text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Launch buttons for interactivity
Because Jupyter Books are built with Jupyter Notebooks, you can allow users to launch
live Jupyter sessions in the cloud directly from your book. This lets readers quickly interact
with your content in a traditional coding interface using either JupyterHub or BinderHub.
This page describes a few ways to accomplish this.
In each case, you'll need to tell Jupyter Book where your book content lives online.
To do so, use this configuration in `_config.yml`:
```yaml
# Information about where the book exists on the web
repository:
url : https://github.com/yourusername/yourbookrepo # Online location of your book
book_path : path/to/book # Optional path to your book, relative to the repository root
branch : master # Which branch of the repository should be used when creating links (optional)
```
```{admonition} MyST Markdown and launch buttons
:class: tip
If you're writing your notebooks [in MyST Markdown](../content-types/myst-notebooks.md) then launch buttons
won't open notebooks out-of-the-box. If you want notebooks to open immediately upon load, you have two options:
- Write your content in `.ipynb` files instead of MyST markdown files
- Install `jupytext` in the Binder/JupyterHub for your book, and instruct readers to right-click the markdown
file and click "Open in notebook editor".
```
## {term}`Binder` buttons for your pages
{term}`BinderHub` can be used to build the environment needed to run a repository, and provides
a link that lets others interact with that repository. If your Jupyter Book is hosted online
on GitHub, you can automatically insert buttons that link to the Jupyter Notebook running in a BinderHub.
When a user clicks the button, they'll be taken to a live version of the page. If your code
doesn't require a significant amount of CPU or RAM, you can use the free, public BinderHub running
at https://mybinder.org.
To automatically include Binder link buttons in each page of your Jupyter Book, use the following
configuration in `_config.yml`:
```yaml
launch_buttons:
binderhub_url: "https://mybinder.org" # The URL for your BinderHub (e.g., https://mybinder.org)
```
By adding this configuration, along with the repository url configuration above, Jupyter Book
will insert Binder links to any pages that were built from notebook content.
## {term}`Google Colab` buttons for your pages
If your Jupyter Book is hosted online on GitHub, you can automatically insert buttons that link to the Jupyter Notebook running on [Google Colab](https://colab.research.google.com/). When a user clicks the button, they'll be taken to a live version of the page.
Similar to Binder link buttons, you can automatically include Google Colab link buttons with the following configuration in `_config.yml`:
```yaml
launch_buttons:
colab_url: "https://colab.research.google.com"
```
```{note}
Google Colab links will only work for pages that have the `.ipynb` extension.
```
## Creating interact buttons for JupyterHub
JupyterHub lets you host an online service that gives users their own Jupyter servers
with an environment that you specify for them. It allows you to give users access to
resources and hardware that you provision in the cloud, and allows you to authenticate users
in order to control who has access to your hardware.
Similar to Binder link buttons, you can also automatically include interact links that send
your readers to a JupyterHub that is running a live, interactive version of your page. This
is accomplished using the [nbgitpuller](https://github.com/jupyterhub/nbgitpuller) server
extension.
You can configure the location of the JupyterHub (which you may set up on your own using a guide
such as [zero to jupyterhub for kubernetes](https://z2jh.jupyter.org) or [the littlest jupyterhub](https://tljh.jupyter.org)) with the following configuration.
```yaml
launch_buttons:
jupyterhub_url: "your-hub-url" # The URL for your JupyterHub. (e.g., https://datahub.berkeley.edu)
```
(launch/thebelab)=
## Live interactive pages with Thebelab
This page describes how to bring interactivity to your book. This lets users
run code and see outputs *without leaving the page*. Interactivity is provided
by a kernel running on the public [**MyBinder**](https://mybinder.org) service.
```{warning}
This is an experimental feature, and may change in the future or work unexpectedly.
```
To make your content interactive without requiring readers to leave the current page,
you can use a project called [Thebelab](https://github.com/minrk/thebelab).
This provides you a button that, when clicked, will convert each code cell into
an **interactive** cell that can be edited. It also adds a "run" button to each cell,
and connects to a Binder kernel running in the cloud.
As an alternative to pressing the Thebelab button at the top of the page, you
can press the <img src="../images/logo/edit-button.svg" alt="" style="width: 20px; display: inline;" /> symbol in the top right corner of each code cell to start the
interactive mode.
To add a Thebelab button to your Jupyter Book pages, use the following configuration:
```yaml
launch_buttons:
thebelab : true
```
In addition, you can configure the Binder settings that are used to provide a kernel for
Thebelab to run the code. These use the same configuration fields as the BinderHub interact
buttons described above.
For an example, click the **Thebelab** button above on this page, and run the code below.
```
import numpy as np
import matplotlib.pyplot as plt
plt.ion()
x = np.arange(500)
y = np.random.randn(500)
fig, ax = plt.subplots()
ax.scatter(x, y, c=y, s=x)
```
### Running cells in Thebelab when it is initialized
Sometimes you'd like to initialize the kernel that Thebelab uses by running
some code ahead of time. This might be code that you then hide from the user
in order to narrow the focus of what they interact with. This is possible
by using Jupyter Notebook tags.
Adding the tag `thebelab-init` to any code cell will cause Thebelab to
run this cell after it has received a kernel. Any subsequent Thebelab cells
will have access to the same environment (e.g. any module imports made in the
initialization cell).
You can then pair this with something like `hide-input` in order to run
initialization code that your user doesn't immediately see. For example,
below we'll initialize a variable in a hidden cell, and then tell another
cell to print the output of that variable.
```
my_hidden_variable = 'wow, it worked!'
# The variable for this is defined in the cell above!
print(my_hidden_variable)
```
| github_jupyter |
---
layout: page
title: Método Científico (Incompleto)
nav_order: 12
---
[<img src="./colab_favicon_small.png" style="float: right;">](https://colab.research.google.com/github/icd-ufmg/icd-ufmg.github.io/blob/master/_lessons/12-causalidade.ipynb)
# Método Científico (Incompleto)
{: .no_toc .mb-2 }
Juntando o método científico com o conceito de causalidade
{: .fs-6 .fw-300 }
{: .no_toc .text-delta }
Resultados Esperados
1. Entender como testes de hipótese ligam com causalidade
1. Entender o método científico
---
**Sumário**
1. TOC
{:toc}
---
```
# -*- coding: utf8
from scipy import stats as ss
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
plt.style.use('seaborn-colorblind')
plt.rcParams['figure.figsize'] = (16, 10)
plt.rcParams['axes.labelsize'] = 20
plt.rcParams['axes.titlesize'] = 20
plt.rcParams['legend.fontsize'] = 20
plt.rcParams['xtick.labelsize'] = 20
plt.rcParams['ytick.labelsize'] = 20
plt.rcParams['lines.linewidth'] = 4
plt.ion()
def despine(ax=None):
if ax is None:
ax = plt.gca()
# Hide the right and top spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
```
## Introdução
Aqui, vamos brincar um pouco com dados onde posso falar algo de causalidade. Isto é, foi feito um experimento controlado e randomizado. Note que minha ferramenta é a mesma de antes, permutação, porém a forma que os dados foram coletados mudara. Esta é a diferença.
Abaixo tenho uma função simples que permuta uma coluna de um dataframe. Vamos usar ela para implementar o nosso teste de permutação. Este exemplo é bem similar ao teste de permutação já feito. Use o mesmo para revisar!
```
def permuta(df, coluna):
'''
Permuta um dataframe com base e uma coluna categórica.
Este código é mais lento pois cria uma cópia.
Parâmetros
----------
df: o dataframe
coluna: uma coluna categórica
Retorna
-------
um novo df permutado
'''
novo = df.copy() # Cópia dos dados
dados = df[coluna].copy() # Copia da coluna, evitar um warning pandas. Deve ter forma melhor de fazer.
np.random.shuffle(dados) # Faz o shuffle
novo[coluna] = dados # Faz overwrite da coluna
return novo
```
## Dados
O DataFrame consiste de dois grupos. Um de controle, outro de tratamento. No primeiro, foi medicado placebo. No segundo, foi utilizado um novo medicamento. Quando o resultado é 1, dizemos que houve melhoria nos pacientes.
```
df = pd.read_csv('https://media.githubusercontent.com/media/icd-ufmg/material/master/aulas/13-CausalidadeRCT/bta.csv')
df.head()
control = df.query('Group == "Control"')
control.head()
medicados = df.query('Group != "Control"')
medicados.head()
```
Ao somar o resultado de cada caso, vejo quantos melhoraram. Note que o valor é bem maior nos medicados.
```
control['Result'].sum()
medicados['Result'].sum()
```
Como os dados são 1/0, a média aqui vira uma proporção. Cada observação, $x_i \in \{0, 1\}$. 0 é quando não temos um efeito positivo e 1 quando temos.
$$\sum_{i=1}^{N} x_i/n$$
```
control['Result'].mean()
medicados['Result'].mean()
```
Aqui tenho o efeito nos dados reais, mensurados em abs.
```
abs(medicados['Result'].mean() - control['Result'].mean())
tobs = abs(medicados['Result'].mean() - control['Result'].mean())
tobs
```
## Dados permutados
```
p1 = permuta(df, 'Group')
p1.head()
p1.query('Group == "Control"').mean()
p1.query('Group == "Treatment"').mean()
df.head()
```
## Teste de Permutação abaixo.
```
valores = []
for _ in range(10000):
novo = permuta(df, 'Group')
controle = novo.query('Group == "Control"')['Result']
medicados = novo.query('Group != "Control"')['Result']
valores.append(abs(controle.mean() - medicados.mean()))
valores = np.array(valores)
valores
bins = np.arange(0.15, 0.75, 0.05)
print(bins)
plt.hist(valores, bins=10, edgecolor='k')
plt.ylabel('# de Permutações')
plt.xlabel('Diferança Absoluta ao Permutar')
plt.plot([tobs], [0], 'ro', ms=15)
despine()
valor_p = (valores > tobs).mean()
valor_p
```
## Diferença de outros exemplos
Neste exemplo foi feito uma intervenção. Isto é, medicamos parte dos dados. Não estamos observando dados apenas. Por isto um exemplo como este é causal, vemos um efeito real em um experimento controlado! Mais importante do que isto, em 2011 foi averiguado que este pequeno estudo é um dos mais corretos quando se trata de dor crônica lombar! O ferramental aqui foram testes simples + uma boa amostra!
| github_jupyter |
```
from transformers import GPT2Tokenizer, GPT2LMHeadModel, AutoTokenizer, AutoModelWithLMHead, BertTokenizer, LongformerTokenizer, LongformerModel
import torch, json, random
import numpy as np
percentage = '20'
path = '../data/multiwiz/agent/'+percentage+'p/'
db_path = '../createData/multiwoz21/'
artificial_data_path = '../artificial_data/'
save_path = '../data/multiwiz/agent/'
# original ratio is the ratio of generate_artificial_data
original_ratio = 1
# final_ratio is what we want while training. Original should always be greater
final_ratio = 1
current_ratio = 1
with open(path + 'train_input.json') as f:
contexts = json.load(f)
with open(path + 'train_tgt.json') as f:
responses = json.load(f)
with open(path + 'train_query.json') as f:
queries = json.load(f)
with open(path + 'train_kb.json') as f:
kbs = json.load(f)
with open(path + 'valid_input.json') as f:
contexts_valid = json.load(f)
with open(path + 'valid_tgt.json') as f:
responses_valid = json.load(f)
with open(path + 'valid_query.json') as f:
queries_valid = json.load(f)
with open(path + 'valid_kb.json') as f:
kbs_valid = json.load(f)
# length = {}
# for t in ['restaurant', 'train', 'hotel', 'taxi', 'attraction']:
# length[t] = len(contexts[t])
# final_contexts = contexts
# final_responses = responses
# final_queries = queries
# final_kbs = kbs
# final_contexts_valid = contexts_valid
# final_responses_valid = responses_valid
# final_queries_valid = queries_valid
# final_kbs_valid = kbs_valid
# print(len(final_contexts))
# final_contexts = contexts['police']
# final_contexts.extend(contexts['hospital'])
final_contexts = contexts.copy()
final_responses=responses.copy()
final_queries=queries.copy()
# final_kbs = kbs['police']
# final_kbs.extend(kbs['hospital'])
final_kbs=kbs.copy()
# print(final_kbs[-1])
# final_contexts_valid = contexts_valid['police']
# final_contexts_valid.extend(contexts_valid['hospital'])
final_contexts_valid = contexts_valid.copy()
final_responses_valid = responses_valid.copy()
# final_queries_valid = queries_valid['police']
# final_queries_valid.extend(queries_valid['hospital'])
final_queries_valid=queries_valid.copy()
# final_kbs_valid.extend(kbs_valid['hospital'])
final_kbs_valid = kbs_valid.copy()
d_train= json.load(open(db_path + 'db/train_db.json'))
d_rest = json.load(open(db_path + 'db/restaurant_db.json'))
d_hotel = json.load(open(db_path + 'db/hotel_db.json'))
d_police = json.load(open(db_path + 'db/police_db.json'))
d_hosp = json.load(open(db_path + 'db/hospital_db.json'))
d_attr = json.load(open(db_path + 'db/attraction_db.json'))
d_taxi = [{
"taxi_colors" : ["black","white","red","yellow","blue","grey"],
"taxi_types": ["toyota","skoda","bmw","honda","ford","audi","lexus","volvo","volkswagen","tesla"],
"taxi_phone": ["^[0-9]{10}$"]
}]
entity_db_map = {'train':d_train, 'restaurant': d_rest, 'police': d_police, 'hospital': d_hosp, 'attraction': d_attr, 'taxi':d_taxi,'hotel':d_hotel}
query_key = {'train' : list(d_train[0].keys()),
'restaurant' : list(d_rest[0].keys()),
'hotel' : list(d_hotel[0].keys()),
'police' : list(d_police[0].keys()),
'hospital' : list(d_hosp[0].keys()),
'attraction' : list(d_attr[0].keys()),
'taxi' : ['taxi_colors', 'taxi_types', 'taxi_phone'],
}
def getKB(query, utt_no, domain_key):
not_kb = ['people', 'time', 'stay']
if domain_key != 'train':
not_kb.append('day')
db = entity_db_map[domain_key]
final_query = {}
for q in query.split(' | ')[1:]:
q = q.split(' = ')
for k in query_key[domain_key]:
if q[0].find(k) != -1 and q[1] != 'not mentioned' and q[1]!='dontcare' and q[1]!='none' and q[1]!='' and q[1]!='*':
final_query[k] = q[1]
ret = []
for row in db:
match = True
for k in final_query.keys():
if(k == "arriveBy"):
try:
if int(final_query[k][:2]) > int(row[k][:2]):
match=True
elif int(final_query[k][:2]) == int(row[k][:2]) and int(final_query[k][3:]) >= int(row[k][3:]):
match=True
else:
match=False
break
except:
match=True
elif(k == "leaveAt"):
try:
if int(row[k][:2]) > int(final_query[k][:2]):
match=True
elif int(row[k][:2]) == int(final_query[k][:2]) and int(row[k][3:]) >= int(final_query[k][3:]):
match=True
else:
match=False
break
except:
match=True
elif(row[k]!=final_query[k]):
match=False
break
if(match):
ret.append(row)
if len(ret) == 0:
return "[KB] " + " Total = 0 [KB]", {}
else:
# return '[KB] Total : ' + str(len(ret)) + ' ' + str(getStringKB(ret[0])), ret[0]
return "[KB] " + " Total = " + str(len(ret)) + ' [KB]', ret[0]
contexts_artificial = []
responses_artificial = []
queries_artificial = []
kbs_artificial = []
with open(artificial_data_path + 'data_all_'+percentage+'p.json') as f:
data = json.load(f)
print(len(data))
with open(artificial_data_path + 'query_all_'+percentage+'p.json') as f:
query = json.load(f)
print(len(query))
with open(artificial_data_path + 'kb_all_'+percentage+'p.json') as f:
kb = json.load(f)
print(len(kb))
for i in range(len(data)):
conversation = data[i]
conv_context = []
conv_response = []
conv_query = []
conv_kb = []
utt_no = 0
for j in range(2, len(conversation), 2):
conv_context.append(conversation[:j])
conv_response.append(conversation[j])
conv_query.append(query[i][utt_no])
conv_kb.append(kb[i][utt_no])
utt_no+=1
if i == 0:
print(conv_context)
print(conv_response)
print(conv_query)
print(conv_kb)
contexts_artificial.append(conv_context)
responses_artificial.append(conv_response)
queries_artificial.append(conv_query)
kbs_artificial.append(conv_kb)
final_contexts.extend(contexts_artificial)
final_responses.extend(responses_artificial)
final_queries.extend(queries_artificial)
final_kbs.extend(kbs_artificial)
save_dir = save_path + "augmented_mixed_goal_"+percentage+"p/"
with open(save_dir + "train_input.json", "w") as f:
json.dump(final_contexts,f,indent=4)
with open(save_dir + "train_tgt.json", "w") as f:
json.dump(final_responses,f,indent=4)
with open(save_dir + "train_kb.json", "w") as f:
json.dump(final_kbs,f,indent=4)
with open(save_dir + "train_query.json", "w") as f:
json.dump(final_queries,f,indent=4)
with open(save_dir + "valid_input.json", "w") as f:
json.dump(final_contexts_valid,f,indent=4)
with open(save_dir + "valid_tgt.json", "w") as f:
json.dump(final_responses_valid,f,indent=4)
with open(save_dir + "valid_kb.json", "w") as f:
json.dump(final_kbs_valid,f,indent=4)
with open(save_dir + "valid_query.json", "w") as f:
json.dump(final_queries_valid,f,indent=4)
len(final_contexts), len(final_responses), len(final_queries), len(final_kbs)
final_contexts[-1], final_responses[-1]
```
| github_jupyter |
# Prompt Tuning
```
import torch
colab = 'google.colab' in str(get_ipython())
# You need a T4. A K80 will not work.
if colab:
!nvidia-smi
gpu_type = torch.cuda.get_device_name(0)
if gpu_type != 'Tesla T4':
raise ValueError("I don't know about this, chief")
# Setup for Colab only
if colab:
!pip install git+https://github.com/finetuneanon/transformers@gpt-neo-localattention3
!pip install git+https://github.com/corolla-johnson/mkultra.git#egg=mkultra --log PIP_LOG
!pip install gdown
!pip install datasets
!pip install tqdm
# If on Colab, mount your Google Drive first!
if colab:
from google.colab import drive
drive.mount('/content/drive')
# Decide the length of your training blocks in tokens.
# Sizes with headroom for gpt-neo-2.7B-halved:
# - 700 on a Colab T4 (16GB)
# - 400 on a Colab K80 (12GB)
# - 32 on a GTX1080 (8GB)
# If it seems a bit small, don't worry!
# Soft prompts can be moved forward in context for the best effect.
if colab:
if gpu_type == 'Tesla T4':
block_size = 700
else:
block_size = 400
else:
block_size = 32
# Name your soft prompt project.
sp_name = 'neuromancer-x-gpt2-optuna'
# Specify the model directory or huggingface name.
if colab:
model_dir = "/content/drive/MyDrive/models/gpt-neo-2.7B-halved/"
else:
model_dir = "D:/Git Repos/mkultra/models/gpt-neo-2.7B-halved/"
# Specify the path to the text file used for training.
if colab:
text_path = "/content/drive/MyDrive/datasets/neuromancer_reformatted.txt"
else:
text_path = "datasets/neuromancer_reformatted.txt"
# Specify the project directory bases.
if colab:
project_dir_root = f"/content/drive/MyDrive/soft_prompts/{sp_name}/"
else:
project_dir_root = f"soft_prompts/{sp_name}/"
shuffle_seed = 1234567890
eval_percentage = 1.0
from mkultra.tuning import GPT2PromptTuningLM
model = GPT2PromptTuningLM.from_pretrained(model_dir).half().to("cuda")
import optuna
def objective(trial):
import os
from mkultra.trainers import SoftPromptTrainer
from transformers import Adafactor
project_dir = os.path.join(project_dir_root, f"{sp_name}-trial-{trial.number}")
optimizer = Adafactor(
params=[model.get_soft_params()],
clip_threshold=trial.suggest_float("clip_threshold", 1.0, 2.0),
decay_rate=(trial.suggest_float("decay_rate", 0.0, 0.8)-0.8),
weight_decay=trial.suggest_float("weight_decay", 1e-6, 0.5, log=True),
beta1=trial.suggest_float("beta1", 0.0, 0.99),
warmup_init=trial.suggest_categorical("warmup_init", [True, False]))
n_tokens = trial.suggest_int("n_tokens", 20, 100)
trainer = SoftPromptTrainer(
model=model,
optimizer=optimizer,
project_dir=project_dir,
text_path=text_path,
block_size=1024-n_tokens,
n_tokens=n_tokens,
shuffle_seed=shuffle_seed)
for i in range(5):
try:
trainer.shuffle_seed = shuffle_seed+i
trainer.train()
trainer.shuffle_seed = shuffle_seed
eval_loss = trainer.evaluate(eval_percentage=eval_percentage)
trial.report(eval_loss, i)
if trial.should_prune():
raise optuna.TrialPruned()
except ValueError:
# Return 10.0 for a NaN
return 10.0
return eval_loss
import optuna
study_db = "sqlite:///" + os.path.join(project_dir_root, f"trials.db")
study = optuna.create_study(direction="minimize", storage=study_db, load_if_exists=True)
study.optimize(objective, n_trials=20)
```
| github_jupyter |
# Clickstream Analysis using Apache Spark and Apache Kafka(or Message hub).
[Message Hub: Apache Kafka as a Service](https://developer.ibm.com/messaging/2016/03/14/message-hub-apache-kafka-as-a-service/), is well integrated into the IBM Data Science Experience.
Before running the notebook, you will need to setup a [Message hub](https://developer.ibm.com/messaging/message-hub/) service.
**Note:** Message hub is a paid service.
* For creating a Message hub service, go to `Data services-> Services` tab on the dashboard. Select the option to create a message hub service following the on screen instructions. Post creating service instance, select it and create a topic `clicks` with all defaults or as per your preference.
* Once the service is running, it has to be added to the current notebook. For this, first we need to create a connection for this message hub service instance. Go to `Data services-> connections` tab on dashboard and create new connection filling in the details and referring to the above created service instance in the Service instance section and then select topic `clicks`. Once done, go to `Assets` tab on the project dashboard and then click `+New data asset`. Then locate the created message hub service connection under the connections tab and click `Apply`.
* Once the service is added to the notebook, credentials to access it can be auto inserted. Please follow comments in the next step, for instructions on how to insert credentials. Once the credentials are inserted it is ready for execution.
### Credentials Section.
```
// In a Project create a Connection to a Message Hub topic and then in a Notebook in the Project use 'Insert to Code'
// to get the connection info pasted into the code cell. And rename it to credentials_1.
// Just like this:
// @hidden_cell
/*var credentials_1 = scala.collection.mutable.HashMap[String, String](
"instance_id"->"***",
"mqlight_lookup_url"->"***",
"api_key"->"***",
"kafka_admin_url"->"***",
"kafka_rest_url"->"***",
"kafka_brokers_sasl"->"ip:port",
"user"->"***",
"password"->"""****""",
"topic"->"test"
)*/
// @hidden_cell
spark.version
```
Specify the schema of the incoming wikipedia clickstream and the parse method:
```
import scala.util.Try
case class Click(prev: String, curr: String, link: String, n: Long)
def parseVal(x: Array[Byte]): Option[Click] = {
val split: Array[String] = new Predef.String(x).split("\\t")
if (split.length == 4) {
Try(Click(split(0), split(1), split(2), split(3).toLong)).toOption
} else
None
}
val user = credentials_1("user")
val pass = credentials_1("password")
val topic = credentials_1("topic")
val saslConfig =
s"""org.apache.kafka.common.security.plain.PlainLoginModule required
|debug=true
|username="$user"
|password="$pass";""".stripMargin
```
Setup structured streaming to read from Kafka:
```
val records = spark.readStream.format("kafka").option("kafka.sasl.jaas.config", saslConfig).option("subscribe", topic).option("kafka.sasl.mechanism", "PLAIN").option("kafka.security.protocol", "SASL_SSL").option("failOnDataLoss", "false").option("kafka.bootstrap.servers", credentials_1("kafka_brokers_sasl")).load()
```
Process the records:
```
val messages = records.select("value").as[Array[Byte]].flatMap(x => parseVal(x)).groupBy("curr").agg(Map("n" -> "sum")).sort($"sum(n)".desc)
```
Output to the console and start streaming:
```
val query = messages.writeStream.outputMode("complete").option("truncate", "false").format("console").start()
```
| github_jupyter |
# Welcome to hent-AI colab!
This colab can utilize Googles vast resources for super fast decensoring using this project. All you need is a Google Drive and a good amount of free space on it.
hent-AI git project page: https://github.com/natethegreate/hentAI
# Prereqs
In your Google Drive, make a folder called hent-AI. Inside that folder, make a folder called videos.
Don't worry about getting the weights or models. This repo will auto download them to this Google Drive folder for you, so make sure your drive isn't full.
# Tutorial:
Now, you can start running this notebook.
* Under the runtime option above, hit '**Change runtime type**' and make sure Hardware Accelerator is set to **GPU**.
* Then, start running the code cells below by hitting the play buttons on their top left. (Or hit Runtime->Run all). They each have comments and instructions if needed.
* *Some of the cells will require input, as a y/n box. Make sure to do those or else nothing will continue.*
* When you mount the google drive and have to authorize it, be sure to select your google account that you wish to place the models and videos on.
* When decensoring finished, the output video will be called `(video name)_decensored.avi`
* The filesystem is the window looking button on the left. Click on it, and you'll see the local hent-AI folder dropdown, and the drive folder above it.
* Expand the hent-AI folder, then the expand drive / My Drive folders
* Simply drag the decensored video avi from the hent-AI folder to the drive/My Drive folder. This will transfer the decensored video from this instance to your actual Google Drive, and is the fastest way to get the video.
* Or, you can right-click the video and download from here, but it will be much slower.
# Notes
Colab **automatically disconnects** if you dont do anything for 30 minutes, and sessions last **at most** 12 hours. Whenever you launch a new instance, You will have to run all these steps again. But, everything on your Google drive (like the can stay and it does not have to be repeated.
So, its best if you have all the clips you want to decensor ready so you can run them all at once.
```
!nvidia-smi #First check what GPU have. Tesla T4 will not work. P100 and K80s are confirmed working.
# Install conda and python 3.5.2
!pip3 install conda
!wget https://repo.anaconda.com/archive/Anaconda3-5.2.0-Linux-x86_64.sh && bash Anaconda3-5.2.0-Linux-x86_64.sh -bfp /usr/local
# You will need to confirm the yes/no prompt when installing python 3.5.2
!conda list python
!conda install python=3.5.2
# Get to cuda 9
!wget https://developer.nvidia.com/compute/cuda/9.0/Prod/local_installers/cuda-repo-ubuntu1604-9-0-local_9.0.176-1_amd64-deb
!dpkg -i cuda-repo-ubuntu1604-9-0-local_9.0.176-1_amd64-deb
!apt-key add /var/cuda-repo-9-0-local/7fa2af80.pub
!apt-get update
!apt-get install cuda=9.0.176-1
# Get hent-AI repo
!git clone https://github.com/natethegreate/hent-AI.git
%cd hent-AI/
!git checkout master
# Get ffmpeg just in case
!pip install ffmpeg-python
!add-apt-repository ppa:jon-severinsson/ffmpeg
!apt-get update
!apt-get install ffmpeg
# Mount Google Drive. Follow authentication below
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# Create directories, you'll only need to do this if you dont already have them in your drive
!mkdir /content/drive/My\ Drive/hent-AI/
!mkdir /content/drive/My\ Drive/hent-AI/videos
!mkdir /content/drive/My\ Drive/hent-AI/images
# Get models
%cd "/content/drive/My Drive/"
!wget --no-check-certificate "https://de-next.owncube.com/index.php/s/mDGmi7NgdyyQRXL/download?path=%2F&files=4x_FatalPixels_340000_G.pth&downloadStartSecret=r4q3aw60ijm" -O hent-AI/4x_FatalPixels_340000_G.pth
!wget --no-check-certificate "https://www.dropbox.com/s/zvf6vbx3hnm9r31/weights268.zip?dl=0" -O hent-AI/weights.zip
# Get requirements. This will take some time and lots of disk space. MAKE SURE TO PRESS THE "RESTART RUNTIME" BUTTON AT THE BOTTOM OF THE OUTPUT HERE
%cd /content/hent-AI/
!pip install -r requirements-gpu.txt
%cd /content/hent-AI/
!git checkout master
# Install mask rcnn
!python setup.py install
# Create folders if they are not already made. Ignore errors if they show up here.
!mkdir ESR_temp/
!mkdir ESR_temp/temp/
!mkdir ESR_temp/ESR_out/
!mkdir ESR_temp/temp2/
# Extract both the hent-AI weights and the ESRGAN weights
!unzip /content/drive/My\ Drive/hent-AI/weights.zip
# !7z x /content/drive/My\ Drive/hent-AI/4x_FatalPixels_340000_G.7z
!cp /content/drive/My\ Drive/hent-AI/4x_FatalPixels_340000_G.pth . # Auto downloader will download .pth, so no need to extract it
!ls # Verify models are inside this hent-AI folder
# Ignore this cell
# Remove tensorflow normal to operate on GPU only? NOTE: You will need to authorize both uninstalls. MAKE SURE TO PRESS THE "RESTART RUNTIME" BUTTON AT THE BOTTOM OF THE OUTPUT HERE
# !pip uninstall tensorflow
# !pip uninstall protobuf
# !pip install tensorflow==1.8.0
# !pip install --force-reinstall tensorflow-gpu==1.9.0
# Runtime may be restarted.
%cd hent-AI/
!git checkout master
!git pull #ignore me
# Make sure videos are in the videos folder inside hent-AI. You may need to confirm y/n if a video will be overwritten.
!python samples/hentai/hentai.py inference --weights=weights.h5 --sources=/content/drive/My\ Drive/hent-AI/videos/ --dtype=esrgan
# Use this if you want to detect bars on images for use with DCP. Make sure to comment-out all other lines.
# !python samples/hentai/hentai.py inference --weights=weights.h5 --sources=/content/drive/My\ Drive/hent-AI/videos/ --dtype=bar --dcpdir=/path/to/dcpdir
# Use this if you want to detect mosaics on images for use with DCP. Make sure to comment-out all other lines.
# !python samples/hentai/hentai.py inference --weights=weights.h5 --sources=/content/drive/My\ Drive/hent-AI/videos/ --dtype=mosaic --dcpdir=/path/to/dcpdir
```
Now, use the filesystem on the left to manually drag decensored videos back into your drive folder. Then they will show up in your Google drive.
| github_jupyter |
# About this kernel
The `cost_function` in this kernel is roughly 300x faster compared to the original kernel. Each function call takes roughly 37 µs.
## Reference
* (Excellent) Original Kernel: https://www.kaggle.com/inversion/santa-s-2019-starter-notebook
* First kernel that had the idea to use Numba: https://www.kaggle.com/nickel/250x-faster-cost-function-with-numba-jit
* Another great cost function optimization: https://www.kaggle.com/sekrier/fast-scoring-using-c-52-usec
```
import os
from numba import njit
import numpy as np
import pandas as pd
from tqdm import tqdm_notebook as tqdm
from lapsolver import solve_dense
import matplotlib.pyplot as plt
```
## Read in the family information and sample submission
```
fpath = './Data/family_data.csv'
data = pd.read_csv(fpath, index_col='family_id')
fpath = './Data/sample_submission.csv'
submission = pd.read_csv(fpath, index_col='family_id')
```
### Constants
```
N_DAYS = 100
MAX_OCCUPANCY = 300
MIN_OCCUPANCY = 125
family_size = data.n_people.values
days_array = np.arange(N_DAYS, 0, -1)
choice_dict = data.loc[:, 'choice_0': 'choice_9'].T.to_dict()
choice_array_num = np.full((data.shape[0], N_DAYS + 1), -1)
for i, choice in enumerate(data.loc[:, 'choice_0': 'choice_9'].values):
for d, day in enumerate(choice):
choice_array_num[i, day] = d
penalties_array = np.array([
[
0,
50,
50 + 9 * n,
100 + 9 * n,
200 + 9 * n,
200 + 18 * n,
300 + 18 * n,
300 + 36 * n,
400 + 36 * n,
500 + 36 * n + 199 * n,
500 + 36 * n + 398 * n
]
for n in range(family_size.max() + 1)
])
```
## Cost Function
```
@njit
def cost_function(prediction, penalties_array, family_size, days):
penalty = 0
# We'll use this to count the number of people scheduled each day
daily_occupancy = np.zeros((len(days)+1))
N = family_size.shape[0]
# Looping over each family; d is the day, n is size of that family,
# and choice is their top choices
for i in range(N):
# add the family member count to the daily occupancy
n = family_size[i]
d = prediction[i]
choice = choice_array_num[i]
daily_occupancy[d] += n
# Calculate the penalty for not getting top preference
penalty += penalties_array[n, choice[d]]
# for each date, check total occupancy
# (using soft constraints instead of hard constraints)
relevant_occupancy = daily_occupancy[1:]
incorrect_occupancy = np.any(
(relevant_occupancy > MAX_OCCUPANCY) |
(relevant_occupancy < MIN_OCCUPANCY)
)
if incorrect_occupancy:
penalty += 100000000
# Calculate the accounting cost
# The first day (day 100) is treated special
init_occupancy = daily_occupancy[days[0]]
accounting_cost = (init_occupancy - 125.0) / 400.0 * init_occupancy**(0.5)
# using the max function because the soft constraints might allow occupancy to dip below 125
accounting_cost = max(0, accounting_cost)
# Loop over the rest of the days, keeping track of previous count
yesterday_count = init_occupancy
for day in days[1:]:
today_count = daily_occupancy[day]
diff = np.abs(today_count - yesterday_count)
accounting_cost += max(0, (today_count - 125.0) / 400.0 * today_count**(0.5 + diff / 50.0))
yesterday_count = today_count
penalty += accounting_cost
return penalty
```
## Hungarian
```
# Start with the sample submission values
best = submission['assigned_day'].values
start_score = cost_function(best, penalties_array, family_size, days_array)
print(start_score)
def get_choice_cost(choice, n):
return penalties_array[n, choice]
def get_weights(n):
preference_mat = np.zeros((5000, 5000), dtype=int)
for i, row in data.iterrows():
preference_mat[i, :] = get_choice_cost(10,row['n_people'])
for family_id, row in data.iterrows():
for i in range(10):
choice = row["choice_{}".format(i)]
choice_cost = get_choice_cost(i, row['n_people'])
for k in range ((choice-1)*n, (choice) * n):
preference_mat[family_id, k] = choice_cost
return preference_mat
def run_hungarian(n):
weights = get_weights(n)
rids, cids = solve_dense(weights)
for r,c in zip(rids, cids):
best[r] = int(c/n)+1
score = int(cost_function(best, penalties_array, family_size, days_array))
# hist = plt.hist(best, 100, density=False, facecolor='g', alpha=0.75, weights=family_size)
return score
score = run_hungarian(50)
submission['assigned_day'] = best
submission.to_csv(f'submission_{score}.csv')
```
| github_jupyter |
# Part 3: Advanced Remote Execution Tools
In the last section we trained a toy model using Federated Learning. We did this by calling .send() and .get() on our model, sending it to the location of training data, updating it, and then bringing it back. However, at the end of the example we realized that we needed to go a bit further to protect people privacy. Namely, we want to average the gradients **before** calling .get(). That way, we won't ever see anyone's exact gradient (thus better protecting their privacy!!!)
But, in order to do this, we need a few more pieces:
- use a pointer to send a Tensor directly to another worker
And in addition, while we're here, we're going to learn about a few more advanced tensor operations as well which will help us both with this example and a few in the future!
Authors:
- Andrew Trask - Twitter: [@iamtrask](https://twitter.com/iamtrask)
```
import torch
import syft as sy
hook = sy.TorchHook(torch)
```
# Section 3.1 - Pointers to Pointers
As you know, PointerTensor objects feel just like normal tensors. In fact, they are _so much like tensors_ that we can even have pointers **to** the pointers. Check it out!
```
bob = sy.VirtualWorker(hook, id='bob')
alice = sy.VirtualWorker(hook, id='alice')
# this is a local tensor
x = torch.tensor([1,2,3,4])
x
# this sends the local tensor to Bob
x_ptr = x.send(bob)
# this is now a pointer
x_ptr
# now we can SEND THE POINTER to alice!!!
pointer_to_x_ptr = x_ptr.send(alice)
pointer_to_x_ptr
```
### What happened?
So, in the previous example, we created a tensor called `x` and send it to Bob, creating a pointer on our local machine (`x_ptr`).
Then, we called `x_ptr.send(alice)` which **sent the pointer** to Alice.
Note, this did NOT move the data! Instead, it moved the pointer to the data!!
```
# As you can see above, Bob still has the actual data (data is always stored in a LocalTensor type).
bob._objects
# Alice, on the other hand, has x_ptr!! (notice how it points at bob)
alice._objects
```
```
# and we can use .get() to get x_ptr back from Alice
x_ptr = pointer_to_x_ptr.get()
x_ptr
# and then we can use x_ptr to get x back from Bob!
x = x_ptr.get()
x
```
### Arithmetic on Pointer -> Pointer -> Data Object
And just like with normal pointers, we can perform arbitrary PyTorch operations across these tensors
```
bob._objects
alice._objects
p2p2x = torch.tensor([1,2,3,4,5]).send(bob).send(alice)
y = p2p2x + p2p2x
bob._objects
alice._objects
y.get().get()
bob._objects
alice._objects
p2p2x.get().get()
bob._objects
alice._objects
```
# Section 3.2 - Pointer Chain Operations
So in the last section whenever we called a .send() or a .get() operation, it called that operation directly on the tensor on our local machine. However, if you have a chain of pointers, sometimes you want to call operations like .get() or .send() on the **last** pointer in the chain (such as sending data directly from one worker to another). To accomplish this, you want to use functions which are especially designed for this privacy preserving operation.
These operations are:
- `my_pointer2pointer.move(another_worker)`
```
# x is now a pointer to the data which lives on Bob's machine
x = torch.tensor([1,2,3,4,5]).send(bob)
print(' bob:', bob._objects)
print('alice:',alice._objects)
x = x.move(alice)
print(' bob:', bob._objects)
print('alice:',alice._objects)
x
```
Excellent! Now we're equiped with the tools to perform remote **gradient averaging** using a trusted aggregator!
# Congratulations!!! - Time to Join the Community!
Congratulations on completing this notebook tutorial! If you enjoyed this and would like to join the movement toward privacy preserving, decentralized ownership of AI and the AI supply chain (data), you can do so in the following ways!
### Star PySyft on GitHub
The easiest way to help our community is just by starring the Repos! This helps raise awareness of the cool tools we're building.
- [Star PySyft](https://github.com/OpenMined/PySyft)
### Join our Slack!
The best way to keep up to date on the latest advancements is to join our community! You can do so by filling out the form at [http://slack.openmined.org](http://slack.openmined.org)
### Join a Code Project!
The best way to contribute to our community is to become a code contributor! At any time you can go to PySyft GitHub Issues page and filter for "Projects". This will show you all the top level Tickets giving an overview of what projects you can join! If you don't want to join a project, but you would like to do a bit of coding, you can also look for more "one off" mini-projects by searching for GitHub issues marked "good first issue".
- [PySyft Projects](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3AProject)
- [Good First Issue Tickets](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3A%22good+first+issue%22)
### Donate
If you don't have time to contribute to our codebase, but would still like to lend support, you can also become a Backer on our Open Collective. All donations go toward our web hosting and other community expenses such as hackathons and meetups!
[OpenMined's Open Collective Page](https://opencollective.com/openmined)
| github_jupyter |
# Visual GPU Log Analytics Part I: CPU Baseline in Python Pandas
Graphistry is great -- Graphistry and RAPIDS/BlazingDB is better!
This tutorial series visually analyzes Zeek/Bro network connection logs using different compute engines:
* Part I: [CPU Baseline in Python Pandas](./part_i_cpu_pandas.ipynb)
* Part II: [GPU Dataframse with RAPIDS Python cudf bindings](./part_ii_gpu_cudf)
**Part I Contents:**
Time using CPU-based Python Pandas and Graphistry for a full ETL & visual analysis flow:
1. Load data
2. Analyze data
3. Visualize data
```
#!pip install graphistry -q
import pandas as pd
import graphistry
#graphistry.register(key='MY_KEY', protocol='https', server='graphistry.site.com')
graphistry.__version__
```
## 1. Load data
```
%%time
!curl https://www.secrepo.com/maccdc2012/conn.log.gz | gzip -d > conn.log
!head -n 3 conn.log
# OPTIONAL: For slow devices, work on a subset
#!awk 'NR % 20 == 0' < conn.log > conn-5pc.log
df = pd.read_csv("./conn.log", sep="\t", header=None,
names=["time", "uid", "id.orig_h", "id.orig_p", "id.resp_h", "id.resp_p", "proto", "service",
"duration", "orig_bytes", "resp_bytes", "conn_state", "local_orig", "missed_bytes",
"history", "orig_pkts", "orig_ip_bytes", "resp_pkts", "resp_ip_bytes", "tunnel_parents"],
na_values=['-'], index_col=False)
df.sample(3)
```
## 2. Analyze Data
Summarize network activities between every communicating src/dst IP, split by connection state
```
df_summary = df\
.assign(
sum_bytes=df.apply(lambda row: row['orig_bytes'] + row['resp_bytes'], axis=1))\
.groupby(['id.orig_h', 'id.resp_h', 'conn_state'])\
.agg({
'time': ['min', 'max', 'size'],
'id.resp_p': ['nunique'],
'uid': ['nunique'],
'duration': ['min', 'max', 'mean'],
'orig_bytes': ['min', 'max', 'sum', 'mean'],
'resp_bytes': ['min', 'max', 'sum', 'mean'],
'sum_bytes': ['min', 'max', 'sum', 'mean']
}).reset_index()
df_summary.columns = [' '.join(col).strip() for col in df_summary.columns.values]
df_summary = df_summary\
.rename(columns={'time size': 'count'})\
.assign(
conn_state_uid=df_summary.apply(lambda row: row['id.orig_h'] + '_' + row['id.resp_h'] + '_' + row['conn_state'], axis=1))
print ('# rows', len(df_summary))
df_summary.sample(3)
```
## 3. Visualize data
* Nodes:
* IPs
* Bigger when more sessions (split by connection state) involving them
* Edges:
* src_ip -> dest_ip, split by connection state
```
hg = graphistry.hypergraph(
df_summary,
['id.orig_h', 'id.resp_h'],
direct=True,
opts={
'CATEGORIES': {
'ip': ['id.orig_h', 'id.resp_h']
}
})
hg['graph'].plot()
```
## Next Steps
* Part I: [CPU Baseline in Python Pandas](./part_i_cpu_pandas.ipynb)
* Part II: [GPU Dataframe with RAPIDS Python cudf bindings](./part_ii_gpu_cudf.ipynb)
| github_jupyter |
     
     
     
     
     
  
[Home Page](Start_Here.ipynb)
[Previous Notebook](Introduction_to_Performance_analysis.ipynb)
     
     
     
     
[1](Introduction_to_Performance_analysis.ipynb)
[2](Performance_Analysis_using_NSight_systems.ipynb)
[3]
     
     
     
     
In the previous notebooks, we have optimized the Multi-stream version of DeepStream Test App 2.In this notebook, we will work on a different pipeline to optimize it further.
- [Case 2:COVID-19 Social Distancing Application.](#Case-2:-COVID-19-Social-Distancing-Application.)
- [Finding distance between 2 people](#Finding-distance-between-2-people)
- [Solving the computational bottleneck](#Solving-the-computational-bottleneck)
- [Jetson specific optimizations](#Jetson-specific-optimizations)
- [Summary](#Summary)
#### Case 2: COVID-19 Social Distancing Application.
The COVID-19 Social Distance application can be constructed from `deepstream-test-app1` by adding suitable Metadata processing to determine whether two people have come in close contact and violated the social distancing norms.

##### Finding distance between 2 people
As we view people from a camera, it is necessary to have a perspective correction as far people would look smaller and appear less distant in pixel space. So, it's important to approximate world-space distance between persons.
We **assume** that avg human is of height 170 cm. We normalize bounding box height with this value. This normalized value is then further used to normalize pixel-space distance between objects.
We define the distance between two persons given their BBOX centroid(x,y) and BBOX height (h) as follows.
```python
# Pixel distance
dx = x2 - x1;
dy = y2 - y1;
# Pixel to real-world conversion using avg height as 170cm
lx = dx * 170 * (1/h1 + 1/h2) / 2;
ly = dy * 170 * (1/h1 + 1/h2) / 2;
l = sqrt(lx*lx + ly*ly);
```
Limitations: Above method tries to approximate 3d distance between persons from a 2d camera. As expected, this has limitations and works best only if the persons' body is perpendicular to the camera. These limitations can be removed if we use multiple cameras and camera calibration data to approximate persons' 3d location.
Let us now start building our pipeline considering this assumtion.
```
# Import Required Libraries
import sys
sys.path.append('../source_code')
import gi
import time
gi.require_version('Gst', '1.0')
from gi.repository import GObject, Gst , GLib
from common.bus_call import bus_call
import pyds
import math
# Defining the Class Labels
PGIE_CLASS_ID_VEHICLE = 0
PGIE_CLASS_ID_BICYCLE = 1
PGIE_CLASS_ID_PERSON = 2
PGIE_CLASS_ID_ROADSIGN = 3
# Defining the input output video file
INPUT_VIDEO_NAME = 'file:///opt/nvidia/deepstream/deepstream-5.0/python/source_code/dataset/wt.mp4'
OUTPUT_VIDEO_NAME = "../source_code/N3/ds_out.mp4"
def cb_newpad(decodebin, decoder_src_pad,data):
print("In cb_newpad\n")
caps=decoder_src_pad.get_current_caps()
gststruct=caps.get_structure(0)
gstname=gststruct.get_name()
source_bin=data
features=caps.get_features(0)
# Need to check if the pad created by the decodebin is for video and not
# audio.
print("gstname=",gstname)
if(gstname.find("video")!=-1):
# Link the decodebin pad only if decodebin has picked nvidia
# decoder plugin nvdec_*. We do this by checking if the pad caps contain
# NVMM memory features.
print("features=",features)
if features.contains("memory:NVMM"):
# Get the source bin ghost pad
bin_ghost_pad=source_bin.get_static_pad("src")
if not bin_ghost_pad.set_target(decoder_src_pad):
sys.stderr.write("Failed to link decoder src pad to source bin ghost pad\n")
else:
sys.stderr.write(" Error: Decodebin did not pick nvidia decoder plugin.\n")
def decodebin_child_added(child_proxy,Object,name,user_data):
print("Decodebin child added:", name, "\n")
if(name.find("decodebin") != -1):
Object.connect("child-added",decodebin_child_added,user_data)
def create_source_bin(index,uri):
print("Creating source bin")
# Create a source GstBin to abstract this bin's content from the rest of the
# pipeline
bin_name="source-bin-%02d" %index
print(bin_name)
nbin=Gst.Bin.new(bin_name)
if not nbin:
sys.stderr.write(" Unable to create source bin \n")
# Source element for reading from the uri.
# We will use decodebin and let it figure out the container format of the
# stream and the codec and plug the appropriate demux and decode plugins.
uri_decode_bin=Gst.ElementFactory.make("uridecodebin", "uri-decode-bin")
if not uri_decode_bin:
sys.stderr.write(" Unable to create uri decode bin \n")
# We set the input uri to the source element
uri_decode_bin.set_property("uri",uri)
# Connect to the "pad-added" signal of the decodebin which generates a
# callback once a new pad for raw data has beed created by the decodebin
uri_decode_bin.connect("pad-added",cb_newpad,nbin)
uri_decode_bin.connect("child-added",decodebin_child_added,nbin)
# We need to create a ghost pad for the source bin which will act as a proxy
# for the video decoder src pad. The ghost pad will not have a target right
# now. Once the decode bin creates the video decoder and generates the
# cb_newpad callback, we will set the ghost pad target to the video decoder
# src pad.
Gst.Bin.add(nbin,uri_decode_bin)
bin_pad=nbin.add_pad(Gst.GhostPad.new_no_target("src",Gst.PadDirection.SRC))
if not bin_pad:
sys.stderr.write(" Failed to add ghost pad in source bin \n")
return None
return nbin
## Make Element or Print Error and any other detail
def make_elm_or_print_err(factoryname, name, printedname, detail=""):
print("Creating", printedname)
elm = Gst.ElementFactory.make(factoryname, name)
if not elm:
sys.stderr.write("Unable to create " + printedname + " \n")
if detail:
sys.stderr.write(detail)
return elm
# Standard GStreamer initialization
Gst.init(None)
# Create gstreamer elements
# Create Pipeline element that will form a connection of other elements
print("Creating Pipeline \n ")
pipeline = Gst.Pipeline()
if not pipeline:
sys.stderr.write(" Unable to create Pipeline \n")
########### Create Elements required for the Pipeline ###########
# Create nvstreammux instance to form batches from one or more sources.
streammux = make_elm_or_print_err("nvstreammux", "Stream-muxer","Stream-muxer")
pipeline.add(streammux)
num_sources = 1
for i in range(num_sources):
print("Creating source_bin ",i," \n ")
uri_name=INPUT_VIDEO_NAME
if uri_name.find("rtsp://") == 0 :
is_live = True
source_bin=create_source_bin(i, uri_name)
if not source_bin:
sys.stderr.write("Unable to create source bin \n")
pipeline.add(source_bin)
padname="sink_%u" %i
sinkpad = streammux.get_request_pad(padname)
if not sinkpad:
sys.stderr.write("Unable to create sink pad bin \n")
srcpad = source_bin.get_static_pad("src")
if not srcpad:
sys.stderr.write("Unable to create src pad bin \n")
srcpad.link(sinkpad)
# Use nvinfer to run inferencing on decoder's output, behaviour of inferencing is set through config file
pgie = make_elm_or_print_err("nvinfer", "primary-inference" ,"pgie")
# Use convertor to convert from NV12 to RGBA as required by nvosd
nvvidconv = make_elm_or_print_err("nvvideoconvert", "convertor","nvvidconv")
# Create OSD to draw on the converted RGBA buffer
nvosd = make_elm_or_print_err("nvdsosd", "onscreendisplay","nvosd")
# Finally encode and save the osd output
queue = make_elm_or_print_err("queue", "queue", "Queue")
# Use convertor to convert from NV12 to RGBA as required by nvosd
nvvidconv2 = make_elm_or_print_err("nvvideoconvert", "convertor2","nvvidconv2")
# Place an encoder instead of OSD to save as video file
encoder = make_elm_or_print_err("avenc_mpeg4", "encoder", "Encoder")
# Parse output from Encoder
codeparser = make_elm_or_print_err("mpeg4videoparse", "mpeg4-parser", 'Code Parser')
# Create a container
container = make_elm_or_print_err("qtmux", "qtmux", "Container")
# Create Sink for storing the output
sink = make_elm_or_print_err("filesink", "filesink", "Sink")
############ Set properties for the Elements ############
print("Playing file ",INPUT_VIDEO_NAME)
# Set Input Width , Height and Batch Size
streammux.set_property('width', 1920)
streammux.set_property('height', 1080)
streammux.set_property('batch-size', 1)
# Timeout in microseconds to wait after the first buffer is available
# to push the batch even if a complete batch is not formed.
streammux.set_property('batched-push-timeout', 4000000)
# Set Congifuration file for nvinfer
pgie.set_property('config-file-path', "../source_code/N3/dstest1_pgie_config.txt")
# Set Encoder bitrate for output video
encoder.set_property("bitrate", 2000000)
# Set Output file name and disable sync and async
sink.set_property("location", OUTPUT_VIDEO_NAME)
sink.set_property("sync", 0)
sink.set_property("async", 0)
########## Add and Link ELements in the Pipeline ##########
print("Adding elements to Pipeline \n")
pipeline.add(pgie)
pipeline.add(nvvidconv)
pipeline.add(nvosd)
pipeline.add(queue)
pipeline.add(nvvidconv2)
pipeline.add(encoder)
pipeline.add(codeparser)
pipeline.add(container)
pipeline.add(sink)
# Linking elements to the Pipeline
print("Linking elements to Pipeline \n")
streammux.link(pgie)
pgie.link(nvvidconv)
nvvidconv.link(nvosd)
nvosd.link(queue)
queue.link(nvvidconv2)
nvvidconv2.link(encoder)
encoder.link(codeparser)
codeparser.link(container)
container.link(sink)
# create an event loop and feed gstreamer bus mesages to it
loop = GLib.MainLoop()
bus = pipeline.get_bus()
bus.add_signal_watch()
bus.connect ("message", bus_call, loop)
print("Created event loop")
############# Define Computation required for our pipeline #################
def compute_dist(p1, p2):
(x1, y1, h1) = p1;
(x2, y2, h2) = p2;
dx = x2 - x1;
dy = y2 - y1;
lx = dx * 170 * (1/h1 + 1/h2) / 2;
ly = dy * 170 * (1/h1 + 1/h2) / 2;
l = math.sqrt(lx*lx + ly*ly);
return l
def get_min_distances(centroids):
mini=[]
for i in range(len(centroids)):
distance=[]
for j in range(len(centroids)):
distance.append(compute_dist(centroids[i],centroids[j]))
distance[i]=10000000
mini.append(min(distance))
return mini
def visualize(objs):
violations = 0
dist_threshold = 160 # Distance in cms
for obj in objs:
min_dist = obj["min_dist"]
redness_factor = 1.5
r_channel = max(255 * (dist_threshold - min_dist) / dist_threshold, 0) * redness_factor
g_channel = 255 - r_channel
b_channel = 0
obj_meta = obj["obj_meta"]
obj_meta.rect_params.border_color.red = r_channel
obj_meta.rect_params.border_color.green = g_channel
obj_meta.rect_params.border_color.blue = b_channel
obj["violated"] = (min_dist < dist_threshold)
violations = violations + int(min_dist < dist_threshold)
return violations
def get_centroid(rect):
xmin = rect.left
xmax = rect.left + rect.width
ymin = rect.top
ymax = rect.top + rect.height
centroid_x = (xmin + xmax) / 2
centroid_y = (ymin + ymax) / 2
return (centroid_x, centroid_y, rect.height)
def compute_min_distances_cpp(objs):
centroids = [o["centroid"] for o in objs]
min_distances = get_min_distances(centroids)
for o in range(len(objs)):
objs[o]["min_dist"] = min_distances[o]
############## Working with the Metadata ################
def osd_sink_pad_buffer_probe(pad,info,u_data):
#Intiallizing object counter with 0.
obj_counter = {
PGIE_CLASS_ID_VEHICLE:0,
PGIE_CLASS_ID_PERSON:0,
PGIE_CLASS_ID_BICYCLE:0,
PGIE_CLASS_ID_ROADSIGN:0
}
# Set frame_number & rectangles to draw as 0
frame_number=0
num_rects=0
gst_buffer = info.get_buffer()
if not gst_buffer:
print("Unable to get GstBuffer ")
return
# Retrieve batch metadata from the gst_buffer
# Note that pyds.gst_buffer_get_nvds_batch_meta() expects the
# C address of gst_buffer as input, which is obtained with hash(gst_buffer)
batch_meta = pyds.gst_buffer_get_nvds_batch_meta(hash(gst_buffer))
l_frame = batch_meta.frame_meta_list
while l_frame is not None:
try:
# Note that l_frame.data needs a cast to pyds.NvDsFrameMeta
frame_meta = pyds.NvDsFrameMeta.cast(l_frame.data)
except StopIteration:
break
objects=[]
# Get frame number , number of rectables to draw and object metadata
frame_number=frame_meta.frame_num
num_rects = frame_meta.num_obj_meta
l_obj=frame_meta.obj_meta_list
while l_obj is not None:
try:
# Casting l_obj.data to pyds.NvDsObjectMeta
obj_meta=pyds.NvDsObjectMeta.cast(l_obj.data)
except StopIteration:
break
# Increment Object class by 1 and Set Box border to Red color
obj_counter[obj_meta.class_id] +=1
obj_meta.rect_params.border_color.set(0.0, 0.0, 1.0, 0.0)
if (obj_meta.class_id == PGIE_CLASS_ID_PERSON):
obj = {}
obj["tracker_id"] = obj_meta.object_id
obj["unique_id"] = obj_meta.unique_component_id
obj["centroid"] = get_centroid(obj_meta.rect_params)
obj["obj_meta"] = obj_meta
objects.append(obj)
else:
obj_meta.rect_params.border_width = 0
try:
l_obj=l_obj.next
except StopIteration:
break
# Get the number of violations
compute_min_distances_cpp(objects)
violations = visualize(objects)
################## Setting Metadata Display configruation ###############
# Acquiring a display meta object.
display_meta=pyds.nvds_acquire_display_meta_from_pool(batch_meta)
display_meta.num_labels = 1
py_nvosd_text_params = display_meta.text_params[0]
# Setting display text to be shown on screen
py_nvosd_text_params.display_text = "Frame Number={} Number of Objects={} Vehicle_count={} Person_count={} Violations={}".format(frame_number, num_rects, obj_counter[PGIE_CLASS_ID_VEHICLE], obj_counter[PGIE_CLASS_ID_PERSON],violations)
# Now set the offsets where the string should appear
py_nvosd_text_params.x_offset = 10
py_nvosd_text_params.y_offset = 12
# Font , font-color and font-size
py_nvosd_text_params.font_params.font_name = "Serif"
py_nvosd_text_params.font_params.font_size = 10
# Set(red, green, blue, alpha); Set to White
py_nvosd_text_params.font_params.font_color.set(1.0, 1.0, 1.0, 1.0)
# Text background color
py_nvosd_text_params.set_bg_clr = 1
# Set(red, green, blue, alpha); set to Black
py_nvosd_text_params.text_bg_clr.set(0.0, 0.0, 0.0, 1.0)
# Using pyds.get_string() to get display_text as string to print in notebook
print(pyds.get_string(py_nvosd_text_params.display_text))
pyds.nvds_add_display_meta_to_frame(frame_meta, display_meta)
############################################################################
try:
l_frame=l_frame.next
except StopIteration:
break
return Gst.PadProbeReturn.OK
# Lets add probe to get informed of the meta data generated, we add probe to the sink pad
# of the osd element, since by that time, the buffer would have had got all the metadata.
osdsinkpad = nvosd.get_static_pad("sink")
if not osdsinkpad:
sys.stderr.write(" Unable to get sink pad of nvosd \n")
osdsinkpad.add_probe(Gst.PadProbeType.BUFFER, osd_sink_pad_buffer_probe, 0)
print("Probe added")
# start play back and listen to events
print("Starting pipeline \n")
start_time = time.time()
pipeline.set_state(Gst.State.PLAYING)
try:
loop.run()
except:
pass
# cleanup
pipeline.set_state(Gst.State.NULL)
print("--- %s seconds ---" % (time.time() - start_time))
# Convert video profile to be compatible with Jupyter notebook
!ffmpeg -loglevel panic -y -an -i ../source_code/N3/ds_out.mp4 -vcodec libx264 -pix_fmt yuv420p -profile:v baseline -level 3 ../source_code/N3/output.mp4
# Display the Output
from IPython.display import HTML
HTML("""
<video width="640" height="480" controls>
<source src="../source_code/N3/output.mp4"
</video>
""".format())
```
Let us now run multiple streams concurrently and benchmark the performance we obtain from this.
```
!python3 ../source_code/utils/deepstream-covid-19.py --num-sources 32
!nsys profile --force-overwrite true -o ../source_code/reports/report4 python3 ../source_code/utils/deepstream-covid-19.py --num-sources 32 --prof True
```
[Click Here](../source_code/reports/report4.qdrep) to download the report file.

#### Solving the computational bottleneck
Here we can notice that the bottleneck is now shifted to the NV Decode. For Hardware capable of decoding multiple inputs concurrently, such as the A100, NV Decode would not be a bottleneck. We can see that the `queue3`, which works on the computation of the distance between people, would become the bottleneck as it takes a long time to execute. (In this case ~48 ms),in such cases, we can reduce the computation time to C++ or CUDA to make the computation faster. Here is one such example where we use C++ to run the distancing algorithm.
```
! cd ../source_code/distancing && cmake . && make
!python3 ../source_code/utils/deepstream-covid-19-cpp.py --num-sources 32
!nsys profile --force-overwrite true -o ../source_code/reports/report5 python3 ../source_code/utils/deepstream-covid-19-cpp.py --num-sources 32 --prof True
```
[Click Here](../source_code/reports/report5.qdrep) to download the report file.

Here we can notice that there has been a reduction in computation time when we shift it from Python to C++ (~48 ms to ~32ms), this can further be optimized in this case and even can be extended to CUDA if necessary.
### Jetson specific optimizations
#### Power
For Jetson devices, it is recommended to be set the mode to use Max power.
The Max power mode can be enabled using the following command
```bash
$ sudo nvpmodel -m 0
```
The GPU clocks can be stepped to maximum using the following command
```bash
$ sudo jetson_clocks
```
For information about supported power modes, see "Supported Modes and Power Efficiency" in the power management topics of NVIDIA Tegra Linux Driver Package Development Guide, e.g., "Power Management for Jetson AGX Xavier Devices."
For Jetson devices, the details regarding the memory and compute usage can be queried using the following command.
```bash
$ tegrastats
```
This command cannot be run inside the Docker container and needs to be run in a separate terminal.
#### Deep Learning Accelerators
Jetson AGX Xavier and Jetson NX supports 2 DLA engines. DeepStream does support inferencing using GPU and DLAs in parallel. You can do this in separate processes or a single process. You will need three separate sets of configs, configured to run on GPU, DLA0, and DLA1.
More details on this can be found [here](https://docs.nvidia.com/metropolis/deepstream/dev-guide/text/DS_Performance.html#running-applications-using-dla)
### Summary
In this notebook we learnt some techniques to optimize the Deepstream application and deal with computational bottlenecks that a may user can encounter and discussed one such way of solving them.
## Licensing
This material is released by NVIDIA Corporation under the Creative Commons Attribution 4.0 International (CC BY 4.0).
[Previous Notebook](Introduction_to_Performance_analysis.ipynb)
     
     
     
     
[1](Introduction_to_Performance_analysis.ipynb)
[2](Performance_Analysis_using_NSight_systems.ipynb)
[3]
     
     
     
     
     
     
     
     
     
