: lf.reduction = v
def forward(self, outputs, *targets):
loss = 0.
for i, loss_func, weights, output, target in zip(range(len(outputs)),
self.loss_funcs, self.weights,
outputs, targets):
loss += weights * loss_func(output, target)
return loss
def activation(self, outs):
acts = [ self.loss_funcs[i].activation(o) for i, o in enumerate(outs) ]
return acts
def decodes(self, outs):
decodes = [ self.loss_funcs[i].decodes(o) for i, o in enumerate(outs) ]
return decodes
model = HF_BaseModelWrapper(hf_model)
learn = Learner(dls,
model,
opt_func=partial(Adam, decouple_wd=True),
cbs=[HF_QstAndAnsModelCallback],
splitter=hf_splitter)
learn.loss_func=MultiTargetLoss()
learn.create_opt() # -> will create your layer groups based on your "splitter" function
learn.freeze()
```
Notice above how I had to define the loss function *after* creating the `Learner` object. I'm not sure why, but the `MultiTargetLoss` above prohibits the learner from being exported if I do.
```
learn.summary()
print(len(learn.opt.param_groups))
x, y_start, y_end = dls.one_batch()
preds = learn.model(x)
len(preds),preds[0].shape
#slow
learn.lr_find(suggest_funcs=[minimum, steep, valley, slide])
#slow
learn.fit_one_cycle(3, lr_max=1e-3)
```
### Showing results
Below we'll add in additional functionality to more intuitively show the results of our model.
```
#export
@typedispatch
def show_results(x:HF_QuestionAnswerInput, y, samples, outs, learner, skip_special_tokens=True,
ctxs=None, max_n=6, trunc_at=None, **kwargs):
hf_before_batch_tfm = get_blurr_tfm(learner.dls.before_batch)
hf_tokenizer = hf_before_batch_tfm.hf_tokenizer
res = L()
for sample, input_ids, start, end, pred in zip(samples, x, *y, outs):
txt = hf_tokenizer.decode(sample[0], skip_special_tokens=True)[:trunc_at]
ans_toks = hf_tokenizer.convert_ids_to_tokens(input_ids, skip_special_tokens=False)[start:end]
pred_ans_toks = hf_tokenizer.convert_ids_to_tokens(input_ids, skip_special_tokens=False)[int(pred[0]):int(pred[1])]
res.append((txt,
(start.item(),end.item()), hf_tokenizer.convert_tokens_to_string(ans_toks),
(int(pred[0]),int(pred[1])), hf_tokenizer.convert_tokens_to_string(pred_ans_toks)))
df = pd.DataFrame(res, columns=['text', 'start/end', 'answer', 'pred start/end', 'pred answer'])
display_df(df[:max_n])
return ctxs
learn.show_results(learner=learn, skip_special_tokens=True, max_n=2, trunc_at=500)
```
... and lets see how `Learner.blurr_predict` works with question/answering tasks
```
inf_df = pd.DataFrame.from_dict([{
'question': 'What did George Lucas make?',
'context': 'George Lucas created Star Wars in 1977. He directed and produced it.'
}],
orient='columns')
learn.blurr_predict(inf_df.iloc[0])
inf_df = pd.DataFrame.from_dict([
{
'question': 'What did George Lucas make?',
'context': 'George Lucas created Star Wars in 1977. He directed and produced it.'
}, {
'question': 'What year did Star Wars come out?',
'context': 'George Lucas created Star Wars in 1977. He directed and produced it.'
}, {
'question': 'What did George Lucas do?',
'context': 'George Lucas created Star Wars in 1977. He directed and produced it.'
}],
orient='columns')
learn.blurr_predict(inf_df)
inp_ids = hf_tokenizer.encode('What did George Lucas make?',
'George Lucas created Star Wars in 1977. He directed and produced it.')
hf_tokenizer.convert_ids_to_tokens(inp_ids, skip_special_tokens=False)[11:13]
```
Note that there is a bug currently in fastai v2 (or with how I'm assembling everything) that currently prevents us from seeing the decoded predictions and probabilities for the "end" token.
```
inf_df = pd.DataFrame.from_dict([{
'question': 'When was Star Wars made?',
'context': 'George Lucas created Star Wars in 1977. He directed and produced it.'
}],
orient='columns')
test_dl = dls.test_dl(inf_df)
inp = test_dl.one_batch()[0]['input_ids']
probs, _, preds = learn.get_preds(dl=test_dl, with_input=False, with_decoded=True)
hf_tokenizer.convert_ids_to_tokens(inp.tolist()[0],
skip_special_tokens=False)[torch.argmax(probs[0]):torch.argmax(probs[1])]
```
We can unfreeze and continue training like normal
```
learn.unfreeze()
#slow
learn.fit_one_cycle(3, lr_max=slice(1e-7, 1e-4))
learn.recorder.plot_loss()
learn.show_results(learner=learn, max_n=2, trunc_at=100)
learn.blurr_predict(inf_df.iloc[0])
preds, pred_classes, probs = zip(*learn.blurr_predict(inf_df.iloc[0]))
preds
inp_ids = hf_tokenizer.encode('When was Star Wars made?',
'George Lucas created Star Wars in 1977. He directed and produced it.')
hf_tokenizer.convert_ids_to_tokens(inp_ids, skip_special_tokens=False)[int(preds[0][0]):int(preds[0][1])]
```
### Inference
Note that I had to replace the loss function because of the above-mentioned issue to exporting the model with the `MultiTargetLoss` loss function. After getting our inference learner, we put it back and we're good to go!
```
export_name = 'q_and_a_learn_export'
learn.loss_func = CrossEntropyLossFlat()
learn.export(fname=f'{export_name}.pkl')
inf_learn = load_learner(fname=f'{export_name}.pkl')
inf_learn.loss_func = MultiTargetLoss()
inf_df = pd.DataFrame.from_dict([
{
'question': 'What did George Lucas make?',
'context': 'George Lucas created Star Wars in 1977. He directed and produced it.'
}, {
'question': 'What year did Star Wars come out?',
'context': 'George Lucas created Star Wars in 1977. He directed and produced it.'
}, {
'question': 'What did George Lucas do?',
'context': 'George Lucas created Star Wars in 1977. He directed and produced it.'
}],
orient='columns')
inf_learn.blurr_predict(inf_df)
inp_ids = hf_tokenizer.encode('What did George Lucas make?',
'George Lucas created Star Wars in 1977. He directed and produced it.')
hf_tokenizer.convert_ids_to_tokens(inp_ids, skip_special_tokens=False)[11:13]
```
... and onnx works here too
```
# #slow
# learn.blurr_to_onnx(export_name)
# #slow
# onnx_inf = blurrONNX(export_name)
# #slow
# onnx_inf.predict(inf_df)
```
## Cleanup
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
<img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# Gmail - Schedule mailbox cleaning
<a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Gmail/Gmail_Schedule_mailbox_cleaning.ipynb" target="_parent"><img src="https://img.shields.io/badge/-Open%20in%20Naas-success?labelColor=000000&logo=data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iMTAyNHB4IiBoZWlnaHQ9IjEwMjRweCIgdmlld0JveD0iMCAwIDEwMjQgMTAyNCIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayIgdmVyc2lvbj0iMS4xIj4KIDwhLS0gR2VuZXJhdGVkIGJ5IFBpeGVsbWF0b3IgUHJvIDIuMC41IC0tPgogPGRlZnM+CiAgPHRleHQgaWQ9InN0cmluZyIgdHJhbnNmb3JtPSJtYXRyaXgoMS4wIDAuMCAwLjAgMS4wIDIyOC4wIDU0LjUpIiBmb250LWZhbWlseT0iQ29tZm9ydGFhLVJlZ3VsYXIsIENvbWZvcnRhYSIgZm9udC1zaXplPSI4MDAiIHRleHQtZGVjb3JhdGlvbj0ibm9uZSIgZmlsbD0iI2ZmZmZmZiIgeD0iMS4xOTk5OTk5OTk5OTk5ODg2IiB5PSI3MDUuMCI+bjwvdGV4dD4KIDwvZGVmcz4KIDx1c2UgaWQ9Im4iIHhsaW5rOmhyZWY9IiNzdHJpbmciLz4KPC9zdmc+Cg=="/></a>
**Tags:** #gmail #productivity
## Input
### Import libraries
```
import naas
from naas_drivers import email
import pandas as pd
import numpy as np
import plotly.express as px
```
### Account credentials
```
username = "naas.sanjay22@gmail.com"
password = "atsuwkylwfhucugw"
smtp_server = "imap.gmail.com"
box = "INBOX"
```
Note: You need to create an application password following this procedure - https://support.google.com/mail/answer/185833?hl=en
## Model
### Setting the scheduler
```
naas.scheduler.add(recurrence="0 9 * * *") # Scheduler set for 9 am
```
### Connect to email box
```
emails = naas_drivers.email.connect(username,
password,
username,
smtp_server)
```
### Get email list
```
dataframe = emails.get(criteria="seen")
dataframe
```
### Creating dataframe and inserting values
```
sender_name = []
sender_email = []
for df in dataframe["from"]:
sender_name.append(df['name'])
sender_email.append(df['email'])
result = pd.DataFrame(columns = ['SENDER_NAME','SENDER_EMAIL','COUNT','PERCENTAGE'])
name_unique = np.unique(sender_name)
email_unique = np.unique(sender_email)
total_email = len(emails.get(criteria="seen")) + len(emails.get(criteria="unseen"))
c = 0
for i in np.unique(sender_name):
new_row = {'SENDER_NAME':i,'SENDER_EMAIL':sender_email[c],'COUNT':sender_name.count(i),'PERCENTAGE':round(((sender_name.count(i))/total_email)*100)}
result = result.append(new_row, ignore_index=True)
c+=1
result
```
### Email graph plot
```
fig = px.bar(x=result['COUNT'], y=result['SENDER_NAME'], orientation='h')
fig.show()
```
## Output
### Deleting using email id
```
d_email = "notifications@naas.ai" # email id to be deleted
data_from = dataframe['from']
data_uid = dataframe['uid']
uid = []
```
### Updating the uid values
```
for i in range(len(dataframe)):
if data_from[i]['email'] == d_email:
uid.append(data_uid[i])
print(uid)
```
### Deleting the emails
```
for i in uid:
attachments = emails.set_flag(i, "DELETED", True)
```
### Showing the upated email list
```
dataframe = emails.get(criteria="seen")
dataframe
```
| github_jupyter |
# k-Nearest Neighbor (kNN) exercise
*Complete and hand in this completed worksheet (including its outputs and any supporting code outside of the worksheet) with your assignment submission. For more details see the [assignments page](http://vision.stanford.edu/teaching/cs231n/assignments.html) on the course website.*
The kNN classifier consists of two stages:
- During training, the classifier takes the training data and simply remembers it
- During testing, kNN classifies every test image by comparing to all training images and transfering the labels of the k most similar training examples
- The value of k is cross-validated
In this exercise you will implement these steps and understand the basic Image Classification pipeline, cross-validation, and gain proficiency in writing efficient, vectorized code.
```
# Run some setup code for this notebook.
import random
import numpy as np
from cs231n.data_utils import load_CIFAR10
import matplotlib.pyplot as plt
# This is a bit of magic to make matplotlib figures appear inline in the notebook
# rather than in a new window.
%matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
%load_ext autoreload
%autoreload 2
# Load the raw CIFAR-10 data.
cifar10_dir = 'cs231n/datasets/cifar-10-batches-py'
X_train, y_train, X_test, y_test = load_CIFAR10(cifar10_dir)
# As a sanity check, we print out the size of the training and test data.
print 'Training data shape: ', X_train.shape
print 'Training labels shape: ', y_train.shape
print 'Test data shape: ', X_test.shape
print 'Test labels shape: ', y_test.shape
# Visualize some examples from the dataset.
# We show a few examples of training images from each class.
classes = ['plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
num_classes = len(classes)
samples_per_class = 7
for y, cls in enumerate(classes):
idxs = np.flatnonzero(y_train == y)
idxs = np.random.choice(idxs, samples_per_class, replace=False)
for i, idx in enumerate(idxs):
plt_idx = i * num_classes + y + 1
plt.subplot(samples_per_class, num_classes, plt_idx)
plt.imshow(X_train[idx].astype('uint8'))
plt.axis('off')
if i == 0:
plt.title(cls)
plt.show()
# Subsample the data for more efficient code execution in this exercise
num_training = 5000
mask = range(num_training)
X_train = X_train[mask]
y_train = y_train[mask]
num_test = 500
mask = range(num_test)
X_test = X_test[mask]
y_test = y_test[mask]
# Reshape the image data into rows
X_train = np.reshape(X_train, (X_train.shape[0], -1))
X_test = np.reshape(X_test, (X_test.shape[0], -1))
print X_train.shape, X_test.shape
from cs231n.classifiers import KNearestNeighbor
# Create a kNN classifier instance.
# Remember that training a kNN classifier is a noop:
# the Classifier simply remembers the data and does no further processing
classifier = KNearestNeighbor()
classifier.train(X_train, y_train)
```
We would now like to classify the test data with the kNN classifier. Recall that we can break down this process into two steps:
1. First we must compute the distances between all test examples and all train examples.
2. Given these distances, for each test example we find the k nearest examples and have them vote for the label
Lets begin with computing the distance matrix between all training and test examples. For example, if there are **Ntr** training examples and **Nte** test examples, this stage should result in a **Nte x Ntr** matrix where each element (i,j) is the distance between the i-th test and j-th train example.
First, open `cs231n/classifiers/k_nearest_neighbor.py` and implement the function `compute_distances_two_loops` that uses a (very inefficient) double loop over all pairs of (test, train) examples and computes the distance matrix one element at a time.
```
# Open cs231n/classifiers/k_nearest_neighbor.py and implement
# compute_distances_two_loops.
# Test your implementation:
dists = classifier.compute_distances_two_loops(X_test)
print dists.shape
# We can visualize the distance matrix: each row is a single test example and
# its distances to training examples
plt.imshow(dists, interpolation='none')
plt.show()
```
**Inline Question #1:** Notice the structured patterns in the distance matrix, where some rows or columns are visible brighter. (Note that with the default color scheme black indicates low distances while white indicates high distances.)
- What in the data is the cause behind the distinctly bright rows?
- What causes the columns?
**Your Answer**: *fill this in.*
```
# Now implement the function predict_labels and run the code below:
# We use k = 1 (which is Nearest Neighbor).
y_test_pred = classifier.predict_labels(dists, k=1)
# Compute and print the fraction of correctly predicted examples
num_correct = np.sum(y_test_pred == y_test)
accuracy = float(num_correct) / num_test
print 'Got %d / %d correct => accuracy: %f' % (num_correct, num_test, accuracy)
```
You should expect to see approximately `27%` accuracy. Now lets try out a larger `k`, say `k = 5`:
```
y_test_pred = classifier.predict_labels(dists, k=5)
num_correct = np.sum(y_test_pred == y_test)
accuracy = float(num_correct) / num_test
print 'Got %d / %d correct => accuracy: %f' % (num_correct, num_test, accuracy)
```
You should expect to see a slightly better performance than with `k = 1`.
```
# Now lets speed up distance matrix computation by using partial vectorization
# with one loop. Implement the function compute_distances_one_loop and run the
# code below:
dists_one = classifier.compute_distances_one_loop(X_test)
# To ensure that our vectorized implementation is correct, we make sure that it
# agrees with the naive implementation. There are many ways to decide whether
# two matrices are similar; one of the simplest is the Frobenius norm. In case
# you haven't seen it before, the Frobenius norm of two matrices is the square
# root of the squared sum of differences of all elements; in other words, reshape
# the matrices into vectors and compute the Euclidean distance between them.
difference = np.linalg.norm(dists - dists_one, ord='fro')
print 'Difference was: %f' % (difference, )
if difference < 0.001:
print 'Good! The distance matrices are the same'
else:
print 'Uh-oh! The distance matrices are different'
# Now implement the fully vectorized version inside compute_distances_no_loops
# and run the code
dists_two = classifier.compute_distances_no_loops(X_test)
# check that the distance matrix agrees with the one we computed before:
difference = np.linalg.norm(dists - dists_two, ord='fro')
print 'Difference was: %f' % (difference, )
if difference < 0.001:
print 'Good! The distance matrices are the same'
else:
print 'Uh-oh! The distance matrices are different'
# Let's compare how fast the implementations are
def time_function(f, *args):
"""
Call a function f with args and return the time (in seconds) that it took to execute.
"""
import time
tic = time.time()
f(*args)
toc = time.time()
return toc - tic
two_loop_time = time_function(classifier.compute_distances_two_loops, X_test)
print 'Two loop version took %f seconds' % two_loop_time
one_loop_time = time_function(classifier.compute_distances_one_loop, X_test)
print 'One loop version took %f seconds' % one_loop_time
no_loop_time = time_function(classifier.compute_distances_no_loops, X_test)
print 'No loop version took %f seconds' % no_loop_time
# you should see significantly faster performance with the fully vectorized implementation
```
### Cross-validation
We have implemented the k-Nearest Neighbor classifier but we set the value k = 5 arbitrarily. We will now determine the best value of this hyperparameter with cross-validation.
```
num_folds = 5
k_choices = [1, 3, 5, 8, 10, 12, 15, 20, 50, 100]
X_train_folds = []
y_train_folds = []
################################################################################
# TODO: #
# Split up the training data into folds. After splitting, X_train_folds and #
# y_train_folds should each be lists of length num_folds, where #
# y_train_folds[i] is the label vector for the points in X_train_folds[i]. #
# Hint: Look up the numpy array_split function. #
################################################################################
pass
# split self.X_train to 5 folds
avg_size = int(X_train.shape[0] / num_folds) # will abandon the rest if not divided evenly.
for i in range(num_folds):
X_train_folds.append(X_train[i * avg_size : (i+1) * avg_size])
y_train_folds.append(y_train[i * avg_size : (i+1) * avg_size])
################################################################################
# END OF YOUR CODE #
################################################################################
# A dictionary holding the accuracies for different values of k that we find
# when running cross-validation. After running cross-validation,
# k_to_accuracies[k] should be a list of length num_folds giving the different
# accuracy values that we found when using that value of k.
k_to_accuracies = {}
################################################################################
# TODO: #
# Perform k-fold cross validation to find the best value of k. For each #
# possible value of k, run the k-nearest-neighbor algorithm num_folds times, #
# where in each case you use all but one of the folds as training data and the #
# last fold as a validation set. Store the accuracies for all fold and all #
# values of k in the k_to_accuracies dictionary. #
################################################################################
pass
for k in k_choices:
accuracies = []
print k
for i in range(num_folds):
X_train_cv = np.vstack(X_train_folds[0:i] + X_train_folds[i+1:])
y_train_cv = np.hstack(y_train_folds[0:i] + y_train_folds[i+1:])
X_valid_cv = X_train_folds[i]
y_valid_cv = y_train_folds[i]
classifier.train(X_train_cv, y_train_cv)
dists = classifier.compute_distances_no_loops(X_valid_cv)
accuracy = float(np.sum(classifier.predict_labels(dists, k) == y_valid_cv)) / y_valid_cv.shape[0]
accuracies.append(accuracy)
k_to_accuracies[k] = accuracies
################################################################################
# END OF YOUR CODE #
################################################################################
# Print out the computed accuracies
for k in sorted(k_to_accuracies):
for accuracy in k_to_accuracies[k]:
print 'k = %d, accuracy = %f' % (k, accuracy)
# plot the raw observations
for k in k_choices:
accuracies = k_to_accuracies[k]
plt.scatter([k] * len(accuracies), accuracies)
# plot the trend line with error bars that correspond to standard deviation
accuracies_mean = np.array([np.mean(v) for k,v in sorted(k_to_accuracies.items())])
accuracies_std = np.array([np.std(v) for k,v in sorted(k_to_accuracies.items())])
plt.errorbar(k_choices, accuracies_mean, yerr=accuracies_std)
plt.title('Cross-validation on k')
plt.xlabel('k')
plt.ylabel('Cross-validation accuracy')
plt.show()
# Based on the cross-validation results above, choose the best value for k,
# retrain the classifier using all the training data, and test it on the test
# data. You should be able to get above 28% accuracy on the test data.
best_k = 10
classifier = KNearestNeighbor()
classifier.train(X_train, y_train)
y_test_pred = classifier.predict(X_test, k=best_k)
# Compute and display the accuracy
num_correct = np.sum(y_test_pred == y_test)
accuracy = float(num_correct) / num_test
print 'Got %d / %d correct => accuracy: %f' % (num_correct, num_test, accuracy)
```
| github_jupyter |
```
import torch
import torch.nn as nn
from torch2trt import torch2trt
from torch.nn.utils import weight_norm
class TensorCache(nn.Module):
def __init__(self, tensor):
super(TensorCache, self).__init__()
# self.register_buffer('cache', tensor)
self.cache = tensor
def forward(self, x):
# assert x.size() == self.cache[:,:,0:1].size()
cache_update = torch.cat((self.cache[:,:,1:], x.detach()), dim=2)
self.cache[:,:,:] = cache_update
return self.cache
tc = TensorCache(torch.zeros(1,1,10))
tc_script = torch.jit.script(tc)
# tc_trace = torch.jit.trace(tc, torch.tensor([[[0.0]]))
print('Original:')
for inp in [1.,2.,3.,4.]:
print(tc(torch.tensor([[[inp]]])))
print()
print('Convert using script:')
for inp in [1.,2.,3.,4.]:
print(tc_script(torch.tensor([[[inp]]])))
# print('Original:')
# for inp in [1.,2.,3.,4.]:
# print(tc(torch.tensor([[[inp]]])))
# print()
# print('Convert using script:')
# for inp in [1.,2.,3.,4.]:
# print(tc_script(torch.tensor([[[inp]]])))
# print('')
# print('Convert using trace:')
# for inp in [1.,2.,3.,4.]:
# print(tc_trace(torch.tensor([[[inp]]])))
ex_in = torch.tensor([[[0.0]]])
ex_in.size()
tc_trt = torch2trt(tc, ex_in)
t = torch.zeros(10)
torch.cat((t[1:], torch.tensor([1])))
class TemporalInferenceBlock(nn.Module):
# class TemporalInferenceBlock(torch.jit.ScriptModule):
def __init__(self, n_inputs, n_outputs, kernel_size, stride, dilation, batch_size=1):
super(TemporalInferenceBlock, self).__init__()
self.in_ch, self.k, self.d = n_inputs, kernel_size, dilation
self.conv1 = weight_norm(nn.Conv1d(n_inputs, n_outputs, kernel_size,
stride=stride, padding=0, dilation=dilation))
self.relu1 = nn.ReLU()
self.conv2 = weight_norm(nn.Conv1d(n_outputs, n_outputs, kernel_size,
stride=stride, padding=0, dilation=dilation))
self.relu2 = nn.ReLU()
self.batch_size = batch_size
# self.cache1 = torch.jit.script(TensorCache(torch.zeros(
# batch_size,
# self.conv1.in_channels,
# (self.conv1.kernel_size[0]-1)*self.conv1.dilation[0] + 1
# )))
# self.cache2 = torch.jit.script(TensorCache(torch.zeros(
# batch_size,
# self.conv2.in_channels,
# (self.conv2.kernel_size[0]-1)*self.conv2.dilation[0] + 1
# )))
self.cache1 = torch.zeros(
batch_size,
self.conv1.in_channels,
(self.conv1.kernel_size[0]-1)*self.conv1.dilation[0] + 1
)
self.cache2 = torch.zeros(
batch_size,
self.conv2.in_channels,
(self.conv2.kernel_size[0]-1)*self.conv2.dilation[0] + 1
)
self.stage1 = nn.Sequential(self.conv1, self.relu1)
self.stage2 = nn.Sequential(self.conv2, self.relu2)
self.downsample = nn.Conv1d(n_inputs, n_outputs, 1) if n_inputs != n_outputs else nn.Identity()
self.relu = nn.ReLU()
self.init_weights()
def init_weights(self):
self.conv1.weight.data.normal_(0, 0.01)
self.conv2.weight.data.normal_(0, 0.01)
if isinstance(self.downsample, nn.modules.conv.Conv1d):
self.downsample.weight.data.normal_(0, 0.01)
def reset_cache(self):
self.cache1.zero_cache()
self.cache2.zero_cache()
# def forward(self, x):
# '''
# x is of shape (B, CH, 1)
# '''
# # out = self.stage1(self.cache1(x)[:x.size()[0], :, :])
# # out = self.stage2(self.cache2(out)[:x.size()[0], :, :])
# # out1 = self.stage1(self.cache1(x))
# # out2 = self.stage2(self.cache2(out1))
# out1 = self.relu1(self.conv1(self.cache1(x)))
# out2 = self.relu2(self.conv2(self.cache2(out1)))
# # self.cache1.zero_cache()
# # out1 = self.cache1()
# # self.cache2.zero_cache()
# # out2 = self.cache2()
# res = self.downsample(x)
# out = self.relu(out2 + res)
# # print(f'\t res shape: {res.size()}')
# # print(f'x: {x} \n c1: {self.cache1.cache} \n out1: {out1} \n c2: {self.cache2.cache} \n out2: {out2} \n \n')
# return x, self.cache1.cache, out1, self.cache2.cache, out2, res, out
# @torch.jit.script_method
def forward(self, x):
'''
x is of shape (B, CH, 1)
'''
# out = self.stage1(self.cache1(x)[:x.size()[0], :, :])
# out = self.stage2(self.cache2(out)[:x.size()[0], :, :])
cache_update = torch.cat((self.cache1[:,:,1:], x.detach()), dim=2)
self.cache1[:,:,:] = cache_update
out1 = self.stage1(self.cache1)
cache_update = torch.cat((self.cache2[:,:,1:], out1), dim=2)
self.cache2[:,:,:] = cache_update
out2 = self.stage2(self.cache2)
res = self.downsample(x)
out = self.relu(out2 + res)
# print(f'\t res shape: {res.size()}')
# print(f'x: {x} \n c1: {self.cache1.cache} \n out1: {out1} \n c2: {self.cache2.cache} \n out2: {out2} \n \n')
return x, self.cache1, out1, self.cache2, out2, res, out
tblock = TemporalInferenceBlock(1,1,7,1,1,1)
tblock.eval()
tblock.cuda()
tblock_script = torch.jit.script(tblock)
tblock_trace = torch.jit.trace(tblock, torch.tensor([[[0]]]).cuda())
print('Original:')
for inp in [1,2,3,4]:
print(tc(torch.tensor([[[inp]]])))
print()
print('Convert using script:')
for inp in [1,2,3,4]:
print(tc_script(torch.tensor([[[inp]]])))
```
| github_jupyter |
<img src="./pictures/DroneApp_logo.png" style="float:right; max-width: 180px; display: inline" alt="INSA" /></a>
<img src="./pictures/logo_sizinglab.png" style="float:right; max-width: 100px; display: inline" alt="INSA" /></a>
# Application of First Monotonicity Principle to the optimization of MRAV
*Created by Aitor Ochotorena (INSA Toulouse), Toulouse, France.*
Based on the differentiability of continuous mathematical functions, in this Notebook we present a guide to **reduce the excess of constraints** in optimization problems.
The standard expression of an optimization problem has the following form:
<math>\begin{align}
&\underset{\mathbf{x}}{\operatorname{minimize}}& & f(\mathbf{x}) \\
&\operatorname{subject\ to}
& &g_i(\mathbf{x}) \leq 0, \quad i = 1, \dots, m \\
&&&h_i(\mathbf{x}) = 0, \quad i = 1, \dots, p,
\end{align}</math>
where $x \in \mathbb{R}^n$ is the optimization variable, the functions $f, g_1, \ldots, g_m$ are convex, and the functions $h_1, \ldots, h_p$ are equality constraints.
In this notation, the function $f$ is the objective function of the problem, and the functions $g_i$ and $h_i$are referred to as the constraint functions.
**Find out which set of inequality constraints can be turned to equality ones has an enormous importance to reduce the complexity of the problem and the calculation time.**
The applicacion of the 'First Monotonicity Principle',
> P. Y. Papalambros, D. J. Wilde, Principles of Optimal Design, Cambridge University Press, 2018.
permits to evaluate if the objective meets the minimum values w.r.t. the considered variable when the constraint acts an equality. We refer to such constraints as *active constraints* and they are identified by studying the monotonicity behaviour of both the objective and the constraint. In a well constrainted objective function, every (strictly) increasing (decreasing) variable is bounded below (above) by at least one active constraint. For a well constrained minimization problem, there exist at least one $x$ that satisfies the optimality conditions:
<math>\begin{align}
$\displaystyle (\frac{\partial{ f}}{\partial{ x_i}})_*+\sum_{j}\mu_j(\frac{\partial{ g_j}}{\partial{ x_i}})_*=0$
\end{align}</math>
where $\mu_j \geq 0$.
In the case in which the sign of a variable in the objective function is
uniquely opposite to the sign of the same variable for a single constraint, this
constraint can be turned to active and as equality. If the sign of the variable
is opposite to that of the objective function in several cosntraints, then no
evident statement can be given and in that case, the constraints will be left
as inequalities.

*Representative plot of an optimization problem with three constraints. Two of them: $g_1$ and $g_2$ are acting as active constraints, since they bound the function objective as equality.*
## 1. Import Sizing Code from a .py code
Here Python reads the sizing code with all equations used for the drone saved in the folder :
``` '.\SizingCode'```. Our file is called: ``` DroneSystem.py```.
Design variables are defined as symbols using the symbolic calculation of Sympy:
**(This part is specific for every sizing code)**
```
import sys
sys.path.insert(0, 'SizingCode')
from DroneSystem import *
```
## 2. Problem definition
Once the equations are imported from the .py file, we define here the main parameters for the optimization problem: objective ```Obj```, design variables ```Vars```, constraints ```Const``` and bounds ```bounds```. We work in forwards on with symbolic mathematics (SymPy):
**(This part is specific for every sizing code)**
- Objective:
```
Obj=Mtotal_final
```
- Design Variables:
```
Vars=[ Mtotal,ND,Tmot,Ktmot,P_esc,V_bat,C_bat,beta, J, D_ratio, Lbra,Dout]
```
- Constraints:
```
Const=[
-Tmot_max+Qpro_max ,
-Tmot_max+Qpro_cl,
-Tmot+Qpro_hover,
-V_bat+Umot_max,
-V_bat+Umot_cl,
-V_bat+Vesc,
-V_bat*Imax+Umot_max*Imot_max*Npro/0.95,
-V_bat*Imax+Umot_cl*Imot_cl*Npro/0.95,
-P_esc+P_esc_max,
-P_esc+P_esc_cl,
-J*n_pro_cl*Dpro+V_cl,
+J*n_pro_cl*Dpro-V_cl-0.05,
-NDmax+ND,
-NDmax+n_pro_cl*Dpro,
-Lbra+Dpro/2/(math.sin(pi/Narm)),
(-Sigma_max+Tpro_max*Lbra/(pi*(Dout**4-(D_ratio*Dout)**4)/(32*Dout)))
]
```
- Bounds:
```
bounds=[(0,100),#M_total
(0,105000/60*.0254),#ND
(0.01,10),#Tmot
(0,1),#Ktmot
(0,1500),#P_esc
(0,150),#V_bat
(0,20*3600),#C_bat
(0.3,0.6),#beta
(0,0.5),#J
(0,0.99),#D_ratio
(0.01,1),#Lb
(0.001,0.1),#Dout
(1,15),#Nred
]
```
## 3. Monotonicity algorithm
The next step is to evaluate the monotonicity of the functions. This will be done through the study of the differentiability of the functions. We will follow this procedure: A constraint is passed to ```is_increasing()``` or ```is_decreasing()```, which return a predicate of ```lambda x: x > 0``` or ```lambda x: x < 0``` respectively. This method calls ```compute_min_and_max``` which differentiates the constraint with respect to the desired variable, creates a series of random points defined within the bounds and substitute such values into the derivative of the constraint. If the predicate match the output, this method returns a True
To run the design of experiments satisfactorily, update pyDOE: `pip install --upgrade pyDOE`:
This algorithm is saved under the file `Monotonicity.ipynb` :
**(This part is reusable)**
```
# Note the python import here
import reuse, sys
# This is the Ipython hook
sys.meta_path.append(reuse.NotebookFinder())
import Monotonicity
```
## 3. Construction of table of Monotonicity
For each constraint and variable we will study the monotonicity behaviour calling the previous methods defined. If the constraint has an increasing behaviour, a ```+``` will be printed, in case where it is decreasing ```-```, and in the case where both increases and decreases, a ```?``` is displayed. The objective will be studied as well.
**(This part is reusable)**
```
import pandas as pd
M=[["" for x in Vars] for y in Const];
ObjVector=["" for x in Vars];
print('Monotonicity Analysis for the constraints w.r.t. the following variables:')
for Cnumber,C in enumerate(Const): #loop for constraints
print('Constant %d out of %d' %(Cnumber+1,len(Const)))
for Anumber,A in enumerate(Vars): #loop for variables
#print(C,A,bounds,Vars)
if Monotonicity.is_increasing(C,A,bounds,Vars)=='ZERO':
M[Cnumber][Anumber]=' '
elif Monotonicity.is_increasing(C,A,bounds,Vars):
print('* %s is increasing' %A)
M[Cnumber][Anumber]='+'
elif Monotonicity.is_decreasing(C,A,bounds,Vars):
print('* %s is decreasing' %A)
M[Cnumber][Anumber]='-'
else: M[Cnumber][Anumber]='?'
print('\n')
print('Monotonicity Analysis for the objective w.r.t. the following variables:')
for Anumber,A in enumerate(Vars):
if Monotonicity.is_increasing(Obj,A,bounds,Vars)=='ZERO':
ObjVector[Anumber]=' '
elif Monotonicity.is_increasing(Obj,A,bounds,Vars):
print('* %s is increasing' %A)
ObjVector[Anumber]='+'
elif Monotonicity.is_decreasing(Obj,A,bounds,Vars):
print('* %s is decreasing' %A)
ObjVector[Anumber]='-'
else: ObjVector[Anumber]='? '
M.append(ObjVector)
```
Create from np.array a DataFrame:
```
import pandas as pd
indexcol=[i for i in range(len(Const))]
indexcol.append('Objective')
pd.DataFrame(M, columns=Vars, index=indexcol)
```
Last point is the comparison of every constraint's variable to objective's oneActive constraints decision:
```
for index,objvalue in enumerate(ObjVector):
counter=0;
for j,constvalue in enumerate([i[index] for i in M]):
# print(objvalue);
# print(constvalue)
if objvalue=='+' and constvalue=='-':
counter+=1;
x=j #to save the constraint
elif objvalue=='-' and constvalue=='+':
counter+=1;
x=j; #to save the constraint
if counter==1:
print('Const %d w.r.t. %s can be eliminated'%(x,Vars[index]))
```
<a id='section_3'></a>
| github_jupyter |
<center>
<img src="https://tensorflowkorea.files.wordpress.com/2020/12/4.-e18492e185a9e186abe1848ce185a1-e18480e185a9e186bce18487e185aee18492e185a1e18482e185b3e186ab-e18486e185a5e18489e185b5e186abe18485e185a5e18482e185b5e186bce18483e185b5e186b8e18485e185a5e.png?w=972" width="200" height="200"><br>
</center>
# chaper.5 트리 알고리즘
## 05-1 결정 트리
### - 로지스틱 회귀로 와인 분류하기
와인 데이터를 먼저 불러오자.
데이터 출처 : https://github.com/rickiepark/hg-mldl
```
import pandas as pd
wine = pd.read_csv('/home/jaeyoon89/hg-mldl/wine.csv')
wine.head()
```
데이터를 잘 불러왔다 처음의 열 3개는 각각 알코올 도수,당도,PH 값을 나타낸다. 네번째 열은 타깃값으로 0이면 레드와인 1이면 화이트 와인이다. 레드 와인과 화이트 와인을 구분하는 이진 분류 문제이고, 화이트 와인이 양성 클래스이다. 즉 전체 와인에서 화이트 와인을 골라내는 문제이다. 로지스틱 회귀 모델을 바로 훈련하기 전에 판다스 데이터프레임의 유용한 메서드 2개를 먼저 알아보자.
먼저 info() 메서드이다. 이 메서드는 데이터프레임의 각 열의 데이터 타입과 누락된 데이터가 있는지 확인하는데 유용하다.
```
wine.info()
```
출력 결과를 보면 총 6497개의 샘플이 있고 누락된 값은 없는 것 같다.
다음 알아볼 메서드는 describe() 메서드이다. 이 메서드는 열에 대한 간략한 통계를 출력한다.
```
wine.describe()
```
평균, 표준편차, 최소, 최대값과 중간값, 1사분위수, 3사분위수를 알려준다.
여기서 알 수 있는 것은 알코올 도수와 당도, PH 값의 스케일이 다르다는 것이다. 이전에 배웠던 사이킷런의 StandardScaler 클래스를 사용해 특성을 표준화하자. 그전에 먼저 데이터프레임을 넘파이 배열로 바꾸고 훈련 세트와 테스트 세트로 나누자.
```
data = wine[['alcohol', 'sugar', 'pH']].to_numpy()
target = wine['class'].to_numpy()
```
그 다음으로 훈련 세트와 테스트 세트로 나누자.
```
from sklearn.model_selection import train_test_split
train_input, test_input, train_target, test_target = train_test_split(
data, target, test_size=0.2, random_state=42)
```
train_test_split() 함수는 설정값을 지정하지 않으면 25%를 테스트 세트로 지정한다. 샘플개수가 충분히 많으므로 20% 정도만 테스트 세트로 나누었다. 코드의 test_size=0.2가 이런 의미이다.
```
print(train_input.shape, test_input.shape)
```
이제 StandardScaler 클래스를 사용해 훈련 세트를 전처리 해보자.
```
from sklearn.preprocessing import StandardScaler
ss = StandardScaler()
ss.fit(train_input)
train_scaled = ss.transform(train_input)
test_scaled = ss.transform(test_input)
```
이제 표준점수로 변환된 train_scaled와 test_scaled를 사용해 로지스틱 회귀 모델을 훈련하자.
```
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
lr.fit(train_scaled, train_target)
print(lr.score(train_scaled, train_target))
print(lr.score(test_scaled, test_target))
```
점수가 높지 않다. 훈련 세트와 테스트 세트의 점수가 모두 낮아서 모델이 과소적합이라 볼 수 있다.이 문제를 해결하기 위해 규제 매개 변수 C의 값을 바꾸던지 solver 매개변수에서 다른 알고리즘을 선택할 수도 있다. 또한 다항 특성을 만들어 추가할 수도 있다.
### - 설명하기 쉬운 모델과 어려운 모델
로지스틱 회귀가 학습한 계수와 절편을 출력해 보자.
```
print(lr.coef_, lr.intercept_)
```
사실 이 모델이 왜 저런 계수 값을 학습했는지 정확히 이해하기 어렵다. 아마도 알코올 도수와 당도가 높을수록 화이트 와인일 가능성이 높고, PH가 높을수록 레드 와인일 가능성이 높은 것 같다. 하지만 정확히 이 숫자가 어떤 의미인지 설명하긴 어렵다. 더군다나 다항 특성을 추가한다면 설명하기가 더 어려울 것이다. 대부분 러닝머신 모델은 이렇게 학습의 결과를 설명하기 어렵다. 쉬운 방법으로 설명할 수 있는 모델을 알아보자.
### - 결정 트리
결정 트리 모델이 이유를 설명하기가 쉽다. 결정 트리 모델은 스무고개와 같다. 데이터를 잘 나눌 수 있는 질문을 찾는다면 계속 질문을 추가해서 분류 정확도를 높일 수 있다. 사이킷런의 DecisionTreeClassifier 클래스를 사용해 결정 트리 모델을 훈련해 보자. fit() 메서드를 호출해서 모델을 훈련한 다음 score() 메서드로 정확도를 평가해 보자.
```
from sklearn.tree import DecisionTreeClassifier
dt = DecisionTreeClassifier(random_state=42)
dt.fit(train_scaled, train_target)
print(dt.score(train_scaled, train_target))
print(dt.score(test_scaled, test_target))
```
훈련 세트에 대한 점수가 엄청 높게 나왔다. 테스트 세트의 성능은 그에 비해 조금 낮다. 과대적합된 모델이라고 볼 수 있다. 이 모델을 그림으로 표현하려면 plot_tree() 함수를 사용해 결정 트리를 이해하기 쉬운 트리 그림으로 출력해 보자.
```
import matplotlib.pyplot as plt
from sklearn.tree import plot_tree
plt.figure(figsize=(10,7))
plot_tree(dt)
plt.show()
```
맨 위의 노드를 루프노드라 부르고 맨 아래 끝에 달린 노드를 리프 노드라고 한다.
너무 복잡하니 plot_tree() 함수에서 트리의 깊이를 제한해서 출력해 보자. max_depth 매개변수를 1로 주면 루트 노드를 제외하고 하나의 노드를 더 확장하여 그린다. 또 filled 매개변수에서 클래스에 맞게 노드의 색을 칠할 수 있다. feature_names 매개변수에는 특성의 이름을 전달할 수 있다. 다음의 코드를 따라해 보자.
```
plt.figure(figsize=(10,7))
plot_tree(dt, max_depth=1, filled=True, feature_names=['alcohol','sugar','pH'])
plt.show()
```
루트 노드는 sugar가 -0.239 이하인지 질문을 한다. 만약 어떤 샘플의 당도가 -0.239와 같거나 작으면 왼쪽 가지로 이동한다. 그렇지 않으면 오른쪽으로 이동한다. 왼쪽이 yes 오른쪽이 no이다. 루트 노드의 총 샘플수는 5197개이다. 이 중에서 음성 클래스(레드와인)이 1258개이고, 양성 클래스(화이트 와인)는 3939개 이다. 이 값이 value에 나타나 있다.
이어서 왼쪽 노드를 살펴보자. 이 노드는 sugar가 더 낮은지 물어본다. -0.802 보다 같거나 낮으면 다시 왼쪽 가지로, 그렇지 않으면 오른쪽 가지로 이동한다. 이 노드에서 음성 클래스와 양성 클래스의 샘플 개수는 각각 1177개와 1745개이다. 루트 노드보다 양성 클래스, 즉 화이트 와인의 비율이 크게 줄었다. 그 이유는 오른쪽 노드를 보면 알 수 있다.
오른쪽 노드는 음성 클래스가 81개, 양성 클래스가 2194개로 대부분의 화이트 와인 샘플이 이 노드로 이동했다. 노드의 바탕 색깔을 유심히 보면 루트 노드보다 이 노드가 더 진하고 왼쪽 노드는 더 연해졌다. plot_tree() 함수에서 filled=True로 지정하면 클래스마다 색깔을 부여하고 어떤 클래스의 비율이 높아지면 점점 진한 색으로 표시한다. 결정 트리에서 예측하는 방법은 간단하다. 리프 노드에서 가장 많은 클래스가 예측 클래스가 된다. 앞에서 보았던 k-최근접 이웃과 매우 유사하다. 만약 이 결정트리의 성장을 여기서 멈춘다면 왼쪽 노드에 도달한 샘플과 오른쪽 노드에 도달한 샘플은 모두 양성 클래스로 예측된다. 두 노드 모두 양성 클래스의 개수가 많기 때문이다.
이번엔 노드 상자 안에 있는 gini에 대해 알아보자.
### - 불순도
gini는 지니 불순도(gini impurity)를 의미한다. DecisionTreeClassifier 클래스의 criterion 매개변수의 기본값이 gini이다. creterion 매개변수의 용도는 노드에서 데이터를 분할할 기준을 정하는 것이다. 앞의 루트 노드에서 -0.239를 기준으로 왼쪽과 오른쪽 노드로 나누었다. 나눈 방법은 바로 criterion 매개변수에 지정한 지니 불순도를 사용한 것이다. 지니 불순도는 클래스의 비율을 제곱해서 더한 다음 1에서 빼면된다.
- 지니 불순도 = 1 - (음성 클래스 비율의 제곱 + 양성 클래스 비율의 제곱)
### - 가지치기
예를 들어 열매를 잘 맺기 위해 과수원에서 가지치기를 하는 것처럼 결정 트리도 가지치기를 해야 한다. 그렇지 않으면 무작정 끝까지 자라나는 트리가 만들어진다. 훈련 세트에는 아주 잘 맞겠지만 테스트 세트에서 점수는 그에 못 미칠 것이다. 이것을 일반화가 잘 되지 않았다고 한다.
이제 가지치기를 해보자. 결정 트리에서 가지치기를 하는 가장 간단한 방법은 자라날 수 있는 트리의 최대 깊이를 지정하는 것이다. DecisionTreeClassifier 클래스의 max_depth 매개변수를 3으로 지정하여 모델을 만들어 보자.
```
dt = DecisionTreeClassifier(max_depth=3, random_state=42)
dt.fit(train_scaled, train_target)
print(dt.score(train_scaled, train_target))
print(dt.score(test_scaled, test_target))
```
훈련 세트의 성능은 낮아졌지만 테스트 세트의 성능은 거의 그대로이다. plot_tree() 함수로 그려보자.
```
plt.figure(figsize=(20,15))
plot_tree(dt, filled=True, feature_names = ['alcohol','sugar','pH'])
plt.show()
```
앞서 불순도를 기준으로 샘플을 나눈다고 했다. 불순도는 클래스별 비율을 가지고 계산하였다. 샘플을 어떤 클래스 비율로 나누는지 계산할 때 특성값의 스케일이 계산에 영향을 미칠까? 아니다. 특성값의 스케일은 결정 트리 알고리즘에 아무런 영향을 미치지 않는다. 따라서 표준화 전처리를 할 필요가 없다. 이것이 결정 트리 알고리즘의 또 다른 장점 중 하나이다.
그럼 앞서 전처리하기 전의 훈련 세트와 테스트 세트로 결정 트리모델을 다시 훈련해 보자.
```
dt = DecisionTreeClassifier(max_depth=3, random_state=42)
dt.fit(train_input, train_target)
print(dt.score(train_input, train_target))
print(dt.score(test_input, test_target))
```
결과가 정확히 같다. 이번에는 트리를 그려보자.
```
plt.figure(figsize=(20,15))
plot_tree(dt, filled=True, feature_names = ['alcohol','sugar','pH'])
plt.show()
```
결과를 보면 같은 트리지만, 특성값을 표준점수로 바꾸지 않은 터라 이해하기가 훨씬 쉽다. 당도가 1.625보다 크고 4.325보다 작은 와인중 알코올 도수가 11.025와 같거나 작은 것이 레드 와인이다. 그 이외에는 모두 화이트 와인으로 예측했다.
마지막으로 결정 트리는 어떤 특성이 가장 유용한지 나타내는 특성 중요도를 계산해 준다. 이 트리의 루트 노드와 깊이 1에서 sugar를 사용했기 때문에 아마도 sugar가 가장 유용한 특성중 하나일 것 같다. 특성 중요도는 결정 트리 모델의 feature_importances_ 속성에 저장되어 있다. 이 값을 출력해 보자.
```
print(dt.feature_importances_)
```
역시 두 번째 특성인 sugar가 0.87 정도로 특성 중요도가 가장 높다. 특성 중요도는 각 노드의 정보 이득과 전체 샘플에 대한 비율을 곱한 후 특성별로 더하여 계산한다. 특성 중요도를 활용하면 결정 트리 모델을 특성 선택에 활용할 수 있다. 이것이 결정 트리 알고리즘의 또 다른 장점중 하나이다.
출처 : 혼자 공부하는 머신러닝 + 딥러닝
| github_jupyter |
<!--BOOK_INFORMATION-->
<img align="left" style="padding-right:10px;" src="fig/cover-small.jpg">
*This notebook contains an excerpt from the [Whirlwind Tour of Python](http://www.oreilly.com/programming/free/a-whirlwind-tour-of-python.csp) by Jake VanderPlas; the content is available [on GitHub](https://github.com/jakevdp/WhirlwindTourOfPython).*
*The text and code are released under the [CC0](https://github.com/jakevdp/WhirlwindTourOfPython/blob/master/LICENSE) license; see also the companion project, the [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook).*
<!--NAVIGATION-->
< [Defining and Using Functions](08-Defining-Functions.ipynb) | [Contents](Index.ipynb) | [Iterators](10-Iterators.ipynb) >
# Errors and Exceptions
No matter your skill as a programmer, you will eventually make a coding mistake.
Such mistakes come in three basic flavors:
- *Syntax errors:* Errors where the code is not valid Python (generally easy to fix)
- *Runtime errors:* Errors where syntactically valid code fails to execute, perhaps due to invalid user input (sometimes easy to fix)
- *Semantic errors:* Errors in logic: code executes without a problem, but the result is not what you expect (often very difficult to track-down and fix)
Here we're going to focus on how to deal cleanly with *runtime errors*.
As we'll see, Python handles runtime errors via its *exception handling* framework.
## Runtime Errors
If you've done any coding in Python, you've likely come across runtime errors.
They can happen in a lot of ways.
For example, if you try to reference an undefined variable:
```
print(Q)
```
Or if you try an operation that's not defined:
```
1 + 'abc'
```
Or you might be trying to compute a mathematically ill-defined result:
```
2 / 0
```
Or maybe you're trying to access a sequence element that doesn't exist:
```
L = [1, 2, 3]
L[1000]
```
Note that in each case, Python is kind enough to not simply indicate that an error happened, but to spit out a *meaningful* exception that includes information about what exactly went wrong, along with the exact line of code where the error happened.
Having access to meaningful errors like this is immensely useful when trying to trace the root of problems in your code.
## Catching Exceptions: ``try`` and ``except``
The main tool Python gives you for handling runtime exceptions is the ``try``...``except`` clause.
Its basic structure is this:
```
try:
print("this gets executed first")
except:
print("this gets executed only if there is an error")
```
Note that the second block here did not get executed: this is because the first block did not return an error.
Let's put a problematic statement in the ``try`` block and see what happens:
```
try:
print("let's try something:")
x = 1 / 0 # ZeroDivisionError
except:
print("something bad happened!")
```
Here we see that when the error was raised in the ``try`` statement (in this case, a ``ZeroDivisionError``), the error was caught, and the ``except`` statement was executed.
One way this is often used is to check user input within a function or another piece of code.
For example, we might wish to have a function that catches zero-division and returns some other value, perhaps a suitably large number like $10^{100}$:
```
def safe_divide(a, b):
try:
return a / b
except:
return 1E100
safe_divide(1, 2)
safe_divide(2, 0)
```
There is a subtle problem with this code, though: what happens when another type of exception comes up? For example, this is probably not what we intended:
```
safe_divide (1, '2')
```
Dividing an integer and a string raises a ``TypeError``, which our over-zealous code caught and assumed was a ``ZeroDivisionError``!
For this reason, it's nearly always a better idea to catch exceptions *explicitly*:
```
def safe_divide(a, b):
try:
return a / b
except ZeroDivisionError:
return 1E100
safe_divide(1, 0)
safe_divide(1, '2')
```
We're now catching zero-division errors only, and letting all other errors pass through un-modified.
## Raising Exceptions: ``raise``
We've seen how valuable it is to have informative exceptions when using parts of the Python language.
It's equally valuable to make use of informative exceptions within the code you write, so that users of your code (foremost yourself!) can figure out what caused their errors.
The way you raise your own exceptions is with the ``raise`` statement. For example:
```
raise RuntimeError("my error message")
```
As an example of where this might be useful, let's return to our ``fibonacci`` function that we defined previously:
```
def fibonacci(N):
L = []
a, b = 0, 1
while len(L) < N:
a, b = b, a + b
L.append(a)
return L
```
One potential problem here is that the input value could be negative.
This will not currently cause any error in our function, but we might want to let the user know that a negative ``N`` is not supported.
Errors stemming from invalid parameter values, by convention, lead to a ``ValueError`` being raised:
```
def fibonacci(N):
if N < 0:
raise ValueError("N must be non-negative")
L = []
a, b = 0, 1
while len(L) < N:
a, b = b, a + b
L.append(a)
return L
fibonacci(10)
fibonacci(-10)
```
Now the user knows exactly why the input is invalid, and could even use a ``try``...``except`` block to handle it!
```
N = -10
try:
print("trying this...")
print(fibonacci(N))
except ValueError:
print("Bad value: need to do something else")
```
### Exercise
1. Let's come back to one of our previous exercise contexts. Create a function out of one of them and raise an error in this function.
2. Call the above function for an input, which will raise an exception, catch the error that you raised in your function and redefine the reaction to it.
## Diving Deeper into Exceptions
Briefly, I want to mention here some other concepts you might run into.
I'll not go into detail on these concepts and how and why to use them, but instead simply show you the syntax so you can explore more on your own.
### Accessing the error message
Sometimes in a ``try``...``except`` statement, you would like to be able to work with the error message itself.
This can be done with the ``as`` keyword:
```
try:
x = 1 / 0
except ZeroDivisionError as err:
print("Error class is: ", type(err))
print("Error message is:", err)
```
With this pattern, you can further customize the exception handling of your function.
### Defining custom exceptions
In addition to built-in exceptions, it is possible to define custom exceptions through *class inheritance*.
For instance, if you want a special kind of ``ValueError``, you can do this:
```
class MySpecialError(ValueError):
pass
raise MySpecialError("here's the message")
```
This would allow you to use a ``try``...``except`` block that only catches this type of error:
```
try:
print("do something")
raise MySpecialError("[informative error message here]")
except MySpecialError:
print("do something else")
```
You might find this useful as you develop more customized code.
## ``try``...``except``...``else``...``finally``
In addition to ``try`` and ``except``, you can use the ``else`` and ``finally`` keywords to further tune your code's handling of exceptions.
The basic structure is this:
```
try:
print("try something here")
except:
print("this happens only if it fails")
else:
print("this happens only if it succeeds")
finally:
print("this happens no matter what")
```
The utility of ``else`` here is clear, but what's the point of ``finally``?
Well, the ``finally`` clause really is executed *no matter what*: I usually see it used to do some sort of cleanup after an operation completes.
<!--NAVIGATION-->
< [Defining and Using Functions](08-Defining-Functions.ipynb) | [Contents](Index.ipynb) | [Iterators](10-Iterators.ipynb) >
| github_jupyter |
```
import random
import numpy as np
class NeuralNetwork(object):
def __init__(self, sizes):
self.num_layers = len(sizes)
self.sizes = sizes
self.weights = [np.random.randn(y, x+1) \
for x, y in zip((sizes[:-1]), sizes[1:])] # biases are included in weights
def feedforward(self, a):
for w in self.weights:
a = np.concatenate((a,np.array([1]).reshape(1,1))) # add bias neuron
a = sigmoid(np.dot(w, a))
return a
def SGD(self, training_data, epochs, mini_batch_size, eta, test_data=None):
if test_data: n_test = len(test_data)
n = len(training_data)
for j in range(epochs):
random.shuffle(training_data)
mini_batches = [training_data[k:k+mini_batch_size] \
for k in range(0, n, mini_batch_size)]
for mini_batch in mini_batches:
self.update_mini_batch(mini_batch, eta)
if test_data:
print ("Epoch {0}: {1} / {2}".format( \
j, self.evaluate_0(test_data), n_test))
else:
print ("Epoch {0} complete".format(j))
def update_mini_batch(self, mini_batch, eta):
nabla_w = [np.zeros(w.shape) for w in self.weights]
for x, y in mini_batch:
delta_nabla_w = self.backprop(x, y)
nabla_w = [nw+dnw for nw, dnw in zip(nabla_w, delta_nabla_w)]
self.weights = [w-(eta/len(mini_batch))*nw
for w, nw in zip(self.weights, nabla_w)]
def backprop(self, x, y):
nabla_w = [np.zeros(w.shape) for w in self.weights]
# feedforward
activation = x
activations = [activation] # list to store all the activations, layer by layer
zs = [] # list to store all the z vectors, layer by layer
for w in self.weights:
activation = np.concatenate((activation,np.array([1]).reshape(1,1)))
activations[-1]=activation
z = np.dot(w, activation)
zs.append(z)
activation = sigmoid(z)
activations.append(activation)
# backward pass
delta = self.cost_derivative(activations[-1], y) * \
sigmoid_prime(zs[-1])
nabla_w[-1] = np.dot(delta, activations[-2].transpose())
for l in range(2, self.num_layers):
z = zs[-l]
sp = sigmoid_prime(z)
delta = np.dot(self.weights[-l+1].transpose(), delta)[:-1]
delta = delta * sp
nabla_w[-l] = np.dot(delta, activations[-l-1].transpose())
return nabla_w
def evaluate_0(self, test_data):
test_results = [(int(self.feedforward(x)[1][0]>0.1), y)
for (x, y) in test_data]
return sum(int(x == y) for (x, y) in test_results)
def evaluate(self, test_data):
test_results = [(np.argmax(self.feedforward(x)), y)
for (x, y) in test_data]
return sum(int(x == y) for (x, y) in test_results)
def cost_derivative(self, output_activations, y):
return (output_activations-y)
#### Miscellaneous functions
def sigmoid(z):
return 1.0/(1.0+np.exp(-z))
def sigmoid_prime(z):
return sigmoid(z)*(1-sigmoid(z))
import pandas as pd
car_data=pd.read_csv('car.csv')
car_data = car_data.reindex(columns=['IsBadBuy','Size','Make','VNST','IsOnlineSale','VehicleAge','Transmission',
'WheelType','Auction'])
shuffler= np.random.permutation(len(car_data))
car_shuffle = car_data.take(shuffler) # pandas' shuffling method in comparison of random.shuffle
# X preparation
Size = pd.get_dummies(car_data['Size'],prefix='Size') # generate dummy varibles from categorical varible
Make = pd.get_dummies(car_data['Make'],prefix='Make')
VNST = pd.get_dummies(car_data['VNST'],prefix='VNST')
VehicleAge = pd.get_dummies(car_data['VehicleAge'],prefix='VehicleAge')
WheelType = pd.get_dummies(car_data['WheelType'],prefix='WheelType')
Auction = pd.get_dummies(car_data['Auction'],prefix='Auction')
IsOnlineSale =(car_data.IsOnlineSale=='Yes').apply(float)
X= Size.join(Make).join(VNST).join(IsOnlineSale).join(VehicleAge).join(WheelType).join(Auction)
Y=pd.get_dummies(car_data['IsBadBuy'],prefix='IsBadbuy')
car_training=[(X.iloc[i].values.reshape(93,1),Y.iloc[i].values.reshape(2,1)) for i in X.index]
#test data preparing, as did with training data
car_test=pd.read_csv('car_test.csv')
car_test = car_test.reindex(columns=['IsBadBuy','Size','Make','VNST','IsOnlineSale','VehicleAge','Transmission',
'WheelType','Auction'])
Size = pd.get_dummies(car_test['Size'],prefix='Size') # generate dummy varibles from categorical varible
Make = pd.get_dummies(car_test['Make'],prefix='Make')
VNST = pd.get_dummies(car_test['VNST'],prefix='VNST')
VehicleAge = pd.get_dummies(car_test['VehicleAge'],prefix='VehicleAge')
WheelType = pd.get_dummies(car_test['WheelType'],prefix='WheelType')
Auction = pd.get_dummies(car_test['Auction'],prefix='Auction')
IsOnlineSale =(car_test.IsOnlineSale=='Yes').apply(float)
X= Size.join(Make).join(VNST).join(IsOnlineSale).join(VehicleAge).join(WheelType).join(Auction)
Y=car_test['IsBadBuy']
car_test=[(X.iloc[i].values.reshape(93,1),Y.iloc[i]) for i in X.index]
# set of net for Car training
net = NeuralNetwork([93, 10, 2])
net.SGD(car_training, 10, 50, 1.0)
net.SGD(car_training, 30, 50, 1.0,test_data=car_test)
ProbIsGoodBuy=[net.feedforward(x)[0][0] for (x,y) in car_test]
ProbIsBadBuy=[net.feedforward(x)[1][0] for (x,y) in car_test]
import matplotlib.pyplot as plt
plt.hist(ProbIsBadBuy,bins=30,color='red',alpha=0.3)
plt.hist(ProbIsGoodBuy,bins=30,color='blue',alpha=0.5)
plt.show()
test_result=pd.read_csv('car_test.csv')
test_result = test_result.reindex(columns=['IsBadBuy','Size','Make','VNST','IsOnlineSale','VehicleAge','Transmission',
'WheelType','Auction'])
test_result['ProbIsBadBuy']=ProbIsBadBuy
test_result["ProbCat"]=pd.qcut(ProbIsBadBuy,10,precision=1)
#test_result=test_result.sort_values('ProbIsBadBuy')
test_result.groupby("ProbCat").count()
test_result.groupby("ProbCat").sum()
import psycopg2
con= psycopg2.connect(database="cqbus",user="gpadmin",password="gpadmin",host="192.168.0.93",port="5432")
import pandas as pd
df = pd.read_sql('select * from point_data where lng != 0 limit 10;', con=con)
df
#con.close()
df = pd.read_sql('select * from cqbus2017 limit 10;', con=con)
df
#con.close()
```
| github_jupyter |
# Automated Gradual Pruning Schedule
Michael Zhu and Suyog Gupta, ["To prune, or not to prune: exploring the efficacy of pruning for model compression"](https://arxiv.org/pdf/1710.01878), 2017 NIPS Workshop on Machine Learning of Phones and other Consumer Devices<br>
<br>
After completing sensitivity analysis, decide on your pruning schedule.
## Table of Contents
1. [Implementation of the gradual sparsity function](#Implementation-of-the-gradual-sparsity-function)
2. [Visualize pruning schedule](#Visualize-pruning-schedule)
3. [References](#References)
```
import numpy
import matplotlib.pyplot as plt
from functools import partial
import torch
from torch.autograd import Variable
from ipywidgets import widgets, interact
```
## Implementation of the gradual sparsity function
The function ```sparsity_target``` implements the gradual sparsity schedule from [[1]](#zhu-gupta):<br><br>
<b><i>"We introduce a new automated gradual pruning algorithm in which the sparsity is increased from an initial sparsity value $s_i$ (usually 0) to a final sparsity value $s_f$ over a span of $n$ pruning steps, starting at training step $t_0$ and with pruning frequency $\Delta t$."</i></b><br>
<br>
<div id="eq:zhu_gupta_schedule"></div>
<center>
$\large
\begin{align}
s_t = s_f + (s_i - s_f) \left(1- \frac{t-t_0}{n\Delta t}\right)^3
\end{align}
\ \ for
\large \ \ t \in \{t_0, t_0+\Delta t, ..., t_0+n\Delta t\}
$
</center>
<br>
Pruning happens once at the beginning of each epoch, until the duration of the pruning (the number of epochs to prune) is exceeded. After pruning ends, the training continues without pruning, but the pruned weights are kept at zero.
```
def sparsity_target(starting_epoch, ending_epoch, initial_sparsity, final_sparsity, current_epoch):
if final_sparsity < initial_sparsity:
return current_epoch
if current_epoch < starting_epoch:
return current_epoch
span = ending_epoch - starting_epoch
target_sparsity = ( final_sparsity +
(initial_sparsity - final_sparsity) *
(1.0 - ((current_epoch-starting_epoch)/span))**3)
return target_sparsity
```
## Visualize pruning schedule
When using the Automated Gradual Pruning (AGP) schedule, you may want to visualize how the pruning schedule will look as a function of the epoch number. This is called the *sparsity function*. The widget below will help you do this.<br>
There are three knobs you can use to change the schedule:
- ```duration```: this is the number of epochs over which to use the AGP schedule ($n\Delta t$).
- ```initial_sparsity```: $s_i$
- ```final_sparsity```: $s_f$
- ```frequency```: this is the pruning frequency ($\Delta t$).
