text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# Import helpful libraries
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import numpy as np # linear algebra
import matplotlib.pyplot as plt
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn import feature_extraction, linear_model, model_selection, preprocessing
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
# Load the data, and separate the target
f='../input/newsdata/data/Fake.csv'
t='../input/newsdata/data/True.csv'
fake = pd.read_csv(f)
true = pd.read_csv(t)
# Get the shape of fake and true data
fake.shape
true.shape
# Preview the first 5 rows of the fake data
fake.head()
# Preview the first 5 rows of the true data
true.head()
# Add flag to track fake and real
fake['target'] = 'fake'
true['target'] = 'true'
# Concatenate dataframes
data = pd.concat([fake, true]).reset_index(drop = True)
data.shape
# Shuffle the data
from sklearn.utils import shuffle
data = shuffle(data)
data = data.reset_index(drop=True)
# preview first 5 rows of data
data.head()
# Removing the date and title column (we won't use it for the analysis)
data.drop(["date"],axis=1,inplace=True)
data.drop(["title"],axis=1,inplace=True)
data.head()
# Removing punctuation
import string
def punctuation_removal(text):
all_list = [char for char in text if char not in string.punctuation]
clean_str = ''.join(all_list)
return clean_str
data['text'] = data['text'].apply(punctuation_removal)
data.head()
# Convert to lowercase
data['text'] = data['text'].apply(lambda x: x.lower())
data.head()
# Removing stopwords
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
stop = stopwords.words('english')
data['text'] = data['text'].apply(lambda x: ' '.join([word for word in x.split() if word not in (stop)]))
data.head()
# DATA EXPLORATION
# How many articles per subject?
print(data.groupby(['subject'])['text'].count())
data.groupby(['subject'])['text'].count().plot(kind="bar")
plt.show()
# How many fake and real articles?
print(data.groupby(['target'])['text'].count())
data.groupby(['target'])['text'].count().plot(kind="bar")
plt.show()
# Word cloud for fake news
from wordcloud import WordCloud
fake_data = data[data["target"] == "fake"]
all_words = ' '.join([text for text in fake_data.text])
wordcloud = WordCloud(width= 800, height= 500,
max_font_size = 110,
collocations = False).generate(all_words)
plt.figure(figsize=(12,10))
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
# Word cloud for real news
from wordcloud import WordCloud
real_data = data[data["target"] == "true"]
all_words = ' '.join([text for text in real_data.text])
wordcloud = WordCloud(width= 800, height= 500,
max_font_size = 110,
collocations = False).generate(all_words)
plt.figure(figsize=(12,10))
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
# Most frequent words counter (Code adapted from https://www.kaggle.com/rodolfoluna/fake-news-detector)
from nltk import tokenize
token_space = tokenize.WhitespaceTokenizer()
def counter(text, column_text, quantity):
all_words = ' '.join([text for text in text[column_text]])
token_phrase = token_space.tokenize(all_words)
frequency = nltk.FreqDist(token_phrase)
df_frequency = pd.DataFrame({"Word": list(frequency.keys()),
"Frequency": list(frequency.values())})
df_frequency = df_frequency.nlargest(columns = "Frequency", n = quantity)
plt.figure(figsize=(12,8))
ax = sns.barplot(data = df_frequency, x = "Word", y = "Frequency", color = 'blue')
ax.set(ylabel = "Count")
plt.xticks(rotation='vertical')
plt.show()
# Most frequent words in fake news
counter(data[data["target"] == "fake"], "text", 20)
# Most frequent words in real news
counter(data[data["target"] == "true"], "text", 20)
#MODELLING
# Function to plot the confusion matrix
from sklearn import metrics
import itertools
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# Split the data
X_train,X_test,y_train,y_test = train_test_split(data['text'], data.target, test_size=0.2, random_state=42)
#Tryung different modelling technique to get better prediction
#LOGISTIC REGRESSION
# Vectorizing and applying TF-IDF
from sklearn.linear_model import LogisticRegression
pipe = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('model_1', LogisticRegression())])
# Fitting the model
model_1 = pipe.fit(X_train, y_train)
# Accuracy
prediction = model_1.predict(X_test)
print("Accuracy using Logistic Regression: {}%".format(round(accuracy_score(y_test, prediction)*100,2)))
model1_cm=metrics.confusion_matrix(y_test, prediction)
plot_confusion_matrix(model1_cm, classes=['Fake', 'Real'])
#RANDOM FOREST CLASSIFIER
from sklearn.ensemble import RandomForestClassifier
pipe = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('model_2', RandomForestClassifier(n_estimators=50, criterion="entropy"))])
#fitting the model
model_2 = pipe.fit(X_train, y_train)
#prediction and accuracy
prediction = model_2.predict(X_test)
print("Accuracy using Random Forest Classifier: {}%".format(round(accuracy_score(y_test, prediction)*100,2)))
model2_cm = metrics.confusion_matrix(y_test, prediction)
plot_confusion_matrix(model2_cm, classes=['Fake', 'Real'])
#DECISION TREE CLASSIFIER
from sklearn.tree import DecisionTreeClassifier
# Vectorizing and applying TF-IDF
pipe = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('model_3', DecisionTreeClassifier(criterion= 'entropy',
max_depth = 20,
splitter='best',
random_state=42))])
# Fitting the model
model_3 = pipe.fit(X_train, y_train)
# Accuracy
prediction = model_3.predict(X_test)
print("Accuracy using Decision Tree Classifier: {}%".format(round(accuracy_score(y_test, prediction)*100,2)))
model3_cm = metrics.confusion_matrix(y_test, prediction)
plot_confusion_matrix(model3_cm, classes=['Fake', 'Real'])
# Run the code to save predictions in the format used for competition scoring
output = pd.DataFrame(model3_cm)
output.to_csv('submission.csv', index=False)
```
| github_jupyter |
# exma quick start
In this tutorial we will take the typical molecular dynamics case of a Lennard-Jones (LJ) fluid, in its solid phase and in its liquid phase, and we will see how to obtain different properties of them using this library.
This first part of the code will be common to all three sections. We are going to import the necessary libraries and define the variables that we will use next.
```
import matplotlib.pyplot as plt
import numpy as np
```
We are going to define the _strings_ that will direct us to where the files with the paths are located (assuming we are in _exma/docs/source_).
```
fsolid = "tutorial_data/solid.xyz"
fliquid = "tutorial_data/liquid.xyz"
```
We leave these variables defined as strings and we do not read them since `exma` is in charge of reading xyz or lammpstrj files.
These trajectories were generated with a [homemade code](https://github.com/fernandezfran/tiny_md), in both cases there are 201 frames and 500 atoms. Next we define the parameters of the simulation cell for each case (this is necessary since the file is xyz, if it was lammpstrj we could skip this cell).
```
solid_box = np.full(3, 7.46901)
liquid_box = np.full(3, 8.54988)
```
In this case the distances are given in Lennard-Jones units.
## Mean Square Displacement (MSD)
The mean square displacement (MSD) is a measure of the deviation of the position of the particles with respect to a reference positions over time. From it, it is possible to obtain, through a linear regression, the trace diffusion coefficient. For more information you can start reading the Wikipedia article of [mean square displacement](https://en.wikipedia.org/wiki/Mean_squared_displacement).
We start by importing the `MeanSquareDisplacement` class from exma.
```
from exma import MeanSquareDisplacement
```
As in every Pair Analyzer, we have dedicated `calculate`, `plot` and `save` methods, the latter will not be used in this tutorial but it is useful when we want to save the results in a different file without the need to re-run the calculations which, for long time simulations, can be demanding.
```
# for both structures we discard the first 10 equilibration frames
solid_msd = MeanSquareDisplacement(fsolid, 0.05, "Ar", start=10, xyztype="image")
liquid_msd = MeanSquareDisplacement(fliquid, 0.05, "Ar", start=10, xyztype="image")
```
At this point we just instantiate the class, we are able to calculate, for which it is necessary to pass the optional argument of the cell length in each direction.
```
solid_msd.calculate(box=solid_box)
liquid_msd.calculate(box=solid_box)
```
We can see directly from the numbers how in the liquid phase a larger quadratic displacement is obtained already in the first steps. But to be more illustrative, we can plot both curves on the same graph, using `plt.gca()` of matplotlib.
```
ax = plt.gca()
# we pass the same axis to both plots and define the labels offered by the wrapper to the plot function
solid_msd.plot(ax=ax, plot_kws={"label": "solid"})
liquid_msd.plot(ax=ax, plot_kws={"label": "liquid"})
# we add the legend to the plot
plt.legend()
```
We obtain the expected response for a LJ fluid where the liquid phase diffuses with the expected linear behaivor and the solid phase does not diffuse.
## Radial Distribution Function (RDF)
The pair radial distribution function (RDF), _g(r)_, characterizes the local structure of a fluid, and describes the probability to find an atom in a shell at distance _r_ from a reference atom. This quantity is calculated as the ratio between the average density at distance _r_ from the reference atom and the density at that same distance of an ideal gas. For more information you can start reading the Wikipedia article of [radial distribution function](https://en.wikipedia.org/wiki/Radial_distribution_function).
We start importing the `RadialDistributionFunction` class from exma.
```
from exma import RadialDistributionFunction
```
And the mode of use is quite analogous to that of the MSD, some parameters of the inizializer are changed.
```
# for both structures we discard the first 10 equilibration frames
solid_rdf = RadialDistributionFunction(fsolid, "Ar", "Ar", start=10, rmax=solid_box[0] / 2)
liquid_rdf = RadialDistributionFunction(fliquid, "Ar", "Ar", start=10, rmax=liquid_box[0] / 2)
```
In this case we declare that the RDF is calculated up to half the distance from the box, i.e. atoms at a greater distance are ignored.
```
solid_rdf.calculate(box=solid_box)
liquid_rdf.calculate(box=liquid_box)
```
As before, we can obtain the corresponding graph.
```
ax = plt.gca()
# we pass the same axis to both plots and define the labels offered by the wrapper to the plot function
solid_rdf.plot(ax=ax, plot_kws={"label": "solid"})
liquid_rdf.plot(ax=ax, plot_kws={"label": "liquid"})
# we add the legend to the plot
plt.legend()
```
We get the expected results. For the solid phase we have the defined peaks of an fcc crystal with noise given by the temperature and for the liquid phase we get the usual behavior of a liquid. For both systems we have that the g(r) oscillates around 1 for larger distances.
## Coordination Number (CN)
The coordination number (CN), also called ligancy, of a given atom in a chemical system is defined as the number of atoms, molecules or ions bonded to it. This quantity is calculated considered the number of neighbors surrounding a given atom type a cutoff distance.
From the previous graph we can define the cut-off radius to consider only the first neighbors.
```
solid_rcut = 1.29
liquid_rcut = 1.56
```
Now we import the `CoordinationNumber` class.
```
from exma import CoordinationNumber
```
We have the same conduct as in the previos classes.
```
solid_cn = CoordinationNumber(fsolid, "Ar", "Ar", solid_rcut, start=10)
liquid_cn = CoordinationNumber(fliquid, "Ar", "Ar", liquid_rcut, start=10)
```
In this case there is no point in making a plot, so the `calculate` method directly gives us the mean and standard deviation of the number of coordination calculated over all the production frames.
```
solid_cn.calculate(box=solid_box)
liquid_cn.calculate(box=liquid_box)
```
For both cases we have roughly the same result of the CN close to 12 typical value of a compact packing structure, as is the _fcc_ crystal and its amorphization.
| github_jupyter |
<h1> Scaling up ML using Cloud ML Engine </h1>
In this notebook, we take a previously developed TensorFlow model to predict taxifare rides and package it up so that it can be run in Cloud MLE. For now, we'll run this on a small dataset. The model that was developed is rather simplistic, and therefore, the accuracy of the model is not great either. However, this notebook illustrates *how* to package up a TensorFlow model to run it within Cloud ML.
Later in the course, we will look at ways to make a more effective machine learning model.
<h2> Environment variables for project and bucket </h2>
Note that:
<ol>
<li> Your project id is the *unique* string that identifies your project (not the project name). You can find this from the GCP Console dashboard's Home page. My dashboard reads: <b>Project ID:</b> cloud-training-demos </li>
<li> Cloud training often involves saving and restoring model files. If you don't have a bucket already, I suggest that you create one from the GCP console (because it will dynamically check whether the bucket name you want is available). A common pattern is to prefix the bucket name by the project id, so that it is unique. Also, for cost reasons, you might want to use a single region bucket. </li>
</ol>
<b>Change the cell below</b> to reflect your Project ID and bucket name.
```
import os
PROJECT = 'cloud-training-demos' # REPLACE WITH YOUR PROJECT ID
BUCKET = 'cloud-training-demos-ml' # REPLACE WIHT YOUR BUCKET NAME
REGION = 'us-central1' # REPLACE WITH YOUR BUCKET REGION e.g. us-central1
# for bash
os.environ['PROJECT'] = PROJECT
os.environ['BUCKET'] = BUCKET
os.environ['REGION'] = REGION
os.environ['TFVERSION'] = '1.4' # Tensorflow version
%bash
gcloud config set project $PROJECT
gcloud config set compute/region $REGION
```
Allow the Cloud ML Engine service account to read/write to the bucket containing training data.
```
%bash
PROJECT_ID=$PROJECT
AUTH_TOKEN=$(gcloud auth print-access-token)
SVC_ACCOUNT=$(curl -X GET -H "Content-Type: application/json" \
-H "Authorization: Bearer $AUTH_TOKEN" \
https://ml.googleapis.com/v1/projects/${PROJECT_ID}:getConfig \
| python -c "import json; import sys; response = json.load(sys.stdin); \
print response['serviceAccount']")
echo "Authorizing the Cloud ML Service account $SVC_ACCOUNT to access files in $BUCKET"
gsutil -m defacl ch -u $SVC_ACCOUNT:R gs://$BUCKET
gsutil -m acl ch -u $SVC_ACCOUNT:R -r gs://$BUCKET # error message (if bucket is empty) can be ignored
gsutil -m acl ch -u $SVC_ACCOUNT:W gs://$BUCKET
```
<h2> Packaging up the code </h2>
Take your code and put into a standard Python package structure. <a href="taxifare/trainer/model.py">model.py</a> and <a href="taxifare/trainer/task.py">task.py</a> contain the Tensorflow code from earlier (explore the <a href="taxifare/trainer/">directory structure</a>).
```
!find taxifare
!cat taxifare/trainer/model.py
```
<h2> Find absolute paths to your data </h2>
Note the absolute paths below. /content is mapped in Datalab to where the home icon takes you
```
%bash
echo $PWD
rm -rf $PWD/taxi_trained
head -1 $PWD/taxi-train.csv
head -1 $PWD/taxi-valid.csv
```
<h2> Running the Python module from the command-line </h2>
```
%bash
rm -rf taxifare.tar.gz taxi_trained
export PYTHONPATH=${PYTHONPATH}:${PWD}/taxifare
python -m trainer.task \
--train_data_paths="${PWD}/taxi-train*" \
--eval_data_paths=${PWD}/taxi-valid.csv \
--output_dir=${PWD}/taxi_trained \
--train_steps=1000 --job-dir=./tmp
%bash
ls $PWD/taxi_trained/export/exporter/
%writefile ./test.json
{"pickuplon": -73.885262,"pickuplat": 40.773008,"dropofflon": -73.987232,"dropofflat": 40.732403,"passengers": 2}
%bash
model_dir=$(ls ${PWD}/taxi_trained/export/exporter)
gcloud ml-engine local predict \
--model-dir=${PWD}/taxi_trained/export/exporter/${model_dir} \
--json-instances=./test.json
```
<h2> Running locally using gcloud </h2>
```
%bash
rm -rf taxifare.tar.gz taxi_trained
gcloud ml-engine local train \
--module-name=trainer.task \
--package-path=${PWD}/taxifare/trainer \
-- \
--train_data_paths=${PWD}/taxi-train.csv \
--eval_data_paths=${PWD}/taxi-valid.csv \
--train_steps=1000 \
--output_dir=${PWD}/taxi_trained
```
When I ran it (due to random seeds, your results will be different), the ```average_loss``` (Mean Squared Error) on the evaluation dataset was 187, meaning that the RMSE was around 13.
```
from google.datalab.ml import TensorBoard
TensorBoard().start('./taxi_trained')
for pid in TensorBoard.list()['pid']:
TensorBoard().stop(pid)
print 'Stopped TensorBoard with pid {}'.format(pid)
```
If the above step (to stop TensorBoard) appears stalled, just move on to the next step. You don't need to wait for it to return.
```
!ls $PWD/taxi_trained
```
<h2> Submit training job using gcloud </h2>
First copy the training data to the cloud. Then, launch a training job.
After you submit the job, go to the cloud console (http://console.cloud.google.com) and select <b>Machine Learning | Jobs</b> to monitor progress.
<b>Note:</b> Don't be concerned if the notebook stalls (with a blue progress bar) or returns with an error about being unable to refresh auth tokens. This is a long-lived Cloud job and work is going on in the cloud. Use the Cloud Console link (above) to monitor the job.
```
%bash
echo $BUCKET
gsutil -m rm -rf gs://${BUCKET}/taxifare/smallinput/
gsutil -m cp ${PWD}/*.csv gs://${BUCKET}/taxifare/smallinput/
%%bash
OUTDIR=gs://${BUCKET}/taxifare/smallinput/taxi_trained
JOBNAME=lab3a_$(date -u +%y%m%d_%H%M%S)
echo $OUTDIR $REGION $JOBNAME
gsutil -m rm -rf $OUTDIR
gcloud ml-engine jobs submit training $JOBNAME \
--region=$REGION \
--module-name=trainer.task \
--package-path=${PWD}/taxifare/trainer \
--job-dir=$OUTDIR \
--staging-bucket=gs://$BUCKET \
--scale-tier=BASIC \
--runtime-version=$TFVERSION \
-- \
--train_data_paths="gs://${BUCKET}/taxifare/smallinput/taxi-train*" \
--eval_data_paths="gs://${BUCKET}/taxifare/smallinput/taxi-valid*" \
--output_dir=$OUTDIR \
--train_steps=10000
```
Don't be concerned if the notebook appears stalled (with a blue progress bar) or returns with an error about being unable to refresh auth tokens. This is a long-lived Cloud job and work is going on in the cloud.
<b>Use the Cloud Console link to monitor the job and do NOT proceed until the job is done.</b>
<h2> Deploy model </h2>
Find out the actual name of the subdirectory where the model is stored and use it to deploy the model. Deploying model will take up to <b>5 minutes</b>.
```
%bash
gsutil ls gs://${BUCKET}/taxifare/smallinput/taxi_trained/export/exporter
%bash
MODEL_NAME="taxifare"
MODEL_VERSION="v1"
MODEL_LOCATION=$(gsutil ls gs://${BUCKET}/taxifare/smallinput/taxi_trained/export/exporter | tail -1)
echo "Run these commands one-by-one (the very first time, you'll create a model and then create a version)"
#gcloud ml-engine versions delete ${MODEL_VERSION} --model ${MODEL_NAME}
#gcloud ml-engine models delete ${MODEL_NAME}
gcloud ml-engine models create ${MODEL_NAME} --regions $REGION
gcloud ml-engine versions create ${MODEL_VERSION} --model ${MODEL_NAME} --origin ${MODEL_LOCATION} --runtime-version $TFVERSION
```
<h2> Prediction </h2>
```
%bash
gcloud ml-engine predict --model=taxifare --version=v1 --json-instances=./test.json
from googleapiclient import discovery
from oauth2client.client import GoogleCredentials
import json
credentials = GoogleCredentials.get_application_default()
api = discovery.build('ml', 'v1', credentials=credentials,
discoveryServiceUrl='https://storage.googleapis.com/cloud-ml/discovery/ml_v1_discovery.json')
request_data = {'instances':
[
{
'pickuplon': -73.885262,
'pickuplat': 40.773008,
'dropofflon': -73.987232,
'dropofflat': 40.732403,
'passengers': 2,
}
]
}
parent = 'projects/%s/models/%s/versions/%s' % (PROJECT, 'taxifare', 'v1')
response = api.projects().predict(body=request_data, name=parent).execute()
print "response={0}".format(response)
```
<h2> Train on larger dataset </h2>
I have already followed the steps below and the files are already available. <b> You don't need to do the steps in this comment. </b> In the next chapter (on feature engineering), we will avoid all this manual processing by using Cloud Dataflow.
Go to http://bigquery.cloud.google.com/ and type the query:
<pre>
SELECT
(tolls_amount + fare_amount) AS fare_amount,
pickup_longitude AS pickuplon,
pickup_latitude AS pickuplat,
dropoff_longitude AS dropofflon,
dropoff_latitude AS dropofflat,
passenger_count*1.0 AS passengers,
'nokeyindata' AS key
FROM
[nyc-tlc:yellow.trips]
WHERE
trip_distance > 0
AND fare_amount >= 2.5
AND pickup_longitude > -78
AND pickup_longitude < -70
AND dropoff_longitude > -78
AND dropoff_longitude < -70
AND pickup_latitude > 37
AND pickup_latitude < 45
AND dropoff_latitude > 37
AND dropoff_latitude < 45
AND passenger_count > 0
AND ABS(HASH(pickup_datetime)) % 1000 == 1
</pre>
Note that this is now 1,000,000 rows (i.e. 100x the original dataset). Export this to CSV using the following steps (Note that <b>I have already done this and made the resulting GCS data publicly available</b>, so you don't need to do it.):
<ol>
<li> Click on the "Save As Table" button and note down the name of the dataset and table.
<li> On the BigQuery console, find the newly exported table in the left-hand-side menu, and click on the name.
<li> Click on "Export Table"
<li> Supply your bucket name and give it the name train.csv (for example: gs://cloud-training-demos-ml/taxifare/ch3/train.csv). Note down what this is. Wait for the job to finish (look at the "Job History" on the left-hand-side menu)
<li> In the query above, change the final "== 1" to "== 2" and export this to Cloud Storage as valid.csv (e.g. gs://cloud-training-demos-ml/taxifare/ch3/valid.csv)
<li> Download the two files, remove the header line and upload it back to GCS.
</ol>
<p/>
<p/>
<h2> Run Cloud training on 1-million row dataset </h2>
This took 60 minutes and uses as input 1-million rows. The model is exactly the same as above. The only changes are to the input (to use the larger dataset) and to the Cloud MLE tier (to use STANDARD_1 instead of BASIC -- STANDARD_1 is approximately 10x more powerful than BASIC). At the end of the training the loss was 32, but the RMSE (calculated on the validation dataset) was stubbornly at 9.03. So, simply adding more data doesn't help.
```
%%bash
XXXXX this takes 60 minutes. if you are sure you want to run it, then remove this line.
OUTDIR=gs://${BUCKET}/taxifare/ch3/taxi_trained
JOBNAME=lab3a_$(date -u +%y%m%d_%H%M%S)
CRS_BUCKET=cloud-training-demos # use the already exported data
echo $OUTDIR $REGION $JOBNAME
gsutil -m rm -rf $OUTDIR
gcloud ml-engine jobs submit training $JOBNAME \
--region=$REGION \
--module-name=trainer.task \
--package-path=${PWD}/taxifare/trainer \
--job-dir=$OUTDIR \
--staging-bucket=gs://$BUCKET \
--scale-tier=STANDARD_1 \
--runtime-version=$TFVERSION \
-- \
--train_data_paths="gs://${CRS_BUCKET}/taxifare/ch3/train.csv" \
--eval_data_paths="gs://${CRS_BUCKET}/taxifare/ch3/valid.csv" \
--output_dir=$OUTDIR \
--train_steps=100000
```
## Challenge Exercise
Modify your solution to the challenge exercise in d_trainandevaluate.ipynb appropriately. Make sure that you implement training and deployment. Increase the size of your dataset by 10x since you are running on the cloud. Does your accuracy improve?
Copyright 2016 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
# Train-validation tagging
This notebook shows how to split a training dataset into train and validation folds using tags
**Input**:
- Source project
- Train-validation split ratio
**Output**:
- New project with images randomly tagged by `train` or `val`, based on split ration
## Configuration
Edit the following settings for your own case
```
import supervisely_lib as sly
from tqdm import tqdm
import random
import os
team_name = "jupyter_tutorials"
workspace_name = "cookbook"
project_name = "tutorial_project"
dst_project_name = "tutorial_project_tagged"
validation_fraction = 0.4
tag_meta_train = sly.TagMeta('train', sly.TagValueType.NONE)
tag_meta_val = sly.TagMeta('val', sly.TagValueType.NONE)
# Obtain server address and your api_token from environment variables
# Edit those values if you run this notebook on your own PC
address = os.environ['SERVER_ADDRESS']
token = os.environ['API_TOKEN']
# Initialize API object
api = sly.Api(address, token)
```
## Verify input values
Test that context (team / workspace / project) exists
```
# Get IDs of team, workspace and project by names
team = api.team.get_info_by_name(team_name)
if team is None:
raise RuntimeError("Team {!r} not found".format(team_name))
workspace = api.workspace.get_info_by_name(team.id, workspace_name)
if workspace is None:
raise RuntimeError("Workspace {!r} not found".format(workspace_name))
project = api.project.get_info_by_name(workspace.id, project_name)
if project is None:
raise RuntimeError("Project {!r} not found".format(project_name))
print("Team: id={}, name={}".format(team.id, team.name))
print("Workspace: id={}, name={}".format(workspace.id, workspace.name))
print("Project: id={}, name={}".format(project.id, project.name))
```
## Get Source ProjectMeta
```
meta_json = api.project.get_meta(project.id)
meta = sly.ProjectMeta.from_json(meta_json)
print("Source ProjectMeta: \n", meta)
```
## Construct Destination ProjectMeta
```
dst_meta = meta.add_img_tag_metas([tag_meta_train, tag_meta_val])
print("Destination ProjectMeta:\n", dst_meta)
```
## Create Destination project
```
# check if destination project already exists. If yes - generate new free name
if api.project.exists(workspace.id, dst_project_name):
dst_project_name = api.project.get_free_name(workspace.id, dst_project_name)
print("Destination project name: ", dst_project_name)
dst_project = api.project.create(workspace.id, dst_project_name)
api.project.update_meta(dst_project.id, dst_meta.to_json())
print("Destination project has been created: id={}, name={!r}".format(dst_project.id, dst_project.name))
```
## Iterate over all images, tag them and add to destination project
```
for dataset in api.dataset.get_list(project.id):
print('Dataset: {}'.format(dataset.name), flush=True)
dst_dataset = api.dataset.create(dst_project.id, dataset.name)
images = api.image.get_list(dataset.id)
with tqdm(total=len(images), desc="Process annotations") as progress_bar:
for batch in sly.batched(images):
image_ids = [image_info.id for image_info in batch]
image_names = [image_info.name for image_info in batch]
ann_infos = api.annotation.download_batch(dataset.id, image_ids)
anns_to_upload = []
for ann_info in ann_infos:
ann = sly.Annotation.from_json(ann_info.annotation, meta)
tag = sly.Tag(tag_meta_val) if random.random() <= validation_fraction else sly.Tag(tag_meta_train)
ann = ann.add_tag(tag)
anns_to_upload.append(ann)
dst_image_infos = api.image.upload_ids(dst_dataset.id, image_names, image_ids)
dst_image_ids = [image_info.id for image_info in dst_image_infos]
api.annotation.upload_anns(dst_image_ids, anns_to_upload)
progress_bar.update(len(batch))
print("Project {!r} has been sucessfully uploaded".format(dst_project.name))
print("Number of images: ", api.project.get_images_count(dst_project.id))
```
| github_jupyter |
## Train GPT on addition
Train a GPT model on a dedicated addition dataset to see if a Transformer can learn to add.
```
# set up logging
import logging
logging.basicConfig(
format="%(asctime)s - %(levelname)s - %(name)s - %(message)s",
datefmt="%m/%d/%Y %H:%M:%S",
level=logging.INFO,
)
# make deterministic
from mingpt.utils import set_seed
set_seed(42)
import numpy as np
import torch
import string
import os
from tqdm.auto import tqdm
import torch.nn as nn
from torch.nn import functional as F
%load_ext autoreload
%autoreload 2
test = []
for i in range(7):
test.append([])
test[0]
class PrepareData:
""" Tokenizer helper functions """
def __init__(self, mem_slots):
self.mem_slots = mem_slots
self.vocab = ['pad', 'answer', 'end'] + list(' ' + string.punctuation + string.digits + string.ascii_uppercase + string.ascii_lowercase)
self.vocab_size = len(self.vocab) # 10 possible digits 0..9
# Max input characters plus max answer characters
self.src_max_size = 30
self.max_trg = 5
self.block_size = 160 + 32
self.t = {k: v for v, k in enumerate(self.vocab)} # Character to ID
self.idx = {v: k for k, v in self.t.items()} # ID to Character
def initiate_mem_slot_data(self, fname):
# split up all addition problems into either training data or test data
# head_tail = os.path.split(fname)
src, trg = [], []
with open(fname, "r") as file:
text = file.read()[:-1] # Excluding the final linebreak
text_list = text.split('\n')
src = text_list[0:][::2]
trg = text_list[1:][::2]
os.remove(fname)
with open(fname, "a") as file:
for src, trg in zip(src, trg):
file.write(src + '\n')
file.write(trg + '\n')
for _ in range(self.mem_slots):
file.write('\n')
def prepare_data(self, fname):
# split up all addition problems into either training data or test data
# head_tail = os.path.split(fname)
dataset = []
for _ in range(self.mem_slot + 2):
dataset.append([])
with open(fname, "r") as file:
text = file.read()[:-1] # Excluding the final linebreak
text_list = text.split('\n')
for i in range(self.mem_slot + 2):
dataset[i] = text_list[i:][::self.mem_slot + 2]
self.max_src = len(max(dataset[0], key=len))
self.max_trg = len(max(dataset[1], key=len))
return dataset
def sort_data_by_len(self, indexes, data):
test_data_by_length = []
for index in indexes:
test_data_by_length.append([index, len(data[index])])
test_data_by_length = sorted(test_data_by_length, key=lambda x: x[1])
return [i[0] for i in test_data_by_length]
def src2Canvas(self, src):
x = self.t['pad']] * self.src_max_size
return x[:len(src_trg[:-1])] = src_trg[:-1]
def trg2Canvas(self):
y = [self.t['pad']] * self.trg_max_size
return [self.t['pad']] * self.trg_max_size
def tensor2string(self, tensor):
return ''.join([self.idx[tok] for tok in tensor.tolist()])
def string2digits(self, string):
return ''.join([self.t[tok] for tok in string])
def mask_padding(self, digits):
return [-100 if tok == self.t['pad'] else tok for tok in digits]
def mask_question(self, digits, src):
return digits[:len(src)] = -100
def locate_token(self, token, tensor):
return None if self.t[token] not in tensor.tolist() else tensor.tolist().index(self.t[token])
from torch.utils.data import Dataset
class AdditionDataset(Dataset):
"""
Returns addition problems of up to some number of digits in the inputs. Recall
that all GPT cares about are sequences of integers, and completing them according to
patterns in the data. Therefore, we have to somehow encode addition problems
as a sequence of integers.
"""
def __init__(self, fname, split):
self.split = split # train/test
self.vocab = ['pad', 'answer', 'end', 'right', 'wrong'] + list(' ' + string.punctuation + string.digits + string.ascii_uppercase + string.ascii_lowercase)
self.vocab_size = len(self.vocab) # 10 possible digits 0..9
# Max input characters plus max answer characters
# self.block_size = 160 + 32
self.t = {k: v for v, k in enumerate(self.vocab)} # Character to ID
self.idx = {v: k for k, v in self.t.items()} # ID to Character
# split up all addition problems into either training data or test data
with open(fname, "r") as file:
text = file.read()[:-1] # Excluding the final linebreak
text_list = text.split('\n')
self.src = text_list[0:][::2]
self.trg = text_list[1:][::2]
self.src_trg = [src+trg for src,trg in zip(self.src,self.trg)]
self.max_trg = np.ceil((sum(map(len, self.trg)) / len(self.trg)))
self.block_size = len(max(self.src_trg, key=len)) + 1
data_len = len(self.src) # total number of possible combinations
r = np.random.RandomState(1337) # make deterministic
perm = r.permutation(data_len)
num_test = int(data_len*0.1) # 20% of the whole dataset, or only up to 1000
# Sort test data by lenght to batch predictions
test_data_by_length = []
for index in perm[:num_test]:
test_data_by_length.append([index, len(self.src[index])])
test_data_by_length = sorted(test_data_by_length, key=lambda x: x[1])
test_data_by_length = [i[0] for i in test_data_by_length]
self.ixes = np.array(test_data_by_length) if split == 'test' else perm[num_test:]
def __len__(self):
return self.ixes.size
def __getitem__(self, idx):
# given a problem index idx, first recover the associated a + b
idx = self.ixes[idx]
src = self.src[idx]
trg = self.trg[idx]
src_trg = list(src) + ['answer'] + list(trg) + ['end']
src_trg = [self.t[tok] for tok in src_trg] # convert each character to its token index
# x will be input to GPT and y will be the associated expected outputs
x = [self.t['pad']] * self.block_size
y = [self.t['pad']] * self.block_size
x[:len(src_trg[:-1])] = src_trg[:-1]
y[:len(src_trg[1:])] = src_trg[1:] # predict the next token in the sequence
y = [-100 if tok == self.t['pad'] else tok for tok in y] # -100 will mask loss to zero
x = torch.tensor(x, dtype=torch.long)
y = torch.tensor(y, dtype=torch.long)
y[:len(src)] = -100 # we will only train in the output locations. -100 will mask loss to zero
return x, y
# create a dataset
easy = 'data/numbers__place_value.txt'
medium = 'data/numbers__is_prime.txt'
hard = 'data/numbers__list_prime_factors.txt'
train_dataset = AdditionDataset(fname=easy, split='train')
test_dataset = AdditionDataset(fname=easy, split='test')
# for i in range(0, len(train_dataset)):
# if len(train_dataset[i][0]) != 52 or len(train_dataset[i][1]) != 52:
# print(train_dataset.block_size)
# print(len(train_dataset[i][0]))
# print(len(train_dataset[i][1]))
# print(train_dataset[i])
train_dataset[0] # sample a training instance just to see what one raw example looks like
from mingpt.model import GPT, GPTConfig, GPT1Config
# initialize a baby GPT model
mconf = GPTConfig(train_dataset.vocab_size, train_dataset.block_size,
n_layer=2, n_head=4, n_embd=128)
model = GPT(mconf)
from mingpt.trainer import Trainer, TrainerConfig
# initialize a trainer instance and kick off training
tconf = TrainerConfig(max_epochs=1, batch_size=512, learning_rate=6e-4,
lr_decay=True, warmup_tokens=1024, final_tokens=50*len(train_dataset)*(14+1),
num_workers=0)
trainer = Trainer(model, train_dataset, test_dataset, tconf)
trainer.train()
trainer.save_checkpoint()
# now let's give the trained model an addition exam
from torch.utils.data.dataloader import DataLoader
from mingpt.utils import sample, Tokenizer
def give_exam(dataset, batch_size=1, max_batch_size=512, max_batches=-1):
t = Tokenizer(dataset)
results, examples = [], []
loader = DataLoader(dataset, batch_size=batch_size, shuffle=False)
prev_src_len, predict, batch, x_in = 0, 0, 0, 0
pbar = tqdm(enumerate(loader), total=len(loader))
for b, (x, y) in pbar:
src_len = t.locateToken('answer', x[0])
x_in = leftover if prev_src_len == -1 else x_in
# Concat input source with same length
if prev_src_len == src_len:
x_in = torch.cat((x_in, x), 0)
elif prev_src_len == 0:
x_in = x
else:
prev_src_len = -1
predict = 1
leftover = x
prev_src_len = src_len
batch += 1
# Make prediction when the size increses or it reaches max_batch
if predict or batch == max_batch_size:
src_len = t.locateToken('answer', x_in[0]) + 1
batch, predict, prev_src_len, = 0, 0, 0
x_cut = x_in[:, :src_len]
pred = x_cut.to(trainer.device)
pred = sample(model, pred, int(dataset.max_trg+1))
for i in range(x_in.size(0)):
pad, end= t.locateToken('pad', x_in[i]), t.locateToken('end', pred[i])
x, out = x_in[i][src_len:pad], pred[i][src_len:end]
x, out = t.tensor2string(x), t.tensor2string(out)
correct = 1 if x == out else 0
results.append(correct)
question = x_in[i][:src_len-1]
question_str = tensor2string(dataset.idx, question)
if not correct:
examples.append([question_str, x, out, t.tensor2string(x_cut[i]), t.tensor2string(x_in[i]), t.tensor2string(pred[i]), pad, end])
if max_batches >= 0 and b+1 >= max_batches:
break
print("final score: %d/%d = %.2f%% correct" % (np.sum(results), len(results), 100*np.mean(results)))
return examples
# training set: how well did we memorize?
examples = give_exam(test_dataset, batch_size=1, max_batches=-1)
print("Q: %s\nX:%s\nO:%s\n" % (examples[0][0], examples[0][1] , examples[0][2]))
for item in examples:
print("Question:", item[0])
print("X:", item[1])
print("Out:", item[2])
# test set: how well did we generalize?
give_exam(train_dataset, batch_size=1024, max_batches=1)
# well that's amusing... our model learned everything except 55 + 45
import itertools as it
f = ['-1', '-1', '2', '1', '1']
it.takewhile(lambda x: x!='2', f)
f
```
| github_jupyter |
# Demo 1: a demo based on visual-92-categories-task MEG data
Here is a demo based on the publicly available visual-92-categories-task MEG datasets. (Reference: Cichy, R. M., Pantazis, D., & Oliva, A. “Resolving human object recognition in space and time.” Nature neuroscience (2014): 17(3), 455-462.) MNE-Python has been used to load this dataset.
```
# -*- coding: utf-8 -*-
' a demo based on visual-92-categories-task MEG data '
# Users can learn how to use Neurora to do research based on EEG/MEG etc data.
__author__ = 'Zitong Lu'
import numpy as np
import os.path as op
from pandas import read_csv
import mne
from mne.io import read_raw_fif
from mne.datasets import visual_92_categories
from neurora.nps_cal import nps
from neurora.rdm_cal import eegRDM
from neurora.rdm_corr import rdm_correlation_spearman
from neurora.corr_cal_by_rdm import rdms_corr
from neurora.rsa_plot import plot_rdm, plot_corrs_by_time, plot_nps_hotmap, plot_corrs_hotmap
```
## Section 1: loading example data
Here, we use MNE-Python toolbox for loading data and processing. You can learn this process from MNE-Python (https://mne-tools.github.io/stable/index.html).
```
data_path = visual_92_categories.data_path()
fname = op.join(data_path, 'visual_stimuli.csv')
conds = read_csv(fname)
conditions = []
for c in conds.values:
cond_tags = list(c[:2])
cond_tags += [('not-' if i == 0 else '') + conds.columns[k]
for k, i in enumerate(c[2:], 2)]
conditions.append('/'.join(map(str, cond_tags)))
event_id = dict(zip(conditions, conds.trigger + 1))
print(event_id)
sub_id = [0, 1, 2]
megdata = np.zeros([3, 92, 306, 1101], dtype=np.float32)
subindex = 0
for id in sub_id:
fname = op.join(data_path, 'sample_subject_'+str(id)+'_tsss_mc.fif')
raw = read_raw_fif(fname)
events = mne.find_events(raw, min_duration=.002)
events = events[events[:, 2] <= 92]
subdata = np.zeros([92, 306, 1101], dtype=np.float32)
for i in range(92):
epochs = mne.Epochs(raw, events=events, event_id=i + 1, baseline=None,
tmin=-0.1, tmax=1, preload=True)
data = epochs.average().data
subdata[i] = data
megdata[subindex] = subdata
subindex = subindex + 1
# the shape of MEG data: megdata is [3, 92, 306, 1101]
# n_subs = 3, n_conditions = 92, n_channels = 306, n_timepoints = 1101 (-100ms to 1000ms)
```
## Section 2: Preprocessing
```
# shape of megdata: [n_subs, n_cons, n_chls, n_ts] -> [n_cons, n_subs, n_chls, n_ts]
megdata = np.transpose(megdata, (1, 0, 2, 3))
# shape of megdata: [n_cons, n_subs, n_chls, n_ts] -> [n_cons, n_subs, n_trials, n_chls, n_ts]
# here data is averaged, so set n_trials = 1
megdata = np.reshape(megdata, [92, 3, 1, 306, 1101])
```
## Section 3: Calculating the neural pattern similarity
```
# Get data under different condition
# Here we calculate the neural pattern similarity (NPS) between two stimulus
# Seeing Humanface vs. Seeing Non-Humanface
# get data under "humanface" condtion
megdata_humanface = megdata[12:24]
# get data under "nonhumanface" condition
megdata_nonhumanface = megdata[36:48]
# Average the data
avg_megdata_humanface = np.average(megdata_humanface, axis=0)
avg_megdata_nonhumanface = np.average(megdata_nonhumanface, axis=0)
# Create NPS input data
# Here we extract the data from first 5 channels between 0ms and 1000ms
nps_data = np.zeros([2, 3, 1, 5, 1000]) # n_cons=2, n_subs=3, n_chls=5, n_ts=1000
nps_data[0] = avg_megdata_humanface[:, :, :5, 100:1100] # the start time of the data is -100ms
nps_data[1] = avg_megdata_nonhumanface[:, :, :5, 100:1100] # so 100:1200 corresponds 0ms-1000ms
# Calculate the NPS with a 10ms time-window
# (raw sampling requency is 1000Hz, so here time_win=10ms/(1s/1000Hz)/1000=10)
nps = nps(nps_data, time_win=10, time_step=10, sub_opt=0)
# Plot the NPS results
plot_nps_hotmap(nps[:, :, 0], time_unit=[0, 0.01], abs=True)
# Smooth the results and plot
plot_nps_hotmap(nps[:, :, 0], time_unit=[0, 0.01], abs=True, smooth=True)
```
## Section 4: Calculating single RDM and Plotting
```
# Calculate the RDM based on the data during 190ms-210ms
rdm = eegRDM(megdata[:, :, :, :, 290:310], sub_opt=0)
# Plot this RDM
plot_rdm(rdm, percentile=True)
```
## Section 5: Calculating RDMs and Plotting
```
# Calculate the RDMs by a 10ms time-window
# (raw sampling requency is 1000Hz, so here time_win=10ms/(1s/1000Hz)/1000=10)
rdms = eegRDM(megdata, time_opt=1, time_win=10, time_step=10, sub_opt=0)
# Plot the RDM of -100ms, 0ms, 50ms, 100ms, 150ms, 200ms
times = [0, 10, 20, 30, 40, 50]
for t in times:
plot_rdm(rdms[t], percentile=True)
```
## Section 6: Calculating the Similarity between two RDMs
```
# RDM of 200ms
rdm_sample1 = rdms[30]
# RDM of 800ms
rdm_sample2 = rdms[90]
# calculate the correlation coefficient between these two RDMs
corr = rdm_correlation_spearman(rdm_sample1, rdm_sample2)
print(corr)
```
## Section 7: Calculating the Similarity and Plotting
```
# Calculate the representational similarity between 200ms and all the time points
corrs1 = rdms_corr(rdm_sample1, rdms)
# Plot the corrs1
corrs1 = np.reshape(corrs1, [1, 110, 2])
plot_corrs_by_time(corrs1, time_unit=[-0.1, 0.01])
# Calculate and Plot multi-corrs
corrs2 = rdms_corr(rdm_sample2, rdms)
corrs = np.zeros([2, 110, 2])
corrs[0] = corrs1
corrs[1] = corrs2
labels = ["by 200ms's data", "by 800ms's data"]
plot_corrs_by_time(corrs, labels=labels, time_unit=[-0.1, 0.01])
```
## Section 8: Calculating the RDMs for each channels
```
# Calculate the RDMs for the first six channels by a 10ms time-window between 0ms and 1000ms
rdms_chls = eegRDM(megdata[:, :, :, :6, 100:1100], chl_opt=1, time_opt=1, time_win=10, time_step=10, sub_opt=0)
# Create a 'human-related' coding model RDM
model_rdm = np.ones([92, 92])
for i in range(92):
for j in range(92):
if (i < 24) and (j < 24):
model_rdm[i, j] = 0
model_rdm[i, i] = 0
# Plot this coding model RDM
plot_rdm(model_rdm)
# Calculate the representational similarity between the neural activities and the coding model for each channel
corrs_chls = rdms_corr(model_rdm, rdms_chls)
# Plot the representational similarity results
plot_corrs_hotmap(corrs_chls, time_unit=[0, 0.01])
# Set more parameters and re-plot
plot_corrs_hotmap(corrs_chls, time_unit=[0, 0.01], lim=[-0.15, 0.15], smooth=True, cmap='bwr')
```
| github_jupyter |
# Detailed Steps Example
#### This notebook demonstrates how the data cleaning, peak fitting and descriptors generation works step by step, serving as a detailed example of the `ProcessData_PlotDescriptors_Examples.ipynb`.
## Packages and Needed Python Files Preparation
### First we import packages we need:
```
import glob
import itertools
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import peakutils
import scipy
import sqlite3 as sql
from diffcapanalyzer import chachifuncs as ccf
from diffcapanalyzer import descriptors as dct
```
### Then, import the data we want to process:
```
df = pd.read_csv(os.path.join('../data/ARBIN/CS2_33/CS2_33_8_30_10.csv'))
database = 'example_db.db'
base_filename = 'CS2_33_8_30_10'
datatype = 'ARBIN'
```
## Processing Data, including Data Cleaning, Peak Fitting and Descriptors Generation
### Data Cleaning
First, we import the raw data of cycle 1
```
raw_data = ccf.load_sep_cycles(base_filename, database, datatype)
raw_df = raw_data[1]
# user can change the index of raw_df to see other cycles
fig1 = plt.figure(figsize = (8,8), facecolor = 'w', edgecolor= 'k')
plt.plot(raw_df['Voltage(V)'], raw_df['dQ/dV'], c = 'black', linewidth = 2, label = 'Raw Cycle Data')
plt.ylabel('dQ/dV (Ah/V)', fontsize =20)
plt.xlabel('Voltage(V)', fontsize = 20)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.tick_params(size = 10, width = 1)
plt.title('Raw Data of Cycle 1', fontsize = 24)
# plt.xlim(0, 4)
plt.ylim(-10,10)
# Uncomment the following line if you would like to save the plot.
# plt.savefig(fname = 'MyExampleCycle_Raw Data.png', bbox_inches='tight', dpi = 600)
```
As we can see in the Raw data, there are several noise and jumps on both ends, So in order to eliminate those noise and jumps, we need to remove those data points which the dV value is about zero.
Therefore, we excute fuction `drop_inf_nan_dqdv` to drop rows where dV=0 (or about 0) in a dataframe that has already had dv calculated, then recalculates dV and calculates dQ/dV.
```
rawdf = ccf.init_columns(raw_df, datatype)
rawdf1 = ccf.calc_dq_dqdv(rawdf, datatype)
clean_df = ccf.drop_inf_nan_dqdv(rawdf1, datatype)
# Clean charge and discharge cycles separately:
charge, discharge = ccf.sep_char_dis(clean_df, datatype)
charge = ccf.clean_charge_discharge_separately(charge, datatype)
discharge = ccf.clean_charge_discharge_separately(discharge, datatype)
fig2 = plt.figure(figsize = (8,8), facecolor = 'w', edgecolor= 'k')
plt.plot(charge['Voltage(V)'], charge['dQ/dV'], c = 'green', linewidth = 2, label = 'Clean Charge Cycle Data')
plt.plot(discharge['Voltage(V)'], discharge['dQ/dV'], c = 'green', linewidth = 2, label = 'Clean Dishcarge Cycle Data')
plt.ylabel('dQ/dV (Ah/V)', fontsize =20)
plt.xlabel('Voltage(V)', fontsize = 20)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.tick_params(size = 10, width = 1)
plt.title('Clean Data of Cycle 1', fontsize = 24)
# plt.xlim(0, 4)
plt.ylim(-10,10)
# Uncomment the following line if you would like to save the plot.
# plt.savefig(fname = 'MyExampleCycle-Clean Data.png', bbox_inches='tight', dpi = 600)
```
In order to help the computer to recognize peaks more easily later, we need to apply the Savitzky-Golay Filter to our data to get a nice and smooth curve.
Then, sperate cycles into charge and discharge cycles first, and apply the Savitzky–Golay filter to smooth the data:
```
windowlength = 9
polyorder = 3
# apply Savitzky–Golay filter
if len(discharge) > windowlength:
smooth_discharge = ccf.my_savgolay(discharge, windowlength, polyorder)
else:
discharge['Smoothed_dQ/dV'] = discharge['dQ/dV']
smooth_discharge = discharge
# this if statement is for when the datasets have less datapoints
# than the windowlength given to the sav_golay filter.
# without this if statement, the sav_golay filter throws an error
# when given a dataset with too few points. This way, we simply
# forego the smoothing function.
if len(charge) > windowlength:
smooth_charge = ccf.my_savgolay(charge, windowlength, polyorder)
else:
charge['Smoothed_dQ/dV'] = charge['dQ/dV']
smooth_charge = charge
# same as above, but for charging cycles.
fig3 = plt.figure(figsize = (8,8), facecolor = 'w', edgecolor= 'k')
plt.plot(charge['Voltage(V)'], charge['Smoothed_dQ/dV'], c = 'red', linewidth = 2, label = 'Smooth Charge Cycle Data')
plt.plot(discharge['Voltage(V)'], discharge['Smoothed_dQ/dV'], c = 'red', linewidth = 2, label = 'Smooth Dishcarge Cycle Data')
plt.ylabel('dQ/dV (Ah/V)', fontsize =20)
plt.xlabel('Voltage(V)', fontsize = 20)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.tick_params(size = 10, width = 1)
plt.title('Smooth Data of Cycle 1', fontsize = 24)
#plt.xlim(2.8, 4.2)
plt.ylim(-10,10)
# Uncomment the following line if you would like to save the plot.
# plt.savefig(fname = 'MyExampleCycle_Smooth Data.png', bbox_inches='tight', dpi = 600)
```
### Peak Finding
Once we got the smooth data, we then apply the "peak_finder" function, which is based on an open source package "Peakutils" to locate the peaks for both charge and discharge data.
```
# we first create the column of the dataframe according to the datatype
(cycle_ind_col, data_point_col, volt_col, curr_col,
dis_cap_col, char_cap_col, charge_or_discharge) = ccf.col_variables(datatype)
chargeloc_dict = {}
param_df = pd.DataFrame(columns=['Cycle','Model_Parameters_charge','Model_Parameters_discharge'])
# and we determine the max length of the dataframe
if len(clean_df[cycle_ind_col].unique()) > 1:
length_list = [len(clean_df[df_clean[cycle_ind_col] == cyc])
for cyc in clean_df[cycle_ind_col].unique() if cyc != 1]
lenmax = max(length_list)
else:
length_list = 1
lenmax = len(clean_df)
import peakutils
import scipy.signal
peak_thresh=2/max(charge['Smoothed_dQ/dV'])
# user can change the above peak_thresh to adjust the sensitivity of the peak_finder
# apply peak_finder
i_charge, volts_i_ch, peak_heights_c = dct.peak_finder(charge,'c', windowlength, polyorder, datatype, lenmax, peak_thresh)
i_discharge, volts_i_dic, peak_heights_d = dct.peak_finder(discharge,'d', windowlength, polyorder, datatype, lenmax, peak_thresh)
# set up a figure
fig4 = plt.figure(figsize = (8,8), facecolor = 'w', edgecolor= 'k')
plt.plot(charge['Voltage(V)'], charge['Smoothed_dQ/dV'], 'r')
plt.plot(discharge['Voltage(V)'], discharge['Smoothed_dQ/dV'], 'r')
plt.xlabel('Voltage(V)', fontsize = 20)
plt.ylabel('dQ/dV (Ah/V)', fontsize =20)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.tick_params(size = 10, width = 1)
plt.title('Peak Location of Cycle 1', fontsize = 24)
plt.plot(charge['Voltage(V)'][i_charge], charge['Smoothed_dQ/dV'][i_charge], 'o', c='b')
plt.plot(discharge['Voltage(V)'][i_discharge], discharge['Smoothed_dQ/dV'][i_discharge], 'o', c='b')
# Uncomment the following line if you would like to save the plot.
# plt.savefig(fname = 'MyExampleCycle_peak_finder.png', bbox_inches='tight', dpi = 600)
```
### Model Generation
Following that, we continued to apply functions "model_gen" and "model_eval" to generate models that fit those peaks, which generates a mixture of Pseudo-Voigt distributions with the gaussian function that is fitted to the peak.
```
# change the number of cyc according to the cycle number
cyc = 1
# we first assign some variables in charge cycle
V_series_c = smooth_charge[volt_col]
dQdV_series_c = smooth_charge['Smoothed_dQ/dV']
# apply model_gen and model_eval to generate model and iritate from inital guess to the best fit
par_c, mod_c, indices_c = dct.model_gen(V_series_c, dQdV_series_c, 'c', i_charge, cyc, peak_thresh)
model_c = dct.model_eval(V_series_c, dQdV_series_c, 'c', par_c, mod_c)
if model_c is not None:
mod_y_c = mod_c.eval(params=model_c.params, x=V_series_c)
myseries_c = pd.Series(mod_y_c)
myseries_c = myseries_c.rename('Model')
model_c_vals = model_c.values
new_df_mody_c = pd.concat([myseries_c, V_series_c, dQdV_series_c, smooth_charge[cycle_ind_col]], axis=1)
else:
mod_y_c = None
new_df_mody_c = None
model_c_vals = None
# now the discharge
V_series_d = smooth_discharge[volt_col]
dQdV_series_d = smooth_discharge['Smoothed_dQ/dV']
par_d, mod_d, indices_d = dct.model_gen(V_series_d, dQdV_series_d, 'd', i_discharge, cyc, peak_thresh)
model_d = dct.model_eval(V_series_d, dQdV_series_d, 'd', par_d, mod_d)
if model_d is not None:
mod_y_d = mod_d.eval(params=model_d.params, x=V_series_d)
myseries_d = pd.Series(mod_y_d)
myseries_d = myseries_d.rename('Model')
new_df_mody_d = pd.concat([-myseries_d, V_series_d, dQdV_series_d, smooth_discharge[cycle_ind_col]], axis=1)
model_d_vals = model_d.values
else:
mod_y_d = None
new_df_mody_d = None
model_d_vals = None
if new_df_mody_c is not None or new_df_mody_d is not None:
new_df_mody = pd.concat([new_df_mody_c, new_df_mody_d], axis=0)
else:
new_df_mody = None
new_df_mody
# plots data
fig5 = plt.figure(figsize = (18,8), facecolor = 'w', edgecolor= 'k')
plt.subplot(1, 2, 1)
plt.plot(smooth_charge['Voltage(V)'], smooth_charge['Smoothed_dQ/dV'], c = 'red', linewidth = 2, label = 'Smooth Data')
plt.plot(smooth_charge['Voltage(V)'], model_c.init_fit, 'k--')
plt.plot(smooth_charge['Voltage(V)'], model_c.best_fit, 'b-')
plt.xlabel('Voltage(V)', fontsize = 20)
plt.ylabel('dQ/dV (Ah/V)', fontsize =20)
plt.rcParams.update({'font.size':20})
plt.title('Charge Peak Fitting of Cycle 1', fontsize = 24)
plt.legend(['Raw Data', 'Initial Model', 'Fitted Model'], loc=2, fontsize=10)
plt.subplot(1, 2, 2)
plt.plot(smooth_discharge['Voltage(V)'], -smooth_discharge['Smoothed_dQ/dV'], c = 'red', linewidth = 2, label = 'Smooth Data')
plt.plot(smooth_discharge['Voltage(V)'], model_d.init_fit[::-1], 'k--')
plt.plot(smooth_discharge['Voltage(V)'], model_d.best_fit[::-1], 'b-')
plt.xlabel('Voltage(V)', fontsize = 20)
plt.ylabel('dQ/dV (Ah/V)', fontsize =20)
plt.rcParams.update({'font.size':20})
plt.title('Discharge Peak Fitting of Cycle 1', fontsize = 24)
plt.legend(['Raw Data', 'Initial Model', 'Fitted Model'], loc=2, fontsize=10)
# Uncomment the following line if you would like to save the plot.
# plt.savefig(fname = 'MyExampleCycle_generate model', bbox_inches='tight', dpi = 600)
```
So, the overall fit model with the raw data is shown below
```
smooth_df = smooth_charge.append(smooth_discharge)
fig6 = plt.figure(figsize = (8,8), facecolor = 'w', edgecolor= 'k')
plt.plot(raw_df['Voltage(V)'], raw_df['dQ/dV'], 'k-', label = 'Raw Data')
plt.plot(smooth_df['Voltage(V)'], new_df_mody['Model'], 'r--')
plt.xlabel('Voltage(V)', fontsize = 20)
plt.ylabel('dQ/dV (Ah/V)', fontsize =20)
plt.rcParams.update({'font.size':20})
plt.title('Model Generation of Cycle 1', fontsize = 24)
plt.legend(['Raw Data', 'Fitted Model'], loc=2, fontsize=10)
plt.ylim(-10,10)
# Uncomment the following line if you would like to save the plot.
# plt.savefig(fname = 'MyExampleCycle_final generate model', bbox_inches='tight', dpi = 600)
```
Moreover, from the model generation, we can also obtain the descriptors of the peak, the below shows the extraction of the descriptors:
```
# extract the peak location from the descriptors dictionary
center_c = [value for key, value in model_c_vals.items() if 'center' in key.lower()]
center_d = [value for key, value in model_d_vals.items() if 'center' in key.lower()]
center_c = center_c[0:-1]
center_d = center_d[0:-1]
print("The location(V) of the charge peak of cycle", cyc, "are", center_c)
print("The location(V) of the discharge peak of cycle", cyc, "are", center_d)
print("The height(Ah/V) of the charge peak of cycle", cyc, "are", peak_heights_c)
print("The height(Ah/V) of the discharge peak of cycle", cyc, "are", peak_heights_d)
```
| github_jupyter |
# Indexing
Okay guys today's lecture is indexing.
> What is indexing?
At heart, indexing is the ability to inspect a value inside a object. So basically if we have a list, X, of 100 items and our index is 'i' then 'i of X' returns the *ith value* inside the list (p.s. we can index strings too).
Okay, so what is the Syntax for this? Glad you asked:
{variable} [{integer}]
So if we wanted to index into something called "a_string" in code it would look something like:
a_string[integer]
Now, the integer in question cannot be any number from -infinity to +infinity. Rather, it is bounded by the size of the variable. For example, if the size of the variable is 5 that means our integer has to be in the range -5 to 4. Or more generally:
Index Range ::
Lower Bound = -len(variable)
Upper Bound = len(variable) - 1
Anything outside this range = IndexError
Just as a quick explanation, len() is a built-in command that gets the size of the object and adding a "-" sign infront of an integer 'flips' its sign:
```
# flipping signs of numbers...
a = 5
b = -5
print(-a, -b)
# len function
x1 = []
x2 = "12"
x3 = [1,2,3]
print(len(x1), len(x2), len(x3))
x = [1,2,3]
print(x[100]) # <--- IndexError! 100 is waayyy out of bounds
```
Now, those bounds I have just given might sound a bit arbitrary, but actually I can explain exactly how they work. Consider the following picture:

So in this picture we have the string ‘hello’. The two rows of numbers represent the indexes of this string. In Python we start counting from 0 which means the first item in a list/string always has an index of 0. And since we start counting at zero then that means the last item in the list/string is len(item)-1 like so:
```
string = "hello"
print(string[0]) # first item
print(string[len(string)-1]) # last item
```
So that explains the first row of numbers in the image. What about the second row? Well, in Python not only can you index forwards you can also index backwards.
## Readabily counts...
So basically index [0] will always be the start of the list/string and an index of [-1] will always be the end. If you wanted the middle "l" in "hello" have a choice; either [2] or [-3] will work. **And, as a general rule, if code ends up being equivalent your choice should be to go with whatever is more readable.**
> There should be one-- and preferably only one --obvious way to do it. ~ Zen of Python
For example:
```
a_string = "Hello"
# indexing first item...
print(a_string[0]) # Readable
print(a_string[-len(a_string)]) # Less readable
print(a_string[-1]) # Readable
print(a_string[len(a_string)-1]) # Less readable
print(a_string[4]) # Avoid this whereever possible! BAD BAD BAD!!
```
You might wonder what is wrong with index[4] to reference the end of the list.
The problem with using index[4] instead of [-1] is that the former way of doing things is considerably less readable. Without actually checking the length of the input the meaning of index[4] is somewhat ambiguous; is this the end? Near the beginning/middle? Meanwhile [-1] **always** refers to the end regardless of input size, and so therefore its meaning is always clear **even when** we don’t know the size of the input.
Index[len(a_string)-1] meanwhile always refers to the end of the list but it is considerably more verbose and less readable than the simple [-1].
## The Index Method
The string class AND the list class both have an index method, and now that we have just covered indexing we are in a position to understand its output.
Basically, we ask if an item is in a string/list. And if it is, the method returns an index for that item. For example:
```
a_list = ["qwerty", "dave", "magic johnson", "qwerty"]
a_string = "Helllllllo how ya doin fam?"
# notice that Python returns the index of the first match.
print(a_list.index("qwerty"))
print(a_string.index("l"))
# if item is not in the list, you get an value error:
print(a_list.index("chris"))
```
## What can we do with indexing?
Obviously we can do a lot with indexing, in the cases of lists, for example, we change the value of the list at position ‘i’. Its simple to do that:
```
a_list = [1,2,3]
print(a_list)
a_list[-1] = "a"
print(a_list)
a_list[0] = "c"
print(a_list)
a_list[1] = "b"
print(a_list)
```
### Can we change the values inside strings?
Lets try!
```
a_string = "123"
a_string[0] = "a" # <-- Error; strings are an "immutable" data type in Python.
```
In python strings are immutable, which is a fancy way of saying that they are set in stone; once created you just can't change them. Your only option is to create new strings with data you want.
If we create a new string we can use the old variable if we want. But in this case, you didn't change the value of the string. Rather what you did was create a new string and give it a variable name, and thats allowed.
Here is one way we can change the value of 'a_string':
```
a_string = "123"
a_string = "a" + a_string[1:] # slicing, see below.
print(a_string)
```
## Making Grids
> "Flat is better than nested". ~ Zen of Python
Talking of lists, remember that we can go all "inception-like" with lists and shove lists inside lists inside lists. How can we index a beast like that? Well, with difficulty...
```
this_is_insane = [ [[[[[[[[[[[[100]]]]]]]]]]]] ] # WTF !!??
print(this_is_insane[0][0][0][0][0][0][0][0][0][0][0][0][0])
```
To index a list inside a list the syntax is to add another [{integer}] on the end. Repeat until you get to the required depth.
list[{integer}][{integer}]
In the case of the above the value 100 was nested inside so many lists that it took a lot of effort to tease it out. Structures like this are hard to work with, which is why the usual advice is to 'flatten' your lists wherever possible.
With this said, nested structures are not all bad. A really common way of representing a grid in Python is to use nested lists. In which case, we can index any square we want by first indexing the 'row' and then the 'column'. Like so:
grid[row][column]
If you ever want to build simple board games (chess, connect 4, etc) you might find the representation useful. In code:
```
grid = [ ["0"] * 5 for _ in range(5) ] # building a nested list, in style. 'List Comprehensions' are not covered in this course.
print("The Grid looks like this...:", grid[2:], "\n")
# Note: "grid[2:]" above is a 'slice' (more on slicing below), in this case I'm using slicing to truncate the results,
# observe that three lists get printed, not five.
def print_grid():
"""This function simply prints grid, row by row."""
for row in grid: # This is a for-loop, more on these later!
print(row)
print_grid()
print("\n")
grid[0][0] = "X" # Top-left corner
grid[0][-1] = "Y" # Top-right corner
grid[-1][0] = "W" # Bottom-left corner
grid[-1][-1] = "Z" # Bottom-right corner
grid[2][2] = "A" # Somewhere near the middle
print_grid()
# Quick note, since the corners index are defined by 0 and -1, these numbers should work for all nxn grids.
```
Anyway, thats enough about indexing for now, let's move onto the topic of slicing...
## Slicing
What is slicing? Well it is a bit like indexing, only instead of returning point 'X' we return all the values between the points (x, y). Just as with indexing, you can slice strings as well as lists.
Note: start points are *inclusive* and endpoints are *exclusive*.
{variable} [{start} : {end} : {step}]
* Where start, end and step are all integer values.
It is also worth noting that each of start, end and step are optional arguments, when nothing is given they default to the start of the list, end of the list and the default step is 1.
If you give start/step an integer Python will treat that number as an index. Thus, a_list[2:10] says "Hey Python, go fetch me all the values in 'a_list' starting at index 2 up-to **(but not including)** index 10.
Unlike indexing however, if you try to slice outside of range you won't get an error message. If you have a list of length five and try to slice with values 0 and 100 Python will just return the whole list. If you try to slice the list at 100 and 200 an empty list '[]' will be the result. Lets see a few examples:
```
lst = list(range(1,21)) # list(range) just makes a list of numbers 1 to 20
# The below function just makes it faster for me to type out the test cases below.
def printer(start, end, lst):
""" Helper function, takes two integers (start, end) and a list/string.
Function returns a formated string that contains: start, end and lst[start:end]"""
if start:
if end:
sliced = lst[start:end]
else:
sliced = lst[start:]
elif end:
sliced = lst[:end]
else:
sliced = lst[:]
return "slice is '[{}:{}]', which returns: {}".format(start, end, sliced)
print("STARTING LIST IS:", lst)
print("")
# Test cases
print("SLICING LISTS...")
print(printer("","", lst)) # [:] is sometimes called a 'shallow copy' of a list.
print(printer("", 5, lst )) # first 5 items.
print(printer(14,"", lst)) # starting at index 14, go to the end.
print(printer(200,500,lst)) # No errors for indexes that should be "out of bounds".
print(printer(5,10, lst))
print(printer(4,5, lst))
# Negative numbers work too. In the case below we start at the 5th last item and move toward the 2nd to last item.
print(printer(-5,-2, lst))
print(printer(-20,-1, lst)) # note that this list finishes at 19, not 20.
# and for good measure, a few strings:
print("\nSLICING STRINGS...")
a_string = "Hello how are you?"
print(printer("","", a_string)) # The whole string aka a 'shallow copy'
print(printer(0,5, a_string))
print(printer(6,9, a_string))
print(printer(10,13, a_string))
print(printer(14, 17, a_string))
print(printer(17, "", a_string))
```
Alright, so that's the basics of slicing covered, the only remaining question is what the final "step" argument does. Well basically, the step allows us to 'skip' every *nth *element of the list/string.
For example, suppose that I have (just as before) a list of numbers 1-to-20, but this time I want to return the EVEN numbers between 15 and 19. Intuitively we know that the result should be [16,18] but how can we do this in code?
```
a_list = list(range(1,21))
sliced_list = a_list[15:19:2]
print(sliced_list)
print(a_list[17])
```
How does this work? Well, index 15 is the number 16 (remember we count from 0 in Python), and then we skip index 16 (an odd number) and go straight to index 17 (which is the number 18). The next index to look at is 20, but since that is larger than our end step (19) we terminate.
On last thing I'd like to note is that we got even numbers in this case because we started with an even number (index 15= 16). Had we of started with an odd number, this process would have returned odd numbers. For example:
```
a_list = list(range(0,206))
slice1 = a_list[::10] # every 10th element starting from zero = [0, 10, 20, ...]
slice2 = a_list[5::10] # every 10th element starting from 5 = [5, 15, 25,...]
a_string = "a123a123a123a123a123a123a123" # this pattern has a period of 4.
slice3 = a_string[::4] # starts at a, returns aaaaaa
slice4 = a_string[3::4] # starts at 3, returns 333333
print(slice1, slice2, slice3, slice4, sep="\n")
```
In both of the above cases we are using a step of size 10. If we start at 0 that means we get:
10,20,30...
but if we start at 5 then the sequence we get is
5, 15, 25...
In the case of the string example above, the patten has a length of four and then repeats. Thus, if we start with n charater and have a step of 4 the resulting pattern with be "nnnnnn".
## Reversing lists with step
The very last thing I want to show you about a the step argument is that if you set step to -1 it will reverse the string/list.
For example:
```
a_list = list(range(1, 11))
print(a_list)
print(a_list[::-1]) # reverses the list
```
## The Range Function
You maybe have observed that I use the 'range' function in some of the above examples. This function doesn't have anything to do with indexing or slicing, but I thought I would briefly talk about it here because although the syntax is different this function works in a very similar way to slicing. More specifically, the range function takes 3 arguments; start, end, step (optional). And these arguments work in a similar way to how start, end and step work with regards slicing. Allow me to demonstrate:
```
list_1 = list(range(1,21))
list_1 = list_1[2::3]
print(list_1)
# The above 3 lines can be refactored to:
list_2 = list(range(3, 21, 3))
print(list_2)
```
You will note a small difference between the two ways of doing things. When we slice we start the the count at 2 whereas with range we start the count at 3. The difference is the result of the fact the range function is dealing with numbers, whereas the slice is using indexing (e.g. list_1[2] is the number 3).
And just as with slicing, a step of -1 counts backwards...
```
list_3 = list(range(10, -1, -1)) # this says: "start at the number 10 and count backwards to 0
# please remember that start points are inclusive BUT endpoints are exclusive,
# if we want to include 0 in the results we must have an endpoint +1 of our target.
# in this case the number one past zero (when counting backwards) is -1.
print(list_3)
```
| github_jupyter |
```
from fastai.text.all import *
chunked??
```
Let's look at how long it takes to tokenize a sample of 1000 IMDB review.
```
path = untar_data(URLs.IMDB_SAMPLE)
df = pd.read_csv(path/'texts.csv')
df.head(2)
ss = L(list(df.text))
ss[0]
```
We'll start with the simplest approach:
```
def delim_tok(s, delim=' '): return L(s.split(delim))
s = ss[0]
delim_tok(s)
```
...and a general way to tokenize a bunch of strings:
```
def apply(func, items): return list(map(func, items))
```
Let's time it:
```
%%timeit -n 2 -r 3
global t
t = apply(delim_tok, ss)
```
...and the same thing with 2 workers:
```
%%timeit -n 2 -r 3
parallel(delim_tok, ss, n_workers=2, progress=False)
```
How about if we put half the work in each worker?
```
batches32 = [L(list(o)).map(str) for o in np.array_split(ss, 32)]
batches8 = [L(list(o)).map(str) for o in np.array_split(ss, 8 )]
batches = [L(list(o)).map(str) for o in np.array_split(ss, 2 )]
%%timeit -n 2 -r 3
parallel(partial(apply, delim_tok), batches, progress=False, n_workers=2)
```
So there's a lot of overhead in using parallel processing in Python. :(
Let's see why. What if we do nothing interesting in our function?
```
%%timeit -n 2 -r 3
global t
t = parallel(noop, batches, progress=False, n_workers=2)
```
That's quite fast! (Although still slower than single process.)
What if we don't return much data?
```
def f(x): return 1
%%timeit -n 2 -r 3
global t
t = parallel(f, batches, progress=False, n_workers=2)
```
That's a bit faster still.
What if we don't actually return the lists of tokens, but create them still?
```
def f(items):
o = [s.split(' ') for s in items]
return [s for s in items]
```
So creating the tokens, isn't taking the time, but returning them over the process boundary is.
```
%%timeit -n 2 -r 3
global t
t = parallel(f, batches, progress=False, n_workers=2)
```
Is numpy any faster?
```
sarr = np.array(ss)
%%timeit -n 2 -r 3
global t
t = np.char.split(sarr)
```
## Spacy
```
from spacy.lang.en import English
def conv_sp(doc): return L(doc).map(str)
class SpTok:
def __init__(self):
nlp = English()
self.tok = nlp.Defaults.create_tokenizer(nlp)
def __call__(self, x): return L(self.tok(str(x))).map(conv_sp)
```
Let's see how long it takes to create a tokenizer in Spacy:
```
%%timeit -n 2 -r 3
SpTok()
nlp = English()
sp_tokenizer = nlp.Defaults.create_tokenizer(nlp)
def spacy_tok(s): return L(sp_tokenizer(str(s))).map(str)
```
Time tokenize in Spacy using a loop:
```
%%timeit -r 3
global t
t = apply(spacy_tok, ss)
```
...and the same thing in parallel:
```
%%timeit -r 3
global t
t = parallel(partial(apply, spacy_tok), batches, progress=False, n_workers=2)
```
...and with more workers:
```
%%timeit -r 3
global t
t = parallel(partial(apply, spacy_tok), batches8, progress=False, n_workers=8)
```
...and with creating the tokenizer in the child process:
```
def f(its):
tok = SpTok()
return [[str(o) for o in tok(p)] for p in its]
%%timeit -r 3
global t
t = parallel(f, batches8, progress=False, n_workers=8)
```
Let's try `pipe`
```
%%timeit -r 3
global t
t = L(nlp.tokenizer.pipe(ss)).map(conv_sp)
def f(its): return L(nlp.tokenizer.pipe(its)).map(conv_sp)
%%timeit -r 3
global t
t = parallel(f, batches8, progress=False, n_workers=8)
test_eq(chunked(range(12),n_chunks=4), [[0, 1, 2], [3, 4, 5], [6, 7, 8], [9, 10, 11]])
test_eq(chunked(range(11),n_chunks=4), [[0, 1, 2], [3, 4, 5], [6, 7, 8], [9, 10]])
test_eq(chunked(range(10),n_chunks=4), [[0, 1, 2], [3, 4, 5], [6, 7, 8], [9]])
test_eq(chunked(range( 9),n_chunks=3), [[0, 1, 2], [3, 4, 5], [6, 7, 8]])
%%timeit -r 3
global t
t = parallel_chunks(f, ss, n_workers=8, progress=False)
def array_split(arr, n): return chunked(arr, math.floor(len(arr)/n))
```
| github_jupyter |
# Lab 05 : Train with mini-batches -- solution
```
# For Google Colaboratory
import sys, os
if 'google.colab' in sys.modules:
# mount google drive
from google.colab import drive
drive.mount('/content/gdrive')
# find automatically the path of the folder containing "file_name" :
file_name = 'minibatch_training_solution.ipynb'
import subprocess
path_to_file = subprocess.check_output('find . -type f -name ' + str(file_name), shell=True).decode("utf-8")
path_to_file = path_to_file.replace(file_name,"").replace('\n',"")
# if previous search failed or too long, comment the previous line and simply write down manually the path below :
#path_to_file = '/content/gdrive/My Drive/CS5242_2021_codes/codes/labs_lecture03/lab05_minibatch_training'
print(path_to_file)
# change current path to the folder containing "file_name"
os.chdir(path_to_file)
!pwd
import torch
import torch.nn as nn
import torch.optim as optim
from random import randint
import utils
```
### Download the data and print the sizes
```
from utils import check_fashion_mnist_dataset_exists
data_path=check_fashion_mnist_dataset_exists()
train_data=torch.load(data_path+'fashion-mnist/train_data.pt')
print(train_data.size())
train_label=torch.load(data_path+'fashion-mnist/train_label.pt')
print(train_label.size())
test_data=torch.load(data_path+'fashion-mnist/test_data.pt')
print(test_data.size())
```
### Make a one layer net class
```
class one_layer_net(nn.Module):
def __init__(self, input_size, output_size):
super(one_layer_net , self).__init__()
self.linear_layer = nn.Linear( input_size, output_size , bias=False)
def forward(self, x):
y = self.linear_layer(x)
prob = torch.softmax(y, dim=1)
return prob
```
### Build the net
```
net=one_layer_net(784,10)
print(net)
```
### Choose the size of the mini-batches
```
bs=200
```
### Train the network (only 5000 iterations) on the train set
```
criterion = nn.NLLLoss()
optimizer=torch.optim.SGD(net.parameters() , lr=0.01 )
for iter in range(1,5000):
# create a minibatch
indices=torch.LongTensor(bs).random_(0,60000)
minibatch_data = train_data[indices]
minibatch_label= train_label[indices]
#reshape them to fit the network
inputs=minibatch_data.view(bs,784)
# feed the input to the net
inputs.requires_grad_()
prob=net(inputs)
# update the weights (all the magic happens here -- we will discuss it later)
log_prob=torch.log(prob)
loss = criterion(log_prob, minibatch_label)
optimizer.zero_grad()
loss.backward()
optimizer.step()
```
### Choose image at random from the test set and see how good/bad are the predictions
```
# choose a picture at random
idx=randint(0, 10000-1)
im=test_data[idx]
# diplay the picture
utils.show(im)
# feed it to the net and display the confidence scores
prob = net( im.view(1,784))
utils.show_prob_fashion_mnist(prob)
```
| github_jupyter |
# Lab 3 - Distance Metrics and Clustering
### Non-Euclidean Distance Metrics
We are most familiar with the typical Euclidian distance metric, ie: given two vectors $\overline{v_1} = [x_1, y_1]$ and $\overline{v_2} = [x_2, y_2]$, the distance $D$ between them is $\sqrt{(x_2 - x_1)^2 + (y_2 - y_1)^2}$. This is generalized for $N$-sized vectors $\overline{a} = [a_1, ..., a_N]$ and $\overline{b} = [b_1, ..., b_N]$ with the following formula:
$$
D = \sqrt{\sum_{i = 1}^{N}(a_i - b_i)^2}
$$
There are other metrics which may be useful, depending on the type of analysis you wish to do. The *Hamming Distance*, $D_H$ is useful when analyzing categorical data:
$$
D_H = \frac{\sum_{i = 1}^{N}|a_i - b_i|}{N}
$$
The *Block Distance*, $D_B$, is useful for calculating distance when only straight paths parallel to your axes are allowed, eg: while navigating through blocks in a large city:
$$
D_B = \sum_{i = 1}^{N}|a_i - b_i|
$$
The *Minkowski Distance*, $D_M$, is a generalized metric that reduces to the block and euclidean distances for $p=1$ and $p=2$, respectively. The value of $p$ can vary by application and is generally tuned by experiment - it can be used for applications ranging from analysing the quality of mibile networks to calculating distances in curved space:
$$
D_M = \big(\sum_{i = 1}^{N}(a_i - b_i)^p\big)^{1/p}
$$
#### Example 1: Distance calculation with varying metrics
Use the **Euclidean**, **Block**, **Hamming**, and **Minkowski** (p=3 and p=4) distance formulas to calculate the distance between the two vectors given below. You may change the values contained in the vectors to see how their distances respond relative to eachother.
```
import numpy as np
vec1 = np.array([1.2, 5.7, 6.1, 2.8, 1.1, 14.8])
vec2 = np.array([5.1, 6.0, 1.3, 28.3, 14.4, -11.9])
```
### Clustering with K-Means
"Clustering" is a method by which a set of *N* vectors can be subdivided into groups, based on the relative "closeness" of one vector to the surrounding vectors. This is done with the "*k*-means algorigthm", and can be seperated into three main parts:
- Check if the data can be clustered. If so, how many clusters are there?
- Determine a point's association with surrounding clusters.
- Repeat until clustering is optimized as much as possible.
Consider *N* vectors of length *n*, $\overline{x_1} = [x_{1, 1}, ..., x{1, n}], ..., \overline{x_N} = [x_{N, 1}, ..., x{N, n}]$. Let there be *k* clusters, labeled $1, ..., k$. We specify the cluster assignment of a given vector by defining an *N*-sized vector, $\overline{c} = [c_1, ..., c_N]$, where $c_i = 1, ..., k$ is the cluster assignment for vector $\overline{x_i}$. The contents of a given cluster can then be defined as $G_j = \{i : c_i = j\}$, with $i$ the number of the cluster.
For each group $G_i$, define a "group-representative $n$-vector" $\overline{z_i}$. We want this representative to be as close to other points in the cluster as possible, so we seek to minimize the distance between representative and cluster points (ie: minimize the quantity $||\overline{x_i} - \overline{z_{c_i}}||$).
To evaluate a given choice of clustering, we use the mean square distance from the vectors to their associated representatives:
$$
J^{clust} = \frac{(||\overline{x_1} - \overline{z_{c_1}}||^2 + ... + ||\overline{x_N} - \overline{z_{c_N}}||^2)}{N}
$$
For optimal clustering, we seek to make $J^{clust}$ as small as possible. We can select $c_i$ to minimize each term: for each data vector $x_i$, select $c_i$ such that $||x_i - z_j||$ is minimized over all $j$. ie: assign each vector $x_i$ to its nearest neighbor amongst the group representatives.
The best group assignment is then the one that minimizes $J^{clust}$. By the above logic, we have
$$
||x_i - z_{c_i}|| = min_{j = 1, ..., k}||x_i - z_j||
$$
so then
$$
J^{clust} = \frac{(min_{j = 1, ..., k}||x_1 - z_j||^2 + ... + min_{j = 1, ..., k}||x_N - z_j||^2)}{N}
$$
### SciKit-Learn `KMeans()`
The `KMeans()` function in the `sklearn.cluster` package can help to easily identify the cluster centers. The documentation for this function can be found [here](https://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html).
Given a `numpy` array, you can run the `KMeans()` function to produce an initial cluster grouping. You will need to specify a guess at a number of clusters (`n_clusters`) and a random state (`random_state`). An example of how to do this is shown in the documentation.
Once the *k*-means operation is run, you can determine cluster membership using the `.labels_` attribute (ie: if you set `kmeans = KMeans(...).fit(X)` with point-array `X`, `kmeans.labels_` returns an array with the group numbers for each point). The `.cluster_centers_` attribute returns the cluster center location for each cluster. The `.predict()` function will predict the cluster membership of any additional points you want to add in or test.
#### Example 2: Find the number of darts players.
Several darts players are throwing darts at a $10\times10$ meter board. The points they hit are stored in the `darts` array below as *x*-*y* coordinate pairs. Guess the number of clusters, *k* - this corresponds to the number of darts players there are. You may want to plot the data to confirm your guess visually.
Run the *k*-means algorithm to calculate and minimize the mean square distance from the vectors to their associated representatives, $J^{clust}$. The goal is to get $J^{clust}$ as small as possible, so that each point is in its most optimal grouping. After optimizing the groups, plot the points and use colors to show which cluster a given point is a member of.
Try this for your chosen *k*-value and other nearby *k*-values to confirm your choice of *k*.
```
%matplotlib inline
import numpy as np
from sklearn.cluster import KMeans
import matplotlib.pyplot as plt
darts = np.array([[1.369, 1.764], [1.250, 2.148], [1.369, 2.046], [1.408, 2.276],
[1.547, 2.199], [1.805, 2.480], [1.765, 3.094], [1.289, 3.120],
[0.833, 2.736], [1.686, 1.585], [2.023, 2.046], [1.289, 2.685],
[1.706, 1.841], [0.952, 1.227], [3.631, 8.030], [3.769, 7.519],
[3.571, 7.749], [3.968, 8.056], [3.769, 6.879], [4.404, 7.059],
[4.365, 7.621], [2.579, 7.340], [3.333, 6.061], [3.333, 7.391],
[3.412, 8.286], [3.948, 8.439], [5.694, 2.659], [5.813, 3.069],
[5.932, 2.634], [5.000, 2.890], [5.436, 2.148], [5.456, 2.762],
[5.218, 2.506], [5.654, 2.097], [6.111, 2.327], [5.575, 2.480],
[5.555, 1.687], [6.726, 1.739], [6.428, 2.915], [5.714, 3.887],
[5.753, 4.322], [5.992, 3.810], [5.436, 3.554], [5.218, 4.884],
[5.019, 4.578], [5.337, 4.373], [5.694, 4.833], [2.420, 2.659],
[1.785, 3.478], [2.202, 2.838], [2.162, 2.455], [1.448, 3.938],
[0.813, 5.063], [0.793, 4.629], [0.992, 3.836], [3.789, 6.010],
[4.146, 6.138], [3.809, 6.342], [1.329, 9.053], [1.825, 8.746],
[1.964, 9.053], [2.341, 8.388], [2.956, 9.130], [5.138, 4.169],
[7.341, 9.028], [7.400, 8.695], [7.480, 8.874], [6.924, 8.746],
[7.261, 8.388], [7.559, 8.516], [8.015, 8.005], [7.876, 8.491],
[7.460, 8.056], [5.615, 3.094], [5.654, 3.478], [5.952, 3.120],
[5.337, 2.915], [5.635, 2.736], [3.274, 8.030], [3.452, 8.107],
[3.392, 7.774], [3.115, 8.132], [7.083, 8.235], [7.400, 7.749],
[7.837, 8.849], [6.686, 9.207], [7.202, 8.670], [7.361, 9.258],
[7.619, 8.363], [7.599, 9.130], [8.829, 8.542], [8.849, 9.232],
[8.551, 8.849], [8.392, 7.877], [8.888, 7.595], [8.869, 6.700],
[8.174, 7.109], [8.452, 7.263], [8.214, 6.828], [7.698, 7.544]])
kmeans = KMeans(n_clusters=4, random_state=0, max_iter=3).fit(darts)
clusters = kmeans.labels_
k=0
plt_color = ['r','b','k','y','g','c']
for i in np.unique(clusters):
indx = np.where(clusters==i)[0]
plt.scatter(darts[indx,0],darts[indx,1],c=plt_color[k])
k = k + 1
plt.title('Clustering dart positions')
plt.show()
```
### Regression with K-Nearest Neighbors
Now that we know how to determine clusters, let's develop a method by which additional points can be classified as belonging to one of the groups. The "K-Nearest Neighbors" algorithm (or KNN for short) can be used for both classification and regression - it can be used to guess the value of a new point based on its similarity to surrounding points. The algorithm is as follows:
- First, find the distance between the new point and all surrounding points.
- Select the closest *k* data points - this is a different *k*-value than the one used to number groups in the previous section!
- Use the average of these *k* datapoints to calculate the predicted value of the new point.
The *k*-value is key in determining an accurate prediction based on the data. This value can be optimized using the training data - by plugging in various given points to the KNN algorithm, we can calculate the error for our chosen *k*. This is simply the difference between the actual and predicted value.
#### Example 3: Different Classes of Irises
Consider the following dataset containing data about irises. The attributes contained in the dataset are as follows:
1. sepal length in cm
2. sepal width in cm
3. petal length in cm
4. petal width in cm
5. class:
- Iris Setosa
- Iris Versicolour
- Iris Virginica
The data has been imported for your convenience below, as well as several steps to walk you through how the classification is done. Try changing the *k*-value (the number of nearest neighbors evaluated, preset as `n_neighbors=5`) to see how the class predictions change. Predict the class of a new iris with sepal length of $5.01cm$, sepal width of $3.72cm$, petal length of $1.23cm$, and petal width of $0.31cm$. This should be the only bit of wholly new code you need to add at the bottom.
The data was used as a courtesy of the UCI Machine Learning Repository, from the [Irises Dataset](https://archive.ics.uci.edu/ml/datasets/Iris).
**References**:
Blake, C. L. and Merz, C. J. 1998. “UCI repository of machine learning databases”. University of California. Available online at: http://www.ics.uci.edu/∼mlearn/MLRepository.html
Robinson, S. "K-Nearest Neighbors Algorithm in Python and Scikit-Learn". StackAbuse.com. Available online at: https://stackabuse.com/k-nearest-neighbors-algorithm-in-python-and-scikit-learn/
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data"
# Assign column names to the dataset
names = ['sepal-length', 'sepal-width', 'petal-length', 'petal-width', 'Class']
# Read dataset to pandas dataframe
dataset = pd.read_csv(url, names=names)
# Shows the first 5 entries in the dataset (only works if code below the following line is commented out)
dataset.head()
# Split dataset into values and labels
my_data = np.array(dataset.iloc[:, :-1].values)
classifications = np.array(dataset.iloc[:, 4].values)
# Split the values into train and test groups - 80% train and 20% test
from sklearn.model_selection import train_test_split
data_train, data_test, class_train, class_test = train_test_split(my_data, classifications, test_size=0.20)
# Use Sci-Kit to create a classification of the training data
# here, we set k=5 to classify points based on 5 nearest neighbors
from sklearn.neighbors import KNeighborsClassifier
classifier = KNeighborsClassifier(n_neighbors=5)
classifier.fit(data_train, class_train)
# Now lets make predictions of the classifications of our test data
predicted_class = classifier.predict(data_test)
# Predicted versus test classes
# print(predicted_class)
# print(class_test)
### Now predict the class of a new iris with the following attributes:
# - sepal length of 5.01cm
# - sepal width of 3.72cm
# - petal length of 1.23cm
# - petal width of 0.31cm
# You will need to input the values into a numpy array, then use the classifier.predict() function as above
'''
Insert Code Here
'''
```
| github_jupyter |
# Deep $Q$-learning
In this notebook, we'll build a neural network that can learn to play games through reinforcement learning. More specifically, we'll use $Q$-learning to train an agent to play a game called [Cart-Pole](https://gym.openai.com/envs/CartPole-v0). In this game, a freely swinging pole is attached to a cart. The cart can move to the left and right, and the goal is to keep the pole upright as long as possible.

We can simulate this game using [OpenAI Gym](https://github.com/openai/gym). First, let's check out how OpenAI Gym works. Then, we'll get into training an agent to play the Cart-Pole game.
```
import gym
import numpy as np
# Create the Cart-Pole game environment
env = gym.make('CartPole-v1')
# Number of possible actions
print('Number of possible actions:', env.action_space.n)
[2018-01-22 23:10:02,350] Making new env: CartPole-v1
Number of possible actions: 2
```
We interact with the simulation through `env`. You can see how many actions are possible from `env.action_space.n`, and to get a random action you can use `env.action_space.sample()`. Passing in an action as an integer to `env.step` will generate the next step in the simulation. This is general to all Gym games.
In the Cart-Pole game, there are two possible actions, moving the cart left or right. So there are two actions we can take, encoded as 0 and 1.
Run the code below to interact with the environment.
```
actions = [] # actions that the agent selects
rewards = [] # obtained rewards
state = env.reset()
while True:
action = env.action_space.sample() # choose a random action
state, reward, done, _ = env.step(action)
rewards.append(reward)
actions.append(action)
if done:
break
```
We can look at the actions and rewards:
```
print('Actions:', actions)
print('Rewards:', rewards)
Actions: [0, 1, 1, 0, 1, 1, 1, 1, 1, 1, 1, 0, 0, 1, 0, 0]
Rewards: [1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0]
```
The game resets after the pole has fallen past a certain angle. For each step while the game is running, it returns a reward of 1.0. The longer the game runs, the more reward we get. Then, our network's goal is to maximize the reward by keeping the pole vertical. It will do this by moving the cart to the left and the right.
## $Q$-Network
To keep track of the action values, we'll use a neural network that accepts a state $s$ as input. The output will be $Q$-values for each available action $a$ (i.e., the output is **all** action values $Q(s,a)$ _corresponding to the input state $s$_).
<img src="assets/q-network.png" width=550px>
For this Cart-Pole game, the state has four values: the position and velocity of the cart, and the position and velocity of the pole. Thus, the neural network has **four inputs**, one for each value in the state, and **two outputs**, one for each possible action.
As explored in the lesson, to get the training target, we'll first use the context provided by the state $s$ to choose an action $a$, then simulate the game using that action. This will get us the next state, $s'$, and the reward $r$. With that, we can calculate $\hat{Q}(s,a) = r + \gamma \max_{a'}{Q(s', a')}$. Then we update the weights by minimizing $(\hat{Q}(s,a) - Q(s,a))^2$.
Below is one implementation of the $Q$-network. It uses two fully connected layers with ReLU activations. Two seems to be good enough, three might be better. Feel free to try it out.
```
import tensorflow as tf
class QNetwork:
def __init__(self, learning_rate=0.01, state_size=4,
action_size=2, hidden_size=10,
name='QNetwork'):
# state inputs to the Q-network
with tf.variable_scope(name):
self.inputs_ = tf.placeholder(tf.float32, [None, state_size], name='inputs')
# One hot encode the actions to later choose the Q-value for the action
self.actions_ = tf.placeholder(tf.int32, [None], name='actions')
one_hot_actions = tf.one_hot(self.actions_, action_size)
# Target Q values for training
self.targetQs_ = tf.placeholder(tf.float32, [None], name='target')
# ReLU hidden layers
self.fc1 = tf.contrib.layers.fully_connected(self.inputs_, hidden_size)
self.fc2 = tf.contrib.layers.fully_connected(self.fc1, hidden_size)
# Linear output layer
self.output = tf.contrib.layers.fully_connected(self.fc2, action_size,
activation_fn=None)
### Train with loss (targetQ - Q)^2
# output has length 2, for two actions. This next line chooses
# one value from output (per row) according to the one-hot encoded actions.
self.Q = tf.reduce_sum(tf.multiply(self.output, one_hot_actions), axis=1)
self.loss = tf.reduce_mean(tf.square(self.targetQs_ - self.Q))
self.opt = tf.train.AdamOptimizer(learning_rate).minimize(self.loss)
```
## Experience replay
Reinforcement learning algorithms can have stability issues due to correlations between states. To reduce correlations when training, we can store the agent's experiences and later draw a random mini-batch of those experiences to train on.
Here, we'll create a `Memory` object that will store our experiences, our transitions $<s, a, r, s'>$. This memory will have a maximum capacity, so we can keep newer experiences in memory while getting rid of older experiences. Then, we'll sample a random mini-batch of transitions $<s, a, r, s'>$ and train on those.
Below, I've implemented a `Memory` object. If you're unfamiliar with `deque`, this is a double-ended queue. You can think of it like a tube open on both sides. You can put objects in either side of the tube. But if it's full, adding anything more will push an object out the other side. This is a great data structure to use for the memory buffer.
```
from collections import deque
class Memory():
def __init__(self, max_size=1000):
self.buffer = deque(maxlen=max_size)
def add(self, experience):
self.buffer.append(experience)
def sample(self, batch_size):
idx = np.random.choice(np.arange(len(self.buffer)),
size=batch_size,
replace=False)
return [self.buffer[ii] for ii in idx]
```
## $Q$-Learning training algorithm
We will use the below algorithm to train the network. For this game, the goal is to keep the pole upright for 195 frames. So we can start a new episode once meeting that goal. The game ends if the pole tilts over too far, or if the cart moves too far the left or right. When a game ends, we'll start a new episode. Now, to train the agent:
* Initialize the memory $D$
* Initialize the action-value network $Q$ with random weights
* **For** episode $\leftarrow 1$ **to** $M$ **do**
* Observe $s_0$
* **For** $t \leftarrow 0$ **to** $T-1$ **do**
* With probability $\epsilon$ select a random action $a_t$, otherwise select $a_t = \mathrm{argmax}_a Q(s_t,a)$
* Execute action $a_t$ in simulator and observe reward $r_{t+1}$ and new state $s_{t+1}$
* Store transition $<s_t, a_t, r_{t+1}, s_{t+1}>$ in memory $D$
* Sample random mini-batch from $D$: $<s_j, a_j, r_j, s'_j>$
* Set $\hat{Q}_j = r_j$ if the episode ends at $j+1$, otherwise set $\hat{Q}_j = r_j + \gamma \max_{a'}{Q(s'_j, a')}$
* Make a gradient descent step with loss $(\hat{Q}_j - Q(s_j, a_j))^2$
* **endfor**
* **endfor**
You are welcome (and encouraged!) to take the time to extend this code to implement some of the improvements that we discussed in the lesson, to include fixed $Q$ targets, double DQNs, prioritized replay, and/or dueling networks.
## Hyperparameters
One of the more difficult aspects of reinforcement learning is the large number of hyperparameters. Not only are we tuning the network, but we're tuning the simulation.
```
train_episodes = 1000 # max number of episodes to learn from
max_steps = 200 # max steps in an episode
gamma = 0.99 # future reward discount
# Exploration parameters
explore_start = 1.0 # exploration probability at start
explore_stop = 0.01 # minimum exploration probability
decay_rate = 0.0001 # exponential decay rate for exploration prob
# Network parameters
hidden_size = 64 # number of units in each Q-network hidden layer
learning_rate = 0.0001 # Q-network learning rate
# Memory parameters
memory_size = 10000 # memory capacity
batch_size = 20 # experience mini-batch size
pretrain_length = batch_size # number experiences to pretrain the memory
tf.reset_default_graph()
mainQN = QNetwork(name='main', hidden_size=hidden_size, learning_rate=learning_rate)
```
## Populate the experience memory
Here we re-initialize the simulation and pre-populate the memory. The agent is taking random actions and storing the transitions in memory. This will help the agent with exploring the game.
```
# Initialize the simulation
env.reset()
# Take one random step to get the pole and cart moving
state, reward, done, _ = env.step(env.action_space.sample())
memory = Memory(max_size=memory_size)
# Make a bunch of random actions and store the experiences
for ii in range(pretrain_length):
# Make a random action
action = env.action_space.sample()
next_state, reward, done, _ = env.step(action)
if done:
# The simulation fails so no next state
next_state = np.zeros(state.shape)
# Add experience to memory
memory.add((state, action, reward, next_state))
# Start new episode
env.reset()
# Take one random step to get the pole and cart moving
state, reward, done, _ = env.step(env.action_space.sample())
else:
# Add experience to memory
memory.add((state, action, reward, next_state))
state = next_state
```
## Training
Below we'll train our agent.
```
# Now train with experiences
saver = tf.train.Saver()
rewards_list = []
with tf.Session() as sess:
# Initialize variables
sess.run(tf.global_variables_initializer())
step = 0
for ep in range(1, train_episodes):
total_reward = 0
t = 0
while t < max_steps:
step += 1
# Uncomment this next line to watch the training
# env.render()
# Explore or Exploit
explore_p = explore_stop + (explore_start - explore_stop)*np.exp(-decay_rate*step)
if explore_p > np.random.rand():
# Make a random action
action = env.action_space.sample()
else:
# Get action from Q-network
feed = {mainQN.inputs_: state.reshape((1, *state.shape))}
Qs = sess.run(mainQN.output, feed_dict=feed)
action = np.argmax(Qs)
# Take action, get new state and reward
next_state, reward, done, _ = env.step(action)
total_reward += reward
if done:
# the episode ends so no next state
next_state = np.zeros(state.shape)
t = max_steps
print('Episode: {}'.format(ep),
'Total reward: {}'.format(total_reward),
'Training loss: {:.4f}'.format(loss),
'Explore P: {:.4f}'.format(explore_p))
rewards_list.append((ep, total_reward))
# Add experience to memory
memory.add((state, action, reward, next_state))
# Start new episode
env.reset()
# Take one random step to get the pole and cart moving
state, reward, done, _ = env.step(env.action_space.sample())
else:
# Add experience to memory
memory.add((state, action, reward, next_state))
state = next_state
t += 1
# Sample mini-batch from memory
batch = memory.sample(batch_size)
states = np.array([each[0] for each in batch])
actions = np.array([each[1] for each in batch])
rewards = np.array([each[2] for each in batch])
next_states = np.array([each[3] for each in batch])
# Train network
target_Qs = sess.run(mainQN.output, feed_dict={mainQN.inputs_: next_states})
# Set target_Qs to 0 for states where episode ends
episode_ends = (next_states == np.zeros(states[0].shape)).all(axis=1)
target_Qs[episode_ends] = (0, 0)
targets = rewards + gamma * np.max(target_Qs, axis=1)
loss, _ = sess.run([mainQN.loss, mainQN.opt],
feed_dict={mainQN.inputs_: states,
mainQN.targetQs_: targets,
mainQN.actions_: actions})
saver.save(sess, "checkpoints/cartpole.ckpt")
Episode: 1 Total reward: 13.0 Training loss: 1.0202 Explore P: 0.9987
Episode: 2 Total reward: 13.0 Training loss: 1.0752 Explore P: 0.9974
Episode: 3 Total reward: 9.0 Training loss: 1.0600 Explore P: 0.9965
Episode: 4 Total reward: 17.0 Training loss: 1.0429 Explore P: 0.9949
Episode: 5 Total reward: 16.0 Training loss: 1.0519 Explore P: 0.9933
Episode: 6 Total reward: 15.0 Training loss: 1.0574 Explore P: 0.9918
Episode: 7 Total reward: 12.0 Training loss: 1.0889 Explore P: 0.9906
Episode: 8 Total reward: 27.0 Training loss: 1.0859 Explore P: 0.9880
Episode: 9 Total reward: 24.0 Training loss: 1.2007 Explore P: 0.9857
Episode: 10 Total reward: 17.0 Training loss: 1.1116 Explore P: 0.9840
Episode: 11 Total reward: 12.0 Training loss: 1.0739 Explore P: 0.9828
Episode: 12 Total reward: 25.0 Training loss: 1.0805 Explore P: 0.9804
Episode: 13 Total reward: 23.0 Training loss: 1.0628 Explore P: 0.9782
Episode: 14 Total reward: 31.0 Training loss: 1.0248 Explore P: 0.9752
Episode: 15 Total reward: 15.0 Training loss: 0.9859 Explore P: 0.9737
Episode: 16 Total reward: 12.0 Training loss: 1.0983 Explore P: 0.9726
Episode: 17 Total reward: 16.0 Training loss: 1.4343 Explore P: 0.9710
Episode: 18 Total reward: 21.0 Training loss: 1.2696 Explore P: 0.9690
Episode: 19 Total reward: 15.0 Training loss: 1.3542 Explore P: 0.9676
Episode: 20 Total reward: 15.0 Training loss: 1.2635 Explore P: 0.9661
Episode: 21 Total reward: 16.0 Training loss: 1.3648 Explore P: 0.9646
Episode: 22 Total reward: 43.0 Training loss: 1.6088 Explore P: 0.9605
Episode: 23 Total reward: 7.0 Training loss: 1.5027 Explore P: 0.9599
Episode: 24 Total reward: 13.0 Training loss: 1.7275 Explore P: 0.9586
Episode: 25 Total reward: 18.0 Training loss: 1.3902 Explore P: 0.9569
Episode: 26 Total reward: 27.0 Training loss: 2.5874 Explore P: 0.9544
Episode: 27 Total reward: 32.0 Training loss: 1.5907 Explore P: 0.9513
Episode: 28 Total reward: 17.0 Training loss: 2.1144 Explore P: 0.9497
Episode: 29 Total reward: 34.0 Training loss: 1.7340 Explore P: 0.9466
Episode: 30 Total reward: 18.0 Training loss: 2.5100 Explore P: 0.9449
Episode: 31 Total reward: 15.0 Training loss: 2.0166 Explore P: 0.9435
Episode: 32 Total reward: 11.0 Training loss: 1.8675 Explore P: 0.9424
Episode: 33 Total reward: 18.0 Training loss: 4.0481 Explore P: 0.9408
Episode: 34 Total reward: 10.0 Training loss: 4.0895 Explore P: 0.9398
Episode: 35 Total reward: 15.0 Training loss: 2.1252 Explore P: 0.9384
Episode: 36 Total reward: 14.0 Training loss: 4.7765 Explore P: 0.9371
Episode: 37 Total reward: 16.0 Training loss: 3.3848 Explore P: 0.9357
Episode: 38 Total reward: 21.0 Training loss: 3.9125 Explore P: 0.9337
Episode: 39 Total reward: 16.0 Training loss: 2.6183 Explore P: 0.9322
Episode: 40 Total reward: 20.0 Training loss: 5.4929 Explore P: 0.9304
Episode: 41 Total reward: 18.0 Training loss: 3.6606 Explore P: 0.9287
Episode: 42 Total reward: 17.0 Training loss: 4.5812 Explore P: 0.9272
Episode: 43 Total reward: 10.0 Training loss: 3.7633 Explore P: 0.9263
Episode: 44 Total reward: 8.0 Training loss: 4.6176 Explore P: 0.9255
Episode: 45 Total reward: 39.0 Training loss: 4.2732 Explore P: 0.9220
Episode: 46 Total reward: 18.0 Training loss: 4.0041 Explore P: 0.9203
Episode: 47 Total reward: 11.0 Training loss: 4.4035 Explore P: 0.9193
Episode: 48 Total reward: 25.0 Training loss: 5.4287 Explore P: 0.9171
Episode: 49 Total reward: 19.0 Training loss: 9.6972 Explore P: 0.9153
Episode: 50 Total reward: 11.0 Training loss: 16.3460 Explore P: 0.9143
Episode: 51 Total reward: 11.0 Training loss: 13.4854 Explore P: 0.9133
Episode: 52 Total reward: 12.0 Training loss: 12.8016 Explore P: 0.9123
Episode: 53 Total reward: 13.0 Training loss: 5.8589 Explore P: 0.9111
Episode: 54 Total reward: 12.0 Training loss: 8.5924 Explore P: 0.9100
Episode: 55 Total reward: 19.0 Training loss: 8.6204 Explore P: 0.9083
Episode: 56 Total reward: 36.0 Training loss: 14.2701 Explore P: 0.9051
Episode: 57 Total reward: 9.0 Training loss: 4.5481 Explore P: 0.9043
Episode: 58 Total reward: 22.0 Training loss: 12.9695 Explore P: 0.9023
Episode: 59 Total reward: 36.0 Training loss: 11.2639 Explore P: 0.8991
Episode: 60 Total reward: 16.0 Training loss: 7.7648 Explore P: 0.8977
Episode: 61 Total reward: 31.0 Training loss: 4.6997 Explore P: 0.8949
Episode: 62 Total reward: 13.0 Training loss: 5.9755 Explore P: 0.8938
Episode: 63 Total reward: 10.0 Training loss: 39.1040 Explore P: 0.8929
Episode: 64 Total reward: 14.0 Training loss: 23.2767 Explore P: 0.8917
Episode: 65 Total reward: 12.0 Training loss: 9.3477 Explore P: 0.8906
Episode: 66 Total reward: 20.0 Training loss: 6.4336 Explore P: 0.8888
Episode: 67 Total reward: 29.0 Training loss: 17.1522 Explore P: 0.8863
Episode: 68 Total reward: 13.0 Training loss: 39.3250 Explore P: 0.8852
Episode: 69 Total reward: 20.0 Training loss: 6.2099 Explore P: 0.8834
Episode: 70 Total reward: 15.0 Training loss: 20.9229 Explore P: 0.8821
Episode: 71 Total reward: 27.0 Training loss: 24.7817 Explore P: 0.8797
Episode: 72 Total reward: 12.0 Training loss: 20.7842 Explore P: 0.8787
Episode: 73 Total reward: 15.0 Training loss: 12.3202 Explore P: 0.8774
Episode: 74 Total reward: 31.0 Training loss: 9.2270 Explore P: 0.8747
Episode: 75 Total reward: 13.0 Training loss: 19.8264 Explore P: 0.8736
Episode: 76 Total reward: 20.0 Training loss: 72.9411 Explore P: 0.8719
Episode: 77 Total reward: 27.0 Training loss: 5.2214 Explore P: 0.8695
Episode: 78 Total reward: 14.0 Training loss: 39.3913 Explore P: 0.8683
Episode: 79 Total reward: 16.0 Training loss: 7.9491 Explore P: 0.8670
Episode: 80 Total reward: 18.0 Training loss: 10.8364 Explore P: 0.8654
Episode: 81 Total reward: 16.0 Training loss: 22.2031 Explore P: 0.8641
Episode: 82 Total reward: 21.0 Training loss: 23.6590 Explore P: 0.8623
Episode: 83 Total reward: 13.0 Training loss: 8.4819 Explore P: 0.8612
Episode: 84 Total reward: 10.0 Training loss: 13.3548 Explore P: 0.8603
Episode: 85 Total reward: 13.0 Training loss: 18.0272 Explore P: 0.8592
Episode: 86 Total reward: 24.0 Training loss: 42.1243 Explore P: 0.8572
Episode: 87 Total reward: 9.0 Training loss: 30.8526 Explore P: 0.8564
Episode: 88 Total reward: 22.0 Training loss: 36.6084 Explore P: 0.8546
Episode: 89 Total reward: 7.0 Training loss: 10.5430 Explore P: 0.8540
Episode: 90 Total reward: 12.0 Training loss: 25.5808 Explore P: 0.8529
Episode: 91 Total reward: 17.0 Training loss: 47.3073 Explore P: 0.8515
Episode: 92 Total reward: 21.0 Training loss: 7.9998 Explore P: 0.8498
Episode: 93 Total reward: 15.0 Training loss: 66.6464 Explore P: 0.8485
Episode: 94 Total reward: 17.0 Training loss: 95.6354 Explore P: 0.8471
Episode: 95 Total reward: 23.0 Training loss: 57.4714 Explore P: 0.8451
Episode: 96 Total reward: 11.0 Training loss: 40.7717 Explore P: 0.8442
Episode: 97 Total reward: 13.0 Training loss: 43.3380 Explore P: 0.8431
Episode: 98 Total reward: 9.0 Training loss: 10.8368 Explore P: 0.8424
Episode: 99 Total reward: 21.0 Training loss: 57.7325 Explore P: 0.8406
Episode: 100 Total reward: 11.0 Training loss: 9.7291 Explore P: 0.8397
Episode: 101 Total reward: 10.0 Training loss: 10.4052 Explore P: 0.8389
Episode: 102 Total reward: 26.0 Training loss: 60.4829 Explore P: 0.8368
Episode: 103 Total reward: 34.0 Training loss: 9.0924 Explore P: 0.8339
Episode: 104 Total reward: 30.0 Training loss: 178.0664 Explore P: 0.8315
Episode: 105 Total reward: 14.0 Training loss: 9.0423 Explore P: 0.8303
Episode: 106 Total reward: 18.0 Training loss: 126.9380 Explore P: 0.8289
Episode: 107 Total reward: 11.0 Training loss: 58.9921 Explore P: 0.8280
Episode: 108 Total reward: 20.0 Training loss: 9.1945 Explore P: 0.8263
Episode: 109 Total reward: 9.0 Training loss: 9.2887 Explore P: 0.8256
Episode: 110 Total reward: 29.0 Training loss: 20.7970 Explore P: 0.8232
Episode: 111 Total reward: 17.0 Training loss: 144.6258 Explore P: 0.8218
Episode: 112 Total reward: 15.0 Training loss: 82.4089 Explore P: 0.8206
Episode: 113 Total reward: 15.0 Training loss: 39.9963 Explore P: 0.8194
Episode: 114 Total reward: 8.0 Training loss: 9.8394 Explore P: 0.8188
Episode: 115 Total reward: 29.0 Training loss: 76.9930 Explore P: 0.8164
Episode: 116 Total reward: 21.0 Training loss: 25.0172 Explore P: 0.8147
Episode: 117 Total reward: 24.0 Training loss: 143.5481 Explore P: 0.8128
Episode: 118 Total reward: 35.0 Training loss: 86.5429 Explore P: 0.8100
Episode: 119 Total reward: 28.0 Training loss: 8.4315 Explore P: 0.8078
Episode: 120 Total reward: 13.0 Training loss: 25.7062 Explore P: 0.8067
Episode: 121 Total reward: 9.0 Training loss: 6.5005 Explore P: 0.8060
Episode: 122 Total reward: 32.0 Training loss: 90.7984 Explore P: 0.8035
Episode: 123 Total reward: 21.0 Training loss: 130.2779 Explore P: 0.8018
Episode: 124 Total reward: 15.0 Training loss: 167.6294 Explore P: 0.8006
Episode: 125 Total reward: 15.0 Training loss: 74.7611 Explore P: 0.7994
Episode: 126 Total reward: 20.0 Training loss: 119.3178 Explore P: 0.7978
Episode: 127 Total reward: 18.0 Training loss: 196.5175 Explore P: 0.7964
Episode: 128 Total reward: 8.0 Training loss: 45.2131 Explore P: 0.7958
Episode: 129 Total reward: 24.0 Training loss: 86.0374 Explore P: 0.7939
Episode: 130 Total reward: 11.0 Training loss: 7.8129 Explore P: 0.7931
Episode: 131 Total reward: 11.0 Training loss: 76.8442 Explore P: 0.7922
Episode: 132 Total reward: 28.0 Training loss: 196.6863 Explore P: 0.7900
Episode: 133 Total reward: 9.0 Training loss: 45.7586 Explore P: 0.7893
Episode: 134 Total reward: 21.0 Training loss: 5.8484 Explore P: 0.7877
Episode: 135 Total reward: 10.0 Training loss: 7.3919 Explore P: 0.7869
Episode: 136 Total reward: 17.0 Training loss: 12.3142 Explore P: 0.7856
Episode: 137 Total reward: 16.0 Training loss: 75.7170 Explore P: 0.7843
Episode: 138 Total reward: 12.0 Training loss: 145.3568 Explore P: 0.7834
Episode: 139 Total reward: 27.0 Training loss: 121.1114 Explore P: 0.7813
Episode: 140 Total reward: 26.0 Training loss: 7.3243 Explore P: 0.7793
Episode: 141 Total reward: 40.0 Training loss: 10.6523 Explore P: 0.7762
Episode: 142 Total reward: 14.0 Training loss: 6.9482 Explore P: 0.7752
Episode: 143 Total reward: 24.0 Training loss: 137.7784 Explore P: 0.7733
Episode: 144 Total reward: 12.0 Training loss: 98.8381 Explore P: 0.7724
Episode: 145 Total reward: 8.0 Training loss: 14.1739 Explore P: 0.7718
Episode: 146 Total reward: 51.0 Training loss: 69.1545 Explore P: 0.7679
Episode: 147 Total reward: 29.0 Training loss: 249.9989 Explore P: 0.7657
Episode: 148 Total reward: 9.0 Training loss: 140.8663 Explore P: 0.7651
Episode: 149 Total reward: 13.0 Training loss: 141.5930 Explore P: 0.7641
Episode: 150 Total reward: 19.0 Training loss: 12.6228 Explore P: 0.7627
Episode: 151 Total reward: 19.0 Training loss: 136.3315 Explore P: 0.7612
Episode: 152 Total reward: 10.0 Training loss: 110.3699 Explore P: 0.7605
Episode: 153 Total reward: 18.0 Training loss: 8.3900 Explore P: 0.7591
Episode: 154 Total reward: 18.0 Training loss: 96.3717 Explore P: 0.7578
Episode: 155 Total reward: 7.0 Training loss: 6.0889 Explore P: 0.7573
Episode: 156 Total reward: 15.0 Training loss: 126.7419 Explore P: 0.7561
Episode: 157 Total reward: 15.0 Training loss: 67.2544 Explore P: 0.7550
Episode: 158 Total reward: 26.0 Training loss: 12.2839 Explore P: 0.7531
Episode: 159 Total reward: 20.0 Training loss: 5.8118 Explore P: 0.7516
Episode: 160 Total reward: 10.0 Training loss: 96.4570 Explore P: 0.7509
Episode: 161 Total reward: 23.0 Training loss: 7.6207 Explore P: 0.7492
Episode: 162 Total reward: 18.0 Training loss: 66.5249 Explore P: 0.7478
Episode: 163 Total reward: 18.0 Training loss: 111.3273 Explore P: 0.7465
Episode: 164 Total reward: 21.0 Training loss: 11.5292 Explore P: 0.7450
Episode: 165 Total reward: 17.0 Training loss: 6.3130 Explore P: 0.7437
Episode: 166 Total reward: 22.0 Training loss: 153.8167 Explore P: 0.7421
Episode: 167 Total reward: 17.0 Training loss: 7.0915 Explore P: 0.7408
Episode: 168 Total reward: 34.0 Training loss: 228.3831 Explore P: 0.7384
Episode: 169 Total reward: 13.0 Training loss: 8.5996 Explore P: 0.7374
Episode: 170 Total reward: 13.0 Training loss: 90.8898 Explore P: 0.7365
Episode: 171 Total reward: 20.0 Training loss: 4.8179 Explore P: 0.7350
Episode: 172 Total reward: 9.0 Training loss: 6.2508 Explore P: 0.7344
Episode: 173 Total reward: 14.0 Training loss: 5.2401 Explore P: 0.7334
Episode: 174 Total reward: 12.0 Training loss: 3.9268 Explore P: 0.7325
Episode: 175 Total reward: 12.0 Training loss: 5.6376 Explore P: 0.7316
Episode: 176 Total reward: 11.0 Training loss: 44.5308 Explore P: 0.7308
Episode: 177 Total reward: 12.0 Training loss: 4.9717 Explore P: 0.7300
Episode: 178 Total reward: 9.0 Training loss: 181.0085 Explore P: 0.7293
Episode: 179 Total reward: 11.0 Training loss: 73.1134 Explore P: 0.7285
Episode: 180 Total reward: 13.0 Training loss: 87.3085 Explore P: 0.7276
Episode: 181 Total reward: 12.0 Training loss: 121.6627 Explore P: 0.7267
Episode: 182 Total reward: 12.0 Training loss: 58.1967 Explore P: 0.7259
Episode: 183 Total reward: 13.0 Training loss: 85.1540 Explore P: 0.7249
Episode: 184 Total reward: 16.0 Training loss: 5.1214 Explore P: 0.7238
Episode: 185 Total reward: 19.0 Training loss: 69.1839 Explore P: 0.7224
Episode: 186 Total reward: 7.0 Training loss: 63.2256 Explore P: 0.7219
Episode: 187 Total reward: 17.0 Training loss: 73.2788 Explore P: 0.7207
Episode: 188 Total reward: 15.0 Training loss: 78.6213 Explore P: 0.7197
Episode: 189 Total reward: 11.0 Training loss: 88.5211 Explore P: 0.7189
Episode: 190 Total reward: 14.0 Training loss: 60.1332 Explore P: 0.7179
Episode: 191 Total reward: 15.0 Training loss: 135.7724 Explore P: 0.7168
Episode: 192 Total reward: 15.0 Training loss: 156.9691 Explore P: 0.7158
Episode: 193 Total reward: 17.0 Training loss: 93.3756 Explore P: 0.7146
Episode: 194 Total reward: 12.0 Training loss: 3.0462 Explore P: 0.7137
Episode: 195 Total reward: 9.0 Training loss: 119.2650 Explore P: 0.7131
Episode: 196 Total reward: 13.0 Training loss: 66.6383 Explore P: 0.7122
Episode: 197 Total reward: 9.0 Training loss: 113.7849 Explore P: 0.7116
Episode: 198 Total reward: 13.0 Training loss: 54.6072 Explore P: 0.7106
Episode: 199 Total reward: 19.0 Training loss: 54.8980 Explore P: 0.7093
Episode: 200 Total reward: 20.0 Training loss: 155.5480 Explore P: 0.7079
Episode: 201 Total reward: 10.0 Training loss: 45.8685 Explore P: 0.7072
Episode: 202 Total reward: 14.0 Training loss: 53.5145 Explore P: 0.7062
Episode: 203 Total reward: 8.0 Training loss: 107.9623 Explore P: 0.7057
Episode: 204 Total reward: 21.0 Training loss: 40.2749 Explore P: 0.7042
Episode: 205 Total reward: 32.0 Training loss: 43.2627 Explore P: 0.7020
Episode: 206 Total reward: 9.0 Training loss: 55.5398 Explore P: 0.7014
Episode: 207 Total reward: 15.0 Training loss: 1.9959 Explore P: 0.7004
Episode: 208 Total reward: 12.0 Training loss: 105.3751 Explore P: 0.6995
Episode: 209 Total reward: 11.0 Training loss: 40.8319 Explore P: 0.6988
Episode: 210 Total reward: 10.0 Training loss: 89.7147 Explore P: 0.6981
Episode: 211 Total reward: 10.0 Training loss: 1.1946 Explore P: 0.6974
Episode: 212 Total reward: 10.0 Training loss: 80.6916 Explore P: 0.6967
Episode: 213 Total reward: 24.0 Training loss: 88.0977 Explore P: 0.6951
Episode: 214 Total reward: 14.0 Training loss: 46.4105 Explore P: 0.6941
Episode: 215 Total reward: 11.0 Training loss: 40.3726 Explore P: 0.6933
Episode: 216 Total reward: 11.0 Training loss: 3.0770 Explore P: 0.6926
Episode: 217 Total reward: 8.0 Training loss: 1.7495 Explore P: 0.6921
Episode: 218 Total reward: 16.0 Training loss: 1.5615 Explore P: 0.6910
Episode: 219 Total reward: 17.0 Training loss: 2.0250 Explore P: 0.6898
Episode: 220 Total reward: 18.0 Training loss: 37.8432 Explore P: 0.6886
Episode: 221 Total reward: 17.0 Training loss: 1.9049 Explore P: 0.6874
Episode: 222 Total reward: 16.0 Training loss: 1.9652 Explore P: 0.6863
Episode: 223 Total reward: 16.0 Training loss: 1.4384 Explore P: 0.6853
Episode: 224 Total reward: 27.0 Training loss: 66.0615 Explore P: 0.6834
Episode: 225 Total reward: 9.0 Training loss: 0.8478 Explore P: 0.6828
Episode: 226 Total reward: 14.0 Training loss: 1.0319 Explore P: 0.6819
Episode: 227 Total reward: 17.0 Training loss: 97.6957 Explore P: 0.6808
Episode: 228 Total reward: 8.0 Training loss: 68.0521 Explore P: 0.6802
Episode: 229 Total reward: 9.0 Training loss: 110.8437 Explore P: 0.6796
Episode: 230 Total reward: 19.0 Training loss: 1.6856 Explore P: 0.6783
Episode: 231 Total reward: 9.0 Training loss: 2.0634 Explore P: 0.6777
Episode: 232 Total reward: 11.0 Training loss: 32.0714 Explore P: 0.6770
Episode: 233 Total reward: 9.0 Training loss: 2.0387 Explore P: 0.6764
Episode: 234 Total reward: 13.0 Training loss: 66.9349 Explore P: 0.6755
Episode: 235 Total reward: 14.0 Training loss: 110.6725 Explore P: 0.6746
Episode: 236 Total reward: 18.0 Training loss: 1.0585 Explore P: 0.6734
Episode: 237 Total reward: 11.0 Training loss: 117.0101 Explore P: 0.6727
Episode: 238 Total reward: 7.0 Training loss: 2.6115 Explore P: 0.6722
Episode: 239 Total reward: 10.0 Training loss: 124.7320 Explore P: 0.6716
Episode: 240 Total reward: 18.0 Training loss: 2.5475 Explore P: 0.6704
Episode: 241 Total reward: 37.0 Training loss: 2.1454 Explore P: 0.6679
Episode: 242 Total reward: 11.0 Training loss: 23.6042 Explore P: 0.6672
Episode: 243 Total reward: 32.0 Training loss: 1.4344 Explore P: 0.6651
Episode: 244 Total reward: 9.0 Training loss: 1.5328 Explore P: 0.6645
Episode: 245 Total reward: 14.0 Training loss: 84.7870 Explore P: 0.6636
Episode: 246 Total reward: 12.0 Training loss: 2.7292 Explore P: 0.6628
Episode: 247 Total reward: 26.0 Training loss: 40.6692 Explore P: 0.6611
Episode: 248 Total reward: 12.0 Training loss: 22.0901 Explore P: 0.6603
Episode: 249 Total reward: 15.0 Training loss: 37.9304 Explore P: 0.6594
Episode: 250 Total reward: 20.0 Training loss: 1.4137 Explore P: 0.6581
Episode: 251 Total reward: 16.0 Training loss: 1.7831 Explore P: 0.6570
Episode: 252 Total reward: 9.0 Training loss: 38.0640 Explore P: 0.6565
Episode: 253 Total reward: 17.0 Training loss: 21.7703 Explore P: 0.6554
Episode: 254 Total reward: 24.0 Training loss: 40.3204 Explore P: 0.6538
Episode: 255 Total reward: 30.0 Training loss: 43.4179 Explore P: 0.6519
Episode: 256 Total reward: 11.0 Training loss: 60.9330 Explore P: 0.6512
Episode: 257 Total reward: 14.0 Training loss: 66.6886 Explore P: 0.6503
Episode: 258 Total reward: 15.0 Training loss: 2.5639 Explore P: 0.6493
Episode: 259 Total reward: 19.0 Training loss: 2.6969 Explore P: 0.6481
Episode: 260 Total reward: 10.0 Training loss: 2.6837 Explore P: 0.6475
Episode: 261 Total reward: 30.0 Training loss: 20.6603 Explore P: 0.6456
Episode: 262 Total reward: 17.0 Training loss: 32.1585 Explore P: 0.6445
Episode: 263 Total reward: 15.0 Training loss: 1.0833 Explore P: 0.6435
Episode: 264 Total reward: 13.0 Training loss: 81.4551 Explore P: 0.6427
Episode: 265 Total reward: 17.0 Training loss: 3.3823 Explore P: 0.6416
Episode: 266 Total reward: 11.0 Training loss: 36.3942 Explore P: 0.6409
Episode: 267 Total reward: 11.0 Training loss: 1.6628 Explore P: 0.6402
Episode: 268 Total reward: 33.0 Training loss: 26.9925 Explore P: 0.6382
Episode: 269 Total reward: 18.0 Training loss: 45.8608 Explore P: 0.6370
Episode: 270 Total reward: 20.0 Training loss: 2.7911 Explore P: 0.6358
Episode: 271 Total reward: 10.0 Training loss: 35.9215 Explore P: 0.6352
Episode: 272 Total reward: 14.0 Training loss: 2.5923 Explore P: 0.6343
Episode: 273 Total reward: 16.0 Training loss: 41.2339 Explore P: 0.6333
Episode: 274 Total reward: 18.0 Training loss: 46.7318 Explore P: 0.6322
Episode: 275 Total reward: 14.0 Training loss: 2.7245 Explore P: 0.6313
Episode: 276 Total reward: 8.0 Training loss: 16.2681 Explore P: 0.6308
Episode: 277 Total reward: 10.0 Training loss: 21.6856 Explore P: 0.6302
Episode: 278 Total reward: 12.0 Training loss: 1.7879 Explore P: 0.6294
Episode: 279 Total reward: 10.0 Training loss: 97.1567 Explore P: 0.6288
Episode: 280 Total reward: 16.0 Training loss: 3.4710 Explore P: 0.6278
Episode: 281 Total reward: 14.0 Training loss: 65.8457 Explore P: 0.6270
Episode: 282 Total reward: 21.0 Training loss: 32.4442 Explore P: 0.6257
Episode: 283 Total reward: 17.0 Training loss: 48.0136 Explore P: 0.6246
Episode: 284 Total reward: 11.0 Training loss: 2.8833 Explore P: 0.6239
Episode: 285 Total reward: 16.0 Training loss: 92.6062 Explore P: 0.6230
Episode: 286 Total reward: 16.0 Training loss: 19.1051 Explore P: 0.6220
Episode: 287 Total reward: 7.0 Training loss: 1.8220 Explore P: 0.6216
Episode: 288 Total reward: 16.0 Training loss: 41.3844 Explore P: 0.6206
Episode: 289 Total reward: 18.0 Training loss: 50.0580 Explore P: 0.6195
Episode: 290 Total reward: 13.0 Training loss: 83.2142 Explore P: 0.6187
Episode: 291 Total reward: 14.0 Training loss: 70.2605 Explore P: 0.6178
Episode: 292 Total reward: 16.0 Training loss: 53.9664 Explore P: 0.6169
Episode: 293 Total reward: 17.0 Training loss: 3.2764 Explore P: 0.6158
Episode: 294 Total reward: 18.0 Training loss: 17.7963 Explore P: 0.6147
Episode: 295 Total reward: 17.0 Training loss: 32.3772 Explore P: 0.6137
Episode: 296 Total reward: 32.0 Training loss: 18.3755 Explore P: 0.6118
Episode: 297 Total reward: 29.0 Training loss: 17.1377 Explore P: 0.6100
Episode: 298 Total reward: 12.0 Training loss: 14.2922 Explore P: 0.6093
Episode: 299 Total reward: 14.0 Training loss: 29.2226 Explore P: 0.6085
Episode: 300 Total reward: 17.0 Training loss: 38.9089 Explore P: 0.6075
Episode: 301 Total reward: 9.0 Training loss: 62.2483 Explore P: 0.6069
Episode: 302 Total reward: 22.0 Training loss: 2.3240 Explore P: 0.6056
Episode: 303 Total reward: 16.0 Training loss: 0.9979 Explore P: 0.6047
Episode: 304 Total reward: 8.0 Training loss: 67.9231 Explore P: 0.6042
Episode: 305 Total reward: 13.0 Training loss: 33.0928 Explore P: 0.6034
Episode: 306 Total reward: 20.0 Training loss: 1.3173 Explore P: 0.6022
Episode: 307 Total reward: 23.0 Training loss: 50.2106 Explore P: 0.6009
Episode: 308 Total reward: 17.0 Training loss: 52.5245 Explore P: 0.5999
Episode: 309 Total reward: 20.0 Training loss: 32.5832 Explore P: 0.5987
Episode: 310 Total reward: 19.0 Training loss: 29.0224 Explore P: 0.5976
Episode: 311 Total reward: 19.0 Training loss: 29.8863 Explore P: 0.5965
Episode: 312 Total reward: 27.0 Training loss: 34.4016 Explore P: 0.5949
Episode: 313 Total reward: 9.0 Training loss: 1.1433 Explore P: 0.5944
Episode: 314 Total reward: 20.0 Training loss: 28.8137 Explore P: 0.5932
Episode: 315 Total reward: 24.0 Training loss: 48.5379 Explore P: 0.5918
Episode: 316 Total reward: 28.0 Training loss: 45.2671 Explore P: 0.5902
Episode: 317 Total reward: 13.0 Training loss: 45.9822 Explore P: 0.5894
Episode: 318 Total reward: 12.0 Training loss: 86.3972 Explore P: 0.5887
Episode: 319 Total reward: 10.0 Training loss: 11.2909 Explore P: 0.5881
Episode: 320 Total reward: 11.0 Training loss: 36.5474 Explore P: 0.5875
Episode: 321 Total reward: 13.0 Training loss: 1.1439 Explore P: 0.5867
Episode: 322 Total reward: 8.0 Training loss: 12.6978 Explore P: 0.5863
Episode: 323 Total reward: 20.0 Training loss: 31.7664 Explore P: 0.5851
Episode: 324 Total reward: 8.0 Training loss: 29.4243 Explore P: 0.5847
Episode: 325 Total reward: 13.0 Training loss: 12.2373 Explore P: 0.5839
Episode: 326 Total reward: 19.0 Training loss: 24.2228 Explore P: 0.5828
Episode: 327 Total reward: 68.0 Training loss: 0.7256 Explore P: 0.5790
Episode: 328 Total reward: 11.0 Training loss: 1.2313 Explore P: 0.5783
Episode: 329 Total reward: 15.0 Training loss: 1.3319 Explore P: 0.5775
Episode: 330 Total reward: 53.0 Training loss: 9.9350 Explore P: 0.5745
Episode: 331 Total reward: 74.0 Training loss: 1.4366 Explore P: 0.5703
Episode: 332 Total reward: 16.0 Training loss: 11.2724 Explore P: 0.5694
Episode: 333 Total reward: 34.0 Training loss: 10.6128 Explore P: 0.5675
Episode: 334 Total reward: 27.0 Training loss: 14.9559 Explore P: 0.5660
Episode: 335 Total reward: 31.0 Training loss: 16.6541 Explore P: 0.5643
Episode: 336 Total reward: 49.0 Training loss: 23.3966 Explore P: 0.5616
Episode: 337 Total reward: 40.0 Training loss: 45.3419 Explore P: 0.5594
Episode: 338 Total reward: 71.0 Training loss: 0.8244 Explore P: 0.5555
Episode: 339 Total reward: 56.0 Training loss: 41.4562 Explore P: 0.5525
Episode: 340 Total reward: 18.0 Training loss: 1.2548 Explore P: 0.5515
Episode: 341 Total reward: 56.0 Training loss: 1.5400 Explore P: 0.5485
Episode: 342 Total reward: 34.0 Training loss: 12.0206 Explore P: 0.5466
Episode: 343 Total reward: 67.0 Training loss: 1.4189 Explore P: 0.5430
Episode: 344 Total reward: 27.0 Training loss: 1.3138 Explore P: 0.5416
Episode: 345 Total reward: 42.0 Training loss: 1.1650 Explore P: 0.5394
Episode: 346 Total reward: 23.0 Training loss: 23.1743 Explore P: 0.5382
Episode: 347 Total reward: 54.0 Training loss: 0.6971 Explore P: 0.5353
Episode: 348 Total reward: 34.0 Training loss: 27.2789 Explore P: 0.5335
Episode: 349 Total reward: 25.0 Training loss: 37.4133 Explore P: 0.5322
Episode: 350 Total reward: 20.0 Training loss: 1.6443 Explore P: 0.5312
Episode: 351 Total reward: 26.0 Training loss: 12.6839 Explore P: 0.5298
Episode: 352 Total reward: 40.0 Training loss: 13.3593 Explore P: 0.5278
Episode: 353 Total reward: 18.0 Training loss: 1.7079 Explore P: 0.5268
Episode: 354 Total reward: 47.0 Training loss: 32.5788 Explore P: 0.5244
Episode: 355 Total reward: 20.0 Training loss: 1.6101 Explore P: 0.5234
Episode: 356 Total reward: 53.0 Training loss: 2.5321 Explore P: 0.5207
Episode: 357 Total reward: 15.0 Training loss: 1.6396 Explore P: 0.5199
Episode: 358 Total reward: 76.0 Training loss: 20.8058 Explore P: 0.5160
Episode: 359 Total reward: 12.0 Training loss: 13.0315 Explore P: 0.5154
Episode: 360 Total reward: 42.0 Training loss: 10.1313 Explore P: 0.5133
Episode: 361 Total reward: 53.0 Training loss: 25.4319 Explore P: 0.5106
Episode: 362 Total reward: 33.0 Training loss: 26.4256 Explore P: 0.5090
Episode: 363 Total reward: 85.0 Training loss: 20.2429 Explore P: 0.5048
Episode: 364 Total reward: 23.0 Training loss: 16.1083 Explore P: 0.5036
Episode: 365 Total reward: 30.0 Training loss: 1.6888 Explore P: 0.5022
Episode: 366 Total reward: 66.0 Training loss: 2.0408 Explore P: 0.4989
Episode: 367 Total reward: 37.0 Training loss: 18.6438 Explore P: 0.4971
Episode: 368 Total reward: 50.0 Training loss: 20.1544 Explore P: 0.4947
Episode: 369 Total reward: 78.0 Training loss: 23.8497 Explore P: 0.4909
Episode: 370 Total reward: 83.0 Training loss: 20.6897 Explore P: 0.4869
Episode: 371 Total reward: 44.0 Training loss: 25.4317 Explore P: 0.4849
Episode: 372 Total reward: 44.0 Training loss: 1.5212 Explore P: 0.4828
Episode: 373 Total reward: 14.0 Training loss: 1.5019 Explore P: 0.4821
Episode: 374 Total reward: 31.0 Training loss: 1.8348 Explore P: 0.4806
Episode: 375 Total reward: 25.0 Training loss: 19.7533 Explore P: 0.4795
Episode: 376 Total reward: 51.0 Training loss: 1.5433 Explore P: 0.4771
Episode: 377 Total reward: 23.0 Training loss: 12.9174 Explore P: 0.4760
Episode: 378 Total reward: 67.0 Training loss: 27.2318 Explore P: 0.4729
Episode: 379 Total reward: 26.0 Training loss: 1.9319 Explore P: 0.4717
Episode: 380 Total reward: 35.0 Training loss: 43.2445 Explore P: 0.4701
Episode: 381 Total reward: 33.0 Training loss: 1.5195 Explore P: 0.4686
Episode: 382 Total reward: 30.0 Training loss: 15.4622 Explore P: 0.4672
Episode: 383 Total reward: 12.0 Training loss: 1.8349 Explore P: 0.4666
Episode: 384 Total reward: 25.0 Training loss: 47.7600 Explore P: 0.4655
Episode: 385 Total reward: 36.0 Training loss: 29.6753 Explore P: 0.4639
Episode: 386 Total reward: 50.0 Training loss: 1.1244 Explore P: 0.4616
Episode: 387 Total reward: 35.0 Training loss: 1.0955 Explore P: 0.4600
Episode: 388 Total reward: 52.0 Training loss: 24.9624 Explore P: 0.4577
Episode: 389 Total reward: 52.0 Training loss: 28.2028 Explore P: 0.4554
Episode: 390 Total reward: 132.0 Training loss: 30.5190 Explore P: 0.4495
Episode: 391 Total reward: 25.0 Training loss: 10.3908 Explore P: 0.4484
Episode: 392 Total reward: 56.0 Training loss: 14.1483 Explore P: 0.4460
Episode: 393 Total reward: 110.0 Training loss: 2.0169 Explore P: 0.4412
Episode: 394 Total reward: 78.0 Training loss: 1.2122 Explore P: 0.4379
Episode: 395 Total reward: 44.0 Training loss: 56.4728 Explore P: 0.4360
Episode: 396 Total reward: 90.0 Training loss: 65.6667 Explore P: 0.4322
Episode: 397 Total reward: 36.0 Training loss: 0.9032 Explore P: 0.4307
Episode: 398 Total reward: 40.0 Training loss: 0.8414 Explore P: 0.4290
Episode: 399 Total reward: 109.0 Training loss: 42.5467 Explore P: 0.4244
Episode: 400 Total reward: 37.0 Training loss: 2.6053 Explore P: 0.4229
Episode: 401 Total reward: 62.0 Training loss: 1.2301 Explore P: 0.4203
Episode: 402 Total reward: 42.0 Training loss: 1.1384 Explore P: 0.4186
Episode: 403 Total reward: 71.0 Training loss: 1.6765 Explore P: 0.4157
Episode: 404 Total reward: 88.0 Training loss: 2.4000 Explore P: 0.4122
Episode: 405 Total reward: 55.0 Training loss: 24.7748 Explore P: 0.4100
Episode: 406 Total reward: 33.0 Training loss: 13.5934 Explore P: 0.4087
Episode: 407 Total reward: 44.0 Training loss: 14.6865 Explore P: 0.4069
Episode: 408 Total reward: 40.0 Training loss: 2.0898 Explore P: 0.4053
Episode: 409 Total reward: 98.0 Training loss: 2.1043 Explore P: 0.4015
Episode: 410 Total reward: 63.0 Training loss: 11.3562 Explore P: 0.3990
Episode: 411 Total reward: 50.0 Training loss: 14.1151 Explore P: 0.3971
Episode: 412 Total reward: 44.0 Training loss: 1.2370 Explore P: 0.3954
Episode: 413 Total reward: 56.0 Training loss: 2.1136 Explore P: 0.3932
Episode: 414 Total reward: 61.0 Training loss: 2.2578 Explore P: 0.3909
Episode: 415 Total reward: 49.0 Training loss: 1.3966 Explore P: 0.3890
Episode: 416 Total reward: 55.0 Training loss: 10.2836 Explore P: 0.3869
Episode: 417 Total reward: 121.0 Training loss: 2.2477 Explore P: 0.3824
Episode: 418 Total reward: 46.0 Training loss: 2.3118 Explore P: 0.3807
Episode: 419 Total reward: 70.0 Training loss: 27.3952 Explore P: 0.3781
Episode: 420 Total reward: 72.0 Training loss: 45.7570 Explore P: 0.3755
Episode: 421 Total reward: 41.0 Training loss: 59.1887 Explore P: 0.3740
Episode: 422 Total reward: 67.0 Training loss: 27.0998 Explore P: 0.3716
Episode: 423 Total reward: 46.0 Training loss: 43.1971 Explore P: 0.3699
Episode: 424 Total reward: 52.0 Training loss: 2.0718 Explore P: 0.3680
Episode: 425 Total reward: 92.0 Training loss: 96.7074 Explore P: 0.3647
Episode: 426 Total reward: 60.0 Training loss: 2.0684 Explore P: 0.3626
Episode: 427 Total reward: 106.0 Training loss: 54.1831 Explore P: 0.3589
Episode: 428 Total reward: 76.0 Training loss: 1.9612 Explore P: 0.3563
Episode: 429 Total reward: 42.0 Training loss: 1.6153 Explore P: 0.3548
Episode: 430 Total reward: 77.0 Training loss: 3.9801 Explore P: 0.3522
Episode: 431 Total reward: 123.0 Training loss: 2.0505 Explore P: 0.3480
Episode: 432 Total reward: 150.0 Training loss: 19.4217 Explore P: 0.3430
Episode: 433 Total reward: 57.0 Training loss: 1.5850 Explore P: 0.3411
Episode: 434 Total reward: 74.0 Training loss: 2.4292 Explore P: 0.3386
Episode: 435 Total reward: 97.0 Training loss: 23.5709 Explore P: 0.3354
Episode: 436 Total reward: 99.0 Training loss: 2.0727 Explore P: 0.3322
Episode: 437 Total reward: 101.0 Training loss: 22.3250 Explore P: 0.3290
Episode: 438 Total reward: 46.0 Training loss: 2.0320 Explore P: 0.3275
Episode: 439 Total reward: 51.0 Training loss: 4.8099 Explore P: 0.3259
Episode: 440 Total reward: 111.0 Training loss: 68.3524 Explore P: 0.3224
Episode: 441 Total reward: 167.0 Training loss: 2.3045 Explore P: 0.3173
Episode: 442 Total reward: 80.0 Training loss: 0.8798 Explore P: 0.3148
Episode: 443 Total reward: 170.0 Training loss: 48.6270 Explore P: 0.3097
Episode: 444 Total reward: 77.0 Training loss: 2.2555 Explore P: 0.3074
Episode: 445 Total reward: 84.0 Training loss: 3.0428 Explore P: 0.3049
Episode: 447 Total reward: 12.0 Training loss: 2.8022 Explore P: 0.2987
Episode: 448 Total reward: 66.0 Training loss: 120.3442 Explore P: 0.2968
Episode: 449 Total reward: 152.0 Training loss: 6.2880 Explore P: 0.2925
Episode: 450 Total reward: 141.0 Training loss: 2.8015 Explore P: 0.2885
Episode: 451 Total reward: 99.0 Training loss: 64.0921 Explore P: 0.2858
Episode: 452 Total reward: 79.0 Training loss: 1.5581 Explore P: 0.2836
Episode: 453 Total reward: 49.0 Training loss: 113.2557 Explore P: 0.2823
Episode: 455 Total reward: 106.0 Training loss: 210.4934 Explore P: 0.2741
Episode: 456 Total reward: 109.0 Training loss: 81.6662 Explore P: 0.2712
Episode: 457 Total reward: 56.0 Training loss: 3.2287 Explore P: 0.2697
Episode: 458 Total reward: 138.0 Training loss: 2.5795 Explore P: 0.2662
Episode: 459 Total reward: 93.0 Training loss: 3.4260 Explore P: 0.2638
Episode: 460 Total reward: 71.0 Training loss: 139.3341 Explore P: 0.2620
Episode: 461 Total reward: 106.0 Training loss: 2.6074 Explore P: 0.2594
Episode: 462 Total reward: 63.0 Training loss: 2.8252 Explore P: 0.2578
Episode: 463 Total reward: 71.0 Training loss: 25.8917 Explore P: 0.2560
Episode: 464 Total reward: 79.0 Training loss: 3.8067 Explore P: 0.2541
Episode: 465 Total reward: 86.0 Training loss: 1.6050 Explore P: 0.2520
Episode: 466 Total reward: 88.0 Training loss: 44.2827 Explore P: 0.2499
Episode: 467 Total reward: 72.0 Training loss: 0.7160 Explore P: 0.2482
Episode: 468 Total reward: 152.0 Training loss: 75.7239 Explore P: 0.2446
Episode: 469 Total reward: 122.0 Training loss: 7.4345 Explore P: 0.2417
Episode: 470 Total reward: 81.0 Training loss: 101.0922 Explore P: 0.2399
Episode: 471 Total reward: 38.0 Training loss: 1.6301 Explore P: 0.2390
Episode: 472 Total reward: 79.0 Training loss: 72.4920 Explore P: 0.2372
Episode: 473 Total reward: 190.0 Training loss: 1.3869 Explore P: 0.2329
Episode: 474 Total reward: 197.0 Training loss: 1.5386 Explore P: 0.2286
Episode: 476 Total reward: 42.0 Training loss: 0.8364 Explore P: 0.2233
Episode: 477 Total reward: 134.0 Training loss: 88.3979 Explore P: 0.2205
Episode: 478 Total reward: 128.0 Training loss: 94.1007 Explore P: 0.2178
Episode: 479 Total reward: 79.0 Training loss: 103.1366 Explore P: 0.2162
Episode: 480 Total reward: 169.0 Training loss: 58.8788 Explore P: 0.2127
Episode: 481 Total reward: 160.0 Training loss: 1.1934 Explore P: 0.2095
Episode: 482 Total reward: 81.0 Training loss: 2.9244 Explore P: 0.2079
Episode: 484 Total reward: 68.0 Training loss: 77.9688 Explore P: 0.2027
Episode: 486 Total reward: 17.0 Training loss: 0.6864 Explore P: 0.1985
Episode: 488 Total reward: 62.0 Training loss: 1.9978 Explore P: 0.1937
Episode: 489 Total reward: 178.0 Training loss: 228.6335 Explore P: 0.1904
Episode: 490 Total reward: 114.0 Training loss: 0.4453 Explore P: 0.1884
Episode: 491 Total reward: 127.0 Training loss: 1.6523 Explore P: 0.1861
Episode: 492 Total reward: 124.0 Training loss: 120.2207 Explore P: 0.1840
Episode: 493 Total reward: 184.0 Training loss: 0.5913 Explore P: 0.1808
Episode: 494 Total reward: 129.0 Training loss: 56.3829 Explore P: 0.1786
Episode: 495 Total reward: 95.0 Training loss: 1.9883 Explore P: 0.1770
Episode: 496 Total reward: 129.0 Training loss: 1.2513 Explore P: 0.1749
Episode: 497 Total reward: 176.0 Training loss: 1.0322 Explore P: 0.1720
Episode: 498 Total reward: 132.0 Training loss: 0.9320 Explore P: 0.1699
Episode: 499 Total reward: 146.0 Training loss: 289.4379 Explore P: 0.1675
Episode: 500 Total reward: 147.0 Training loss: 0.5124 Explore P: 0.1652
Episode: 501 Total reward: 166.0 Training loss: 0.9444 Explore P: 0.1627
Episode: 503 Total reward: 31.0 Training loss: 1.4756 Explore P: 0.1592
Episode: 504 Total reward: 129.0 Training loss: 0.6077 Explore P: 0.1573
Episode: 505 Total reward: 127.0 Training loss: 1.2508 Explore P: 0.1554
Episode: 506 Total reward: 123.0 Training loss: 0.8265 Explore P: 0.1537
Episode: 507 Total reward: 159.0 Training loss: 260.4604 Explore P: 0.1514
Episode: 508 Total reward: 136.0 Training loss: 0.9311 Explore P: 0.1495
Episode: 509 Total reward: 198.0 Training loss: 0.9262 Explore P: 0.1467
Episode: 511 Total reward: 40.0 Training loss: 2.3126 Explore P: 0.1435
Episode: 512 Total reward: 130.0 Training loss: 1.2985 Explore P: 0.1418
Episode: 514 Total reward: 25.0 Training loss: 1.1655 Explore P: 0.1388
Episode: 515 Total reward: 149.0 Training loss: 214.9246 Explore P: 0.1369
Episode: 516 Total reward: 200.0 Training loss: 0.8085 Explore P: 0.1344
Episode: 517 Total reward: 172.0 Training loss: 24.9451 Explore P: 0.1323
Episode: 519 Total reward: 52.0 Training loss: 61.7503 Explore P: 0.1293
Episode: 520 Total reward: 176.0 Training loss: 0.4361 Explore P: 0.1272
Episode: 521 Total reward: 160.0 Training loss: 1.8377 Explore P: 0.1253
Episode: 522 Total reward: 180.0 Training loss: 1.5684 Explore P: 0.1233
Episode: 523 Total reward: 174.0 Training loss: 58.6258 Explore P: 0.1213
Episode: 525 Total reward: 10.0 Training loss: 222.1836 Explore P: 0.1190
Episode: 527 Total reward: 32.0 Training loss: 0.4007 Explore P: 0.1165
Episode: 529 Total reward: 33.0 Training loss: 0.3054 Explore P: 0.1140
Episode: 530 Total reward: 185.0 Training loss: 0.7425 Explore P: 0.1121
Episode: 531 Total reward: 171.0 Training loss: 0.3441 Explore P: 0.1104
Episode: 532 Total reward: 199.0 Training loss: 0.2333 Explore P: 0.1084
Episode: 534 Total reward: 95.0 Training loss: 0.4929 Explore P: 0.1056
Episode: 536 Total reward: 8.0 Training loss: 0.5416 Explore P: 0.1036
Episode: 538 Total reward: 42.0 Training loss: 163.3946 Explore P: 0.1014
Episode: 539 Total reward: 180.0 Training loss: 0.2803 Explore P: 0.0997
Episode: 540 Total reward: 193.0 Training loss: 0.4929 Explore P: 0.0980
Episode: 542 Total reward: 36.0 Training loss: 0.7983 Explore P: 0.0960
Episode: 544 Total reward: 152.0 Training loss: 41.4165 Explore P: 0.0930
Episode: 546 Total reward: 30.0 Training loss: 0.7570 Explore P: 0.0911
Episode: 548 Total reward: 50.0 Training loss: 0.3215 Explore P: 0.0891
Episode: 550 Total reward: 79.0 Training loss: 0.5349 Explore P: 0.0869
Episode: 552 Total reward: 38.0 Training loss: 0.3635 Explore P: 0.0851
Episode: 553 Total reward: 131.0 Training loss: 0.3965 Explore P: 0.0841
Episode: 554 Total reward: 135.0 Training loss: 0.2453 Explore P: 0.0831
Episode: 555 Total reward: 111.0 Training loss: 0.9434 Explore P: 0.0823
Episode: 556 Total reward: 136.0 Training loss: 0.7058 Explore P: 0.0814
Episode: 557 Total reward: 106.0 Training loss: 0.4755 Explore P: 0.0806
Episode: 558 Total reward: 98.0 Training loss: 0.4107 Explore P: 0.0799
Episode: 559 Total reward: 102.0 Training loss: 62.3874 Explore P: 0.0792
Episode: 560 Total reward: 92.0 Training loss: 0.4026 Explore P: 0.0786
Episode: 561 Total reward: 86.0 Training loss: 0.3649 Explore P: 0.0780
Episode: 562 Total reward: 83.0 Training loss: 0.5843 Explore P: 0.0774
Episode: 563 Total reward: 100.0 Training loss: 0.1493 Explore P: 0.0768
Episode: 564 Total reward: 102.0 Training loss: 0.4021 Explore P: 0.0761
Episode: 565 Total reward: 60.0 Training loss: 0.3445 Explore P: 0.0757
Episode: 566 Total reward: 63.0 Training loss: 0.2461 Explore P: 0.0753
Episode: 567 Total reward: 59.0 Training loss: 0.2115 Explore P: 0.0749
Episode: 568 Total reward: 73.0 Training loss: 3.2738 Explore P: 0.0744
Episode: 569 Total reward: 70.0 Training loss: 0.4267 Explore P: 0.0740
Episode: 570 Total reward: 53.0 Training loss: 0.8779 Explore P: 0.0736
Episode: 571 Total reward: 88.0 Training loss: 187.5536 Explore P: 0.0731
Episode: 572 Total reward: 54.0 Training loss: 0.3208 Explore P: 0.0727
Episode: 573 Total reward: 87.0 Training loss: 0.2894 Explore P: 0.0722
Episode: 574 Total reward: 58.0 Training loss: 0.2578 Explore P: 0.0718
Episode: 575 Total reward: 85.0 Training loss: 0.3401 Explore P: 0.0713
Episode: 576 Total reward: 73.0 Training loss: 0.2245 Explore P: 0.0709
Episode: 577 Total reward: 114.0 Training loss: 0.3640 Explore P: 0.0702
Episode: 578 Total reward: 94.0 Training loss: 0.7954 Explore P: 0.0696
Episode: 579 Total reward: 114.0 Training loss: 0.2615 Explore P: 0.0689
Episode: 580 Total reward: 80.0 Training loss: 0.4812 Explore P: 0.0685
Episode: 581 Total reward: 125.0 Training loss: 0.8818 Explore P: 0.0677
Episode: 582 Total reward: 109.0 Training loss: 0.2953 Explore P: 0.0671
Episode: 583 Total reward: 98.0 Training loss: 0.4371 Explore P: 0.0665
Episode: 584 Total reward: 119.0 Training loss: 0.4685 Explore P: 0.0659
Episode: 585 Total reward: 96.0 Training loss: 0.3440 Explore P: 0.0653
Episode: 586 Total reward: 172.0 Training loss: 0.1414 Explore P: 0.0644
Episode: 587 Total reward: 104.0 Training loss: 0.3309 Explore P: 0.0638
Episode: 588 Total reward: 85.0 Training loss: 0.2262 Explore P: 0.0634
Episode: 590 Total reward: 104.0 Training loss: 0.3231 Explore P: 0.0618
Episode: 591 Total reward: 148.0 Training loss: 0.4431 Explore P: 0.0610
Episode: 592 Total reward: 135.0 Training loss: 0.1894 Explore P: 0.0603
Episode: 595 Total reward: 99.0 Training loss: 0.2376 Explore P: 0.0579
Episode: 597 Total reward: 172.0 Training loss: 0.4083 Explore P: 0.0561
Episode: 600 Total reward: 99.0 Training loss: 0.1152 Explore P: 0.0539
Episode: 603 Total reward: 99.0 Training loss: 0.3594 Explore P: 0.0518
Episode: 604 Total reward: 149.0 Training loss: 0.1398 Explore P: 0.0511
Episode: 607 Total reward: 99.0 Training loss: 0.3337 Explore P: 0.0491
Episode: 608 Total reward: 180.0 Training loss: 7.4786 Explore P: 0.0484
Episode: 611 Total reward: 28.0 Training loss: 0.1953 Explore P: 0.0468
Episode: 614 Total reward: 38.0 Training loss: 0.3152 Explore P: 0.0453
Episode: 617 Total reward: 50.0 Training loss: 0.4420 Explore P: 0.0437
Episode: 620 Total reward: 9.0 Training loss: 0.3347 Explore P: 0.0423
Episode: 623 Total reward: 99.0 Training loss: 0.1405 Explore P: 0.0408
Episode: 626 Total reward: 78.0 Training loss: 0.1488 Explore P: 0.0393
Episode: 628 Total reward: 198.0 Training loss: 0.9185 Explore P: 0.0382
Episode: 631 Total reward: 99.0 Training loss: 0.3505 Explore P: 0.0368
Episode: 633 Total reward: 142.0 Training loss: 0.1654 Explore P: 0.0359
Episode: 635 Total reward: 134.0 Training loss: 0.3178 Explore P: 0.0351
Episode: 637 Total reward: 104.0 Training loss: 0.3331 Explore P: 0.0343
Episode: 639 Total reward: 73.0 Training loss: 66.8497 Explore P: 0.0337
Episode: 641 Total reward: 35.0 Training loss: 0.1411 Explore P: 0.0331
Episode: 643 Total reward: 83.0 Training loss: 0.2136 Explore P: 0.0325
Episode: 645 Total reward: 62.0 Training loss: 0.2303 Explore P: 0.0319
Episode: 647 Total reward: 28.0 Training loss: 0.2552 Explore P: 0.0314
Episode: 649 Total reward: 4.0 Training loss: 0.1967 Explore P: 0.0310
Episode: 650 Total reward: 194.0 Training loss: 0.1424 Explore P: 0.0306
Episode: 651 Total reward: 170.0 Training loss: 0.1509 Explore P: 0.0302
Episode: 652 Total reward: 150.0 Training loss: 0.2699 Explore P: 0.0299
Episode: 653 Total reward: 161.0 Training loss: 0.2821 Explore P: 0.0296
Episode: 654 Total reward: 148.0 Training loss: 0.4859 Explore P: 0.0293
Episode: 655 Total reward: 142.0 Training loss: 0.1541 Explore P: 0.0290
Episode: 656 Total reward: 140.0 Training loss: 0.0963 Explore P: 0.0288
Episode: 657 Total reward: 144.0 Training loss: 0.3165 Explore P: 0.0285
Episode: 658 Total reward: 160.0 Training loss: 0.2059 Explore P: 0.0282
Episode: 659 Total reward: 127.0 Training loss: 0.0918 Explore P: 0.0280
Episode: 660 Total reward: 124.0 Training loss: 431.4700 Explore P: 0.0278
Episode: 661 Total reward: 127.0 Training loss: 0.2660 Explore P: 0.0275
Episode: 662 Total reward: 133.0 Training loss: 0.4122 Explore P: 0.0273
Episode: 663 Total reward: 119.0 Training loss: 0.2070 Explore P: 0.0271
Episode: 664 Total reward: 114.0 Training loss: 0.3453 Explore P: 0.0269
Episode: 665 Total reward: 130.0 Training loss: 0.3865 Explore P: 0.0267
Episode: 666 Total reward: 125.0 Training loss: 0.2518 Explore P: 0.0265
Episode: 667 Total reward: 138.0 Training loss: 0.1668 Explore P: 0.0263
Episode: 669 Total reward: 42.0 Training loss: 0.3241 Explore P: 0.0259
Episode: 671 Total reward: 105.0 Training loss: 0.1787 Explore P: 0.0254
Episode: 674 Total reward: 99.0 Training loss: 0.2393 Explore P: 0.0246
Episode: 677 Total reward: 99.0 Training loss: 0.2190 Explore P: 0.0239
Episode: 680 Total reward: 99.0 Training loss: 2.5996 Explore P: 0.0232
Episode: 683 Total reward: 99.0 Training loss: 0.3376 Explore P: 0.0226
Episode: 686 Total reward: 99.0 Training loss: 0.5884 Explore P: 0.0220
Episode: 689 Total reward: 99.0 Training loss: 0.1356 Explore P: 0.0214
Episode: 692 Total reward: 99.0 Training loss: 0.0920 Explore P: 0.0208
Episode: 695 Total reward: 99.0 Training loss: 0.1880 Explore P: 0.0203
Episode: 698 Total reward: 99.0 Training loss: 0.0951 Explore P: 0.0198
Episode: 701 Total reward: 99.0 Training loss: 0.1050 Explore P: 0.0193
Episode: 704 Total reward: 99.0 Training loss: 0.1234 Explore P: 0.0189
Episode: 707 Total reward: 99.0 Training loss: 0.1407 Explore P: 0.0185
Episode: 709 Total reward: 160.0 Training loss: 0.0913 Explore P: 0.0182
Episode: 712 Total reward: 99.0 Training loss: 0.1815 Explore P: 0.0178
Episode: 715 Total reward: 99.0 Training loss: 0.1191 Explore P: 0.0174
Episode: 718 Total reward: 99.0 Training loss: 0.1073 Explore P: 0.0170
Episode: 721 Total reward: 99.0 Training loss: 0.1133 Explore P: 0.0167
Episode: 724 Total reward: 99.0 Training loss: 0.0898 Explore P: 0.0164
Episode: 727 Total reward: 99.0 Training loss: 0.1217 Explore P: 0.0160
Episode: 730 Total reward: 99.0 Training loss: 0.2150 Explore P: 0.0158
Episode: 733 Total reward: 99.0 Training loss: 0.0678 Explore P: 0.0155
Episode: 736 Total reward: 99.0 Training loss: 0.0872 Explore P: 0.0152
Episode: 739 Total reward: 99.0 Training loss: 0.1330 Explore P: 0.0150
Episode: 742 Total reward: 99.0 Training loss: 0.1116 Explore P: 0.0147
Episode: 745 Total reward: 99.0 Training loss: 0.1611 Explore P: 0.0145
Episode: 748 Total reward: 99.0 Training loss: 0.1307 Explore P: 0.0143
Episode: 751 Total reward: 99.0 Training loss: 0.0875 Explore P: 0.0141
Episode: 754 Total reward: 99.0 Training loss: 0.1344 Explore P: 0.0139
Episode: 757 Total reward: 99.0 Training loss: 0.0911 Explore P: 0.0137
Episode: 760 Total reward: 99.0 Training loss: 0.1224 Explore P: 0.0135
Episode: 763 Total reward: 99.0 Training loss: 0.0572 Explore P: 0.0133
Episode: 766 Total reward: 99.0 Training loss: 0.0757 Explore P: 0.0132
Episode: 769 Total reward: 99.0 Training loss: 0.0381 Explore P: 0.0130
Episode: 772 Total reward: 99.0 Training loss: 0.1698 Explore P: 0.0129
Episode: 775 Total reward: 99.0 Training loss: 0.0365 Explore P: 0.0127
Episode: 778 Total reward: 99.0 Training loss: 0.1805 Explore P: 0.0126
Episode: 781 Total reward: 99.0 Training loss: 0.1017 Explore P: 0.0125
Episode: 784 Total reward: 99.0 Training loss: 0.1112 Explore P: 0.0123
Episode: 787 Total reward: 99.0 Training loss: 0.0930 Explore P: 0.0122
Episode: 790 Total reward: 99.0 Training loss: 0.0693 Explore P: 0.0121
Episode: 793 Total reward: 99.0 Training loss: 0.0431 Explore P: 0.0120
Episode: 796 Total reward: 99.0 Training loss: 0.1168 Explore P: 0.0119
Episode: 799 Total reward: 99.0 Training loss: 0.1071 Explore P: 0.0118
Episode: 802 Total reward: 99.0 Training loss: 0.1360 Explore P: 0.0117
Episode: 805 Total reward: 99.0 Training loss: 0.0872 Explore P: 0.0117
Episode: 808 Total reward: 99.0 Training loss: 0.1197 Explore P: 0.0116
Episode: 811 Total reward: 99.0 Training loss: 0.0848 Explore P: 0.0115
Episode: 814 Total reward: 99.0 Training loss: 0.0515 Explore P: 0.0114
Episode: 817 Total reward: 99.0 Training loss: 0.1590 Explore P: 0.0114
Episode: 820 Total reward: 99.0 Training loss: 0.2080 Explore P: 0.0113
Episode: 823 Total reward: 99.0 Training loss: 0.1532 Explore P: 0.0112
Episode: 826 Total reward: 99.0 Training loss: 0.0622 Explore P: 0.0112
Episode: 829 Total reward: 99.0 Training loss: 0.0553 Explore P: 0.0111
Episode: 832 Total reward: 99.0 Training loss: 0.0746 Explore P: 0.0111
Episode: 835 Total reward: 99.0 Training loss: 0.1045 Explore P: 0.0110
Episode: 838 Total reward: 99.0 Training loss: 0.0929 Explore P: 0.0110
Episode: 841 Total reward: 99.0 Training loss: 0.1053 Explore P: 0.0109
Episode: 844 Total reward: 99.0 Training loss: 0.0877 Explore P: 0.0109
Episode: 847 Total reward: 99.0 Training loss: 0.0783 Explore P: 0.0108
Episode: 850 Total reward: 99.0 Training loss: 0.0724 Explore P: 0.0108
Episode: 853 Total reward: 99.0 Training loss: 0.1745 Explore P: 0.0107
Episode: 856 Total reward: 99.0 Training loss: 0.0334 Explore P: 0.0107
Episode: 859 Total reward: 99.0 Training loss: 0.0205 Explore P: 0.0107
Episode: 862 Total reward: 99.0 Training loss: 0.0674 Explore P: 0.0106
Episode: 865 Total reward: 99.0 Training loss: 0.1149 Explore P: 0.0106
Episode: 868 Total reward: 99.0 Training loss: 191.0773 Explore P: 0.0106
Episode: 871 Total reward: 99.0 Training loss: 0.1013 Explore P: 0.0106
Episode: 873 Total reward: 156.0 Training loss: 0.2140 Explore P: 0.0105
Episode: 874 Total reward: 185.0 Training loss: 0.1837 Explore P: 0.0105
Episode: 876 Total reward: 45.0 Training loss: 0.1855 Explore P: 0.0105
Episode: 877 Total reward: 55.0 Training loss: 0.0789 Explore P: 0.0105
Episode: 880 Total reward: 99.0 Training loss: 0.0890 Explore P: 0.0105
Episode: 882 Total reward: 185.0 Training loss: 0.0788 Explore P: 0.0105
Episode: 885 Total reward: 99.0 Training loss: 0.1397 Explore P: 0.0104
Episode: 888 Total reward: 99.0 Training loss: 0.0400 Explore P: 0.0104
Episode: 891 Total reward: 99.0 Training loss: 0.0962 Explore P: 0.0104
Episode: 894 Total reward: 99.0 Training loss: 0.1356 Explore P: 0.0104
Episode: 897 Total reward: 99.0 Training loss: 0.2037 Explore P: 0.0104
Episode: 900 Total reward: 99.0 Training loss: 0.0486 Explore P: 0.0103
Episode: 903 Total reward: 99.0 Training loss: 0.2492 Explore P: 0.0103
Episode: 906 Total reward: 99.0 Training loss: 0.1467 Explore P: 0.0103
Episode: 909 Total reward: 99.0 Training loss: 0.2217 Explore P: 0.0103
Episode: 912 Total reward: 99.0 Training loss: 0.1772 Explore P: 0.0103
Episode: 915 Total reward: 99.0 Training loss: 0.0898 Explore P: 0.0103
Episode: 918 Total reward: 99.0 Training loss: 0.0552 Explore P: 0.0103
Episode: 921 Total reward: 99.0 Training loss: 0.1267 Explore P: 0.0102
Episode: 924 Total reward: 99.0 Training loss: 0.3037 Explore P: 0.0102
Episode: 927 Total reward: 99.0 Training loss: 0.1654 Explore P: 0.0102
Episode: 930 Total reward: 99.0 Training loss: 0.1975 Explore P: 0.0102
Episode: 933 Total reward: 99.0 Training loss: 0.2122 Explore P: 0.0102
Episode: 936 Total reward: 99.0 Training loss: 0.0754 Explore P: 0.0102
Episode: 939 Total reward: 99.0 Training loss: 0.1481 Explore P: 0.0102
Episode: 942 Total reward: 99.0 Training loss: 0.0895 Explore P: 0.0102
Episode: 945 Total reward: 99.0 Training loss: 0.0690 Explore P: 0.0102
Episode: 948 Total reward: 99.0 Training loss: 0.0942 Explore P: 0.0102
Episode: 951 Total reward: 99.0 Training loss: 0.0567 Explore P: 0.0101
Episode: 954 Total reward: 99.0 Training loss: 0.0665 Explore P: 0.0101
Episode: 957 Total reward: 99.0 Training loss: 0.0645 Explore P: 0.0101
Episode: 960 Total reward: 99.0 Training loss: 224.4461 Explore P: 0.0101
Episode: 963 Total reward: 99.0 Training loss: 0.0508 Explore P: 0.0101
Episode: 966 Total reward: 99.0 Training loss: 0.0792 Explore P: 0.0101
Episode: 969 Total reward: 99.0 Training loss: 0.0754 Explore P: 0.0101
Episode: 972 Total reward: 99.0 Training loss: 0.0655 Explore P: 0.0101
Episode: 975 Total reward: 99.0 Training loss: 0.0686 Explore P: 0.0101
Episode: 978 Total reward: 99.0 Training loss: 0.0361 Explore P: 0.0101
Episode: 981 Total reward: 99.0 Training loss: 0.1777 Explore P: 0.0101
Episode: 984 Total reward: 99.0 Training loss: 0.0633 Explore P: 0.0101
Episode: 987 Total reward: 99.0 Training loss: 0.0559 Explore P: 0.0101
Episode: 990 Total reward: 99.0 Training loss: 0.0543 Explore P: 0.0101
Episode: 993 Total reward: 99.0 Training loss: 0.0833 Explore P: 0.0101
Episode: 996 Total reward: 99.0 Training loss: 0.1037 Explore P: 0.0101
Episode: 997 Total reward: 45.0 Training loss: 0.0619 Explore P: 0.0101
```
## Visualizing training
Below we plot the total rewards for each episode. The rolling average is plotted in blue.