  
[Home Page](Start_Here.ipynb)
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
# Standard
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from scipy import signal
from PIL import Image
import scipy
import os
import cv2
# # Tensorflow and Keras
# from keras.datasets import mnist
# from keras.models import Sequential, Model, Input
# from keras.layers import Dense, Dropout, Activation
# from keras.layers import Conv2D, MaxPooling2D, Flatten
# from tensorflow.keras.optimizers import SGD
# from keras.regularizers import l2
# import tensorflow as tf
# Xception because other model not working
from tensorflow.keras.applications.xception import preprocess_input
from tensorflow.keras.applications import Xception, MobileNetV2, VGG16, VGG19
from tensorflow.keras.preprocessing.image import img_to_array, load_img
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import SGD, RMSprop
from keras_preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
os.chdir('../')
from src import image_manip
from src import my_models as models
```
## Start with histology data
for each 40X, 100X, 200X, 400X, have 644 items for benign and 1300 for malignant
Augment our benign data since we only have ~650 images. This might not be necessary.
```
files = [f for f in os.listdir('data/Histology/100X/benign')]
for f in files:
img = Image.open(os.path.join('data/Histology/100X/benign',f))
x = ip.reshape_image(img)
ip.create_new_images(x)
root_dir = '/home/maureen/Documents/Galvanize/Capstone1/Capstone3/Cancer_Prediction'
```
## Make the model using simplecnn.py
```
# Assuming input shape = (width, height, channels) NOT (rows, cols, channels)
model = simple_cnn.create_model(input_size=(500,328,3), n_categories=2)
train_path = 'data/Histology/100X/train'
val_path = 'data/Histology/100X/validation'
test_path = 'data/Histology/100X/test'
# Data Generators
training_datagen = ImageDataGenerator(preprocessing_function=preprocess_input,
horizontal_flip=True)
validation_datagen = ImageDataGenerator(preprocessing_function=preprocess_input)
test_datagen = ImageDataGenerator(preprocessing_function=preprocess_input)
# Data generators from directory
train_generator = training_datagen.flow_from_directory(train_path,
target_size=(500, 328),
batch_size=16)
validation_generator = validation_datagen.flow_from_directory(val_path,
target_size=(500,328),
batch_size=16)
test_generator = test_datagen.flow_from_directory(test_path,
target_size=(500,328),
batch_size=16)
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
from tensorflow.keras.callbacks import TensorBoard
%load_ext tensorboard
callback = keras.callbacks.ModelCheckpoint(filepath='models/simplecnn_whc.h5', save_best_only=True)
tensorboard_callback = keras.callbacks.TensorBoard(log_dir="logs")
model.fit_generator(train_generator,
steps_per_epoch=1666//16,
epochs=3,
validation_data=validation_generator,
validation_steps=415//16,
callbacks=[callback, tensorboard_callback])
```
## Try Transfer Learning!
### Histology Data
```
# Run on Histology data
mag = ['40X', '100X', '200X', '400X']
predictions = []
filenames = []
for m in mag:
train_path = f'data/Histology/{m}/train'
val_path = f'data/Histology/{m}/validation'
test_path = f'data/Histology/{m}/test'
model = models.Xception_model(500,368)
model.compile_model(train_path, val_path)
model.fit()
files, predict = model.predict(test_path)
filenames.append(files)
predictions.append(predict)
predictions = np.asarray(predictions)
predictions.shape
np.save('filenames.txt', filenames)
# with open('predictionsrmsprop.txt', 'w') as f:
# f.write('\n'.join(predictions))
filenames
```
### Mammograms
```
# Regular MLO mammograms
# Using 500,277, which is rows, cols gives better results (.65 val vs .35 val)
# Rescale effect on train and val generator -> val accuracy changes with no rescale
# Getting same val score for all epochs. Not sure why. Didn't have this last week.
train_path = f'data/Mammograms/Tensor2/train'
val_path = f'data/Mammograms/Tensor2/validation'
test_path = f'data/Mammograms/Tensor2/test'
model = models.Xception_model(700,420) # best with 500,277. this is rows, cols. Val accuracy the samefor all epochs
model.compile_model(train_path, val_path)
model.fit()
files, predict = model.predict(test_path)
y_pred = np.argmax(predict, axis=1)
y_true = []
for f in files:
if 'cancer' in f:
y_true.append(0)
else:
y_true.append(1)
y_proba0 = predict[:,0]
y_proba1 = predict[:,1]
data = pd.DataFrame({'Files': files, 'y_true': y_true, 'y_pred': y_pred, 'y_proba0':y_proba0, 'y_proba1': y_proba1})
plt.style.use('ggplot')
from src import model_analysis
fig, ax = plt.subplots()
model_analysis.plot_roc(ax, 'Mammograms', y_true, predict)
plt.savefig('nn_roc_cc.png', dpi=350)
model_analysis.make_confusion('nn_cc', y_true, class_label)
data.to_csv('nn_mam_cc.csv')
data['y_true'].sum(), data['y_pred'].sum()
# Sinograms
train_path = f'data/Mammograms/Tensor/train_sino'
val_path = f'data/Mammograms/Tensor/validation_sino/'
test_path = f'data/Mammograms/Tensor/test_sino/'
model = models.Xception_model(400,720)
model.compile_model(train_path, val_path)
model.fit()
files, predict = model.predict(test_path)
predict
```
## All the code for Xception Class
```
# Try new model because previous isn't working
# Using softmax so that we'll get probabilities for predictions
# Regular MLO mammograms
train_path = 'data/Mammograms/Tensor/train'
val_path = 'data/Mammograms/Tensor/validation'
test_path = 'data/Mammograms/Tensor/test'
width = 500
height = 277
# Data Generators
training_datagen = ImageDataGenerator(preprocessing_function=preprocess_input,
# rescale=1./255,
horizontal_flip=True)
validation_datagen = ImageDataGenerator(preprocessing_function=preprocess_input)
# rescale=1./255,)
test_datagen = ImageDataGenerator(preprocessing_function=preprocess_input)
# Data generators from directory
train_generator = training_datagen.flow_from_directory(train_path,
target_size=(width, height),
batch_size=16)
validation_generator = validation_datagen.flow_from_directory(val_path,
target_size=(width,height),
batch_size=16)
test_generator = test_datagen.flow_from_directory(test_path,
target_size=(width,height),
batch_size=16)
from tensorflow.keras.layers import Dense, GlobalAveragePooling2D, Flatten, Dropout
from tensorflow.keras.models import Model
def create_transfer_model(input_size, n_categories, weights = 'imagenet'):
# note that the "top" is not included in the weights below
base_model = Xception(weights=weights,
include_top=False,
input_shape=input_size)
model = base_model.output
model = GlobalAveragePooling2D()(model)
predictions = Dense(n_categories, activation='softmax')(model)
model = Model(inputs=base_model.input, outputs=predictions)
return model
model = create_transfer_model((width,height,3),2)
def print_model_properties(model, indices = 0):
for i, layer in enumerate(model.layers[indices:]):
print(i+indices, layer.name,layer.trainable)
print_model_properties(model)
```
Change head
```
def change_trainable_layers(model, trainable_index):
for layer in model.layers[:trainable_index]:
layer.trainable = False
for layer in model.layers[trainable_index:]:
layer.trainable = True
_ = change_trainable_layers(model, 132)
print_model_properties(model, 120)
# Compile model. Changed lr from 0.0005 to 0.005
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# Train model can we use fit generator?
model.fit(train_generator,
steps_per_epoch=915//16,
epochs=4,
validation_data=validation_generator,
validation_steps=227//16)
#model.save_weights('models/weights.h5')
#model.save('models/transfermodel.h5')
## Shap
import json
import shap
# SHAP doesn't play nice with tensorflow 2.0
import tensorflow
from tensorflow.compat.v1.keras.backend import get_session
tensorflow.compat.v1.disable_v2_behavior()
# explain how the input to the 7th layer of the model explains the top two classes
def map2layer(x, layer):
feed_dict = dict(zip([model.layers[0].input], [preprocess_input(x.copy())]))
return get_session().run(model.layers[layer].input, feed_dict)
model.layers[0].input
# Magic starts here. Layer number is important
# Get the image
train_images = 'data/Mammograms/Tensor/train/cancer/'
files = [f for f in os.listdir(train_images)]
lst = [cv2.imread(os.path.join(train_images,f)) for f in files]
# X is array of all the image inputs to train the model
X = np.asarray(lst)
to_explain = X[[39,41]]
print(X.shape)
l = 17
e = shap.GradientExplainer((model.layers[l].input, model.layers[-1].output),
map2layer(X, l),
local_smoothing=0 # std dev of smoothing noise
)
shap_values,indexes = e.shap_values(map2layer(to_explain, l), ranked_outputs=2)
# get the names for the classes
#index_names = np.vectorize(lambda x: class_names[str(x)][1])(indexes)
# plot the explanations
#shap.image_plot(shap_values, to_explain, index_names)
```
## Make predictions!
```
# Single image
#path = 'data/Histology/100X/test/test/SOB_B_TA-14-13200-100-001.png'
img = Image.open(img_path)
x = img_to_array(img)
images = np.array([x])
images.shape
model.predict(images)
# predict on entire folder
predictions = model.predict(test_generator)
predictions, test_generator.filenames
```
| github_jupyter |
```
%matplotlib notebook
import control as c
import ipywidgets as w
import numpy as np
from IPython.display import display, HTML
import matplotlib.pyplot as plt
import matplotlib.animation as animation
#display(HTML('<script> $(document).ready(function() { $(\"div.input\").hide(); }); </script>'))
# Toggle cell visibility
from IPython.display import HTML
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide()
} else {
$('div.input').show()
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
Promijeni vidljivost <a href="javascript:code_toggle()">ovdje</a>.''')
display(tag)
```
## Stvaranje PI-regulatora korištenjem operacijskih pojačala
U analognoj elektronici, operacijska pojačala uobičajeno se koriste za realizaciju proporcionalno-integracijski-derivacijskih (PID) regulatora. Dok matematički modeli linearnih vremenski-nepromjenjivih (LTI) sustava pretpostavljaju idealne uvjete, realni sklopovi možda im ne odgovaraju u potpunosti.
U većini slučajeva idealni model daje prihvatljive rezultate, ali frekvencijske karakteristike mogu se bolje aproksimirati proširivanjem modela s pojačanjem otvorene petlje:
<br><br>
$$G_{ideal}(s)=\frac{V_{out}}{V_{in}}=-\frac{Z_F}{Z_G}\qquad\qquad G_{approx}(s)=\frac{V_{out}}{V_{in}}=-\frac{\frac{-A\cdot Z_F}{Z_G+Z_F}}{1+\frac{A\cdot Z_G}{Z_G+Z_F}}$$
<br>
U ovom ćemo primjeru istražiti neke od konfiguracija PI regulatora zasnovanih na operacijskim pojačalima.<br>
<b>Prvo, odaberite vrijednost pojačanja otvorene petlje za prikazane izračune!</b>
```
# Model selector
opampGain = w.ToggleButtons(
options=[('10 000', 10000), ('50 000', 50000), ('200 000', 200000),],
description='Pojačanje operacijskog pojačala: ', style={'description_width':'30%'})
display(opampGain)
```
PI regulator može se implementirati pomoću otpornika u unaprijednoj vezi i kondenzatora u povratnoj vezi. Idealni model točno odgovara matematičkom obliku regulatora. Ali, nakon uključivanja pojačanja otvorene petlje, integrator se zamjenjuje sustavom prvog reda s ogromnom vremenskom konstantom, ograničavajući na taj način amplitudu na niskim frekvencijama.
<br><br>
<img src="Images/int1.png" width="30%" />
<br>
<b>Prilagodite pasivne komponente tako da neidealni model bude najbliži idealnom! Gdje karakteristike značajno odstupaju od idealnih? Što se može reći o grafu faze?</b>
```
# Figure definition
fig1, ((f1_ax1), (f1_ax2)) = plt.subplots(2, 1)
fig1.set_size_inches((9.8, 5))
fig1.set_tight_layout(True)
l1 = f1_ax1.plot([], [], color='red')
l2 = f1_ax2.plot([], [], color='red')
l3 = f1_ax1.plot([], [], color='blue')
l4 = f1_ax2.plot([], [], color='blue')
f1_line1 = l1[0]
f1_line2 = l2[0]
f1_line3 = l3[0]
f1_line4 = l4[0]
f1_ax1.legend(l1+l3, ['Ne-idealno', 'Idealno'], loc=1)
f1_ax2.legend(l2+l4, ['Ne-idealno', 'Idealno'], loc=1)
f1_ax1.grid(which='both', axis='both', color='lightgray')
f1_ax2.grid(which='both', axis='both', color='lightgray')
f1_ax1.autoscale(enable=True, axis='x', tight=True)
f1_ax2.autoscale(enable=True, axis='x', tight=True)
f1_ax1.autoscale(enable=True, axis='y', tight=False)
f1_ax2.autoscale(enable=True, axis='y', tight=False)
f1_ax1.set_title('Bodeov graf amplitude', fontsize=11)
f1_ax1.set_xscale('log')
f1_ax1.set_xlabel(r'$f\/$[Hz]', labelpad=0, fontsize=10)
f1_ax1.set_ylabel(r'$A\/$[dB]', labelpad=0, fontsize=10)
f1_ax1.tick_params(axis='both', which='both', pad=0, labelsize=8)
f1_ax2.set_title('Bodeov graf faze', fontsize=11)
f1_ax2.set_xscale('log')
f1_ax2.set_xlabel(r'$f\/$[Hz]', labelpad=0, fontsize=10)
f1_ax2.set_ylabel(r'$\phi\/$[°]', labelpad=0, fontsize=10)
f1_ax2.tick_params(axis='both', which='both', pad=0, labelsize=8)
# System model
def system_model(rg, cf, a):
Rg = rg / 1000 # Convert to Ohm
Cf = cf * 1000000 # Convert to Farad
W_ideal = c.tf([-1], [Rg*Cf, 0])
W_ac = c.tf([-a], [Cf*Rg*(1+a), 1])
global f1_line1, f1_line2, f1_line3, f1_line4
f1_ax1.lines.remove(f1_line1)
f1_ax2.lines.remove(f1_line2)
f1_ax1.lines.remove(f1_line3)
f1_ax2.lines.remove(f1_line4)
mag, phase, omega = c.bode_plot(W_ac, Plot=False) # Non-ideal Bode-plot
f1_line1, = f1_ax1.plot(omega/2/np.pi, 20*np.log10(mag), lw=1, color='red')
f1_line2, = f1_ax2.plot(omega/2/np.pi, phase*180/np.pi, lw=1, color='red')
mag, phase, omega = c.bode_plot(W_ideal, omega=omega, Plot=False) # Ideal Bode-plot at the non-ideal points
f1_line3, = f1_ax1.plot(omega/2/np.pi, 20*np.log10(mag), lw=1, color='blue')
f1_line4, = f1_ax2.plot(omega/2/np.pi, phase*180/np.pi, lw=1, color='blue')
f1_ax1.relim()
f1_ax2.relim()
f1_ax1.autoscale_view()
f1_ax2.autoscale_view()
print('Prijenosna funkcija za idealni PI:')
print(W_ideal)
print('\nPrijenosna funkcija za ne-idealni PI:')
print(W_ac)
# GUI widgets
rg_slider = w.FloatLogSlider(value=1, base=10, min=-3, max=3, description=r'$R_g\ [k\Omega]\ :$', continuous_update=False,
layout=w.Layout(width='75%'), style={'description_width':'30%'})
cf_slider = w.FloatLogSlider(value=1, base=10, min=-3, max=3, description=r'$C_f\ [\mu H]\ :$', continuous_update=False,
layout=w.Layout(width='75%'), style={'description_width':'30%'})
input_data = w.interactive_output(system_model, {'rg':rg_slider, 'cf':cf_slider, 'a':opampGain})
display(w.HBox([rg_slider, cf_slider]), input_data)
```
Ovaj PI regulator može biti jednostavan, ali nije moguće kontrolirati DC pojačanje putem pasivnih komponenata. Zbog toga se obično u povratnu vezu paralelno spaja odgovarajući otpornik.
<br><br>
<img src="Images/int2.png" width="30%" />
<br>
<b>Prilagodite pasivne komponente tako da neidealni model bude najbliži idealnom! Koje su razlike s obzirom na prethodni model?</b>
```
# Filtered PI - parallel
fig2, ((f2_ax1), (f2_ax2)) = plt.subplots(2, 1)
fig2.set_size_inches((9.8, 5))
fig2.set_tight_layout(True)
l1 = f2_ax1.plot([], [], color='red')
l2 = f2_ax2.plot([], [], color='red')
l3 = f2_ax1.plot([], [], color='blue')
l4 = f2_ax2.plot([], [], color='blue')
f2_line1 = l1[0]
f2_line2 = l2[0]
f2_line3 = l3[0]
f2_line4 = l4[0]
f2_ax1.legend(l1+l3, ['Ne-idealno', 'Idealno'], loc=1)
f2_ax2.legend(l2+l4, ['Ne-idealno', 'Idealno'], loc=1)
f2_ax1.grid(which='both', axis='both', color='lightgray')
f2_ax2.grid(which='both', axis='both', color='lightgray')
f2_ax1.autoscale(enable=True, axis='x', tight=True)
f2_ax2.autoscale(enable=True, axis='x', tight=True)
f2_ax1.autoscale(enable=True, axis='y', tight=False)
f2_ax2.autoscale(enable=True, axis='y', tight=False)
f2_ax1.set_title('Bodeov graf amplitude', fontsize=11)
f2_ax1.set_xscale('log')
f2_ax1.set_xlabel(r'$f\/$[Hz]', labelpad=0, fontsize=10)
f2_ax1.set_ylabel(r'$A\/$[dB]', labelpad=0, fontsize=10)
f2_ax1.tick_params(axis='both', which='both', pad=0, labelsize=8)
f2_ax2.set_title('Bodeov graf faze', fontsize=11)
f2_ax2.set_xscale('log')
f2_ax2.set_xlabel(r'$f\/$[Hz]', labelpad=0, fontsize=10)
f2_ax2.set_ylabel(r'$\phi\/$[°]', labelpad=0, fontsize=10)
f2_ax2.tick_params(axis='both', which='both', pad=0, labelsize=8)
# System model
def system2_model(rg, rf, cf, a):
Rg = rg / 1000 # Convert to Ohm
Rf = rf / 1000
Cf = cf * 1000000 # Convert to Farad
W_ideal = c.tf([-1], [Rg*Cf, Rg/Rf])
W_ac = c.tf([-a], [Cf*Rg*(a+1), Rg*(a+1)/Rf+1])
global f2_line1, f2_line2, f2_line3, f2_line4
f2_ax1.lines.remove(f2_line1)
f2_ax2.lines.remove(f2_line2)
f2_ax1.lines.remove(f2_line3)
f2_ax2.lines.remove(f2_line4)
mag, phase, omega = c.bode_plot(W_ac, Plot=False) # Non-ideal Bode-plot
f2_line1, = f2_ax1.plot(omega/2/np.pi, 20*np.log10(mag), lw=1, color='red')
f2_line2, = f2_ax2.plot(omega/2/np.pi, phase*180/np.pi, lw=1, color='red')
mag, phase, omega = c.bode_plot(W_ideal, omega=omega, Plot=False) # Ideal Bode-plot at the non-ideal points
f2_line3, = f2_ax1.plot(omega/2/np.pi, 20*np.log10(mag), lw=1, color='blue')
f2_line4, = f2_ax2.plot(omega/2/np.pi, phase*180/np.pi, lw=1, color='blue')
f2_ax1.relim()
f2_ax2.relim()
f2_ax1.autoscale_view()
f2_ax2.autoscale_view()
print('Prijenosna funckija za idealni filtrirani PI:')
print(W_ideal)
print('\nPrijenosna funckija za ne-idealni filtrirani PI::')
print(W_ac)
# GUI widgets
rg2_slider = w.FloatLogSlider(value=1, base=10, min=-3, max=3, description=r'$R_g$ [k$\Omega$]', continuous_update=False,
layout=w.Layout(width='75%'), style={'description_width':'30%'})
rf2_slider = w.FloatLogSlider(value=1, base=10, min=-3, max=3, description=r'$R_f$ [k$\Omega$]', continuous_update=False,
layout=w.Layout(width='75%'), style={'description_width':'30%'})
cf2_slider = w.FloatLogSlider(value=1, base=10, min=-3, max=3, description=r'$C_f$ [$\mu$H]', continuous_update=False,
layout=w.Layout(width='75%'), style={'description_width':'30%'})
input_data = w.interactive_output(system2_model, {'rg':rg2_slider, 'rf':rf2_slider, 'cf':cf2_slider, 'a':opampGain})
display(w.HBox([rg2_slider, rf2_slider, cf2_slider]), input_data)
```
| github_jupyter |
## Instructions
0. If you haven't already, follow [the setup instructions here](https://jennselby.github.io/MachineLearningCourseNotes/#setting-up-python3) to get all necessary software installed.
0. Install the Gensim word2vec Python implementation: `python3 -m pip install --upgrade gensim`
0. Get the trained model (1billion_word_vectors.zip) from Canvas and put it in the same folder as this ipynb file.
0. Unzip the trained model file. You should now have three files in the folder (if zip created a new folder, move these files out of that separate folder into the same folder as this ipynb file):
* 1billion_word_vectors
* 1billion_word_vectors.syn1neg.npy
* 1billion_word_vectors.wv.syn0.npy
0. Read through the code in the following sections:
* [Load trained word vectors](#Load-Trained-Word-Vectors)
* [Explore word vectors](#Explore-Word-Vectors)
0. Optionally, complete [Exercise: Explore Word Vectors](#Exercise:-Explore-Word-Vectors)
0. Read through the code in the following sections:
* [Use Word Vectors in an Embedding Layer of a Keras Model](#Use-Word-Vectors-in-an-Embedding-Layer-of-a-Keras-Model)
* [IMDB Dataset](#IMDB-Dataset)
* [Train IMDB Word Vectors](#Train-IMDB-Word-Vectors)
* [Process Dataset](#Process-Dataset)
* [Classification With Word Vectors Trained With Model](#Classification-With-Word-Vectors-Trained-With-Model)
0. Complete one of the two [Exercises](#Exercises). Remember to keep notes about what you do!
## Extra Details -- Do Not Do This
This took awhile, which is why I'm giving you the trained file rather than having you do this. But just in case you're curious, here is how to create the trained model file.
1. Download the corpus of sentences from [http://www.statmt.org/lm-benchmark/1-billion-word-language-modeling-benchmark-r13output.tar.gz](http://www.statmt.org/lm-benchmark/1-billion-word-language-modeling-benchmark-r13output.tar.gz)
1. Unzip and unarchive the file: `tar zxf 1-billion-word-language-modeling-benchmark-r13output.tar.gz`
1. Run the following Python code:
```
from gensim.models import word2vec
import os
corpus_dir = '1-billion-word-language-modeling-benchmark-r13output/training-monolingual.tokenized.shuffled'
sentences = word2vec.PathLineSentences(corpus_dir)
model = word2vec.Word2Vec(sentences) # just use all of the default settings for now
model.save('1billion_word_vectors')
```
## Documentation/Sources
* [https://radimrehurek.com/gensim/models/word2vec.html](https://radimrehurek.com/gensim/models/word2vec.html) for more information about how to use gensim word2vec in general
* _Blog post has been removed_ [https://codekansas.github.io/blog/2016/gensim.html](https://codekansas.github.io/blog/2016/gensim.html) for information about using it to create embedding layers for neural networks.
* [https://machinelearningmastery.com/sequence-classification-lstm-recurrent-neural-networks-python-keras/](https://machinelearningmastery.com/sequence-classification-lstm-recurrent-neural-networks-python-keras/) for information on sequence classification with keras
* [https://blog.keras.io/using-pre-trained-word-embeddings-in-a-keras-model.html](https://blog.keras.io/using-pre-trained-word-embeddings-in-a-keras-model.html) for using pre-trained embeddings with keras (though the syntax they use for the model layers is different than most other tutorials).
* [https://keras.io/](https://keras.io/) Keras API documentation
## Load Trained Word Vectors
```
from gensim.models import word2vec
```
Load the trained model file into memory
```
wv_model = word2vec.Word2Vec.load('1billion_word_vectors')
```
Since we do not need to continue training the model, we can save memory by keeping the parts we need (the word vectors themselves) and getting rid of the rest of the model.
```
wordvec = wv_model.wv
del wv_model
```
## Explore Word Vectors
Now we can look at some of the relationships between different words.
Like [the gensim documentation](https://radimrehurek.com/gensim/models/word2vec.html), let's start with a famous example: king + woman - man
```
wordvec.most_similar(positive=['king', 'woman'], negative=['man'])
```
This next one does not work as well as I'd hoped, but it gets close. Maybe you can find a better example.
```
wordvec.most_similar(positive=['panda', 'eucalyptus'], negative=['bamboo'])
```
Which one of these is not like the others?
Note: It looks like the gensim code needs to be updated to meet the requirements of later versions of numpy. You can ignore the warning.
```
wordvec.doesnt_match(['red', 'purple', 'laptop', 'turquoise', 'ruby'])
```
How far apart are different words?
```
wordvec.distances('laptop', ['computer', 'phone', 'rabbit'])
```
Let's see what one of these vectors actually looks like.
```
wordvec['textbook']
```
What other methods are available to us?
```
help(wordvec)
```
# Exercise: Explore Word Vectors
## Optional
What other interesting relationship can you find, using the methods used in the examples above or anything you find in the help message?
## Use Word Vectors in an Embedding Layer of a Keras Model
```
from keras.models import Sequential
import numpy
```
You may have noticed in the help text for wordvec that it has a built-in method for converting into a Keras embedding layer.
Since for this experimentation, we'll just be giving the embedding layer one word at a time, we can set the input length to 1.
```
test_embedding_layer = wordvec.get_keras_embedding()
test_embedding_layer.input_length = 1
embedding_model = Sequential()
embedding_model.add(test_embedding_layer)
```
But how do we actually use this? If you look at the [Keras Embedding Layer documentation](https://keras.io/layers/embeddings/) you might notice that it takes numerical input, not strings. How do we know which number corresponds to a particular word? In addition to having a vector, each word has an index:
```
wordvec.vocab['python'].index
```
Let's see if we get the same vector from the embedding layer as we get from our word vector object.
```
wordvec['python']
embedding_model.predict(numpy.array([[30438]]))
```
Looks good, right? But let's not waste our time when the computer could tell us definitively and quickly:
```
embedding_model.predict(numpy.array([[wordvec.vocab['python'].index]]))[0][0] == wordvec['python']
```
Now we have a way to turn words into word vectors with Keras layers. Yes! Time to get some data.
## IMDB Dataset
The [IMDB dataset](https://keras.io/datasets/#imdb-movie-reviews-sentiment-classification) consists of movie reviews that have been marked as positive or negative. (There is also a built-in dataset of [Reuters newswires](https://keras.io/datasets/#reuters-newswire-topics-classification) that have been classified by topic.)
```
from keras.datasets import imdb
(x_train, y_train), (x_test, y_test) = imdb.load_data()
```
It looks like our labels consist of 0 or 1, which makes sense for positive and negative.
```
print(y_train[0:9])
print(max(y_train))
print(min(y_train))
```
But x is a bit more trouble. The words have already been converted to numbers -- numbers that have nothing to do with the word embeddings we spent time learning!
```
x_train[0]
```
Looking at the help page for imdb, it appears there is a way to get the word back. Phew.
```
help(imdb)
imdb_offset = 3
imdb_map = dict((index + imdb_offset, word) for (word, index) in imdb.get_word_index().items())
imdb_map[0] = 'PADDING'
imdb_map[1] = 'START'
imdb_map[2] = 'UNKNOWN'
```
The knowledge about the initial indices and offset came from [this stack overflow post](https://stackoverflow.com/questions/42821330/restore-original-text-from-keras-s-imdb-dataset) after I got gibberish when I tried to translate the first review, below. It looks coherent now!
```
' '.join([imdb_map[word_index] for word_index in x_train[0]])
```
## Train IMDB Word Vectors
The word vectors from the 1 billion words dataset might work for us when trying to classify the IMDB data. Word vectors trained on the IMDB data itself might work better, though.
```
train_sentences = [['PADDING'] + [imdb_map[word_index] for word_index in review] for review in x_train]
test_sentences = [['PADDING'] + [imdb_map[word_index] for word_index in review] for review in x_test]
# min count says to put any word that appears at least once into the vocabulary
# size sets the dimension of the output vectors
imdb_wv_model = word2vec.Word2Vec(train_sentences + test_sentences + ['UNKNOWN'], min_count=1, size=100)
imdb_wordvec = imdb_wv_model.wv
del imdb_wv_model
```
## Process Dataset
For this exercise, we're going to keep all inputs the same length (we'll see how to do variable-length later). This means we need to choose a maximum length for the review, cutting off longer ones and adding padding to shorter ones. What should we make the length? Let's understand our data.
```
lengths = [len(review) for review in x_train + x_test]
print('Longest review: {} Shortest review: {}'.format(max(lengths), min(lengths)))
```
2697 words! Wow. Well, let's see how many reviews would get cut off at a particular cutoff.
```
cutoff = 500
print('{} reviews out of {} are over {}.'.format(
sum([1 for length in lengths if length > cutoff]),
len(lengths),
cutoff))
from keras.preprocessing import sequence
x_train_padded = sequence.pad_sequences(x_train, maxlen=cutoff)
x_test_padded = sequence.pad_sequences(x_test, maxlen=cutoff)
```
## Classification With Word Vectors Trained With Model
```
from keras.models import Sequential
from keras.layers import Embedding, Conv1D, Dense, Flatten
```
Model definition. The embedding layer here learns the 100-dimensional vector embedding within the overall classification problem training. That is usually what we want, unless we have a bunch of un-tagged data that could be used to train word vectors but not a classification model.
```
not_pretrained_model = Sequential()
not_pretrained_model.add(Embedding(input_dim=len(imdb_map), output_dim=100, input_length=cutoff))
not_pretrained_model.add(Conv1D(filters=32, kernel_size=5, activation='relu'))
not_pretrained_model.add(Conv1D(filters=32, kernel_size=5, activation='relu'))
not_pretrained_model.add(Flatten())
not_pretrained_model.add(Dense(units=128, activation='relu'))
not_pretrained_model.add(Dense(units=1, activation='sigmoid')) # because at the end, we want one yes/no answer
not_pretrained_model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['binary_accuracy'])
```
Train the model. __This takes awhile. You might not want to re-run it.__
```
not_pretrained_model.fit(x_train_padded, y_train, epochs=1, batch_size=64)
```
Assess the model. __This takes awhile. You might not want to re-run it.__
```
not_pretrained_scores = not_pretrained_model.evaluate(x_test_padded, y_test)
print('loss: {} accuracy: {}'.format(*not_pretrained_scores))
```
# Exercises
## These exercises will help you learn more about how to use word vectors in a model and how to translate between data representations.
## For any model that you try in these exercises, take notes about the performance you see and anything you notice about the differences between the models.
## Exercise Option #1 - Advanced Difficulty
Using the details above about how the imdb dataset and the keras embedding layer represent words, define a model that uses the pre-trained word vectors from the imdb dataset rather than an embedding that keras learns as it goes along. You'll need to replace the embedding layer and feed in different training data.
## Exercise Option #2 - Advanced Difficulty
Same as option 1, but try using the 1billion vector word embeddings instead of the imdb vectors. If you also did option 1, comment on how the performance changes.
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
```
Training and Testing Data
=====================================
To evaluate how well our supervised models generalize, we can split our data into a training and a test set:
<img src="figures/train_test_split_matrix.svg" width="100%">
```
from sklearn.datasets import load_iris
iris = load_iris()
X, y = iris.data, iris.target
```
Thinking about how machine learning is normally performed, the idea of a train/test split makes sense. Real world systems train on the data they have, and as other data comes in (from customers, sensors, or other sources) the classifier that was trained must predict on fundamentally *new* data. We can simulate this during training using a train/test split - the test data is a simulation of "future data" which will come into the system during production.
Specifically for iris, the 150 labels in iris are sorted, which means that if we split the data using a proportional split, this will result in fudamentally altered class distributions. For instance, if we'd perform a common 2/3 training data and 1/3 test data split, our training dataset will only consists of flower classes 0 and 1 (Setosa and Versicolor), and our test set will only contain samples with class label 2 (Virginica flowers).
Under the assumption that all samples are independent of each other (in contrast time series data), we want to **randomly shuffle the dataset before we split the dataset** as illustrated above.
```
y
```
Now we need to split the data into training and testing. Luckily, this is a common pattern in machine learning and scikit-learn has a pre-built function to split data into training and testing sets for you. Here, we use 50% of the data as training, and 50% testing. 80% and 20% is another common split, but there are no hard and fast rules. The most important thing is to fairly evaluate your system on data it *has not* seen during training!
```
from sklearn.model_selection import train_test_split
train_X, test_X, train_y, test_y = train_test_split(X, y,
train_size=0.5,
test_size=0.5,
random_state=123)
print("Labels for training data:")
print(train_y)
print("Labels for test data:")
print(test_y)
```
---
**Tip: Stratified Split**
Especially for relatively small datasets, it's better to stratify the split. Stratification means that we maintain the original class proportion of the dataset in the test and training sets. For example, after we randomly split the dataset as shown in the previous code example, we have the following class proportions in percent:
```
print('All:', np.bincount(y) / float(len(y)) * 100.0)
print('Training:', np.bincount(train_y) / float(len(train_y)) * 100.0)
print('Test:', np.bincount(test_y) / float(len(test_y)) * 100.0)
```
So, in order to stratify the split, we can pass the label array as an additional option to the `train_test_split` function:
```
train_X, test_X, train_y, test_y = train_test_split(X, y,
train_size=0.5,
test_size=0.5,
random_state=123,
stratify=y)
print('All:', np.bincount(y) / float(len(y)) * 100.0)
print('Training:', np.bincount(train_y) / float(len(train_y)) * 100.0)
print('Test:', np.bincount(test_y) / float(len(test_y)) * 100.0)
```
---
By evaluating our classifier performance on data that has been seen during training, we could get false confidence in the predictive power of our model. In the worst case, it may simply memorize the training samples but completely fails classifying new, similar samples -- we really don't want to put such a system into production!
Instead of using the same dataset for training and testing (this is called "resubstitution evaluation"), it is much much better to use a train/test split in order to estimate how well your trained model is doing on new data.
```
from sklearn.neighbors import KNeighborsClassifier
classifier = KNeighborsClassifier().fit(train_X, train_y)
pred_y = classifier.predict(test_X)
print("Fraction Correct [Accuracy]:")
print(np.sum(pred_y == test_y) / float(len(test_y)))
```
We can also visualize the correct predictions ...
```
print('Samples correctly classified:')
correct_idx = np.where(pred_y == test_y)[0]
print(correct_idx)
```
... as well as the failed predictions
```
print('Samples incorrectly classified:')
incorrect_idx = np.where(pred_y != test_y)[0]
print(incorrect_idx)
# Plot two dimensions
for n in np.unique(test_y):
idx = np.where(test_y == n)[0]
plt.scatter(test_X[idx, 1], test_X[idx, 2], label="Class %s" % str(iris.target_names[n]))
plt.scatter(test_X[incorrect_idx, 1], test_X[incorrect_idx, 2], color="darkred")
plt.xlabel('sepal width [cm]')
plt.ylabel('petal length [cm]')
plt.legend(loc=3)
plt.title("Iris Classification results")
plt.show()
```
We can see that the errors occur in the area where green (class 1) and orange (class 2) overlap. This gives us insight about what features to add - any feature which helps separate class 1 and class 2 should improve classifier performance.
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>
Print the true labels of 3 wrong predictions and modify the scatterplot code, which we used above, to visualize and distinguish these three samples with different markers in the 2D scatterplot. Can you explain why our classifier made these wrong predictions?
</li>
</ul>
</div>
```
# %load solutions/04_wrong-predictions.py
```
| github_jupyter |
# Mutivariate Regression Analysis
***
**Videos can be found at: https://www.youtube.com/channel/UCBsTB02yO0QGwtlfiv5m25Q**
In our previous tutorial, we explored the topic of Linear Regression Analysis which attempts to model the relationship between two variables by fitting a linear equation to the observed data. In this simple regression analysis, we have one explanatory variable and one dependent variable. However, what happens if we believe there is more than one explanatory variable that impacts the dependent variable? How would we model this?
Welcome to the world of multiple regression analysis. In this type of model, we attempt to model the relationship between multiple explanatory variables to a single dependent variable. While adding more variables allows us to model more complex phenomenons there are also additional steps we must take to make sure our model is sound and robust.
In this tutorial, we will be performing a multiple regression analysis on South Korea's GDP growth. South Korea in the 1950s came out of the Korean War, which left it's country ravaged and in extreme poverty. However, South Korea would go through one most significant economic developments the World has seen, taking it from a country in poverty to one of the top 15 economies in the World today.
Our goal is to be able to predict what the GDP growth rate will be in any year, given a few explanatory variables that we will define below.
***
## Assumptions of the Model
It's essential to understand the assumptions of the model before we start building and coding. Each assumption if violated means we may have to take extra steps to improve our model or in some cases dump the model altogether. Here is a list of the assumptions of the model:
- Regression residuals must be normally distributed.
- A linear relationship is assumed between the dependent variable and the independent variables.
- The residuals are homoscedastic and approximately rectangular-shaped.
- Absence of multicollinearity is expected in the model, meaning that independent variables are not too highly correlated.
- No Autocorrelation of the residuals.
I will be explaining these assumptions in more detail as we arrive at each of them in the tutorial. At this point, however, we need to have an idea of what they are.
***
## Section One: Import our Libraries
The first thing we need to do is import the libraries we will be using in this tutorial. To visualize our data, we will be using `matplotlib` and `seaborn` to create heatmaps and a scatter matrix. To build our model, we will be using the `sklearn` library, and the evaluation will be taking place with the `statsmodels` library. I've also added a few additional modules to help calculate certain metrics.
```
import numpy as np
import pandas as pd
import seaborn as sns
from scipy import stats
import matplotlib.pyplot as plt
import statsmodels.api as sm
from statsmodels.stats import diagnostic as diag
from statsmodels.stats.outliers_influence import variance_inflation_factor
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error, r2_score, mean_absolute_error
%matplotlib inline
```
## Section Two: Load the Data into Pandas
After we've loaded our libraries, we can begin the process of importing and exploring our data. I've created an excel file with all the data we will be using in this tutorial. It contains 10 explanatory variables and 1 dependent variable. After we've loaded the data into the data frame, we will need to replace all the `..` values with `nan` as these represent missing values in our dataset.
This dataset was downloaded from the World Bank website; if you would like to visit the site yourself, I encourage you to visit the link provided below. There is a tremendous amount of data available for free, that can be used across a wide range of models.
Link: https://data.worldbank.org/
From here, we will set the index of our data frame using the `set_index()` function to the `Year` column. The reasoning behind this is because it will make selecting the data easier. After we've defined the index, we convert the entire data frame to a `float` data type and then select years `1969 to 2016`. **These years were selected because they do not contain any missing values.**
To make selecting the columns a little easier, we will rename all of our columns. I'll create a dictionary where the keys represent the old column names and the values associated with those keys are the new column names. I'll then call the `rename()` method and pass through the new columns dictionary.
Finally, I'll check one last time for any missing values using `isnull().any()`, which will return true for a given column if any values are missing, and then print the head of the data frame.
```
# load the data and replace the '..' with nan
econ_df = pd.read_excel('korea_data.xlsx')
econ_df = econ_df.replace('..','nan')
# set the index to the year column
econ_df = econ_df.set_index('Year')
# set the data type and select rows up to 2016
econ_df = econ_df.astype(float)
econ_df = econ_df.loc['1969':'2016']
column_names = {'Unemployment, total (% of total labor force) (national estimate)':'unemployment',
'GDP growth (annual %)': 'gdp_growth',
'Gross capital formation (% of GDP)':'gross_capital_formation',
'Population growth (annual %)':'pop_growth',
'Birth rate, crude (per 1,000 people)':'birth_rate',
'Broad money growth (annual %)':'broad_money_growth',
'Final consumption expenditure (% of GDP)':'final_consum_gdp',
'Final consumption expenditure (annual % growth)':'final_consum_growth',
'General government final consumption expenditure (annual % growth)':'gov_final_consum_growth',
'Gross capital formation (annual % growth)':'gross_cap_form_growth',
'Households and NPISHs Final consumption expenditure (annual % growth)':'hh_consum_growth'}
# rename columns
econ_df = econ_df.rename(columns = column_names)
# check for nulls
display('-'*100)
display(econ_df.isnull().any())
# display the first five rows
display('-'*100)
display(econ_df.head())
```
## Section Three: Check for Perfect Multicollinearity
One of the first things we can do after loading our data is to validate one of the assumptions of our model; in this case, we will be checking for multicollinearity.
### What is multicollinearity?
One of the assumptions of our model is that there isn't any Perfect multicollinearity. Multicollinearity is where one of the explanatory variables is highly correlated with another explanatory variable. **In essence, one of the X variables is almost perfectly correlated with another or multiple X variables.**
### What is the problem with multicollinearity?
The problem with multicollinearity, from a math perspective, is that the coefficient estimates themselves tend to be unreliable. Additionally, the standard errors of slope coefficients become artificially inflated. **Because the standard error is used to help calculate the p-value, this leads to a higher probability that we will incorrectly conclude that a variable is not statistically significant.**
Another way we can look at this problem is by using an analogy. Imagine we ask you to go to a concert and determine who was the best singer. This task would become very challenging if you couldn't distinguish the two singers because they are singing at the same volume. **The idea is the same in our analysis, how can we determine which variable is playing a role in our model if we can't distinguish the two? The problem is we can't.**
Now a little correlation is fine, but if it gets too high, we can effectively distinguish the two variables. The other issue that arises is that when we have highly correlated exploratory variables is that, in a sense, we have duplicates. This means that we can remove one of them and we haven't lost anything; the model would still perform the same.
### How to test for multicollinearity?
Because of these drawbacks, we should always check for multicollinearity in our data. Now, in the step above I purposely pull in variables that I knew would be highly correlated with each other; that way we could see some examples of variables that would cause some issues.
The first thing we can do is create a correlation matrix using the `corr()` function; this will create a matrix with each variable having its correlation calculated for all the other variables. Keep in mind, if you travel diagonally down the matrix all the associations should be one, as it is calculating the correlation of the variable with itself. When we have multiple variables as we do, I sometimes prefer to use a correlation heatmap this way I can quickly identify the highly correlated variables, by just looking for the darker colors.
```
# calculate the correlation matrix
corr = econ_df.corr()
# display the correlation matrix
display(corr)
# plot the correlation heatmap
sns.heatmap(corr, xticklabels=corr.columns, yticklabels=corr.columns, cmap='RdBu')
```
***
Looking at the heatmap along with the correlation matrix we can identify a few highly correlated variables. For example, if you look at the correlation between `birth_rate` and `pop_growth` it ends up at almost .98. This is an extremely high correlation and marks it as a candidate to be removed. Logically it makes sense that these two are highly correlated; if you're having more babies, then the population should be increasing.
However, we should be more systematic in our approach to removing highly correlated variables. One method we can use is the `variance_inflation_factor` which **is a measure of how much a particular variable is contributing to the standard error in the regression model. When significant multicollinearity exists, the variance inflation factor will be huge for the variables in the calculation.**
A general recommendation is that if any of our variables come back with a **value of 5 or higher, then they should be removed from the model.** I decided to show you how the VFI comes out before we drop the highly correlated variables and after we remove the highly correlated variables. Going forward in the tutorial we will only be using the `econ_df_after` data frame.
```
# define two data frames one before the drop and one after the drop
econ_df_before = econ_df
econ_df_after = econ_df.drop(['gdp_growth','birth_rate', 'final_consum_growth','gross_capital_formation'], axis = 1)
# the VFI does expect a constant term in the data, so we need to add one using the add_constant method
X1 = sm.tools.add_constant(econ_df_before)
X2 = sm.tools.add_constant(econ_df_after)
# create the series for both
series_before = pd.Series([variance_inflation_factor(X1.values, i) for i in range(X1.shape[1])], index=X1.columns)
series_after = pd.Series([variance_inflation_factor(X2.values, i) for i in range(X2.shape[1])], index=X2.columns)
# display the series
print('DATA BEFORE')
print('-'*100)
display(series_before)
print('DATA AFTER')
print('-'*100)
display(series_after)
```
Looking at the data above we now get some confirmation about our suspicion. It makes sense to remove either `birth_rate` or `pop_growth` and some of the consumption growth metrics. Once we remove those metrics and recalculate the VFI, we get a passing grade and can move forward.
***
I also want to demonstrate another way to visualize our data to check for multicollinearity. Inside of `pandas`, there is a `scatter_matrix` chart that will create a scatter plot for each variable in our dataset against another variable. This is a great tool for visualizing the correlation of one variable across all the other variables in the dataset. I'll take my econ_df_after and pass it through the `scatter_matrix` method. What you're looking for is a more random distribution, there shouldn't be any strong trends in the scatter matrix as this would be identifying correlated variables. Now, for our explanatory variable, we want to see trends!
```
# define the plot
pd.plotting.scatter_matrix(econ_df_after, alpha = 1, figsize = (30, 20))
# show the plot
plt.show()
```
***
## Section Four: Describe the Data Set
Before we get to an in-depth exploration of the data or even building the model, we should explore the data a little more and see how the data is distributed and if there are any outliers. I will be adding a few more metrics to the `summary data frame`, sp that it now includes a metric for three standard deviations below and above the mean.
I'll store my information in a new variable called `desc_df`.
```
# get the summary
desc_df = econ_df.describe()
# add the standard deviation metric
desc_df.loc['+3_std'] = desc_df.loc['mean'] + (desc_df.loc['std'] * 3)
desc_df.loc['-3_std'] = desc_df.loc['mean'] - (desc_df.loc['std'] * 3)
# display it
desc_df
```
***
One thing that I want to mention is that we have only 50 observations, but 6 (minus the 3 we dropped) exploratory variables. Many people would argue that we need more data to have this many exploratory variables and to be honest, they are correct. **Generally we should aim for at least 20 instances for each variable; however, some argue only 10 would do.** Regardless, we will see at the end of our model that we only end up with 4 exploratory variables so that we will satisfy that rule.
Looking at the data frame up above, a few values are standing out, for example, the maximum value in the `broad_money_growth` column is almost four standard deviations above the mean. Such an enormous value would qualify as an outlier.
### Filtering the Dataset
To drop or not to drop, that is the question. Generally, if we believe the data has been entered in error, we should remove it. However, in this situation, the values that are being identified as outliers are correct values and are not errors. Both of these values were produced during specific moments in time. The one in 1998 was right after the Asian Financial Crisis, and the one in 2001 is right after the DotCom Bubble, so it's entirely conceivable that these values were produced in extreme albeit rare conditions. **For this reason, I will NOT be removing these values from the dataset as they recognize actual values that took place.**
Imagine if we wanted to remove the values that have an amount exceeding three standard deviations. How would we approach this? Well, if we leverage the `numpy` module and the `scipy` module we can filter out the rows using the `stats.zscore` function. The Z-score is the number of standard deviations from the mean a data point is, so if it's less than 3 we keep it otherwise we drop it. From here, I also provided a way to let us know what rows were removed by using the `index.difference` the function which will show the difference between the two datasets.
```
# filter the data frame to remove the values exceeding 3 standard deviations
econ_remove_df = econ_df[(np.abs(stats.zscore(econ_df)) < 3).all(axis=1)]
# what rows were removed
econ_df.index.difference(econ_remove_df.index)
```
***
## Section Five: Build the Model
Okay, now that we've loaded, cleaned, and explored the data we can proceed to the next part, building the model. The first thing we need to do is, define our exploratory variables and our explanatory variable. From here, let's split the data into a training and testing set; a healthy ratio is 20% testing and 80% training but a 30% 70% split is also ok.
After splitting the data, we will create an instance of the linear regression model and pass through the `X_train` and `y_train` variables using the `fit()` function.
```
# define our input variable (X) & output variable
econ_df_after = econ_df.drop(['birth_rate', 'final_consum_growth','gross_capital_formation'], axis = 1)
X = econ_df_after.drop('gdp_growth', axis = 1)
Y = econ_df_after[['gdp_growth']]
# Split X and y into X_
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.20, random_state=1)
# create a Linear Regression model object
regression_model = LinearRegression()
# pass through the X_train & y_train data set
regression_model.fit(X_train, y_train)
```
***
### Exploring the Output
With the data now fitted to the model, we can explore the output. The first thing we should do is look at the intercept of the model, and then we will print out each of the coefficients of the model. I print everything out using a loop to make it more efficient.
```
# let's grab the coefficient of our model and the intercept
intercept = regression_model.intercept_[0]
coefficent = regression_model.coef_[0][0]
print("The intercept for our model is {:.4}".format(intercept))
print('-'*100)
# loop through the dictionary and print the data
for coef in zip(X.columns, regression_model.coef_[0]):
print("The Coefficient for {} is {:.2}".format(coef[0],coef[1]))
```
**The intercept term is the value of the dependent variable when all the independent variables are equal to zero. For each slope coefficient, it is the estimated change in the dependent variable for a one unit change in that particular independent variable, holding the other independent variables constant.**
For example, if all the independent variables were equal to zero, then the `gdp_growth` would be 2.08%. If we looked at the `gross_cap_form_growth` while *holding all the other independent variables constant*, then we would say for a 1 unit increase in `gross_cap_form_growth` would lead to a 0.14% increase in GDP growth.
***
We can also now make predictions with our newly trained model. The process is simple; we call the `predict` method and then pass through some values. In this case, we have some values predefined with the `x_test` variable so we will pass that through. Once we do that, we can select the predictions by slicing the array.
```
# Get multiple predictions
y_predict = regression_model.predict(X_test)
# Show the first 5 predictions
y_predict[:5]
```
***
## Section Six: Evaluating the Model
### Using the `Statsmodel`
To make diagnosing the model easier, we will, from this point forward, be using the `statsmodel` module. This module has built-in functions that will make calculating metrics quick. However, we will need "rebuild" our model using the `statsmodel` module. We do this by creating a constant variable, call the `OLS()` method and then the `fit()` method. We now have a new model, and the first thing we need to do is to make sure that the assumptions of our model hold. This means checking the following:
- Regression residuals must be normally distributed.
- The residuals are homoscedastic
- Absence of multicollinearity (we did this above).
- No Autocorrelation.
```
# define our intput
X2 = sm.add_constant(X)
# create a OLS model
model = sm.OLS(Y, X2)
# fit the data
est = model.fit()
```
## Checking for Heteroscedasticity
### What is Heteroscedasticity?
One of the assumptions of our model is that there is no heteroscedasticity. What exactly does this mean? Well, to give a simple definition it merely means the standard errors of a variable, monitored over a specific amount of time, are non-constant. Let's imagine a situation where heteroscedasticity could exist.
Imagine we modeled household consumption based on income, something we would probably notice is how the variability of expenditures changes depending on how much income you have. In simple terms, we would see that households with more income spend money on a broader set of items compared to lower income households that would only be able to focus on the main staples. This results in standard errors that change over income levels.
***
### What is the problem with heteroscedasticity?
There are two big reasons why you want homoscedasticity:
1. While heteroscedasticity does not cause bias in the coefficient estimates, **it causes the coefficient estimates to be less precise.** The Lower precision increases the likelihood that the coefficient estimates are further from the correct population value.
2. **Heteroscedasticity tends to produce p-values that are smaller than they should be.** This effect occurs because heteroscedasticity increases the variance of the coefficient estimates, but the OLS procedure does not detect this increase. Consequently, OLS calculates the t-values and F-values using an underestimated amount of variance. This problem can lead you to conclude that a model term is statistically significant when it is not significant.
***
### How to test for heteroscedasticity?
To check for heteroscedasticity, we can leverage the `statsmodels.stats.diagnostic` module. This module will give us to a few test functions we can run, the Breusch-Pagan and the White test for heteroscedasticity. The **Breusch-Pagan is a more general test for heteroscedasticity while the White test is a unique case.**
- The null hypothesis for both the White’s test and the Breusch-Pagan test is that the variances for the errors are equal:
- **H0 = σ2i = σ2**
- The alternate hypothesis (the one you’re testing), is that the variances are not equal:
- **H1 = σ2i ≠ σ2**
Our goal is to fail to reject the null hypothesis, have a high p-value because that means we have no heteroscedasticity.
```
# Run the White's test
_, pval, __, f_pval = diag.het_white(est.resid, est.model.exog, retres = False)
print(pval, f_pval)
print('-'*100)
# print the results of the test
if pval > 0.05:
print("For the White's Test")
print("The p-value was {:.4}".format(pval))
print("We fail to reject the null hypthoesis, so there is no heterosecdasticity. \n")
else:
print("For the White's Test")
print("The p-value was {:.4}".format(pval))
print("We reject the null hypthoesis, so there is heterosecdasticity. \n")
# Run the Breusch-Pagan test
_, pval, __, f_pval = diag.het_breuschpagan(est.resid, est.model.exog)
print(pval, f_pval)
print('-'*100)
# print the results of the test
if pval > 0.05:
print("For the Breusch-Pagan's Test")
print("The p-value was {:.4}".format(pval))
print("We fail to reject the null hypthoesis, so there is no heterosecdasticity.")
else:
print("For the Breusch-Pagan's Test")
print("The p-value was {:.4}".format(pval))
print("We reject the null hypthoesis, so there is heterosecdasticity.")
```
## Checking for Autocorrelation
### What is autocorrelation?
Autocorrelation is a characteristic of data in which the correlation between the values of the same variables is based on related objects. It violates the assumption of instance independence, which underlies most of conventional models.
When you have a series of numbers, and there is a pattern such that values in the series can be predicted based on preceding values in the series, the set of numbers is said to exhibit autocorrelation. This is also known as serial correlation and serial dependence. It generally exists in those types of data-sets in which the data, instead of being randomly selected, are from the same source.
***
### What is the problem with autocorrelation?
The existence of autocorrelation means that computed standard errors, and consequently p-values, are misleading. Autocorrelation in the residuals of a model is also a sign that the model may be unsound. A workaround is we can compute more robust standard errors.
***
### How to test for autocorrelation?
Again, we will go to our favorite module the `statsmodels.stats.diagnostic` module, and use the Ljung-Box test for no autocorrelation of residuals. Here:
- **H0: The data are random.**
- **Ha: The data are not random.**
That means we want to fail to reject the null hypothesis, have a large p-value because then it means we have no autocorrelation. To use the Ljung-Box test, we will call the `acorr_ljungbox` function, pass through the `est.resid` and then define the lags.
The lags can either be calculated by the function itself, or we can calculate them. If the function handles it the max lag will be `min((num_obs // 2 - 2), 40)`, however, there is a rule of thumb that for non-seasonal time series the lag is ` min(10, (num_obs // 5))`.
We also can visually check for autocorrelation by using the `statsmodels.graphics` module to plot a graph of the autocorrelation factor.
```
# test for autocorrelation
from statsmodels.stats.stattools import durbin_watson
# calculate the lag, optional
lag = min(10, (len(X)//5))
print('The number of lags will be {}'.format(lag))
print('-'*100)
# run the Ljung-Box test for no autocorrelation of residuals
# test_results = diag.acorr_breusch_godfrey(est, nlags = lag, store = True)
test_results = diag.acorr_ljungbox(est.resid, lags = lag)
# grab the p-values and the test statistics
ibvalue, p_val = test_results
# print the results of the test
if min(p_val) > 0.05:
print("The lowest p-value found was {:.4}".format(min(p_val)))
print("We fail to reject the null hypthoesis, so there is no autocorrelation.")
print('-'*100)
else:
print("The lowest p-value found was {:.4}".format(min(p_val)))
print("We reject the null hypthoesis, so there is autocorrelation.")
print('-'*100)
# plot autocorrelation
sm.graphics.tsa.plot_acf(est.resid)
plt.show()
```
## Checking For Normally Distributed Residuals
This one is easy to check for; we will do it visually. **This will require using a QQ pplot which help us assess if a set of data plausibly came from some theoretical distribution such as a Normal or exponential.** It’s just a visual check, not an air-tight proof, so it is somewhat subjective.
Visually what we are looking for is the data hugs the line tightly; this would give us confidence in our assumption that the residuals are normally distributed. Now, it is highly unlikely that the data will perfectly hug the line, so this is where we have to be subjective.
## Checking the Mean of the Residuals Equals 0
Additionally, we need to check another assumption, that the mean of the residuals is equal to zero. If the mean is very close to zero, then we are good to proceed. Just a side note, it's not uncommon to get a mean that isn't exactly zero; this is because of rounding errors. However, if it's very close to zero, it's ok. In the example below, you will see that it doesn't come out exactly to zero.
```
import pylab
# check for the normality of the residuals
sm.qqplot(est.resid, line='s')
pylab.show()
# also check that the mean of the residuals is approx. 0.
mean_residuals = sum(est.resid)/ len(est.resid)
print("The mean of the residuals is {:.4}".format(mean_residuals))
```
### Measures of Error
We can examine how well our data fit the model, so we will take `y_predictions` and compare them to our `y_actuals` these will be our residuals. From here we can calculate a few metrics to help quantify how well our model fits the data. Here are a few popular metrics:
- **Mean Absolute Error (MAE):** Is the mean of the absolute value of the errors. This gives an idea of magnitude but no sense of direction (too high or too low).
- **Mean Squared Error (MSE):** Is the mean of the squared errors. MSE is more popular than MAE because MSE "punishes" more significant errors.
- **Root Mean Squared Error (RMSE):** Is the square root of the mean of the squared errors. RMSE is even more favored because it allows us to interpret the output in y-units.
Luckily for us, `sklearn` and `statsmodel` both contain functions that will calculate these metrics for us. The examples below were calculated using the `sklearn` library and the `math` library.
```
import math
# calculate the mean squared error
model_mse = mean_squared_error(y_test, y_predict)
# calculate the mean absolute error
model_mae = mean_absolute_error(y_test, y_predict)
# calulcate the root mean squared error
model_rmse = math.sqrt(model_mse)
# display the output
print("MSE {:.3}".format(model_mse))
print("MAE {:.3}".format(model_mae))
print("RMSE {:.3}".format(model_rmse))
```
***
### R-Squared
The R-Squared metric provides us a way to measure the goodness of fit or, in other words, how well our data fits the model. The higher the R-Squared metric, the better the data fit our model. However, one limitation is that R-Square increases as the number of features increase in our model, so if I keep adding variables even if they're poor choices R-Squared will still go up! **A more popular metric is the adjusted R-Square which penalizes more complex models, or in other words models with more exploratory variables.** In the example below, I calculate the regular R-Squared value, however, the `statsmodel` summary will calculate the Adjusted R-Squared below.
```
model_r2 = r2_score(y_test, y_predict)
print("R2: {:.2}".format(model_r2))
```
***
### Confidence Intervals
Let's look at our confidence intervals. Keep in mind that by default confidence intervals are calculated using 95% intervals. We interpret confidence intervals by saying if the population from which this sample was drawn was sampled 100 times. **Approximately 95 of those confidence intervals would contain the "true" coefficient.**
Why do we provide a confidence range? Well, it comes from the fact that we only have a sample of the population, not the entire population itself. Because of this, it means that the "true" coefficient could exist in the interval below or it couldn't, but we cannot say for sure. We provide some uncertainty by providing a range, usually 95%, where the coefficient is probably in.
- Want a narrower range? **Decrease your confidence**.
- Want a wider range? **Increase your confidence**.
```
# make some confidence intervals, 95% by default
est.conf_int()
```
### Hypothesis Testing
With hypothesis testing, we are trying to determine the statistical significance of the coefficient estimates. This test is outlined as the following.
- **Null Hypothesis:** There is no relationship between the exploratory variables and the explanatory variable.
- **Alternative Hypothesis:** There is a relationship between the exploratory variables and the explanatory variable.
***
- If we **reject the null**, we are saying there is a relationship, and the coefficients do not equal 0.
- If we **fail to reject the null**, we are saying there is no relationship, and the coefficients do equal 0
```
# estimate the p-values
est.pvalues
```
Here it's a little hard to tell, but we have a few insignificant coefficients. The first is the constant itself, so technically this should be dropped. However, we will see that once we remove the irrelevant variables that the intercept becomes significant. **If it still wasn't significant, we could have our intercept start at 0 and assume that the cumulative effect of X on Y begins from the origin (0,0).** Along with the constant, we have `unemployment` and `broad_money_growth` both come out as insignificant.
***
### Create a Summary of the Model Output
Let's create a summary of some of our keep metrics, `sklearn` does not have a good way of creating this output so we would have to calculate all the parameters ourselves. Let's avoid this and use the `statsmodel.api` library as we can create the same model we did up above, but we can also leverage the `summary()` method to create an output for us. Some of the metrics might differ slightly, but they generally should be the same.
```
# print out a summary
print(est.summary())
```
The first thing we notice is that the p-values from up above are now easier to read and we can now determine that the coefficients that have a p-value greater than 0.05 can be removed. We also have our 95% confidence interval (described up above), our coefficient estimates (described up above), the standard errors, and t-values.
The other metric that stands out is our Adjusted R-Squared value which is .878, lower than our R-Squared value. This makes sense as we were probably docked for the complexity of our model. However, an R-Squared over .878 is still very strong.
The only additional metrics we will describe here is the t-value which is the coefficient divided by the standard error. The higher the t-value, the more evidence we have to reject the null hypothesis. Also the standard error, the standard error is the approximate standard deviation of a statistical sample population.
***
## Section Seven: Remove the Insignificant Variables.
Now that we know which variables are insignificant we should remove them from the model and refit the data to see what we get, the steps are the same the only thing I'm changing is that I am removing some additional columns from the data frame.
```
# define our input variable (X) & output variable
econ_df_after = econ_df.drop(['birth_rate', 'final_consum_growth','gross_capital_formation','broad_money_growth',
'unemployment'], axis = 1)
X = econ_df_after.drop('gdp_growth', axis = 1)
Y = econ_df_after[['gdp_growth']]
# Split X and y into X_
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.20, random_state=1)
# create a Linear Regression model object
regression_model = LinearRegression()
# pass through the X_train & y_train data set
regression_model.fit(X_train, y_train)
# define our intput
X2 = sm.add_constant(X)
# create a OLS model
model = sm.OLS(Y, X2)
# fit the data
est = model.fit()
print(est.summary())
```
**Looking at the output, we now see that all of the independent variables are significant and even our constant is significant.** We could rerun our test for autocorrelation and, but the tests will take you to the same conclusions we found above so I decided to leave that out of the tutorial. At this point, we can interpret our formula and begin making predictions. Looking at the coefficents, we would say `pop_growth`, `gross_cap_form_growth`, and `hh_consum_growth` all have a positive effect on GDP growth. Additionally, we would say that `gov_final_consum_growth` has a negative effect on GDP growth. That's a little surprising to see, but we would have to see why that might be the case.
***
## Section Eight: Save the Model for Future Use
We will probably want to use this model in the future, so let us save our work so we can use it later. Saving the model can be achieved by storing our model in a pickle which is storing a python object as a character stream in a file which can be reloaded later to use.
```
import pickle
# pickle the model
with open('my_mulitlinear_regression.sav','wb') as f:
pickle.dump(regression_model, f)
# load it back in
with open('my_mulitlinear_regression.sav', 'rb') as pickle_file:
regression_model_2 = pickle.load(pickle_file)
# make a new prediction
regression_model_2.predict([X_test.loc[2002]])
```
| github_jupyter |
# PINN: Heat equation with variable diffusion
Solving the heat equation in 2D for variable diffusion D using the PINN-concept.
```
import torch
import torchphysics as tp
import math
```
First, we create the spaces for our problem. These define the variable names which will be used in the remaining part of this code.
In this example, x is the space variable, t corresponds to the time, D is an interval of diffusions and u is the variable for the (1D-)solution.
```
X = tp.spaces.R2('x')
T = tp.spaces.R1('t')
D = tp.spaces.R1('D')
U = tp.spaces.R1('u')
```
As a next step, we build the domain of the problem. There are multiple options to build multi-dimensional domains - in this case, we simply create a rectangle in space and intervals in time and diffusion which will later be multiplied to obtain the cartesian product.
```
h, w = 20, 20
A_x = tp.domains.Parallelogram(X, [0, 0], [w, 0], [0, h])
A_t = tp.domains.Interval(T, 0, 40)
A_D = tp.domains.Interval(D, 0.1, 1.0)
```
Before we visualize the created domain, we create Sampler objects which are iterators that sample points from the domain during the optimization task. There are again various options to sample from the domains, an easy way would be to sample uniformly distributed random points. In this example, we choose an adaptive sampler to sample points in the inner of the domain. It will sample more points in points where the loss is large.
The amount of sampled points is defined by their density in the 3/2-dim subset, it could be increased to achieve better training results.
```
inner_sampler = tp.samplers.AdaptiveRandomRejectionSampler(A_x*A_t*A_D, density=1)
# initial values should be sampled on the left boundary of the time interval and for every x and D
initial_v_sampler = tp.samplers.RandomUniformSampler(A_x*A_t.boundary_left*A_D, density=1)
boundary_v_sampler = tp.samplers.RandomUniformSampler(A_x.boundary*A_t*A_D, density=1)
```
We visualize the domain through the points created by the samplers using matplotlib:
```
tp.utils.scatter(X*T, inner_sampler, initial_v_sampler, boundary_v_sampler)
```
In the next step we define the NN-model we want to fit to the PDE. A normalization can improve convergence for large or small domains.
```
model = tp.models.Sequential(
tp.models.NormalizationLayer(A_x*A_t*A_D),
tp.models.FCN(input_space=X*T*D, output_space=U, hidden=(50,50,50))
)
```
Now, we define a condition which aims to minimze the mean squared error of the residual of the poisson equation.
```
def heat_residual(u, x, t, D):
return D*tp.utils.laplacian(u, x) - tp.utils.grad(u, t)
pde_condition = tp.conditions.PINNCondition(module=model,
sampler=inner_sampler,
residual_fn=heat_residual,
name='pde_condition')
```
Additionally, we add a boundary condition at the boundary of the domain:
```
def boundary_v_residual(u):
return u
boundary_v_condition = tp.conditions.PINNCondition(module=model,
sampler=boundary_v_sampler,
residual_fn=boundary_v_residual,
name='boundary_condition')
```
The initial condition can be defined via a data function. Again, we minimize the mean squared error over the sampled points.
```
def f(x):
return torch.sin(math.pi/w*x[:, :1])*torch.sin(math.pi/h*x[:,1:])
def initial_v_residual(u, f):
return u-f
initial_v_condition = tp.conditions.PINNCondition(module=model,
sampler=initial_v_sampler,
residual_fn=initial_v_residual,
data_functions={'f': f},
name='initial_condition')
```
For comparison, we compute the solution via a finite difference scheme.
```
import sys
sys.path.append('..')
from fdm_heat_equation import FDM, transform_to_points
fdm_domain, fdm_time_domains, fdm_solution = FDM([0, w, 0, h], 2*[2e-1], [0,5], [1.0, 1.5, 2.0, 2.5, 3.0, 3.5, 4.0], f)
fdm_inp, fdm_out = transform_to_points(fdm_domain, fdm_time_domains, fdm_solution, [1.0, 1.5, 2.0, 2.5, 3.0, 3.5, 4.0], True)
```
Comparsion to measured or computed data can be performed via a DataCondition using data supplied via a PointsDataLoader.
```
val_condition = tp.conditions.DataCondition(module=model,
dataloader=tp.utils.PointsDataLoader((fdm_inp, fdm_out), batch_size=80000),
norm='inf')
```
Finally, we optimize the conditions using a pytorch-lightning.LightningModule Solver and running the training. In the Solver, the training and validation conditions, as well as all optimizer options can be specified.
```
solver = tp.solver.Solver([pde_condition,
boundary_v_condition,
initial_v_condition], [val_condition])
import pytorch_lightning as pl
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
trainer = pl.Trainer(gpus=1, # or None for CPU
max_steps=2000,
logger=False,
benchmark=True,
val_check_interval=400,
checkpoint_callback=False)
trainer.fit(solver)
```
Finally, we plot the obtained solution:
```
anim_sampler = tp.samplers.AnimationSampler(A_x, A_t, 100, n_points=400, data_for_other_variables={'D': 1.0})
anim = tp.utils.animate(model, lambda u: u[:, 0], anim_sampler, ani_speed=10)
```
| github_jupyter |
```
import numpy as np
import torch
import pandas as pd
from transformers import PreTrainedTokenizerFast
import re
import spacy
nlp = spacy.load("en_core_web_sm")
tokenizer_bert = PreTrainedTokenizerFast.from_pretrained('bert-base-uncased', do_lower_case=True,return_offsets_mapping = True, max_length=512,truncate=True,add_special_tokens=False,return_token_type_ids=False,return_attention_mask=False)
vocab_sorted = {k: v for k, v in sorted(tokenizer_bert.vocab.items(), key=lambda item: item[1])}
```
## Picking adjectives
```
words=[]
for item in vocab_sorted.items():
if re.match('[a-z]{2,}$',item[0]):
words.append(item[0])
len(words)
nouns = []
adjs = []
for ix,word in enumerate(words):
if nlp(word)[0].pos_ == 'NOUN' and len(nouns) < 1000:
nouns.append(nlp(word)[0].text)
elif nlp(word)[0].pos_ == 'ADJ' and len(adjs) < 2000:
adjs.append(nlp(word)[0].text)
```
## Finding gradable adjectives
```
from collections import defaultdict
import textacy
import textacy.datasets
cw = textacy.datasets.CapitolWords()
cw.download()
adjectives_encountered = []
unique_adjectives_encountered = set()
for text,record in cw.records():
processed = nlp(text)
adjectives_encountered += [token for token in processed if token.text in adjs]
for token in processed:
if token.text in adjs:
unique_adjectives_encountered |= set([token.text])
len(adjectives_encountered),len(unique_adjectives_encountered)
gradable = defaultdict(int)
non_gradable = defaultdict(int)
modifiers = ['somewhat','very','really','extremely','rather']
for adj in adjectives_encountered:
if len([x for x in adj.children if x.text in modifiers])>0:
gradable[adj.text] += 1
else:
non_gradable[adj.text]+=1
combined = defaultdict(list)
for adj in unique_adjectives_encountered:
toAdd = []
toAdd.append(gradable[adj])
toAdd.append(non_gradable[adj])
combined[adj] = toAdd
adjs = defaultdict(list)
for adj in combined:
occurences = sum(combined[adj])
gradability_score = round(float((combined[adj][0])/occurences) * 100, 3)
if occurences > 100 and gradability_score > 0.5:
adjs[adj] = gradability_score
adjs = [k for k, v in sorted(adjs.items(), key=lambda item: item[1],reverse=True)][:200]
with open('gradable_adjectives.txt', 'w') as f:
for item in adjs:
f.write("%s\n" % item)
```
## Generating sentences
```
sentences = []
for noun in nouns:
for adj in adjs:
sentences.append('The '+noun+' is '+adj+'.')
sentences.append('The '+noun+' are '+adj+'.')
len(sentences)
sentences[:10]
```
## Filtering by GPT perplexity
```
from pytorch_pretrained_bert import GPT2LMHeadModel, GPT2Tokenizer
device = torch.device('cuda:0')
model_id = 'gpt2'
model_gpt = GPT2LMHeadModel.from_pretrained(model_id).to(device)
tokenizer_gpt = GPT2Tokenizer.from_pretrained(model_id)
def process_gpt(sentence):
tokens = ["[CLS]"] + tokenizer_gpt.tokenize(sentence)
tokens_ids = tokenizer_gpt.convert_tokens_to_ids(tokens)
tokens_ids = torch.tensor([tokens_ids,], dtype=torch.long).to(device)
with torch.no_grad():
outputs = model_gpt(tokens_ids, lm_labels=tokens_ids)
log_likelihood = outputs.item()
return np.exp(log_likelihood)
pairs = {}
for sentence in sentences:
pairs[sentence] = process_gpt(sentence)
df = pd.DataFrame.from_dict(pairs, orient='index').reset_index()
df = df.rename(columns={"index": "sentence", 0: "perplexity"})
df.sort_values(by='perplexity', ascending=True)
df.sort_values(by='perplexity', ascending=True).to_csv('/home/lisa/hobbies/modifiers_all.csv')
df.sort_values(by='perplexity', ascending=True).head(10000).to_csv('/home/lisa/hobbies/modifiers_top10k.csv')
```
## Producing negative vs positive sentence pairs
```
ten_k = df.sort_values(by='perplexity', ascending=True).head(10000)
pos_10k = []
neg_10k = []
for sentence in ten_k['sentence'].values:
words = sentence.split(' ')
aff = ' '.join(words[:3]+['[MASK]']+words[3:])
if words[2] == 'is':
neg = ' '.join(words[:2]+["isn't [MASK]"]+words[3:])
else:
neg = ' '.join(words[:2]+["aren't [MASK]"]+words[3:])
pos_10k.append(aff)
neg_10k.append(neg)
with open('10k_aff.txt', 'a') as f:
for sentence in pos_10k:
f.write(sentence+'\n')
with open('10k_neg.txt', 'a') as f:
for sentence in neg_10k:
f.write(sentence+'\n')
```
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# Two Formulations of Maxwell's equations in Cartesian Coordinates
## Authors: Patrick Nelson & Zach Etienne
### Formatting improvements courtesy Brandon Clark
[comment]: <> (Abstract: TODO)
**Notebook Status:** <font color='orange'><b> Self-Validated </b></font>
**Validation Notes:** This tutorial notebook has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented below, [here](#code_validation_sys1) and [here](#code_validation). **Additional validation tests may have been performed, but are as yet, undocumented. (TODO)**
### NRPy+ Source Code for this module:
* [Maxwell/MaxwellCartesian_Evol.py](../edit/Maxwell/MaxwellCartesian_Evol.py)
* [Maxwell/MaxwellCartesian_ID.py](../edit/Maxwell/MaxwellCartesian_ID.py)
## Introduction:
This tutorial will draw on previous work done by Ian Ruchlin on [Maxwell's equations in Curvilinear Coordinates](Tutorial-MaxwellCurvilinear.ipynb), which itself drew on the two formulations described in [Illustrating Stability Properties of Numerical Relativity in Electrodynamics](https://arxiv.org/abs/gr-qc/0201051). This will be done to aid construction of an Einstein Toolkit thorn, which will itself be built in the [Tutorial-ETK_thorn-Maxwell](Tutorial-ETK_thorn-Maxwell.ipynb) tutorial notebook.
The construction of our equations here will be nearly identical; however, by assuming Cartesian coordinates, we are able to make several simplifications and eliminate the need for [reference_metric.py](../edit/reference_metric.py) as a dependency.
We will begin with their System I. While the Curvilinear version of this code assumed flat spacetime, we will be constructing the equations in a general spacetime. This allows us to simply replace the reference metric "hatted" quantities with their general counterparts.
<a id='toc'></a>
# Table of Contents:
$$\label{toc}$$
This notebook is organized as follows
1. [Step 1](#initializenrpy): Import core NRPy+ modules and set NRPy+ parameters
1. [Step 2](#laplacian): Constructing the Covariant Laplacian
1. [Step 3](#violation): Measuring the Constraint violation
1. [Step 4](#code_validation_sys1): Code Validation against `Maxwell.MaxwellCartesian_Evol` NRPy+ Module (System I)
1. [Step 5](#code_validation_sys2): Code Validation against `Maxwell.MaxwellCartesian_Evol` NRPy+ Module (System II)
1. [Step 6](#id): Constructing the Initial Data
1. [Step 7](#code_validation): Code Validation against `Maxwell.MaxwellCartesian_ID` NRPy+ Module
1. [Step 8](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
<a id='initializenrpy'></a>
# Step 1: Import core NRPy+ modules and set NRPy+ parameters \[Back to [top](#toc)\]
$$\label{initializenrpy}$$
```
#Step P1: Import needed Python modules
import NRPy_param_funcs as par # NRPy+: Parameter interface
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import finite_difference as fin # NRPy+: Finite difference C code generation module
import grid as gri # NRPy+: Functions having to do with numerical grids
#Step P2: Name this Python module
thismodule = "MaxwellCartesian"
#Step 0: Set the spatial dimension parameter to 3.
par.set_parval_from_str("grid::DIM", 3)
DIM = par.parval_from_str("grid::DIM")
# Step 1: Set the finite differencing order to 4.
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER", 4)
# Step 2: Register gridfunctions that are needed as input.
psi = gri.register_gridfunctions("EVOL", ["psi"])
# Step 3a: Declare the rank-1 indexed expressions E_{i}, A_{i},
# and \partial_{i} \psi. Derivative variables like these
# must have an underscore in them, so the finite
# difference module can parse the variable name properly.
ED = ixp.register_gridfunctions_for_single_rank1("EVOL", "ED")
AD = ixp.register_gridfunctions_for_single_rank1("EVOL", "AD")
psi_dD = ixp.declarerank1("psi_dD")
x,y,z = gri.register_gridfunctions("AUX",["x","y","z"])
## Step 3b: Declare the conformal metric tensor and its first
# derivative. These are needed to find the Christoffel
# symbols, which we need for covariant derivatives.
gammaDD = ixp.register_gridfunctions_for_single_rank2("AUX","gammaDD", "sym01") # The AUX or EVOL designation is *not*
# used in diagnostic modules.
gammaDD_dD = ixp.declarerank3("gammaDD_dD","sym01")
gammaDD_dDD = ixp.declarerank4("gammaDD_dDD","sym01_sym23")
gammaUU, detgamma = ixp.symm_matrix_inverter3x3(gammaDD)
gammaUU_dD = ixp.declarerank3("gammaDD_dD","sym01")
# Define the Christoffel symbols
GammaUDD = ixp.zerorank3(DIM)
for i in range(DIM):
for k in range(DIM):
for l in range(DIM):
for m in range(DIM):
GammaUDD[i][k][l] += (sp.Rational(1,2))*gammaUU[i][m]*\
(gammaDD_dD[m][k][l] + gammaDD_dD[m][l][k] - gammaDD_dD[k][l][m])
# Step 3b: Declare the rank-2 indexed expression \partial_{j} A_{i},
# which is not symmetric in its indices.
# Derivative variables like these must have an underscore
# in them, so the finite difference module can parse the
# variable name properly.
AD_dD = ixp.declarerank2("AD_dD", "nosym")
# Step 3c: Declare the rank-3 indexed expression \partial_{jk} A_{i},
# which is symmetric in the two {jk} indices.
AD_dDD = ixp.declarerank3("AD_dDD", "sym12")
# Step 4: Calculate first and second covariant derivatives, and the
# necessary contractions.
# First covariant derivative
# D_{j} A_{i} = A_{i,j} - \Gamma^{k}_{ij} A_{k}
AD_dcovD = ixp.zerorank2()
for i in range(DIM):
for j in range(DIM):
AD_dcovD[i][j] = AD_dD[i][j]
for k in range(DIM):
AD_dcovD[i][j] -= GammaUDD[k][i][j] * AD[k]
```
<a id='laplacian'></a>
# Step 2: Constructing the Covariant Laplacian \[Back to [top](#toc)\]
$$\label{laplacian}$$
One particular difficulty we will encounter here is taking the covariant Laplacian of a vector. This will here take the form of $D_j D^j A_i$. We will start with the outer derivative after lowering the index on the second operator. So, we see that
\begin{align}
D_j D^j A_i &= D_j (\gamma^{jk} D_k A_i) \\
&= (D_k A_i) D_j \gamma^{jk} + \gamma^{jk} D_j D_k A_i \\
&= \gamma^{jk} [\partial_j (D_k A_i) - \Gamma^l_{ij} D_k A_l - \Gamma^l_{jk} D_l A_i],
\end{align}
dropping the first term from the second line because $D_j \gamma^{jk} = 0$ Next, we will again apply the covariant derivative to $A_i$. First, however, we should consider that
\begin{align}
D_k A_i &= \partial_k A_i - \Gamma^l_{ik} A_l \\
& = \partial_k A_i - \gamma^{lm} \Gamma_{mik} A_l \\
& = \partial_k A_i - \Gamma_{mik} A^m, \\
\end{align}
where $\Gamma_{ljk} = \frac{1}{2} (\partial_k \gamma_{lj} + \partial_j \gamma_{kl} - \partial_l \gamma_{jk})$ is the Christoffel symbol of the first kind. Note how we were able to use the raising operator to switch the height of two indices; we will use this in upcoming steps. Thus, our expression becomes
\begin{align}
D_j D^j A_i &= \gamma^{jk} [\partial_j \partial_k A_i - \partial_j (\Gamma_{lik} A^l) - \Gamma^l_{ij} \partial_k A_l + \Gamma^m_{ij} \Gamma_{lmk} A^l - \Gamma^l_{jk} \partial_l A_i + \Gamma^m_{jk} \Gamma_{lim} A^l] \\
&= \gamma^{jk} [\partial_j \partial_k A_i - \underbrace{\partial_j (\Gamma_{lik} A^l)}_{\text{sub-term}} - \Gamma^l_{ij} \partial_k A_l + \Gamma^m_{ij} \Gamma^l_{mk} A_l - \Gamma^l_{jk} \partial_l A_i + \Gamma^m_{jk} \Gamma^l_{im} A_l].
\end{align}
Let's focus on the underbraced sub-term for a moment. Expanding this using the product rule and the definition of the Christoffel symbol,
\begin{align}
\partial_j (\Gamma_{lik} A^l) &= A^l \partial_j \Gamma_{lik} + \Gamma_{lik} \partial_j A^l \\
&= A^l \partial_j (\partial_k \gamma_{li} + \partial_i \gamma_{kl} - \partial_l \gamma_{ik}) + \Gamma_{lik} \partial_j (\gamma^{lm} A_m) \\
&= A^l (\gamma_{li,kj} + \gamma_{kl,ij} - \gamma_{ik,lj}) + \Gamma_{lik} (\gamma^{lm} A_{m,j} + A_m \gamma^{lm}{}_{,j}), \\
\end{align}
where commas in subscripts denote partial derivatives.
So, the Laplacian becomes
\begin{align}
D_j D^j A_i &= \gamma^{jk} [A_{i,jk} -
\underbrace{A^l (\gamma_{li,kj} + \gamma_{kl,ij} - \gamma_{ik,lj})}_{\text{Term 1}} +
\underbrace{\Gamma_{lik} (\gamma^{lm} A_{m,j} + A_m \gamma^{lm}{}_{,j})}_{\text{Term 2}} -
\underbrace{(\Gamma^l_{ij} A_{l,k} + \Gamma^l_{jk} A_{i,l})}_{\text{Term 3}} +
\underbrace{(\Gamma^m_{ij} \Gamma^l_{mk} A_l + \Gamma ^m_{jk} \Gamma^l_{im} A_l)}_{\text{Term 4}}]; \\
\end{align}
we will now begin to contruct these terms individually.
```
# First, we must construct the lowered Christoffel symbols:
# \Gamma_{ijk} = \gamma_{il} \Gamma^l_{jk}
# And raise the index on A:
# A^j = \gamma^{ij} A_i
GammaDDD = ixp.zerorank3()
AU = ixp.zerorank1()
for i in range(DIM):
for j in range(DIM):
AU[j] += gammaUU[i][j] * AD[i]
for k in range(DIM):
for l in range(DIM):
GammaDDD[i][j][k] += gammaDD[i][l] * GammaUDD[l][j][k]
# Covariant second derivative (the bracketed terms):
# D_j D^j A_i = \gamma^{jk} [A_{i,jk} - A^l (\gamma_{li,kj} + \gamma_{kl,ij} - \gamma_{ik,lj})
# + \Gamma_{lik} (\gamma^{lm} A_{m,j} + A_m \gamma^{lm}{}_{,j})
# - (\Gamma^l_{ij} A_{l,k} + \Gamma^l_{jk} A_{i,l})
# + (\Gamma^m_{ij} \Gamma^l_{mk} A_l + \Gamma ^m_{jk} \Gamma^l_{im} A_l)
AD_dcovDD = ixp.zerorank3()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
AD_dcovDD[i][j][k] = AD_dDD[i][j][k]
for l in range(DIM):
# Terms 1 and 3
AD_dcovDD[i][j][k] -= AU[l] * (gammaDD_dDD[l][i][k][j] + gammaDD_dDD[k][l][i][j] - \
gammaDD_dDD[i][k][l][j]) \
+ GammaUDD[l][i][j] * AD_dD[l][k] + GammaUDD[l][j][k] * AD_dD[i][l]
for m in range(DIM):
# Terms 2 and 4
AD_dcovDD[i][j][k] += GammaDDD[l][i][k] * (gammaUU[l][m] * AD_dD[m][j] + AD[m] * gammaUU_dD[l][m][j]) \
+ GammaUDD[m][i][j] * GammaUDD[l][m][k] * AD[l] \
+ GammaUDD[m][j][k] * GammaUDD[l][i][m] * AD[l]
# Covariant divergence
# D_{i} A^{i} = \gamma^{ij} D_{j} A_{i}
DivA = 0
# Gradient of covariant divergence
# DivA_dD_{i} = \gamma^{jk} A_{k;\hat{j}\hat{i}}
DivA_dD = ixp.zerorank1()
# Covariant Laplacian
# LapAD_{i} = \gamma^{jk} A_{i;\hat{j}\hat{k}}
LapAD = ixp.zerorank1()
for i in range(DIM):
for j in range(DIM):
DivA += gammaUU[i][j] * AD_dcovD[i][j]
for k in range(DIM):
DivA_dD[i] += gammaUU[j][k] * AD_dcovDD[k][j][i]
LapAD[i] += gammaUU[j][k] * AD_dcovDD[i][j][k]
# Step 5: Define right-hand sides for the evolution.
ArhsD = ixp.zerorank1()
ErhsD = ixp.zerorank1()
for i in range(DIM):
ArhsD[i] = -ED[i] - psi_dD[i]
ErhsD[i] = -LapAD[i] + DivA_dD[i]
psi_rhs = -DivA
# Step 6: Generate C code for System I Maxwell's evolution equations,
# print output to the screen (standard out, or stdout).
#lhrh_list = []
#for i in range(DIM):
# lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "AD" + str(i)), rhs=ArhsD[i]))
# lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "ED" + str(i)), rhs=ErhsD[i]))
#lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "psi"), rhs=psi_rhs))
#fin.FD_outputC("stdout", lhrh_list)
```
<a id='violation'></a>
# Step 3: Measuring the Constraint violation \[Back to [top](#toc)\]
$$\label{violation}$$
To evaluate our results, we will need to measure how much our simulation differs from what should be physically possible. We will do this with the constraint equation $D_i E^i = 4 \pi \rho_e$; specifically, we will measure the constraint violation
\begin{align}
\mathcal{C} &= D_i E^i - 4 \pi \rho_e \\
&= D_i (\gamma^{ij} E_j) - 4 \pi \rho_e \\
&= \gamma^{ij} D_i E_j + E_j D_i \gamma^{ij} - 4 \pi \rho_e \\
&= \gamma^{ij} D_i E_j,
\end{align}
since the covariant derivative of the metric tensor is $0$ and $\rho_e=0$ in free space. So, $\mathcal{C} = \gamma^{ij} (E_{j,i} - \Gamma^b_{ij} E_b)$, which will be valid for both systems.
```
ED_dD = ixp.declarerank2("ED_dD","nosym")
Cviolation = gri.register_gridfunctions("AUX", ["Cviolation"])
Cviolation = sp.sympify(0)
for i in range(DIM):
for j in range(DIM):
Cviolation += gammaUU[i][j] * ED_dD[j][i]
for b in range(DIM):
Cviolation -= gammaUU[i][j] * GammaUDD[b][i][j] * ED[b]
```
<a id='code_validation_sys1'></a>
# Step 4: Code Validation against `Maxwell.MaxwellCartesian_Evol` NRPy+ Module (System I) \[Back to [top](#toc)\]
$$\label{code_validation_sys1}$$
Here, as a code validation check, we verify agreement in the SymPy expressions for the RHSs of Maxwell's equations (in System I) between
1. this tutorial and
2. the NRPy+ [Maxwell.MaxwellCartesian](../edit/Maxwell/MaxwellCartesian_Evol.py) module.
```
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
# Step 18: Call the MaxwellCartesian_Evol() function from within the
# Maxwell/MaxwellCartesian_Evol.py module,
# which should do exactly the same as in Steps 1-16 above.
import Maxwell.MaxwellCartesian_Evol as mwevol
par.set_parval_from_str("System_to_use","System_I")
mwevol.MaxwellCartesian_Evol()
print("Consistency check between MaxwellCartesian tutorial and NRPy+ module: ALL SHOULD BE ZERO.")
print("psi_rhs - mwevol.psi_rhs = " + str(psi_rhs - mwevol.psi_rhs))
for i in range(DIM):
print("ArhsD["+str(i)+"] - mwevol.ArhsD["+str(i)+"] = " + str(ArhsD[i] - mwevol.ArhsD[i]))
print("ErhsD["+str(i)+"] - mwevol.ErhsD["+str(i)+"] = " + str(ErhsD[i] - mwevol.ErhsD[i]))
print("Cviolation - mwevol.Cviolation = " + str(Cviolation - mwevol.Cviolation))
```
We will now build the equations for System II.
```
# We inherit here all of the definitions from System I, above
# Step 7a: Register the scalar auxiliary variable \Gamma
Gamma = gri.register_gridfunctions("EVOL", ["Gamma"])
# Step 7b: Declare the ordinary gradient \partial_{i} \Gamma
Gamma_dD = ixp.declarerank1("Gamma_dD")
# Step 8a: Construct the second covariant derivative of the scalar \psi
# \psi_{;\hat{i}\hat{j}} = \psi_{,i;\hat{j}}
# = \psi_{,ij} - \Gamma^{k}_{ij} \psi_{,k}
psi_dDD = ixp.declarerank2("psi_dDD", "sym01")
psi_dcovDD = ixp.zerorank2()
for i in range(DIM):
for j in range(DIM):
psi_dcovDD[i][j] = psi_dDD[i][j]
for k in range(DIM):
psi_dcovDD[i][j] += - GammaUDD[k][i][j] * psi_dD[k]
# Step 8b: Construct the covariant Laplacian of \psi
# Lappsi = ghat^{ij} D_{j} D_{i} \psi
Lappsi = 0
for i in range(DIM):
for j in range(DIM):
Lappsi += gammaUU[i][j] * psi_dcovDD[i][j]
# Step 9: Define right-hand sides for the evolution.
ArhsD = ixp.zerorank1()
ErhsD = ixp.zerorank1()
for i in range(DIM):
ArhsD[i] = -ED[i] - psi_dD[i]
ErhsD[i] = -LapAD[i] + Gamma_dD[i]
psi_rhs = -Gamma
Gamma_rhs = -Lappsi
# Step 10: Generate C code for System II Maxwell's evolution equations,
# print output to the screen (standard out, or stdout).
#lhrh_list = []
#for i in range(DIM):
# lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "AD" + str(i)), rhs=ArhsD[i]))
# lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "ED" + str(i)), rhs=ErhsD[i]))
#lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "psi"), rhs=psi_rhs))
#lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "Gamma"), rhs=Gamma_rhs))
#fin.FD_outputC("stdout", lhrh_list)
```
<a id='code_validation_sys2'></a>
# Step 5: Code Validation against `Maxwell.MaxwellCartesian_Evol` NRPy+ Module (System II) \[Back to [top](#toc)\]
$$\label{code_validation_sys2}$$
Here, as a code validation check, we verify agreement in the SymPy expressions for the RHSs of Maxwell's equations (in System II) between
1. this tutorial and
2. the NRPy+ [Maxwell.MaxwellCartesian](../edit/Maxwell/MaxwellCartesian_Evol.py) module.
```
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
# Step 18: Call the MaxwellCartesian_Evol() function from within the
# Maxwell/MaxwellCartesian_Evol.py module,
# which should do exactly the same as in Steps 1-16 above.
par.set_parval_from_str("System_to_use","System_II")
mwevol.MaxwellCartesian_Evol()
print("Consistency check between MaxwellCartesian tutorial and NRPy+ module: ALL SHOULD BE ZERO.")
print("psi_rhs - mwevol.psi_rhs = " + str(psi_rhs - mwevol.psi_rhs))
print("Gamma_rhs - mwevol.Gamma_rhs = " + str(Gamma_rhs - mwevol.Gamma_rhs))
for i in range(DIM):
print("ArhsD["+str(i)+"] - mwevol.ArhsD["+str(i)+"] = " + str(ArhsD[i] - mwevol.ArhsD[i]))
print("ErhsD["+str(i)+"] - mwevol.ErhsD["+str(i)+"] = " + str(ErhsD[i] - mwevol.ErhsD[i]))
print("Cviolation - mwevol.Cviolation = " + str(Cviolation - mwevol.Cviolation))
```
<a id='id'></a>
# Step 6: Constructing the Initial Data \[Back to [top](#toc)\]
$$\label{id}$$
Now that we have evolution equations in place, we must construct the initial data that will be evolved by the solver of our choice. We will start from the analytic solution to this system of equations, given in [Illustrating Stability Properties of Numerical Relativity in Electrodynamics](https://arxiv.org/abs/gr-qc/0201051) as
\begin{align}
A^{\hat{\phi}} &= \mathcal{A} \sin \theta \left( \frac{e^{-\lambda v^2}-e^{-\lambda u^2}}{r^2} - 2 \lambda \frac{ve^{-\lambda v^2}-ue^{-\lambda u^2}}{r} \right), \\
\end{align}
for vanishing scalar potential $\psi$, where $\mathcal{A}$ gives the amplitude, $\lambda$ describes the size of the wavepacket, $u = t+r$, and $v = t-r$. Other components of this field are $0$.To get initial data, then, we simply set $t=0$; since $\psi=0$, $E_i = \partial_t A_i$. Thus, our initial data becomes the equations
\begin{align}
A^{\hat{\phi}} &= 0 \\
E^{\hat{\phi}} &= 8 \mathcal{A} r \sin \theta \lambda^2 e^{-\lambda r^2} \\
\psi &= 0
\end{align}
where the non-$\hat{\phi}$ components are set to 0. We still will need to convert $E^i$ in spherical-like coordinates and then lower its index. Using the standard transformations for coordinates and unit vectors,
\begin{align}
E^{\hat{x}} &= -\frac{y E^{\hat{\phi}}(x,y,z)}{\sqrt{x^2+y^2}} \\
E^{\hat{y}} &= \frac{x E^{\hat{\phi}}(x,y,z)}{\sqrt{x^2+y^2}} \\
E^{\hat{z}} &= 0. \\
\end{align}
We can lower the index in the usual way.
For system II, we will also need to set initial data for $\Gamma$. Since $\Gamma = -\partial_t \psi$ and we have chosen $\psi(t=0) = 0$, $\Gamma(t=0) = 0$.
```
# Step 1: Declare free parameters intrinsic to these initial data
amp,lam = par.Cparameters("REAL",thismodule,
["amp","lam"],
[1.0,1.0]) # __name__ = "MaxwellCartesian_ID", this module's name
# Step 2: Set the initial data
AidD = ixp.zerorank1()
EidD = ixp.zerorank1()
EidU = ixp.zerorank1()
# Set the coordinate transformations:
radial = sp.sqrt(x*x + y*y + z*z)
polar = sp.atan2(sp.sqrt(x*x + y*y),z)
EU_phi = 8*amp*radial*sp.sin(polar)*lam*lam*sp.exp(-lam*radial*radial)
EidU[0] = -(y * EU_phi)/sp.sqrt(x*x + y*y)
EidU[1] = (x * EU_phi)/sp.sqrt(x*x + y*y)
# The z component (2)is zero.
for i in range(DIM):
for j in range(DIM):
EidD[i] += gammaDD[i][j] * EidU[j]
psi_ID = sp.sympify(0)
Gamma_ID = sp.sympify(0)
```
<a id='code_validation'></a>
# Step 7: Code Validation against `Maxwell.MaxwellCartesian_ID` NRPy+ Module \[Back to [top](#toc)\]
$$\label{code_validation}$$
Here, as a code validation check, we verify agreement in the SymPy expressions for the initial data we intend to use between
1. this tutorial and
2. the NRPy+ [Maxwell.MaxwellCartesian_ID](../edit/Maxwell/MaxwellCartesian_ID.py) module.
Since the initial data is identical between the two systems for $E_i$, $A_i$, and $\psi$, so checking system I should be redundant; we will do it anyways, to be sure.
```
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
par.set_parval_from_str("System_to_use","System_I")
import Maxwell.MaxwellCartesian_ID as mwid
mwid.MaxwellCartesian_ID()
print("System I consistency check between MaxwellCartesian tutorial and NRPy+ module;\n ALL SHOULD BE ZERO:")
print("psi_ID - mwid.psi_ID = " + str(psi_ID - mwid.psi_ID))
for i in range(DIM):
print("AidD["+str(i)+"] - mwid.AidD["+str(i)+"] = " + str(AidD[i] - mwid.AidD[i]))
print("EidD["+str(i)+"] - mwid.EidD["+str(i)+"] = " + str(EidD[i] - mwid.EidD[i]))
```
Finally, we will repeat the check with system II initial data.
```
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
par.set_parval_from_str("System_to_use","System_II")
mwid.MaxwellCartesian_ID()
print("System II consistency check between MaxwellCartesian tutorial and NRPy+ module;\n ALL SHOULD BE ZERO:")
print("psi_ID - mwid.psi_ID = " + str(psi_ID - mwid.psi_ID))
print("Gamma_ID - mwid.Gamma_ID = " + str(Gamma_ID - mwid.Gamma_ID))
for i in range(DIM):
print("AidD["+str(i)+"] - mwid.AidD["+str(i)+"] = " + str(AidD[i] - mwid.AidD[i]))
print("EidD["+str(i)+"] - mwid.EidD["+str(i)+"] = " + str(EidD[i] - mwid.EidD[i]))
```
<a id='latex_pdf_output'></a>
# Step 8: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-MaxwellCartesian.pdf](Tutorial-MaxwellCartesian.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-MaxwellCartesian")
```
| github_jupyter |
```
import csv
import pandas as pd
import os
import scipy.stats
import numpy as np
from datetime import date,timedelta,datetime
def read_data(file):
df = pd.read_csv(file)
df = pd.DataFrame(df)
return df
def mofunc(row):
if row['Severity'] > 0.8 or row['Hazard_Score'] > 80:
return 'Warning'
elif 0.6 < row['Severity'] < 0.80 or 60 < row['Hazard_Score'] < 80:
return 'Watch'
elif 0.35 < row['Severity'] < 0.6 or 35 < row['Hazard_Score'] < 60:
return 'Advisory'
elif 0 < row['Severity'] < 0.35 or 0 < row['Hazard_Score'] < 35:
return 'Information'
forcast_date = date.today()
cur_year, cur_month,cur_day = map(str,[forcast_date.today().year,forcast_date.today().month,forcast_date.today().day])
cur_month = cur_month.zfill(2)
cur_day=cur_day.zfill(2)
MOMOutput='Final_Attributes_'+cur_year+cur_month+str(int(cur_day)-1)+'18.csv'
VIIRS="VIIRS_Flood_"+cur_year+cur_month+str(int(cur_day)-1)+'.csv'
#MOMOutput='Final_Attributes_20210701_DFOUpdated.csv'
#VIIRS="VIIRS_Flood_20210701.csv"
weightage = read_data('VIIRS_Weightages.csv')
Attributes=read_data('Attributes.csv')
PDC_resilience = read_data('Copy of Resilience_Index.csv')
add_field_VIIRS=['VIIRS_area_1day_score', 'VIIRS_percarea_1day_score', 'VIIRS_area_5day_score', 'VIIRS_percarea_5day_score','VIIRSTotal_Score']
#Read VIIRS Processing data and calculate score
with open(VIIRS, 'r', encoding='UTF-8') as VIIRS_file:
VIIRS_reader = csv.reader(VIIRS_file)
csvfile = open('VIIRS_w_score.csv', 'w', newline='\n', encoding='utf-8')
VIIRS_w_score = csv.writer(csvfile)
row_count = 1
# csv_writer = csv.writer(write_obj)
for row in VIIRS_reader:
if row_count == 1:
for x in add_field_VIIRS:
row.append(x)
row_count = row_count + 1
else:
if float(row[1]) / float(weightage.VIIRS_Area_wt) > float(weightage.VIIRS_Area_max_pt):
VIIRS_area_1day_score = str(float(weightage.VIIRS_Area_max_pt)*float(weightage.one_Day_Multiplier))
else:
VIIRS_area_1day_score = str(float(weightage.VIIRS_Area_Min_pt) * float(weightage.one_Day_Multiplier)* float(row[1]) / float(weightage.VIIRS_Area_wt))
if float(row[2]) / float(weightage.VIIRS_percArea_wt) > float(weightage.VIIRS_percArea_Maxpt):
VIIRS_perc_area_1day_score = str(float(weightage.VIIRS_percArea_Maxpt)*float(weightage.one_Day_Multiplier))
else:
VIIRS_perc_area_1day_score = str(float(weightage.VIIRS_percArea_Minpt)*float(weightage.one_Day_Multiplier)* float(row[2]) / float(weightage.VIIRS_percArea_wt))
if float(row[3]) / float(weightage.VIIRS_Area_wt) > float(weightage.VIIRS_Area_max_pt):
VIIRS_area_5day_score = str(float(weightage.VIIRS_Area_max_pt)*float(weightage.five_Day_Multiplier))
else:
VIIRS_area_5day_score = str(float(weightage.VIIRS_Area_Min_pt) * float(weightage.five_Day_Multiplier)* float(row[3]) / float(weightage.VIIRS_Area_wt))
if float(row[4]) / float(weightage.VIIRS_percArea_wt) > float(weightage.VIIRS_percArea_Maxpt):
VIIRS_perc_area_5day_score = str(float(weightage.VIIRS_percArea_Maxpt)*float(weightage.five_Day_Multiplier))
else:
VIIRS_perc_area_5day_score = str(float(weightage.VIIRS_percArea_Minpt)*float(weightage.five_Day_Multiplier)* float(row[4]) / float(weightage.VIIRS_percArea_wt))
Sum_Score = str(
(float(VIIRS_area_1day_score) + float(VIIRS_perc_area_1day_score) + float(VIIRS_area_5day_score) + float(VIIRS_perc_area_5day_score)))
score_field = [VIIRS_area_1day_score, VIIRS_perc_area_1day_score, VIIRS_area_5day_score, VIIRS_perc_area_5day_score, Sum_Score]
for x in score_field:
row.append(x)
VIIRS_w_score.writerow(row)
csvfile.close()
VIIRS = read_data('VIIRS_w_score.csv')
VIIRS = VIIRS[VIIRS.VIIRSTotal_Score > 0.1]
MOM = read_data(MOMOutput)
MOM.drop(columns=['area_km2','ISO','Admin0','Admin1','rfr_score','cfr_score','Resilience_Index',' NormalizedLackofResilience ','Severity','Alert'], inplace=True)
Final_Output_0= pd.merge(MOM.set_index('pfaf_id'), VIIRS.set_index('pfaf_id'), on='pfaf_id', how='outer')
join1 = pd.merge(Attributes, PDC_resilience[['ISO', 'Resilience_Index', ' NormalizedLackofResilience ']], on='ISO', how='inner')
Final_Output=pd.merge(join1.set_index('pfaf_id'), Final_Output_0, on='pfaf_id', how='right')
Final_Output[['Hazard_Score']] = Final_Output[['Hazard_Score']].fillna(value=0)
Final_Output.loc[(Final_Output['Hazard_Score']<Final_Output['VIIRSTotal_Score']),'Flag']=3
Final_Output['Hazard_Score'] =Final_Output[['Hazard_Score', 'VIIRSTotal_Score']].max(axis=1)
Final_Output = Final_Output[Final_Output.Hazard_Score != 0]
Final_Output.drop(Final_Output.index[(Final_Output['rfr_score']==0) & (Final_Output['cfr_score']==0)], inplace=True)
Final_Output = Final_Output.assign(
Scaled_Riverine_Risk=lambda x: Final_Output['rfr_score'] * 20)
Final_Output = Final_Output.assign(
Scaled_Coastal_Risk=lambda x: Final_Output['cfr_score'] * 20)
Final_Output = Final_Output.assign(
Severity=lambda x: scipy.stats.norm(np.log(100 - Final_Output[['Scaled_Riverine_Risk', 'Scaled_Coastal_Risk']].max(axis=1)), 1).cdf(
np.log(Final_Output['Hazard_Score'])))
Final_Output['Alert'] = Final_Output.apply(mofunc, axis=1)
Final_Output.loc[Final_Output['Alert']=="Information",'Flag']=''
Final_Output.loc[Final_Output['Alert']=="Advisory",'Flag']=''
Final_Output.to_csv('Final_Attributes_'+cur_year+cur_month+str(float(cur_day)-1)+'18_VIIRSUpdated.csv', encoding='utf-8-sig')
#Final_Output.to_csv('Final_Attributes_20210701_HWRFDFOandVIIRSUpdated.csv', encoding='utf-8-sig')
join1 = pd.merge(Attributes, PDC_resilience[['ISO', 'Resilience_Index', ' NormalizedLackofResilience ']], on='ISO', how='inner')
Attributes_Clean_VIIRS_Updated = pd.merge(join1.set_index('pfaf_id'), Final_Output[['Alert','Flag']], on='pfaf_id', how='right')
Attributes_Clean_VIIRS_Updated.to_csv('Attributes_Clean'+cur_year+cur_month+str(float(cur_day)-1)+'18_VIIRSUpdated.csv', encoding='utf-8-sig')
os.remove('VIIRS_w_score.csv')
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/gdrive')
!pip install -q efficientnet
import math, re, os
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import tensorflow as tf
import tensorflow_probability as tfp
import tensorflow.keras.layers as L
import tensorflow.keras.backend as K
import efficientnet.tfkeras as efn
from sklearn import metrics
from sklearn.model_selection import train_test_split
import random
from sklearn.model_selection import GroupKFold
import pickle
# Detect hardware, return appropriate distribution strategy
try:
# TPU detection. No parameters necessary if TPU_NAME environment variable is
# set: this is always the case on Kaggle.
tpu = tf.distribute.cluster_resolver.TPUClusterResolver()
print('Running on TPU ', tpu.master())
except ValueError:
tpu = None
if tpu:
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)
strategy = tf.distribute.experimental.TPUStrategy(tpu)
else:
# Default distribution strategy in Tensorflow. Works on CPU and single GPU.
strategy = tf.distribute.get_strategy()
print("REPLICAS: ", strategy.num_replicas_in_sync)
EFNS = [efn.EfficientNetB0,efn.EfficientNetB1,efn.EfficientNetB2,efn.EfficientNetB3,
efn.EfficientNetB4,efn.EfficientNetB5,efn.EfficientNetB6,efn.EfficientNetB7]
def get_model_v1(IMAGE_SIZE, NUM_CLASSES, EMB_SIZE, EFF_VER, order=0,weight_path=None):
class ArcMarginProduct_v2(tf.keras.layers.Layer):
def __init__(self, num_classes):
super(ArcMarginProduct_v2, self).__init__()
self.num_classes= num_classes
def build(self, input_shape):
self.w = self.add_variable(
"weights", shape=[int(input_shape[-1]), self.num_classes])
def call(self, input):
cosine = tf.matmul(tf.nn.l2_normalize(input, axis=1), tf.nn.l2_normalize(self.w, axis=0))
return cosine
def getefn():
pretrained_model = EFNS[EFF_VER](weights=None, include_top=False ,input_shape=[*IMAGE_SIZE, 3])
pretrained_model.trainable = True
return pretrained_model
def ArcFaceResNet():
x= inputs = tf.keras.Input([*IMAGE_SIZE, 3])
x = getefn()(x)
x = L.GlobalAveragePooling2D()(x)
x = L.Dense(EMB_SIZE, activation='swish')(x)
target = ArcMarginProduct_v2(NUM_CLASSES)(x)
model = tf.keras.Model(inputs, target)
model.get_layer('efficientnet-b'+str(EFF_VER))._name='efficientnet-b'+str(EFF_VER)+str(order)
return model
model = ArcFaceResNet()
model.summary()
if weight_path is not None:
model.load_weights(weight_path)
return model
#single model
model = get_model_v1([640,640],203094,512,6,1,'/content/gdrive/My Drive/eff6_640_notclean_0.5_1.1931.hdf5')
_model= tf.keras.Model(inputs= model.input,
outputs =model.get_layer('dense').output)
def export_model_v1(model, outdir):
@tf.function(input_signature=[
tf.TensorSpec(
shape=[None, None, 3],
dtype=tf.uint8,
name='input_image')
])
def serving(input_image):
image = tf.image.resize(input_image, [640,640])
image -= tf.constant([0.485 * 255, 0.456 * 255, 0.406 * 255]) # RGB
image /= tf.constant([0.229 * 255, 0.224 * 255, 0.225 * 255]) # RGB
image = tf.reshape(image, [640,640,3])
outputs = model(image[tf.newaxis])
features = tf.math.l2_normalize(outputs[0])
return {
'global_descriptor': tf.identity(features, name='global_descriptor')
}
tf.saved_model.save(
obj=model,
export_dir=outdir,
signatures={'serving_default': serving})
export_model_v1(_model,'/content/gdrive/My Drive/landmark_export_model/eff6_640_notclean05_11931')
#ensemble
model1 = get_model_v1([640,640],203094,512,7,1,'/content/gdrive/My Drive/eff7_640_notclean_0.5_1.1411.hdf5')
model2 = get_model_v1([640,640],203094,512,6,2,'/content/gdrive/My Drive/eff6_640_notclean_0.5_1.1931.hdf5')
model3 = get_model_v1([640,640],203094,512,7,3,'/content/gdrive/My Drive/landmark/eff7_512_ver2_10293_NotClean_0.5_640/shuffle_weights.epoch44.loss1.5736.valid_loss1.2554.hdf5')
model4 = get_model_v1([512,512],203094,512,7,4,'/content/gdrive/My Drive/eff7_512_ver1_notclean0.5_1.2580.hdf5')
_model= tf.keras.Model(inputs= [model1.input,
model2.input,
model3.input,
model4.input,
],
outputs =[model1.get_layer('dense').output,
model2.get_layer('dense_1').output,
model3.get_layer('dense_2').output,
model4.get_layer('dense_3').output,
])
def export_model_v1(model, outdir):
@tf.function(input_signature=[
tf.TensorSpec(
shape=[None, None, 3],
dtype=tf.uint8,
name='input_image')
])
def serving(input_image):
image2 = tf.image.resize(input_image, [640,640])
image2 -= tf.constant([0.485 * 255, 0.456 * 255, 0.406 * 255]) # RGB
image2 /= tf.constant([0.229 * 255, 0.224 * 255, 0.225 * 255]) # RGB
image2 = tf.reshape(image2, [640,640,3])
image3 = tf.image.resize(input_image, [512,512])
image3 -= tf.constant([0.485 * 255, 0.456 * 255, 0.406 * 255]) # RGB
image3 /= tf.constant([0.229 * 255, 0.224 * 255, 0.225 * 255]) # RGB
image3 = tf.reshape(image3, [512,512,3])
outputs = model((image2[tf.newaxis],image2[tf.newaxis],image2[tf.newaxis],image3[tf.newaxis]))
output1 = tf.math.l2_normalize(outputs[0][0])
output2 = 0.8*tf.math.l2_normalize(outputs[1][0])
output3 = 0.55*tf.math.l2_normalize(outputs[2][0])
output4 = 0.5*tf.math.l2_normalize(outputs[3][0])
features = tf.concat([output1,output2,output3, output4],axis=-1)
return {
'global_descriptor': tf.identity(features, name='global_descriptor')
}
tf.saved_model.save(
obj=model,
export_dir=outdir,
signatures={'serving_default': serving})
export_model_v1(_model,'/content/gdrive/My Drive/landmark_export_model/0816_notclean05_640_776_512_7')
```
| github_jupyter |
```
%cd ..
import pandas as pd
import pickle
import numpy as np
import matplotlib.pyplot as plt
import sys
sys.path.append(".")
from src.factory import *
from src.utils import *
from sklearn.metrics import log_loss
DATADIR = Path("../input/rsna-str-pulmonary-embolism-detection/")
train = pd.read_csv(DATADIR / "train.csv")
pre = pd.read_csv(DATADIR / "split.csv")
train = train.merge(pre, on="StudyInstanceUID")
portion = pd.read_csv(DATADIR / "study_pos_portion.csv")
train = train.merge(portion, on="StudyInstanceUID")
z_pos_df = pd.read_csv(DATADIR / "sop_to_prefix.csv").rename(columns={'img_prefix': 'z_pos'})
train = train.merge(z_pos_df, on="SOPInstanceUID")
### train = train.query("fold == 0 or fold == 1") # now I have fold0,1 only
studies = train.StudyInstanceUID.unique()
def get_pred(_path):
res = load_pickle(_path)
raw_pred = pd.DataFrame({
"SOPInstanceUID": res["ids"],
**res["outputs"],
})
return raw_pred
def calib_p(arr, factor): # set factor>1 to enhance positive prob
return arr * factor / (arr * factor + (1-arr))
# check
# oof_f0 = get_pred("output_yuji/b3_non_weight//oof_fold0_ep0.pkl")
# plt.hist( oof_f0.pe_present_on_image, bins=300 )
# plt.show()
# ! SET claibration value. Run src/oof_optpy to calculate the value
oof_f0, fold0_calib_f = get_pred("output/final_image_level/oof_fold0.pkl"), 3.8250639579850194
oof_f1, fold1_calib_f = get_pred("output/final_image_level/oof_fold1.pkl"), 8.555037588568537
oof_f2, fold2_calib_f = get_pred("output/final_image_level/oof_fold2.pkl"), 4.374239635034443
oof_f3, fold3_calib_f = get_pred("output/final_image_level/oof_fold3.pkl"), 7.480972390526775
oof_f4, fold4_calib_f = get_pred("output/final_image_level/oof_fold4.pkl"), 5.002262078458348
# BAD
if False: # pick best one which yields weighted-logloss after calib
oof_f3, fold3_calib_f = get_pred("output/035_pe_present___448___apex/fold3_ep0.pt.valid.pickle"), 6.541753595870311
oof_f4, fold4_calib_f = get_pred("output/035_pe_present___448___apex/fold4_ep0.pt.valid.pickle"), 3.8250639579850194
if True: ### ==== do calib for each fold
oof_f0["pe_present_on_image"] = calib_p(oof_f0["pe_present_on_image"], fold0_calib_f)
oof_f1["pe_present_on_image"] = calib_p(oof_f1["pe_present_on_image"], fold1_calib_f)
oof_f2["pe_present_on_image"] = calib_p(oof_f2["pe_present_on_image"], fold2_calib_f)
oof_f3["pe_present_on_image"] = calib_p(oof_f3["pe_present_on_image"], fold3_calib_f)
oof_f4["pe_present_on_image"] = calib_p(oof_f4["pe_present_on_image"], fold4_calib_f)
oof = pd.concat([oof_f0, oof_f1, oof_f2, oof_f3, oof_f4]).rename(columns={'pe_present_on_image': 'pred0'})
train = train.merge(oof[['pred0', 'SOPInstanceUID']], on="SOPInstanceUID") # add pred
train_copyed = train.copy()
train.columns
""" feature engineer """
train = train.sort_values(['StudyInstanceUID', 'z_pos'])
train_current_z_pos = train.groupby('StudyInstanceUID')['z_pos'].shift(0)
### for i in range(1, 20):
for i in range(1, 10):
train[f'pred0_pre{i}'] = train.groupby('StudyInstanceUID')['pred0'].shift(i)
train[f'pred0_post{i}'] = train.groupby('StudyInstanceUID')['pred0'].shift(-i)
# NORMALIZED Z POS
z_max = train.groupby('StudyInstanceUID').z_pos.max().rename('z_pos_max')
train = train.merge(z_max, on='StudyInstanceUID')
train['z_pos_norm'] = train['z_pos'] / train['z_pos_max']
train = train.drop('z_pos_max', axis=1)
train.tail()
ids = [c for c in list(train) if 'UID' in c]
targets = [
'negative_exam_for_pe',
'indeterminate',
'chronic_pe',
'acute_and_chronic_pe',
'central_pe',
'leftsided_pe',
'rightsided_pe',
'rv_lv_ratio_gte_1',
'rv_lv_ratio_lt_1',
]
other_targets = [c for c in list(train) if 'pe_present_on_image' in c]
### remove_cols = ['fold', 'path', 'weight', 'qa_contrast', 'qa_motion'] + targets + ids + other_targets
### remove_cols = ['fold', 'path', 'weight', 'qa_contrast', 'qa_motion'] + ['exam_type','flow_artifact','pe_present_portion', 'true_filling_defect_not_pe'] + targets + ids + other_targets
remove_cols = ['fold', 'path', 'weight', 'qa_contrast', 'qa_motion'] + ['exam_type','flow_artifact','pe_present_portion', 'true_filling_defect_not_pe'] + targets + ids + other_targets + ['z_pos']
features = sorted(list(set(list(train)) - set(remove_cols)))
print(features)
features_copyed = features.copy()
def fobj(pred, data):
true = data.get_label()
label = 2*true - 1
weights = data.weights
response = -label / (1 + np.exp(label * pred))
abs_response = np.abs(response)
grad = response
hess = abs_response * (1 - abs_response)
return grad*weights, hess*weights
import torch
bce_func = torch.nn.BCELoss(reduction='none')
def feval2(preds, data):
scores = bce_func(torch.FloatTensor(preds), torch.FloatTensor(data.label))
scores = scores * torch.FloatTensor(data.weights)
return 'weighted logloss', torch.mean(scores), False
import torch
bce_func_logit = torch.nn.BCEWithLogitsLoss(reduction='none')
def feval(preds, data):
scores = bce_func_logit(torch.FloatTensor(preds), torch.FloatTensor(data.label))
scores = scores * torch.FloatTensor(data.weights)
return 'weighted logloss', torch.mean(scores), False
import torch
def sigmoid(x):
return 1 / (1 + np.exp(-x))
# features = ['pred'] # for test
features = features_copyed
import lightgbm as lgb
import numpy as np
from sklearn.metrics import roc_auc_score, average_precision_score
import warnings
warnings.simplefilter('ignore')
import pickle
oof_preds_list = []
models_list = []
target = 'pe_present_on_image'
for i in range(1):
print(f'=================={i}================')
if i % 4 == 0:
params = {'boosting_type': 'gbdt',
'objective': 'binary',
# 'metric': 'None',
'subsample': 0.75,
'subsample_freq': 1,
'learning_rate': 0.1,
'feature_fraction': 0.9,
'max_depth': 15,
'lambda_l1': 1,
'lambda_l2': 1,
'verbose': 100,
'early_stopping_rounds': 100,
'verbose': -1,
}
elif i % 4 == 1:
params = {
'max_depth': 4,
'max_leave': int(0.2 * 2 ** 4),
'reg_lambda': 1,
'reg_alpha': 1,
'subsamples': 0.8,
'colsample_bytree': 0.7,
'objective': 'binary',
'min_data_in_leaf': 0,
'boosting': 'gbdt',
'metric': 'None',
'learning_rate': 0.1,
}
elif i % 4 == 2:
params = {
'num_leaves': 19,
'min_data_in_leaf': 160,
'min_child_weight': 0.03,
'bagging_fraction' : 0.7,
'feature_fraction' : 0.8,
'learning_rate' : 0.1,
'max_depth': -1,
'reg_alpha': 0.02,
'reg_lambda': 0.12,
'objective': 'binary',
'verbose': 100,
'boost_from_average': False,
'metric': 'None',
}
else:
params = {
'objective': "binary",
'metric': 'None',
'boost_from_average': "false",
'tree_learner': "serial",
'max_depth': -1,
'learning_rate': 0.1,
'num_leaves': 197,
'feature_fraction': 0.3,
'bagging_freq': 1,
'bagging_fraction': 0.7,
'min_data_in_leaf': 100,
'bagging_seed': 11,
'max_bin': 255,
'verbosity': -1}
oof_preds = np.zeros(train.shape[0])
val_results = {}
models = []
params['random_state'] = i
iter = 100000
# for n_fold, (trn_idx, val_idx) in enumerate(kf.split(train, train[target])):
for n_fold in range(5):
### for n_fold in range(2):
print(f' ==============fold{n_fold}================')
tr = train.query(f'fold != {n_fold}')
val = train.query(f'fold == {n_fold}')
trn_data = lgb.Dataset(tr[features], label=tr[target])
trn_data.weights = tr.pe_present_portion.values
val_data = lgb.Dataset(val[features], label=val[target])
val_data.weights = val.pe_present_portion.values
clf = lgb.train(params, trn_data, num_boost_round=iter, valid_sets=[trn_data, val_data], valid_names=['train', 'val'],
# verbose_eval=200, early_stopping_rounds = 10/params['learning_rate'], evals_result=val_results,)
feval=feval, fobj = fobj, verbose_eval=2000, early_stopping_rounds = 10/params['learning_rate'], evals_result=val_results, )
file = f'lgbs/lgb_seed{i}_fold{n_fold}.pkl'
pickle.dump(clf, open(file, 'wb'))
oof_preds[train.fold==n_fold] = clf.predict(val[features])
oof_preds_list.append(oof_preds)
print(f'-------------------------------------------------------------------------roc_auc: {roc_auc_score(train[target], np.mean(oof_preds_list, axis=0))}')
print(f'----------------------------------------------------------roc_auc using raw pred: {roc_auc_score(train[target], train["pred0"])}')
print(f'------------------------------------------------------------------------------AP: {average_precision_score(train[target], np.mean(oof_preds_list, axis=0))}')
print(f'---------------------------------------------------------------AP using raw pred: {average_precision_score(train[target], train["pred0"])}')
lgb_oof = np.mean(oof_preds_list, axis=0)
train['lgb_preds'] = sigmoid(lgb_oof)
bce_func = torch.nn.BCELoss(reduction='none')
lgb_losses = bce_func(torch.FloatTensor(sigmoid(lgb_oof)), torch.FloatTensor(train['pe_present_on_image']))
### torch.mean(lgb_losses*train['weight'].values)
torch.mean(lgb_losses*train['pe_present_portion'].values).item()
# no stacking result
lgb_losses = bce_func(torch.FloatTensor(train['pred0']), torch.FloatTensor(train['pe_present_on_image']))
torch.mean(lgb_losses*train['pe_present_portion'].values).item()
raise "BELOW IS NOT USED FOR FINAL SUB"
```
# <<< BELOW IS NOT USED FOR FINAL SUB >>> stacking for PE_REPESNT -> POS_EXAM
```
DATADIR = Path("../input/rsna-str-pulmonary-embolism-detection/")
train = pd.read_csv(DATADIR / "train.csv")
pre = pd.read_csv(DATADIR / "split.csv")
train = train.merge(pre, on="StudyInstanceUID")
portion = pd.read_csv(DATADIR / "study_pos_portion.csv")
train = train.merge(portion, on="StudyInstanceUID")
z_pos_df = pd.read_csv(DATADIR / "sop_to_prefix.csv").rename(columns={'img_prefix': 'z_pos'})
train = train.merge(z_pos_df, on="SOPInstanceUID")
studies = train.StudyInstanceUID.unique()
oof = pd.concat([oof_f0, oof_f1, oof_f2, oof_f3, oof_f4]).rename(columns={'pe_present_on_image': 'pred'})
train = train.merge(oof[['pred', 'SOPInstanceUID']], on="SOPInstanceUID") # add pred
from functools import partial
def grouping(df):
grouped = pd.DataFrame(df.groupby('StudyInstanceUID')['pred'].mean())
grouped = grouped.rename(columns={'pred': 'mean'})
count = df.groupby('StudyInstanceUID')['pred'].count()
grouped['count_total'] = count
for i in range(1,10):
count = df[df.pred>i/10].groupby('StudyInstanceUID')['pred'].count()
grouped[f'count_over{i/10}'] = count
grouped[f'count_over{i/10}_ratio'] = count / grouped['count_total']
for q in [30, 50, 70, 80, 90, 95, 99]:
# for q in [95]:
grouped[f'percentile{q}'] = df.groupby('StudyInstanceUID')['pred'].apply(lambda arr: np.percentile(arr, q))
ma = pd.DataFrame(df.groupby('StudyInstanceUID')['pred'].max())
grouped['max'] = ma.pred
grouped = grouped.reset_index().fillna(0)
return grouped
train_grouped = grouping(train)
train_grouped['fold'] = train.groupby('StudyInstanceUID')['fold'].first().values
train_grouped['negative_exam_for_pe'] = train.groupby('StudyInstanceUID')['negative_exam_for_pe'].first().values
train_grouped['positive_exam_for_pe'] = (1 - train.groupby('StudyInstanceUID')['negative_exam_for_pe'].first().values) * (1 - train.groupby('StudyInstanceUID')['indeterminate'].first().values)
# test_grouped = grouping(test)
# features = list(set(list(train_grouped)) - set(['StudyInstanceUID', 'negative_exam_for_pe']))
# target = 'negative_exam_for_pe'
### features = list(set(list(train_grouped)) - set(['StudyInstanceUID', 'positive_exam_for_pe', 'negative_exam_for_pe', 'fold']))
features = list(set(list(train_grouped)) - set(['StudyInstanceUID', 'positive_exam_for_pe', 'negative_exam_for_pe', 'fold']) - set(['count_total']))
features = sorted(features)
target = 'positive_exam_for_pe'
# target = 'negative_exam_for_pe'
print(features)
train_grouped.head()
import lightgbm as lgb
import numpy as np
from sklearn.metrics import roc_auc_score
import warnings
warnings.simplefilter('ignore')
import pickle
from sklearn.model_selection import KFold, StratifiedKFold
oof_preds_list = []
test_preds_list = []
models_list = []
for i in range(1):
print(f'=================={i}================')
if i % 4 == 0:
params = {'boosting_type': 'gbdt',
'objective': 'binary',
# 'metric': 'None',
'subsample': 0.75,
'subsample_freq': 1,
'learning_rate': 0.1,
'feature_fraction': 0.9,
'max_depth': 15,
'lambda_l1': 1,
'lambda_l2': 1,
'verbose': 100,
'early_stopping_rounds': 100,
'verbose': -1,
}
elif i % 4 == 1:
params = {
'max_depth': 4,
'max_leave': int(0.2 * 2 ** 4),
'reg_lambda': 1,
'reg_alpha': 1,
'subsamples': 0.8,
'colsample_bytree': 0.7,
'objective': 'binary',
'min_data_in_leaf': 0,
'boosting': 'gbdt',
# 'metric': 'None',
'learning_rate': 0.1,
}
elif i % 4 == 2:
params = {
'num_leaves': 19,
'min_data_in_leaf': 160,
'min_child_weight': 0.03,
'bagging_fraction' : 0.7,
'feature_fraction' : 0.8,
'learning_rate' : 0.1,
'max_depth': -1,
'reg_alpha': 0.02,
'reg_lambda': 0.12,
'objective': 'binary',
'verbose': 100,
'boost_from_average': False,
# 'metric': 'None',
}
else:
params = {
'objective': "binary",
# 'metric': 'None',
'boost_from_average': "false",
'tree_learner': "serial",
'max_depth': -1,
'learning_rate': 0.1,
'num_leaves': 197,
'feature_fraction': 0.3,
'bagging_freq': 1,
'bagging_fraction': 0.7,
'min_data_in_leaf': 100,
'bagging_seed': 11,
'max_bin': 255,
'verbosity': -1}
oof_preds = np.zeros(train_grouped.shape[0])
### test_preds = np.zeros(test_grouped.shape[0])
val_results = {}
models = []
params['random_state'] = i
iter = 100000
kf = KFold(n_splits=5, shuffle=True, random_state=72)
#for n_fold, (trn_idx, val_idx) in enumerate(kf.split(train_grouped, train_grouped[target])):
# tr = train_grouped.iloc[trn_idx]
# val = train_grouped.iloc[val_idx]
for n_fold in range( 5 ):
tr = train_grouped[train_grouped.fold != n_fold]
val = train_grouped[train_grouped.fold == n_fold]
trn_data = lgb.Dataset(tr[features], label=tr[target])
val_data = lgb.Dataset(val[features], label=val[target])
clf = lgb.train(params, trn_data, num_boost_round=iter, valid_sets=[trn_data, val_data], valid_names=['train', 'val'],
# feval=feval, verbose_eval=10, early_stopping_rounds = 10/params['learning_rate'], evals_result=val_results,)
verbose_eval=200, early_stopping_rounds = 10/params['learning_rate'], evals_result=val_results,)
file = f'lgbs/posexam_lgb_seed{i}_fold{n_fold}.pkl'
pickle.dump(clf, open(file, 'wb'))
models.append(clf)
oof_preds[train_grouped.fold == n_fold] = clf.predict(val[features])
### oof_preds[val_idx] = clf.predict(val[features])
### test_preds += clf.predict(test_grouped[features]) / 5
# models_list.append(models)
oof_preds_list.append(oof_preds)
###test_preds_list.append(test_preds)
print(f'--------------------------------------------------------------------roc: {roc_auc_score(train_grouped[target], np.mean(oof_preds_list, axis=0))}')
print(f'----------------------------------------------------------------logloss: {log_loss(train_grouped[target], np.mean(oof_preds_list, axis=0))}')
lgb_oof_exam = np.mean(oof_preds_list, axis=0)
### lgb_preds = np.mean(test_preds_list, axis=0)
bce_func = torch.nn.BCELoss(reduction='mean')
lgb_losses = bce_func(torch.FloatTensor(oof_preds), torch.FloatTensor(train_grouped['positive_exam_for_pe']))
torch.mean(lgb_losses).item()
bce_func = torch.nn.BCELoss(reduction='mean')
lgb_losses = bce_func(
( 1 - torch.FloatTensor(oof_preds) ) * (4911) / (4911 + 157),
torch.FloatTensor(train_grouped['negative_exam_for_pe']))
torch.mean(lgb_losses).item()
# # current yama's pipeline for fold0-ep1
# def calib_p(arr, factor): # set factor>1 to enhance positive prob
# return arr * factor / (arr * factor + (1-arr))
# def post_yama(arr):
# return calib_p( np.percentile(arr, 95), factor=1/8.5550)
# lgb_losses = bce_func(torch.FloatTensor(train[['StudyInstanceUID','pred']].groupby('StudyInstanceUID').apply(post_yama)), torch.FloatTensor(train_grouped['positive_exam_for_pe'])).item()
# lgb_losses
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import json
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
```
# Coordinate Ascent - AUC only
```
coats_df_auc_only = pd.read_csv("../output_data/coordinate_ascent_run_AUConly.csv")
coats_df_auc_only[coats_df_auc_only["loss"] == coats_df_auc_only["loss"].max()].index[0]
coats_df_auc_only.plot("Unnamed: 0", "loss")
max_loss = max(coats_df_auc_only["loss"])
max_loss_iteration = coats_df_auc_only[coats_df_auc_only["loss"] == max_loss].index.values[0]
plt.xlabel("Iterace algoritmu COATS")
plt.ylabel("Nákladová funkce")
plt.xticks(coats_df_auc_only.index.values)
plt.locator_params(axis='x', nbins=10)
plt.ylim(0, 1)
def get_dict(dict_str):
return json.loads(dict_str.replace("'", '"'))
lambda_data = [ get_dict(x) for x in coats_df_auc_only["current_params"].to_list() ]
lambda_data_df = pd.DataFrame(lambda_data)
lambda_data_df.head()
lambda_data_df.columns = ["hodnota $\lambda_1$", "hodnota $\lambda_2$", "hodnota $\lambda_3$",
"hodnota $\lambda_4$", "hodnota $\lambda_5$", "hodnota $\lambda_6$", "hodnota $\lambda_7$"]
axes = lambda_data_df.plot(subplots=True, figsize=(12, 14), sharey=False, sharex=False, layout=(3, 3), xticks=lambda_data_df.index.values)
print(axes)
for ax0 in axes.reshape(9):
ax0.set_xlabel("Iterace algoritmu COATS")
ax0.xaxis.set_major_locator(ticker.MaxNLocator(4))
ax0.set_ylim(0, 1000)
```
# Coordinate Ascent - AUC and interpretability bounds
```
coats_df_auc = pd.read_csv("../output_data/coordinate_ascent_run_AUC_interpretability.csv")
coats_df_auc.plot("Unnamed: 0", "loss")
plt.xlabel("Iterace algoritmu COATS")
plt.ylabel("Nákladová funkce")
plt.xticks(coats_df_auc.index.values)
plt.locator_params(axis='x', nbins=10)
plt.ylim(0, 1)
lambda_data = [ get_dict(x) for x in coats_df_auc_only["current_params"].to_list() ]
lambda_data_df = pd.DataFrame(lambda_data)
lambda_data_df.head()
lambda_data_df.columns = ["hodnota $\lambda_1$", "hodnota $\lambda_2$", "hodnota $\lambda_3$",
"hodnota $\lambda_4$", "hodnota $\lambda_5$", "hodnota $\lambda_6$", "hodnota $\lambda_7$"]
axes = lambda_data_df.plot(subplots=True, figsize=(12, 14), sharey=False, sharex=False, layout=(3, 3), xticks=lambda_data_df.index.values)
print(axes)
for ax0 in axes.reshape(9):
ax0.set_xlabel("Iterace algoritmu COATS")
ax0.xaxis.set_major_locator(ticker.MaxNLocator(4))
ax0.set_ylim(0, 1000)
```
# Coordinate Ascent - AUC and interpretability bounds (distance sum penalty)
```
coats_df_auc = pd.read_csv("../output_data/coordinate_ascent_run_AUC_interpretability_distance_sum.csv")
coats_df_auc.plot("Unnamed: 0", "loss")
plt.xlabel("Iterace algoritmu COATS")
plt.ylabel("Nákladová funkce")
plt.xticks(coats_df_auc.index.values)
plt.locator_params(axis='x', nbins=10)
plt.hlines([0], xmin=0, xmax=21, color="red")
lambda_data = [ get_dict(x) for x in coats_df_auc_only["current_params"].to_list() ]
lambda_data_df = pd.DataFrame(lambda_data)
lambda_data_df.head()
lambda_data_df.columns = ["hodnota $\lambda_1$", "hodnota $\lambda_2$", "hodnota $\lambda_3$",
"hodnota $\lambda_4$", "hodnota $\lambda_5$", "hodnota $\lambda_6$", "hodnota $\lambda_7$"]
axes = lambda_data_df.plot(subplots=True, figsize=(12, 14), sharey=False, sharex=False, layout=(3, 3), xticks=lambda_data_df.index.values)
print(axes)
for ax0 in axes.reshape(9):
ax0.set_xlabel("Iterace algoritmu COATS")
ax0.xaxis.set_major_locator(ticker.MaxNLocator(4))
ax0.set_ylim(0, 1000)
```
# Coordinate Ascent - AUC and interpretability bounds (distance euclidean)
```
coats_df_auc = pd.read_csv("../output_data/coordinate_ascent_run_AUC_interpretability_distance_sum_euclidean.csv")
coats_df_auc.plot("Unnamed: 0", "loss")
plt.xlabel("Iterace algoritmu COATS")
plt.ylabel("Nákladová funkce")
plt.xticks(coats_df_auc.index.values)
plt.locator_params(axis='x', nbins=10)
plt.hlines([0], xmin=0, xmax=21, color="red")
lambda_data = [ get_dict(x) for x in coats_df_auc_only["current_params"].to_list() ]
lambda_data_df = pd.DataFrame(lambda_data)
lambda_data_df.head()
lambda_data_df.columns = ["hodnota $\lambda_1$", "hodnota $\lambda_2$", "hodnota $\lambda_3$",
"hodnota $\lambda_4$", "hodnota $\lambda_5$", "hodnota $\lambda_6$", "hodnota $\lambda_7$"]
axes = lambda_data_df.plot(subplots=True, figsize=(12, 14), sharey=False, sharex=False, layout=(3, 3), xticks=lambda_data_df.index.values)
print(axes)
for ax0 in axes.reshape(9):
ax0.set_xlabel("Iterace algoritmu COATS")
ax0.xaxis.set_major_locator(ticker.MaxNLocator(4))
ax0.set_ylim(0, 1000)
```
| github_jupyter |
```
import math
import torch
import matplotlib.pyplot as plt
fpath = "./"
range_ = 10.0
n_pts = 25
fname = "high_loss_" + str(range_) + "_" + str(n_pts)
fname = fname.replace(".", "_")
high_loss = torch.load(fpath + fname, map_location=("cpu"))
fname = "low_loss_" + str(range_) + "_" + str(n_pts)
fname = fname.replace(".", "_")
low_loss = torch.load(fpath + fname, map_location=("cpu"))
fname = "full_loss_" + str(range_) + "_" + str(n_pts)
fname = fname.replace(".", "_")
full_loss = torch.load(fpath + fname, map_location=("cpu"))
ymax = 2500
plt.imshow(full_loss, vmax=ymax)
plt.colorbar()
plt.imshow(low_loss, vmax=ymax)
plt.colorbar()
plt.imshow(high_loss.log())
plt.colorbar()
ymax=high_loss.log().max()
import numpy as np
import matplotlib.cm as cm
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
top = cm.get_cmap('Oranges_r', 128)
bottom = cm.get_cmap('Blues', 128)
newcolors = np.vstack((bottom(np.linspace(1, 0, 128)),
top(np.linspace(1, 0, 128))))
# newcolors[:, -1] = np.linspace(0.25, 0.75, 256)
newcmp = ListedColormap(newcolors, name='OrangeBlue')
def set_params(ax):
ax.set_yticks([0, 12, 24])
ax.set_yticklabels(["-10", "0", "10"])
ax.set_xticks([0, 12, 24])
ax.set_xticklabels(["-10", "0", "10"])
ax.tick_params("both", labelsize=tick_size)
from mpl_toolkits.axes_grid1 import make_axes_locatable, axes_size
title_fs = 28
tick_size=24
ax_fs = 24
fig, ax = plt.subplots(1, 2, figsize=(12, 5))
im = ax[0].imshow(high_loss.log().detach(), cmap=newcmp)
# cbar=fig.colorbar(im, ax=ax[1])
# cbar.ax.tick_params(labelsize=20)
# ax[1].autoscale(False)
ax[0].set_title("Top 2 Eigenvectors",
fontsize=title_fs)
set_params(ax[0])
divider2 = make_axes_locatable(ax[0])
cax2 = divider2.append_axes("right", size="5%", pad=0.05)
cbar = fig.colorbar(im, cax=cax2)
cbar.ax.tick_params(labelsize=tick_size)
# cbar.set_label('Loss', rotation=270, fontsize=ax_fs, labelpad=15)
im = ax[1].imshow(low_loss.log().detach(), cmap=newcmp, vmin=6.8, vmax=6.9)
# ax[0].autoscale(False)
# cbar=fig.colorbar(im, ax=ax[0])
# cbar.ax.tick_params(labelsize=20)
# cbar.ax.yaxis.offsetText.set(size=20)
ax[1].set_title("Degenerate Directions",
fontsize=title_fs)
set_params(ax[1])
divider2 = make_axes_locatable(ax[1])
cax2 = divider2.append_axes("right", size="5%", pad=0.05)
cbar = fig.colorbar(im, cax=cax2)
cbar.ax.tick_params(labelsize=tick_size)
cbar.set_label('Loss', rotation=270, fontsize=ax_fs, labelpad=25)
# fig.subplots_adjust(right=0.8)
# cbar_ax = fig.add_axes([0.82, 0.13, 0.02, 0.75])
# cbar = fig.colorbar(im, cax=cbar_ax)
# cbar.ax.tick_params(labelsize=tick_size)
plt.savefig("./cifar-loss-surface.pdf", bbox_inches="tight")
plt.show()
```
| github_jupyter |
# Lightweight On-line Detector of Anomalies with MinMaxScaler
This code template is for Anomaly detection/outlier analysis using the LODA Algorithm implemented using pyod library and feature scaling using MinMaxScaler.
### Required Packages
```
!pip install plotly
!pip install pyod
import time
import warnings
import pandas as pd
import numpy as np
from scipy import stats
import seaborn as sns
import plotly.express as px
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.manifold import Isomap
from pyod.models.loda import LODA
from sklearn.preprocessing import LabelEncoder, MinMaxScaler
from sklearn.model_selection import train_test_split
warnings.filterwarnings("ignore")
```
### Initialization
Filepath of CSV file
```
file_path= ''
```
List of features which are required for model training
```
features = []
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X.
```
X=df[features]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
X.head()
```
### Data Rescaling
`MinMaxScaler` subtracts the minimum value in the feature and then divides by the range, where range is the difference between the original maximum and original minimum.
[For more Reference](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html)
```
X_Scaled=MinMaxScaler().fit_transform(X)
X_Scaled=pd.DataFrame(data = X_Scaled,columns = X.columns)
X_Scaled.head()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test=train_test_split(X_Scaled,test_size=0.2,random_state=123)
```
### Model
Loda: Lightweight on-line detector of anomalies Adapted from tilitools
#### Tuning parameters
**contamination** – The amount of contamination of the data set, i.e. the proportion of outliers in the data set. Used when fitting to define the threshold on the decision function.
**n_bins** – The number of bins for the histogram.
**n_random_cuts** – The number of random cuts.
[For more information](https://pyod.readthedocs.io/en/latest/pyod.models.html#module-pyod.models.loda)
```
model = LODA(contamination=0.001)
model.fit(x_train)
```
### Anomaly Prediction
```
result=x_test.copy(deep=True)
result['Anomaly']=model.predict(x_test)
result.head()
```
### Anomaly Visualization
#### Bar Plot
```
result['Anomaly'].value_counts().plot(kind='bar',color=['green','red'])
```
#### Pie Chart
```
fig = px.pie(result['Anomaly'],names=result['Anomaly'], title='Anomaly rate',)
fig.show()
```
#### Anomalies
In this part we will perform Dimensionality Reduction technique to visualize data. This can be performed using technique such as PCA or TSNE algorithms.
```
pca = PCA(n_components=2)
pca_results = pca.fit_transform(result.drop('Anomaly',axis=1))
plt.rcParams["figure.figsize"] = (20,10)
plt.scatter(x=pca_results[:,0],y=pca_results[:,1],c=result.iloc[:,result.columns.get_loc('Anomaly')])
plt.show()
```
#### Creator: Vamsi Mukkamala , Github: [Profile](https://github.com/vmc99)
| github_jupyter |
```
import cv2
import PIL
import kornia
import glob
import torch
import numpy as np
import imgaug as ia
import imgaug.augmenters as iaa
import matplotlib.pyplot as plt
from torchvision import transforms as T
from networks.ResnetFaceSTN import ResnetFaceSTN
class RowImage:
def __init__(self, resize_dim=None):
self.imgs = []
self.resize_dim = resize_dim
def add_img(self, img):
if isinstance(self.resize_dim, tuple):
img = cv2.resize(img, self.resize_dim,
interpolation = cv2.INTER_AREA)
if len(self.imgs) == 0:
self.imgs = img
else:
self.imgs = np.hstack((self.imgs, img))
def get_row_img(self):
return self.imgs
def __call__(self):
return self.imgs
```
## Transform Definitions
```
class Transform:
def __init__(self):
self.aug = iaa.Sequential([
iaa.Fliplr(0.5),
iaa.Affine(
scale=(0.8, 1.2),
translate_percent={"x": (-0.1, 0.1), "y": (-0.1, 0.1)},
order=[0, 1],
mode='edge'
),
iaa.Resize({"height": 128, "width": 128})
])
def __call__(self, img):
img = np.asarray(img)
return self.aug.augment_image(img)
transform_train = T.Compose([
Transform(),
T.ToTensor(),
T.Normalize(mean=[0.5,0.5,0.5], std=[0.5,0.5,0.5])
])
class UnNormalize(object):
def __init__(self, mean, std):
self.mean = mean
self.std = std
def __call__(self, tensor):
"""
Args:
tensor (Tensor): Tensor image of size (C, H, W) to be normalized.
Returns:
Tensor: Normalized image.
"""
for t, m, s in zip(tensor, self.mean, self.std):
t.mul_(s).add_(m)
return tensor
transform = T.Compose(
[
T.Resize((128, 128)),
T.ToTensor(),
T.Normalize(mean=[0.5,0.5,0.5], std=[0.5,0.5,0.5])
]
)
```
## Natural Sorting
```
import re
def atoi(text):
return int(text) if text.isdigit() else text
def natural_keys(text):
'''
alist.sort(key=natural_keys) sorts in human order
http://nedbatchelder.com/blog/200712/human_sorting.html
(See Toothy's implementation in the comments)
'''
return [ atoi(c) for c in re.split(r'(\d+)', text) ]
```
## Weight Paths and Image Paths
Weight paths
```
w_glob = "weights/mask_exp19-resnetSTN/epoch_*/mask_exp19-resnetSTN_ep*.pth"
wpaths = glob.glob(w_glob)
wpaths.sort(key=natural_keys)
wpaths = wpaths[2:20:3]
weight_count = len(wpaths)
print(wpaths)
print(weight_count)
```
Image paths
```
sample_images = glob.glob('image_samples/LFW-Masked/*.jpg')
sample_images.sort(key=natural_keys)
print(sample_images)
```
## Preview
```
torch.set_printoptions(precision=10)
np.set_printoptions(suppress=True)
row = 1
col = weight_count + 1
start_2 = col + 1
ep_start = 1
# eps = [i for i in range(ep_start, ep_start+weight_count)]
eps = ['3', '6', '9', '12', '15', '18']
print(eps)
row_imgs = [RowImage(resize_dim=(112, 112)) for i in range(5)]
net = ResnetFaceSTN(stn_mode='resnet')
unorm = UnNormalize(mean=[0.5,0.5,0.5], std=[0.5,0.5,0.5])
for n, img_path in enumerate(sample_images, 1):
img = PIL.Image.open(img_path)
img_t = transform(img).unsqueeze(0)
img_p = kornia.tensor_to_image(unorm(img_t.clone().detach()))
# plt.figure() #figsize=(3 * col, 5)
# plt.subplot(row, col, 1)
# plt.imshow(np.asarray(img_p))
# plt.tight_layout()
row_imgs[n-1].add_img(img_p)
for i, weight in enumerate(wpaths, 2):
net.load_state_dict(torch.load(weight))
net.eval()
with torch.no_grad():
img_t2 = img_t.clone().detach()
img_stn = net.stn(img_t2)
img_p = kornia.tensor_to_image(unorm(img_stn.clone().detach()))
row_imgs[n-1].add_img(img_p)
# plt.subplot(row, col, i)
# plt.xlabel(f'ep {eps[i-2]}')
# plt.yticks([])
# plt.imshow(np.asarray(img_p))
# plt.tight_layout()
img = PIL.Image.open(img_path)
# plt.tight_layout()
# plt.subplots_adjust(left=None, bottom=0., right=None, top=None, wspace=0., hspace=0.)
# plt.show()
fig = np.vstack([r() for r in row_imgs])
plt.imshow(fig)
plt.show()
plt.imsave("pair_sample_result/stn_epoch_comparison.jpg", fig)
```
| github_jupyter |
[Table of Contents](./table_of_contents.ipynb)
# Multivariate Gaussians
Modeling Uncertainty in Multiple Dimensions
```
#format the book
%matplotlib inline
from __future__ import division, print_function
from book_format import load_style
load_style()
```
## Introduction
The techniques in the last chapter are very powerful, but they only work with one variable or dimension. They provide no way to represent multidimensional data, such as the position and velocity of a dog in a field. Position and velocity are related to each other, and as we learned in the g-h chapter we should never throw away information. In this chapter we learn how to describe this relationship probabilistically. Through this key insight we will achieve markedly better filter performance.
## Multivariate Normal Distributions
We've been using Gaussians for a scalar random variable, expressed as $\mathcal{N}(\mu, \sigma^2)$. A more formal term for this is *univariate normal*, where univariate means 'one variable'. The probability distribution of the Gaussian is known as the *univariate normal distribution*.
What might a *multivariate normal distribution* be? *Multivariate* means multiple variables. Our goal is to be able to represent a normal distribution with multiple dimensions. I don't necessarily mean spatial dimensions - if we track the position, velocity, and acceleration of an aircraft in (x, y, z) that gives us a nine dimensional problem. Consider a two dimensional case. It might be the *x* and *y* coordinates of a robot, it might be the position and velocity of a dog on the x-axis, or milk production and feed rate at a dairy. It doesn't really matter. We can see that for $N$ dimensions, we need $N$ means, which we will arrange in a column matrix (vector) like so:
$$
\mu = \begin{bmatrix}\mu_1\\\mu_2\\ \vdots \\\mu_n\end{bmatrix}
$$
Let's say we believe that $x = 2$ and $y = 17$. We would have
$$
\mu = \begin{bmatrix}2\\17\end{bmatrix}
$$
The next step is representing our variances. At first blush we might think we would also need N variances for N dimensions. We might want to say the variance for x is 10 and the variance for y is 4, like so.
$$\sigma^2 = \begin{bmatrix}10\\4\end{bmatrix}$$
This is incomplete because it does not consider the more general case. In the **Gaussians** chapter we computed the variance in the heights of students. That is a measure of how the heights vary relative to each other. If all students are the same height, then the variance is 0, and if their heights are wildly different, then the variance will be large.
There is also a relationship between height and weight. In general, a taller person weighs more than a shorter person. Height and weight are *correlated*. We want a way to express not only what we think the variance is in the height and the weight, but also the degree to which they are correlated. In other words, we want to know how weight varies compared to the heights. We call that the *covariance*.
Before we can understand multivariate normal distributions we need to understand the mathematics behind correlations and covariances.
## Correlation and Covariance
*Covariance* describes how much two variables vary together. Covariance is short for *correlated variances*. In other words, *variance* is a measure for how a population vary amongst themselves, and *covariance* is a measure for how much two variables change in relation to each other. For example, as height increases weight also generally increases. These variables are *correlated*. They are *positively correlated* because as one variable gets larger so does the other. As the outdoor temperature decreases home heating bills increase. These are *inversely correlated* or *negatively correlated* because as one variable gets larger the other variable lowers. The price of tea and the number of tail wags my dog makes have no relation to each other, and we say they are *uncorrelated* or *independent*- each can change independent of the other.
Correlation allows prediction. If you are significantly taller than me I can predict that you also weigh more than me. As winter comes I predict that I will be spending more to heat my house. If my dog wags his tail more I don't conclude that tea prices will be changing.
For example, here is a plot of height and weight of students on the school's track team. If a student is 68 inches tall I can predict they weigh roughly 160 pounds. Since the correlation is not perfect neither is my prediction.
```
from kf_book.gaussian_internal import plot_correlated_data
height = [60, 62, 63, 65, 65.1, 68, 69, 70, 72, 74]
weight = [95, 120, 127, 119, 151, 143, 173, 171, 180, 210]
plot_correlated_data(height, weight, 'Height (in)', 'Weight (lbs)', False)
```
In this book we only consider *linear correlation*. We assume that the relationship between variables is linear. That is, a straight line is a good fit for the data. I've fit a straight line through the data in the above chart. The concept of *nonlinear correlation* exists, but we will not be using it.
The equation for the covariance between $X$ and $Y$ is
$$ COV(X, Y) = \sigma_{xy} = \mathbb E\big[(X-\mu_x)(Y-\mu_y)\big]$$
Where $\mathbb E[X]$ is the *expected value* of X, defined as
$$\mathbb E[X] = \begin{cases} \sum_{i=1}^n p_ix_i & \mbox{discrete}\\ \int_{-\infty}^\infty f(x)\, x & \mbox{continuous}\end{cases}$$
We assume each data point is equally likely, so the probability of each is $\frac{1}{N}$, giving
$$\mathbb E[X] = \frac{1}{N}\sum_{i=1}^n x_i$$
for the discrete case we will be considering.
Compare the covariance equation to the equation for the variance. As you can see they are very similar:
$$\begin{aligned}VAR(X) = \sigma_x^2 &= \mathbb E[(X - \mu)^2]\\
COV(X, Y) = \sigma_{xy} &= \mathbb E\big[(X-\mu_x)(Y-\mu_y)\big]\end{aligned}$$
In particular, if you compute $COV(X, X)$ you get the equation for $VAR(X)$, which supports my statement that the variance computes how a random variable varies amongst itself.
We use a *covariance matrix* to denote covariances of a multivariate normal distribution, and it looks like this:
$$
\Sigma = \begin{bmatrix}
\sigma_1^2 & \sigma_{12} & \cdots & \sigma_{1n} \\
\sigma_{21} &\sigma_2^2 & \cdots & \sigma_{2n} \\
\vdots & \vdots & \ddots & \vdots \\
\sigma_{n1} & \sigma_{n2} & \cdots & \sigma_n^2
\end{bmatrix}
$$
The diagonal contains the variance for each variable, and the off-diagonal elements contain the covariance between the $i^{th}$ and $j^{th}$ variables. So $\sigma_3^2$ is the variance of the third variable, and $\sigma_{13}$ is the covariance between the first and third variables.
A covariance of 0 indicates no correlation. If the variance for $x$ is 10, the variance for $y$ is 4, and there is no linear correlation between $x$ and $y$, then we would write
$$\Sigma = \begin{bmatrix}10&0\\0&4\end{bmatrix}$$
If there was a small amount of positive correlation between $x$ and $y$ we might have
$$\Sigma = \begin{bmatrix}10&1.2\\1.2&4\end{bmatrix}$$
where 1.2 is the covariance between $x$ and $y$. I say the correlation is "small" because the covariance of 1.2 is small relative to the variances of 10.
If there was a large amount of negative correlation between between $x$ and $y$ we might have
$$\Sigma = \begin{bmatrix}10&-9.7\\-9.7&4\end{bmatrix}$$
The covariance matrix is symmetric. After all, the covariance between $x$ and $y$ is always equal to the covariance between $y$ and $x$. That is, $\sigma_{xy}=\sigma_{yx}$ for any $x$ and $y$.
I fear I might be losing you, so let's work an example. In the **Gaussians** chapter we had a class of students with heights H=[1.8, 2.0, 1.7, 1.9, 1.6] meters. We computed:
$$\begin{aligned}
\mathit{VAR}(H) &= E[(H - \mu_H)^2] \\
&= \frac{1}{N}\sum_{i=1}^n (H_i - \mu_H)^2 \\
&= \frac{1}{5}\left[(1.8-1.8)^2 + (2-1.8)^2 + (1.7-1.8)^2 + (1.9-1.8)^2 + (1.6-1.8)^2\right] \\
&= 0.02
\end{aligned}$$
Easy, right? If we weigh the students we might find their weights to be W = [70.1, 91.2, 59.5, 93.2, 53.5]. Can we use the covariance equation to create the covariance matrix? Sure. It will look like:
$$\Sigma = \begin{bmatrix}\sigma_H^2 & \sigma_{H,W} \\
\sigma_{W,H} & \sigma_{W}^2\end{bmatrix}$$
We just computed the variance of the height, and it will go in the upper left hand corner of the matrix. The lower right corner contains the variance in weights. Using the same equation we get:
$$\begin{aligned}
\mu_W &= \frac{1}{5}(70.1 + 91.2 + 59.5 + 93.2 + 53.5) = 73.5 \\
\sigma_W^2 &= \frac{1}{5}\left[(70.1-73.5)^2 + (91.2-73.5)^2 + (59.5-73.5)^2 + (93.2-73.5)^2 + (53.5-73.5)^2\right] \\
&= 261.8
\end{aligned}$$
Now the covariances. Using the formula above, we compute:
$$\begin{aligned}
\sigma_{H,W} &= \mathbb E\big[(H-\mu_H)(W-\mu_W)\big] \\
&= \frac{1}{N}\sum_{i=1}^n (H_i-\mu_H)(W_i-\mu_W) \\
&= \frac{1}{5}[(1.8-1.8)(70.1-73.5) + (2-1.8)(91.2-73.5) + (1.7-1.8)(59.5-73.5)\, +\\
&\, \, \, \, \, (1.9-1.8)(93.2-73.5) + (1.6-1.8)(53.5-73.5)] \\
&= 2.18
\end{aligned}$$
That was tedious, but easy enough. We will never do that again because, of course, NumPy will compute it for you.
```
import numpy as np
W = [70.1, 91.2, 59.5, 93.2, 53.5]
H = [1.8, 2.0, 1.7, 1.9, 1.6]
np.cov(H, W)
```
That doesn't agree with our calculation! What went wrong? Nothing. NumPy applies a correction for small sample sizes; it uses $\frac{1}{N-1}$ as the normalization term instead of $\frac{1}{N}$.
This is a bit beyond the scope of this book. Briefly, suppose the actual class size is 200 students, and we took a sample of 5 students to perform this computation because we couldn't afford to measure and weigh all 200 students. It is nearly certain that there will be some error in our estimator because the sample is unlikely to perfectly represent the class. As our sample size approaches 200 the error will approach 0. We say there is no *bias* in the latter, and that we have an *unbiased estimator*. In contrast, when we take a small sample there is bias (error is nonzero), and we have a *biased estimator*.
If the error is zero it makes sense to divide by $N$. I will not prove why, but for biased estimators we use $\frac{1}{N-1}$ to correct for the small sample size. NumPy does this by default because in practice we are almost always working from data samples from a larger collection. If you want the unbiased estimator, which we computed above, use `bias=1` in the call to `np.cov'.
```
np.cov(H, W, bias=1)
```
This agrees with our computation. We will not use `bias=1` again in this book since we are using *random variables* which are sampling from the infinite set of positions of the objects we are tracking. Here we are computing the variance and covariance for the entire population, so `bias=1` is correct.
What does this matrix tell us? It tells us the variance in heights is 0.02 $m^2$ and the variance in weights is 261.788 $kg^2$. Furthermore, it tells us the weights and heights are positively correlated - as heights increase so do the weights.
Let's create perfectly correlated data. By this I mean that the data perfectly fits on a line - there is no variance from the line.
```
X = np.linspace(1, 10, 100)
Y = np.linspace(1, 10, 100)
np.cov(X, Y)
```
We can see from the covariance matrix that the covariance is equal to the variance in x and in y.
Now let's add some noise to one of the variables so that they are no longer perfectly correlated. I will make $Y$ negative to create a negative correlation.
```
X = np.linspace(1, 10, 100)
Y = -(np.linspace(1, 5, 100) + np.sin(X)*.2)
plot_correlated_data(X, Y)
print(np.cov(X, Y))
```
The data no longer forms a straight line. The covariance is $\sigma_{xy}=-3.08$. It is not close to zero compared to the magnitudes of $\sigma_x^2$ and $\sigma_y^2$, and so we know there is still a high degree of correlation. We can verify this by looking at the chart. The data forms nearly a straight line.
Now I will add random noise to a straight line.
```
from numpy.random import randn
X = np.linspace(1, 10, 1000) + randn(1000)*2
Y = np.linspace(1, 5, 1000) + randn(1000)
plot_correlated_data(X, Y)
print(np.cov(X, Y))
```
We see that the covariance is smaller in relation to the variances, reflecting the lower correlation between $X$ and $Y$. We can still fit a straight line through this data, but there is much greater variation in the data.
Finally, here is the covariance between completely random data.
```
X = randn(100000)
Y = randn(100000)
plot_correlated_data(X, Y)
print(np.cov(X, Y))
```
Here the covariances are very near zero. As you can see with the plot, there is no clear way to draw a line to fit the data. A vertical line would be as unconvincing as the horizontal line I've shown.
## Multivariate Normal Distribution Equation
Recall the equation for the normal distribution from the **Gaussians** chapter:
$$
f(x, \mu, \sigma) = \frac{1}{\sqrt{2\pi\sigma^2}} \exp \Big [{-\frac{1}{2}}{(x-\mu)^2}/\sigma^2 \Big ]
$$
Here is the multivariate normal distribution in $n$ dimensions.
$$
f(\mathbf{x},\, \mu,\,\Sigma) = \frac{1}{\sqrt{(2\pi)^n|\Sigma|}}\, \exp \Big [{ -\frac{1}{2}(\mathbf{x}-\mu)^\mathsf{T}\Sigma^{-1}(\mathbf{x}-\mu) \Big ]}
$$
The multivariate version merely replaces the scalars of the univariate equations with matrices. If you are reasonably well-versed in linear algebra this equation should look quite manageable. If not, don't worry, both FilterPy and SciPy provide functions to compute it for you. Let's ignore the computation for a moment and plot it to see what it looks like.
```
import kf_book.mkf_internal as mkf_internal
mean = [2., 17.]
cov = [[10., 0.],
[0., 4.]]
mkf_internal.plot_3d_covariance(mean, cov)
```
This is a plot of multivariate Gaussian with a mean of $\mu=[\begin{smallmatrix}2\\17\end{smallmatrix}]$ and a covariance of $\Sigma=[\begin{smallmatrix}10&0\\0&4\end{smallmatrix}]$. The three dimensional shape shows the probability density for any value of $(X, Y)$ in the z-axis. I have projected the variance for x and y onto the walls of the chart - you can see that they take on the Gaussian bell curve shape. The curve for $X$ is wider than the curve for $Y$, which is explained by $\sigma_x^2=10$ and $\sigma_y^2=4$. The highest point of the 3D surface is at the the means for $X$ and $Y$.
All multivariate Gaussians have this shape. If we think of this as the Gaussian for the position of a dog, the z-value at each point of ($X, Y$) is the probability density of the dog being at that position. Strictly speaking this is the *joint probability density function*, which I will define soon. So, the dog has the highest probability of being near (2, 17), a modest probability of being near (5, 14), and a very low probability of being near (10, 10). As with the univariate case this is a *probability density*, not a *probability*. Continuous distributions have an infinite range, and so the probability of being exactly at (2, 17), or any other point, is 0%. We can compute the probability of being within a given range by computing the volume under the surface with an integral.
FilterPy [2] implements the equation with the function `multivariate_gaussian()` in the `filterpy.stats.` module. SciPy's `stats` module implements the multivariate normal equation with `multivariate_normal()`. It implements a 'frozen' form where you set the mean and covariance once, and then calculate the probability density for any number of values for x over any arbitrary number of calls. I named my function `multivariate_gaussian()` to ensure it is never confused with the SciPy version.
> The <a href="http://docs.scipy.org/doc/scipy/reference/tutorial/stats.html">tutorial</a>[1] for the `scipy.stats` module explains 'freezing' distributions and other very useful features.
```
from filterpy.stats import gaussian, multivariate_gaussian
```
I'll demonstrate using it, and then move on to more interesting things.
First, let's find the probability density for our dog being at (2.5, 7.3) if we believe he is at (2, 7) with a variance of 8 for $x$ and a variance of 3 for $y$.
Start by setting $x$ to (2.5, 7.3). You can use a tuple, list, or NumPy array.
```
x = [2.5, 7.3]
```
Next, we set the mean of our belief:
```
mu = [2.0, 7.0]
```
Finally, we have to define our covariance matrix. In the problem statement we did not mention any correlation between $x$ and $y$, and we will assume there is none. This makes sense; a dog can choose to independently wander in either the $x$ direction or $y$ direction without affecting the other. I will use the variable name `P`. Kalman filters use the name $\textbf{P}$ for the covariance matrix, and we need to become familiar with the conventions.
```
P = [[8., 0.],
[0., 3.]]
```
Now call the function
```
%precision 4
multivariate_gaussian(x, mu, P)
```
We can get the same result from the `scipy.stats` module.
```
import scipy
try:
print('{:.4f}'.format(scipy.stats.multivariate_normal(mu, P).pdf(x)))
except:
print('you have an old version of scipy, upgrade it now!')
```
It's time to define some terms. The *joint probability*, denoted $P(x,y)$, is the probability of both $x$ and $y$ happening. For example, if you roll two die $P(2,5)$ is the probability of the first die rolling a 2 and the second die rolling a 5. Assuming the die are six sided and fair, the probability $P(2,5) = \frac{1}{6}\times \frac{1}{6}=\frac{1}{36}$. The 3D chart above shows the *joint probability density function*.
The *marginal probability* is the probability of an event happening without regard of any other event. In the chart above the Gaussian curve drawn to the left is the marginal for $Y$. This is the probability for the dog being at any position in $Y$ disregarding the value for $X$. Earlier I wrote "I have projected the variance for x and y onto the walls of the chart"; these are the marginal probabilities for $x$ and $y$. Another computational benefit of Gaussians is that the marginal of a multivariate Gaussian is another Gaussian!
Let's look at this in a slightly different way. Instead of plotting a surface showing the probability distribution I will generate 1,000 points with the distribution of $[\begin{smallmatrix}8&0\\0&3\end{smallmatrix}]$.
```
mkf_internal.plot_3d_sampled_covariance(mu, P)
```
We can think of the sampled points as being possible locations for our dog given those particular mean and covariances. The contours on the side show the marginal probability for $X$ and $Y$. We can see that he is far more likely to be at (2, 7) where there are many points, than at (-5, 5) where there are few.
As beautiful as these plots are, it is hard to get useful information from them. For example, it is not easy to tell if $X$ and $Y$ both have the same variance, and how much they are correlated. In most of the book I'll display Gaussians as contour plots.
The contour plots display the range of values that the multivariate Gaussian takes for a specific standard deviation. This is like taking a horizontal slice out of the 3D plot.
These plots show the shape of the slice for 3 standard deviations.
```
mkf_internal.plot_3_covariances()
```
For those of you viewing this online or in Juptyer Notebook on your computer, here is an animation of varying the covariance while holding the variance constant.
<img src='animations/multivariate_ellipse.gif'>
(source: http://git.io/vqxLS)
These plots look like circles and ellipses. Indeed, it turns out that any slice through the multivariate Gaussian is an ellipse. Hence, in statistics we do not call these 'contour plots', but either *error ellipses* or *confidence ellipses*; the terms are interchangable.
This code uses the function `plot_covariance_ellipse()` from `filterpy.stats`. By default the function displays one standard deviation, but you can use either the `variance` or `std` parameter to control what is displayed. For example, `variance=3**2` or `std=3` would display the 3rd standard deviation, and `variance=[1,4,9]` or `std=[1,2,3]` would display the 1st, 2nd, and 3rd standard deviations.
```
from filterpy.stats import plot_covariance_ellipse
import matplotlib.pyplot as plt
P = [[2, 0], [0, 6]]
plot_covariance_ellipse((2, 7), P, fc='g', alpha=0.2,
std=[1, 2, 3],
title='|2 0|\n|0 6|')
plt.gca().grid(b=False);
```
The solid colors may suggest to you that the probability distribution is constant between the standard deviations. This is not true, as you can tell from the 3D plot of the Gaussian. Here is a 2D shaded representation of the probability distribution for the covariance ($\begin{smallmatrix}2&1.2\\1.2&1.3\end{smallmatrix})$. Darker gray corresponds to higher probability density.
```
from kf_book.nonlinear_plots import plot_cov_ellipse_colormap
plot_cov_ellipse_colormap(cov=[[2, 1.2], [1.2, 1.3]]);
```
Thinking about the physical interpretation of these plots clarifies their meaning. The mean and covariance of the first plot is
$$
\mathbf{\mu} =\begin{bmatrix}2\\7\end{bmatrix},\, \,
\Sigma = \begin{bmatrix}2&0\\0&2 \end{bmatrix}
$$
```
x = [2, 7]
P = [[2, 0], [0, 2]]
plot_covariance_ellipse(x, P, fc='g', alpha=0.2,
title='|2 0|\n|0 2|')
plt.gca().grid(b=False)
```
A Bayesian way of thinking about this is that the ellipse shows us the amount of error in our belief. A tiny circle would indicate that we have a very small error, and a very large circle indicates a lot of error in our belief. The shape of the ellipse shows us the geometric relationship of the errors in $X$ and $Y$. Here we have a circle so errors in $X$ and $Y$ are equally likely.
The mean and covariance of the second plot are
$$
\mu =\begin{bmatrix}2\\7\end{bmatrix}, \, \, \,
\Sigma = \begin{bmatrix}2&0\\0&6\end{bmatrix}
$$
```
x = [2, 7]
P = [[2, 0], [0, 6]]
plot_covariance_ellipse(x, P, fc='g', alpha=0.2,
title='|2 0|\n|0 6|')
plt.gca().grid(b=False)
```
This time we use a different variance for $X$ ($\sigma_x^2=2$) vs $Y$ ($\sigma^2_y=6$). The result is a tall and narrow ellipse. We can see that a lot more uncertainty in $Y$ vs $X$. In both cases we believe the dog is at (2, 7), but the uncertainties are different.
The third plot shows the mean and covariance
$$
\mu =\begin{bmatrix}2\\7\end{bmatrix}, \, \, \,
\Sigma = \begin{bmatrix}2&1.2\\1.2&2\end{bmatrix}
$$
```
x = [2, 7]
P = [[2, 1.2], [1.2, 2]]
plot_covariance_ellipse(x, P, fc='g', alpha=0.2,
title='|2 1.2|\n|1.2 2|')
```
This is the first contour that has values in the off-diagonal elements of the covariance, and this is the first contour plot with a slanted ellipse. This is not a coincidence. The two facts are telling us the same thing. A slanted ellipse tells us that the $x$ and $y$ values are somehow correlated. The off-diagonal elements in the covariance matrix are non-zero, indicating that a correlation exists.
Recall the plot for height versus weight. It formed a slanted grouping of points. We can use NumPy's `cov()` function to compute the covariance of two or more variables by placing them into a 2D array. Let's do that, then plot the $2\sigma$ covariance ellipse on top of the data. We will need to use `bias=1` because the data represents the entire population; it is not a sample.
```
cov_hw = np.cov(np.vstack((height, weight)), bias=1)
cov_hw
plt.scatter(height, weight, s=120, marker='s')
plt.title('Track Team Height vs. Weight')
plt.xlabel('Height (in)'); plt.ylabel('Weight (lbs)')
plot_covariance_ellipse((np.mean(height), np.mean(weight)), cov_hw, fc='g',
alpha=0.2, axis_equal=False, std=2)
```
This should help you form a strong intuition on the meaning and use of covariances. The covariance ellipse shows you how the data is 'scattered' in relation to each other. A narrow ellipse like this tells you that the data is very correlated. There is only a narrow range of weights for any given height. The ellipse leans towards the right, telling us there is a positive correlation - as x increases y also increases. If the ellipse leaned towards the left then the correlation would be negative - as x increases y decreases. We can see this in the following plot:
```
max_temp = [200, 250, 300, 400, 450, 500]
lifespan = [10, 9.7, 5, 5.4, 4.3, 0.3]
plt.scatter(max_temp, lifespan, s=80)
cov = np.cov(np.vstack((max_temp, lifespan)))
plot_covariance_ellipse((np.mean(max_temp), np.mean(lifespan)), cov, fc='g',
alpha=0.2, axis_equal=False, std=2)
plt.title('Engine Temperature vs Lifespan')
plt.xlabel('Temperature (C)'); plt.ylabel('Years');
```
The relationships between variances and covariances can be hard to puzzle out by inspection, so here is an interactive plot. (If you are reading this in a static form instructions to run this online are here: https://git.io/vza7b)
```
from ipywidgets import interact
from kf_book.book_plots import figsize, FloatSlider
fig = None
def plot_covariance(var_x, var_y, cov_xy):
global fig
if fig: plt.close(fig)
fig = plt.figure(figsize=(4,4))
P1 = [[var_x, cov_xy], [cov_xy, var_y]]
plot_covariance_ellipse((10, 10), P1, axis_equal=False,
show_semiaxis=True)
plt.xlim(4, 16)
plt.gca().set_aspect('equal')
plt.ylim(4, 16)
with figsize(y=6):
interact (plot_covariance,
var_x=FloatSlider(5, min=0, max=20),
var_y=FloatSlider(5, min=0, max=20),
cov_xy=FloatSlider(1.5, min=0, max=50, step=.2));
```
### Pearson's Correlation Coefficient
We will not be using this coefficient in this book, but you may see it elsewhere. You can safely skip this section if uninterested.
The correlation between two variables can be given a numerical value with *Pearson's Correlation Coefficient*. It is defined as
$$\rho_{xy} = \frac{COV(X, Y)}{\sigma_x \sigma_y}$$
This value can range in value from -1 to 1. If the covariance is 0 than $\rho=0$. A value greater than 0 indicates that the relationship is a positive correlation, and a negative value indicates that there is a negative correlation. Values near -1 or 1 indicate a very strong correlation, and values near 0 indicate a very weak correlation.
Correlation and covariance are very closely related. Covariance has units associated with it, and correlation is a unitless ratio. For example, for our dog $\sigma_{xy}$ has units of meters squared.
We can use `scipy.stats.pearsonr` function to compute the Pearson coefficient. It returns a tuple of the Pearson coefficient and of the 2 tailed p-value. The latter is not used in this book. Here we compute $\rho$ for height vs weight of student athletes:
```
from scipy.stats import pearsonr
pearsonr(height, weight)[0]
```
Here we compute the correlation between engine temperature and lifespan.
```
pearsonr(max_temp, lifespan)[0]
```
## Using Correlations to Improve Estimates
Suppose we believe our dog is at position (5, 10) with some given covariance. If the standard deviation in x and y is each 2 meters, but they are strongly correlated, the covariance contour would look something like this.
```
P = [[4, 3.9], [3.9, 4]]
plot_covariance_ellipse((5, 10), P, ec='k', std=[1, 2, 3])
plt.xlabel('X')
plt.ylabel('Y');
```
Now suppose I were to tell you that we know that $x=7.5$. What can we infer about the value for $y$? The position is extremely likely to lie within the 3$\sigma$ covariance ellipse. We can infer the position in *y* based on the covariance matrix because there is a correlation between *x* and *y*. I've illustrated the likely range of values for y as a blue filled circle.
```
mkf_internal.plot_correlation_covariance()
```
The circle not mathematically correct, but it gets the idea across. We will tackle the mathematics in the next section. For now recognize that we can predict that $y$ is likely near 12. A value of $y=-10$ is extremely improbable.
A word about *correlation* and *independence*. If variables are *independent* they can vary separately. If you walk in an open field, you can move in the $x$ direction (east-west), the $y$ direction(north-south), or any combination thereof. Independent variables are always also *uncorrelated*. Except in special cases, the reverse does not hold true. Variables can be uncorrelated, but dependent. For example, consider $y=x^2$. Correlation is a linear measurement, so $x$ and $y$ are uncorrelated. However, $y$ is dependent on $x$.
## Multiplying Multidimensional Gaussians
In the previous chapter we incorporated an uncertain measurement with an uncertain estimate by multiplying their Gaussians together. The result was another Gaussian with a smaller variance. If two pieces of uncertain information corroborate each other we should be more certain in our conclusion. The graphs look like this:
```
mkf_internal.plot_gaussian_multiply()
```
The combination of measurements 1 and 2 yields more certainty, so the new Gaussian is taller and narrower - the variance became smaller. The same happens in multiple dimensions with multivariate Gaussians.
Here are the equations for multiplying multivariate Gaussians. The capital sigma ($\Sigma$) indicates that these are matrices, not scalars. Specifically, they are covariance matrices:
$$\begin{aligned}
\mu &= \Sigma_2(\Sigma_1 + \Sigma_2)^{-1}\mu_1 + \Sigma_1(\Sigma_1 + \Sigma_2)^{-1}\mu_2 \\
\Sigma &= \Sigma_1(\Sigma_1+\Sigma_2)^{-1}\Sigma_2
\end{aligned}$$
They are generated by plugging the multivariate Gaussians for the prior and the estimate into Bayes Theorem. I gave you the algebra for the univariate case in the **Gaussians** chapter.
You will not need to remember these equations as they are computed by Kalman filter equations that will be presented shortly. This computation is also available in FilterPy using the `multivariate_multiply()` method, which you can import from `filterpy.stats`.
To give you some intuition about this, recall the equations for multiplying univariate Gaussians:
$$\begin{aligned}
\mu &=\frac{\sigma_1^2 \mu_2 + \sigma_2^2 \mu_1} {\sigma_1^2 + \sigma_2^2}, \\
\sigma^2 &= \frac{\sigma_1^2\sigma_2^2}{\sigma_1^2+\sigma_2^2}
\end{aligned}$$
This looks similar to the equations for the multivariate equations. This will be more obvious if you recognize that matrix inversion, denoted by the -1 power, is *like* a reciprocal since $AA^{-1} =I$. I will rewrite the inversions as divisions - this is not a mathematically correct thing to do as division for matrices is not defined, but it does help us compare the equations.
$$\begin{aligned}
\mu &\approx \frac{\Sigma_2\mu_1 + \Sigma_1\mu_2}{\Sigma_1 + \Sigma_2} \\ \\
\Sigma &\approx \frac{\Sigma_1\Sigma_2}{(\Sigma_1+\Sigma_2)}
\end{aligned}$$
In this form the relationship between the univariate and multivariate equations is clear.
Now let's explore multivariate Gaussians in terms of a concrete example. Suppose that we are tracking an aircraft with two radar systems. I will ignore altitude so I can use two dimensional plots. Radar provides the range and bearing to a target. We start out being uncertain about the position of the aircraft, so the covariance, which is our uncertainty about the position, might look like this. In the language of Bayesian statistics this is our *prior*.
```
P0 = [[6, 0], [0, 6]]
plot_covariance_ellipse((10, 10), P0, fc='y', alpha=0.6)
```
Now suppose that there is a radar to the lower left of the aircraft. Further suppose that the radar's bearing measurement is accurate, but the range measurement is inaccurate. The covariance for the error in the measurement might look like this (plotted in green on top of the yellow prior):
```
P1 = [[2, 1.9], [1.9, 2]]
plot_covariance_ellipse((10, 10), P0, fc='y', alpha=0.6)
plot_covariance_ellipse((10, 10), P1, fc='g', alpha=0.9)
```
Recall that Bayesian statistics calls this the *evidence*. The ellipse points towards the radar. It is very long because the range measurement is inaccurate, and the aircraft could be within a considerable distance of the measured range. It is very narrow because the bearing estimate is very accurate and thus the aircraft must be very close to the bearing estimate.
We want to find the *posterior* - the mean and covariance that results from incorporating the evidence into the prior. As in every other chapter we combine evidence by multiplying them together.
```
from filterpy.stats import multivariate_multiply
P2 = multivariate_multiply((10, 10), P0, (10, 10), P1)[1]
plot_covariance_ellipse((10, 10), P0, ec='k', fc='y', alpha=0.2)
plot_covariance_ellipse((10, 10), P1, ec='k', fc='g', alpha=0.9)
plot_covariance_ellipse((10, 10), P2, ec='k', fc='b')
```
I have plotted the original estimate (prior) in a very transparent yellow, the radar reading in green (evidence), and the finale estimate (posterior) in blue.
The posterior retained the same shape and position as the radar measurement, but is smaller. We've seen this with one dimensional Gaussians. Multiplying two Gaussians makes the variance smaller because we are incorporating more information, hence we are less uncertain. Another point to recognize is that the covariance shape reflects the physical layout of the aircraft and the radar system. The importance of this will become clear in the next step.
Now let's say we get a measurement from a second radar, this one to the lower right. The posterior from the last step becomes our new prior, which I plot in yellow. The new measurement is plotted in green.
```
P3 = [[2, -1.9], [-1.9, 2.2]]
plot_covariance_ellipse((10, 10), P2, ec='k', fc='y', alpha=0.6)
plot_covariance_ellipse((10, 10), P3, ec='k', fc='g', alpha=0.6)
```
We incorporate this information by multiplying the Gaussians:
```
P4 = multivariate_multiply((10, 10), P2, (10, 10), P3)[1]
plot_covariance_ellipse((10, 10), P2, ec='k', fc='y', alpha=0.6)
plot_covariance_ellipse((10, 10), P3, ec='k', fc='g', alpha=0.6)
plot_covariance_ellipse((10, 10), P4, ec='k', fc='b')
```
The only likely place for the aircraft is where the two ellipses intersect. The intersection, formed by multiplying the prior and measurement, is a new Gaussian. The shapes reflects the geometry of the problem. This allows us to *triangulate* on the aircraft, resulting in a very accurate estimate. We didn't explicitly write any code to perform triangulation; it was a natural outcome of multiplying the Gaussians of each measurement together.
Think back to the **g-h Filter** chapter where we displayed the error bars of two weighings on a scale. The estimate must fall somewhere within the region where the error bars overlap. Here the estimate must fall between 161 to 163 pounds.
```
import kf_book.book_plots as book_plots
book_plots.plot_errorbars([(160, 8, 'A'), (170, 8, 'B')], xlims=(150, 180))
```
Let's consider a different layout. Suppose the first radar is directly to the left of the aircraft. I can model the measurement error with
$$\Sigma = \begin{bmatrix}2&0\\0&0.2\end{bmatrix}$$
Here we see the result of multiplying the prior with the measurement.
```
P1 = [[2, 0], [0, .2]]
P2 = multivariate_multiply((10, 10), P0, (10, 10), P1)[1]
plot_covariance_ellipse((10, 10), P0, ec='k', fc='y', alpha=0.2)
plot_covariance_ellipse((10, 10), P1, ec='k', fc='g', alpha=0.6)
plot_covariance_ellipse((10, 10), P2, ec='k', fc='b')
```
Now we can incorporate the measurement from the second radar system, which we will leave in the same position as before.
```
P3 = [[2, -1.9], [-1.9, 2.2]]
P4 = multivariate_multiply((10, 10), P2, (10, 10), P3)[1]
plot_covariance_ellipse((10, 10), P2, ec='k', fc='y', alpha=0.2)
plot_covariance_ellipse((10, 10), P3, ec='k', fc='g', alpha=0.6)
plot_covariance_ellipse((10, 10), P4, ec='k', fc='b')
```
Our estimate is not as accurate as the previous example. The two radar stations are no longer orthogonal to each other relative to the aircraft's position so the triangulation is not optimal.
For a final example, imagine taking two measurements from the same radar a short time apart. The covariance ellipses will nearly overlap, leaving a very large error in our new estimate:
```
P5 = multivariate_multiply((10,10), P2, (10.1, 9.97), P2)
plot_covariance_ellipse((10, 10), P2, ec='k', fc='y', alpha=0.2)
plot_covariance_ellipse((10.1, 9.97), P2, ec='k', fc='g', alpha=0.6)
plot_covariance_ellipse(P5[0], P5[1], ec='k', fc='b')
```
## Hidden Variables
You can already see why a multivariate Kalman filter can perform better than a univariate one. Correlations between variables can significantly improve our estimates. We can take this much further. **This section contains the key insight to this chapter, so read carefully**.
Let's say we are tracking an aircraft and we get the following data for the $x$ and $y$ coordinates at time $t$=1, 2, and 3 seconds. What does your intuition tell you the value of $x$ will be at time $t$=4 seconds?
```
mkf_internal.show_position_chart()
```
It appears that the aircraft is flying in a straight line and we know that aircraft cannot turn on a dime. The most reasonable guess is that at $t$=4 the aircraft is at (4,4). I will depict that with a green arrow.
```
mkf_internal.show_position_prediction_chart()
```
You made this inference because you *inferred* a constant velocity for the airplane. The reasonable
assumption is that the aircraft is moving one unit each in *x* and *y* per time step.
Think back to the **g-h Filter** chapter when we were trying to improve the weight predictions of a noisy scale. We incorporated *weight gain* into the equations because it allowed us to make a better prediction of the weight the next day. The g-h filter uses the $g$ parameter to scale the amount of significance given to the current weight measurement, and the $h$ parameter scaled the amount of significance given to the weight gain.
We are going to do the same thing with our Kalman filter. After all, the Kalman filter is a form of a g-h filter. In this case we are tracking an airplane, so instead of weight and weight gain we need to track position and velocity. Weight gain is the *derivative* of weight, and of course velocity is the derivative of position. It's impossible to plot and understand the 4D chart that would be needed to plot *x* and *y* and their respective velocities so let's do it for $x$, knowing that the math generalizes to more dimensions.
At time 1 we might be fairly certain about the position (x=0) but have no idea about the velocity. We can plot that with a covariance matrix like this. The narrow width expresses our relative certainty about position, and the tall height expresses our lack of knowledge about velocity.
```
mkf_internal.show_x_error_chart(1)
```
Now after one second we get a position update of x=5.
```
mkf_internal.show_x_error_chart(2)
```
This implies that our velocity is roughly 5 m/s. But of course position and velocity are correlated. If the velocity is 5 m/s the position would be 5, but if the velocity was 10 m/s the position would be 10. So let's draw a covariance matrix in red showing the relationship between the position and velocity.
```
mkf_internal.show_x_error_chart(3)
```
It won't be clear until the next chapter how I calculate this covariance. Ignore the calculation, and think about what this implies. We have no easy way to say where the object really is because we are so uncertain about the velocity. Hence the ellipse stretches very far in the x-axis. Our uncertainty in velocity of course means it is also very spread in the y-axis. But as I said in the last paragraph, position is correlated to velocity. If the velocity is 5 m/s the next position would be 5, and if the velocity is 10 the next position would be 10. They are very correlated, so the ellipse must be very narrow.
This superposition of the two covariances is where the magic happens. The only reasonable estimate at time t=1 (where position=5) is roughly the intersection between the two covariance matrices! More exactly, we can use the math from the last section and multiply the two covariances together. From a Bayesian point of view we multiply the prior with the probability of the evidence (the *likelihood*) to get the posterior. If we multiply the position covariance with the velocity covariance using the Bayesian equations we get this result:
```
mkf_internal.show_x_error_chart(4)
```
The new covariance (the posterior) lies at the intersection of the position covariance and the velocity covariance. It is slightly tilted, showing that there is some correlation between the position and velocity. Far more importantly, it is much smaller than either the position or velocity covariances. In the previous chapter our variance would get smaller each time we performed an `update()` because the previous estimate was multiplied by the new measurement. The same happens here. However, here the improvement is markedly better. This is because we are using two different pieces of information which are nevertheless correlated. Knowing the velocity approximately and their correlation and the position approximately allows us to make a very accurate estimate.
This is a key point, so read carefully! The radar is only detecting the position of the aircraft. This is called an *observed variable*. Based on the position estimates we can compute velocity. We call the velocity a *hidden variable*. Hidden means what it sounds like - there is no sensor that is measuring velocity, thus its value is hidden from us. We are able to use the correlation between position and velocity to infer its value very accurately.
To round out the terminology there are also *unobserved variables*. For example, the aircraft's state includes things such as as heading, engine RPM, weight, color, the first name of the pilot, and so on. We cannot sense these directly using the position sensor so they are not *observed*. There is no way to *infer* them from the sensor measurements and correlations (red planes don't go faster than white planes), so they are not *hidden*. Instead, they are *unobservable*. If you include an unobserved variable in your filter state the estimate for that variable will be nonsense.
What makes this possible? Imagine for a moment that we superimposed the velocity from a different airplane over the position graph. Clearly the two are not related, and there is no way that combining the two could possibly yield any additional information. In contrast, the velocity of this airplane tells us something very important - the direction and speed of travel. So long as the aircraft does not alter its velocity the velocity allows us to predict where the next position is. After a relatively small amount of error in velocity the probability that it is a good match with the position is very small. Think about it - if you suddenly change direction your position is also going to change a lot. If the measurement of the position is not in the direction of the velocity change it is very unlikely to be true. The two are correlated, so if the velocity changes so must the position, and in a predictable way.
It is important to understand that we are taking advantage of the fact that velocity and position are correlated. We get a rough estimate of velocity from the distance and time between two measurements, and use Bayes theorem to produce very accurate estimates after only a few observations. Please reread this section if you have any doubts. If you do not understand this you will quickly find it impossible to reason about what you will learn in the following chapters.
## Higher Dimensions
So far I have shown you two dimensional Gaussians, but the math does not limit you to two dimensions. In later chapters we will be working in 9, or even 12 dimensions. If you work in areas such as weather prediction, you can end up with thousands of dimensions.
What do these higher dimensions 'look like? Well, a two dimensional Gaussian can be represented by an error ellipse, so it stands to reason a three dimensional Gaussian could be represented by a 3D error ellipsoid. We won't delve into the math here, but this turns out to be true. `FilterPy` provides a function to plot this ellipse.
First, let's make some noisy data with a given covariance, just so we can plot it inside the ellipsoid.
```
from filterpy.stats import plot_3d_covariance
mu = [0.3, 5., 10.]
C = np.array([[1.0, .03, .2],
[.03, 4.0, .0],
[.2, .0, 16.1]])
sample = np.random.multivariate_normal(mu, C, size=1000)
```
Now we plot the ellipsoid with the `FilterPy` function `plot_3d_covariance`, and then scatter plot the samples.
```
ax = plot_3d_covariance(mu, C, alpha=.4, std=3, limit_xyz=True)
ax.scatter(sample[:, 0], sample[:, 1], zs=sample[:, 2],);
```
Theory states that roughly 99% of a distribution will fall withing 3 standard deviations, and this appears to be the case.
Nine dimensions? I haven't quite figured out how to plot a 9D ellipsoid on a 2D screen, so there will be no graphs. The concept is the same; the standard deviation error of the distribution can be described by a 9D ellipsoid.
## Summary
We have taken advantage of the geometry and correlations of the system to produce a very accurate estimate. The math does not care whether we are working with two positions, or a position and a correlated velocity, or if these are spatial dimensions. If floor space is correlated to house price you can write a Kalman filter to track house prices. If age is correlated to disease incidence you can write a Kalman filter to track diseases. If the zombie population is inversely correlated with the number of shotguns then you can write a Kalman filter to track zombie populations. I showed you this in terms of geometry and talked about *triangulation*. That was just to build your intuition. You can write a Kalman filter for state variables that have no geometric representation, such as filters for stock prices or milk production of cows (I received an email from someone tracking milk production!) Get used to thinking of these as Gaussians with correlations. If we can express our uncertainties as a multidimensional Gaussian we can then multiply the prior with the likelihood and get a much more accurate result.
## References
- [1] http://docs.scipy.org/doc/scipy/reference/tutorial/stats.html
- [2] `FilterPy` library. Roger Labbe.
https://github.com/rlabbe/filterpy
| github_jupyter |
$\newcommand{\vct}[1]{\boldsymbol{#1}}
\newcommand{\mtx}[1]{\mathbf{#1}}
\newcommand{\tr}{^\mathrm{T}}
\newcommand{\reals}{\mathbb{R}}
\newcommand{\lpa}{\left(}
\newcommand{\rpa}{\right)}
\newcommand{\lsb}{\left[}
\newcommand{\rsb}{\right]}
\newcommand{\lbr}{\left\lbrace}
\newcommand{\rbr}{\right\rbrace}
\newcommand{\fset}[1]{\lbr #1 \rbr}
\newcommand{\pd}[2]{\frac{\partial #1}{\partial #2}}$
# Single layer models
In this lab we will implement a single-layer network model consisting of solely of an affine transformation of the inputs. The relevant material for this was covered in [the slides of the first lecture](http://www.inf.ed.ac.uk/teaching/courses/mlp/2016/mlp01-intro.pdf).
We will first implement the forward propagation of inputs to the network to produce predicted outputs. We will then move on to considering how to use gradients of an error function evaluated on the outputs to compute the gradients with respect to the model parameters to allow us to perform an iterative gradient-descent training procedure. In the final exercise you will use an interactive visualisation to explore the role of some of the different hyperparameters of gradient-descent based training methods.
#### A note on random number generators
It is generally a good practice (for machine learning applications **not** for cryptography!) to seed a pseudo-random number generator once at the beginning of each experiment. This makes it easier to reproduce results as the same random draws will produced each time the experiment is run (e.g. the same random initialisations used for parameters). Therefore generally when we need to generate random values during this course, we will create a seeded random number generator object as we do in the cell below.
```
import numpy as np
seed = 27092016
rng = np.random.RandomState(seed)
```
## Exercise 1: linear and affine transforms
Any *linear transform* (also called a linear map) of a finite-dimensional vector space can be parametrised by a matrix. So for example if we consider $\vct{x} \in \reals^{D}$ as the input space of a model with $D$ dimensional real-valued inputs, then a matrix $\mtx{W} \in \reals^{K\times D}$ can be used to define a prediction model consisting solely of a linear transform of the inputs
\begin{equation}
\vct{y} = \mtx{W} \vct{x}
\qquad
\Leftrightarrow
\qquad
y_k = \sum_{d=1}^D \lpa W_{kd} x_d \rpa \quad \forall k \in \fset{1 \dots K}
\end{equation}
with here $\vct{y} \in \reals^K$ the $K$-dimensional real-valued output of the model. Geometrically we can think of a linear transform doing some combination of rotation, scaling, reflection and shearing of the input.
An *affine transform* consists of a linear transform plus an additional translation parameterised by a vector $\vct{b} \in \reals^K$. A model consisting of an affine transformation of the inputs can then be defined as
\begin{equation}
\vct{y} = \mtx{W}\vct{x} + \vct{b}
\qquad
\Leftrightarrow
\qquad
y_k = \sum_{d=1}^D \lpa W_{kd} x_d \rpa + b_k \quad \forall k \in \fset{1 \dots K}
\end{equation}
In machine learning we will usually refer to the matrix $\mtx{W}$ as a *weight matrix* and the vector $\vct{b}$ as a *bias vector*.
Generally rather than working with a single data vector $\vct{x}$ we will work with batches of datapoints $\fset{\vct{x}^{(b)}}_{b=1}^B$. We could calculate the outputs for each input in the batch sequentially
\begin{align}
\vct{y}^{(1)} &= \mtx{W}\vct{x}^{(1)} + \vct{b}\\
\vct{y}^{(2)} &= \mtx{W}\vct{x}^{(2)} + \vct{b}\\
\dots &\\
\vct{y}^{(B)} &= \mtx{W}\vct{x}^{(B)} + \vct{b}\\
\end{align}
by looping over each input in the batch and calculating the output. However in general loops in Python are slow (particularly compared to compiled and typed languages such as C). This is due at least in part to the large overhead in dynamically inferring variable types. In general therefore wherever possible we want to avoid having loops in which such overhead will become the dominant computational cost.
For array based numerical operations, one way of overcoming this bottleneck is to *vectorise* operations. NumPy `ndarrays` are typed arrays for which operations such as basic elementwise arithmetic and linear algebra operations such as computing matrix-matrix or matrix-vector products are implemented by calls to highly-optimised compiled libraries. Therefore if you can implement code directly using NumPy operations on arrays rather than by looping over array elements it is often possible to make very substantial performance gains.
As a simple example we can consider adding up two arrays `a` and `b` and writing the result to a third array `c`. First lets initialise `a` and `b` with arbitrary values by running the cell below.
```
size = 1000
a = np.arange(size)
b = np.ones(size)
```
Now let's time how long it takes to add up each pair of values in the two array and write the results to a third array using a loop-based implementation. We will use the `%%timeit` magic briefly mentioned in the previous lab notebook specifying the number of times to loop the code as 100 and to give the best of 3 repeats. Run the cell below to get a print out of the average time taken.
```
%%timeit -n 100 -r 3
c = np.empty(size)
for i in range(size):
c[i] = a[i] + b[i]
```
And now we will perform the corresponding summation with the overloaded addition operator of NumPy arrays. Again run the cell below to get a print out of the average time taken.
```
%%timeit -n 100 -r 3
c = a + b
```
The first loop-based implementation should have taken on the order of milliseconds ($10^{-3}$s) while the vectorised implementation should have taken on the order of microseconds ($10^{-6}$s), i.e. a $\sim1000\times$ speedup. Hopefully this simple example should make it clear why we want to vectorise operations whenever possible!
Getting back to our affine model, ideally rather than individually computing the output corresponding to each input we should compute the outputs for all inputs in a batch using a vectorised implementation. As you saw last week, data providers return batches of inputs as arrays of shape `(batch_size, input_dim)`. In the mathematical notation used earlier we can consider this as a matrix $\mtx{X}$ of dimensionality $B \times D$, and in particular
\begin{equation}
\mtx{X} = \lsb \vct{x}^{(1)} ~ \vct{x}^{(2)} ~ \dots ~ \vct{x}^{(B)} \rsb\tr
\end{equation}
i.e. the $b^{\textrm{th}}$ input vector $\vct{x}^{(b)}$ corresponds to the $b^{\textrm{th}}$ row of $\mtx{X}$. If we define the $B \times K$ matrix of outputs $\mtx{Y}$ similarly as
\begin{equation}
\mtx{Y} = \lsb \vct{y}^{(1)} ~ \vct{y}^{(2)} ~ \dots ~ \vct{y}^{(B)} \rsb\tr
\end{equation}
then we can express the relationship between $\mtx{X}$ and $\mtx{Y}$ using [matrix multiplication](https://en.wikipedia.org/wiki/Matrix_multiplication) and addition as
\begin{equation}
\mtx{Y} = \mtx{X} \mtx{W}\tr + \mtx{B}
\end{equation}
where $\mtx{B} = \lsb \vct{b} ~ \vct{b} ~ \dots ~ \vct{b} \rsb\tr$ i.e. a $B \times K$ matrix with each row corresponding to the bias vector. The weight matrix needs to be transposed here as the inner dimensions of a matrix multiplication must match i.e. for $\mtx{C} = \mtx{A} \mtx{B}$ then if $\mtx{A}$ is of dimensionality $K \times L$ and $\mtx{B}$ is of dimensionality $M \times N$ then it must be the case that $L = M$ and $\mtx{C}$ will be of dimensionality $K \times N$.
The first exercise for this lab is to implement *forward propagation* for a single-layer model consisting of an affine transformation of the inputs in the `fprop` function given as skeleton code in the cell below. This should work for a batch of inputs of shape `(batch_size, input_dim)` producing a batch of outputs of shape `(batch_size, output_dim)`.
You will probably want to use the NumPy `dot` function and [broadcasting features](http://docs.scipy.org/doc/numpy/user/basics.broadcasting.html) to implement this efficiently. If you are not familiar with either / both of these you may wish to read the [hints](#Hints:-Using-the-dot-function-and-broadcasting) section below which gives some details on these before attempting the exercise.
```
def fprop(inputs, weights, biases):
"""Forward propagates activations through the layer transformation.
For inputs `x`, outputs `y`, weights `W` and biases `b` the layer
corresponds to `y = W x + b`.
Args:
inputs: Array of layer inputs of shape (batch_size, input_dim).
weights: Array of weight parameters of shape
(output_dim, input_dim).
biases: Array of bias parameters of shape (output_dim, ).
Returns:
outputs: Array of layer outputs of shape (batch_size, output_dim).
"""
return inputs.dot(weights.T) + biases
```
Once you have implemented `fprop` in the cell above you can test your implementation by running the cell below.
```
inputs = np.array([[0., -1., 2.], [-6., 3., 1.]])
weights = np.array([[2., -3., -1.], [-5., 7., 2.]])
biases = np.array([5., -3.])
true_outputs = np.array([[6., -6.], [-17., 50.]])
if not np.allclose(fprop(inputs, weights, biases), true_outputs):
print('Wrong outputs computed.')
else:
print('All outputs correct!')
```
### Hints: Using the `dot` function and broadcasting
For those new to NumPy below are some details on the `dot` function and broadcasting feature of NumPy that you may want to use for implementing the first exercise. If you are already familiar with these and have already completed the first exercise you can move on straight to [second exercise](#Exercise-2:-visualising-random-models).
#### `numpy.dot` function
Matrix-matrix, matrix-vector and vector-vector (dot) products can all be computed in NumPy using the [`dot`](http://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html) function. For example if `A` and `B` are both two dimensional arrays, then `C = np.dot(A, B)` or equivalently `C = A.dot(B)` will both compute the matrix product of `A` and `B` assuming `A` and `B` have compatible dimensions. Similarly if `a` and `b` are one dimensional arrays then `c = np.dot(a, b)` / `c = a.dot(b)` will compute the [scalar / dot product](https://en.wikipedia.org/wiki/Dot_product) of the two arrays. If `A` is a two-dimensional array and `b` a one-dimensional array `np.dot(A, b)` / `A.dot(b)` will compute the matrix-vector product of `A` and `b`. Examples of all three of these product types are shown in the cell below:
```
# Initiliase arrays with arbitrary values
A = np.arange(9).reshape((3, 3))
B = np.ones((3, 3)) * 2
a = np.array([-1., 0., 1.])
b = np.array([0.1, 0.2, 0.3])
print(A.dot(B)) # Matrix-matrix product
print(B.dot(A)) # Reversed product of above A.dot(B) != B.dot(A) in general
print(A.dot(b)) # Matrix-vector product
print(b.dot(A)) # Again A.dot(b) != b.dot(A) unless A is symmetric i.e. A == A.T
print(a.dot(b)) # Vector-vector scalar product
```
#### Broadcasting
Another NumPy feature it will be helpful to get familiar with is [broadcasting](http://docs.scipy.org/doc/numpy/user/basics.broadcasting.html). Broadcasting allows you to apply operations to arrays of different shapes, for example to add a one-dimensional array to a two-dimensional array or multiply a multidimensional array by a scalar. The complete set of rules for broadcasting as explained in the official documentation page just linked to can sound a bit complex: you might find the [visual explanation on this page](http://www.scipy-lectures.org/intro/numpy/operations.html#broadcasting) more intuitive. The cell below gives a few examples:
```
# Initiliase arrays with arbitrary values
A = np.arange(6).reshape((3, 2))
b = np.array([0.1, 0.2])
c = np.array([-1., 0., 1.])
print(A + b) # Add b elementwise to all rows of A
print((A.T + c).T) # Add b elementwise to all columns of A
print(A * b) # Multiply each row of A elementise by b
```
## Exercise 2: visualising random models
In this exercise you will use your `fprop` implementation to visualise the outputs of a single-layer affine transform model with two-dimensional inputs and a one-dimensional output. In this simple case we can visualise the joint input-output space on a 3D axis.
For this task and the learning experiments later in the notebook we will use a regression dataset from the [UCI machine learning repository](http://archive.ics.uci.edu/ml/index.html). In particular we will use a version of the [Combined Cycle Power Plant dataset](http://archive.ics.uci.edu/ml/datasets/Combined+Cycle+Power+Plant), where the task is to predict the energy output of a power plant given observations of the local ambient conditions (e.g. temperature, pressure and humidity).
The original dataset has four input dimensions and a single target output dimension. We have preprocessed the dataset by [whitening](https://en.wikipedia.org/wiki/Whitening_transformation) it, a common preprocessing step. We will only use the first two dimensions of the whitened inputs (corresponding to the first two principal components of the inputs) so we can easily visualise the joint input-output space.
The dataset has been wrapped in the `CCPPDataProvider` class in the `mlp.data_providers` module and the data included as a compressed file in the data directory as `ccpp_data.npz`. Running the cell below will initialise an instance of this class, get a single batch of inputs and outputs and import the necessary `matplotlib` objects.
```
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from mlp.data_providers import CCPPDataProvider
%matplotlib notebook
data_provider = CCPPDataProvider(
which_set='train',
input_dims=[0, 1],
batch_size=5000,
max_num_batches=1,
shuffle_order=False
)
input_dim, output_dim = 2, 1
inputs, targets = data_provider.next()
```
Here we used the `%matplotlib notebook` magic command rather than the `%matplotlib inline` we used in the previous lab as this allows us to produce interactive 3D plots which you can rotate and zoom in/out by dragging with the mouse and scrolling the mouse-wheel respectively. Once you have finished interacting with a plot you can close it to produce a static inline plot using the <i class="fa fa-power-off"></i> button in the top-right corner.
Now run the cell below to plot the predicted outputs of a randomly initialised model across the two dimensional input space as well as the true target outputs. This sort of visualisation can be a useful method (in low dimensions) to assess how well the model is likely to be able to fit the data and to judge appropriate initialisation scales for the parameters. Each time you re-run the cell a new set of random parameters will be sampled
Some questions to consider:
* How do the weights and bias initialisation scale affect the sort of predicted input-output relationships?
* Magnitude of weights initialisation scale determines how steep along the two input directions the plane predictions lie on is. The magnitude of the bias initialisation scale determines the typical offset of the plane from the `output = 0.` plane.
* Does the linear form of the model seem appropriate for the data here?
* While it appears a linear-model will not be able to fully capture the input-output relationship evident in the data with there some degree of non-linearity seeming to be present, as a first approximation a linear model seems a reasonable choice as a simple model for the data.
```
weights_init_range = 0.5
biases_init_range = 0.1
# Randomly initialise weights matrix
weights = rng.uniform(
low=-weights_init_range,
high=weights_init_range,
size=(output_dim, input_dim)
)
# Randomly initialise biases vector
biases = rng.uniform(
low=-biases_init_range,
high=biases_init_range,
size=output_dim
)
# Calculate predicted model outputs
outputs = fprop(inputs, weights, biases)
# Plot target and predicted outputs against inputs on same axis
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(111, projection='3d')
ax.plot(inputs[:, 0], inputs[:, 1], targets[:, 0], 'r.', ms=2)
ax.plot(inputs[:, 0], inputs[:, 1], outputs[:, 0], 'b.', ms=2)
ax.set_xlabel('Input dim 1')
ax.set_ylabel('Input dim 2')
ax.set_zlabel('Output')
ax.legend(['Targets', 'Predictions'], frameon=False)
fig.tight_layout()
```
## Exercise 3: computing the error function and its gradient
Here we will consider the task of regression as covered in the first lecture slides. The aim in a regression problem is given inputs $\fset{\vct{x}^{(n)}}_{n=1}^N$ to produce outputs $\fset{\vct{y}^{(n)}}_{n=1}^N$ that are as 'close' as possible to a set of target outputs $\fset{\vct{t}^{(n)}}_{n=1}^N$. The measure of 'closeness' or distance between target and predicted outputs is a design choice.
A very common choice is the squared Euclidean distance between the predicted and target outputs. This can be computed as the sum of the squared differences between each element in the target and predicted outputs. A common convention is to multiply this value by $\frac{1}{2}$ as this gives a slightly nicer expression for the error gradient. The error for the $n^{\textrm{th}}$ training example is then
\begin{equation}
E^{(n)} = \frac{1}{2} \sum_{k=1}^K \lbr \lpa y^{(n)}_k - t^{(n)}_k \rpa^2 \rbr.
\end{equation}
The overall error is then the *average* of this value across all training examples
\begin{equation}
\bar{E} = \frac{1}{N} \sum_{n=1}^N \lbr E^{(n)} \rbr.
\end{equation}
*Note here we are using a slightly different convention from the lectures. There the overall error was considered to be the sum of the individual error terms rather than the mean. To differentiate between the two we will use $\bar{E}$ to represent the average error here as opposed to sum of errors $E$ as used in the slides with $\bar{E} = \frac{E}{N}$. Normalising by the number of training examples is helpful to do in practice as this means we can more easily compare errors across data sets / batches of different sizes, and more importantly it means the size of our gradient updates will be independent of the number of training examples summed over.*
The regression problem is then to find parameters of the model which minimise $\bar{E}$. For our simple single-layer affine model here that corresponds to finding weights $\mtx{W}$ and biases $\vct{b}$ which minimise $\bar{E}$.
As mentioned in the lecture, for this simple case there is actually a closed form solution for the optimal weights and bias parameters. This is the linear least-squares solution those doing MLPR will have come across.
However in general we will be interested in models where closed form solutions do not exist. We will therefore generally use iterative, gradient descent based training methods to find parameters which (locally) minimise the error function. A basic requirement of being able to do gradient-descent based training is (unsuprisingly) the ability to evaluate gradients of the error function.
In the next exercise we will consider how to calculate gradients of the error function with respect to the model parameters $\mtx{W}$ and $\vct{b}$, but as a first step here we will consider the gradient of the error function with respect to the model outputs $\fset{\vct{y}^{(n)}}_{n=1}^N$. This can be written
\begin{equation}
\pd{\bar{E}}{\vct{y}^{(n)}} = \frac{1}{N} \lpa \vct{y}^{(n)} - \vct{t}^{(n)} \rpa
\qquad \Leftrightarrow \qquad
\pd{\bar{E}}{y^{(n)}_k} = \frac{1}{N} \lpa y^{(n)}_k - t^{(n)}_k \rpa \quad \forall k \in \fset{1 \dots K}
\end{equation}
i.e. the gradient of the error function with respect to the $n^{\textrm{th}}$ model output is just the difference between the $n^{\textrm{th}}$ model and target outputs, corresponding to the $\vct{\delta}^{(n)}$ terms mentioned in the lecture slides.
The third exercise is, using the equations given above, to implement functions computing the mean sum of squared differences error and its gradient with respect to the model outputs. You should implement the functions using the provided skeleton definitions in the cell below.
```
def error(outputs, targets):
"""Calculates error function given a batch of outputs and targets.
Args:
outputs: Array of model outputs of shape (batch_size, output_dim).
targets: Array of target outputs of shape (batch_size, output_dim).
Returns:
Scalar error function value.
"""
return 0.5 * ((outputs - targets)**2).sum() / outputs.shape[0]
def error_grad(outputs, targets):
"""Calculates gradient of error function with respect to model outputs.
Args:
outputs: Array of model outputs of shape (batch_size, output_dim).
targets: Array of target outputs of shape (batch_size, output_dim).
Returns:
Gradient of error function with respect to outputs.
This will be an array of shape (batch_size, output_dim).
"""
return (outputs - targets) / outputs.shape[0]
```
Check your implementation by running the test cell below.
```
outputs = np.array([[1., 2.], [-1., 0.], [6., -5.], [-1., 1.]])
targets = np.array([[0., 1.], [3., -2.], [7., -3.], [1., -2.]])
true_error = 5.
true_error_grad = np.array([[0.25, 0.25], [-1., 0.5], [-0.25, -0.5], [-0.5, 0.75]])
if not error(outputs, targets) == true_error:
print('Error calculated incorrectly.')
elif not np.allclose(error_grad(outputs, targets), true_error_grad):
print('Error gradient calculated incorrectly.')
else:
print('Error function and gradient computed correctly!')
```
## Exercise 4: computing gradients with respect to the parameters
In the previous exercise you implemented a function computing the gradient of the error function with respect to the model outputs. For gradient-descent based training, we need to be able to evaluate the gradient of the error function with respect to the model parameters.
Using the [chain rule for derivatives](https://en.wikipedia.org/wiki/Chain_rule#Higher_dimensions) we can write the partial deriviative of the error function with respect to single elements of the weight matrix and bias vector as
\begin{equation}
\pd{\bar{E}}{W_{kj}} = \sum_{n=1}^N \lbr \pd{\bar{E}}{y^{(n)}_k} \pd{y^{(n)}_k}{W_{kj}} \rbr
\quad \textrm{and} \quad
\pd{\bar{E}}{b_k} = \sum_{n=1}^N \lbr \pd{\bar{E}}{y^{(n)}_k} \pd{y^{(n)}_k}{b_k} \rbr.
\end{equation}
From the definition of our model at the beginning we have
\begin{equation}
y^{(n)}_k = \sum_{d=1}^D \lbr W_{kd} x^{(n)}_d \rbr + b_k
\quad \Rightarrow \quad
\pd{y^{(n)}_k}{W_{kj}} = x^{(n)}_j
\quad \textrm{and} \quad
\pd{y^{(n)}_k}{b_k} = 1.
\end{equation}
Putting this together we get that
\begin{equation}
\pd{\bar{E}}{W_{kj}} =
\sum_{n=1}^N \lbr \pd{\bar{E}}{y^{(n)}_k} x^{(n)}_j \rbr
\quad \textrm{and} \quad
\pd{\bar{E}}{b_{k}} =
\sum_{n=1}^N \lbr \pd{\bar{E}}{y^{(n)}_k} \rbr.
\end{equation}
Although this may seem a bit of a roundabout way to get to these results, this method of decomposing the error gradient with respect to the parameters in terms of the gradient of the error function with respect to the model outputs and the derivatives of the model outputs with respect to the model parameters, will be key when calculating the parameter gradients of more complex models later in the course.
Your task in this exercise is to implement a function calculating the gradient of the error function with respect to the weight and bias parameters of the model given the already computed gradient of the error function with respect to the model outputs. You should implement this in the `grads_wrt_params` function in the cell below.
```
def grads_wrt_params(inputs, grads_wrt_outputs):
"""Calculates gradients with respect to model parameters.
Args:
inputs: array of inputs to model of shape (batch_size, input_dim)
grads_wrt_to_outputs: array of gradients of with respect to the model
outputs of shape (batch_size, output_dim).
Returns:
list of arrays of gradients with respect to the model parameters
`[grads_wrt_weights, grads_wrt_biases]`.
"""
grads_wrt_weights = grads_wrt_outputs.T.dot(inputs)
grads_wrt_biases = grads_wrt_outputs.sum(0)
return [grads_wrt_weights, grads_wrt_biases]
```
Check your implementation by running the test cell below.
```
inputs = np.array([[1., 2., 3.], [-1., 4., -9.]])
grads_wrt_outputs = np.array([[-1., 1.], [2., -3.]])
true_grads_wrt_weights = np.array([[-3., 6., -21.], [4., -10., 30.]])
true_grads_wrt_biases = np.array([1., -2.])
grads_wrt_weights, grads_wrt_biases = grads_wrt_params(
inputs, grads_wrt_outputs)
if not np.allclose(true_grads_wrt_weights, grads_wrt_weights):
print('Gradients with respect to weights incorrect.')
elif not np.allclose(true_grads_wrt_biases, grads_wrt_biases):
print('Gradients with respect to biases incorrect.')
else:
print('All parameter gradients calculated correctly!')
```
## Exercise 5: wrapping the functions into reusable components
In exercises 1, 3 and 4 you implemented methods to compute the predicted outputs of our model, evaluate the error function and its gradient on the outputs and finally to calculate the gradients of the error with respect to the model parameters. Together they constitute all the basic ingredients we need to implement a gradient-descent based iterative learning procedure for the model.
Although you could implement training code which directly uses the functions you defined, this would only be usable for this particular model architecture. In subsequent labs we will want to use the affine transform functions as the basis for more interesting multi-layer models. We will therefore wrap the implementations you just wrote in to reusable components that we can build more complex models with later in the course.
* In the [`mlp.layers`](/edit/mlp/layers.py) module, use your implementations of `fprop` and `grad_wrt_params` above to implement the corresponding methods in the skeleton `AffineLayer` class provided.
* In the [`mlp.errors`](/edit/mlp/errors.py) module use your implementation of `error` and `error_grad` to implement the `__call__` and `grad` methods respectively of the skeleton `SumOfSquaredDiffsError` class provided. Note `__call__` is a special Python method that allows an object to be used with a function call syntax.
Run the cell below to use your completed `AffineLayer` and `SumOfSquaredDiffsError` implementations to train a single-layer model using batch gradient descent on the CCCP dataset.
```
from mlp.layers import AffineLayer
from mlp.errors import SumOfSquaredDiffsError
from mlp.models import SingleLayerModel
from mlp.initialisers import UniformInit, ConstantInit
from mlp.learning_rules import GradientDescentLearningRule
from mlp.optimisers import Optimiser
import logging
# Seed a random number generator
seed = 27092016
rng = np.random.RandomState(seed)
# Set up a logger object to print info about the training run to stdout
logger = logging.getLogger()
logger.setLevel(logging.INFO)
logger.handlers = [logging.StreamHandler()]
# Create data provider objects for the CCPP training set
train_data = CCPPDataProvider('train', [0, 1], batch_size=100, rng=rng)
input_dim, output_dim = 2, 1
# Create a parameter initialiser which will sample random uniform values
# from [-0.1, 0.1]
param_init = UniformInit(-0.1, 0.1, rng=rng)
# Create our single layer model
layer = AffineLayer(input_dim, output_dim, param_init, param_init)
model = SingleLayerModel(layer)
# Initialise the error object
error = SumOfSquaredDiffsError()
# Use a basic gradient descent learning rule with a small learning rate
learning_rule = GradientDescentLearningRule(learning_rate=1e-2)
# Use the created objects to initialise a new Optimiser instance.
optimiser = Optimiser(model, error, learning_rule, train_data)
# Run the optimiser for 5 epochs (full passes through the training set)
# printing statistics every epoch.
stats, keys, run_time = optimiser.train(num_epochs=10, stats_interval=1)
# Plot the change in the error over training.
fig = plt.figure(figsize=(8, 4))
ax = fig.add_subplot(111)
ax.plot(np.arange(1, stats.shape[0] + 1), stats[:, keys['error(train)']])
ax.set_xlabel('Epoch number')
ax.set_ylabel('Error')
```
Using similar code to previously we can now visualise the joint input-output space for the trained model. If you implemented the required methods correctly you should now see a much improved fit between predicted and target outputs when running the cell below.
```
data_provider = CCPPDataProvider(
which_set='train',
input_dims=[0, 1],
batch_size=5000,
max_num_batches=1,
shuffle_order=False
)
inputs, targets = data_provider.next()
# Calculate predicted model outputs
outputs = model.fprop(inputs)[-1]
# Plot target and predicted outputs against inputs on same axis
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(111, projection='3d')
ax.plot(inputs[:, 0], inputs[:, 1], targets[:, 0], 'r.', ms=2)
ax.plot(inputs[:, 0], inputs[:, 1], outputs[:, 0], 'b.', ms=2)
ax.set_xlabel('Input dim 1')
ax.set_ylabel('Input dim 2')
ax.set_zlabel('Output')
ax.legend(['Targets', 'Predictions'], frameon=False)
fig.tight_layout()
```
## Exercise 6: visualising training trajectories in parameter space
Running the cell below will display an interactive widget which plots the trajectories of gradient-based training of the single-layer affine model on the CCPP dataset in the three dimensional parameter space (two weights plus bias) from random initialisations. Also shown on the right is a plot of the evolution of the error function (evaluated on the current batch) over training. By moving the sliders you can alter the training hyperparameters to investigate the effect they have on how training procedes.
Some questions to explore:
* Are there multiple local minima in parameter space here? Why?
* In this case there is a single unique global minima, as suggested by the fact random parameter initialisations consistently converge to the same point in parameter space. As mentioned previously there is a closed form solution for the optimal weights and biases for this simple single-layer affine model <a href='https://en.wikipedia.org/wiki/Linear_least_squares_(mathematics)'>linear least squares</a>) and the error function is [convex](https://en.wikipedia.org/wiki/Convex_function).
* What happens to learning for very small learning rates? And very large learning rates?
* For very small learning rates, the training proceeds very slowly and the parameters do not tend to converge to the global optimum unless a lot of training epochs are used. For very large learning rates, the gradient descent dynamic becomes increasingly instable, leading to large oscillations or at extreme values divergence in the parameter space.
* How does the batch size affect learning?
* Smaller batch sizes generally lead to quicker initial learning as the parameters are updated more frequently (more batches in an epoch) however as the batch becomes smaller the error estimate calculated from the batch and its gradients become increasingly noisy estimates for the true error function / gradients. This can be observed by the less smooth trajectories in parameter space for lower batch sizes and greater noise in the batch error curves.
**Note:** You don't need to understand how the code below works. The idea of this exercise is to help you understand the role of the various hyperparameters involved in gradient-descent based training methods.
```
from ipywidgets import interact
%matplotlib inline
def setup_figure():
# create figure and axes
fig = plt.figure(figsize=(12, 6))
ax1 = fig.add_axes([0., 0., 0.5, 1.], projection='3d')
ax2 = fig.add_axes([0.6, 0.1, 0.4, 0.8])
# set axes properties
ax2.spines['right'].set_visible(False)
ax2.spines['top'].set_visible(False)
ax2.yaxis.set_ticks_position('left')
ax2.xaxis.set_ticks_position('bottom')
ax2.set_yscale('log')
ax1.set_xlim((-2, 2))
ax1.set_ylim((-2, 2))
ax1.set_zlim((-2, 2))
#set axes labels and title
ax1.set_title('Parameter trajectories over training')
ax1.set_xlabel('Weight 1')
ax1.set_ylabel('Weight 2')
ax1.set_zlabel('Bias')
ax2.set_title('Batch errors over training')
ax2.set_xlabel('Batch update number')
ax2.set_ylabel('Batch error')
return fig, ax1, ax2
def visualise_training(n_epochs=1, batch_size=200, log_lr=-1., n_inits=5,
w_scale=1., b_scale=1., elev=30., azim=0.):
fig, ax1, ax2 = setup_figure()
# create seeded random number generator
rng = np.random.RandomState(1234)
# create data provider
data_provider = CCPPDataProvider(
input_dims=[0, 1],
batch_size=batch_size,
shuffle_order=False,
)
learning_rate = 10 ** log_lr
n_batches = data_provider.num_batches
weights_traj = np.empty((n_inits, n_epochs * n_batches + 1, 1, 2))
biases_traj = np.empty((n_inits, n_epochs * n_batches + 1, 1))
errors_traj = np.empty((n_inits, n_epochs * n_batches))
# randomly initialise parameters
weights = rng.uniform(-w_scale, w_scale, (n_inits, 1, 2))
biases = rng.uniform(-b_scale, b_scale, (n_inits, 1))
# store initial parameters
weights_traj[:, 0] = weights
biases_traj[:, 0] = biases
# iterate across different initialisations
for i in range(n_inits):
# iterate across epochs
for e in range(n_epochs):
# iterate across batches
for b, (inputs, targets) in enumerate(data_provider):
outputs = fprop(inputs, weights[i], biases[i])
errors_traj[i, e * n_batches + b] = error(outputs, targets)
grad_wrt_outputs = error_grad(outputs, targets)
weights_grad, biases_grad = grads_wrt_params(inputs, grad_wrt_outputs)
weights[i] -= learning_rate * weights_grad
biases[i] -= learning_rate * biases_grad
weights_traj[i, e * n_batches + b + 1] = weights[i]
biases_traj[i, e * n_batches + b + 1] = biases[i]
# choose a different color for each trajectory
colors = plt.cm.jet(np.linspace(0, 1, n_inits))
# plot all trajectories
for i in range(n_inits):
lines_1 = ax1.plot(
weights_traj[i, :, 0, 0],
weights_traj[i, :, 0, 1],
biases_traj[i, :, 0],
'-', c=colors[i], lw=2)
lines_2 = ax2.plot(
np.arange(n_batches * n_epochs),
errors_traj[i],
c=colors[i]
)
ax1.view_init(elev, azim)
plt.show()
w = interact(
visualise_training,
elev=(-90, 90, 2),
azim=(-180, 180, 2),
n_epochs=(1, 5),
batch_size=(100, 1000, 100),
log_lr=(-3., 1.),
w_scale=(0., 2.),
b_scale=(0., 2.),
n_inits=(1, 10)
)
for child in w.widget.children:
child.layout.width = '100%'
```
| github_jupyter |
```
# default_exp data.acquisition
```
# Data Acquisition
> This is a script which invokes `pybaseball`'s [`statcast()`](https://github.com/jldbc/pybaseball#statcast-pull-advanced-metrics-from-major-league-baseballs-statcast-system) function to retrieve pitch-level data from statcast.
```
#hide
# documentation
from nbdev.showdoc import *
# testing
import pytest
# exporti
from pybaseball import statcast
import pandas as pd
from fastscript import *
import sqlite3
from os import path
# export
@call_parse
def query_statcast(
start_dt: Param(help="Beginning date to pull data from", type=str) = None,
end_dt: Param(help="End date to pull data from", type=str) = None,
team: Param(help="Abbreviation for team of interest", type=str) = None,
verbose: Param(help="Whether or not to print verbose updates", type=bool_arg) = True,
output_type: Param(help="What format to save data in", type=str) = "db",
overwrite: Param(help="Whether or not to overwrite the db table if it already exists", type=bool_arg,) = False,
output_path: Param(help="path to location that data should be saved", type=str) = "."):
"""
Callable from the command-line or in Python. Pulls pitch-level MLB data from [statcast](https://baseballsavant.mlb.com/statcast_search).
Saves as either a sqlite db file, or csv.
* inputs:
- `start_dt`: `str`, Beginning date to pull data from = None
- `end_dt`: `str`, End date to pull data from = None
- `team`: `str`, abbreviation for team of interest = None
- `verbose`: `bool`, Whether or not to print verbose updates
- `output_type`: `str`, What format to save data in (must be one of {'db', 'csv'}) = 'db'
- `overwrite`: `bool`, Whether or not to overwrite the db table if it already exists = False
- `output_path`: `str`, Path to the location that the data should be saved at = '.'
* outputs:
- None
"""
# checking for correct output type
if output_type not in ("db", "csv"):
raise ValueError("output_type must be one of {'db', 'csv'}")
if output_type == "db":
# creating db connection
conn = sqlite3.connect(f"{output_path}/statcast_pitches.db")
# Checking if year is already in db
cursor = conn.execute(f"select name from sqlite_master where type='table' and name='statcast_{start_dt[:4]}'")
# if table exists in db
if cursor.fetchone():
if overwrite:
conn.execute(f"DROP TABLE IF EXISTS statcast_{start_dt[:4]}")
else:
# don't want to overwrite, pop out of function
print(f"Table named 'statcast_{start_dt[:4]}' already exists in db saved at `{output_path}/statcast_{start_dt[:4]}`.")
return None
# if table does not already exist in db or it was just dropped
# pulling data from statcast
data = statcast(start_dt=start_dt, end_dt=end_dt, team=team, verbose=verbose)
data.to_sql(f"statcast_{start_dt[:4]}", conn)
conn.close()
# output type is csv
else:
# Checking if file is already saved as csv
if path.exists(f"{output_path}/statcast_{start_dt[:4]}.csv"):
print(f"File named `{output_path}/statcast_{start_dt[:4]}.csv` already exists.")
return None
# pulling data from statcast
data = statcast(start_dt=start_dt, end_dt=end_dt, team=team, verbose=verbose)
# saving to csv
data.to_csv(f"{output_path}/statcast_{start_dt[:4]}.csv", index=False)
return None
! rm /tmp/*.db /tmp/*.pkl /tmp/*.csv
# output type must be db or csv
with pytest.raises(ValueError):
query_statcast(output_type=None)
# making sure a db file is created
output_path = "/tmp"
start_dt = end_dt = "2019-07-07"
query_statcast(
start_dt=start_dt, end_dt=end_dt, output_type="db", output_path=output_path
)
assert path.exists(f"{output_path}/statcast_pitches.db")
# making sure the db file will be over-written without error
query_statcast(
start_dt=start_dt,
end_dt=end_dt,
team="BOS",
output_type="db",
overwrite=True,
output_path=output_path,
)
# making sure db file will not be overwritten
query_statcast(
start_dt=start_dt,
end_dt=end_dt,
team="BOS",
output_type="db",
output_path=output_path,
)
# making sure a csv file is created
query_statcast(
start_dt=start_dt, end_dt=end_dt, output_type="csv", output_path=output_path
)
assert path.exists(f"{output_path}/statcast_{start_dt[:4]}.csv")
# export
def query_db(
db_path: str = "../data/raw/statcast_pitches.db",
year: str = "2019",
columns: str = "*",
limit: int = None,
verbose: bool = True,
):
"""
Queries a sqlite db file. Assumes that it's been created by `query_statcast`.
Only queries for a single year at a time.
* intputs:
- `db_path`: `str`, path that db file is located at
- `year`: `str`, year of data to query
- `columns`: `str`, which columns from the [statcast data](https://baseballsavant.mlb.com/csv-docs) to include in table
- `limit`: `int`, the maximum number of rows to retrieve ([postgresql documentation](https://www.postgresql.org/docs/8.1/queries-limit.html))
- `verbose`: `bool`, Whether or not to print verbose updates
* output:
- `df`: `pd.DataFrame`, DataFrame populated with data queried from database
"""
if verbose:
print(f"querying db at {db_path} now.")
conn = sqlite3.connect(db_path)
query = f"""select {columns}
from statcast_{year}"""
if limit:
query += f" limit {round(limit)}"
# making sure year is in db
cursor = conn.execute(f"select name from sqlite_master where type='table' and name='statcast_{year}'")
if cursor.fetchone():
df = pd.read_sql_query(query, conn)
else:
df = pd.DataFrame()
conn.close()
return df
# BOS @ DET on 7/7/19
db_path = f"{output_path}/statcast_pitches.db"
df = query_db(db_path=db_path)
assert df["away_team"].unique().item() == "BOS"
# checking consistent rows and columns (extra column because index is included)
assert df.shape == (339, 91)
# year not present in db gives empty DataFrame
df = query_db(db_path=db_path,
year="2012")
assert df.empty
# also testing that csv file is of expected size
df = pd.read_csv(f"{output_path}/statcast_{start_dt[:4]}.csv")
assert df.shape == (4457, 90)
# clean-up
! rm {output_path}/statcast_*
! ls /tmp
```
## Usage
### From the command-line
```shell
$ query_statcast --start_dt 2019-05-07 --end_dt 2019-06-09 --output_type db --output_path /tmp
This is a large query, it may take a moment to complete
Completed sub-query from 2019-05-07 to 2019-05-12
Completed sub-query from 2019-05-13 to 2019-05-18
Completed sub-query from 2019-05-19 to 2019-05-24
Completed sub-query from 2019-05-25 to 2019-05-30
Completed sub-query from 2019-05-31 to 2019-06-05
Completed sub-query from 2019-06-06 to 2019-06-09
$ ls /tmp/ | grep statcast_pitches
statcast_pitches.db
```
### Using Python
```python
>>> query_statcast(
start_dt="2019-06-07", end_dt="2019-06-09", output_type="csv", output_path="/tmp"
)
```
```shell
$ ls /tmp/ | grep statcast
```
| github_jupyter |
```
#hide
#default_exp vis.gen
```
# Visualisation Generation
<br>
### Imports
```
#exports
import json
import pandas as pd
import typer
import croniter
import importlib
from tqdm import tqdm
import matplotlib.pyplot as plt
from IPython.display import JSON
#exports
def rgb_2_plt_tuple(r, g, b):
"""converts a standard rgb set from a 0-255 range to 0-1"""
plt_tuple = tuple([x/255 for x in (r, g, b)])
return plt_tuple
vis_configs = [
{
'cron': '0 * * * 1', # midnight every monday
'function': 'ElexonDataPortal.vis.ei.generate_GB_decarb_progess',
'kwargs': {
'dt_col': 'local_datetime',
'dt_tz': 'Europe/London',
'url': 'https://api.github.com/repos/AyrtonB/Electric-Insights/git/trees/master?recursive=1',
'raw_file_prefix': 'https://raw.githubusercontent.com/AyrtonB/Electric-Insights/master/',
'update_time': pd.Timestamp.now().round('5min').strftime('%Y-%m-%d %H:%M'),
'dpi': 250,
'freq': '7D',
'use_preloaded_ei_df': True,
'fuel_colour_dict': {
'Imports & Storage' : rgb_2_plt_tuple(121,68,149),
'nuclear' : rgb_2_plt_tuple(77,157,87),
'biomass' : rgb_2_plt_tuple(168,125,81),
'gas' : rgb_2_plt_tuple(254,156,66),
'coal' : rgb_2_plt_tuple(122,122,122),
'hydro' : rgb_2_plt_tuple(50,120,196),
'wind' : rgb_2_plt_tuple(72,194,227),
'solar' : rgb_2_plt_tuple(255,219,65),
},
'docs_dir': 'docs',
'update_time': None,
}
},
{
'cron': '0 * * * 1', # midnight every monday
'function': 'ElexonDataPortal.vis.ei.generate_moe',
'kwargs': {
'dt_col': 'local_datetime',
'dt_tz': 'Europe/London',
'url': 'https://api.github.com/repos/AyrtonB/Electric-Insights/git/trees/master?recursive=1',
'raw_file_prefix': 'https://raw.githubusercontent.com/AyrtonB/Electric-Insights/master/',
'reg_dates_start': '2010-01-01',
'reg_dates_end': None,
'reg_dates_freq': '13W',
'num_fits': 15,
'x_pred': None,
'dt_idx': None,
'dpi': 250,
'use_preloaded_ei_df': True,
'img_name': 'moe_surface',
'docs_dir': 'docs',
'update_time': None,
}
},
{
'cron': '0,30 * * * *', # every half-hour
'function': 'ElexonDataPortal.vis.lolp.generate_lolpdrm_imgs_text',
'kwargs': {
'api_key': None,
'fcst_horizons': [8, 4, 2, 1],
'update_time': None,
'docs_dir': 'docs',
'update_time': None,
}
},
{
'cron': '0,30 * * * *', # every half-hour
'function': 'ElexonDataPortal.vis.map.generate_map',
'kwargs': {
'data_dir': 'data/PN',
'api_key': None,
'update_time': None,
'powerdict_url': 'https://raw.githubusercontent.com/OSUKED/Power-Station-Dictionary/main/data/output/power_stations.csv',
'js_template_fp': 'templates/map.js',
'js_docs_fp': 'docs/js/map.js',
'md_template_fp': 'templates/map.md',
'plants_geojson_fp': 'data/power_plants.json',
'plants_geojson_url': 'https://raw.githubusercontent.com/OSUKED/ElexonDataPortal/master/data/power_plants.json',
'routes_geojson_url': 'https://raw.githubusercontent.com/OSUKED/ElexonDataPortal/master/data/network_routes.json'
}
}
]
JSON(vis_configs)
save_vis_configs = False
if save_vis_configs == True:
with open('../data/vis_configs.json', 'w') as f:
json.dump(vis_configs, f)
#exports
def get_vis_func(func_path):
*lib_path, func_name = func_path.split('.')
lib_obj = importlib.import_module('.'.join(lib_path))
func = getattr(lib_obj, func_name)
return func
def get_vis_md_text(vis_config, docs_dir=None, update_time=None):
func_path = vis_config['function']
kwargs = vis_config['kwargs']
if (docs_dir is not None) and ('docs_dir' in kwargs.keys()):
kwargs['docs_dir'] = docs_dir
if (update_time is not None) and ('update_time' in kwargs.keys()):
kwargs['update_time'] = update_time
vis_func = get_vis_func(func_path)
vis_md_text = vis_func(**kwargs)
plt.close()
return vis_md_text
docs_dir = '../docs'
vis_config = vis_configs[0]
vis_md_text = get_vis_md_text(vis_config, docs_dir=docs_dir)
print(vis_md_text)
#exports
def get_rerun_vis_bool(vis_config):
if 'last_update_time' not in vis_config.keys():
return True
else:
last_update_time = pd.to_datetime(vis_config['last_update_time']).tz_localize('Europe/London')
cron = croniter.croniter(vis_config['cron'], pd.Timestamp.now()-pd.Timedelta(weeks=1))
cron_dts = pd.to_datetime([cron.get_next() for i in range(10*48*7)], unit='s').tz_localize('UTC').tz_convert('Europe/London')
s_cron_dts_time_delta_to_now = pd.Series((cron_dts - pd.Timestamp.now(tz='Europe/London')).total_seconds())
assert (s_cron_dts_time_delta_to_now<0).sum()>0 and (s_cron_dts_time_delta_to_now>0).sum()>0, 'The cron dts being assessed do not cross the current time'
s_cron_dts_time_delta_to_last_update_time = pd.Series((cron_dts - last_update_time).total_seconds())
if s_cron_dts_time_delta_to_now.abs().idxmin() == s_cron_dts_time_delta_to_last_update_time.abs().idxmin():
return False
avg_adj_dt_time_delta_s = pd.Series(cron_dts).diff(1).dropna().dt.total_seconds().mean()
min_time_delta_s = s_cron_dts_time_delta_to_now.abs().min()
rerun_vis = avg_adj_dt_time_delta_s >= min_time_delta_s
return rerun_vis
rerun_vis = get_rerun_vis_bool(vis_config)
rerun_vis
#exports
def update_vis_configs(
vis_configs,
docs_dir: str='docs',
override_rerun_vis_bool: bool=False
):
for i, vis_config in enumerate(vis_configs):
update_time = pd.Timestamp.now().round('5min').strftime('%Y-%m-%d %H:%M')
rerun_vis = get_rerun_vis_bool(vis_config)
if override_rerun_vis_bool == True:
rerun_vis = True
if rerun_vis == True:
vis_md_text = get_vis_md_text(vis_config, docs_dir=docs_dir, update_time=update_time)
vis_configs[i]['md_text'] = vis_md_text
vis_configs[i]['last_update_time'] = update_time
return vis_configs
docs_dir = '../docs'
data_dir = '../data'
with open(f'{data_dir}/vis_configs.json', 'r') as f:
vis_configs = json.load(f)
vis_configs = update_vis_configs(vis_configs, docs_dir=docs_dir)
with open(f'{data_dir}/vis_configs.json', 'w') as f:
json.dump(vis_configs, f)
# add in a mini example func template
all_vis_md_texts = ['# Visualisations'] + [vis_config['md_text'] for vis_config in vis_configs]
combined_md_text = '\n\n<br>\n\n'.join(all_vis_md_texts)
print(combined_md_text)
with open('../docs/visualisations.md', 'w', encoding='utf-8') as f:
f.write(combined_md_text)
```
<br>
`python -m ElexonDataPortal.vis.gen`
```
#exports
app = typer.Typer()
@app.command()
def update_vis(
docs_dir: str='docs',
data_dir: str='data',
override_rerun_vis_bool: bool=False
):
with open(f'{data_dir}/vis_configs.json', 'r') as f:
vis_configs = json.load(f)
vis_configs = update_vis_configs(vis_configs, docs_dir=docs_dir, override_rerun_vis_bool=override_rerun_vis_bool)
with open(f'{data_dir}/vis_configs.json', 'w') as f:
json.dump(vis_configs, f)
prefix_text = """# Visualisations
On this page you can view visualisations of key phenomena in the GB power sector, ranging from long-term trends in the generation-mix and market prices to information on excess capacity in the grid. All data used in these visualisations was either sourced directly from BMRS using the `ElexonDataPortal` client, or has been derived from BMRS data streams. As with the other components of the `ElexonDataPortal` the code to generate these visualisations is open-source and users are welcome to contribute their own visualisations, for more detail on how to do this please refer to the [user contribution guide](#contributor-guide)
"""
suffix_text = """### Contributor Guide
We encourage users to contribute their own visualisations which the `ElexonDataPortal` will then update automatically. To this end the library adopts a standardised format for generating visualisations, the core component of which is the `data/vis_configs.json` file to which you will have to add detail on your visualisation function:
```javascript
[
...
{
"cron": "0 * * * *", # the update schedule, in this instance to run at midnight every sunday
"function": "path_to_function", # e.g. ElexonDataPortal.vis.generate_vis
"kwargs": {
'api_key': null, # if no api_key is passed then the client will try and look for the `BMRS_API_KEY` environment variable
'update_time': null, # if no update_time is passed you should generate it yourself, e.g. with `pd.Timestamp.now().round('5min').strftime('%Y-%m-%d %H:%M')`
'docs_dir': 'docs', # in almost all circumstances this should just be `docs`
"optional_kwarg": "optional_value" # you can specify any additional keyword arguments that your function requires
}
},
...
]
```
<br>
The other core component is writing the function that generates the visualisation. This function should require parameters for the `docs_dir`, `api_key`, and `update_time` but can include optional parameters that you wish to specify, it should then return markdown text which will be used to populate the *Visualisations* page. These functions will normally contain three steps: data retrieval, generating the visualisation, and generating the accompanying text - an example can be seen below.
```python
import pandas as pd
import matplotlib.pyplot as plt
from ElexonDataPortal.api import Client
def generate_vis(
docs_dir: str='docs',
api_key: str=None,
update_time: str=pd.Timestamp.now().round('5min').strftime('%Y-%m-%d %H:%M'),
) -> str:
# Data Retrieval
client = Client(api_key=api_key)
df = client.get_data_stream(param1, param2)
# Generating the Visualisation
fig, ax = plt.subplots(dpi=150)
df.plot(ax=ax)
fig.savefig(f'{docs_dir}/img/vis/great_vis_name.png')
# Generating the Text
md_text = f\"\"\"### Title
Explanation of what your visualisation shows