```
def draw_pruning(duration, initial_sparsity, final_sparsity, frequency):
epochs = []
sparsity_levels = []
# The derivative of the sparsity (i.e. sparsity rate of change)
d_sparsity = []
if frequency=='':
frequency = 1
else:
frequency = int(frequency)
for epoch in range(0,40):
epochs.append(epoch)
current_epoch=Variable(torch.FloatTensor([epoch]), requires_grad=True)
if epoch<duration and epoch%frequency == 0:
sparsity = sparsity_target(
starting_epoch=0,
ending_epoch=duration,
initial_sparsity=initial_sparsity,
final_sparsity=final_sparsity,
current_epoch=current_epoch
)
sparsity_levels.append(sparsity)
sparsity.backward()
d_sparsity.append(current_epoch.grad.item())
current_epoch.grad.data.zero_()
else:
sparsity_levels.append(sparsity)
d_sparsity.append(0)
plt.plot(epochs, sparsity_levels, epochs, d_sparsity)
plt.ylabel('sparsity (%)')
plt.xlabel('epoch')
plt.title('Pruning Rate')
plt.ylim(0, 100)
plt.draw()
duration_widget = widgets.IntSlider(min=0, max=100, step=1, value=28)
si_widget = widgets.IntSlider(min=0, max=100, step=1, value=0)
interact(draw_pruning,
duration=duration_widget,
initial_sparsity=si_widget,
final_sparsity=(0,100,1),
frequency='2');
```
<div id="toc"></div>
## References
1. <div id="zhu-gupta"></div> **Michael Zhu and Suyog Gupta**.
[*To prune, or not to prune: exploring the efficacy of pruning for model compression*](https://arxiv.org/pdf/1710.01878),
NIPS Workshop on Machine Learning of Phones and other Consumer Devices,
2017.
| github_jupyter |
# Coursework 2: Neural Networks
This coursework covers the topics covered in class regarding neural networks for image classification.
This coursework includes both coding questions as well as written ones. Please upload the notebook, which contains your code, results and answers as a pdf file onto Cate.
Dependencies: If you work on a college computer in the Computing Lab, where Ubuntu 18.04 is installed by default, you can use the following virtual environment for your work, where relevant Python packages are already installed.
`source /vol/bitbucket/wbai/virt/computer_vision_ubuntu18.04/bin/activate`
Alternatively, you can use pip, pip3 or anaconda etc to install Python packages.
**Note 1:** please read the both the text and code comment in this notebook to get an idea what you are supposed to implement.
**Note 2:** If you are using the virtual environment in the Computing Lab, please run the following command in the command line before opening jupyter-notebook and importing tensorflow. This will tell tensorflow where the Nvidia CUDA libariries are.
`export LD_LIBRARY_PATH=/vol/cuda/9.0.176/lib64/:"${LD_LIBRARY_PATH}}"`
```
# Import libraries
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import tensorflow as tf
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout
```
## Question 1 (20 points)
Throughout this coursework you will be working with the Fashion-MNIST dataset. If you are interested, you may find relevant information regarding the dataset in this paper.
[1] Fashion-MNIST: A novel image dataset for benchmarking machine learning algorithms. Han Xiao, Kashif Rasul, Roland Vollgraf. [arXiv:1708.07747](https://arxiv.org/abs/1708.07747)
Be sure that you have the following files in your working directory: data.tar.gz and reader.py. Loading the data can be done as follows:
`from reader import get_images
(x_train, y_train), (x_test, y_test) = get_images()`
The dataset is already split into a set of 60,000 training images and a set of 10,000 test images. The images are of size 28x28 pixels and stored as 784-D vector. So if you would like to visualise the images, you need to reshape the array.
There are in total 10 label classes, which are:
* 0: T-shirt/top
* 1: Trousers
* 2: Pullover
* 3: Dress
* 4: Coat
* 5: Sandal
* 6: Shirt
* 7: Sneaker
* 8: Bag
* 9: Ankle boot
### 1.1 Load data (6 points)
Load the dataset and print the dimensions of the training set and the test set.
```
from reader import get_images
(x_train, y_train), (x_test, y_test) = get_images()
print('dimensions of the training set:',x_train.shape,y_train.shape)
print('dimensions of the test set:',x_test.shape,y_test.shape)
```
### 1.2 Visualize data (6 points)
Visualise 3 training images (T-shirt, trousers and pullover) and 3 test images (dress, coat and sandal).
```
num=0
class_name=['T-shirt','Trousers','Pullover','Dress','Coat','Sandal','Shirt','Sneaker','Bag','Ankle B']
image=[np.reshape(x_train[1],(28,28))]*6
for index in range(np.random.randint(9000),10000):
if num<3 and y_train[index]==num:
image[num]=np.reshape(x_train[index],(28,28))
num+=1
if num>=3 and y_test[index]==num:
image[num]=np.reshape(x_test[index],(28,28))
num+=1
if num==6: break
plt.figure
for i in range(6):
plt.subplot(2,3,i+1)
plt.imshow(image[i],cmap='gray')
plt.title(class_name[i])
```
### 1.3 Data balance (4 points)
Print out the number of training samples for each class.
```
dict = {}
for class_ in y_train:
dict[class_] = dict.get(class_, 0) + 1
dictlist=sorted(dict.items(), key = lambda x: x[0], reverse=False)
for i in range(10):
print('Sample Number of No.',dictlist[i][0],' ',class_name[i],'=',dictlist[i][1],sep='')
```
### 1.4 Discussion (4 points)
Is the dataset balanced? What would happen if the dataset is not balanced in the context of image classification?
Well, we can know from the output above that the number of training samples for each class is equal, which is 6000. Traditional classification algorithm, which pay more attention on overall classification accuracy, focus too much on the majority of the class. In result, performance degradation of minority classification can not be inevitable.
## Question 2 (40 points)
Build a neural network and train it with the Fashion-MNIST dataset. Here, we use the keras library, which is a high-level neural network library built upon tensorflow.
```
# Convert the label class into a one-hot representation
num_classes = 10
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# normalization from 0-255 to 0-1
x_train=x_train.astype('float32')/255
x_test=x_test.astype('float32')/255
```
### 2.1 Build a multi-layer perceptron, also known as multi-layer fully connected network. You need to define the layers, the loss function, the optimiser and evaluation metric. (30 points)
```
model = keras.models.Sequential()
# as input layer in a sequential model:
model.add(Dense(512,activation='relu',input_shape=(784,)))
model.add(Dropout(0.25))
#as hidden layer in the model
model.add(Dense(144,activation='relu'))
model.add(Dropout(0.20))
#as output layer in model
model.add(Dense(num_classes,activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='sgd', metrics=["accuracy"])
print(model.summary())
```
### 2.2 Define the optimisation parameters including the batch size and the number of epochs and then run the optimiser. (10 points)
We have tested that for an appropriate network architecture, on a personal laptop and with only CPU, it takes about a few seconds per epoch to train the network. For 100 epochs, it takes about a coffee break's time to finish the training. If you run it on a powerful GPU, it would be even much faster.
```
batch_size = 32
epochs = 20
model.fit(x_train, y_train,epochs=epochs,batch_size=batch_size)
```
## Question 3 (20 points)
Evaluate the performance of your network with the test data.
Visualize the performance using appropriate metrics and graphs (eg. confusion matrix).
Comment on your per class performance and how it could be better.
```
# This function is provided for you to display the confusion matrix.
# For more information about the confusion matrix, you can read at
# https://en.wikipedia.org/wiki/Confusion_matrix
import itertools
def plot_confusion_matrix(cm, classes, normalize=False, title='Confusion matrix', cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
cm: confusion matrix, default to be np.int32 data type
classes: a list of the class labels or class names
normalize: normalize the matrix so that each row amounts to one
cmap: color map
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.tight_layout()
```
### 3.1 Evaluate the classification accuracy on the test set (10 points)
```
score = model.evaluate(x_test, y_test)
print('Test Loss','%.4f' %score[0])
print('Test Accuracy',score[1])
```
### 3.2 Calculate and plot the confusion matrix (10 points)
```
from sklearn.metrics import confusion_matrix
y_pred = model.predict(x_test)
y_pred = np.argmax(y_pred, axis=1)
y_test = np.argmax(y_test, axis=1)
# confustion matrix
cm=confusion_matrix(y_test, y_pred)
plot_confusion_matrix(cm,class_name)
```
## Question 4 (20 points)
Take two photos, one of your clothes or shoes that belongs to one of 10 classes, the other that does not belong to any class.
Use either Python or other software (Photoshop, Gimp, or any image editer) to convert the photos into grayscale, crop the region of interest and reshape into the size of 28x28.
### 4.1 Load and visualise your own images (6 points)
```
import matplotlib.image
def rgb2gray(rgb):
r, g, b = rgb[:,:,0], rgb[:,:,1], rgb[:,:,2]
gray = 0.2989 * r + 0.5870 * g + 0.1140 * b
return gray
image_name=["Queen's_Tower","T-shirt"]
image_reshape=[]
for i in range(len(image_name)):
img_colour = matplotlib.image.imread(image_name[i]+'.png')
img_grey = rgb2gray(img_colour)
plt.subplot(1,2,i+1)
plt.imshow(img_grey,cmap='gray')
plt.title(image_name[i])
image_reshape.append(np.reshape(img_grey,(1,784)))
```
### 4.2 Test your network on the two images and show the classification results (10 points)
```
for i in range(len(image_reshape)):
pred=model.predict(image_reshape[i])
# print(pred)
class_index=pred.argmax(axis=1)[0]
print('Prediction of',image_name[i]+':',class_name[class_index])
```
### 4.3 Discuss the classification results and provide one method to improve real life performance of the network (4 points)
Well, this classification algorithm identified T-shirt sucsessfully but class the Qieen's Tower as a bag, which is wrong withou suspense. According to the result of this test, we can say that
* This algorithm is good enough to class thoese 10 classes related to clothing and wearing.
* Cannot identify other unlabel classes under structure of muilt-layer connected network and the limited traning data.
There something we can do to bring it into reallife application
1. A large number of sample data and various label classes are needed to adapt to the reallife. Affine transformation can be used to increase the number of data.
2. Combined with neural networks to construct a more complex model is a good way to deal with more data with less parameters.
3. Adding regularisation term is Another method to improve the accuracy of classification
## 5. Survey
How long did the coursework take you to solve?
The whole afternoon of lovely Thursday
| github_jupyter |
```
!python ../input/jigsawsrc/inference.py \
--num_folds 10 \
--base_model ../input/deberta/deberta-large \
--base_model_name microsoft/deberta-large \
--weights_dir ../input/ranking-30-deberta-large \
--data_path ../input/jigsaw-toxic-severity-rating/comments_to_score.csv \
--save_path preds_30.csv
!python ../input/jigsawsrc/inference.py \
--num_folds 5 \
--base_model ../input/deberta/deberta-large \
--base_model_name microsoft/deberta-large \
--weights_dir ../input/ranking-52-deberta-large-e2 \
--data_path ../input/jigsaw-toxic-severity-rating/comments_to_score.csv \
--save_path preds_52.csv
!python ../input/jigsawsrc/inference.py \
--num_folds 5 \
--base_model ../input/rembert/rembert \
--base_model_name google/rembert \
--weights_dir ../input/ranking-58-rembert \
--data_path ../input/jigsaw-toxic-severity-rating/comments_to_score.csv \
--save_path preds_58.csv
!python ../input/jigsawsrc/inference.py \
--num_folds 5 \
--base_model ../input/deberta-v3/deberta-v3-base \
--base_model_name microsoft/deberta-v3-base \
--weights_dir ../input/ranking-61-deberta-v3-base-with-oof/ranking_61_deberta_v3_base \
--data_path ../input/jigsaw-toxic-severity-rating/comments_to_score.csv \
--save_path preds_61.csv
!python ../input/jigsawsrc/inference.py \
--num_folds 5 \
--base_model ../input/deberta-v3/deberta-v3-large \
--base_model_name microsoft/deberta-v3-large \
--weights_dir ../input/ranking-63-deberta-v3-large \
--data_path ../input/jigsaw-toxic-severity-rating/comments_to_score.csv \
--save_path preds_63.csv
!python ../input/jigsawsrc/inference.py \
--num_folds 5 \
--base_model ../input/k/amontgomerie/roberta/distilroberta-base \
--base_model_name distilroberta-base \
--weights_dir ../input/ranking-64-distilroberta-base-with-oof \
--data_path ../input/jigsaw-toxic-severity-rating/comments_to_score.csv \
--save_path preds_64.csv
import pandas as pd
weights = {
'weight_30': 0.4024099870709701,
'weight_52': 0.9011229226395379,
'weight_58': 0.755155406930562,
'weight_61': 0.27330107235558226,
'weight_63': 0.02807554189147518,
'weight_64': 0.029575587654601887,
}
model_preds = {
"preds_30": pd.read_csv("preds_30.csv")["score"],
"preds_52": pd.read_csv("preds_52.csv")["score"],
"preds_58": pd.read_csv("preds_58.csv")["score"],
"preds_61": pd.read_csv("preds_61.csv")["score"],
"preds_63": pd.read_csv("preds_63.csv")["score"],
"preds_64": pd.read_csv("preds_64.csv")["score"],
}
mean_preds = (
(model_preds["preds_30"] * weights["weight_30"])
+ (model_preds["preds_52"] * weights["weight_52"])
+ (model_preds["preds_58"] * weights["weight_58"])
+ (model_preds["preds_61"] * weights["weight_61"])
+ (model_preds["preds_63"] * weights["weight_63"])
+ (model_preds["preds_64"] * weights["weight_64"])
)
test_data = pd.read_csv("../input/jigsaw-toxic-severity-rating/comments_to_score.csv")
submission = pd.DataFrame({"comment_id": test_data.comment_id, "score": mean_preds})
submission["score"] = submission.score.rank()
submission.to_csv("./submission.csv", index=False)
submission
```
| github_jupyter |
```
import os
import pandas as pd
import numpy as np
import seaborn as sns
sns.set()
import matplotlib.pyplot as plt
sns.set_style('darkgrid')
#print curent dir
print("Current Working Directory " , os.getcwd())
df = pd.read_csv ('WA_Fn-UseC_-Telco-Customer-Churn.csv')
df.dtypes
df.shape
df.head(5)
df.customerID.nunique()
df.PhoneService.unique()
pd.crosstab(df.PhoneService, df.MultipleLines)
df.SeniorCitizen.unique()
```
1.Services that are – phone, multiple lines, internet, online security, online backup, device protection, tech support, and streaming TV and movies
2.Demographic info about customers – gender, age range, and if they have partners and dependents
```
df.dtypes
df.PhoneService.unique()
df_phones=df[df.PhoneService=="Yes"]
pd.crosstab(df_phones.PhoneService, df_phones.MultipleLines)
```
1. QUESTION 1 (Part a)
```
fig, axes = plt.subplots(1, 4, figsize=(20, 5), sharey=True)
fig.suptitle('Effect on Phone Service')
sns.countplot(ax=axes[0] , x ='gender', hue = "PhoneService", data = df)
axes[0].set_title("gender")
sns.countplot(ax=axes[1] , x ='SeniorCitizen', hue = "PhoneService", data = df)
axes[1].set_title("Senior Citizen")
sns.countplot(ax=axes[2] , x ='Partner', hue = "PhoneService", data = df)
axes[2].set_title("Partner")
sns.countplot(ax=axes[3] , x ='Dependents', hue = "PhoneService", data = df)
axes[3].set_title("Dependents")
fig, axes = plt.subplots(1, 4, figsize=(20, 5), sharey=True)
fig.suptitle('Effect on Phone Service (with phones only)')
sns.countplot(ax=axes[0] , x ='gender', hue = "MultipleLines", data = df_phones)
axes[0].set_title("gender")
sns.countplot(ax=axes[1] , x ='SeniorCitizen', hue = "MultipleLines", data = df_phones)
axes[1].set_title("Senior Citizen")
sns.countplot(ax=axes[2] , x ='Partner', hue = "MultipleLines", data = df_phones)
axes[2].set_title("Partner")
sns.countplot(ax=axes[3] , x ='Dependents', hue = "MultipleLines", data = df_phones)
axes[3].set_title("Dependents")
fig, axes = plt.subplots(1, 4, figsize=(20, 5), sharey=True)
fig.suptitle('Effect on Internet Service')
sns.countplot(ax=axes[0] , x ='gender', hue = "InternetService", data = df)
axes[0].set_title("gender")
sns.countplot(ax=axes[1] , x ='SeniorCitizen', hue = "InternetService", data = df)
axes[1].set_title("Senior Citizen")
sns.countplot(ax=axes[2] , x ='Partner', hue = "InternetService", data = df)
axes[2].set_title("Partner")
sns.countplot(ax=axes[3] , x ='Dependents', hue = "InternetService", data = df)
axes[3].set_title("Dependents")
df_Internet=df[df.InternetService!="No"]
fig, axes = plt.subplots(1, 4, figsize=(20, 5), sharey=True)
fig.suptitle('Effect on Online Security')
sns.countplot(ax=axes[0] , x ='gender', hue = "OnlineSecurity", data = df_Internet)
axes[0].set_title("gender")
sns.countplot(ax=axes[1] , x ='SeniorCitizen', hue = "OnlineSecurity", data = df_Internet)
axes[1].set_title("Senior Citizen")
sns.countplot(ax=axes[2] , x ='Partner', hue = "OnlineSecurity", data = df_Internet)
axes[2].set_title("Partner")
sns.countplot(ax=axes[3] , x ='Dependents', hue = "OnlineSecurity", data = df_Internet)
axes[3].set_title("Dependents")
fig, axes = plt.subplots(1, 4, figsize=(20, 5), sharey=True)
fig.suptitle('Effect on Online Backup')
sns.countplot(ax=axes[0] , x ='gender', hue = "OnlineBackup", data = df_Internet)
axes[0].set_title("gender")
sns.countplot(ax=axes[1] , x ='SeniorCitizen', hue = "OnlineBackup", data = df_Internet)
axes[1].set_title("Senior Citizen")
sns.countplot(ax=axes[2] , x ='Partner', hue = "OnlineBackup", data = df_Internet)
axes[2].set_title("Partner")
sns.countplot(ax=axes[3] , x ='Dependents', hue = "OnlineBackup", data = df_Internet)
axes[3].set_title("Dependents")
```
QUESTION 1 PART B
```
fig, axes = plt.subplots(1, 4, figsize=(20, 5), sharey=True)
fig.suptitle('Effect on churn')
sns.countplot(ax=axes[0] , x ='gender', hue = "Churn", data = df)
axes[0].set_title("gender")
sns.countplot(ax=axes[1] , x ='SeniorCitizen', hue = "Churn", data = df)
axes[1].set_title("Senior Citizen")
sns.countplot(ax=axes[2] , x ='Partner', hue = "Churn", data = df)
axes[2].set_title("Partner")
sns.countplot(ax=axes[3] , x ='Dependents', hue = "Churn", data = df)
axes[3].set_title("Dependents")
g = sns.FacetGrid(df, col="gender", row="Partner", hue = "SeniorCitizen")
g.map(sns.countplot, "Churn")
g.add_legend()
```
Answer: Senior citizens
df.dtypes
```
df.dtypes
g = sns.FacetGrid(df, col="PhoneService", row="Partner", hue = "InternetService")
g.map(sns.countplot, "Churn")
g.add_legend()
```
| github_jupyter |
## 1. Where are the old left-handed people?
<p><img src="https://s3.amazonaws.com/assets.datacamp.com/production/project_479/img/Obama_signs_health_care-20100323.jpg" alt="Barack Obama signs the Patient Protection and Affordable Care Act at the White House, March 23, 2010"></p>
<p>Barack Obama is left-handed. So are Bill Gates and Oprah Winfrey; so were Babe Ruth and Marie Curie. A <a href="https://www.nejm.org/doi/full/10.1056/NEJM199104043241418">1991 study</a> reported that left-handed people die on average nine years earlier than right-handed people. Nine years! Could this really be true? </p>
<p>In this notebook, we will explore this phenomenon using age distribution data to see if we can reproduce a difference in average age at death purely from the changing rates of left-handedness over time, refuting the claim of early death for left-handers. This notebook uses <code>pandas</code> and Bayesian statistics to analyze the probability of being a certain age at death given that you are reported as left-handed or right-handed.</p>
<p>A National Geographic survey in 1986 resulted in over a million responses that included age, sex, and hand preference for throwing and writing. Researchers Avery Gilbert and Charles Wysocki analyzed this data and noticed that rates of left-handedness were around 13% for people younger than 40 but decreased with age to about 5% by the age of 80. They concluded based on analysis of a subgroup of people who throw left-handed but write right-handed that this age-dependence was primarily due to changing social acceptability of left-handedness. This means that the rates aren't a factor of <em>age</em> specifically but rather of the <em>year you were born</em>, and if the same study was done today, we should expect a shifted version of the same distribution as a function of age. Ultimately, we'll see what effect this changing rate has on the apparent mean age of death of left-handed people, but let's start by plotting the rates of left-handedness as a function of age.</p>
<p>This notebook uses two datasets: <a href="https://www.cdc.gov/nchs/data/statab/vs00199_table310.pdf">death distribution data</a> for the United States from the year 1999 (source website <a href="https://www.cdc.gov/nchs/nvss/mortality_tables.htm">here</a>) and rates of left-handedness digitized from a figure in this <a href="https://www.ncbi.nlm.nih.gov/pubmed/1528408">1992 paper by Gilbert and Wysocki</a>. </p>
```
# import libraries
# ... YOUR CODE FOR TASK 1 ...
import pandas as pd
import matplotlib.pyplot as plt
# load the data
data_url_1 = "https://gist.githubusercontent.com/mbonsma/8da0990b71ba9a09f7de395574e54df1/raw/aec88b30af87fad8d45da7e774223f91dad09e88/lh_data.csv"
lefthanded_data = pd.read_csv(data_url_1)
# plot male and female left-handedness rates vs. age
%matplotlib inline
fig, ax = plt.subplots() # create figure and axis objects
ax.plot("Age", "Female", data=lefthanded_data, marker = 'o') # plot "Female" vs. "Age"
ax.plot("Age", "Male", data=lefthanded_data, marker = 'x') # plot "Male" vs. "Age"
ax.legend() # add a legend
ax.set_xlabel("Sex")
ax.set_ylabel("Age")
```
## 2. Rates of left-handedness over time
<p>Let's convert this data into a plot of the rates of left-handedness as a function of the year of birth, and average over male and female to get a single rate for both sexes. </p>
<p>Since the study was done in 1986, the data after this conversion will be the percentage of people alive in 1986 who are left-handed as a function of the year they were born. </p>
```
# create a new column for birth year of each age
# ... YOUR CODE FOR TASK 2 ...
lefthanded_data["Birth_year"] = 1986 - lefthanded_data["Age"]
# create a new column for the average of male and female
# ... YOUR CODE FOR TASK 2 ...
lefthanded_data["Mean_lh"] = lefthanded_data[["Female","Male"]].mean(axis=1)
# create a plot of the 'Mean_lh' column vs. 'Birth_year'
fig, ax = plt.subplots()
ax.plot("Birth_year", "Mean_lh", data=lefthanded_data) # plot 'Mean_lh' vs. 'Birth_year'
ax.set_xlabel("Mean_lh") # set the x label for the plot
ax.set_ylabel("Birth_year") # set the y label for the plot
```
## 3. Applying Bayes' rule
<p><strong>Bayes' rule</strong> or <strong>Bayes' theorem</strong> is a statement about conditional probability which allows us to update our beliefs after seeing evidence. The probability of outcome or event A, given that outcome or event B has happened (or is true) is not the same as the probability of outcome B given that outcome A has happened. We need to take into account the <strong>prior</strong> probability that A has happened (the probability that A has happened is written P(A)). Bayes' rule can be written as follows:</p>
<p>$$P(A | B) = \frac{P(B|A) P(A)}{P(B)}$$</p>
<p>The quantity we ultimately want to calculate is the probability of dying at a particular age A, <em>given that</em> your family thinks you are left-handed. Let's write this in shorthand as P(A | LH). We also want the same quantity for right-handers: P(A | RH). As we go, we will figure out or approximate the other three quantities to find out what difference in age of death we might expect purely from the changing rates of left-handedness plotted above.</p>
<p>Here's Bayes' rule in the context of our discussion:</p>
<p>$$P(A | LH) = \frac{P(LH|A) P(A)}{P(LH)}$$</p>
<p>P(LH | A) is the probability that you are left-handed <em>given that</em> you died at age A. P(A) is the overall probability of dying at age A, and P(LH) is the overall probability of being left-handed. We will now calculate each of these three quantities, beginning with P(LH | A).</p>
<p>To calculate P(LH | A) for ages that might fall outside the original data, we will need to extrapolate the data to earlier and later years. Since the rates flatten out in the early 1900s and late 1900s, we'll use a few points at each end and take the mean to extrapolate the rates on each end. The number of points used for this is arbitrary, but we'll pick 10 since the data looks flat-ish until about 1910. </p>
```
# import library
# ... YOUR CODE FOR TASK 3 ...
import numpy as np
# create a function for P(LH | A)
def P_lh_given_A(ages_of_death, study_year = 1990):
""" P(Left-handed | ages of death), calculated based on the reported rates of left-handedness.
Inputs: numpy array of ages of death, study_year
Returns: probability of left-handedness given that subjects died in `study_year` at ages `ages_of_death` """
# Use the mean of the 10 last and 10 first points for left-handedness rates before and after the start
early_1900s_rate = lefthanded_data["Mean_lh"][-10:].mean()
late_1900s_rate = lefthanded_data["Mean_lh"][:10].mean()
middle_rates = lefthanded_data.loc[lefthanded_data['Birth_year'].isin(study_year - ages_of_death)]['Mean_lh']
youngest_age = study_year - 1986 + 10 # the youngest age is 10
oldest_age = study_year - 1986 + 86 # the oldest age is 86
P_return = np.zeros(ages_of_death.shape) # create an empty array to store the results
# extract rate of left-handedness for people of ages 'ages_of_death'
P_return[ages_of_death > oldest_age] = early_1900s_rate/100
P_return[ages_of_death < youngest_age] = late_1900s_rate/100
P_return[np.logical_and((ages_of_death <= oldest_age), (ages_of_death >= youngest_age))] = middle_rates/100
return P_return
```
## 4. When do people normally die?
<p>To estimate the probability of living to an age A, we can use data that gives the number of people who died in a given year and how old they were to create a distribution of ages of death. If we normalize the numbers to the total number of people who died, we can think of this data as a probability distribution that gives the probability of dying at age A. The data we'll use for this is from the entire US for the year 1999 - the closest I could find for the time range we're interested in. </p>
<p>In this block, we'll load in the death distribution data and plot it. The first column is the age, and the other columns are the number of people who died at that age. </p>
```
# Death distribution data for the United States in 1999
data_url_2 = "https://gist.githubusercontent.com/mbonsma/2f4076aab6820ca1807f4e29f75f18ec/raw/62f3ec07514c7e31f5979beeca86f19991540796/cdc_vs00199_table310.tsv"
# load death distribution data
# ... YOUR CODE FOR TASK 4 ...
death_distribution_data = pd.read_csv(data_url_2, sep = "\t", skiprows=[1])
# drop NaN values from the `Both Sexes` column
# ... YOUR CODE FOR TASK 4 ...
death_distribution_data = death_distribution_data.dropna(subset = ["Both Sexes"])
# plot number of people who died as a function of age
fig, ax = plt.subplots()
ax.plot("Age", "Both Sexes", data = death_distribution_data, marker='o') # plot 'Both Sexes' vs. 'Age'
ax.set_xlabel("Both Sexes")
ax.set_ylabel("Age")
```
## 5. The overall probability of left-handedness
<p>In the previous code block we loaded data to give us P(A), and now we need P(LH). P(LH) is the probability that a person who died in our particular study year is left-handed, assuming we know nothing else about them. This is the average left-handedness in the population of deceased people, and we can calculate it by summing up all of the left-handedness probabilities for each age, weighted with the number of deceased people at each age, then divided by the total number of deceased people to get a probability. In equation form, this is what we're calculating, where N(A) is the number of people who died at age A (given by the dataframe <code>death_distribution_data</code>):</p>
<p><img src="https://i.imgur.com/gBIWykY.png" alt="equation" width="220"></p>
<!--- $$P(LH) = \frac{\sum_{\text{A}} P(LH | A) N(A)}{\sum_{\text{A}} N(A)}$$ -->
```
def P_lh(death_distribution_data, study_year = 1990): # sum over P_lh for each age group
""" Overall probability of being left-handed if you died in the study year
Input: dataframe of death distribution data, study year
Output: P(LH), a single floating point number """
p_list = death_distribution_data["Both Sexes"]*P_lh_given_A(death_distribution_data["Age"], study_year) # multiply number of dead people by P_lh_given_A
p = np.sum(p_list) # calculate the sum of p_list
return p/np.sum(death_distribution_data["Both Sexes"]) # normalize to total number of people (sum of death_distribution_data['Both Sexes'])
print(P_lh(death_distribution_data, 1990))
```
## 6. Putting it all together: dying while left-handed (i)
<p>Now we have the means of calculating all three quantities we need: P(A), P(LH), and P(LH | A). We can combine all three using Bayes' rule to get P(A | LH), the probability of being age A at death (in the study year) given that you're left-handed. To make this answer meaningful, though, we also want to compare it to P(A | RH), the probability of being age A at death given that you're right-handed. </p>
<p>We're calculating the following quantity twice, once for left-handers and once for right-handers.</p>
<p>$$P(A | LH) = \frac{P(LH|A) P(A)}{P(LH)}$$</p>
<p>First, for left-handers.</p>
<!--Notice that I was careful not to call these "probability of dying at age A", since that's not actually what we're calculating: we use the exact same death distribution data for each. -->
```
def P_A_given_lh(ages_of_death, death_distribution_data, study_year = 1990):
""" The overall probability of being a particular `age_of_death` given that you're left-handed """
P_A = death_distribution_data["Both Sexes"][ages_of_death]/np.sum(death_distribution_data["Both Sexes"])
P_left = P_lh(death_distribution_data, study_year) # use P_lh function to get probability of left-handedness overall
P_lh_A = P_lh_given_A(ages_of_death, study_year) # use P_lh_given_A to get probability of left-handedness for a certain age
return P_lh_A*P_A/P_left
```
## 7. Putting it all together: dying while left-handed (ii)
<p>And now for right-handers.</p>
```
def P_A_given_rh(ages_of_death, death_distribution_data, study_year = 1990):
""" The overall probability of being a particular `age_of_death` given that you're right-handed """
P_A = death_distribution_data["Both Sexes"][ages_of_death]/np.sum(death_distribution_data["Both Sexes"])
P_right = 1 - P_lh(death_distribution_data, study_year)# either you're left-handed or right-handed, so P_right = 1 - P_left
P_rh_A = 1 - P_lh_given_A(ages_of_death, study_year) # P_rh_A = 1 - P_lh_A
return P_rh_A*P_A/P_right
```
## 8. Plotting the distributions of conditional probabilities
<p>Now that we have functions to calculate the probability of being age A at death given that you're left-handed or right-handed, let's plot these probabilities for a range of ages of death from 6 to 120. </p>
<p>Notice that the left-handed distribution has a bump below age 70: of the pool of deceased people, left-handed people are more likely to be younger. </p>
```
ages = np.arange(6, 120) # make a list of ages of death to plot
# calculate the probability of being left- or right-handed for each
left_handed_probability = P_A_given_lh(ages, death_distribution_data)
right_handed_probability = P_A_given_rh(ages, death_distribution_data)
# create a plot of the two probabilities vs. age
fig, ax = plt.subplots() # create figure and axis objects
ax.plot(ages, left_handed_probability, label = "Left-handed")
ax.plot(ages, right_handed_probability, label = "Right-handed")
ax.legend() # add a legend
ax.set_xlabel("Age at death")
ax.set_ylabel(r"Probability of being age A at death")
```
## 9. Moment of truth: age of left and right-handers at death
<p>Finally, let's compare our results with the original study that found that left-handed people were nine years younger at death on average. We can do this by calculating the mean of these probability distributions in the same way we calculated P(LH) earlier, weighting the probability distribution by age and summing over the result.</p>
<p>$$\text{Average age of left-handed people at death} = \sum_A A P(A | LH)$$</p>
<p>$$\text{Average age of right-handed people at death} = \sum_A A P(A | RH)$$</p>
```
# calculate average ages for left-handed and right-handed groups
# use np.array so that two arrays can be multiplied
average_lh_age = np.nansum(ages*np.array(left_handed_probability))
average_rh_age = np.nansum(ages*np.array(right_handed_probability))
# print the average ages for each group
# ... YOUR CODE FOR TASK 9 ...
print("Average age of lefthanded" + str(average_lh_age))
print("Average age of righthanded" + str(average_rh_age))
# print the difference between the average ages
print("The difference in average ages is " + str(round(average_lh_age - average_rh_age, 1)) + " years.")
```
## 10. Final comments
<p>We got a pretty big age gap between left-handed and right-handed people purely as a result of the changing rates of left-handedness in the population, which is good news for left-handers: you probably won't die young because of your sinisterness. The reported rates of left-handedness have increased from just 3% in the early 1900s to about 11% today, which means that older people are much more likely to be reported as right-handed than left-handed, and so looking at a sample of recently deceased people will have more old right-handers.</p>
<p>Our number is still less than the 9-year gap measured in the study. It's possible that some of the approximations we made are the cause: </p>
<ol>
<li>We used death distribution data from almost ten years after the study (1999 instead of 1991), and we used death data from the entire United States instead of California alone (which was the original study). </li>
<li>We extrapolated the left-handedness survey results to older and younger age groups, but it's possible our extrapolation wasn't close enough to the true rates for those ages. </li>
</ol>
<p>One thing we could do next is figure out how much variability we would expect to encounter in the age difference purely because of random sampling: if you take a smaller sample of recently deceased people and assign handedness with the probabilities of the survey, what does that distribution look like? How often would we encounter an age gap of nine years using the same data and assumptions? We won't do that here, but it's possible with this data and the tools of random sampling. </p>
<!-- I did do this if we want to add more tasks - it would probably take three more blocks.-->
<p>To finish off, let's calculate the age gap we'd expect if we did the study in 2018 instead of in 1990. The gap turns out to be much smaller since rates of left-handedness haven't increased for people born after about 1960. Both the National Geographic study and the 1990 study happened at a unique time - the rates of left-handedness had been changing across the lifetimes of most people alive, and the difference in handedness between old and young was at its most striking. </p>
```
# Calculate the probability of being left- or right-handed for all ages
left_handed_probability_2018 = P_A_given_lh(ages, death_distribution_data, 2018)
right_handed_probability_2018 = P_A_given_rh(ages, death_distribution_data, 2018)
# calculate average ages for left-handed and right-handed groups
average_lh_age_2018 = np.nansum(ages*np.array(left_handed_probability_2018))
average_rh_age_2018 = np.nansum(ages*np.array(right_handed_probability_2018))
# print the average ages for each group
print("Average age of lefthanded" + str(average_lh_age_2018))
print("Average age of righthanded" + str(average_rh_age_2018))
print("The difference in average ages is " +
str(round(average_lh_age_2018 - average_rh_age_2018, 1)) + " years.")
```
| github_jupyter |
This notebook was prepared by [Donne Martin](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# Challenge Notebook
## Problem: Given a list of stock prices on each consecutive day, determine the max profits with k transactions.
* [Constraints](#Constraints)
* [Test Cases](#Test-Cases)
* [Algorithm](#Algorithm)
* [Code](#Code)
* [Unit Test](#Unit-Test)
* [Solution Notebook](#Solution-Notebook)
## Constraints
* Is k the number of sell transactions?
* Yes
* Can we assume the prices input is an array of ints?
* Yes
* Can we assume the inputs are valid?
* No
* If the prices are all decreasing and there is no opportunity to make a profit, do we just return 0?
* Yes
* Should the output be the max profit and days to buy and sell?
* Yes
* Can we assume this fits memory?
* Yes
## Test Cases
<pre>
* Prices: None or k: None -> None
* Prices: [] or k <= 0 -> []
* Prices: [0, -1, -2, -3, -4, -5]
* (max profit, list of transactions)
* (0, [])
* Prices: [2, 5, 7, 1, 4, 3, 1, 3] k: 3
* (max profit, list of transactions)
* (10, [Type.SELL day: 7 price: 3,
Type.BUY day: 6 price: 1,
Type.SELL day: 4 price: 4,
Type.BUY day: 3 price: 1,
Type.SELL day: 2 price: 7,
Type.BUY day: 0 price: 2])
</pre>
## Algorithm
Refer to the [Solution Notebook](). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
## Code
```
from enum import Enum # Python 2 users: Run pip install enum34
class Type(Enum):
SELL = 0
BUY = 1
class Transaction(object):
def __init__(self, type, day, price):
self.type = type
self.day = day
self.price = price
def __eq__(self, other):
return self.type == other.type and \
self.day == other.day and \
self.price == other.price
def __repr__(self):
return str(self.type) + ' day: ' + \
str(self.day) + ' price: ' + \
str(self.price)
class StockTrader(object):
def find_max_profit(self, prices, k):
# TODO: Implement me
pass
```
## Unit Test
**The following unit test is expected to fail until you solve the challenge.**
```
# %load test_max_profit.py
import unittest
class TestMaxProfit(unittest.TestCase):
def test_max_profit(self):
stock_trader = StockTrader()
self.assertRaises(TypeError, stock_trader.find_max_profit, None, None)
self.assertEqual(stock_trader.find_max_profit(prices=[], k=0), [])
prices = [5, 4, 3, 2, 1]
k = 3
self.assertEqual(stock_trader.find_max_profit(prices, k), (0, []))
prices = [2, 5, 7, 1, 4, 3, 1, 3]
profit, transactions = stock_trader.find_max_profit(prices, k)
self.assertEqual(profit, 10)
self.assertTrue(Transaction(Type.SELL,
day=7,
price=3) in transactions)
self.assertTrue(Transaction(Type.BUY,
day=6,
price=1) in transactions)
self.assertTrue(Transaction(Type.SELL,
day=4,
price=4) in transactions)
self.assertTrue(Transaction(Type.BUY,
day=3,
price=1) in transactions)
self.assertTrue(Transaction(Type.SELL,
day=2,
price=7) in transactions)
self.assertTrue(Transaction(Type.BUY,
day=0,
price=2) in transactions)
print('Success: test_max_profit')
def main():
test = TestMaxProfit()
test.test_max_profit()
if __name__ == '__main__':
main()
```
## Solution Notebook
Review the [Solution Notebook]() for a discussion on algorithms and code solutions.
| github_jupyter |
# Non-ideal equations of state
This example demonstrates a comparison between ideal and non-ideal equations of state (EoS) using Cantera and CoolProp. The following equations of state are used to evaluate thermodynamic properties in this example
1. Ideal-gas EoS from Cantera
2. Non-ideal Redlich-Kwong EoS from Cantera
3. Helmholtz energy EoS from CoolProp
#### Import required packages (Cantera and CoolProp)
[CoolProp](http://coolprop.org) [1] is an open-source package that contains a highly-accurate database for thermophysical properties. The thermodynamic properties are obtained using pure and pseudo-pure fluid equations of state implemented for 122 components.
>1. I.H. Bell, J.Wronski, S. Quoilin, V. Lemort, 'Pure and Pseudo-pure Fluid Thermophysical Property Evaluation and the Open-Source Thermophysical Property Library CoolProp,' Industrial & Engineering Chemistry Research 53 (2014), https://pubs.acs.org/doi/10.1021/ie4033999
```
# Import Cantera
import cantera as ct
import numpy as np
%matplotlib notebook
import matplotlib.pyplot as plt
import time
print(f'Running Cantera version: {ct.__version__}')
from CoolProp.CoolProp import PropsSI
```
### Helper functions
This example uses CO$_2$ as the only species. The function `get_thermo_Cantera` calculates thermodynamic properties based on the thermodynamic state ($T$, $p$) of the species using Cantera. Applicable phases are `Ideal-gas` and `Redlich-Kwong`. The ideal-gas equation can be stated as
\begin{equation}
pv = RT,
\end{equation}
where $p$, $v$ and $T$ represent thermodynamic pressure, molar volume, and the temperature of the gas-phase. $R$ is the universal gas constant.
The function `get_thermo_CoolProp` utilizes the CoolProp package to evaluate thermodynamic properties based on the gas-phase, thermodynamic state ($T$, $p$) for a given fluid. The standard-reference thermodynamic states are different for Cantera and CoolProp, it is necessary to convert these values in an appropriate scale before comparison. Therefore, both functions `get_thermo_Cantera` and `get_thermo_CoolProp` return the thermodynamic values relative to the reference state obtained at 1 Bar, 300 K.
To plot the comparison of thermodynamic properties among three EoS, the `plot` function is used.
```
def get_thermo_Cantera(gas, T, p) :
states = ct.SolutionArray(gas, len(p))
X = 'CO2:1.0'
states.TPX = T, p, X
u = states.u
h = states.h
s = states.s
cp = states.cp
cv = states.cv
# Get the relative enthalpy, entropy and int. energy with referance to the first point
u = u - u[0]
s = s - s[0]
h = h - h[0]
return h, u, s, cp, cv
def get_thermo_CoolProp(T, p) :
n = len(p)
u = np.zeros(n)
h = np.zeros(n)
s = np.zeros(n)
cp = np.zeros(n)
cv = np.zeros(n)
for i in range(n):
#PropsSI("T","P",101325,"Q",0,"REFPROP::CO2")
u[i] = PropsSI("U","P", p[i],"T", T,"HEOS::CO2") # kJ/kg/K
h[i] = PropsSI("H","P", p[i],"T", T,"HEOS::CO2")
s[i] = PropsSI("S","P", p[i],"T", T,"HEOS::CO2")
cp[i] = PropsSI("C","P", p[i],"T", T,"HEOS::CO2")
cv[i] = PropsSI("O","P", p[i],"T", T,"HEOS::CO2")
# Get the relative enthalpy, entropy and int. energy with referance to the first point
u = u - u[1]
s = s - s[1]
h = h - h[1]
return h, u, s, cp, cv
def plot(T, p, a_Ideal, a_RK, a_CoolProp, name) :
line_width = 3
n = len(p)
plt.figure()
plt.plot(p/1e5, a_Ideal,'-',color='b', linewidth = line_width)
plt.plot(p/1e5, a_RK,'-',color='r', linewidth = line_width)
plt.plot(p/1e5, a_CoolProp,'-',color='k', linewidth = line_width)
plt.xlabel('Pressure [Bar]')
plt.ylabel(name)
plt.legend(['Ideal EoS','RK EoS', 'CoolProp'], prop={'size': 14}, frameon=False)
```
## 1. EoS Comparison based on thermodynamic properties
This is the main subroutine that compares and plots the thermodynamic values obtained using three equations of state.
```
# Main function
# Input parameters
T = 300 # Temperature is constant [unit:K]
p = 1e5*np.linspace(1, 100, 1000) # Pressure is varied from 1 to 100 bar [unit:Pa]
# Read the ideal gas-phase
gasIdeal = ct.Solution('data/co2-thermo.yaml', 'CO2-Ideal', transport=None)
[hIdeal, uIdeal, sIdeal, cpIdeal, cvIdeal] = get_thermo_Cantera(gasIdeal, T, p)
# Read the R-K gas-phase
gasRK = ct.Solution('data/co2-thermo.yaml', 'CO2-RK', transport=None)
[hRK, uRK, sRK, cpRK, cvRK] = get_thermo_Cantera(gasRK, T, p)
# Read the thermo data using CoolProp
[hCoolProp, uCoolProp, sCoolProp, cpCoolProp, cvCoolProp] = get_thermo_CoolProp(T, p)
# Plot the result
# Internal energy
plot(T, p, uIdeal, uRK, uCoolProp, "Relative Internal Energy [J/kg]")
# Enthalpy
plot(T, p, hIdeal, hRK, hCoolProp, "Relative Enthalpy [J/kg]")
# Entropy
plot(T, p, sIdeal, sRK, sCoolProp, "Relative Entropy [J/K]")
```
The thermodynamic properties such as internal energy, enthalpy, and entropy are plotted against the operating pressure at a constant temperature $T = 300$ K. The three equations follow each other closely at low pressures ($P < 10$ Bar). However, the ideal gas EoS departs significantly from the observed behavior of gases near the critical regime ($P_{\rm {crit}} = 73.77$ Bar).
The ideal gas EoS does not consider inter-molecular interactions and the volume occupied by individual gas particles. At low temperatures and high pressures, inter-molecular forces become particularly significant due to a reduction in inter-molecular distances. Additionally, at high pressures, the volume of individual molecules becomes significant. Both of these factors contribute to the deviation from ideal behavior at high pressures. The cubic Redlich-Kwong EoS, on the other hand, predicts thermodynamic properties accurately near the critical regime.
```
# Specific heat at constant pressure
plot(T, p, cpIdeal, cpRK, cpCoolProp, "Cp [J/kg/K]")
# Specific heat at constant volume
plot(T, p, cvIdeal, cvRK, cvCoolProp, "Cv [J/kg/K]")
```
In the case of Ideal gas EoS, the specific heats at constant pressure ($C_{\rm p}$) and constant volume ($C_{\rm v}$) are independent of the pressure. Hence, $C_{\rm p}$ and $C_{\rm v}$ for ideal EoS do not change as the pressure is varied from $1$ Bar to $100$ Bar in this study.
Specific heat at constant pressure ($C_{\rm p}$) for R-K EoS follow the trend closely with the Helmholtz EoS to the critical regime. Although the specific heat at constant pressure ($C_{\rm p}$) shows reasonable agreement with the Helmholtz EoS in sub-critical and supercritical regimes, it inaccurately predicts a very high value near the critical point. The heat capacity at constant pressure ($C_{\rm p}$) at the critical point is finite. The sudden rise in $C_{\rm p}$ value in the case of R-K EoS is just a numerical artifact and not a real singularity.
The specific heat at constant volume ($C_{\rm v}$), on the other hand, predicts smaller values in the subcritical and critical regime. However, it shows completely wrong values in the super-critical region, making it invalid at very high pressures. It is well known that the cubic equations typically fail to predict accurate constant-volume heat capacity in the transcritical region [2]. Certain cubic EoS models have been extended to resolve this discrepancy using crossover models. For further information, see the work of Span [2] and Saeed et al [3].
>2. R. Span, 'Multiparameter Equations of State - An Accurate Source of Thermodynamic Property Data', Springer Berlin Heidelberg (2000), http://dx.doi.org/10.1007/978-3-662-04092-8
>3. A. Saeed, S. Ghader,'Calculation of density, vapor pressure and heat capacity near the critical point by incorporating cubic SRK EoS and crossover translation', Fluid Phase Equilibria (2019) 493, https://doi.org/10.1016/j.fluid.2019.03.027
## 2. Temperature-Density plots
Following function plots the $T-\rho$ diagram over a wide pressure and temperature range. The temperature is varied from 250 K to 400 K. The pressure is changed from 1 Bar to 600 Bar.
```
# Input parameters
# Set up arrays for pressure and temperature
p_array = [1] + [i for i in np.arange(10,101,30)] + [i for i in np.arange(200,601,100)] # Pressure is varied from 1 to 600 bar
T_array = [i for i in np.arange(250,401,10)] # Temperature is varied from 250K to 400K
# Read ideal and non-ideal phases
gasIdeal = ct.Solution('data/co2-thermo.yaml', 'CO2-Ideal', transport=None) # ideal gas-phase
gasRK = ct.Solution('data/co2-thermo.yaml', 'CO2-RK', transport=None) # R-K gas-phase
# Loop over temperature and pressure
density_coolprop = np.zeros(len(T_array))
# Plot
plt.rcParams['axes.labelsize'] = 14
plt.figure(figsize=(10,5))
cmap= plt.get_cmap()
colors = iter(cmap(np.linspace(0,1,len(p_array))))
for i in range(len(p_array)):
p = 1e5* p_array[i]
states = ct.SolutionArray(gasIdeal, len(T_array))
states.TP = T_array, p
density_Ideal = states.density_mass
states = ct.SolutionArray(gasRK, len(T_array))
states.TP = T_array, p
density_RK = states.density_mass
for j in range(len(T_array)):
density_coolprop[j] = PropsSI('D','P', p,'T',T_array[j],'HEOS::CO2')
colr = next(colors)
pBar = p/1e5;
plt.plot(density_Ideal, T_array,'--',color=colr, label= pBar)
plt.plot(density_RK, T_array,'o',color=colr)
plt.plot(density_coolprop, T_array,'-',color=colr)
plt.text(-50, 320, 'p = 1 [Bar]', color='darkblue', rotation = 'vertical', size = '14')
plt.text(300, 308, 'p = 70 [Bar]', color= 'c', size = '14', rotation = '-5')
plt.text(950, 320, 'p = 600 [Bar]', color=colr, size = '14', rotation = '-55')
plt.xlabel('Density [kg/m3]')
plt.ylabel('Temperature [K]')
plt.legend(['Ideal EoS','RK EoS', 'CoolProp'], prop={'size': 14}, frameon=False)
```
The figure compares $T-\rho$ plots for ideal, R-K, and Helmholtz EoS at different operating pressures. All three EoS yield the same plots at low pressures (0 Bar and 10 Bar). However, the Ideal gas EoS departs significantly at high pressures ($P > 10$ Bar), where non-ideal effects are prominent. The R-K EoS closely matches the Helmholtz EoS at supercritical pressures ($P \ge 70$ Bar). However, it does depart in the liquid-vapor region that exists at $P < P_{\rm {crit}}$ and low temperatures (~$T_{\rm {crit}}$).
| github_jupyter |
#SHDOM single-scattering adjoint
This folder conatains the most recent (3 May 2015) single-scattering adjoint calculations from frank.
>This has the log output files (.log), radiance output files (.arad),
and optical property adjoint files (.adj) for the "true" and "guess"
cases. The first set of output is for the single scattering forward radiance calculations
>>les0822nh15t13y135_ocaer1_w0.646_ns1true.*,
>>les0822nh15t13y135_ocaer1_w0.646_ns1guess.*,
>and the second set of outputs is for the full multiple scattering forward radiance calculations,
>>les0822nh15t13y135_ocaer1_w0.646_ns1true2.*,
>>les0822nh15t13y135_ocaer1_w0.646_ns1guess2.*.
The cloud field used in these simulations is a 2D slice taken at $y_{(i_y=135)}=8.375 \mathrm{km}$ of the 320x320 high resolution simulation shown below.

The adjoint calculation is for the gradient of the misfit function,
\begin{align}
\frac{\partial \Phi(\sigma, \omega)}{\partial(\sigma)} &= \left < \Delta p, \mathcal{U}_{\sigma}[\Delta f] \right >
\end{align}
Only the first order of scattering completed so the results are approximate. In the calculation labelled "1" the forward and adjoint calculations are only single-scattering. In the calculation labelled "2", the Forward calculation includes multiple scatterng while the adjoint does not.
```
%pylab inline
# Imports
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
import pandas as pd
# Utility functions
def print_head(fname, Nlines=10, indent="\t"):
"Print the head of the file."
# Print a message and then the first N lines
print("Showing head: {}".format(fname))
for i, line in zip(range(Nlines), open(fname, 'r')):
print(indent + line.strip())
print('\n')
ls
```
# Looking at the data files
```
# The log file
flog = "les0822nh15t13y135_w0.646_ns1true2.log"
print_head(flog, Nlines=10)
# The radiance file
frad = "les0822nh15t13y135_w0.646_ns1true2.arad"
print_head(frad, Nlines=25)
# The adjoint file
fadj_true = "les0822nh15t13y135_w0.646_ns1true2.adj"
fadj = "les0822nh15t13y135_w0.646_ns1guess2.adj"
print_head(fadj, Nlines=35)
# The adjoint Radiance file
farad = "les0822nh15t13y135_w0.646_ns1guess2.arad"
farad_true = "les0822nh15t13y135_w0.646_ns1true2.arad"
print_head(farad, Nlines=35)
# Load the adjoint files into memory
adj_frame = pd.read_csv(fadj, quoting=3, delim_whitespace=True, skiprows=2)
adj_true_frame = pd.read_csv(fadj_true, quoting=3, delim_whitespace=True, skiprows=2)
adj_arad_frame = pd.read_csv(farad, quoting=3, delim_whitespace=True, skiprows=19)
adj_arad_true_frame = pd.read_csv(farad_true, quoting=3, delim_whitespace=True, skiprows=19)
# Get variables from the array
nx = adj_frame['X'].unique().size
nz = adj_frame["Z"].unique().size
# Get the guess case adjoint field properties
adj_x = np.array(adj_frame["X"], dtype='f8').reshape(nx, nz)
adj_z = np.array(adj_frame["Z"], dtype='f8').reshape(nx, nz)
adj_ext = np.array(adj_frame["Extinct"], dtype='f8').reshape(nx, nz)
adj_alb = np.array(adj_frame["SSalbedo"], dtype='f8').reshape(nx, nz)
adj_dMFdext = np.array(adj_frame["dMF/dExt"], dtype='f8').reshape(nx, nz)
adj_dMFdalb = np.array(adj_frame["dMF/dSSalb"], dtype='f8').reshape(nx, nz)
# Get the True case adjoint extinction field
adj_ext_true = np.array(adj_true_frame["Extinct"], dtype='f8').reshape(nx, nz)
# Define the adjoint radiance parameters
adj_arad = np.array(adj_arad_frame['RADIANCE'], dtype='f8').reshape(nx)
adj_arad_true = np.array(adj_arad_true_frame['RADIANCE'], dtype='f8').reshape(nx)
# Check the sign of dMF/dExt for Z near the top of the atmosphere
adj_frame[abs(adj_frame["Z"]-15) < .2]
# Plot uniformity parameters
H = 20 #5.5 # top of the atmosphere plots
mask_clouds = (adj_ext>= .75e-5) #* (adj_z <= 2.83)
cloud_outline = (adj_ext>= .75e-1) #* (adj_z <= 2.83)
# Scaling of the verticle coordinate
Zscale = adj_z[0,:]
DZscale = 0 * Zscale
DZscale_other = 0 * Zscale
DZscale[:-1] += 1 / (Zscale[1:] - Zscale[:-1])
DZscale_other[1:] += 1 / (Zscale[1:] - Zscale[:-1])
alpha = .99725
DZscale = alpha * DZscale + (1-alpha) * DZscale_other
# Compute the residual
adj_residual = adj_arad_true - adj_arad
# Make a figure to plot the true extinction field
f0 = plt.figure(0, (15,8), facecolor='white')
ax0 = f0.add_axes([.1, .1, .8, .8])
contour0 = ax0.contourf(adj_x, adj_z, adj_ext_true, 100, cmap=cm.gray)
ax0.set_ybound((0,8))
ax0.set_title(" Guess for the extinction field at y=8.35")
f0.colorbar(contour0, ax=ax0)
# Make a figure to plot the true extinction field
f0 = plt.figure(0, (15,8), facecolor='white')
ax0 = f0.add_axes([.1, .1, .8, .8])
fvals = adj_ext_true - adj_ext
contour0 = ax0.contourf(adj_x, adj_z, fvals, 100, cmap=cm.gray)
ax0.set_ybound((0,8))
ax0.set_title("Difference $\sigma_{\mathrm{true}} - \sigma_{\mathrm{guess}}$")
f0.colorbar(contour0, ax=ax0)
# Make a figure to plot the true single-scattering albedo field
f1 = plt.figure(1, (15,8), facecolor='white')
ax1 = f1.add_axes([.1, .1, .8, .8])
levels = np.linspace(.999995, 1, 100)
contour1 = ax1.contourf(adj_x, adj_z, adj_alb, levels=levels, cmap=cm.gray)
ax1.set_ybound((0,13))
ax1.set_title("SSAlbedo field at y=8.35")
f1.colorbar(contour1, ax=ax1)
# Make a figure to plot the true single-scattering albedo field
f2 = plt.figure(1, (10,6), facecolor='white')
ax2 = f2.add_axes([.1, .1, .8, .8])
fvals = adj_dMFdext * mask_clouds * DZscale * adj_ext# * .1 * adj_ext
fmax = abs(fvals).max() / 20.
levels = np.linspace(-fmax, fmax, 400)
contour2 = ax2.contourf(adj_x, adj_z, fvals, levels=levels, cmap=cm.RdBu_r, extend="both") #levels=linspace(-16, 16, 100),
contour2a = ax2.contour(adj_x, adj_z, cloud_outline, 1, cmap=cm.gray) #levels=linspace(-16, 16, 100),
#contour2b = ax2.contourf(adj_x, adj_z, cloud_outline * fvals, 10 , cmap=cm.RdBu_r, linewidth=3) #levels=linspace(-16, 16, 100),
# Add the residual to the plot
ftrans = lambda y: 200 * y + .8*H
plot_overlay_origin = ftrans(0 * adj_residual)
plot_overlay_arad = ftrans(adj_residual)
plot_res_pos = ax2.fill_between(adj_x[:,0], plot_overlay_origin, plot_overlay_arad,
where=(plot_overlay_origin<plot_overlay_arad),
color='b')#, linewidth=3)
plot_res_neg = ax2.fill_between(adj_x[:,0], plot_overlay_origin, plot_overlay_arad,
where=(plot_overlay_origin>plot_overlay_arad),
color='r',)#, linewidth=3)
plot_res_line = ax2.plot(adj_x[:,0], plot_overlay_arad, 'k', linewidth=2)
plot_origin = ax2.plot(adj_x[:,0], plot_overlay_origin, 'k,', linewidth=1)
ax2.set_ybound((0,H))
ax2.set_ylabel("Z Height [km]", fontsize='x-large', fontweight='bold')
ax2.set_xlabel('X [km]', fontsize='large', fontweight='bold')
ax2.set_title("""
Risidual ($y_{\mathrm{data}} - y_{\mathrm{model}}(\mathbf{a})$) and derivative (-$\mathrm{d}\Phi(\sigma) / \mathrm{d}\log\sigma$)
""", fontsize='xx-large', fontweight='bold')
cb = f2.colorbar(contour2, ax=ax2)
cb.set_ticks([-fmax, 0, fmax])
f2.savefig("adjoint-derivative-single-scattering.jpg", dpi=300)
#cb.set_ticklabels(['-2', '-1', '0', '1', '2'])
# Make a figure to plot the true single-scattering albedo field
f2 = plt.figure(1, (10,6), facecolor='white')
ax2 = f2.add_axes([.1, .1, .8, .8])
fvals = -adj_dMFdext * mask_clouds * DZscale * adj_ext# * .1 * adj_ext
fvals = - (adj_ext_true - adj_ext) #* adj_ext
fmax = abs(fvals).max() / 1.
levels = np.linspace(-fmax, fmax, 200)
contour2 = ax2.contourf(adj_x, adj_z, fvals, levels=levels, cmap=cm.RdBu_r, extend="both") #levels=linspace(-16, 16, 100),
contour2a = ax2.contour(adj_x, adj_z, cloud_outline, 1, cmap=cm.gray) #levels=linspace(-16, 16, 100),
#contour2b = ax2.contourf(adj_x, adj_z, cloud_outline * fvals, 10 , cmap=cm.RdBu_r, linewidth=3) #levels=linspace(-16, 16, 100),
# Add the residual to the plot
ftrans = lambda y: 200 * y + .8*H
plot_overlay_origin = ftrans(0 * adj_residual)
plot_overlay_arad = ftrans(adj_residual)
plot_res_pos = ax2.fill_between(adj_x[:,0], plot_overlay_origin, plot_overlay_arad,
where=(plot_overlay_origin<plot_overlay_arad),
color='b')#, linewidth=3)
plot_res_neg = ax2.fill_between(adj_x[:,0], plot_overlay_origin, plot_overlay_arad,
where=(plot_overlay_origin>plot_overlay_arad),
color='r',)#, linewidth=3)
plot_res_line = ax2.plot(adj_x[:,0], plot_overlay_arad, 'k', linewidth=2)
plot_origin = ax2.plot(adj_x[:,0], plot_overlay_origin, 'k,', linewidth=1)
ax2.set_ybound((0,H))
ax2.set_ylabel("Z Height [km]", fontsize='x-large', fontweight='bold')
ax2.set_xlabel('X [km]', fontsize='large', fontweight='bold')
ax2.set_title("""
Risidual ($y_{\mathrm{data}} - y_{\mathrm{model}}(\mathbf{a})$) and extinction error ($\sigma_{\mathrm{true}}-\sigma_{\mathrm{guess}})$)
""", fontsize='xx-large', fontweight='bold')
cb = f2.colorbar(contour2, ax=ax2)
cb.set_ticks([-fmax, 0, fmax])
f2.savefig("adjoint-extinction-difference-and-residual.jpg", dpi=300)
#cb.set_ticklabels(['-2', '-1', '0', '1', '2'])
plot(Zscale, DZscale)
```
| github_jupyter |
```
<h6>Corrections to Udacity Course</h6>
The DL Udacity class was originally created by Vincent Vanhoucke, a research scientist and tech lead for Google Brain.
This was a free class. Udacity removed some of Vincent's content, cut it up into 5 modules and added their own content
from presented by youtube stars.
<h6>Corrections to Uddacity Section1 Introduction</6>
Welcome, Anaconda, Jupyter, Applying DL, Regression Videos.
<h6>Corrections to Udacity Section2 Neural Networks</h6>
<p>
There are 6 sections here, 1) Matrix Math and NumPy Refresher, 2) Intro to NN,
3) your First Neural Network assignment, 4) Model Evaluation and Validation,
5) Sentiment Analysis with Andrew Trask, 6)MiniFlow.
I viewed the Intro to NN, First NN assignment, Model Evaluation and Validation videos
</p>
<h6>Corrections for Section 2.2 NN - Intro to NN</h6>
<p>This section is added by Udacity. There are no presentations by Vincent here. </p>
</p>In this section Udacity presents examples for AND/OR/NOT/XOR perceptrons. A better idea would have
been to write a MLP for a XOR gate and then to the IRIS example. This gives you a way to see numerical
values and to use them to build intuition and debug tools. Difficult if not impossible to do this w/image data.
They do not include code for a working XOR gate.</p>
<p> Gradient descent; explain and code Gradient Descent. This is useless and misleading. DL does not use GD,
it uses SGD which is very different. SGD is covered in the CNN-Intro to Tensorflow section.
</p>
<p>Model Evaluation and Validation : This presentation uses linear regression for both regression and classification.
They present a numerical dataset and not a categorical one.
They present an exercise in filling out Confusion matrix for regression. There is no mention of AUC/ROC.
Logistic regression more popular here. Doesn't seem apparant the presenter knows the difference.
with a probability interpretation. Logistic Regression is important to develop to explain softmax and to understand
the FC layer at the end of CNNs.
They cover accuracy vs. errors.
They present over and underfitting and k-fold cross validation as a way to fix.
</p>
<p>Miniflow. Did not look at this. Code up a python version of tf. If the strategy is to train someone to become an open
source contributor we should focus on gradients and how to calculate them especially in a distributed
environement. </p>
<h6>Corrections for Section2 CNN </h6>
There are 10 modules here, 1) Intro to TF, 2) Cloud Computing, 3) Deep Neural Networks, 4) Convolutional Networks,
5) Siraj's Image Classification,
6) Weight Initialization, 7) Image Classification, 8) Siraj's Image Generation,
9) AutoEncoders, 10) Transfer Learning in tf.
I did not comment on all of the sections above. Only selected ones.
<h6>Section 1: Intro to TF</h6>
<p>There are 29 sections here. </p>
<h6>Section 3: Deep Neural Networks</h6>
<p>There are 13 sections in this module. All of these lectures are from Vincent. Udacity does the intro lecture
and adds in some sample code sections. </p>
<p>Module 3 is a mix between Udacity doing an intro lecture and Vincent</p>
<p>Module 3.2-3.3 cover a 2 layer network and RELUs. They don't cover why this solves the vanishing
gradient problem with logistic/sigmoids. The goal of this module seems to be programming NNs in TF.