```
%matplotlib inline
import matplotlib.pyplot as plt
def running_mean(x, N):
cumsum = np.cumsum(np.insert(x, 0, 0))
return (cumsum[N:] - cumsum[:-N]) / N
eps, rews = np.array(rewards_list).T
smoothed_rews = running_mean(rews, 10)
plt.plot(eps[-len(smoothed_rews):], smoothed_rews)
plt.plot(eps, rews, color='grey', alpha=0.3)
plt.xlabel('Episode')
plt.ylabel('Total Reward')
Text(0,0.5,'Total Reward')
```

## Playing Atari Games
So, Cart-Pole is a pretty simple game. However, the same model can be used to train an agent to play something much more complicated like Pong or Space Invaders. Instead of a state like we're using here though, you'd want to use convolutional layers to get the state from the screen images.

I'll leave it as a challenge for you to use deep Q-learning to train an agent to play Atari games. Here's the original paper which will get you started: http://www.davidqiu.com:8888/research/nature14236.pdf.
| github_jupyter |
## Week 2 Optional Thoery Discussion
The following problems are for those of you looking to challenge yourself beyond the required problem sets and programming questions. Most of these have been given in Stanford's CS161 course, Design and Analysis of Algorithms, at some point. They are completely optional and will not be graded. While they vary in level, many are pretty challenging, and we strongly encourage you to discuss ideas and approaches with your fellow students on the "Theory Problems" discussion form.
1. You are given as input an unsorted array of n distinct numbers, where n is a power of 2. Give an algorithm that identifies the second-largest number in the array, and that uses at most $n + log_2 n − 2$ comparisons.
2. You are a given a unimodal array of n distinct elements, meaning that its entries are in increasing order up until its maximum element, after which its elements are in decreasing order. Give an algorithm to compute the maximum element that runs in O(log n) time.
3. You are given a sorted (from smallest to largest) array A of n distinct integers which can be positive, negative, or zero. You want to decide whether or not there is an index i such that ```A[i] = i```. Design the fastest algorithm that you can for solving this problem.
4. You are given an n by n grid of distinct numbers. A number is a local minimum if it is smaller than all of its neighbors. (A neighbor of a number is one immediately above, below, to the left, or the right. Most numbers have four neighbors; numbers on the side have three; the four corners have two.) Use the divide-and-conquer algorithm design paradigm to compute a local minimum with only O(n) comparisons between pairs of numbers. (Note: since there are $n^2$ numbers in the input, you cannot afford to look at all of them. Hint: Think about what types of recurrences would give you the desired upper bound.)
## Week 4 Optional Problems
The following problems are for those of you looking to challenge yourself beyond the required problem sets and programming questions. Most of these have been given in Stanford's CS161 course, Design and Analysis of Algorithms, at some point. They are completely optional and will not be graded. While they vary in level, many are pretty challenging, and we strongly encourage you to discuss ideas and approaches with your fellow students on the "Theory Problems" discussion form.
1. Prove that the worst-case expected running time of every randomized comparison-based sorting algorithm is $\Omega (nlogn)$. (Here the worst-case is over inputs, and the expectation is over the random coin flips made by the algorithm.)
2. Suppose we modify the deterministic linear-time selection algorithm by grouping the elements into groups of 7, rather than groups of 5. (Use the "median-of-medians" as the pivot, as before.) Does the algorithm still run in O(n)O(n) time? What if we use groups of 3?
3. Given an array of n distinct (but unsorted) elements $x_1,x_2,\ldots,x_n$ with positive weights $w_1,w_2,\ldots,w_n$ such that $\sum_{i=1}^n w_i = W$, a weighted median is an element $x_k$ for which the total weight of all elements with value less than $x_k$ (i.e., $\sum_{x_i<x_k} w_i $) is at most $W/2$ and also the total weight of elements with value larger than $x_k$ (i.e., $\sum_{x_i>x_k} w_i$) is at most $W/2$. Observe that there are at most two weighted medians. Show how to compute all weighted medians in O(n) worst-case time.
4. We showed in an optional video lecture that every undirected graph has only polynomially (in the number nn of vertices) different minimum cuts. Is this also true for directed graphs? Prove it or give a counterexample.
5. For a parameter $\alpha \ge 1$, an $\alpha$-minimum cut is one for which the number of crossing edges is at most \alphaα times that of a minimum cut. How many $\alpha$-minimum cuts can an undirected graph have, as a function of $\alpha$ and the number nn of vertices? Prove the best upper bound that you can.
| github_jupyter |
# Multi-Label Classification Tutorial
This tutorial shows how to use Tribuo's MultiLabel package to perform [multi-label classification](https://en.wikipedia.org/wiki/Multi-label_classification) tasks. Multi-label classification is the task of assigning a *set* of labels to a given example from a specific label domain, as opposed to multi-class classification which is assigning a *single* label to a given example.
Tribuo provides linear model and factorization machine algorithms for native multi-label prediction, along with ensemble methods that either predict each label independently or as part of a [classifier chain](http://www.cs.waikato.ac.nz/~ml/publications/2009/chains.pdf), using any of Tribuo's classification algorithms as the base learners. Both the linear models, factorization machines and the `IndependentMultiLabelTrainer` use the *Binary Relevance* approach to multi-label prediction, where each label is predicted independently. The `ClassifierChainTrainer` and `CCEnsembleTrainer` use classifier chains which incorporate label structure into the prediction. In this tutorial we'll cover loading in multi-label data, performing predictions using several binary relevance based classifiers along with some classifier chains, and finally we'll evaluate the multi-label models using Tribuo's multi-label evaluation package.
## Setup
First you'll need a copy of the multi-label yeast dataset (we'll download these from the [LibSVM](https://www.csie.ntu.edu.tw/~cjlin/libsvm/) dataset repo):
```
wget https://www.csie.ntu.edu.tw/~cjlin/libsvmtools/datasets/multilabel/yeast_train.svm.bz2
wget https://www.csie.ntu.edu.tw/~cjlin/libsvmtools/datasets/multilabel/yeast_test.svm.bz2
```
Then you should extract it using your preferred method. On macOS and Linux you can use `bunzip2`, and on Windows there are several packages which can extract bz2 files (e.g., [7-zip](https://www.7-zip.org/)).
This dataset has 14 labels which represent different functional groups and the task is to predict the functional groups a gene belongs in based on micro-array expression measurements. Fortunately we don't need a PhD in Genetics to use this dataset as a benchmark, though obviously domain knowledge would be critical if we wanted to actually deploy any model based on this data.
Now we'll load in the necessary jars and import some packages from the JDK and Tribuo.
```
%jars ./tribuo-multilabel-sgd-4.2.0-SNAPSHOT-jar-with-dependencies.jar
%jars ./tribuo-classification-experiments-4.2.0-SNAPSHOT-jar-with-dependencies.jar
import java.nio.file.Paths;
import org.tribuo.*;
import org.tribuo.classification.Label;
import org.tribuo.classification.dtree.CARTClassificationTrainer;
import org.tribuo.classification.dtree.impurity.*;
import org.tribuo.datasource.*;
import org.tribuo.math.optimisers.*;
import org.tribuo.multilabel.*;
import org.tribuo.multilabel.baseline.*;
import org.tribuo.multilabel.ensemble.*;
import org.tribuo.multilabel.evaluation.*;
import org.tribuo.multilabel.sgd.linear.*;
import org.tribuo.multilabel.sgd.objectives.*;
import org.tribuo.util.Util;
```
## Loading the data
There are two main forms for multi-label data in columnar representations. Either the dataset stores the labels in a single column using some delimiter (e.g., "first_label,third_label"), resulting in a sparse representation of the labels, or each label is stored in it's own column with a flag representing if that label is present (e.g., "TRUE" or "1"), resulting in a dense representation of the labels. Tribuo can load both formats, though currently the `MultiLabelFactory` only supports comma separated labels when parsing inputs directly from a `String`. When processing multi-label values through a `RowProcessor` then the factory receives a `List<String>` and processes each separate label appropriately.
The yeast dataset we downloaded is stored in libsvm format which uses a sparse representation of the labels, so we'll use Tribuo's `LibSVMDataSource` to load it in, and process the outputs through a `MultiLabelFactory`.
```
var factory = new MultiLabelFactory();
var trainSource = new LibSVMDataSource<>(Paths.get(".","yeast_train.svm"),factory);
var testSource = new LibSVMDataSource<>(Paths.get(".","yeast_test.svm"),factory,trainSource.isZeroIndexed(),trainSource.getMaxFeatureID());
var train = new MutableDataset<>(trainSource);
var test = new MutableDataset<>(testSource);
System.out.println(String.format("Training data size = %d, number of features = %d, number of classes = %d",train.size(),train.getFeatureMap().size(),train.getOutputInfo().size()));
System.out.println(String.format("Testing data size = %d, number of features = %d, number of classes = %d",test.size(),test.getFeatureMap().size(),test.getOutputInfo().size()));
```
In Tribuo we represent a multi-label task using the `org.tribuo.multilabel.MultiLabel` output, which internally uses a set of `org.tribuo.classification.Label` objects to store the individual labels. This means that unlike most Tribuo prediction type packages, `tribuo-multilabel-core` depends on another output core package `tribuo-classification-core`. `MultiLabel` is a sparse representation of the labels, only the `Label`s which are active are stored in the `MultiLabel` object.
We can inspect the first output from the training dataset to see this:
```
System.out.println("First output = " + train.getExample(0).getOutput());
System.out.println("Second output = " + train.getExample(1).getOutput());
```
This first example is tagged with labels 2 & 3, and the second one is tagged with 6, 7, 11 and 12. Unfortunately the LibSVM format we loaded in uses numbers rather than names for the labels, but if there are more descriptive names present when the data is loaded in then those would be used as the label names.
## The task of multi-label prediction
Multi-label problems can be approached in several different ways. A common approach is to treat each label as an independent function of the input features, this leads to the *binary relevance* approach where each label is independent from each other, and multi-label classification can be thought of as a set of standard binary classification problems. This approach scales well, but if there is underlying structure in the label space (e.g., the label "human" implies the label "animal", but the label "animal" does not imply "human", so they are not independent), then this approach ignores useful information from the training data and may underperform more complicated approaches. Another popular way to convert a multi-label problem into a standard classification problem is via a *label powerset*, where each unique combination of the individual labels is treated as a single label in a large multi-class classification problem. While this allows the learning algorithm to fully capture any interactions between the labels, the label powerset is exponential in the number of labels, which rapidly makes this approach intractable as the number of labels increases though it can be useful in small label spaces. Tribuo currently focuses on binary relevance and other approaches which don't require exponential computation, though we're happy to discuss incorporating label powerset methods if people have need for them.
## Training Binary Relevance models
Now we'll train a few different binary relevance models (i.e., independent predictions of each label). First we'll use Tribuo's multi-label `LinearSGDModel` which natively makes multi-label predictions, then we'll wrap a binary classification decision tree into a multi-label predictor using `IndependentMultiLabelTrainer` and `IndependentMultiLabelModel`. Note: Tribuo has three classes called `LinearSGDModel`, one each for `Label`, `MultiLabel`, and `Regressor`, so the `LinearSGDModel` used in this tutorial is `org.tribuo.multilabel.sgd.linear.LinearSGDModel`, and the one used in the *multi-class* classification tutorials is `org.tribuo.classification.sgd.linear.LinearSGDModel`.
Tribuo's multi-label SGD package supports two different objective functions, Binary Cross-Entropy and Hinge loss. The BCE loss produces probabilitistic outputs thresholded at 0.5, whereas the hinge loss produces scores thresholded at 0. As Tribuo usually produces scores for each possible label, these thresholds are used to determine when a particular label is present in the `MultiLabel` object representing the predicted label set. As you may have seen in other tutorials, Tribuo uses stochastic gradient descent to fit it's linear models, so we'll define a gradient optimizer along with the loss function.
```
var linTrainer = new LinearSGDTrainer(new BinaryCrossEntropy(),new AdaGrad(0.1,0.1),5,1000,1,Trainer.DEFAULT_SEED);
```
We train the model the same way we train the rest of Tribuo's models.
```
var linStartTime = System.currentTimeMillis();
var linModel = linTrainer.train(train);
var linEndTime = System.currentTimeMillis();
System.out.println();
System.out.println("Linear model training took " + Util.formatDuration(linStartTime,linEndTime));
```
Tribuo doesn't have a native implementation of a multi-label decision tree, but it does have a multi-class decision tree, which we can convert into a multi-label predictor using `IndependentMultiLabelTrainer`. Now let's train a model using a decision tree to predict each label independently. First we define the binary classification trainer, then we'll use `IndependentMultiLabelTrainer` to wrap that `Trainer<Label>` and convert it into a `Trainer<MultiLabel>`, before training as usual.
```
Trainer<Label> treeTrainer = new CARTClassificationTrainer(6,10,0.0f,1.0f,false,new Entropy(),1L);
Trainer<MultiLabel> dtTrainer = new IndependentMultiLabelTrainer(treeTrainer);
var dtStartTime = System.currentTimeMillis();
var dtModel = dtTrainer.train(train);
var dtEndTime = System.currentTimeMillis();
System.out.println();
System.out.println("Tree model training took " + Util.formatDuration(dtStartTime,dtEndTime));
```
We've now got two different models, so let's measure their performance.
## Evaluating multi-class problems
Multi-label problems have many evaluation options available, as many standard classification evaluation measures like accuracy, precision, recall and F1 can be applied at the label level, and there are also many set level metric such as the [Jaccard Index](https://en.wikipedia.org/wiki/Jaccard_index) which can be used to compare the predicted label set against the ground truth one. In Tribuo we have access to most of the metrics available for multi-class classification problems, and v4.2 began adding set level metrics as well. If there are useful metrics that aren't implemented in Tribuo raise an issue on Tribuo's [Github page](https://github.com/oracle/tribuo).
If you want to use the predicted scores for each of the labels separately (e.g., to analyse the model's performance) then as usual the `Map<String,MultiLabel>` available from `Prediction.getOutputScores()` has the full distribution. This map behaves slightly counter-intuitively, as each value is a `MultiLabel` object containing a single `Label`, and the key is the output of `Label.toString()`. This allows the labels to be inspected individually, but it is a little uncomfortable if you're used to working with a multi-label specific API. However it maintains conformity across all of Tribuo's different prediction APIs, both for predictions and evaluations, which makes it easier to incorporate lots of ML models into a larger system.
We use the same evaluation paradigm as other Tribuo prediction tasks, first we construct an `Evaluator` and then feed it a model and some test data to produce an `Evaluation` which contains the appropriate performance metrics.
```
var eval = new MultiLabelEvaluator();
```
First we'll look at the linear model:
```
var linTStartTime = System.currentTimeMillis();
var linEval = eval.evaluate(linModel,test);
var linTEndTime = System.currentTimeMillis();
System.out.println();
System.out.println("Linear model evaluation took " + Util.formatDuration(linTStartTime,linTEndTime));
System.out.println(linEval);
```
Next, the decision tree:
```
var dtTStartTime = System.currentTimeMillis();
var dtEval = eval.evaluate(dtModel,test);
var dtTEndTime = System.currentTimeMillis();
System.out.println();
System.out.println("Tree model evaluation took " + Util.formatDuration(dtTStartTime,dtTEndTime));
System.out.println(dtEval);
```
We can see the native multi-label linear model outperformed the wrapped decision tree in terms of Jaccard Score, though the picture is more mixed in the other metrics, and the linear model is ignoring some of the labels.
Unfortunately some of the metrics we might like to examine for regular multi-class classification aren't as easy to use in the multi-label case. For example, a multi-class confusion matrix has no direct analogue in the multi-label case, as there could be an arbitrary number of labels predicted for each output, meaning there is no notion of a label being mispredicted as another label. This means a multi-label confusion matrix is best presented as a series of binary confusion matrices, one per label. This tends to take up a lot of space, so we'll skip inspecting them in this tutorial, though they are accessible on the `MultiLabelEvaluation` object.
Now let's look at a more complicated multi-label classification approach, using *Classifier Chains*.
## Training Classifier Chains
A [classifier chain](http://www.cs.waikato.ac.nz/~ml/publications/2009/chains.pdf) is similar to a binary relevance model, except there is a sequential order to the predictions (forming a chain), and each member of the chain receives extra features in the form of the predictions of earlier members of the chain. This means that if the chain is correctly ordered according to the causal structure of the labels (which is tricky to do) then it can start with the most independent label first, and then predict each label in sequence so it can use the earlier predictions to improve predictions for each subsequent label (e.g., we could predict if the example is an "animal" first, and then when we come to predict if it's a "human" we know that humans are animals making the prediction task easier).
In practice we don't usually know the correct ordering of the labels as the causal structure is unknown, and if we supply the incorrect structure then we can reduce performance back to the level of the binary relevance models. Fortunately in Machine Learning we have a trick we can use when we need to deal with uncertain data, which is to randomize it many times, and take an average. So we could take many different classifier chains each with an random label order, and then each chain votes on the labels that should be predicted. This improves statistical performance over a single chain with a random order, and over a single chain with a poorly chosen order, though it's unlikely to beat a single classifier chain with the correct label ordering (if such an ordering exists). Unfortunately the classifier chain ensemble is more expensive computationally than the single chain, which is already relatively expensive compared to a single classifier like `LinearSGDModel`, but the chains can be straightforwardly parallelized (and we'll add support for this to a future version of Tribuo).
We're going to use a single classifier chain with a random order, and then an ensemble of 20 classifier chains each using random orders to see how they perform.
```
var ccTrainer = new ClassifierChainTrainer(treeTrainer,1L);
var ccEnsembleTrainer = new CCEnsembleTrainer(treeTrainer,20,1L);
```
First we'll train and evaluate the single chain:
```
// train the model
var ccStartTime = System.currentTimeMillis();
var ccModel = ccTrainer.train(train);
var ccEndTime = System.currentTimeMillis();
System.out.println();
System.out.println("Classifier Chain model training took " + Util.formatDuration(ccStartTime,ccEndTime));
// evaluate the model
var ccTStartTime = System.currentTimeMillis();
var ccEval = eval.evaluate(ccModel,test);
var ccTEndTime = System.currentTimeMillis();
System.out.println("Classifier Chain model evaluation took " + Util.formatDuration(ccTStartTime,ccTEndTime));
System.out.println(ccEval);
```
We can see the classifier chain improved over the binary relevance model when using trees as the base learner, and took roughly the same amount of time to train and evaluate. It's still not quite up to the linear model, but let's try the chain ensemble and see how it does.
Now we'll train and evaluate the ensemble:
```
// train the model
var ccEnsembleStartTime = System.currentTimeMillis();
var ccEnsembleModel = ccEnsembleTrainer.train(train);
var ccEnsembleEndTime = System.currentTimeMillis();
System.out.println();
System.out.println("Classifier Chain Ensemble model training took " + Util.formatDuration(ccEnsembleStartTime,ccEnsembleEndTime));
// evaluate the model
var ccETStartTime = System.currentTimeMillis();
var ccEnsembleEval = eval.evaluate(ccEnsembleModel,test);
var ccETEndTime = System.currentTimeMillis();
System.out.println("Classifier Chain Ensemble model evaluation took " + Util.formatDuration(ccETStartTime,ccETEndTime));
System.out.println(ccEnsembleEval);
```
As expected the classifier chain ensemble outperformed the binary relevance model and the single classifier chain when using trees as the base learner, at the cost of the greatest runtime. It did this by significantly decreasing the number of false positives, at the cost of a small increase in false negatives. We didn't quite beat the performance of the linear model in terms of Jaccard score, but in general classifier chains are a powerful multi-label approach, and we could always use a the linear model as a base learner (and if you do, then you do improve the Jaccard score above 0.497). We leave the implementation of that as an exercise for the reader.
## Conclusion
We looked at Tribuo's multi-label classification package, trying out several different models using different approaches to the multi-label problem, namely binary relevance models and classifier chains. We're interested in expanding Tribuo's support for multi-label problems, so if there are algorithms or metrics Tribuo is missing head over to our [Github page](https://github.com/oracle/tribuo) and contributions are always welcome.
| github_jupyter |
# Copying an ArcGIS StoryMap item to another organization
## Introduction
Esri provides two models for telling stories with maps: The [Classic Story Map](https://storymaps-classic.arcgis.com/en/) and the newer [ArcGIS StoryMap](https://www.esri.com/en-us/arcgis/products/arcgis-storymaps/overview). Each offers the infrastructure to utilize ArcGIS Platform items such as [`Web Maps`](https://developers.arcgis.com/documentation/core-concepts/web-maps/) combined with supporting resources like images, text and videos for impactful storytelling. To answer your next question, please see [What's the Difference?](https://storymaps.arcgis.com/stories/6d3aff3f321f4f14b2f4ee29873c891b).
The platform stores each model differently, which leads to this sample document. The ArcGIS API for Python [`clone_items()`](https://developers.arcgis.com/python/api-reference/arcgis.gis.toc.html#arcgis.gis.ContentManager.clone_items) function equips you with all you need to effectively transfer many [Item](https://developers.arcgis.com/python/api-reference/arcgis.gis.toc.html#item) types between organizations, regardless of the organization's deployment. The `ArcGIS StoryMap` item is an exception to that currently. While there are plans to update `clone_items()` to handle the modern architecture behind `ArcGIS StoryMaps`, the sample below details a procedure to use immediately to transfer these items between organizations.
Let's proceed.
## Import Libraries
```
import os
import uuid
import json
import shutil
import tempfile
from arcgis.gis import GIS
from arcgis import __version__
```
Assign a variable to store appropriate version values to differentiate beween each story model.
```
_version = [int(i) for i in __version__.split('.')]
```
## Define function to export all the Story Map's resources to a zip file
Given its novel architecture and resource storage, we'll define a function to return the supporting resources for the ArcGIS StoryMap model.
```
def export_resources(item, save_path=None, file_name=None):
"""Export's the data's resources as a zip file"""
url = \
f'{item._gis._portal.resturl}content/users/{item._user_id}/items/{item.itemid}/resources/export'
if save_path is None:
save_path = tempfile.gettempdir()
if file_name is None:
file_name = f"{uuid.uuid4().hex[:6]}.zip"
params = {'f' : 'zip'}
con = item._gis._portal.con
resources = con.get(url, params=params,
out_folder=save_path,
file_name=file_name,
try_json=False)
return resources
```
## Connect to the source and destination GIS organizations
```
gis = GIS(profile='your_online_profile', verify_cert=False)
dest_gis = GIS(profile="your_online_admin_profile", verify_cert=False)
```
## Step 1. Get the `ArcGIS StoryMap` and export its resources
Once we have the item, we can use the `_version` variable we created earlier to run the `export_resources()` function we wrote earlier for `ArcGIS StoryMaps`, or the item's `resources` instance to export the supporting resources for `Classic Story Maps`. In this instance, we'll run `export_resources()` since we're copying an `ArcGIS Story Map`.
```
story_map_id = "358b83b5f776402fa726cfa316aa197c"
story_map = gis.content.get(story_map_id)
if _version <= [1,8,2]:
resource = export_resources(item=story_map)
else:
resource = story_map.resources.export()
# Visualize the Story Map item details
story_map
```
Examine the resources used by the Story Map
```
resource
```
We can see the `resource` variable stores a path to a zip file containing all the supporting resources needed to reconstruct our original `ArcGIS StoryMap`.
## Step 2. Get the `StoryMap` Item's [data](https://developers.arcgis.com/rest/users-groups-and-items/item-data.htm) to extract maps
`ArcGIS StoryMaps` utilize `Web Maps` and/or [Express Maps](https://doc.arcgis.com/en/arcgis-storymaps/author-and-share/add-maps.htm#ESRI_SECTION1_C30D73392D964D51A8B606128A8A6E8F) to contextualize the story's geography and allow direct interation with its mapped data. We can use the [`get_data()`](https://developers.arcgis.com/python/api-reference/arcgis.gis.toc.html#arcgis.gis.Item.get_data) method to extract each from the `StoryMap`.
```
print(f"{'-'*80}")
# Collect the Web Maps and Express Maps using the StoryMap's data. Use the set
# operator each item is collected only once for cloning.
story_map_json = story_map.get_data(try_json=True)
web_maps = set([v['data']['itemId'] for k, v in story_map_json['resources'].items() \
if v['type'].lower().find('webmap')>-1])
express_maps = set([v['data']['itemId'] for k, v in story_map_json['resources'].items() \
if v['type'].lower().find('expressmap')>-1])
```
Clone each `Web Map` from the `StoryMap` and assign a dictionary with the source `Web Map` id as the key, and the cloned `Web Map` id as the value. We'll use this dictionary to replace the source `Web Map` id with the cloned `Web Map` id in the new item we create in the destination GIS.
Let's examine the Web Map(s) from the set we created above.
```
webmap_mapper = {}
for wm in web_maps:
webmap_to_copy = gis.content.get(wm)
cloned_webmaps = dest_gis.content.clone_items([webmap_to_copy]) # Clones the WebMap
webmap_mapper[webmap_to_copy.id] = [i for i in cloned_webmaps if i.type == 'Web Map'][0].id
```
The `clone_items()` function used above duplicates not only the `Web Map`, but also any [`Layers`](https://doc.arcgis.com/en/arcgis-online/reference/layers.htm) contained in it. Depending upon the type of layer in the `Web Map`, cloning will create corresponding items in the destination.
Let's examine the cloned output and quickly compare the item details to the original `Web Map`.
```
cloned_webmaps
cloned_webmaps[2]
```
Let's also look at our dictionary and then use it to remap the original `Web Map` id to the cloned `Web Map` id in the json structure resulting from the `get_data()` function run earlier.
```
webmap_mapper
```
Remap the OLD ItemId to the New Item ID
```
story_map_text = json.dumps(story_map_json)
for k, v in webmap_mapper.items():
story_map_text = story_map_text.replace(k, v) # replace the IDs
```
## Step 3. Create a new StoryMap item in the Destination `GIS`
We'll use the original `story_map` variable properties to create a new `item` in our destination `GIS`. We'll eventually add the `resource` zip file we created earlier to the item to essetially duplicate the original `StoryMap`.
```
new_item = dest_gis.content.add({'type' : story_map.type,
'tags' : story_map.tags,
'title' : story_map.title,
'description' : story_map.description,
'typeKeywords' : story_map.typeKeywords,
'extent' : story_map.extent,
'text' :story_map_text}
)
new_item
```
Let's also download the original `item` thumbnail to use to update our new `item'.
```
# orig_thumbnail = story_map.download_thumbnail(r"your/file/path")
orig_thumbnail = story_map.download_thumbnail(r"C:/Job/sftriage/thumbnails/")
new_item.update(thumbnail=orig_thumbnail)
new_item
```
## Step 4. Add the `StoryMap` resources
We exported the images, expressmaps, text or other resources associated with the `StoryMap` earlier in this sample. Let's add those using the `resource` variable where we stored that output.
```
new_item.resources.add(resource,
archive=True)
```
## Step 5. Update the original `StoryMap` url
Set the `id` component of the new_`item`'s url to the new item `id` property.
```
new_item.update({'url': story_map.url.replace(story_map.id, new_item.id)})
```
## Step 6. Transfer Draft Resources
`ArcGIS StoryMaps` support a workflow that enables you to make changes to a published story, preview those changes, and then republish to make those changes to the existing story. Such updates to published stories are stored as unpublished drafts and are not visible to the audience until you are ready to republish the story.
The following code creates the supporting file to store draft resources and adds it as a supporting file to the destination `StoryMap`.
```
with tempfile.NamedTemporaryFile(mode='w', suffix='.json',
dir=tempfile.gettempdir(),
delete=False) as jsonfile:
jsonfile.write(json.dumps(new_item.get_data()))
new_item.resources.add(file=jsonfile.name)
type_keywords = [tk for tk in new_item.typeKeywords if 'smdraftresourceid:' not in tk]
type_keywords.append(f'smdraftresourceid:{os.path.basename(jsonfile.name)}')
new_item.update({'typeKeywords' : type_keywords})
```
Draft express map resources are handled separately and added as a resource.
```
if len(express_maps) > 0:
with tempfile.TemporaryDirectory() as d:
shutil.unpack_archive(filename=resource, extract_dir=d)
for expmap in express_maps:
express_draft = os.path.join(d, "draft_"+ expmap)
express_pub = os.path.join(d, "pub_" + expmap)
if os.path.isfile(express_pub):
shutil.copy(express_pub, express_draft)
new_item.resources.add(express_draft)
```
## Use the new `StoryMap`!
```
print("your new item can be found here: " + new_item.homepage)
new_item
```
| github_jupyter |
# 实战 Kaggle 比赛:图像分类 (CIFAR-10)
:label:`sec_kaggle_cifar10`
之前几节中,我们一直在使用深度学习框架的高级API直接获取张量格式的图像数据集。
但是在实践中,图像数据集通常以图像文件的形式出现。
在本节中,我们将从原始图像文件开始,然后逐步组织、读取并将它们转换为张量格式。
我们在 :numref:`sec_image_augmentation`中对CIFAR-10数据集做了一个实验。CIFAR-10是计算机视觉领域中的一个重要的数据集。
在本节中,我们将运用我们在前几节中学到的知识来参加CIFAR-10图像分类问题的Kaggle竞赛,(**比赛的网址是https://www.kaggle.com/c/cifar-10**)。
:numref:`fig_kaggle_cifar10`显示了竞赛网站页面上的信息。
为了能提交结果,你需要首先注册Kaggle账户。

:width:`600px`
:label:`fig_kaggle_cifar10`
首先,导入竞赛所需的包和模块。
```
import collections
import math
import os
import shutil
import pandas as pd
import torch
import torchvision
from torch import nn
from d2l import torch as d2l
```
## 获取并组织数据集
比赛数据集分为训练集和测试集,其中训练集包含50000张、测试集包含300000张图像。
在测试集中,10000张图像将被用于评估,而剩下的290000张图像将不会被进行评估,包含它们只是为了防止手动标记测试集并提交标记结果。
两个数据集中的图像都是png格式,高度和宽度均为32像素并有三个颜色通道(RGB)。
这些图片共涵盖10个类别:飞机、汽车、鸟类、猫、鹿、狗、青蛙、马、船和卡车。
:numref:`fig_kaggle_cifar10`的左上角显示了数据集中飞机、汽车和鸟类的一些图像。
### 下载数据集
登录Kaggle后,我们可以点击 :numref:`fig_kaggle_cifar10`中显示的CIFAR-10图像分类竞赛网页上的“Data”选项卡,然后单击“Download All”按钮下载数据集。
在`../data`中解压下载的文件并在其中解压缩`train.7z`和`test.7z`后,你将在以下路径中找到整个数据集:
* `../data/cifar-10/train/[1-50000].png`
* `../data/cifar-10/test/[1-300000].png`
* `../data/cifar-10/trainLabels.csv`
* `../data/cifar-10/sampleSubmission.csv`
`train`和`test`文件夹分别包含训练和测试图像,`trainLabels.csv`含有训练图像的标签,
`sample_submission.csv`是提交文件的范例。
为了便于入门,[**我们提供包含前1000个训练图像和5个随机测试图像的数据集的小规模样本**]。
要使用Kaggle竞赛的完整数据集,你需要将以下`demo`变量设置为`False`。
```
#@save
d2l.DATA_HUB['cifar10_tiny'] = (d2l.DATA_URL + 'kaggle_cifar10_tiny.zip',
'2068874e4b9a9f0fb07ebe0ad2b29754449ccacd')
# 如果你使用完整的Kaggle竞赛的数据集,设置demo为False
demo = True
if demo:
data_dir = d2l.download_extract('cifar10_tiny')
else:
data_dir = '../data/cifar-10/'
```
### [**整理数据集**]
我们需要整理数据集来训练和测试模型。
首先,我们用以下函数读取CSV文件中的标签,它返回一个字典,该字典将文件名中不带扩展名的部分映射到其标签。
```
#@save
def read_csv_labels(fname):
"""读取fname来给标签字典返回一个文件名"""
with open(fname, 'r') as f:
# 跳过文件头行(列名)
lines = f.readlines()[1:]
tokens = [l.rstrip().split(',') for l in lines]
return dict(((name, label) for name, label in tokens))
labels = read_csv_labels(os.path.join(data_dir, 'trainLabels.csv'))
print('# 训练样本 :', len(labels))
print('# 类别 :', len(set(labels.values())))
```
接下来,我们定义`reorg_train_valid`函数来[**将验证集从原始的训练集中拆分出来**]。
此函数中的参数`valid_ratio`是验证集中的样本数与原始训练集中的样本数之比。
更具体地说,令$n$等于样本最少的类别中的图像数量,而$r$是比率。
验证集将为每个类别拆分出$\max(\lfloor nr\rfloor,1)$张图像。
让我们以`valid_ratio=0.1`为例,由于原始的训练集有50000张图像,因此`train_valid_test/train`路径中将有45000张图像用于训练,而剩下5000张图像将作为路径`train_valid_test/valid`中的验证集。
组织数据集后,同类别的图像将被放置在同一文件夹下。
```
#@save
def copyfile(filename, target_dir):
"""将文件复制到目标目录"""
os.makedirs(target_dir, exist_ok=True)
shutil.copy(filename, target_dir)
#@save
def reorg_train_valid(data_dir, labels, valid_ratio):
"""将验证集从原始的训练集中拆分出来"""
# 训练数据集中样本最少的类别中的样本数
n = collections.Counter(labels.values()).most_common()[-1][1]
# 验证集中每个类别的样本数
n_valid_per_label = max(1, math.floor(n * valid_ratio))
label_count = {}
for train_file in os.listdir(os.path.join(data_dir, 'train')):
label = labels[train_file.split('.')[0]]
fname = os.path.join(data_dir, 'train', train_file)
copyfile(fname, os.path.join(data_dir, 'train_valid_test',
'train_valid', label))
if label not in label_count or label_count[label] < n_valid_per_label:
copyfile(fname, os.path.join(data_dir, 'train_valid_test',
'valid', label))
label_count[label] = label_count.get(label, 0) + 1
else:
copyfile(fname, os.path.join(data_dir, 'train_valid_test',
'train', label))
return n_valid_per_label
```
下面的`reorg_test`函数用来[**在预测期间整理测试集,以方便读取**]。
```
#@save
def reorg_test(data_dir):
"""在预测期间整理测试集,以方便读取"""
for test_file in os.listdir(os.path.join(data_dir, 'test')):
copyfile(os.path.join(data_dir, 'test', test_file),
os.path.join(data_dir, 'train_valid_test', 'test',
'unknown'))
```
最后,我们使用一个函数来[**调用前面定义的函数**]`read_csv_labels`、`reorg_train_valid`和`reorg_test`。
```
def reorg_cifar10_data(data_dir, valid_ratio):
labels = read_csv_labels(os.path.join(data_dir, 'trainLabels.csv'))
reorg_train_valid(data_dir, labels, valid_ratio)
reorg_test(data_dir)
```
在这里,我们只将样本数据集的批量大小设置为32。
在实际训练和测试中,应该使用Kaggle竞赛的完整数据集,并将`batch_size`设置为更大的整数,例如128。
我们将10%的训练样本作为调整超参数的验证集。
```
batch_size = 32 if demo else 128
valid_ratio = 0.1
reorg_cifar10_data(data_dir, valid_ratio)
```
## [**图像增广**]
我们使用图像增广来解决过拟合的问题。例如在训练中,我们可以随机水平翻转图像。
我们还可以对彩色图像的三个RGB通道执行标准化。
下面,我们列出了其中一些可以调整的操作。
```
transform_train = torchvision.transforms.Compose([
# 在高度和宽度上将图像放大到40像素的正方形
torchvision.transforms.Resize(40),
# 随机裁剪出一个高度和宽度均为40像素的正方形图像,
# 生成一个面积为原始图像面积0.64到1倍的小正方形,
# 然后将其缩放为高度和宽度均为32像素的正方形
torchvision.transforms.RandomResizedCrop(32, scale=(0.64, 1.0),
ratio=(1.0, 1.0)),
torchvision.transforms.RandomHorizontalFlip(),
torchvision.transforms.ToTensor(),
# 标准化图像的每个通道
torchvision.transforms.Normalize([0.4914, 0.4822, 0.4465],
[0.2023, 0.1994, 0.2010])])
```
在测试期间,我们只对图像执行标准化,以消除评估结果中的随机性。
```
transform_test = torchvision.transforms.Compose([
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize([0.4914, 0.4822, 0.4465],
[0.2023, 0.1994, 0.2010])])
```
## 读取数据集
接下来,我们[**读取由原始图像组成的数据集**],每个样本都包括一张图片和一个标签。
```
train_ds, train_valid_ds = [torchvision.datasets.ImageFolder(
os.path.join(data_dir, 'train_valid_test', folder),
transform=transform_train) for folder in ['train', 'train_valid']]
valid_ds, test_ds = [torchvision.datasets.ImageFolder(
os.path.join(data_dir, 'train_valid_test', folder),
transform=transform_test) for folder in ['valid', 'test']]
```
在训练期间,我们需要[**指定上面定义的所有图像增广操作**]。
当验证集在超参数调整过程中用于模型评估时,不应引入图像增广的随机性。
在最终预测之前,我们根据训练集和验证集组合而成的训练模型进行训练,以充分利用所有标记的数据。
```
train_iter, train_valid_iter = [torch.utils.data.DataLoader(
dataset, batch_size, shuffle=True, drop_last=True)
for dataset in (train_ds, train_valid_ds)]
valid_iter = torch.utils.data.DataLoader(valid_ds, batch_size, shuffle=False,
drop_last=True)
test_iter = torch.utils.data.DataLoader(test_ds, batch_size, shuffle=False,
drop_last=False)
```
## 定义[**模型**]
我们定义了 :numref:`sec_resnet`中描述的Resnet-18模型。
```
def get_net():
num_classes = 10
net = d2l.resnet18(num_classes, 3)
return net
loss = nn.CrossEntropyLoss(reduction="none")
```
## 定义[**训练函数**]
我们将根据模型在验证集上的表现来选择模型并调整超参数。
下面我们定义了模型训练函数`train`。
```
def train(net, train_iter, valid_iter, num_epochs, lr, wd, devices, lr_period,
lr_decay):
trainer = torch.optim.SGD(net.parameters(), lr=lr, momentum=0.9,
weight_decay=wd)
scheduler = torch.optim.lr_scheduler.StepLR(trainer, lr_period, lr_decay)
num_batches, timer = len(train_iter), d2l.Timer()
legend = ['train loss', 'train acc']
if valid_iter is not None:
legend.append('valid acc')
animator = d2l.Animator(xlabel='epoch', xlim=[1, num_epochs],
legend=legend)
net = nn.DataParallel(net, device_ids=devices).to(devices[0])
for epoch in range(num_epochs):
net.train()
metric = d2l.Accumulator(3)
for i, (features, labels) in enumerate(train_iter):
timer.start()
l, acc = d2l.train_batch_ch13(net, features, labels,
loss, trainer, devices)
metric.add(l, acc, labels.shape[0])
timer.stop()
if (i + 1) % (num_batches // 5) == 0 or i == num_batches - 1:
animator.add(epoch + (i + 1) / num_batches,
(metric[0] / metric[2], metric[1] / metric[2],
None))
if valid_iter is not None:
valid_acc = d2l.evaluate_accuracy_gpu(net, valid_iter)
animator.add(epoch + 1, (None, None, valid_acc))
scheduler.step()
measures = (f'train loss {metric[0] / metric[2]:.3f}, '
f'train acc {metric[1] / metric[2]:.3f}')
if valid_iter is not None:
measures += f', valid acc {valid_acc:.3f}'
print(measures + f'\n{metric[2] * num_epochs / timer.sum():.1f}'
f' examples/sec on {str(devices)}')
```
## [**训练和验证模型**]
现在,我们可以训练和验证模型了,而以下所有超参数都可以调整。
例如,我们可以增加周期的数量。当`lr_period`和`lr_decay`分别设置为4和0.9时,优化算法的学习速率将在每4个周期乘以0.9。
为便于演示,我们在这里只训练20个周期。
```
devices, num_epochs, lr, wd = d2l.try_all_gpus(), 20, 2e-4, 5e-4
lr_period, lr_decay, net = 4, 0.9, get_net()
train(net, train_iter, valid_iter, num_epochs, lr, wd, devices, lr_period,
lr_decay)
```
## 在 Kaggle 上[**对测试集进行分类并提交结果**]
在获得具有超参数的满意的模型后,我们使用所有标记的数据(包括验证集)来重新训练模型并对测试集进行分类。
```
net, preds = get_net(), []
train(net, train_valid_iter, None, num_epochs, lr, wd, devices, lr_period,
lr_decay)
for X, _ in test_iter:
y_hat = net(X.to(devices[0]))
preds.extend(y_hat.argmax(dim=1).type(torch.int32).cpu().numpy())
sorted_ids = list(range(1, len(test_ds) + 1))
sorted_ids.sort(key=lambda x: str(x))
df = pd.DataFrame({'id': sorted_ids, 'label': preds})
df['label'] = df['label'].apply(lambda x: train_valid_ds.classes[x])
df.to_csv('submission.csv', index=False)
```
向Kaggle提交结果的方法与 :numref:`sec_kaggle_house`中的方法类似,上面的代码将生成一个
`submission.csv`文件,其格式符合Kaggle竞赛的要求。
## 小结
* 将包含原始图像文件的数据集组织为所需格式后,我们可以读取它们。
* 我们可以在图像分类竞赛中使用卷积神经网络和图像增广。
## 练习
1. 在这场Kaggle竞赛中使用完整的CIFAR-10数据集。将超参数设为`batch_size = 128`,`num_epochs = 100`,`lr = 0.1`,`lr_period = 50`,`lr_decay = 0.1`。看看你在这场比赛中能达到什么准确度和排名。或者你能进一步改进吗?
1. 不使用图像增广时,你能获得怎样的准确度?
[Discussions](https://discuss.d2l.ai/t/2831)
| github_jupyter |
# Assignment 1
Welcome to the first programming assigment for the course. This assignments will help to familiarise you with qiskit while revisiting the topics discussed in this week's lectures.
### Submission Guidelines
For final submission, and to ensure that you have no errors in your solution, please use the 'Restart and Run All' option availble in the Kernel menu at the top of the page.
To submit your solution, run the completed notebook and then copy the two strings at the bottom of the notebook (which will be generated from your answers) and paste them in the google form associated with the assignment. In addition to this, please also attach the solved notebook as a file using the 'Add or Create' option under the 'Your Work' heading on the assignment page.
```
%matplotlib inline
import hashlib
import numpy as np
import matplotlib.pyplot as plt
# Importing standard Qiskit libraries and configuring account
from qiskit import QuantumCircuit, execute
from qiskit.circuit import Parameter
from qiskit.providers.aer import QasmSimulator, StatevectorSimulator
from qiskit.visualization import *
from qiskit.quantum_info import *
from qiskit.circuit.library import HGate
success_msg = 'Your answer is correct and has been saved. Please continue to the next section.'
fail_msg = 'Your answer is not correct. Please try again.'
```
## Rotations on the Bloch sphere
A general single-qubit state on the Bloch sphere is denoted by the statevector $$ |\psi\rangle = \cos{(\tfrac{\theta}{2})}|0\rangle + e^{i\phi}\sin{\tfrac{\theta}{2}}|1\rangle $$
We have seen the phase-shift operation $R_{\phi}$ corresponding to rotation by some angle $\phi$ about the $z$-axis. The statevector on the Bloch sphere traces a horizontal circle (a line of latitude) by $\phi$ radians. This operation has the matrix representation
$$
R_{\phi} =
\begin{pmatrix}
1 & 0 \\
0 & e^{i\phi}
\end{pmatrix}
$$
This operation changes the relative phase of the statevector, hence the name.
It stands to reason that there are operations which rotate a vector about other axes too. We have already seen that the phase-shift gate is a generalisation of the Pauli $Z$ gate. So it is reasonable to think that rotation about the $x$-axis and $y$-axis might be related to the Pauli $X$ and $Y$ operations. This is indeed the case. In general a rotation about a Pauli axis (these are synonymous with the $x$, $y$ and $z$ axes for our purposes) is represented by $$R_{P}(\theta) = \exp(-i\theta P/2) = \cos(\theta/2)I -i \sin(\theta/2)P$$
For the purposes of this assigment, a generalised rotation about the $x$-axis can be performed using the operation
$$
R_x(\theta) =
\begin{pmatrix}
\cos(\theta/2) & -i\sin(\theta/2)\\
-i\sin(\theta/2) & \cos(\theta/2)
\end{pmatrix}
$$
A generalised rotation about the $y$-axis can be performed using the operation
$$
R_y(\theta) =
\begin{pmatrix}
\cos(\theta/2) & - \sin(\theta/2)\\
\sin(\theta/2) & \cos(\theta/2).
\end{pmatrix}
$$
And a generalised rotation about the $z$-axis can be performed using the operation
$$
R_z(\phi) =
\begin{pmatrix}
e^{-i \phi/2} & 0 \\
0 & e^{i \phi/2}
\end{pmatrix}
$$
Note that here we have used an equivalent as it is different to $R_{\phi}$ by a global phase $e^{-i \phi/2}$.
If we start in the $|0\rangle$ state, as all quantum circuits in qiskit do, we can get to the general state $|\psi\rangle$ by first performing a rotation about the $y$-axis by an angle $\theta$, followed by a rotation about the $z$-axis by an angle $\phi$.
$$ |\psi\rangle = R_z(\phi) R_y(\theta) |0\rangle$$
You can find a summary of all the gates available in qiskit [here](https://qiskit.org/documentation/tutorials/circuits/3_summary_of_quantum_operations.html#Single-Qubit-Gates). For this assignment, you are only allowed to use these gates.
<div class="alert alert-block alert-warning"><b>Conventions:</b> Conventions are important and results can be confusing if the incorrect convention is used. In qiskit, both $\theta$ and $\phi$ are assumed to be positive in the counterclockwise direction. </div>
## Measurement in different bases
Qiskit allows measurement only in the computational basis directly. However, we can also perform meaurements in other bases.
For example, consider the Hadamard basis $\{|+\rangle, |-\rangle\}$. When we measure in this basis, we get an outcome that is one of the two basis vectors. This can be done by projecting a general state $|\psi\rangle$ onto each of the basis states $|+\rangle$ and $|-\rangle$. The probability of obtaining the $+$ outcome is $ P_{+}(|\psi\rangle) = |\langle +|\psi\rangle |^{2}$ and similarly for $-$. We note here that $|+\rangle = H|0\rangle$. Taking the adoint of this equation, we get $\langle +| = \langle 0|H^{\dagger} = \langle 0|H$ where we have used the fact that the Hadamard transformation is equal to it's adjoint (easily verified from the matrix representation). So we can write $ P_{+}(|\psi\rangle) = |\langle +|\psi\rangle |^{2} = |\langle 0| H\psi\rangle |^{2}$. It seems that the probability of measuring the $+$ outcome in the Hadamard basis is the same as the probability of measuring $0$ in the computational basis after applying the Hadamard transformation to the statevector $|\psi\rangle$. So, if we want to measure in the Hadamard basis, we need only to apply a Hadamard gate to our qubit and measure in the computational basis. Similar arguments apply to measuring in other bases too. If we can transform the standard basis states into the basis states for some basis $\mathcal{B}$ via some transformation, applying the inverse transformation allows us to measure in the basis $\mathcal{B}$ using standard basis measurements.
## **Problem 1**
Prepare the state $|i\rangle$ in a quantum circuit and measure it in the Hadamard basis $\{ |+\rangle, |-\rangle \}$
Below we have provided you with some code to create a quantum circuit. Remember that a qubit in a quantum circuit always begins in the $|0\rangle$ state. Add appropriate gates to prepare the $|i\rangle$ state and then add the appropriate gates and a standard basis measurement to measure required it in the Hadamard basis. You are required to use separate gates for both these actions.
```
qc1 = QuantumCircuit(1)
# Insert gates below to create the state
qc1.rx(-np.pi/2,0)
# Insert the necessary gates to change to the Hadamard basis below
qc1.h(0)
# Do not change below this line
qc1.measure_all()
qc1.draw('mpl')
```
## **Solution**
The easiest way to prepare the $|i\rangle$ state is to apply a rotation about the $x$-axis by $-\pi/2$ radians. This can be done using The $R_x(-\pi/2)$ gate as shown above. Alternatively, following the discussion in the single-qubit states and operations notebook, $H$ followed by $S$ will also prepare the $|i\rangle$ state. As mentioned above, to measure in the Hadamard basis, you need to apply an $H$ gate and then measure using a standard basis measurement
<div class="alert alert-block alert-info"><b>Instructions:</b>Once your circuit is ready, run the cell below to check and save your answer. You can change your answer by running these two cells in order again. </div>
```
basis_gates = ['id', 'x', 'y', 'z', 's', 't', 'sdg', 'tdg', 'h', 'p', 'sx' ,'r', 'rx', 'ry', 'rz', 'u', 'u1', 'u2', 'u3', 'barrier', 'measure']
assert list(qc1.count_ops()) != [], "Circuit cannot be empty"
assert set(qc1.count_ops().keys()).difference(basis_gates) == set(), "Only basic gates are allowed"
job = execute(qc1, backend=QasmSimulator(), shots=1024, seed_simulator=0)
counts = job.result().get_counts()
sv_check = Statevector.from_instruction(qc1.remove_final_measurements(inplace=False)).evolve(HGate()).equiv(Statevector.from_label('r'))
op_check_dict = qc1.count_ops()
_ = op_check_dict.pop('measure', None)
_ = op_check_dict.pop('barrier', None)
op_check = len(op_check_dict) > 1
print(success_msg if (sv_check and op_check) else fail_msg)
answer1 = hashlib.sha256((str(counts)+str(sv_check and op_check)).encode()).hexdigest()
plot_histogram(counts)
```
## Simulating polarisation of light on a quantum computer
In this exercise, we will use quantum computing to simulate a toy model of photon polarisation.
For a brief refresher on polarisation, and a demonstration of the experiment we will be simulating, watch the segment of the YouTube video below.
Reference:
Michael Melloch. "Polarization of Electromagnetic Waves" _YouTube_, Jun 20, 2019 https://www.youtube.com/watch?v=6N3bJ7Uxpp0
```
from IPython.display import YouTubeVideo, display
polariser_exp = YouTubeVideo('6N3bJ7Uxpp0', end=93, height=405, width=720, modestbranding=1)
display(polariser_exp)
```
To simulate the action of a polariser on plane-polarised light, we have to somehow map the problem onto the Bloch sphere. Here is one way to do it. As shown in the video, the light exiting the first polariser is plane-polarised, which means it is polarised in a plane at some angle to the horizontal and vertical axes. After that, the second polariser essentially _projects_ this onto the pass-axis orientation of the polariser, which can be seen as a kind of measurement. So in our quantum circuit, we will consider only this part of the experiment, and see what fraction of the plane polarised light is transmitted through the second polariser as a function of their relative orientation.
To do this, we need to consider only plane-polarised light. Let us assume that the light exiting the first polarizer is horizontally polarised. We know that if we place the second polariser with it's pass-axis vertical, no light will be transmitted. So these two states are orthogonal. Let us map these two states to $|0\rangle$ and $|1\rangle$ respectively (since these are also orthogonal). However, note that while the angle between the two orthogonal polarisation states is $\pi/2$, the angle on the Bloch sphere between $|0\rangle$ and $|1\rangle$ is $\pi$. We know that any other orientation of linearly polarised light can be written as a superposition of these two basis vectors. Moreover, since we are only considering linearly polarised light, we know that the relative phase is $0$. So we can restrict ourselves to the great circle on the Bloch sphere passing through $|0\rangle$, $|+\rangle$, $|1\rangle$ and $|-\rangle$.
## **Problem 2**
For this experiment, we will need to create a parameterised circuit, where the rotation angle is a parameter. Qiskit enables this using `Parameter()`. We will define a parameter $\beta$ which is the relative angle between the two polarisers. Given below is a quantum circuit where the qubit starts in the $|0\rangle$ state, the equivalent of horizontally plane polarised light. Change the measurement basis using a rotation gate with an angle of rotation in terms of the variable `beta` and measure in this basis. This basis change should correspond to rotating the second polariser by an angle $\beta$.
<div class="alert alert-block alert-warning"><b>Instructions:</b> We take $\beta$ to be positive in the counter-clockwise direction, as is the convention in qiskit. </div>
```
beta = Parameter('β')
qc2 = QuantumCircuit(1)
# Enter your code below this line
qc2.ry(-2*beta, 0)
qc2.measure_all()
# Do not change below this line
qc2.draw(output='mpl')
```
## **Solution**
First we need to visualise the part of the Bloch sphere onto which we're mapping linearly polarised states. As mentioned above, we can create any state on the Bloch sphere starting from $|0\rangle$ like so $$ |\psi\rangle = R_z(\phi) R_y(\theta) |0\rangle$$
We have also mentioned above that we will be restricting ourselves to the great circle passing through $|0\rangle$, $|+\rangle$, $|1\rangle$ and $|-\rangle$ and that the relative phase is $0$, so we are restricted to the $\phi=0$ circle on the Bloch sphere. The relevant region is highlighted in the diagram below.
<center><img style="display:block; width: 200px" src="https://raw.githubusercontent.com/deadbeatfour/quantum-computing-course/master/img/bloch_pol.png" alt="Mapping Linearly Polarised states onto the Bloch sphere"></center>
We note that due to the mapping mentioned above, $\theta = 2\beta$, so a rotation of the polariser in the range $0\le\beta\le\pi$ corresponds to $0\le\theta\le2\pi$. So the, polarisation state of a photon after the second polariser (with the polariser at an angle $\beta$) can be constructed using
$$ |\beta\rangle = R_y(2\beta) |0\rangle$$
However, we want to measure using this basis (this spans all states on the great circle). To do that, we use the same method as illustrated in the section above regarding measurement in different bases. $$P_{\beta}(|\psi\rangle) = |\langle \beta|\psi\rangle |^{2}$$
Taking the adjoint of the equation to prepate the state $|\beta\rangle$, we get $ \langle \beta| = \langle 0| R_y^{\dagger}(2\beta) = \langle 0| R_y(-2\beta)$, where the last equality is evident from the form of the matrix given above. With this,
$$P_{\beta}(|\psi\rangle) = |\langle \beta|\psi\rangle |^{2} = |\langle 0| R_y(-2\beta)|\psi\rangle |^{2}$$
So the appropriate gate to apply would be $R_y(-2\beta)$ and then perform a standard basis measurement.
<div class="alert alert-block alert-info"><b>Instructions:</b>Refer to the lecture slides for the theoretical transmission probability. Fill in that expression in the function below.</div>
```
def theoretical_prob(beta):
'''
Definition of theoretical transmission probability.
The expression for transmitted probability between two polarisers
with a relative angle `beta` given in radians
'''
# Fill in the correct expression for this probability and assign it to the variable tp below
# You may use numpy function like so: np.func_name()
tp = np.cos(-beta)**2
return tp
```
<div class="alert alert-block alert-info"><b>Instructions:</b>When you have defined the expression for the transmission probability, run the cell below to save your answer. You can change your answer by running these two cells in order again. </div>
We will rotate the second polariser through an angle of $\pi$. This means that $0\le\beta\le\pi$. We can repeat this experiment for $\beta$ values in this range and count the number of times we measured a photon passing through the second polariser.
<div class="alert alert-block alert-info"><b>Instructions:</b>When you have done that, run the cell below to perform the experiment</div>
```
beta_range = np.linspace(0, np.pi, 50)
num_shots = 1024
basis_gates = ['id', 'x', 'y', 'z', 's', 't', 'sdg', 'tdg', 'h', 'p', 'sx' ,'r', 'rx', 'ry', 'rz', 'u', 'u1', 'u2', 'u3', 'barrier', 'measure']
assert set(qc1.count_ops().keys()).difference(basis_gates) == set(), "Only basic gates are allowed"
job = execute(qc2,
backend=QasmSimulator(),
shots = num_shots,
parameter_binds=[{beta: beta_val} for beta_val in beta_range],
seed_simulator=0) # For consistent results
counts = job.result().get_counts()
# Calculating the probability of photons passing through
probabilities = list(map(lambda c: c.get('0', 0)/num_shots, counts))
pol_checks = [Statevector.from_instruction(qc2.bind_parameters({beta: beta_val})
.remove_final_measurements(inplace=False))
.equiv(Statevector([np.cos(-beta_val), np.sin(-beta_val)]))
for beta_val in beta_range]
print(success_msg if all(pol_checks) else fail_msg)
answer2 = hashlib.sha256((str(probabilities)+str(pol_checks)).encode()).hexdigest()
fig = plt.figure(figsize=(8,6))
ax = fig.add_subplot(111)
ax.plot(beta_range, probabilities, 'o', label='Experimental')
ax.plot(beta_range, theoretical_prob(beta_range), '-', label='Theoretical')
ax.set_xticks([i * np.pi / 4 for i in range(5)])
ax.set_xticklabels(['β', r'$\frac{\pi}{4}$', r'$\frac{\pi}{2}$', r'$\frac{3\pi}{4}$', r'$\pi$'], fontsize=14)
ax.set_xlabel('β', fontsize=14)
ax.set_ylabel('Probability of Transmission', fontsize=14)
ax.legend(fontsize=14)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import csv
f = open('find inter thres_300.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
find_inter_thres_300 = pd.DataFrame(data_list)
new_col = ['rep', 0.05, 0.1, 0.15, 0.2, 0.25, 0.3, 0.35, 0.4, 0.45, 0.5, 0.55, 0.6, 0.65, 0.7, 0.75, 0.8, 0.85, 0.9, 0.95, 1.0]
find_inter_thres_300.columns = new_col
find_inter_thres_300 = find_inter_thres_300.drop(find_inter_thres_300.index[0])
f = open('find inter thres_500.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
find_inter_thres_500 = pd.DataFrame(data_list)
new_col = ['rep', 0.05, 0.1, 0.15, 0.2, 0.25, 0.3, 0.35, 0.4, 0.45, 0.5, 0.55, 0.6, 0.65, 0.7, 0.75, 0.8, 0.85, 0.9, 0.95, 1.0]
find_inter_thres_500.columns = new_col
find_inter_thres_500 = find_inter_thres_500.drop(find_inter_thres_500.index[0])
f = open('find inter thres_1000.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
find_inter_thres_1000 = pd.DataFrame(data_list)
new_col = ['rep', 0.05, 0.1, 0.15, 0.2, 0.25, 0.3, 0.35, 0.4, 0.45, 0.5, 0.55, 0.6, 0.65, 0.7, 0.75, 0.8, 0.85, 0.9, 0.95, 1.0]
find_inter_thres_1000.columns = new_col
find_inter_thres_1000 = find_inter_thres_1000.drop(find_inter_thres_1000.index[0])
f = open('rep5_04_002.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
rep5_04_002 = pd.DataFrame(data_list)
rep5_04_002 = rep5_04_002.fillna(0)
cur_col = rep5_04_002.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
index += 1
l += 2
rep5_04_002.columns = new_col
rep5_04_002 = rep5_04_002.drop(rep5_04_002.index[0])
f = open('Buldy_RGG_30_rep5_04_002.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_rep5_04_002 = pd.DataFrame(data_list)
buldy_RGG_rep5_04_002 = buldy_RGG_rep5_04_002.fillna(0)
cur_col = buldy_RGG_rep5_04_002.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
index += 1
l += 2
buldy_RGG_rep5_04_002.columns = new_col
f = open('Buldy_RGG_rep30_03_0005.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_rep30_03_0005 = pd.DataFrame(data_list)
buldy_RGG_rep30_03_0005 = buldy_RGG_rep30_03_0005.fillna(0)
cur_col = buldy_RGG_rep30_03_0005.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
index += 1
l += 2
buldy_RGG_rep30_03_0005.columns = new_col
f = open('Buldy_RGG_100_rep30_03_0005.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_100_rep30_03_0005 = pd.DataFrame(data_list)
buldy_RGG_100_rep30_03_0005 = buldy_RGG_100_rep30_03_0005.fillna(0)
cur_col = buldy_RGG_100_rep30_03_0005.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
index += 1
l += 2
buldy_RGG_100_rep30_03_0005.columns = new_col
f = open('Buldy_RGG_200_rep30_03_0005.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_200_rep30_03_0005 = pd.DataFrame(data_list)
buldy_RGG_200_rep30_03_0005 = buldy_RGG_200_rep30_03_0005.fillna(0)
cur_col = buldy_RGG_200_rep30_03_0005.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
index += 1
l += 2
buldy_RGG_200_rep30_03_0005.columns = new_col
f = open('Buldy_RGG_200_rep30_035_0006.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_200_rep30_035_0006 = pd.DataFrame(data_list)
buldy_RGG_200_rep30_035_0006 = buldy_RGG_200_rep30_035_0006.fillna(0)
cur_col = buldy_RGG_200_rep30_035_0006.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
index += 1
l += 2
buldy_RGG_200_rep30_035_0006.columns = new_col
f = open('Buldy_RGG_30_rep30_04_0007.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_30_rep30_04_0007 = pd.DataFrame(data_list)
buldy_RGG_30_rep30_04_0007 = buldy_RGG_30_rep30_04_0007.fillna(0)
cur_col = buldy_RGG_30_rep30_04_0007.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
index += 1
l += 2
buldy_RGG_30_rep30_04_0007.columns = new_col
f = open('Buldy_RGG_1000_rep30_02_0005.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_1000_rep30_02_0005 = pd.DataFrame(data_list)
buldy_RGG_1000_rep30_02_0005 = buldy_RGG_1000_rep30_02_0005.fillna(0)
cur_col = buldy_RGG_1000_rep30_02_0005.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
index += 1
l += 2
buldy_RGG_1000_rep30_02_0005.columns = new_col
f = open('Buldy_RGG_50_rep100_045.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_50_rep100_045 = pd.DataFrame(data_list)
buldy_RGG_50_rep100_045 = buldy_RGG_50_rep100_045.fillna(0)
cur_col = buldy_RGG_50_rep100_045.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
new_col.append('step%d_far_dead_node' % index)
index += 1
l += 3
buldy_RGG_50_rep100_045.columns = new_col
f = open('Buldy_RGG_50_rep100_067.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_50_rep100_067 = pd.DataFrame(data_list)
buldy_RGG_50_rep100_067 = buldy_RGG_50_rep100_067.fillna(0)
cur_col = buldy_RGG_50_rep100_067.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
new_col.append('step%d_far_dead_node' % index)
index += 1
l += 3
buldy_RGG_50_rep100_067.columns = new_col
f = open('Buldy_RGG_200_rep100_0685.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_200_rep100_0685 = pd.DataFrame(data_list)
buldy_RGG_200_rep100_0685 = buldy_RGG_200_rep100_0685.fillna(0)
buldy_RGG_200_rep100_0685 = buldy_RGG_200_rep100_0685.replace(np.nan, 0)
buldy_RGG_200_rep100_0685 = buldy_RGG_200_rep100_0685.replace('', 0)
cur_col = buldy_RGG_200_rep100_0685.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
new_col.append('step%d_far_dead_node' % index)
index += 1
l += 3
buldy_RGG_200_rep100_0685.columns = new_col
f = open('Buldy_RGG_200_rep100_095.csv', 'rt')
reader = csv.reader(f)
data_list = []
for line in reader:
data_list.append(line)
f.close()
buldy_RGG_200_rep100_095 = pd.DataFrame(data_list)
buldy_RGG_200_rep100_095 = buldy_RGG_200_rep100_095.fillna(0)
buldy_RGG_200_rep100_095 = buldy_RGG_200_rep100_095.replace(np.nan, 0)
buldy_RGG_200_rep100_095 = buldy_RGG_200_rep100_095.replace('', 0)
cur_col = buldy_RGG_200_rep100_095.columns
new_col = ['rep', 'intra_thres', 'init_intra_edge', 'init_supp_edge', 'fin_intra_edge', 'fin_supp_edge', 'alive_nodes', 'tot_isol_node', 'tot_unsupp_node', 'cas_steps', 'init_far_node', 'fin_far_node', 'init_clust', 'fin_clust', 'init_mean_deg', 'fin_mean_deg', 'init_larg_comp', 'fin_larg_comp', 'deg_assort', 'dist_deg_cent', 'dist_bet_cent']
needed = len(cur_col) - len(new_col)
l = 0
index = 1
while l < needed:
new_col.append('step%d_isol' % index)
new_col.append('step%d_unsupp' % index)
new_col.append('step%d_far_dead_node' % index)
index += 1
l += 3
buldy_RGG_200_rep100_095.columns = new_col
find_inter_thres
rep5_04_002
buldy_RGG_rep5_04_002
buldy_RGG_rep30_03_0005
buldy_RGG_100_rep30_03_0005
buldy_RGG_200_rep30_03_0005
buldy_RGG_30_rep30_04_0007
buldy_RGG_1000_rep30_02_0005
buldy_RGG_50_rep100_045
buldy_RGG_50_rep100_067
buldy_RGG_200_rep100_0685
buldy_RGG_200_rep100_095
find_inter_thres_300.to_csv('proc_find_inter_thres_300.csv')
find_inter_thres_500.to_csv('proc_find_inter_thres_500.csv')
find_inter_thres_1000.to_csv('proc_find_inter_thres_1000.csv')
rep5_04_002.to_csv('proc_rep5_04_002.csv')
buldy_RGG_rep5_04_002.to_csv('proc_buldy_RGG_30_rep5_04_002.csv')
buldy_RGG_rep30_03_0005.to_csv('proc_buldy_RGG_rep30_03_0005.csv')
buldy_RGG_100_rep30_03_0005.to_csv('proc_buldy_RGG_100_rep30_03_0005.csv')
buldy_RGG_200_rep30_03_0005.to_csv('proc_buldy_RGG_200_rep30_03_0005.csv')
buldy_RGG_200_rep30_035_0006.to_csv('proc_buldy_RGG_200_rep30_035_0006.csv')
buldy_RGG_30_rep30_04_0007.to_csv('proc_buldy_RGG_30_rep30_04_0007.csv')
buldy_RGG_1000_rep30_02_0005.to_csv('proc_buldy_RGG_1000_rep30_02_0005.csv')
buldy_RGG_50_rep100_045.to_csv('proc_buldy_RGG_50_rep100_045.csv')
buldy_RGG_50_rep100_067.to_csv('proc_buldy_RGG_50_rep100_067.csv')
buldy_RGG_200_rep100_0685.to_csv('proc_buldy_RGG_200_rep100_0685.csv')
buldy_RGG_200_rep100_095.to_csv('proc_buldy_RGG_200_rep100_095.csv')
```
| github_jupyter |
### Regression results
```
import numpy as np
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.ERROR)
import tensorflow.contrib.slim as slim
import matplotlib.pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
from cn_reg_class import cn_reg_class
from mlp_reg_class import mlp_reg_class
from sklearn.datasets import load_boston
from util import nzr,get_train_test_datasets,gpusession,print_n_txt
print ("Packages loaded")
```
### Summarize results
```
# Configurations
dataset,dataset_name = load_boston(),'boston' # select dataset to use
test_size = 0.2
outlier_rates = [0,0.05,0.1,0.15,0.2,0.3,0.4,0.5]
rseeds = [0,1,2,3,4,5]
h_dims = [256,128]
actv = tf.nn.relu
l2_reg_coef = 1e-4
# Flags
RUN_CN = 1
RUN_L2 = 1
RUN_L1 = 1
RUN_RBST = 1
RUN_LRBST = 1
SAVE_TXT = True
SAVE_FINAL = False
SAVE_BEST = True
REMOVE_PREVS = True
# RMSEs to save
rmses_cn = np.zeros(shape=(len(outlier_rates),len(rseeds)))
rmses_l2 = np.zeros(shape=(len(outlier_rates),len(rseeds)))
rmses_l1 = np.zeros(shape=(len(outlier_rates),len(rseeds)))
rmses_robust = np.zeros(shape=(len(outlier_rates),len(rseeds)))
rmses_leaky_robust = np.zeros(shape=(len(outlier_rates),len(rseeds)))
# Save to txt
txt_name = 'res/robust_regression_results.txt'
f = open(txt_name,'w')
for s_idx,rseed in enumerate(rseeds): # for differnt random seeds
for o_idx,outlier_rate in enumerate(outlier_rates): # for different outlier rates
x_train,x_test,y_train,y_test = get_train_test_datasets(
dataset,dataset_name,_test_size=test_size,_outlier_rate=outlier_rate,
_seed=rseed,_SAVE_MAT=False,_VERBOSE=False) # get training data with outliers
# 1. Run ChoiceNet
if RUN_CN:
tf.reset_default_graph(); sess = gpusession()
tf.set_random_seed(rseed); np.random.seed(rseed) # fix seeds
CN = cn_reg_class(_name='cn_%s_err%02d_seed%d'%(dataset_name,outlier_rate*100,rseed),_x_dim=np.shape(x_train)[1],_y_dim=1,
_h_dims=h_dims,_k_mix=10,_actv=actv,_bn=slim.batch_norm,
_rho_ref_train=0.95,_tau_inv=1e-2,_var_eps=1e-4,
_pi1_bias=0.0,_log_sigma_Z_val=-1,
_kl_reg_coef=1e-6,_l2_reg_coef=l2_reg_coef,
_SCHEDULE_MDN_REG=0,_GPU_ID=1,_VERBOSE=0)
sess.run(tf.global_variables_initializer()) # initialize variables
CN.restore_from_npz(sess,_loadname='net/net_%s_best.npz'%(CN.name))
rmse = CN.test(_sess=sess,_x_train=x_train,_y_train=y_train,_x_test=x_test,_y_test=y_test,
_PLOT_TRAIN=0,_PLOT_TEST=0,_SAVE_FIG=0,
_title_str4data='Outlier rate:[%.1f]'%(outlier_rate),
_x_dim4plot=5,_x_name4plot='Average number of rooms per dwelling') # test
rmses_cn[o_idx,s_idx] = rmse
chars = ("rseed:[%d] outlier rate:[%.2f] choicenet:[%.4f]"%(rseed,outlier_rate,rmse))
print_n_txt(_f=f,_chars=chars,_DO_PRINT=True)
# 2. Run MLP with L2 loss
if RUN_L2:
tf.reset_default_graph(); sess = gpusession()
tf.set_random_seed(rseed); np.random.seed(rseed) # fix seeds
MLP = mlp_reg_class(_name='l2_%s_err%02d_seed%d'%(dataset_name,outlier_rate*100,rseed),_x_dim=np.shape(x_train)[1],_y_dim=1,
_h_dims=h_dims,_actv=actv,_bn=slim.batch_norm,#slim.batch_norm/None
_l2_reg_coef=l2_reg_coef,_GPU_ID=1,_L1_LOSS=0,_ROBUST_LOSS=0,_LEAKY_ROBUST_LOSS=0,_VERBOSE=0)
sess.run(tf.global_variables_initializer()) # initialize variables
MLP.restore_from_npz(sess,_loadname='net/net_%s_best.npz'%(MLP.name))
rmse = MLP.test(_sess=sess,_x_train=x_train,_y_train=y_train,_x_test=x_test,_y_test=y_test,
_PLOT_TRAIN=0,_PLOT_TEST=0,_SAVE_FIG=0,
_title_str4data='Outlier rate:[%.1f]'%(outlier_rate),
_x_dim4plot=5,_x_name4plot='Average number of rooms per dwelling') # test
rmses_l2[o_idx,s_idx] = rmse
chars = ("rseed:[%d] outlier rate:[%.2f] L2 loss:[%.4f]"%(rseed,outlier_rate,rmse))
print_n_txt(_f=f,_chars=chars,_DO_PRINT=True)
# 3. Run MLP with L1 loss
if RUN_L1:
tf.reset_default_graph(); sess = gpusession()
tf.set_random_seed(rseed); np.random.seed(rseed) # fix seeds
MLP = mlp_reg_class(_name='l1_%s_err%02d_seed%d'%(dataset_name,outlier_rate*100,rseed),_x_dim=np.shape(x_train)[1],_y_dim=1,
_h_dims=h_dims,_actv=actv,_bn=slim.batch_norm,#slim.batch_norm/None
_l2_reg_coef=l2_reg_coef,_GPU_ID=1,_L1_LOSS=1,_ROBUST_LOSS=0,_LEAKY_ROBUST_LOSS=0,_VERBOSE=0)
sess.run(tf.global_variables_initializer()) # initialize variables
MLP.restore_from_npz(sess,_loadname='net/net_%s_best.npz'%(MLP.name))
rmse = MLP.test(_sess=sess,_x_train=x_train,_y_train=y_train,_x_test=x_test,_y_test=y_test,
_PLOT_TRAIN=0,_PLOT_TEST=0,_SAVE_FIG=0,
_title_str4data='Outlier rate:[%.1f]'%(outlier_rate),
_x_dim4plot=5,_x_name4plot='Average number of rooms per dwelling') # test
rmses_l1[o_idx,s_idx] = rmse
chars = ("rseed:[%d] outlier rate:[%.2f] L1 loss:[%.4f]"%(rseed,outlier_rate,rmse))
print_n_txt(_f=f,_chars=chars,_DO_PRINT=True)
# 4. Run MLP with the robust loss
if RUN_RBST:
tf.reset_default_graph(); sess = gpusession()
tf.set_random_seed(rseed); np.random.seed(rseed) # fix seeds
MLP = mlp_reg_class(_name='rbst_%s_err%02d_seed%d'%(dataset_name,outlier_rate*100,rseed),_x_dim=np.shape(x_train)[1],_y_dim=1,
_h_dims=h_dims,_actv=actv,_bn=slim.batch_norm,#slim.batch_norm/None
_l2_reg_coef=l2_reg_coef,_GPU_ID=1,_L1_LOSS=0,_ROBUST_LOSS=1,_LEAKY_ROBUST_LOSS=0,_VERBOSE=0)
sess.run(tf.global_variables_initializer()) # initialize variables
MLP.restore_from_npz(sess,_loadname='net/net_%s_best.npz'%(MLP.name))
rmse = MLP.test(_sess=sess,_x_train=x_train,_y_train=y_train,_x_test=x_test,_y_test=y_test,
_PLOT_TRAIN=0,_PLOT_TEST=0,_SAVE_FIG=0,
_title_str4data='Outlier rate:[%.1f]'%(outlier_rate),
_x_dim4plot=5,_x_name4plot='Average number of rooms per dwelling') # test
rmses_robust[o_idx,s_idx] = rmse
chars = ("rseed:[%d] outlier rate:[%.2f] robust loss:[%.4f]"%(rseed,outlier_rate,rmse))
print_n_txt(_f=f,_chars=chars,_DO_PRINT=True)
# 5. Run MLP with the leaky robust loss
if RUN_LRBST:
tf.reset_default_graph(); sess = gpusession()
tf.set_random_seed(rseed); np.random.seed(rseed) # fix seeds
MLP = mlp_reg_class(_name='lrbst_%s_err%02d_seed%d'%(dataset_name,outlier_rate*100,rseed),_x_dim=np.shape(x_train)[1],_y_dim=1,
_h_dims=h_dims,_actv=actv,_bn=slim.batch_norm,#slim.batch_norm/None
_l2_reg_coef=l2_reg_coef,_GPU_ID=1,_L1_LOSS=0,_ROBUST_LOSS=0,_LEAKY_ROBUST_LOSS=1,_VERBOSE=0)
sess.run(tf.global_variables_initializer()) # initialize variables
MLP.restore_from_npz(sess,_loadname='net/net_%s_best.npz'%(MLP.name))
rmse = MLP.test(_sess=sess,_x_train=x_train,_y_train=y_train,_x_test=x_test,_y_test=y_test,
_PLOT_TRAIN=0,_PLOT_TEST=0,_SAVE_FIG=0,
_title_str4data='Outlier rate:[%.1f]'%(outlier_rate),
_x_dim4plot=5,_x_name4plot='Average number of rooms per dwelling') # test
rmses_leaky_robust[o_idx,s_idx] = rmse
chars = ("rseed:[%d] outlier rate:[%.2f] leaky robust loss:[%.4f]"%(rseed,outlier_rate,rmse))
print_n_txt(_f=f,_chars=chars,_DO_PRINT=True)
chars = ''
print_n_txt(_f=f,_chars=chars,_DO_PRINT=True)
print ("Done.")
plt.figure(figsize=(8,6))
h_cn,=plt.plot(outlier_rates,rmses_cn.mean(axis=1),'o-',color='r')
h_l2,=plt.plot(outlier_rates,rmses_l2.mean(axis=1),'o-',color='g')
h_l1,=plt.plot(outlier_rates,rmses_l1.mean(axis=1),'o-',color='b')
h_robust,=plt.plot(outlier_rates,rmses_robust.mean(axis=1),'o-',color='k')
h_leaky,=plt.plot(outlier_rates,rmses_leaky_robust.mean(axis=1),'o-',color='m')
plt.legend([h_cn,h_l2,h_l1,h_robust,h_leaky],
['ChoiceNet','L2 cost','L1 cost','Robust cost','Leaky robust cost'],
fontsize=15)
plt.xlabel('Outlier rate',fontsize=15)
plt.ylabel('RMSE',fontsize=15)
plt.savefig('fig/fig_%s_results.png'%(dataset_name))
plt.show()
```
### Print result
```
print (outlier_rates)
print ('ChoiceNet:',rmses_cn.mean(axis=1))
print ('L2:',rmses_l2.mean(axis=1))
print ('L1:',rmses_l1.mean(axis=1))
print ('Robust:',rmses_robust.mean(axis=1))
print ('LeakyRobust:',rmses_leaky_robust.mean(axis=1))
```
| github_jupyter |
```
from mayavi import mlab
mlab.init_notebook()
from pyscf import gto, scf, lo
import numpy as np
from functools import reduce
from pyscf.lo.orth import pre_orth_ao_atm_scf
import ase, scipy
from pyscf import lo
import itertools as itl
import ase.visualize as av
T,F=True,False
np.set_printoptions(precision=2,suppress=True)
def calc(aseobj, bst='cc-pvdz', icab=F):
zs = aseobj.numbers
nh = (zs==1).sum()
coords = aseobj.positions
assert zs[0]>1 and np.all(zs[1:]==1) #zs = []; coords = []
spin = sum(zs)%2
atom = ''
na = len(aseobj)
for i in range(na):
x,y,z = coords[i]
ai = aseobj[i]
atom += '%s %.8f %.8f %.8f;'%(ai.symbol, x,y,z)
if icab:
basis = {'H':'sto-3g'}
else:
basis = {'H':bst}
for si in ['C','N','O','F']: basis[si] = bst
mol = gto.M(atom=atom, basis=basis, verbose=0, spin=spin)
mf = None
if not icab:
mf = scf.RHF(mol)
mf.kernel()
return mol, mf
def get_hao(mol):
zs = mol.atom_charges()
nh = (zs==1).sum()
s1 = mol.intor_symmetric('int1e_ovlp')
b1 = pre_orth_ao_atm_scf(mol)
sb = reduce( np.dot, (b1.conjugate().T, s1, b1) )
aolbs = mol.ao_labels(); nao = len(aolbs)
sb_hx = sb[-nh:,:-nh] # overlap matrix H-X
u,d,vh = np.linalg.svd(sb_hx, full_matrices=False, compute_uv=True)
a1 = np.dot(vh.T, u.T)
# now Schmidt orthogonalization
n1 = nh
n2 = nao - nh
t = np.eye(n2)
t[:,:nh] = a1
for i in range(nh,n2):
for j in range(i):
cj = t[i,j]
t[:,i] -= cj*t[:,j]
t[:,i] /= np.linalg.norm(t[:,i])
for i in range(n2):
csi = t[i,:6]
so = ' '.join(['%10.2f '%si for si in csi])
print(aolbs[i], so)
return t
def get_new_dm1(mol, mf, t):
cs = mf.mo_coeff
return cs
def get_nho(m,bst='sto-3g',new_idx=None,debug=F):
mol, _ = calc(m, bst=bst, icab=T)
mf = scf.RHF(mol)
mf.kernel()
A1 = pre_orth_ao_atm_scf(mol)
s = mol.intor_symmetric('int1e_ovlp_sph')
s1 = reduce(np.dot, (A1.T,s,A1)) # under ANO basis
if debug: print('s1=',s1)
B1 = np.linalg.solve(A1,mf.mo_coeff)
dm1 = reduce( np.dot, (B1, np.diag(mf.mo_occ), B1.T) ) ##
if debug: print('dm1=',dm1)
p1 = dm1 # reduce(np.dot, (s, mf.make_rdm1(), s))
zs = mol.atom_charges()
nh = (zs==1).sum()
e1,v1 = scipy.linalg.eigh(p1[:-nh,:-nh], s1[:-nh,:-nh])
eigs1 = e1[::-1]; vs1 = v1[:,::-1]
if debug: print('eigs=',eigs1)
# exchange ao idx
if new_idx is None:
new_idx = np.arange(vs1.shape[0])
# = [0,3,2,1,4]
vs1u = vs1[:,new_idx]
c1 = np.eye(mol.nao)
c1[:-nh,:-nh] = vs1u # ANO basis
a = np.linalg.solve(c1,B1)
if debug: print('a=',a)
return eigs1,vs1,a
np.set_printoptions(precision=4,suppress=True)
bst = 'sto-3g' # 'cc-pvdz'
zs = [9,1]; coords = [[0.,0.,0],[0.,0.,0.98]]
m = ase.Atoms(zs,coords)
#av.view(m)
eigs, vs, a = get_nho(m,bst=bst,debug=T)
m.rotate(60, [1,1,1])
#av.view(m)
idxs = list(itl.permutations([1,2,3]))
for idx in idxs[:1]:
print('idx=',idx)
eigs2, vs2, a2 = get_nho(m,bst=bst,new_idx=[0]+list(idx)+[4], debug=F)
print('ddm: ', np.max(a2-a), np.min(a2-a), np.abs(a2)-np.abs(a))
print(np.abs(a2)-np.abs(a))
nao = mol.nao
aols = mol.ao_labels()
for i in range(nao-1):
si = aols[i]
for j in range(nao-1):
si += '%6.2f '%vs1[i,j]
print(si)
# write HAO
import interfaces._pyscf as pscf
#reload(pscf)
oo = pscf.io(mol)
c1 = np.eye(mol.nao)
c1[:-1,:-1] = vs1
orig, cell, dt = oo.orbital(c1, grids=[100,100,100], label=None)#'ch4-carbon')
a
#from ase.io.cube import read_cube_data
#data, atoms = read_cube_data('ch4-carbon_01.cube')
import visualization.mayavi as mv
_atoms = mv.draw_molecule(m, dt[0], cell, orig)
#_atoms
```
| github_jupyter |
## Text Classification using torchflare.
***
* Dataset: https://www.kaggle.com/columbine/imdb-dataset-sentiment-analysis-in-csv-format
```
import pandas as pd
import numpy as np
import torch
import torch.nn as nn
from sklearn.model_selection import train_test_split
import transformers
import torchflare.metrics as metrics
import torchflare.criterion as crit
import torchflare.callbacks as cbs
from torchflare.datasets import SimpleDataloader
from torchflare.experiments import Experiment
%load_ext nb_black
df = pd.read_csv("Train.csv")
df.text = df.text.apply(lambda x: x.lower())
train_df, valid_df = train_test_split(df, test_size=0.3)
tokenizer = transformers.AutoTokenizer.from_pretrained("prajjwal1/bert-tiny")
train_dl = SimpleDataloader.text_data_from_df(
df=train_df, input_col="text", label_cols="label", tokenizer=tokenizer, max_len=128
).get_loader(batch_size=16, shuffle=True, num_workers=0)
valid_dl = SimpleDataloader.text_data_from_df(
df=valid_df, input_col="text", label_cols="label", tokenizer=tokenizer, max_len=128
).get_loader(batch_size=16, shuffle=False)
class Model(torch.nn.Module):
def __init__(self):
super(Model, self).__init__()
self.bert = transformers.BertModel.from_pretrained(
"prajjwal1/bert-tiny", return_dict=False
)
self.bert_drop = nn.Dropout(0.3)
self.out = nn.Linear(128, 1)
def forward(self, input_ids, attention_mask, token_type_ids):
_, o_2 = self.bert(
input_ids, attention_mask=attention_mask, token_type_ids=token_type_ids
)
b_o = self.bert_drop(o_2)
output = self.out(b_o)
return output
model = Model()
metric_list = [metrics.Accuracy(num_classes=2, multilabel=False, threshold=0.6)]
callbacks = [
cbs.EarlyStopping(monitor="accuracy", patience=5),
cbs.ModelCheckpoint(monitor="accuracy"),
]
param_optimizer = list(model.named_parameters())
no_decay = ["bias", "LayerNorm.bias"]
optimizer_parameters = [
{
"params": [
p for n, p in param_optimizer if not any(nd in n for nd in no_decay)
],
"weight_decay": 0.001,
},
{
"params": [p for n, p in param_optimizer if any(nd in n for nd in no_decay)],
"weight_decay": 0.0,
},
]
exp = Experiment(
num_epochs=3,
save_dir="./models",
model_name="text_cls.bin",
fp16=False,
using_batch_mixers=False,
device="cuda",
compute_train_metrics=True,
seed=42,
)
exp.compile_experiment(
model=model,
optimizer="AdamW",
optimizer_params=dict(model_params=optimizer_parameters, lr=3e-4),
callbacks=callbacks,
scheduler="ReduceLROnPlateau",
scheduler_params=dict(mode="max", patience=2),
criterion=crit.BCEWithLogitsFlat,
metrics=metric_list,
main_metric="accuracy",
)
exp.perform_sanity_check(train_dl)
exp.run_experiment(train_dl=train_dl, valid_dl=valid_dl)
test_df = pd.read_csv("test.csv")
test_df.text = test_df.text.apply(lambda x: x.lower())
test_dl = SimpleDataloader.text_data_from_df(
df=test_df, input_col="text", label_cols=None, tokenizer=tokenizer, max_len=128
).get_loader(batch_size=16, shuffle=True)
ops = []
for op in exp.infer(path="./models/text_cls.bin", test_loader=test_dl):
z = torch.sigmoid(op).numpy()
ops.extend(z)
ops = np.concatenate(ops)
```
| github_jupyter |
<img src="https://raw.githubusercontent.com/google/jax/main/images/jax_logo_250px.png" width="300" height="300" align="center"/><br>
Welcome to another JAX tutorial. I hope you all have been enjoying the JAX Tutorials so far. We have already completed three tutorials on JAX each of which introduced an important concept.
In the first tutorial, we discussed **DeviceArray**, the core Data Structure in JAX. In the second tutorial, we looked into **Pure Functions** and their pros and cons. In the third tutorial, we looked into **Pseudo-Random Number Generation** in JAX, and how they are different from Numpy's PRNG. If you haven't gone through the previous tutorials, I highly suggest going through them. Here are the links:
1. [TF_JAX_Tutorials - Part 1](https://www.kaggle.com/aakashnain/tf-jax-tutorials-part1)
2. [TF_JAX_Tutorials - Part 2](https://www.kaggle.com/aakashnain/tf-jax-tutorials-part2)
3. [TF_JAX_Tutorials - Part 3](https://www.kaggle.com/aakashnain/tf-jax-tutorials-part3)
4. [TF_JAX_Tutorials - Part 4 (JAX and DeviceArray)](https://www.kaggle.com/aakashnain/tf-jax-tutorials-part-4-jax-and-devicearray)
5. [TF_JAX_Tutorials - Part 5 (Pure Functions in JAX)](https://www.kaggle.com/aakashnain/tf-jax-tutorials-part-5-pure-functions-in-jax/)
6. [TF_JAX_Tutorials - Part 6 (PRNG in JAX)](https://www.kaggle.com/aakashnain/tf-jax-tutorials-part-6-prng-in-jax/)
Today we will look into another important concepts: **Just In Time Compilation (JIT)** in JAX
# What is Just In Time (JIT) Compilation?
If we go by the [definition](https://en.wikipedia.org/wiki/Just-in-time_compilation) of JIT, then JIT is a way of compiling your code during the execution. A system implementing a JIT compiler typically continuously analyses the code being executed and identifies parts of the code where the speedup gained from compilation or recompilation would outweigh the overhead of compiling that code.
# JIT in JAX
As we discussed in the first chapter that JAX uses XLA for compilation. The `jax.jit(...)` transform does the just-in-time compilation and **transforms** your normal JAX Python functions so that they can be executed **more efficiently** in XLA.
Let's see a few examples of it before discussing the details
```
import os
import time
import requests
import jax
import jax.numpy as jnp
from jax import jit, grad, random
from jax.config import config
%config IPCompleter.use_jedi = False
def apply_activation(x):
return jnp.maximum(0.0, x)
def get_dot_product(W, X):
return jnp.dot(W, X)
# Always use a seed
key = random.PRNGKey(1234)
W = random.normal(key=key, shape=[1000, 10000], dtype=jnp.float32)
# Never reuse the key
key, subkey = random.split(key)
X = random.normal(key=subkey, shape=[10000, 20000], dtype=jnp.float32)
# JIT the functions we have
dot_product_jit = jit(get_dot_product)
activation_jit = jit(apply_activation)
for i in range(3):
start = time.time()
# Don't forget to use `block_until_ready(..)`
# else you will be recording dispatch time only
Z = dot_product_jit(W, X).block_until_ready()
end = time.time()
print(f"Iteration: {i+1}")
print(f"Time taken to execute dot product: {end - start:.2f} seconds", end="")
start = time.time()
A = activation_jit(Z).block_until_ready()
print(f", activation function: {time.time()-start:.2f} seconds")
```
Let's break down the above example into steps to know in detail what happened under the hood.
1. We defined two functions namely, `get_dot_product(...)` that does a dot product of weights and the inputs, and `apply_activation(...)` that applies `relu` on the previous result.
2. We then defined two transformations using `jit(function_name)`, and got the **compiled** versions of our functions
3. When you call the compiled function for the first time with the specified arguments, the execution time is pretty high. Why? Because the first call serves as the `warmup` phase. The warmup phase is nothing but the time taken by JAX **tracing**. Depending on the inputs, the tracers convert the code into an intermediate language, **`jaxprs`** (we will talk about this in a bit) which, is then compiled for execution in XLA
4. The subsequent calls run the compiled version of the code
**Note:** If you are benchmarking `jit` version of your function with something else, do a warmup first for a fair comparison else you will include the compilation time in the benchmarks
Before continuing further on JIT transformations, we will take a break here and try to understand the concept of **`jaxprs`** first
# Jaxprs
Jaxpr is an intermediate language for representing the normal Python functions. When you transform a function the function is first converted to simple statically-typed intermediate expressions by Jaxpr language, then the transformations are directly applied on these jaxprs.
1. A jaxpr instance represents a function with one or more typed parameters (input variables) and one or more typed results
2. The inputs and outputs have `types` and are represented as abstract values
3. Not all Python programs can be represented by jaxprs but many scientific computations and machine learning programs can
## Should you learn about Jaxprs?
Every transformation in JAX materializes to some form of `jaxpr`. If you want to understand how JAX works internally, or if you want to understand the result of JAX tracing, then yes, it is useful to understand jaxprs.
Let's take a few examples of how jaxpr works. We will first see how the functions we defined above are expressed by jaxpr
```
# Make jaxpr for the activation function
print(jax.make_jaxpr(activation_jit)(Z))
```
How to interpret this jaxpr?
1. The first line tells you that the function receives one argument `a`
2. The second line tells you that this is what would be executed on XLA, the max of (0, `a`)
3. The last line tells you the output being returned
Let's look at the jaxpr of our function that applies dot product
```
# Make jaxpr for the activation function
print(jax.make_jaxpr(dot_product_jit)(W, X))
```
Simlar to above, here:
1. The first line is telling that the function receives two input variables `a` and `b`, corresponding to our `W` and `X`
2. The second line is an XLA call where we perform the dot operation. (Check the dimesions numbers used for dot product)
3. The last line is the result to be returned denoted by `c`
Let's take another interesting example
```
# We know that `print` introduces but impurity but it is
# also very useful to print values while debugging. How does
# jaxprs interpret that?
def number_squared(num):
print("Received: ", num)
return num ** 2
# Compiled version
number_squared_jit = jit(number_squared)
# Make jaxprs
print(jax.make_jaxpr(number_squared_jit)(2))
```
Notice how the `num` inside the print statement is traced. Nothing stops you from running an impure function but you should be ready to encounter such side effects. The fact that the print statement is traced on the first call but may not be on the subsequent calls is because your python code will run at least once. Let's see that in action as well
```
# Subsequent calls to the jitted function
for i, num in enumerate([2, 4, 8]):
print("Iteration: ", i+1)
print("Result: ", number_squared_jit(num))
print("="*50)
```
We will take one more example to appreciate the beauty of `jaxprs` before moving on to JIT again
```
squared_numbers = []
# An impure function (using a global state)
def number_squared(num):
global squared_numbers
squared = num ** 2
squared_numbers.append(squared)
return squared
# Compiled verison
number_squared_jit = jit(number_squared)
# Make jaxpr
print(jax.make_jaxpr(number_squared_jit)(2))
```
A few things to notice:
1. The first line stats as usual and shows that we have an input variable `a`, corresponding to the `num` argument
2. The second line is an XLA call that squares the input number.
3. The last line returns the results of the XLA call denoted by `b`
**The side effect isn't captured by jaxpr**. jaxpr depends on **`tracing`**. The behavior of any transformed function is dependent on the traced values. You may notice the side effect on the first run but not necessarily on the subsequent calls. Hence jaxpr isn't even bothered about the global list in this case.
**Note:** One more important thing to note is the `device` value in the jaxprs. Although this argument is there unless you specify the device during jit transform like this `jit(fn_name, device=)`, no device would be listed here. This can be confusing sometimes because your computation would be running on some accelerator but here the device name won't be reflected. The logic behind this is that jaxpr is just an expression, independent of the logic where it is going to run. It is more concerned about the layout of the representation for XLA rather than the device on which the expression will be made to run
```
# Subsequent calls to the jitted function
for i, num in enumerate([4, 8, 16]):
print("Iteration: ", i+1)
print("Result: ", number_squared_jit(num))
print("="*50)
# What's in the list?
print("\n Results in the global list")
squared_numbers
```
You might be wondering that if the side effect was to appear on the first call, why there are two traced values in the global list. The reason is that the side effect may or may not appear on the subsequent calls. It is an unpredictable behavior.
# How much to JIT?
Before diving into the nuances related to JIT, let's assume that you have two functions that can be jitted with no problems, for example, our `get_dot_product(...)` and `apply_activation(..)` functions. Should you jit them both or should you use them into one function or module and jit that function/module? Le's see that in action
```
# Calling the two functions into a single function
# so that we can jit this function instead of jitting them
def forward_pass(W, X):
Z = get_dot_product(W, X)
A = apply_activation(Z)
return Z, A
# Always use a seed
key = random.PRNGKey(1234)
# We will use much bigger array this time
W = random.normal(key=key, shape=[2000, 10000], dtype=jnp.float32)
# Never reuse the key
key, subkey = random.split(key)
X = random.normal(key=subkey, shape=[10000, 20000], dtype=jnp.float32)
# JIT the functions we have individually
dot_product_jit = jit(get_dot_product)
activation_jit = jit(apply_activation)
# JIT the function that wraps both the functions
forward_pass_jit = jit(forward_pass)
for i in range(3):
start = time.time()
# Don't forget to use `block_until_ready(..)`
# else you will be recording dispatch time only
Z = dot_product_jit(W, X).block_until_ready()
end = time.time()
print(f"Iteration: {i+1}")
print(f"Time taken to execute dot product: {end - start:.2f} seconds", end="")
start = time.time()
A = activation_jit(Z).block_until_ready()
print(f", activation function: {time.time()- start:.2f} seconds")
# Now measure the time with a single jitted function that calls
# the other two functions
Z, A = forward_pass_jit(W, X)
Z, A = Z.block_until_ready(), A.block_until_ready()
print(f"Time taken by the forward pass function: {time.time()- start:.2f} seconds")
print("")
print("="*50)
```
Which approach to follow? That's up to you. Also, I don't have a confirmation whether the second approach always works but a Twitter user, who is a heavy JAX user, pointed this out.
# JIT and Python Control Flow
A natural question that comes to mind at this stage is `Why don't we just JIT everything? That would give a massive gain in terms of execution`. Though true in some sense, you can't jit everything. There are certain scenarios where jitting wouldn't work out of the box. Let's take a few examples to understand this
```
def square_or_cube(x):
if x % 2 == 0:
return x ** 2
else:
return x * x * x
# JIT transformation
square_or_cube_jit = jit(square_or_cube)
# Run the jitted version on some sample data
try:
val = square_or_cube_jit(2)
except Exception as ex:
print(type(ex).__name__, ex)
```
So why this code didn't work? Let's break down the whole process of JIT once again, including the one we have here
1. When we `jit` a function, we aim to get a compiled version of that function, so that we can cache and reuse the compiled code for different values of the arguments.
2. To achieve this, JAX traces it on abstract values that represent sets of possible inputs
3. There are [different levels of abstractions](https://github.com/google/jax/blob/main/jax/_src/abstract_arrays.py) that are used during tracing, and the kind of abstraction used for a particular function tracing depends on the kind of transformation is done.
4. By default, jit traces your code on the **`ShapedArray`** abstraction level, where each abstract value represents the set of all array values with a fixed shape and dtype. For example, if we trace using the abstract value ShapedArray((3,), jnp.float32), we get a view of the function that can be reused for any concrete value in the corresponding set of arrays. That means we can save on compile time.
Coming to the above code and why it failed, in this case, the value of `x` isn't concrete while tracing. As a result when we hit a line like `if x % 2 == 0`, the expression `x % 2` evaluates to an abstract `ShapedArray((), jnp.bool_)` that represents the set {True, False}. **When Python attempts to coerce that to a concrete True or False, we get an error: we don’t know which branch to take, and can’t continue tracing!**
Let's take one more example, this time involving a loop
```
def multiply_n_times(x, n):
count = 0
res = 1
while count < n:
res = res * x
count +=1
return x
try:
val = jit(multiply_n_times)(2, 5)
except Exception as ex:
print(type(ex).__name__, ex)
```
If the computation inside the loop is pretty expensive, you can still jit some part of the function body. Let's see it in action
```
# Jitting the expensive computational part
def multiply(x, i):
return x * i
# Specifying the static args
multiply_jit = jit(multiply, static_argnums=0)
# Leaving it as it as
def multiply_n_times(x, n):
count = 0
res = 1
while count < n:
res = multiply_jit(x, res)
count += 1
return res
%timeit multiply_n_times(2, 5)
```
# Caching
When you `jit` a function, it gets compiled on the first call. Any subsequent calls to the jitted function reuse the cached code. You pay the price once!
If we need to JIT a function that has a condition on the value of an input, we can tell JAX to make a less abstract tracer for a particular input by specifying `static_argnums`. The cost of this is that the resulting jaxpr is less flexible, so JAX will have to re-compile the function for every new value of the specified input. It is only a good strategy if the function is guaranteed to get limited different values.
<div class="alert alert-warning">
<strong>Warning: Don't do this!</strong>
</div>
```python
def multiply(x, i):
return x * i
def multiply_n_times(x, n):
count = 0
res = 1
while count < n:
res = jit(multiply)(x, res)
count += 1
return res
print(multiply_n_times(2, 5))
```
Doing that effectively creates a new jit transformed object at each call that will get compiled each time instead of reusing the same cached function.
That's it for Part-7! More in the next tutorial!
| github_jupyter |
# pyGemPick Tutorial 3: Outputing Detected Gold Particle Centers
## How to Output Gold Particle Centers To Use In Future Spatial-Statistical Analysis of Gold Particle Cross Correlation
In this tutorial we'll be using the **bin2df( )** function found in the pygempick.spatialstats package to record the x,y centers of each detected keypoint in each image!
```python
def bin2df(images):
i = 0 #image counter
j = 0 #total particles
#difine filtering paramaters
pclap = 25 #HCLAP anchor value
plog = 20 #HLOG anchor value
#make an empty dataframe, which will be filled with x,y centers of keypoints detected
data = pd.DataFrame()
#Change picking parameters per test set...
#Will ask you to fill in these parameters - keep it same as parameters used in Tutorial2
minArea = np.int(input('Min Area to Detect: '))
minCirc = np.float(input('Min Circularity: '))
minCon = np.float(input('Min Concavity: '))
minIner = np.float(input('Min Inertial Ratio: '))
for image in images:
orig_img = cv2.imread(image) ##reads specific test file
output1 = py.hclap_filt(pclap, orig_img, 'no') #High Contrast Laplace Filtering!
output2 = py.hlog_filt(plog, orig_img, 'no')
#image, minArea, minCirc, minConv, minIner, minThres
#use these parameters >> keypoints1 = py.pick(output1, 37, .71, .5 , .5, 0)
keypoints1 = py.pick(output1, minArea, minCirc, minCon , minIner, 0)
keypoints2 = py.pick(output2, minArea, minCirc, minCon , minIner, 0)
#this function removes duplicated detections
keypoints1, dup1 = py.key_filt(keypoints1, keypoints2)
#combine the two lists of keypoints
keypoints = keypoints1 + keypoints2
#record the subsequent keypoint centers and update the pandas dataframe
data, k = record_kp(i,keypoints,data)
j += k
i+=1
return data, j #returns data as df and total particles accounted...