\"\"\"
return md_text
```
N.b. the path to the image should be relative to the `docs` directory.
If you require any assistance in this process please start a discussion [here](https://github.com/OSUKED/ElexonDataPortal/discussions) and we'll endeavour to help as best we can.
"""
all_vis_md_texts = [prefix_text] + [vis_config['md_text'] for vis_config in vis_configs] + [suffix_text]
combined_md_text = '\n\n<br>\n\n'.join(all_vis_md_texts)
with open(f'{docs_dir}/visualisations.md', 'w', encoding='utf-8') as f:
f.write(combined_md_text)
return
if __name__ == '__main__' and '__file__' in globals():
app()
update_vis(docs_dir='../docs', data_dir='../data', override_rerun_vis_bool=True)
# need to write a func that will be run by the GH action
# should check the time between now and any cron jobs descs, then run those that are within 24hrs
# no cron jobs will run more frequently than every 24hrs
# How to contribute?
# * Create a function that writes an image to the `docs/img/vis` directory
# * Ensure that same function returns a markdown string which will render the desired text and images if loaded from the `docs` directory
# * Add the function name, schedule (in cron notation), and any kwargs to be passed to the function as a new item in the `data/vis_configs.json` file
#hide
from ElexonDataPortal.dev.nbdev import notebook2script
notebook2script('vis-00-gen.ipynb')
```
| github_jupyter |
SOP013 - Create secret for azdata login (inside cluster)
========================================================
Description
-----------
Create a secret in the Kubernetes Secret Store, to:
- Run app-deploys (i.e. `azdata app run`)
- Save results in HDFS at /app-deploy
- Enable SOP028 to perform `azdata login` when run from inside the Big
Data Cluster. This is needed for example, when running notebooks in
an app-deploy (which is inside a Big Data Cluster)
### Parameters
```
import os
azdata_login_username = "<INSERT USERNAME>"
azdata_login_password = "<INSERT PASSWORD>"
# If an Active Directory (secure) cluster, provide a domain account domain name, i.e. username@domain_name
#
azdata_login_domain_name = "<INSERT DOMAIN>" # This should be UPPER CASE
azdata_login_secret_name = "azdata-login-notebook-run-secret"
print("Parameters set for user name: "+ azdata_login_username)
```
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
def run(cmd, return_output=False, no_output=False, retry_count=0, base64_decode=False, return_as_json=False, regex_mask=None):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportability, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
# Display an install HINT, so the user can click on a SOP to install the missing binary
#
if which_binary == None:
print(f"The path used to search for '{cmd_actual[0]}' was:")
print(sys.path)
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
cmd_display = cmd
if regex_mask is not None:
regex = re.compile(regex_mask)
cmd_display = re.sub(regex, '******', cmd)
print(f"START: {cmd_display} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd_display} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd_display} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
# Hints for tool retry (on transient fault), known errors and install guide
#
retry_hints = {'azdata': ['Endpoint sql-server-master does not exist', 'Endpoint livy does not exist', 'Failed to get state for cluster', 'Endpoint webhdfs does not exist', 'Adaptive Server is unavailable or does not exist', 'Error: Address already in use', 'Login timeout expired (0) (SQLDriverConnect)', 'SSPI Provider: No Kerberos credentials available', ], 'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond', ], 'python': [ ], }
error_hints = {'azdata': [['Please run \'azdata login\' to first authenticate', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['The token is expired', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Reason: Unauthorized', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Max retries exceeded with url: /api/v1/bdc/endpoints', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Look at the controller logs for more details', 'TSG027 - Observe cluster deployment', '../diagnose/tsg027-observe-bdc-create.ipynb'], ['provided port is already allocated', 'TSG062 - Get tail of all previous container logs for pods in BDC namespace', '../log-files/tsg062-tail-bdc-previous-container-logs.ipynb'], ['Create cluster failed since the existing namespace', 'SOP061 - Delete a big data cluster', '../install/sop061-delete-bdc.ipynb'], ['Failed to complete kube config setup', 'TSG067 - Failed to complete kube config setup', '../repair/tsg067-failed-to-complete-kube-config-setup.ipynb'], ['Data source name not found and no default driver specified', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Can\'t open lib \'ODBC Driver 17 for SQL Server', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Control plane upgrade failed. Failed to upgrade controller.', 'TSG108 - View the controller upgrade config map', '../diagnose/tsg108-controller-failed-to-upgrade.ipynb'], ['NameError: name \'azdata_login_secret_name\' is not defined', 'SOP013 - Create secret for azdata login (inside cluster)', '../common/sop013-create-secret-for-azdata-login.ipynb'], ['ERROR: No credentials were supplied, or the credentials were unavailable or inaccessible.', 'TSG124 - \'No credentials were supplied\' error from azdata login', '../repair/tsg124-no-credentials-were-supplied.ipynb'], ['Please accept the license terms to use this product through', 'TSG126 - azdata fails with \'accept the license terms to use this product\'', '../repair/tsg126-accept-license-terms.ipynb'], ], 'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb'], ], 'python': [['Library not loaded: /usr/local/opt/unixodbc', 'SOP012 - Install unixodbc for Mac', '../install/sop012-brew-install-odbc-for-sql-server.ipynb'], ['WARNING: You are using pip version', 'SOP040 - Upgrade pip in ADS Python sandbox', '../install/sop040-upgrade-pip.ipynb'], ], }
install_hint = {'azdata': [ 'SOP063 - Install azdata CLI (using package manager)', '../install/sop063-packman-install-azdata.ipynb' ], 'kubectl': [ 'SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb' ], }
print('Common functions defined successfully.')
```
### Get the Kubernetes namespace for the big data cluster
Get the namespace of the Big Data Cluster use the kubectl command line
interface .
**NOTE:**
If there is more than one Big Data Cluster in the target Kubernetes
cluster, then either:
- set \[0\] to the correct value for the big data cluster.
- set the environment variable AZDATA\_NAMESPACE, before starting
Azure Data Studio.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
if "AZDATA_NAMESPACE" in os.environ:
namespace = os.environ["AZDATA_NAMESPACE"]
else:
try:
namespace = run(f'kubectl get namespace --selector=MSSQL_CLUSTER -o jsonpath={{.items[0].metadata.name}}', return_output=True)
except:
from IPython.display import Markdown
print(f"ERROR: Unable to find a Kubernetes namespace with label 'MSSQL_CLUSTER'. SQL Server Big Data Cluster Kubernetes namespaces contain the label 'MSSQL_CLUSTER'.")
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print(f'The SQL Server Big Data Cluster Kubernetes namespace is: {namespace}')
```
### Establish if cluster is Active Directory enabled
An Active Directory enabled cluster will have a `dns` pod. Non Active
Directory enabled clusters do not have a `dns` pod.
```
dns_pod = run(f'kubectl get pods -n {namespace} -o name -l app=dns', return_output=True)
if len(dns_pod) > 0:
is_ad_enabled_cluster = True
print(f"Cluster {namespace} is an Active Directory enabled cluster")
else:
is_ad_enabled_cluster = False
print(f"Cluster {namespace} is not an Active Directory enabled cluster")
```
### Is notebook being run inside a Kubernetes cluster
When this is notebook is running inside a Kubernetes cluster, such as
when running inside an App-Deploy pod, there is no KUBECONFIG present,
therefore azdata login needs to use the -e (endpoint) approach to login.
```
import os
if "KUBERNETES_SERVICE_PORT" in os.environ and "KUBERNETES_SERVICE_HOST" in os.environ:
inside_kubernetes_cluster = True
print("This notebook is running inside a Kubernetes cluster")
else:
inside_kubernetes_cluster = False
print("This notebook is not running inside a Kubernetes cluster")
```
### Set username/password from environment variables if not already provided
```
if azdata_login_username == "<INSERT USERNAME>":
if is_ad_enabled_cluster:
if "DOMAIN_SERVICE_ACCOUNT_USERNAME" in os.environ and "DOMAIN_SERVICE_ACCOUNT_PASSWORD" in os.environ and "DOMAIN_SERVICE_ACCOUNT_DOMAIN_NAME" in os.environ:
azdata_login_username = os.environ["DOMAIN_SERVICE_ACCOUNT_USERNAME"]
azdata_login_password = os.environ["DOMAIN_SERVICE_ACCOUNT_PASSWORD"]
azdata_login_domain_name = os.environ["DOMAIN_SERVICE_ACCOUNT_DOMAIN_NAME"]
else:
if "AZDATA_USERNAME" in os.environ and "AZDATA_PASSWORD" in os.environ:
azdata_login_username = os.environ["AZDATA_USERNAME"]
azdata_login_password = os.environ["AZDATA_PASSWORD"]
```
### Verify parameter values have been supplied
```
if is_ad_enabled_cluster:
if azdata_login_username == "<INSERT USERNAME>":
raise SystemExit("Please provide the 'azdata_login_username' parameter value (or set the DOMAIN_SERVICE_ACCOUNT_USERNAME environment variable")
if azdata_login_password == "<INSERT PASSWORD>":
raise SystemExit("Please provide the 'azdata_login_password' parameter value (or set the DOMAIN_SERVICE_ACCOUNT_PASSWORD environment variable")
if azdata_login_domain_name == "<INSERT DOMAIN>":
raise SystemExit("Please provide the 'azdata_login_domain_name' parameter value (or set the DOMAIN_SERVICE_ACCOUNT_DOMAIN_NAME environment variable")
else:
if azdata_login_username == "<INSERT USERNAME>":
raise SystemExit("Please provide the 'azdata_login_username' parameter value (or set the AZDATA_USERNAME environment variable")
if azdata_login_password == "<INSERT PASSWORD>":
raise SystemExit("Please provide the 'azdata_login_password' parameter value (or set the AZDATA_PASSWORD environment variable")
```
### Verify `azdata login` does work with these credentials
```
if not inside_kubernetes_cluster and not is_ad_enabled_cluster:
os.environ["AZDATA_USERNAME"] = azdata_login_username
os.environ["AZDATA_PASSWORD"] = azdata_login_password
print(f'Verifying login for user: {azdata_login_username}')
try:
run(f"azdata login --namespace {namespace} --auth basic")
finally:
del os.environ["AZDATA_USERNAME"]
del os.environ["AZDATA_PASSWORD"]
else:
print("SKIPPED: Can't test the credentials if running inside a Kubernetes cluster, because SOP028 will try to find the secret that hasn't been created yet, or if an AD (secure) cluster, because the client will use the current credentials, not the credentials provided above in the Parameters.")
```
### Create Secret
```
# Delete K8s secret if previously created
#
secret = run(f"kubectl get secrets --field-selector metadata.name={azdata_login_secret_name} -n {namespace} --no-headers -o jsonpath={{.items}}", return_output=True)
if secret != "[]":
run(f"kubectl delete secret {azdata_login_secret_name} -n {namespace}")
if is_ad_enabled_cluster:
print(f"Cluster {namespace} is an Active Directory enabled cluster, create username@domain credential")
if len(azdata_login_username) == 0 or \
len(azdata_login_domain_name) == 0 or \
len(azdata_login_password) == 0 or \
azdata_login_username == "<INSERT USERNAME>" or \
azdata_login_domain_name == "<INSERT PASSWORD>" or \
azdata_login_password == "<INSERT DOMAIN>":
raise SystemExit("This is an Active Directory (secure) cluster, please provide a domain account that has required permissions to run app-deploys and place the executed notebook files in HDFS (variables: azdata_login_username/azdata_login_domain_name/azdata_login_password)")
run(f"""kubectl create secret generic {azdata_login_secret_name} -n {namespace} --from-literal=azdata_login_username={azdata_login_username} --from-literal=azdata_login_domain_name={azdata_login_domain_name} --from-literal=azdata_login_password={azdata_login_password}""")
else:
print(f"Cluster {namespace} is not an Active Directory enabled cluster, create a username/password credential")
if len(azdata_login_username) == 0 or \
len(azdata_login_password) == 0 or \
azdata_login_username == "<INSERT USERNAME>" or \
azdata_login_domain_name == "<INSERT PASSWORD>":
raise SystemExit("Please provide a username/password account that has required permissions to run app-deploys and place the executed notebook files in HDFS (variables: azdata_login_username/azdata_login_password)")
run(f"""kubectl create secret generic {azdata_login_secret_name} -n {namespace} --from-literal=azdata_login_username={azdata_login_username} --from-literal=azdata_login_password={azdata_login_password}""")
```
### Create role
```
role = run(f"kubectl get role --field-selector metadata.name={azdata_login_secret_name}-reader --no-headers -o jsonpath={{.items}} --namespace {namespace}", return_output=True)
if role == "[]": # does not exist
run(f"kubectl create role {azdata_login_secret_name}-reader --verb=get --resource=secrets --resource-name={azdata_login_secret_name} --namespace {namespace}")
```
### Create role-binding
```
role_binding = run(f"kubectl get rolebindings --field-selector metadata.name={azdata_login_secret_name}-reader-binding --no-headers -o jsonpath={{.items}} --namespace={namespace}", return_output=True)
if role_binding == "[]": # does not exist
run(f"kubectl create rolebinding {azdata_login_secret_name}-reader-binding --role={azdata_login_secret_name}-reader --user=system:serviceaccount:test:default --namespace={namespace}")
print("Notebook execution is complete.")
```
| github_jupyter |
# Training a Score Estimator (SALLY)
```
import sys
import os
madminer_src_path = "/home/shomiller/madminer"
sys.path.append(madminer_src_path)
from __future__ import absolute_import, division, print_function, unicode_literals
import logging
import numpy as np
import math
import matplotlib
from matplotlib import pyplot as plt
from scipy.optimize import curve_fit
% matplotlib inline
from madminer.sampling import SampleAugmenter
from madminer import sampling
from madminer.ml import ScoreEstimator, Ensemble
import madminer.__version__
print( 'MadMiner version: {}'.format(madminer.__version__) )
# MadMiner output
logging.basicConfig(
format='%(asctime)-5.5s %(name)-20.20s %(levelname)-7.7s %(message)s',
datefmt='%H:%M',
level=logging.INFO
)
# Output of all other modules (e.g. matplotlib)
for key in logging.Logger.manager.loggerDict:
if "madminer" not in key:
logging.getLogger(key).setLevel(logging.WARNING)
```
## Setup
Here we define a function `augment_and_train`, which creates the augmented (unweighted) training and test samples, then runs the score estimator (using the `SALLY` method) to create a model for a given dataset. With the form of all our datafiles from the previous notebooks, we can run this with just two arguments, `channel` (e.g., `wph_mu`) and `observables` (e.g, `full` or `met`)
```
def augment_and_train(channel, observables, nsamples, is_signal_only=False):
n_estimators = 5
print('Creating Training Samples...\n')
if observables == 'ptw' or '2d':
sampler_obs = 'met'
else:
sampler_obs = observables
# Make (unweighted) training and test samples with augmented data
if is_signal_only:
sampler = SampleAugmenter('data/{}/signal/{}_lhedata_{}.h5'.format(sampler_obs, channel, sampler_obs))
else:
sampler = SampleAugmenter('data/{}/{}_lhedata_{}.h5'.format(sampler_obs, channel, sampler_obs))
#create training samples (the same number as the number of estimators we want)
for i in range(n_estimators):
x, theta, t_xz, _ = sampler.sample_train_local(
theta=sampling.benchmark('sm'),
n_samples=int(nsamples/2.),
folder='./samples/{}/samples_{}_{}'.format(observables, channel, observables),
filename='train_score_{}'.format(i),
sample_only_from_closest_benchmark=False,
)
print('Creating Testing Samples...\n')
#create test sample
_ = sampler.sample_test(
theta=sampling.benchmark('sm'),
n_samples=int(nsamples/2.),
folder='./samples/{}/samples_{}_{}'.format(observables, channel, observables),
filename='test',
sample_only_from_closest_benchmark=False,
)
#Choose which features to train on
# if 'met' or 'full', we use all of them (None),
# otherwise we select the correct indices
if observables == 'met' or 'full':
my_features = None
elif observables == 'ptw':
my_features = [18]
elif observables == '2d':
my_features = [18, 39]
#Create a list of ScoreEstimator objects to add to the ensemble
estimators = [ ScoreEstimator(features=my_features, n_hidden=(50,)) for _ in range(n_estimators) ]
ensemble = Ensemble(estimators)
print('Training Ensemble...\n')
# Run the Training
ensemble.train_all(
method='sally',
x=[ 'samples/{}/samples_{}_{}/x_train_score_{}.npy'.format(observables, channel, observables, i) for i in range(n_estimators) ],
t_xz=[ 'samples/{}/samples_{}_{}/t_xz_train_score_{}.npy'.format(observables, channel, observables, i) for i in range(n_estimators) ],
)
#Finally, save our SALLY model to a file we can load later
ensemble.save('models/{}/sally_ensemble_{}_{}'.format(observables, channel, observables))
```
## Run Augmentation and Training (with Backgrounds)
### Full Observables
```
augment_and_train('wph_mu_wbkgs','full',100*50000)
augment_and_train('wph_e_wbkgs','full',100*50000)
augment_and_train('wmh_mu_wbkgs','full',100*50000)
augment_and_train('wmh_e_wbkgs','full',100*50000)
```
### MET Observables
```
augment_and_train('wph_mu_wbkgs','met',100*50000)
augment_and_train('wph_e_wbkgs','met',100*50000)
augment_and_train('wmh_mu_wbkgs','met',100*50000)
augment_and_train('wmh_e_wbkgs','met',100*50000)
```
### $p_{T,W}$ Only
```
augment_and_train('wph_mu_wbkgs','ptw',100*50000)
augment_and_train('wph_e_wbkgs','ptw',100*50000)
augment_and_train('wmh_mu_wbkgs','ptw',100*50000)
augment_and_train('wmh_e_wbkgs','ptw',100*50000)
```
### $p_{T,W}$ and $m_{T,\mathrm{tot}}$ Only
```
augment_and_train('wph_mu_wbkgs','short_2d',100*50000)
augment_and_train('wph_e_wbkgs','short_2d',100*50000)
augment_and_train('wmh_mu_wbkgs','short_2d',100*50000)
augment_and_train('wmh_e_wbkgs','short_2d',100*50000)
```
## Background Free Runs
### Full Observables
```
augment_and_train('wph_mu_smeftsim','full',20*50000,is_signal_only=True)
augment_and_train('wph_e_smeftsim','full',20*50000,is_signal_only=True)
augment_and_train('wmh_mu_smeftsim','full',20*50000,is_signal_only=True)
augment_and_train('wmh_e_smeftsim','full',20*50000,is_signal_only=True)
```
### MET Observables
```
augment_and_train('wph_mu_smeftsim','met',20*50000,is_signal_only=True)
augment_and_train('wph_e_smeftsim','met',20*50000,is_signal_only=True)
augment_and_train('wmh_mu_smeftsim','met',20*50000,is_signal_only=True)
augment_and_train('wmh_e_smeftsim','met',20*50000,is_signal_only=True)
```
### $p_{T,W}$ Only
```
augment_and_train('wph_mu_smeftsim','ptw',20*50000,is_signal_only=True)
augment_and_train('wph_e_smeftsim','ptw',20*50000,is_signal_only=True)
augment_and_train('wmh_mu_smeftsim','ptw',20*50000,is_signal_only=True)
augment_and_train('wmh_e_smeftsim','ptw',20*50000,is_signal_only=True)
```
### $p_{T,W}$ and $m_{T,\mathrm{tot}}$
```
augment_and_train('wph_mu_smeftsim','2d',20*50000,is_signal_only=True)
augment_and_train('wph_e_smeftsim','2d',20*50000,is_signal_only=True)
augment_and_train('wmh_mu_smeftsim','2d',20*50000,is_signal_only=True)
augment_and_train('wmh_e_smeftsim','2d',20*50000,is_signal_only=True)
```
## Runs with Systematics on Signal Only
### Full Observables
```
augment_and_train('wph_mu_wbkgs_sigsystonly','full',100*50000)
augment_and_train('wph_e_wbkgs_sigsystonly','full',100*50000)
augment_and_train('wmh_mu_wbkgs_sigsystonly','full',100*50000)
augment_and_train('wmh_e_wbkgs_sigsystonly','full',100*50000)
```
### MET Observables
```
augment_and_train('wph_mu_wbkgs_sigsystonly','met',100*50000)
augment_and_train('wph_e_wbkgs_sigsystonly','met',100*50000)
augment_and_train('wmh_mu_wbkgs_sigsystonly','met',100*50000)
augment_and_train('wmh_e_wbkgs_sigsystonly','met',100*50000)
```
### $p_{T,W}$ Only
```
augment_and_train('wph_mu_wbkgs_sigsystonly','ptw',100*50000)
augment_and_train('wph_e_wbkgs_sigsystonly','ptw',100*50000)
augment_and_train('wmh_mu_wbkgs_sigsystonly','ptw',100*50000)
augment_and_train('wmh_e_wbkgs_sigsystonly','ptw',100*50000)
```
### $p_{T,W}$ and $m_{T,\mathrm{tot}}$ Only
```
augment_and_train('wph_mu_wbkgs_sigsystonly','2d',100*50000)
augment_and_train('wph_e_wbkgs_sigsystonly','2d',100*50000)
augment_and_train('wmh_mu_wbkgs_sigsystonly','2d',100*50000)
augment_and_train('wmh_e_wbkgs_sigsystonly','2d',100*50000)
```
| github_jupyter |
```
# Copyright 2021 Google LLC
# Use of this source code is governed by an MIT-style
# license that can be found in the LICENSE file or at
# https://opensource.org/licenses/MIT.
# Notebook authors: Kevin P. Murphy (murphyk@gmail.com)
# and Mahmoud Soliman (mjs@aucegypt.edu)
# This notebook reproduces figures for chapter 10 from the book
# "Probabilistic Machine Learning: An Introduction"
# by Kevin Murphy (MIT Press, 2021).
# Book pdf is available from http://probml.ai
```
<a href="https://opensource.org/licenses/MIT" target="_parent"><img src="https://img.shields.io/github/license/probml/pyprobml"/></a>
<a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/book1/figures/chapter10_logistic_regression_figures.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Figure 10.1:<a name='10.1'></a> <a name='iris-logreg-2d'></a>
(a) Visualization of a 2d plane in a 3d space with surface normal $\mathbf w $ going through point $\mathbf x _0=(x_0,y_0,z_0)$. See text for details. (b) Visualization of optimal linear decision boundary induced by logistic regression on a 2-class, 2-feature version of the iris dataset.
Figure(s) generated by [iris_logreg.py](https://github.com/probml/pyprobml/blob/master/scripts/iris_logreg.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./iris_logreg.py")
%run iris_logreg.py
```
## Figure 10.2:<a name='10.2'></a> <a name='sigmoidPlot2d'></a>
Plots of $\sigma (w_1 x_1 + w_2 x_2)$. Here $\mathbf w = (w_1,w_2)$ defines the normal to the decision boundary. Points to the right of this have $\sigma (\mathbf w ^\top \mathbf x )>0.5$, and points to the left have $\sigma (\mathbf w ^\top \mathbf x ) < 0.5$. Adapted from Figure 39.3 of <a href='#MacKay03'>[Mac03]</a> .
Figure(s) generated by [sigmoid_2d_plot.py](https://github.com/probml/pyprobml/blob/master/scripts/sigmoid_2d_plot.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./sigmoid_2d_plot.py")
%run sigmoid_2d_plot.py
```
## Figure 10.3:<a name='10.3'></a> <a name='kernelTrickQuadratic'></a>
Illustration of how we can transform a quadratic decision boundary into a linear one by transforming the features from $\mathbf x =(x_1,x_2)$ to $\boldsymbol \phi (\mathbf x )=(x_1^2,x_2^2)$. Used with kind permission of Jean-Philippe Vert
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
show_image("/pyprobml/book1/figures/images/Figure_10.3.png")
```
## Figure 10.4:<a name='10.4'></a> <a name='logregPoly'></a>
Polynomial feature expansion applied to a two-class, two-dimensional logistic regression problem. (a) Degree $K=1$. (b) Degree $K=2$. (c) Degree $K=4$. (d) Train and test error vs degree.
Figure(s) generated by [logreg_poly_demo.py](https://github.com/probml/pyprobml/blob/master/scripts/logreg_poly_demo.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./logreg_poly_demo.py")
%run logreg_poly_demo.py
```
## Figure 10.5:<a name='10.5'></a> <a name='irisLossSurface'></a>
NLL loss surface for binary logistic regression applied to Iris dataset with 1 feature and 1 bias term. The goal is to minimize the function.
Figure(s) generated by [iris_logreg_loss_surface.py](https://github.com/probml/pyprobml/blob/master/scripts/iris_logreg_loss_surface.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./iris_logreg_loss_surface.py")
%run iris_logreg_loss_surface.py
```
## Figure 10.6:<a name='10.6'></a> <a name='logregPolyRidge'></a>
Weight decay with variance $C$ applied to two-class, two-dimensional logistic regression problem with a degree 4 polynomial. (a) $C=1$. (b) $C=316$. (c) $C=100,000$. (d) Train and test error vs $C$.
Figure(s) generated by [logreg_poly_demo.py](https://github.com/probml/pyprobml/blob/master/scripts/logreg_poly_demo.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./logreg_poly_demo.py")
%run logreg_poly_demo.py
```
## Figure 10.7:<a name='10.7'></a> <a name='logregMultinom3class'></a>
Example of 3-class logistic regression with 2d inputs. (a) Original features. (b) Quadratic features.
Figure(s) generated by [logreg_multiclass_demo.py](https://github.com/probml/pyprobml/blob/master/scripts/logreg_multiclass_demo.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./logreg_multiclass_demo.py")
%run logreg_multiclass_demo.py
```
## Figure 10.8:<a name='10.8'></a> <a name='labelTree'></a>
A simple example of a label hierarchy. Nodes within the same ellipse have a mutual exclusion relationship between them.
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
show_image("/pyprobml/book1/figures/images/Figure_10.8.png")
```
## Figure 10.9:<a name='10.9'></a> <a name='hierSoftmax'></a>
A flat and hierarchical softmax model $p(w|C)$, where $C$ are the input features (context) and $w$ is the output label (word). Adapted from https://www.quora.com/What-is-hierarchical-softmax .
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
show_image("/pyprobml/book1/figures/images/Figure_10.9_A.png")
show_image("/pyprobml/book1/figures/images/Figure_10.9_B.png")
```
## Figure 10.10:<a name='10.10'></a> <a name='logregRobust'></a>
(a) Logistic regression on some data with outliers (denoted by x). Training points have been (vertically) jittered to avoid overlapping too much. Vertical line is the decision boundary, and its posterior credible interval. (b) Same as (a) but using robust model, with a mixture likelihood. Adapted from Figure 4.13 of <a href='#Martin2018'>[Mar18]</a> .
Figure(s) generated by [logreg_iris_bayes_robust_1d_pymc3.py](https://github.com/probml/pyprobml/blob/master/scripts/logreg_iris_bayes_robust_1d_pymc3.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./logreg_iris_bayes_robust_1d_pymc3.py")
%run logreg_iris_bayes_robust_1d_pymc3.py
```
## Figure 10.11:<a name='10.11'></a> <a name='bitemperedLoss'></a>
(a) Illustration of logistic and tempered logistic loss with $t_1=0.8$. (b) Illustration of sigmoid and tempered sigmoid transfer function with $t_2=2.0$. From https://ai.googleblog.com/2019/08/bi-tempered-logistic-loss-for-training.html . Used with kind permission of Ehsan Amid.
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
show_image("/pyprobml/book1/figures/images/Figure_10.11_A.png")
show_image("/pyprobml/book1/figures/images/Figure_10.11_B.png")
```
## Figure 10.12:<a name='10.12'></a> <a name='bitempered'></a>
Illustration of standard and bi-tempered logistic regression on data with label noise. From https://ai.googleblog.com/2019/08/bi-tempered-logistic-loss-for-training.html . Used with kind permission of Ehsan Amid.
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
show_image("/pyprobml/book1/figures/images/Figure_10.12.png")
```
## Figure 10.13:<a name='10.13'></a> <a name='logregLaplaceGirolamiPost'></a>
(a) Illustration of the data. (b) Log-likelihood for a logistic regression model. The line is drawn from the origin in the direction of the MLE (which is at infinity). The numbers correspond to 4 points in parameter space, corresponding to the lines in (a). (c) Unnormalized log posterior (assuming vague spherical prior). (d) Laplace approximation to posterior. Adapted from a figure by Mark Girolami.
Figure(s) generated by [logregLaplaceGirolamiDemo.py](https://github.com/probml/pyprobml/blob/master/scripts/logregLaplaceGirolamiDemo.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./logregLaplaceGirolamiDemo.py")
%run logregLaplaceGirolamiDemo.py
```
## Figure 10.14:<a name='10.14'></a> <a name='logregLaplaceDemoPred'></a>
Posterior predictive distribution for a logistic regression model in 2d. Top left: contours of $p(y=1|\mathbf x , \mathbf w _ map )$. Top right: samples from the posterior predictive distribution. Bottom left: Averaging over these samples. Bottom right: moderated output (probit approximation). Adapted from a figure by Mark Girolami.
Figure(s) generated by [logregLaplaceGirolamiDemo.py](https://github.com/probml/pyprobml/blob/master/scripts/logregLaplaceGirolamiDemo.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./logregLaplaceGirolamiDemo.py")
%run logregLaplaceGirolamiDemo.py
```
## Figure 10.15:<a name='10.15'></a> <a name='ridgeLassoOLS'></a>
(a) Data for logistic regression question. (b) Plot of $ w _k$ vs amount of correlation $c_k$ for three different estimators.
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
show_image("/pyprobml/book1/figures/images/Figure_10.15_A.png")
show_image("/pyprobml/book1/figures/images/Figure_10.15_B.png")
```
## References:
<a name='MacKay03'>[Mac03]</a> D. MacKay "Information Theory, Inference, and Learning Algorithms". (2003).
<a name='Martin2018'>[Mar18]</a> O. Martin "Bayesian analysis with Python". (2018).
| github_jupyter |
# MLOps with Seldon and Jenkins Classic
This repository shows how you can build a Jenkins Classic pipeline to enable Continuous Integration and Continuous Delivery (CI/CD) on your Machine Learning models leveraging Seldon for deployment.
This CI/CD pipeline will allow you to:
- Run unit tests using Jenkins Classic.
- Run end-to-end tests for your model with KIND (Kubernetes in Docker).
- Promote your model as a across multiple (staging / prod) environments.
To showcase these features we will implement add continuous integration and delivery to three different models.
You can find these under the `/models` folder.
As we shall see, each of them will require a [different approach to deployment](#Use-Cases).
## CI/CD Pipeline
The diagram below provides a high level overview of the CI/CD pipeline.
It includes an overview of all the different types of repositories, together with the stakeholders that are the primary contributors of each, as well as the Kubernetes environments in which the applications are deployed.
The key pieces to note on the diagram are:
- There are different types of environments with different restrictions and behaviours, e.g. staging and production.
- It’s possible to have more than one environment for each type (as the type is just what would give it a specific type of config/behaviour).
- The environments are by default in the same cluster (as namespaces), however it’s also possible to configure them across different clusters.
- Each of the green boxes is a single repository, but it can also have a mono-repo approach, whereby each of the white boxes is a folder within a repo.

### Model implementation repository
From a high-level point of view, when a model implementation repository is updated by a Data Scientist or ML Engineer, the Jenkins CI will push changes to the [GitOps repository](#gitops-repository). This enables the following workflow:
1. A Data Scientist or ML Engineer trains a new model.
2. The Data Scientist or ML Engineer pushes the updated configuration to the model implementation repository.
3. The CI tool automatically builds and tests the model implementation.
4. The CI tool automatically pushes the change into the GitOps staging repository.
5. The CI tool automatically opens a PR into the GitOps production repository.
One key point to highlight which may not be obvious by just looking at the diagram is that in this phase of model implementation, the example above showcases how we can leverage a re-usable model server - that is, reusing a pre-built docker image instead of building one every time.
If there are more custom requirements, the user is in full control of the steps performed by the CI Platform Jenkins.
This means that it is also possible to build s2i wrapped components which may require training the image every time.
To gain a better understanding of how the CI/CD pipeline is implemented on each model implementation repository you can check the documented [deep dive](#diving-into-our-cicd-pipeline).
#### Why a new repo for every model?
A new model implementation repo is currently created because it provides us with a way to separate the “Model Deployment” phase and the “Model Training/Experimentation” phase, and allows us to use the repo as the integration between any frameworks that can serve as sources of models (MLFlow, Kubeflow, Spark, etc).
The repo is able to store any metadata, IDs, and configuration files required, and is processed through the CI pipeline every time it is modified.
#### Building a docker image in model implementation repository
Whilst most of the times users of this approach will be leveraging re-usable model servers such as the SKLearn model server, it is also possible to build a docker image every single time (i.e. build a non-reusable model every time a model changes).
This can be be done by adding the relevant steps which would most often include the s2i utility.
This may be desired if there are non-standard linux libraries or non-standard depdencies that need to be re-installed every time.
### GitOps repository
The state of each of our environments (e.g. production or staging) is stored on a GitOps repository.
This repository contains all the different Kubernetes resources that have been deployed to each cluster.
It is linked through [ArgoCD](#ArgoCD) to each of our Kubernetes clusters (or namespaces) so that a change in the repository triggers an update of our environment.
When the deployment configuration of a machine learning model implementation is updated, this will automatically make the changes available through a PR to the respective manager/tech-lead/approver.
This step will enable the end to end machine learning model promotion to be reviewed and approved by the respective individual.
The manager/tech-lead will have to approve the PR before it can be merged.
Once it’s approved, it will be merged into the GitOps repo, which will immediately trigger the update in the production namespace/cluster.
You can see an example of a GitOps repository in the [SeldonIO/seldon-gitops](https://github.com/SeldonIO/seldon-gitops) repository.
#### Re-usable model server repository
If there is a need for a new reusable model server, then it’s possible to do so by creating a repository which would follow a different path.
This would be different to the model implementation repository as it would only be built once in a while, whilst the model server would be built multiple times.
### Set up
As a pre-requisite you need to ensure that have access to a Kubernetes cluster.
In particular, this guide requires the following pre-requisites:
- A Kubernetes cluster running v1.13+.
- Jenkins Classic installed in your cluster. You can find instructions on how to install and configure it on the [Installing Jenkins on your K8s cluster](#Installing-Jenkins-on-your-K8s-cluster) section.
- Seldon Core v0.5.1 installed in your cluster.
### Use cases
This guide goes through three different methods to build and deploy your model.
Each of these can be found under the `./models/` of this repository.
- Using Seldon pre-built re-usable model servers (`./models/news_classifier`).
- Using custom re-usable servers (`./models/images_classifier`).
- Using custom servers with an embedded model.
## Diving into our CI/CD Pipeline
On this section we will dive into the internals of the CI/CD pipeline for our [model implementation repositories](#Model-implementation-repository).
This includes a detailed description of the `Jenkinsfile`, as well as a look into our suggested testing methodology.
Note that this will cover a generic example.
However, as we shall see, specialising this approach into any of our [three main use cases](#Use-cases) will be straightforward.
We leverage [Jenkins Pipelines](https://jenkins.io/doc/book/pipeline/) in order to run our continous integration and delivery automation.
From a high-level point of view, the pipeline configuration will be responsible for:
- Define a **replicable** test and build environment.
- Run the unit and integration tests (if applicable).
- Promote the application into our staging and production environments.
We can see a `Jenkinsfile` below taken from the `./models/news_classifier` example.
This `Jenkinsfile` defines a pipeline which takes into account all of the points mentioned above.
The following sections will dive into each of the sections in a much higher detail.
```
%%writefile ./models/news_classifier/Jenkinsfile
pipeline {
agent {
kubernetes {
defaultContainer 'core-builder'
yamlFile 'models/news_classifier/podTemplate.yaml'
}
}
stages {
stage('Test') {
steps {
sh '''
cd models/news_classifier
make install_dev test
'''
}
}
stage('Test integration') {
steps {
sh '''
cd models/news_classifier
./integration/kind_test_all.sh
'''
}
}
stage('Promote application') {
steps {
withCredentials([[$class: 'UsernamePasswordMultiBinding',
credentialsId: 'github-access',
usernameVariable: 'GIT_USERNAME',
passwordVariable: 'GIT_PASSWORD']]) {
sh '''
cd models/news_classifier
./promote_application.sh
'''
}
}
}
}
}
%%writefile ./models/news_classifier/podTemplate.yaml
apiVersion: v1
kind: Pod
metadata:
name: test-pod
spec:
containers:
- name: core-builder
image: seldonio/core-builder:0.8
resources:
limits:
cpu: 500m
memory: 1500Mi
ephemeral-storage: "15Gi"
requests:
cpu: 200m
memory: 1500Mi
ephemeral-storage: "15Gi"
securityContext:
privileged: true
tty: true
volumeMounts:
- mountPath: /lib/modules
name: modules
readOnly: true
- mountPath: /sys/fs/cgroup
name: cgroup
- mountPath: /var/lib/docker
name: dind-storage
volumes:
- name: modules
hostPath:
path: /lib/modules
- name: cgroup
hostPath:
path: /sys/fs/cgroup
- name: dind-storage
emptyDir: {}
```
### Replicable test and build environment
In order to ensure that our test environments are versioned and replicable, we make use of the [Jenkins Kubernetes plugin](https://github.com/jenkinsci/kubernetes-plugin).
This will allow us to create a Docker image with all the necessary tools for testing and building our models.
Using this image, we will then spin up a separate pod, where all our build instructions will be ran.
We will use the `podTemplate()` object in the Jenkins Pipeline configuration to define the requirements of this pod
Since it leverages Kubernetes underneath, this also ensure that our CI/CD pipelines are easily scalable.
### Integration tests
Now that we have a model that we want to be able to deploy, we want to make sure that we run end-to-end tests on that model to make sure everything works as expected.
For this we will leverage the same framework that the Kubernetes team uses to test Kubernetes itself: [KIND](https://kind.sigs.k8s.io/).
KIND stands for Kubernetes-in-Docker, and is used to isolate a Kubernetes environent for end-to-end tests.
In our case, we will use this isolated environment to test our model.
The steps we'll have to carry out include:
1. Enable Docker within your CI/CD pod.
2. Add an integration test stage.
3. Leverage the `kind_test_all.sh` script that creates a KIND cluster and runs the tests.
#### Add integration stage to Jenkins
We can leverage Jenkins Pipelines to manage the different stages of our CI/CD pipeline.
In particular, to add an integration stage, we can use the `stage()` object:
```groovy
stage('Test integration') {
steps {
sh '''
cd models/news_classifier
./integration/kind_test_all.sh
'''
}
}
```
#### Enable Docker
To test our models, we will need to build their respective containers, for which we will need Docker.
In order to do so, we will first need to mount a few volumes into the CI/CD container.
These basically consist of the core components that docker will need to be able to run.
To mount them we will add these entries into the `podTemplate.yaml` file.
Please also note that we set container to run in `privileged` mode.
```yaml
ApiVersion: v1
...
spec:
containers:
- name: core-builder
...
securityContext:
privileged: true
...
volumeMounts:
- mountPath: /lib/modules
name: modules
readOnly: true
- mountPath: /sys/fs/cgroup
name: cgroup
- mountPath: /var/lib/docker
name: dind-storage
volumes:
- name: modules
hostPath:
path: /lib/modules
- name: cgroup
hostPath:
path: /sys/fs/cgroup
- name: dind-storage
emptyDir: {}
```
#### Run tests in Kind
The `kind_run_all.sh` may seem complicated at first, but it's actually quite simple.
All the script does is set-up a kind cluster with all dependencies, deploy the model and clean everything up.
Let's break down each of the components within the script.
We first start the docker daemon and wait until Docker is running (using `docker ps q` for guidance.
```bash
## FIRST WE START THE DOCKER DAEMON
service docker start
## the service can be started but the docker socket not ready, wait for ready
WAIT_N=0
while true; do
# docker ps -q should only work if the daemon is ready
docker ps -q > /dev/null 2>&1 && break
if [[ ${WAIT_N} -lt 5 ]]; then
WAIT_N=$((WAIT_N+1))
echo "[SETUP] Waiting for Docker to be ready, sleeping for ${WAIT_N} seconds ..."
sleep ${WAIT_N}
else
echo "[SETUP] Reached maximum attempts, not waiting any longer ..."
break
fi
done
```
Once we're running a docker daemon, we can run the command to create our KIND cluster, and install all the components.
This will set up a Kubernetes cluster using the docker daemon (using containers as Nodes), and then install Ambassador + Seldon Core.
```bash
########################################
## AVOID EXIT ON ERROR FOR FOLLOWING CMDS
set +o errexit
## START CLUSTER
make kind_create_cluster
KIND_EXIT_VALUE=$?
## Ensure we reach the kubeconfig path
export KUBECONFIG=$(kind get kubeconfig-path)
## ONLY RUN THE FOLLOWING IF SUCCESS
if [[ ${KIND_EXIT_VALUE} -eq 0 ]]; then
# KIND CLUSTER SETUP
make kind_setup
SETUP_EXIT_VALUE=$?
```
We can now run the tests; for this we run all the dev installations and kick off our tests (which we'll add inside of the integration folder).
```bash
# BUILD S2I BASE IMAGES
make build
S2I_EXIT_VALUE=$?
## INSTALL ALL REQUIRED DEPENDENCIES
make install_integration_dev
INSTALL_EXIT_VALUE=$?
## RUNNING TESTS AND CAPTURING ERROR
make test
TEST_EXIT_VALUE=$?
fi
```
Finally we just clean everything, including the cluster, the containers and the docker daemon.
```bash
## DELETE KIND CLUSTER
make kind_delete_cluster
DELETE_EXIT_VALUE=$?
########################################
## EXIT STOPS COMMANDS FROM HERE ONWARDS
set -o errexit
## CLEANING DOCKER
docker ps -aq | xargs -r docker rm -f || true
service docker stop || true
```
### Promote your application
After running our integration tests, the last step is to promote our model to our staging and production environments.
For that, we will leverage our [GitOps repository](#GitOps-repository) where the state of each environment is stored.
In particular, we will:
- Push a change to the staging GitOps repository, which will update the staging environment instantly.
- Submit a PR to the production GitOps repository, which will wait for a Tech Lead / Manager approval.
This will be handled by the `promote_application.sh` script, which can be seen below.
```
%%writefile ./models/news_classifier/promote_application.sh
##!/bin/bash
## ENSURE WE ARE IN THE DIR OF SCRIPT
cd -P -- "$(dirname -- "$0")"
## SO WE CAN MOVE RELATIVE TO THE ACTUAL BASE DIR
export GITOPS_REPO="seldon-gitops"
export GITOPS_ORG="adriangonz"
export STAGING_FOLDER="staging"
export PROD_FOLDER="production"
## This is the user that is going to be assigned to PRs
export GIT_MANAGER="adriangonz"
export UUID=$(cat /proc/sys/kernel/random/uuid)
git clone https://${GIT_USERNAME}:${GIT_PASSWORD}@github.com/${GITOPS_ORG}/${GITOPS_REPO}
cd ${GITOPS_REPO}
cp -r ../charts/* ${STAGING_FOLDER}/.
ls ${STAGING_FOLDER}
## Check if any modifications identified
git add -N ${STAGING_FOLDER}/
git --no-pager diff --exit-code --name-only origin/master ${STAGING_FOLDER}
STAGING_MODIFIED=$?
if [[ $STAGING_MODIFIED -eq 0 ]]; then
echo "Staging env not modified"
exit 0
fi
## Adding changes to staging repo automatically
git add ${STAGING_FOLDER}/
git commit -m '{"Action":"Deployment created","Message":"","Author":"","Email":""}'
git push https://${GIT_USERNAME}:${GIT_PASSWORD}@github.com/${GITOPS_ORG}/${GITOPS_REPO}
## Add PR to prod
cp -r ../charts/* production/.
## Create branch and push
git checkout -b ${UUID}
git add ${PROD_FOLDER}/
git commit -m '{"Action":"Moving deployment to production repo","Message":"","Author":"","Email":""}'
git push https://${GIT_USERNAME}:${GIT_PASSWORD}@github.com/${GITOPS_ORG}/${GITOPS_REPO} ${UUID}
## Create pull request
export PR_RESULT=$(curl \
-u ${GIT_USERNAME}:${GIT_PASSWORD} \
-v -H "Content-Type: application/json" \
-X POST -d "{\"title\": \"SeldonDeployment Model Promotion Request - UUID: ${UUID}\", \"body\": \"This PR contains the deployment for the Seldon Deploy model and has been allocated for review and approval for relevant manager.\", \"head\": \"${UUID}\", \"base\": \"master\" }" \
https://api.github.com/repos/$GITOPS_ORG/$GITOPS_REPO/pulls)
export ISSUE_NUMBER=$(echo \
$PR_RESULT |
python -c 'import json,sys;obj=json.load(sys.stdin);print(obj["number"])')
## Assign PR to relevant user
curl \
-u ${GIT_USERNAME}:${GIT_PASSWORD} \
-v -H "Content-Type: application/json" \
-X POST -d "{\"assignees\": [\"${GIT_MANAGER}\"] }" \
https://api.github.com/repos/$GITOPS_ORG/$GITOPS_REPO/issues/$ISSUE_NUMBER
```
## Creating a CI/CD pipeline
In order to add a pipeline to Jenkins, you just have to go to the "Manage Jenkins" configuration dashboard, and click on "New Item" to create a new pipeline.

In the first menu, we'll add a name.
For example, we can create a new pipeline with name `news_classifier`.
We will then be able to add the specific details.
Most of these will remain on "default", but we will need to change a couple of them to add a GitHub trigger, Docker access and to point to the right folder within the repository.
Firstly, we will change the following:
* GitHub hook trigger for GITScm polling.
* Tick "This project is parameterised", and then when you see the next dialog:
* Click on the "Add parameter" dropdown, and select "Credential Parameter".
* This will open yet another box, where you want to provide the following details:
* name: `docker-access`
* Credential type "Username and Password"
* Tick: required
* Default value: Click on the "Add" dropdown, and then on "Jenkins provider":
* This has opened another dialog box, where you want to add your docker credentials.
* For this you need to make sure that the current selected option is "Username and Password".
* There you have to enter your Docker username, and for password it's advised to use a Docker API Key.

Lastly, we will need to point to the right `Jenkinsfile`.
Note that since we are working with a monorepository, where multiple model implementations are tracked, we will need to point our pipeline to the `./models/news_classifier` folder.
If we were working with a single model implementation repository, we would only need to point to the global repo.
* Select "Pipeline script from SCM" from dropdown.
* Add the repository as SCM (in this case https://github.com/SeldonIO/sig-mlops-jenkins-classic/)
* Point to the right `Jenkinsfile` under "Script Path". In this case, `models/news_classifier/Jenkinsfile`.
* If needed, add credentials that will allow to access private repos.

### Running pipeline
In order to trigger a new build, we can do it manually by clicking on "Build with Parameters" and then on "Build" or we can just push a new change to our GitHub repo.
This will take us to a view where we can see some details about each of the stages of the latest builds.

## Installing Jenkins on your K8s cluster
If you already have access to a cluster but which doesn't have Jenkins installed, you can do so easily using Helm.
In particular, you will need to run the following:
```
%%bash
helm install \
--name "jenkins" stable/jenkins \
--namespace "jenkins" \
--set "rbac.create=true" \
--set "master.adminUser=admin" \
--set "master.adminPassword=admin" \
--set "master.serviceType=LoadBalancer"
```
This will install Jenkins and all the required services in the cluster.
To get the Load Balancer where it can be accessed you can run:
```
%%bash
kubectl get svc -n jenkins | grep jenkins
```
### Further configuration
If you wish to set up automated pipeline triggers, you will have to install the "GitHub" plugin (there are quite a few github related ones but the one you want is the one called plainly "GitHub", which then will allow for triggering pipelines automatically on commit.
- Install the GitHub Plugin [(for automated webhook triggers)](https://support.cloudbees.com/hc/en-us/articles/115003015691-GitHub-Webhook-Non-Multibranch-Jobs).
- Provide a GitHub token with read access so it can clone relevant repositories.
- Set-up webhooks so that GitHub can send push requests.
Additionally, you will need to configure your Git's `name` and `email` as part of Jenkins settings.

#### Make sure plugins are updated
If you try to run a pipeline and you get an error such as "No Such DSL Method", or any strange Java exception when running a pipeline, the most probably reason is due to current plugins not being up to date.
Updating your plugins can be done by going to "Manage Jenkins" -> "Plugins", and then selecct all the plugins and click "Update and load after restart". This will take you to another screen - there you should tick the checkbox that reads "restart after plugins are downloaded and installed".
Once you update our plugins you should be ready to go.
## ArgoCD
A key point of this approach to MLOps relies on having a GitOps repository which gets synced with our Kubernetes cluster.
To achieve this we leverage [ArgoCD](https://argoproj.github.io/argo-cd/), which will take care of setting up webhooks with your GitOps repository so that on every change it triggers a synchronisation between the resources you've pushed and what's deployed on the cluster.
### Installation
If you don't have it already, you can install ArgoCD following the [official documentation](https://argoproj.github.io/argo-cd/getting_started/#1-install-argo-cd):
```
%%bash
kubectl create namespace argocd
kubectl apply -n argocd -f https://raw.githubusercontent.com/argoproj/argo-cd/stable/manifests/install.yaml
```
Additionally, you will need to install the accompanying CLI tool.
This tool will allow you to easily link your GitOps repository taking care of the entire process.
The instructions to install it will vary between different platforms.
The official documentation shows the [recommended method](https://argoproj.github.io/argo-cd/cli_installation/) on each of the major ones.
### Setting up GitOps repository
To set up the GitOps repository so that it's tracked by ArgoCD we will use the `argocd` CLI tool.
We will assume that the `GITHUB_ORG` and `REPONAME` environment variables have been created and that the repository has already been created and can be found in the `https://github.com/$GITHUB_ORG/$REPONAME` url.
```
%%bash
export GITHUB_ORG=SeldonIO
export REPONAME=seldon-gitops
```
#### Private repositories (optional)
If your repository is private, we will first need to provide the right credentials for ArgoCD to use.
We can do so either using a [user / password login](https://argoproj.github.io/argo-cd/user-guide/private-repositories/#https-username-and-password-credential) or using [SSH keys](https://argoproj.github.io/argo-cd/user-guide/private-repositories/#tls-client-certificates-for-https-repositories).
Note that, for the former, we can also use a [personal access token](https://help.github.com/en/github/authenticating-to-github/creating-a-personal-access-token-for-the-command-line) instead of the password.
As an example, we will add our GitOps repository using a personal access token.
We will assume that the environment variables `GITHUB_USER` and `GITHUB_TOKEN` are set.
```
%%bash
export GITHUB_USER=john.doe
export GITHUB_TOKEN=12341234
argocd repo add https://github.com/$GITHUB_ORG/$REPONAME --username $GITHUB_USER --password $GITHUB_TOKEN
```
#### Create ArgoCD projects
The next step is to create two projects within ArgoCD to manage the staging and production environments respectively.
Each of them will be linked to a folder within our GitOps repository.
```
%%bash
argocd app create seldon-staging \
--repo https://github.com/$GITHUB_ORG/$REPONAME \
--path staging \
--dest-namespace staging
argocd app create seldon-production \
--repo https://github.com/$GITHUB_ORG/$REPONAME \
--path production \
--dest-namespace production
```
Note that we could also sync our `staging` and `production` environment differently.
For example, we could have them on separate repositories or separate branches.
In this case we would also need to update the `promote_application.sh` script so that it knows how it should promote the respective model between environments.
| github_jupyter |
El objetivo de este documento es explorar alternativas ofrecidas por la el modulo scipy.interpolate para interpolar datos 2d correspondientes a curvas de $C_L~vs.~\alpha$, $C_M~vs.~\alpha$ y $C_D~vs.~C_L$ para distintos reynolds. Los datos son estan extraidos del Report NACA 824 Gregory P.D. Siemens en 1994 cuando estaba en la Universidad de Saskatchewan.
```
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import pyplot as plt
from scipy import interpolate
import numpy as np
import profile_characteristics
%matplotlib inline
```
Creamos un objeto perfil usando el modulo profile_characteristics, que requiere como inicialización el nombre de archivo con los datos antes mencionados. En este caso analizamos el perfil NACA 0009
```
name = 'TR824-Digitized/0009.txt'
airfoil = profile_characteristics.Airfoil(name)
aoa_l = [[aoa for aoa,cl in airfoil.AIRFOIL_DATA["Re{}".format(re)]["AoA_Cl"]] for re in [3,6,9]]
cl_l = [[cl for aoa,cl in airfoil.AIRFOIL_DATA["Re{}".format(re)]["AoA_Cl"]] for re in [3,6,9]]
re_l = [[re for i in range(len(aoa))] for aoa,re in zip(aoa_l,[3e6,6e6,9e6])]
aoa_points = aoa_l[0]+aoa_l[1]+aoa_l[2]
cl_points = cl_l[0]+cl_l[1]+cl_l[2]
re_points = re_l[0]+re_l[1]+re_l[2]
grid_aoa, grid_re = np.mgrid[-20:25.5:0.5,1.5e6:10.5e6:0.5e6]
```
Como primer función de interpolación usamos la función bisplrep, que es la base de las otras funciones de interpolación 2d, y que usa como base la libreria FORTRAN fitpack. Usamos el comando plot_surface de pyplot, indicandole a los ejes que son 3d (para esto hace falta tener el paquete Axes3D de mpl_toolkits.mplot3d). La desventaja que tiene esta función es que hace falta usar la función bisplev para evaluarla.
```
bisplrep = interpolate.bisplrep(aoa_points, re_points, cl_points, s=0.5, kx=4, ky=2)
grid_cl = np.array([[interpolate.bisplev(aoa,re, bisplrep) for aoa,re in zip(aoa_line,re_line)] for aoa_line,re_line in zip(grid_aoa, grid_re)])
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(grid_aoa, grid_re,grid_cl)
ax.scatter(aoa_points, re_points, zs=cl_points, zdir='z')
plt.show()
```
Como segunda opción usamos interp2d, que es un rapper de bisplrep, y devuelve una función lista para usar. La desventaja es que es menos configurable, por lo que hay menos parametros con los que jugar para obtener el resultado deseado, y evitar mensajes de advertencia como el siguiente.
```
interp2d = interpolate.interp2d(aoa_points, re_points, cl_points, kind='cubic')
grid_cl = np.array([[float(interp2d(aoa,re)) for aoa,re in zip(aoa_line,re_line)] for aoa_line,re_line in zip(grid_aoa, grid_re)])
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(grid_aoa, grid_re, grid_cl)
ax.scatter(aoa_points, re_points, zs=cl_points, zdir='z')
fig.show()
```
```
rbf = interpolate.Rbf(aoa_points, re_points, cl_points, epsilon=2, function='linear', smooth=1)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(grid_aoa, grid_re, rbf(grid_aoa, grid_re))
ax.scatter(aoa_points, re_points, zs=cl_points, zdir='z')
plt.show()
```
| github_jupyter |
# SETUP AND DEPS
```
! git clone https://github.com/SwapnilDreams100/calling-out-bluff.git
! pip install alibi xhtml2pdf
from google.colab import drive
drive.mount('/content/drive')
! cp ./drive/My\ Drive/glove.6B.300d.txt ./
essay_type = '7'
import keras.layers as klayers
from keras.preprocessing.text import text_to_word_sequence
from keras.preprocessing.sequence import pad_sequences
from keras.layers import Dense, LSTM, Input, Embedding, GlobalAveragePooling1D, Concatenate, Activation, Lambda, BatchNormalization, Convolution1D, Dropout
from keras.preprocessing.text import Tokenizer
import numpy as np
import nltk
nltk.download('punkt')
# from quadratic_weighted_kappa import QWK
from sklearn.metrics import cohen_kappa_score
from keras.models import Model
from keras import backend as K
from keras.engine.topology import Layer, InputSpec
from keras.optimizers import Adam
from keras.callbacks import EarlyStopping
from keras import regularizers
from keras import initializers
from scipy import stats
import matplotlib as mpl
from IPython.display import HTML
from alibi.explainers import IntegratedGradients
import tensorflow.compat.v1 as tf
tf.enable_eager_execution()
import gc
import matplotlib.pyplot as plt
from collections import Counter
# tf.disable_v2_behavior()
```
# MODEL ARCH
```
class Neural_Tensor_layer(Layer):
def __init__(self,output_dim,input_dim=None, **kwargs):
self.output_dim=output_dim
self.input_dim=input_dim
if self.input_dim:
kwargs['input_shape']=(self.input_dim,)
# print("YAYY", input_dim, output_dim)
super(Neural_Tensor_layer,self).__init__(**kwargs)
def call(self,inputs,mask=None):
e1=inputs[0]
e2=inputs[1]
batch_size=K.shape(e1)[0]
k=self.output_dim
feed_forward=K.dot(K.concatenate([e1,e2]),self.V)
bilinear_tensor_products = [ K.sum((e2 * K.dot(e1, self.W[0])) + self.b, axis=1) ]
for i in range(k)[1:]:
btp=K.sum((e2*K.dot(e1,self.W[i]))+self.b,axis=1)
bilinear_tensor_products.append(btp)
result=K.tanh(K.reshape(K.concatenate(bilinear_tensor_products,axis=0),(batch_size,k))+feed_forward)
return result
def build(self,input_shape):
mean=0.0
std=1.0
k=self.output_dim
d=self.input_dim
##truncnorm generate continuous random numbers in given range
W_val=stats.truncnorm.rvs(-2 * std, 2 * std, loc=mean, scale=std, size=(k,d,d))
V_val=stats.truncnorm.rvs(-2 * std, 2 * std, loc=mean, scale=std, size=(2*d,k))
self.W=K.variable(W_val)
self.V=K.variable(V_val)
self.b=K.zeros((self.input_dim,))
self.trainable_weights.append([self.W,self.V,self.b])
def compute_output_shape(self, input_shape):
batch_size=input_shape[0][0]
return(batch_size,self.output_dim)
class Temporal_Mean_Pooling(Layer): # conversion from (samples,timesteps,features) to (samples,features)
def __init__(self, **kwargs):
super(Temporal_Mean_Pooling,self).__init__(**kwargs)
# masked values in x (number_of_samples,time)
self.supports_masking=True
# Specifies number of dimensions to each layer
self.input_spec=InputSpec(ndim=3)
def call(self,x,mask=None):
if mask is None:
mask=K.mean(K.ones_like(x),axis=-1)
mask=K.cast(mask,K.floatx())
#dimension size single vec/number of samples
return K.sum(x,axis=-2)/K.sum(mask,axis=-1,keepdims=True)
def compute_mask(self,input,mask):
return None
def compute_output_shape(self,input_shape):
return (input_shape[0],input_shape[2])
main_path = './calling-out-bluff/Model3(SkipFlow)/'
EMBEDDING_DIM=300
MAX_NB_WORDS=4000
MAX_SEQUENCE_LENGTH=500
VALIDATION_SPLIT=0.20
DELTA=20
texts=[]
labels=[]
originals = []
fp1=open("glove.6B.300d.txt","r", encoding="utf-8")
glove_emb={}
for line in fp1:
temp=line.split(" ")
try:
glove_emb[temp[0]]=np.asarray([float(i) for i in temp[1:]])
except Exception as e:
pass
fp=open(main_path+"data/training_set_rel3.tsv",'r', encoding="ascii", errors="ignore")
fp.readline()
originals = []
for line in fp:
temp=line.split("\t")
if(temp[1]==essay_type): ## why only 4 ?? - evals in prompt specific fashion
originals.append(float(temp[6]))
# print(originals)
fp.close()
# print(originals)
print("range min - ", min(originals) , " ; range max - ", max(originals))
range_min = min(originals)
range_max = max(originals)
fp=open(main_path+"data/training_set_rel3.tsv",'r', encoding="ascii", errors="ignore")
fp.readline()
sentences=[]
for line in fp:
temp=line.split("\t")
if(temp[1]==essay_type): ## why only 4 ?? - evals in prompt specific fashion
texts.append(temp[2])
labels.append((float(temp[6])-range_min)/(range_max-range_min)) ## why ?? - normalize to range [0-1]
line=temp[2].strip()
sentences.append(nltk.tokenize.word_tokenize(line))
fp.close()
# MAIN LABELS
orig_labels = labels
for i in sentences:
temp1=np.zeros((1, EMBEDDING_DIM))
for w in i:
if(w in glove_emb):
temp1+=glove_emb[w]
temp1/=len(i)
tokenizer=Tokenizer(nb_words = MAX_NB_WORDS) #num_words=MAX_NB_WORDS) #limits vocabulory size
tokenizer.fit_on_texts(texts) #encoding the text
sequences=tokenizer.texts_to_sequences(texts) #returns list of sequences
word_index=tokenizer.word_index #dictionary mapping, word and specific token for that word...
print('Found %s unique tokens.' % len(word_index))
data = pad_sequences(sequences, maxlen=MAX_SEQUENCE_LENGTH) #padding to max_length
# CREATE VALIDATION SET
np.random.seed(0)
indices=np.arange(data.shape[0]) #with one argument, start=0, step =1
print(data.shape)
np.random.shuffle(indices)
data=data[indices]
print(data.shape)
labels=np.asarray(labels)
labels=labels[indices]
# np.reshape(labels, ())
print(labels.shape)
validation_size=int(VALIDATION_SPLIT*data.shape[0])
print(validation_size)
x_train=data[:-validation_size] #data-validation data
print(x_train.shape)
# print(x_train)
# print(labels)
y_train=labels[:-validation_size]
# print(y_train.transpose)
print(y_train.shape)
# y_train = np.reshape(y_train, (1427, 1))
# print(y_train_new)
# print(y_train)
x_val=data[-validation_size:]
print(x_val.shape)
y_val=labels[-validation_size:]
embedding_matrix = np.zeros((len(word_index), EMBEDDING_DIM))
for word,i in word_index.items():
if(i>=len(word_index)):
continue
if word in glove_emb:
embedding_matrix[i]=glove_emb[word]
vocab_size=len(word_index)
print(vocab_size)
embedding_layer=Embedding(vocab_size,EMBEDDING_DIM,weights=[embedding_matrix],
input_length=MAX_SEQUENCE_LENGTH,
mask_zero=True,
trainable=False)
# print(embedding_layer.shape)
side_embedding_layer=Embedding(vocab_size,EMBEDDING_DIM,weights=[embedding_matrix],
input_length=MAX_SEQUENCE_LENGTH,
mask_zero=False,
trainable=False)
def SKIPFLOW(lstm_dim=50, lr=1e-4, lr_decay=1e-6, k=4, eta=3, delta=50, activation="relu", maxlen=MAX_SEQUENCE_LENGTH, seed=None):
e = Input(name='essay',shape=(maxlen,))
print("e", e)
# trad_feats=Input(shape=(7,))
# print("trad_feats", trad_feats)
embed = embedding_layer(e)
print(embed.shape)
lstm_layer=LSTM(lstm_dim,return_sequences=True)
# print(lstm_layer)
hidden_states=lstm_layer(embed)
htm=Temporal_Mean_Pooling()(hidden_states)
side_embed = side_embedding_layer(e)
side_hidden_states=lstm_layer(side_embed)
tensor_layer=Neural_Tensor_layer(output_dim=k,input_dim=500)
# print(input_dim, output_dim)
pairs = [((eta + i * delta) % maxlen, (eta + i * delta + delta) % maxlen) for i in range(maxlen // delta)]
hidden_pairs = [ (Lambda(lambda t: t[:, p[0], :])(side_hidden_states), Lambda(lambda t: t[:, p[1], :])(side_hidden_states)) for p in pairs]
sigmoid = Dense(1, activation="sigmoid", kernel_initializer=initializers.glorot_normal(seed=seed))
coherence = [sigmoid(tensor_layer([hp[0], hp[1]])) for hp in hidden_pairs]
co_tm=Concatenate()(coherence[:]+[htm])
dense = Dense(256, activation=activation,kernel_initializer=initializers.glorot_normal(seed=seed))(co_tm)
dense = Dense(128, activation=activation,kernel_initializer=initializers.glorot_normal(seed=seed))(dense)
dense = Dense(64, activation=activation,kernel_initializer=initializers.glorot_normal(seed=seed))(dense)
out = Dense(1, activation="sigmoid")(dense)
model = Model(inputs=[e], outputs=[out])
print("input", [e])
print("outputs", out)
adam = Adam(lr=lr, decay=lr_decay)
model.compile(loss="mean_squared_error", optimizer=adam, metrics=["MSE"])
return model
earlystopping = EarlyStopping(monitor="val_mean_squared_error", patience=5)
sf = SKIPFLOW(lstm_dim=500, lr=2e-4, lr_decay=2e-6, k=4, eta=13, delta=50, activation="relu", seed=None)
# sf.summary()
# sf.load_weights(main_path+'weights_final/'+essay_type+'_weights.h5')
# load weights
import pickle
pklfile= "./drive/My Drive/sf_models/"+essay_type+"_weights.pkl"
fpkl= open(pklfile, 'rb')
sf.set_weights(pickle.load(fpkl))
fpkl.close()
import pandas as pd
import nltk
from nltk import tokenize
import pickle
from random import shuffle
import random
import numpy as np
from scipy import spatial
from xhtml2pdf import pisa
import math
from alibi.explainers import IntegratedGradients as alibi_ig
import matplotlib as mpl
from collections import Counter
from sklearn.metrics import cohen_kappa_score
import matplotlib.pyplot as plt
import os
import copy
import gc
from google_drive_downloader import GoogleDriveDownloader as gdd
gdd.download_file_from_google_drive(file_id='1CIEpiDmzLmJ6LMCVSOmCKw_eOg4ocuS4', dest_path='/content/AES.zip', unzip=True)
# get val predictions
preds_main = sf.predict(x_val)
preds_main =[int(round(a*(range_max-range_min)+range_min)) for a in preds_main.flatten().tolist()]
# MAIN ADVERSARIAL CLASS - WITH IG METHOD AND ADV SAMPLE GENERATION
class gen_adv_examples:
def __init__(self, maxlen, tokenizer, prompt, model, input_name, type_ig, model_name, val_preds):
self.MAX_SEQUENCE_LENGTH = maxlen
self.tokenizer = tokenizer
self.word_map = tokenizer.word_index
self.reversed_word_map = dict(map(reversed, self.word_map.items()))
self.prompt = prompt
self.model = model
self.input_name = input_name
self.type_ig = type_ig
prompt_to_range = {'1':[2,12],'2':[1,6],'3':[0,3],'4':[0,3],'5':[0,4],'6':[0,4],'7':[0,30],'8':[0,60]}
self.range_min = prompt_to_range[self.prompt][0]
self.range_max = prompt_to_range[self.prompt][1]
self.model_name = model_name
self.ATTRS_DIR = '/content/drive/My Drive/IG RESULTS/'+self.model_name+'/P'+self.prompt+'/'
self.ATTRS_TSV = self.ATTRS_DIR + 'attrs.tsv'
self.small_preds_orig = []
self.big_preds_orig = val_preds
def normalize(self, labels):
if type(labels) == 'numpy.ndarray':
labels_new = labels.flatten().tolist()
else:
labels_new = copy.deepcopy(labels)
l = [0]*len(labels_new)
for i, label in enumerate(labels_new):
l[i] = (float(label)-self.range_min)/(self.range_max-self.range_min)
return l
def denormalize(self, labels):
if type(labels) == 'numpy.ndarray':
labels_new = labels.flatten().tolist()
else:
labels_new = copy.deepcopy(labels)
l = [0]*len(labels_new)
for i, label in enumerate(labels_new):
if math.isnan(labels_new[i]):
l[i] = self.range_min
else:
l[i] = int(label*(self.range_max-self.range_min) + self.range_min)
return l
def vectorize(self, text_array, is_text = False, pad='pre'):
texts = copy.deepcopy(text_array)
if is_text:
texts = self.tokenizer.texts_to_sequences(texts)
padded_seq = pad_sequences(texts, maxlen = self.MAX_SEQUENCE_LENGTH, padding = pad, truncating = pad)
return padded_seq
def gen_igs(self, n_steps=50, method="riemann_trapezoid", batch_size=100):
self.ig = alibi_ig(self.model,
layer=self.model.get_layer(self.input_name),
n_steps=n_steps,
method=method,
internal_batch_size=batch_size)
def save_data(self, data, name, is_text=False):
all_texts = []
for i,tokens in enumerate(data):
d={}
if not is_text:
text,_ = self.sequence_to_text(tokens)
else:
text = tokens
d['text'] = ' '.join(text)
d['label_orig'] = self.labels_orig[i]
all_texts.append(d)
df = pd.DataFrame(all_texts)
df.to_csv('/content/drive/My Drive/IG RESULTS/'+name+'_'+self.prompt+'.csv')
def make_glove(self, glove_emb):
self.glove_emb = glove_emb
def find_closest_embeddings(self, embedding):
return sorted(self.glove_emb.keys(), key=lambda word: spatial.distance.euclidean(self.glove_emb[word], embedding))[1]
def top_k_attrs(self, tokens, attrs, k):
k = min(k, len(tokens))
return ([tokens[i].strip() for i in np.argpartition(attrs, -k)[-k:]])
def rindex(self, lst, value):
lst.reverse()
i = lst.index(value)
lst.reverse()
return len(lst) - i - 1
def normal(self, path = '/content/drive/My Drive/IG RESULTS/'):
norm = pd.read_csv(path+self.prompt+'_normal.csv')['text'].tolist()
normal_data = self.vectorize(norm, pad = 'pre', is_text=True)
preds = self.predict_and_norm(np.array(normal_data))
self.small_preds_orig = preds
# self.general_proc(normal_data, self.small_preds_orig, save = True, NAME = 'normal')
def adv_add_song(self, path = '/content/drive/My Drive/IG RESULTS/'):
song_beg = pd.read_csv(path+self.prompt+'_song_beg.csv')['text'].tolist()
song_end = pd.read_csv(path+self.prompt+'_song_end.csv')['text'].tolist()
song_beg_data = self.vectorize(song_beg, pad = 'pre', is_text=True)
song_end_data = self.vectorize(song_end, pad = 'post', is_text=True)
self.general_proc(song_beg_data, self.small_preds_orig, save = True, NAME = 'song_beg')
self.general_proc(song_end_data, self.small_preds_orig, save = True, NAME = 'song_end')
def adv_del(self, path = '/content/drive/My Drive/IG RESULTS/'):
del_beg = pd.read_csv(path+self.prompt+'_del_beg.csv')['text'].tolist()
del_end = pd.read_csv(path+self.prompt+'_del_end.csv')['text'].tolist()
del_beg_data = self.vectorize(del_beg, pad = 'pre', is_text=True)
del_end_data = self.vectorize(del_end, pad = 'pre', is_text=True)
self.general_proc(del_beg_data, self.small_preds_orig, save = True, NAME = 'del_beg')
self.general_proc(del_end_data, self.small_preds_orig, save = True, NAME = 'del_end')
def adv_modify_syn(self, top_k=10, path = '/content/drive/My Drive/IG RESULTS/'):
result = []
norm = pd.read_csv(path+self.prompt+'_normal.csv')['text'].tolist()
data_test = self.vectorize(norm, pad = 'pre', is_text=True)
for i,r in enumerate(data_test):
attrs = self.get_attrs_alibi(np.array([r]))[0]
res_words,counts = self.sequence_to_text(r)
assert (len(res_words[counts:]) == len(attrs[counts:]))
high_attr_tokens = self.top_k_attrs(res_words[counts:],attrs[counts:], top_k)
high_attr_tokens = [x for x in high_attr_tokens if x in self.glove_emb.keys()]
high_attr_tokens = list(set(list(high_attr_tokens)))
print(high_attr_tokens)
syn_dict = {}
for token in high_attr_tokens:
syn = self.find_closest_embeddings(self.glove_emb[token])
# if syn in self.reversed_word_map.keys():
syn_dict[token] = syn
# else:
# continue
print(syn_dict)
for i, token in enumerate(res_words):
if token in high_attr_tokens:
res_words[i] = syn_dict[token]
result.append(res_words[counts:])
# res_data = self.vectorize(result, pad='pre', is_text = True)
self.save_data(result, 'syn', is_text = True)
self.general_proc(res_data, self.small_preds_orig, save = True, NAME = 'syn')
def adv_syn_all(self, percent=0.1, top_k= None):
import tqdm
result = []
syn_dict = {}
with open(self.ATTRS_TSV) as f:
for line in tqdm.tqdm(f):
line = line.strip()
all_attrs = line.split('\t')[0]
tokens = []
attrs = []
for word_attr in all_attrs.split('||'):
if word_attr == 'done':
break
word, attr = word_attr.split('|')
tokens.append(word)
attrs.append(float(attr))
if top_k == None:
top_k = int(percent*len(tokens))
high_attr_tokens = self.top_k_attrs(tokens, attrs, top_k)
high_attr_tokens = list(set(high_attr_tokens))
for token in high_attr_tokens:
if token not in syn_dict.keys():
try:
syn_dict[token] = self.find_closest_embeddings(self.glove_emb[token])
except Exception as e:
pass
c = 0
for i, token in enumerate(tokens):
if token in high_attr_tokens and c<=top_k and token in syn_dict.keys():
tokens[i] = syn_dict[token]
c+=1
result.append(tokens)
res_data = self.vectorize(result, pad='pre', is_text = True)
self.general_proc(res_data, self.big_preds_orig, save = False, NAME = 'syn_all')
def adv_babel(self, path):
babel_csv = pd.read_csv(path, names = ['text'])
result = babel_csv['text'].tolist()[:2]
self.save_data(result, 'babel')
res_data = self.vectorize(result, is_text = True)
self.general_proc(res_data, self.small_preds_orig, save = True, NAME = 'babel')
def general_proc(self, result, preds_orig, save = False, NAME = None):
print(NAME+': PREDICTING')
res_pred = self.predict_and_norm(result)
if save:
print(NAME+': SAVING')
if NAME == 'babel':
res_pred = [self.range_min]*len(result)
self.save_attrs_pdf(result, preds_orig, res_pred, NAME)
else:
print(NAME+': GETTING STATS')
self.get_pred_stats(preds_orig, res_pred, NAME)
print(NAME+': DONE')
def remove_tokens(self, data, counts_list, MAX_LEN = 10):
t_new= []
l = 0
for i,v in enumerate(data):
x_new = []
l_max = 0
for w in v:
if l_max < MAX_LEN:
if w==0:
pass
elif w in counts_list[i]:
l_max += 1
pass
else:
x_new.append(w)
else:
x_new.append(w)
l+=len(x_new)
t_new.append(x_new)
avg_len = l/len(t_new)
t_new = self.vectorize(t_new, is_text = True)
return t_new , avg_len
def get_data_from_tsv(self):
data= []
with open(self.ATTRS_TSV) as f:
for line in f:
line = line.strip()
all_attrs = line.split('\t')[0]
tokens = []
for word_attr in all_attrs.split('||'):
if word_attr == 'done':
break
word, _ = word_attr.split('|')
tokens.append(word)
data.append(tokens)
return np.array(data)
def init_test(self, n = 1, is_abs= False):
top_counts_list,_ = self.get_top_bottom_attrs(n, is_top = True, is_abs= is_abs)
bottom_counts_list,_ = self.get_top_bottom_attrs(n, is_top = False, is_abs= is_abs)
top_counts_list = list(top_counts_list)
bottom_counts_list = list(bottom_counts_list)
data = self.get_data_from_tsv()
assert len(top_counts_list) == len(bottom_counts_list)
new_top,top_len = self.remove_tokens(data, top_counts_list, MAX_LEN =n)
new_bottom,bottom_len = self.remove_tokens(data, bottom_counts_list, MAX_LEN =n)
print(top_len,bottom_len)
top_pred = self.predict_and_norm(new_top)
bottom_pred = self.predict_and_norm(new_bottom)
self.get_pred_stats(self.big_preds_orig, top_pred, 'top')
self.get_pred_stats(self.big_preds_orig, bottom_pred, 'bottom')
def get_attrs_alibi(self, v = x_val[0:1]):
baseline = np.zeros(x_val[0:1].shape)
baseline[0][0] = tokenizer.word_index['a']
explanation = self.ig.explain(v, baselines=baseline)
attrs = explanation.attributions
attrs = attrs.sum(axis=2)
return attrs
def sequence_to_text(self, list_of_indices):
count = 0
words = [self.reversed_word_map.get(letter) for letter in list_of_indices]
for x in words:
if x == None:
count+=1
return(words, count)
def explain(self, essay):
attrs = self.get_attrs_alibi(np.array([essay]))[0]
words,count = self.sequence_to_text(essay)
assert len(words[count:]) == len(attrs[count:])
html = self.visualize_token_attrs(words[count:], attrs[count:])
return attrs, words, count
def visualize_token_attrs(self, tokens, attrs):
cmap='PiYG'
cmap_bound = np.abs(attrs).max()
norm = mpl.colors.Normalize(vmin=-cmap_bound, vmax=cmap_bound)
cmap = mpl.cm.get_cmap(cmap)
html_text = ""
for i, tok in enumerate(tokens):
if tok is not None:
color = mpl.colors.rgb2hex(cmap(norm(attrs[i])))
html_text += " <mark style='background-color:{}'>{}</mark>".format(color, tok)
return (html_text)
def convert_html_to_pdf(self, source_html, output_filename):
result_file = open(output_filename, "w+b")
pisa_status = pisa.CreatePDF(source_html, dest=result_file)
result_file.close()
def save_attrs_pdf(self, data, labels_orig, labels_new, essay_type):
dir = self.ATTRS_DIR+essay_type+'/'
if not os.path.exists(dir):
os.makedirs(dir)
for i,essay in enumerate(data):
attrs = self.get_attrs_alibi(np.array([essay]))[0]
words,count = self.sequence_to_text(essay)
assert len(words[count:]) == len(attrs[count:])
question_attrs = []
html = self.visualize_token_attrs(words[count:], attrs[count:])
self.convert_html_to_pdf(html, dir+str(i)+'_'+str(labels_orig[i])+'_'+str(labels_new[i])+'.pdf')
def save_attrs_tsv(self, data):
self.ATTRS_TSV = self.ATTRS_DIR + 'attrs.tsv'
if os.path.isdir(self.ATTRS_DIR):
pass
else:
os.mkdir(self.ATTRS_DIR)
n = len(data)
batch = 1
with open(self.ATTRS_TSV, 'a') as outf:
c=0
while c<n:
ans = ''
for i,v in enumerate(data[c:c+batch]):
tsv_string = ''
attrs = self.get_attrs_alibi(np.array([v]))[0]
words,count = self.sequence_to_text(v)
assert len(words[count:]) == len(attrs[count:])
question_attrs = []
for ind in range(len(words[count:])):
if words[count:][ind] != None and str(attrs[count:][ind])!=None:
question_attrs.append(
'|'.join([ words[count:][ind], str(attrs[count:][ind]) ])
)
tsv_string = ['||'.join(question_attrs)]
ans += '\t'.join(tsv_string) + '\n'
del attrs, words, question_attrs, tsv_string
gc.collect()
c+=batch
print(c)
outf.write(ans)
outf.flush()
del ans
gc.collect()
print('DONE ALL')
def top_k_attrs(self, tokens, attrs, k):
k = min(k, len(tokens))
return ([tokens[i].strip() for i in np.argpartition(attrs, -k)[-k:]])
def bottom_k_attrs(self, tokens, attrs, k):
k = min(k, len(tokens))
return [tokens[i].strip() for i in np.argpartition(attrs, k)[:k]]
def get_top_bottom_attrs(self, top_k, is_top = True, is_abs=False): #top k attributions from each line
counts_list = []
with open(self.ATTRS_TSV) as f:
for line in f:
line = line.strip()
all_attrs = line.split('\t')[0]
tokens = []
attrs = []
for word_attr in all_attrs.split('||'):
if word_attr == 'done':
break
word, attr = word_attr.split('|')
tokens.append(word)
if is_abs:
attrs.append(abs(float(attr)))
else:
attrs.append(float(attr))
try:
if is_top:
counts_list.append(self.top_k_attrs(tokens, attrs, top_k))
else:
counts_list.append(self.bottom_k_attrs(tokens, attrs, top_k))
except Exception as e:
pass
flat_counts_list = [item for sublist in counts_list for item in sublist]
frequent_attributions = Counter(flat_counts_list)
if is_top:
with open(self.ATTRS_DIR+'highest_attrs.txt', 'w') as f:
attr_to_save = frequent_attributions.most_common(10)
f.write(str(attr_to_save))
# frequent_attributions = Counter(counts_list)
return counts_list, frequent_attributions
def npos(self, orig, new):
count = 0
for i in range(len(orig)):
if new[i]>orig[i]:
count+=1
return count
def nneg(self, orig, new):
count = 0
for i in range(len(orig)):
if new[i]<orig[i]:
count+=1
return count
def nsame(self, orig, new):
count = 0
for i in range(len(orig)):
if new[i]==orig[i]:
count+=1
return count
def get_pred_stats(self, orig, new, name):
# a = ('kappa', cohen_kappa_score(orig, new, weights='quadratic'))
b = ('NPOS', self.npos(orig, new))
c = ('NNEG', self.nneg(orig, new))
d = ('NSAME', self.nsame(orig, new))
with open(self.ATTRS_DIR+ 'results_'+name+'.txt', 'w') as f:
f.write(str([b,c,d]))
def get_overstability_data(self, data, labels, top_k=100, spacing = 10):
curve_data = {}
counts_list,_ = self.get_top_bottom_attrs(top_k, is_top=True)
if type(counts_list[0]) == list:
counts_list = [item for sublist in counts_list for item in sublist]
preds_orig = self.predict_and_norm(np.array(data))
# print(preds_orig, labels)
orig_acc = cohen_kappa_score(preds_orig, labels, weights='quadratic')
for K in np.append(0, np.unique(np.floor(np.geomspace(1, len(Counter(counts_list)), spacing)))):
# take K most top attributed words
if K in curve_data:
continue
whitelist = set([self.word_map[w] for w in counts_list[:int(K)]]) if K > 0 else set()
print('wh len', len(whitelist))
num_batches = 0
avg_question_length_orig = 0
avg_question_length_new = 0
num_questions = 0
pred_array = []
batch_size = 10
# iterator over the validation dataset
for ind in range(0, len(data), batch_size):
test = np.array(data[ind:ind+batch_size])
new_test = np.zeros(test.shape)
curr_batch_size = int(test.shape[0])
for batch_i in range(curr_batch_size):
len_counter = 0
# avg_question_length_orig += int(test[batch_i])
for word_i, w in enumerate(test[batch_i]):
if int(w) in whitelist:
new_test[batch_i, len_counter] = int(w)
len_counter += 1
if len_counter == 0:
len_counter = 1
avg_question_length_new += int(len_counter)
num_questions += 1
input_df = [new_test]
pred = self.predict_and_norm(input_df)
pred_array.extend(pred)
acc = cohen_kappa_score(pred_array, labels, weights='quadratic')
print(acc, orig_acc)
curve_data[len(whitelist)] = acc
self.plot_overstability(curve_data, orig_acc)
def plot_overstability(self, curve_data, orig_acc):
OVERSTABILITY_CURVE_FILE = self.ATTRS_DIR + 'over_'+self.prompt+'.eps'
plt.plot(list(curve_data.keys()), list(curve_data.values())/orig_acc)
plt.xscale('symlog')
plt.xlabel('num. words in vocab')
plt.ylabel('relative accuracy')
plt.savefig(OVERSTABILITY_CURVE_FILE, format='eps')
plt.savefig(OVERSTABILITY_CURVE_FILE.replace('eps','png'), format='png')
def predict_and_norm(self, data):
preds = self.model.predict(data)
new_labels = self.denormalize(preds)
return new_labels
#INIT CLASS
adv = gen_adv_examples(MAX_SEQUENCE_LENGTH, tokenizer, essay_type, sf, 'embedding','alibi', 'SKIPFLOW', preds_main)
# CREATE IG CLASS
adv.gen_igs()
# data, labels = adv.choose_examples(x_val, y_val)
# labels_main = adv.denormalize(y_val)
# adv.make_glove(path = '')
```
# generate ADVERSARIAL EXAMPLES
```
pattern_none = [0, 0, 0, 0, 0]
def subfinder(l, sl):
sll=len(sl)
for ind in (i for i,e in enumerate(l) if e==sl[0]):
if l[ind:ind+sll]==sl:
return ind,ind+sll-1
def get_loc(w):
count = 0
for x in w:
if x == 0:
count+=1
return count
a = []
ids = []
count = 0
for i,essay in enumerate(data):
c = get_loc(essay)
c2 = len(essay[c:])
if c2>200 and c2<250 and count<10:
a.append(c2)
ids.append(i)
count+=1
lab = adv.denormalize(labels)
min_val = min(lab)
max_val = max(lab)
avg_val = (min_val+max_val)//2
min_idxs = [i for i, value in enumerate(lab) if value == min_val][:3]
max_idxs = [i for i, value in enumerate(lab) if value == max_val][:3]
avg_idxs = [i for i, value in enumerate(lab) if value == avg_val][:3]
indices = min_idxs + max_idxs + avg_idxs
print(len(indices))
labels_orig = [min_val]*len(min_idxs) + [max_val]*len(max_idxs) + [avg_val]*len(avg_idxs)
data = [texts[i] for i in indices]
def denormalize(labels, max_val, min_val):
if type(labels) == 'numpy.ndarray':
labels_new = labels.flatten().tolist()
else:
labels_new = copy.deepcopy(labels)
l = [0]*len(labels_new)
for i, label in enumerate(labels_new):
if math.isnan(labels_new[i]):
l[i] = max_val+1
else:
l[i] = int(label*(max_val-min_val) + min_val)
return l
lab = denormalize(orig_labels, 30 , 0)
word_map = tokenizer.word_index
reversed_word_map = dict(map(reversed, word_map.items()))
ig = alibi_ig(sf,
layer=sf.get_layer('embedding'),
n_steps=50,
method='riemann_trapezoid',
internal_batch_size=100)
def vectorize(text_array, is_text = False, pad='pre'):
texts = copy.deepcopy(text_array)
if is_text:
texts = tokenizer.texts_to_sequences(texts)
padded_seq = pad_sequences(texts, maxlen = MAX_SEQUENCE_LENGTH, padding = pad, truncating = pad)
return padded_seq
# save the data in csv
def save_data(data, name, labels_orig, is_text=False):
all_texts = []
for i,tokens in enumerate(data):
d={}
if not is_text:
text,_ = sequence_to_text(tokens)
else:
text = tokens
d['text'] = ' '.join(text)
d['label_orig'] = labels_orig[i]
all_texts.append(d)
df = pd.DataFrame(all_texts)
df.to_csv('/content/drive/My Drive/IG RESULTS/'+name+'_'+self.prompt+'.csv')
def get_attrs_alibi(v = x_val[0:1]):
baseline = np.zeros(x_val[0:1].shape)
baseline[0][0] = tokenizer.word_index['a']
explanation = ig.explain(v, baselines=baseline)
attrs = explanation.attributions
attrs = attrs.sum(axis=2)
return attrs
def sequence_to_text(list_of_indices):
count = 0
words = [reversed_word_map.get(letter) for letter in list_of_indices]
for x in words:
if x == None:
count+=1
return(words, count)
def find_closest_embeddings(embedding):
return sorted(glove_emb.keys(), key=lambda word: spatial.distance.euclidean(glove_emb[word], embedding))[1]
def top_k_attrs(tokens, attrs, k):
k = min(k, len(tokens))
return ([tokens[i].strip() for i in np.argpartition(attrs, -k)[-k:]])
def bottom_k_attrs(tokens, attrs, k):
k = min(k, len(tokens))
return [tokens[i].strip() for i in np.argpartition(attrs, k)[:k]]
def adv_modify_syn(percent=0.10, path = '/content/drive/My Drive/IG RESULTS/', prompt= '7', small= True):
result = []
if small:
norm = pd.read_csv(path+prompt+'_normal.csv')['text'].tolist()
else:
norm = pd.read_csv(path+'big_'+prompt+'_normal.csv')['text'].tolist()
data_test = vectorize(norm, pad = 'pre', is_text=True)
for i,r in enumerate(data_test):
attrs = get_attrs_alibi(np.array([r]))[0]
res_words,counts = sequence_to_text(r)
top_k = int(percent*len(res_words[counts:]))
high_attr_tokens = top_k_attrs(res_words[counts:],attrs[counts:], top_k)
high_attr_tokens = [x for x in high_attr_tokens if x in glove_emb.keys()]
high_attr_tokens = list(set(list(high_attr_tokens)))
attrs_abs = [abs(x) for x in attrs[counts:]]
low_attr_tokens = bottom_k_attrs(res_words[counts:],attrs_abs, top_k)
low_attr_tokens = [x for x in low_attr_tokens if x in glove_emb.keys()]
low_attr_tokens = list(set(list(low_attr_tokens)))
syn_dict = {}
for token in high_attr_tokens:
syn = find_closest_embeddings(glove_emb[token])
syn_dict[token] = syn
for token in low_attr_tokens:
syn = find_closest_embeddings(glove_emb[token])
syn_dict[token] = syn
c_high = 0
c_low = 0
for i, token in enumerate(res_words):
if token in high_attr_tokens and c_high<=top_k:
res_words[i] = syn_dict[token]
c_high += 1
elif token in low_attr_tokens and c_low<=top_k:
res_words[i] = syn_dict[token]
c_low +=1
text = (res_words[counts:])
result.append(' '.join(text))
print(syn_dict)
return result
import random
random.seed(42)
# save the SMALL - 8 sample data
def save(data, out, name):
all_texts = []
for i,text in enumerate(data):
d={}
d['text'] = text
d['label_orig'] = out[i]
all_texts.append(d)
df = pd.DataFrame(all_texts)
df.to_csv('/content/drive/My Drive/IG RESULTS/7_'+name+'.csv')
def create_small_data(texts, lab):
min_val = min(lab)
max_val = max(lab)
avg_val = (min_val+max_val)//2
min_idxs = [i for i, value in enumerate(lab) if value == min_val][:3]
max_idxs = [i for i, value in enumerate(lab) if value == max_val][:3]
avg_idxs = [i for i, value in enumerate(lab) if value == avg_val][:3]
ids = min_idxs + max_idxs + avg_idxs
data = [texts[i] for i in ids]
labs_new = [lab[i] for i in ids]
path = 'song.pickle'
with open('/content/calling-out-bluff/'+path, 'rb') as handle:
songs= pickle.load(handle)
song_tot = songs[:4]+songs[10:16]
false_tot = [ 'There are more submarines in lakes right now than there are in the oceans. ', \
'Apples are not fruits. ',\
'The world is flat. ']
song_beg = []
song_end = []
for i in range(len(data)):
random_song = random.choice(song_tot)
song_beg.append(random_song + data[i])
song_end.append(data[i] + random_song)
false_beg = []
false_end = []
for i in range(len(data)):
random_false = random.choice(false_tot)
false_beg.append(random_false +' '+ data[i])
false_end.append(data[i] + ' ' + random_false)
def scramble(sentence):
split = tokenize.sent_tokenize(sentence)
shuffle(split) # This shuffles the list in-place.
return ''.join(split) # Turn the list back into a string
shuffled=[]
for i in range(len(data)):
shuffled.append( scramble( data[i] ) )
out = labs_new
save(data, out, 'normal')
save(song_beg, out, 'song_beg')
save(song_end, out, 'song_end')
save(false_beg, out, 'false_beg')
save(false_end, out, 'false_end')
save(shuffled, out, 'shuffle')
return out
out = create_small_data(texts, lab)
syn_result = adv_modify_syn(prompt = '7')
save(syn_result, out, 'syn')
# save the BIG data of 100 samples
def save_big(data, out, name):
all_texts = []
for i,text in enumerate(data):
d={}
d['text'] = text
d['label_orig'] = out[i]
all_texts.append(d)
df = pd.DataFrame(all_texts)
df.to_csv('/content/drive/My Drive/IG RESULTS/big_7_'+name+'.csv')
def create_big_data(texts, lab, n = 100):
data = texts[:n]
labs_new = lab[:n]
path = 'song.pickle'
with open('/content/calling-out-bluff/'+path, 'rb') as handle:
songs= pickle.load(handle)
song_tot = songs[:4]+songs[10:16]
false_tot = [ 'There are more submarines in lakes right now than there are in the oceans. ', \
'Apples are not fruits. ',\
'The world is flat. ']
song_beg = []
song_end = []
for i in range(len(data)):
random_song = random.choice(song_tot)
song_beg.append(random_song + data[i])
song_end.append(data[i] + random_song)
false_beg = []
false_end = []
for i in range(len(data)):
random_false = random.choice(false_tot)
false_beg.append(random_false +' '+ data[i])
false_end.append(data[i] + ' ' + random_false)
def scramble(sentence):
split = tokenize.sent_tokenize(sentence)
shuffle(split) # This shuffles the list in-place.
return ''.join(split) # Turn the list back into a string
shuffled=[]
for i in range(len(data)):
shuffled.append( scramble( data[i] ) )
out = labs_new
save_big(data, out, 'normal')
save_big(song_beg, out, 'song_beg')
save_big(song_end, out, 'song_end')
save_big(false_beg, out, 'false_beg')
save_big(false_end, out, 'false_end')
save_big(shuffled, out, 'shuffle')
return out
out = create_big_data(texts, lab)
syn_result = adv_modify_syn(small = False, prompt = '7')
save_big(syn_result, out, 'syn')
```
# SAVE ATTRIBUTION
```
import gc
import multiprocessing
word_map = tokenizer.word_index
reversed_word_map = dict(map(reversed, word_map.items()))
ig = alibi_ig(sf,
layer=sf.get_layer('embedding'),
n_steps=50,
method='riemann_trapezoid',
internal_batch_size=100)
def vectorize(text_array, is_text = False, pad='pre'):
texts = copy.deepcopy(text_array)
if is_text:
texts = tokenizer.texts_to_sequences(texts)
padded_seq = pad_sequences(texts, maxlen = MAX_SEQUENCE_LENGTH, padding = pad, truncating = pad)
return padded_seq
def sequence_to_text(list_of_indices):
count = 0
words = [reversed_word_map.get(letter) for letter in list_of_indices]
for x in words:
if x == None:
count+=1
return(words, count)
def get_attrs_alibi(v = x_val[0:1]):
baseline = np.zeros(x_val[0:1].shape)
baseline[0][0] = tokenizer.word_index['a']
explanation = ig.explain(v, baselines=baseline)
attrs = explanation.attributions
attrs = attrs.sum(axis=2)
return attrs
def create_model_and_train( attr_type = 'add'):
ATTRS_DIR = './drive/My Drive/IG RESULTS/'
ATTRS_TSV = ATTRS_DIR+'SKIPFLOW/P7/attrs_'+attr_type+'.tsv'
data = pd.read_csv(ATTRS_DIR+'big_7_'+attr_type+'.csv')['text'].tolist()
data_test = vectorize(data, pad = 'pre', is_text=True)
with open(ATTRS_TSV, 'w') as outf:
ans = ''
for i,v in enumerate(data_test):
tsv_string = ''
attrs = get_attrs_alibi(np.array([v]))[0]
words,count = sequence_to_text(v)
assert len(words[count:]) == len(attrs[count:])
question_attrs = []
for ind in range(len(words[count:])):
if words[count:][ind] is not None:
question_attrs.append(
'|'.join([ words[count:][ind], str(attrs[count:][ind]) ])
)
tsv_string = ['||'.join(question_attrs)]
ans += '\t'.join(tsv_string) + '\n'
del attrs, words, question_attrs, tsv_string
gc.collect()
outf.write(ans)
outf.flush()
del ans
gc.collect()
outf.write('done')
print('DONE')
attr_type = 'song_beg'
create_model_and_train(attr_type)
gc.collect()
attr_type = 'song_end'
create_model_and_train(attr_type)
gc.collect()
attr_type = 'false_beg'
create_model_and_train(attr_type)
gc.collect()
attr_type = 'false_end'
create_model_and_train(attr_type)
gc.collect()
attr_type = 'normal'
create_model_and_train(attr_type)
# attr_type = 'shuffle'
# create_model_and_train(attr_type)
gc.collect()
attr_type = 'syn'
create_model_and_train(attr_type)
gc.collect()
```
# LOAD ADVERSARIAL EXAMPLES AND ATTRIBUTIONS
```
ATTRS_DIR = '/content/drive/My Drive/IG RESULTS/SKIPFLOW/P7/'
ATTRS_TSV = '/content/drive/My Drive/IG RESULTS/SKIPFLOW/P7/attrs.tsv'
import pandas as pd
essay_set_id = '7'
names = ['song_beg', 'song_end', 'false_beg','false_end','normal','shuffle', 'syn','incomp_data']
adv_data_list = {}
for name in names:
adv_data = pd.read_csv('/content/drive/My Drive/IG RESULTS/'+str(essay_set_id)+'_'+name+'.csv')
adv_data_list[name] = adv_data
E_list = {}
labels_true = []
labels_syn = []
for adv_data in adv_data_list.keys():
texts = adv_data_list[adv_data]['text'].tolist()
if adv_data == 'normal' or adv_data == 'incomp_data':
labels_true = adv_data_list[adv_data]['label_orig'].tolist()
elif adv_data == 'syn':
labels_syn = adv_data_list[adv_data]['label_orig'].tolist()
sequences=tokenizer.texts_to_sequences(texts) #returns list of sequences
data = pad_sequences(sequences, maxlen=MAX_SEQUENCE_LENGTH) #padding to max_length
E_list[adv_data]= data
e = E_list['normal']
labels_orig = adv.predict_and_norm(e)
labels_orig
```
### PROCESS ADV SAMPLES AND SAVE POPULATION STATISTICS
```
from xhtml2pdf import pisa
def convert_html_to_pdf(source_html, output_filename):
result_file = open(output_filename, "w+b")
pisa_status = pisa.CreatePDF(source_html, dest=result_file)
result_file.close()
def save_stats_add(diff, diff_attr, word_list, ratio,output_filename):
result = open(output_filename, 'w')
result.write('\ndiff in scores: '+str(diff))
result.write('\ndiff in attrs: '+ str(diff_attr))
result.write('\nnew words in top 10%: '+ ', '.join(word_list))
result.write('\npercent of top words in added words: '+ str(ratio))
result.close()
def save_stats_mod(diff, diff_attr, changed_no,output_filename):
result = open(output_filename, 'w')
result.write('\ndiff in scores: '+str(diff))
result.write('\ndiff in attrs: '+ str(diff_attr))
result.write('\nno of words which changed attr: '+ str(changed_no))
result.close()
def save_stats_gen(babel_total, babel_unattrib, output_filename):
result = open(output_filename, 'w')
result.write('\ntop attributed words: '+ str(babel_total))
result.write('\ntop unattributed words: '+ str(babel_unattrib))
result.close()
def top_k_attrs(tokens, attrs, sign = None, k=None):
k = min(k, len(tokens))
if sign != None:
tokens_list = []
signs_list = []
for i in np.argpartition(attrs, -k)[-k:]:
tokens_list.append(tokens[i].strip())
signs_list.append(sign[i])
return tokens_list , signs_list
else:
return ([tokens[i].strip() for i in np.argpartition(attrs, -k)[-k:]])
def bottom_k_attrs(tokens, attrs, k):
k = min(k, len(tokens))
return [tokens[i].strip() for i in np.argpartition(attrs, k)[:k]]
def save_normal_attrs(data, labels_orig, essay_type):
dir = ATTRS_DIR+essay_type+'/'
if not os.path.exists(dir):
os.makedirs(dir)
a_total = []
w_total= []
for i,essay in enumerate(data):
attrs, words, c= adv.explain(data[i])
label_new = adv.predict_and_norm(data[i:i+1])[0]
html = adv.visualize_token_attrs(words[c:], attrs[c:])
convert_html_to_pdf(html, dir+str(i)+'_'+str(labels_orig[i])+'_'+str(label_new+0)+'.pdf')
a_total.append(attrs)
w_total.append(words)
return a_total, w_total
def get_loc(w):
count = 0
for x in w:
if x == None:
count+=1
return count
def save_attrs_pdf(data, data_normal, labels_orig, essay_type, type_add = False, type_mod = False, type_gen = False):
dir = ATTRS_DIR+essay_type+'/'
if not os.path.exists(dir):
os.makedirs(dir)
a,w = data_normal
babel_total = {}
babel_unattrib = {}
for i,essay in enumerate(data):
attrs, words,c_data = adv.explain(data[i])
labels_new = adv.predict_and_norm(data[i:i+1])[0]
if not type_gen:
c_normal = get_loc(w[i])
a_normal = a[i][c_normal:]
w_normal = w[i][c_normal:]
a_data = attrs[c_data:]
w_data = words[c_data:]
if type_add:
diff = labels_orig[i] - labels_new
diff_attr = sum(attrs) - sum(a[i])
if essay_type == 'song_beg' or essay_type =='false_beg' :
pattern = w_normal[:4]
try:
loc = subfinder(w_data, pattern)[0]
except Exception as e:
loc = len(w_data)
new_w = w_data[:loc]
new_a = a_data[:loc]
elif essay_type == 'song_end' or essay_type =='false_end':
loc = len(w_normal[i])
new_w = w_data[loc:]
new_a = a_data[loc:]
top_words_normal = top_k_attrs(w_normal, a_normal, k = int(0.1*len(w_normal)))
top_words_data = top_k_attrs(w_data, a_data, k = int(0.1*len(w_data)))
top_words_final = [x for x in top_words_data if x not in top_words_normal]
html = adv.visualize_token_attrs(w_data, a_data)
convert_html_to_pdf(html, dir+str(i)+'_'+str(labels_orig[i])+'_'+str(labels_new+0)+'.pdf')
save_stats_add(diff, diff_attr, top_words_final, len(top_words_final)/len(new_w), dir+'stats_'+str(i)+'_'+str(labels_orig[i])+'_'+str(labels_new+0)+'.txt')
elif type_gen:
attrs_sign= []
for x in a_data:
if x>0:
attrs_sign.append('+')
else:
attrs_sign.append('-')
top_words, top_words_signs = top_k_attrs(w_data, a_data, attrs_sign, k = int(0.1*len(w_data)))
for j,x in enumerate(top_words):
if x in babel_total.keys():
if top_words_signs[j] == '+':
babel_total[x][0]+=1
else:
babel_total[x][1]+=1
else:
if top_words_signs[j] == '+':
babel_total[x] = [1,0]
else:
babel_total[x] = [0,1]
attrs_abs = [abs(x) for x in a_data]
bottom_words = bottom_k_attrs(w_data, attrs_abs, k = int(0.1*len(w_data)))
for j,x in enumerate(bottom_words):
if x in babel_unattrib.keys():
babel_unattrib[x]+= 1
else:
babel_unattrib[x] = 1
html = adv.visualize_token_attrs(w_data, a_data)
convert_html_to_pdf(html, dir+str(i)+'_'+str(labels_new+0)+'.pdf')
elif type_mod:
diff = labels_orig[i] - labels_new
diff_attr = sum(attrs) - sum(a[i])
if essay_type == 'shuffle' or essay_type == 'syn':
attrs_abs=[]
attrs_sign =[]
for x in a_normal:
attrs_abs.append(abs(x))
if x>0:
attrs_sign.append('+')
else:
attrs_sign.append('-')
top_words_orig, token_signs_orig = top_k_attrs(w_normal, attrs_abs, attrs_sign, k = int(0.1*len(w_normal)))
attrs_abs=[]
attrs_sign =[]
for x in a_data:
attrs_abs.append(abs(x))
if x>0:
attrs_sign.append('+')
else:
attrs_sign.append('-')
top_words, token_signs = top_k_attrs(w_data, attrs_abs, attrs_sign, k = int(0.1*len(w_data)))
changed_count = 0
for orig_index,t in enumerate(top_words_orig):
try:
ind = top_words.index(t)
except Exception as e:
ind = -1
if ind!=-1:
if token_signs_orig[orig_index] != token_signs[ind]:
changed_count+=1
html = adv.visualize_token_attrs(w_data, a_data)
convert_html_to_pdf(html, dir+str(i)+'_'+str(labels_orig[i])+'_'+str(labels_new+0)+'.pdf')
# save_stats_mod(diff, diff_attr, changed_count, dir+'stats_'+str(i)+'_'+str(labels_orig[i])+'_'+str(labels_new+0)+'.txt')
if type_gen:
# del babel_unattrib[None]
save_stats_gen(babel_total, babel_unattrib, dir+'stats_word_attributions.txt')
# NORMAL DATA
e_normal = E_list['normal']
data_normal = save_normal_attrs(e_normal, labels_orig, 'normal')
# WORD SOUP SAVE
e_normal = E_list['incomp_data']
data_normal = save_normal_attrs(e_normal, labels_true, 'incomp_data')
# ADDITION DATA
gc.collect()
e_add_song = E_list['song_beg']
save_attrs_pdf(e_add_song, data_normal, labels_orig, 'song_beg', type_add=True)
gc.collect()
e_add_song = E_list['song_end']
save_attrs_pdf(e_add_song, data_normal, labels_orig, 'song_end', type_add=True)
gc.collect()
e_add_song = E_list['false_beg']
save_attrs_pdf(e_add_song, data_normal, labels_orig, 'false_beg', type_add=True)
gc.collect()
e_add_song = E_list['false_end']
save_attrs_pdf(e_add_song, data_normal, labels_orig, 'false_end', type_add=True)
gc.collect()
# SYNONYMS
gc.collect()
e_add_song = E_list['syn']
save_attrs_pdf(e_add_song, data_normal, labels_orig, 'syn', type_mod = True)
gc.collect()
e_add_song = E_list['shuffle']
save_attrs_pdf(e_add_song, data_normal, labels_orig, 'shuffle', type_mod = True)
gc.collect()
e = E_list['incomp_data']
labels_incomp = adv.predict_and_norm(e)
labels_incomp
```
### BABEL DATA
```
from google_drive_downloader import GoogleDriveDownloader as gdd
gdd.download_file_from_google_drive(file_id='1CIEpiDmzLmJ6LMCVSOmCKw_eOg4ocuS4', dest_path='/content/AES.zip', unzip=True)
gc.collect()
babel_data = pd.read_csv('/content/AES_testcases/prompt'+str(essay_set_id)+'/prompt 7 babel - Sheet1.csv', names= ['text'])
babel_data['label_orig'] = 0
texts = babel_data['text'].tolist()
sequences=tokenizer.texts_to_sequences(texts) #returns list of sequences
data = pad_sequences(sequences, maxlen=MAX_SEQUENCE_LENGTH) #padding to max_length
E_babel= data
e_add_song = E_babel
save_attrs_pdf(e_add_song, (1,2), babel_data['label_orig'].tolist(), 'babel', type_gen = True)
## ATTRIBUTIONS PROCESSING
def get_w_a(essay_type):
ATTRS_DIR = '/content/drive/MyDrive/IG RESULTS/SKIPFLOW/P7/'
### NORMAL
path = ATTRS_DIR + 'attrs_'+essay_type+'.tsv'
w = []
a = []
with open(path, 'r') as f:
for line in f:
line = line.strip()
question_attrs = line.split('\t')[0]
question_tokens = []
attrs = []
for word_attr in question_attrs.split('||'):
if word_attr != 'done':
word, attr = word_attr.split('|')
question_tokens.append(word)
attrs.append(float(attr))
question_tokens = question_tokens
attrs = attrs + [0]*(500 - len(attrs))
w.append(question_tokens)
a.append(attrs)
return a[:-1], w[:-1]
def save_stats_add(diff, diff_attr, percent, output_filename):
result = open(output_filename, 'w')
result.write('\ndiff in scores: '+str(diff))
result.write('\ndiff in attrs: '+ str(diff_attr))
result.write('\npercent of top words in added words: '+ str(percent))
result.close()
def save_stats_mod_shuffle(diff, diff_attr, changed_no,output_filename):
result = open(output_filename, 'w')
result.write('\ndiff in scores: '+str(diff))
result.write('\ndiff in attrs: '+ str(diff_attr))
result.write('\nno of top words which changed attr: '+ str(changed_no))
result.close()
def save_stats_mod_syn(diff, diff_attr, changed_top_no,changed_bottom_no, top_words, bottom_words, output_filename):
result = open(output_filename, 'w')
result.write('\n diff in scores: '+str(diff))
result.write('\n diff in attrs: '+ str(diff_attr))
result.write('\n no of top words which changed attr: '+ str(changed_top_no))
result.write('\n no of bottom words which changed attr: '+ str(changed_bottom_no))
result.write('\n top words which changed attr: '+ str(top_words))
result.write('\n bottom words which changed attr: '+ str(bottom_words))
result.close()
def subfinder(l, sl):
sll=len(sl)
for ind in (i for i,e in enumerate(l) if e==sl[0]):
if l[ind:ind+sll]==sl:
return ind,ind+sll-1
def top_k_attrs(tokens, attrs, sign = None, k=None):
k = min(k, len(tokens))
if sign != None:
tokens_list = []
signs_list = []
for i in np.argpartition(attrs, -k)[-k:]:
tokens_list.append(tokens[i])
signs_list.append(sign[i])
return tokens_list , signs_list
else:
return ([tokens[i] for i in np.argpartition(attrs, -k)[-k:]])
def bottom_k_attrs(tokens, attrs, sign = None, k=None):
k = min(k, len(tokens))
if sign != None:
tokens_list = []
signs_list = []
for i in np.argpartition(attrs, k)[:k]:
tokens_list.append(tokens[i])
signs_list.append(sign[i])
return tokens_list , signs_list
else:
return ([tokens[i] for i in np.argpartition(attrs, k)[:k]])
adv.predict_and_norm()[0]
def save_big_attrs_pdf(essay_type, type_add = False, type_mod = False, type_gen = False):
ATTRS_DIR = '/content/drive/MyDrive/IG RESULTS/SKIPFLOW/P7/'
dir = ATTRS_DIR
if not os.path.exists(dir):
os.makedirs(dir)
pattern_none = [None, None, None, None, None]
if type_add:
diff_array = []
diff_attr_array = []
percent_array = []
if type_mod:
diff_array = []
diff_attr_array = []
changed_count_array = []
changed_count_top_array = []
changed_count_bottom_array = []
changed_top_words = {}
changed_bottom_words = {}
### NORMAL
a, w = get_w_a('normal')
w_normal = []
for i,x in enumerate(w):
w_normal.append(x)
w_normal = tokenizer.texts_to_sequences(w_normal)
if 'beg' in essay_type:
w_normal = pad_sequences(w_normal, maxlen=500, padding = 'post', truncating = 'post')
else:
w_normal = pad_sequences(w_normal, maxlen=500, padding = 'pre', truncating = 'pre')
labs_normal = adv.predict_and_norm(w_normal)
### ADV
a_new, w_new = get_w_a(essay_type)
w_adv = []
for i,x in enumerate(w_new):
w_adv.append(x)
w_adv = tokenizer.texts_to_sequences(w_adv)
if 'beg' in essay_type:
w_adv = pad_sequences(w_adv, maxlen=500, padding = 'post', truncating = 'post')
else:
w_adv = pad_sequences(w_adv, maxlen=500, padding = 'pre', truncating = 'pre')
labs_adv = adv.predict_and_norm(w_adv)
for i,essay in enumerate(w_new):
attrs, words= a_new[i], essay
words = words + [None]*(500 - len(words))
w[i] = w[i] + [None]*(500 - len(w[i]))
if type_add:
try:
loc = subfinder(w[i], pattern_none)[0]
except Exception as e:
loc = len(w[i])
try:
loc2 = subfinder(words, pattern_none)[0]
except Exception as e:
loc2 = len(words)
# print(loc, loc2, w[i])
if essay_type == 'song_beg' or essay_type =='false_beg' :
pattern = w[i][:5]
loc_patt = subfinder(words, pattern)[0]
# print(loc_patt)
new_w = words[:loc_patt]
new_a = attrs[:loc_patt]
other_a = attrs[loc_patt:loc2]
elif essay_type == 'song_end' or essay_type =='false_end':
new_w = words[loc:loc2]
new_a = attrs[loc:loc2]
other_a = attrs[:loc]
# print(len(new_w))
if new_w != []:
top_words_orig = top_k_attrs(w[i], a[i], k = int(0.2*loc))
top_words = top_k_attrs(words , attrs, k = int(0.2*loc2))
top_words_final = [x for x in top_words if x not in top_words_orig]
diff_attr_frac = []
for i_attrs in range(0,len(other_a), len(new_a)):
if i_attrs+len(new_a) < len(other_a):
diff_attr_frac.append( sum(other_a[ i_attrs:i_attrs+len(new_a) ]) )
else:
diff_attr_frac.append( sum(other_a[ i_attrs:len(other_a) ]) )
break
new_diff_frac = sum(diff_attr_frac) / len(diff_attr_frac)
diff_attr = sum(new_a)/new_diff_frac
diff_attr_array.append(diff_attr)
percent = len(top_words_final)/len(new_w)
percent_array.append(percent)
elif type_mod:
diff_attr = ((sum(attrs) - sum(a[i])) / sum(attrs)) *100
diff_attr_array.append(diff_attr)
###### SYN
if essay_type == 'syn':
# normal
try:
loc = subfinder(w[i], pattern_none)[0]
except Exception as e:
loc = len(w[i])
attrs_abs=[]
attrs_sign_orig =[]
for x in a[i]:
attrs_abs.append(abs(x))
if x>0:
attrs_sign_orig.append('+')
else:
attrs_sign_orig.append('-')
top_words_orig, token_signs_orig = top_k_attrs(w[i][:loc], attrs_abs[:loc], attrs_sign_orig[:loc], k = int(0.2*loc))
bottom_words_orig, bottom_token_signs_orig = bottom_k_attrs(w[i][:loc], attrs_abs[:loc], attrs_sign_orig[:loc], k = int(0.2*loc))
# adv
try:
loc2 = subfinder(words, pattern_none)[0]
except Exception as e:
loc2 = len(words)
attrs_abs=[]
attrs_sign =[]
for x in attrs:
attrs_abs.append(abs(x))
if x>0:
attrs_sign.append('+')
else:
attrs_sign.append('-')
top_words, token_signs = top_k_attrs(words[:loc2], attrs_abs[:loc2], attrs_sign[:loc2], k = int(0.2*loc2))
bottom_words, bottom_token_signs = bottom_k_attrs(words[:loc2], attrs_abs[:loc2], attrs_sign[:loc2], k = int(0.2*loc2))
changed_count_top = 0
changed_count_bottom = 0
# top
for orig_index,t in enumerate(top_words_orig):
try:
ind = w[i].index(t)
except Exception as e:
ind = -1
if ind!=-1:
if attrs_sign[ind] != attrs_sign_orig[ind]:
changed_count_top+=1
changed_top_words[t] = words[ind]
# bottom
for orig_index,t in enumerate(bottom_words_orig):
try:
ind = w[i].index(t)
except Exception as e:
ind = -1
if ind!=-1:
if attrs_sign[ind] != attrs_sign_orig[ind]:
changed_count_bottom+=1
changed_bottom_words[t] = words[ind]
changed_count_top_array.append(changed_count_top)
changed_count_bottom_array.append(changed_count_bottom)
###### SHUFFLE
if essay_type == 'shuffle':
# print(i, len(w[i]), w[i])
try:
loc = subfinder(w[i], pattern_none)[0]
except Exception as e:
loc = len(w[i])
attrs_abs=[]
attrs_sign =[]
for x in a[i]:
attrs_abs.append(abs(x))
if x>0:
attrs_sign.append('+')
else:
attrs_sign.append('-')
top_words_orig, token_signs_orig = top_k_attrs(w[i], attrs_abs, attrs_sign, k = int(0.2*loc))
try:
loc = subfinder(words, pattern_none)[0]
except Exception as e:
loc = len(words)
attrs_abs=[]
attrs_sign =[]
for x in attrs:
attrs_abs.append(abs(x))
if x>0:
attrs_sign.append('+')
else:
attrs_sign.append('-')
top_words, token_signs = top_k_attrs(words, attrs_abs, attrs_sign, k = int(0.2*loc))
changed_count = 0
for orig_index,t in enumerate(top_words_orig):
try:
ind = top_words.index(t)
except Exception as e:
ind = -1
if ind!=-1:
if token_signs_orig[orig_index] != token_signs[ind]:
changed_count+=1
changed_count_array.append(changed_count)
def Average(lst):
return sum(lst) / len(lst)
if type_add:
diff_array = [abs(labs_normal[i] -labs_adv[i]) for i in range(len(labs_adv))]
save_stats_add(Average(diff_array), Average(diff_attr_array), Average(percent_array), dir+'stats_'+essay_type+'.txt')
if type_mod and essay_type == 'syn':
diff_array = [abs(labs_normal[i] -labs_adv[i]) for i in range(len(labs_adv))]
save_stats_mod_syn(Average(diff_array), Average(diff_attr_array), Average(changed_count_top_array), \
Average(changed_count_bottom_array), set(changed_top_words), set(changed_bottom_words), \
dir+'stats_'+essay_type+'.txt')
if type_mod and essay_type == 'shuffle':
diff_array = [abs(labs_normal[i] -labs_adv[i]) for i in range(len(labs_adv))]
save_stats_mod_shuffle(Average(diff_array), Average(diff_attr_array), Average(changed_count_array), dir+'stats_'+essay_type+'.txt')
save_big_attrs_pdf('song_beg', type_add=True)
save_big_attrs_pdf('false_beg', type_add=True)
save_big_attrs_pdf('song_end', type_add=True)
save_big_attrs_pdf('false_end', type_add=True)
save_big_attrs_pdf('shuffle', type_mod=True)
save_big_attrs_pdf('syn', type_mod=True)
###NORMAL ATTRIBUTIONS PROCESSING:
def top_k_attrs(tokens, attrs, sign = None, k=None):
k = min(k, len(tokens))
if sign != None:
tokens_list = []
signs_list = []
for i in np.argpartition(attrs, -k)[-k:]:
tokens_list.append(tokens[i].strip())
signs_list.append(sign[i])
return tokens_list , signs_list
else:
return ([tokens[i].strip() for i in np.argpartition(attrs, -k)[-k:]])
# ATTRS_TSV = '/content/drive/My Drive/IG RESULTS/P1/MEMORY NET/attrs.tsv'
lines = []
with open(ATTRS_TSV) as f:
for line in f:
lines.append(line)
lines = list(set(lines[-313:]))
print(len(lines))
# GET TOP AND BOTTOM ATTRIBUTIONS
def get_counts_list_normal( top_k = 10, is_abs = False, is_sign = False, is_percent = None):
essay_list = []
counts_list = []
signs_list = []
for line in lines:
line = line.strip()
question_attrs = line.split('\t')[0]
question_tokens = []
attrs = []
for word_attr in question_attrs.split('||'):
word, attr = word_attr.split('|')
question_tokens.append(word)
if is_abs:
attrs.append(abs(float(attr)))
else:
attrs.append(float(attr))
essay_list.append(question_tokens)
if is_percent!=None:
top_k = int(is_percent*len(question_tokens))
if top_k == None:
k = len(question_tokens)
else:
k = min(top_k, len(question_tokens))
if is_sign:
signs = []
for i in attrs:
if i>0:
signs.append('+')
else:
signs.append('-')
# get top k words by attribution
c_list, sign_list = top_k_attrs(question_tokens , attrs, signs, k = k)
counts_list.extend(c_list)
signs_list.extend(sign_list)
return counts_list, signs_list, essay_list
counts_list, signs_list, essay_list = get_counts_list_normal(is_percent = 0.1, is_sign = True)
signs_dict= {}
frequent_attributions = Counter(counts_list).most_common(50)
for i in frequent_attributions:
w = i[0]
for j,w2 in enumerate(counts_list):
if w ==w2 :
if w not in signs_dict.keys():
signs_dict[w]=[]
else:
signs_dict[w].append(signs_list[j])
for k,v in signs_dict.items():
signs_dict[k] = Counter(v)
with open(ATTRS_DIR+'NORMAL_word_importance.txt','w') as f:
f.write(str(signs_dict))
###NORMAL UNATTRIBUTED STUFF:
def bottom_k_attrs(tokens, attrs, k=None):
k = min(k, len(tokens))
return ([tokens[i].strip() for i in np.argpartition(attrs, k)[:k]])
def neg_k_attrs(tokens, attrs, k=None):
k = min(k, len(tokens))
return ([tokens[i].strip() for i in np.argpartition(attrs, k)[:k] if attrs[i]<0 ])
lines = []
with open(ATTRS_TSV) as f:
for line in f:
lines.append(line)
lines = list(set(lines[-271:]))
def get_bottom_list_normal( top_k = 10, is_abs = True, is_percent = None, is_neg= False):
counts_list = []
signs_list = []
for line in lines:
line = line.strip()
question_attrs = line.split('\t')[0]
question_tokens = []
attrs = []
for word_attr in question_attrs.split('||'):
word, attr = word_attr.split('|')
question_tokens.append(word)
if is_abs:
attrs.append(abs(float(attr)))
else:
attrs.append(float(attr))
if is_percent!=None:
top_k = int(is_percent*len(question_tokens))
if top_k == None:
k = len(question_tokens)
else:
k = min(top_k, len(question_tokens))
# get top k words by attribution
if is_neg:
c_list = neg_k_attrs(question_tokens , attrs, k = k)
else:
c_list = bottom_k_attrs(question_tokens , attrs, k = k)
counts_list.extend(c_list)
return counts_list
counts_list_unattrib = get_bottom_list_normal(is_percent = 0.1)
counts_list_neg = get_bottom_list_normal(is_percent = 0.1, is_abs = False, is_neg = True)
with open(ATTRS_DIR+'NORMAL_bottom_attributed_words.txt','w') as f:
f.write(str(Counter(counts_list_unattrib).most_common(50)))
with open(ATTRS_DIR+'NORMAL_negative_attributed_words.txt','w') as f:
f.write(str(Counter(counts_list_neg).most_common(50)))
ATTRS_DIR = '/content/drive/My Drive/IG RESULTS/SKIPFLOW/P7/'
ATTRS_TSV = '/content/drive/My Drive/IG RESULTS/SKIPFLOW/P7/attrs.tsv'
def plot_and_save(curve_data, filename, x = 'num. words in vocab', y= 'relative accuracy', title= 'title'):
import matplotlib.pyplot as plt
OVERSTABILITY_CURVE_FILE = filename
plt.plot(list(curve_data.keys()), list(curve_data.values()))
# plt.xscale('symlog')
plt.xlabel(x)
plt.ylabel(y)
plt.title(title)
plt.savefig(OVERSTABILITY_CURVE_FILE, format='eps')
plt.savefig(OVERSTABILITY_CURVE_FILE.replace('eps','png'), format='png')
plt.tight_layout()
plt.show()
def plot_and_save_both(a,b, filename, x = 'num. words in vocab', y= 'relative accuracy', title= 'title'):
import matplotlib.pyplot as plt
OVERSTABILITY_CURVE_FILE = filename
plt.plot(a,b)
# plt.xscale('symlog')
ax = plt.gca()
ax.invert_xaxis()
plt.xlabel(x)
plt.title(title)
plt.ylabel(y)
plt.savefig(OVERSTABILITY_CURVE_FILE, format='eps')
plt.savefig(OVERSTABILITY_CURVE_FILE.replace('eps','png'), format='png')
plt.tight_layout()
plt.show()
### NORMAL ATTRS VARIATION
data = E_list['normal']
for i in range(len(data)):
attrs, words,c_data = adv.explain(data[i])
attrs = attrs[c_data:]
attrs = [abs(x) for x in attrs]
words = words[c_data:]
s= {}
l = len(attrs)
for j in range(0, l, l//5):
s[(j)] = sum(attrs[j: j+ l//5])
dir = ATTRS_DIR+'normal/'
plot_and_save(s, dir+str(i)+'_attrs_variation.txt', x= 'word number', y='absolute attribution')
# with open('/content/drive/My Drive/IG RESULTS/MEM MODELS/tokenizer.pkl', 'rb') as f:
word_to_idx = tokenizer.word_index
idx_to_word = {v: k for k, v in word_to_idx.items()}
ATTRS_DIR = '/content/drive/My Drive/IG RESULTS/SKIPFLOW/P7/'
ATTRS_TSV = '/content/drive/My Drive/IG RESULTS/SKIPFLOW/P7/attrs.tsv'
lines = []
with open(ATTRS_TSV) as f:
for line in f:
lines.append(line)
lines = list(set(lines[-313:]))
def get_essay_list(tsv_lines):
essay_list = []
attrs_list = []
l = []
for line in tsv_lines:
line = line.strip()
question_attrs = line.split('\t')[0]
question_tokens = []
attrs = []
for word_attr in question_attrs.split('||'):
word, attr = word_attr.split('|')
question_tokens.append(word)
attrs.append(float(attr))
question_tokens_idx = tokenizer.texts_to_sequences([question_tokens])
l.append(len(question_tokens_idx[0]))
question_tokens_idx = pad_sequences(question_tokens_idx, maxlen=MAX_SEQUENCE_LENGTH)[0]
essay_list.append(question_tokens_idx)
# attrs_padded = [0.0]*(MAX_SEQUENCE_LENGTH - l) + attrs
attrs_list.append(attrs)
return essay_list, attrs_list, l
essay_list, attrs_list, l = get_essay_list(lines)
# STATISTICS
def npos(orig, new):
count = 0
for i in range(len(orig)):
if new[i]>orig[i]:
count+=1
return (count/len(orig))*100
def nneg(orig, new):
count = 0
for i in range(len(orig)):
if new[i]<orig[i]:
count+=1
return (count/len(orig))*100
def nsame(orig, new):
count = 0
for i in range(len(orig)):
if new[i]==orig[i]:
count+=1
return (count/len(orig))*100
def mu(orig, new):
s=0
n = len(orig)
for i in range(n):
s+=(orig[i] - new[i])
return (s/n)/30
def absmu(orig, new):
s=0
n = len(orig)
for i in range(n):
s+=(orig[i] - new[i])
return ((abs(s)/n)/30)*100
def sd(orig, new):
mu_val = mu(orig, new)
s=0
n = len(orig)
for i in range(n):
s+=(orig[i] - new[i] - mu_val)**2
return (((s/n)**(1/2))/30)*100
def mupos(orig, new):
s=0
n = len(orig)
for i in range(n):
if new[i]>orig[i]:
s+=(new[i] - orig[i])
return ((s/n)/30)*100
def muneg(orig, new):
s=0
n = len(orig)
for i in range(n):
if orig[i] > new[i]:
s+=(orig[i] - new[i])
return ((s/n)/30)*100
def get_pred_stats(orig, new, filename, K):
b = ('NPOS', npos(orig, new))
c = ('NNEG', nneg(orig, new))
d = ('NSAME', nsame(orig, new))
e = ('MU', mu(orig, new))
f = ('ABSMU', absmu(orig, new))
g = ('SD', sd(orig, new))
h = ('MUPOS', mupos(orig, new))
i = ('MUNEG', muneg(orig, new))
with open(filename, 'a') as file:
file.write(str(K)+"___"+str([h,i,b,c,g])+ "\n")
pred_array_orig = []
pred = adv.predict_and_norm(np.array(essay_list))
pred_array_orig.extend(pred)
# from xhtml2pdf import pisa
def convert_html_to_pdf(source_html, output_filename):
result_file = open(output_filename, "w+b")
pisa_status = pisa.CreatePDF(source_html, dest=result_file)
result_file.close()
def save_attrs(data, K, essay_type):
dir = ATTRS_DIR+essay_type+'/'
if not os.path.exists(dir):
os.makedirs(dir)
attrs, words,c_data = adv.explain(data)
label_new = adv.predict_and_norm([data])[0]
html = adv.visualize_token_attrs(words[c_data:], attrs[c_data:])
convert_html_to_pdf(html, dir+str(K)+'_'+str(int(label_new))+'.pdf')
html = IG.visualize_token_attrs(words, attrs)
# def get_attrs_alibi(v):
# baseline = np.zeros(x_val[0:1].shape)
# baseline[0][0] = adv.tokenizer.word_index['a']
# explanation = ig.explain(v, baselines=baseline)
# attrs = explanation.attributions
# attrs = attrs.sum(axis=2)
# return attrs
def sequence_to_text( list_of_indices):
count = 0
words = [adv.reversed_word_map.get(letter) for letter in list_of_indices]
for x in words:
if x == None:
count+=1
return(words, count)
# # def explain(self, essay):
# # attrs = get_attrs_alibi(np.array([essay]))[0]
# # words,count = sequence_to_text(essay)
# # assert len(words[count:]) == len(attrs[count:])
# # html = visualize_token_attrs(words[count:], attrs[count:])
# # return attrs, words, count
# GET TOP WORDS ADDITION GRAPH
from sklearn.metrics import cohen_kappa_score
import numpy as np
import random
from random import randint
random.seed(42)
def top_k_attrs(tokens, attrs,k=None):
k = min(k, len(tokens))
return ([tokens[i] for i in np.argpartition(attrs, -k)[-k:]])
d = {}
labels_inc = []
inc_essay = []
p_list = {}
for K in range(0,6):
percent = K*0.2
preds_new = []
new_essay_list = []
for id, essay in enumerate(essay_list):
top_k = int(percent* len(attrs_list[id]))
attrs = attrs_list[id]
question_tokens = essay_list[id]
try:
c_list = top_k_attrs(question_tokens[-len(attrs):] , attrs, k = top_k)
# print(top_k, len(c_list), c_list)
except Exception as e:
print(e)
c_list = list(set(question_tokens))
if top_k==0:
c_list = []
count = 0
new_essay = []
for i in range(len(question_tokens[-len(attrs):])):
if question_tokens[-len(attrs):][i] in c_list and count<top_k:
new_essay.append(question_tokens[-len(attrs):][i])
count+=1
# print(len(new_essay)/ len(question_tokens[-len(attrs):]), new_essay, question_tokens[-len(attrs):])
new_essay_padded = pad_sequences([new_essay], maxlen=MAX_SEQUENCE_LENGTH)[0]
pred = adv.predict_and_norm(np.array([new_essay_padded]))[0]
if round(abs(pred - pred_array_orig[id])) == 1:
# print('in')
if id not in p_list.keys():
p_list[id] = percent*100
else:
pass
new_essay_list.append(new_essay_padded)
pred = adv.predict_and_norm(np.array(new_essay_list))
preds_new.extend(pred)
acc = cohen_kappa_score(preds_new, pred_array_orig, weights='quadratic')
get_pred_stats(pred_array_orig, preds_new, ATTRS_DIR+'stats_top.txt', int(percent*100))
d[percent*100] = acc
l = p_list.values()
sum(l)/len(l)
d
plot_and_save(d,ATTRS_DIR+'adding_top', x = '% length of response', y ='relative QWK', title= 'iteratively adding words(in order of importance)')
# GET BOTTOM WORDS REMOVAL GRAPH
def bottom_k_attrs(tokens, attrs,k=None):
k = min(k, len(tokens))
return ([tokens[i] for i in np.argpartition(attrs, k)[:k]])
d2_keys = []
d2_vals = []
p_list = {}
for K in range(0,6):
percent = K*0.2
preds_new = []
new_essay_list = []
for id, essay in enumerate(essay_list):
top_k = int(percent* len(attrs_list[id]))
attrs = attrs_list[id]
question_tokens = essay_list[id]
attrs = [abs(x) for x in attrs]
try:
c_list = bottom_k_attrs(question_tokens[-len(attrs):] , attrs, k = top_k)
# print(top_k, c_list)
except Exception as e:
c_list = list(set(question_tokens))
if top_k==0:
c_list = []
count = 0
new_essay = []
for i in range(len(question_tokens[-len(attrs):])):
if question_tokens[-len(attrs):][i] in c_list and count<top_k:
count +=1
pass
else:
new_essay.append( question_tokens[-len(attrs):][i])
# print(len(new_essay)/ len(question_tokens[-len(attrs):]))
new_essay_padded = pad_sequences([new_essay], maxlen=MAX_SEQUENCE_LENGTH)[0]
pred = adv.predict_and_norm(np.array([new_essay_padded]))[0]
if round(abs(pred - pred_array_orig[id])) == 1:
# print('in')
if id not in p_list.keys():
p_list[id] = percent*100
else:
pass
new_essay_list.append(new_essay_padded)
pred = adv.predict_and_norm(np.array(new_essay_list))
preds_new.extend(pred)
acc = cohen_kappa_score(preds_new, pred_array_orig, weights='quadratic')
# print(acc)
get_pred_stats(pred_array_orig, preds_new, ATTRS_DIR+'stats_bottom.txt', int(100-percent*100))
d2_keys.append(100-percent*100)
d2_vals.append(acc)
l = p_list.values()
sum(l)/len(l)
plot_and_save_both(d2_keys, d2_vals, ATTRS_DIR+'removing_bottom', x = '% length of response', y ='relative QWK', title= 'iteratively removing words(in reverse order of importance)')
```
| github_jupyter |
```
%load_ext sql
%sql sqlite://
# Create tables & insert some random numbers
# Note: in Postgresql, try the generate_series function...
%sql DROP TABLE IF EXISTS R; DROP TABLE IF EXISTS S; DROP TABLE IF EXISTS T;
%sql CREATE TABLE R (A int); CREATE TABLE S (A int); CREATE TABLE T (A int);
for i in range(1,6):
%sql INSERT INTO R VALUES (:i)
for i in range(1,10,2):
%sql INSERT INTO S VALUES (:i)
for i in range(1,11,3):
%sql INSERT INTO T VALUES (:i)
%%sql
drop table if exists product; -- This needs to be dropped if exists, see why further down!
drop table if exists company;
pragma foreign_keys = ON; -- WARNING by default off in sqlite
create table company (
cname varchar primary key, -- company name uniquely identifies the company.
stockprice money, -- stock price is in money
country varchar); -- country is just a string
insert into company values ('ToyWorks', 25.0, 'USA');
insert into company values ('ToyFriends', 65.0, 'China');
insert into company values ('ToyCo', 15.0, 'China');
create table product(
pname varchar, -- name of the product
price money, -- price of the product
category varchar, -- category
manufacturer varchar, -- manufacturer
primary key (pname, manufacturer),
foreign key (manufacturer) references company(cname));
insert into product values('Pikachu', 19.99, 'Toy', 'ToyWorks');
insert into product values('Pikachu', 19.99, 'Toy', 'ToyFriends');
insert into product values('Pokeball', 29.99, 'Electronic', 'ToyCo');
insert into product values('Bulbasaur', 149.99, 'Toy', 'ToyFriends');
insert into product values('Charizard', 203.99, 'Toy', 'ToyCo');
insert into product values('PokeCamera', 19.99, 'Electronic', 'ToyWorks');
```
Activity 2-3:
-------------
Multi-table queries
Exercise #1:
-----------
For three tables $R,S,T$ that only have one attribute $A$:
* R = {1,2,3,4,5}
* S = {1,3,5,7,9}
* T = {1,4,7,10}
Can you write a query to select $R \cap (S \cup T)$- in other words elements that are in $R$ and either $S$ or $T$?
Write your query here:
```
%sql SELECT * FROM R WHERE A IN S OR A IN T;
```
Now test your query above for the case where $S = \emptyset$- what happens and why?
Execute the below, then re-run your query above
```
%%sql
delete from S;
%sql SELECT * FROM R WHERE A IN S OR A IN T;
```
Exercise #2
-----------
* Schema is same as before
> Product (<u>pname</u>, price, category, manufacturer)<br>
> Company (<u>cname</u>, stockPrice, country)
* Our goal is to answer the following question:
> Find all categories of products that are made by Chinese companies
Write your query here:
```
%sql SELECT DISTINCT category FROM product,company WHERE manufacturer = cname AND country = "China";
```
| github_jupyter |
<a href="https://colab.research.google.com/github/DanIulian/BookStore/blob/master/02_rainbow(1)(1).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Not Quite Rainbow
```
# !apt install xvfb python-opengl ffmpeg -y > /dev/null 2>&1
# !pip install pyvirtualdisplay > /dev/null 2>&1
# !pip install -U torch > /dev/null 2>&1
# !pip install git+git://github.com/maximecb/gym-minigrid.git@master#egg=gym-minigrid > /dev/null 2>&1
!pip uninstall gym-minigrid gym
!pip install git+git://github.com/floringogianu/gym-minigrid.git@poli#egg=gym-minigrid > /dev/null 2>&1
print("\nRuntime > Restart Runtime after this cell executes!")
import itertools
import random
from argparse import Namespace
from collections import deque, defaultdict
from copy import deepcopy
import numpy as np
import torch
import torch.nn as nn
import torch.optim as O
from torchvision import transforms as T
from PIL import Image
import gym
import gym_minigrid
from gym_minigrid.wrappers import RGBImgPartialObsWrapper, ImgObsWrapper, ReseedWrapper
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
sns.set()
DEVICE = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(f"OpenAI Gym: {gym.__version__}. \t\tShould be: ~0.15.x")
print(f"PyTorch : {torch.__version__}. \tShould be: >=1.2.x+cu100")
print(f"DEVICE : {DEVICE}. \t\tShould be: cuda")
def reset_rng(seed=42):
print(f"Setting all rngs to seed={seed}")
torch.manual_seed(seed)
np.random.seed(seed)
random.seed(seed)
reset_rng()
envs = Namespace(
easy="MiniGrid-Empty-5x5-v0",
maze="MiniGrid-SimpleCrossingS9N1-v0",
two_maze="MiniGrid-SimpleCrossingS9N2-v0",
large_maze="MiniGrid-SimpleCrossingS11N1-v0",
overestimation="MiniGrid-OverEstimation-9x9-v0",
random_overestimation="MiniGrid-OverEstimation-Random-9x9-v0",
fetch="MiniGrid-Fetch-8x8-N3-v0"
)
# Define some helpers: Gym Wrappers and visualization functions
class TorchWrapper(gym.ObservationWrapper):
""" Applies a couple of transformations depending on the mode.
Receives numpy arrays and returns torch tensors.
"""
def __init__(self, env):
super().__init__(env)
self._transform = T.Compose([
lambda obs: (obs * int(255 / 9)).swapaxes(1, 0),
lambda obs: torch.from_numpy(obs).permute(2, 1, 0)
])
def observation(self, obs):
return self._transform(obs).unsqueeze(0).to(DEVICE)
class FrameStack(gym.Wrapper):
"""Stack k last frames. """
def __init__(self, env, k, verbose=False):
super().__init__(env)
self.k = k
self.frames = deque([], maxlen=k)
def reset(self):
observation = self.env.reset()
for _ in range(self.k):
self.frames.append(observation)
return self._get_ob()
def step(self, action):
observation, reward, done, info = self.env.step(action)
self.frames.append(observation)
return self._get_ob(), reward, done, info
def _get_ob(self):
assert len(self.frames) == self.k
if self.k == 1:
return self.frames.pop()
return np.concatenate(list(self.frames), axis=2)
def show_representations(env_name="MiniGrid-SimpleCrossingS9N1-v0", tile_size=8):
seed = torch.randint(100, (1,)).item()
env = ImgObsWrapper(RGBImgPartialObsWrapper(gym.make(env_name), tile_size=tile_size))
print("Action-space: ", env.action_space.n)
env.seed(seed)
rgb_obs = env.reset()
env = ImgObsWrapper(gym.make(env_name))
env.seed(seed)
sym_obs = env.reset()
print("RGB:", rgb_obs.shape)
print("SYM:", sym_obs.shape)
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(12, 36))
ax1.imshow(rgb_obs)
ax2.imshow((sym_obs * int(255 / 9)).swapaxes(1, 0))
ax3.imshow(env.render(mode="rgb_array"))
def plot_stats(stats, y="ep_rewards", hue=None, window=10):
df = pd.DataFrame(stats)
if window:
new_col = f"avg_{y}"
if hue is not None:
df[new_col] = df.groupby(hue)[y].rolling(window=window).mean().reset_index(0,drop=True)
else:
df[new_col] = df[y].rolling(window=window).mean()
y = f"avg_{y}" if window else y
with matplotlib.rc_context({'figure.figsize':(10, 6)}):
sns.lineplot(x="step_idx", y=y, hue=hue, data=df)
```
## Let's take a look at the environments
```
for k, v in vars(envs).items():
print(f"{k:<24}: {v}")
# you can execute this a few times to get an idea about the two possible
# views of the agent
show_representations(envs.overestimation, tile_size=8)
```
## Let's define the training routine.
It takes an agent and an environment and implements the action-perception loop.
```
def train(agent, env, step_num=100_000):
stats, N = {"step_idx": [0], "ep_rewards": [0.0], "ep_steps": [0.0]}, 0
state, done = env.reset().clone(), False
for step in range(step_num):
action = agent.step(state)
state_, reward, done, _ = env.step(action)
agent.learn(state, action, reward, state_, done)
# some envs just update the state and are not returning a new one
state = state_.clone()
# stats
stats["ep_rewards"][N] += reward
stats["ep_steps"][N] += 1
if done:
# episode done, reset env!
state, done = env.reset().clone(), False
# some more stats
if N % 10 == 0:
print("[{0:3d}][{1:6d}], R/ep={2:6.2f}, steps/ep={3:2.0f}.".format(
N, step,
torch.tensor(stats["ep_rewards"][-10:]).mean().item(),
torch.tensor(stats["ep_steps"][-10:]).mean().item(),
))
stats["ep_rewards"].append(0.0) # reward accumulator for a new episode
stats["ep_steps"].append(0.0) # reward accumulator for a new episode
stats["step_idx"].append(step)
N += 1
print("[{0:3d}][{1:6d}], R/ep={2:6.2f}, steps/ep={3:2.0f}.".format(
N, step, torch.tensor(stats["ep_rewards"][-10:]).mean().item(),
torch.tensor(stats["ep_steps"][-10:]).mean().item(),
))
stats["agent"] = [agent.__class__.__name__ for _ in range(N+1)]
return stats
```
## Start implementing the DQN agent
## 1. Experience Replay
#### TASK 1: implement `sample` method.
```
class ReplayMemory:
def __init__(self, size=1000, batch_size=32):
self._buffer = deque(maxlen=size)
self._batch_size = batch_size
def push(self, transition):
self._buffer.append(transition)
def sample(self):
""" Sample from self._buffer
Should return a tuple of tensors of size:
(
states: N * (C*K) * H * W, (torch.uint8)
actions: N * 1, (torch.int64)
rewards: N * 1, (torch.float32)
states_: N * (C*K) * H * W, (torch.uint8)
done: N * 1, (torch.uint8)
)
where N is the batch_size, C is the number of channels = 3 and
K is the number of stacked states.
"""
# sample
s_list, a_list, r_list, s_list_, d_list = zip(*random.sample(self._buffer, self._batch_size))
s = torch.cat(s_list).to(DEVICE)
s_ = torch.cat(s_list_).to(DEVICE)
a = torch.LongTensor(a_list).unsqueeze(-1).to(DEVICE)
r = torch.DoubleTensor(r_list).unsqueeze(-1).to(DEVICE)
d = torch.zeros((self._batch_size, 1), dtype=torch.uint8).to(DEVICE)
d[np.array(d_list) == True] = 1
return (s, a, r, s_, d)
def __len__(self):
return len(self._buffer)
```
## 2. $\epsilon$-greedy schedule.
#### TASK 2: Implement the epsilon-greedy schedule
```
def get_epsilon_schedule(start=1.0, end=0.1, steps=500):
""" Returns either:
- a generator of epsilon values
- a function that receives the current step and returns an epsilon
The epsilon values returned by the generator or function need
to be degraded from the `start` value to the `end` within the number
of `steps` and then continue returning the `end` value indefinetly.
You can pick any schedule (exp, poly, etc.). I tested with linear decay.
"""
t = 0
while True:
if t >= steps:
yield end
else:
eps = start - (start - end) / steps * t
t += 1
yield eps
# test it, it needs to look nice
epsilon = get_epsilon_schedule(1.0, 0.1, 100)
plt.plot([next(epsilon) for _ in range(500)])
# or if you prefer a function
# epsilon_fn = get_epsilon_schedule(1.0, 0.1, 100)
# plt.plot([epsilon(step_idx) for step_idx in range(500)])
```
### Define a Neural Network Approximator for your Agents
```
class ByteToFloat(nn.Module):
""" Converts ByteTensor to FloatTensor and rescales.
"""
def forward(self, x):
assert (
x.dtype == torch.uint8
), "The model expects states of type ByteTensor."
return x.float().div_(255)
class View(nn.Module):
def forward(self, x):
return x.view(x.size(0), -1)
def get_estimator(action_num, input_ch=3, lin_size=32):
return nn.Sequential(
ByteToFloat(),
nn.Conv2d(input_ch, 16, kernel_size=3),
nn.ReLU(inplace=True),
nn.Conv2d(16, 16, kernel_size=2),
nn.ReLU(inplace=True),
nn.Conv2d(16, 16, kernel_size=2),
nn.ReLU(inplace=True),
View(),
nn.Linear(9 * 16, lin_size),
nn.ReLU(inplace=True),
nn.Linear(lin_size, action_num),
).to(DEVICE)
```
## 3. DQN Agent, finally
#### TASK 3:
- implement the `step()` method
- implement the `learn()` method
- implement the `_update()` method
```
class DQN:
def __init__(
self,
estimator,
buffer,
optimizer,
epsilon_schedule,
action_num,
gamma=0.92,
update_steps=4,
update_target_steps=10,
warmup_steps=100,
):
self._estimator = estimator
self._target_estimator = deepcopy(estimator)
self._buffer = buffer
self._optimizer = optimizer
self._epsilon = epsilon_schedule
self._action_num = action_num
self._gamma = gamma
self._update_steps=update_steps
self._update_target_steps=update_target_steps
self._warmup_steps = warmup_steps
self._step_cnt = 0
assert warmup_steps > self._buffer._batch_size, (
"You should have at least a batch in the ER.")
def step(self, state):
# implement an epsilon greedy policy using the
# estimator and epsilon schedule attributes.
# warning, you should make sure you are not including
# this step into torch computation graph
if self._step_cnt < self._warmup_steps:
return torch.randint(self._action_num, (1,)).item()
# return the action according to the self._epsilon schedule
# you defined earlier
if np.random.random() < next(self._epsilon):
return torch.randint(self._action_num, (1,)).item()
else:
with torch.no_grad():
acts = self._estimator(state)
return acts.argmax(dim=1)
def learn(self, state, action, reward, state_, done):
# TODO: add transition to the experience replay
self._buffer.push((state, action, reward, state_, done))
if self._step_cnt < self._warmup_steps:
self._step_cnt += 1
return
if self._step_cnt % self._update_steps == 0:
# TODO: sample from experience replay and do an update
s, a, r, s_, d = self._buffer.sample()
self._update(s, a, r, s_, d)
if self._step_cnt % self._update_target_steps == 0:
# TODO: update the target estimator (hint, use pytorch state_dict methods)
self._target_estimator.load_state_dict(self._estimator.state_dict())
self._step_cnt += 1
def _update(self, states, actions, rewards, states_, done):
# compute the DeepQNetwork update. Carefull not to include the
# target network in the computational graph.
# Compute Q(s, * | θ) and Q(s', . | θ^)
q_values = self._estimator(states)
with torch.no_grad():
q_values_ = self._target_estimator(states_)
qsa = torch.gather(q_values, dim=1, index=actions)
qsa_, _ = torch.max(q_values_, dim=1)
qsa_ = qsa_.unsqueeze(-1)
# compute Q(s, a) and max_a' Q(s', a')
# qsa =
# qsa_ =
# compute target values
target_qsa = rewards + (1 - done) * self._gamma * qsa_
# at this step you should check the target values
# are looking about right :). You can use this code.
# if rewards.squeeze().sum().item() > 0.0:
# print("R: ", rewards.squeeze())
# print("T: ", target_qsa.squeeze())
# print("D: ", done.squeeze())
# compute the loss and average it over the entire batch
loss = nn.functional.mse_loss(qsa.float(), target_qsa.float())
self._optimizer.zero_grad()
loss.backward()
self._optimizer.step()
# backprop and optimize
env = gym.make(envs.easy)
env = TorchWrapper(ImgObsWrapper(env))
net = get_estimator(env.action_space.n)
stats = train(
DQN(
net,
ReplayMemory(size=1000, batch_size=32),
O.Adam(net.parameters(), lr=1e-3, eps=1e-4),
get_epsilon_schedule(start=1.0, end=0.1, steps=4000),
env.action_space.n,
warmup_steps=100,
update_steps=2,
),
env,
step_num=7_000 # change the experiment length if it's learning but not reaching about .95
)
plot_stats(stats)
```
## 4. Train on a partial observable maze
```
show_representations(envs.maze)
# show_representations(envs.large_maze)
step_num=30_000
seeds = [2, 5] # add more map configurations
hist_len = [1, 2, 3] # increase it to two and compare
common_seed = np.random.randint(1000)
stats = defaultdict(list)
for K in hist_len:
print(f"Started training with hist_len={K}.")
# 102 worked nicely here :))
reset_rng(common_seed) # we want each experiment to have the same starting conditions
env = gym.make(envs.maze)
env = TorchWrapper(FrameStack(ImgObsWrapper(ReseedWrapper(env, seeds=seeds)), k=K))
net = get_estimator(env.action_space.n, input_ch=K*3, lin_size=64)
stats_ = train(
DQN(
net,
ReplayMemory(size=10_000, batch_size=32),
O.Adam(net.parameters(), lr=1e-3, eps=1e-4),
get_epsilon_schedule(start=1.0, end=0.1, steps=10_000),
env.action_space.n,
warmup_steps=1000,
update_steps=2,
update_target_steps=8
),
env,
step_num=step_num
)
stats_["hist_len"] = [K] * len(stats_["ep_rewards"])
for k, v in stats_.items():
stats[k] += v
plot_stats(stats, hue="hist_len")
```
## 5. Double DQN
#### TASK 4: Implement the _update() method for DoubleDQN
```
class DoubleDQN(DQN):
def _update(self, states, actions, rewards, states_, done):
# compute the DeepQNetwork update. Carefull not to include the
# target network in the computational graph.
# Compute Q(s, * | θ) and Q(s', . | θ^)
# q_values =
# q_values_ =
#... and the rest of it
# Compute Q(s, * | θ) and Q(s', . | θ^)
with torch.no_grad():
q_values_target = self._target_estimator(states_)
q_values_doubledqn = self._estimator(states_)
q_values_online_net = self._estimator(states)
qsa_online_net = torch.gather(q_values_online_net, dim=1, index=actions)
actions_max = torch.argmax(q_values_doubledqn, dim=1).unsqueeze(-1)
qsa_target_net = torch.gather(q_values_target, dim=1, index=actions_max)
#qsa = torch.gather(q_values, dim=1, index=actions)
#qsa_, _ = torch.max(q_values_, dim=1)
#qsa_ = qsa_.unsqueeze(-1)
# compute Q(s, a) and max_a' Q(s', a')
# qsa =
# qsa_ =
# compute target values
target_qsa = rewards + (1 - done) * self._gamma * qsa_target_net
# at this step you should check the target values
# are looking about right :). You can use this code.
# if rewards.squeeze().sum().item() > 0.0:
# print("R: ", rewards.squeeze())
# print("T: ", target_qsa.squeeze())
# print("D: ", done.squeeze())
# compute the loss and average it over the entire batch
loss = nn.functional.mse_loss(qsa_online_net.float(), target_qsa.float())
self._optimizer.zero_grad()
loss.backward()
self._optimizer.step()
# backprop and optimize
env = gym.make(envs.easy)
env = TorchWrapper(ImgObsWrapper(env))
net = get_estimator(env.action_space.n)
stats = train(
DoubleDQN(
net,
ReplayMemory(size=1000, batch_size=32),
O.Adam(net.parameters(), lr=1e-3, eps=1e-4),
get_epsilon_schedule(start=1.0, end=0.1, steps=4000),
env.action_space.n,
warmup_steps=100,
update_steps=2,
update_target_steps=16
),
env,
step_num=7_000 # change the experiment length if it's learning but not reaching about .95
)
plot_stats(stats)
```
## 6. DQN vs Double DQN: OverEstimation environment
```
show_representations(envs.overestimation)
step_num = 50_000 # change the experiment length
stats = defaultdict(list)
common_seed = np.random.randint(1000)
for Agent in [DQN, DoubleDQN]:
# 256 worked nicely here :))
reset_rng(common_seed) # we want each experiment to have the same starting conditions
env = TorchWrapper(ImgObsWrapper(gym.make(envs.overestimation)))
net = get_estimator(env.action_space.n)
agent_name = Agent.__name__
print(f"\n{agent_name} started training.")
stats_ = train(
Agent(
net,
ReplayMemory(size=5000, batch_size=32),
O.Adam(net.parameters(), lr=1e-3, eps=1e-4),
get_epsilon_schedule(start=1.0, end=0.1, steps=5000),
env.action_space.n,
warmup_steps=3000,
update_steps=2,
update_target_steps=256
),
env,
step_num=step_num
)
for k, v in stats_.items():
stats[k] += v
plot_stats(stats, hue="agent")
```
## 7. DQN vs DoubleDQN: Maze Environment
```
show_representations(envs.maze)
env_name = envs.maze
step_num = 50_000 # change the experiment length
seeds = [2, 5] # add more maps
K = 2
stats = defaultdict(list)
common_seed = np.random.randint(1000)
for Agent in [DQN, DoubleDQN]:
# maybe 621? :))
reset_rng(common_seed) # we want each experiment to have the same starting conditions
env = gym.make(env_name)
env = TorchWrapper(FrameStack(ImgObsWrapper(ReseedWrapper(env, seeds=seeds)), k=K))
net = get_estimator(env.action_space.n, input_ch=K*3, lin_size=64)
agent_name = Agent.__name__
print(f"\n{agent_name} started training.")
stats_ = train(
Agent(
net,
ReplayMemory(size=10_000, batch_size=32),
O.Adam(net.parameters(), lr=1e-3, eps=1e-4),
get_epsilon_schedule(start=1.0, end=0.1, steps=10_000),
env.action_space.n,
warmup_steps=1000,
update_steps=2,
update_target_steps=256
),
env,
step_num=step_num
)
for k, v in stats_.items():
stats[k] += v
plot_stats(stats, hue="agent")
```
## 8. Dueling DQN
### BONUS TASK: Implement Dueling DQN and compare it on any env you want with the other two methods.
Extra-credits for showing statistically-relevant graphs, unlike I did in
this lab.
```
# Dueling DQN can be implemented fairly easy by defining a new
# nn.Module instead of the one returned by `get_estimator()`.
# It can **probably** be used with both DQN and DoubleDQN as they are.
class DuelingNet(nn.Module):
def __init__(self, num_inputs, num_outputs):
super(DuelingNet, self).__init__()
# define convolutional feature extractor
# advantage layers
# value layers
self.num_inputs = num_inputs
self.num_outputs = num_outputs
self.conv = nn.Sequential(
ByteToFloat(),
nn.Conv2d(self.num_inputs, 16, kernel_size=3),
nn.ReLU(inplace=True),
nn.Conv2d(16, 16, kernel_size=2),
nn.ReLU(inplace=True),
nn.Conv2d(16, 16, kernel_size=2),
nn.ReLU(inplace=True)
)
self.value_stream = nn.Sequential(
nn.Linear(9 * 16, 32),
nn.ReLU(inplace=True),
nn.Linear(32, 1)
)
self.advantage_stream = nn.Sequential(
nn.Linear(9 * 16, 32),
nn.ReLU(inplace=True),
nn.Linear(32, self.num_outputs)
)
def forward(self, x):
features = self.conv(x)
features = features.view(features.size(0), -1)
value_estimate = self.value_stream(features)
advantage_estimate = self.advantage_stream(features)
qvals = values + (advantages - advantages.mean())
return qvals
class DuelingDQN(DoubleDQN):
def __init__(
self,
estimator,
buffer,
optimizer,
epsilon_schedule,
action_num,
gamma=0.92,
update_steps=4,
update_target_steps=10,
warmup_steps=100,
):
super(DuelingDQN, self).__init__(
estimator,
buffer,
optimizer,
epsilon_schedule,
action_num,
gamma,
update_steps,
update_target_steps,
warmup_steps
)
env = gym.make(envs.easy)
env = TorchWrapper(ImgObsWrapper(env))
net = get_estimator(env.action_space.n)
stats = train(
DuelingDQN(
net,
ReplayMemory(size=1000, batch_size=32),
O.Adam(net.parameters(), lr=1e-3, eps=1e-4),
get_epsilon_schedule(start=1.0, end=0.1, steps=4000),
env.action_space.n,
warmup_steps=100,
update_steps=2,
update_target_steps=16
),
env,
step_num=7_000 # change the experiment length if it's learning but not reaching about .95
)
plot_stats(stats)
env_name = envs.maze
step_num = 50_000 # change the experiment length
seeds = [2, 5] # add more maps
K = 2
stats = defaultdict(list)
common_seed = np.random.randint(1000)
for Agent in [DQN, DoubleDQN, DuelingDQN]:
# maybe 621? :))
reset_rng(526) # we want each experiment to have the same starting conditions
env = gym.make(env_name)
env = TorchWrapper(FrameStack(ImgObsWrapper(ReseedWrapper(env, seeds=seeds)), k=K))
net = get_estimator(env.action_space.n, input_ch=K*3, lin_size=64)
agent_name = Agent.__name__
print(f"\n{agent_name} started training.")
stats_ = train(
Agent(
net,
ReplayMemory(size=10_000, batch_size=32),
O.Adam(net.parameters(), lr=1e-3, eps=1e-4),
get_epsilon_schedule(start=1.0, end=0.1, steps=10_000),
env.action_space.n,
warmup_steps=1000,
update_steps=2,
update_target_steps=256
),
env,
step_num=step_num
)
for k, v in stats_.items():
stats[k] += v
```
| github_jupyter |
## Importing packages
Throughout this tutorial, we will use the following common Python packages:
```
# Use these packages to easily access files on your hard drive
import os, sys, glob
# The Numpy package allows you to manipulate data (mainly numerical)
import numpy as np
# The Pandas package allows more advanced data manipulation e.g. in structured data frames
import pandas as pd
# The Matplotlib package is for plotting - uses the same syntax as plotting in Matlab (figures, axes etc)
import matplotlib.pyplot as plt
# Seaborn is a higher-level package for plotting that calls functions in Matplotlib,
# you can usually input your Pandas dataframes to get pretty plots in 1 or 2 lines
import seaborn as sns
# We will use Scipy for advanced computation like model fitting
import scipy
```
## Solutions
#### 1. Create two lists that separate numbers (eg. from 1-100) divisible by 3 and numbers not divisible by 3.
```
y1 = []
y2 = []
for x in np.arange(1,100):
if x%3 == 0:
#if divisible by 3, put the number in y1
y1.append(x)
else:
#if not divisible by 3, put the number in y1
y2.append(x)
# display(y1)
# display(y2)
```
#### 2. Keep generating random numbers until a generated number is greater than 0.8 and store the number of times it takes you to get this number
```
u = np.random.rand()
n = 1
while u < 0.8:
u = np.random.rand()
n = n+1
display(u)
display(n)
```
#### 3. Generate some random data in two variables of equal length and make a scatter plot using matplotlib
```
var1 = np.random.rand(100)
var2 = np.random.rand(100)
plt.scatter(var1,var2)
```
#### 4. Generate some data for a linear relationship between two variables (e.g. age and height of schoolchildren), put them in a Pandas dataframe with 2 named columns, and use Seaborn to create a scatterplot with regression line
```
age = 5 + np.random.rand(100)*7
height = 108 + (152-108)*((age-5)/7) + np.random.randn(100)*20
age_height = pd.DataFrame.from_dict({'age':age,'height':height}).sort_values(by=['age','height'])
display(age_height.head())
sns.regplot(data = age_height, x = 'age', y = 'height')
```
#### 5. Create a Pandas dataframe with height data for 5 age groups and use Seaborn to turn this into a barplot with errorbars and an overlaid stripplot or swarmplot.
```
age_height['group'] = age_height['age'].apply(lambda x: np.floor(x)-4)
age_height
sns.barplot(data = age_height.query('group < 6'), x = 'group', y = 'height', alpha = .5)
sns.swarmplot(data = age_height.query('group < 6'), x = 'group', y = 'height')
```
| github_jupyter |
```
### First imports and default parameters
%matplotlib inline
import warnings
warnings.filterwarnings("ignore")
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# Overwritting matplotlib default linestyle of negative contours
matplotlib.rcParams['contour.negative_linestyle'] = 'solid'
SEED = 42
```
# 1. First steps with unsupervised anomaly detection algorithms
The goal of this section is to get familiar with different unsupervised anomaly detection approaches and algorithms. In order to visualise the output of the different algorithms we consider a toy data set consisting in a two-dimensional Gaussian mixture.
### Generating the data set
```
from utils import GaussianMixture
n_samples = 500
n_features = 2
weight_1 = 0.5
weight_2 = 0.5
mean_1 = np.zeros(n_features)
mean_2 = -1 * np.ones(n_features)
cov_1 = np.array([[2., 2.,], [2., 4.]])
cov_2 = 2 * np.identity(n_features)
weights = np.array([weight_1, weight_2])
means = np.array([mean_1, mean_2])
covars = np.array([cov_1, cov_2])
gm = GaussianMixture(weights, means, covars, random_state=SEED)
X = gm.sample(n_samples)
```
### Plot the samples and levels set of the density
```
X_range = np.zeros((n_features, 2))
X_range[:, 0] = np.min(X, axis=0)
X_range[:, 1] = np.max(X, axis=0)
h = 0.1 # step size of the mesh
x_min, x_max = X_range[0, 0] - 0.1, X_range[0, 1] + 0.1
y_min, y_max = X_range[1, 0] - 0.1, X_range[1, 1] + 0.1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
grid = np.c_[xx.ravel(), yy.ravel()]
Z = gm.density(grid)
Z = Z.reshape(xx.shape)
plt.figure()
plt.contour(xx, yy, Z, 10, cmap=plt.cm.Blues_r)
plt.scatter(X[:, 0], X[:, 1], s=1.)
plt.show()
```
The goal is to estimate a Minimum volume set with mass at least 0.95. We know that under regularity assumption of the underlying distribution a Minimum volume set is a density level set of the density $f$.
$$
\{x, f(x) \geq F_f^{-1}(1-\alpha) \}
$$
where $F_f^{-1}(1-\alpha)$ is the quantile of order $1-\alpha$ of the distribution of the random variable $f(X)$.
1. Draw a sample $\mathcal{S}$ from the previous Gaussian mixture of size $n = 1,000,000$.
2. Using the sample $\mathcal{S}$ and the true density $f$ given by `gm.density` compute the empirical quantile of order $1-\alpha$ for $\alpha = 0.95$ and $\alpha = 0.99$. You can use for instance the `scipy.stats.mstats.mquantiles` function.
3. Plot the corresponding Minimum Volume set in a figure like the previous one, using `plt.contour` and replacing the number of levels (10) by the threshold you computed: `levels=threshold`.
4. Emphasize the outliers (for instance use a different color) in the previous plot where an outlier is a sample outside the Minimum Volume set.
```
alpha_set = 0.95
from scipy.stats.mstats import mquantiles
n_quantile = 1000000
Xq = gm.sample(n_quantile)
density_q = gm.density(Xq)
tau = mquantiles(density_q, 1 - alpha_set)
X_outliers = X[gm.density(X) < tau]
plt.figure()
c_0 = plt.contour(xx, yy, Z, levels=tau, colors='green', linewidths=3)
plt.clabel(c_0, inline=1, fontsize=15, fmt={tau[0]: str(alpha_set)})
plt.scatter(X[:, 0], X[:, 1], s=1.)
plt.scatter(X_outliers[:, 0], X_outliers[:, 1], color='red', s=4.)
plt.show()
```
## 1.1. Density estimation
We are going to use the plug-in approach to estimate the Minimum Volume set with mass at least 0.95. The plug-in approach means that we replace the density $f$ by an estimate $\hat f$.
1. Equation of the corresponding Minimum Volume set estimate.
2. Using a Gaussian kernel, compute a kernel density estimator of $f$. To stick with `sklearn` you can use `sklearn.neighbors.kde.KernelDensity` with the default bandwidth.
3. Compute the empirical quantile $\widehat F_{\hat f}^{-1}(1-\alpha)$ from the original sample for $\alpha = 0.95$ (still using the function `mquantiles`).
4. Add the corresponding Minimum Volume set estimate to the previous plot to visualize the difference between the true MV set and the estimated one.
```
from sklearn.neighbors.kde import KernelDensity
# Estimate density with a Gaussian kernel density estimator
kde = KernelDensity(kernel='gaussian')
kde = kde.fit(X)
kde_X = kde.score_samples(X)
tau_kde = mquantiles(kde_X, 1 - alpha_set)
Z_kde = kde.score_samples(grid)
Z_kde = Z_kde.reshape(xx.shape)
plt.figure()
c_0 = plt.contour(xx, yy, Z_kde, levels=tau_kde, colors='red', linewidths=3)
plt.clabel(c_0, inline=1, fontsize=15, fmt={tau_kde[0]: str(alpha_set)})
c_1 = plt.contour(xx, yy, Z, levels=tau, colors='green', linewidths=2, label='True')
plt.clabel(c_1, inline=1, fontsize=15, fmt={tau[0]: 'True'})
plt.scatter(X[:, 0], X[:, 1], s=1.)
plt.legend()
plt.show()
```
### Bandwidth selection with cross validation
We used the default bandwidth.
1. Use cross validation to learn the bandwidth from the data. You may use `sklearn.model_selection.GridSearchCV` to perform a 3-fold cross validation.
2. Plot the Minimum Volume set estimate obtained with the learnt bandwidth.
```
from sklearn.model_selection import GridSearchCV
grid_cv = GridSearchCV(KernelDensity(kernel='gaussian'), {'bandwidth': np.linspace(0.1, 2.0, 30)}, cv=3)
grid_cv.fit(X)
grid_cv.best_params_
kde_best = grid_cv.best_estimator_
kde_best_X = kde_best.score_samples(X)
tau_kde_best = mquantiles(kde_best_X, 1 - alpha_set)
Z_kde_best = kde_best.score_samples(grid)
Z_kde_best = Z_kde_best.reshape(xx.shape)
plt.figure()
c_0 = plt.contour(xx, yy, Z_kde_best, levels=tau_kde_best, colors='red', linewidths=3)
plt.clabel(c_0, inline=1, fontsize=15, fmt={tau_kde_best[0]: str(alpha_set)})
c_1 = plt.contour(xx, yy, Z, levels=tau, colors='green', linewidths=2, label='True')
plt.clabel(c_1, inline=1, fontsize=15, fmt={tau[0]: 'True'})
plt.scatter(X[:, 0], X[:, 1], s=1.)
plt.legend()
plt.show()
```
## 1.2. One-Class SVM
The One-Class SVM separates the data from the origin by solving the following optimization problem.
\begin{equation}
\min_{\boldsymbol{w},\boldsymbol{\xi},\rho} \quad \frac{1}{2}\Vert \boldsymbol{w} \Vert^2 - \rho + \frac{1}{\nu n}\sum_{i=1}^n\xi_i\\
\text{s.t.} \quad \langle \boldsymbol{w}, \Phi(x_i) \rangle \geqslant \rho - \xi_i, \quad \xi_i \geqslant 0 \quad \quad 1 \leqslant i \leqslant n
\end{equation}
where the parameter $\nu \in (0,1)$ has to be set by the user and controls the proportion of outliers.
It is in general solved via the dual given by
\begin{equation}
\min_{\boldsymbol{\alpha}} \quad \frac{1}{2}\sum_{i,j} k(x_i, x_j)\\
\text{s.t.} \quad 0 \leqslant \alpha_i \leqslant \frac{1}{\nu n},\quad \sum_{i}\alpha_i = 1 \quad \quad 1 \leqslant i \leqslant n
\end{equation}
1. Show that when using a Gaussian kernel, all data in the feature space lies on the same hypersphere. Show that data are always linearly separable from the origin. Hint: what's the formula of $k(x, x')$ in terms of the feature map $\Phi$.
```
from sklearn.svm import OneClassSVM
```
Under mild assumptions the following inequality holds true for all sample size $n$:
$$ \frac{\text{Outliers}}{n} \leq \nu \leq \frac{\text{Support Vectors}}{n} $$.
Futhermore the two fractions on the left-hand side and right-hand converge almost surely towards $\nu$.
1. If we want to estimate a Minimum Volume set with mass at least $\alpha$, how can we set $\nu$?
2. Use `sklearn.svm.OneClassSVM` with the default parameters (except the parameter $\nu$ set according to the previous question) to learn a Minimum Volume set with mass at least $0.95$.
3. Plot the Minimum Volume set estimate.
```
nu = 0.05
ocsvm = OneClassSVM(kernel='rbf', gamma=0.05, nu=nu)
#ocsvm = OneClassSVM(kernel='rbf', gamma=50., nu=nu)
#ocsvm = OneClassSVM(kernel='rbf', nu=nu)
ocsvm.fit(X)
X_outliers = X[ocsvm.predict(X) == -1]
Z_ocsvm = ocsvm.decision_function(grid)
Z_ocsvm = Z_ocsvm.reshape(xx.shape)
plt.figure()
c_0 = plt.contour(xx, yy, Z_ocsvm, levels=[0], colors='red', linewidths=3)
plt.clabel(c_0, inline=1, fontsize=15, fmt={0: str(alpha_set)})
c_1 = plt.contour(xx, yy, Z, levels=tau, colors='green', linewidths=2, label='True')
plt.clabel(c_1, inline=1, fontsize=15, fmt={tau[0]: 'True'})
plt.scatter(X[:, 0], X[:, 1], s=1.)
plt.scatter(X_outliers[:, 0], X_outliers[:, 1], s=4., color='red')
plt.legend()
plt.show()
```
1. What do you notice? The kernel used by the One-Class SVM is given by $$ k(x, x') = \exp(-\gamma\Vert x - x' \Vert^2). $$
2. What is the default gamma used when we trained the One-Class SVM? Do we need to increase or decrease its value?
```
ocsvm?
```
### Support vectors - Outliers
1. Check that we indeed have $$ \frac{\text{Outliers}}{n} \leq \nu \leq \frac{\text{Support Vectors}}{n} $$.
2. Use `gamma=10`. Is the inequality always satisfied? Can you guess why?
```
X_SV = X[ocsvm.support_]
n_SV = len(X_SV)
n_outliers = len(X_outliers)
print('{0:.2f} <= {1:.2f} <= {2:.2f}?'.format(1./n_samples*n_outliers, nu, 1./n_samples*n_SV))
```
Only the support vectors are involved in the decision function of the One-Class SVM.
1. Plot the level sets of the One-Class SVM decision function as we dit for the true density.
2. Emphasize the Support vectors.
```
plt.figure()
plt.contourf(xx, yy, Z_ocsvm, 10, cmap=plt.cm.Blues_r)
plt.scatter(X[:, 0], X[:, 1], s=1.)
plt.scatter(X_SV[:, 0], X_SV[:, 1], s=20., color='orange')
plt.show()
```
1. Concerning the support vectors, what is the main difference between the OneClass SVM and a Kernel Density estimator when using a Gaussian Kernel?
2. When is it particularly advantageous to use the OneClass SVM?
### What if we change the value of $\nu$?
1. Set $\nu = 0.4$. How can we estimate a Minimum Volume set with mass at least $0.95$. Hint: Use the `decision_function`.
```
nu = 0.4
ocsvm = OneClassSVM(kernel='rbf', gamma=0.5, nu=nu)
ocsvm.fit(X)
Z_ocsvm = ocsvm.decision_function(grid)
Z_ocsvm = Z_ocsvm.reshape(xx.shape)
ocsvm_X = ocsvm.decision_function(X)
tau_ocsvm = mquantiles(ocsvm_X, 1 - alpha_set)
plt.figure()
c_0 = plt.contour(xx, yy, Z_ocsvm, levels=tau_ocsvm, colors='red', linewidths=3)
plt.clabel(c_0, inline=1, fontsize=15, fmt={tau_ocsvm[0]: str(alpha_set)})
c_1 = plt.contour(xx, yy, Z, levels=tau, colors='green', linewidths=2, label='True')
plt.clabel(c_1, inline=1, fontsize=15, fmt={tau[0]: 'True'})
plt.scatter(X[:, 0], X[:, 1], s=1.)
plt.legend()
plt.show()
```
### Equivalence with SVDD
Support Vector Data Description consists in finding the ball of minimum volume containing an arbitrary proportion of data. This is obtained by minimizing the following optimization problem.
\begin{equation}
\min_{R, \boldsymbol{\xi},\boldsymbol{c}} \quad R^2 + \frac{1}{\nu n}\sum_{i=1}^{n}\xi_i\\
\text{s.t.} \quad \Vert \boldsymbol{x}_i - \boldsymbol{c}\Vert^2 \leqslant R^2 + \xi_i, \quad \xi_i \geqslant 0 \quad \quad 1\leqslant i \leqslant n\\
\end{equation}
It is in general solved via the dual given by
\begin{equation}
\min_{\boldsymbol{\alpha}} \quad \sum_{i,j} k(x_i, x_j) + \sum_{i}\alpha_i k(x_i,x_i)\\
\text{s.t.} \quad 0 \leqslant \alpha_i \leqslant \frac{1}{\nu n},\quad \sum_{i}\alpha_i = 1 \quad \quad 1 \leqslant i \leqslant n
\end{equation}
1. When are SVDD and the One-Class SVM equivalent?
## 1.3. Isolation Forest
1. Run Isolation with `n_estimators=1` , i.e., only one tree will be built.
2. Plot the contour of the set with the $95\%$ most normal observations. Hint: use the `decision_function` and the `contamination` parameter. Compare it with the true Minimum Volume set with mass at least $0.95$.
3. Increase `n_estimators` to 50, 100 and then 500.
4. What does the `max_sample` parameter change?
```
from sklearn.ensemble import IsolationForest
# iforest = IsolationForest(n_estimators=1, contamination=0.05)
iforest = IsolationForest(n_estimators=100, contamination=0.05)
iforest = iforest.fit(X)
Z_iforest = iforest.decision_function(grid)
Z_iforest = Z_iforest.reshape(xx.shape)
plt.figure()
c_0 = plt.contour(xx, yy, Z_iforest, levels=[iforest.threshold_], colors='red', linewidths=3)
plt.clabel(c_0, inline=1, fontsize=15, fmt={iforest.threshold_: str(alpha_set)})
c_1 = plt.contour(xx, yy, Z, levels=tau, colors='green', linewidths=2, label='True')
plt.clabel(c_1, inline=1, fontsize=15, fmt={tau[0]: 'True'})
plt.scatter(X[:, 0], X[:, 1], s=1.)
plt.legend()
plt.show()
```
## 1.4. Local Outlier Factor
k-NN based anomaly detection algorithm. Available on the dev version of Scikit-Learn.
## 1.5. Unsupervised as supervised using SVM
Let $P$ be a distribution with finite support $C$. Let $\mu$ be the uniform distribution over the support $C$: $\mu(A) = \lambda(A)/\lambda(C)$. We now consider the joint distribution $Q$ of $(X, Y) \in \mathbb{R}^d \times \{-1, +1\}$ given by the class conditionals distributions:
$$
X \vert Y= +1 \sim P \quad \text{and} \quad X \vert Y=-1 \sim \mu
$$
and a priori class probabilities $p = \mathbb{P}(Y=1)$.
We are as before interested in finding a given density level set $\{ f \geq \tau \}$ where $f$ is the density of $P$ with respect to the Lebesgue measure $\lambda$.
The marginal distribution of $X$, $P_X$, is given by
$$
P_X = p P + (1-p) Q.
$$
Assuming that $P$ has a density $f$. It can be shown that
$$\eta(x) = P(Y=1\vert X=x) = \frac{pf(x)}{pf(x) + (1-p)/\lambda(C)}$$
The optimal classification set is given by $\{ \eta \geq 1/2 \}$
1. Show that if $p=1/(1 + \lambda(C)\tau)$ then $\lambda(\{\eta \geq 1/2 \} \Delta \{f \geq \tau \} ) = 0$
Thus solving a binary classification problem assigning weights $p$ and $1-p$ to the samples can lead to a density level set.
### Generate uniform data in the range of X
```
### Generating artificial second class
rng = np.random.RandomState(SEED)
U = np.zeros((n_samples, n_features))
for i in range(n_features):
U [:, i] = rng.uniform(X_range[i, 0], X_range[i, 1], n_samples)
vol = np.prod(X_range[:, 1] - X_range[:, 0])
X_bin = np.r_[X, U]
y_bin = np.r_[np.ones(n_samples), np.zeros(n_samples)].astype(int)
from sklearn.svm import SVC
# SVM parameters
C = 1.
lambda_reg = 0.00005
gamma = 0.01
p = 1. / (1. + vol * tau[0])
weight_1 = p * 1. / (lambda_reg * n_samples)
weight_0 = (1 - p) * 1. / (lambda_reg * n_samples)
class_weight = {1: weight_1, 0: weight_0}
clf = SVC(C=C, kernel='rbf', gamma=gamma, class_weight=class_weight)
clf = clf.fit(X_bin, y_bin)
Z_svm = clf.decision_function(grid)
Z_svm = Z_svm.reshape(xx.shape)
plt.figure()
c_0 = plt.contour(xx, yy, Z_svm, levels=[0], colors='red', linewidths=3)
plt.clabel(c_0, inline=1, fontsize=15, fmt={0: str(alpha_set)})
c_1 = plt.contour(xx, yy, Z, levels=tau, colors='green', linewidths=2, label='True')
plt.clabel(c_1, inline=1, fontsize=15, fmt={tau[0]: 'True'})
plt.scatter(X[:, 0], X[:, 1], s=1.)
plt.legend()
plt.show()
```
# 2. Anomaly scoring and Mass Volume curve
Usually unsupervised anomaly detection algorithms returns a scoring function $s$ such that the smaller $s(x)$ the more abnormal $x$ is. To assess the performance of such a scoring function we can use the Mass Volume curve. The MV curve of a scoring function $s$ is given for all $\alpha \in (0,1)$ by
$$
MV_s(\alpha) = \lambda\{x, s(x) \geq F_s^{-1}(1-\alpha) \}
$$
where $F_s^{-1}(1-\alpha)$ is the quantile of order $1-\alpha$ of the distribution of the random variable $s(X)$.
Under regularity assumptions, the MV curve is also given by the parametric curve
$$
(\mathbb{P}(s(X) \geq t), \lambda(x, s(x) \geq t))
$$
Assume that the random variable $X$ with distribution $P$ has a density $f$ with respect to the Lebesgue measure. Furthermore assume that for all $\alpha \in (0,1)$ the Minimum Volume set optimization problem
$$
\min_{\Omega \in \mathcal{M}} \lambda(\Omega) \quad \text{ s.t } \quad P(\Omega) \geq \alpha .
$$
has a unique solution given by
$$
\Omega^*_{\alpha} = \{x, f(x) \geq F_f^{-1}(1-\alpha) \}.
$$
1. Show that for all $\alpha \in (0,1)$, $MV_s(\alpha) - MV_f(\alpha) \leq \lambda(\Omega^*_{\alpha} \Delta \{x, s(x) \geq F_s^{-1}(1-\alpha) \})$.
2. Show that for all $\alpha \in (0,1)$, $MV_f(\alpha) \leq MV_s(\alpha)$. Hint: use the following property: for all $\alpha \in (0,1)$, $\mathbb{P}(s(X) < F_s^{-1}(1-\alpha)) \leq 1-\alpha$.
3. In what sense a scoring function minimizing the area under the Mass Volume curve is optimal?
## 2.1. Using Mass Volume curve for performance evaluation
The lower $MV_s$ the better the scoring function $s$. The goal is to compare the performance of scoring functions given by the One-Class SVM for different values of $\gamma$ using the Mass Volume curve. To easily plot the Mass Volume we are going to use the ROC curve function given by `sklearn`. The idea is to use a uniform sample as the positive class and to consider the Gaussian mixture data set as the negative class. To plot the MV curve we want to plot the volume against the mass $\alpha$. This will be given by $1-TPR$ against $1-FPR$.
1. Using `sklearn.model_selection.ShuffleSplit` split the data set into a training set and a test set.
2. Generate `n_sim=100000` uniform random variables in the hypercube enclosing the data using `numpy.random.uniform`.
3. Build a training set consisting in the concatenation of the training set of the Gaussian mixture data set (obtained in 1.) and the uniform sample.
4. Similarly, build the test set by concatenating the test set of the Gaussian mixture data set (obtained in 1.) and the uniform sample.
```
from sklearn.model_selection import ShuffleSplit
X_range = np.zeros((n_features, 2))
X_range[:, 0] = np.min(X, axis=0)
X_range[:, 1] = np.max(X, axis=0)
n_sim = 100000
U = np.zeros((n_sim, n_features))
for i in range(n_features):
U [:, i] = rng.uniform(X_range[i, 0], X_range[i, 1], n_sim)
### Training on half the data
ss = ShuffleSplit(n_splits=1, test_size=0.5, random_state=SEED)
ss = ss.split(X)
train, test = ss.next()
X_train = X[train]
X_test = X[test]
X_train_all = np.r_[U, X_train]
y_train_all = np.r_[np.zeros(n_sim), np.ones(len(train))]
X_test_all = np.r_[U, X_test]
y_test_all = np.r_[np.zeros(n_sim), np.ones(len(test))]
```
1. Compute the Mass Volume curve, estimated on the training set, of the scoring function obtained with the One-Class SVM for $\gamma=0.05$ and $\gamma=10$. Compute the area under the Mass Volume curve for each value of $\gamma$.
2. Compute the Mass Volume curve estimated on the test set for each value of $\gamma$. Compute the area under the Mass Volume curve for each value of $\gamma$.
```
from sklearn.metrics import roc_curve, auc
colors = ['red', 'blue']
plt.figure(figsize=(12, 6))
for g, gamma in enumerate([0.05, 10]):
ocsvm = OneClassSVM(gamma=gamma, nu=0.05)
### Train algorithm (fit), compute decision function on train and test, compute ROC and AUC, draw ROC
ocsvm.fit(X_train)
ocsvm_train_all = ocsvm.decision_function(X_train_all)
ocsvm_test_all = ocsvm.decision_function(X_test_all)
fpr_train_, tpr_train_, _ = roc_curve(y_train_all, -ocsvm_train_all, pos_label=0)
ocsvm_auc_train = auc(1 - fpr_train_, 1 - tpr_train_)
fpr_test_, tpr_test_, _ = roc_curve(y_test_all, -ocsvm_test_all, pos_label=0)
ocsvm_auc_test = auc(1 - fpr_test_, 1 - tpr_test_)
plt.subplot(1, 2, 1)
plt.title('Performance on training set')
plt.plot(1 - fpr_train_, 1 - tpr_train_, color=colors[g], label= 'Gamma: {0:.2f} - AMV: {1:.3f}'.format(gamma, ocsvm_auc_train))
plt.subplot(1, 2, 2)
plt.title('Performance on test set')
plt.plot(1 - fpr_test_, 1 - tpr_test_, color=colors[g], label= 'Gamma: {0:.2f} - AMV: {1:.3f}'.format(gamma, ocsvm_auc_test))
plt.subplot(1, 2, 1)
plt.legend(loc=0)
plt.xlim((-0.05, 1.05))
plt.ylim((-0.05, 1.05))
plt.subplot(1, 2, 2)
plt.legend(loc=0)
plt.xlim((-0.05, 1.05))
plt.ylim((-0.05, 1.05))
plt.show()
```
# 3. Performance evaluation on real data sets
## 3.1. Shuttle data set
```
from sklearn.datasets import fetch_mldata
dataset = fetch_mldata('shuttle')
X = dataset.data
y = dataset.target
### Usual setup when using this data set for anomaly detection
# Instances with label 4 are removed
# Normal data: label 1
ind = (y != 4)
X = X[ind, :]
y = y[ind]
y = (y == 1).astype(int)
n_samples, n_features = X.shape
n_samples, n_features
```
1. What's the proportion of anomalies in the data set (label 0)?
2. Split the data set into a training a test set using `StratifiedShuffleSplit` to preserve the proportion of each class in the training and test sets.
3. Compare the performance of the OneClass SVM and Isolation Forest using the ROC curve (computed with the true labels) on the train set and the set set:
- train the algorithm on the training set (without the labels), compute the ROC curve on the training set using the labels.
- train the algorithm on the training set (without the labels), compute the ROC curve on the test set using the labels
4. Compare the performance of the OneClass SVM and Isolation Forest using the Mass Volume curve on the training and test sets
```
'Percentage of anomalies: {0}'.format(1 - np.mean(y))
from sklearn.model_selection import StratifiedShuffleSplit
sss = StratifiedShuffleSplit(n_splits=1, test_size=0.5, random_state=SEED)
train, test = sss.split(X, y).next()
X_train, y_train = X[train], y[train]
X_test, y_test = X[test], y[test]
print('Percentage of anomalies in train : {0}'.format(1 - np.mean(y_train)))
print('Percentage of anomalies in test : {0}'.format(1 - np.mean(y_test)))
```
### Performance on training set and test set - ROC curve - AUC
```
### !!!! This can take a few minutes. You might consider running it with Isolation Forest only at first
algorithms = [OneClassSVM(), IsolationForest()]
name = ['Isolation Forest', 'Ocsvm']
colors = ['green', 'red']
plt.figure(figsize=(12, 6))
for a, algo in enumerate(algorithms):
algo.fit(X_train)
algo_train = algo.decision_function(X_train)
algo_test = algo.decision_function(X_test)
fpr_train_, tpr_train_, _ = roc_curve(y_train, -algo_train, pos_label=0)
algo_auc_train = auc(fpr_train_, tpr_train_)
fpr_test_, tpr_test_, _ = roc_curve(y_test, -algo_test, pos_label=0)
algo_auc_test = auc(fpr_test_, tpr_test_)
plt.subplot(1, 2, 1)
plt.title('Performance on training set')
plt.plot(fpr_train_, tpr_train_, color=colors[a], label= '{0} - AUC: {1:.3f}'.format(name[a], algo_auc_train))
plt.subplot(1, 2, 2)
plt.title('Performance on test set')
plt.plot(fpr_test_, tpr_test_, color=colors[a], label= '{0} - AUC: {1:.3f}'.format(name[a], algo_auc_test))
plt.subplot(1, 2, 1)
plt.legend(loc=0)
plt.xlim((-0.05, 1.05))
plt.ylim((-0.05, 1.05))
plt.subplot(1, 2, 2)
plt.legend(loc=0)
plt.xlim((-0.05, 1.05))
plt.ylim((-0.05, 1.05))
plt.show()
```
### Performance - MV curve - AMV
```
X_range = np.zeros((n_features, 2))
X_range[:, 0] = np.min(X, axis=0)
X_range[:, 1] = np.max(X, axis=0)
n_sim = 100000
U = np.zeros((n_sim, n_features))
for i in range(n_features):
U [:, i] = rng.uniform(X_range[i, 0], X_range[i, 1], n_sim)
### We add uniform data to compute volumes
X_train_all = np.r_[U, X_train]
X_test_all = np.r_[U, X_test]
# Assigning same label to all data (normal and anomalies): the anomalies are the uniform samples
y_train_all = np.r_[np.zeros(n_sim), np.ones(len(X_train))]
y_test_all = np.r_[np.zeros(n_sim), np.ones(len(X_test))]
### !!!! This can take a few minutes. You might consider running it with Isolation Forest only at first
algorithms = [OneClassSVM(), IsolationForest()]
name = ['Isolation Forest', 'Ocsvm']
colors = ['green', 'red']
# algorithms = [IsolationForest()]
# name = ['Isolation Forest']
# colors = ['green']
plt.figure(figsize=(12, 6))
for a, algo in enumerate(algorithms):
### Train algorithm (fit), compute decision function on train and test, compute ROC and AUC, draw ROC
algo.fit(X_train)
algo_train_all = algo.decision_function(X_train_all)
algo_test_all = algo.decision_function(X_test_all)
fpr_train_, tpr_train_, _ = roc_curve(y_train_all, -algo_train_all, pos_label=0)
algo_auc_train = auc(1 - fpr_train_, 1 - tpr_train_)
fpr_test_, tpr_test_, _ = roc_curve(y_test_all, -algo_test_all, pos_label=0)
algo_auc_test = auc(1 - fpr_test_, 1 - tpr_test_)
plt.subplot(1, 2, 1)
plt.title('Performance on training set')
plt.plot(1 - fpr_train_, 1 - tpr_train_, color=colors[a], label= '{0} - AMV: {1:.3f}'.format(name[a], algo_auc_train))
plt.subplot(1, 2, 2)
plt.title('Performance on test set')
plt.plot(1 - fpr_test_, 1 - tpr_test_, color=colors[a], label= '{0} - AUC: {1:.3f}'.format(name[a], algo_auc_test))
plt.subplot(1, 2, 1)
plt.legend(loc=0)
plt.xlim((-0.05, 1.05))
plt.ylim((-0.05, 1.05))
plt.subplot(1, 2, 2)
plt.legend(loc=0)
plt.xlim((-0.05, 1.05))
plt.ylim((-0.05, 1.05))
plt.show()
```
## 3.2. Http data set - Network intrusion detection
1. Same questions with the http data set
```
from sklearn.datasets import fetch_kddcup99
dataset = fetch_kddcup99(subset='http', shuffle=True,
percent10=True, random_state=SEED)
X = dataset['data']
X = X.astype('float')
y = dataset['target']
y = (y == 'normal.').astype(int)
n_samples, n_features = X.shape
```
### Novelty detection
Now we only train the model on normal data. For instance let's train the model on half the normal data.
1. Split the data set into a set with the normal data and a set with the anomalies.
2. Randomly split the normal data set into a two data sets: a training set and a test set.
3. Build a test set from the normal test set and the anomalies.
4. Train the OneClass SVM and Isolation Forest on the normal training set and compute the ROC curve on the test set.
```
X_normal = X[y == 1]
X_ano = X[y != 1]
y_normal = y[y == 1]
y_ano = y[y != 1]
from sklearn.model_selection import ShuffleSplit
ss = ShuffleSplit(n_splits=1, test_size=0.5, random_state=SEED)
ss = ss.split(X_normal, y_normal)
train_normal, test_normal = ss.next()
X_train = X_normal[train_normal]
X_test = np.r_[X_normal[test_normal], X_ano]
y_test = np.r_[y_normal[test_normal], y_ano]
```
#### ROC - AUC on test set
```
### !!!! This can take a few minutes. You might consider running it with Isolation Forest only at first
algorithms = [OneClassSVM(), IsolationForest()]
name = ['Isolation Forest', 'Ocsvm']
colors = ['green', 'red']
# algorithms = [IsolationForest()]
# name = ['Isolation Forest']
# colors = ['green']
plt.figure(figsize=(12, 6))
for a, algo in enumerate(algorithms):
### Train algorithm (fit), compute decision function on train and test, compute ROC and AUC, draw ROC
algo.fit(X_train)
algo_test = algo.decision_function(X_test)
fpr_test_, tpr_test_, _ = roc_curve(y_test, -algo_test, pos_label=0)
algo_auc_test = auc(fpr_test_, tpr_test_)
plt.subplot(1, 2, 2)
plt.title('Performance on test set')
plt.plot(fpr_test_, tpr_test_, color=colors[a], label= '{0} - AUC: {1:.3f}'.format(name[a], algo_auc_test))
plt.xlim((-0.05, 1.05))
plt.ylim((-0.05, 1.05))
plt.legend(loc=0)
plt.show()
```
#### MV curve on test set
1. Plot the MV curve estimated on the test set for the OneClass SVM and Isolation Forest.
```
X_range = np.zeros((n_features, 2))
X_range[:, 0] = np.min(X, axis=0)
X_range[:, 1] = np.max(X, axis=0)
n_sim = 100000
U = np.zeros((n_sim, n_features))
for i in range(n_features):
U [:, i] = rng.uniform(X_range[i, 0], X_range[i, 1], n_sim)
### We add uniform data to compute volumes
X_test_all = np.r_[U, X_test]
# Assigning same label to all data (normal and anomalies): the anomalies are the uniform samples
y_test_all = np.r_[np.zeros(n_sim), np.ones(len(X_test))]
### !!!! This can take a few minutes. You might consider running it with Isolation Forest only at first
algorithms = [OneClassSVM(), IsolationForest()]
name = ['Isolation Forest', 'Ocsvm']
colors = ['green', 'red']
# algorithms = [IsolationForest()]
# name = ['Isolation Forest']
# colors = ['green']
plt.figure(figsize=(12, 6))
for a, algo in enumerate(algorithms):
### Train algorithm (fit), compute decision function on train and test, compute ROC and AUC, draw ROC
algo.fit(X_train)
algo_test_all = algo.decision_function(X_test_all)
fpr_test_all_, tpr_test_all_, _ = roc_curve(y_test_all, -algo_test_all, pos_label=0)
algo_auc_test = auc(1 - fpr_test_all_, 1 - tpr_test_all_)
plt.subplot(1, 2, 2)
plt.title('Performance on test set')
plt.plot(1 - fpr_test_all_, 1 - tpr_test_all_, color=colors[a], label= '{0} - AUC: {1:.3f}'.format(name[a], algo_auc_test))
plt.xlim((-0.05, 1.05))
plt.ylim((-0.05, 1.05))
plt.legend(loc=0)
plt.show()
```
# 4. Optional
1. Show that a density level set $\{f \geq \tau \}$ is a Minimum volume set with mass $\alpha = \mathbb{P}(f(X) \geq \tau)$.
2. Show that if the density has not flat parts, the minimum volume set optimization problem has a solution.
# 5. Overfitting for Minimum volume set estimation
Let $\lambda$ denotes the Lebesgue measure of $\mathbb{R}^d$ and let $\mathcal{M}$ be the class of all measurable sets. A Minimum Volume set with mass at least $\alpha$ is the solution of the following optimization problem:
$$
\min_{\Omega \in \mathcal{M}} \lambda(\Omega) \quad \text{ s.t } \quad P(\Omega) \geq \alpha .
$$
We do not know the distribution $P$ but only have a sample $X_1, \dots, X_n$ drawn from this distribution.
1. What is the solution for all $\alpha \in (0,1)$ of the empirical optimization problem if we solve the following empirical version of the Minimum Volume set optimization problem:
$$
\min_{\Omega \in \mathcal{M}} \lambda(\Omega) \quad \text{ s.t } \quad \widehat P(\Omega) \geq \alpha
$$
where for all $\Omega$, $\widehat P(\Omega) = \frac{1}{n}\sum_{i=1}^n \mathbb{I}\{X_i \in \Omega \}$.
# 6. Illustration of IsolationForest on Digits data set
```
from sklearn import datasets
digits = datasets.load_digits()
```
The digits data set consits in images (8 x 8) of digits.
```
images = digits.images
labels = digits.target
images.shape
i = 0
plt.figure()
plt.title('{0}'.format(labels[i]))
plt.axis('off')
plt.imshow(images[i], cmap=plt.cm.gray_r, interpolation='nearest')
plt.show()
```
To use the images as a training set we need to flatten the images.
```
n_samples = len(digits.images)
data = digits.images.reshape((n_samples, -1))
data.shape
X = data
y = digits.target
```
We are going to focus on digit 5.
```
X_5 = X[y == 5]
```
1. Use IsolationForest to find the top 5% most abnormal images.
2. Plot them using `plt.imshow(outlier.reshape(8, 8)), cmap=plt.cm.gray_r, interpolation='nearest')`
```
from sklearn.ensemble import IsolationForest
iforest = IsolationForest(contamination=0.05)
iforest = iforest.fit(X_5)
iforest_X = iforest.decision_function(X_5)
X_outliers = X_5[iforest.predict(X_5)==-1]
for i in range(len(X_outliers)):
plt.subplot(2, int(len(X_outliers) / 2.) + 1, i + 1)
plt.axis('off')
plt.imshow(X_outliers[i].reshape((8, 8)), cmap=plt.cm.gray_r, interpolation='nearest')
plt.show()
```
| github_jupyter |
In this notebook, we show how we can train a model with Scikit-learn and save it as a TileDB array on TileDB-Cloud.
Firstly, let's import what we need.
```
import numpy as np
import tiledb.cloud
import os
from sklearn import preprocessing
from sklearn.linear_model import LogisticRegression
from tiledb.ml.models.sklearn import SklearnTileDBModel
```
We then have to export and load our TileDB-Cloud credentials. For TileDB cloud you can also use a token.
You have to also set up your AWS credentials on your TileDB-Cloud account.
```
# This is also our namespace on TileDB-Cloud.
TILEDB_USER_NAME = os.environ.get('TILEDB_USER_NAME')
TILEDB_PASSWD = os.environ.get('TILEDB_PASSWD')
```
We then create a TileDB-Cloud context and set up our communication with TileDB-Cloud.
```
ctx = tiledb.cloud.Ctx()
tiledb.cloud.login(username=TILEDB_USER_NAME, password=TILEDB_PASSWD)
```
And move on with training a sklearn model with some random data.
```
X_train = np.random.random((1000, 784))
y_train = np.random.randint(9, size=1000)
X_test = np.random.random((500, 784))
y_test = np.random.randint(9, size=500)
scaler = preprocessing.StandardScaler().fit(X_train)
scaled_X_train = scaler.transform(X_train)
scaled_X_test = scaler.transform(X_test)
print("Model fit...")
model = LogisticRegression(random_state=0).fit(scaled_X_train, y_train)
print("Model score...")
sparsity = np.mean(model.coef_ == 0) * 100
score = model.score(scaled_X_test, y_test)
print("Sparsity with L1 penalty: %.2f%%" % sparsity)
print("Test score with L1 penalty: %.4f" % score)
```
We can move on by defining a TileDB Sklearn model and use model save functionality in order to save it directly to
our bucket on S3 (defined with AWS credentials in your TileDB-Cloud account) and register it on TileDB-Cloud.
```
# Define array model uri.
uri = "tiledb-sklearn-model"
print('Defining SklearnTileDBModel model...')
# In order to save our model on S3 and register it on TileDB-Cloud we have to pass our Namespace and TileDB Context.
tiledb_model = SklearnTileDBModel(uri=uri, namespace=TILEDB_USER_NAME, ctx=ctx, model=model)
print(tiledb_model.uri)
# We will need the uri that was created from our model class
# (and follows pattern tiledb://my_username/s3://my_bucket/my_array),
# in order to interact with our model on TileDB-Cloud.
tiledb_cloud_model_uri = tiledb_model.uri
print('Saving model on S3 and registering on TileDB-Cloud...')
tiledb_model.save(meta={"Sparsity_with_L1_penalty": sparsity, "score": score})
```
Finally, we can use TileDB-Cloud API as described in our [cloud documentation](https://docs.tiledb.com/cloud/), in order
to list our models, get information and deregister them.
```
# List all our models. Here, we filter with file_type = 'ml_model'. All machine learning model TileDB arrays are of type
# 'ml_model'
print(
tiledb.cloud.client.list_arrays(
file_type=['ml_model'],
namespace=TILEDB_USER_NAME))
# Get model's info
print(tiledb.cloud.array.info(tiledb_cloud_model_uri))
# Load our model for inference
loaded_tiledb_model = SklearnTileDBModel(uri=tiledb_cloud_model_uri, ctx=ctx).load()
print(score == loaded_tiledb_model.score(X_test, y_test))
# Deregister model
tiledb.cloud.deregister_array(tiledb_cloud_model_uri)
```
| github_jupyter |
# Riskfolio-Lib Tutorial:
<br>__[Financionerioncios](https://financioneroncios.wordpress.com)__
<br>__[Orenji](https://www.orenj-i.net)__
<br>__[Riskfolio-Lib](https://riskfolio-lib.readthedocs.io/en/latest/)__
<br>__[Dany Cajas](https://www.linkedin.com/in/dany-cajas/)__
<a href='https://ko-fi.com/B0B833SXD' target='_blank'><img height='36' style='border:0px;height:36px;' src='https://cdn.ko-fi.com/cdn/kofi1.png?v=2' border='0' alt='Buy Me a Coffee at ko-fi.com' /></a>
## Tutorial 14: Mean [Ulcer Index](https://en.wikipedia.org/wiki/Ulcer_index) Portfolio Optimization
## 1. Downloading the data:
```
import numpy as np
import pandas as pd
import yfinance as yf
import warnings
warnings.filterwarnings("ignore")
yf.pdr_override()
pd.options.display.float_format = '{:.4%}'.format
# Date range
start = '2016-01-01'
end = '2019-12-30'
# Tickers of assets
assets = ['JCI', 'TGT', 'CMCSA', 'CPB', 'MO', 'APA', 'MMC', 'JPM',
'ZION', 'PSA', 'BAX', 'BMY', 'LUV', 'PCAR', 'TXT', 'TMO',
'DE', 'MSFT', 'HPQ', 'SEE', 'VZ', 'CNP', 'NI', 'T', 'BA']
assets.sort()
# Downloading data
data = yf.download(assets, start = start, end = end)
data = data.loc[:,('Adj Close', slice(None))]
data.columns = assets
# Calculating returns
Y = data[assets].pct_change().dropna()
display(Y.head())
```
## 2. Estimating Mean Ulcer Index Portfolios
### 2.1 Calculating the portfolio that maximizes Ulcer Performance Index (UPI) ratio.
```
import riskfolio.Portfolio as pf
# Building the portfolio object
port = pf.Portfolio(returns=Y)
# Calculating optimum portfolio
# Select method and estimate input parameters:
method_mu='hist' # Method to estimate expected returns based on historical data.
method_cov='hist' # Method to estimate covariance matrix based on historical data.
port.assets_stats(method_mu=method_mu, method_cov=method_cov, d=0.94)
# Estimate optimal portfolio:
model='Classic' # Could be Classic (historical), BL (Black Litterman) or FM (Factor Model)
rm = 'UCI' # Risk measure used, this time will be variance
obj = 'Sharpe' # Objective function, could be MinRisk, MaxRet, Utility or Sharpe
hist = True # Use historical scenarios for risk measures that depend on scenarios
rf = 0 # Risk free rate
l = 0 # Risk aversion factor, only useful when obj is 'Utility'
w = port.optimization(model=model, rm=rm, obj=obj, rf=rf, l=l, hist=hist)
display(w.T)
```
### 2.2 Plotting portfolio composition
```
import riskfolio.PlotFunctions as plf
# Plotting the composition of the portfolio
ax = plf.plot_pie(w=w, title='Sharpe Mean Ulcer Index', others=0.05, nrow=25, cmap = "tab20",
height=6, width=10, ax=None)
```
### 2.3 Calculate efficient frontier
```
points = 40 # Number of points of the frontier
frontier = port.efficient_frontier(model=model, rm=rm, points=points, rf=rf, hist=hist)
display(frontier.T.head())
# Plotting the efficient frontier
label = 'Max Risk Adjusted Return Portfolio' # Title of point
mu = port.mu # Expected returns
cov = port.cov # Covariance matrix
returns = port.returns # Returns of the assets
ax = plf.plot_frontier(w_frontier=frontier, mu=mu, cov=cov, returns=returns, rm=rm,
rf=rf, alpha=0.05, cmap='viridis', w=w, label=label,
marker='*', s=16, c='r', height=6, width=10, ax=None)
# Plotting efficient frontier composition
ax = plf.plot_frontier_area(w_frontier=frontier, cmap="tab20", height=6, width=10, ax=None)
```
## 3. Estimating Risk Parity Portfolios for Ulcer Index
### 3.1 Calculating the risk parity portfolio for Ulcer Index.
```
b = None # Risk contribution constraints vector
w_rp = port.rp_optimization(model=model, rm=rm, rf=rf, b=b, hist=hist)
display(w.T)
```
### 3.2 Plotting portfolio composition
```
ax = plf.plot_pie(w=w_rp, title='Risk Parity Ulcer Index', others=0.05, nrow=25, cmap = "tab20",
height=6, width=10, ax=None)
```
### 3.3 Plotting Risk Composition
```
ax = plf.plot_risk_con(w_rp, cov=port.cov, returns=port.returns, rm=rm, rf=0, alpha=0.01,
color="tab:blue", height=6, width=10, ax=None)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/neurorishika/PSST/blob/master/Tutorial/Day%203%20Cells%20in%20Silicon/Day%203.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a> <a href="https://kaggle.com/kernels/welcome?src=https://raw.githubusercontent.com/neurorishika/PSST/master/Tutorial/Day%203%20Cells%20in%20Silicon/Day%203.ipynb" target="_parent"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" alt="Open in Kaggle"/></a>
## Day 3: Cells in Silicon
Welcome to Day 3! Today, we start with our discussion of Hodgkin Huxley Neurons and how we can simulate them in Python using Tensorflow and Numerical Integration.
The electric potential measured across the membranes of excitable cells, such as neurons or heart cells, can undergo transient changes when perturbed by external inputs. When the inputs to a neuron are sufficiently large these transient changes can regeneratively build up into a large deviation from the resting state known as an action potential. Action potentials propagate undiminished along the axon and perturb post-synaptic neurons. The Hodgkin-Huxley model is a system of differential equations that describe the generation an action potential and its propagation along the axon. We provide only a brief overview of the Hodgkin-Huxley model. A number of classic references (Dayan 2005, Johnston 1995) and the original papers by Hodgkin and Huxley (Huxley 1952) chronicle the history and the details of the model. An excellent set of MOOCS and the accompanying textbooks (Gerstner 2014, Dayan 2005) give an accessible introduction to the topic
### What is the Hodgkin Huxley Neuron Model?
The cell membrane, a 5nm thick lipid bilayer, separates the inside from the outside of the neuron. The membrane is largely impermeable to charged ions present on either side. The concentration of $\text{Na}^{+}$ ions outside the cell is greater than its concentration inside, while $\text{K}^{+}$ ions are relatively abundant inside compared to the outside. In addition to these there are chloride ($\text{Cl}^{-}$), calcium ($\text{Ca}^{2+}$) and magnesium ions ($\text{Mg}^{+}$) that populate the cellular milieu. The differences in ionic abundances across the membrane cause a net accumulation of positive ions on one side of the membrane and negative ions on the other, and thus a potential difference across the membrane. Embedded on the membrane are ion channels that are highly selective to the ion species it lets across. In the squid axon, Hodgkin and Huxley found that there were only two types of ion channels ($\text{Na}^{+}$ and $\text{K}^{+}$), in addition to a non-specific leak channel. The Hodgkin-Huxley model of neurons can be understood with the help of an equivalent electrical circuit given below. The cell membrane acts as a capacitor. The total injected current ($I$) can be written as the sum of the capacitive current $I_{C}$, ionic currents $I_{Na}$ and $I_{K}$ and the leak current $I_L$.
<img src="https://raw.githubusercontent.com/neurorishika/PSST/master/Tutorial/Day%203%20Cells%20in%20Silicon/circuit.svg" width="800"/>
\begin{equation}
I = I_{C}(t) + I_{Na}(t) + I_{K}(t)
\end{equation}
where,
\begin{eqnarray}
C_m = 1 \mu F/cm^2 \\
I_{Na} = g_{Na}(u-E_{Na})\\
I_{K} = g_{k}(u-E_K)\\
I_{L} = g_{L}(u-E_L)
\end{eqnarray}
The equation describing the membrane potential can thus be written as follows,
\begin{eqnarray}
C_m\frac{dV}{dt}=−I_{Na}(t)−I_{K}(t)−I_{L}(t)+I(t)
\end{eqnarray}
Hodgkin and Huxley discovered that the $Na$ and the $K$ channels do not act as Ohmic conductances, but are modulated by the potential across the membrane.