</p>
<p>Module 3.4 no lecture, covers TF code to classify MNIST digits. My preference is to start with a simpler example
using XOR gates or IRIS b/c it allows for debugging and separates the formatting of convolutional layers
away from NNs. The key is to focus first on a simple numerical example both with regression and classification. These
are common interview questions. </p>
<p>Vincent Module 3.5. Training a DNN and how adding layers for CNNS increase the capacity and is more efficient
in terms of parameter efficiency and the deeper layers capture hierarchial/neighboring image effects. Initial layers
in a CNN capture lines, higher layers capture partial shapes and next layers capture representations of objects.</p>
<img src="vincent35.png" /> Note to self: add resnets and skip layers and show deep vanishing gradients,
<p>Module 3.6 no lecture covers model saving and restoring. Add details on graph/saving restoring. The simplifiction
is they only cover saving and restoring variables, specifically initial weights and biases. They cover saving
the final model and then restoring the final model and running an accuracy operation on test data. Saving variables is
never used. They fail to explain the difference between graphs/models/variables and when/why you would do this.</p>
<p>>Module 3.7 Finetuning, unclear if this is relevant or usable. Goal is to finetune a model. They present to do this
you have to name the weights and biases with names and not let TF create variable names. If tf saves and restores
a session there is no way for tf to know Variable0 is a saved variable so it fails to load a valid value. Udacity added
content. </p>
<p>Vincent Module 3.8 Regularization Intro. Make model bigger than necessary and take steps to not overfit. Hard
to design exactly size of NN for data. Skinny Jeans problem, hard to get network to exactly fit like jeans. Try bigger
pants and then prevent overfitting. </p>
<p>Vincent Module 3.9 - Regularization. 1) Early termination still best way, 2) regularization, adding
artificial constraints. L2 regulartization,add another term to prevent large weights. Some of the value of the
weights go into the regularization term. </p>
<p>Vincent Module 3.10 derivative of regularizatoin term beta*1/2*norm weight^2 = w. </p>
<p>Vincent Module 3.11 - dropout. Dropout some of the activations. Set 1/2 to 0. Random. Hintons idea. The network
can not rely on any given activation so it has to add some redundancies. IN practice this makes things more robust
makes the network seem like ensemble. If dropout not working then you should probably using a bigger network. </p>
<p>Vincent Module 3.12 - Dropout trick. Not only do you zero out 1/2 the activatoins but you scale the remaining
ones by a factor of 2. 2x the ones not zeroed out. </p><img src="vincentdropout2.png">
<p>No mention if dropout is as good as early termination although most code samples seem to prefer dropout
over early termination. </p>
<h6>Section 4: Convolutional Networks</h6>
<p>There are 35 sections in this module. </p>
Intro to CNNs, Color, Statistical Invariance, Convolutional Networks, Intuition, Filters, Feature Map Sizes, Convolutions
continued, Parameteres, Convolution Output Shape(2), Number of parameters(2), parameter sharing, Visualizing CNNs,
Tensorflow Convolution Layer, Explore Design Space, TF Max Pooling, Pooling intuition(2), Pooling mechanics(2), Pooling
practice(2), Average Pooling(2),1x1 convolutions, Inception module, Convolution Network in TF, TF convolution Layer(2),
TF pooling layer(2), additional resources.
<h6>Section 4: Weight Initialization</h6>
<p>There are 6 sections here presented by Udacity. This material is covered in Intro to TF by Vincent. There
are gaps which we can fill in from the Hinton Coursera material. Weight histograms and how to debug if weights
are not small enough. </p>
<h6>Section 9: Siraj's Image Generation</h6>
<p>Siraj covers VAEs. There is no mention of generative models, calls autoencoders unsupervised models. Says AE is
used for compression but lossy. Says VAEs are bayesian inference. All points of uncertainity are expressed w/probability.
He presents Bayes rule and describes prior, posterior and marginal probabilities. There is no mention of what the
variables actually are in his presentation. No mention of a posterior, prior, likelihood in his verbal presentation.
At any time there is evidence for and against something. When you learn something
new you have to fold this new evidence into what you already know. You create a new probability. Siraj quote for his VAE
code which says he uses relu to squash the dimensionality. This is an inaccurate statement. The relu is a substibute
for a logistic which creates a probablity distribution between 0-1. Squashing is not a technical term. </p>
<img src="siraj_imagegen1.png">
<img src="siraj_imagegen2.png">
<img src="siraj_imagegen3.png">
<h6>Module 4: RNN</h6>
<p>There are 12 sections in this module, Intro to RNN, Siraj's Stock predictin, Hyperparameters, Embeddings and
Word2Vec, Siraj's Style transfer, Q/A with Floydhub, Sentiment prediction RNN, Siraj's text summaratin, generate
TV Scripts, Sequence to Sequence, Siraj;s chatbot, Translation project </p>
<p></p>
<h6>Corrections to Udacity Lesson11</h6>
The png below lists links listing chatbots and machine traslation. Those are 2 fundamentally different implementations
A chatbot closely resembles a QA system while a ML application is a different architecture, similiar to performing
text alignment across language models.
The links also include a dynamic memory network and attention which they do not cover in the lectures.
<img src="UdacityCorrectionDeepLearningLesson12.png">
<h6>Corrections to Udacity Section4 Recurrent Neural Networks</h6>
<p>Missing an overview of sequence to sequence architectures</p>
<p>Missing an explanation of how LSTM solves vanishing/exploding gradient problem. </p>
<h6>Lesson 1 section 2: vanishing and exploding gradients when values are less than or greater than one.
</h6><p>This is not accurate. These are matricies so you have to talk about singular values of matricies not
numbers like they are scalars. </p>
<><>
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/gdrive')
```
#**Importing Packages**
```
import pandas as pd
import numpy as np
from keras.models import load_model
from keras.models import Sequential
from keras.layers import Dense, Conv1D, GRU, Bidirectional, Dropout
from keras.utils import plot_model
import matplotlib.pyplot as plt
import time
import matplotlib.pyplot as plt
%matplotlib inline
from IPython.display import display
import datetime
import math
import warnings
warnings.filterwarnings("ignore")
import glob
from sklearn.metrics import mean_squared_error, mean_absolute_error
```
### *Initializing Window size*
```
window_size = 10
```
# **Functions**
```
def read_label():
label = {}
for i in range(1,7):
hi = '/content/gdrive/MyDrive/energy_disaggregation/windowGRU/data/house_{}/labels.dat'.format(i)
label[i] = {}
with open(hi) as f:
for line in f:
splitted_line = line.split(' ')
label[i][int(splitted_line[0])] = splitted_line[1].strip() + '_' + splitted_line[0]
return label
#-------------------------------------------------------------------------------------------------------------------
def read_merge_data(house):
path = '/content/gdrive/MyDrive/energy_disaggregation/windowGRU/data/house_{}/'.format(house)
file = path + 'channel_1.dat'
df = pd.read_table(file, sep = ' ', names = ['unix_time', labels[house][1]],
dtype = {'unix_time': 'int64', labels[house][1]:'float64'})
num_apps = len(glob.glob(path + 'channel*'))
for i in range(2, num_apps + 1):
file = path + 'channel_{}.dat'.format(i)
data = pd.read_table(file, sep = ' ', names = ['unix_time', labels[house][i]],
dtype = {'unix_time': 'int64', labels[house][i]:'float64'})
df = pd.merge(df, data, how = 'inner', on = 'unix_time')
df['timestamp'] = df['unix_time'].astype("datetime64[s]")
df = df.set_index(df['timestamp'].values)
df.drop(['unix_time','timestamp'], axis=1, inplace=True)
return df
#-------------------------------------------------------------------------------------------------------------------
def uniform_label(df):
if 'kitchen_outlets_7' in df.columns:
df = df.rename(columns={'kitchen_outlets_7': 'kitchen'})
if 'kitchen_outlets_5' in df.columns:
df = df.rename(columns={'kitchen_outlets_5': 'kitchen'})
if 'kitchen_outlets_3' in df.columns:
df = df.rename(columns={'kitchen_outlets_3': 'kitchen'})
if 'kitchen_outlets_24' in df.columns:
df = df.rename(columns={'kitchen_outlets_24': 'kitchen'})
return df
#----------------------------------------------------------------------------------------------------------------------
def add_houses(df_list, appliance):
merged_df = pd.DataFrame()
for df in df_list:
df = uniform_label(df)
df["aggregate_mains"] = df['mains_1'] + df['mains_2']
df = df[['aggregate_mains',appliance]]
merged_df = pd.concat([merged_df,df])
return merged_df
#-----------------------------------------------------------------------------------------------------------------------
def max_value(df):
return max(df['aggregate_mains'])
#-----------------------------------------------------------------------------------------------------------------------
def preprocess_data(df, appliance, nmax):
df['aggregate_mains'] = df['aggregate_mains']/ nmax
df[appliance] = df[appliance]/ nmax
return df
#----------------------------------------------------------------------------------------------------------------------
def create_window_chunk(df, sequence_length):
result = []
for index in range(len(df) - sequence_length):
result.append(df[index: index + sequence_length])
return result
#----------------------------------------------------------------------------------------------------------------------
def _create_model(window_size):
'''Creates the GRU architecture described in the paper
'''
model = Sequential()
# 1D Conv
model.add(Conv1D(16, 4, activation='relu', input_shape=(window_size,1), padding="same", strides=1))
#Bi-directional GRUs
model.add(Bidirectional(GRU(64, activation='relu', return_sequences=True), merge_mode='concat'))
model.add(Dropout(0.5))
model.add(Bidirectional(GRU(128, activation='relu', return_sequences=False), merge_mode='concat'))
model.add(Dropout(0.5))
# Fully Connected Layers
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(1, activation='linear'))
model.compile(loss='mse', optimizer='adam')
print(model.summary())
return model
#---------------------------------------------------------------------------------------------------------------------
```
# **Data Preprocessing**
```
df_dic = {}
dates = {}
```
### *Reading and storing labels of all the houses in a dictionary*
```
labels = read_label()
labels
```
### *Converting raw dataset of a house into Pandas Dataframe and storing all the dataframes in a dictionary*
```
for i in range(1,7):
df_dic[i] = read_merge_data(i)
print(df_dic)
```
### *Checking the Number of days of available data of different houses*
```
for i in range(1,7):
dates[i] = [str(time)[:10] for time in df_dic[i].index.values]
dates[i] = sorted(list(set(dates[i])))
print('House {0} data contain {1} days from {2} to {3}.'.format(i,len(dates[i]),dates[i][0], dates[i][-1]))
print(dates[i], '\n')
for i in range(1,7):
print('House {} data has shape: '.format(i), df_dic[i].shape)
display(df_dic[i].tail(3))
```
### *Preparing training data - house 1 & 4, Appliance- Kitchen_outlet*
```
df_list = [df_dic[1], df_dic[4]]
dfs = add_houses(df_list, 'kitchen')
dfs
```
### *Creating window chunk and normalizing dataframe*
```
nmax = max_value(dfs)
df = preprocess_data(dfs, 'kitchen', nmax)
df
sequence_length = window_size
x = create_window_chunk(df['aggregate_mains'], sequence_length)
y = df['kitchen'][sequence_length:]
x = np.array(x)
y = np.array(y)
```
### *reshaping the data*
```
x_train = np.reshape(x, (x.shape[0], x.shape[1], 1))
```
# **Creating Model**
```
model = _create_model(window_size)
```
###*training model*
```
start = time.time()
model.fit(x_train, y, epochs=2, batch_size=128, shuffle=True, validation_split=0.33)
print('Finish trainning. Time: ', time.time() - start)
```
###*Saving the trained model*
```
model.save("gdrive/MyDrive/saved_model/kitchen_outlet_train_1_4_test_5_6_epochs_2_batch_128_window_10_validation_split_0.33_model.h5")
```
### *Preparing and preprocessing test data - house 5 & 6*
```
df_list = [df_dic[5],df_dic[6]]
dfs = add_houses(df_list, 'kitchen')
df_test = preprocess_data(dfs, "kitchen", nmax)
print(df_test.head())
sequence_length = window_size
x_test = create_window_chunk(df_test['aggregate_mains'], sequence_length)
y_test = df_test['kitchen'][sequence_length:]
x_test = np.array(x_test)
y_test = np.array(y_test)
x_test = np.reshape(x_test, (x_test.shape[0], x_test.shape[1], 1))
```
###*Predicting on the test*
```
pred_test = model.predict(x_test, batch_size=128)
pred_test = np.reshape(pred_test, (len(pred_test)))
```
### *Denormalized the test and predicted data*
```
y_test_d = y_test * nmax
y_pred_d = pred_test* nmax
```
#**Data Visualization**
```
plt.figure(figsize = (30, 8))
plt.plot(y_test, color='green', label="ground kitchen outlet")
plt.plot(pred_test, color='blue', label="predicted kitchen outlet")
plt.legend
plt.show
```
###*More Zooming into the graph*
```
plt.figure(figsize = (30, 8))
plt.plot(pred_test[50000:100000], color='green', label="predicted kitchen outlet")
plt.plot(y_test[50000:100000], color='blue', label="ground kitchen outlet")
plt.legend
plt.show
```
#**Evaluation Metrics**
```
# Calculating MAE on test data
mean_absolute_error(y_test_d, y_pred_d)
# Calculating RMSE on test data
math.sqrt(mean_squared_error(y_test_d, y_pred_d))
```
| github_jupyter |
# GRU 231
* Operate on 16000 GenCode 34 seqs.
* 5-way cross validation. Save best model per CV.
* Report mean accuracy from final re-validation with best 5.
* Use Adam with a learn rate decay schdule.
```
NC_FILENAME='ncRNA.gc34.processed.fasta'
PC_FILENAME='pcRNA.gc34.processed.fasta'
DATAPATH=""
try:
from google.colab import drive
IN_COLAB = True
PATH='/content/drive/'
drive.mount(PATH)
DATAPATH=PATH+'My Drive/data/' # must end in "/"
NC_FILENAME = DATAPATH+NC_FILENAME
PC_FILENAME = DATAPATH+PC_FILENAME
except:
IN_COLAB = False
DATAPATH=""
EPOCHS=200
SPLITS=5
K=1
VOCABULARY_SIZE=4**K+1 # e.g. K=3 => 64 DNA K-mers + 'NNN'
EMBED_DIMEN=128
FILENAME='GRU231'
NEURONS=32
DROP=0.5
ACT="tanh"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import RepeatedKFold
from sklearn.model_selection import StratifiedKFold
import tensorflow as tf
from tensorflow import keras
from keras.wrappers.scikit_learn import KerasRegressor
from keras.models import Sequential
from keras.layers import Bidirectional
from keras.layers import GRU
from keras.layers import Dense
from keras.layers import LayerNormalization
import time
dt='float32'
tf.keras.backend.set_floatx(dt)
```
## Build model
```
def compile_model(model):
adam_default_learn_rate = 0.001
schedule = tf.keras.optimizers.schedules.ExponentialDecay(
initial_learning_rate = adam_default_learn_rate*10,
#decay_steps=100000, decay_rate=0.96, staircase=True)
decay_steps=10000, decay_rate=0.99, staircase=True)
# learn rate = initial_learning_rate * decay_rate ^ (step / decay_steps)
alrd = tf.keras.optimizers.Adam(learning_rate=schedule)
bc=tf.keras.losses.BinaryCrossentropy(from_logits=False)
print("COMPILE...")
#model.compile(loss=bc, optimizer=alrd, metrics=["accuracy"])
model.compile(loss=bc, optimizer="adam", metrics=["accuracy"])
print("...COMPILED")
return model
def build_model():
embed_layer = keras.layers.Embedding(
#VOCABULARY_SIZE, EMBED_DIMEN, input_length=1000, input_length=1000, mask_zero=True)
#input_dim=[None,VOCABULARY_SIZE], output_dim=EMBED_DIMEN, mask_zero=True)
input_dim=VOCABULARY_SIZE, output_dim=EMBED_DIMEN, mask_zero=True)
#rnn1_layer = keras.layers.Bidirectional(
# keras.layers.GRU(NEURONS, return_sequences=True,
# input_shape=[1000,EMBED_DIMEN], activation=ACT, dropout=DROP) )#bi
#rnn2_layer = keras.layers.Bidirectional(
# keras.layers.GRU(NEURONS, return_sequences=False,
# activation=ACT, dropout=DROP) )#bi
rnn0_layer = keras.layers.GRU(NEURONS, return_sequences=False,
input_shape=[1000,EMBED_DIMEN], activation=ACT, dropout=DROP)# )#bi
dense1_layer = keras.layers.Dense(NEURONS, activation=ACT,dtype=dt)
drop1_layer = keras.layers.Dropout(DROP)
#dense2_layer = keras.layers.Dense(NEURONS, activation=ACT,dtype=dt)
#drop2_layer = keras.layers.Dropout(DROP)
output_layer = keras.layers.Dense(1, activation="sigmoid", dtype=dt)
mlp = keras.models.Sequential()
mlp.add(embed_layer)
#mlp.add(rnn1_layer)
#mlp.add(rnn2_layer)
mlp.add(rnn0_layer)
mlp.add(dense1_layer)
mlp.add(drop1_layer)
#mlp.add(dense2_layer)
#mlp.add(drop2_layer)
mlp.add(output_layer)
mlpc = compile_model(mlp)
return mlpc
```
## Load and partition sequences
```
# Assume file was preprocessed to contain one line per seq.
# Prefer Pandas dataframe but df does not support append.
# For conversion to tensor, must avoid python lists.
def load_fasta(filename,label):
DEFLINE='>'
labels=[]
seqs=[]
lens=[]
nums=[]
num=0
with open (filename,'r') as infile:
for line in infile:
if line[0]!=DEFLINE:
seq=line.rstrip()
num += 1 # first seqnum is 1
seqlen=len(seq)
nums.append(num)
labels.append(label)
seqs.append(seq)
lens.append(seqlen)
df1=pd.DataFrame(nums,columns=['seqnum'])
df2=pd.DataFrame(labels,columns=['class'])
df3=pd.DataFrame(seqs,columns=['sequence'])
df4=pd.DataFrame(lens,columns=['seqlen'])
df=pd.concat((df1,df2,df3,df4),axis=1)
return df
def separate_X_and_y(data):
y= data[['class']].copy()
X= data.drop(columns=['class','seqnum','seqlen'])
return (X,y)
```
## Make K-mers
```
def make_kmer_table(K):
npad='N'*K
shorter_kmers=['']
for i in range(K):
longer_kmers=[]
for mer in shorter_kmers:
longer_kmers.append(mer+'A')
longer_kmers.append(mer+'C')
longer_kmers.append(mer+'G')
longer_kmers.append(mer+'T')
shorter_kmers = longer_kmers
all_kmers = shorter_kmers
kmer_dict = {}
kmer_dict[npad]=0
value=1
for mer in all_kmers:
kmer_dict[mer]=value
value += 1
return kmer_dict
KMER_TABLE=make_kmer_table(K)
def strings_to_vectors(data,uniform_len):
all_seqs=[]
for seq in data['sequence']:
i=0
seqlen=len(seq)
kmers=[]
while i < seqlen-K+1 -1: # stop at minus one for spaced seed
#kmer=seq[i:i+2]+seq[i+3:i+5] # SPACED SEED 2/1/2 for K=4
kmer=seq[i:i+K]
i += 1
value=KMER_TABLE[kmer]
kmers.append(value)
pad_val=0
while i < uniform_len:
kmers.append(pad_val)
i += 1
all_seqs.append(kmers)
pd2d=pd.DataFrame(all_seqs)
return pd2d # return 2D dataframe, uniform dimensions
def make_kmers(MAXLEN,train_set):
(X_train_all,y_train_all)=separate_X_and_y(train_set)
X_train_kmers=strings_to_vectors(X_train_all,MAXLEN)
# From pandas dataframe to numpy to list to numpy
num_seqs=len(X_train_kmers)
tmp_seqs=[]
for i in range(num_seqs):
kmer_sequence=X_train_kmers.iloc[i]
tmp_seqs.append(kmer_sequence)
X_train_kmers=np.array(tmp_seqs)
tmp_seqs=None
labels=y_train_all.to_numpy()
return (X_train_kmers,labels)
def make_frequencies(Xin):
Xout=[]
VOCABULARY_SIZE= 4**K + 1 # plus one for 'NNN'
for seq in Xin:
freqs =[0] * VOCABULARY_SIZE
total = 0
for kmerval in seq:
freqs[kmerval] += 1
total += 1
for c in range(VOCABULARY_SIZE):
freqs[c] = freqs[c]/total
Xout.append(freqs)
Xnum = np.asarray(Xout)
return (Xnum)
def make_slice(data_set,min_len,max_len):
slice = data_set.query('seqlen <= '+str(max_len)+' & seqlen>= '+str(min_len))
return slice
```
## Cross validation
```
def do_cross_validation(X,y,given_model):
cv_scores = []
fold=0
splitter = ShuffleSplit(n_splits=SPLITS, test_size=0.1, random_state=37863)
for train_index,valid_index in splitter.split(X):
fold += 1
X_train=X[train_index] # use iloc[] for dataframe
y_train=y[train_index]
X_valid=X[valid_index]
y_valid=y[valid_index]
# Avoid continually improving the same model.
model = compile_model(keras.models.clone_model(given_model))
bestname=DATAPATH+FILENAME+".cv."+str(fold)+".best"
mycallbacks = [keras.callbacks.ModelCheckpoint(
filepath=bestname, save_best_only=True,
monitor='val_accuracy', mode='max')]
print("FIT")
start_time=time.time()
history=model.fit(X_train, y_train, # batch_size=10, default=32 works nicely
epochs=EPOCHS, verbose=1, # verbose=1 for ascii art, verbose=0 for none
callbacks=mycallbacks,
validation_data=(X_valid,y_valid) )
end_time=time.time()
elapsed_time=(end_time-start_time)
print("Fold %d, %d epochs, %d sec"%(fold,EPOCHS,elapsed_time))
pd.DataFrame(history.history).plot(figsize=(8,5))
plt.grid(True)
plt.gca().set_ylim(0,1)
plt.show()
best_model=keras.models.load_model(bestname)
scores = best_model.evaluate(X_valid, y_valid, verbose=0)
print("%s: %.2f%%" % (best_model.metrics_names[1], scores[1]*100))
cv_scores.append(scores[1] * 100)
print()
print("%d-way Cross Validation mean %.2f%% (+/- %.2f%%)" % (fold, np.mean(cv_scores), np.std(cv_scores)))
```
## Train on RNA lengths 200-1Kb
```
MINLEN=200
MAXLEN=1000
print("Load data from files.")
nc_seq=load_fasta(NC_FILENAME,0)
pc_seq=load_fasta(PC_FILENAME,1)
train_set=pd.concat((nc_seq,pc_seq),axis=0)
nc_seq=None
pc_seq=None
print("Ready: train_set")
#train_set
subset=make_slice(train_set,MINLEN,MAXLEN)# One array to two: X and y
print ("Data reshape")
(X_train,y_train)=make_kmers(MAXLEN,subset)
#print ("Data prep")
#X_train=make_frequencies(X_train)
print ("Compile the model")
model=build_model()
print ("Summarize the model")
print(model.summary()) # Print this only once
model.save(DATAPATH+FILENAME+'.model')
print ("Cross valiation")
do_cross_validation(X_train,y_train,model)
print ("Done")
```
| github_jupyter |
```
import pandas as pd
import numpy as np
#from plotnine import *
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix
from sklearn.metrics import precision_score, recall_score
from sklearn.metrics import cohen_kappa_score
from sklearn import preprocessing
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import PolynomialFeatures
from sklearn.model_selection import train_test_split
from sklearn.svm import LinearSVC
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.tree import export_graphviz
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import log_loss
from sklearn.pipeline import Pipeline
from sklearn.ensemble import VotingClassifier
import statsmodels.api as sm
import itertools
from scipy.stats import mode
import random
import matplotlib.pyplot as plt
plt.style.use('classic')
%matplotlib inline
import seaborn as sns
sns.set()
import xgboost as xgb
naif = pd.read_excel('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\featuresTrain.xlsx')
eeg = pd.read_excel('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\spectrogram_eeg_features30Train.xlsx')
acc = pd.read_excel('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\acc_featuresTrain.xlsx')
pulse = pd.read_csv('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\pulse_featuresTrain.csv')
#Pulse coliumn 0 is index (pd.write_csv / pd.read_csv) cause overfitting. So removing it.
pulse = pulse.iloc[:, 1:]
pulse.columns
print(eeg[eeg.eeg1_Above100Hz0!=0].shape)
print(eeg[eeg.eeg2_Above100Hz0!=0].shape)
print(eeg[eeg.eeg3_Above100Hz0!=0].shape)
print(eeg[eeg.eeg4_Above100Hz0!=0].shape)
```
Frequency aboive 100Hz are always null, we don't need the columns
```
eeg.drop(columns=["eeg1_Above100Hz0", "eeg2_Above100Hz0", "eeg3_Above100Hz0", "eeg4_Above100Hz0"], inplace=True)
eeg.head()
df = pd.concat([eeg, acc, pulse, naif], axis=1)
df.head()
#training, test = np.split(df.sample(frac=1, random_state=42), [int(.8*len(df))])
training, test = train_test_split(df, test_size=0.2, random_state=42)
X = training.iloc[:,:-1]
y = training.iloc[:,-1]
X_test = test.iloc[:,:-1]
y_true = test.iloc[:,-1]
```
Subdatat set:
- naif
- all but
```
naif.columns[:-1]
Xbaseline = X.drop(columns=naif.columns[:-1], inplace=False)
X_testbaseline = X_test.drop(columns=naif.columns[:-1], inplace=False)
Xbaseline.head()
L= list(eeg.columns) + list(acc.columns) + list(pulse.columns)#eeg.columns + acc.columns + pulse.columns
Xnaif = X.drop(columns=L, inplace=False)
X_testnaif = X_test.drop(columns=L, inplace=False)
Xnaif.head()
L= list(naif.columns[:-1]) + list(acc.columns) + list(pulse.columns)
Xeeg = X.drop(columns=L, inplace=False)
X_testeeg = X_test.drop(columns=L, inplace=False)
Xeeg.head()
L= list(naif.columns[:-1]) + list(acc.columns) + list(eeg.columns)
Xpulse= X.drop(columns=L, inplace=False)
X_testpulse = X_test.drop(columns=L, inplace=False)
Xpulse.head()
L= list(naif.columns[:-1]) + list(pulse.columns) + list(eeg.columns)
Xacc= X.drop(columns=L, inplace=False)
X_testacc = X_test.drop(columns=L, inplace=False)
Xacc.head()
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# %load "C:\\Users\\i053131\Desktop\\Epilepsie\\Dreem\\src\\utils\\error.py"
def AnalyzeError(y_true, y_pred):
fig, ax = plt.subplots(figsize=(20,10))
plt.subplot(1,2, 1)
sns.countplot(x=0, data=pd.DataFrame(y_true))
plt.ylim(0, 4000)
plt.subplot(1,2, 2)
sns.countplot(x=0, data=pd.DataFrame(y_pred))
plt.ylim(0, 4000)
fig.suptitle("Actual and predicted distribution", size = 'x-large')
plt.show()
df_ = pd.DataFrame()
df_["Test"]= y_true
df_["Pred"] = y_pred
df_['error'] = df_.Test != df_.Pred
#sns.countplot(x="Test", data=df_[df_.error])
error0 = df_[(df_.error) & (df_.Test==0)].count()[0] / df_[df_.Test==0].count()[0]
error1 = df_[(df_.error) & (df_.Test==1)].count()[0] / df_[df_.Test==1].count()[0]
error2 = df_[(df_.error) & (df_.Test==2)].count()[0] / df_[df_.Test==2].count()[0]
error3 = df_[(df_.error) & (df_.Test==3)].count()[0] / df_[df_.Test==3].count()[0]
error4 = df_[(df_.error) & (df_.Test==4)].count()[0] / df_[df_.Test==4].count()[0]
Lerror = [error0, error1, error2, error3, error4]
sns.barplot(x=[0, 1, 2, 3, 4], y=Lerror)
plt.title('Wrongly classified in a phase in percent of the test population for this phase')
plt.show()
```
## for comparaison only on naif features
Only naive features with gradient boosting
n_estimators = 100
kappa: 0.683
accurancy: 0.777
{'learning_rate': 0.1, 'max_depth': 15, 'min_samples_leaf': 1, 'min_samples_split': 2, 'subsample': 0.8}
- kappa: 0.6790102154750923
- accurancy for n_estimators = 0.7741273100616016
{'learning_rate': 0.1, 'max_depth': 10, 'min_samples_leaf': 1, 'min_samples_split': 2, 'subsample': 0.8}
- kappa: 0.6832914749130093
- accurancy for n_estimators = 0.7767510837326033
"Seleccted)
i=110
gbc = GradientBoostingClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 10,
min_samples_leaf= 1, min_samples_split= 2, subsample= 0.8)
- for n_estimators= 110
- log loss = 0.5953633123652049
- kappa = 0.685
- accuracy = 0.778
```
gbc = GradientBoostingClassifier(n_estimators = 100, random_state=42)
r1 = [1]
r2 = [2]
parametres = {'max_depth': [15, 20] ,'learning_rate': [0.01, 0.1], "min_samples_leaf" : r1,
"min_samples_split" : r2, 'subsample': [0.8, 1.0]}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=gbc, param_grid=parametres, scoring=ck_score, n_jobs=-1, verbose=2)
grid_fitted = grid.fit(Xnaif,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_testnaif)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy for n_estimators = " , accuracy_score(y_true, y_pred))
gbc = GradientBoostingClassifier(n_estimators = 100, random_state=42)
r1 = [1]
r2 = [2]
parametres = {'max_depth': [10, 15] ,'learning_rate': [0.1], "min_samples_leaf" : r1,
"min_samples_split" : r2, 'subsample': [0.6, 0.8]}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=gbc, param_grid=parametres, scoring=ck_score, n_jobs=-1, verbose=2)
grid_fitted = grid.fit(Xnaif,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_testnaif)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy for n_estimators = " , accuracy_score(y_true, y_pred))
r = range(10, 200 , 10)
errors = []
Lk = []
La = []
for i in r:
gbc = GradientBoostingClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 10,
min_samples_leaf= 1, min_samples_split= 2, subsample= 0.8)
gbc.fit(Xnaif, y)
ll = log_loss(y_true, gbc.predict_proba(X_testnaif))
errors.append(ll)
y_pred = gbc.predict(X_testnaif)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators= ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
plt.plot(r, errors, label = "log loss")
plt.plot(r, La, label = "accuracy")
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
```
go for 110
- for n_estimators= 110
- log loss = 0.5953633123652049
- kappa = 0.685
- accuracy = 0.778
```
errors = []
Lk = []
La = []
i=110
gbc = GradientBoostingClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 10,
min_samples_leaf= 1, min_samples_split= 2, subsample= 0.8)
gbc.fit(Xnaif, y)
ll = log_loss(y_true, gbc.predict_proba(X_testnaif))
y_pred = gbc.predict(X_testnaif)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators= ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
```
confusion matrix has to be (re) computed on best hyparameters
```
cnf_matrix = confusion_matrix(y_true, y_pred)
np.set_printoptions(precision=2)
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], title='Confusion matrix, without normalization')
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], normalize=True,
title='Normalized confusion matrix')
plt.show()
importances = gbc.feature_importances_
feature_importances = pd.DataFrame(importances, index = Xnaif.columns,
columns=['importance']).sort_values('importance', ascending=False)
plt.bar(feature_importances.index, feature_importances["importance"])
plt.show()
feature_importances.head(50)
```
## Features when naive are dropped
- original with "index features" was
{'learning_rate': 0.1, 'max_depth': 15, 'min_samples_leaf': 1, 'min_samples_split': 2, 'subsample': 0.8}
kappa: 0.747
accurancy : 0.821
- when index features is dropped, this drop to
{'learning_rate': 0.1, 'max_depth': 15, 'min_samples_leaf': 1, 'min_samples_split': 2, 'subsample': 0.8}
kappa: 0.697
accurancy for n_estimators = 0.784
- when correcting the bug on pulse data generation (now ir captor is relevant)
{'learning_rate': 0.1, 'max_depth': 15, 'min_samples_leaf': 1, 'min_samples_split': 2, 'subsample': 0.8}
kappa: 0.720
accurancy for n_estimators = 0.800
- when correcting the second bug on pulse (both ir and r captors are relevant
for n_estimators= 50 (being more conservative on parcimonie). performance are comparable
kappa = 0.712
accuracy = 0.795
- as a comparaison, only on naive
Only naive features
kappa: 0.683
accurancy for n_estimators = 0.777
```
gbc = GradientBoostingClassifier(n_estimators = 30, random_state=42)
r1 = [1]
r2 = [2]
parametres = {'max_depth': [5, 10, 15] ,'learning_rate': [0.1], "min_samples_leaf" : r1,
"min_samples_split" : r2, 'subsample': [0.6, 0.8]}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=gbc, param_grid=parametres, scoring=ck_score, n_jobs=-1, verbose=2)
grid_fitted = grid.fit(Xbaseline,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_testbaseline)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy for n_estimators = " , accuracy_score(y_true, y_pred))
errors = []
Lk = []
La = []
r = range(10, 100 , 10)
for i in r:
gbc = GradientBoostingClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 15,
min_samples_leaf= 1, min_samples_split= 2, subsample= 0.8)
gbc.fit(Xbaseline, y)
ll = log_loss(y_true, gbc.predict_proba(X_testbaseline))
errors.append(ll)
y_pred = gbc.predict(X_testbaseline)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators= ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
plt.plot(r, errors, label = "log loss")
plt.plot(r, La, label = "accuracy")
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
```
Go for 50
```
errors = []
Lk = []
La = []
i=50
gbc = GradientBoostingClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 15,
min_samples_leaf= 1, min_samples_split= 2, subsample= 0.8)
gbc.fit(Xbaseline, y)
ll = log_loss(y_true, gbc.predict_proba(X_testbaseline))
y_pred = gbc.predict(X_testbaseline)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators= ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
cnf_matrix = confusion_matrix(y_true, y_pred)
np.set_printoptions(precision=2)
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], title='Confusion matrix, without normalization')
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], normalize=True,
title='Normalized confusion matrix')
plt.show()
importances = gbc.feature_importances_
feature_importances = pd.DataFrame(importances, index = Xbaseline.columns,
columns=['importance']).sort_values('importance', ascending=False)
plt.bar(feature_importances.index, feature_importances["importance"])
plt.show()
feature_importances.head(50)
```
### with all features (naive + baseline),
starting with Random forest
```
rnd_clf = RandomForestClassifier(max_depth=12, max_features= None, random_state=42, n_estimators=100, n_jobs=-2)
rnd_clf.fit(X, y)
y_pred_rf = rnd_clf.predict(X_test)
kappa_rf = cohen_kappa_score(y_true, y_pred_rf)
print("kappa: ", kappa_rf)
print("accurancy: ", accuracy_score(y_true, y_pred_rf))
```
with GridSearchCV
parametres = {"max_depth": [7, 8, 9, 10, 11, 12], "min_samples_leaf" : r1, "class_weight": [None, "balanced"],
"min_samples_split" : r2, "n_estimators" :[100]}
- original
{'class_weight': None, 'max_depth': 12, 'min_samples_leaf': 1, 'min_samples_split': 2, 'n_estimators': 100}
kappa: 0.707
accurancy: 0.792
- dropping "index feature" this decreased to
{'class_weight': None, 'max_depth': 12, 'min_samples_leaf': 1, 'min_samples_split': 2, 'n_estimators': 100}
kappa: 0.6959086083496023
accurancy: 0.783937942048825
- first pulse feature correction
{'class_weight': None, 'max_depth': 12, 'min_samples_leaf': 1, 'min_samples_split': 2, 'n_estimators': 100}
kappa: 0.6988175987857627
accurancy: 0.7859913301391741
```
rfc = RandomForestClassifier(random_state=42, max_features= None)
#RandomForestClassifier(max_depth=9, min_samples_leaf=5, random_state=42, min_samples_split=64, n_estimators=n
r1 = [1, 10]
r2 = [2, 10]
parametres = {"max_depth": [10, 12, 15], "min_samples_leaf" : r1, "class_weight": [None, "balanced"],
"min_samples_split" : r2, "n_estimators" :[100]}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=rfc,param_grid=parametres,scoring=ck_score, n_jobs=-1, verbose = 2)
grid_fitted = grid.fit(X,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_test)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy: ", accuracy_score(y_true, y_pred))
```
{'class_weight': 'balanced', 'max_depth': 20, 'min_samples_leaf': 1, 'min_samples_split': 10, 'n_estimators': 100}
kappa: 0.725
accurancy: 0.802
with index feature dropped
{'class_weight': 'balanced', 'max_depth': 20, 'min_samples_leaf': 1, 'min_samples_split': 10, 'n_estimators': 100}
kappa: 0.720
accurancy: 0.799
{'class_weight': 'balanced', 'max_depth': 15, 'min_samples_leaf': 1, 'min_samples_split': 2, 'n_estimators': 100}
{'class_weight': 'balanced', 'max_depth': 15, 'min_samples_leaf': 1, 'min_samples_split': 2, 'n_estimators': 100}
kappa: 0.7146082490800214
accurancy: 0.7942048825005704
```
print("accurancy: ", accuracy_score(y_true, y_pred))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_true, y_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], title='Confusion matrix, without normalization')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], normalize=True,
title='Normalized confusion matrix')
plt.show()
```
Random forrest with
{'class_weight': None, 'max_depth': 30, 'min_samples_leaf': 1, 'min_samples_split': 2, 'n_estimators': 100}
kappa: 0.733
accurancy: 0.810
Class_weigth = None correcrly predicted 1 drop from 22% to 11%
improvement in kappa and accuracy probably does not work the extra tree depth
After discarding index 'features'
{'class_weight': 'balanced', 'max_depth': 30, 'min_samples_leaf': 1, 'min_samples_split': 10, 'n_estimators': 100}
kappa: 0.723
accurancy: 0.802
```
rfc = RandomForestClassifier(random_state=42, max_features= None)
r1 = [1, 10]
r2 = [2, 10]
parametres = {"max_depth": [20, 25, 30], "min_samples_leaf" : r1, "class_weight": [None, "balanced"],
"min_samples_split" : r2, "n_estimators" :[100]}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=rfc,param_grid=parametres,scoring=ck_score, n_jobs=-1)
grid_fitted = grid.fit(X,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_test)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy: ", accuracy_score(y_true, y_pred))
cnf_matrix = confusion_matrix(y_true, y_pred)
np.set_printoptions(precision=2)
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], title='Confusion matrix, without normalization')
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], normalize=True,
title='Normalized confusion matrix')
plt.show()
```
Trying best parameters with different growing values for n_estimators
{'class_weight': None, 'max_depth': 30, 'min_samples_leaf': 1, 'min_samples_split': 2, 'n_estimators': 100}
from gridcv with n_estimators=100
kappa: 0.733
accurancy: 0.809719370294319
for n_estimators= 800
kappa = 0.738
accuracy = 0.813
- After discaeding index "features"
for n_estimators= 1400
log loss = 0.5523685672967809
kappa = 0.7233808409352107
accuracy = 0.8028747433264887
- after first correction of pulse features
for n_estimators = 1500
kappa = 0.730
accuracy = 0.810
- after second correction
kappa = 0.730
accuracy = 0.805
Analysis of with n-estimators = 1500
(with pulse bugs corrected)
rnd_clf = RandomForestClassifier(n_estimators=1500, n_jobs=-1, max_depth=30, class_weight= "balanced", min_samples_leaf= 1,
min_samples_split= 10, random_state=42, max_features= None)
- kappa = 0.730
- accuracy = 0.81
```
errors = []
Lk = []
La = []
r = range(200, 1600 , 100)
rnd_clf = RandomForestClassifier(n_estimators=100, n_jobs=-1, max_depth=30, class_weight= None, min_samples_leaf= 1,
min_samples_split= 10, warm_start=True, random_state=42, max_features= None)
for i in r:
rnd_clf.fit(X, y)
ll = log_loss(y_true, rnd_clf.predict_proba(X_test))
errors.append(ll)
y_pred = rnd_clf.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators= ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
rnd_clf.fit(X, y).n_estimators += 100
plt.plot(r, errors, label = "log loss")
plt.plot(r, La, label = "accuracy")
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
```
Analysis of with n-estimators = 1500
(with pulse bugs corrected)
rnd_clf = RandomForestClassifier(n_estimators=1500, n_jobs=-1, max_depth=30, class_weight= "balanced", min_samples_leaf= 1,
min_samples_split= 10, random_state=42, max_features= None)
- kappa = 0.730
- accuracy = 0.81
```
rnd_clf = RandomForestClassifier(n_estimators=1500, n_jobs=-1, max_depth=30, class_weight= "balanced", min_samples_leaf= 1,
min_samples_split= 10, random_state=42, max_features= None)
rnd_clf.fit(X, y)
y_pred = rnd_clf.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators= 1500")
print("kappa = ", k)
print("accuracy = ", a)
cnf_matrix = confusion_matrix(y_true, y_pred)
np.set_printoptions(precision=2)
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], title='Confusion matrix, without normalization')
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], normalize=True,
title='Normalized confusion matrix')
plt.show()
importances = rnd_clf.feature_importances_
feature_importances = pd.DataFrame(importances, index = X.columns,
columns=['importance']).sort_values('importance', ascending=False)
plt.bar(feature_importances.index, feature_importances["importance"])
plt.show()
feature_importances.head(50)
```
### Boosting
for n_estimators = 70
log loss = 0.48672557810125716
kappa = 0.7634687814405141
accuracy = 0.8316221765913757
```
gbc = GradientBoostingClassifier(n_estimators=100, learning_rate=0.1, max_depth=15, random_state=42)
gbc.fit(X, y)
y_pred = gbc.predict(X_test)
kappa = cohen_kappa_score(y_true, y_pred)
print("kappa for n_estimators = " , kappa)
print("accurancy for n_estimators = " , accuracy_score(y_true, y_pred))
```
{'learning_rate': 0.1, 'max_depth': 15, 'subsample': 0.8}
- kappa: 0.785
- accurancy = 0.847
after dropping index
- kappa: 0.766
- accurancy for n_estimators = 0.834
after fixing pulse bug with n estimators = 30
{'learning_rate': 0.1, 'max_depth': 13, 'min_samples_leaf': 1, 'min_samples_split': 2, 'subsample': 0.7}
- kappa: 0.744
- accurancy for n_estimators = 0.82
as a comparaison
Features when naive are dropped
{'learning_rate': 0.1, 'max_depth': 15, 'min_samples_leaf': 1, 'min_samples_split': 2, 'subsample': 0.8}
kappa: 0.747
accurancy : 0.821
Only naive features
kappa: 0.683
accurancy for n_estimators = 0.777
```
gbc = GradientBoostingClassifier(n_estimators = 30, random_state=42)
r1 = [1]
r2 = [2]
parametres = {'max_depth': [13, 15] ,'learning_rate': [0.1], "min_samples_leaf" : r1,
"min_samples_split" : r2, 'subsample': [0.7, 0.8]}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=gbc, param_grid=parametres, scoring=ck_score, n_jobs=-1, verbose=2)
grid_fitted = grid.fit(X,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_test)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy for n_estimators = " , accuracy_score(y_true, y_pred))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_true, y_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], title='Confusion matrix, without normalization')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], normalize=True,
title='Normalized confusion matrix')
plt.show()
```
trying with parameter from gridCV
{'learning_rate': 0.1, 'max_depth': 13, 'min_samples_leaf': 1, 'min_samples_split': 2, 'subsample': 0.7}
kappa: 0.7433764686380122
accurancy for n_estimators = 0.8174766141911932
Overfitting after 100 n_estimators. (from 200 on)
for 100:
kappa = 0.786
accuracy = 0.847
as a comparaison
Features when naive are dropped
{'learning_rate': 0.1, 'max_depth': 15, 'min_samples_leaf': 1, 'min_samples_split': 2, 'subsample': 0.8}
kappa: 0.747
accurancy : 0.821
Only naive features
kappa: 0.683
accurancy for n_estimators = 0.777
```
errors = []
Lk = []
La = []
r = range(10, 100 , 10)
for i in r:
gbc = GradientBoostingClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 13,
min_samples_leaf= 1, min_samples_split= 2, subsample= 0.7)
gbc.fit(X, y)
ll = log_loss(y_true, gbc.predict_proba(X_test))
errors.append(ll)
y_pred = gbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators = ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
plt.plot(r, errors, label = "log loss")
plt.plot(r, La, label = "accuracy")
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
```
go for 50
- log loss = 0.4857512636168815
- kappa = 0.7586408515510799
- accuracy = 0.8281998631074606
```
errors = []
Lk = []
La = []
i=50
gbc = GradientBoostingClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 13,
min_samples_leaf= 1, min_samples_split= 2, subsample= 0.7)
gbc.fit(X, y)
ll = log_loss(y_true, gbc.predict_proba(X_test))
y_pred = gbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators= ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
cnf_matrix = confusion_matrix(y_true, y_pred)
np.set_printoptions(precision=2)
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], title='Confusion matrix, without normalization')
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], normalize=True,
title='Normalized confusion matrix')
plt.show()
importances = gbc.feature_importances_
feature_importances = pd.DataFrame(importances, index = X.columns,
columns=['importance']).sort_values('importance', ascending=False)
plt.bar(feature_importances.index, feature_importances["importance"])
plt.show()
feature_importances.head(50)
```
## Xgboost
Performance à battre (regular gradien boosting)
- kappa = 0.7586408515510799
- accuracy = 0.8281998631074606
nouveau record a battre for n_estimators = 115
- log loss = 0.4354704374970457
- kappa = 0.7797652198456537
- accuracy = 0.8428017339721652
```
model = xgb.XGBClassifier()
model.fit(X, y)
#y_pred = gbc.predict(X_test)
#k=cohen_kappa_score(y_true, y_pred)
#a= accuracy_score(y_true, y_pred)
y_pred = model.predict(X_test)
accuracy_score(y_true, y_pred)
cohen_kappa_score(y_true, y_pred)
xbc = xgb.XGBClassifier(n_estimators = 30, random_state=42)
parametres = {'max_depth': [10, 13, 15] ,'learning_rate': [0.1], 'subsample': [0.7, 0.8, 1]}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=xbc, param_grid=parametres, scoring=ck_score, n_jobs=-1, verbose=2)
grid_fitted = grid.fit(X,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_test)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy for n_estimators = " , accuracy_score(y_true, y_pred))
errors = []
Lk = []
La = []
r = range(10, 100 , 10)
for i in r:
xbc = xgb.XGBClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 13, subsample= 0.7,
n_jobs=-2)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators = ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
plt.plot(r, errors, label = "log loss")
plt.plot(r, La, label = "accuracy")
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
errors = []
Lk = []
La = []
r = range(100, 200, 20)
for i in r:
xbc = xgb.XGBClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 13, subsample= 0.7,
n_jobs=-2)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators = ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
plt.plot(r, errors, label = "log loss")
plt.plot(r, La, label = "accuracy")
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
xbc = xgb.XGBClassifier(n_estimators = 120, random_state=42)
parametres = {'max_depth': [12, 13, 14] ,'learning_rate': [0.1], 'subsample': [0.7]}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=xbc, param_grid=parametres, scoring=ck_score, n_jobs=-1, verbose=2)
grid_fitted = grid.fit(X,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_test)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
errors = []
Lk = []
La = []
r = range(110, 140, 5)
for i in r:
xbc = xgb.XGBClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 13, subsample= 0.7,
n_jobs=-2)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators = ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
plt.plot(r, errors, label = "log loss")
plt.plot(r, La, label = "accuracy")
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
```
Go for 115
```
i = 115
xbc = xgb.XGBClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 13, subsample= 0.7,
n_jobs=-2)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators = ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
cnf_matrix = confusion_matrix(y_true, y_pred)
np.set_printoptions(precision=2)
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], title='Confusion matrix, without normalization')
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=[0, 1, 2, 3, 4], normalize=True,
title='Normalized confusion matrix')
plt.show()
importances = xbc.feature_importances_
feature_importances = pd.DataFrame(importances, index = X.columns,
columns=['importance']).sort_values('importance', ascending=False)
plt.bar(feature_importances.index, feature_importances["importance"])
plt.show()
feature_importances.head(50)
```
### SVM (Support Vector Machine classifier)
Linear kernel {'C': 1, 'loss': 'hinge'}
- kappa: 0.2702593710539586
- accurancy: 0.4498060689025781
Polynomial kernel {'C': 10, 'coef0': 1, 'degree': 4}
- kappa: 0.40
- accurancy: 0.60
RBF kernel 'C': 100
- kappa: 0.46
- accurancy: 0.63
```
scaler = StandardScaler()
scaler.fit(X)
X_scaled = pd.DataFrame(scaler.transform(X), columns = X.columns)
X_test_scaled = pd.DataFrame(scaler.transform(X_test), columns = X_test.columns)
lsvc = LinearSVC(penalty='l2', random_state=42)
parametres = {'C': [0.01, 0.1, 1, 10], 'loss': ['hinge', 'squared_hinge']}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=lsvc, param_grid=parametres, scoring=ck_score, n_jobs=-1, verbose=2)
grid_fitted = grid.fit(X_scaled,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_test_scaled)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy: " , accuracy_score(y_true, y_pred))
poly_svc = SVC(kernel="poly", degree=2, random_state=42)
parametres = {'C': [0.01, 0.1, 1, 10], 'coef0': [0, 1], 'degree' : [2, 3, 4]}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=poly_svc, param_grid=parametres, scoring=ck_score, n_jobs=-1, verbose=2)
grid_fitted = grid.fit(X_scaled,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_test_scaled)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy: " , accuracy_score(y_true, y_pred))
#for degree = 2
#{'C': 10, 'coef0': 1}
#kappa: 0.31617419927367196
#accurancy: 0.543235227013461
rbf_svc = SVC(kernel="rbf", random_state=42)
parametres = {'C': [0.01, 0.1, 1, 10, 100]}
ck_score = make_scorer(cohen_kappa_score)
grid = GridSearchCV(estimator=rbf_svc, param_grid=parametres, scoring=ck_score, n_jobs=-1, verbose=2)
grid_fitted = grid.fit(X_scaled,y)
print(grid_fitted.best_params_)
y_pred = grid.predict(X_test_scaled)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy: " , accuracy_score(y_true, y_pred))
errors = []
Lk = []
La = []
L = [0.1, 1, 10, 100]
for i in L:
rbfc = SVC(kernel="rbf", random_state=42, probability=True, C=i)
rbfc.fit(X_scaled, y)
ll = log_loss(y_true, rbfc.predict_proba(X_test_scaled))
errors.append(ll)
y_pred = rbfc.predict(X_test_scaled)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("for n_estimators = ", i)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
plt.plot(r, errors, label = "log loss")
plt.plot(r, La, label = "accuracy")
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
scaler = StandardScaler()
svc_rbf = SVC(kernel="rbf", random_state=42, C=100, probability=True)
clf_svm = Pipeline([('std_scaler', scaler), ('svc', svc_rbf)])
clf_svm.fit(X, y)
y_pred = clf_svm.predict(X_test)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy: " , accuracy_score(y_true, y_pred))
```
### Putting everything together to vote
```
clf_svm = Pipeline([('std_scaler', scaler), ('svc', svc_rbf)])
rnd_clf = RandomForestClassifier(n_estimators=1500, n_jobs=-2, max_depth=30, class_weight= "balanced", min_samples_leaf= 1,
min_samples_split= 10, random_state=42, max_features= None)
xbc = xgb.XGBClassifier(n_estimators = 115, random_state=42, learning_rate= 0.1, max_depth= 13, subsample= 0.7,
n_jobs=-2)
log_clf = LogisticRegression()
rnd_clf = RandomForestClassifier()
svm_clf = SVC()
voting_clf = VotingClassifier( estimators=[('lr', clf_svm), ('rf', rnd_clf), ('svc', xbc)], voting='soft' )
voting_clf.fit(X, y)
y_pred = voting_clf.predict(X_test)
print("kappa: ", cohen_kappa_score(y_true, y_pred))
print("accurancy: " , accuracy_score(y_true, y_pred))
```
| github_jupyter |
# Extras
If we have time: Changing the colormap with widgets in two ways -- The Michigan Depth Map
Order is:
1. Review of Michigan map
1. More complicated layout
1. Even more complicated layout and placement (perhaps unnecessarily so :D )
Import our usual things:
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
```
Also import ipywidgets:
```
import ipywidgets
```
## 1. Review of Michigan map: Michigan colormap and scale with interact
More info for `Output`: https://ipywidgets.readthedocs.io/en/latest/examples/Output%20Widget.html
**Note:** If we are short on time, we might only get through the color-map-by-hand portion of this.
We'll need a few extra functions to do this sort of thing "by hand":
Last week we also started working with the Michigan Depth Map which we loaded with Numpy and performed some data cleaning on:
```
data_filename = '/Users/jillnaiman/Downloads/michigan_lld.flt'
michigan = np.fromfile(data_filename, dtype='f4').reshape((5365, 4201))
michigan[michigan == -9999] = np.nan # set flagged bad data to NaN
# quick plot of depths:
plt.hist(michigan.flat)
plt.show()
```
Neat! Let's look at this data more in the way that it was intended -- as an image. We can use `matplotlib`'s `imshow` function to do this:
```
plt.imshow(michigan)
plt.colorbar()
plt.show()
```
### Question:
Now that we've had a chance to look at our data a bit, what do we think the values represent? What does a positive value mean? Negative value? Where do we think, spatially, these things will lie?
```
plt.imshow(michigan)
plt.clim(0, 100) # only plot from depths of 0->100
plt.colorbar(extend = 'both') # add little arrow ends
plt.show()
```
Let's see if we can't get a colormap that shows this outline better. Turns out there is an actual "terrain" map:
```
plt.imshow(michigan, cmap="terrain")
plt.colorbar()
plt.show()
```
So, while this is starting to look better, intutatively, we want our map to look bluish in the lake, and brownish on the land. We can do this by doing a symmetric log color normalization:
```
import matplotlib.colors as colors
plt.imshow(michigan, cmap="terrain", norm = colors.SymLogNorm(10))
plt.colorbar()
plt.show()
```
We can even set the color limits to be symmetric so that the yellow bit is right at zero elevation:
```
np.nanmin(michigan), np.nanmax(michigan)
```
So, we'll make sure we make our colormap to include these limits:
```
plt.imshow(michigan, cmap="terrain", norm = colors.SymLogNorm(10))
plt.clim(-352, 352)
plt.colorbar()
plt.show()
```
If we now look at our image, we see some interesting things. So, now there is a sharp contrast between negative & positive depths/heights and there is not as much contrast between blue/green or brown/white.
But why? Let's check out the docs for `SymLogNorm`:
```
colors.SymLogNorm?
```
This is a symmetrical log scale so it logs things both in the negative & positive directions.
Example:
```
np.log10([1,10,50]),np.log10(np.abs([-1,-10,-50]))
```
We see that 1 and 10 are mapped to a jump of 1 but 1->50 is mapped only to a jump of 0.7 instead of 40.
The lake Michigan data is a very high resolution map, so we can zoom in to see some cool details:
```
plt.imshow(michigan, cmap="terrain", norm = colors.SymLogNorm(10))
plt.clim(-352, 352)
plt.colorbar()
plt.xlim(2700, 3300)
plt.ylim(3300, 3900)
```
This shows us one of the rivers that feed into lake Michigan.
And just for fun, here is how it looks with our bad "jet" colormap:
```
plt.imshow(michigan, cmap="jet", norm = colors.SymLogNorm(10))
plt.clim(-352, 352)
plt.colorbar()
# ew.
```
One natural thing we might want to do is change color scheme and be able to toggle on and off the SymLogNorm color remapper. We can do this 2 ways - by using our widget `@interact` decorator function again, and by explicitly laying out widgets. Let's try to first way first:
```
@ipywidgets.interact(colormap = plt.colormaps(), color_range = (1.0, 352.0, 1.0),
sym_log=True)
def plot(colormap = 'terrain', color_range = 352, sym_log = True):
if sym_log:
norm = colors.SymLogNorm(10)
else:
norm = colors.Normalize()
fig, ax = plt.subplots(figsize=(6,8))
# calling colorbar in a different way:
CAX = ax.imshow(michigan, cmap=colormap, norm = norm)
CAX.set_clim(-color_range, color_range)
plt.colorbar(CAX, extend = 'both')
plt.show()
```
## 2. More complicated layout
We can mess with the layout of our widgets by creating them externally, and then using them to plot.
Let's start with creating a dropdown widget for all of the colormaps:
```
cmap_widget = ipywidgets.Dropdown(options=plt.colormaps())
```
Let's take a quick look:
```
cmap_widget
```
Ok! So we just have the stand-alone widget. Since we know that some of the color maps work well/less well for this dataset, let's set a default of the "terrain" colormap to this widget:
```
cmap_widget = ipywidgets.Dropdown(options=plt.colormaps(), value='terrain')
cmap_widget
```
Finally, let's ad a description to this widget that is different from the default in the `@interacts` call above.
```
cmap_widget = ipywidgets.Dropdown(options=plt.colormaps(), value='terrain', description='Select colormap:')
cmap_widget
```
We note that now our description sort of "runs off" the page. Because we have access to the individual widget, we can mess with the "layout" of this widget -- i.e., how it looks.
```
cmap_widget.keys
cmap_widget.layout.width = '500px' # changes the box size
cmap_widget.style.keys # here is where the description width is hidden
cmap_widget.style.description_width = 'initial'
cmap_widget
cmap_widget.layout.width = '200px' # back to not so large of a box
cmap_widget
```
Let's now make our checkbox button widget:
```
log_check = ipywidgets.Checkbox(value=True, description='Take log of colormap? ')
log_check
```
Note that we could also use a toggle button (see: https://ipywidgets.readthedocs.io/en/latest/examples/Widget%20List.html#ToggleButton).
We can use these as inputs to an `@interacts` call:
```
@ipywidgets.interact(colormap = cmap_widget, sym_log = log_check)
def plot(colormap = 'terrain', color_range = 352, sym_log = True): # hard-coding color_range here
if sym_log:
norm = colors.SymLogNorm(10)
else:
norm = colors.Normalize()
fig, ax = plt.subplots(figsize=(6,8))
# calling colorbar in a different way:
CAX = ax.imshow(michigan, cmap=colormap, norm = norm)
CAX.set_clim(-color_range, color_range)
plt.colorbar(CAX, extend = 'both')
plt.show()
```
So, now we've messed with how our widgets look, but how about where they are placed?
One option is the "Even more complicated layout and placement" section below, OR what we will cover with `bqplot` next week (for example, see `bqplot` examples in https://ipywidgets.readthedocs.io/en/latest/examples/Layout%20Templates.html#2x2-Grid)
## 3. Even more complicated layout and placement
```
#plt.close('all')
from IPython.display import display, clear_output
%config InlineBackend.close_figures=False
# If you get "double" displays over the next 2 cells, make sure you have this "config" statement there
# This stops the auto-creation of figures until we say "plt.show" or "display"
# Read more here: https://github.com/jupyter-widgets/ipywidgets/issues/1940
fig = plt.figure()
# add axes by hand like last week
# order here is: left, bottom, width, height
plt.ioff()
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
image = ax.imshow(michigan, cmap='terrain')
fig.colorbar(image, extend = 'both')
out = ipywidgets.Output() ### NEW WIDGET CALL
display(out)
```
The `Output` widget sort of "captures" the display until we explictly call it in context:
```
with out:
display(fig)
```
We use the `Layout` widget call, along with `figsize` in matplotlib to change the size of our image:
```
fig = plt.figure(figsize=(8,8))
# add axes by hand like last week
# order here is: left, bottom, width, height
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
image = ax.imshow(michigan, cmap='terrain')
fig.colorbar(image, extend = 'both')
#del out
out = ipywidgets.Output(layout=ipywidgets.Layout(height='500px', width = '500px')) ### NEW WIDGET CALL
display(out) # hold our output...
with out: # until we explicitly say display!
display(fig) # only display the fig when we explitly say to!
```
Why would we bother making our lives more complicated like this instead of just using `@interact` like we did before? So that we can start placing our widgets where we want and start to have a little more control over what we are displaying and how. For example, let's add a dropdown menu by hand:
```
dropdown = ipywidgets.Dropdown(options=plt.colormaps())
dropdown.keys
```
`dropdown.index` gives us the # of the color map from `plt.colormaps()`. Let's add this in:
```
fig = plt.figure(figsize=(8,8))
# add axes by hand like last week
# order here is: left, bottom, width, height
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
image = ax.imshow(michigan, cmap='terrain')
fig.colorbar(image, extend = 'both')
#del out
out = ipywidgets.Output(layout=ipywidgets.Layout(height='500px', width = '500px'))
dropdown = ipywidgets.Dropdown(options=plt.colormaps())
hbox=ipywidgets.HBox([out, dropdown])
display(hbox)
with out:
display(fig)
```
So now we can start placing our interactive widgets how we want! Note that if update the dropdown, nothing happens because its not connected to the plot anyway. Let's work on connecting our dropdown menu to our plot using an `.observe` traitlets call:
```
fig = plt.figure(figsize=(8,8))
# add axes by hand like last week
# order here is: left, bottom, width, height
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
image = ax.imshow(michigan, cmap='terrain')
fig.colorbar(image, extend = 'both')
#del out
out = ipywidgets.Output(layout=ipywidgets.Layout(height='500px', width = '500px'))
dropdown = ipywidgets.Dropdown(options=plt.colormaps())
hbox=ipywidgets.HBox([out, dropdown])
display(hbox)
#with out:
# display(fig)
def updateDropdown(change):
print(change) # first just print
with out:
clear_output(wait=True) # clear everything on the display - don't keep making figures!
display(fig)
dropdown.observe(updateDropdown)
```
Let's use `change['owner'].index` to grab the index of the colormap we want:
```
fig = plt.figure(figsize=(8,8))
# add axes by hand like last week
# order here is: left, bottom, width, height
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
image = ax.imshow(michigan, cmap='terrain')
fig.colorbar(image, extend = 'both')
out = ipywidgets.Output(layout=ipywidgets.Layout(height='500px', width = '500px'))
dropdown = ipywidgets.Dropdown(options=plt.colormaps())
hbox=ipywidgets.HBox([out, dropdown])
display(hbox)
def updateDropdown(change):
cmap=plt.colormaps()[change['owner'].index] # grab our new cmap
# let's start by clearing out all our previous axes and starting with a fresh canvas
for a in fig.axes:
fig.delaxes(a)
# draw on our axes like before
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
# make an image and assign a color map
image = ax.imshow(michigan, cmap=cmap)
fig.colorbar(image, extend = 'both')
# display with an output widget
with out:
clear_output(wait=True) # clear everything on the display - don't keep making figures!
display(fig)
dropdown.observe(updateDropdown)
```
So, it's a little annoying that we have to wait to select something to display, so let's reorganize our function a bit to make it look nice:
```
#plt.close('all') # if you get a "too many figures open" warning
fig = plt.figure(figsize=(8,8))
# add axes by hand like last week
# order here is: left, bottom, width, height
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
image = ax.imshow(michigan, cmap='terrain')
fig.colorbar(image, extend = 'both')
out = ipywidgets.Output(layout=ipywidgets.Layout(height='500px', width = '500px'))
dropdown = ipywidgets.Dropdown(options=plt.colormaps())
hbox=ipywidgets.HBox([out, dropdown])
display(hbox)
def updateDropdown(change):
if change is not None:
cmap=plt.colormaps()[change['owner'].index]
for a in fig.axes:
fig.delaxes(a)
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
image = ax.imshow(michigan, cmap=cmap)
fig.colorbar(image, extend = 'both')
with out:
clear_output(wait=True)
display(fig)
dropdown.observe(updateDropdown)
updateDropdown(None)
```
Let's keep going and add in our toggle box!