```
```
#import required modules
import glob
import pygempick.spatialstats as spa
images = glob.glob('/home/joseph/Documents/pygempick/samples/compressed/*.jpg')
N = len(images)
data, gold = spa.bin2df(images)
file = input("Name Your File!")
data.to_csv('/home/joseph/Documents/pygempick/supdocs/{}'.format(file), index=False)
print('Counted {} gold particle in {} images.'.format(gold,N))
# this outputs a csv file which you cn find in the folder above or save to the folder of your choice!
# this dataframe (or file) can be used to calculate the spatial-point distribution of the gold
# particle positions on these EM mircrographs!
```
| github_jupyter |
# <font color='blue'>Monte Carlo Simulation</font>
# <font color='blue'>Monte Carlo Simulation and Time Series for Financial Modeling</font>
### Loading the Packages
```
# Python Version
from platform import python_version
print('Python Version:', python_version())
# Imports for data manipulation
import numpy as np
import pandas as pd
# Imports for viewing
import matplotlib.pyplot as plt
import matplotlib as m
import seaborn as sns
# Imports for statistical calculations
import scipy
from scipy.stats import kurtosis, skew, shapiro
import warnings
warnings.filterwarnings("ignore")
# Imports for formatting graphics
plt.style.use('fivethirtyeight')
m.rcParams['axes.labelsize'] = 14
m.rcParams['xtick.labelsize'] = 12
m.rcParams['ytick.labelsize'] = 12
m.rcParams['text.color'] = 'k'
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 20,10
```
### Loading Data
```
df = pd.read_csv("YOUR_PATH", parse_dates = True, index_col = "Date")
# View of the first lines
df.head()
# Data Types
df.dtypes
# Shape
df.shape
# Summary
df.describe()
```
## Viewing the Daily Share Closing Price
```
# Plot
plt.plot(df["Close"])
plt.title("Daily Share Closing Price", size = 14)
plt.show()
```
Calculation of the Daily Return of the Series.
```
#Calculating the percentage change in the daily closing quote of the shares
daily_return = df["Close"].pct_change().dropna()
daily_return.head()
```
Calculation of the Accumulated Return of the Series.
```
# Daily Return
accumulated_daily_return = (1 + daily_return).cumprod() - 1
accumulated_daily_return.max()
```
### Exploratory Analysis and Descriptive Statistics
Calculation of Average Return and Variation.
```
# Daily closing average
av_return_daily = np.mean(daily_return)
# Standard Deviation of Daily Closed
dev_daily_return = np.std(daily_return)
# Mean and Standard Deviation
print("Average Closing Return:", av_return_daily)
print("Standard Deviation of Closing Return:", dev_daily_return)
```
Note: Considering 252 Days of Trading on the United States Stock Exchange.
```
# Mean and Standard Deviation Per Year
print("Yearly Average Closing Return:", (1 + av_return_daily) ** 252 - 1)
print("Yearly Standard Deviation of Closing:", dev_daily_return*np.sqrt(252))
```
Although the stock's performance has been good in recent years, the average gain is low, but positive. Thus, the investor has not lost money.
```
# Plot
plt.plot(daily_return)
plt.title("Daily Return", size = 14)
plt.show()
```
The daily return has been constant over time, with only two major variations.
```
# Plot
plt.hist(daily_return, bins = 75)
plt.title("Daily Return Histogram", size = 14)
plt.show()
```
### Kurtosis and Skewness
```
print("Kurtosis:", kurtosis(daily_return))
print("Skewness:", skew(daily_return))
```
Although Kurtosis indicates that the records are close to the mean, Skewness demonstrates a distortion and a non-Normal Distribution of the data.
### Shapiro-Wilk Test
```
# Run the normality test for the series
is_normal_test_01 = shapiro(daily_return)[1]
# Check return based on p-value of 0.05
if is_normal_test_01 <= 0.05:
print("Rejects the Null Hypothesis of Data Normality.")
else:
print("Failure to reject the Null Hypothesis of Data Normality.")
```
There is no Normal Distribution.
To calculate the Daily Return Amount:
Log transformation to the series and then apply the differencing technique to remove the trend patterns and leave only the real data.
```
# Log Transformation and Differentiation
log_daily_return = (np.log(df["Close"]) - np.log(df["Close"]).shift(-1)).dropna()
# Mean and Standard Deviation After Transformation
log_av_daily_return = np.mean(log_daily_return)
log_dev_daily_return = np.std(log_daily_return)
# Plot
plt.plot(log_daily_return)
plt.title("Daily Return (Log Transformation)", size = 14)
plt.show()
# Plot
plt.hist(log_daily_return, bins = 75)
plt.title("Daily Return Histogram (Log Transformation)", size = 14)
plt.show()
# Kurtosis and Skewness
print("Kurtosis:", kurtosis(log_daily_return))
print("Skewness:", skew(log_daily_return))
# Normality Test for the Series
is_normal_test_02 = shapiro(log_daily_return)[1]
# Return Based on p-Value of 0.05
if is_normal_test_02 <= 0.05:
print("Rejects the Null Hypothesis of Data Normality.")
else:
print("Failure to reject the Null Hypothesis of Data Normality.")
```
Note: The data is still not normal, despite the reduction of data distortion. There is room for other transformations.
### Historical value
Historical value of the share.
```
# Variance Level
var_level = 95
var = np.percentile(log_daily_return, 100 - var_level)
print("Assurance that daily losses will not exceed VaR%.")
print("VaR 95%:", var)
# Var for the next 5 days
var * np.sqrt(5)
```
### Conditional Historical Value
```
# Variance Level
var_level = 95
var = np.percentile(log_daily_return, 100 - var_level)
cvar = log_daily_return[log_daily_return < var].mean()
print("In the worst 5% of cases on average losses were higher than the historical percentage.")
print("CVaR 95%:", cvar)
```
### Monte Carlo Simulation
```
# Number of Days Ahead
ahead_days = 252
# Number of Simulations
sim = 2500
# Last Share Value
last_price = 270.3
# Empy Array with the dimensions
res = np.empty((sim, ahead_days))
# Loop por cada simulação
for s in range(sim):
# Calculates the return with random data following a Normal Distribution
random_returns = 1 + np.random.normal(loc = log_av_daily_return,
scale = log_dev_daily_return,
size = ahead_days)
result = last_price * (random_returns.cumprod())
res[s, :] = result
# Defining the Simulated Series Index
index = pd.date_range("2020-03-11", periods = ahead_days, freq = "D")
results_all = pd.DataFrame(result.T, index = index)
average_results = results_all.apply("mean", axis = 1)
```
## Monte Carlo Simulation Result
```
fig, ax = plt.subplots(nrows = 2, ncols = 1)
# Plot
ax[0].plot(df["Close"][:"2018-12-31"])
ax[0].plot(results_all)
ax[0].axhline(270.30, c = "orange")
ax[0].set_title(f"Monte Carlo {sim} Simulation", size = 14)
ax[0].legend(["Historical Price", "Last Price = 270.30"])
ax[1].plot(df["Close"][:"2018-12-31"])
ax[1].plot(results_all.apply("mean", axis = 1), lw = 2)
ax[1].plot(average_results.apply((lambda x: x * (1+1.96 * log_dev_daily_return))),
lw = 2, linestyle = "dotted", c = "gray")
ax[1].plot(average_results, lw = 2, c = "orange")
ax[1].plot(average_results.apply((lambda x: x * (1-1.96 * log_dev_daily_return))),
lw = 2, linestyle = "dotted", c = "gray")
ax[1].set_title(f"Average Result Monte Carlo {sim} Simulation", size = 14)
ax[1].legend(["Price", "Average Forecast", "2x Standard Deviation"])
plt.show()
```
- Positive Forecast.
- Stocks tend to appreciate in the long term.
- Do not expect expressive returns.
| github_jupyter |
## 2020년 2월 6일 금요일
### 백준 6588번: 골드바흐의 추측문제
### 문제 : https://www.acmicpc.net/problem/6588
### 블로그 : https://somjang.tistory.com/entry/BaeKJoon-6588%EB%B2%88-%EA%B3%A8%EB%93%9C%EB%B0%94%ED%9D%90%EC%9D%98-%EC%B6%94%EC%B8%A1-%EB%AC%B8%EC%A0%9C-%ED%92%80%EC%9D%B4
### 첫번째 시도
먼저 입력 받은 수보다 작은 소수를 모두 구하고 가장 큰 소수와 가장 작은 소수와 더한 값이
입력받은 값이 나올때 그 두개의 값을 가지고 출력값을 만드는 것을 생각해보았습니다.
#### 입력받은 수 까지 존재하는 소수를 구하는 함수
```
def getPrimaryNums(N):
a = [2, 3, 5]
for i in range(6, N):
if i % 2 != 0 and i % 3 != 0 and i % 5 != 0:
a.append(i)
return a
while(True):
N = int(input())
if N == 0:
break
primary_nums = getPrimaryNums(N)
primary_nums_len = len(primary_nums)
break_flag = False
for i in primary_nums[::-1]:
if break_flag == True:
break
for j in range(len(primary_nums)):
if N == (i + primary_nums[j]):
print("{} = {} + {}".format(N, primary_nums[j], i))
break_flag=True
if break_flag == False:
print("Goldbach's conjecture is wrong.")
```
각각 입력받은 값에 대해서 가장 큰 값과 가장 작은 값을 더해보며 값이 나올경우
출력 후 break_flag를 True로 변경하여 반복문을 중단하였습니다.
만약 반복문이 종료가 되었는데도 원하는 조건의 값이 없을때 break_flag가 False일 경우
Goldbach의 추측은 틀렸다라고 출력해주도록 하였습니다.
시간초과로 통과를 하지 못하였습니다.
생각해보니 GoldBach의 추측이 틀렸을 경우 이중 반복문이 끝까지 실행되어
O(N²)의 시간복잡도를 가지게 됩니다.
---
### 두번째 시도
입력받은 N 이하의 모든 소수를 구하고
combination 라이브러리를 활용하여 N이하의 소수 두 개로 만들 수 있는 조합을 만들고
그 조합중에서 두 수의 합이 N인 것을 찾은 뒤
두 수의 차가 가장 큰 조합을 찾아 출력까지 해주는 방법을 생각해 보았습니다.
```
from itertools import combinations
while(True):
N = int(input())
if N == 0:
break
primary_nums = getPrimaryNums(N)
combs = list(combinations(primary_nums, 2))
new_combs = []
for comb in combs:
if sum(comb) == N:
new_combs.append(comb)
sub = []
for comb in new_combs:
sub.append(comb[1] - comb[0])
max_sub = max(sub)
for i in range(len(new_combs)):
if max_sub == sub[i]:
print("{} = {} + {}".format(N, new_combs[i][0], new_combs[i][1]))
```
결과가 나오기는 하나 테스트에서 1,000,000을 입력하게 되면...
너무 많은 조합을 만들게되어 메모리를 많이 점유하는 문제가 발생했습니다.
역시나 시간초과!
---
### 세번째 시도
에라토스테네스의 체의 방법을 쓰면 해결할 수 있다고하여 먼저 에라토스테네스의 체의 방법 문제를 풀어보았습니다.
```
def getPrimaryNum_Eratos(N):
nums = [True] * (N + 1)
for i in range(2, len(nums) // 2 + 1):
if nums[i] == True:
for j in range(i+i, N, i):
nums[j] = False
return [i for i in range(2, N) if nums[i] == True]
getPrimaryNum_Eratos(10)
def getPrimaryNum_Eratos(N):
nums = [True] * (N + 1)
for i in range(2, len(nums) // 2 + 1):
if nums[i] == True:
for j in range(i+i, N, i):
nums[j] = False
return [i for i in range(2, N) if nums[i] == True]
while(True):
N = int(input())
if N == 0:
break
primary_nums = getPrimaryNum_Eratos(N)
primary_nums_len = len(primary_nums)
break_flag = False
for i in primary_nums[::-1]:
if break_flag == True:
break
for j in range(len(primary_nums)):
if N == (i + primary_nums[j]):
print("{} = {} + {}".format(N, primary_nums[j], i))
break_flag=True
if break_flag == False:
print("Goldbach's conjecture is wrong.")
```
### 결과는 시간초과
소수를 구하는 법이 문제가아닌 뒤 쪽에 문제가 있는 것 같습니다.
이번에는 조금 다르게 접근하여 보기로 했습니다.
N 이하의 소수를 모두 찾고 가장 작은 소수부터 소수의 개수만큼 반복문을 돌면서 N - 소수 를 했을때
그 값도 소수인 경우 print를 해주면 될 것이라고 생각했습니다.
하지만 결과는 처참했습니다.
이제 어디서 더 줄일 수 있을까?
고민해보니 이 문제는 반복문을 돌면서 0이 나올때까지 계속 값을 구해야합니다.
근데 지금의 방식은 while 반복문이 실행될때마다 새로 소수를 구하고있습니다.
애초에 1,00,000 이하의 모든 소수를 구해두고 하면 어떨까? 생각해보았습니다.
---
### 마지막 시도
```
def getPrimaryNum_Eratos(N):
nums = [True] * (N + 1)
for i in range(2, len(nums) // 2 + 1):
if nums[i] == True:
for j in range(i+i, N, i):
nums[j] = False
return [[i for i in range(2, N) if nums[i] == True], nums]
primary_nums = getPrimaryNum_Eratos(1000000)[0]
primary_bools = getPrimaryNum_Eratos(1000000)[1]
while(True):
N = int(input())
if N == 0:
break
for i in range(N // 2):
if primary_bools[N-primary_nums[i]] == True:
print("{} = {} + {}".format(N, primary_nums[i], N-primary_nums[i]))
break
```
| github_jupyter |
# Transforms and Multi-Table Relational Databases
* This notebook shows how to run transforms directly on a mutli-table relational database while keeping the referential integrity of primary and foreign keys intact.
* This notebook also contains instructions on how to transform data residing in CSV files.
* Primary and foreign keys must originally be IDs.
* This notebook is discussed in ths [blog](https://gretel.ai/blog/transforms-and-multi-table-relational-databases)
## Our ecommerce database
* Execute the below cell to see a diagram of the database we'll be using in this blueprint. The lines in the diagram show connections between primary and foreign keys
```
from IPython.display import Image
Image("https://gretel-blueprints-pub.s3.us-west-2.amazonaws.com/rdb/ecommerce_db.png",width = 600, height = 600)
```
## Getting started
```
!pip install pyyaml Faker pandas
!pip install SQLAlchemy
!pip install -U gretel-client
# Specify your Gretel API key
from getpass import getpass
import pandas as pd
from gretel_client import configure_session, ClientConfig
pd.set_option('max_colwidth', None)
configure_session(ClientConfig(api_key=getpass(prompt="Enter Gretel API key"),
endpoint="https://api.gretel.cloud"))
```
## Gather data and schema relationships directly from a database
* For demonstration purposes, we'll first grab our ecommerce SQLite database from S3
* This notebook can be run on any database SQLAlchemy supports such as Postgresql or MySQL
* For example, if you have a postgres database, simply swap the `sqlite:///` connection string for a `postgres://` one in the `create_engine` command
* Using SQLAlchemy's reflection extension, we will crawl the schema, gather table data and produce a list of relationships by table primary key.
```
from sqlalchemy import create_engine
!wget https://gretel-blueprints-pub.s3.amazonaws.com/rdb/ecom.db
engine = create_engine("sqlite:///ecom.db")
# Gather the table data from the database
from sqlalchemy import MetaData, text
# This is the directory where we will temporarily store csv files for the transformer model
base_path = "./"
metadata = MetaData()
metadata.reflect(engine)
rdb_config = {}
rdb_config["table_data"] = {}
rdb_config["table_files"] = {}
for name, table in metadata.tables.items():
df = pd.read_sql_table(name, engine)
rdb_config["table_data"][name] = df
filename = name + ".csv"
df.to_csv(filename, index=False, header=True)
rdb_config["table_files"][name] = filename
# Extract primary/foriegn key relationshihps
from collections import defaultdict
rels_by_pkey = defaultdict(list)
for name, table in metadata.tables.items():
for col in table.columns:
for f_key in col.foreign_keys:
rels_by_pkey[(f_key.column.table.name, f_key.column.name)].append((name, col.name))
list_of_rels_by_pkey = []
for p_key, f_keys in rels_by_pkey.items():
list_of_rels_by_pkey.append([p_key] + f_keys)
rdb_config["relationships"] = list_of_rels_by_pkey
# Extract each table's primary key
rdb_config["primary_keys"] = {}
for name, table in metadata.tables.items():
for col in table.columns:
if col.primary_key:
rdb_config["primary_keys"][name] = col.name
```
## Alternatively, specify primary/foreign key relationships and locations of data csv files
* This is an alternative to the above three cells that work directly with a database
* First, assign `base_path` to the directory where the csv files are located.
* Then, add a name/key pair for each table name/filename to `rdb_config["table_files"]`
* Add all primary keys for each table to `rdb_config["primary_keys"]`
* Add all foreign key/primary keys that connect to the same set under `rdb_config["relationshipts"]`
```
# base_path is the directory where your csv files can be found
base_path = "https://gretel-blueprints-pub.s3.amazonaws.com/rdb/"
rdb_config = {
"table_files": {
"users": "users.csv",
"order_items": "order_items.csv",
"events": "events.csv",
"inventory_items": "inventory_items.csv",
"products": "products.csv",
"distribution_center": "distribution_center.csv"
},
# List the primary keys for each table
"primary_keys": {
"users": "id",
"order_items": "id",
"events": "id",
"inventory_items": "id",
"products": "id",
"distribution_center": "id"
},
# List the (table, field) relationships between primary and foreign keys
"relationships": [
{("users","id"),
("order_items","user_id"),
("events","user_id")
},
{("inventory_items","id"),
("order_items","inventory_item_id")
},
{("products","id"),
("inventory_items","product_id")
},
{("distribution_center","id"),
("products","distribution_center_id"),
("inventory_items", "product_distribution_center_id")
}
]
}
# Gather the table data using the filenames entered above
rdb_config["table_data"] = {}
for table in rdb_config["table_files"]:
filename = base_path + rdb_config["table_files"][table]
df = pd.read_csv(filename)
rdb_config["table_data"][table] = df
```
## Take a look at your data by joining two tables
* Note that every record in the table "order_items" matches to an entry in the table "users"
* An "inner" join will take the intersection of two tables
```
table_to_view1 = "order_items"
table_to_view2 = "users"
df1 = rdb_config["table_data"][table_to_view1]
df2 = rdb_config["table_data"][table_to_view2]
joined_data = df1.join(df2.set_index('id'), how='inner', on='user_id', lsuffix='_order_items', rsuffix='_users')
print("Number of records in order_items table is " + str(len(df1)))
print("Number of records in user table is " + str(len(df2)))
print("Number of records in joined data is " + str(len(joined_data)))
show_fields = ['id', 'user_id', 'inventory_item_id', 'sale_price', 'shipped_at', 'delivered_at', 'first_name', 'last_name', 'age', 'latitude', 'longitude']
joined_data.filter(show_fields).head()
```
## Define your transform policies
* Define one policy per table which transforms any PII or sensitive information that could be used to reidentify a user.
* You needn't include a transform for any of the primary/foreign key combinations. We'll be handling those seperately in order to maintain referential integrity.
* However, if a table contains a primary key that does not match to a foreign key, that field should be included in the transforms.
* Note the tables inventory_items, products and distribution center contain only public information so there will be no transformation.
* To run this notebook on a different database, simply enter the table names and policy files below. We will assume all policy files are located in the `policy_dir` defined below
```
policy_dir = "https://gretel-blueprints-pub.s3.amazonaws.com/rdb/"
transform_policies = {}
transform_policies["users"] = "users_policy.yaml"
transform_policies["order_items"] = "order_items_policy.yaml"
transform_policies["events"] = "events_policy.yaml"
transform_policies["inventory_items"] = None
transform_policies["products"] = None
transform_policies["distribution_center"] = None
```
## Policy detail
* Let's take a detailed look at the transforms for the users table.
* Within the `rules` section, we define each type of transformation we want, each one beginning with `- name`.
* We start by replacing any field classified as a person name or email address with a fake version.
* Note, we choose to leave "city", "state", "country" and "zip" as is since it's public knowledge that this database is about user ecommerce transactions in Arizona.
* We then transform the "created_at" timestamp using a random date shift.
* And finally, we transform the numeric fields of age, latitude and longitude with a random numeric shift.
* Note, we do not transform "id" because it is a primary key that matches to a foreign key. We'll take care of that later.
```
from smart_open import open
policy_file = transform_policies["users"]
policy_file_path = policy_dir + policy_file
yaml_file = open(policy_file_path, "r")
policy = yaml_file.read()
yaml_file.close()
print(policy)
```
## Define functions to train models and generate transformed data
```
import yaml
import numpy as np
from smart_open import open
from sklearn import preprocessing
from gretel_client import create_project
from gretel_client.helpers import poll
def create_model(table:str, project):
# Read in the transform policy
policy_file = transform_policies[table]
policy_file_path = policy_dir + policy_file
yaml_file = open(policy_file_path, "r")
policy = yaml_file.read()
yaml_file.close()
# Get the dataset_file_path
dataset_file = rdb_config["table_files"][table]
dataset_file_path = base_path + dataset_file
# Create the transform model
model = project.create_model_obj(model_config=yaml.safe_load(policy))
# Upload the training data. Train the model.
model.data_source = dataset_file_path
model.submit(upload_data_source=True)
print("Model training started for " + table)
return model
def generate_data(table:str, model):
record_handler = model.create_record_handler_obj()
# Get the dataset_file_path
dataset_file = rdb_config["table_files"][table]
dataset_file_path = base_path + dataset_file
# Submit the generation job
record_handler.submit(
action = "transform",
data_source = dataset_file_path,
upload_data_source = True
)
print("Generation started for " + table)
return record_handler
```
## Set off training and generation and monitor progress
* Note, jobs will be set off in parallel then we will monitor them until completion
```
# Initiate model training and generation
import pandas as pd
# model_progress will hold the status of each model during training and generation
model_progress = {}
# transformed_tables will hold the final transformed tables
transformed_tables = {}
# Create a new project
project = create_project(display_name="rdb_transform3")
for table in rdb_config["table_files"]:
if transform_policies[table] is not None:
model = create_model(table, project)
model_progress[table] = {
"model": model,
"model_status": "pending",
"record_handler": "",
"record_handler_status": "",
}
# If there is no transform on a table, copy it over as is
else:
transformed_tables[table] = rdb_config["table_data"][table]
# Monitor model progression
more_to_do = True
while more_to_do:
# Check status of training
more_to_do = False
for table in model_progress:
status = model_progress[table]["model_status"]
if (status == 'created') or (status == 'pending') or (status == "active"):
more_to_do = True
model = model_progress[table]["model"]
model._poll_job_endpoint()
status = model.__dict__['_data']['model']['status']
# If status is now complete, submit the generation job
if status == 'completed':
print("Training completed for " + table)
rh = generate_data(table, model)
model_progress[table]["record_handler"] = rh
model_progress[table]["record_handler_status"] = "pending"
model_progress[table]["model_status"] = status
# If training status was already complete, check status of the generation job
elif status == 'completed':
status = model_progress[table]["record_handler_status"]
if (status == 'created') or (status == 'pending') or (status == 'active'):
rh = model_progress[table]["record_handler"]
rh._poll_job_endpoint()
status = rh.__dict__['_data']['handler']['status']
# If generation is now complete, get the synthetic data
if status == 'completed':
transform_df = pd.read_csv(rh.get_artifact_link("data"), compression='gzip')
transformed_tables[table] = transform_df
model_progress[table]["record_handler_status"] = status
print("Generation completed for " + table)
elif status != 'error':
more_to_do = True
else:
print("\nGeneration for " + table + " ended in error")
more_to_do = False
model_progress[table]["record_handler_status"] = status
elif status == 'error':
print("\nTraining for " + table + " ended in error")
model_progress[table]["model_status"] = status
more_to_do = False
if status != 'error':
print("\nModel training and initial generation all complete!")
```
## Transform primary/foreign key combinations
* To ensure referential integrity on each primary key/foreign key table set, we will first fit a Label Encoder on the combined set of unique values in each table.
* We then run the Label Encoder on the key field in each table in the set.
* This both de-identifies the keys as well as serves to ensure referential integrity.
```
primary_keys_processed = []
def transform_keys(key_set):
# Get array of unique values from each table, can use dfs in transformed_tables
first = True
field_values = set()
for table_field_pair in key_set:
table, field = table_field_pair
# The first pair is a primary key
if first:
primary_keys_processed.append(table)
first = False
field_values = field_values.union(set(rdb_config["table_data"][table][field]))
# Train a label encoder
field_values_list = list(set(field_values))
le = preprocessing.LabelEncoder()
le.fit(field_values_list)
# Run the label encoder on dfs in transformed_tables
for table_field_pair in key_set:
table, field = table_field_pair
transformed_tables[table][field] = le.transform(rdb_config["table_data"][table][field])
# Run our transform_keys function on each key set
for key_set in rdb_config["relationships"]:
transform_keys(key_set)
# Process the remaining primary keys
for table in rdb_config["primary_keys"]:
if table not in primary_keys_processed:
le = preprocessing.LabelEncoder()
key_field = rdb_config["primary_keys"][table]
le.fit(rdb_config["table_data"][table][key_field])
transformed_tables[table][key_field] = le.transform(rdb_config["table_data"][table][key_field])
```
## View the transformed content
* We'll again join the order_items and users tables
```
pd.set_option("display.max_columns", None)
table_to_view1 = "order_items"
table_to_view2 = "users"
df1 = transformed_tables[table_to_view1]
df2 = transformed_tables[table_to_view2]
joined_data = df1.join(df2.set_index('id'), how='inner', on='user_id', lsuffix='_order_items', rsuffix='_users')
print("Number of records in order_items table is " + str(len(df1)))
print("Number of records in user table is " + str(len(df2)))
print("Number of records in joined data is " + str(len(joined_data)))
show_fields = ['id', 'user_id', 'inventory_item_id', 'sale_price', 'shipped_at', 'delivered_at', 'first_name', 'last_name', 'age', 'latitude', 'longitude']
joined_data.filter(show_fields).head()
```
## Save the transformed data back into an SQLite database
```
!cp ecom.db ecom_xf.db
engine_xf = create_engine("sqlite:///ecom_xf.db")
with engine_xf.connect() as conn:
for table in transformed_tables:
command = "DELETE FROM " + table
conn.execute(command)
transformed_tables[table].to_sql(table, con=conn, if_exists='append', index=False)
```
## Alternatively save the transformed data back into a Postgres database
```
from sqlalchemy import text
with engine.connect() as conn:
result = conn.execute(text("create database ecom_xf with template ecom"))
engine_xf = create_engine("postgres:///ecom_xf.db")
for table in transformed_tables:
transformed_tables[table].to_sql(table, con=engine_xf, if_exists='replace', index=False)
conn.commit()
```
## Alterntively, save the transformed content into CSV files
```
# Change final_dir to be the location where you'd like your csv files saved
final_dir = "./"
for table in transformed_tables:
df = transformed_tables[table]
filename = final_dir + table + '_transform.csv'
df.to_csv(filename, index=False, header=True)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/yohanesnuwara/reservoir-engineering/blob/master/Unit%203%20Reservoir%20Statics/notebook/3_examples.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **Unit 3 Reservoir Statics (Examples)**
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
!git clone https://github.com/yohanesnuwara/reservoir-engineering
import sys, os
sys.path.append('/content/reservoir-engineering/Unit 3 Reservoir Statics/functions')
```
# Example 3.1 Calculating Oil Density in a Reservoir from Well Test Data
```
"Question. calculate oil density in the reservoir based on well data"
# known
qo = 250 # in STB/day
qg = 150 # Mscf/day
api = 45
sg = 0.82
temp = 240 # in fahrenheit
# standard conditions
temp_sc = 60 # in fahrenheit
pressure_sc = 14.73 # 1 atm = 14.73 psia
z_sc = 1 # gas z factor at standard condition
# calculate gas-oil ratio
Rs = qg * 1000 / qo # in scf gas/STB oil
Rs_converted = Rs * (1 / 5.6148) # convert to scf gas/scf oil
print("Gas-oil ratio:", Rs_converted, "scf gas/scf oil")
# oil FVF at surface/standard condition using Standing correlation (Equation 2.37 See Unit 2)
so = 141.5 / (api + 131.5)
F = Rs * ((sg / so)**0.5) + (1.25 * temp_sc) # Rs must be in scf/STB
Bo = 0.972 + (0.000147*(F**1.1756))
print("Oil FVF at standard condition:", Bo, "RB/STB")
# oil density at surface/standard condition
rhowater = 62.366 # 1 g/cm3 = 62.366 lbm/ft3
rhooil_sc = so * rhowater
print("Oil density at standard condition:", rhooil_sc, "lbm/ft3")
# gas density at surface/standard condition (Eq 2.23, real-gas law)
R = 10.732 # gas constant in (ft3*psi)/(lb-mol*R)
rhogas_sc = (28.97 * sg * pressure_sc) / (z_sc * R * (temp_sc + 459)) # temp converted to Rankine
print("Gas density at standard condition:", rhogas_sc, "lbm/ft3")
# finally, oil density at reservoir condition
rhooil = (rhooil_sc + (rhogas_sc * Rs_converted)) / Bo
print("Oil density at reservoir condition:", rhooil, "lbm/ft3")
# oil density gradient
rhooil_grad = rhooil / 144 # 144 is factor conversion from density lbm/ft3 to psi/ft
print("Oil density gradient:", rhooil_grad, "psi/ft")
```
# Example 3.2 Investigating Whether Two Oil Wells Are in Pressure Communication
```
# Question. Are the two wells in pressure communication?
# known
api = 32
Rs = 500 # in scf/STB
temp = 200 # in fahrenheit
sg = 0.72
Bo = 1.3 # in RB/STB
well1_elev = 7134; well2_elev = 7028 # well elevation in ft
well1_depthkb = 5652; well2_depthkb = 5426 # well depth from KB in ft
well1_pressure = 2453; well2_pressure = 2306 # well pressure in psia
# standard conditions
temp_sc = 60 # in fahrenheit
pressure_sc = 14.73 # 1 atm = 14.73 psia
z_sc = 1 # gas z factor at standard condition
# calculate oil density at standard condition
so = 141.5 / (131.5 + api)
rhowater = 62.366
rhooil_sc = so * rhowater
# print("Oil density at standard condition:", rhooil_sc, "lbm/ft3")
# calculate gas density at standard condition
R = 10.732 # gas constant in (ft3*psi)/(lb-mol*R)
rhogas_sc = (28.97 * sg * pressure_sc) / (z_sc * R * (temp_sc + 459)) # temp converted to Rankine
# print("Gas density at standard condition:", rhogas_sc, "lbm/ft3")
# calculate oil density at reservoir condition
Rs_converted = Rs * (1 / 5.6148) # convert to scf gas/scf oil
rhooil = (rhooil_sc + (rhogas_sc * Rs_converted)) / Bo
print("Oil density at reservoir condition:", rhooil, "lbm/ft3")
# oil density gradient
rhooil_grad = rhooil / 144 # 144 is factor conversion from density lbm/ft3 to psi/ft
# print("Oil density gradient:", rhooil_grad, "psi/ft")
# check if the two wells are communicated
well1_depthmsl = well1_elev - well1_depthkb
well2_depthmsl = well2_elev - well2_depthkb
delta_depth = np.abs(well2_depthmsl - well1_depthmsl)
if well1_depthmsl > well2_depthmsl:
print("well 1 is shallower", delta_depth, "ft than well 2")
else:
print("well 1 is deeper", delta_depth, "ft than well 2")
well1_pressure_extrapolated = well2_pressure + rhooil_grad * delta_depth # pressure extrapolation, Eq 3.1
print("From extrapolation, well 1 is supposed to be", well1_pressure_extrapolated, "psi")
delta_extrapolated_actual = np.abs(well1_pressure_extrapolated - well1_pressure)
percent_delta = delta_extrapolated_actual / well1_pressure * 100
print("Difference of extrapolated well 1 pressure to actual pressure:", delta_extrapolated_actual, "psi, or percent difference:", percent_delta, "%")
if percent_delta < 1:
print("The two wells are most likely in pressure communication")
else:
print("The two wells are most likely NOT in pressure communication")
```
# Example 2.3 Extrapolating Pressures in a Gas Reservoir
```
# Question 1. Extrapolate pressure 100 ft above and below datum using simplified integration formula (Eq 3.8, 3.9)
# known
sg = 0.65
pressure = 2500
temp_average = 170
z_average = 0.87
delta = 100
pressure_extrapolated_below = pressure*(np.exp((0.01877 * sg * delta) / (z_average * (temp_average + 459)))) # temp in Rankine
pressure_extrapolated_above = pressure*(np.exp(-(0.01877 * sg * delta) / (z_average * (temp_average + 459)))) # temp in Rankine
print("Using Eq 3.8, extrapolated pressure at", delta, "ft above datum:", pressure_extrapolated_above, "psi, and", delta, "ft below datum:", pressure_extrapolated_below, "psi")
# Question 2. Extrapolate pressure 100 ft above and below datum using general Eq 3.1 from density Eq 3.5
# Eq 3.5
R = 10.732
rhogas = (28.97 * sg * pressure) / (z_average * R * (temp_average + 459)) # temp convert to Rankine
print("Gas density at", pressure, "psi:", rhogas, "lbm/ft3")
# gas density gradient
rhogas_grad = rhogas / 144
# extrapolate using Eq 3.1
pressure_extrapolated_below2 = pressure + rhogas_grad * delta
pressure_extrapolated_above2 = pressure - rhogas_grad * delta
print("Using Eq 3.1, extrapolated pressure at", delta, "ft above datum:", pressure_extrapolated_above2, "psi, and", delta, "ft below datum:", pressure_extrapolated_below2, "psi")
```
| github_jupyter |
# Fitting BERT Classifier to Twitter MBTI
```
import tensorflow as tf
import torch
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
from sklearn.model_selection import train_test_split
from transformers import BertTokenizer, BertConfig
from transformers import AdamW, BertForSequenceClassification
from tqdm import tqdm, trange
import pandas as pd
import io
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score
from sklearn.metrics import roc_curve
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
def expand_frame(df, length):
ndf = pd.DataFrame()
for i in range(1,30):
adf = df.copy(deep= True)
adf['text'] = adf['text'].apply(lambda x: x[0 + length*5*i : length * 5 * (i+1)])
ndf = ndf.append(adf)
ndf.reset_index(drop=True, inplace= True)
return ndf
```
# Loading the Twitter personality dataset:
# OPTION 1 : Load raw without tokenized:
```
ofile = open('Data/personality_likes_large.csv', encoding = 'cp1252', mode='r' )
raw_df = pd.read_csv(ofile ,index_col=0)
ofile.close()
wifile = open('train_tokenized_large_exp.csv', mode = 'w+')
wtfile = open('test_tokenized_large_exp.csv', mode = 'w+')
wifile.truncate(0)
wtfile.truncate(0)
train_df, test_df = train_test_split(raw_df, stratify = raw_df['type'], random_state= 1729, test_size= 0.12)
train_df = expand_frame(train_df, 256)
train_df.dropna(inplace = True)
test_df.to_csv(wtfile)
wtfile.close()
# Create sentence and label lists
sentences = train_df.text.values
# We need to add special tokens at the beginning and end of each sentence for BERT to work properly
sentences = ["[CLS] " + str(sentence) + " [SEP]" for sentence in sentences]
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased', do_lower_case=True)
tokenized_texts = [tokenizer.tokenize(sent) for sent in sentences]
train_df['tokenized_texts'] = tokenized_texts
train_df.to_csv(wifile)
wifile.close()
ofile = open('train_tokenized_large_exp.csv', mode='r' )
edf = pd.read_csv(ofile, index_col = 0)
```
# OPTION 2: Load Dataframe from disk
```
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased', do_lower_case=True)
test_file = open('test_tokenized_large_exp.csv')
ofile = open('train_tokenized_large_exp.csv')
edf = pd.read_csv(ofile, index_col = 0)
edf.dropna(inplace= True)
#edf, test_df = train_test_split(edf, random_state=2020, test_size=0.10)
test_df = pd.read_csv(test_file, index_col=0)
test_df = test_df.reset_index(drop = True)
def fourType(x):
if ((x=='INFJ') | (x=='INFP') | (x=='ENFJ') | (x=='ENFP')):
return 1
elif ((x=='INTJ') | (x=='INTP') | (x=='ENTJ') | (x=='ENTP')):
return 2
elif ((x=='ISFJ') | (x=='ISTJ') | (x=='ESFJ') | (x=='ESTJ')):
return 3
else:
return 0
edf['role'] = edf['type'].map(fourType)
test_df['role'] = test_df['type'].map(fourType)
df = edf[edf['role']==1].sample(10808, random_state = 34)
df = df.append(edf[edf['role']==2].sample(10808, random_state = 35))
df = df.append(edf[edf['role']==3].sample(10808, random_state = 35))
df = df.append(edf[edf['role']==0].sample(10808, random_state = 35))
from ast import literal_eval
df = df.sample(frac=1).reset_index(drop=True)
tokenized_texts = df['tokenized_texts'].map(literal_eval)
test_df = test_df.reset_index(drop = True)
tokenized_texts
# Set the maximum sequence length. The longest sequence in our training set is 47, but we'll leave room on the end anyway.
# In the original paper, the authors used a length of 512.
MAX_LEN = 256
import os
os.environ['KERAS_BACKEND'] = 'tensorflow'
from keras.preprocessing.sequence import pad_sequences
# Pad our input tokens
input_ids = [tokenizer.convert_tokens_to_ids(x) for x in tokenized_texts]
input_ids = pad_sequences(input_ids, maxlen=MAX_LEN, dtype="long", truncating="post", padding="post")
input_ids
# Create attention masks
attention_masks = []
# Create a mask of 1s for each token followed by 0s for padding
for seq in input_ids:
seq_mask = [float(i>0) for i in seq]
attention_masks.append(seq_mask)
# Use train_test_split to split our data into train and validation sets for training
labels = df.role.values
train_inputs, validation_inputs, train_labels, validation_labels = train_test_split(input_ids, labels,
random_state=2020, test_size=0.01)
train_masks, validation_masks, _, _ = train_test_split(attention_masks, input_ids,
random_state=2020, test_size=0.01)
```
# **OPTION 3: Load v2 Dataframe from disk**
```
!pip install transformers
import ast
import sklearn
import pandas as pd
import transformers
import torch
from sklearn.model_selection import train_test_split
import numpy as np
from ast import literal_eval
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
from sklearn.model_selection import train_test_split
from transformers import BertTokenizer, BertConfig
from transformers import AdamW, BertForSequenceClassification
from tqdm import tqdm, trange
import pandas as pd
import io
import pickle
from google.colab import drive
drive.mount('/content/drive')
def from_np_array(array_string):
array_string = ','.join(array_string.replace('[ ', '[').split())
return np.array(ast.literal_eval(array_string))
folder_path = '/content/drive/MyDrive/MBTI_DL_Data/'
ofile = open(folder_path + 'Dataframes/master_v2.3.csv', mode = 'r', encoding= 'windows-1252')
df = pd.read_pickle(folder_path + 'Dataframes/master_v2.3.csv' )
df
adf = df[['extravert', 'thinking', 'intuitive', 'judging', 'lang', 'liked_tweets']]
adf['liked_tweets']
def trim_liked(x):
r = ''
s = x.split()
for word in s:
if (len(word)<20) and (word[0] != '@'):
r+= ' ' + word
return r
adf['liked_tweets'] = adf['liked_tweets'].apply(lambda x: '. '.join(ast.literal_eval(x)))
adf['liked_tweets'] = adf['liked_tweets'].apply(lambda x: trim_liked(x))
adf
target_mode = 'intuitive'
sample_size = min(len(adf[adf[target_mode]==1]), len(adf[adf[target_mode]==0] ))
typeA = adf[adf[target_mode]==1].sample(sample_size)
typeB = adf[adf[target_mode]==0].sample(sample_size)
sample_size
data_A = typeA[['liked_tweets', target_mode]]
data_B = typeB[['liked_tweets', target_mode]]
data_df = data_A.append(data_B).sample(frac = 1, random_state = 42).reset_index(drop = True)
train_df, test_df = train_test_split(data_df, random_state= 1729, test_size= 0.10)
train_df.reset_index(inplace = True, drop = True)
test_df.reset_index(inplace = True, drop = True)
train_df
test_df
def expand_frame(df, length):
ndf = pd.DataFrame()
cdf = df.copy(deep= True)
cdf['split_tweets'] = cdf['liked_tweets'].apply(lambda x: x.split())
print('1')
for i in range(0, 21):
ccdf = cdf[cdf['split_tweets'].apply(lambda x: (len(x) > (0 + i * length)) == 1)]
ccdf['split_tweets'] = ccdf['split_tweets'].apply(lambda x: x[0 + length*i : length * (i+1)])
ndf = ndf.append(ccdf)
ndf.reset_index(drop=True, inplace= True)
print('1')
ndf['liked_tweets'] = ndf['split_tweets'].apply(lambda x: ' '.join(x))
return ndf
train_df = expand_frame(train_df, 512)
train_df = train_df.sample(frac = 1, random_state= 153)
train_df.reset_index(inplace= True, drop = True)
train_df
train_df
test_df
import seaborn as sns
sns.histplot(train_df['liked_tweets'].apply(lambda x: len(x.split())))
# Create sentence and label lists
sentences = train_df.liked_tweets.values
# We need to add special tokens at the beginning and end of each sentence for BERT to work properly
sentences = ["[CLS] " + str(sentence) + " [SEP]" for sentence in sentences]
labels = train_df.intuitive.values
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased', do_lower_case=True)
tokenized_texts = [tokenizer.tokenize(sent) for sent in sentences]
print ("Tokenize the first sentence:")
print (tokenized_texts[0])
# Set the maximum sequence length. The longest sequence in our training set is 47, but we'll leave room on the end anyway.
# In the original paper, the authors used a length of 512.
MAX_LEN = 512
input_ids = [tokenizer.convert_tokens_to_ids(x) for x in tokenized_texts]
import os
os.environ['KERAS_BACKEND'] = 'tensorflow'
from keras.preprocessing.sequence import pad_sequences
# Pad our input tokens
input_ids = pad_sequences(input_ids, maxlen=MAX_LEN, dtype="long", truncating="post", padding="post")
# Create attention masks
attention_masks = []
# Create a mask of 1s for each token followed by 0s for padding
for seq in input_ids:
seq_mask = [float(i>0) for i in seq]
attention_masks.append(seq_mask)
# Use train_test_split to split our data into train and validation sets for training
train_inputs, validation_inputs, train_labels, validation_labels = train_test_split(input_ids, labels,
random_state=2020, test_size=0.01)
train_masks, validation_masks, _, _ = train_test_split(attention_masks, input_ids,
random_state=2020, test_size=0.01)
```
# BERT
```
# Convert all of our data into torch tensors, the required datatype for our model
train_inputs = torch.tensor(train_inputs)
validation_inputs = torch.tensor(validation_inputs)
train_labels = torch.tensor(train_labels)
validation_labels = torch.tensor(validation_labels)
train_masks = torch.tensor(train_masks)
validation_masks = torch.tensor(validation_masks)
# Select a batch size for training. For fine-tuning BERT on a specific task, the authors recommend a batch size of 16 or 32
batch_size = 8
# Create an iterator of our data with torch DataLoader. This helps save on memory during training because, unlike a for loop,
# with an iterator the entire dataset does not need to be loaded into memory
train_data = TensorDataset(train_inputs, train_masks, train_labels)
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=batch_size)
validation_data = TensorDataset(validation_inputs, validation_masks, validation_labels)
validation_sampler = SequentialSampler(validation_data)
validation_dataloader = DataLoader(validation_data, sampler=validation_sampler, batch_size=batch_size)
model = BertForSequenceClassification.from_pretrained("bert-base-uncased", num_labels=2)
model.cuda()
param_optimizer = list(model.named_parameters())
no_decay = ['bias', 'gamma', 'beta']
optimizer_grouped_parameters = [
{'params': [p for n, p in param_optimizer if not any(nd in n for nd in no_decay)],
'weight_decay_rate': 0.01},
{'params': [p for n, p in param_optimizer if any(nd in n for nd in no_decay)],
'weight_decay_rate': 0.0}
]
# This variable contains all of the hyperparemeter information our training loop needs
optimizer = AdamW(model.parameters(),
lr = 1e-5, # args.learning_rate - default is 5e-5, our notebook had 2e-5
eps = 1e-8 # args.adam_epsilon - default is 1e-8.
)
from transformers import get_linear_schedule_with_warmup
# Number of training epochs (authors recommend between 2 and 4)
epochs = 3
# Total number of training steps is number of batches * number of epochs.
total_steps = len(train_dataloader) * epochs
# Create the learning rate scheduler.
scheduler = get_linear_schedule_with_warmup(optimizer,
num_warmup_steps = 0, # Default value in run_glue.py
num_training_steps = total_steps)
# Function to calculate the accuracy of our predictions vs labels
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
import time
import datetime
def format_time(elapsed):
'''
Takes a time in seconds and returns a string hh:mm:ss
'''
# Round to the nearest second.
elapsed_rounded = int(round((elapsed)))
# Format as hh:mm:ss
return str(datetime.timedelta(seconds=elapsed_rounded))
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
device
import random
torch.cuda.empty_cache()
# This training code is based on the `run_glue.py` script here:
# https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L128
# Set the seed value all over the place to make this reproducible.
seed_val = 44
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# Store the average loss after each epoch so we can plot them.
loss_values = []
# For each epoch...
for epoch_i in range(0, epochs):
# ========================================
# Training
# ========================================
# Perform one full pass over the training set.
print("")
print('======== Epoch {:} / {:} ========'.format(epoch_i + 1, epochs))
print('Training...')
# Measure how long the training epoch takes.
t0 = time.time()
# Reset the total loss for this epoch.
total_loss = 0
# Put the model into training mode. Don't be mislead--the call to
# `train` just changes the *mode*, it doesn't *perform* the training.
# `dropout` and `batchnorm` layers behave differently during training
# vs. test (source: https://stackoverflow.com/questions/51433378/what-does-model-train-do-in-pytorch)
model.train()
# For each batch of training data...
for step, batch in enumerate(train_dataloader):
# Progress update every 40 batches.
if step % 40 == 0 and not step == 0:
# Calculate elapsed time in minutes.
elapsed = format_time(time.time() - t0)
# Report progress.
print(' Batch {:>5,} of {:>5,}. Elapsed: {:}.'.format(step, len(train_dataloader), elapsed))
# Unpack this training batch from our dataloader.
#
# As we unpack the batch, we'll also copy each tensor to the GPU using the
# `to` method.
#
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1]: attention masks
# [2]: labels
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
# Always clear any previously calculated gradients before performing a
# backward pass. PyTorch doesn't do this automatically because
# accumulating the gradients is "convenient while training RNNs".
# (source: https://stackoverflow.com/questions/48001598/why-do-we-need-to-call-zero-grad-in-pytorch)
model.zero_grad()
# Perform a forward pass (evaluate the model on this training batch).
# This will return the loss (rather than the model output) because we
# have provided the `labels`.
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
b_labels = torch.nn.functional.one_hot(b_labels.to(torch.int64), num_classes=2)
outputs = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels.to(torch.float64))
# The call to `model` always returns a tuple, so we need to pull the
# loss value out of the tuple.
loss = outputs[0]
# Accumulate the training loss over all of the batches so that we can
# calculate the average loss at the end. `loss` is a Tensor containing a
# single value; the `.item()` function just returns the Python value
# from the tensor.
loss = loss.mean()
total_loss += loss.item()
# Perform a backward pass to calculate the gradients.
loss.backward()
# Clip the norm of the gradients to 1.0.
# This is to help prevent the "exploding gradients" problem.
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
# Update parameters and take a step using the computed gradient.
# The optimizer dictates the "update rule"--how the parameters are
# modified based on their gradients, the learning rate, etc.
optimizer.step()
# Update the learning rate.
scheduler.step()
# Calculate the average loss over the training data.
avg_train_loss = total_loss / len(train_dataloader)
# Store the loss value for plotting the learning curve.
loss_values.append(avg_train_loss)
print("")
print(" Average training loss: {0:.2f}".format(avg_train_loss))
print(" Training epcoh took: {:}".format(format_time(time.time() - t0)))
# ========================================
# Validation
# ========================================
# After the completion of each training epoch, measure our performance on
# our validation set.
print("")
print("Running Validation...")
t0 = time.time()
# Put the model in evaluation mode--the dropout layers behave differently
# during evaluation.
model.eval()
# Tracking variables
eval_loss, eval_accuracy = 0, 0
nb_eval_steps, nb_eval_examples = 0, 0
# Evaluate data for one epoch
for batch in validation_dataloader:
# Add batch to GPU
batch = tuple(t.to(device) for t in batch)
# Unpack the inputs from our dataloader
b_input_ids, b_input_mask, b_labels = batch
# Telling the model not to compute or store gradients, saving memory and
# speeding up validation
with torch.no_grad():
# Forward pass, calculate logit predictions.
# This will return the logits rather than the loss because we have
# not provided labels.
# token_type_ids is the same as the "segment ids", which
# differentiates sentence 1 and 2 in 2-sentence tasks.
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
outputs = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask)
# Get the "logits" output by the model. The "logits" are the output
# values prior to applying an activation function like the softmax.
logits = outputs[0]
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Calculate the accuracy for this batch of test sentences.
tmp_eval_accuracy = flat_accuracy(logits, label_ids)
# Accumulate the total accuracy.
eval_accuracy += tmp_eval_accuracy
# Track the number of batches
nb_eval_steps += 1
# Report the final accuracy for this validation run.
print(" Accuracy: {0:.2f}".format(eval_accuracy/nb_eval_steps))
print(" Validation took: {:}".format(format_time(time.time() - t0)))
print("")
print("Training complete!")
```
# Testing
## Helper function for predicting whether label matches the person's prediction, for one person:
```
import statistics
def predict_person(text, label, length):
# print(len(text))
split_text = text.split()
# print('split_text: ', len(split_text))
sentences = []
for i in range(0, int(len(split_text)/length)+1):
sentences.append(' '.join(split_text[0 + i*length : (i+1) * length]))
# print('sentences: ', len(sentences))
labels = [label] * len(sentences)
# print(len(labels))
# print(len(sentences))
input_ids = []
for sent in sentences:
# `encode` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
encoded_sent = tokenizer.encode(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
)
input_ids.append(encoded_sent)
input_ids = pad_sequences(input_ids, maxlen=MAX_LEN,
dtype="long", truncating="post", padding="post")
# Create attention masks
attention_masks = []
# Create a mask of 1s for each token followed by 0s for padding
for seq in input_ids:
seq_mask = [float(i>0) for i in seq]
attention_masks.append(seq_mask)
# Convert to tensors.
prediction_inputs = torch.tensor(input_ids)
prediction_masks = torch.tensor(attention_masks)
prediction_labels = torch.tensor(labels)
# Set the batch size.
batch_size = 32
# Create the DataLoader.
prediction_data = TensorDataset(prediction_inputs, prediction_masks, prediction_labels)
prediction_sampler = SequentialSampler(prediction_data)
prediction_dataloader = DataLoader(prediction_data, sampler=prediction_sampler, batch_size=batch_size)
# print('Predicting labels for {:,} test sentences...'.format(len(prediction_inputs)))
# Put model in evaluation mode
model.eval()
# Tracking variables
predictions , true_labels = [], []
# Predict
for batch in prediction_dataloader:
# Add batch to GPU
batch = tuple(t.to(device) for t in batch)
# Unpack the inputs from our dataloader
b_input_ids, b_input_mask, b_labels = batch
# Telling the model not to compute or store gradients, saving memory and
# speeding up prediction
with torch.no_grad():
# Forward pass, calculate logit predictions
outputs = model(b_input_ids, token_type_ids=None,
attention_mask=b_input_mask)
logits = outputs[0]
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Store predictions and true labels
predictions.append(logits)
true_labels.append(label_ids)
#for i in range(len(true_labels)):
# The predictions for this batch are a 2-column ndarray (one column for "0"
# and one column for "1"). Pick the label with the highest value and turn this
# in to a list of 0s and 1s.
correct = 0
num = 0
final_predict_list = []
for i in range(len(true_labels)):
pred_labels_i = np.argmax(predictions[i], axis=1).flatten()
[final_predict_list.append(i) for i in pred_labels_i]
# if accuracy_score(pred_labels_i, [0] * len(pred_labels_i)) < 0.5 :
# correct += 1
# num+=1
# print('list: ', final_predict_list)
# print(label)
try:
if (statistics.mode(final_predict_list)==label):
return 1
else:
return 0
except:
return random.randint(0,1)
```
## Final predictive accuracy score:
```
test_df
correct = 0
num = 0
for row in tqdm(range(0, len(test_df.index)), "testing.."):
pred_text = test_df.at[row, 'liked_tweets']
pred_label = test_df.at[row, target_mode]
if pred_label == 0:
#print('Skipped')
continue
num+=1
pred = predict_person(pred_text, pred_label, 512)
# print(pred)
if pred == 1:
correct+=1
print("ACC: ", correct/num)
print("ACC: ", correct/num)
num
```
## Saving model to disk
```
import os
# Saving best-practices: if you use defaults names for the model, you can reload it using from_pretrained()
output_dir = 'BERT_roles'
# Create output directory if needed
if not os.path.exists(output_dir):
os.makedirs(output_dir)
print("Saving model to %s" % output_dir)
# Save a trained model, configuration and tokenizer using `save_pretrained()`.
# They can then be reloaded using `from_pretrained()`
model_to_save = model.module if hasattr(model, 'module') else model # Take care of distributed/parallel training
model_to_save.save_pretrained(output_dir)
tokenizer.save_pretrained(output_dir)
# Good practice: save your training arguments together with the trained model
# torch.save(args, os.path.join(output_dir, 'training_args.bin'))
```
## Sandbox (the outputs from these code chunks are not up to date and were performed ad hoc)
```
raw_df = raw_df.drop_duplicates()
raw_df
raw_set = set(raw_df['liked_by'])
print(len(raw_set))
train_df, test_df = train_test_split(raw_df, stratify = raw_df['type'], random_state= 1729, test_size= 0.12)
train_set = set(edf['liked_by'])
print(len(train_set))
test_set = set(test_df['liked_by'])
print(len(test_set))
i = 0
for val in set(test_df['liked_by']):
if val in train_set:
i+=1
print(val)
print('dude wtf')
print(i)
df[df['liked_by']==856944637575090176]
testing_df[testing_df['liked_by']==856944637575090176]
predict_person(testing_df.at[45, 'text'], testing_df.at[45, 'thinking'], 256)
for row in range(300,340):
print(predict_person(testing_df.at[row, 'text'], testing_df.at[row, 'intuitive'], 256))
print(test_df.at[row, 'intuitive'])
print(" ")
# Report the number of sentences.
print('Number of persons in test set: {:,}\n'.format(test_df.shape[0]))
print('Positive samples: %d of %d (%.2f%%)' % (test_df.judging.sum(), len(test_df.judging), (test_df.judging.sum() / len(test_df.judging) * 100.0)))
# from sklearn.metrics import matthews_corrcoef
#
# matthews_set = []
#
# # Evaluate each test batch using Matthew's correlation coefficient
# print('Calculating Matthews Corr. Coef. for each batch...')
#
# # For each input batch...
# for i in range(len(true_labels)):
#
# # The predictions for this batch are a 2-column ndarray (one column for "0"
# # and one column for "1"). Pick the label with the highest value and turn this
# # in to a list of 0s and 1s.
# pred_labels_i = np.argmax(predictions[i], axis=1).flatten()
#
# # Calculate and store the coef for this batch.
# matthews = matthews_corrcoef(true_labels[i], pred_labels_i)
# matthews_set.append(matthews)
#
#matthews_set
# Combine the predictions for each batch into a single list of 0s and 1s.
# flat_predictions = [item for sublist in predictions for item in sublist]
# flat_predictions = np.argmax(flat_predictions, axis=1).flatten()
#
# # Combine the correct labels for each batch into a single list.
# flat_true_labels = [item for sublist in true_labels for item in sublist]
#
# # Calculate the MCC
# mcc = matthews_corrcoef(flat_true_labels, flat_predictions)
#
# print('MCC: %.3f' % mcc)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import seaborn as sns
from google.colab import drive
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import classification_report, plot_confusion_matrix, plot_roc_curve, accuracy_score, confusion_matrix
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
drive.mount('/content/drive', force_remount=True)
from google.colab import drive
drive.mount('/content/drive')
# define project root / change to match your path
gdrive_project_root = "drive/MyDrive/01-Education/03-PhD/2021/Courses/HCI/hci_code/"
df = pd.read_csv(gdrive_project_root + '/data/processed/cancer_data_preprocessed.csv', delimiter=',')
df.head()
list(df.columns)
X_cols = ['Study ID',
'Mutation Count',
'Overall Survival (Months)',
'Overall Survival Status',
'Number of Samples Per Patient',
'Sex',
'Somatic Status',
'Diagnosis Age',
'Additional_cancer',
'Age Group']
y_col = ['Oncotree Code']
X = df[X_cols]
y = df[y_col]
len(X.columns)
y['Oncotree Code'].unique()
x_train, x_test, y_train, y_test = train_test_split(X, y ,test_size = 0.2, random_state=42)
y_train.hist()
y_test.hist()
y_test.value_counts()
y_test['Oncotree Code'].sort_values().unique()
clf = LogisticRegression().fit(x_train, y_train)
predictions = clf.predict(x_test)
print(classification_report(y_test, predictions))
clf = RandomForestClassifier(n_estimators=1000, random_state=42)
clf.fit(x_train, y_train)
predictions = clf.predict(x_test)
# raise Exception("Takes time. Comment the line to run" )
accuracy = []
for i in range(10, 800, 10):
clf = RandomForestClassifier(n_estimators=i, random_state=42, n_jobs=-1)
clf.fit(x_train, y_train)
predictions = clf.predict(x_test)
accuracy.append(accuracy_score(y_test, predictions))
print(np.max(accuracy), " - ", np.argmax(accuracy, axis=0))
pd.Series(accuracy).plot()
clf = RandomForestClassifier(n_estimators=330, random_state=42, n_jobs=-1)
clf.fit(x_train, y_train)
predictions = clf.predict(x_test)
print(accuracy_score(y_test, predictions))
print(classification_report(y_test, predictions))
plot_confusion_matrix(clf, x_test, y_test,cmap=plt.cm.Blues)
print(clf.predict_proba([x_test.values[1]]))
print(y_test.values[1])
# pd.Series(clf.predict_proba(x_test.values[3])).plot(kind='bar')
# print(y_test.values[3])
feat_importances = pd.Series(clf.feature_importances_, index=x_train.columns)
print(feat_importances.nlargest(100).to_string())
```
# Removing Study ID
---
Not sure if Study ID is connected with the type of cancer
```
x_train, x_test, y_train, y_test = train_test_split(X[X.columns.difference(['Study ID'])], y ,test_size = 0.2, random_state=42)
clf = RandomForestClassifier(n_estimators=330, random_state=42)
clf.fit(x_train, y_train)
predictions = clf.predict(x_test)
print(accuracy_score(y_test, predictions))
print(classification_report(y_test, predictions))
# plot_confusion_matrix(clf, x_test, y_test, cmap=plt.cm.Blues)
confusion_matrix(y_test, predictions)
```
# Testing with top 6 classes
```
dft6 = pd.read_csv(gdrive_project_root + '/data/processed/cancer_data_preprocessed_top_6classes.csv', delimiter=',')
dft6.head()
Xt6 = dft6[X_cols].drop(['Study ID'], axis=1)
yt6 = dft6[y_col]
x_train, x_test, y_train, y_test = train_test_split(Xt6, yt6, test_size = 0.2, random_state=42)
yt6.hist()
y_train.hist()
y_test.hist()
clf = RandomForestClassifier(n_estimators=330, random_state=42, n_jobs=-1)
clf.fit(x_train, y_train)
predictions = clf.predict(x_test)
print(accuracy_score(y_test, predictions))
print(classification_report(y_test, predictions))
```
- 'GBM': 1,
- 'PHC': 2,
- 'AASTR': 3,
- 'ODG': 4,
- 'OAST': 5,
- 'ASTR': 6,
- 'AOAST': -1,
'PGNG': -1,
'DASTR': -1,
'AODG': -1,
'GSARC': -1,
'MNET': -1,
'DIFG': -1,
'PXA': -1,
'RGNT': -1
```
plot_confusion_matrix(clf, x_test, y_test,cmap=plt.cm.Blues, display_labels=["Other", "GBM", "PHC", "AASTR", "ODG", "OAST", "ASTR", "AOAST"])
feat_importances = pd.Series(clf.feature_importances_, index=x_train.columns)
print(feat_importances.nlargest(100).to_string())
# Grid search
from sklearn.model_selection import RandomizedSearchCV
# Number of trees in random forest
n_estimators = [int(x) for x in np.linspace(start = 200, stop = 2000, num = 10)]
# Number of features to consider at every split
max_features = ['auto', 'sqrt']
# Maximum number of levels in tree
max_depth = [int(x) for x in np.linspace(10, 110, num = 11)]
max_depth.append(None)
# Minimum number of samples required to split a node
min_samples_split = [2, 5, 10]
# Minimum number of samples required at each leaf node
min_samples_leaf = [1, 2, 4]
# Method of selecting samples for training each tree
bootstrap = [True, False]
# Create the random grid
random_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'bootstrap': bootstrap}
# Use the random grid to search for best hyperparameters
# First create the base model to tune
rf = RandomForestClassifier()
# Random search of parameters, using 3 fold cross validation,
# search across 100 different combinations, and use all available cores
rf_random = RandomizedSearchCV(estimator = rf, param_distributions = random_grid, n_iter = 100, cv = 3, verbose=2, random_state=42, n_jobs = -1)
# Fit the random search model
rf_random.fit(x_train, y_train)
rf_random.best_estimator_
'''
RandomForestClassifier(bootstrap=True, ccp_alpha=0.0, class_weight=None,
criterion='gini', max_depth=20, max_features='sqrt',
max_leaf_nodes=None, max_samples=None,
min_impurity_decrease=0.0, min_impurity_split=None,
min_samples_leaf=1, min_samples_split=10,
min_weight_fraction_leaf=0.0, n_estimators=1600,
n_jobs=None, oob_score=False, random_state=None,
verbose=0, warm_start=False)
'''
clf = RandomForestClassifier(bootstrap=True, ccp_alpha=0.0, class_weight=None,
criterion='gini', max_depth=20, max_features='sqrt',
max_leaf_nodes=None, max_samples=None,
min_impurity_decrease=0.0, min_impurity_split=None,
min_samples_leaf=1, min_samples_split=10,
min_weight_fraction_leaf=0.0, n_estimators=1600,
n_jobs=None, oob_score=False, random_state=None,
verbose=0, warm_start=False)
clf.fit(x_train, y_train)
predictions = clf.predict(x_test)
print(accuracy_score(y_test, predictions))
print(classification_report(y_test, predictions))
plot_confusion_matrix(clf, x_test, y_test,cmap=plt.cm.Blues, display_labels=["Other", "GBM", "PHC", "AASTR", "ODG", "OAST", "ASTR", "AOAST"])
### CV
from sklearn.model_selection import cross_val_score
clf = RandomForestClassifier(n_estimators=330, n_jobs=-1, random_state=42)
scores = cross_val_score(clf, X, y, cv=10)
scores
y_test.value_counts()
dft6_j6 = dft6[dft6['Oncotree Code'] != -1]
print(dft6_j6.shape)
print(dft6.shape)
Xt6_j6 = dft6_j6[X_cols].drop(['Study ID'], axis=1)
yt6_j6 = dft6_j6[y_col]
x_train, x_test, y_train, y_test = train_test_split(Xt6_j6, yt6_j6, test_size = 0.2, random_state=42)
clf = RandomForestClassifier(n_estimators=330, random_state=42, n_jobs=-1)
clf.fit(x_train, y_train)
predictions = clf.predict(x_test)
print(accuracy_score(y_test, predictions))
print(classification_report(y_test, predictions))
plot_confusion_matrix(clf, x_test, y_test,cmap=plt.cm.Blues, display_labels=["GBM", "PHC", "AASTR", "ODG", "OAST", "ASTR", "AOAST"])
```
# Top 3 classes
```
dft3 = dft6[(dft6['Oncotree Code'] > 0) & (dft6['Oncotree Code'] < 4)]
print(dft3.shape)
print(dft3.shape)
Xt3 = dft3[X_cols].drop(['Study ID'], axis=1)
yt3 = dft3[y_col]
x_train, x_test, y_train, y_test = train_test_split(Xt3, yt3, test_size = 0.2, random_state=42)
clf = RandomForestClassifier(n_estimators=330, random_state=42, n_jobs=-1)
clf.fit(x_train, y_train)
predictions = clf.predict(x_test)
print(accuracy_score(y_test, predictions))
print(classification_report(y_test, predictions))
plot_confusion_matrix(clf, x_test, y_test,cmap=plt.cm.Blues, display_labels=["GBM", "PHC", "AASTR"])
```
## Exporting model & data
```
import joblib
# gdrive_project_root + '/data/processed/cancer_data_preprocessed.csv'
joblib.dump(clf, gdrive_project_root+'models/CLF_3classes_GBM-PHC-AASTR_RandomForest_330estimators_42_random_state.pkl', compress=9)
Xt3.to_csv(gdrive_project_root + 'data/processed/3_cls_model_input/X_3cls.csv', index=False)
yt3.to_csv(gdrive_project_root + 'data/processed/3_cls_model_input/y_3cls.csv', index=False)
x_train.to_csv(gdrive_project_root + 'data/processed/3_cls_model_input/X_train_3cls.csv', index=False)
x_test.to_csv(gdrive_project_root + 'data/processed/3_cls_model_input/X_test_3cls.csv', index=False)
y_train.to_csv(gdrive_project_root + 'data/processed/3_cls_model_input/y_train_3cls.csv', index=False)
y_test.to_csv(gdrive_project_root + 'data/processed/3_cls_model_input/y_test_3cls.csv', index=False)
```
# ___Archive
```
clf = GradientBoostingClassifier(random_state=0)
clf.fit(x_train, y_train)
predictions = clf.predict(x_test)
print(classification_report(y_test, predictions))
```
| github_jupyter |
```
# from google.colab import drive
# drive.mount('/content/drive')
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]].type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx].type("torch.DoubleTensor"))
label = foreground_label[fg_idx]- fg1 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=12, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=12, out_channels=6, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=12, out_channels=32, kernel_size=3, padding=0)
self.fc1 = nn.Linear(1014, 512)
self.fc2 = nn.Linear(512, 64)
# self.fc3 = nn.Linear(512, 64)
# self.fc4 = nn.Linear(64, 10)
self.fc3 = nn.Linear(64,1)
def forward(self,z): #y is avg image #z batch of list of 9 images
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
y = y.to("cuda")
x = x.to("cuda")
for i in range(9):
x[:,i] = self.helper(z[:,i])[:,0]
x = F.softmax(x,dim=1)
x1 = x[:,0]
torch.mul(x1[:,None,None,None],z[:,0])
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],z[:,i])
return x, y
def helper(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = (F.relu(self.conv2(x)))
# print(x.shape)
# x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
# x = F.relu(self.fc3(x))
# x = F.relu(self.fc4(x))
x = self.fc3(x)
return x
focus_net = Focus().double()
focus_net = focus_net.to("cuda")
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=6, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=6, out_channels=6, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=12, out_channels=20, kernel_size=3, padding=0)
self.fc1 = nn.Linear(1014, 512)
self.fc2 = nn.Linear(512, 64)
# self.fc3 = nn.Linear(512, 64)
# self.fc4 = nn.Linear(64, 10)
self.fc3 = nn.Linear(64,3)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = (F.relu(self.conv2(x)))
# print(x.shape)
# x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
# x = F.relu(self.fc3(x))
# x = F.relu(self.fc4(x))
x = self.fc3(x)
return x
classify = Classification().double()
classify = classify.to("cuda")
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer_classify = optim.Adam(classify.parameters(), lr=0.001)#, betas=(0.9, 0.999), eps=1e-08, weight_decay=0, amsgrad=False)
optimizer_focus = optim.Adam(focus_net.parameters(), lr=0.001)#, betas=(0.9, 0.999), eps=1e-08, weight_decay=0, amsgrad=False)
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
col1.append(0)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
nos_epochs = 200
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
epoch_loss = []
cnt=0
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
# zero the parameter gradients
optimizer_focus.zero_grad()
optimizer_classify.zero_grad()
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion(outputs, labels)
loss.backward()
optimizer_focus.step()
optimizer_classify.step()
running_loss += loss.item()
mini = 60
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
if epoch % 5 == 0:
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
if(np.mean(epoch_loss) <= 0.005):
break;
if epoch % 5 == 0:
# focus_net.eval()
# classify.eval()
col1.append(epoch+1)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
#************************************************************************
#testing data set
with torch.no_grad():
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
print('Finished Training')
# torch.save(focus_net.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn_3layers/"+name+"_focus_net.pt")
# torch.save(classify.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn_3layers/"+name+"_classify.pt")
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
df_train[columns[0]] = col1
df_train[columns[1]] = col2
df_train[columns[2]] = col3
df_train[columns[3]] = col4
df_train[columns[4]] = col5
df_train[columns[5]] = col6
df_train[columns[6]] = col7
df_test[columns[0]] = col1
df_test[columns[1]] = col8
df_test[columns[2]] = col9
df_test[columns[3]] = col10
df_test[columns[4]] = col11
df_test[columns[5]] = col12
df_test[columns[6]] = col13
df_train
# plt.figure(12,12)
plt.plot(col1,col2, label='argmax > 0.5')
plt.plot(col1,col3, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.title("On Training set")
plt.show()
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.savefig("train_ftpt.pdf", bbox_inches='tight')
plt.show()
df_test
# plt.figure(12,12)
plt.plot(col1,col8, label='argmax > 0.5')
plt.plot(col1,col9, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.title("On Testing set")
plt.show()
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.savefig("test_ftpt.pdf", bbox_inches='tight')
plt.show()
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
max_alpha =[]
alpha_ftpt=[]
argmax_more_than_half=0
argmax_less_than_half=0
for i, data in enumerate(test_loader):
inputs, labels,fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
alphas, avg = focus_net(inputs)
outputs = classify(avg)
mx,_ = torch.max(alphas,1)
max_alpha.append(mx.cpu().detach().numpy())
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if (focus == fore_idx[j] and predicted[j] == labels[j]):
alpha_ftpt.append(alphas[j][focus].item())
max_alpha = np.concatenate(max_alpha,axis=0)
print(max_alpha.shape)
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(max_alpha,bins=50,color ="c")
plt.title("alpha values histogram")
plt.savefig("alpha_hist.pdf")
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(np.array(alpha_ftpt),bins=50,color ="c")
plt.title("alpha values in ftpt")
plt.savefig("alpha_hist_ftpt.pdf")
```
| github_jupyter |
# Josephson Junction QComponent Demo Notebook
This demo notebook describes two types of Josephson Junction (JJ) qcomponents available in Qiskit Metal, including a "Manhattan"-style JJ and a "Dolan"-style JJ. In addition, we demonstrate how to insert these realistic JJ structures in between the capacative pads of the transmon pocket qcomponent, forming a realistic qubit design.
Let's start to importing some key modules:
```
import qiskit_metal as metal
from qiskit_metal import designs, draw
from qiskit_metal import MetalGUI, Dict #, open_docs
```
Next, we'll fire up the Qiskit Metal GUI:
```
design = designs.DesignPlanar()
gui = MetalGUI(design)
```
Next, let's see what a Manhattan-style JJ looks like. We'll import the qcomponent and place it in our design, calling it "JJ1":
```
from qiskit_metal.qlibrary.qubits.JJ_Manhattan import jj_manhattan
design.overwrite_enabled = True
jj1 = jj_manhattan(design, 'JJ1', options=dict())
gui.rebuild()
gui.autoscale()
```
We can take a screenshot of the GUI to see what our JJ looks like up close::
```
gui.rebuild()
gui.autoscale()
gui.edit_component('JJ1')
gui.zoom_on_components(['JJ1'])
gui.highlight_components(['JJ1'])
gui.screenshot()
```
Note that you'll have to zoom in to see the JJ qcomponent since the default dimensions are quite small (micrometers instead of mm!) For the Manhattan-style junction, we see two crossing rectangles each connected to a larger rectangular pad. All of the metal is part of the same layer, as opposed to the two-layer Dolan-style design that we'll see below.
Also note that for both JJ qcomponents, the coordinates (x_pos, y_pos) define the lower-left corner of the bottom rectangle. Since the default values of these coordinates are zero, we see that the bottom left corner is at the origin.
Now, let's take a look at a Dolan-style JJ qcomponent design. We'll call this one "JJ2" and we'll place it such that the bottom left corner is at the coordinate (0.1, 0.0) so that we can look at the two qcomponents side-by-side in our design:
```
from qiskit_metal.qlibrary.qubits.JJ_Dolan import jj_dolan
design.overwrite_enabled = True
jj2 = jj_dolan(design, 'JJ2', options=dict(x_pos="0.1", y_pos="0.0"))
gui.rebuild()
gui.autoscale()
```
We can again take a GUI screenshot to see how this JJ differs from the Manhattan-style one:
```
gui.rebuild()
gui.autoscale()
gui.edit_component('JJ2')
gui.zoom_on_components(['JJ2'])
gui.highlight_components(['JJ2'])
gui.screenshot()
```
You'll notice that the Dolan-style JJ has a similar structure as the Manhattan-style JJ but the second set of rectangles is translated so that they no longer overlap with the first pair of rectangles. In addition, we see a smaller rectangle drawn which is actually defined to be on a different metal layer than the other rectangles. This reflects the fact that in Dolan-style JJ fabrication, there are two separate metal levels used in the shadow masking process.
### Inserting Josephson Junctions into a Transmon Qubit Design
In a realistic quantum chip design, the JJs would be located in between the large pads of the transmon pocket. Before we show what this would actually look like, let's delete the two JJs that we've drawn and start from scratch.
```
jj1.delete()
jj2.delete()
```
You may have to hit the "Replot" button in the GUI after deleting. Now, let's create an instance of a basic transmon pocket qcomponent, using all default settings:
```
from qiskit_metal.qlibrary.qubits.transmon_pocket import TransmonPocket
design.overwrite_enabled = True
q1 = TransmonPocket(design, 'qubit1', options=dict())
gui.rebuild()
gui.autoscale()
```
When we zoom in and see that the middle of the "JJ region" of the transmon pocket is at the origin. This is the region we will place our actual JJ qcomponent. Since we want the larger rectangular pads of the JJ qcomponent to overlap with the large rectangular pads of the transmon pocket, we'll have to pick a suitable (x,y) coordinate for the JJ. Here's an example of placing the Manhattan-style JJ in this region:
```
from qiskit_metal.qlibrary.qubits.JJ_Manhattan import jj_manhattan
design.overwrite_enabled = True
jj1 = jj_manhattan(design, 'JJ1', options=dict(x_pos="-0.015", y_pos="-0.02"))
gui.rebuild()
gui.autoscale()
```
We can take a GUI screenshot to see how the transmon qubit looks with the JJ inserted:
```
gui.rebuild()
gui.autoscale()
gui.edit_component('qubit1')
gui.zoom_on_components(['qubit1'])
gui.screenshot()
```
We can zoom in on the actual JJ to see how the pads overlap with the large rectangular pads of the transmon:
```
gui.rebuild()
gui.autoscale()
gui.edit_component('JJ1')
gui.zoom_on_components(['JJ1'])
gui.screenshot()
```
In the same way, we can place another transmon pocket at (1,1) and then place a Dolan-style JJ inside:
```
design.overwrite_enabled = True
q2 = TransmonPocket(design, 'qubit2', options=dict(pos_x="1.0"))
jj2 = jj_dolan(design, 'JJ2', options=dict(x_pos="0.985", y_pos="-0.02"))
gui.rebuild()
gui.autoscale()
```
Let's take a screenshot to see what the transmon qubit looks like with the JJ inserted:
```
gui.rebuild()
gui.autoscale()
gui.edit_component('qubit2')
gui.zoom_on_components(['qubit2'])
gui.screenshot()
```
Lastly, we can zoom in even more to see how the Dolan-style JJ looks while connected to the pads of the transmon:
```
gui.rebuild()
gui.autoscale()
gui.edit_component('JJ2')
gui.zoom_on_components(['JJ2'])
gui.screenshot()
```
| github_jupyter |
# データサイエンス100本ノック(構造化データ加工編) - SQL
## はじめに
- データベースはPostgreSQL13です
- 初めに以下のセルを実行してください
- セルに %%sql と記載することでSQLを発行することができます
- jupyterからはdescribeコマンドによるテーブル構造の確認ができないため、テーブル構造を確認する場合はlimitを指定したSELECTなどで代用してください
- 使い慣れたSQLクライアントを使っても問題ありません(接続情報は以下の通り)
- IPアドレス:Docker Desktopの場合はlocalhost、Docker toolboxの場合は192.168.99.100
- Port:5432
- database名: dsdojo_db
- ユーザ名:padawan
- パスワード:padawan12345
- 大量出力を行うとJupyterが固まることがあるため、出力件数は制限することを推奨します(設問にも出力件数を記載)
- 結果確認のために表示させる量を適切にコントロールし、作業を軽快にすすめる技術もデータ加工には求められます
- 大量結果が出力された場合は、ファイルが重くなり以降開けなくなることもあります
- その場合、作業結果は消えますがファイルをGitHubから取り直してください
- vimエディタなどで大量出力範囲を削除することもできます
- 名前、住所等はダミーデータであり、実在するものではありません
```
%load_ext sql
import os
pgconfig = {
'host': 'db',
'port': os.environ['PG_PORT'],
'database': os.environ['PG_DATABASE'],
'user': os.environ['PG_USER'],
'password': os.environ['PG_PASSWORD'],
}
dsl = 'postgresql://{user}:{password}@{host}:{port}/{database}'.format(**pgconfig)
# MagicコマンドでSQLを書くための設定
%sql $dsl
```
# 使い方
- セルの先頭に%%sqlと記載し、2行目以降にSQLを記述することでJupyterからPostgreSQLに対しSQLを実行できます。
```
%%sql
select 'このように実行できます' as sample
```
# データ加工100本ノック
---
> S-001: レシート明細テーブル(receipt)から全項目の先頭10件を表示し、どのようなデータを保有しているか目視で確認せよ。
```
%%sql
SELECT * FROM receipt LIMIT 10
```
---
> S-002: レシート明細のテーブル(receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、10件表示させよ。
```
%%sql
SELECT sales_ymd, customer_id, product_cd, amount FROM receipt LIMIT 10
```
---
> S-003: レシート明細のテーブル(receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、10件表示させよ。ただし、sales_ymdはsales_dateに項目名を変更しながら抽出すること。
```
%%sql
SELECT sales_ymd as sales_date, customer_id, product_cd, amount
FROM receipt LIMIT 10
```
---
> S-004: レシート明細のテーブル(receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
> - 顧客ID(customer_id)が"CS018205000001"
```
%%sql
SELECT
sales_ymd as sales_date, customer_id, product_cd, amount
FROM
receipt
WHERE
customer_id = 'CS018205000001'
```
---
> S-005: レシート明細のテーブル(receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
> - 顧客ID(customer_id)が"CS018205000001"
> - 売上金額(amount)が1,000以上
```
%%sql
SELECT
sales_ymd as sales_date, customer_id, product_cd, amount
FROM
receipt
WHERE
customer_id = 'CS018205000001'
and
amount >= 1000
```
---
> S-006: レシート明細テーブル(receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上数量(quantity)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
> - 顧客ID(customer_id)が"CS018205000001"
> - 売上金額(amount)が1,000以上または売上数量(quantity)が5以上
```
%%sql
SELECT
sales_ymd as sales_date, customer_id, product_cd, quantity, amount
FROM
receipt
WHERE
customer_id = 'CS018205000001'
and
(
amount >= 1000
or
quantity >= 5
)
```
---
> S-007: レシート明細のテーブル(receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
> - 顧客ID(customer_id)が"CS018205000001"
> - 売上金額(amount)が1,000以上2,000以下
```
%%sql
SELECT
sales_ymd as sales_date, customer_id, product_cd, amount
FROM
receipt
WHERE
customer_id = 'CS018205000001'
and
amount between 1000 and 2000
```
---
> S-008: レシート明細テーブル(receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
> - 顧客ID(customer_id)が"CS018205000001"
> - 商品コード(product_cd)が"P071401019"以外
```
%%sql
SELECT
sales_ymd as sales_date, customer_id, product_cd, amount
FROM
receipt
WHERE
customer_id = 'CS018205000001'
and
product_cd != 'P071401019'
```
---
> S-009: 以下の処理において、出力結果を変えずにORをANDに書き換えよ。
`select * from store where not (prefecture_cd = '13' or floor_area > 900)`
```
%%sql
SELECT * FROM store WHERE prefecture_cd != '13' and floor_area <= 900
```
---
> S-010: 店舗テーブル(store)から、店舗コード(store_cd)が"S14"で始まるものだけ全項目抽出し、10件だけ表示せよ。
```
%%sql
SELECT * FROM store WHERE store_cd like 'S14%' LIMIT 10
```
---
> S-011: 顧客テーブル(customer)から顧客ID(customer_id)の末尾が1のものだけ全項目抽出し、10件だけ表示せよ。
```
%%sql
SELECT * FROM customer WHERE customer_id like '%1' LIMIT 10
```
---
> S-012: 店舗テーブル(store)から横浜市の店舗だけ全項目表示せよ。
```
%%sql
SELECT * FROM store WHERE address LIKE '%横浜市%'
```
---
> S-013: 顧客テーブル(customer)から、ステータスコード(status_cd)の先頭がアルファベットのA〜Fで始まるデータを全項目抽出し、10件だけ表示せよ。
```
%%sql
SELECT * FROM customer WHERE status_cd ~ '^[A-F]' LIMIT 10
```
---
> S-014: 顧客テーブル(customer)から、ステータスコード(status_cd)の末尾が数字の1〜9で終わるデータを全項目抽出し、10件だけ表示せよ。
```
%%sql
SELECT * FROM customer WHERE status_cd ~ '[1-9]$' LIMIT 10
```
---
> S-015: 顧客テーブル(customer)から、ステータスコード(status_cd)の先頭がアルファベットのA〜Fで始まり、末尾が数字の1〜9で終わるデータを全項目抽出し、10件だけ表示せよ。
```
%%sql
SELECT * FROM customer WHERE status_cd ~ '^[A-F].*[1-9]$' LIMIT 10
```
---
> S-016: 店舗テーブル(store)から、電話番号(tel_no)が3桁-3桁-4桁のデータを全項目表示せよ。
```
%%sql
SELECT * FROM store WHERE tel_no ~ '^[0-9]{3}-[0-9]{3}-[0-9]{4}$'
```
---
> S-017: 顧客テーブル(customer)を生年月日(birth_day)で高齢順にソートし、先頭10件を全項目表示せよ。
```
%%sql
SELECT * from customer ORDER BY birth_day LIMIT 10
```
---
> S-018: 顧客テーブル(customer)を生年月日(birth_day)で若い順にソートし、先頭10件を全項目表示せよ。
```
%%sql
SELECT * from customer ORDER BY birth_day DESC LIMIT 10
```
---
> S-019: レシート明細テーブル(receipt)に対し、1件あたりの売上金額(amount)が高い順にランクを付与し、先頭10件を抽出せよ。項目は顧客ID(customer_id)、売上金額(amount)、付与したランクを表示させること。なお、売上金額(amount)が等しい場合は同一順位を付与するものとする。
```
%%sql
SELECT customer_id, amount, RANK() OVER(ORDER BY amount DESC) AS ranking
FROM receipt
LIMIT 10
```
---
> S-020: レシート明細テーブル(receipt)に対し、1件あたりの売上金額(amount)が高い順にランクを付与し、先頭10件を抽出せよ。項目は顧客ID(customer_id)、売上金額(amount)、付与したランクを表示させること。なお、売上金額(amount)が等しい場合でも別順位を付与すること。
```
%%sql
SELECT customer_id, amount, ROW_NUMBER() OVER(ORDER BY amount DESC) AS ranking
FROM receipt
LIMIT 10
```
---
> S-021: レシート明細テーブル(receipt)に対し、件数をカウントせよ。
```
%%sql
SELECT count(1) FROM receipt
```
---
> S-022: レシート明細テーブル(receipt)の顧客ID(customer_id)に対し、ユニーク件数をカウントせよ。
```
%%sql
SELECT count(distinct customer_id) FROM receipt
```
---
> S-023: レシート明細テーブル(receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)と売上数量(quantity)を合計せよ。
```
%%sql
SELECT store_cd
, SUM(amount) as amount
, SUM(quantity) as quantity
FROM receipt
group by store_cd
```
---
> S-024: レシート明細テーブル(receipt)に対し、顧客ID(customer_id)ごとに最も新しい売上日(sales_ymd)を求め、10件表示せよ。
```
%%sql
SELECT customer_id, MAX(sales_ymd)
FROM receipt
GROUP BY customer_id
LIMIT 10
```
---
> S-025: レシート明細テーブル(receipt)に対し、顧客ID(customer_id)ごとに最も古い売上日(sales_ymd)を求め、10件表示せよ。
```
%%sql
SELECT customer_id, MIN(sales_ymd)
FROM receipt
GROUP BY customer_id
LIMIT 10
```
---
> S-026: レシート明細テーブル(receipt)に対し、顧客ID(customer_id)ごとに最も新しい売上日(sales_ymd)と古い売上日を求め、両者が異なるデータを10件表示せよ。
```
%%sql
SELECT customer_id, MAX(sales_ymd), MIN(sales_ymd)
FROM receipt
GROUP BY customer_id
HAVING MAX(sales_ymd) != MIN(sales_ymd)
LIMIT 10
```
---
> S-027: レシート明細テーブル(receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の平均を計算し、降順でTOP5を表示せよ。
```
%%sql
SELECT store_cd, AVG(amount) as avr_amount
FROM receipt
GROUP BY store_cd
ORDER BY avr_amount DESC
LIMIT 5
```
---
> S-028: レシート明細テーブル(receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の中央値を計算し、降順でTOP5を表示せよ。
```
%%sql
SELECT
store_cd,
PERCENTILE_CONT(0.5) WITHIN GROUP(ORDER BY amount) as amount_50per
FROM receipt
GROUP BY store_cd
ORDER BY amount_50per desc
LIMIT 5
```
---
> S-029: レシート明細テーブル(receipt)に対し、店舗コード(store_cd)ごとに商品コード(product_cd)の最頻値を求めよ。
```
%%sql
-- コード例1: 分析関数でmodeを計算する
WITH product_mode AS (
SELECT store_cd,product_cd, COUNT(1) as mode_cnt,
RANK() OVER(PARTITION BY store_cd ORDER BY COUNT(1) DESC) AS rnk
FROM receipt
GROUP BY store_cd,product_cd
)
SELECT store_cd,product_cd, mode_cnt
FROM product_mode
WHERE rnk = 1
ORDER BY store_cd,product_cd;
%%sql
-- コード例2:mode()を使う簡易ケース(早いが最頻値が複数の場合は一つだけ選ばれる)
SELECT store_cd, mode() WITHIN GROUP(ORDER BY product_cd)
FROM receipt
GROUP BY store_cd
ORDER BY store_cd
```
---
> S-030: レシート明細テーブル(receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の標本分散を計算し、降順でTOP5を表示せよ。
```
%%sql
SELECT store_cd, var_samp(amount) as vars_amount
FROM receipt
GROUP BY store_cd
ORDER BY vars_amount desc
LIMIT 5
```
---
> S-031: レシート明細テーブル(receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の標本標準偏差を計算し、降順でTOP5を表示せよ。
```
%%sql
SELECT store_cd, stddev_samp(amount) as stds_amount
FROM receipt
GROUP BY store_cd
ORDER BY stds_amount desc
LIMIT 5
```
---
> S-032: レシート明細テーブル(receipt)の売上金額(amount)について、25%刻みでパーセンタイル値を求めよ。
```
%%sql
SELECT
PERCENTILE_CONT(0.25) WITHIN GROUP(ORDER BY amount) as amount_25per,
PERCENTILE_CONT(0.50) WITHIN GROUP(ORDER BY amount) as amount_50per,
PERCENTILE_CONT(0.75) WITHIN GROUP(ORDER BY amount) as amount_75per,
PERCENTILE_CONT(1.0) WITHIN GROUP(ORDER BY amount) as amount_100per
FROM receipt
```
---
> S-033: レシート明細テーブル(receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の平均を計算し、330以上のものを抽出せよ。
```
%%sql
SELECT store_cd, AVG(amount) as avg_amount
FROM receipt
GROUP BY store_cd
HAVING AVG(amount) >= 330
```
---
> S-034: レシート明細テーブル(receipt)に対し、顧客ID(customer_id)ごとに売上金額(amount)を合計して全顧客の平均を求めよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。
```
%%sql
WITH customer_amount AS (
SELECT customer_id, SUM(amount) AS sum_amount
FROM receipt
WHERE customer_id not like 'Z%'
GROUP BY customer_id
)
SELECT AVG(sum_amount) from customer_amount
```
---
> S-035: レシート明細テーブル(receipt)に対し、顧客ID(customer_id)ごとに売上金額(amount)を合計して全顧客の平均を求め、平均以上に買い物をしている顧客を抽出せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。なお、データは10件だけ表示させれば良い。
```
%%sql
WITH customer_amount AS (
SELECT customer_id, SUM(amount) AS sum_amount
FROM receipt
WHERE customer_id not like 'Z%'
GROUP BY customer_id
)
SELECT customer_id, sum_amount
FROM customer_amount
WHERE sum_amount >= (SELECT AVG(sum_amount) from customer_amount)
limit 10
```
---
> S-036: レシート明細テーブル(receipt)と店舗テーブル(store)を内部結合し、レシート明細テーブルの全項目と店舗テーブルの店舗名(store_name)を10件表示させよ。
```
%%sql
SELECT r.*, s.store_name
FROM receipt r
JOIN store s
ON r.store_cd = s.store_cd
LIMIT 10
```
---
> S-037: 商品テーブル(product)とカテゴリテーブル(category)を内部結合し、商品テーブルの全項目とカテゴリテーブルの小区分名(category_small_name)を10件表示させよ。
```
%%sql
SELECT p.*, c.category_small_name
FROM product p
JOIN category c
on p.category_small_cd = c.category_small_cd
LIMIT 10
```
---
> S-038: 顧客テーブル(customer)とレシート明細テーブル(receipt)から、各顧客ごとの売上金額合計を求めよ。ただし、買い物の実績がない顧客については売上金額を0として表示させること。また、顧客は性別コード(gender_cd)が女性(1)であるものを対象とし、非会員(顧客IDが'Z'から始まるもの)は除外すること。なお、結果は10件だけ表示させれば良い。
```
%%sql
WITH customer_amount AS (
SELECT customer_id, SUM(amount) AS sum_amount
FROM receipt
GROUP BY customer_id
)
SELECT c.customer_id, COALESCE(a.sum_amount,0)
FROM customer c
LEFT JOIN customer_amount a
ON c.customer_id = a.customer_id
WHERE c.gender_cd = '1'
and c.customer_id not like 'Z%'
LIMIT 10
```
---
> S-039: レシート明細テーブル(receipt)から売上日数の多い顧客の上位20件と、売上金額合計の多い顧客の上位20件を抽出し、完全外部結合せよ。ただし、非会員(顧客IDが'Z'から始まるもの)は除外すること。
```
%%sql
WITH customer_days AS (
select customer_id, count(distinct sales_ymd) come_days
FROM receipt
WHERE customer_id NOT LIKE 'Z%'
GROUP BY customer_id
ORDER BY come_days DESC LIMIT 20
),
customer_amount AS (
SELECT customer_id, sum(amount) buy_amount
FROM receipt
WHERE customer_id NOT LIKE 'Z%'
GROUP BY customer_id
ORDER BY buy_amount DESC LIMIT 20
)
SELECT COALESCE(d.customer_id, a.customer_id), d.come_days, a.buy_amount
FROM customer_days d
FULL JOIN customer_amount a
ON d.customer_id = a.customer_id;
```
---
> S-040: 全ての店舗と全ての商品を組み合わせると何件のデータとなるか調査したい。店舗(store)と商品(product)を直積した件数を計算せよ。
```
%%sql
SELECT COUNT(1) FROM store CROSS JOIN product;
```
---
> S-041: レシート明細テーブル(receipt)の売上金額(amount)を日付(sales_ymd)ごとに集計し、前日からの売上金額増減を計算せよ。なお、計算結果は10件表示すればよい。
```
%%sql
WITH sales_amount_by_date AS (
SELECT sales_ymd, SUM(amount) as amount FROM receipt
GROUP BY sales_ymd
ORDER BY sales_ymd
)
SELECT sales_ymd, LAG(sales_ymd, 1) OVER(ORDER BY sales_ymd) lag_ymd,
amount,
LAG(amount, 1) OVER(ORDER BY sales_ymd) as lag_amount,
amount - LAG(amount, 1) OVER(ORDER BY sales_ymd) as diff_amount
FROM sales_amount_by_date
LIMIT 10;
```
---
> S-042: レシート明細テーブル(receipt)の売上金額(amount)を日付(sales_ymd)ごとに集計し、各日付のデータに対し、1日前、2日前、3日前のデータを結合せよ。結果は10件表示すればよい。
```
%%sql
-- コード例1:縦持ちケース
WITH sales_amount_by_date AS (
SELECT sales_ymd, SUM(amount) as amount FROM receipt
GROUP BY sales_ymd
ORDER BY sales_ymd
),
sales_amount_lag_date AS (
SELECT sales_ymd,
COALESCE(LAG(sales_ymd, 3) OVER (ORDER BY sales_ymd),
MIN(sales_ymd) OVER (PARTITION BY NULL)) as lag_date_3,
amount
FROM sales_amount_by_date
)
SELECT a.sales_ymd, b.sales_ymd as lag_ymd,
a.amount as amount, b.amount as lag_amount
FROM sales_amount_lag_date a
JOIN sales_amount_lag_date b
ON b.sales_ymd >= a.lag_date_3
and b.sales_ymd < a.sales_ymd
ORDER BY sales_ymd, lag_ymd
LIMIT 10;
%%sql
-- コード例2:横持ちケース
WITH sales_amount_by_date AS (
SELECT sales_ymd, SUM(amount) as amount FROM receipt
GROUP BY sales_ymd
ORDER BY sales_ymd
), sales_amount_with_lag AS(
SELECT sales_ymd, amount,
LAG(sales_ymd, 1) OVER (ORDER BY sales_ymd) as lag_ymd_1,
LAG(amount, 1) OVER (ORDER BY sales_ymd) as lag_amount_1,
LAG(sales_ymd, 2) OVER (ORDER BY sales_ymd) as lag_ymd_2,
LAG(amount, 2) OVER (ORDER BY sales_ymd) as lag_amount_2,
LAG(sales_ymd, 3) OVER (ORDER BY sales_ymd) as lag_ymd_3,
LAG(amount, 3) OVER (ORDER BY sales_ymd) as lag_amount_3
FROM sales_amount_by_date
)
SELECT * FROM sales_amount_with_lag
WHERE lag_ymd_3 IS NOT NULL
ORDER BY sales_ymd
LIMIT 10;
```
---
> S-043: レシート明細テーブル(receipt)と顧客テーブル(customer)を結合し、性別(gender)と年代(ageから計算)ごとに売上金額(amount)を合計した売上サマリテーブル(sales_summary)を作成せよ。性別は0が男性、1が女性、9が不明を表すものとする。
>
>ただし、項目構成は年代、女性の売上金額、男性の売上金額、性別不明の売上金額の4項目とすること(縦に年代、横に性別のクロス集計)。また、年代は10歳ごとの階級とすること。
```
%%sql
-- SQL向きではないため、やや強引に記載する(カテゴリ数が多いときはとても長いSQLとなってしまう点に注意)
DROP TABLE IF EXISTS sales_summary;
CREATE TABLE sales_summary AS
WITH gender_era_amount AS (
SELECT c.gender_cd,
TRUNC(age/ 10) * 10 AS era,
SUM(r.amount) AS amount
FROM customer c
JOIN receipt r
ON c.customer_id = r.customer_id
GROUP BY c.gender_cd, era
)
select era,
MAX(CASE gender_cd WHEN '0' THEN amount ELSE 0 END) AS male ,
MAX(CASE gender_cd WHEN '1' THEN amount ELSE 0 END) AS female,
MAX(CASE gender_cd WHEN '9' THEN amount ELSE 0 END) AS unknown
FROM gender_era_amount
GROUP BY era
ORDER BY era
;
%%sql
SELECT * FROM sales_summary;
```
---
> S-044: 前設問で作成した売上サマリテーブル(sales_summary)は性別の売上を横持ちさせたものであった。このテーブルから性別を縦持ちさせ、年代、性別コード、売上金額の3項目に変換せよ。ただし、性別コードは男性を'00'、女性を'01'、不明を'99'とする。
```
%%sql
-- SQL向きではないため、やや強引に記載する(カテゴリ数が多いときはとても長いSQLとなってしまう点に注意)
SELECT era, '00' as gender_cd , male AS amount FROM sales_summary
UNION ALL
SELECT era, '01' as gender_cd, female AS amount FROM sales_summary
UNION ALL
SELECT era, '99' as gender_cd, unknown AS amount FROM sales_summary
```
---
> S-045: 顧客テーブル(customer)の生年月日(birth_day)は日付型でデータを保有している。これをYYYYMMDD形式の文字列に変換し、顧客ID(customer_id)とともに抽出せよ。データは10件を抽出すれば良い。
```
%%sql
SELECT customer_id, TO_CHAR(birth_day, 'YYYYMMDD') FROM customer LIMIT 10;
```
---
> S-046: 顧客テーブル(customer)の申し込み日(application_date)はYYYYMMDD形式の文字列型でデータを保有している。これを日付型に変換し、顧客ID(customer_id)とともに抽出せよ。データは10件を抽出すれば良い。
```
%%sql
SELECT customer_id, TO_DATE(application_date, 'YYYYMMDD')
FROM customer LIMIT 10;
```
---
> S-047: レシート明細テーブル(receipt)の売上日(sales_ymd)はYYYYMMDD形式の数値型でデータを保有している。これを日付型に変換し、レシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。データは10件を抽出すれば良い。
```
%%sql
SELECT
TO_DATE(CAST(sales_ymd AS VARCHAR), 'YYYYMMDD'),
receipt_no,
receipt_sub_no
FROM receipt
LIMIT 10;
```
---
> S-048: レシート明細テーブル(receipt)の売上エポック秒(sales_epoch)は数値型のUNIX秒でデータを保有している。これを日付型に変換し、レシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。データは10件を抽出すれば良い。
```
%%sql
SELECT
TO_TIMESTAMP(sales_epoch) as sales_date,
receipt_no, receipt_sub_no
FROM receipt
LIMIT 10;
```
---
> S-049: レシート明細テーブル(receipt)の販売エポック秒(sales_epoch)を日付型に変換し、「年」だけ取り出してレシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。データは10件を抽出すれば良い。
```
%%sql
SELECT
TO_CHAR(EXTRACT(YEAR FROM TO_TIMESTAMP(sales_epoch)),'FM9999') as sales_year,
receipt_no,
receipt_sub_no
FROM receipt
LIMIT 10;
```
---
> S-050: レシート明細テーブル(receipt)の売上エポック秒(sales_epoch)を日付型に変換し、「月」だけ取り出してレシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。なお、「月」は0埋め2桁で取り出すこと。データは10件を抽出すれば良い。
```
%%sql
SELECT
TO_CHAR(EXTRACT(
MONTH FROM TO_TIMESTAMP(sales_epoch)
), 'FM00') as sales_month,
receipt_no, receipt_sub_no
FROM receipt LIMIT 10;
```
---
> S-051: レシート明細テーブル(receipt)の売上エポック秒を日付型に変換し、「日」だけ取り出してレシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。なお、「日」は0埋め2桁で取り出すこと。データは10件を抽出すれば良い。
```
%%sql
SELECT
receipt_no, receipt_sub_no,
TO_CHAR(EXTRACT(DAY FROM TO_TIMESTAMP(sales_epoch)), 'FM00') as sales_day
FROM receipt LIMIT 10;
```
---
> S-052: レシート明細テーブル(receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計の上、売上金額合計に対して2,000円以下を0、2,000円より大きい金額を1に2値化し、顧客ID、売上金額合計とともに10件表示せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。
```
%%sql
SELECT
customer_id,
SUM(amount) AS sum_amount,
CASE
WHEN SUM(amount) > 2000 THEN 1
WHEN SUM(amount) <= 2000 THEN 0
END as amount_flg
FROM receipt
WHERE customer_id not like 'Z%'
GROUP BY customer_id
LIMIT 10
```
---
> S-053: 顧客テーブル(customer)の郵便番号(postal_cd)に対し、東京(先頭3桁が100〜209のもの)を1、それ以外のものを0に2値化せよ。さらにレシート明細テーブル(receipt)と結合し、全期間において買い物実績のある顧客数を、作成した2値ごとにカウントせよ。
```
%%sql
WITH cust AS (
SELECT
customer_id,
postal_cd,
CASE
WHEN 100 <= CAST(SUBSTR(postal_cd, 1, 3) AS INTEGER)
AND CAST(SUBSTR(postal_cd, 1, 3) AS INTEGER) <= 209 THEN 1
ELSE 0
END AS postal_flg
FROM customer
),
rect AS(
SELECT
customer_id,
SUM(amount)
FROM
receipt
GROUP BY
customer_id
)
SELECT
c.postal_flg, count(1)
FROM
rect r
JOIN
cust c
ON
r.customer_id = c.customer_id
GROUP BY
c.postal_flg
```
---
> S-054: 顧客テーブル(customer)の住所(address)は、埼玉県、千葉県、東京都、神奈川県のいずれかとなっている。都道府県毎にコード値を作成し、顧客ID、住所とともに抽出せよ。値は埼玉県を11、千葉県を12、東京都を13、神奈川県を14とすること。結果は10件表示させれば良い。
```
%%sql
-- SQL向きではないため、やや強引に記載する(カテゴリ数が多いときはとても長いSQLとなってしまう点に注意)
SELECT
customer_id,
-- 確認用に住所も表示
address,
CASE SUBSTR(address,1, 3)
WHEN '埼玉県' THEN '11'
WHEN '千葉県' THEN '12'
WHEN '東京都' THEN '13'
WHEN '神奈川' THEN '14'
END AS prefecture_cd
FROM
customer
LIMIT 10
```
---
> S-055: レシート明細テーブル(receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、その合計金額の四分位点を求めよ。その上で、顧客ごとの売上金額合計に対して以下の基準でカテゴリ値を作成し、顧客ID、売上金額合計とともに表示せよ。カテゴリ値は上から順に1〜4とする。結果は10件表示させれば良い。
>
> - 最小値以上第一四分位未満
> - 第一四分位以上第二四分位未満
> - 第二四分位以上第三四分位未満
> - 第三四分位以上
```
%%sql
WITH sales_amount AS(
SELECT
customer_id,
SUM(amount) as sum_amount
FROM
receipt
GROUP BY
customer_id
),
sales_pct AS (
SELECT
PERCENTILE_CONT(0.25) WITHIN GROUP(ORDER BY sum_amount) AS pct25,
PERCENTILE_CONT(0.50) WITHIN GROUP(ORDER BY sum_amount) AS pct50,
PERCENTILE_CONT(0.75) WITHIN GROUP(ORDER BY sum_amount) AS pct75
FROM
sales_amount
)
SELECT
a.customer_id,
a.sum_amount,
CASE
WHEN a.sum_amount < pct25 THEN 1
WHEN pct25 <= a.sum_amount and a.sum_amount < pct50 THEN 2
WHEN pct50 <= a.sum_amount and a.sum_amount < pct75 THEN 3
WHEN pct75 <= a.sum_amount THEN 4
END as pct_flg
FROM sales_amount a
CROSS JOIN sales_pct p
LIMIT 10
```
---
> S-056: 顧客テーブル(customer)の年齢(age)をもとに10歳刻みで年代を算出し、顧客ID(customer_id)、生年月日(birth_day)とともに抽出せよ。ただし、60歳以上は全て60歳代とすること。年代を表すカテゴリ名は任意とする。先頭10件を表示させればよい。
```
%%sql
SELECT
customer_id,
birth_day,
LEAST(CAST(TRUNC(age / 10) * 10 AS INTEGER), 60) AS era
FROM
customer
GROUP BY
customer_id,
birth_day
-- 確認用の条件
--HAVING LEAST(CAST(TRUNC(age / 10) * 10 AS INTEGER), 60) >= 60
LIMIT 10
```
---
> S-057: 前問題の抽出結果と性別(gender)を組み合わせ、新たに性別×年代の組み合わせを表すカテゴリデータを作成せよ。組み合わせを表すカテゴリの値は任意とする。先頭10件を表示させればよい。
```
%%sql
SELECT
customer_id,
birth_day,
gender_cd || LEAST(CAST(TRUNC(age / 10) * 10 AS INTEGER), 60) AS era
FROM
customer
GROUP BY
customer_id,
birth_day
HAVING LEAST(CAST(TRUNC(age / 10) * 10 AS INTEGER), 60) >= 60
LIMIT 10
```
---
> S-058: 顧客テーブル(customer)の性別コード(gender_cd)をダミー変数化し、顧客ID(customer_id)とともに抽出せよ。結果は10件表示させれば良い。
```
%%sql
-- SQL向きではないため、やや強引に記載する(カテゴリ数が多いときはとても長いSQLとなってしまう点に注意)
SELECT
customer_id,
CASE WHEN gender_cd = '0' THEN '1' ELSE '0' END AS gender_male,
CASE WHEN gender_cd = '1' THEN '1' ELSE '0' END AS gender_female,
CASE WHEN gender_cd = '9' THEN '1' ELSE '0' END AS gender_unknown
FROM
customer
LIMIT 10
```
---
> S-059: レシート明細テーブル(receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を平均0、標準偏差1に標準化して顧客ID、売上金額合計とともに表示せよ。標準化に使用する標準偏差は、不偏標準偏差と標本標準偏差のどちらでも良いものとする。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。結果は10件表示させれば良い。
```
%%sql
WITH sales_amount AS(
SELECT
customer_id,
SUM(amount) as sum_amount
FROM
receipt
WHERE
customer_id NOT LIKE 'Z%'
GROUP BY
customer_id
),
stats_amount AS (
SELECT
AVG(sum_amount) as avg_amount,
stddev_samp(sum_amount) as std_amount
FROM
sales_amount
)
SELECT
customer_id,
sum_amount,
(sum_amount - avg_amount) / std_amount as normal_amount
FROM sales_amount
CROSS JOIN stats_amount
LIMIT 10
```
---
> S-060: レシート明細テーブル(receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を最小値0、最大値1に正規化して顧客ID、売上金額合計とともに表示せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。結果は10件表示させれば良い。
```
%%sql
WITH sales_amount AS(
SELECT
customer_id,
SUM(amount) as sum_amount
FROM
receipt
WHERE
customer_id NOT LIKE 'Z%'
GROUP BY
customer_id
),
stats_amount AS (
SELECT
max(sum_amount) as max_amount,
min(sum_amount) as min_amount
FROM
sales_amount
)
SELECT
customer_id,
sum_amount,
(sum_amount - min_amount) * 1.0
/ (max_amount - min_amount) * 1.0 AS scale_amount
FROM sales_amount
CROSS JOIN stats_amount
LIMIT 10
```
---
> S-061: レシート明細テーブル(receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を常用対数化(底=10)して顧客ID、売上金額合計とともに表示せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。結果は10件表示させれば良い。
```
%%sql
SELECT
customer_id,
SUM(amount),
LOG(SUM(amount) + 1) as log_amount
FROM
receipt
WHERE
customer_id NOT LIKE 'Z%'
GROUP BY
customer_id
LIMIT 10
```
---
> S-062: レシート明細テーブル(receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を自然対数化(底=e)して顧客ID、売上金額合計とともに表示せよ(ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること)。結果は10件表示させれば良い。
```
%%sql
SELECT
customer_id,
SUM(amount),
LN(SUM(amount) + 1) as log_amount
FROM
receipt
WHERE
customer_id NOT LIKE 'Z%'
GROUP BY
customer_id
LIMIT 10
```
---
> S-063: 商品テーブル(product)の単価(unit_price)と原価(unit_cost)から、各商品の利益額を算出せよ。結果は10件表示させれば良い。
```
%%sql
SELECT
product_cd,
unit_price,
unit_cost,
unit_price - unit_cost as unit_profit
FROM
product
LIMIT 10
```
---
> S-064: 商品テーブル(product)の単価(unit_price)と原価(unit_cost)から、各商品の利益率の全体平均を算出せよ。 ただし、単価と原価にはNULLが存在することに注意せよ。
```
%%sql
SELECT
AVG((unit_price * 1.0 - unit_cost) / unit_price) as unit_profit_rate
FROM
product
LIMIT 10
```
---
> S-065: 商品テーブル(product)の各商品について、利益率が30%となる新たな単価を求めよ。ただし、1円未満は切り捨てること。そして結果を10件表示させ、利益率がおよそ30%付近であることを確認せよ。ただし、単価(unit_price)と原価(unit_cost)にはNULLが存在することに注意せよ。
```
%%sql
SELECT
product_cd,
unit_price,
unit_cost,
TRUNC(unit_cost / 0.7) as new_price,
((TRUNC(unit_cost / 0.7) - unit_cost)
/ TRUNC(unit_cost / 0.7)) as new_profit
FROM
product
LIMIT 10
```
---
> S-066: 商品テーブル(product)の各商品について、利益率が30%となる新たな単価を求めよ。今回は、1円未満を四捨五入すること。そして結果を10件表示させ、利益率がおよそ30%付近であることを確認せよ。ただし、単価(unit_price)と原価(unit_cost)にはNULLが存在することに注意せよ。
```
%%sql
SELECT ROUND(2.5)
%%sql
SELECT
product_cd,
unit_price,
unit_cost,
ROUND(unit_cost / 0.7) as new_price,
((ROUND(unit_cost / 0.7) - unit_cost)
/ ROUND(unit_cost / 0.7)) as new_profit
FROM
product
LIMIT 10
```
---
> S-067: 商品テーブル(product)の各商品について、利益率が30%となる新たな単価を求めよ。今回は、1円未満を切り上げること。そして結果を10件表示させ、利益率がおよそ30%付近であることを確認せよ。ただし、単価(unit_price)と原価(unit_cost)にはNULLが存在することに注意せよ。
```
%%sql
SELECT
product_cd,
unit_price,
unit_cost,
CEIL(unit_cost / 0.7) as new_price,
((CEIL(unit_cost / 0.7) - unit_cost) / CEIL(unit_cost / 0.7)) as new_profit
FROM
product
LIMIT 10
```
---
> S-068: 商品テーブル(product)の各商品について、消費税率10%の税込み金額を求めよ。 1円未満の端数は切り捨てとし、結果は10件表示すれば良い。ただし、単価(unit_price)にはNULLが存在することに注意せよ。
```
%%sql
SELECT
product_cd,
unit_price,
TRUNC(unit_price * 1.1) as tax_price
FROM
product
LIMIT 10
```
---
> S-069: レシート明細テーブル(receipt)と商品テーブル(product)を結合し、顧客毎に全商品の売上金額合計と、カテゴリ大区分(category_major_cd)が"07"(瓶詰缶詰)の売上金額合計を計算の上、両者の比率を求めよ。抽出対象はカテゴリ大区分"07"(瓶詰缶詰)の購入実績がある顧客のみとし、結果は10件表示させればよい。
```
%%sql
WITH amount_all AS(
SELECT
customer_id,
sum(amount) AS sum_all
FROM
receipt
GROUP BY
customer_id
),
amount_07 AS (
SELECT
r.customer_id,
sum(r.amount) AS sum_07
FROM
receipt r
JOIN
product p
ON
r.product_cd = p.product_cd
and p.category_major_cd = '07'
GROUP BY
customer_id
)
SELECT
amount_all.customer_id,
sum_all,
sum_07,
sum_07 * 1.0 / sum_all as sales_rate
FROM
amount_all
JOIN
amount_07
ON
amount_all.customer_id = amount_07.customer_id
LIMIT 10
```
---
> S-070: レシート明細テーブル(receipt)の売上日(sales_ymd)に対し、顧客テーブル(customer)の会員申込日(application_date)からの経過日数を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。
```
%%sql
WITH receit_distinct AS (
SELECT distinct
customer_id,
sales_ymd
FROM
receipt
)
SELECT
c.customer_id,
r.sales_ymd,
c.application_date,
EXTRACT(DAY FROM (TO_TIMESTAMP(CAST(r.sales_ymd AS VARCHAR), 'YYYYMMDD')
- TO_TIMESTAMP(c.application_date, 'YYYYMMDD'))) AS elapsed_days
FROM
receit_distinct r
JOIN
customer c
ON
r.customer_id = c.customer_id
LIMIT 10
```
---
> S-071: レシート明細テーブル(receipt)の売上日(sales_ymd)に対し、顧客テーブル(customer)の会員申込日(application_date)からの経過月数を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。1ヶ月未満は切り捨てること。
```
%%sql
WITH receit_distinct AS (
SELECT distinct
customer_id,
sales_ymd
FROM
receipt
),
time_age_tbl AS(
SELECT
c.customer_id,
r.sales_ymd,
c.application_date,
AGE(TO_TIMESTAMP(CAST(r.sales_ymd AS VARCHAR), 'YYYYMMDD'),
TO_TIMESTAMP(c.application_date, 'YYYYMMDD')) AS time_age
FROM
receit_distinct r
JOIN
customer c
ON
r.customer_id = c.customer_id
)
SELECT
customer_id,
sales_ymd, application_date,
extract(year from time_age) * 12
+ extract(month from time_age) AS elapsed_months
FROM
time_age_tbl
LIMIT 10
```
---
> S-072: レシート明細テーブル(receipt)の売上日(sales_ymd)に対し、顧客テーブル(customer)の会員申込日(application_date)からの経過年数を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。1年未満は切り捨てること。
```
%%sql
WITH receit_distinct AS (
SELECT distinct
customer_id,
sales_ymd
FROM
receipt
)
SELECT
c.customer_id,
r.sales_ymd,
c.application_date,
EXTRACT(YEAR FROM AGE(
TO_TIMESTAMP(CAST(r.sales_ymd AS VARCHAR), 'YYYYMMDD'),
TO_TIMESTAMP(c.application_date, 'YYYYMMDD'))) AS elapsed_years
FROM
receit_distinct r
JOIN
customer c
ON
r.customer_id = c.customer_id
LIMIT 10
```
---
> S-073: レシート明細テーブル(receipt)の売上日(sales_ymd)に対し、顧客テーブル(customer)の会員申込日(application_date)からのエポック秒による経過時間を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。なお、時間情報は保有していないため各日付は0時0分0秒を表すものとする。
```
%%sql
WITH receit_distinct AS (
SELECT distinct
customer_id,
sales_ymd
FROM
receipt
)
SELECT
c.customer_id,
r.sales_ymd,
c.application_date,
EXTRACT(
EPOCH FROM TO_TIMESTAMP(CAST(r.sales_ymd AS VARCHAR), 'YYYYMMDD'))
- EXTRACT(
EPOCH FROM TO_TIMESTAMP(c.application_date, 'YYYYMMDD')
) AS elapsed_epoch
FROM
receit_distinct r
JOIN
customer c
ON
r.customer_id = c.customer_id
LIMIT 10
```
---
> S-074: レシート明細テーブル(receipt)の売上日(sales_ymd)に対し、当該週の月曜日からの経過日数を計算し、売上日、当該週の月曜日付とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値でデータを保持している点に注意)。
```
%%sql
SELECT
customer_id,
TO_DATE(CAST(sales_ymd AS VARCHAR), 'YYYYMMDD'),
EXTRACT(DOW FROM (
TO_DATE(CAST(sales_ymd AS VARCHAR), 'YYYYMMDD') - 1)) AS elapsed_years,
TO_DATE(CAST(sales_ymd AS VARCHAR), 'YYYYMMDD')
- CAST(EXTRACT(
DOW FROM (TO_DATE(CAST(sales_ymd AS VARCHAR), 'YYYYMMDD') - 1)
) AS INTEGER) AS monday
FROM
receipt
LIMIT 10
```
---
> S-075: 顧客テーブル(customer)からランダムに1%のデータを抽出し、先頭から10件データを抽出せよ。
```
%%sql
-- コード例1(シンプルにやるなら)
SELECT * FROM customer WHERE RANDOM() <= 0.01
LIMIT 10
%%sql
-- コード例2(丁寧にやるなら)
WITH customer_tmp AS(
SELECT
*
,ROW_NUMBER() OVER() as row
,COUNT(*) OVER() as count
FROM customer
ORDER BY random()
)
SELECT
customer_id
,customer_name
,gender_cd
,gender
,birth_day
,age
,postal_cd
,address
,application_store_cd
,application_date
,status_cd
FROM customer_tmp
WHERE row < count * 0.01
LIMIT 10
```
---
> S-076: 顧客テーブル(customer)から性別(gender_cd)の割合に基づきランダムに10%のデータを層化抽出し、性別ごとに件数を集計せよ。
```
%%sql
-- カテゴリ数が少ない場合はそれぞれサンプリングしUNIONするほうが簡単だが、カテゴリ数が多いケースを考慮して以下のSQLとした
-- RANDOMでORDER BYしているため、大量データを扱う場合は注意が必要
WITH cusotmer_random AS (
SELECT customer_id, g_cd, cnt
FROM (
SELECT
ARRAY_AGG(customer ORDER BY RANDOM()) AS customer_r,
gender_cd as g_cd, count(1) as cnt
FROM
customer
GROUP BY gender_cd
)sample, UNNEST(customer_r)
),
cusotmer_rownum AS(
SELECT * , ROW_NUMBER() OVER(PARTITION BY g_cd) AS rn FROM cusotmer_random
)
SELECT
g_cd,
count(1)
FROM
cusotmer_rownum
WHERE rn <= cnt * 0.1
GROUP BY g_cd
```
---
> S-077: レシート明細テーブル(receipt)の売上金額(amount)を顧客単位に合計し、合計した売上金額の外れ値を抽出せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。なお、ここでは外れ値を平均から3σ以上離れたものとする。結果は10件表示させれば良い。
```
%%sql
WITH sales_amount AS(
SELECT customer_id, SUM(amount) AS sum_amount
FROM receipt
WHERE customer_id NOT LIKE 'Z%'
GROUP BY customer_id
)
SELECT customer_id, sum_amount
FROM sales_amount
CROSS JOIN (
SELECT AVG(sum_amount) AS avg_amount, STDDEV_SAMP(sum_amount) AS std_amount
FROM sales_amount
) stats_amount
WHERE ABS(sum_amount - avg_amount) / std_amount > 3
LIMIT 10
```
---
> S-078: レシート明細テーブル(receipt)の売上金額(amount)を顧客単位に合計し、合計した売上金額の外れ値を抽出せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。なお、ここでは外れ値を第一四分位と第三四分位の差であるIQRを用いて、「第一四分位数-1.5×IQR」よりも下回るもの、または「第三四分位数+1.5×IQR」を超えるものとする。結果は10件表示させれば良い。
```
%%sql
WITH sales_amount AS(
SELECT customer_id, SUM(amount) AS sum_amount
FROM receipt
WHERE customer_id NOT LIKE 'Z%'
GROUP BY customer_id
)
SELECT customer_id, sum_amount
FROM sales_amount
CROSS JOIN (
SELECT
PERCENTILE_CONT(0.25) WITHIN GROUP(ORDER BY sum_amount) as amount_25per,
PERCENTILE_CONT(0.75) WITHIN GROUP(ORDER BY sum_amount) as amount_75per
FROM sales_amount
) stats_amount
WHERE sum_amount < amount_25per - (amount_75per - amount_25per) * 1.5
OR amount_75per + (amount_75per - amount_25per) * 1.5 < sum_amount
LIMIT 10
```
---
> S-079: 商品テーブル(product)の各項目に対し、欠損数を確認せよ。
```
%%sql
SELECT
SUM(
CASE WHEN product_cd IS NULL THEN 1 ELSE 0 END
) AS product_cd,
SUM(
CASE WHEN category_major_cd IS NULL THEN 1 ELSE 0 END
) AS category_major_cd,
SUM(
CASE WHEN category_medium_cd IS NULL THEN 1 ELSE 0 END
) AS category_medium_cd,
SUM(
CASE WHEN category_small_cd IS NULL THEN 1 ELSE 0 END
) AS category_small_cd,
SUM(
CASE WHEN unit_price IS NULL THEN 1 ELSE 0 END
) AS unit_price,
SUM(
CASE WHEN unit_cost IS NULL THEN 1 ELSE 0 END
) AS unit_cost
FROM product LIMIT 10
```
---
> S-080: 商品テーブル(product)のいずれかの項目に欠損が発生しているレコードを全て削除した新たなproduct_1を作成せよ。なお、削除前後の件数を表示させ、前設問で確認した件数だけ減少していることも確認すること。
```
%%sql
SELECT COUNT(1) FROM product;
%%sql
DROP TABLE IF EXISTS product_1;
CREATE TABLE product_1 AS (
SELECT * FROM product
WHERE unit_price IS NOT NULL OR unit_cost IS NOT NULL
);
SELECT COUNT(1) FROM product_1;
```
---
> S-081: 単価(unit_price)と原価(unit_cost)の欠損値について、それぞれの平均値で補完した新たなproduct_2を作成せよ。なお、平均値について1円未満は四捨五入とする。補完実施後、各項目について欠損が生じていないことも確認すること。
```
%%sql
DROP TABLE IF EXISTS product_2;
CREATE TABLE product_2 AS (
SELECT
product_cd,
category_major_cd,
category_medium_cd,
category_small_cd,
COALESCE(unit_price, unit_avg) as unit_price,
COALESCE(unit_cost, cost_avg) as unit_cost
FROM
product
CROSS JOIN (
SELECT
ROUND(AVG(unit_price)) AS unit_avg,
ROUND(AVG(unit_cost)) AS cost_avg
FROM
product
) stats_product
)
%%sql
SELECT
SUM(CASE WHEN unit_price IS NULL THEN 1 ELSE 0 END) AS unit_price,
SUM(CASE WHEN unit_cost IS NULL THEN 1 ELSE 0 END) AS unit_cost
FROM product_2 LIMIT 10
```
---
> S-082: 単価(unit_price)と原価(unit_cost)の欠損値について、それぞれの中央値で補完した新たなproduct_3を作成せよ。なお、中央値について1円未満は四捨五入とする。補完実施後、各項目について欠損が生じていないことも確認すること。
```
%%sql
DROP TABLE IF EXISTS product_3;
CREATE TABLE product_3 AS (
SELECT
product_cd,
category_major_cd,
category_medium_cd,
category_small_cd,
COALESCE(unit_price, unit_med) as unit_price,
COALESCE(unit_cost, cost_med) as unit_cost
FROM
product
CROSS JOIN (
SELECT
ROUND(
PERCENTILE_CONT(0.5) WITHIN GROUP(ORDER BY unit_price)
) AS unit_med,
ROUND(
PERCENTILE_CONT(0.5) WITHIN GROUP(ORDER BY unit_cost)
) AS cost_med
FROM
product
) stats_product
)
%%sql
SELECT
SUM(CASE WHEN unit_price IS NULL THEN 1 ELSE 0 END) AS unit_price,
SUM(CASE WHEN unit_cost IS NULL THEN 1 ELSE 0 END) AS unit_cost
FROM product_3 LIMIT 10
```
---
> S-083: 単価(unit_price)と原価(unit_cost)の欠損値について、各商品の小区分(category_small_cd)ごとに算出した中央値で補完した新たなproduct_4を作成せよ。なお、中央値について1円未満は四捨五入とする。補完実施後、各項目について欠損が生じていないことも確認すること。
```
%%sql
DROP TABLE IF EXISTS product_4;
CREATE TABLE product_4 AS (
WITH category_median AS(
SELECT
category_small_cd,
ROUND(
PERCENTILE_CONT(0.5) WITHIN GROUP(ORDER BY unit_price)
) AS unit_med,
ROUND(
PERCENTILE_CONT(0.5) WITHIN GROUP(ORDER BY unit_cost)
) AS cost_med
FROM product
GROUP BY category_small_cd
)
SELECT
product_cd,
category_major_cd,
category_medium_cd,
category_small_cd,
COALESCE(unit_price, unit_med) as unit_price,
COALESCE(unit_cost, cost_med) as unit_cost
FROM
product
JOIN
category_median
USING(category_small_cd)
)
%%sql
SELECT
SUM(CASE WHEN unit_price IS NULL THEN 1 ELSE 0 END) AS unit_price,
SUM(CASE WHEN unit_cost IS NULL THEN 1 ELSE 0 END) AS unit_cost
FROM product_4 LIMIT 10
```
---
> S-084: 顧客テーブル(customer)の全顧客に対し、全期間の売上金額に占める2019年売上金額の割合を計算せよ。ただし、販売実績のない場合は0として扱うこと。そして計算した割合が0超のものを抽出せよ。 結果は10件表示させれば良い。
```
%%sql
WITH sales_amount_2019 AS (
SELECT
customer_id,
SUM(amount) AS sum_amount_2019
FROM
receipt
WHERE
20190101 <= sales_ymd AND sales_ymd <= 20191231
GROUP BY
customer_id
),
sales_amount_all AS (
SELECT
customer_id,
SUM(amount) AS sum_amount_all
FROM
receipt
GROUP BY
customer_id
)
SELECT
a.customer_id,
COALESCE(b.sum_amount_2019, 0) AS sales_amount_2019,
COALESCE(c.sum_amount_all, 0) AS sales_amount_all,
CASE COALESCE(c.sum_amount_all, 0)
WHEN 0 THEN 0
ELSE COALESCE(b.sum_amount_2019, 0) * 1.0 / c.sum_amount_all
END AS sales_rate
FROM
customer a
LEFT JOIN
sales_amount_2019 b
ON a.customer_id = b.customer_id
LEFT JOIN
sales_amount_all c
ON a.customer_id = c.customer_id
WHERE CASE COALESCE(c.sum_amount_all, 0)
WHEN 0 THEN 0
ELSE COALESCE(b.sum_amount_2019, 0) * 1.0 / c.sum_amount_all
END > 0
LIMIT 10
```
---
> S-085: 顧客テーブル(customer)の全顧客に対し、郵便番号(postal_cd)を用いて経度緯度変換用テーブル(geocode)を紐付け、新たなcustomer_1を作成せよ。ただし、複数紐づく場合は経度(longitude)、緯度(latitude)それぞれ平均を算出すること。
```
%%sql
DROP TABLE IF EXISTS customer_1;
CREATE TABLE customer_1 AS (
WITH geocode_avg AS(
SELECT
postal_cd,
AVG(longitude) as m_longitude,
AVG(latitude) as m_latitude
FROM
geocode
GROUP BY
postal_cd
)
SELECT
*
FROM
customer c
JOIN
geocode_avg g
USING(postal_cd)
);
%%sql
SELECT * FROM customer_1 LIMIT 3
```
---
> S-086: 前設問で作成した緯度経度つき顧客テーブル(customer_1)に対し、申込み店舗コード(application_store_cd)をキーに店舗テーブル(store)と結合せよ。そして申込み店舗の緯度(latitude)・経度情報(longitude)と顧客の緯度・経度を用いて距離(km)を求め、顧客ID(customer_id)、顧客住所(address)、店舗住所(address)とともに表示せよ。計算式は簡易式で良いものとするが、その他精度の高い方式を利用したライブラリを利用してもかまわない。結果は10件表示すれば良い。
$$
緯度(ラジアン):\phi \\
経度(ラジアン):\lambda \\
距離L = 6371 * arccos(sin \phi_1 * sin \phi_2
+ cos \phi_1 * cos \phi_2 * cos(\lambda_1 − \lambda_2))
$$
```
%%sql
SELECT
c.customer_id,
c.address AS customer_address,
s.address AS store_address,
(
6371 * ACOS(
SIN(RADIANS(c.m_latitude))
* SIN(RADIANS(s.latitude))
+ COS(RADIANS(c.m_latitude))
* COS(RADIANS(s.latitude))
* COS(RADIANS(c.m_longitude) - RADIANS(s.longitude))
)
) AS distance
FROM
customer_1 c
JOIN
store s
ON
c.application_store_cd = s.store_cd
limit 10
```
---
> S-087: 顧客テーブル(customer)では、異なる店舗での申込みなどにより同一顧客が複数登録されている。名前(customer_name)と郵便番号(postal_cd)が同じ顧客は同一顧客とみなし、1顧客1レコードとなるように名寄せした名寄顧客テーブル(customer_u)を作成せよ。ただし、同一顧客に対しては売上金額合計が最も高いものを残すものとし、売上金額合計が同一もしくは売上実績の無い顧客については顧客ID(customer_id)の番号が小さいものを残すこととする。
```
%%sql
DROP TABLE IF EXISTS customer_u;
CREATE TABLE customer_u AS (
WITH sales_amount AS(
SELECT
c.customer_id,
c.customer_name,
c.postal_cd,
SUM(r.amount) as sum_amount
FROM
customer c
LEFT JOIN
receipt r
ON c.customer_id = r.customer_id
GROUP by
c.customer_id, c.customer_name, c.postal_cd
),
sales_ranking AS(
SELECT
*,
ROW_NUMBER() OVER(
PARTITION BY customer_name, postal_cd
ORDER BY sum_amount desc, customer_ID ) as rank
FROM sales_amount
)
SELECT c.*
FROM
customer c
JOIN
sales_ranking r
ON
c.customer_id = r.customer_id
and r.rank = 1
)
%%sql
SELECT
cnt,
cnt_u,
cnt - cnt_u AS diff
FROM
(SELECT count(1) as cnt FROM customer) customer
CROSS JOIN (SELECT count(1) as cnt_u FROM customer_u) customer_u
```
---
> S-088: 前設問で作成したデータを元に、顧客テーブルに統合名寄IDを付与したテーブル(customer_n)を作成せよ。ただし、統合名寄IDは以下の仕様で付与するものとする。
>
>- 重複していない顧客:顧客ID(customer_id)を設定
>- 重複している顧客:前設問で抽出したレコードの顧客IDを設定
```
%%sql
DROP TABLE IF EXISTS customer_n;
CREATE TABLE customer_n AS (
SELECT
c.*,
u.customer_id as integration_id
FROM
customer c
JOIN
customer_u u
ON c.customer_name = u.customer_name
and c.postal_cd = u.postal_cd
)
%%sql
SELECT count(1) FROM customer_n
WHERE customer_id != integration_id
```
---
> S-089: 売上実績のある顧客に対し、予測モデル構築のため学習用データとテスト用データに分割したい。それぞれ8:2の割合でランダムにデータを分割せよ。
```
%%sql
SELECT SETSEED(0.1);
CREATE TEMP TABLE IF NOT EXISTS sales_record_customer_id AS (
SELECT customer_id ,ROW_NUMBER()OVER(ORDER BY RANDOM()) AS row
FROM customer
LEFT JOIN receipt USING(customer_id)
GROUP BY customer_id
HAVING SUM(amount) IS NOT NULL
);
DROP TABLE IF EXISTS customer_train;
CREATE TABLE customer_train AS
SELECT customer.*
FROM sales_record_customer_id
LEFT JOIN customer USING(customer_id)
WHERE sales_record_customer_id.row < (SELECT
COUNT(0)
FROM sales_record_customer_id) *0.8
;
DROP TABLE IF EXISTS customer_test;
CREATE TABLE customer_test AS
SELECT customer.*
FROM sales_record_customer_id
LEFT JOIN customer USING(customer_id)
EXCEPT
SELECT * from customer_train
;
```
---
> S-090: レシート明細テーブル(receipt)は2017年1月1日〜2019年10月31日までのデータを有している。売上金額(amount)を月次で集計し、学習用に12ヶ月、テスト用に6ヶ月のモデル構築用データを3テーブルとしてセット作成せよ。データの持ち方は自由とする。
```
%%sql
-- SQL向きではないため、やや強引に記載する(分割数が多くなる場合はSQLが長くなるため現実的ではない)
-- また、秒単位のデータなど時系列が細かく、かつ長期間に渡る場合はデータが膨大となるため注意(そのようなケースではループ処理でモデル学習ができる言語が望ましい)
-- 学習データ(0)とテストデータ(1)を区別するフラグを付与する
DROP TABLE IF EXISTS sales_amount ;
CREATE TABLE sales_amount AS (
SELECT
SUBSTR(CAST(sales_ymd AS VARCHAR), 1, 6) AS sales_ym,
SUM(amount) AS sum_amount,
row_number() OVER(PARTITION BY NULL ORDER BY
SUBSTR(CAST(sales_ymd AS VARCHAR), 1, 6)) AS rn
FROM
receipt
GROUP BY sales_ym
);
-- SQLでは限界があるが、作成データセットの増加に伴いなるべく使いまわしができるものにする
-- WITH句内のLAG関数について、ラグ期間を変えれば使い回せるよう記述
DROP TABLE IF EXISTS series_data_1 ;
CREATE TABLE series_data_1 AS (
WITH lag_amount AS (
SELECT sales_ym, sum_amount, LAG(rn, 0) OVER (ORDER BY rn) AS rn
FROM sales_amount
)
SELECT
sales_ym, sum_amount,
CASE WHEN rn <= 12 THEN 0 WHEN 12 < rn THEN 1 END as test_flg
FROM lag_amount
WHERE rn <= 18);
DROP TABLE IF EXISTS series_data_2 ;
CREATE TABLE series_data_2 AS (
WITH lag_amount AS (
SELECT
sales_ym,
sum_amount,
LAG(rn, 6) OVER (ORDER BY rn) AS rn
FROM sales_amount
)
SELECT
sales_ym,
sum_amount,
CASE WHEN rn <= 12 THEN 0 WHEN 12 < rn THEN 1 END as test_flg
FROM lag_amount WHERE rn <= 18);
DROP TABLE IF EXISTS series_data_3 ;
CREATE TABLE series_data_3 AS (
WITH lag_amount AS (
SELECT sales_ym, sum_amount, LAG(rn, 12) OVER (ORDER BY rn) AS rn
FROM sales_amount
)
SELECT
sales_ym,
sum_amount,
CASE WHEN rn <= 12 THEN 0 WHEN 12 < rn THEN 1 END as test_flg
FROM lag_amount WHERE rn <= 18);
%%sql
SELECT * FROM series_data_1
```
---
> S-091: 顧客テーブル(customer)の各顧客に対し、売上実績のある顧客数と売上実績のない顧客数が1:1となるようにアンダーサンプリングで抽出せよ。
```
%%sql
SELECT SETSEED(0.1);
WITH pre_table_1 AS(
SELECT
c.*
,COALESCE(r.amount,0) AS r_amount
FROM
customer c
LEFT JOIN
receipt r
ON
c.customer_id=r.customer_id
)
,pre_table_2 AS(
SELECT
customer_id
,CASE WHEN SUM(r_amount)>0 THEN 1 ELSE 0 END AS is_buy_flag
,CASE WHEN SUM(r_amount)=0 THEN 1 ELSE 0 END AS is_not_buy_flag
FROM
pre_table_1
GROUP BY
customer_id
)
,pre_table_3 AS(
SELECT
*
,ROW_NUMBER() OVER(PARTITION BY is_buy_flag ORDER BY RANDOM())
FROM
pre_table_2
CROSS JOIN
(SELECT SUM(is_buy_flag) AS buying FROM pre_table_2) AS t1
CROSS JOIN
(SELECT SUM(is_not_buy_flag) AS not_buying FROM pre_table_2) AS t2
)
,pre_table_4 AS(
SELECT
*
FROM
pre_table_3
WHERE
row_number<=buying
AND
row_number<=not_buying
)
SELECT COUNT(*) FROM pre_table_4 GROUP BY is_buy_flag;
```
---
> S-092: 顧客テーブル(customer)では、性別に関する情報が非正規化の状態で保持されている。これを第三正規化せよ。
```
%%sql
DROP TABLE IF EXISTS customer_std;
CREATE TABLE customer_std AS (
SELECT
customer_id,
customer_name,
gender_cd,
birth_day,
age,
postal_cd,
application_store_cd,
application_date,
status_cd
FROM
customer
);
DROP TABLE IF EXISTS gender_std;
CREATE TABLE gender_std AS (
SELECT distinct
gender_cd, gender
FROM
customer
)
%%sql
SELECT * FROM gender_std
```
---
> S-093: 商品テーブル(product)では各カテゴリのコード値だけを保有し、カテゴリ名は保有していない。カテゴリテーブル(category)と組み合わせて非正規化し、カテゴリ名を保有した新たな商品テーブルを作成せよ。
```
%%sql
DROP TABLE IF EXISTS product_full;
CREATE TABLE product_full AS (
SELECT
p.product_cd,
p.category_major_cd,
c.category_major_name,
p.category_medium_cd,
c.category_medium_name,
p.category_small_cd,
c.category_small_name,
p.unit_price,
p.unit_cost
FROM
product p
JOIN
category c
USING(category_small_cd)
)
%%sql
SELECT * FROM product_full LIMIT 10
```
---
> S-094: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。出力先のパスは"/tmp/data"を指定することでJupyterの"/work/data"と共有されるようになっている。なお、COPYコマンドの権限は付与済みである。
>
> - ファイル形式はCSV(カンマ区切り)
> - ヘッダ有り
> - 文字コードはUTF-8
```
%%sql
COPY product_full TO '/tmp/data/S_product_full_UTF-8_header.csv'
WITH CSV HEADER encoding 'UTF-8'
```
---
> S-095: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。出力先のパスは"/tmp/data"を指定することでJupyterの"/work/data"と共有されるようになっている。なお、COPYコマンドの権限は付与済みである。
>
> - ファイル形式はCSV(カンマ区切り)
> - ヘッダ有り
> - 文字コードはSJIS
```
%%sql
COPY product_full TO '/tmp/data/S_product_full_SJIS_header.csv'
WITH CSV HEADER encoding 'SJIS'
```
---
> S-096: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。出力先のパスは"/tmp/data"を指定することでJupyterの"/work/data"と共有されるようになっている。なお、COPYコマンドの権限は付与済みである。
>
> - ファイル形式はCSV(カンマ区切り)
> - ヘッダ無し
> - 文字コードはUTF-8
```
%%sql
COPY product_full TO '/tmp/data/S_product_full_UTF-8_noh.csv'
WITH CSV encoding 'UTF-8'
```
---
> S-097: 先に作成した以下形式のファイルを読み込み、テーブルを作成せよ。また、先頭3件を表示させ、正しくとりまれていることを確認せよ。
>
> - ファイル形式はCSV(カンマ区切り)
> - ヘッダ有り
> - 文字コードはUTF-8
```
%%sql
DROP TABLE IF EXISTS product_full;
CREATE TABLE product_full (
product_cd VARCHAR(10),
category_major_cd VARCHAR(2),
category_major_name VARCHAR(20),
category_medium_cd VARCHAR(4),
category_medium_name VARCHAR(20),
category_small_cd VARCHAR(6),
category_small_name VARCHAR(20),
unit_price INTEGER,
unit_cost INTEGER
);
%%sql
COPY product_full FROM '/tmp/data/S_product_full_UTF-8_header.csv'
WITH CSV HEADER encoding 'UTF-8'
%%sql
SELECT * FROM product_full LIMIT 3
```
---
> S-098: 先に作成した以下形式のファイルを読み込み、テーブルを作成せよ。また、先頭3件を表示させ、正しくとりまれていることを確認せよ。
>
> - ファイル形式はCSV(カンマ区切り)
> - ヘッダ無し
> - 文字コードはUTF-8
```
%%sql
DROP TABLE IF EXISTS product_full;
CREATE TABLE product_full (
product_cd VARCHAR(10),
category_major_cd VARCHAR(2),
category_major_name VARCHAR(20),
category_medium_cd VARCHAR(4),
category_medium_name VARCHAR(20),
category_small_cd VARCHAR(6),
category_small_name VARCHAR(20),
unit_price INTEGER,
unit_cost INTEGER
);
%%sql
COPY product_full FROM '/tmp/data/S_product_full_UTF-8_noh.csv'
WITH CSV encoding 'UTF-8'
%%sql
SELECT * FROM product_full LIMIT 3
```
---
> S-099: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。出力先のパスは"/tmp/data"を指定することでJupyterの"/work/data"と共有されるようになっている。なお、COPYコマンドの権限は付与済みである。
>
> - ファイル形式はTSV(タブ区切り)
> - ヘッダ有り
> - 文字コードはUTF-8
```
%%sql
COPY product_full TO '/tmp/data/S_product_full_UTF-8_header.tsv'
WITH CSV HEADER DELIMITER E'\t' encoding 'UTF-8'
```
---
> S-100: 先に作成した以下形式のファイルを読み込み、テーブルを作成せよ。また、先頭10件を表示させ、正しくとりまれていることを確認せよ。
>
> - ファイル形式はTSV(タブ区切り)
> - ヘッダ有り
> - 文字コードはUTF-8
```
%%sql
DROP TABLE IF EXISTS product_full;
CREATE TABLE product_full (
product_cd VARCHAR(10),
category_major_cd VARCHAR(2),
category_major_name VARCHAR(20),
category_medium_cd VARCHAR(4),
category_medium_name VARCHAR(20),
category_small_cd VARCHAR(6),
category_small_name VARCHAR(20),
unit_price INTEGER,
unit_cost INTEGER
);
%%sql
COPY product_full FROM '/tmp/data/S_product_full_UTF-8_header.tsv'
WITH CSV HEADER DELIMITER E'\t' encoding 'UTF-8'
%%sql
SELECT * FROM product_full LIMIT 10
```
# これで100本終わりです。おつかれさまでした!
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
# %matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
import src.solver_helper as helper
from src.traffic_world import TrafficWorld
from src.car_plotting_multiple import plot_multiple_cars, plot_cars, animate, plot_single_frame
from src.multiagent_mpc import MultiMPC, generate_warm_starts
from src.vehicle import Vehicle
from src.idm import IDM_acceleration, IDM_trajectory_prediction, get_lead_vehicle, MOBIL_lanechange
from contextlib import redirect_stdout
params = {}
params["T"] = 3
params["dt"] = 0.2
params["p_exec"] = 0.4
params["n_lanes"] = 2
params["n_other"] = 4
params["car_density"] = 5000
params["seed"] = 0
params["random_svo"] = 1
params["wall_CA"] = 0
i_mpc_start = 0
params["N"] = max(1, int(params["T"] / params["dt"]))
params["number_ctrl_pts_executed"] = max(1, int(np.floor(params["N"] * params["p_exec"])))
### Create the world and vehicle objects
world = TrafficWorld(params["n_lanes"], 0, 999999)
### Create the vehicle placement based on a Poisson distribution
MAX_VELOCITY = 25 * 0.447 # m/s
VEHICLE_LENGTH = 4.5 # m
time_duration_s = (params["n_other"] * 3600.0 /
params["car_density"]) * 10 # amount of time to generate traffic
initial_vehicle_positions = helper.poission_positions(params["car_density"],
int(time_duration_s),
params["n_lanes"],
MAX_VELOCITY,
VEHICLE_LENGTH,
position_random_seed=params["seed"])
position_list = initial_vehicle_positions[:params["n_other"]]
### Create the SVOs for each vehicle
if params["random_svo"] == 1:
list_of_svo = [np.random.choice([0, np.pi / 4.0, np.pi / 2.01]) for i in range(params["n_other"])]
else:
list_of_svo = [params["svo_theta"] for i in range(params["n_other"])]
(ambulance, amb_x0, all_other_vehicles,
all_other_x0) = helper.initialize_cars_from_positions(params["N"], params["dt"], world, True, position_list,
list_of_svo)
from matplotlib.patches import Circle, Rectangle
from matplotlib.transforms import Affine2D
class CenterRectangle(Rectangle):
def __init__(self, xy, width, height, angle=0.0, **kwargs):
lower_xy = (xy[0] - width/2.0, xy[1]-height/2.0)
Rectangle.__init__(self, lower_xy, width, height, angle=angle, **kwargs)
fig, axs = plt.subplots(2,1)
def add_steering_wheel(ax, angle_radians):
# rotation = Affine2D().rotate_around(0.0, 0.0, angle)
# rotation = None
c1 = Circle((0,0), radius = 5, fill = None, linewidth=10)
c2 = Circle((0,0), radius = 1, facecolor = 'black')
angle_deg = np.rad2deg(angle_radians)
right_rectangle = CenterRectangle((2.5,0), 5, 1, angle_deg, facecolor='black')
left_rectangle = CenterRectangle((-2.5,0),5, 1, angle_deg, facecolor='black')
# left_rectangle = Rectangle((-5*np.sin(angle),0), 5, 1, facecolor='black')
bottom_rectangle = CenterRectangle((0, -2.5), 1, 5, angle_deg, facecolor='black')
# c2 = Circle((0,0), radius = 3, fill = 'white')
ax.add_patch(c1)
ax.add_patch(c2)
ax.add_patch(left_rectangle)
ax.add_patch(right_rectangle)
ax.add_patch(bottom_rectangle)
add_steering_wheel(axs[0], np.pi/2.0)
axs[0].set_xlim([-10,10])
axs[0].set_ylim([-10, 10])
axs[0].axis('square')
plt.show()
plot_multiple_cars(0, world, ambulance, amb_x0.reshape(6,1), [x.reshape(6,1) for x in all_other_x0],
None, car_labels = [str(i) for i in range(len(all_other_x0))])
plt.show()
import ipywidgets as widgets
from IPython.display import display
import time
class acceleration_button:
def __init__(self):
self.accel = 0
self.steering = 0
def on_button_clicked(self, delta_accel):
self.accel += delta_accel
print("new a", self.accel)
def on_steer_clicked(self, delta_steering):
self.steering -= delta_steering
print("new steeer", self.steering)
def on_speed_up_clicked(self, button):
self.on_button_clicked(delta_accel = 1.0)
def on_speed_down_clicked(self, button):
self.on_button_clicked(delta_accel = -1.0)
def on_steer_left_clicked(self, b):
self.on_steer_clicked(-1)
def on_steer_right_clicked(self, b):
self.on_steer_clicked(1)
button_speed_up = widgets.Button(description="Speed up!")