Changes in potential had a nonlinear effect on flow of ionic currents. Based in their experimental results they obtained a system of differential equations that described the temporal evolution of the membrane potential in terms of changes in ionic currents (chiefly $\text{Na}^{+}$ and $\text{K}^{+}$).
\begin{eqnarray}
I = g_{Na}m^3h(u−E_{Na}) \\
I_K = g_Kn^4(u−E_K)\\
I_L = g_L(u−E_L)
\end{eqnarray}
where $E_{Na}=50\ mV$, $E_K = -95\ mV$ and $E_L=-55\ mV$ are the reversal potentials; $g_{Na} = 100\ \mu S/cm^2$, $g_K = 10\ \mu S/cm^2$ and $g_L = 0.15\ \mu S/cm^2$ are the channel conductances; and m,h, and n are gating variables that follow the dynamics given by:
\begin{eqnarray}
\frac{dm}{dt} = - \frac{1}{\tau_m}(m-m_0)\\
\frac{dh}{dt} = - \frac{1}{\tau_h}(h-h_0)\\
\frac{dn}{dt} = - \frac{1}{\tau_n}(n-n_0)
\end{eqnarray}
where $\tau_m$, $\tau_h$ and $\tau_n$ are empirically determined voltage dependent time constants and $m_0$, $h_0$ and $n_0$ are voltage dependent asymptotic gating values.
<img src="https://raw.githubusercontent.com/neurorishika/PSST/master/Tutorial/Day%203%20Cells%20in%20Silicon/mhn.svg" width="800"/>
On day 2, we had created a RK4 based numerical integrator. Recall this implementation:
```
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
## OR for Tensorflow 2.0 and above ##
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
%matplotlib inline
def tf_check_type(t, y0): # Ensure Input is Correct
if not (y0.dtype.is_floating and t.dtype.is_floating):
raise TypeError('Error in Datatype')
class _Tf_Integrator():
def integrate(self, func, y0, t):
time_delta_grid = t[1:] - t[:-1]
def scan_func(y, t_dt):
t, dt = t_dt
dy = self._step_func(func,t,dt,y) # Make code more modular.
return y + dy
y = tf.scan(scan_func, (t[:-1], time_delta_grid),y0)
return tf.concat([[y0], y], axis=0)
def _step_func(self, func, t, dt, y):
k1 = func(y, t)
half_step = t + dt / 2
dt_cast = tf.cast(dt, y.dtype) # Failsafe
k2 = func(y + dt_cast * k1 / 2, half_step)
k3 = func(y + dt_cast * k2 / 2, half_step)
k4 = func(y + dt_cast * k3, t + dt)
return tf.add_n([k1, 2 * k2, 2 * k3, k4]) * (dt_cast / 6)
def odeint(func, y0, t):
t = tf.convert_to_tensor(t, name='t')
y0 = tf.convert_to_tensor(y0, name='y0')
tf_check_type(y0,t)
return _Tf_Integrator().integrate(func,y0,t)
```
#### Implementing the Hodgkin-Huxley neuron model
The variables of the Hodgkin Huxley neuron model that are updated at each integration time step are, the membrane potential, $V$, the sodium activation gating variable, $m$, the sodium inactivation gating variable, $h$, and the potassium channel gating variable, $n$. The dynamics are given by Equations above. In the following code, we define the parameters associated with the conductances, including the formulae for $\tau_{m}$, $\tau_{h}$, $\tau_{n}$ and the voltage dependent steady state values of the gating variables.
##### Step 1: Defining Parameters of the Neuron
```
C_m = 1 # Membrane Capacitance
g_K = 10
E_K = -95
g_Na = 100
E_Na = 50
g_L = 0.15
E_L = -55
```
##### Step 2: Defining functions that calculate $\tau_m$, $\tau_h$, $\tau_n$, $m_0$, $h_0$, $n_0$
Note: Always use Tensorflow functions for all mathematical operations.
For our Hodgkin Huxley Model, we will determine the values of $\tau_m$, $\tau_h$, $\tau_n$, $m_0$, $h_0$, $n_0$ by the following equations:
<img src="https://raw.githubusercontent.com/neurorishika/PSST/master/Tutorial/Day%203%20Cells%20in%20Silicon/eqn1.svg" width="800"/>
```
def K_prop(V):
T = 22
phi = 3.0**((T-36.0)/10)
V_ = V-(-50)
alpha_n = 0.02*(15.0 - V_)/(tf.exp((15.0 - V_)/5.0) - 1.0)
beta_n = 0.5*tf.exp((10.0 - V_)/40.0)
t_n = 1.0/((alpha_n+beta_n)*phi)
n_0 = alpha_n/(alpha_n+beta_n)
return n_0, t_n
def Na_prop(V):
T = 22
phi = 3.0**((T-36)/10)
V_ = V-(-50)
alpha_m = 0.32*(13.0 - V_)/(tf.exp((13.0 - V_)/4.0) - 1.0)
beta_m = 0.28*(V_ - 40.0)/(tf.exp((V_ - 40.0)/5.0) - 1.0)
alpha_h = 0.128*tf.exp((17.0 - V_)/18.0)
beta_h = 4.0/(tf.exp((40.0 - V_)/5.0) + 1.0)
t_m = 1.0/((alpha_m+beta_m)*phi)
t_h = 1.0/((alpha_h+beta_h)*phi)
m_0 = alpha_m/(alpha_m+beta_m)
h_0 = alpha_h/(alpha_h+beta_h)
return m_0, t_m, h_0, t_h
```
##### Step 3: Defining function that calculate Neuronal currents
<img src="https://raw.githubusercontent.com/neurorishika/PSST/master/Tutorial/Day%203%20Cells%20in%20Silicon/eqn2.svg" width="800"/>
```
def I_K(V, n):
return g_K * n**4 * (V - E_K)
def I_Na(V, m, h):
return g_Na * m**3 * h * (V - E_Na)
def I_L(V):
return g_L * (V - E_L)
```
##### Step 4: Define the function dX/dt where X is the State Vector
```
def dXdt(X, t):
V = X[0:1]
m = X[1:2]
h = X[2:3]
n = X[3:4]
dVdt = (5 - I_Na(V, m, h) - I_K(V, n) - I_L(V)) / C_m
# Here the current injection I_injected = 5 uA
m0,tm,h0,th = Na_prop(V)
n0,tn = K_prop(V)
dmdt = - (1.0/tm)*(m-m0)
dhdt = - (1.0/th)*(h-h0)
dndt = - (1.0/tn)*(n-n0)
out = tf.concat([dVdt,dmdt,dhdt,dndt],0)
return out
```
##### Step 5: Define Initial Condition and Integrate
```
y0 = tf.constant([-71,0,0,0], dtype=tf.float64)
epsilon = 0.01
t = np.arange(0,200,epsilon)
state = odeint(dXdt,y0,t)
with tf.Session() as sess:
state = sess.run(state)
```
##### Step 6: Plot Output
```
plt.plot(t,state.T[0,:])
plt.xlabel("Time (in ms)")
plt.ylabel("Voltage (in mV)")
plt.show()
```
#### Simulating Multiple Independent HH Neurons at the Same Time
Here we illustrate some simple steps that can be used to simulate populations of neurons efficiently. Key to setting up the equations is to order it in a manner that utilizes TensorFlow's algorithms that distribute vector, matrix and tensor computations over multiple cores. Consider a system of 20 independent HH neurons with different input currents that characterise the firing rates.
##### Methods of Parallelization
TensorFlow has built-in functions that speed up Tensor computations using available multi-cores, and GPU/TPU setups. There are two major parts of the code where such a speed-up can be effected
1. **RK4 iterations** Our implementation of the integrator utilizes Tensors as inputs.
2. **Functional Evaluations:** The form of the equations that describe the neuronal dynamics, are common across neurons. Only the parameters for differ across neurons. This can be used to `vectorize' the equations.
Say $\vec{X}=[V,m,n,h]$ is the state vector of a single neuron and its dynamics are defined using parameters $C_m,g_K,...E_L$ equations of the form:
\begin{eqnarray}\frac{d\vec{X}}{dt} = [f_1(\vec{X},C_m,g_K,...E_L),f_2(\vec{X},C_m,g_K,...E_L)...f_m(\vec{X},C_m,g_K,...E_L)]\end{eqnarray}
We can convert these equations to a form in which all evaluations are done as vector calculations and NOT scalar calculations. Despite the parameters being different, the functional forms of the equations are similar for the same state variable of different neurons. Thus, the trick is to reorganize $\mathbf{X}$ as $\mathbf{X'}=[(V_1,V_2,...V_n),(m_1,m_2,...m_n),(h_1,h_2,...h_n),(n_1,n_2,...n_n)]=[\vec{V},\vec{m},\vec{h},\vec{n}]$. And the parameters as $[\vec{C_m},\vec{g_K}] = [C_{m_{1}}\dots C_{m_{n}},g_{K_{1}}\dots g_{K_{n}}]$ and so on.
The advantage of this re-ordering is that the differential equation of the form,
\begin{eqnarray}\frac{dV_i}{dt}=f(V_i,m_i,h_i,n_i,C_{m_i},g_{K_i}...)\end{eqnarray}
is now easily parallelizable using a vector computation of the form,
\begin{eqnarray}\frac{d\vec{V}}{dt}=f(\vec{V},\vec{m},\vec{h},\vec{n},\vec{C_m},\vec{g_K}...)\end{eqnarray}
The equations can now be written in the form,
\begin{eqnarray}\frac{d\mathbf{X'}}{dt}= \Big[\frac{d\vec{V}}{dt},\frac{d\vec{m}}{dt},\frac{d\vec{h}}{dt},\frac{d\vec{n}}{dt}\Big]\end{eqnarray}
```
n_n = 20 # number of simultaneous neurons to simulate
# parameters will now become n_n-vectors
C_m = [1.0]*n_n
g_K = [10.0]*n_n
E_K = [-95.0]*n_n
g_Na = [100]*n_n
E_Na = [50]*n_n
g_L = [0.15]*n_n
E_L = [-55.0]*n_n
def K_prop(V):
T = 22
phi = 3.0**((T-36.0)/10)
V_ = V-(-50)
alpha_n = 0.02*(15.0 - V_)/(tf.exp((15.0 - V_)/5.0) - 1.0)
beta_n = 0.5*tf.exp((10.0 - V_)/40.0)
t_n = 1.0/((alpha_n+beta_n)*phi)
n_0 = alpha_n/(alpha_n+beta_n)
return n_0, t_n
def Na_prop(V):
T = 22
phi = 3.0**((T-36)/10)
V_ = V-(-50)
alpha_m = 0.32*(13.0 - V_)/(tf.exp((13.0 - V_)/4.0) - 1.0)
beta_m = 0.28*(V_ - 40.0)/(tf.exp((V_ - 40.0)/5.0) - 1.0)
alpha_h = 0.128*tf.exp((17.0 - V_)/18.0)
beta_h = 4.0/(tf.exp((40.0 - V_)/5.0) + 1.0)
t_m = 1.0/((alpha_m+beta_m)*phi)
t_h = 1.0/((alpha_h+beta_h)*phi)
m_0 = alpha_m/(alpha_m+beta_m)
h_0 = alpha_h/(alpha_h+beta_h)
return m_0, t_m, h_0, t_h
def I_K(V, n):
return g_K * n**4 * (V - E_K)
def I_Na(V, m, h):
return g_Na * m**3 * h * (V - E_Na)
def I_L(V):
return g_L * (V - E_L)
def dXdt(X, t):
V = X[:1*n_n] # First n_n values are Membrane Voltage
m = X[1*n_n:2*n_n] # Next n_n values are Sodium Activation Gating Variables
h = X[2*n_n:3*n_n] # Next n_n values are Sodium Inactivation Gating Variables
n = X[3*n_n:] # Last n_n values are Potassium Gating Variables
dVdt = (np.linspace(0,10,n_n) - I_Na(V, m, h) - I_K(V, n) -I_L(V)) / C_m
# Input current is linearly varied between 0 and 10
m0,tm,h0,th = Na_prop(V)
n0,tn = K_prop(V)
dmdt = - (1.0/tm)*(m-m0)
dhdt = - (1.0/th)*(h-h0)
dndt = - (1.0/tn)*(n-n0)
out = tf.concat([dVdt,dmdt,dhdt,dndt],0)
return out
y0 = tf.constant([-71]*n_n+[0,0,0]*n_n, dtype=tf.float64)
epsilon = 0.01
t = np.arange(0,200,epsilon)
state = odeint(dXdt,y0,t)
with tf.Session() as sess:
state = sess.run(state)
plt.figure(figsize=(12,17))
for i in range(20):
plt.subplot(10,2,i+1)
plt.plot(t,state[:,i])
plt.title("Injected Current = {:0.1f}".format(i/2))
plt.ylim([-90,60])
plt.xlabel("Time (in ms)")
plt.ylabel("Voltage (in mV)")
plt.tight_layout()
plt.show()
```
#### Quantifying the Firing Rates against Input Current
The firing frequency as a function of the input is shown in the figure below. The code to generate the firing rate is below.
```
plt.plot(np.linspace(0,10,20),np.bitwise_and(state[:-1,:20]<0,state[1:,:20]>0).sum(axis=0)/0.2,"o")
plt.xlabel("Injected Current(mA)")
plt.ylabel("Firing Rate (Hz)")
plt.show()
```
# References
(<a id="cit-Dayan2005" href="#call-Dayan2005">Dayan and Abbott, 2005</a>) Peter Dayan and Larry F. Abbott, ``Theoretical Neuroscience - Computational and Mathematical Modeling of Neural Systems``, 2005.
(<a id="cit-Johnston1995" href="#call-Johnston1995">Johnston and Wu, 1995</a>) D. Johnston and S. M.S. Wu, ``Foundations of cellular neurophysiology``, 1995.
(<a id="cit-Huxley1952" href="#call-Huxley1952">Huxley and Hodgkin, 1952</a>) Huxley A. L. and Hodgkin A. F., ``Quantitative description of nerve current``, Journal of Physiology, vol. , number , pp. , 1952.
(<a id="cit-gerstnerMOOC" href="#call-gerstnerMOOC">MOOC</a>) , ``Neuronal dynamics``, . [online](https://www.edx.org/course/neuronal-dynamics)
(<a id="cit-compneuroMOOC" href="#call-compneuroMOOC">MOOC</a>) , ``Computational Neuroscience``, . [online](https://www.coursera.org/learn/computational-neuroscience)
(<a id="cit-Gerstner2014" href="#call-Gerstner2014">Gerstner, Kistler <em>et al.</em>, 2014</a>) Wulfram Gerstner, Werner M. Kistler, Richard Naud <em>et al.</em>, ``Neuronal dynamics: From single neurons to networks and models of cognition``, 2014.
| github_jupyter |
<a href="https://colab.research.google.com/github/nnuncert/nnuncert/blob/master/notebooks/DNNC_toy.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Git + Repo Installs
```
!git clone https://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@github.com/nnuncert/nnuncert
%cd nnuncert
!pip install -r requirements.txt
```
# Imports
```
# %cd nnuncert
# general imports
import numpy as np
import numexpr as ne
import tensorflow as tf
import matplotlib.pyplot as plt
# thesis code
import nnuncert
from nnuncert.models import make_model, type2name
from nnuncert.app.toy import make_toy_data, make_toy_plot, gen_2d_gaussian_samples, input2grid, contour_plot_2d
from nnuncert.utils.traintest import TrainTestSplit
from nnuncert.utils.dist import Dist
```
# Toy 1D
## Make data
Generate input features on (-4, 4) uniformly and calculate noisy targets with true function 'x**3' and additive homoscedastic noise N(0, 3).
```
# set seed for reproducibility and make random number generator
seed = 21
rng = np.random.default_rng(seed)
# define function that generates true relationship, possible to use any kind of expression such as "sin(x)", "exp(x)", "3*x**2 - 8x + 14", ...
def reg_func(x):
reg_func = "x**3"
return ne.evaluate(reg_func)
# generate input data (x) uniformly from -4 to 4
x = rng.random((20, ))*8 - 4
# alternatively: setup (multiple) clusters and sample uniformly in between
# ppc = 20 # points per cluster
# clusters = [[-4, 4]] # list of cluster bounds (left, right)
# x = np.array([rng.choice(np.linspace(x1, x2, 1000), ppc)
# for (x1, x2) in clusters]).ravel()
# generate responses with function
noise_std = 3
data_1d = make_toy_data(x, reg_func, noise_std, seed=rng)
# have a look at the data
data_1d.head()
# plot ground truth
fig, ax = plt.subplots(figsize=(8, 4))
minx, maxx = [-6, 6]
x0 = np.linspace(minx, maxx, 80)
ax.scatter(data_1d["x1"], data_1d["y"])
ax.plot(x0, reg_func(x0), "--", color="black")
```
## Fit model
Fit a model to all training samples that were generated.
Get predictitve mean, variance for 100 inputs in (-6, 6), evenly spaced.
```
class TrainTestSplitToy1D(TrainTestSplit):
def __init__(self, df, train_id = None, test_id = None, test_ratio=0.1, norm_x=False, rng=None):
non_norm = []
if norm_x is False:
non_norm = ["x1"]
super(TrainTestSplitToy1D, self).__init__(df, "y", non_norm=non_norm,train_id=train_id, test_id=test_id, ratio=test_ratio, rng=rng)
# standardize features x
toy_1d = TrainTestSplitToy1D(data_1d, test_ratio=0, norm_x=True, rng=rng)
input_shape = toy_1d.x_train.shape[1]
# get empirical marginal desnity of y (not meaningful for toy data)
# method = "gauss" or "ssv"
dist = Dist._from_values(data_1d.y, method="gauss")
y0 = np.linspace(data_1d.y.min(), data_1d.y.max(), 100)
plt.plot(y0, dist.pdf(y0))
_ = plt.hist(data_1d.y, density=True, bins=12, color="grey")
# handle general settings
arch = [[50, "relu", 0]] # list of hidden layer description (size, act. func, dropout rate)
epochs = 40
verbose = 0
learning_rate = 0.01
# make model and compile
model = make_model("DNNC-R", input_shape, arch)
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate), metrics=["mae", "mse"])
# fit model to all samples (test_ratio in TrainTestSplitToy1D was set to 0)
model.fit(toy_1d.x_train, toy_1d.y_train, epochs=epochs, verbose=verbose, dist=dist)
# get prediction on scaled test input features
xlim = (-6, 6)
x0 = np.linspace(*xlim, 100)
pred = model.make_prediction(toy_1d.scale_x(x0))
```
## Plot
Plot mean prediction and predictive uncertainty (2 standard deviations).
```
# setup some colors and setup limits for y axis
colors = ["mediumblue", "tab:blue", "#b3cde0"]
ylim = (-150, 150)
fig, ax = plt.subplots(figsize=(8, 5))
plt.setp(ax, xlim=xlim, ylim=ylim)
make_toy_plot(pred, toy_1d.x_train_us, toy_1d.y_train_us, x0=x0, reg_func=reg_func, std_devs=2, colors=colors, ax=ax)
```
# Toy 2D
## Make data
Create input data clusters located at (-1, -1) and (1, 1) and generate noise response with standard deviation = 0.01 (very low noise).
```
# set seed for reproducibility and make random number generator
seed = 21
rng = np.random.default_rng(seed)
# define function that generates true relationship
def reg_func_2d(x):
x1, x2 = x.T
reg_func = "0.8*x1 + 0.8*x2"
return ne.evaluate(reg_func)
# generate input data (x), defined by cluster centers muh and noise in the data
muh = [[-1, -1], [1, 1]]
x = gen_2d_gaussian_samples(muh, var=0.02, ppc=50, seed=rng)
# make y data
toy2d_noise_std = 0.1
data2d = make_toy_data(x, reg_func_2d, toy2d_noise_std, seed=rng)
# show data
data2d.head()
# plot noisy data in 3D
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.scatter(data2d.x1, data2d.x2, data2d.y, c="red")
```
## Fit model
Fit a model to all training samples that were generated.
Get predictitve mean, variance for inputs on a [-2, 2] x [-2, 2] grid.
```
class TrainTestSplitToy2D(TrainTestSplit):
def __init__(self, df, train_id = None, test_id = None, test_ratio=0.1, norm_x=False, rng=None):
non_norm = ["x1", "x2"]
if norm_x is True:
non_norm = []
super(TrainTestSplitToy2D, self).__init__(df, "y", non_norm=non_norm, train_id=train_id, test_id=test_id, ratio=test_ratio, rng=rng)
# standardize features x
toy_2d = TrainTestSplitToy2D(data2d, test_ratio=0, norm_x=True, rng=rng)
input_shape = toy_2d.x_train.shape[1]
# get empirical marginal desnity of y (not meaningful for toy data)
# method = "gauss" or "ssv"
dist = Dist._from_values(data2d.y, method="ssv")
y0 = np.linspace(data2d.y.min(), data2d.y.max(), 100)
plt.plot(y0, dist.pdf(y0))
_ = plt.hist(data2d.y, density=True, bins=12, color="grey")
# handle general settings
arch = [[50, "relu", 0]] # list of hidden layer description (size, act. func, dropout rate)
epochs = 40
verbose = 0
learning_rate = 0.01
# make model and compile
model = make_model("DNNC-R", input_shape, arch)
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate), metrics=["mae", "mse"])
# fit model to all samples (test_ratio in TrainTestSplitToy1D was set to 0)
model.fit(toy_2d.x_train, toy_2d.y_train, epochs=epochs, verbose=verbose, dist=dist)
# generate scaled test features
ppa = 100 # points per axis -> 100*100=10,000 test points
grid_in, x1, x2 = input2grid([-2, 2], [-2, 2], ppa)
x_test = toy_2d.scale_x(grid_in)
# get prediction of model for test features
pred = model.make_prediction(x_test)
```
## Plot
Contour plot of predictive uncertainty (std. deviation) for input features.
```
# init plot and define colormap
fig, ax = plt.subplots(figsize=(8, 8))
cmap = plt.get_cmap("viridis")
# retrieve predictive standard deviation in proper form
std = pred.std_total.reshape(ppa, ppa)
# plot std as contour to ax
contour_plot_2d(x1, x2, std, x_train=(data2d[["x1", "x2"]].values, "white"), levels=20, cmap=cmap, ax=ax, fig_=fig, make_colbar=True)
```
| github_jupyter |
# Goals
My goal with this dataset is it use a regression machine learning model to accurately predict the price of a house dependent on features. I will also look through some data analysis and a few other features such as PCA
## Process
I’ll be following a typical data science pipeline, “OSEMN”.
1. Obtaining the data is the first approach in solving the problem.
2. Scrubbing or cleaning the data is the next step. This includes data imputation of missing or invalid data and fixing column names.
3. Exploring the data will follow right after and allow further insight of what our dataset contains. Looking for any outliers or weird data. Understanding the relationships between features.
4. Modeling the data will give us our predictive power on whether an employee will leave.
5. INterpreting the data is last. With all the results and analysis of the data, what conclusion is made?
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
sns.set_style('darkgrid')
```
## 1.0 Obtaining the data
The data I'm using has been taken from Kaggle and it contains apartment sales data in Daegu, South Korea from 2007 to 2017. This dataset is particularly interesting as it contains many interesting features for me to analyse. Another reason this dataset is interesting is due to the fact that it contains 10 years of data, which allows me to undertake some interesting time analysis.
```
df = pd.read_csv('daegu_housing.csv')
df.head()
```
## 2.0 Scrubbing the data
The first stage of the process is scrub the data. This means I will be searching the data for any missing data, and making sure the data frame is in an easily readable format.
```
# Checking to see if any of the features have missing data
df.isnull().any()
```
## 3.0 Exploring the data
I have confirmed that there is no missing data and am happy with the layout of the rest of the data frame. Now I will explore a few areas that intrigue me most. This stage of the process is incredibly important, especially if it's your first time using it, as it allows you to understand the data more and the relationships held within.
```
# To begin with I will look at the distribution of prices across the dataset
df['SalePrice'].hist()
plt.xlabel('Sale Price')
plt.ylabel('Count')
plt.title('Histogram of Sale Price');
sns.jointplot(x='SalePrice',y='Size(sqf)',data=df,kind='reg',xlim=(0,600000),ylim=(0,2500))
plt.xlabel('Sale Price')
plt.ylabel('Size (sqf)')
plt.title('Sale Price against Size');
```
The relationship between sale price and square foot is always an interesting one to plot. Here we can see that there is a strong correlation, and also from this we can see that the square foot data is trenched. The same square footage can be sold for a wide range of prices. There is a trench at approx. 900 sqft which is sold for between 50,000 and 450,000. This helps us understand that there are other significant factors in play here.
```
plt.figure(figsize=(12,6))
sns.boxplot(x='YrSold',y='SalePrice',data=df)
plt.xlabel('Year Sold')
plt.ylabel('Sale Price')
plt.title('Year Sold against Sale Price');
```
### 3.1 Deeper exploration
Now that I have looked at some overarching trends within the data, I would like to explore two themes in more depth. Those are:
1. The affordable housing within the city
2. The price according to which subway station you are near
### 3.1.1 House affordability
I will be looking at house affordability in 2017. I was unable to find reliable data for the average salary in Daegu from 2007 onwards so it would be unfair and crude to just adjust for inflation as there are many other factors at play here. I instead will be looking at the distribution of affordability in just 2017.
To do this I found the UN's housing affordability index which is calculated as house price/average salary.
```
# The mean salary in Daegu for 2017 in dollars
avg_sal_2017 = 39000
def affordability(price):
# UN housing affordability index formula
index = price/39000
if index >= 5.1:
return 'Severely Unaffordable'
if 4.1 <= index <= 5.0:
return 'Seriously Unaffordable'
if 3.1 <= index <= 4.0:
return 'Moderately Unaffordable'
if index <= 3.0:
return 'Affordable'
df['Affordability_index'] = df['SalePrice'].apply(lambda x: affordability(x))
df[df['YrSold'] == 2017]['Affordability_index'].value_counts()
```
From my rough calculations I can see that only a small proportion of apartments sold in Daegu in 2017 were classes as affordable.
Now that I have created this new feature, I have added it to my dataset to increase the efficiency of the mode later on.
### 3.1.2 Price relating to subway station
One area of the data that I'm particularly interested in is the nearest subway station. I'm particularly interested to see if certain subway stations relate to a higher price point, and the development of these prices over time.
First I will look at the stations in the data and the frequency count
```
df['SubwayStation'].value_counts()
```
There are several ways to undertake this analysis. I have decided to make a data frame for each station and then undertake the analysis off of this.
```
Kyungbuk = pd.DataFrame(data=df[df['SubwayStation'] == 'Kyungbuk_uni_hospital'],columns=['SalePrice','YrSold'])
Myung_duk = pd.DataFrame(data=df[df['SubwayStation'] == 'Myung-duk'],columns=['SalePrice','YrSold'])
Banwoldang = pd.DataFrame(data=df[df['SubwayStation'] == 'Banwoldang'],columns=['SalePrice','YrSold'])
Bangoge = pd.DataFrame(data=df[df['SubwayStation'] == 'Bangoge'],columns=['SalePrice','YrSold'])
Sin_nam = pd.DataFrame(data=df[df['SubwayStation'] == 'Sin-nam'],columns=['SalePrice','YrSold'])
no_subway_nearby = pd.DataFrame(data=df[df['SubwayStation'] == 'no_subway_nearby'],columns=['SalePrice','YrSold'])
Chil_sung_market = pd.DataFrame(data=df[df['SubwayStation'] == 'Chil-sung-market'],columns=['SalePrice','YrSold'])
Daegu = pd.DataFrame(data=df[df['SubwayStation'] == 'Daegu'],columns=['SalePrice','YrSold'])
fig, ax = plt.subplots(figsize=(12,8))
Kyungbuk.groupby('YrSold').mean().plot(ax=ax,color='y')
Myung_duk.groupby('YrSold').mean().plot(ax=ax,color='m',linewidth=3)
Banwoldang.groupby('YrSold').mean().plot(ax=ax,color='c')
Bangoge.groupby('YrSold').mean().plot(ax=ax,color='r')
Sin_nam.groupby('YrSold').mean().plot(ax=ax,color='g')
no_subway_nearby.groupby('YrSold').mean().plot(ax=ax,color='b')
Chil_sung_market.groupby('YrSold').mean().plot(ax=ax,color='pink')
Daegu.groupby('YrSold').mean().plot(ax=ax,color='k',linewidth=3)
plt.xlabel('Year Sold')
plt.ylabel('Mean Price Sold')
plt.title('Mean price over time per subway station')
ax.legend(['Kyungbuk','Myung duk','Banwoldang','Bangoge','Sin nam','No subway','Chil sung market','Daegu'],loc='upper center',bbox_to_anchor=(1.1, 0.6));
```
There are two stations here which seemingly have grown in popularity over the past 10 years. They are: Daegu and Myung Duk, both highlighted in the plot. The mean house price in those areas has increased dramatically over the past 10 years.
# 4.0 Modelling the data
I have undertook some analysis into the data, now I will fit my machine learning model. In this project I have chosen to use the linear regression model.
```
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
```
## 4.1 First iteration of the model
This is my first iteration of the model, the only adjustments I will be making are that I will be removing the features that are strings as the machine learning model cannot take this data in. I will drop these columns from the data frame, split the data into training and testing data, and then fit the model.
```
df_non_string = df.drop(['Affordability_index','HallwayType', 'HeatingType', 'AptManageType', 'SubwayStation', 'TimeToBusStop', 'TimeToSubway'],axis=1)
x = df_non_string.drop('SalePrice',axis=1)
y = df_non_string['SalePrice']
x_train, x_test, y_train, y_test = train_test_split(x,y,test_size=0.33)
lr = LinearRegression()
lr.fit(x_train,y_train)
predictions = lr.predict(x_test)
```
Now that I have created my predictions for the test data it is important to see how effective it was. One way to do this is to look at it visually. To do so I will create a scatter plot of Predictions against the actual Test Data. The more correlated this data is, the more accurate my model is.
```
plt.scatter(predictions,y_test)
plt.xlabel('Predictions')
plt.ylabel('Test Data')
plt.title('Predictions against Test Data');
```
This plot is fairly correlated, indicating a high level of accuracy. The model seems to struggle the higher up the price bracket it goes, however that is not unexpected considering the bulk of my data is for houses towards the least expensive end.
To test the effectiveness of the model in an analytical manner I will use the linear regression score (Mean Accuracy).
```
print('The Linear Regression score for the training data is: {}'.format(lr.score(x_train,y_train)))
print('The Linear Regression score for the test data is: {}'.format(lr.score(x_test,y_test)))
```
## 4.2 Improving the model
To improve the accuracy of this model I will be using 3 methods.
1. Categorical features
2. Rescale data
3. PCA
### 4.2.1 Adding categorical features
I believe that the best way to improve the accuracy of this model is to add categorical features. In the first iteration I dropped the string based features from the model, however to improve accuracy I will convert these to categorical features.
```
hallway = pd.get_dummies(df['HallwayType'])
heating = pd.get_dummies(df['HeatingType'])
manager_type = pd.get_dummies(df['AptManageType'])
bus_time = pd.get_dummies(df['TimeToBusStop'])
subway_time = pd.get_dummies(df['TimeToSubway'])
subway = pd.get_dummies(df['SubwayStation'])
affordability = pd.get_dummies(df['Affordability_index'])
ready_data = pd.concat([df,affordability,hallway,heating,manager_type,bus_time,subway_time,subway],axis=1)
ready_data.drop(['Affordability_index','HallwayType', 'HeatingType', 'AptManageType','TimeToBusStop', 'TimeToSubway', 'SubwayStation'],inplace=True,axis=1)
ready_data.head()
x = ready_data.drop('SalePrice',axis=1)
y = ready_data['SalePrice']
x_train, x_test, y_train, y_test = train_test_split(x,y,test_size=0.33)
lr = LinearRegression()
lr.fit(x_train,y_train)
predictions = lr.predict(x_test)
print('The Linear Regression score for the training data is: {}'.format(lr.score(x_train,y_train)))
print('The Linear Regression score for the test data is: {}'.format(lr.score(x_test,y_test)))
```
### 4.2.2 Rescale the data
To further improve the accuracy I believe that rescaling the data will help. I will use the StandardScaler to standardise my data and center it around 0.
```
from sklearn.preprocessing import StandardScaler
x_train, x_test, y_train, y_test = train_test_split(ready_data.drop('SalePrice',axis=1),ready_data['SalePrice'],test_size=0.33)
scaler = StandardScaler().fit(x_train)
standardised_x_train = scaler.transform(x_train)
standardised_x_test = scaler.transform(x_test)
lr = LinearRegression()
lr.fit(standardised_x_train,y_train)
predictions = lr.predict(standardised_x_test)
print('The Linear Regression score for the training data is: {}'.format(lr.score(standardised_x_train,y_train)))
print('The Linear Regression score for the test data is: {}'.format(lr.score(standardised_x_test,y_test)))
```
There doesn't appear to have been a significant change in accuracy by scaling the data.
### 4.2.3 What effect would PCA have?
Principal Component Analysis (PCA) is a method to reduce the number of components within a dataframe whilst still maintaining a high level of accuracy. I don't think this will improve the accuracy of the model, however I am interested to see the impact. I am also aware that if this dataset were to have a large number of features, this would be an important step to speed up training.
```
from sklearn.decomposition import PCA
df_data = ready_data.drop('SalePrice',axis=1)
df_target = ready_data['SalePrice']
pca = PCA(n_components=.95) #We want to cover 95% of variance
pca.fit(df_data)
pca_data = pd.DataFrame(pca.transform(df_data))
print("95% of variance can be accounted for with {} components".format(pca.n_components_))
x_train, x_test, y_train, y_test = train_test_split(pca_data,df_target,test_size=0.33)
scaler = StandardScaler().fit(x_train)
standardised_x_train = scaler.transform(x_train)
standardised_x_test = scaler.transform(x_test)
lr = LinearRegression()
lr.fit(standardised_x_train,y_train)
predictions = lr.predict(x_test)
print('The Linear Regression score for the training data is: {}'.format(lr.score(standardised_x_train,y_train)))
print('The Linear Regression score for the test data is: {}'.format(lr.score(standardised_x_test,y_test)))
```
By using PCA with 3 components our accuracy has reduced drastically from 90% to 63%. The more components that I introduce however, the higher this number would be, however by adding too many you are defeating the original idea of using PCA.
## 5.0 Interpreting the results
I have gone through all the stages of my data exploration, and I believe that I have been able to pull out some interesting insights and been able to create an accurate machine learning model for predicting the price of an apartment in Daegu, South Korea. Some interesting points from this exploration:
1. Affordable areas of the city have, in the past 10 years, become unaffordable (3.1.1)
2. Affordable housing in Daegu is lacking (3.1.2)
3. The largest increase in model accuracy was by adding categorical features (4.2.1)
| github_jupyter |
# Hatch Template!
## Dandelion Voting
Note: What are peoples goal target raise?
1. Percentage of total tokens that have to vote 'yes' to `something` for it to pass.
```
import param
import panel as pn
import pandas as pd
import hvplot.pandas
import holoviews as hv
import numpy as np
pn.extension()
class DandelionVoting(param.Parameterized):
total_tokens = param.Number(1e6, constant=True)
support_required = param.Number(0.6, bounds=(0.5,0.9), step=0.01)
acceptance_quorum = param.Number(0.02, bounds=(0.01,1), step=0.01)
vote_duration_days = param.Number(3, bounds=(1,14), step=1)
vote_buffer_hours = param.Number(8, bounds=(1,48), step=1)
rage_quit_hours = param.Number(24, bounds=(1, 48), step=1)
tollgate_fee_xdai = param.Number(3, bounds=(1,100), step=1)
def vote_pass_view(self):
x = np.linspace(0, self.total_tokens, num=100)
y = [a*self.support_required for a in x]
df = pd.DataFrame(zip(x,y))
y_fill = [a for a in x]
df_fill = pd.DataFrame(zip(x,y_fill))
y_fill_quorum = [a for i, a in enumerate(x) if i < self.acceptance_quorum*len(x)]
df_fill_q = pd.DataFrame(zip(x,y_fill_quorum))
return df_fill.hvplot.area(x='0', y='1', xformatter='%.0f', yformatter='%.0f', color='green', xlabel='Total Token Votes', ylabel='Yes Token Votes') * \
df.hvplot.area(x='0', y='1', xformatter='%.0f', yformatter='%.0f', color='red') * \
df_fill_q.hvplot.area(x='0', y='1', xformatter='%.0f', yformatter='%.0f', color='red')
d = DandelionVoting()
pn.Row(d, d.vote_pass_view)
import numpy as np
x = np.linspace(0, d.total_tokens, num=100)
y = [a*0.6 for a in x]
df = pd.DataFrame(zip(x,y))
df_fill = pd.DataFrame(zip(x,x))
y_fill_quorum = [a for i, a in enumerate(x) if i < 0.5*len(x)]
df_fill_q = pd.DataFrame(zip(x,y_fill_quorum))
df_fill.hvplot.area(x='0', y='1', xformatter='%.0f', yformatter='%.0f', color='green') * \
df.hvplot.area(x='0', y='1', xformatter='%.0f', yformatter='%.0f', color='red') * \
df_fill_q.hvplot.area(x='0', y='1', xformatter='%.0f', yformatter='%.0f', color='red')
df_fill_q.hvplot.area(x='0', y='1', xformatter='%.0f', yformatter='%.0f', color='red')
df_fill_q
class DandelionVoting(param.Parameterized):
total_tokens = param.Number(1e6, constant=True)
minimum_quorum = param.Number(0.02, bounds=(0,1), step=0.01)
support_required = param.Number(0.5, bounds=(0.5,0.9), step=0.01)
days_to_vote_on_proposal = param.Integer(3 + 8 + 24, bounds=(0,100))
days_to_exit_hatch = param.Integer(8)
# vote_buffer_blocks = param.Integer(8, bounds=(0,None))
# vote_execution_delay_blocks = param.Integer(24, bounds=(0,None))
cost_to_make_a_proposal = param.Number(3, step=1, doc="cost to make a proposal")
maximum_number_proposals_per_month = param.Number(10, bounds=(1, 100))
def view(self):
min_yes_tokens = self.support_required * self.minimum_quorum * self.total_tokens
min_blockers = (1 - self.support_required) * self.minimum_quorum * self.total_tokens
votes = pd.DataFrame.from_dict({'Votes': [min_yes_tokens, min_blockers]}, orient='index', columns=['Minimum Tokens to Pass', 'Minimum Tokens for Quorum'])
vote_plot = votes.hvplot.bar(stacked=True, ylim=(0,self.total_tokens)).opts(color=hv.Cycle(['#0F2EEE', '#0b0a15', '#DEFB48']))
return pn.Row(vote_plot, pn.Column("Minimum Tokens to Meet Quorum: ", int(self.minimum_quorum * self.total_tokens), "Minimum Tokens to Pass a Vote: ", int(min_yes_tokens), "Minimum Tokens to Block a Vote: ", int(min_blockers)))
d = DandelionVoting()
pn.Pane(d)
pn.Column(d, d.view)
class Hatch(param.Parameterized):
# CSTK Ratio
total_cstk_tokens = param.Number(700000, constant=True)
hatch_oracle_ratio = param.Number(0.005, constant=True)
@param.depends('hatch_oracle_ratio', 'total_cstk_tokens')
def wxdai_range(self):
return pn.Row(pn.Pane("Cap on wxdai staked: "), self.hatch_oracle_ratio * self.total_cstk_tokens)
# Min and Target Goals
min_goal = param.Number(5, bounds=(1,100), step=10)
max_goal = param.Number(1000, bounds=(100,10000), step=50) # Something to consider -> target goal or max goal
# Hatch params
hatch_period = param.Integer(15, bounds=(5, 30), step=2)
hatch_exchange_rate = param.Number() # This needs to be tested and explained -> See the forum post
hatch_tribute = param.Number(0.05, bounds=(0,1))
h = Hatch()
pn.Pane(h)
pip install openpyxl
import pandas as pd
import panel as pn
import os
import hvplot.pandas
APP_PATH = './'
sheets = [
"Total Impact Hours so far",
"IH Predictions",
"#8 Jan 1",
"#7 Dec 18",
"#6 Dec 4",
"#5 Nov 20",
"#4 Nov 6",
"#3 Oct 23",
"#2 Oct 9",
"#1 Sept 24",
"#0 Sept 7 (historic)",
] + [f"#{i} IH Results" for i in range(9)]
sheets = {i:sheet for i, sheet in enumerate(sheets)}
def read_excel(sheet_name="Total Impact Hours so far", header=1, index_col=0, usecols=None) -> pd.DataFrame:
data = pd.read_excel(
os.path.join(APP_PATH, "data", "TEC Praise Quantification.xlsx"),
sheet_name=sheet_name,
engine='openpyxl',
header=header,
index_col=index_col,
usecols=usecols
).reset_index().dropna(how='any')
return data
## Tests
total_impact_hours = read_excel()
impact_hour_data = read_excel(sheet_name="IH Predictions", header=0, index_col=0, usecols='A:I').drop(index=19)
pn.Row(impact_hour_data.hvplot.table(), total_impact_hours.hvplot.table())
import numpy as np
class ImpactHours(param.Parameterized):
max_ih_rate = param.Number(0.01, bounds=(0,200))
expected_raise_per_ih = param.Number(0.012, bounds=(0,20))
@param.depends('max_ih_rate', 'expected_raise_per_ih')
def impact_hours_rewards(self):
x = np.linspace(h.min_goal, h.max_goal)
R = self.max_ih_rate
m = self.expected_raise_per_ih
H = total_impact_hours['Impact Hours'].sum()
y = [R* (x / (x + m*H)) for x in x]
df = pd.DataFrame([x,y]).T
df.columns = ['x','y']
return df.hvplot(x='x')
i = ImpactHours()
pn.Row(i, i.impact_hours_rewards)
pn.Pane(h)
pn.Row(d, h, i)
pn.Row(d.view, i.impact_hours_rewards)
```
# Target/Expected Goals
```
class CommunityParticipation(param.Parameterized)
```
| github_jupyter |
# Walkthrough: Multi Device Plugin and the DevCloud
This notebook is a demonstration showing you how to request an edge node with an Intel i5 CPU and load a model on the CPU, GPU, and VPU (Intel® Neural Compute Stick 2) at the same time using the Multi Device Plugin on Udacity's workspace integration with Intel's DevCloud. This notebook is just to give you an overview of the process (you won't be writing any code). In the next workspace, you'll be given TODO items to complete.
Below are the six steps we'll walk through in this notebook:
1. Creating a Python script to load the model
2. Creating a job submission script
3. Submitting a job using the `qsub` command
4. Checking the job status using the `liveQStat` function
5. Retrieving the output files using the `getResults` function
6. Viewing the resulting output
Click the **Introduction to Multi Device Plugin and the DevCloud** button below for a quick overview of the overall process. We'll then walk through each step of the process.
<span class="graffiti-highlight graffiti-id_u5l9o8a-id_9e2xr8h"><i></i><button>Introduction to Multi Device Plugin and the DevCloud</button></span>
#### IMPORTANT: Set up paths so we can run Dev Cloud utilities
You *must* run this every time you enter a Workspace session.
```
%env PATH=/opt/conda/bin:/opt/spark-2.4.3-bin-hadoop2.7/bin:/opt/conda/bin:/usr/local/sbin:/usr/local/bin:/usr/sbin:/usr/bin:/sbin:/bin:/opt/intel_devcloud_support
import os
import sys
sys.path.insert(0, os.path.abspath('/opt/intel_devcloud_support'))
sys.path.insert(0, os.path.abspath('/opt/intel'))
```
## The Model
We will be using the `vehicle-license-plate-detection-barrier-0106` model for this exercise.
Remember to use the appropriate model precisions for each device:
* IGPU - `FP16`
* VPU - `FP16`
* CPU - It is prefered to use `FP32`, but we have to use `FP16` since **GPU** and **VPU** use `FP16`
The model has already been downloaded for you in the `/data/models/intel` directory on Intel's DevCloud.
We will be running inference on an image of a car. The path to the image is `/data/resources/car.png`.
# Step 1: Creating a Python Script
The first step is to create a Python script that you can use to load the model and perform an inference. I have used the `%%writefile` magic command to create a Python file called `load_model_to_device.py`. This will create a new Python file in the working directory.
**Note**: The advantage of using the **Multi device plugin** is that it does not require us to change our application code. So we will be using the same Python script we used in the previous VPU walkthrough.
Click the **Writing a Python Script** button below for a demonstration.
<span class="graffiti-highlight graffiti-id_i2z9e6u-id_yqhna2v"><i></i><button>Writing a Python Script</button></span>
```
%%writefile load_model_to_device.py
import time
from openvino.inference_engine import IENetwork
from openvino.inference_engine import IEPlugin
import argparse
def main(args):
model=args.model_path
model_weights=model+'.bin'
model_structure=model+'.xml'
start=time.time()
model=IENetwork(model_structure, model_weights)
plugin = IEPlugin(device=args.device)
net = plugin.load(network=model, num_requests=1)
print(f"Time taken to load model = {time.time()-start} seconds")
if __name__=='__main__':
parser=argparse.ArgumentParser()
parser.add_argument('--model_path', required=True)
parser.add_argument('--device', default=None)
args=parser.parse_args()
main(args)
```
## Step 2: Creating a Job Submission Script
To submit a job to the DevCloud, we need to create a shell script. Similar to the Python script above, I have used the `%%writefile` magic command to create a shell script called `load_multi_model_job.sh`.
This script does a few things.
1. Writes stdout and stderr to their respective .log files
2. Creates the `/output` directory
3. Creates `DEVICE ` and `MODELPATH` variables and assigns their value as the first and second argument passed to the shell script
4. Calls the Python script using the `MODELPATH` and `DEVICE` variable values as the command line argument
5. Changes to the `/output` directory
6. Compresses the stdout.log and stderr.log files to `output.tgz`
**Note**: Just like our Python script, our job submission script also does not need to change when using the **Multi device plugin**. Step 3, where we submit our job to the DevCloud, is where we have to make a minor change.
Click the **Creating a Job Submission Script** button below for a demonstration.
<span class="graffiti-highlight graffiti-id_2c3xj24-id_z7gc9v1"><i></i><button>Creating a Job Submission Script</button></span>
```
%%writefile load_multi_model_job.sh
exec 1>/output/stdout.log 2>/output/stderr.log
mkdir -p /output
DEVICE=$1
MODELPATH=$2
# Run the load model python script
python3 load_model_to_device.py --model_path ${MODELPATH} --device ${DEVICE}
cd /output
tar zcvf output.tgz stdout.log stderr.log
```
## Step 3: Submitting a Job to Intel's DevCloud
The code below will submit a job to an **IEI Tank-870** edge node with the following three devices:
* **Intel Core i5 6500TE**
* **Intel HD Graphics 530**
* **Intel Neural Compute Stick 2**
**Note**: We'll pass in a device type argument of `MULTI:MYRIAD,GPU,CPU` to load our model on all three devices at the same time. We'll need to use `FP16` as the model precision since we're loading our model on a GPU and VPU even though the recommended model precison is `FP32` for CPU.
The `!qsub` command takes a few command line arguments:
1. The first argument is the shell script filename - `load_multi_model_job.sh`. This should always be the first argument.
2. The `-d` flag designates the directory where we want to run our job. We'll be running it in the current directory as denoted by `.`.
3. The `-l` flag designates the node and quantity we want to request. The default quantity is 1, so the **1** after `nodes` is optional.
4. The `-F` flag let's us pass in a string with all command line arguments we want to pass to our Python script.
**Note**: There is an optional flag, `-N`, you may see in a few exercises. This is an argument that only works on Intel's DevCloud that allows you to name your job submission. This argument doesn't work in Udacity's workspace integration with Intel's DevCloud.
In the cell below, we assign the returned value of the `!qsub` command to a variable `job_id_core`. This value is an array with a single string.
Once the cell is run, this queues up a job on Intel's DevCloud and prints out the first value of this array below the cell, which is the job id.
Click the **Submitting a Job to Intel's DevCloud** button below for a demonstration.
<span class="graffiti-highlight graffiti-id_hox95hs-id_9pd5z9z"><i></i><button>Submitting a Job to Intel's DevCloud</button></span>
```
job_id_core = !qsub load_multi_model_job.sh -d . -l nodes=1:tank-870:i5-6500te:intel-hd-530:intel-ncs2 -F "MULTI:MYRIAD,GPU,CPU /data/models/intel/vehicle-license-plate-detection-barrier-0106/FP16/vehicle-license-plate-detection-barrier-0106" -N store_core
print(job_id_core[0])
```
## Step 4: Running liveQStat
Running the `liveQStat` function, we can see the live status of our job. Running the this function will lock the cell and poll the job status 10 times. The cell is locked until this finishes polling 10 times or you can interrupt the kernel to stop it by pressing the stop button at the top: 
* `Q` status means our job is currently awaiting an available node
* `R` status means our job is currently running on the requested node
**Note**: In the demonstration, it is pointed out that `W` status means your job is done. This is no longer accurate. Once a job has finished running, it will no longer show in the list when running the `liveQStat` function.
Click the **Running liveQStat** button below for a demonstration.
<span class="graffiti-highlight graffiti-id_lnsl6m2-id_lauyzu5"><i></i><button>Running LiveQStat</button></span>
```
import liveQStat
liveQStat.liveQStat()
```
## Step 5: Retrieving Output Files
In this step, we'll be using the `getResults` function to retrieve our job's results. This function takes a few arguments.
1. `job id` - This value is stored in the `job_id_core` variable we created during **Step 3**. Remember that this value is an array with a single string, so we access the string value using `job_id_core[0]`.
2. `filename` - This value should match the filename of the compressed file we have in our `load_multi_model_job.sh` shell script. In this example, filename shoud be set to `output.tgz`.
3. `blocking` - This is an optional argument and is set to `False` by default. If this is set to `True`, the cell is locked while waiting for the results to come back. There is a status indicator showing the cell is waiting on results.
**Note**: The `getResults` function is unique to Udacity's workspace integration with Intel's DevCloud. When working on Intel's DevCloud environment, your job's results are automatically retrieved and placed in your working directory.
Click the **Retrieving Output Files** button below for a demonstration.
<span class="graffiti-highlight graffiti-id_v3k1sjd-id_emzwj1d"><i></i><button>Retrieving Output Files</button></span>
```
import get_results
get_results.getResults(job_id_core[0], filename="output.tgz", blocking=True)
```
## Step 6: Viewing the Outputs
In this step, we unpack the compressed file using `!tar zxf` and read the contents of the log files by using the `!cat` command.
`stdout.log` should contain the printout of the print statement in our Python script.
```
!tar zxf output.tgz
!cat stdout.log
!cat stderr.log
```
| github_jupyter |
```
import tensorflow as tf
import cv2
import functools
import json
import math
import matplotlib.pyplot as plt
import numpy as np
import os
import random
import time
import xml.etree.ElementTree as ET
import yaml
from object_detection.utils import dataset_util
from PIL import Image
from PIL import ImageDraw
from PIL import ImageColor
from PIL import ImageFilter
from scipy.stats import norm
from shapely.geometry import Polygon
from shapely import affinity
%matplotlib inline
plt.style.use('ggplot')
def adjust_gamma(img, gamma=1.0):
inv_gamma = 1.0 / gamma
table = np.array([
((i / 255.0) ** inv_gamma) * 255
for i in np.arange(0, 256)])
return cv2.LUT(img.astype(np.uint8), table.astype(np.uint8))
def adjust_contrast(img):
clahe = cv2.createCLAHE(clipLimit=4.0, tileGridSize=(6,6))
img = cv2.cvtColor(img, cv2.COLOR_RGB2LAB)
img[:,:,0] = clahe.apply(img[:,:,0])
img = cv2.cvtColor(img, cv2.COLOR_LAB2RGB)
return img
def preprocess(img, gamma):
img = adjust_contrast(img)
img = adjust_gamma(img, gamma)
return img
```
## Create synthetic training data
```
sim_backgrounds = [
'synthetic-data/backgrounds/sim/bg-1.jpg',
'synthetic-data/backgrounds/sim/bg-2.jpg',
'synthetic-data/backgrounds/sim/bg-3.jpg',
'synthetic-data/backgrounds/sim/bg-4.jpg',
'synthetic-data/backgrounds/sim/bg-5.jpg',
'synthetic-data/backgrounds/sim/bg-6.jpg',
'synthetic-data/backgrounds/sim/bg-7.jpg',
'synthetic-data/backgrounds/sim/bg-8.jpg',
'synthetic-data/backgrounds/sim/bg-9.jpg',
'synthetic-data/backgrounds/sim/bg-10.jpg',
'synthetic-data/backgrounds/sim/bg-11.jpg',
'synthetic-data/backgrounds/sim/bg-12.jpg',
'synthetic-data/backgrounds/sim/bg-13.jpg',
'synthetic-data/backgrounds/sim/bg-14.jpg',
'synthetic-data/backgrounds/sim/bg-15.jpg',
'synthetic-data/backgrounds/sim/bg-16.jpg',
'synthetic-data/backgrounds/sim/bg-17.jpg',
'synthetic-data/backgrounds/sim/bg-18.jpg',
'synthetic-data/backgrounds/sim/bg-19.jpg',
'synthetic-data/backgrounds/sim/bg-20.jpg',
'synthetic-data/backgrounds/sim/bg-21.jpg',
'synthetic-data/backgrounds/sim/bg-22.jpg',
'synthetic-data/backgrounds/sim/bg-23.jpg',
'synthetic-data/backgrounds/sim/bg-24.jpg',
'synthetic-data/backgrounds/sim/bg-25.jpg',
'synthetic-data/backgrounds/sim/bg-26.jpg',
'synthetic-data/backgrounds/sim/bg-27.jpg',
'synthetic-data/backgrounds/sim/bg-28.jpg',
'synthetic-data/backgrounds/sim/bg-29.jpg',
'synthetic-data/backgrounds/sim/bg-30.jpg'
]
sim_red = [
'synthetic-data/elements/sim/red-large.jpg',
'synthetic-data/elements/sim/red-medium.jpg',
'synthetic-data/elements/sim/red-small.jpg'
]
sim_green = [
'synthetic-data/elements/sim/green-large.jpg',
'synthetic-data/elements/sim/green-medium.jpg',
'synthetic-data/elements/sim/green-small.jpg'
]
sim_yellow = [
'synthetic-data/elements/sim/yellow-large.jpg',
'synthetic-data/elements/sim/yellow-medium.jpg',
'synthetic-data/elements/sim/yellow-small.jpg'
]
site_backgrounds = [
# 'synthetic-data/backgrounds/site/bg-1.jpg',
# 'synthetic-data/backgrounds/site/bg-2.jpg',
# 'synthetic-data/backgrounds/site/bg-3.jpg',
'synthetic-data/backgrounds/site/bg-4.jpg',
'synthetic-data/backgrounds/site/bg-5.jpg',
'synthetic-data/backgrounds/site/bg-6.jpg',
'synthetic-data/backgrounds/site/bg-7.jpg',
'synthetic-data/backgrounds/site/bg-8.jpg',
'synthetic-data/backgrounds/site/bg-9.jpg',
'synthetic-data/backgrounds/site/bg-10.jpg',
'synthetic-data/backgrounds/site/bg-11.jpg',
'synthetic-data/backgrounds/site/bg-12.jpg',
'synthetic-data/backgrounds/site/bg-13.jpg',
'synthetic-data/backgrounds/site/bg-14.jpg',
'synthetic-data/backgrounds/site/bg-15.jpg',
'synthetic-data/backgrounds/site/bg-16.jpg',
'synthetic-data/backgrounds/site/bg-17.png',
'synthetic-data/backgrounds/site/bg-18.png',
'synthetic-data/backgrounds/site/bg-19.png',
'synthetic-data/backgrounds/site/bg-20.png',
'synthetic-data/backgrounds/site/bg-21.png',
'synthetic-data/backgrounds/site/bg-22.png',
'synthetic-data/backgrounds/site/bg-23.png',
'synthetic-data/backgrounds/site/bg-24.png',
'synthetic-data/backgrounds/site/bg-25.png',
'synthetic-data/backgrounds/site/bg-26.png',
'synthetic-data/backgrounds/site/bg-27.png',
'synthetic-data/backgrounds/site/bg-28.png',
]
site_red = [
# 'synthetic-data/elements/site/red-1.jpg',
# 'synthetic-data/elements/site/red-2.jpg',
# 'synthetic-data/elements/site/red-3.jpg',
# 'synthetic-data/elements/site/red-4.jpg',
'synthetic-data/elements/site/red-5.jpg',
'synthetic-data/elements/site/red-6.jpg',
'synthetic-data/elements/site/red-7.jpg',
'synthetic-data/elements/site/red-8.jpg',
'synthetic-data/elements/site/red-9.jpg',
'synthetic-data/elements/site/red-10.jpg',
'synthetic-data/elements/site/red-11.jpg',
'synthetic-data/elements/site/red-12.jpg',
'synthetic-data/elements/site/red-13.jpg',
'synthetic-data/elements/site/red-14.jpg',
'synthetic-data/elements/site/red-15.png',
'synthetic-data/elements/site/red-16.png',
'synthetic-data/elements/site/red-17.png',
'synthetic-data/elements/site/red-18.png',
'synthetic-data/elements/site/red-19.png',
'synthetic-data/elements/site/red-20.png',
'synthetic-data/elements/site/red-21.png',
'synthetic-data/elements/site/red-22.png',
'synthetic-data/elements/site/red-23.png',
'synthetic-data/elements/site/red-24.png',
'synthetic-data/elements/site/red-25.png',
'synthetic-data/elements/site/red-26.png',
'synthetic-data/elements/site/red-27.png',
'synthetic-data/elements/site/red-28.png',
'synthetic-data/elements/site/red-29.png',
'synthetic-data/elements/site/red-30.png',
'synthetic-data/elements/site/red-31.png',
'synthetic-data/elements/site/red-32.png',
'synthetic-data/elements/site/red-33.png',
'synthetic-data/elements/site/red-34.png',
]
site_green = [
# 'synthetic-data/elements/site/green-1.jpg',
# 'synthetic-data/elements/site/green-2.jpg',
# 'synthetic-data/elements/site/green-3.jpg',
'synthetic-data/elements/site/green-4.jpg',
'synthetic-data/elements/site/green-5.jpg',
'synthetic-data/elements/site/green-6.jpg',
'synthetic-data/elements/site/green-7.jpg',
'synthetic-data/elements/site/green-8.jpg',
'synthetic-data/elements/site/green-9.jpg',
'synthetic-data/elements/site/green-10.jpg',
'synthetic-data/elements/site/green-11.jpg',
'synthetic-data/elements/site/green-12.jpg',
'synthetic-data/elements/site/green-13.png',
'synthetic-data/elements/site/green-14.png',
'synthetic-data/elements/site/green-15.png',
'synthetic-data/elements/site/green-16.png',
'synthetic-data/elements/site/green-17.png',
'synthetic-data/elements/site/green-18.png',
'synthetic-data/elements/site/green-19.png',
'synthetic-data/elements/site/green-20.png',
'synthetic-data/elements/site/green-21.png',
'synthetic-data/elements/site/green-22.png',
'synthetic-data/elements/site/green-23.png',
'synthetic-data/elements/site/green-24.png',
'synthetic-data/elements/site/green-25.png',
'synthetic-data/elements/site/green-26.png',
'synthetic-data/elements/site/green-27.png',
'synthetic-data/elements/site/green-28.png',
'synthetic-data/elements/site/green-29.png',
'synthetic-data/elements/site/green-30.png',
'synthetic-data/elements/site/green-31.png',
'synthetic-data/elements/site/green-32.png',
'synthetic-data/elements/site/green-33.png',
'synthetic-data/elements/site/green-34.png',
'synthetic-data/elements/site/green-35.png',
'synthetic-data/elements/site/green-36.png',
'synthetic-data/elements/site/green-37.png',
]
site_yellow = [
# 'synthetic-data/elements/site/yellow-1.jpg',
# 'synthetic-data/elements/site/yellow-2.jpg',
# 'synthetic-data/elements/site/yellow-3.jpg',
'synthetic-data/elements/site/yellow-4.jpg',
'synthetic-data/elements/site/yellow-5.jpg',
'synthetic-data/elements/site/yellow-6.jpg',
'synthetic-data/elements/site/yellow-7.jpg',
'synthetic-data/elements/site/yellow-8.jpg',
'synthetic-data/elements/site/yellow-9.jpg',
'synthetic-data/elements/site/yellow-10.jpg',
'synthetic-data/elements/site/yellow-11.jpg',
'synthetic-data/elements/site/yellow-12.png',
'synthetic-data/elements/site/yellow-13.png',
'synthetic-data/elements/site/yellow-14.png',
'synthetic-data/elements/site/yellow-15.png',
'synthetic-data/elements/site/yellow-16.png',
'synthetic-data/elements/site/yellow-17.png',
'synthetic-data/elements/site/yellow-18.png',
'synthetic-data/elements/site/yellow-19.png',
'synthetic-data/elements/site/yellow-20.png',
'synthetic-data/elements/site/yellow-21.png',
'synthetic-data/elements/site/yellow-22.png',
'synthetic-data/elements/site/yellow-23.png',
'synthetic-data/elements/site/yellow-24.png',
'synthetic-data/elements/site/yellow-25.png',
'synthetic-data/elements/site/yellow-26.png',
'synthetic-data/elements/site/yellow-27.png',
'synthetic-data/elements/site/yellow-28.png',
'synthetic-data/elements/site/yellow-29.png',
'synthetic-data/elements/site/yellow-30.png',
'synthetic-data/elements/site/yellow-31.png',
'synthetic-data/elements/site/yellow-32.png',
'synthetic-data/elements/site/yellow-33.png',
'synthetic-data/elements/site/yellow-34.png',
'synthetic-data/elements/site/yellow-35.png',
]
def create_object(wrapper, label, bounding_box):
obj = ET.SubElement(wrapper, 'object')
name = ET.SubElement(obj, 'name')
name.text = label
pose = ET.SubElement(obj, 'pose')
pose.text = 'Unspecified'
truncated = ET.SubElement(obj, 'truncated')
truncated.text = str(0)
difficult = ET.SubElement(obj, 'difficult')
difficult.text = str(0)
bndbox = ET.SubElement(obj, 'bndbox')
xmin = ET.SubElement(bndbox, 'xmin')
xmin.text = str(bounding_box[0])
ymin = ET.SubElement(bndbox, 'ymin')
ymin.text = str(bounding_box[1])
xmax = ET.SubElement(bndbox, 'xmax')
xmax.text = str(bounding_box[2])
ymax = ET.SubElement(bndbox, 'ymax')
ymax.text = str(bounding_box[3])
def create_xml(name, width, height, bounding_boxes, labels):
annotation = ET.Element('annotation')
filename = ET.SubElement(annotation, 'filename')
filename.text = name
size = ET.SubElement(annotation, 'size')
w = ET.SubElement(size, 'width')
w.text = str(width)
h = ET.SubElement(size, 'height')
h.text = str(height)
depth = ET.SubElement(size, 'depth')
depth.text = str(3)
segmented = ET.SubElement(annotation, 'segmented')
segmented.text = str(0)
for i in range(len(bounding_boxes)):
create_object(annotation, labels[i], bounding_boxes[i])
return annotation
def find_overlapping_bounding_box(bounding_boxes, new_bounding_box):
max_intersection = 0
for i in range(len(bounding_boxes)):
min_x, min_y, max_x, max_y = bounding_boxes[i]
new_min_x, new_min_y, new_max_x, new_max_y = new_bounding_box
# print(new_bounding_box)
existing_bb = Polygon([
(min_x, min_y),
(max_x, min_y),
(max_x, max_y),
(min_x, max_y)
])
new_bb= Polygon([
(new_min_x, new_min_y),
(new_max_x, new_min_y),
(new_max_x, new_max_y),
(new_min_x, new_max_y)
])
intersection = existing_bb.intersection(new_bb)
area_of_overlap = intersection.area / new_bb.area if new_bb.area < existing_bb.area else intersection.area / existing_bb.area
if area_of_overlap > max_intersection:
max_intersection = area_of_overlap
return max_intersection
def rotate_bb(bb, degrees):
min_x, min_y, max_x, max_y = bb
poly = Polygon([
(min_x, min_y),
(max_x, min_y),
(max_x, max_y),
(min_x, max_y)
])
modified = affinity.rotate(poly, degrees)
min_x, min_y, max_x, max_y = modified.bounds
return [int(min_x), int(max_y), int(max_x), int(min_y)]
def rescale_bb(bb, ratio):
min_x, min_y, max_x, max_y = bb
poly = Polygon([
(min_x, min_y),
(max_x, min_y),
(max_x, max_y),
(min_x, max_y)
])
modified = affinity.scale(poly, xfact=ratio, yfact=ratio)
min_x, min_y, max_x, max_y = modified.bounds
return [int(min_x), int(max_y), int(max_x), int(min_y)]
def add_to_image(bg, img_set, num, other_bounding_boxes):
bounding_boxes = []
for num_light in range(random.randint(1, num)):
# 0 = large, 1 = medium, 2 = small
img_num = random.randint(0, len(img_set) - 1)
synth_image = Image.open(img_set[img_num]).convert('RGBA')
# 50 percent chance of flipping the image horizontally
flip_chance = random.uniform(0.0, 1.0)
if flip_chance > 0.5:
synth_image.transpose(Image.FLIP_LEFT_RIGHT)
# get the element dimensions
light_width, light_height = synth_image.size
bg_width, bg_height = bg.size
# random rotation/rescale rtios
degrees = random.uniform(-5.0, 5.0)
ratio = random.uniform(0.9, 1.8)
# if img_num == 0:
# # large images
# ratio = random.uniform(0.5, 1.5)
# elif img_num == 1:
# # medium images
# ratio = random.uniform(0.5, 1.5)
# else:
# # small images
# ratio = random.uniform(0.7, 1.0)
while True:
upper_left_x = random.randint(-light_width / 2, bg_width - light_width / 2)
upper_left_y = random.randint(-light_width / 2, bg_height + light_height / 2 - float(bg_height) * 0.4)
bounding_box = rescale_bb(rotate_bb([
upper_left_x,
upper_left_y + light_height,
upper_left_x + light_width,
upper_left_y
], degrees), ratio)
overlap = find_overlapping_bounding_box(bounding_boxes + other_bounding_boxes, bounding_box)
# check to make sure this isn't osverlapping an existing image, and isn't over 50% off screen
if overlap < 0.15 and bounding_box[3] < bg.size[1] - light_height / 2 and bounding_box[2] < bg.size[0] - light_width / 2:
break
# apply random rotation
rotated_image = synth_image.rotate(degrees, expand=1)
# apply random resizing
rescaled_image = rotated_image.resize(
(int(rotated_image.width * ratio), int(rotated_image.height * ratio)),
resample=Image.BICUBIC
)
final_image = rescaled_image
final_light_width, final_light_height = final_image.size
# apply random gaussian blur to larger images
# print('final_light_height', final_light_height)
if final_light_height > 150:
final_image = rescaled_image.filter(ImageFilter.GaussianBlur(radius=random.randint(0,1)))
bg.paste(
final_image,
(bounding_box[0], bounding_box[3]),
final_image
)
# overlay = Image.open(site_overlays[random.randint(0, len(site_overlays) - 1)])
# bg = Image.blend(bg, overlay, alpha=random.uniform(0.0, 0.2))
# bounding box is in the format xmin, ymin, xmax, ymax
bounding_boxes.append(bounding_box)
return bg, bounding_boxes
# at most we want the light half-obscured
# currently this outputs images and labels to a directory called output/ (model-playground/output/)
all_images = []
all_labels = []
entries = []
dest_folder = './evaluation-dataset/site/'
yaml_path = './evaluation-dataset/site/images.yaml'
backgrounds = site_backgrounds
green = site_green
yellow = site_yellow
red = site_red
num_backgrounds = len(backgrounds)
for i in range(2000):
if i % 100 == 0:
print('creating image ' + str(i))
bg = Image.open(backgrounds[random.randint(0, num_backgrounds - 1)])
filename = 'synthetic-' + str(i)
bounding_boxes = []
labels = []
bg, new_bounding_boxes = add_to_image(bg, green, 3, bounding_boxes)
for i in range(len(new_bounding_boxes)):
labels.append('Green_light')
bounding_boxes.append(new_bounding_boxes[i])
bg, new_bounding_boxes = add_to_image(bg, yellow, 3, bounding_boxes)
for i in range(len(new_bounding_boxes)):
labels.append('Yellow_light')
bounding_boxes.append(new_bounding_boxes[i])
bg, new_bounding_boxes = add_to_image(bg, red, 3, bounding_boxes)
for i in range(len(new_bounding_boxes)):
labels.append('Red_light')
bounding_boxes.append(new_bounding_boxes[i])
bg_width, bg_height = bg.size
annotation = create_xml(filename + '.jpg', bg_width, bg_height, bounding_boxes, labels)
ET.ElementTree(annotation).write(dest_folder + filename + '.xml')
image_filename = dest_folder + filename + '.jpg'
image = np.asarray(bg)
image = preprocess(np.asarray(bg), random.uniform(0.3, 0.6))
bg = Image.fromarray(image)
bg.save(image_filename)
# everything below is for writing yaml file for the tfrecord conversion
all_labels.append([bounding_boxes, labels])
entry = {
'path': '',
'boxes': []
}
entry['path'] = image_filename
for i in range(len(bounding_boxes)):
x_min, y_min, x_max, y_max = bounding_boxes[i]
entry['boxes'].append({
'label': labels[i],
'x_min': x_min,
'x_max': x_max,
# reverse y min and max because the Y axis flips
'y_min': y_max,
'y_max': y_min,
})
entries.append(entry)
print('Done with images and labels')
with open(yaml_path, 'w') as file:
yaml.dump(entries, file)
print('Done with yaml')
# plt.imshow(bg)
```
# Analysis of content of existing database
```
directoryGreen = "./alex-data/simulator_dataset_rgb/Green/labels/"
directoryYellow = "./alex-data/simulator_dataset_rgb/Yellow/labels/"
directoryRed = "./alex-data/simulator_dataset_rgb/Red/labels/"
directory = directoryRed
with open("evaluation_simulator_training_dataset.csv", "a") as myfile:
for filename in os.listdir(directory):
if filename.endswith(".xml"):
root = ET.parse(directory+filename).getroot()
for obj in root.findall('object'):
myfile.write(obj.find('name').text + ",")
bbox = obj.find('bndbox')
myfile.write(bbox.find('xmin').text + ",")
myfile.write(bbox.find('xmax').text + ",")
myfile.write(bbox.find('ymin').text + ",")
myfile.write(bbox.find('ymax').text)
myfile.write("\n")
```
### Create TFRecord
```
INPUT_YAML = "./evaluation-dataset/site/images.yaml"
TFRECORD_DESTINATION = './evaluation-dataset/site/images.record'
LABEL_DICT = {
"Green_light" : 1,
"Red_light" : 2,
"Yellow_light" : 3,
}
def create_tf_record(example):
height = 600 # Image height
width = 800 # Image width
filename = example['path'] # Filename of the image. Empty if image is not from file
filename = filename.encode()
with tf.gfile.GFile(example['path'], 'rb') as fid:
encoded_image = fid.read()
image_format = 'jpg'.encode()
xmins = [] # List of normalized left x coordinates in bounding box (1 per box)
xmaxs = [] # List of normalized right x coordinates in bounding box
# (1 per box)
ymins = [] # List of normalized top y coordinates in bounding box (1 per box)
ymaxs = [] # List of normalized bottom y coordinates in bounding box
# (1 per box)
classes_text = [] # List of string class name of bounding box (1 per box)
classes = [] # List of integer class id of bounding box (1 per box)
for box in example['boxes']:
#if box['occluded'] is False:
#print("adding box")
xmins.append(float(float(box['x_min']) / float(width)))
xmaxs.append(float(float(box['x_max']) / float(width)))
ymins.append(float(float(box['y_min']) / float(height)))
ymaxs.append(float(float(box['y_max']) / float(height)))
classes_text.append(box['label'].encode())
classes.append(int(LABEL_DICT[box['label']]))
tf_record = tf.train.Example(features=tf.train.Features(feature={
'image/height': dataset_util.int64_feature(height),
'image/width': dataset_util.int64_feature(width),
'image/filename': dataset_util.bytes_feature(filename),
'image/source_id': dataset_util.bytes_feature(filename),
'image/encoded': dataset_util.bytes_feature(encoded_image),
'image/format': dataset_util.bytes_feature(image_format),
'image/object/bbox/xmin': dataset_util.float_list_feature(xmins),
'image/object/bbox/xmax': dataset_util.float_list_feature(xmaxs),
'image/object/bbox/ymin': dataset_util.float_list_feature(ymins),
'image/object/bbox/ymax': dataset_util.float_list_feature(ymaxs),
'image/object/class/text': dataset_util.bytes_list_feature(classes_text),
'image/object/class/label': dataset_util.int64_list_feature(classes),
}))
return tf_record
print(tf)
writer = tf.python_io.TFRecordWriter(TFRECORD_DESTINATION)
print("Reading yaml...")
examples = yaml.load(open(INPUT_YAML, 'rb').read())
len_examples = len(examples)
print("Loaded " + str(len(examples)) + " examples...")
for i in range(2000):
examples[i]['path'] = os.path.abspath(examples[i]['path'])
example = examples[i]
# print(example)
tf_record = create_tf_record(example)
writer.write(tf_record.SerializeToString())
writer.close()
print("Done!")
```
### Test detection code
```
# the following function is to add data in a dictionary in order to analyse the detection boxes based on the bbox size
INDEX_DETECT_CORRECTLY = 0
INDEX_DETECT_INCORRECTLY = 1
INDEX_NOT_DETECTED = 2
INDEX_TOTAL_SAMPLE = 3
def add_bbox_sample(dictionary, bbox_size, light_color_id, is_detected, is_detected_correctly):
if not ((bbox_size,light_color_id) in dictionary):
dictionary.update({(bbox_size,light_color_id):[0,0,0,0]})
#update
if is_detected:
if is_detected_correctly:
dictionary[(bbox_size,light_color_id)][INDEX_DETECT_CORRECTLY] += 1
else:
dictionary[(bbox_size,light_color_id)][INDEX_DETECT_INCORRECTLY] += 1
else:
dictionary[(bbox_size,light_color_id)][INDEX_NOT_DETECTED] += 1
dictionary[(bbox_size,light_color_id)][INDEX_TOTAL_SAMPLE] += 1
def load_graph(graph_file):
"""Loads a frozen inference graph"""
graph = tf.Graph()
with graph.as_default():
od_graph_def = tf.GraphDef()
with tf.gfile.GFile(graph_file, 'rb') as fid:
serialized_graph = fid.read()
od_graph_def.ParseFromString(serialized_graph)
tf.import_graph_def(od_graph_def, name='')
return graph
def filter_boxes(min_score, boxes, scores, classes):
"""Return boxes with a confidence >= `min_score`"""
n = len(classes)
idxs = []
for i in range(n):
if scores[i] >= min_score:
idxs.append(i)
filtered_boxes = boxes[idxs, ...]
filtered_scores = scores[idxs, ...]
filtered_classes = classes[idxs, ...]
return filtered_boxes, filtered_scores, filtered_classes
def to_image_coords(boxes, height, width):
"""
The original box coordinate output is normalized, i.e [0, 1].
This converts it back to the original coordinate based on the image
size.
"""
box_coords = np.zeros_like(boxes)
box_coords[:, 0] = boxes[:, 0] * height
box_coords[:, 1] = boxes[:, 1] * width
box_coords[:, 2] = boxes[:, 2] * height
box_coords[:, 3] = boxes[:, 3] * width
return box_coords
def draw_boxes(image, boxes, classes, colors, thickness=3):
"""Draw bounding boxes on the image"""
draw = ImageDraw.Draw(image)
for i in range(len(boxes)):
bot, left, top, right = boxes[i, ...]
class_id = int(classes[i])
color = colors[class_id - 1]
draw.line([(left, top), (left, bot), (right, bot), (right, top), (left, top)], width=thickness, fill=color)
detection_graph = load_graph('../ros/src/tl_detector/light_classification/site/ssd_inception_v2_coco_17000_gamma_new/frozen_inference_graph.pb')
# The input placeholder for the image.
# `get_tensor_by_name` returns the Tensor with the associated name in the Graph.
image_tensor = detection_graph.get_tensor_by_name('image_tensor:0')
# Each box represents a part of the image where a particular object was detected.
detection_boxes = detection_graph.get_tensor_by_name('detection_boxes:0')
# Each score represent how level of confidence for each of the objects.
# Score is shown on the result image, together with the class label.
detection_scores = detection_graph.get_tensor_by_name('detection_scores:0')
# The classification of the object (integer id).
detection_classes = detection_graph.get_tensor_by_name('detection_classes:0')
# image = Image.open('./Site_Training_Test_Dataset/site-green-151.jpg')
image = Image.open('./assets/site-green-102.jpg')
image = np.asarray(image)
image = preprocess(image, 0.4)
# image = np.asarray(image)
# image = adjust_gamma(image, 0.5)
# image = Image.fromarray(image)
image = Image.fromarray(image)
image_np = np.expand_dims(np.asarray(image, dtype=np.uint8), 0)
with tf.Session(graph=detection_graph) as sess:
# Actual detection.
(boxes, scores, classes) = sess.run([detection_boxes, detection_scores, detection_classes],
feed_dict={image_tensor: image_np})
# Remove unnecessary dimensions
boxes = np.squeeze(boxes)
scores = np.squeeze(scores)
classes = np.squeeze(classes)
confidence_cutoff = 0.5
# Filter boxes with a confidence score less than `confidence_cutoff`
boxes, scores, classes = filter_boxes(confidence_cutoff, boxes, scores, classes)
print(scores)
print(classes)
# The current box coordinates are normalized to a range between 0 and 1.
# This converts the coordinates actual location on the image.
width, height = image.size
box_coords = to_image_coords(boxes, height, width)
# Each class with be represented by a differently colored box
draw_boxes(image, box_coords, classes, ['green', 'yellow', 'red', 'gray'])
# print(detection_classes)
# print(classes)
plt.figure(figsize=(12, 8))
plt.imshow(image)
for x in xrange(1,6,1):
print x
import os
#to store in the file where we record data for evaluation
MODEL_NAME = "ssd_inception_v2_coco_17000_gamma_new"
NUM_STEPS = 17000
DATABASE_TRAIN_NAME = "test_train"
DATABASE_EVAL_NAME = "Site_Training_Test_Dataset"
SYNTHETIC_DATA = "No"
evaluation_folder = './evaluation-dataset/site-original-new/'
def distance(x1, y1, x2, y2):
return math.sqrt(pow(x2 - x1, 2) + pow(y2 - y1, 2))
label_mapping = {
'Green_light': 1,
'Red_light': 2,
'Yellow_light': 3
}
# image = preprocess(image)
def evaluate(images, frozen_graph, labels):
# correct is when an expected bounding box is detected and is labelled correctly
correct = {
'1': 0,
'2': 0,
'3': 0,
}
# incorrect is when an expected bounding box is detected and is labelled incorrectly
incorrect = {
'1': 0,
'2': 0,
'3': 0,
}
# non-detected is when an expected bounding box is not detected
not_detected = {
'1': 0,
'2': 0,
'3': 0,
}
# background_as_traffic_light is when we have detected a bounding box but there is not any traffic light (i.e. when the bounding box detected falls into background)
background_as_traffic_light = {
'1': 0,
'2': 0,
'3': 0,
}
total = {
'1': 0,
'2': 0,
'3': 0,
}
bbox_analysis = {}
# The input placeholder for the image.
# `get_tensor_by_name` returns the Tensor with the associated name in the Graph.
image_tensor = frozen_graph.get_tensor_by_name('image_tensor:0')
# Each box represents a part of the image where a particular object was detected.
detection_boxes = frozen_graph.get_tensor_by_name('detection_boxes:0')
# Each score represent how level of confidence for each of the objects.
# Score is shown on the result image, together with the class label.
detection_scores = frozen_graph.get_tensor_by_name('detection_scores:0')
# The classification of the object (integer id).
detection_classes = frozen_graph.get_tensor_by_name('detection_classes:0')
with tf.Session(graph=frozen_graph) as sess:
for idx_image in range(len(images)):
image_name = images[idx_image]
# sample_name = 'site-red-' + str(idx_image)
exists = os.path.isfile(evaluation_folder + image_name + '.jpg')
if not exists:
continue
image = Image.open(evaluation_folder + image_name + '.jpg')
image = np.asarray(image)
image = preprocess(image, 0.4)
image = Image.fromarray(image)
image_np = np.expand_dims(np.asarray(image, dtype=np.uint8), 0)
(boxes, scores, classes) = sess.run([detection_boxes, detection_scores, detection_classes],
feed_dict={image_tensor: image_np})
# Remove unnecessary dimensions
boxes = np.squeeze(boxes)
scores = np.squeeze(scores)
classes = np.squeeze(classes)
confidence_cutoff = 0.5
# Filter boxes with a confidence score less than `confidence_cutoff`
boxes, scores, classes = filter_boxes(confidence_cutoff, boxes, scores, classes)
# print(boxes)
# print(scores)
# The current box coordinates are normalized to a range between 0 and 1.
# This converts the coordinates actual location on the image.
width, height = image.size
box_coords = to_image_coords(boxes, height, width)
exists = os.path.isfile(evaluation_folder + image_name + '.xml')
if not exists:
continue
label_xml = ET.parse(evaluation_folder + image_name + '.xml')
expected_bbs = []
expected_classes = []
correpondent_detected = [False] * len(boxes)
for label in label_xml.findall('object'):
expected = label_mapping[label.find('name').text]
expected_classes.append(expected)
# print(expected)
box = label.find('bndbox')
min_x = int(box.find('xmin').text)
min_y = int(box.find('ymin').text)
max_x = int(box.find('xmax').text)
max_y = int(box.find('ymax').text)
#print(str(min_x) + ", " + str(min_y) + ", " + str(max_x) + ", " + str(max_y))
bbox_size = abs(max_y-min_y)*abs(max_x-min_x)
expected_bbs.append([min_y, min_x, max_y, max_x])
# expected_bb = Polygon([
# (min_y, min_x),
# (max_x, min_y),
# (max_x, max_y),
# (min_x, max_y)
# ])
total[str(expected)] += 1
# print('\ntesting')
# print([max_y, min_x, min_y, max_x])
# print('expected class: ' + str(expected))
# print('\n')
detected = False
for i in range(len(box_coords)):
new_max_y, new_min_x, new_min_y, new_max_x = box_coords[i]
#print(str(new_min_x) + ", " + str(new_min_y) + ", " + str(new_max_x) + ", " + str(new_max_y))
# actual_bb = Polygon([
# (new_min_x, new_min_y),
# (new_max_x, new_min_y),
# (new_max_x, new_max_y),
# (new_min_x, new_max_y)
# ])
# intersection = expected_bb.intersection(actual_bb)
# area_of_overlap = intersection.area / actual_bb.area if actual_bb.area < expected_bb.area else intersection.area / expected_bb.area
# print(box_coords[i])
# print('overlap: ' + str(area_of_overlap))
# consider it a match if there is 75% overlap
# distance of top left corners
#dist_tl = distance(min_x, max_y, new_min_x, new_min_y)
dist_tl = distance(min_x, max_y, new_min_x, new_max_y)
#print(dist_tl)
# print(dist_tl)
# distance of bottom right corners
#dist_br = distance(max_x, min_y, new_max_x, new_max_y)
dist_br = distance(max_x, min_y, new_max_x, new_min_y)
#print(dist_br)
# print(dist_br)
# print('predicted class: ' + str(int(classes[i])))
avg_corner_distance = (dist_tl + dist_br) / 2
#print(avg_corner_distance)
# print('avg corner distance: ' + str(avg_corner_distance))
if avg_corner_distance < 50:# and area_of_overlap > 0.7:
correpondent_detected[i] = True
detected = True
if int(classes[i]) == expected:
#print("correct")
correct[str(expected)] += 1
add_bbox_sample(bbox_analysis, bbox_size, str(expected), True, True)
else:
#print("incorrect")
incorrect[str(expected)] += 1
add_bbox_sample(bbox_analysis, bbox_size, str(expected), True, False)
if not detected:
not_detected[str(expected)] += 1
add_bbox_sample(bbox_analysis, bbox_size, str(expected), False, False)
for i in range(len(box_coords)):
if correpondent_detected[i] is False:
background_as_traffic_light[str(int(classes[i]))] += 1
#d = dict(zip(unique, counts))
#non_correspondance[int(classes[i])] += d[False]
# print(expected_bbs)
# print(np.array(expected_bbs))
# print(box_coords)
# draw_boxes(image, np.array(expected_bbs), expected_classes, ['#c2f6a8', '#f68d8d', '#f6eaa8', 'gray'])
# draw_boxes(image, box_coords, classes, ['green', 'red', 'yellow', 'gray'])
# print(detection_classes)
# print(classes)
# plt.figure(figsize=(12, 8))
# plt.imshow(image)
# print('Note: it is possible that total expected lights is different from the detected correct plus detected incorrect\nsince it is possible that we have two overlapping bounding boxes')
return total, correct, incorrect, not_detected, background_as_traffic_light
# with open("evaluation_model_accuracy_site_13nov_site_real_adj.csv", "a") as myfile:
# for i in range(3): # for all the colors
# myfile.write(MODEL_NAME + ",")
# myfile.write(str(NUM_STEPS) + ",")
# myfile.write(DATABASE_TRAIN_NAME + ",")
# myfile.write(DATABASE_EVAL_NAME + ",")
# myfile.write(SYNTHETIC_DATA + ",")
# myfile.write(str(i+1) + ",") #colour
# myfile.write(str(total[str(i+1)]) + ", ")
# myfile.write(str(correct[str(i+1)]) + ", ")
# myfile.write(str(incorrect[str(i+1)]) + ", ")
# myfile.write(str(not_detected[str(i+1)]) + ", ")
# myfile.write(str(background_as_traffic_light[str(i+1)]))
# myfile.write("\n")
# with open("evaluation_model_accuracy_bbox_site_13nov_site_real_adj.csv", "a") as myfile:
# for x in bbox_analysis.keys():
# myfile.write(MODEL_NAME + ",")
# myfile.write(str(NUM_STEPS) + ",")
# myfile.write(DATABASE_TRAIN_NAME + ",")
# myfile.write(DATABASE_EVAL_NAME + ",")
# myfile.write(str(x[1]) + ",") # traffic light color
# myfile.write(str(x[0]) + ",") # bbox size
# myfile.write(str(bbox_analysis[x][INDEX_DETECT_CORRECTLY]) + ",") # detection correct
# myfile.write(str(bbox_analysis[x][INDEX_DETECT_INCORRECTLY]) + ",") # detection incorrect
# myfile.write(str(bbox_analysis[x][INDEX_NOT_DETECTED]) + ",") # not detected
# myfile.write(str(bbox_analysis[x][INDEX_TOTAL_SAMPLE])) # total samples
# myfile.write("\n")
root_dir = '../ros/src/tl_detector/light_classification/site'
networks = ['ssd_mobilenet_v1_coco_20000_gamma', 'ssd_inception_v2_coco_17000_gamma_new']
colors = [[0.5, 0.5, 0.5], [0.9, 0.8, 0]]
num_images = 1000
image_list = []
for i in range(num_images):
image_list.append('synthetic-' + str(i))
stats = []
for i in range(len(networks)):
network_name = networks[i]
graph_filename = root_dir + '/' + network_name + '/frozen_inference_graph.pb'
total, correct, incorrect, not_detected, background_as_traffic_light = evaluate(
image_list,
load_graph(graph_filename),
label_mapping
)
stats.append({
'network_name': network_name,
'total': total,
'correct': correct,
'background_as_traffic_light': background_as_traffic_light,
'incorrect': incorrect,
'not_detected': not_detected,
'color': colors[i]
})
# print('Green:')
# print('Total expected lights: ' + str(total['1']))
# print('Detected Correct: ' + str(correct['1']))
# print('Detected Incorrect: ' + str(incorrect['1']))
# print('Not detected: ' + str(not_detected['1']))
# print('Background as traffic light: ' + str(background_as_traffic_light['1']))
# print('Accuracy: '+ str(correct['1'] / float(total['1'])))
# print('\nRed:')
# print('Total expected lights: ' + str(total['2']))
# print('Detected Correct: ' + str(correct['2']))
# print('Detected Incorrect: ' + str(incorrect['2']))
# print('Not detected: ' + str(not_detected['2']))
# print('Background as traffic light: ' + str(background_as_traffic_light['2']))
# print('Accuracy: '+ str(correct['2'] / float(total['2'])))
# print('\nYellow:')
# print('Total expected lights: ' + str(total['3']))
# print('Detected Correct: ' + str(correct['3']))
# print('Detected Incorrect: ' + str(incorrect['3']))
# print('Not detected: ' + str(not_detected['3']))
# print('Background as traffic light: ' + str(background_as_traffic_light['3']))
# print('Accuracy: '+ str(correct['3'] / float(total['3'])))
labels = ['Red', 'Yellow', 'Green']
x = np.arange(len(labels))
y = [0.0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 1.1, 1.2, 1.3]
bar_width = 0.25
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2, 2)
# plot accuracy
for i in range(len(stats)):
network_name = stats[i]['network_name']
correct = stats[i]['correct']
total = stats[i]['total']
color = stats[i]['color']
ax1.bar(x + bar_width * i - bar_width / 2, [
correct['1'] / float(total['1']),
correct['3'] / float(total['3']),
correct['2'] / float(total['2'])
], bar_width, label=network_name, color=color)
ax1.set_ylabel('Correct %')
ax1.set_title('Network accuracy by color')
# plot not_detected
for i in range(len(stats)):
network_name = stats[i]['network_name']
correct = stats[i]['correct']
total = stats[i]['total']
not_detected = stats[i]['not_detected']
color = stats[i]['color']
ax2.bar(x + bar_width * i - bar_width / 2, [
not_detected['1'] / float(total['1']),
not_detected['3'] / float(total['3']),
not_detected['2'] / float(total['2'])
], bar_width, label=network_name, color=color)
ax2.set_ylabel('Missed %')
ax2.set_title('Missed lights by color')
# plot incorrect
for i in range(len(stats)):
network_name = stats[i]['network_name']
incorrect = stats[i]['incorrect']
total = stats[i]['total']
not_detected = stats[i]['not_detected']
color = stats[i]['color']
ax3.bar(x + bar_width * i - bar_width / 2, [
incorrect['1'] / float(total['1']),
incorrect['3'] / float(total['3']),
incorrect['2'] / float(total['2'])
], bar_width, label=network_name, color=color)
ax3.set_ylabel('Incorrect %')
ax3.set_title('Incorrect lights by color')
# plot background detected as light
for i in range(len(stats)):
network_name = stats[i]['network_name']
incorrect = stats[i]['incorrect']
total = stats[i]['total']
background_as_traffic_light = stats[i]['background_as_traffic_light']
color = stats[i]['color']
ax4.bar(x + bar_width * i - bar_width / 2, [
background_as_traffic_light['1'] / float(total['1']),
background_as_traffic_light['3'] / float(total['3']),
background_as_traffic_light['2'] / float(total['2'])
], bar_width, label=network_name, color=color)
ax4.set_ylabel('Phantom light %')
ax4.set_title('Phantom lights by color')
ax1.set_xticks(x)
ax1.set_yticks(y)
ax1.legend()
ax1.set_xticklabels(labels)
ax2.set_xticks(x)
ax2.set_yticks(y)
ax2.set_xticklabels(labels)
ax2.legend()
ax3.set_xticks(x)
ax3.set_yticks(y)
ax3.set_xticklabels(labels)
ax3.legend()
ax4.set_xticks(x)
ax4.set_yticks(y)
ax4.set_xticklabels(labels)
ax4.legend()
# fig.tight_layout()
fig.set_size_inches(16, 16)
plt.show()
```
# Analysis with different gamma steps
```
#to store in the file where we record data for evaluation
MODEL_NAME = "ssd_mobilenet_v1_coco_20000_gamma_test"
NUM_STEPS = 10000
DATABASE_TRAIN_NAME = "test_train"
DATABASE_EVAL_NAME = "SyntheticDataOriginalGamma"
SYNTHETIC_DATA = "No"
evaluation_folder = './evaluation-dataset/site-original/'
root_dir = '../ros/src/tl_detector/light_classification/site'
graph_filename = root_dir + '/ssd_inception_v2_coco_17000_gamma_new/frozen_inference_graph.pb'
detection_graph = load_graph(graph_filename)
def distance(x1, y1, x2, y2):
return math.sqrt(pow(x2 - x1, 2) + pow(y2 - y1, 2))
# The input placeholder for the image.
# `get_tensor_by_name` returns the Tensor with the associated name in the Graph.
image_tensor = detection_graph.get_tensor_by_name('image_tensor:0')
# Each box represents a part of the image where a particular object was detected.
detection_boxes = detection_graph.get_tensor_by_name('detection_boxes:0')
# Each score represent how level of confidence for each of the objects.
# Score is shown on the result image, together with the class label.
detection_scores = detection_graph.get_tensor_by_name('detection_scores:0')
# The classification of the object (integer id).
detection_classes = detection_graph.get_tensor_by_name('detection_classes:0')
label_mapping = {
'Green_light': 1,
'Red_light': 2,
'Yellow_light': 3
}
# image = preprocess(image)
with tf.Session(graph=detection_graph) as sess:
for i in xrange(30,100,5):
# correct is when an expected bounding box is detected and is labelled correctly
correct = {
'1': 0,
'2': 0,
'3': 0,
}
# incorrect is when an expected bounding box is detected and is labelled incorrectly
incorrect = {
'1': 0,
'2': 0,
'3': 0,
}
# non-detected is when an expected bounding box is not detected
not_detected = {
'1': 0,
'2': 0,
'3': 0,
}
# background_as_traffic_light is when we have detected a bounding box but there is not any traffic light (i.e. when the bounding box detected falls into background)
background_as_traffic_light = {
'1': 0,
'2': 0,
'3': 0,
}
total = {
'1': 0,
'2': 0,
'3': 0,
}
bbox_analysis = {}
gammaValue = i/100.
print('testing gamma ' + str(gammaValue))
for idx_image in range(500):
sample_name = 'synthetic-' + str(idx_image)
image = Image.open(evaluation_folder + sample_name + '.jpg')
image = np.asarray(image)
image = preprocess(image, gammaValue)
image = Image.fromarray(image)
image_np = np.expand_dims(np.asarray(image, dtype=np.uint8), 0)
(boxes, scores, classes) = sess.run([detection_boxes, detection_scores, detection_classes],
feed_dict={image_tensor: image_np})
# Remove unnecessary dimensions
boxes = np.squeeze(boxes)
scores = np.squeeze(scores)
classes = np.squeeze(classes)
confidence_cutoff = 0.4
# Filter boxes with a confidence score less than `confidence_cutoff`
boxes, scores, classes = filter_boxes(confidence_cutoff, boxes, scores, classes)
# print(boxes)
# print(scores)
# The current box coordinates are normalized to a range between 0 and 1.
# This converts the coordinates actual location on the image.
width, height = image.size
box_coords = to_image_coords(boxes, height, width)
label_xml = ET.parse(evaluation_folder + sample_name + '.xml')
expected_bbs = []
expected_classes = []
correpondent_detected = [False] * len(boxes)
for label in label_xml.findall('object'):
expected = label_mapping[label.find('name').text]
expected_classes.append(expected)
# print(expected)
box = label.find('bndbox')
min_x = int(box.find('xmin').text)
min_y = int(box.find('ymin').text)
max_x = int(box.find('xmax').text)
max_y = int(box.find('ymax').text)
bbox_size = abs(max_y-min_y)*abs(max_x-min_x)
expected_bbs.append([min_y, min_x, max_y, max_x])
# expected_bb = Polygon([
# (min_y, min_x),
# (max_x, min_y),
# (max_x, max_y),
# (min_x, max_y)
# ])
total[str(expected)] += 1
# print('\ntesting')
# print([max_y, min_x, min_y, max_x])
# print('expected class: ' + str(expected))
# print('\n')
detected = False
for i in range(len(box_coords)):
new_min_y, new_min_x, new_max_y, new_max_x = box_coords[i]
# actual_bb = Polygon([
# (new_min_x, new_min_y),
# (new_max_x, new_min_y),
# (new_max_x, new_max_y),
# (new_min_x, new_max_y)
# ])
# intersection = expected_bb.intersection(actual_bb)
# area_of_overlap = intersection.area / actual_bb.area if actual_bb.area < expected_bb.area else intersection.area / expected_bb.area
# print(box_coords[i])
# print('overlap: ' + str(area_of_overlap))
# consider it a match if there is 75% overlap
# distance of top left corners
dist_tl = distance(min_x, max_y, new_min_x, new_min_y)
# print(dist_tl)
# distance of bottom right corners
dist_br = distance(max_x, min_y, new_max_x, new_max_y)
# print(dist_br)
# print('predicted class: ' + str(int(classes[i])))
avg_corner_distance = (dist_tl + dist_br) / 2
# print('avg corner distance: ' + str(avg_corner_distance))
if avg_corner_distance < 40:# and area_of_overlap > 0.7:
correpondent_detected[i] = True
detected = True
if int(classes[i]) == expected:
correct[str(expected)] += 1
add_bbox_sample(bbox_analysis, bbox_size, str(expected), True, True)
else:
incorrect[str(expected)] += 1
add_bbox_sample(bbox_analysis, bbox_size, str(expected), True, False)
if not detected:
not_detected[str(expected)] += 1
add_bbox_sample(bbox_analysis, bbox_size, str(expected), False, False)
for i in range(len(box_coords)):
if correpondent_detected[i] is False:
background_as_traffic_light[str(int(classes[i]))] += 1
#d = dict(zip(unique, counts))
#non_correspondance[int(classes[i])] += d[False]
# print(expected_bbs)
# print(np.array(expected_bbs))
# print(box_coords)
# draw_boxes(image, np.array(expected_bbs), expected_classes, ['#c2f6a8', '#f68d8d', '#f6eaa8', 'gray'])
# draw_boxes(image, box_coords, classes, ['green', 'red', 'yellow', 'gray'])
# print(detection_classes)
# print(classes)
# plt.figure(figsize=(12, 8))
# plt.imshow(image)
with open("evaluation_model_accuracy_gamma_25nov.csv", "a") as myfile:
for i in range(3): # for all the colors
myfile.write(MODEL_NAME + ",")
myfile.write(str(NUM_STEPS) + ",")
myfile.write(DATABASE_TRAIN_NAME + ",")
myfile.write(DATABASE_EVAL_NAME + ",")
myfile.write(SYNTHETIC_DATA + ",")
myfile.write(str(i+1) + ",") #colour
myfile.write(str(total[str(i+1)]) + ", ")
myfile.write(str(correct[str(i+1)]) + ", ")
myfile.write(str(incorrect[str(i+1)]) + ", ")
myfile.write(str(not_detected[str(i+1)]) + ", ")
myfile.write(str(background_as_traffic_light[str(i+1)]) + ", ")
myfile.write(str(gammaValue))
myfile.write(str(correct[str(i+1)]/total[str(i+1)]))
myfile.write("\n")
```
# Analsysis of site data
## Timing Detection
The model zoo comes with a variety of models, each its benefits and costs. Below you'll time some of these models. The general tradeoff being sacrificing model accuracy for seconds per frame (SPF).
```
def time_detection(sess, runs=10):
image_tensor = sess.graph.get_tensor_by_name('image_tensor:0')
detection_boxes = sess.graph.get_tensor_by_name('detection_boxes:0')
detection_scores = sess.graph.get_tensor_by_name('detection_scores:0')
detection_classes = sess.graph.get_tensor_by_name('detection_classes:0')
numImages = 100
times = []
for idx_image in range(numImages):
sample_name = 'synthetic-' + str(idx_image)
image = Image.open(evaluation_folder + sample_name + '.jpg')
image_np = np.expand_dims(np.asarray(image, dtype=np.uint8), 0)
for i in range(runs):
t0 = time.time()
sess.run([detection_boxes, detection_scores, detection_classes], feed_dict={image_tensor: image_np})
t1 = time.time()
times.append((t1 - t0) * 1000)
return times
#to store in the file where we record data for evaluation
MODEL_NAME = "ssd_mobilenet_v1_coco_10000"
evaluation_folder = './synthetic-dataset-oct-30-with-labels/'
detection_graph = load_graph('./graphsModels/ssd_mobilenet_v1_coco_10000.pb')
with tf.Session(graph=detection_graph) as sess:
times = time_detection(sess, runs=10)
with open("evaluation_model_object_detection_timings.csv", "a") as myfile:
myfile.write(MODEL_NAME + ",")
myfile.write(str(np.mean(times)) + ",")
myfile.write(str(np.std(times)))
myfile.write("\n")
# Create a figure instance
fig = plt.figure(1, figsize=(9, 6))
# Create an axes instance
ax = fig.add_subplot(111)
plt.title("Object Detection Timings")
plt.ylabel("Time (ms)")
# Create the boxplot
plt.style.use('fivethirtyeight')
bp = ax.boxplot(times)
np.mean(times)
np.std(times)
```
### Exercise 4 - Model Tradeoffs
Download a few models from the [model zoo](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/detection_model_zoo.md) and compare the timings.
## Detection on a Video
Finally run your pipeline on [this short video](https://s3-us-west-1.amazonaws.com/udacity-selfdrivingcar/advanced_deep_learning/driving.mp4).
```
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
HTML("""
<video width="960" height="600" controls>
<source src="{0}" type="video/mp4">
</video>
""".format('driving.mp4'))
```
### Exercise 5 - Object Detection on a Video
Run an object detection pipeline on the above clip.
```
clip = VideoFileClip('driving.mp4')
# TODO: Complete this function.
# The input is an NumPy array.
# The output should also be a NumPy array.
def pipeline(img):
draw_img = Image.fromarray(img)
# Actual detection.
(boxes, scores, classes) = sess.run([detection_boxes, detection_scores, detection_classes],
feed_dict={image_tensor: np.expand_dims(img, 0)})
# Remove unnecessary dimensions
boxes = np.squeeze(boxes)
scores = np.squeeze(scores)
classes = np.squeeze(classes)
confidence_cutoff = 0.3
# Filter boxes with a confidence score less than `confidence_cutoff`
boxes, scores, classes = filter_boxes(confidence_cutoff, boxes, scores, classes)
# The current box coordinates are normalized to a range between 0 and 1.
# This converts the coordinates actual location on the image.
width, height = draw_img.size
box_coords = to_image_coords(boxes, height, width)
# Each class with be represented by a differently colored box
draw_boxes(draw_img, box_coords, classes)
return np.array(draw_img)
```
**[Sample solution](./exercise-solutions/e5.py)**
```
with tf.Session(graph=detection_graph) as sess:
image_tensor = sess.graph.get_tensor_by_name('image_tensor:0')
detection_boxes = sess.graph.get_tensor_by_name('detection_boxes:0')
detection_scores = sess.graph.get_tensor_by_name('detection_scores:0')
detection_classes = sess.graph.get_tensor_by_name('detection_classes:0')
new_clip = clip.fl_image(pipeline)
# write to file
new_clip.write_videofile('result.mp4', audio=False)
HTML("""
<video width="960" height="600" controls>
<source src="{0}" type="video/mp4">
</video>
""".format('result.mp4'))
```
## Further Exploration
Some ideas to take things further:
* Finetune the model on a new dataset more relevant to autonomous vehicles. Instead of loading the frozen inference graph you'll load the checkpoint.
* Optimize the model and get the FPS as low as possible.
* Build your own detector. There are several base model pretrained on ImageNet you can choose from. [Keras](https://keras.io/applications/) is probably the quickest way to get setup in this regard.
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W2D3_BiologicalNeuronModels/student/W2D3_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Tutorial 1: The Leaky Integrate-and-Fire (LIF) Neuron Model
**Week 2, Day 3: Biological Neuron Models**
**By Neuromatch Academy**
__Content creators:__ Qinglong Gu, Songtin Li, John Murray, Richard Naud, Arvind Kumar
__Content reviewers:__ Maryam Vaziri-Pashkam, Ella Batty, Lorenzo Fontolan, Richard Gao, Matthew Krause, Spiros Chavlis, Michael Waskom, Ethan Cheng
**Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
<p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
---
# Tutorial Objectives
*Estimated timing of tutorial: 1 hour, 10 min*
This is Tutorial 1 of a series on implementing realistic neuron models. In this tutorial, we will build up a leaky integrate-and-fire (LIF) neuron model and study its dynamics in response to various types of inputs. In particular, we are going to write a few lines of code to:
- simulate the LIF neuron model
- drive the LIF neuron with external inputs, such as direct currents, Gaussian white noise, and Poisson spike trains, etc.
- study how different inputs affect the LIF neuron's output (firing rate and spike time irregularity)
Here, we will especially emphasize identifying conditions (input statistics) under which a neuron can spike at low firing rates and in an irregular manner. The reason for focusing on this is that in most cases, neocortical neurons spike in an irregular manner.
```
# @title Tutorial slides
# @markdown These are the slides for the videos in all tutorials today
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/8djsm/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
```
---
# Setup
```
# Imports
import numpy as np
import matplotlib.pyplot as plt
# @title Figure Settings
import ipywidgets as widgets # interactive display
%config InlineBackend.figure_format = 'retina'
# use NMA plot style
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
my_layout = widgets.Layout()
# @title Plotting Functions
def plot_volt_trace(pars, v, sp):
"""
Plot trajetory of membrane potential for a single neuron
Expects:
pars : parameter dictionary
v : volt trajetory
sp : spike train
Returns:
figure of the membrane potential trajetory for a single neuron
"""
V_th = pars['V_th']
dt, range_t = pars['dt'], pars['range_t']
if sp.size:
sp_num = (sp / dt).astype(int) - 1
v[sp_num] += 20 # draw nicer spikes
plt.plot(pars['range_t'], v, 'b')
plt.axhline(V_th, 0, 1, color='k', ls='--')
plt.xlabel('Time (ms)')
plt.ylabel('V (mV)')
plt.legend(['Membrane\npotential', r'Threshold V$_{\mathrm{th}}$'],
loc=[1.05, 0.75])
plt.ylim([-80, -40])
def plot_GWN(pars, I_GWN):
"""
Args:
pars : parameter dictionary
I_GWN : Gaussian white noise input
Returns:
figure of the gaussian white noise input
"""
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.plot(pars['range_t'][::3], I_GWN[::3], 'b')
plt.xlabel('Time (ms)')
plt.ylabel(r'$I_{GWN}$ (pA)')
plt.subplot(122)
plot_volt_trace(pars, v, sp)
plt.tight_layout()
def my_hists(isi1, isi2, cv1, cv2, sigma1, sigma2):
"""
Args:
isi1 : vector with inter-spike intervals
isi2 : vector with inter-spike intervals
cv1 : coefficient of variation for isi1
cv2 : coefficient of variation for isi2
Returns:
figure with two histograms, isi1, isi2
"""
plt.figure(figsize=(11, 4))
my_bins = np.linspace(10, 30, 20)
plt.subplot(121)
plt.hist(isi1, bins=my_bins, color='b', alpha=0.5)
plt.xlabel('ISI (ms)')
plt.ylabel('count')
plt.title(r'$\sigma_{GWN}=$%.1f, CV$_{\mathrm{isi}}$=%.3f' % (sigma1, cv1))
plt.subplot(122)
plt.hist(isi2, bins=my_bins, color='b', alpha=0.5)
plt.xlabel('ISI (ms)')
plt.ylabel('count')
plt.title(r'$\sigma_{GWN}=$%.1f, CV$_{\mathrm{isi}}$=%.3f' % (sigma2, cv2))
plt.tight_layout()
plt.show()
```
---
# Section 1: The Leaky Integrate-and-Fire (LIF) model
```
# @title Video 1: Reduced Neuron Models
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="av456396195", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="rSExvwCVRYg", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
This video introduces the reduction of a biological neuron to a simple leaky-integrate-fire (LIF) neuron model.
<details>
<summary> <font color='blue'>Click here for text recap of video </font></summary>
Now, it's your turn to implement one of the simplest mathematical model of a neuron: the leaky integrate-and-fire (LIF) model. The basic idea of LIF neuron was proposed in 1907 by Louis Édouard Lapicque, long before we understood the electrophysiology of a neuron (see a translation of [Lapicque's paper](https://pubmed.ncbi.nlm.nih.gov/17968583/) ). More details of the model can be found in the book [**Theoretical neuroscience**](http://www.gatsby.ucl.ac.uk/~dayan/book/) by Peter Dayan and Laurence F. Abbott.
The subthreshold membrane potential dynamics of a LIF neuron is described by
\begin{eqnarray}
C_m\frac{dV}{dt} = -g_L(V-E_L) + I,\quad (1)
\end{eqnarray}
where $C_m$ is the membrane capacitance, $V$ is the membrane potential, $g_L$ is the leak conductance ($g_L = 1/R$, the inverse of the leak resistance $R$ mentioned in previous tutorials), $E_L$ is the resting potential, and $I$ is the external input current.
Dividing both sides of the above equation by $g_L$ gives
\begin{align}
\tau_m\frac{dV}{dt} = -(V-E_L) + \frac{I}{g_L}\,,\quad (2)
\end{align}
where the $\tau_m$ is membrane time constant and is defined as $\tau_m=C_m/g_L$.
Note that dividing capacitance by conductance gives units of time!
Below, we will use Eqn.(2) to simulate LIF neuron dynamics.
If $I$ is sufficiently strong such that $V$ reaches a certain threshold value $V_{\rm th}$, $V$ is reset to a reset potential $V_{\rm reset}< V_{\rm th}$, and voltage is clamped to $V_{\rm reset}$ for $\tau_{\rm ref}$ ms, mimicking the refractoriness of the neuron during an action potential:
\begin{eqnarray}
\mathrm{if}\quad V(t_{\text{sp}})\geq V_{\rm th}&:& V(t)=V_{\rm reset} \text{ for } t\in(t_{\text{sp}}, t_{\text{sp}} + \tau_{\text{ref}}]
\end{eqnarray}
where $t_{\rm sp}$ is the spike time when $V(t)$ just exceeded $V_{\rm th}$.
(__Note__: in the lecture slides, $\theta$ corresponds to the threshold voltage $V_{th}$, and $\Delta$ corresponds to the refractory time $\tau_{\rm ref}$.)
</details>
Note that you have seen the LIF model before if you looked at the pre-reqs Python or Calculus days!
The LIF model captures the facts that a neuron:
- performs spatial and temporal integration of synaptic inputs
- generates a spike when the voltage reaches a certain threshold
- goes refractory during the action potential
- has a leaky membrane
The LIF model assumes that the spatial and temporal integration of inputs is linear. Also, membrane potential dynamics close to the spike threshold are much slower in LIF neurons than in real neurons.
## Coding Exercise 1: Python code to simulate the LIF neuron
We now write Python code to calculate our equation for the LIF neuron and simulate the LIF neuron dynamics. We will use the Euler method, which you saw in the linear systems case yesterday to numerically integrate this equation:
\begin{align*}
\tau_m\frac{dV}{dt} = -(V-E_L) + \frac{I}{g_L}\,
\end{align*}
where $V$ is the membrane potential, $g_L$ is the leak conductance, $E_L$ is the resting potential, $I$ is the external input current, and $\tau_m$ is membrane time constant.
The cell below initializes a dictionary that stores parameters of the LIF neuron model and the simulation scheme. You can use `pars=default_pars(T=simulation_time, dt=time_step)` to get the parameters. Note that, `simulation_time` and `time_step` have the unit `ms`. In addition, you can add the value to a new parameter by `pars['New_param'] = value`.
```
# @markdown Execute this code to initialize the default parameters
def default_pars(**kwargs):
pars = {}
# typical neuron parameters#
pars['V_th'] = -55. # spike threshold [mV]
pars['V_reset'] = -75. # reset potential [mV]
pars['tau_m'] = 10. # membrane time constant [ms]
pars['g_L'] = 10. # leak conductance [nS]
pars['V_init'] = -75. # initial potential [mV]
pars['E_L'] = -75. # leak reversal potential [mV]
pars['tref'] = 2. # refractory time (ms)
# simulation parameters #
pars['T'] = 400. # Total duration of simulation [ms]
pars['dt'] = .1 # Simulation time step [ms]
# external parameters if any #
for k in kwargs:
pars[k] = kwargs[k]
pars['range_t'] = np.arange(0, pars['T'], pars['dt']) # Vector of discretized time points [ms]
return pars
pars = default_pars()
print(pars)
```
Complete the function below to simulate the LIF neuron when receiving external current inputs. You can use `v, sp = run_LIF(pars, Iinj)` to get the membrane potential (`v`) and spike train (`sp`) given the dictionary `pars` and input current `Iinj`.
```
def run_LIF(pars, Iinj, stop=False):
"""
Simulate the LIF dynamics with external input current
Args:
pars : parameter dictionary
Iinj : input current [pA]. The injected current here can be a value
or an array
stop : boolean. If True, use a current pulse
Returns:
rec_v : membrane potential
rec_sp : spike times
"""
# Set parameters
V_th, V_reset = pars['V_th'], pars['V_reset']
tau_m, g_L = pars['tau_m'], pars['g_L']
V_init, E_L = pars['V_init'], pars['E_L']
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
tref = pars['tref']
# Initialize voltage
v = np.zeros(Lt)
v[0] = V_init
# Set current time course
Iinj = Iinj * np.ones(Lt)
# If current pulse, set beginning and end to 0
if stop:
Iinj[:int(len(Iinj) / 2) - 1000] = 0
Iinj[int(len(Iinj) / 2) + 1000:] = 0
# Loop over time
rec_spikes = [] # record spike times
tr = 0. # the count for refractory duration
for it in range(Lt - 1):
if tr > 0: # check if in refractory period
v[it] = V_reset # set voltage to reset
tr = tr - 1 # reduce running counter of refractory period
elif v[it] >= V_th: # if voltage over threshold
rec_spikes.append(it) # record spike event
v[it] = V_reset # reset voltage
tr = tref / dt # set refractory time
########################################################################
## TODO for students: compute the membrane potential v, spike train sp #
# Fill out function and remove
raise NotImplementedError('Student Exercise: calculate the dv/dt and the update step!')
########################################################################
# Calculate the increment of the membrane potential
dv = ...
# Update the membrane potential
v[it + 1] = ...
# Get spike times in ms
rec_spikes = np.array(rec_spikes) * dt
return v, rec_spikes
# Get parameters
pars = default_pars(T=500)
# Simulate LIF model
v, sp = run_LIF(pars, Iinj=100, stop=True)
# Visualize
plot_volt_trace(pars, v, sp)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial1_Solution_60a1e954.py)
*Example output:*
<img alt='Solution hint' align='left' width=1106.0 height=828.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D3_BiologicalNeuronModels/static/W2D3_Tutorial1_Solution_60a1e954_0.png>
---
# Section 2: Response of an LIF model to different types of input currents
*Estimated timing to here from start of tutorial: 20 min*
In the following section, we will learn how to inject direct current and white noise to study the response of an LIF neuron.
```
# @title Video 2: Response of the LIF neuron to different inputs
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="av541417171", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="preNGdab7Kk", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
## Section 2.1: Direct current (DC)
*Estimated timing to here from start of tutorial: 30 min*
### Interactive Demo 2.1: Parameter exploration of DC input amplitude
Here's an interactive demo that shows how the LIF neuron behavior changes for DC input (constant current) with different amplitudes. We plot the membrane potential of an LIF neuron. You may notice that the neuron generates a spike. But this is just a cosmetic spike only for illustration purposes. In an LIF neuron, we only need to keep track of times when the neuron hit the threshold so the postsynaptic neurons can be informed of the spike.
How much DC is needed to reach the threshold (rheobase current)? How does the membrane time constant affect the frequency of the neuron?
```
# @title
# @markdown Make sure you execute this cell to enable the widget!
my_layout.width = '450px'
@widgets.interact(
I_dc=widgets.FloatSlider(50., min=0., max=300., step=10.,
layout=my_layout),
tau_m=widgets.FloatSlider(10., min=2., max=20., step=2.,
layout=my_layout)
)
def diff_DC(I_dc=200., tau_m=10.):
pars = default_pars(T=100.)
pars['tau_m'] = tau_m
v, sp = run_LIF(pars, Iinj=I_dc)
plot_volt_trace(pars, v, sp)
plt.show()
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial1_Solution_1058324c.py)
## Section 2.2: Gaussian white noise (GWN) current
*Estimated timing to here from start of tutorial: 38 min*
Given the noisy nature of neuronal activity _in vivo_, neurons usually receive complex, time-varying inputs.
To mimic this, we will now investigate the neuronal response when the LIF neuron receives Gaussian white noise $\xi(t)$ with mean 0 ($\mu = 0$) and some standard deviation $\sigma$.
Note that the GWN has zero mean, that is, it describes only the fluctuations of the input received by a neuron. We can thus modify our definition of GWN to have a nonzero mean value $\mu$ that equals the DC input, since this is the average input into the cell. The cell below defines the modified gaussian white noise currents with nonzero mean $\mu$.
### Interactive Demo 2.2: LIF neuron Explorer for noisy input
The mean of the Gaussian white noise (GWN) is the amplitude of DC. Indeed, when $\sigma = 0$, GWN is just a DC.
So the question arises how does $\sigma$ of the GWN affect the spiking behavior of the neuron. For instance we may want to know
1. how does the minimum input (i.e. $\mu$) needed to make a neuron spike change with increase in $\sigma$
2. how does the spike regularity change with increase in $\sigma$
To get an intuition about these questions you can use the following interactive demo that shows how the LIF neuron behavior changes for noisy input with different amplitudes (the mean $\mu$) and fluctuation sizes ($\sigma$). We use a helper function to generate this noisy input current: `my_GWN(pars, mu, sig, myseed=False)`. Note that fixing the value of the random seed (e.g., `myseed=2020`) will allow you to obtain the same result every time you run this. We then use our `run_LIF` function to simulate the LIF model.
```
# @markdown Execute to enable helper function `my_GWN`
def my_GWN(pars, mu, sig, myseed=False):
"""
Function that generates Gaussian white noise input
Args:
pars : parameter dictionary
mu : noise baseline (mean)
sig : noise amplitute (standard deviation)
myseed : random seed. int or boolean
the same seed will give the same
random number sequence
Returns:
I : Gaussian white noise input
"""
# Retrieve simulation parameters
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# Set random seed
if myseed:
np.random.seed(seed=myseed)
else:
np.random.seed()
# Generate GWN
# we divide here by 1000 to convert units to sec.
I_gwn = mu + sig * np.random.randn(Lt) / np.sqrt(dt / 1000.)
return I_gwn
help(my_GWN)
# @title
# @markdown Make sure you execute this cell to enable the widget!
my_layout.width = '450px'
@widgets.interact(
mu_gwn=widgets.FloatSlider(200., min=100., max=300., step=5.,
layout=my_layout),
sig_gwn=widgets.FloatSlider(2.5, min=0., max=5., step=.5,
layout=my_layout)
)
def diff_GWN_to_LIF(mu_gwn, sig_gwn):
pars = default_pars(T=100.)
I_GWN = my_GWN(pars, mu=mu_gwn, sig=sig_gwn)
v, sp = run_LIF(pars, Iinj=I_GWN)
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.plot(pars['range_t'][::3], I_GWN[::3], 'b')
plt.xlabel('Time (ms)')
plt.ylabel(r'$I_{GWN}$ (pA)')
plt.subplot(122)
plot_volt_trace(pars, v, sp)
plt.tight_layout()
plt.show()
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial1_Solution_2de5d8a9.py)
### Think! 2.2: Analyzing GWN Effects on Spiking
- As we increase the input average ($\mu$) or the input fluctuation ($\sigma$), the spike count changes. How much can we increase the spike count, and what might be the relationship between GWN mean/std or DC value and spike count?
- We have seen above that when we inject DC, the neuron spikes in a regular manner (clock like), and this regularity is reduced when GWN is injected. The question is, how irregular can we make the neurons spiking by changing the parameters of the GWN?
We will see the answers to these questions in the next section but discuss first!
---
# Section 3: Firing rate and spike time irregularity
*Estimated timing to here from start of tutorial: 48 min*
When we plot the output firing rate as a function of GWN mean or DC value, it is called the input-output transfer function of the neuron (so simply F-I curve).
Spike regularity can be quantified as the **coefficient of variation (CV) of the inter-spike-interval (ISI)**:
\begin{align}
\text{CV}_{\text{ISI}} = \frac{std(\text{ISI})}{mean(\text{ISI})}
\end{align}
A Poisson train is an example of high irregularity, in which $\textbf{CV}_{\textbf{ISI}} \textbf{= 1}$. And for a clocklike (regular) process we have $\textbf{CV}_{\textbf{ISI}} \textbf{= 0}$ because of **std(ISI)=0**.
## Interactive Demo 3A: F-I Explorer for different `sig_gwn`
How does the F-I curve of the LIF neuron change as we increase the $\sigma$ of the GWN? We can already expect that the F-I curve will be stochastic and the results will vary from one trial to another. But will there be any other change compared to the F-I curved measured using DC?
Here's an interactive demo that shows how the F-I curve of a LIF neuron changes for different levels of fluctuation $\sigma$.
```
# @title
# @markdown Make sure you execute this cell to enable the widget!
my_layout.width = '450px'
@widgets.interact(
sig_gwn=widgets.FloatSlider(3.0, min=0., max=6., step=0.5,
layout=my_layout)
)
def diff_std_affect_fI(sig_gwn):
pars = default_pars(T=1000.)
I_mean = np.arange(100., 400., 10.)
spk_count = np.zeros(len(I_mean))
spk_count_dc = np.zeros(len(I_mean))
for idx in range(len(I_mean)):
I_GWN = my_GWN(pars, mu=I_mean[idx], sig=sig_gwn, myseed=2020)
v, rec_spikes = run_LIF(pars, Iinj=I_GWN)
v_dc, rec_sp_dc = run_LIF(pars, Iinj=I_mean[idx])
spk_count[idx] = len(rec_spikes)
spk_count_dc[idx] = len(rec_sp_dc)
# Plot the F-I curve i.e. Output firing rate as a function of input mean.
plt.figure()
plt.plot(I_mean, spk_count, 'k',
label=r'$\sigma_{\mathrm{GWN}}=%.2f$' % sig_gwn)
plt.plot(I_mean, spk_count_dc, 'k--', alpha=0.5, lw=4, dashes=(2, 2),
label='DC input')
plt.ylabel('Spike count')
plt.xlabel('Average injected current (pA)')
plt.legend(loc='best')
plt.show()
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial1_Solution_eba2370f.py)
## Coding Exercise 3: Compute $CV_{ISI}$ values
As shown above, the F-I curve becomes smoother while increasing the amplitude of the fluctuation ($\sigma$). In addition, the fluctuation can also change the irregularity of the spikes. Let's investigate the effect of $\mu=250$ with $\sigma=0.5$ vs $\sigma=3$.
Fill in the code below to compute ISI, then plot the histogram of the ISI and compute the $CV_{ISI}$. Note that, you can use `np.diff` to calculate ISI.
```
def isi_cv_LIF(spike_times):
"""
Calculates the inter-spike intervals (isi) and
the coefficient of variation (cv) for a given spike_train
Args:
spike_times : (n, ) vector with the spike times (ndarray)
Returns:
isi : (n-1,) vector with the inter-spike intervals (ms)
cv : coefficient of variation of isi (float)
"""
########################################################################
## TODO for students: compute the membrane potential v, spike train sp #
# Fill out function and remove
raise NotImplementedError('Student Exercise: calculate the isi and the cv!')
########################################################################
if len(spike_times) >= 2:
# Compute isi
isi = ...
# Compute cv
cv = ...
else:
isi = np.nan
cv = np.nan
return isi, cv
# Set parameters
pars = default_pars(T=1000.)
mu_gwn = 250
sig_gwn1 = 0.5
sig_gwn2 = 3.0
# Run LIF model for sigma = 0.5
I_GWN1 = my_GWN(pars, mu=mu_gwn, sig=sig_gwn1, myseed=2020)
_, sp1 = run_LIF(pars, Iinj=I_GWN1)
# Run LIF model for sigma = 3
I_GWN2 = my_GWN(pars, mu=mu_gwn, sig=sig_gwn2, myseed=2020)
_, sp2 = run_LIF(pars, Iinj=I_GWN2)
# Compute ISIs/CV
isi1, cv1 = isi_cv_LIF(sp1)
isi2, cv2 = isi_cv_LIF(sp2)
# Visualize
my_hists(isi1, isi2, cv1, cv2, sig_gwn1, sig_gwn2)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial1_Solution_27d69c89.py)
*Example output:*
<img alt='Solution hint' align='left' width=1552.0 height=544.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D3_BiologicalNeuronModels/static/W2D3_Tutorial1_Solution_27d69c89_0.png>
## Interactive Demo 3B: Spike irregularity explorer for different `sig_gwn`
In the above illustration, we see that the CV of inter-spike-interval (ISI) distribution depends on $\sigma$ of GWN. What about the mean of GWN, should that also affect the CV$_{\rm ISI}$? If yes, how? Does the efficacy of $\sigma$ in increasing the CV$_{\rm ISI}$ depend on $\mu$?
In the following interactive demo, you will examine how different levels of fluctuation $\sigma$ affect the CVs for different average injected currents ($\mu$).
1. Does the standard deviation of the injected current affect the F-I curve in any qualitative manner?
2. Why does increasing the mean of GWN reduce the $CV_{ISI}$?
3. If you plot spike count (or rate) vs. $CV_{ISI}$, should there be a relationship between the two? Try out yourself.
```
#@title
#@markdown Make sure you execute this cell to enable the widget!
my_layout.width = '450px'
@widgets.interact(
sig_gwn=widgets.FloatSlider(0.0, min=0., max=10.,
step=0.5, layout=my_layout)
)
def diff_std_affect_fI(sig_gwn):
pars = default_pars(T=1000.)
I_mean = np.arange(100., 400., 20)
spk_count = np.zeros(len(I_mean))
cv_isi = np.empty(len(I_mean))
for idx in range(len(I_mean)):
I_GWN = my_GWN(pars, mu=I_mean[idx], sig=sig_gwn)
v, rec_spikes = run_LIF(pars, Iinj=I_GWN)
spk_count[idx] = len(rec_spikes)
if len(rec_spikes) > 3:
isi = np.diff(rec_spikes)
cv_isi[idx] = np.std(isi) / np.mean(isi)
# Plot the F-I curve i.e. Output firing rate as a function of input mean.
plt.figure()
plt.plot(I_mean[spk_count > 5], cv_isi[spk_count > 5], 'bo', alpha=0.5)
plt.xlabel('Average injected current (pA)')
plt.ylabel(r'Spike irregularity ($\mathrm{CV}_\mathrm{ISI}$)')
plt.ylim(-0.1, 1.5)
plt.grid(True)
plt.show()
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial1_Solution_c6f1c4a2.py)
---
# Summary
*Estimated timing of tutorial: 1 hour, 10 min*
Congratulations! You've just built a leaky integrate-and-fire (LIF) neuron model from scratch, and studied its dynamics in response to various types of inputs, having:
- simulated the LIF neuron model
- driven the LIF neuron with external inputs, such as direct current and Gaussian white noise
- studied how different inputs affect the LIF neuron's output (firing rate and spike time irregularity),
with a special focus on low rate and irregular firing regime to mimc real cortical neurons. The next tutorial will look at how spiking statistics may be influenced by a neuron's input statistics.
If you have extra time, look at the bonus sections below to explore a different type of noise input and learn about extensions to integrate-and-fire models.
---
# Bonus
---
## Bonus Section 1: Orenstein-Uhlenbeck Process
When a neuron receives spiking input, the synaptic current is Shot Noise -- which is a kind of colored noise and the spectrum of the noise determined by the synaptic kernel time constant. That is, a neuron is driven by **colored noise** and not GWN.
We can model colored noise using the Ohrenstein-Uhlenbeck process - filtered white noise.
We next study if the input current is temporally correlated and is modeled as an Ornstein-Uhlenbeck process $\eta(t)$, i.e., low-pass filtered GWN with a time constant $\tau_{\eta}$:
$$\tau_\eta \frac{d}{dt}\eta(t) = \mu-\eta(t) + \sigma_\eta\sqrt{2\tau_\eta}\xi(t).$$
**Hint:** An OU process as defined above has
$$E[\eta(t)]=\mu$$
and autocovariance
$$[\eta(t)\eta(t+\tau)]=\sigma_\eta^2e^{-|t-\tau|/\tau_\eta},$$
which can be used to check your code.
```
# @markdown Execute this cell to get helper function `my_OU`
def my_OU(pars, mu, sig, myseed=False):
"""
Function that produces Ornstein-Uhlenbeck input
Args:
pars : parameter dictionary
sig : noise amplitute
myseed : random seed. int or boolean
Returns:
I_ou : Ornstein-Uhlenbeck input current
"""
# Retrieve simulation parameters
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
tau_ou = pars['tau_ou'] # [ms]
# set random seed
if myseed:
np.random.seed(seed=myseed)
else:
np.random.seed()
# Initialize
noise = np.random.randn(Lt)
I_ou = np.zeros(Lt)
I_ou[0] = noise[0] * sig
# generate OU
for it in range(Lt-1):
I_ou[it+1] = I_ou[it] + (dt / tau_ou) * (mu - I_ou[it]) + np.sqrt(2 * dt / tau_ou) * sig * noise[it + 1]
return I_ou
help(my_OU)
```
### Bonus Interactive Demo 1: LIF Explorer with OU input
In the following, we will check how a neuron responds to a noisy current that follows the statistics of an OU process.
- How does the OU type input change neuron responsiveness?
- What do you think will happen to the spike pattern and rate if you increased or decreased the time constant of the OU process?
```
# @title
# @markdown Remember to enable the widget by running the cell!
my_layout.width = '450px'
@widgets.interact(
tau_ou=widgets.FloatSlider(10.0, min=5., max=20.,
step=2.5, layout=my_layout),
sig_ou=widgets.FloatSlider(10.0, min=5., max=40.,
step=2.5, layout=my_layout),
mu_ou=widgets.FloatSlider(190.0, min=180., max=220.,
step=2.5, layout=my_layout)
)
def LIF_with_OU(tau_ou=10., sig_ou=40., mu_ou=200.):
pars = default_pars(T=1000.)
pars['tau_ou'] = tau_ou # [ms]
I_ou = my_OU(pars, mu_ou, sig_ou)
v, sp = run_LIF(pars, Iinj=I_ou)
plt.figure(figsize=(12, 4))
plt.subplot(121)
plt.plot(pars['range_t'], I_ou, 'b', lw=1.0)
plt.xlabel('Time (ms)')
plt.ylabel(r'$I_{\mathrm{OU}}$ (pA)')
plt.subplot(122)
plot_volt_trace(pars, v, sp)
plt.tight_layout()
plt.show()
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial1_Solution_cf5b6a80.py)
---
## Bonus Section 2: Generalized Integrate-and-Fire models
LIF model is not the only abstraction of real neurons. If you want to learn about more realistic types of neuronal models, watch the Bonus Video!
```
# @title Video 3 (Bonus): Extensions to Integrate-and-Fire models
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="G0b6wLhuQxE", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
| github_jupyter |
```
from sympy import symbols, pprint
from sympy import diff
from sympy.solvers import solve
import numpy as np
from scipy import optimize
import string
import random
from autodp.transformer_zoo import Composition
from functools import lru_cache
# data subject
class Entity():
def __init__(self, name="", id=None):
self.name = name
self.id = id
scalar_name2obj = {}
def individual_RDP_gaussian(params, alpha):
"""
:param params:
'sigma' --- is the normalized noise level: std divided by global L2 sensitivity
:param alpha: The order of the Renyi Divergence
:return: Evaluation of the RDP's epsilon
"""
sigma = params['sigma']
value = params['value']
L = params['L']
assert(sigma > 0)
assert(alpha >= 0)
return (alpha * (L ** 2) * (value**2)) / (2 * (sigma**2))
import math
from autodp.autodp_core import Mechanism
from autodp import rdp_bank, dp_bank, fdp_bank, utils
from autodp import transformer_zoo
from scipy.optimize import minimize_scalar
# Example of a specific mechanism that inherits the Mechanism class
class iDPGaussianMechanism(Mechanism):
def __init__(self, sigma, value, L, entity, name='Gaussian',
RDP_off=False, approxDP_off=False, fdp_off=True,
use_basic_RDP_to_approxDP_conversion=False,
use_fDP_based_RDP_to_approxDP_conversion=False):
# the sigma parameter is the std of the noise divide by the l2 sensitivity
Mechanism.__init__(self)
self.name = name # When composing
self.params = {'sigma': sigma, 'value':value, 'L':L} # This will be useful for the Calibrator
self.entity = entity
# TODO: should a generic unspecified mechanism have a name and a param dictionary?
self.delta0 = 0
if not RDP_off:
new_rdp = lambda x: individual_RDP_gaussian({'sigma': sigma,
'value': value,
'L':L}, x)
if use_fDP_based_RDP_to_approxDP_conversion:
# This setting is slightly more complex, which involves converting RDP to fDP,
# then to eps-delta-DP via the duality
self.propagate_updates(new_rdp, 'RDP', fDP_based_conversion=True)
elif use_basic_RDP_to_approxDP_conversion:
self.propagate_updates(new_rdp, 'RDP', BBGHS_conversion=False)
else:
# This is the default setting with fast computation of RDP to approx-DP
self.propagate_updates(new_rdp, 'RDP')
if not approxDP_off: # Direct implementation of approxDP
new_approxdp = lambda x: dp_bank.get_eps_ana_gaussian(sigma, x)
self.propagate_updates(new_approxdp,'approxDP_func')
if not fdp_off: # Direct implementation of fDP
fun1 = lambda x: fdp_bank.log_one_minus_fdp_gaussian({'sigma': sigma}, x)
fun2 = lambda x: fdp_bank.log_neg_fdp_grad_gaussian({'sigma': sigma}, x)
self.propagate_updates([fun1,fun2],'fDP_and_grad_log')
# overwrite the fdp computation with the direct computation
self.fdp = lambda x: fdp_bank.fDP_gaussian({'sigma': sigma}, x)
# the fDP of gaussian mechanism is equivalent to analytical calibration of approxdp,
# so it should have been automatically handled numerically above
# Discussion: Sometimes delta as a function of eps has a closed-form solution
# while eps as a function of delta does not
# Shall we represent delta as a function of eps instead?
class AdversarialAccountant():
def __init__(self, max_budget=10, delta=1e-6):
self.entity2ledger = {}
self.max_budget = max_budget
self.delta = delta
def append(self, entity2mechanisms):
for key, ms in entity2mechanisms.items():
if key not in self.entity2ledger.keys():
self.entity2ledger[key] = list()
for m in ms:
self.entity2ledger[key].append(m)
def get_eps_for_entity(self, entity_name):
# compose them with the transformation: compose.
compose = Composition()
mechanisms = self.entity2ledger[entity_name]
composed_mech = compose(mechanisms, [1]*len(mechanisms))
# Query for eps given delta
return Scalar(value=composed_mech.get_approxDP(self.delta),
min_val=0,
max_val=self.max_budget,
ent=Entity(name=entity_name))
def has_budget(self, entity_name):
return self.get_eps_for_entity(entity_name)._value < self.max_budget
@property
def entities(self):
return self.entity2ledger.keys()
@property
def overbudgeted_entities(self):
entities = set()
for ent in self.entities:
if not self.has_budget(ent):
entities.add(ent)
return entities
def print_ledger(self, delta=1e-6):
for entity, mechanisms in self.entity2ledger.items():
print(entity + "\t" + str(self.get_eps_for_entity(entity)._value))
def run(func, **kwargs):
return func.subs(kwargs)
def run_specific(f, **kwargs):
"""pass in kwargs to run in fixed polynomial because this is what
optimize.brute expects"""
return run(f, **kwargs)
@lru_cache(maxsize=None)
def search(run_specific_args, rranges, full_output, finish, disp):
return optimize.shgo(run_specific_args, rranges)
class Scalar():
def __init__(self, value, min_val=None, max_val=None, poly=None, ent=None, name=None):
if name is None:
lower_upper_alphabet = string.ascii_letters
name = ''.join([random.choice(lower_upper_alphabet) for i in range(5)])
self.name = name
self._value = value
self._min_val = min_val
self._max_val = max_val
self.enabled = True
if poly is not None:
# if this Scalar is being formed as a function of other Scalar objects
self._poly = poly
elif ent is not None:
# if you're creating a Scalar for the first time (no parents)
self.scalar_name = self.name + "_" + ent.name
self._poly = symbols(self.scalar_name)
scalar_name2obj[self.scalar_name] = self
else:
raise Exception("Poly or ent must be not None")
@property
def poly(self):
return self._poly
@property
def value(self):
if(self._value is not None):
return self._value
sy_names = self.poly.free_symbols
sym = list()
for sy_name in sy_names:
sym.append(scalar_name2obj[str(sy_name)])
run_specific_args, index2symbol, symbol2index = self.create_run_specific_args(f=self.poly)
inputs = list()
for sym in index2symbol:
inputs.append(scalar_name2obj[sym]._value)
return run_specific_args(inputs)
@property
def min_val(self):
return self._min_val
@property
def max_val(self):
return self._max_val
def __mul__(self, other):
result_poly = self.poly * other.poly
result = Scalar(value=None, poly=result_poly)
return result
def __add__(self, other):
result_poly = self.poly + other.poly
result = Scalar(value=None, poly=result_poly)
return result
def __sub__(self, other):
result_poly = self.poly - other.poly
result = Scalar(value=None, poly=result_poly)
return result
def __str__(self):
return str(self.poly) + "=" + str(self.value)
def __repr__(self):
return str(self)
@property
def sens(self):
if self.min_val is not None and self.max_val is not None:
return self.max_val - self.min_val
def neg_deriv(self, name):
obj = scalar_name2obj[name]
return -diff(self.poly, obj.poly)
def create_run_specific_args(self, f):
free_symbols_list = list(self.poly.free_symbols)
index2symbol = list(map(lambda x:str(x), free_symbols_list))
symbol2index = {}
for i,sym in enumerate(index2symbol):
symbol2index[sym] = i
def _run_specific_args(tuple_of_args, *params):
kwargs = {}
for sym,i in symbol2index.items():
kwargs[sym] = tuple_of_args[i]
return run_specific(f=f, **kwargs)
return _run_specific_args, index2symbol, symbol2index
def get_mechanism(self,
symbol_name = 'b',
sigma = 0.1):
# Step 1: get derivative we want to maximize
z = self.neg_deriv(symbol_name)
# Step 2: prepare metadata for optimize.brute() function
sy_names = z.free_symbols
sym = list()
for sy_name in sy_names:
sym.append(scalar_name2obj[str(sy_name)])
run_specific_args, index2symbol, symbol2index = self.create_run_specific_args(f=z)
rranges = list()
for i,sym in enumerate(index2symbol):
obj = scalar_name2obj[sym]
rranges.append(tuple([obj.min_val, obj.max_val]))
# Step 3: maximize the derivative over a bounded range of <entity_name>
resbrute = search(run_specific_args, tuple(rranges), full_output=False, finish=None, disp=True)
resbrute = resbrute.x
if isinstance(resbrute, np.float64):
L = resbrute
else:
L = resbrute[symbol2index[symbol_name]]
input_obj = scalar_name2obj[symbol_name]
# Step 4: create the gaussian mechanism object
gm1 = iDPGaussianMechanism(sigma=sigma, value=input_obj._value, L=L, entity=symbol_name.split("_")[1], name='gm_'+symbol_name)
return gm1
def get_all_entity_mechanisms(self,sigma=0.1):
sy_names = self.poly.free_symbols
entity2mechanisms = {}
for sy_name in sy_names:
mechanism = self.get_mechanism(symbol_name=str(sy_name), sigma=sigma)
split_name = str(sy_name).split("_")
entity_name = split_name[1]
var_name = split_name[0]
if entity_name not in entity2mechanisms.keys():
entity2mechanisms[entity_name] = list()
entity2mechanisms[entity_name].append(mechanism)
return entity2mechanisms
@property
def entities(self):
entities = set()
sy_names = self.poly.free_symbols
for sy_name in sy_names:
entities.add(str(sy_name).split("_")[1])
return entities
def publish(self, acc, sigma=1.5):
acc_original = acc
assert sigma > 0
acc_temp = deepcopy(acc_original)
# get mechanisms for new publish event
ms = self.get_all_entity_mechanisms(sigma=sigma)
acc_temp.append(ms)
overbudgeted_entities = acc_temp.overbudgeted_entities
sample = random.gauss(0,sigma)
while len(overbudgeted_entities) > 0:
for sy_name in self.poly.free_symbols:
entity_name = str(sy_name).split("_")[1]
if(entity_name in overbudgeted_entities):
sym = scalar_name2obj[str(sy_name)]
self._poly = self.poly.subs(sym.poly, 0)
acc_temp = deepcopy(acc_original)
# get mechanisms for new publish event
ms = self.get_all_entity_mechanisms(sigma=sigma)
acc_temp.append(ms)
overbudgeted_entities = acc_temp.overbudgeted_entities
output = self.value + sample
acc_original.entity2ledger = deepcopy(acc_temp.entity2ledger)
return output
from copy import deepcopy
bob = Scalar(value=1, min_val=-2, max_val=2, ent=Entity(name="Bob"))
bobby = Scalar(value=1, min_val=-2, max_val=2, ent=Entity(name="Bob"))
alice = Scalar(value=1, min_val=-1, max_val=1, ent=Entity(name="Alice"))
charlie = Scalar(value=2, min_val=-2, max_val=2, ent=Entity(name="Charlie"))
david = Scalar(value=2, min_val=-2, max_val=2, ent=Entity(name="David"))
acc = AdversarialAccountant(max_budget=70)
# PhiScalar()
bob2 = bob + bob
bobby2 = bobby + bobby
alice2 = alice + alice
charlie2 = charlie + charlie
(alice2 * bobby2) + (alice2 * bob2)
def __add__(self, other):
if self.gama==False and other.gamma== False and self.entity == other.entity
return Scalar(entity=self.entity,
value=self.value + other.value,
min_val=self.min_val + other.min_value,
max_val=self.max_val + other.max_val,
symbol_name=entity_name + "_" + random_hash)
else:
self.gama = True
self.poly = symbol(self.symbol_name)
other.gama = True
other.poly = symbol(other.symbol_name)
# GammaScalar()
out = bob2**2 + alice2*0.5 + bobby2**3
#
%%timeit -n1 -r1
public_out = out.publish(acc=acc, sigma=0.5)
acc.print_ledger()
result = optimize.shgo(eggholder, bounds)
result.x, result.fun
```
| github_jupyter |
# Self-orginizing maps
Self-orginizing map (SOM) is a type of neural network which is trained using unsupervised learning algorithms. One of the basic abilities of SOM is to project high-dimensional data to lower dimension (1D, 2D, 3D obviously). SOM can be considered as a general cluster analysis tool.
Scheme of 2D SOM:
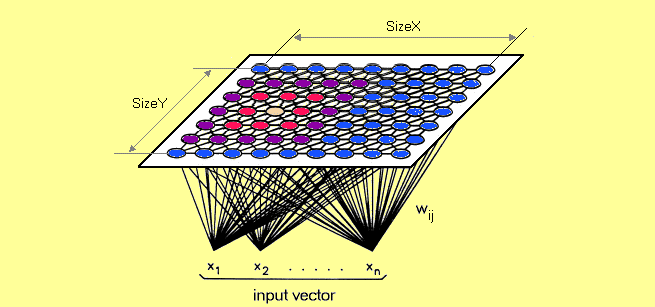
(Source of the scheme - https://miro.medium.com/max/655/1*QG7afWQKjY3IpezhNQMzBg.png)
2D SOM consists of neurons in a plane, each neuron has *n* number of weights, where *n* equals to input data dimension.
Positions of neurons are determined by any kind of distance metrics. For our pupose the Euclidean distance is sufficient.
Euclidean distance between two neurons with weights vector $\boldsymbol{w_1}$ and $\boldsymbol{w_2}$:
$$||\boldsymbol{w_1}-\boldsymbol{w_2}||= \sqrt{\sum_{j=1}^{p}(w_{1j}-w_{2j})^2}$$
## Learning algorithm
(Source of the gif - https://upload.wikimedia.org/wikipedia/commons/3/35/TrainSOM.gif)
1. The weights $\boldsymbol{w(0)}$ have random values in initialization step.
2. Random observation $\boldsymbol{x}(k)$ is chosen from input data
3. Calculate the distance (e.g. Euclidean) between chosen observation and all neuron weights
4. Find which neuron is the closest to chosen observation. This is our winning neuron $v = arg min ||\boldsymbol{x}(k) - \boldsymbol{w}||$
5. Update winning neuron weights $\boldsymbol{w_v}$: $$\boldsymbol{w_v}(k+1) = \boldsymbol{w_v}(k) + \alpha(k) [\boldsymbol{x}(k) - \boldsymbol{w_v}(k)]$$ where $\alpha(k)$ is a learning parameter which should be high at the beginning and low at the end of learning algorithm. We want to make bigger steps at the beginning of learning and then finer and finer: $$ \alpha(k) = \alpha(0) \exp\left({- \frac{k}{\lambda_\alpha}}\right),$$
6. However it is important to update also the neighbour neurons $\boldsymbol{w_n}(k)$ of the winning neuron. This is crucial for finding similar clusters of neurons (this is the basic formula of learning algorithm): $$\boldsymbol{w_n}(k+1) = \boldsymbol{w_n}(k) + \alpha(k)\eta(v,k,n) [\boldsymbol{x}(k) - \boldsymbol{w_v}(k)]$$ The update is similar to winning neuron update but it has additional neighbourhood parameter $\eta(v,k,n)$, which determines how much are the neighbours influenced by the winning neuron. The closest neighbours are influenced the most and the influence is gettig weaker with bigger distance according to: $$\eta = exp\left( -\frac{||n-v||^2}{2 \sigma(k)^2} \right)$$ which is a gaussian function where $||n-v||$ is distance between winning neuron and the neighbour and $\sigma(k)$ is a parameter which determines how wide is the neighbourhood area. $\sigma(k)$ should be also high at the beginning and low at the end of the algorithm: $$ \sigma(k) = \sigma(0) \exp\left({- \frac{k}{\lambda_\sigma}}\right),$$
7. Go to step 2. and repeat
## Simple 2D SOM example
```
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
Define a function for 2D SOM creation. SOM has some number of neurons determined by number of rows and columns in SOM plane and is also determined by number of weights of each neuron
```
def new_SOM(nRows, nCols, nWeights):
som = np.random.rand(nRows, nCols, nWeights) #random initialization
return som
```
Define a function which finds the winning neuron and returns its position and distance between the winner and the observation from input data
```
def find_winner(som, x):
#initialization
dist_win = np.inf
row_win = np.nan
col_win = np.nan
nRowsSOM = som.shape[0]
nColsSOM = som.shape[1]
#go through the som
for row in range(nRowsSOM):
for col in range(nColsSOM):
#calculate euclidean distance between neuron and input data
currentCentroidDistance = np.sqrt(sum((som[row, col, :]-x)**2))
#find the closest one
if currentCentroidDistance < dist_win:
dist_win = currentCentroidDistance
row_win = row
col_win = col
return row_win, col_win, dist_win
```
Define a function for Euclidean distance between neurons
```
def distance_in_map(row1, col1, row2, col2):
return np.sqrt((row1-row2)**2+(col1-col2)**2)
```
Finally the learning algorithm. Inputs are our SOM, input datax, iteration index and parameters of learning:
* sigma0 is initial size of neighbourhood
* alpha0 is initial learning parameter
* lambdaSigma determines how quickly will neighbourhood influence decrease during the algorithm
* lambaAlpha determines how quickly will learning parameter decrease during the algorithm
```
def training(som, x, iteration, sigma0, alpha0, lambdaSigma, lambdaAlpha):
row_win, col_win, dist_win = find_winner(som, x)
nRowsSOM = som.shape[0]
nColsSOM = som.shape[1]
#alpha parameter is getting lower during the algorithm
alphaIteration = alpha0 * np.exp(-iteration/float(lambdaAlpha))
#Sigma parameter is getting lower during the algorithm
sigmaIteration = sigma0 * np.exp(-iteration/float(lambdaSigma))
#Go through rows and columns
for row in range(nRowsSOM):
for col in range(nColsSOM):
#this is the main formula as in step 6.
deltaWeights = (x - som[row, col, :]) * \
alphaIteration * np.exp(-distance_in_map(row, col, row_win, col_win)**2 / float(sigmaIteration)**2)
som[row, col, :] = som[row, col, :] + deltaWeights
return som
```
## Let's apply the SOM!
We would like to distinguish color clusters. Input data will have 3 dimensions (RGB values)
```
colors = [[1,0,0], [0,1,0], [0,0,1]] #three basic colors
data = []
#create a noisy RGB input data
for i in range (150):
#choose one of the colors and add noise
data.append(colors[np.random.randint(0,3)]+np.random.rand(3)/5.)
#color value should be in range <0;1>
data[-1] = np.clip(data[-1],0.0,1.0)
```
Now create a SOM and plot it as an image. You will see random pixels
```
som = new_SOM(10,10,3)
plt.imshow(som)
plt.show()
```
Try 10 learning iterations. SOM weights will get closer to input data and will cluster similar pixels
```
#You can try to play with the parameters and test the learning behaviour
sigma0 = 10
alpha0 = 0.5
lambdaSigma = 30
lambdaAlpha = 20
for i in range(10):
som = training(som, data[i], i+1, sigma0, alpha0, lambdaSigma, lambdaAlpha)
plt.imshow(som)
plt.show()
```
More and more interations
```
for i in range(40):
som = training(som, data[i], i+1, sigma0, alpha0, lambdaSigma, lambdaAlpha)
plt.imshow(som)
plt.show()
for i in range(100):
som = training(som, data[i], i+1, sigma0, alpha0, lambdaSigma, lambdaAlpha)
plt.imshow(som)
plt.show()
```
We made a visualization of neuron weights using RGB.
We will plot the winning neurons now.
```
plt.figure()
#for each color (RGB) find the winner
for colour in [[1,0,0],[0,1,0],[0,0,1]]:
X = []
Y = []
for i in range(10):
#find the winner from noisy data
row, col, d = find_winner(som, np.array(colour)+np.random.rand(3)/5.)
X.append(col)
Y.append(row)
plt.plot(Y,X,"x", label=str(colour))
plt.legend()
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/daveluo/covid19-healthsystemcapacity/blob/master/nbs/usa_beds_capacity_analysis_20200313_v2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!apt-get install python3-rtree
!pip install geopandas
import geopandas as gpd
import pandas as pd
import numpy as np
!wget https://raw.githubusercontent.com/daveluo/covid19-healthsystemcapacity/master/data/usa_hospital_beds_hcris2018_cleaned3.geojson
hosp_gdf = gpd.read_file('usa_hospital_beds_hcris2018_cleaned3.geojson')
hosp_gdf.head()
hosp_gdf.shape
hosp_gdf['Total Beds'].sort_values(ascending=False)
hosp_gdf[['Total Beds', 'ICU Total Beds']].sum()
hosp_county_gdf = hosp_gdf.groupby(['State','County'], as_index='False')['ICU Total Beds', 'Total Beds'].sum().reset_index()
hosp_county_gdf.head()
# thank you https://eric.clst.org/tech/usgeojson/
!wget https://eric.clst.org/assets/wiki/uploads/Stuff/gz_2010_us_050_00_20m.json
import json, os
us_county_path = 'gz_2010_us_050_00_20m.json'
cur_json = json.load(open(us_county_path, encoding='ISO-8859-1'))
path,ext = os.path.splitext(us_county_path)
new_path =path+"_new"+ext
with open(new_path,"w", encoding='utf-8') as jsonfile:
json.dump(cur_json,jsonfile,ensure_ascii=False)
us_county = gpd.read_file(new_path, driver='GeoJSON')
us_county.plot()
hosp_gdf.plot()
hosp_gdf_countyjoin = gpd.sjoin(hosp_gdf, us_county, how='inner', op='intersects')
hosp_gdf_countyjoin.head()
hosp_county_gdf = hosp_gdf_countyjoin.groupby(['STATE','COUNTY','GEO_ID'], as_index='False')['ICU Total Beds', 'Total Beds','Total Bed Days Available','Total Inpt Days', 'ICU Total Bed Days Available', 'ICU Total Inpt Days'].sum().reset_index()
hosp_county_gdf.shape
hosp_county_gdf = pd.merge(hosp_county_gdf, us_county)
hosp_county_gdf['ICU Occupancy Rate'] = hosp_county_gdf['ICU Total Inpt Days']/hosp_county_gdf['ICU Total Bed Days Available']
hosp_county_gdf['Total Bed Occupancy Rate'] = hosp_county_gdf['Total Inpt Days']/hosp_county_gdf['Total Bed Days Available']
hosp_county_gdf.head()
hosp_county_gdf = gpd.GeoDataFrame(hosp_county_gdf, crs=4326)
hosp_county_gdf.plot(figsize=(15,15))
# get latest census data for population demographics by us county at https://www.census.gov/data/tables/time-series/demo/popest/2010s-counties-detail.html
# data description here: https://www2.census.gov/programs-surveys/popest/technical-documentation/file-layouts/2010-2018/cc-est2018-alldata.pdf
!wget https://www2.census.gov/programs-surveys/popest/datasets/2010-2018/counties/asrh/cc-est2018-alldata.csv
census_df = pd.read_csv('cc-est2018-alldata.csv', encoding='unicode_escape')
census_df.head()
census_df['fips_code'] = census_df['STATE'].apply(lambda x: str(x).zfill(2)) + census_df['COUNTY'].apply(lambda x: str(x).zfill(3))
census_df.head()
# YEAR 11 = 7/1/2018 population estimate
census_df[census_df['YEAR'] == 11].head()
census2018_df = census_df[census_df['YEAR'] == 11]
# pop of 15+ year olds by county
# AGEGRP 4 = Age 15 to 19 years
# AGEGRP 0 = All, need to exclude or double counts
adult_totpop_countywise = census2018_df[census2018_df['AGEGRP']>=4].groupby(['fips_code'])['TOT_POP'].sum()
all_totpop_countywise = census2018_df[census2018_df['AGEGRP']>=1].groupby(['fips_code'])['TOT_POP'].sum()
adult_totpop_countywise.sum(), all_totpop_countywise.sum()
adult_totpop_countywise
hosp_county_gdf.shape
hosp_county_gdf['fips_code'] = hosp_county_gdf['GEO_ID'].apply(lambda x: x[-5:])
hosp_county_census_gdf = hosp_county_gdf.join(adult_totpop_countywise, how='inner', on='fips_code')
hosp_county_census_gdf.rename({'TOT_POP':'15 and Older Population'}, axis=1, inplace=True)
hosp_county_census_gdf = hosp_county_census_gdf.join(all_totpop_countywise, how='inner', on='fips_code')
hosp_county_census_gdf.rename({'TOT_POP':'All Population'}, axis=1, inplace=True)
hosp_county_census_gdf.head()
hosp_county_census_gdf['ICU Beds per 1000 Adults (15+)'] = hosp_county_census_gdf['ICU Total Beds']/(hosp_county_census_gdf['15 and Older Population']/1000)
hosp_county_census_gdf['Total Beds per 1000 Adults (15+)'] = hosp_county_census_gdf['Total Beds']/(hosp_county_census_gdf['15 and Older Population']/1000)
hosp_county_census_gdf['ICU Beds per 1000 People'] = hosp_county_census_gdf['ICU Total Beds']/(hosp_county_census_gdf['All Population']/1000)
hosp_county_census_gdf['Total Beds per 1000 People'] = hosp_county_census_gdf['Total Beds']/(hosp_county_census_gdf['All Population']/1000)
hosp_county_census_gdf.head()
# sanity check
hosp_county_census_gdf['ICU Total Beds'].sum() / (hosp_county_census_gdf['All Population'].sum()/100000)
hosp_county_census_gdf['Total Beds'].sum() / (hosp_county_census_gdf['All Population'].sum()/1000)
```
## Check bed stats against:
https://www.kff.org/other/state-indicator/beds-by-ownership/?currentTimeframe=0&sortModel=%7B%22colId%22:%22Location%22,%22sort%22:%22asc%22%7D
- 2.4 staffed beds per 1,000 pop across USA
https://www.sccm.org/Communications/Critical-Care-Statistics
AHA data: According to the AHA 2015 annual survey, the United States had
- 4862 acute care registered hospitals;
- 2814 of these had at least 10 acute care beds and at least 1 ICU bed.
- These hospitals had a total of 540,668 staffed beds and 94,837 ICU beds (14.3% ICU beds/total beds) in 5229 ICUs.
- There were 46,490 medical-surgical beds in 2644 units,
- 14,731 cardiac beds in 976 units,
- 6588 other beds in 379 units,
- 4698 pediatric beds in 307 units, and
- 22,330 neonatal beds in 920 units.
- The median number of beds in medical-surgical, cardiac, and other units was 12, with 10 beds in pediatrics and 18 in neonatal. Fifty-two percent of hospitals had 1 unit, 24% had 2 units, and 24% had 3 or more units.
HCRIS data:
- In 2010 there were 2977 acute care hospitals with ICU beds.
- In these, there were 641,395 total acute care beds with 103,900 ICU beds (16.2% ICU beds/total beds).
- From 2000 to 2010, the number of critical care beds in the United States increased by 17.8%, from 88,235 to 103,900. However, the majority of the growth in critical care bed supply is occurring in a small number of U.S. regions that tend to have large populations, fewer baseline ICUs per 100,000 capita, higher baseline ICU occupancy, and increased market competition. Additionally, between 2000 and 2010, the greatest percentage increases were in neonatal beds (29%), followed by adult beds (26%); there were minimal changes in pediatric beds (2.7%).
- Of the 103,900 ICU beds in 2010,
- 83,417 (80.3%) were adult,
- 1917 (1.8%) were pediatric, and
- 18,567 (17.9%) were neonatal.
- In total, there were 33.6 beds per 100,000 population,
- 35.5 beds per 100,000 adult beds (age > 18 years),
- 2.7 beds/100,000 pediatric beds (age 1-17 years),
- and 470 beds/100,000 neonatal beds (age < 1 year).
ICU days: HCRIS analysis showed that there were 150.9 million hospital days, including 25 million ICU days in 2010 (16.5% ICU days/total days). Medicare accounted for 7.9 million ICU days (31.4%) and Medicaid 4.3 million ICU days (17.2%).
Occupancy: Occupancy rates were calculated from HCRIS (days/possible days) data. In 2010, hospital and ICU occupancy rates were 64.6% and 68%, respectively. Occupancy rates vary by hospital size, with higher occupancy rates associated with larger hospitals.
```
hosp_county_census_gdf.to_file('usa_county_hospital_bedcapacity2018.geojson', driver='GeoJSON')
hosp_county_census_gdf.to_csv('usa_county_hospital_bedcapacity2018.csv')
```
| github_jupyter |
```
#Load libraries
import warnings
warnings.filterwarnings("ignore")
import numpy as np
import pandas as pd
from pandas import read_csv
from pandas import set_option
from matplotlib import pyplot
from pandas import read_csv
from pandas import set_option
from matplotlib import pyplot as plt
import seaborn
HOME_PATH = '' #home path of the project
FILENAME = 'F_IndianLiverPatient_Data_Real.csv'
```
## 1. Load the dataset
```
dataset = pd.read_csv(HOME_PATH + FILENAME)
dataset
```
## 2. Analyze data
```
categorical_cols = ['gender','class']
categorical_cols
#dimensions of the dataset
dataset.shape
#data types of each attribute
dataset.dtypes
#peak of the data
dataset.head(20)
#summarize the distribution of each attribute
set_option('precision', 2)
dataset.describe()
```
## 3. Data visualization
```
for col in dataset.columns :
# Multiple box plots on one Axes
data = dataset[col]
if col in categorical_cols :
data = data.astype("category").cat.codes
fig, ax = plt.subplots()
ax.boxplot(data)
ax.set_title(col)
for col in dataset.columns :
# Multiple box plots on one Axes
data = dataset[col]
if col in categorical_cols :
data = data.astype("category").cat.codes
fig, ax = plt.subplots()
ax.hist(data, density=False, histtype='bar')
ax.set_title(col)
#Correlation matrix
set_option('precision', 2)
pyplot.figure(figsize=(20,10))
cors = abs(dataset.corr(method='pearson'))
seaborn.heatmap(cors, mask=np.triu(np.ones_like(cors, dtype=bool)), vmin=0, vmax=1, cmap='Blues', annot=True)
pyplot.show()
```
## 4. Edit data
```
for col in dataset.columns :
if not dataset[col].isnull().values.any() :
print(col, ':', 'NO NaN values')
else :
print(col, ':', 'NaN values finded')
print('Number of NaN values: ', dataset[col].isnull().sum())
#quick look at the breakdown of class values
for col in categorical_cols :
dataset[col] = dataset[col].astype('category')
print('###########################')
print(dataset.groupby(col).size())
```
## 5. Data split (train and test)
```
from sklearn.model_selection import train_test_split
#Split data indixes in train and test
idx_train, idx_test = train_test_split(dataset.index.tolist(), train_size=0.8, random_state=42, shuffle=True)
print('Train data length: ', len(idx_train))
print('Test data length: ', len(idx_test))
print('Total data length: ', len(idx_train) + len(idx_test))
#Select train data and save locally
diabetes_train_data = dataset.loc[idx_train]
diabetes_train_data.to_csv(HOME_PATH + 'TRAIN DATASETS/F_IndianLiverPatient_Real_Train.csv', index=False)
#Select test data and save locally
diabetes_test_data = dataset.loc[idx_test]
diabetes_test_data.to_csv(HOME_PATH + 'TEST DATASETS/F_IndianLiverPatient_Real_Test.csv', index=False)
print('Train data size: ', diabetes_train_data.shape)
print('Test data length: ', diabetes_test_data.shape)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from urllib.request import Request, urlopen
from IPython.display import Markdown as md
%matplotlib inline
```
# Data explorations
(c) Carlos Contreras, August 2021
## Load data
```
df_comor = pd.read_csv('../../data/AHS/Restricted/demo_comorb_cdom.csv', true_values=["Yes"])
df_hosps = pd.read_csv('../../data/AHS/Restricted/hosps.csv')
df_comor['Year_Month'] = pd.to_datetime(df_comor['Year_Month'], format='%Y_%m')
md("""The data is contained in two datasets:
- `demo_comorb_cdom`: contains personal information, COVID-19 infection information, comorbidities, and symptoms.
- Number of entries: {}
- Number of features: {}
- Observations range: {} to {}
- `hosps`: hospitalizations (one patient can be admited more than once) with admission and discharge days, and ICU, ventilation and death flags.
- Number of observations: {}
- Number of features: {}
- Several patients ({}) have been admitted more than once to the hospital. The count of unique values in `demo_comorb_cdom` and `hosps` coincide.
""".format(df_comor.shape[0],
df_comor.shape[1],
min(df_comor['Year_Month']).strftime("%B %Y"),
max(df_comor['Year_Month']).strftime("%B %Y"),
df_hosps.shape[0],
df_hosps.shape[1],
sum(df_hosps.pivot_table(columns=['PHN_ENC'], aggfunc='size').value_counts()[1:])))
pd.DataFrame(df_hosps.pivot_table(columns=['PHN_ENC'], aggfunc='size').value_counts(),
columns=['Number of patients admited once or more'])
```
## Featured variables
- Number of comorbidities.
- Number of symptoms, including symptoms listed in other.
- Dead flag: 1 (True) if death days is non-empty (greather than zero)
- Number of hospitalizations
```
df_comor['Num. comorbidities'] = df_comor.iloc[:, 4:20].sum(axis=1)
df_comor['Num. symptoms'] = df_comor.iloc[:, 22:53].sum(axis=1) + \
df_comor.iloc[:, 54].str.split(',').apply(lambda x: len(x) if type(x)==list else 0)
df_comor['Num. symptoms'] = df_comor['Num. symptoms'].astype(int)
df_hosps['Dead'] = (df_hosps.Death_days >= 0)
df_comor['Age group'] = pd.cut(df_comor['age'], bins=[0, 1, 5, 10, 20, 30, 40, 50, 60, 70, 80, np.inf],
right=True, include_lowest=True,
labels=['Under 1 year', '1-4 years', '5-9 years', '10-19 years',
'20-29 years', '30-39 years', '40-49 years', '50-59 years',
'60-69 years', '70-79 years', '80+ years'])
# number of hospitalizations
df_numhosps = pd.DataFrame(df_hosps.pivot_table(columns=['PHN_ENC'], aggfunc='size'), columns=['Num. hospitalizations'])
df_numhosps.reset_index(inplace=True)
```
Merged data frame. Each entry corrresponds to one hospitalization and personal information is thus duplicated in cases with multiple hospitalizations. This means each entry is considered a separate patient. **(WARNING: duplication bias?)**
```
df = pd.merge(df_comor, pd.merge(df_hosps, df_numhosps, on='PHN_ENC'), on='PHN_ENC')
df = df.rename(columns={"age": "Age",
"Year_Month": "Year Month",
"MI": "Myocardial infarction",
"CHF": "Congestive Heart Failure",
"PVD": "Peripheral Vascular Disease",
"CEVD": "Cerebrovascular Disease",
"Dementia": "Dementia",
"CPD": "Chronic Pulmonary Disease",
"Rheumatic": "Rheumatic Disease",
"PUD": "Peptic Ulcer Disease",
"MILDLD": "Liver disease – mild",
"DIAB_UC": "Diabetes without complications",
"DIAB_C": "Diabetes with complications",
"Paraplegia": "Paraplegia and Hemiplegia",
"RD": "Renal Disease",
"Cancer": "Cancer",
"MSLD": "Metastatic Carcinoma",
"METS": "Liver disease – moderate/severe",
"SSHX_DATA_ANOREXIA": "Anorexia",
"SSHX_DATA_ARTHRALGIA": "Arthralgia",
"SSHX_DATA_CHEST_PAIN": "Chest pain",
"SSHX_DATA_FEVERISH_CHILLS": "Feverish chills",
"SSHX_DATA_CONJUNCTIVITAL_INJECTI": "Conjunctivital injecti",
"SSHX_DATA_CONJUNCTIVITIS": "Conjunctivitis",
"SSHX_DATA_COUGH": "Cough",
"SSHX_DATA_DECREASED_APPETITE": "Decreased appetite",
"SSHX_DATA_DIARRHEA": "Diarrhea",
"SSHX_DATA_DIZZINESS": "Dizziness",
"SSHX_DATA_ENCEPHALITIS": "Encephalitis",
"SSHX_DATA_FEVER": "Fever",
"SSHX_DATA_HEADACHE": "Headache",
"SSHX_DATA_HYPOTENSION": "Hypotension",
"SSHX_DATA_IRRITABILITY_CNFSN": "Irritability cnfsn",
"SSHX_DATA_LOSS_OF_TASTE_SMELL": "Loss of taste smell",
"SSHX_DATA_MALAISE": "Malaise",
"SSHX_DATA_MYALGIA": "Myalgia",
"SSHX_DATA_NASAL_CONGESTION": "Nasal congestion",
"SSHX_DATA_NAUSEA": "Nausea",
"SSHX_DATA_NOSE_BLEED": "Nose bleed",
"SSHX_DATA_PAIN": "Pain",
"SSHX_DATA_PHARYNGEAL_EXUDATE": "Pharyngeal exudate",
"SSHX_DATA_PROSTRATION": "Prostration",
"SSHX_DATA_RHINORRHEA": "Rhinorrhea",
"SSHX_DATA_SEIZURES": "Seizures",
"SSHX_DATA_DIFFICULTY_BREATHING": "Difficulty breathing",
"SSHX_DATA_SNEEZING": "Sneezing",
"SSHX_DATA_SORE_THROAT": "Sore throat",
"SSHX_DATA_TACHYPNEA": "Tachypnea",
"SSHX_DATA_VOMITING": "Vomiting",
"SSHX_DATA_ALTERED_MENTAL_STATE": "Altered mental state",
"SSHX_DATA_ABN_LUNG_ASC": "Abn lung asc",
"SSHX_DATA_OTHER": "Other",
"SIGNSYMPHX_IF_OTHER": "List of other symptoms",
"ICU_flag": "ICU",
"Disch days": "Days to discharge",
"LOS": "Length of stay",
"ICU_flag": "ICU",
"Vent_flag": "Ventilation",
"Death_days": "Days to death",
"Admit_days": "Days to admission"})
df = df.replace({"M":"Male", "F": "Female"})
idx = list(range(4,20))+[61, 62]
df.iloc[:, idx] = df.iloc[:, idx].astype('bool')
df.info()
```
Some summary tables bellow.
```
df.describe(include=['bool', 'object', 'category']).transpose()
df.describe(include=['int', 'float'])
```
## Exclusions
- Two (2) entries have age greater than 140. Unrealistic: removing from the table.
```
print("Number of entries (original): ", df.shape[0])
# Incorrect age entries
print("\nSorted values of age:")
print(df.Age.sort_values(ascending=False).head())
df.drop(df[df['Age'] >= 140].index, inplace=True)
print("\nEntries remaining: ", df.shape[0])
# Multiple hospitalizations?
# df.drop(df[df['Dead_flag'] & df['NUM_HOSPS']>2].index, inplace=True)
df.to_csv('../../data/AHS/Restricted/analysis.csv')
```
## Charts and plots
```
def counttable(df):
def Count(x):
return x
def Percent(x):
return (x/x.sum()*100).round(2)
temp = df.apply([Count, Percent], axis=0)
return temp.append(pd.Series(temp.sum(), name='Total'))
```
### Sex and age
```
df_temp = df
temp = df_temp['Sex'].value_counts()
temp = temp.rename(index={"M":"Male", "F": "Female"})
temp.index.name = "Sex"
temp.plot.bar()
plt.xlabel("Sex", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Gender of patients", fontsize=16)
plt.xticks(rotation=0)
counttable(temp)
df_temp = df
temp = df_temp['Age group'].value_counts()
temp = temp.reindex(index=df_comor['Age group'].cat.categories.values)
temp.index.name = "Age group"
temp.plot.bar()
plt.xlabel("Age group", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Age distribution", fontsize=16)
counttable(temp)
temp = df.groupby('Sex')['Age group'].value_counts().unstack(0)
temp = temp.reindex(index=df_comor['Age group'].cat.categories.values)
temp = temp.rename(columns={"M":"Male", "F": "Female"})
temp.plot.bar()
plt.xlabel("Age group", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Age distribution by gender", fontsize=16)
counttable(temp)
```
### Comorbidities
```
temp = df.iloc[:, 4:20].sum()
temp.index.name = "Comorbidity"
temp.sort_values(ascending=False).plot.pie(y='Percent')
plt.ylabel("")
counttable(temp).sort_values('Percent', ascending=False)
df_temp = df
temp = df_temp['Num. comorbidities'].value_counts()
temp.index.name = "Num. comorbidities"
temp.plot.bar()
plt.xlabel("Number of comorbidities", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Distribution of number of comorbidities", fontsize=16)
plt.xticks(rotation=0)
counttable(temp)
df_temp = df
temp = df_temp[['Num. comorbidities', 'Sex']].groupby('Sex')['Num. comorbidities'].value_counts().unstack(0)
temp = temp.rename(columns={"M":"Male", "F": "Female"})
temp.plot.bar()
plt.xlabel("Number of comobidities", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Distribution of number of comorbidities", fontsize=16)
plt.xticks(rotation=0)
counttable(temp)
df_temp = df
temp = df_temp[['Num. comorbidities', 'Age group']].groupby('Num. comorbidities')['Age group'].value_counts().unstack(0)
temp = temp.reindex(index=df_comor['Age group'].cat.categories.values)
temp = temp.fillna(0).astype(int)
sns.heatmap(temp, cmap='Blues', annot=True, fmt="d");
```
### Symptoms
```
df.iloc[:,54].str.split(',').apply(lambda x: list(map(str.lower, x)).count('hypoxia') if type(x)==list else 0).sum()
temp = df.iloc[:, 20:54].sum()
temp.index.name = "Symptoms"
temp = pd.concat([temp, pd.Series(data={"Hypoxia": df.iloc[:,54].str.split(',').apply(lambda x: list(map(str.lower, x)).count('hypoxia') if type(x)==list else 0).sum()})])
temp2 = (temp/temp.sum()*100).copy()
temp3 = temp2[temp2>1]
temp3 = pd.concat([temp3.sort_values(ascending=False), pd.Series(data={"Other": temp2[temp2<=1].sum()})])
temp3.plot.pie()
plt.ylabel("")
counttable(temp).sort_values('Count', ascending=False)
df_temp = df
temp = df_temp[['Num. symptoms', 'Sex']].groupby('Sex')['Num. symptoms'].value_counts().unstack(0)
temp = temp.rename(columns={"M":"Male", "F": "Female"})
temp.plot.bar()
plt.xlabel("Number of symptoms", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Distribution of number of symptoms", fontsize=16)
plt.xticks(rotation=0)
temp.transpose()
df_temp = df
temp = df_temp[['Num. symptoms', 'Age group']].groupby('Num. symptoms')['Age group'].value_counts().unstack(0)
temp = temp.reindex(index=df_comor['Age group'].cat.categories.values)
temp = temp.fillna(0).astype(int)
plt.figure(figsize=(12, 4))
sns.heatmap(temp, cmap='Blues', annot=True, fmt="d")
plt.title("Heat map of number of cases", fontsize=15);
df_temp = df
temp = df_temp[['Num. symptoms', 'Num. comorbidities']].groupby('Num. symptoms')['Num. comorbidities'].value_counts().unstack(0)
temp = temp.fillna(0).astype(int)
plt.figure(figsize=(12, 4))
sns.heatmap(temp, cmap='Blues', annot=True, fmt="d")
plt.title("Heat map of number of cases", fontsize=15);
```
### Deaths, ICU hopitalizations and ventilation required
```
temp = df[['Ventilation', 'ICU', 'Dead', 'Sex']].groupby('Sex').agg(sum)
temp = temp.rename(columns={'Vent_flag':'Ventilation', 'ICU_flag':"ICU", 'Dead flag':'Dead'},
index={'M':'Male', 'F':'Female'})
temp.plot.bar()
plt.xlabel("Sex", fontsize=15)
plt.xticks(rotation=0)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Gender of patients", fontsize=16)
counttable(temp)
temp = df.groupby('Dead').agg('sum').iloc[:, 2:18].transpose().iloc[:, 1]
temp.index.name = "Comorbidity"
counttable(temp).sort_values('Percent', ascending=False)
df_temp = df[df['Dead']]
temp = df_temp[['Num. comorbidities', 'Sex']].groupby('Sex')['Num. comorbidities'].value_counts().unstack(0)
temp = temp.rename(columns={"M":"Male", "F": "Female"})
temp.plot.bar()
plt.xlabel("Number of comobidities", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Distribution of number of comorbidities", fontsize=16)
plt.xticks(rotation=0)
counttable(temp)
df_temp = df
temp = df_temp[['Num. symptoms', 'Dead']].groupby('Dead')['Num. symptoms'].value_counts().unstack(0)
temp.columns.name = "Dead"
temp.plot.bar()
plt.xlabel("Number of symptoms", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Distribution of number of symptoms", fontsize=16)
plt.xticks(rotation=0)
temp.transpose()
df_temp = df
temp = df_temp[['Num. symptoms', 'ICU']].groupby('ICU')['Num. symptoms'].value_counts().unstack(0)
temp.plot.bar()
plt.xlabel("Number of symptoms", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Distribution of number of symptoms", fontsize=16)
plt.xticks(rotation=0)
temp.transpose()
df_temp = df
temp = df_temp[['Num. symptoms', 'Ventilation']].groupby('Ventilation')['Num. symptoms'].value_counts().unstack(0)
temp.plot.bar()
plt.xlabel("Number of symptoms", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Distribution of number of symptoms", fontsize=16)
plt.xticks(rotation=0)
temp.transpose()
temp = df[['Ventilation', 'ICU', 'Dead', 'Age group']].groupby('Age group').agg(sum)
temp.index = temp.index.astype('object')
temp.plot.bar()
plt.xlabel("Age group", fontsize=15)
plt.ylabel("Number of cases", fontsize=15)
plt.title("Age distribution", fontsize=16)
counttable(temp)
temp = df[df['Dead']].groupby('Sex')['Age group'].value_counts().unstack(0)
temp = temp.reindex(index=df_comor['Age group'].cat.categories.values)
temp.plot.bar()
plt.xlabel("Age group", fontsize=15)
plt.ylabel("Number of deaths", fontsize=15)
plt.title("Age distribution by gender", fontsize=16)
counttable(temp)
df_temp = df[df['Dead']]
temp = df_temp[['Num. symptoms', 'Num. comorbidities']].groupby('Num. symptoms')['Num. comorbidities'].value_counts().unstack(0)
temp = temp.fillna(0).astype(int)
plt.figure(figsize=(12, 4))
sns.heatmap(temp, cmap='Blues', annot=True, fmt="d")
plt.title("Heat map of number of deaths", fontsize=15);
```
### Days to deaths
```
sns.displot(df, x='Days to death', col='Sex');
```
### Deaths over time
```
temp = df[df['Dead']].groupby('Sex')['Year Month']
temp = temp.value_counts().unstack(0).sort_index().fillna(0).fillna(method="pad")
fig, ax = plt.subplots(2, 1, figsize=(6, 6), sharex=True)
temp.plot.area(stacked=False, ax=ax[0])
ax[0].set_ylabel('Deaths', fontsize=15)
temp1 = (temp['Male'] )/(temp['Male'] + temp['Female'])
temp1.plot(ax=ax[1])
plt.hlines(0.5, temp1.index.values.min(), temp1.index.values.max(), linestyle='dotted', color='black')
plt.hlines(temp1.mean(), temp1.index.values.min(), temp1.index.values.max(), linestyle='dashed', color='tab:blue')
ax[1].set_xlabel('Date', fontsize=15)
ax[1].set_ylim([0, 1])
ax[1].set_ylabel('Proportion of males', fontsize=15)
fig.suptitle("Number of deaths by gender", fontsize=16);
temp = df[df['Dead']].groupby('Sex')['Year Month']
temp = temp.value_counts().unstack(0).sort_index().fillna(0).fillna(method="pad")
fig, ax = plt.subplots(2, 1, figsize=(6, 6), sharex=True)
temp.plot.area(stacked=False, ax=ax[0])
ax[0].set_ylabel('Deaths', fontsize=15)
temp1 = (temp['Male'] )/(temp['Male'] + temp['Female'])
temp1.plot(ax=ax[1])
plt.hlines(0.5, temp1.index.values.min(), temp1.index.values.max(), linestyle='dotted', color='black')
plt.hlines(temp1.mean(), temp1.index.values.min(), temp1.index.values.max(), linestyle='dashed', color='tab:blue')
ax[1].set_xlabel('Date', fontsize=15)
ax[1].set_ylim([0, 1])
ax[1].set_ylabel('Proportion of males', fontsize=15)
fig.suptitle("Number of deaths by gender", fontsize=16);
```
| github_jupyter |
<!--
# Copyright Amazon.com, Inc. or its affiliates. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License").
# You may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
-->
# Creating EBS Volume for Teams to use
## Content
1. Admin Operations
1. [Parameters](#Parameters)
2. [Cleanup](#Cleanup)
3. [Creating the EBS Volume](#Creating-the-EBS-Volume)
4. [Creating the K8 Volume](#Creating-the-K8-Volume)
2. User Operations
1. [Creating the K8 Volume Claim](#Creating-the-K8-Volume-Claim)
2. [Creating the Profile with the required AZ](#Creating-the-Profile-with-the-required-AZ)
3. [Running the container](#Running-the-container)
---
---
```
from aws_orbit_sdk import controller
from aws_orbit_sdk.magics.orbit import OrbitWorkbenchMagics
import json
import boto3
from aws_orbit_sdk.common import get_workspace
# we will need the team kms key from workspace
workspace = get_workspace()
team_kms_key = workspace['TeamKmsKeyArn']
image = workspace['BaseImageAddress']
%cd ebs
env_name = %env AWS_ORBIT_ENV
team_name = %env AWS_ORBIT_TEAM_SPACE
region = %env AWS_DEFAULT_REGION
(env_name,team_name,region)
```
## Parameters
```
pv_name = 'my-pv1'
pvc_name = 'my-pvc1'
az = str(region+'a')
volume_size = 20 #gb
```
## Cleanup
```
!kubectl delete pvc $pvc_name --force
!kubectl delete pv $pv_name --force
```
## Creating the EBS Volume
```
!echo aws ec2 create-volume --availability-zone=$az --encrypted \
--size=$volume_size --volume-type=gp2 --kms-key-id $team_kms_key
res = !aws ec2 create-volume --availability-zone=$az --encrypted \
--size=$volume_size --volume-type=gp2
res
ebs_vol = json.loads('\n'.join(res))
ebs_vol
volume_id = ebs_vol['VolumeId']
volume_id
!aws ec2 wait volume-available --volume-ids $volume_id
```
## Creating the K8 Volume
```
with open("pv.yaml", "w") as file:
file.write("""
apiVersion: v1
kind: PersistentVolume
metadata:
name: {pv_name}
labels:
type: {pv_name}
spec:
storageClassName: ebs-{team_name}-gp2
capacity:
storage: 5Gi
accessModes:
- ReadWriteOnce
awsElasticBlockStore:
volumeID: {volume_id}
fsType: xfs
""".format(team_name=team_name,pv_name=pv_name,volume_id=volume_id)
)
!cat pv.yaml
!kubectl apply -f pv.yaml
```
## User Section
## Creating the K8 Volume Claim
```
with open("pvc.yaml", "w") as file:
file.write("""
kind: PersistentVolumeClaim
apiVersion: v1
metadata:
name: {pvc_name}
labels:
type: {pvc_name}
spec:
accessModes:
- ReadWriteOnce
storageClassName: ebs-{team_name}-gp2
resources:
requests:
storage: 5Gi
selector:
matchLabels:
type: {pv_name}
""".format(team_name=team_name,pv_name=pv_name,pvc_name=pvc_name)
)
!cat pvc.yaml
!kubectl apply -f pvc.yaml
```
## Creating the PodSetting with the required AZ
```
import json
customname = "orbit-custom-volumes-"+team_name
with open("podsetting_ebs.yaml", "w") as file:
file.write("""
kind: PodSetting
apiVersion: orbit.aws/v1
metadata:
labels:
orbit/env: {env_name}
orbit/space: team
orbit/team: {team_name}
name: {customname}
namespace: {team_name}
spec:
affinity:
nodeAffinity:
preferredDuringSchedulingIgnoredDuringExecution:
- preference:
matchExpressions:
- key: topology.kubernetes.io/zone
operator: In
values:
- {az}
weight: 1
containerSelector:
jsonpath: metadata.labels.app
desc: Example EFS orbit-{customname}
env:
- name: custom_name
value: custom_value
image: >-
{image}
podSelector:
matchExpressions:
- key: orbit/{customname}
operator: Exists
resources:
limits:
cpu: '1.0'
memory: 1Gi
requests:
cpu: '1.0'
memory: 1Gi
securityContext:
runAsUser: 1000
volumeMounts:
- mountPath: /ebs
name: ebs-volume
volumes:
- name: ebs-volume
persistentVolumeClaim:
claimName: {pvc_name}
""".format(team_name=team_name,env_name=env_name,pvc_name=pvc_name,customname=customname,image=image,az=az)
)
!kubectl apply -f podsetting_ebs.yaml -n {team_name}
```
## Running the container
```
run = {
"tasks": [
{
"notebookName": "test-ebs.ipynb",
"sourcePath": "shared/samples/notebooks/M-Admin/ebs",
"targetPath": "shared/regression/notebooks/M-Admin/ebs",
"params": {
"test" : "1"
}
}
],
"compute": {
"container" : {
"p_concurrent": "1"
},
"node_type": "ec2",
"podsetting":customname,
"labels": {
"my-jobid": "1"
}
}
}
with open("run.json", 'w') as f:
json.dump(run, f)
%%time
!orbit run notebook --env $env_name --team $team_name --user testing --wait --tail-logs run.json
```
## Cleanup
```
# Using our label to delete the job
!kubectl delete job -l my-jobid=1
!kubectl delete podsetting -n {team_name} {customname}
!kubectl delete pvc $pvc_name --force
!kubectl delete pv $pv_name --force
!aws ec2 delete-volume --volume-id $volume_id
```
| github_jupyter |
An example showing univariate feature selection.
Noisy (non informative) features are added to the iris data and univariate feature selection is applied. For each feature, we plot the p-values for the univariate feature selection and the corresponding weights of an SVM. We can see that univariate feature selection selects the informative features and that these have larger SVM weights.
In the total set of features, only the 4 first ones are significant. We can see that they have the highest score with univariate feature selection. The SVM assigns a large weight to one of these features, but also Selects many of the non-informative features. Applying univariate feature selection before the SVM increases the SVM weight attributed to the significant features, and will thus improve classification.
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
### Version
```
import sklearn
sklearn.__version__
```
### Imports
This tutorial imports [SelectPercentile](http://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.SelectPercentile.html#sklearn.feature_selection.SelectPercentile) and [f_classif](http://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.f_classif.html#sklearn.feature_selection.f_classif).
```
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
from sklearn import datasets, svm
from sklearn.feature_selection import SelectPercentile, f_classif
```
### Calculations
Import some data
```
# The iris dataset
iris = datasets.load_iris()
# Some noisy data not correlated
E = np.random.uniform(0, 0.1, size=(len(iris.data), 20))
# Add the noisy data to the informative features
X = np.hstack((iris.data, E))
y = iris.target
X_indices = np.arange(X.shape[-1])
```
### Plot Results
Univariate feature selection with F-test for feature scoring We use the default selection function: the 10% most significant features
```
selector = SelectPercentile(f_classif, percentile=10)
selector.fit(X, y)
scores = -np.log10(selector.pvalues_)
scores /= scores.max()
trace = go.Bar(x=X_indices - .45,
y=scores, width=.2,
name=r'Univariate score (<i>-Log(p_{value})</i>)',
marker=dict(color='darkorange',
line=dict(color='black', width=1))
)
py.iplot([trace])
```
Compare to the weights of an SVM
```
clf = svm.SVC(kernel='linear')
clf.fit(X, y)
svm_weights = (clf.coef_ ** 2).sum(axis=0)
svm_weights /= svm_weights.max()
trace1 = go.Bar(x=X_indices - .25,
y=svm_weights,
name='SVM weight',
marker=dict(color='navy',
line=dict(color='black', width=1))
)
clf_selected = svm.SVC(kernel='linear')
clf_selected.fit(selector.transform(X), y)
svm_weights_selected = (clf_selected.coef_ ** 2).sum(axis=0)
svm_weights_selected /= svm_weights_selected.max()
trace2 = go.Bar(x=X_indices[selector.get_support()] - .05,
y=svm_weights_selected,
name='SVM weights after selection',
marker=dict(color='cyan',
line=dict(color='black', width=1))
)
data = [trace1, trace2]
layout = go.Layout(title="Comparing feature selection",
xaxis=dict(title='Feature number'),
barmode='grouped'
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig)
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'Univariate Feature Selection.ipynb', 'scikit-learn/plot-feature-selection/', 'Univariate Feature Selection | plotly',
' ',
title = 'Univariate Feature Selection | plotly',
name = 'Univariate Feature Selection',
has_thumbnail='true', thumbnail='thumbnail/ufs.jpg',
language='scikit-learn', page_type='example_index',
display_as='feature_selection', order=6,
ipynb= '~Diksha_Gabha/3093')
```
| github_jupyter |
# AWS Glue Notebook for Serverless Data Lake Workshop
This notebook contains the PySpark scripts run in AWS Glue to transform the data in the data lake. Each section refers a section in the lab.
## Initialization
The first two sections initialize the Spark environment and only need to be run once. The first block may take a few seconds as it negotiates the Spark session.
```
## @ Import the AWS Glue libraries, pySpark we'll need
import sys
from awsglue.transforms import *
from awsglue.utils import getResolvedOptions
from pyspark.context import SparkContext
from awsglue.context import GlueContext
from awsglue.job import Job
from pyspark.sql.functions import *
from awsglue.dynamicframe import DynamicFrame
## @ set up a single GlueContext.
sc = SparkContext.getOrCreate()
glueContext = GlueContext(sc)
```
## Lab - Transform / Decode data with AWS Glue
The first script is a straight forward transformation of the user activity table. The request column contains a number of fields embedded within it
```
GET /petstore/Cats/Treats
```
The column is split out to get the request type, top level, top page, and sub page.
The timestamp field is also parsed out to extract the date components: date, time, month, and year.
17/Jan/2018:10:43:54
Date: 01/17/2018
Time: 10:43:54
Year: 2018
Month: 10
| ip_address | username | timestamp | request | http | bytes | requesttype | topdomain | subpage | date | time | year | month | toppage |
|------|------|------|------|------|------|------|------|------|------|------|------|------|------|
| 0.32.193.205 | grldmccfdm8 | 11/Oct/2018:23:36:54 | DELETE | /petstore/Bird/Treats | 500 | 927 | DELETE | petstore | Treats | 10/11/2018 | 23:36:54 | 2018 | 10 | Bird |
| 0.32.193.205 | grldmccfdm8 | 6/Jun/2017:13:03:54 | PUT | /petstore/Bird/Food | 500 | 927 | DELETE | petstore | Food | 06/06/2017 | 13:03:54 | 2017 | 6 | Bird |
The data is then written out in a partitioned parquet format.
*This process takes 3-5 minutes.*
```
spark = glueContext.spark_session
job = Job(glueContext)
job.init('^stackname^-exercise1')
## @ create the Glue DynamicFrame from table schema. A DynamicFrame is similar to a DataFrame, except that each record is
## @ self-describing, so no schema is required initially.
useractivity = glueContext.create_dynamic_frame.from_catalog(database = "weblogs", table_name = "useractivity", transformation_ctx = "useractivity")
## @ ApplyMapping is one of the built in transforms that maps source columns and data types from a DynamicFrame to target columns
## @ and data types in a returned DynamicFrame. You specify the mapping argument, which is a list of tuples that contain source column,
## @ source type, target column, and target type.
useractivityApplyMapping = ApplyMapping.apply(frame = useractivity, mappings = [("ip_address", "string", "ip_address", "string"), ("username", "string", "username", "string"), ("timestamp", "string", "timestamp", "string"), ("request", "string", "request", "string"), ("http", "long", "http", "long"), ("bytes", "long", "bytes", "long")], transformation_ctx = "applymapping1")
## @ ResolveChoice is another built in transform that you can use to specify how a column should be handled when it contains values of
## @ multiple types. You can choose to either cast the column to a single data type, discard one or more of the types, or retain all
## @ types in either separate columns or a structure. You can select a different resolution policy for each column or specify a global
## @ policy that is applied to all columns.
resolvechoice2 = ResolveChoice.apply(frame = useractivityApplyMapping, choice = "make_struct", transformation_ctx = "resolvechoice2")
## @ DropNullFields transform removes null fields from a DynamicFrame. The output DynamicFrame does not contain fields of the null type
## @ in the schema.
useractivity = DropNullFields.apply(frame = resolvechoice2, transformation_ctx = "dropnullfields3")
## @ We will leverage PySpark functions to manipulate our data, starting with converting glue DynamicFrame to DataFrame
dataframe0 = DynamicFrame.toDF(useractivity)
## @ Use PySpark functions to split request columns on '/'
split_column = split(dataframe0['request'], '/')
dataframe0 = dataframe0.withColumn('requesttype', split_column.getItem(0))
dataframe0 = dataframe0.withColumn('topdomain', split_column.getItem(1))
dataframe0 = dataframe0.withColumn('toppage', split_column.getItem(2))
dataframe0 = dataframe0.withColumn('subpage', split_column.getItem(3))
## @ split timestamp column into date, time, year and month
dataframe0 = dataframe0.withColumn('date',date_format(from_unixtime(unix_timestamp('timestamp', 'd/MMM/yyyy:HH:mm:ss')), 'MM/dd/yyy'))
dataframe0 = dataframe0.withColumn('time',date_format(from_unixtime(unix_timestamp('timestamp', 'd/MMM/yyyy:HH:mm:ss')), 'HH:mm:ss'))
dataframe0 = dataframe0.withColumn('year', year(from_unixtime(unix_timestamp('timestamp', 'd/MMM/yyyy:HH:mm:ss'))))
dataframe0 = dataframe0.withColumn('month', month(from_unixtime(unix_timestamp('timestamp', 'd/MMM/yyyy:HH:mm:ss'))))
## @ convert dataframe to glue DynamicFrame and write the output in Parquet format partitioned on toppage column
useractivity = DynamicFrame.fromDF(dataframe0, glueContext, "name1")
writeUseractivityToS3 = glueContext.write_dynamic_frame.from_options(frame = useractivity, connection_type = "s3", connection_options = {"path": 's3://^ingestionbucket^/weblogs/useractivityconverted', "partitionKeys" :["toppage"]}, format = "parquet", transformation_ctx = "writeUseractivityToS3")
job.commit()
dataframe0.show()
```
## Results
To to the S3 bucket to view the results of the transformation and continue with the lab instructions.
## Lab - Join and relationalize data with AWS Glue
In this lab we will take two different datasets from different source systems and merge them to prepare a table that combines both useractivity and user profile datasets.
```
job = Job(glueContext)
job.init('^stackname^-exercise2')
## @ useractivity dynamicframe
useractivity = glueContext.create_dynamic_frame.from_catalog(database = "weblogs", table_name = "useractivity", transformation_ctx = "useractivity")
## @ applymappings to the dynamicframe to make sure we have the correct data types and column names
applymapping1 = ApplyMapping.apply(frame = useractivity, mappings = [("ip_address", "string", "ip_address", "string"), ("username", "string", "username", "string"), ("timestamp", "string", "timestamp", "string"), ("request", "string", "request", "string"), ("http", "long", "http", "long"), ("bytes", "long", "bytes", "long")], transformation_ctx = "applymapping1")
## @ resolve any issues with column data types
resolvechoice2 = ResolveChoice.apply(frame = applymapping1, choice = "make_struct", transformation_ctx = "resolvechoice2")
## @ drop any null fields
useractivity = DropNullFields.apply(frame = resolvechoice2, transformation_ctx = "useractivity")
## @ create userprofile dynamicframe
userprofile = glueContext.create_dynamic_frame.from_catalog(database="weblogs", table_name="userprofile")
## @ we will only keep the fields that we want and drop the rest and rename username to dy_username
userprofile = userprofile.drop_fields(['cc', 'password', 'ssn', 'email', 'phone','ip_address'])
userprofile = userprofile.rename_field('username','dy_username')
## @ as the data types in different datasets are different we are going to convert all column to string
## @ The Glue build in transform ApplyMapping, Maps source columns and data types from a DynamicFrame to target columns and data types
## @ in a returned DynamicFrame. You specify the mapping argument, which is a list of tuples that contain source column, source type,
## @ target column, and target type. In the below case we are converting the data types for zip and age to string and updating the column
## @ names for first_name & last_name
userprofile = ApplyMapping.apply(frame = userprofile,
mappings = [("first_name", "string", "firstname", "string"),
("dy_username", "string", "dy_username", "string"),
("zip", "bigint", "zip", "string"),
("age", "bigint", "age", "string"),
("gender", "string", "gender", "long"),
("last_name", "string", "lastname", "long")
], transformation_ctx = "userprofile")
## @join useractivity and userprofile datasets to create one file and drop the duplicate column dy_username
joined = Join.apply(userprofile, useractivity, 'dy_username', 'username').drop_fields(['dy_username'])
glueContext.write_dynamic_frame.from_options(frame = joined,
connection_type = "s3",
connection_options = {"path": 's3://^ingestionbucket^/weblogs/joindatasets'},
format = "parquet")
job.commit()
print 'Job Complete'
df = DynamicFrame.toDF(joined)
df.show()
```
## Results
Continue with the lab to explore the resulting table in the data lake.
## Create a UDF to simplify apply a hash function to columns
```
from pyspark.sql.functions import udf
from pyspark.sql.types import StringType
import hashlib
from dateutil.parser import parse
def hash_cc(s):
return hashlib.sha256(s).hexdigest()
job = Job(glueContext)
job.init('^stackname^-exercise3')
datasource0 = glueContext.create_dynamic_frame.from_catalog(database = "weblogs", table_name = "userprofile", transformation_ctx = "datasource0")
## @convert glue DynamicFrame to DataFrame to manipulate the columns
dataframe0 = DynamicFrame.toDF(datasource0)
hash_cc_f = udf(lambda x: hash_cc(x), StringType())
dataframe0 = dataframe0.withColumn("hash_cc", hash_cc_f(dataframe0["cc"])).withColumn("hash_ssn", hash_cc_f(dataframe0["ssn"]))
dataframe0 = dataframe0.drop('cc').drop('ssn').drop('password')
## @convert dataframe to glue DynamicFrame and write the output in Parquet format
datasource1 = DynamicFrame.fromDF(dataframe0, glueContext, "name1")
datasink4 = glueContext.write_dynamic_frame.from_options(frame = datasource1, connection_type = "s3", connection_options = {"path": 's3://^ingestionbucket^/weblogs/userprofile-secure'}, format = "parquet", transformation_ctx = "datasink4")
job.commit()
print 'Job Complete'
dataframe0.show()
```
# Exercise 4 - Lookup
You are on your own now! Using you pyspark skills, create a simple UDF that performs a lookup on the data.
In this case, we're going create a lookup for the geocode of the IP Address. Instead of calling out to a geocoding service, we'll just return US if the first value is less than 100 and UK if the value is 100 or greater.
Using this function, create a lookup that creates a copy of the useractivity table that includes the country.
```
def geocode(ip):
if int(ip.split('.')[0]) < 100:
return "US"
else:
return "UK"
job = Job(glueContext)
job.init('Job4')
# Write Transformation Code here
job.commit()
```
| github_jupyter |
```
import os
import sys
import glob
import torch
import numpy as np
import pydicom as dicom
from skimage.draw import polygon
import matplotlib.pyplot as plt
%matplotlib inline
def read_structure(structure):
contours = []
for i, ri in enumerate(structure.ROIContourSequence):
contour = {}
#return ri
contour['color'] = ri.ROIDisplayColor
contour['number'] = ri.ReferencedROINumber
#contour['name'] = ri.ROIName
#assert contour['number'] == ri.ROINumber
contour['contours'] = [s.ContourData for s in ri.ContourSequence]
contours.append(contour)
return contours
def get_mask(contours, slices):
z = np.around(np.array([float(s.ImagePositionPatient[2]) for s in slices]), 1)
pos_r = slices[0].ImagePositionPatient[1]
spacing_r = slices[0].PixelSpacing[1]
pos_c = slices[0].ImagePositionPatient[0]
spacing_c = slices[0].PixelSpacing[0]
label = np.zeros_like(image, dtype=np.uint8)
for con in contours:
num = int(con['number'])
for c in con['contours']:
nodes = np.array(c).reshape((-1, 3))
assert np.amax(np.abs(np.diff(nodes[:, 2]))) == 0
#z_index = z.index(nodes[0, 2])
try:
z_index = np.where(z == float(np.around(nodes[0, 2], 1)))[0] # fix in later comments -JH
except:
print(z)
print(nodes[0,2])
raise
r = (nodes[:, 1] - pos_r) / spacing_r
c = (nodes[:, 0] - pos_c) / spacing_c
rr, cc = polygon(r, c)
label[rr, cc, z_index] = num
colors = tuple(np.array([con['color'] for con in contours]) / 255.0)
return label, colors
train_data_path = '/mnt/USB/AAPM17CTSegChallenge/LCTSC/DOI' #"./DOI" # point to our data -JH
preprocessing_imgdir = "/home/ygx/data/aapm17/preprocessing/imgs"
preprocessing_labeldir = "/home/ygx/data/aapm17/preprocessing/labels"
train_patients = [os.path.join(train_data_path, name)
for name in os.listdir(train_data_path) if os.path.isdir(os.path.join(train_data_path, name))]
print(f'First Patient: {train_patients[0]}')
for i, patient in enumerate(train_patients):
print(f"Patient {i}: {patient}")
image = None
slices = None
contours = None
for subdir, dirs, files in os.walk(patient):
dcms = glob.glob(os.path.join(subdir, "*.dcm"))
if len(dcms) == 1:
structure = dicom.read_file(os.path.join(subdir, files[0]))
contours = read_structure(structure)
elif len(dcms) > 1:
slices = [dicom.read_file(dcm) for dcm in dcms]
slices.sort(key = lambda x: float(x.ImagePositionPatient[2]))
image = np.stack([s.pixel_array for s in slices], axis=-1)
if image is not None:
torch.save(torch.Tensor(image.astype(np.float32)), f"{preprocessing_imgdir}/image_{i}.pth")
if contours is not None:
label, colors = get_mask(contours, slices)
print(f'label: {label.shape}')
print(f'color: {type(colors)}, len: {len(colors)}, First index: {colors[0]}')
label_int = label.astype(np.uint8)
print(f'Integer label: {label_int.shape}')
torch.save(torch.Tensor(label_int), f"{preprocessing_labeldir}/label_{i}.pth")
break
x = np.array([0., 1.5, .1])
x
x.astype(np.uint)
```
| github_jupyter |
# Interactive experimentation
```
!pip install --upgrade lightgbm scikit-learn pandas adlfs
```
## Setup cloud tracking
```
import mlflow
from azureml.core import Workspace
ws = Workspace.from_config()
mlflow.set_tracking_uri(ws.get_mlflow_tracking_uri())
mlflow.set_experiment("untitled")
```
## Load data
You can read directly from public URIs into Pandas. For private Blob or ADLS data, consider using [adlfs](https://github.com/dask/adlfs).
```
data_uri = "https://azuremlexamples.blob.core.windows.net/datasets/iris.csv"
import pandas as pd
df = pd.read_csv(data_uri)
df.head()
```
## Define functions
```
# imports
import time
import lightgbm as lgb
from sklearn.metrics import log_loss, accuracy_score
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
# define functions
def preprocess_data(df):
X = df.drop(["species"], axis=1)
y = df["species"]
enc = LabelEncoder()
y = enc.fit_transform(y)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=42
)
return X_train, X_test, y_train, y_test, enc
def train_model(params, num_boost_round, X_train, X_test, y_train, y_test):
t1 = time.time()
train_data = lgb.Dataset(X_train, label=y_train)
test_data = lgb.Dataset(X_test, label=y_test)
model = lgb.train(
params,
train_data,
num_boost_round=num_boost_round,
valid_sets=[test_data],
valid_names=["test"],
)
t2 = time.time()
return model, t2 - t1
def evaluate_model(model, X_test, y_test):
y_proba = model.predict(X_test)
y_pred = y_proba.argmax(axis=1)
loss = log_loss(y_test, y_proba)
acc = accuracy_score(y_test, y_pred)
return loss, acc
```
## Run a trial
```
# preprocess data
X_train, X_test, y_train, y_test, enc = preprocess_data(df)
# set training parameters
params = {
"objective": "multiclass",
"num_class": 3,
"learning_rate": 0.1,
"metric": "multi_logloss",
"colsample_bytree": 1.0,
"subsample": 1.0,
"seed": 42,
}
num_boost_round = 32
# start run
run = mlflow.start_run()
# enable automatic logging
mlflow.lightgbm.autolog()
# train model
model, train_time = train_model(
params, num_boost_round, X_train, X_test, y_train, y_test
)
mlflow.log_metric("training_time", train_time)
# evaluate model
loss, acc = evaluate_model(model, X_test, y_test)
mlflow.log_metrics({"loss": loss, "accuracy": acc})
# end run
mlflow.end_run()
```
| github_jupyter |
# AttnGAN
## Fine-Grained Text to Image Generation with Attentional Generative Adversarial Networks
http://openaccess.thecvf.com/content_cvpr_2018/papers/Xu_AttnGAN_Fine-Grained_Text_CVPR_2018_paper.pdf
https://github.com/taoxugit/AttnGAN
---
## TODO
- run le code en debug dans IntelliJ
- indiquer les shapes dans le code de ce notebook
- copier et analyser la fonction train DAMSM
---