```
plt.close('all') # if you get a "too many figures open" warning
fig = plt.figure(figsize=(8,8))
# add axes by hand like last week
# order here is: left, bottom, width, height
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
image = ax.imshow(michigan, cmap='terrain')
fig.colorbar(image, extend = 'both')
out = ipywidgets.Output(layout=ipywidgets.Layout(height='500px', width = '500px'))
#dropdown = ipywidgets.Dropdown(options=plt.colormaps())
# just so that we can start with 'terrain'
dropdown = ipywidgets.Dropdown(options=plt.colormaps(), index=plt.colormaps().index('terrain'))
toggleButton = ipywidgets.ToggleButton(value=False,description='Log Norm?')
controls = ipywidgets.VBox([dropdown, toggleButton])
hbox=ipywidgets.HBox([out, controls])
display(hbox)
# (2) update figure based on toggle on/off
def updateToggle(change):
if change is not None:
#print(change)
#print(change['owner'])
#print(change['owner'].value)
# grab base colormap from other widget
cmap=plt.colormaps()[dropdown.index]
for a in fig.axes:
fig.delaxes(a)
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
# pick norm based on toggle button
if change['owner'].value:
norm = mpl_colors.SymLogNorm(10)
else:
norm = mpl_colors.Normalize()
image = ax.imshow(michigan, cmap=cmap, norm=norm)
fig.colorbar(image, extend = 'both')
with out:
clear_output(wait=True)
display(fig)
toggleButton.observe(updateToggle)
# (1) update figure based on dropdown
# AND MAKE SURE WE ADD IN RESULTS OF TOGGLE BUTTON!
def updateDropdown(change):
if change is not None:
cmap=plt.colormaps()[change['owner'].index]
for a in fig.axes:
fig.delaxes(a)
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
# pick norm based on toggle button
if toggleButton.value:
norm = mpl_colors.SymLogNorm(10)
else:
norm = mpl_colors.Normalize()
image = ax.imshow(michigan, cmap=cmap, norm=norm)
fig.colorbar(image, extend = 'both')
with out:
clear_output(wait=True)
display(fig)
dropdown.observe(updateDropdown)
updateDropdown(None)
updateToggle(None)
```
Finally, note that we can now move around each widget individually on our plot:
```
plt.close('all') # if you get a "too many figures open" warning
fig = plt.figure(figsize=(8,8))
# add axes by hand like last week
# order here is: left, bottom, width, height
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
image = ax.imshow(michigan, cmap='terrain')
fig.colorbar(image, extend = 'both')
out = ipywidgets.Output(layout=ipywidgets.Layout(height='500px', width = '500px'))
#dropdown = ipywidgets.Dropdown(options=plt.colormaps())
# just so that we can start with 'terrain'
dropdown = ipywidgets.Dropdown(options=plt.colormaps(), index=plt.colormaps().index('terrain'))
toggleButton = ipywidgets.ToggleButton(value=False,description='Log Norm?')
controls = ipywidgets.VBox([dropdown, toggleButton])
controls.layout.top = '200px' # UPDATED
hbox=ipywidgets.HBox([out, controls])
display(hbox)
# (2) update figure based on toggle on/off
def updateToggle(change):
if change is not None:
# grab base colormap from other widget
cmap=plt.colormaps()[dropdown.index]
for a in fig.axes:
fig.delaxes(a)
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
# pick norm based on toggle button
if change['owner'].value:
norm = mpl_colors.SymLogNorm(10)
else:
norm = mpl_colors.Normalize()
image = ax.imshow(michigan, cmap=cmap, norm=norm)
fig.colorbar(image, extend = 'both')
with out:
clear_output(wait=True)
display(fig)
toggleButton.observe(updateToggle)
# (1) update figure based on dropdown
# AND MAKE SURE WE ADD IN RESULTS OF TOGGLE BUTTON!
def updateDropdown(change):
if change is not None:
cmap=plt.colormaps()[change['owner'].index]
for a in fig.axes:
fig.delaxes(a)
ax = fig.add_axes([0.0, 0.15, 1.0, 0.8])
# pick norm based on toggle button
if toggleButton.value:
norm = mpl_colors.SymLogNorm(10)
else:
norm = mpl_colors.Normalize()
image = ax.imshow(michigan, cmap=cmap, norm=norm)
fig.colorbar(image, extend = 'both')
with out:
clear_output(wait=True)
display(fig)
dropdown.observe(updateDropdown)
updateDropdown(None)
updateToggle(None)
```
This seems a lot more complicated: why would we bother?
1. You don't have to if you don't want to! (At least for this week...)
1. It gives us finer-grained control over where to place things when we start building up multi-panel dashboards.
Taking some time to understand widgets in this context will help you design custom dashboards for your analysis & visualization needs.
`bqplot`, which we will use next week, uses this sort of layout options to link figures with widgets, but makes this sort of design a lot easier then what we just did!
| github_jupyter |
# Example of semi-supervised gan for mnist
Based on https://machinelearningmastery.com/semi-supervised-generative-adversarial-network/
```
from numpy import expand_dims
from numpy import zeros
from numpy import ones
from numpy import asarray
from numpy.random import randn
from numpy.random import randint
from keras.datasets.mnist import load_data
from keras.optimizers import Adam
from keras.models import Model
from keras.layers import Input
from keras.layers import Dense
from keras.layers import Reshape
from keras.layers import Flatten
from keras.layers import Conv2D
from keras.layers import Conv2DTranspose
from keras.layers import LeakyReLU
from keras.layers import Dropout
from keras.layers import Lambda
from keras.layers import Activation
from matplotlib import pyplot
from keras import backend
%matplotlib notebook
# custom activation function
def custom_activation(output):
logexpsum = backend.sum(backend.exp(output), axis=-1, keepdims=True)
result = logexpsum / (logexpsum + 1.0)
return result
# define the standalone supervised and unsupervised discriminator models
def define_discriminator(in_shape=(28,28,1), n_classes=10):
# image input
in_image = Input(shape=in_shape)
# downsample
fe = Conv2D(128, (3,3), strides=(2,2), padding='same')(in_image)
fe = LeakyReLU(alpha=0.2)(fe)
# downsample
fe = Conv2D(128, (3,3), strides=(2,2), padding='same')(fe)
fe = LeakyReLU(alpha=0.2)(fe)
# downsample
fe = Conv2D(128, (3,3), strides=(2,2), padding='same')(fe)
fe = LeakyReLU(alpha=0.2)(fe)
# flatten feature maps
fe = Flatten()(fe)
# dropout
fe = Dropout(0.4)(fe)
# output layer nodes
fe = Dense(n_classes)(fe)
# supervised output
c_out_layer = Activation('softmax')(fe)
# define and compile supervised discriminator model
c_model = Model(in_image, c_out_layer)
c_model.compile(loss='sparse_categorical_crossentropy', optimizer=Adam(lr=0.0002, beta_1=0.5), metrics=['accuracy'])
# unsupervised output
d_out_layer = Lambda(custom_activation)(fe)
# define and compile unsupervised discriminator model
d_model = Model(in_image, d_out_layer)
d_model.compile(loss='binary_crossentropy', optimizer=Adam(lr=0.0002, beta_1=0.5))
return d_model, c_model
# define the standalone generator model
def define_generator(latent_dim):
# image generator input
in_lat = Input(shape=(latent_dim,))
# foundation for 7x7 image
n_nodes = 128 * 7 * 7
gen = Dense(n_nodes)(in_lat)
gen = LeakyReLU(alpha=0.2)(gen)
gen = Reshape((7, 7, 128))(gen)
# upsample to 14x14
gen = Conv2DTranspose(128, (4,4), strides=(2,2), padding='same')(gen)
gen = LeakyReLU(alpha=0.2)(gen)
# upsample to 28x28
gen = Conv2DTranspose(128, (4,4), strides=(2,2), padding='same')(gen)
gen = LeakyReLU(alpha=0.2)(gen)
# output
out_layer = Conv2D(1, (7,7), activation='tanh', padding='same')(gen)
# define model
model = Model(in_lat, out_layer)
return model
# define the combined generator and discriminator model, for updating the generator
def define_gan(g_model, d_model):
# make weights in the discriminator not trainable
d_model.trainable = False
# connect image output from generator as input to discriminator
gan_output = d_model(g_model.output)
# define gan model as taking noise and outputting a classification
model = Model(g_model.input, gan_output)
# compile model
opt = Adam(lr=0.0002, beta_1=0.5)
model.compile(loss='binary_crossentropy', optimizer=opt)
return model
# load the images
def load_real_samples():
# load dataset
(trainX, trainy), (_, _) = load_data()
# expand to 3d, e.g. add channels
X = expand_dims(trainX, axis=-1)
# convert from ints to floats
X = X.astype('float32')
# scale from [0,255] to [-1,1]
X = (X - 127.5) / 127.5
print(X.shape, trainy.shape)
return [X, trainy]
# select a supervised subset of the dataset, ensures classes are balanced
def select_supervised_samples(dataset, n_samples=100, n_classes=10):
X, y = dataset
X_list, y_list = list(), list()
n_per_class = int(n_samples / n_classes)
for i in range(n_classes):
# get all images for this class
X_with_class = X[y == i]
# choose random instances
ix = randint(0, len(X_with_class), n_per_class)
# add to list
[X_list.append(X_with_class[j]) for j in ix]
[y_list.append(i) for j in ix]
return asarray(X_list), asarray(y_list)
# select real samples
def generate_real_samples(dataset, n_samples):
# split into images and labels
images, labels = dataset
# choose random instances
ix = randint(0, images.shape[0], n_samples)
# select images and labels
X, labels = images[ix], labels[ix]
# generate class labels
y = ones((n_samples, 1))
return [X, labels], y
# generate points in latent space as input for the generator
def generate_latent_points(latent_dim, n_samples):
# generate points in the latent space
z_input = randn(latent_dim * n_samples)
# reshape into a batch of inputs for the network
z_input = z_input.reshape(n_samples, latent_dim)
return z_input
# use the generator to generate n fake examples, with class labels
def generate_fake_samples(generator, latent_dim, n_samples):
# generate points in latent space
z_input = generate_latent_points(latent_dim, n_samples)
# predict outputs
images = generator.predict(z_input)
# create class labels
y = zeros((n_samples, 1))
return images, y
# generate samples and save as a plot and save the model
def summarize_performance(step, g_model, c_model, latent_dim, dataset, n_samples=100):
# prepare fake examples
X, _ = generate_fake_samples(g_model, latent_dim, n_samples)
# scale from [-1,1] to [0,1]
X = (X + 1) / 2.0
# plot images
for i in range(100):
# define subplot
pyplot.subplot(10, 10, 1 + i)
# turn off axis
pyplot.axis('off')
# plot raw pixel data
pyplot.imshow(X[i, :, :, 0], cmap='gray_r')
# save plot to file
filename1 = 'generated_plot_%04d.png' % (step+1)
pyplot.savefig(filename1)
pyplot.close()
# evaluate the classifier model
X, y = dataset
_, acc = c_model.evaluate(X, y, verbose=0)
print('Classifier Accuracy: %.3f%%' % (acc * 100))
# save the generator model
filename2 = 'g_model_%04d.h5' % (step+1)
g_model.save(filename2)
# save the classifier model
filename3 = 'c_model_%04d.h5' % (step+1)
c_model.save(filename3)
print('>Saved: %s, %s, and %s' % (filename1, filename2, filename3))
# train the generator and discriminator
def train(g_model, d_model, c_model, gan_model, dataset, latent_dim, n_epochs=20, n_batch=100):
# select supervised dataset
X_sup, y_sup = select_supervised_samples(dataset)
print(X_sup.shape, y_sup.shape)
# calculate the number of batches per training epoch
bat_per_epo = int(dataset[0].shape[0] / n_batch)
# calculate the number of training iterations
n_steps = bat_per_epo * n_epochs
# calculate the size of half a batch of samples
half_batch = int(n_batch / 2)
print('n_epochs=%d, n_batch=%d, 1/2=%d, b/e=%d, steps=%d' % (n_epochs, n_batch, half_batch, bat_per_epo, n_steps))
# manually enumerate epochs
for i in range(n_steps):
# update supervised discriminator (c)
[Xsup_real, ysup_real], _ = generate_real_samples([X_sup, y_sup], half_batch)
c_loss, c_acc = c_model.train_on_batch(Xsup_real, ysup_real)
# update unsupervised discriminator (d)
[X_real, _], y_real = generate_real_samples(dataset, half_batch)
d_loss1 = d_model.train_on_batch(X_real, y_real)
X_fake, y_fake = generate_fake_samples(g_model, latent_dim, half_batch)
d_loss2 = d_model.train_on_batch(X_fake, y_fake)
# update generator (g)
X_gan, y_gan = generate_latent_points(latent_dim, n_batch), ones((n_batch, 1))
g_loss = gan_model.train_on_batch(X_gan, y_gan)
# summarize loss on this batch
print('>%d, c[%.3f,%.0f], d[%.3f,%.3f], g[%.3f]' % (i+1, c_loss, c_acc*100, d_loss1, d_loss2, g_loss))
# evaluate the model performance every so often
if (i+1) % (bat_per_epo * 1) == 0:
summarize_performance(i, g_model, c_model, latent_dim, dataset)
# size of the latent space
latent_dim = 100
# create the discriminator models
d_model, c_model = define_discriminator()
# create the generator
g_model = define_generator(latent_dim)
# create the gan
gan_model = define_gan(g_model, d_model)
# load image data
dataset = load_real_samples()
# train model
train(g_model, d_model, c_model, gan_model, dataset, latent_dim)
```
| github_jupyter |
# Facial Keypoint Detection
This project will be all about defining and training a convolutional neural network to perform facial keypoint detection, and using computer vision techniques to transform images of faces. The first step in any challenge like this will be to load and visualize the data you'll be working with.
Let's take a look at some examples of images and corresponding facial keypoints.
<img src='images/key_pts_example.png' width=50% height=50%/>
Facial keypoints (also called facial landmarks) are the small magenta dots shown on each of the faces in the image above. In each training and test image, there is a single face and **68 keypoints, with coordinates (x, y), for that face**. These keypoints mark important areas of the face: the eyes, corners of the mouth, the nose, etc. These keypoints are relevant for a variety of tasks, such as face filters, emotion recognition, pose recognition, and so on. Here they are, numbered, and you can see that specific ranges of points match different portions of the face.
<img src='images/landmarks_numbered.jpg' width=30% height=30%/>
---
## Load and Visualize Data
The first step in working with any dataset is to become familiar with your data; you'll need to load in the images of faces and their keypoints and visualize them! This set of image data has been extracted from the [YouTube Faces Dataset](https://www.cs.tau.ac.il/~wolf/ytfaces/), which includes videos of people in YouTube videos. These videos have been fed through some processing steps and turned into sets of image frames containing one face and the associated keypoints.
#### Training and Testing Data
This facial keypoints dataset consists of 5770 color images. All of these images are separated into either a training or a test set of data.
* 3462 of these images are training images, for you to use as you create a model to predict keypoints.
* 2308 are test images, which will be used to test the accuracy of your model.
The information about the images and keypoints in this dataset are summarized in CSV files, which we can read in using `pandas`. Let's read the training CSV and get the annotations in an (N, 2) array where N is the number of keypoints and 2 is the dimension of the keypoint coordinates (x, y).
---
```
# import the required libraries
import glob
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import cv2
key_pts_frame = pd.read_csv('data/training_frames_keypoints.csv')
n = 0
image_name = key_pts_frame.iloc[n, 0]
key_pts = key_pts_frame.iloc[n, 1:].as_matrix()
key_pts = key_pts.astype('float').reshape(-1, 2)
print('Image name: ', image_name)
print('Landmarks shape: ', key_pts.shape)
print('First 4 key pts: {}'.format(key_pts[:4]))
# print out some stats about the data
print('Number of images: ', key_pts_frame.shape[0])
```
## Look at some images
Below, is a function `show_keypoints` that takes in an image and keypoints and displays them. As you look at this data, **note that these images are not all of the same size**, and neither are the faces! To eventually train a neural network on these images, we'll need to standardize their shape.
```
def show_keypoints(image, key_pts):
"""Show image with keypoints"""
plt.imshow(image)
plt.scatter(key_pts[:, 0], key_pts[:, 1], s=20, marker='.', c='m')
# Display a few different types of images by changing the index n
# select an image by index in our data frame
n = 0
image_name = key_pts_frame.iloc[n, 0]
key_pts = key_pts_frame.iloc[n, 1:].as_matrix()
key_pts = key_pts.astype('float').reshape(-1, 2)
plt.figure(figsize=(5, 5))
show_keypoints(mpimg.imread(os.path.join('data/training/', image_name)), key_pts)
plt.show()
```
## Dataset class and Transformations
To prepare our data for training, we'll be using PyTorch's Dataset class. Much of this this code is a modified version of what can be found in the [PyTorch data loading tutorial](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html).
#### Dataset class
``torch.utils.data.Dataset`` is an abstract class representing a
dataset. This class will allow us to load batches of image/keypoint data, and uniformly apply transformations to our data, such as rescaling and normalizing images for training a neural network.
Your custom dataset should inherit ``Dataset`` and override the following
methods:
- ``__len__`` so that ``len(dataset)`` returns the size of the dataset.
- ``__getitem__`` to support the indexing such that ``dataset[i]`` can
be used to get the i-th sample of image/keypoint data.
Let's create a dataset class for our face keypoints dataset. We will
read the CSV file in ``__init__`` but leave the reading of images to
``__getitem__``. This is memory efficient because all the images are not
stored in the memory at once but read as required.
A sample of our dataset will be a dictionary
``{'image': image, 'keypoints': key_pts}``. Our dataset will take an
optional argument ``transform`` so that any required processing can be
applied on the sample. We will see the usefulness of ``transform`` in the
next section.
```
from torch.utils.data import Dataset, DataLoader
class FacialKeypointsDataset(Dataset):
"""Face Landmarks dataset."""
def __init__(self, csv_file, root_dir, transform=None):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.key_pts_frame = pd.read_csv(csv_file)
self.root_dir = root_dir
self.transform = transform
def __len__(self):
return len(self.key_pts_frame)
def __getitem__(self, idx):
image_name = os.path.join(self.root_dir,
self.key_pts_frame.iloc[idx, 0])
image = mpimg.imread(image_name)
# if image has an alpha color channel, get rid of it
if(image.shape[2] == 4):
image = image[:,:,0:3]
key_pts = self.key_pts_frame.iloc[idx, 1:].as_matrix()
key_pts = key_pts.astype('float').reshape(-1, 2)
sample = {'image': image, 'keypoints': key_pts}
if self.transform:
sample = self.transform(sample)
return sample
```
Now that we've defined this class, let's instantiate the dataset and display some images.
```
# Construct the dataset
face_dataset = FacialKeypointsDataset(csv_file='data/training_frames_keypoints.csv',
root_dir='data/training/')
# print some stats about the dataset
print('Length of dataset: ', len(face_dataset))
# Display a few of the images from the dataset
num_to_display = 3
for i in range(num_to_display):
# define the size of images
fig = plt.figure(figsize=(20,10))
# randomly select a sample
rand_i = np.random.randint(0, len(face_dataset))
sample = face_dataset[rand_i]
# print the shape of the image and keypoints
print(i, sample['image'].shape, sample['keypoints'].shape)
ax = plt.subplot(1, num_to_display, i + 1)
ax.set_title('Sample #{}'.format(i))
# Using the same display function, defined earlier
show_keypoints(sample['image'], sample['keypoints'])
```
## Transforms
Now, the images above are not of the same size, and neural networks often expect images that are standardized; a fixed size, with a normalized range for color ranges and coordinates, and (for PyTorch) converted from numpy lists and arrays to Tensors.
Therefore, we will need to write some pre-processing code.
Let's create four transforms:
- ``Normalize``: to convert a color image to grayscale values with a range of [0,1] and normalize the keypoints to be in a range of about [-1, 1]
- ``Rescale``: to rescale an image to a desired size.
- ``RandomCrop``: to crop an image randomly.
- ``ToTensor``: to convert numpy images to torch images.
We will write them as callable classes instead of simple functions so
that parameters of the transform need not be passed everytime it's
called. For this, we just need to implement ``__call__`` method and
(if we require parameters to be passed in), the ``__init__`` method.
We can then use a transform like this:
tx = Transform(params)
transformed_sample = tx(sample)
Observe below how these transforms are generally applied to both the image and its keypoints.
```
import torch
from torchvision import transforms, utils
# tranforms
class Normalize(object):
"""Convert a color image to grayscale and normalize the color range to [0,1]."""
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
image_copy = np.copy(image)
key_pts_copy = np.copy(key_pts)
# convert image to grayscale
image_copy = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
# scale color range from [0, 255] to [0, 1]
image_copy= image_copy/255.0
# scale keypoints to be centered around 0 with a range of [-1, 1]
# mean = 100, sqrt = 50, so, pts should be (pts - 100)/50
key_pts_copy = (key_pts_copy - 100)/50.0
return {'image': image_copy, 'keypoints': key_pts_copy}
class Rescale(object):
"""Rescale the image in a sample to a given size.
Args:
output_size (tuple or int): Desired output size. If tuple, output is
matched to output_size. If int, smaller of image edges is matched
to output_size keeping aspect ratio the same.
"""
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
self.output_size = output_size
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
h, w = image.shape[:2]
if isinstance(self.output_size, int):
if h > w:
new_h, new_w = self.output_size * h / w, self.output_size
else:
new_h, new_w = self.output_size, self.output_size * w / h
else:
new_h, new_w = self.output_size
new_h, new_w = int(new_h), int(new_w)
img = cv2.resize(image, (new_w, new_h))
# scale the pts, too
key_pts = key_pts * [new_w / w, new_h / h]
return {'image': img, 'keypoints': key_pts}
class RandomCrop(object):
"""Crop randomly the image in a sample.
Args:
output_size (tuple or int): Desired output size. If int, square crop
is made.
"""
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
if isinstance(output_size, int):
self.output_size = (output_size, output_size)
else:
assert len(output_size) == 2
self.output_size = output_size
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
h, w = image.shape[:2]
new_h, new_w = self.output_size
top = np.random.randint(0, h - new_h)
left = np.random.randint(0, w - new_w)
image = image[top: top + new_h,
left: left + new_w]
key_pts = key_pts - [left, top]
return {'image': image, 'keypoints': key_pts}
class ToTensor(object):
"""Convert ndarrays in sample to Tensors."""
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
# if image has no grayscale color channel, add one
if(len(image.shape) == 2):
# add that third color dim
image = image.reshape(image.shape[0], image.shape[1], 1)
# swap color axis because
# numpy image: H x W x C
# torch image: C X H X W
image = image.transpose((2, 0, 1))
return {'image': torch.from_numpy(image),
'keypoints': torch.from_numpy(key_pts)}
```
## Test out the transforms
Let's test these transforms out to make sure they behave as expected. As you look at each transform, note that, in this case, **order does matter**. For example, you cannot crop a image using a value smaller than the original image (and the orginal images vary in size!), but, if you first rescale the original image, you can then crop it to any size smaller than the rescaled size.
```
# test out some of these transforms
rescale = Rescale(100)
crop = RandomCrop(50)
composed = transforms.Compose([Rescale(250),
RandomCrop(224)])
# apply the transforms to a sample image
test_num = 500
sample = face_dataset[test_num]
fig = plt.figure()
for i, tx in enumerate([rescale, crop, composed]):
transformed_sample = tx(sample)
ax = plt.subplot(1, 3, i + 1)
plt.tight_layout()
ax.set_title(type(tx).__name__)
show_keypoints(transformed_sample['image'], transformed_sample['keypoints'])
plt.show()
```
## Create the transformed dataset
Apply the transforms in order to get grayscale images of the same shape. Verify that your transform works by printing out the shape of the resulting data (printing out a few examples should show you a consistent tensor size).
```
# define the data tranform
# order matters! i.e. rescaling should come before a smaller crop
data_transform = transforms.Compose([Rescale(250),
RandomCrop(224),
Normalize(),
ToTensor()])
# create the transformed dataset
transformed_dataset = FacialKeypointsDataset(csv_file='data/training_frames_keypoints.csv',
root_dir='data/training/',
transform=data_transform)
# print some stats about the transformed data
print('Number of images: ', len(transformed_dataset))
# make sure the sample tensors are the expected size
for i in range(5):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['keypoints'].size())
```
## Data Iteration and Batching
Right now, we are iterating over this data using a ``for`` loop, but we are missing out on a lot of PyTorch's dataset capabilities, specifically the abilities to:
- Batch the data
- Shuffle the data
- Load the data in parallel using ``multiprocessing`` workers.
``torch.utils.data.DataLoader`` is an iterator which provides all these
features, and we'll see this in use in the *next* notebook, Notebook 2, when we load data in batches to train a neural network!
---
## Ready to Train!
Now that you've seen how to load and transform our data, you're ready to build a neural network to train on this data.
In the next notebook, you'll be tasked with creating a CNN for facial keypoint detection.
| github_jupyter |
# Address Segmentation
Conversion of address points into segmented address ranges along a road network.
**Notes:** The following guide assumes data has already been preprocessed including data scrubbing and filtering.
```
import contextily as ctx
import geopandas as gpd
import math
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import re
import shapely
from bisect import bisect
from collections import OrderedDict
from IPython.display import display_html
from matplotlib_scalebar.scalebar import ScaleBar
from operator import itemgetter
from shapely.geometry import LineString, Point
# Define index of example roadseg segment.
ex_idx = 264
ex_place = "City of Yellowknife"
# Define join fields.
join_roadseg = "roadname"
join_addresses = "street"
# Define helper functions.
def groupby_to_list(df, group_field, list_field):
"""
Helper function: faster alternative to pandas groupby.apply/agg(list).
Groups records by one or more fields and compiles an output field into a list for each group.
"""
if isinstance(group_field, list):
for field in group_field:
if df[field].dtype.name != "geometry":
df[field] = df[field].astype("U")
transpose = df.sort_values(group_field)[[*group_field, list_field]].values.T
keys, vals = np.column_stack(transpose[:-1]), transpose[-1]
keys_unique, keys_indexes = np.unique(keys.astype("U") if isinstance(keys, np.object) else keys,
axis=0, return_index=True)
else:
keys, vals = df.sort_values(group_field)[[group_field, list_field]].values.T
keys_unique, keys_indexes = np.unique(keys, return_index=True)
vals_arrays = np.split(vals, keys_indexes[1:])
return pd.Series([list(vals_array) for vals_array in vals_arrays], index=keys_unique).copy(deep=True)
```
## Step 1. Load dataframes and configure attributes
Loads dataframes into geopandas and separates address numbers and suffixes, if required.
```
# Load dataframes.
addresses = gpd.read_file("C:/scratch/City_Of_Yellowknife.gpkg", layer="addresses")
roadseg = gpd.read_file("C:/scratch/City_Of_Yellowknife.gpkg", layer="roads")
# Configure attributes - number and suffix.
addresses["suffix"] = addresses["number"].map(lambda val: re.sub(pattern="\\d+", repl="", string=val, flags=re.I))
addresses["number"] = addresses["number"].map(lambda val: re.sub(pattern="[^\\d]", repl="", string=val, flags=re.I)).map(int)
addresses.head()
roadseg.head()
```
## Preview data
**Note:** this code block is for visual purposes only.
```
# Fetch basemaps.
# Note: basemaps are retrieved in EPSG:3857 and, therefore, dataframes should also use this crs.
basemaps = list()
basemaps.append(ctx.bounds2img(*roadseg.total_bounds, ll=False, source=ctx.providers.Esri.WorldImagery))
basemaps.append(ctx.bounds2img(*roadseg.loc[roadseg.index==ex_idx].total_bounds, ll=False,
source=ctx.providers.Esri.WorldImagery))
# Configure local positional distortion (for scalebar dx parameter).
ymin, ymax = itemgetter(1, 3)(roadseg[roadseg.index==ex_idx].to_crs("EPSG:4617").total_bounds)
lat = ymin + ((ymax - ymin) / 2)
dx = math.cos(math.radians(lat))
# Create data for viewing.
starting_pt = gpd.GeoDataFrame(geometry=[Point(roadseg.loc[roadseg.index==ex_idx]["geometry"].iloc[0].coords[0])],
crs=addresses.crs)
# Configure plots.
fig, ax = plt.subplots(1, 2, figsize=(12, 7), tight_layout=True)
for plt_idx, title in enumerate(["All Data", f"roadseg={ex_idx}"]):
ax[plt_idx].imshow(basemaps[plt_idx][0], extent=basemaps[plt_idx][1])
if plt_idx == 0:
addresses.plot(ax=ax[plt_idx], color="red", label="addresses", markersize=2)
roadseg.plot(ax=ax[plt_idx], color="cyan", label="roadseg", linewidth=1)
else:
addresses.plot(ax=ax[plt_idx], color="red", label="addresses", linewidth=2)
starting_pt.plot(ax=ax[plt_idx], color="gold", label=f"roadseg={ex_idx}, 1st point", linewidth=2)
roadseg.loc[roadseg.index==ex_idx].plot(ax=ax[plt_idx], color="yellow", label=f"roadseg={ex_idx}", linewidth=2)
roadseg.loc[roadseg.index!=ex_idx].plot(ax=ax[plt_idx], color="cyan", label="roadseg", linewidth=1)
ax[plt_idx].add_artist(ScaleBar(dx=dx, units="m", location="lower left", pad=0.5, color="black"))
ax[plt_idx].axes.xaxis.set_visible(False)
ax[plt_idx].axes.yaxis.set_visible(False)
ax[plt_idx].set_title(title, fontsize=12)
ax[plt_idx].set_xlim(itemgetter(0, 1)(basemaps[plt_idx][1]))
ax[plt_idx].set_ylim(itemgetter(2, 3)(basemaps[plt_idx][1]))
plt.suptitle(ex_place, fontsize=12)
plt.legend(loc="center left", bbox_to_anchor=(1.0, 0.5), fontsize=12)
plt.show()
```
## Step 2. Configure address to roadseg linkages
Links addresses to the nearest, matching road segment.
```
# Link addresses and roadseg on join fields.
addresses["addresses_index"] = addresses.index
roadseg["roadseg_index"] = roadseg.index
merge = addresses.merge(roadseg[[join_roadseg, "roadseg_index"]], how="left", left_on=join_addresses, right_on=join_roadseg)
addresses["roadseg_index"] = groupby_to_list(merge, "addresses_index", "roadseg_index")
addresses.drop(columns=["addresses_index"], inplace=True)
roadseg.drop(columns=["roadseg_index"], inplace=True)
# Discard non-linked addresses.
addresses.drop(addresses[addresses["roadseg_index"].map(itemgetter(0)).isna()].index, axis=0, inplace=True)
# Convert linkages to integer tuples, if possible.
def as_int(val):
try:
return int(val)
except ValueError:
return val
addresses["roadseg_index"] = addresses["roadseg_index"].map(lambda vals: tuple(set(map(as_int, vals))))
addresses.head()
# Reduce linkages to one roadseg index per address.
# Configure roadseg geometry lookup dictionary.
roadseg_geom_lookup = roadseg["geometry"].to_dict()
def get_nearest_linkage(pt, roadseg_indexes):
"""Returns the roadseg index associated with the nearest roadseg geometry to the given address point."""
# Get roadseg geometries.
roadseg_geometries = itemgetter(*roadseg_indexes)(roadseg_geom_lookup)
# Get roadseg distances from address point.
roadseg_distances = tuple(map(lambda road: pt.distance(road), roadseg_geometries))
# Get the roadseg index associated with the smallest distance.
roadseg_index = roadseg_indexes[roadseg_distances.index(min(roadseg_distances))]
return roadseg_index
# Flag plural linkages.
flag_plural = addresses["roadseg_index"].map(len) > 1
# Reduce plural linkages to the road segment with the lowest (nearest) geometric distance.
addresses.loc[flag_plural, "roadseg_index"] = addresses[flag_plural][["geometry", "roadseg_index"]].apply(
lambda row: get_nearest_linkage(*row), axis=1)
# Unpack first tuple element for singular linkages.
addresses.loc[~flag_plural, "roadseg_index"] = addresses[~flag_plural]["roadseg_index"].map(itemgetter(0))
# Compile linked roadseg geometry for each address.
addresses["roadseg_geometry"] = addresses.merge(
roadseg["geometry"], how="left", left_on="roadseg_index", right_index=True)["geometry_y"]
addresses.head()
```
## Step 3. Configure address parity
Computes address-roadseg parity (left / right side).
```
def get_parity(pt, vector):
"""
Determines the parity (left or right side) of an address point relative to a roadseg vector.
Parity is derived from the determinant of the vectors formed by the road segment and the address-to-roadseg
vectors. A positive determinant indicates 'left' parity and negative determinant indicates 'right' parity.
"""
det = (vector[1][0] - vector[0][0]) * (pt.y - vector[0][1]) - \
(vector[1][1] - vector[0][1]) * (pt.x - vector[0][0])
sign = np.sign(det)
return "l" if sign == 1 else "r"
def get_road_vector(pt, segment):
"""
Returns the following:
a) the distance of the address intersection along the road segment.
b) the vector comprised of the road segment coordinates immediately before and after the address
intersection point.
"""
# For all road segment points and the intersection point, calculate the distance along the road segment.
# Note: always use the length as the distance for the last point to avoid distance=0 for looped roads.
node_distance = (*map(lambda coord: segment.project(Point(coord)), segment.coords[:-1]), segment.length)
intersection_distance = segment.project(pt)
# Compute the index of the intersection point within the road segment points, based on distances.
intersection_index = bisect(node_distance, intersection_distance)
# Conditionally compile the road segment points, as a vector, immediately bounding the intersection point.
# Intersection matches a pre-existing road segment point.
if intersection_distance in node_distance:
# Intersection matches the first road segment point.
if intersection_index == 1:
vector = itemgetter(intersection_index - 1, intersection_index)(segment.coords)
# Intersection matches the last road segment point.
elif intersection_index == len(node_distance):
vector = itemgetter(intersection_index - 2, intersection_index - 1)(segment.coords)
# Intersection matches an interior road segment point.
else:
vector = itemgetter(intersection_index - 2, intersection_index)(segment.coords)
# Intersection matches no pre-existing road segment point.
else:
vector = itemgetter(intersection_index - 1, intersection_index)(segment.coords)
return intersection_distance, vector
# Get point of intersection between each address and the linked road segment.
addresses["intersection"] = addresses[["geometry", "roadseg_geometry"]].apply(
lambda row: itemgetter(-1)(shapely.ops.nearest_points(*row)), axis=1)
# Get the following:
# a) the distance of the intersection point along the linked road segment.
# b) the road segment vector which bounds the intersection point.
# i.e. vector formed by the coordinates immediately before and after the intersection point.
results = addresses[["intersection", "roadseg_geometry"]].apply(lambda row: get_road_vector(*row), axis=1)
addresses["distance"] = results.map(itemgetter(0))
addresses["roadseg_vector"] = results.map(itemgetter(1))
# Get address parity.
addresses["parity"] = addresses[["geometry", "roadseg_vector"]].apply(
lambda row: get_parity(*row), axis=1)
addresses[["geometry", "roadseg_geometry", "intersection", "distance", "roadseg_vector", "parity"]].head()
```
## View relationship between parity variables
View the relationship between address points, bounding roadseg vectors, address-roadseg intersection points, and the computed parity.
**Note:** this code block is for visual purposes only.
```
# Create geometries for viewing.
bounding_vectors = gpd.GeoDataFrame(geometry=addresses["roadseg_vector"].map(LineString), crs=addresses.crs)
intersection = gpd.GeoDataFrame(addresses["parity"], geometry=addresses[["geometry", "intersection"]].apply(
lambda row: LineString([pt.coords[0][:2] for pt in row]), axis=1), crs=addresses.crs)
# Configure plots.
fig, ax = plt.subplots(1, 2, figsize=(14.5, 7), tight_layout=True)
for plt_idx, title in enumerate(["Parity Input", "Parity Output"]):
ax[plt_idx].imshow(basemaps[1][0], extent=basemaps[1][1])
addresses.plot(ax=ax[plt_idx], color="red", label="addresses", linewidth=2)
starting_pt.plot(ax=ax[plt_idx], color="gold", label=f"roadseg={ex_idx}, 1st point", linewidth=2)
roadseg.loc[roadseg.index==ex_idx].plot(ax=ax[plt_idx], color="yellow", label=f"roadseg={ex_idx}", linewidth=2)
roadseg.loc[roadseg.index!=ex_idx].plot(ax=ax[plt_idx], color="cyan", label="roadseg", linewidth=1)
if plt_idx == 0:
intersection.plot(ax=ax[plt_idx], color="orange", label="address-roadseg intersection", linewidth=2)
bounding_vectors.plot(ax=ax[plt_idx], color="magenta", label="bounding roadseg vectors", linewidth=2)
else:
intersection.loc[intersection["parity"]=="l"].plot(
ax=ax[plt_idx], color="blue", label="address-roadseg intersection (left)", linewidth=2)
intersection.loc[intersection["parity"]=="r"].plot(
ax=ax[plt_idx], color="lime", label="address-roadseg intersection (right)", linewidth=2)
ax[plt_idx].add_artist(ScaleBar(dx=dx, units="m", location="lower left", pad=0.5, color="black"))
ax[plt_idx].axes.xaxis.set_visible(False)
ax[plt_idx].axes.yaxis.set_visible(False)
ax[plt_idx].set_title(title, fontsize=12)
ax[plt_idx].set_xlim(itemgetter(0, 1)(basemaps[1][1]))
ax[plt_idx].set_ylim(itemgetter(2, 3)(basemaps[1][1]))
plt.suptitle(ex_place, fontsize=12)
legend_icons = list()
legend_labels = list()
for axis in ax:
legend_items = list(zip(*[items for items in zip(*axis.get_legend_handles_labels()) if items[1] not in legend_labels]))
legend_icons.extend(legend_items[0])
legend_labels.extend(legend_items[1])
plt.legend(legend_icons, legend_labels, loc="center left", bbox_to_anchor=(1.0, 0.5), fontsize=12)
plt.show()
```
## Step 4. Configure address ranges (addrange) and attributes
Groups addresses into ranges then computes the addrange attributes.
```
def get_digdirfg(sequence):
"""Returns the digdirfg attribute for the given sequence of address numbers."""
sequence = list(sequence)
# Return digitizing direction for single addresses.
if len(sequence) == 1:
return "Not Applicable"
# Derive digitizing direction from sequence sorting direction.
if sequence == sorted(sequence):
return "Same Direction"
else:
return "Opposite Direction"
def get_hnumstr(sequence):
"""Returns the hnumstr attribute for the given sequence of address numbers."""
sequence = list(sequence)
# Validate structure for single addresses.
if len(sequence) == 1:
return "Even" if (sequence[0] % 2 == 0) else "Odd"
# Configure sequence sort status.
if sequence == sorted(sequence) or sequence == sorted(sequence, reverse=True):
# Configure sequence parities.
parities = tuple(map(lambda number: number % 2 == 0, sequence))
# Validate structure for sorted address ranges.
if all(parities):
return "Even"
elif not any(parities):
return "Odd"
else:
return "Mixed"
# Return structure for unsorted address ranges.
else:
return "Irregular"
def get_number_sequence(addresses):
"""Returns the filtered number sequence for the given addresses."""
# Separate address components.
numbers, suffixes, distances = tuple(zip(*addresses))
# Reduce addresses at a duplicated intersection distance to only the first instance.
if len(distances) == len(set(distances)):
sequence = numbers
else:
sequence = pd.DataFrame({"number": numbers, "suffix": suffixes, "distance": distances}).drop_duplicates(
subset="distance", keep="first")["number"].to_list()
# Remove duplicated addresses.
sequence = list(OrderedDict.fromkeys(sequence))
return sequence
def sort_addresses(numbers, suffixes, distances):
"""
Sorts the addresses successively by:
1) distance - the distance of the intersection point along the road segment.
2) number
3) suffix
Taking into account the directionality of the addresses relative to the road segment.
"""
# Create individual address tuples from separated address components.
addresses = tuple(zip(numbers, suffixes, distances))
# Apply initial sorting, by distance, to identify address directionality.
addresses_sorted = sorted(addresses, key=itemgetter(2))
directionality = -1 if addresses_sorted[0][0] > addresses_sorted[-1][0] else 1
# Sort addresses - same direction.
if directionality == 1:
return tuple(sorted(addresses, key=itemgetter(2, 1, 0)))
# Sort addresses - opposite direction.
else:
return tuple(sorted(sorted(sorted(
addresses, key=itemgetter(1), reverse=True),
key=itemgetter(0), reverse=True),
key=itemgetter(2)))
```
### Step 4.1. Group and sort addresses
Groups addresses by roadseg index and parity and sorts each grouping prior to configuring addrange attributes.
```
# Split address dataframe by parity.
addresses_l = addresses[addresses["parity"] == "l"].copy(deep=True)
addresses_r = addresses[addresses["parity"] == "r"].copy(deep=True)
# Create dataframes from grouped addresses.
cols = ("number", "suffix", "distance")
addresses_l = pd.DataFrame({col: groupby_to_list(addresses_l, "roadseg_index", col) for col in cols})
addresses_r = pd.DataFrame({col: groupby_to_list(addresses_r, "roadseg_index", col) for col in cols})
# Sort addresses.
addresses_l = addresses_l.apply(lambda row: sort_addresses(*row), axis=1)
addresses_r = addresses_r.apply(lambda row: sort_addresses(*row), axis=1)
```
### View example address grouping
**Note:** this code block is for visual purposes only.
```
# View data.
vals_l = list(zip(*addresses_l.loc[addresses_l.index==ex_idx].iloc[0]))
vals_r = list(zip(*addresses_r.loc[addresses_r.index==ex_idx].iloc[0]))
cols = ("number", "suffix", "distance")
left = pd.DataFrame({("Left Parity", cols[idx]): vals for idx, vals in enumerate(vals_l)})
right = pd.DataFrame({("Right Parity", cols[idx]): vals for idx, vals in enumerate(vals_r)})
display_html(left.style.set_table_attributes("style='display:inline'")._repr_html_()+
"<pre style='display:inline'> </pre>"+
right.style.set_table_attributes("style='display:inline'")._repr_html_(), raw=True)
```
### Step 4.2. Configure addrange attributes
```
# Configure addrange attributes.
addrange = pd.DataFrame(index=map(int, {*addresses_l.index, *addresses_r.index}))
# Configure addrange attributes - hnumf, hnuml.
addrange.loc[addresses_l.index, "l_hnumf"] = addresses_l.map(lambda addresses: addresses[0][0])
addrange.loc[addresses_l.index, "l_hnuml"] = addresses_l.map(lambda addresses: addresses[-1][0])
addrange.loc[addresses_r.index, "r_hnumf"] = addresses_r.map(lambda addresses: addresses[0][0])
addrange.loc[addresses_r.index, "r_hnuml"] = addresses_r.map(lambda addresses: addresses[-1][0])
# Configuring addrange attributes - hnumsuff, hnumsufl.
addrange.loc[addresses_l.index, "l_hnumsuff"] = addresses_l.map(lambda addresses: addresses[0][1])
addrange.loc[addresses_l.index, "l_hnumsufl"] = addresses_l.map(lambda addresses: addresses[-1][1])
addrange.loc[addresses_r.index, "r_hnumsuff"] = addresses_r.map(lambda addresses: addresses[0][1])
addrange.loc[addresses_r.index, "r_hnumsufl"] = addresses_r.map(lambda addresses: addresses[-1][1])
# Configuring addrange attributes - hnumtypf, hnumtypl.
addrange.loc[addresses_l.index, "l_hnumtypf"] = addresses_l.map(lambda addresses: "Actual Located")
addrange.loc[addresses_l.index, "l_hnumtypl"] = addresses_l.map(lambda addresses: "Actual Located")
addrange.loc[addresses_r.index, "r_hnumtypf"] = addresses_r.map(lambda addresses: "Actual Located")
addrange.loc[addresses_r.index, "r_hnumtypl"] = addresses_r.map(lambda addresses: "Actual Located")
# Get address number sequence.
address_sequence_l = addresses_l.map(get_number_sequence)
address_sequence_r = addresses_r.map(get_number_sequence)
# Configure addrange attributes - hnumstr.
addrange.loc[addresses_l.index, "l_hnumstr"] = address_sequence_l.map(get_hnumstr)
addrange.loc[addresses_r.index, "r_hnumstr"] = address_sequence_r.map(get_hnumstr)
# Configure addrange attributes - digdirfg.
addrange.loc[addresses_l.index, "l_digdirfg"] = address_sequence_l.map(get_digdirfg)
addrange.loc[addresses_r.index, "r_digdirfg"] = address_sequence_r.map(get_digdirfg)
```
## Step 5. Merge addrange attributes with roadseg
```
# Merge addrange attributes with roadseg.
roadseg = roadseg.merge(addrange, how="left", left_index=True, right_index=True)
```
## View Results
**Note:** this code block is for visual purposes only.
```
# Create data for viewing.
addresses_filtered = addresses.loc[addresses["roadseg_index"]==ex_idx]
labels = addresses_filtered[["number", "suffix", "geometry", "parity"]].apply(
lambda row: (f"{row[0]}{row[1]}", row[2].x, row[2].y, row[3]), axis=1)
# Configure plots.
fig, ax = plt.subplots(1, 1, figsize=(6, 7), tight_layout=False)
ax.imshow(basemaps[1][0], extent=basemaps[1][1])
addresses_filtered.loc[addresses_filtered["parity"]=="l"].plot(ax=ax, color="blue", label="addresses (left)", linewidth=2)
addresses_filtered.loc[addresses_filtered["parity"]=="r"].plot(ax=ax, color="lime", label="addresses(right)", linewidth=2)
starting_pt.plot(ax=ax, color="gold", label=f"roadseg={ex_idx}, 1st point", linewidth=2)
roadseg.loc[roadseg.index==ex_idx].plot(ax=ax, color="yellow", label=f"roadseg={ex_idx}", linewidth=2)
roadseg.loc[roadseg.index!=ex_idx].plot(ax=ax, color="cyan", label="roadseg", linewidth=1)
ax.add_artist(ScaleBar(dx=dx, units="m", location="lower left", pad=0.5, color="black"))
ax.axes.xaxis.set_visible(False)
ax.axes.yaxis.set_visible(False)
ax.set_title("Parity Output", fontsize=12)
ax.set_xlim(itemgetter(0, 1)(basemaps[1][1]))
ax.set_ylim(itemgetter(2, 3)(basemaps[1][1]))
for label_params in labels:
label, x, y, parity = label_params
if parity == "l":
kwargs = {"xytext": (x-10, y+10), "ha": "right"}
else:
kwargs = {"xytext": (x+10, y+10), "ha": "left"}
plt.annotate(label, xy=(x, y), textcoords="data", va="bottom", fontsize=10, color="red", fontweight="bold",
bbox=dict(pad=0.3, fc="black"), **kwargs)
plt.legend(loc="center left", bbox_to_anchor=(1.0, 0.5), fontsize=12)
plt.savefig("temp.png", bbox_inches='tight', pad_inches=0)
plt.close()
display_html(f"""
<div class=\"container\" style=\"width:100%;\">
<img src=\"temp.png\" style=\"float:left;max-width:59%;\">
{pd.DataFrame(roadseg.loc[roadseg.index==ex_idx].iloc[0]).style.set_table_styles([
{'selector': '', 'props': [('float', 'right'), ('width', '40%')]},
{'selector': 'td', 'props': [('overflow', 'hidden'), ('text-overflow', 'ellipsis'), ('white-space', 'nowrap')]}
])._repr_html_()}
</div>
""", raw=True)
```
| github_jupyter |
<img src="https://raw.githubusercontent.com/dask/dask/main/docs/source/images/dask_horizontal_no_pad.svg"
width="30%"
alt="Dask logo\" />
# Parallel and Distributed Machine Learning
The material in this notebook was based on the open-source content from [Dask's tutorial repository](https://github.com/dask/dask-tutorial) and the [Machine learning notebook](https://github.com/coiled/data-science-at-scale/blob/master/3-machine-learning.ipynb) from data science at scale from coiled
So far we have seen how Dask makes data analysis scalable with parallelization via Dask DataFrames. Let's now see how [Dask-ML](https://ml.dask.org/) allows us to do machine learning in a parallel and distributed manner. Note, machine learning is really just a special case of data analysis (one that automates analytical model building), so the 💪 Dask gains 💪 we've seen will apply here as well!
(If you'd like a refresher on the difference between parallel and distributed computing, [here's a good discussion on StackExchange](https://cs.stackexchange.com/questions/1580/distributed-vs-parallel-computing).)
## Types of scaling problems in machine learning
There are two main types of scaling challenges you can run into in your machine learning workflow: scaling the **size of your data** and scaling the **size of your model**. That is:
1. **CPU-bound problems**: Data fits in RAM, but training takes too long. Many hyperparameter combinations, a large ensemble of many models, etc.
2. **Memory-bound problems**: Data is larger than RAM, and sampling isn't an option.
Here's a handy diagram for visualizing these problems:
<img src="https://raw.githubusercontent.com/coiled/data-science-at-scale/master/images/dimensions_of_scale.svg"
width="60%"
alt="scaling problems\" />
In the bottom-left quadrant, your datasets are not too large (they fit comfortably in RAM) and your model is not too large either. When these conditions are met, you are much better off using something like scikit-learn, XGBoost, and similar libraries. You don't need to leverage multiple machines in a distributed manner with a library like Dask-ML. However, if you are in any of the other quadrants, distributed machine learning is the way to go.
Summarizing:
* For in-memory problems, just use scikit-learn (or your favorite ML library).
* For large models, use `dask_ml.joblib` and your favorite scikit-learn estimator.
* For large datasets, use `dask_ml` estimators.
## Scikit-learn in five minutes
<img src="https://raw.githubusercontent.com/coiled/data-science-at-scale/master/images/scikit_learn_logo_small.svg"
width="30%"
alt="sklearn logo\" />
In this section, we'll quickly run through a typical scikit-learn workflow:
* Load some data (in this case, we'll generate it)
* Import the scikit-learn module for our chosen ML algorithm
* Create an estimator for that algorithm and fit it with our data
* Inspect the learned attributes
* Check the accuracy of our model
Scikit-learn has a nice, consistent API:
* You instantiate an `Estimator` (e.g. `LinearRegression`, `RandomForestClassifier`, etc.). All of the models *hyperparameters* (user-specified parameters, not the ones learned by the estimator) are passed to the estimator when it's created.
* You call `estimator.fit(X, y)` to train the estimator.
* Use `estimator` to inspect attributes, make predictions, etc.
Here `X` is an array of *feature variables* (what you're using to predict) and `y` is an array of *target variables* (what we're trying to predict).
### Generate some random data
```
from sklearn.datasets import make_classification
# Generate data
X, y = make_classification(n_samples=10000, n_features=4, random_state=0)
```
**Refreshing some ML concepts**
- `X` is the samples matrix (or design matrix). The size of `X` is typically (`n_samples`, `n_features`), which means that samples are represented as rows and features are represented as columns.
- A "feature" (also called an "attribute") is a measurable property of the phenomenon we're trying to analyze. A feature for a dataset of employees might be their hire date, for example.
- `y` are the target values, which are real numbers for regression tasks, or integers for classification (or any other discrete set of values). For unsupervized learning tasks, `y` does not need to be specified. `y` is usually 1d array where the `i`th entry corresponds to the target of the `i`th sample (row) of `X`.
```
# Let's take a look at X
X[:8]
# Let's take a look at y
y[:8]
```
### Fitting and SVC
For this example, we will fit a [Support Vector Classifier](https://scikit-learn.org/stable/modules/generated/sklearn.svm.SVC.html).
```
from sklearn.svm import SVC
estimator = SVC(random_state=0)
estimator.fit(X, y)
```
We can inspect the learned features by taking a look a the `support_vectors_`:
```
estimator.support_vectors_[:4]
```
And we check the accuracy:
```
estimator.score(X, y)
```
There are [3 different approaches](https://scikit-learn.org/0.15/modules/model_evaluation.html) to evaluate the quality of predictions of a model. One of them is the **estimator score method**. Estimators have a score method providing a default evaluation criterion for the problem they are designed to solve, which is discussed in each estimator's documentation.
### Hyperparameter Optimization
There are a few ways to learn the best *hyper*parameters while training. One is `GridSearchCV`.
As the name implies, this does a brute-force search over a grid of hyperparameter combinations. scikit-learn provides tools to automatically find the best parameter combinations via cross-validation (which is the "CV" in `GridSearchCV`).
```
from sklearn.model_selection import GridSearchCV
%%time
estimator = SVC(gamma='auto', random_state=0, probability=True)
param_grid = {
'C': [0.001, 10.0],
'kernel': ['rbf', 'poly'],
}
# Brute-force search over a grid of hyperparameter combinations
grid_search = GridSearchCV(estimator, param_grid, verbose=2, cv=2)
grid_search.fit(X, y)
grid_search.best_params_, grid_search.best_score_
```
## Compute Bound: Single-machine parallelism with Joblib
<img src="https://raw.githubusercontent.com/coiled/data-science-at-scale/master/images/joblib_logo.svg"
alt="Joblib logo"
width="50%"/>
In this section we'll see how [Joblib](https://joblib.readthedocs.io/en/latest/) ("*a set of tools to provide lightweight pipelining in Python*") gives us parallelism on our laptop. Here's what our grid search graph would look like if we set up six training "jobs" in parallel:
<img src="https://raw.githubusercontent.com/coiled/data-science-at-scale/master/images/unmerged_grid_search_graph.svg"
alt="grid search graph"
width="100%"/>
With Joblib, we can say that scikit-learn has *single-machine* parallelism.
Any scikit-learn estimator that can operate in parallel exposes an `n_jobs` keyword, which tells you how many tasks to run in parallel. Specifying `n_jobs=-1` jobs means running the maximum possible number of tasks in parallel.
```
%%time
grid_search = GridSearchCV(estimator, param_grid, verbose=2, cv=2, n_jobs=-1)
grid_search.fit(X, y)
```
Notice that the computation above it is faster than before. If you are running this computation on binder, you might not see a speed-up and the reason for that is that binder instances tend to have only one core with no threads so you can't see any parallelism.
## Compute Bound: Multi-machine parallelism with Dask
In this section we'll see how Dask (plus Joblib and scikit-learn) gives us multi-machine parallelism. Here's what our grid search graph would look like if we allowed Dask to schedule our training "jobs" over multiple machines in our cluster:
<img src="https://raw.githubusercontent.com/coiled/data-science-at-scale/master/images/merged_grid_search_graph.svg"
alt="merged grid search graph"
width="100%"/>
We can say that Dask can talk to scikit-learn (via Joblib) so that our *cluster* is used to train a model.
If we run this on a laptop, it will take quite some time, but the CPU usage will be satisfyingly near 100% for the duration. To run faster, we would need a distributed cluster. For details on how to create a LocalCluster you can check the Dask documentation on [Single Machine: dask.distributed](https://docs.dask.org/en/latest/setup/single-distributed.html).
Let's instantiate a Client with `n_workers=4`, which will give us a `LocalCluster`.
```
import dask.distributed
client = dask.distributed.Client(n_workers=4)
client
```
**Note:** Click on Cluster Info, to see more details about the cluster. You can see the configuration of the cluster and some other specs.
We can expand our problem by specifying more hyperparameters before training, and see how using `dask` as backend can help us.
```
param_grid = {
'C': [0.001, 0.1, 1.0, 2.5, 5, 10.0],
'kernel': ['rbf', 'poly', 'linear'],
'shrinking': [True, False],
}
grid_search = GridSearchCV(estimator, param_grid, verbose=2, cv=2, n_jobs=-1)
```
### Dask parallel backend
We can fit our estimator with multi-machine parallelism by quickly *switching to a Dask parallel backend* when using joblib.
```
import joblib
%%time
with joblib.parallel_backend("dask", scatter=[X, y]):
grid_search.fit(X, y)
```
**What did just happen?**
Dask-ML developers worked with the scikit-learn and Joblib developers to implement a Dask parallel backend. So internally, scikit-learn now talks to Joblib, and Joblib talks to Dask, and Dask is what handles scheduling all of those tasks on multiple machines.
The best parameters and best score:
```
grid_search.best_params_, grid_search.best_score_
```
## Memory Bound: Single/Multi machine parallelism with Dask-ML
We have seen how to work with larger models, but sometimes you'll want to train on a larger than memory dataset. `dask-ml` has implemented estimators that work well on Dask `Arrays` and `DataFrames` that may be larger than your machine's RAM.
```
import dask.array as da
import dask.delayed
from sklearn.datasets import make_blobs
import numpy as np
```
We'll make a small (random) dataset locally using scikit-learn.
```
n_centers = 12
n_features = 20
X_small, y_small = make_blobs(n_samples=1000, centers=n_centers, n_features=n_features, random_state=0)
centers = np.zeros((n_centers, n_features))
for i in range(n_centers):
centers[i] = X_small[y_small == i].mean(0)
centers[:4]
```
**Note**: The small dataset will be the template for our large random dataset.
We'll use `dask.delayed` to adapt `sklearn.datasets.make_blobs`, so that the actual dataset is being generated on our workers.
If you are not in binder and you machine has 16GB of RAM you can make `n_samples_per_block=200_000` and the computations takes around 10 min. If you are in binder the resources are limited and the problem below is big enough.
```
n_samples_per_block = 60_000 #on binder replace this for 15_000
n_blocks = 500
delayeds = [dask.delayed(make_blobs)(n_samples=n_samples_per_block,
centers=centers,
n_features=n_features,
random_state=i)[0]
for i in range(n_blocks)]
arrays = [da.from_delayed(obj, shape=(n_samples_per_block, n_features), dtype=X.dtype)
for obj in delayeds]
X = da.concatenate(arrays)
X
```
### KMeans from Dask-ml
The algorithms implemented in Dask-ML are scalable. They handle larger-than-memory datasets just fine.
They follow the scikit-learn API, so if you're familiar with scikit-learn, you'll feel at home with Dask-ML.
```
from dask_ml.cluster import KMeans
clf = KMeans(init_max_iter=3, oversampling_factor=10)
%time clf.fit(X)
clf.labels_
clf.labels_[:10].compute()
client.close()
```
## Multi-machine parallelism in the cloud with Coiled
<br>
<img src="https://raw.githubusercontent.com/coiled/data-science-at-scale/master/images/Coiled-Logo_Horizontal_RGB_Black.png"
alt="Coiled logo"
width=25%/>
<br>
In this section we'll see how Coiled allows us to solve machine learning problems with multi-machine parallelism in the cloud.
Coiled, [among other things](https://coiled.io/product/), provides hosted and scalable Dask clusters. The biggest barriers to entry for doing machine learning at scale are "Do you have access to a cluster?" and "Do you know how to manage it?" Coiled solves both of those problems.
We'll spin up a Coiled cluster (with 10 workers in this case), then instantiate a Dask Client to use with that cluster.
If you are running on your local machine and not in binder, and you want to give Coiled a try, you can signup [here](https://cloud.coiled.io/login?redirect_uri=/) and you will get some free credits. If you installed the environment by following the steps on the repository's [README](https://github.com/coiled/dask-mini-tutorial/blob/main/README.md) you will have `coiled` installed. You will just need to login, by following the steps on the [setup page](https://docs.coiled.io/user_guide/getting_started.html), and you will be ready to go.
To learn more about how to set up an environment you can visit Coiled documentation on [Creating software environments](https://docs.coiled.io/user_guide/software_environment_creation.html). But for now you can use the envioronment we set up for this tutorial.
```
import coiled
from dask.distributed import Client
# Spin up a Coiled cluster, instantiate a Client
cluster = coiled.Cluster(n_workers=10, software="ncclementi/dask-mini-tutorial",)
client = Client(cluster)
client
```
### Memory bound: Dask-ML
We can use Dask-ML estimators on the cloud to work with larger datasets.
```
n_centers = 12
n_features = 20
X_small, y_small = make_blobs(n_samples=1000, centers=n_centers, n_features=n_features, random_state=0)
centers = np.zeros((n_centers, n_features))
for i in range(n_centers):
centers[i] = X_small[y_small == i].mean(0)
n_samples_per_block = 200_000
n_blocks = 500
delayeds = [dask.delayed(make_blobs)(n_samples=n_samples_per_block,
centers=centers,
n_features=n_features,
random_state=i)[0]
for i in range(n_blocks)]
arrays = [da.from_delayed(obj, shape=(n_samples_per_block, n_features), dtype=X.dtype)
for obj in delayeds]
X = da.concatenate(arrays)
X = X.persist()
from dask_ml.cluster import KMeans
clf = KMeans(init_max_iter=3, oversampling_factor=10)
%time clf.fit(X)
```
Computing the labels:
```
clf.labels_[:10].compute()
client.close()
```
## Extra resources:
- [Dask-ML documentation](https://ml.dask.org/)
- [Getting started with Coiled](https://docs.coiled.io/user_guide/getting_started.html)
| github_jupyter |
## Нейронные сети: зависимость ошибки и обучающей способности от числа нейронов
В этом задании вы будете настраивать двуслойную нейронную сеть для решения задачи многоклассовой классификации. Предлагается выполнить процедуры загрузки и разбиения входных данных, обучения сети и подсчета ошибки классификации. Предлагается определить оптимальное количество нейронов в скрытом слое сети. Нужно так подобрать число нейронов, чтобы модель была с одной стороны несложной, а с другой стороны давала бы достаточно точный прогноз и не переобучалась. Цель задания -- показать, как зависит точность и обучающая способность сети от ее сложности.
Для решения задачи многоклассовой классификации предлагается воспользоваться библиотекой построения нейронных сетей [pybrain](http://pybrain.org/). Библиотека содержит основные модули инициализации двуслойной нейронной сети прямого распространения, оценки ее параметров с помощью метода обратного распространения ошибки (backpropagation) и подсчета ошибки.
Установить библиотеку pybrain можно с помощью стандартной системы управления пакетами pip:
```
pip install pybrain
```
Кроме того, для установки библиотеки можно использовать и другие способы, приведенные в [документации](https://github.com/pybrain/pybrain/wiki/installation).
### Используемые данные
Рассматривается задача оценки качества вина по его физико-химическим свойствам [1]. Данные размещены в [открытом доступе](https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv) в репозитории UCI и содержат 1599 образцов красного вина, описанных 11 признаками, среди которых -- кислотность, процентное содержание сахара, алкоголя и пр. Кроме того, каждому объекту поставлена в соответствие оценка качества по шкале от 0 до 10. Требуется восстановить оценку качества вина по исходному признаковому описанию.
[1] P. Cortez, A. Cerdeira, F. Almeida, T. Matos and J. Reis. Modeling wine preferences by data mining from physicochemical properties. In Decision Support Systems, Elsevier, 47(4):547-553, 2009.
```
# Выполним инициализацию основных используемых модулей
%matplotlib inline
import random
import matplotlib.pyplot as plt
from sklearn.preprocessing import normalize
import numpy as np
```
Выполним загрузку данных
```
with open('winequality-red.csv') as f:
f.readline() # пропуск заголовочной строки
data = np.loadtxt(f, delimiter=';')
```
В качестве альтернативного варианта, можно выполнить загрузку данных напрямую из репозитория UCI, воспользовавшись библиотекой urllib.
```
import urllib
# URL for the Wine Quality Data Set (UCI Machine Learning Repository)
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv"
# загрузка файла
f = urllib.urlopen(url)
f.readline() # пропуск заголовочной строки
data = np.loadtxt(f, delimiter=';')
```
Выделим из данных целевую переменную. Классы в задаче являются несбалинсированными: основной доле объектов поставлена оценка качества от 5 до 7. Приведем задачу к трехклассовой: объектам с оценкой качества меньше пяти поставим оценку 5, а объектам с оценкой качества больше семи поставим 7.
```
TRAIN_SIZE = 0.7 # Разделение данных на обучающую и контрольную части в пропорции 70/30%
from sklearn.cross_validation import train_test_split
y = data[:, -1]
np.place(y, y < 5, 5)
np.place(y, y > 7, 7)
y -= min(y)
X = data[:, :-1]
X = normalize(X)
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=TRAIN_SIZE, random_state=0)
```
### Двуслойная нейронная сеть
Двуслойная нейронная сеть представляет собой функцию распознавания, которая може быть записана в виде следующей суперпозиции:
$f(x,W)=h^{(2)}\left(\sum\limits_{i=1}^D w_i^{(2)}h^{(1)}\left(\sum\limits_{j=1}^n w_{ji}^{(1)}x_j+b_i^{(1)}\right)+b^{(2)}\right)$, где
$x$ -- исходный объект (сорт вина, описанный 11 признаками), $x_j$ -- соответствующий признак,
$n$ -- количество нейронов во входном слое сети, совпадающее с количеством признаков,
$D$ -- количество нейронов в скрытом слое сети,
$w_i^{(2)}, w_{ji}^{(1)}, b_i^{(1)}, b^{(2)}$ -- параметры сети, соответствующие весам нейронов,
$h^{(1)}, h^{(2)}$ -- функции активации.