button_slow_down = widgets.Button(description="Slow down!")
button_steer_left = widgets.Button(description="Steer left")
button_steer_right = widgets.Button(description="Steer right")
output = widgets.Output()
display(button_speed_up, button_slow_down, button_steer_left, button_steer_right,
output)
a_button = acceleration_button()
button_speed_up.on_click(a_button.on_speed_up_clicked)
button_slow_down.on_click(a_button.on_speed_down_clicked)
button_steer_left.on_click(a_button.on_steer_left_clicked)
button_steer_right.on_click(a_button.on_steer_right_clicked)
w = widgets.IntSlider()
display(w, output)
print(w.value)
from src.human_keyboard import SteeringKeyboard
N_total = 300
X_other = [np.zeros((6, N_total+1)) for i in range(len(all_other_vehicles))]
X_amb = np.zeros((6,N_total+1))
X_amb[:,0] = amb_x0
for i in range(len(X_other)):
X_other[i][:,0] = all_other_x0[i]
idm_params = {
"desired_time_gap": 0.1,
"jam_distance": 4,
}
# out_file = open('/dev/null','rb')
for t in range(N_total):
print(t)
current_other_x0 = [X_other[i][:,t] for i in range(len(X_other))]
current_amb_x0 = X_amb[:,t]
current_other_veh = [all_other_vehicles[i] for i in range(len(X_other))]
# Mobile lane change
driver_x0 = current_amb_x0
driver_veh = ambulance
all_other_x0 = current_other_x0
MOBIL_params = {
"politeness_factor": 0.1,
}
# Compute whether lane change occurs
desired_lane = True
new_lane, accel = MOBIL_lanechange(driver_x0, driver_veh, all_other_x0, current_other_veh, world, desired_lane,
MOBIL_params, idm_params)
if new_lane is not None:
driver_veh.update_desired_lane(world, new_lane, True)
# Compute IDM acceleration (ambulance)
lead_veh = get_lead_vehicle(current_amb_x0, current_other_x0, world)
if lead_veh is None:
x_lead = current_amb_x0 + 999999
v_lead = 999999
else:
x_lead = current_other_x0[lead_veh]
v_lead = x_lead[4] * np.cos(x_lead[2])
v_current = current_amb_x0[4] * np.cos(current_amb_x0[2])
v_desired = ambulance.max_v
bumper_distance = x_lead[0] - current_amb_x0[0]
idm_params["maximum_acceleration"] = ambulance.max_acceleration
a_IDM = IDM_acceleration(bumper_distance, v_lead, v_current, v_desired, idm_params)
# Solve for steering angle
ambulance.k_lat = 5.0
ambulance.k_lan = 2.0
ambulance.k_x_dot = 0.0
ambulance.k_final = 0.0
solver_params = {
"k_slack" : 1000,
"k_CA": 0.05,
"k_CA_power": 1.0,
# "constant_v": True,
}
warm_starts = generate_warm_starts(ambulance, world, current_amb_x0, [], params)
u=None
for k_warm in warm_starts:
try:
steering_mpc = MultiMPC(ambulance, [], [], world, solver_params)
n_mpc = 5
steering_mpc.generate_optimization(params["N"], current_amb_x0, [], [], params=params, ipopt_params = {'print_level':0})
u_warm, x_warm, x_des_warm = warm_starts[k_warm]
steering_mpc.opti.set_initial(steering_mpc.u_ego, u_warm)
steering_mpc.opti.set_initial(steering_mpc.x_ego, x_warm)
steering_mpc.opti.set_initial(steering_mpc.xd_ego, x_des_warm)
# with redirect_stdout(out_file):
steering_mpc.solve(None, None)
_, u, _ = steering_mpc.get_bestresponse_solution()
except RuntimeError:
print("Solver didn't work")
if u is not None:
break
if u is None:
raise Exception("Solver didn't solve")
# Update control and step the simulator
u_ego = np.array([[u[0,0]],[a_IDM*ambulance.dt]])
# Update state with new control inputs
x_ego_traj, _ = ambulance.forward_simulate_all(current_amb_x0, u_ego)
X_amb[:,t+1] = x_ego_traj[:, 1]
# Other vehicles
for ego_idx in range(len(all_other_vehicles)):
N = 1
# Mobile lane change
driver_x0 = current_other_x0[ego_idx]
driver_veh = all_other_vehicles[ego_idx]
all_other_x0 = current_other_x0[:ego_idx] + current_other_x0[ego_idx+1:] + [current_amb_x0]
all_other_veh = all_other_vehicles[:ego_idx] + all_other_vehicles[ego_idx+1:] + [ambulance]
MOBIL_params = {
"politeness_factor": 0.5,
}
new_lane, accel = MOBIL_lanechange(driver_x0, driver_veh, all_other_x0, all_other_veh, world, desired_lane,
MOBIL_params, idm_params)
if new_lane is not None:
driver_veh.update_desired_lane(world, new_lane, True)
driver_x0 = current_other_x0[ego_idx]
dummy_x0 = current_other_x0[ego_idx] - 10000
ado_x0s = current_other_x0[:ego_idx] + [dummy_x0] + current_other_x0[ego_idx+1:] + [current_amb_x0]
lead_veh = get_lead_vehicle(current_other_x0[ego_idx], ado_x0s, world)
if lead_veh is None:
x_lead = driver_x0 + 999999
v_lead = 999999
else:
x_lead = ado_x0s[lead_veh]
v_lead = x_lead[4] * np.cos(x_lead[2])
v_current = driver_x0[4] * np.cos(driver_x0[2])
v_desired = driver_veh.max_v
# v_desired = 0.001
v_desired = driver_veh.max_v * .5
bumper_distance = x_lead[0] - driver_x0[0]
idm_params["maximum_acceleration"] = driver_veh.max_acceleration
a_IDM = IDM_acceleration(bumper_distance, v_lead, v_current, v_desired, idm_params)
# Solve for steering angle
driver_veh.k_lat = 5.0
driver_veh.k_lan = 2.0
driver_veh.k_x_dot = 0.0
driver_veh.k_final = 0.0
solver_params = {
"k_slack" : 1000,
"k_CA": 0.05,
"k_CA_power": 1.0,
# "constant_v": True,
}
warm_starts = generate_warm_starts(driver_veh, world, driver_x0, [], params)
u = None
for k_warm in warm_starts:
try:
steering_mpc = MultiMPC(driver_veh, [], [], world, solver_params)
n_mpc = 5
steering_mpc.generate_optimization(params["N"], driver_x0, [], [], params=params, ipopt_params = {'print_level':0})
u_warm, x_warm, x_des_warm = warm_starts[k_warm]
steering_mpc.opti.set_initial(steering_mpc.u_ego, u_warm)
steering_mpc.opti.set_initial(steering_mpc.x_ego, x_warm)
steering_mpc.opti.set_initial(steering_mpc.xd_ego, x_des_warm)
# with redirect_stdout(out_file):
steering_mpc.solve(None, None)
_, u, _ = steering_mpc.get_bestresponse_solution()
except RuntimeError:
print("Solver didn't work")
if u is not None:
break
if u is None:
raise Exception("Solver didn't solve")
# Update control and step the simulator
u_ego = np.array([[u[0,0]],[a_IDM*driver_veh.dt]])
# Update state with new control inputs
x_ego_traj, _ = driver_veh.forward_simulate_all(driver_x0, u_ego)
X_other[ego_idx][:,t+1] = x_ego_traj[:, 1]
# print(ego_idx, X_ego[:,1])
X_amb = X_amb[:, :t+1]
X_other = [x[:,:t+1] for x in X_other]
dummy_veh = Vehicle(0.2)
temp_folder = "/home/nbuckman/mpc-multiple-vehicles/jupyter_notebooks/temp/"
plot_cars(world, dummy_veh, X_amb, X_other, temp_folder, "image")
vid_fname = temp_folder + "imgs/MOBIL_halfspeed.mp4"
animate(temp_folder, vid_fname, fps=16)
from IPython.display import Video
Video(vid_fname, embed=True, width=1024)
vid_fname = temp_folder + "MOBIL_Egoistic.mp4"
animate(temp_folder, vid_fname, fps=16)
plot_multiple_cars(0, world, ambulance, np.zeros((6,1)), [np.array([[6, 0,0,0,0,0]]).T],
None, car_plot_shape = 'both')
plt.show()
plot_multiple_cars(0, world, ambulance, np.zeros((6,1)), [np.array([[ambulance.L, 0,0,0,0,0]]).T],
None, car_plot_shape = 'image')
plt.show()
np.array([[6, 0,0,0,0,0]]).shape
# driver_idx = 1
# driver_x0 = X_other[driver_idx][:,-1]
# driver_veh = all_other_vehicles[driver_idx]
# all_other_x0 = [X_other[idx][:,-1] for idx in range(1,len(X_other)) if idx!=driver_idx] + [amb_x0]
# all_other_veh = [all_other_vehicles[idx] for idx in range(1,len(X_other)) if idx!=driver_idx] + [ambulance]
driver_x0 = X_amb[:, -1]
driver_veh = ambulance
all_other_x0 = [X_other[idx][:,-1] for idx in range(1,len(X_other))]
all_other_veh = [all_other_vehicles[idx] for idx in range(1,len(X_other))]
plot_multiple_cars(0, world, ambulance, driver_x0.reshape(6,1), [x.reshape(6,1) for x in all_other_x0],
None, car_labels = [str(i) for i in range(len(all_other_x0))])
plt.show()
MOBIL_params = {}
lane, accel = helper.MOBIL_lanechange(driver_x0, driver_veh, all_other_x0, all_other_veh, world, MOBIL_params, idm_params)
lane
lane
accel
default_MOBIL_params = {
"politeness_factor": 0.5,
"changing_threshold": 0.1,
"maximum_safe_deceleration": 4,
"bias_for_right_lane": 0.3
}
if MOBIL_params:
for param in MOBIL_params:
try:
default_MOBIL_params[param] = MOBIL_params[param]
except KeyError:
raise Exception("Key Error: Check if MOBIL Param is correct")
p = default_MOBIL_params["politeness_factor"]
a_thr = default_MOBIL_params["changing_threshold"]
b_safe = default_MOBIL_params["maximum_safe_deceleration"]
a_bias = default_MOBIL_params["bias_for_right_lane"]
safety_criteria = accel["newfollower_after"] >= -b_safe
driver_incentive = accel["driver_after"] - accel["driver_before"]
new_follower_incentive = accel["newfollower_after"] - accel["newfollower_before"]
old_follower_incentive = accel["oldfollower_after"] - accel["oldfollower_before"]
incentive_criteria = (driver_incentive + p * (new_follower_incentive + old_follower_incentive)) >= (a_thr)
driver_incentive
((new_follower_incentive + old_follower_incentive)*p + driver_incentive >= a_thr)
a_thr
```
| github_jupyter |
```
import os
import sys
import gym
from gym import wrappers, logger
import gridworld
import random
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
matplotlib.use("TkAgg")
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from collections import namedtuple, deque
```
## utils
```
Transition = namedtuple('Transition', ('state', 'next_state', 'action', 'reward', 'mask'))
class Memory(object):
def __init__(self, capacity):
self.memory = deque(maxlen=capacity)
self.capacity = capacity
def push(self, state, next_state, action, reward, mask):
self.memory.append(Transition(state, next_state, action, reward, mask))
def sample(self, batch_size):
transitions = random.sample(self.memory, batch_size)
batch = Transition(*zip(*transitions))
return batch
def __len__(self):
return len(self.memory)
def get_action(state, online_net, epsilon, env):
if np.random.rand() <= epsilon:
return env.action_space.sample()
else:
return online_net.get_action(state)
def update_target_model(online_net, target_net):
target_net.load_state_dict(online_net.state_dict())
class FeaturesExtractor(object):
def __init__(self,outSize):
super().__init__()
self.outSize=outSize*3
def __call__(self, obs):
state=np.zeros((3,np.shape(obs)[0],np.shape(obs)[1]))
state[0]=np.where(obs == 2,1,state[0])
state[1]=np.where(obs == 4,1,state[1])
state[2]=np.where(obs == 6,1,state[2])
return state.reshape(1,-1)
```
## DQN
```
class QNet(nn.Module):
def __init__(self, num_inputs, num_outputs):
super(QNet, self).__init__()
self.num_inputs = num_inputs
self.num_outputs = num_outputs
self.fc1 = nn.Linear(num_inputs, 256)
self.fc2 = nn.Linear(256, 30)
self.fc3 = nn.Linear(30, num_outputs)
for m in self.modules():
if isinstance(m, nn.Linear):
nn.init.xavier_uniform_(m.weight)
def forward(self, x):
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
qvalue = self.fc3(x)
return qvalue
@classmethod
def train_model(cls, online_net, target_net, optimizer, batch):
states = torch.stack(batch.state)
next_states = torch.stack(batch.next_state)
actions = torch.Tensor(batch.action).float()
rewards = torch.Tensor(batch.reward)
masks = torch.Tensor(batch.mask)
pred = online_net(states).squeeze(1)
next_pred = target_net(next_states).squeeze(1)
pred = torch.sum(pred.mul(actions), dim=1)
target = rewards + masks * gamma * next_pred.max(1)[0]
loss = F.mse_loss(pred, target.detach())
optimizer.zero_grad()
loss.backward()
optimizer.step()
return loss
def get_action(self, input):
qvalue = self.forward(input)
_, action = torch.max(qvalue, 1)
return action.numpy()[0]
```
### hyper parameters
```
gamma = 0.99
batch_size = 64
lr = 0.0001
update_target = 20
replay_memory_capacity = 1000
count_epsds = 500
epsilon = 1.0
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
```
##### train
```
env_name = "gridworld-v0"
env = gym.make(env_name)
env.seed(0)
torch.manual_seed(0)
# log file plusieurs scenarios
outdir = 'gridworld-v0/random-agent-results'
envm = wrappers.Monitor(env, directory=outdir, force=True , video_callable=False )
env.setPlan("gridworldPlans/plan1.txt", {0: -0.1, 3: 1, 4: 1, 5: -1, 6: -1})
env.seed() # randomness init
num_actions = env.action_space.n
Phi_s = FeaturesExtractor(num_actions)
num_inputs = Phi_s(env.reset()).shape[1]
online_net = QNet(num_inputs, num_actions)
target_net = QNet(num_inputs, num_actions)
update_target_model(online_net, target_net)
optimizer = optim.Adam(online_net.parameters(), lr=lr)
online_net.to(device)
target_net.to(device)
online_net.train()
target_net.train()
memory = Memory(replay_memory_capacity)
steps = 0
nbr_actions_list, score_list, loss_list = [],[],[]
for e in range(count_epsds):
done = False
score = 0
state = env.reset()
state = torch.Tensor(state).to(device)
state = torch.Tensor(Phi_s(state))
nbr_actions = 0
avg_loss = 0
while not done:
steps += 1
action = get_action(state, online_net, epsilon, env)
next_state, reward, done, _ = env.step(action)
nbr_actions += 1
next_state = torch.Tensor(Phi_s(next_state))
mask = 0 if done else 1
action_one_hot = np.zeros(4)
action_one_hot[action] = 1
memory.push(state, next_state, action_one_hot, reward, mask)
score += reward
state = next_state
if ( e % 100 ):
env.render(0.0001)
if steps > replay_memory_capacity:
epsilon -= 0.00005
epsilon = max(epsilon, 0.01)
batch = memory.sample(batch_size)
loss = QNet.train_model(online_net, target_net, optimizer, batch)
if steps % update_target == 0 :
update_target_model(online_net, target_net)
avg_loss += loss
epsilon -=0.02
epsilon = max(epsilon, 0.01)
score_list.append(score)
if ( e % 50 == 0 and e > 0 ):
optimizer.param_groups[0]["lr"] /= 2
nbr_actions_list.append(nbr_actions)
if ( steps > replay_memory_capacity ) :
loss_list.append(avg_loss/nbr_actions)
print('episode {}| score: {:.4f}| Avg Loss: {:.4f}| epsilon_greedy : {:.2f}'.format(e, score,
avg_loss/nbr_actions, epsilon))
else :
print('episode {}| score: {:.4f}| epsilon_greedy : {:.2f}'.format(e, score, epsilon))
env.close()
%matplotlib inline
plt.figure(figsize=(12,6))
plt.subplot(1,2,1)
plt.plot( np.arange(count_epsds), score_list, "g")
plt.xlabel('epsiodes')
plt.title("score")
plt.subplot(1,2,2)
plt.plot( np.arange(count_epsds), nbr_actions_list,"r")
plt.xlabel('episodes')
plt.title("nbr actions")
plt.show()
```
| github_jupyter |
# Getting started with ReColorAdv
This file contains instructions for experimenting with the ReColorAdv attack, by itself and combined with other attacks. This tutorial is based on the [first tutorial](https://github.com/revbucket/mister_ed/blob/master/notebooks/tutorial_1.ipynb) of `mister_ed`. See the README to make sure all dependencies are installed.
## Imports
First let's make sure that you can import everything you need:
```
# EXTERNAL LIBRARIES
import numpy as np
import re
import torch
import torch.nn as nn
import torch.optim as optim
# mister_ed
import recoloradv.mister_ed.loss_functions as lf
import recoloradv.mister_ed.utils.pytorch_utils as utils
import recoloradv.mister_ed.utils.image_utils as img_utils
import recoloradv.mister_ed.cifar10.cifar_loader as cifar_loader
import recoloradv.mister_ed.cifar10.cifar_resnets as cifar_resnets
import recoloradv.mister_ed.adversarial_training as advtrain
import recoloradv.mister_ed.utils.checkpoints as checkpoints
import recoloradv.mister_ed.adversarial_perturbations as ap
import recoloradv.mister_ed.adversarial_attacks as aa
import recoloradv.mister_ed.spatial_transformers as st
import recoloradv.mister_ed.config as config
# ReColorAdv
import recoloradv.perturbations as pt
import recoloradv.color_transformers as ct
import recoloradv.color_spaces as cs
from recoloradv import norms
from recoloradv.utils import load_pretrained_cifar10_model, get_attack_from_name
```
# Generating adversarial examples
Here, we will demonstrate how to generate a single minibatch of adversarial examples using ReColorAdv on CIFAR-10.
To set up, let's start by collecting a minibatch worth of data and loading up our classifier to attack.
```
cifar_valset = cifar_loader.load_cifar_data('val', batch_size=16)
examples, labels = next(iter(cifar_valset))
model, normalizer = load_pretrained_cifar10_model('pretrained_models/normal.resnet32.pt')
if utils.use_gpu():
examples = examples.cuda()
labels = labels.cuda()
model.cuda()
```
Let's take a look at what our original images look like:
```
img_utils.show_images(examples)
```
## ReColorAdv
Now let's attack all of these examples with a ReColorAdv attack that changes every pixel using the same function.
```
# This threat model defines the regularization parameters of the attack.
recoloradv_threat = ap.ThreatModel(pt.ReColorAdv, {
'xform_class': ct.FullSpatial,
'cspace': cs.CIELUVColorSpace(), # controls the color space used
'lp_style': 'inf',
'lp_bound': [0.06, 0.06, 0.06], # [epsilon_1, epsilon_2, epsilon_3]
'xform_params': {
'resolution_x': 16, # R_1
'resolution_y': 32, # R_2
'resolution_z': 32, # R_3
},
'use_smooth_loss': True,
})
# Now, we define the main optimization term (the Carlini & Wagner f6 loss).
adv_loss = lf.CWLossF6(model, normalizer)
# We also need the smoothness loss.
smooth_loss = lf.PerturbationNormLoss(lp=2)
# We combine them with a RegularizedLoss object.
attack_loss = lf.RegularizedLoss({'adv': adv_loss, 'smooth': smooth_loss},
{'adv': 1.0, 'smooth': 0.05}, # lambda = 0.05
negate=True) # Need this true for PGD type attacks
# PGD is used to optimize the above loss.
pgd_attack_obj = aa.PGD(model, normalizer, recoloradv_threat, attack_loss)
# We run the attack for 10 iterations at learning rate 0.01.
perturbation = pgd_attack_obj.attack(examples, labels, num_iterations=10, signed=False,
optimizer=optim.Adam, optimizer_kwargs={'lr': 0.01},
verbose=True)
# Now, we can collect the successful adversarial examples and display them.
successful_advs, successful_origs = perturbation.collect_successful(model, normalizer)
successful_diffs = ((successful_advs - successful_origs) * 3 + 0.5).clamp(0, 1)
img_utils.show_images([successful_origs, successful_advs, successful_diffs])
```
In the above image, the first row is the original images; the second row is the adversarial examples; and the third row is the magnified difference between them.
## Combined Attacks
Now that we've seen how to use the ReColorAdv attack, we can combine it with an additive delta attack.
```
# First, we define the additive threat model.
additive_threat = ap.ThreatModel(ap.DeltaAddition, {
'lp_style': 'inf',
'lp_bound': 0.03,
})
# Combine it with the ReColorAdv functional threat model.
combined_threat = ap.ThreatModel(
ap.SequentialPerturbation,
[recoloradv_threat, additive_threat],
ap.PerturbationParameters(norm_weights=[1.0, 0.0]),
)
# Again, define the optimization terms.
adv_loss = lf.CWLossF6(model, normalizer)
smooth_loss = lf.PerturbationNormLoss(lp=2)
attack_loss = lf.RegularizedLoss({'adv': adv_loss, 'smooth': smooth_loss},
{'adv': 1.0, 'smooth': 0.05},
negate=True) # Need this true for PGD type attacks
# Setup and run PGD over both perturbations at once.
pgd_attack_obj = aa.PGD(model, normalizer, combined_threat, attack_loss)
perturbation = pgd_attack_obj.attack(examples, labels, num_iterations=10, signed=False,
optimizer=optim.Adam, optimizer_kwargs={'lr': 0.01},
verbose=True)
# Display the successful adversarial examples.
successful_advs, successful_origs = perturbation.collect_successful(model, normalizer)
successful_diffs = ((successful_advs - successful_origs) * 3 + 0.5).clamp(0, 1)
img_utils.show_images([successful_origs, successful_advs, successful_diffs])
```
Note that the resulting adversarial examples have been both recolored using ReColorAdv and had some additive adversarial noise applied from the delta attack.
## Prebuilt Attacks
The convenience function `get_attack_from_name` allows you to easily instantiate one of the attacks used in the paper. For instance, to use the combined ReColorAdv, StAdv, and delta attacks:
```
attack = get_attack_from_name('recoloradv+stadv+delta', model, normalizer, verbose=True)
perturbation = attack.attack(examples, labels)
# Display the successful adversarial examples.
successful_advs, successful_origs = perturbation.collect_successful(model, normalizer)
successful_diffs = ((successful_advs - successful_origs) * 3 + 0.5).clamp(0, 1)
img_utils.show_images([successful_origs, successful_advs, successful_diffs])
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Partial Differential Equations
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r1/tutorials/non-ml/pdes.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r1/tutorials/non-ml/pdes.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
> Note: This is an archived TF1 notebook. These are configured
to run in TF2's
[compatbility mode](https://www.tensorflow.org/guide/migrate)
but will run in TF1 as well. To use TF1 in Colab, use the
[%tensorflow_version 1.x](https://colab.research.google.com/notebooks/tensorflow_version.ipynb)
magic.
TensorFlow isn't just for machine learning. Here you will use TensorFlow to simulate the behavior of a [partial differential equation](https://en.wikipedia.org/wiki/Partial_differential_equation). You'll simulate the surface of a square pond as a few raindrops land on it.
## Basic setup
A few imports you'll need.
```
#Import libraries for simulation
import tensorflow.compat.v1 as tf
import numpy as np
#Imports for visualization
import PIL.Image
from io import BytesIO
from IPython.display import clear_output, Image, display
```
A function for displaying the state of the pond's surface as an image.
```
def DisplayArray(a, fmt='jpeg', rng=[0,1]):
"""Display an array as a picture."""
a = (a - rng[0])/float(rng[1] - rng[0])*255
a = np.uint8(np.clip(a, 0, 255))
f = BytesIO()
PIL.Image.fromarray(a).save(f, fmt)
clear_output(wait = True)
display(Image(data=f.getvalue()))
```
Here you start an interactive TensorFlow session for convenience in playing around. A regular session would work as well if you were doing this in an executable .py file.
```
sess = tf.InteractiveSession()
```
## Computational convenience functions
```
def make_kernel(a):
"""Transform a 2D array into a convolution kernel"""
a = np.asarray(a)
a = a.reshape(list(a.shape) + [1,1])
return tf.constant(a, dtype=1)
def simple_conv(x, k):
"""A simplified 2D convolution operation"""
x = tf.expand_dims(tf.expand_dims(x, 0), -1)
y = tf.nn.depthwise_conv2d(x, k, [1, 1, 1, 1], padding='SAME')
return y[0, :, :, 0]
def laplace(x):
"""Compute the 2D laplacian of an array"""
laplace_k = make_kernel([[0.5, 1.0, 0.5],
[1.0, -6., 1.0],
[0.5, 1.0, 0.5]])
return simple_conv(x, laplace_k)
```
## Define the PDE
Our pond is a perfect 500 x 500 square, as is the case for most ponds found in nature.
```
N = 500
```
Here you create a pond and hit it with some rain drops.
```
# Initial Conditions -- some rain drops hit a pond
# Set everything to zero
u_init = np.zeros([N, N], dtype=np.float32)
ut_init = np.zeros([N, N], dtype=np.float32)
# Some rain drops hit a pond at random points
for n in range(40):
a,b = np.random.randint(0, N, 2)
u_init[a,b] = np.random.uniform()
DisplayArray(u_init, rng=[-0.1, 0.1])
```
Now you specify the details of the differential equation.
```
# Parameters:
# eps -- time resolution
# damping -- wave damping
eps = tf.placeholder(tf.float32, shape=())
damping = tf.placeholder(tf.float32, shape=())
# Create variables for simulation state
U = tf.Variable(u_init)
Ut = tf.Variable(ut_init)
# Discretized PDE update rules
U_ = U + eps * Ut
Ut_ = Ut + eps * (laplace(U) - damping * Ut)
# Operation to update the state
step = tf.group(
U.assign(U_),
Ut.assign(Ut_))
```
## Run the simulation
This is where it gets fun -- running time forward with a simple for loop.
```
# Initialize state to initial conditions
tf.global_variables_initializer().run()
# Run 1000 steps of PDE
for i in range(1000):
# Step simulation
step.run({eps: 0.03, damping: 0.04})
# Show final image
DisplayArray(U.eval(), rng=[-0.1, 0.1])
```
Look! Ripples!
| github_jupyter |
# Classical Music Recommendation Playground
This notebook will show how to implement simple recommender system follwing two different approaches: **Collaborative Filtering** (user based) and **Content Based** recommendation.
> DISCLAIMER:
> The used dataset is NOT a real dataset, but it has been artificially generated for the Tutorial purposes.
> It absolutely should NOT be used as training data for any application.
```
import pandas as pd
import numpy as np
import scipy.spatial.distance as distance
```
## Data import
### data.csv
User listening experience dataset. 100 **users** -- numeric identifiers from 0 to 99 -- interact (or not) with 100 **items** (classical composers).
1 = interaction, 0 = no interaction (implicit feedback)
```
data = pd.read_csv('data.csv', index_col=0)
data.head()
```
Here a (quite-predictable) list of the top 10 most popular composer.
```
data.sum().sort_values(ascending=False)[0:10]
```
### artists.csv
The 100 involved artists with label, uris and 17 embedding dimension coming from the [music embeddings repo](https://github.com/DOREMUS-ANR/music-embeddings).
```
artists = pd.read_csv('artists.csv', index_col=0)
artists.head()
```
## Utils
```
# returns the list of items which a given user has interacted to
def get_items(id):
user = data.loc[id]
return user[user[:] == 1].axes[0].tolist()
# example user 7
get_items(7)
# retrieve the embeddings for an artists given a label
def get_emb(label):
a = artists.loc[artists['label'] == label]
embs = a.drop('label', axis=1).values[0]
return np.ma.array(embs, mask=embs == -2.)
# example
get_emb('Wolfgang Amadeus Mozart')
```
## Collaborative filtering
```
# Find most similar users to the given one
def most_similar_users(user, k):
user_vec = user.values # user listening
# search among all the other users
pool = data.drop(user.name)
# apply the cosine distance to each element of the pool, and sort accordingly
pool['distance'] = data.apply(lambda u: distance.cosine(user_vec, u.values), axis=1)
pool = pool.sort_values('distance').drop('distance', 1)
# return the first k users
return pool[:k]
# Select the most popular artists among a subset of users
def most_popular_among(user_subset, k=10):
return user_subset.sum().sort_values(ascending=False).index.tolist()
# Recommend artists by looking at similar users
def collaborative_filtering(user, k=10):
_user = data.loc[user]
# find k most similar users
similar_users = most_similar_users(_user, k)
# get k closest items
most_popular = most_popular_among(similar_users)
# remove the ones already in the list
prediction = [x for x in most_popular if x not in get_items(user)]
return prediction[:k]
```
Which artist would be recommended to our user? He already listened these ones:
```
user_example = 8
get_items(user_example)
```
The recommendation proposes other Germans composers
```
collaborative_filtering(user_example)
```
## Content based recommendation
Define similarity metric
```
def compute_similarity(seed, target, w=1):
b1 = np.where(seed.mask==True)[0]
b2 = np.where(target.mask==True)[0]
bad_pos = np.unique(np.concatenate([b1, b2]))
_seed = np.delete(seed, bad_pos, axis=0)
_target = np.delete(target, bad_pos, axis=0)
_w = np.delete(w, bad_pos, axis=0)
if len(_seed) == 0:
return 0
# distance
d = weighted_l2(_seed, _target, _w)
# how much info I am not finding
penalty = len([x for x in b2 if x not in b1]) / len(seed)
# score
s = (max_distance - d) / max_distance
return s * (1 - penalty)
def weighted_l2(a, b, w=1):
q = a - b
return np.sqrt((w * q * q).sum())
_ones = np.ones(17)
max_distance = weighted_l2(_ones,-_ones, _ones)
```
Compute all the similarity scores between couple of artists and put them in a Data Frame.
```
similarity_matrix = pd.DataFrame(index=artists['label'], columns=artists['label'])
for i in np.arange(len(similarity_matrix)):
seed = artists.iloc[i]['label']
for j in np.arange(len(similarity_matrix)):
if i == j:
similarity_matrix.iloc[i][j] = 1
continue
target = artists.iloc[j]['label']
similarity_matrix.iloc[i][j] = compute_similarity(get_emb(seed), get_emb(target))
similarity_matrix.head()
def content_based(user, k=10):
_items = get_items(user)
# remove the items already in the list
candidates = similarity_matrix.drop(labels=_items, axis=1)
# choose the artists that maximise the similarity among all the items
candidates = candidates.loc[_items]
return candidates.sum().sort_values(ascending=False).index.tolist()[0:k]
get_items(user_example)
content_based(user_example)
```
## Bonus
What happens with a user that appreciate just a particular composer?
```
new_user = np.zeros(len(data.loc[0]))
new_user_id = len(data)
data.loc[new_user_id] = new_user
data.loc[new_user_id]['Antonio Vivaldi'] = 1
data.loc[[new_user_id]]
collaborative_filtering(new_user_id)
content_based(new_user_id)
```
| github_jupyter |
```
import numpy as np
class LinearRegression:
def __init__(self, fit_intercept=True):
self.coef_ = None
self.intercept_ = None
self._fit_intercept = fit_intercept
def fit(self, X, y):
"""Fit model coefficients.
Arguments:
X -- 1D or 2D numpy array
y -- 1D numpy array
"""
# check if X is 1D or 2D array
if len(X.shape) == 1:
X = X.reshape(-1,1)
# add bias if fit_intercept
if self._fit_intercept:
X = np.c_[np.ones(X.shape[0]), X]
# closed form solution
xTx = np.dot(X.T, X)
inverse_xTx = np.linalg.inv(xTx)
xTy = np.dot(X.T, y)
coef = np.dot(inverse_xTx, xTy)
# set attributes
if self._fit_intercept:
self.intercept_ = coef[0]
self.coef_ = coef[1:]
else:
self.intercept_ = 0
self.coef_ = coef
def predict(self, X):
"""Output model prediction.
Arguments:
X -- 1D or 2D numpy array
"""
# check if X is 1D or 2D array
if len(X.shape) == 1:
X = X.reshape(-1,1)
return np.dot(X, self.coef_) + self.intercept_
class Metrics:
def mean_squared_error(self, y_true, y_predicted):
error = np.subtract(y_true, y_predicted)
squared_error = error ** 2
return np.mean(squared_error)
def root_mean_squared_error(self, y_true, y_predicted):
return np.sqrt(mean_squared_error)
def mean_absolute_error(self, y_true, y_predicted):
error = np.subtract(y_true, y_predicted)
abs_error = np.abs(error)
return np.mean(abs_error)
class MachineLearning(LinearRegression, Metrics):
def __init__(self, fit_intercept=True):
self.coef_ = None
self.intercept_ = None
self._fit_intercept = fit_intercept
ml = MachineLearning()
ml.fit(data, target)
target[:10]
ml.mean_squared_error(target, ml.predict(data))
ml.mean_absolute_error(target, ml.predict(data))
ml.root_mean_squared_error(target, ml.predict(data))
class LinearRegression(MachineLearning):
def __init__(self, fit_intercept=True):
self.coef_ = None
self.intercept_ = None
self._fit_intercept = fit_intercept
def fit(self, X, y):
"""Fit model coefficients.
Arguments:
X -- 1D or 2D numpy array
y -- 1D numpy array
"""
# check if X is 1D or 2D array
if len(X.shape) == 1:
X = X.reshape(-1,1)
# add bias if fit_intercept
if self._fit_intercept:
X = np.c_[np.ones(X.shape[0]), X]
# closed form solution
xTx = np.dot(X.T, X)
inverse_xTx = np.linalg.inv(xTx)
xTy = np.dot(X.T, y)
coef = np.dot(inverse_xTx, xTy)
# set attributes
if self._fit_intercept:
self.intercept_ = coef[0]
self.coef_ = coef[1:]
else:
self.intercept_ = 0
self.coef_ = coef
def predict(self, X):
"""Output model prediction.
Arguments:
X -- 1D or 2D numpy array
"""
# check if X is 1D or 2D array
if len(X.shape) == 1:
X = X.reshape(-1,1)
return np.dot(X, self.coef_) + self.intercept_
```
## Testing
```
from sklearn.datasets import load_boston
boston = load_boston()
data = boston.data
target = boston.target
lr = LinearRegression(fit_intercept=True)
lr.fit(data, target)
lr.coef_
lr.intercept_
lr.metrics.mean_squared_error(target, lr.predict(data))
from sklearn.linear_model import LinearRegression
lrsk = LinearRegression(fit_intercept=True)
lrsk.fit(data, target)
lrsk.coef_
lrsk.intercept_
from sklearn.metrics import mean_squared_error, mean_absolute_error
mean_squared_error(target, lrsk.predict(data))
mean_absolute_error(target, lrsk.predict(data))
np.sqrt(mean_squared_error(target, lrsk.predict(data)))
```
| github_jupyter |
```
import numpy
import imp
from re import sub
import logging
from datetime import datetime, timedelta
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import os
try:
changed
except NameError:
os.chdir('..')
os.system('find -name *.pyc | xargs rm')
changed = True
import sys
stdout = sys.stdout
from hyperstream import HyperStream, UTC, TimeInterval, __version__
sys.stdout = stdout
%matplotlib inline
%load_ext autoreload
%autoreload 2
sns.set_style("dark")
minute = timedelta(minutes=1)
t1 = datetime(2016, 4, 28, 20, 0, 0, 0, UTC)
t2 = t1 + minute
ti = TimeInterval(t1, t2)
module_file_old = '/Users/td6301/dev/IRC-SPHERE_public/HyperStream/hyperstream/tools/percentiles_from_list/2016-12-02_v0.0.1.py'
module_file_new = '/Users/td6301/dev/IRC-SPHERE_public/HyperStream/hyperstream/tools/percentiles_from_list/2017-07-18_v0.0.2.py'
print(__version__)
hs = HyperStream()
def load_module(module_file):
module_file_components = module_file[:-3].split('/')
with open(module_file, 'rb') as fp:
module_name = '_'.join(map(lambda pp: sub(r'[^a-zA-Z0-9]', '_', pp), module_file_components))
if module_name in sys.modules:
print("module {} already loaded ...".format(module_name))
del sys.modules[module_name]
mod = imp.load_module(
module_name, fp, module_file,
('.py', 'rb', imp.PY_SOURCE)
)
return mod
# Generate some random date
dg = hs.plugins.data_generators
ticker = hs.channel_manager.memory.get_or_create_stream('ticker')
random = hs.channel_manager.memory.get_or_create_stream('random')
hs.tools.clock().execute(sources=[], sink=ticker, interval=ti)
dg.tools.random(seed=1234).execute(sources=[], sink=random, interval=ti, alignment_stream=ticker)
print(random.window().values()[:5])
def print_percentiles(stream_old, stream_new):
data_old = stream_old.window().items()
data_new = stream_new.window().items()
for i, (old, new) in enumerate(zip(data_old, data_new)):
print('{0:2.0f}, {1:2.0f}, {2}, {3}, {4}'.format(
old.timestamp.minute, old.timestamp.second, old.value, new.value, np.array_equal(old.value, new.value)))
if not np.array_equal(old.value, new.value):
raise
# Force loading of tool modules
mod_old = load_module(module_file_old)
mod_new = load_module(module_file_new)
q = [25, 50, 75]
p_old = mod_old.PercentilesFromList(percentiles=q)
p_new = mod_new.PercentilesFromList(percentiles=q)
perc_old = hs.channel_manager.memory.get_or_create_stream('perc_old')
perc_new = hs.channel_manager.memory.get_or_create_stream('perc_new')
hs.channel_manager.memory.purge_stream(perc_old.stream_id)
hs.channel_manager.memory.purge_stream(perc_new.stream_id)
p_old.execute(sources=[random], sink=perc_old, interval=ti)
p_new.execute(sources=[random], sink=perc_new, interval=ti)
print_percentiles(perc_old, perc_new)
# Force loading of tool modules
mod_old = load_module(module_file_old)
mod_new = load_module(module_file_new)
p_old = mod_old.PercentilesFromList(n_segments=11)
p_new = mod_new.PercentilesFromList(n_segments=11)
perc_old = hs.channel_manager.memory.get_or_create_stream('perc_old')
perc_new = hs.channel_manager.memory.get_or_create_stream('perc_new')
hs.channel_manager.memory.purge_stream(perc_old.stream_id)
hs.channel_manager.memory.purge_stream(perc_new.stream_id)
p_old.execute(sources=[random], sink=perc_old, interval=ti)
p_new.execute(sources=[random], sink=perc_new, interval=ti)
print_percentiles(perc_old, perc_new)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import os
import re
```
# Encoding of categorical variables
In this notebook, we will present typical ways of dealing with
**categorical variables** by encoding them, namely **ordinal encoding** and
**one-hot encoding**.
Let us first load the entire adult dataset containing both numerical and catagorical data
```
adult_census = pd.read_pickle('../data/adult_census.pkl')
adult_census = adult_census.drop(columns = 'education-num')
target_name = 'class'
target = adult_census[target_name]
data = adult_census.drop(columns = [target_name])
```
## Identify categorical variables
As we saw in the previous section, a numerical variable is a
quantity represented by a real or integer number. These variables can be
naturally handled by machine learning algorithms that are typically composed
of a sequence of arithmetic instructions such as additions and
multiplications.
In contrast, categorical variables have discrete values, typically
represented by string labels (but not only) taken from a finite list of
possible choices. For instance, the variable `native-country` in our dataset
is a categorical variable because it encodes the data using a finite list of
possible countries (along with the `?` symbol when this information is
missing):
```
data['native-country'].value_counts().sort_index()
```
How can we easily recognize categorical columns among the dataset? Part of
the answer lies in the columns' data type:
```
data.dtypes
```
If we look at the `"native-country"` column, we observe its data type is
`object`, meaning it contains string values.
## Select features based on their data type
In the previous notebook, we manually defined the numerical columns. We could
do a similar approach. Instead, we will use the scikit-learn helper function
`make_column_selector`, which allows us to select columns based on
their data type. We will illustrate how to use this helper.
```
from sklearn.compose import make_column_selector
categorical_columns_selector = make_column_selector(dtype_include = object)
categorical_columns = categorical_columns_selector(data)
categorical_columns
```
Here, we created the selector by passing the data type to include; we then
passed the input dataset to the selector object, which returned a list of
column names that have the requested data type. We can now filter out the
unwanted columns:
```
data_categorical = data[categorical_columns]
data_categorical.head()
print(f"The Categorical dataset is composed of {data_categorical.shape[1]} features")
```
In the remainder of this section, we will present different strategies to
encode categorical data into numerical data which can be used by a
machine-learning algorithm.
## Strategies to encode categories
### Encoding ordinal categories
The most intuitive strategy is to encode each category with a different
number. The `OrdinalEncoder` will transform the data in such manner.
We will start by encoding a single column to understand how the encoding
works.
```
from sklearn.preprocessing import OrdinalEncoder
education_column = data_categorical[['education']]
encoder = OrdinalEncoder()
education_encoded = encoder.fit_transform(education_column)
education_encoded
```
We see that each category in "education" has been replaced by a numeric value. We could check the mapping between the categories and the numerical values by checking the fitted attribute `categories_`.
```
encoder.categories_
education_column
data_encoded = encoder.fit_transform(data_categorical)
data_encoded
encoder.categories_
print(
f"The dataset encoded contains {data_encoded.shape[1]} features")
```
We see that the categories have been encoded for each feature (column)
independently. We also note that the number of features before and after the
encoding is the same.
However, be careful when applying this encoding strategy:
using this integer representation leads downstream predictive models
to assume that the values are ordered (0 < 1 < 2 < 3... for instance).
By default, `OrdinalEncoder` uses a lexicographical strategy to map string
category labels to integers. This strategy is arbitrary and often
meaningless. For instance, suppose the dataset has a categorical variable
named `"size"` with categories such as "S", "M", "L", "XL". We would like the
integer representation to respect the meaning of the sizes by mapping them to
increasing integers such as `0, 1, 2, 3`.
However, the lexicographical strategy used by default would map the labels
"S", "M", "L", "XL" to 2, 1, 0, 3, by following the alphabetical order.
The `OrdinalEncoder` class accepts a `categories` constructor argument to
pass categories in the expected ordering explicitly. You can find more
information in the
[scikit-learn documentation](https://scikit-learn.org/stable/modules/preprocessing.html#encoding-categorical-features)
if needed.
If a categorical variable does not carry any meaningful order information
then this encoding might be misleading to downstream statistical models and
you might consider using one-hot encoding instead (see below).
### Encoding nominal categories (without assuming any order)
`OneHotEncoder` is an alternative encoder that prevents the downstream
models to make a false assumption about the ordering of categories. For a
given feature, it will create as many new columns as there are possible
categories. For a given sample, the value of the column corresponding to the
category will be set to `1` while all the columns of the other categories
will be set to `0`.
We will start by encoding a single feature (e.g. `"education"`) to illustrate
how the encoding works.
```
from sklearn.preprocessing import OneHotEncoder
encoder = OneHotEncoder(sparse = False)
education_encoded = encoder.fit_transform(education_column)
education_encoded
```
<div class="admonition note alert alert-info">
<p class="first admonition-title" style="font-weight: bold;">Note</p>
<p><tt class="docutils literal">sparse=False</tt> is used in the <tt class="docutils literal">OneHotEncoder</tt> for didactic purposes, namely
easier visualization of the data.</p>
<p class="last">Sparse matrices are efficient data structures when most of your matrix
elements are zero. They won't be covered in details in this course. If you
want more details about them, you can look at
<a class="reference external" href="https://scipy-lectures.org/advanced/scipy_sparse/introduction.html#why-sparse-matrices">this</a>.</p>
</div>
We see that encoding a single feature will give a NumPy array full of zeros
and ones. We can get a better understanding using the associated feature
names resulting from the transformation.
```
feature_names = encoder.get_feature_names(input_features=['education'])
education_encoded = pd.DataFrame(education_encoded, columns = feature_names)
education_encoded
```
As we can see, each category (unique value) became a column; the encoding
returned, for each sample, a 1 to specify which category it belongs to.
Let's apply this encoding on the full dataset.
```
print(
f"The dataset is composed of {data_categorical.shape[1]} features")
data_categorical.head()
data_encoded = encoder.fit_transform(data_categorical)
data_encoded[:5]
data_encoded.shape
print(f'The encoded dataset has got {data_encoded.shape[1]} features')
```
Let's wrap this NumPy array in a dataframe with informative column names as
provided by the encoder object:
```
columns_encoded = encoder.get_feature_names(data_categorical.columns)
pd.DataFrame(data_encoded, columns = columns_encoded).head()
```
Look at how the `"workclass"` variable of the 3 first records has been
encoded and compare this to the original string representation.
The number of features after the encoding is more than 10 times larger than
in the original data because some variables such as `occupation` and
`native-country` have many possible categories.
### Choosing an encoding strategy
Choosing an encoding strategy will depend on the underlying models and the
type of categories (i.e. ordinal vs. nominal).
Indeed, using an `OrdinalEncoder` will output ordinal categories. It means
that there is an order in the resulting categories (e.g. `0 < 1 < 2`). The
impact of violating this ordering assumption is really dependent on the
downstream models. Linear models will be impacted by misordered categories
while tree-based models will not be.
Thus, in general `OneHotEncoder` is the encoding strategy used when the
downstream models are **linear models** while `OrdinalEncoder` is used with
**tree-based models**.
You still can use an `OrdinalEncoder` with linear models but you need to be
sure that:
- the original categories (before encoding) have an ordering;
- the encoded categories follow the same ordering than the original
categories.
The next exercise highlight the issue of misusing `OrdinalEncoder` with a
linear model.
Also, there is no need to use an `OneHotEncoder` even if the original
categories do not have an given order with tree-based model. It will be
the purpose of the final exercise of this sequence.
## Evaluate our predictive pipeline
We can now integrate this encoder inside a machine learning pipeline like we
did with numerical data: let's train a linear classifier on the encoded data
and check the statistical performance of this machine learning pipeline using
cross-validation.
Before we create the pipeline, we have to linger on the `native-country`.
Let's recall some statistics regarding this column.
```
data['native-country'].value_counts()
```
We see that the `Holand-Netherlands` category is occurring rarely. This will
be a problem during cross-validation: if the sample ends up in the test set
during splitting then the classifier would not have seen the category during
training and will not be able to encode it.
In scikit-learn, there are two solutions to bypass this issue:
* list all the possible categories and provide it to the encoder via the
keyword argument `categories`;
* use the parameter `handle_unknown`.
Here, we will use the latter solution for simplicity.
<div class="admonition tip alert alert-warning">
<p class="first admonition-title" style="font-weight: bold;">Tip</p>
<p class="last">Be aware the <tt class="docutils literal">OrdinalEncoder</tt> exposes as well a parameter
<tt class="docutils literal">handle_unknown</tt>. It can be set to <tt class="docutils literal">use_encoded_value</tt> and by setting
<tt class="docutils literal">unknown_value</tt> to handle rare categories. You are going to use these
parameters in the next exercise.</p>
</div>
We can now create our machine learning pipeline.
```
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import LogisticRegression
model = make_pipeline(
OneHotEncoder(handle_unknown='ignore'), LogisticRegression(max_iter = 500)
)
```
<div class="admonition note alert alert-info">
<p class="first admonition-title" style="font-weight: bold;">Note</p>
<p class="last">Here, we need to increase the maximum number of iterations to obtain a fully
converged <tt class="docutils literal">LogisticRegression</tt> and silence a <tt class="docutils literal">ConvergenceWarning</tt>. Contrary
to the numerical features, the one-hot encoded categorical features are all
on the same scale (values are 0 or 1), so they would not benefit from
scaling. In this case, increasing <tt class="docutils literal">max_iter</tt> is the right thing to do.</p>
</div>
```
from sklearn.model_selection import cross_validate
cv_results = cross_validate(model, data_categorical, target)
cv_results
scores = cv_results['test_score']
print(f'The accuracy id: {scores.mean():.3f} +/- {scores.std():.3f}')
```
As you can see, this representation of the categorical variables is
slightly more predictive of the revenue than the numerical variables
that we used previously.
In this notebook we have:
* seen two common strategies for encoding categorical features: **ordinal
encoding** and **one-hot encoding**;
* used a **pipeline** to use a **one-hot encoder** before fitting a logistic
regression.
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df=pd.read_csv('only_road_accidents_data3.csv')
df.head()
print(df['STATE/UT'].unique())
df2=df[df.columns[2:10]].sum(axis=0)
#print(df2)
df2.plot.pie(title='Time slot distribution of all accidents in India(2001-14)',autopct='%1.1f%%')
df3=df.loc[:,['STATE/UT','Total']].groupby(['STATE/UT']).sum(axis=0)
#df3.plot.bar(title="State wise accidents")
plt.figure(figsize=(15,5))
df3.plot(kind='bar')
plt.xlabel("STATE/UT")
plt.ylabel("Number of Road Accidents ")
plt.title("Number of Road Accidents in State/UT")
plt.xticks(rotation=90)
df4=df.loc[:,['YEAR','Total']].groupby(['YEAR']).sum(axis=0)
df4.plot.bar(title="Year wise number of accidents")
def top_n_state_over_year(top, year):
df_state = df.loc[:, ['YEAR', 'STATE/UT', 'Total']]
df_state = df_state[df_state['YEAR'] == year].groupby(['STATE/UT']).sum()\
.sort_values(by=['YEAR', 'Total'], ascending=[True,False]).head(top)
plt.figure(figsize=(10,5))
sns.barplot(x="STATE/UT", y="Total", data=df_state)
plt.title(f"Top {top} States/UT (Road Accidents in Year {year})")
plt.xlabel("STATE/UT")
plt.ylabel("Number of Road Accidents")
plt.xticks(rotation=90)
return;
top_n_state_over_year(5, 2001)
def least_n_state_over_year(least, year):
df_state = df.loc[:, ['YEAR', 'STATE/UT', 'Total']]
df_state = df_state[df_state['YEAR'] == year].groupby(['STATE/UT']).sum()\
.sort_values(by=['YEAR', 'Total'], ascending=[True,True]).head(least)
plt.figure(figsize=(10,5))
sns.barplot(x="STATE/UT", y="Total", data=df_state)
plt.title(f"Least {least} States/UT (Road Accidents in Year {year})")
plt.xlabel("STATE/UT")
plt.ylabel("Number of Road Accidents")
plt.xticks(rotation=90)
return;
least_n_state_over_year(5, 2001)
UT=['A & N Islands' ,'Chandigarh' , 'D & N Haveli' ,'Daman & Diu' ,'Delhi (Ut)','Jammu & Kashmir', 'Lakshadweep', 'Puducherry']
df7=df.loc[df['STATE/UT'].isin(UT)]
df7=df7.loc[:,['STATE/UT','Total']].groupby(['STATE/UT'])['Total'].sum().reset_index()
index=np.arange(len(UT))
plt.figure(figsize=(15,5))
plt.barh(index,df7['Total'])
plt.ylabel('Union Territories')
plt.xlabel('Accidents')
plt.title(' Accidents in union territories')
plt.yticks(index,UT )
for index, value in enumerate(df7['Total']):
plt.text(value, index, str(value))
plt.show()
North=['Haryana', 'Himachal Pradesh' ,'Jammu & Kashmir','Punjab' ,'Uttar Pradesh', 'Uttarakhand']
East=['Bihar', 'Jharkhand', 'Odisha', 'West Bengal']
South=[ 'Andhra Pradesh' , 'Karnataka', 'Kerala', 'Tamil Nadu']
West=[ 'Goa', 'Gujarat','Maharashtra' , 'Rajasthan']
Central=['Chhattisgarh' ,'Madhya Pradesh']
North_East=[ 'Assam', 'Manipur', 'Meghalaya', 'Mizoram' ,'Nagaland', 'Sikkim' ,'Tamil Nadu', 'Tripura']
df8=df.loc[df['STATE/UT'].isin(North)]
x1=df8.loc[:,['Total']].sum(axis=0)
df8=df.loc[df['STATE/UT'].isin(East)]
x2=df8.loc[:,['Total']].sum(axis=0)
df8=df.loc[df['STATE/UT'].isin(West)]
x3=df8.loc[:,['Total']].sum(axis=0)
df8=df.loc[df['STATE/UT'].isin(South)]
x4=df8.loc[:,['Total']].sum(axis=0)
df8=df.loc[df['STATE/UT'].isin(North_East)]
x5=df8.loc[:,['Total']].sum(axis=0)
df8=df.loc[df['STATE/UT'].isin(Central)]
x6=df8.loc[:,['Total']].sum(axis=0)
y=[x1['Total'],x2['Total'],x3['Total'],x4['Total'],x5['Total'],x6['Total']]
x=('North','East','West','South','North_East','Central' )
index = np.arange(len(x))
plt.bar(index, y,label="Total Accidents")
plt.xlabel('Regions')
plt.ylabel('Accidents')
plt.title('Region-wise Accidents')
plt.xticks(index,x )
plt.legend()
plt.tight_layout()
plt.show()
morning=df["3-6 hrs. (Night)"] .sum()+ df["6-9 hrs (Day)"].sum()
noon=df["9-12 hrs (Day)"].sum()+ df["12-15 hrs (Day)"].sum()
evening=df["15-18 hrs (Day)"].sum()+ df["18-21 hrs (Night)"].sum()
night=df["21-24 hrs (Night)"].sum()+ df["0-3 hrs. (Night)"].sum()
y=(morning,noon,evening,night)
x=('Morning','Noon','Evening','Night')
index=np.arange(len(x))
plt.bar(index, y,label="Total Accidents")
plt.xlabel('Time slot')
plt.ylabel('Accidents')
plt.title('Time slot wise accidents')
plt.xticks(index,x )
plt.legend()
plt.tight_layout()
plt.show()
```
| github_jupyter |
# Description
A Colab notebook for generating images using OpenAI's CLIP model.
Heavily influenced by Alexander Mordvintsev's Deep Dream, this work uses CLIP to match an image learned by a SIREN network with a given textual description.
As a good launching point for future directions and to find more related work, see https://distill.pub/2017/feature-visualization/
If you have questions, please see my twitter at https://twitter.com/advadnoun
This is all free! But if you're feeling generous, you can donate to my venmo @rynnn while your "a beautiful Waluigi" loads ;)
# Top
Import the usual libraries
```
import torch
import numpy as np
import torchvision
import torchvision.transforms.functional as TF
import PIL
import matplotlib.pyplot as plt
import os
import random
import imageio
from IPython import display
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import glob
from google.colab import output
```
Check which GPU Colab has assigned to you
```
!nvidia-smi -L
```
# Restart after running this cell!
You must run this cell and then restart and rerun everything for the PyTorch version to be correct. Otherwise the model will run but not produce any meaningful output.
```
import subprocess
CUDA_version = [s for s in subprocess.check_output(["nvcc", "--version"]).decode("UTF-8").split(", ") if s.startswith("release")][0].split(" ")[-1]
print("CUDA version:", CUDA_version)
if CUDA_version == "10.0":
torch_version_suffix = "+cu100"
elif CUDA_version == "10.1":
torch_version_suffix = "+cu101"
elif CUDA_version == "10.2":
torch_version_suffix = ""
else:
torch_version_suffix = "+cu110"
! pip install torch==1.7.1{torch_version_suffix} torchvision==0.8.2{torch_version_suffix} -f https://download.pytorch.org/whl/torch_stable.html ftfy regex
```
# CLIP
Setup CLIP and set it to be the "perceptor" used to determine the loss for the SIREN network.
Thanks to the authors below & OpenAI for sharing! https://github.com/openai/CLIP
Alec Radford \* Jong Wook Kim \* Chris Hallacy Aditya Ramesh Gabriel Goh Sandhini Agarwal
Girish Sastry Amanda Askell Pamela Mishkin Jack Clark Gretchen Krueger
Ilya Sutskever
```
%cd /content/
!git clone https://github.com/openai/CLIP.git
%cd /content/CLIP/
!pip install ftfy
import clip
import numpy as np
# Load the model
perceptor, preprocess = clip.load('ViT-B/32')
```
# Params
Determine the output dimensions of the image and the number of channels.
Set the text to be matched
```
im_shape = [512, 512, 3]
sideX, sideY, channels = im_shape
tx = clip.tokenize("a beautiful Waluigi")
```
# Define
Define some helper functions
```
def displ(img, pre_scaled=True):
img = np.array(img)[:,:,:]
img = np.transpose(img, (1, 2, 0))
if not pre_scaled:
img = scale(img, 48*4, 32*4)
imageio.imwrite(str(3) + '.png', np.array(img))
return display.Image(str(3)+'.png')
def card_padded(im, to_pad=3):
return np.pad(np.pad(np.pad(im, [[1,1], [1,1], [0,0]],constant_values=0), [[2,2], [2,2], [0,0]],constant_values=1),
[[to_pad,to_pad], [to_pad,to_pad], [0,0]],constant_values=0)
```
# SIREN
Thanks to the authors of SIREN! https://github.com/vsitzmann/siren
@inproceedings{sitzmann2019siren,
author = {Sitzmann, Vincent
and Martel, Julien N.P.
and Bergman, Alexander W.
and Lindell, David B.
and Wetzstein, Gordon},
title = {Implicit Neural Representations
with Periodic Activation Functions},
booktitle = {arXiv},
year={2020}
}
The number of layers is 8 right now, but if the machine OOMs (runs out of RAM), it can naturally be tweaked. I've found that 16 layers for the SIREN works best, but I'm not always blessed with a V100 GPU.
```
import torch.nn as nn
class SineLayer(nn.Module):
# See paper sec. 3.2, final paragraph, and supplement Sec. 1.5 for discussion of omega_0.
# If is_first=True, omega_0 is a frequency factor which simply multiplies the activations before the
# nonlinearity. Different signals may require different omega_0 in the first layer - this is a
# hyperparameter.
# If is_first=False, then the weights will be divided by omega_0 so as to keep the magnitude of
# activations constant, but boost gradients to the weight matrix (see supplement Sec. 1.5)
def __init__(self, in_features, out_features, bias=True,
is_first=False, omega_0=30):
super().__init__()
self.omega_0 = omega_0
self.is_first = is_first
self.in_features = in_features
self.linear = nn.Linear(in_features, out_features, bias=bias)
self.init_weights()
def init_weights(self):
with torch.no_grad():
if self.is_first:
self.linear.weight.uniform_(-1 / self.in_features,
1 / self.in_features)
else:
self.linear.weight.uniform_(-np.sqrt(6 / self.in_features) / self.omega_0,
np.sqrt(6 / self.in_features) / self.omega_0)
def forward(self, input):
return torch.sin(self.omega_0 * self.linear(input))
def forward_with_intermediate(self, input):
# For visualization of activation distributions
intermediate = self.omega_0 * self.linear(input)
return torch.sin(intermediate), intermediate
class Siren(nn.Module):
def __init__(self, in_features, hidden_features, hidden_layers, out_features, outermost_linear=True,
first_omega_0=30, hidden_omega_0=30.):
super().__init__()
self.net = []
self.net.append(SineLayer(in_features, hidden_features,
is_first=True, omega_0=first_omega_0))
for i in range(hidden_layers):
self.net.append(SineLayer(hidden_features, hidden_features,
is_first=False, omega_0=hidden_omega_0))
if outermost_linear:
final_linear = nn.Linear(hidden_features, out_features)
with torch.no_grad():
final_linear.weight.uniform_(-np.sqrt(6 / hidden_features) / hidden_omega_0,
np.sqrt(6 / hidden_features) / hidden_omega_0)
self.net.append(final_linear)
else:
self.net.append(SineLayer(hidden_features, out_features,
is_first=False, omega_0=hidden_omega_0))
self.net = nn.Sequential(*self.net)
def forward(self, coords):
coords = coords.clone().detach().requires_grad_(True)
output = self.net(coords.cuda())
return output.view(1, sideX, sideY, 3).permute(0, 3, 1, 2)#.sigmoid_()
def forward_with_activations(self, coords, retain_grad=False):
'''Returns not only model output, but also intermediate activations.
Only used for visualizing activations later!'''
activations = OrderedDict()
activation_count = 0
x = coords.clone().detach().requires_grad_(True)
activations['input'] = x
for i, layer in enumerate(self.net):
if isinstance(layer, SineLayer):
x, intermed = layer.forward_with_intermediate(x)
if retain_grad:
x.retain_grad()
intermed.retain_grad()
activations['_'.join((str(layer.__class__), "%d" % activation_count))] = intermed
activation_count += 1
else:
x = layer(x)
if retain_grad:
x.retain_grad()
activations['_'.join((str(layer.__class__), "%d" % activation_count))] = x
activation_count += 1
return activations
def get_mgrid(sidelen, dim=2):
'''Generates a flattened grid of (x,y,...) coordinates in a range of -1 to 1.
sidelen: int
dim: int'''
tensors = tuple(dim * [torch.linspace(-1, 1, steps=sidelen)])
mgrid = torch.stack(torch.meshgrid(*tensors), dim=-1)
mgrid = mgrid.reshape(-1, dim)
return mgrid
model = Siren(2, 256, 16, 3).cuda()
LLL = []
eps = 0
optimizer = torch.optim.Adam(model.parameters(), .00001)
make sure you're on this page when train first starts running, and it dings
otherwise the thing'll never start
```
# Train
Train and output samples every 150 iterations
We create batches of images at different resolutions in different parts of the SIREN image and resize them with bilinear upsampling. This seems to work very, very well as regularization for visualizing networks with larger images than their usual input resolution.
```
def checkin(loss):
print(loss)
with torch.no_grad():
al = nom(model(get_mgrid(sideX)).cpu()).numpy()
for allls in al:
displ(allls)
display.display(display.Image(str(3)+'.png'))
print('\n')
output.eval_js('new Audio("https://freesound.org/data/previews/80/80921_1022651-lq.ogg").play()')
def ascend_txt():
out = model(get_mgrid(sideX))
cutn = 64
p_s = []
for ch in range(cutn):
size = torch.randint(int(.5*sideX), int(.98*sideX), ())
offsetx = torch.randint(0, sideX - size, ())
offsety = torch.randint(0, sideX - size, ())
apper = out[:, :, offsetx:offsetx + size, offsety:offsety + size]
apper = torch.nn.functional.interpolate(apper, (224,224), mode='bilinear')
p_s.append(nom(apper))
into = torch.cat(p_s, 0)
iii = perceptor.encode_image(into)
t = perceptor.encode_text(tx.cuda())
return -100*torch.cosine_similarity(t, iii, dim=-1).mean()
def train(epoch, i):
loss = ascend_txt()
optimizer.zero_grad()
loss.backward()
optimizer.step()
if itt % 150 == 0:
checkin(loss)
nom = torchvision.transforms.Normalize((0.48145466, 0.4578275, 0.40821073), (0.26862954, 0.26130258, 0.27577711))
itt = 0
for epochs in range(10000):
for i in range(1000):
train(eps, i)
itt+=1
eps+=1
```
# Bot
| github_jupyter |
## Load Data
```
!wget http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz
!tar xfz ./aclImdb_v1.tar.gz
import os
import numpy as np
def get_files(dir):
return [dir + d for d in os.listdir(dir) if os.path.isfile(dir + d)]
train_dir = '/content/aclImdb/train/'
test_dir = '/content/aclImdb/test/'
train_pos_files = get_files(train_dir + 'pos/')
train_neg_files = get_files(train_dir + 'neg/')
test_pos_files = get_files(test_dir + 'pos/')
test_neg_files = get_files(test_dir + 'neg/')
def get_data(files):
temp = []
for c in files:
with open(c, 'r') as f:
temp.append(f.read())
return temp
train_pos = get_data(train_pos_files)
train_neg = get_data(train_neg_files)
test_pos = get_data(test_pos_files)
test_neg = get_data(test_neg_files)
```
## Data Processing
<ul>
<li><b>Convert to lower case</b>
<li><b>Remove Punctuation</b>
</ul>
```
from string import punctuation
def preprocess(data):
for i in range(len(data)):
s = data[i]
s = s.replace('<br />', '')
data[i] = ''.join([c for c in s.lower() if c not in punctuation])
return data
train_pos = preprocess(train_pos)
train_neg = preprocess(train_neg)
test_pos = preprocess(test_pos)
test_neg = preprocess(test_neg)
```
## Tokenize
<li><b>Create word to index mapping dictionary</b>
There is a small trick here, in this mapping index will start from 0 i.e. mapping of ‘the’ will be 0. But later on we are going to do padding for shorter reviews and conventional choice for padding is 0. So we need to start this indexing from 1
```
from collections import Counter
words = ' '.join(train_pos + train_neg + test_pos + test_neg)
words = words.split()
count_words = Counter(words)
total = len(words)
sorted_words = count_words.most_common(total)
word2idx = {w:i+1 for i,(w,c) in enumerate(sorted_words)}
```
<li><b>Encode the words
```
def encode_words(data):
data_idx = []
for review in data:
data_idx.append([word2idx[w] for w in review.split()])
return data_idx
train_pos_idx = encode_words(train_pos)
train_neg_idx = encode_words(train_neg)
test_pos_idx = encode_words(test_pos)
test_neg_idx = encode_words(test_neg)
train_pos_labels = np.full(len(train_pos_idx), 1).tolist()
train_neg_labels = np.full(len(train_neg_idx), 0).tolist()
test_pos_labels = np.full(len(test_pos_idx), 1).tolist()
test_neg_labels = np.full(len(test_neg_idx), 0).tolist()
train_labels = train_pos_labels + train_neg_labels
test_labels = np.array(test_pos_labels + test_neg_labels)
train_reviews = train_pos + train_neg
train_reviews_idx = train_pos_idx + train_neg_idx
test_reviews_idx = test_pos_idx + test_neg_idx
train_reviews_idx = train_reviews_idx[:11250] + train_reviews_idx[13750:]
val_reviews_idx = train_reviews_idx[11250:13750]
train_labels = np.array(train_labels[:11250] + train_labels[13750:])
val_labels = np.array(train_labels[11250:13750])
```
## Analyze Reviews Length
```
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
reviews_len = [len(x) for x in train_reviews]
pd.Series(reviews_len).hist()
plt.show()
pd.Series(reviews_len).describe()
```
<li><b>Padding / Truncating the remaining data
```
import numpy as np
def pad_features(reviews, seq_length):
'''
Return features of review_ints, where each review is padded with 0's or truncated to the input seq_length.
'''
features = np.zeros((len(reviews), seq_length), dtype = int)
for i, review in enumerate(reviews):
review_len = len(review)
if review_len <= seq_length:
zeroes = list(np.zeros(seq_length-review_len))
new = zeroes+review
elif review_len > seq_length:
new = review[0:seq_length]
features[i,:] = np.array(new)
return features
seq_length = 200
train_features = pad_features(train_reviews_idx, seq_length)
val_features = pad_features(val_reviews_idx, seq_length)
test_features = pad_features(test_reviews_idx, seq_length)
```
## Dataloaders and Batching
```
import torch
from torch.utils.data import DataLoader, TensorDataset
# create Tensor datasets
train_data = TensorDataset(torch.from_numpy(train_features), torch.from_numpy(train_labels))
val_data = TensorDataset(torch.from_numpy(val_features), torch.from_numpy(val_labels))
test_data = TensorDataset(torch.from_numpy(test_features), torch.from_numpy(test_labels))
# dataloaders
batch_size = 50
# make sure to SHUFFLE your data
train_loader = DataLoader(train_data, shuffle=True, batch_size=batch_size)
val_loader = DataLoader(val_data, shuffle=True, batch_size=batch_size)
test_loader = DataLoader(test_data, shuffle=True, batch_size=batch_size)
# obtain one batch of training data
dataiter = iter(train_loader)
sample_x, sample_y = dataiter.next()
print('Sample input size: ', sample_x.size()) # batch_size, seq_length
print('Sample input: \n', sample_x)
print()
print('Sample label size: ', sample_y.size()) # batch_size
print('Sample label: \n', sample_y)
```
## LSTM Model
```
import torch.nn as nn
class SentimentLSTM(nn.Module):
def __init__(self, vocab_size, output_size, embedding_dim, hidden_dim, n_layers, drop_prob=0.5):
super(SentimentLSTM, self).__init__()
self.output_size = output_size
self.n_layers = n_layers
self.hidden_dim = hidden_dim
self.embedding = nn.Embedding(vocab_size, embedding_dim)
self.lstm = nn.LSTM(embedding_dim, hidden_dim, n_layers, dropout=drop_prob, batch_first=True)
self.dropout = nn.Dropout(0.3)
self.fc = nn.Linear(hidden_dim, output_size)
self.sig = nn.Sigmoid()
def forward(self, x, hidden):
batch_size = x.size(0)
embeds =self.embedding(x)
lstm_out, hidden = self.lstm(embeds, hidden)
# stack up lstm outputs
lstm_out = lstm_out.contiguous().view(-1, self.hidden_dim)
out = self.dropout(lstm_out)
out = self.fc(out)
# sigmoid function
sig_out = self.sig(out)
# reshape to be batch_size first
sig_out = sig_out.view(batch_size, -1)
sig_out = sig_out[:, -1] # get last batch of labels
# return last sigmoid output and hidden state
return sig_out, hidden
def init_hidden(self, batch_size):
''' Initializes hidden state '''
# Create two new tensors with sizes n_layers x batch_size x hidden_dim,
# initialized to zero, for hidden state and cell state of LSTM
weight = next(self.parameters()).data
if (train_on_gpu):
hidden = (weight.new(self.n_layers, batch_size, self.hidden_dim).zero_().cuda(),
weight.new(self.n_layers, batch_size, self.hidden_dim).zero_().cuda())
else:
hidden = (weight.new(self.n_layers, batch_size, self.hidden_dim).zero_(),
weight.new(self.n_layers, batch_size, self.hidden_dim).zero_())
return hidden
```
## Training
```
# Instantiate the model w/ hyperparams
vocab_size = len(word2idx) + 1 # +1 for the 0 padding
output_size = 1
embedding_dim = 400
hidden_dim = 256
n_layers = 2
train_on_gpu = True
net = SentimentLSTM(vocab_size, output_size, embedding_dim, hidden_dim, n_layers)
lr=0.001
epochs = 4
counter = 0
print_every = 100
clip=5 # gradient clipping
criterion = nn.BCELoss()
optimizer = torch.optim.Adam(net.parameters(), lr=lr)
if(train_on_gpu):
net.cuda()
net.train()
# train for some number of epochs
for e in range(epochs):
# initialize hidden state
h = net.init_hidden(batch_size)
# batch loop
for inputs, labels in train_loader:
counter += 1
if(train_on_gpu):
inputs, labels = inputs.cuda(), labels.cuda()
# Creating new variables for the hidden state, otherwise
# we'd backprop through the entire training history
h = tuple([each.data for each in h])
# zero accumulated gradients
net.zero_grad()
# get the output from the model
inputs = inputs.type(torch.LongTensor).cuda()
output, h = net(inputs, h)
# calculate the loss and perform backprop
loss = criterion(output.squeeze(), labels.float())
loss.backward()
# `clip_grad_norm` helps prevent the exploding gradient problem in RNNs / LSTMs.
nn.utils.clip_grad_norm_(net.parameters(), clip)
optimizer.step()
# loss stats
if counter % print_every == 0:
# Get validation loss
val_h = net.init_hidden(batch_size)
val_losses = []
net.eval()
for inputs, labels in val_loader:
# Creating new variables for the hidden state, otherwise
# we'd backprop through the entire training history
val_h = tuple([each.data for each in val_h])
if(train_on_gpu):
inputs, labels = inputs.cuda(), labels.cuda()
inputs = inputs.type(torch.LongTensor).cuda()
output, val_h = net(inputs, val_h)
val_loss = criterion(output.squeeze(), labels.float())
val_losses.append(val_loss.item())
net.train()
print("Epoch: {}/{}...".format(e+1, epochs),
"Step: {}...".format(counter),
"Loss: {:.6f}...".format(loss.item()),
"Val Loss: {:.6f}".format(np.mean(val_losses)))
```
## Testing
```
# Get test data loss and accuracy
test_losses = [] # track loss
num_correct = 0
# init hidden state
h = net.init_hidden(batch_size)
net.eval()
# iterate over test data
for inputs, labels in test_loader:
# Creating new variables for the hidden state, otherwise
# we'd backprop through the entire training history
h = tuple([each.data for each in h])
if(train_on_gpu):
inputs, labels = inputs.cuda(), labels.cuda()
# get predicted outputs
inputs = inputs.type(torch.LongTensor).cuda()
output, h = net(inputs, h)
# calculate loss
test_loss = criterion(output.squeeze(), labels.float())
test_losses.append(test_loss.item())
# convert output probabilities to predicted class (0 or 1)
pred = torch.round(output.squeeze()) # rounds to the nearest integer
# compare predictions to true label
correct_tensor = pred.eq(labels.float().view_as(pred))
correct = np.squeeze(correct_tensor.numpy()) if not train_on_gpu else np.squeeze(correct_tensor.cpu().numpy())
num_correct += np.sum(correct)
# -- stats! -- ##
# avg test loss
print("Test loss: {:.3f}".format(np.mean(test_losses)))
# accuracy over all test data
test_acc = num_correct/len(test_loader.dataset)
print("Test accuracy: {:.3f}".format(test_acc))
```
## On user-generated data
```
from string import punctuation
def tokenize_review(test_review):
test_review = test_review.lower() # lowercase
# get rid of punctuation
test_text = ''.join([c for c in test_review if c not in punctuation])
# splitting by spaces
test_words = test_text.split()
# tokens
test_ints = []
test_ints.append([vocab_to_int[word] for word in test_words])
return test_ints
# test code and generate tokenized review
test_ints = tokenize_review(test_review_neg)
print(test_ints)
# test sequence padding
seq_length=200
features = pad_features(test_ints, seq_length)
print(features)
# test conversion to tensor and pass into your model
feature_tensor = torch.from_numpy(features)
print(feature_tensor.size())
def predict(net, test_review, sequence_length=200):
net.eval()
# tokenize review
test_ints = tokenize_review(test_review)
# pad tokenized sequence
seq_length=sequence_length
features = pad_features(test_ints, seq_length)
# convert to tensor to pass into your model
feature_tensor = torch.from_numpy(features)
batch_size = feature_tensor.size(0)
# initialize hidden state
h = net.init_hidden(batch_size)
if(train_on_gpu):
feature_tensor = feature_tensor.cuda()
# get the output from the model
output, h = net(feature_tensor, h)
# convert output probabilities to predicted class (0 or 1)
pred = torch.round(output.squeeze())
# printing output value, before rounding
print('Prediction value, pre-rounding: {:.6f}'.format(output.item()))
# print custom response
if(pred.item()==1):
print("Positive review detected!")
else:
print("Negative review detected.")
```
| github_jupyter |
#Part 4 BERT for arithmetic sentiment analysis
Acknowledgement: We used most of the code from https://mccormickml.com/2019/07/22/BERT-fine-tuning/
Most Credit to:
Chris McCormick and Nick Ryan
# Bert Background
**B**idirectional **E**ncoder **R**epresentations from
**T**ransformers (BERT) [Devlin et al., 2019], as the name suggests, is a language model based on the Transformer [Vaswani et al., 2017] encoder architecture that has been pre-trained on a large dataset of unlabeled sentences from Wikipedia and BookCorpus [Zhu et al., 2015]. Given a sequence of tokens representing sentence(s), BERT outputs a ``contextualized representation" vector for each of the token. Now, suppose we are given some down-stream tasks, such as sentence classification or question-answering. We can take the BERT model, add a small layer on top of the BERT representation(s), and then fine-tune the added parameters **and** BERT parameters on the down-stream dataset, which is typically much smaller than the data used to pre-train BERT.
In traditional language modeling task, the objective is to maximize the log likelihood of predicting the current word (or token) in the sentence, given the previous words (to the left of current work) as context. This is called the *autoregressive model*. In BERT, however, we wish to predict the current word given both the words before and after (i.e. to the left and to the right) of the sentence--hence *bidirectional*.
To be able to attend from both directions, BERT uses the encoder Transformer, which does not apply any attention masking unlike the decoder.
We briefly describe how BERT is pre-trained. BERT has 2 task objectives for pre-training: (1) *Masked Language Modeling* (Masked LM), and (2) *Next Sentence Prediction*(NSP). The input to the model is a sequence of tokens of the form:
```
[CLS] Sentence A [SEP] Sentence B,
```
where `[CLS]` ("class") and `[SEP]` ("separator") are special tokens.
In Masked LM, some percentage of the input tokens are converted into `[MASK]` tokens, and the objective is to use the final layer representation for that masked token to predict the correct word that was masked out. For NSP, the task is to use the contextualized representation for the `[CLS]` token to perform binary classification for whether sentence A and sentence B are consecutive sentences in the unlabeled dataset. See Figure 6 (in Handout) for the conceptual picture of BERT pre-training and fine-tuning.
In this assignment, you will be **fine-tuning BERT on a single sentence classification task** (see below about the dataset). Figure 7 (in Handout) illustrates the architecture for fine-tuning on this task. We prepend the tokenized sentence with the `[CLS]` token, then feed the sequence into BERT. We then take the contextualized `[CLS]` token representation at the last layer of BERT and add either a softmax layer on top corresponding to the number of output classes in the task. Alternatively, we can have fully connected hidden layers before the softmax layer for more expressivity for harder tasks. Then, both the new layers and the entire BERT parameters are trained end to end on the task for a few epochs.
# 1. Setup
## Install transformers repo that has Bert
```
!pip install transformers
```
## Download & Extract
Run the following cells to downlaod the dataset files from the CSC413 webpage.
<!-- Download the two csv dataset files from CSC413 webpage, click the folder icon, -->
<!-- and click "upload" to upload them. -->
<!-- https://csc413-2020.github.io/assets/misc/PA03_data_20_train.csv
https://csc413-2020.github.io/assets/misc/PA03_data_20_test.csv -->
```
!pip install wget
import wget
import os
print('Downloading verbal arithmetic dataset')
# The URL for the dataset zip file.
url = 'https://csc413-2020.github.io/assets/misc/'
# Download the file (if we haven't already)
if not os.path.exists('./PA03_data_20_train.csv'):
wget.download(url + 'PA03_data_20_train.csv', './PA03_data_20_train.csv')
print('Done downloading training data')
else:
print('Already downloaded training data')
if not os.path.exists('./PA03_data_20_test.csv'):
wget.download(url + 'PA03_data_20_test.csv', './PA03_data_20_test.csv')
print('Done downloading test data')
else:
print('Already downloaded test data')
```
## Load Training Data
```
import pandas as pd
df = pd.read_csv("./PA03_data_20_train.csv", header=0, names=["index", "input", "label"])
print("Number of data points: ", df.shape[0])
sampled = df.sample(10)
# Display 10 random rows from the data.
df.sample(10)
```
The two properties we actually care about are the the `inputs` and its `label`, which are the questions and the answers.
**label=0** means the result of expression is **negative**
**label=1** means the result of expression is **zero**
**label=2** means the result of expression is **positive**
## BERT Tokenizer
To feed our text to BERT, it must be split into tokens, and then these tokens must be mapped to their index in the tokenizer vocabulary.
```
from transformers import BertTokenizer
# Load the BERT tokenizer.
print('Loading BERT tokenizer...')
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased', do_lower_case=True)
inputs = df.input.values
labels = df.label.values
print("Train data size ", len(inputs))
print(' Original: ', inputs[0])
# Print the sentence split into tokens.
print('Tokenized: ', tokenizer.tokenize(inputs[0]))
# Print the sentence mapped to token ids.
print('Token IDs: ', tokenizer.convert_tokens_to_ids(tokenizer.tokenize(inputs[0])))
```
We can actually use the `tokenize.encode` function to handle both steps, rather than calling `tokenize` and `convert_tokens_to_ids` separately.
## BERT Required Formatting
In a deep learning based NLP pipeline, most of the following preprocessing tricks are frequently needed regardless of whether we use BERT or RNN.
1. Add special tokens to the start and end of each sentence.
2. Pad & truncate all sentences to a single constant length.
3. Explicitly differentiate real tokens from padding tokens with the "attention mask".
### Special Tokens
**`[SEP]`**
At the end of every sentence, we need to append the special `[SEP]` token.
This token is an artifact of two-sentence tasks, where BERT is given two separate sentences and asked to determine something (e.g., can the answer to the question in sentence A be found in sentence B?).
**`[CLS]`**
For classification tasks, we must prepend the special `[CLS]` token to the beginning of every sentence.
This token has special significance. BERT consists of 12 Transformer layers. Each transformer takes in a list of token embeddings, and produces the same number of embeddings on the output.
On the output of the final transformer, *only the first embedding (corresponding to the [CLS] token) is used by the classifier*.
> "The first token of every sequence is always a special classification token (`[CLS]`). The final hidden state
corresponding to this token is used as the aggregate sequence representation for classification
tasks." (from the [BERT paper](https://arxiv.org/pdf/1810.04805.pdf))
Also, because BERT is trained to only use this [CLS] token for classification, we know that the model has been motivated to encode everything it needs for the classification step into that single 768-value embedding vector.
### Sentence Length & Attention Mask
The sentences in our dataset obviously have varying lengths, so how does BERT handle this?
BERT has two constraints:
1. All sentences must be padded or truncated to a single, fixed length.
2. The maximum sentence length is 512 tokens.
Padding is done with a special `[PAD]` token, which is at index 0 in the BERT vocabulary.
The "Attention Mask" is simply an array of 1s and 0s indicating which tokens are padding and which aren't
## Sentences to IDs
The `tokenizer.encode` function combines multiple steps for us:
1. Split the sentence into tokens.
2. Add the special `[CLS]` and `[SEP]` tokens.
3. Map the tokens to their IDs.
Oddly, this function can perform truncating for us, but doesn't handle padding.
```
# For Verbal Arithmetic
input_ids = []
# For every sentence...
for input in inputs:
# `encode` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
encoded_input = tokenizer.encode(
input, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
# This function also supports truncation and conversion
# to pytorch tensors, but we need to do padding, so we
# can't use these features :( .
#max_length = 128, # Truncate all sentences.
#return_tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_input)
# Print sentence 0, now as a list of IDs.
print('Original: ', inputs[0])
print('Token IDs:', input_ids[0])
```
## Padding & Truncating
Pad and truncate our sequences so that they all have the same length, `MAX_LEN`.
First, what's the maximum sentence length in our dataset?
```
print('Max sentence length: ', max([len(sen) for sen in input_ids]))
```
Given that, let's choose MAX_LEN = 7 since our numerical expression is quite short. Then apply the padding.
```
# We'll borrow the `pad_sequences` utility function to do this.
from keras.preprocessing.sequence import pad_sequences
# Set the maximum sequence length.
MAX_LEN = 7
print('\nPadding/truncating all sentences to %d values...' % MAX_LEN)
print('\nPadding token: "{:}", ID: {:}'.format(tokenizer.pad_token, tokenizer.pad_token_id))
# Pad our input tokens with value 0.
# "post" indicates that we want to pad and truncate at the end of the sequence,
# as opposed to the beginning.
input_ids = pad_sequences(input_ids, maxlen=MAX_LEN, dtype="long",
value=0, truncating="post", padding="post")
print('\nDone.')
```
## Attention Masks
The attention mask simply makes it explicit which tokens are actual words versus which are padding.
The BERT vocabulary does not use the ID 0, so if a token ID is 0, then it's padding, and otherwise it's a real token.
```
# Create attention masks
attention_masks = []
# For each sentence...
for sent in input_ids:
# Create the attention mask.
# - If a token ID is 0, then it's padding, set the mask to 0.
# - If a token ID is > 0, then it's a real token, set the mask to 1.
att_mask = [int(token_id > 0) for token_id in sent]
# Store the attention mask for this sentence.
attention_masks.append(att_mask)
```
## Training & Validation Split
Divide up our training set to use 90% for training and 10% for validation.
```
# Use train_test_split to split our data into train and validation sets for
# training
from sklearn.model_selection import train_test_split
# Use 90% for training and 10% for validation.
print(input_ids)
train_inputs, validation_inputs, train_labels, validation_labels = train_test_split(input_ids, labels,
random_state=2018, test_size=0.1)
# Do the same for the masks.
train_masks, validation_masks, _, _ = train_test_split(attention_masks, labels,
random_state=2018, test_size=0.1)
set(labels)
```
## Converting to PyTorch Data Types
Our model expects PyTorch tensors rather than numpy.ndarrays, so convert all of our dataset variables.
```
import torch
train_inputs = torch.tensor(train_inputs)
validation_inputs = torch.tensor(validation_inputs)
train_labels = torch.tensor(train_labels)
validation_labels = torch.tensor(validation_labels)
train_masks = torch.tensor(train_masks)
validation_masks = torch.tensor(validation_masks)
```
We'll also create an iterator for our dataset using the torch DataLoader class. This helps save on memory during training because, unlike a for loop, with an iterator the entire dataset does not need to be loaded into memory.
```
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
batch_size = 32
# Create the DataLoader for our training set.
train_data = TensorDataset(train_inputs, train_masks, train_labels)
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=batch_size)
# Create the DataLoader for our validation set.
validation_data = TensorDataset(validation_inputs, validation_masks, validation_labels)
validation_sampler = SequentialSampler(validation_data)
validation_dataloader = DataLoader(validation_data, sampler=validation_sampler, batch_size=batch_size)
```
# 4. Questions
## Question1 [0pts]
The pre-trained neural network here is the normal BERT model from [BertForSequenceClassification](https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#bertforsequenceclassification). The goal is to add a new classification layer to the pre-trained model. We have provided two example classes to do so.
In this part, you need to make your own `BertCSC413_MLP` class `self.classifier` by, for example, modifying the provided examples: change the number of layers; change the number of hidden neurons; or try a different activation.
```
from transformers import BertForSequenceClassification
import torch.nn as nn
class BertCSC413_Linear(BertForSequenceClassification):
def __init__(self, config):
super(BertCSC413_Linear, self).__init__(config)
self.classifier = nn.Linear(config.hidden_size, self.config.num_labels)
class BertCSC413_MLP_Example(BertForSequenceClassification):
def __init__(self, config):
super(BertCSC413_MLP_Example, self).__init__(config)
self.classifier = nn.Sequential(
nn.Linear(config.hidden_size, config.hidden_size),
nn.ReLU(),
nn.Linear(config.hidden_size, self.config.num_labels)
)
class BertCSC413_MLP(BertForSequenceClassification):
def __init__(self, config):
super(BertCSC413_MLP, self).__init__(config)
# Your own classifier goes here
self.classifier = nn.Sequential(
nn.Linear(config.hidden_size, config.hidden_size),
nn.ReLU(),
nn.Linear(config.hidden_size, config.hidden_size),
nn.ReLU(),
nn.Linear(config.hidden_size, self.config.num_labels)
)
```
## Question2 [0pts]
We instantiated two different BERT models from `BertCSC413_MLP` class, which are called `model_freeze_bert` and `model_finetune_bert` in the notebook.
**Run** the following code to train the models, and attach the training error curves of `model_freeze_bert` and `model_finetune_bert`.
```
from transformers import AdamW, BertConfig
model_freeze_bert = BertCSC413_MLP.from_pretrained(
"bert-base-uncased",
num_labels = 3,
output_attentions = False,
output_hidden_states = False,
)
for name, param in model_freeze_bert.named_parameters():
if 'classifier' not in name: # classifier layer
param.requires_grad = False
model_finetune_bert = BertCSC413_MLP.from_pretrained(
"bert-base-uncased",
num_labels = 3,
output_attentions = False,
output_hidden_states = False,
)
# Model parameters visualization
params = list(model_finetune_bert.named_parameters())
print('The BERT model has {:} different named parameters.\n'.format(len(params)))
print('==== Embedding Layer ====\n')
for p in params[0:5]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== First Transformer ====\n')
for p in params[5:21]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== Output Layer ====\n')
for p in params[-4:]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
```
We use
- Batch size: 32
- Learning rate (Adam): 2e-5
- Number of epochs: 4
```
from transformers import get_linear_schedule_with_warmup
import random
import numpy as np
import time
import datetime
seed_val = 42
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
def format_time(elapsed):
elapsed_rounded = int(round((elapsed)))
return str(datetime.timedelta(seconds=elapsed_rounded))
def train_model(model, learning_rate, epoch_nums):
optimizer = AdamW(model.parameters(),
lr = learning_rate, #2e-5, # args.learning_rate - default is 5e-5, our notebook had 2e-5
eps = 1e-8 # args.adam_epsilon - default is 1e-8.
)
epochs = epoch_nums
total_steps = len(train_dataloader) * epochs
scheduler = get_linear_schedule_with_warmup(optimizer,
num_warmup_steps = 0,
num_training_steps = total_steps)
loss_values = []
for epoch_i in range(0, epochs):
print("")
print('======== Epoch {:} / {:} ========'.format(epoch_i + 1, epochs))
print('Training...')
t0 = time.time()
total_loss = 0
model.train()
for step, batch in enumerate(train_dataloader):
if step % 40 == 0 and not step == 0:
elapsed = format_time(time.time() - t0)
# Report progress.
print(' Batch {:>5,} of {:>5,}. Elapsed: {:}.'.format(step, len(train_dataloader), elapsed))
b_input_ids = batch[0] #.to(device)
b_input_mask = batch[1] #.to(device)
b_labels = batch[2] #.to(device)
model.zero_grad()
# Perform a forward pass (evaluate the model on this training batch).
# This will return the loss (rather than the model output) because we
# have provided the `labels`.
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
outputs = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels)
# The call to `model` always returns a tuple, so we need to pull the
# loss value out of the tuple.
loss = outputs[0]
# Accumulate the training loss over all of the batches so that we can
# calculate the average loss at the end. `loss` is a Tensor containing a
# single value; the `.item()` function just returns the Python value
# from the tensor.
total_loss += loss.item()
# Perform a backward pass to calculate the gradients.
loss.backward()
# Clip the norm of the gradients to 1.0.
# This is to help prevent the "exploding gradients" problem.
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
# Update parameters and take a step using the computed gradient.
# The optimizer dictates the "update rule"--how the parameters are
# modified based on their gradients, the learning rate, etc.
optimizer.step()
# Update the learning rate.
scheduler.step()
# Calculate the average loss over the training data.
avg_train_loss = total_loss / len(train_dataloader)
# Store the loss value for plotting the learning curve.
loss_values.append(avg_train_loss)
print("")
print(" Average training loss: {0:.2f}".format(avg_train_loss))
print(" Training epcoh took: {:}".format(format_time(time.time() - t0)))
print("Running Validation...")
t0 = time.time()
model.eval()
eval_loss, eval_accuracy = 0, 0
nb_eval_steps, nb_eval_examples = 0, 0
# Evaluate data for one epoch
for batch in validation_dataloader:
# batch = tuple(t.to(device) for t in batch)
batch = tuple(t for t in batch)
b_input_ids, b_input_mask, b_labels = batch
with torch.no_grad():
# Forward pass, calculate logit predictions.
# This will return the logits rather than the loss because we have
# not provided labels.
# token_type_ids is the same as the "segment ids", which
# differentiates sentence 1 and 2 in 2-sentence tasks.
outputs = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask)
# Get the "logits" output by the model. The "logits" are the output
# values prior to applying an activation function like the softmax.
logits = outputs[0]
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Calculate the accuracy for this batch of test sentences.
tmp_eval_accuracy = flat_accuracy(logits, label_ids)
# Accumulate the total accuracy.
eval_accuracy += tmp_eval_accuracy
# Track the number of batches
nb_eval_steps += 1
print(" Accuracy: {0:.2f}".format(eval_accuracy/nb_eval_steps))
print(" Validation took: {:}".format(format_time(time.time() - t0)))
print("")
print("Training complete!")
return loss_values
freeze_bert_loss_vals = train_model(model_freeze_bert, 2e-5, 4) # about 1 minute for 4 epochs using CPU
finttune_bert_loss_vals = train_model(model_finetune_bert, 2e-5, 4) # about 5 minutes for 4 epochs using CPU
import matplotlib.pyplot as plt
% matplotlib inline
import seaborn as sns
def plot_loss_vals(loss_vals):
sns.set(style='darkgrid')
sns.set(font_scale=1.5)
plt.rcParams["figure.figsize"] = (12,6)
plt.plot(loss_vals, 'b-o')
plt.title("Training loss")
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.show()
plot_loss_vals(freeze_bert_loss_vals)
plot_loss_vals(finttune_bert_loss_vals)
```
*Run* the evaluation functions. Report the test performances of using trained model\_freeze\_bert and model\_finetune\_bert, and briefly discuss why models are failing under certain target labels
```
import pandas as pd
# Load the dataset into a pandas dataframe.
#df = pd.read_csv("./output.txt_test.csv", delimiter='\t', header=None, names=['sentence_source', 'label', 'label_notes', 'sentence'])
df = pd.read_csv("./PA03_data_20_test.csv", header=0, names=["index", "input", "label"])
# Report the number of sentences.
print('Number of test sentences: {:,}\n'.format(df.shape[0]))
inputs = df.input.values
labels = df.label.values
# Tokenize all of the sentences and map the tokens to thier word IDs.
input_ids = []
input_sent = []
# For every sentence...
for arithmetic_input in inputs:
# `encode` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
encoded_sent = tokenizer.encode(
arithmetic_input,
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
)
input_sent.append(arithmetic_input)
input_ids.append(encoded_sent)
# Pad our input tokens
input_ids = pad_sequences(input_ids, maxlen=MAX_LEN,
dtype="long", truncating="post", padding="post")
# Create attention masks
attention_masks = []
# Create a mask of 1s for each token followed by 0s for padding
for seq in input_ids:
seq_mask = [float(i>0) for i in seq]
attention_masks.append(seq_mask)
# Convert to tensors.
prediction_inputs = torch.tensor(input_ids)
prediction_masks = torch.tensor(attention_masks)
prediction_labels = torch.tensor(labels)
# Set the batch size.
batch_size = 32
# Create the DataLoader.
prediction_data = TensorDataset(prediction_inputs, prediction_masks, prediction_labels)
prediction_sampler = SequentialSampler(prediction_data)
prediction_dataloader = DataLoader(prediction_data, sampler=prediction_sampler, batch_size=batch_size)
def eval_testdata(model_test, show_all_predictions=False):
print('Predicting labels for {:,} test sentences...'.format(len(prediction_inputs)))
# Put model in evaluation mode
model_test.eval()
# Tracking variables
predictions , true_labels, input_sents = [], [], []
# Predict
for batch in prediction_dataloader:
# Unpack the inputs from our dataloader
b_input_ids, b_input_mask, b_labels = batch
decoded_inputs = [tokenizer.decode(b_input_ids[i]).strip("[CLS] ").strip( "[SEP] ").strip(" [PAD] ").strip("[PAD]").strip(" [SE") for i in range(len(b_input_ids))]
# Telling the model not to compute or store gradients, saving memory and
# speeding up prediction
with torch.no_grad():
# Forward pass, calculate logit predictions
outputs = model_test(b_input_ids, token_type_ids=None,
attention_mask=b_input_mask)
logits = outputs[0]
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
input_sents.extend(decoded_inputs)
# Store predictions and true labels
predictions.extend(np.argmax(logits, axis=1))
true_labels.extend(label_ids)
correct = [0,0,0]
totals = [0, 0, 0]
for true_label, prediction in zip(true_labels, predictions):
if true_label == prediction:
correct[true_label] += 1
totals[true_label] += 1
print("Number of expressions with negative result", true_labels.count(0), "\n", correct[0], " predicted correctly", ", accuracy ", correct[0]/totals[0] , "\n")
print("Number of expressions with 0 result", true_labels.count(1), "\n", correct[1], " predicted correctly", ", accuracy ", correct[1]/totals[1], "\n")
print("Number of expressions with positive result", true_labels.count(2),"\n", correct[2], " predicted correctly",", accuracy ", correct[2]/totals[2], "\n")
if show_all_predictions:
index_to_sentiment_map = {0:"negative", 1:"zero", 2:"positive"}
for sent in [sent + "--> "+index_to_sentiment_map[index] for sent, index in zip(input_sents, predictions)]:
print(sent)
eval_testdata(model_freeze_bert, show_all_predictions=False)
eval_testdata(model_finetune_bert, show_all_predictions=False)
```
## Question3 [1pts]
Try a few unseen examples of arithmetic questions using either model\_freeze\_bert or model\_finetune\_bert model, and find 10 interesting results. We will give full marks as long as you provide some comments for why you chose some of the examples. The interesting results can, for example, be both successful extrapolation/interpolation results or surprising failure cases. You can find some examples in our notebook.
```
index_to_sentiment_map = {0:"negative", 1:"zero", 2:"positive"}
model = model_finetune_bert
def what_is(arithmetic_input):
encoded_sent = tokenizer.encode(
arithmetic_input,
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
)
input_sent = [arithmetic_input]
input_ids = [encoded_sent]
input_ids = pad_sequences(input_ids, maxlen=MAX_LEN,
dtype="long", truncating="post", padding="post")
attention_masks = []
for seq in input_ids:
seq_mask = [float(i>0) for i in seq]
attention_masks.append(seq_mask)
with torch.no_grad():
outputs = model(torch.tensor(input_ids), token_type_ids=None,
attention_mask=torch.tensor(attention_masks))
logits = outputs[0]
logits = logits.detach().cpu().numpy()
print(index_to_sentiment_map[np.argmax(logits, axis=1)[0]])
what_is("twelve minus fourteen")
what_is("twelve plus fourteen")
what_is("one plus hundred")
what_is("one minus hundred")
what_is("hundred minus one")
what_is("1 plus 100")
what_is("1 minus 14")
what_is("1 minus two")
what_is("two minus 3")
what_is("three minus two minus eight")
what_is("three minus two plus eight")
what_is("one minus lala")
what_is("one plus lala")
what_is("one minus lala plus lala")
what_is("ten minus lala")
```
## Question4 [1pts]
This is an open question, and we will give marks as long as you show an attempt to try one of the following tasks.
1. Try data augmentation tricks to improve the performances for certain target labels that models were failing to predict.
2. Make a t-sne or PCA plot to visualize the embedding vectors of word tokens related to arithmetic expressions.
3. Try different hyperparameter tunings. E.g. learning rates, optimizer, architecture of the classifier, training epochs, and batch size.
4. Evaluate the Multi-class Matthews correlation score for our imbalanced test dataset.
5. Run a baseline model using MLP without pre-trained BERT. You can assume the sequence length of all the data is 3 in this case.
```
finttune_bert_loss_vals = train_model(model_finetune_bert, 3e-5, 5)
plot_loss_vals(finttune_bert_loss_vals)
what_is("one plus one minus ten")
```
| github_jupyter |
## 第4章 Matplotlibでグラフを 描画しよう
### 4-7: 箱ひげ図
```
import matplotlib.pyplot as plt
# リスト4.7.1:箱ひげ図の描画
plt.style.use("ggplot")
x = [1, 2, 3, 3, 11, 20]
fig = plt.figure()
ax = fig.add_subplot(111)
ax.boxplot(x)
plt.show()
# リスト4.7.2:複数の箱ひげ図の描画
# 複数のリストをリストにセット
x = [[1, 2, 3, 3, 11, 20], [1, 2, 9, 10, 15, 16]]
labels = ["A", "B"]
fig = plt.figure()
ax = fig.add_subplot(111)
# データとラベルを指定
ax.boxplot(x, labels=labels)
plt.show()
# リスト4.7.3:anime_master.csvファイルの読み込み
import os
import pandas as pd
base_url = "https://raw.githubusercontent.com/practical-jupyter/sample-data/master/anime/"
anime_master_csv = os.path.join(base_url, "anime_master.csv")
df = pd.read_csv(anime_master_csv, index_col="anime_id")
df.head(3)
# リスト4.7.4:配給種別ごとのエピソード数を可視化
labels = []
types_list = []
# 配給種別ごとのエピソード数の情報をリスト化
for label, df_per_type in df.groupby("type"):
labels.append(label)
types_list.append(df_per_type["episodes"].tolist())
fig = plt.figure()
ax = fig.add_subplot(111)
ax.boxplot(types_list, labels=labels)
plt.show()
# リスト4.7.5:描画範囲を限定した箱ひげ図
fig = plt.figure(figsize=(8, 6))
ax = fig.add_subplot(111)
ax.boxplot(types_list, labels=labels)
# Y軸の描画範囲を0から100までに限定
ax.set_ylim(0, 100)
plt.show()
# リスト4.7.6:四分位点の値をdescribe()で確認
df["episodes"][df["type"] == "TV"].describe().round(1)
# リスト4.7.7:データセットの作成
import numpy as np
np.random.seed(3)
dataset = [np.random.normal(20 + mu, 5, 1000) for mu in range(1, 5)]
# リスト4.7.8:書式辞書の作成
# はずれ値の書式辞書
flierprop = {"color": "#EC407A",
"marker": "o",
"markerfacecolor": "#2196F3",
"markeredgecolor": "white",
"markersize": 5,
"linestyle": "None",
"linewidth": 0.1}
# 箱の書式辞書
boxprop = {"color": "#2196F3",
"facecolor": "#BBDEFB",
"linewidth": 1,
"linestyle": "-"}
# ヒゲの書式辞書
whiskerprop = {"color": "#2196F3",
"linewidth": 1,
"linestyle": "--"}
# ヒゲ末端キャップの書式辞書
capprop = {"color": "#2196F3",
"linewidth": 1,
"linestyle": ":"}
# 中央値の書式辞書
medianprop = {"color": "#2196F3",
"linewidth": 2,
"linestyle": "-"}
# 平均値の書式辞書
meanprop = {"color": "#2196F3",
"marker": "^",
"markerfacecolor": "#2196F3",
"markeredgecolor": "white",
"markersize": 10,
"linewidth": 1,
"linestyle": ""}
# リスト4.7.9:箱ひげ図の書式一括設定
fig = plt.figure()
ax = fig.add_subplot(111)
ax.boxplot(
dataset,
patch_artist="Patch", # 書式を設定する場合は 「Patch」 を選択
labels=["A", "B", "C", "D"], # 項目ラベル
showmeans=True, # 平均値の描画
flierprops=flierprop, # はずれ値の書式設定
boxprops=boxprop, # ボックスの書式設定
whiskerprops=whiskerprop, # ヒゲの書式設定
capprops=capprop, # ヒゲのキャップの書式設定
medianprops=medianprop, # 中央値の書式設定
meanprops=meanprop, # 平均値の書式設定
)
plt.show()
# リスト4.7.10:ボックスごとに書式を設定する
# 図の描画
fig = plt.figure()
ax = fig.add_subplot(111)
bp = ax.boxplot(
dataset,
patch_artist="Patch",
labels=["A", "B", "C", "D"],
meanline=True,
showmeans=True,
)
# カラーセット
colors1 = ["#2196F3", "#43A047", "#FBC02D", "#FB8C00"]
colors2 = ["#BBDEFB", "#C8E6C9", "#FFF9C4", "#FFE0B2"]
# 上下に分かれた要素に設定するための用の数列
n = [0, 0, 1, 1, 2, 2, 3, 3]
# 書式設定
# ボックスとはずれ値、 中央値の書式設定
for params in zip(bp["boxes"], bp["fliers"], bp["medians"], colors1, colors2):
bpb, bpf, med, color1, color2 = params
# ボックスの書式設定
bpb.set_color(color1)
bpb.set_facecolor(color2)
bpb.set_linewidth(2)
# 外れ値の書式設定
bpf.set(marker="^", color=color2)
bpf.set_markeredgecolor("white")
bpf.set_markerfacecolor(color1)
# 中央値の書式設定
med.set_color(color1)
med.set_linewidth(2)
# ヒゲとヒゲの末端キャップの書式設定
for bpc, bpw, m in zip(bp["caps"], bp["whiskers"], n):
bpc.set_color(colors1[m])
bpc.set_linewidth(2)
bpw.set_color(colors1[m])
bpw.set_linewidth(2)
# 平均値の書式設定
for mean, color2 in zip(bp["means"], colors2):
mean.set_color("grey")
mean.set_linewidth(2)
mean.set_linestyle("--")
plt.show()
```
| github_jupyter |
# Stochastic Gradient Descent Regression
This Code template is for regression analysis using the simple SGDRegressor based on the Stochastic Gradient Descent approach.
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.linear_model import SGDRegressor
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training.
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
### Model
Stochastic Gradient Descent (SGD) is a simple yet very efficient approach to fitting linear classifiers and regressors under convex loss functions such as (linear) Support Vector Machines and Logistic Regression. SGD is merely an optimization technique and does not correspond to a specific family of machine learning models. It is only a way to train a model. Often, an instance of SGDClassifier or SGDRegressor will have an equivalent estimator in the scikit-learn API, potentially using a different optimization technique.
For example, using SGDRegressor(loss='squared_loss', penalty='l2') and Ridge solve the same optimization problem, via different means.
#### Model Tuning Parameters
> - **loss** -> The loss function to be used. The possible values are ‘squared_loss’, ‘huber’, ‘epsilon_insensitive’, or ‘squared_epsilon_insensitive’
> - **penalty** -> The penalty (aka regularization term) to be used. Defaults to ‘l2’ which is the standard regularizer for linear SVM models. ‘l1’ and ‘elasticnet’ might bring sparsity to the model (feature selection) not achievable with ‘l2’.
> - **alpha** -> Constant that multiplies the regularization term. The higher the value, the stronger the regularization. Also used to compute the learning rate when set to learning_rate is set to ‘optimal’.
> - **l1_ratio** -> The Elastic Net mixing parameter, with 0 <= l1_ratio <= 1. l1_ratio=0 corresponds to L2 penalty, l1_ratio=1 to L1. Only used if penalty is ‘elasticnet’.
> - **tol** -> The stopping criterion
> - **learning_rate** -> The learning rate schedule,possible values {'optimal','constant','invscaling','adaptive'}
> - **eta0** -> The initial learning rate for the ‘constant’, ‘invscaling’ or ‘adaptive’ schedules.
> - **power_t** -> The exponent for inverse scaling learning rate.
> - **epsilon** -> Epsilon in the epsilon-insensitive loss functions; only if loss is ‘huber’, ‘epsilon_insensitive’, or ‘squared_epsilon_insensitive’.
```
model=SGDRegressor(random_state=123)
model.fit(x_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
score: The score function returns the coefficient of determination R2 of the prediction.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(x_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
plt.figure(figsize=(14,10))
plt.plot(range(20),y_test[0:20], color = "green")
plt.plot(range(20),model.predict(x_test[0:20]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Thilakraj Devadiga , Github: [Profile](https://github.com/Thilakraj1998)
| github_jupyter |
<a href="https://colab.research.google.com/github/keithvtls/Numerical-Method-Activities/blob/main/Lecture/Week%2015%20-%20Numerical%20Integration/NuMeth_6_Numerical_Integration.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Numerical Integration
$_{\text{©D.J. Lopez | 2021 | Computational Methods for Computer Engineers}}$
Reviving your integral calculus classes, we will be implementing the fundamental concepts of integration to computational and numerical methods. Numerical integration or the quadrature greatly helps, again, with the field of optimziation and estimation. This module will cover:
* The Trapezoidal Rule
* Simpson's 1/4 Integration Rule
* Simpson's 3/8 Integration Rule
* Monte Carlo Simulations/Integration
```
import numpy as np
```
## 6.1 Trapezoidal rule
The concept behind the Trapezoidal rule is a good review on what is integration and how it can be converted to its numerical and computational implementation.
Integration is usually defined as the area under a cruve or area of the function. Like the image below, integration is usually seen as the sum of the areas of the boxes (in this case trapezoids) that make up the area under the curve.
The Trapezoidal rule takes advantage of this concept by summing the areas of those trapezoids. If you would recall, the area of the Trapezoid is given as:
$$A_{trapz}=\frac{h(b-a)}{2} \\ _{\text{(Eq. 6.1)}}$$
Whereas $A_{trapz}$ is the area of a trapezoid, $a$ is the shorter base, $b$ is the longer base, and $h$ is the height of the Trapezoid. Use the image below as a visual reference.

In the trapezoidal rule, we can see that the trapezoids are right trapezoids. And we can formally construct the represtnative equation modelling the concept of the trapezoidal rule as:
$$\int^b_af(x)dx \approx h\left[ \frac{f(x_0)+f(x_n)}{2} +\sum^{n-1}_{i=1}f(x_i) \right]\\ _{\text{(Eq. 6.2)}}$$
For our example, we will mode the equation:
$$\int^{\frac{\pi}{2}}_0x\sin(x)dx = 1$$ and $$\int^{10}_0x^2dx = \frac{1000}{3}$$
```
f = lambda x : x*np.sin(x)
a, b = 0, np.pi/2
n = 5
h = (b-a)/n
A= (f(a)+f(b))/2
for i in range(1,n):
A += f(a+i*h)
S = h*A
S
h*(0.5*(f(a)+f(b))+np.sum(f(a+h*np.arange(1,n))))
def trapz_rule(func,lb,ub,size):
h = (ub-lb)/size
return h*(0.5*(func(lb)+func(ub))+np.sum(func(lb+h*np.arange(1,size))))
f = lambda x: x**2
sum = trapz_rule(f, 0,10,1e4)
sum
```
## Simpson's 1/3 Rule
Simpson's 1/3 Rule, unlike the Trapezoidal Rule, computes more than 2 strips of trapezoids at a time. And rather than trapezoids, Simpson's 1/3 rule uses parabolas ($P(x)$) in approximating areas under the curve.

The Simpson's 1/3 Rule cane be formulated as:
$$\int^b_af(x)dx \approx \frac{(b-a)}{6}\left(f(a)+4f\frac{(a+b)}{2}+f(b)\right)\\ _{\text{(Eq. 6.3)}}$$
It can be discretized as:
$$\int^b_af(x)dx \approx \frac{h}{3}\left[f(x_0)+4*\sum^{n-1}_{i\in odd}+f(x_i)+2*\sum^{n-2}_{i\in even}+f(x_n)\right]\\ _{\text{(Eq. 6.4)}}$$
```
f = lambda x : x*np.sin(x)
a, b = 0, np.pi/2
n = 6
h = (b-a)/n
A= (f(a)+f(b))
for i in range(1,n,2):
A += 4*f(a+i*h)
for i in range(2,n,2):
A += 2*f(a+i*h)
S = h/3*(A)
S
def simp_13(func,lb,ub,divs):
h = (ub-lb)/divs
A = (func(lb)+func(ub))+ \
np.sum(4*func(lb+h*np.arange(1,divs,2)))+ \
np.sum(2*func(lb+h*np.arange(2,divs,2)))
S = (h/3)*A
return S
h = lambda x: x**2
sum = simp_13(h, 0,10,1e4)
sum
```
## Simpson's 3/8 Rule
Simpson's 3/8 rule or Simpson's second rule is ismilar to the 1/3 rule but instead of having a parabolic or quadratic approximation, it uses a cubic approximation.
$$\int^b_af(x)dx \approx \frac{(b-a)}{8}\left(f(a)+3f\frac{(2a+b)}{3}+3f\frac{(a+2b)}{3}+f(b)\right)\\ _{\text{(Eq. 6.5)}}$$
It can be discretized as:
$$\int^b_af(x)dx \approx \frac{3h}{8}\left[f(x_0)+3*\sum^{n-1}_{i=1,4,7,..}+f(x_i)+3*\sum^{n-2}_{i=2,5,8,..}+f(x_i)+2*\sum^{n-3}_{i=3,6,9,..}+f(x_n)\right]\\ _{\text{(Eq. 6.6)}}$$
```
def simp_38(func,lb,ub,divs):
h = (ub-lb)/divs
A = (func(lb)+func(ub))+ \
np.sum(3*(func(lb+h*np.arange(1,divs,3))))+ \
np.sum(3*(func(lb+h*np.arange(2,divs,3))))+ \
np.sum(2*func(lb+h*np.arange(3,divs,3)))
S = (3*h/8)*A
return S
f = lambda x: x*np.sin(x)
sum = simp_38(f, 0,np.pi/2,1e4)
sum
h = lambda x: x**2
sum = simp_38(h, 0,10,1e4)
sum
```
## Monte Carlo Integration
The Monte Carlo Simulation or integration uses a different approach in approximating the area under a curve or function. It differs from the Trapezoidal and Simpson's Rules since it does not use a polynomial for interpolating the curve. The Monte Carlo integration uses the idea of uniform random sampling in a given space and computes the samples that are under the curve of the equation.
In this implementation, we will use the most vanilla version of the Monte Carlo integration. We will use the definition of the mean of a function given as:
$$\left<f(x)\right> = \frac{1}{(b-a)}\int^b_af(x)dx \\ _{\text{(Eq. 6.7)}}$$
We can then perform algebraic manipulation to solve to isolate the integral of the function:
$$(b-a)\left<f(x)\right> = \int^b_af(x)dx \\ _{\text{(Eq. 6.8)}}$$
Then by the definition of means we can use the discretized mean formula and substitute it with $\left< f(x) \right>$:
$$(b-a)\frac{1}{N}\sum^N_{i=0}f(x_i) \approx \int^b_af(x)dx \\ _{\text{(Eq. 6.9)}}$$
```
a, b = 0, np.pi/2`
n = 1e3
samples = np.random.uniform(a,b,int(n))
f = lambda x: x*np.sin(x)
A = np.sum(f(samples))
S = (b-a)/n)
S
```
# End of Module Activity
$\text{Use another notebook to answer the following problems and create a report for the activities in this notebook.}$
1.) Research on the different numerical integration functions implemented in `scipy`. Explain in your report the function/s with three (3) different functions as examples.
2.) Create numerical integration of two sample cases for each of the following functions: higher-order polynomials (degrees greater than 4), trigonometric functions, and logarithmic functions.
> a.) Implement the numerical integration techniques used in this notebook including the `scipy` function/s.
> b.) Measure and compare the errors of each integration technique to the functions you have created.
3.) Research on the "Law of Big Numbers" and explain the law through:
> a.) Testing Simpson's 3/8 Rule by initializing the bin sizes to be arbitrarily large. Run this for 100 iterations while decreasing the bin sizes by a factor of 100. Graph the errors using `matplotlib`.
> b.) Testing the Monte Carlo Simulation with initializing the sample size from an arbitrarily small size. Run this for 100 iterations while increasing the sample sizes by a factor of 100. Graph the errors using `matplotlib`.
```
```
| github_jupyter |
# Geopandas
```
import pandas as pd
import geopandas as gpd
import matplotlib.pyplot as plt
import cartopy
# Cargar el mapa
mapa = gpd.read_file('data/provincias.geojson')
mapa.head(10)
mapa.plot()
natalidad = pd.read_csv('data/natalidad.csv')
natalidad.head()
mapa_data = pd.merge(mapa, natalidad, left_on='NAME_2', right_on='NAME')
mapa_data.head()
# Control del tamaño de la figura del mapa
fig, ax = plt.subplots(figsize=(10, 10))
# Control del título y los ejes
ax.set_title('Natalidad por Provincias en España, 2018',
pad = 20,
fontdict={'fontsize':20, 'color': '#4873ab'})
ax.set_xlabel('Longitud')
ax.set_ylabel('Latitud')
# Mostrar el mapa finalizado
mapa_data.plot(column='NATALIDAD', cmap='Blues', ax=ax, zorder=5)
# Control del tamaño de la figura del mapa
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
# Control del encuadre (área geográfica) del mapa
ax.axis([-12, 5, 32, 48])
# Control del título y los ejes
ax.set_title('Natalidad por Provincias en España, 2018',
pad = 20,
fontdict={'fontsize':20, 'color': '#4873ab'})
ax.set_xlabel('Longitud')
ax.set_ylabel('Latitud')
# Añadir la leyenda separada del mapa
from mpl_toolkits.axes_grid1 import make_axes_locatable
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.2)
# Generar y cargar el mapa
mapa_data.plot(column='NATALIDAD', cmap='Greens', ax=ax,
legend=True, cax=cax, zorder=5)
# Cargar un mapa base con contornos de países
oceanos = "data/ne_50m_ocean.shp"
map_oceanos = gpd.read_file(oceanos)
map_oceanos.plot(ax=ax, color='#89c0e8', zorder=0)
```
# Cartopy
```
plt.figure(figsize=(20,20))
ax = plt.axes(projection=cartopy.crs.PlateCarree())
ax.add_feature(cartopy.feature.LAND, alpha=0.3 )
ax.add_feature(cartopy.feature.COASTLINE, linestyle=':')
#ax.add_feature(cartopy.feature.OCEAN)
#ax.add_feature(cartopy.feature.BORDERS, linestyle=':')
#ax.add_feature(cartopy.feature.LAKES, alpha=0.5)
#ax.add_feature(cartopy.feature.RIVERS)
plt.show()
data = pd.read_csv('data/TweetSurfData.csv', sep=";")
data.head()
plt.figure(figsize=(20,20))
ax = plt.axes(projection=cartopy.crs.PlateCarree())
ax.add_feature(cartopy.feature.LAND, alpha=0.3 )
ax.add_feature(cartopy.feature.COASTLINE, linestyle=':')
# prepare a color for each point depending on the continent.
data['labels_enc'] = pd.factorize(data['homecontinent'])[0]
# Add a point per position
plt.scatter(data['homelon'], data['homelat'], s=data['n']/6, alpha=0.4, c=data['labels_enc'], cmap="Set1")
# copyright and source data info
plt.text( -170, -58,'Where people talk about #Surf\n\nData collected on twitter by @R_Graph_Gallery during 300 days\nPlot realized with Python and the Basemap library', ha='left', va='bottom', size=9, color='#555555' )
#ax.set_extent([-12, 5, 32, 48])
plt.title('Tweting about surf')
plt.show()
municipios = pd.read_csv('data/municipios.csv')
municipios.head()
municipios.Longitud = municipios.Longitud.str.replace(',','.').astype(float)
municipios.Latitud = municipios.Latitud.str.replace(',','.').astype(float)
capitales = municipios.get(municipios.Capital=='Si')
capitales.plot.scatter('Longitud', 'Latitud')
plt.figure(figsize=(20,20))
ax = plt.axes(projection=cartopy.crs.PlateCarree())
ax.add_feature(cartopy.feature.LAND)
ax.add_feature(cartopy.feature.BORDERS, linestyle=':')
ax.add_feature(cartopy.feature.COASTLINE)
ax.set_extent([-12, 5, 32, 48])
plt.scatter(capitales.Longitud, capitales.Latitud, s=capitales.Habitantes/150, alpha=0.4)
plt.title('Capitales de España')
plt.show()
```
| github_jupyter |
# Intro to Python Data Structures
Lists, Tuples, Sets, Dicts
(c) 2019 Joe James
## Sequences: String, List, Tuple
****
**indexing** - access any item in the sequence using its index.
Indexing starts with 0 for the first element.
```
# string
x = 'frog'
print (x[3])
# list
x = ['pig', 'cow', 'horse']
print (x[1])
```
**slicing** - slice out substrings, sublists, subtuples using indexes.
[start : end+1 : step]
```
x = 'computer'
print(x[1:4])
print(x[1:6:2])
print(x[3:])
print(x[:5])
print(x[-1])
print(x[-3:])
print(x[:-2])
print(x[::-1])
```
**adding / concatenating** - combine 2 sequences of the same type by using +
```
# string
x = 'horse' + 'shoe'
print(x)
# list
y = ['pig', 'cow'] + ['horse']
print(y)
# touple
z = ('kevin','nik', 'jen') + ('craig',)
print(z)
```
**multiplying** - multiply a sequence using *
```
# string
x = 'bug' * 3
print(x)
# list
y = [8, 5] * 3
print(y)
```
**checking membership** - test whether an item is or is not in a sequence.
```
# string
x = 'bug'
print('u' in x)
# list
y = ['pig', 'cow', 'horse']
print('cow' not in y)
# tuple
z = ('kevin','nik', 'jen','craig')
print('jen' in z)
```
**iterating** - iterating through the items in a sequence
```
# item
x = [7, 8, 3]
for item in x:
print(item)
# index & item
y = [7, 8, 3]
for index, item in enumerate(y):
print(index, item)
```
**number of items** - count the number of items in a sequence
```
# string
x = 'bug'
print(len(x))
# list
y = ['pig', 'cow', 'horse']
print(len(y))
```
**minimum** - find the minimum item in a sequence lexicographically.
Alpha or numeric types, but cannot mix types.
```
# string
x = 'bug'
print(min(x))
# list
y = ['pig', 'cow', 'horse']
print(min(y))
```
**maximum** - find the maximum item in a sequence lexicographically.
Alpha or numeric types, but cannot mix types.
```
# string
x = 'bug'
print(max(x))
# list
y = ['pig', 'cow', 'horse']
print(max(y))
```
**sum** - find the sum of items in a sequence.
Entire sequence must be numeric.
```
# string -> error
# x = [5, 7, 'bug']
# print(sum(x)) # generates an error
# list
y = [2, 5, 8, 12]
print(sum(y))
print(sum(y[-2:]))
```
**sorting** - returns a new list of items in sorted order.
Does not change the original list.
```
# string
x = 'bug'
print(sorted(x))
# list
y = ['pig', 'cow', 'horse']
print(sorted(y))
```
**count(item)** - returns count of an item
```
# string
x = 'hippo'
print(x.count('p'))
# list
y = ['pig', 'cow', 'horse', 'cow']
print(y.count('cow'))
```
**index(item)** - returns the index of the first occurence of an item.
```
# string
x = 'hippo'
print(x.index('p'))
# list
y = ['pig', 'cow', 'horse', 'cow']
print(y.index('cow'))
```
**unpacking** - unpack the n items of a sequence into n variables
```
x = ['pig', 'cow', 'horse']
a, b, c = x
print(a, b, c)
```
## Lists
****
- General purpose
- Most widely used data structure
- Grow and shrink size as needed
- Sequence type
- Sortable
**constructors** - creating a new list
```
x = list()
y = ['a', 25, 'dog', 8.43]
tuple1 = (10, 20)
z = list(tuple1)
# list comprehension
a = [m for m in range(8)]
print(a)
b = [i**2 for i in range(10) if i>4]
print(b)
```
**delete** - delete a list or an item in a list
```
x = [5, 3, 8, 6]
del(x[1])
print(x)
del(x) # list x no longer exists
```
**append** - append an item to a list
```
x = [5, 3, 8, 6]
x.append(7)
print(x)
```
**extend** - append a sequence to a list
```
x = [5, 3, 8, 6]
y = [12, 13]
x.extend(y)
print(x)
```
**insert** - insert an item at a given index
```
x = [5, 3, 8, 6]
x.insert(1, 7)
print(x)
x.insert(1, ['a', 'm'])
print(x)
```
**pop** - pops last item off list and returns item
```
x = [5, 3, 8, 6]
x.pop() # pop off the 6
print(x)
print(x.pop())
```
**remove** - remove first instance of an item
```
x = [5, 3, 8, 6, 3]
x.remove(3)
print(x)
```
**reverse** - reverse the order of the list. It is an in-place sort, meaning it changes the original list.
```
x = [5, 3, 8, 6]
x.reverse()
print(x)
```
**sort** - sort the list in place.
Note:
sorted(x) returns a new sorted list without changing the original list x.
x.sort() puts the items of x in sorted order (sorts in place).