- **z** - a noise vector usually sampled from a standard normal distribution.
- **F**<sup>*ca*</sup> - represents the Conditioning Augmentation that converts the sentence vector **E** to the conditioning vector.
- **E** is a global sentence vector, and **e** is the matrix of word vectors.
- **F**<sub>i</sub><sup>*attn*</sup> is the proposed attention model at the i<sup>*th*</sup> stage of the AttnGAN.
- **F**<sup>*ca*</sup>, **F**<sub>i</sub><sup>*attn*</sup>, **F**<sub>i</sub> , and **G**<sub>i</sub> are neural networks.
- The attention model **F**<sup>*attn*</sup> has two inputs:
the word features **e** and the image features from the previous hidden layer h.
The word features are first converted into the common semantic space of the image features by adding a new perceptron layer.
Then, a word-context vector is computed for each sub-region of the image based on its hidden features **h** (query). Each column of h is a feature vector of a sub-region of the image. For the j<sup>th</sup> sub-region, its word-context vector is a dynamic representation of word vectors relevant to h<sub>j</sub>.
Finally, image features and the corresponding word-context features are combined to generate images at the next stage.
# Deep Attentional Multimodal Similarity Model (DAMSM)
The DAMSM **learns two neural networks** that map sub-regions of the image and words of the sentence to a common semantic space, thus measures the **image-text similarity** at the word level to compute a fine-grained **loss for image generation**.
## Text Encoder
A **bi-directional LSTM** that extracts semantic vectors from the text description.