В качестве функции активации на скрытом слое сети используется линейная функция. На выходном слое сети используется функция активации softmax, являющаяся обобщением сигмоидной функции на многоклассовый случай:
$y_k=\text{softmax}_k(a_1,...,a_k)=\frac{\exp(a_k)}{\sum_{k=1}^K\exp(a_k)}.$
### Настройка параметров сети
Оптимальные параметры сети $W_{opt}$ определяются путем минимизации функции ошибки:
$W_{opt}=\arg\min\limits_{W}L(W)+\lambda\|W\|^2$.
Здесь $L(W)$ является функцией ошибки многоклассовой классификации,
$L(W)=- \sum^N_{n=1}\sum^K_{k=1} t_{kn} log(y_{kn}),$
$t_{kn}$ -- бинарно закодированные метки классов, $K$ -- количество меток, $N$ -- количество объектов,
а $\lambda\|W\|^2$ является регуляризующим слагаемым, контролирующим суммарный вес параметров сети и предотвращающий эффект переобучения.
Оптимизация параметров выполняется методом обратного распространения ошибки (backpropagation).
Выполним загрузку основных модулей: ClassificationDataSet -- структура данных pybrain, buildNetwork -- инициализация нейронной сети, BackpropTrainer -- оптимизация параметров сети методом backpropagation, SoftmaxLayer -- функция softmax, соответствующая выходному слою сети, percentError -- функцию подсчета ошибки классификации (доля неправильных ответов).
```
from pybrain.datasets import ClassificationDataSet # Структура данных pybrain
from pybrain.tools.shortcuts import buildNetwork
from pybrain.supervised.trainers import BackpropTrainer
from pybrain.structure.modules import SoftmaxLayer
from pybrain.utilities import percentError
```
Инициализируем основные параметры задачи: HIDDEN_NEURONS_NUM -- количество нейронов скрытого слоя, MAX_EPOCHS -- максимальное количество итераций алгоритма оптимизации
```
# Определение основных констант
HIDDEN_NEURONS_NUM = 100 # Количество нейронов, содержащееся в скрытом слое сети
MAX_EPOCHS = 100 # Максимальное число итераций алгоритма оптимизации параметров сети
```
Инициализируем структуру данных ClassificationDataSet, используемую библиотекой pybrain. Для инициализации структура принимает два аргумента: количество признаков *np.shape(X)[1]* и количество различных меток классов *len(np.unique(y))*.
Кроме того, произведем бинаризацию целевой переменной с помощью функции *_convertToOneOfMany( )* и разбиение данных на обучающую и контрольную части.
```
# Конвертация данных в структуру ClassificationDataSet
# Обучающая часть
ds_train = ClassificationDataSet(np.shape(X)[1], nb_classes=len(np.unique(y_train)))
# Первый аргумент -- количество признаков np.shape(X)[1], второй аргумент -- количество меток классов len(np.unique(y_train)))
ds_train.setField('input', X_train) # Инициализация объектов
ds_train.setField('target', y_train[:, np.newaxis]) # Инициализация ответов; np.newaxis создает вектор-столбец
ds_train._convertToOneOfMany( ) # Бинаризация вектора ответов
# Контрольная часть
ds_test = ClassificationDataSet(np.shape(X)[1], nb_classes=len(np.unique(y_train)))
ds_test.setField('input', X_test)
ds_test.setField('target', y_test[:, np.newaxis])
ds_test._convertToOneOfMany( )
```
Инициализируем двуслойную сеть и произведем оптимизацию ее параметров. Аргументами для инициализации являются:
ds.indim -- количество нейронов на входном слое сети, совпадает с количеством признаков (в нашем случае 11),
HIDDEN_NEURONS_NUM -- количество нейронов в скрытом слое сети,
ds.outdim -- количество нейронов на выходном слое сети, совпадает с количеством различных меток классов (в нашем случае 3),
SoftmaxLayer -- функция softmax, используемая на выходном слое для решения задачи многоклассовой классификации.
```
np.random.seed(0) # Зафиксируем seed для получения воспроизводимого результата
# Построение сети прямого распространения (Feedforward network)
net = buildNetwork(ds_train.indim, HIDDEN_NEURONS_NUM, ds_train.outdim, outclass=SoftmaxLayer)
# ds.indim -- количество нейронов входного слоя, равне количеству признаков
# ds.outdim -- количество нейронов выходного слоя, равное количеству меток классов
# SoftmaxLayer -- функция активации, пригодная для решения задачи многоклассовой классификации
init_params = np.random.random((len(net.params))) # Инициализируем веса сети для получения воспроизводимого результата
net._setParameters(init_params)
```
Выполним оптимизацию параметров сети. График ниже показывает сходимость функции ошибки на обучающей/контрольной части.
```
random.seed(0)
# Модуль настройки параметров pybrain использует модуль random; зафиксируем seed для получения воспроизводимого результата
trainer = BackpropTrainer(net, dataset=ds_train) # Инициализируем модуль оптимизации
err_train, err_val = trainer.trainUntilConvergence(maxEpochs=MAX_EPOCHS)
line_train = plt.plot(err_train, 'b', err_val, 'r') # Построение графика
xlab = plt.xlabel('Iterations')
ylab = plt.ylabel('Error')
```
Рассчитаем значение доли неправильных ответов на обучающей и контрольной выборке.
```
res_train = net.activateOnDataset(ds_train).argmax(axis=1) # Подсчет результата на обучающей выборке
print 'Error on train: ', percentError(res_train, ds_train['target'].argmax(axis=1)), '%' # Подсчет ошибки
res_test = net.activateOnDataset(ds_test).argmax(axis=1) # Подсчет результата на тестовой выборке
print 'Error on test: ', percentError(res_test, ds_test['target'].argmax(axis=1)), '%' # Подсчет ошибки
```
### Задание. Определение оптимального числа нейронов.
В задании требуется исследовать зависимость ошибки на контрольной выборке в зависимости от числа нейронов в скрытом слое сети. Количество нейронов, по которому предполагается провести перебор, записано в векторе
```
hidden_neurons_num = [50, 100, 200, 500, 700, 1000]
```
1. Для фиксированного разбиения на обучающую и контрольную части подсчитайте долю неправильных ответов (ошибок) классификации на обучении/контроле в зависимости от количества нейронов в скрытом слое сети. Запишите результаты в массивы ```res_train_vec``` и ```res_test_vec```, соответственно. С помощью функции ```plot_classification_error``` постройте график зависимости ошибок на обучении/контроле от количества нейронов. Являются ли графики ошибок возрастающими/убывающими? При каком количестве нейронов достигается минимум ошибок классификации?
2. С помощью функции ```write_answer_nn``` запишите в выходной файл число: количество нейронов в скрытом слое сети, для которого достигается минимум ошибки классификации на контрольной выборке.
```
random.seed(0) # Зафиксируем seed для получния воспроизводимого результата
np.random.seed(0)
def plot_classification_error(hidden_neurons_num, res_train_vec, res_test_vec):
# hidden_neurons_num -- массив размера h, содержащий количество нейронов, по которому предполагается провести перебор,
# hidden_neurons_num = [50, 100, 200, 500, 700, 1000];
# res_train_vec -- массив размера h, содержащий значения доли неправильных ответов классификации на обучении;
# res_train_vec -- массив размера h, содержащий значения доли неправильных ответов классификации на контроле
plt.figure()
plt.plot(hidden_neurons_num, res_train_vec)
plt.plot(hidden_neurons_num, res_test_vec, '-r')
def write_answer_nn(optimal_neurons_num):
with open("nnets_answer1.txt", "w") as fout:
fout.write(str(optimal_neurons_num))
hidden_neurons_num = [50, 100, 200, 500, 700, 1000]
res_train_vec = list()
res_test_vec = list()
for nnum in hidden_neurons_num:
# Put your code here
# Не забудьте про инициализацию весов командой np.random.random((len(net.params)))
# Постройте график зависимости ошибок на обучении и контроле в зависимости от количества нейронов
plot_classification_error(hidden_neurons_num, res_train_vec, res_test_vec)
# Запишите в файл количество нейронов, при котором достигается минимум ошибки на контроле
write_answer_nn(hidden_neurons_num[res_test_vec.index(min(res_test_vec))])
```
| github_jupyter |
# 초보자를 위한 파이썬 300제
* source : https://wikidocs.net/book/922
## 파이썬 시작하기
* source : https://wikidocs.net/7014
### print 텝과 줄바꿈
* exam : 005, 009
```
print("안녕하세요.\n만나서\t\t반갑습니다.") # \t 텝, \n 줄바꿈 의미
```
```
print("first",end="");print("second") # end='' print 함수는 두 번 사용한다. 세미콜론 (;)은 한줄에 여러 개의 명령을 작성하기 위해 사용한다.
```
### print 기초
* exam : 008, 004
```
print("naver", "kakao", "sk", "samsung", sep="/")
```
```
print('"C:\Windows"')
print('("C:\Windows")')
```
## 파이썬 변수
* source : https://wikidocs.net/7021
### type함수
* exam : 012, 015
```
시가총액 = 298000000000000
현재가 = 50000
PER = 15.79
print(시가총액, type(시가총액))
print(현재가, type(현재가))
print(PER, type(PER))
```
```
a = 128
print(type(a))
```
```
a = "132"
print(type(a))
```
### 문자열, 정수, 실수 변환
* exam : 016, 017, 018, 019
```
num_str = "720"
num_int = int(num_str) # 문자열을 정수로 변환
print(num_int, type(num_int))
```
```
num=100
result=str(num) # 정수를 문자열 100으로 변환
print(result, type(result))
```
```
a = "15.79"
b = float(a) # 문자열을 실수로 변환
print(b,type(b))
```
```
year = "2020"
print(int(year)-3) #문자열을 정수로 변환
print(int(year)-2)
print(int(year)-1)
```
## 파이썬 문자열
*source : https://wikidocs.net/7022
### 문자열 슬라이싱
* exam : 022, 023, 024, 039
```
license_plate = "24rk 2210"
print(license_plate[-4:]) # 음수 값은 문자열의 뒤에서부터 인덱싱 또는 슬라이싱함을 의미한다.
```
```
string = "홀짝홀짝홀짝"
print(string[::2]) # 슬라이싱할 떄 시작인덱스:끝인덱스:오프셋 을 지정할 수 있습니다.
```
```
string = "PYTHON"
print(string[::-1]) # -1은 끝에서부터 라는 뜻...
```
```
분기 = "2020/03(E) (IFRS연결)"
print(분기[:7])
```
### 문자열 인덱싱
* exam : 021
```
letter = 'python'
print(letter[0], letter[2])
```
### 문자열 다루기
*exam : 025, 026, 027
```
phone_number = "010-1111-2222"
phone_number = phone_number.replace("-"," ")
print(phone_number)
```
```
phone_number = "010-1111-2222"
phone_number = phone_number.replace("-","")
print(phone_number)
```
```
url = "http://sharebook.kr"
url_split = url.split('.') # .을 기준으로 분리한다. 분리된 url 중 마지막을 인덱싱 하면 도메인만 출력 가능
print(url_split[-1])
```
### 문자열은 immutable (불변)
* exam : 028
```
lang = 'python'
lang[0] = 'P' #문자열은 수정할 수 없다. 실행 결과를 확인해보면 문자열이 할당(assignment) 메서드를 지원하지 않음을 알 수 있다.
print(lang)
```
### replace 메서드
* exam : 029, 030
```
string = 'abcdfe2a354a32a'
string_replace = string.replace('a','A')
print(string_replace)
```
```
string='abcd'
string.replace('b','B')
print(string) # 문자열은 변경할 수 없는 자료형 이기 때문에 replace 매서드를 사용하면 원본은 그대로 둔채로 변경된 새로운 문자열 객체를 리턴해준다.
```
### Format String
* exam : 035, 036, 037
```
#bad
name = 'python'
#print('이름: ' + name + '나이: ' +3 ) --err! print('이름: ' + name + '나이: ' +str(3) ) -- ok
print('이름: ' , name ,'나이: ' , 3 )
```
```
#good (이 형태 평소에도 많이 씀)
print("이름: %s 나이: %d " % (name, 3)) # %s는 문자열 데이터 타입의 값을, %d는 정수형 데이터 타입 값의 출력을 의미
```
```
name1 = "김민수"
age1 = 10
name2 = "이철희"
age2 = 13
print("이름: {} 나이: {}".format(name1, age1)) # 문자열의 포맷 메서드는 타입과 상관없이 값이 출력될 위치에 {}를 적어주면 된다.
print("이름: {} 나이: {}".format(name2, age2))
```
```
print(f"이름: {name1} 나이: {age1}") # f-string은 문자열 앞에 f가 붙은 형태이다. f-string을 사용하면 {변수}와 같은 형탤호 문자열 사이에 타입과 상관없이 값을 출력 가능
print(f"이름: {name2} 나이: {age2}")
```
### strip 메서드
* exam : 040, 050
```
data = " 삼성전자 "
data1 = data.strip() #문자열에서 strip() 메서드를 사용하면 좌우의 공백을 제거할 수 있다. 이때 원본 문자열은 그대로 유지, 공백이 제거된 새로운 문자열 반환
print(data1)
```
```
data = "039490 " # rstrip() 메서드를 사용하면 오른쪽 고백이 제거된 새로운 문자열 객체가 반환
data = data.rstrip()
```
### split 메서드
* exam : 047, 048, 049
```
a = "hello world" # 문자열의 split() 메서드를 사용하면 문자열에서 공백을 기준으로 분리해준다.
a.split()
```
```
ticker = "btc_krw" # 문자열에서 split() 메서드는 문자열을 분리할 때 사용. 이때 어떤 값을 넘겨주면 그 값을 기준으로 문자열을 분리
ticker.split("_")
```
```
date = "2020-05-01"
date.split("-")
```
## 파이썬 리스트
* source : https://wikidocs.net/7023
### 리스트 생성
* exam : 051
```
movie_rank = ["닥터 스트레인지", "스플릿", "럭키"]
```
### 리스트 추가
* exam : 052, 053
```
movie_rank.append("배트맨")
print(movie_rank)
```
```
movie_rank = ["닥터 스트레인지", "스플릿", "럭키", " 배트맨"]
movie_rank.insert(1, "슈퍼맨") # 리스트의 insert(인덱스,원소) 매서드를 사용하면 특정 위치에 값을 끼워넣기 할 수 있습니다.
print(movie_rank)
```
### 리스트의 최대값, 최소값
* exam : 057
```
nums = [1, 2, 3, 4, 5, 6, 7]
print("max: ", max(nums))
print("min: ", min(nums))
```
### 리스트에 저장된 데이터의 갯수
* exam : 059
```
cook = ["피자", "김밥", "만두", "양념치킨", "족발", "피자", "김치만두", "쫄면", "쏘세지", "라면", "팥빙수", "김치전"]
print(len(cook)) # len은 리스트에 저장된 데이터의 갯수를 구하는 함수이다.
```
### 리스트의 평균
* exam : 060
```
nums = [1, 2, 3, 4, 5]
average = sum(nums) / len(nums) # 리스트의 합을 리스트의 갯수로 나눠 평균을 구함
print(average)
```
### join 메서드
* exam : 066, 067, 068
```
interest = ['삼성전자', 'LG전자', 'Naver', 'SK하이닉스', '미래에셋대우']
print(" ".join(interest)) # 한칸 띄우기
```
```
print("/".join(interest)) # 중간에 / 넣기
```
```
print("\n".join(interest)) # 줄바꿈하기
```
### 리스트 정렬
* exam : 070
```
data = [2, 4, 3, 1, 5, 10, 9]
data.sort() #sort 를 사용하면 오름차순 정렬가능
print(data)
```
## 파이썬 터플
* source : https://wikidocs.net/7027
```
my_variable = ()
print(type(my_variable))
```
### 터플 오류
* 074
```
t = (1, 2, 3)
t[0] = 'a' # 터플은 원소의 값을 변경할 수 없다.
```
### 터플 -> 리스트 변환
* exam : 077
```
interest = ('삼성전자', 'LG전자', 'SK Hynix')
data = list(interest)
print(data)
```
### 리스트 -> 터플 변환
* exam : 078
```
interest = ['삼성전자', 'LG전자', 'SK Hynix']
data = tuple(interest)
print(data)
```
### range 함수
* exam : 080
```
# 1~99까지의 정수 중 짝수만 저장된 튜플을 생성하라
data = tuple(range(2, 100, 2))
print( data )
```
## 파이썬 딕셔너리
* source : https://wikidocs.net/22000
```
temp = { }
print(type(temp))
```
###딕셔너리 추가
*exam : 085, 086
```
ice = {"메로나": 1000, "폴라포": 1200, "빵빠레": 1800}
print(ice)
```
```
# 조스바 1200, 월드콘 1500 추가
ice = {"메로나": 1000, "폴라포": 1200, "빵빠레": 1800}
ice["죠스바"] = 1200
ice["월드콘"] = 1500
print(ice)
```
### 딕셔너리 사용하여 가격 출력, 수정, 삭제
* exam : 087, 088, 089
```
# 가격출력
ice = {'메로나': 1000,
'폴로포': 1200,
'빵빠레': 1800,
'죠스바': 1200,
'월드콘': 1500}
print("메로나 가격: ", ice["메로나"])
```
```
# 가격수정
ice = {'메로나': 1000,
'폴로포': 1200,
'빵빠레': 1800,
'죠스바': 1200,
'월드콘': 1500}
ice["메로나"] = 1300
print(ice)
```
```
# 메로나 삭제
ice = {'메로나': 1000,
'폴로포': 1200,
'빵빠레': 1800,
'죠스바': 1200,
'월드콘': 1500}
del ice["메로나"]
print(ice)
```
### 딕셔너리 생성 및 인덱싱
*exam : 091, 092, 093, 094
```
# 아이스크림 이름을 키값으로, (가격, 재고) 리스트를 딕셔너리의 값으로 저장
inventory = {"메로나": [300, 20],
"비비빅": [400, 3],
"죠스바": [250, 100]}
print(inventory)
```
```
# 가격확인
print(inventory["메로나"][0], "원")
```
```
# 재고확인
print(inventory["메로나"][1], "개")
```
```
# 딕셔너리 추가
inventory = {"메로나": [300, 20],
"비비빅": [400, 3],
"죠스바": [250, 100]}
inventory["월드콘"] = [500, 7]
print(inventory)
```
### 딕셔너리 keys()매서드
* exam 095
```
# 다음의 딕셔너리로부터 key 값으로만 구성된 리스트를 생성하라
icecream = {'탱크보이': 1200, '폴라포': 1200, '빵빠레': 1800, '월드콘': 1500, '메로나': 1000}
ice = list(icecream.keys())
print(ice)
```
## 파이썬 분기문
* source : https://wikidocs.net/7028
```
if 4 < 3:
print("Hello World.")
else:
print("Hi, there.")
```
```
# 사용자로부터 값을 입력받은 후 해당 값에 20을 더한 값을 출력하라.
# 단 사용자가 입력한 값과 20을 더한 계산 값이 255를 초과하는 경우 255를 출력해야 한다.
# 입력값 : 200, 출력값 : 220
# 입력값 : 240, 출력값 : 255
user = input("입력값: ")
num = 20 + int(user)
if num > 255:
print(255)
else:
print(num)
```
```
#사용자로부터 하나의 값을 입력받은 후 해당 값에 20을 뺀 값을 출력하라.
# 단 출력 값의 범위는 0~255이다.
# 예를 들어 결괏값이 0보다 작은 값이되는 경우 0을 출력하고 255보다 큰 값이 되는 경우 255를 출력해야 한다.
user = input("입력값: ")
num = int(user) - 20
if num > 255:
print(255)
elif num < 0:
print(0)
else:
print(num)
```
```
# 아래와 같이 fruit 딕셔너리가 정의되어 있다.
# 사용자가 입력한 값이 딕셔너리 키 (key) 값에 포함되었다면 "정답입니다"를 아닐 경우 "오답입니다" 출력하라.
# >> 제가좋아하는계절은: 봄
# 정답입니다.
fruit = {"봄" : "딸기", "여름" : "토마토", "가을" : "사과"}
user = input ("제가가장좋아하는계절은: ")
if user in fruit:
print("정답입니다.")
else:
print("오답입니다.")
```
```
# 아래와 같이 fruit 딕셔너리가 정의되어 있다.
# 사용자가 입력한 값이 딕셔너리 값 (value)에 포함되었다면 "정답입니다"를 아닐 경우 "오답입니다" 출력하라.
# >> 좋아하는과일은? 한라봉
# 오답입니다.
fruit = {"봄" : "딸기", "여름" : "토마토", "가을" : "사과"}
user = input("좋아하는 과일은?")
if user in fruit.values():
print("정답입니다.")
else:
print("오답입니다.")
```
### 짝수/홀수 판별
* exam : 113
```
user = input("")
if int(user) % 2 == 0: # 전체를 2로 나누었을때 나머지가 0이면 짝수, 아니면 홀수 이기 때문이다.
print("짝수")
else:
print("홀수")
```
### 대문자 소문자 판별 및 변환
* exam : 121
```
user = input("")
if user.islower(): #islower() 함수는 문자의 소문자 여부를 판별
print(user.upper()) #upper() 함수는 대문자로, lower() 함수는 소문자로 변경
else:
print(user.lower())
```
## 파이썬 반복문
* source : https://wikidocs.net/78562
```
# 저장된 문자열의 길이를 다음과 같이 출력하라
# 리스트 = ["SK하이닉스", "삼성전자", "LG전자"]
#6
#4
#4
리스트 = ["SK하이닉스", "삼성전자", "LG전자"]
for 종목명 in 리스트:
길이 = len(종목명)
print(길이)
```
```
리스트 = ['dog', 'cat', 'parrot']
for 이름 in 리스트:
print(이름, len(이름))
```
```
for 이름 in 리스트:
print(이름[0])
```
```
# for 문을 사용해서 반대로 출력하기
리스트 = ["가", "나", "다", "라"]
for 변수 in 리스트[::-1]:
print(변수)
```
### for문을 사용해서 리스트의 음수 출력
* exam : 151
```
리스트 = [3, -20, -3, 44]
for 변수 in 리스트:
if 변수 < 0:
print(변수)
```
### for 문을 사용해서 특정 배수 출력
* exam : 152, 153
```
# 3의 배수만 출력
리스트 = [3, 100, 23, 44]
for 변수 in 리스트:
if 변수 % 3 == 0:
print(변수)
```
```
#리스트에서 20 보다 작은 3의 배수를 출력하라
리스트 = [13, 21, 12, 14, 30, 18]
for 변수 in 리스트:
if 변수 % 3 == 0:
if 변수 < 20:
print(변수)
```
```
리스트 = [13, 21, 12, 14, 30, 18]
for 변수 in 리스트:
if (변수 < 20) and (변수 %3 ==0):
print(변수)
```
### for문 사용해서 대문자 판별
* exam : 155
```
리스트 = ["A", "b", "c", "D"]
for 변수 in 리스트:
if 변수.isupper(): #isupper()은 대문자 판별, islower()은 소문자 판별
print(변수)
```
```
리스트 = ["A", "b", "c", "D"]
for 변수 in 리스트:
if not 변수.isupper(): #대문자 판별 함수 앞에 not 를 사용 가능
print(변수)
```
```
# 파일 이름이 저장된 리스트에서 확장자가 .h 인 파일 이름을 출력하라.
리스트 = ['intra.h', 'intra.c', 'define.h', 'run.py']
for 변수 in 리스트:
split = 변수.split(".")
if split[1] == "h":
print(변수)
```
```
# 파일 이름이 저장된 리스트에서 확장자가 .h나 .c인 파일을 화면에 출력하라.
리스트 = ['intra.h', 'intra.c', 'define.h', 'run.py']
for 변수 in 리스트:
split = 변수.split(".")
if (split[1] == "h") or (split[1] == "c"):
print(변수)
```
### range 함수
* exam : 162, 164
```
# 월드컵은 4년에 한 번 개최된다. range()를 사용하여 2002~2050년까지 중 월드컵이 개최되는 연도를 출력하라
# 참고 : range 3번째 파라미터는 증감폭을 결정합니다.
print (list(range(2002, 2051, 4)))
```
```
for x in range(2002, 2051, 4):
print(x)
```
```
# 99부터 0까지 1씩 감소하는 숫자들을, 한 라인에 하나씩 출력하라.
for i in range(100):
print(99-i)
```
```
# 구구단 3단을 출력하라.
for i in range(1,10):
print(3, "x", i, "=", 3 * i)
```
```
# 구구단 3단을 출력하라. 단 홀수 번째만 출력한다.
num = 3
for i in range(1, 10, 2) : # range 함수는 세번 째 파라미터는 증감폭을 결정한다.
print (num, "x", i, " = ", num * i)
```
## 파이썬 함수
* source : https://wikidocs.net/23906
```
# 두 개의 숫자를 입력받아 합/차/곱/나눗셈을 출력하는 print_arithmetic_operation 함수를 작성하라.
# print_arithmetic_operation(3, 4)
# 3 + 4 = 7
# 3 - 4 = -1
# 3 * 4 = 12
# 3 / 4 = 0.75
print_arithmetic_operation(3, 4)
def print_arithmetic_operation(a, b):
print(a, "+", b, "=", a + b )
print(a, "-", b, "=", a - b )
print(a, "*", b, "=", a * b )
print(a, "/", b, "=", a / b )
```
```
# 하나의 리스트를 입력받아 짝수만 화면에 출력하는 print_even 함수를 정의하라
print_even ([1, 3, 2, 10, 12, 11, 15])
def print_even(my_list):
for v in my_list:
if v % 2 == 0:
print(v)
```
```
def 함수(num):
return num+4
a = 함수(10)
b = 함수(a)
c = 함수(b)
print(c)
```
| github_jupyter |
### Feature Scaling
With any distance based machine learning model (regularized regression methods, neural networks, and now kmeans), you will want to scale your data.
If you have some features that are on completely different scales, this can greatly impact the clusters you get when using K-Means.
In this notebook, you will get to see this first hand. To begin, let's read in the necessary libraries.
```
import numpy as np
import pandas as pd
from sklearn.cluster import KMeans
import matplotlib.pyplot as plt
from sklearn import preprocessing as p
%matplotlib inline
plt.rcParams['figure.figsize'] = (16, 9)
import helpers2 as h
import tests as t
# Create the dataset for the notebook
data = h.simulate_data(200, 2, 4)
df = pd.DataFrame(data)
df.columns = ['height', 'weight']
df['height'] = np.abs(df['height']*100)
df['weight'] = df['weight'] + np.random.normal(50, 10, 200)
```
`1.` Next, take a look at the data to get familiar with it. The dataset has two columns, and it is stored in the **df** variable. It might be useful to get an idea of the spread in the current data, as well as a visual of the points.
```
#Take a look at the data
#use this cell if you would like as well
```
Now that we've got a dataset, let's look at some options for scaling the data. As well as how the data might be scaled. There are two very common types of feature scaling that we should discuss:
**I. MinMaxScaler**
In some cases it is useful to think of your data in terms of the percent they are as compared to the maximum value. In these cases, you will want to use **MinMaxScaler**.
**II. StandardScaler**
Another very popular type of scaling is to scale data so that it has mean 0 and variance 1. In these cases, you will want to use **StandardScaler**.
It is probably more appropriate with this data to use **StandardScaler**. However, to get practice with feature scaling methods in python, we will perform both.
`2.` First let's fit the **StandardScaler** transformation to this dataset. I will do this one so you can see how to apply preprocessing in sklearn.
```
df_ss = p.StandardScaler().fit_transform(df) # Fit and transform the data
df_ss = pd.DataFrame(df_ss) #create a dataframe
df_ss.columns = ['height', 'weight'] #add column names again
plt.scatter(df_ss['height'], df_ss['weight']); # create a plot
```
`3.` Now it's your turn. Try fitting the **MinMaxScaler** transformation to this dataset. You should be able to use the previous example to assist.
```
# fit and transform
#create a dataframe
#change the column names
#plot the data
```
`4.` Now let's take a look at how kmeans divides the dataset into different groups for each of the different scalings of the data. Did you end up with different clusters when the data was scaled differently?
```
def fit_kmeans(data, centers):
'''
INPUT:
data = the dataset you would like to fit kmeans to (dataframe)
centers = the number of centroids (int)
OUTPUT:
labels - the labels for each datapoint to which group it belongs (nparray)
'''
kmeans = KMeans(centers)
labels = kmeans.fit_predict(data)
return labels
labels = fit_kmeans(df, 10) #fit kmeans to get the labels
# Plot the original data with clusters
plt.scatter(df['height'], df['weight'], c=labels, cmap='Set1');
#plot each of the scaled datasets
#another plot of the other scaled dataset
```
Write your response here!
| github_jupyter |
[](https://colab.research.google.com/github/mahdimplus/DeepRetroMoco/blob/main/functions.ipynb)
```
pip install voxelmorph
```
### Requirement libraries.
```
import nibabel as nib
import os
import numpy as np
import random
from nibabel.affines import apply_affine
import time
import voxelmorph as vxm
import pandas as pd
import matplotlib.pyplot as plt
```
### Loading data in our interested shape (64*64)
```
"""
laod_m ==== load data and reshape it
load_with_head ==== loading the data, reshaping it, and creating an appropriate header file
"""
def load_m (file_path):
img = nib.load(file_path)
img_data = img.get_fdata()
if img.shape[0:2]!=(64,64):
img_data = img_data[23:87,23:87,:,:]
if not (file_path.endswith(".nii") or file_path.endswith(".nii.gz")):
raise ValueError(
f"Nifti file path must end with .nii or .nii.gz, got {file_path}."
)
return img_data
def load_with_head (file_path: str):
img = nib.load(file_path)
img_data = img.get_fdata()
if img.shape[0:2]!=(64,64):
img_data = img_data[23:87,23:87,:,:]
header=img.header
## edit the header for shape
header['dim'][1:5]=img_data.shape
if not (file_path.endswith(".nii") or file_path.endswith(".nii.gz")):
raise ValueError(
f"Nifti file path must end with .nii or .nii.gz, got {file_path}."
)
return img_data ,img
```
### Listing data
```
"""
count ==== Listing data:
n= number of data
train_data_num= ID of the data
"""
def count (data_dir):
train_dir = os.path.join(data_dir)
train_data_num = []
for file in os.listdir(train_dir):
train_data_num.append([file])
train_data_num=np.array(train_data_num)
n=train_data_num.shape[0]
return n,train_data_num
```
### Maximum intensity
```
"""
maxx ==== finding maximum intensity among all data
"""
def maxx (data_dir):
n,train_data_num=count(data_dir)
start=0
for i in range(n):
d=load_m(data_dir+'/'+str(train_data_num[i][0]))
maxx=d.max()
if maxx>=start:
start=maxx
return start
```
### Prepare input (moved , fix) and ground truth (ref , deformation map) for training network
```
"""
Generator that takes in data of size [N, H, W], and yields data for
our model. Note that we need to provide numpy data for each
input, and each output.
inputs: moving [bs, H, W, 1], fixed image [bs, H, W, 1]
outputs: moved image [bs, H, W, 1], zero-gradient [bs, H, W, 2]
m= maximum intensity between all subject
split= percent of validation data
batch_size= number of data that take from subject.
### first selects a random subject and then random slice and following it finding random volume based on the number of batch_size ###
"""
def data_generator(data_dir, batch_size,m,split):
n,train_data_num=count(data_dir)
n_train=n-int(split*n)
subject_ID=random.randint(0,n_train-1)
d=load_m(data_dir+'/'+str(train_data_num[subject_ID][0]))
s=d.shape[2]
slice_ID =random.randint(0,s-1)
v=d.shape[3]
# preliminary sizing
vol_shape = d.shape[:2] # extract data shape
ndims = len(vol_shape)
d=d[:,:,slice_ID,:]
d = np.einsum('jki->ijk', d)
# prepare a zero array the size of the deformation
# we'll explain this below
zero_phi = np.zeros([batch_size, *vol_shape, ndims])
while True:
# prepare inputs:
# images need to be of the size [batch_size, H, W, 1]
idx1 = np.random.randint(0, v, size=batch_size)
moving_images = d[idx1, ..., np.newaxis]
moving_images=moving_images/m
idx2 = np.random.randint(0, v, size=batch_size)
fixed_images = d[idx2, ..., np.newaxis]
fixed_images=fixed_images/m
inputs = [moving_images, fixed_images]
# prepare outputs (the 'true' moved image):
# of course, we don't have this, but we know we want to compare
# the resulting moved image with the fixed image.
# we also wish to penalize the deformation field.
outputs = [fixed_images, zero_phi]
yield (inputs, outputs)
```
### prepare data for validation
```
"""
Generator that takes in data of size [N, H, W], and yields data for
our model. Note that we need to provide numpy data for each
input, and each output.
inputs: moving [bs, H, W, 1], fixed image [bs, H, W, 1]
outputs: moved image [bs, H, W, 1], zero-gradient [bs, H, W, 2]
m= maximum intensity between all subject
split= percent of validation data
batch_size= number of data that take from subject.
### first selects a random subject and then random slice and following it finding random volume based on the number of batch_size ###
"""
def val_generator(data_dir, batch_size,m,split):
n,train_data_num=count(data_dir)
n_train=n-int(split*n)
a=n_train
subject_ID=random.randint(a,n-1)
d=load_m(data_dir+'/'+str(train_data_num[subject_ID][0]))
s=d.shape[2]
slice_ID =random.randint(0,s-1)
v=d.shape[3]
# preliminary sizing
vol_shape = d.shape[:2] # extract data shape
ndims = len(vol_shape)
d=d[:,:,slice_ID,:]
d = np.einsum('jki->ijk', d)
# prepare a zero array the size of the deformation
# we'll explain this below
zero_phi = np.zeros([batch_size, *vol_shape, ndims])
# prepare inputs:
# images need to be of the size [batch_size, H, W, 1]
idx1 = np.random.randint(0, v, size=batch_size)
moving_images = d[idx1, ..., np.newaxis]
moving_images=moving_images/m
idx2 = np.random.randint(0, v, size=batch_size)
fixed_images = d[idx2, ..., np.newaxis]
fixed_images=fixed_images/m
inputs = [moving_images, fixed_images]
# prepare outputs (the 'true' moved image):
# of course, we don't have this, but we know we want to compare
# the resulting moved image with the fixed image.
# we also wish to penalize the deformation field.
outputs = [fixed_images,zero_phi]
return (inputs, outputs)
```
### Deformation matrix DOF=4
```
"""
This part creates the deformation matrix based on the:
12mm freedom through the y-direction,
6mm through the x-direction,
and 5 degrees to the left and right direction.
"""
def a(teta):
return (teta*np.pi)/180
def nearest_neighbors(i, j, M, T_inv):
x_max, y_max = M.shape[0] - 1, M.shape[1] - 1
x, y, k = apply_affine(T_inv, np.array([i, j, 1]))
if x<0 or y<0:
x=0
y=0
if x>=x_max+1 or y>=y_max+1:
x=0
y=0
if np.floor(x) == x and np.floor(y) == y:
x, y = int(x), int(y)
return M[x, y]
if np.abs(np.floor(x) - x) < np.abs(np.ceil(x) - x):
x = int(np.floor(x))
else:
x = int(np.ceil(x))
if np.abs(np.floor(y) - y) < np.abs(np.ceil(y) - y):
y = int(np.floor(y))
else:
y = int(np.ceil(y))
if x > x_max:
x = x_max
if y > y_max:
y = y_max
return M[x, y]
def affine_matrix():
t=random.randint(-5, 5)
cos_gamma = np.cos(a(t))
sin_gamma = np.sin(a(t))
x=random.randint(-3, 3)
y=random.randint(-6, 6)
T=np.array([[cos_gamma,-sin_gamma,0,x],
[sin_gamma,cos_gamma,0,y],
[0,0,1,0],
[0,0,0,1]])
return T
```
### Augmentation
```
"""
warps the images based on the deformation matrix
"""
def augsb(ref,volume,affine_matrix):
tdim,xdim,ydim,tdim = ref.shape
img_transformed = np.zeros((xdim, ydim), dtype=np.float64)
for i, row in enumerate(ref[volume,:,:,0]):
for j, col in enumerate(row):
pixel_data = ref[volume,i, j, 0]
input_coords = np.array([i, j, 1])
i_out, j_out,k= apply_affine(affine_matrix, input_coords)
if i_out<0 or j_out<0:
i_out=0
j_out=0
if i_out>=xdim or j_out>=ydim:
i_out=0
j_out=0
img_transformed[int(i_out),int(j_out)] = pixel_data
T_inv = np.linalg.inv(affine_matrix)
img_nn = np.ones((xdim, ydim), dtype=np.float64)
for i, row in enumerate(img_transformed):
for j, col in enumerate(row):
img_nn[i, j] = nearest_neighbors(i, j, ref[volume,:,:,0], T_inv)
return img_nn
```
### Prepare data for augmentation
```
def affine_generator(data_dir,batch_size,m,split):
n,train_data_num=count(data_dir)
n_train=n-int(split*n)
subject_ID=random.randint(0,n_train-1)
d=load_m(data_dir+'/'+str(train_data_num[subject_ID][0]))
s=d.shape[2]
slice_ID =random.randint(0,s-1)
v=d.shape[3]
# preliminary sizing
vol_shape = d.shape[:2] # extract data shape
ndims = len(vol_shape)
d=d[:,:,slice_ID,:]
d = np.einsum('jki->ijk', d)
y=[]
for i in range(batch_size):
y.append(affine_matrix())
y=np.array(y)
# prepare a zero array the size of the deformation
# we'll explain this below
zero_phi = np.zeros([batch_size, *vol_shape, ndims])
# prepare inputs:
# images need to be of the size [batch_size, H, W, 1]
while True:
idx2 = np.random.randint(0, v, size=batch_size)
fixed_images = d[idx2, ..., np.newaxis]
fixed_images=fixed_images/m
moving_images=[]
for i in range(batch_size):
moving_images.append(augsb(fixed_images,i,y[i]))
moving_images=np.array(moving_images)
moving_images=moving_images[... , np.newaxis]
#moving_images=augsb(fixed_images,y)
#idx1 = np.random.randint(0, v, size=batch_size)
#moving_images = d[idx1, ..., np.newaxis]
#moving_images=moving_images/m
inputs = [moving_images, fixed_images]
# prepare outputs (the 'true' moved image):
# of course, we don't have this, but we know we want to compare
# the resulting moved image with the fixed image.
# we also wish to penalize the deformation field.
outputs = [fixed_images,zero_phi]
yield(inputs,outputs)
#y)
```
### Prepare data for validation (based on augmentation data)
```
def label_generator(data_dir,batch_size,m,split):
n,train_data_num=count(data_dir)
n_train=n-int(split*n)
a=n_train
subject_ID=random.randint(a,n-1)
d=load_m(data_dir+'/'+str(train_data_num[subject_ID][0]))
s=d.shape[2]
slice_ID =random.randint(0,s-1)
v=d.shape[3]
# preliminary sizing
vol_shape = d.shape[:2] # extract data shape
ndims = len(vol_shape)
d=d[:,:,slice_ID,:]
d = np.einsum('jki->ijk', d)
y=[]
for i in range(batch_size):
y.append(affine_matrix())
y=np.array(y)
# prepare a zero array the size of the deformation
# we'll explain this below
# prepare inputs:
# images need to be of the size [batch_size, H, W, 1]
while True:
idx2 = np.random.randint(0, v, size=batch_size)
fixed_images = d[idx2, ...]
fixed_images=fixed_images/m
moving_images=[]
for i in range(batch_size):
moving_images.append(augsb(fixed_images,i,y[i]))
moving_images=np.array(moving_images)
#moving_images=moving_images[... ]
c=np.stack([moving_images,fixed_images], axis=2)
inputs = [c]
#inputs=[[moving_images,fixed_images]]
# prepare outputs (the 'true' moved image):
# of course, we don't have this, but we know we want to compare
# the resulting moved image with the fixed image.
# we also wish to penalize the deformation field.
outputs = [y]
yield (inputs, outputs)
```
#### **PREDICTION PHASE**
### Separating and gathering data based on the reference volume
```
"""
Generator that takes in data of size [N, H, W], and yields data for
our model. Note that we need to provide numpy data for each
input, and each output.
inputs: moving [bs, H, W, 1], fixed image [bs, H, W, 1]
outputs: moved image [bs, H, W, 1], zero-gradient [bs, H, W, 2]
m= maximum intensity between all subject
split= percent of validation data
batch_size= number of data that take from subject.
### It works with Slice and volume of data to box the data. ###
"""
def ref(data_dir,m,slice_ID,reference):
d=load_m(data_dir)
#s=d.shape[2]
#slice_ID =random.randint(0,s-1)
v=d.shape[3]
# preliminary sizing
vol_shape = d.shape[:2] # extract data shape
ndims = len(vol_shape)
d=d[:,:,slice_ID,:]
d = np.einsum('jki->ijk', d)
# prepare a zero array the size of the deformation
# we'll explain this below
zero_phi = np.zeros([v, *vol_shape, ndims])
# prepare inputs:
# images need to be of the size [batch_size, H, W, 1]
idx1=[]
for i in range(v):
idx1.append(i)
idx1=np.array(idx1)
moving_images = d[idx1, ..., np.newaxis]
moving_images=moving_images/m
if reference.strip().isdigit():
# print("User input is Number")
reference=int(reference)
idx2=np.ones(v)*reference
idx2=idx2.astype(int)
fixed_images = d[idx2, ..., np.newaxis]
fixed_images=fixed_images/m
else:
# print("User input is string")
img = nib.load(reference)
img_data = img.get_fdata()
if img.shape[0:2]!=(64,64):
img_data = img_data[23:87,23:87,:]
img_data=img_data[np.newaxis,:,:,slice_ID]
idx2=np.zeros(v)
idx2=idx2.astype(int)
fixed_images = img_data[idx2, ..., np.newaxis]
fixed_images=fixed_images/m
inputs = [moving_images, fixed_images]
# prepare outputs (the 'true' moved image):
# of course, we don't have this, but we know we want to compare
# the resulting moved image with the fixed image.
# we also wish to penalize the deformation field.
outputs = [fixed_images,zero_phi]
return (inputs, outputs)
```
### Predict function
```
"""
arg*
input_direction: input Raw data
reference: reference volume
output_direction: output motion corrected data
maximum_intensity: for normalization
loadable_model: loading trained model
"""
def main (input_direction,reference,output_direction,maximum_intensity,loadable_model):
start_time = time.time()
img_data,img=load_with_head(input_direction)
slice_number = img_data.shape[2]
header=img.header
img_mask_affine = img.affine
# configure unet input shape (concatenation of moving and fixed images)
ndim = 2
unet_input_features = 2
# data shape 64*64
s=(64,64)
inshape = (*s, unet_input_features)
# configure unet features
nb_features =[
[64, 64, 64, 64], # encoder features
[64, 64, 64, 64, 64, 32,16] # decoder features
]
# build model using VxmDense
inshape =s
vxm_model = vxm.networks.VxmDense(inshape, nb_features, int_steps=0)
# voxelmorph has a variety of custom loss classes
losses = [vxm.losses.MSE().loss, vxm.losses.Grad('l2').loss]
# usually, we have to balance the two losses by a hyper-parameter
lambda_param = 0.05
loss_weights = [1, lambda_param]
vxm_model.compile(optimizer='Adam', loss=losses, loss_weights=loss_weights, metrics=['accuracy'])
vxm_model.load_weights(loadable_model)
o=np.zeros((img_data.shape[0],img_data.shape[1],img_data.shape[2],img_data.shape[3]))
for i in range(slice_number):
prepare_data=ref(input_direction,maximum_intensity,i,reference)
val_input, _ = prepare_data
val_pred = vxm_model.predict(val_input)
change_order= np.einsum('jki->kij',val_pred[0][:,:,:,0])
o[:, :, i,:] = change_order
img_reg = nib.Nifti1Image(o*maximum_intensity, affine=img_mask_affine, header=header)
nib.save(img_reg,output_direction)
print("--- %s second ---" % (time.time() - start_time))
```
### Calculate SNR parameter
```
def snr (direction):
img = nib.load(direction)
img = img.get_fdata()
mean=[]
for i in range(img.shape[2]):
mean.append(np.mean(img[:,:,i]))
mean=np.array(mean)
deviation=[]
for i in range(img.shape[2]):
deviation.append(np.std(img[:,:,i]))
deviation=np.array(deviation)
return (mean/deviation),mean,deviation
```
### Mean across slice
```
def mean(direction):
img = nib.load(direction)
img = img.get_fdata()
mean=[]
where_are_NaNs = isnan(img)
img[where_are_NaNs] = 0
for i in range(img.shape[2]):
mean.append(np.mean(img[:,:,i]))
mean.append(np.mean(mean))
mean=np.array(mean)
return mean
```
### Mean across a specific region
```
"""
extract mean of segmented part
img[:,:,i][img[:,:,i] != 0
"""
def seg_mean(img):
p=0
for m in range(img.shape[0]):
for n in range(img.shape[1]):
if img[m,n]==0:
p=p+1
s=np.sum(img[:,:])
mean=s/((64*64)-p)
return mean
"""
This function calculate the mean of the ROI mask
note: it remove the non interested area.
"""
def mean_all(direction):
img = nib.load(direction)
img = img.get_fdata()
mean=[]
where_are_NaNs = np.isnan(img)
img[where_are_NaNs] = 0
for i in range(img.shape[2]):
mean.append(seg_mean(img[:,:,i]))
mean=np.mean(mean)
return mean
```
### Shifting images
```
"""
Shifts images across dx and dy
X: source image
"""
def shift_image(X, dx, dy):
X = np.roll(X, dy, axis=0)
X = np.roll(X, dx, axis=1)
if dy>0:
X[:dy, :] = 0
elif dy<0:
X[dy:, :] = 0
if dx>0:
X[:, :dx] = 0
elif dx<0:
X[:, dx:] = 0
return X
```
## DeepRetroMoCO
```
"""
arg*
source_centerline_directory: reference centerline
centerlines_directory: data centerline
main_data_directory: Raw data
center_fix_directory: directory of FC(fix centerline) data------------------ID_c.nifti
final_cplus_directory: final motion corrected data directory----------------ID_cplus.nifti
maximum_intensity: normalization
model: trained model
reference: fix image volume
******
# if reference=0 means reference=first volume
# if reference=-1 means reference=mid volume
# if reference=-2 means reference=mean volume
# if reference>0 means reference=any volume
******
mean_directory: directory of mean image ** if reference is mean**
"""
def cplus(source_centerline_directory,centerlines_directory,main_data_directory,
center_fix_directory,final_cplus_directory,
maximum_intensity,model,reference,mean_directory
):
#############################################
# if reference=0 means reference=first volume
# if reference=-1 means reference=mid volume
# if reference=-2 means reference=mean volume
# if reference>0 means reference=any volume
Xs=[]
Ys=[]
source = pd.read_csv(source_centerline_directory, header=None)
source.columns=['x','y','delete']
source = source[['x','y']]
for s in range(source.shape[0]):
c=source.loc[s]
#xs=int(c['x'])
ys=int(c['y'])
#Xs.append(xs)
Ys.append(ys)
n2,name2=count_endwith(centerlines_directory,'.csv')
dx=[]
dy=[]
for s in range(0,source.shape[0]):
for j in range(n2):
df = pd.read_csv(centerlines_directory+name2[j][0], header=None)
df.columns=['x','y','delete']
df=df[['x','y']]
c=df.loc[s]
#x=int(c['x'])
y=int(c['y'])
#dx.append(Xs[s]-x)
dy.append(Ys[s]-y)
input_direction=main_data_directory
img = nib.load(input_direction)
img_data=img.get_fdata()
img_mask_affine = img.affine
header = img.header
nb_img = header.get_data_shape()
o=np.zeros((nb_img[0],nb_img[1],nb_img[2],nb_img[3]))
DX=np.zeros(len(dy))
start=0
for s in range(0,source.shape[0]):
for v in range(n2):
a= shift_image(img_data[:,:,s,v],dy[v+start],DX[v+start])
o[:,:,s, v] = a
start=start + n2
input_direction=center_fix_directory
img_reg = nib.Nifti1Image(o, affine=img_mask_affine, header=header)
nib.save(img_reg,input_direction)
if reference>0:
reference=str(reference)
if reference==0:
reference='0'
if reference==-1:
y=int(n2/2)
reference=str(y)
if reference==-2:
reference=mean_directory
main(input_direction,reference,final_cplus_directory,maximum_intensity,model)
```
### Tool for finding data
```
"""
arg*
data_dir: input directory
prefix: finding data by this prefix
"""
def count_startwith (data_dir,prefix):
train_dir = os.path.join(data_dir)
train_data_num = []
for file in os.listdir(train_dir):
if file.startswith(prefix):
train_data_num.append([file])
train_data_num=np.array(train_data_num)
n=train_data_num.shape[0]
return n,sorted(train_data_num)
def count_endwith (data_dir,prefix):
train_dir = os.path.join(data_dir)
train_data_num = []
for file in os.listdir(train_dir):
if file.endswith(prefix):
train_data_num.append([file])
train_data_num=np.array(train_data_num)
n=train_data_num.shape[0]
return n,sorted(train_data_num)
```
### Movement plots for one slice
```
"""
arg*
input_direction: "
reference: "
maximum_intensity: "
loadable_model: "
slice_num: Specific slice number to draw
mean_directory: "
title: title of figure
"""
def flow_one_slice(input_direction,reference,maximum_intensity,loadable_model,slice_num,mean_directory,title):
img_data,img=load_with_head(input_direction)
slice_number = img_data.shape[2]
header=img.header
img_mask_affine = img.affine
# configure unet input shape (concatenation of moving and fixed images)
ndim = 2
unet_input_features = 2
# data shape 64*64
s=(64,64)
inshape = (*s, unet_input_features)
# configure unet features
nb_features =[
[64, 64, 64, 64], # encoder features
[64, 64, 64, 64, 64, 32,16] # decoder features
]
# build model using VxmDense
inshape =s
vxm_model = vxm.networks.VxmDense(inshape, nb_features, int_steps=0)
# voxelmorph has a variety of custom loss classes
losses = [vxm.losses.MSE().loss, vxm.losses.Grad('l2').loss]
# usually, we have to balance the two losses by a hyper-parameter
lambda_param = 0.05
loss_weights = [1, lambda_param]
vxm_model.compile(optimizer='Adam', loss=losses, loss_weights=loss_weights, metrics=['accuracy'])
vxm_model.load_weights(loadable_model)
if reference>0:
reference=str(reference)
if reference==0:
reference='0'
if reference==-1:
y=int(img_data.shape[3]/2)
reference=str(y)
if reference==-2:
reference=mean_directory
#for i in range(slice_number):
#slice_number=5
prepare_data=ref(input_direction,maximum_intensity,slice_num,reference)
val_input, _ = prepare_data
val_pred = vxm_model.predict(val_input)
#val_pred=flow(input_direction,reference,maximum_intensity,loadable_model,slice_num)
x=[]
y=[]
for i in range(val_pred[1][:,0,0,0].shape[0]):
x.append(np.mean(val_pred[1][i,...,0]))
y.append(np.mean(val_pred[1][i,...,1]))
x=np.array(x)
y=np.array(y)
volume=range(val_pred[1][:,0,0,0].shape[0])
plt.figure(figsize=(20,5))
plt.plot(volume,x,label = "x")
plt.plot(volume,y,label = "y")
# naming the x axis
plt.xlabel('volumes')
# naming the y axis
plt.ylabel('movement')
# giving a title to my graph
plt.title(title)
# show a legend on the plot
plt.legend()
```
### Movement plot for all slice in one plot
```
"""
arg*
input_direction: "
reference: "
maximum_intensity: "
loadable_model: "
slice_num: Specific slice number to draw
mean_directory: "
title: title of figure
"""
def flow_all_slice(input_direction,reference,maximum_intensity,loadable_model,mean_directory,title):
img_data,img=load_with_head(input_direction)
slice_number = img_data.shape[2]
header=img.header
img_mask_affine = img.affine
# configure unet input shape (concatenation of moving and fixed images)
ndim = 2
unet_input_features = 2
# data shape 64*64
s=(64,64)
inshape = (*s, unet_input_features)
# configure unet features
nb_features =[
[64, 64, 64, 64], # encoder features
[64, 64, 64, 64, 64, 32,16] # decoder features
]
# build model using VxmDense
inshape =s
vxm_model = vxm.networks.VxmDense(inshape, nb_features, int_steps=0)
# voxelmorph has a variety of custom loss classes
losses = [vxm.losses.MSE().loss, vxm.losses.Grad('l2').loss]
# usually, we have to balance the two losses by a hyper-parameter
lambda_param = 0.05
loss_weights = [1, lambda_param]
vxm_model.compile(optimizer='Adam', loss=losses, loss_weights=loss_weights, metrics=['accuracy'])
vxm_model.load_weights(loadable_model)
if reference>0:
reference=str(reference)
if reference==0:
reference='0'
if reference==-1:
y=int(img_data.shape[3]/2)
reference=str(y)
if reference==-2:
reference=mean_directory
x_all_slice=[]
y_all_slice=[]
for i in range(slice_number):
prepare_data=ref(input_direction,maximum_intensity,i,reference)
val_input, _ = prepare_data
val_pred = vxm_model.predict(val_input)
#val_pred=flow(input_direction,reference,maximum_intensity,loadable_model,slice_num)
x=[]
y=[]
for i in range(val_pred[1][:,0,0,0].shape[0]):
x.append(np.mean(val_pred[1][i,...,0]))
y.append(np.mean(val_pred[1][i,...,1]))
x_all_slice.append(x)
y_all_slice.append(y)
x_all_slice=np.array(x_all_slice)
y_all_slice=np.array(y_all_slice)
mean_x=x_all_slice.mean(axis=0)
mean_y=y_all_slice.mean(axis=0)
### for delete the eror for reference to reference
mean_x[int(reference)]=0
mean_y[int(reference)]=0
overal=(mean_x+mean_y)/2
volume=range(val_pred[1][:,0,0,0].shape[0])
plt.figure(figsize=(20,5))
plt.plot(volume,overal,label = "x")
#plt.plot(volume,mean_y,label = "y")
# naming the x axis
plt.xlabel('volumes')
# naming the y axis
plt.ylabel('movement')
# giving a title to my graph
plt.title(title)
# show a legend on the plot
plt.legend()
```
### Showing Flow field map
```
def flow_between_two(input_direction0,input_direction1,reference,
maximum_intensity,loadable_model,mean_directory,
title,label1,label2):
img_data,img=load_with_head(input_direction0)
slice_number = img_data.shape[2]
header=img.header
img_mask_affine = img.affine
# configure unet input shape (concatenation of moving and fixed images)
ndim = 2
unet_input_features = 2
# data shape 64*64
s=(64,64)
inshape = (*s, unet_input_features)
# configure unet features
nb_features =[
[64, 64, 64, 64], # encoder features
[64, 64, 64, 64, 64, 32,16] # decoder features
]
# build model using VxmDense
inshape =s
vxm_model = vxm.networks.VxmDense(inshape, nb_features, int_steps=0)
# voxelmorph has a variety of custom loss classes
losses = [vxm.losses.MSE().loss, vxm.losses.Grad('l2').loss]
# usually, we have to balance the two losses by a hyper-parameter
lambda_param = 0.05
loss_weights = [1, lambda_param]
vxm_model.compile(optimizer='Adam', loss=losses, loss_weights=loss_weights, metrics=['accuracy'])
vxm_model.load_weights(loadable_model)
if reference>0:
reference=str(reference)
if reference==0:
reference='0'
if reference==-1:
y=int(img_data.shape[3]/2)
reference=str(y)
if reference==-2:
reference=mean_directory
x_all_slice=[]
y_all_slice=[]
for i in range(slice_number):
prepare_data=ref(input_direction0,maximum_intensity,i,reference)
val_input, _ = prepare_data
val_pred = vxm_model.predict(val_input)
#val_pred=flow(input_direction,reference,maximum_intensity,loadable_model,slice_num)
x=[]
y=[]
for i in range(val_pred[1][:,0,0,0].shape[0]):
x.append(np.mean(val_pred[1][i,...,0]))
y.append(np.mean(val_pred[1][i,...,1]))
x_all_slice.append(x)
y_all_slice.append(y)
x_all_slice=np.array(x_all_slice)
y_all_slice=np.array(y_all_slice)
mean_x=x_all_slice.mean(axis=0)
mean_y=y_all_slice.mean(axis=0)
### for delete the eror for reference to reference
mean_x[int(reference)]=0
mean_y[int(reference)]=0
overal=(mean_x+mean_y)/2
x_all_slice=[]
y_all_slice=[]
for i in range(slice_number):
prepare_data=ref(input_direction1,maximum_intensity,i,reference)
val_input, _ = prepare_data
val_pred = vxm_model.predict(val_input)
#val_pred=flow(input_direction,reference,maximum_intensity,loadable_model,slice_num)
x=[]
y=[]
for i in range(val_pred[1][:,0,0,0].shape[0]):
x.append(np.mean(val_pred[1][i,...,0]))
y.append(np.mean(val_pred[1][i,...,1]))
x_all_slice.append(x)
y_all_slice.append(y)
x_all_slice=np.array(x_all_slice)
y_all_slice=np.array(y_all_slice)
mean_x=x_all_slice.mean(axis=0)
mean_y=y_all_slice.mean(axis=0)
### for delete the eror for reference to reference
mean_x[int(reference)]=0
mean_y[int(reference)]=0
overal1=(mean_x+mean_y)/2
volume=range(val_pred[1][:,0,0,0].shape[0])
plt.figure(figsize=(25,10))
plt.plot(volume,overal,label = label1)
plt.plot(volume,overal1,label = label2)
# naming the x axis
plt.xlabel('volumes',fontsize=18)
# naming the y axis
plt.ylabel('movement',fontsize=18)
# giving a title to my graph
plt.title(title,fontsize=20)
# show a legend on the plot
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
# show a legend on the plot
plt.legend()
plt.grid()
plt.legend(fontsize=15)
```
| github_jupyter |
```
'''
A Convolutional Network implementation example using TensorFlow library.
This example is using the MNIST database of handwritten digits
(http://yann.lecun.com/exdb/mnist/)
Author: Aymeric Damien
Project: https://github.com/aymericdamien/TensorFlow-Examples/
'''
import tensorflow as tf
# Import MNIST data
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
# Parameters
learning_rate = 0.001
training_iters = 200000
batch_size = 128
display_step = 10
# Network Parameters
n_input = 784 # MNIST data input (img shape: 28*28)
n_classes = 10 # MNIST total classes (0-9 digits)
dropout = 0.75 # Dropout, probability to keep units
# tf Graph input
x = tf.placeholder(tf.float32, [None, n_input])
y = tf.placeholder(tf.float32, [None, n_classes])
keep_prob = tf.placeholder(tf.float32) #dropout (keep probability)
# Create some wrappers for simplicity
def conv2d(x, W, b, strides=1):
# Conv2D wrapper, with bias and relu activation
x = tf.nn.conv2d(x, W, strides=[1, strides, strides, 1], padding='SAME')
x = tf.nn.bias_add(x, b)
return tf.nn.relu(x)
def maxpool2d(x, k=2):
# MaxPool2D wrapper
return tf.nn.max_pool(x, ksize=[1, k, k, 1], strides=[1, k, k, 1],
padding='SAME')
# Create model
def conv_net(x, weights, biases, dropout):
# Reshape input picture
x = tf.reshape(x, shape=[-1, 28, 28, 1])
# Convolution Layer
conv1 = conv2d(x, weights['wc1'], biases['bc1'])
# Max Pooling (down-sampling)
conv1 = maxpool2d(conv1, k=2)
# Convolution Layer
conv2 = conv2d(conv1, weights['wc2'], biases['bc2'])
# Max Pooling (down-sampling)
conv2 = maxpool2d(conv2, k=2)
# Fully connected layer
# Reshape conv2 output to fit fully connected layer input
fc1 = tf.reshape(conv2, [-1, weights['wd1'].get_shape().as_list()[0]])
fc1 = tf.add(tf.matmul(fc1, weights['wd1']), biases['bd1'])
fc1 = tf.nn.relu(fc1)
# Apply Dropout
fc1 = tf.nn.dropout(fc1, dropout)
# Output, class prediction
out = tf.add(tf.matmul(fc1, weights['out']), biases['out'])
return out
# Store layers weight & bias
weights = {
# 5x5 conv, 1 input, 32 outputs
'wc1': tf.Variable(tf.random_normal([5, 5, 1, 32])),
# 5x5 conv, 32 inputs, 64 outputs
'wc2': tf.Variable(tf.random_normal([5, 5, 32, 64])),
# fully connected, 7*7*64 inputs, 1024 outputs
'wd1': tf.Variable(tf.random_normal([7*7*64, 1024])),
# 1024 inputs, 10 outputs (class prediction)
'out': tf.Variable(tf.random_normal([1024, n_classes]))
}
biases = {
'bc1': tf.Variable(tf.random_normal([32])),
'bc2': tf.Variable(tf.random_normal([64])),
'bd1': tf.Variable(tf.random_normal([1024])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
# Construct model
pred = conv_net(x, weights, biases, keep_prob)
# Define loss and optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=pred, labels=y))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
# Evaluate model
correct_pred = tf.equal(tf.argmax(pred, 1), tf.argmax(y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# Initializing the variables
init = tf.global_variables_initializer()
# Launch the graph
with tf.Session() as sess:
sess.run(init)
step = 1
# Keep training until reach max iterations
while step * batch_size < training_iters:
batch_x, batch_y = mnist.train.next_batch(batch_size)
# Run optimization op (backprop)
sess.run(optimizer, feed_dict={x: batch_x, y: batch_y,
keep_prob: dropout})
if step % display_step == 0:
# Calculate batch loss and accuracy
loss, acc = sess.run([cost, accuracy], feed_dict={x: batch_x,
y: batch_y,
keep_prob: 1.})
print "Iter " + str(step*batch_size) + ", Minibatch Loss= " + \
"{:.6f}".format(loss) + ", Training Accuracy= " + \
"{:.5f}".format(acc)
step += 1
print "Optimization Finished!"
# Calculate accuracy for 256 mnist test images
print "Testing Accuracy:", \
sess.run(accuracy, feed_dict={x: mnist.test.images[:256],
y: mnist.test.labels[:256],
keep_prob: 1.})
```
| github_jupyter |
## Import libraries
```
import pandas as pd
import seaborn as sb
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
from sklearn import preprocessing
from itertools import islice
import seaborn as sns
from scipy.stats import binom
from scipy.stats import norm
import statsmodels.stats.api as sms
```
## Load data
```
data_frame = pd.read_csv("adsmart.csv")
#sns.set()
data_frame.info()
```
#### Drop observations where users didn't respond to the questionaire
```
drop_mask = data_frame.query('yes==0 and no==0')
data_frame=data_frame.drop(drop_mask.index)
```
## Count of people that are aware of the smartAd brand for both experiment groups
```
plt.figure(figsize=(9,7))
ax=sns.countplot(x ='experiment', hue='yes', data = data_frame)
ax.set_xlabel('Experiment', fontsize=15)
ax.set_ylabel('Count' , fontsize=15)
ax.set_title('Count of aware people per experiment group', fontsize=15, fontweight='bold')
plt.legend( ['No', 'Yes'])
```
### Count plot of browsers. It is dominated by chrome mobile
```
plt.figure(figsize=(9,7))
ax= sns.countplot(x ='browser', data = data_frame)
ax.set_xticklabels(ax.get_xticklabels(), rotation=60, ha="right")
plt.tight_layout()
ax.set_xlabel('Browsers', fontsize=15)
ax.set_ylabel('Count' , fontsize=15)
ax.set_title('Count of Browsers used', fontsize=15, fontweight='bold')
top_5_device= data_frame['device_make'].value_counts().nlargest(5)
fig1, ax1 = plt.subplots(figsize = (10, 10))
ax1.pie(top_5_device.values, labels=top_5_device.index, autopct='%1.1f%%', shadow=True)
ax1.axis('equal')
ax1.set_title('Pie plot of smart phones', fontsize=15, fontweight='bold')
plt.show()
```
## 85.7% of the recorded data were from an unkonwn device, this makes it difficult for machine learning models to use device_make.