```
x = [5, 3, 8, 6]
x.sort()
print(x)
```
## Tuples
****
- Immutable (can’t add/change)
- Useful for fixed data
- Faster than Lists
- Sequence type
**constructors** - creating new tuples.
```
x = ()
x = (1, 2, 3)
x = 1, 2, 3
x = 2, # the comma tells Python it's a tuple
print(x, type(x))
list1 = [2, 4, 6]
x = tuple(list1)
print(x, type(x))
```
**tuples are immutable**, but member objects may be mutable.
```
x = (1, 2, 3)
# del(x[1]) # fails
# x[1] = 8 # fails
print(x)
y = ([1, 2], 3) # a tuple where the first item is a list
del(y[0][1]) # delete the 2
print(y) # the list within the tuple is mutable
```
## Sets
****
- Store non-duplicate items
- Very fast access vs Lists
- Math Set ops (union, intersect)
- Sets are Unordered
**constructors** - creating new sets
```
x = {3, 5, 3, 5}
print(x)
y = set()
print(y)
list1 = [2, 3, 4]
z = set(list1)
print(z)
```
**set operations**
```
x = {3, 8, 5}
print(x)
x.add(7)
print(x)
x.remove(3)
print(x)
# get length of set x
print(len(x))
# check membership in x
print(5 in x)
# pop random item from set x
print(x.pop(), x)
# delete all items from set x
x.clear()
print(x)
```
**Mathematical set operations**
intersection (AND): set1 & set2
union (OR): set1 | set1
symmetric difference (XOR): set1 ^ set2
difference (in set1 but not set2): set1 - set2
subset (set2 contains set1): set1 <= set2
superset (set1 contains set2): set1 >= set2
```
s1 = {1, 2, 3}
s2 = {3, 4, 5}
print(s1 & s2)
print(s1 | s2)
print(s1 ^ s2)
print(s1 - s2)
print(s1 <= s2)
print(s1 >= s2)
```
## Dictionaries (dict)
****
- Key/Value pairs
- Associative array, like Java HashMap
- Dicts are Unordered
```
x = {'pork':25.3, 'beef':33.8, 'chicken':22.7}
print(x)
x = dict([('pork', 25.3),('beef', 33.8),('chicken', 22.7)])
print(x)
x = dict(pork=25.3, beef=33.8, chicken=22.7)
print(x)
```
**dict operations**
```
x['shrimp'] = 38.2 # add or update
print(x)
# delete an item
del(x['shrimp'])
print(x)
# get length of dict x
print(len(x))
# delete all items from dict x
x.clear()
print(x)
# delete dict x
del(x)
```
**accessing keys and values in a dict**
```
y = {'pork':25.3, 'beef':33.8, 'chicken':22.7}
print(y.keys())
print(y.values())
print(y.items()) # key-value pairs
# check membership in y_keys (only looks in keys, not values)
print('beef' in y)
# check membership in y_values
print('clams' in y.values())
```
**iterating a dict - note, items are in random order**
```
for key in y:
print(key, y[key])
for k, v in y.items():
print(k, v)
```
| github_jupyter |
```
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torchvision.transforms as transforms
import torchvision.datasets as datasets
import torchvision.models.vgg as vgg
import torchvision.models.resnet as resnet
import ocd
device = 'cuda' if torch.cuda.is_available() else 'cpu'
torch.manual_seed(1)
if device == 'cuda':
torch.cuda.manual_seed_all(1)
ocd = ocd.OCD10(num_classes= 10).to(device)
#vgg = vgg.vgg19_bn(num_classes = 196).to(device)
#resnet = resnet.wide_resnet101_2(num_classes = 196).to(device)
test = torch.rand(5, 3, 64, 64).to(device)
ocd_out = ocd(test)
#vgg_out = vgg(test)
#resnet_out = resnet(test)
#print(ocd_out.shape, vgg_out.shape, resnet_out.shape)
batch_size = 128
transform_train = transforms.Compose([
transforms.ToTensor(),
transforms.Resize((64, 64)),
transforms.Normalize((0.25, 0.25, 0.25), (0.25, 0.25, 0.25))
])
transform_test = transforms.Compose([
transforms.ToTensor(),
transforms.Resize((64, 64)),
transforms.Normalize((0.25, 0.25, 0.25), (0.25, 0.25, 0.25))
])
train_set = datasets.CIFAR10(root='cifar10/', transform=transform_train, download = True, train =True)
test_set = datasets.CIFAR10(root='cifar10/', transform=transform_test, download = True, train = False)
train_loader = torch.utils.data.DataLoader(dataset = train_set, shuffle = True, drop_last = True, batch_size = batch_size, num_workers = 1)
test_loader = torch.utils.data.DataLoader(dataset = test_set, shuffle = True, drop_last = True, batch_size = batch_size, num_workers = 1)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horese', 'ship', 'truck')
# transform_train = transforms.Compose([
# transforms.ToTensor(),
# transforms.Resize((150, 150)),
# transforms.Normalize((0.25, 0.25, 0.25), (0.25, 0.25, 0.25))
# ])
# transform_test = transforms.Compose([
# transforms.ToTensor(),
# transforms.Resize((150, 150)),
# transforms.Normalize((0.25, 0.25, 0.25), (0.25, 0.25, 0.25))
# ])
# train_set = datasets.ImageFolder(root='stanford-car/car_data/train/', transform = transform_train)
# test_set = datasets.ImageFolder(root='stanford-car/car_data/test/', transform = transform_test)
# train_loader = torch.utils.data.DataLoader(dataset = train_set, shuffle = True, drop_last= True, batch_size = 32, num_workers= 1)
# test_loader = torch.utils.data.DataLoader(dataset = test_set, shuffle = True, drop_last= True, batch_size = 32, num_workers= 1)
# classes = ('AM General Hummer SUV 2000',
# 'Acura RL Sedan 2012',
# 'Acura TL Sedan 2012',
# 'Acura TL Type-S 2008',
# 'Acura TSX Sedan 2012',
# 'Acura Integra Type R 2001',
# 'Acura ZDX Hatchback 2012',
# 'Aston Martin V8 Vantage Convertible 2012',
# 'Aston Martin V8 Vantage Coupe 2012',
# 'Aston Martin Virage Convertible 2012',
# 'Aston Martin Virage Coupe 2012',
# 'Audi RS 4 Convertible 2008',
# 'Audi A5 Coupe 2012',
# 'Audi TTS Coupe 2012',
# 'Audi R8 Coupe 2012',
# 'Audi V8 Sedan 1994',
# 'Audi 100 Sedan 1994',
# 'Audi 100 Wagon 1994',
# 'Audi TT Hatchback 2011',
# 'Audi S6 Sedan 2011',
# 'Audi S5 Convertible 2012',
# 'Audi S5 Coupe 2012',
# 'Audi S4 Sedan 2012',
# 'Audi S4 Sedan 2007',
# 'Audi TT RS Coupe 2012',
# 'BMW ActiveHybrid 5 Sedan 2012',
# 'BMW 1 Series Convertible 2012',
# 'BMW 1 Series Coupe 2012',
# 'BMW 3 Series Sedan 2012',
# 'BMW 3 Series Wagon 2012',
# 'BMW 6 Series Convertible 2007',
# 'BMW X5 SUV 2007',
# 'BMW X6 SUV 2012',
# 'BMW M3 Coupe 2012',
# 'BMW M5 Sedan 2010',
# 'BMW M6 Convertible 2010',
# 'BMW X3 SUV 2012',
# 'BMW Z4 Convertible 2012',
# 'Bentley Continental Supersports Conv. Convertible 2012',
# 'Bentley Arnage Sedan 2009',
# 'Bentley Mulsanne Sedan 2011',
# 'Bentley Continental GT Coupe 2012',
# 'Bentley Continental GT Coupe 2007',
# 'Bentley Continental Flying Spur Sedan 2007',
# 'Bugatti Veyron 16.4 Convertible 2009',
# 'Bugatti Veyron 16.4 Coupe 2009',
# 'Buick Regal GS 2012',
# 'Buick Rainier SUV 2007',
# 'Buick Verano Sedan 2012',
# 'Buick Enclave SUV 2012',
# 'Cadillac CTS-V Sedan 2012',
# 'Cadillac SRX SUV 2012',
# 'Cadillac Escalade EXT Crew Cab 2007',
# 'Chevrolet Silverado 1500 Hybrid Crew Cab 2012',
# 'Chevrolet Corvette Convertible 2012',
# 'Chevrolet Corvette ZR1 2012',
# 'Chevrolet Corvette Ron Fellows Edition Z06 2007',
# 'Chevrolet Traverse SUV 2012',
# 'Chevrolet Camaro Convertible 2012',
# 'Chevrolet HHR SS 2010',
# 'Chevrolet Impala Sedan 2007',
# 'Chevrolet Tahoe Hybrid SUV 2012',
# 'Chevrolet Sonic Sedan 2012',
# 'Chevrolet Express Cargo Van 2007',
# 'Chevrolet Avalanche Crew Cab 2012',
# 'Chevrolet Cobalt SS 2010',
# 'Chevrolet Malibu Hybrid Sedan 2010',
# 'Chevrolet TrailBlazer SS 2009',
# 'Chevrolet Silverado 2500HD Regular Cab 2012',
# 'Chevrolet Silverado 1500 Classic Extended Cab 2007',
# 'Chevrolet Express Van 2007',
# 'Chevrolet Monte Carlo Coupe 2007',
# 'Chevrolet Malibu Sedan 2007',
# 'Chevrolet Silverado 1500 Extended Cab 2012',
# 'Chevrolet Silverado 1500 Regular Cab 2012',
# 'Chrysler Aspen SUV 2009',
# 'Chrysler Sebring Convertible 2010',
# 'Chrysler Town and Country Minivan 2012',
# 'Chrysler 300 SRT-8 2010',
# 'Chrysler Crossfire Convertible 2008',
# 'Chrysler PT Cruiser Convertible 2008',
# 'Daewoo Nubira Wagon 2002',
# 'Dodge Caliber Wagon 2012',
# 'Dodge Caliber Wagon 2007',
# 'Dodge Caravan Minivan 1997',
# 'Dodge Ram Pickup 3500 Crew Cab 2010',
# 'Dodge Ram Pickup 3500 Quad Cab 2009',
# 'Dodge Sprinter Cargo Van 2009',
# 'Dodge Journey SUV 2012',
# 'Dodge Dakota Crew Cab 2010',
# 'Dodge Dakota Club Cab 2007',
# 'Dodge Magnum Wagon 2008',
# 'Dodge Challenger SRT8 2011',
# 'Dodge Durango SUV 2012',
# 'Dodge Durango SUV 2007',
# 'Dodge Charger Sedan 2012',
# 'Dodge Charger SRT-8 2009',
# 'Eagle Talon Hatchback 1998',
# 'FIAT 500 Abarth 2012',
# 'FIAT 500 Convertible 2012',
# 'Ferrari FF Coupe 2012',
# 'Ferrari California Convertible 2012',
# 'Ferrari 458 Italia Convertible 2012',
# 'Ferrari 458 Italia Coupe 2012',
# 'Fisker Karma Sedan 2012',
# 'Ford F-450 Super Duty Crew Cab 2012',
# 'Ford Mustang Convertible 2007',
# 'Ford Freestar Minivan 2007',
# 'Ford Expedition EL SUV 2009',
# 'Ford Edge SUV 2012',
# 'Ford Ranger SuperCab 2011',
# 'Ford GT Coupe 2006',
# 'Ford F-150 Regular Cab 2012',
# 'Ford F-150 Regular Cab 2007',
# 'Ford Focus Sedan 2007',
# 'Ford E-Series Wagon Van 2012',
# 'Ford Fiesta Sedan 2012',
# 'GMC Terrain SUV 2012',
# 'GMC Savana Van 2012',
# 'GMC Yukon Hybrid SUV 2012',
# 'GMC Acadia SUV 2012',
# 'GMC Canyon Extended Cab 2012',
# 'Geo Metro Convertible 1993',
# 'HUMMER H3T Crew Cab 2010',
# 'HUMMER H2 SUT Crew Cab 2009',
# 'Honda Odyssey Minivan 2012',
# 'Honda Odyssey Minivan 2007',
# 'Honda Accord Coupe 2012',
# 'Honda Accord Sedan 2012',
# 'Hyundai Veloster Hatchback 2012',
# 'Hyundai Santa Fe SUV 2012',
# 'Hyundai Tucson SUV 2012',
# 'Hyundai Veracruz SUV 2012',
# 'Hyundai Sonata Hybrid Sedan 2012',
# 'Hyundai Elantra Sedan 2007',
# 'Hyundai Accent Sedan 2012',
# 'Hyundai Genesis Sedan 2012',
# 'Hyundai Sonata Sedan 2012',
# 'Hyundai Elantra Touring Hatchback 2012',
# 'Hyundai Azera Sedan 2012',
# 'Infiniti G Coupe IPL 2012',
# 'Infiniti QX56 SUV 2011',
# 'Isuzu Ascender SUV 2008',
# 'Jaguar XK XKR 2012',
# 'Jeep Patriot SUV 2012',
# 'Jeep Wrangler SUV 2012',
# 'Jeep Liberty SUV 2012',
# 'Jeep Grand Cherokee SUV 2012',
# 'Jeep Compass SUV 2012',
# 'Lamborghini Reventon Coupe 2008',
# 'Lamborghini Aventador Coupe 2012',
# 'Lamborghini Gallardo LP 570-4 Superleggera 2012',
# 'Lamborghini Diablo Coupe 2001',
# 'Land Rover Range Rover SUV 2012',
# 'Land Rover LR2 SUV 2012',
# 'Lincoln Town Car Sedan 2011',
# 'MINI Cooper Roadster Convertible 2012',
# 'Maybach Landaulet Convertible 2012',
# 'Mazda Tribute SUV 2011',
# 'McLaren MP4-12C Coupe 2012',
# 'Mercedes-Benz 300-Class Convertible 1993',
# 'Mercedes-Benz C-Class Sedan 2012',
# 'Mercedes-Benz SL-Class Coupe 2009',
# 'Mercedes-Benz E-Class Sedan 2012',
# 'Mercedes-Benz S-Class Sedan 2012',
# 'Mercedes-Benz Sprinter Van 2012',
# 'Mitsubishi Lancer Sedan 2012',
# 'Nissan Leaf Hatchback 2012',
# 'Nissan NV Passenger Van 2012',
# 'Nissan Juke Hatchback 2012',
# 'Nissan 240SX Coupe 1998',
# 'Plymouth Neon Coupe 1999',
# 'Porsche Panamera Sedan 2012',
# 'Ram C/V Cargo Van Minivan 2012',
# 'Rolls-Royce Phantom Drophead Coupe Convertible 2012',
# 'Rolls-Royce Ghost Sedan 2012',
# 'Rolls-Royce Phantom Sedan 2012',
# 'Scion xD Hatchback 2012',
# 'Spyker C8 Convertible 2009',
# 'Spyker C8 Coupe 2009',
# 'Suzuki Aerio Sedan 2007',
# 'Suzuki Kizashi Sedan 2012',
# 'Suzuki SX4 Hatchback 2012',
# 'Suzuki SX4 Sedan 2012',
# 'Tesla Model S Sedan 2012',
# 'Toyota Sequoia SUV 2012',
# 'Toyota Camry Sedan 2012',
# 'Toyota Corolla Sedan 2012',
# 'Toyota 4Runner SUV 2012',
# 'Volkswagen Golf Hatchback 2012',
# 'Volkswagen Golf Hatchback 1991',
# 'Volkswagen Beetle Hatchback 2012',
# 'Volvo C30 Hatchback 2012',
# 'Volvo 240 Sedan 1993',
# 'Volvo XC90 SUV 2007',
# 'smart fortwo Convertible 2012',
# ),
training_epochs = 200
learning_rate_ocd = 0.1
learning_rate_vgg = 0.01
learning_rate_resnet = 0.5
optimizer_ocd = optim.Adam(ocd.parameters(), lr = learning_rate_ocd)
#optimizer_vgg = optim.SGD(vgg.parameters(), momentum =0.9, lr = learning_rate_vgg)
#optimizer_resnet = optim.SGD(resnet.parameters(), momentum =0.9, lr = learning_rate_resnet)
lr_sche_ocd = optim.lr_scheduler.StepLR(optimizer_ocd, step_size =100, gamma = 0.9)
#lr_sche_vgg = optim.lr_scheduler.StepLR(optimizer_vgg, step_size =100, gamma = 0.9)
#lr_sche_resnet = optim.lr_scheduler.StepLR(optimizer_resnet, step_size =100, gamma = 0.9)
import visdom
vis = visdom.Visdom()
vis.close(env = "main")
ocd_loss_plt = vis.line(Y=torch.Tensor(1).zero_(), opts = dict(title = "ocd_loss_tracker", legend = ['loss'], showlegend = True))
ocd_acc_plt = vis.line(Y = torch.Tensor(1).zero_(), opts = dict(title = 'ocd_Accuracy', legend = ['Acc'], showlegend = True))
#vgg_loss_plt = vis.line(Y=torch.Tensor(1).zero_(), opts = dict(title = "vgg_loss_tracker", legend = ['loss'], showlegend = True))
#vgg_acc_plt = vis.line(Y = torch.Tensor(1).zero_(), opts = dict(title = 'vgg_Accuracy', legend = ['Acc'], showlegend = True))
#resnet_loss_plt = vis.line(Y=torch.Tensor(1).zero_(), opts = dict(title = "resnet_loss_tracker", legend = ['loss'], showlegend = True))
#resnet_acc_plt = vis.line(Y = torch.Tensor(1).zero_(), opts = dict(title = 'resnet_Accuracy', legend = ['Acc'], showlegend = True))
def value_tracker(value_plot, value, index):
'''num, loss_value are Tensor'''
vis.line(X = index, Y = value, win = value_plot, update = 'append')
def acc_check(net, name, test_loader, epoch, save = 1):
correct = 0
total = 0
with torch.no_grad():
for X, Y in test_loader:
X = X.to(device)
Y = Y.to(device)
#prediction
pred = net(X)
_, predicted = torch.max(pred, 1)
total += Y.size(0)
correct += (predicted == Y).sum().item()
acc = (100 * correct / total)
print('correct: ', correct)
print('total: ', total)
print('Accuracy of the network on the 10000 test images: {}'.format(acc))
if epoch % 5 == 4:
if save:
torch.save(net.state_dict(), "./models/cifar10/light_v/{}_model_epoch_{}_acc_{:5f}.pth".format(name, epoch, acc))
return acc
total_batch = len(train_loader)
for epoch in range(training_epochs):
ocd.train()
#vgg.train()
#resnet.train()
running_loss_ocd = 0.0
#running_loss_vgg = 0.0
#running_loss_resnet = 0.0
for i, data in enumerate(train_loader):
X, Y = data
X = X.to(device)
Y = Y.to(device)
#prediction
pred_ocd = ocd(X)
#pred_vgg = vgg(X)
#pred_resnet = resnet(X)
#cost
cost_ocd = F.cross_entropy(pred_ocd, Y).to(device)
#cost_vgg = F.cross_entropy(pred_vgg, Y).to(device)
#cost_resnet = F.cross_entropy(pred_resnet, Y).to(device)
#Reduce the cost
optimizer_ocd.zero_grad()
cost_ocd.backward()
optimizer_ocd.step()
#optimizer_vgg.zero_grad()
#cost_vgg.backward()
#optimizer_vgg.step()
#optimizer_resnet.zero_grad()
#cost_resnet.backward()
#optimizer_resnet.step()
#print statistics
running_loss_ocd += cost_ocd.item()
#running_loss_vgg += cost_vgg.item()
#running_loss_resnet += cost_resnet.item()
if i % 5 == 4:
print('Epoch: {} / {}, MiniBatch: {} / {}, Cost: {}'.format(epoch + 1, 100, i, len(train_loader), cost_ocd.item()))
value_tracker(ocd_loss_plt, torch.Tensor([running_loss_ocd / 5]), torch.Tensor([i + epoch * len(train_loader)]))
#print('Epoch: {} / {}, MiniBatch: {} / {}, Cost: {}'.format(epoch + 1, 100, i, len(train_loader), cost_vgg.item()))
#value_tracker(vgg_loss_plt, torch.Tensor([running_loss_vgg / 5]), torch.Tensor([i + epoch * len(train_loader)]))
#print('Epoch: {} / {}, MiniBatch: {} / {}, Cost: {}'.format(epoch + 1, 100, i, len(train_loader), cost_resnet.item()))
#value_tracker(resnet_loss_plt, torch.Tensor([running_loss_resnet / 5]), torch.Tensor([i + epoch * len(train_loader)]))
running_loss_ocd = 0
#running_loss_resnet = 0
#running_loss_vgg = 0
#Check accuracy
acc_ocd = acc_check(ocd, 'ocd', test_loader, epoch, save = 1)
value_tracker(ocd_acc_plt, torch.Tensor([acc_ocd]), torch.Tensor([epoch]))
#acc_vgg = acc_check(vgg, 'vgg', test_loader, epoch, save = 1)
#value_tracker(vgg_acc_plt, torch.Tensor([acc_vgg]), torch.Tensor([epoch]))
#acc_resnet = acc_check(resnet, 'resnet', test_loader, epoch, save = 1)
#value_tracker(resnet_acc_plt, torch.Tensor([acc_resnet]), torch.Tensor([epoch]))
print('Finshed Learning')
```
| github_jupyter |
```
# Import pyNBS modules
from pyNBS import data_import_tools as dit
from pyNBS import network_propagation as prop
from pyNBS import pyNBS_core as core
from pyNBS import pyNBS_single
from pyNBS import consensus_clustering as cc
from pyNBS import pyNBS_plotting as plot
# Import other needed packages
import os
import time
import pandas as pd
import numpy as np
from IPython.display import Image
```
# Load Data
First, we must load the somatic mutation and network data for running pyNBS. We will also set an output directory location to save our results.
### Load binary somatic mutation data
The binary somatic mutation data file can be represented in two file formats:
The default format for the binary somatic mutation data file is the ```list``` format. This file format is a 2-column csv or tsv list where the 1st column is a sample/patient and the 2nd column is a gene mutated in the sample/patient. There are no headers in this file format. Loading data with the list format is typically faster than loading data from the matrix format.The following text is the list representation of the matrix above.
```
TCGA-04-1638 A2M
TCGA-23-1029 A1CF
TCGA-23-2647 A2BP1
TCGA-24-1847 A2M
TCGA-42-2589 A1CF
```
The ```matrix``` binary somatic mutation data format is the format that data for this example is currently represented. This file format is a binary csv or tsv matrix with rows represent samples/patients and columns represent genes. The following table is a small excerpt of a matrix somatic mutation data file:
||A1CF|A2BP1|A2M|
|-|-|-|-|
|TCGA-04-1638|0|0|1|
|TCGA-23-1029|1|0|0|
|TCGA-23-2647|0|1|0|
|TCGA-24-1847|0|0|1|
|TCGA-42-2589|1|0|0|
__Note:__ The default file type is defined as ```'list'```, but if the user would like to specify the 'matrix' type, the user needs to simply pass the string ```'matrix'``` to the ```filetype``` optional parameter (as below). The delimiter for the file is passed similarly to the optional parameter ```delimiter```
For more examples and definitions in the somatic mutation data file format, please see our Github Wiki page:
https://github.com/huangger/pyNBS/wiki/Somatic-Mutation-Data-File-Format
```
sm_data_filepath = './Example_Data/Mutation_Files/HNSC_sm_data.txt'
sm_mat = dit.load_binary_mutation_data(sm_data_filepath, filetype='list', delimiter='\t')
```
### Load molecular network
The network file is a 2-column text file representing an unweighted network. Each row represents a single edge in the molecular network.
Notes about the network file:
- The default column delimiter is a tab character '\t' but a different delimiter can be defined by the user here or in the parameter file with the "net_filedelim" parameter.
- The network must not contain duplicate edges (e.g. TP53\tMDM2 is equivalent to MDM2\tTP53)
- The network must not contain self-edges (e.g. TP53\tTP53)
- Only the first two columns of a network file are read as edges for the network, all other columns will be ignored.
- The load_network function also includes options to read in edge- or label-shuffled versions of the network, but by default, these options are turned off.
An excerpt of the first five rows of the PID network file is given below:
```
A1BG A2M
A1BG AKT1
A1BG GRB2
A1BG PIK3CA
A1BG PIK3R1
```
For more examples and definitions in the network file format, please see our Github Wiki page:
https://github.com/huangger/pyNBS/wiki/Molecular-Network-File-Format
```
# The only required parameter for this function is the network file path
network_filepath = './Example_Data/Network_Files/CancerSubnetwork.txt'
network = dit.load_network_file(network_filepath)
```
### Setting result output options
The following code is completely optional for the user. Allows users to pre-define a directory to save intermediate and final results to and establishes a file name prefix for those files in the output directory folder. Also creates the output directory if it does not already exist. The result of this cell will be a dictionary that can be passed optionally to functions to save results.
**Note:** The key assumption here is that if the user passes **save_args to the function that contains a valid file path to a directory in ```outdir```, the result of that particular call of the function will be saved to the given ```outdir```
```
# Optional: Setting the output directory for files to be saved in
outdir = './Results/via_notebook/CancerSubnetwork_HNSC/'
# Optional: Creating above output directory if it doesn't already exist
if not os.path.exists(outdir):
os.makedirs(outdir)
# Optional: Setting a filename prefix for all files saved to outdir
job_name = 'CancerSubnetwork_HNSC'
# Constructs dictionary to be passed as "save_args" to functions if output to be saved
save_args = {'outdir': outdir, 'job_name': job_name}
```
# Construct regularization graph for use in network-regularized NMF
In this step, we will construct the graph used in the network-regularized non-negative matrix factorization (netNMF) step of pyNBS. This network is a K-nearest neighbor (KNN) network constructed from the network influence matrix (Vandin et al 2011*) of the molecular network being used to stratify tumor samples. The graph laplacian of this KNN network (knnGlap) is used as the regularizer in the following netNMF steps. This step uses the ```network_inf_KNN_glap``` function in the pyNBS_core module.
For additional notes on the graph laplacian construction method, please visit our GitHub wiki for this function:
https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_core.network_inf_KNN_glap
---
**Note: ** This step is technically optional. No regularization network laplacian has to be constructed if the user would like to run the NMF step without a network regularizer. The user simply has to pass ```None``` into the optional parameter ```regNet_glap``` or remove the optional parameter in the ```pyNBS_single()``` function call below. This will cause pyNBS to run a non-network regularized NMF procedure. However, given the implementation of the multiplicative update steps, the results may not be exactly the same as some other NMF implementations (e.g. from scikit-learn).
```
# Constructing knnGlap
knnGlap = core.network_inf_KNN_glap(network)
##########################################################################################################
# The resulting matrix can be very large, so we choose not to save the intermediate result here
# To run this function and save the KNN graph laplaican to the output directory 'outdir' given above:
# Uncomment and run the following line instead:
# knnGlap = core.network_inf_KNN_glap(network, **save_args)
##########################################################################################################
```
# Construct network propagation kernel matrix
Due to the multiple subsampling and propagation steps used in pyNBS, we have found that the algorithm can be significantly sped up for large numbers of subsampling and propagation iterations if a gene-by-gene matrix describing the influence of each gene on every other gene in the network by the random-walk propagation operation is pre-computed. We refer to this matrix as the "network propagation kernel". Here we compute this propagation kernel by propagating the all genes in the molecular network independently of one another. The propagation profile of each tumor is then simply the column sum vector of the resulting network propagation kernel selected for only the rows of genes marked as mutated in each tumor, rather than having to perform the full network propagation step again after each subsampling of the data.
For additional notes on the propagation methods used, please visit our GitHub wiki for this function:
https://github.com/huangger/pyNBS/wiki/pyNBS.network_propagation.network_propagation
### Calibrating the network propagation coefficient
The current network propagation coefficient ($\alpha$) is currently set to 0.7 and must range between 0 and 1. This parameter can be tuned and changing it may have a result on the final propagation results. Previous results from [Hofree et al 2013](https://www.nature.com/articles/nmeth.2651) suggest that values between 0.5 and 0.8 produce relatively robust results, but we suspect that the optimal value may be dependent on certain network properties such as edge density.
```
# Set or change network propagation coefficient if desired
alpha = 0.7
# Construct identity matrix of network
network_nodes = network.nodes()
network_I = pd.DataFrame(np.identity(len(network_nodes)), index=network_nodes, columns=network_nodes)
# Construct network propagation kernel
kernel = prop.network_propagation(network, network_I, alpha=alpha, symmetric_norm=False)
##########################################################################################################
# The resulting matrix can be very large, so we choose not to save the intermediate result here
# To run this function and save the propagation kernel to the output directory 'outdir' given above,
# Uncomment and run the following two lines instead of the above line:
# save_args['iteration_label']='kernel'
# kernel = prop.network_propagation(network, network_I, alpha=alpha, symmetric_norm=True, **save_args)
##########################################################################################################
```
# Subsampling, propagation, and netNMF
After the pre-computation of the regularization graph laplacian and the network propagation kernel, we perform the following core steps of the NBS algorithm multiple times (default=100x) to produce multiple patient clusterings that will be used in the later consensus clustering step. Each patient clustering is performed with the following steps:
1. **Subsample binary somatic mutation data.** (See the documentation for the [```subsample_sm_mat```](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_core.subsample_sm_mat) function for more details.)
2. **Propagate binary somatic mutation data over network.** (See the documentation for the [```network_propagation```](https://github.com/huangger/pyNBS/wiki/pyNBS.network_propagation.network_propagation) or [```network_kernel_propagation```](https://github.com/huangger/pyNBS/wiki/pyNBS.network_propagation.network_propagation) function for more details.)
3. **Quantile normalize the network-smoothed mutation data.** (See the documentation for the [```qnorm```](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_core.qnorm) function for more details.)
4. **Use netNMF to decompose network data into k clusters.** (See the documentation for the [```mixed_netNMF```](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_core.mixed_netNMF) function for more details.)
These functions for each step here are wrapped by the [```NBS_single```](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_single.NBS_single) function, which calls each step above in sequence to perform a single iteration of the pyNBS algorithm.
### Number of pyNBS clusters
The default number of clusters constructed by pyNBS is k=3. We change that definition explicitly below or in the parameters for [```NBS_single```](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_single.NBS_single), and in this example we choose 4 clusters. Other parameters such as the subsampling parameters and the propagation coefficient (when no kernel is pre-computed) can also be changed using \*\*kwargs. \*\*kwargs can also will hold the values of \*\*save_args as seen in previous functions if the user would like to save the resulting dimension reduced patient profiles. All documentation of \*\*kwargs definitions are given in the Github wiki page for [```NBS_single```](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_single.NBS_single)
```
clusters = 4
```
### Number of pyNBS iterations
The consensus clustering step of the pyNBS algorithm will improve if the data is subsampled, and re-clustered multiple times. The default number of times we perform the aforementioned operation (```niter```) is 100 times. The number can be reduced for faster run-time, but may produce less robust results. Increasing ```niter``` will increase overall runtime, but should produce more robust cluster assignments during consensus clustering.
```
# Set the number of times to perform pyNBS core steps
niter = 100
```
### Minimum number of mutations
In order to better utilize more patients in the cohort, lowering the minimum number of mutations for patients required for propagation (especially when using smaller networks) may be more appropriate. The default number of minimum mutations for a patient to have after subsampling is 10, but for smaller networks, using patients with less mutations, especially if they fall within the network can still be appropriate. We lower the threshold of minimum mutations for patients below.
```
params = {'min_muts':'5'}
# Optional: Saving the intermediate propagation step (from subsampled data) to file
# save_args['save_prop'] = True
# Run pyNBS 'niter' number of times
Hlist = []
for i in range(niter):
netNMF_time = time.time()
# Run pyNBS core steps and save resulting H matrix to Hlist
Hlist.append(pyNBS_single.NBS_single(sm_mat, knnGlap, propNet=network, propNet_kernel=kernel, k=clusters, **params))
##########################################################################################################
# Optional: If the user is saving intermediate outputs (propagation results or H matrices),
# a different 'iteration_label' should be used for each call of pyNBS_single().
# Otherwise, the user will overwrite each H matrix at each call of pyNBS_single()
# Uncomment and run the two lines below to save intermediate steps instead of the previous line
# save_args['iteration_label']=str(i+1)
# Hlist.append(pyNBS_single.NBS_single(sm_mat, propNet=network, propNet_kernel=kernel, regNet_glap=knnGlap,
# k=clusters, **save_args))
##########################################################################################################
# Report run time of each pyNBS iteration
t = time.time()-netNMF_time
print 'NBS iteration:', i+1, 'complete:', t, 'seconds'
```
# Consensus Clustering
In order to produce robust patient clusters, the sub-sampling and re-clustering steps as done above are needed. After the patient data is subsampled multiple times (default ```niter```=100), we perform the [```consensus_hclust_hard```](https://github.com/huangger/pyNBS/wiki/pyNBS.consensus_clustering.consensus_hclust_hard) function in the conensus_clustering module. It accepts a list of pandas dataframes as generated in the previous step. If the H matrices were generated separately and saved to a directory, the user will need to manually import those H matrices into a python list first before passing the list to the function below.
For more information on how the consensus clustering is performed, please see our wiki page on this function:
https://github.com/huangger/pyNBS/wiki/pyNBS.consensus_clustering.consensus_hclust_hard
```
NBS_cc_table, NBS_cc_linkage, NBS_cluster_assign = cc.consensus_hclust_hard(Hlist, k=clusters, **save_args)
```
# Co-Clustering Map
To visualize the clusters formed by the pyNBS algorithm, we can plot a similarity map using the objects created in the previous step. This step uses the [`cluster_color_assign`](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_plotting.cluster_color_assign) and [`plot_cc_map()`](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_plotting.plot_cc_map) functions in the [`pyNBS_plotting`](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_plotting) module.
```
# Assign colors to clusters from pyNBS
pyNBS_HNSC_clust_cmap = plot.cluster_color_assign(NBS_cluster_assign, name='pyNBS HNSC Cluster Assignments')
# Plot and save co-cluster map figure
plot.plot_cc_map(NBS_cc_table, NBS_cc_linkage, col_color_map=pyNBS_HNSC_clust_cmap, **save_args)
Image(filename = save_args['outdir']+save_args['job_name']+'_cc_map.png', width=600, height=600)
```
# Survival analysis
To determine if the patient clusters are prognostically relevant, we perform a standard survival analysis using a multi-class logrank test to evaluate the significance of survival separation between patient clusters. This data is plotted using a Kaplan-Meier plot using the [`cluster_KMplot()`](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_plotting.cluster_KMplot) in the [`pyNBS_plotting`](https://github.com/huangger/pyNBS/wiki/pyNBS.pyNBS_plotting) module.
In order to plot the survival differences between clusters, we will need to load survival data for each patient. This data was extracted from TCGA clinical data. The survival data is given in a 5-column delimited table with the specific headings described below (the columns must be in the same order as shown below). The following is an example of a few lines of the a survival table:
||vital_status|days_to_death|days_to_last_followup|overall_survival|
|-|-|-|-|-|
|TCGA-2E-A9G8|0|0|1065|1065|
|TCGA-A5-A0GI|0|0|1750|1750|
|TCGA-A5-A0GM|0|0|1448|1448|
|TCGA-A5-A1OK|0|0|244|244|
|TCGA-A5-AB3J|0|0|251|251|
Additional details on the survival data file format is also describe on our Github wiki at:
https://github.com/huangger/pyNBS/wiki/Patient-Survival-Data-File-Format
Note: The default setting for pyNBS is that no survival curves are drawn because the survival data is not a required parameter. The path to valid survival data must be explicitly defined.
```
# Load survival Data
surv_data = './Example_Data/Clinical_Files/HNSC.clin.merged.surv.txt'
# Plot KM Plot for patient clusters
plot.cluster_KMplot(NBS_cluster_assign, surv_data, delimiter=',', **save_args)
Image(filename = save_args['outdir']+save_args['job_name']+'_KM_plot.png', width=600, height=600)
```
| github_jupyter |
# 6 - Transformers for Sentiment Analysis
In this notebook we will be using the transformer model, first introduced in [this](https://arxiv.org/abs/1706.03762) paper. Specifically, we will be using the BERT (Bidirectional Encoder Representations from Transformers) model from [this](https://arxiv.org/abs/1810.04805) paper.
Transformer models are considerably larger than anything else covered in these tutorials. As such we are going to use the [transformers library](https://github.com/huggingface/transformers) to get pre-trained transformers and use them as our embedding layers. We will freeze (not train) the transformer and only train the remainder of the model which learns from the representations produced by the transformer. In this case we will be using a multi-layer bi-directional GRU, however any model can learn from these representations.
## Preparing Data
First, as always, let's set the random seeds for deterministic results.
```
import torch
import random
import numpy as np
SEED = 1234
random.seed(SEED)
np.random.seed(SEED)
torch.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
```
The transformer has already been trained with a specific vocabulary, which means we need to train with the exact same vocabulary and also tokenize our data in the same way that the transformer did when it was initially trained.
Luckily, the transformers library has tokenizers for each of the transformer models provided. In this case we are using the BERT model which ignores casing (i.e. will lower case every word). We get this by loading the pre-trained `bert-base-uncased` tokenizer.
```
from transformers import BertTokenizer
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
```
The `tokenizer` has a `vocab` attribute which contains the actual vocabulary we will be using. We can check how many tokens are in it by checking its length.
```
len(tokenizer.vocab)
```
Using the tokenizer is as simple as calling `tokenizer.tokenize` on a string. This will tokenize and lower case the data in a way that is consistent with the pre-trained transformer model.
```
tokens = tokenizer.tokenize('Hello WORLD how ARE yoU?')
print(tokens)
```
We can numericalize tokens using our vocabulary using `tokenizer.convert_tokens_to_ids`.
```
indexes = tokenizer.convert_tokens_to_ids(tokens)
print(indexes)
```
The transformer was also trained with special tokens to mark the beginning and end of the sentence, detailed [here](https://huggingface.co/transformers/model_doc/bert.html#transformers.BertModel). As well as a standard padding and unknown token. We can also get these from the tokenizer.
**Note**: the tokenizer does have a beginning of sequence and end of sequence attributes (`bos_token` and `eos_token`) but these are not set and should not be used for this transformer.
```
init_token = tokenizer.cls_token
eos_token = tokenizer.sep_token
pad_token = tokenizer.pad_token
unk_token = tokenizer.unk_token
print(init_token, eos_token, pad_token, unk_token)
```
We can get the indexes of the special tokens by converting them using the vocabulary...
```
init_token_idx = tokenizer.convert_tokens_to_ids(init_token)
eos_token_idx = tokenizer.convert_tokens_to_ids(eos_token)
pad_token_idx = tokenizer.convert_tokens_to_ids(pad_token)
unk_token_idx = tokenizer.convert_tokens_to_ids(unk_token)
print(init_token_idx, eos_token_idx, pad_token_idx, unk_token_idx)
```
...or by explicitly getting them from the tokenizer.
```
init_token_idx = tokenizer.cls_token_id
eos_token_idx = tokenizer.sep_token_id
pad_token_idx = tokenizer.pad_token_id
unk_token_idx = tokenizer.unk_token_id
print(init_token_idx, eos_token_idx, pad_token_idx, unk_token_idx)
```
Another thing we need to handle is that the model was trained on sequences with a defined maximum length - it does not know how to handle sequences longer than it has been trained on. We can get the maximum length of these input sizes by checking the `max_model_input_sizes` for the version of the transformer we want to use. In this case, it is 512 tokens.
```
max_input_length = tokenizer.max_model_input_sizes['bert-base-uncased']
print(max_input_length)
```
Previously we have used the `spaCy` tokenizer to tokenize our examples. However we now need to define a function that we will pass to our `TEXT` field that will handle all the tokenization for us. It will also cut down the number of tokens to a maximum length. Note that our maximum length is 2 less than the actual maximum length. This is because we need to append two tokens to each sequence, one to the start and one to the end.
```
def tokenize_and_cut(sentence):
tokens = tokenizer.tokenize(sentence)
tokens = tokens[:max_input_length-2]
return tokens
```
Now we define our fields. The transformer expects the batch dimension to be first, so we set `batch_first = True`. As we already have the vocabulary for our text, provided by the transformer we set `use_vocab = False` to tell torchtext that we'll be handling the vocabulary side of things. We pass our `tokenize_and_cut` function as the tokenizer. The `preprocessing` argument is a function that takes in the example after it has been tokenized, this is where we will convert the tokens to their indexes. Finally, we define the special tokens - making note that we are defining them to be their index value and not their string value, i.e. `100` instead of `[UNK]` This is because the sequences will already be converted into indexes.
We define the label field as before.
```
from torchtext import data
TEXT = data.Field(batch_first = True,
use_vocab = False,
tokenize = tokenize_and_cut,
preprocessing = tokenizer.convert_tokens_to_ids,
init_token = init_token_idx,
eos_token = eos_token_idx,
pad_token = pad_token_idx,
unk_token = unk_token_idx)
LABEL = data.LabelField(dtype = torch.float)
```
We load the data and create the validation splits as before.
```
from torchtext import datasets
train_data, test_data = datasets.IMDB.splits(TEXT, LABEL)
train_data, valid_data = train_data.split(random_state = random.seed(SEED))
print(f"Number of training examples: {len(train_data)}")
print(f"Number of validation examples: {len(valid_data)}")
print(f"Number of testing examples: {len(test_data)}")
```
We can check an example and ensure that the text has already been numericalized.
```
print(vars(train_data.examples[6]))
```
We can use the `convert_ids_to_tokens` to transform these indexes back into readable tokens.
```
tokens = tokenizer.convert_ids_to_tokens(vars(train_data.examples[6])['text'])
print(tokens)
```
Although we've handled the vocabulary for the text, we still need to build the vocabulary for the labels.
```
LABEL.build_vocab(train_data)
print(LABEL.vocab.stoi)
```
As before, we create the iterators. Ideally we want to use the largest batch size that we can as I've found this gives the best results for transformers.
```
BATCH_SIZE = 128
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size = BATCH_SIZE,
device = device)
```
## Build the Model
Next, we'll load the pre-trained model, making sure to load the same model as we did for the tokenizer.
```
from transformers import BertTokenizer, BertModel
bert = BertModel.from_pretrained('bert-base-uncased')
```
Next, we'll define our actual model.
Instead of using an embedding layer to get embeddings for our text, we'll be using the pre-trained transformer model. These embeddings will then be fed into a GRU to produce a prediction for the sentiment of the input sentence. We get the embedding dimension size (called the `hidden_size`) from the transformer via its config attribute. The rest of the initialization is standard.
Within the forward pass, we wrap the transformer in a `no_grad` to ensure no gradients are calculated over this part of the model. The transformer actually returns the embeddings for the whole sequence as well as a *pooled* output. The [documentation](https://huggingface.co/transformers/model_doc/bert.html#transformers.BertModel) states that the pooled output is "usually not a good summary of the semantic content of the input, you’re often better with averaging or pooling the sequence of hidden-states for the whole input sequence", hence we will not be using it. The rest of the forward pass is the standard implementation of a recurrent model, where we take the hidden state over the final time-step, and pass it through a linear layer to get our predictions.
```
import torch.nn as nn
class BERTGRUSentiment(nn.Module):
def __init__(self,
bert,
hidden_dim,
output_dim,
n_layers,
bidirectional,
dropout):
super().__init__()
self.bert = bert
embedding_dim = bert.config.to_dict()['hidden_size']
self.rnn = nn.GRU(embedding_dim,
hidden_dim,
num_layers = n_layers,
bidirectional = bidirectional,
batch_first = True,
dropout = 0 if n_layers < 2 else dropout)
self.out = nn.Linear(hidden_dim * 2 if bidirectional else hidden_dim, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, text):
#text = [batch size, sent len]
with torch.no_grad():
embedded = self.bert(text)[0]
#embedded = [batch size, sent len, emb dim]
_, hidden = self.rnn(embedded)
#hidden = [n layers * n directions, batch size, emb dim]
if self.rnn.bidirectional:
hidden = self.dropout(torch.cat((hidden[-2,:,:], hidden[-1,:,:]), dim = 1))
else:
hidden = self.dropout(hidden[-1,:,:])
#hidden = [batch size, hid dim]
output = self.out(hidden)
#output = [batch size, out dim]
return output
```
Next, we create an instance of our model using standard hyperparameters.
```
HIDDEN_DIM = 256
OUTPUT_DIM = 1
N_LAYERS = 2
BIDIRECTIONAL = True
DROPOUT = 0.25
model = BERTGRUSentiment(bert,
HIDDEN_DIM,
OUTPUT_DIM,
N_LAYERS,
BIDIRECTIONAL,
DROPOUT)
```
We can check how many parameters the model has. Our standard models have under 5M, but this one has 112M! Luckily, 110M of these parameters are from the transformer and we will not be training those.
```
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
```
In order to freeze paramers (not train them) we need to set their `requires_grad` attribute to `False`. To do this, we simply loop through all of the `named_parameters` in our model and if they're a part of the `bert` transformer model, we set `requires_grad = False`.
```
for name, param in model.named_parameters():
if name.startswith('bert'):
param.requires_grad = False
```
We can now see that our model has under 3M trainable parameters, making it almost comparable to the `FastText` model. However, the text still has to propagate through the transformer which causes training to take considerably longer.
```
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
```
We can double check the names of the trainable parameters, ensuring they make sense. As we can see, they are all the parameters of the GRU (`rnn`) and the linear layer (`out`).
```
for name, param in model.named_parameters():
if param.requires_grad:
print(name)
```
## Train the Model
As is standard, we define our optimizer and criterion (loss function).
```
import torch.optim as optim
optimizer = optim.Adam(model.parameters())
criterion = nn.BCEWithLogitsLoss()
```
Place the model and criterion onto the GPU (if available)
```
model = model.to(device)
criterion = criterion.to(device)
```
Next, we'll define functions for: calculating accuracy, performing a training epoch, performing an evaluation epoch and calculating how long a training/evaluation epoch takes.
```
def binary_accuracy(preds, y):
"""
Returns accuracy per batch, i.e. if you get 8/10 right, this returns 0.8, NOT 8
"""
#round predictions to the closest integer
rounded_preds = torch.round(torch.sigmoid(preds))
correct = (rounded_preds == y).float() #convert into float for division
acc = correct.sum() / len(correct)
return acc
def train(model, iterator, optimizer, criterion):
epoch_loss = 0
epoch_acc = 0
model.train()
for batch in iterator:
optimizer.zero_grad()
predictions = model(batch.text).squeeze(1)
loss = criterion(predictions, batch.label)
acc = binary_accuracy(predictions, batch.label)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
def evaluate(model, iterator, criterion):
epoch_loss = 0
epoch_acc = 0
model.eval()
with torch.no_grad():
for batch in iterator:
predictions = model(batch.text).squeeze(1)
loss = criterion(predictions, batch.label)
acc = binary_accuracy(predictions, batch.label)
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
import time
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
```
Finally, we'll train our model. This takes considerably longer than any of the previous models due to the size of the transformer. Even though we are not training any of the transformer's parameters we still need to pass the data through the model which takes a considerable amount of time on a standard GPU.
```
N_EPOCHS = 5
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss, train_acc = train(model, train_iterator, optimizer, criterion)
valid_loss, valid_acc = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut6-model.pt')
print(f'Epoch: {epoch+1:02} | Epoch Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}%')
```
We'll load up the parameters that gave us the best validation loss and try these on the test set - which gives us our best results so far!
```
model.load_state_dict(torch.load('tut6-model.pt'))
test_loss, test_acc = evaluate(model, test_iterator, criterion)
print(f'Test Loss: {test_loss:.3f} | Test Acc: {test_acc*100:.2f}%')
```
## Inference
We'll then use the model to test the sentiment of some sequences. We tokenize the input sequence, trim it down to the maximum length, add the special tokens to either side, convert it to a tensor, add a fake batch dimension and then pass it through our model.
```
def predict_sentiment(model, tokenizer, sentence):
model.eval()
tokens = tokenizer.tokenize(sentence)
tokens = tokens[:max_input_length-2]
indexed = [init_token_idx] + tokenizer.convert_tokens_to_ids(tokens) + [eos_token_idx]
tensor = torch.LongTensor(indexed).to(device)
tensor = tensor.unsqueeze(0)
prediction = torch.sigmoid(model(tensor))
return prediction.item()
predict_sentiment(model, tokenizer, "This film is terrible")
predict_sentiment(model, tokenizer, "This film is great")
```
| github_jupyter |
# FKLearn Tutorial:
* <font size="4"> FKlearn is the nubank functional library for Machine Learning </font>
* <font size="4"> It was created with the idea of scaling machine learning through the company by standardizing model development and implementing an easy interface to allow all users to develop the best practices on Machine Learning </font>
# Input Analysis
## Imports
```
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
import matplotlib
sns.set_style("whitegrid")
sns.set_palette("husl")
import warnings
warnings.filterwarnings('ignore')
```
## Input Dataset
* <font size="4"> This dataset was created with simulated data about users spend behavior on Credit Card </font>
* <font size="4"> The model target is the average spend of the next 2 months and we created some features that can be related to that target </font>
```
# Generate this dataset using the FKLearn Tutorial Dataset.ipynb notebook
df = pd.read_csv("fklearn-tutorial-input-dataset.csv")
df['month_date'] = pd.to_datetime(df.month_date)
df.head()
df.describe().T
features = ["income", "bureau_score", "spend_desire", "random_noise", "monthly_spend", "avg_last_2_months_spend"]
df.isna().sum()
```
## Features and Target:
* <font size="4"> Month (M): Number of months since "2017-01-01" </font>
* <font size="4"> Income (I): N(5000, 2000) E [300, 20000] </font>
* <font size="4"> Phone Type (P): 4 categories </font>
* <font size="4"> Bureau Score (B): N(500 / (Month ** 0.1), 200) E [0, 1000] </font>
* <font size="4"> Spend Desire (W): N(500, 200) </font>
* <font size="4"> Random Noise (R): N(1000, 100) </font>
* <font size="4"> Monthly Spend: Max(0, (S * I + I ** 2 + P * W ** 2 + P * B + N(1, 0.3)) * N(2000, 1000)) </font>
* <font size="4"> Avg Last 2 Months Spend: (Spend(m) + Spend(m-1)) / 2 </font>
* <font size="4"> Target: (Spend(m + 1) + Spend(m + 2)) / 2 </font>
```
plt.plot(sorted(df.month.unique()), df.groupby("month").agg({"id": "count"}))
plt.title("Amount of customers by month")
fig, axes = plt.subplots(2, 2, figsize=(8,8))
axes[0, 0].hist(df.bureau_score, range(0, 1000, 50))
axes[1, 0].hist(df.income, range(300, 20000, 500))
axes[0, 1].hist(df.spend_desire)
axes[1, 1].hist(df.monthly_spend, range(0, 10000, 500))
titles = ["bureau_histogram", "income_histogram", "spend_desire_histogram", "monthly_spend_histogram"]
axes[0, 0].set_title(titles[0])
axes[1, 0].set_title(titles[1])
axes[0, 1].set_title(titles[2])
axes[1, 1].set_title(titles[3])
plt.show()
fig, axes = plt.subplots(2, 2, figsize=(10,10))
sns.pointplot(x="month", y="bureau_score", data=df, ax=axes[0, 0])
sns.pointplot(x="month", y="income", data=df, ax=axes[1, 0])
sns.pointplot(x="month", y="spend_desire", data=df, ax=axes[0, 1])
sns.pointplot(x="month", y="monthly_spend", data=df, ax=axes[1, 1])
plt.show()
```
## Target Analysis
```
pd.DataFrame(df.groupby("month_date").apply(lambda x: x.target.isna().sum()), columns=["null_count_by_month"])
sns.pointplot(x="month", y="target", data=df)
```
# Getting started with fklearn: Creating a simple model
* <font size="4"> Fklearn allows you to get production ready models with ease </font>
* <font size="4"> Tradional steps of training a model like: Train-Test Splitting, Training and Validating are all implemented on fklearn </font>
## Spliting the dataset into train and holdout
* <font size="4"> On real problems we want to validate our model both in the same period and also on a different period to guarantee our model is a good predictor for the future </font>
<img src="imgs/Splitting_Data.png">
```
from fklearn.preprocessing.splitting import space_time_split_dataset
train_set, intime_outspace_hdout, outime_inspace_hdout, outime_outspace_hdout = \
space_time_split_dataset(df,
train_start_date="2017-02-01",
train_end_date="2018-07-01",
holdout_end_date="2018-10-01",
split_seed=42,
space_holdout_percentage=0.2,
space_column="id",
time_column="month_date")
(train_set.shape,
intime_outspace_hdout.shape,
outime_inspace_hdout.shape,
outime_outspace_hdout.shape)
```
## Training:
* <font size="4"> On the training process we want to cap features that have unexpected values </font>
* <font size="4"> We want to encode categorical features </font>
* <font size="4"> We might want to fill missing values </font>
* <font size="4"> We want to train our model </font>
* <font size="4"> We want this transformations to be applied when scoring </font>
* <font size="4"> We want to apply this same transformations when scoring our model, similarly to the fit and transform concept from sklearn </font>
<img src="imgs/Pipeline.png">
```
from fklearn.training.transformation import capper, prediction_ranger, label_categorizer
from fklearn.training.imputation import imputer
from fklearn.training.regression import lgbm_regression_learner
capper_fn = capper(columns_to_cap=["income"], precomputed_caps={"income": 20000.0})
ranger_fn = prediction_ranger(prediction_min=0.0, prediction_max=100000.0, prediction_column="prediction")
label_fn = label_categorizer(columns_to_categorize=["phone_type"])
imputer_fn = imputer(columns_to_impute=["bureau_score"], impute_strategy="median")
regression_fn = lgbm_regression_learner(features=features, target="target", learning_rate=0.1, num_estimators=200)
from fklearn.training.pipeline import build_pipeline
train_fn = build_pipeline(label_fn, capper_fn, imputer_fn, regression_fn, ranger_fn)
predict_fn, scored_train_set, train_logs = train_fn(train_set)
scored_train_set.head()
pd.options.display.max_rows = 120
pd.DataFrame(train_logs['lgbm_regression_learner']['feature_importance'],
index=["importance"]).T.sort_values("importance", ascending=False)
scored_outime_outspace_hdout = predict_fn(outime_outspace_hdout)
scored_intime_outspace_hdout = predict_fn(intime_outspace_hdout)
scored_outime_inspace_hdout = predict_fn(outime_inspace_hdout)
scored_outime_outspace_hdout[["id", "month_date", "income", "bureau_score", "target", "prediction"]].head()
```
## Evaluation:
* <font size="4"> We want to evaluate the performance of our model using multiple metrics for each dataframe (out_of_time, out_of_space, out_of_time_and_space) </font>
<img src="imgs/EvalFn.png">
```
from fklearn.validation.evaluators import r2_evaluator, spearman_evaluator, combined_evaluators
r2_eval_fn = r2_evaluator(prediction_column="prediction", target_column="target")
spearman_eval_fn = spearman_evaluator(prediction_column="prediction", target_column="target")
eval_fn = combined_evaluators(evaluators=[r2_eval_fn, spearman_eval_fn])
outime_outspace_hdout_logs = eval_fn(scored_outime_outspace_hdout)
intime_outspace_hdout_logs = eval_fn(scored_intime_outspace_hdout)
outime_inspace_hdout_logs = eval_fn(scored_outime_inspace_hdout)
{"out_of_time": outime_outspace_hdout_logs,
"in_time": intime_outspace_hdout_logs,
"out_of_time_and_space": outime_inspace_hdout_logs}
```
## Extractors
* <font size="4"> We want to transform this logs into dataframes that we can visualize better </font>
<img src="imgs/Extractor.png" style="width: 500px;">
```
from fklearn.metrics.pd_extractors import evaluator_extractor, combined_evaluator_extractor, extract
r2_extractor = evaluator_extractor(evaluator_name="r2_evaluator__target")
spearman_extractor = evaluator_extractor(evaluator_name="spearman_evaluator__target")
full_extractor = combined_evaluator_extractor(base_extractors=[r2_extractor, spearman_extractor])
pd.concat(
[full_extractor(outime_outspace_hdout_logs).assign(part="out_of_time"),
full_extractor(intime_outspace_hdout_logs).assign(part="in_time_out_of_space"),
full_extractor(outime_inspace_hdout_logs).assign(part="out_of_time_and_space")])
```
# Learning Curves:
<img src="imgs/LearningCurves.png">
<font size="5"> We can reuse our previously defined training function and evaluators! </font>
## Could we improve if we had more data, AKA Spatial Learning Curve
* <font size="4"> If we had more data we could use that for training and see how much the model improves </font>
* <font size="4"> To estimate that we can train with subsamples of the full training set and check how performance changes </font>
<img src="imgs/SpatialLearningCurves.png">
```
from fklearn.validation.splitters import spatial_learning_curve_splitter
split_fn = spatial_learning_curve_splitter(space_column="id",
time_column="month_date",
training_limit="2018-04-01",
train_percentages=[0.1, 0.2, 0.4, 0.6, 0.8, 1.0])
from fklearn.validation.validator import parallel_validator
spatial_learning_curve_logs = parallel_validator(train_set, split_fn, train_fn, eval_fn, n_jobs=8)
spatial_learning_curve_logs
data = extract(spatial_learning_curve_logs['validator_log'], full_extractor)
data
fig, ax = plt.subplots(figsize=(10, 8))
sns.pointplot(data=data, ax=ax, x="percentage", y="r2_evaluator__target")
plt.title("Spatial learning curve (trained from %s to %s)" % (data.train_start.dt.date.min(), data.train_end.dt.date.max()));
```
## Performance over time
* <font size="4"> That metric can be thought as split your dataset month by month and computing the desired metrics </font>
* <font size="4"> We can do that with ease using our split evaluators </font>
```
from fklearn.validation.evaluators import split_evaluator
from fklearn.metrics.pd_extractors import split_evaluator_extractor
out_of_space_holdout = pd.concat([scored_intime_outspace_hdout, scored_outime_outspace_hdout])
monthly_eval_fn = split_evaluator(eval_fn=eval_fn,
split_col="month",
split_values=list(range(0, 25)))
monthly_extractor = split_evaluator_extractor(base_extractor=full_extractor,
split_col="month",
split_values=list(range(0, 25)))
out_of_space_logs = monthly_eval_fn(out_of_space_holdout)
monthly_performance = monthly_extractor(out_of_space_logs)
monthly_performance
fig, ax = plt.subplots(figsize=(10, 8))
sns.pointplot(data=monthly_performance, ax=ax, x="split_evaluator__month", y="r2_evaluator__target")
```
## Impact of More Data on Monthly Performance?
* <font size="4"> We want both our previous way of spliting our dataset and the evaluator we created </font>
* <font size="4"> We can use both to create a new spatial learning curve! </font>
```
monthly_spatial_learning_curve_logs = parallel_validator(train_set,
split_fn,
train_fn,
monthly_eval_fn,
n_jobs=8)
monthly_data = extract(monthly_spatial_learning_curve_logs['validator_log'], monthly_extractor).loc[lambda df: df.r2_evaluator__target.notna()]
monthly_data
fig, ax = plt.subplots(figsize=(10, 8))
sns.pointplot(data=monthly_data, ax=ax, x="percentage", y="r2_evaluator__target", hue="split_evaluator__month")
plt.title("Spatial Learning Curve By Month")
plt.xticks(rotation=45);
```
# What else does fklearn provide me?
* <font size="4"> Several other learning curves that can be used depending on what you want to evaluate </font>
* <font size="4"> Several other algorithms for models </font>
* <font size="4"> Other tools with similar interface for feature selection, parameter tuning </font>
* <font size="4"> All this methods are integrated with similar signatures in a way that is easy to reuse training, spliting and evaluation functions </font>
| github_jupyter |
```
%matplotlib inline
import os
import csv
import codecs
import numpy as np
import pandas as pd
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
from sklearn.utils import shuffle
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import log_loss
import datetime
seed = 111
np.random.seed(seed)
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.utils.np_utils import to_categorical
from keras.layers import Dense, Input, Flatten, Concatenate, LSTM, Lambda, Dropout, Multiply
from keras.layers import Conv1D, MaxPooling1D, Embedding, SpatialDropout1D, GRU
from keras.layers.merge import _Merge
from keras.models import Model
from keras.layers.wrappers import TimeDistributed, Bidirectional
from keras.layers.normalization import BatchNormalization
from keras import backend as K
from keras.utils import plot_model
from keras.callbacks import EarlyStopping, ModelCheckpoint
import sys
from keras_tqdm import TQDMNotebookCallback
BASE_DIR = 'input/'
GLOVE_DIR = 'WordEmbeddings/Glove/'
TRAIN_DATA_FILE = BASE_DIR + 'train.csv'
TEST_DATA_FILE = BASE_DIR + 'test.csv'
MAX_SEQUENCE_LENGTH = 80
MAX_NUM_WORDS = 500000
EMBEDDING_DIM = 300
STATE_DIM = 300
DROP = 0.2
NFOLDS = 10
print('Indexing word vectors.')
embeddings_index = {}
f = codecs.open(os.path.join(GLOVE_DIR, 'glove.840B.300d.txt'), encoding='utf-8')
for line in f:
values = line.split(' ')
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Found %s word vectors.' % len(embeddings_index))
print('Processing text dataset')
train = pd.read_csv(BASE_DIR + 'train.csv', encoding='utf-8')
test = pd.read_csv(BASE_DIR + 'test.csv', encoding='utf-8')
ids = train.id
texts_1 = train.question1.astype(np.str).tolist()
texts_2 = train.question2.astype(np.str).tolist()
labels = train.is_duplicate.tolist() # list of label ids
print('Found %s texts.' % len(texts_1))
test_texts_1 = test.question1.astype(np.str).tolist()
test_texts_2 = test.question2.astype(np.str).tolist()
test_labels = [0] * len(test) # list of label ids
print('Found %s texts.' % len(test_texts_1))
tokenizer = Tokenizer(num_words=MAX_NUM_WORDS)
tokenizer.fit_on_texts(texts_1 + texts_2 + test_texts_1 + test_texts_2)
sequences_1 = tokenizer.texts_to_sequences(texts_1)
sequences_2 = tokenizer.texts_to_sequences(texts_2)
word_index = tokenizer.word_index
print('Found %s unique tokens.' % len(word_index))
test_sequences_1 = tokenizer.texts_to_sequences(test_texts_1)
test_sequences_2 = tokenizer.texts_to_sequences(test_texts_2)
data_1 = pad_sequences(sequences_1, maxlen=MAX_SEQUENCE_LENGTH)
data_2 = pad_sequences(sequences_2, maxlen=MAX_SEQUENCE_LENGTH)
labels = np.array(labels)
print('Shape of data tensor:', data_1.shape)
print('Shape of label tensor:', labels.shape)
test_data_1 = pad_sequences(test_sequences_1, maxlen=MAX_SEQUENCE_LENGTH)
test_data_2 = pad_sequences(test_sequences_2, maxlen=MAX_SEQUENCE_LENGTH)
test_labels = np.array(test_labels)
del test_sequences_1
del test_sequences_2
del sequences_1
del sequences_2
import gc
gc.collect()
print('Preparing embedding matrix.')
# prepare embedding matrix
num_words = min(MAX_NUM_WORDS, len(word_index))
embedding_matrix = np.zeros((num_words, EMBEDDING_DIM))
count = 0
for word, i in word_index.items():
if i >= num_words:
count += 1
continue
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
# words not found in embedding index will be zeros
embedding_matrix[i] = embedding_vector
count += 1
print('Unknown word embeddings:', num_words - count)
# def euc_dist(x):
# 'Merge function: euclidean_distance(u,v)'
# s = x[0] - x[1]
# output = (s ** 2).sum(axis=1)
# output = K.reshape(output, (output.shape[0],1))
# return output
# def euc_dist_shape(input_shape):
# 'Merge output shape'
# shape = list(input_shape)
# outshape = (shape[0][0],1)
# return tuple(outshape)
class Subtract(_Merge):
"""Layer that adds a list of inputs.
It takes as input a list of tensors,
all of the same shape, and returns
a single tensor (also of the same shape).
"""
def _merge_function(self, inputs):
return K.square(inputs[0] - inputs[1])
embedding_layer = Embedding(num_words, EMBEDDING_DIM, weights=[embedding_matrix],
input_length=MAX_SEQUENCE_LENGTH, trainable=False)
def siamese_architecture(seq_len, embed_len, state_len):
inputs = Input(shape=(seq_len, embed_len))
x = Bidirectional(GRU(units=state_len, dropout=DROP, recurrent_dropout=DROP,
implementation=2, return_sequences=True))(inputs)
x = Bidirectional(GRU(units=state_len, dropout=DROP, recurrent_dropout=DROP,
implementation=2))(x)
return Model(inputs=inputs, outputs=x)
# Model Architecture #
def create_model():
siamese_arch = siamese_architecture(MAX_SEQUENCE_LENGTH, EMBEDDING_DIM, STATE_DIM)
sequence_1_input = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
embedded_sequences_1 = embedding_layer(sequence_1_input)
x1 = siamese_arch(embedded_sequences_1)
sequence_2_input = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
embedded_sequences_2 = embedding_layer(sequence_2_input)
y1 = siamese_arch(embedded_sequences_2)
# merged = Concatenate()([x1, y1])
merged_sub = Subtract()([x1, y1])
merged_mult = Multiply()([x1, y1])
merged_comb = Concatenate()([x1, y1, merged_sub, merged_mult])
merged = BatchNormalization()(merged_comb)
merged = Dense(512, activation='relu')(merged)
merged = BatchNormalization()(merged)
merged = Dense(128, activation='relu')(merged)
# merged = Dropout(DROP)(merged)
merged = BatchNormalization()(merged)
preds = Dense(1, activation='sigmoid')(merged)
model = Model(inputs=[sequence_1_input, sequence_2_input], outputs=preds)
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['acc'])
return model
model = create_model()
print(data_1.shape, data_2.shape, labels.shape)
model.summary()
plot_model(model, to_file='model.png')
img = mpimg.imread('model.png')
fig = plt.figure(figsize=(15, 15))
plt.imshow(img)
BATCH_SIZE = 512
skf = StratifiedKFold(n_splits=NFOLDS, shuffle=True, random_state=seed).split(data_1, labels)
test_preds = np.zeros((len(test_data_1), NFOLDS))
val_preds = np.zeros(len(data_1))
train_rnn_vals = np.zeros((len(data_1), STATE_DIM * 4), dtype=np.float32)
test_rnn_vals = np.zeros((len(test_data_1), STATE_DIM * 4), dtype=np.float32)
X_test = [test_data_1, test_data_2]
now = datetime.datetime.now()
for i, (idx_train, idx_val) in enumerate(skf):
print('Fold', i+1)
X_train = [data_1[idx_train], data_2[idx_train]]
X_val = [data_1[idx_val], data_2[idx_val]]
y_train = labels[idx_train]
y_val = labels[idx_val]
callbacks = [TQDMNotebookCallback(),
EarlyStopping(patience=0),
ModelCheckpoint('weights_gru{}.hdf5'.format(i), save_best_only=True, save_weights_only=True)]
model = create_model()
model.fit(X_train, y_train, validation_data=(X_val, y_val), epochs=50, batch_size=BATCH_SIZE,
callbacks=callbacks, shuffle=True, verbose=0)
model.load_weights('weights_gru{}.hdf5'.format(i))
# feature_extractor = Model(inputs=[model.layers[0].input, model.layers[1].input],
# outputs=[model.layers[6].input[0], model.layers[6].input[1], model.layers[-1].output])
# feature_extractor.predict_function = K.function(feature_extractor._feed_inputs + [K.learning_phase()],
# feature_extractor.outputs,
# updates=feature_extractor.state_updates,
# name='predict_function',
# **(getattr(feature_extractor, '_function_kwargs', {})))
# outs = feature_extractor.predict(X_val, batch_size=BATCH_SIZE)
# train_rnn_vals[idx_val, :600] = outs[0]
# train_rnn_vals[idx_val, 600:] = outs[1]
val_preds[idx_val] = model.predict(X_val, batch_size=BATCH_SIZE)
# outs = feature_extractor.predict(X_test, batch_size=BATCH_SIZE)
# test_rnn_vals[:, :600] = outs[0]
# test_rnn_vals[:, 600:] = outs[1]
test_preds[:, i] = model.predict(X_test, batch_size=BATCH_SIZE).reshape((-1,))
# pd.to_pickle(test_rnn_vals, 'GRU_hidden_states_test_{}_{:%Y%m%d_%H%M}.pkl'.format(i+1, now))
# del model, feature_extractor, outs, test_rnn_vals
# gc.collect()
a = 0.165 / 0.369191399096
b = (1 - 0.165) / (1 - 0.369191399096)
def pred_transform(preds):
return a * preds / (a * preds + b * (1 - preds))
from sklearn.metrics import log_loss
test_df = pd.DataFrame({"test_id": np.arange(len(test_data_1)), "is_duplicate":test_preds.mean(1)})
test_preds_mod = pred_transform(test_preds)
test_df_mod = pd.DataFrame({"test_id": np.arange(len(test_data_1)), "is_duplicate":test_preds_mod.mean(1)})
val_df = pd.DataFrame({"id": np.arange(len(data_1)), "is_duplicate": val_preds})
loss = log_loss(train.is_duplicate, val_preds)
print('Log Loss:', loss)
# now = datetime.datetime.now()
test_pred_filename = "model_out/test_preds_gru_{:.4f}_{:%Y%m%d_%H%M}.csv.gz".format(loss, now)
test_df.to_csv(test_pred_filename, index=False, compression='gzip')
test_pred_mod_filename = "model_out/test_preds_gru_{:.4f}_{:%Y%m%d_%H%M}_mod.csv.gz".format(loss, now)
test_df_mod.to_csv(test_pred_mod_filename, index=False, compression='gzip')
val_pred_filename = "model_out/train_preds_gru_{:.4f}_{:%Y%m%d_%H%M}.csv.gz".format(loss, now)
val_df.to_csv(val_pred_filename, index=False, compression='gzip')
# pd.to_pickle(train_rnn_vals, 'GRU_hidden_states_train_{:%Y%m%d_%H%M}.pkl'.format(now))
```
| github_jupyter |
```
# Install the latest Tensorflow version.
!pip3 install --quiet "tensorflow>=1.7"
# Install TF-Hub.
!pip3 install --quiet "tensorflow-hub>=0.7.0"
!pip3 install --quiet seaborn
from absl import logging
import tensorflow.compat.v1 as tf
import tensorflow_hub as hub
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import re
import seaborn as sns
import json
import pandas as pd
from tqdm import tqdm_notebook
module_url = "https://tfhub.dev/google/universal-sentence-encoder/2" #@param ["https://tfhub.dev/google/universal-sentence-encoder/2", "https://tfhub.dev/google/universal-sentence-encoder-large/3"]
tf.disable_v2_behavior()
tf.compat.v1.disable_eager_execution()
stocknet_dataset_filepath = './stocknet-dataset-master'
preprocessed_prices_filepath = stocknet_dataset_filepath + '/price/preprocessed'
preprocessed_tweets_filepath = stocknet_dataset_filepath + '/tweet/preprocessed'
company_to_price_df = {}
company_to_tweets = {}
for filename in os.listdir(preprocessed_prices_filepath):
with open(preprocessed_prices_filepath + '/' + filename) as file:
company_name = filename.split('.')[0]
# Not enough data for GMRE
if company_name == 'GMRE':
continue
df = pd.read_csv(file, sep='\t')
df.columns = ['date', 'open', 'high', 'low', 'close', 'adjust_close', 'volume']
company_to_price_df[company_name] = df
for filename in tqdm_notebook(os.listdir(preprocessed_tweets_filepath)):
company_name = filename.split('.')[0]
dates_to_tweets = {}
for tweet_filename in os.listdir(preprocessed_tweets_filepath + '/' + filename):
with open(preprocessed_tweets_filepath + '/' + filename + '/' + tweet_filename) as file:
list_of_tweets = []
for line in file:
tweet_json = json.loads(line)
list_of_tweets.append(tweet_json)
dates_to_tweets[tweet_filename] = list_of_tweets
company_to_tweets[company_name] = dates_to_tweets
#print(company_to_tweets.keys())
#print(dates_to_tweets.keys())
print(company_to_tweets['AAPL']['2015-10-02'][0])
# company_to_tweets maps each company name to a dictionary of dates and list of tweets. Schema:
{
company_name:
{
date: [list of tweets + metadata]
...
}
...
}
# Import the Universal Sentence Encoder's TF Hub module
embed = hub.Module(module_url)
# Reduce logging output.
logging.set_verbosity(logging.ERROR)
tf.get_logger().setLevel(logging.ERROR)
tf.compat.v1.logging.set_verbosity(tf.compat.v1.logging.ERROR)
with tf.Session() as session:
session.run([tf.global_variables_initializer(), tf.tables_initializer()])
for company in ['AAPL']:
#for company in company_to_tweets.keys():
for date in tqdm_notebook(company_to_tweets[company].keys()):
messages = []
for j in range(len(company_to_tweets[company][date])):
messages.append(' '.join(company_to_tweets[company][date][j]['text']))
message_embeddings = session.run(embed(messages))
for k in range(len(company_to_tweets[company][date])):
company_to_tweets[company][date][k]['embedding'] = list(message_embeddings[k])
```
| github_jupyter |
# Machine Learning Engineer Nanodegree
## Supervised Learning
## Project 2: Building a Student Intervention System
Welcome to the second project of the Machine Learning Engineer Nanodegree! In this notebook, some template code has already been provided for you, and it will be your job to implement the additional functionality necessary to successfully complete this project. Sections that begin with **'Implementation'** in the header indicate that the following block of code will require additional functionality which you must provide. Instructions will be provided for each section and the specifics of the implementation are marked in the code block with a `'TODO'` statement. Please be sure to read the instructions carefully!
In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
>**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
### Question 1 - Classification vs. Regression
*Your goal for this project is to identify students who might need early intervention before they fail to graduate. Which type of supervised learning problem is this, classification or regression? Why?*
**Answer: ** This is a classification problem. We need to identify students that need intervention before fail to graduate. It means that our output must be something like "yes" or "no", 0 or 1, "need intervention" or "doesn't need intervention". These are a set of limited outputs, i.e., these are discrete ou categorical values for our output. If this were a regression problem, the output would be continuous values.
## Exploring the Data
Run the code cell below to load necessary Python libraries and load the student data. Note that the last column from this dataset, `'passed'`, will be our target label (whether the student graduated or didn't graduate). All other columns are features about each student.
```
# Import libraries
import numpy as np
import pandas as pd
from time import time
from sklearn.metrics import f1_score
from __future__ import division
# Read student data
student_data = pd.read_csv("student-data.csv")
print "Student data read successfully!"
```
### Implementation: Data Exploration
Let's begin by investigating the dataset to determine how many students we have information on, and learn about the graduation rate among these students. In the code cell below, you will need to compute the following:
- The total number of students, `n_students`.
- The total number of features for each student, `n_features`.
- The number of those students who passed, `n_passed`.
- The number of those students who failed, `n_failed`.
- The graduation rate of the class, `grad_rate`, in percent (%).
```
student_data.columns
# TODO: Calculate number of students
n_students = len(student_data)
# TODO: Calculate number of features
n_features = len(student_data.columns) -1
# TODO: Calculate passing students
n_passed = len(student_data[student_data.passed=="yes"])
# TODO: Calculate failing students
n_failed = len(student_data[student_data.passed=="no"])
# TODO: Calculate graduation rate
grad_rate =n_passed/n_students * 100
# Print the results
print "Total number of students: {}".format(n_students)
print "Number of features: {}".format(n_features)
print "Number of students who passed: {}".format(n_passed)
print "Number of students who failed: {}".format(n_failed)
print "Graduation rate of the class: {:.2f}%".format(grad_rate)
```
## Preparing the Data
In this section, we will prepare the data for modeling, training and testing.
### Identify feature and target columns
It is often the case that the data you obtain contains non-numeric features. This can be a problem, as most machine learning algorithms expect numeric data to perform computations with.
Run the code cell below to separate the student data into feature and target columns to see if any features are non-numeric.
```
# Extract feature columns
feature_cols = list(student_data.columns[:-1])
# Extract target column 'passed'
target_col = student_data.columns[-1]
# Show the list of columns
print "Feature columns:\n{}".format(feature_cols)
print "\nTarget column: {}".format(target_col)
# Separate the data into feature data and target data (X_all and y_all, respectively)
X_all = student_data[feature_cols]
y_all = student_data[target_col]
# Show the feature information by printing the first five rows
print "\nFeature values:"
print X_all.head()
```
### Preprocess Feature Columns
As you can see, there are several non-numeric columns that need to be converted! Many of them are simply `yes`/`no`, e.g. `internet`. These can be reasonably converted into `1`/`0` (binary) values.
Other columns, like `Mjob` and `Fjob`, have more than two values, and are known as _categorical variables_. The recommended way to handle such a column is to create as many columns as possible values (e.g. `Fjob_teacher`, `Fjob_other`, `Fjob_services`, etc.), and assign a `1` to one of them and `0` to all others.
These generated columns are sometimes called _dummy variables_, and we will use the [`pandas.get_dummies()`](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.get_dummies.html?highlight=get_dummies#pandas.get_dummies) function to perform this transformation. Run the code cell below to perform the preprocessing routine discussed in this section.
```
def preprocess_features(X):
''' Preprocesses the student data and converts non-numeric binary variables into
binary (0/1) variables. Converts categorical variables into dummy variables. '''
# Initialize new output DataFrame
output = pd.DataFrame(index = X.index)
# Investigate each feature column for the data
for col, col_data in X.iteritems():
# If data type is non-numeric, replace all yes/no values with 1/0
if col_data.dtype == object:
col_data = col_data.replace(['yes', 'no'], [1, 0])
# If data type is categorical, convert to dummy variables
if col_data.dtype == object:
# Example: 'school' => 'school_GP' and 'school_MS'
col_data = pd.get_dummies(col_data, prefix = col)
# Collect the revised columns
output = output.join(col_data)
return output
X_all = preprocess_features(X_all)
print "Processed feature columns ({} total features):\n{}".format(len(X_all.columns), list(X_all.columns))
X_all.head()
```
### Implementation: Training and Testing Data Split
So far, we have converted all _categorical_ features into numeric values. For the next step, we split the data (both features and corresponding labels) into training and test sets. In the following code cell below, you will need to implement the following:
- Randomly shuffle and split the data (`X_all`, `y_all`) into training and testing subsets.
- Use 300 training points (approximately 75%) and 95 testing points (approximately 25%).
- Set a `random_state` for the function(s) you use, if provided.
- Store the results in `X_train`, `X_test`, `y_train`, and `y_test`.
```
# TODO: Import any additional functionality you may need here
from sklearn.model_selection import train_test_split
# TODO: Set the number of training points
num_train = 300
# Set the number of testing points
num_test = X_all.shape[0] - num_train
# TODO: Shuffle and split the dataset into the number of training and testing points above
# X_train = None
# X_test = None
# y_train = None
# y_test = None
X_train, X_test, y_train, y_test = train_test_split(X_all, y_all, test_size=num_test, random_state=1973, stratify=y_all)
# Show the results of the split
print "Training set has {} samples.".format(X_train.shape[0])
print "Testing set has {} samples.".format(X_test.shape[0])
```
## Training and Evaluating Models
In this section, you will choose 3 supervised learning models that are appropriate for this problem and available in `scikit-learn`. You will first discuss the reasoning behind choosing these three models by considering what you know about the data and each model's strengths and weaknesses. You will then fit the model to varying sizes of training data (100 data points, 200 data points, and 300 data points) and measure the F<sub>1</sub> score. You will need to produce three tables (one for each model) that shows the training set size, training time, prediction time, F<sub>1</sub> score on the training set, and F<sub>1</sub> score on the testing set.
**The following supervised learning models are currently available in** [`scikit-learn`](http://scikit-learn.org/stable/supervised_learning.html) **that you may choose from:**
- **Gaussian Naive Bayes (GaussianNB)**
- Decision Trees
- **Ensemble Methods (Bagging, AdaBoost, Random Forest, Gradient Boosting)**
- K-Nearest Neighbors (KNeighbors)
- Stochastic Gradient Descent (SGDC)
- Support Vector Machines (SVM)
- **Logistic Regression**
### Question 2 - Model Application
*List three supervised learning models that are appropriate for this problem. For each model chosen*
- Describe one real-world application in industry where the model can be applied. *(You may need to do a small bit of research for this — give references!)*
- What are the strengths of the model; when does it perform well?
- What are the weaknesses of the model; when does it perform poorly?
- What makes this model a good candidate for the problem, given what you know about the data?
**Answer: **
#### Naive Bayes ####
- Naive Bayes has been used in cases such as face recognition, sentiment analysis, spam detection and text sorting.
- **Strengths:**
- Easy and fast to implement
- Easy to understand
- Can be used for online learning
- **Weaknesses:**
- Fails estimating rare occurrences
- Suffers from irrelevant features
- Strong feature indepence assumptions
- **Applicability: **
- Naive Bayes is very simple and very cheap in terms of computation and, since it is good with small datasets for training I thing it is a good candidate to solve our problem.
- _ref: Building Machine Learning with Python, Willi Richert and Luis Pedro Coelho - Packt Publishing. Page 146_
#### Ensemble Methods - Random Forest ####
- Applications include classification of internet traffic, bioinformatics like , gene expression classification, mass spectrum protein expression analysis, biomarker discovery, sequence annotation, protein-protein interaction prediction.
- **Strengths:**
- Classification tree ensembles perform very well in practice. They are robust to outliers, scalable, and able to naturally model non-linear decision boundaries thanks to their hierarchical structure.
- **Weaknesses:**
- Difficult to interpret
- Weaker on regression whe estimating values at the extremities of the distribution of response values
- Biased in multiclass problems toward more frequent classes
- **Applicability: **
- Decision trees are very good in classification problems, and using Random Forest we have more consistent results. Since random forest will create a bunch of random decision trees, it can deal with these results to pick the best. Another point is that in our case, we didn't even create the dummy variables because random forest is able to work with discrete and categorical variables. Besides that, random forest is not prone to overfit the model like a single decision tree.
#### Logistic Regression ####
- Applications include ordering results by probability, modelling marketing responses.
- **Strengths**:
- Outputs have a nice probabilistic interpretation, and the algorithm can be regularized to avoid overfitting. Logistic models can be updated easily with new data using stochastic gradient descent.
- **Weaknesses:**
- Logistic regression tends to underperform when there are multiple or non-linear decision boundaries. They are not flexible enough to naturally capture more complex relationships.
- **Applicability: **
- We have a binary classification with most part our features as continuous and/or categorical data.
_Refs:_
[Machine Learning for Dummies Cheat Sheet](http://www.dummies.com/programming/big-data/data-science/machine-learning-dummies-cheat-sheet/)
[Modern Machine Learning Algorithms: Strengths and Weaknesses](https://elitedatascience.com/machine-learning-algorithms)
### Setup
Run the code cell below to initialize three helper functions which you can use for training and testing the three supervised learning models you've chosen above. The functions are as follows:
- `train_classifier` - takes as input a classifier and training data and fits the classifier to the data.
- `predict_labels` - takes as input a fit classifier, features, and a target labeling and makes predictions using the F<sub>1</sub> score.
- `train_predict` - takes as input a classifier, and the training and testing data, and performs `train_clasifier` and `predict_labels`.
- This function will report the F<sub>1</sub> score for both the training and testing data separately.
```
def train_classifier(clf, X_train, y_train):
''' Fits a classifier to the training data. '''
# Start the clock, train the classifier, then stop the clock
start = time()
clf.fit(X_train, y_train)
end = time()
# Print the results
print "Trained model in {:.4f} seconds".format(end - start)
def predict_labels(clf, features, target):
''' Makes predictions using a fit classifier based on F1 score. '''
# Start the clock, make predictions, then stop the clock
start = time()
y_pred = clf.predict(features)
end = time()
# Print and return results
print "Made predictions in {:.4f} seconds.".format(end - start)
return f1_score(target.values, y_pred, pos_label='yes')
def train_predict(clf, X_train, y_train, X_test, y_test):
''' Train and predict using a classifer based on F1 score. '''
# Indicate the classifier and the training set size
print "Training a {} using a training set size of {}. . .".format(clf.__class__.__name__, len(X_train))
# Train the classifier
train_classifier(clf, X_train, y_train)
# Print the results of prediction for both training and testing
print "F1 score for training set: {:.4f}.".format(predict_labels(clf, X_train, y_train))
print "F1 score for test set: {:.4f}.".format(predict_labels(clf, X_test, y_test))
```
### Implementation: Model Performance Metrics
With the predefined functions above, you will now import the three supervised learning models of your choice and run the `train_predict` function for each one. Remember that you will need to train and predict on each classifier for three different training set sizes: 100, 200, and 300. Hence, you should expect to have 9 different outputs below — 3 for each model using the varying training set sizes. In the following code cell, you will need to implement the following:
- Import the three supervised learning models you've discussed in the previous section.
- Initialize the three models and store them in `clf_A`, `clf_B`, and `clf_C`.
- Use a `random_state` for each model you use, if provided.
- **Note:** Use the default settings for each model — you will tune one specific model in a later section.
- Create the different training set sizes to be used to train each model.
- *Do not reshuffle and resplit the data! The new training points should be drawn from `X_train` and `y_train`.*
- Fit each model with each training set size and make predictions on the test set (9 in total).
**Note:** Three tables are provided after the following code cell which can be used to store your results.
```
# TODO: Import the three supervised learning models from sklearn
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import GaussianNB
# TODO: Initialize the three models
clf_A = GaussianNB()
clf_B = LogisticRegression(random_state=1973)
clf_C = RandomForestClassifier(random_state=1973)
# TODO: Set up the training set sizes
X_train_100 = X_train[:100]
y_train_100 = y_train[:100]
X_train_200 = X_train[:200]
y_train_200 = y_train[:200]
X_train_300 = X_train
y_train_300 = y_train
# TODO: Execute the 'train_predict' function for each classifier and each training set size
# train_predict(clf, X_train, y_train, X_test, y_test)
for clf in [clf_A, clf_B, clf_C]:
for size in [100, 200, 300]:
train_predict(clf, X_train[:size], y_train[:size], X_test, y_test)
print '='*80 # imprime linha
```
### Tabular Results
Edit the cell below to see how a table can be designed in [Markdown](https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet#tables). You can record your results from above in the tables provided.
** Classifer 1 - GaussianNB**
| Training Set Size | Training Time | Prediction Time (test) | F1 Score (train) | F1 Score (test) |
| :---------------: | :---------------------: | :--------------------: | :--------------: | :-------------: |
| 100 | 0.0060 seconds | 0.0020 seconds | 0.3218 | 0.2078 |
| 200 | 0.0060 seconds | 0.0030 seconds | 0.7925 | 0.7727 |
| 300 | 0.0060 seconds | 0.0030 seconds | 0.0761 | 0.7852 |
** Classifer 2 - Logistic Regression **
| Training Set Size | Training Time | Prediction Time (test) | F1 Score (train) | F1 Score (test) |
| :---------------: | :---------------------: | :--------------------: | :--------------: | :-------------: |
| 100 | 0.0090 seconds | 0.0000 seconds | 0.9114 | 0.7500 |
| 200 | 0.0190 seconds | 0.0000 seconds | 0.8293 | 0.7606 |
| 300 | 0.0320 seconds | 0.0000 seconds | 0.8435 | 0.8138 |
** Classifer 3 - Random Forest **
| Training Set Size | Training Time | Prediction Time (test) | F1 Score (train) | F1 Score (test) |
| :---------------: | :---------------------: | :--------------------: | :--------------: | :-------------: |
| 100 | 0.1890 seconds | 0.0010 seconds | 1.0000 | 0.7671 |
| 200 | 0.1300 seconds | 0.0160 seconds | 0.9924 | 0.6970 |
| 300 | 0.1730 seconds | 0.0000 seconds | 0.9875 | 0.7445 |
## Choosing the Best Model
In this final section, you will choose from the three supervised learning models the *best* model to use on the student data. You will then perform a grid search optimization for the model over the entire training set (`X_train` and `y_train`) by tuning at least one parameter to improve upon the untuned model's F<sub>1</sub> score.
### Question 3 - Choosing the Best Model
*Based on the experiments you performed earlier, in one to two paragraphs, explain to the board of supervisors what single model you chose as the best model. Which model is generally the most appropriate based on the available data, limited resources, cost, and performance?*
**Answer: ** I've decide to use Random Forest despite GaussianNB got better results. In my researches, GaussianNB doesn't have ways to improve the results with GridSearchCV. The Random Forest has a bigger training time, but it seems very fast in prediction time with high train and test F1 scores. Beyond that, RandomForestClassifier has many hyperparameters to tuning the algorithm.
### Question 4 - Model in Layman's Terms
*In one to two paragraphs, explain to the board of directors in layman's terms how the final model chosen is supposed to work. Be sure that you are describing the major qualities of the model, such as how the model is trained and how the model makes a prediction. Avoid using advanced mathematical or technical jargon, such as describing equations or discussing the algorithm implementation.*
**Answer: ** Random Forest is just a bunch of decision trees. Decision tree is an algorithm that make simple questions in order to categorize the data. For example, if a bank needs to decide whether to lend money to a customer, it will check its historical data and start to make a serie of questions. Based on answers the bank can conclude if it will lend money or not.
However trust in a single decision tree could be a weak decision. Then we used Random Forest, that I said before, it is a bunch of decision trees. The algorithm combine many trees to get a better result. The bunch of decision trees are generated randomly. In our case we have a dataset with 30 features. Not all features are used in each of decision trees. Let's suppose that you use 20% of features of each decision tree. That means the algorithm will select randomly 6 features for each tree. This technique guarantees to train the data in numerous different decision trees. With the results of this series of decision trees, the algorithm can decide the best classification for specific input. The decision is made by vote. For example: we run the algorithm with 100 decision trees. A specific student was classified as "passed" in 85 decision trees and "not passed" in 15 trees. In this case, the majority vote for this student is "passed" and that is the result of the algorithm.
### Implementation: Model Tuning
Fine tune the chosen model. Use grid search (`GridSearchCV`) with at least one important parameter tuned with at least 3 different values. You will need to use the entire training set for this. In the code cell below, you will need to implement the following:
- Import [`sklearn.grid_search.gridSearchCV`](http://scikit-learn.org/stable/modules/generated/sklearn.grid_search.GridSearchCV.html) and [`sklearn.metrics.make_scorer`](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.make_scorer.html).
- Create a dictionary of parameters you wish to tune for the chosen model.
- Example: `parameters = {'parameter' : [list of values]}`.
- Initialize the classifier you've chosen and store it in `clf`.
- Create the F<sub>1</sub> scoring function using `make_scorer` and store it in `f1_scorer`.
- Set the `pos_label` parameter to the correct value!
- Perform grid search on the classifier `clf` using `f1_scorer` as the scoring method, and store it in `grid_obj`.
- Fit the grid search object to the training data (`X_train`, `y_train`), and store it in `grid_obj`.
```
# TODO: Import 'GridSearchCV' and 'make_scorer'
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import f1_score, make_scorer
# TODO: Create the parameters list you wish to tune
parameters = {"n_estimators":[10, 50, 100],
"criterion":["gini", "entropy"]}
# TODO: Initialize the classifier
clf = RandomForestClassifier(random_state=1973)
# TODO: Make an f1 scoring function using 'make_scorer'
f1_scorer = make_scorer(f1_score, pos_label='yes')
# TODO: Perform grid search on the classifier using the f1_scorer as the scoring method
grid_obj = GridSearchCV(clf, param_grid=parameters, scoring=f1_scorer)
# TODO: Fit the grid search object to the training data and find the optimal parameters
grid_obj = grid_obj.fit(X_train, y_train)
print(grid_obj.best_params_)
# Report the final F1 score for training and testing after parameter tuning
print "Tuned model has a training F1 score of {:.4f}.".format(predict_labels(grid_obj, X_train, y_train))
print "Tuned model has a testing F1 score of {:.4f}.".format(predict_labels(grid_obj, X_test, y_test))
print ("--------------------------------------------------------------------------")
```
### Question 5 - Final F<sub>1</sub> Score
*What is the final model's F<sub>1</sub> score for training and testing? How does that score compare to the untuned model?*
**Answer: **
- Tuned model has a training F1 score of 1.0000.
- Tuned model has a testing F1 score of 0.8322.
The tuned Random Forest model had a good increase in performance comparing the F1 score. The original F1 test score for 300 points was 0.7445 and now it is 0.8322. The original F1 train score for 300 points was 0.9875 and now it is 1.0
The final parameters that GridSearchCV foud are:
- n_estimators: 50
- criterion: entropy
> **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to
**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
```
import seaborn as sns # Install using 'pip install seaborn'
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
%matplotlib inline
cm_test = confusion_matrix(y_test, grid_obj.predict(X_test))
plt.figure(figsize=(7,5))
sns.heatmap(cm_test, annot=True, cmap='Greys', xticklabels=['No', 'Yes'], yticklabels=['No', 'Yes'])
plt.title('Confusion Matrix for the Test Set')
plt.ylabel('True')
plt.xlabel('Predicted')
```
| github_jupyter |
# Machine Learning Part I
## 1. Introduction
### 1.1 What is Machine Learning?
[Machine learning](https://en.wikipedia.org/wiki/Machine_learning) was described by Arthur Samuel as the "field of study that gives computers the ability to learn without being explicitly programmed".
The following three principles are central to Machine Learning:
1. **a pattern exists**: machine learning will not work on data that is completely random.
2. **the pattern cannot be "pinned down" mathematically**: if a pattern can be pinned down mathematically (for example Newton's equations), it is probably more appropriate to use the original equations
3. **we have data on that pattern**: machine learning algorithms require relevant data, in order to search for patterns in that data.
The last point raises the following question - how much data is needed in order to learn?
### 1.2 The Feasibility of Learning - how much data do we need in order to learn?
Two related concepts can give us insight as to why we are able to learn from data, and how does having more data help us to learn--these are [Hoeffding's Inequality](https://en.wikipedia.org/wiki/Hoeffding%27s_inequality) and the [law of large numbers](https://en.wikipedia.org/wiki/Law_of_large_numbers).
#### 1.2.1 Hoeffding's Inequality
Consider an infinite bin, from which we take a sample of size $N$, which we find to have a sample frequency, $\nu$. However, the bin frequency, $\mu$, is unknown. However, **Hoeffding's Inequality** allows us to calculate a probability bound between these two quantities, i.e.:
$$\mathbb{P} \left[ \left| \nu - \mu \right| > \epsilon \right] \le 2e^{-2\epsilon^2 N}$$
This inequality applies to a *single bin*, and is valid for all $N$ and $\epsilon$ and the bound does not depend on $\mu$.
It illustrates the tradeoff between $N$, $\epsilon$, and the bound, i.e. the larger the sample size, $N$, the smaller our probability bound. On the other hand, the smaller the tolerance, $\epsilon$, the harder it will be to keep the probability small.
#### 1.2.2 The Law of Large Numbers
A related concept is the weak **law of large numbers**, given by:
$$\lim_{m\rightarrow\infty} \mathbb{P} \left[ \left| \underset{X \sim P}{\mathbb{E}}[\mathbf{X}] - \frac{1}{m}\sum\limits_{i=1}^m x_i \right| > \epsilon \right] = 0$$
As the sample size, $m$, approaches infinity, the probability approaches zero.
**Further Reading**:
1. [Machine Learning Theory - Part 1: Introduction](https://mostafa-samir.github.io/ml-theory-pt1/) | Mostafa Samir
2. [Machine Learning Theory - Part 2: Generalization Bounds](https://mostafa-samir.github.io/ml-theory-pt2/) | Mostafa Samir
### 1.3 Libraries used in this exercise
The following Python libraries are used in this exercise
```
from __future__ import print_function, division
import numpy as np
import matplotlib.pyplot as plt
import itertools
np.set_printoptions(edgeitems=3,infstr='inf',linewidth=75,nanstr='nan',precision=4,suppress=False,threshold=1000,formatter=None)
%matplotlib inline
```
## 2. Machine Learning Example
### 2.1 Learning Boolean Target Function
In this example, we intend to "learn" a Boolean target function. The function takes in input vector of size 3, say [0, 1, 0] or [0, 1, 1] and outputs a single output, $y_n$, which can be zero or 1. If we enumerate the set of all possible target functions, we would have $2^{2^3}=256$ distinct Boolean functions on 3 Boolean inputs.
In this example, we wish to score 4 different hypotheses, $g_a, g_b, g_c, g_d$ based on how well they perform on "out of sample points". The scoring system is 1 point per correct point, i.e. if the function gets 3 "out of sample points" correct, it will get a score of 3. We wish to calcuate the total score when enumerating over the entire set of all possible functions.
In the code below, the 256 distinct Boolean functions are enumerated using `itertools.product([0,1],repeat=8)` and a score is calculated for each of 4 possible cases. It is determined that the score for each of these cases is identical.
```
def calculate_score_last3(y,g):
total_score=0
for y_last in y[:,5:]:
total_score += np.sum(y_last==g)
return total_score
g_a = np.vstack(np.array([1,1,1,1,1]))
g_b = np.vstack(np.array([0,0,0,0,0]))
g_c = np.vstack(np.array([0,1,1,0,1]))
g_d = np.logical_not(g_a).astype(int)
y = np.array(list(itertools.product([0,1],repeat=8)))
print("Scores:\n(a) {}\n(b) {}\n(c) {}\n(d) {}".format(
calculate_score_last3(y,[1,1,1]),
calculate_score_last3(y,[0,0,0]),
calculate_score_last3(y,[0,0,1]),
calculate_score_last3(y,[1,1,0])))
```
## 3. The Perceptron
### 3.1 Introduction & Background
The [Perceptron](https://en.wikipedia.org/wiki/Perceptron) is attributed to the work of Frank Rosenblatt in 1957. It is a binary classifier that works on the basis of whether a dot product, $\mathbf{w \cdot x}$, exceeds a certain threshold:
$$f(\mathbf{x}) = \left\{ \begin{array}{cl} 1, & \text{if } \mathbf{w \cdot x} > 0\\0, & \text{otherwise}
\end{array}\right.$$
This can be written in the form of a hypothesis given by:
$$h(\mathbf{x}) = \text{sign}(\mathbf{w^T x}) = \text{sign}\left(\sum_{i=1}^N w_i x_i\right)$$
### 3.2 Generating Sample Data
To generate the sample data for the perceptron classifier above, a three-column random matrix with the first column as ones is created as follows:
```
def generate_data(n,seed=None):
if seed is not None:
np.random.seed(seed)
x0 = np.ones(n)
x1 = np.random.uniform(low=-1,high=1,size=(2,n))
return np.vstack((x0,x1)).T
```
### 3.3 Creating a random line inside the region of interest
The region of interest is $\mathcal{X}=\left[-1,1\right] \times \left[-1,1\right]$,
where $\times$ denotes the [Cartesian Product](https://en.wikipedia.org/wiki/Cartesian_product).
A random line is created, and to ensure that it falls within the region of interest, it is created from two random points, $(x_0,y_0)$ and $(x_1,y_1)$ which are generated within $\mathcal{X}$. The equation for this line in *slope-intercept* form and in the *hypothesis / weights* form are derived as follows.
#### 3.3.1 Equation of the random line in the slope-intercept form, $y = mx + c$
$$\text{Slope, }m = \frac{y_1 - y_0}{x_1 - x_0}$$
$$\text{Intercept, }c = y_0 - m x_0$$
#### 3.3.2 Equation of the random line in hypothesis [weights] form
The hypothesis, in the two dimensional case ($N=2$), and since $x_0=1$:
$$h(\mathbf{x}) = \text{sign}(w_0 + w_1 x_1 + w_2 x_2)$$
The decision boundary, $h(\mathbf{x})=0$ is given by:
$$w_0 + w_1 x_1 + w_2 x_2 = 0$$
This can be converted to the slope-intercept form if we take $x_1$ to be the $x$ axis, and $x_2$ to be the $y$ axis, i.e.:
$$w_0 + w_1 x + w_2 y = 0$$
$$y = - \frac{w_1}{w_2} x - \frac{w_0}{w_2}$$
Comparison with the equation $y = mx + c$ yields the following relationships:
$$m = - \frac{w_1}{w_2}$$
$$c = - \frac{w_0}{w_2}$$
If we arbitrarily set $w_2=1$, we arrive at the following set of weights, which is consistent with the decision boundary denoted by $y = mx + c$:
$$\mathbf{w} = \left(-c,-m,1\right)$$
```
def get_random_line(seed=None):
X = generate_data(2,seed=seed)
x = X[:,1]
y = X[:,2]
m = (y[1]-y[0])/(x[1]-x[0])
c = y[0] - m*x[0]
return np.array([-c,-m,1])
def draw_line(ax,w,marker='g--',label=None):
m = -w[1]/w[2]
c = -w[0]/w[2]
x = np.linspace(-1,1,20)
y = m*x + c
if label is None:
ax.plot(x,y,marker)
else:
ax.plot(x,y,marker,label=label)
def get_hypothesis(X,w):
h=np.dot(X,w)
return np.sign(h).astype(int)
```
### 3.4 Generating a Training Dataset
The following code generates a training dataset (separated into positive and negative classes according to a random line in $\mathcal{X}$) and plots it.
```
def plot_data(fig,plot_id,X,y=None,w_arr=None,my_x=None,title=None):
ax = fig.add_subplot(plot_id)
if y is None:
ax.plot(X[:,1],X[:,2],'gx')
else:
ax.plot(X[y > 0,1],X[y > 0,2],'b+',label='Positive (+)')
ax.plot(X[y < 0,1],X[y < 0,2],'ro',label='Negative (-)')
ax.set_xlim(-1,1)
ax.set_ylim(-1,1)
ax.grid(True)
if w_arr is not None:
if isinstance(w_arr,list) is not True:
w_arr=[w_arr]
for i,w in enumerate(w_arr):
if i==0:
draw_line(ax,w,'g-',label='Theoretical')
else:
draw_line(ax,w,'g--')
if my_x is not None:
ax.plot([my_x[0]],[my_x[1]],'kx',markersize=10)
if title is not None:
ax.set_title(title)
ax.legend(loc='best',frameon=True)
def create_dataset(N,make_plot=True,seed=None):
X = generate_data(N,seed=seed)
w_theoretical = get_random_line()
y = get_hypothesis(X,w_theoretical)
if make_plot is True:
fig = plt.figure(figsize=(7,5))
plot_data(fig,111,X,y,w_theoretical,title="Initial Dataset")
return X,y,w_theoretical
```
### 3.5 The Perceptron Learning Algorithm
The Preceptron Learning Algorithm (PLA) is implemented in the following steps
1. Calculate $h\left(\mathbf{x}\right)=\text{sign}\left(\mathbf{w^T x}\right)$ which can take on values of -1, 0, or 1 for each sample.
2. Compare $h\left(\mathbf{x}\right)$ with $y$ to find misclassified point(s) if any.
3. Pick one misclassified point at random.
4. Iterate the weights according to the PLA: $w = w + y_n x_n$, where $y_n$ is the correct classification for the misclassified point, and $x_n$ is the misclassified point.
The code below also keeps track of the weights, misclassification error, and misclassified point at each iteration.
```
def PLA(w,X,y0,n_iterations=10,verbose=True):
assert len(y0)==X.shape[0]
n=len(y0)
x_arr = list()
w_arr = list()
m_arr = list()
for i in range(n_iterations):
h = get_hypothesis(X,w)
bad = h != y0
bad = np.argwhere(bad).flatten()
if len(bad) > 0:
idx = np.random.choice(bad,1)[0]
my_x = X[idx,:]
m_arr.append(100.0*len(bad)/n)
x_arr.append(my_x)
w_arr.append(np.copy(w))
w += np.dot(y0[idx],my_x)
if verbose is True:
print("iter {}: {}% misclassified, w={}" \
.format(i,m_arr[-1],w_arr[-1]))
else:
m_arr.append(0.0)
x_arr.append(np.array([1.0, np.nan,np.nan]))
w_arr.append(np.copy(w))
if verbose is True:
print("iter {}: zero misclassified (PLA has converged)".format(i))
return w,w_arr,m_arr,x_arr
print("PLA failed to converge after {} iterations".format(i))
return None,None,None,None
```
### 3.6 Implementing the PLA
Here, we generate a sample dataset of 10 points and plot it. The perceptron learning algorithm, starting from an initial weight of $(0,0,0)$, converges in less than 10 iterations.
```
X,y,w_theoretical = create_dataset(N=10,make_plot=True,seed=247)
w0 = np.array([0,0,0],dtype=float)
w,w_arr,m_arr,x_arr = PLA(w0,X,y,n_iterations=100,verbose=True)
def draw_plot_steps(fig,plot_id,X,y,w_theoretical,w_arr,x_arr,idx_arr):
assert len(idx_arr) <= 9
for idx in idx_arr:
print("w_arr[{}] = {}, x_arr[{}] = {}".format(idx,w_arr[idx],idx,x_arr[idx][1:]))
plot_data(fig,plot_id,X,y,[w_theoretical] + [w_arr[idx]],
x_arr[idx][1:],title="iteration {}".format(idx))
plot_id += 1
fig = plt.figure(figsize=(10,15))
draw_plot_steps(fig,421,X,y,w_theoretical,w_arr,x_arr,np.arange(len(w_arr)-1)+1)
def plot_convergence(m_arr):
fig = plt.figure(figsize=(7,5))
ax = fig.add_subplot(111)
ax.plot(m_arr,'g+-',markersize=10)
ax.grid(True)
ax.set_title("Convergence")
ax.set_xlabel("Iterations")
ax.set_ylabel("Misclassification Error (%)")
plot_convergence(m_arr)
```
### 3.7 Number of iterations required for convergence
The following code calculates the number of iterations required for convergence and plots its distribution.
```
def plot_histogram(my_count,bins=200,x_max=80):
fig = plt.figure(figsize=(7,5))
ax = fig.add_subplot(111)
ax.hist(my_count,bins=bins);
ax.set_xlim(0,x_max)
ax.grid(True)
def get_iteration_distribution(N,n_trials=1000,max_iterations=10000,summary=True):
n_iterations=np.zeros(n_trials,dtype=int)
w0 = np.array([0,0,0],dtype=float)
for i in range(n_trials):
X,y,w_theoretical = create_dataset(N=N,make_plot=False,seed=None)
w,w_arr,m_arr,x_arr = PLA(w0,X,y,n_iterations=max_iterations,verbose=False)
n_iterations[i]=len(w_arr)
if summary is True:
print("Minumum iterations: {}".format(np.min(n_iterations)))
print("Maximum iterations: {}".format(np.max(n_iterations)))
print("Mean iterations: {}".format(np.mean(n_iterations)))
return n_iterations
n_iterations = get_iteration_distribution(N=10,n_trials=1000)
plot_histogram(n_iterations,bins=200,x_max=50)
```
### 3.8 Calculate the misclassification error for the converged weights
If we know the theoretical decision boundary, `w_theoretical`, that 'knows' the correct classification of points, we can calculate the number of points misclassified by `w` (the final weights after convergence using the PLA) via random sampling. The misclassification error is slightly less than 20%.
```
def calculate_misclassification(w_theoretical,w,n_samples=1000,verbose=True):
X = generate_data(n_samples,seed=None)
y0 = get_hypothesis(X,w_theoretical)
y = get_hypothesis(X,w)
n_correct = np.sum(y == y0)
if verbose is True:
if w_theoretical[0] != 0.0:
print("Theoretical Weights : {}".format(w_theoretical/w_theoretical[0]))
else:
print("Theoretical Weights : {}".format(w_theoretical))
print("PLA Predicted Weights: {}".format(w))
print("Correct points = {}".format(n_correct))
print("Incorrect points = {}".format(n_samples-n_correct))
print("Misclassification= {}%".format(np.round(100 * (n_samples-n_correct)/n_samples, 4)))
fig = plt.figure(figsize=(7,5))
plot_data(fig,111,X,y0,[w_theoretical,w])
return (n_samples-n_correct)/n_samples
misclassification = calculate_misclassification(w_theoretical,w,verbose=True)
misclassification
def plot_misclassification(my_count,bins=20,x_max=0.25):
fig = plt.figure(figsize=(7,5))
ax = fig.add_subplot(111)
ax.hist(my_count,bins=bins);
ax.set_xlim(0,x_max)
ax.grid(True)
def get_misclassification_distribution(N=10,n_trials=1000,max_iterations=10000,summary=True):
w0 = np.array([0,0,0],dtype=float)
misclassification=np.zeros(n_trials)
for i in range(n_trials):
X,y,w_theoretical = create_dataset(N,make_plot=False,seed=None)
w,w_arr,m_arr,x_arr = PLA(w0,X,y,n_iterations=max_iterations,verbose=False)
misclassification[i]=calculate_misclassification(w_theoretical,w,n_samples=1000,verbose=False)
if summary is True:
print("Minumum misclassification: {}".format(np.min(misclassification)))
print("Maximum misclassification: {}".format(np.max(misclassification)))
print("Mean misclassification: {}".format(np.mean(misclassification)))
return misclassification
```
### 3.9 Iteration distribution and misclassification distribution for N=10
Here, we find that for $N$=10, an average of about 10 iterations is required for convergence. The average misclassification error is about 10%.
```
n_iterations = get_iteration_distribution(N=10,n_trials=1000)
plot_histogram(n_iterations,bins=100,x_max=40)
misclassification = get_misclassification_distribution(N=10)
plot_misclassification(misclassification,bins=20,x_max=0.4)
```
### 3.10 Iteration distribution and misclassification distribution for N=100
For $N$=100, an average of about 80~100 iterations is required for convergence. The average misclassification error is slightly over ~1%.
```
n_iterations = get_iteration_distribution(N=100,n_trials=1000)
plot_histogram(n_iterations,bins=300,x_max=300)
misclassification = get_misclassification_distribution(N=100)
plot_misclassification(misclassification,bins=40,x_max=0.05)
```
### 3.11 Convergence Plot for N=100
```
X,y,w_theoretical = create_dataset(N=100,make_plot=True,seed=12345)
w0 = np.array([0,0,0],dtype=float)
w,w_arr,m_arr,x_arr = PLA(w0,X,y,n_iterations=1000,verbose=False)
plot_convergence(m_arr)
fig = plt.figure(figsize=(8,7))
plot_data(fig,111,X,y,[w_theoretical] + w_arr)
```
| github_jupyter |
# CStreet: a computed <ins>C</ins>ell <ins>S</ins>tate <ins>tr</ins>ajectory inf<ins>e</ins>r<ins>e</ins>nce method for <ins>t</ins>ime-series single-cell RNA-seq data
## This is a tutorial written using Jupyter Notebook.
### Step 1. CStreet installation following the [tutorial](https://github.com/yw-Hua/CStreet).
### Step 2. Input preparation
CStreet utilizes time-series expression levels in tab-delimited format or AnnData format as input.
The cell state information can be generated using the built-in clustering function of CStreet or input by the user.
We provided a small [test dataset](https://github.com/yw-Hua/CStreet/tree/master/test/test_data) containing normalized expression levels and the state information at three time points.
### Step 3. Operation of CStreet
### 3.1 Load required packages
```
from cstreet import *
import pandas as pd
```
### 3.2 Read expression matrix and cell state information
Expression data: Expression matrix containing the time-series expression level as reads counts or normalized values in tab delimited format, and anndata format are accepted as the input of CStreet. (For example: ExpressionMatrix_t1.txt ExpressionMatrix_t2.txt ExpressionMatrix_t3.txt)
```
data_t1=pd.read_table('ExpressionMatrix_t1.txt',header=0, sep="\t",index_col=0)
data_t2=pd.read_table('ExpressionMatrix_t2.txt',header=0, sep="\t",index_col=0)
data_t3=pd.read_table('ExpressionMatrix_t3.txt',header=0, sep="\t",index_col=0)
```
Cell states info: The cell states information can be inputted by the user or generated using the internal clustering function of CStreet. (For example: CellStates_t1.txt CellStates_t2.txt CellStates_t3.txt)
```
state_t1=pd.read_table('CellStates_t1.txt',header=None, sep="\t",index_col=0)
state_t2=pd.read_table('CellStates_t2.txt',header=None, sep="\t",index_col=0)
state_t3=pd.read_table('CellStates_t3.txt',header=None, sep="\t",index_col=0)
```
### 3.3 Create a new CStreet object
```
cdata=CStreetData()
```
### 3.4 Add data into CStreet object
```
cdata.add_new_timepoint_scdata(data_t1,list(state_t1[1]))
cdata.add_new_timepoint_scdata(data_t2,list(state_t2[1]))
cdata.add_new_timepoint_scdata(data_t3,list(state_t3[1]))
```
AnnData object can be inputted as well.
```
#cdata.timepoint_scdata_dict[1]=anndata_t1
#cdata.timepoint_scdata_dict[2]=anndata_t2
#cdata.timepoint_scdata_dict[3]=anndata_t3
cdata.timepoint_scdata_dict[1]
```
### 3.5 Custom parameters
```
cdata.params
cdata.params.FigureParam_LabelBoxWidth=8
```
### 3.6 Run CStreet
#### 3.6.1 step by step
```
cdata.cell_clusters(CellClusterParam_PCAn=10,CellClusterParam_k=15,CellClusterParam_Resolution=1,Switch_Normalize=True,Switch_LogTransform=True)
cdata.filter_dead_cell(Threshold_MitoPercent=0.2)
cdata.filter_lowcell_gene(Threshold_LowCellNum=3)
cdata.filter_lowgene_cells(Threshold_LowGeneNum=200)
cdata.normalize_data(Threshold_NormalizeBase=1000000)
cdata.log_transform()
cdata.get_knn_within(WithinTimePointParam_PCAn=10,WithinTimePointParam_k=15)
cdata.get_knn_between(BetweenTimePointParam_PCAn=10,BetweenTimePointParam_k=15)
cdata.get_knn_graph()
cdata.filter_knn_graph()
cdata.get_state_trajectory(ProbParam_SamplingSize=5,ProbParam_RandomSeed=0)
cdata.get_knn_nxG(Threshold_MaxOutDegree=10,Threshold_MinCellNumofStates=1)
cdata.draw_nxG(FigureParam_FigureSize=(6,7),FigureParam_LabelBoxWidth=8)
cdata.output_results()
```
#### 3.6.2 one step
```
cdata=CStreetData()
state_t1=pd.read_table('CellStates_t1.txt',header=None, sep="\t",index_col=0)
state_t2=pd.read_table('CellStates_t2.txt',header=None, sep="\t",index_col=0)
state_t3=pd.read_table('CellStates_t3.txt',header=None, sep="\t",index_col=0)
cdata.run_cstreet()
```
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import dataframe_image as dfi
import matplotlib.ticker
import numpy as np
import pandas as pd
import seaborn as sns
%matplotlib inline
sns.set()
matplotlib.rcParams['figure.figsize'] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
```
### COVID-19 Cases
```
confirmed_df = pd.read_csv("https://covid19tracking.narrativa.com/csv/confirmed.csv")
radar_covid_countries = {"Spain"}
# radar_covid_regions = { ... }
confirmed_df = confirmed_df[confirmed_df["Country_EN"].isin(radar_covid_countries)]
# confirmed_df = confirmed_df[confirmed_df["Region"].isin(radar_covid_regions)]
# set(confirmed_df.Region.tolist()) == radar_covid_regions
confirmed_country_columns = list(filter(lambda x: x.startswith("Country_"), confirmed_df.columns))
confirmed_regional_columns = confirmed_country_columns + ["Region"]
confirmed_df.drop(columns=confirmed_regional_columns, inplace=True)
confirmed_df = confirmed_df.sum().to_frame()
confirmed_df.tail()
confirmed_df.reset_index(inplace=True)
confirmed_df.columns = ["sample_date_string", "cumulative_cases"]
confirmed_df.sort_values("sample_date_string", inplace=True)
confirmed_df["new_cases"] = confirmed_df.cumulative_cases.diff()
confirmed_df["rolling_mean_new_cases"] = confirmed_df.new_cases.rolling(7).mean()
confirmed_df.tail()
extraction_date_confirmed_df = \
confirmed_df[confirmed_df.sample_date_string == extraction_date]
extraction_previous_date_confirmed_df = \
confirmed_df[confirmed_df.sample_date_string == extraction_previous_date].copy()
if extraction_date_confirmed_df.empty and \
not extraction_previous_date_confirmed_df.empty:
extraction_previous_date_confirmed_df["sample_date_string"] = extraction_date
extraction_previous_date_confirmed_df["new_cases"] = \
extraction_previous_date_confirmed_df.rolling_mean_new_cases
extraction_previous_date_confirmed_df["cumulative_cases"] = \
extraction_previous_date_confirmed_df.new_cases + \
extraction_previous_date_confirmed_df.cumulative_cases
confirmed_df = confirmed_df.append(extraction_previous_date_confirmed_df)
confirmed_df.tail()
confirmed_df[["new_cases", "rolling_mean_new_cases"]].plot()
```
### Extract API TEKs
```
from Modules.RadarCOVID import radar_covid
exposure_keys_df = radar_covid.download_last_radar_covid_exposure_keys(days=14)
exposure_keys_df[[
"sample_date_string", "source_url", "region", "key_data"]].head()
exposure_keys_summary_df = \
exposure_keys_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "tek_count"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = exposure_keys_df[["sample_date_string", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
"sample_date").tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
"Data/TEKs/Current/RadarCOVID-TEKs.json",
lines=True, orient="records")
tek_list_df.drop(columns=["extraction_date_with_hour"]).to_json(
"Data/TEKs/Daily/RadarCOVID-TEKs-" + extraction_date + ".json",
lines=True, orient="records")
tek_list_df.to_json(
"Data/TEKs/Hourly/RadarCOVID-TEKs-" + extraction_date_with_hour + ".json",
lines=True, orient="records")
tek_list_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame()
paths = list(reversed(sorted(glob.glob(f"Data/TEKs/{mode}/RadarCOVID-TEKs-*.json"))))
if limit:
paths = paths[:limit]
for path in paths:
logging.info(f"Loading TEKs from '{path}'...")
iteration_extracted_teks_df = pd.read_json(path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
return extracted_teks_df
```
### Daily New TEKs
```
daily_extracted_teks_df = load_extracted_teks(mode="Daily", limit=14)
daily_extracted_teks_df.head()
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "new_tek_count",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.head()
new_tek_devices_df = daily_extracted_teks_df.copy()
new_tek_devices_df["new_sample_extraction_date"] = \
pd.to_datetime(new_tek_devices_df.sample_date) + datetime.timedelta(1)
new_tek_devices_df["extraction_date"] = pd.to_datetime(new_tek_devices_df.extraction_date)
new_tek_devices_df = new_tek_devices_df[
new_tek_devices_df.new_sample_extraction_date == new_tek_devices_df.extraction_date]
new_tek_devices_df.head()
new_tek_devices_df.set_index("extraction_date", inplace=True)
new_tek_devices_df = new_tek_devices_df.tek_list.apply(lambda x: len(set(x))).to_frame()
new_tek_devices_df.reset_index(inplace=True)
new_tek_devices_df.rename(columns={
"extraction_date": "sample_date_string",
"tek_list": "new_tek_devices"}, inplace=True)
new_tek_devices_df["sample_date_string"] = new_tek_devices_df.sample_date_string.dt.strftime("%Y-%m-%d")
new_tek_devices_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(mode="Hourly", limit=24)
hourly_extracted_teks_df.head()
hourly_tek_list_df = hourly_extracted_teks_df.groupby("extraction_date_with_hour").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
hourly_tek_list_df = hourly_tek_list_df.set_index("extraction_date_with_hour").sort_index(ascending=True)
hourly_new_tek_df = hourly_tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
hourly_new_tek_df.rename(columns={
"tek_list": "new_tek_count"}, inplace=True)
hourly_new_tek_df.tail()
hourly_new_tek_devices_df = hourly_extracted_teks_df.copy()
hourly_new_tek_devices_df["new_sample_extraction_date"] = \
pd.to_datetime(hourly_new_tek_devices_df.sample_date) + datetime.timedelta(1)
hourly_new_tek_devices_df["extraction_date"] = pd.to_datetime(hourly_new_tek_devices_df.extraction_date)
hourly_new_tek_devices_df = hourly_new_tek_devices_df[
hourly_new_tek_devices_df.new_sample_extraction_date == hourly_new_tek_devices_df.extraction_date]
hourly_new_tek_devices_df.set_index("extraction_date_with_hour", inplace=True)
hourly_new_tek_devices_df_ = pd.DataFrame()
for i, chunk_df in hourly_new_tek_devices_df.groupby("extraction_date"):
chunk_df = chunk_df.copy()
chunk_df.sort_index(inplace=True)
chunk_tek_count_df = chunk_df.tek_list.apply(lambda x: len(set(x)))
chunk_df = chunk_tek_count_df.diff().fillna(chunk_tek_count_df).to_frame()
hourly_new_tek_devices_df_ = hourly_new_tek_devices_df_.append(chunk_df)
hourly_new_tek_devices_df = hourly_new_tek_devices_df_
hourly_new_tek_devices_df.reset_index(inplace=True)
hourly_new_tek_devices_df.rename(columns={
"tek_list": "new_tek_devices"}, inplace=True)
hourly_new_tek_devices_df.tail()
hourly_summary_df = hourly_new_tek_df.merge(
hourly_new_tek_devices_df, on=["extraction_date_with_hour"], how="outer")
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df.tail()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(new_tek_devices_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(confirmed_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["tek_count_per_new_case"] = \
result_summary_df.tek_count / result_summary_df.rolling_mean_new_cases
result_summary_df["new_tek_count_per_new_case"] = \
result_summary_df.new_tek_count / result_summary_df.rolling_mean_new_cases
result_summary_df["new_tek_devices_per_new_case"] = \
result_summary_df.new_tek_devices / result_summary_df.rolling_mean_new_cases
result_summary_df["new_tek_count_per_new_tek_device"] = \
result_summary_df.new_tek_count / result_summary_df.new_tek_devices
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
result_summary_df.set_index("sample_date", inplace=True)
result_summary_df = result_summary_df.sort_index(ascending=False)
```
## Report Results
### Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[[
"tek_count",
"new_tek_count",
"new_cases",
"rolling_mean_new_cases",
"tek_count_per_new_case",
"new_tek_count_per_new_case",
"new_tek_devices",
"new_tek_devices_per_new_case",
"new_tek_count_per_new_tek_device"]]
result_summary_df
```
### Summary Plots
```
summary_ax_list = result_summary_df[[
"rolling_mean_new_cases",
"tek_count",
"new_tek_count",
"new_tek_devices",
"new_tek_count_per_new_tek_device",
"new_tek_devices_per_new_case"
]].sort_index(ascending=True).plot.bar(
title="Summary", rot=45, subplots=True, figsize=(15, 22))
summary_ax_list[-1].yaxis.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df.plot.bar(
title="Last 24h Summary", rot=45, subplots=True)
```
### Publish Results
```
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
media_path = get_temporary_image_path()
dfi.export(df, media_path)
return media_path
summary_plots_image_path = save_temporary_plot_image(ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(df=result_summary_df)
hourly_summary_plots_image_path = save_temporary_plot_image(ax=hourly_summary_ax_list)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(report_resources_path_prefix + "Summary-Table.html")
_ = shutil.copyfile(summary_plots_image_path, report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(summary_table_image_path, report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(hourly_summary_plots_image_path, report_resources_path_prefix + "Hourly-Summary-Plots.png")
report_daily_url_pattern = \
"https://github.com/pvieito/RadarCOVID-Report/blob/master/Notebooks/" \
"RadarCOVID-Report/{report_type}/RadarCOVID-Report-{report_date}.ipynb"
report_daily_url = report_daily_url_pattern.format(
report_type="Daily", report_date=extraction_date)
report_hourly_url = report_daily_url_pattern.format(
report_type="Hourly", report_date=extraction_date_with_hour)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
summary_table_html = result_summary_df.to_html()
readme_contents = readme_contents.format(
summary_table_html=summary_table_html,
report_url_with_hour=report_hourly_url,
extraction_date_with_hour=extraction_date_with_hour)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule":
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
hourly_summary_plots_media = api.media_upload(hourly_summary_plots_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
hourly_summary_plots_media.media_id,
]
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
new_teks = extraction_date_result_summary_df.new_tek_count.sum().astype(int)
new_teks_last_hour = extraction_date_result_hourly_summary_df.new_tek_count.sum().astype(int)
new_devices = extraction_date_result_summary_df.new_tek_devices.sum().astype(int)
new_devices_last_hour = extraction_date_result_hourly_summary_df.new_tek_devices.sum().astype(int)
new_tek_count_per_new_tek_device = \
extraction_date_result_summary_df.new_tek_count_per_new_tek_device.sum()
new_tek_devices_per_new_case = \
extraction_date_result_summary_df.new_tek_devices_per_new_case.sum()
status = textwrap.dedent(f"""
Report Update – {extraction_date_with_hour}
#ExposureNotification #RadarCOVID
Shared Diagnoses Day Summary:
- New TEKs: {new_teks} ({new_teks_last_hour:+d} last hour)
- New Devices: {new_devices} ({new_devices_last_hour:+d} last hour, {new_tek_count_per_new_tek_device:.2} TEKs/device)
- Usage Ratio: {new_tek_devices_per_new_case:.2%} devices/case
Report Link: {report_hourly_url}
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
# D&D Name generator
```
%matplotlib inline
import os
import sys
PROJECT_ROOT = os.path.dirname(os.getcwd())
sys.path.append(PROJECT_ROOT)
from collections import defaultdict
import numpy as np
import matplotlib.pyplot as plt
from data import DnDCharacterNameDataset
from train import RNNLayerTrainer
from generator import RNNLayerGenerator
from utils import read_log
```
## Dataset
Basic information about dataset
```
# Read the dataset
dataset = DnDCharacterNameDataset(os.path.join(PROJECT_ROOT, "data"))
# Print sample from dataset
train, target = dataset[4]
print("Name: {}".format(train['name']))
print("Gender: {}".format(train['gender'])) # List genders and races are just convinient way of storing information per
print("Race: {}".format(train['race'])) # letter for later transformation to indices
print("Target letters: {}".format(target))
```
**How many names per race?**
```
counter = defaultdict(int)
for train, target in dataset:
race = train['race'][0]
counter[race] += 1
print(counter)
x = np.arange(len(counter))
labels, y = zip(*counter.items())
plt.bar(x, y, tick_label=labels)
plt.ylabel("Count")
plt.show()
```
**How mana names with same first later?**
```
counter = defaultdict(int)
for train, target in dataset:
first_letter = train['name'][0]
counter[first_letter] += 1
x = np.arange(len(counter))
labels, y = zip(*sorted(counter.items()))
print(counter)
plt.bar(x, y, tick_label=labels)
plt.ylabel("Count")
plt.show()
```
## Training
Training configuration
```
epochs = 300
batch_size = 128
hidden_size = 128
learning_rate = 0.0001
device = "cuda"
logfile = "train_loss.log"
verbose = 0 # Removing outputs during training
```
Initialize trainer
```
trainer = RNNLayerTrainer(os.path.join(PROJECT_ROOT, "data"),
epochs=epochs,
batch_size=batch_size,
hidden_size=hidden_size,
lr=learning_rate,
device=device,
logfile=logfile,
verbose=verbose)
```
Run training loop for number of `epochs` and log losses. Models will be saved on 25 epoch intervals.
```
trainer.run_train_loop()
```
Plot losses during training
```
epochs, losses = read_log(os.path.join(PROJECT_ROOT, "train_loss.log"))
assert len(epochs) == len(losses), "Mismatch lengths. {} != {}".format(len(epochs), len(losses))
plt.plot(np.arange(len(losses)), losses)
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.show()
```
## Evaluation
* Examination of name quality over epochs
* Comparision between generated names and real examples
Create helper function that is going to find most similar name in dataset
```
import distance
def most_similar_name(name):
"""Find most similar name in dataset based on Levenshtein distance"""
min_score = 1
similar_name = None
for train, _ in dataset:
n = "".join(train['name'])
score = distance.levenshtein(n, name, normalized=True)
if score < min_score:
min_score = score
similar_name = n
return similar_name, min_score
```
To validate how good generated names are, let us focus on only one specific letter/race/gender combination. This will make tracking changes across epochs easier. In this case, we'll sample an **Elven**, **Female** character name with start letter __B__. Since the model is trained on 300 epochs, let's take models at epochs **15**, **50**, **100**, **200**, **300**.
```
model_nums = [15, 50, 100, 200, 300] # Take models from these epochs
for model_num in model_nums:
model_path = os.path.join(PROJECT_ROOT, "models", "rnn_layer_epoch_{}.pt".format(model_num))
generator = RNNLayerGenerator(model_path)
letter, race, gender = "B", "elf", "female"
name = generator.sample(letter, race, gender)
similar_name, score = most_similar_name(name)
print("Samples from model on epoch {}".format(model_num))
print("Generated name: {}".format(name))
print("Most similar name: {} with score {:.2f}\n".format(similar_name, score))
```
**Comments**
To give a better perspective, these are the only Female Elf names that start with letter B:
Bethrynna
Birel
Baelitae
Bethrynna
From the previous example, you can see that the model is not trying to reproduce exact names but actually combine with others, for example, Bival (generated name) vs Mival (real name). Which doesn't sound so bad... Of course, there is some repetition with Bethrynna, but there is always a chance for that to happen.