**word features**: The feature matrix of words indicated by **e**. Its i<sup>*th*</sup> column **e**<sub>i</sub> is feature vector for the i<sup>*th*</sup> word.
**sentence feature**: The last hidden states of the bi-directional LSTM are concatenated to be the global sentence vector, denoted by **E**.
```
class RNN_ENCODER(nn.Module):
def __init__(self, ntoken=2932, ninput=300, drop_prob=0.5, nhidden=256, nlayers=1, bidirectional=True):
'''
ntoken -- size of the dictionary computed from the dataset captions; 2932 tokens in FashionGen2
nhidden -- TEXT.EMBEDDING_DIM = 256 for FashionGen2
ninput -- size of each embedding vector (300 by default)
nlayers -- Number of recurrent layers
'''
self.n_steps = cfg.TEXT.WORDS_NUM # 10 in FashionGen2 (caption max number of words)
# ...
if bidirectional:
self.num_directions = 2
# number of features in the hidden state (hidden nodes in the LSTM layer)
self.nhidden = nhidden // self.num_directions # 128 = 256 / 2 (1 Bi-LSTM layer of 128 nodes)
def define_module(self):
# ...
self.encoder = nn.Embedding(self.ntoken, self.ninput) # nn.Embedding(2932, 300)
if self.rnn_type == 'LSTM':
self.rnn = nn.LSTM(self.ninput, self.nhidden,
self.nlayers, batch_first=True,
dropout=self.drop_prob,
bidirectional=self.bidirectional)
def forward(self, captions, cap_lens, hidden, mask=None):
# input: torch.LongTensor of size batch x n_steps --> emb: batch x n_steps x ninput
# input (bs, 10) --> emb (bs, 10, 300)
emb = self.drop(self.encoder(captions))
# Returns: a PackedSequence object
cap_lens = cap_lens.data.tolist()
# cap_lens -> <type 'list'>: [10, 10, 10, 10, 10, 10, 10, 9, 9, 8, 8, 8, 7, 7, 7,
# 7, 7, 7, 7, 7, 7, 6, 6, 6, 6, 6, 6, 6, 5, 5, 5, 5]
emb = pack_padded_sequence(emb, cap_lens, batch_first=True)
# emb -> PackedSequence(tensor([[-0.0112, 0.0000, -0.0002, ..., 0.5088, 0.0000, -0.1240],
# [-0.0000, 0.0782, 0.3802, ..., -0.3164, 0.4351, 0.0748],
# [-0.1120, -0.0000, -0.1069, ..., -0.0000, 0.0000, -0.0000],
# ...,
# [ 0.0000, -0.1407, -0.6452, ..., 0.0000, -0.0000, 0.4360],
# [-0.0000, -0.0074, 0.0000, ..., 0.1719, 0.0000, 0.0082],
# [ 0.0000, -0.0000, -0.0000, ..., -0.0000, 0.0000, 2.9084]]
# #hidden and memory (num_layers * num_directions, batch, hidden_size):
# tensor containing the initial hidden state for each element in batch.
# #output (batch, seq_len, hidden_size * num_directions) or a PackedSequence object:
# tensor containing output features (h_t) from the last layer of RNN
output, hidden = self.rnn(emb, hidden)
# PackedSequence object --> (batch, seq_len, hidden_size * num_directions)
output = pad_packed_sequence(output, batch_first=True)[0]
# output = self.drop(output) --> batch x hidden_size * num_directions x seq_len
words_emb = output.transpose(1, 2)
# words_emb.shape --> torch.Size([32, 256, 10])
# --> batch x num_directions * hidden_size
if self.rnn_type == 'LSTM':
sentence_emb = hidden[0].transpose(0, 1).contiguous()
sentence_emb = sentence_emb.view(-1, self.nhidden * self.num_directions) # (-1, 128*2)
# words_emb.shape --> torch.Size([32, 256, 10])
# sentence_emb --> torch.Size([32, 256])
return words_emb, sentence_emb
RNN_ENCODER(
(encoder): Embedding(2932, 300)
(drop): Dropout(p=0.5)
(rnn): LSTM(300, 128, batch_first=True, dropout=0.5, bidirectional=True)
)
```
---
# Image Encoder
A pretrained **Inception v3** CNN (input image of 299×299) that maps images to semantic vectors.