### The following distribution plot shows people use the internet (view the ad) throughout the day. A peak is observed at 16:00
```
plt.figure(figsize=(9,7))
top_5_device= data_frame['device_make'].value_counts().nlargest(5)
ax=sns.distplot(data_frame['hour'],bins=20)
ax.set_xlabel('Hour', fontsize=15)
ax.set_ylabel('Probability' , fontsize=15)
plt.title('Distribution of Hour')
# sns.distplot()
```
## Split the dataframe to the two experiment groups. The number of total people and aware people are also counted.
```
exposed_yes = data_frame.query('experiment == "exposed"').yes
exposed_count = len(exposed_yes)
exposed_yes_count = exposed_yes.sum(axis=0)
control_yes = data_frame.query('experiment == "control"').yes
control_count = len(control_yes)
control_yes_count = control_yes.sum(axis=0)
print('Converted Control:',control_yes_count,' Total Control:',control_count)
print('Converted Exposed:',exposed_yes_count,' Total Exposed:',exposed_count)
```
## Determine the required sample size.
```
baseline_rate = control_yes_count / control_count
practical_significance = 0.01
confidence_level = 0.05
sensitivity = 0.8
effect_size = sms.proportion_effectsize(baseline_rate, baseline_rate + practical_significance)
sample_size = sms.NormalIndPower().solve_power(effect_size = effect_size, power = sensitivity,
alpha = confidence_level, ratio=1)
print("Required sample size: ", round(sample_size), " per group")
```
## As shown above, we are short of the required sample size since we have only 586 and 657 in each group
## Determine the probability of having x number of click throughs
```
cv_rate_control, cv_rate_exposed = control_yes_count / control_count, exposed_yes_count / exposed_count
range = np.arange(200, 360)
cv_prob_control = binom(control_count, cv_rate_control).pmf(range)
cv_prob_exposed = binom(exposed_count, cv_rate_exposed).pmf(range)
fig, ax = plt.subplots(figsize=(12,10))
plt.bar(range, cv_prob_control, label="Control",color='red')
plt.bar(range, cv_prob_exposed, label="Exposed",color='green')
plt.legend()
plt.xlabel("Conversions"); plt.ylabel("Probability");
```
### So we can see here that the exposed group has an edge.
## Calculate standard deviations for each experiment group. This will help us change our binomial distribution to normal by using the central limit theorem. We can then calculate z-scores.
```
std_dev_control = np.sqrt(cv_rate_control * (1 - cv_rate_control) / control_count)
std_dev_exposed = np.sqrt(cv_rate_exposed * (1 - cv_rate_exposed) / exposed_count)
```
## Calculate conversion rates for each experiment group and plot the probability distribution
```
conversion_rate = np.linspace(0, 0.9, 200)
prob_a = norm(cv_rate_control, std_dev_control).pdf(conversion_rate)
prob_b = norm(cv_rate_exposed, std_dev_exposed).pdf(conversion_rate)
plt.figure(figsize=(10,5))
plt.plot(conversion_rate, prob_a, label="A")
plt.plot(conversion_rate, prob_b, label="B")
plt.legend(frameon=False)
plt.xlabel("Conversion rate"); plt.ylabel("Probability");
```
### So we can see here that the exposed group has an edge.
## Calculate z-score and p-value.
```
z_score = (cv_rate_exposed - cv_rate_control) / np.sqrt(std_dev_control**2 + std_dev_exposed**2)
p = norm(cv_rate_exposed - cv_rate_control, np.sqrt(std_dev_control**2 + std_dev_exposed**2))
x = np.linspace(-0.05, 0.15, 1000)
y = p.pdf(x)
area_under_curve = p.sf(0)
plt.figure(figsize=(12,8))
plt.plot(x, y, label="Density Function")
plt.fill_between(x, 0, y, where=x>0, label="Prob(b>a)", alpha=0.3)
plt.annotate(f"Area={area_under_curve:0.3f}", (0.02, 5))
plt.legend()
plt.xlabel("Difference in conversion rate"); plt.ylabel("Prob");
print(f"zscore is {z_score:0.3f}, with p-value {norm().sf(z_score):0.3f}")
```
## Since the p-value(0.259)>0.05, we accept the null hypothesis.
### From our A/B test we can see that even if we saw a rise in conversion rate in the exposed group for this particular experiment, it may be a result of random chance. So we cannot conclude that the experiment group has more awareness.
### But if we analyze the distribution graph and the power value we understand that the sample size taken to condact this testing is very small. There is a very high probability that we will make a type-II error. We need to gather more data to make a sounding conclusion from this testing.
| github_jupyter |
## Dependencies
```
import json, warnings, shutil
from jigsaw_utility_scripts import *
from transformers import TFXLMRobertaModel, XLMRobertaConfig
from tensorflow.keras.models import Model
from tensorflow.keras import optimizers, metrics, losses, layers
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
SEED = 0
seed_everything(SEED)
warnings.filterwarnings("ignore")
```
## TPU configuration
```
strategy = set_up_strategy()
print("REPLICAS: ", strategy.num_replicas_in_sync)
AUTO = tf.data.experimental.AUTOTUNE
```
# Load data
```
database_base_path = '/kaggle/input/jigsaw-dataset-split-toxic-roberta-base-192/'
k_fold = pd.read_csv(database_base_path + '5-fold.csv')
display(k_fold.head())
# Unzip files
!tar -xvf /kaggle/input/jigsaw-dataset-split-toxic-roberta-base-192/fold_1.tar.gz
!tar -xvf /kaggle/input/jigsaw-dataset-split-toxic-roberta-base-192/fold_2.tar.gz
!tar -xvf /kaggle/input/jigsaw-dataset-split-toxic-roberta-base-192/fold_3.tar.gz
# !tar -xvf /kaggle/input/jigsaw-dataset-split-toxic-roberta-base-192/fold_4.tar.gz
# !tar -xvf /kaggle/input/jigsaw-dataset-split-toxic-roberta-base-192/fold_5.tar.gz
```
# Model parameters
```
base_path = '/kaggle/input/jigsaw-transformers/XLM-RoBERTa/'
config = {
"MAX_LEN": 192,
"BATCH_SIZE": 16 * strategy.num_replicas_in_sync,
"EPOCHS": 3,
"LEARNING_RATE": 1e-5,
"ES_PATIENCE": 1,
"N_FOLDS": 3,
"base_model_path": base_path + 'tf-xlm-roberta-base-tf_model.h5',
"config_path": base_path + 'xlm-roberta-base-config.json'
}
with open('config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
```
# Model
```
module_config = XLMRobertaConfig.from_pretrained(config['config_path'], output_hidden_states=False)
def model_fn(MAX_LEN):
input_ids = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='input_ids')
attention_mask = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='attention_mask')
base_model = TFXLMRobertaModel.from_pretrained(config['base_model_path'], config=module_config)
sequence_output = base_model({'input_ids': input_ids, 'attention_mask': attention_mask})
last_state = sequence_output[0]
cls_token = last_state[:, 0, :]
output = layers.Dense(1, activation='sigmoid', name='output')(cls_token)
model = Model(inputs=[input_ids, attention_mask], outputs=output)
model.compile(optimizers.Adam(lr=config['LEARNING_RATE']),
loss=losses.BinaryCrossentropy(),
metrics=[metrics.BinaryAccuracy(), metrics.AUC()])
return model
```
# Train
```
history_list = []
for n_fold in range(config['N_FOLDS']):
print('\nFOLD: %d' % (n_fold+1))
# Load data
base_data_path = 'fold_%d/' % (n_fold+1)
x_train = np.load(base_data_path + 'x_train.npy')
y_train = np.load(base_data_path + 'y_train.npy')
x_valid = np.load(base_data_path + 'x_valid.npy')
y_valid = np.load(base_data_path + 'y_valid.npy')
step_size = x_train.shape[0] // config['BATCH_SIZE']
### Delete data dir
shutil.rmtree(base_data_path)
# Train model
model_path = 'model_fold_%d.h5' % (n_fold+1)
es = EarlyStopping(monitor='val_loss', mode='min', patience=config['ES_PATIENCE'],
restore_best_weights=True, verbose=1)
checkpoint = ModelCheckpoint(model_path, monitor='val_loss', mode='min',
save_best_only=True, save_weights_only=True, verbose=1)
with strategy.scope():
model = model_fn(config['MAX_LEN'])
history = model.fit(list(x_train), y_train,
validation_data=(list(x_valid), y_valid),
callbacks=[checkpoint, es],
epochs=config['EPOCHS'],
batch_size=config['BATCH_SIZE'],
steps_per_epoch=step_size,
verbose=2).history
history_list.append(history)
# Make predictions
train_preds = model.predict(list(x_train))
valid_preds = model.predict(list(x_valid))
k_fold.loc[k_fold['fold_%d' % (n_fold+1)] == 'train', 'pred_%d' % (n_fold+1)] = np.round(train_preds)
k_fold.loc[k_fold['fold_%d' % (n_fold+1)] == 'validation', 'pred_%d' % (n_fold+1)] = np.round(valid_preds)
```
## Model loss graph
```
sns.set(style="whitegrid")
for n_fold in range(config['N_FOLDS']):
print('Fold: %d' % (n_fold+1))
plot_metrics(history_list[n_fold])
```
# Model evaluation
```
display(evaluate_model(k_fold, config['N_FOLDS']).style.applymap(color_map))
```
# Confusion matrix
```
for n_fold in range(config['N_FOLDS']):
print('Fold: %d' % (n_fold+1))
train_set = k_fold[k_fold['fold_%d' % (n_fold+1)] == 'train']
validation_set = k_fold[k_fold['fold_%d' % (n_fold+1)] == 'validation']
plot_confusion_matrix(train_set['toxic'], train_set['pred_%d' % (n_fold+1)],
validation_set['toxic'], validation_set['pred_%d' % (n_fold+1)])
```
# Visualize predictions
```
display(k_fold[['comment_text', 'toxic'] + [c for c in k_fold.columns if c.startswith('pred')]].head(15))
```
| github_jupyter |
## CIFAR 10
```
%matplotlib inline
%reload_ext autoreload
%autoreload 2
```
You can get the data via:
wget http://pjreddie.com/media/files/cifar.tgz
```
from fastai.conv_learner import *
from pathlib import Path
if os.name == 'nt':
PATH = str(Path.home()) + "\\data\\cifar10\\"
else:
PATH = "data/cifar10/"
os.makedirs(PATH, exist_ok=True)
def moveFilesToSubDirsFromFileName(path):
files = os.listdir(path)
for f in files:
if os.path.isdir(os.path.join(path, f)):
continue
filename, file_extension = os.path.splitext(f)
regex = re.compile('[^a-zA-Z]')
file_label = regex.sub('', filename)
target_folder = os.path.join(path, file_label)
if not os.path.exists(target_folder): os.makedirs(target_folder)
os.rename(os.path.join(path, f), os.path.join(target_folder, f))
moveFilesToSubDirsFromFileName(os.path.join(PATH, "train"))
moveFilesToSubDirsFromFileName(os.path.join(PATH, "test"))
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
stats = (np.array([ 0.4914 , 0.48216, 0.44653]), np.array([ 0.24703, 0.24349, 0.26159]))
def get_data(sz,bs):
tfms = tfms_from_stats(stats, sz, aug_tfms=[RandomFlip()], pad=sz//8)
return ImageClassifierData.from_paths(PATH, val_name='test', tfms=tfms, bs=bs)
bs=256
```
### Look at data
```
data = get_data(32, 4)
x,y=next(iter(data.trn_dl))
plt.imshow(data.trn_ds.denorm(x)[0]);
plt.imshow(data.trn_ds.denorm(x)[1]);
```
## Fully connected model
```
data = get_data(32,bs)
lr=1e-2
```
From [this notebook](https://github.com/KeremTurgutlu/deeplearning/blob/master/Exploring%20Optimizers.ipynb) by our student Kerem Turgutlu:
```
class SimpleNet(nn.Module):
def __init__(self, layers):
super().__init__()
self.layers = nn.ModuleList([
nn.Linear(layers[i], layers[i + 1]) for i in range(len(layers) - 1)])
def forward(self, x):
x = x.view(x.size(0), -1)
for l in self.layers:
l_x = l(x)
x = F.relu(l_x)
return F.log_softmax(l_x, dim=-1)
learn = ConvLearner.from_model_data(SimpleNet([32*32*3, 40,10]), data)
learn, [o.numel() for o in learn.model.parameters()]
learn.summary()
%time learn.lr_find()
learn.sched.plot()
%time learn.fit(lr, 2)
%time learn.fit(lr, 2, cycle_len=1)
```
## CNN
```
class ConvNet(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.layers = nn.ModuleList([
nn.Conv2d(layers[i], layers[i + 1], kernel_size=3, stride=2)
for i in range(len(layers) - 1)])
self.pool = nn.AdaptiveMaxPool2d(1)
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
for l in self.layers: x = F.relu(l(x))
x = self.pool(x)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvNet([3, 20, 40, 80], 10), data)
learn.summary()
learn.lr_find(end_lr=100)
learn.sched.plot()
%time learn.fit(1e-1, 2)
%time learn.fit(1e-1, 4, cycle_len=1)
```
## Refactored
```
class ConvLayer(nn.Module):
def __init__(self, ni, nf):
super().__init__()
self.conv = nn.Conv2d(ni, nf, kernel_size=3, stride=2, padding=1)
def forward(self, x): return F.relu(self.conv(x))
class ConvNet2(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.layers = nn.ModuleList([ConvLayer(layers[i], layers[i + 1])
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
for l in self.layers: x = l(x)
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvNet2([3, 20, 40, 80], 10), data)
learn.summary()
%time learn.fit(1e-1, 2)
%time learn.fit(1e-1, 2, cycle_len=1)
```
## BatchNorm
```
class BnLayer(nn.Module):
def __init__(self, ni, nf, stride=2, kernel_size=3):
super().__init__()
self.conv = nn.Conv2d(ni, nf, kernel_size=kernel_size, stride=stride,
bias=False, padding=1)
self.a = nn.Parameter(torch.zeros(nf,1,1))
self.m = nn.Parameter(torch.ones(nf,1,1))
def forward(self, x):
x = F.relu(self.conv(x))
x_chan = x.transpose(0,1).contiguous().view(x.size(1), -1)
if self.training:
self.means = x_chan.mean(1)[:,None,None]
self.stds = x_chan.std (1)[:,None,None]
return (x-self.means) / self.stds *self.m + self.a
class ConvBnNet(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.conv1 = nn.Conv2d(3, 10, kernel_size=5, stride=1, padding=2)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i + 1])
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
x = self.conv1(x)
for l in self.layers: x = l(x)
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvBnNet([10, 20, 40, 80, 160], 10), data)
learn.summary()
%time learn.fit(3e-2, 2)
%time learn.fit(1e-1, 4, cycle_len=1)
```
## Deep BatchNorm
```
class ConvBnNet2(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.conv1 = nn.Conv2d(3, 10, kernel_size=5, stride=1, padding=2)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i+1])
for i in range(len(layers) - 1)])
self.layers2 = nn.ModuleList([BnLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
x = self.conv1(x)
for l,l2 in zip(self.layers, self.layers2):
x = l(x)
x = l2(x)
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(ConvBnNet2([10, 20, 40, 80, 160], 10), data)
%time learn.fit(1e-2, 2)
%time learn.fit(1e-2, 2, cycle_len=1)
```
## Resnet
```
class ResnetLayer(BnLayer):
def forward(self, x): return x + super().forward(x)
class Resnet(nn.Module):
def __init__(self, layers, c):
super().__init__()
self.conv1 = nn.Conv2d(3, 10, kernel_size=5, stride=1, padding=2)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i+1])
for i in range(len(layers) - 1)])
self.layers2 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.layers3 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
def forward(self, x):
x = self.conv1(x)
for l,l2,l3 in zip(self.layers, self.layers2, self.layers3):
x = l3(l2(l(x)))
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(Resnet([10, 20, 40, 80, 160], 10), data)
wd=1e-5
%time learn.fit(1e-2, 2, wds=wd)
%time learn.fit(1e-2, 3, cycle_len=1, cycle_mult=2, wds=wd)
%time learn.fit(1e-2, 8, cycle_len=4, wds=wd)
```
## Resnet 2
```
class Resnet2(nn.Module):
def __init__(self, layers, c, p=0.5):
super().__init__()
self.conv1 = BnLayer(3, 16, stride=1, kernel_size=7)
self.layers = nn.ModuleList([BnLayer(layers[i], layers[i+1])
for i in range(len(layers) - 1)])
self.layers2 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.layers3 = nn.ModuleList([ResnetLayer(layers[i+1], layers[i + 1], 1)
for i in range(len(layers) - 1)])
self.out = nn.Linear(layers[-1], c)
self.drop = nn.Dropout(p)
def forward(self, x):
x = self.conv1(x)
for l,l2,l3 in zip(self.layers, self.layers2, self.layers3):
x = l3(l2(l(x)))
x = F.adaptive_max_pool2d(x, 1)
x = x.view(x.size(0), -1)
x = self.drop(x)
return F.log_softmax(self.out(x), dim=-1)
learn = ConvLearner.from_model_data(Resnet2([16, 32, 64, 128, 256], 10, 0.2), data)
wd=1e-6
%time learn.fit(1e-2, 2, wds=wd)
%time learn.fit(1e-2, 3, cycle_len=1, cycle_mult=2, wds=wd)
%time learn.fit(1e-2, 8, cycle_len=4, wds=wd)
learn.save('tmp3')
log_preds,y = learn.TTA()
preds = np.mean(np.exp(log_preds),0)
metrics.log_loss(y,preds), accuracy(preds,y)
```
### End
| github_jupyter |
# Graphing Data with matplotlib
We will extract data series to plot from pandas DataFrames but, first, let's do a very simple example of a matplotlib line chart and scatterplot.
The statement below causes the matplotlib graphs to appear within the Jupter notebook rather than in a separate window that pops up. This can be very convenient because, among other reasons, the image will be retained in the notebook and computation will not be paused when many graphs are constructed within a loop. Statements that start with '%' are called magic functions. I think it is also fair to say that these are system-level commands that you might type in a DOS Command Window. The '%' indicates a system command so that it is executed properly. This web page talks about magis functions: https://stackoverflow.com/questions/20961287/what-is-pylab
```
%matplotlib inline
```
The next statement imports the graphing package we will use and uses a very popular, if not ubiquitous alias, plt, for this matplotlib package.
```
import matplotlib.pyplot as plt
```
## Simple Examples First: Line and Scatterplot
The contents of matplotlib_eg_line.py are contained in the cell below. The plt.show() command is required if you are working in Spyder, but not if you are working in Jupyter and have executed this command: %matplotlib inline
```
xs = [0,82.0442626658045,164.140532825633,249.101916493353,332.182520454561,416.254670963008,495.330844802651,575.152803567278,653.205144040749,738.002602107511,818.027625347191,898.118246791891,980.079475188722,1061.14169245002,1139.27372549306,1222.46124841828,1300.35875796241,1375.56916894815,1450.58381247379,1528.61926864683,1605.58397275636,1690.73526419816,1765.8607155748,1844.76374052086,1927.68455634244,2007.86892722346,2086.83606915306,2166.71913598734,2251.80135023691,2335.70816516882,2420.79263834332,2499.86354911873,2577.05890165442,2654.19208140119,2730.09283946298,2809.07577275313,2894.08624312766,2969.05670691318,3049.93958564477,3129.75861740531,3206.68277248806,3283.60346903165,3364.58552562287,3449.40362446521,3525.4406380069,3608.13685182861,3690.05929028829,3773.14656467608,3850.22786804579,3930.30737684189,4015.46434790825,4093.50879464788,4172.64841837878,4252.66450124259,4335.57956650011,4420.52164467734,4495.52991250898,4575.39742270308,4650.49642074232,4730.52574677055,4805.68851119975,4880.60019974235,4960.70865715944,5043.67168991259,5118.73724644963,5203.54753389807,5283.38430140739,5359.51383129058,5438.3504224767,5519.19106748314,5597.38879699402]
ys = [69,70.5331166725557,71.5580608629997,71.7925736363559,71.3468902075964,70.8280365103217,70.6414982437025,70.4451053764375,70.3795952412711,69.7645648576213,69.5426154537094,69.3004779067882,68.9026747371149,68.599154906399,68.519069857217,68.3772433121902,69.1486631584102,68.7548264957532,68.3609585242701,68.5575021969504,68.5786288310157,70.1801398605569,69.7914710297875,70.1491959166127,71.3308639026079,71.9361568322099,71.7841530239482,71.5632100706621,70.9532872882161,70.4120650812618,69.7956890967353,69.6193830861858,69.6154149236862,69.6130427165012,69.7102463368457,69.5264209638887,68.9050613087613,69.1051712486311,68.8193037564598,68.6066819000086,68.6092749335593,68.6204785117934,68.3360360051156,68.5812312774717,68.3713770000006,69.5440095449469,70.5260768620176,72.0285425812187,72.0349254824144,71.8158332177273,71.1892654017154,71.1169346409977,70.8948191898231,70.6895661675202,70.2298913978404,69.6255849641529,69.7340391269025,69.5070018796742,69.6617353391113,69.4627544694931,69.630774569752,69.8026157834872,69.5269217792645,69.0549816241028,69.1984712324886,68.3562401007611,68.1422906748129,68.2062726166005,68.0498033914326,68.6018257745021,68.7606159741864]
plt.plot(xs,ys)
plt.show()
```
The contents of matplotlib_eg_scatter.py are contained in the cell below.
```
import matplotlib.pyplot as plt
xs = [0,82.0442626658045,164.140532825633,249.101916493353,332.182520454561,416.254670963008,495.330844802651, \
575.152803567278,653.205144040749,738.002602107511,818.027625347191,898.118246791891,980.079475188722, \
1061.14169245002,1139.27372549306,1222.46124841828,1300.35875796241,1375.56916894815,1450.58381247379, \
1528.61926864683,1605.58397275636,1690.73526419816,1765.8607155748,1844.76374052086,1927.68455634244, \
2007.86892722346,2086.83606915306,2166.71913598734,2251.80135023691,2335.70816516882,2420.79263834332, \
2499.86354911873,2577.05890165442,2654.19208140119,2730.09283946298,2809.07577275313,2894.08624312766, \
2969.05670691318,3049.93958564477,3129.75861740531,3206.68277248806,3283.60346903165,3364.58552562287, \
3449.40362446521,3525.4406380069,3608.13685182861,3690.05929028829,3773.14656467608,3850.22786804579, \
3930.30737684189,4015.46434790825,4093.50879464788,4172.64841837878,4252.66450124259,4335.57956650011, \
4420.52164467734,4495.52991250898,4575.39742270308,4650.49642074232,4730.52574677055,4805.68851119975, \
4880.60019974235,4960.70865715944,5043.67168991259,5118.73724644963,5203.54753389807,5283.38430140739, \
5359.51383129058,5438.3504224767,5519.19106748314,5597.38879699402]
ys = [69,70.5331166725557,71.5580608629997,71.7925736363559,71.3468902075964,70.8280365103217,70.6414982437025, \
70.4451053764375,70.3795952412711,69.7645648576213,69.5426154537094,69.3004779067882,68.9026747371149,68.599154906399, \
68.519069857217,68.3772433121902,69.1486631584102,68.7548264957532,68.3609585242701,68.5575021969504,68.5786288310157, \
70.1801398605569,69.7914710297875,70.1491959166127,71.3308639026079,71.9361568322099,71.7841530239482,71.5632100706621, \
70.9532872882161,70.4120650812618,69.7956890967353,69.6193830861858,69.6154149236862,69.6130427165012,69.7102463368457, \
69.5264209638887,68.9050613087613,69.1051712486311,68.8193037564598,68.6066819000086,68.6092749335593,68.6204785117934, \
68.3360360051156,68.5812312774717,68.3713770000006,69.5440095449469,70.5260768620176,72.0285425812187,72.0349254824144, \
71.8158332177273,71.1892654017154,71.1169346409977,70.8948191898231,70.6895661675202,70.2298913978404,69.6255849641529, \
69.7340391269025,69.5070018796742,69.6617353391113,69.4627544694931,69.630774569752,69.8026157834872,69.5269217792645, \
69.0549816241028,69.1984712324886,68.3562401007611,68.1422906748129,68.2062726166005,68.0498033914326,68.6018257745021, \
68.7606159741864]
plt.scatter(xs,ys)
plt.show()
```
The data for the previous graphs was typed, which is rearely done. We'll need to input data automatically from files, databases, and other sources. The pandas package provides a great resource for reading data files, and so we'll use pandas to input the data and manage it so we'll need to import that package.
# Inputting and Handling Data with the pandas package
We'll use pandas to input the data and manage it so we'll need to import that package.
```
import pandas as pd
```
These are the data we will be working with using the pandas DataFrame data type
df is often used as a variable name to denote a entity of the pandas DataFrame type
The pandas .read_csv() method is very useful. Note that the data files must have column headings in the first row. You'll, possibly, need to adjust the path to the location of your data file if it is not in the same folder as your default Jupyter folder.
The first data set below (stored in the df_oz DataFrame) is from William S. Cleveland and it is included in a typical R installation, whcih is my sources. These data describe temperature, wind speed, ozone, and radiation levels over time. The second data set, which is from the US Census web site, gives time series data for public (governmental) construction spending and private construction spending. The last data set, which is stored in the df_test DataFrame, gives high school standard of learning (SOL) scores for six course sections of a social studies course.
The double backslash, \\\, is used in the file paths below to avoid issues with escape characters. Alternately, a forward slash could have been used, /.
```
#df_oz = pd.read_csv('D:\\TeachingMaterials\\BusinessAnalytics\\Visualization\\VizData\\ozone.csv')
df_oz = pd.read_csv('ozone.csv')
df_oz
type(df_oz)
df_oz.head(n=3)
```
Before we start plotting, let's look at how you would find out about some of the properties of a DataFrame, atarting with descriptive statistics.
```
df_oz.describe()
```
The .shape command gives a tuple where the element in the 0th position is the number of rows in the DataFrame and the element in the 1st position indicates the number of columns.
```
df_oz.shape
```
The df_oz DataFrame contains four climatic data series for New York City. Retrieve the column heading to see what they are.
```
df_oz.columns
df_oz.columns.values
```
Extract a column of data from a DataFrame (a pandas Series data type)
```
df_oz['temperature']
type(df_oz['temperature'])
```
Extract a row from a DataFrame by index label.
```
df_oz.loc[50]
```
Extract a particular row element form a particular column.
```
df_oz['temperature'].loc[50]
type(df_oz['temperature'].loc[50])
type(df_oz.loc[50]['temperature'])
df_oz.loc[50]['temperature']
```
Let's investigate the relationship of ozone level to wind speed with a scatter plot.
```
""" Format of the scatterplot method is as follows: ax.scatter(x-series, y-series) """
fig, ax = plt.subplots()
"""
ax.scatter() statement creates scatterplot
The 'alpha' parameter controls dot transparency: 1 = solid, <1 = various transparency levels, 0 = no mark
The c parameter designates color of the dots: 'b' stands for blue
"""
ax.scatter(df_oz['wind'], df_oz['ozone'], alpha=0.5, c = 'b')
fig.suptitle('Ozone vs. Wind Speed') # Graph title
ax.xaxis.set_label_text('Wind Speed') # x-axis caption
ax.yaxis.set_label_text('Ozone') # y-axis caption
fig.set_size_inches(7,5)
plt.show()
```
Here's ozone level versus solar radiation. What relationship do you see?
```
""" Format of the scatterplot method is as follows: ax.scatter(x-series, y-series) """
fig, ax = plt.subplots()
"""
ax.scatter() statement creates scatterplot
The 'alpha' parameter controls dot transparency: 1 = solid, <1 = various transparency levels, 0 = no mark
The c parameter designates color of the dots: 'b' stands for blue
"""
ax.scatter(df_oz['radiation'], df_oz['ozone'], alpha=0.5, c = 'b') # the 'alpha' parameter controls dot opacity
fig.suptitle('Ozone vs. Radiation')
ax.xaxis.set_label_text('Radiation (langleys)')
ax.yaxis.set_label_text('Ozone (ppb)')
ax.set_xlim(0,400) # Set min and max for y axis
ax.set_ylim(0,200)
fig.set_size_inches(7,5)
plt.show()
```
## In-Class Examples
```
""" Format of the scatterplot method is as follows: ax.scatter(x-series, y-series) """
fig, ax = plt.subplots()
"""
ax.scatter() statement creates scatterplot
The 'alpha' parameter controls dot transparency: 1 = solid, <1 = various transparency levels, 0 = no mark
The c parameter designates color of the dots: 'b' stands for blue
"""
ax.scatter(df_oz['temperature'], df_oz['ozone'], alpha=0.5, c = 'b') # the 'alpha' parameter controls dot opacity
fig.suptitle('Ozone vs. Temperature')
ax.xaxis.set_label_text('Temperature')
ax.yaxis.set_label_text('Ozone (ppb)')
#ax.set_xlim(0,400) # Set min and max for y axis
ax.set_ylim(0,200)
fig.set_size_inches(7,5)
plt.show()
""" Format of the scatterplot method is as follows: ax.scatter(x-series, y-series) """
fig, ax = plt.subplots()
"""
ax.scatter() statement creates scatterplot
The 'alpha' parameter controls dot transparency: 1 = solid, <1 = various transparency levels, 0 = no mark
The c parameter designates color of the dots: 'b' stands for blue
"""
ax.plot(df_oz.index, df_oz['temperature'], alpha=0.5, c = 'b')
fig.suptitle('Temperature') # Graph title
ax.xaxis.set_label_text('Index') # x-axis caption
ax.yaxis.set_label_text('Temperature') # y-axis caption
fig.set_size_inches(7,5)
plt.show()
""" Format of the scatterplot method is as follows: ax.scatter(x-series, y-series) """
fig, ax = plt.subplots()
"""
ax.scatter() statement creates scatterplot
The 'alpha' parameter controls dot transparency: 1 = solid, <1 = various transparency levels, 0 = no mark
The c parameter designates color of the dots: 'b' stands for blue
"""
ax.plot(df_oz.idexdf_oz['temperature'], alpha=0.5, c = 'b') # the 'alpha' parameter controls dot opacity
fig.suptitle('Temperature')
ax.xaxis.set_label_text('Index')
ax.yaxis.set_label_text('Temperature')
#ax.set_xlim(0,400) # Set min and max for y axis
#ax.set_ylim(0,200)
fig.set_size_inches(7,5)
plt.show()
import matplotlib.pyplot as plt
# Data
bdata = [1.28,1.05,0.6093,0.22195,0.16063,0.1357,0.10226,0.08499,0.06148,0.05022,0.04485,0.02981]
blabels = ['Unemp','Health','Mil.','Interest', 'Veterans','Agri.','Edu','Trans','Housing','Intl','EnergyEnv','Science']
xs = range(len(bdata))
bdata_cum = []
for i in range(len(bdata)):
bdata_cum.append(sum(bdata[0:i+1])/sum(bdata))
fig, ax = plt.subplots(nrows=1,ncols=2)
fig.suptitle('United States Budget Analysis')
# Set bar chart parameters
ax[0].bar(xs,bdata, align='center')
ax[0].set_ylim(0,sum(bdata))
ax[0].set_xticks(xs)
ax[0].set_xticklabels(blabels, rotation = 45)
ax[0].grid(False)
ax[0].tick_params(axis = 'y', which = 'both', direction = 'in', width = 2, color = 'black')
# Set line chart paramters and assign the second y axis
#ax1 = ax.twinx()
ax[1].plot(xs,bdata_cum,color='k')
ax[1].set_ylim(0,1)
ax[1].set_yticklabels(['{:1.1f}%'.format(x*100) for x in ax1.get_yticks()])
ax[1].grid(False)
#fig.set_figwidth(9)
#fig.set_figheight(5)
fig.set_size_inches(9,5)
fig.savefig('Section1a.jpg')
plt.show()
ax[1]
df_oz.head()
fig, ax = plt.subplots()
ax.hist(x = df_oz['temperature'], bins = 20, facecolor='g', alpha=0.75, rwidth=0.75)
fig.suptitle('End of Year test Score Frequency Histogram')
fig.set_size_inches(7,5)
ax.xaxis.set_label_text('End Year Test Score')
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn top x axis tick marks off
ax.yaxis.set_label_text('Frequency of Scores')
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn right y axis tick marks off
#ax.set_xlim(64, 85)
#ax.set_ylim(0, 8)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.grid(False)
```
# Scatterplots with Nonlinear Trendlines
```
import numpy as np
""" Format of the scatterplot method is as follows: plt.scatter(x-series, y-series) """
plt.figure(figsize=(24,5))
plt.scatter(df_oz['wind'], df_oz['ozone'], alpha=0.5, c = 'b')
plt.plot(np.unique(df_oz['wind']), np.poly1d(np.polyfit(df_oz['wind'], df_oz['ozone'],3))(np.unique(df_oz['wind'])))
# alpha is a parameter that controls the transparency of the dots: 1 = solid, <1 = various transparency levels, 0 = no mark
plt.title('Ozone vs. Wind Speed')
plt.xlabel('Wind Speed')
plt.ylabel('Ozone')
plt.show()
```
Change dimensions of the graph to increase the contrast between varying slopes.
```
""" Format of the scatterplot method is as follows: plt.scatter(x-series, y-series) """
plt.figure(figsize=(8,5))
plt.scatter(df_oz['wind'], df_oz['ozone'], alpha=0.5, c='b')
plt.plot(np.unique(df_oz['wind']), np.poly1d(np.polyfit(df_oz['wind'], df_oz['ozone'],3))(np.unique(df_oz['wind'])))
# alpha is a parameter that controls the transparency of the dots: 1 = solid, <1 = various transparency levels, 0 = no mark
plt.title('Ozone vs. Wind Speed')
plt.xlabel('Wind Speed')
plt.ylabel('Ozone')
plt.show()
```
# Distributional Data: Histograms and Bar Charts
Let's look at some distributional data from a database that contains student scores for two tests, one taken at the beginning of the year and one at the end of the year. Each student's scores are recorded along with their isntructor, school number, and course section number. Here are the column names:
```
#df_test = pd.read_csv('D:/TeachingMaterials/BusinessAnalytics/Visualization/MSBAST1DataPrograms\\test.csv')
df_test = pd.read_csv('test.csv')
df_test.head()
df_test.columns.values
```
Here's what the data look like:
```
df_test.head()
```
Before we proceed, let's clean things up a bit by using the StudentIdentifier data as the index. It is unnecessary to have both the (automatic) integer index and the StudentIdentifier.
```
df_test1 = df_test.set_index('StudentIdentifier')
df_test1.head()
```
pandas provides an easy way, using the .unique() method, to find the distinct entries in each column, for example to find what instructors are represented in the database
```
df_test1['InstructorName'].unique()
```
Let's filter the rows, choosing only those rows where Smith is the instructor
```
df_test1.loc[df_test1['InstructorName'] == 'Smith' ]
```
Let's plot a frequency histogram of this data for the year end test scores.
```
df_test1.loc[df_test1['InstructorName'] == 'Smith']['EndYearTestScore']
```
Here's a frequency histogram for Instructor Smith's students at the end of the school year.
```
fig, ax = plt.subplots()
ax.hist(x = df_test1.loc[df_test1['InstructorName'] == 'Smith']['EndYearTestScore'], bins = 20, facecolor='g', alpha=0.75)
fig.suptitle('End of Year test Score Frequency Histogram')
fig.set_size_inches(7,5)
ax.xaxis.set_label_text('End Year Test Score')
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn top x axis tick marks off
ax.yaxis.set_label_text('Frequency of Scores')
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn right y axis tick marks off
ax.set_xlim(64, 85)
ax.set_ylim(0, 8)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.grid(False)
```
A bar chart can be used to create a histogram also, but using this chart type requires that the data be processed into frequency histogram format before plotting.
```
""" Create frequency histogram data from DataFrame """
scores = df_test1.loc[df_test1['InstructorName'] == 'Smith']['EndYearTestScore']
scores = dict(scores.value_counts())
scoresminx = min(scores)
maxx = max(scores)
minx = min(scores)
x = [i for i in range(minx,maxx+1)]
freq = [scores[i] for i in range(minx,maxx+1)]
""" Plot frequency histogram data """
fig, ax = plt.subplots()
ax.bar(x=x,height=freq, color='g', alpha=0.75)
fig.suptitle('End of Year test Score Frequency Histogram')
fig.set_size_inches(7,5)
ax.xaxis.set_label_text('End Year Test Score')
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn top x axis tick marks off
ax.yaxis.set_label_text('Frequency of Scores')
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn right y axis tick marks off
ax.set_xlim(64, 85)
ax.set_ylim(0, 8)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.grid(False)
```
Block of code from teh cell above that creates a dictionary with the frequency histogram data.
```
scores = df_test1.loc[df_test1['InstructorName'] == 'Smith']['EndYearTestScore']
scores = dict(scores.value_counts())
scores
```
Here's one way to show a comparison of Smith's and Green's student scores at the end of the year. Do you like the graph? Is it easy to read? Do you ahve any suggestions for improvement?
```
fig, ax = plt.subplots()
ax.hist(x = df_test1.loc[df_test1['InstructorName'] == 'Smith']['EndYearTestScore'], bins = 20, facecolor='r')
ax.hist(x = df_test1.loc[df_test1['InstructorName'] == 'Green']['EndYearTestScore'], bins = 20, facecolor='g')
fig.suptitle('End of Year test Score Frequency Histogram')
fig.set_size_inches(7,5)
ax.xaxis.set_label_text('End Year Test Score')
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn top x axis tick marks off
ax.yaxis.set_label_text('Frequency of Scores')
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn right y axis tick marks off
ax.set_xlim(64, 85)
ax.set_ylim(0, 8)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.grid(True)
fig, ax = plt.subplots()
ax.hist(x = df_test1.loc[df_test1['InstructorName'] == 'Smith']['EndYearTestScore'], bins = 20, facecolor='r', alpha=0.5)
ax.hist(x = df_test1.loc[df_test1['InstructorName'] == 'Green']['EndYearTestScore'], bins = 20, facecolor='g', alpha=0.5)
fig.suptitle('End of Year test Score Frequency Histogram')
fig.set_size_inches(7,5)
ax.xaxis.set_label_text('End Year Test Score')
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn top x axis tick marks off
ax.yaxis.set_label_text('Frequency of Scores')
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn right y axis tick marks off
ax.set_xlim(64, 85)
ax.set_ylim(0, 8)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.grid(True)
fig, ax = plt.subplots()
n, bins, patches = ax.hist((df_test1.loc[df_test1['InstructorName'] == 'Smith']['EndYearTestScore'], \
df_test1.loc[df_test1['InstructorName'] == 'Green']['EndYearTestScore']), bins = 20, stacked = False)
for i in range(len(patches)):
if i%2 == 0:
plt.setp(patches[i],'facecolor','r')
else:
plt.setp(patches[i],'facecolor','g')
fig.suptitle('End of Year test score Frequency Histogram')
fig.set_size_inches(7,5)
ax.xaxis.set_label_text('End Year Test Score')
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn top x axis tick marks off
ax.yaxis.set_label_text('Frequency of Scores')
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn right y axis tick marks off
ax.set_xlim(64, 85)
ax.set_ylim(0, 8)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.grid(True)
ax.legend(['Smith','Green'], loc=2)
import numpy as np
df_test2 = pd.concat([df_test1.loc[df_test1['InstructorName'] == 'Smith']['EndYearTestScore'],df_test1.loc[df_test1['InstructorName'] == 'Green']['EndYearTestScore']], axis = 1)
df_test2.columns = ['Smith','Green']
df_test1.loc[df_test1['InstructorName'] == 'Green']['EndYearTestScore']
df_test2
```
Here's a boxplot of Smith's end of year student test scores.
```
fig, ax = plt.subplots()
ax.boxplot(df_test1.loc[df_test1['InstructorName'] == 'Smith']['EndYearTestScore'])
ax.yaxis.axes.set_ylim(60,85)
ax.set_xticklabels(['Smith'])
# Reducing clutter
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn top x axis tick marks off
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn right y axis tick marks off
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
```
Code for comparing Smith and Green
```
# Get data
data = [] # create an empty Python list; sublists will be appended for each boxplot
data.append(df_test1.loc[df_test1['InstructorName'] == 'Smith']['EndYearTestScore'])
data.append(df_test1.loc[df_test1['InstructorName'] == 'Green']['EndYearTestScore'])
data_min = min([min(sublist) for sublist in data]) # this and the following 3 lines automatically size the graph to the data
data_max = max([max(sublist) for sublist in data]) # while providing buffer space
fig, ax = plt.subplots()
fig.set_figheight(5)
fig.set_figwidth(7)
fig.add_axes
ax.boxplot(data)
ax.yaxis.axes.set_ylim(data_min - 2, data_max + 2)
ax.set_xticklabels(['Smith','Green'])
ax.set_ylabel('Test Score')
ax.set_xlabel('Instructor')
# Reducing clutter
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn top x axis tick marks off
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn right y axis tick marks off
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
```
Code for comparing Smith, Jones, and Green
```
# Get data
instructors = ['Smith','Green','Jones']
data = [] # create an empty Python list; sublists will be appended for each boxplot
for instructor in instructors:
data.append(df_test1.loc[df_test1['InstructorName'] == instructor]['EndYearTestScore'])
data_min = min([min(sublist) for sublist in data]) # this and the following 3 lines automatically size the graph to the data
data_max = max([max(sublist) for sublist in data]) # while providing buffer space
fig, ax = plt.subplots()
fig.set_figheight(5)
fig.set_figwidth(7)
fig.add_axes
ax.boxplot(data)
ax.yaxis.axes.set_ylim(data_min - 2, data_max + 2)
ax.set_xticklabels(instructors)
ax.set_ylabel('Test Score')
ax.set_xlabel('Instructor')
ax.set_xticks(range(1,len(instructors)+1),instructors)
# Reducing clutter
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn top x axis tick marks off
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn right y axis tick marks off
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
data = [] # create an empty Python list; sublists will be appended for each boxplot
instructors = ['Smith','Green','Jones']
for instructor in instructors:
data.append(list(df_test1.loc[df_test1['InstructorName'] == instructor]['EndYearTestScore']))
plt.boxplot(data)
plt.xticks(range(1,len(instructors)+1),instructors)
data_min = min([min(sublist) for sublist in data]) # this and the following 3 lines automatically size the graph to the data
data_max = max([max(sublist) for sublist in data]) # while providing buffer space
plt.ylim([data_min - 2, data_max + 2])
plt.show()
```
# The seaborn Package
seaborn is written 'on top of' matplotlib and automates many forms of multi-graph figures. Here's is one example where each one-to-one relationships in the four fields in the ozone data are shown via a grid of scatter plots. seaborn in this way takes care of a lot of detailed work that you would need to do if you used matplotlib to construct this graph.
```
import seaborn as sns
sns.set(style="ticks", color_codes=True)
g = sns.pairplot(df_oz)
g.savefig('ozone.jpg')
sns.set(style="whitegrid")
xs = [0,82.0442626658045,164.140532825633,249.101916493353,332.182520454561,416.254670963008,495.330844802651,575.152803567278,653.205144040749,738.002602107511,818.027625347191,898.118246791891,980.079475188722,1061.14169245002,1139.27372549306,1222.46124841828,1300.35875796241,1375.56916894815,1450.58381247379,1528.61926864683,1605.58397275636,1690.73526419816,1765.8607155748,1844.76374052086,1927.68455634244,2007.86892722346,2086.83606915306,2166.71913598734,2251.80135023691,2335.70816516882,2420.79263834332,2499.86354911873,2577.05890165442,2654.19208140119,2730.09283946298,2809.07577275313,2894.08624312766,2969.05670691318,3049.93958564477,3129.75861740531,3206.68277248806,3283.60346903165,3364.58552562287,3449.40362446521,3525.4406380069,3608.13685182861,3690.05929028829,3773.14656467608,3850.22786804579,3930.30737684189,4015.46434790825,4093.50879464788,4172.64841837878,4252.66450124259,4335.57956650011,4420.52164467734,4495.52991250898,4575.39742270308,4650.49642074232,4730.52574677055,4805.68851119975,4880.60019974235,4960.70865715944,5043.67168991259,5118.73724644963,5203.54753389807,5283.38430140739,5359.51383129058,5438.3504224767,5519.19106748314,5597.38879699402]
ys = [69,70.5331166725557,71.5580608629997,71.7925736363559,71.3468902075964,70.8280365103217,70.6414982437025,70.4451053764375,70.3795952412711,69.7645648576213,69.5426154537094,69.3004779067882,68.9026747371149,68.599154906399,68.519069857217,68.3772433121902,69.1486631584102,68.7548264957532,68.3609585242701,68.5575021969504,68.5786288310157,70.1801398605569,69.7914710297875,70.1491959166127,71.3308639026079,71.9361568322099,71.7841530239482,71.5632100706621,70.9532872882161,70.4120650812618,69.7956890967353,69.6193830861858,69.6154149236862,69.6130427165012,69.7102463368457,69.5264209638887,68.9050613087613,69.1051712486311,68.8193037564598,68.6066819000086,68.6092749335593,68.6204785117934,68.3360360051156,68.5812312774717,68.3713770000006,69.5440095449469,70.5260768620176,72.0285425812187,72.0349254824144,71.8158332177273,71.1892654017154,71.1169346409977,70.8948191898231,70.6895661675202,70.2298913978404,69.6255849641529,69.7340391269025,69.5070018796742,69.6617353391113,69.4627544694931,69.630774569752,69.8026157834872,69.5269217792645,69.0549816241028,69.1984712324886,68.3562401007611,68.1422906748129,68.2062726166005,68.0498033914326,68.6018257745021,68.7606159741864]
#import seaborn as sns
sns.set(style="darkgrid", color_codes=True)
tips = sns.load_dataset("tips")
g = sns.jointplot("total_bill", "tip", data=tips, kind="reg",
xlim=(0, 60), ylim=(0, 12), color="r", size=7)
```
# Pareto Diagrams
The next cell shows a Pareto package that is freely available. I will also show my own code below.
```
import matplotlib.pyplot as plt
# Data
bdata = [1.28,1.05,0.6093,0.22195,0.16063,0.1357,0.10226,0.08499,0.06148,0.05022,0.04485,0.02981]
blabels = ['Unemp','Health','Mil.','Interest', 'Veterans','Agri.','Edu','Trans','Housing','Intl','EnergyEnv','Science']
xs = range(len(bdata))
bdata_cum = []
for i in range(len(bdata)):
bdata_cum.append(sum(bdata[0:i+1])/sum(bdata))
fig, ax = plt.subplots()
fig.suptitle('United States Budget Analysis')
# Set bar chart parameters
ax.bar(xs,bdata, align='center')
ax.set_ylim(0,sum(bdata))
ax.set_xticks(xs)
ax.set_xticklabels(blabels, rotation = 45)
ax.grid(False)
ax.tick_params(axis = 'y', which = 'both', direction = 'in', width = 2, color = 'black')
# Set line chart paramters and assign the second y axis
ax1 = ax.twinx()
ax1.plot(xs,bdata_cum,color='k')
ax1.set_ylim(0,1)
ax1.set_yticklabels(['{:1.1f}%'.format(x*100) for x in ax1.get_yticks()])
ax1.grid(False)
#fig.set_figwidth(9)
#fig.set_figheight(5)
fig.set_size_inches(9,5)
fig.savefig('Section1.jpg')
plt.show()
```
In the following code a typical version of a Pareto chart is shown where bars are shown for only the most frequent items.
```
import matplotlib.pyplot as plt
# Data
blabels1 = ['SS','Health','Mil.','Interest', 'Vet.','Agri.','Other']
bindex = 6
bother = sum(bdata[bindex:])
bdata1 = bdata[:bindex] + [bother]
xs = range(len(bdata1))
bdata_cum = []
for i in range(len(bdata1)):
bdata_cum.append(sum(bdata1[0:i+1])/sum(bdata1))
fig, ax = plt.subplots()
fig.set_figwidth(9)
fig.set_figheight(5)
# Bar chart settings
ax.set_xticks(xs)
ax.set_xticklabels(blabels1)
ax.bar(xs,bdata1, align='center')
ax.set_ylim(0,sum(bdata1))
# Line chart settings
ax1 = ax.twinx()
ax1.plot(xs,bdata_cum,color='k')
ax1.set_ylim(0,1)
ax1.set_yticklabels(['{:1.0f}'.format(x*100) + '%' for x in ax1.get_yticks()])
plt.show()
```
# Construction Data
This is the data that you will use for your Storytelling & Visualization aassignment due at the end of the course.
```
import pandas as pd
#df_con = pd.read_csv('C:/Users/jrbrad/CTBA/ConstructionTimeSeriesDataV2.csv')
df_con = pd.read_csv('ConstructionTimeSeriesDataV2.csv')
df_con.head()
df_con['Total Construction']
x = df_con.index
y = df_con['Total Construction']
fig, ax = plt.subplots()
fig.set_size_inches(6,4) # figure must be resized prior to .plot() statement
ax.plot(x,y,label='Total Construction')
ax.set_xlabel('Month',fontsize=14) # Title for the horizontal axis
ax.set_ylabel('Construction Spending',fontsize=14) # Title for the vertical axis
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
#ax.spines['left'].set_edgecolor('b')
#ax.spines['bottom'].set_position(('outward', 10))
plt.axis([0,max(df_con['Month'])+1,0,max(df_con['Total Construction'])*1.05],fontsize=14) # Set ranges of axes
plt.xticks(fontsize = 12)
plt.grid(False)
plt.show()
x = df_con.Month
y = df_con['Total Construction']
y1 = df_con['Private Construction']
plt.plot(x,y,label='Total Construction') # The label parameter is a label for the y axis data that will be used in the legend
plt.plot(x,y1,label='Private Construction')
plt.xlabel('Month') # Title for the horizontal axis
plt.ylabel('Construction Spending') # Title for the vertical axis
plt.axis([x.min(),x.max(),0,1.1*y.max()])
plt.legend()
plt.savefig('sample.jpg')
plt.show()
import matplotlib.pyplot as plt
# Extract data for graphing
x = df_con.Month[0:12]
y1 = df_con['Total Construction'][0:12]
y2 = df_con['Private Construction'][0:12]
# My variables
x_labels = ['','J','F','M','A','M','J','J','A','S','O','N','D']
# Create graph and assign Figure and Axes objects to variables fig and ax
fig, ax = plt.subplots()
# Plot the data and set other Axes attributes
ax.plot(x,y1,label='Total Construction') # Add y1 data to graph and create label for the legend
ax.plot(x,y2,label='Private Construction') # Add y2 data to graph and create label for the legend
ax.spines['right'].set_visible(False) # Remove right spine
ax.spines['top'].set_visible(False) # Remove top spine
ax.legend(loc = 'lower center', prop = {'family':'Times New Roman', 'size':'large'}) # Add legend and format it
ax.set_xlim(0,x.max()+1) # Set min and max for y axis
ax.set_ylim(0,1.1*y1.max()) # Set min and max for y axis
# Set x-axis attributes
ax.xaxis.set_label_text('Month',fontsize = 18, fontname = 'Times New Roman')
ax.xaxis.set_ticks(range(0,13))
ax.xaxis.set_ticklabels(x_labels)
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn tick marks off for top x axis
# Set y-axis attributes: the parameter 'both' refers to both major and minor tick marks
ax.yaxis.set_label_text('Construction Spending',fontsize = 18, fontname = 'Times New Roman') # Title for the vertical axis
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn tick marks off for right y axis
# Set Figure attributes
fig.set_size_inches(7,5) # Set size of figure
fig.suptitle('Private vs. Total Construction',fontsize='xx-large', fontname = 'Times New Roman')
fig.tight_layout() # Helps with formatting and fitting everything into the figure
plt.savefig('sample3.jpg') # Save jpg of figure
plt.show()
# Extract data for graphing
x = df_con.Month[0:12]
y1 = df_con['Total Construction'][0:12]
y2 = df_con['Private Construction'][0:12]
# My variables
x_labels = ['','J','F','M','A','M','J','J','A','S','O','N','D']
# Create graph and assign Figure and Axes objects to variables fig and Axes variables ax1 and ax2
fig, (ax1, ax2) = plt.subplots(1,2,sharey=True) # Create 2 Axes (subplots) and assign to ax1 and ax2 respectively
# Plot the data and set other Axes attributes
ax1.plot(x,y1,label='Total Construction') # Add y1 data to graph and create label for the legend
ax2.plot(x,y2,label='Private Construction') # Add y2 data to graph and create label for the legend
# Set common axes attributes
for ax in fig.axes:
ax.spines['right'].set_visible(False) # Remove right spine
ax.spines['top'].set_visible(False) # Remove top spine
ax.legend(loc = 'upper right', prop = {'family':'Times New Roman', 'size':'large'}) # Add legend and format it
ax.set_xlim(0,x.max()+1) # Set min and max for y axis
ax.set_ylim(0,1.1*y1.max()) # Set min and max for y axis
# Set x-axis attributes
for ax in fig.axes:
ax.xaxis.set_label_text('Month',fontsize = 18)
ax.xaxis.set_ticks(range(0,13))
ax.xaxis.set_ticklabels(x_labels)
ax.xaxis.set_tick_params(which = 'both', top = False, bottom = True, labelbottom = True) # Turn top x axis tick marks off
# Set y-axis attributes: the parameter 'both' refers to both major and minor tick marks
for ax in fig.axes:
ax.yaxis.set_label_text('Construction Spending',fontsize = 18, fontname = 'Times New Roman') # Title for the vertical axis
ax.yaxis.set_tick_params(which = 'both', right = False, left = True, labelleft = True) # Turn right y axis tick marks off
# Set Figure attributes
fig.set_size_inches(12,5) # Set size of figure
fig.tight_layout() # Helps with formatting and fitting everything into the figure
plt.savefig('sample3.jpg')
plt.show()
x = df_con.index
y = df_con['Total Construction']
plt.figure(figsize=(24,4)) # figure must be resized prior to .plot() statement
plt.plot(x,y,label='Total Construction')
plt.xlabel('Month',fontsize=14) # Title for the horizontal axis
plt.ylabel('Construction Spending',fontsize=14) # Title for the vertical axis
plt.axis([0,max(df_con['Month'])+1,0,max(df_con['Total Construction'])*1.05],fontsize=14) # Set ranges of axes
plt.xticks(fontsize = 14)
plt.show()
```
| github_jupyter |
# Lesson 1
In the screencast for this lesson I go through a few scenarios for time series. This notebook contains the code for that with a few little extras! :)
# Setup
```
# !pip install -U tf-nightly-2.0-preview
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
def plot_series(time, series, format="-", start=0, end=None, label=None):
plt.plot(time[start:end], series[start:end], format, label=label)
plt.xlabel("Time")
plt.ylabel("Value")
if label:
plt.legend(fontsize=14)
plt.grid(True)
```
# Trend and Seasonality
```
def trend(time, slope=0):
return slope * time
```
Let's create a time series that just trends upward:
```
time = np.arange(4 * 365 + 1)
baseline = 10
series = trend(time, 0.1)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
```
Now let's generate a time series with a seasonal pattern:
```
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.4,
np.cos(season_time * 2 * np.pi),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
baseline = 10
amplitude = 40
series = seasonality(time, period=365, amplitude=amplitude)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
```
Now let's create a time series with both trend and seasonality:
```
slope = 0.05
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
```
# Noise
In practice few real-life time series have such a smooth signal. They usually have some noise, and the signal-to-noise ratio can sometimes be very low. Let's generate some white noise:
```
def white_noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
noise_level = 5
noise = white_noise(time, noise_level, seed=42)
plt.figure(figsize=(10, 6))
plot_series(time, noise)
plt.show()
```
Now let's add this white noise to the time series:
```
series += noise
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
```
All right, this looks realistic enough for now. Let's try to forecast it. We will split it into two periods: the training period and the validation period (in many cases, you would also want to have a test period). The split will be at time step 1000.
```
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
def autocorrelation(time, amplitude, seed=None):
rnd = np.random.RandomState(seed)
φ1 = 0.5
φ2 = -0.1
ar = rnd.randn(len(time) + 50)
ar[:50] = 100
for step in range(50, len(time) + 50):
ar[step] += φ1 * ar[step - 50]
ar[step] += φ2 * ar[step - 33]
return ar[50:] * amplitude
series = autocorrelation(time, 10, seed=42)
plot_series(time[:200], series[:200])
plt.show()
def autocorrelation(time, amplitude, seed=None):
rnd = np.random.RandomState(seed)
φ = 0.8
ar = rnd.randn(len(time) + 1)
for step in range(1, len(time) + 1):
ar[step] += φ * ar[step - 1]
return ar[1:] * amplitude
series = autocorrelation(time, 10, seed=42)
plot_series(time[:200], series[:200])
plt.show()
series = autocorrelation(time, 10, seed=42) + trend(time, 2)
plot_series(time[:200], series[:200])
plt.show()
series = autocorrelation(time, 10, seed=42) + seasonality(time, period=50, amplitude=150) + trend(time, 2)
plot_series(time[:200], series[:200])
plt.show()
series = autocorrelation(time, 10, seed=42) + seasonality(time, period=50, amplitude=150) + trend(time, 2)
series2 = autocorrelation(time, 5, seed=42) + seasonality(time, period=50, amplitude=2) + trend(time, -1) + 550
series[200:] = series2[200:]
#series += noise(time, 30)
plot_series(time[:300], series[:300])
plt.show()
def impulses(time, num_impulses, amplitude=1, seed=None):
rnd = np.random.RandomState(seed)
impulse_indices = rnd.randint(len(time), size=10)
series = np.zeros(len(time))
for index in impulse_indices:
series[index] += rnd.rand() * amplitude
return series
series = impulses(time, 10, seed=42)
plot_series(time, series)
plt.show()
def autocorrelation(source, φs):
ar = source.copy()
max_lag = len(φs)
for step, value in enumerate(source):
for lag, φ in φs.items():
if step - lag > 0:
ar[step] += φ * ar[step - lag]
return ar
signal = impulses(time, 10, seed=42)
series = autocorrelation(signal, {1: 0.99})
plot_series(time, series)
plt.plot(time, signal, "k-")
plt.show()
signal = impulses(time, 10, seed=42)
series = autocorrelation(signal, {1: 0.70, 50: 0.2})
plot_series(time, series)
plt.plot(time, signal, "k-")
plt.show()
series_diff1 = series[1:] - series[:-1]
plot_series(time[1:], series_diff1)
from pandas.plotting import autocorrelation_plot
autocorrelation_plot(series)
from statsmodels.tsa.arima_model import ARIMA
model = ARIMA(series, order=(5, 1, 0))
model_fit = model.fit(disp=0)
print(model_fit.summary())
df = pd.read_csv("sunspots.csv", parse_dates=["Date"], index_col="Date")
series = df["Monthly Mean Total Sunspot Number"].asfreq("1M")
series.head()
series.plot(figsize=(12, 5))
series["1995-01-01":].plot()
series.diff(1).plot()
plt.axis([0, 100, -50, 50])
from pandas.plotting import autocorrelation_plot
autocorrelation_plot(series)
autocorrelation_plot(series.diff(1)[1:])
autocorrelation_plot(series.diff(1)[1:].diff(11 * 12)[11*12+1:])
plt.axis([0, 500, -0.1, 0.1])
autocorrelation_plot(series.diff(1)[1:])
plt.axis([0, 50, -0.1, 0.1])
116.7 - 104.3
[series.autocorr(lag) for lag in range(1, 50)]
pd.read_csv(filepath_or_buffer, sep=',', delimiter=None, header='infer', names=None, index_col=None, usecols=None, squeeze=False, prefix=None, mangle_dupe_cols=True, dtype=None, engine=None, converters=None, true_values=None, false_values=None, skipinitialspace=False, skiprows=None, skipfooter=0, nrows=None, na_values=None, keep_default_na=True, na_filter=True, verbose=False, skip_blank_lines=True, parse_dates=False, infer_datetime_format=False, keep_date_col=False, date_parser=None, dayfirst=False, iterator=False, chunksize=None, compression='infer', thousands=None, decimal=b'.', lineterminator=None, quotechar='"', quoting=0, doublequote=True, escapechar=None, comment=None, encoding=None, dialect=None, tupleize_cols=None, error_bad_lines=True, warn_bad_lines=True, delim_whitespace=False, low_memory=True, memory_map=False, float_precision=None)
Read a comma-separated values (csv) file into DataFrame.
from pandas.plotting import autocorrelation_plot
series_diff = series
for lag in range(50):
series_diff = series_diff[1:] - series_diff[:-1]
autocorrelation_plot(series_diff)
import pandas as pd
series_diff1 = pd.Series(series[1:] - series[:-1])
autocorrs = [series_diff1.autocorr(lag) for lag in range(1, 60)]
plt.plot(autocorrs)
plt.show()
```
| github_jupyter |
Engineering Features and Preparing Data to Use in Modeling
Objective
Now that you understand your data, your next step is to engineer features and prepare two sets of data to use for modeling: one that contains only numeric values, and one that contains numeric and categorical values, so you can leverage the advantage of H2O Random Forests over scikit-learn Random Forests.
Workflow
Load your data from Milestone 1, and make sure it looks OK.
Engineer and clean features from your string and datetime columns and examine their relationship to the target column “LOAN_DEFAULT.”
Parse a few columns that have date spans and are strings (e.g., AVERAGE_ACCT_AGE_MONTHS) for the appropriate data type (e.g., integer, datetime) using string methods or regex (the re library).
Examine the relationship of new features to the target using appropriate methods such as Pearson correlation or mutual information.
Create a copy of the DataFrame for use in scikit-learn to one-hot-encode (OHE) categorical columns and convert all columns to numeric types.
Perform any missing value imputation .
Take care to simplify the data if necessary. For example, if more than 50% of the data can be represented with 3 categories, you might group all your other data into an ”other” category to simplify it.
If unique values in a column/feature are too spread out (e.g., each unique value is less than 1% of the total), it won’t work well for OHE. But if some unique values in the column make up ~5% of the total, you can use the risk ratio to evaluate if that column might be useful.
Save the OHE and H2O DataFrames to disk (e.g., as CSV files) for your next step; also save about 10% or less of down-sampled versions of the data for developing your solution to Milestone 3.
If you find the .apply() method in pandas is slow, you can parallelize it using swifter.
```
import pandas as pd
from pandas_profiling import ProfileReport
from sklearn.model_selection import train_test_split
import pathlib
import h2o
df = pd.read_csv('loan_data_cleaned.csv')
df.head().T
employ_df = pd.get_dummies(df['employment_type'])
employ_df.columns = [col.lower().replace(' ', '_') for col in employ_df.columns]
df = pd.merge(df, employ_df, left_index=True, right_index=True)
del df['employment_type']
```
# Impute missing values
Should some of these values be normalized at all??? I'm unclear.