Interestingly, it's hard to determine from these examples which of the models is actually the best here... I guess a deeper investigation on how the probabilities are distributed when sampling new letter might give more insight. Did we just gut lucky with randomness or some letters have much higher probability?
| github_jupyter |
```
import json
import time
import sys
import pandas as pd
import numpy as np
import scipy.sparse as sp
import pickle as pkl
import collections
import gc
with open('../data/index_item_map.pkl', 'rb') as f:
data_map = pkl.load(f)
paper_id_title = data_map['paper_id_title']
author_id_name = data_map['author_id_name']
venue_id_name = data_map['venue_id_name']
keywords_id_name = data_map['keywords_id_name']
paper_title_id = data_map['paper_title_id']
author_name_id = data_map['author_name_id']
venue_name_id = data_map['venue_name_id']
keywords_name_id = data_map['keywords_name_id']
keywords_set = data_map['keywords_set']
venue_set = data_map['venue_set']
with open ('../data/cumulative_labels.pkl','rb') as f:
cumulative_labels = pkl.load(f)
full_pair = pd.read_csv('../data/whole_list.txt',names=['P','O','type'])
def construct_index_from_one_df(df,item_id,flag):
if flag == 'P':
max_len = 100
elif flag == 'A':
max_len = 20
elif flag == 'V':
max_len = 1
elif flag == 'K':
max_len = 15
dic = collections.defaultdict(lambda : [])
keys = list(df['P'])
values = list(df['O'])
for i in range(len(keys)):
try:
if len(dic[keys[i]])<max_len:
index_id = item_id[values[i]]
dic[keys[i]].append(index_id)
cnt = cnt + 1
except:
pass
for i in range(len(keys)):
need_len = max_len-len(dic[keys[i]])
for j in range(need_len):
dic[keys[i]].append(-1)
return dic
def construct_index_5_years(df,item_list,flag):
dic_1 = construct_index_from_one_df(df,item_list[0],flag)
dic_2 = construct_index_from_one_df(df,item_list[1],flag)
dic_3 = construct_index_from_one_df(df,item_list[2],flag)
dic_4 = construct_index_from_one_df(df,item_list[3],flag)
dic_5 = construct_index_from_one_df(df,item_list[4],flag)
return [dic_1,dic_2,dic_3,dic_4,dic_5]
def index_mapping_one(paper_lst,id_dic):
return [id_dic[paper] for paper in paper_lst]
def index_mapping_five(paper_lst,id_dic_list):
return np.array([index_mapping_one(paper_lst,id_dic) for id_dic in id_dic_list])
with open('../individual_data/graph_2000.pkl','rb') as f:
graph_2000 = pkl.load(f)
item_id_2000 = graph_2000['item_id']
with open('../individual_data/graph_2001.pkl','rb') as f:
graph_2001 = pkl.load(f)
item_id_2001 = graph_2001['item_id']
with open('../individual_data/graph_2002.pkl','rb') as f:
graph_2002 = pkl.load(f)
item_id_2002 = graph_2002['item_id']
with open('../individual_data/graph_2003.pkl','rb') as f:
graph_2003 = pkl.load(f)
item_id_2003 = graph_2003['item_id']
with open('../individual_data/graph_2004.pkl','rb') as f:
graph_2004 = pkl.load(f)
item_id_2004 = graph_2004['item_id']
t = time.time()
P1Y_2005 = full_pair[full_pair.O==2005]
all_paper_2005 = pd.DataFrame({"P":list(set(P1Y_2005['P']))})
full_pair_2005 = pd.merge(all_paper_2005,full_pair,how="left")
full_pair_2005_P1P = full_pair_2005[full_pair_2005.type=='P1P']
full_pair_2005_P1A = full_pair_2005[full_pair_2005.type=='P1A']
full_pair_2005_P1V = full_pair_2005[full_pair_2005.type=='P1V']
full_pair_2005_P1K = full_pair_2005[full_pair_2005.type=='P1K']
graph_lookip_list = [item_id_2000,item_id_2001,item_id_2002,item_id_2003,item_id_2004]
P2005_1_P1P_list = construct_index_5_years(full_pair_2005_P1P,graph_lookip_list,'P')
P2005_1_P1A_list = construct_index_5_years(full_pair_2005_P1A,graph_lookip_list,'A')
P2005_1_P1V_list = construct_index_5_years(full_pair_2005_P1V,graph_lookip_list,'V')
P2005_1_P1K_list = construct_index_5_years(full_pair_2005_P1K,graph_lookip_list,'K')
P2005_label_cumu = list(cumulative_labels['P2005_label']['P'])
P2005_1_P1P_list = index_mapping_five(P2005_label_cumu,P2005_1_P1P_list)
P2005_1_P1A_list = index_mapping_five(P2005_label_cumu,P2005_1_P1A_list)
P2005_1_P1V_list = index_mapping_five(P2005_label_cumu,P2005_1_P1V_list)
P2005_1_P1K_list = index_mapping_five(P2005_label_cumu,P2005_1_P1K_list)
print ((P2005_1_P1P_list).shape)
print ((P2005_1_P1A_list).shape)
print ((P2005_1_P1V_list).shape)
print ((P2005_1_P1K_list).shape)
with open('../individual_data/index_2005.pkl','wb') as f:
pkl.dump([P2005_1_P1P_list,P2005_1_P1A_list,P2005_1_P1V_list,P2005_1_P1K_list],f,0)
time.time()-t
with open('../individual_data/graph_2005.pkl','rb') as f:
graph_2005 = pkl.load(f)
item_id_2005 = graph_2005['item_id']
t = time.time()
P1Y_2006 = full_pair[full_pair.O==2006]
all_paper_2006 = pd.DataFrame({"P":list(set(P1Y_2006['P']))})
full_pair_2006 = pd.merge(all_paper_2006,full_pair,how="left")
full_pair_2006_P1P = full_pair_2006[full_pair_2006.type=='P1P']
full_pair_2006_P1A = full_pair_2006[full_pair_2006.type=='P1A']
full_pair_2006_P1V = full_pair_2006[full_pair_2006.type=='P1V']
full_pair_2006_P1K = full_pair_2006[full_pair_2006.type=='P1K']
graph_lookip_list = [item_id_2001,item_id_2002,item_id_2003,item_id_2004,item_id_2005]
P2006_1_P1P_list = construct_index_5_years(full_pair_2006_P1P,graph_lookip_list,'P')
P2006_1_P1A_list = construct_index_5_years(full_pair_2006_P1A,graph_lookip_list,'A')
P2006_1_P1V_list = construct_index_5_years(full_pair_2006_P1V,graph_lookip_list,'V')
P2006_1_P1K_list = construct_index_5_years(full_pair_2006_P1K,graph_lookip_list,'K')
P2006_label_cumu = list(cumulative_labels['P2006_label']['P'])
P2006_1_P1P_list = index_mapping_five(P2006_label_cumu,P2006_1_P1P_list)
P2006_1_P1A_list = index_mapping_five(P2006_label_cumu,P2006_1_P1A_list)
P2006_1_P1V_list = index_mapping_five(P2006_label_cumu,P2006_1_P1V_list)
P2006_1_P1K_list = index_mapping_five(P2006_label_cumu,P2006_1_P1K_list)
print ((P2006_1_P1P_list).shape)
print ((P2006_1_P1A_list).shape)
print ((P2006_1_P1V_list).shape)
print ((P2006_1_P1K_list).shape)
with open('../individual_data/index_2006.pkl','wb') as f:
pkl.dump([P2006_1_P1P_list,P2006_1_P1A_list,P2006_1_P1V_list,P2006_1_P1K_list],f,0)
time.time()-t
with open('../individual_data/graph_2006.pkl','rb') as f:
graph_2006 = pkl.load(f)
item_id_2006 = graph_2006['item_id']
t = time.time()
P1Y_2007 = full_pair[full_pair.O==2007]
all_paper_2007 = pd.DataFrame({"P":list(set(P1Y_2007['P']))})
full_pair_2007 = pd.merge(all_paper_2007,full_pair,how="left")
full_pair_2007_P1P = full_pair_2007[full_pair_2007.type=='P1P']
full_pair_2007_P1A = full_pair_2007[full_pair_2007.type=='P1A']
full_pair_2007_P1V = full_pair_2007[full_pair_2007.type=='P1V']
full_pair_2007_P1K = full_pair_2007[full_pair_2007.type=='P1K']
graph_lookip_list = [item_id_2002,item_id_2003,item_id_2004,item_id_2005,item_id_2006]
P2007_1_P1P_list = construct_index_5_years(full_pair_2007_P1P,graph_lookip_list,'P')
P2007_1_P1A_list = construct_index_5_years(full_pair_2007_P1A,graph_lookip_list,'A')
P2007_1_P1V_list = construct_index_5_years(full_pair_2007_P1V,graph_lookip_list,'V')
P2007_1_P1K_list = construct_index_5_years(full_pair_2007_P1K,graph_lookip_list,'K')
P2007_label_cumu = list(cumulative_labels['P2007_label']['P'])
P2007_1_P1P_list = index_mapping_five(P2007_label_cumu,P2007_1_P1P_list)
P2007_1_P1A_list = index_mapping_five(P2007_label_cumu,P2007_1_P1A_list)
P2007_1_P1V_list = index_mapping_five(P2007_label_cumu,P2007_1_P1V_list)
P2007_1_P1K_list = index_mapping_five(P2007_label_cumu,P2007_1_P1K_list)
print ((P2007_1_P1P_list).shape)
print ((P2007_1_P1A_list).shape)
print ((P2007_1_P1V_list).shape)
print ((P2007_1_P1K_list).shape)
with open('../individual_data/index_2007.pkl','wb') as f:
pkl.dump([P2007_1_P1P_list,P2007_1_P1A_list,P2007_1_P1V_list,P2007_1_P1K_list],f,0)
time.time()-t
with open('../individual_data/graph_2007.pkl','rb') as f:
graph_2007 = pkl.load(f)
item_id_2007 = graph_2007['item_id']
t = time.time()
P1Y_2008 = full_pair[full_pair.O==2008]
all_paper_2008 = pd.DataFrame({"P":list(set(P1Y_2008['P']))})
full_pair_2008 = pd.merge(all_paper_2008,full_pair,how="left")
full_pair_2008_P1P = full_pair_2008[full_pair_2008.type=='P1P']
full_pair_2008_P1A = full_pair_2008[full_pair_2008.type=='P1A']
full_pair_2008_P1V = full_pair_2008[full_pair_2008.type=='P1V']
full_pair_2008_P1K = full_pair_2008[full_pair_2008.type=='P1K']
graph_lookip_list = [item_id_2003,item_id_2004,item_id_2005,item_id_2006,item_id_2007]
P2008_1_P1P_list = construct_index_5_years(full_pair_2008_P1P,graph_lookip_list,'P')
P2008_1_P1A_list = construct_index_5_years(full_pair_2008_P1A,graph_lookip_list,'A')
P2008_1_P1V_list = construct_index_5_years(full_pair_2008_P1V,graph_lookip_list,'V')
P2008_1_P1K_list = construct_index_5_years(full_pair_2008_P1K,graph_lookip_list,'K')
P2008_label_cumu = list(cumulative_labels['P2008_label']['P'])
P2008_1_P1P_list = index_mapping_five(P2008_label_cumu,P2008_1_P1P_list)
P2008_1_P1A_list = index_mapping_five(P2008_label_cumu,P2008_1_P1A_list)
P2008_1_P1V_list = index_mapping_five(P2008_label_cumu,P2008_1_P1V_list)
P2008_1_P1K_list = index_mapping_five(P2008_label_cumu,P2008_1_P1K_list)
print ((P2008_1_P1P_list).shape)
print ((P2008_1_P1A_list).shape)
print ((P2008_1_P1V_list).shape)
print ((P2008_1_P1K_list).shape)
with open('../individual_data/index_2008.pkl','wb') as f:
pkl.dump([P2008_1_P1P_list,P2008_1_P1A_list,P2008_1_P1V_list,P2008_1_P1K_list],f,0)
time.time()-t
with open('../individual_data/graph_2008.pkl','rb') as f:
graph_2008 = pkl.load(f)
item_id_2008 = graph_2008['item_id']
t = time.time()
P1Y_2009 = full_pair[full_pair.O==2009]
all_paper_2009 = pd.DataFrame({"P":list(set(P1Y_2009['P']))})
full_pair_2009 = pd.merge(all_paper_2009,full_pair,how="left")
full_pair_2009_P1P = full_pair_2009[full_pair_2009.type=='P1P']
full_pair_2009_P1A = full_pair_2009[full_pair_2009.type=='P1A']
full_pair_2009_P1V = full_pair_2009[full_pair_2009.type=='P1V']
full_pair_2009_P1K = full_pair_2009[full_pair_2009.type=='P1K']
graph_lookip_list = [item_id_2004,item_id_2005,item_id_2006,item_id_2007,item_id_2008]
P2009_1_P1P_list = construct_index_5_years(full_pair_2009_P1P,graph_lookip_list,'P')
P2009_1_P1A_list = construct_index_5_years(full_pair_2009_P1A,graph_lookip_list,'A')
P2009_1_P1V_list = construct_index_5_years(full_pair_2009_P1V,graph_lookip_list,'V')
P2009_1_P1K_list = construct_index_5_years(full_pair_2009_P1K,graph_lookip_list,'K')
P2009_label_cumu = list(cumulative_labels['P2009_label']['P'])
P2009_1_P1P_list = index_mapping_five(P2009_label_cumu,P2009_1_P1P_list)
P2009_1_P1A_list = index_mapping_five(P2009_label_cumu,P2009_1_P1A_list)
P2009_1_P1V_list = index_mapping_five(P2009_label_cumu,P2009_1_P1V_list)
P2009_1_P1K_list = index_mapping_five(P2009_label_cumu,P2009_1_P1K_list)
print ((P2009_1_P1P_list).shape)
print ((P2009_1_P1A_list).shape)
print ((P2009_1_P1V_list).shape)
print ((P2009_1_P1K_list).shape)
with open('../individual_data/index_2009.pkl','wb') as f:
pkl.dump([P2009_1_P1P_list,P2009_1_P1A_list,P2009_1_P1V_list,P2009_1_P1K_list],f,0)
time.time()-t
with open('../individual_data/graph_2009.pkl','rb') as f:
graph_2009 = pkl.load(f)
item_id_2009 = graph_2009['item_id']
t = time.time()
P1Y_2010 = full_pair[full_pair.O==2010]
all_paper_2010 = pd.DataFrame({"P":list(set(P1Y_2010['P']))})
full_pair_2010 = pd.merge(all_paper_2010,full_pair,how="left")
full_pair_2010_P1P = full_pair_2010[full_pair_2010.type=='P1P']
full_pair_2010_P1A = full_pair_2010[full_pair_2010.type=='P1A']
full_pair_2010_P1V = full_pair_2010[full_pair_2010.type=='P1V']
full_pair_2010_P1K = full_pair_2010[full_pair_2010.type=='P1K']
graph_lookip_list = [item_id_2005,item_id_2006,item_id_2007,item_id_2008,item_id_2009]
P2010_1_P1P_list = construct_index_5_years(full_pair_2010_P1P,graph_lookip_list,'P')
P2010_1_P1A_list = construct_index_5_years(full_pair_2010_P1A,graph_lookip_list,'A')
P2010_1_P1V_list = construct_index_5_years(full_pair_2010_P1V,graph_lookip_list,'V')
P2010_1_P1K_list = construct_index_5_years(full_pair_2010_P1K,graph_lookip_list,'K')
P2010_label_cumu = list(cumulative_labels['P2010_label']['P'])
P2010_1_P1P_list = index_mapping_five(P2010_label_cumu,P2010_1_P1P_list)
P2010_1_P1A_list = index_mapping_five(P2010_label_cumu,P2010_1_P1A_list)
P2010_1_P1V_list = index_mapping_five(P2010_label_cumu,P2010_1_P1V_list)
P2010_1_P1K_list = index_mapping_five(P2010_label_cumu,P2010_1_P1K_list)
print ((P2010_1_P1P_list).shape)
print ((P2010_1_P1A_list).shape)
print ((P2010_1_P1V_list).shape)
print ((P2010_1_P1K_list).shape)
with open('../individual_data/index_2010.pkl','wb') as f:
pkl.dump([P2010_1_P1P_list,P2010_1_P1A_list,P2010_1_P1V_list,P2010_1_P1K_list],f,0)
time.time()-t
with open('../individual_data/graph_2010.pkl','rb') as f:
graph_2010 = pkl.load(f)
item_id_2010 = graph_2010['item_id']
t = time.time()
P1Y_2011 = full_pair[full_pair.O==2011]
all_paper_2011 = pd.DataFrame({"P":list(set(P1Y_2011['P']))})
full_pair_2011 = pd.merge(all_paper_2011,full_pair,how="left")
full_pair_2011_P1P = full_pair_2011[full_pair_2011.type=='P1P']
full_pair_2011_P1A = full_pair_2011[full_pair_2011.type=='P1A']
full_pair_2011_P1V = full_pair_2011[full_pair_2011.type=='P1V']
full_pair_2011_P1K = full_pair_2011[full_pair_2011.type=='P1K']
graph_lookip_list = [item_id_2006,item_id_2007,item_id_2008,item_id_2009,item_id_2010]
P2011_1_P1P_list = construct_index_5_years(full_pair_2011_P1P,graph_lookip_list,'P')
P2011_1_P1A_list = construct_index_5_years(full_pair_2011_P1A,graph_lookip_list,'A')
P2011_1_P1V_list = construct_index_5_years(full_pair_2011_P1V,graph_lookip_list,'V')
P2011_1_P1K_list = construct_index_5_years(full_pair_2011_P1K,graph_lookip_list,'K')
P2011_label_cumu = list(cumulative_labels['P2011_label']['P'])
P2011_1_P1P_list = index_mapping_five(P2011_label_cumu,P2011_1_P1P_list)
P2011_1_P1A_list = index_mapping_five(P2011_label_cumu,P2011_1_P1A_list)
P2011_1_P1V_list = index_mapping_five(P2011_label_cumu,P2011_1_P1V_list)
P2011_1_P1K_list = index_mapping_five(P2011_label_cumu,P2011_1_P1K_list)
print ((P2011_1_P1P_list).shape)
print ((P2011_1_P1A_list).shape)
print ((P2011_1_P1V_list).shape)
print ((P2011_1_P1K_list).shape)
with open('../individual_data/index_2011.pkl','wb') as f:
pkl.dump([P2011_1_P1P_list,P2011_1_P1A_list,P2011_1_P1V_list,P2011_1_P1K_list],f,0)
time.time()-t
with open('../data/graph_2011.pkl','rb') as f:
graph_2011 = pkl.load(f)
item_id_2011 = graph_2011['item_id']
t = time.time()
P1Y_2012 = full_pair[full_pair.O==2012]
all_paper_2012 = pd.DataFrame({"P":list(set(P1Y_2012['P']))})
full_pair_2012 = pd.merge(all_paper_2012,full_pair,how="left")
full_pair_2012_P1P = full_pair_2012[full_pair_2012.type=='P1P']
full_pair_2012_P1A = full_pair_2012[full_pair_2012.type=='P1A']
full_pair_2012_P1V = full_pair_2012[full_pair_2012.type=='P1V']
full_pair_2012_P1K = full_pair_2012[full_pair_2012.type=='P1K']
graph_lookip_list = [item_id_2007,item_id_2008,item_id_2009,item_id_2010,item_id_2011]
P2012_1_P1P_list = construct_index_5_years(full_pair_2012_P1P,graph_lookip_list,'P')
P2012_1_P1A_list = construct_index_5_years(full_pair_2012_P1A,graph_lookip_list,'A')
P2012_1_P1V_list = construct_index_5_years(full_pair_2012_P1V,graph_lookip_list,'V')
P2012_1_P1K_list = construct_index_5_years(full_pair_2012_P1K,graph_lookip_list,'K')
P2012_label_cumu = list(cumulative_labels['P2012_label']['P'])
P2012_1_P1P_list = index_mapping_five(P2012_label_cumu,P2012_1_P1P_list)
P2012_1_P1A_list = index_mapping_five(P2012_label_cumu,P2012_1_P1A_list)
P2012_1_P1V_list = index_mapping_five(P2012_label_cumu,P2012_1_P1V_list)
P2012_1_P1K_list = index_mapping_five(P2012_label_cumu,P2012_1_P1K_list)
print ((P2012_1_P1P_list).shape)
print ((P2012_1_P1A_list).shape)
print ((P2012_1_P1V_list).shape)
print ((P2012_1_P1K_list).shape)
with open('../data/index_2012.pkl','wb') as f:
pkl.dump([P2012_1_P1P_list,P2012_1_P1A_list,P2012_1_P1V_list,P2012_1_P1K_list],f,0)
time.time()-t
with open('../data/graph_2012.pkl','rb') as f:
graph_2012 = pkl.load(f)
item_id_2012 = graph_2012['item_id']
t = time.time()
P1Y_2013 = full_pair[full_pair.O==2013]
all_paper_2013 = pd.DataFrame({"P":list(set(P1Y_2013['P']))})
full_pair_2013 = pd.merge(all_paper_2013,full_pair,how="left")
full_pair_2013_P1P = full_pair_2013[full_pair_2013.type=='P1P']
full_pair_2013_P1A = full_pair_2013[full_pair_2013.type=='P1A']
full_pair_2013_P1V = full_pair_2013[full_pair_2013.type=='P1V']
full_pair_2013_P1K = full_pair_2013[full_pair_2013.type=='P1K']
graph_lookip_list = [item_id_2008,item_id_2009,item_id_2010,item_id_2011,item_id_2012]
P2013_1_P1P_list = construct_index_5_years(full_pair_2013_P1P,graph_lookip_list,'P')
P2013_1_P1A_list = construct_index_5_years(full_pair_2013_P1A,graph_lookip_list,'A')
P2013_1_P1V_list = construct_index_5_years(full_pair_2013_P1V,graph_lookip_list,'V')
P2013_1_P1K_list = construct_index_5_years(full_pair_2013_P1K,graph_lookip_list,'K')
P2013_label_cumu = list(cumulative_labels['P2013_label']['P'])
P2013_1_P1P_list = index_mapping_five(P2013_label_cumu,P2013_1_P1P_list)
P2013_1_P1A_list = index_mapping_five(P2013_label_cumu,P2013_1_P1A_list)
P2013_1_P1V_list = index_mapping_five(P2013_label_cumu,P2013_1_P1V_list)
P2013_1_P1K_list = index_mapping_five(P2013_label_cumu,P2013_1_P1K_list)
print ((P2013_1_P1P_list).shape)
print ((P2013_1_P1A_list).shape)
print ((P2013_1_P1V_list).shape)
print ((P2013_1_P1K_list).shape)
with open('../data/index_2013.pkl','wb') as f:
pkl.dump([P2013_1_P1P_list,P2013_1_P1A_list,P2013_1_P1V_list,P2013_1_P1K_list],f,0)
time.time()-t
```
| github_jupyter |
```
"""
Convert netCDF files to geotiff.
Update 2019 06 26: rerun using updated inundation layers
Author: Rutger Hofste
Date: 20180816
Kernel: python35
Docker: rutgerhofste/gisdocker:ubuntu16.04
Args:
TESTING (boolean) : Toggle testing mode
SCRIPT_NAME (string) : Script name
OUTPUT_VERSION (integer) : output version for ec2 and s3.
GCS_OUTPUT_PATH (string) : Output path for Google Cloud Storage.
Returns:
Result:
Geotiff and pickled dictionaries in ec2_output folder.
"""
TESTING = 0
SCRIPT_NAME = "Y2018M08D16_RH_Convertt_Geotiff_V01"
OUTPUT_VERSION = 5
EC2_INPUT_PATH = "/volumes/data/Y2018M08D08_RH_S3_EC2_V01/output_V02/"
GCS_OUTPUT_PATH = "gs://aqueduct30_v01/{}/output_V{:02.0f}/".format(SCRIPT_NAME,OUTPUT_VERSION)
ec2_output_path = "/volumes/data/{}/output_V{:02.0f}/".format(SCRIPT_NAME,OUTPUT_VERSION)
s3_output_path = "s3://wri-projects/Aqueduct30/processData/{}/output_V{:02.0f}/".format(SCRIPT_NAME,OUTPUT_VERSION)
print("EC2_INPUT_PATH: " + EC2_INPUT_PATH +
"\nec2_output_path: " + ec2_output_path,
"\ns3_output_path: " + s3_output_path,
"\nGCS_OUTPUT_PATH:" + GCS_OUTPUT_PATH)
import time, datetime, sys
dateString = time.strftime("Y%YM%mD%d")
timeString = time.strftime("UTC %H:%M")
start = datetime.datetime.now()
print(dateString,timeString)
sys.version
!rm -r {ec2_output_path}
!mkdir -p {ec2_output_path}
import os
import pandas as pd
import netCDF4
import pickle
import multiprocessing as mp
import numpy as np
try:
from osgeo import ogr, osr, gdal
except:
sys.exit('ERROR: cannot find GDAL/OGR modules')
if 'GDAL_DATA' not in os.environ:
os.environ['GDAL_DATA'] = r'/usr/share/gdal/2.1'
def filename_to_dict(filename):
values = filename.split("_")
value_length = len(values)
if value_length == 5:
keys = ["floodtype","climate","model","year","returnperiod"]
dictje = dict(zip(keys,values))
elif value_length == 6:
keys = ["floodtype","climate","subsidence","year","returnperiod","returnperiod_decimal"]
dictje = dict(zip(keys,values))
elif value_length == 8:
keys = ["floodtype","climate","subsidence","year","returnperiod","returnperiod_decimal","model","sea_level_rise_scenario"]
dictje = dict(zip(keys,values))
else:
print("error")
return dictje
def ncdump(nc_fid):
'''ncdump outputs dimensions, variables and their attribute information.
-------------------------------------------------------------------------------
The information is similar to that of NCAR's ncdump utility.
ncdump requires a valid instance of Dataset.
Args:
nc_fid (netCDF4.Dataset) : A netCDF4 dateset object
Returns:
nc_attrs (list) : A Python list of the NetCDF file global attributes.
nc_dims (list) : A Python list of the NetCDF file dimensions.
nc_vars (list) : A Python list of the NetCDF file variables.
'''
nc_attrs = nc_fid.ncattrs()
nc_dims = [dim for dim in nc_fid.dimensions] # list of nc dimensions
nc_vars = [var for var in nc_fid.variables] # list of nc variables
return nc_attrs, nc_dims, nc_vars
def get_global_attributes(dictje):
""" Get global attributes from netcdf
Args:
dictionary with root, filename and properties.
"""
input_path = os.path.join(dictje["root"],dictje["filename"])
nc_fid = netCDF4.Dataset(input_path, 'r')
nc_attrs, nc_dims, nc_vars = ncdump(nc_fid)
global_attributes_dict = {}
for nc_attr in nc_attrs:
global_attributes_dict[nc_attr] = nc_fid.getncattr(nc_attr)
return global_attributes_dict
def get_variable_attributes(dictje):
""" Get global attributes from netcdf
Args:
dictionary with root, filename and properties.
"""
input_path = os.path.join(dictje["root"],dictje["filename"])
nc_fid = netCDF4.Dataset(input_path, 'r')
nc_attrs, nc_dims, nc_vars = ncdump(nc_fid)
parameter = nc_vars[-1] #warning, project dependent
variable_attrs = nc_fid.variables[parameter].ncattrs()
variable_attributes_dict = {}
for variable_attr in variable_attrs:
variable_attributes_dict[parameter+"_"+variable_attr] = nc_fid.variables[parameter].getncattr(variable_attr)
return variable_attributes_dict
def write_geotiff(output_path,geotransform,geoprojection,data,nodata_value=-9999,datatype=gdal.GDT_Float32):
""" Write data to geotiff file
-------------------------------------------------------------------------------
Args:
output_path (string) : output_path
geotransform (tuple) : geotransform
geoprojection (string) : geoprojection in osr format
data (np.array) : numpy array
nodata_value (integer) : NoData value
datatype (GDAL datatype)
"""
(x,y) = data.shape
format = "GTiff"
driver = gdal.GetDriverByName(format)
# you can change the dataformat but be sure to be able to store negative values including -9999
dst_ds = driver.Create(output_path,y,x,1,datatype, [ 'COMPRESS=LZW' ])
dst_ds.GetRasterBand(1).SetNoDataValue(nodata_value)
dst_ds.GetRasterBand(1).WriteArray(data)
dst_ds.SetGeoTransform(geotransform)
dst_ds.SetProjection(geoprojection)
dst_ds = None
return 1
def get_global_georeference(array):
""" Get the geotransform and projection for a numpy array
-------------------------------------------------------------------------------
Returns a geotransform and projection for a global extent in epsg 4326
projection.
Args:
array (np.array) : numpy array
Returns:
geotransform (tuple) : geotransform
geoprojection (string) : geoprojection in osr format
"""
y_dimension = array.shape[0] #rows, lat
x_dimension = array.shape[1] #cols, lon
geotransform = (-180,360.0/x_dimension,0,90,0,-180.0/y_dimension)
srs = osr.SpatialReference()
srs.ImportFromEPSG(4326)
geoprojection = srs.ExportToWkt()
if len(geoprojection) == 0:
warnings.warn("GDAL_DATA path not set correctly. Assert os.environ " \
"contains GDAL_DATA \n" \
"Code will execute without projection set")
return geotransform, geoprojection
def standardize_time(time_unit,times):
""" Append standardize time to list
-------------------------------------------------------------------------------
The netCDF results of the university of Utrecht consist of multiple time
formats.
Args:
time_unit (string) : units as provided by the netCDF4 file.
times (list) : list of time in units provided in time_units (e.g. days).
Returns:
standardized_time (list) : list of normalized times in datetime format.
"""
standardized_time =[]
for time in times:
if time_unit == ("days since 1900-01-01 00:00:00") or (time_unit =="Days since 1900-01-01"):
standardized_time.append(datetime.datetime(1900,1,1) + datetime.timedelta(days=time))
elif time_unit == "days since 1901-01-01 00:00:00" or time_unit == "Days since 1901-01-01":
standardized_time.append(datetime.datetime(1901,1,1) + datetime.timedelta(days=time))
elif time_unit == "Days since 1960-01-01 00:00:00":
standardized_time.append(datetime.datetime(1960,1,1) + datetime.timedelta(days=time))
else:
raise("Error, unknown format:",time_unit)
standardized_time.append(-9999)
return standardized_time
def convert_netcdf_geotiff(dictje):
""" Convert netcdf to geotiff
Args:
dictionary with root, filename and properties.
"""
input_path = os.path.join(dictje["root"],dictje["filename"])
nc_fid = netCDF4.Dataset(input_path, 'r')
nc_attrs, nc_dims, nc_vars = ncdump(nc_fid)
parameter = nc_vars[-1]
lats = nc_fid.variables['lat'][:] # extract/copy the data
lons = nc_fid.variables['lon'][:]
times = nc_fid.variables['time'][:]
time_unit = nc_fid.variables["time"].getncattr("units")
standardized_time = standardize_time(time_unit,times)
i = 0 # single time step
Z = nc_fid.variables[parameter][i, :, :]
Z[Z<-9990]= -9999
Z[Z>1e19] = -9999
Z = np.flipud(Z) #depending on NetCDF type.
base_filename, extension = dictje["filename"].split(".")
output_filename = base_filename + ".tif"
output_path_geotiff = os.path.join(ec2_output_path,output_filename)
geotransform, geoprojection = get_global_georeference(Z)
write_geotiff(output_path_geotiff,geotransform,geoprojection,Z,nodata_value=-9999,datatype=gdal.GDT_Float32)
return standardized_time
def pickle_dictionary(dictje):
base_filename, extension = dictje["filename"].split(".")
output_filename = base_filename + ".pickle"
output_path_pickle = os.path.join(ec2_output_path,output_filename)
with open(output_path_pickle, 'wb') as handle:
pickle.dump(dictje, handle, protocol=pickle.HIGHEST_PROTOCOL)
ID = 0
master_dict = {}
for root, dirs, files in os.walk(EC2_INPUT_PATH):
for one_file in files:
if one_file.endswith("nc"):
file_dict = {}
file_dict["root"] = root
file_dict["filename"] = one_file
filename , extension = one_file.split(".")
file_dict["properties_from_filename"] = filename_to_dict(filename)
master_dict[ID] = file_dict
ID += 1
# number of files that need to be converted
print(len(master_dict.keys()))
for ID, dictje in master_dict.items():
master_dict[ID]["global_attributes"] = get_global_attributes(dictje)
master_dict[ID]["variable_attributes"] = get_variable_attributes(dictje)
def process_items(dictje):
try:
convert_netcdf_geotiff(dictje)
pickle_dictionary(dictje)
print(dictje["filename"])
except:
print("error",dictje["filename"])
if TESTING:
n = 10
master_dict = {k: master_dict[k] for k in list(master_dict)[:n]}
cpu_count = mp.cpu_count()
p= mp.Pool(3) # Memory issues, limiting processes to 3, using appr 60-80% memory
processed_values= p.map( process_items, master_dict.values())
p.close()
p.join()
!gsutil -m cp -r {ec2_output_path} {GCS_OUTPUT_PATH}
!aws s3 cp --recursive {ec2_output_path} {s3_output_path}
end = datetime.datetime.now()
elapsed = end - start
print(elapsed)
```
Previous runs:
2:27:01.588096
| github_jupyter |
# Temporal Difference: On-policy n-Tuple Sarsa, Stochastic
```
import numpy as np
```
## Create environment
```
def create_environment_states():
"""Creates environment states.
Returns:
num_states: int, number of states.
num_terminal_states: int, number of terminal states.
num_non_terminal_states: int, number of non terminal states.
"""
num_states = 16
num_terminal_states = 2
num_non_terminal_states = num_states - num_terminal_states
return num_states, num_terminal_states, num_non_terminal_states
def create_environment_actions(num_non_terminal_states):
"""Creates environment actions.
Args:
num_non_terminal_states: int, number of non terminal states.
Returns:
max_num_actions: int, max number of actions possible.
num_actions_per_non_terminal_state: array[int], number of actions per
non terminal state.
"""
max_num_actions = 4
num_actions_per_non_terminal_state = np.repeat(
a=max_num_actions, repeats=num_non_terminal_states)
return max_num_actions, num_actions_per_non_terminal_state
def create_environment_successor_counts(num_states, max_num_actions):
"""Creates environment successor counts.
Args:
num_states: int, number of states.
max_num_actions: int, max number of actions possible.
Returns:
num_state_action_successor_states: array[int], number of successor
states s' that can be reached from state s by taking action a.
"""
num_state_action_successor_states = np.repeat(
a=1, repeats=num_states * max_num_actions)
num_state_action_successor_states = np.reshape(
a=num_state_action_successor_states,
newshape=(num_states, max_num_actions))
return num_state_action_successor_states
def create_environment_successor_arrays(
num_non_terminal_states, max_num_actions):
"""Creates environment successor arrays.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
Returns:
sp_idx: array[int], state indices of new state s' of taking action a
from state s.
p: array[float], transition probability to go from state s to s' by
taking action a.
r: array[float], reward from new state s' from state s by taking
action a.
"""
sp_idx = np.array(
object=[1, 0, 14, 4,
2, 1, 0, 5,
2, 2, 1, 6,
4, 14, 3, 7,
5, 0, 3, 8,
6, 1, 4, 9,
6, 2, 5, 10,
8, 3, 7, 11,
9, 4, 7, 12,
10, 5, 8, 13,
10, 6, 9, 15,
12, 7, 11, 11,
13, 8, 11, 12,
15, 9, 12, 13],
dtype=np.int64)
p = np.repeat(
a=1.0, repeats=num_non_terminal_states * max_num_actions * 1)
r = np.repeat(
a=-1.0, repeats=num_non_terminal_states * max_num_actions * 1)
sp_idx = np.reshape(
a=sp_idx,
newshape=(num_non_terminal_states, max_num_actions, 1))
p = np.reshape(
a=p,
newshape=(num_non_terminal_states, max_num_actions, 1))
r = np.reshape(
a=r,
newshape=(num_non_terminal_states, max_num_actions, 1))
return sp_idx, p, r
def create_environment():
"""Creates environment.
Returns:
num_states: int, number of states.
num_terminal_states: int, number of terminal states.
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
num_actions_per_non_terminal_state: array[int], number of actions per
non terminal state.
num_state_action_successor_states: array[int], number of successor
states s' that can be reached from state s by taking action a.
sp_idx: array[int], state indices of new state s' of taking action a
from state s.
p: array[float], transition probability to go from state s to s' by
taking action a.
r: array[float], reward from new state s' from state s by taking
action a.
"""
(num_states,
num_terminal_states,
num_non_terminal_states) = create_environment_states()
(max_num_actions,
num_actions_per_non_terminal_state) = create_environment_actions(
num_non_terminal_states)
num_state_action_successor_states = create_environment_successor_counts(
num_states, max_num_actions)
(sp_idx,
p,
r) = create_environment_successor_arrays(
num_non_terminal_states, max_num_actions)
return (num_states,
num_terminal_states,
num_non_terminal_states,
max_num_actions,
num_actions_per_non_terminal_state,
num_state_action_successor_states,
sp_idx,
p,
r)
```
## Set hyperparameters
```
def set_hyperparameters():
"""Sets hyperparameters.
Returns:
num_episodes: int, number of episodes to train over.
maximum_episode_length: int, max number of timesteps for an episode.
num_qs: int, number of state-action-value functions Q_i(s, a).
alpha: float, alpha > 0, learning rate.
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
gamma: float, 0 <= gamma <= 1, amount to discount future reward.
"""
num_episodes = 10000
maximum_episode_length = 200
num_qs = 3
alpha = 0.1
epsilon = 0.1
gamma = 1.0
return num_episodes, maximum_episode_length, num_qs, alpha, epsilon, gamma
```
## Create value function and policy arrays
```
def create_value_function_arrays(num_qs, num_states, max_num_actions):
"""Creates value function arrays.
Args:
num_qs: int, number of state-action-value functions Q_i(s, a).
num_states: int, number of states.
max_num_actions: int, max number of actions possible.
Returns:
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
"""
q = np.repeat(a=0.0, repeats=num_qs * num_states * max_num_actions)
q = np.reshape(a=q, newshape=(num_qs, num_states, max_num_actions))
return q
def create_policy_arrays(num_non_terminal_states, max_num_actions):
"""Creates policy arrays.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
Returns:
policy: array[float], learned stochastic policy of which
action a to take in state s.
"""
policy = np.repeat(
a=1.0 / max_num_actions,
repeats=num_non_terminal_states * max_num_actions)
policy = np.reshape(
a=policy,
newshape=(num_non_terminal_states, max_num_actions))
return policy
```
## Create algorithm
```
# Set random seed so that everything is reproducible
np.random.seed(seed=0)
def initialize_epsiode(
num_non_terminal_states, max_num_actions, q, epsilon, policy):
"""Initializes epsiode with initial state, initial action, and policy.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
policy: array[float], learned stochastic policy of which
action a to take in state s.
Returns:
init_s_idx: int, initial state index from set of non terminal states.
init_a_idx: int, initial action index from set of actions of state
init_s_idx.
policy: array[float], learned stochastic policy of which
action a to take in state s.
"""
# Randomly choose an initial state from all non-terminal states
init_s_idx = np.random.randint(
low=0, high=num_non_terminal_states, dtype=np.int64)
# Choose policy for chosen state by epsilon-greedy choosing from the
# state-action-value function
policy = epsilon_greedy_policy_from_state_action_function(
max_num_actions, q, epsilon, init_s_idx, policy)
# Get initial action
init_a_idx = np.random.choice(
a=max_num_actions, p=policy[init_s_idx, :])
return init_s_idx, init_a_idx, policy
def epsilon_greedy_policy_from_state_action_function(
max_num_actions, q, epsilon, s_idx, policy):
"""Create epsilon-greedy policy from state-action value function.
Args:
max_num_actions: int, max number of actions possible.
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
s_idx: int, current state index.
policy: array[float], learned stochastic policy of which action a to
take in state s.
Returns:
policy: array[float], learned stochastic policy of which action a to
take in state s.
"""
# Combine state-action value functions
q = np.sum(a=q[:, s_idx, :], axis=0)
# Save max state-action value and find the number of actions that have the
# same max state-action value
max_action_value = np.max(a=q)
max_action_count = np.count_nonzero(a=q == max_action_value)
# Apportion policy probability across ties equally for state-action pairs
# that have the same value and zero otherwise
if max_action_count == max_num_actions:
max_policy_prob_per_action = 1.0 / max_action_count
remain_prob_per_action = 0.0
else:
max_policy_prob_per_action = (1.0 - epsilon) / max_action_count
remain_prob_per_action = epsilon / (max_num_actions - max_action_count)
policy[s_idx, :] = np.where(
q == max_action_value,
max_policy_prob_per_action,
remain_prob_per_action)
return policy
def loop_through_episode(
num_non_terminal_states,
max_num_actions,
num_state_action_successor_states,
sp_idx,
p,
r,
num_qs,
q,
policy,
alpha,
epsilon,
gamma,
maximum_episode_length,
s_idx,
a_idx):
"""Loops through episode to iteratively update policy.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
num_state_action_successor_states: array[int], number of successor
states s' that can be reached from state s by taking action a.
sp_idx: array[int], state indices of new state s' of taking action a
from state s.
p: array[float], transition probability to go from state s to s' by
taking action a.
r: array[float], reward from new state s' from state s by taking
action a.
num_qs: int, number of state-action-value functions Q_i(s, a).
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
policy: array[float], learned stochastic policy of which
action a to take in state s.
alpha: float, alpha > 0, learning rate.
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
gamma: float, 0 <= gamma <= 1, amount to discount future reward.
maximum_episode_length: int, max number of timesteps for an episode.
s_idx: int, current state index.
a_idx: int, current action index.
Returns:
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
policy: array[float], learned stochastic policy of which
action a to take in state s.
"""
# Loop through episode steps until termination
for t in range(0, maximum_episode_length):
# Get reward
successor_state_transition_idx = np.random.choice(
a=num_state_action_successor_states[s_idx, a_idx],
p=p[s_idx, a_idx, :])
reward = r[s_idx, a_idx, successor_state_transition_idx]
# Get next state
next_s_idx = sp_idx[s_idx, a_idx, successor_state_transition_idx]
# Update state action value equally randomly selecting from the
# state-action-value functions
updating_q_idx = np.random.randint(low=0, high=num_qs, dtype=np.int64)
q, policy, s_idx, a_idx = update_q(
num_non_terminal_states,
max_num_actions,
policy,
alpha,
epsilon,
gamma,
s_idx,
a_idx,
reward,
next_s_idx,
updating_q_idx,
num_qs,
q)
if next_s_idx >= num_non_terminal_states:
break # episode terminated since we ended up in a terminal state
return q, policy
def update_q(
num_non_terminal_states,
max_num_actions,
policy,
alpha,
epsilon,
gamma,
s_idx,
a_idx,
reward,
next_s_idx,
updating_q_idx,
num_qs,
q):
"""Updates state-action-value function using multiple estimates.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
policy: array[float], learned stochastic policy of which
action a to take in state s.
alpha: float, alpha > 0, learning rate.
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
gamma: float, 0 <= gamma <= 1, amount to discount future reward.
s_idx: int, current state index.
a_idx: int, current action index.
reward: float, current reward from taking action a_idx in state s_idx.
next_s_idx: int, next state index.
updating_q_idx: int, index to which Q_i(s, a) we'll be updating.
num_qs: int, number of state-action-value functions Q_i(s, a).
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
Returns:
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
policy: array[float], learned stochastic policy of which
action a to take in state s.
s_idx: int, new current state index.
a_idx: int, new current action index.
"""
# Check to see if we actioned into a terminal state
if next_s_idx >= num_non_terminal_states:
delta = reward - q[updating_q_idx, s_idx, a_idx]
q[updating_q_idx, s_idx, a_idx] += alpha * delta
else:
# Choose policy for chosen state by epsilon-greedy choosing from the
# state-action-value function
policy = epsilon_greedy_policy_from_state_action_function(
max_num_actions, q, epsilon, next_s_idx, policy)
# Get next action
next_a_idx = np.random.choice(
a=max_num_actions, p=policy[next_s_idx, :])
# Calculate state-action-function using quintuple SARSA
q_indices = np.arange(num_qs)
not_updating_q_idx = np.random.choice(
a=np.extract(condition=q_indices != updating_q_idx, arr=q_indices))
delta = gamma * q[not_updating_q_idx, next_s_idx, next_a_idx]
delta -= q[updating_q_idx, s_idx, a_idx]
q[updating_q_idx, s_idx, a_idx] += alpha * (reward + delta)
# Update state and action to next state and action
s_idx = next_s_idx
a_idx = next_a_idx
return q, policy, s_idx, a_idx
def on_policy_temporal_difference_n_tuple_sarsa(
num_non_terminal_states,
max_num_actions,
num_state_action_successor_states,
sp_idx,
p,
r,
num_qs,
q,
policy,
alpha,
epsilon,
gamma,
maximum_episode_length,
num_episodes):
"""Loops through episodes to iteratively update policy.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
num_state_action_successor_states: array[int], number of successor
states s' that can be reached from state s by taking action a.
sp_idx: array[int], state indices of new state s' of taking action a
from state s.
p: array[float], transition probability to go from state s to s' by
taking action a.
r: array[float], reward from new state s' from state s by taking
action a.
num_qs: int, number of state-action-value functions Q_i(s, a).
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
policy: array[float], learned stochastic policy of which
action a to take in state s.
alpha: float, alpha > 0, learning rate.
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
gamma: float, 0 <= gamma <= 1, amount to discount future reward.
maximum_episode_length: int, max number of timesteps for an episode.
num_episodes: int, number of episodes to train over.
Returns:
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
policy: array[float], learned stochastic policy of which
action a to take in state s.
"""
for episode in range(0, num_episodes):
# Initialize episode to get initial state and action
init_s_idx, init_a_idx, policy = initialize_epsiode(
num_non_terminal_states, max_num_actions, q, epsilon, policy)
# Loop through episode and update the policy
q, policy = loop_through_episode(
num_non_terminal_states,
max_num_actions,
num_state_action_successor_states,
sp_idx,
p,
r,
num_qs,
q,
policy,
alpha,
epsilon,
gamma,
maximum_episode_length,
init_s_idx,
init_a_idx)
return q, policy
```
## Run algorithm
```
def run_algorithm():
"""Runs the algorithm."""
(num_states,
num_terminal_states,
num_non_terminal_states,
max_num_actions,
num_actions_per_non_terminal_state,
num_state_action_successor_states,
sp_idx,
p,
r) = create_environment()
(num_episodes,
maximum_episode_length,
num_qs,
alpha,
epsilon,
gamma) = set_hyperparameters()
q = create_value_function_arrays(num_qs, num_states, max_num_actions)
policy = create_policy_arrays(num_non_terminal_states, max_num_actions)
# Print initial arrays
print("\nInitial state-action value function")
print(q)
print("\nInitial policy")
print(policy)
# Run on policy temporal difference n-tuple sarsa
q, policy = on_policy_temporal_difference_n_tuple_sarsa(
num_non_terminal_states,
max_num_actions,
num_state_action_successor_states,
sp_idx,
p,
r,
num_qs,
q,
policy,
alpha,
epsilon,
gamma,
maximum_episode_length,
num_episodes)
# Print final results
print("\nFinal state-action value function")
print(q)
print("\nFinal policy")
print(policy)
run_algorithm()
```
| github_jupyter |
# Classification on Iris dataset with sklearn and DJL
In this notebook, you will try to use a pre-trained sklearn model to run on DJL for a general classification task. The model was trained with [Iris flower dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set).
## Background
### Iris Dataset
The dataset contains a set of 150 records under five attributes - sepal length, sepal width, petal length, petal width and species.
Iris setosa | Iris versicolor | Iris virginica
:-------------------------:|:-------------------------:|:-------------------------:
 |  | 
The chart above shows three different kinds of the Iris flowers.
We will use sepal length, sepal width, petal length, petal width as the feature and species as the label to train the model.
### Sklearn Model
You can find more information [here](http://onnx.ai/sklearn-onnx/). You can use the sklearn built-in iris dataset to load the data. Then we defined a [RandomForestClassifer](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html) to train the model. After that, we convert the model to onnx format for DJL to run inference. The following code is a sample classification setup using sklearn:
```python
# Train a model.
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
iris = load_iris()
X, y = iris.data, iris.target
X_train, X_test, y_train, y_test = train_test_split(X, y)
clr = RandomForestClassifier()
clr.fit(X_train, y_train)
```
## Preparation
This tutorial requires the installation of Java Kernel. To install the Java Kernel, see the [README](https://github.com/awslabs/djl/blob/master/jupyter/README.md).
These are dependencies we will use. To enhance the NDArray operation capability, we are importing ONNX Runtime and PyTorch Engine at the same time. Please find more information [here](https://github.com/awslabs/djl/blob/master/docs/onnxruntime/hybrid_engine.md#hybrid-engine-for-onnx-runtime).
```
// %mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
%maven ai.djl:api:0.8.0
%maven ai.djl.onnxruntime:onnxruntime-engine:0.8.0
%maven ai.djl.pytorch:pytorch-engine:0.8.0
%maven org.slf4j:slf4j-api:1.7.26
%maven org.slf4j:slf4j-simple:1.7.26
%maven com.microsoft.onnxruntime:onnxruntime:1.4.0
%maven ai.djl.pytorch:pytorch-native-auto:1.6.0
import ai.djl.inference.*;
import ai.djl.modality.*;
import ai.djl.ndarray.*;
import ai.djl.ndarray.types.*;
import ai.djl.repository.zoo.*;
import ai.djl.translate.*;
import java.util.*;
```
## Step 1 create a Translator
Inference in machine learning is the process of predicting the output for a given input based on a pre-defined model.
DJL abstracts away the whole process for ease of use. It can load the model, perform inference on the input, and provide
output. DJL also allows you to provide user-defined inputs. The workflow looks like the following:
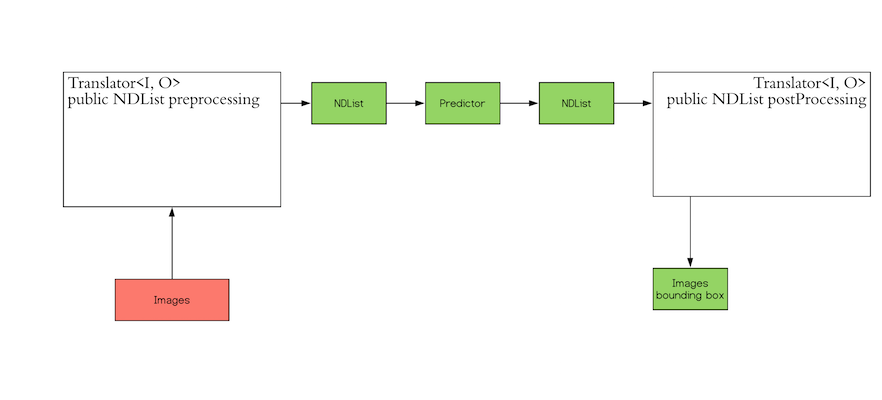
The `Translator` interface encompasses the two white blocks: Pre-processing and Post-processing. The pre-processing
component converts the user-defined input objects into an NDList, so that the `Predictor` in DJL can understand the
input and make its prediction. Similarly, the post-processing block receives an NDList as the output from the
`Predictor`. The post-processing block allows you to convert the output from the `Predictor` to the desired output
format.
In our use case, we use a class namely `FlowerInfo` as our input class type. We will use [`Classifications`](https://javadoc.io/doc/ai.djl/api/latest/ai/djl/modality/Classifications.html) as our output class type.
```
public static class FlowerInfo {
public float sepalLength;
public float sepalWidth;
public float petalLength;
public float petalWidth;
public FlowerInfo(float sepalLength, float sepalWidth, float petalLength, float petalWidth) {
this.sepalLength = sepalLength;
this.sepalWidth = sepalWidth;
this.petalLength = petalLength;
this.petalWidth = petalWidth;
}
}
```
Let's create a translator
```
public static class MyTranslator implements Translator<FlowerInfo, Classifications> {
private final List<String> synset;
public MyTranslator() {
// species name
synset = Arrays.asList("setosa", "versicolor", "virginica");
}
@Override
public NDList processInput(TranslatorContext ctx, FlowerInfo input) {
float[] data = {input.sepalLength, input.sepalWidth, input.petalLength, input.petalWidth};
NDArray array = ctx.getNDManager().create(data, new Shape(1, 4));
return new NDList(array);
}
@Override
public Classifications processOutput(TranslatorContext ctx, NDList list) {
return new Classifications(synset, list.get(1));
}
@Override
public Batchifier getBatchifier() {
return null;
}
}
```
## Step 2 Prepare your model
We will load a pretrained sklearn model into DJL. We defined a [`ModelZoo`](https://javadoc.io/doc/ai.djl/api/latest/ai/djl/repository/zoo/ModelZoo.html) concept to allow user load model from varity of locations, such as remote URL, local files or DJL pretrained model zoo. We need to define `Criteria` class to help the modelzoo locate the model and attach translator. In this example, we download a compressed ONNX model from S3.
```
String modelUrl = "https://alpha-djl-demos.s3.amazonaws.com/model/sklearn/rf_iris.zip";
Criteria<FlowerInfo, Classifications> criteria = Criteria.builder()
.setTypes(FlowerInfo.class, Classifications.class)
.optModelUrls(modelUrl)
.optTranslator(new MyTranslator())
.optEngine("OnnxRuntime") // use OnnxRuntime engine by default
.build();
ZooModel<FlowerInfo, Classifications> model = ModelZoo.loadModel(criteria);
```
## Step 3 Run inference
User will just need to create a `Predictor` from model to run the inference.
```
Predictor<FlowerInfo, Classifications> predictor = model.newPredictor();
FlowerInfo info = new FlowerInfo(1.0f, 2.0f, 3.0f, 4.0f);
predictor.predict(info);
```
| github_jupyter |
<a href="https://colab.research.google.com/github/martin-fabbri/colab-notebooks/blob/master/deeplearning.ai/tf/trax_ner_reformer.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# NER Reformer
## Named Entity Recognition
Named-entity recognition (NER) (also known as entity identification, entity chunking and entity extraction) is a subtask of information extraction that seeks to locate and classify named entities mentioned in unstructured text into pre-defined categories such as person names, organizations, locations, medical codes, time expressions, quantities, monetary values, percentages, etc.
```
#@title ## Install Dependencies
#@markdown - trax
#@markdown - kaggle client: downloads dataset
%%capture --no-stdout --no-stderr
!pip install -Uqq trax
!pip install -Uqq kaggle
# %%python
print("Dependencies successfully installed.")
#@title ## Download Kaggle Dataset
#@markdown Dataset: Annotated Corpus for Named Entity Recognition <br>
#@markdown [https://www.kaggle.com/abhinavwalia95/entity-annotated-corpus](https://www.kaggle.com/abhinavwalia95/entity-annotated-corpus)
#@markdown <br><br>
#@markdown This is the extract from GMB corpus which is tagged, annotated and built specifically to train the classifier to predict named entities such as name, location, etc.
from google.colab import drive
drive.mount('/content/drive')
!mkdir -p ~/.kaggle
!cp /content/drive/MyDrive/kaggle/kaggle.json ~/.kaggle/kaggle.json
!chmod 600 ~/.kaggle/kaggle.json
!kaggle datasets download -d abhinavwalia95/entity-annotated-corpus
!unzip -o /content/entity-annotated-corpus
#@title ## Import packages
#@markdown DL framework: trax<br>
#@markdown Data Manipulation: pandas<br>
import random as rnd
import numpy as np
import pandas as pd
import trax
from trax import layers as tl
#print('trax:', trax.__version__)
print('numpy:', np.__version__)
print('pandas:', pd.__version__)
```
## Preprocessing
Padding tokens
```
PAD_TOKEN = "PAD"
PAD_INDEX = 0
PAD_TAG = "O"
```
Loading dataset
```
data = pd.read_csv("ner_dataset.csv", encoding="ISO-8859-1", error_bad_lines=False)
#data = data.rename(columns={"Sentence #": "sentence_id", "Word": "word", "Tag": "tag"})
#data = data[["sentence_id", "word", "tag"]]
data = data.fillna(method= "ffill")
data.head(3)
```
Tag Entities
```
#data["tag"].value_counts()
```
Build Vocab
```
## Extract the 'Word' column from the dataframe
words = data.loc[:, "Word"]
!touch words.txt
vocab = {}
with open('words.txt') as f:
for i, l in enumerate(f.read().splitlines()):
vocab[l] = i
print("Number of words:", len(vocab))
vocab['<PAD>'] = len(vocab)
## Convert into a text file using the .savetxt() function
np.savetxt(r'words.txt', words.values, fmt="%s")
class Get_sentence(object):
def __init__(self,data):
self.n_sent=1
self.data = data
agg_func = lambda s:[(w,p,t) for w,p,t in zip(s["Word"].values.tolist(),
s["POS"].values.tolist(),
s["Tag"].values.tolist())]
self.grouped = self.data.groupby("Sentence #").apply(agg_func)
self.sentences = [s for s in self.grouped]
getter = Get_sentence(data)
sentence = getter.sentences
words = list(set(data["Word"].values))
words_tag = list(set(data["Tag"].values))
word_idx = {w : i+1 for i ,w in enumerate(words)}
tag_idx = {t : i for i ,t in enumerate(words_tag)}
X = [[word_idx[w[0]] for w in s] for s in sentence]
y = [[tag_idx[w[2]] for w in s] for s in sentence]
def data_generator(batch_size, x, y,pad, shuffle=False, verbose=False):
num_lines = len(x)
lines_index = [*range(num_lines)]
if shuffle:
rnd.shuffle(lines_index)
index = 0
while True:
buffer_x = [0] * batch_size
buffer_y = [0] * batch_size
max_len = 0
for i in range(batch_size):
if index >= num_lines:
index = 0
if shuffle:
rnd.shuffle(lines_index)
buffer_x[i] = x[lines_index[index]]
buffer_y[i] = y[lines_index[index]]
lenx = len(x[lines_index[index]])
if lenx > max_len:
max_len = lenx
index += 1
X = np.full((batch_size, max_len), pad)
Y = np.full((batch_size, max_len), pad)
for i in range(batch_size):
x_i = buffer_x[i]
y_i = buffer_y[i]
for j in range(len(x_i)):
X[i, j] = x_i[j]
Y[i, j] = y_i[j]
if verbose: print("index=", index)
yield((X,Y))
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test = train_test_split(X,y,test_size = 0.1,random_state=1)
```
## Model
```
!pip install --upgrade jax # install jax(base)
def NERmodel(tags, vocab_size=35178, d_model=50):
model = tl.Serial(
trax.models.reformer.Reformer(vocab_size, d_model, ff_activation=tl.LogSoftmax),
tl.Dense(tags),
tl.LogSoftmax()
)
return model
model = NERmodel(tags=17)
#print(model)
from trax.supervised import training
rnd.seed(33)
batch_size = 64
train_generator = trax.data.inputs.add_loss_weights(
data_generator(batch_size, x_train, y_train,vocab['<PAD>'], True),
id_to_mask=vocab['<PAD>'])
eval_generator = trax.data.inputs.add_loss_weights(
data_generator(batch_size, x_test, y_test,vocab['<PAD>'] ,True),
id_to_mask=vocab['<PAD>'])
def train_model(model, train_generator, eval_generator, train_steps=1, output_dir='model'):
train_task = training.TrainTask(
train_generator,
loss_layer = tl.CrossEntropyLoss(),
optimizer = trax.optimizers.Adam(0.01),
n_steps_per_checkpoint=10
)
eval_task = training.EvalTask(
labeled_data = eval_generator,
metrics = [tl.CrossEntropyLoss(), tl.Accuracy()],
n_eval_batches = 10
)
training_loop = training.Loop(
model,
train_task,
eval_tasks = eval_task,
output_dir = output_dir)
training_loop.run(n_steps = train_steps)
return training_loop
train_steps = 100
training_loop = train_model(model, train_generator, eval_generator, train_steps)
train_steps
```
| github_jupyter |
# Transfer Learning
In this notebook, you'll learn how to use pre-trained networks to solved challenging problems in computer vision. Specifically, you'll use networks trained on [ImageNet](http://www.image-net.org/) [available from torchvision](http://pytorch.org/docs/0.3.0/torchvision/models.html).
ImageNet is a massive dataset with over 1 million labeled images in 1000 categories. It's used to train deep neural networks using an architecture called convolutional layers. I'm not going to get into the details of convolutional networks here, but if you want to learn more about them, please [watch this](https://www.youtube.com/watch?v=2-Ol7ZB0MmU).
Once trained, these models work astonishingly well as feature detectors for images they weren't trained on. Using a pre-trained network on images not in the training set is called transfer learning. Here we'll use transfer learning to train a network that can classify our cat and dog photos with near perfect accuracy.
With `torchvision.models` you can download these pre-trained networks and use them in your applications. We'll include `models` in our imports now.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models
```
Most of the pretrained models require the input to be 224x224 images. Also, we'll need to match the normalization used when the models were trained. Each color channel was normalized separately, the means are `[0.485, 0.456, 0.406]` and the standard deviations are `[0.229, 0.224, 0.225]`.
```
data_dir = 'Cat_Dog_data'
# TODO: Define transforms for the training data and testing data
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],[0.229, 0.224, 0.225])])
test_transforms = transforms.Compose([transforms.Resize(225),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],[0.229, 0.224, 0.225])])
# Pass transforms in here, then run the next cell to see how the transforms look
train_data = datasets.ImageFolder(data_dir + '/train', transform=train_transforms)
test_data = datasets.ImageFolder(data_dir + '/test', transform=test_transforms)
trainloader = torch.utils.data.DataLoader(train_data, batch_size=64, shuffle=True)
testloader = torch.utils.data.DataLoader(test_data, batch_size=64)
```
We can load in a model such as [DenseNet](http://pytorch.org/docs/0.3.0/torchvision/models.html#id5). Let's print out the model architecture so we can see what's going on.
```
model = models.densenet121(pretrained=True)
model
```
This model is built out of two main parts, the features and the classifier. The features part is a stack of convolutional layers and overall works as a feature detector that can be fed into a classifier. The classifier part is a single fully-connected layer `(classifier): Linear(in_features=1024, out_features=1000)`. This layer was trained on the ImageNet dataset, so it won't work for our specific problem. That means we need to replace the classifier, but the features will work perfectly on their own. In general, I think about pre-trained networks as amazingly good feature detectors that can be used as the input for simple feed-forward classifiers.
```
# Freeze parameters so we don't backprop through them
for param in model.parameters():
param.requires_grad =False
from collections import OrderedDict
classifier = nn.Sequential(OrderedDict([
('fc1', nn.Linear(1024, 500)),
('relu', nn.ReLU()),
('fc2', nn.Linear(500, 2)),
('output', nn.LogSoftmax(dim=1))
]))
model.classifier = classifier
```
With our model built, we need to train the classifier. However, now we're using a **really deep** neural network. If you try to train this on a CPU like normal, it will take a long, long time. Instead, we're going to use the GPU to do the calculations. The linear algebra computations are done in parallel on the GPU leading to 100x increased training speeds. It's also possible to train on multiple GPUs, further decreasing training time.
PyTorch, along with pretty much every other deep learning framework, uses [CUDA](https://developer.nvidia.com/cuda-zone) to efficiently compute the forward and backwards passes on the GPU. In PyTorch, you move your model parameters and other tensors to the GPU memory using `model.to('cuda')`. You can move them back from the GPU with `model.to('cpu')` which you'll commonly do when you need to operate on the network output outside of PyTorch. As a demonstration of the increased speed, I'll compare how long it takes to perform a forward and backward pass with and without a GPU.
```
import time
for device in ['cpu', 'cuda']:
criterion = nn.NLLLoss()
# Only train the classifier parameters, feature parameters are frozen
optimizer = optim.Adam(model.classifier.parameters(), lr=0.001)
model.to(device)
for ii, (inputs, labels) in enumerate(trainloader):
# Move input and label tensors to the GPU
inputs, labels = inputs.to(device), labels.to(device)
start = time.time()
outputs = model.forward(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
if ii==3:
break
print(f"Device = {device}; Time per batch: {(time.time() - start)/3:.3f} seconds")
```
You can write device agnostic code which will automatically use CUDA if it's enabled like so:
```python
# at beginning of the script
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
...
# then whenever you get a new Tensor or Module
# this won't copy if they are already on the desired device
input = data.to(device)
model = MyModule(...).to(device)
```
From here, I'll let you finish training the model. The process is the same as before except now your model is much more powerful. You should get better than 95% accuracy easily.
>**Exercise:** Train a pretrained models to classify the cat and dog images. Continue with the DenseNet model, or try ResNet, it's also a good model to try out first. Make sure you are only training the classifier and the parameters for the features part are frozen.
```
# Use GPU if it's available
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model = models.densenet121(pretrained=True)
# Freeze parameters so we don't backprop through them
for param in model.parameters():
param.requires_grad = False
model.classifier = nn.Sequential(nn.Linear(1024, 256),
nn.ReLU(),
nn.Dropout(0.2),
nn.Linear(256, 2),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
# Only train the classifier parameters, feature parameters are frozen
optimizer = optim.Adam(model.classifier.parameters(), lr=0.003)
model.to(device);
epochs = 1
steps = 0
running_loss = 0
print_every = 5
for epoch in range(epochs):
for inputs, labels in trainloader:
steps += 1
# Move input and label tensors to the default device
inputs, labels = inputs.to(device), labels.to(device)
optimizer.zero_grad()
logps = model.forward(inputs)
loss = criterion(logps, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
if steps % print_every == 0:
test_loss = 0
accuracy = 0
model.eval()
with torch.no_grad():
for inputs, labels in testloader:
inputs, labels = inputs.to(device), labels.to(device)
logps = model.forward(inputs)
batch_loss = criterion(logps, labels)
test_loss += batch_loss.item()
# Calculate accuracy
ps = torch.exp(logps)
top_p, top_class = ps.topk(1, dim=1)
equals = top_class == labels.view(*top_class.shape)
accuracy += torch.mean(equals.type(torch.FloatTensor)).item()
print(f"Epoch {epoch+1}/{epochs}.. "
f"Train loss: {running_loss/print_every:.3f}.. "
f"Test loss: {test_loss/len(testloader):.3f}.. "
f"Test accuracy: {accuracy/len(testloader):.3f}")
running_loss = 0
model.train()
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
a = np.zeros(3, dtype=int)
a
z = np.zeros(10)
print(z)
print(z.shape)
z.shape = (10,1)
print(z)
z = np.zeros(4)
z.shape = (2,2)
print(z)
z = np.zeros((2,2))
print(z)
z = np.ones((2,3))
print(z)
z = np.empty((3,3))
print(z)
z = np.identity(3)
print(z)
blalist = [222, 22, 2, 0.2]
bla2list = (222, 22, 2, 0.2)
bla3list = (111, 11, 1, 0.1)
bla4list = (0, 0, 0, 0)
a = np.array(blalist)
b = np.array(bla2list)
c = np.array([bla3list, bla4list])
a.shape = (2,2)
b.shape = (2,2)
c.shape = (8,1)
print(a)
print(b)
print(c)
z = np.linspace(1,2,5)
print(z[0])
print(z[0:5])
print(z[-3])
z = np.array([[2,3], [4,5]])
print(z[0,0])
print(z[0,1])
print(z[1,0])
print(z[1,1])
z = np.array([[2,3], [4,5]])
print(z[0,:])
print(z[1,:])
print(z[:,0])
print(z[:,1])
z = np.linspace(1,2,5)
a = np.array((0,2,3))
print(z[a])
b = np.array((1,0,1,1,0), dtype=bool)
print(z[b])
z = np.empty((3,3))
z[:] = 42
z
z = np.array([4,3,2,1])
z.sort()
z
z = np.array([4,3,2,1])
z.sum()
z = np.array([4,3,2,1])
z.mean()
z = np.array([4,3,2,1])
z.max()
z = np.array([4,3,2,1])
z.argmax()
z = np.array([4,3,2,1])
z.cumsum()
z = np.array([4,3,2,1])
z.cumprod()
z = np.array([4,3,2,1])
z.var()
z = np.array([4,3,2,1])
z.std()
z = np.linspace(1,9,9)
z.shape = (3,3)
print(z)
print(z.T)
z = np.array([1,2,3,4])
z.searchsorted(2)
z = np.array([4,3,2,1])
np.sum(z)
z = np.array([4,3,2,1])
np.mean(z)
z = np.array([4,3,2,1])
np.max(z)
z = np.array([4,3,2,1])
np.var(z)
z = np.array([4,3,2,1])
np.std(z)
z = np.array([[1,2], [1,2]])
y = np.array([[1,2], [1,2]])
print(z+y)
print(z-y)
print(z*y)
print(z/y)
print(z**y)
print(z@y)
z = np.array([4,3,2,1])
y = np.copy(z)
y is z
z = np.array([4,3,2,1])
np.sin(z)
np.where(z<=1, 1, 0)
def f(x):
return 1 if x>=1 else 0
f = np.vectorize(f)
z = np.array([4,3,2,1])
%%time
for i in range(100000):
f(z)
%%time
for i in range(100000):
np.where(z>=1, 1, 0)
z = np.array([1,2,3,4])
y = np.copy(z)
y == z
y != z
y >= z
a = y > 3
y[a]
#easier:
y[y>3]
z = np.random.randn(1000)
y = np.random.binomial(1, 0.5, size=100)
def p(x, coeff):
a = np.ones(len(coeff))
a[1:] = x
return np.sum(np.cumprod(a) * coeff)
coeff = np.array([0,0,0,1])
a = -10
b = 10
res = 112
xrange = np.linspace(a,b,res)
for x in xrange:
plt.scatter(x, p(x, coeff), alpha=0.4, c='r', s=20)
%%time
class ecdf:
def __init__(self, observations): #initialise ecdf object with sample s which contains a set of observations
self.observations = np.asarray(observations)
def __call__(self, x): #call ecdf value-calculation at x
self.x = x
functionvalue = np.mean(self.observations <= x)
return functionvalue
def plot(self, a, b, res):
self.a, self.b, self.res = a, b, res
f = np.vectorize(self.__call__)
xvals = np.linspace(self.a, self.b, self.res)
plt.scatter(xvals, f(xvals), alpha=0.2)
import numpy as np
import matplotlib.pyplot as plt
trials=50 #how many picks from distribution?
resolution=100 #Number of calculations between every natural number (minimum 1)
mean = 0
std = 1
a = mean - 4*std
b = mean + 4*std
sample = np.random.normal(loc=mean, scale=std, size=trials)
#sample through a distribution
#and pick a trial number of values
f_n = ecdf(sample)
f_n.plot(a, b, resolution)
```
| github_jupyter |
```
#IMPORT SEMUA LIBARARY
#IMPORT LIBRARY PANDAS
import pandas as pd
#IMPORT LIBRARY UNTUK POSTGRE
from sqlalchemy import create_engine
import psycopg2
#IMPORT LIBRARY CHART
from matplotlib import pyplot as plt
from matplotlib import style
#IMPORT LIBRARY BASE PATH
import os
import io
#IMPORT LIBARARY PDF
from fpdf import FPDF
#IMPORT LIBARARY CHART KE BASE64
import base64
#IMPORT LIBARARY EXCEL
import xlsxwriter
#FUNGSI UNTUK MENGUPLOAD DATA DARI CSV KE POSTGRESQL
def uploadToPSQL(columns, table, filePath, engine):
#FUNGSI UNTUK MEMBACA CSV
df = pd.read_csv(
os.path.abspath(filePath),
names=columns,
keep_default_na=False
)
#APABILA ADA FIELD KOSONG DISINI DIFILTER
df.fillna('')
#MENGHAPUS COLUMN YANG TIDAK DIGUNAKAN
del df['kategori']
del df['jenis']
del df['pengiriman']
del df['satuan']
#MEMINDAHKAN DATA DARI CSV KE POSTGRESQL
df.to_sql(
table,
engine,
if_exists='replace'
)
#DIHITUNG APABILA DATA YANG DIUPLOAD BERHASIL, MAKA AKAN MENGEMBALIKAN KELUARAN TRUE(BENAR) DAN SEBALIKNYA
if len(df) == 0:
return False
else:
return True
#FUNGSI UNTUK MEMBUAT CHART, DATA YANG DIAMBIL DARI DATABASE DENGAN MENGGUNAKAN ORDER DARI TANGGAL DAN JUGA LIMIT
#DISINI JUGA MEMANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
def makeChart(host, username, password, db, port, table, judul, columns, filePath, name, subjudul, limit, negara, basePath):
#TEST KONEKSI DATABASE
try:
#KONEKSI KE DATABASE
connection = psycopg2.connect(user=username,password=password,host=host,port=port,database=db)
cursor = connection.cursor()
#MENGAMBL DATA DARI TABLE YANG DIDEFINISIKAN DIBAWAH, DAN DIORDER DARI TANGGAL TERAKHIR
#BISA DITAMBAHKAN LIMIT SUPAYA DATA YANG DIAMBIL TIDAK TERLALU BANYAK DAN BERAT
postgreSQL_select_Query = "SELECT * FROM "+table+" ORDER BY tanggal ASC LIMIT " + str(limit)
cursor.execute(postgreSQL_select_Query)
mobile_records = cursor.fetchall()
uid = []
lengthx = []
lengthy = []
#MELAKUKAN LOOPING ATAU PERULANGAN DARI DATA YANG SUDAH DIAMBIL
#KEMUDIAN DATA TERSEBUT DITEMPELKAN KE VARIABLE DIATAS INI
for row in mobile_records:
uid.append(row[0])
lengthx.append(row[1])
if row[2] == "":
lengthy.append(float(0))
else:
lengthy.append(float(row[2]))
#FUNGSI UNTUK MEMBUAT CHART
#bar
style.use('ggplot')
fig, ax = plt.subplots()
#MASUKAN DATA ID DARI DATABASE, DAN JUGA DATA TANGGAL
ax.bar(uid, lengthy, align='center')
#UNTUK JUDUL CHARTNYA
ax.set_title(judul)
ax.set_ylabel('Total')
ax.set_xlabel('Tanggal')
ax.set_xticks(uid)
#TOTAL DATA YANG DIAMBIL DARI DATABASE, DIMASUKAN DISINI
ax.set_xticklabels((lengthx))
b = io.BytesIO()
#CHART DISIMPAN KE FORMAT PNG
plt.savefig(b, format='png', bbox_inches="tight")
#CHART YANG SUDAH DIJADIKAN PNG, DISINI DICONVERT KE BASE64
barChart = base64.b64encode(b.getvalue()).decode("utf-8").replace("\n", "")
#CHART DITAMPILKAN
plt.show()
#line
#MASUKAN DATA DARI DATABASE
plt.plot(lengthx, lengthy)
plt.xlabel('Tanggal')
plt.ylabel('Total')
#UNTUK JUDUL CHARTNYA
plt.title(judul)
plt.grid(True)
l = io.BytesIO()
#CHART DISIMPAN KE FORMAT PNG
plt.savefig(l, format='png', bbox_inches="tight")
#CHART YANG SUDAH DIJADIKAN PNG, DISINI DICONVERT KE BASE64
lineChart = base64.b64encode(l.getvalue()).decode("utf-8").replace("\n", "")
#CHART DITAMPILKAN
plt.show()
#pie
#UNTUK JUDUL CHARTNYA
plt.title(judul)
#MASUKAN DATA DARI DATABASE
plt.pie(lengthy, labels=lengthx, autopct='%1.1f%%',
shadow=True, startangle=180)
plt.axis('equal')
p = io.BytesIO()
#CHART DISIMPAN KE FORMAT PNG
plt.savefig(p, format='png', bbox_inches="tight")
#CHART YANG SUDAH DIJADIKAN PNG, DISINI DICONVERT KE BASE64
pieChart = base64.b64encode(p.getvalue()).decode("utf-8").replace("\n", "")
#CHART DITAMPILKAN
plt.show()
#MENGAMBIL DATA DARI CSV YANG DIGUNAKAN SEBAGAI HEADER DARI TABLE UNTUK EXCEL DAN JUGA PDF
header = pd.read_csv(
os.path.abspath(filePath),
names=columns,
keep_default_na=False
)
#MENGHAPUS COLUMN YANG TIDAK DIGUNAKAN
header.fillna('')
del header['tanggal']
del header['total']
#MEMANGGIL FUNGSI EXCEL
makeExcel(mobile_records, header, name, limit, basePath)
#MEMANGGIL FUNGSI PDF
makePDF(mobile_records, header, judul, barChart, lineChart, pieChart, name, subjudul, limit, basePath)
#JIKA GAGAL KONEKSI KE DATABASE, MASUK KESINI UNTUK MENAMPILKAN ERRORNYA
except (Exception, psycopg2.Error) as error :
print (error)
#KONEKSI DITUTUP
finally:
if(connection):
cursor.close()
connection.close()
#FUNGSI MAKEEXCEL GUNANYA UNTUK MEMBUAT DATA YANG BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH XLSXWRITER
def makeExcel(datarow, dataheader, name, limit, basePath):
#MEMBUAT FILE EXCEL
workbook = xlsxwriter.Workbook(basePath+'jupyter/BLOOMBERG/SektorIndustri/excel/'+name+'.xlsx')
#MENAMBAHKAN WORKSHEET PADA FILE EXCEL TERSEBUT
worksheet = workbook.add_worksheet('sheet1')
#SETINGAN AGAR DIBERIKAN BORDER DAN FONT MENJADI BOLD
row1 = workbook.add_format({'border': 2, 'bold': 1})
row2 = workbook.add_format({'border': 2})
#MENJADIKAN DATA MENJADI ARRAY
data=list(datarow)
isihead=list(dataheader.values)
header = []
body = []
#LOOPING ATAU PERULANGAN, KEMUDIAN DATA DITAMPUNG PADA VARIABLE DIATAS
for rowhead in dataheader:
header.append(str(rowhead))
for rowhead2 in datarow:
header.append(str(rowhead2[1]))
for rowbody in isihead[1]:
body.append(str(rowbody))
for rowbody2 in data:
body.append(str(rowbody2[2]))
#MEMASUKAN DATA DARI VARIABLE DIATAS KE DALAM COLUMN DAN ROW EXCEL
for col_num, data in enumerate(header):
worksheet.write(0, col_num, data, row1)
for col_num, data in enumerate(body):
worksheet.write(1, col_num, data, row2)
#FILE EXCEL DITUTUP
workbook.close()
#FUNGSI UNTUK MEMBUAT PDF YANG DATANYA BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH FPDF
def makePDF(datarow, dataheader, judul, bar, line, pie, name, subjudul, lengthPDF, basePath):
#FUNGSI UNTUK MENGATUR UKURAN KERTAS, DISINI MENGGUNAKAN UKURAN A4 DENGAN POSISI LANDSCAPE
pdf = FPDF('L', 'mm', [210,297])
#MENAMBAHKAN HALAMAN PADA PDF
pdf.add_page()
#PENGATURAN UNTUK JARAK PADDING DAN JUGA UKURAN FONT
pdf.set_font('helvetica', 'B', 20.0)
pdf.set_xy(145.0, 15.0)
#MEMASUKAN JUDUL KE DALAM PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=judul, border=0)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_font('arial', '', 14.0)
pdf.set_xy(145.0, 25.0)
#MEMASUKAN SUB JUDUL KE PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=subjudul, border=0)
#MEMBUAT GARIS DI BAWAH SUB JUDUL
pdf.line(10.0, 30.0, 287.0, 30.0)
pdf.set_font('times', '', 10.0)
pdf.set_xy(17.0, 37.0)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_font('Times','',10.0)
#MENGAMBIL DATA HEADER PDF YANG SEBELUMNYA SUDAH DIDEFINISIKAN DIATAS
datahead=list(dataheader.values)
pdf.set_font('Times','B',12.0)
pdf.ln(0.5)
th1 = pdf.font_size
#MEMBUAT TABLE PADA PDF, DAN MENAMPILKAN DATA DARI VARIABLE YANG SUDAH DIKIRIM
pdf.cell(100, 2*th1, "Kategori", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][0], border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Jenis", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][1], border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Pengiriman", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][2], border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Satuan", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][3], border=1, align='C')
pdf.ln(2*th1)
#PENGATURAN PADDING
pdf.set_xy(17.0, 75.0)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_font('Times','B',11.0)
data=list(datarow)
epw = pdf.w - 2*pdf.l_margin
col_width = epw/(lengthPDF+1)
#PENGATURAN UNTUK JARAK PADDING
pdf.ln(0.5)
th = pdf.font_size
#MEMASUKAN DATA HEADER YANG DIKIRIM DARI VARIABLE DIATAS KE DALAM PDF
pdf.cell(50, 2*th, str("Negara"), border=1, align='C')
for row in data:
pdf.cell(40, 2*th, str(row[1]), border=1, align='C')
pdf.ln(2*th)
#MEMASUKAN DATA ISI YANG DIKIRIM DARI VARIABLE DIATAS KE DALAM PDF
pdf.set_font('Times','B',10.0)
pdf.set_font('Arial','',9)
pdf.cell(50, 2*th, negara, border=1, align='C')
for row in data:
pdf.cell(40, 2*th, str(row[2]), border=1, align='C')
pdf.ln(2*th)
#MENGAMBIL DATA CHART, KEMUDIAN CHART TERSEBUT DIJADIKAN PNG DAN DISIMPAN PADA DIRECTORY DIBAWAH INI
#BAR CHART
bardata = base64.b64decode(bar)
barname = basePath+'jupyter/BLOOMBERG/SektorIndustri/img/'+name+'-bar.png'
with open(barname, 'wb') as f:
f.write(bardata)
#LINE CHART
linedata = base64.b64decode(line)
linename = basePath+'jupyter/BLOOMBERG/SektorIndustri/img/'+name+'-line.png'
with open(linename, 'wb') as f:
f.write(linedata)
#PIE CHART
piedata = base64.b64decode(pie)
piename = basePath+'jupyter/BLOOMBERG/SektorIndustri/img/'+name+'-pie.png'
with open(piename, 'wb') as f:
f.write(piedata)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_xy(17.0, 75.0)
col = pdf.w - 2*pdf.l_margin
widthcol = col/3
#MEMANGGIL DATA GAMBAR DARI DIREKTORY DIATAS
pdf.image(barname, link='', type='',x=8, y=100, w=widthcol)
pdf.set_xy(17.0, 75.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(linename, link='', type='',x=103, y=100, w=widthcol)
pdf.set_xy(17.0, 75.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(piename, link='', type='',x=195, y=100, w=widthcol)
pdf.ln(2*th)
#MEMBUAT FILE PDF
pdf.output(basePath+'jupyter/BLOOMBERG/SektorIndustri/pdf/'+name+'.pdf', 'F')
#DISINI TEMPAT AWAL UNTUK MENDEFINISIKAN VARIABEL VARIABEL SEBELUM NANTINYA DIKIRIM KE FUNGSI
#PERTAMA MANGGIL FUNGSI UPLOADTOPSQL DULU, KALAU SUKSES BARU MANGGIL FUNGSI MAKECHART
#DAN DI MAKECHART MANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
#DEFINISIKAN COLUMN BERDASARKAN FIELD CSV
columns = [
"kategori",
"jenis",
"tanggal",
"total",
"pengiriman",
"satuan",
]
#UNTUK NAMA FILE
name = "SektorIndustri6_1"
#VARIABLE UNTUK KONEKSI KE DATABASE
host = "localhost"
username = "postgres"
password = "1234567890"
port = "5432"
database = "bloomberg_SektorIndustri"
table = name.lower()
#JUDUL PADA PDF DAN EXCEL
judul = "Data Sektor Industri"
subjudul = "Badan Perencanaan Pembangunan Nasional"
#LIMIT DATA UNTUK SELECT DI DATABASE
limitdata = int(8)
#NAMA NEGARA UNTUK DITAMPILKAN DI EXCEL DAN PDF
negara = "Indonesia"
#BASE PATH DIRECTORY
basePath = 'C:/Users/ASUS/Documents/bappenas/'
#FILE CSV
filePath = basePath+ 'data mentah/BLOOMBERG/SektorIndustri/' +name+'.csv';
#KONEKSI KE DATABASE
engine = create_engine('postgresql://'+username+':'+password+'@'+host+':'+port+'/'+database)
#MEMANGGIL FUNGSI UPLOAD TO PSQL
checkUpload = uploadToPSQL(columns, table, filePath, engine)
#MENGECEK FUNGSI DARI UPLOAD PSQL, JIKA BERHASIL LANJUT MEMBUAT FUNGSI CHART, JIKA GAGAL AKAN MENAMPILKAN PESAN ERROR
if checkUpload == True:
makeChart(host, username, password, database, port, table, judul, columns, filePath, name, subjudul, limitdata, negara, basePath)
else:
print("Error When Upload CSV")
```
| github_jupyter |
# ECCB2020 Tutorial T05: Computational modelling of cellular processes: regulatory vs metabolic systems
## Part 3: Introductions to constrainat-based modeling using cobrapy
### Instructors:
* Miguel Ponce de León from (Barcelona Supercomputing Center)
* Marta Cascante (Universidad de Barcelona)
1 Septembar, 2020
```
import cobra
from IPython.display import Image
```
### [Full cobrapy documentation](https://cobrapy.readthedocs.io/en/latest/)
### Cobrapy Documentation
[https://cobrapy.readthedocs.io](https://cobrapy.readthedocs.io) <br>
### References
* [Advances in Flux Balance Analysis](https://www.sciencedirect.com/science/article/abs/pii/S0958166903001174)
* [What is Flux Balance Analysis](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3108565/)
### Models repositories
* Manually reconstructed genome-scale models repository: [http://bigg.ucsd.edu/](http://bigg.ucsd.edu/)
* Human metabolic model: [https://www.vmh.life/](https://www.vmh.life/)
# Part 1: Warming up.
## Objective:
To get familiar with COBRA library by creating a toy model from Kauffman et al (figure 1) and manipulating it.

Figure 1. Toy model with three metabolites (A, B y C), four reactions
(v1-v4) and three exchange fluxes (b1-b3). a) Model chart; b) Stoichiometric matrix
### Let's create the model of figure to understand basic cobra objects
```
# Importing the required classes from core package
from cobra.core import Model
from cobra.core import Reaction
from cobra.core import Metabolite
# Creating the model with id Toymodel
toy_model = Model('Toymodel')
toy_model.description = 'Just a toy model'
print("We have created a model with ID:", toy_model)
print("List of model metabolites:", toy_model.metabolites)
print("List of model reactions:", toy_model.reactions)
# Creating two metabolites and set them a comparment
A = Metabolite("A", name="I'm A")
A.compartment = 'cytosol'
B = Metabolite("B", name="I'm B")
B.compartment = 'cytosol'
```
## Excercise 1.1:
Create Metabolite C
```
######################
# Create Metabolite C
######################
## TODO
## Write your code below
## Add the metabolites to the model
toy_model.add_metabolites([A, B, C])
print("List of model metabolites:")
print(toy_model.metabolites)
# We can access metabolties from the model
A = toy_model.metabolites.A
# This is the safer way to access metabolties
A = toy_model.metabolites.get_by_id("A")
# Print the reactions of a given metabolite
print(A.name)
# Print the reactions of a given metabolite
print(A.reactions)
# We get an empty set because we haven't created any reaction yet
# Creating the reactions b1, b2 ... v3
# Create reaction with id b1
b1 = Reaction("b1")
# To add metabolites to the reactions we need to pass
# a dictionary with metabolites as keys
# and the stoichiometric coefficients as values
b1.add_metabolites({A: 1})
# The same is done for the other reactions
b2 = Reaction("b2")
# Metabolites are added to the reaction by passing a dictionary:
# {metabolite_1: stoich_coef_1, ... , metabolite_n: stoich_coef_n}
b2.add_metabolites({B: -1})
b3 = Reaction("b3")
b3.add_metabolites({C: -1})
v1 = Reaction("v1")
v1.add_metabolites({A:-1, B:1})
v2 = Reaction("v2")
v2.add_metabolites({A:-1, C:1})
v3 = Reaction("v3")
v3.add_metabolites({A:1, C:-1})
```
## Excercice 1.2:
#### Create reactions v4 (and add its metabolites)
<br>Hint 1: v4 is C --> B
<br>Hint 2: check the stoich matrix of figure 1 to find the stoichiometrix coefficient
```
########################
# Create reactions v4
########################
## TODO
## Write your code beloW
# Adding the reactions to the toy model
toy_model.add_reactions([b1,b2,b3,v1,v2,v3,v4])
# print the balance equations of a metabolite
A = toy_model.metabolites.A
for r in toy_model.reactions:
if A not in r.metabolites:
stoich = 0
else:
stoich = r.get_coefficient(A)
print(A.id, r.id, stoich)
################################
# print stoichiometirc matrix
################################
## TODO
## write
```
## Flux Balance Analysis
* N: stoichiometric matrix
* v: flux vector
* $J_{irrev}$: set of irreversible reactions
* $ lb_i $: lower bound of the ith reaction flux
* $ ub_i$: upper bound of the ith reaction flux
### Objective function
$Max~Z: v_{bioamss}$
### Subject to constraints:
#### Mass Balance constraint (pseudo steady state)
$N . v = 0$
#### Thermodynamic constraints
Irreversible reactions cannot carry negative flux (reverse reaction) <br><br>
$v_i >= 0 ~~~~ {\forall i} \in J_{irrev} $
#### Capacity constraints
$lb_i <= v_i <= ub_i ~~~~ {\forall i} \in J $
```
# Setting the limits on the inputs
toy_model.reactions.b1.upper_bound = 1
toy_model.reactions.b2.upper_bound = 2
# Setting a reaction to be optimized as the objective or target
toy_model.reactions.v1.objective_coefficient = 1
toy_model.reactions.v4.objective_coefficient = 2
# To compute a FBA on the model we use the following function:
solution = toy_model.optimize()
# The result is a solution object which containsthe the following attributes:
# objective_value: the objective value of the optimized function (biomass!)
print("Objective value: %.2f\n" % solution.objective_value)
# solution.status: shows the status of the solution, it should be optimal
# if it is unfeasible, this means that there is no feasible solution
print("Status: %s\n" % solution.status)
# solution.fluxes: is a datagrame (pandas) storing the reactions (index) and
# their flux values found in the optimal solution.
print("Fluxes:\n")
print(solution.fluxes)
```
### Write the model in SBML format (to do in localhost)
1. First import the corresponding function
2. Write the model
3. Optional: You can inspect the SBML using some plain text editor.
```
import cobra.io
from cobra.io import write_sbml_model
# Saving the model to a file in SBML format
write_sbml_model(toy_model, "out/toymodel.sbml")
# Saving the solution into tab-separed-value (tsv) format (plain text)
solution.fluxes.to_csv("out/toymodel_fba.tsv", sep="\t")
# Inspect the file
```
# Part 2: Genome-scale modelling.
In this part we are gonna use a genome-scale metabolic model of Escherichia coli named iJO1366
The file has already been stored in the data folder and its path is data/iJO1366.xml
Alternatively, you can also access it here:
- [http://bigg.ucsd.edu/models/iJO1366](http://bigg.ucsd.edu/models/iJO1366)
to download the model and to see other metadata (citation, description, etc)
## Part 2.1: Studying the model.
### Read the SBML model
First we need to import the function read_sbml_model from the cobra.io modules
```
from cobra.io import read_sbml_model
# State the path to the file iJO1366.xml
sbml_fname = './data/iJO1366.xml'
# Read the model
model = read_sbml_model(sbml_fname)
```
### Exercise 2.1: Inspecting the model's numbers
First print model description
1. print(model)
2. Print the total number of reactions: print len(model.reactions)
3. Print the total number of metabolties: print len(model.metabolites)
4. Print the total number of genes: print len(model.genes)
5. Access a particular reaction:
* You can do it directly with: rxn = model.reactions.ENO
* Or you can do use the function get_by_id: rxn = model.reactions.get_by_id('ENO')
6. Inspect the reaction by printing:
1. rxn.name
2. rxn.id
3. rxn.bounds
4. rxn.reaction
5. rxn.gene_reaction_rule
```
## TODO
## Write your code below
```
### Exercise 2.2: Inspecting the genes
First print the model description
1. Access a particular reaction:
* You can do it directly with: gene = model.genes.b0720
* Or you can use the function get_by_id: gene = model.genes.get_by_id('b0720')
6. Inspect the reaction by printing:
1. gene.name
2. gene.id
3. gene.reactions
```
## TODO
## Write your code below
```
### Exercise 2.3: Inspecting the systems' boundaries
* see the exchange fluxes
* see the objective function (the reaction set to be optimized)
Use print(model.summary())
You can also find the objective function using the following filtering technique:
* [r for r in model.reactions if r.objective_coefficient == 1]
* the reaction id of the biomass is Ec_biomass_iJO1366_WT_53p95M
and the exchange fluxes can be accessed using:
* model.boundary
```
## TODO
## Write your code below
```
## Part 2.2 Running a Flux Balance Analysis (FBA).
Documentation: [https://cobrapy.readthedocs.io/en/latest/simulating.html](https://cobrapy.readthedocs.io/en/latest/simulating.html)
By default, the model boundary condition (growth medium) is M9 aerobic (glucose minimal)
1. Check the medium by inspecting the lower_bound of the following reactions:
* EX\_glc\_e\_.lower_bound
* EX\_o2\_e\_.lower_bound
2. Optimize biomass using:
* solution = model.optimize()
3. Inspect the solution as we did previously in Part 1.2 Optimization.
```
solution = model.optimize()
print("Objective value: %.2f\n" % solution.objective_value)
print("Status: %s\n" % solution.status)
print("Fluxes:\n")
print(solution.fluxes)
# Converting the solution into a pandas dataframe
df = solution.to_frame()
# Saving the solution into tab-separed-value (tsv) format (plain text)
df.to_csv("out/iJO1366_fba.tsv", sep="\t")
```
### Exercise 2.4: Identify reactions in the iJO1366 simulation:
Inspect the flux value of the following reactions
* The glucose consumption: EX_glc_e_
* The oxygen consumption: EX_o2_e_
* The biomass reaction: Ec_biomass_iJO1366_WT_53p95M
HINT 1: use the solution object -> solution.fluxes.reaction_id <br>
HINT 2: use model.summary()
```
## TODO
## Write your code below
```
## Visualizing flux distributions using Escher
[Escher documentation](https://escher.readthedocs.io/en/latest/)
Escher online WebApp: [https://escher.github.io/](https://escher.github.io/#/)
```
import escher
from escher import Builder
# Lets crate a builder by passing our model as well a given map name to tell escher how to represent the network
# Check the escher web to see other maps https://escher.github.io/#/
builder = Builder(organism='Escherichia coli', map_name='iJO1366.Central metabolism')
# Add the optimal flux distribution to our map builder
builder.reaction_data = solution.fluxes
builder
```
## Part 3: Knock out studies.
### Exercise 3.1: Single knock out study.
Documentation: [https://cobrapy.readthedocs.io/en/latest/deletions.html#Knocking-out-single-genes-and-reactions](https://cobrapy.readthedocs.io/en/latest/deletions.html#Knocking-out-single-genes-and-reactions)
We will use gene b0720 as an example
```
from cobra.manipulation import find_gene_knockout_reactions
# Pick a gene of interest
gene = model.genes.b0720
# Inspect the reactions associated to b0720
print("id\treaction_name")
for r in gene.reactions:
print("%s \t%s" % (r.id,r.name))
print()
# We can also check the genes associated to this reaction
reaction = model.reactions.CS
print("GPR:",reaction.gene_reaction_rule)
```
Alternatively, COBRA can find the proper reaction to be disabled when a gene is knocked out as follows:
gene = model.genes.b0720
with model:
gene.knock_out()
ko_solution = model.optimize()
This codes knocks out the gene b0720, recalculates the FBA and stores the new solution in ko_solution.
```
# If we perform the knockout in the "with" block we don't need to care
# about restoring the knocked out gene afterwards; it is automatically restored out of the "with" block.
with model:
gene.knock_out()
ko_solution = model.optimize()
########################################################################################
# TODO
# Check the growth value (Hint: ko_solution.fluxes.Ec_biomass_iJO1366_WT_53p95M or ko_solution.objective_values)
# What happened?
## write your code below
```
Go to the Ecocyc database and check the in vivo experimental result for the knockout of b0720
by accessing the following link:
* [https://ecocyc.org/gene?orgid=ECOLI&id=EG10402](https://ecocyc.org/gene?orgid=ECOLI&id=EG10402)
Is b0720 essential or not?
### Exercise 3.2: Systems-wide knock out study of *E. coli*.
COBRA has a special function to run the single gene knock outs of a list of genes.
The function's name is single_gene_deletion
First import the function
```
# Import the function single_gene_deletion
from cobra.flux_analysis import single_gene_deletion
# Then get the list of all the genes
all_genes = [g.id for g in model.genes]
# Running in silico (takes a while)
knockout = single_gene_deletion(model, gene_list=all_genes)
# This is a fix to get the gene's id as the index
if cobra.__version__ == '0.19.0':
knockout['ids'] = [list(i)[0] for i in knockout.ids]
knockout = knockout.set_index('ids')
else:
index_mapper = {i:list(i)[0] for i in knockout.index}
knockout = knockout.rename(mapper=index_mapper, axis=0)
# The output of the function single_gene_deletion is a dataframe
knockout.head()
# We define a threshold to define whether the reduction on the biomass flux is considered lethal.
threshold = 0.01
# Use this threshold to find which set of genes' knock out reduce the predicted growth below the threshold.
insilico_lethals = set(knockout.index[knockout.growth< threshold])
# The set of non-essential genes are the genes with a growth value above the threshold.
insilico_non_lethals = set(knockout.index[knockout.growth > threshold])
print("in-silico lethals:", len(insilico_lethals))
print("in-silico non lethals:", len(insilico_non_lethals))
# Now we need to experimentally verify essential and non-essential gene sets.
# Read the set of essential genes in vivo
import json
fname = './data/m9_invivo_lethals.json'
with open(fname) as fh:
invivo_lethals = json.load(fh)
invivo_lethals = set(invivo_lethals)
# Convert the list of all model genes into a set.
all_genes = set(all_genes)
# We can use the difference to obtain the list of in vivo non-lethals
invivo_non_lethals = all_genes - invivo_lethals
# Print the size of both sets
print("in-vivo lethals:", len(invivo_lethals))
print("in-vivo non lethals:", len(invivo_non_lethals))
# https://en.wikipedia.org/wiki/Receiver_operating_characteristic
# True Positives, genes predicted as essentials that are essentials in vivo (correctly predicted)
TP = insilico_lethals & invivo_lethals
# True Negatives, genes predicted as NON-essentials that are NON-essential in vivo (correctly predicted)
TN = insilico_non_lethals & invivo_non_lethals
# False Positives, wrongly predicted as NON-essential genes
FN = insilico_non_lethals & invivo_lethals
# False Positives, wrongly predicted as essential genes
FP = insilico_lethals & invivo_non_lethals
# True in vivo esssential genes
P = TP | FN
# True in vivo NON-esssential genes
N = TN | FP
len(TN)
```
### Excercise 3.3:
Complete the following table using the values from Exercise 3.2 (*E. coli*)
| In vivo \ In silico | in silico lethal | in silico non-lethal |
| -------------------------- |:----------------:| --------------------:|
| <b>in vivo lethal</b> | ? | ? |
| <b>in vivo non-lehtal</b> | ? | ? |
Total negatives = {{N}}
### Excercise 3.4:
Acces the following link:
https://en.wikipedia.org/wiki/Sensitivity_and_specificity
Get the formulas and calculate the following metrics:
* sensitivity
* specificity
* precision
* accuracy
```
# Sensitivity, recall, hit rate, or true positive rate (TPR)
# We computed the sensitivity as follows
sensitivity = len(TP) / len(P)
# TODO
# complete the following code
# Specificity, selectivity or true negative rate (TNR)
specificity = None ## COMPLETE HERE
# Precision or positive predictive value (PPV)
precision = None ## COMPLETE HERE
# Accuracy (ACC)
accuracy = None ## COMPLETE HERE
# Print the different values and discuss their meanings
```
| github_jupyter |
# Particle Filter on Episode
千葉工業大学 上田 隆一
(c) 2017 Ryuichi Ueda
This software is released under the MIT License, see LICENSE.
## はじめに
このコードは、上田が https://link.springer.com/chapter/10.1007/978-3-319-48036-7_54 で公表した「particle filter on episode」というアルゴリズムです。簡単なタスクを学習できますが、まだ弱いです。
```
%matplotlib inline
import numpy as np
from copy import copy
import math, random
import matplotlib.pyplot as plt # for plotting data
from matplotlib.patches import Ellipse # for drawing
```
## イベント、エピソードのクラス
記憶をモデル化します。行動に必要な情報は、結局のところどう動いて何を見て、どんな良いこと・悪いことがあったか、だけになります。
```
class Event:
TIME = 0
def __init__(self,action,reward,observation):
self.action = action
self.reward = reward
self.observation = observation
self.time = TIME
TIME += 1
def __str__(self):
return "action:" + str(self.action) + "\t" + "reward:" + str(self.reward) + "\t" + "observation:" + str(self.observation)
class Episodes: #強化学習では一つの試行をエピソードと呼ぶ。このクラスは複数のエピソードをずーっとシーケンシャルに持つので、Episodesと複数形に。
def __init__(self):
self.events = []
def __str__(self):
es = []
for e in self.events:
es.append(str(e))
return "\n".join(es)
```
## ロボットのクラス
環境と相互作用しながら記憶を蓄積していきます。
```
class Action:
def __init__(self,name,fw,rot):
self.name = name
self.fw = fw
self.rot = rot
class Robot:
def __init__(self):
self.episodes = Episodes()
self.actions = [Action("fw",100.0,0.0), Action("cw",0.0,-90.0), Action("ccw",0.0,90.0)]
def observe(self):
return None
def __motion(self, pos, fw, rot):
actual_fw = random.gauss(fw,fw/10) #標準偏差にしてfwの10%だけ移動距離に雑音を入れる
dir_error = random.gauss(0.0, math.pi / 180.0 * 3.0) # 前進時にロボットの進路が曲がる雑音。標準偏差3[deg]
px, py, pt = pos
#前進させる
x = px + actual_fw * math.cos(pt + dir_error)
y = py + actual_fw * math.sin(pt + dir_error)
#回転。回転角にもrotの10%だけ雑音を入れる
t = pt + dir_error + random.gauss(rot,rot/10)
return np.array([x,y,t])
def move(self,pos,action):
a = self.actions[action]
return self.__motion(pos,a.fw,a.rot)
def one_step(self,action):
pass
```
## 環境のクラス
```
class Wall:
def __init__(self,x1,y1,x2,y2):
self.p1 = np.array([x1,y1]).T
self.p2 = np.array([x2,y2]).T
class Environment:
def __init__(self,robot):
self.walls = [Wall(-90.0,0.0,90.0,0.0),
Wall(90.0,0.0,90.0,180.0), Wall(90.0,180.0,90.0,360.0),
Wall(90.0,360.0,270.0,360.0),
Wall(270.0,360.0, 270.0, 540.0),
Wall(270.0, 540.0,90.0,540.0), Wall(90.0,540.0,-90.0,540.0), Wall(-90.0,540.0,-270,540.0),
Wall(-270.0, 540.0, -270.0, 360.0),
Wall( -270.0, 360.0, -90.0, 360.0,),
Wall(-90.0,0.0,-90.0,180.0), Wall(-90.0,180.0,-90.0,360.0),
]
self.robot = robot
#the robot never knows its position by itself.
#so the pose of the robot should be here, not in a robot instance.
self.robot_pos = np.array([0,100,0])
def draw(self):
fig = plt.figure(0,figsize=(8, 8))
p = fig.add_subplot(111, aspect='equal')
p.set_xlim(-300,300)
p.set_ylim(-30,570)
for w in self.walls:
p.plot([w.p1[0],w.p2[0]],[w.p1[1],w.p2[1]],color="red")
r = self.robot_pos
p.add_artist(plt.Circle((r[0], r[1]), 25, color='black', fill=False))
p.plot([r[0],r[0] + 25.0*math.cos(r[2])],[r[1],r[1] + 25.0*math.sin(r[2])],color="black")
```
## 実行
```
env = Environment(Robot())
env.draw()
```
| github_jupyter |
# Backtesting Sentiment Pairs
<b>Summary: </b>
<br>
For rolling average and rolling standard deviation of the lenght 7 days, NLP was calcualted for all possible combinations. A trading fee was assumed 0.0075 (taken from Bitmex). Calculations show that the best pairs are Bots/Whitepaper, Announcement/Bearish, Shilling/Team, and FOMO/Whales. Not surprisingly, increasing fees and window sizes changes to outcome of top performing pairs of topics.
<br>
Furthermore, NLP of the top performers was plotted on with different windows values (up to 30). It's interesting to see that for some pairs, 7 days for rolling mean and standard deviation is not an optimal window. For example, for whales/FOMO, a window of 24 yields a much higher PNL.
<br>
Finally a heat map for various moving standard deviation and average windows sizes shows that an optimal value is concentrated within a specific area.
<br>
In conclusion, due to a very high amount of combinations of windows sizes and sentiment tags, finding the "best" pair is hard. Every pair will have its highest NLP concentrated in different areas. Changing a size of one of the windows might change results drastically.
```
import sys
sys.path.insert(0, "../src")
import example_helper as eh
import analysis_helper as ah
import msgpack
import zlib
import numpy as np
import datetime
import time
import matplotlib.pyplot as plt
import matplotlib.dates as md
from matplotlib.pyplot import figure
import pandas as pd
import seaborn as sns; sns.set()
```
# Get Data
```
# define the location of the input file
filename_augmento_topics = "../data/example_data/augmento_topics.msgpack.zlib"
filename_augmento_data = "../data/example_data/augmento_data.msgpack.zlib"
filename_bitmex_data = "../data/example_data/bitmex_data.msgpack.zlib"
# load the example data
all_data = eh.load_example_data(filename_augmento_topics,
filename_augmento_data,
filename_bitmex_data)
aug_topics, aug_topics_inv, t_aug_data, aug_data, t_price_data, price_data = all_data
all_topics = aug_data.T.astype(float)
# calculate PNL for a given strategy
# if sentiment positive go long, else go short
# fees are assumed to be 0.75% (taker fee from BITMEX)
def strategy(price_data, signal_a, signal_b, window_1 = 24 * 7, window_2 = 24*7,buy_sell_fee = 0.0075, pnl_0 = 1.0):
sent_score = ah.nb_calc_sentiment_score_b(signal_a,signal_b,window_1,window_2)
pnl = ah.nb_backtest_a(price_data, sent_score, 1.0, buy_sell_fee)
return pnl
# PNL of various moving window size for a given combination of topics
```
### Given window size 7 in rolling average and standard deviation, calculate PNL for every possible pair of strategies.
It will give 8649 NLP values calculated from 2017 until the beginning of 2019
```
# for each combination of signals, generate PNL for the last period in data
total = np.zeros(shape=(93,93))
print("calculating... might take a minute or two...")
for i in range(0,len(all_topics)):
for j in range(0,len(all_topics)):
sent_score = ah.nb_calc_sentiment_score_b(all_topics[i],all_topics[j],ra_win_size_short=24*7,ra_win_size_long=24*7)
pnl = ah.nb_backtest_a(price_data, sent_score, 1.0, buy_sell_fee=0.0075)
total[i][j] = pnl[-1]
#print("Row " + str(i+1) + " out of 93...")
print("done")
```
### Impossible to see all 8649 values
Chose top 30
```
# get all PNL in a dataframe
data = pd.DataFrame(total).rename(columns=aug_topics,index=aug_topics)
# given all combinations of signals, show the combinations that yield the highest PNL
c = data.abs()
s = c.unstack()
so_st = s.sort_values(kind="quicksort")
# specify n, a number of top combinations to be shown
t = so_st.tail(n=30).index
# labels for graphs and tables
columns_t = dict((y, x) for x, y in t).keys()
rows_t = dict((x, y) for x, y in t).keys()
# pick from the dataframes only the pairs of strategies that are within the top list
top = data[rows_t].loc[columns_t]
so_st.tail(n=20)
```
# Heat Map for top 30 pairs
```
# a sorted dataframe to get highest PNLs in the first rows
idx = pd.unique([i[1] for i in np.flip(t.values)])
col = pd.unique([i[0] for i in np.flip(t.values)])
sorted_df = data[col].loc[idx]
```
# Testing for different windows sizes
### Before the backtesting was for only one window size. It's also interesting to see how the strategy would work with different windows sizes
# Different rolling mean and std window sizes
From a chosen pair of topics, compute NLP for various rolling average and rolling std. It's interesting to see, whether the "optimal" values are concentrated withing a specific range.
### Example for 'Bots' and 'Whitepaper'
```
def window_combination(price_data,top_a,top_b,end_day_x,end_day_y,start_day_x=0,start_day_y=0,buy_sell_fee=0.0075):
total_comb = np.zeros(shape=(end_day_x,end_day_y))
print("Calculating...")
for i in range(start_day_x,end_day_x):
for j in range(start_day_y,end_day_y):
if i<j:
total_comb[i][j] = strategy(price_data,top_a,top_b,window_1=24*(i+1),window_2=24*(j+1),buy_sell_fee = 0.0075)[-1]
else:
pass
print("Done.")
return total_comb[start_day_x:end_day_x,start_day_y:end_day_y]
#specify tags
ix = 0 # specify startpoint number of rolling mean
iy = 10 # specify startpoint of rolling std
end_x = 20 # specify endpoint number of rolling mean
end_y = 30 # specify endpoint of rolling std
topic_a = 'Bots'
topic_b = 'Whitepaper'
top_b = all_topics[aug_topics_inv[topic_b]]
top_a = all_topics[aug_topics_inv[topic_a]]
total_s = window_combination(price_data,top_a,top_b,end_x,end_y,start_day_x=ix,start_day_y=iy)
# plot
cmap = sns.cubehelix_palette(50, hue=0.05, rot=0, light=0.0, dark=1.2, as_cmap=True)
figure(num=None, figsize=(10, 7), dpi=80, facecolor='w', edgecolor='k')
ax = sns.heatmap(total_s, linewidth=0.00, cmap="RdYlGn",yticklabels=np.arange(ix+1,end_x+1),xticklabels=np.arange(iy+1,end_y+1))
plt.show()
```
#### example for 'Positive' and 'Bearish'
```
#specify tags
ix = 0
iy = 0
end_x = 60
end_y = 60
topic_a = 'Positive'
topic_b = 'Bearish'
top_b = all_topics[aug_topics_inv[topic_b]]
top_a = all_topics[aug_topics_inv[topic_a]]
total_s = window_combination(price_data,top_a,top_b,end_x,end_y,start_day_x=ix,start_day_y=iy)
# plot
figure(num=None, figsize=(16, 12), dpi=80, facecolor='w', edgecolor='black')
ax = sns.heatmap(total_s, linewidth=0.00, cmap="RdYlGn",yticklabels=np.arange(ix+1,end_x+1),xticklabels=np.arange(iy+1,end_y+1))
ax.set_title('Bearish/Positive')
ax.set_ylabel('First Moving Average')
ax.set_xlabel('Second Moving Average')
plt.show()
```
### Plotted 4 points
```
# Pick Topics
aug_signal_a = aug_data[:, aug_topics_inv["Positive"]].astype(np.float64)
aug_signal_b = aug_data[:, aug_topics_inv["Bearish"]].astype(np.float64)
# generate the sentiment score
sent_score = ah.nb_calc_sentiment_score_b(aug_signal_a, aug_signal_b, 7*24, 39*24)
sent_score1 = ah.nb_calc_sentiment_score_b(aug_signal_a, aug_signal_b, 18*24, 38*24)
sent_score2 = ah.nb_calc_sentiment_score_b(aug_signal_a, aug_signal_b, 18*24, 39*24)
sent_score3 = ah.nb_calc_sentiment_score_b(aug_signal_a, aug_signal_b, 7*24, 40*24)
# define some parameters for the backtest
start_pnl = 1.0
buy_sell_fee = 0.0075
# run the backtest
pnl = ah.nb_backtest_a(price_data, sent_score, start_pnl, buy_sell_fee)
pnl1 = ah.nb_backtest_a(price_data, sent_score1, start_pnl, buy_sell_fee)
pnl2 = ah.nb_backtest_a(price_data, sent_score2, start_pnl, buy_sell_fee)
pnl3 = ah.nb_backtest_a(price_data, sent_score3, start_pnl, buy_sell_fee)
# set up the figure
fig, ax = plt.subplots(2, 1, sharex=True, sharey=False, figsize=(20,10))
# initialise some labels for the plot
datenum_aug_data = [md.date2num(datetime.datetime.fromtimestamp(el)) for el in t_aug_data]
datenum_price_data = [md.date2num(datetime.datetime.fromtimestamp(el)) for el in t_price_data]
# plot stuff
ax[0].grid(linewidth=0.4)
ax[1].grid(linewidth=0.4)
ax[0].plot(datenum_price_data, price_data, linewidth=0.5)
ax[1].plot(datenum_aug_data, pnl, linewidth=0.5)
ax[1].plot(datenum_price_data, pnl1, linewidth=0.5)
ax[1].plot(datenum_price_data, pnl2, linewidth=0.5)
ax[1].plot(datenum_price_data, pnl3, linewidth=0.5)
ax[1].legend(("A","B","C","D"))
# label axes
ax[0].set_ylabel("Price")
ax[1].set_ylabel("PnL")
ax[0].set_title("Profit and Loss.py")
# generate the time axes
plt.subplots_adjust(bottom=0.2)
plt.xticks( rotation=25 )
ax[0]=plt.gca()
xfmt = md.DateFormatter('%Y-%m-%d')
ax[0].xaxis.set_major_formatter(xfmt)
# show the plot
plt.show()
```
| github_jupyter |
# Hybrid quantum-classical Neural Networks with PyTorch and Qiskit
Machine learning (ML) has established itself as a successful interdisciplinary field which seeks to mathematically extract generalizable information from data. Throwing in quantum computing gives rise to interesting areas of research which seek to leverage the principles of quantum mechanics to augment machine learning or vice-versa. Whether you're aiming to enhance classical ML algorithms by outsourcing difficult calculations to a quantum computer or optimise quantum algorithms using classical ML architectures - both fall under the diverse umbrella of quantum machine learning (QML).
In this chapter, we explore how a classical neaural network can be partially quantized to create a hybrid quantum-classical neural network. We will code up a simple example which integrates qiskit with a state-of-the-art open source software package - [PyTorch](https://pytorch.org/). The purpose of this example is to demonstrate the ease of integrating Qiskit with existing ML tools and to encourage ML practitioners to explore what is possible with quantum computing.
## How does it work?
<img src="hybridnetwork.png" width="800"/>
**Fig.1** Illustrates the framework we will construct in this chapter. Ultimately, we will create a hybrid quantum-classical neural network that seeks to classify hand drawn digits. Note that the edges shown in this image are all directed downward; however, the directionality is not visually indicated.
### Preliminaries:
The background presented here on classical neural networks is included to establish relevant ideas and shared terminology; however, it is still extremely high-level. __If you'd like to dive one step deeper into classical neural networks, see the well made video series by youtuber__ [3Blue1Brown](https://youtu.be/aircAruvnKk). Alternatively, if you are already familiar with classical networks, you can [skip to the next section](#quantumlayer).
###### Neurons and Weights
A neural network is ultimately just an elaborate function that is built by composing smaller building blocks called neurons. A ***neuron*** is typically a simple, easy-to-compute, and nonlinear function that maps one or more inputs to a single real number. The single output of a neuron is typically copied and fed as input into other neurons. Graphically, we represent neurons as nodes in a graph and we draw directed edges between nodes to indicate how the output of one neuron will be used as input to other neurons. It's also important to note that each edge in our graph is often associated with a scalar-value called a [***weight***](https://en.wikipedia.org/wiki/Artificial_neural_network#Connections_and_weights). The idea here is that each of the inputs to a neuron will be multiplied by a different scalar before being collected and processed into a single value. The objective when training a neural network consists primarily of choosing our weights such that the network behaves in a particular way.
###### Feed Forward Neural Networks
It is also worth noting that the particular type of neural network we will concern ourselves with is called a **[feed-forward neural network (FFNN)](https://en.wikipedia.org/wiki/Feedforward_neural_network)**. This means that as data flows through our neural network, it will never return to a neuron it has already visited. Equivalently, you could say that the graph which describes our neural network is a **[directed acyclic graph (DAG)](https://en.wikipedia.org/wiki/Directed_acyclic_graph)**. Furthermore, we will stipulate that neurons within the same layer of our neural network will not have edges between them.
###### IO Structure of Layers
The input to a neural network is a classical (real-valued) vector. Each component of the input vector is multiplied by a different weight and fed into a layer of neurons according to the graph structure of the network. After each neuron in the layer has been evaluated, the results are collected into a new vector where the i'th component records the output of the i'th neuron. This new vector can then treated as input for a new layer, and so on. We will use the standard term ***hidden layer*** to describe all but the first and last layers of our network.
## So how does quantum enter the picture? <a id='quantumlayer'> </a>
To create a quantum-classical neural network, one can implement a hidden layer for our neural network using a parameterized quantum circuit. By "parameterized quantum circuit", we mean a quantum circuit where the rotation angles for each gate are specified by the components of a classical input vector. The outputs from our neural network's previous layer will be collected and used as the inputs for our parameterized circuit. The measurement statistics of our quantum circuit can then be collected and used as inputs for the following layer. A simple example is depicted below:
<img src="neuralnetworkQC.png" width="800"/>
Here, $\sigma$ is a [nonlinear function](https://en.wikipedia.org/wiki/Activation_function) and $h_i$ is the value of neuron $i$ at each hidden layer. $R(h_i)$ represents any rotation gate about an angle equal to $h_i$ and $y$ is the final prediction value generated from the hybrid network.
### What about backpropagation?
If you're familiar with classical ML, you may immediately be wondering *how do we calculate gradients when quantum circuits are involved?* This would be necessary to enlist powerful optimisation techniques such as **[gradient descent](https://en.wikipedia.org/wiki/Gradient_descent)**. It gets a bit technical, but in short, we can view a quantum circuit as a black box and the gradient of this black box with respect to its parameters can be calculated as follows:
<img src="quantumgradient.png" width="800"/>
where $\theta$ represents the parameters of the quantum circuit and $s$ is a macroscopic shift. The gradient is then simply the difference between our quantum circuit evaluated at $\theta+s$ and $\theta - s$. Thus, we can systematically differentiate our quantum circuit as part of a larger backpropogation routine. This closed form rule for calculating the gradient of quantum circuit parameters is known as **[the parameter shift rule](https://arxiv.org/pdf/1905.13311.pdf)**.
# Let's code!
### Imports
First, we import some handy packages that we will need, including Qiskit and PyTorch.
```
import numpy as np
import torch
from torch.autograd import Function
import torch.optim as optim
import torch.nn as nn
import torch.nn.functional as F
import torchvision
from torchvision import datasets, transforms
from qiskit import QuantumRegister, QuantumCircuit, ClassicalRegister, execute
from qiskit.circuit import Parameter
from qiskit import Aer
from tqdm import tqdm
from matplotlib import pyplot as plt
%matplotlib inline
```
### Tensors to lists
Next we create an additional function that converts a tensor to a list in Python. This is needed to connect Qiskit and PyTorch objects. In particular, we will use this function to convert tensors produced by PyTorch to a list, such that they can be fed into quantum circuits in Qiskit.
```
def to_numbers(tensor_list):
num_list = []
for tensor in tensor_list:
num_list += [tensor.item()]
return num_list
```
### Create a "quantum class" with Qiskit
We can conveniently put our Qiskit quantum functions into a class. First, we specify how many trainable quantum parameters and how many shots we wish to use in our quantum circuit. In this example, we will keep it simple and use a 1-qubit circuit with one trainable quantum parameter $\theta$. We hard code the circuit for simplicity and use a $RY-$rotation by the angle $\theta$ to train the output of our circuit. The circuit looks like this:
<img src="1qubitcirc.png" width="400"/>
In order to measure the output in the $z-$basis, we create a Python function to obtain the $\sigma_z$ expectation. Lastly, we create a "bind" function to convert our parameter to a list and run the circuit on the Aer simulator. We will see later how this all ties into the hybrid neural network.
```
class QiskitCircuit():
# Specify initial parameters and the quantum circuit
def __init__(self,shots):
self.theta = Parameter('Theta')
self.shots = shots
def create_circuit():
qr = QuantumRegister(1,'q')
cr = ClassicalRegister(1,'c')
ckt = QuantumCircuit(qr,cr)
ckt.h(qr[0])
ckt.barrier()
ckt.ry(self.theta,qr[0])
ckt.barrier()
ckt.measure(qr,cr)
return ckt
self.circuit = create_circuit()
def N_qubit_expectation_Z(self, counts, shots, nr_qubits):
expects = np.zeros(nr_qubits)
for key in counts.keys():
perc = counts[key]/shots
check = np.array([(float(key[i])-1/2)*2*perc for i in range(nr_qubits)])
expects += check
return expects
def bind(self, parameters):
[self.theta] = to_numbers(parameters)
self.circuit.data[2][0]._params = to_numbers(parameters)
def run(self, i):
self.bind(i)
backend = Aer.get_backend('qasm_simulator')
job_sim = execute(self.circuit,backend,shots=self.shots)
result_sim = job_sim.result()
counts = result_sim.get_counts(self.circuit)
return self.N_qubit_expectation_Z(counts,self.shots,1)
```
### Create a "quantum-classical class" with PyTorch
Now that our quantum circuit is defined, we can create the functions needed for backpropagation using PyTorch. [The forward and backward passes](http://www.ai.mit.edu/courses/6.034b/backprops.pdf) contain elements from our Qiskit class. The backward pass directly computes the analytical gradients using the finite difference formula we introduced above.
```
class TorchCircuit(Function):
@staticmethod
def forward(ctx, i):
if not hasattr(ctx, 'QiskitCirc'):
ctx.QiskitCirc = QiskitCircuit(shots=100)
exp_value = ctx.QiskitCirc.run(i[0])
result = torch.tensor([exp_value]) # store the result as a torch tensor
ctx.save_for_backward(result, i)
return result
@staticmethod
def backward(ctx, grad_output):
s = np.pi/2
forward_tensor, i = ctx.saved_tensors
# Obtain paramaters
input_numbers = to_numbers(i[0])
gradient = []
for k in range(len(input_numbers)):
input_plus_s = input_numbers
input_plus_s[k] = input_numbers[k] + s # Shift up by s
exp_value_plus = ctx.QiskitCirc.run(torch.tensor(input_plus_s))[0]
result_plus_s = torch.tensor([exp_value_plus])
input_minus_s = input_numbers
input_minus_s[k] = input_numbers[k] - s # Shift down by s
exp_value_minus = ctx.QiskitCirc.run(torch.tensor(input_minus_s))[0]
result_minus_s = torch.tensor([exp_value_minus])
gradient_result = (result_plus_s - result_minus_s)
gradient.append(gradient_result)
result = torch.tensor([gradient])
return result.float() * grad_output.float()
```
Putting this all together
We will create a simple hybrid neural network to classify images of two types of digits (0 or 1) from the [MNIST dataset](http://yann.lecun.com/exdb/mnist/). We first load MNIST and filter for pictures containing 0's and 1's. These will serve as inputs for our neural network to classify.
### Data loading and preprocessing
```
transform = torchvision.transforms.Compose([torchvision.transforms.ToTensor()]) # transform images to tensors/vectors
mnist_trainset = datasets.MNIST(root='./data', train=True, download=True, transform=transform)
labels = mnist_trainset.targets # get the labels for the data
labels = labels.numpy()
idx1 = np.where(labels == 0) # filter on zeros
idx2 = np.where(labels == 1) # filter on ones
# Specify number of datapoints per class (i.e. there will be n pictures of 1 and n pictures of 0 in the training set)
n=100
# concatenate the data indices
idx = np.concatenate((idx1[0][0:n],idx2[0][0:n]))
# create the filtered dataset for our training set
mnist_trainset.targets = labels[idx]
mnist_trainset.data = mnist_trainset.data[idx]
train_loader = torch.utils.data.DataLoader(mnist_trainset, batch_size=1, shuffle=True)
```
The data will consist of images belonging to two classes: 0 and 1. An example image from both classes looks like this:
<img src="MNISTplot.png" width="100"/>
So far, we have loaded the data and coded a class that creates our quantum circuit which contains 1 trainable parameter. This quantum parameter will be inserted into a classical neural network along with the other classical parameters to form the hybrid neural network. We also created backward and forward pass functions that allow us to do backpropagation and optimise our neural network. Lastly, we need to specify our neural network architecture such that we can begin to train our parameters using optimisation techniques provided by PyTorch.
### Creating the hybrid neural network
We can use a neat PyTorch pipeline to create a neural network architecture. The network will need to be compatible in terms of its dimensionality when we insert the quantum layer (i.e. our quantum circuit). Since our quantum in this example contains 1 parameter, we must ensure the network condenses neurons down to size 1. We create a network consisting of 3 hidden layers with 320, 50 and 1 neurons respectively. The value of the last neuron is fed as the parameter $\theta$ into our quantum circuit. The circuit measurement then serves as the final prediction for 0 or 1 as provided by a $\sigma_z$ measurement. The measurement outcomes are -1 which implies a predicted label of 0 and 1 which implies a predicted label of 1.
```
qc = TorchCircuit.apply
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 10, kernel_size=5)
self.conv2 = nn.Conv2d(10, 20, kernel_size=5)
self.conv2_drop = nn.Dropout2d()
self.h1 = nn.Linear(320, 50)
self.h2 = nn.Linear(50, 1)
def forward(self, x):
x = F.relu(F.max_pool2d(self.conv1(x), 2))
x = F.relu(F.max_pool2d(self.conv2_drop(self.conv2(x)), 2))
x = x.view(-1, 320)
x = F.relu(self.h1(x))
x = F.dropout(x, training=self.training)
x = self.h2(x)
x = qc(x)
x = (x+1)/2 # Normalise the inputs to 1 or 0
x = torch.cat((x, 1-x), -1)
return x
```
### Training the network
We now have all the ingredients to train our hybrid network! We can specify any [PyTorch optimiser](https://pytorch.org/docs/stable/optim.html), [learning rate](https://en.wikipedia.org/wiki/Learning_rate) and [cost/loss function](https://en.wikipedia.org/wiki/Loss_function) in order to train over multiple epochs. In this instance, we use the [Adam optimiser](https://arxiv.org/abs/1412.6980), a learning rate of 0.001 and the [negative log-likelihood loss function](https://pytorch.org/docs/stable/_modules/torch/nn/modules/loss.html).
```
network = Net()
optimizer = optim.Adam(network.parameters(), lr=0.001)
epochs = 30
loss_list = []
for epoch in range(epochs):
total_loss = []
target_list = []
for batch_idx, (data, target) in enumerate(train_loader):
target_list.append(target.item())
optimizer.zero_grad()
output = network(data)
loss = F.nll_loss(output, target)
loss.backward()
optimizer.step()
total_loss.append(loss.item())
loss_list.append(sum(total_loss)/len(total_loss))
print(loss_list[-1])
# Normalise the loss between 0 and 1
for i in range(len(loss_list)):
loss_list[i] += 1
# Plot the loss per epoch
plt.plot(loss_list)
plt.title("Hybrid NN Training Convergence")
plt.xlabel('Training Iterations')
plt.ylabel('Loss')
```
# What now?
While it is totally possible to create hybrid neural networks, does this actually have any benefit? In fact, the classical layers of this network train perfectly fine (in fact, better) without the quantum layer. Furthermore, you may have noticed that the quantum layer we trained here **generates no entanglement**, and will therefore continue to be classically simulable as we scale up this particular architecture. This means that if you hope to achieve a quantum advantage using hybrid nerural networks, you'll need to start by extending this code to include a more sophisticated quantum layer.
The point of this exercise was to get you thinking about integrating techniques from ML and quantum computing in order to investigate if there is indeed some element of interest - and thanks to PyTorch and Qiskit, this becomes a little bit easier.
| github_jupyter |
```
!curl -o datasets 'https://ftp.ncbi.nlm.nih.gov/pub/datasets/command-line/LATEST/linux-amd64/datasets'
!curl -o dataformat 'https://ftp.ncbi.nlm.nih.gov/pub/datasets/command-line/LATEST/linux-amd64/dataformat'
!chmod +x datasets
!./datasets --help
!./datasets version
!./datasets download virus genome taxon SARS-CoV-2 --filename SARS2-all.zip
corona=""" gatttaagtg aatagcttgg ctatctcact tcccctcgtt ctcttgcaga actttgattt
taacgaactt aaataaaagc cctgttgttt agcgtattgt tgcacttgtc tggtgggatt
gtggcattaa tttgcctgct catctaggca gtggacatat gctcaacact gggtataatt
181 ctaattgaat actatttttc agttagagcg tcgtgtctct tgtacgtctc ggtcacaata
241 cacggtttcg tccggtgcgt ggcaattcgg ggcacatcat gtctttcgtg gctggtgtga
301 ccgcgcaagg tgcgcgcggt acgtatcgag cagcgctcaa ctctgaaaaa catcaagacc
361 atgtgtctct aactgtgcca ctctgtggtt caggaaacct ggttgaaaaa ctttcaccat
421 ggttcatgga tggcgaaaat gcctatgaag tggtgaaggc catgttactt aaaaaggagc
481 cacttctcta tgtgcccatc cggctggctg gacacactag acacctccca ggtcctcgtg
541 tatacctggt tgagaggctc attgcttgtg aaaatccatt catggttaac caattggctt
601 atagctctag tgcaaatggc agcttggttg gcacaatttt gcagggcaag cctattggta
661 tgttcttccc ttatgacatc gaacttgtca caggaaagca aaatattctc ctgcgcaagt
721 atggccgtgg tggttaccac tacaccccat tccactatga gcgagacaac acctcttgcc
781 ctgagtggat ggacgatttt gaggcggatc ctaaaggcaa atatgcccag aatctgctta
841 agaagttgat tggcggtgat gtcactccag ttgaccaata catgtgtggc gttgatggaa
901 aacccattag tgcctacgca tttttaatgg ccaaggatgg aataaccaaa ctggctgatg
961 ttgaagcgga cgtcgcagca cgtgctgatg acgaaggctt catcacatta aagaacaatc
1021 tatatagatt ggtttggcat gttgagcgta aagacgttcc atatcctaag caatctattt
1081 ttactattaa tagtgtggtc caaaaggatg gtgttgaaaa cactcctcct cactatttta
1141 ctcttggatg caaaatttta acgctcaccc cacgcaacaa gtggagtggc gtttctgact
1201 tgtccctcaa acaaaaactc ctttacacct tctatggtaa ggagtcactt gagaacccaa
1261 cctacattta ccactccgca ttcattgagt gtggaagttg tggtaatgat tcctggctta
1321 cagggaatgc tatccaaggg tttgcctgtg gatgtggggc atcatataca gctaatgatg
1381 tcgaagtcca atcatctggc atgattaagc caaatgctct tctttgtgct acttgcccct
1441 ttgctaaggg tgatagctgt tcttctaatt gcaaacattc agttgctcag ttggttagtt
1501 acctttctga acgttgtaat gttattgctg attctaagtc cttcacactt atctttggtg
1561 gcgtagctta cgcctacttt ggatgtgagg aaggtactat gtactttgtg cctagagcta
1621 agtctgttgt ctcaaggatt ggagactcca tctttacagg ctgtactggc tcttggaaca
1681 aggtcactca aattgctaac atgttcttgg aacagactca gcattccctt aactttgtgg
1741 gagagttcgt tgtcaacgat gttgtcctcg caattctctc tggaaccaca actaatgttg
1801 acaaaatacg ccagcttctc aaaggtgtca cccttgacaa gttgcgtgat tatttagctg
1861 actatgacgt agcagtcact gccggcccat tcatggataa tgctattaat gttggtggta
1921 caggattaca gtatgccgcc attactgcac cttatgtagt tctcactggc ttaggtgagt
1981 cctttaagaa agttgcaacc ataccgtaca aggtttgcaa ctctgttaag gatactctga
2041 cttattatgc tcacagcgtg ttgtacagag tttttcctta tgacatggat tctggtgtgt
2101 catcctttag tgaactactt tttgattgcg ttgatctttc agtagcttct acctattttt
2161 tagtccgcct cttgcaagat aagactggcg actttatgtc tacaattatt acttcctgcc
2221 aaactgctgt tagtaagctt ctagatacat gttttgaagc tacagaagca acatttaact
2281 tcttgttaga tttggcagga ttgttcagaa tctttcttcg caatgcctat gtgtacactt
2341 cacaagggtt tgtggtggtc aatggcaaag tttctacact tgtcaaacaa gtgttagact
2401 tgcttaataa gggtatgcaa cttttgcata caaaggtctc ctgggctggt tctaatatca
2461 gtgctgttat ctacagcggc agggagtctt taatattccc atcgggaacc tattactgtg
2521 tcaccactaa ggctaagtcc gttcaacaag atcttgacgt tattttgcct ggtgagtttt
2581 ccaagaagca gttaggactg ctccaaccta ctgacaattc tacaactgtt agtgttactg
2641 tatccagtaa catggttgaa actgttgtgg gtcaacttga gcaaactaat atgcatagtc
2701 ctgatgttat agtaggtgac tatgtcatta ttagtgaaaa attgtttgtg cgtagtaagg
2761 aagaagacgg atttgccttc taccctgctt gcactaatgg tcatgctgta ccgactctct
2821 ttagacttaa gggaggtgca cctgtaaaaa aagtagcctt tggcggtgat caagtacatg
2881 aggttgctgc tgtaagaagt gttactgtcg agtacaacat tcatgctgta ttagacacac
2941 tacttgcttc ttctagtctt agaacctttg ttgtagataa gtctttgtca attgaggagt
3001 ttgctgacgt agtaaaggaa caagtctcag acttgcttgt taaattactg cgtggaatgc
3061 cgattccaga ttttgattta gacgatttta ttgacgcacc atgctattgc tttaacgctg
3121 agggtgatgc atcttggtct tctactatga tcttctctct tcaccccgtc gagtgtgacg
3181 aggagtgttc tgaagtagag gcttcagatt tagaagaagg tgaatcagag tgcatttctg
3241 agacttcaac tgaacaagtt gacgtttctc atgagatttc tgacgacgag tgggctgctg
3301 cagttgatga agcgttccct ctcgatgaag cagaagatgt tactgaatct gtgcaagaag
3361 aagcacaacc agtagaagta cctgttgaag atattgcgca ggttgtcata gctgacacct
3421 tacaggaaac tcctgttgtg tctgatactg ttgaagtccc accgcaagtg gtgaaacttc
3481 cgtctgaacc tcagactatc cagcccgagg taaaagaagt tgcacctgtc tatgaggctg
3541 ataccgaaca gacacagagt gttactgtta aacctaagag gttacgcaaa aagcgtaatg
3601 ttgacccttt gtccaatttt gaacataagg ttattacaga gtgcgttacc atagttttag
3661 gtgacgcaat tcaagtagcc aagtgctatg gtgagtctgt gttagttaat gctgctaaca
3721 cacatcttaa gcatggcggt ggtatcgctg gtgctattaa tgcggcttca aaaggggctg
3781 tccaaaaaga gtcagatgag tatattctgg ctaaagggcc gttacaagta ggagattcag
3841 ttctcttgca aggccattct ctagctaaga atatcctgca tgtcgtaggc ccagatgccc
3901 gcgctaaaca ggatgtttct ctccttagta agtgctataa ggctatgaat gcatatcctc
3961 ttgtagtcac tcctcttgtt tcagcaggca tatttggtgt aaaaccagct gtgtcttttg
4021 attatcttat tagagaggct aagactagag ttttagtcgt cgttaattcc caagatgtct
4081 ataagagtct taccatagtt gacattccac agagtttgac tttttcatat gatgggttac
4141 gtggcgcaat acgtaaagct aaagattatg gttttactgt ttttgtgtgc acagacaact
4201 ctgctaacac taaagttctt aggaacaagg gtgttgatta tactaagaag tttcttacag
4261 ttgacggtgt gcaatattat tgctacacgt ctaaggacac tttagatgat atcttacaac
4321 aggctaataa gtctgttggt attatatcta tgcctttggg atatgtgtct catggtttag
4381 acttaattca agcagggagt gtcgtgcgta gagttaacgt gccctacgtg tgtctcctag
4441 ctaataaaga gcaagaagct attttgatgt ctgaagacgt taagttaaac ccttcagaag
4501 attttataaa gcacgtccgc actaatggtg gttacaattc ttggcattta gtcgagggtg
4561 aactattggt gcaagactta cgcttaaata agctcctgca ttggtctgat caaaccatat
4621 gctacaagga tagtgtgttt tatgttgtaa agaatagtac agcttttcca tttgaaacac
4681 tttcagcatg tcgtgcgtat ttggattcac gcacgacaca gcagttaaca atcgaagtct
4741 tagtgactgt cgatggtgta aattttagaa cagtcgttct aaataataag aacacttata
4801 gatcacagct tggatgcgtt ttctttaatg gtgctgatat ttctgatacc attcctgatg
4861 agaaacagaa tggtcacagt ttatatctag cagacaattt gactgctgat gaaacaaagg
4921 cgcttaaaga gttatatggc cccgttgatc ctactttctt acacagattc tattcactta
4981 aggctgcagt ccataagtgg aagatggttg tgtgtgataa ggtacgttct ctcaaattga
5041 gtgataataa ttgttatctt aatgcagtta ttatgacact tgatttattg aaggacatta
5101 aatttgttat acctgctcta cagcatgcat ttatgaaaca taagggcggt gattcaactg
5161 acttcatagc cctcattatg gcttatggca attgcacatt tggtgctcca gatgatgcct
5221 ctcggttact tcataccgtg cttgcaaagg ctgagttatg ctgttctgca cgcatggttt
5281 ggagagagtg gtgcaatgtc tgtggcataa aagatgttgt tctacaaggc ttaaaagctt
5341 gttgttacgt gggtgtgcaa actgttgaag atctgcgtgc tcgcatgaca tatgtatgcc
5401 agtgtggtgg tgaacgtcat cggcaaatag tcgaacacac caccccctgg ttgctgctct
5461 caggcacacc aaatgaaaaa ttggtgacaa cctccacggc gcctgatttt gtagcgttta
5521 atgtctttca gggcattgaa acggctgttg gccattatgt tcatgctcgc ctgaagggtg
5581 gtcttatttt aaagtttgac tctggcaccg ttagcaagac ttcagactgg aagtgcaagg
5641 tgacagatgt acttttctcc ggccaaaaat acagtagcga ttgtaatgtc gtacggtatt
5701 ctttggacgg taatttcaga acagaggttg atcccgacct atctgctttc tatgttaagg
5761 atggtaaata ctttacaagt gaaccacccg taacatattc accagctaca attttagctg
5821 gtagtgtcta cactaatagc tgccttgtat cgtctgatgg acaacctggc ggtgatgcta
5881 ttagtttgag ttttaataac cttttagggt ttgattctag taaaccagtc actaagaaat
5941 acacttactc cttcttgcct aaagaagacg gcgatgtgtt gttggctgag tttgacactt
6001 atgaccctat ttataagaat ggtgccatgt ataaaggcaa accaattctt tgggtcaaca
6061 aagcatctta tgatactaat cttaataagt tcaatagagc tagtttgcgt caaatttttg
6121 acgtagcccc cattgaactc gaaaataaat tcacaccttt gagtgtggag tctacaccag
6181 ttgaacctcc aactgtagat gtggtagcac ttcaacagga aatgacaatt gtcaaatgta
6241 agggtttaaa taaacctttc gtgaaggaca atgtcagttt cgttgctgat gactcaggta
6301 ctcccgttgt tgagtatctg tctaaagaag atctacatac attgtatgta gaccctaagt
6361 atcaagtcat tgtcttaaaa gacaatgtac tttcttctat gcttagattg cacaccgttg
6421 agtcaggtga tattaacgtt gttgcagctt ccggatcttt gacacgtaaa gtgaaattac
6481 tatttagggc ttcattttat ttcaaagaat ttgctacccg cactttcact gctaccactg
6541 ctgtaggtag ttgtataaag agtgtagtgc ggcatctagg tgttactaaa ggcatattga
6601 caggctgttt tagttttgtc aagatgttat ttatgcttcc actagcttac tttagtgatt
6661 caaaactcgg caccacagag gttaaagtga gtgctttgaa aacagctggc gttgtgacag
6721 gtaatgttgt aaaacagtgt tgcactgctg ctgttgattt aagtatggat aagttgcgcc
6781 gtgtggattg gaaatcaacc ctacggttgt tacttatgtt atgcacaact atggtattgt
6841 tgtcttctgt gtatcacttg tatgtcttca atcaggtctt atcaagtgat gttatgtttg
6901 aagatgccca aggtttgaaa aagttctaca aagaagttag agcttaccta ggaatctctt
6961 ctgcttgtga cggtcttgct tcagcttata gggcgaattc ctttgatgta cctacattct
7021 gcgcaaaccg ttctgcaatg tgtaattggt gcttgattag ccaagattcc ataactcact
7081 acccagctct taagatggtt caaacacatc ttagccacta tgttcttaac atagattggt
7141 tgtggtttgc atttgagact ggtttggcat acatgctcta tacctcggcc ttcaactggt
7201 tgttgttggc aggtacattg cattatttct ttgcacagac ttccatattt gtagactggc
7261 ggtcatacaa ttatgctgtg tctagtgcct tctggttatt cacccacatt ccaatggcgg
7321 gtttggtacg aatgtataat ttgttagcat gcctttggct tttacgcaag ttttatcagc
7381 atgtaatcaa tggttgcaaa gatacggcat gcttgctctg ctataagagg aaccgactta
7441 ctagagttga agcttctacc gttgtctgtg gtggaaaacg tacgttttat atcacagcaa
7501 atggcggtat ttcattctgt cgtaggcata attggaattg tgtggattgt gacactgcag
7561 gtgtggggaa taccttcatc tgtgaagaag tcgcaaatga cctcactacc gccctacgca
7621 ggcctattaa cgctacggat agatcacatt attatgtgga ttccgttaca gttaaagaga
7681 ctgttgttca gtttaattat cgtagagacg gtcaaccatt ctacgagcgg tttcccctct
7741 gcgcttttac aaatctagat aagttgaagt tcaaagaggt ctgtaaaact actactggta
7801 tacctgaata caactttatc atctacgact catcagatcg tggccaggaa agtttagcta
7861 ggtctgcatg tgtttattat tctcaagtct tgtgtaaatc aattcttttg gttgactcaa
7921 gtttggttac ttctgttggt gattctagtg aaatcgccac taaaatgttt gattcctttg
7981 ttaatagttt cgtctcgctg tataatgtca cacgcgataa gttggaaaaa cttatctcta
8041 ctgctcgtga tggcgtaagg cgaggcgata acttccatag tgtcttaaca acattcattg
8101 acgcagcacg aggccccgca ggtgtggagt ctgatgttga gaccaatgaa attgttgact
8161 ctgtgcagta tgctcataaa catgacatac aaattactaa tgagagttac aataattatg
8221 taccctcata tgttaaacct gatagtgtgt ctaccagtga tttaggtagt ctcattgatt
8281 gtaatgcggc ttcagttaac caaattgtct tgcgtaattc taatggtgct tgtatttgga
8341 acgctgctgc atatatgaaa ctctcggatg cacttaaacg acagattcgc attgcatgcc
8401 gtaagtgtaa tttagctttc cggttaacca cctcaaagct acgcgctaat gataatatct
8461 tatcagttag attcactgct aacaaaattg ttggtggtgc tcctacatgg tttaatgcgt
8521 tgcgtgactt tacgttaaag ggttacgttc ttgctaccat tattgtgttt ctgtgtgctg
8581 tactgatgta tttgtgttta cctacatttt ctatggtacc tgttgaattt tatgaagacc
8641 gcatcttgga ctttaaagtt cttgataatg gtatcattag ggatgtaaat cctgatgata
8701 agtgctttgc taataagcac cggtccttca cacaatggta tcatgagcat gttggtggtg
8761 tctatgacaa ctctatcaca tgcccattga cagttgcagt aattgctgga gttgctggtg
8821 ctcgcattcc agacgtacct actacattgg cttgggtgaa caatcagata attttctttg
8881 tttctcgagt ctttgctaat acaggcagtg tttgctacac tcctatagat gagataccct
8941 ataagagttt ctctgatagt ggttgcattc ttccatctga gtgcactatg tttagggatg
9001 cagagggccg tatgacacca tactgccatg atcctactgt tttgcctggg gcttttgcgt
9061 acagtcagat gaggcctcat gttcgttacg acttgtatga tggtaacatg tttattaaat
9121 ttcctgaagt agtatttgaa agtacactta ggattactag aactctgtca actcagtact
9181 gccggttcgg tagttgtgag tatgcacaag agggtgtttg tattaccaca aatggctcgt
9241 gggccatttt taatgaccac catcttaata gacctggtgt ctattgtggc tctgatttta
9301 ttgacattgt caggcggtta gcagtatcac tgttccagcc tattacttat ttccaattga
9361 ctacctcatt ggtcttgggt ataggtttgt gtgcgttcct gactttgctc ttctattata
9421 ttaataaagt aaaacgtgct tttgcagatt acacccagtg tgctgtaatt gctgttgttg
9481 ctgctgttct taatagcttg tgcatctgct ttgttgcctc tataccattg tgtatagtac
9541 cttacactgc attgtactat tatgctacat tctattttac taatgagcct gcatttatta
9601 tgcatgtttc ttggtacatt atgttcgggc ctatcgttcc catatggatg acctgcgttt
9661 atacagttgc aatgtgcttt agacacttct tctgggtttt agcttatttt agtaagaaac
9721 atgtagaagt ttttactgat ggtaagctta attgtagttt ccaggacgct gcctctaata
9781 tctttgttat taacaaggac acttatgcag ctcttagaaa ctctttaact aatgatgcct
9841 attcacgatt tttggggttg tttaacaagt ataagtactt ctctggtgct atggaaacag
9901 ccgcttatcg tgaagctgca gcatgtcatc ttgctaaagc cttacaaaca tacagcgaga
9961 ctggtagtga tcttctttac caaccaccca actgtagcat aacctctggc gtgttgcaaa
10021 gcggtttggt gaaaatgtca catcccagtg gagatgttga ggcttgtatg gttcaggtta
10081 cctgcggtag catgactctt aatggtcttt ggcttgacaa cacagtctgg tgcccacgac
10141 acgtaatgtg cccggctgac cagttgtctg atcctaatta tgatgccttg ttgatttcta
10201 tgactaatca tagtttcagt gtgcaaaaac acattggcgc tccagcaaac ttgcgtgttg
10261 ttggtcatgc catgcaaggc actcttttga agttgactgt cgatgttgct aaccctagca
10321 ctccagccta cacttttaca acagtgaaac ctggcgcagc atttagtgtg ttagcatgct
10381 ataatggtcg tccgactggt acattcactg ttgtaatgcg ccctaactac acaattaagg
10441 gttcctttct gtgtggttct tgtggtagtg ttggttacac caaggagggt agtgtgatca
10501 atttttgtta catgcatcaa atggaacttg ctaatggtac acataccggt tcagcatttg
10561 atggtactat gtatggtgcc tttatggata aacaagtgca ccaagttcag ttaacagaca
10621 aatactgcag tgttaatgta gtagcttggc tttacgcagc aatacttaat ggttgcgctt
10681 ggtttgtaaa acctaatcgc actagtgttg tttcttttaa tgaatgggct cttgccaacc
10741 aattcactga atttgttggc actcaatccg ttgacatgtt agctgtcaaa acaggcgttg
10801 ctattgaaca gctgctttat gcgatccaac aactttatac tgggttccag ggaaagcaaa
10861 tccttggcag tactatgttg gaagatgaat tcacacctga ggatgttaat atgcagatta
10921 tgggtgtggt tatgcagagt ggtgtgagaa aagttacata tggtactgcg cattggttgt
10981 tcgcgaccct tgtctcaacc tatgtgataa tcttacaagc cactaaattt actttgtgga
11041 actacttgtt tgagactatt cccacacagt tgttcccact cttatttgtg actatggcct
11101 tcgttatgtt gttggttaaa cacaaacaca cctttttgac acttttcttg ttgcctgtgg
11161 ctatttgttt gacttatgca aacatagtct acgagcccac tactcccatt tcgtcagcgc
11221 tgattgcagt tgcaaattgg cttgccccca ctaatgctta tatgcgcact acacatactg
11281 atattggtgt ctacattagt atgtcacttg tattagtcat tgtagtgaag agattgtaca
11341 acccatcact ttctaacttt gcgttagcat tgtgcagtgg tgtaatgtgg ttgtacactt
11401 atagcattgg agaagcctca agccccattg cctatctggt ttttgtcact acactcacta
11461 gtgattatac gattacagtc tttgttactg tcaaccttgc aaaagtttgc acttatgcca
11521 tctttgctta ctcaccacag cttacacttg tgtttccgga agtgaagatg atacttttat
11581 tatacacatg tttaggtttc atgtgtactt gctattttgg tgtcttctct tttttgaacc
11641 ttaagcttag agcacctatg ggtgtctatg actttaaggt ctcaacacaa gagttcagat
11701 tcatgactgc taacaatcta actgcaccta gaaattcttg ggaggctatg gctctgaact
11761 ttaagttaat aggtattggc ggtacacctt gtataaaggt tgctgctatg cagtctaaac
11821 ttacagatct taaatgcaca tctgtggttc tcctctctgt gctccaacag ttacacttag
11881 aggctaatag tagggcctgg gctttctgtg ttaaatgcca taatgatata ttggcagcaa
11941 cagaccccag tgaggctttc gagaaattcg taagtctctt tgccacttta atgacttttt
12001 ctggtaatgt agatcttgat gcgttagcta gtgatatttt tgacactcct agcgtacttc
12061 aagctactct ttctgagttt tcacacttag ctacctttgc tgagttggaa gctgcgcaga
12121 aagcctatca ggaagctatg gactctggtg acacctcacc acaagttctt aaggctttgc
12181 agaaggctgt taatatagct aaaaacgcct atgagaagga taaggcagtg gcccgtaagt
12241 tagaacgtat ggctgaccag gctatgactt ctatgtataa gcaagcacgt gctgaagaca
12301 agaaagcaaa aattgtcagt gctatgcaaa ctatgttgtt tggtatgatt aagaagctcg
12361 acaacgatgt tcttaatggt atcatttcta acgctaggaa tggttgtata cctcttagtg
12421 tcattccact gtgtgcttca aataaacttc gcgttgtaat tcctgacttc accgtctgga
12481 atcaggtagt cacatatccc tcgcttaact acgctggggc tttgtgggac attacagtta
12541 taaacaatgt ggacaatgaa attgttaagt cttcagatgt tgtagacagc aatgaaaatt
12601 taacatggcc acttgtttta gaatgcacta gggcatccac ttctgccgtt aagttgcaaa
12661 ataatgagat caaaccttca ggtttaaaaa ccatggttgt gtctgcaggt caagagcaaa
12721 ctaactgtaa tactagttcc ttagcttatt acgaacctgt gcagggtcgt aaaatgctga
12781 tggctcttct ttctgataat gcctatctca aatgggcgcg tgttgaaggt aaggacggat
12841 ttgttagtgt agagctacaa cctccttgca aattcttgat tgcgggacca aaaggacctg
12901 aaatccgata tctctatttt gttaaaaatc ttaacaacct tcatcgcggg caagtgttag
12961 ggcacattgc tgcgactgtt agattgcaag ctggttctaa caccgagttt gcctctaatt
13021 cttcggtgtt gtcacttgtt aacttcaccg ttgatcctca aaaagcttat ctcgattttg
13081 tcaatgcggg aggtgcccca ttgacaaatt gtgttaagat gcttactcct aaaactggta
13141 caggtatagc tatatctgtt aaaccagaga gtacagctga tcaagagact tatggtggag
13201 cttcagtgtg tctctattgc cgtgcgcata tagaacatcc tgatgtctct ggtgtttgta
13261 aatataaggg taagtttgtc caaatccctg ctcagtgtgt ccgtgaccct gtgggatttt
13321 gtttgtcaaa taccccctgt aatgtctgtc aatattggat tggatatggg tgcaattgtg
13381 actcgcttag gcaagcagca ctgccccaat ctaaagattc caatttttta aacgagtccg
13441 gggttctatt gtaaatgccc gaatagaacc ctgttcaagt ggtttgtcca ctgatgtcgt
13501 ctttagggca tttgacatct gcaactataa ggctaaggtt gctggtattg gaaaatacta
13561 caagactaat acttgtaggt ttgtagaatt agatgaccaa gggcatcatt tagactccta
13621 ttttgtcgtt aagaggcata ctatggagaa ttatgaacta gagaagcact gttacgattt
13681 gttacgtgac tgtgatgctg tagctcccca tgatttcttc atctttgatg tagacaaagt
13741 taaaacacct catattgtac gtcagcgttt aactgagtac actatgatgg atcttgtata
13801 tgccctgagg cactttgatc aaaatagcga agtgcttaag gctatcttag tgaagtatgg
13861 ttgctgtgat gttacctact ttgaaaataa actctggttt gattttgttg aaaatcccag
13921 tgttattggt gtttatcata aacttggaga acgtgtacgc caagctatct taaacactgt
13981 taaattttgt gaccacatgg tcaaggctgg tttagtcggt gtgctcacac tagacaacca
14041 ggaccttaat ggcaagtggt atgattttgg tgacttcgta atcactcaac ctggttcagg
14101 agtagctata gttgatagct actattctta tttgatgcct gtgctctcaa tgaccgattg
14161 tctggccgct gagacacata gggattgtga ttttaataaa ccactcattg agtggccact
14221 tactgagtat gattttactg attataaggt acaactcttt gagaagtact ttaaatattg
14281 ggatcagacg tatcacgcaa attgcgttaa ttgtactgat gaccgttgtg tgttacattg
14341 tgctaatttc aatgtattgt ttgctatgac catgcctaag acttgtttcg gacccatagt
14401 ccgaaagatc tttgttgatg gcgtgccatt tgtagtatct tgtggttatc actacaaaga
14461 attaggttta gtcatgaata tggatgttag tctccataga cataggctct ctcttaagga
14521 gttgatgatg tatgccgctg atccagccat gcacattgcc tcctctaacg cttttcttga
14581 tttgaggaca tcatgtttta gtgtcgctgc acttacaact ggtttgactt ttcaaactgt
14641 gcggcctggc aattttaacc aagacttcta tgatttcgtg gtatctaaag gtttctttaa
14701 ggagggctct tcagtgacgc tcaaacattt tttctttgct caagatggta atgctgctat
14761 tacagattat aattactatt cttataatct gcctactatg tgtgacatca aacaaatgtt
14821 gttctgcatg gaagttgtaa acaagtactt cgaaatctat gacggtggtt gtcttaatgc
14881 ttctgaagtg gttgttaata atttagacaa gagtgctggc catcctttta ataagtttgg
14941 caaagctcgt gtctattatg agagcatgtc ttaccaggag caagatgaac tctttgccat
15001 gacaaagcgt aacgtcattc ctaccatgac tcaaatgaat ctaaaatatg ctattagtgc
15061 taagaataga gctcgcactg ttgcaggcgt gtccatactt agcacaatga ctaatcgcca
15121 gtaccatcag aaaatgctta agtccatggc tgcaactcgt ggagcgactt gcgtcattgg
15181 tactacaaag ttctatggtg gctgggattt catgcttaaa acattgtaca aagatgttga
15241 taatccgcat cttatgggtt gggattaccc taagtgtgat agagctatgc ctaatatgtg
15301 tagaatcttc gcttcactca tattagctcg taaacatggc acttgttgta ctacaaggga
15361 cagattttat cgcttggcaa atgagtgtgc tcaggtgcta agcgaatatg ttctatgtgg
15421 tggtggttac tacgtcaaac ctggaggtac cagtagcgga gatgccacca ctgcatatgc
15481 caatagtgtc tttaacattt tgcaggcgac aactgctaat gtcagtgcac ttatgggtgc
15541 taatggcaac aagattgttg acaaagaagt taaagacatg cagtttgatt tgtatgtcaa
15601 tgtttacagg agcactagcc cagaccccaa atttgttgat aaatactatg cttttcttaa
15661 taagcacttt tctatgatga tactgtctga tgacggtgtc gtttgctata atagtgatta
15721 tgcagctaag ggttacattg ctggaataca gaattttaag gaaacgctgt attatcagaa
15781 caatgtcttt atgtctgaag ctaaatgctg ggtggaaacc gatctgaaga aagggccaca
15841 tgaattctgt tcacagcata cgctttatat taaggatggc gacgatggtt acttccttcc
15901 ttatccagac ccttcaagaa ttttgtctgc cggttgcttt gtagatgata tcgttaagac
15961 tgacggtaca ctcatggtag agcggtttgt gtctttggct atagatgctt accctctcac
16021 aaagcatgaa gatatagaat accagaatgt attctgggtc tacttacagt atatagaaaa
16081 actgtataaa gaccttacag gacacatgct tgacagttat tctgtcatgc tatgtggtga
16141 taattctgct aagttttggg aagaggcatt ctacagagat ctctatagtt cgcctaccac
16201 tttgcaggct gtcggttcat gcgttgtatg ccattcacag acttccctac gctgtgggac
16261 atgcatccgt agaccatttc tctgctgtaa atgctgctat gatcatgtta tagcaactcc
16321 acataagatg gttttgtctg tttctcctta cgtttgtaat gcccctggtt gtggcgtttc
16381 agacgttact aagctatatt taggtggtat gagctacttt tgtgtagatc atagacctgt
16441 gtgtagtttt ccactttgcg ctaatggtct tgtattcggc ttatacaaga atatgtgcac
16501 aggtagtcct tctatagttg aatttaatag gttggctacc tgtgactgga ctgaaagtgg
16561 tgattacacc cttgccaata ctacaacaga accactcaaa ctttttgctg ctgagacttt
16621 acgtgccact gaagaggcgt ctaagcagtc ttatgctatt gccaccatca aagaaattgt
16681 tggtgagcgc caactattac ttgtgtggga ggctggcaag tccaaaccac cactcaatcg
16741 taattatgtt tttactggtt atcatataac caaaaatagt aaagtgcagc tcggtgagta
16801 catcttcgag cgcattgatt atagtgatgc tgtatcctac aagtctagta caacgtataa
16861 actgactgta ggtgacatct tcgtacttac ctctcactct gtggctacct tgacggcgcc
16921 cacaattgtg aatcaagaga ggtatgttaa aattactggg ttgtacccaa ccattacggt
16981 acctgaagag ttcgcaagtc atgttgccaa cttccaaaaa tcaggttata gtaaatatgt
17041 cactgttcag ggaccacctg gcactggcaa aagtcatttt gctatagggt tagcgattta
17101 ctaccctaca gcacgtgttg tttatacagc atgttcacac gcagctgttg atgctttgtg
17161 tgaaaaagct tttaaatatt tgaacattgc taaatgttcc cgtatcattc ctgcaaaggc
17221 acgtgttgag tgctatgaca ggtttaaagt taatgagaca aattctcaat atttgtttag
17281 tactattaat gctctaccag aaacttctgc cgatattctg gtggttgatg aggttagtat
17341 gtgcactaat tatgatcttt caattattaa tgcacgtatt aaagctaagc acattgtcta
17401 tgtaggagat ccagcacagt tgccagctcc taggactttg ttgactagag gcacattgga
17461 accagaaaat ttcaatagtg tcactagatt gatgtgtaac ttaggtcctg acatattttt
17521 aagtatgtgc tacaggtgtc ctaaggaaat agtaagcact gtgagcgctc ttgtctacaa
17581 taataaattg ttagccaaga aggagctttc aggccagtgc tttaaaatac tctataaggg
17641 caatgtgacg catgatgcta gctctgccat taatagacca caactcacat ttgtgaagaa
17701 ttttattact gccaatccgg catggagtaa ggcagtcttt atttcgcctt ataattcaca
17761 gaatgctgtg gctcgttcaa tgctgggtct tactactcag actgttgatt cctcacaggg
17821 ttcagaatac cagtatgtta tcttctgtca aacagcagat acggcacatg ctaacaacat
17881 taacagattt aatgttgcaa tcactcgtgc ccaaaaaggt attctttgtg ttatgacatc
17941 tcaggcactc tttgagtcct tagagtttac tgaattgtct tttactaatt acaagctcca
18001 gtctcagatt gtaactggcc tttttaaaga ttgctctaga gaaacttctg gcctctcacc
18061 tgcttatgca ccaacatacg ttagtgttga tgacaagtat aagacgagtg atgagctttg
18121 cgtgaatctt aatttacccg caaatgtccc atactctcgt gttatttcca ggatgggctt
18181 taaactcgat gcaacagttc ctggatatcc taagcttttc attactcgtg aagaggctgt
18241 aaggcaagtt cgaagctgga taggcttcga tgttgagggt gctcatgctt cccgtaatgc
18301 atgtggcacc aatgtgcctc tacaattagg attttcaact ggtgtgaact ttgttgttca
18361 gccagttggt gttgtagaca ctgagtgggg taacatgtta acgggcattg ctgcccgtcc
18421 tccaccaggt gaacagttta agcacctcgt gcctcttatg cataaggggg ctgcgtggcc
18481 tattgttaga cgacgtatag tgcaaatgtt gtcagacact ttagacaaat tgtctgatta
18541 ctgtacgttt gtttgttggg ctcatggctt tgaattaacg tctgcatcat acttttgcaa
18601 gataggtaag gaacagaagt gttgcatgtg caatagacgc gctgcagcgt actcttcacc
18661 tctgcaatct tatgcctgct ggactcattc ctgcggttat gattatgtct acaacccttt
18721 ctttgtcgat gttcaacagt ggggttatgt aggcaatctt gctactaatc acgatcgtta
18781 ttgctctgtc catcaaggag ctcatgtggc ttctaatgat gcaataatga ctcgttgttt
18841 agctattcat tcttgtttta tagaacgtgt ggattgggat atagagtatc cttatatctc
18901 acatgaaaag aaattgaatt cctgttgtag aatcgttgag cgcaacgtcg tacgtgctgc
18961 tcttcttgcc ggttcatttg acaaagtcta tgatattggc aatcctaaag gaattcctat
19021 tgttgatgac cctgtggttg attggcatta ttttgatgca cagcccttga ccagaaaggt
19081 acaacagctt ttctatacag aggacatggc ctcaagattt gctgatgggc tctgcttatt
19141 ttggaactgt aatgtaccaa aatatcctaa taatgcaatt gtatgcaggt ttgacacacg
19201 tgtgcattct gagttcaatt tgccaggttg tgatggcggt agtttgtatg ttaacaagca
19261 cgcttttcat acaccagcat atgatgtgag tgcattccgt gatctgaaac ctttaccatt
19321 cttttattat tctactacac catgtgaagt gcatggtaat ggtagtatga tagaggatat
19381 tgattatgta cccctaaaat ctgcagtctg tattacagct tgtaatttag ggggcgctgt
19441 ttgtaggaag catgctacag agtacagaga gtatatggaa gcatataatc ttgtctctgc
19501 atcaggtttc cgcctttggt gttataagac ctttgatatt tataatctct ggtctacttt
19561 tacaaaagtt caaggtttgg aaaacattgc ttttaatgtt gttaaacaag gccattttat
19621 tggtgttgag ggtgaactac ctgtagctgt agtcaatgat aagatcttca ccaagagtgg
19681 cgttaatgac atttgtatgt ttgagaataa aaccactttg cctactaata tagcttttga
19741 actctatgct aagcgtgctg tacgctcgca tcccgatttc aaattgctac acaatttaca
19801 agcagacatt tgctacaagt tcgtcctttg ggattatgaa cgtagcaata tttatggtac
19861 tgctactatt ggtgtatgta agtacactga tattgatgtt aattcagctt tgaatatatg
19921 ttttgacata cgcgataatg gttcattgga gaagttcatg tctactccca atgccatctt
19981 tatttctgat agaaaaatta agaaataccc ttgtatggta ggtcctgatt atgcttactt
20041 caatggtgct atcatccgtg atagtgatgt tgttaaacaa ccagtgaagt tctacttgta
20101 taagaaagtc aataatgagt ttattgatcc tactgagtgt atttacactc agagtcgctc
20161 ttgtagtgac ttcctacccc tgtctgacat ggagaaagac tttctatctt ttgatagtga
20221 tgttttcatt aagaagtatg gcttggaaaa ctatgctttt gagcacgtag tctatggaga
20281 cttctctcat actacgttag gcggtcttca cttgcttatt ggtttataca agaagcaaca
20341 ggaaggtcat attattatgg aagaaatgct aaaaggtagc tcaactattc ataactattt
20401 tattactgag actaacacag cggcttttaa ggcggtgtgt tctgttatag atttaaagct
20461 tgacgacttt gttatgattt taaagagtca agaccttggc gtagtatcca aggttgtcaa
20521 ggttcctatt gacttaacaa tgattgagtt tatgttatgg tgtaaggatg gacaggttca
20581 aaccttctac cctcgactcc aggcttctgc agattggaaa cctggtcatg caatgccatc
20641 cctctttaaa gttcaaaatg taaaccttga acgttgtgag cttgctaatt acaagcaatc
20701 tattcctatg cctcgcggtg tgcacatgaa catcgctaaa tatatgcaat tgtgccagta
20761 tttaaatact tgcacattag ccgtgcctgc caatatgcgt gttatacatt ttggcgctgg
20821 ttctgataaa ggtatcgctc ctggtacatc agttttacga cagtggcttc ctacagatgc
20881 cattattata gataatgatt taaatgagtt cgtgtcagat gctgacataa ctttatttgg
20941 agattgtgta actgtacgtg tcggccaaca agtggatctt gttatttccg acatgtatga
21001 tcctactact aagaatgtaa caggtagtaa tgagtcaaag gctttattct ttacttacct
21061 gtgtaacctc attaataata atcttgctct tggtgggtct gttgctatta aaataacaga
21121 acactcttgg agcgttgaac tttatgaact tatgggaaaa tttgcttggt ggactgtttt
21181 ctgcaccaat gcaaatgcat cctcatctga aggattcctc ttaggtatta attacttggg
21241 tactattaaa gaaaatatag atggtggtgc tatgcacgcc aactatatat tttggagaaa
21301 ttccactcct atgaatctga gtacttactc actttttgat ttatccaagt ttcaattaaa
21361 attaaaagga acaccagttc ttcaattaaa ggagagtcaa attaacgaac tcgtaatatc
21421 tctcctgtcg cagggtaagt tacttatccg tgacaatgat acactcagtg tttctactga
21481 tgttcttgtt aacacctaca gaaagttacg ttgatgtagg gccagattct gttaagtctg
21541 cttgtattga ggttgatata caacagactt tctttgataa aacttggcct aggccaattg
21601 atgtttctaa ggctgacggt attatatacc ctcaaggccg tacatattct aacataacta
21661 tcacttatca aggtcttttt ccctatcagg gagaccatgg tgatatgtat gtttactctg
21721 caggacatgc tacaggcaca actccacaaa agttgtttgt agctaactat tctcaggacg
21781 tcaaacagtt tgctaatggg tttgtcgtcc gtataggagc agctgccaat tccactggca
21841 ctgttattat tagcccatct accagcgcta ctatacgaaa aatttaccct gcttttatgc
21901 tgggttcttc agttggtaat ttctcagatg gtaaaatggg ccgcttcttc aatcatactc
21961 tagttctttt gcccgatgga tgtggcactt tacttagagc tttttattgt attctagagc
22021 ctcgctctgg aaatcattgt cctgctggca attcctatac ttcttttgcc acttatcaca
22081 ctcctgcaac agattgttct gatggcaatt acaatcgtaa tgccagtctg aactctttta
22141 aggagtattt taatttacgt aactgcacct ttatgtacac ttataacatt accgaagatg
22201 agattttaga gtggtttggc attacacaaa ctgctcaagg tgttcacctc ttctcatctc
22261 ggtatgttga tttgtacggc ggcaatatgt ttcaatttgc caccttgcct gtttatgata
22321 ctattaagta ttattctatc attcctcaca gtattcgttc tatccaaagt gatagaaaag
22381 cttgggctgc cttctacgta tataaacttc aaccgttaac tttcctgttg gatttttctg
22441 ttgatggtta tatacgcaga gctatagact gtggttttaa tgatttgtca caactccact
22501 gctcatatga atccttcgat gttgaatctg gagtttattc agtttcgtct ttcgaagcaa
22561 aaccttctgg ctcagttgtg gaacaggctg aaggtgttga atgtgatttt tcacctcttc
22621 tgtctggcac acctcctcag gtttataatt tcaagcgttt ggtttttacc aattgcaatt
22681 ataatcttac caaattgctt tcactttttt ctgtgaatga ttttacttgt agtcaaatat
22741 ctccagcagc aattgctagc aactgttatt cttcactgat tttggattat ttttcatacc
22801 cacttagtat gaaatccgat ctcagtgtta gttctgctgg tccaatatcc cagtttaatt
22861 ataaacagtc cttttctaat cccacatgtt tgattttagc gactgttcct cataacctta
22921 ctactattac taagcctctt aagtacagct atattaacaa gtgctctcgt cttctttctg
22981 atgatcgtac tgaagtacct cagttagtga acgctaatca atactcaccc tgtgtatcca
23041 ttgtcccatc cactgtgtgg gaagacggtg attattatag gaaacaacta tctccacttg
23101 aaggtggtgg ctggcttgtt gctagtggct caactgttgc catgactgag caattacaga
23161 tgggctttgg tattacagtt caatatggta cagacaccaa tagtgtttgc cccaagcttg
23221 aatttgctaa tgacacaaaa attgcctctc aattaggcaa ttgcgtggaa tattccctct
23281 atggtgtttc gggccgtggt gtttttcaga attgcacagc tgtaggtgtt cgacagcagc
23341 gctttgttta tgatgcgtac cagaatttag ttggctatta ttctgatgat ggcaactact
23401 actgtttgcg tgcttgtgtt agtgttcctg tttctgtcat ctatgataaa gaaactaaaa
23461 cccacgctac tctatttggt agtgttgcat gtgaacacat tttctctacc atgtctcaat
23521 actcccgttc tacgcgatca atgcttaaac ggcgagattc tacatatggt ccccttcaga
23581 cacctgttgg ttgtgtccta ggacttgtta attcctcttt gttcgtagag gactgcaagt
23641 tgcctcttgg tcaatctctc tgtgctcttc ctgacacacc tagtactctc acacctcgca
23701 gtgtgcgctc tgttccaggt gaaatgcgct tggcatccat tgcttttaat catcctattc
23761 aggttgatca acttaatagt agttatttta aattaagtat acctactaat ttttcctttg
23821 gtgtgactca ggagtacatt cagacaacca ttcagaaagt tactgttgat tgtaaacagt
23881 acgtttgcaa tggtttccag aagtgtgagc aattactgcg cgagtatggc cagttttgtt
23941 ccaaaataaa ccgggctctc catggtgcca atttacgcca ggatgattct gtacgtaatt
24001 tgtttgcgag cgtgaaaagc tctcaatcat ctcctatcat accaggtttt ggaggtgact
24061 ttaatttgac acttctagaa cctgtttcta tatctactgg cagtcgtagt gcacgtagtg
24121 ctattgagga tttgctattt gacaaagtca ctatagctga tcctggttat atgcaaggtt
24181 acgatgattg catgcagcaa ggtccagcat cagctcgtga tcttatttgt gctcaatatg
24241 tggctggtta caaagtatta cctcctctta tggatgttaa tatggaagcc gcgtatacct
24301 catctttgct tggcagcata gcaggtgttg gctggactgc tggcttatcc tcctttgctg
24361 ctattccatt tgcacagagt atcttttata ggttaaacgg tgttggcatt actcaacagg
24421 ttctttcaga gaaccaaaag cttattgcca ataagtttaa tcaggctctg ggagctatgc
24481 aaacaggctt cactacaact aatgaagctt ttcggaaggt tcaggatgct gtgaacaaca
24541 atgcacaggc tctatccaaa ttagctagcg agctatctaa tacttttggt gctatttccg
24601 cctctattgg agacatcata caacgtcttg atgttctcga acaggacgcc caaatagaca
24661 gacttattaa tggccgtttg acaacactaa atgcttttgt tgcacagcag cttgttcgtt
24721 ccgaatcagc tgctctttcc gctcaattgg ctaaagataa agtcaatgag tgtgtcaagg
24781 cacaatccaa gcgttctgga ttttgcggtc aaggcacaca tatagtgtcc tttgttgtaa
24841 atgcccctaa tggcctttac ttcatgcatg ttggttatta ccctagcaac cacattgagg
24901 ttgtttctgc ttatggtctt tgcgatgcag ctaaccctac taattgtata gcccctgtta
24961 atggctactt tattaaaact aataacacta ggattgttga tgagtggtca tatactggct
25021 cgtccttcta tgcacctgag cccatcacct ctcttaatac taagtatgtt gcaccacagg
25081 tgacatacca aaacatttct actaacctcc ctcctcctct tctcggcaat tccaccggga
25141 ttgacttcca agatgagttg gatgagtttt tcaaaaatgt tagcaccagt atacctaatt
25201 ttggttctct aacacagatt aatactacat tactcgatct tacctacgag atgttgtctc
25261 ttcaacaagt tgttaaagcc cttaatgagt cttacataga ccttaaagag cttggcaatt
25321 atacttatta caacaaatgg ccgtggtaca tttggcttgg tttcattgct gggcttgttg
25381 ccttagctct atgcgtcttc ttcatactgt gctgcactgg ttgtggcaca aactgtatgg
25441 gaaaacttaa gtgtaatcgt tgttgtgata gatacgagga atacgacctc gagccgcata
25501 aggttcatgt tcactaatta acgaactatc aatgagagtt caaagaccac ccactctctt
25561 gttagtgttc tcactctctc ttttggtcac tgcattttca aaacctctct atgtacctga
25621 gcattgtcag aattattctg gttgcatgct tagggcttgt attaaaactg cccaagctga
25681 tacagctggt ctttatacaa attttcgaat tgacgtccca tctgcagaat caactggtac
25741 tcaatcagtt tctgtcgatc gtgagtcaac ttcaactcat gatggtccta ccgaacatgt
25801 tactagtgtg aatctttttg acgttggtta ctcagttaat taacgaactc tatggattac
25861 gtgtctctgc ttaatcaaat ttggcagaag taccttaatt caccgtatac tacttgtttg
25921 tatatcccta aacccacagc taagtataca cctttagttg gcacttcatt gcaccctgtg
25981 ctgtggaact gtcagctatc ctttgctggt tatactgaat ctgctgttaa ttctacaaaa
26041 gctttggcca aacaggacgc agctcagcga atcgcttggt tgctacataa ggatggagga
26101 atccctgatg gatgttccct ctacctccgg cactcaagtt tattcgcgca aagcgaggaa
26161 gaggagtcat tctccaacta agaaactgcg ctacgttaag cgtagatttt ctcttctgcg
26221 ccctgaagac cttagtgtta ttgtccaacc aacacactat gtcagggtta cattttcaga
26281 ccccaacatg tggtatctac gttcgggtca tcatttacac tcagttcaca attggcttaa
26341 accttatggc ggccaacctg tttctgagta ccatattact ctagctttgc taaatctcac
26401 tgatgaagat ttagctagag atttttcacc cattgcgctc tttttgcgca atgtcagatt
26461 tgagctacat gagttcgcct tgctgcgcaa aactcttgtt cttaatgcat cagagatcta
26521 ctgtgctaac atacatagat ttaagcctgt gtatagagtt aacacggcaa tccctactat
26581 taaggattgg cttctcgttc agggattttc cctttaccat agtggcctcc ctttacatat
26641 gtcaatctct aaattgcatg cactggatga tgttactcgc aattacatca ttacaatgcc
26701 atgctttaga acttatcctc aacaaatgtt tgttactcct ttggccgtag atgttgtctc
26761 catacggtct tccaatcagg gtaataaaca aattgttcat tcttacccca ttttacatca
26821 tccaggattt taacgaacta tggctttctc ggcgtcttta tttaaacccg tccagctagt
26881 cccagtttct cctgcatttc atcgcattga gtctactgac tctattgttt tcacatacat
26941 tcctgctagc ggctatgtag ctgctttagc tgtcaatgtg tgtctcattc ccctattatt
27001 actgctacgt caagatactt gtcgtcgcag cattatcaga actatggttc tctatttcct
27061 tgttctgtat aactttttat tagccattgt actagtcaat ggtgtacatt atccaactgg
27121 aagttgcctg atagccttct tagttatcct cataatactt tggtttgtag atagaattcg
27181 tttctgtctc atgctgaatt cctacattcc actgtttgac atgcgttctc actttattcg
27241 tgttagtaca gtttcttctc atggtatggt ccctgtcata cacaccaaac cattatttat
27301 tagaaacttc gatcagcgtt gcagctgttc tcgttgtttt tatttgcact cttccactta
27361 tatagagtgc acttatatta gccgttttag taagattagc ctagtttctg taactgactt
27421 ctccttaaac ggcaatgttt ccactgtttt cgtgcctgca acgcgcgatt cagttcctct
27481 tcacataatc gccccgagct cgcttatcgt ttaagcagct ctgcgctact atgggtcccg
27541 tgtagaggct aatccattag tctctctttg gacatatgga aaacgaacta tgttaccctt
27601 tgtccaagaa cgaatagggt tgttcatagt aaactttttc atttttaccg tagtatgtgc
27661 tataacactc ttggtgtgta tggctttcct tacggctact agattatgtg tgcaatgtat
27721 gacaggcttc aataccctgt tagttcagcc cgcattatac ttgtataata ctggacgttc
27781 agtctatgta aaattccagg atagtaaacc ccctctacca cctgacgagt gggtttaacg
27841 aactccttca taatgtctaa tatgacgcaa ctcactgagg cgcagattat tgccattatt
27901 aaagactgga actttgcatg gtccctgatc tttctcttaa ttactatcgt actacagtat
27961 ggatacccat cccgtagtat gactgtctat gtctttaaaa tgtttgtttt atggctccta
28021 tggccatctt ccatggcgct atcaatattt agcgccattt atccaattga tctagcttcc
28081 cagataatct ctggcattgt agcagctgtt tcagctatga tgtggatttc ctactttgtg
28141 cagagtatcc ggctgtttat gagaactgga tcatggtggt cattcaatcc tgagactaat
28201 tgccttttga acgttccatt tggtggtaca actgtcgtac gtccactcgt agaggactct
28261 accagtgtaa ctgctgttgt aaccaatggc cacctcaaaa tggctggcat gcatttcggt
28321 gcttgtgact acgacagact tcctaatgaa gtcaccgtgg ccaaacccaa tgtgctgatt
28381 gctttaaaaa tggtgaagcg gcaaagctac ggaactaatt ccggcgttgc catttaccat
28441 agatataagg caggtaatta caggagtccg cctattacgg cggatattga acttgcattg
28501 cttcgagctt aggctcttta gtaagagtat cttaattgat tttaacgaat ctcaatttca
28561 ttgttatggc atcccctgct gcacctcgtg ctgtttcctt tgccgataac aatgatataa
28621 caaatacaaa cctgtctcga ggtagaggac gtaatccaaa accacgagct gcaccaaata
28681 acactgtctc ttggtacact gggcttaccc aacacgggaa agtccctctt acctttccac
28741 ctgggcaggg tgtacctctt aatgccaatt ccaccccagc gcaaaatgct gggtattggc
28801 ggagacagga cagaaaaatt aataccggga atggaattaa gcaactggct cccaggtggt
28861 acttctacta cactggaacc ggacccgaag cagcactccc attccgggct gttaaggatg
28921 gcatcgtttg ggtccatgaa catggcgcca ctgatgctcc ttcaactttt gggacgcgga
28981 accctaacaa tgattcagct attgttacac aattcgcgcc cggtactaag cttcctaaaa
29041 acttccacat tgaggggact ggaggcaata gtcaatcatc ttcaagagcc tctagcgtaa
29101 gcagaaactc ttccagatct agttcacaag gttcaagatc aggaaactct acccgcggca
29161 cttctccagg tccatctgga atcggagcag taggaggtga tctactttac cttgatcttc
29221 tgaacagact acaagccctt gagtctggca aagtaaagca atcgcagcca aaagtaatca
29281 ctaagaaaga tgctgctgct gctaaaaata agatgcgcca caagcgcact tccaccaaaa
29341 gtttcaacat ggtgcaagct tttggtcttc gcggaccagg agacctccag ggaaactttg
29401 gtgatcttca attgaataaa ctcggcactg aggacccacg ttggccccaa attgctgagc
29461 ttgctcctac agccagtgct tttatgggta tgtcgcaatt taaacttacc catcagaaca
29521 atgatgatca tggcaaccct gtgtacttcc ttcggtacag tggagccatt aaacttgacc
29581 caaagaatcc caactacaat aagtggttgg agcttcttga gcaaaatatt gatgcctaca
29641 aaaccttccc taagaaggaa aagaaacaaa aggcaccaaa agaagaatca acagaccaaa
29701 tgtctgaacc tcctaaggag cagcgtgtgc aaggtagcat cactcagcgc actcgcaccc
29761 gtccaagtgt tcagcctggt ccaatgattg atgttaacac tgattagtgt cactcaaagt
29821 aacaagatcg cggcaatcgt ttgtgtttgg taaccccatc tcaccatcgc ttgtccactc
29881 ttgcacagaa tggaatcatg ttgtaattac agtgcaataa ggtaattata acccatttaa
29941 ttgatagcta tgctttatta aagtgtgtag ctgtagagag aatgttaaag actgtcacct
30001 ctgcctgatt gcaagtgaac agtgcccccc gggaagagct ctacagtgtg aaatgtaaat
30061 aaaaatagct attattcaat tagattaggc taattagatg atttgcaaaa aaaaaaaaaa
30121 aaa
"""
for s in " \n0123456789":
corona = corona.replace(s, "")
corona
corona
!unzip /content/SARS2-all.zip
```
| github_jupyter |
# 2A.algo - L'énigme d'Einstein et sa résolution
Résolution de l'énigme [L'énigme d'Einstein](http://fr.wikipedia.org/wiki/%C3%89nigme_d'Einstein). Implémentatin d'une solution à base de règles.
```
from io import StringIO
from pandas import read_csv
```
[L'énigme d'Einstein](http://fr.wikipedia.org/wiki/%C3%89nigme_d'Einstein) est une énigme comme celle que résoud [Hermionne](http://www.encyclopedie-hp.org/hogwarts/chamber_of_stone.php) dans le premier tome de Harry Potter. Je la reproduis ici :
Il y a cinq maisons de cinq couleurs différentes. Dans chacune de ces maisons, vit une personne de nationalité différente. Chacune de ces personnes boit une boisson différente, fume un cigare différent et a un animal domestique différent.
1. L'Anglais vit dans la maison rouge.
2. Le Suédois a des chiens.
3. Le Danois boit du thé.
4. La maison verte est à gauche de la maison blanche.
5. Le propriétaire de la maison verte boit du café.
6. La personne qui fume des Pall Mall a des oiseaux.
7. Le propriétaire de la maison jaune fume des Dunhill.
8. La personne qui vit dans la maison du centre boit du lait.
9. Le Norvégien habite dans la première maison.
10. L'homme qui fume des Blend vit à côté de celui qui a des chats.
11. L'homme qui a un cheval est le voisin de celui qui fume des Dunhill.
12. Le propriétaire qui fume des Blue Master boit de la bière.
13. L'Allemand fume des prince.
14. Le Norvégien vit juste à côté de la maison bleue.
15. L'homme qui fume des Blend a un voisin qui boit de l'eau.
Question : Qui a le poisson ?
Après quelques essais, une bonne feuille de papier, on arrive à reconstituer la solution après de nombreuses déductions logiques et quelques essais. On peut voir aussi ce jeu comme un puzzle : chaque configuration est un pièce du puzzle dont la forme des bords est définie par toutes ces règles. Il faut trouver le seul emboîtement possible sachant que parfois, une pièce peut s'emboîter avec plusieurs mais qu'il n'existe qu'une façon de les emboîter toutes ensemble. Ecrire un programme qui résoud ce problème revient à s'intéresser à deux problèmes :
1. Comment définir une pièce du puzzle ?
2. Comment parcourir toutes les combinaisons possibles ?
Chaque règle ou pièce de puzzle peut être exprimer comme une [clause](http://fr.wikipedia.org/wiki/Clause_de_Horn). Pour notre problème, chaque pièce du puzzle est simplement décrite par un attribut (rouge, norvégien) et un numéro de maisons (1 à 5). Les règles définissent la compatibilité de deux pièces. On peut regrouper ces règles en cinq catégories :
1. Un attribut est à la position p (règle 9).
2. Deux attributs sont équivalents (règle 1).
3. Deux attributs sont voisins (règle 11).
4. Deux attributs sont ordonnés par rapport aux positions (règle 4).
5. Deux attributs font partie du même ensemble et sont exclusives : on ne peut pas être l'un et l'autre à la fois (rouge et jaune par exemple).
Une fois que chaque règle a été exprimée dans une de ces cinq catégories, il faut définir l'association de deux règles (ou clause) pour former une clause plus complexe. Trois cas possibles :
1. Deux clauses sont compatibles : on peut avoir l'une et l'autre.
2. Deux clauses sont incompatibles : on ne peut avoir l'une et l'autre.
Dans le premier cas, la clause résultante est simplement qu'on peut la clause A et la clause B : $A \, et\, B$. Dans le second cas, il existe deux possibilités, on peut avoir l'une et l'opposé de l'autre ou l'inverse : $(A \, et\, non \, B) \, ou\, (non \, A \, et\, B)$.
Avec cette description, il est plus facile d'exprimer le problème avec des objets informatiques ce que fait le programme suivant. Il explicite ensuite toutes les configurations compatibles avec une règle donnée (mais pas toutes ensembles).
On commence par la fonction [permutation](http://www.xavierdupre.fr/app/ensae_teaching_cs/helpsphinx/ensae_teaching_cs/special/einstein_prolog.html?highlight=permutation#ensae_teaching_cs.special.einstein_prolog.permutation) qui énumère les permutations d'un ensemble :
```
def permutation(nb):
per = []
p = [i for i in range(0, nb)]
while p[0] < nb:
cont = False
for i in range(1, nb):
if p[i] in p[0:i]:
cont = True
break
if not cont:
per.append(copy.copy(p))
p[nb-1] += 1
for j in range(nb-1, 0, -1):
if p[j] >= nb:
p[j] = 0
p[j-1] += 1
return per
import copy
ttcouleur = ["jaune", "bleu", "rouge", "blanc", "vert"]
ttnationalite = ["danois", "norvegien", "anglais", "allemand", "suedois"]
ttboisson = ["eau", "the", "lait", "cafe", "biere"]
ttcigare = ["Dunhill", "Blend", "Pall Mall", "Prince", "Bluemaster"]
ttanimal = ["chats", "cheval", "oiseaux", "poisson", "chiens"]
ensemble = [ttcouleur, ttnationalite, ttboisson, ttcigare, ttanimal]
class Rule:
"""
This class defines a constraint of the problem
or a clause (see `http://en.wikipedia.org/wiki/Clause_(logic)`)
There are 5 different types of clauses described by Einstein's enigma
each of them is described by a different class. There are defined by classes:
@ref cl RulePosition, @ref cl RuleEquivalence, @ref cl RuleVoisin,
@ref cl RuleAvant, @ref cl RuleEnsemble.
"""
def __init__(self):
"""
constructor
"""
#: name of the rule
self.name = None
#: set of clauses
self.set = None
def genere(self):
"""
Generates all possible clauses (list of lists)
(`l[0][0]` et `l[0][1]`) ou (`l[1][0]` et `l[1][1]`),
a clause is a triplet of
`(person, (property, category))`.
"""
return None
def __str__(self):
"""
display
"""
if self.name != None:
if "clauses" not in self.__dict__:
s = self.name + " \t: "
a = self.genere()
for al in a:
st = "\n ou " + str(al)
if len(st) > 260:
st = st[:260] + "..."
s += st
if len(s) > 1000:
break
return s
else:
s = self.name + " \t: " + str(self.set)
for al in self.clauses:
st = "\n ou " + str(al)
if len(st) > 260:
st = st[:260] + "..."
s += st
if len(s) > 1000:
break
return s
else:
return "None"
def combine(self, cl1, cl2):
"""
Combine two clauses, two cases:
1. nothing in common or everything in common --> concatenation of clauses
2. a position or a property in common --> null clause
:param cl1: clause 1
:param cl2: clause 2
:return: the new clause
A clause is a @ref cl Rule.
"""
# incompatibility
for p1 in cl1:
for p2 in cl2:
if p1[1][0] == p2[1][0]: # same property
if p1[0] != p2[0]: # but different positions
return None
if p1[0] == p2[0]: # same person
if p1[1][1] == p2[1][1] and p1[1][0] != p2[1][0]:
# same category but different properties
return None
# compatibility
r = copy.deepcopy(cl1)
for c in cl2:
if c not in r:
r.append(c)
return r
def combine_cross_sets(self, set1, set2):
"""
Combines two sets of clauses.
:param set1: set of clauses 1
:param set2: set of clauses 2
:return: combination
"""
if len(set1) == 0:
return copy.deepcopy(set2)
if len(set2) == 0:
return copy.deepcopy(set1)
res = []
for cl1 in set1:
for cl2 in set2:
r = self.combine(cl1, cl2)
if r != None:
res.append(r)
return res
class RulePosition (Rule):
"""
p1 at position
"""
def __init__(self, p1, pos):
self.set = [p1]
self.name = "position"
self.position = pos
def genere(self):
"""
overrides method ``genere``
"""
return [[(self.position, self.set[0])]]
class RuleEquivalence (Rule):
"""
p1 equivalent to p2
"""
def __init__(self, p1, p2):
self.set = [p1, p2]
self.name = "equivalence"
def genere(self):
"""
overrides method ``genere``
"""
l = []
for i in range(0, 5):
l.append([(i, self.set[0]), (i, self.set[1])])
return l
class RuleVoisin (Rule):
"""
p1 and p2 are neighbors
"""
def __init__(self, p1, p2):
self.set = [p1, p2]
self.name = "voisin"
def genere(self):
"""
overrides method ``genere``
"""
l = []
for i in range(0, 4):
l.append([(i, self.set[0]), (i+1, self.set[1])])
l.append([(i+1, self.set[0]), (i, self.set[1])])
return l
class RuleAvant (Rule):
"""
p1 before p2
"""
def __init__(self, p1, p2):
self.set = [p1, p2]
self.name = "avant"
def genere(self):
"""
overrides method ``genere``
"""
l = []
for j in range(1, 5):
for i in range(0, j):
l.append([(i, self.set[0]), (j, self.set[1])])
return l
class RuleEnsemble (Rule):
"""
permutation of the elements of a category
"""
def __init__(self, set, categorie):
self.set = [(s, categorie) for s in set]
self.name = "ensemble"
def genere(self):
"""
overrides method ``genere``
"""
l = []
per = permutation(5)
for p in per:
tl = []
for i in range(0, len(p)):
tl.append((i, self.set[p[i]]))
l.append(tl)
return l
def find(p):
for i in range(0, len(ensemble)):
if p in ensemble[i]:
return (p, i)
return None
regle = []
regle.append(RulePosition(find("lait"), 2))
regle.append(RulePosition(find("norvegien"), 0))
regle.append(RuleEquivalence(find("Pall Mall"), find("oiseaux")))
regle.append(RuleEquivalence(find("anglais"), find("rouge")))
regle.append(RuleEquivalence(find("suedois"), find("chiens")))
regle.append(RuleEquivalence(find("danois"), find("the")))
regle.append(RuleEquivalence(find("vert"), find("cafe")))
regle.append(RuleEquivalence(find("jaune"), find("Dunhill")))
regle.append(RuleEquivalence(find("biere"), find("Bluemaster")))
regle.append(RuleEquivalence(find("allemand"), find("Prince")))
regle.append(RuleVoisin(find("Dunhill"), find("cheval")))
regle.append(RuleVoisin(find("norvegien"), find("bleu")))
regle.append(RuleVoisin(find("Blend"), find("eau")))
regle.append(RuleVoisin(find("Blend"), find("chats")))
regle.append(RuleAvant(find("vert"), find("blanc")))
regle.append(RuleEnsemble(ttcouleur, 0))
regle.append(RuleEnsemble(ttnationalite, 1))
regle.append(RuleEnsemble(ttboisson, 2))
regle.append(RuleEnsemble(ttcigare, 3))
regle.append(RuleEnsemble(ttanimal, 4))
for r in regle:
print(r)
```
Parmi tous ces cas possibles, beaucoup sont incompatibles. L'objectif est d'éliminer tous ceux qui sont incompatibles pour ne garer que les 25 qui constituent la solution. L'algorithme est inspiré de la [logique des prédicats](http://fr.wikipedia.org/wiki/Calcul_des_pr%C3%A9dicats). De manière récursive, la fonction ``solve`` combine les clauses jusqu'à ce qu'il ne puisse plus continuer :
1. Soit le même attribut apparaît à deux positions différentes : incompatibilité.
2. Soit deux attributs apparaissent à la même position : incompatibilité.
3. Soit il ne reste plus qu'une seule clause : c'est la solution.
```
class Enigma:
"""
This class solves the enigma.
"""
def __init__(self, display=True):
"""
We describe the enigma using the classes we defined above.
:param display: if True, use print to print some information
"""
self.regle = []
self.regle.append(RulePosition(self.find("lait"), 2))
self.regle.append(RulePosition(self.find("norvegien"), 0))
self.regle.append(RuleEquivalence(self.find("Pall Mall"), self.find("oiseaux")))
self.regle.append(RuleEquivalence(self.find("anglais"), self.find("rouge")))
self.regle.append(RuleEquivalence(self.find("suedois"), self.find("chiens")))
self.regle.append(RuleEquivalence(self.find("danois"), self.find("the")))
self.regle.append(RuleEquivalence(self.find("vert"), self.find("cafe")))
self.regle.append(RuleEquivalence(self.find("jaune"), self.find("Dunhill")))
self.regle.append(RuleEquivalence(self.find("biere"), self.find("Bluemaster")))
self.regle.append(RuleEquivalence(self.find("allemand"), self.find("Prince")))
self.regle.append(RuleVoisin(self.find("Dunhill"), self.find("cheval")))
self.regle.append(RuleVoisin(self.find("norvegien"), self.find("bleu")))
self.regle.append(RuleVoisin(self.find("Blend"), self.find("eau")))
self.regle.append(RuleVoisin(self.find("Blend"), self.find("chats")))
self.regle.append(RuleAvant(self.find("vert"), self.find("blanc")))
self.regle.append(RuleEnsemble(ttcouleur, 0))
self.regle.append(RuleEnsemble(ttnationalite, 1))
self.regle.append(RuleEnsemble(ttboisson, 2))
self.regle.append(RuleEnsemble(ttcigare, 3))
self.regle.append(RuleEnsemble(ttanimal, 4))
for r in self.regle:
r.clauses = r.genere()
r.utilise = False
self.count = 0
def find(self, p):
"""
Finds a clause in the different sets of clause (houses, colors, ...).
:param p: clause
:return: tuple (clause, position)
"""
for i in range(0, len(ensemble)):
if p in ensemble[i]:
return (p, i)
return None
def to_dataframe(self):
sr = []
matrix = [list(" " * 5) for _ in range(0, 5)]
for row in self.solution:
i = row[0]
j = row[1][1]
s = row[1][0]
matrix[i][j] = s
for row in matrix:
sr.append(", ".join(row))
text = "\n".join(sr)
return read_csv(StringIO(text), header=None)
def solve(self, solution=[], logf=print): # solution = [ ]) :
"""
Solves the enigma by eploring in deepness,
the method is recursive
:param solution: `[]` empty at the beginning, recursively used then
:return: solution
"""
self.count += 1
if self.count % 10 == 0:
logf("*", self.count, " - properties in place : ", len(solution)-1)
if len(solution) == 25:
# we know the solution must contain 25 clauses,
# if are here than the problem is solved unless some incompatibility
for r in self.regle:
cl = r.combine_cross_sets([solution], r.clauses)
if cl == None or len(cl) == 0:
# the solution is incompatible with a solution
return None
self.solution = solution
return solution
# we are looking for the rule which generates the least possible clauses
# in order to reduce the number of possibilities as much as possible
# the research could be represented as a tree, we avoid creating two many paths
best = None
rule = None
for r in self.regle:
cl = r.combine_cross_sets([solution], r.clauses)
if cl == None:
# the solution is incompatible with a solution
return None
# we check rule r is bringing back some results
for c in cl:
if len(c) > len(solution):
break
else:
cl = None
if cl != None and (best == None or len(best) > len(cl)):
best = cl
rule = r
if best == None:
# the solution is incompatible with a solution
return None
rule.utilise = True
# we test all clauses
for c in best:
r = self.solve(c, logf=logf)
if r != None:
# we found
return r
rule.utilise = False # impossible
return None
en = Enigma()
en.solve()
en.to_dataframe()
```
| github_jupyter |
```
import os
import math
import pandas as pd
import numpy as np
import seaborn as sns
from pandas import datetime
from matplotlib import pyplot as plt
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import r2_score
## convert one to multiple series
def lag_ahead_series(data, n_in=1, n_out=1, n_vars = 1,dropnan=True):
df = pd.DataFrame(data)
cols, names = list(), list()
# input sequence (t-n, ... t-1)
for j in range(n_vars):
for i in range(n_in, 0, -1):
cols.append(df.iloc[:,j].shift(i))
names.append('{}{}(t-{})'.format(df.columns[0],j+1, i))
# forecast sequence (t+1, ... t+n)
for j in range(n_vars):
for i in range(0, n_out):
cols.append(df.iloc[:,j].shift(-i))
names += [('{}{}(t+{})'.format(df.columns[0],j+1, i)) ]
# put it all together
agg = pd.concat(cols, axis=1)
agg.columns = names
#drop rows with NaN values
if dropnan:
agg.dropna(inplace=True)
return agg
## Distribution plot funciton
def distri_plot(df):
f, axes = plt.subplots(3, 3, figsize=(15, 11), sharex=False)
for idx, col_name in enumerate(df.columns, 0):
idx = int(idx)
## jump to plotting energy
if(col_name == "rain"):
sns.distplot(df["energy"],ax=axes[2,2])
return
sns.distplot(df[col_name],ax=axes[idx//3,idx%3])
## plot
plt.tight_layout()
## Scatter plot function
def scatter_plot(df):
f, axes = plt.subplots(4, 2, figsize=(15, 11), sharex=False)
for idx, col_name in enumerate(df.columns, 0):
idx = int(idx)
if(idx >= 8):
return
## jump to plotting energy
sns.scatterplot(x= col_name,y = "energy", data = df, ax=axes[idx//2,idx%2])
## plot
plt.tight_layout()
## plot dataframe creation
def plot_df(arr, name):
plot_df = pd.DataFrame()
i = 0
for row in arr:
plot_df.insert(i, "{}_{}".format(name,i), row, True)
i += 1
return plot_df
def get_eval(y, yhat):
print("MSE: {}".format(mean_squared_error(y,yhat)))
print("MAE: {}".format(mean_absolute_error(y,yhat)))
print("r2_score: {}".format(r2_score(y,yhat, multioutput = "variance_weighted")))
## feature/ target construction fucntion with lag variable
def feature_target_construct(df, load_lag, target_ahead, temp_ahead, wd_on = False):
tempcols = ['temperature','temperature.1','temperature.2','temperature.3']
load = df['TotalLoad']
f_temp = pd.DataFrame()
for col in tempcols:
temp = lag_ahead_series(df[col],
n_in = 0,
n_out = temp_ahead + 1,
n_vars = 1,
dropnan = True)
f_temp = pd.concat([f_temp, temp], axis = 1)
t = lag_ahead_series(load,
n_in = 0,
n_out = target_ahead + 1,
n_vars = 1,
dropnan = True)
if(target_ahead > temp_ahead):
num_ahead = target_ahead
f_temp = f_temp.iloc[load_lag:-num_ahead + temp_ahead,:]
t = t.iloc[load_lag:,:]
elif(target_ahead < temp_ahead):
num_ahead = temp_ahead
f_temp = f_temp.iloc[load_lag:,:]
t = t.iloc[load_lag:-num_ahead + target_ahead,:]
else:
num_ahead = temp_ahead
f_temp = f_temp.iloc[load_lag:,:]
t = t.iloc[load_lag:,:]
## load lag series
f_load = lag_ahead_series(load,
n_in = load_lag,
n_out = 0,
n_vars = 1,
dropnan = True).iloc[:-num_ahead,:]
## feature concatanation
if wd_on:
weekday = pd.get_dummies(df.iloc[load_lag:-num_ahead,-1])
f = pd.concat([weekday, f_temp, f_load], axis = 1)
else:
f = pd.concat([f_temp, f_load], axis = 1)
## target load values
return f, t
train = pd.read_csv("../data/train_elia.csv", index_col= 'time')
test = pd.read_csv("../data/test_elia.csv", index_col= 'time')
# load_lag, target_ahead , temp_ahead, weekday_on
f_train, t_train = feature_target_construct(train, 4, 4, 4, True)
f_test, t_test = feature_target_construct(test, 4, 4, 4, True)
```
### TPOT: 6 hour ahead, 15 mins
```
from tpot import TPOTRegressor
from mmm import mul_reg_config_dict
from sklearn.model_selection import TimeSeriesSplit
#import mul_config as mc
train_X, val_X, train_y, val_y = train_test_split(f_train, t_train, train_size = 0.1, shuffle = False)
tpot_reg = TPOTRegressor(generations=20,
population_size=60,
offspring_size=None,
mutation_rate=0.9,
crossover_rate=0.1,
scoring='neg_mean_squared_error',
cv=TimeSeriesSplit(n_splits = 5),
subsample=1.0,
n_jobs=4,
max_time_mins=None,
max_eval_time_mins=5,
random_state=123,
config_dict=mul_reg_config_dict,
template=None,
warm_start=False,
memory=None,
use_dask=False,
periodic_checkpoint_folder=None,
early_stop=None,
verbosity=0,
disable_update_check=False)
tpot_reg.fit(train_X , train_y)
#val_X, val_y = train.iloc[500:600,:8], ahead_e.iloc[500:600,:]
yhat = tpot_reg.predict(f_test)
```
### Result Evaluation
```
get_eval(t_test, yhat)
## assignment
real = t_test.to_numpy()
guess = yhat
real = real[1:2,:]
guess = guess[1:2,:]
rpdf = plot_df(real, "observed")
gpdf = plot_df(guess, "prediction")
#plot
ax = plt.gca()
gpdf.plot(figsize=(25,10), colormap = 'plasma',style='--x',legend = True, ax = ax)
rpdf.plot(figsize=(25,10), color = 'g',style ='-o',legend = True, ax = ax, lw = 4)
ax.legend(frameon=False, loc='upper right', ncol=6, prop={'size': 16})
plt.show()
```
### TPOT: 24 hour ahead, half-hourly
```
from tpot import TPOTRegressor
from tpot_mulr import mul_reg_config_dict
from sklearn.model_selection import train_test_split
#import mul_config as mc
train_X, val_X, train_y, val_y = train_test_split(ahead_w, ahead_e, train_size = 0.05, test_size = 0.95, random_state = 123)
tpot_reg = TPOTRegressor(generations=30,
population_size=60,
n_jobs=4,
verbosity=2,
random_state=123,
subsample= 0.8,
config_dict=mul_reg_config_dict)
tpot_reg.fit(train_X , train_y)
yhat = tpot_reg.predict(test_X)
mean_squared_error(test_y, yhat)
mean_absolute_error(test_y, yhat)
r2_score(test_y, yhat)
## assignment
real = test_y.to_numpy()
guess = yhat
real = real[50:51,:49]
guess = guess[50:51,:49]
rpdf = plot_df(real, "observed")
gpdf = plot_df(guess, "prediction")
#plot
ax = plt.gca()
gpdf.plot(figsize=(25,10), colormap = 'plasma',style='--',legend = True, ax = ax)
rpdf.plot(figsize=(25,10), color = 'g',style ='-o',legend = True, ax = ax, lw = 4)
ax.legend(frameon=False, loc='upper right', ncol=6, prop={'size': 16})
plt.show()
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# NumPy API on TensorFlow
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/tf_numpy"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/tf_numpy.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/tf_numpy.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/tf_numpy.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
## Overview
TensorFlow implements a subset of the [NumPy API](https://numpy.org/doc/1.16), available as `tf.experimental.numpy`. This allows running NumPy code, accelerated by TensorFlow, while also allowing access to all of TensorFlow's APIs.
## Setup
```
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import tensorflow.experimental.numpy as tnp
import timeit
print("Using TensorFlow version %s" % tf.__version__)
```
## TensorFlow NumPy ND array
An instance of `tf.experimental.numpy.ndarray`, called **ND Array**, represents a multidimensional dense array of a given `dtype` placed on a certain device. Each one of these objects internally wraps a `tf.Tensor`. Check out the ND array class for useful methods like `ndarray.T`, `ndarray.reshape`, `ndarray.ravel` and others.
First create an ND array object, and then invoke different methods.
```
# Create an ND array and check out different attributes.
ones = tnp.ones([5, 3], dtype=tnp.float32)
print("Created ND array with shape = %s, rank = %s, "
"dtype = %s on device = %s\n" % (
ones.shape, ones.ndim, ones.dtype, ones.data.device))
# Check out the internally wrapped `tf.Tensor` object.
print("The ND array wraps a tf.Tensor: %s\n" % ones.data)
# Try commonly used member functions.
print("ndarray.T has shape %s" % str(ones.T.shape))
print("narray.reshape(-1) has shape %s" % ones.reshape(-1).shape)
```
### Type promotion
TensorFlow NumPy APIs have well-defined semantics for converting literals to ND array, as well as for performing type promotion on ND array inputs. Please see [`np.result_type`](https://numpy.org/doc/1.16/reference/generated/numpy.result_type.html) for more details. When converting literals to ND array, NumPy prefers wide types like `tnp.int64` and `tnp.float64`.
In contrast, `tf.convert_to_tensor` prefers `tf.int32` and `tf.float32` types for converting constants to `tf.Tensor`. TensorFlow APIs leave `tf.Tensor` inputs unchanged and do not perform type promotion on them.
In the next example, you will perform type promotion. First, run addition on ND array inputs of different types and note the output types. None of these type promotions would be allowed on straight `tf.Tensor` objects. Finally,
convert literals to ND array using `ndarray.asarray` and note the resulting type.
```
print("Type promotion for operations")
values = [tnp.asarray(1, dtype=d) for d in
(tnp.int32, tnp.int64, tnp.float32, tnp.float64)]
for i, v1 in enumerate(values):
for v2 in values[i + 1:]:
print("%s + %s => %s" % (v1.dtype, v2.dtype, (v1 + v2).dtype))
print("Type inference during array creation")
print("tnp.asarray(1).dtype == tnp.%s" % tnp.asarray(1).dtype)
print("tnp.asarray(1.).dtype == tnp.%s\n" % tnp.asarray(1.).dtype)
```
### Broadcasting
Similar to TensorFlow, NumPy defines rich semantics for "broadcasting" values.
You can check out the [NumPy broadcasting guide](https://numpy.org/doc/1.16/user/basics.broadcasting.html) for more information and compare this with [TensorFlow broadcasting semantics](https://www.tensorflow.org/guide/tensor#broadcasting).
```
x = tnp.ones([2, 3])
y = tnp.ones([3])
z = tnp.ones([1, 2, 1])
print("Broadcasting shapes %s, %s and %s gives shape %s" % (
x.shape, y.shape, z.shape, (x + y + z).shape))
```
### Indexing
NumPy defines very sophisticated indexing rules. See the [NumPy Indexing guide](https://numpy.org/doc/1.16/reference/arrays.indexing.html). Note the use of ND arrays as indices below.
```
x = tnp.arange(24).reshape(2, 3, 4)
print("Basic indexing")
print(x[1, tnp.newaxis, 1:3, ...], "\n")
print("Boolean indexing")
print(x[:, (True, False, True)], "\n")
print("Advanced indexing")
print(x[1, (0, 0, 1), tnp.asarray([0, 1, 1])])
# Mutation is currently not supported
try:
tnp.arange(6)[1] = -1
except TypeError:
print("Currently, TensorFlow NumPy does not support mutation.")
```
### Example Model
Next, you can see how to create a model and run inference on it. This simple model applies a relu layer followed by a linear projection. Later sections will show how to compute gradients for this model using TensorFlow's `GradientTape`.
```
class Model(object):
"""Model with a dense and a linear layer."""
def __init__(self):
self.weights = None
def predict(self, inputs):
if self.weights is None:
size = inputs.shape[1]
# Note that type `tnp.float32` is used for performance.
stddev = tnp.sqrt(size).astype(tnp.float32)
w1 = tnp.random.randn(size, 64).astype(tnp.float32) / stddev
bias = tnp.random.randn(64).astype(tnp.float32)
w2 = tnp.random.randn(64, 2).astype(tnp.float32) / 8
self.weights = (w1, bias, w2)
else:
w1, bias, w2 = self.weights
y = tnp.matmul(inputs, w1) + bias
y = tnp.maximum(y, 0) # Relu
return tnp.matmul(y, w2) # Linear projection
model = Model()
# Create input data and compute predictions.
print(model.predict(tnp.ones([2, 32], dtype=tnp.float32)))
```
## TensorFlow NumPy and NumPy
TensorFlow NumPy implements a subset of the full NumPy spec. While more symbols will be added over time, there are systematic features that will not be supported in the near future. These include NumPy C API support, Swig integration, Fortran storage order, views and `stride_tricks`, and some `dtype`s (like `np.recarray` and `np.object`). For more details, please see the [TensorFlow NumPy API Documentation](https://www.tensorflow.org/api_docs/python/tf/experimental/numpy).
### NumPy interoperability
TensorFlow ND arrays can interoperate with NumPy functions. These objects implement the `__array__` interface. NumPy uses this interface to convert function arguments to `np.ndarray` values before processing them.
Similarly, TensorFlow NumPy functions can accept inputs of different types including `tf.Tensor` and `np.ndarray`. These inputs are converted to an ND array by calling `ndarray.asarray` on them.
Conversion of the ND array to and from `np.ndarray` may trigger actual data copies. Please see the section on [buffer copies](#buffer-copies) for more details.
```
# ND array passed into NumPy function.
np_sum = np.sum(tnp.ones([2, 3]))
print("sum = %s. Class: %s" % (float(np_sum), np_sum.__class__))
# `np.ndarray` passed into TensorFlow NumPy function.
tnp_sum = tnp.sum(np.ones([2, 3]))
print("sum = %s. Class: %s" % (float(tnp_sum), tnp_sum.__class__))
# It is easy to plot ND arrays, given the __array__ interface.
labels = 15 + 2 * tnp.random.randn(1000)
_ = plt.hist(labels)
```
### Buffer copies
Intermixing TensorFlow NumPy with NumPy code may trigger data copies. This is because TensorFlow NumPy has stricter requirements on memory alignment than those of NumPy.
When a `np.ndarray` is passed to TensorFlow NumPy, it will check for alignment requirements and trigger a copy if needed. When passing an ND array CPU buffer to NumPy, generally the buffer will satisfy alignment requirements and NumPy will not need to create a copy.
ND arrays can refer to buffers placed on devices other than the local CPU memory. In such cases, invoking a NumPy function will trigger copies across the network or device as needed.
Given this, intermixing with NumPy API calls should generally be done with caution and the user should watch out for overheads of copying data. Interleaving TensorFlow NumPy calls with TensorFlow calls is generally safe and avoids copying data. See the section on [TensorFlow interoperability](#tensorflow-interoperability) for more details.
### Operator precedence
TensorFlow NumPy defines an `__array_priority__` higher than NumPy's. This means that for operators involving both ND array and `np.ndarray`, the former will take precedence, i.e., `np.ndarray` input will get converted to an ND array and the TensorFlow NumPy implementation of the operator will get invoked.
```
x = tnp.ones([2]) + np.ones([2])
print("x = %s\nclass = %s" % (x, x.__class__))
```
## TF NumPy and TensorFlow
TensorFlow NumPy is built on top of TensorFlow and hence interoperates seamlessly with TensorFlow.
### `tf.Tensor` and ND array
ND array is a thin wrapper on `tf.Tensor`. These types can be converted cheaply to one another without triggering actual data copies.
```
x = tf.constant([1, 2])
# Convert `tf.Tensor` to `ndarray`.
tnp_x = tnp.asarray(x)
print(tnp_x)
# Convert `ndarray` to `tf.Tensor` can be done in following ways.
print(tnp_x.data)
print(tf.convert_to_tensor(tnp_x))
# Note that tf.Tensor.numpy() will continue to return `np.ndarray`.
print(x.numpy(), x.numpy().__class__)
```
### TensorFlow interoperability
An ND array can be passed to TensorFlow APIs. These calls internally convert ND array inputs to `tf.Tensor`. As mentioned earlier, such conversion does not actually do data copies, even for data placed on accelerators or remote devices.
Conversely, `tf.Tensor` objects can be passed to `tf.experimental.numpy` APIs. These inputs will internally be converted to an ND array without performing data copies.
```
# ND array passed into TensorFlow function.
# This returns a `tf.Tensor`.
tf_sum = tf.reduce_sum(tnp.ones([2, 3], tnp.float32))
print("Output = %s" % tf_sum)
# `tf.Tensor` passed into TensorFlow NumPy function.
# This returns an ND array.
tnp_sum = tnp.sum(tf.ones([2, 3]))
print("Output = %s" % tnp_sum)
```
#### Operator precedence
When ND array and `tf.Tensor` objects are combined using operators, a precedence rule is used to determine which object executes the operator. This is controlled by the `__array_priority__` value defined by these classes.
`tf.Tensor` defines an `__array_priority__` higher than that of ND array. This means that the ND array input will be converted to `tf.Tensor` and the `tf.Tensor` version of the operator will be called.
The code below demonstrates how that affects the output type.
```
x = tnp.ones([2, 2]) + tf.ones([2, 1])
print("x = %s\nClass = %s" % (x, x.__class__))
```
### Gradients and Jacobians: tf.GradientTape
TensorFlow's GradientTape can be used for backpropagation through TensorFlow and TensorFlow NumPy code. GradientTape APIs can also return ND array outputs.
Use the model created in [Example Model](#example-model) section, and compute gradients and jacobians.
```
def create_batch(batch_size=32):
"""Creates a batch of input and labels."""
return (tnp.random.randn(batch_size, 32).astype(tnp.float32),
tnp.random.randn(batch_size, 2).astype(tnp.float32))
def compute_gradients(model, inputs, labels):
"""Computes gradients of squared loss between model prediction and labels."""
with tf.GradientTape() as tape:
assert model.weights is not None
# Note that `model.weights` need to be explicitly watched since they
# are not tf.Variables.
tape.watch(model.weights)
# Compute prediction and loss
prediction = model.predict(inputs)
loss = tnp.sum(tnp.square(prediction - labels))
# This call computes the gradient through the computation above.
return tape.gradient(loss, model.weights)
inputs, labels = create_batch()
gradients = compute_gradients(model, inputs, labels)
# Inspect the shapes of returned gradients to verify they match the
# parameter shapes.
print("Parameter shapes:", [w.shape for w in model.weights])
print("Gradient shapes:", [g.shape for g in gradients])
# Verify that gradients are of type ND array.
assert isinstance(gradients[0], tnp.ndarray)
# Computes a batch of jacobians. Each row is the jacobian of an element in the
# batch of outputs w.r.t. the corresponding input batch element.
def prediction_batch_jacobian(inputs):
with tf.GradientTape() as tape:
tape.watch(inputs)
prediction = model.predict(inputs)
return prediction, tape.batch_jacobian(prediction, inputs)
inp_batch = tnp.ones([16, 32], tnp.float32)
output, batch_jacobian = prediction_batch_jacobian(inp_batch)
# Note how the batch jacobian shape relates to the input and output shapes.
print("Output shape: %s, input shape: %s" % (output.shape, inp_batch.shape))
print("Batch jacobian shape:", batch_jacobian.shape)
```
### Trace compilation: tf.function
TensorFlow's `tf.function` works by "trace compiling" the code and then optimizing these traces for much faster performance. See the [Introduction to Graphs and Functions](./intro_to_graphs.ipynb).
`tf.function` can be used to optimize TensorFlow NumPy code as well. Here is a simple example to demonstrate the speedups. Note that the body of `tf.function` code includes calls to TensorFlow NumPy APIs, and the inputs and output are ND arrays.
```
inputs, labels = create_batch(512)
print("Eager performance")
compute_gradients(model, inputs, labels)
print(timeit.timeit(lambda: compute_gradients(model, inputs, labels),
number=10) * 100, "ms")
print("\ntf.function compiled performance")
compiled_compute_gradients = tf.function(compute_gradients)
compiled_compute_gradients(model, inputs, labels) # warmup
print(timeit.timeit(lambda: compiled_compute_gradients(model, inputs, labels),
number=10) * 100, "ms")
```
### Vectorization: tf.vectorized_map
TensorFlow has inbuilt support for vectorizing parallel loops, which allows speedups of one to two orders of magnitude. These speedups are accessible via the `tf.vectorized_map` API and apply to TensorFlow NumPy code as well.
It is sometimes useful to compute the gradient of each output in a batch w.r.t. the corresponding input batch element. Such computation can be done efficiently using `tf.vectorized_map` as shown below.
```
@tf.function
def vectorized_per_example_gradients(inputs, labels):
def single_example_gradient(arg):
inp, label = arg
return compute_gradients(model,
tnp.expand_dims(inp, 0),
tnp.expand_dims(label, 0))
# Note that a call to `tf.vectorized_map` semantically maps
# `single_example_gradient` over each row of `inputs` and `labels`.
# The interface is similar to `tf.map_fn`.
# The underlying machinery vectorizes away this map loop which gives
# nice speedups.
return tf.vectorized_map(single_example_gradient, (inputs, labels))
batch_size = 128
inputs, labels = create_batch(batch_size)
per_example_gradients = vectorized_per_example_gradients(inputs, labels)
for w, p in zip(model.weights, per_example_gradients):
print("Weight shape: %s, batch size: %s, per example gradient shape: %s " % (
w.shape, batch_size, p.shape))
# Benchmark the vectorized computation above and compare with
# unvectorized sequential computation using `tf.map_fn`.
@tf.function
def unvectorized_per_example_gradients(inputs, labels):
def single_example_gradient(arg):
inp, label = arg
return compute_gradients(model,
tnp.expand_dims(inp, 0),
tnp.expand_dims(label, 0))
return tf.map_fn(single_example_gradient, (inputs, labels),
fn_output_signature=(tf.float32, tf.float32, tf.float32))
print("Running vectorized computation")
print(timeit.timeit(lambda: vectorized_per_example_gradients(inputs, labels),
number=10) * 100, "ms")
print("\nRunning unvectorized computation")
per_example_gradients = unvectorized_per_example_gradients(inputs, labels)
print(timeit.timeit(lambda: unvectorized_per_example_gradients(inputs, labels),
number=10) * 100, "ms")
```
### Device placement
TensorFlow NumPy can place operations on CPUs, GPUs, TPUs and remote devices. It uses standard TensorFlow mechanisms for device placement. Below a simple example shows how to list all devices and then place some computation on a particular device.
TensorFlow also has APIs for replicating computation across devices and performing collective reductions which will not be covered here.
#### List devices
`tf.config.list_logical_devices` and `tf.config.list_physical_devices` can be used to find what devices to use.
```
print("All logical devices:", tf.config.list_logical_devices())
print("All physical devices:", tf.config.list_physical_devices())
# Try to get the GPU device. If unavailable, fallback to CPU.
try:
device = tf.config.list_logical_devices(device_type="GPU")[0]
except IndexError:
device = "/device:CPU:0"
```
#### Placing operations: **`tf.device`**
Operations can be placed on a device by calling it in a `tf.device` scope.
```
print("Using device: %s" % str(device))
# Run operations in the `tf.device` scope.
# If a GPU is available, these operations execute on the GPU and outputs are
# placed on the GPU memory.
with tf.device(device):
prediction = model.predict(create_batch(5)[0])
print("prediction is placed on %s" % prediction.data.device)
```
#### Copying ND arrays across devices: **`tnp.copy`**
A call to `tnp.copy`, placed in a certain device scope, will copy the data to that device, unless the data is already on that device.
```
with tf.device("/device:CPU:0"):
prediction_cpu = tnp.copy(prediction)
print(prediction.data.device)
print(prediction_cpu.data.device)
```
## Performance comparisons
TensorFlow NumPy uses highly optimized TensorFlow kernels that can be dispatched on CPUs, GPUs and TPUs. TensorFlow also performs many compiler optimizations, like operation fusion, which translate to performance and memory improvements. See [TensorFlow graph optimization with Grappler](./graph_optimization.ipynb) to learn more.
However TensorFlow has higher overheads for dispatching operations compared to NumPy. For workloads composed of small operations (less than about 10 microseconds), these overheads can dominate the runtime and NumPy could provide better performance. For other cases, TensorFlow should generally provide better performance.
Run the benchmark below to compare NumPy and TensorFlow NumPy performance for different input sizes.
```
def benchmark(f, inputs, number=30, force_gpu_sync=False):
"""Utility to benchmark `f` on each value in `inputs`."""
times = []
for inp in inputs:
def _g():
if force_gpu_sync:
one = tnp.asarray(1)
f(inp)
if force_gpu_sync:
with tf.device("CPU:0"):
tnp.copy(one) # Force a sync for GPU case
_g() # warmup
t = timeit.timeit(_g, number=number)
times.append(t * 1000. / number)
return times
def plot(np_times, tnp_times, compiled_tnp_times, has_gpu, tnp_times_gpu):
"""Plot the different runtimes."""
plt.xlabel("size")
plt.ylabel("time (ms)")
plt.title("Sigmoid benchmark: TF NumPy vs NumPy")
plt.plot(sizes, np_times, label="NumPy")
plt.plot(sizes, tnp_times, label="TF NumPy (CPU)")
plt.plot(sizes, compiled_tnp_times, label="Compiled TF NumPy (CPU)")
if has_gpu:
plt.plot(sizes, tnp_times_gpu, label="TF NumPy (GPU)")
plt.legend()
# Define a simple implementation of `sigmoid`, and benchmark it using
# NumPy and TensorFlow NumPy for different input sizes.
def np_sigmoid(y):
return 1. / (1. + np.exp(-y))
def tnp_sigmoid(y):
return 1. / (1. + tnp.exp(-y))
@tf.function
def compiled_tnp_sigmoid(y):
return tnp_sigmoid(y)
sizes = (2 ** 0, 2 ** 5, 2 ** 10, 2 ** 15, 2 ** 20)
np_inputs = [np.random.randn(size).astype(np.float32) for size in sizes]
np_times = benchmark(np_sigmoid, np_inputs)
with tf.device("/device:CPU:0"):
tnp_inputs = [tnp.random.randn(size).astype(np.float32) for size in sizes]
tnp_times = benchmark(tnp_sigmoid, tnp_inputs)
compiled_tnp_times = benchmark(compiled_tnp_sigmoid, tnp_inputs)
has_gpu = len(tf.config.list_logical_devices("GPU"))
if has_gpu:
with tf.device("/device:GPU:0"):
tnp_inputs = [tnp.random.randn(size).astype(np.float32) for size in sizes]
tnp_times_gpu = benchmark(compiled_tnp_sigmoid, tnp_inputs, 100, True)
else:
tnp_times_gpu = None
plot(np_times, tnp_times, compiled_tnp_times, has_gpu, tnp_times_gpu)
```
## Further reading
- [TensorFlow NumPy: Distributed Image Classification Tutorial](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/ops/numpy_ops/g3doc/TensorFlow_Numpy_Distributed_Image_Classification.ipynb)
- [TensorFlow NumPy: Keras and Distribution Strategy](
https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/ops/numpy_ops/g3doc/TensorFlow_NumPy_Keras_and_Distribution_Strategy.ipynb)
- [Sentiment Analysis with Trax and TensorFlow NumPy](
https://github.com/google/trax/blob/master/trax/tf_numpy_and_keras.ipynb)
| github_jupyter |
```
from pymongo import MongoClient
import pymatgen
import bson
import itertools
import numpy as np
import collections
from bson import ObjectId
from pymatgen.core.structure import Structure
from pymatgen.vis.structure_vtk import EL_COLORS
M = MongoClient()
distortions = M.ferroelectric_dataset.distortions
workflow_data = M.ferroelectric_dataset.workflow_data
def create_metadata_for_unitcell(structure):
matrix = structure.lattice.matrix
lines = []
for i in itertools.product([0, 1], repeat=3):
for j in itertools.product([0, 1], repeat=3):
if ((np.max(np.array(j) - np.array(i)) == 1) and
(np.min(np.array(j) - np.array(i)) == 0) and
(np.sum(np.array(j) - np.array(i)) == 1)):
x1, y1, z1 = np.einsum('mn,m->n', matrix, i)
x2, y2, z2 = np.einsum('mn,m->n', matrix, j)
x, y, z = [x1, x2], [y1, y2], [z1, z2]
lines.append([x, y, z])
coords = structure.cart_coords
species = structure.species
atomic_numbers = [specie.number for specie in species]
species = list(map(str, species))
colors = ["rgb({},{},{})".format(*EL_COLORS["VESTA"][s]) for s in species]
xs, ys, zs = coords[:, 0].tolist(), coords[:, 1].tolist(), coords[:, 2].tolist()
data = zip(xs, ys, zs, species, colors)
atoms = collections.defaultdict(lambda: collections.defaultdict(list))
for d in data:
x, y, z, s, c = d
atoms[s]['x'].append(x)
atoms[s]['y'].append(y)
atoms[s]['z'].append(z)
# atoms = {'x': xs, 'y': ys, 'z': zs, 'species': species, 'atomic_numbers': atomic_numbers}
legend = list(set(zip(species, colors)))
legend = {s: c for s,c in legend}
return {'unitcell': lines, 'atoms': atoms, 'legend': legend }
def get_polarization_plot_data(polarization,
with_quanta=True,
abs_ylim_min=5.,
convert_to_muC_per_cm2=True):
polarization_plot_data = {axis: {} for axis in "abc"}
same_branch = np.asarray(polarization.get_same_branch_polarization_data(
convert_to_muC_per_cm2=convert_to_muC_per_cm2))
quanta = np.asarray(polarization.get_lattice_quanta(
convert_to_muC_per_cm2=convert_to_muC_per_cm2))
splines = polarization.same_branch_splines()
for i, axis in enumerate("abc"):
# Make symmetric axis range around zero.
ax_max = max(max(same_branch[:, i]), abs_ylim_min)
ax_min = min(min(same_branch[:, i]), -abs_ylim_min)
ax_range = max(abs(ax_min), ax_max)
polarization_plot_data[axis].update({"plot_ylim": (-ax_range, ax_range)})
polarization_plot_data[axis].update({"plot_xlim": (0, len(same_branch[:, i]) - 1)})
# Plot same branch polarization with quanta
polarization_plot_data[axis].update({'quanta': {'data': [], 'color': []}})
num_copies_quanta = 2 * int(np.ceil((ax_max - ax_min) / quanta[0, i]))
for j in range(-num_copies_quanta, num_copies_quanta + 1):
color = 'ro' if j == 0 else 'bo'
polarization_plot_data[axis]['quanta']['data'].append(
(same_branch[:, i].flatten() + j * np.array(quanta[:, i].flatten())).tolist())
polarization_plot_data[axis]['quanta']['color'].append(color)
# Plot same branch polarization spline
polarization_plot_data[axis].update({'splines': {}})
if splines[i]:
xs = np.linspace(0, len(same_branch[:, i]) - 1, 1000)
polarization_plot_data[axis]['splines'].update(
{'x': xs.tolist(), 'y': splines[i](xs).tolist()})
return polarization_plot_data
def get_polarization_from_workflow_data(wfid_str):
import pymatgen.analysis.ferroelectricity.polarization as polarization
entry = workflow_data.find_one({'wfid': wfid_str})
vasp_structs = list(map(pymatgen.Structure.from_dict, entry['structures']))
vasp_pelecs = entry['raw_electron_polarization']
vasp_pions = entry['raw_ionic_polarization']
return polarization.Polarization(vasp_pelecs, vasp_pions, vasp_structs,p_elecs_in_cartesian=False)
def get_scalar_plot_dict(scalars):
import pymatgen.analysis.ferroelectricity.polarization as polarization
energy_trend = polarization.EnergyTrend(scalars)
spline = energy_trend.spline()
spline_xs = np.linspace(0, len(scalars)-1, num=50)
spline_ys = spline(spline_xs)
return {'scalars': scalars, 'spline': {'x': spline_xs.tolist(), 'y': spline_ys.tolist()}}
def get_task_tables(wfid_str):
all_labels = collections.OrderedDict()
# Create default table of False values
relaxation_task_labels = collections.OrderedDict()
relaxation_task_labels.update({'_nonpolar_relaxation': False})
relaxation_task_labels.update({'_polar_relaxation': False})
static_task_labels = collections.OrderedDict()
static_task_labels.update({'_nonpolar_static': False})
for i in range(8,0,-1):
static_task_labels.update({'_interpolation_{}_static'.format(i): False})
static_task_labels.update({'_polar_static': False})
polarization_task_labels = collections.OrderedDict()
polarization_task_labels.update({'_nonpolar_polarization': False})
for i in range(8,0,-1):
polarization_task_labels.update({'_interpolation_{}_polarization'.format(i): False})
polarization_task_labels.update({'_polar_polarization': False})
all_labels.update({'relaxation_task_labels': relaxation_task_labels})
all_labels.update({'static_task_labels': static_task_labels})
all_labels.update({'polarization_task_labels': polarization_task_labels})
# Update default table with Trues
entry = workflow_data.find_one({'wfid': wfid_str})
relax_labels = [] if 'relaxation_task_labels' not in entry else entry['relaxation_task_labels']
static_labels = [] if 'static_task_labels' not in entry else entry['static_task_labels']
pol_labels = [] if 'polarization_task_labels' not in entry else entry['polarization_task_labels']
for label in relax_labels:
all_labels['relaxation_task_labels'][label] = True
for label in static_labels:
all_labels['static_task_labels'][label] = True
for i, label in enumerate(pol_labels):
if entry['raw_electron_polarization'][i] is not None:
all_labels['polarization_task_labels'][label] = True
json_table_data = {key: [] for key in all_labels}
for key in all_labels:
for subkey in all_labels[key]:
json_table_data[key].append({"task": subkey, "complete": all_labels[key][subkey]})
return json_table_data
```
We use the following categories for the dataset:
smooth -- workflows with a high quality reconstructed polarization and energy trend
not smooth -- workflows that have all polarization calculations completed but reconstructed trends are not smooth
static_only -- workflows that are missing polarization data but otherwise completed
not_complete -- all other incomplete workflows (Includes DEFUSED, FIZZLED, and RUNNING workflow states)
```
smooth = workflow_data.find({'polarization_len' : 10,
'polarization_max_spline_jumps': {"$exists": True, "$not" : { "$gt" : 1 }},
'energies_per_atom_max_spline_jumps': {'$lte': 1e-2}})
not_smooth = workflow_data.find({'polarization_len' : 10,
'polarization_change_norm': {'$exists': True},
'$or': [{'polarization_max_spline_jumps': { "$gt" : 1 }},
{'energies_per_atom_max_spline_jumps': {'$gt': 1e-2}}]})
static = workflow_data.find({'polarization_change_norm': {'$exists': False},
'static_len': 10,
'$or': [{'workflow_status': 'COMPLETED'}, {'workflow_status': 'DEFUSED'}]})
defused_or_fizzled_or_running = workflow_data.find(
{'$and': [{'$or': [{'polarization_len' : {'$lt': 10}},
{'polarization_change_norm': {'$exists': False}}]},
{'$or': [
{'$and': [{'workflow_status': 'DEFUSED'}, {'static_len' : {'$lt': 10}}]},
{'workflow_status': 'FIZZLED'},
{'workflow_status': 'RUNNING'}]}]})
print ("smooth: ", smooth.count())
print ("not_smooth: ", not_smooth.count())
print ("static_only: ", static.count())
print ("not_complete: ", defused_or_fizzled_or_running.count())
def get_alphabetical_formula(pretty_formula):
from pymatgen.core.composition import Composition
alphabetical_formula = Composition(pretty_formula).alphabetical_formula.split()
# Remove 1s
alphabetical_formula = [string[:-1] if (string[-1] == "1" and string[-2] not in "0123456789") else string for string in alphabetical_formula]
# Stringify
alphabetical_formula = "".join(alphabetical_formula)
return alphabetical_formula
def make_candidate_json(pymongo_result, category):
for workflow_entry in pymongo_result:
distort = None
entry_dict = {workflow_entry['wfid']: {}}
#Magnetization data
if 'total_magnetization' in workflow_entry:
entry_dict[workflow_entry['wfid']].update({'total_magnetization': workflow_entry['total_magnetization']})
#Polarization data
if 'polarization_change_norm' in workflow_entry:
# Need to reach into distortion database to get structures
# for incomplete 'structures' array in workflow data.
entry_polarization = get_polarization_from_workflow_data(workflow_entry['wfid'])
entry_pol_plot_data = get_polarization_plot_data(entry_polarization)
entry_dict[workflow_entry['wfid']].update({'polarization': entry_pol_plot_data})
entry_dict[workflow_entry['wfid']].update({'polarization_change_norm': workflow_entry['polarization_change_norm']})
# Distortion data
if 'structures' not in workflow_entry or len(workflow_entry['structures']) < 10:
cid = workflow_entry['cid']
distort = distortions.find_one({'_id': ObjectId(cid[4:])})
nonpolar = Structure.from_dict(distort['distortion']['high_low_setting'])
polar = Structure.from_dict(distort['distortion']['low_symm'])
all_structs = nonpolar.interpolate(polar, nimages=9, interpolate_lattices=True)
else:
all_structs = list(map(pymatgen.core.structure.Structure.from_dict, workflow_entry['structures']))
entry_dict[workflow_entry['wfid']].update({'distortion' : list(map(create_metadata_for_unitcell, all_structs))})
# Energy data
if ('energies_per_atom' in workflow_entry and
'static_len' in workflow_entry and
workflow_entry['static_len'] == 10):
entry_dict[workflow_entry['wfid']].update(
{'energy_per_atom': get_scalar_plot_dict(workflow_entry['energies_per_atom'])})
entry_dict[workflow_entry['wfid']].update(
{
'nonpolar_id': workflow_entry['nonpolar_id'],
'polar_id': workflow_entry['polar_id'],
'nonpolar_spacegroup': workflow_entry['nonpolar_spacegroup'],
'polar_spacegroup': workflow_entry['polar_spacegroup'],
'pretty_formula': distort['pretty_formula'] if distort is not None else workflow_entry['pretty_formula'],
'alphabetical_formula': get_alphabetical_formula(distort['pretty_formula']) if distort is not None else workflow_entry['alphabetical_formula'],
'hubbards': None if 'hubbards' not in workflow_entry else workflow_entry['hubbards'],
'workflow_status': workflow_entry['workflow_status'],
'search_id': workflow_entry['search_id'],
})
entry_dict[workflow_entry['wfid']].update({'tasks' : get_task_tables(workflow_entry['wfid'])})
entry_dict[workflow_entry['wfid']].update({'category' : category})
import json
with open("../json/"+ workflow_entry['wfid']+'.json', 'w') as fp:
json.dump(entry_dict, fp, indent=2)
categories = ["smooth", "not_smooth", "static_only", "not_complete"]
pymongo_results = [smooth, not_smooth, static, defused_or_fizzled_or_running]
for pymongo_result, category in zip(pymongo_results, categories):
print ("category: ", category)
make_candidate_json(pymongo_result, category)
```
| github_jupyter |
## Chapter 11.3
```
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.layers import Embedding, LSTM, Dense, Bidirectional, Dropout
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import Adam
def plotGraph(history,title):
loss,acc = history.history.values()
epochs = range(1,len(loss)+1)
# Plot graph
plt.plot(epochs,acc,label = 'Training Accuracy')
plt.title('Training Accuracy of %s Model'%title)
plt.legend()
plt.figure()
plt.plot(epochs,loss,label='Training Loss')
plt.title('Training Loss of %s Model'%title)
plt.legend()
plt.figure()
def textGenerator(model, title):
seed_text = "I've got a bad feeling about this"
next_words = 100
for _ in range(next_words):
token_list = tokenizer.texts_to_sequences([seed_text])[0]
token_list = pad_sequences([token_list], maxlen=max_sequence_len-1, padding='pre')
predicted = model.predict_classes(token_list, verbose=0)
output_word = ""
for word, index in tokenizer.word_index.items():
if index == predicted:
output_word = word
break
seed_text += " " + output_word
print(title,'model:',seed_text)
data = open('irish-lyrics-eof.txt').read()
corpus = data.lower().split("\n")
tokenizer = Tokenizer()
tokenizer.fit_on_texts(corpus)
total_words = len(tokenizer.word_index) + 1
# print(tokenizer.word_index)
# print(total_words)
input_sequences = []
for line in corpus:
token_list = tokenizer.texts_to_sequences([line])[0]
for i in range(1, len(token_list)):
n_gram_sequence = token_list[:i+1]
input_sequences.append(n_gram_sequence)
# pad sequences
max_sequence_len = max([len(x) for x in input_sequences])
input_sequences = np.array(pad_sequences(input_sequences, maxlen=max_sequence_len, padding='pre'))
# create predictors and label
xs, labels = input_sequences[:,:-1],input_sequences[:,-1]
ys = tf.keras.utils.to_categorical(labels, num_classes=total_words)
model_1 = Sequential([
Embedding(total_words, 100, input_length=max_sequence_len-1),
Bidirectional(LSTM(150)),
Dense(total_words, activation='softmax'),
])
model_1.compile(loss='categorical_crossentropy', optimizer=Adam(lr=0.01), metrics=['accuracy'])
#earlystop = EarlyStopping(monitor='val_loss', min_delta=0, patience=5, verbose=0, mode='auto')
from tensorflow.keras import regularizers
# 还可以在全连接层中使用正则化。
model_2 = Sequential([
Embedding(total_words, 100, input_length=max_sequence_len-1),
Bidirectional(LSTM(150, return_sequences = True)),
Dense(total_words, activation='softmax'),
Dropout(0.2),
LSTM(100),
Dense(total_words/2, activation='relu', kernel_regularizer=regularizers.l2(0.01)),
Dense(total_words, activation='softmax')
])
model_2.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
history1 = model_1.fit(xs, ys, epochs=150, verbose=1)
plotGraph(history1,'Regularized')
textGenerator(model_1,'Regularized')
history2 = model_2.fit(xs, ys, epochs=150, verbose=1)
plotGraph(history2,'Regularized')
textGenerator(model_2,'Regularized')
```
| github_jupyter |
<img src="https://avatars.githubusercontent.com/u/74911464?s=200&v=4"
alt="OpenEO Platform logo"
style="float: left; margin-right: 10px;" />
## openEO Platform UC6 - Near Real Time Forest Dynamics
### Author michele.claus@eurac.edu
### Date: 2021/09/10
```
from eo_utils import *
```
## Connect to openEO
```
backend = "https://openeo.cloud"
conn = openeo.connect(backend)
```
### Authenticate
```
conn = conn.authenticate_oidc(provider_id="egi")
```
### Select the area of interest from the map using the rectangle tool. The selected area will be the one where you want to look for changes.
```
center = [46.35, 11.45]
zoom = 12
eoMap = openeoMap(center,zoom)
eoMap.map
bbox = eoMap.getBbox()
print("Coordinates selected from map:",'\n west',bbox[0],'\n east',bbox[2],'\n south',bbox[1],'\n north',bbox[3])
```
## Sentinel-2 data pre-processing (cloud masking, data clipping)
Sentinel-2 data loading
```
spatial_extent = {'west':bbox[0],'east':bbox[2],'south':bbox[1],'north':bbox[3]}
temporal_extent = ["2016-09-01", "2018-09-01"] # This is the training period
collection = 'boa_sentinel_2'
bands = ["B02","B03","B04","B05","B08"] # We could have also more bands
l2a_bands = conn.load_collection(collection,spatial_extent=spatial_extent,bands=bands,temporal_extent=temporal_extent)
```
Clip the data to avoid unmasked clouds
```
l2a_bands_clipped = l2a_bands.apply(lambda x: x.clip(0,5000))
```
# Seasonal curve fitting
The following formula models a seasonal harmonic behavior, typical of vegetation:
```
a0+a1*cos(2*pi/31557600*x)+a2*sin(2*pi/31557600*x)
```
31557600 are the seconds in one year. The values that x will assume are Unix timesteps (seconds from 01-01-1970).
The forests should follow this seasonal behaviour and the fitted function should get close to the real signal like in the following example:

## Curve fitting
```
def fitFunction(x:ProcessBuilder, parameters:ProcessBuilder):
t = 2 * math.pi / 31557600 * x
return parameters[0] + parameters[1] * cos(t) + parameters[2] * sin(t)
curve_fitting = l2a_bands_clipped.fit_curve(
parameters=[1,1,1], # Initial guess of the parameters
dimension="t", # Fit the function along the temporal dimension
function=fitFunction
)
```
Save the result parameters as a netCDF for later use and start the batch job
```
fitted_parameters = curve_fitting.save_result(format="NetCDF")
job = fitted_parameters.send_job(title = "S2_curve_fitting_SRR2")
jobIdFitting = job.job_id
job.start_job()
```
### Check the fitting job status and when it is marked as 'finished', you can continue
```
conn.job(jobIdFitting)
```
## Curve prediction and change detection threshold
Given the computed parameters stored in a netCDF from the previous job, we can now reuse them, predicting the values in the training period when following the sinuisoidal behavior.
With the difference between predicted and real data, we compute the RMSE over the training period and finally the standard deviation over time, resulting in a final raster with a single band and no time dimension.
Load result parameters from previous job
```
curve_fitting_loaded = conn.load_result(id=jobIdFitting)
```
Compute the predicted values
```
temporal_labels = l2a_bands_clipped.dimension_labels('t')
curve_prediction = l2a_bands_clipped.predict_curve(parameters=curve_fitting_loaded,dimension='t',function=fitFunction,labels=temporal_labels)
```
Mask out the cloud covered areas as in the original training data
Create a cloud mask based on the cloud masked areas produced by FORCE
```
force_cloud_mask = l2a_bands.apply(lambda x: x.gt(0))
predicted_masked = curve_prediction.merge_cubes(force_cloud_mask,overlap_resolver=multiply)
```
Subtract the predicted values from the pre-processed S-2 data
```
merged_cubes = l2a_bands_clipped.merge_cubes(predicted_masked,overlap_resolver=subtract)
```
Compute the RMSE
```
from change_detection_utils import compute_residual
bands = ["B02","B03","B04","B05","B08"]
rmse = compute_residual(merged_cubes,bands)
```
Compute the standard deviation of the residual error over time
```
rmse_std = rmse.reduce_dimension(dimension='t',reducer=sd)
```
Save the resulting layer as netCDF and start the batch job
```
rmse_std_netcdf = rmse_std.save_result(format="NetCDF")
job = rmse_std_netcdf.send_job(title = "S2_curve_predicting_SRR2")
jobIdPredicting = job.job_id
job.start_job()
```
### Sample result from the curve fitting and prediction steps
The following plot shows a pixel time series from the infrared band. In blue the raw signal and in orange the fitted seasonal trend

### Check the predicting job status and when it is marked as 'finished', you can continue
```
conn.job(jobIdPredicting)
# jobIdPredicting = ""
# jobIdFitting = ""
```
# Change detection
Given the fitted function to each pixel, we can find out how much the signal deviates from the model and flag it as a change when it goes beyond a threshold.
The following example shows how noticeable is the change in the SAR (Sentinel-1) signal after the Vaia storm of a forest area affected from it.

TIP: If you want to use the same bbox as a previously executed job, you can use the following function to retrieve the spatial extent, using the corresponding job id:
```
from change_detection_utils import get_bbox_from_job
job = conn.job(jobIdFitting)
spatial_extent = get_bbox_from_job(job)
print(spatial_extent)
```
Load the precomputed parameters
```
curve_fitting_loaded = conn.load_result(id=jobIdFitting)
```
Load the testing data and filter it in the same way as we did for training
```
spatial_extent = {'west':bbox[0],'east':bbox[2],'south':bbox[1],'north':bbox[3]}
temporal_extent = ["2018-10-01", "2019-03-01"] # This is the testing period
collection = 'boa_sentinel_2'
bands = ["B02","B03","B04","B05","B08"]
l2a_bands_test = conn.load_collection(collection,spatial_extent=spatial_extent,bands=bands,temporal_extent=temporal_extent)
```
Clip the data to avoid unmasked clouds
```
l2a_bands_test_clipped = l2a_bands_test.apply(lambda x: x.clip(0,5000))
```
## Curve prediction
Predict the values that the test data should have following the seasonal trend
```
temporal_labels_test = l2a_bands_test_clipped.dimension_labels('t')
curve_prediction_test = l2a_bands_test_clipped.predict_curve(parameters=curve_fitting_loaded,dimension='t',function=fitFunction,labels=temporal_labels_test)
```
Create a cloud mask based on the cloud masked areas produced by FORCE
```
force_cloud_mask_test = l2a_bands_test.apply(lambda x: x.gt(0))
predicted_masked_test = curve_prediction_test.merge_cubes(force_cloud_mask_test,overlap_resolver=multiply)
merged_cubes_test = l2a_bands_test_clipped.merge_cubes(predicted_masked_test,overlap_resolver=subtract)
```
Compute the residual error for each time step in the testing period
```
from change_detection_utils import compute_residual
bands = ["B02","B03","B04","B05","B08"]
residual = compute_residual(merged_cubes,bands)
```
### Set the thresold for the change detection
We raise and alarm if residual error > 3*std
It is not mandatory to use 3 as multiplicative factor, it depends on the signal and how big are deviations from the model that we want to detect.
Load precomputed threshold layer
```
training_std_loaded = conn.load_result(id=jobIdPredicting)
alarm_threshhold = training_std_loaded.apply(lambda x: x * 3)
alarm_cubes = residual.merge_cubes(alarm_threshhold,overlap_resolver=gt)
```
Save the resulting timeseries of change alarms as netCDF and start the batch job
```
alarm_cubes_netcdf = alarm_cubes.save_result(format="NetCDF")
job = alarm_cubes_netcdf.send_job(title = "S2_change_detection_SRR2")
jobIdAlarms = job.job_id
job.start_job()
#jobIdAlarms = "jb-3131a9f0-7caf-464d-befd-d52359c79839"
```
Check the change detection job status and when it is marked as 'finished', you can continue
```
conn.job(jobIdAlarms)
```
Download the results. Please note: each time step will be downloaded as a separate file
```
jobAlarms = conn.job(jobIdAlarms)
jobResults = jobAlarms.get_results()
jobResults.download_files('./results')
```
# Visualization of detected changes
We consider and visualize a detected change only if there are 3 subsequent changes detected. You can modify the value of subsequentAlarms with a bigger or smaller amount and see what changes.
If monthlyAggregate=True, we compute the aggregation of the detected changes for each month. If your testing period covers more than 12 months, the aggregation will be performed over all the data for the month (Jan,Feb,...), independently from which year.
```
from change_detection_utils import plot_detected_changes
plot_detected_changes(netcdfPath='./results/out_*.nc',monthlyAggregate=False,subsequentAlarms=3,backgroundTiles='OSM') #Try 'ESRI' as backgroundTiles for satellite RGB background
```
| github_jupyter |
# Introduction to Deep Learning with PyTorch
In this notebook, you'll get introduced to [PyTorch](http://pytorch.org/), a framework for building and training neural networks. PyTorch in a lot of ways behaves like the arrays you love from Numpy. These Numpy arrays, after all, are just tensors. PyTorch takes these tensors and makes it simple to move them to GPUs for the faster processing needed when training neural networks. It also provides a module that automatically calculates gradients (for backpropagation!) and another module specifically for building neural networks. All together, PyTorch ends up being more coherent with Python and the Numpy/Scipy stack compared to TensorFlow and other frameworks.
## Neural Networks
Deep Learning is based on artificial neural networks which have been around in some form since the late 1950s. The networks are built from individual parts approximating neurons, typically called units or simply "neurons." Each unit has some number of weighted inputs. These weighted inputs are summed together (a linear combination) then passed through an activation function to get the unit's output.
<img src="assets/simple_neuron.png" width=400px>
Mathematically this looks like:
$$
\begin{align}
y &= f(w_1 x_1 + w_2 x_2 + b) \\
y &= f\left(\sum_i w_i x_i +b \right)
\end{align}
$$
With vectors this is the dot/inner product of two vectors:
$$
h = \begin{bmatrix}
x_1 \, x_2 \cdots x_n
\end{bmatrix}
\cdot
\begin{bmatrix}
w_1 \\
w_2 \\
\vdots \\
w_n
\end{bmatrix}
$$
## Tensors
It turns out neural network computations are just a bunch of linear algebra operations on *tensors*, a generalization of matrices. A vector is a 1-dimensional tensor, a matrix is a 2-dimensional tensor, an array with three indices is a 3-dimensional tensor (RGB color images for example). The fundamental data structure for neural networks are tensors and PyTorch (as well as pretty much every other deep learning framework) is built around tensors.
<img src="assets/tensor_examples.svg" width=600px>
With the basics covered, it's time to explore how we can use PyTorch to build a simple neural network.
```
# First, import PyTorch
import torch
def activation(x):
""" Sigmoid activation function
Arguments
---------
x: torch.Tensor
"""
return 1/(1+torch.exp(-x))
### Generate some data
torch.manual_seed(7) # Set the random seed so things are predictable
# Features are 5 random normal variables
features = torch.randn((1, 5))
# True weights for our data, random normal variables again
weights = torch.randn_like(features)
# and a true bias term
bias = torch.randn((1, 1))
```
Above I generated data we can use to get the output of our simple network. This is all just random for now, going forward we'll start using normal data. Going through each relevant line:
`features = torch.randn((1, 5))` creates a tensor with shape `(1, 5)`, one row and five columns, that contains values randomly distributed according to the normal distribution with a mean of zero and standard deviation of one.
`weights = torch.randn_like(features)` creates another tensor with the same shape as `features`, again containing values from a normal distribution.
Finally, `bias = torch.randn((1, 1))` creates a single value from a normal distribution.
PyTorch tensors can be added, multiplied, subtracted, etc, just like Numpy arrays. In general, you'll use PyTorch tensors pretty much the same way you'd use Numpy arrays. They come with some nice benefits though such as GPU acceleration which we'll get to later. For now, use the generated data to calculate the output of this simple single layer network.
> **Exercise**: Calculate the output of the network with input features `features`, weights `weights`, and bias `bias`. Similar to Numpy, PyTorch has a [`torch.sum()`](https://pytorch.org/docs/stable/torch.html#torch.sum) function, as well as a `.sum()` method on tensors, for taking sums. Use the function `activation` defined above as the activation function.
```
## Calculate the output of this network using the weights and bias tensors
output = activation(torch.sum(weights*features) + bias)
output
```
You can do the multiplication and sum in the same operation using a matrix multiplication. In general, you'll want to use matrix multiplications since they are more efficient and accelerated using modern libraries and high-performance computing on GPUs.
Here, we want to do a matrix multiplication of the features and the weights. For this we can use [`torch.mm()`](https://pytorch.org/docs/stable/torch.html#torch.mm) or [`torch.matmul()`](https://pytorch.org/docs/stable/torch.html#torch.matmul) which is somewhat more complicated and supports broadcasting. If we try to do it with `features` and `weights` as they are, we'll get an error
```python
>> torch.mm(features, weights)
---------------------------------------------------------------------------
RuntimeError Traceback (most recent call last)
<ipython-input-13-15d592eb5279> in <module>()
----> 1 torch.mm(features, weights)
RuntimeError: size mismatch, m1: [1 x 5], m2: [1 x 5] at /Users/soumith/minicondabuild3/conda-bld/pytorch_1524590658547/work/aten/src/TH/generic/THTensorMath.c:2033
```
As you're building neural networks in any framework, you'll see this often. Really often. What's happening here is our tensors aren't the correct shapes to perform a matrix multiplication. Remember that for matrix multiplications, the number of columns in the first tensor must equal to the number of rows in the second column. Both `features` and `weights` have the same shape, `(1, 5)`. This means we need to change the shape of `weights` to get the matrix multiplication to work.
**Note:** To see the shape of a tensor called `tensor`, use `tensor.shape`. If you're building neural networks, you'll be using this method often.
There are a few options here: [`weights.reshape()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.reshape), [`weights.resize_()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.resize_), and [`weights.view()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.view).
* `weights.reshape(a, b)` will return a new tensor with the same data as `weights` with size `(a, b)` sometimes, and sometimes a clone, as in it copies the data to another part of memory.
* `weights.resize_(a, b)` returns the same tensor with a different shape. However, if the new shape results in fewer elements than the original tensor, some elements will be removed from the tensor (but not from memory). If the new shape results in more elements than the original tensor, new elements will be uninitialized in memory. Here I should note that the underscore at the end of the method denotes that this method is performed **in-place**. Here is a great forum thread to [read more about in-place operations](https://discuss.pytorch.org/t/what-is-in-place-operation/16244) in PyTorch.
* `weights.view(a, b)` will return a new tensor with the same data as `weights` with size `(a, b)`.
I usually use `.view()`, but any of the three methods will work for this. So, now we can reshape `weights` to have five rows and one column with something like `weights.view(5, 1)`.
> **Exercise**: Calculate the output of our little network using matrix multiplication.
```
## Calculate the output of this network using matrix multiplication
output = activation(torch.mm(weights, features.view(5,1)) + bias)
output
```
### Stack them up!
That's how you can calculate the output for a single neuron. The real power of this algorithm happens when you start stacking these individual units into layers and stacks of layers, into a network of neurons. The output of one layer of neurons becomes the input for the next layer. With multiple input units and output units, we now need to express the weights as a matrix.
<img src='assets/multilayer_diagram_weights.png' width=450px>
The first layer shown on the bottom here are the inputs, understandably called the **input layer**. The middle layer is called the **hidden layer**, and the final layer (on the right) is the **output layer**. We can express this network mathematically with matrices again and use matrix multiplication to get linear combinations for each unit in one operation. For example, the hidden layer ($h_1$ and $h_2$ here) can be calculated
$$
\vec{h} = [h_1 \, h_2] =
\begin{bmatrix}
x_1 \, x_2 \cdots \, x_n
\end{bmatrix}
\cdot
\begin{bmatrix}
w_{11} & w_{12} \\
w_{21} &w_{22} \\
\vdots &\vdots \\
w_{n1} &w_{n2}
\end{bmatrix}
$$
The output for this small network is found by treating the hidden layer as inputs for the output unit. The network output is expressed simply
$$
y = f_2 \! \left(\, f_1 \! \left(\vec{x} \, \mathbf{W_1}\right) \mathbf{W_2} \right)
$$
```
### Generate some data
torch.manual_seed(7) # Set the random seed so things are predictable
# Features are 3 random normal variables
features = torch.randn((1, 3))
# Define the size of each layer in our network
n_input = features.shape[1] # Number of input units, must match number of input features
n_hidden = 2 # Number of hidden units
n_output = 1 # Number of output units
# Weights for inputs to hidden layer
W1 = torch.randn(n_input, n_hidden)
# Weights for hidden layer to output layer
W2 = torch.randn(n_hidden, n_output)
# and bias terms for hidden and output layers
B1 = torch.randn((1, n_hidden))
B2 = torch.randn((1, n_output))
```
> **Exercise:** Calculate the output for this multi-layer network using the weights `W1` & `W2`, and the biases, `B1` & `B2`.
```
## Your solution here
```
If you did this correctly, you should see the output `tensor([[ 0.3171]])`.
The number of hidden units a parameter of the network, often called a **hyperparameter** to differentiate it from the weights and biases parameters. As you'll see later when we discuss training a neural network, the more hidden units a network has, and the more layers, the better able it is to learn from data and make accurate predictions.
## Numpy to Torch and back
Special bonus section! PyTorch has a great feature for converting between Numpy arrays and Torch tensors. To create a tensor from a Numpy array, use `torch.from_numpy()`. To convert a tensor to a Numpy array, use the `.numpy()` method.
```
import numpy as np
a = np.random.rand(4,3)
a
b = torch.from_numpy(a)
b
b.numpy()
```
The memory is shared between the Numpy array and Torch tensor, so if you change the values in-place of one object, the other will change as well.
```
# Multiply PyTorch Tensor by 2, in place
b.mul_(2)
# Numpy array matches new values from Tensor
a
```
| github_jupyter |
# Automated Air-Liquid Interface Cell Culture Analysis Using Deep Optical Flow
## Autogenerated Report: RAINBOW image series analysis results
#### Author: Alphons G
#### Website: https://github.com/AlphonsGwatimba/Automated-Air-Liquid-Interface-Cell-Culture-Analysis-Using-Deep-Optical-Flow
```
# import required packages
import json
import os
import cv2
from IPython.display import Image, Video
import pandas as pd
```
## Source directory
```
print(os.getcwd())
```
## Image Series Type
```
# Load metadata
with open('metadata.json') as f:
metadata = json.load(f)
print(metadata['type'])
```
## RAINBOW Analysis Timestamp
```
print(metadata['analysis_timestamp'])
```
## Image Series Metadata
```
print(json.dumps(metadata, sort_keys=True, indent=4))
```
## Magnitude Statistics
```
df = pd.read_csv('mag_stats.csv', sep=',')
print(df)
```
## Direction Statistics
```
df = pd.read_csv('dirn_stats.csv', sep=',')
print(df)
```
## Raw Image Series (left) and RAINBOW Optical Flow Visualisation (Right)
```
# video resizing function
def video_reshp(vid_path, set_wdh=None):
cap = cv2.VideoCapture(vid_path)
hgt, wdh, _ = cap.read()[1].shape
dsp_wdh = set_wdh if set_wdh is not None else wdh
dsp_hgt = dsp_wdh * (hgt / wdh) if wdh is not None else hgt
return dsp_wdh, dsp_hgt
vid_path = 'Combined Images/Combined.mp4v'
dsp_wdh, dsp_hgt = video_reshp(vid_path, 1280)
Video(vid_path, embed=True, width=dsp_wdh, height=dsp_hgt, html_attributes='controls loop')
```
a) The direction of motion at any position within RAINBOW generated optical flow images is measured clockwise from the initial horizontal position of a unit circle (left) and is shown using hue values (right).
b) The magnitude of motion at any position within RAINBOW generated optical flow images is shown using saturation values. For instance, high saturation corresponds to high magnitude of motion and low saturation corresponds to low magnitude of motion. For instance, high saturation (100%) corresponds to high magnitude of motion and low saturation (25%) corresponds to low magnitude of motion.

## Magnitude Heatmaps Across Image Series
```
vid_path = 'Heatmaps/heatmap.mp4v'
dsp_wdh, dsp_hgt = video_reshp(vid_path, 640)
Video(vid_path, embed=True, width=dsp_wdh, height=dsp_hgt, html_attributes='controls loop')
```
Visualization of motion magnitiude using heatmap with hot colour mapping.
## Quiver Plots Across Image Series
```
vid_path = 'Quiver Plots/quiver_plot.mp4v'
dsp_wdh, dsp_hgt = video_reshp(vid_path, 640)
Video(vid_path, embed=True, width=dsp_wdh, height=dsp_hgt, html_attributes='controls loop')
```
Visualisation of optical flow using quiver plots containing vector arrows.
| github_jupyter |
```
from skimage import io
import numpy as np
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from skimage.transform import pyramid_gaussian
from torch.autograd import Variable
from math import exp
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
from google.colab import drive
drive.mount('/content/drive')
I = io.imread("/content/drive/My Drive/MM803/Assignment2/knee1.bmp").astype(np.float32)/255.0 # fixed image
J = io.imread("/content/drive/My Drive/MM803/Assignment2/knee2.bmp").astype(np.float32)/255.0 # moving image
%matplotlib inline
fig=plt.figure()
fig.add_subplot(1,2,1)
plt.imshow(I,cmap="gray")
plt.title("Fixed Image")
fig.add_subplot(1,2,2)
plt.imshow(J,cmap="gray")
plt.title("Moving Image")
L = 6 # Gaussian pyramid level
downscale = 2.0 # downscale factor for the gaussian pyramid
pyramid_I = tuple(pyramid_gaussian(I, downscale=downscale, multichannel=False))
pyramid_J = tuple(pyramid_gaussian(J, downscale=downscale, multichannel=False))
# create a list of necessary objects you will need and commit to GPU
I_lst,J_lst,h_lst,w_lst,xy_lst,ind_lst=[],[],[],[],[],[]
for s in range(L):
I_, J_ = torch.tensor(pyramid_I[s].astype(np.float32)).to(device), torch.tensor(pyramid_J[s].astype(np.float32)).to(device)
I_lst.append(I_)
J_lst.append(J_)
h_, w_ = I_lst[s].shape[0], I_lst[s].shape[1]
ind_ = torch.randperm(h_*w_).to(device)
ind_lst.append(ind_)
print(h_,w_,len(ind_))
h_lst.append(h_)
w_lst.append(w_)
y_, x_ = torch.meshgrid([torch.arange(0,h_).float().to(device), torch.arange(0,w_).float().to(device)])
y_, x_ = 2.0*y_/(h_-1) - 1.0, 2.0*x_/(w_-1) - 1.0
xy_ = torch.stack([x_,y_],2)
xy_lst.append(xy_)
class HomographyNet(nn.Module):
def __init__(self):
super(HomographyNet, self).__init__()
# perspective transform basis matrices
self.B = torch.zeros(8,3,3).to(device)
self.B[0,0,2] = 1.0
self.B[1,1,2] = 1.0
self.B[2,0,1] = 1.0
self.B[3,1,0] = 1.0
self.B[4,0,0], self.B[4,1,1] = 1.0, -1.0
self.B[5,1,1], self.B[5,2,2] = -1.0, 1.0
self.B[6,2,0] = 1.0
self.B[7,2,1] = 1.0
self.v = torch.nn.Parameter(torch.zeros(8,1,1).to(device), requires_grad=True)
def forward(self):
return MatrixExp(self.B,self.v)
def MatrixExp(B,v):
C = torch.sum(B*v,0)
A = torch.eye(3).to(device)
H = torch.eye(3).to(device)
for i in torch.arange(1,10):
A = torch.mm(A/i,C)
H = H + A
return H
def PerspectiveWarping(I, H, xv, yv):
# apply transformation in the homogeneous coordinates
xvt = (xv*H[0,0]+yv*H[0,1]+H[0,2])/(xv*H[2,0]+yv*H[2,1]+H[2,2])
yvt = (xv*H[1,0]+yv*H[1,1]+H[1,2])/(xv*H[2,0]+yv*H[2,1]+H[2,2])
J = F.grid_sample(I,torch.stack([xvt,yvt],2).unsqueeze(0),align_corners=False).squeeze()
return J
def ncc_val(I,J):
I_mean = torch.mean(I)
J_mean = torch.mean(J)
I_std = torch.std(I)
J_std = torch.std(J)
return torch.mean((I-I_mean)*(J-J_mean)/(I_std*J_std))
def multi_resolution_NCC_loss():
loss=0.0
for s in np.arange(L-1,-1,-1):
Jw_ = PerspectiveWarping(J_lst[s].unsqueeze(0).unsqueeze(0), homography_net(), xy_lst[s][:,:,0], xy_lst[s][:,:,1]).squeeze()
ncc_value = ncc_val(I_lst[s], Jw_)
loss = loss - (1. / L) * ncc_value
#print(histogram_mutual_information(I_lst[s].cpu().detach().numpy(), Jw_.cpu().detach().numpy()))
return loss, ncc_value
homography_net = HomographyNet().to(device)
optimizer = optim.Adam([{'params': homography_net.v, 'lr': 1e-4}], amsgrad=True)
for itr in range(2200):
optimizer.zero_grad()
loss,ncc_value = multi_resolution_NCC_loss()
if itr%200 == 0:
print("Itr:",itr,"NCC value:","{:.4f}".format(ncc_value))
print("NCC loss:", "{:.4f}".format(loss))
loss.backward()
optimizer.step()
print("Itr:",itr+1,"NCC value:","{:.4f}".format(ncc_value))
def histogram_mutual_information(image1, image2):
hgram, x_edges, y_edges = np.histogram2d(image1.ravel(), image2.ravel(), bins=100)
pxy = hgram / float(np.sum(hgram))
px = np.sum(pxy, axis=1)
py = np.sum(pxy, axis=0)
px_py = px[:, None] * py[None, :]
nzs = pxy > 0
return np.sum(pxy[nzs] * np.log(pxy[nzs] / px_py[nzs]))
I_t = torch.tensor(I).to(device)
J_t = torch.tensor(J).to(device)
H = homography_net()
J_w = PerspectiveWarping(J_t.unsqueeze(0).unsqueeze(0), H, xy_lst[0][:, :, 0], xy_lst[0][:, :, 1]).squeeze()
D = J_t - I_t
D_w = J_w - I_t
print("Mutual information before registration:", "{:.4f}".format(histogram_mutual_information(I, J_t.cpu().detach().numpy())))
print("Mutual information after registration:", "{:.4f}".format(histogram_mutual_information(I, J_w.cpu().detach().numpy())))
print("Transformation matrix:")
print(H.cpu().detach().numpy())
print("")
Ra = I_t.clone()
Rb = I_t.clone()
b = 50
for i in torch.arange(0, I_t.shape[0] / b, 1).int():
for j in torch.arange(i % 2, np.floor(I_t.shape[1] / b), 2).int():
Rb[i * b:(i + 1) * b, j * b:(j + 1) * b] = J_t[i * b:(i + 1) * b, j * b:(j + 1) * b].clone()
Ra[i * b:(i + 1) * b, j * b:(j + 1) * b] = J_w[i * b:(i + 1) * b, j * b:(j + 1) * b].clone()
fig = plt.figure(figsize=(10, 10))
fig.add_subplot(1, 2, 1)
plt.imshow(Rb.cpu().data, cmap="gray")
plt.title("Images before registration")
fig.add_subplot(1, 2, 2)
plt.imshow(Ra.cpu().data, cmap="gray")
plt.title("Images after registration")
plt.show()
```
| github_jupyter |
# Conservation Analysis and Epitope Prediction
#### Author: C. Mazzaferro, K. Fisch
#### Email: cmazzafe@ucsd.edu
#### Date: October 2016
## Outline of Notebook
<a id = "toc"></a>
1. <a href = "#background">Background</a>
2. <a href = "#Cons">High Affinity Binding Prediction </a>
* <a href = "#Agg">Data Aggregation</a>
* <a href = "#Sim">Similarty Score Calculation</a>
* <a href = "#Plot">Visualize</a>
Directions (from Mali's e-mail sent a couple of weeks ago):
Determine Cas9 orthologs that are orthogonal in the 'immunogenicity space'. This will allow us to prescribe a sequential regimen of Cas9s for therapeutic interventions. I've been thinking some more about how to do this, and I believe a simple strategy as follows might work:
- step i: build a database of Cas9 orthologs; #n.
- step ii: pick a seed Cas9 ortholog, say Sp, and then do a pairwise alignment with every other ortholog (n-1 alignments total). note that aligning all together will not be useful for our objective.
- step iii: overlay the immunogenicity profiles and score direct peptide overlaps (adding +1 for each overlap and 0 otherwise). sum the scores and retain orthologs (#m) for which the summation score = 0.
- step iv: randomly pick a second Cas9 from the orthologs retained in step iii and iterate back to step ii but now do just m-1 alignments with the retained orthologs.
This should quickly converge and give a handful of orthologs. We should ideally be able to dial the immunogenicity thresholds in the workflow as we could potentially land up with too many or too few options. We might also get different orthogonal lists based on the choice of seed ortholog -- but their should still be substantial overlaps, and so we should ideally iterate through all orthologs in step i to establish robustness of the lists.
This is just a suggestion for a potential workflow, so let me know what you'll think and absolutely feel free to pick this apart!
```
- Phylogenetic tree
- craig venture; provean software
```
<a id = "background"></a>
# Background
CRISPR (clustered regularly interspaced short palindromic repeat) is an adaptive immune system that provides protection against mobile genetic elements (viruses, transposable elements and conjugative plasmids).
CRISPR clusters contain spacers, sequences complementary to antecedent mobile elements, and target invading nucleic acids. CRISPR clusters are transcribed and processed into CRISPR RNA (crRNA). In type II CRISPR systems correct processing of pre-crRNA requires a trans-encoded small RNA (tracrRNA), endogenous ribonuclease 3 (rnc) and this protein. The tracrRNA serves as a guide for ribonuclease 3-aided processing of pre-crRNA; Cas9 only stabilizes the pre-crRNA:tracrRNA interaction and has no catalytic function in RNA processing. Subsequently Cas9/crRNA/tracrRNA endonucleolytically cleaves linear or circular dsDNA target complementary to the spacer; Cas9 is inactive in the absence of the 2 guide RNAs (gRNA). The target strand not complementary to crRNA is first cut endonucleolytically, then trimmed 3'-5' exonucleolytically. DNA-binding requires protein and both gRNAs, as does nuclease activity. Cas9 recognizes the protospacer adjacent motif (PAM) in the CRISPR repeat sequences to help distinguish self versus nonself, as targets within the bacterial CRISPR locus do not have PAMs. DNA strand separation and heteroduplex formation starts at PAM sites; PAM recognition is required for catalytic activity (PubMed:24476820). Confers immunity against a plasmid with homology to the appropriate CRISPR spacer sequences (CRISPR interference) (PubMed:21455174)
```
from IPython.display import Image
Image("/Users/carlomazzaferro/Desktop/BINF Int Rand Pics/workflow_epitope.png")
```
The pipeline presented here is aimed at identify the most immunogenic peptides in Cas9. By looking at multiple sequence alignment scores for the possible peptides and their associated predicted MHC affinity, we can determine which peptides within a Cas9 orthologue can be swapped in order to reduce overall immunogenecity.
The steps are constructed as follows:
1. Select proteins [from reference papaer](http://nar.oxfordjournals.org/content/suppl/2013/10/29/gkt1074.DC1/nar-02672-z-2013-File008.pdf)
2. Predict the MHC affinity of each peptide using [CBS's prediction services](http://www.cbs.dtu.dk/services/), in particular [netMHCcons Server](http://www.cbs.dtu.dk/services/NetMHCcons/)
3. Rank proteins according to their similarity bases on a seed*
*See attached file: `workflow_and_results.txt` for a lengthier description of how the scoring was done
<a id = "Cons"></a>
# MHC Binding Affinity Prediction
### Run on netMHCcons for n-mers of 8, 9, 10, 11 amino acids and for the 12 allele supertype families
Run for each protein sequence retrieved from reference paper.