```
# Treat 0s As missing as a separate variable, not sure if these should be imputed
df[df['perform_cns_score'] != 0]['perform_cns_score'].median()
df['perform_cns_score_cat'] = df['perform_cns_score']
df['missing_cns_score'] = df['perform_cns_score'] == 0
df.loc[
df['missing_cns_score'] == 1,
'perform_cns_score'
] = df[df['perform_cns_score'] != 0]['perform_cns_score'].median()
df['perform_cns_score'].hist()
# profile = ProfileReport(df, explorative=True)
# profile.to_file('loan_data.html')
```
# Correlations
```
df.corrwith(df['loan_default'])
# correlations too low
del df['sec_instal_amt']
del df['sec_current_balance']
```
# Sample Data
```
development_df, validation_df = train_test_split(
df,
train_size=0.25,
stratify=df['loan_default'],
random_state=42
)
training_df, hyperparameter_df = train_test_split(
development_df,
train_size=0.05,
stratify=development_df['loan_default'],
random_state=42
)
```
# Output as Dataframe and H2O
```
datadir = pathlib.Path('data')
datadir.mkdir(exist_ok=True)
training_df.drop(columns='perform_cns_score_cat').to_csv(datadir / 'training.ohe.csv', index=False)
hyperparameter_df.drop(columns='perform_cns_score_cat').to_csv(datadir / 'hyperparameter.ohe.csv', index=False)
validation_df.drop(columns='perform_cns_score_cat').to_csv(datadir / 'validation.ohe.csv', index=False)
h2o.init()
h2o.export_file(h2o.H2OFrame(training_df), str(datadir / 'training.h2o'))
h2o.export_file(h2o.H2OFrame(hyperparameter_df), str(datadir / 'hyperparameter.h2o'))
h2o.export_file(h2o.H2OFrame(validation_df), str(datadir / 'validation.h2o'))
h2o.shutdown()
```
| github_jupyter |
# `rlplay`-ing around with Policy Gradients
```
import torch
import numpy
import matplotlib.pyplot as plt
%matplotlib inline
import gym
# hotfix for gym's unresponsive viz (spawns gl threads!)
import rlplay.utils.integration.gym
```
See example.ipynb for the overview of `rlplay`
<br>
## Sophisticated CartPole with PG
### The environment
The environment factory
```
from rlplay.zoo.env import NarrowPath
class FP32Observation(gym.ObservationWrapper):
def observation(self, observation):
obs = observation.astype(numpy.float32)
obs[0] = 0. # mask the position info
return obs
# def step(self, action):
# obs, reward, done, info = super().step(action)
# reward -= abs(obs[1]) / 10 # punish for non-zero speed
# return obs, reward, done, info
class OneHotObservation(gym.ObservationWrapper):
def observation(self, observation):
return numpy.eye(1, self.env.observation_space.n,
k=observation, dtype=numpy.float32)[0]
def base_factory(seed=None):
# return gym.make("LunarLander-v2")
return FP32Observation(gym.make("CartPole-v0").unwrapped)
# return OneHotObservation(NarrowPath())
```
<br>
### the Actor
A procedure and a layer, which converts the input integer data into its
little-endian binary representation as float $\{0, 1\}^m$ vectors.
```
def onehotbits(input, n_bits=63, dtype=torch.float):
"""Encode integers to fixed-width binary floating point vectors"""
assert not input.dtype.is_floating_point
assert 0 < n_bits < 64 # torch.int64 is signed, so 64-1 bits max
# n_bits = {torch.int64: 63, torch.int32: 31, torch.int16: 15, torch.int8 : 7}
# get mask of set bits
pow2 = torch.tensor([1 << j for j in range(n_bits)]).to(input.device)
x = input.unsqueeze(-1).bitwise_and(pow2).to(bool)
# upcast bool to float to get one-hot
return x.to(dtype)
class OneHotBits(torch.nn.Module):
def __init__(self, n_bits=63, dtype=torch.float):
assert 1 <= n_bits < 64
super().__init__()
self.n_bits, self.dtype = n_bits, dtype
def forward(self, input):
return onehotbits(input, n_bits=self.n_bits, dtype=self.dtype)
```
A special module dictionary, which applies itself to the input dict of tensors
```
from typing import Optional, Mapping
from torch.nn import Module, ModuleDict as BaseModuleDict
class ModuleDict(BaseModuleDict):
"""The ModuleDict, that applies itself to the input dicts."""
def __init__(
self,
modules: Optional[Mapping[str, Module]] = None,
dim: Optional[int]=-1
) -> None:
super().__init__(modules)
self.dim = dim
def forward(self, input):
# enforce concatenation in the order of the declaration in __init__
return torch.cat([
m(input[k]) for k, m in self.items()
], dim=self.dim)
```
An $\ell_2$ normalization layer.
```
from torch.nn.functional import normalize
class Normalize(torch.nn.Module):
def __init__(self, dim=-1):
super().__init__()
self.dim = dim
def forward(self, input):
return normalize(input, dim=self.dim)
```
A more sophisticated policy learner
```
from rlplay.engine import BaseActorModule
from rlplay.utils.common import multinomial
from torch.nn import Sequential, Linear, ReLU, LogSoftmax
class CartPoleActor(BaseActorModule):
def __init__(self, lstm='none'):
assert lstm in ('none', 'loop', 'cudnn')
super().__init__()
self.use_lstm = self.use_cudnn = False
# blend the policy with a uniform distribution, determined by
# the exploration epsilon. We update it in the actor clones via a buffer
# self.register_buffer('epsilon', torch.tensor(epsilon))
# XXX isn't the stochastic policy random enough by itself?
self.baseline = Sequential(
Linear(4, 20),
ReLU(),
Linear(20, 1),
)
self.policy = Sequential(
Linear(4, 20),
ReLU(),
Linear(20, 2),
LogSoftmax(dim=-1),
)
def forward(self, obs, act, rew, fin, *, hx=None, stepno=None, virtual=False):
# value must not have any trailing dims, i.e. T x B
logits = self.policy(obs)
value = self.baseline(obs).squeeze(-1)
if not self.training:
actions = logits.argmax(dim=-1)
else:
actions = multinomial(logits.detach().exp())
return actions, (), dict(value=value, logits=logits)
```
<br>
### PPO/GAE A2C and V-trace A2C algos
Service functions for the algorithms
```
from plyr import apply, suply, xgetitem
def timeshift(state, *, shift=1):
"""Get current and shifted slices of nested objects."""
# use `xgetitem` to let None through
# XXX `curr[t]` = (x_t, a_{t-1}, r_t, d_t), t=0..T-H
curr = suply(xgetitem, state, index=slice(None, -shift))
# XXX `next[t]` = (x_{t+H}, a_{t+H-1}, r_{t+H}, d_{t+H}), t=0..T-H
next = suply(xgetitem, state, index=slice(shift, None))
return curr, next
```
The Advantage Actor-Critic algo
```
import torch.nn.functional as F
from rlplay.algo.returns import pyt_gae, pyt_returns, pyt_multistep
# @torch.enable_grad()
def a2c(
fragment, module, *, gamma=0.99, gae=1., ppo=0.,
C_entropy=1e-2, C_value=0.5, c_rho=1.0, multistep=0,
):
r"""The Advantage Actor-Critic algorithm (importance-weighted off-policy).
Close to REINFORCE, but uses separate baseline value estimate to compute
advantages in the policy gradient:
$$
\nabla_\theta J(s_t)
= \mathbb{E}_{a \sim \beta(a\mid s_t)}
\frac{\pi(a\mid s_t)}{\beta(a\mid s_t)}
\bigl( r_{t+1} + \gamma G_{t+1} - v(s_t) \bigr)
\nabla_\theta \log \pi(a\mid s_t)
\,, $$
where the critic estimates the state's value under the current policy
$$
v(s_t)
\approx \mathbb{E}_{\pi_{\geq t}}
G_t(a_t, s_{t+1}, a_{t+1}, ... \mid s_t)
\,. $$
"""
state, state_next = timeshift(fragment.state)
# REACT: (state[t], h_t) \to (\hat{a}_t, h_{t+1}, \hat{A}_t)
_, _, info = module(
state.obs, state.act, state.rew, state.fin,
hx=fragment.hx, stepno=state.stepno)
# info['value'] = V(`.state[t]`)
# <<-->> v(x_t)
# \approx \mathbb{E}( G_t \mid x_t)
# \approx \mathbb{E}( r_{t+1} + \gamma r_{t+2} + ... \mid x_t)
# <<-->> npv(`.state[t+1:]`)
# info['logits'] = \log \pi(... | .state[t] )
# <<-->> \log \pi( \cdot \mid x_t)
# `.actor[t]` is actor's extra info in reaction to `.state[t]`, t=0..T
bootstrap = fragment.actor['value'][-1]
# `bootstrap` <<-->> `.value[-1]` = V(`.state[-1]`)
# XXX post-mul by `1 - \gamma` fails to train, but seems appropriate
# for the continuation/survival interpretation of the discount factor.
# <<-- but who says this is a good interpretation?
# ret.mul_(1 - gamma)
# \pi is the target policy, \mu is the behaviour policy
log_pi, log_mu = info['logits'], fragment.actor['logits']
# Future rewards after `.state[t]` are recorded in `.state[t+1:]`
# G_t <<-->> ret[t] = rew[t] + gamma * (1 - fin[t]) * (ret[t+1] or bootstrap)
if multistep > 0:
ret = pyt_multistep(state_next.rew, state_next.fin,
info['value'].detach(),
gamma=gamma, n_lookahead=multistep,
bootstrap=bootstrap.unsqueeze(0))
else:
ret = pyt_returns(state_next.rew, state_next.fin,
gamma=gamma, bootstrap=bootstrap)
# the critic's mse score (min)
# \frac1{2 T} \sum_t (G_t - v(s_t))^2
value = info['value']
critic_mse = F.mse_loss(value, ret, reduction='mean') / 2
# v(x_t) \approx \mathbb{E}( G_t \mid x_t )
# \approx G_t (one-point estimate)
# <<-->> ret[t]
# compute the advantages $G_t - v(s_t)$
# or GAE [Schulman et al. (2016)](http://arxiv.org/abs/1506.02438)
# XXX sec 6.1 in the GAE paper uses V from the `current` value
# network, not the one used during the rollout.
# value = fragment.actor['value'][:-1]
if gae < 1.:
# the positional arguments are $r_{t+1}$, $d_{t+1}$, and $v(s_t)$,
# respectively, for $t=0..T-1$. The bootstrap is $v(S_T)$ from
# the rollout.
adv = pyt_gae(state_next.rew, state_next.fin, value.detach(),
gamma=gamma, C=gae, bootstrap=bootstrap)
else:
adv = ret.sub(value.detach())
# adv.sub_(adv.mean())
# adv.div_(adv.std(dim=0))
# Assume `.act` is unstructured: `act[t]` = a_{t+1} -->> T x B x 1
act = state_next.act.unsqueeze(-1) # actions taken during the rollout
# the importance weights
log_pi_a = log_pi.gather(-1, act).squeeze(-1)
log_mu_a = log_mu.gather(-1, act).squeeze(-1)
# the policy surrogate score (max)
if ppo > 0:
# the PPO loss is the properly clipped rho times the advantage
ratio = log_pi_a.sub(log_mu_a).exp()
a2c_score = torch.min(
ratio * adv,
ratio.clamp(1. - ppo, 1. + ppo) * adv
).mean()
else:
# \exp{- ( \log \mu - \log \pi )}, evaluated at $a_t \mid z_t$
rho = log_mu_a.sub_(log_pi_a.detach()).neg_()\
.exp_().clamp_(max=c_rho)
# \frac1T \sum_t \rho_t (G_t - v_t) \log \pi(a_t \mid z_t)
a2c_score = log_pi_a.mul(adv.mul_(rho)).mean()
# the policy's neg-entropy score (min)
# - H(\pi(•\mid s)) = - (-1) \sum_a \pi(a\mid s) \log \pi(a\mid s)
f_min = torch.finfo(log_pi.dtype).min
negentropy = log_pi.exp().mul(log_pi.clamp(min=f_min)).sum(dim=-1).mean()
# breakpoint()
# maximize the entropy and the reinforce score, minimize the critic loss
objective = C_entropy * negentropy + C_value * critic_mse - a2c_score
return objective.mean(), dict(
entropy=-float(negentropy),
policy_score=float(a2c_score),
value_loss=float(critic_mse),
)
```
A couple of three things:
* a2c is on-policy and no importance weight could change this!
* L72-80: [stable_baselines3](./common/on_policy_algorithm.py#L183-192)
and [rlpyt](./algos/pg/base.py#L49-58) use rollout data, when computing the GAE
* L61-62: [stable_baselines3](./stable_baselines3/a2c/a2c.py#L147-156) uses `vf_coef=0.5`,
and **unhalved** `F.mse-loss`, while [rlpyt](./rlpyt/rlpyt/algos/pg/a2c.py#L93-94)
uses `value_loss_coeff=0.5`, and **halved** $\ell_2$ loss!
The off-policy actor-critic algorithm for the learner, called V-trace,
from [Espeholt et al. (2018)](http://proceedings.mlr.press/v80/espeholt18a.html).
```
from rlplay.algo.returns import pyt_vtrace
# @torch.enable_grad()
def vtrace(fragment, module, *, gamma=0.99, C_entropy=1e-2, C_value=0.5):
# REACT: (state[t], h_t) \to (\hat{a}_t, h_{t+1}, \hat{A}_t)
_, _, info = module(
fragment.state.obs, fragment.state.act,
fragment.state.rew, fragment.state.fin,
hx=fragment.hx, stepno=fragment.state.stepno)
# Assume `.act` is unstructured: `act[t]` = a_{t+1} -->> T x B x 1
state, state_next = timeshift(fragment.state)
act = state_next.act.unsqueeze(-1) # actions taken during the rollout
# \pi is the target policy, \mu is the behaviour policy (T+1 x B x ...)
log_pi, log_mu = info['logits'], fragment.actor['logits']
# the importance weights
log_pi_a = log_pi.gather(-1, act).squeeze(-1)
log_mu_a = log_mu.gather(-1, act).squeeze(-1)
log_rho = log_mu_a.sub_(log_pi_a.detach()).neg_()
# `.actor[t]` is actor's extra info in reaction to `.state[t]`, t=0..T
val = fragment.actor['value'] # info['value'].detach()
# XXX Although Esperholt et al. (2018, sec.~4.2) use the value estimate of
# the rollout policy for the V-trace target in eq. (1), it makes more sense
# to use the estimates of the current policy, as has been done in monobeast.
# https://hackernoon.com/intuitive-rl-intro-to-advantage-actor-critic-a2c-4ff545978752
val, bootstrap = val[:-1], val[-1]
target = pyt_vtrace(state_next.rew, state_next.fin, val,
gamma=gamma, bootstrap=bootstrap,
omega=log_rho, r_bar=1., c_bar=1.)
# the critic's mse score against v-trace targets (min)
critic_mse = F.mse_loss(info['value'][:-1], target, reduction='mean') / 2
# \delta_t = r_{t+1} + \gamma \nu(s_{t+1}) 1_{\neg d_{t+1}} - v(s_t)
adv = torch.empty_like(state_next.rew).copy_(bootstrap)
adv[:-1].copy_(target[1:]) # copy the v-trace targets \nu(s_{t+1})
adv.masked_fill_(state_next.fin, 0.).mul_(gamma)
adv.add_(state_next.rew).sub_(val)
# XXX note `val` here, not `target`! see sec.~4.2 in (Esperholt et al.; 2018)
# the policy surrogate score (max)
# \rho_t = \min\{ \bar{\rho}, \frac{\pi_t(a_t)}{\mu_t(a_t)} \}
rho = log_rho.exp_().clamp_(max=1.)
vtrace_score = log_pi_a.mul(adv.mul_(rho)).mean()
# the policy's neg-entropy score (min)
f_min = torch.finfo(log_pi.dtype).min
negentropy = log_pi.exp().mul(log_pi.clamp(min=f_min)).sum(dim=-1).mean()
# maximize the entropy and the reinforce score, minimize the critic loss
objective = C_entropy * negentropy + C_value * critic_mse - vtrace_score
return objective.mean(), dict(
entropy=-float(negentropy),
policy_score=float(vtrace_score),
value_loss=float(critic_mse),
)
```
<br>
### Run!
Initialize the learner and the environment factories
```
from functools import partial
factory_eval = partial(base_factory)
factory = partial(base_factory)
learner = CartPoleActor(lstm='none')
learner.train()
device_ = torch.device('cpu') # torch.device('cuda:0')
learner.to(device=device_)
optim = torch.optim.Adam(learner.parameters(), lr=1e-3)
```
Initialize the sampler
```
T, B = 25, 4
sticky = learner.use_cudnn
```
```
from rlplay.engine.rollout import multi
batchit = multi.rollout(
factory,
learner,
n_steps=T,
n_actors=6,
n_per_actor=B,
n_buffers=15,
n_per_batch=2,
sticky=sticky,
pinned=False,
clone=True,
close=False,
device=device_,
start_method='fork', # fork in notebook for macos, spawn in linux
)
```
A generator of evaluation rewards
```
from rlplay.engine.rollout.evaluate import evaluate
test_it = evaluate(factory_eval, learner, n_envs=4, n_steps=500,
clone=False, device=device_, start_method='fork')
```
Implement your favourite training method
```
n_epochs = 100
use_vtrace = True
# gamma, gae, ppo = 0.99, 0.92, 0.2
gamma, gae, ppo, multistep = 0.99, 1., 0.2, 0
import tqdm
from torch.nn.utils import clip_grad_norm_
torch.set_num_threads(1)
losses, rewards = [], []
for epoch in tqdm.tqdm(range(n_epochs)):
for j, batch in zip(range(100), batchit):
if use_vtrace:
loss, info = vtrace(batch, learner, gamma=gamma)
else:
loss, info = a2c(batch, learner, gamma=gamma, gae=gae, ppo=ppo, multistep=multistep)
optim.zero_grad()
loss.backward()
grad = clip_grad_norm_(learner.parameters(), max_norm=1.0)
optim.step()
losses.append(dict(
loss=float(loss), grad=float(grad), **info
))
# fetch the evaluation results lagged by one inner loop!
rewards.append(next(test_it))
# close the generators
batchit.close()
test_it.close()
```
<br>
```
def collate(records):
"""collate identically keyed dicts"""
out, n_records = {}, 0
for record in records:
for k, v in record.items():
out.setdefault(k, []).append(v)
return out
data = {k: numpy.array(v) for k, v in collate(losses).items()}
if 'value_loss' in data:
plt.semilogy(data['value_loss'])
if 'entropy' in data:
plt.plot(data['entropy'])
if 'policy_score' in data:
plt.plot(data['policy_score'])
plt.semilogy(data['grad'])
rewards = numpy.stack(rewards, axis=0)
rewards
m, s = numpy.median(rewards, axis=-1), rewards.std(axis=-1)
fi, ax = plt.subplots(1, 1, figsize=(4, 2), dpi=300)
ax.plot(numpy.mean(rewards, axis=-1))
ax.plot(numpy.median(rewards, axis=-1))
ax.plot(numpy.min(rewards, axis=-1))
ax.plot(numpy.std(rewards, axis=-1))
# ax.plot(m+s * 1.96)
# ax.plot(m-s * 1.96)
plt.show()
```
<br>
The ultimate evaluation run
```
from rlplay.engine import core
with factory_eval() as env:
learner.eval()
eval_rewards, info = core.evaluate([
env
], learner, render=True, n_steps=1e4, device=device_)
print(sum(eval_rewards))
```
<br>
Let's analyze the performance
```
from rlplay.algo.returns import npy_returns, npy_deltas
td_target = eval_rewards + gamma * info['value'][1:]
td_error = td_target - info['value'][:-1]
# td_error = npy_deltas(
# eval_rewards, numpy.zeros_like(eval_rewards, dtype=bool), info['value'][:-1],
# gamma=gamma, bootstrap=info['value'][-1])
fig, ax = plt.subplots(1, 1, figsize=(4, 2), dpi=300)
ax.semilogy(abs(td_error) / abs(td_target))
ax.set_title('relative td(1)-error');
from rlplay.algo.returns import npy_returns, npy_deltas
# plt.plot(
# npy_returns(eval_rewards, numpy.zeros_like(eval_rewards, dtype=bool),
# gamma=gamma, bootstrap=info['value'][-1]))
fig, ax = plt.subplots(1, 1, figsize=(4, 2), dpi=300)
ax.plot(info['value'])
ax.axhline(1 / (1 - gamma), c='k', alpha=0.5, lw=1);
import math
from scipy.special import softmax, expit, entr
*head, n_actions = info['logits'].shape
proba = softmax(info['logits'], axis=-1)
fig, ax = plt.subplots(1, 1, figsize=(4, 2), dpi=300)
ax.plot(entr(proba).sum(-1)[:, 0])
ax.axhline(math.log(n_actions), c='k', alpha=0.5, lw=1);
fig, ax = plt.subplots(1, 1, figsize=(4, 2), dpi=300)
ax.hist(info['logits'][..., 1] - info['logits'][..., 0], bins=51); # log-ratio
```
<br>
```
assert False
```
<br>
### Other agents
An agent that uses other inputs, beside `obs`.
```
class CartPoleActor(BaseActorModule):
def __init__(self, epsilon=0.1, lstm='none'):
assert lstm in ('none', 'loop', 'cudnn')
super().__init__()
self.use_lstm = (lstm != 'none')
self.use_cudnn = (lstm == 'cudnn')
# for updating the exploration epsilon in the actor clones
self.register_buffer('epsilon', torch.tensor(epsilon))
# the features
n_output_dim = dict(obs=64, act=8, stepno=0)
self.features = torch.nn.Sequential(
ModuleDict(dict(
obs=Linear(4, n_output_dim['obs']),
act=Embedding(2, n_output_dim['act']),
stepno=Sequential(
OneHotBits(32),
Linear(32, n_output_dim['stepno']),
),
)),
ReLU(),
)
# the core
n_features = sum(n_output_dim.values())
if self.use_lstm:
self.core = LSTM(n_features, 64, 1)
else:
self.core = Sequential(
Linear(n_features, 64, bias=True),
ReLU(),
)
# the rest of the actor's model
self.baseline = Linear(64, 1)
self.policy = Sequential(
Linear(64, 2),
LogSoftmax(dim=-1),
)
def forward(self, obs, act, rew, fin, *, hx=None, stepno=None, virtual=False):
# Everything is [T x B x ...]
input = self.features(locals())
# `input` is T x B x F, `hx` is either `None`, or a proper recurrent state
n_steps, n_envs, *_ = fin.shape
if not self.use_lstm:
# update `hx` into an empty container
out, hx = self.core(input), ()
elif not self.use_cudnn:
outputs = []
for x, m in zip(input.unsqueeze(1), ~fin.unsqueeze(-1)):
# `m` indicates if NO reset took place, otherwise
# multiply by zero to stop the grads
if hx is not None:
hx = suply(m.mul, hx)
# one LSTM step [1 x B x ...]
output, hx = self.core(x, hx)
outputs.append(output)
# compile the output
out = torch.cat(outputs, dim=0)
else:
# sequence padding (MUST have sampling with `sticky=True`)
if n_steps > 1:
lengths = 1 + (~fin[1:]).sum(0).cpu()
input = pack_padded_sequence(input, lengths, enforce_sorted=False)
out, hx = self.core(input, hx)
if n_steps > 1:
out, lens = pad_packed_sequence(
out, batch_first=False, total_length=n_steps)
# apply relu after the core and get the policy
logits = self.policy(out)
# value must not have any trailing dims, i.e. T x B
value = self.baseline(out).squeeze(-1)
if not self.training:
actions = logits.argmax(dim=-1)
else:
# blend the policy with a uniform distribution
prob = logits.detach().exp().mul_(1 - self.epsilon)
prob.add_(self.epsilon / logits.shape[-1])
actions = multinomial(prob)
return actions, hx, dict(value=value, logits=logits)
```
A non-recurrent actor with features shared between the policy and the baseline.
```
class CartPoleActor(BaseActorModule):
def __init__(self, epsilon=0.1, lstm='none'):
assert lstm in ('none', 'loop', 'cudnn')
super().__init__()
self.use_lstm = self.use_cudnn = False
# for updating the exploration epsilon in the actor clones
self.register_buffer('epsilon', torch.tensor(epsilon))
# the features
self.features = Sequential(
Linear(4, 20),
ReLU(),
)
self.baseline = Linear(20, 1)
self.policy = Sequential(
Linear(20, 2),
LogSoftmax(dim=-1),
)
def forward(self, obs, act, rew, fin, *, hx=None, stepno=None, virtual=False):
x = self.features(obs)
# value must not have any trailing dims, i.e. T x B
logits = self.policy(x)
value = self.baseline(x).squeeze(-1)
if not self.training:
actions = logits.argmax(dim=-1)
else:
# blend the policy with a uniform distribution
prob = logits.detach().exp().mul_(1 - self.epsilon)
prob.add_(self.epsilon / logits.shape[-1])
actions = multinomial(prob)
return actions, (), dict(value=value, logits=logits)
```
<br>
```
# stepno = batch.state.stepno
stepno = torch.arange(256)
with torch.no_grad():
out = learner.features[0]['stepno'](stepno)
out = F.linear(F.relu(out), learner.core[1].weight[:, -8:],
bias=learner.core[1].bias)
# out = F.linear(F.relu(out), learner.core.weight_ih_l0[:, -8:],
# bias=learner.core.bias_ih_l0)
# out = F.relu(out)
fig, axes = plt.subplots(3, 3, figsize=(8, 8), dpi=200,
sharex=True, sharey=True)
for j, ax in zip(range(out.shape[1]), axes.flat):
ax.plot(out[:, j], lw=1)
fig.tight_layout(pad=0, h_pad=0, w_pad=0)
with torch.no_grad():
plt.imshow(abs(learner.core[1].weight[:, -8:]).T)
lin = learner.features.stepno[1]
with torch.no_grad():
plt.imshow(abs(lin.weight))
```
| github_jupyter |
```
import numpy as np
from LSTM_Learning_Lib import Model
from FeatureSetCalculation_Lib import ComputeMultiLevelLogsig1dBM
import matplotlib.pyplot as plt
import time
from sklearn.model_selection import ParameterGrid
from sklearn import preprocessing
import random
from GetSeqMnistData import GetSeqPenandCalLogSig, GetSeqPenDigit,GetSeqPenNormCalLogSig,GetSeqPenNorm
```
## Hyperparameters
```
# Parameters grid
param_grid = {'deg_of_sig': [2,3,4,5,6], 'number_of_segment': [8],
'learning_rate': [0.001]}
Param = list(ParameterGrid(param_grid))
# Parameters
training_iters = 60
batch_size = 200
sig_comp_time = []
test_result = []
test_time = []
```
## train RNN with different feature sets
```
for i in range(np.size(Param)):
# Raw data feature set generator
if Param[i]['deg_of_sig']==0:
start = time.time()
train_X, train_Y = GetSeqPenDigit('pendigits-orig.tra.txt')
test_X, test_Y = GetSeqPenDigit('pendigits-orig.tes.txt')
trainsampleClip = len(train_Y)
testsampleClip = len(test_Y)
max_interval = 0
for j in range(trainsampleClip):
if max_interval < len(train_X[j]):
max_interval = len(train_X[j])
for j in range(testsampleClip):
if max_interval < len(test_X[j]):
max_interval = len(test_X[j])
n_input = int(max_interval/Param[i]['number_of_segment'])+1
if n_input % 2 != 0:
n_input += 1
train_data = np.zeros((trainsampleClip, n_input*Param[i]['number_of_segment']))
test_data = np.zeros((testsampleClip, n_input*Param[i]['number_of_segment']))
for sn in range(trainsampleClip):
tmplen = len(train_X[sn])
train_data[sn, :tmplen] = train_X[sn][:]
for sn in range(testsampleClip):
tmplen = len(test_X[sn])
test_data[sn, :tmplen] = test_X[sn][:]
train_data = train_data.reshape(trainsampleClip, Param[i]['number_of_segment'], n_input)
test_data = test_data.reshape(testsampleClip, Param[i]['number_of_segment'], n_input)
elapsed = time.time()-start
sig_comp_time.append(elapsed)
model3 = Model( Param[i]['learning_rate'], training_iters, batch_size, n_input, Param[i]['number_of_segment'], Param[i]['deg_of_sig'], train_data, train_Y, test_data, test_Y)
# Folded raw data feature set generator
elif Param[i]['deg_of_sig']==1:
start = time.time()
train_X, train_Y = GetSeqPenDigit('pendigits-orig.tra.txt')
test_X, test_Y = GetSeqPenDigit('pendigits-orig.tes.txt')
trainsampleClip = len(train_Y)
testsampleClip = len(test_Y)
max_interval = 0
train_increment = [[] for k in range(trainsampleClip)]
test_increment = [[] for k in range(testsampleClip)]
for x in train_X:
if max_interval < len(x):
max_interval = len(x)
for x in test_X:
if max_interval < len(x):
max_interval = len(x)
print(max_interval)
train_data = np.zeros((trainsampleClip, max_interval))
test_data = np.zeros((testsampleClip, max_interval))
for sn in range(trainsampleClip):
tmplen = len(train_X[sn])
train_data[sn, :tmplen] = train_X[sn][:]
for sn in range(testsampleClip):
tmplen = len(test_X[sn])
test_data[sn, :tmplen] = test_X[sn][:]
n_input = 2
train_data = train_data.reshape(trainsampleClip, int(max_interval/2), 2)
test_data = test_data.reshape(testsampleClip, int(max_interval/2), 2)
print(train_data[0])
elapsed = time.time()-start
sig_comp_time.append(elapsed)
model3 = Model( Param[i]['learning_rate'], training_iters, batch_size, n_input, int(max_interval/2), Param[i]['deg_of_sig'], train_data, train_Y, test_data, test_Y)
# Logsig feature set generator
else:
start = time.time()
X_logsig_start, Y = GetSeqPenandCalLogSig(Param[i]['deg_of_sig'], Param[i]['number_of_segment'],'pendigits-orig.tra.txt')
test_X_logsig_start, test_Y = GetSeqPenandCalLogSig(Param[i]['deg_of_sig'], Param[i]['number_of_segment'],'pendigits-orig.tes.txt')
print(X_logsig_start.shape)
print(test_X_logsig_start.shape)
n_input = np.shape(X_logsig_start)[2]
# number_of_samples = np.shape(X_logsig_start)[0]
elapsed = time.time()-start
sig_comp_time.append(elapsed)
model3 = Model( Param[i]['learning_rate'], training_iters, batch_size, n_input, Param[i]['number_of_segment'], Param[i]['deg_of_sig'], X_logsig_start, Y, test_X_logsig_start, test_Y)
# Model built and train
fixed_error_result_model3 = model3.BuildModelKerasMn()
print("Time = " + str(time.time()-start))
print("Testing loss = " + str(fixed_error_result_model3['Loss']))
# model3.KerasPredict()
test_result.append(fixed_error_result_model3 ['Loss'])
test_time.append(fixed_error_result_model3 ['Time'])
# results save
np.save('error_tol'+str(error_tol)+'deg_logsig'+str(Param[i]['deg_of_sig'])+'_test_result', test_result)
np.save('error_tol'+str(error_tol)+'deg_logsig'+str(Param[i]['deg_of_sig'])+'_test_time', test_time)
np.save('error_tol'+str(error_tol)+'deg_logsig'+str(Param[i]['deg_of_sig'])+'_sig_comp_time', elapsed)
print(test_time)
print(test_result)
print(sig_comp_time)
```
| github_jupyter |
```
# !wget https://raw.githubusercontent.com/huseinzol05/Malaya-Dataset/master/dependency/gsd-ud-train.conllu.txt
# !wget https://raw.githubusercontent.com/huseinzol05/Malaya-Dataset/master/dependency/gsd-ud-test.conllu.txt
# !wget https://raw.githubusercontent.com/huseinzol05/Malaya-Dataset/master/dependency/gsd-ud-dev.conllu.txt
# !wget https://raw.githubusercontent.com/huseinzol05/Malaya-Dataset/master/dependency/augmented-dependency.json
with open('gsd-ud-train.conllu.txt') as fopen:
corpus = fopen.read().split('\n')
with open('gsd-ud-test.conllu.txt') as fopen:
corpus.extend(fopen.read().split('\n'))
with open('gsd-ud-dev.conllu.txt') as fopen:
corpus.extend(fopen.read().split('\n'))
import malaya
import re
from malaya.texts._text_functions import split_into_sentences
from malaya.texts import _regex
import numpy as np
import itertools
import tensorflow as tf
from tensorflow.keras.preprocessing.sequence import pad_sequences
tokenizer = malaya.preprocessing._tokenizer
splitter = split_into_sentences
def is_number_regex(s):
if re.match("^\d+?\.\d+?$", s) is None:
return s.isdigit()
return True
def preprocessing(w):
if is_number_regex(w):
return '<NUM>'
elif re.match(_regex._money, w):
return '<MONEY>'
elif re.match(_regex._date, w):
return '<DATE>'
elif re.match(_regex._expressions['email'], w):
return '<EMAIL>'
elif re.match(_regex._expressions['url'], w):
return '<URL>'
else:
w = ''.join(''.join(s)[:2] for _, s in itertools.groupby(w))
return w
def process_string(string):
splitted = string.split()
return [preprocessing(w) for w in splitted]
word2idx = {'PAD': 0,'UNK':1, '_ROOT': 2}
tag2idx = {'PAD': 0, '_<ROOT>': 1}
char2idx = {'PAD': 0,'UNK':1, '_ROOT': 2}
word_idx = 3
tag_idx = 2
char_idx = 3
special_tokens = ['<NUM>', '<MONEY>', '<DATE>', '<URL>', '<EMAIL>']
for t in special_tokens:
word2idx[t] = word_idx
word_idx += 1
char2idx[t] = char_idx
char_idx += 1
word2idx, char2idx
PAD = "_PAD"
PAD_POS = "_PAD_POS"
PAD_TYPE = "_<PAD>"
PAD_CHAR = "_PAD_CHAR"
ROOT = "_ROOT"
ROOT_POS = "_ROOT_POS"
ROOT_TYPE = "_<ROOT>"
ROOT_CHAR = "_ROOT_CHAR"
END = "_END"
END_POS = "_END_POS"
END_TYPE = "_<END>"
END_CHAR = "_END_CHAR"
def process_corpus(corpus, until = None):
global word2idx, tag2idx, char2idx, word_idx, tag_idx, char_idx
sentences, words, depends, labels, pos, chars = [], [], [], [], [], []
temp_sentence, temp_word, temp_depend, temp_label, temp_pos = [], [], [], [], []
first_time = True
for sentence in corpus:
try:
if len(sentence):
if sentence[0] == '#':
continue
if first_time:
print(sentence)
first_time = False
sentence = sentence.split('\t')
for c in sentence[1]:
if c not in char2idx:
char2idx[c] = char_idx
char_idx += 1
if sentence[7] not in tag2idx:
tag2idx[sentence[7]] = tag_idx
tag_idx += 1
sentence[1] = preprocessing(sentence[1])
if sentence[1] not in word2idx:
word2idx[sentence[1]] = word_idx
word_idx += 1
temp_word.append(word2idx[sentence[1]])
temp_depend.append(int(sentence[6]))
temp_label.append(tag2idx[sentence[7]])
temp_sentence.append(sentence[1])
temp_pos.append(sentence[3])
else:
if len(temp_sentence) < 2 or len(temp_word) != len(temp_label):
temp_word = []
temp_depend = []
temp_label = []
temp_sentence = []
temp_pos = []
continue
words.append(temp_word)
depends.append(temp_depend)
labels.append(temp_label)
sentences.append( temp_sentence)
pos.append(temp_pos)
char_ = []
for w in temp_sentence:
if w in char2idx:
char_.append([char2idx[w]])
else:
char_.append([char2idx[c] for c in w])
chars.append(char_)
temp_word = []
temp_depend = []
temp_label = []
temp_sentence = []
temp_pos = []
except Exception as e:
print(e, sentence)
return sentences[:-1], words[:-1], depends[:-1], labels[:-1], pos[:-1], chars[:-1]
sentences, words, depends, labels, _, _ = process_corpus(corpus)
import json
with open('augmented-dependency.json') as fopen:
augmented = json.load(fopen)
text_augmented = []
for a in augmented:
text_augmented.extend(a[0])
depends.extend((np.array(a[1]) - 1).tolist())
labels.extend((np.array(a[2]) + 1).tolist())
def parse_XY(texts):
global word2idx, tag2idx, char2idx, word_idx, tag_idx, char_idx
outside, sentences = [], []
for no, text in enumerate(texts):
s = process_string(text)
sentences.append(s)
inside = []
for w in s:
for c in w:
if c not in char2idx:
char2idx[c] = char_idx
char_idx += 1
if w not in word2idx:
word2idx[w] = word_idx
word_idx += 1
inside.append(word2idx[w])
outside.append(inside)
return outside, sentences
outside, new_sentences = parse_XY(text_augmented)
words.extend(outside)
sentences.extend(new_sentences)
idx2word = {v:k for k, v in word2idx.items()}
idx2tag = {v:k for k, v in tag2idx.items()}
len(idx2word)
from sklearn.model_selection import train_test_split
sentences_train, sentences_test, words_train, words_test, depends_train, depends_test, labels_train, labels_test \
= train_test_split(sentences, words, depends, labels, test_size = 0.2)
len(sentences_train), len(sentences_test)
def generate_char_seq(batch, UNK = 2):
maxlen_c = max([len(k) for k in batch])
x = [[len(i) for i in k] for k in batch]
maxlen = max([j for i in x for j in i])
temp = np.zeros((len(batch),maxlen_c,maxlen),dtype=np.int32)
for i in range(len(batch)):
for k in range(len(batch[i])):
for no, c in enumerate(batch[i][k]):
temp[i,k,-1-no] = char2idx.get(c, UNK)
return temp
generate_char_seq(sentences_train[:5]).shape
train_X = words_train
train_Y = labels_train
train_depends = depends_train
train_char = sentences_train
test_X = words_test
test_Y = labels_test
test_depends = depends_test
test_char = sentences_test
class BiAAttention:
def __init__(self, input_size_encoder, input_size_decoder, num_labels):
self.input_size_encoder = input_size_encoder
self.input_size_decoder = input_size_decoder
self.num_labels = num_labels
self.W_d = tf.get_variable("W_d", shape=[self.num_labels, self.input_size_decoder],
initializer=tf.contrib.layers.xavier_initializer())
self.W_e = tf.get_variable("W_e", shape=[self.num_labels, self.input_size_encoder],
initializer=tf.contrib.layers.xavier_initializer())
self.U = tf.get_variable("U", shape=[self.num_labels, self.input_size_decoder, self.input_size_encoder],
initializer=tf.contrib.layers.xavier_initializer())
def forward(self, input_d, input_e, mask_d=None, mask_e=None):
batch = tf.shape(input_d)[0]
length_decoder = tf.shape(input_d)[1]
length_encoder = tf.shape(input_e)[1]
out_d = tf.expand_dims(tf.matmul(self.W_d, tf.transpose(input_d, [0, 2, 1])), 3)
out_e = tf.expand_dims(tf.matmul(self.W_e, tf.transpose(input_e, [0, 2, 1])), 2)
output = tf.matmul(tf.expand_dims(input_d, 1), self.U)
output = tf.matmul(output, tf.transpose(tf.expand_dims(input_e, 1), [0, 1, 3, 2]))
output = output + out_d + out_e
if mask_d is not None:
d = tf.expand_dims(tf.expand_dims(mask_d, 1), 3)
e = tf.expand_dims(tf.expand_dims(mask_e, 1), 2)
output = output * d * e
return output
class Model:
def __init__(
self,
dim_word,
dim_char,
dropout,
learning_rate,
hidden_size_char,
hidden_size_word,
num_layers
):
def cells(size, reuse = False):
return tf.contrib.rnn.DropoutWrapper(
tf.nn.rnn_cell.LSTMCell(
size,
initializer = tf.orthogonal_initializer(),
reuse = reuse,
),
output_keep_prob = dropout,
)
self.word_ids = tf.placeholder(tf.int32, shape = [None, None])
self.char_ids = tf.placeholder(tf.int32, shape = [None, None, None])
self.labels = tf.placeholder(tf.int32, shape = [None, None])
self.depends = tf.placeholder(tf.int32, shape = [None, None])
self.maxlen = tf.shape(self.word_ids)[1]
self.lengths = tf.count_nonzero(self.word_ids, 1)
self.mask = tf.math.not_equal(self.word_ids, 0)
float_mask = tf.cast(self.mask, tf.float32)
self.arc_h = tf.layers.Dense(hidden_size_word)
self.arc_c = tf.layers.Dense(hidden_size_word)
self.attention = BiAAttention(hidden_size_word, hidden_size_word, 1)
self.word_embeddings = tf.Variable(
tf.truncated_normal(
[len(word2idx), dim_word], stddev = 1.0 / np.sqrt(dim_word)
)
)
self.char_embeddings = tf.Variable(
tf.truncated_normal(
[len(char2idx), dim_char], stddev = 1.0 / np.sqrt(dim_char)
)
)
word_embedded = tf.nn.embedding_lookup(
self.word_embeddings, self.word_ids
)
char_embedded = tf.nn.embedding_lookup(
self.char_embeddings, self.char_ids
)
s = tf.shape(char_embedded)
char_embedded = tf.reshape(
char_embedded, shape = [s[0] * s[1], s[-2], dim_char]
)
for n in range(num_layers):
(out_fw, out_bw), (
state_fw,
state_bw,
) = tf.nn.bidirectional_dynamic_rnn(
cell_fw = cells(hidden_size_char),
cell_bw = cells(hidden_size_char),
inputs = char_embedded,
dtype = tf.float32,
scope = 'bidirectional_rnn_char_%d' % (n),
)
char_embedded = tf.concat((out_fw, out_bw), 2)
output = tf.reshape(
char_embedded[:, -1], shape = [s[0], s[1], 2 * hidden_size_char]
)
word_embedded = tf.concat([word_embedded, output], axis = -1)
for n in range(num_layers):
(out_fw, out_bw), (
state_fw,
state_bw,
) = tf.nn.bidirectional_dynamic_rnn(
cell_fw = cells(hidden_size_word),
cell_bw = cells(hidden_size_word),
inputs = word_embedded,
dtype = tf.float32,
scope = 'bidirectional_rnn_word_%d' % (n),
)
word_embedded = tf.concat((out_fw, out_bw), 2)
logits = tf.layers.dense(word_embedded, len(idx2tag))
log_likelihood, transition_params = tf.contrib.crf.crf_log_likelihood(
logits, self.labels, self.lengths
)
arc_h = tf.nn.elu(self.arc_h(word_embedded))
arc_c = tf.nn.elu(self.arc_c(word_embedded))
out_arc = tf.squeeze(self.attention.forward(arc_h, arc_h, mask_d=float_mask, mask_e=float_mask), axis = 1)
batch = tf.shape(out_arc)[0]
batch_index = tf.range(0, batch)
max_len = tf.shape(out_arc)[1]
sec_max_len = tf.shape(out_arc)[2]
minus_inf = -1e8
minus_mask = (1 - float_mask) * minus_inf
out_arc = out_arc + tf.expand_dims(minus_mask, axis = 2) + tf.expand_dims(minus_mask, axis = 1)
loss_arc = tf.nn.log_softmax(out_arc, dim=1)
loss_arc = loss_arc * tf.expand_dims(float_mask, axis = 2) * tf.expand_dims(float_mask, axis = 1)
num = tf.reduce_sum(float_mask) - tf.cast(batch, tf.float32)
child_index = tf.tile(tf.expand_dims(tf.range(0, max_len), 1), [1, batch])
t = tf.transpose(self.depends)
broadcasted = tf.broadcast_to(batch_index, tf.shape(t))
concatenated = tf.transpose(tf.concat([tf.expand_dims(broadcasted, axis = 0),
tf.expand_dims(t, axis = 0),
tf.expand_dims(child_index, axis = 0)], axis = 0))
loss_arc = tf.gather_nd(loss_arc, concatenated)
loss_arc = tf.transpose(loss_arc, [1, 0])[1:]
loss_arc = tf.reduce_sum(-loss_arc) / num
self.cost = tf.reduce_mean(-log_likelihood) + loss_arc
self.optimizer = tf.train.AdamOptimizer(
learning_rate = learning_rate
).minimize(self.cost)
mask = tf.sequence_mask(self.lengths, maxlen = self.maxlen)
self.tags_seq, _ = tf.contrib.crf.crf_decode(
logits, transition_params, self.lengths
)
out_arc = out_arc + tf.linalg.diag(tf.fill([max_len], -np.inf))
minus_mask = tf.expand_dims(tf.cast(1.0 - float_mask, tf.bool), axis = 2)
minus_mask = tf.tile(minus_mask, [1, 1, sec_max_len])
out_arc = tf.where(minus_mask, tf.fill(tf.shape(out_arc), -np.inf), out_arc)
self.heads = tf.argmax(out_arc, axis = 1)
self.prediction = tf.boolean_mask(self.tags_seq, mask)
mask_label = tf.boolean_mask(self.labels, mask)
correct_pred = tf.equal(self.prediction, mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
self.prediction = tf.cast(tf.boolean_mask(self.heads, mask), tf.int32)
mask_label = tf.boolean_mask(self.depends, mask)
correct_pred = tf.equal(self.prediction, mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy_depends = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
tf.reset_default_graph()
sess = tf.InteractiveSession()
dim_word = 128
dim_char = 256
dropout = 0.8
learning_rate = 1e-3
hidden_size_char = 128
hidden_size_word = 128
num_layers = 2
model = Model(dim_word,dim_char,dropout,learning_rate,hidden_size_char,hidden_size_word,num_layers)
sess.run(tf.global_variables_initializer())
batch_x = train_X[:5]
batch_x = pad_sequences(batch_x,padding='post')
batch_char = train_char[:5]
batch_char = generate_char_seq(batch_char)
batch_y = train_Y[:5]
batch_y = pad_sequences(batch_y,padding='post')
batch_depends = train_depends[:5]
batch_depends = pad_sequences(batch_depends,padding='post')
sess.run([model.accuracy, model.accuracy_depends, model.cost],
feed_dict = {model.word_ids: batch_x,
model.char_ids: batch_char,
model.labels: batch_y,
model.depends: batch_depends})
from tqdm import tqdm
batch_size = 64
epoch = 10
for e in range(epoch):
train_acc, train_loss = [], []
test_acc, test_loss = [], []
train_acc_depends, test_acc_depends = [], []
pbar = tqdm(
range(0, len(train_X), batch_size), desc = 'train minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(train_X))
batch_x = train_X[i: index]
batch_x = pad_sequences(batch_x,padding='post')
batch_char = train_char[i: index]
batch_char = generate_char_seq(batch_char)
batch_y = train_Y[i: index]
batch_y = pad_sequences(batch_y,padding='post')
batch_depends = train_depends[i: index]
batch_depends = pad_sequences(batch_depends,padding='post')
acc_depends, acc, cost, _ = sess.run(
[model.accuracy_depends, model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.word_ids: batch_x,
model.char_ids: batch_char,
model.labels: batch_y,
model.depends: batch_depends
},
)
train_loss.append(cost)
train_acc.append(acc)
train_acc_depends.append(acc_depends)
pbar.set_postfix(cost = cost, accuracy = acc, accuracy_depends = acc_depends)
pbar = tqdm(
range(0, len(test_X), batch_size), desc = 'test minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(test_X))
batch_x = test_X[i: index]
batch_x = pad_sequences(batch_x,padding='post')
batch_char = test_char[i: index]
batch_char = generate_char_seq(batch_char)
batch_y = test_Y[i: index]
batch_y = pad_sequences(batch_y,padding='post')
batch_depends = test_depends[i: index]
batch_depends = pad_sequences(batch_depends,padding='post')
acc_depends, acc, cost = sess.run(
[model.accuracy_depends, model.accuracy, model.cost],
feed_dict = {
model.word_ids: batch_x,
model.char_ids: batch_char,
model.labels: batch_y,
model.depends: batch_depends
},
)
test_loss.append(cost)
test_acc.append(acc)
test_acc_depends.append(acc_depends)
pbar.set_postfix(cost = cost, accuracy = acc, accuracy_depends = acc_depends)
print(
'epoch: %d, training loss: %f, training acc: %f, training depends: %f, valid loss: %f, valid acc: %f, valid depends: %f\n'
% (e, np.mean(train_loss),
np.mean(train_acc),
np.mean(train_acc_depends),
np.mean(test_loss),
np.mean(test_acc),
np.mean(test_acc_depends)
))
tags_seq, heads = sess.run(
[model.tags_seq, model.heads],
feed_dict = {
model.word_ids: batch_x,
model.char_ids: batch_char
},
)
tags_seq[0], heads[0], batch_depends[0]
def evaluate(heads_pred, types_pred, heads, types, lengths,
symbolic_root=False, symbolic_end=False):
batch_size, _ = heads_pred.shape
ucorr = 0.
lcorr = 0.
total = 0.
ucomplete_match = 0.
lcomplete_match = 0.
corr_root = 0.
total_root = 0.
start = 1 if symbolic_root else 0
end = 1 if symbolic_end else 0
for i in range(batch_size):
ucm = 1.
lcm = 1.
for j in range(start, lengths[i] - end):
total += 1
if heads[i, j] == heads_pred[i, j]:
ucorr += 1
if types[i, j] == types_pred[i, j]:
lcorr += 1
else:
lcm = 0
else:
ucm = 0
lcm = 0
if heads[i, j] == 0:
total_root += 1
corr_root += 1 if heads_pred[i, j] == 0 else 0
ucomplete_match += ucm
lcomplete_match += lcm
return ucorr / total, lcorr / total, corr_root / total_root
arc_accuracy, type_accuracy, root_accuracy = evaluate(heads, tags_seq, batch_depends, batch_y,
np.count_nonzero(batch_x, axis = 1))
arc_accuracy, type_accuracy, root_accuracy
arcs, types, roots = [], [], []
pbar = tqdm(
range(0, len(test_X), batch_size), desc = 'test minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(test_X))
batch_x = test_X[i: index]
batch_x = pad_sequences(batch_x,padding='post')
batch_char = test_char[i: index]
batch_char = generate_char_seq(batch_char)
batch_y = test_Y[i: index]
batch_y = pad_sequences(batch_y,padding='post')
batch_depends = test_depends[i: index]
batch_depends = pad_sequences(batch_depends,padding='post')
tags_seq, heads = sess.run(
[model.tags_seq, model.heads],
feed_dict = {
model.word_ids: batch_x,
model.char_ids: batch_char
},
)
arc_accuracy, type_accuracy, root_accuracy = evaluate(heads, tags_seq, batch_depends, batch_y,
np.count_nonzero(batch_x, axis = 1))
pbar.set_postfix(arc_accuracy = arc_accuracy, type_accuracy = type_accuracy,
root_accuracy = root_accuracy)
arcs.append(arc_accuracy)
types.append(type_accuracy)
roots.append(root_accuracy)
print('arc accuracy:', np.mean(arcs))
print('types accuracy:', np.mean(types))
print('root accuracy:', np.mean(roots))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/AI4Finance-Foundation/FinRL/blob/master/FinRL_Raytune_for_Hyperparameter_Optimization_RLlib%20Models.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#Installing FinRL
%%capture
!pip install git+https://github.com/AI4Finance-LLC/FinRL-Library.git
%%capture
!pip install "ray[tune]" optuna
%%capture
!pip install int_date==0.1.8
```
#Importing libraries
```
#Importing the libraries
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# matplotlib.use('Agg')
import datetime
import optuna
%matplotlib inline
from finrl import config
from finrl.finrl_meta.preprocessor.yahoodownloader import YahooDownloader
from finrl.finrl_meta.preprocessor.preprocessors import FeatureEngineer, data_split
from finrl.finrl_meta.env_stock_trading.env_stocktrading_np import StockTradingEnv as StockTradingEnv_numpy
from finrl.agents.rllib.models import DRLAgent as DRLAgent_rllib
from stable_baselines3.common.vec_env import DummyVecEnv
from finrl.finrl_meta.data_processor import DataProcessor
from finrl.plot import backtest_stats, backtest_plot, get_daily_return, get_baseline
import ray
from pprint import pprint
from ray.rllib.agents.ppo import PPOTrainer
from ray.rllib.agents.ddpg import DDPGTrainer
from ray.rllib.agents.a3c import A2CTrainer
from ray.rllib.agents.a3c import a2c
from ray.rllib.agents.ddpg import ddpg, td3
from ray.rllib.agents.ppo import ppo
from ray.rllib.agents.sac import sac
import sys
sys.path.append("../FinRL-Library")
import os
import itertools
from ray import tune
from ray.tune.suggest import ConcurrencyLimiter
from ray.tune.schedulers import AsyncHyperBandScheduler
from ray.tune.suggest.optuna import OptunaSearch
from ray.tune.registry import register_env
import time
import psutil
psutil_memory_in_bytes = psutil.virtual_memory().total
ray._private.utils.get_system_memory = lambda: psutil_memory_in_bytes
from typing import Dict, Optional, Any
import os
if not os.path.exists("./" + config.DATA_SAVE_DIR):
os.makedirs("./" + config.DATA_SAVE_DIR)
if not os.path.exists("./" + config.TRAINED_MODEL_DIR):
os.makedirs("./" + config.TRAINED_MODEL_DIR)
if not os.path.exists("./" + config.TENSORBOARD_LOG_DIR):
os.makedirs("./" + config.TENSORBOARD_LOG_DIR)
if not os.path.exists("./" + config.RESULTS_DIR):
os.makedirs("./" + config.RESULTS_DIR)
# if not os.path.exists("./" + "tuned_models"):
# os.makedirs("./" + "tuned_models")
```
##Defining the hyperparameter search space
1. You can look up [here](https://docs.ray.io/en/latest/tune/key-concepts.html#search-spaces) to learn how to define hyperparameter search space
2. Jump over to this [link](https://github.com/DLR-RM/rl-baselines3-zoo/blob/master/utils/hyperparams_opt.py) to find the range of different hyperparameter
3. To learn about different hyperparameters for different algorithms for RLlib models, jump over to this [link](https://docs.ray.io/en/latest/rllib-algorithms.html)
```
def sample_ddpg_params():
return {
"buffer_size": tune.choice([int(1e4), int(1e5), int(1e6)]),
"lr": tune.loguniform(1e-5, 1),
"train_batch_size": tune.choice([32, 64, 128, 256, 512])
}
def sample_a2c_params():
return{
"lambda": tune.choice([0.1,0.3,0.5,0.7,0.9,1.0]),
"entropy_coeff": tune.loguniform(0.00000001, 0.1),
"lr": tune.loguniform(1e-5, 1)
}
def sample_ppo_params():
return {
"entropy_coeff": tune.loguniform(0.00000001, 0.1),
"lr": tune.loguniform(5e-5, 1),
"sgd_minibatch_size": tune.choice([ 32, 64, 128, 256, 512]),
"lambda": tune.choice([0.1,0.3,0.5,0.7,0.9,1.0])
}
MODELS = {"a2c": a2c, "ddpg": ddpg, "td3": td3, "sac": sac, "ppo": ppo}
```
## Getting the training and testing environment
```
def get_train_env(start_date, end_date, ticker_list, data_source, time_interval,
technical_indicator_list, env, model_name, if_vix = True,
**kwargs):
#fetch data
DP = DataProcessor(data_source, **kwargs)
data = DP.download_data(ticker_list, start_date, end_date, time_interval)
data = DP.clean_data(data)
data = DP.add_technical_indicator(data, technical_indicator_list)
if if_vix:
data = DP.add_vix(data)
price_array, tech_array, turbulence_array = DP.df_to_array(data, if_vix)
train_env_config = {'price_array':price_array,
'tech_array':tech_array,
'turbulence_array':turbulence_array,
'if_train':True}
return train_env_config
#Function to calculate the sharpe ratio from the list of total_episode_reward
def calculate_sharpe(episode_reward:list):
perf_data = pd.DataFrame(data=episode_reward,columns=['reward'])
perf_data['daily_return'] = perf_data['reward'].pct_change(1)
if perf_data['daily_return'].std() !=0:
sharpe = (252**0.5)*perf_data['daily_return'].mean()/ \
perf_data['daily_return'].std()
return sharpe
else:
return 0
def get_test_config(start_date, end_date, ticker_list, data_source, time_interval,
technical_indicator_list, env, model_name, if_vix = True,
**kwargs):
DP = DataProcessor(data_source, **kwargs)
data = DP.download_data(ticker_list, start_date, end_date, time_interval)
data = DP.clean_data(data)
data = DP.add_technical_indicator(data, technical_indicator_list)
if if_vix:
data = DP.add_vix(data)
price_array, tech_array, turbulence_array = DP.df_to_array(data, if_vix)
test_env_config = {'price_array':price_array,
'tech_array':tech_array,
'turbulence_array':turbulence_array,'if_train':False}
return test_env_config
def val_or_test(test_env_config,agent_path,model_name,env):
episode_total_reward = DRL_prediction(model_name,test_env_config,
env = env,
agent_path=agent_path)
return calculate_sharpe(episode_total_reward),episode_total_reward
TRAIN_START_DATE = '2014-01-01'
TRAIN_END_DATE = '2019-07-30'
VAL_START_DATE = '2019-08-01'
VAL_END_DATE = '2020-07-30'
TEST_START_DATE = '2020-08-01'
TEST_END_DATE = '2021-10-01'
technical_indicator_list =config.INDICATORS
model_name = 'a2c'
env = StockTradingEnv_numpy
ticker_list = ['TSLA']
data_source = 'yahoofinance'
time_interval = '1D'
train_env_config = get_train_env(TRAIN_START_DATE, VAL_END_DATE,
ticker_list, data_source, time_interval,
technical_indicator_list, env, model_name)
```
## Registering the environment
```
from ray.tune.registry import register_env
env_name = 'StockTrading_train_env'
register_env(env_name, lambda config: env(train_env_config))
```
## Running tune
```
MODEL_TRAINER = {'a2c':A2CTrainer,'ppo':PPOTrainer,'ddpg':DDPGTrainer}
if model_name == "ddpg":
sample_hyperparameters = sample_ddpg_params()
elif model_name == "ppo":
sample_hyperparameters = sample_ppo_params()
elif model_name == "a2c":
sample_hyperparameters = sample_a2c_params()
def run_optuna_tune():
algo = OptunaSearch()
algo = ConcurrencyLimiter(algo,max_concurrent=4)
scheduler = AsyncHyperBandScheduler()
num_samples = 10
training_iterations = 100
analysis = tune.run(
MODEL_TRAINER[model_name],
metric="episode_reward_mean", #The metric to optimize for tuning
mode="max", #Maximize the metric
search_alg = algo,#OptunaSearch method which uses Tree Parzen estimator to sample hyperparameters
scheduler=scheduler, #To prune bad trials
config = {**sample_hyperparameters,
'env':'StockTrading_train_env','num_workers':1,
'num_gpus':1,'framework':'torch'},
num_samples = num_samples, #Number of hyperparameters to test out
stop = {'training_iteration':training_iterations},#Time attribute to validate the results
verbose=1,local_dir="./tuned_models",#Saving tensorboard plots
# resources_per_trial={'gpu':1,'cpu':1},
max_failures = 1,#Extra Trying for the failed trials
raise_on_failed_trial=False,#Don't return error even if you have errored trials
keep_checkpoints_num = num_samples-5,
checkpoint_score_attr ='episode_reward_mean',#Only store keep_checkpoints_num trials based on this score
checkpoint_freq=training_iterations#Checpointing all the trials
)
print("Best hyperparameter: ", analysis.best_config)
return analysis
analysis = run_optuna_tune()
```
## Best config, directory and checkpoint for hyperparameters
```
best_config = analysis.get_best_config(metric='episode_reward_mean',mode='max')
best_config
best_logdir = analysis.get_best_logdir(metric='episode_reward_mean',mode='max')
best_logdir
best_checkpoint = analysis.best_checkpoint
best_checkpoint
# sharpe,df_account_test,df_action_test = val_or_test(TEST_START_DATE, TEST_END_DATE, ticker_list, data_source, time_interval,
# technical_indicator_list, env, model_name,best_checkpoint, if_vix = True)
test_env_config = get_test_config(TEST_START_DATE, TEST_END_DATE, ticker_list, data_source, time_interval,
technical_indicator_list, env, model_name)
sharpe,account,actions = val_or_test(test_env_config,agent_path,model_name,env)
def DRL_prediction(
model_name,
test_env_config,
env,
model_config,
agent_path,
env_name_test='StockTrading_test_env'
):
env_instance = env(test_env_config)
register_env(env_name_test, lambda config: env(test_env_config))
model_config['env'] = env_name_test
# ray.init() # Other Ray APIs will not work until `ray.init()` is called.
if model_name == "ppo":
trainer = MODELS[model_name].PPOTrainer(config=model_config)
elif model_name == "a2c":
trainer = MODELS[model_name].A2CTrainer(config=model_config)
elif model_name == "ddpg":
trainer = MODELS[model_name].DDPGTrainer(config=model_config)
elif model_name == "td3":
trainer = MODELS[model_name].TD3Trainer(config=model_config)
elif model_name == "sac":
trainer = MODELS[model_name].SACTrainer(config=model_config)
try:
trainer.restore(agent_path)
print("Restoring from checkpoint path", agent_path)
except BaseException:
raise ValueError("Fail to load agent!")
# test on the testing env
state = env_instance.reset()
episode_returns = list() # the cumulative_return / initial_account
episode_total_assets = list()
episode_total_assets.append(env_instance.initial_total_asset)
done = False
while not done:
action = trainer.compute_single_action(state)
state, reward, done, _ = env_instance.step(action)
total_asset = (
env_instance.amount
+ (env_instance.price_ary[env_instance.day] * env_instance.stocks).sum()
)
episode_total_assets.append(total_asset)
episode_return = total_asset / env_instance.initial_total_asset
episode_returns.append(episode_return)
ray.shutdown()
print("episode return: " + str(episode_return))
print("Test Finished!")
return episode_total_assets
episode_total_assets = DRL_prediction(
model_name,
test_env_config,
env,
best_config,
best_checkpoint,
env_name_test='StockTrading_test_env')
print('The test sharpe ratio is: ',calculate_sharpe(episode_total_assets))
df_account_test = pd.DataFrame(data=episode_total_assets,columns=['account_value'])
```
| github_jupyter |
# Introduction
*Cole Plum*
This notebook performs a very simple analysis of some data about **earthquakes**. This earthquake data originally comes from the United States Geological Survey. The data was downloaded from the CORGIS project, a collection of datasets. More information about the dataset can be found on the [CORGIS site](https://corgis-edu.github.io/corgis/json/earthquakes/).
For this analysis, I will explore the following research question: *"What is the distribution of earthquake magnitudes?"* I will answer this question by doing the following:
1. Downloading a JSON file of earthquake data
2. Printing the first element of the dataset, to learn its structure
3. Extracting out the magnitude information from all the earthquakes
4. Plotting the distribution of magnitudes
5. Discussing the plot and its implications for society
# Downloading Data
```
import requests
url = "https://corgis-edu.github.io/corgis/datasets/json/earthquakes/earthquakes.json"
data = requests.get(url).json()
```
# Previewing the Structure
```
from pprint import pprint
# The pretty print (pprint) module is great for
# quickly visualizing structured data!
pprint(data[0])
```
# Extract Magnitudes
```
magnitudes = [earthquake['impact']['magnitude']
for earthquake in data]
```
# Plotting Magnitudes
```
import matplotlib.pyplot as plt
plt.hist(magnitudes)
plt.title("Distribution of Earthquake Magnitudes")
plt.xlabel("Magnitude")
plt.ylabel("Number of Earthquakes")
plt.show()
```
# Discussion
The histogram above shows that most earthquakes are not very big, with a magnitude of less than 2. However, some earthquakes are very severe, reaching up to a 7 or 8 for their magnitudes. The dataset collects data from all over the world for about a month, so in general, there are quite a few earthquakes that occur.
A number of stakeholders could be interested in these results:
- People building houses would want to know whether they need to regularly plan for very strong earthquakes.
- Emergency relief effort planners would want to know how often they need to plan for earthquakes in general.
- Scientists studying tectonic plate activity could use this to determine if a particular earthquake was unusual.
A potential conflict between these stakeholders could be between emergency relief planners and builders; builders might want to conserve stronger materials since strong earthquakes are relatively rare, while relief planners would rather they always use strong materials since earthquakes are so frequent. According to the ethical theory of [Least Harm](http://www.dsef.org/wp-content/uploads/2012/07/EthicalTheories.pdf), which states that decision makers should always attempt to harm the fewest people possible, the builders should just use the stronger materials to minimize the threat to the populace.