```
import pandas
idd = pandas.read_csv('/Users/carlomazzaferro/Desktop/single_swap_orig_proteins.csv').drop(['Matches Loc'], 1)
idd.to_csv('/Users/carlomazzaferro/Desktop/single_swap_orig_protein.csv')
```
## Load file
```
from nepitope import mhc_utils
import pandas
import glob
import importlib
importlib.reload(mhc_utils)
#files saved from netMHC in multiple different files, one per each query (nmer, allele)
fasta_file = '/Users/carlomazzaferro/Desktop/Test_Workflow_From_Scratch/fasta_from_study_dealigned.fasta'
filepath = '/Users/carlomazzaferro/Desktop/Test_Workflow_From_Scratch/mhc_preds_fasta_from_study_dealigned/'
aggregate_all = mhc_utils.FileConsolidation(filepath, fasta_file)
```
<a id = "Agg"></a>
### Data Aggregation
Join data: return data in a list of dataframes, where each dataframe contains data about a protein (May take some time)
```
#returns a dataframe per file used (just one in this case)
df_list = aggregate_all.return_df_list()
#Would run this. But too slow. Th e 3 cells below are speed-optimized.
lsss_1 = aggregate_all.optimized_list_df_by_prot(df_list)
sss = pandas.concat(lsss_1)
high_int_aa = sss.loc[sss['Affinity Level'] == ('High' or 'Intermediate')]
high_int_aa.to_csv('/Users/carlomazzaferro/Desktop/Test_Workflow_From_Scratch/all_high_aff_aas.csv')
```
## Load original proteins
```
from nepitope import pep_utils
fasta_all_prots = '/Users/carlomazzaferro/Desktop/Test_Workflow_From_Scratch/fasta_from_study_dealigned.fasta'
from nepitope import pairwise_comp
import importlib
importlib.reload(pairwise_comp)
```
### Pipeline subprocess call
```
pwcomp = pairwise_comp.PairwiseComp(lsss_1, 5, fasta_all_prots)
df_pairwise_comps = pwcomp.pipe_run()
basepath = '/Users/carlomazzaferro/Desktop/Test_IEDB/newer_pipeline_files/consistent_aws_netMHC40/High AA Per Protein/'
df_pairwise_comps.drop(['Matches Loc'], 1).to_csv('/Users/carlomazzaferro/Desktop/Test_Workflow_From_Scratch/df_pairwise_comps_no_swap.csv')
importlib.reload(merge_workflows)
from nepitope import merge_workflows
swaps_df_file = '/Users/carlomazzaferro/Desktop/Test_IEDB/newer_pipeline_files/consistent_aws_netMHC40/summary_results_per_prediction.csv'
swaps_df = pandas.read_csv(swaps_df_file)
```
## Merge workflows
```
############
merged = merge_workflows.MergeSwapsAndComp(df_pairwise_comps, swaps_df, 1)
df_top = merged.top_swap_df
df_ordered_filtered = merged.final_sort_and_filtering(df_top)
importlib.reload(modify_data)
from nepitope import modify_data
mod_fasta = '/Users/carlomazzaferro/Desktop/Test_Workflow_From_Scratch/mod_fasta/'
orig_fasta = '/Users/carlomazzaferro/Desktop/Test_Workflow_From_Scratch/fasta_from_study_dealigned.fasta'
mod_data = modify_data.ModifyData(df_ordered_filtered, fasta_file, mod_fasta, lsss_1)
mod_data.get_modified_fastas_single_swap()
exchange_pairs = []
for i,j in mod_data.swaps_dic.items():
exchange_pairs.append([i,j])
pair = exchange_pairs[0]
pair[0] = pair[0][0:-1] + 'D'
pair[0] = 'ISTFRIPYYV'
pair[1] = 'ISTFRIPYYV'
pair
my_fasta = '/Users/carlomazzaferro/Desktop/Test_Workflow_From_Scratch/mod_fasta/ISTFRIPYYV-ILTFRIPYYD.fasta'
#ls_mod = mod_data.get_singly_modified_df_list(pair[0], pair[1])
pwcomp_pair_swap = pairwise_comp.PairwiseComp(ls_mod, 5, my_fasta)
df_swap = pwcomp_pair_swap.pipe_run()
df_swap = df_swap.drop('Matches Loc', 1)
df_swap.to_csv(csv_path + pair[0] + '-' + pair[1] + '.csv')
list_dfs.append(df_swap)
list_dfs[-1].loc['StreptococcusPyogenes_reference']
list_dfs[1]
idk = lsss_1[-1][lsss_1[-1]['Peptide'] == 'MIKFRGHF']
idk['Affinity Level'] = 'No'
import re
mystring = list(merged.top_swap_list['top scoring peptides'][0:3].values)
list(filter(None, re.sub('[^A-Za-z0-9]+', ',', mystring).split(',')))[-3]
lsss_1[-1][lsss_1[-1]['Peptide'] == 'MIKFRGHF']
ix = df[df['original peptide'].isin(orig_peps)].index
df.set_value(ix, 'issubset', 'No')
df = merged.top_swap_list
_get_swap_peps(df, 5)
n_orig_peptide = 5
peps_to_swap = _get_swap_peps(df, n_orig_peptide)
orig_peps = _get_orig_peps(df, n_orig_peps)
orig_and_swaps = dict(zip(orig_peps, peps_to_swap))
orig_and_swaps
mod_df = modify_df(lsss_1, 1, df)
md = mod_df[-1]
md['Peptide'] = md['Peptide'].apply(lambda x: str(x))
md.Peptide = md.Peptide.replace({'3': 'abc'})
md
#ix = md.loc[md.Peptide.isin(peps_to_swap)].index
#md.set_value(ix, 'Affinity Level', 'No')
#md
### reference = 'StreptococcusPyogenes_reference'
n_orig_peps = 5
reference = 'StreptococcusPyogenes_reference'
from nepitope import pep_utils
def _get_swap_peps(df, n_swaps):
swaps = []
list_swaps = list(df['top scoring peptides'][0:n_swaps].values)
for i in list_swaps:
swap = list(filter(None, re.sub('[^A-Za-z0-9]+', ',', i).split(',')))[-3]
swaps.append(swap)
return swaps
def _get_orig_peps(df, n_swaps):
return df['original peptide'].values.tolist()[0:n_swaps]
def modify_df(list_df, n_orig_peptide, df):
peps_to_swap = _get_swap_peps(df, n_orig_peptide)
orig_peps = _get_orig_peps(df, n_orig_peps)
orig_and_swaps = dict(zip(orig_peps, peps_to_swap))
for idx, df in enumerate(list_df):
if df.ID.unique()[0] == reference:
new_df = df
new_df.Peptide = new_df.replace({'Peptide' : {'DIKFRGHF': 'MIKFRGHF'}})
ix = new_df.loc[new_df.Peptide.isin(peps_to_swap)].index
new_df.set_value(ix, 'Affinity Level', 'No')
list_df[idx] = new_df
return list_df
fasta = '/Users/carlomazzaferro/Desktop/Test_IEDB/newer_pipeline_files/fasta_from_study_dealigned.fasta'
fasta_out_dir = '/Users/carlomazzaferro/Desktop/Test_IEDB/newer_pipeline_files/swapped_fastas/'
def modify_fasta(fasta, fasta_our_dir, orig_peptide, swap_pep, reference, df):
n_orig_peptide = 5
idx, seq = pep_utils.create_separate_lists(fasta)
zipped = list(zip(idx, seq))
zipped = [list(x) for x in zipped]
peps_to_swap = _get_swap_peps(df, n_orig_peptide)
print(peps_to_swap)
orig_peps = _get_orig_peps(df, n_orig_peps)
print(orig_peps)
orig_and_swaps = dict(zip(orig_peps, peps_to_swap))
swapped_fasta = []
for i in zipped:
if i[0][1::] == reference:
i[1] = replace_all(i[1], orig_and_swaps)
swapped_fasta.append(i)
with open(fasta_out_dir + 'n_swaps' + '.fasta', 'w') as out:
for i in swapped_fasta:
out.write(i[0] + '\n')
out.write(i[1] + '\n')
def replace_all(text, dic):
for i, j in dic.items():
text = text.replace(i, j)
return text
modify_fasta(fasta, fasta_out_dir, orig_pep, swap_pep, reference, df)
ls = modify_df(lsss_1, orig_pep, swap_pep)
mod_fasta = '/Users/carlomazzaferro/Desktop/Test_IEDB/newer_pipeline_files/swapped_fastas/MIKFPGHF.fasta'
pwcomp_1_swap = pairwise_comp.PairwiseComp(ls, 5, mod_fasta)
df_pairwise_comps_1 = pwcomp_1_swap.pipe_run()
df_pairwise_comps_1.loc[reference]
df_pairwise_comps.loc[reference]
df_pairwise_comps_1.to_csv('/Users/carlomazzaferro/Desktop/df_single_swap.csv')
```
| github_jupyter |
# Setup
```
%%capture
%pip install poetry
%pip install git+https://github.com/oughtinc/ergo.git@f5646b672eb0d60c58e7de850eea5f43a4feaacc
%pip install xlrd
%load_ext google.colab.data_table
%%capture
import ergo
import numpy as np
import pandas as pd
import ssl
import warnings
import requests
from datetime import timedelta, date
from ergo.contrib.el_paso import texas_data, onlyasith, krismoore, brachbach, shaman
from ergo.contrib.utils import plot_question, question, rejection_sample, sample_from_ensemble, samplers, summarize_question_samples, daterange
warnings.filterwarnings(action="ignore", category=FutureWarning)
warnings.filterwarnings(module="plotnine", action="ignore")
warnings.filterwarnings(module="jax", action="ignore")
ssl._create_default_https_context = ssl._create_unverified_context
metaculus = ergo.Metaculus(
username="oughtpublic",
password="123456",
api_domain = "pandemic"
)
```
# Retrieve external data and models
## Texas government cases data
```
el_paso_cases = texas_data.get_el_paso_data()
el_paso_cases
```
## @KrisMoore's compiled data
Pulled from
[here](https://docs.google.com/spreadsheets/d/1eGF9xYmDmvAkr-dCmd-N4efHzPyYEfVl0YmL9zBvH9Q/edit#gid=1694267458)
```
compiled_data = krismoore.get_krismoore_data()
krismoore.graph_compiled_data(compiled_data)
```
## @onlyasith's cases model
Pulled from
[here](https://docs.google.com/spreadsheets/d/1L6pzFAEJ6MfnUwt-ea6tetKyvdi0YubnK_70SGm436c/edit#gid=1807978187)
```
projected_cases = onlyasith.get_onlyasith_results()
projected_cases
```
## Shaman et al. cases model
Pulled from [here](https://github.com/shaman-lab/COVID-19Projection)
([paper](https://www.medrxiv.org/content/10.1101/2020.03.21.20040303v2))
```
cu_projections = shaman.load_cu_projections("El Paso County TX")
```
## @brachbach model: cases -> hospitalized
```
get_daily_hospital_confirmed = brachbach.get_daily_hospital_confirmed
```
## Access historical data
```
def get_historical_data(date: date, column_name):
"""
Look up the value of a parameter on a given date
in the historical data we've loaded.
Return the value or raise a KeyError if we don't have it.
"""
# prefer Texas government data over @KrisMoore compiled data
try:
value = el_paso_cases.loc[date, column_name]
if np.isnan(value):
raise KeyError(f"value for {column_name} in el_paso_cases is NaN")
except KeyError:
value = compiled_data.loc[date, column_name]
if np.isnan(value):
raise KeyError(f"value for {column_name} in compiled_data is NaN")
return value
```
# Model components
In this notebook, we model some aspects of how the COVID-19 pandemic
will play out in El Paso. Our goal is to answer the questions in the
[El
Paso Metaculus series](https://pandemic.metaculus.com/questions/?search=cat:internal--el-paso).
In this section, we model some key variables that we'll use to answer
the Metaculus questions in the next section. We sometimes ensemble
multiple models to try to get a better estimate of a variable.
```
START_DATE = date(2020, 4, 1)
```
## Daily COVID Infections
### Shaman Model
```
@ergo.mem
def cu_model_scenario():
"""Which of the model scenarios are we in?"""
return ergo.random_choice([s for s in cu_projections.keys()])
@ergo.mem
def cu_model_quantile():
"""Where in the distribution of model outputs are we for this model run?
Want to be consistent across time, so we sample it once per model run"""
return ergo.uniform()
def cu_projection(param: str, date: date) -> int:
"""
Get the Columbia model's prediction
of the param for the date
"""
scenario = cu_model_scenario()
quantile = cu_model_quantile()
# Extract quantiles of the model distribution
xs = np.array([0.025, 0.25, 0.5, 0.75, 0.975])
scenario_df = cu_projections[scenario]
param_df = scenario_df[scenario_df["var"] == param]
date_df = param_df[param_df["Date"] == date]
if date_df.empty:
raise KeyError(f"No Columbia project for param: {param}, date: {date}")
ys = np.array(date_df[["2.5", "25", "50", "75", "97.5"]].iloc[0])
# Linearly interpolate
return int(round(np.interp(quantile, xs, ys)))
```
### @onlyasith model+
```
@ergo.mem
def daily_infections_onlyasith(date: date) -> int:
"""
What is the number of reported (new) Covid-19 infections on [date]?
"""
try:
# Look up projections from @onlyasith's model
cases = projected_cases.loc[date, "New cases"]
if np.isnan(cases):
raise KeyError
# Add some (fairly arbitrary) uncertainty around this point estimate
if cases == 0:
return cases
cases_estimate = ergo.lognormal_from_interval(cases * 0.8, cases * 1.2)
return int(np.clip(cases_estimate, cases * 0.5, cases * 2).round())
except KeyError:
# We're beyond the time range for data and model
return 0
```
### Get historical data or predict using ensemble
```
@ergo.mem
def daily_infections(date: date) -> int:
"""
What is the number of reported (new) Covid-19 infections on [date]?
"""
try:
return get_historical_data(date, "New cases")
except KeyError: # if we don't have historical data, use our ensemble of models
return sample_from_ensemble([
lambda date: cu_projection("cases", date),
daily_infections_onlyasith
], {"date": date}, [.8, .2], fallback=True, default=0)
```
## Daily COVID infections: mean, sma, peak
```
@ergo.mem
def mean_infections(start_date: date, end_date: date):
"""
What is the average number of reported new infections for this range of
dates? (Including start date, excluding end date)
"""
days = daterange(start_date, end_date)
return np.mean([daily_infections(day) for day in days])
@ergo.mem
def sma_infections(date: date):
"""
The simple moving average of infections for a date.
Defined in https://pandemic.metaculus.com/questions/4128:
'The 2-day SMA is defined as the unweighted average (arithmetic mean)
over the current day and the previous day.'
"""
return mean_infections(date - timedelta(1), date + timedelta(1))
@ergo.mem
def peak_compatible_with_historical_data(peak_date):
"""
Check whether it's possible that some date in the past
was the peak date of new COVID infections in El Paso,
given the historical data we have about COVID infections
"""
if not peak_date in el_paso_cases.index:
return True
for comparison_date in daterange(START_DATE, peak_date + timedelta(11)):
if comparison_date not in el_paso_cases.index:
continue
if sma_infections(comparison_date) > sma_infections(peak_date):
return False
if sma_infections(comparison_date) == sma_infections(peak_date) and comparison_date > peak_date:
return False
return True
@ergo.mem
def peak_infection_date_community():
"""
The community assigns probability to some dates in the past
that we already know were not the peak.
So instead of sampling from the full community distribution,
sample from the portion of the community distribution
that is plausibly correct.
"""
peak_date = rejection_sample(
peak_infection_date.question.sample_community,
peak_compatible_with_historical_data)
return peak_date
```
## Patients with COVID in the hospital
### @brachbach model+
```
br_confirmed_from_infected_model = get_daily_hospital_confirmed(compiled_data, daily_infections)
def brachbach_model_plus(date: date) -> int:
"""
Get a point estimate from @brachbach's model,
then add some (fairly arbitrary) uncertainty
"""
cases = br_confirmed_from_infected_model(date)
if cases == 0:
return cases
cases_estimate = ergo.lognormal_from_interval(cases * 0.8, cases * 1.2)
return np.clip(cases_estimate, cases * 0.5, cases * 2)
```
### Get historical data or predict using ensemble
```
@ergo.mem
def hospital_confirmed_for_date(date: date) -> int:
"""
The total number of lab-confirmed COVID-19 patients in El Paso County in
the hospital on this date
"""
try:
return get_historical_data(date, "In hospital confirmed")
except KeyError: # if we don't have historical data, use our ensemble of models
return sample_from_ensemble([
lambda date: cu_projection("hosp", date),
brachbach_model_plus
], {"date": date}, [.2, .8], fallback=True, default=0)
```
## Proportion ICU admissions requiring ventilation
```
@ergo.mem
def frac_icu_ventilation():
"""
Proportion of ICU admissions requiring ventilation
Approach (PabloStafforini et al):
https://pandemic.metaculus.com/questions/4154/#comment-28155
TODO:
- Improve how we use case data
- Add qualitative adjustments
"""
ventilation_pseudocounts = 25 + 17 + 0.05 * 1150 + 0.1 * 132
icu_pseudocounts = 100 + 36 + 0.05 * 1300 + 0.1 * 196
return ergo.beta_from_hits(ventilation_pseudocounts, icu_pseudocounts)
```
# El Paso questions
```
@question(metaculus, 4128, community_weight=0.3, community_fn=peak_infection_date_community, start_date=START_DATE)
def peak_infection_date() -> date:
"""
When will El Paso County, Texas, experience its first peak number of COVID
infections?
From https://pandemic.metaculus.com/questions/4128:
'This question resolves as the date for which
the 2-day simple moving average(SMA) of the number of reported new infections
is strictly greater than the 2-day SMA over the subsequent 10 days.'
"""
end_date = date(2020, 7, 1)
for today in daterange(START_DATE, end_date):
sma_today = sma_infections(today)
future_smas = [sma_infections(today + timedelta(i)) for i in range(1,11)]
if sma_today > max(future_smas):
return today
return end_date
plot_question(peak_infection_date, start_date=START_DATE)
@question(metaculus, 4137, community_weight=0.5)
def peak_infections():
"""
How many new infections will be reported in El Paso on the day on which
the number of new reported infections peaks?
"""
peak = peak_infection_date()
return daily_infections(peak)
plot_question(peak_infections)
@question(metaculus, 4152, community_weight=0.5)
def mean_infections_peak345():
"""
What will the average number of reported daily infections be in El Paso,
over the 3rd, 4th and 5th days after the first "peak"?
"""
peak = peak_infection_date()
return mean_infections(peak + timedelta(3), peak + timedelta(6))
plot_question(mean_infections_peak345)
@question(metaculus, 4170, community_weight=0.8)
def mean_infections_peak678():
"""
What will the average number of reported daily infections be in El Paso,
over the 6th, 7th and 8th days after the first "peak"?
"""
peak = peak_infection_date()
return mean_infections(peak + timedelta(6), peak + timedelta(9))
plot_question(mean_infections_peak678)
@question(metaculus, 4155, community_weight=0.7)
def frac_patients_icu():
"""
What portion of in-hospital cases in El Paso County will require admission
to the ICU?
Following @katifish's approach:
https://pandemic.metaculus.com/questions/4155/#comment-28054
TODO: Add others from katifish comment
"""
alpha = 0.1 # Rescaling counts becase we're more uncertain than implied by counts
return ergo.random_choice([
ergo.beta_from_hits(alpha * 121, alpha * 508),
ergo.beta_from_hits(alpha * 181, alpha * 507),
])
plot_question(frac_patients_icu)
@question(metaculus, 4154, community_weight=0.3)
def frac_patients_invasive():
"""
What portion of in-hospital patients with Covid-19 in El Paso County will
require invasive ventilation?
Following @PabloStafforini's indirect estimation approach:
https://pandemic.metaculus.com/questions/4154/#comment-28155
TODO:
- Combine with direct estimate
direct_estimate = ergo.beta_from_hits(0.1 * 130, 0.1 * 393)
"""
return frac_patients_icu() * frac_icu_ventilation()
plot_question(frac_patients_invasive)
@ergo.mem
def peak_hospitalized_date():
"""
What will be the date when there are the max number of COVID patients in the hospital
within 15 days before or after the date of the first peak in confirmed cases?
"""
infection_peak_date = peak_infection_date()
days = list(daterange(infection_peak_date - timedelta(15), infection_peak_date + timedelta(16)))
hospitalization_peak_date = days[0]
hospitalized_peak = 0
for day in days:
hospitalized_for_day = hospital_confirmed_for_date(day)
# if there are 2 different dates
# with the same peak number of hospitalized patients,
# return the first date:
# https://pandemic.metaculus.com/questions/4204#comment-30023
if hospitalized_for_day > hospitalized_peak:
hospitalization_peak_date = day
hospitalized_peak = hospitalized_for_day
return hospitalization_peak_date
@question(metaculus, 4153, community_weight=0.3)
def max_30d_hospital_confirmed_for_peak():
"""
What will the maximum number of in-hospital lab-confirmed COVID-19
patients in El Paso County, in the 30-day period during which the "peak"
occurs?
"""
return hospital_confirmed_for_date(peak_hospitalized_date())
plot_question(max_30d_hospital_confirmed_for_peak, bw=0.01)
@question(metaculus, 4204)
def peak_icu_patients():
"""
How many patients with Covid-19 in El Paso County will be in the
ICU on the day when the number of hospital admissions of cases peak?
"""
peak_date = peak_hospitalized_date()
try:
return get_historical_data(peak_date, "in_icu")
# if we don't have historical data, sample from the Columbia model or from the community
except KeyError:
return sample_from_ensemble([
lambda date: cu_projection("ICU", date),
lambda date: peak_icu_patients.question.sample_community()
], {"date": peak_date}, [.65, .35], fallback=True)
plot_question(peak_icu_patients, bw=0.1)
@question(metaculus, 4201)
def peak_invasive_ventilation():
"""
How many patients with Covid-19 in El Paso County will be on invasive
ventilation on the day when the number of hospital admissions of cases
peak?
"""
peak_date = peak_hospitalized_date()
try:
return get_historical_data(peak_hospitalized_date(), "on_ventilator")
# if we don't have historical data, sample from the Columbia model or from the community
except KeyError:
return sample_from_ensemble([
lambda date: cu_projection("ICU", date),
lambda date: peak_icu_patients.question.sample_community()
], {"date": peak_date}, [.65, .35], fallback=True)
plot_question(peak_invasive_ventilation, bw=0.1)
```
# Generate predictions for all questions
```
def model():
for sampler in samplers.values():
sampler()
samples = ergo.run(model, num_samples=2000)
summarize_question_samples(samples)
```
# Compare predictions to community
This takes a while since we're fitting a mixture of logistic
distributions to our samples before visualizing (and submitting) them.
These may look a little different from the plots for the questions shown
above, because:
1. we've taken more samples from the distribution
2. rather than showing raw samples, we're fitting logistic distributions that we can submit them to metaculus
```
submissions = {}
for sampler in samplers.values():
q = sampler.question
q_samples = samples[sampler.__name__]
if q.id == 4128: # Date question: Need to convert back to date from days (https://github.com/oughtinc/ergo/issues/144)
q_samples = np.array([START_DATE + timedelta(s) for s in q_samples])
if q.id in [4201, 4204, 4137, 4152, 4170, 4153]:
# Clip extreme values for questions that we had issues fitting
(sample_min, sample_max) = np.quantile(q_samples, [0.08, 0.94])
q_samples = q_samples[(q_samples >= sample_min) & (q_samples <= sample_max)]
submission = q.get_submission_from_samples(q_samples)
submissions[q] = submission
# the graph for this question will be too zoomed out unless we cut off more of the graph
if q.id == 4153:
q.show_prediction(q_samples, plot_samples=False, plot_fitted=True, show_community=True, percent_kept=0.7)
else:
q.show_prediction(q_samples, plot_samples=False, plot_fitted=True, show_community=True, percent_kept=0.9)
# Should we submit this to Metaculus? If so, uncomment the following lines:
# for q, submission in submissions.items():
# try:
# print(q.submit(submission))
# except requests.HTTPError as e:
# print(e)
```
| github_jupyter |
```
%matplotlib inline
import pymc3 as pm
import numpy as np
import matplotlib.pyplot as plt
import pystan
import pystan.chains
from collections import OrderedDict
import pandas as pd
plt.style.use('seaborn-darkgrid')
print('Runing on PyMC3 v{}'.format(pm.__version__))
```
# Effective sample size in PyStan
Reference implementation in PyStan: [pystan/_chains.pyx
](https://github.com/stan-dev/pystan/blob/develop/pystan/_chains.pyx)
(related PR: https://github.com/stan-dev/pystan/pull/415)
```
f1 = '/usr/local/lib/python3.5/dist-packages/pystan/tests/data/blocker.1.csv'
f2 = '/usr/local/lib/python3.5/dist-packages/pystan/tests/data/blocker.2.csv'
# f1 = '/Library/Frameworks/Python.framework/Versions/3.5/lib/python3.5/site-packages/pystan/tests/data/blocker.1.csv'
# f2 = '/Library/Frameworks/Python.framework/Versions/3.5/lib/python3.5/site-packages/pystan/tests/data/blocker.2.csv'
# read csv using numpy
c1 = np.loadtxt(f1, skiprows=41, delimiter=',')[:, 4:]
c1_colnames = open(f1, 'r').readlines()[36].strip().split(',')[4:]
np.testing.assert_equal(c1_colnames[0], 'd')
c2 = np.loadtxt(f2, skiprows=41, delimiter=',')[:, 4:]
c2_colnames = open(f2, 'r').readlines()[36].strip().split(',')[4:]
np.testing.assert_equal(c1_colnames, c2_colnames)
np.testing.assert_equal(len(c1_colnames), c1.shape[1])
n_samples = len(c1)
np.testing.assert_equal(n_samples, 1000)
c1 = OrderedDict((k, v) for k, v in zip(c1_colnames, c1.T))
c2 = OrderedDict((k, v) for k, v in zip(c2_colnames, c2.T))
lst = dict(fnames_oi=c1_colnames, samples=[{'chains': c1}, {'chains': c2}],
n_save=np.repeat(n_samples, 2), permutation=None,
warmup=0, warmup2=[0, 0], chains=2, n_flatnames=len(c1))
n_eff = [
466.099, 136.953, 1170.390, 541.256,
518.051, 589.244, 764.813, 688.294,
323.777, 502.892, 353.823, 588.142,
654.336, 480.914, 176.978, 182.649,
642.389, 470.949, 561.947, 581.187,
446.389, 397.641, 338.511, 678.772,
1442.250, 837.956, 869.865, 951.124,
619.336, 875.805, 233.260, 786.568,
910.144, 231.582, 907.666, 747.347,
720.660, 195.195, 944.547, 767.271,
723.665, 1077.030, 470.903, 954.924,
497.338, 583.539, 697.204, 98.421
]
ess = []
for i in range(len(n_eff)):
ess.append(pystan.chains.ess(lst, i))
np.testing.assert_almost_equal(ess[i], n_eff[i], 2)
df_neff = pd.DataFrame(data=dict(Target=n_eff, PyStan=ess),
columns=['Target', 'PyStan'])
df_neff
lst.keys()
slst = lst['samples'][0]['chains']
slst.keys()
param_names = list(slst.keys())
slst['d'].shape
from copy import deepcopy
sim = deepcopy(lst)
m = sim['chains']
ns_save = sim['n_save']
ns_warmup2 = sim['warmup2']
ns_kept = [s - w for s, w in zip(sim['n_save'], sim['warmup2'])]
n_samples = min(ns_kept)
for im in range(m):
plt.plot(sim['samples'][im]['chains']['d']);
```
# Validate PyStan autocov implementation
```
# test autocovariance function
chain = 0
var_id = 5
stan_acov = pystan._chains._test_autocovariance(sim, chain, var_id)
len(stan_acov)
print(pystan.chains.ess(sim, var_id))
"""get_kept_samples function in pystan"""
# xtrace = []
# nv = slst[param_names[var_id]]
# for i in range(nv.shape[0] - ns_warmup2[chain]):
# xtrace.append(nv[ns_warmup2[chain] + i])
# xtrace = np.asarray(xtrace)
varname = param_names[var_id]
xtrace = sim['samples'][chain]['chains'][varname]
pymc_acov = [pm.stats.autocov(xtrace, lag)[0, 1] for lag in range(1, len(xtrace))]
import scipy.fftpack as fft
def autocorr(x):
"""
Compute autocorrelation using FFT
https://en.wikipedia.org/wiki/Autocorrelation#Efficient_computation
"""
x = np.asarray(x)
x = x - x.mean()
N = len(x)
s = fft.fft(x, N * 2 - 1)
result = np.real(fft.ifft(s * np.conjugate(s), N * 2 - 1))
result = result[:N]
result /= (N - np.arange(N))
result /= result[0]
return result
def autocov(x):
acorr = autocorr(x)
varx = np.var(x) * (len(x) - 1) / len(x)
return acorr * varx
from scipy.signal import fftconvolve
def autocorr2(x):
x -= x.mean()
N = len(x)
result = fftconvolve(x, x[::-1])
result = result[len(result) // 2:]
result /= (N - np.arange(N))
result /= result[0]
return result
def autocov2(x):
acorr = autocorr2(x)
varx = np.var(x) * (len(x) - 1) / len(x)
return acorr * varx
acov_ = autocov2(xtrace)
acov_.shape
# plt.plot(autocorr(xtrace))
# plt.plot(autocorr2(xtrace));
plt.plot(autocorr(xtrace)-autocorr2(xtrace));
x = np.random.randn(50000)
%timeit autocorr(x)
%timeit autocorr2(x)
plt.plot(np.array(stan_acov), alpha=.5)
plt.plot(np.array(pymc_acov), alpha=.5)
# plt.plot(autocov(xtrace), alpha=.5)
plt.plot(autocov2(xtrace), alpha=.5)
plt.legend();
varx = np.var(xtrace) * (len(xtrace) - 1) / len(xtrace)
plt.plot(autocorr(xtrace))
plt.acorr(xtrace, maxlags=len(xtrace)-1);
plt.plot(np.array(stan_acov) - acov_);
np.array(stan_acov)[:10]-acov_[:10]
np.array(stan_acov)[:10]-np.array(pymc_acov)[:10]
```
# PyMC3 translation of PyStan effect_n
```
from scipy.signal import fftconvolve
def autocorr(x):
"""
Compute autocorrelation using FFTfor every lag for the input array
https://en.wikipedia.org/wiki/Autocorrelation#Efficient_computation
Parameters
----------
x : Numpy array
A 1D array containing MCMC samples
Returns
-------
acorr: Numpy array same size as the input array
"""
y = x - x.mean()
N = len(y)
result = fftconvolve(y, y[::-1])
acorr = result[len(result) // 2:]
acorr /= (N - np.arange(N))
acorr /= acorr[0]
return acorr
def autocov(x):
"""Compute autocovariance estimates for every lag for
the input array
Parameters
----------
x : Numpy array
A 1D array containing MCMC samples
Returns
-------
acov: Numpy array same size as the input array
"""
acorr = autocorr(x)
varx = np.var(x, ddof=1) * (len(x) - 1) / len(x)
return acorr * varx
var_id = 13 #22
stan_acov_ = deepcopy([pystan._chains._test_autocovariance(sim, im, var_id) for im in range(m)])
trace_values = deepcopy([sim['samples'][im]['chains'][param_names[var_id]]
for im in range(m)])
x = np.array(trace_values, copy=True)
shape = x.shape
# Make sure to handle scalars correctly, adding extra dimensions if
# needed. We could use np.squeeze here, but we don't want to squeeze
# out dummy dimensions that a user inputs.
if len(shape) == 2:
x = np.atleast_3d(trace_values)
# Transpose all dimensions, which makes the loop below
# easier by moving the axes of the variable to the front instead
# of the chain and sample axes.
x = x.transpose()
for tup in np.ndindex(*list(x.shape[:-2])):
print(x[tup].shape)
trace_value = x[0].T
nchain, n_samples = trace_value.shape
acov = np.asarray([autocov(trace_value[chain]) for chain in range(nchain)])
# acov = np.asarray([stan_acov_[chain] for chain in range(nchain)])
chain_mean = np.mean(trace_value, axis=1)
chain_var = acov[:, 0] * n_samples / (n_samples - 1.)
acov_t = acov[:, 1] * n_samples / (n_samples - 1.)
mean_var = np.mean(chain_var)
var_plus = mean_var * (n_samples - 1.) / n_samples
var_plus += np.var(chain_mean, ddof=1)
rho_hat_t = np.zeros(n_samples)
rho_hat_even = 1.
rho_hat_t[0] = rho_hat_even
rho_hat_odd = 1. - (mean_var - np.mean(acov_t)) / var_plus
rho_hat_t[1] = rho_hat_odd
# Geyer's initial positive sequence
max_t = 1
t = 1
while t < (n_samples - 2) and (rho_hat_even + rho_hat_odd) >= 0:
rho_hat_even = 1. - (mean_var - np.mean(acov[:, t + 1])) / var_plus
rho_hat_odd = 1. - (mean_var - np.mean(acov[:, t + 2])) / var_plus
if (rho_hat_even + rho_hat_odd) >= 0:
rho_hat_t[t + 1] = rho_hat_even
rho_hat_t[t + 2] = rho_hat_odd
max_t = t + 2
t += 2
# Geyer's initial monotone sequence
t = 3
while t <= max_t - 2:
if (rho_hat_t[t + 1] + rho_hat_t[t + 2]) > (rho_hat_t[t - 1] + rho_hat_t[t]):
rho_hat_t[t + 1] = (rho_hat_t[t - 1] + rho_hat_t[t]) / 2.
rho_hat_t[t + 2] = rho_hat_t[t + 1]
t += 2
ess = nchain * n_samples / (-1. + 2. * np.sum(rho_hat_t))
t = 3
rho_hat_t[t + 1:t+2+1]
ess
pystan._chains.effective_sample_size(sim, var_id)
_, axes = plt.subplots(3, 1, figsize=(15, 10), sharex=True)
axes[0].plot(np.asarray(trace_values).T)
stan_acov = [pystan._chains._test_autocovariance(sim, im, var_id) for im in range(m)]
pymc_acov = [autocov(x) for x in trace_values]
for im in range(m):
axes[im+1].plot(stan_acov[im])
axes[im+1].plot(pymc_acov[im])
plt.tight_layout()
```
# New implementation
```
from scipy.signal import fftconvolve
def autocorr(x):
"""
Compute autocorrelation using FFTfor every lag for the input array
https://en.wikipedia.org/wiki/Autocorrelation#Efficient_computation
Parameters
----------
x : Numpy array
An array containing MCMC samples
Returns
-------
acorr: Numpy array same size as the input array
"""
y = x - x.mean()
N = len(y)
result = fftconvolve(y, y[::-1])
acorr = result[len(result) // 2:]
acorr /= (N - np.arange(N))
acorr /= acorr[0]
return acorr
def autocov(x):
"""Compute autocovariance estimates for every lag for the input array
Parameters
----------
x : Numpy array
An array containing MCMC samples
Returns
-------
acov: Numpy array same size as the input array
"""
acorr = autocorr(x)
varx = np.var(x, ddof=1) * (len(x) - 1) / len(x)
return acorr * varx
def get_neff(x):
trace_value = x.T
nchain, n_samples = trace_value.shape
acov = np.asarray([autocov(trace_value[chain]) for chain in range(nchain)])
chain_mean = trace_value.mean(axis=1)
chain_var = acov[:, 0] * n_samples / (n_samples - 1.)
acov_t = acov[:, 1] * n_samples / (n_samples - 1.)
mean_var = np.mean(chain_var)
var_plus = mean_var * (n_samples - 1.) / n_samples
var_plus += np.var(chain_mean, ddof=1)
rho_hat_t = np.zeros(n_samples)
rho_hat_even = 1.
rho_hat_t[0] = rho_hat_even
rho_hat_odd = 1. - (mean_var - np.mean(acov_t)) / var_plus
rho_hat_t[1] = rho_hat_odd
# Geyer's initial positive sequence
max_t = 1
t = 1
while t < (n_samples - 2) and (rho_hat_even + rho_hat_odd) >= 0.:
rho_hat_even = 1. - (mean_var - np.mean(acov[:, t + 1])) / var_plus
rho_hat_odd = 1. - (mean_var - np.mean(acov[:, t + 2])) / var_plus
if (rho_hat_even + rho_hat_odd) >= 0:
rho_hat_t[t + 1] = rho_hat_even
rho_hat_t[t + 2] = rho_hat_odd
max_t = t + 2
t += 2
# Geyer's initial monotone sequence
t = 3
while t <= max_t - 2:
if rho_hat_t[t + 1] + rho_hat_t[t + 2] > rho_hat_t[t - 1] + rho_hat_t[t]:
rho_hat_t[t + 1] = (rho_hat_t[t - 1] + rho_hat_t[t]) / 2.
rho_hat_t[t + 2] = rho_hat_t[t + 1]
t += 2
ess = nchain * n_samples
ess = ess / (-1. + 2. * np.sum(rho_hat_t))
return ess
def generate_neff(trace_values):
x = np.array(trace_values)
shape = x.shape
# Make sure to handle scalars correctly, adding extra dimensions if
# needed. We could use np.squeeze here, but we don't want to squeeze
# out dummy dimensions that a user inputs.
if len(shape) == 2:
x = np.atleast_3d(x)
# Transpose all dimensions, which makes the loop below
# easier by moving the axes of the variable to the front instead
# of the chain and sample axes.
x = x.transpose()
# Get an array the same shape as the var
_n_eff = np.zeros(x.shape[:-2])
# Iterate over tuples of indices of the shape of var
for tup in np.ndindex(*list(x.shape[:-2])):
_n_eff[tup] = get_neff(x[tup])
if len(shape) == 2:
return _n_eff[0]
return np.transpose(_n_eff)
# trace_values = tr.get_values('x', combine=False)
trace_values = [lst['samples'][0]['chains']['d'], lst['samples'][1]['chains']['d']]
generate_neff(trace_values)
ess3 = []
for varname in param_names:
trace_values = [sim['samples'][im]['chains'][varname] for im in range(m)]
ess3.append(generate_neff(trace_values))
df_neff['PyMC3_new'] = pd.Series(ess3)
df_neff
```
# Current implementation in PyMC3
```
from pymc3.stats import autocorr
def get_vhat(x):
# Chain samples are second to last dim (-2)
num_samples = x.shape[-2]
# Calculate between-chain variance
B = num_samples * np.var(np.mean(x, axis=-2), axis=-1, ddof=1)
# Calculate within-chain variance
W = np.mean(np.var(x, axis=-2, ddof=1), axis=-1)
# Estimate marginal posterior variance
Vhat = W * (num_samples - 1) / num_samples + B / num_samples
return Vhat
def get_neff(x, Vhat):
# Number of chains is last dim (-1)
num_chains = x.shape[-1]
# Chain samples are second to last dim (-2)
num_samples = x.shape[-2]
negative_autocorr = False
rho = np.ones(num_samples)
t = 1
# Iterate until the sum of consecutive estimates of autocorrelation is
# negative
while not negative_autocorr and (t < num_samples):
variogram = np.mean((x[t:, :] - x[:-t, :])**2)
rho[t] = 1. - variogram / (2. * Vhat)
negative_autocorr = sum(rho[t - 1:t + 1]) < 0
t += 1
if t % 2:
t -= 1
neff = num_chains * num_samples / (1. + 2 * rho[1:t-1].sum())
return min(num_chains * num_samples, np.floor(neff))
def generate_neff(trace_values):
x = np.array(trace_values)
shape = x.shape
# Make sure to handle scalars correctly, adding extra dimensions if
# needed. We could use np.squeeze here, but we don't want to squeeze
# out dummy dimensions that a user inputs.
if len(shape) == 2:
x = np.atleast_3d(trace_values)
# Transpose all dimensions, which makes the loop below
# easier by moving the axes of the variable to the front instead
# of the chain and sample axes.
x = x.transpose()
Vhat = get_vhat(x)
# Get an array the same shape as the var
_n_eff = np.zeros(x.shape[:-2])
# Iterate over tuples of indices of the shape of var
for tup in np.ndindex(*list(x.shape[:-2])):
_n_eff[tup] = get_neff(x[tup], Vhat[tup])
if len(shape) == 2:
return _n_eff[0]
return np.transpose(_n_eff)
#generate_neff(trace_values)
# trace_values = tr.get_values('x', combine=False)
trace_values = [lst['samples'][im]['chains'][varname] for im in range(m)]
generate_neff(trace_values)
ess2 = []
for varname in param_names:
trace_values = [lst['samples'][im]['chains'][varname] for im in range(m)]
ess2.append(generate_neff(trace_values))
df_neff['PyMC3_old'] = pd.Series(ess2)
df_neff
```
# New validation
```
cov1 = np.eye(3) * np.diag([.9, 1.2, 1.3])
M = np.random.randn(3, 3)
cov2 = M.dot(M.T)
cov2
with pm.Model() as m:
x = pm.Normal('x', shape=(3, ))
pm.MvNormal('x1', mu=x, cov=cov2, shape=(5, 3))
tr = pm.sample(cores=4)
pm.traceplot(tr);
from pymc3.util import get_default_varnames
from pymc3.backends.base import MultiTrace
from scipy.signal import fftconvolve
def autocorr(x, lag=None):
"""
Compute autocorrelation using FFT for every lag for the input array
https://en.wikipedia.org/wiki/Autocorrelation#Efficient_computation
Parameters
----------
x : Numpy array
An array containing MCMC samples
Returns
-------
acorr: Numpy array same size as the input array
"""
y = x - x.mean()
n = len(y)
result = fftconvolve(y, y[::-1])
acorr = result[len(result) // 2:]
acorr /= np.arange(n, 0, -1)
acorr /= acorr[0]
if lag is None:
return acorr
else:
warnings.warn(
"The `lag` argument has been deprecated. If you want to get "
"the value of a specific lag please call `autocorr(x)[lag]`.",
DeprecationWarning)
return acorr[lag]
def autocov(x, lag=None):
"""Compute autocovariance estimates for every lag for the input array
Parameters
----------
x : Numpy array
An array containing MCMC samples
Returns
-------
acov: Numpy array same size as the input array
"""
acorr = autocorr(x)
varx = np.var(x, ddof=1) * (len(x) - 1) / len(x)
acov = acorr * varx
if lag is None:
return acov
else:
warnings.warn(
"The `lag` argument has been deprecated. If you want to get "
"the value of a specific lag please call `autocov(x)[lag]`.",
DeprecationWarning)
return acov[lag]
def effective_n(mtrace, varnames=None, include_transformed=False):
R"""Returns estimate of the effective sample size of a set of traces.
Parameters
----------
mtrace : MultiTrace or trace object
A MultiTrace object containing parallel traces (minimum 2)
of one or more stochastic parameters.
varnames : list
Names of variables to include in the effective_n report
include_transformed : bool
Flag for reporting automatically transformed variables in addition
to original variables (defaults to False).
Returns
-------
n_eff : dictionary of floats (MultiTrace) or float (trace object)
Return the effective sample size, :math:`\hat{n}_{eff}`
Notes
-----
The diagnostic is computed by:
.. math:: \hat{n}_{eff} = \frac{mn}{1 + 2 \sum_{t=1}^T \hat{\rho}_t}
where :math:`\hat{\rho}_t` is the estimated autocorrelation at lag t, and T
is the first odd positive integer for which the sum
:math:`\hat{\rho}_{T+1} + \hat{\rho}_{T+1}` is negative.
The current implementation is similar to Stan, which uses Geyer's initial
monotone sequence criterion (Geyer, 1992; Geyer, 2011).
References
----------
Gelman et al. BDA (2014)"""
def get_neff(x):
"""Compute the effective sample size for a 2D array
"""
trace_value = x.T
nchain, n_samples = trace_value.shape
acov = np.asarray([autocov(trace_value[chain])
for chain in range(nchain)])
chain_mean = trace_value.mean(axis=1)
chain_var = acov[:, 0] * n_samples / (n_samples - 1.)
acov_t = acov[:, 1] * n_samples / (n_samples - 1.)
mean_var = np.mean(chain_var)
var_plus = mean_var * (n_samples - 1.) / n_samples
var_plus += np.var(chain_mean, ddof=1)
rho_hat_t = np.zeros(n_samples)
rho_hat_even = 1.
rho_hat_t[0] = rho_hat_even
rho_hat_odd = 1. - (mean_var - np.mean(acov_t)) / var_plus
rho_hat_t[1] = rho_hat_odd
# Geyer's initial positive sequence
max_t = 1
t = 1
while t < (n_samples - 2) and (rho_hat_even + rho_hat_odd) >= 0.:
rho_hat_even = 1. - (mean_var - np.mean(acov[:, t + 1])) / var_plus
rho_hat_odd = 1. - (mean_var - np.mean(acov[:, t + 2])) / var_plus
if (rho_hat_even + rho_hat_odd) >= 0:
rho_hat_t[t + 1] = rho_hat_even
rho_hat_t[t + 2] = rho_hat_odd
max_t = t + 2
t += 2
# Geyer's initial monotone sequence
t = 3
while t <= max_t - 2:
if (rho_hat_t[t + 1] + rho_hat_t[t + 2]) > (rho_hat_t[t - 1] + rho_hat_t[t]):
rho_hat_t[t + 1] = (rho_hat_t[t - 1] + rho_hat_t[t]) / 2.
rho_hat_t[t + 2] = rho_hat_t[t + 1]
t += 2
ess = nchain * n_samples
ess = ess / (-1. + 2. * np.sum(rho_hat_t))
return ess
def generate_neff(trace_values):
x = np.array(trace_values)
shape = x.shape
# Make sure to handle scalars correctly, adding extra dimensions if
# needed. We could use np.squeeze here, but we don't want to squeeze
# out dummy dimensions that a user inputs.
if len(shape) == 2:
x = np.atleast_3d(trace_values)
# Transpose all dimensions, which makes the loop below
# easier by moving the axes of the variable to the front instead
# of the chain and sample axes.
x = x.transpose()
# Get an array the same shape as the var
_n_eff = np.zeros(x.shape[:-2])
# Iterate over tuples of indices of the shape of var
for tup in np.ndindex(*list(x.shape[:-2])):
_n_eff[tup] = get_neff(x[tup])
if len(shape) == 2:
return _n_eff[0]
return np.transpose(_n_eff)
if not isinstance(mtrace, MultiTrace):
# Return neff for non-multitrace array
return generate_neff(mtrace)
if mtrace.nchains < 2:
raise ValueError(
'Calculation of effective sample size requires multiple chains '
'of the same length.')
if varnames is None:
varnames = get_default_varnames(
mtrace.varnames, include_transformed=include_transformed)
n_eff = {}
for var in varnames:
n_eff[var] = generate_neff(mtrace.get_values(var, combine=False))
return n_eff
%timeit effective_n(tr)
%timeit pm.effective_n(tr)
effective_n(tr)
pm.effective_n(tr)
```
| github_jupyter |
# Highly Performant TensorFlow Batch Inference on TFRecord Data Using the SageMaker CLI
In this notebook, we'll show how to use SageMaker batch transform to get inferences on a large datasets. To do this, we'll use a TensorFlow Serving model to do batch inference on a large dataset of images encoded in TFRecord format, using the AWS command-line interface. We'll show how to use the new pre-processing and post-processing feature of the TensorFlow Serving container on Amazon SageMaker so that your TensorFlow model can make inferences directly on data in S3, and save post-processed inferences to S3.
The dataset we'll be using is the [“Challenge 2018/2019"](https://github.com/cvdfoundation/open-images-dataset#download-the-open-images-challenge-28182019-test-set)” subset of the [Open Images V5 Dataset](https://storage.googleapis.com/openimages/web/index.html). This subset consists of 100,00 images in .jpg format, for a total of 10GB. For demonstration, the [model](https://github.com/tensorflow/models/tree/master/official/resnet#pre-trained-model) we'll be using is an image classification model based on the ResNet-50 architecture that has been trained on the ImageNet dataset, and which has been exported as a TensorFlow SavedModel.
We will use this model to predict the class that each model belongs to. We'll write a pre- and post-processing script and package the script with our TensorFlow SavedModel, and demonstrate how to get inferences on large datasets with SageMaker batch transform quickly, efficiently, and at scale, on GPU-accelerated instances.
## Setup
We'll begin with some necessary imports, and get an Amazon SageMaker session to help perform certain tasks, as well as an IAM role with the necessary permissions.
```
import numpy as np
import os
import sagemaker
from sagemaker import get_execution_role
sagemaker_session = sagemaker.Session()
role = get_execution_role()
region = sagemaker_session.boto_region_name
bucket = sagemaker_session.default_bucket()
prefix = 'sagemaker/DEMO-tf-batch-inference-jpeg-images-python-sdk'
uri_suffix = 'amazonaws.com'
account_id = 520713654638
account_id_cn = {
'cn-north-1': 422961961927,
'cn-northwest-1': 423003514399,
}
if region in ['cn-north-1', 'cn-northwest-1']:
uri_suffix = 'amazonaws.com.cn'
account_id = account_id_cn[region]
print('Region: {}'.format(region))
print('S3 URI: s3://{}/{}'.format(bucket, prefix))
print('Role: {}'.format(role))
```
## Inspecting the SavedModel
In order to make inferences, we'll have to preprocess our image data in S3 to match the serving signature of our TensorFlow SavedModel (https://www.tensorflow.org/guide/saved_model), which we can inspect using the saved_model_cli (https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/tools/saved_model_cli.py). This is the serving signature of the ResNet-50 v2 (NCHW, JPEG) (https://github.com/tensorflow/models/tree/master/official/resnet#pre-trained-model) model:
```
!aws s3 cp s3://sagemaker-sample-data-{region}/batch-transform/open-images/model/resnet_v2_fp32_savedmodel_NCHW_jpg.tar.gz .
!tar -zxf resnet_v2_fp32_savedmodel_NCHW_jpg.tar.gz
!saved_model_cli show --dir resnet_v2_fp32_savedmodel_NCHW_jpg/1538687370/ --all
```
The SageMaker TensorFlow Serving Container uses the model’s SignatureDef named serving_default , which is declared when the TensorFlow SavedModel is exported. This SignatureDef says that the model accepts a string of arbitrary length as input, and responds with classes and their probabilities. With our image classification model, the input string will be a base-64 encoded string representing a JPEG image, which our SavedModel will decode.
## Writing a pre- and post-processing script
We will package up our SavedModel with a Python script named `inference.py`, which will pre-process input data going from S3 to our TensorFlow Serving model, and post-process output data before it is saved back to S3:
```
!pygmentize code/inference.py
```
Additionally, we add a `requirements.txt` file, which contains additional dependencies to install from the Python Package Index:
```
!cat code/requirements.txt
```
The input_handler intercepts inference requests, base-64 encodes the request body, and formats the request body to conform to TensorFlow Serving’s REST API (https://www.tensorflow.org/tfx/serving/api_rest). The return value of the input_handler function is used as the request body in the TensorFlow Serving request.
Binary data must use key "b64", according to the TFS REST API (https://www.tensorflow.org/tfx/serving/api_rest#encoding_binary_values), and since our serving signature’s input tensor has the suffix "\_bytes", the encoded image data under key "b64" will be passed to the "image\_bytes" tensor. Some serving signatures may accept a tensor of floats or integers instead of a base-64 encoded string, but for binary data (including image data), it is recommended that your SavedModel accept a base-64 encoded string for binary data, since JSON representations of binary data can be large.
Each incoming request originally contains a serialized JPEG image in its request body, and after passing through the input_handler, the request body contains the following, which our TensorFlow Serving accepts for inference:
`{"instances": [{"b64":"[base-64 encoded JPEG image]"}]}`
The first field in the return value of `output_handler` is what SageMaker Batch Transform will save to S3 as this example’s prediction. In this case, our `output_handler` passes the content on to S3 unmodified.
Pre- and post-processing functions let you perform inference with TensorFlow Serving on any data format, not just images. To learn more about the `input_handler` and `output_handler`, consult the SageMaker TensorFlow Serving Container README (https://github.com/aws/sagemaker-tensorflow-serving-container/blob/master/README.md).
## Packaging a Model
After writing a pre- and post-processing script, you’ll need to package your TensorFlow SavedModel along with your script into a `model.tar.gz` file, which we’ll upload to S3 for the SageMaker TensorFlow Serving Container to use. Let's package the SavedModel with the `inference.py` script and examine the expected format of the `model.tar.gz` file:
```
!tar -cvzf model.tar.gz code --directory=resnet_v2_fp32_savedmodel_NCHW_jpg 1538687370
```
`1538687370` refers to the model version number of the SavedModel, and this directory contains our SavedModel artifacts. The code directory contains our pre- and post-processing script, which must be named `inference.py`. I can also include an optional `requirements.txt` file, which is used to install dependencies with `pip` from the Python Package Index before the Transform Job starts, but we don’t need any additional dependencies in this case, so we don't include a requirements file.
We will use this `model.tar.gz` when we create a SageMaker Model, which we will use to run Transform Jobs. To learn more about packaging a model, you can consult the SageMaker TensorFlow Serving Container [README](https://github.com/aws/sagemaker-tensorflow-serving-container/blob/master/README.md).
## Run a Batch Transform job
Next, we'll run a Batch Transform job using our data processing script and GPU-based Amazon SageMaker Model. More specifically, we'll perform inference on a cluster of two instances, though we can choose more or fewer. The objects in the S3 path will be distributed between the instances.
Before we create a Transform Job, let's inspect some of our input data. Here's an example, the first image in our dataset. We can inspect the format of each TFRecord file. The first record in the object named "train-00001-of-00100" refers to object "785877fb88018e89.jpg":
<img src="sample_image/785877fb88018e89.jpg">
The data in the input path consists of 100 TFRecord files, each with 1,000 JPEG images of varying sizes and shapes. Here is a subset:
```
!aws s3 ls s3://sagemaker-sample-data-{region}/batch-transform/open-images/tfrecord/ --human-readable
```
### Creating a Model and Running a Transform Job
The code below creates a SageMaker Model entity that will be used for Batch inference, and runs a Transform Job using that Model. The Model contains a reference to the TFS container, and the `model.tar.gz` containing our TensorFlow SavedModel and the pre- and post-processing `inference.py` script.
After we create a SageMaker Model, we use it to run batch predictions using Batch Transform. We specify the input S3 data, content type of the input data, the output S3 data, and instance type and count.
### Performance
For improved performance, we specify two additional parameters `max_concurrent_transforms` and `max_payload`, which control the maximum number of parallel requests that can be sent to each instance in a transform job at a time, and the maximum size of each request body.
When performing inference on entire S3 objects that cannot be split by newline characters, such as images, it is recommended that you set `max_payload` to be slightly larger than the largest S3 object in your dataset, and that you experiment with the `max_concurrent_transforms` parameter in powers of two to find a value that maximizes throughput for your model. For example, we’ve set `max_concurrent_transforms` to 64 after experimenting with powers of two, and we set `max_payload` to 1, since the largest object in our S3 input is less than one megabyte.
In addition to performance parameters, we specify AssembleWith to be “Line”, to instruct our Transform Job to assemble the individual predictions in each object by newline characters rather than concatenating them.
Furthermore, we specify certain environment variables, which are passed to the TensorFlow Serving Container, and are used to enable request batching, a TensorFlow Serving feature that allows records from multiple requests be batched together. When carefully configured, this can improve throughput, especially with GPU-accelerated inference. You can learn more about the request batching environment variables in the [SageMaker TensorFlow Serving Container repository](https://github.com/aws/sagemaker-tensorflow-serving-container#enabling-batching).
```
%%bash -s "$bucket" "$prefix" "$role" "$region" "$uri_suffix" "$account_id"
# For convenience, we pass in bucket, prefix, role, region, uri suffix and algo account id set in first Python set-up cell
BUCKET=$1
PREFIX=$2
ROLE_ARN=$3
REGION=$4
URI_SUFFIX=$5
ACCOUNT_ID=$6
timestamp() {
date +%Y-%m-%d-%H-%M-%S
}
# Creating the SageMaker Model:
MODEL_NAME="image-classification-tfs-$(timestamp)"
MODEL_DATA_URL="s3://$BUCKET/$PREFIX/model/tfrecord/model.tar.gz"
aws s3 cp model.tar.gz $MODEL_DATA_URL
# This image is maintained at https://github.com/aws/sagemaker-tensorflow-serving-container
TFS_VERSION="1.13"
PROCESSOR_TYPE="gpu"
IMAGE="$ACCOUNT_ID.dkr.ecr.$REGION.$URI_SUFFIX/sagemaker-tensorflow-serving:$TFS_VERSION-$PROCESSOR_TYPE"
aws sagemaker create-model \
--model-name $MODEL_NAME \
--primary-container Image=$IMAGE,ModelDataUrl=$MODEL_DATA_URL \
--execution-role-arn $ROLE_ARN
# Creating the Transform Job:
TRANSFORM_JOB_NAME="tfs-image-classification-job-$(timestamp)"
# Specify where to get input data and where to get output data:
TRANSFORM_S3_INPUT="s3://sagemaker-sample-data-$REGION/batch-transform/open-images/tfrecord"
TRANSFORM_S3_OUTPUT="s3://$BUCKET/$PREFIX/output"
# This configures Batch to split TFRecord files into individual records for each request
# Other options for SPLIT_TYPE include "Line" to split by newline character, and "MultiRecord"
# for BATCH_STRATEGY to include multiple records per request.
# We choose "SingleRecord" so that our own pre-processing code doesn't have to manually split TFRecords.
SPLIT_TYPE="TFRecord"
BATCH_STRATEGY="SingleRecord"
# Join outputs by newline characters. This will make JSONLines output, since each output is JSON.
ASSEMBLE_WITH="Line"
# The Data Source tells Batch to get all objects under the S3 prefix.
TRANSFORM_INPUT_DATA_SOURCE={S3DataSource={S3DataType="S3Prefix",S3Uri=$TRANSFORM_S3_INPUT}}
CONTENT_TYPE="application/x-tfexample"
DATA_SOURCE=$TRANSFORM_INPUT_DATA_SOURCE,ContentType=$CONTENT_TYPE,SplitType=$SPLIT_TYPE
# Specify resources used to transform the job
INSTANCE_TYPE="ml.p3.2xlarge"
INSTANCE_COUNT=2
# Performance parameters. MaxPayloadInMB specifies how large each request body can be.
# Our images happen to be less than 1MB, so we set MaxPayloadInMB to 1MB.
# MaxConcurrentTransforms configures the number of concurrent requests made to the container at once.
# The ideal number depends on the payload size, instance type, and model, so some experimentation
# may be beneficial.
MAX_PAYLOAD_IN_MB=1
MAX_CONCURRENT_TRANSFORMS=64
ENVIRONMENT=SAGEMAKER_TFS_ENABLE_BATCHING="true",SAGEMAKER_TFS_BATCH_TIMEOUT_MICROS="50000",SAGEMAKER_TFS_MAX_BATCH_SIZE="16"
aws sagemaker create-transform-job \
--model-name $MODEL_NAME \
--transform-input DataSource=$DATA_SOURCE \
--batch-strategy $BATCH_STRATEGY \
--transform-output S3OutputPath=$TRANSFORM_S3_OUTPUT,AssembleWith=$ASSEMBLE_WITH \
--transform-resources InstanceType=$INSTANCE_TYPE,InstanceCount=$INSTANCE_COUNT \
--max-payload-in-mb $MAX_PAYLOAD_IN_MB \
--max-concurrent-transforms $MAX_CONCURRENT_TRANSFORMS \
--transform-job-name $TRANSFORM_JOB_NAME \
--environment $ENVIRONMENT
echo "Model name: $MODEL_NAME"
echo "Transform job name: $TRANSFORM_JOB_NAME"
echo "Transform job input path: $TRANSFORM_S3_INPUT"
echo "Transform job output path: $TRANSFORM_S3_OUTPUT"
# Wait for the transform job to finish.
aws sagemaker wait transform-job-completed-or-stopped \
--transform-job-name $TRANSFORM_JOB_NAME
# Examine the output.
aws s3 ls $TRANSFORM_S3_OUTPUT --human-readable
# Copy an output example locally.
aws s3 cp $TRANSFORM_S3_OUTPUT/train-00001-of-00100.out .
```
We see that after our transform job finishes, we find one S3 object in the output path for each object in the input path. This object contains the inferences from our model for that object, and has the same name as the corresponding input object, but with `.out` appended to it.
Inspecting one of the output objects, we find the prediction from our TensorFlow Serving model. This is from the example image displayed above:
```
!head -n 1 train-00001-of-00100.out
```
## Conclusion
SageMaker batch transform can transform large datasets quickly and scalably. We used the SageMaker TensorFlow Serving Container to demonstrate how to quickly get inferences on a hundred thousand images using GPU-accelerated instances.
The Amazon SageMaker TFS container supports CSV and JSON data out of the box. The pre- and post-processing feature of the container lets you run transform jobs on data of any format. The same container can be used for real-time inference as well using an Amazon SageMaker hosted model endpoint.
| github_jupyter |
# Read in redrock output for SV1 Blanc Deep Exposures
```
!pip install git+https://github.com/desi-bgs/bgs-cmxsv.git --upgrade --user
import numpy as np
import matplotlib.pyplot as plt
from bgs_sv import sv1
# get TileIDs of Blanc deep exposures
deep_exp = sv1.blanc_deep_exposures()
deep_exp
# get redrock zbest file for deep exposure
zbest = sv1.get_zbest(deep_exp['TILEID'][9], 'deep', targetclass='bright')
zbest
# redshift success criteria
def zsuccess_criteria(zbest):
crit_zwarn = (zbest['ZWARN'] == 0)
crit_dchi2 = (zbest['DELTACHI2'] > 40.)
crit_stype = (zbest['SPECTYPE'] != "STAR") # only galaxy spectra
crit_z_lim = (zbest['Z'] > 0.0) & (zbest['Z'] < 0.6) # rough BGS redshift limit
crit_z_err = (zbest['ZERR'] < (0.0005 * (1. + zbest['Z'])))
# combine all criteria
zsuccess = crit_zwarn & crit_dchi2 & crit_stype & crit_z_lim & crit_z_err
print('%i of %i pass the redshift success criteria' % (np.sum(zsuccess), len(zsuccess)))
print('%.2f redshift success rate' % (np.sum(zsuccess)/len(zsuccess)))
return zsuccess
def zsuccess_rate(prop, zsuccess_cond, range=None, nbins=20, bin_min=2):
''' measure the redshift success rate along with property `prop`
:params prop:
array of properties (i.e. Legacy r-band magnitude)
:params zsuccess_cond:
boolean array indicating redshift success
:params range: (default: None)
range of the `prop`
:params nbins: (default: 20)
number of bins to divide `prop` by
:params bin_min: (default: 2)
minimum number of objects in bin to exlcude it
:return wmean:
weighted mean of `prop` in the bins
:return e1:
redshift success rate in the bins
:return ee1:
simple poisson error on the success rate
'''
h0, bins = np.histogram(prop, bins=nbins, range=range)
hv, _ = np.histogram(prop, bins=bins, weights=prop)
h1, _ = np.histogram(prop[zsuccess_cond], bins=bins)
good = h0 > bin_min
hv = hv[good]
h0 = h0[good]
h1 = h1[good]
wmean = hv / h0 # weighted mean
rate = h1.astype("float") / (h0.astype('float') + (h0==0))
e_rate = np.sqrt(rate * (1 - rate)) / np.sqrt(h0.astype('float') + (h0 == 0))
return wmean, rate, e_rate
# convert r-band flux to magnitude
r_mag = 22.5 - 2.5 * np.log10(zbest['FLUX_R'])
r_mid, zs, zs_err = zsuccess_rate(r_mag, zsuccess_criteria(zbest))
fig = plt.figure(figsize=(8,6))
sub = fig.add_subplot(111)
sub.errorbar(r_mid, zs, yerr=zs_err, fmt='.C0')
sub.plot(r_mid, zs)
sub.set_xlabel('$r$ mag', fontsize=25)
sub.set_ylabel('$z$ success rate', fontsize=25)
fig = plt.figure(figsize=(8,6))
sub = fig.add_subplot(111)
for i, tileid in enumerate(deep_exp['TILEID']):
try:
zbest = sv1.get_zbest(tileid, 'deep', targetclass='bright')
except:
continue
if len(zbest) < 200: continue
# convert r-band flux to magnitude
r_mag = 22.5 - 2.5 * np.log10(zbest['FLUX_R'])
r_mid, zs, zs_err = zsuccess_rate(r_mag, zsuccess_criteria(zbest))
print('tileid = %i' % tileid)
sub.errorbar(r_mid, zs, yerr=zs_err, fmt='.C%i' % (i % 10))
sub.plot(r_mid, zs, c='C%i' % (i % 10), label=str(tileid))
print()
sub.legend(loc='lower right')
sub.set_xlabel('$r$ mag', fontsize=25)
sub.set_ylabel('$z$ success rate', fontsize=25)
```
# Why do TILEID=80613 and 80618 have < 90% redshift success rate?
```
all_exps = sv1.blanc_nexp1_exposures()
all_exps[all_exps['TILEID'] == 80613]
all_exps[all_exps['TILEID'] == 80618]
```
| github_jupyter |
```
# !wget https://github.com/gouthamcm/recruit/raw/master/Entity%20Recognition%20in%20Resumes.tsv
# import pandas as pd
# df = pd.read_csv('/content/Entity Recognition in Resumes.tsv', sep='\t')
# df.head()
# from tqdm.notebook import tqdm
# ids = []
# for i, text in tqdm(enumerate(df.Abhishek)):
# if not str(text).strip():
# ids.append(i)
len(ids)
# data = df.drop(index=ids)
data.shape, df.shape
# data.Name.replace(to_replace='Can Relocate to', value='Relocate to', regex=False, inplace=True)
# data.Name.replace(to_replace='des', value='Designation', regex=False, inplace=True)
# data.Name.replace(to_replace='abc', value='O', regex=False, inplace=True)
# data.Name.replace(to_replace='work experience', value='Years of Experience', regex=False, inplace=True)
# data.Name.replace(to_replace='College Name', value='College', regex=False, inplace=True)
# data.Name.replace(to_replace='University', value='College', regex=False, inplace=True)
# data.Name.replace(to_replace='state', value='Location', regex=False, inplace=True)
# data.Name.replace(to_replace='Links', value='links', regex=False, inplace=True)
# data[data.Abhishek == '.'].head()
# data = data[:5000]
import pandas as pd
data = pd.read_csv('/content/drive/My Drive/Ideas/Resume_Analyser/Engineering/CV_Dataset/bert_relabled_v2.csv')
train_df = data[0:int(len(data)*0.8)]
test_df = data[int(len(data)*0.8):int(len(data)*0.9)]
dev_df = data[int(len(data)*0.9):]
from google.colab import drive
drive.mount('/content/drive')
int(len(data)*0.8)
train_df.shape, test_df.shape, dev_df.shape
data.head()
```
#Prepare datasets
```
from tqdm.notebook import tqdm
tokens = []
labels = []
ids = []
prev_label = ""
count = 0
sen_id = 0
for i in tqdm(range(len(dev_df))):
row = dev_df.iloc[i]
token = row.Tokens
label = row.Labels
if count > 100:
if prev_label != label:
tokens.append(" ")
labels.append(' ')
# sen_id += 1
count = 0
elif label == 'O':
tokens.append(" ")
labels.append(' ')
count = 0
# sen_id += 1
tokens.append(token)
labels.append(label)
# ids.append(sen_id)
prev_label = label
count += 1
len(tokens), len(labels)
di = {
"Tokens": tokens,
"Labels": labels
}
devd = pd.DataFrame(di)
devd.Labels.unique()
devd.to_csv("dev.txt", sep='\t', index=False)
devd[95:110]
```
Prepare train and Test
```
def process(dev_df, name):
tokens = []
labels = []
ids = []
prev_label = ""
count = 0
sen_id = 0
for i in tqdm(range(len(dev_df))):
row = dev_df.iloc[i]
token = row.Tokens
label = row.Labels
if count > 100:
if prev_label != label:
tokens.append(" ")
labels.append(' ')
# sen_id += 1
count = 0
elif label == 'O':
tokens.append(" ")
labels.append(' ')
count = 0
# sen_id += 1
tokens.append(token)
labels.append(label)
# ids.append(sen_id)
prev_label = label
count += 1
print(len(tokens), len(labels))
di = {
"Tokens": tokens,
"Labels": labels
}
devd = pd.DataFrame(di)
print(devd.Labels.unique())
devd.to_csv(name, sep='\t', index=False)
process(train_df, 'train.txt')
process(test_df, 'test.txt')
```
### Copy files
### Lets clean up memory
```
del test_df
del data
del df
del train_df
del dev_df
del tokens
del labels
```
# Install Flair
```
!pip install flair
from flair.data import Corpus
from flair.datasets import ColumnCorpus
# define columns
columns = {0: 'text', 1:'ner'}
# this is the folder in which train, test and dev files reside
data_folder = '/content/drive/My Drive/flair/data/'
# init a corpus using column format, data folder and the names of the train, dev and test files
corpus: Corpus = ColumnCorpus(data_folder, columns,
train_file='train.txt',
test_file='test.txt',
dev_file='dev.txt', # column_delimiter='\t', or use other reg pattern, use this parameters if tokens and labels not works as expected
in_memory=False,
column_delimiter='\t') # use false then it will fix ram issues
!cp dev.txt '/content/drive/My Drive/flair/data/'
len(corpus.train), len(corpus.test), len(corpus.dev)
print(corpus.train[0].to_tagged_string('ner'))
len((corpus.train[0].to_tagged_string('ner')))
tag_type = 'ner'
# 3. make the tag dictionary from the corpus
tag_dictionary = corpus.make_tag_dictionary(tag_type=tag_type)
# remember our total labels 24+ some internal labels
print(tag_dictionary)
```
if size is huge, then reduce labels
```
from flair.data import Dictionary
tags = Dictionary()
top=tag_dictionary.get_items()[:17]
bottom = tag_dictionary.get_items()[-2:]
for i in top:
tags.add_item(i)
tags.add_item('Address')
tags.add_item('projects')
for i in bottom:
tags.add_item(i)
tag_dictionary.get_items()[:17]
tags.get_items()
from flair.data import Corpus
from flair.datasets import UD_ENGLISH
from flair.embeddings import TokenEmbeddings, WordEmbeddings, StackedEmbeddings, FlairEmbeddings, TransformerWordEmbeddings, CharacterEmbeddings
embedding_types = [
WordEmbeddings('glove'),
# comment in this line to use character embeddings
# CharacterEmbeddings(), # must try
# comment in these lines to use flair embeddings
# FlairEmbeddings('news-forward'), # must try
# FlairEmbeddings('news-backward'), # must try
# TransformerWordEmbeddings('bert-base-multilingual-cased'),
]
embeddings: StackedEmbeddings = StackedEmbeddings(embeddings=embedding_types)
from flair.models import SequenceTagger
tagger: SequenceTagger = SequenceTagger(hidden_size=256,
embeddings=embeddings,
tag_dictionary=tags,
tag_type=tag_type,
use_crf=True) # use True, if gpu uses is too small
# 6. initialize trainer
from flair.trainers import ModelTrainer
trainer: ModelTrainer = ModelTrainer(tagger, corpus)
# accuracy lr hidden_size embd
# 0.21 1 256 (globe, bert)
#
# 7. start training
history = trainer.train('/content/drive/My Drive/flair/model', # saved on drive
learning_rate=0.1,
mini_batch_size=32,
patience=1,
max_epochs=30)
# check other parameters and use more epochs
from matplotlib import pyplot as plt
plt.figure(figsize=(10,5))
plt.plot(history['train_loss_history'], label="train_loss_history")
plt.plot(history['dev_loss_history'], label='dev_loss_history')
plt.legend(loc="upper right")
plt.show()
plt.figure(figsize=(10,5))
plt.plot(history['dev_score_history'])
plt.show()
```
# Test
```
st = """
Abhishek Jain
Phone: +91 8359838129
+91 7987116381
Data Scientist
E-Mail: abhishek33200jain@gmail.com
Career Objective: Seeking a position in a fast-paced,
growth-oriented organization that enables me to utilize my skills and grow as a professional and provides me the opportunity to acquire new skills through which I can prove myself to be an asset to the company.
Career Summary: Passionate, focused and confident professional with 2.5 Years of experience as Data Scientist. Currently working as a Data Science Consultant with Tiarion Software since March 2018. Before the current Role I have worked as Senior Analyst at Capgemini India Pvt Ldt.
Expertise in Python libraries NumPy, Pandas, Matplotlib, Seaborn, SciKit-Learn, TensorFlow, Keras, NLTK
Perform exploratory data analysis to understand the problem on finance Data. Used machine learning, data mining, predictive modelling & statistical techniques to create new scalable models for business requirements.
Responsible for creating actuals vs. forecast report for finance product.
Research and implement data mining machine learning algorithms in supervised and unsupervised learning areas along with that I have experience with CNN and ANN algorithms. Understating of Hadoop ecosystem and exposure to work on Pyspark.
Technical Skills: Languages :Python, SQL Machine Learning : Regression, SVM, KNN, Ensemble Learning, XGBoost, K-Means Cluster, ANN, CNN, LSTM, Text mining, Word2Vec. Tools :Jupyter, Spyder, PGAdmin, Tableau, IBM Watson(chatbot), MS Excel, Salesforce (Reporting)
Projects: Client Morgan Stanly Capital International ( MSCI) Industry Role Finance Analysis of data and build ML model Predict the future cancellation for finance product and developed the model that can predict cancellation either product will cancel or not and predict the amount of cancellation.
Description - Analyze the data set and other strategies that optimize statistical efficiency and quality. Responsibilities - Identify, analyze, and interpret trends or patterns in complex data sets. - Filter and “clean” data by reviewing visualization reports. - Work with management to prioritize business and information needs.
- When the dataset was ready, divide the data into Train and Test dataset with the ratio 70 and 30. - Implement the algorithm on Train dataset to train the model and test it with Test dataset. -Find out the confusion matrix and performance by measuring Accuracy, recall and Precision Project Name Industry Auto Ticket Tagging System.
ITSM Language Python, NLTK , Word2vec Description In this project, goal was to create the model that can automatically assignment the ticket to respective team as per the issue on email. Responsibilities - Importing the data and aggregate subject and body of the email. - Clean the text, remove the unwanted punctuation, stopwrods, extra spaces. - Implement MultiLabelBinarizer to convert the tags to one hot encoder.
- Feature extraction using word2vec model.
- When the dataset was ready, divide the data into Train and Test dataset - - Implement the algorithm on Train dataset to train the model and test it with Test dataset. Find out the performance by measuring Recall, Precision and F1 score. Work Experience : Capgemini – Senior Analyst May 2017 – March 2019 Tiarion Software- Data Scientist March 2019- Present Professional Qualification: ••••Data Science training from DataMites Bangalore Certification from International Association of Business Analytics Certification (IABAC).
Deep Learning training and Certification from Edureka. Natural Language Processing (NLP) training and certification from Analytics Vidhya. Academic Qualification: Course College/School CGPA/% 7.6 CGPA 70% Year of Passing 2016 B.E.(E.C) Intermediate Saraswathi Higher secondary School Jabalpur High School Deepak Memorial School Jabalpur Gyan Ganga Collage of Technology Jabalpur 2012 2010 77% Hobbies: Excursion, Cricket, Music Self-Declaration: I hereby declare that the above-mentioned information is correct up to my knowledge and I bear the responsibility for the correctness of the above-mentioned. Abhishek Jain
"""
from flair.data import Sentence
sentence = Sentence(st)
from flair.models import SequenceTagger
model = SequenceTagger.load('/content/drive/My Drive/flair/model/best-model.pt')
model.predict(sentence)
for entity in sentence.get_spans('ner'):
print(entity)
```
| github_jupyter |
# Auxiliary layers - DEV
Here we create
* a raster that is empty - is this useful ?
* a raster with the distance to the raster border - used for selecting pixels in a multi-tile project
* a raster with the distance to the polygon border - useful for selecting clean training samples
**TODO**: Create a Snakemake task.
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
# - - - - - - - - - - - - - - - - - - - -
# DEFAULT IMPORTS - IN ALL NOTEBOKS
from src import configs
prjconf = configs.ProjectConfigParser()
# - - - - - - - - - - - - - - - - - - - -
# NOTEBOOK SPECIFIC IMPORTS
import numpy as np
from pathlib import Path
import pandas as pd
import rasterio
from tqdm import tqdm
from eobox.raster import cube
from eobox.raster import gdalutils
from eobox.raster.rasterprocessing import create_distance_to_raster_border
from eobox.vector import calc_distance_to_border
def create_empty_raster(tile, overwrite=False):
"""Create an empty raster in the *timeless* folder from first *__QA.tif* match in 'Interim' 'hls'."""
path_empty_raster = prjconf.get_path("Interim", "timeless_empty_raster", tile=tile)
path_empty_raster.parent.mkdir(exist_ok=True, parents=True)
if not path_empty_raster.exists() or overwrite:
# get the first QA layer as a raster template. that could be anything
for pth in (prjconf.get_path("Interim", "hls") / tile).rglob("**/*__QA.tif"):
path_template_raster = pth
break
# create an empty raster
path_empty_raster.parent.mkdir(exist_ok=True, parents=True)
with rasterio.open(path_template_raster) as src:
meta = src.meta
meta.update(dtype='uint8')
arr = (src.read() * 0).astype('uint8')
with rasterio.open(path_empty_raster, 'w', **meta) as dst:
dst.write(arr)
return path_empty_raster
tilenames = prjconf.get("Params", "tiles").split(" ")
tilenames
```
## Inputs
### Parameters
```
path_ref_vector_all_tiles = {}
for tile in tilenames:
print(tile)
path_ref_vector = prjconf.get_path("Interim", "clc_subset5", tile=tile)
# create an empty raster to be used as template
create_empty_raster(tile)
if False: # NOT USED ANYMORE: THIS IS NOW DONE DURING EXTRACTION !!!
# create distance to raster border to be used in a multi-tile project
# after extraction, this auxiliar layer togehter with the coordinates helps to decide
# which pixel to take, given that they have been extracted multiple times in overlapping areas
# reasoning: we want the pixels which are most distance from the raster border
path_empty_raster = prjconf.get_path("Interim", "timeless_empty_raster", tile=tile)
path_dist2rb = prjconf.get_path("Interim", "timeless_dist2rb", tile=tile)
create_distance_to_raster_border(src_raster = path_empty_raster, # could be any raster
dst_raster = path_dist2rb,
maxdist=None, # None means we calculate distances for all pixels
overwrite=False)
# create distance to polygon border layer useful for filtering training samples / select purer inner polygon pixels
path_dist2pb = prjconf.get_path("Interim", "timeless_dist2pb", tile=tile)
path_dist2pb = str(path_dist2pb).replace("VECTORNAME", path_ref_vector.stem)
calc_distance_to_border(polygons=path_ref_vector,
template_raster=path_empty_raster,
dst_raster=path_dist2pb,
overwrite=False,
keep_interim_files=False)
# CREATED FILES
print(path_empty_raster)
print(path_dist2rb)
print(path_dist2pb)
```
| github_jupyter |
**[Data Visualization: From Non-Coder to Coder Micro-Course Home Page](https://www.kaggle.com/learn/data-visualization-from-non-coder-to-coder)**
---
Now it's time for you to demonstrate your new skills with a project of your own!
In this exercise, you will work with a dataset of your choosing. Once you've selected a dataset, you'll design and create your own plot to tell interesting stories behind the data!
## Setup
Run the next cell to import and configure the Python libraries that you need to complete the exercise.
```
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
print("Setup Complete")
```
The questions below will give you feedback on your work. Run the following cell to set up the feedback system.
```
# Set up code checking
from learntools.core import binder
binder.bind(globals())
from learntools.data_viz_to_coder.ex7 import *
print("Setup Complete")
```
## Step 1: Attach a dataset to the notebook
Begin by selecting a CSV dataset from [Kaggle Datasets](https://www.kaggle.com/datasets). If you're unsure how to do this or would like to work with your own data, please revisit the instructions in the previous tutorial.
Once you have selected a dataset, click on the **[+ ADD DATASET]** option in the top right corner. This will generate a pop-up window that you can use to search for your chosen dataset.

Once you have found the dataset, click on the **[Add]** button to attach it to the notebook. You can check that it was successful by looking at the **Workspace** dropdown menu to the right of the notebook -- look for an **input** folder containing a subfolder that matches the name of the dataset.

You can click on the carat to the right of the name of the dataset to double-check that it contains a CSV file. For instance, the image below shows that the example dataset contains two CSV files: (1) **dc-wikia-data.csv**, and (2) **marvel-wikia-data.csv**.

Once you've uploaded a dataset with a CSV file, run the code cell below **without changes** to receive credit for your work!
```
# Check for a dataset with a CSV file
step_1.check()
```
## Step 2: Specify the filepath
Now that the dataset is attached to the notebook, you can find its filepath. To do this, use the **Workspace** menu to list the set of files, and click on the CSV file you'd like to use. This will open the CSV file in a tab below the notebook. You can find the filepath towards the top of this new tab.

After you find the filepath corresponding to your dataset, fill it in as the value for `my_filepath` in the code cell below, and run the code cell to check that you've provided a valid filepath. For instance, in the case of this example dataset, we would set
```
my_filepath = "../input/dc-wikia-data.csv"
```
Note that **you must enclose the filepath in quotation marks**; otherwise, the code will return an error.
Once you've entered the filepath, you can close the tab below the notebook by clicking on the **[X]** at the top of the tab.
```
# Fill in the line below: Specify the path of the CSV file to read
my_filepath = "../input/spotify.csv"
# Check for a valid filepath to a CSV file in a dataset
step_2.check()
```
## Step 3: Load the data
Use the next code cell to load your data file into `my_data`. Use the filepath that you specified in the previous step.
```
# Fill in the line below: Read the file into a variable my_data
my_data = pd.read_csv(my_filepath)
# Check that a dataset has been uploaded into my_data
step_3.check()
```
**_After the code cell above is marked correct_**, run the code cell below without changes to view the first five rows of the data.
```
# Print the first five rows of the data
my_data.head()
```
## Step 4: Visualize the data
Use the next code cell to create a figure that tells a story behind your dataset. You can use any chart type (_line chart, bar chart, heatmap, etc_) of your choosing!
```
# Create a plot
sns.lineplot(x='Date', y='Shape of You', data=my_data) # Your code here
# Check that a figure appears below
step_4.check()
```
## Keep going
Learn how to use your skills after completing the micro-course to create data visualizations in a **[final tutorial](https://www.kaggle.com/alexisbcook/creating-your-own-notebooks)**.
---
**[Data Visualization: From Non-Coder to Coder Micro-Course Home Page](https://www.kaggle.com/learn/data-visualization-from-non-coder-to-coder)**
| github_jupyter |
# Weighted Least Squares
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import statsmodels.api as sm
from scipy import stats
from statsmodels.iolib.table import SimpleTable, default_txt_fmt
np.random.seed(1024)
```
## WLS Estimation
### Artificial data: Heteroscedasticity 2 groups
Model assumptions:
* Misspecification: true model is quadratic, estimate only linear
* Independent noise/error term
* Two groups for error variance, low and high variance groups
```
nsample = 50
x = np.linspace(0, 20, nsample)
X = np.column_stack((x, (x - 5) ** 2))
X = sm.add_constant(X)
beta = [5.0, 0.5, -0.01]
sig = 0.5
w = np.ones(nsample)
w[nsample * 6 // 10 :] = 3
y_true = np.dot(X, beta)
e = np.random.normal(size=nsample)
y = y_true + sig * w * e
X = X[:, [0, 1]]
```
### WLS knowing the true variance ratio of heteroscedasticity
In this example, `w` is the standard deviation of the error. `WLS` requires that the weights are proportional to the inverse of the error variance.
```
mod_wls = sm.WLS(y, X, weights=1.0 / (w ** 2))
res_wls = mod_wls.fit()
print(res_wls.summary())
```
## OLS vs. WLS
Estimate an OLS model for comparison:
```
res_ols = sm.OLS(y, X).fit()
print(res_ols.params)
print(res_wls.params)
```
Compare the WLS standard errors to heteroscedasticity corrected OLS standard errors:
```
se = np.vstack(
[
[res_wls.bse],
[res_ols.bse],
[res_ols.HC0_se],
[res_ols.HC1_se],
[res_ols.HC2_se],
[res_ols.HC3_se],
]
)
se = np.round(se, 4)
colnames = ["x1", "const"]
rownames = ["WLS", "OLS", "OLS_HC0", "OLS_HC1", "OLS_HC3", "OLS_HC3"]
tabl = SimpleTable(se, colnames, rownames, txt_fmt=default_txt_fmt)
print(tabl)
```
Calculate OLS prediction interval:
```
covb = res_ols.cov_params()
prediction_var = res_ols.mse_resid + (X * np.dot(covb, X.T).T).sum(1)
prediction_std = np.sqrt(prediction_var)
tppf = stats.t.ppf(0.975, res_ols.df_resid)
pred_ols = res_ols.get_prediction()
iv_l_ols = pred_ols.summary_frame()["obs_ci_lower"]
iv_u_ols = pred_ols.summary_frame()["obs_ci_upper"]
```
Draw a plot to compare predicted values in WLS and OLS:
```
pred_wls = res_wls.get_prediction()
iv_l = pred_wls.summary_frame()["obs_ci_lower"]
iv_u = pred_wls.summary_frame()["obs_ci_upper"]
fig, ax = plt.subplots(figsize=(8, 6))
ax.plot(x, y, "o", label="Data")
ax.plot(x, y_true, "b-", label="True")
# OLS
ax.plot(x, res_ols.fittedvalues, "r--")
ax.plot(x, iv_u_ols, "r--", label="OLS")
ax.plot(x, iv_l_ols, "r--")
# WLS
ax.plot(x, res_wls.fittedvalues, "g--.")
ax.plot(x, iv_u, "g--", label="WLS")
ax.plot(x, iv_l, "g--")
ax.legend(loc="best")
```
## Feasible Weighted Least Squares (2-stage FWLS)
Like `w`, `w_est` is proportional to the standard deviation, and so must be squared.
```
resid1 = res_ols.resid[w == 1.0]
var1 = resid1.var(ddof=int(res_ols.df_model) + 1)
resid2 = res_ols.resid[w != 1.0]
var2 = resid2.var(ddof=int(res_ols.df_model) + 1)
w_est = w.copy()
w_est[w != 1.0] = np.sqrt(var2) / np.sqrt(var1)
res_fwls = sm.WLS(y, X, 1.0 / ((w_est ** 2))).fit()
print(res_fwls.summary())
```
| github_jupyter |
# **Introduction to Competitive Programming**
---
Date and Time: 8th July 2019 Monday 5-7pm
Venue: Matthews Bldg RM232
Handlers: Payton Yao (Canva), Kathrina Ondap (Google)
Coordinator: Luke Sy
Repository: https://github.com/ieeeunswsb/cpworkshop
```
print("Welcome to IEEE UNSW student branch's introduction to competitive programming!")
print("We'll be having Payton and Kathy (and to some extent Luke) to guide us through this workshop!")
from urllib.request import urlretrieve
import os
def download(url, file):
if not os.path.isfile(file):
print("Download file... " + file + " ...")
urlretrieve(url,file)
print("File downloaded")
Repo = 'https://raw.githubusercontent.com/ieeeunswsb/cpworkshop/master/'
Files = ['sample_data/s01-foregone.txt',
'sample_data/s01-maxpathsum1.txt',
'sample_data/s01-maxpathsum2.txt',
'sample_data/s01-gorosort.txt',
'sample_data/s01-mutual-friend-zone.txt',
'sample_data/s01-cryptopangrams.txt']
for i in Files:
download(Repo+i, i)
```
## Problem 01: Foregone Solution (GCJ2019 Qualification)
Link: https://codingcompetitions.withgoogle.com/codejam/round/0000000000051705/0000000000088231?fbclid=IwAR2hq-J3PzLTiDXzKInwkT8CdnhJZjpdAnWH70qcojFxugkDm4HlguJRtQs
```
import sys
# comment the first line (read file) and uncomment the 2nd line (read stdin)
# when submitting your answer
with open('sample_data/s01-foregone.txt', 'r') as f:
# with sys.stdin as f:
T = int(f.readline())
for TIdx in range(T):
N = f.readline().strip()
A = ''
B = ''
for i in range(len(N)-1,-1,-1):
if N[i] == '4':
A = '2'+A
B = '2'+B
else:
A = N[i]+A
B = '0'+B
print("Case #{}: {} {}".format(TIdx+1, A.lstrip('0'), B.lstrip('0')))
```
## Problem 02: Max Sum Path I and II (Project Euler 18 and 67)
Link1: https://projecteuler.net/problem=18
Link2: https://projecteuler.net/problem=67
```
import sys
with open('sample_data/s01-maxpathsum1.txt', 'r') as f:
N = 15 # max sum path I
#N = 100 # max sum path II
t = [[0]*N for i in range(N)]
s = [[0]*N for i in range(N)]
for i in range(N-1,-1,-1):
for j0, j1 in enumerate(f.readline().split(' ')):
t[i][j0] = int(j1)
for i in range(N):
s[0][i] = t[0][i]
for i in range(1,N):
for j in range(N-i):
s[i][j] = max(s[i-1][j], s[i-1][j+1]) + t[i][j]
print(s[N-1][0])
```
## Problem 03: Gorosort (GCJ 2011 Qualification)
Link: https://code.google.com/codejam/contest/dashboard?c=975485&fbclid=IwAR1nckjG1Wpmddyb0xb1tUaeLi9hnOgVq-uY-J9P4dL-Cg9QlUUZxEmV5S0#s=p3
```
import sys
# comment the first line (read file) and uncomment the 2nd line (read stdin)
# when submitting your answer
with open('sample_data/s01-gorosort.txt', 'r') as f:
# with sys.stdin as f:
T = int(f.readline())
for TIdx in range(T):
N = int(f.readline().strip())
vals = [int(i) for i in f.readline().split(' ')]
misplaced = 0
for i in range(N):
if vals[i] != (i+1):
misplaced = misplaced+1
print('Case #{}: {:.2f}'.format(TIdx+1, misplaced))
```
## Problem 04: Mutual Friend Zone
Link: https://www.hackerrank.com/contests/noi-ph-practice-page/challenges/mutual-friendzone-not-hacked?fbclid=IwAR0Q8J0CzHcDeAfGpENaQZBjOsMvpLwbiFlT5jCajKImhqxsBkV4uH-qxQ0
Hiding the solution for now so as not to spoil it. If you did try it and wanted to see the solution, contact l.sy@unsw.edu.au.
## (EXTRA) Problem 05: Cryptopangrams
In case you find problem 1-4 TOO easy
Link: https://codingcompetitions.withgoogle.com/codejam/round/0000000000051705/000000000008830b?fbclid=IwAR3dyzboFg7sMafZ5uvGulwGLL7ow00_zr9l3z9jQcDickWXkzi_8v66wyE
Hiding the solution for now so as not to spoil it. If you did try it and wanted to see the solution, contact l.sy@unsw.edu.au.
```
```
| github_jupyter |
# Elaborate statistics features for Silvereye
## Dependencies imports
```
import xarray as xr
import os
import sys
import pandas as pd
from functools import wraps
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns # noqa, pandas aware plotting library
from datetime import date
from dateutil.relativedelta import relativedelta # $ pip install python-dateutil
```
The last section of this notebook investigates ipyleaflet for visualisation.
```
from ipyleaflet import Map, basemaps, basemap_to_tiles, ImageOverlay
import PIL
from io import StringIO, BytesIO
from base64 import b64encode
if ('SP_SRC' in os.environ):
root_src_dir = os.environ['SP_SRC']
elif sys.platform == 'win32':
root_src_dir = r'C:\src\csiro\stash\silverpieces'
else:
root_src_dir = '/silverpieces'
pkg_src_dir = root_src_dir
sys.path.append(pkg_src_dir)
from silverpieces import *
from silverpieces.functions import *
if ('SP_DATA' in os.environ):
root_data_dir = os.environ['SP_DATA']
elif sys.platform == 'win32':
root_data_dir = r'C:\data\silverpieces'
else:
root_data_dir = '/silverpieces/notebooks/'
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
# the default cmap_sequential for xarray is viridis. 'RdBu' is divergent, but works better for wetness concepts
# # https://matplotlib.org/3.1.0/tutorials/colors/colormaps.html
xr.set_options(cmap_sequential='bwr_r')
# Can get tassie_silo_rain.nc data from https://cloudstor.aarnet.edu.au/plus/s/nj2RevvD1EUD77n
fn = os.path.join(root_data_dir, 'tassie_silo_rain.nc')
tassie = xr.open_mfdataset([fn])
tassie
daily_rain = tassie.daily_rain
daily_rain.isel(time=4300).plot()
```
## Use case - inter-period statistical comparisons.
We want to be able to compare a grid of statistics for a period compared to all periods of similar lengths.
The start and end of the period should be as arbitrary as possible. The sliding window could however be limited or fixed to a year: it is probably moot to compare windows with shifted seasonality.
### How does the cumulated rainfall 2016-2018 over TAS compare with all 3 year periods over the whole record?
```
s = SpatialTemporalDataArrayStat()
start_time = pd.to_datetime('2016-01-01')
end_time = pd.to_datetime('2018-12-31')
daily_rain.load()
three_years_rains = s.periods_stat_yearly(daily_rain, start_time, end_time, func = np.sum)
three_years_rains.name = '3yrs cumulated rain'
TIME_DIMNAME = 'time'
g_simple = three_years_rains.plot(x='lon', y='lat', col=TIME_DIMNAME, col_wrap=3)
```
Let's define "percentiles" of interest as boundaries to classify against
```
q = np.array([.1, .5, .9])
y = s.quantile_over_time_dim(three_years_rains, q=q)
y.name = '3-yr cumulated rain quantiles'
```
The following color scheme may not be the best to see the map of quantile values, but should give an idea
```
y.plot(x='lon', y='lat', col='quantile', col_wrap=3, cmap='gist_ncar')
```
So now we want a map that tells us where the last three years, for every grid cell, sits (which side of every quantile)
```
last_three_years_cumrain = three_years_rains[-1,:,:]
cat_q = s.searchsorted(y, last_three_years_cumrain)
cat_q.name = 'Quantile categories 10/50/90'
cat_q.plot(cmap='bwr_r')
```
So, the three years 2016 to 2018 have been the wettest on most of the mountainous areas of the state compared to the last decade, except for the south west National Park which has been the driest comparatively.
That said, the deviation from mean may still be quite small and it may not be a "drought" as such.
Now let's look at inter-annual variability rather than 3-year moving windows.
```
cat_q
cat_q.values.shape
yearly_rain = s.periods_stat_yearly(daily_rain, '2016-01-01', '2016-12-31', func = np.sum) # Yes, xarray vanilla would suffice in this case.
yearly_rain.name = 'yearly rainfall'
yearly_rain.plot(x='lon', y='lat', col=TIME_DIMNAME, col_wrap=3)
yearly_cat_q = yearly_rain.copy()
y = s.quantile_over_time_dim(yearly_rain, q=q)
for yr in range(len(yearly_rain.time)):
x = yearly_rain[yr,:,:]
cat_q = s.searchsorted(y, x)
yearly_cat_q[yr,:,:] = cat_q
yearly_cat_q.name = 'Quantile categories yearly rain 10/50/90'
yearly_cat_q.plot(x='lon', y='lat', col=TIME_DIMNAME, col_wrap=3, cmap='bwr_r')
```
## ipyleaflet
```
from silverpieces.vis import *
bounds = make_bounds(cat_q)
imgurl = to_embedded_png(cat_q)
io = ImageOverlay(url=imgurl, bounds=bounds)
center = center_from_bounds(bounds)
zoom = 7
m = Map(center=center, zoom=zoom, interpolation='nearest')
m.layout.height = '600px'
m
m.add_layer(io)
io.interact(opacity=(0.0,1.0,0.01))
```
Try to add a colorscale legend.
```
from branca.colormap import linear
# Could not get something that displays in the widget from matplotlib colormaps
legend = linear.RdBu_09.scale(0,3)
plt.cm.bwr_r
io = ImageOverlay(url=imgurl, bounds=bounds)
io.colormap=legend
from ipywidgets.widgets import Output
out = Output(layout={'border': '1px solid black'})
with out:
display(legend)
from ipyleaflet import WidgetControl
m = Map(center=center, zoom=zoom, interpolation='nearest')
m.layout.height = '600px'
m.add_layer(io)
io.interact(opacity=(0.0,1.0,0.01))
widget_control = WidgetControl(widget=out, position='topright')
m.add_control(widget_control)
display(m)
```
| github_jupyter |
# Convolutional Layer
In this notebook, we visualize four filtered outputs (a.k.a. activation maps) of a convolutional layer.
In this example, *we* are defining four filters that are applied to an input image by initializing the **weights** of a convolutional layer, but a trained CNN will learn the values of these weights.
<img src='notebook_ims/conv_layer.gif' height=60% width=60% />
### Import the image
```
import cv2
import matplotlib.pyplot as plt
%matplotlib inline
# TODO: Feel free to try out your own images here by changing img_path
# to a file path to another image on your computer!
img_path = 'data/udacity_sdc.png'
# load color image
bgr_img = cv2.imread(img_path)
# convert to grayscale
gray_img = cv2.cvtColor(bgr_img, cv2.COLOR_BGR2GRAY)
# normalize, rescale entries to lie in [0,1]
gray_img = gray_img.astype("float32")/255
# plot image
plt.imshow(gray_img, cmap='gray')
plt.show()
```
### Define and visualize the filters
```
import numpy as np
## TODO: Feel free to modify the numbers here, to try out another filter!
filter_vals = np.array([[-1, -1, 1, 1], [-1, -1, 1, 1], [-1, -1, 1, 1], [-1, -1, 1, 1]])
print('Filter shape: ', filter_vals.shape)
# Defining four different filters,
# all of which are linear combinations of the `filter_vals` defined above
# define four filters
filter_1 = filter_vals
filter_2 = -filter_1
filter_3 = filter_1.T
filter_4 = -filter_3
filters = np.array([filter_1, filter_2, filter_3, filter_4])
# For an example, print out the values of filter 1
print('Filter 1: \n', filter_1)
print('Filter 2: \n', filter_2)
print('Filter 3: \n', filter_3)
print('Filter 4: \n', filter_4)
# visualize all four filters
fig = plt.figure(figsize=(10, 5))
for i in range(4):
ax = fig.add_subplot(1, 4, i+1, xticks=[], yticks=[])
ax.imshow(filters[i], cmap='gray')
ax.set_title('Filter %s' % str(i+1))
width, height = filters[i].shape
for x in range(width):
for y in range(height):
ax.annotate(str(filters[i][x][y]), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if filters[i][x][y]<0 else 'black')
```
## Define a convolutional layer
The various layers that make up any neural network are documented, [here](http://pytorch.org/docs/stable/nn.html). For a convolutional neural network, we'll start by defining a:
* Convolutional layer
Initialize a single convolutional layer so that it contains all your created filters. Note that you are not training this network; you are initializing the weights in a convolutional layer so that you can visualize what happens after a forward pass through this network!
#### `__init__` and `forward`
To define a neural network in PyTorch, you define the layers of a model in the function `__init__` and define the forward behavior of a network that applies those initialized layers to an input (`x`) in the function `forward`. In PyTorch we convert all inputs into the Tensor datatype, which is similar to a list data type in Python.
Below, I define the structure of a class called `Net` that has a convolutional layer that can contain four 4x4 grayscale filters.
```
import torch
import torch.nn as nn
import torch.nn.functional as F
# define a neural network with a single convolutional layer with four filters
class Net(nn.Module):
def __init__(self, weight):
super(Net, self).__init__()
# initializes the weights of the convolutional layer to be the weights of the 4 defined filters
k_height, k_width = weight.shape[2:]
# assumes there are 4 grayscale filters
self.conv = nn.Conv2d(1, 4, kernel_size=(k_height, k_width), bias=False)
self.conv.weight = torch.nn.Parameter(weight)
def forward(self, x):
# calculates the output of a convolutional layer
# pre- and post-activation
conv_x = self.conv(x)
activated_x = F.relu(conv_x)
# returns both layers
return conv_x, activated_x
# instantiate the model and set the weights
weight = torch.from_numpy(filters).unsqueeze(1).type(torch.FloatTensor)
model = Net(weight)
# print out the layer in the network
print(model)
```
### Visualize the output of each filter
First, we'll define a helper function, `viz_layer` that takes in a specific layer and number of filters (optional argument), and displays the output of that layer once an image has been passed through.
```
# helper function for visualizing the output of a given layer
# default number of filters is 4
def viz_layer(layer, n_filters= 4):
fig = plt.figure(figsize=(20, 20))
for i in range(n_filters):
ax = fig.add_subplot(1, n_filters, i+1, xticks=[], yticks=[])
# grab layer outputs
ax.imshow(np.squeeze(layer[0,i].data.numpy()), cmap='gray')
ax.set_title('Output %s' % str(i+1))
```
Let's look at the output of a convolutional layer, before and after a ReLu activation function is applied.
```
# plot original image
plt.imshow(gray_img, cmap='gray')
# visualize all filters
fig = plt.figure(figsize=(12, 6))
fig.subplots_adjust(left=0, right=1.5, bottom=0.8, top=1, hspace=0.05, wspace=0.05)
for i in range(4):
ax = fig.add_subplot(1, 4, i+1, xticks=[], yticks=[])
ax.imshow(filters[i], cmap='gray')
ax.set_title('Filter %s' % str(i+1))
# convert the image into an input Tensor
gray_img_tensor = torch.from_numpy(gray_img).unsqueeze(0).unsqueeze(1)
# get the convolutional layer (pre and post activation)
conv_layer, activated_layer = model(gray_img_tensor)
# visualize the output of a conv layer
viz_layer(conv_layer)
```
#### ReLu activation
In this model, we've used an activation function that scales the output of the convolutional layer. We've chose a ReLu function to do this, and this function simply turns all negative pixel values in 0's (black). See the equation pictured below for input pixel values, `x`.
<img src='notebook_ims/relu_ex.png' height=50% width=50% />
```
# after a ReLu is applied
# visualize the output of an activated conv layer
viz_layer(activated_layer)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.model_selection import train_test_split
from sklearn.datasets import fetch_openml
import matplotlib.pyplot as plt
import time
import warnings
warnings.filterwarnings('ignore')
```
# PCA para acelerar algoritmos de machine learning
#### Vamos a ver una prueba de concepto donde nuestro objetivo es ver como PCA puede reducir nuestro tiempo de procesamiento
Vamos a utilizar MINST dataset que se compone de imagenes de 28x28 pixeles de numeros escritos a mano.
Cada data point de MNIST es una imagen. Cada imagen es un array de 28x28 con numeros que describen cuan negro es el pixel.
Si lo ponemos en un vector cada imagen tiene una dimensionalirdad de 784 (28x28) donde cada uno tiene un valor entre zero y uno.
Mas informacion del dataset y como analizarlo:
https://colah.github.io/posts/2014-10-Visualizing-MNIST/
```
mnist = fetch_openml('mnist_784')
#Vemos que mnist es un diccionario donde los datos se encuentran en "data"
mnist
X = mnist.data[:10000]
mnist.data.shape
#Sabemos que hay 70000 imagenes con la dimensionalidad antes mencionada
np.unique(mnist.target)
#Los labels son valores que van del 0 al 9
y = mnist.target[:10000]
mnist.target.shape
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=1/7.0, random_state=0)
# Scalamos nuestros datos
scaler = StandardScaler()
scaler.fit(x_train)
x_train = scaler.transform(x_train)
x_test = scaler.transform(x_test)
```
### Vamos reducir el dataset original en diferentes proporciones y ver cuales son los tiempos de entrenamientos
#### 1 - Entendemos la relacion entre la varianza explicada y la cantidad de componentes elegidos
```
from sklearn import decomposition
pca = decomposition.PCA()
pca.n_components = 784
pca_data = pca.fit_transform(x_train)
percentage_var_explained = pca.explained_variance_ / np.sum(pca.explained_variance_);
cum_var_explained = np.cumsum(percentage_var_explained)
# Plot the PCA spectrum
plt.figure(1, figsize=(6, 4))
plt.clf()
plt.plot(cum_var_explained, linewidth=2)
plt.axis('tight')
plt.grid()
plt.xlabel('n_components')
plt.ylabel('Cumulative_explained_variance')
plt.show()
```
#### 2 - Entrenamos la regresion logistica sin aplicar PCA
```
from sklearn.linear_model import LogisticRegression
# all parameters not specified are set to their defaults
# default solver is incredibly slow which is why it was changed to 'lbfgs'
start = time.time()
logisticRegr = LogisticRegression(solver = 'lbfgs')
logisticRegr.fit(x_train, y_train)
stop = time.time()
time_100 = stop - start
print(f"Training time: {time_100}s")
score_100 = logisticRegr.score(x_test, y_test)
print("Score:", score_100)
```
#### 3 - Entrenamos la regresion logistica PCA 95% de varianza
```
# Make an instance of the Model
pca_95 = PCA(.95)
pca_95.fit(x_train)
n_95 = pca_95.n_components_
print(pca_95.n_components_ )
x_train_95 = pca_95.transform(x_train)
x_test_95 = pca_95.transform(x_test)
start_95 = time.time()
logisticRegr = LogisticRegression(solver = 'lbfgs')
logisticRegr.fit(x_train_95, y_train)
stop_95 = time.time()
time_95 = stop_95 - start_95
print(f"Training time: {time_95}s")
score_95 = logisticRegr.score(x_test_95, y_test)
print("Score:", score_95)
```
#### 4 - Entrenamos la regresion logistica PCA 80% de varianza
```
# Make an instance of the Model
pca_80 = PCA(.80)
pca_80.fit(x_train)
x_train_80 = pca_80.transform(x_train)
x_test_80 = pca_80.transform(x_test)
n_80 = pca_80.n_components_
print(n_80)
# all parameters not specified are set to their defaults
# default solver is incredibly slow which is why it was changed to 'lbfgs'
start_80 = time.time()
logisticRegr = LogisticRegression(solver = 'lbfgs')
logisticRegr.fit(x_train_80, y_train)
stop_80 = time.time()
time_80 = stop_80 - start_80
print(f"Training time: {time_80}s")
score_80 = logisticRegr.score(x_test_80, y_test)
print("Score:", score_80)
```
#### 5 - Entrenamos la regresion logistica PCA 70% de varianza
```
# Make an instance of the Model
pca_70 = PCA(.70)
pca_70.fit(x_train)
x_train_70 = pca_70.transform(x_train)
x_test_70 = pca_70.transform(x_test)
n_70 = pca_70.n_components_
print(n_70)
# all parameters not specified are set to their defaults
# default solver is incredibly slow which is why it was changed to 'lbfgs'
start_70 = time.time()
logisticRegr = LogisticRegression(solver = 'lbfgs')
logisticRegr.fit(x_train_70, y_train)
stop_70 = time.time()
time_70 = stop_70 - start_70
print(f"Training time: {time_70}s")
score_70 = logisticRegr.score(x_test_70, y_test)
print("Score:", score_70)
```
#### 6 - Resumen de los resultados encontramos. Que conclusiones sacamos?
```
pd.DataFrame(data = [[1.00, 784, time_100,score_100],
[.95, n_95, time_95, score_95],
[.80, n_80, time_80, score_80],
[.70, n_70, time_70, score_70]],
columns = ['Variance Retained',
'Number of Components',
'Time (seconds)',
'Accuracy'])
```
## Agisnacion: Obtener la tabla anterior para el iris dataset usando un algoritmo de clasificacion a eleccion
### Comparar los resultados
```
from sklearn.datasets import load_iris
data = load_iris()
data
data.target.shape
list(data.target_names)
```
| github_jupyter |
## NLP model creation and training
```
from fastai.gen_doc.nbdoc import *
from fastai.text import *
```
The main thing here is [`RNNLearner`](/text.learner.html#RNNLearner). There are also some utility functions to help create and update text models.
## Quickly get a learner
```
show_doc(language_model_learner)
```
The model used is given by `arch` and `config`. It can be:
- an [<code>AWD_LSTM</code>](/text.models.html#AWD_LSTM)([Merity et al.](https://arxiv.org/abs/1708.02182))
- a [<code>Transformer</code>](/text.models.html#Transformer) decoder ([Vaswani et al.](https://arxiv.org/abs/1706.03762))
- a [<code>TransformerXL</code>](/text.models.html#TransformerXL) ([Dai et al.](https://arxiv.org/abs/1901.02860))
They each have a default config for language modelling that is in <code>{lower_case_class_name}\_lm\_config</code> if you want to change the default parameter. At this stage, only the AWD LSTM and Tranformer support `pretrained=True` but we hope to add more pretrained models soon. `drop_mult` is applied to all the dropouts weights of the `config`, `learn_kwargs` are passed to the [`Learner`](/basic_train.html#Learner) initialization.
If your [`data`](/text.data.html#text.data) is backward, the pretrained model downloaded will also be a backard one (only available for [`AWD_LSTM`](/text.models.awd_lstm.html#AWD_LSTM)).
```
jekyll_note("Using QRNN (change the flag in the config of the AWD LSTM) requires to have cuda installed (same version as pytorch is using).")
path = untar_data(URLs.IMDB_SAMPLE)
data = TextLMDataBunch.from_csv(path, 'texts.csv')
learn = language_model_learner(data, AWD_LSTM, drop_mult=0.5)
show_doc(text_classifier_learner)
```
Here again, the backbone of the model is determined by `arch` and `config`. The input texts are fed into that model by bunch of `bptt` and only the last `max_len` activations are considered. This gives us the backbone of our model. The head then consists of:
- a layer that concatenates the final outputs of the RNN with the maximum and average of all the intermediate outputs (on the sequence length dimension),
- blocks of ([`nn.BatchNorm1d`](https://pytorch.org/docs/stable/nn.html#torch.nn.BatchNorm1d), [`nn.Dropout`](https://pytorch.org/docs/stable/nn.html#torch.nn.Dropout), [`nn.Linear`](https://pytorch.org/docs/stable/nn.html#torch.nn.Linear), [`nn.ReLU`](https://pytorch.org/docs/stable/nn.html#torch.nn.ReLU)) layers.
The blocks are defined by the `lin_ftrs` and `drops` arguments. Specifically, the first block will have a number of inputs inferred from the backbone arch and the last one will have a number of outputs equal to data.c (which contains the number of classes of the data) and the intermediate blocks have a number of inputs/outputs determined by `lin_ftrs` (of course a block has a number of inputs equal to the number of outputs of the previous block). The dropouts all have a the same value ps if you pass a float, or the corresponding values if you pass a list. Default is to have an intermediate hidden size of 50 (which makes two blocks model_activation -> 50 -> n_classes) with a dropout of 0.1.
```
path = untar_data(URLs.IMDB_SAMPLE)
data = TextClasDataBunch.from_csv(path, 'texts.csv')
learn = text_classifier_learner(data, AWD_LSTM, drop_mult=0.5)
show_doc(RNNLearner)
```
Handles the whole creation from <code>data</code> and a `model` with a text data using a certain `bptt`. The `split_func` is used to properly split the model in different groups for gradual unfreezing and differential learning rates. Gradient clipping of `clip` is optionally applied. `alpha` and `beta` are all passed to create an instance of [`RNNTrainer`](/callbacks.rnn.html#RNNTrainer). Can be used for a language model or an RNN classifier. It also handles the conversion of weights from a pretrained model as well as saving or loading the encoder.
```
show_doc(RNNLearner.get_preds)
```
If `ordered=True`, returns the predictions in the order of the dataset, otherwise they will be ordered by the sampler (from the longest text to the shortest). The other arguments are passed [`Learner.get_preds`](/basic_train.html#Learner.get_preds).
```
show_doc(TextClassificationInterpretation,title_level=3)
```
The darker the word-shading in the below example, the more it contributes to the classification. Results here are without any fitting. After fitting to acceptable accuracy, this class can show you what is being used to produce the classification of a particular case.
```
import matplotlib.cm as cm
txt_ci = TextClassificationInterpretation.from_learner(learn)
test_text = "Zombiegeddon was perhaps the GREATEST movie i have ever seen!"
txt_ci.show_intrinsic_attention(test_text,cmap=cm.Purples)
```
You can also view the raw attention values with `.intrinsic_attention(text)`
```
txt_ci.intrinsic_attention(test_text)[1]
```
Create a tabulation showing the first `k` texts in top_losses along with their prediction, actual,loss, and probability of actual class. `max_len` is the maximum number of tokens displayed. If `max_len=None`, it will display all tokens.
```
txt_ci.show_top_losses(5)
```
### Loading and saving
```
show_doc(RNNLearner.load_encoder)
show_doc(RNNLearner.save_encoder)
show_doc(RNNLearner.load_pretrained)
```
Opens the weights in the `wgts_fname` of `self.model_dir` and the dictionary in `itos_fname` then adapts the pretrained weights to the vocabulary of the <code>data</code>. The two files should be in the models directory of the `learner.path`.
## Utility functions
```
show_doc(convert_weights)
```
Uses the dictionary `stoi_wgts` (mapping of word to id) of the weights to map them to a new dictionary `itos_new` (mapping id to word).
## Get predictions
```
show_doc(LanguageLearner, title_level=3)
show_doc(LanguageLearner.predict)
```
If `no_unk=True` the unknown token is never picked. Words are taken randomly with the distribution of probabilities returned by the model. If `min_p` is not `None`, that value is the minimum probability to be considered in the pool of words. Lowering `temperature` will make the texts less randomized.
```
show_doc(LanguageLearner.beam_search)
```
## Basic functions to get a model
```
show_doc(get_language_model)
show_doc(get_text_classifier)
```
This model uses an encoder taken from the `arch` on `config`. This encoder is fed the sequence by successive bits of size `bptt` and we only keep the last `max_seq` outputs for the pooling layers.
The decoder use a concatenation of the last outputs, a `MaxPooling` of all the outputs and an `AveragePooling` of all the outputs. It then uses a list of `BatchNorm`, `Dropout`, `Linear`, `ReLU` blocks (with no `ReLU` in the last one), using a first layer size of `3*emb_sz` then following the numbers in `n_layers`. The dropouts probabilities are read in `drops`.
Note that the model returns a list of three things, the actual output being the first, the two others being the intermediate hidden states before and after dropout (used by the [`RNNTrainer`](/callbacks.rnn.html#RNNTrainer)). Most loss functions expect one output, so you should use a Callback to remove the other two if you're not using [`RNNTrainer`](/callbacks.rnn.html#RNNTrainer).
## Undocumented Methods - Methods moved below this line will intentionally be hidden
## New Methods - Please document or move to the undocumented section
```
show_doc(MultiBatchEncoder.forward)
show_doc(LanguageLearner.show_results)
show_doc(MultiBatchEncoder.concat)
show_doc(MultiBatchEncoder)
show_doc(decode_spec_tokens)
show_doc(MultiBatchEncoder.reset)
```
| github_jupyter |
# Exchanging assignment files manually
After an assignment has been created using `nbgrader generate_assignment`, the instructor must actually release that assignment to students. This page describes how to do that using your institution's existing learning management system, assuming that the students will fetch the assignments from - and submit their assignments to - the learning management system.
If this is not the case and you are using nbgrader in a shared server environment (e.g. JupyterHub), you can do this with an exchange implementation (see :doc:`managing_assignment_files`).
Distributing assignments to students and collecting them can be a logistical nightmare. The previous page discussed the built-in exchange directory, but that is not the only option (and in fact, was added later on). One can also distribute and collect files by other means, such as though your institution's learning management system. If you are relying on your institution's learning management system to get the submitted versions of notebooks back from students, ``nbgrader`` has some built-in functionality to make theat easier (putting the files in the right place into the course directory via an importer).
One can also do this fully manually, by sending files around. This may be useful during the testing phase.
## Releasing assignments
In short, to release an assignment, send the files at ``release/{assignment_id}/*`` to your students. For example, you might post the files on your course page.
## Submitting assignments
When an assignment is submitted, it needs to be placed in ``submitted/{student_id}/{assignment_id}/*``. The rest of this page describes the built-in ways to do this, if students upload them to a learning management system and you can download them all at once in an archive. This is called **collecting** the assignment.
## Collecting assignments
Once the students have submitted their assignments and you have downloaded these assignment files from your institution's learning management system, you can get theses files back into ``nbgrader`` by using the ``nbgrader zip_collect`` sub-command.
### Directory Structure:
### Workflow
### Step 1: Download submission files or archives
For demo purposes we have already created the directories needed by the ``nbgrader zip_collect`` sub-command and placed the downloaded assignment submission files and archive (zip) files in there. For example we have one ``.zip`` file and one ``.ipynb`` file:
```
%%bash
ls -l downloaded/ps1/archive
```
But before we can run the ``nbgrader zip_collect`` sub-command we first need to specify a few config options:
```
%%file nbgrader_config.py
c = get_config()
# Only set for demo purposes so as to not mess up the other documentation
c.CourseDirectory.submitted_directory = 'submitted_zip'
# Only collect submitted notebooks with valid names
c.ZipCollectApp.strict = True
# Apply this regular expression to the extracted file filename (absolute path)
c.FileNameCollectorPlugin.named_regexp = (
'.*_(?P<student_id>\w+)_attempt_'
'(?P<timestamp>[0-9\-]+)_'
'(?P<file_id>.*)'
)
```
Setting the ``strict`` flag ``True`` skips any submitted notebooks with invalid names.
By default the ``nbgrader zip_collect`` sub-command uses the ``FileNameCollectorPlugin`` to collect files from the ``extracted_directory``. This is done by sending each filename (**absolute path**) through to the ``FileNameCollectorPlugin``, which in turn applies a named group regular expression (``named_regexp``) to the filename.
The ``FileNameCollectorPlugin`` returns ``None`` if the given file should be skipped or it returns an object that must contain the ``student_id`` and ``file_id`` data, and can optionally contain the ``timestamp``, ``first_name``, ``last_name``, and ``email`` data.
Thus if using the default ``FileNameCollectorPlugin`` you must at least supply the ``student_id`` and ``file_id`` named groups. This plugin assumes all extracted files have the same filename or path structure similar to the downloaded notebook:
```
%%bash
ls -l downloaded/ps1/archive
```
Before we extract the files, we also need to have run ``nbgrader generate_assignment``:
```
%%bash
nbgrader generate_assignment "ps1" --IncludeHeaderFooter.header=source/header.ipynb --force
```
### Step 2: Extract, collect, and copy files
```
%%bash
nbgrader zip_collect ps1 --force
```
After running the ``nbgrader zip_collect`` sub-command, the archive (zip) files were extracted - and the non-archive files were copied - to the ``extracted_directory``:
```
%%bash
ls -l downloaded/ps1/extracted/
ls -l downloaded/ps1/extracted/notebooks/
```
```
%%bash
ls -l submitted_zip
%%bash
ls -l submitted_zip/hacker/ps1/
```
## Custom plugins
```
%%bash
cat submitted_zip/hacker/ps1/timestamp.txt
```
This is an issue with the underlying ``dateutils`` package used by ``nbgrader``. But not to worry, we can easily create a custom collector plugin to correct the timestamp strings when the files are collected, for example:
```
%%file plugin.py
from nbgrader.plugins import FileNameCollectorPlugin
class CustomPlugin(FileNameCollectorPlugin):
def collect(self, submission_file):
info = super(CustomPlugin, self).collect(submission_file)
if info is not None:
info['timestamp'] = '{}-{}-{} {}:{}:{}'.format(
*tuple(info['timestamp'].split('-'))
)
return info
%%bash
# Use force flag to overwrite existing files
nbgrader zip_collect --force --collector=plugin.CustomPlugin ps1
```
The ``--force`` flag is used this time to overwrite existing extracted and submitted files. Now if we check the timestamp we see it parsed correctly:
```
%%bash
cat submitted_zip/hacker/ps1/timestamp.txt
```
Note that there should only ever be *one* instructor who runs the ``nbgrader zip_collect`` command (and there should probably only be one instructor -- the same instructor -- who runs `nbgrader generate_assignment`, `nbgrader autograde` and `nbgrader formgrade` as well). However this does not mean that only one instructor can do the grading, it just means that only one instructor manages the assignment files. Other instructors can still perform grading by accessing the formgrader URL.
| github_jupyter |
# Unit 5 - Financial Planning
```
# Initial imports
import os
import requests
import pandas as pd
from dotenv import load_dotenv
import alpaca_trade_api as tradeapi
from MCForecastTools import MCSimulation
import json
%matplotlib inline
# Load .env enviroment variables
load_dotenv()
```
## Part 1 - Personal Finance Planner
### Collect Crypto Prices Using the `requests` Library
```
# Set current amount of crypto assets
my_btc = 1.2
my_eth = 5.3
crypto_data = {
"Hodlings": [1.2, 5.3]
}
# Set crypto tickers
crypto_tickers = ["BTC", "ETH"]
#display sample data
df_Hodlings = pd.DataFrame(crypto_data, index=crypto_tickers)
# Display holdings
df_Hodlings
# Crypto API URLs
import requests
btc_url = "https://api.alternative.me/v2/ticker/Bitcoin/?convert=CAD"
eth_url = "https://api.alternative.me/v2/ticker/Ethereum/?convert=CAD"
# Fetch current BTC price
my_btc_value = requests.get(btc_url).json()
# Fetch current ETH price
my_eth_value = requests.get(eth_url).json()
# Parse API JSON request and store in designated values:
print(json.dumps(my_btc_value, indent=4))
print(json.dumps(my_eth_value, indent=4))
# Parse the data for BTC and ETH
btc_price = my_btc_value["data"]["1"]["quotes"]["USD"]["price"]
eth_price = my_eth_value["data"]["1027"]["quotes"]["USD"]["price"]
# Compute current value of my crypto
my_btc_value = my_btc * btc_price
my_eth_value = my_eth * eth_price
# Print current crypto wallet balance
print(f"The current value of your {my_btc} BTC is ${my_btc_value:0.2f}")
print(f"The current value of your {my_eth} ETH is ${my_eth_value:0.2f}")
```
### Collect Investments Data Using Alpaca: `SPY` (stocks) and `AGG` (bonds)
```
# Current amount of shares
my_agg = 200
my_spy = 50
shares_data = {
"Shares": [200, 50]
}
#set the tickers
tickers = ["AGG", "SPY"]
# Display sample data
df_Shares = pd.DataFrame(shares_data, index=tickers)
# Display shares data
df_Shares
# Set Alpaca API key and secret
alpaca_api_key = os.getenv("ALPACA_API_KEY")
alpaca_secret_key = os.getenv("ALPACA_SECRET_KEY")
# Create the Alpaca API object
alpaca = tradeapi.REST(
alpaca_api_key,
alpaca_secret_key,
api_version="v2")
alpaca
# Format current date as ISO format
today = pd.Timestamp("2020-07-14", tz="America/New_York").isoformat()
today
# Set the tickers
tickers = ["AGG", "SPY"]
# Set timeframe to '1D' for Alpaca API
timeframe = "1D"
# Get current closing prices for SPY and AGG
df_portfolio = alpaca.get_barset(
tickers,
timeframe,
start = today,
end = today
).df
# Preview DataFrame
df_portfolio
# Pick AGG and SPY close prices
df_closing_prices = pd.DataFrame()
# fetch closing prices
df_closing_prices["AGG"] = df_portfolio["AGG"]["close"]
df_closing_prices["SPY"] = df_portfolio["SPY"]["close"]
# Drop the time component of the date
df_closing_prices.index = df_closing_prices.index.date
# Display sample data
df_closing_prices.head()
# Compute the current value of shares
agg_close_price = df_closing_prices["AGG"][0]
spy_close_price = df_closing_prices["SPY"][0]
my_agg_value = my_agg * agg_close_price
my_spy_value = my_spy * spy_close_price
# Print the current value of shares
print(f"The current value of your {my_agg} AGG shares is: ${my_agg_value:0.2f}")
print(f"The current value of your {my_spy}: SPY shares is: ${my_spy_value:0.2f}")
```
### Savings Health Analysis
```
# Set monthly household income
monthly_income = 12000
# add crypto and share values together:
crypto = my_btc_value + my_eth_value
shares = my_agg_value + my_spy_value
# Create savings DataFrame
data = {"Amount":[crypto, shares]}
# Create the pandas DataFrame
df_savings = pd.DataFrame(data, index = ['Crypto', 'Shares'])
# print dataframe.
print(df_savings)
# Plot savings pie chart
df_savings.plot.pie(subplots = True)
# Set ideal emergency fund
emergency_fund = monthly_income * 3
# Calculate total amount of savings
total_savings = crypto + shares
total_savings
# Validate saving health
if total_savings > emergency_fund:
print(f"Congratulations! You have enough money in your emergency fund.")
elif total_savings == emergency_fund:
print(f"Congratulations! You have enough money in your emergency fund.")
else:
print(f"Consider making a plan to save funds for an emergency.")
```
## Part 2 - Retirement Planning
### Monte Carlo Simulation
```
# Set start and end dates of five years back from today.
# Sample results may vary from the solution based on the time frame chosen
start_date = pd.Timestamp('2015-08-07', tz='America/New_York').isoformat()
end_date = pd.Timestamp('2020-08-07', tz='America/New_York').isoformat()
# Get 5 years' worth of historical data for SPY and AGG
df_portfolio = alpaca.get_barset(
tickers,
timeframe,
start = start_date,
end = end_date,
limit = 1000
).df
# Preview DataFrame
df_portfolio
# Display sample data
df_portfolio.head()
# Configuring a Monte Carlo simulation to forecast 30 years cumulative returns
?MCSimulation
# Printing the simulation input data
MC_thirty_year = MCSimulation(
portfolio_data = df_portfolio,
weights = [.60,.40],
num_simulation = 500,
num_trading_days = 252*30
)
# Running a Monte Carlo simulation to forecast 30 years cumulative returns
MC_thirty_year.portfolio_data.head()
MC_thirty_year.calc_cumulative_return()
# Plot simulation outcomes
line_plot = MC_thirty_year.plot_simulation()
# Save the plot for future usage
line_plot.get_figure().savefig("MC_thirty_year_sim_plot.png", bbox_inches="tight")
# Plot probability distribution and confidence intervals
dist_plot = MC_thirty_year.plot_distribution()
# Save the plot for future usage
dist_plot.get_figure().savefig('MC_thirty_year_dist_plot.png',bbox_inches='tight')
```
### Retirement Analysis
```
# Fetch summary statistics from the Monte Carlo simulation results
tbl = MC_thirty_year.summarize_cumulative_return()
# Print summary statistics
print(tbl)
```
### Calculate the expected portfolio return at the 95% lower and upper confidence intervals based on a `$20,000` initial investment.
```
# Set initial investment
initial_investment = 20000
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $20,000
ci_lower = round(tbl[8]*20000,2)
ci_upper = round(tbl[9]*20000,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 30 years will end within in the range of"
f" ${ci_lower} and ${ci_upper}")
```
### Calculate the expected portfolio return at the `95%` lower and upper confidence intervals based on a `50%` increase in the initial investment.
```
# Set initial investment
initial_investment = 20000 * 1.5
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $30,000
ci_lower = round(tbl[8]*30000,2)
ci_upper = round(tbl[9]*30000,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 30 years will end within in the range of"
f" ${ci_lower} and ${ci_upper}")
```
## Optional Challenge - Early Retirement
### Five Years Retirement Option
```
# Configuring a Monte Carlo simulation to forecast 5 years cumulative returns
MC_five_year = MCSimulation(
portfolio_data = df_portfolio,
weights = [.60,.40],
num_simulation = 500,
num_trading_days = 252*5
)
# Running a Monte Carlo simulation to forecast 5 years cumulative returns
MC_five_year.portfolio_data.head()
# Plot simulation outcomes
line_plot = MC_five_year.plot_simulation()
# Save the plot for future usage
line_plot.get_figure().savefig("MC_five_year_sim_plot.png", bbox_inches="tight")
# Plot probability distribution and confidence intervals
dist_plot = MC_five_year.plot_distribution()
# Save the plot for future usage
dist_plot.get_figure().savefig('MC_five_year_dist_plot.png',bbox_inches='tight')
# Fetch summary statistics from the Monte Carlo simulation results
tbl = MC_five_year.summarize_cumulative_return()
# Print summary statistics
print(tbl)
# Set initial investment
initial_investment = 120000
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $120,000
ci_lower_five = round(tbl[8]*120000,2)
ci_upper_five = round(tbl[9]*120000,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 5 years will end within in the range of"
f" ${ci_lower_five} and ${ci_upper_five}")
```
### Ten Years Retirement Option
```
# Configuring a Monte Carlo simulation to forecast 10 years cumulative returns
MC_ten_year = MCSimulation(
portfolio_data = df_portfolio,
weights = [.60,.40],
num_simulation = 500,
num_trading_days = 252*10
)
# Running a Monte Carlo simulation to forecast 10 years cumulative returns
MC_ten_year.portfolio_data.head()
# Run simulation
MC_ten_year.calc_cumulative_return()
# Plot simulation outcomes
line_plot = MC_ten_year.plot_simulation()
# Save the plot for future usage
line_plot.get_figure().savefig("MC_ten_year_sim_plot.png", bbox_inches="tight")
# Plot probability distribution and confidence intervals
dist_plot = MC_ten_year.plot_distribution()
# Save the plot for future usage
dist_plot.get_figure().savefig('MC_ten_year_dist_plot.png',bbox_inches='tight')
# Fetch summary statistics from the Monte Carlo simulation results
tbl = MC_ten_year.summarize_cumulative_return()
# Print summary statistics
print(tbl)
# Set initial investment
initial_investment = 60000
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $60,000
ci_lower_ten = round(tbl[8]*60000,2)
ci_upper_ten = round(tbl[9]*60000,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 10 years will end within in the range of"
f" ${ci_lower_ten} and ${ci_upper_ten}")
```
| github_jupyter |
# Example of optimizing a convex function
# Goal is to test the objective values found by Mango
- Search space size: Uniform
- Number of iterations to try: 40
- domain size: 5000
- Initial Random: 5
# Benchmarking test with different iterations for serial executions
```
from mango.tuner import Tuner
from scipy.stats import uniform
import math
def get_param_dict():
param_dict = {
'a': uniform(-2, 4),
'b': uniform(-2, 4)
}
return param_dict
def get_objective(x,y):
x2 = math.pow(x,2)
x4 = math.pow(x,4)
y2 = math.pow(y,2)
return ((4.0 - 2.1 * x2 + (x4 / 3.0)) * x2 + x*y + (-4.0 + 4.0 * y2) * y2)
def objfunc(args_list):
results = []
for hyper_par in args_list:
a = hyper_par['a']
b = hyper_par['b']
result = -1.0*get_objective(a,b)
results.append(result)
return results
def get_conf():
conf = dict()
conf['batch_size'] = 5
conf['initial_random'] = 5
conf['num_iteration'] = 100
conf['domain_size'] = 1000
return conf
def get_optimal_x():
param_dict = get_param_dict()
conf = get_conf()
tuner = Tuner(param_dict, objfunc,conf)
results = tuner.maximize()
return results
Store_Optimal_X = []
Store_Results = []
num_of_tries = 20
for i in range(num_of_tries):
results = get_optimal_x()
Store_Results.append(results)
print(i,":",results['best_objective'])
#results['best_objective']
len(Store_Results)
#Store_Results[0]['objective_values'][:15]
#len(Store_Results[0]['params_tried'])
```
# Extract from the results returned the true optimal values for each iteration
```
import numpy as np
total_experiments = 20
initial_random = 5
plotting_itr =[10, 20,30,40,50,60,70,80,90,100]
plotting_list = []
for exp in range(total_experiments): #for all exp
local_list = []
for itr in plotting_itr: # for all points to plot
# find the value of optimal parameters in itr+ initial_random
max_value = np.array(Store_Results[exp]['objective_values'][:itr*5+initial_random]).max()
local_list.append(max_value)
plotting_list.append(local_list)
plotting_array = np.array(plotting_list)
plotting_array.shape
#plotting_array
Y = []
#count range between -1 and 1 and show it
for i in range(len(plotting_itr)):
y_value = plotting_array[:,i].mean()
Y.append(y_value)
Y
import numpy as np
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(10,10))
plt.plot(plotting_itr,Y,label = 'Mango(Batch=5)',linewidth=4.0) #x, y
plt.xlabel('Number of Iterations',fontsize=25)
plt.ylabel('Mean optimal achieved',fontsize=25)
#plt.title('Variation of Optimal Value of X with iterations',fontsize=20)
plt.xticks(fontsize=20)
plt.yticks(fontsize=20)
#plt.yticks(np.arange(10, 110, step=10))
#plt.xticks(np.arange(10, 110, step=10))
plt.grid(True)
plt.legend(fontsize=20)
plt.show()
```
$Best_{hyper}$ = $f(Model,Dataset)$
$Best_{hyper}$, $Best_{model}$ = $f(Dataset)|_{Set\ of\ models}$
$Best_{hyper}$, $Best_{model}$ = $f(Dataset|_{Results\ on \ Similar\ datasets})|_{Set\ of\ models}$
| github_jupyter |
<img src="../../../images/qiskit-heading.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# _*Qiskit Chemistry, Programmatic Approach*_
The latest version of this notebook is available on https://github.com/Qiskit/qiskit-tutorial.
***
### Contributors
Richard Chen<sup>[1]</sup>, Antonio Mezzacapo<sup>[1]</sup>, Marco Pistoia<sup>[1]</sup>, Stephen Wood<sup>[1]</sup>
### Affiliation
- <sup>[1]</sup>IBMQ
### Introduction
In the [declarative_approach](declarative_approach.ipynb) example, we show how to configure different parameters in an input dictionary for different experiments in Qiskit Chemistry. However, many users might be intersted in experimenting with new algorithms or algorithm components, or in programming an experiment step by step using the Qiskit Chemistry APIs. This notebook illustrates how to use Qiskit Chemistry's programmatic APIs.
In this notebook, we decompose the computation of the ground state energy of a molecule into 4 steps:
1. Define a molecule and get integrals from a computational chemistry driver (PySCF in this case)
2. Construct a Fermionic Hamiltonian and map it onto a qubit Hamiltonian
3. Instantiated and initialize dynamically-loaded algorithmic components, such as the quantum algorithm VQE, the optimizer and variational form it will use, and the initial_state to initialize the variational form
4. Run the algorithm on a quantum backend and retrieve the results
```
# import common packages
import numpy as np
from qiskit import Aer
# lib from Qiskit Aqua
from qiskit.aqua import Operator, QuantumInstance
from qiskit.aqua.algorithms import VQE, ExactEigensolver
from qiskit.aqua.components.optimizers import COBYLA
# lib from Qiskit Aqua Chemistry
from qiskit.chemistry import FermionicOperator
from qiskit.chemistry.drivers import PySCFDriver, UnitsType
from qiskit.chemistry.aqua_extensions.components.variational_forms import UCCSD
from qiskit.chemistry.aqua_extensions.components.initial_states import HartreeFock
```
### Step 1: define a molecule
Here, we use LiH in sto3g basis with PySCF driver as an example.
The `molecule` object records the information from the PySCF driver.
```
# using driver to get fermionic Hamiltonian
# PySCF example
driver = PySCFDriver(atom='Li .0 .0 .0; H .0 .0 1.6', unit=UnitsType.ANGSTROM,
charge=0, spin=0, basis='sto3g')
molecule = driver.run()
```
### Step 2: Prepare qubit Hamiltonian
Here, we setup the **to-be-frozen** and **to-be-removed** orbitals to reduce the problem size when we mapping to qubit Hamiltonian. Furthermore, we define the **mapping type** for qubit Hamiltonian.
For the particular `parity` mapping, we can further reduce the problem size.
```
# please be aware that the idx here with respective to original idx
freeze_list = [0]
remove_list = [-3, -2] # negative number denotes the reverse order
map_type = 'parity'
h1 = molecule.one_body_integrals
h2 = molecule.two_body_integrals
nuclear_repulsion_energy = molecule.nuclear_repulsion_energy
num_particles = molecule.num_alpha + molecule.num_beta
num_spin_orbitals = molecule.num_orbitals * 2
print("HF energy: {}".format(molecule.hf_energy - molecule.nuclear_repulsion_energy))
print("# of electrons: {}".format(num_particles))
print("# of spin orbitals: {}".format(num_spin_orbitals))
# prepare full idx of freeze_list and remove_list
# convert all negative idx to positive
remove_list = [x % molecule.num_orbitals for x in remove_list]
freeze_list = [x % molecule.num_orbitals for x in freeze_list]
# update the idx in remove_list of the idx after frozen, since the idx of orbitals are changed after freezing
remove_list = [x - len(freeze_list) for x in remove_list]
remove_list += [x + molecule.num_orbitals - len(freeze_list) for x in remove_list]
freeze_list += [x + molecule.num_orbitals for x in freeze_list]
# prepare fermionic hamiltonian with orbital freezing and eliminating, and then map to qubit hamiltonian
# and if PARITY mapping is selected, reduction qubits
energy_shift = 0.0
qubit_reduction = True if map_type == 'parity' else False
ferOp = FermionicOperator(h1=h1, h2=h2)
if len(freeze_list) > 0:
ferOp, energy_shift = ferOp.fermion_mode_freezing(freeze_list)
num_spin_orbitals -= len(freeze_list)
num_particles -= len(freeze_list)
if len(remove_list) > 0:
ferOp = ferOp.fermion_mode_elimination(remove_list)
num_spin_orbitals -= len(remove_list)
qubitOp = ferOp.mapping(map_type=map_type, threshold=0.00000001)
qubitOp = qubitOp.two_qubit_reduced_operator(num_particles) if qubit_reduction else qubitOp
qubitOp.chop(10**-10)
```
We use the classical eigen decomposition to get the smallest eigenvalue as a reference.
```
# Using exact eigensolver to get the smallest eigenvalue
exact_eigensolver = ExactEigensolver(qubitOp, k=1)
ret = exact_eigensolver.run()
print('The computed energy is: {:.12f}'.format(ret['eigvals'][0].real))
print('The total ground state energy is: {:.12f}'.format(ret['eigvals'][0].real + energy_shift + nuclear_repulsion_energy))
```
### Step 3: Initiate and config dynamically-loaded instances
To run VQE with UCCSD variational form, we require
- VQE algorithm
- Classical Optimizer
- UCCSD variational form
- Prepare the initial state into HartreeFock state
### [Optional] Setup token to run the experiment on a real device
If you would like to run the experiement on a real device, you need to setup your account first.
Note: If you do not store your token yet, use `IBMQ.save_accounts()` to store it first.
```
# from qiskit import IBMQ
# IBMQ.load_accounts()
backend = Aer.get_backend('statevector_simulator')
# setup COBYLA optimizer
max_eval = 200
cobyla = COBYLA(maxiter=max_eval)
# setup HartreeFock state
HF_state = HartreeFock(qubitOp.num_qubits, num_spin_orbitals, num_particles, map_type,
qubit_reduction)
# setup UCCSD variational form
var_form = UCCSD(qubitOp.num_qubits, depth=1,
num_orbitals=num_spin_orbitals, num_particles=num_particles,
active_occupied=[0], active_unoccupied=[0, 1],
initial_state=HF_state, qubit_mapping=map_type,
two_qubit_reduction=qubit_reduction, num_time_slices=1)
# setup VQE
vqe = VQE(qubitOp, var_form, cobyla, 'matrix')
quantum_instance = QuantumInstance(backend=backend)
```
### Step 4: Run algorithm and retrieve the results
The smallest eigenvalue is stored in the first entry of the `eigvals` key.
```
results = vqe.run(quantum_instance)
print('The computed ground state energy is: {:.12f}'.format(results['eigvals'][0]))
print('The total ground state energy is: {:.12f}'.format(results['eigvals'][0] + energy_shift + nuclear_repulsion_energy))
print("Parameters: {}".format(results['opt_params']))
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.