*I have neither given nor received unauthorized assistance on this assignment.*
| github_jupyter |
```
A = {1,2,3,8}
B = {3,4}
print(1 in A)
print(4 in A)
flag = 4 in A
type(flag)
print(B.issubset(A))
def f_issubset(A,B):
for e in A:
if e in B:
pass
else:
return False
return True
print(f_issubset(B,A))
print(f_issubset({2,3,4},{1,2,3,4,5,6}))
import numpy as np
Omg = set(np.arange(10))
type(Omg)
Omg
A = set(np.arange(0,9,2))
A
B = set(np.arange(1,9,3))
B
A.union(B)
A.intersection(B)
B.add(6)
B
A.intersection(B)
A
B
A.difference(B)
A_complement = Omg.difference(A)
A_complement
B_complement = Omg.difference(B)
Omg.difference(A.union(B))
A_complement.intersection(B_complement)
Omg.difference(A.intersection(B))
A_complement.union(B_complement)
import numpy as np
def BernoulliTrail(p=0.5):
X = int(np.random.rand()<=p)
return X
print(BernoulliTrail(0.7))
countOnes = 0
countZeros = 0
p = 0.611334
n = 1000000
for i in range(n):
x = BernoulliTrail(p)
if x == 1:
countOnes += 1
else:
countZeros += 1
print("Success Fraction is : ",countOnes/n)
def GT(p):
X = 1
while True:
if BernoulliTrail(p) == 1:
return X
else:
X+=1
p = 0.2
print(GT(p))
n = 1000
p = 0.6
G = np.zeros(n)
for i in range(n):
G[i] = GT(p)
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
n = 100000
p = 0.02
G = np.zeros(n)
for i in range(n):
G[i] = GT(p)
plt.hist(G,density=True,bins=100)
sns.kdeplot(G,shade=True)
def BT(n,p):
X = (np.random.rand(n)<=p).sum()
return X
print(BT(100,0.2))
N = 10000
n = 1000
p = 0.1
B = np.zeros(N)
for i in range(N):
B[i] = BT(n,p)
plt.hist(B,density=True,bins=20)
sns.kdeplot(B,shade=True)
import pandas as pd
iris = sns.load_dataset('iris')
iris.head()
iris.species.unique()
T = sns.load_dataset('titanic')
T.head()
T.survived.unique()
T.pclass.unique()
B = pd.read_csv(r'C:\Users\DeLL\Desktop\ML\fullcourse\PS\Data\Bias_correction_ucl.csv')
B.shape
B.head()
X = 100*np.random.rand(100000)+20
plt.hist(X,density=True,bins = 50)
sns.kdeplot(X)
x = np.linspace(0,10,1000)
lmdaz = np.linspace(1,2,4)
for i in lmdaz:
fx = i*np.exp(-i*x)
plt.plot(x,fx,label=str(i))
plt.legend()
x = np.linspace(-20,20,10000)
#muz = np.arange(-20,20,10)
mu = 0
sgmaz = np.arange(1,10,2)
for sgma in sgmaz:
#sgma = 1
fx = np.exp((-(x-mu)**2)/(2*(sgma**2)))/((2*np.pi*sgma**2)**0.5)
plt.plot(x,fx,label=str(sgma))
plt.legend()
X = np.random.normal(0,1,1000)
Y = np.random.normal(10,10,1000)
Z = np.random.normal(-10,4,1000)
W = np.random.exponential(scale=1,size=1000)
plt.hist(X,density=True,alpha=0.5,bins = 30)
#plt.hist(Y,density=True,alpha=0.5,bins = 30)
plt.hist(Z,density=True,alpha=0.5,bins = 30)
plt.hist(W,density=True,alpha=0.5,bins = 30)
iris = sns.load_dataset('iris')
X = iris.sepal_length
plt.hist(X,density=True,bins = 30)
sns.pairplot(iris,hue='species')
p = 0.73
N = 1000000
(np.random.rand(N)<=p).mean()
X = np.random.geometric(p,N)
X.mean()
n = 330
B = np.random.binomial(n,p,N)
B.mean()
B[0]
B[1]
B[3]
```
# Bernoulli
```
# E[X] = p
p = 0.812
sMeanX = []
for N in range(1,10000,100):
sMeanX.append((np.random.rand(N)<=p).mean())
y = np.array(sMeanX)
x = np.arange(y.size)
plt.plot(x,y,linestyle='--')
```
# Geometric(p), $E[X]=1/p$
```
# E[X] = 1/p
p = 0.312
print(1/p)
sMeanX = []
for N in range(1,10000,100):
sMeanX.append((np.random.geometric(p,N).mean()))
y = np.array(sMeanX)
x = np.arange(y.size)
plt.plot(x,y,linestyle='--')
y[-1]
np.random.geometric(p,1000000).mean()
```
# Binomial(n,p) $E[X]=np$
```
# E[X] = np
n = 350
p = 0.312
print(n*p)
sMeanX = []
for N in range(1,10000,100):
sMeanX.append((np.random.binomial(n,p,N).mean()))
y = np.array(sMeanX)
x = np.arange(y.size)
plt.plot(x,y,linestyle='--')
y[-1]
```
# Normal($\mu,\sigma$), $E[X]=\mu$
```
# E[X] = mu
mu = 2.4
sgma = 3.1
sMeanX = []
for N in range(1,10000,100):
sMeanX.append((np.random.normal(mu,sgma,N).mean()))
y = np.array(sMeanX)
x = np.arange(y.size)
plt.plot(x,y,linestyle='--')
#import seaborn as sns
iris = sns.load_dataset('iris')
iris.head()
iris.species.unique()
sns.pairplot(iris,hue='species')
X = iris.petal_length
X.shape
y = iris.species
y.unique()
y[y=='setosa'] = '0'
y[y=='versicolor'] = '1'
y[y=='verginica'] = '2'
y
L = y.unique()
L[-1]
y[y==L[-1]] = '2'
y.unique()
X
y
idx = np.random.permutation(np.arange(y.size))
nTrain = 120
nTest = idx.size-nTrain
Xtrain=X[idx[:nTrain]]
ytrain=y[idx[:nTrain]]
Xtest =X[idx[nTrain:]]
ytest =y[idx[nTrain:]]
ytrain.size
ytest.size
Py = np.zeros(3)
Py[0] = (ytrain=='0').sum()/ytrain.size
Py[1] = (ytrain=='1').sum()/ytrain.size
Py[2] = (ytrain=='2').sum()/ytrain.size
Py
fx_given_y = [[0,0],[0,0],[0,0]]
X_0 = Xtrain[ytrain=='0']
X_1 = Xtrain[ytrain=='1']
X_2 = Xtrain[ytrain=='2']
mu_0 = X_0.mean()
mu_1 = X_1.mean()
mu_2 = X_2.mean()
sgma_0 = X_0.std()
sgma_1 = X_1.std()
sgma_2 = X_2.std()
fx_given_y[0][0] = mu_0
fx_given_y[0][1] = sgma_0
fx_given_y[1][0] = mu_1
fx_given_y[1][1] = sgma_1
fx_given_y[2][0] = mu_2
fx_given_y[2][1] = sgma_2
x = np.linspace(-2,10,10000)
for i in range(3):
mu = fx_given_y[i][0]
sgma = fx_given_y[i][1]
fx = np.exp((-(x-mu)**2)/(2*(sgma**2)))/((2*np.pi*sgma**2)**0.5)
plt.plot(x,fx,label=str(i))
plt.legend()
type(Xtest)
Xtest = np.asarray(Xtest)
type(Xtest)
xt = Xtest[20]
print(xt)
i = 0
mu = fx_given_y[i][0]
sgma = fx_given_y[i][1]
Py_0_given_x = Py[0]*np.exp((-(xt-mu)**2)/(2*(sgma**2)))/((2*np.pi*sgma**2)**0.5)
Py_0_given_x
i = 1
mu = fx_given_y[i][0]
sgma = fx_given_y[i][1]
Py_1_given_x = Py[0]*np.exp((-(xt-mu)**2)/(2*(sgma**2)))/((2*np.pi*sgma**2)**0.5)
i = 2
mu = fx_given_y[i][0]
sgma = fx_given_y[i][1]
Py_2_given_x = Py[0]*np.exp((-(xt-mu)**2)/(2*(sgma**2)))/((2*np.pi*sgma**2)**0.5)
print(Py_0_given_x,Py_1_given_x,Py_2_given_x)
ytest = np.asarray(ytest)
ytest[20]
iris.head()
X = np.array(iris.petal_length)
X
plt.hist(X,bins=30,density=True)
sns.kdeplot(X)
X = np.random.exponential(1,10)
X
```
| github_jupyter |
## Welcome to Aequitas
The Aequitas toolkit is a flexible bias-audit utility for algorithmic decision-making models, accessible via Python API, command line interface (CLI), and through our [web application](http://aequitas.dssg.io/).
Use Aequitas to evaluate model performance across several bias and fairness metrics, and utilize the [most relevant metrics](https://dsapp.uchicago.edu/wp-content/uploads/2018/05/metrictree-1200x750.png) to your process in model selection.
Aequitas will help you:
- Understand where biases exist in your model(s)
- Compare the level of bias between groups in your sample population (bias disparity)
- Visualize absolute bias metrics and their related disparities for rapid comprehension and decision-making
Our goal is to support informed and equitable action for both machine learnining practitioners and the decision-makers who rely on them.
Aequitas is compatible with: **Python 3.6+**
<a id='getting_started'></a>
# Getting started
You can audit your risk assessment system for two types of biases:
- Biased actions or interventions that are not allocated in a way that’s representative of the population.
- Biased outcomes through actions or interventions that are a result of your system being wrong about certain groups of people.
For both audits, you need the following data:
- Data about the specific attributes (race, gender, age, income, etc.) you want to audit for the the overall population considered for interventions
- The set of individuals in the above population that your risk assessment system recommended/ selected for intervention or action. _It’s important t this set come from the assessments made after the system has been built, and not from the data the machine learning system was “trained” on if you're using the audit as a factor in model selection._
If you want to audit for biases due to model or system errors, you also need to include actual outcomes (label values) for all individuals in the overall population.
Input data has slightly different requirements depending on whether you are using Aequitas via the webapp, CLI or Python package. In general, input data is a single table with the following columns:
- `score`
- `label_value` (for error-based metrics only)
- at least one attribute e.g. `race`, `sex` and `age_cat` (attribute categories defined by user)
## Bias measures tailored to your problem
### Input machine learning predictions
After [installing on your computer](./installation.html)
Run `aequitas-report` on [COMPAS data](https://github.com/dssg/aequitas/tree/master/examples):
`compas_for_aequitas.csv` excerpt:
| score | label_value| race | sex | age_cat |
| --------- |------------| -----| --- | ------- |
| 0 | 1 | African-American | Male | 25 - 45 |
| 1 | 1 | Native American | Female | Less than 25 |
```
aequitas-report --input compas_for_aequitas.csv
```
**Note:** Disparites are always defined in relation to a reference group. By default, Aequitas uses the majority group within each attribute as the reference group. [Defining a reference group](./config.html)
### The Bias Report output
The Bias Report produces a pdf that returns descriptive interpretation of the results along with three sets of tables.
* Fairness Measures Results
* Bias Metrics Results
* Group Metrics Results
Additionally, a csv is produced that contains the relevant data. More information about output [here](./output_data.html).
### Command Line output
In the command line you will see The Bias Report, which returns counts for each attribute by group and then computes various fairness metrics. This is the same information that is captured in the csv output.
```
___ _ __
/ | ___ ____ ___ __(_) /_____ ______
/ /| |/ _ \/ __ `/ / / / / __/ __ `/ ___/
/ ___ / __/ /_/ / /_/ / / /_/ /_/ (__ )
/_/ |_\___/\__, /\__,_/_/\__/\__,_/____/
/_/
____________________________________________________________________________
Bias and Fairness Audit Tool
____________________________________________________________________________
Welcome to Aequitas-Audit
Fairness measures requested: Statistical Parity,Impact Parity,FDR Parity,FPR Parity,FNR Parity,FOR Parity
model_id, score_thresholds 1 {'rank_abs': [3317]}
COUNTS::: race
African-American 3696
Asian 32
Caucasian 2454
Hispanic 637
Native American 18
Other 377
dtype: int64
COUNTS::: sex
Female 1395
Male 5819
dtype: int64
COUNTS::: age_cat
25 - 45 4109
Greater than 45 1576
Less than 25 1529
dtype: int64
audit: df shape from the crosstabs: (11, 26)
get_disparity_major_group()
number of rows after bias majority ref group: 11
Any NaN?: False
bias_df shape: (11, 38)
Fairness Threshold: 0.8
Fairness Measures: ['Statistical Parity', 'Impact Parity', 'FDR Parity', 'FPR Parity', 'FNR Parity', 'FOR Parity']
...
```
| github_jupyter |
# 定义目标
# 数据获取
## 训练数据
```
import pandas as pd
import numpy as np
import seaborn as sb
from matplotlib import pyplot as plt
%matplotlib inline
data_train = pd.read_csv("./data/train.csv")
data_train.head()
```
## 测试数据
```
data_test = pd.read_csv("./data/test.csv")
data_test.head()
```
# 数据理解
## 数据集名称
PassengerId 乘客ID<br>
Survived 获救与否(0死亡,1生存)<br>
Pclass 乘客等级(1/2/3等舱位)<br>
Name 乘客姓名<br>
Sex 性别<br>
Age 年龄<br>
SibSp 堂兄弟/妹个数<br>
Parch 父母与小孩个数<br>
Ticket 船票信息<br>
Fare 票价<br>
Cabin 客舱<br>
Embarked 登船港口
## 缺失值分析
```
data_train.info()
```
* <font color=red>Age和Cabin都有大幅度缺失值,Embarked有2个缺失值<font>
## 数值型特征分布
```
data_train.describe()
```
* <font color=red>Survived中值(mean)表明仅有0.383838的人获救<font>
* <font color=red>平均年龄在29.7岁(缺失值不计入)<font>
* <font color=red>乘客普遍为2\3等级,1等乘客数量少<font>
* <font color=red>绝大多数乘客为独生子女<font>
## 部分特征与标签的关系
```
import matplotlib.pyplot as plt
plt.rcParams['font.sans-serif'] = ['SimHei'] #用来正常显示中文标签
plt.rcParams['axes.unicode_minus'] = False #用来正常显示负号
fig = plt.figure(figsize=(18,8))
fig.set(alpha=20) # 设定图表颜色alpha参数(透明)
plt.subplot2grid((2,3),(0,0)) # 在一张大图里分列几个小图
data_train.Survived.value_counts().plot(kind='bar') # 对获救人数做条形图统计
plt.title(u"获救情况 (1为获救)")
plt.ylabel(u"人数")
plt.grid(True)
plt.subplot2grid((2,3),(0,1))
data_train.Pclass.value_counts().plot(kind="bar") # 对乘客等级数量做统计
plt.ylabel(u"人数")
plt.title(u"乘客等级分布")
plt.grid(True)
plt.subplot2grid((2,3),(0,2))
plt.scatter(data_train.Survived, data_train.Age)
plt.ylabel(u"年龄")
plt.grid(b=True, which='major', axis='y') # 查看按年龄分布的获救情况散点图
plt.title(u"按年龄看获救分布 (1为获救)")
plt.subplot2grid((2,3),(1,0), colspan=2)
data_train.Age[data_train.Pclass == 1].plot(kind='kde') # 查看各等级乘客年龄分布
data_train.Age[data_train.Pclass == 2].plot(kind='kde')
data_train.Age[data_train.Pclass == 3].plot(kind='kde')
plt.xlabel(u"年龄")# plots an axis lable
plt.ylabel(u"密度")
plt.title(u"各等级的乘客年龄分布")
plt.legend((u'头等舱', u'2等舱',u'3等舱'),loc='best') # 设置图例
plt.grid(True)
plt.subplot2grid((2,3),(1,2))
data_train.Embarked.value_counts().plot(kind='bar') # 查看各登船口岸上船人数
plt.title(u"各登船口岸上船人数")
plt.ylabel(u"人数")
plt.grid(True)
plt.show()
```
* <font color=red>获救人数刚刚300多一点<font>
* <font color=red>3等乘客人数最多,其次是1等<font>
* <font color=red>头等舱乘客年龄普遍年轻,其次是2等舱,最后是3等舱<font>
* <font color=red>明显口岸上岸人数不均等,可能跟地位有关<font>
## 查看按乘客等级分布的获救情况
```
import matplotlib.pyplot as plt
plt.rcParams['font.sans-serif'] = ['SimHei'] #用来正常显示中文标签
plt.rcParams['axes.unicode_minus'] = False #用来正常显示负号
#看看各乘客等级的获救情况
fig = plt.figure(figsize=(20,10))
fig.set(alpha=0.2) # 设定图表颜色alpha参数
Survived_0 = data_train.Pclass[data_train.Survived == 0].value_counts()
Survived_1 = data_train.Pclass[data_train.Survived == 1].value_counts()
df=pd.DataFrame({u'获救':Survived_1, u'未获救':Survived_0})
df.plot(kind='bar', stacked=True)
plt.title(u"各乘客等级的获救情况")
plt.xlabel(u"乘客等级")
plt.ylabel(u"人数")
plt.show()
```
* <font color=red>有钱就是希望大<font>
## 查看按性别分布的获救情况
```
import matplotlib.pyplot as plt
plt.rcParams['font.sans-serif'] = ['SimHei'] #用来正常显示中文标签
plt.rcParams['axes.unicode_minus'] = False #用来正常显示负号
#看看各乘客性别的获救情况
fig = plt.figure(figsize=(20,10))
fig.set(alpha=0.2)
Survived_0 = data_train.Sex[data_train.Survived == 0].value_counts()
Survived_1 = data_train.Sex[data_train.Survived == 1].value_counts()
df=pd.DataFrame({u'获救':Survived_1, u'未获救':Survived_0})
df.plot(kind='bar', stacked=True, rot=0)
plt.title(u"各乘客性别的获救情况")
plt.xlabel(u"乘客性别")
plt.ylabel(u"人数")
plt.show()
```
* <font color=red>听船长的话,妇女先走<font>
## 登录口岸划分的获救情况
```
import matplotlib.pyplot as plt
plt.rcParams['font.sans-serif'] = ['SimHei'] #用来正常显示中文标签
plt.rcParams['axes.unicode_minus'] = False #用来正常显示负号
#看看各乘客性别的获救情况
fig = plt.figure(figsize=(20,10))
fig.set(alpha=0.2)
Survived_1 = data_train.Embarked[data_train.Survived == 1].value_counts()
Survived_0 = data_train.Embarked[data_train.Survived == 0].value_counts()
df=pd.DataFrame({u'获救':Survived_1, u'未获救':Survived_0})
df.plot(kind='bar', stacked=True, rot=0)
plt.title(u"登录口岸划分的获救情况")
plt.xlabel(u"口岸")
plt.ylabel(u"人数")
plt.show()
```
* <font color=red>C口岸可能是给头等舱提供的入口<font>
## 查看按乘客等级分布的获救情况
```
# 然后我们再来看看各种舱级别情况下各性别的获救情况
fig=plt.figure(figsize=(16,10))
fig.set(alpha=0.5) # 设置图像透明度,无所谓
plt.title(u"根据舱等级和性别的获救情况")
ax1=fig.add_subplot(141)
data_train.Survived[data_train.Sex == 'female'][data_train.Pclass != 3].value_counts().plot(kind='bar', label="female highclass", color='#FA2479')
ax1.set_xticklabels([u"获救", u"未获救"], rotation=0)
ax1.legend([u"女性/高级舱"], loc='best')
ax2=fig.add_subplot(142, sharey=ax1)
data_train.Survived[data_train.Sex == 'female'][data_train.Pclass == 3].value_counts().plot(kind='bar', label='female, low class', color='pink')
ax2.set_xticklabels([u"未获救", u"获救"], rotation=0)
plt.legend([u"女性/低级舱"], loc='best')
ax3=fig.add_subplot(143, sharey=ax1)
data_train.Survived[data_train.Sex == 'male'][data_train.Pclass != 3].value_counts().plot(kind='bar', label='male, high class',color='lightblue')
ax3.set_xticklabels([u"未获救", u"获救"], rotation=0)
plt.legend([u"男性/高级舱"], loc='best')
ax4=fig.add_subplot(144, sharey=ax1)
data_train.Survived[data_train.Sex == 'male'][data_train.Pclass == 3].value_counts().plot(kind='bar', label='male low class', color='steelblue')
ax4.set_xticklabels([u"未获救", u"获救"], rotation=0)
plt.legend([u"男性/低级舱"], loc='best')
plt.show()
```
* <font color=red>有钱就是希望大<font>
## 家庭成员与获救的关系
```
g = data_train.groupby(['Parch','Survived'])
df = pd.DataFrame(g.count()['PassengerId'])
df
```
* <font color=red>没有太大的关联性<font>
## 查看兄弟姐妹人数和获救的关系
```
g = data_train.groupby(['SibSp','Survived'])
df = pd.DataFrame(g.count()['PassengerId'])
df
```
* <font color=red>家里兄弟人数少的,获救希望更大<font>
# 特征工程
## 对于缺失数据补全
```
from sklearn.ensemble import RandomForestRegressor
### 使用 RandomForestClassifier 填补缺失的年龄属性
def set_missing_ages(df):
# 把已有的数值型特征取出来丢进Random Forest Regressor中
age_df = df[['Age','Fare', 'Parch', 'SibSp', 'Pclass']]
# 乘客分成已知年龄和未知年龄两部分
known_age = age_df[age_df.Age.notnull()].as_matrix()
unknown_age = age_df[age_df.Age.isnull()].as_matrix()
# y即目标年龄
train_know_age_y = known_age[:, 0]
# X即特征属性值
train_know_age_X = known_age[:, 1:] # 这里与源代码比做了修改
# fit到RandomForestRegressor之中
rfr = RandomForestRegressor(random_state=0, n_estimators=2000, n_jobs=-1)
rfr.fit(train_know_age_X, train_know_age_y)
# 用得到的模型进行未知年龄结果预测
predictedAges = rfr.predict(unknown_age[:, 1:])
# 用得到的预测结果填补原缺失数据
df.loc[ (df.Age.isnull()), 'Age' ] = predictedAges
return df, rfr
def set_Cabin_type(df):
df.loc[ (df.Cabin.notnull()), 'Cabin' ] = "Yes"
df.loc[ (df.Cabin.isnull()), 'Cabin' ] = "No"
return df
data_train, rfr = set_missing_ages(data_train)
data_train = set_Cabin_type(data_train)
data_train
```
## 数据离散化
* <font color=red>因为逻辑回归建模时,需要输入的特征都是数值型特征,我们通常会先对类目型的特征因子化/one-hot编码<font>
```
# 因为逻辑回归建模时,需要输入的特征都是数值型特征
# 我们先对类目型的特征离散/因子化
# 以Cabin为例,原本一个属性维度,因为其取值可以是['yes','no'],而将其平展开为'Cabin_yes','Cabin_no'两个属性
# 原本Cabin取值为yes的,在此处的'Cabin_yes'下取值为1,在'Cabin_no'下取值为0
# 原本Cabin取值为no的,在此处的'Cabin_yes'下取值为0,在'Cabin_no'下取值为1
# 我们使用pandas的get_dummies来完成这个工作,并拼接在原来的data_train之上,如下所示
dummies_Cabin = pd.get_dummies(data_train['Cabin'], prefix= 'Cabin')
dummies_Embarked = pd.get_dummies(data_train['Embarked'], prefix= 'Embarked')
dummies_Sex = pd.get_dummies(data_train['Sex'], prefix= 'Sex')
dummies_Pclass = pd.get_dummies(data_train['Pclass'], prefix= 'Pclass')
df = pd.concat([data_train, dummies_Cabin, dummies_Embarked, dummies_Sex, dummies_Pclass], axis=1)
df.drop(['Pclass', 'Name', 'Sex', 'Ticket', 'Cabin', 'Embarked'], axis=1, inplace=True)
df
```
## 数据归一化
* <font color=red>我们还得做一些处理,仔细看看Age和Fare两个属性,乘客的数值幅度变化,也忒大了吧!!如果大家了解逻辑回归与梯度下降的话,会知道,各属性值之间scale差距太大,将对收敛速度造成几万点伤害值!甚至不收敛!所以我们先用scikit-learn里面的preprocessing模块对这俩货做一个scaling,所谓scaling,其实就是将一些变化幅度较大的特征化到[-1,1]之内。<font>
```
# 接下来我们要接着做一些数据预处理的工作,比如scaling,将一些变化幅度较大的特征化到[-1,1]之内
# 这样可以加速logistic regression的收敛
import sklearn.preprocessing as preprocessing
scaler = preprocessing.StandardScaler()
age_scale_param = scaler.fit(df['Age'].values.reshape(-1,1))
df['Age_scaled'] = scaler.fit_transform(df['Age'].values.reshape(-1,1), age_scale_param)
fare_scale_param = scaler.fit(df['Fare'].values.reshape(-1,1))
df['Fare_scaled'] = scaler.fit_transform(df['Fare'].values.reshape(-1,1), fare_scale_param)
df.head()
```
## 特征选择
```
# 我们把需要的feature字段取出来,转成numpy格式
train_df = df.filter(regex='Survived|Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass_.*')
train_np = train_df.as_matrix()
train_np
```
# 数据保存
# 模型选择
## 选择模型库训练获取模型
```
# 使用scikit-learn中的LogisticRegression建模
from sklearn import linear_model
# y即Survival结果
train_y = train_np[:, 0]
# X即特征属性值
train_X = train_np[:, 1:]
# fit 到 LogisticRegression 之中
clf = linear_model.LogisticRegression(C=1.0, penalty='l1', tol=1e-6)
clf.fit(train_X, train_y)
clf
```
## 对训练集做相同操作
```
from sklearn.ensemble import RandomForestRegressor
data_test.loc[ (data_test.Fare.isnull()), 'Fare' ] = 0
# 接着我们对test_data做和train_data中一致的特征变换
# 首先用同样的RandomForestRegressor模型填上丢失的年龄
tmp_df = data_test[['Age','Fare', 'Parch', 'SibSp', 'Pclass']]
null_age = tmp_df[data_test.Age.isnull()].as_matrix()
# 根据特征属性X预测年龄并补上
test_age_null_X = null_age[:, 1:]
predictedAges = rfr.predict(test_age_null_X)
data_test.loc[ (data_test.Age.isnull()), 'Age' ] = predictedAges
data_test = set_Cabin_type(data_test)
dummies_Cabin = pd.get_dummies(data_test['Cabin'], prefix= 'Cabin')
dummies_Embarked = pd.get_dummies(data_test['Embarked'], prefix= 'Embarked')
dummies_Sex = pd.get_dummies(data_test['Sex'], prefix= 'Sex')
dummies_Pclass = pd.get_dummies(data_test['Pclass'], prefix= 'Pclass')
df_test = pd.concat([data_test, dummies_Cabin, dummies_Embarked, dummies_Sex, dummies_Pclass], axis=1)
df_test.drop(['Pclass', 'Name', 'Sex', 'Ticket', 'Cabin', 'Embarked'], axis=1, inplace=True)
df_test['Age_scaled'] = scaler.fit_transform(df_test['Age'].values.reshape(-1,1), age_scale_param)
df_test['Fare_scaled'] = scaler.fit_transform(df_test['Fare'].values.reshape(-1,1), fare_scale_param)
df_test
```
**注意训练一次可以用,第二次会对"rfr.predict(X)"报错.其次源代码中应该修改**
* 特征选择
```
test = df_test.filter(regex='Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass_.*')
predictions = clf.predict(test)
result = pd.DataFrame({'PassengerId':data_test['PassengerId'].as_matrix(), 'Survived':predictions.astype(np.int32)})
result.to_csv("logistic_regression_predictions.csv", index=False)
pd.read_csv("logistic_regression_predictions.csv")
```
<font color=red>提交后结果为0.76555,恩,结果还不错。毕竟,这只是我们简单分析过后出的一个baseline系统嘛</font>
# 性能评估
## 画出学习曲线
性能可视化
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn.learning_curve import learning_curve
plt.rcParams['font.sans-serif'] = ['SimHei'] # 用来正常显示中文标签
plt.rcParams['axes.unicode_minus'] = False # 用来正常显示负号
# 用sklearn的learning_curve得到training_score和cv_score,使用matplotlib画出learning curve
def plot_learning_curve(estimator, title, X, y, ylim=None, cv=None, n_jobs=1,
train_sizes=np.linspace(.05, 1., 20), verbose=0, plot=True): # train_sizes=np.linspace(.05, 1., 20)做出切割选择
"""
画出data在某模型上的learning curve.
参数解释
----------
estimator : 你用的分类器。
title : 表格的标题。
X : 输入的feature,numpy类型
y : 输入的target vector
ylim : tuple格式的(ymin, ymax), 设定图像中纵坐标的最低点和最高点
cv : 做cross-validation的时候,数据分成的份数,其中一份作为cv集,其余n-1份作为training(默认为3份)
n_jobs : 并行的的任务数(默认1)
train_sizes : 对数据集进行切割的具体量
"""
train_sizes, train_scores, test_scores = learning_curve(
estimator, X, y, cv=cv, n_jobs=n_jobs, train_sizes=train_sizes, verbose=verbose)
# print(train_sizes)
# print(train_scores)
# print(test_scores)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
# print(train_scores_mean)
# print(train_scores_std)
# print(test_scores_mean)
# print(test_scores_std)
if plot:
plt.figure(figsize=(12,8))
plt.title(title)
if ylim is not None:
plt.ylim(*ylim)
plt.xlabel(u"训练样本数")
plt.ylabel(u"得分")
plt.gca().invert_yaxis() # 翻转坐标轴
plt.grid()
plt.fill_between(train_sizes, train_scores_mean - train_scores_std, train_scores_mean + train_scores_std,
alpha=0.1, color="b") # 颜色填充,下图中淡蓝色和淡红色的部分
plt.fill_between(train_sizes, test_scores_mean - test_scores_std, test_scores_mean + test_scores_std,
alpha=0.1, color="r") # 颜色填充,下图中淡蓝色和淡红色的部分
plt.plot(train_sizes, train_scores_mean, 'o-', color="b", label=u"训练集上得分")
plt.plot(train_sizes, test_scores_mean, 'o-', color="r", label=u"交叉验证集上得分")
plt.legend(loc="best")
plt.draw()
plt.gca().invert_yaxis()
plt.show()
midpoint = ((train_scores_mean[-1] + train_scores_std[-1]) + (test_scores_mean[-1] - test_scores_std[-1])) / 2
diff = (train_scores_mean[-1] + train_scores_std[-1]) - (test_scores_mean[-1] - test_scores_std[-1])
return midpoint, diff
plot_learning_curve(clf, u"学习曲线", train_X, train_y)
```
<font color=red>在实际数据上看,我们得到的learning curve没有理论推导的那么光滑哈,但是可以大致看出来,训练集和交叉验证集上的得分曲线走势还是符合预期的。<font><br>
<font color=red>目前的曲线看来,我们的model并不处于overfitting的状态(overfitting的表现一般是训练集上得分高,而交叉验证集上要低很多,中间的gap比较大)。因此我们可以再做些feature engineering的工作,添加一些新产出的特征或者组合特征到模型中。<font><br>
将变量传递给另一个页面
```
%store train_X
%store train_y
%store clf
```
## 其他模型构建
```
import numpy as np
import pandas as pd
from pandas import DataFrame
from patsy import dmatrices
import string
from operator import itemgetter
import json
from sklearn.ensemble import RandomForestClassifier
from sklearn.cross_validation import cross_val_score
from sklearn.pipeline import Pipeline
from sklearn.grid_search import GridSearchCV
from sklearn.cross_validation import train_test_split,StratifiedShuffleSplit,StratifiedKFold
from sklearn import preprocessing
from sklearn.metrics import classification_report
from sklearn.externals import joblib
##Read configuration parameters
train_file="./data/train.csv"
MODEL_PATH="./"
test_file="./data/test.csv"
SUBMISSION_PATH="./"
seed= 0
print(train_file,seed)
# 输出得分
def report(grid_scores, n_top=3):
top_scores = sorted(grid_scores, key=itemgetter(1), reverse=True)[:n_top]
for i, score in enumerate(top_scores):
print("Model with rank: {0}".format(i + 1))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
score.mean_validation_score,
np.std(score.cv_validation_scores)))
print("Parameters: {0}".format(score.parameters))
print("")
#清理和处理数据
def substrings_in_string(big_string, substrings):
for substring in substrings:
if str.find(big_string, substring) != -1:
return substring
print(big_string)
return np.nan
le = preprocessing.LabelEncoder()
enc=preprocessing.OneHotEncoder()
def clean_and_munge_data(df):
#处理缺省值
df.Fare = df.Fare.map(lambda x: np.nan if x==0 else x)
#处理一下名字,生成Title字段
title_list=['Mrs', 'Mr', 'Master', 'Miss', 'Major', 'Rev',
'Dr', 'Ms', 'Mlle','Col', 'Capt', 'Mme', 'Countess',
'Don', 'Jonkheer']
df['Title']=df['Name'].map(lambda x: substrings_in_string(x, title_list))
#处理特殊的称呼,全处理成mr, mrs, miss, master
def replace_titles(x):
title=x['Title']
if title in ['Mr','Don', 'Major', 'Capt', 'Jonkheer', 'Rev', 'Col']:
return 'Mr'
elif title in ['Master']:
return 'Master'
elif title in ['Countess', 'Mme','Mrs']:
return 'Mrs'
elif title in ['Mlle', 'Ms','Miss']:
return 'Miss'
elif title =='Dr':
if x['Sex']=='Male':
return 'Mr'
else:
return 'Mrs'
elif title =='':
if x['Sex']=='Male':
return 'Master'
else:
return 'Miss'
else:
return title
df['Title']=df.apply(replace_titles, axis=1)
#看看家族是否够大,咳咳
df['Family_Size']=df['SibSp']+df['Parch']
df['Family']=df['SibSp']*df['Parch']
df.loc[ (df.Fare.isnull())&(df.Pclass==1),'Fare'] =np.median(df[df['Pclass'] == 1]['Fare'].dropna())
df.loc[ (df.Fare.isnull())&(df.Pclass==2),'Fare'] =np.median( df[df['Pclass'] == 2]['Fare'].dropna())
df.loc[ (df.Fare.isnull())&(df.Pclass==3),'Fare'] = np.median(df[df['Pclass'] == 3]['Fare'].dropna())
df['Gender'] = df['Sex'].map( {'female': 0, 'male': 1} ).astype(int)
df['AgeFill']=df['Age']
mean_ages = np.zeros(4)
mean_ages[0]=np.average(df[df['Title'] == 'Miss']['Age'].dropna())
mean_ages[1]=np.average(df[df['Title'] == 'Mrs']['Age'].dropna())
mean_ages[2]=np.average(df[df['Title'] == 'Mr']['Age'].dropna())
mean_ages[3]=np.average(df[df['Title'] == 'Master']['Age'].dropna())
df.loc[ (df.Age.isnull()) & (df.Title == 'Miss') ,'AgeFill'] = mean_ages[0]
df.loc[ (df.Age.isnull()) & (df.Title == 'Mrs') ,'AgeFill'] = mean_ages[1]
df.loc[ (df.Age.isnull()) & (df.Title == 'Mr') ,'AgeFill'] = mean_ages[2]
df.loc[ (df.Age.isnull()) & (df.Title == 'Master') ,'AgeFill'] = mean_ages[3]
df['AgeCat']=df['AgeFill']
df.loc[ (df.AgeFill<=10) ,'AgeCat'] = 'child'
df.loc[ (df.AgeFill>60),'AgeCat'] = 'aged'
df.loc[ (df.AgeFill>10) & (df.AgeFill <=30) ,'AgeCat'] = 'adult'
df.loc[ (df.AgeFill>30) & (df.AgeFill <=60) ,'AgeCat'] = 'senior'
df.Embarked = df.Embarked.fillna('S')
df.loc[ df.Cabin.isnull()==True,'Cabin'] = 0.5
df.loc[ df.Cabin.isnull()==False,'Cabin'] = 1.5
df['Fare_Per_Person']=df['Fare']/(df['Family_Size']+1)
#Age times class
df['AgeClass']=df['AgeFill']*df['Pclass']
df['ClassFare']=df['Pclass']*df['Fare_Per_Person']
df['HighLow']=df['Pclass']
df.loc[ (df.Fare_Per_Person<8) ,'HighLow'] = 'Low'
df.loc[ (df.Fare_Per_Person>=8) ,'HighLow'] = 'High'
le.fit(df['Sex'] )
x_sex=le.transform(df['Sex'])
df['Sex']=x_sex.astype(np.float)
le.fit( df['Ticket'])
x_Ticket=le.transform( df['Ticket'])
df['Ticket']=x_Ticket.astype(np.float)
le.fit(df['Title'])
x_title=le.transform(df['Title'])
df['Title'] =x_title.astype(np.float)
le.fit(df['HighLow'])
x_hl=le.transform(df['HighLow'])
df['HighLow']=x_hl.astype(np.float)
le.fit(df['AgeCat'])
x_age=le.transform(df['AgeCat'])
df['AgeCat'] =x_age.astype(np.float)
le.fit(df['Embarked'])
x_emb=le.transform(df['Embarked'])
df['Embarked']=x_emb.astype(np.float)
df = df.drop(['PassengerId','Name','Age','Cabin'], axis=1) #remove Name,Age and PassengerId
return df
#读取数据
traindf=pd.read_csv(train_file)
##清洗数据
df=clean_and_munge_data(traindf)
########################################formula################################
formula_ml='Survived~Pclass+C(Title)+Sex+C(AgeCat)+Fare_Per_Person+Fare+Family_Size'
y_train, x_train = dmatrices(formula_ml, data=df, return_type='dataframe')
y_train = np.asarray(y_train).ravel()
print(y_train.shape,x_train.shape)
##选择训练和测试集
X_train, X_test, Y_train, Y_test = train_test_split(x_train, y_train, test_size=0.2,random_state=seed)
#初始化分类器
clf=RandomForestClassifier(n_estimators=500, criterion='entropy', max_depth=5, min_samples_split=2,
min_samples_leaf=1, max_features='auto', bootstrap=False, oob_score=False, n_jobs=1, random_state=seed,
verbose=0)
#grid search找到最好的参数
param_grid = dict( )
##创建分类pipeline
pipeline=Pipeline([('clf',clf)])
grid_search = GridSearchCV(pipeline, param_grid=param_grid, verbose=3,scoring='accuracy',\
cv=StratifiedShuffleSplit(Y_train, n_iter=10, test_size=0.2, train_size=None, \
random_state=seed)).fit(X_train, Y_train)
# 对结果打分
print("Best score: %0.3f" % grid_search.best_score_)
print(grid_search.best_estimator_)
report(grid_search.grid_scores_)
print('-----grid search end------------')
print('on all train set')
scores = cross_val_score(grid_search.best_estimator_, x_train, y_train,cv=3,scoring='accuracy')
print(scores.mean(),scores)
print ('on test set')
scores = cross_val_score(grid_search.best_estimator_, X_test, Y_test,cv=3,scoring='accuracy')
print(scores.mean(),scores)
# 对结果打分
print(classification_report(Y_train, grid_search.best_estimator_.predict(X_train) ))
print('test data')
print(classification_report(Y_test, grid_search.best_estimator_.predict(X_test) ))
model_file=MODEL_PATH+'model-rf.pkl'
joblib.dump(grid_search.best_estimator_, model_file)
```
# 模型优化
## 根据模型的系数调优
<font color=red>接下来,我们就该看看如何优化baseline系统了<br>
我们还有些特征可以再挖掘挖掘<br><br>
1. 比如说Name和Ticket两个属性被我们完整舍弃了(好吧,其实是一开始我们对于这种,每一条记录都是一个完全不同的值的属性,并没有很直接的处理方式)<br>
2. 比如说,我们想想,年龄的拟合本身也未必是一件非常靠谱的事情<br>
3. 另外,以我们的日常经验,小盆友和老人可能得到的照顾会多一些,这样看的话,年龄作为一个连续值,给一个固定的系数,似乎体现不出两头受照顾的实际情况,所以,说不定我们把年龄离散化,按区段分作类别属性会更合适一些<br>
那怎么样才知道,哪些地方可以优化,哪些优化的方法是promising的呢?<br>
是的<br><br>
要做交叉验证(cross validation)!<br>
要做交叉验证(cross validation)!<br>
要做交叉验证(cross validation)!<br><br>
重要的事情说3编!!!<br>
因为test.csv里面并没有Survived这个字段(好吧,这是废话,这明明就是我们要预测的结果),我们无法在这份数据上评定我们算法在该场景下的效果。。。<br>
我们通常情况下,这么做cross validation:把train.csv分成两部分,一部分用于训练我们需要的模型,另外一部分数据上看我们预测算法的效果。<br>
我们可以用scikit-learn的cross_validation来完成这个工作</font>
<font color=red>在此之前,咱们可以看看现在得到的模型的系数,因为系数和它们最终的判定能力强弱是正相关的</font>
```
%store train_df
%store clf
%store df
%store origin_data_train
%store data_train
pd.DataFrame({"columns":list(train_df.columns)[1:], "coef":list(clf.coef_.T)})
```
<font color=red>上面的系数和最后的结果是一个正相关的关系<br>
我们先看看那些权重绝对值非常大的feature,在我们的模型上:<br>
* Sex属性,如果是female会极大提高最后获救的概率,而male会很大程度拉低这个概率。
* Pclass属性,1等舱乘客最后获救的概率会上升,而乘客等级为3会极大地拉低这个概率。
* 有Cabin值会很大程度拉升最后获救概率(这里似乎能看到了一点端倪,事实上从最上面的有无Cabin记录的Survived分布图上看出,即使有Cabin记录的乘客也有一部分遇难了,估计这个属性上我们挖掘还不够)
* Age是一个负相关,意味着在我们的模型里,年龄越小,越有获救的优先权(还得回原数据看看这个是否合理)
* 有一个登船港口S会很大程度拉低获救的概率,另外俩港口压根就没啥作用(这个实际上非常奇怪,因为我们从之前的统计图上并没有看到S港口的获救率非常低,所以也许可以考虑把登船港口这个feature去掉试试)。
* 船票Fare有小幅度的正相关(并不意味着这个feature作用不大,有可能是我们细化的程度还不够,举个例子,说不定我们得对它离散化,再分至各个乘客等级上?)
噢啦,观察完了,我们现在有一些想法了,但是怎么样才知道,哪些优化的方法是promising的呢?<br>
恩,要靠交叉验证
```
from sklearn import cross_validation
# 简单看看打分情况
clf = linear_model.LogisticRegression(C=1.0, penalty='l1', tol=1e-6)
all_data = df.filter(regex='Survived|Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass_.*')
X = all_data.as_matrix()[:,1:]
y = all_data.as_matrix()[:,0]
print(cross_validation.cross_val_score(clf, X, y, cv=5))
# 分割数据
split_train, split_cv = cross_validation.train_test_split(df, test_size=0.3, random_state=0) # random_state保存随机切割的数据量,确保下次使用还能切分成一样的数据
train_df = split_train.filter(regex='Survived|Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass_.*')
# 生成模型
clf = linear_model.LogisticRegression(C=1.0, penalty='l1', tol=1e-6)
clf.fit(train_df.as_matrix()[:,1:], train_df.as_matrix()[:,0])
# 对cross validation数据进行预测
cv_df = split_cv.filter(regex='Survived|Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass_.*')
predictions = clf.predict(cv_df.as_matrix()[:,1:])
split_cv[predictions != cv_df.as_matrix()[:,0]]
# 去除预测错误的case看原始dataframe数据
#split_cv['PredictResult'] = predictions
origin_data_train = pd.read_csv("./data/train.csv")
bad_cases = origin_data_train.loc[origin_data_train['PassengerId'].isin(split_cv[predictions != cv_df.as_matrix()[:,0]]['PassengerId'].values)]
bad_cases
```
<font color=red>对比bad case,我们仔细看看我们预测错的样本,到底是哪些特征有问题,咱们处理得还不够细?<br>
我们随便列一些可能可以做的优化操作:<br>
* Age属性不使用现在的拟合方式,而是根据名称中的『Mr』『Mrs』『Miss』等的平均值进行填充。
* Age不做成一个连续值属性,而是使用一个步长进行离散化,变成离散的类目feature。
* Cabin再细化一些,对于有记录的Cabin属性,我们将其分为前面的字母部分(我猜是位置和船层之类的信息) 和 后面的数字部分(应该是房间号,有意思的事情是,如果你仔细看看原始数据,你会发现,这个值大的情况下,似乎获救的可能性高一些)。
* **Pclass和Sex俩太重要了,我们试着用它们去组出一个组合属性来试试,这也是另外一种程度的细化。**
* 单加一个Child字段,Age<=12的,设为1,其余为0(你去看看数据,确实小盆友优先程度很高啊)
* 如果名字里面有『Mrs』,而Parch>1的,我们猜测她可能是一个母亲,应该获救的概率也会提高,因此可以多加一个Mother字段,此种情况下设为1,其余情况下设为0
* 登船港口可以考虑先去掉试试(Q和C本来就没权重,S有点诡异)
* 把堂兄弟/兄妹 和 Parch 还有自己 个数加在一起组一个Family_size字段(考虑到大家族可能对最后的结果有影响)
* Name是一个我们一直没有触碰的属性,我们可以做一些简单的处理,比如说男性中带某些字眼的(‘Capt’, ‘Don’, ‘Major’, ‘Sir’)可以统一到一个Title,女性也一样。
大家接着往下挖掘,可能还可以想到更多可以细挖的部分。我这里先列这些了,然后我们可以使用手头上的”train_df”和”cv_df”开始试验这些feature engineering的tricks是否有效了。
```
data_train[data_train['Name'].str.contains("Major")]
data_train = pd.read_csv("./data/train.csv")
data_train['Sex_Pclass'] = data_train.Sex + "_" + data_train.Pclass.map(str) # 对每个Pclass执行str函数(数字变为字符串)
from sklearn.ensemble import RandomForestRegressor
# 使用 RandomForestClassifier 填补缺失的年龄属性
def set_missing_ages(df):
# 把已有的数值型特征取出来丢进Random Forest Regressor中
age_df = df[['Age','Fare', 'Parch', 'SibSp', 'Pclass']]
# 乘客分成已知年龄和未知年龄两部分
known_age = age_df[age_df.Age.notnull()].as_matrix()
unknown_age = age_df[age_df.Age.isnull()].as_matrix()
# y即目标年龄
y = known_age[:, 0]
# X即特征属性值
X = known_age[:, 1:]
# fit到RandomForestRegressor之中
rfr = RandomForestRegressor(random_state=0, n_estimators=2000, n_jobs=-1)
rfr.fit(X, y)
# 用得到的模型进行未知年龄结果预测
predictedAges = rfr.predict(unknown_age[:, 1:])
# 用得到的预测结果填补原缺失数据
df.loc[ (df.Age.isnull()), 'Age' ] = predictedAges
return df, rfr
def set_Cabin_type(df):
df.loc[ (df.Cabin.notnull()), 'Cabin' ] = "Yes"
df.loc[ (df.Cabin.isnull()), 'Cabin' ] = "No"
return df
data_train, rfr = set_missing_ages(data_train)
data_train = set_Cabin_type(data_train)
dummies_Cabin = pd.get_dummies(data_train['Cabin'], prefix= 'Cabin')
dummies_Embarked = pd.get_dummies(data_train['Embarked'], prefix= 'Embarked')
dummies_Sex = pd.get_dummies(data_train['Sex'], prefix= 'Sex')
dummies_Pclass = pd.get_dummies(data_train['Pclass'], prefix= 'Pclass')
dummies_Sex_Pclass = pd.get_dummies(data_train['Sex_Pclass'], prefix= 'Sex_Pclass')
df = pd.concat([data_train, dummies_Cabin, dummies_Embarked, dummies_Sex, dummies_Pclass, dummies_Sex_Pclass], axis=1)
df.drop(['Pclass', 'Name', 'Sex', 'Ticket', 'Cabin', 'Embarked', 'Sex_Pclass'], axis=1, inplace=True)
import sklearn.preprocessing as preprocessing
scaler = preprocessing.StandardScaler()
age_scale_param = scaler.fit(df['Age'].values.reshape(-1,1))
df['Age_scaled'] = scaler.fit_transform(df['Age'].values.reshape(-1,1), age_scale_param)
fare_scale_param = scaler.fit(df['Fare'].values.reshape(-1,1))
df['Fare_scaled'] = scaler.fit_transform(df['Fare'].values.reshape(-1,1), fare_scale_param)
from sklearn import linear_model
train_df = df.filter(regex='Survived|Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass.*')
train_np = train_df.as_matrix()
# y即Survival结果
y = train_np[:, 0]
# X即特征属性值
X = train_np[:, 1:]
# fit到RandomForestRegressor之中
clf = linear_model.LogisticRegression(C=1.0, penalty='l1', tol=1e-6)
clf.fit(X, y)
clf
```
对test也做同样的变换
```
data_test = pd.read_csv("./data/test.csv")
data_test.loc[ (data_test.Fare.isnull()), 'Fare' ] = 0
data_test['Sex_Pclass'] = data_test.Sex + "_" + data_test.Pclass.map(str)
# 接着我们对test_data做和train_data中一致的特征变换
# 首先用同样的RandomForestRegressor模型填上丢失的年龄
tmp_df = data_test[['Age','Fare', 'Parch', 'SibSp', 'Pclass']]
null_age = tmp_df[data_test.Age.isnull()].as_matrix()
# 根据特征属性X预测年龄并补上
X = null_age[:, 1:]
predictedAges = rfr.predict(X)
data_test.loc[ (data_test.Age.isnull()), 'Age' ] = predictedAges
data_test = set_Cabin_type(data_test)
dummies_Cabin = pd.get_dummies(data_test['Cabin'], prefix= 'Cabin')
dummies_Embarked = pd.get_dummies(data_test['Embarked'], prefix= 'Embarked')
dummies_Sex = pd.get_dummies(data_test['Sex'], prefix= 'Sex')
dummies_Pclass = pd.get_dummies(data_test['Pclass'], prefix= 'Pclass')
dummies_Sex_Pclass = pd.get_dummies(data_test['Sex_Pclass'], prefix= 'Sex_Pclass')
df_test = pd.concat([data_test, dummies_Cabin, dummies_Embarked, dummies_Sex, dummies_Pclass, dummies_Sex_Pclass], axis=1)
df_test.drop(['Pclass', 'Name', 'Sex', 'Ticket', 'Cabin', 'Embarked', 'Sex_Pclass'], axis=1, inplace=True)
df_test['Age_scaled'] = scaler.fit_transform(df_test['Age'].values.reshape(-1,1), age_scale_param)
df_test['Fare_scaled'] = scaler.fit_transform(df_test['Fare'].values.reshape(-1,1), fare_scale_param)
df_test
test = df_test.filter(regex='Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass.*')
predictions = clf.predict(test)
result = pd.DataFrame({'PassengerId':data_test['PassengerId'].as_matrix(), 'Survived':predictions.astype(np.int32)})
result.to_csv("logistic_regression_predictions2.csv", index=False)
pd.read_csv("logistic_regression_predictions2.csv")
```
<font color=red>一般做到后期,咱们要进行模型优化的方法就是模型融合啦<br>
先解释解释啥叫模型融合哈,我们还是举几个例子直观理解一下好了。<br><br>
大家都看过知识问答的综艺节目中,求助现场观众时候,让观众投票,最高的答案作为自己的答案的形式吧,每个人都有一个判定结果,最后我们相信答案在大多数人手里。<br>
再通俗一点举个例子。你和你班某数学大神关系好,每次作业都『模仿』他的,于是绝大多数情况下,他做对了,你也对了。突然某一天大神脑子犯糊涂,手一抖,写错了一个数,于是…恩,你也只能跟着错了。 <br>
我们再来看看另外一个场景,你和你班5个数学大神关系都很好,每次都把他们作业拿过来,对比一下,再『自己做』,那你想想,如果哪天某大神犯糊涂了,写错了,but另外四个写对了啊,那你肯定相信另外4人的是正确答案吧?<br>
最简单的模型融合大概就是这么个意思,比如分类问题,当我们手头上有一堆在同一份数据集上训练得到的分类器(比如logistic regression,SVM,KNN,random forest,神经网络),那我们让他们都分别去做判定,然后对结果做投票统计,取票数最多的结果为最后结果。<br>
bingo,问题就这么完美的解决了。<br>
模型融合可以比较好地缓解,训练过程中产生的过拟合问题,从而对于结果的准确度提升有一定的帮助。<br>
话说回来,回到我们现在的问题。你看,我们现在只讲了logistic regression,如果我们还想用这个融合思想去提高我们的结果,我们该怎么做呢?<br>
既然这个时候模型没得选,那咱们就在数据上动动手脚咯。大家想想,如果模型出现过拟合现在,一定是在我们的训练上出现拟合过度造成的对吧。<br>
那我们干脆就不要用全部的训练集,每次取训练集的一个subset,做训练,这样,我们虽然用的是同一个机器学习算法,但是得到的模型却是不一样的;同时,因为我们没有任何一份子数据集是全的,因此即使出现过拟合,也是在子训练集上出现过拟合,而不是全体数据上,这样做一个融合,可能对最后的结果有一定的帮助。对,这就是常用的Bagging。<br>
我们用scikit-learn里面的Bagging来完成上面的思路,过程非常简单。代码如下:<br><br><font>
## 使用模型融合进行优化
```
from sklearn.ensemble import BaggingRegressor
train_df = df.filter(regex='Survived|Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass.*|Mother|Child|Family|Title')
train_np = train_df.as_matrix()
# y即Survival结果
y = train_np[:, 0]
# X即特征属性值
X = train_np[:, 1:]
# fit到BaggingRegressor之中
clf = linear_model.LogisticRegression(C=1.0, penalty='l1', tol=1e-6)
bagging_clf = BaggingRegressor(clf, n_estimators=10, max_samples=0.8, max_features=1.0, bootstrap=True, bootstrap_features=False, n_jobs=-1)
bagging_clf.fit(X, y)
test = df_test.filter(regex='Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass.*|Mother|Child|Family|Title')
predictions = bagging_clf.predict(test)
result = pd.DataFrame({'PassengerId':data_test['PassengerId'].as_matrix(), 'Survived':predictions.astype(np.int32)})
result.to_csv("logistic_regression_predictions3.csv", index=False)
pd.read_csv("logistic_regression_predictions2.csv")
```
# 模型使用
## 模型保存输出
```
from sklearn.externals import joblib
model_file=MODEL_PATH+'model-rf.pkl'
joblib.dump(grid_search.best_estimator_, model_file)
```
| github_jupyter |
```
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '2'
import bert
from bert import run_classifier
from bert import optimization
from bert import tokenization
from bert import modeling
import numpy as np
import tensorflow as tf
import pandas as pd
from tqdm import tqdm
# !wget https://github.com/huseinzol05/Malaya/raw/master/pretrained-model/preprocess/sp10m.cased.bert.model
# !wget https://github.com/huseinzol05/Malaya/raw/master/pretrained-model/preprocess/sp10m.cased.bert.vocab
from prepro_utils import preprocess_text, encode_ids, encode_pieces
import sentencepiece as spm
sp_model = spm.SentencePieceProcessor()
sp_model.Load('sp10m.cased.bert.model')
with open('sp10m.cased.bert.vocab') as fopen:
v = fopen.read().split('\n')[:-1]
v = [i.split('\t') for i in v]
v = {i[0]: i[1] for i in v}
class Tokenizer:
def __init__(self, v):
self.vocab = v
pass
def tokenize(self, string):
return encode_pieces(sp_model, string, return_unicode=False, sample=False)
def convert_tokens_to_ids(self, tokens):
return [sp_model.PieceToId(piece) for piece in tokens]
def convert_ids_to_tokens(self, ids):
return [sp_model.IdToPiece(i) for i in ids]
tokenizer = Tokenizer(v)
from glob import glob
import json
left, right, label = [], [], []
for file in glob('../text-similarity/*k.json'):
with open(file) as fopen:
x = json.load(fopen)
for i in x:
splitted = i[0].split(' <> ')
if len(splitted) != 2:
continue
left.append(splitted[0])
right.append(splitted[1])
label.append(i[1])
l = {'contradiction': 0, 'entailment': 1}
snli = glob('../text-similarity/part*.json')
for file in snli:
with open(file) as fopen:
x = json.load(fopen)
for i in x:
splitted = i[1].split(' <> ')
if len(splitted) != 2:
continue
if i[0] not in l:
continue
left.append(splitted[0])
right.append(splitted[1])
try:
label.append(l[i[0]])
except Exception as e:
print(e)
print(splitted, i[0])
mnli = glob('../text-similarity/translated-*.json')
mnli
for file in mnli:
with open(file) as fopen:
x = json.load(fopen)
for i in x:
if len(i) != 3:
continue
splitted = i[2].split(' <> ')
if len(splitted) != 3:
continue
if i[1] not in l:
continue
left.append(splitted[0])
right.append(splitted[1])
try:
label.append(l[i[1]])
except Exception as e:
print(e)
print(splitted, i)
BERT_INIT_CHKPNT = 'bert-base-2020-03-19/model.ckpt-2000002'
BERT_CONFIG = 'bert-base-2020-03-19/bert_config.json'
from tqdm import tqdm
MAX_SEQ_LENGTH = 200
def _truncate_seq_pair(tokens_a, tokens_b, max_length):
while True:
total_length = len(tokens_a) + len(tokens_b)
if total_length <= max_length:
break
if len(tokens_a) > len(tokens_b):
tokens_a.pop()
else:
tokens_b.pop()
def get_inputs(left, right):
input_ids, input_masks, segment_ids = [], [], []
for i in tqdm(range(len(left))):
tokens_a = tokenizer.tokenize(left[i])
tokens_b = tokenizer.tokenize(right[i])
_truncate_seq_pair(tokens_a, tokens_b, MAX_SEQ_LENGTH - 3)
tokens = []
segment_id = []
tokens.append("[CLS]")
segment_id.append(0)
for token in tokens_a:
tokens.append(token)
segment_id.append(0)
tokens.append("[SEP]")
segment_id.append(0)
for token in tokens_b:
tokens.append(token)
segment_id.append(1)
tokens.append("[SEP]")
segment_id.append(1)
input_id = tokenizer.convert_tokens_to_ids(tokens)
input_mask = [1] * len(input_id)
while len(input_id) < MAX_SEQ_LENGTH:
input_id.append(0)
input_mask.append(0)
segment_id.append(0)
input_ids.append(input_id)
input_masks.append(input_mask)
segment_ids.append(segment_id)
return input_ids, input_masks, segment_ids
input_ids, input_masks, segment_ids = get_inputs(left, right)
bert_config = modeling.BertConfig.from_json_file(BERT_CONFIG)
epoch = 20
batch_size = 60
warmup_proportion = 0.1
num_train_steps = int(len(left) / batch_size * epoch)
num_warmup_steps = int(num_train_steps * warmup_proportion)
class Model:
def __init__(
self,
dimension_output,
learning_rate = 2e-5,
training = True
):
self.X = tf.placeholder(tf.int32, [None, None])
self.segment_ids = tf.placeholder(tf.int32, [None, None])
self.input_masks = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None])
model = modeling.BertModel(
config=bert_config,
is_training=training,
input_ids=self.X,
input_mask=self.input_masks,
token_type_ids=self.segment_ids,
use_one_hot_embeddings=False)
output_layer = model.get_pooled_output()
self.logits = tf.layers.dense(output_layer, dimension_output)
self.logits = tf.identity(self.logits, name = 'logits')
self.cost = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
logits = self.logits, labels = self.Y
)
)
self.optimizer = optimization.create_optimizer(self.cost, learning_rate,
num_train_steps, num_warmup_steps, False)
correct_pred = tf.equal(
tf.argmax(self.logits, 1, output_type = tf.int32), self.Y
)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
dimension_output = 2
learning_rate = 2e-5
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(
dimension_output,
learning_rate
)
sess.run(tf.global_variables_initializer())
var_lists = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, scope = 'bert')
saver = tf.train.Saver(var_list = var_lists)
saver.restore(sess, BERT_INIT_CHKPNT)
from sklearn.model_selection import train_test_split
train_input_ids, test_input_ids, train_input_masks, test_input_masks, train_segment_ids, test_segment_ids, train_Y, test_Y = train_test_split(
input_ids, input_masks, segment_ids, label, test_size = 0.2)
from tqdm import tqdm
import time
EARLY_STOPPING, CURRENT_CHECKPOINT, CURRENT_ACC, EPOCH = 1, 0, 0, 0
while True:
lasttime = time.time()
if CURRENT_CHECKPOINT == EARLY_STOPPING:
print('break epoch:%d\n' % (EPOCH))
break
train_acc, train_loss, test_acc, test_loss = [], [], [], []
pbar = tqdm(
range(0, len(train_input_ids), batch_size), desc = 'train minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(train_input_ids))
batch_x = train_input_ids[i: index]
batch_masks = train_input_masks[i: index]
batch_segment = train_segment_ids[i: index]
batch_y = train_Y[i: index]
acc, cost, _ = sess.run(
[model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
model.segment_ids: batch_segment,
model.input_masks: batch_masks
},
)
assert not np.isnan(cost)
train_loss.append(cost)
train_acc.append(acc)
pbar.set_postfix(cost = cost, accuracy = acc)
pbar = tqdm(range(0, len(test_input_ids), batch_size), desc = 'test minibatch loop')
for i in pbar:
index = min(i + batch_size, len(test_input_ids))
batch_x = test_input_ids[i: index]
batch_masks = test_input_masks[i: index]
batch_segment = test_segment_ids[i: index]
batch_y = test_Y[i: index]
acc, cost = sess.run(
[model.accuracy, model.cost],
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
model.segment_ids: batch_segment,
model.input_masks: batch_masks
},
)
test_loss.append(cost)
test_acc.append(acc)
pbar.set_postfix(cost = cost, accuracy = acc)
train_loss = np.mean(train_loss)
train_acc = np.mean(train_acc)
test_loss = np.mean(test_loss)
test_acc = np.mean(test_acc)
if test_acc > CURRENT_ACC:
print(
'epoch: %d, pass acc: %f, current acc: %f'
% (EPOCH, CURRENT_ACC, test_acc)
)
CURRENT_ACC = test_acc
CURRENT_CHECKPOINT = 0
else:
CURRENT_CHECKPOINT += 1
print('time taken:', time.time() - lasttime)
print(
'epoch: %d, training loss: %f, training acc: %f, valid loss: %f, valid acc: %f\n'
% (EPOCH, train_loss, train_acc, test_loss, test_acc)
)
EPOCH += 1
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'bert-base-similarity/model.ckpt')
dimension_output = 2
learning_rate = 2e-5
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(
dimension_output,
learning_rate,
training = False
)
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver(tf.trainable_variables())
saver.restore(sess, 'bert-base-similarity/model.ckpt')
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or 'logits' in n.name
or 'alphas' in n.name
or 'self/Softmax' in n.name)
and 'adam' not in n.name
and 'beta' not in n.name
and 'global_step' not in n.name
]
)
strings.split(',')
real_Y, predict_Y = [], []
pbar = tqdm(
range(0, len(test_input_ids), batch_size), desc = 'validation minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(test_input_ids))
batch_x = test_input_ids[i: index]
batch_masks = test_input_masks[i: index]
batch_segment = test_segment_ids[i: index]
batch_y = test_Y[i: index]
predict_Y += np.argmax(sess.run(model.logits,
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
model.segment_ids: batch_segment,
model.input_masks: batch_masks
},
), 1, ).tolist()
real_Y += batch_y
from sklearn import metrics
print(
metrics.classification_report(
real_Y, predict_Y, target_names = ['not similar', 'similar'],
digits = 5
)
)
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('bert-base-similarity', strings)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.