The **intermediate layers** of the CNN learn **local features** of different **sub-regions of the image**, while the later layers learn global features of the image.
We extract the **local feature** matrix **f** ∈ R768⇥289 (reshaped from 768×17×17, 17x17=289) from the **“mixed 6e” layer** of Inception-v3.
Each column of **f** is the **feature vector** of a **sub-region of the image**.
**f** shape is (768, 289):
768 is the dimension of the local feature vector, and
289 is the number of sub-regions in the image.
```
class CNN_ENCODER(nn.Module):
def __init__(self, nef):
'''
nef <-- TEXT.EMBEDDING_DIM = 256 (does 'nef' stands for 'Number Embedding Features'?)
'''
super(CNN_ENCODER, self).__init__()
if cfg.TRAIN.FLAG:
self.nef = nef
else:
self.nef = 256 # define a uniform ranker
model = models.inception_v3()
url = 'https://download.pytorch.org/models/inception_v3_google-1a9a5a14.pth'
model.load_state_dict(model_zoo.load_url(url))
for param in model.parameters():
param.requires_grad = False
self.define_module(model)
self.init_trainable_weights()
def define_module(self, model):
self.Conv2d_1a_3x3 = model.Conv2d_1a_3x3
# ...
self.Mixed_5d = model.Mixed_5d
self.Mixed_6a = model.Mixed_6a
self.Mixed_6b = model.Mixed_6b
# ...
self.emb_features = conv1x1(768, self.nef)
self.emb_cnn_code = nn.Linear(2048, self.nef)
def init_trainable_weights(self):
initrange = 0.1
self.emb_features.weight.data.uniform_(-initrange, initrange)
self.emb_cnn_code.weight.data.uniform_(-initrange, initrange)
def forward(self, x):
features = None
# --> fixed-size input: batch x 3 x 299 x 299
x = nn.Upsample(size=(299, 299), mode='bilinear')(x)
# 299 x 299 x 3
x = self.Conv2d_1a_3x3(x)
# 149 x 149 x 32
x = self.Conv2d_2a_3x3(x)
# ...
x = self.Mixed_6e(x)
# 17 x 17 x 768
# --- image region features ---
features = x
# 17 x 17 x 768
x = self.Mixed_7a(x)
# 8 x 8 x 1280
x = self.Mixed_7b(x)
# 8 x 8 x 2048
x = self.Mixed_7c(x)
# 8 x 8 x 2048
x = F.avg_pool2d(x, kernel_size=8)
# 1 x 1 x 2048
x = x.view(x.size(0), -1)
# 2048
# --- global image features ---
cnn_code = self.emb_cnn_code(x) # self.emb_cnn_code = nn.Linear(2048, self.nef)
# 512
if features is not None:
features = self.emb_features(features) # self.emb_features = conv1x1(768, self.nef)
return features, cnn_code
CNN_ENCODER(
(Conv2d_1a_3x3): BasicConv2d(
(conv): Conv2d(3, 32, kernel_size=(3, 3), stride=(2, 2), bias=False)
(bn): BatchNorm2d(32, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(Conv2d_2a_3x3): BasicConv2d(
(conv): Conv2d(32, 32, kernel_size=(3, 3), stride=(1, 1), bias=False)
(bn): BatchNorm2d(32, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(Conv2d_2b_3x3): BasicConv2d(
(conv): Conv2d(32, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn): BatchNorm2d(64, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(Conv2d_3b_1x1): BasicConv2d(
(conv): Conv2d(64, 80, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(80, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(Conv2d_4a_3x3): BasicConv2d(
(conv): Conv2d(80, 192, kernel_size=(3, 3), stride=(1, 1), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(Mixed_5b): InceptionA(
(branch1x1): BasicConv2d(
(conv): Conv2d(192, 64, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(64, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch5x5_1): BasicConv2d(
(conv): Conv2d(192, 48, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(48, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch5x5_2): BasicConv2d(
(conv): Conv2d(48, 64, kernel_size=(5, 5), stride=(1, 1), padding=(2, 2), bias=False)
(bn): BatchNorm2d(64, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch3x3dbl_1): BasicConv2d(
(conv): Conv2d(192, 64, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(64, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch3x3dbl_2): BasicConv2d(
(conv): Conv2d(64, 96, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn): BatchNorm2d(96, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch3x3dbl_3): BasicConv2d(
(conv): Conv2d(96, 96, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn): BatchNorm2d(96, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch_pool): BasicConv2d(
(conv): Conv2d(192, 32, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(32, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
)
...
(Mixed_6e): InceptionC(
(branch1x1): BasicConv2d(
(conv): Conv2d(768, 192, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch7x7_1): BasicConv2d(
(conv): Conv2d(768, 192, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch7x7_2): BasicConv2d(
(conv): Conv2d(192, 192, kernel_size=(1, 7), stride=(1, 1), padding=(0, 3), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch7x7_3): BasicConv2d(
(conv): Conv2d(192, 192, kernel_size=(7, 1), stride=(1, 1), padding=(3, 0), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch7x7dbl_1): BasicConv2d(
(conv): Conv2d(768, 192, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch7x7dbl_2): BasicConv2d(
(conv): Conv2d(192, 192, kernel_size=(7, 1), stride=(1, 1), padding=(3, 0), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch7x7dbl_3): BasicConv2d(
(conv): Conv2d(192, 192, kernel_size=(1, 7), stride=(1, 1), padding=(0, 3), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch7x7dbl_4): BasicConv2d(
(conv): Conv2d(192, 192, kernel_size=(7, 1), stride=(1, 1), padding=(3, 0), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch7x7dbl_5): BasicConv2d(
(conv): Conv2d(192, 192, kernel_size=(1, 7), stride=(1, 1), padding=(0, 3), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch_pool): BasicConv2d(
(conv): Conv2d(768, 192, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
)
...
(Mixed_7c): InceptionE(
(branch1x1): BasicConv2d(
(conv): Conv2d(2048, 320, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(320, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch3x3_1): BasicConv2d(
(conv): Conv2d(2048, 384, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(384, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch3x3_2a): BasicConv2d(
(conv): Conv2d(384, 384, kernel_size=(1, 3), stride=(1, 1), padding=(0, 1), bias=False)
(bn): BatchNorm2d(384, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch3x3_2b): BasicConv2d(
(conv): Conv2d(384, 384, kernel_size=(3, 1), stride=(1, 1), padding=(1, 0), bias=False)
(bn): BatchNorm2d(384, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch3x3dbl_1): BasicConv2d(
(conv): Conv2d(2048, 448, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(448, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch3x3dbl_2): BasicConv2d(
(conv): Conv2d(448, 384, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn): BatchNorm2d(384, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch3x3dbl_3a): BasicConv2d(
(conv): Conv2d(384, 384, kernel_size=(1, 3), stride=(1, 1), padding=(0, 1), bias=False)
(bn): BatchNorm2d(384, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch3x3dbl_3b): BasicConv2d(
(conv): Conv2d(384, 384, kernel_size=(3, 1), stride=(1, 1), padding=(1, 0), bias=False)
(bn): BatchNorm2d(384, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
(branch_pool): BasicConv2d(
(conv): Conv2d(2048, 192, kernel_size=(1, 1), stride=(1, 1), bias=False)
(bn): BatchNorm2d(192, eps=0.001, momentum=0.1, affine=True, track_running_stats=True)
)
)
(emb_features): Conv2d(768, 256, kernel_size=(1, 1), stride=(1, 1), bias=False)
(emb_cnn_code): Linear(in_features=2048, out_features=256, bias=True)
)
```
# Training
```
cfg.TEXT.EMBEDDING_DIM = 256
dataset.n_words = 2932 # for FashionGen subset. Computed by dataset.load_text_data(): parsing all captions
def build_models():
text_encoder = RNN_ENCODER(dataset.n_words, nhidden=cfg.TEXT.EMBEDDING_DIM)
image_encoder = CNN_ENCODER(cfg.TEXT.EMBEDDING_DIM)
```
---
```
batch_size = 32
for step, data in enumerate(dataloader, 0):
rnn_model.zero_grad()
cnn_model.zero_grad()
imgs, captions, cap_lens, class_ids, keys = prepare_data(data)
#
# imgs -- list of 1 image tensor
# imgs[0].shape --> torch.Size([32, 3, 299, 299])
#
# cap_lens.shape(32) --> tensor([10, 10, 10, 10, 10, 10, 10, 9, 9, 8, 8, 8, 7, 7, 7, 7,
# 7, 7, 7, 7, 7, 6, 6, 6, 6, 6, 6, 6, 5, 5, 5, 5]
#
#
# captions.shape(32, 10) --> tensor([[1151, 16, 526, 241, 1240, 1944, 1443, 303, 526, 1147],
# [2331, 195, 1624, 1151, 1078, 2859, 1837, 16, 526, 1147],
# ...
# [2153, 1837, 2538, 526, 1147, 0, 0, 0, 0, 0]]
#
# nef -- cfg.TEXT.EMBEDDING_DIM = 256 (for FashionGen)
# words_features: batch_size x nef x 17 x 17
# sentence_feature: batch_size x nef
words_features, sentence_feature = cnn_model(imgs[-1])
# words_features.shape --> torch.Size([32, 256, 17, 17])
# sentence_feature.shape --> torch.Size([32, 256])
# --> batch_size x nef x 17*17
nef, att_sze = words_features.size(1), words_features.size(2)
# nef -> 256
# att_sze -> 17
# words_features = words_features.view(batch_size, nef, -1)
hidden = rnn_model.init_hidden(batch_size)
# hidden -> list of size 2
# hidden[0].shape -> torch.Size([2, 32, 128]) -- all zeros
# hidden[1].shape -> torch.Size([2, 32, 128]) -- all zeros
# words_emb: batch_size x nef x seq_len
# sent_emb: batch_size x nef
words_emb, sent_emb = rnn_model(captions, cap_lens, hidden)
w_loss0, w_loss1, attn_maps = words_loss(words_features, words_emb, labels, cap_lens, class_ids, batch_size)
w_total_loss0 += w_loss0.data
w_total_loss1 += w_loss1.data
loss = w_loss0 + w_loss1
s_loss0, s_loss1 = sent_loss(sentence_feature, sent_emb, labels, class_ids, batch_size)
loss += s_loss0 + s_loss1
s_total_loss0 += s_loss0.data
s_total_loss1 += s_loss1.data
loss.backward()
# `clip_grad_norm` helps prevent the exploding gradient problem in RNNs / LSTMs.
torch.nn.utils.clip_grad_norm_(rnn_model.parameters(), cfg.TRAIN.RNN_GRAD_CLIP)
optimizer.step()
```
| github_jupyter |
# Adversarial Attacks with parametrized DPR on VGGFace2
```
from torch.autograd import Variable
%load_ext autoreload
%autoreload 2
import os.path
import sys
sys.path.append(os.path.join(os.path.dirname(os.path.realpath('__file__')), '..'))
from relighters.DPR.model.defineHourglass_512_gray_skip import HourglassNet
from torch import nn
import torch
import torchvision
import os
from relighters.DPR.face_utils import plot_face_attack, get_sh_with_relighter
from relighters.DPR.spherical_harmonics import get_random_spherical_harmonics
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
from facenet_pytorch import MTCNN
from torchvision import transforms
from torchvision.datasets import ImageFolder
from classifiers.VGGFace2.VGGFace2Classifier import VGGFace2Classifier
from relighters.DPR.preparation import np_rgb_to_torch_lab
from utils.kornia_lab import RgbToLab, LabToRgb
from utils import labels_util
# Classifier
model = VGGFace2Classifier()
model.eval()
def collate_fn(x):
return x[0]
# Image standardization used when the vggface2 classifer was trained
def fixed_image_standardization(image_tensor):
processed_tensor = (image_tensor - 127.5) / 128.0
return processed_tensor
path = "../data/vggface2-80"
preprocess = transforms.Compose([
transforms.Resize((224, 224)),
transforms.ToTensor()
])
dataset = ImageFolder(path)
dataset.idx_to_class = {i: c for c, i in dataset.class_to_idx.items()}
batch_size = 1
loader = torch.utils.data.DataLoader(dataset, batch_size=batch_size, shuffle=False, collate_fn=collate_fn)
# Load helper files that deal with the different labels
complete_to_subset = labels_util.load_idx_to_label('vggface2')
subset_to_complete = {value: key for key, value in complete_to_subset.items()}
# Cropper
mtcnn = MTCNN(image_size=224, margin=0, min_face_size=20, thresholds=[0.6, 0.7, 0.7],
factor=0.709, post_process=False)
# Initialize classes for doing LAB transformations
r2l = RgbToLab()
l2r = LabToRgb()
%%capture
modelFolder = '../relighters/DPR/trained_model/'
my_network = HourglassNet()
my_network.load_state_dict(torch.load(os.path.join(modelFolder, 'trained_model_03.t7')))
my_network.train(False)
relighting = my_network
learning_rate = 0.015
max_steps = 50
# Set targeted or untargeted
targeted = True
target_label = 1
sign = -1 if targeted else 1
target_label = torch.tensor(target_label).unsqueeze(0)
successful_iterations = []
#show_image_every = 50
total, ad, f = 0, 0, 0
reg = 0.5
# loop over test data
for id, (image, gt_label) in enumerate(loader):
gt_label = subset_to_complete[gt_label]
loss_history = []
# Get cropped image
cropped = mtcnn(image)
if cropped is None:
print("No Face detected")
continue
# Standardize image for classifier
standardized = fixed_image_standardization(cropped)
# Predict on clean image
logits = model(standardized.unsqueeze(0))
#print(logits.size())
probs = torch.softmax(logits, dim=1)
# Get predicted original label
orig_prob, orig_predicted_label = torch.max(probs, dim=1)
if orig_predicted_label != gt_label:
print("Unperturbed image was misclassified")
continue
# relighter expects image in range 0;1
cropped = cropped / 255
# l-space transformations
img_lab = np_rgb_to_torch_lab(cropped.numpy().squeeze())
input_l = img_lab[0,:, :]
# DPR expects values between 0 and 1, whereas our lab transformation returns the common L values between 0 and 100
input_l = (input_l/100.0)
input_l = input_l[None,None, ...]
input_l = input_l.float()
input_ab = img_lab[1:,:,: ]
# initialize shade params
estimated_sh = get_sh_with_relighter(input_l, relighting).detach()
# Our sh parameter we optimize over
sh = get_random_spherical_harmonics()
sh = Variable(torch.from_numpy(sh), requires_grad=True).float()
sh.retain_grad()
gt_label = torch.tensor(gt_label).unsqueeze(0)
model.eval()
mtcnn.eval()
i = 0
# optimization loop to find optimal shade parameters
with torch.enable_grad():
for i in range(max_steps):
model.zero_grad()
mtcnn.zero_grad()
if sh.grad is not None:
sh.grad.zero_()
sh = Variable(sh, requires_grad=True)
sh.retain_grad()
# relight the current image
out_l, out_sh = relighting(input_l, sh, 0)
out_l_perm = out_l[0]
out_l_scaled = (out_l_perm*100.0)
output_lab = torch.cat([out_l_scaled.double(), input_ab.double()], dim=0)
# Transform back to RGB space for Classifier
output_rgb = l2r.lab_to_rgb(output_lab)
# After the relighting on the cropped image we do the standardization that the classifier expects
# The *255 because classifer expects different range
standardized = fixed_image_standardization(output_rgb * 255)
logits = model(standardized.unsqueeze(0).float())
probs = torch.softmax(logits, dim=1)
# Get predicted original label
probability, prediction = torch.max(probs, dim=1)
nll = nn.functional.nll_loss(logits, target_label)
loss = nll - sign * nll * reg * torch.dist(estimated_sh, sh, p=2)
loss_history.append(loss)
# Breaking conditions
if (targeted and prediction == target_label) or (
not targeted and prediction != gt_label):
# Generated image is adversarial
break
loss.backward()
grad = sh.grad
if grad is None:
print("Error")
# Perform gradient ascent or descent update step based on whether we do targeted attack or not
sh = sh + sign * learning_rate * grad
# plot results
total += 1
if torch.mean(output_rgb) < 0.1 or torch.mean(output_rgb) > 0.9:
f += 1
print("This is a failed relighting:")
plot_face_attack(torch.clamp(cropped, 0, 1), torch.clamp(output_rgb, 0, 1), loss_history, sh, out_sh, gt_label,
orig_prob, prediction, probability)
else:
if not targeted:
if gt_label != prediction:
ad += 1
successful_iterations.append((True, i))
plot_face_attack(torch.clamp(transforms.ToTensor()(image), 0, 1), torch.clamp(output_rgb, 0, 1), loss_history, sh, out_sh, gt_label,
orig_prob, prediction, probability)
else:
if target_label == prediction:
ad += 1
successful_iterations.append((True, i))
plot_face_attack(torch.clamp(transforms.ToTensor()(image), 0, 1), torch.clamp(output_rgb, 0, 1), loss_history, sh, out_sh, gt_label,
orig_prob, prediction, probability)
print("total images processed: ", total)
print("adversarial examples: ", ad)
print("failed relightings: ", f)
print("iterations: ", i)
print(f"Overall success rate: {ad / float(total)}")
print(f"Average number of iterations: {torch.mean(torch.tensor(successful_iterations))}")
```
| github_jupyter |
```
import numpy as np
import pandas as pd
from sklearn.preprocessing import LabelEncoder
from rules import normalized_chars
import random
import re
from unidecode import unidecode
laughing = {
'huhu',
'haha',
'gagaga',
'hihi',
'wkawka',
'wkwk',
'kiki',
'keke',
'huehue',
'hshs',
'hoho',
'hewhew',
'uwu',
'sksk',
'ksks',
'gituu',
'gitu',
'mmeeooww',
'meow',
'alhamdulillah',
'muah',
'mmuahh',
'hehe',
'salamramadhan',
'happywomensday',
'jahagaha',
'ahakss',
'ahksk'
}
def make_cleaning(s, c_dict):
s = s.translate(c_dict)
return s
def cleaning(string):
"""
use by any transformer model before tokenization
"""
string = unidecode(string)
string = ' '.join(
[make_cleaning(w, normalized_chars) for w in string.split()]
)
string = re.sub('\(dot\)', '.', string)
string = (
re.sub(re.findall(r'\<a(.*?)\>', string)[0], '', string)
if (len(re.findall(r'\<a (.*?)\>', string)) > 0)
and ('href' in re.findall(r'\<a (.*?)\>', string)[0])
else string
)
string = re.sub(
r'\w+:\/{2}[\d\w-]+(\.[\d\w-]+)*(?:(?:\/[^\s/]*))*', ' ', string
)
chars = '.,/'
for c in chars:
string = string.replace(c, f' {c} ')
string = re.sub(r'[ ]+', ' ', string).strip().split()
string = [w for w in string if w[0] != '@']
x = []
for word in string:
word = word.lower()
if any([laugh in word for laugh in laughing]):
if random.random() >= 0.5:
x.append(word)
else:
x.append(word)
string = [w.title() if w[0].isupper() else w for w in x]
return ' '.join(string)
labels = """
1. severe toxic
2. obscene
3. identity attack
4. insult
5. threat
6. asian
7. atheist
8. bisexual
9. black
10. buddhist
11. christian
12. female
13. heterosexual
14. indian
15. homosexual, gay or lesbian
16. intellectual or learning disability
17. jewish
18. latino
19. male
20. muslim
21. other disability
22. other gender
23. other race or ethnicity
24. other religion
25. other sexual orientation
26. physical disability
27. psychiatric or mental illness
28. transgender
29. white
30. malay
31. chinese
"""
labels = [l.split('. ')[1].strip() for l in labels.split('\n') if len(l)]
labels
import glob
files = glob.glob('../toxicity/translated*')
files
import json
X, Y = [], []
for file in files:
print(file)
with open(file) as fopen:
f = json.load(fopen)
for row in f:
if len(row[1]) == 29:
X.append(row[0])
Y.append(row[1] + [0, 0])
len(X)
rejected_labels = ['black', 'white', 'jewish', 'latino']
[labels.index(l) for l in rejected_labels]
labels = [l for l in labels if l not in rejected_labels]
ydf = pd.DataFrame(np.array(Y))
ydf = ydf.loc[(ydf[8] == 0) & (ydf[28] == 0) & (ydf[16] == 0) & (ydf[17] == 0)]
ydf = ydf.drop([8, 28, 16, 17], axis = 1)
ix = ydf.index.tolist()
Y = ydf.values.tolist()
X = [X[i] for i in ix]
mapping = {'severe_toxic': 'severe toxic', 'identity_hate': 'identity attack',
'toxic': 'severe toxic', 'melayu': 'malay', 'cina': 'chinese', 'india': 'indian'}
def generate_onehot(tags, depth = len(labels)):
onehot = [0] * depth
for tag in tags:
onehot[labels.index(tag)] = 1
return onehot
with open('../toxicity/kaum.json') as fopen:
kaum = json.load(fopen)
for k, v in kaum.items():
print(k, len(v))
with open('../toxicity/weak-learning-toxicity.json') as fopen:
scores = json.load(fopen)
for k, v in scores.items():
for no in range(len(v)):
tags = []
for l, v_ in v[no].items():
if round(v_) == 1:
tags.append(mapping.get(l, l))
tags.append(mapping[k])
Y.append(generate_onehot(tags))
X.append(kaum[k][no])
from tqdm import tqdm
for i in tqdm(range(len(X))):
X[i] = cleaning(X[i])
actual_t, actual_l = [], []
for i in tqdm(range(len(X))):
if len(X[i]) > 2:
actual_t.append(X[i])
actual_l.append(Y[i])
with open('combined.txt', 'w') as fopen:
fopen.write('\n'.join(actual_t))
import youtokentome as yttm
%%time
bpe = yttm.BPE.train(data='combined.txt',
vocab_size=60000, model='toxic.model')
vocab = {v: i for i, v in enumerate(bpe.vocab())}
rev_vocab = {i: v for i, v in enumerate(bpe.vocab())}
len(vocab)
from sklearn.feature_extraction.text import TfidfVectorizer
import re
r = re.compile(r'[\S]+').findall
subs = [' '.join(s) for s in bpe.encode(actual_t, output_type=yttm.OutputType.SUBWORD)]
tfidf = TfidfVectorizer(vocabulary = vocab, token_pattern = r'[\S]+').fit(subs)
import pickle
with open('tfidf-toxic.pkl','wb') as fopen:
pickle.dump(tfidf,fopen)
vector = tfidf.transform(subs)
Y = np.around(np.array(actual_l))
from sklearn.model_selection import train_test_split
train_X, test_X, train_Y, test_Y = train_test_split(vector, Y, test_size = 0.2)
train_X.shape, test_X.shape
from sklearn.naive_bayes import ComplementNB
from sklearn.multiclass import OneVsRestClassifier
multinomial = OneVsRestClassifier(ComplementNB()).fit(train_X, train_Y)
from sklearn import metrics
print(
metrics.classification_report(
train_Y,
multinomial.predict(train_X),
target_names=labels,digits=5
)
)
print(
metrics.classification_report(
test_Y,
multinomial.predict(test_X),
target_names=labels,digits=5
)
)
with open('multinomial-toxic.pkl','wb') as fopen:
pickle.dump(multinomial,fopen)
import boto3
s3 = boto3.client('s3')
bucketName = 'huseinhouse-storage'
Key = 'multinomial-toxic.pkl'
outPutname = "v34/toxicity/multinomial.pkl"
s3.upload_file(Key,bucketName,outPutname)
Key = 'tfidf-toxic.pkl'
outPutname = "v34/toxicity/tfidf.pkl"
s3.upload_file(Key,bucketName,outPutname)
Key = 'toxic.model'
outPutname = "v34/toxicity/youtokentome.model"
s3.upload_file(Key,bucketName,outPutname)
```
| github_jupyter |
```
%matplotlib notebook
import matplotlib.pyplot as plt
import numpy as np
import scipy.stats
```
# Tiempo de mezcla (mixing time)
Previamente hemos visto como diseñar una cadena de Markov finita tal que converja a una distribución estacionaria de nuestro interés
Pero ¿Cuánto debemos esperar para que ocurra la convergencia?
El tiempo de mezcla para una cadena de Markov irreducible y aperiodica se define como
$$
t_{mix}(\epsilon) = \min \left(n > 0: \|s(n) - \pi\|_{TV} < \epsilon \right)
$$
es decir el mínimo tiempo (número de pasos) tal que estemos a una distancia $\epsilon$ de la distribución estacionaria $\pi$
El operador $\|p - q\|_{TV} = \max_{x\in\mathcal{\Omega}} \|p(x) - q(x)\|$ se conoce como la distancia de variación total entre dos distribuciones
Se tienen algunas garantías con respecto a este tiempo, en particular la siguiente cota superior
$$
t_{mix}(\epsilon) < \log \left(\frac{1}{\epsilon \sqrt{\min_j \pi_j}} \right) \frac{1}{1-\lambda_*}
$$
donde $\lambda_*$ es el segundo valor propio más grande de la matriz de transición $P$ de la cadena.
La descomposición en valores propios de la matriz de transición de una cadena irreducible y de $\mathcal{S}$ estados se puede escribir como
$$
P^n = \sum_{i=1}^\mathcal{S} \alpha_i \lambda_i^n = \pi + \sum_{i=2}^\mathcal{S} \alpha_i \lambda_i^n
$$
Por propiedad de la cadena de Markov irreducible su valor propio más grande siempre es igual a uno y su vector propio asociado es la distribución estacionaria.
Todos los demás valores propios se harán eventualmente cero cuando $n \to \infty$, siendo el segundo valor propio más grande y distinto de uno el que más se demore
# Autocorrelación en la cadena y número de muestras efectivo
¿Cómo podemos confirmar si nuestro algoritmo MCMC ha convergido?
Por construcción, las muestras de nuestra traza son dependientes, pues $\theta_{t+1}$ se calcula a partir de $\theta_t$.
Sin embargo, luego de un periódo de *burn-in*, las probabilidades de transición de la cadena debería converger a la distribución estacionaria y volverse independientes del tiempo
Es decir que podemos confirmar la convergencia estudiando la autocorrelación de la traza
$$
\rho(\tau) = \mathbb{E}\left[(\theta_t - \bar \theta)(\theta_{t+\tau} - \bar \theta)\right]
$$
La autocorrelación nos indica que tanto las muestras de una serie de tiempo dependen de muestras pasadas.
En este caso, al graficar $\rho$ en función de $\tau$ buscamos una autocorrelación que converja rapidamente y que luego fluctue en torno a cero
<img src="images/autocorr.png" width="700">
## Ejemplo
La lección pasada vimos como el valor de $\sigma_\epsilon$ repercutía fuertemente en la convergencia del algoritmo de Metropolis
Usemos la función `np.correlate` para estudiar la autocorrelación de las trazas para distintos valores de $\sigma_\epsilon$
```
x = np.array([9.37, 10.18, 9.16, 11.60, 10.33])
prior = lambda theta : scipy.stats.norm(loc=5, scale=np.sqrt(10)).pdf(theta)
likelihood = lambda theta : np.prod([scipy.stats.norm(loc=theta, scale=1.).pdf(x_) for x_ in x])
r = lambda ts, tt : likelihood(ts)*prior(ts)/(likelihood(tt)*prior(tt))
def metropolis(mix_time=5000, sigma_eps=1.):
thetas = np.zeros(shape=(mix_time, ))
thetas[0] = np.random.randn()
ar = 0.
qs = scipy.stats.norm(loc=0, scale=sigma_eps).rvs(size=mix_time)
us = scipy.stats.uniform.rvs(size=mix_time)
for n in range(1, mix_time):
theta_star = thetas[n-1] + qs[n]
if us[n] < np.amin([1, r(theta_star, thetas[n-1])]):
thetas[n] = theta_star
ar += 1.
else:
thetas[n] = thetas[n-1]
return thetas, ar/mix_time
def autocorr(thetas):
thetas_norm = (thetas-np.mean(thetas))/np.std(thetas)
rho = np.correlate(thetas_norm,
thetas_norm, mode='full')
return rho[len(rho) // 2:]/len(thetas)
def neff(rho):
T = np.where(rho < 0.)[0][0]
return len(rho)/(1 + 2*np.sum(rho[:T]))
%%time
np.random.seed(12345)
fig, ax = plt.subplots(1, 2, figsize=(7, 3), tight_layout=True)
for sigma in [0.1, 1., 2., 10.]:
thetas, ar = metropolis(mix_time=2000, sigma_eps=sigma)
ax[0].plot(thetas)
ax[1].plot(autocorr(thetas), label=str(sigma))
print(f"Número efectivo de muestras para sigma {sigma}: {neff(autocorr(thetas)):0.4f}")
print(f"Fracción de aceptación: {ar:0.4f}")
ax[0].set_title('Traza')
ax[1].set_title('Autocorrelación');
ax[1].legend();
```
De la figura podemos ver que
- Si el paso de las propuestas es muy corto, todos los pasos son aceptados, pero la correlación entre los mismos es alta por ser poco diversos
- Si el paso de las propuestas es muy largo, se proponen demasiadas propuestas malas que terminan siendo rechazadas. La fracción de aceptación disminuye y la autocorrelación aumenta
La fracción de aceptación es la cantidad de propuestas aceptadas dividido las propuestas totales. La sugerencia de los expertos es calibrar el algoritmo de Metropolis tal que alcance [una fracción de aceptación cercana a 23.4%](https://www.maths.lancs.ac.uk/~sherlocc/Publications/rwm.final.pdf)
Una figura de mérito muy utilizada que se basa en la función de autocorrelación es el número de muestras efectivas
$$
n_{eff} = \frac{N}{1 + 2 \sum_{\tau=1}^T \rho(\tau)}
$$
donde $N$ es la cantidad de muestras de la cadena y $T$ es el instante en que la autocorrelación se vuelve por primera vez negativa
Idealmente quisieramos que $n_{eff} = N$, pero debido a que las muestras no son independientes en realidad tendremos $n_{eff} < N$
Podemos calibrar nuestro algoritmo MCMC tal que maximicemos $n_{eff}$
## Thinning (adelgazamiento)
Es una técnica para disminuir la autocorrelación de la traza.
Consiste en submuestrear la traza, retendiendo sólo cada $t$ muestras, donde $t$ es el "intervalo de adelgazamiento". La idea es escoger este intervalo estudiando la función de autocorrelación
Debido a la gran cantidad de muestras que se podrían descartar es preferible ajustar adecuadamente el paso de las propuestas por sobre esta técnica
## Estadístico Gelman-Rubin
Supongamos que no entrenamos una sino $M$ cadenas
Otra forma de estudiar la convergencia en este caso es el estadístico Gelman Rubin o $\hat r$
Este estadístico compara la varianza (dispersión) dentro de la cadena con la varianza entre las $M$ cadenas.
Si el valor de $\hat r$ es cercano a uno es indicativo de que la cadena ha convergido
# Material extra
- [Valores propios](https://www.cl.cam.ac.uk/teaching/1819/Probablty/materials/Lecture9_handout.pdf) y [cotas en los tiempos de mezcla](https://www.cl.cam.ac.uk/teaching/1920/Probablty/materials/Lecture10.pdf)
- [Parallel tempering](https://www.pas.rochester.edu/~sybenzvi/courses/phy403/2016s/p403_18_mcmc.pdf)
- [Artículo "Una introducción conceptual a MCMC"](https://arxiv.org/pdf/1909.12313.pdf)
| github_jupyter |
```
import pandas as pd
import os
df = pd.read_table('linhas_dr.txt', delim_whitespace = True)
df.head(5)
columns = ["Hbeta", "OIII.4959", "OIII.5007", "NII.6548", "Halpha", "NII.6584", "SII.6716", "SII.6731", "mag_r"]
for column in columns:
if column == "Hbeta":
df_final = df["Hbeta"]
else:
df_final = pd.concat([df_final,df[column]],axis=1)
df_final.shape
df_final.head(5)
for column in columns:
df_final = df_final[df_final[column] != -999]
df_final.shape
df_final.head(5)
df = df_final
data = []
indexes = []
for index, row in df.iterrows():
temp = row.tolist()
if 16<temp[8]<17:
data.append(temp)
indexes.append(index)
print(len(data))
print(len(indexes))
import numpy as np
data = np.asarray(data)
data[0]
#data = np.delete(data, np.s_[:2],1)
data = np.delete(data, np.s_[-1],1)
data[0]
print(data.shape)
data_normalized = np.zeros(data.shape)
for feature in range(data.shape[1]):
Min = min(data[:,feature])
Max = max(data[:,feature])
for training_exemple in range(data.shape[0]):
data_normalized[training_exemple][feature] = (data[training_exemple][feature] - Min) / (Max - Min)
print(data_normalized[:3])
print('Valor Mínimo:{} \nValor Máximo:{}'.format(np.min(data_normalized[:,0]),np.max(data_normalized[:,0])))
contador_minimo = 0
contador_maximo = 0
for x in range(data_normalized.shape[1]):
valor_minimo = np.min(data_normalized[:,0])
valor_maximo = np.max(data_normalized[:,0])
if valor_minimo == 0.0:
contador_minimo += 1
if valor_maximo == 1.0:
contador_maximo += 1
if contador_minimo and contador_maximo == 8:
print('[INFO] Dados Normalizados')
df = pd.DataFrame(data_normalized)
df.head(5)
df.to_csv("Training_Data", index = False)
df.shape
dff = pd.read_csv("Training_Data")
dff.head(5)
dff.shape
import pandas as pd
import numpy as np
data_normalized = pd.read_csv("Training_Data")
data_normalized = np.asarray(data_normalized)
classes = pd.read_csv("Classes.csv")
features = []
for i in range(data_normalized.shape[1]):
features.append('Feature {}'.format(i+1))
features.append("type1")
features.append("whan")
type1 = np.copy(classes.values[:,1])
type1 = np.reshape(type1, (type1.shape[0],1))
concat = np.concatenate([data_normalized, type1], axis = 1)
whan = np.copy(classes.values[:,-1])
whan = np.reshape(whan, (whan.shape[0],1))
concat = np.concatenate([concat, whan], axis = 1)
data_df = pd.DataFrame(data = concat , columns = features)
data_df.to_csv("Dataset.csv", index = False)
data_df.head(5)
classes.shape
seed = 7
np.random.seed(seed)
#train_df = data_df
train_df = data_df.sample(frac=1)
train_df.to_csv("Training_Data", index = False)
import pandas as pd
train_df = pd.read_csv("Training_Data")
train_df.head(5)
import pandas as pd
import numpy as np
import astropy
df = pd.read_csv("Training_Data")
df.head(5)
mat = df.values[:,:-2]
zeros = np.zeros((mat.shape[0],6),dtype="float32")
mat = np.concatenate([mat,zeros], axis=1)
mat = np.concatenate([mat,df.values[:,-2:]], axis=1)
features = []
count = 0
for i in df.columns:
if count == 10:
break
else:
features.append(i)
count += 1
for i in range(zeros.shape[1]):
features.append("pad")
features.append('BPT_class_n')
features.append('WHAN_class_n')
df = pd.DataFrame(data = mat, columns = features)
df.head(5)
df.to_csv("Training_Data", index=False)
line_list = [3727, 3869, 4101, 4340, 4363, 4471, 4861, 4959, 5007, 6300, 6548, \
6563, 6584, 6717, 6731]
features = np.array([sample[str(line)+'_ew'].data.data for line in line_list])
from astropy.table import Table
df = Table.read('DR7_lines_ML1.fits')
df = df[df["scienceprimary"] == 1]
df = df[df["WHAN_flag"] == True]
df = df[df["BPT_flag"] == True]
df
import numpy as np
line_list = ['specobjid','3727_ew', '4340_ew', '4861_ew', '5007_ew', '6300_ew', '6548_ew', \
'6563_ew', '6584_ew', '6717_ew', '6731_ew']
features = np.array([df[line].data.data for line in line_list])
features.shape
features = features.T
zeros = np.zeros((features.shape[0], 6))
features = np.concatenate([features,zeros], axis = 1)
labels = ["BPT_flag", "WHAN_flag"]
classe = np.array([df[label].data.data for label in labels])
features = np.concatenate([features,classe.T], axis=1)
import pandas as pd
for i in range(zeros.shape[1]):
line_list.extend(["pad"])
line_list.extend(labels)
df = pd.DataFrame(data = features, columns = line_list)
df.head(5)
line_list = ['3727_ew', '4340_ew', '4861_ew', '5007_ew', '6300_ew', '6548_ew', \
'6563_ew', '6584_ew', '6717_ew', '6731_ew']
for line in line_list:
df = df[df[line] != -999]
df.shape
df.to_csv("Dataset", index=False)
import pandas as pd
import numpy as np
d = pd.read_csv("Dataset")
e = pd.read_csv("Training_Data")
print(d.shape, e.shape)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn as sk
from sklearn.naive_bayes import MultinomialNB
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import f1_score
from sklearn.preprocessing import MinMaxScaler
# Набор данных взят с https://www.kaggle.com/aungpyaeap/fish-market
# Параметры нескольких популярных промысловых рыб
# length 1 = Body height
# length 2 = Total Length
# length 3 = Diagonal Length
fish_data = pd.read_csv("datasets/Fish.csv", delimiter=',')
print(fish_data)
# Выделим параметры и метки классов
x_labels = ['Weight', 'Length1', 'Length2', 'Length3', 'Height', 'Width']
y_label = 'Species'
data = fish_data[x_labels + [y_label]]
print(data)
# Определим размер валидационной и тестовой выборок
val_test_size = round(0.2*len(data))
print(val_test_size)
# Генерируем уникальный seed
my_code = "Махраби"
seed_limit = 2 ** 32
my_seed = int.from_bytes(my_code.encode(), "little") % seed_limit
# Создадим обучающую, валидационную и тестовую выборки
random_state = my_seed
train_val, test = train_test_split(data, test_size=val_test_size, random_state=random_state)
train, val = train_test_split(train_val, test_size=val_test_size, random_state=random_state)
print(len(train), len(val), len(test))
# Выделим обучающую, валидационную и тестовую выборки
train_x = train[x_labels]
train_y = np.array(train[y_label])
val_x = val[x_labels]
val_y = np.array(val[y_label])
test_x = test[x_labels]
test_y = np.array(test[y_label])
test_y1 = np.array(test[y_label]).reshape(-1,1)
# Нормируем значения параметров
scaler_x = MinMaxScaler()
scaler_x.fit(train_x)
scaled_train_x = scaler_x.transform(train_x)
scaled_val_x = scaler_x.transform(val_x)
scaled_test_x = scaler_x.transform(test_x)
# Нарисуем график распределения классов
plt.hist(train_y)
plt.show()
plt.hist(val_y)
plt.show()
plt.hist(test_y)
plt.show()
# Создадим модель наивного Байесовского классификатора и обучим ее на ненормированных данных.
model1 = MultinomialNB()
model1.fit(train_x, train_y)
# Создадим модель наивного Байесовского классификатора и обучим ее на нормированных данных.
model2 = MultinomialNB()
model2.fit(scaled_train_x, train_y)
# Проверим результат на валидационной выборке. модель1 - на ненормированных данных. модель2 на нормированных
val_predicted1 = model1.predict(val_x)
f1_1 = f1_score(val_y, val_predicted1, average = 'weighted')
print(f1_1)
val_predicted2 = model2.predict(scaled_val_x)
f1_2 = f1_score(val_y, val_predicted2, average = 'weighted')
print(f1_2)
# Создадим модель логистической регрессии и обучим ее на нормированных данных.
model1 = LogisticRegression()
model1.fit(scaled_train_x, train_y)
# Проверим результат на валидационной выборке
val_predicted = model1.predict(scaled_val_x)
f1_1 = f1_score(val_y, val_predicted, average = 'weighted')
print(f1_1)
# модель логистической регрессии на валидационной выборке имеет лучший результат
test_predicted = model1.predict(scaled_test_x)
f1_1 = f1_score(test_y, test_predicted, average = 'weighted')
print(f1_1)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/michelucci/zhaw-dlcourse-spring2019/blob/master/Week%203%20-%20Computational%20graphs/Week%203%20-%20Exercises%20Solutions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Neural Networks and Deep Learning for Life Sciences and Health Applications - An introductory course about theoretical fundamentals, case studies and implementations in python and tensorflow
(C) Umberto Michelucci 2018 - umberto.michelucci@gmail.com
github repository: https://github.com/michelucci/zhaw-dlcourse-spring2019
Spring Semester 2019
```
import tensorflow as tf
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
```
# Solutions to exercises
## Exercise 1 (Difficulty: easy)
Draw and develop in tensorflow with ```tf.constant``` the computational graphs for the following operations
A) ```w1*x1+w2*x2+x1*x1```
B) ```A*x1+3+x2/2```
Use as input values ```x1 = 5``` and ```x2 = 6```
## A)
There are several ways of solving this exercise. This is one possible
```
# Building Phase
x1 = tf.constant(5.)
x2 = tf.constant(6.)
w1 = 10.
w2 = 20.
z1 = tf.multiply(w1, x1)
z2 = tf.multiply(w2, x2)
z3 = tf.multiply(x1, x1)
result = z1 + z2 + z3
# Evaluation Phase
with tf.Session() as sess:
print(result.eval())
```
A second way of doing that is the following
```
# Building Phase
x1 = tf.constant(5.)
x2 = tf.constant(6.)
w1 = 10.
w2 = 20.
z1 = tf.multiply(w1, x1)
z2 = tf.multiply(w2, x2)
z3 = tf.multiply(x1, x1)
result = z1 + z2 + z3
# Evaluation Phase
sess = tf.Session()
print(sess.run(result))
sess.close()
```
But you can also define ```w1``` and ```w2``` as constants too
```
# Building Phase
x1 = tf.constant(5.)
x2 = tf.constant(6.)
w1 = tf.constant(10.)
w2 = tf.constant(20.)
z1 = tf.multiply(w1, x1)
z2 = tf.multiply(w2, x2)
z3 = tf.multiply(x1, x1)
result = z1 + z2 + z3
# Evaluation Phase
sess = tf.Session()
print(sess.run(result))
sess.close()
```
### B)
```
# Building Phase
x1 = tf.constant(5.)
x2 = tf.constant(6.)
A = tf.constant(10.)
result = tf.multiply(A, x1) + tf.constant(3.) + tf.divide(x2, 2.)
# Evaluation Phase
sess = tf.Session()
print(sess.run(result))
sess.close()
```
or you can define the ```result``` in multiple steps
```
# Building Phase
z1 = tf.multiply(A, x1)
z2 = tf.add(z1, 3.)
z3 = tf.add(z2, tf.divide(x2,2.))
# Evaluation Phase
sess = tf.Session()
print(sess.run(result))
sess.close()
```
## Exercise 2 (Difficulty: medium)
Draw and develop in tensorflow with ```tf.Variable``` the computational graph for the following operation ```A*(w1*x1+w2*x2)```
build the computational graph and then evaluate it two times (without re-building it) with the initial values in the same session
A) ```x1 = 3, x2 = 4```
B) ```x1 = 5, x2 = 7```
```
# Building Phase
x1 = tf.Variable(3.)
x2 = tf.Variable(4.)
w1 = tf.constant(10.)
w2 = tf.constant(20.)
A = tf.constant(30.)
init = tf.global_variables_initializer()
z1 = tf.multiply(w1,x1)
z2 = tf.multiply(w2,x2)
z3 = tf.add(z1, z2)
result = tf.multiply(A, z3)
```
To run the same graph twice in the same session you can do the following
```
sess = tf.Session()
print(sess.run(result, feed_dict = {x1: 3, x2: 4}))
print(sess.run(result, feed_dict = {x1: 5, x2: 7}))
sess.close()
```
Or you can write a function that creates a session, evaluates a node, and then close it.
```
def run_evaluation(x1_, x2_):
sess = tf.Session()
print(sess.run(result, feed_dict = {x1: x1_, x2: x2_}))
sess.close()
```
And then you can evalute the node with a call to your function.
```
run_evaluation(3,4)
run_evaluation(5,7)
```
## Exercise 3 (Difficulty: FUN)
Consider two vectors
``` x1 = [1,2,3,4,5], x2 = [6,7,8,9,10]```
draw and build in tensorflow the computational graph for the dot-product operation between the two vectors. If you don't know what a dot-product is you can check it here (we covered that in our introductory week) [](https://en.wikipedia.org/wiki/Dot_product).
Build it in two different ways:
A) Do it with loops. Build a computational graph that takes as input scalars and in the session/evaluation phase build a loop to go over all the inputs and then sums the results
B) Do it in one shot with tensorflow. Build a computational graph that takes as input vectors and do the entire operation directly in tensorflow.
Hint: you can use in tensorflow two methods: ```tf.reduce_sum(tf.multiply(x1, x2))``` or ```tf.matmul(tf.reshape(x1,[1,5]), tf.reshape(x2, [-1, 1]))```. Try to understand why they work checking the official documentation.
## a)
```
first = tf.Variable(0.)
second = tf.Variable(0.)
mult = tf.multiply(first, second)
x1 = [1,2,3,4,5]
x2 = [6,7,8,9,10]
sess = tf.Session()
total = 0
for i in range(0,len(x1)):
total = total + sess.run(mult, feed_dict = {first: x1[i], second: x2[i]})
print(total)
```
Note that you can do that easily in numpy
```
np.dot(x1, x2)
```
## b)
Another way, and much more efficient, is the following
```
x1 = tf.placeholder(tf.int32, None) # Let's assume we work with integers
x2 = tf.placeholder(tf.int32, None) # Let's assume we work with integers
result = tf.reduce_sum(tf.multiply(x1, x2))
sess = tf.Session()
print(sess.run(result, feed_dict = {x1: [1,2,3,4,5], x2:[6,7,8,9,10]}))
sess.close()
```
Or in with matrices
```
x1 = tf.placeholder(tf.int32, None) # Let's assume we work with integers
x2 = tf.placeholder(tf.int32, None) # Let's assume we work with integers
result = tf.matmul(tf.reshape(x1,[1,5]), tf.reshape(x2, [-1, 1]))
sess = tf.Session()
print(sess.run(result, feed_dict = {x1: [1,2,3,4,5], x2:[6,7,8,9,10]}))
sess.close()
```
Note that the result is different in the two cases! In the first we get a scalar, in the second a matrix that has dimensions ```1x1```, because the second method is a matrix multiplication function that will return a matrix (or better a tensor).
## c) (even another way) (BONUS Solution)
There is actually another way. Tensorflow can perform the dot product directly
```
x1 = tf.placeholder(tf.int32, None) # Let's assume we work with integers
x2 = tf.placeholder(tf.int32, None) # Let's assume we work with integers
result = tf.tensordot(x1, x2, axes = 1)
sess = tf.Session()
print(sess.run(result, feed_dict = {x1: [1,2,3,4,5], x2:[6,7,8,9,10]}))
sess.close()
```
## Exercise 4 (Difficulty: medium)
Write a function that build a computational graph for the operation ```x1+x2``` where the input ```x1``` and ```x2``` are input with given dimensions. Your ```x1``` and ```x2``` should be declared as ```tf.placeholder```.
Your functions should accept as input:
- dimensions of ```x1``` as list, for example ```[3]```
- dimensions of ```x2``` as list, for example ```[3]```
The function should return a tensor ```z = x1 + x2```.
Then open a session and evaluate ```z``` with the following inputs:
- ```x1 = [4,6,7], x2 = [1,2,9]```
- ```x1 = [1,2,....., 1000], x2 = [10001, 10002, ...., 11000]```
and print the result.
```
def build_graph(dim1, dim2):
tf.reset_default_graph()
x1 = tf.placeholder(tf.float32, dim1)
x2 = tf.placeholder(tf.float32, dim2)
z = tf.add(x1, x2)
return z, x1, x2
x1list = [4,6,7]
x2list = [1,2,9]
# Building Phase
z, x1, x2 = build_graph(len(x1list), len(x2list))
sess = tf.Session()
print(sess.run(z, feed_dict = {x1: x1list, x2: x2list}))
sess.close()
```
**Note that since you refer to the tensors ```x1``` and ```x2``` in the ```feed_dict``` dictionary you need to have the tensors visible, otherwise you will get an error, therefore you need your function to return no only ```z``` but also ```x1``` and ```x2```.**
```
x1list = np.arange(1, 1001, 1)
x2list = np.arange(10001, 11001, 1)
# Building Phase
z, x1, x2 = build_graph(len(x1list), len(x2list))
sess = tf.Session()
print(sess.run(z, feed_dict = {x1: x1list, x2: x2list}))
sess.close()
```
## Exercise 5 (Difficult: FUN)
### Linear Regression with tensorflow
https://onlinecourses.science.psu.edu/stat501/node/382/
Consider the following dataset
```
x = [4.0, 4.5, 5.0, 5.5, 6.0, 6.5, 7.0]
y = [33, 42, 45, 51, 53, 61, 62]
```
We want to find the best parameters $p_0$ and $p_1$ that minimise the MSE (mean squared error) for the data given, in other words we want to do a linear regression on the data $(x,y)$. Given that a matrix solution to find the best parameter is
$$
{\bf p} =(X^TX)^{-1} X^T Y
$$
where $X^T$ is the transpose of the matrix $X$. The matrix $X$ is defined as
$$
X =
\begin{bmatrix}
1 & x_1 \\
... & ... \\
1 & x_n
\end{bmatrix}
$$
The matrix $Y$ is simply a matrix $n\times 1$ containing the values $y_i$.
dimensions are:
- $X$ has dimensions $n\times 2$
- $Y$ has dimensions $n\times 1$
- ${\bf p}$ has dimensions $2\times 1$
Build a computational graph that evaluates $\bf p$ as given above, given the matrices $X$ and $Y$. Note you will have to build the matrices from the data given at the beginning. If you need more information a beatifully long explanation can be found here https://onlinecourses.science.psu.edu/stat501/node/382/
Let's convert ```y``` to a floating list... **Remeber tensorflow is really strict with datatypes**.
```
y = [float(i) for i in y]
y
x = pd.DataFrame(x)
y = pd.DataFrame(y)
x['b'] = 1
x.head()
cols = x.columns.tolist()
cols = cols[-1:] + cols[:-1]
print(cols)
x = x[cols]
x.head()
```
Let's build the computational graph:
**NOTE: if you use tf.float32 you will get results that are slightly different than numpy. So be aware. To be safe you can use ```float64```.**
Always try to be as specific
as you can with dimensions
The first dimensions is defined as "None" so that we use, in necessary,
with different number of observations without rebuilding the graph.
```
tf.reset_default_graph()
xinput = tf.placeholder(tf.float64, [None,2])
yinput = tf.placeholder(tf.float64, [None,1])
```
Multiplication between tensors is somewhat complicated, especially when dealing
with tensors with more dimensions. So we use the method
https://www.tensorflow.org/api_docs/python/tf/einsum
check it out to get more information.
```
tmp = tf.einsum('ij,jk->ik',tf.transpose(xinput) , xinput)
part1 = tf.linalg.inv(tmp)
part2 = tf.einsum('ij,jk->ik',tf.transpose(xinput), yinput)
pout = tf.einsum('ij,jk->ik', part1, part2)
# Reference: https://www.tensorflow.org/api_docs/python/tf/einsum
sess = tf.Session()
print("The best parameters p are:")
print(sess.run(pout, feed_dict = {xinput: x, yinput: y}))
sess.close()
```
If you remember the first week (check https://github.com/michelucci/dlcourse2018_students/blob/master/Week%201%20-%20Mathematic%20introduction/Week%201%20-%20Solution%20to%20exercises.ipynb) you can do the same with ```numpy```
```
part1np = np.linalg.inv(np.matmul(x.transpose() , x))
part2np = np.matmul(x.transpose(), y)
pnp = np.matmul(part1np, part2np)
print(pnp)
```
## Computational Graph for predictions
The same result we got with tensorflow. Now we can build a graph that will use the ```p``` we have found for predictions
```
p = tf.placeholder(tf.float32, [2,1])
xnode = tf.placeholder(tf.float32, [None, 2]) # This time let's be specific with dimensions
pred = tf.tensordot(xnode, p, axes = 1)
sess = tf.Session()
pred_y = sess.run(pred, feed_dict = {p: pnp, xnode: x})
pred_y
```
And those are the **true** values
```
y
```
## Plot of the results
```
plt.rc('font', family='arial')
plt.rc('xtick', labelsize='x-small')
plt.rc('ytick', labelsize='x-small')
plt.tight_layout()
fig = plt.figure(figsize=(8, 5))
ax = fig.add_subplot(1, 1, 1)
ax.scatter(y, pred_y, lw = 0.3, s = 80)
ax.plot([y.min(), y.max()], [y.min(), y.max()], 'k--', lw = 3)
ax.set_xlabel('Measured Target Value', fontsize = 16);
ax.set_ylabel('Predicted Target Value', fontsize = 16);
plt.tick_params(labelsize=16)
```
| github_jupyter |
# Systematic correction of protein distribution moments
(c) 2020 Manuel Razo. This work is licensed under a [Creative Commons Attribution License CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/). All code contained herein is licensed under an [MIT license](https://opensource.org/licenses/MIT)
---
```
import os
import pickle
import cloudpickle
import itertools
import glob
import git
# Our numerical workhorses
import numpy as np
import scipy as sp
import pandas as pd
import statsmodels.api as sm
# Import libraries to parallelize processes
from joblib import Parallel, delayed
# Import matplotlib stuff for plotting
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import matplotlib as mpl
# Seaborn, useful for graphics
import seaborn as sns
# Import the project utils
import ccutils
# Magic function to make matplotlib inline; other style specs must come AFTER
%matplotlib inline
# This enables SVG graphics inline
%config InlineBackend.figure_format = 'retina'
# Find home directory for repo
repo = git.Repo("./", search_parent_directories=True)
homedir = repo.working_dir
tmpdir = f'{homedir}/tmp/'
figdir = f'{homedir}/fig/moment_dynamics_numeric/'
datadir = f'{homedir}/data/csv_maxEnt_dist/'
# Set PBoC plotting format
ccutils.viz.set_plotting_style()
# Increase dpi
mpl.rcParams['figure.dpi'] = 110
```
### $\LaTeX$ macros
$\newcommand{kpon}{k^{(p)}_{\text{on}}}$
$\newcommand{kpoff}{k^{(p)}_{\text{off}}}$
$\newcommand{kron}{k^{(r)}_{\text{on}}}$
$\newcommand{kroff}{k^{(r)}_{\text{off}}}$
$\newcommand{rm}{r _m}$
$\newcommand{gm}{\gamma _m}$
$\newcommand{rp}{r _p}$
$\newcommand{gp}{\gamma _p}$
$\newcommand{mm}{\left\langle m \right\rangle}$
$\newcommand{ee}[1]{\left\langle #1 \right\rangle}$
$\newcommand{bb}[1]{\mathbf{#1}}$
$\newcommand{foldchange}{\text{fold-change}}$
$\newcommand{\ee}[1]{\left\langle #1 \right\rangle}$
$\newcommand{\bb}[1]{\mathbf{#1}}$
$\newcommand{\dt}[1]{{\partial{#1} \over \partial t}}$
$\newcommand{\Km}{\bb{K}}$
$\newcommand{\Rm}{\bb{R}_m}$
$\newcommand{\Gm}{\bb{\Gamma}_m}$
$\newcommand{\Rp}{\bb{R}_p}$
$\newcommand{\Gp}{\bb{\Gamma}_p}$
## Systematic exploration of correction for protein distribution moments
As we showed in the `moment_dynamics_cell_division.ipynb` notebook there is a systematic deviation between our model predictions of the noise in gene expression and the experimental measurements. As a reminder we define the noise in protein copy number $p$ as
$$
\text{noise} \equiv {\sqrt{\ee{p^2} - \ee{p}^2} \over \ee{p}},
\tag{1}
$$
To gain some intuition we'll first take a quick look at the plot shown before of our predictions of the noise in gene expression. For this we'll first load the experimental data.
```
df_noise = pd.read_csv(
f'{homedir}/data/csv_microscopy/microscopy_noise_bootstrap.csv',
index_col=0
)
df_noise = df_noise[df_noise.percentile == 0.95]
df_noise.head()
```
As well as the theoretical predictions
```
# Read moment predictions for fine IPTG grid
df_mom_iptg = pd.read_csv(datadir + 'MaxEnt_multi_prom_IPTG_range.csv')
# Find the mean unregulated levels to compute the fold-change
mean_m_delta = np.mean(
df_mom_iptg[df_mom_iptg.repressor==0].m1p0
)
mean_p_delta = np.mean(
df_mom_iptg[df_mom_iptg.repressor==0].m0p1
)
# Compute the noise for the multi-promoter data
df_mom_iptg = df_mom_iptg.assign(
m_noise=np.sqrt(df_mom_iptg.m2p0 - df_mom_iptg.m1p0**2) /
df_mom_iptg.m1p0,
p_noise=np.sqrt(df_mom_iptg.m0p2 - df_mom_iptg.m0p1**2) /
df_mom_iptg.m0p1,
m_fold_change=df_mom_iptg.m1p0 / mean_m_delta,
p_fold_change=df_mom_iptg.m0p1 / mean_p_delta
)
# Read moment predictions for fine repressor grid
df_mom_rep = pd.read_csv(datadir + 'MaxEnt_multi_prom_constraints.csv')
# Find the mean unregulated levels to compute the fold-change
mean_m_delta = np.mean(
df_mom_rep[df_mom_rep.repressor==0].m1p0
)
mean_p_delta = np.mean(
df_mom_rep[df_mom_rep.repressor==0].m0p1
)
df_mom_rep = df_mom_rep.assign(
m_noise=np.sqrt(df_mom_rep.m2p0 - df_mom_rep.m1p0**2) /
df_mom_rep.m1p0,
p_noise=np.sqrt(df_mom_rep.m0p2 - df_mom_rep.m0p1**2) /
df_mom_rep.m0p1,
m_fold_change=df_mom_rep.m1p0 / mean_m_delta,
p_fold_change=df_mom_rep.m0p1 / mean_p_delta
)
df_mom_rep.head()
```
Having loaded the predictions and the data let's compare them both. But first let's generate the groups that we will need, as well as the color palettes that we will use.
```
# Define repressor copy numbers to include
rep = [22, 260, 1740]
# Generate index for each opeartor
operators = ['O1', 'O2', 'O3']
op_idx = dict(zip(operators, np.arange(3)))
# Define energies to go along operators
energies = [-15.3, -13.9, -9.7]
# Extract regulated promoter information
df_noise_reg = df_noise[df_noise.repressor > 0]
# Define repressor copy numbers to include
rep = df_noise_reg["repressor"].unique()
# Group moments by operator and repressor
df_group_exp = (
df_noise_reg[df_noise_reg.noise > 0]
.sort_values("IPTG_uM")
.groupby(["operator", "repressor"])
)
df_group = (
df_mom_iptg[df_mom_iptg["repressor"].isin(rep)]
.sort_values("inducer_uM")
.groupby(["operator", "repressor"])
)
# Generate index for each opeartor
operators = ["O1", "O2", "O3"]
op_idx = dict(zip(operators, np.arange(3)))
# Generate list of colors
col_list = ["Blues_r", "Oranges_r", "Greens_r"]
# Loop through operators generating dictionary of colors for each
col_dict = {}
for i, op in enumerate(operators):
col_dict[op] = dict(
zip(rep, sns.color_palette(col_list[i], n_colors=len(rep) + 1)[0:3])
)
# Define threshold to separate log scale from linear scale
thresh = 1e-1
```
Now let's plot the noise as a function of the inducer (IPTG) concentration.
```
# Initialize figure
fig, ax = plt.subplots(
2,
3,
figsize=(7, 2.5),
sharex=True,
sharey="row",
gridspec_kw={"height_ratios": [1, 5], "wspace": 0.05, "hspace": 0},
)
ax = ax.ravel()
# Loop through groups on multi-promoter
for i, (group, data) in enumerate(df_group):
# Log scale
ax[op_idx[group[0]] + 3].plot(
data[data.inducer_uM >= thresh].inducer_uM,
data[data.inducer_uM >= thresh].p_noise,
color=col_dict[group[0]][group[1]],
label=int(group[1]),
)
# Linear scale
ax[op_idx[group[0]] + 3].plot(
data[data.inducer_uM <= thresh].inducer_uM,
data[data.inducer_uM <= thresh].p_noise,
color=col_dict[group[0]][group[1]],
label="",
linestyle=":",
)
# Set threshold for data
dthresh = 10
# Loop through groups on experimental data
for i, (group, data) in enumerate(df_group_exp):
# Plot data points on lower plot
ax[op_idx[group[0]] + 3].errorbar(
x=data.IPTG_uM,
y=data.noise,
yerr=[data.noise - data.noise_lower, data.noise_upper - data.noise],
fmt="o",
ms=3.5,
color=col_dict[group[0]][group[1]],
label="",
)
# Plot same data points with different plotting style on the upper row
ax[op_idx[group[0]]].plot(
data[data.noise > dthresh].IPTG_uM,
data[data.noise > dthresh].noise,
linestyle="--",
color="w",
label="",
lw=0,
marker="o",
markersize=3,
markeredgecolor=col_dict[group[0]][group[1]],
)
# Set scales of reference plots and the other ones will follow
ax[0].set_xscale("symlog", linthreshx=thresh, linscalex=1)
ax[0].set_yscale("log")
ax[3].set_yscale("log")
# Set limits of reference plots and the rest will folow
ax[3].set_ylim(top=6)
ax[0].set_ylim([6, 5e2])
# Set ticks for the upper plot
ax[0].set_yticks([1e1, 1e2])
# Define location for secondary legend
leg2_loc = ["lower left"] * 2 + ["upper left"]
for i in range(3):
# Set title
label = r"$\Delta\epsilon_r$ = {:.1f} $k_BT$".format(energies[i])
ax[i].set_title(label, bbox=dict(facecolor="#ffedce"))
# Label axis
ax[i + 3].set_xlabel(r"IPTG ($\mu$M)")
# Set legend
leg = ax[i + 3].legend(title="rep./cell", fontsize=7)
# Set legend font size
plt.setp(leg.get_title(), fontsize=8)
ax[3].set_ylabel(r"noise");
```
We can see that the model has the right scaling buy systematically fails to pass through the points. Let's take a different look of this. We'll plot the predicted noise vs the measured noise. For this we'll first add an extra column to the experimental data `DataFrame` containing the theoreticla prediction.
```
# Initialize list to save theoretical noise
# and theoretical mean
thry_noise = list()
thry_mean = list()
# Iterate through rows
for idx, row in df_noise.iterrows():
# Extract information
rep = float(row.repressor)
op = row.operator
if np.isnan(row.IPTG_uM):
iptg = 0
else:
iptg = row.IPTG_uM
# Extract equivalent theoretical prediction
thry = df_mom_rep[(df_mom_rep.repressor == rep) &
(df_mom_rep.operator == op) &
(df_mom_rep.inducer_uM == iptg)].p_noise
# Append to list
thry_noise.append(thry.iloc[0])
# Extract equivalent theoretical prediction
thry = df_mom_rep[(df_mom_rep.repressor == rep) &
(df_mom_rep.operator == op) &
(df_mom_rep.inducer_uM == iptg)].m0p1
# Append to list
thry_mean.append(thry.iloc[0])
df_noise = df_noise.assign(
noise_theory=thry_noise,
mean_theory=thry_mean
)
df_noise.head()
```
Let's now plot predicted vs. measured noise. We'll display both a linear and a log-log plot.
```
# Initialize figure
fig, ax = plt.subplots(1, 2, figsize=(7, 3))
# Linear scale
# Plot reference line
ax[0].plot([1e-2, 1e2], [1e-2, 1e2], "--", color="gray")
# Plot error bars
ax[0].errorbar(
x=df_noise.noise_theory,
y=df_noise.noise,
yerr=[
df_noise.noise - df_noise.noise_lower,
df_noise.noise_upper - df_noise.noise,
],
color="gray",
alpha=0.5,
mew=0,
zorder=0,
fmt=".",
)
# Plot data with color depending on log fold-change
ax[0].scatter(
df_noise.noise_theory,
df_noise.noise,
c=np.log10(df_noise.fold_change),
cmap="viridis",
s=10,
)
ax[0].set_xlabel("theoretical noise")
ax[0].set_ylabel("experimental noise")
ax[0].set_title("linear scale")
# ax[0].set_xticks([0, 1, 2, 3, 4])
# ax[0].set_yticks([0, 1, 2, 3, 4])
ax[0].set_xlim(0, 2)
ax[0].set_ylim(0, 2)
# Log scale
# Plot reference line
line = [1e-1, 1e2]
ax[1].loglog(line, line, "--", color="gray")
# Plot data with color depending on log fold-change
ax[1].errorbar(
x=df_noise.noise_theory,
y=df_noise.noise,
yerr=[
df_noise.noise - df_noise.noise_lower,
df_noise.noise_upper - df_noise.noise,
],
color="gray",
alpha=0.5,
mew=0,
zorder=0,
fmt=".",
)
plot = ax[1].scatter(
df_noise.noise_theory,
df_noise.noise,
c=np.log10(df_noise.fold_change),
cmap="viridis",
s=10,
)
ax[1].set_xlabel("theoretical noise")
ax[1].set_ylabel("experimental noise")
ax[1].set_title("log scale")
ax[1].set_xlim([0.1, 10])
# show color scale
fig.subplots_adjust(right=0.8)
cbar_ax = fig.add_axes([0.82, 0.15, 0.02, 0.7])
cbar = fig.colorbar(plot, cax=cbar_ax, ticks=[0, -1, -2, -3])
cbar.ax.set_ylabel("fold-change")
cbar.ax.set_yticklabels(["1", "0.1", "0.01", "0.001"])
cbar.ax.tick_params(width=0)
plt.subplots_adjust(wspace=0.3)
```
Again, the scaling of the predictions is in agreement with the data, but the bulk of the measurements, i.e. the noise measurements between 0 and 1, are all systematically above the curve. We'll now try some empirical adjustments to guide our intuition of the origin of these deviations.
### Empirical multiplicative constant
The first thing that we will attempt to fix this data is a simple empirical constant. What this means is that the experimental noise and the theoretical noise are related as
$$
\text{noise}_{\exp} = \alpha \cdot \text{noise}_{\text{theory}},
\tag{2}
$$
where $\alpha$ is a multiplicative constant that accounts for the systematic deviation between our predictions and the data. Since our predictions fall below the measured noise we expect $\alpha > 1$. To find this value we will perform a linear regression with a **fixed intercept at zero**. To give less weight to the measurements with extremely large noise values (associated with measurement errors since the expression of these highly repressed cells is very close to the autofluorescence background) we will perform the regression in log scale. We'll also weigh each of the datum by the width of the bootstrap confidence interval on the noise measurement. In this way we force points with high noise and and high confidence interval to influence less in the regression.
Notice that fitting the slope in linear scale is equivalent to finding an intercept for a fix slope in log scale. Let's go ahead and perform this regression. We will weight each of the datum by the width of their confidence interval and their fold-change value.
```
# Select data with experimetal noise < 10
data = df_noise[df_noise.noise < 10]
# Define the weights for each of the datum to be the width
# of their bootstrap confidence interval.
noise_range = (data.noise_upper.values - data.noise_lower.values)
weights = noise_range
# Assign the non-zero minimum value to all zero weights
weights[weights == 0] = min(weights[weights > 0])
# Normalize weights
weights = weights / weights.sum()
def mult_factor(x, a):
'''
Function to find additive constant used with scipy curve_fit
'''
return a * x
popt, pcov = sp.optimize.curve_fit(
mult_factor,
data.noise_theory.values,
data.noise.values,
sigma=weights,
)
multiplicative = popt[0]
# Print result
print(
f"Multiplicative factor: {multiplicative}"
)
```
So the multiplicative factor is ≈ 1.5. Let's plot the predictions multiplied by this factor vs the data to see if there is an improvement.
```
# Initialize figure
fig, ax = plt.subplots(1, 2, figsize=(7, 3))
# Linear scale
# Plot reference line
ax[0].plot([1E-2, 1E2], [1E-2, 1E2], '--', color='gray')
# Plot error bars
ax[0].errorbar(x=df_noise.noise_theory * multiplicative,
y=df_noise.noise,
yerr=[df_noise.noise - df_noise.noise_lower,
df_noise.noise_upper - df_noise.noise],
color='gray',
alpha=0.5,
mew=0,
zorder=0,
fmt='.')
# Plot data with color depending on log fold-change
ax[0].scatter(df_noise.noise_theory * multiplicative, df_noise.noise,
c=np.log10(df_noise.fold_change), cmap='viridis',
s=10)
ax[0].set_xlabel('theoretical noise')
ax[0].set_ylabel('experimental noise')
ax[0].set_title('linear scale')
ax[0].set_xlim(0, 2)
ax[0].set_ylim(0, 2);
# ax[0].set_xticks([0, 1, 2, 3, 4, 6])
# ax[0].set_yticks([0, 1, 2, 3, 4, 6])
# Log scale
# Plot reference line
line = [1E-1, 1E2]
ax[1].loglog(line, line, '--', color='gray')
# Plot data with color depending on log fold-change
ax[1].errorbar(x=df_noise.noise_theory * multiplicative,
y=df_noise.noise,
yerr=[df_noise.noise - df_noise.noise_lower,
df_noise.noise_upper - df_noise.noise],
color='gray',
alpha=0.5,
mew=0,
zorder=0,
fmt='.')
plot = ax[1].scatter(df_noise.noise_theory * multiplicative,
df_noise.noise,
c=np.log10(df_noise.fold_change), cmap='viridis',
s=10)
ax[1].set_xlabel('theoretical noise')
ax[1].set_ylabel('experimental noise')
ax[1].set_title('log scale')
ax[1].set_xlim([0.1, 10])
# show color scale
fig.subplots_adjust(right=0.8)
cbar_ax = fig.add_axes([0.82, 0.15, 0.02, 0.7])
cbar = fig.colorbar(plot, cax=cbar_ax, ticks=[0, -1, -2, -3])
cbar.ax.set_ylabel('fold-change')
cbar.ax.set_yticklabels(['1', '0.1', '0.01', '0.001'])
cbar.ax.tick_params(width=0)
plt.subplots_adjust(wspace=0.3)
```
there is definitely an improvement. Let's take a different look. We'll plot again noise as a function of inducer concentration.
```
# Initialize figure
fig, ax = plt.subplots(
2,
3,
figsize=(7, 2.5),
sharex=True,
sharey="row",
gridspec_kw={"height_ratios": [1, 5], "wspace": 0.05, "hspace": 0},
)
ax = ax.ravel()
# Loop through groups on multi-promoter
for i, (group, data) in enumerate(df_group):
# Log scale
ax[op_idx[group[0]] + 3].plot(
data[data.inducer_uM >= thresh].inducer_uM,
data[data.inducer_uM >= thresh].p_noise * multiplicative,
color=col_dict[group[0]][group[1]],
label=int(group[1]),
)
# Linear scale
ax[op_idx[group[0]] + 3].plot(
data[data.inducer_uM <= thresh].inducer_uM,
data[data.inducer_uM <= thresh].p_noise * multiplicative,
color=col_dict[group[0]][group[1]],
label="",
linestyle=":",
)
# Set threshold for data
dthresh = 10
# Loop through groups on experimental data
for i, (group, data) in enumerate(df_group_exp):
# Plot data points on lower plot
ax[op_idx[group[0]] + 3].errorbar(
x=data.IPTG_uM,
y=data.noise,
yerr=[data.noise - data.noise_lower, data.noise_upper - data.noise],
fmt="o",
ms=3.5,
color=col_dict[group[0]][group[1]],
label="",
)
# Plot same data points with different plotting style on the upper row
ax[op_idx[group[0]]].plot(
data[data.noise > dthresh].IPTG_uM,
data[data.noise > dthresh].noise,
linestyle="--",
color="w",
label="",
lw=0,
marker="o",
markersize=3,
markeredgecolor=col_dict[group[0]][group[1]],
)
# Set scales of reference plots and the other ones will follow
ax[0].set_xscale("symlog", linthreshx=thresh, linscalex=1)
ax[0].set_yscale("log")
ax[3].set_yscale("log")
# Set limits of reference plots and the rest will folow
ax[3].set_ylim(top=8)
ax[0].set_ylim([8, 5e2])
# Set ticks for the upper plot
ax[0].set_yticks([1e1, 1e2])
# Define location for secondary legend
leg2_loc = ["lower left"] * 2 + ["upper left"]
for i in range(3):
# Set title
label = r"$\Delta\epsilon_r$ = {:.1f} $k_BT$".format(energies[i])
ax[i].set_title(label, bbox=dict(facecolor="#ffedce"))
# Label axis
ax[i + 3].set_xlabel(r"IPTG ($\mu$M)")
# Set legend
leg = ax[i + 3].legend(title="rep./cell", fontsize=7)
# Set legend font size
plt.setp(leg.get_title(), fontsize=8)
ax[3].set_ylabel(r"noise");
```
There is a notorious improvement for our noise predictions. For completeness let's test another potential empirical fix.
### Empirical additive constant
Another possible empirical improvement from our predictions would come from an additive constant. What this means is that the experimental and theoretical noise are related as
$$
\text{noise}_{\exp} = \beta + \text{noise}_{\text{theory}},
\tag{3}
$$
where $\beta$ is our empirical additive constant. Since there is no easy way to do this in log scale, let's try it in linear scale. Again we will weight each of the datum by their error and their fold-change values.
```
# Select data with experimetal noise < 10
data = df_noise[df_noise.noise < 10]
# Define the weights for each of the datum to be the width
# of their bootstrap confidence interval.
noise_range = (data.noise_upper.values - data.noise_lower.values)
weights = noise_range
# Assign the non-zero minimum value to all zero weights
weights[weights == 0] = min(weights[weights > 0])
# Normalize weights
weights = weights / weights.sum()
def add_factor(x, a):
'''
Function to find additive constant used with scipy curve_fit
'''
return a + x
popt, pcov = sp.optimize.curve_fit(
add_factor,
data.noise_theory.values,
data.noise.values,
sigma=weights,
)
additive = popt[0]
# Print result
print(
f"Additive factor: {additive}"
)
```
Just as before, let's take a look a what the additive constant does to the data.
```
# Initialize figure
fig, ax = plt.subplots(1, 2, figsize=(7, 3))
# Linear scale
# Plot reference line
ax[0].plot([1E-2, 1E2], [1E-2, 1E2], '--', color='gray')
# Plot error bars
ax[0].errorbar(x=df_noise.noise_theory + additive,
y=df_noise.noise,
yerr=[df_noise.noise - df_noise.noise_lower,
df_noise.noise_upper - df_noise.noise],
color='gray',
alpha=0.5,
mew=0,
zorder=0,
fmt='.')
# Plot data with color depending on log fold-change
ax[0].scatter(df_noise.noise_theory + additive, df_noise.noise,
c=np.log10(df_noise.fold_change), cmap='viridis',
s=10)
ax[0].set_xlabel('theoretical noise')
ax[0].set_ylabel('experimental noise')
ax[0].set_title('linear scale')
ax[0].set_xlim(0, 2)
ax[0].set_ylim(0, 2);
# ax[0].set_xticks([0, 1, 2, 3, 4, 6])
# ax[0].set_yticks([0, 1, 2, 3, 4, 6])
# Log scale
# Plot reference line
line = [1E-1, 1E2]
ax[1].loglog(line, line, '--', color='gray')
# Plot data with color depending on log fold-change
ax[1].errorbar(x=df_noise.noise_theory + additive,
y=df_noise.noise,
yerr=[df_noise.noise - df_noise.noise_lower,
df_noise.noise_upper - df_noise.noise],
color='gray',
alpha=0.5,
mew=0,
zorder=0,
fmt='.')
plot = ax[1].scatter(df_noise.noise_theory + additive,
df_noise.noise,
c=np.log10(df_noise.fold_change), cmap='viridis',
s=10)
ax[1].set_xlabel('theoretical noise')
ax[1].set_ylabel('experimental noise')
ax[1].set_title('log scale')
ax[1].set_xlim([0.1, 10])
# show color scale
fig.subplots_adjust(right=0.8)
cbar_ax = fig.add_axes([0.82, 0.15, 0.02, 0.7])
cbar = fig.colorbar(plot, cax=cbar_ax, ticks=[0, -1, -2, -3])
cbar.ax.set_ylabel('fold-change')
cbar.ax.set_yticklabels(['1', '0.1', '0.01', '0.001'])
cbar.ax.tick_params(width=0)
plt.subplots_adjust(wspace=0.3)
```
For completeness, let's look at the noise as a function of inducer concentration.
```
# Initialize figure
fig, ax = plt.subplots(
2,
3,
figsize=(7, 2.5),
sharex=True,
sharey="row",
gridspec_kw={"height_ratios": [1, 5], "wspace": 0.05, "hspace": 0},
)
ax = ax.ravel()
# Loop through groups on multi-promoter
for i, (group, data) in enumerate(df_group):
# Log scale
ax[op_idx[group[0]] + 3].plot(
data[data.inducer_uM >= thresh].inducer_uM,
data[data.inducer_uM >= thresh].p_noise + additive,
color=col_dict[group[0]][group[1]],
label=int(group[1]),
)
# Linear scale
ax[op_idx[group[0]] + 3].plot(
data[data.inducer_uM <= thresh].inducer_uM,
data[data.inducer_uM <= thresh].p_noise + additive,
color=col_dict[group[0]][group[1]],
label="",
linestyle=":",
)
# Set threshold for data
dthresh = 10
# Loop through groups on experimental data
for i, (group, data) in enumerate(df_group_exp):
# Plot data points on lower plot
ax[op_idx[group[0]] + 3].errorbar(
x=data.IPTG_uM,
y=data.noise,
yerr=[data.noise - data.noise_lower, data.noise_upper - data.noise],
fmt="o",
ms=3.5,
color=col_dict[group[0]][group[1]],
label="",
)
# Plot same data points with different plotting style on the upper row
ax[op_idx[group[0]]].plot(
data[data.noise > dthresh].IPTG_uM,
data[data.noise > dthresh].noise,
linestyle="--",
color="w",
label="",
lw=0,
marker="o",
markersize=3,
markeredgecolor=col_dict[group[0]][group[1]],
)
# Set scales of reference plots and the other ones will follow
ax[0].set_xscale("symlog", linthreshx=thresh, linscalex=1)
ax[0].set_yscale("log")
ax[3].set_yscale("log")
# Set limits of reference plots and the rest will folow
ax[3].set_ylim(top=8)
ax[0].set_ylim([8, 5e2])
# Set ticks for the upper plot
ax[0].set_yticks([1e1, 1e2])
# Define location for secondary legend
leg2_loc = ["lower left"] * 2 + ["upper left"]
for i in range(3):
# Set title
label = r"$\Delta\epsilon_r$ = {:.1f} $k_BT$".format(energies[i])
ax[i].set_title(label, bbox=dict(facecolor="#ffedce"))
# Label axis
ax[i + 3].set_xlabel(r"IPTG ($\mu$M)")
# Set legend
leg = ax[i + 3].legend(title="rep./cell", fontsize=7)
# Set legend font size
plt.setp(leg.get_title(), fontsize=8)
ax[3].set_ylabel(r"noise");
```
One could argue that this looks even better than the multiplicative constant. As a way of comparison we will compute an $R^2$ for each of the cases. Notices that although the noise as a function of inducer concentration is a non-linear fit for which we couldn't compute an $R^2$, both the multiplicative and the additive constant as we inferred them are simple linear linear regressions for which we can compute such numbers. Only large differences in these numbers will be revealing.
```
data = df_noise[df_noise.noise < 10]
# Original regression
r_sqr_original = np.corrcoef(
data.noise_theory.values,
data.noise.values,
)[0, 1]**2
# Multiplicative constant
r_sqr_mult = np.corrcoef(
data.noise_theory.values * multiplicative,
data.noise.values,
)[0, 1]**2
# Additive constant
r_sqr_add = np.corrcoef(
data.noise_theory.values + additive,
data.noise.values,
)[0, 1]**2
print(
f'''
Original R**2 = {r_sqr_original}
Multiplicative factor R**2 = {r_sqr_mult}
Additive factor R**2 = {r_sqr_add}
'''
)
```
Completely indistinguishable as far as this metric is concerned.
| github_jupyter |
# QuakeMigrate - Example - Icequake detection
## Overview:
This notebook shows how to run QuakeMigrate for icequake detection, using a 2 minute window of continuous seismic data from Hudson et al (2019). Please refer to this paper for details and justification of the settings used.
Here, we detail how to:
1. Create a travel-times lookup table for the example seismometer network
2. Run the detect stage to coalesce energy through time
3. Run the trigger stage to determine events above a threshold value
4. Run the locate stage to refine the earthquake location
We also provide an outline of some of the key outputs
```
# Import necessary modules:
import QMigrate.core.model as qmod
import QMigrate.signal.scan as qscan
import QMigrate.io.data as qdata
import QMigrate.io.quakeio as qio
import QMigrate.signal.trigger as qtrigger
# Set i/o paths:
station_file = "./inputs/stations.txt"
data_in = "./inputs/mSEED"
lut_out = "./outputs/lut/icequake.LUT"
out_path = "./outputs/runs"
run_name = "icequake_example"
```
## 1. Create a travel-times lookup table (LUT)
```
# Read in station information
stations = qio.stations(station_file)
# Set the parameters for the travel-times lookup table (LUT)
# Cell count (x,y,z); cell size (x,y,z in metres)
lut = qmod.LUT(stations, cell_count=[20, 20, 140], cell_size=[100, 100, 20])
lut.lonlat_centre(-17.224, 64.328)
# Set the LUT projection (here we use the Lambert Conformal Conic projection)
lut.lcc_standard_parallels = (64.32, 64.335)
lut.projections(grid_proj_type="LCC")
lut.elevation=1400 # Defining the elevation of the top of the grid in m
# Compute for a homogeneous velocity model
v_p_homo_model = 3630
v_s_homo_model = 1833
lut.compute_homogeneous_vmodel(v_p_homo_model, v_s_homo_model)
# Save the LUT
lut.save(lut_out)
```
## 2. Coalesce the seismic energy through time
```
# Create a new instance of the MSEED class and set path structure
data = qdata.Archive(station_file=station_file, archive_path=data_in)
data.path_structure(archive_format="YEAR/JD/*_STATION_*")
# Create a new instance of the SeisScan class
scan = qscan.QuakeScan(data, lut_out, output_path=out_path, run_name=run_name)
# Set detect parameters
scan.sampling_rate = 500 # Sampling rate of data, in Hz
scan.p_bp_filter = [10, 125, 4] # The band-pass filter parameters for the P-phase (10 to 125 Hz, with 4th order corners)
scan.s_bp_filter = [10, 125, 4] # The band-pass filter parameters for the P-phase (10 to 125 Hz, with 4th order corners)
scan.p_onset_win = [0.01, 0.25] # Length of the STA and LTA time windows for the P-phase
scan.s_onset_win = [0.05, 0.5] # Length of the STA and LTA time windows for the S-phase
scan.time_step = 0.75 # The length of the time-step
scan.decimate = [1, 1, 1] # Decimation factors in x,y,z (no decimation here)
scan.n_cores = 12 # Number of cores/processors to use
# Defining the start and end times
starttime = "2014-06-29T18:41:55.0"
endtime = "2014-06-29T18:42:20.0"
# Run the detect stage to find the coalescence of energy through time:
scan.detect(starttime, endtime)
```
## 3. Run the trigger stage, to detect and output individual icequakes
nb: We can use the same SeisScan object here because we are not using a different decimation. If running trigger and locate on grids with different levels of decimation, a new SeisScan object must be initialised.
```
trig = qtrigger.Trigger(out_path, run_name, stations)
trig.normalise_coalescence = True
trig.marginal_window = 2.75
trig.minimum_repeat = 6.
trig.detection_threshold = 1.8
# Run trigger
trig.trigger(starttime, endtime, savefig=True)
```
## 4. Run the locate stage, to relocate triggered events on a less decimated grid
```
# Set locate parameters:
scan.marginal_window = 2.75
# Turn on plotting features
scan.plot_coal_video = False
scan.plot_coal_grid = False
scan.plot_coal_picture = True
scan.plot_coal_trace = False
# Run the locate stage to determine the location of any triggered events
scan.locate(starttime, endtime)
```
## 4. Some of the key outputs
```
# Show the .event file, containing event origin time and location:
icequake_event_fname = "./outputs/runs/icequake_example/events/20140629184210330000.event"
with open(icequake_event_fname) as f:
lines = f.readlines()
for line in lines:
print(line)
# Show the .stn file, containing station time picks:
icequake_stn_fname = "outputs/runs/icequake_example/picks/20140629184210330000.picks"
with open(icequake_stn_fname) as f:
lines = f.readlines()
for line in lines:
print(line)
# Show the coalescence pdf file, containing event origin time and location:
icequake_coal_image_fname = "outputs/runs/icequake_example/summaries/icequake_example_20140629184210330000_EventSummary.pdf"
from IPython.display import IFrame # For plotting pdf
IFrame(icequake_coal_image_fname, width=800, height=400) # Plot pdf
```
References:
Hudson, T.S., Smith, J., Brisbourne, A.M., and White R.S. (2019). Automated detection of basal icequakes and discrimination from surface crevassing. Annals of Glaciology, 79
| github_jupyter |
# Homework 1
*This notebook includes both coding and written questions. Please hand in this notebook file with all the outputs and your answers to the written questions.*
This assignment covers linear filters, convolution and correlation
```
# Setup
import numpy as np
import matplotlib.pyplot as plt
from time import time
from skimage import io
from __future__ import print_function
%matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading extenrnal modules
%load_ext autoreload
%autoreload 2
```
## Part 1: Convolutions
### 1.1 Commutative Property (10 points)
Recall that the convolution of an image $f:\mathbb{R}^2\rightarrow \mathbb{R}$ and a kernel $h:\mathbb{R}^2\rightarrow\mathbb{R}$ is defined as follows:
$$(f*h)[m,n]=\sum_{i=-\infty}^\infty\sum_{j=-\infty}^\infty f[i,j]\cdot h[m-i,n-j]$$
Or equivalently,
\begin{align}
(f*h)[m,n] &= \sum_{i=-\infty}^\infty\sum_{j=-\infty}^\infty h[i,j]\cdot f[m-i,n-j]\\
&= (h*f)[m,n]
\end{align}
Show that this is true (i.e. prove that the convolution operator is commutative: $f*h = h*f$).
**Your Answer:** *Write your solution in this markdown cell. Please write your equations in [LaTex equations](http://jupyter-notebook.readthedocs.io/en/latest/examples/Notebook/Typesetting%20Equations.html).*
### 1.2 Linear and Shift Invariance (10 points)
Let $f$ be a function $\mathbb{R}^2\rightarrow\mathbb{R}$. Consider a system $f\xrightarrow{s}g$, where $g=(f*h)$ with some kernel $h:\mathbb{R}^2\rightarrow\mathbb{R}$. Show that $S$ defined by any kernel $h$ is a Linear Shift Invariant (LSI) system. In other words, for any $h$, show that $S$ satisfies both of the following:
- $S[a\cdot{f_1}+b\cdot{f_2}]= a\cdot{S[f_1]}+b\cdot{S[f_2]}$
- If $f[m,n]\xrightarrow{s}g[m,n]$ then $f[m-m_0,n-n_0]\xrightarrow{s}g[m-m_0,n-n_0]$
**Your Answer:** *Write your solution in this markdown cell. Please write your equations in [LaTex equations](http://jupyter-notebook.readthedocs.io/en/latest/examples/Notebook/Typesetting%20Equations.html).*
### 1.3 Implementation (30 points)
In this section, you will implement two versions of convolution:
- `conv_nested`
- `conv_fast`
First, run the code cell below to load the image to work with.
```
# Open image as grayscale
img = io.imread('dog.jpg', as_grey=True)
# Show image
plt.imshow(img)
plt.axis('off')
plt.title("Isn't he cute?")
plt.show()
```
Now, implement the function **`conv_nested`** in **`filters.py`**. This is a naive implementation of convolution which uses 4 nested for-loops. It takes an image $f$ and a kernel $h$ as inputs and outputs the convolved image $(f*h)$ that has the same shape as the input image. This implementation should take a few seconds to run.
*- Hint: It may be easier to implement $(h*f)$*
We'll first test your `conv_nested` function on a simple input.
```
from filters import conv_nested
# Simple convolution kernel.
kernel = np.array(
[
[1,0,1],
[0,0,0],
[1,0,1]
])
# Create a test image: a white square in the middle
test_img = np.zeros((9, 9))
test_img[3:6, 3:6] = 1
# Run your conv_nested function on the test image
test_output = conv_nested(test_img, kernel)
# Build the expected output
expected_output = np.zeros((9, 9))
expected_output[2:7, 2:7] = 1
expected_output[4, 2:7] = 2
expected_output[2:7, 4] = 2
expected_output[4, 4] = 4
# Plot the test image
plt.subplot(1,3,1)
plt.imshow(test_img)
plt.title('Test image')
plt.axis('off')
# Plot your convolved image
plt.subplot(1,3,2)
plt.imshow(test_output)
plt.title('Convolution')
plt.axis('off')
# Plot the exepected output
plt.subplot(1,3,3)
plt.imshow(expected_output)
plt.title('Exepected output')
plt.axis('off')
plt.show()
# Test if the output matches expected output
assert np.max(test_output - expected_output) < 1e-10, "Your solution is not correct."
```
Now let's test your `conv_nested` function on a real image.
```
from filters import conv_nested
# Simple convolution kernel.
# Feel free to change the kernel and to see different outputs.
kernel = np.array(
[
[1,0,-1],
[2,0,-2],
[1,0,-1]
])
out = conv_nested(img, kernel)
# Plot original image
plt.subplot(2,2,1)
plt.imshow(img)
plt.title('Original')
plt.axis('off')
# Plot your convolved image
plt.subplot(2,2,3)
plt.imshow(out)
plt.title('Convolution')
plt.axis('off')
# Plot what you should get
solution_img = io.imread('convoluted_dog.jpg', as_grey=True)
plt.subplot(2,2,4)
plt.imshow(solution_img)
plt.title('What you should get')
plt.axis('off')
plt.show()
```
Let us implement a more efficient version of convolution using array operations in numpy. As shown in the lecture, a convolution can be considered as a sliding window that computes sum of the pixel values weighted by the flipped kernel. The faster version will i) zero-pad an image, ii) flip the kernel horizontally and vertically, and iii) compute weighted sum of the neighborhood at each pixel.
First, implement the function **`zero_pad`** in **`filters.py`**.
```
from filters import zero_pad
pad_width = 20 # width of the padding on the left and right
pad_height = 40 # height of the padding on the top and bottom
padded_img = zero_pad(img, pad_height, pad_width)
# Plot your padded dog
plt.subplot(1,2,1)
plt.imshow(padded_img)
plt.title('Padded dog')
plt.axis('off')
# Plot what you should get
solution_img = io.imread('padded_dog.jpg', as_grey=True)
plt.subplot(1,2,2)
plt.imshow(solution_img)
plt.title('What you should get')
plt.axis('off')
plt.show()
```
Next, complete the function **`conv_fast`** in **`filters.py`** using `zero_pad`. Run the code below to compare the outputs by the two implementations. `conv_fast` should run significantly faster than `conv_nested`.
Depending on your implementation and computer, `conv_nested` should take a few seconds and `conv_fast` should be around 5 times faster.
```
from filters import conv_fast
t0 = time()
out_fast = conv_fast(img, kernel)
t1 = time()
out_nested = conv_nested(img, kernel)
t2 = time()
# Compare the running time of the two implementations
print("conv_nested: took %f seconds." % (t2 - t1))
print("conv_fast: took %f seconds." % (t1 - t0))
# Plot conv_nested output
plt.subplot(1,2,1)
plt.imshow(out_nested)
plt.title('conv_nested')
plt.axis('off')
# Plot conv_fast output
plt.subplot(1,2,2)
plt.imshow(out_fast)
plt.title('conv_fast')
plt.axis('off')
# Make sure that the two outputs are the same
if not (np.max(out_fast - out_nested) < 1e-10):
print("Different outputs! Check your implementation.")
```
### Extra Credit 1 (1% of final grade)
Devise a faster version of convolution and implement **`conv_faster`** in **`filters.py`**. You will earn extra credit only if the `conv_faster` runs faster (by a fair margin) than `conv_fast` **and** outputs the same result.
```
from filters import conv_faster
t0 = time()
out_fast = conv_fast(img, kernel)
t1 = time()
out_faster = conv_faster(img, kernel)
t2 = time()
# Compare the running time of the two implementations
print("conv_fast: took %f seconds." % (t1 - t0))
print("conv_faster: took %f seconds." % (t2 - t1))
# Plot conv_nested output
plt.subplot(1,2,1)
plt.imshow(out_fast)
plt.title('conv_fast')
plt.axis('off')
# Plot conv_fast output
plt.subplot(1,2,2)
plt.imshow(out_faster)
plt.title('conv_faster')
plt.axis('off')
# Make sure that the two outputs are the same
if not (np.max(out_fast - out_faster) < 1e-10):
print("Different outputs! Check your implementation.")
```
---
## Part 2: Cross-correlation
Cross-correlation of two 2D signals $f$ and $g$ is defined as follows:
$$(f\star{g})[m,n]=\sum_{i=-\infty}^\infty\sum_{j=-\infty}^\infty f[i,j]\cdot g[i-m,j-n]$$
### 2.1 Template Matching with Cross-correlation (12 points)
Suppose that you are a clerk at a grocery store. One of your responsibilites is to check the shelves periodically and stock them up whenever there are sold-out items. You got tired of this laborious task and decided to build a computer vision system that keeps track of the items on the shelf.
Luckily, you have learned in CS131 that cross-correlation can be used for template matching: a template $g$ is multiplied with regions of a larger image $f$ to measure how similar each region is to the template.
The template of a product (`template.jpg`) and the image of shelf (`shelf.jpg`) is provided. We will use cross-correlation to find the product in the shelf.
Implement **`cross_correlation`** function in **`filters.py`** and run the code below.
*- Hint: you may use the `conv_fast` function you implemented in the previous question.*
```
from filters import cross_correlation
# Load template and image in grayscale
img = io.imread('shelf.jpg')
img_grey = io.imread('shelf.jpg', as_grey=True)
temp = io.imread('template.jpg')
temp_grey = io.imread('template.jpg', as_grey=True)
# Perform cross-correlation between the image and the template
out = cross_correlation(img_grey, temp_grey)
# Find the location with maximum similarity
y,x = (np.unravel_index(out.argmax(), out.shape))
# Display product template
plt.figure(figsize=(25,20))
plt.subplot(3, 1, 1)
plt.imshow(temp)
plt.title('Template')
plt.axis('off')
# Display cross-correlation output
plt.subplot(3, 1, 2)
plt.imshow(out)
plt.title('Cross-correlation (white means more correlated)')
plt.axis('off')
# Display image
plt.subplot(3, 1, 3)
plt.imshow(img)
plt.title('Result (blue marker on the detected location)')
plt.axis('off')
# Draw marker at detected location
plt.plot(x, y, 'bx', ms=40, mew=10)
plt.show()
```
#### Interpretation
How does the output of cross-correlation filter look like? Was it able to detect the product correctly? Explain what might be the problem with using raw template as a filter.
**Your Answer:** *Write your solution in this markdown cell.*
---
### 2.2 Zero-mean cross-correlation (6 points)
A solution to this problem is to subtract off the mean value of the template so that it has zero mean.
Implement **`zero_mean_cross_correlation`** function in **`filters.py`** and run the code below.
```
from filters import zero_mean_cross_correlation
# Perform cross-correlation between the image and the template
out = zero_mean_cross_correlation(img_grey, temp_grey)
# Find the location with maximum similarity
y,x = (np.unravel_index(out.argmax(), out.shape))
# Display product template
plt.figure(figsize=(30,20))
plt.subplot(3, 1, 1)
plt.imshow(temp)
plt.title('Template')
plt.axis('off')
# Display cross-correlation output
plt.subplot(3, 1, 2)
plt.imshow(out)
plt.title('Cross-correlation (white means more correlated)')
plt.axis('off')
# Display image
plt.subplot(3, 1, 3)
plt.imshow(img)
plt.title('Result (blue marker on the detected location)')
plt.axis('off')
# Draw marker at detcted location
plt.plot(x, y, 'bx', ms=40, mew=10)
plt.show()
```
You can also determine whether the product is present with appropriate scaling and thresholding.
```
def check_product_on_shelf(shelf, product):
out = zero_mean_cross_correlation(shelf, product)
# Scale output by the size of the template
out = out / float(product.shape[0]*product.shape[1])
# Threshold output (this is arbitrary, you would need to tune the threshold for a real application)
out = out > 0.025
if np.sum(out) > 0:
print('The product is on the shelf')
else:
print('The product is not on the shelf')
# Load image of the shelf without the product
img2 = io.imread('shelf_soldout.jpg')
img2_grey = io.imread('shelf_soldout.jpg', as_grey=True)
plt.imshow(img)
plt.axis('off')
plt.show()
check_product_on_shelf(img_grey, temp_grey)
plt.imshow(img2)
plt.axis('off')
plt.show()
check_product_on_shelf(img2_grey, temp_grey)
```
---
### 2.3 Normalized Cross-correlation (12 points)
One day the light near the shelf goes out and the product tracker starts to malfunction. The `zero_mean_cross_correlation` is not robust to change in lighting condition. The code below demonstrates this.
```
from filters import normalized_cross_correlation
# Load image
img = io.imread('shelf_dark.jpg')
img_grey = io.imread('shelf_dark.jpg', as_grey=True)
# Perform cross-correlation between the image and the template
out = zero_mean_cross_correlation(img_grey, temp_grey)
# Find the location with maximum similarity
y,x = (np.unravel_index(out.argmax(), out.shape))
# Display image
plt.imshow(img)
plt.title('Result (red marker on the detected location)')
plt.axis('off')
# Draw marker at detcted location
plt.plot(x, y, 'rx', ms=25, mew=5)
plt.show()
```
A solution is to normalize the pixels of the image and template at every step before comparing them. This is called **normalized cross-correlation**.
The mathematical definition for normalized cross-correlation of $f$ and template $g$ is:
$$(f\star{g})[m,n]=\sum_{i,j} \frac{f[i,j]-\overline{f_{m,n}}}{\sigma_{f_{m,n}}} \cdot \frac{g[i-m,j-n]-\overline{g}}{\sigma_g}$$
where:
- $f_{m,n}$ is the patch image at position $(m,n)$
- $\overline{f_{m,n}}$ is the mean of the patch image $f_{m,n}$
- $\sigma_{f_{m,n}}$ is the standard deviation of the patch image $f_{m,n}$
- $\overline{g}$ is the mean of the template $g$
- $\sigma_g$ is the standard deviation of the template $g$
Implement **`normalized_cross_correlation`** function in **`filters.py`** and run the code below.
```
from filters import normalized_cross_correlation
# Perform normalized cross-correlation between the image and the template
out = normalized_cross_correlation(img_grey, temp_grey)
# Find the location with maximum similarity
y,x = (np.unravel_index(out.argmax(), out.shape))
# Display image
plt.imshow(img)
plt.title('Result (red marker on the detected location)')
plt.axis('off')
# Draw marker at detcted location
plt.plot(x, y, 'rx', ms=25, mew=5)
plt.show()
```
## Part 3: Separable Filters
### 3.1 Theory (10 points)
Consider a $M_1\times{N_1}$ image $I$ and a $M_2\times{N_2}$ filter $F$. A filter $F$ is **separable** if it can be written as a product of two 1D filters: $F=F_1F_2$.
For example,
$$F=
\begin{bmatrix}
1 & -1 \\
1 & -1
\end{bmatrix}
$$
can be written as a matrix product of
$$F_1=
\begin{bmatrix}
1 \\
1
\end{bmatrix},
F_2=
\begin{bmatrix}
1 & -1
\end{bmatrix}
$$
Therefore $F$ is a separable filter.
Prove that for any separable filter $F=F_1F_2$,
$$I*F=(I*F_1)*F_2$$
**Your Answer:** *Write your solution in this markdown cell. Please write your equations in [LaTex equations](http://jupyter-notebook.readthedocs.io/en/latest/examples/Notebook/Typesetting%20Equations.html).*
### 3.2 Complexity comparison (10 points)
(i) How many multiplications do you need to do a direct 2D convolution (i.e. $I*F$?)<br>
(ii) How many multiplications do you need to do 1D convolutions on rows and columns (i.e. $(I*F_1)*F_2$)<br>
(iii) Use Big-O notation to argue which one is more efficient in general: direct 2D convolution or two successive 1D convolutions?
**Your Answer:** *Write your solution in this markdown cell. Please write your equations in [LaTex equations](http://jupyter-notebook.readthedocs.io/en/latest/examples/Notebook/Typesetting%20Equations.html).*
Now, we will empirically compare the running time of a separable 2D convolution and its equivalent two 1D convolutions. Gaussian kernel, widely used for blurring images, is one example of a separable filter. Run the code below to see its effect.
```
# Load image
img = io.imread('dog.jpg', as_grey=True)
# 5x5 Gaussian blur
kernel = np.array(
[
[1,4,6,4,1],
[4,16,24,16,4],
[6,24,36,24,6],
[4,16,24,16,4],
[1,4,6,4,1]
])
t0 = time()
out = conv_nested(img, kernel)
t1 = time()
t_normal = t1 - t0
# Plot original image
plt.subplot(1,2,1)
plt.imshow(img)
plt.title('Original')
plt.axis('off')
# Plot convolved image
plt.subplot(1,2,2)
plt.imshow(out)
plt.title('Blurred')
plt.axis('off')
plt.show()
```
In the below code cell, define the two 1D arrays (`k1` and `k2`) whose product is equal to the Gaussian kernel.
```
# The kernel can be written as outer product of two 1D filters
k1 = None # shape (5, 1)
k2 = None # shape (1, 5)
### YOUR CODE HERE
pass
### END YOUR CODE
# Check if kernel is product of k1 and k2
if not np.all(k1 * k2 == kernel):
print('k1 * k2 is not equal to kernel')
assert k1.shape == (5, 1), "k1 should have shape (5, 1)"
assert k2.shape == (1, 5), "k2 should have shape (1, 5)"
```
We now apply the two versions of convolution to the same image, and compare their running time. Note that the outputs of the two convolutions must be the same.
```
# Perform two convolutions using k1 and k2
t0 = time()
out_separable = conv_nested(img, k1)
out_separable = conv_nested(out_separable, k2)
t1 = time()
t_separable = t1 - t0
# Plot normal convolution image
plt.subplot(1,2,1)
plt.imshow(out)
plt.title('Normal convolution')
plt.axis('off')
# Plot separable convolution image
plt.subplot(1,2,2)
plt.imshow(out_separable)
plt.title('Separable convolution')
plt.axis('off')
plt.show()
print("Normal convolution: took %f seconds." % (t_normal))
print("Separable convolution: took %f seconds." % (t_separable))
# Check if the two outputs are equal
assert np.max(out_separable - out) < 1e-10
```
| github_jupyter |
## Stereo Vision
```
# import libraries
import cv2
import numpy as np
import matplotlib.pyplot as plt
from numba import jit
from math import sqrt
# Read sample images
left = cv2.imread('images/l4.png', 0)
right = cv2.imread('images/r4.png', 0)
plt.figure(figsize=(15,10))
ax1 = plt.subplot(121)
ax1.imshow(left, cmap='gray')
ax2 = plt.subplot(122)
ax2.imshow(right, cmap='gray')
ax1.axis('off')
ax2.axis('off')
plt.show()
```
### SAD (Sum of Absolute Differences)
```
@jit
def sad(left, right, kernel, offset):
result = np.zeros( right.shape, dtype=np.int )
h,w = right.shape
range_depth = 255 / offset;
max_v = kernel * kernel * 255
min_v = 0
for x in range(h):
for y in range(w - kernel):
offset_id = 0
min_sad = kernel * kernel * 255
for o in range(offset):
if y + kernel + o >= w:
break
curr_sad = 0
for yr in range(kernel):
curr_sad += abs(right[x,y + yr] - left[x,y + yr + o])
if curr_sad < min_sad:
min_sad = curr_sad
offset_id = o
result[x,y] = offset_id * range_depth
#result[x,y] = (min_sad / max_v) * 255
return result
result = sad(left, right, 10, 50)
plt.figure(figsize=(15,10))
ax1 = plt.subplot(131)
ax1.imshow(left, cmap='gray')
ax2 = plt.subplot(132)
ax2.imshow(right, cmap='gray')
ax3 = plt.subplot(133)
ax3.imshow(result, cmap='gray')
ax1.axis('off')
ax2.axis('off')
ax3.axis('off')
plt.show()
```
### SSD (Sum of Squared Differences)
```
@jit
def ssd(left, right, kernel, offset):
result = np.zeros( right.shape, dtype=np.int )
h,w = right.shape
range_depth = 255 / offset;
max_v = kernel * kernel * 255
min_v = 0
for x in range(h):
for y in range(w - kernel):
offset_id = 0
min_sad = kernel * kernel * 255
for o in range(offset):
if y + kernel + o >= w:
break
curr_sad = 0
for yr in range(kernel):
curr_sad += (right[x,y + yr] - left[x,y + yr + o]) ** 2
if curr_sad < min_sad:
min_sad = curr_sad
offset_id = o
result[x,y] = offset_id * range_depth
#result[x,y] = (min_sad / max_v) * 255
return result
result = ssd(left, right, 20, 70)
plt.figure(figsize=(15,10))
ax1 = plt.subplot(131)
ax1.imshow(left, cmap='gray')
ax2 = plt.subplot(132)
ax2.imshow(right, cmap='gray')
ax3 = plt.subplot(133)
ax3.imshow(result, cmap='gray')
ax1.axis('off')
ax2.axis('off')
ax3.axis('off')
plt.show()
```
### NCC (Normalized Cross Correlation)
```
@jit
def ncc(left, right, kernel, offset):
result = np.zeros( right.shape, dtype=np.int )
h,w = right.shape
range_depth = 255 / offset;
max_v = kernel * kernel * 255
min_v = 0
for x in range(h):
for y in range(w - kernel):
min_sad = kernel * kernel * 255
mean_right = np.mean(right[ x, y : y + kernel ])
sum_right = right[ x, y : y + kernel ] - mean_right
sum_right_squared = np.sum((right[ x, y : y + kernel ] - mean_right) ** 2)
#print(mean_left, sum_left, sum_left_squared)
offset_id = 0
for o in range(offset):
if y + kernel + o >= w:
break
curr_sad = 0
curr_sum = 0
mean_left = np.mean(left[ x, y + o: y + kernel + o])
sum_left = left[ x, y + o: y + kernel + o] - mean_left
curr_sum = np.sum(np.multiply(sum_left,sum_right))
sum_left_squared = np.sum((left[ x, y + o: y + kernel + o] - mean_left) ** 2)
root = sqrt( sum_left_squared * sum_right_squared )
if root == 0:
curr_sad = 255
else:
curr_sad = curr_sum / root
if curr_sad < min_sad:
min_sad = curr_sad
offset_id = o
result[x,y] = (offset_id) * range_depth
#result[x,y] = (min_sad / max_v) * 255
return result
result = ncc(left, right, 10, 50)
plt.figure(figsize=(15,10))
ax1 = plt.subplot(131)
ax1.imshow(left, cmap='gray')
ax2 = plt.subplot(132)
ax2.imshow(right, cmap='gray')
ax3 = plt.subplot(133)
ax3.imshow(result, cmap='gray')
ax1.axis('off')
ax2.axis('off')
ax3.axis('off')
plt.show()
```
| github_jupyter |
# Implementing an RNN in TensorFlow
----------------------------------
This script implements an RNN in TensorFlow to predict spam/ham from texts.
We start by loading the necessary libraries and initializing a computation graph in TensorFlow.
```
import os
import re
import io
import requests
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from zipfile import ZipFile
from tensorflow.python.framework import ops
ops.reset_default_graph()
# Start a graph
sess = tf.Session()
```
Next we set the parameters for the RNN model.
```
# Set RNN parameters
epochs = 20
batch_size = 250
max_sequence_length = 25
rnn_size = 10
embedding_size = 50
min_word_frequency = 10
learning_rate = 0.0005
dropout_keep_prob = tf.placeholder(tf.float32)
```
We download and save the data next. First we check if we have saved it before and load it locally, if not, we load it from the internet (UCI machine learning data repository).
```
# Download or open data
data_dir = 'temp'
data_file = 'text_data.txt'
if not os.path.exists(data_dir):
os.makedirs(data_dir)
if not os.path.isfile(os.path.join(data_dir, data_file)):
zip_url = 'http://archive.ics.uci.edu/ml/machine-learning-databases/00228/smsspamcollection.zip'
r = requests.get(zip_url)
z = ZipFile(io.BytesIO(r.content))
file = z.read('SMSSpamCollection')
# Format Data
text_data = file.decode()
text_data = text_data.encode('ascii', errors='ignore')
text_data = text_data.decode().split('\n')
# Save data to text file
with open(os.path.join(data_dir, data_file), 'w') as file_conn:
for text in text_data:
file_conn.write("{}\n".format(text))
else:
# Open data from text file
text_data = []
with open(os.path.join(data_dir, data_file), 'r') as file_conn:
for row in file_conn:
text_data.append(row)
text_data = text_data[:-1]
text_data = [x.split('\t') for x in text_data if len(x) >= 1]
[text_data_target, text_data_train] = [list(x) for x in zip(*text_data)]
```
Next, we process the texts and turn them into numeric representations (words --> indices).
```
# Create a text cleaning function
def clean_text(text_string):
text_string = re.sub(r'([^\s\w]|_|[0-9])+', '', text_string)
text_string = " ".join(text_string.split())
text_string = text_string.lower()
return text_string
# Clean texts
text_data_train = [clean_text(x) for x in text_data_train]
# Change texts into numeric vectors
vocab_processor = tf.contrib.learn.preprocessing.VocabularyProcessor(max_sequence_length,
min_frequency=min_word_frequency)
text_processed = np.array(list(vocab_processor.fit_transform(text_data_train)))
```
> Note: there will be a WARNING:... use tensorflow/transform or tf.data. Ignore this for now- there is an issue with getting tensorflow/transform to work. Hopefully this will be fixed soon and the code here will be updated.
Now we shuffle and split the texts into train/tests (80% training, 20% testing).
```
# Shuffle and split data
text_processed = np.array(text_processed)
text_data_target = np.array([1 if x == 'ham' else 0 for x in text_data_target])
shuffled_ix = np.random.permutation(np.arange(len(text_data_target)))
x_shuffled = text_processed[shuffled_ix]
y_shuffled = text_data_target[shuffled_ix]
# Split train/test set
ix_cutoff = int(len(y_shuffled)*0.80)
x_train, x_test = x_shuffled[:ix_cutoff], x_shuffled[ix_cutoff:]
y_train, y_test = y_shuffled[:ix_cutoff], y_shuffled[ix_cutoff:]
vocab_size = len(vocab_processor.vocabulary_)
print("Vocabulary Size: {:d}".format(vocab_size))
print("80-20 Train Test split: {:d} -- {:d}".format(len(y_train), len(y_test)))
```
Here we can define our RNN model. We create the placeholders for the data, word embedding matrices (and embedding lookups), and define the rest of the model.
The rest of the RNN model will create a dynamic RNN cell (regular RNN type), which will vary the number of RNNs needed for variable input length (different amount of words for input texts), and then output into a fully connected logistic layer to predict spam or ham as output.
```
# Create placeholders
x_data = tf.placeholder(tf.int32, [None, max_sequence_length])
y_output = tf.placeholder(tf.int32, [None])
# Create embedding
embedding_mat = tf.Variable(tf.random_uniform([vocab_size, embedding_size], -1.0, 1.0))
embedding_output = tf.nn.embedding_lookup(embedding_mat, x_data)
# Define the RNN cell
# tensorflow change >= 1.0, rnn is put into tensorflow.contrib directory. Prior version not test.
if tf.__version__[0] >= '1':
cell = tf.contrib.rnn.BasicRNNCell(num_units=rnn_size)
else:
cell = tf.nn.rnn_cell.BasicRNNCell(num_units=rnn_size)
output, state = tf.nn.dynamic_rnn(cell, embedding_output, dtype=tf.float32)
output = tf.nn.dropout(output, dropout_keep_prob)
# Get output of RNN sequence
output = tf.transpose(output, [1, 0, 2])
last = tf.gather(output, int(output.get_shape()[0]) - 1)
weight = tf.Variable(tf.truncated_normal([rnn_size, 2], stddev=0.1))
bias = tf.Variable(tf.constant(0.1, shape=[2]))
logits_out = tf.matmul(last, weight) + bias
```
Next we declare the loss function (softmax cross entropy), an accuracy function, and optimization function (RMSProp).
```
# Loss function
losses = tf.nn.sparse_softmax_cross_entropy_with_logits(logits=logits_out, labels=y_output)
loss = tf.reduce_mean(losses)
accuracy = tf.reduce_mean(tf.cast(tf.equal(tf.argmax(logits_out, 1), tf.cast(y_output, tf.int64)), tf.float32))
optimizer = tf.train.RMSPropOptimizer(learning_rate)
train_step = optimizer.minimize(loss)
```
> You may ignore the warning, as the texts are small and our batch size is only 100. If you increase the batch size and/or have longer sequences of texts, this model may consume too much memory.
Next we initialize the variables in the computational graph.
```
init = tf.global_variables_initializer()
sess.run(init)
train_loss = []
test_loss = []
train_accuracy = []
test_accuracy = []
# Start training
for epoch in range(epochs):
# Shuffle training data
shuffled_ix = np.random.permutation(np.arange(len(x_train)))
x_train = x_train[shuffled_ix]
y_train = y_train[shuffled_ix]
num_batches = int(len(x_train)/batch_size) + 1
# TO DO CALCULATE GENERATIONS ExACTLY
for i in range(num_batches):
# Select train data
min_ix = i * batch_size
max_ix = np.min([len(x_train), ((i+1) * batch_size)])
x_train_batch = x_train[min_ix:max_ix]
y_train_batch = y_train[min_ix:max_ix]
# Run train step
train_dict = {x_data: x_train_batch, y_output: y_train_batch, dropout_keep_prob:0.5}
sess.run(train_step, feed_dict=train_dict)
# Run loss and accuracy for training
temp_train_loss, temp_train_acc = sess.run([loss, accuracy], feed_dict=train_dict)
train_loss.append(temp_train_loss)
train_accuracy.append(temp_train_acc)
# Run Eval Step
test_dict = {x_data: x_test, y_output: y_test, dropout_keep_prob:1.0}
temp_test_loss, temp_test_acc = sess.run([loss, accuracy], feed_dict=test_dict)
test_loss.append(temp_test_loss)
test_accuracy.append(temp_test_acc)
print('Epoch: {}, Test Loss: {:.2}, Test Acc: {:.2}'.format(epoch+1, temp_test_loss, temp_test_acc))
```
Here is matplotlib code to plot the loss and accuracy over the training generations for both the train and test sets.
```
%matplotlib inline
# Plot loss over time
epoch_seq = np.arange(1, epochs+1)
plt.plot(epoch_seq, train_loss, 'k--', label='Train Set')
plt.plot(epoch_seq, test_loss, 'r-', label='Test Set')
plt.title('Softmax Loss')
plt.xlabel('Epochs')
plt.ylabel('Softmax Loss')
plt.legend(loc='upper left')
plt.show()
# Plot accuracy over time
plt.plot(epoch_seq, train_accuracy, 'k--', label='Train Set')
plt.plot(epoch_seq, test_accuracy, 'r-', label='Test Set')
plt.title('Test Accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend(loc='lower right')
plt.show()
```
| github_jupyter |
# Classificador de Raças de Cachorros usando Tensorflow e Keras
Neste notebook iremos implementadar um modelo para classificação de imagens. Classificação é uma das "tarefas" em que podemos utilizar Machine Learning, nesta tarefa o ensino é **supervisionado**, em outras palavras nós vamos ensinar ao modelo através de exemplos com gabarito.
Nosso modelo deverá receber imagens de veículos e não-veículos e identificar a que **classe** (raça de cachorro) o cachorro pertence.
## Dados
Os dados são oriundos da competição [Dog Breed Indentification do Kaggle](https://www.kaggle.com/c/dog-breed-identification), na qual fornece aproximadamente 10 mil imagens de cachorros de 120 classes.
## Modelo
Iremos utilizar a [arquitetura da InceptionV3](https://arxiv.org/abs/1512.00567), ela está implementada no [Keras](https://keras.io/applications/#inceptionv3)
## Conseguindo os dados
Para ter acesso aos dados é necessário uma conta no Kaggle, e ter que entrar na [competição](https://www.kaggle.com/c/dog-breed-identification), e ir na aba Data na competição a baixá-los
### Avisos
#### Aviso #1
Para fazer o treinamento da InceptionV3 é necessário um grande poder computacional, na qual a maioria das pessoas não possuem. Mas não será por isso que não utilizaremos a Inception, graças ao Kaggle, temos a opção de rodar Kernels (que são muito similares aos notebooks do jupyter) na infraestrutura do próprio Kaggle, para mais informações sobre o suporte a GPU's do Kaggle veja [esse notebook](https://www.kaggle.com/dansbecker/running-kaggle-kernels-with-a-gpu) do [Dan Becker](https://twitter.com/dan_s_becker)
#### Aviso #2
Esse notebook não foi executado na minha máquina, eu rodei ele nos kernels do Kaggle. Por isso não temos as saídas das células, se você quiser visualizar as saídas clique [aqui](https://www.kaggle.com/igorslima/inception)
```
import pandas as pd
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
np.random.seed(0)
input_folder = '/kaggle/input'
# lendo input
df_train = pd.read_csv(input_folder+'/labels.csv')
df_test = pd.read_csv(input_folder+'/sample_submission.csv')
df_train.breed.value_counts().plot(kind='bar', figsize=(15,15), title="Quantidade de imagens por raça no treino");
df_train.head()
df_test.head()
```
## Transormando os dados para a "notação" one-hot-encoding
Para mais informações sobre o One Hot Enconding leia este [post](https://hackernoon.com/what-is-one-hot-encoding-why-and-when-do-you-have-to-use-it-e3c6186d008f)
```
targets_series = pd.Series(df_train['breed'])
one_hot = pd.get_dummies(targets_series, sparse = True)
one_hot_labels = np.asarray(one_hot)
im_size = 224
```
## Lendo as imagens
Para treinar a rede é necessário peger as imagens do disco e colocar elas em memória. Não entendeu um 'a' do que eu disse? Tudo bem, é normal. O que eu quis dizer foi que vamos ter que pegar as imagens do HD e colocar elas na memória RAM.
```
from tqdm import tqdm # bliblioteca para colocar a porcentagem de andamento do for
import cv2 # biblioteca para visão computacional
x_train = []
y_train = []
x_test = []
i = 0
for f, breed in tqdm(df_train.values):
img = cv2.imread(input_folder+'/train/{}.jpg'.format(f))
x_train.append(cv2.resize(img, (im_size, im_size)))
label = one_hot_labels[i]
y_train.append(label)
i += 1
del df_train # apagando uma variável pra diminuir consumo de memória
for f in tqdm(df_test['id'].values):
img = cv2.imread(input_folder+'/test/{}.jpg'.format(f))
x_test.append(cv2.resize(img, (im_size, im_size)))
```
## Dividindo dataset
Geralmente em dividimos os dados em treino, validação e teste.
1. Treino: conjunto para treinar o modelo
2. Validação: conjunto para escolher os melhores hiperparâmetros do modelo (mais tarde falo sobre hiperparâmetros, ok?)
3. Teste: conjunto para coletar as métricas finais do modelo
```
from sklearn.model_selection import train_test_split # biblioteca para fazer a divisão dos dados em treino e teste
num_class = 120
X_train, X_valid, Y_train, Y_valid = train_test_split(x_train, y_train, shuffle=True, test_size=0.2, random_state=1)
```
## Data augmentation
Nós temos dados o suficiente para travar nossas máquinas XD, mas não o suficiente para treinar modelos bastantes robustos, temos poucas imagens por classe.
Para ameninzar esse problema iremos utilizar uma técnica chamada data augmentations, ela transforma uma imagem em diversas, como por exemplo dar um giro vertical, ou horizontal. Como nesse exemplo:

Links legais (em inglês, desculpem):
[Link para a documentação](https://keras.io/preprocessing/image/)
[Tutorial massa do keras](https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html)
[Outro tutorial massa, mas não é do Keras, esse é do Dan Becker](https://www.kaggle.com/dansbecker/data-augmentation)[](http://)
```
from keras.preprocessing.image import ImageDataGenerator # biblioteca para data augmetantaion
datagen = ImageDataGenerator(width_shift_range=0.2,
height_shift_range=0.2,
zoom_range=0.2,
rotation_range=30,
vertical_flip=False,
horizontal_flip=True) # aqui eu defino os parâmetros que irei
# utilizar para gerar as imagens
train_generator = datagen.flow(np.array(X_train), np.array(Y_train),
batch_size=32)
valid_generator = datagen.flow(np.array(X_valid), np.array(Y_valid),
batch_size=32)
```
## Criação da Inception
A partir de agora iremos criar a rede propriamente dita, iremos utilizar a arquitetura da rede Inception, e os pesos pré-treinada sobre os dados do ImageNet.
```
from keras.applications.inception_v3 import InceptionV3
from keras.layers import Dense, Dropout, Flatten
from keras import regularizers
from keras.models import Model
base_model = InceptionV3(weights="imagenet",include_top=False, input_shape=(im_size, im_size, 3))
dropout = base_model.output
dropout = Dropout(0.5)(dropout)
model_with_dropout = Model(inputs=base_model.input, outputs=dropout)
x = model_with_dropout.output
x = Flatten()(x)
predictions = Dense(num_class, activation='softmax',
kernel_regularizer=regularizers.l2(0.0015),
activity_regularizer=regularizers.l1(0.0015))(x)
my_model = Model(inputs=model_with_dropout.input, outputs=predictions)
my_model.compile(optimizer='sgd',
loss='categorical_crossentropy',
metrics=['accuracy'])
```
## Treinando o modelo
```
my_model.fit_generator(
train_generator,
epochs=10, steps_per_epoch=len(X_train) / 18,
validation_data=valid_generator, validation_steps=len(X_valid) / 18) # reali
```
## Fazendo predições
```
preds = my_model.predict(np.array(x_test), verbose=1)
sub = pd.DataFrame(preds)
col_names = one_hot.columns.values
sub.columns = col_names
sub.insert(0, 'id', df_test['id'])
sub.head(5)
sub.to_csv("submission.csv")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/graviraja/100-Days-of-NLP/blob/applications%2Fgeneration/applications/generation/utterance_generation/Basic%20Utterance%20Generation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
TASK_DATA_DIR = 'glue_data/QQP'
!test -d glue_data || git clone https://gist.github.com/60c2bdb54d156a41194446737ce03e2e.git glue_data
!test -d $TASK_DATA_DIR || python glue_data/download_glue_data.py --data_dir glue_data --tasks=QQP
!ls -alh $TASK_DATA_DIR
import os
import time
import math
import random
import pandas as pd
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
from torchtext import data, vocab
import matplotlib.pyplot as plt
import seaborn as sns
SEED = 42
np.random.seed(SEED)
torch.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
device
train_df = pd.read_csv(TASK_DATA_DIR + '/train.tsv', sep='\t', error_bad_lines=False)
valid_df = pd.read_csv(TASK_DATA_DIR + '/dev.tsv', sep='\t', error_bad_lines=False)
train_df.head()
len(train_df), len(valid_df)
sns.countplot(train_df['is_duplicate'])
plt.xlabel('Train data distribution')
sns.countplot(valid_df['is_duplicate'])
plt.xlabel('Valid data distribution')
train_data = train_df[train_df['is_duplicate'] == 1]
valid_data = valid_df[valid_df['is_duplicate'] == 1]
train_data.head()
len(train_data), len(valid_data)
train_data = train_data[['question1', 'question2']]
valid_data = valid_data[['question1', 'question2']]
train_data.head()
sample_train_data = train_data.sample(50000)
sample_valid_data = valid_data.sample(5000)
sample_train_data.to_csv('train_ds.csv')
sample_valid_data.to_csv('valid_ds.csv')
!ls -lah
tokenizer = data.get_tokenizer('spacy')
TEXT = data.Field(tokenize=tokenizer, lower=True, init_token='<sos>', eos_token='<eos>')
fields = [(None, None), ("source", TEXT), ("target", TEXT)]
train_dataset, valid_dataset = data.TabularDataset.splits(path='.',
train='train_ds.csv', validation='valid_ds.csv',
format='csv', skip_header=True, fields=fields)
print(f"Number of training examples: {len(train_dataset)}")
print(f"Number of validation examples: {len(valid_dataset)}")
print(vars(train_dataset.examples[1]))
TEXT.build_vocab(train_dataset, min_freq=5)
print(f"Number of tokens in vocabulary: {len(TEXT.vocab)}")
BATCH_SIZE = 64
train_iterator, valid_iterator = data.BucketIterator.splits(
(train_dataset, valid_dataset),
batch_size=BATCH_SIZE,
sort_key=lambda x: len(x.source),
device=device
)
# sample checking
temp = next(iter(train_iterator))
temp.source.shape, temp.target.shape
class Encoder(nn.Module):
def __init__(self, input_dim, emb_dim, hidden_dim, n_layers, dropout):
super().__init__()
self.n_layers = n_layers
self.hidden_dim = hidden_dim
self.embedding = nn.Embedding(input_dim, emb_dim)
self.rnn = nn.LSTM(emb_dim, hidden_dim, bidirectional=True, num_layers=n_layers, dropout=dropout)
self.fc = nn.Linear(hidden_dim * 2, hidden_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, src):
# src => [seq_len, batch_size]
embedded = self.dropout(self.embedding(src))
# embedded => [seq_len, batch_size, hidden_dim]
outputs, (hidden, cell) = self.rnn(embedded)
# outputs => [seq_len, batch_size, hidden_dim * 2]
# hidden, cell => [num_layers * num_dir, batch_size, hidden_dim]
hidden = hidden.view(self.n_layers, 2, -1, self.hidden_dim)
cell = cell.view(self.n_layers, 2, -1, self.hidden_dim)
# hidden, cell => [num_layers, num_dir, batch_size, hidden_dim]
final_forward_hidden = hidden[:, 0, :, :]
final_backward_hidden = hidden[:, 1, :, :]
# final_hiddens => [num_layers, batch_size, hidden_dim]
final_forward_cell = cell[:, 0, :, :]
final_backward_cell = cell[:, 1, :, :]
# final_cells => [num_layers, batch_size, hidden_dim]
combined_hidden = torch.cat((final_forward_hidden, final_backward_hidden), dim=2)
combined_cell = torch.cat((final_forward_cell, final_backward_cell), dim=2)
# combined_hidden, combined_cell => [num_layers, batch_size, hidden_dim * 2]
decoder_initial_hidden = self.fc(combined_hidden)
decoder_initial_cell = self.fc(combined_cell)
# decoder_initial_states => [num_layers, batch_size, hidden_dim]
return decoder_initial_hidden, decoder_initial_cell
class Decoder(nn.Module):
def __init__(self, input_dim, emb_dim, hidden_dim, n_layers, dropout):
super().__init__()
self.n_layers = n_layers
self.hidden_dim = hidden_dim
self.input_dim = input_dim
self.rnn = nn.LSTM(emb_dim, hidden_dim, num_layers=n_layers, dropout=dropout)
self.fc = nn.Linear(hidden_dim, input_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, input, hidden, cell):
# input => [seq_len, batch_size, emb_dim]
# => [1, batch_size, emb_dim]
# hidden => [num_layers, batch_size, hidden_dim]
# cell => [num_layers, batch_size, hidden_dim]
output, (hidden, cell) = self.rnn(input, (hidden, cell))
# output => [1, batch_size, hidden_dim]
# hidden => [num_layers, batch_size, hidden_dim]
# cell => [num_layers, batch_size, hidden_dim]
logits = self.fc(self.dropout(output.squeeze(0)))
# logits => [batch_size, output_dim]
return logits, hidden, cell
class Seq2Seq(nn.Module):
def __init__(self, encoder, decoder, device):
super().__init__()
self.encoder = encoder
self.decoder = decoder
self.device = device
def forward(self, src, trg, teacher_forcing_ratio=0.5):
# src => [seq_len, batch_size]
# trg => [trg_len, batch_size]
batch_size = src.shape[1]
trg_len = trg.shape[0]
trg_vocab_size = self.decoder.input_dim
# outputs: to store the predictions of the decoder
outputs = torch.zeros(trg_len, batch_size, trg_vocab_size).to(self.device)
hidden, cell = self.encoder(src)
# hidden, cell => [num_layers, batch_size, hidden_dim]
dec_inp = trg[0, :]
for t in range(1, trg_len):
dec_inp_emb = self.encoder.embedding(dec_inp.unsqueeze(0))
# dec_inp_emb => [1, batch_size, emb_dim]
output, hidden, cell = self.decoder(dec_inp_emb, hidden, cell)
# save the output
outputs[t] = output
# to decide whether to use teacher force or not
teacher_force = random.random() < teacher_forcing_ratio
top1 = output.argmax(1)
dec_inp = trg[t] if teacher_force else top1
return outputs
INPUT_DIM = len(TEXT.vocab)
EMB_DIM = 256
HID_DIM = 512
N_LAYERS = 2
DROPOUT = 0.5
enc = Encoder(INPUT_DIM, EMB_DIM, HID_DIM, N_LAYERS, DROPOUT)
dec = Decoder(INPUT_DIM, EMB_DIM, HID_DIM, N_LAYERS, DROPOUT)
model = Seq2Seq(enc, dec, device).to(device)
def init_weights(model):
for name, param in model.named_parameters():
nn.init.uniform_(param.data, -0.08, 0.08)
model.apply(init_weights)
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model)} trainable parameters')
PAD_IDX = TEXT.vocab.stoi[TEXT.pad_token]
optimizer = optim.Adam(model.parameters())
criterion = nn.CrossEntropyLoss(ignore_index=PAD_IDX)
def train(model, iterator, criterion, optimizer, clip):
epoch_loss = 0
# keep the model in train mode
model.train()
# iterate over train data
for i, batch in enumerate(iterator):
src = batch.source
trg = batch.target
# src => [seq_len, batch_size]
# trg => [seq_len, batch_size]
# zero the gradients
optimizer.zero_grad()
# forward pass
output = model(src, trg)
# reshaping the output to make it compatible to cal. loss
# can also do without reshaping
output_dim = output.shape[-1]
output = output[1:].view(-1, output_dim)
trg = trg[1:].view(-1)
loss = criterion(output, trg)
# backward pass
loss.backward()
# gradient clipping
torch.nn.utils.clip_grad_norm_(model.parameters(), clip)
# update the parameters of the model
optimizer.step()
# update the loss
epoch_loss += loss.item()
return epoch_loss / len(iterator)
def evaluate(model, iterator, criterion):
epoch_loss = 0
# keep the model in eval mode
model.eval()
# do not calculate gradients
with torch.no_grad():
# iterate over the data
for batch in iterator:
src = batch.source
trg = batch.target
# src => [seq_len, batch_size]
# trg => [seq_len, batch_size]
# forward pass
# make sure the teacher_forcing_ratio is 0 in eval
output = model(src, trg, 0)
# reshaping for loss calculation
output_dim = output.shape[-1]
output = output[1:].view(-1, output_dim)
trg = trg[1:].view(-1)
# loss
loss = criterion(output, trg)
# update loss
epoch_loss += loss.item()
return epoch_loss / len(iterator)
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = elapsed_time - (elapsed_mins * 60)
return elapsed_mins, elapsed_secs
N_EPOCHS = 10
CLIP = 1
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss = train(model, train_iterator, criterion, optimizer, CLIP)
valid_loss = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'model.pt')
print(f"Epoch {epoch + 1} | Time: {epoch_mins}m {epoch_secs}s")
print(f"\tTrain Loss: {train_loss:.3f} | Train PPL: {math.exp(train_loss):7.3f} |")
print(f"\tValid Loss: {valid_loss:.3f} | Valid PPL: {math.exp(valid_loss):7.3f} |")
```
| github_jupyter |
### BCO-DMO Knowledge Graph Data Exploration Prototype
This is a prototype demonstrating how python can be used to interactively explore oceanographic data within the BCO-DMO Knowledge Graph. This demonstration was developed for SciPy 2020.
**WARNING** This is just a prototype and will likely be updated (or abandoned ¯\\_(ツ)_/¯). In addition, the BCO-DMO Knowledge Graph is also under construction, so stability of this prototype is far from guaranteed. This is mainly just an example of how we might leverage the Knowledge Graph to facilitate interactive exploration of oceanographic data. Hope to have some amazing (and stable) tools for exploration of the Graph in the future. Brutal honesty moment: This tool which was just intended for a viz example has been great for revealing some issues with tagging of data in the Graph -- side bonus for us at BCO-DMO so we can fix these, but this does result in some issues visualizing some datasets.
**WARNING \#2:** Some of the datasets within BCO-DMO are very large. Therefore, for performance reasons, a limit on datasets displayed is set below. Feel free to change.
```
MAX_DATASET_SHOW = 5
from bqplot import Lines, Figure, LinearScale, DateScale, Axis
from ipyleaflet import Map, GeoJSON, basemaps, WidgetControl, Marker, MarkerCluster
from ipywidgets import link, HTML
import json
import os
import sys
import requests
import geopandas
import pandas as pd
import numpy as np
import rdflib
from SPARQLWrapper import SPARQLWrapper, JSON
from ipywidgets import Layout, IntText, Dropdown, Combobox, VBox, IntSlider
#credit: Doug Fils
def get_sparql_dataframe(service, query):
"""
Helper function to convert SPARQL results into a Pandas data frame.
"""
sparql = SPARQLWrapper(service)
sparql.setQuery(query)
sparql.setReturnFormat(JSON)
result = sparql.query()
processed_results = json.load(result.response)
cols = processed_results['head']['vars']
out = []
for row in processed_results['results']['bindings']:
item = []
for c in cols:
#item.append(str(row.get(c, {}).get('value')))
item.append(row.get(c, {}).get('value'))
out.append(item)
# could simply return 'out' which is a list of lists
return pd.DataFrame(out, columns=cols)
BCODMO_SERVE = "https://lod.bco-dmo.org/sparql" #BCO-DMO SPARQL Endpoint
BCODMO_PREF = "http://lod.bco-dmo.org/id/" #BCO-DMO URI prefix
## Paramter options
###### Dataset Description Widget
#SPARQL query for BCO-DMO dataset information
masterParameterQuery = """
SELECT DISTINCT ?masterParamtersId ?shortDesc ?label #?datasetParameter #?url
WHERE {
<http://lod.bco-dmo.org/id/parameters> ?property ?masterParamtersId .
?masterParamtersId owl:deprecated "0"^^xsd:boolean . #remove deprecated master parameters
?masterParamtersId odo:hasParameterShortDescription ?shortDesc .
?masterParamtersId skos:prefLabel ?label .
?datasetParameter odo:isInstanceOf ?masterParametersId .
?dataset odo:storesValuesFor ?datasetParameter .
#Select only those master params that have depths
?datasetParameter odo:isInstanceOf <http://lod.bco-dmo.org/id/parameter/808> .
#end
?affordance schema:subjectOf ?dataset .
?affordance rdf:type ?action_type .
?affordance schema:target ?target .
?target schema:contentType "application/geo+json"^^xsd:token .
#?target schema:url ?url .
}
ORDER BY ?shortDesc ?label ?masterParametersId
"""
df_masterParams_with_geoJson = get_sparql_dataframe(BCODMO_SERVE, masterParameterQuery)
#Displays dropdown selector for dataset parameters
masterParamsOptions = df_masterParams_with_geoJson["label"].values.tolist()
style = {'description_width': 'initial'}
masterParamsMenu = Combobox(
options=masterParamsOptions,
description='Search Parameter:',
disabled=False,
layout=Layout(width='80%'),
continuous_update=False,
style=style
)
###### Update Dataset Parameter Description with Menu Selection
def handle_masterParam_change(change):
if change.new != change.old:
masterParamsMenu.value = change.new
masterParamsMenu.observe(handle_masterParam_change, names='value') #observer for change
masterParamsMenu
parameterSelected = df_masterParams_with_geoJson["masterParamtersId"].loc[df_masterParams_with_geoJson["label"]\
== masterParamsMenu.value]
parameterSelected = parameterSelected.to_string(index=False).strip()
###Select Nitrite and see what happens - http://lod.bco-dmo.org/id/parameter/1192
nitriteQuery = """
SELECT DISTINCT ?masterParam ?nanValue ?unit ?parameterName ?datasetID ?url
WHERE {
VALUES ?masterParam {<""" + parameterSelected + """>}
?dataset_parameter odo:isInstanceOf ?masterParam .
?dataset_parameter odo:hasNoDataValue ?nanValue .
?dataset_parameter odo:hasUnitOfMeasure ?nodeUnit .
?nodeUnit rdf:value ?unit .
?dataset_parameter skos:prefLabel ?parameterName .
?dataset odo:storesValuesFor ?dataset_parameter .
#?dataset_parameter odo:isInstanceOf <http://lod.bco-dmo.org/id/parameter/808> .
?dataset dcterms:identifier ?datasetID .
#check GeoJSON
?affordance schema:subjectOf ?dataset .
?affordance rdf:type ?action_type .
?affordance schema:target ?target .
?target schema:contentType "text/csv"^^xsd:token .
?target schema:url ?url .
}
"""
df_parameter = get_sparql_dataframe(BCODMO_SERVE, nitriteQuery)
#df_parameter
# All the datasets that have the target parameter
listDataSetIDs = df_parameter["datasetID"].astype("str").values.tolist()
listDataSetIDsStr = ' '.join(listDataSetIDs)
nitriteDepthQuery = """
SELECT DISTINCT ?masterParamDepth ?nanValueDepth ?unitDepth ?col_nameDepth ?datasetID ?url
WHERE {
VALUES ?datasetID {""" + listDataSetIDsStr + """}
VALUES ?masterParamDepth { <http://lod.bco-dmo.org/id/parameter/808>} #808 is the parameter for Depth
?dataset_parameter odo:isInstanceOf ?masterParamDepth .
?dataset_parameter odo:hasNoDataValue ?nanValueDepth .
?dataset_parameter odo:hasUnitOfMeasure ?nodeUnit .
?nodeUnit rdf:value ?unitDepth .
?dataset_parameter skos:prefLabel ?col_nameDepth .
?dataset odo:storesValuesFor ?dataset_parameter .
#?dataset_parameter odo:isInstanceOf <http://lod.bco-dmo.org/id/parameter/808> .
?dataset dcterms:identifier ?datasetID .
#check GeoJSON
?affordance schema:subjectOf ?dataset .
?affordance rdf:type ?action_type .
?affordance schema:target ?target .
?target schema:contentType "text/csv"^^xsd:token .
?target schema:url ?url .
}
"""
df_parameterDepth = get_sparql_dataframe(BCODMO_SERVE, nitriteDepthQuery)
#parameterDepth_df
#df_parameterDepth.style.set_properties(subset=['url'], **{'width': '600px'})
#Find all datasets that have the target parameter and associated depth data
df_dataSetsWithParameterAndDepth = df_parameter.loc[df_parameter["datasetID"].isin(df_parameterDepth["datasetID"].unique())].reset_index()
df_dataSetsWithParameterAndDepth.drop_duplicates(subset="url", inplace=True)
urlTest = df_dataSetsWithParameterAndDepth[["url", "datasetID"]].values[0:MAX_DATASET_SHOW] #limiting to 5 datasets max right now
#create groupbys on specific datasets
df_parameterDepth["parameterType"] = "depth"
try:
df_dataSetsWithParameterAndDepth = df_dataSetsWithParameterAndDepth.drop(columns=["index"])
except:
print("no column named index")
df_dataSetsWithParameterAndDepth["parameterType"] = str(masterParamsMenu.value)
df_parameterDepth = df_parameterDepth.rename(columns={"masterParamDepth":"masterParam", \
"col_nameDepth":"parameterName", \
"unitDepth":"unit", \
"nanValueDepth":"nanValue"})
df_paramsAndDepths = pd.concat([df_parameterDepth, df_dataSetsWithParameterAndDepth])
gb_paramDepth = df_paramsAndDepths.groupby("datasetID")
dfg_big = geopandas.GeoDataFrame(columns=['datasetID', 'geometry', 'parameterName', 'value', 'parameterType', 'unit'])
dfg_points = geopandas.GeoDataFrame(columns=["datasetID", "geometry"])
for url, datasetID in urlTest:
#generate lists of parameters and depth dataset-specific column names
subdf = gb_paramDepth.get_group(datasetID)
paramCols = subdf["parameterName"].loc[subdf["parameterType"] == masterParamsMenu.value].unique().tolist()
depthCols = subdf["parameterName"].loc[subdf["parameterType"] == "depth"].unique().tolist()
depthCols = depthCols + ["depth"]
#Adding some common nan issues -- need to update KG where there are multiple NaNs/dataset parameter
nanValues = subdf["nanValue"].unique().tolist() + ["n.a.", "nan", "-9999", "-999.0", "-999", "mix", ""]# \
# "bdl", 'Below_detection_limit', 'ND', 'DNP', 'BDL']# coerce all these fun strings in float cols
df_units = subdf[["unit", "parameterName"]]
data = pd.read_csv(url, low_memory=False)
data = data.drop([0])
colsKeep = ["latitude", "longitude"] + depthCols + paramCols
checkParamInFile = all(item in data.columns for item in colsKeep)
if checkParamInFile is False:
colsKeep = [s for s in colsKeep if s in data.columns]
paramCols = [s for s in paramCols if s in data.columns]
depthCols = [s for s in depthCols if s in data.columns]
#Drop dataset if latitude or longitude don't exist
checkCoords = all(item in data.columns for item in ["latitude", "longitude"])
if checkCoords is False:
continue
dfg = data[colsKeep].copy()
dfg["datasetID"] = datasetID
[dfg.replace(x, np.nan, inplace=True) for x in nanValues]
#add unique subset of location points to points dataframe for mapping
dfg["longitude"] = dfg["longitude"].astype("float")
dfg["latitude"] = dfg["latitude"].astype("float")
dfg_geometry = geopandas.GeoDataFrame(dfg, geometry=geopandas.points_from_xy(dfg["longitude"], dfg["latitude"]))
#Drop rows that don't have data for the selected parameter
dfg_geometry[paramCols] = dfg_geometry[paramCols].fillna(1).apply(lambda x: pd.to_numeric(x, errors='coerce'))
dfg_geometry = dfg_geometry.dropna(subset=paramCols)
dfg_points = dfg_points.append(dfg_geometry[["datasetID", "geometry"]].drop_duplicates())
paramsList = paramCols + depthCols
dfg_melt = pd.melt(dfg_geometry, id_vars=["datasetID", "geometry"], \
value_vars=[c for c in dfg_geometry.columns if c in paramsList],\
var_name='parameterName')
dfg_melt["parameterType"] = "depth"
dfg_melt.loc[dfg_melt["parameterName"].isin(paramCols), ['parameterType']] = 'parameter'
dfg_melt = pd.merge(dfg_melt, df_units, how="left")
dfg_big = dfg_big.append(dfg_melt)
#convert dfg_points to geoJson
dfg_points = dfg_points.reset_index().drop(columns='index')
point_geoJson = dfg_points.to_file("points.geojson", driver='GeoJSON')
with open('points.geojson', 'r') as f:
data = json.load(f)
dfp = pd.DataFrame(dfg_big) #convert to pandas dataframe to do more
dfp["geometry_str"] = dfp["geometry"].astype("str").str.replace("POINT ", "").str.replace("(", "").str.replace(")", "")
dfp[['longitude','latitude']] = dfp["geometry_str"].str.split(expand=True)
dfp["value"] = dfp["value"].astype("float").round(2)
dfp["value"].loc[dfp["parameterName"] == masterParamsMenu.value].astype("float").round(0)
dfp[['longitude','latitude']] = dfp[['longitude','latitude']].astype("float")
#dfp[['longitude','latitude']] = dfp[['longitude','latitude']].round(5)
dfp["lon_lat"] = dfp["latitude"].astype("str") + " " + dfp["longitude"].astype("str")
gb_dfp = dfp.groupby(["datasetID"])
m = Map(center=(0, 0), zoom=1, basemap=basemaps.Esri.NatGeoWorldMap)
geo_json = GeoJSON(data=data, style = {
})
m.add_layer(geo_json)
html1 = HTML('''
<h4>Dataset Info</h4>
Click on a point
''')
html1.layout.margin = '0px 20px 20px 20px'
control1 = WidgetControl(widget=html1, position='bottomleft')
def update_html(feature, **kwargs):
html1.value = '''
<b>Dataset: {}</b></br>
<a>https://www.bco-dmo.org/dataset/{}</a>
'''.format(feature['properties']['datasetID'], feature['properties']['datasetID'])
geo_json.on_click(update_html)
#add minimap
#minimap = Map(
# zoom_control=True, attribution_control=False,
# zoom=-2, center=m.center, basemap=basemaps.Esri.WorldImagery
#)
#minimap.layout.width = '250px'
#minimap.layout.height = '200px'
#### Changed the datatype of dfg_points, so would need to update this in order for it to work
#minimap.add_layer(MarkerCluster(markers=[Marker(location=geolocation.coords[0][::-1]) \
# for geolocation in dfg_points.geometry.unique()]))
#link((minimap, 'center'), (m, 'center'))
#minimap_control = WidgetControl(widget=minimap, position='bottomleft')
#m.add_control(minimap_control)
m.add_control(control1)
#m
x_data = []
depth_data = []
x_sc = LinearScale()
y_sc = LinearScale(reverse=True)
line = Lines(x=x_data,
y=depth_data,
scales={'x': x_sc, 'y': y_sc},
colors=['orange', 'red', 'blue', 'black'])
ax_x = Axis(label="", scale=x_sc, tick_format='0.1f', num_ticks=5)
ax_y = Axis(label="", scale=y_sc,
orientation='vertical', tick_format='0.0f', side='left')
figure = Figure(axes=[ax_x, ax_y], marks=[line], animation_duration=300,
layout={'max_height': '270px'}, title=masterParamsMenu.value)
#Make the Widgets for selecting parameters
paramOptions = [""]
style = {'description_width': 'initial'}
paramPlotMenu = Dropdown(
options=paramOptions,
description='Parameter to plot:',
disabled=False,
layout=Layout(width='80%'),
continuous_update=False,
value="",
style=style
)
depthOptions = [""]
style = {'description_width': 'initial'}
depthPlotMenu = Dropdown(
options=depthOptions,
description='Depth to plot:',
disabled=False,
layout=Layout(width='80%'),
continuous_update=False,
value="",
style=style
)
figureDisplay = VBox([figure, paramPlotMenu, depthPlotMenu])
#figureDisplay
def update_figure(datasetID, dfp_paramType, gb_dfp_sub_point):
paramPlotMenu.options = dfp_paramType["parameterName"].loc[dfp_paramType["parameterType"] == "parameter"].unique()
depthPlotMenu.options = dfp_paramType["parameterName"].loc[dfp_paramType["parameterType"] == "depth"].unique()
def paramSelect_changed(change):
if change.new != change.old:
paramPlotMenu.value = change.new
def depthSelect_changed(change):
if change.new != change.old:
depthPlotMenu.value = change.new
paramPlotMenu.observe(paramSelect_changed, names='value')
depthPlotMenu.observe(depthSelect_changed, names='value')
parameter = gb_dfp_sub_point[paramPlotMenu.value].dropna().values
depth = gb_dfp_sub_point[depthPlotMenu.value].dropna().values
if len(parameter) == len(depth): #Need at least 2 points for a depth profile
line.x = parameter
line.y = depth
figure.title = paramPlotMenu.value
ax_x.label = dfp_paramType["unit"].loc[dfp_paramType["parameterName"] == paramPlotMenu.value].to_string(index=False)
ax_y.label = dfp_paramType["unit"].loc[dfp_paramType["parameterName"] == depthPlotMenu.value].to_string(index=False)
else:
figure.title = "Incompatible parameter & depth"
line.x = [0]
line.y = [0]
widget_control1 = WidgetControl(widget=figureDisplay, position='topright')
m.add_control(widget_control1)
def plot_on_click(event, feature, **kwargs):
global datasetID, dfp_paramType, gb_dfp_sub_point
coordsList = feature["geometry"]["coordinates"]
datasetID = feature["properties"]["datasetID"]
#point = str('%.5f' % coordsList[1]) + " " + str('%.5f' % coordsList[0])
point = str(coordsList[1]) + " " + str(coordsList[0])
dfp_sub = gb_dfp.get_group(str(datasetID))
dfp_paramType = dfp_sub[["parameterType", "parameterName", "unit"]].drop_duplicates()
dfp_sub = dfp_sub.drop(columns=["unit", "parameterType", "geometry", "datasetID", \
"longitude", "latitude"])
#print(dfp_sub)
gb_dfp_sub = dfp_sub.groupby(["lon_lat"])
#print(gb_dfp_sub.groups)
gb_dfp_sub_point = gb_dfp_sub.get_group(point).reset_index()
gb_dfp_sub_point = gb_dfp_sub_point.drop(columns=["lon_lat"])
gb_dfp_sub_point = gb_dfp_sub_point.pivot(index=None, columns='parameterName', values=["value"])
gb_dfp_sub_point.columns = gb_dfp_sub_point.columns.droplevel(level=0)
update_figure(datasetID, dfp_paramType, gb_dfp_sub_point)#add back point
geo_json.on_click(plot_on_click)
m
```
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# Initial Data for Solving Maxwell's Equations in Flat Spacetime
## Authors: Terrence Pierre Jacques, Zachariah Etienne and Ian Ruchlin
## This module constructs the initial data for Maxwell's equations as symbolic (SymPy) expressions, for a purely toriodal dipole field, as defined in [Knapp, Walker & Baumgarte (2002)](https://arxiv.org/abs/gr-qc/0201051).
**Notebook Status:** <font color='green'><b> Validated </b></font>
**Validation Notes:** All expressions generated in this module have been validated, against the [Dendro code Maxwell initial data](https://github.com/paralab/Dendro-GR), and have satisfied the contraints given in [Tutorial-VacuumMaxwell_Curvilinear_RHS-Rescaling](Tutorial-VacuumMaxwell_Curvilinear_RHS-Rescaling.ipynb), as well as the wave equation for the electric field and the vector potential.
### NRPy+ Source Code for this module: [Maxwell/InitialData.py](../edit/Maxwell/InitialData.py), [reference_metric.py](../edit/reference_metric.py)
[comment]: <> (Introduction: TODO)
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
1. [Step 1](#initializenrpy): Initialize needed Python/NRPy+ modules and set destination basis
1. [Step 2](#step2): A Purely Toriodal Dipole Field
1. [Step 2.a](#cart_basis): Converting to Cartesian Basis
1. [Step 2.b](#dst_basis): Converting to Destination Basis
1. [Step 3](#step3): Checks
1. [Step 3.a](#lorentz): Lorentz Gauge Condition & Divergence Constraint
1. [Step 3.b](#wave_eq): Check that $A^i$ satisfies the wave equation
1. [Step 4](#step4): Code Validation
1. [Step 5](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
<a id='initializenrpy'></a>
# Step 1: Initialize needed Python/NRPy+ modules and set destination basis \[Back to [top](#toc)\]
$$\label{initializenrpy}$$
```
# Import needed Python modules
import NRPy_param_funcs as par # NRPy+: Parameter interface
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
#Step 0: Set the spatial dimension parameter to 3.
par.set_parval_from_str("grid::DIM", 3)
DIM = par.parval_from_str("grid::DIM")
# Choices are: Spherical, SinhSpherical, SinhSphericalv2, Cylindrical, SinhCylindrical,
# SymTP, SinhSymTP
dst_basis = "Cylindrical"
# To help with simplifications, we tell Sympy that
# the coordinate xx0 is radial like (positive real)
radial_like_dst_xx0 = True
# Set coordinate system to Cartesian
par.set_parval_from_str("reference_metric::CoordSystem","Cartesian")
rfm.reference_metric()
```
<a id='step2'></a>
# Step 2: A Purely Toriodal Dipole Field \[Back to [top](#toc)\]
$$\label{step2}$$
Having the evolution equations from [Knapp, Walker & Baumgarte (2002)](https://arxiv.org/abs/gr-qc/0201051) written in [Tutorial-VacuumMaxwell_Cartesian_RHSs](Tutorial-VacuumMaxwell_Cartesian_RHSs.ipynb), we must construct the initial data that will then be time evolved. Beginning from the analytic solution to this system of equation given by equation 16 of [Knapp, Walker & Baumgarte (2002)](https://arxiv.org/abs/gr-qc/0201051),
\begin{align}
A^{\hat{\phi}} &= \mathcal{A} \sin \theta \left( \frac{e^{-\lambda v^2}-e^{-\lambda u^2}}{r^2} - 2 \lambda \frac{ve^{-\lambda v^2}-ue^{-\lambda u^2}}{r} \right), \\
\end{align}
where $A^{\hat{\phi}} = A^{\phi} r\sin\theta$, $\mathcal{A}$ gives the amplitude, $\lambda$ describes the size of the wavepacket, $u = t+r$, and $v = t-r$. Other components of the vector potential are $0$. Note that these expressions repesent the exact solution to both systems of equations at any time $t \geq 0$, at all points on our numerical grid. Thus, to get initial data we set $t=0$.
For system II, we will also need to set initial data for $\Gamma$. Since $\Gamma = -\partial_t \psi$ and we have chosen $\psi(t=0) = 0$, $\Gamma(t=0) = 0$.
We may calculate $E^i$ using
\begin{align}
E^i = -\partial_t A^i.
\end{align}
**Inputs for initial data**:
* amp - $A$
* lam - $\lambda$
* time - $t$
Below we define the Cartesian coordinates $x, y$ and $z$. We then define the vector potential $A^i$ in spherical coordinates, but each component is written in terms of Cartesian coordinates. This makes the subsequent basis changes easier.
```
x = rfm.xx_to_Cart[0]
y = rfm.xx_to_Cart[1]
z = rfm.xx_to_Cart[2]
# Step 1: Declare free parameters intrinsic to these initial data
# Amplitude
amp = par.Cparameters("REAL",__name__,"amp", default_vals=1.0)
# lambda
lam = par.Cparameters("REAL",__name__,"lam", default_vals=1.0)
time = par.Cparameters("REAL",__name__,"time", default_vals=0.0)
wavespeed = par.Cparameters("REAL",__name__,"wavespeed", default_vals=1.0)
psi_ID = sp.sympify(0)
Gamma_ID = sp.sympify(0)
```
\begin{align}
A^{\hat{\phi}} &= \mathcal{A} \sin \theta \left( \frac{e^{-\lambda v^2}-e^{-\lambda u^2}}{r^2} - 2 \lambda \frac{ve^{-\lambda v^2}-ue^{-\lambda u^2}}{r} \right), \\
A^{\phi} &= \frac{A^{\hat{\phi}}} {r\sin\theta}
\end{align}
```
AidU_Sph = ixp.zerorank1()
# Set coordinate transformations:
r = sp.sqrt(x*x + y*y + z*z)
sin_theta = z / r
u = time + r
v = time - r
e_lam_u = sp.exp(-lam*u**2)
e_lam_v = sp.exp(-lam*v**2)
# Equation 16 from https://arxiv.org/abs/gr-qc/0201051
AU_phi_hat = (amp*sin_theta)*( ((e_lam_v - e_lam_u)/r**2) - \
2*lam*(v*e_lam_v + u*e_lam_u)/r )
AidU_Sph[2] = AU_phi_hat/(r*sin_theta)
```
<a id='cart_basis'></a>
## Step 2.a: Converting to Cartesian Basis \[Back to [top](#toc)\]
$$\label{cart_basis}$$
Note that $A^i$ is defined in sperical coordinates, so we must therefore transform to Cartesian coordinates using the [Jacobian](https://en.wikipedia.org/wiki/Jacobian_matrix_and_determinant#Example_3:_spherical-Cartesian_transformation). Here we will use the coordinate transformation definitions provided by [reference_metric.py](../edit/reference_metric.py) to build the Jacobian:
\begin{align}
\frac{\partial x_{\rm Cart}^i}{\partial x_{\rm Sph}^j},
\end{align}
where $x_{\rm Sph}^j \in \{r,\theta,\phi\}$ and $x_{\rm Cart}^i \in \{x,y,z\}$. We then apply it to $A^i$ to transform into Cartesian coordinates, via
\begin{align}
A^i_{\rm Cart} = \frac{\partial x_{\rm Cart}^i}{\partial x_{\rm Sph}^j} A^j_{\rm Sph}.
\end{align}
```
# Coordinate transformation from spherical to Cartesian
AidU_Cart = ixp.zerorank1()
Jac_dxSphU_dxCartD = ixp.zerorank2()
for i in range(DIM):
for j in range(DIM):
Jac_dxSphU_dxCartD[i][j] = sp.diff(rfm.xxSph[i],rfm.xx_to_Cart[j])
# Jac_dxCartU_dxSphD[i][j] = sp.diff(rfm.xx_to_Cart[i],rfm.xx[j])
Jac_dxCartU_dxSphD,dummy = ixp.generic_matrix_inverter3x3(Jac_dxSphU_dxCartD)
for i in range(DIM):
for j in range(DIM):
AidU_Cart[i] += Jac_dxCartU_dxSphD[i][j]*AidU_Sph[j]
for i in range(DIM):
AidU_Cart[i] = sp.simplify(AidU_Cart[i])
```
<a id='dst_basis'></a>
## Step 2.b: Converting to Destination Basis \[Back to [top](#toc)\]
$$\label{dst_basis}$$
Here we prepare to convert $A^i$ from the Cartesian basis to the destination basis. To do so, we first rewrite each component of $A^i$ in terms of the destination coordinates. This is done by first re-labelling the NRPy+ coordinates $xx0, xx1, xx2$ as $cart_{xx0}, cart_{xx1}, cart_{xx2}$. Then, each $cart_{xxi}$ is replaced by its counterpart expression in the destination basis using [reference_metric.py](../edit/reference_metric.py).
Note that for algebraic simplification, we tell sympy that the coordinate $xx0$ is radial like and thus positive and real (if the destination coordinates are curvilinear).
```
# rfm is still defined in Cartesian coordinates
cart_xx = ixp.declarerank1("cart_xx")
for i in range(DIM):
for k in range(DIM):
AidU_Cart[i] = AidU_Cart[i].subs(rfm.xx[k], cart_xx[k])
# Set coordinate system to dst_basis
par.set_parval_from_str("reference_metric::CoordSystem",dst_basis)
rfm.reference_metric()
for i in range(DIM):
for k in range(DIM):
AidU_Cart[i] = AidU_Cart[i].subs(cart_xx[k], rfm.xx_to_Cart[k])
if radial_like_dst_xx0:
for j in range(DIM):
AidU_Cart[j] = sp.refine(sp.simplify(AidU_Cart[j]), sp.Q.positive(rfm.xx[0]))
```
We define Jacobians relative to the center of the destination grid, at a point $x^j_{\rm dst}=$(`xx0,xx1,xx2`)${}_{\rm dst}$ on the destination grid:
$$
{\rm Jac\_dUCart\_dDdstUD[i][j]} = \frac{\partial x^i_{\rm Cart}}{\partial x^j_{\rm dst}},
$$
via exact differentiation (courtesy SymPy), and the inverse Jacobian
$$
{\rm Jac\_dUdst\_dDCartUD[i][j]} = \frac{\partial x^i_{\rm dst}}{\partial x^j_{\rm Cart}},
$$
using NRPy+'s `generic_matrix_inverter3x3()` function. In terms of these, the transformation of BSSN tensors from Cartesian to the destination grid's `"reference_metric::CoordSystem"` coordinates may be written:
$$
A^i_{\rm dst} = \frac{\partial x^i_{\rm dst}}{\partial x^\ell_{\rm Cart}} A^\ell_{\rm Cart}
$$
```
# Step 3: Transform BSSN tensors in Cartesian basis to destination grid basis, using center of dest. grid as origin
# Step 3.a: Next construct Jacobian and inverse Jacobian matrices:
Jac_dUCart_dDrfmUD,Jac_dUrfm_dDCartUD = rfm.compute_Jacobian_and_inverseJacobian_tofrom_Cartesian()
# Step 3.b: Convert basis of all BSSN *vectors* from Cartesian to destination basis
AidU = rfm.basis_transform_vectorU_from_Cartesian_to_rfmbasis(Jac_dUrfm_dDCartUD, AidU_Cart)
# Define electric field --> E^i = -\partial_t A^i
EidU = ixp.zerorank1()
for j in range(DIM):
EidU[j] = -sp.diff(AidU[j], time)
```
<a id='step3'></a>
# Step 3: Checks \[Back to [top](#toc)\]
$$\label{step3}$$
Here we validate the initial data. Specifically, we check that the constraints from [Tutorial-VacuumMaxwell_formulation_Curvilinear](Tutorial-VacuumMaxwell_formulation_Curvilinear.ipynb) are satisfied;
\begin{align}
\mathcal{G} &\equiv \Gamma - \partial_i A^i - \hat{\Gamma}^i_{ji} A^j &= 0, \quad &\text{Lorenz gauge condition} \\
\mathcal{C} &\equiv \partial_i E^i + \hat{\Gamma}^i_{ji} E^j &= 0, \quad &\text{Divergence Constraint}.
\end{align}
Note that the above simply to their usual forms in Cartesian coordinates.
Finally, we check that $A^i$ satisfies the covariant wave equation,
\begin{align}
\partial_t^2 A^i - \hat{\gamma}^{jk} \hat{\nabla}_j \hat{\nabla}_k A^i = 0,
\end{align}
where $\hat{\nabla}_j$ is the covariant derivative associated with the spatial metric $\hat{\gamma}_{jk}$.
```
AidU_dD = ixp.zerorank2()
for i in range(DIM):
for j in range(DIM):
AidU_dD[i][j] += sp.diff(AidU[i], rfm.xx[j])
AidU_dDD = ixp.zerorank3()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
AidU_dDD[i][j][k] += sp.diff(AidU[i], rfm.xx[j], rfm.xx[k])
```
<a id='lorentz'></a>
## Step 3.a: Lorentz Gauge Condition & Divergence Constraint \[Back to [top](#toc)\]
$$\label{lorentz}$$
\begin{align}
\mathcal{G} &\equiv \Gamma - \partial_i A^i - \hat{\Gamma}^i_{ji} A^j &= 0, \quad &\text{Lorenz gauge condition} \\
\mathcal{C} &\equiv \partial_i E^i + \hat{\Gamma}^i_{ji} E^j &= 0, \quad &\text{Divergence Constraint}
\end{align}
```
# \mathcal{G} \equiv \Gamma - \partial_i A^i - \hat{\Gamma}^i_{ji} A^j
G = Gamma_ID
for i in range(DIM):
G -= AidU_dD[i][i]
for j in range(DIM):
G -= rfm.GammahatUDD[i][j][i]*AidU[j]
print('G should evaluate to zero:', sp.simplify(G), '\n')
# \mathcal{C} \equiv \partial_i E^i + \hat{\Gamma}^i_{ji} E^j
C = sp.sympify(0)
for i in range(DIM):
C += sp.diff(EidU[i], rfm.xx[i], 1)
for j in range(DIM):
C += rfm.GammahatUDD[i][j][i]*EidU[j]
print('C should evaluate to zero:', sp.simplify(C))
```
<a id='wave_eq'></a>
## Step 3.b: Check that $A^i$ satisfies the wave equation \[Back to [top](#toc)\]
$$\label{wave_eq}$$
Based on the definition of covariant derivative, we have
$$
\hat{\nabla}_{k} A^{i} = A^i_{,k} + \hat{\Gamma}^i_{mk} A^m
$$
Since $\hat{\nabla}_{k} A^{i}$ is a tensor, the covariant derivative of this will have the same indexing as a tensor $T_k^i$:
$$
\hat{\nabla}_{j} T^i_k = T^i_{k,j} + \hat{\Gamma}^i_{dj} T^d_k - \hat{\Gamma}^d_{kj} T^i_d.
$$
Therefore,
\begin{align}
\hat{\nabla}_{j} \left(\hat{\nabla}_{k} A^{i}\right) &= \left(A^i_{,k} + \hat{\Gamma}^i_{mk} A^m\right)_{,j} + \hat{\Gamma}^i_{dj} \left(A^d_{,k} + \hat{\Gamma}^d_{mk} A^m\right) - \hat{\Gamma}^d_{kj} \left(A^i_{,d} + \hat{\Gamma}^i_{md} A^m\right) \\
&= A^i_{,kj} + \hat{\Gamma}^i_{mk,j} A^m + \hat{\Gamma}^i_{mk} A^m_{,j} + \hat{\Gamma}^i_{dj}A^d_{,k} + \hat{\Gamma}^i_{dj}\hat{\Gamma}^d_{mk} A^m - \hat{\Gamma}^d_{kj} A^i_{,d} - \hat{\Gamma}^d_{kj} \hat{\Gamma}^i_{md} A^m \\
&= {\underbrace {\textstyle A^i_{,kj}}_{\text{Term 1}}}+
{\underbrace {\textstyle \hat{\Gamma}^i_{mk,j} A^m + \hat{\Gamma}^i_{mk} A^m_{,j} + \hat{\Gamma}^i_{dj} A^d_{,k} - \hat{\Gamma}^d_{kj} A^i_{,d}}_{\text{Term 2}}} +
{\underbrace {\textstyle \hat{\Gamma}^i_{dj}\hat{\Gamma}^d_{mk} A^m - \hat{\Gamma}^d_{kj} \hat{\Gamma}^i_{md} A^m}_{\text{Term 3}}}.
\end{align}
Thus
$$
\hat{\gamma}^{jk} \hat{\nabla}_{j} \left(\hat{\nabla}_{k} A^{i}\right) = \hat{\gamma}^{jk} \left(\text{Term 1} + \text{Term 2} + \text{Term 3}\right).
$$
We use the above to confirm
\begin{align}
\partial_t^2 A^i - \hat{\gamma}^{jk} \hat{\nabla}_j \hat{\nabla}_k A^i = 0,
\end{align}
$$
\text{Term 1} = A^i_{,kj}
$$
```
# Term 1: A^i_{,kj}
Term1UDD = ixp.zerorank3()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
Term1UDD[i][j][k] += AidU_dDD[i][k][j]
```
$$
\text{Term 2} = \hat{\Gamma}^i_{mk,j} A^m + \hat{\Gamma}^i_{mk} A^m_{,j} + \hat{\Gamma}^i_{dj}A^d_{,k} - \hat{\Gamma}^d_{kj} A^i_{,d}
$$
```
# Term 2: \hat{\Gamma}^i_{mk,j} A^m + \hat{\Gamma}^i_{mk} A^m_{,j}
# + \hat{\Gamma}^i_{dj}A^d_{,k} - \hat{\Gamma}^d_{kj} A^i_{,d}
Term2UDD = ixp.zerorank3()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
for m in range(DIM):
Term2UDD[i][j][k] += rfm.GammahatUDDdD[i][m][k][j]*AidU[m] \
+ rfm.GammahatUDD[i][m][k]*AidU_dD[m][j] \
+ rfm.GammahatUDD[i][m][j]*AidU_dD[m][k] \
- rfm.GammahatUDD[m][k][j]*AidU_dD[i][m]
```
$$
\text{Term 3} = \hat{\Gamma}^i_{dj}\hat{\Gamma}^d_{mk} A^m - \hat{\Gamma}^d_{kj} \hat{\Gamma}^i_{md} A^m
$$
```
# Term 3: \hat{\Gamma}^i_{dj}\hat{\Gamma}^d_{mk} A^m -
# \hat{\Gamma}^d_{kj} \hat{\Gamma}^i_{md} A^m
Term3UDD = ixp.zerorank3()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
for m in range(DIM):
for d in range(DIM):
Term3UDD[i][j][k] += ( rfm.GammahatUDD[i][d][j]*rfm.GammahatUDD[d][m][k] \
-rfm.GammahatUDD[d][k][j]*rfm.GammahatUDD[i][m][d])*AidU[m]
```
$$
\hat{\gamma}^{jk} \hat{\nabla}_{j} \left(\hat{\nabla}_{k} A^{i}\right) = \hat{\gamma}^{jk} \left(\text{Term 1} + \text{Term 2} + \text{Term 3}\right),
$$
$$
\partial_t^2 A^i - \hat{\gamma}^{jk} \hat{\nabla}_j \hat{\nabla}_k A^i = 0
$$
```
# A^i_{,kj} + \hat{\Gamma}^i_{mk,j} A^m + \hat{\Gamma}^i_{mk} A^m_{,j} +
# \hat{\Gamma}^i_{dj}A^d_{,k} + \hat{\Gamma}^i_{dj}\hat{\Gamma}^d_{mk} A^m -
# \hat{\Gamma}^d_{kj} A^i_{,d} - \hat{\Gamma}^d_{kj} \hat{\Gamma}^i_{md} A^m
Difference = ixp.zerorank1()
for i in range(DIM):
Difference[i] = sp.diff(AidU[i], time, 2)
for j in range(DIM):
for k in range(DIM):
Difference[i] += -rfm.ghatUU[k][j]*(Term1UDD[i][j][k] + Term2UDD[i][j][k] + Term3UDD[i][j][k])
for i in range(DIM):
print(str(i)+"th component of A-field equation. Should be zero (takes a bit, please be patient): ")
print(" "+str(sp.simplify(Difference[i])))
```
<a id='step4'></a>
# Step 4: NRPy+ Module Code Validation \[Back to [top](#toc)\]
$$\label{step4}$$
Here, as a code validation check, we verify agreement in the SymPy expressions for the initial data we intend to use between
1. this tutorial and
2. the NRPy+ [InitialData](../edit/Maxwell/InitialData.py) module.
Since the initial data is identical between the two systems for $E^i, A^i$, and $\psi$, we also set and validate initial data for $\Gamma$.
```
import Maxwell.InitialData as mwid
par.set_parval_from_str("Maxwell.InitialData::System_to_use","System_II")
mwid.InitialData()
# Again, to help sympy with simplifications
if radial_like_dst_xx0:
for j in range(DIM):
mwid.AidU[j] = sp.refine(sp.simplify(mwid.AidU[j]), sp.Q.positive(rfm.xx[0]))
mwid.EidU[j] = sp.refine(sp.simplify(mwid.EidU[j]), sp.Q.positive(rfm.xx[0]))
print("Consistency check between this tutorial and NRPy+ module: ALL SHOULD BE ZERO.")
print("psi_ID - mwid.psi_ID = " + str(sp.simplify(psi_ID) - mwid.psi_ID))
print("Gamma_ID - mwid.Gamma_ID = " + str(Gamma_ID - mwid.Gamma_ID))
for i in range(DIM):
print("AidU["+str(i)+"] - mwid.AidU["+str(i)+"] = " + str(sp.simplify(AidU[i] - mwid.AidU[i])))
print("EidU["+str(i)+"] - mwid.EidU["+str(i)+"] = " + str(sp.simplify(EidU[i] - mwid.EidU[i])))
```
<a id='latex_pdf_output'></a>
# Step 5: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-VacuumMaxwell_InitialData.pdf](Tutorial-VacuumMaxwell_InitialData.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-VacuumMaxwell_InitialData")
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import pandas as pd
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import math
import seaborn as sns
import matplotlib.colors as mcolors
import statsmodels.api as sm
import statsmodels.formula.api as smf
from statsmodels.formula.api import ols
from statsmodels.formula.api import mixedlm
import os
colors = list(mcolors.TABLEAU_COLORS.keys())*2
full_names = {
'AU': 'Australia',
'BR': 'Brazil',
'CA': 'Canada',
'FR': 'France',
'DE': 'Germany',
'IN': 'India',
'IT': 'Italy',
'MX': 'Mexico',
'ES': 'Spain',
'GB': 'United Kingdom',
'US': 'United States',
'DK': 'Denmark',
'KE': 'Kenya',
'NG': 'Nigeria',
'JP': 'Japan',
'SE': 'Sweden',
'ID': 'Indonesia',
'EG': 'Egypt'
}
event_dicts = [{'country': 'AU',
'end_md_1': '2020-06-07',
'start_md_1': '2020-03-27',
'start_md_2': np.nan},
{'country': 'BR',
'end_md_1': '2020-08-09',
'start_md_1': '2020-03-23',
'start_md_2': np.nan},
{'country': 'CA',
'end_md_1': '2020-06-21',
'start_md_1': '2020-03-19',
'start_md_2': '2020-10-12'},
{'country': 'DE',
'end_md_1': '2020-05-09',
'start_md_1': '2020-03-21',
'start_md_2': '2020-12-18'},
{'country': 'DK',
'end_md_1': '2020-05-07',
'start_md_1': '2020-03-17',
'start_md_2': np.nan},
{'country': 'EG',
'end_md_1': '2020-07-01',
'start_md_1': '2020-03-24',
'start_md_2': np.nan},
{'country': 'ES',
'end_md_1': '2020-06-14',
'start_md_1': '2020-03-17',
'start_md_2': '2020-11-07'},
{'country': 'FR',
'end_md_1': '2020-06-08',
'start_md_1': '2020-03-18',
'start_md_2': '2020-11-01'},
{'country': 'GB',
'end_md_1': '2020-08-03',
'start_md_1': '2020-03-23',
'start_md_2': '2020-10-21'},
{'country': 'ID',
'end_md_1': '2020-08-10',
'start_md_1': '2020-03-24',
'start_md_2': np.nan},
{'country': 'IN',
'end_md_1': '2020-10-29',
'start_md_1': '2020-03-24',
'start_md_2': np.nan},
{'country': 'IT',
'end_md_1': '2020-06-06',
'start_md_1': '2020-03-11',
'start_md_2': '2020-11-06'},
{'country': 'JP',
'end_md_1': '2020-05-30',
'start_md_1': '2020-04-12',
'start_md_2': np.nan},
{'country': 'KE',
'end_md_1': '2020-10-04',
'start_md_1': '2020-03-24',
'start_md_2': np.nan},
{'country': 'MX',
'end_md_1': '2020-10-06',
'start_md_1': '2020-03-25',
'start_md_2': np.nan},
{'country': 'NG',
'end_md_1': '2020-08-09',
'start_md_1': '2020-03-27',
'start_md_2': np.nan},
{'country': 'SE',
'end_md_1': '2020-04-09',
'start_md_1': '2020-04-03',
'start_md_2': np.nan},
{'country': 'US',
'end_md_1': '2020-06-11',
'start_md_1': '2020-03-21',
'start_md_2': '2020-11-26'}]
parentDirectory = os.path.abspath(os.path.join(os.path.join(os.path.join(os.getcwd(), os.pardir), os.pardir),os.pardir))
DATA_DIR = parentDirectory +'/data/'
FIGURES_DIR = parentDirectory +'/figures/'
df_events = pd.DataFrame(event_dicts)
df_events['start_md_1'] = pd.to_datetime(df_events['start_md_1'])
df_events['end_md_1'] = pd.to_datetime(df_events['end_md_1'])
df_events['start_md_2'] = pd.to_datetime(df_events['start_md_2'])
df_agg = pd.read_pickle(DATA_DIR+'df_agg_modes.pickle')
df_agg
weeks_2019 = list(df_agg.iloc[0]['volume_weekly_total'].index)[:52]
weeks_2020 = list(df_agg.iloc[0]['volume_weekly_total'].index)[52:]
l = []
for cnt, row in df_agg.iterrows():
start_md = df_events.loc[df_events['country'] == row['country']].iloc[0]['start_md_1']
end_md = df_events.loc[df_events['country'] == row['country']].iloc[0]['end_md_1']
start_md2 = df_events.loc[df_events['country'] == row['country']].iloc[0]['start_md_2']
for week in zip(row['volume_weekly_total'].index,row['volume_weekly_total'].values,row['volume_percent_weekly_total'].values):
entry = {}
entry['country'] = row['country']
entry['category'] = row['category']
if week[0] in weeks_2020:
date = pd.to_datetime(week[0])
if type(start_md2)!=pd._libs.tslibs.nattype.NaTType and date > start_md2:
continue
entry['k'] = math.floor(((date - start_md).days +7) / 7)
entry['volume_total'] = week[1]
entry['volume_percent'] = week[2]
entry['year'] = '2020'
l.append(entry)
elif week[0] in weeks_2019:
date = pd.to_datetime(weeks_2020[weeks_2019.index(week[0])])
if type(start_md2)!=pd._libs.tslibs.nattype.NaTType and date > start_md2:
continue
entry['k'] = math.floor(((date - start_md).days +7) / 7)
entry['volume_total'] = week[1]
entry['volume_percent'] = week[2]
entry['year'] = '2019'
l.append(entry)
df = pd.DataFrame(l)
df
k = 30
df = df.loc[(df['k'] >= -30) & (df['k'] <= 30)].copy()
df['intervention_flag'] = df['k'].apply(lambda x: 1 if x >= 0 else 0)
df
def generate_equation(order):
if order == 'Cubic':
eq = "volume_total ~ intervention_flag*k*year + intervention_flag*np.power(k,2)*year + intervention_flag*np.power(k,3)*year"
elif order == "Quadratic":
eq = "volume_total ~ intervention_flag*k*year + intervention_flag*np.power(k,2)*year"
elif order == "Linear":
eq = "volume_total ~ intervention_flag*k*year"
elif order == 'Constant':
eq = "volume_total ~ intervention_flag*year"
return eq
def generate_equation_interactions(order):
if order == 'Cubic':
eq = "volume_total ~ intervention_flag*k*year*C(country)*C(category) + intervention_flag*np.power(k,2)*year*C(country)*C(category) + intervention_flag*np.power(k,3)*year*C(country)*C(category)"
elif order == "Quadratic":
eq = "volume_total ~ intervention_flag*k*year*C(country)*C(category) + intervention_flag*np.power(k,2)*year*C(country)*C(category)"
elif order == "Linear":
eq = "volume_total ~ intervention_flag*k*year*C(country)*C(category)"
elif order == 'Constant':
eq = "volume_total ~ intervention_flag*year*C(country)*C(category)"
return eq
df_temp = df.loc[(df['k'] >= -k) & (df['k'] <= k)].copy()
df_temp['volume_total'] = df_temp['volume_total'].apply(lambda x: np.log(x + 0.001))
mod = smf.ols(generate_equation_interactions('Linear'), data = df_temp)
result_interactions = mod.fit(cov_type='hc0')
def get_standard_error_sum(covariates):
'''
#95CI is approximated with +- 2 sum_variance_standard_error
'''
#get the variance covariance matrix
vcov = result_interactions.cov_params()\
.loc[covariates,covariates].values
#calculate the sum of all pair wise covariances by summing up
m_sum = np.sum(vcov)
#variance of a sum of variables is the square root
return np.sqrt((m_sum))
cats = ['Mode 1','Mode 2','Mode 3','Mode 4']
default_country = 'AU'
default_category = 'Mode 1'
alpha_baseline = 'intervention_flag:year[T.2020]'
beta_baseline = 'intervention_flag:k:year[T.2020]'
list_results = []
for country in full_names.keys():
for c in cats:
entry = {}
entry['country'] = country
entry['category'] = c
suffix_country = (':C(country)[T.'+country+']')
suffix_category = (':C(category)[T.'+c+']')
if country == default_country and c == default_category:
total_alpha = (result_interactions.params[alpha_baseline])
total_alpha_error = (result_interactions.bse[alpha_baseline])
total_beta = (result_interactions.params[beta_baseline])
total_beta_error = (result_interactions.bse[beta_baseline])
elif country == default_country and c != default_category:
total_alpha = (result_interactions.params[alpha_baseline]) \
+ (result_interactions.params[alpha_baseline + suffix_category])
total_alpha_error = (get_standard_error_sum([alpha_baseline,
alpha_baseline + suffix_category]))
total_beta = (result_interactions.params[beta_baseline]) \
+ (result_interactions.params[beta_baseline + suffix_category])
total_beta_error = (get_standard_error_sum([beta_baseline,
beta_baseline + suffix_category]))
elif country != default_country and c == default_category:
total_alpha = (result_interactions.params[alpha_baseline]) \
+ (result_interactions.params[alpha_baseline + suffix_country])
total_alpha_error = (get_standard_error_sum([alpha_baseline,
alpha_baseline + suffix_country]))
total_beta = (result_interactions.params[beta_baseline]) \
+ (result_interactions.params[beta_baseline + suffix_country])
total_beta_error = (get_standard_error_sum([beta_baseline,
beta_baseline + suffix_country]))
else:
total_alpha = (result_interactions.params[alpha_baseline]) \
+ (result_interactions.params[alpha_baseline + suffix_country]) \
+ (result_interactions.params[alpha_baseline + suffix_category]) \
+ (result_interactions.params[alpha_baseline + suffix_country + suffix_category])
total_alpha_error = (get_standard_error_sum([alpha_baseline,
alpha_baseline + suffix_category,
alpha_baseline + suffix_country,
alpha_baseline + suffix_country + suffix_category]))
total_beta = (result_interactions.params[beta_baseline]) \
+ (result_interactions.params[beta_baseline + suffix_country]) \
+ (result_interactions.params[beta_baseline + suffix_category]) \
+ (result_interactions.params[beta_baseline + suffix_country + suffix_category])
total_beta_error = (get_standard_error_sum([beta_baseline,
beta_baseline + suffix_category,
beta_baseline + suffix_country,
beta_baseline + suffix_country + suffix_category]))
entry['alpha'] = total_alpha
entry['alpha_ste'] = total_alpha_error
entry['beta'] = total_beta
entry['beta_ste'] = total_beta_error
list_results.append(entry)
df_results = pd.DataFrame(list_results)
countries_sorted = list(df_results.loc[df_results['category'] == 'Mode 1'].\
sort_values(by = 'alpha', ascending = False)['country'].values)
cats_sorted = list(df_results.groupby('category')['alpha'].agg('mean').sort_values(ascending = False).index)
#countries_sorted = list(df_results.groupby('country')['alpha'].\
# agg('mean').sort_values(ascending = False).index)
sorterIndex = dict(zip(countries_sorted, range(len(countries_sorted))))
def sort_pd(key=None,reverse=False):
def sorter(series):
series_list = list(series)
return [series_list.index(i)
for i in sorted(series_list,key=key,reverse=reverse)]
return sorter
sort_by_custom_dict = sort_pd(key=sorterIndex.get)
dict_annotate = {'Mode 1': 'Recipe, cooking, baking, grocery\n store, supermarket',
'Mode 2': 'Food delivery, take-out,\n drive-in',
'Mode 3': 'Restaurant, careteria, cafe,\n diner, food festival',
'Mode 4': 'Picnic, barbecue, \nlunchbox'}
fig, axes = plt.subplots(2,2, figsize = (6,6), sharey = True)
for cnt,c in enumerate(['Mode 1','Mode 2','Mode 4','Mode 3']):
sbplt = axes[math.floor(cnt/2), cnt%2]
x = df_results.loc[df_results['category'] == c].iloc[sort_by_custom_dict(df_results.loc[df_results['category'] == c]['country'])][['alpha','country','alpha_ste']]
colors_bars = []
for i in range(18):
if x['alpha'].values[i]>0 and x['alpha'].values[i]-2*x['alpha_ste'].values[i]>0:
colors_bars.append('darkmagenta')
elif x['alpha'].values[i]<0 and x['alpha'].values[i]+2*x['alpha_ste'].values[i]<0:
colors_bars.append('darkgoldenrod')
else:
colors_bars.append('silver')
#sbplt.bar(range(12),x['alpha'].apply(lambda x: np.exp(x)-1), yerr = 2*x['alpha_ste'].apply(lambda x: np.exp(x)-1), color = colors_bars)
sbplt.bar(range(18),x['alpha'].apply(lambda x: np.exp(x)-1),
#here we convert errors back to linear scale
yerr = np.array([x['alpha'].apply(lambda x: np.exp(x)-1) - (x['alpha']-2*x['alpha_ste']).apply(lambda x: np.exp(x)-1),
(x['alpha']+2*x['alpha_ste']).apply(lambda x: np.exp(x)-1) - x['alpha'].apply(lambda x: np.exp(x)-1)]),
color = colors_bars)
sbplt.set_xticks(range(18))
sbplt.set_xticklabels(x['country'], fontsize= 7)
sbplt.set_title(dict_annotate[c], size= 11, style='italic')
sbplt.set_yticks([-2,-1,0,1,2])
sbplt.set_yticklabels(["-200%","-100%","0","+100%","+200%"])
sbplt.set_ylim([-2.5,2.5])
#fig.suptitle("α", position = (0.5, 1.05))
size_l = 12
fig.text(0.55, -0.14, 'Prepared by whom?', ha='center', fontsize= size_l)
fig.text(0.32, -0.08, 'By persons within the\nhousehold or social group', ha='center', fontsize= size_l)
fig.text(0.77, -0.08, 'By a third party\n', ha='center', fontsize= size_l)
fig.text(-0.1, 0.5, 'Consumed where?', va='center', rotation='vertical', fontsize= size_l)
fig.text(-0.04, 0.25, 'Outside of home', va='center', rotation='vertical', fontsize= size_l)
fig.text(-0.04, 0.75, 'At home', va='center', rotation='vertical', fontsize= size_l)
plt.tight_layout()
plt.savefig(FIGURES_DIR+"appendix_linear_modes.pdf", bbox_inches='tight')
```
| github_jupyter |
# Convolutional Neural Networks
*by Marvin Bertin*
<img src="../../images/keras-tensorflow-logo.jpg" width="400">
## Convolutional Neural Networks (CNNs)
Convolutional Neural Networks are very similar to ordinary (fully connected) Neural Networks. They are made up of neurons that have learnable weights and biases. Each neuron receives some inputs, performs a dot product and optionally follows it with a non-linearity function. The whole network represents a single differentiable score function. This function takes in as input raw image pixels on one end and computes class scores at the other. The output scores are fed into a loss function to compute the multinomial class probabilities.
### What makes CNN special?
CNN architectures make the explicit assumption that the inputs are images, which allows us to encode certain properties into the architecture related to the image features. These then make the forward function more efficient to implement and vastly reduce the amount of parameters in the network. Below is a list of all the main components in a CNN.
### Main components found in CNNs
- inputs matrix - image 4D tensor
- output class scores
- convolutional layers
- fully-connected layers
- activation functions
- max-pooling layers
- dropout layers
- softmax layer for multinomial class probabilities
- loss function
## Fully Connected (Dense) Layer
<div style="float:right;margin-right:5px;">
<img src="../../images/SingleNeuron.png" width="300" />
<p style="text-align:center;">*Single feedforward neuron*</p>
</div>
<br>
**Feedforward computation**
$\textstyle h_{W,b}(x) = f(W^Tx) = f(\sum_{i=1}^3 W_{i}x_i +b)$ <br>
$f =$ activation function <br>
$W =$ weight vector/matrix <br>
$b =$ bias scalar/vector <br>
A fully connected (dense) layer is the most basic layer in neural networks.
Neurons in a fully connected layer have full connections to all activations in the previous layer. The layer basically computes a weighted sum of the previous layer followed by an activation function for each neuron in the output layer. The weighted sum across all neurons can be computed with a matrix multiplication between the input vector and weight matrix followed by a bias offset.
Fully connected layers are not spatially located since there is no weight sharing(as we'll see with convolutional layers). Therefore the input to a fully connect layer must be reshaped to a vector.
### Fully Connected Neural Network
<br>
<img src="../../images/NN1.gif" width="500">
## Convolutional Layer
The convolutional layer is the core building block of a CNN. The layer's parameters consist of a set of learnable filters (or kernels), which have a small receptive field, but extend through the full depth of the input volume. During the forward pass, each filter is convolved across the width and height of the input volume, computing the dot product between the entries of the filter and the input and producing a 2-dimensional activation map of that filter. As a result, the network learns filters that activate when it detects some specific type of feature at some spatial position in the input. Stacking the activation maps for all filters along the depth dimension forms the full output volume of the convolution layer.
<br>
<div style="float:left;margin-right:5px;">
<img src="../../images/Conv3.jpeg" width="300" />
<p style="text-align:center;">*2D Convolution on color image*</p>
</div>
<div style="float:center;margin-right:5px;">
<img src="../../images/neuron_model.jpeg" width="350" />
<p style="text-align:center;">*A Neural Network "neuron"*</p>
</div>
Convolutional Neural Networks take advantage of the fact that the input consists of images and they constrain the architecture based in this assumption. In particular, unlike a regular Neural Network, the layers of a ConvNet have neurons arranged in 3 dimensions: width, height, depth.
Convolutional layers provide 3 big benefits for Computer Vision:
1. **Location Invariance** - because of the sliding filters, the exact location of important features is not important, which allows the model to generalize better to unseen images (pooling also provides invariance)
2. **Local connectivity** - Convolutional networks exploit spatially local correlation by enforcing a local connectivity pattern (receptive field) between neurons of adjacent layers. This is in contrast to fully connected layers that do not take into account the spatial structure of the input.
3. **Compositionality** - CNN layers are generally stacked on top of eachother. Allowing the model to construct incrementally higher-level representation of the image, making the classification task easier at the last layer.
<img src="../../images/Convolution.gif" width="400">
<center>Convolution with 3×3 Filter</center>
## Activation Layer
The activation function is a layer, which applies an elementwise non-linearity to the output of a parameterized layer (ie conv, dense). The most commonly used activation for CNNs is the ReLu function.
ReLU is the abbreviation of Rectified Linear Units. This is a layer of neurons that applies the non-saturating activation function $f(x)=\max(0,x)$. It increases the nonlinear properties of the decision function and of the overall network without affecting the receptive fields of the convolution layer.
Other functions are also used to increase nonlinearity, for example the saturating hyperbolic tangent $f(x)=\tanh(x)$ and the sigmoid function $f(x)=(1+e^{-x})^{-1}$. Compared to other functions the usage of ReLU is preferable, because it results in the neural network training several times faster, without making a significant difference to generalisation accuracy.
<img src="../../images/activations.png" width="600">
## Max-Pooling Layer
A pooling layer is a type of downsampling layer. In this category, there are several layer options, with maxpooling being the most popular. This basically takes a filter (usually of size 2x2) and a stride of the same length. It then applies it to the input spacial features and outputs the maximum number in every subregion that the filter convolves around.
Other options for pooling layers are average pooling and L2-norm pooling. The intuitive reasoning behind this layer is that once we know that a specific feature is in the original input volume (there will be a high activation value), its exact location is not as important as its relative location to the other features. This layer drastically reduces the spatial dimension (the length and the width change but not the depth) of the input tensor.
This serves two main purposes:
1. Reduce the amount of parameters and computation in the network
2. Control overfitting (high train accuracy but low test accuracy)
<br>
<div style="float:left;margin-right:5px;">
<img src="../../images/pool.jpeg" width="300" />
<p style="text-align:center;">*Spatial downsampling with filter size 2, stride 2*</p>
</div>
<div style="float:center;margin-right:5px;">
<img src="../../images/maxpool.jpeg" width="400" />
<p style="text-align:center;">*Maxpooling operation*</p>
</div>
Pooling layer downsamples the volume spatially, independently in each depth slice of the input volume. In this example, the input volume of size [224x224x64] is pooled with filter size 2, stride 2 into output volume of size [112x112x64].
## Dropout Layer
Neural network models can quickly become very expressive which allows them to represent very complex functions. This expressiveness is needed in image recognition due to their high dimensional and non-linear nature. However, overly complex models can lead to the problem of overfitting. Where after training, the weights of the network are so tuned to the training examples that the network doesn’t perform well and can't generalize to new unseen examples.
The solution is to apply regularization. There are many ways to regularize a neural network. A common method is to add the L2-norm of the model's weights to the cost function. The L2 regularization has the intuitive interpretation of heavily penalizing peaky weight vectors and preferring diffuse weight vectors. This forces the network to use all of its inputs a little rather that some of its inputs a lot.
<img src="../../images/dropout1.png" width="600">
Dropout is an extremely effective and simple regularization technique that complements the other regularization methods. This layer “drops out” a random set of activations in that layer by setting them to zero in the forward pass. it forces the network to be redundant. That means that the network should be able to provide the right classification or output for a specific example even if some of the activations are dropped out. During testing there is no dropout applied, with the interpretation of evaluating an averaged prediction across the exponentially-sized ensemble of all sub-networks
## Convolutional Neural Network Architecture
A CNN architecture is formed by a stack of distinct layers that transform the input volume into an output volume ( holding the class scores) through a differentiable function.
The four main type of layers are:
1. Convolutional Layer (contains trainable parameters)
2. Fully-Connected Layer (contains trainable parameters)
3. Pooling Layer (fixed function)
4. Activation Layer (fixed function)
<img src="../../images/convnet.jpeg" width="800">
## Next Lesson
### CNN layers in TF-Keras
- You will learn aboutthe different layers in TF-Keras
<img src="../../images/divider.png" width="100">
| github_jupyter |
# Recurrent Neural Networks
When working with sequential data (time-series, sentences, etc.) the order of the inputs is crucial for the task at hand. Recurrent neural networks (RNNs) process sequential data by accounting for the current input and also what has been learned from previous inputs. In this notebook, we'll learn how to create and train RNNs on sequential data.
<img src="figures/rnn.png" width=550>
# Overview
* **Objective:** Process sequential data by accounting for the currend input and also what has been learned from previous inputs.
* **Advantages:**
* Account for order and previous inputs in a meaningful way.
* Conditioned generation for generating sequences.
* **Disadvantages:**
* Each time step's prediction depends on the previous prediction so it's difficult to parallelize RNN operations.
* Processing long sequences can yield memory and computation issues.
* Interpretability is difficult but there are few [techniques](https://arxiv.org/abs/1506.02078) that use the activations from RNNs to see what parts of the inputs are processed.
* **Miscellaneous:**
* Architectural tweaks to make RNNs faster and interpretable is an ongoing area of research.
<img src="figures/rnn2.png" width=650>
RNN forward pass for a single time step $X_t$:
$h_t = tanh(W_{hh}h_{t-1} + W_{xh}X_t+b_h)$
$y_t = W_{hy}h_t + b_y $
$ P(y) = softmax(y_t) = \frac{e^y}{\sum e^y} $
*where*:
* $X_t$ = input at time step t | $\in \mathbb{R}^{NXE}$ ($N$ is the batch size, $E$ is the embedding dim)
* $W_{hh}$ = hidden units weights| $\in \mathbb{R}^{HXH}$ ($H$ is the hidden dim)
* $h_{t-1}$ = previous timestep's hidden state $\in \mathbb{R}^{NXH}$
* $W_{xh}$ = input weights| $\in \mathbb{R}^{EXH}$
* $b_h$ = hidden units bias $\in \mathbb{R}^{HX1}$
* $W_{hy}$ = output weights| $\in \mathbb{R}^{HXC}$ ($C$ is the number of classes)
* $b_y$ = output bias $\in \mathbb{R}^{CX1}$
You repeat this for every time step's input ($X_{t+1}, X_{t+2}, ..., X_{N})$ to the get the predicted outputs at each time step.
**Note**: At the first time step, the previous hidden state $h_{t-1}$ can either be a zero vector (unconditioned) or initialize (conditioned). If we are conditioning the RNN, the first hidden state $h_0$ can belong to a specific condition or we can concat the specific condition to the randomly initialized hidden vectors at each time step. More on this in the subsequent notebooks on RNNs.
Let's see what the forward pass looks like with an RNN for a synthetic task such as processing reviews (a sequence of words) to predict the sentiment at the end of processing the review.
```
# Let's make sure the libraries are installed
#!pip install numpy
#!pip install torch
#!pip install matplotlib
#!pip install pandas
# Now import the libraries
import torch
import torch.nn as nn
import torch.nn.functional as F
import warnings
warnings.filterwarnings('ignore')
batch_size = 5
seq_size = 10 # max length per input (masking will be used for sequences that aren't this max length)
x_lengths = [8, 5, 4, 10, 5] # lengths of each input sequence
embedding_dim = 100
rnn_hidden_dim = 256
output_dim = 4
# Initialize synthetic inputs
x_in = torch.randn(batch_size, seq_size, embedding_dim)
x_lengths = torch.tensor(x_lengths)
print (x_in.size())
# Initialize hidden state
hidden_t = torch.zeros((batch_size, rnn_hidden_dim))
print (hidden_t.size())
# Initialize RNN cell
rnn_cell = nn.RNNCell(embedding_dim, rnn_hidden_dim)
print (rnn_cell)
# Forward pass through RNN
x_in = x_in.permute(1, 0, 2) # RNN needs batch_size to be at dim 1
# Loop through the inputs time steps
hiddens = []
for t in range(seq_size):
hidden_t = rnn_cell(x_in[t], hidden_t)
hiddens.append(hidden_t)
hiddens = torch.stack(hiddens)
hiddens = hiddens.permute(1, 0, 2) # bring batch_size back to dim 0
print (hiddens.size())
# We also could've used a more abstracted layer
x_in = torch.randn(batch_size, seq_size, embedding_dim)
rnn = nn.RNN(embedding_dim, rnn_hidden_dim, batch_first=True)
out, h_n = rnn(x_in) #h_n is the last hidden state
print ("out: ", out.size())
print ("h_n: ", h_n.size())
def gather_last_relevant_hidden(hiddens, x_lengths):
x_lengths = x_lengths.long().detach().cpu().numpy() - 1
out = []
for batch_index, column_index in enumerate(x_lengths):
out.append(hiddens[batch_index, column_index])
return torch.stack(out)
# Gather the last relevant hidden state
z = gather_last_relevant_hidden(hiddens, x_lengths)
print (z.size())
# Forward pass through FC layer
fc1 = nn.Linear(rnn_hidden_dim, output_dim)
y_pred = fc1(z)
y_pred = F.softmax(y_pred, dim=1)
print (y_pred.size())
print (y_pred)
```
# Sequential data
There are a variety of different sequential tasks that RNNs can help with.
1. **One to one**: there is one input and produces one output.
* Ex. Given a word predict it's class (verb, noun, etc.).
2. **One to many**: one input generates many outputs.
* Ex. Given a sentiment (positive, negative, etc.) generate a review.
3. **Many to one**: Many inputs are sequentially processed to generate one output.
* Ex. Process the words in a review to predict the sentiment.
4. **Many to many**: Many inputs are sequentially processed to generate many outputs.
* Ex. Given a sentence in French, processes the entire sentence and then generate the English translation.
* Ex. Given a sequence of time-series data, predict the probability of an event (risk of disease) at each time step.
<img src="figures/seq2seq.jpeg" width=700>
# Issues with vanilla RNNs
There are several issues with the vanilla RNN that we've seen so far.
1. When we have an input sequence that has many time steps, it becomes difficult for the model to retain information seen earlier as we process more and more of the downstream timesteps. The goals of the model is to retain the useful components in the previously seen time steps but this becomes cumbersome when we have so many time steps to process.
2. During backpropagation, the gradient from the loss has to travel all the way back towards the first time step. If our gradient is larger than 1 (${1.01}^{1000} = 20959$) or less than 1 (${0.99}^{1000} = 4.31e-5$) and we have lot's of time steps, this can quickly spiral out of control.
To address both these issues, the concept of gating was introduced to RNNs. Gating allows RNNs to control the information flow between each time step to optimize on the task. Selectively allowing information to pass through allows the model to process inputs with many time steps. The most common RNN gated varients are the long short term memory ([LSTM](https://pytorch.org/docs/stable/nn.html#torch.nn.LSTM)) units and gated recurrent units ([GRUs](https://pytorch.org/docs/stable/nn.html#torch.nn.GRU)). You can read more about how these units work [here](http://colah.github.io/posts/2015-08-Understanding-LSTMs/).
<img src="figures/gates.png" width=900>
```
# GRU in PyTorch
gru = nn.GRU(input_size=embedding_dim, hidden_size=rnn_hidden_dim,
batch_first=True)
# Initialize synthetic input
x_in = torch.randn(batch_size, seq_size, embedding_dim)
print (x_in.size())
# Forward pass
out, h_n = gru(x_in)
print ("out:", out.size())
print ("h_n:", h_n.size())
```
**Note**: Choosing whether to use GRU or LSTM really depends on the data and empirical performance. GRUs offer comparable performance with reduce number of parameters while LSTMs are more efficient and may make the difference in performance for your particular task.
# Bidirectional RNNs
There have been many advancements with RNNs ([attention](https://www.oreilly.com/ideas/interpretability-via-attentional-and-memory-based-interfaces-using-tensorflow), Quasi RNNs, etc.) that we will cover in later lessons but one of the basic and widely used ones are bidirectional RNNs (Bi-RNNs). The motivation behind bidirectional RNNs is to process an input sequence by both directions. Accounting for context from both sides can aid in performance when the entire input sequence is known at time of inference. A common application of Bi-RNNs is in translation where it's advantageous to look at an entire sentence from both sides when translating to another language (ie. Japanese → English).
<img src="figures/birnn.png" width=700>
```
# BiGRU in PyTorch
bi_gru = nn.GRU(input_size=embedding_dim, hidden_size=rnn_hidden_dim,
batch_first=True, bidirectional=True)
# Forward pass
out, h_n = bi_gru(x_in)
print ("out:", out.size()) # collection of all hidden states from the RNN for each time step
print ("h_n:", h_n.size()) # last hidden state from the RNN
```
Notice that the output for each sample at each timestamp has size 512 (double the hidden dim). This is because this includes both the forward and backward directions from the BiRNN.
# Document classification with RNNs
Let's apply RNNs to the document classification task from the emebddings notebook (12_Embeddings.ipynb) where we want to predict an article's category given its title.
## Set up
```
import os
from argparse import Namespace
import collections
import copy
import json
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import re
import torch
# Set Numpy and PyTorch seeds
def set_seeds(seed, cuda):
np.random.seed(seed)
torch.manual_seed(seed)
if cuda:
torch.cuda.manual_seed_all(seed)
# Creating directories
def create_dirs(dirpath):
if not os.path.exists(dirpath):
os.makedirs(dirpath)
# Arguments
args = Namespace(
seed=1234,
cuda=True,
shuffle=True,
data_file="data/news.csv",
vectorizer_file="vectorizer.json",
model_state_file="model.pth",
save_dir="news",
train_size=0.7,
val_size=0.15,
test_size=0.15,
pretrained_embeddings=None,
cutoff=25, # token must appear at least <cutoff> times to be in SequenceVocabulary
num_epochs=5,
early_stopping_criteria=5,
learning_rate=1e-3,
batch_size=64,
embedding_dim=100,
rnn_hidden_dim=128,
hidden_dim=100,
num_layers=1,
bidirectional=False,
dropout_p=0.1,
)
# Set seeds
set_seeds(seed=args.seed, cuda=args.cuda)
# Create save dir
create_dirs(args.save_dir)
# Expand filepaths
args.vectorizer_file = os.path.join(args.save_dir, args.vectorizer_file)
args.model_state_file = os.path.join(args.save_dir, args.model_state_file)
# Check CUDA
if not torch.cuda.is_available():
args.cuda = False
args.device = torch.device("cuda" if args.cuda else "cpu")
print("Using CUDA: {}".format(args.cuda))
```
## Data
```
import re
import urllib
# Raw data
df = pd.read_csv(args.data_file, header=0)
df.head()
# Split by category
by_category = collections.defaultdict(list)
for _, row in df.iterrows():
by_category[row.category].append(row.to_dict())
for category in by_category:
print ("{0}: {1}".format(category, len(by_category[category])))
# Create split data
final_list = []
for _, item_list in sorted(by_category.items()):
if args.shuffle:
np.random.shuffle(item_list)
n = len(item_list)
n_train = int(args.train_size*n)
n_val = int(args.val_size*n)
n_test = int(args.test_size*n)
# Give data point a split attribute
for item in item_list[:n_train]:
item['split'] = 'train'
for item in item_list[n_train:n_train+n_val]:
item['split'] = 'val'
for item in item_list[n_train+n_val:]:
item['split'] = 'test'
# Add to final list
final_list.extend(item_list)
# df with split datasets
split_df = pd.DataFrame(final_list)
split_df["split"].value_counts()
# Preprocessing
def preprocess_text(text):
text = ' '.join(word.lower() for word in text.split(" "))
text = re.sub(r"([.,!?])", r" \1 ", text)
text = re.sub(r"[^a-zA-Z.,!?]+", r" ", text)
text = text.strip()
return text
split_df.title = split_df.title.apply(preprocess_text)
split_df.head()
```
## Vocabulary
```
class Vocabulary(object):
def __init__(self, token_to_idx=None):
# Token to index
if token_to_idx is None:
token_to_idx = {}
self.token_to_idx = token_to_idx
# Index to token
self.idx_to_token = {idx: token \
for token, idx in self.token_to_idx.items()}
def to_serializable(self):
return {'token_to_idx': self.token_to_idx}
@classmethod
def from_serializable(cls, contents):
return cls(**contents)
def add_token(self, token):
if token in self.token_to_idx:
index = self.token_to_idx[token]
else:
index = len(self.token_to_idx)
self.token_to_idx[token] = index
self.idx_to_token[index] = token
return index
def add_tokens(self, tokens):
return [self.add_token[token] for token in tokens]
def lookup_token(self, token):
return self.token_to_idx[token]
def lookup_index(self, index):
if index not in self.idx_to_token:
raise KeyError("the index (%d) is not in the Vocabulary" % index)
return self.idx_to_token[index]
def __str__(self):
return "<Vocabulary(size=%d)>" % len(self)
def __len__(self):
return len(self.token_to_idx)
# Vocabulary instance
category_vocab = Vocabulary()
for index, row in df.iterrows():
category_vocab.add_token(row.category)
print (category_vocab) # __str__
print (len(category_vocab)) # __len__
index = category_vocab.lookup_token("Business")
print (index)
print (category_vocab.lookup_index(index))
```
## Sequence vocabulary
Next, we're going to create our Vocabulary classes for the article's title, which is a sequence of tokens.
```
from collections import Counter
import string
class SequenceVocabulary(Vocabulary):
def __init__(self, token_to_idx=None, unk_token="<UNK>",
mask_token="<MASK>", begin_seq_token="<BEGIN>",
end_seq_token="<END>"):
super(SequenceVocabulary, self).__init__(token_to_idx)
self.mask_token = mask_token
self.unk_token = unk_token
self.begin_seq_token = begin_seq_token
self.end_seq_token = end_seq_token
self.mask_index = self.add_token(self.mask_token)
self.unk_index = self.add_token(self.unk_token)
self.begin_seq_index = self.add_token(self.begin_seq_token)
self.end_seq_index = self.add_token(self.end_seq_token)
# Index to token
self.idx_to_token = {idx: token \
for token, idx in self.token_to_idx.items()}
def to_serializable(self):
contents = super(SequenceVocabulary, self).to_serializable()
contents.update({'unk_token': self.unk_token,
'mask_token': self.mask_token,
'begin_seq_token': self.begin_seq_token,
'end_seq_token': self.end_seq_token})
return contents
def lookup_token(self, token):
return self.token_to_idx.get(token, self.unk_index)
def lookup_index(self, index):
if index not in self.idx_to_token:
raise KeyError("the index (%d) is not in the SequenceVocabulary" % index)
return self.idx_to_token[index]
def __str__(self):
return "<SequenceVocabulary(size=%d)>" % len(self.token_to_idx)
def __len__(self):
return len(self.token_to_idx)
# Get word counts
word_counts = Counter()
for title in split_df.title:
for token in title.split(" "):
if token not in string.punctuation:
word_counts[token] += 1
# Create SequenceVocabulary instance
title_vocab = SequenceVocabulary()
for word, word_count in word_counts.items():
if word_count >= args.cutoff:
title_vocab.add_token(word)
print (title_vocab) # __str__
print (len(title_vocab)) # __len__
index = title_vocab.lookup_token("general")
print (index)
print (title_vocab.lookup_index(index))
```
## Vectorizer
Something new that we introduce in this Vectorizer is calculating the length of our input sequence. We will use this later on to extract the last relevant hidden state for each input sequence.
```
class NewsVectorizer(object):
def __init__(self, title_vocab, category_vocab):
self.title_vocab = title_vocab
self.category_vocab = category_vocab
def vectorize(self, title):
indices = [self.title_vocab.lookup_token(token) for token in title.split(" ")]
indices = [self.title_vocab.begin_seq_index] + indices + \
[self.title_vocab.end_seq_index]
# Create vector
title_length = len(indices)
vector = np.zeros(title_length, dtype=np.int64)
vector[:len(indices)] = indices
return vector, title_length
def unvectorize(self, vector):
tokens = [self.title_vocab.lookup_index(index) for index in vector]
title = " ".join(token for token in tokens)
return title
@classmethod
def from_dataframe(cls, df, cutoff):
# Create class vocab
category_vocab = Vocabulary()
for category in sorted(set(df.category)):
category_vocab.add_token(category)
# Get word counts
word_counts = Counter()
for title in df.title:
for token in title.split(" "):
word_counts[token] += 1
# Create title vocab
title_vocab = SequenceVocabulary()
for word, word_count in word_counts.items():
if word_count >= cutoff:
title_vocab.add_token(word)
return cls(title_vocab, category_vocab)
@classmethod
def from_serializable(cls, contents):
title_vocab = SequenceVocabulary.from_serializable(contents['title_vocab'])
category_vocab = Vocabulary.from_serializable(contents['category_vocab'])
return cls(title_vocab=title_vocab, category_vocab=category_vocab)
def to_serializable(self):
return {'title_vocab': self.title_vocab.to_serializable(),
'category_vocab': self.category_vocab.to_serializable()}
# Vectorizer instance
vectorizer = NewsVectorizer.from_dataframe(split_df, cutoff=args.cutoff)
print (vectorizer.title_vocab)
print (vectorizer.category_vocab)
vectorized_title, title_length = vectorizer.vectorize(preprocess_text(
"Roger Federer wins the Wimbledon tennis tournament."))
print (np.shape(vectorized_title))
print ("title_length:", title_length)
print (vectorized_title)
print (vectorizer.unvectorize(vectorized_title))
```
## Dataset
```
from torch.utils.data import Dataset, DataLoader
class NewsDataset(Dataset):
def __init__(self, df, vectorizer):
self.df = df
self.vectorizer = vectorizer
# Data splits
self.train_df = self.df[self.df.split=='train']
self.train_size = len(self.train_df)
self.val_df = self.df[self.df.split=='val']
self.val_size = len(self.val_df)
self.test_df = self.df[self.df.split=='test']
self.test_size = len(self.test_df)
self.lookup_dict = {'train': (self.train_df, self.train_size),
'val': (self.val_df, self.val_size),
'test': (self.test_df, self.test_size)}
self.set_split('train')
# Class weights (for imbalances)
class_counts = df.category.value_counts().to_dict()
def sort_key(item):
return self.vectorizer.category_vocab.lookup_token(item[0])
sorted_counts = sorted(class_counts.items(), key=sort_key)
frequencies = [count for _, count in sorted_counts]
self.class_weights = 1.0 / torch.tensor(frequencies, dtype=torch.float32)
@classmethod
def load_dataset_and_make_vectorizer(cls, df, cutoff):
train_df = df[df.split=='train']
return cls(df, NewsVectorizer.from_dataframe(train_df, cutoff))
@classmethod
def load_dataset_and_load_vectorizer(cls, df, vectorizer_filepath):
vectorizer = cls.load_vectorizer_only(vectorizer_filepath)
return cls(df, vectorizer)
def load_vectorizer_only(vectorizer_filepath):
with open(vectorizer_filepath) as fp:
return NewsVectorizer.from_serializable(json.load(fp))
def save_vectorizer(self, vectorizer_filepath):
with open(vectorizer_filepath, "w") as fp:
json.dump(self.vectorizer.to_serializable(), fp)
def set_split(self, split="train"):
self.target_split = split
self.target_df, self.target_size = self.lookup_dict[split]
def __str__(self):
return "<Dataset(split={0}, size={1})".format(
self.target_split, self.target_size)
def __len__(self):
return self.target_size
def __getitem__(self, index):
row = self.target_df.iloc[index]
title_vector, title_length = self.vectorizer.vectorize(row.title)
category_index = self.vectorizer.category_vocab.lookup_token(row.category)
return {'title': title_vector, 'title_length': title_length,
'category': category_index}
def get_num_batches(self, batch_size):
return len(self) // batch_size
def generate_batches(self, batch_size, collate_fn, shuffle=True,
drop_last=False, device="cpu"):
dataloader = DataLoader(dataset=self, batch_size=batch_size,
collate_fn=collate_fn, shuffle=shuffle,
drop_last=drop_last)
for data_dict in dataloader:
out_data_dict = {}
for name, tensor in data_dict.items():
out_data_dict[name] = data_dict[name].to(device)
yield out_data_dict
# Dataset instance
dataset = NewsDataset.load_dataset_and_make_vectorizer(df=split_df,
cutoff=args.cutoff)
print (dataset) # __str__
input_ = dataset[5] # __getitem__
print (input_['title'], input_['title_length'], input_['category'])
print (dataset.vectorizer.unvectorize(input_['title']))
print (dataset.class_weights)
```
## Model
input → embedding → RNN → FC
```
import torch.nn as nn
import torch.nn.functional as F
def gather_last_relevant_hidden(hiddens, x_lengths):
x_lengths = x_lengths.long().detach().cpu().numpy() - 1
out = []
for batch_index, column_index in enumerate(x_lengths):
out.append(hiddens[batch_index, column_index])
return torch.stack(out)
class NewsModel(nn.Module):
def __init__(self, embedding_dim, num_embeddings, rnn_hidden_dim,
hidden_dim, output_dim, num_layers, bidirectional, dropout_p,
pretrained_embeddings=None, freeze_embeddings=False,
padding_idx=0):
super(NewsModel, self).__init__()
if pretrained_embeddings is None:
self.embeddings = nn.Embedding(embedding_dim=embedding_dim,
num_embeddings=num_embeddings,
padding_idx=padding_idx)
else:
pretrained_embeddings = torch.from_numpy(pretrained_embeddings).float()
self.embeddings = nn.Embedding(embedding_dim=embedding_dim,
num_embeddings=num_embeddings,
padding_idx=padding_idx,
_weight=pretrained_embeddings)
# Conv weights
self.gru = nn.GRU(input_size=embedding_dim, hidden_size=rnn_hidden_dim,
num_layers=num_layers, batch_first=True,
bidirectional=bidirectional)
# FC weights
self.dropout = nn.Dropout(dropout_p)
self.fc1 = nn.Linear(rnn_hidden_dim, hidden_dim)
self.fc2 = nn.Linear(hidden_dim, output_dim)
if freeze_embeddings:
self.embeddings.weight.requires_grad = False
def forward(self, x_in, x_lengths, apply_softmax=False):
# Embed
x_in = self.embeddings(x_in)
# Feed into RNN
out, h_n = self.gru(x_in)
# Gather the last relevant hidden state
out = gather_last_relevant_hidden(out, x_lengths)
# FC layers
z = self.dropout(out)
z = self.fc1(z)
z = self.dropout(z)
y_pred = self.fc2(z)
if apply_softmax:
y_pred = F.softmax(y_pred, dim=1)
return y_pred
```
## Training
```
import torch.optim as optim
class Trainer(object):
def __init__(self, dataset, model, model_state_file, save_dir, device, shuffle,
num_epochs, batch_size, learning_rate, early_stopping_criteria):
self.dataset = dataset
self.class_weights = dataset.class_weights.to(device)
self.model = model.to(device)
self.save_dir = save_dir
self.device = device
self.shuffle = shuffle
self.num_epochs = num_epochs
self.batch_size = batch_size
self.loss_func = nn.CrossEntropyLoss(self.class_weights)
self.optimizer = optim.Adam(self.model.parameters(), lr=learning_rate)
self.scheduler = optim.lr_scheduler.ReduceLROnPlateau(
optimizer=self.optimizer, mode='min', factor=0.5, patience=1)
self.train_state = {
'done_training': False,
'stop_early': False,
'early_stopping_step': 0,
'early_stopping_best_val': 1e8,
'early_stopping_criteria': early_stopping_criteria,
'learning_rate': learning_rate,
'epoch_index': 0,
'train_loss': [],
'train_acc': [],
'val_loss': [],
'val_acc': [],
'test_loss': -1,
'test_acc': -1,
'model_filename': model_state_file}
def update_train_state(self):
# Verbose
print ("[EPOCH]: {0} | [LR]: {1} | [TRAIN LOSS]: {2:.2f} | [TRAIN ACC]: {3:.1f}% | [VAL LOSS]: {4:.2f} | [VAL ACC]: {5:.1f}%".format(
self.train_state['epoch_index'], self.train_state['learning_rate'],
self.train_state['train_loss'][-1], self.train_state['train_acc'][-1],
self.train_state['val_loss'][-1], self.train_state['val_acc'][-1]))
# Save one model at least
if self.train_state['epoch_index'] == 0:
torch.save(self.model.state_dict(), self.train_state['model_filename'])
self.train_state['stop_early'] = False
# Save model if performance improved
elif self.train_state['epoch_index'] >= 1:
loss_tm1, loss_t = self.train_state['val_loss'][-2:]
# If loss worsened
if loss_t >= self.train_state['early_stopping_best_val']:
# Update step
self.train_state['early_stopping_step'] += 1
# Loss decreased
else:
# Save the best model
if loss_t < self.train_state['early_stopping_best_val']:
torch.save(self.model.state_dict(), self.train_state['model_filename'])
# Reset early stopping step
self.train_state['early_stopping_step'] = 0
# Stop early ?
self.train_state['stop_early'] = self.train_state['early_stopping_step'] \
>= self.train_state['early_stopping_criteria']
return self.train_state
def compute_accuracy(self, y_pred, y_target):
_, y_pred_indices = y_pred.max(dim=1)
n_correct = torch.eq(y_pred_indices, y_target).sum().item()
return n_correct / len(y_pred_indices) * 100
def pad_seq(self, seq, length):
vector = np.zeros(length, dtype=np.int64)
vector[:len(seq)] = seq
vector[len(seq):] = self.dataset.vectorizer.title_vocab.mask_index
return vector
def collate_fn(self, batch):
# Make a deep copy
batch_copy = copy.deepcopy(batch)
processed_batch = {"title": [], "title_length": [], "category": []}
# Get max sequence length
get_length = lambda sample: len(sample["title"])
max_seq_length = max(map(get_length, batch))
# Pad
for i, sample in enumerate(batch_copy):
padded_seq = self.pad_seq(sample["title"], max_seq_length)
processed_batch["title"].append(padded_seq)
processed_batch["title_length"].append(sample["title_length"])
processed_batch["category"].append(sample["category"])
# Convert to appropriate tensor types
processed_batch["title"] = torch.LongTensor(
processed_batch["title"])
processed_batch["title_length"] = torch.LongTensor(
processed_batch["title_length"])
processed_batch["category"] = torch.LongTensor(
processed_batch["category"])
return processed_batch
def run_train_loop(self):
for epoch_index in range(self.num_epochs):
self.train_state['epoch_index'] = epoch_index
# Iterate over train dataset
# initialize batch generator, set loss and acc to 0, set train mode on
self.dataset.set_split('train')
batch_generator = self.dataset.generate_batches(
batch_size=self.batch_size, collate_fn=self.collate_fn,
shuffle=self.shuffle, device=self.device)
running_loss = 0.0
running_acc = 0.0
self.model.train()
for batch_index, batch_dict in enumerate(batch_generator):
# zero the gradients
self.optimizer.zero_grad()
# compute the output
y_pred = self.model(batch_dict['title'], batch_dict['title_length'])
# compute the loss
loss = self.loss_func(y_pred, batch_dict['category'])
loss_t = loss.item()
running_loss += (loss_t - running_loss) / (batch_index + 1)
# compute gradients using loss
loss.backward()
# use optimizer to take a gradient step
self.optimizer.step()
# compute the accuracy
acc_t = self.compute_accuracy(y_pred, batch_dict['category'])
running_acc += (acc_t - running_acc) / (batch_index + 1)
self.train_state['train_loss'].append(running_loss)
self.train_state['train_acc'].append(running_acc)
# Iterate over val dataset
# # initialize batch generator, set loss and acc to 0; set eval mode on
self.dataset.set_split('val')
batch_generator = self.dataset.generate_batches(
batch_size=self.batch_size, collate_fn=self.collate_fn,
shuffle=self.shuffle, device=self.device)
running_loss = 0.
running_acc = 0.
self.model.eval()
for batch_index, batch_dict in enumerate(batch_generator):
# compute the output
y_pred = self.model(batch_dict['title'], batch_dict['title_length'])
# compute the loss
loss = self.loss_func(y_pred, batch_dict['category'])
loss_t = loss.to("cpu").item()
running_loss += (loss_t - running_loss) / (batch_index + 1)
# compute the accuracy
acc_t = self.compute_accuracy(y_pred, batch_dict['category'])
running_acc += (acc_t - running_acc) / (batch_index + 1)
self.train_state['val_loss'].append(running_loss)
self.train_state['val_acc'].append(running_acc)
self.train_state = self.update_train_state()
self.scheduler.step(self.train_state['val_loss'][-1])
if self.train_state['stop_early']:
break
def run_test_loop(self):
# initialize batch generator, set loss and acc to 0; set eval mode on
self.dataset.set_split('test')
batch_generator = self.dataset.generate_batches(
batch_size=self.batch_size, collate_fn=self.collate_fn,
shuffle=self.shuffle, device=self.device)
running_loss = 0.0
running_acc = 0.0
self.model.eval()
for batch_index, batch_dict in enumerate(batch_generator):
# compute the output
y_pred = self.model(batch_dict['title'], batch_dict['title_length'])
# compute the loss
loss = self.loss_func(y_pred, batch_dict['category'])
loss_t = loss.item()
running_loss += (loss_t - running_loss) / (batch_index + 1)
# compute the accuracy
acc_t = self.compute_accuracy(y_pred, batch_dict['category'])
running_acc += (acc_t - running_acc) / (batch_index + 1)
self.train_state['test_loss'] = running_loss
self.train_state['test_acc'] = running_acc
def plot_performance(self):
# Figure size
plt.figure(figsize=(15,5))
# Plot Loss
plt.subplot(1, 2, 1)
plt.title("Loss")
plt.plot(trainer.train_state["train_loss"], label="train")
plt.plot(trainer.train_state["val_loss"], label="val")
plt.legend(loc='upper right')
# Plot Accuracy
plt.subplot(1, 2, 2)
plt.title("Accuracy")
plt.plot(trainer.train_state["train_acc"], label="train")
plt.plot(trainer.train_state["val_acc"], label="val")
plt.legend(loc='lower right')
# Save figure
plt.savefig(os.path.join(self.save_dir, "performance.png"))
# Show plots
plt.show()
def save_train_state(self):
self.train_state["done_training"] = True
with open(os.path.join(self.save_dir, "train_state.json"), "w") as fp:
json.dump(self.train_state, fp)
# Initialization
dataset = NewsDataset.load_dataset_and_make_vectorizer(df=split_df,
cutoff=args.cutoff)
dataset.save_vectorizer(args.vectorizer_file)
vectorizer = dataset.vectorizer
model = NewsModel(embedding_dim=args.embedding_dim,
num_embeddings=len(vectorizer.title_vocab),
rnn_hidden_dim=args.rnn_hidden_dim,
hidden_dim=args.hidden_dim,
output_dim=len(vectorizer.category_vocab),
num_layers=args.num_layers,
bidirectional=args.bidirectional,
dropout_p=args.dropout_p,
pretrained_embeddings=None,
padding_idx=vectorizer.title_vocab.mask_index)
print (model.named_modules)
# Train
trainer = Trainer(dataset=dataset, model=model,
model_state_file=args.model_state_file,
save_dir=args.save_dir, device=args.device,
shuffle=args.shuffle, num_epochs=args.num_epochs,
batch_size=args.batch_size, learning_rate=args.learning_rate,
early_stopping_criteria=args.early_stopping_criteria)
trainer.run_train_loop()
# Plot performance
trainer.plot_performance()
# Test performance
trainer.run_test_loop()
print("Test loss: {0:.2f}".format(trainer.train_state['test_loss']))
print("Test Accuracy: {0:.1f}%".format(trainer.train_state['test_acc']))
# Save all results
trainer.save_train_state()
```
## Inference
```
class Inference(object):
def __init__(self, model, vectorizer, device="cpu"):
self.model = model.to(device)
self.vectorizer = vectorizer
self.device = device
def predict_category(self, dataset):
# Batch generator
batch_generator = dataset.generate_batches(
batch_size=len(dataset), shuffle=False, device=self.device)
self.model.eval()
# Predict
for batch_index, batch_dict in enumerate(batch_generator):
# compute the output
y_pred = self.model(batch_dict['title'], batch_dict["title_length"],
apply_softmax=True)
# Top k nationalities
y_prob, indices = torch.topk(y_pred, k=len(self.vectorizer.category_vocab))
probabilities = y_prob.detach().to('cpu').numpy()[0]
indices = indices.detach().to('cpu').numpy()[0]
results = []
for probability, index in zip(probabilities, indices):
category = self.vectorizer.category_vocab.lookup_index(index)
results.append({'category': category, 'probability': probability})
return results
# Load vectorizer
with open(args.vectorizer_file) as fp:
vectorizer = NewsVectorizer.from_serializable(json.load(fp))
# Load the model
model = NewsModel(embedding_dim=args.embedding_dim,
num_embeddings=len(vectorizer.title_vocab),
rnn_hidden_dim=args.rnn_hidden_dim,
hidden_dim=args.hidden_dim,
output_dim=len(vectorizer.category_vocab),
num_layers=args.num_layers,
bidirectional=args.bidirectional,
dropout_p=args.dropout_p,
pretrained_embeddings=None,
padding_idx=vectorizer.title_vocab.mask_index)
model.load_state_dict(torch.load(args.model_state_file))
print (model.named_modules)
# Initialize
inference = Inference(model=model, vectorizer=vectorizer, device=args.device)
class InferenceDataset(Dataset):
def __init__(self, df, vectorizer):
self.df = df
self.vectorizer = vectorizer
self.target_size = len(self.df)
def __str__(self):
return "<Dataset(size={1})>".format(self.target_size)
def __len__(self):
return self.target_size
def __getitem__(self, index):
row = self.df.iloc[index]
title_vector, title_length = self.vectorizer.vectorize(row.title)
return {'title': title_vector, 'title_length': title_length}
def get_num_batches(self, batch_size):
return len(self) // batch_size
def generate_batches(self, batch_size, shuffle=True, drop_last=False, device="cpu"):
dataloader = DataLoader(dataset=self, batch_size=batch_size,
shuffle=shuffle, drop_last=drop_last)
for data_dict in dataloader:
out_data_dict = {}
for name, tensor in data_dict.items():
out_data_dict[name] = data_dict[name].to(device)
yield out_data_dict
# Inference
title = input("Enter a title to classify: ")
infer_df = pd.DataFrame([title], columns=['title'])
infer_df.title = infer_df.title.apply(preprocess_text)
infer_dataset = InferenceDataset(infer_df, vectorizer)
results = inference.predict_category(dataset=infer_dataset)
results
```
| github_jupyter |
# FormantNet Configuration Code
This code is used to parse the configuration file, if one exists, and save the global variables used by FormantNet into one object, referred to as **cfg** in the other scripts and passed around from function to function.
```
import configparser
class configuration(object):
def __init__(self):
self.TESTRUN = False
self.NFORMANTS = 6
self.NZEROS = 1
self.DIFFWEIGHT = 0.15
self.SAMPLERATE = 16000.0
self.MAX_ANALYSIS_FREQ = self.SAMPLERATE / 2.0
self.MAXFREQ = 8000.0
self.MINFREQ = 0.0
self.MAXBW = 5000.0
self.MINBW = 20.0
self.MAXAMP = 100.0
self.MINAMP = -100.0
self.WINDOW_LENGTH_MSEC = 32.0
self.FRAME_STRIDE_MSEC = 5.0
self.PREEMPH = 0.98
self.SMOOTH_LINEAR = True
self.ENV_SMOOTH_PASSES = 6
self.FLOOR = 0.001
self.SEQUENCE_LENGTH = 64
self.SEQUENCE_STRIDE = self.SEQUENCE_LENGTH
self.BATCH_SIZE = 32
self.LSTM_LAYERS = 1
self.DENSE_LAYERS = 1
self.LSTM_UNITS = 512
self.DENSE_UNITS = 512
self.DENSE_ACTIVATION = 'relu'
self.TOP_ACTIVATION = 'sigmoid'
self.LEARNING_RATE = 0.0001
self.ALLOW_RETRAIN = True
self.EPOCHS = 200
self.PATIENCE = 20
self.DELETE_OLDER_MODELS = True
self.GET_TEST_LOSS = False
self.OUT_EXT = 'txt'
self.REAL_AMPLITUDES = True
self.FREQUENCIES_FIRST = True
self.BIN_SMOOTH_PASSES = 10
def configure(self, configFile=None):
if configFile is not None:
config = configparser.ConfigParser()
config.read(configFile)
self.TESTRUN = config['DEFAULT'].getboolean('TESTRUN', self.TESTRUN)
self.NFORMANTS = config['DEFAULT'].getint('NFORMANTS', self.NFORMANTS)
self.NZEROS = config['DEFAULT'].getint('NZEROS', self.NZEROS)
self.DIFFWEIGHT = config['DEFAULT'].getfloat('DIFFWEIGHT', self.DIFFWEIGHT)
self.SAMPLERATE = config['DEFAULT'].getfloat('SAMPLERATE', self.SAMPLERATE)
self.MAX_ANALYSIS_FREQ = config['DEFAULT'].getfloat('MAX_ANALYSIS_FREQ', self.SAMPLERATE / 2.0)
self.MAXFREQ = config['DEFAULT'].getfloat('MAXFREQ', self.MAXFREQ)
self.MINFREQ = config['DEFAULT'].getfloat('MINFREQ', self.MINFREQ)
self.MAXBW = config['DEFAULT'].getfloat('MAXBW', self.MAXBW)
self.MINBW = config['DEFAULT'].getfloat('MINBW', self.MINBW)
self.MAXAMP = config['DEFAULT'].getfloat('MAXAMP', self.MAXAMP)
self.MINAMP = config['DEFAULT'].getfloat('MINAMP', self.MINAMP)
self.WINDOW_LENGTH_MSEC = config['DEFAULT'].getfloat('WINDOW_LENGTH_MSEC', self.WINDOW_LENGTH_MSEC)
self.FRAME_STRIDE_MSEC = config['DEFAULT'].getfloat('FRAME_STRIDE_MSEC', self.FRAME_STRIDE_MSEC)
self.PREEMPH = config['DEFAULT'].getfloat('PREEMPH', self.PREEMPH)
self.SMOOTH_LINEAR = config['DEFAULT'].getboolean('SMOOTH_LINEAR', self.SMOOTH_LINEAR)
self.ENV_SMOOTH_PASSES = config['DEFAULT'].getint('ENV_SMOOTH_PASSES', self.ENV_SMOOTH_PASSES)
self.FLOOR = config['DEFAULT'].getfloat('FLOOR', self.FLOOR)
self.SEQUENCE_LENGTH = config['DEFAULT'].getint('SEQUENCE_LENGTH', self.SEQUENCE_LENGTH)
self.SEQUENCE_STRIDE = config['DEFAULT'].getint('SEQUENCE_STRIDE', self.SEQUENCE_LENGTH)
self.BATCH_SIZE = config['DEFAULT'].getint('BATCH_SIZE', self.BATCH_SIZE)
self.LSTM_LAYERS = config['DEFAULT'].getint('LSTM_LAYERS', self.LSTM_LAYERS)
self.DENSE_LAYERS = config['DEFAULT'].getint('DENSE_LAYERS', self.DENSE_LAYERS)
self.LSTM_UNITS = config['DEFAULT'].getint('LSTM_UNITS', self.LSTM_UNITS)
self.DENSE_UNITS = config['DEFAULT'].getint('DENSE_UNITS', self.DENSE_UNITS)
self.DENSE_ACTIVATION = config['DEFAULT'].get('DENSE_ACTIVATION', self.DENSE_ACTIVATION)
self.TOP_ACTIVATION = config['DEFAULT'].get('TOP_ACTIVATION', self.TOP_ACTIVATION)
self.LEARNING_RATE = config['DEFAULT'].getfloat('LEARNING_RATE', self.LEARNING_RATE)
self.ALLOW_RETRAIN = config['DEFAULT'].getboolean('ALLOW_RETRAIN', self.ALLOW_RETRAIN)
self.EPOCHS = config['DEFAULT'].getint('EPOCHS', self.EPOCHS)
self.PATIENCE = config['DEFAULT'].getint('PATIENCE', self.PATIENCE)
self.DELETE_OLDER_MODELS = config['DEFAULT'].getboolean('DELETE_OLDER_MODELS', self.DELETE_OLDER_MODELS)
self.GET_TEST_LOSS = config['DEFAULT'].getboolean('GET_TEST_LOSS', self.GET_TEST_LOSS)
self.OUT_EXT = config['DEFAULT'].get('OUT_EXT', self.OUT_EXT)
self.REAL_AMPLITUDES = config['DEFAULT'].getboolean('REAL_AMPLITUDES', self.REAL_AMPLITUDES)
self.FREQUENCIES_FIRST = config['DEFAULT'].getboolean('FREQUENCIES_FIRST', self.FREQUENCIES_FIRST)
self.BIN_SMOOTH_PASSES = config['DEFAULT'].getint('BIN_SMOOTH_PASSES', self.BIN_SMOOTH_PASSES)
if self.MAX_ANALYSIS_FREQ > self.SAMPLERATE / 2.0:
print("MAX_ANALYSIS_FREQ value", self.MAX_ANALYSIS_FREQ,
"is too high; it must be less than or equal to half the SAMPLERATE.")
self.MAX_ANALYSIS_FREQ = self.SAMPLERATE / 2.0
print("Reset MAX_ANALYSIS_FREQ to", self.MAX_ANALYSIS_FREQ)
if self.MINBW <= 0:
print("MINBW value", self.MINBW, "is too low; it must be greater than 0.")
self.MINBW = 1.0
print("Reset self.MINBW to 1.0")
if self.TESTRUN:
self.EPOCHS = 3
# Width of signal analysis window in samples
self.WINDOW_LENGTH_SAMPLES = int(self.WINDOW_LENGTH_MSEC * self.SAMPLERATE / 1000.0)
# Number of samples between analysis window start points
self.FRAME_STRIDE_SAMPLES = int(self.FRAME_STRIDE_MSEC * self.SAMPLERATE / 1000.0)
# Spectral resolution: Number of points per input spectrum
self.SPECTRUM_NPOINTS = int(self.WINDOW_LENGTH_MSEC * self.MAX_ANALYSIS_FREQ / 1000.0) + 1
self.NSUM = self.NFORMANTS + self.NZEROS # Total number of resonances (poles + zeros)
self.NPARAMS = self.NFORMANTS*3 + self.NZEROS*2 # Total number of model output features
def report_status(self):
print("\n\nSUMMARY OF CONFIGURATION SETTINGS:")
print("Test Run:", self.TESTRUN)
print("\n# Formants (poles) to be modeled:", self.NFORMANTS)
print("# Antiformants (zeros) to be modeled:", self.NZEROS)
print("Total # of model output parameters:", self.NPARAMS)
print("\nDelta-frequency weight:", self.DIFFWEIGHT)
print("Wavefile sampling rate:", self.SAMPLERATE, "Hz")
print("Frequency analysis range: 0 -", self.MAX_ANALYSIS_FREQ, "Hz")
print("\nLower and upper limits on formant parameter predictions:")
print("Frequencies:", self.MINFREQ, "-", self.MAXFREQ, "Hz")
print("Bandwidths:", self.MINBW, "-", self.MAXBW, "Hz")
print("Amplitude correction factors:", self.MINAMP, "-", self.MAXAMP, "dB")
print("\nAnalysis window length:", self.WINDOW_LENGTH_MSEC,
"msec (" + str(self.WINDOW_LENGTH_SAMPLES), "samples)")
print("Spectral resolution (model input size):", self.SPECTRUM_NPOINTS, "bins")
print("Analysis window spacing: Once every", self.FRAME_STRIDE_MSEC,
"msec (" + str(self.FRAME_STRIDE_SAMPLES), "samples)")
print("Pre-emphasis factor:", self.PREEMPH)
print("Perform smoothing on linear-scale envelopes (rather than dB-scale):", self.SMOOTH_LINEAR)
print("# of envelope smoothing passes:", self.ENV_SMOOTH_PASSES)
print("Floor value added to linear envelopes before conversion to dB:", self.FLOOR)
print("\nTraining sequence length:", self.SEQUENCE_LENGTH, "frames")
print("Training sequence spacing: every", self.SEQUENCE_STRIDE, "frames")
print("Batch size (sequences per training batch):", self.BATCH_SIZE)
print("\n# of LSTM layers:", self.LSTM_LAYERS)
print("LSTM layer size:", self.LSTM_UNITS, "units")
print("# of Dense layers:", self.DENSE_LAYERS, "(including output layer)")
if self.DENSE_LAYERS > 1:
print("Dense hidden layer size:", self.DENSE_UNITS, "units")
print("Activation function of dense hidden layers:", self.DENSE_ACTIVATION)
print("Activation function of output layer:", self.TOP_ACTIVATION)
print("\nAllow retraining of pre-existing model?:", self.ALLOW_RETRAIN)
print("Optimizer learning rate:", self.LEARNING_RATE)
print("Maximum # of training epochs:", self.EPOCHS)
print("Convergence patience:", self.PATIENCE, "epochs")
print("Delete older models after training?:", self.DELETE_OLDER_MODELS)
print("\nCalculate test set stats and loss?:", self.GET_TEST_LOSS)
print("Output file extension: ." + self.OUT_EXT)
print("Output real (predicted) amplitudes?:", self.REAL_AMPLITUDES)
print("Output in frequencies-first order?:", self.FREQUENCIES_FIRST)
print("# of binomial smoothing passes on output:", self.BIN_SMOOTH_PASSES)
print("\n")
if __name__ == "__main__":
cfg = configuration()
cfg.configure(None)
cfg.report_status()
```
| github_jupyter |
```
##################################################################
#《Python机器学习及实践:从零开始通往Kaggle竞赛之路(2023年度版)》开源代码
#-----------------------------------------------------------------
# @章节号:6.8.2.1(批量标准化的PyTorch实践)
# @作者:范淼
# @电子邮箱:fanmiao.cslt.thu@gmail.com
# @微博:https://weibo.com/fanmiaothu
# @官方交流QQ群号:561500762
##################################################################
from torch import nn, optim
#设定超参数。
INPUT_SIZE = 784
HIDDEN_SIZE = 256
NUM_CLASSES = 10
EPOCHS = 5
BATCH_SIZE = 64
LEARNING_RATE = 1e-3
class FFN_BN(nn.Module):
'''
自定义带有批量标准化功能的前馈神经网络类,继承自nn.Module。
'''
def __init__(self, input_size, hidden_size, num_classes):
super(FFN_BN, self).__init__()
self.l1 = nn.Linear(input_size, hidden_size)
self.relu = nn.ReLU()
self.bn = nn.BatchNorm1d(hidden_size)
self.l2 = nn.Linear(hidden_size, num_classes)
def forward(self, x):
#添加有256个神经元的隐藏层。
out = self.l1(x)
#设定激活函数为ReLU。
out = self.relu(out)
#添加批量标准化层。
out = self.bn(out)
#添加有10个神经元的输出层。
out = self.l2(out)
return out
#初始化带有批量标准化功能的前馈神经网络模型。
model = FFN_BN(INPUT_SIZE, HIDDEN_SIZE, NUM_CLASSES)
#设定神经网络的损失函数。
criterion = nn.CrossEntropyLoss()
#设定神经网络的优化方法。
optimizer = optim.Adam(model.parameters(), lr = LEARNING_RATE)
import pandas as pd
#使用pandas,读取fashion_mnist的训练和测试数据文件。
train_data = pd.read_csv('../datasets/fashion_mnist/fashion_mnist_train.csv')
test_data = pd.read_csv('../datasets/fashion_mnist/fashion_mnist_test.csv')
#从训练数据中,拆解出训练特征和类别标签。
X_train = train_data[train_data.columns[1:]]
y_train = train_data['label']
#从测试数据中,拆解出测试特征和类别标签。
X_test = test_data[train_data.columns[1:]]
y_test = test_data['label']
from sklearn.preprocessing import StandardScaler
#初始化数据标准化处理器。
ss = StandardScaler()
#标准化训练数据特征。
X_train = ss.fit_transform(X_train)
#标准化测试数据特征。
X_test = ss.transform(X_test)
import torch
from torch.utils.data import TensorDataset, DataLoader
#构建适用于PyTorch模型训练的数据结构。
train_tensor = TensorDataset(torch.tensor(X_train.astype('float32')), torch.tensor(y_train.values))
#构建适用于PyTorch模型训练的数据读取器。
train_loader = DataLoader(dataset = train_tensor, batch_size = BATCH_SIZE, shuffle = True)
n_total_steps = len(train_loader)
#开启模型训练。
model.train()
for epoch in range(EPOCHS):
for i, (features, labels) in enumerate(train_loader):
outputs = model(features)
loss = criterion(outputs, labels)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if (i+1) % 300 == 0:
print (f'Epoch [{epoch+1}/{EPOCHS}], Step[{i+1}/{n_total_steps}], Loss: {loss.item():.4f}')
#构建适用于PyTorch模型测试的数据结构。
test_tensor = TensorDataset(torch.tensor(X_test.astype('float32')), torch.tensor(y_test.values))
#构建适用于PyTorch模型测试的数据读取器。
test_loader = DataLoader(dataset = test_tensor, batch_size = BATCH_SIZE, shuffle = False)
#开启模型测试。
model.eval()
n_correct = 0
n_samples = 0
for features, labels in test_loader:
outputs = model(features)
_, predictions = torch.max(outputs.data, 1)
n_samples += labels.size(0)
n_correct += (predictions == labels).sum().item()
acc = 100.0 * n_correct / n_samples
print('带有批量标准化功能的前馈神经网络(PyTorch版本)在fashion_mnist测试集上的准确率为: %.2f%%。' %acc)
```
| github_jupyter |
```
import keras
from keras.models import Sequential, Model, load_model
from keras.layers import Dense, Dropout, Activation, Flatten, Input, Lambda
from keras.layers import Conv2D, MaxPooling2D, AveragePooling2D, Conv1D, MaxPooling1D, LSTM, ConvLSTM2D, GRU, CuDNNLSTM, CuDNNGRU, BatchNormalization, LocallyConnected2D, Permute, TimeDistributed, Bidirectional
from keras.layers import Concatenate, Reshape, Softmax, Conv2DTranspose, Embedding, Multiply
from keras.callbacks import ModelCheckpoint, EarlyStopping, Callback
from keras import regularizers
from keras import backend as K
from keras.utils.generic_utils import Progbar
from keras.layers.merge import _Merge
import keras.losses
from functools import partial
from collections import defaultdict
import tensorflow as tf
from tensorflow.python.framework import ops
import isolearn.keras as iso
import numpy as np
import tensorflow as tf
import logging
logging.getLogger('tensorflow').setLevel(logging.ERROR)
import pandas as pd
import os
import pickle
import numpy as np
import scipy.sparse as sp
import scipy.io as spio
import matplotlib.pyplot as plt
import isolearn.io as isoio
import isolearn.keras as isol
from sequence_logo_helper_protein import plot_protein_logo
import pandas as pd
from keras.backend.tensorflow_backend import set_session
def contain_tf_gpu_mem_usage() :
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
set_session(sess)
contain_tf_gpu_mem_usage()
class EpochVariableCallback(Callback) :
def __init__(self, my_variable, my_func) :
self.my_variable = my_variable
self.my_func = my_func
def on_epoch_begin(self, epoch, logs={}) :
K.set_value(self.my_variable, self.my_func(K.get_value(self.my_variable), epoch))
class IdentityEncoder(iso.SequenceEncoder) :
def __init__(self, seq_len, channel_map) :
super(IdentityEncoder, self).__init__('identity', (seq_len, len(channel_map)))
self.seq_len = seq_len
self.n_channels = len(channel_map)
self.encode_map = channel_map
self.decode_map = {
val : key for key, val in channel_map.items()
}
def encode(self, seq) :
encoding = np.zeros((self.seq_len, self.n_channels))
for i in range(len(seq)) :
if seq[i] in self.encode_map :
channel_ix = self.encode_map[seq[i]]
encoding[i, channel_ix] = 1.
return encoding
def encode_inplace(self, seq, encoding) :
for i in range(len(seq)) :
if seq[i] in self.encode_map :
channel_ix = self.encode_map[seq[i]]
encoding[i, channel_ix] = 1.
def encode_inplace_sparse(self, seq, encoding_mat, row_index) :
raise NotImplementError()
def decode(self, encoding) :
seq = ''
for pos in range(0, encoding.shape[0]) :
argmax_nt = np.argmax(encoding[pos, :])
max_nt = np.max(encoding[pos, :])
if max_nt == 1 :
seq += self.decode_map[argmax_nt]
else :
seq += self.decode_map[-1]
return seq
def decode_sparse(self, encoding_mat, row_index) :
encoding = np.array(encoding_mat[row_index, :].todense()).reshape(-1, 4)
return self.decode(encoding)
class NopTransformer(iso.ValueTransformer) :
def __init__(self, n_classes) :
super(NopTransformer, self).__init__('nop', (n_classes, ))
self.n_classes = n_classes
def transform(self, values) :
return values
def transform_inplace(self, values, transform) :
transform[:] = values
def transform_inplace_sparse(self, values, transform_mat, row_index) :
transform_mat[row_index, :] = np.ravel(values)
#Re-load cached dataframe (shuffled)
dataset_name = "coiled_coil_binders"
experiment = "baker_big_set_5x_negatives"
pair_df = pd.read_csv("pair_df_" + experiment + "_in_shuffled.csv", sep="\t")
print("len(pair_df) = " + str(len(pair_df)))
print(pair_df.head())
#Generate training and test set indexes
valid_set_size = 0.0005
test_set_size = 0.0995
data_index = np.arange(len(pair_df), dtype=np.int)
train_index = data_index[:-int(len(pair_df) * (valid_set_size + test_set_size))]
valid_index = data_index[train_index.shape[0]:-int(len(pair_df) * test_set_size)]
test_index = data_index[train_index.shape[0] + valid_index.shape[0]:]
print('Training set size = ' + str(train_index.shape[0]))
print('Validation set size = ' + str(valid_index.shape[0]))
print('Test set size = ' + str(test_index.shape[0]))
#Sub-select smaller dataset
n_train_pos = 20000
n_train_neg = 20000
n_test_pos = 2000
n_test_neg = 2000
orig_n_train = train_index.shape[0]
orig_n_valid = valid_index.shape[0]
orig_n_test = test_index.shape[0]
train_index_pos = np.nonzero((pair_df.iloc[train_index]['interacts'] == 1).values)[0][:n_train_pos]
train_index_neg = np.nonzero((pair_df.iloc[train_index]['interacts'] == 0).values)[0][:n_train_neg]
train_index = np.concatenate([train_index_pos, train_index_neg], axis=0)
np.random.shuffle(train_index)
test_index_pos = np.nonzero((pair_df.iloc[test_index]['interacts'] == 1).values)[0][:n_test_pos] + orig_n_train + orig_n_valid
test_index_neg = np.nonzero((pair_df.iloc[test_index]['interacts'] == 0).values)[0][:n_test_neg] + orig_n_train + orig_n_valid
test_index = np.concatenate([test_index_pos, test_index_neg], axis=0)
np.random.shuffle(test_index)
print('Training set size = ' + str(train_index.shape[0]))
print('Test set size = ' + str(test_index.shape[0]))
#Calculate sequence lengths
pair_df['amino_seq_1_len'] = pair_df['amino_seq_1'].str.len()
pair_df['amino_seq_2_len'] = pair_df['amino_seq_2'].str.len()
pair_df.head()
#Initialize sequence encoder
seq_length = 81
residue_map = {'D': 0, 'E': 1, 'V': 2, 'K': 3, 'R': 4, 'L': 5, 'S': 6, 'T': 7, 'N': 8, 'H': 9, 'A': 10, 'I': 11, 'G': 12, 'P': 13, 'Q': 14, 'Y': 15, 'W': 16, 'M': 17, 'F': 18, '#': 19}
encoder = IdentityEncoder(seq_length, residue_map)
#Construct data generators
class CategoricalRandomizer :
def __init__(self, case_range, case_probs) :
self.case_range = case_range
self.case_probs = case_probs
self.cases = 0
def get_random_sample(self, index=None) :
if index is None :
return self.cases
else :
return self.cases[index]
def generate_random_sample(self, batch_size=1, data_ids=None) :
self.cases = np.random.choice(self.case_range, size=batch_size, replace=True, p=self.case_probs)
def get_amino_seq(row, index, flip_randomizer, homodimer_randomizer, max_seq_len=seq_length) :
is_flip = True if flip_randomizer.get_random_sample(index=index) == 1 else False
is_homodimer = True if homodimer_randomizer.get_random_sample(index=index) == 1 else False
amino_seq_1, amino_seq_2 = row['amino_seq_1'], row['amino_seq_2']
if is_flip :
amino_seq_1, amino_seq_2 = row['amino_seq_2'], row['amino_seq_1']
if is_homodimer and row['interacts'] < 0.5 :
amino_seq_2 = amino_seq_1
return amino_seq_1, amino_seq_2
flip_randomizer = CategoricalRandomizer(np.arange(2), np.array([0.5, 0.5]))
homodimer_randomizer = CategoricalRandomizer(np.arange(2), np.array([0.95, 0.05]))
batch_size = 32
data_gens = {
gen_id : iso.DataGenerator(
idx,
{ 'df' : pair_df },
batch_size=(idx.shape[0] // batch_size) * batch_size,
inputs = [
{
'id' : 'amino_seq_1',
'source_type' : 'dataframe',
'source' : 'df',
#'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: (get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[0] + "#" * seq_length)[:seq_length],
'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[0],
'encoder' : IdentityEncoder(seq_length, residue_map),
'dim' : (1, seq_length, len(residue_map)),
'sparsify' : False
},
{
'id' : 'amino_seq_2',
'source_type' : 'dataframe',
'source' : 'df',
#'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: (get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[1] + "#" * seq_length)[:seq_length],
'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[1],
'encoder' : IdentityEncoder(seq_length, residue_map),
'dim' : (1, seq_length, len(residue_map)),
'sparsify' : False
},
{
'id' : 'amino_seq_1_len',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: len(get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[0]),
'encoder' : lambda t: t,
'dim' : (1,),
'sparsify' : False
},
{
'id' : 'amino_seq_2_len',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: len(get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[1]),
'encoder' : lambda t: t,
'dim' : (1,),
'sparsify' : False
}
],
outputs = [
{
'id' : 'interacts',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index: row['interacts'],
'transformer' : NopTransformer(1),
'dim' : (1,),
'sparsify' : False
}
],
randomizers = [flip_randomizer, homodimer_randomizer],
shuffle = True
) for gen_id, idx in [('train', train_index), ('valid', valid_index), ('test', test_index)]
}
#Load data matrices
[x_1_train, x_2_train, l_1_train, l_2_train], [y_train] = data_gens['train'][0]
[x_1_test, x_2_test, l_1_test, l_2_test], [y_test] = data_gens['test'][0]
print("x_1_train.shape = " + str(x_1_train.shape))
print("x_2_train.shape = " + str(x_2_train.shape))
print("x_1_test.shape = " + str(x_1_test.shape))
print("x_2_test.shape = " + str(x_2_test.shape))
print("l_1_train.shape = " + str(l_1_train.shape))
print("l2_train.shape = " + str(l_2_train.shape))
print("l_1_test.shape = " + str(l_1_test.shape))
print("l2_test.shape = " + str(l_2_test.shape))
print("y_train.shape = " + str(y_train.shape))
print("y_test.shape = " + str(y_test.shape))
#Define sequence templates
sequence_templates = [
'$' * i + '@' * (seq_length - i)
for i in range(seq_length+1)
]
sequence_masks = [
np.array([1 if sequence_templates[i][j] == '$' else 0 for j in range(len(sequence_templates[i]))])
for i in range(seq_length+1)
]
#Calculate background distributions
pseudo_count = 0.1
x_means = []
x_mean_logits = []
for i in range(seq_length + 1) :
x_train_len = x_1_train[np.ravel(l_1_train) == i, ...]
if x_train_len.shape[0] > 0 :
x_mean_len = (np.sum(x_train_len, axis=(0, 1)) + pseudo_count) / (np.sum(x_train_len, axis=(0, 1, 3)).reshape(-1, 1) + 20. * pseudo_count)
x_mean_logits_len = np.log(x_mean_len)
x_means.append(x_mean_len)
x_mean_logits.append(x_mean_logits_len)
else :
x_means.append(np.zeros((x_1_train.shape[2], x_1_train.shape[3])))
x_mean_logits.append(np.zeros((x_1_train.shape[2], x_1_train.shape[3])))
#Visualize a few background sequence distributions
visualize_len = 67
plot_protein_logo(residue_map, np.copy(x_means[visualize_len]), sequence_template=sequence_templates[visualize_len], figsize=(12, 1), logo_height=1.0, plot_start=0, plot_end=81)
visualize_len = 72
plot_protein_logo(residue_map, np.copy(x_means[visualize_len]), sequence_template=sequence_templates[visualize_len], figsize=(12, 1), logo_height=1.0, plot_start=0, plot_end=81)
visualize_len = 81
plot_protein_logo(residue_map, np.copy(x_means[visualize_len]), sequence_template=sequence_templates[visualize_len], figsize=(12, 1), logo_height=1.0, plot_start=0, plot_end=81)
#Calculate mean training set kl-divergence against background
mean_kl_divs = []
for i in range(seq_length + 1) :
x_train_len = x_1_train[np.ravel(l_1_train) == i, ...]
if x_train_len.shape[0] > 0 :
x_train_clipped_len = np.clip(np.copy(x_train_len[:, 0, :, :]), 1e-8, 1. - 1e-8)
kl_divs = np.sum(x_train_clipped_len * np.log(x_train_clipped_len / np.tile(np.expand_dims(x_means[i], axis=0), (x_train_clipped_len.shape[0], 1, 1))), axis=-1) / np.log(2.0)
x_mean_kl_divs = np.sum(kl_divs * sequence_masks[i], axis=-1) / np.sum(sequence_masks[i])
x_mean_kl_div = np.mean(x_mean_kl_divs)
mean_kl_divs.append(x_mean_kl_div)
print("[Length = " + str(i) + "] Mean KL Div against background (bits) = " + str(x_mean_kl_div))
else :
mean_kl_divs.append(0)
from tensorflow.python.framework import ops
#Stochastic Binarized Neuron helper functions (Tensorflow)
#ST Estimator code adopted from https://r2rt.com/beyond-binary-ternary-and-one-hot-neurons.html
#See Github https://github.com/spitis/
def st_sampled_softmax(logits):
with ops.name_scope("STSampledSoftmax") as namescope :
nt_probs = tf.nn.softmax(logits)
onehot_dim = logits.get_shape().as_list()[1]
sampled_onehot = tf.one_hot(tf.squeeze(tf.multinomial(logits, 1), 1), onehot_dim, 1.0, 0.0)
with tf.get_default_graph().gradient_override_map({'Ceil': 'Identity', 'Mul': 'STMul'}):
return tf.ceil(sampled_onehot * nt_probs)
def st_hardmax_softmax(logits):
with ops.name_scope("STHardmaxSoftmax") as namescope :
nt_probs = tf.nn.softmax(logits)
onehot_dim = logits.get_shape().as_list()[1]
sampled_onehot = tf.one_hot(tf.argmax(nt_probs, 1), onehot_dim, 1.0, 0.0)
with tf.get_default_graph().gradient_override_map({'Ceil': 'Identity', 'Mul': 'STMul'}):
return tf.ceil(sampled_onehot * nt_probs)
@ops.RegisterGradient("STMul")
def st_mul(op, grad):
return [grad, grad]
#Gumbel Distribution Sampler
def gumbel_softmax(logits, temperature=0.5) :
gumbel_dist = tf.contrib.distributions.RelaxedOneHotCategorical(temperature, logits=logits)
batch_dim = logits.get_shape().as_list()[0]
onehot_dim = logits.get_shape().as_list()[1]
return gumbel_dist.sample()
#PWM Masking and Sampling helper functions
def mask_pwm(inputs) :
pwm, onehot_template, onehot_mask = inputs
return pwm * onehot_mask + onehot_template
def sample_pwm_st(pwm_logits) :
n_sequences = K.shape(pwm_logits)[0]
seq_length = K.shape(pwm_logits)[2]
flat_pwm = K.reshape(pwm_logits, (n_sequences * seq_length, 20))
sampled_pwm = st_sampled_softmax(flat_pwm)
return K.reshape(sampled_pwm, (n_sequences, 1, seq_length, 20))
def sample_pwm_gumbel(pwm_logits) :
n_sequences = K.shape(pwm_logits)[0]
seq_length = K.shape(pwm_logits)[2]
flat_pwm = K.reshape(pwm_logits, (n_sequences * seq_length, 20))
sampled_pwm = gumbel_softmax(flat_pwm, temperature=0.5)
return K.reshape(sampled_pwm, (n_sequences, 1, seq_length, 20))
#Generator helper functions
def initialize_sequence_templates(generator, encoder, sequence_templates, background_matrices) :
embedding_templates = []
embedding_masks = []
embedding_backgrounds = []
for k in range(len(sequence_templates)) :
sequence_template = sequence_templates[k]
onehot_template = encoder(sequence_template).reshape((1, len(sequence_template), 20))
for j in range(len(sequence_template)) :
if sequence_template[j] not in ['$', '@'] :
nt_ix = np.argmax(onehot_template[0, j, :])
onehot_template[:, j, :] = -4.0
onehot_template[:, j, nt_ix] = 10.0
onehot_mask = np.zeros((1, len(sequence_template), 20))
for j in range(len(sequence_template)) :
if sequence_template[j] == '$' :
onehot_mask[:, j, :] = 1.0
embedding_templates.append(onehot_template.reshape(1, -1))
embedding_masks.append(onehot_mask.reshape(1, -1))
embedding_backgrounds.append(background_matrices[k].reshape(1, -1))
embedding_templates = np.concatenate(embedding_templates, axis=0)
embedding_masks = np.concatenate(embedding_masks, axis=0)
embedding_backgrounds = np.concatenate(embedding_backgrounds, axis=0)
generator.get_layer('template_dense').set_weights([embedding_templates])
generator.get_layer('template_dense').trainable = False
generator.get_layer('mask_dense').set_weights([embedding_masks])
generator.get_layer('mask_dense').trainable = False
generator.get_layer('background_dense').set_weights([embedding_backgrounds])
generator.get_layer('background_dense').trainable = False
#Generator construction function
def build_sampler(batch_size, seq_length, n_classes=1, n_samples=1, sample_mode='st') :
#Initialize Reshape layer
reshape_layer = Reshape((1, seq_length, 20))
#Initialize background matrix
onehot_background_dense = Embedding(n_classes, seq_length * 20, embeddings_initializer='zeros', name='background_dense')
#Initialize template and mask matrices
onehot_template_dense = Embedding(n_classes, seq_length * 20, embeddings_initializer='zeros', name='template_dense')
onehot_mask_dense = Embedding(n_classes, seq_length * 20, embeddings_initializer='ones', name='mask_dense')
#Initialize Templating and Masking Lambda layer
masking_layer = Lambda(mask_pwm, output_shape = (1, seq_length, 20), name='masking_layer')
background_layer = Lambda(lambda x: x[0] + x[1], name='background_layer')
#Initialize PWM normalization layer
pwm_layer = Softmax(axis=-1, name='pwm')
#Initialize sampling layers
sample_func = None
if sample_mode == 'st' :
sample_func = sample_pwm_st
elif sample_mode == 'gumbel' :
sample_func = sample_pwm_gumbel
upsampling_layer = Lambda(lambda x: K.tile(x, [n_samples, 1, 1, 1]), name='upsampling_layer')
sampling_layer = Lambda(sample_func, name='pwm_sampler')
permute_layer = Lambda(lambda x: K.permute_dimensions(K.reshape(x, (n_samples, batch_size, 1, seq_length, 20)), (1, 0, 2, 3, 4)), name='permute_layer')
def _sampler_func(class_input, raw_logits) :
#Get Template and Mask
onehot_background = reshape_layer(onehot_background_dense(class_input))
onehot_template = reshape_layer(onehot_template_dense(class_input))
onehot_mask = reshape_layer(onehot_mask_dense(class_input))
#Add Template and Multiply Mask
pwm_logits = masking_layer([background_layer([raw_logits, onehot_background]), onehot_template, onehot_mask])
#Compute PWM (Nucleotide-wise Softmax)
pwm = pwm_layer(pwm_logits)
#Tile each PWM to sample from and create sample axis
pwm_logits_upsampled = upsampling_layer(pwm_logits)
sampled_pwm = sampling_layer(pwm_logits_upsampled)
sampled_pwm = permute_layer(sampled_pwm)
sampled_mask = permute_layer(upsampling_layer(onehot_mask))
return pwm_logits, pwm, sampled_pwm, onehot_mask, sampled_mask
return _sampler_func
#Scrambler network definition
def make_resblock(n_channels=64, window_size=8, dilation_rate=1, group_ix=0, layer_ix=0, drop_rate=0.0) :
#Initialize res block layers
batch_norm_0 = BatchNormalization(name='scrambler_resblock_' + str(group_ix) + '_' + str(layer_ix) + '_batch_norm_0')
relu_0 = Lambda(lambda x: K.relu(x, alpha=0.0))
conv_0 = Conv2D(n_channels, (1, window_size), dilation_rate=dilation_rate, strides=(1, 1), padding='same', activation='linear', kernel_initializer='glorot_normal', name='scrambler_resblock_' + str(group_ix) + '_' + str(layer_ix) + '_conv_0')
batch_norm_1 = BatchNormalization(name='scrambler_resblock_' + str(group_ix) + '_' + str(layer_ix) + '_batch_norm_1')
relu_1 = Lambda(lambda x: K.relu(x, alpha=0.0))
conv_1 = Conv2D(n_channels, (1, window_size), dilation_rate=dilation_rate, strides=(1, 1), padding='same', activation='linear', kernel_initializer='glorot_normal', name='scrambler_resblock_' + str(group_ix) + '_' + str(layer_ix) + '_conv_1')
skip_1 = Lambda(lambda x: x[0] + x[1], name='scrambler_resblock_' + str(group_ix) + '_' + str(layer_ix) + '_skip_1')
drop_1 = None
if drop_rate > 0.0 :
drop_1 = Dropout(drop_rate)
#Execute res block
def _resblock_func(input_tensor) :
batch_norm_0_out = batch_norm_0(input_tensor)
relu_0_out = relu_0(batch_norm_0_out)
conv_0_out = conv_0(relu_0_out)
batch_norm_1_out = batch_norm_1(conv_0_out)
relu_1_out = relu_1(batch_norm_1_out)
if drop_rate > 0.0 :
conv_1_out = drop_1(conv_1(relu_1_out))
else :
conv_1_out = conv_1(relu_1_out)
skip_1_out = skip_1([conv_1_out, input_tensor])
return skip_1_out
return _resblock_func
def load_scrambler_network(n_groups=1, n_resblocks_per_group=4, n_channels=32, window_size=8, dilation_rates=[1], drop_rate=0.0) :
#Discriminator network definition
conv_0 = Conv2D(n_channels, (1, 1), strides=(1, 1), padding='same', activation='linear', kernel_initializer='glorot_normal', name='scrambler_conv_0')
skip_convs = []
resblock_groups = []
for group_ix in range(n_groups) :
skip_convs.append(Conv2D(n_channels, (1, 1), strides=(1, 1), padding='same', activation='linear', kernel_initializer='glorot_normal', name='scrambler_skip_conv_' + str(group_ix)))
resblocks = []
for layer_ix in range(n_resblocks_per_group) :
resblocks.append(make_resblock(n_channels=n_channels, window_size=window_size, dilation_rate=dilation_rates[group_ix], group_ix=group_ix, layer_ix=layer_ix, drop_rate=drop_rate))
resblock_groups.append(resblocks)
last_block_conv = Conv2D(n_channels, (1, 1), strides=(1, 1), padding='same', activation='linear', kernel_initializer='glorot_normal', name='scrambler_last_block_conv')
skip_add = Lambda(lambda x: x[0] + x[1], name='scrambler_skip_add')
final_conv = Conv2D(1, (1, 1), strides=(1, 1), padding='same', activation='softplus', kernel_initializer='glorot_normal', name='scrambler_final_conv')
onehot_to_logits = Lambda(lambda x: 2. * x - 1., name='scrambler_onehot_to_logits')
scale_logits = Lambda(lambda x: x[1] * K.tile(x[0], (1, 1, 1, 20)), name='scrambler_logit_scale')
def _scrambler_func(sequence_input) :
conv_0_out = conv_0(sequence_input)
#Connect group of res blocks
output_tensor = conv_0_out
#Res block group execution
skip_conv_outs = []
for group_ix in range(n_groups) :
skip_conv_out = skip_convs[group_ix](output_tensor)
skip_conv_outs.append(skip_conv_out)
for layer_ix in range(n_resblocks_per_group) :
output_tensor = resblock_groups[group_ix][layer_ix](output_tensor)
#Last res block extr conv
last_block_conv_out = last_block_conv(output_tensor)
skip_add_out = last_block_conv_out
for group_ix in range(n_groups) :
skip_add_out = skip_add([skip_add_out, skip_conv_outs[group_ix]])
#Final conv out
final_conv_out = final_conv(skip_add_out)
#Scale logits by importance scores
scaled_logits = scale_logits([final_conv_out, onehot_to_logits(sequence_input)])
return scaled_logits, final_conv_out
return _scrambler_func
#Keras loss functions
def get_sigmoid_kl_divergence() :
def _kl_divergence(y_true, y_pred) :
y_true = K.clip(y_true, K.epsilon(), 1.0 - K.epsilon())
y_pred = K.clip(y_pred, K.epsilon(), 1.0 - K.epsilon())
return K.mean(y_true * K.log(y_true / y_pred) + (1.0 - y_true) * K.log((1.0 - y_true) / (1.0 - y_pred)), axis=-1)
return _kl_divergence
def get_margin_entropy_ame_masked(pwm_start, pwm_end) :
def _margin_entropy_ame_masked(pwm, pwm_mask, pwm_background, max_bits) :
conservation = pwm[:, 0, pwm_start:pwm_end, :] * K.log(K.clip(pwm[:, 0, pwm_start:pwm_end, :], K.epsilon(), 1. - K.epsilon()) / pwm_background[:, 0, pwm_start:pwm_end, :]) / K.log(2.0)
conservation = K.sum(conservation, axis=-1)
mask = K.max(pwm_mask[:, 0, pwm_start:pwm_end, :], axis=-1)
n_unmasked = K.sum(mask, axis=-1)
mean_conservation = K.sum(conservation * mask, axis=-1) / n_unmasked
margin_conservation = K.switch(mean_conservation > K.constant(max_bits[:, 0], shape=(1,)), mean_conservation - K.constant(max_bits, shape=(1,)), K.zeros_like(mean_conservation))
return margin_conservation
return _margin_entropy_ame_masked
def get_target_entropy_sme_masked(pwm_start, pwm_end) :
def _target_entropy_sme_masked(pwm, pwm_mask, pwm_background, target_bits) :
conservation = pwm[:, 0, pwm_start:pwm_end, :] * K.log(K.clip(pwm[:, 0, pwm_start:pwm_end, :], K.epsilon(), 1. - K.epsilon()) / pwm_background[:, 0, pwm_start:pwm_end, :]) / K.log(2.0)
conservation = K.sum(conservation, axis=-1)
mask = K.max(pwm_mask[:, 0, pwm_start:pwm_end, :], axis=-1)
n_unmasked = K.sum(mask, axis=-1)
mean_conservation = K.sum(conservation * mask, axis=-1) / n_unmasked
return (mean_conservation - target_bits[:, 0])**2
return _target_entropy_sme_masked
def get_weighted_loss(loss_coeff=1.) :
def _min_pred(y_true, y_pred) :
return loss_coeff * y_pred
return _min_pred
#Initialize Encoder and Decoder networks
batch_size = 32
seq_length = 81
n_samples = 32
sample_mode = 'gumbel'
#Resnet parameters
resnet_n_groups = 5
resnet_n_resblocks_per_group = 4
resnet_n_channels = 48
resnet_window_size = 3
resnet_dilation_rates = [1, 2, 4, 2, 1]
resnet_drop_rate = 0.0
#Load scrambler
scrambler = load_scrambler_network(
n_groups=resnet_n_groups,
n_resblocks_per_group=resnet_n_resblocks_per_group,
n_channels=resnet_n_channels, window_size=resnet_window_size,
dilation_rates=resnet_dilation_rates,
drop_rate=resnet_drop_rate
)
#Load sampler
sampler = build_sampler(batch_size, seq_length, n_classes=seq_length+1, n_samples=n_samples, sample_mode=sample_mode)
#Load predictor
predictor_path = 'saved_models/ppi_rnn_baker_big_set_5x_negatives_classifier_symmetric_drop_25_5x_negatives_balanced_partitioned_data_epoch_10.h5'
predictor = load_model(predictor_path, custom_objects={ 'sigmoid_nll' : get_sigmoid_kl_divergence() })
predictor.trainable = False
predictor.compile(loss='mean_squared_error', optimizer=keras.optimizers.SGD(lr=0.1))
#Build scrambler model
scrambler_class = Input(shape=(1,), name='scrambler_class')
scrambler_input = Input(shape=(1, seq_length, 20), name='scrambler_input')
scrambled_logits, importance_scores = scrambler(scrambler_input)
pwm_logits, pwm, sampled_pwm, _, sampled_mask = sampler(scrambler_class, scrambled_logits)
zeropad_layer = Lambda(lambda x: x[0] * x[1], name='zeropad')
sampled_pwm_zeropad = zeropad_layer([sampled_pwm, sampled_mask])
scrambler_model = Model([scrambler_input, scrambler_class], [pwm_logits, pwm, sampled_pwm_zeropad, importance_scores])
#Initialize Sequence Templates and Masks
initialize_sequence_templates(scrambler_model, encoder, sequence_templates, x_mean_logits)
scrambler_model.compile(
optimizer=keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999),
loss='mean_squared_error'
)
#Set target bits
conservation_target_bits = np.zeros(seq_length+1)
conservation_target_bits[:] = 0.25
conservation_target_bits = conservation_target_bits.tolist()
entropy_target_bits = np.zeros(seq_length+1)
entropy_target_bits[:] = 0.25
entropy_target_bits = entropy_target_bits.tolist()
#Helper function for setting sequence-length-specific parameters
def initialize_sequence_length_params(model, background_matrix_list, conservation_target_bits_list, entropy_target_bits_list) :
flat_background_matrix_list = []
flat_conservation_target_bits_list = []
flat_entropy_target_bits_list = []
for k in range(len(background_matrix_list)) :
flat_background_matrix_list.append(background_matrix_list[k].reshape(1, -1))
flat_conservation_target_bits_list.append(np.array([conservation_target_bits_list[k]]).reshape(1, -1))
flat_entropy_target_bits_list.append(np.array([entropy_target_bits_list[k]]).reshape(1, -1))
flat_background_matrix_list = np.concatenate(flat_background_matrix_list, axis=0)
flat_conservation_target_bits_list = np.concatenate(flat_conservation_target_bits_list, axis=0)
flat_entropy_target_bits_list = np.concatenate(flat_entropy_target_bits_list, axis=0)
model.get_layer('x_mean_dense').set_weights([flat_background_matrix_list])
model.get_layer('x_mean_dense').trainable = False
model.get_layer('conservation_target_bits_dense').set_weights([flat_conservation_target_bits_list])
model.get_layer('conservation_target_bits_dense').trainable = False
model.get_layer('entropy_target_bits_dense').set_weights([flat_entropy_target_bits_list])
model.get_layer('entropy_target_bits_dense').trainable = False
#Build Auto-scrambler pipeline
#Define model inputs
ae_scrambler_class_1 = Input(shape=(1,), name='ae_scrambler_class_1')
ae_scrambler_input_1 = Input(shape=(1, seq_length, 20), name='ae_scrambler_input_1')
ae_scrambler_class_2 = Input(shape=(1,), name='ae_scrambler_class_2')
ae_scrambler_input_2 = Input(shape=(1, seq_length, 20), name='ae_scrambler_input_2')
#ae_label_input = Input(shape=(1,), name='ae_label_input')
#Run encoder and decoder
_, scrambled_pwm_1, scrambled_sample_1, pwm_mask_1, sampled_mask_1 = sampler(ae_scrambler_class_1, scrambler(ae_scrambler_input_1)[0])
_, scrambled_pwm_2, scrambled_sample_2, pwm_mask_2, sampled_mask_2 = sampler(ae_scrambler_class_2, scrambler(ae_scrambler_input_2)[0])
zeropad_layer_1 = Lambda(lambda x: x[0] * x[1], name='zeropad_1')
zeropad_layer_2 = Lambda(lambda x: x[0] * x[1], name='zeropad_2')
scrambled_sample_1_zeropad = zeropad_layer_1([scrambled_sample_1, sampled_mask_1])
scrambled_sample_2_zeropad = zeropad_layer_2([scrambled_sample_2, sampled_mask_2])
#Define layer to deflate sample axis
deflate_scrambled_sample = Lambda(lambda x: K.reshape(x, (batch_size * n_samples, 1, seq_length, 20)), name='deflate_scrambled_sample')
#Deflate sample axis
scrambled_sample_deflated_1 = deflate_scrambled_sample(scrambled_sample_1_zeropad)
scrambled_sample_deflated_2 = deflate_scrambled_sample(scrambled_sample_2_zeropad)
#Make reference prediction on non-scrambled input sequence
collapse_input_layer_non_scrambled = Lambda(lambda x: x[:, 0, :, :], output_shape=(seq_length, 20))
collapsed_in_1_non_scrambled = collapse_input_layer_non_scrambled(ae_scrambler_input_1)
collapsed_in_2_non_scrambled = collapse_input_layer_non_scrambled(ae_scrambler_input_2)
y_pred_non_scrambled = predictor([collapsed_in_1_non_scrambled, collapsed_in_2_non_scrambled])#ae_label_input
#Make prediction on scrambled sequence samples
collapse_input_layer = Lambda(lambda x: x[:, 0, :, :], output_shape=(seq_length, 20))
collapsed_in_1 = collapse_input_layer(scrambled_sample_deflated_1)
collapsed_in_2 = collapse_input_layer(scrambled_sample_deflated_2)
y_pred_scrambled_deflated = predictor([collapsed_in_1, collapsed_in_2])
#Define layer to inflate sample axis
inflate_scrambled_prediction = Lambda(lambda x: K.reshape(x, (batch_size, n_samples)), name='inflate_scrambled_prediction')
#Inflate sample axis
y_pred_scrambled = inflate_scrambled_prediction(y_pred_scrambled_deflated)
#Cost function parameters
pwm_start = 0
pwm_end = 81
#Define background matrix embeddings and target bits
seq_reshape_layer = Reshape((1, seq_length, 20))
flatten_bit_layer = Reshape((1,))
x_mean_dense = Embedding(seq_length+1, seq_length * 20, embeddings_initializer='zeros', name='x_mean_dense')
conservation_target_bits_dense = Embedding(seq_length+1, 1, embeddings_initializer='zeros', name='conservation_target_bits_dense')
entropy_target_bits_dense = Embedding(seq_length+1, 1, embeddings_initializer='zeros', name='entropy_target_bits_dense')
x_mean_len_1 = seq_reshape_layer(x_mean_dense(ae_scrambler_class_1))
x_mean_len_2 = seq_reshape_layer(x_mean_dense(ae_scrambler_class_2))
conservation_target_bits_len_1 = flatten_bit_layer(conservation_target_bits_dense(ae_scrambler_class_1))
conservation_target_bits_len_2 = flatten_bit_layer(conservation_target_bits_dense(ae_scrambler_class_2))
entropy_target_bits_len_1 = flatten_bit_layer(entropy_target_bits_dense(ae_scrambler_class_1))
entropy_target_bits_len_2 = flatten_bit_layer(entropy_target_bits_dense(ae_scrambler_class_2))
#NLL cost
nll_loss_func = get_sigmoid_kl_divergence()
#Conservation cost
conservation_loss_func = get_target_entropy_sme_masked(pwm_start=pwm_start, pwm_end=pwm_end)
#Entropy cost
entropy_loss_func = get_target_entropy_sme_masked(pwm_start=pwm_start, pwm_end=pwm_end)
#entropy_loss_func = get_margin_entropy_ame_masked(pwm_start=pwm_start, pwm_end=pwm_end)
#Define annealing coefficient
anneal_coeff = K.variable(1.0)
#Execute NLL cost
nll_loss = Lambda(lambda x: nll_loss_func(K.tile(x[0], (1, K.shape(x[1])[1])), x[1]), name='nll')([
y_pred_non_scrambled,
y_pred_scrambled
])
#Execute conservation cost
conservation_loss = Lambda(lambda x: anneal_coeff * (0.5 * conservation_loss_func(x[0], x[1], x[2], x[3]) + 0.5 * conservation_loss_func(x[4], x[5], x[6], x[7])), name='conservation')([
scrambled_pwm_1,
pwm_mask_1,
x_mean_len_1,
conservation_target_bits_len_1,
scrambled_pwm_2,
pwm_mask_2,
x_mean_len_2,
conservation_target_bits_len_2
])
#Execute entropy cost
entropy_loss = Lambda(lambda x: (1. - anneal_coeff) * (0.5 * entropy_loss_func(x[0], x[1], x[2], x[3]) + 0.5 * entropy_loss_func(x[4], x[5], x[6], x[7])), name='entropy')([
scrambled_pwm_1,
pwm_mask_1,
x_mean_len_1,
entropy_target_bits_len_1,
scrambled_pwm_2,
pwm_mask_2,
x_mean_len_2,
entropy_target_bits_len_2
])
loss_model = Model(
[ae_scrambler_class_1, ae_scrambler_input_1, ae_scrambler_class_2, ae_scrambler_input_2], #ae_label_input
[nll_loss, conservation_loss, entropy_loss]
)
#Initialize Sequence Templates and Masks
initialize_sequence_templates(loss_model, encoder, sequence_templates, x_mean_logits)
#Initialize Sequence Length Parameters
initialize_sequence_length_params(loss_model, x_means, conservation_target_bits, entropy_target_bits)
loss_model.compile(
optimizer=keras.optimizers.Adam(lr=0.0001, beta_1=0.5, beta_2=0.9),
loss={
'nll' : get_weighted_loss(loss_coeff=1.0),
'conservation' : get_weighted_loss(loss_coeff=1.0),
'entropy' : get_weighted_loss(loss_coeff=10.0)
}
)
scrambler_model.summary()
loss_model.summary()
#Training configuration
#Define number of training epochs
n_epochs = 20
#Define experiment suffix (optional)
experiment_suffix = "_kl_divergence_zeropad_gumbel"
#Define anneal function
def _anneal_func(val, epoch, n_epochs=n_epochs) :
if epoch in [0] :
return 1.0
return 0.0
architecture_str = "resnet_" + str(resnet_n_groups) + "_" + str(resnet_n_resblocks_per_group) + "_" + str(resnet_n_channels) + "_" + str(resnet_window_size) + "_" + str(resnet_drop_rate).replace(".", "")
model_name = "autoscrambler_dataset_" + dataset_name + "_sample_mode_" + sample_mode + "_n_samples_" + str(n_samples) + "_" + architecture_str + "_n_epochs_" + str(n_epochs) + "_target_bits_" + str(entropy_target_bits[0]).replace(".", "") + experiment_suffix
print("Model save name = " + model_name)
#Execute training procedure
callbacks =[
#ModelCheckpoint("model_checkpoints/" + model_name + "_epoch_{epoch:02d}.hdf5", monitor='val_loss', mode='min', period=10, save_weights_only=True),
EpochVariableCallback(anneal_coeff, _anneal_func)
]
s_train = np.zeros((x_1_train.shape[0], 1))
s_test = np.zeros((x_1_test.shape[0], 1))
# train the autoencoder
train_history = loss_model.fit(
[l_1_train, x_1_train, l_2_train, x_2_train], #y_train
[s_train, s_train, s_train],
shuffle=True,
epochs=n_epochs,
batch_size=batch_size,
validation_data=(
[l_1_test, x_1_test, l_2_test, x_2_test], #y_test
[s_test, s_test, s_test]
),
callbacks=callbacks
)
f, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(3 * 4, 3))
n_epochs_actual = len(train_history.history['nll_loss'])
ax1.plot(np.arange(1, n_epochs_actual + 1), train_history.history['nll_loss'], linewidth=3, color='green')
ax1.plot(np.arange(1, n_epochs_actual + 1), train_history.history['val_nll_loss'], linewidth=3, color='orange')
plt.sca(ax1)
plt.xlabel("Epochs", fontsize=14)
plt.ylabel("NLL", fontsize=14)
plt.xlim(1, n_epochs_actual)
plt.xticks([1, n_epochs_actual], [1, n_epochs_actual], fontsize=12)
plt.yticks(fontsize=12)
ax2.plot(np.arange(1, n_epochs_actual + 1), train_history.history['entropy_loss'], linewidth=3, color='green')
ax2.plot(np.arange(1, n_epochs_actual + 1), train_history.history['val_entropy_loss'], linewidth=3, color='orange')
plt.sca(ax2)
plt.xlabel("Epochs", fontsize=14)
plt.ylabel("Entropy Loss", fontsize=14)
plt.xlim(1, n_epochs_actual)
plt.xticks([1, n_epochs_actual], [1, n_epochs_actual], fontsize=12)
plt.yticks(fontsize=12)
ax3.plot(np.arange(1, n_epochs_actual + 1), train_history.history['conservation_loss'], linewidth=3, color='green')
ax3.plot(np.arange(1, n_epochs_actual + 1), train_history.history['val_conservation_loss'], linewidth=3, color='orange')
plt.sca(ax3)
plt.xlabel("Epochs", fontsize=14)
plt.ylabel("Conservation Loss", fontsize=14)
plt.xlim(1, n_epochs_actual)
plt.xticks([1, n_epochs_actual], [1, n_epochs_actual], fontsize=12)
plt.yticks(fontsize=12)
plt.tight_layout()
plt.show()
# Save model and weights
save_dir = 'saved_models'
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
model_path = os.path.join(save_dir, model_name + '.h5')
scrambler_model.save(model_path)
print('Saved scrambler model at %s ' % (model_path))
#Load models
save_dir = 'saved_models'
#model_name = "autoscrambler_dataset_coiled_coil_binders_inverted_scores_sample_mode_st_n_samples_32_resnet_5_4_48_3_00_n_epochs_20_target_bits_24_kl_divergence_log_prob"
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
model_path = os.path.join(save_dir, model_name + '.h5')
scrambler_model = load_model(model_path, custom_objects={
'gumbel_softmax' : gumbel_softmax
})
print('Loaded scrambler model %s ' % (model_path))
#Visualize a few reconstructed sequence patterns
logits_test, pwm_test, sample_test, importance_scores = scrambler_model.predict_on_batch(x=[x_1_test[:32], l_1_test[:32]])
subtracted_logits_test = (2. * np.array(x_1_test[:32], dtype=np.float64) - 1.) * np.maximum(np.array(importance_scores, dtype=np.float64), 1e-7)
subtracted_pwm_test = np.exp(subtracted_logits_test) / np.expand_dims(np.sum(np.exp(subtracted_logits_test), axis=-1), axis=-1)
for plot_i in range(0, 5) :
print("Test sequence " + str(plot_i) + ":")
plot_protein_logo(residue_map, x_1_test[plot_i, 0, :, :], sequence_template=sequence_templates[l_1_test[plot_i, 0]], figsize=(12, 1), plot_start=0, plot_end=96)
plot_protein_logo(residue_map, pwm_test[plot_i, 0, :, :], sequence_template=sequence_templates[l_1_test[plot_i, 0]], figsize=(12, 1), plot_start=0, plot_end=96)
plot_protein_logo(residue_map, subtracted_pwm_test[plot_i, 0, :, :], sequence_template=sequence_templates[l_1_test[plot_i, 0]], figsize=(12, 1), plot_start=0, plot_end=96)
#Binder DHD_154
#seq_1 = ("TAEELLEVHKKSDRVTKEHLRVSEEILKVVEVLTRGEVSSEVLKRVLRKLEELTDKLRRVTEEQRRVVEKLN" + "#" * seq_length)[:81]
#seq_2 = ("DLEDLLRRLRRLVDEQRRLVEELERVSRRLEKAVRDNEDERELARLSREHSDIQDKHDKLAREILEVLKRLLERTE" + "#" * seq_length)[:81]
seq_1 = "TAEELLEVHKKSDRVTKEHLRVSEEILKVVEVLTRGEVSSEVLKRVLRKLEELTDKLRRVTEEQRRVVEKLN"[:81]
seq_2 = "DLEDLLRRLRRLVDEQRRLVEELERVSRRLEKAVRDNEDERELARLSREHSDIQDKHDKLAREILEVLKRLLERTE"[:81]
print("Seq 1 = " + seq_1)
print("Seq 2 = " + seq_2)
encoder = IdentityEncoder(81, residue_map)
test_onehot_1 = np.tile(np.expand_dims(np.expand_dims(encoder(seq_1), axis=0), axis=0), (batch_size, 1, 1, 1))
test_onehot_2 = np.tile(np.expand_dims(np.expand_dims(encoder(seq_2), axis=0), axis=0), (batch_size, 1, 1, 1))
test_len_1 = np.tile(np.array([[len(seq_1)]]), (batch_size, 1))
test_len_2 = np.tile(np.array([[len(seq_2)]]), (batch_size, 1))
pred_interacts = predictor.predict(x=[test_onehot_1[:, 0, ...], test_onehot_2[:, 0, ...]])[0, 0]
print("Predicted interaction prob = " + str(round(pred_interacts, 4)))
#Visualize a few reconstructed sequence patterns
save_figs = False
pair_name = "DHD_154"
logits_test_1, pwm_test_1, sample_test_1, importance_scores_1 = scrambler_model.predict_on_batch(x=[test_onehot_1, test_len_1])
logits_test_2, pwm_test_2, sample_test_2, importance_scores_2 = scrambler_model.predict_on_batch(x=[test_onehot_2, test_len_2])
scrambled_pred_interacts = predictor.predict(x=[sample_test_1[0, :, 0, ...], sample_test_2[0, :, 0, ...]])[:, 0]
print("Scrambler predictions = " + str(np.round(scrambled_pred_interacts[:10], 2)))
def get_subtracted_pwm(test_onehot, importance_scores_test) :
subtracted_logits_test = (2. * np.array(test_onehot, dtype=np.float64) - 1.) * np.maximum(np.array(importance_scores_test, dtype=np.float64), 1e-7)
subtracted_pwm_test = np.exp(subtracted_logits_test) / np.expand_dims(np.sum(np.exp(subtracted_logits_test), axis=-1), axis=-1)
return subtracted_pwm_test
subtracted_pwm_test_1 = get_subtracted_pwm(test_onehot_1, importance_scores_1)
subtracted_pwm_test_2 = get_subtracted_pwm(test_onehot_2, importance_scores_2)
print("Binder 1:")
plot_protein_logo(residue_map, test_onehot_1[0, 0, :, :], sequence_template=sequence_templates[test_len_1[0, 0]], figsize=(12, 1), plot_start=0, plot_end=96, save_figs=save_figs, fig_name=model_name + "_original_example_" + pair_name + "_binder_1")
plot_protein_logo(residue_map, pwm_test_1[0, 0, :, :], sequence_template=sequence_templates[test_len_1[0, 0]], figsize=(12, 1), plot_start=0, plot_end=96, save_figs=save_figs, fig_name=model_name + "_scrambled_example_" + pair_name + "_binder_1")
plot_protein_logo(residue_map, subtracted_pwm_test_1[0, 0, :, :], sequence_template=sequence_templates[test_len_1[0, 0]], figsize=(12, 1), plot_start=0, plot_end=96, save_figs=save_figs, fig_name=model_name + "_subtracted_example_" + pair_name + "_binder_1")
print("Binder 2:")
plot_protein_logo(residue_map, test_onehot_2[0, 0, :, :], sequence_template=sequence_templates[test_len_2[0, 0]], figsize=(12, 1), plot_start=0, plot_end=96, save_figs=save_figs, fig_name=model_name + "_original_example_" + pair_name + "_binder_2")
plot_protein_logo(residue_map, pwm_test_2[0, 0, :, :], sequence_template=sequence_templates[test_len_2[0, 0]], figsize=(12, 1), plot_start=0, plot_end=96, save_figs=save_figs, fig_name=model_name + "_scrambled_example_" + pair_name + "_binder_2")
plot_protein_logo(residue_map, subtracted_pwm_test_2[0, 0, :, :], sequence_template=sequence_templates[test_len_2[0, 0]], figsize=(12, 1), plot_start=0, plot_end=96, save_figs=save_figs, fig_name=model_name + "_subtracted_example_" + pair_name + "_binder_2")
#Binder DHD_154
test_onehot_1 = np.tile(np.expand_dims(np.expand_dims(encoder(seq_1), axis=0), axis=0), (batch_size, 1, 1, 1))
test_onehot_2 = np.tile(np.expand_dims(np.expand_dims(encoder(seq_2), axis=0), axis=0), (batch_size, 1, 1, 1))
test_len_1 = np.tile(np.array([[len(seq_1)]]), (batch_size, 1))
test_len_2 = np.tile(np.array([[len(seq_2)]]), (batch_size, 1))
bg = np.tile(np.expand_dims(np.expand_dims(np.concatenate([
x_means[test_len_1[0, 0]],
x_means[test_len_2[0, 0]]
], axis=0), axis=0), axis=0), (batch_size, 1, 1, 1))
seq_mask = np.concatenate([
np.max(test_onehot_1[0, 0, ...], axis=-1, keepdims=True),
np.max(test_onehot_2[0, 0, ...], axis=-1, keepdims=True)
], axis=0)
x_curr = np.concatenate([test_onehot_1, test_onehot_2], axis=2)[0, 0, ...]
bg_curr = bg[0, 0, ...]
importance_scores = np.concatenate([importance_scores_1[:1, ...], importance_scores_2[:1, ...]], axis=2)
importance_scores = importance_scores * seq_mask
n_imp = len(np.nonzero(importance_scores > 1.0)[0])
q_imp = 1. - n_imp / x_curr.shape[0]
print("Quantile = " + str(round(q_imp, 3)) + " (" + str(n_imp) + " deemed important).")
chosen_importance_scores = np.zeros(importance_scores.shape)
chosen_importance_scores[importance_scores >= np.quantile(importance_scores, q=q_imp)] = 1.
x_curr[np.sum(chosen_importance_scores, axis=(0, 1, 3)) <= 0.,:] = -1
def _mask_and_template_proper(onehot, bg) :
indicator = np.min(onehot, axis=-1)
sampled_mask = np.ones(onehot.shape)
sampled_template = np.zeros(onehot.shape)
for j in range(indicator.shape[0]) :
if indicator[j] == -1 :
sampled_mask[j, :] = 0.
sampled_ix = np.random.choice(np.arange(20), p=bg[j, :])
sampled_template[j, sampled_ix] = 1.
new_onehot = onehot * sampled_mask + sampled_template
return new_onehot
sample_curr = np.expand_dims(np.expand_dims(_mask_and_template_proper(x_curr, bg_curr), axis=0), axis=0)
sample_curr = sample_curr * np.expand_dims(np.expand_dims(seq_mask, axis=0), axis=0)
pred_interacts = predictor.predict(x=[sample_curr[:, 0, :81, ...], sample_curr[:, 0, 81:, ...]])[0, 0]
print("Predicted interaction prob = " + str(round(pred_interacts, 4)))
#Re-do test a number of times
n_test_samples = 1000
pred_interacts = []
for i in range(n_test_samples) :
sample_curr = np.expand_dims(np.expand_dims(_mask_and_template_proper(x_curr, bg_curr), axis=0), axis=0)
sample_curr = sample_curr * np.expand_dims(np.expand_dims(seq_mask, axis=0), axis=0)
pred_interacts.append(predictor.predict(x=[sample_curr[:, 0, :81, ...], sample_curr[:, 0, 81:, ...]])[0, 0])
pred_interacts = np.array(pred_interacts)
#Plot distribution of binding predictions on samples
target_prob = 0.8533
mean_kl = target_prob * np.log(target_prob / pred_interacts) + (1. - target_prob) * np.log((1. - target_prob) / (1. - pred_interacts))
print("Mean predited prob = " + str(round(np.mean(pred_interacts), 3)))
print("Mean KL = " + str(round(np.mean(mean_kl), 3)))
f = plt.figure(figsize=(6, 4))
plt.hist(pred_interacts, bins=50, edgecolor='black', color='red', linewidth=2)
plt.xlabel("Predicted Binding Prob.", fontsize=12)
plt.ylabel("Sample Count", fontsize=12)
plt.xticks(fontsize=12, rotation=45)
plt.yticks(fontsize=12)
plt.xlim(0, 1)
plt.ylim(0)
plt.tight_layout()
plt.show()
#Re-load cached dataframe (shuffled)
dataset_name = "coiled_coil_binders"
experiment = "coiled_coil_binders_alyssa"
data_df = pd.read_csv(experiment + ".csv", sep="\t")
print("len(data_df) = " + str(len(data_df)))
test_df = data_df.copy().reset_index(drop=True)
batch_size = 32
test_df = test_df.iloc[:(len(test_df) // batch_size) * batch_size].copy().reset_index(drop=True)
print("len(test_df) = " + str(len(test_df)))
print(test_df.head())
#Construct test data
batch_size = 32
test_gen = iso.DataGenerator(
np.arange(len(test_df), dtype=np.int),
{ 'df' : test_df },
batch_size=(len(test_df) // batch_size) * batch_size,
inputs = [
{
'id' : 'amino_seq_1',
'source_type' : 'dataframe',
'source' : 'df',
#'extractor' : lambda row, index: (row['amino_seq_1'] + "#" * seq_length)[:seq_length],
'extractor' : lambda row, index: row['amino_seq_1'],
'encoder' : IdentityEncoder(seq_length, residue_map),
'dim' : (1, seq_length, len(residue_map)),
'sparsify' : False
},
{
'id' : 'amino_seq_2',
'source_type' : 'dataframe',
'source' : 'df',
#'extractor' : lambda row, index: row['amino_seq_2'] + "#" * seq_length)[:seq_length],
'extractor' : lambda row, index: row['amino_seq_2'],
'encoder' : IdentityEncoder(seq_length, residue_map),
'dim' : (1, seq_length, len(residue_map)),
'sparsify' : False
},
{
'id' : 'amino_seq_1_len',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index: len(row['amino_seq_1']),
'encoder' : lambda t: t,
'dim' : (1,),
'sparsify' : False
},
{
'id' : 'amino_seq_2_len',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index: len(row['amino_seq_2']),
'encoder' : lambda t: t,
'dim' : (1,),
'sparsify' : False
}
],
outputs = [
{
'id' : 'interacts',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index: row['interacts'],
'transformer' : NopTransformer(1),
'dim' : (1,),
'sparsify' : False
}
],
randomizers = [],
shuffle = False
)
#Load data matrices
[x_1_test, x_2_test, l_1_test, l_2_test], [y_test] = test_gen[0]
print("x_1_test.shape = " + str(x_1_test.shape))
print("x_2_test.shape = " + str(x_2_test.shape))
print("l_1_test.shape = " + str(l_1_test.shape))
print("l_2_test.shape = " + str(l_2_test.shape))
print("y_test.shape = " + str(y_test.shape))
#Predict on test set
_, _, sample_test_1, importance_scores_1 = scrambler_model.predict(x=[x_1_test, l_1_test], batch_size=32, verbose=True)
_, _, sample_test_2, importance_scores_2 = scrambler_model.predict(x=[x_2_test, l_2_test], batch_size=32, verbose=True)
unscrambled_preds = predictor.predict(x=[x_1_test[:, 0, ...], x_2_test[:, 0, ...]], batch_size=32, verbose=True)[:, 0]
scrambled_preds = []
for i in range(sample_test_1.shape[0]) :
if i % 100 == 0 :
print("Predicting scrambled samples for sequence " + str(i) + "...")
scrambled_pred_samples = predictor.predict(x=[sample_test_1[i, :, 0, ...], sample_test_2[i, :, 0, ...]], batch_size=32, verbose=False)[:, 0]
scrambled_preds.append(np.mean(scrambled_pred_samples))
scrambled_preds = np.array(scrambled_preds)
min_val = 0.0
max_val = 1.0
max_y_val = 8
n_bins = 25
save_figs = False
figsize = (6, 4)
measurements = [
unscrambled_preds,
scrambled_preds
]
colors = [
'green',
'red'
]
labels = [
'Unscrambled',
'Scrambled'
]
x_label = 'Prediction'
y_label = 'Density'
min_hist_val = np.min(measurements[0])
max_hist_val = np.max(measurements[0])
for i in range(1, len(measurements)) :
min_hist_val = min(min_hist_val, np.min(measurements[i]))
max_hist_val = max(max_hist_val, np.max(measurements[i]))
if min_val is not None :
min_hist_val = min_val
if max_val is not None :
max_hist_val = max_val
hists = []
bin_edges = []
means = []
for i in range(len(measurements)) :
hist, b_edges = np.histogram(measurements[i], range=(min_hist_val, max_hist_val), bins=n_bins, density=True)
hists.append(hist)
bin_edges.append(b_edges)
means.append(np.mean(measurements[i]))
bin_width = bin_edges[0][1] - bin_edges[0][0]
#Compare Log Likelihoods
f = plt.figure(figsize=figsize)
for i in range(len(measurements)) :
plt.bar(bin_edges[i][1:] - bin_width/2., hists[i], width=bin_width, linewidth=2, alpha=0.5, edgecolor='black', color=colors[i], label=labels[i])
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.xlim(min_hist_val, max_hist_val)
if max_y_val is not None :
plt.ylim(0, max_y_val)
plt.xlabel(x_label, fontsize=14)
plt.ylabel(y_label, fontsize=14)
for i in range(len(measurements)) :
plt.axvline(x=means[i], linewidth=2, color=colors[i], linestyle="--")
plt.legend(fontsize=14, loc='upper left')
plt.tight_layout()
if save_figs :
fig_name = experiment + "_model_" + model_name + "_pos_hist"
plt.savefig(fig_name + ".png", dpi=300, transparent=True)
plt.savefig(fig_name + ".eps")
plt.show()
#Store unscrambled and scrambled binding predictions
test_df['pred_interacts'] = np.round(unscrambled_preds, 2)
test_df['pred_interacts_scrambled'] = np.round(scrambled_preds, 2)
flat_importance_scores_1 = importance_scores_1[:, 0, :, 0]
flat_importance_scores_2 = importance_scores_2[:, 0, :, 0]
short_model_name = "inclusion_target_bits_" + str(entropy_target_bits[0]).replace(".", "") + "_epochs_" + str(n_epochs) + experiment_suffix
test_df.to_csv(experiment + "_model_" + short_model_name + "_testset.csv", sep="\t", index=False)
np.save(experiment + "_model_" + short_model_name + "_testset_importance_scores_1", flat_importance_scores_1)
np.save(experiment + "_model_" + short_model_name + "_testset_importance_scores_2", flat_importance_scores_2)
```
| github_jupyter |
## Setup a classification experiment
```
import pandas as pd
from sklearn.datasets import load_boston
from sklearn.model_selection import train_test_split
df = pd.read_csv(
"https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data",
header=None)
df.columns = [
"Age", "WorkClass", "fnlwgt", "Education", "EducationNum",
"MaritalStatus", "Occupation", "Relationship", "Race", "Gender",
"CapitalGain", "CapitalLoss", "HoursPerWeek", "NativeCountry", "Income"
]
train_cols = df.columns[0:-1]
label = df.columns[-1]
X = df[train_cols]
y = df[label].apply(lambda x: 0 if x == " <=50K" else 1) #Turning response into 0 and 1
# We have to transform categorical variables to use sklearn models
X_enc = pd.get_dummies(X, prefix_sep='.')
feature_names = list(X_enc.columns)
seed = 1
X_train, X_test, y_train, y_test = train_test_split(X_enc, y, test_size=0.20, random_state=seed)
```
## Train a blackbox classification system
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.decomposition import PCA
from sklearn.pipeline import Pipeline
#Blackbox system can include preprocessing, not just a classifier!
pca = PCA()
rf = RandomForestClassifier(n_estimators=100, n_jobs=-1)
blackbox_model = Pipeline([('pca', pca), ('rf', rf)])
blackbox_model.fit(X_train, y_train)
```
## Show blackbox model performance
```
from interpret import show
from interpret.perf import ROC
blackbox_perf = ROC(blackbox_model.predict_proba).explain_perf(X_test, y_test, name='Blackbox')
show(blackbox_perf)
```
## Local Explanations: How an individual prediction was made
```
from interpret.blackbox import LimeTabular
from interpret import show
#Blackbox explainers need a predict function, and optionally a dataset
lime = LimeTabular(predict_fn=blackbox_model.predict_proba, data=X_train, random_state=1)
#Pick the instances to explain, optionally pass in labels if you have them
lime_local = lime.explain_local(X_test[:5], y_test[:5], name='LIME')
show(lime_local)
from interpret.blackbox import ShapKernel
import numpy as np
background_val = np.median(X_train, axis=0).reshape(1, -1)
shap = ShapKernel(predict_fn=blackbox_model.predict_proba, data=background_val, feature_names=feature_names)
shap_local = shap.explain_local(X_test[:5], y_test[:5], name='SHAP')
show(shap_local)
```
## Global Explanations: How the model behaves overall
```
from interpret.blackbox import MorrisSensitivity
sensitivity = MorrisSensitivity(predict_fn=blackbox_model.predict_proba, data=X_train)
sensitivity_global = sensitivity.explain_global(name="Global Sensitivity")
show(sensitivity_global)
from interpret.blackbox import PartialDependence
pdp = PartialDependence(predict_fn=blackbox_model.predict_proba, data=X_train)
pdp_global = pdp.explain_global(name='Partial Dependence')
show(pdp_global)
```
## Compare them all in the Dashboard
```
show([blackbox_perf, lime_local, shap_local, sensitivity_global, pdp_global])
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Visualization/ndwi_symbology.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Visualization/ndwi_symbology.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Visualization/ndwi_symbology.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://geemap.org). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('Installing geemap ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
import ee
import geemap
```
## Create an interactive map
The default basemap is `Google Maps`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/basemaps.py) can be added using the `Map.add_basemap()` function.
```
Map = geemap.Map(center=[40,-100], zoom=4)
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
# This function gets NDVI from Landsat 5 imagery.
def getNDWI(image):
return image.normalizedDifference(['B3', 'B5'])
image1 = ee.Image('LANDSAT/LT05/C01/T1_TOA/LT05_044034_19900604')
# Compute NDVI from the scene.
ndvi1 = getNDWI(image1)
ndwiParams = {'palette': ['#ece7f2', '#d0d1e6', '#a6bddb', '#74a9cf', '#3690c0', '#0570b0', '#045a8d', '#023858']}
Map.centerObject(image1, 10)
Map.addLayer(ndvi1, ndwiParams, 'NDWI')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
```
import sys, glob, os
SPARK_HOME=os.environ['SPARK_HOME']
sys.path.append(SPARK_HOME + "/python")
sys.path.append(glob.glob(SPARK_HOME + "/python/lib/py4j*.zip")[0])
from pyspark import SparkConf
from pyspark.sql import SparkSession
from pyspark.sql.functions import *
from pyspark.sql.window import Window, WindowSpec
conf = (SparkConf()
.setAppName("PySpark Application")
.setIfMissing("spark.master", "local[*]")
.setIfMissing("spark.local.dir", "/tmp/spark")
.setIfMissing("spark.driver.memory", "5G")
.setIfMissing("spark.driver.cores", "4")
.setIfMissing("spark.jars.packages", "graphframes:graphframes:0.7.0-spark2.4-s_2.11")
)
spark = SparkSession.builder.config(conf = conf).enableHiveSupport().getOrCreate()
sc = spark.sparkContext
print(sc.uiWebUrl)
sql = spark.sql
from graphframes import GraphFrame
vertices = spark.createDataFrame([('1', 'Carter', 'Derrick', 50),
('2', 'May', 'Derrick', 26),
('3', 'Mills', 'Jeff', 80),
('4', 'Hood', 'Robert', 65),
('5', 'Banks', 'Mike', 93),
('98', 'Berg', 'Tim', 28),
('99', 'Page', 'Allan', 16)],
['id', 'name', 'firstname', 'age'])
edges = spark.createDataFrame([('1', '2', 'friend'),
('2', '1', 'friend'),
('3', '1', 'friend'),
('1', '3', 'friend'),
('2', '3', 'follows'),
('3', '4', 'friend'),
('4', '3', 'friend'),
('5', '3', 'friend'),
('3', '5', 'friend'),
('4', '5', 'follows'),
('98', '99', 'friend'),
('99', '98', 'friend')],
['src', 'dst', 'type'])
g = GraphFrame(vertices, edges)
## Take a look at the DataFrames
print("Veritices")
g.vertices.show()
print("Edges")
g.edges.show()
print("Degrees")
g.degrees.show()
nodes = list(map(lambda r: r.id, g.vertices.select("id").collect()))
nodes
edges = [(r.src, r.dst) for r in g.edges.select("src", "dst").collect()]
edges
```
# Drawing graph using networkx
```
import networkx as nx
G = nx.Graph()
G.add_nodes_from(nodes)
G.add_edges_from(edges)
options = {
'node_color': 'steelblue',
"font_color": "white",
'node_size': 1000,
'width': 3,
'font_weight': 'bold'
}
nx.draw_shell(G, with_labels=True, **options)
edges.where("src<dst").show()
g.edges.filter('type == "friend"').show()
sc.setCheckpointDir("/tmp/spark-checkpoint")
g.connectedComponents().show()
```
# Motif finding
Finding motifs helps to execute queries to discover structural patterns in graphs. Network motifs are patterns that occur repeatedly in the graph and represent the relationships between the vertices. GraphFrames motif finding uses a declarative Domain Specific Language (DSL) for expressing structural queries.
The query can be invoked by using the find-function, where the motif (in quotation marks) is expressed as the first parameter of the function.
The following example will search for pairs of vertices a,b connected by edge e and pairs of vertices b,c connected by edge e2. It will return a DataFrame of all such structures in the graph, with columns for each of the named elements (vertices or edges) in the motif.
```
g.find("(a)-[e]->(b); (b)-[e2]->(c)").show()
mutualFriends = (g
.find("(a)-[]->(b); (b)-[]->(c); (c)-[]->(b); (b)-[]->(a)")
.dropDuplicates()
)
mutualFriends.show(100, False)
mutualFriends.printSchema()
```
To query all the mutual friends between 2 and 3 we can filter the DataFrame.
```
mutualFriends.filter('a.id == 2 and c.id == 3').show(10, False)
```
# Triangle Count
```
g.triangleCount().show()
```
# Page Rank
```
pr = g.pageRank(resetProbability=0.15, tol=0.01)
## look at the pagerank score for every vertex
pr.vertices.show()
## look at the weight of every edge
pr.edges.show()
```
# Shortest Path
```
g.shortestPaths(landmarks=["1", "5"]).show()
```
| github_jupyter |
# pinkfish-challenge
Buy on the close on the SAME day a new 20 day high is set
```
# use future imports for python 3.x forward compatibility
from __future__ import print_function
from __future__ import unicode_literals
from __future__ import division
from __future__ import absolute_import
# other imports
import pandas as pd
import matplotlib.pyplot as plt
import datetime
from talib.abstract import *
# project imports
import pinkfish as pf
# format price data
pd.options.display.float_format = '{:0.2f}'.format
%matplotlib inline
symbol = '^GSPC'
#symbol = 'SPY'
capital = 10000
start = datetime.datetime(2016, 1, 1)
end = datetime.datetime.now()
```
Define high trade periods
```
period = 20
```
Define Strategy Class
```
class Strategy(object):
def __init__(self, symbol, capital, start, end, period,
slippage_per_trade=0, commissions_per_trade=0):
self._symbol = symbol
self._capital = capital
self._start = start
self._end = end
self._period = period
self._slippage_per_trade = slippage_per_trade
self._commissions_per_trade = commissions_per_trade
def _algo(self):
self._tlog.cash = self._capital
start_flag = True
end_flag = False
for i, row in enumerate(self._ts.itertuples()):
date = row.Index.to_pydatetime()
high = row.high
low = row.low
close = row.close
period_high = row.period_high
end_flag = True if (i == len(self._ts) - 1) else False
trade_state = None
if date < self._start:
continue
elif start_flag:
start_flag = False
# set start and end
self._start = date
self._end = self._ts.index[-1]
# buy
if (self._tlog.num_open_trades() == 0
and high > period_high
and not end_flag):
# enter buy in trade log
shares = self._tlog.enter_trade(date, close)
trade_state = pf.TradeState.OPEN
print("{0} BUY {1} {2} @ {3:.2f}".format(
date, shares, self._symbol, close))
# sell
elif end_flag:
# enter sell in trade log
shares = self._tlog.exit_trade(date, close)
trade_state = pf.TradeState.CLOSE
print("{0} SELL {1} {2} @ {3:.2f}".format(
date, shares, self._symbol, close))
# hold
else:
trade_state = pf.TradeState.HOLD
# record daily balance
self._dbal.append(date, high, low, close,
self._tlog.shares, self._tlog.cash,
trade_state)
def run(self):
self._ts = pf.fetch_timeseries(self._symbol)
self._ts = pf.select_tradeperiod(self._ts, self._start,
self._end, use_adj=False)
# Add technical indicator: X day high, and X day low
period_high = pd.Series(self._ts.close).rolling(self._period).max()
self._ts['period_high'] = period_high
self._tlog = pf.TradeLog()
self._dbal = pf.DailyBal()
self._algo()
def get_logs(self):
""" return DataFrames """
tlog = self._tlog.get_log()
dbal = self._dbal.get_log()
return tlog, dbal
```
Run Strategy
```
s = Strategy(symbol, capital, start, end, int(period))
s.run()
s._ts['2016-02-01':'2016-03-01']
```
Retrieve log DataFrames
```
s.tlog, s.dbal = s.get_logs()
s.tlog.tail(100)
```
The first 20 day high occurred on 2/22/2016, so we bought on the close that day.
We held onto the present, then sold the positon.
| github_jupyter |
# Tutorial
This is a very basic tutorial of segmentation and reconstruction in SEM. Here, we use a simple 2-d embedding space as it is easy to visualize. For the purpose of this tutorial, we do not consider structured embedding space (the HRR).
```
# ## un-comment out if running locally
# import os
# os.chdir('../')
## if running locally, comment out the following code
!git clone https://github.com/nicktfranklin/SEM.git
import os
os.chdir('./SEM/')
!pip install tensorflow==1.9
!pip install keras==2.2
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import seaborn as sns
from models import SEM
from sklearn import metrics
sns.set_context('talk')
```
Here, we create a toy data set: 2 events, both linear systems in a simple 2-D space with different ammounts of noise
```
np.random.seed(0) # for consistency, set the seed
def rotation_matrix(theta):
return np.array([[np.cos(theta), -np.sin(theta)], [np.sin(theta), np.cos(theta)]])
A = rotation_matrix( -np.pi/2 * 0.075)
x_train = [ np.array([[-1, 1]]).T / np.sqrt(2)]
for _ in range(19):
x_train.append(np.matmul(A, x_train[-1]))
A = rotation_matrix( np.pi/2 * 0.055)
x_train.append(np.array([[-1, -1]]).T / np.sqrt(2))
for _ in range(19):
x_train.append(np.matmul(A, x_train[-1]))
x_train = np.concatenate(x_train, axis=1).T
# add observation noise
x_train[:20, :] += np.random.randn(20, 2) * 0.04
x_train[20:, :] += np.random.randn(20, 2) * 0.12
plt.figure(figsize=(4,4))
plt.scatter(x_train[:, 0], x_train[:, 1])
plt.plot(x_train[:, 0], x_train[:, 1], alpha=0.25)
# label of the true identies
y = np.concatenate([np.zeros(20), np.ones(20)])
```
# Segmentation
Below, we set the parameters for segmentation. We generally only need to change the sticky-CRP parameters and the prior variance of the event model. The rest is set behind the scenes. Of the two, the prior variance is typically the most imporant
```
lmda = 10.0 # stickyness parameter
alfa = .01 # concentration parameter
# prior over the event varaiance.
var_scale = .6
var_df = 10
# These are parameters of the event model, not SEM, and they are stored in a seperate dictionary
f_opts = dict(var_scale0=var_scale, var_df0=var_df)
# store all of the SEM parameters and the event model parameters in a dictionary
sem_kwargs = dict(lmda=lmda, alfa=alfa, f_opts=f_opts)
help(SEM)
# run segmentation
sem_model = SEM(**sem_kwargs)
post = sem_model.run(x_train)
# plot results
def plot_segmentation(post, y):
cluster_id = np.argmax(post, axis=1)
cc = sns.color_palette('Dark2', post.shape[1])
fig, axes = plt.subplots(1, 2, figsize=(12, 4), gridspec_kw=dict(width_ratios=[1, 2]))
for clt in cluster_id:
idx = np.nonzero(cluster_id == clt)[0]
axes[0].scatter(x_train[idx, 0], x_train[idx, 1], color=cc[clt], alpha=.5)
axes[0].set_xlabel(r'$\mathbf{x}_{s,1}$')
axes[0].set_ylabel(r'$\mathbf{x}_{s,2}$')
axes[1].plot(post)
y_hat = np.argmax(post, axis=1)
axes[1].set_xlabel('Time')
axes[1].set_ylabel('Cluster ID')
plot_segmentation(post, y)
y_hat = np.argmax(post, axis=1)
print "Adjusted Mutual Info:", metrics.adjusted_mutual_info_score(y, y_hat)
print "Adjusted Rand Score: ", metrics.adjusted_rand_score(y, y_hat)
print
print y_hat
```
# Memory
----
Now, we move on to some simple demonstrations of memory smoothing
## Created corrupted memory trace
The software has a buildin function for generating corrupted memory traces consistent with the model
```
from models.memory import create_corrupted_trace
help(create_corrupted_trace)
epsilon_e = 0.25 # event label precision (1 - zero/one loss probability)
tau = 0.08 # feature corruption noise (Guassian)
b = 2 # temporal corruption noise (uniform)
noise_kwargs = dict(tau=tau, epsilon_e=epsilon_e, b=b)
e_true = y_hat # event labels as expereinced by SEM
y_mem = create_corrupted_trace(x_train, e_true, **noise_kwargs)
```
This is a list of corrupted memory traces. Each trace is a tuple, containing
1. a corrupted set of features,
2. a corrupted event label, and
3. a corrupted time index
```
y_mem[:5]
# plot the corrupted features
x_mem = np.concatenate([y_mem0[0].reshape(1, -1) for y_mem0 in y_mem])
# plt.plot()
plt.figure(figsize=(8,8))
plt.scatter(x_train[:, 0], x_train[:, 1], alpha=0.5, label='Original Features')
plt.scatter(x_mem[:, 0], x_mem[:, 1], marker='s', label='Memory Corrupted Features')
plt.legend()
```
## Reconstruction
As before, there is a function in the memory module to perform reconstruction. We just need to set parameters
```
from models.memory import gibbs_memory_sampler
help(gibbs_memory_sampler)
gibbs_memory_kwargs = dict(
y_mem=y_mem, sem_model=sem_model,
memory_alpha = sem_kwargs['alfa'],
memory_lambda = sem_kwargs['lmda'],
memory_epsilon = np.exp(-10),
b = noise_kwargs['b'],
tau = noise_kwargs['tau'],
)
y_samp, e_samp, x_samp = gibbs_memory_sampler(**gibbs_memory_kwargs)
from models.memory import reconstruction_accuracy, evaluate_seg
print "total reconstruction accuracy: {}".format(reconstruction_accuracy(y_samp, y_mem).mean())
# this metric evaluates the proportion of time each memory trace is included in the sample.
# It is a useful diagnositic and sometimes useful to think of this as recall.
print "reconstruction segmentation: {}".format(evaluate_seg(e_samp, e_true))
## How well the memory model recovered the segmentation is also a useful diagnostic
plt.figure(figsize=(8,8))
for ii in range(50):
plt.scatter(x_samp[ii][:, 0], x_samp[ii][:, 1], alpha=0.25, c=e_samp[ii], cmap='Paired')
plt.figure(figsize=(8,8))
for ii in range(25):
plt.scatter(x_samp[ii][:, 0], x_samp[ii][:, 1], alpha=0.05, c='k')
plt.scatter(x_train[:, 0], x_train[:, 1], label='Original Features')
plt.scatter(x_mem[:, 0], x_mem[:, 1], marker='s', label='Memory Corrupted Features')
plt.legend()
```
## Comparison to a Hidden Markov Model (HMM)
We can also compare SEM to a reduced model that infers events but does not learn event dynamics (a.k.a. a Hidden Markov Model). These models typically do well in segmentation, but are less often used as memory models (to our knoweldge).
```
from models.event_models import StationaryEvent
# copy the original segementation parameters and change the event model class to
# a stationary emmision distribution
hmm_kwargs = {k: v for k, v in sem_kwargs.iteritems()}
hmm_kwargs['f_class'] = StationaryEvent
hmm_model = SEM(**hmm_kwargs)
post = hmm_model.run(x_train)
y_hat_hmm = np.argmax(post, axis=1)
print "Adjusted Rand Score: ", metrics.adjusted_rand_score(y, y_hat_hmm)
# copy the gibbs parameters and just update the model
gibbs_memory_kwargs_hmm = {k: v for k, v in gibbs_memory_kwargs.iteritems()}
gibbs_memory_kwargs_hmm['sem_model'] = hmm_model
y_samp_hmm, e_samp_hmm, x_samp_hmm = gibbs_memory_sampler(**gibbs_memory_kwargs_hmm)
```
The HMM does a good job of segmentation but how good is it as a memory model? We can compare the reconstructed scenes for both SEM and the HMM by comparing the distribution of distances between the original scenes and the reconstructed equivalents.
```
def get_dist(a, b):
return np.linalg.norm(a - b, axis=1)
sns.distplot(np.concatenate([get_dist(x0, x_train) for x0 in x_samp]), label='SEM')
sns.distplot(np.concatenate([get_dist(x0, x_train) for x0 in x_samp_hmm]), label='HMM')
plt.xlabel('Reconstruction error\n (Distance to original scene)')
plt.legend()
sns.despine()
```
A key prediction of SEM is that memory traces are regularized towards the event dynamics, and not towards a global average of all scenes within an event.
To calculate this, we get the distance between:
1. each reconstructed feature and both the the original scene
2. each reconstructed feature and the average scene with the (inferred) event.
If (1) is greater than (2), then the model as regularized towards the event trajectory. If (2) is greater than (1), then the model has regularized towards the centroid. For simplicity, we just plot the relative difference between these two here, with a postive value indicating regularization towards the original scene and a negative value indicating regularization towards the centroid.
```
import pandas as pd
event_centroids = np.concatenate([
np.tile(x_train[e_true == 0, :].mean(axis=0), (20, 1)),
np.tile(x_train[e_true == 1, :].mean(axis=0), (20, 1))
])
sem_reld = np.concatenate([-get_dist(x0, x_train) + get_dist(x0, event_centroids) for x0 in x_samp])
red_reld = np.concatenate([-get_dist(x0, x_train) + get_dist(x0, event_centroids) for x0 in x_samp_hmm])
df = pd.DataFrame(
{
'Model': ['SEM'] * len(sem_reld) + ['Reduced'] * len(red_reld),
'Relative Distance': np.concatenate([sem_reld, red_reld])
})
sns.catplot(data=df, x='Model', y='Relative Distance', kind='bar')
plt.axhline(y=0, ls='--', c='k')
```
As we can see, SEM regularizes towards the memory trace and the HMM regularizes toward the avewrage scene
| github_jupyter |
MNIST contains 70,000 images of handwritten digits: 60,000 for training and 10,000 for testing. The images are grayscale, 28x28 pixels.
```
import matplotlib.pyplot as plt
%matplotlib inline
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout
import sys
import tensorflow as tf
import numpy as np
sys.version
batch_size = 128
num_classes = 10
epochs = 12
# input image dimensions
img_rows, img_cols = 28, 28
```
[Dataset](https://s3.amazonaws.com/img-datasets/mnist.npz)
### Load MNIST Dataset
```
myData = np.load('datasets/mnist.npz')
x_train, y_train = myData['x_train'], myData['y_train']
x_test, y_test = myData['x_test'], myData['y_test']
```
### Visualize the Training Images
```
print('Digit Image - {}'.format(y_train[51011]))
plt.imshow(x_train[51011].reshape(28,28), cmap='gray')
plt.show()
# plot first six training images
fig = plt.figure(figsize=(20,20))
for i in range(6):
ax = fig.add_subplot(1, 6, i+1, xticks=[], yticks=[])
ax.imshow(x_train[i].reshape(28,28), cmap='gray')
ax.set_title(str(y_train[i]))
x_train.shape
x_train = x_train.reshape(x_train.shape[0], img_rows * img_cols)
x_test = x_test.reshape(x_test.shape[0], img_rows * img_cols)
input_shape = img_rows * img_cols
x_test.shape
```
### Rescale the Images by Dividing Every Pixel in Every Image by 255
```
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
y_train
```
### Encode Categorical Integer Labels Using a One-Hot Scheme
```
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
y_train
```
### Define the Model Architecture
```
# building a linear stack of layers with the sequential model
model = Sequential()
model.add(Dense(512, input_shape=(784,), activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
print(keras.__version__)
print(tf.__version__)
```
### Compile the Model
```
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
```
### Neural Network
<img src=images/ann.png>
### Calculate the Classification Accuracy on the Test Set (Before Training)
```
# evaluate test accuracy
score = model.evaluate(x_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.2f%%' % accuracy)
```
### Train the Model
```
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
model.save('.\model\myModel_MNIST.h5')
```
### Load the Model
```
# load the weights that yielded the best validation accuracy
model.load_weights('.\model\myModel_MNIST.h5')
```
### Calculate the Classification Accuracy on the Test Set
```
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss: %.4f'% score[0])
print('Test accuracy: %.2f'% score[1])
test_digit = 515
#print('Digit Image - {}'.format(myData['y_test'][test_digit]))
print('Prediction - {}'.format(model.predict_classes(x_test[test_digit].reshape(1,784))[0]))
plt.imshow(x_test[test_digit].reshape(28,28), cmap='gray')
plt.show()
```
| github_jupyter |
# 圖論(Graph Theory)

This work by Jephian Lin is licensed under a [Creative Commons Attribution 4.0 International License](http://creativecommons.org/licenses/by/4.0/).
_Tested on SageMath version 8.7_
## 圖
一個__圖__ $G$ 由
一些**點**
還有一些__邊__構成
這裡邊指的是一些兩個點的集合
**點集**通常記作 $V(G)$
__邊集__通常記作 $E(G)$
```
g = graphs.CycleGraph(4)
print("V(G) = %s"%g.vertices())
print("E(G) = %s"%g.edges(labels=False))
g.show(figsize=[2,2])
```
點沒有一定的位置
(也沒有一定要用 $0,\ldots,n-1$ 標號)
上下兩張圖是一模一樣的
```
pos = {0:(0,0), 1:(1,0), 2:(0,1), 3:(1,1)}
g.set_pos(pos)
g.show(figsize=[2,2])
```
Sage 裡的 `graphs` 裡
內建了很多圖
許多圖也把一個*常用*的點座標設定好了
```
### Path
g = graphs.PathGraph(5)
g.show(figsize=[2,2])
### Cycle
g = graphs.CycleGraph(5)
g.show(figsize=[2,2])
### complete graph
g = graphs.CompleteGraph(5)
g.show(figsize=[2,2])
### star graph
g = graphs.StarGraph(5)
g.show(figsize=[2,2])
```
也可以從頭建造一個圖
設定好點集 `V` 以及邊集 `E`
並用 `Graph([V, E])` 建立一個圖
```
V = [0,1,2,3,4,5]
E = [(0,1), (1,2), (1,4), (3,4), (4,5)]
g = Graph([V, E])
g.show(figsize=[2,2])
```
有需要可以再增加點或邊
(如果邊的點原本不存在
則會自動加入)
```
g.add_vertex(15)
g.add_edge(0,30)
g.show(figsize=[2,2])
```
若 `g` 是 Sage 中的一個圖
則可以用 `g.show()` 來顯示
繪圖有許多參數可以調整
參考[官方說明書](http://doc.sagemath.org/html/en/reference/plotting/sage/graphs/graph_plot.html)
```
g = graphs.CycleGraph(5)
g.show(figsize=[2,2],
vertex_labels=False,
vertex_colors={'red':[1,3], 'blue':[2,4], 'orange':[0]},
edge_style='--'
)
```
### 圖同構
如果兩個圖可以把點和點對起來
使得相對應的邊也對起來
則兩個圖則視為**同構**
```
g = graphs.CubeGraph(3)
pos = g.get_pos()
pos['010'] = (0.7,0.9)
pos['101'] = (0.3,0.7)
g.set_pos(pos)
g.show(figsize=[2,2], vertex_labels=False)
V = [0,1,2,3,4,5,6,7]
E = [(0,1), (1,2), (2,3), (3,0), (0,4), (1,5), (2,6), (3,7), (4,5), (5,6), (6,7), (7,4)]
pos = {0:(0,0), 1:(3,0), 2:(3,3), 3:(0,3), 4:(1,1), 5:(2,1), 6:(2,2), 7:(1,2)}
h = Graph([V, E], pos=pos)
h.show(figsize=[2,2], vertex_labels=False)
Petersen = graphs.PetersenGraph()
Peterson = Petersen.copy()
Peterson.delete_edges([(5,7), (7,9), (9,6), (6,8), (8,5)])
Peterson.add_edges([(5,6), (6,7), (7,8), (8,9), (9,5)])
Petersen.show(figsize=[2,2], vertex_labels=False)
Peterson.show(figsize=[2,2], vertex_labels=False)
```
## 圖的不變量
圖會因為標號的不同而有些差異
但有些性質並不會隨著標號改變
比如說點數、邊數
這些性質叫作圖的**不變量**
它們可以用來幫忙判斷圖是否同構
若 `g` 是 Sage 裡的圖
則 `g.order()` 回傳點數
而 `g.size()` 回傳邊數
```
g = graphs.PetersenGraph()
print(g.order())
print(g.size())
```
### 度數
圖中一個點相連的邊數
稱作這個點的**度數**
比如說下圖中
點 0 的度數為 5
其餘點的度數皆為 1
```
g = graphs.StarGraph(5)
g.show(figsize=[2,2])
```
把每個點的度數收集起來
所形成(元素可重覆)的集合
稱作圖的**度數數列**
同構的圖必須要有相同的度數數列
```
g.degree_sequence()
```
#### 圖論第一定理
圖的度數總合
等於兩倍的邊數
```
print(sum(g.degree_sequence()))
print(2 * g.size())
```
因此度數數列的總合永遠是偶數
而度數為奇數的點一定有偶數個
### 連通
若一個圖中的任兩點
都可以經由一連串的邊接起來
則這個圖為**連通的**
```
g = graphs.CycleGraph(6)
h = graphs.CompleteGraph(3).disjoint_union(graphs.CompleteGraph(3))
g.show(figsize=[2,2], vertex_labels=False)
h.show(figsize=[2,2], vertex_labels=False)
```
大的圖不見得
一眼就可以看出是否連通
後面我們會介紹演算法
來判斷圖的連通性
```
g = graphs.RandomGNP(100,0.05)
g.show(vertex_labels=False, vertex_size=10)
g.is_connected()
```
### 路徑數
一個從點 $i$ 到點 $j$ 的路徑
指的是一群點 $i = v_0\sim v_1\sim\cdots\sim v_k =j$
這裡 $k$ 是路徑的長度
定義 $p_k(i,j)$ 為
$i$ 和 $j$ 中長度為 $k$ 的路徑數
固定一個長度 $k$ 以及一個整數 $d$
圖上有幾組點 $(i,j)$ 滿足 $p_k(i,j)=d$
這樣的組數也是圖的不變量
比如說 $p_1(i,j)=1$ 的組數
就是圖的邊數
#### 相鄰矩陣
一個 $n$ 個點的圖的**相鄰矩陣**
是一個 $n\times n$ 的矩陣
如果點 $i,j$ 有邊則矩陣的 $i,j$-項為 $1$
如果點 $i,j$ 沒邊則矩陣的 $i,j$-項為 $0$
```
g = graphs.PathGraph(4)
g.adjacency_matrix()
```
#### 定理
若 $A$ 為圖 $G$ 的相鄰矩陣
則 $A^k$ 的 $i,j$-項
等於 $p_k(i,j)$
```
g = graphs.PathGraph(4)
A = g.adjacency_matrix()
A**2
```
回顧 `Petersen` 及 `Peterson`
```
A1 = Petersen.adjacency_matrix()
A2 = Peterson.adjacency_matrix()
print("A1^2 =")
print(A1**2)
print("A2^2 =")
print(A2**2)
```
會發現 `Petersen`
完全沒有一組 $(i,j)$ 滿足 $p_2(i,j)=2$
但 `Peterson`
卻有 8 組
因此兩圖不同構
### 最小圈
圖上的一個**圈**
指的是一個起點和終點相同的路徑
而此外其餘任兩點皆不重覆
圈的長度為圈上的邊數
一個圈上的最小圈的長度
也是圈的不變量
可以用 `g.girth()` 來計算
```
print(Petersen.girth())
print(Peterson.girth())
```
再次看到兩圖不同構
### 生成樹
一個圖如果是連通的
而且圖上沒有圈
則被成為**樹**
```
g = graphs.RandomTree(10)
g.show(figsize=[2,2], vertex_labels=False, vertex_size=10)
```
$n$ 個點上的樹一定剛好 $n-1$ 條邊
一個圖上選 $n-1$ 條邊
使得這些邊形成一個樹
則稱為這個圖的**生成樹**
生成樹的個數是圖的不變量
```
g = graphs.CycleGraph(6)
g.show(figsize=[2,2])
```
#### 拉普拉斯矩陣
給定一個圖 $G$
令 $D$ 為一個對角線為 $G$ 的度數列表的對角矩陣
令 $A$ 為 $G$ 的相鄰矩陣
則 $G$ 的拉普拉斯矩陣定義為 $D-A$
```
g = graphs.CycleGraph(6)
L = g.laplacian_matrix()
L
```
#### 矩陣樹定理
令 $L$ 為圖 $G$ 的拉普拉斯矩陣
而 $L'$ 為將 $L$ 的第零行第零列去掉的矩陣
則 $|\det(L')|$ 為圖生成樹的個數
```
Lprime = L[1:,1:]
Lprime.determinant()
```
## 搜尋演算法
圖上的搜尋演算法
目的是在圖上有系統地搜尋所有點
(或是邊、或是都找)
以下是兩種常見的搜尋演算法:
1. 深度優先搜尋 Depth-first search(DFS)
2. 廣度優先搜尋 Breadth-first search(BFS)
這兩種演算法也可以用來判斷一個圖是否連通
```
### 先執行這段程式碼
def DFS_tree(g, v):
searched = []
arcs = []
for new in g.depth_first_search(v):
if searched:
for i in range(1,len(searched)+1):
if g.has_edge(new,searched[-i]):
parent = searched[-i]
break;
arcs.append((parent,new))
searched.append(new)
return arcs
def BFS_tree(g, v):
searched = []
arcs = []
for new in g.breadth_first_search(v):
if searched:
for i in range(len(searched)):
if g.has_edge(new,searched[i]):
parent = searched[i]
break;
arcs.append((parent,new))
searched.append(new)
return arcs
def greedy_coloring(g, color_order=None):
n = g.order()
### ideally, len(color_order) == n
if color_order == None:
color_order = g.vertices()
color_order = list(color_order) ### change the type in case it is a generator
num_c = {k: [] for k in range(n)}
for s in range(n):
new = color_order[s]
for k in range(n):
for u in num_c[k]:
if g.has_edge(u,new):
break;
else:
num_c[k].append(new)
break;
num_c_used = [k for k in range(n) if num_c[k]]
greedy_chi = len(num_c_used)
colors = rainbow(greedy_chi)
c = {colors[k]: num_c[k] for k in range(greedy_chi)}
return c
def illustrate_FS(g, v, alg='DFS', searching_tree=True, coloring=False):
### g should have its position saved
### if not do g.plot(save_pos) first
if alg == 'DFS':
arcs = DFS_tree(g,v)
full_name = 'Depth-First Search at {}'.format(v)
if alg == 'BFS':
arcs = BFS_tree(g,v)
full_name = 'Breadth-First Search at {}'.format(v)
steps = len(arcs)
pic1 = g.plot()
if coloring:
color_order = [v] + [arc[1] for arc in arcs]
c = greedy_coloring(g, color_order)
else:
c = {}
@interact
def _(step=slider(list(range(steps+1))), t = text_control(full_name)):
g.set_pos(g.layout())
g_pos = g.get_pos()
arcs_show = arcs[:step] if searching_tree else []
pic2 = DiGraph([g.vertices(),arcs_show], pos=g_pos).plot(edge_color='red', vertex_colors=c)
unreached = [arc[1] for arc in arcs[step:]]
cover = Graph([unreached,[]], pos={u: g_pos[u] for u in unreached}).plot()
p = pic1 + pic2 + cover
p.axes(False)
p.show()
```
### 深度優先搜尋 Depth-first search(DFS)
能往前就往前
不能再往前時,往回退到還能往前的點繼續
例子:走迷宮
```
g = graphs.PetersenGraph()
v = 0
illustrate_FS(g, v, 'DFS')
```
### 廣度優先搜尋 Breadth-first search(BFS)
先把附近找完
附近沒了,再從第一個附近還沒找完的點繼續
例子:找兩點之間的最短路徑
```
g = graphs.PetersenGraph()
v = 0
illustrate_FS(g, v, 'BFS')
```
## 動手試試看
##### 練習 1
定義一個函數 `spanning_tree_count(g)` 其功能為:
傳入一個圖 `g`
回傳此圖的生成樹的個數
提示:拉普拉斯矩陣
```
### your answer here
```
##### 練習 2
定義一個函數 `is_conn(g)` 其功能為:
傳入一個圖 `g`
若 `g` 連通則回傳 `True` 否則回傳 `False`
(有辦法利用 `spanning_tree_count` 判斷嗎?)
```
### your answer here
```
##### 練習 3
定義一個函數 `is_tree(g)` 其功能為:
傳入一個圖 `g`
若 `g` 為樹則回傳 `True` 否則回傳 `False`
(有辦法利用 `spanning_tree_count` 判斷嗎?)
```
### your answer here
```
##### 練習 4
定義一個函數 `to_tree(g)` 其功能為:
傳入一個連通圖 `g`
將 `g` 多餘的邊移除,使其成為一個樹(可使用 `g.delete_edge(e)` 移除邊)
回傳 `g`
提示:
使用 `is_conn` 幫助判斷哪些邊需要移除
先將某個邊 `e` 移除
若移除之後圖變得不連通,那就把 `e` 加回去(`g.add_edge(e)`)
```
### your answer here
```
##### 獨立集
給定一個圖 $G$ ,令 $S$ 為 $V(G)$ 的子集合
若 $S$ 裡面的任兩個點在圖 $G$ 中都不相鄰,則稱 $S$ 為 $G$ 的獨立集
一個圖 $G$ 可以有很多不同的獨立集
若 $S$ 為圖 $G$ 元素個數最多的獨立集,則稱 $S$ 為 $G$ 的最大獨立集
##### 練習 5
定義一個函數 `is_independent_set(g, s)` 其功能為:
傳入一個圖 `g` 和一個點集 `s`
`s` 為 `V(g)` 的子集合
若 `s` 為 `g` 的獨立集則回傳 `True`,否則回傳 `False`
提示:
使用 `Combinations(s, 2)` 將所有的可能取出
接著用 `g.has_edge(v1, v2)` 判斷是否相鄰
```
### your answer here
```
##### 練習 6
定義一個函數 `all_independent_set(g)` 其功能為:
傳入一個圖 `g`
回傳一個 `list` 為 `g` 的所有獨立集
提示:
1. 開一個空的 `list`
2. 使用 `Combinations(g.vertices(), n)` , n = 0, 1, 2, ..., k 將所有 `V(g)` 的子集合取出
3. 若取出的子集合為 `g` 的獨立集就把它加入到 `list` 中
```
### your answer here
```
##### 練習 7
定義一個函數 `max_independent_set(g)` 其功能為:
傳入一個圖 `g`
回傳一個 `g` 的最大獨立集
```
### your answer here
```
先執行下面這段程式碼,觀察圖 `g` 跟印出的 `list` 之間的關聯性
```
g = graphs.PetersenGraph()
g += g
g.show()
list(g.depth_first_search(0))
```
##### 練習 8
定義一個函數 `has_path(g, v1, v2)` 其功能為:
傳入一個圖 `g` 和兩個點 `v1`, `v2`
若 `v1` 和 `v2` 之間能夠用一串邊連接起來,就回傳 `True`,否則回傳 `False`
提示:想想看能不能使用 `g.depth_first_search` 來幫助判斷
```
### your answer here
```
##### 練習 9
定義一個函數 `to_conn(g)` 其功能為:
傳入一個圖 `g`
若 `g` 不是連通圖,補一些缺失的邊,使其成為連通圖(可使用 `g.add_edge(e)` 補上邊)
回傳 `g`
提示:用上面的 `has_path` 函數判斷要補那些邊
```
### your answer here
```
##### 練習 10
定義一個函數 `find_path(g, v1, v2)` 其功能為:
傳入一個連通圖 `g` 和兩個點 `v1`, `v2`
回傳一個 `list` 裡面為一串將 `v1` 和 `v2` 連接起來的邊
(`list` 的格式必須是 `[(v1,v(1)),(v(1),v(2)), ... ,(v(k-1),v(k)),(v(k),v2)]`,`v(1)`, `v(2)`, ..., `v(k)` 為 `g` 中的點)
請先將 `g` 在函數內複製一份,不要直接改寫 `g` 的內容
```Python
def find_path(g, v1, v2):
h = g.copy()
```
提示:
考慮這幾個函數 `to_tree`, `has_path`, `h.delete_edge`, `h.add_edge`, `h.depth_first_search`
我們先把 `h` 變成一個樹,這樣可以保證任兩點之間只有一種唯一的走法
接下來刪除從 `v1` 走到 `v2` 之間不會經過的邊(想想看要用哪個函數幫忙判斷哪些是要刪除的邊)
此時 `h` 上所剩的邊就是連接 `v1` 和 `v2` 的路徑
最後,試著用 `h.depth_first_search` 造出題目要的 `list`
```
### your answer here
```
完成 **練習10** 後,可以測試以下的程式
```
### 先執行這段程式碼
def illustrate(g, arcs, color='blue'):
g.set_pos(g.layout())
steps = len(arcs)
pic1 = g.plot()
@interact
def _(step=slider(list(range(steps+1)))):
g_pos = g.get_pos()
arcs_show = arcs[:step]
pic2 = DiGraph([g.vertices(),arcs_show], pos=g_pos).plot(edge_color=color)
p = pic1 + pic2
p.axes(False)
p.show()
### 執行看看會有甚麼效果
### g, start_vertex, end_vertex 這三個可以自己改
g = graphs.RandomGNP(25, 0.2)
to_conn(g)
start_vertex = 0
end_vertex = 24
illustrate(g, find_path(g, start_vertex, end_vertex))
```
| github_jupyter |
# Emojify!
Welcome to the second assignment of Week 2. You are going to use word vector representations to build an Emojifier.
Have you ever wanted to make your text messages more expressive? Your emojifier app will help you do that. So rather than writing "Congratulations on the promotion! Lets get coffee and talk. Love you!" the emojifier can automatically turn this into "Congratulations on the promotion! 👍 Lets get coffee and talk. ☕️ Love you! ❤️"
You will implement a model which inputs a sentence (such as "Let's go see the baseball game tonight!") and finds the most appropriate emoji to be used with this sentence (⚾️). In many emoji interfaces, you need to remember that ❤️ is the "heart" symbol rather than the "love" symbol. But using word vectors, you'll see that even if your training set explicitly relates only a few words to a particular emoji, your algorithm will be able to generalize and associate words in the test set to the same emoji even if those words don't even appear in the training set. This allows you to build an accurate classifier mapping from sentences to emojis, even using a small training set.
In this exercise, you'll start with a baseline model (Emojifier-V1) using word embeddings, then build a more sophisticated model (Emojifier-V2) that further incorporates an LSTM.
Lets get started! Run the following cell to load the package you are going to use.
```
import numpy as np
from emo_utils import *
import emoji
import matplotlib.pyplot as plt
%matplotlib inline
```
## 1 - Baseline model: Emojifier-V1
### 1.1 - Dataset EMOJISET
Let's start by building a simple baseline classifier.
You have a tiny dataset (X, Y) where:
- X contains 127 sentences (strings)
- Y contains a integer label between 0 and 4 corresponding to an emoji for each sentence
<img src="images/data_set.png" style="width:700px;height:300px;">
<caption><center> **Figure 1**: EMOJISET - a classification problem with 5 classes. A few examples of sentences are given here. </center></caption>
Let's load the dataset using the code below. We split the dataset between training (127 examples) and testing (56 examples).
```
X_train, Y_train = read_csv('data/train_emoji.csv')
X_test, Y_test = read_csv('data/tesss.csv')
maxLen = len(max(X_train, key=len).split())
```
Run the following cell to print sentences from X_train and corresponding labels from Y_train. Change `index` to see different examples. Because of the font the iPython notebook uses, the heart emoji may be colored black rather than red.
```
index = 1
print(X_train[index], label_to_emoji(Y_train[index]))
```
### 1.2 - Overview of the Emojifier-V1
In this part, you are going to implement a baseline model called "Emojifier-v1".
<center>
<img src="images/image_1.png" style="width:900px;height:300px;">
<caption><center> **Figure 2**: Baseline model (Emojifier-V1).</center></caption>
</center>
The input of the model is a string corresponding to a sentence (e.g. "I love you). In the code, the output will be a probability vector of shape (1,5), that you then pass in an argmax layer to extract the index of the most likely emoji output.
To get our labels into a format suitable for training a softmax classifier, lets convert $Y$ from its current shape current shape $(m, 1)$ into a "one-hot representation" $(m, 5)$, where each row is a one-hot vector giving the label of one example, You can do so using this next code snipper. Here, `Y_oh` stands for "Y-one-hot" in the variable names `Y_oh_train` and `Y_oh_test`:
```
Y_oh_train = convert_to_one_hot(Y_train, C = 5)
Y_oh_test = convert_to_one_hot(Y_test, C = 5)
```
Let's see what `convert_to_one_hot()` did. Feel free to change `index` to print out different values.
```
index = 50
print(Y_train[index], "is converted into one hot", Y_oh_train[index])
```
All the data is now ready to be fed into the Emojify-V1 model. Let's implement the model!
### 1.3 - Implementing Emojifier-V1
As shown in Figure (2), the first step is to convert an input sentence into the word vector representation, which then get averaged together. Similar to the previous exercise, we will use pretrained 50-dimensional GloVe embeddings. Run the following cell to load the `word_to_vec_map`, which contains all the vector representations.
```
word_to_index, index_to_word, word_to_vec_map = read_glove_vecs('data/glove.6B.50d.txt')
```
You've loaded:
- `word_to_index`: dictionary mapping from words to their indices in the vocabulary (400,001 words, with the valid indices ranging from 0 to 400,000)
- `index_to_word`: dictionary mapping from indices to their corresponding words in the vocabulary
- `word_to_vec_map`: dictionary mapping words to their GloVe vector representation.
Run the following cell to check if it works.
```
word = "cucumber"
index = 289846
print("the index of", word, "in the vocabulary is", word_to_index[word])
print("the", str(index) + "th word in the vocabulary is", index_to_word[index])
```
**Exercise**: Implement `sentence_to_avg()`. You will need to carry out two steps:
1. Convert every sentence to lower-case, then split the sentence into a list of words. `X.lower()` and `X.split()` might be useful.
2. For each word in the sentence, access its GloVe representation. Then, average all these values.
```
# GRADED FUNCTION: sentence_to_avg
def sentence_to_avg(sentence, word_to_vec_map):
"""
Converts a sentence (string) into a list of words (strings). Extracts the GloVe representation of each word
and averages its value into a single vector encoding the meaning of the sentence.
Arguments:
sentence -- string, one training example from X
word_to_vec_map -- dictionary mapping every word in a vocabulary into its 50-dimensional vector representation
Returns:
avg -- average vector encoding information about the sentence, numpy-array of shape (50,)
"""
### START CODE HERE ###
# Step 1: Split sentence into list of lower case words (≈ 1 line)
words = None
# Initialize the average word vector, should have the same shape as your word vectors.
avg = None
# Step 2: average the word vectors. You can loop over the words in the list "words".
for w in None:
avg += None
avg = None
### END CODE HERE ###
return avg
avg = sentence_to_avg("Morrocan couscous is my favorite dish", word_to_vec_map)
print("avg = ", avg)
```
**Expected Output**:
<table>
<tr>
<td>
**avg= **
</td>
<td>
[-0.008005 0.56370833 -0.50427333 0.258865 0.55131103 0.03104983
-0.21013718 0.16893933 -0.09590267 0.141784 -0.15708967 0.18525867
0.6495785 0.38371117 0.21102167 0.11301667 0.02613967 0.26037767
0.05820667 -0.01578167 -0.12078833 -0.02471267 0.4128455 0.5152061
0.38756167 -0.898661 -0.535145 0.33501167 0.68806933 -0.2156265
1.797155 0.10476933 -0.36775333 0.750785 0.10282583 0.348925
-0.27262833 0.66768 -0.10706167 -0.283635 0.59580117 0.28747333
-0.3366635 0.23393817 0.34349183 0.178405 0.1166155 -0.076433
0.1445417 0.09808667]
</td>
</tr>
</table>
#### Model
You now have all the pieces to finish implementing the `model()` function. After using `sentence_to_avg()` you need to pass the average through forward propagation, compute the cost, and then backpropagate to update the softmax's parameters.
**Exercise**: Implement the `model()` function described in Figure (2). Assuming here that $Yoh$ ("Y one hot") is the one-hot encoding of the output labels, the equations you need to implement in the forward pass and to compute the cross-entropy cost are:
$$ z^{(i)} = W . avg^{(i)} + b$$
$$ a^{(i)} = softmax(z^{(i)})$$
$$ \mathcal{L}^{(i)} = - \sum_{k = 0}^{n_y - 1} Yoh^{(i)}_k * log(a^{(i)}_k)$$
It is possible to come up with a more efficient vectorized implementation. But since we are using a for-loop to convert the sentences one at a time into the avg^{(i)} representation anyway, let's not bother this time.
We provided you a function `softmax()`.
```
# GRADED FUNCTION: model
def model(X, Y, word_to_vec_map, learning_rate = 0.01, num_iterations = 400):
"""
Model to train word vector representations in numpy.
Arguments:
X -- input data, numpy array of sentences as strings, of shape (m, 1)
Y -- labels, numpy array of integers between 0 and 7, numpy-array of shape (m, 1)
word_to_vec_map -- dictionary mapping every word in a vocabulary into its 50-dimensional vector representation
learning_rate -- learning_rate for the stochastic gradient descent algorithm
num_iterations -- number of iterations
Returns:
pred -- vector of predictions, numpy-array of shape (m, 1)
W -- weight matrix of the softmax layer, of shape (n_y, n_h)
b -- bias of the softmax layer, of shape (n_y,)
"""
np.random.seed(1)
# Define number of training examples
m = Y.shape[0] # number of training examples
n_y = 5 # number of classes
n_h = 50 # dimensions of the GloVe vectors
# Initialize parameters using Xavier initialization
W = np.random.randn(n_y, n_h) / np.sqrt(n_h)
b = np.zeros((n_y,))
# Convert Y to Y_onehot with n_y classes
Y_oh = convert_to_one_hot(Y, C = n_y)
# Optimization loop
for t in range(num_iterations): # Loop over the number of iterations
for i in range(m): # Loop over the training examples
### START CODE HERE ### (≈ 4 lines of code)
# Average the word vectors of the words from the i'th training example
avg = None
# Forward propagate the avg through the softmax layer
z = None
a = None
# Compute cost using the i'th training label's one hot representation and "A" (the output of the softmax)
cost = None
### END CODE HERE ###
# Compute gradients
dz = a - Y_oh[i]
dW = np.dot(dz.reshape(n_y,1), avg.reshape(1, n_h))
db = dz
# Update parameters with Stochastic Gradient Descent
W = W - learning_rate * dW
b = b - learning_rate * db
if t % 100 == 0:
print("Epoch: " + str(t) + " --- cost = " + str(cost))
pred = predict(X, Y, W, b, word_to_vec_map)
return pred, W, b
print(X_train.shape)
print(Y_train.shape)
print(np.eye(5)[Y_train.reshape(-1)].shape)
print(X_train[0])
print(type(X_train))
Y = np.asarray([5,0,0,5, 4, 4, 4, 6, 6, 4, 1, 1, 5, 6, 6, 3, 6, 3, 4, 4])
print(Y.shape)
X = np.asarray(['I am going to the bar tonight', 'I love you', 'miss you my dear',
'Lets go party and drinks','Congrats on the new job','Congratulations',
'I am so happy for you', 'Why are you feeling bad', 'What is wrong with you',
'You totally deserve this prize', 'Let us go play football',
'Are you down for football this afternoon', 'Work hard play harder',
'It is suprising how people can be dumb sometimes',
'I am very disappointed','It is the best day in my life',
'I think I will end up alone','My life is so boring','Good job',
'Great so awesome'])
print(X.shape)
print(np.eye(5)[Y_train.reshape(-1)].shape)
print(type(X_train))
```
Run the next cell to train your model and learn the softmax parameters (W,b).
```
pred, W, b = model(X_train, Y_train, word_to_vec_map)
print(pred)
```
**Expected Output** (on a subset of iterations):
<table>
<tr>
<td>
**Epoch: 0**
</td>
<td>
cost = 1.95204988128
</td>
<td>
Accuracy: 0.348484848485
</td>
</tr>
<tr>
<td>
**Epoch: 100**
</td>
<td>
cost = 0.0797181872601
</td>
<td>
Accuracy: 0.931818181818
</td>
</tr>
<tr>
<td>
**Epoch: 200**
</td>
<td>
cost = 0.0445636924368
</td>
<td>
Accuracy: 0.954545454545
</td>
</tr>
<tr>
<td>
**Epoch: 300**
</td>
<td>
cost = 0.0343226737879
</td>
<td>
Accuracy: 0.969696969697
</td>
</tr>
</table>
Great! Your model has pretty high accuracy on the training set. Lets now see how it does on the test set.
### 1.4 - Examining test set performance
```
print("Training set:")
pred_train = predict(X_train, Y_train, W, b, word_to_vec_map)
print('Test set:')
pred_test = predict(X_test, Y_test, W, b, word_to_vec_map)
```
**Expected Output**:
<table>
<tr>
<td>
**Train set accuracy**
</td>
<td>
97.7
</td>
</tr>
<tr>
<td>
**Test set accuracy**
</td>
<td>
85.7
</td>
</tr>
</table>
Random guessing would have had 20% accuracy given that there are 5 classes. This is pretty good performance after training on only 127 examples.
In the training set, the algorithm saw the sentence "*I love you*" with the label ❤️. You can check however that the word "adore" does not appear in the training set. Nonetheless, lets see what happens if you write "*I adore you*."
```
X_my_sentences = np.array(["i adore you", "i love you", "funny lol", "lets play with a ball", "food is ready", "not feeling happy"])
Y_my_labels = np.array([[0], [0], [2], [1], [4],[3]])
pred = predict(X_my_sentences, Y_my_labels , W, b, word_to_vec_map)
print_predictions(X_my_sentences, pred)
```
Amazing! Because *adore* has a similar embedding as *love*, the algorithm has generalized correctly even to a word it has never seen before. Words such as *heart*, *dear*, *beloved* or *adore* have embedding vectors similar to *love*, and so might work too---feel free to modify the inputs above and try out a variety of input sentences. How well does it work?
Note though that it doesn't get "not feeling happy" correct. This algorithm ignores word ordering, so is not good at understanding phrases like "not happy."
Printing the confusion matrix can also help understand which classes are more difficult for your model. A confusion matrix shows how often an example whose label is one class ("actual" class) is mislabeled by the algorithm with a different class ("predicted" class).
```
print(Y_test.shape)
print(' '+ label_to_emoji(0)+ ' ' + label_to_emoji(1) + ' ' + label_to_emoji(2)+ ' ' + label_to_emoji(3)+' ' + label_to_emoji(4))
print(pd.crosstab(Y_test, pred_test.reshape(56,), rownames=['Actual'], colnames=['Predicted'], margins=True))
plot_confusion_matrix(Y_test, pred_test)
```
<font color='blue'>
**What you should remember from this part**:
- Even with a 127 training examples, you can get a reasonably good model for Emojifying. This is due to the generalization power word vectors gives you.
- Emojify-V1 will perform poorly on sentences such as *"This movie is not good and not enjoyable"* because it doesn't understand combinations of words--it just averages all the words' embedding vectors together, without paying attention to the ordering of words. You will build a better algorithm in the next part.
## 2 - Emojifier-V2: Using LSTMs in Keras:
Let's build an LSTM model that takes as input word sequences. This model will be able to take word ordering into account. Emojifier-V2 will continue to use pre-trained word embeddings to represent words, but will feed them into an LSTM, whose job it is to predict the most appropriate emoji.
Run the following cell to load the Keras packages.
```
import numpy as np
np.random.seed(0)
from keras.models import Model
from keras.layers import Dense, Input, Dropout, LSTM, Activation
from keras.layers.embeddings import Embedding
from keras.preprocessing import sequence
from keras.initializers import glorot_uniform
np.random.seed(1)
```
### 2.1 - Overview of the model
Here is the Emojifier-v2 you will implement:
<img src="images/emojifier-v2.png" style="width:700px;height:400px;"> <br>
<caption><center> **Figure 3**: Emojifier-V2. A 2-layer LSTM sequence classifier. </center></caption>
### 2.2 Keras and mini-batching
In this exercise, we want to train Keras using mini-batches. However, most deep learning frameworks require that all sequences in the same mini-batch have the same length. This is what allows vectorization to work: If you had a 3-word sentence and a 4-word sentence, then the computations needed for them are different (one takes 3 steps of an LSTM, one takes 4 steps) so it's just not possible to do them both at the same time.
The common solution to this is to use padding. Specifically, set a maximum sequence length, and pad all sequences to the same length. For example, of the maximum sequence length is 20, we could pad every sentence with "0"s so that each input sentence is of length 20. Thus, a sentence "i love you" would be represented as $(e_{i}, e_{love}, e_{you}, \vec{0}, \vec{0}, \ldots, \vec{0})$. In this example, any sentences longer than 20 words would have to be truncated. One simple way to choose the maximum sequence length is to just pick the length of the longest sentence in the training set.
### 2.3 - The Embedding layer
In Keras, the embedding matrix is represented as a "layer", and maps positive integers (indices corresponding to words) into dense vectors of fixed size (the embedding vectors). It can be trained or initialized with a pretrained embedding. In this part, you will learn how to create an [Embedding()](https://keras.io/layers/embeddings/) layer in Keras, initialize it with the GloVe 50-dimensional vectors loaded earlier in the notebook. Because our training set is quite small, we will not update the word embeddings but will instead leave their values fixed. But in the code below, we'll show you how Keras allows you to either train or leave fixed this layer.
The `Embedding()` layer takes an integer matrix of size (batch size, max input length) as input. This corresponds to sentences converted into lists of indices (integers), as shown in the figure below.
<img src="images/embedding1.png" style="width:700px;height:250px;">
<caption><center> **Figure 4**: Embedding layer. This example shows the propagation of two examples through the embedding layer. Both have been zero-padded to a length of `max_len=5`. The final dimension of the representation is `(2,max_len,50)` because the word embeddings we are using are 50 dimensional. </center></caption>
The largest integer (i.e. word index) in the input should be no larger than the vocabulary size. The layer outputs an array of shape (batch size, max input length, dimension of word vectors).
The first step is to convert all your training sentences into lists of indices, and then zero-pad all these lists so that their length is the length of the longest sentence.
**Exercise**: Implement the function below to convert X (array of sentences as strings) into an array of indices corresponding to words in the sentences. The output shape should be such that it can be given to `Embedding()` (described in Figure 4).
```
# GRADED FUNCTION: sentences_to_indices
def sentences_to_indices(X, word_to_index, max_len):
"""
Converts an array of sentences (strings) into an array of indices corresponding to words in the sentences.
The output shape should be such that it can be given to `Embedding()` (described in Figure 4).
Arguments:
X -- array of sentences (strings), of shape (m, 1)
word_to_index -- a dictionary containing the each word mapped to its index
max_len -- maximum number of words in a sentence. You can assume every sentence in X is no longer than this.
Returns:
X_indices -- array of indices corresponding to words in the sentences from X, of shape (m, max_len)
"""
m = X.shape[0] # number of training examples
### START CODE HERE ###
# Initialize X_indices as a numpy matrix of zeros and the correct shape (≈ 1 line)
X_indices = None
for i in range(m): # loop over training examples
# Convert the ith training sentence in lower case and split is into words. You should get a list of words.
sentence_words =None
# Initialize j to 0
j = None
# Loop over the words of sentence_words
for w in None:
# Set the (i,j)th entry of X_indices to the index of the correct word.
X_indices[i, j] = None
# Increment j to j + 1
j = None
### END CODE HERE ###
return X_indices
```
Run the following cell to check what `sentences_to_indices()` does, and check your results.
```
X1 = np.array(["funny lol", "lets play baseball", "food is ready for you"])
X1_indices = sentences_to_indices(X1,word_to_index, max_len = 5)
print("X1 =", X1)
print("X1_indices =", X1_indices)
```
**Expected Output**:
<table>
<tr>
<td>
**X1 =**
</td>
<td>
['funny lol' 'lets play football' 'food is ready for you']
</td>
</tr>
<tr>
<td>
**X1_indices =**
</td>
<td>
[[ 155345. 225122. 0. 0. 0.] <br>
[ 220930. 286375. 151266. 0. 0.] <br>
[ 151204. 192973. 302254. 151349. 394475.]]
</td>
</tr>
</table>
Let's build the `Embedding()` layer in Keras, using pre-trained word vectors. After this layer is built, you will pass the output of `sentences_to_indices()` to it as an input, and the `Embedding()` layer will return the word embeddings for a sentence.
**Exercise**: Implement `pretrained_embedding_layer()`. You will need to carry out the following steps:
1. Initialize the embedding matrix as a numpy array of zeroes with the correct shape.
2. Fill in the embedding matrix with all the word embeddings extracted from `word_to_vec_map`.
3. Define Keras embedding layer. Use [Embedding()](https://keras.io/layers/embeddings/). Be sure to make this layer non-trainable, by setting `trainable = False` when calling `Embedding()`. If you were to set `trainable = True`, then it will allow the optimization algorithm to modify the values of the word embeddings.
4. Set the embedding weights to be equal to the embedding matrix
```
# GRADED FUNCTION: pretrained_embedding_layer
def pretrained_embedding_layer(word_to_vec_map, word_to_index):
"""
Creates a Keras Embedding() layer and loads in pre-trained GloVe 50-dimensional vectors.
Arguments:
word_to_vec_map -- dictionary mapping words to their GloVe vector representation.
word_to_index -- dictionary mapping from words to their indices in the vocabulary (400,001 words)
Returns:
embedding_layer -- pretrained layer Keras instance
"""
vocab_len = len(word_to_index) + 1 # adding 1 to fit Keras embedding (requirement)
emb_dim = word_to_vec_map["cucumber"].shape[0] # define dimensionality of your GloVe word vectors (= 50)
### START CODE HERE ###
# Initialize the embedding matrix as a numpy array of zeros of shape (vocab_len, dimensions of word vectors = emb_dim)
emb_matrix = None
# Set each row "index" of the embedding matrix to be the word vector representation of the "index"th word of the vocabulary
for word, index in word_to_index.items():
emb_matrix[index, :] = None
# Define Keras embedding layer with the correct output/input sizes, make it trainable. Use Embedding(...). Make sure to set trainable=False.
embedding_layer = None
### END CODE HERE ###
# Build the embedding layer, it is required before setting the weights of the embedding layer. Do not modify the "None".
embedding_layer.build((None,))
# Set the weights of the embedding layer to the embedding matrix. Your layer is now pretrained.
embedding_layer.set_weights([emb_matrix])
return embedding_layer
embedding_layer = pretrained_embedding_layer(word_to_vec_map, word_to_index)
print("weights[0][1][3] =", embedding_layer.get_weights()[0][1][3])
```
**Expected Output**:
<table>
<tr>
<td>
**weights[0][1][3] =**
</td>
<td>
-0.3403
</td>
</tr>
</table>
## 2.3 Building the Emojifier-V2
Lets now build the Emojifier-V2 model. You will do so using the embedding layer you have built, and feed its output to an LSTM network.
<img src="images/emojifier-v2.png" style="width:700px;height:400px;"> <br>
<caption><center> **Figure 3**: Emojifier-v2. A 2-layer LSTM sequence classifier. </center></caption>
**Exercise:** Implement `Emojify_V2()`, which builds a Keras graph of the architecture shown in Figure 3. The model takes as input an array of sentences of shape (`m`, `max_len`, ) defined by `input_shape`. It should output a softmax probability vector of shape (`m`, `C = 5`). You may need `Input(shape = ..., dtype = '...')`, [LSTM()](https://keras.io/layers/recurrent/#lstm), [Dropout()](https://keras.io/layers/core/#dropout), [Dense()](https://keras.io/layers/core/#dense), and [Activation()](https://keras.io/activations/).
```
# GRADED FUNCTION: Emojify_V2
def Emojify_V2(input_shape, word_to_vec_map, word_to_index):
"""
Function creating the Emojify-v2 model's graph.
Arguments:
input_shape -- shape of the input, usually (max_len,)
word_to_vec_map -- dictionary mapping every word in a vocabulary into its 50-dimensional vector representation
word_to_index -- dictionary mapping from words to their indices in the vocabulary (400,001 words)
Returns:
model -- a model instance in Keras
"""
### START CODE HERE ###
# Define sentence_indices as the input of the graph, it should be of shape input_shape and dtype 'int32' (as it contains indices).
sentence_indices = None
# Create the embedding layer pretrained with GloVe Vectors (≈1 line)
embedding_layer = None
# Propagate sentence_indices through your embedding layer, you get back the embeddings
embeddings = None
# Propagate the embeddings through an LSTM layer with 128-dimensional hidden state
# Be careful, the returned output should be a batch of sequences.
X = None
# Add dropout with a probability of 0.5
X = None
# Propagate X trough another LSTM layer with 128-dimensional hidden state
# Be careful, the returned output should be a single hidden state, not a batch of sequences.
X = None
# Add dropout with a probability of 0.5
X = None
# Propagate X through a Dense layer with softmax activation to get back a batch of 5-dimensional vectors.
X = None
# Add a softmax activation
X = None
# Create Model instance which converts sentence_indices into X.
model = None
### END CODE HERE ###
return model
```
Run the following cell to create your model and check its summary. Because all sentences in the dataset are less than 10 words, we chose `max_len = 10`. You should see your architecture, it uses "20,223,927" parameters, of which 20,000,050 (the word embeddings) are non-trainable, and the remaining 223,877 are. Because our vocabulary size has 400,001 words (with valid indices from 0 to 400,000) there are 400,001\*50 = 20,000,050 non-trainable parameters.
```
model = Emojify_V2((maxLen,), word_to_vec_map, word_to_index)
model.summary()
```
As usual, after creating your model in Keras, you need to compile it and define what loss, optimizer and metrics your are want to use. Compile your model using `categorical_crossentropy` loss, `adam` optimizer and `['accuracy']` metrics:
```
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
```
It's time to train your model. Your Emojifier-V2 `model` takes as input an array of shape (`m`, `max_len`) and outputs probability vectors of shape (`m`, `number of classes`). We thus have to convert X_train (array of sentences as strings) to X_train_indices (array of sentences as list of word indices), and Y_train (labels as indices) to Y_train_oh (labels as one-hot vectors).
```
X_train_indices = sentences_to_indices(X_train, word_to_index, maxLen)
Y_train_oh = convert_to_one_hot(Y_train, C = 5)
```
Fit the Keras model on `X_train_indices` and `Y_train_oh`. We will use `epochs = 50` and `batch_size = 32`.
```
model.fit(X_train_indices, Y_train_oh, epochs = 50, batch_size = 32, shuffle=True)
```
Your model should perform close to **100% accuracy** on the training set. The exact accuracy you get may be a little different. Run the following cell to evaluate your model on the test set.
```
X_test_indices = sentences_to_indices(X_test, word_to_index, max_len = maxLen)
Y_test_oh = convert_to_one_hot(Y_test, C = 5)
loss, acc = model.evaluate(X_test_indices, Y_test_oh)
print()
print("Test accuracy = ", acc)
```
You should get a test accuracy between 80% and 95%. Run the cell below to see the mislabelled examples.
```
# This code allows you to see the mislabelled examples
C = 5
y_test_oh = np.eye(C)[Y_test.reshape(-1)]
X_test_indices = sentences_to_indices(X_test, word_to_index, maxLen)
pred = model.predict(X_test_indices)
for i in range(len(X_test)):
x = X_test_indices
num = np.argmax(pred[i])
if(num != Y_test[i]):
print('Expected emoji:'+ label_to_emoji(Y_test[i]) + ' prediction: '+ X_test[i] + label_to_emoji(num).strip())
```
Now you can try it on your own example. Write your own sentence below.
```
# Change the sentence below to see your prediction. Make sure all the words are in the Glove embeddings.
x_test = np.array(['not feeling happy'])
X_test_indices = sentences_to_indices(x_test, word_to_index, maxLen)
print(x_test[0] +' '+ label_to_emoji(np.argmax(model.predict(X_test_indices))))
```
Previously, Emojify-V1 model did not correctly label "not feeling happy," but our implementation of Emojiy-V2 got it right. (Keras' outputs are slightly random each time, so you may not have obtained the same result.) The current model still isn't very robust at understanding negation (like "not happy") because the training set is small and so doesn't have a lot of examples of negation. But if the training set were larger, the LSTM model would be much better than the Emojify-V1 model at understanding such complex sentences.
### Congratulations!
You have completed this notebook! ❤️❤️❤️
<font color='blue'>
**What you should remember**:
- If you have an NLP task where the training set is small, using word embeddings can help your algorithm significantly. Word embeddings allow your model to work on words in the test set that may not even have appeared in your training set.
- Training sequence models in Keras (and in most other deep learning frameworks) requires a few important details:
- To use mini-batches, the sequences need to be padded so that all the examples in a mini-batch have the same length.
- An `Embedding()` layer can be initialized with pretrained values. These values can be either fixed or trained further on your dataset. If however your labeled dataset is small, it's usually not worth trying to train a large pre-trained set of embeddings.
- `LSTM()` has a flag called `return_sequences` to decide if you would like to return every hidden states or only the last one.
- You can use `Dropout()` right after `LSTM()` to regularize your network.
Congratulations on finishing this assignment and building an Emojifier. We hope you're happy with what you've accomplished in this notebook!
# 😀😀😀😀😀😀
## Acknowledgments
Thanks to Alison Darcy and the Woebot team for their advice on the creation of this assignment. Woebot is a chatbot friend that is ready to speak with you 24/7. As part of Woebot's technology, it uses word embeddings to understand the emotions of what you say. You can play with it by going to http://woebot.io
<img src="images/woebot.png" style="width:600px;height:300px;">
| github_jupyter |
# Машинное обучение, ФКН ВШЭ
## Практическое задание 2. KNN. Exploratory Data Analysis и линейная регрессия
### Оценивание и штрафы
Каждая из задач имеет определенную «стоимость» (указана в скобках около задачи). Максимально допустимая оценка за работу — 10 баллов. Проверяющий имеет право снизить оценку за неэффективную реализацию или неопрятные графики.
**Обратите внимание**, что в каждом разделе домашнего задания есть оцениваниемые задачи и есть вопросы. Вопросы дополняют задачи и направлены на то, чтобы проинтерпретировать или обосновать происходящее. Код без интерпретации не имеет смысла, поэтому отвечать на вопросы обязательно — за отсутствие ответов мы будем снижать баллы за задачи. Если вы ответите на вопросы, но не напишете корректный код к соответствующим оцениваемым задачам, то баллы за такое выставлены не будут.
Сдавать задание после указанного срока сдачи нельзя. При выставлении неполного балла за задание в связи с наличием ошибок на усмотрение проверяющего предусмотрена возможность исправить работу на указанных в ответном письме условиях.
Задание выполняется самостоятельно. «Похожие» решения считаются плагиатом и все задействованные студенты (в том числе те, у кого списали) не могут получить за него больше 0 баллов (подробнее о плагиате см. на странице курса). Если вы нашли решение какого-то из заданий (или его часть) в открытом источнике, необходимо указать ссылку на этот источник в отдельном блоке в конце вашей работы (скорее всего вы будете не единственным, кто это нашел, поэтому чтобы исключить подозрение в плагиате, необходима ссылка на источник).
### Формат сдачи
Задания сдаются через систему Anytask. Инвайт можно найти на странице курса. Присылать необходимо ноутбук с выполненным заданием. Сам ноутбук называйте в формате homework-practice-02-linregr-Username.ipynb, где Username — ваша фамилия.
## 1. KNN (4 балла)
### Задание 1: Визуализация решающих поверхностей в kNN.
В этом задании мы изобразим решающую поверхность для классификатора kNN, чтобы наглядно увидеть, как классификатор принимает решения для новых объектов. Для простоты будем работать со встроенным в `sklearn` набором данных `wine`, содержащим информацию о характеристиках трёх видов вина. Описание набора можно найти [здесь](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_wine.html#sklearn.datasets.load_wine) и [здесь](https://rdrr.io/cran/rattle.data/man/wine.html).
Загрузим набор данных и сохраним информацию о признаках в переменную `X`, а о зависимой переменной – в переменную `y`.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.datasets import load_wine
data = load_wine()
X = pd.DataFrame(data['data'], columns = data['feature_names'])
y = data['target']
X.head(8)
```
**Задача 1.1 (0.5 балла)** Есть ли в наборе данных пропущенные значения? Если да, то удалите их. Есть ли в наборе данных категориальные переменные? Если да, то закодируйте их при помощи OneHot-кодирования.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Задача 1.2 (0.5 балла)** Используя функцию `train_test_split()`, разделите выборку на тренировочную и тестовую, и долю тестовой выборки задайте равной 0.3. Так как разбиение осуществляется случайным образом, не забудьте зафиксировать `np.random.seed()` для воспроизводимости результатов.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Задача 1.3 (1 балл)** На тренировочной выборке обучите шесть классификаторов kNN, отличающихся только числом соседей. Для первого классификатора число соседей поставьте равным 1, для второго - 3, для третьего – 5, для четвертого – 10, для пятого – 15 и для шестого – 25 (обратите внимание на параметр `n_neighbours` класса `KNeighborsClassifier`). Для обучения используйте только два признака: `alcohol` и `magnesium` – и евклидово расстояние. Не забудьте масштабировать признаки, например, при помощи модуля `StandardScaler`.
Выведите долю правильных ответов на тренировочной и тестовой выборках для каждого классификатора.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Задача 1.4 (0 баллов)** Установите библиотеку `mlxtend` командой ниже. Библиотеку также можно установить из терминала при помощи `pip` или `conda`, как указано [здесь](http://rasbt.github.io/mlxtend/installation/).
```
!pip install mlxtend
```
Если всё прошло успешно, то в выводе команды выше вы увидите сообщение вроде "successfully installed", а следующая ячейка выполнится без ошибок.
```
import mlxtend
```
**Задача 1.5 (1.5 балл)** Библиотека `mlxtend` позволяет достаточно просто визуализировать решающие поверхности обученных классификаторов. Изучите [документацию](http://rasbt.github.io/mlxtend/user_guide/plotting/plot_decision_regions/) библиотеки и найдите, как можно построить несколько графиков решающих поверхностей на сетке (decision regions grid). Постройте такую сетку графиков для обученных выше классификаторов.
**Подсказки:**
1. Вы можете использовать готовый код, приведённый в документации, и адаптировать его для нашего случая.
2. Вам могут понадобиться дополнительные библиотеки, которые используются в примере из документации.
3. Обратите внимание на то, как нужно изменить параметры `gridspec.GridSpec()` и `itertools.product()` для нашего числа классификаторов.
4. В функции `plot_decision_region()` используйте `y_train` и нужные столбцы из `X_train`. Возможно, их придётся перевести в формат массива `numpy`.
5. Если в задаче 1.3 вы сохраните обученные классификаторы в список, то не будет необходимости обучать их заново.
6. Построение графика может занять некоторое время – придётся немного подождать!
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Задача 1.6 (0.5 балла)** Прокомментируйте результаты, полученные в задачах 1.3 и 1.5. Какое число соседей оптимально использовать для обучения классификатора? Поясните ваш выбор при помощи описания геометрии данных и получаемой решающей поверхности.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
## 2.Exploratory Data Analysis и линейная регрессия (6 баллов)
### О задании
В этом задании мы попытаемся научиться анализировать данные и выделять из них полезные признаки. Мы также научимся пользоваться `seaborn` и `sklearn`, а заодно привыкнем к основным понятиям машинного обучения.
В этом ноутбуке используется библиотека `folium` для визуализации карт. Она работает в google colab!
```
!pip install folium
import folium
m = folium.Map(location=(55.7522200, 37.6155600), zoom_start=10)
m
```
Если вы всё сделали правильно, то выше должна открыться карта Москвы.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
sns.set(style="darkgrid")
```
## Часть 0. Подготовка (1 балл)
**Задание 1 (1 балл)**. Мы будем работать с данными из соревнования [New York City Taxi Trip Duration](https://www.kaggle.com/c/nyc-taxi-trip-duration/overview), в котором нужно было предсказать длительность поездки на такси. Скачайте обучающую выборку из этого соревнования и загрузите ее:
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Обратите внимание на колонки `pickup_datetime` и `dropoff_datetime`. `dropoff_datetime` был добавлена организаторами только в обучающую выборку, то есть использовать эту колонку нельзя, давайте удалим ее. В `pickup_datetime` записаны дата и время начала поездки. Чтобы с ней было удобно работать, давайте преобразуем даты в `datetime`-объекты
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
В колонке `trip_duration` записано целевое значение, которое мы хотим предсказывать. Давайте посмотрим на распределение таргета в обучающей выборке. Для этого нарисуйте его гистограмму:
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Вопрос**: Что можно сказать о целевой переменной по гистограмме её значений?
В соревновании в качестве метрики качества использовалось RMSLE:
$$\text{RMSLE}(X, y, a) = \sqrt{\frac{1}{\ell}\sum_{i=1}^{\ell} \big(\log{(y_i + 1)} - \log{(a(x_i) + 1)}\big)^2}$$
**Вопрос**: Как вы думаете, почему авторы соревнования выбрали именно RMSLE, а не RMSE?
На семинаре мы рассматривали несколько моделей линейной регрессии в `sklearn`, но каждая из них оптимизировала среднеквадратичную ошибку (MSE), а не RMSLE. Давайте проделаем следующий трюк: будем предсказывать не целевую переменную, а ее *логарифм*. Обозначим $\hat{y}_i = \log{(y_i + 1)}$ — модифицированный таргет, а $\hat{a}(x_i)$ — предсказание модели, которая обучалась на $\hat{y}_i$, то есть логарифм таргета. Чтобы предсказать исходное значение, мы можем просто взять экспоненту от нашего предсказания: $a(x_i) = \exp(\hat{a}(x_i)) - 1$.
**Вопрос**: Покажите, что оптимизация RMSLE для модели $a$ эквивалентна оптимизации MSE для модели $\hat{a}$.
**Доказательство**: ╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
Итак, мы смогли свести задачу оптимизации RMSLE к задаче оптимизации MSE, которую мы умеем решать! Кроме того, у логарифмирования таргета есть еще одно полезное свойство. Чтобы его увидеть, добавьте к нашей выборке колонку `log_trip_duration` (воспользуйтесь `np.log1p`) и нарисуйте гистограмму модифицированного таргета по обучающей выборке. Удалите колонку со старым таргетом.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Чтобы иметь некоторую точку отсчета, давайте посчитаем значение метрики при наилучшем константном предсказании:
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
## Часть 1. Изучаем `pickup_datetime` (3 балла)
**Задание 2 (0.25 баллов)**. Для начала давайте посмотрим, сколько всего было поездок в каждый из дней. Постройте график зависимости количества поездок от дня в году (например, можно воспользоваться `sns.countplot`):
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Вопрос**: Вы, вероятно, заметили, что на графике есть 2 периода с аномально маленькими количествами поездок. Вычислите, в какие даты происходили эти скачки вниз и найдите информацию о том, что происходило в эти дни в Нью-Йорке.
Нарисуйте графики зависимости количества поездок от дня недели и от часов в сутках (воспользуйтесь `sns.relplot`):
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Задание 3 (0.5 баллов)**. Нарисуйте на одном графике зависимости количества поездок от часа в сутках для разных месяцев (разные кривые, соответствующие разным месяцам, окрашивайте в разные цвета, воспользуйтесь `hue` в `sns.relplot`). Аналогично нарисуйте зависимости количества поездок от часа в сутках для разных дней недели.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Вопрос**: Какие выводы можно сделать, основываясь на графиках выше? Выделяются ли какие-нибудь дни недели? Месяца? Время суток? С чем это связано?
**Задание 4 (0.5 баллов)**. Разбейте выборку на обучающую и тестовую в отношении 7:3. По обучающей выборке нарисуйте график зависимости среднего логарифма времени поездки от дня недели. Затем сделайте то же самое, но для часа в сутках и дня в году.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Вопрос**: Похожи ли графики зависимости таргета от дня недели и от часа в сутках на аналогичные графики для количества поездок? Почему? Что происходит со средним таргетом в те два аномальных периода, что мы видели выше? Почему так происходит? Наблюдаете ли вы какой-нибудь тренд на графике зависимости `log_trip_duration` от номера дня в году?
Добавьте следующие признаки на основе `pickup_datetime`:
1. День недели
2. Месяц
3. Час
4. Является ли период аномальным (два бинарных признака, соответствующие двум аномальным периодам)
5. Номер дня в году
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Итак, мы уже создали некоторое количество признаков.
**Вопрос**: Какие из признаков стоит рассматривать как категориальные, а какие - как численные? Почему?
**Задание 5 (1.75 баллов)**. Обучите `Ridge`-регрессию с параметрами по умолчанию, закодировав все категориальные признаки с помощью `OneHotEncoder`. Численные признаки отмасштабируйте с помощью `StandardScaler`. Используйте только признаки, которые мы выделили в этой части задания.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
## Часть 2. Изучаем координаты (2 балла)
Мы уже очень хорошо изучили данные о времени начала поездки, давайте теперь посмотрим на информацию о координатах начала и конца поездки. Мы подготовили для вас функцию, которая на карте рисует точки начала или конца поездки. Примеры ее вызова вы найдете ниже. Обратите внимание, что в эту функцию мы передаем лишь небольшой кусочек данных, посколько иначе функция будет работать очень долго
```
def show_circles_on_map(data, latitude_column, longitude_column, color):
"""
The function draws map with circles on it.
The center of the map is the mean of coordinates passed in data.
data: DataFrame that contains columns latitude_column and longitude_column
latitude_column: string, the name of column for latitude coordinates
longitude_column: string, the name of column for longitude coordinates
color: string, the color of circles to be drawn
"""
location = (data[latitude_column].mean(), data[longitude_column].mean())
m = folium.Map(location=location)
for _, row in data.iterrows():
folium.Circle(
radius=100,
location=(row[latitude_column], row[longitude_column]),
color=color,
fill_color=color,
fill=True
).add_to(m)
return m
show_circles_on_map(df.sample(1000), "pickup_latitude", "pickup_longitude", "blue")
show_circles_on_map(df.sample(1000), "dropoff_latitude", "dropoff_longitude", "blue")
```
**Вопрос**: Какие две точки выделяются на карте?
**Задание 6 (1.5 балл)**. Как мы все прекрасно помним, $t = s / v_{\text{ср}}$, поэтому очевидно, что самым сильным признаком будет расстояние, которое необходимо проехать. Мы не можем посчитать точное расстояние, которое необходимо преодолеть такси, но мы можем его оценить, посчитав кратчайшее расстояние между точками начала и конца поездки. Чтобы корректно посчитать расстояние между двумя точками на Земле, можно использовать функцию `haversine`. Также можно воспользоваться кодом с первого семинара. Посчитайте кратчайшее расстояние для объектов и запишите его в колонку `haversine`:
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Так как мы предсказываем логарифм времени поездки и хотим, чтобы наши признаки были линейно зависимы с этой целевой переменной, нам нужно логарифмировать расстояние: $\log t = \log s - \log{v_{\text{ср}}}$. Запишите логарифм `haversine` в отдельную колонку:
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Убедимся, что логарифм расстояния лучше коррелирует с нашим таргетом, чем просто расстояние:
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Задание 7 (1.5 балла)**. Давайте изучим среднюю скорость движения такси. Посчитайте среднюю скорость для каждого объекта обучающей выборки, разделив `haversine` на `trip_duration`, и нарисуйте гистограмму ее распределения
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Как можно видеть по гистограмме, для некоторых объектов у нас получились очень больше значения скоростей. Нарисуйте гистограмму по объектам, для которых значение скорости получилось разумным (например, можно не включать рассмотрение объекты, где скорость больше некоторой квантили):
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Для каждой пары (день недели, час суток) посчитайте медиану скоростей. Нарисуйте с помощью `sns.heatmap` график, где по осям будут дни недели и часы, а в качестве значения функции - медиана скорости
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Не забудьте удалить колонку со значением скорости из данных!
**Вопрос**: Почему значение скорости нельзя использовать во время обучения?
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Вопрос**: Посмотрите внимательно на график и скажите, в какие моменты времени скорость минимальна; максимальна.
Создайте признаки "поездка совершается в период пробок" и "поездка совершается в период свободных дорог" (естественно, они не должен зависеть от скорости!):
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Задание 8 (0.5 балла)**. Как уже было замечено выше, на карте выделяются две точки вдали от Манхэттена. Для каждой из них добавьте в выборку два признака: началась ли поездка в ней и закончилась ли она в ней.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Для каждого из созданных признаков нарисуйте "ящик с усами" (`sns.boxplot`) распределения логарифма времени поездки
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Вопрос**: судя по графикам, как вы думаете, хорошими ли получились эти признаки?
**Задание 10 (0.5 балла)**. Обучите `Ridge`-регрессию со стандартными параметрами на признаках, которые мы выделили к текущему моменту. Категориальные признаки закодируйте через one-hot-кодирование, числовые признаки отмасштабируйте.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
## Часть 3. Улучшаем модель (бонус 2 балла)
**Задание 13 (1 балл)**. В наших данных есть нетипичные объекты: с аномально маленьким времени поездки, с очень большим пройденным расстоянием или очень большими остатками регрессии. В этом задании предлагается исключить такие объекты из обучающей выборки. Для этого нарисуйте гистограммы распределения упомянутых выше величин, выберите объекты, которые можно назвать выбросами, и очистите обучающую выборку от них.
Отметим, что хотя эти объекты и выглядят как выбросы, в тестовой выборке тоже скорее всего будут объекты с такими же странными значениями целевой переменной и/или признаков. Поэтому, возможно, чистка обучающей выборки приведёт к ухудшению качества на тесте. Тем не менее, всё равно лучше удалять выбросы из обучения, чтобы модель получалась более разумной и интерпретируемой.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Обучите модель на очищенных данных и посчитайте качество на тестовой выборке.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Попробуйте обучить не `Ridge`-, а `Lasso`-регрессию. Какой метод лучше?
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Разбейте обучающую выборку на обучающую и валидационную в отношении 8:2. По валидационной выборке подберите оптимальные значения параметра регуляризации (по логарифмической сетке) для `Ridge` и `Lasso`, на тестовой выборке измерьте качество лучшей полученной модели.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Для каждого перебранного `alpha` для Lasso посчитайте количество нулевых весов в модели и нарисуйте график зависимости его от `alpha`. Как сильно придётся потерять в качестве, если мы хотим с помощью Lasso избавиться хотя бы от половины признаков?
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
**Задание 16 (бонус, 1 балл)**. Где, как не для нашей задачи, считать манхэттенское расстояние?
**Вопрос**: Найдите, что такое манхэттенское расстояние и почему оно так называется. Как оно нам может помочь?
Введите систему координат на нашей карте так, чтобы оси были параллельны улицам Манхэттена, и добавьте сначала в данные признак "манхэттенское расстояние между пунктом отправления и пунктом назначения", а затем и логарифм этого признака. Посчитайте корреляцию между вашим новыми признаком и таргетом; между `log_haversine` и таргетом. В каком случае корреляция больше?
Нарисуйте карту, где покажете выбранные оси. Чтобы мы могли проверить вашу работу, просьба сделать скрин этой карты и приложить картинку (если мы откроем ваш ноутбук, виджеты отображаться не будут).
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Заново обучите модель на новых даннных и посчитайте качество на тестовой выборке. Стало ли лучше? Объясните полученный результат.
```
#╰( ͡° ͜ʖ ͡° )つ──☆*:・゚
```
Вставьте картинку, описывающую ваш опыт выполнения этого ДЗ.
| github_jupyter |
# Variational Auto Encoders using Ignite
This is a tutorial on using Ignite to train neural network models, setup experiments and validate models.
In this experiment, we'll be replicating [Auto-Encoding Variational Bayes](https://arxiv.org/abs/1312.6114) by Kingma and Welling. This paper uses an encoder-decoder architecture to encode images to a vector and then reconstruct the images.
We want to be able to encode and reconstruct MNIST images. MNIST is the classic machine learning dataset, it contains black and white images of digits 0 to 9. There are 50000 training images and 10000 test images. The dataset comprises of image and label pairs.
We'll be using PyTorch to create the model, torchvision to import data and Ignite to train and monitor the models!
Please note that a lot of this code has been borrowed from [official PyTorch example](https://github.com/pytorch/examples/tree/master/vae). Similar to that it uses ReLUs and the adam optimizer, instead of sigmoids and adagrad.
Let's get started!
## Required Dependencies
In this example we only need `torchvision` package, assuming that `torch` and `ignite` are already installed. We can install it using `pip`:
```
pip install torchvision
```
```
!pip install pytorch-ignite torchvision
```
## Import Libraries
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
We import `torch`, `nn` and `functional` modules to create our models! `DataLoader` to create iterators for the downloaded datasets.
The code below also checks whether there are GPUs available on the machine and assigns the device to GPU if there are.
```
import torch
from torch.utils.data import DataLoader
from torch import nn, optim
from torch.nn import functional as F
SEED = 1234
torch.manual_seed(SEED)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
```
`torchvision` is a library that provides multiple datasets for computer vision tasks. Below we import the following:
* **MNIST**: A module to download the MNIST datasets.
* **save_image**: Saves tensors as images.
* **make_grid**: Takes a concatenation of tensors and makes a grid of images.
* **ToTensor**: Converts images to Tensors.
* **Compose**: Collects transformations.
```
from torchvision.datasets import MNIST
from torchvision.utils import save_image, make_grid
from torchvision.transforms import Compose, ToTensor
```
`Ignite` is a High-level library to help with training neural networks in PyTorch. It comes with an `Engine` to setup a training loop, various metrics, handlers and a helpful contrib section!
Below we import the following:
* **Engine**: Runs a given process_function over each batch of a dataset, emitting events as it goes.
* **Events**: Allows users to attach functions to an `Engine` to fire functions at a specific event. Eg: `EPOCH_COMPLETED`, `ITERATION_STARTED`, etc.
* **MeanSquaredError**: Metric to calculate mean squared error.
* **Loss**: General metric that takes a loss function as a parameter, calculate loss over a dataset.
* **RunningAverage**: General metric to attach to Engine during training.
* **ModelCheckpoint**: Handler to checkpoint models.
```
from ignite.engine import Engine, Events
from ignite.metrics import MeanSquaredError, Loss, RunningAverage
```
## Processing Data
Below the only transformation we use is to convert convert the images to Tensor, MNIST downloads the dataset on to your machine.
* `train_data` is a list of tuples of image tensors and labels. `val_data` is the same, just a different number of images.
* `image` is a 28 x 28 tensor with 1 channel, meaning a 28 x 28 grayscale image.
* `label` is a single integer value, denoting what the image is showing.
```
data_transform = Compose([ToTensor()])
train_data = MNIST(download=True, root="/tmp/mnist/", transform=data_transform, train=True)
val_data = MNIST(download=True, root="/tmp/mnist/", transform=data_transform, train=False)
image = train_data[0][0]
label = train_data[0][1]
print ('len(train_data) : ', len(train_data))
print ('len(val_data) : ', len(val_data))
print ('image.shape : ', image.shape)
print ('label : ', label)
img = plt.imshow(image.squeeze().numpy(), cmap='gray')
```
Now let's setup iterators of the training and validation datasets. We can take advantage of PyTorch's `DataLoader` that allows us to specify the dataset, batch size, number of workers, device, and other helpful parameters.
Let's see what the output of the iterators are:
* We see that each batch consists of 32 images and their corresponding labels.
* Examples are shuffled.
* Data is placed on GPU if available, otherwise it uses CPU.
```
kwargs = {'num_workers': 1, 'pin_memory': True} if device == 'cuda' else {}
train_loader = DataLoader(train_data, batch_size=32, shuffle=True, **kwargs)
val_loader = DataLoader(val_data, batch_size=32, shuffle=True, **kwargs)
for batch in train_loader:
x, y = batch
break
print ('x.shape : ', x.shape)
print ('y.shape : ', y.shape)
```
To visualize how well our model reconstruct images, let's save the above value of x as a set of images we can use to compare against the generation reconstructions from our model.
```
fixed_images = x.to(device)
```
## VAE Model
VAE is a model comprised of fully connected layers that take a flattened image, pass them through fully connected layers reducing the image to a low dimensional vector. The vector is then passed through a mirrored set of fully connected weights from the encoding steps, to generate a vector of the same size as the input.
```
class VAE(nn.Module):
def __init__(self):
super(VAE, self).__init__()
self.fc1 = nn.Linear(784, 400)
self.fc21 = nn.Linear(400, 20)
self.fc22 = nn.Linear(400, 20)
self.fc3 = nn.Linear(20, 400)
self.fc4 = nn.Linear(400, 784)
def encode(self, x):
h1 = F.relu(self.fc1(x))
return self.fc21(h1), self.fc22(h1)
def reparameterize(self, mu, logvar):
std = torch.exp(0.5*logvar)
eps = torch.randn_like(std)
return eps.mul(std).add_(mu)
def decode(self, z):
h3 = F.relu(self.fc3(z))
return torch.sigmoid(self.fc4(h3))
def forward(self, x):
mu, logvar = self.encode(x)
z = self.reparameterize(mu, logvar)
return self.decode(z), mu, logvar
```
## Creating Model, Optimizer and Loss
Below we create an instance of the VAE model. The model is placed on a device and then loss functions of Binary Cross Entropy + KL Divergence is used and Adam optimizer are setup.
```
model = VAE().to(device)
optimizer = optim.Adam(model.parameters(), lr=1e-3)
def kld_loss(x_pred, x, mu, logvar):
# see Appendix B from VAE paper:
# Kingma and Welling. Auto-Encoding Variational Bayes. ICLR, 2014
# https://arxiv.org/abs/1312.6114
# 0.5 * sum(1 + log(sigma^2) - mu^2 - sigma^2)
return -0.5 * torch.sum(1 + logvar - mu.pow(2) - logvar.exp())
bce_loss = nn.BCELoss(reduction='sum')
```
## Training and Evaluating using Ignite
### Trainer Engine - process_function
Ignite's `Engine` allows user to define a `process_function` to process a given batch, this is applied to all the batches of the dataset. This is a general class that can be applied to train and validate models! A `process_function` has two parameters engine and batch.
Let's walk through what the function of the trainer does:
* Sets model in train mode.
* Sets the gradients of the optimizer to zero.
* Generate `x` from batch.
* Flattens `x` into shape `(-1, 784)`.
* Performs a forward pass to reconstuct `x` as `x_pred` using model and x. Model also return `mu`, `logvar`.
* Calculates loss using `x_pred`, `x`, `logvar` and `mu`.
* Performs a backward pass using loss to calculate gradients for the model parameters.
* model parameters are optimized using gradients and optimizer.
* Returns scalar loss.
Below is a single operation during the trainig process. This process_function will be attached to the training engine.
```
def process_function(engine, batch):
model.train()
optimizer.zero_grad()
x, _ = batch
x = x.to(device)
x = x.view(-1, 784)
x_pred, mu, logvar = model(x)
BCE = bce_loss(x_pred, x)
KLD = kld_loss(x_pred, x, mu, logvar)
loss = BCE + KLD
loss.backward()
optimizer.step()
return loss.item(), BCE.item(), KLD.item()
```
### Evaluator Engine - process_function
Similar to the training process function, we setup a function to evaluate a single batch. Here is what the `eval_function` does:
* Sets model in eval mode.
* Generates `x` from batch.
* With `torch.no_grad()`, no gradients are calculated for any succeding steps.
* Flattens `x` into shape `(-1, 784)`.
* Performs a forward pass to reconstuct `x` as `x_pred` using model and x. Model also return `mu`, `logvar`.
* Returns `x_pred`, `x`, `mu` and `logvar`.
Ignite suggests attaching metrics to evaluators and not trainers because during the training the model parameters are constantly changing and it is best to evaluate model on a stationary model. This information is important as there is a difference in the functions for training and evaluating. Training returns a single scalar loss. Evaluating returns `y_pred` and `y` as that output is used to calculate metrics per batch for the entire dataset.
All metrics in `Ignite` require `y_pred` and `y` as outputs of the function attached to the `Engine`.
```
def evaluate_function(engine, batch):
model.eval()
with torch.no_grad():
x, _ = batch
x = x.to(device)
x = x.view(-1, 784)
x_pred, mu, logvar = model(x)
kwargs = {'mu': mu, 'logvar': logvar}
return x_pred, x, kwargs
```
### Instantiating Training and Evaluating Engines
Below we create 2 engines, a `trainer` and `evaluator` using the functions defined above. We also define dictionaries to keep track of the history of the metrics on the training and validation sets.
```
trainer = Engine(process_function)
evaluator = Engine(evaluate_function)
training_history = {'bce': [], 'kld': [], 'mse': []}
validation_history = {'bce': [], 'kld': [], 'mse': []}
```
### Metrics - RunningAverage, MeanSquareError and Loss
To start, we'll attach a metric of `RunningAverage` to track a running average of the scalar `loss`, `binary cross entropy` and `KL Divergence` output for each batch.
```
RunningAverage(output_transform=lambda x: x[0]).attach(trainer, 'loss')
RunningAverage(output_transform=lambda x: x[1]).attach(trainer, 'bce')
RunningAverage(output_transform=lambda x: x[2]).attach(trainer, 'kld')
```
Now there are two metrics that we want to use for evaluation - `mean squared error`, `binary cross entropy` and `KL Divergence`. If you noticed earlier, out `eval_function` returns `x_pred`, `x` and a few other values, `MeanSquaredError` only expects two values per batch.
For each batch, the `engine.state.output` will be `x_pred`, `x` and `kwargs`, this is why we use `output_transform` to only extract values from `engine.state.output` based on the the metric need.
As for `Loss`, we pass our defined `loss_function` and simply attach it to the `evaluator` as `engine.state.output` outputs all the parameters needed for `loss_function`.
```
MeanSquaredError(output_transform=lambda x: [x[0], x[1]]).attach(evaluator, 'mse')
Loss(bce_loss, output_transform=lambda x: [x[0], x[1]]).attach(evaluator, 'bce')
Loss(kld_loss).attach(evaluator, 'kld')
```
### Attaching Custom Functions to Engine at specific Events
Below you'll see ways to define your own custom functions and attaching them to various `Events` of the training process.
The first method involves using a decorator, the syntax is simple - `@` `trainer.on(Events.EPOCH_COMPLETED)`, means that the decorated function will be attached to the `trainer` and called at the end of each epoch.
The second method involves using the `add_event_handler` method of `trainer` - `trainer.add_event_handler(Events.EPOCH_COMPLETED, custom_function)`. This achieves the same result as the above.
The function below print the loss during training at the end of each epoch.
```
@trainer.on(Events.EPOCH_COMPLETED)
def print_trainer_logs(engine):
avg_loss = engine.state.metrics['loss']
avg_bce = engine.state.metrics['bce']
avg_kld = engine.state.metrics['kld']
print("Trainer Results - Epoch {} - Avg loss: {:.2f} Avg bce: {:.2f} Avg kld: {:.2f}"
.format(engine.state.epoch, avg_loss, avg_bce, avg_kld))
```
The function below prints the logs of the `evaluator` and updates the history of metrics for training and validation datasets, we see that it takes parameters `DataLoader` and `mode`. Using this way we are repurposing a function and attaching it twice to the `trainer`, once to evaluate of the training dataset and other on the validation dataset.
```
def print_logs(engine, dataloader, mode, history_dict):
evaluator.run(dataloader, max_epochs=1)
metrics = evaluator.state.metrics
avg_mse = metrics['mse']
avg_bce = metrics['bce']
avg_kld = metrics['kld']
avg_loss = avg_bce + avg_kld
print(
mode + " Results - Epoch {} - Avg mse: {:.2f} Avg loss: {:.2f} Avg bce: {:.2f} Avg kld: {:.2f}"
.format(engine.state.epoch, avg_mse, avg_loss, avg_bce, avg_kld))
for key in evaluator.state.metrics.keys():
history_dict[key].append(evaluator.state.metrics[key])
trainer.add_event_handler(Events.EPOCH_COMPLETED, print_logs, train_loader, 'Training', training_history)
trainer.add_event_handler(Events.EPOCH_COMPLETED, print_logs, val_loader, 'Validation', validation_history)
```
The function below uses the set of images (`fixed_images`) and the VAE model to generate reconstructed images, the images are then formed into a grid, saved to your local machine and displayed in the notebook below. We attach this function to the start of the training process and at the end of each epoch, this way we'll be able to visualize how much better the model gets at reconstructing images.
```
def compare_images(engine, save_img=False):
epoch = engine.state.epoch
reconstructed_images = model(fixed_images.view(-1, 784))[0].view(-1, 1, 28, 28)
comparison = torch.cat([fixed_images, reconstructed_images])
if save_img:
save_image(comparison.detach().cpu(), 'reconstructed_epoch_' + str(epoch) + '.png', nrow=8)
comparison_image = make_grid(comparison.detach().cpu(), nrow=8)
fig = plt.figure(figsize=(5, 5));
output = plt.imshow(comparison_image.permute(1, 2, 0));
plt.title('Epoch ' + str(epoch));
plt.show();
trainer.add_event_handler(Events.STARTED, compare_images, save_img=False)
trainer.add_event_handler(Events.EPOCH_COMPLETED(every=5), compare_images, save_img=False)
```
### Run Engine
Next, we'll run the `trainer` for 10 epochs and monitor results.
```
e = trainer.run(train_loader, max_epochs=20)
```
### Plotting Results
Below we see plot the metrics collected on the training and validation sets. We plot the history of `Binary Cross Entropy`, `Mean Squared Error` and `KL Divergence`.
```
plt.plot(range(20), training_history['bce'], 'dodgerblue', label='training')
plt.plot(range(20), validation_history['bce'], 'orange', label='validation')
plt.xlim(0, 20);
plt.xlabel('Epoch')
plt.ylabel('BCE')
plt.title('Binary Cross Entropy on Training/Validation Set')
plt.legend();
plt.plot(range(20), training_history['kld'], 'dodgerblue', label='training')
plt.plot(range(20), validation_history['kld'], 'orange', label='validation')
plt.xlim(0, 20);
plt.xlabel('Epoch')
plt.ylabel('KLD')
plt.title('KL Divergence on Training/Validation Set')
plt.legend();
plt.plot(range(20), training_history['mse'], 'dodgerblue', label='training')
plt.plot(range(20), validation_history['mse'], 'orange', label='validation')
plt.xlim(0, 20);
plt.xlabel('Epoch')
plt.ylabel('MSE')
plt.title('Mean Squared Error on Training/Validation Set')
plt.legend();
```
| github_jupyter |
# Analysis of single-cell transcriptomics
This tutorial demonstrates how to analyze single-cell transcriptomics data using LANTSA including
* Clustering & visualization
* Cell type marker genes
```
import numpy as np
import pandas as pd
import scanpy as sc
import matplotlib.pyplot as plt
import lantsa
```
## Read the data
We will use an annotated single-cell transcriptomics dataset from [Alex Pollen et al.](http://dx.doi.org/10.1038/nbt.2967), which can be downloaded [here](https://s3.amazonaws.com/scrnaseq-public-datasets/manual-data/pollen/NBT_hiseq_linear_tpm_values.txt).
Firstly, we read the data table and convert it into an [AnnData](https://anndata.readthedocs.io/en/latest/anndata.AnnData.html) object.
```
X = pd.read_table('./data/NBT_hiseq_linear_tpm_values.txt', index_col=0).T
celltype = X.index.str.split('_', expand=True).to_frame().to_numpy()[:, 1]
adata = sc.AnnData(X)
adata.obs['celltype'] = pd.Categorical(celltype)
adata
```
## Preprocessing
Then, we perform basic preprocessing including log transformation and finding highly variable genes.
```
sc.pp.log1p(adata)
sc.pp.highly_variable_genes(adata, n_top_genes=2000, flavor='seurat')
```
## Subspace analysis
Since this is a small dataset, we do not need to select landmarks.
Now, we perform subspace analysis to learn representation for clustering.
```
lantsa.subspace_analysis(adata, Lambda=0.1, n_neighbors=40)
```
## Clustering and visualization
The resulting `adata` is compatible with [scanpy.tl.leiden()](https://scanpy.readthedocs.io/en/stable/generated/scanpy.tl.leiden.html) for clustering and [scanpy.tl.umap()](https://scanpy.readthedocs.io/en/stable/generated/scanpy.tl.umap.html) for visualization.
```
sc.tl.leiden(adata, resolution=2.5, neighbors_key='subspace_analysis')
sc.pp.pca(adata)
sc.tl.umap(adata, init_pos='random', neighbors_key='subspace_analysis')
```
We visualize the inferred clusters in UMAP space.
```
fig, axs = plt.subplots(figsize=(8, 7))
sc.pl.umap(
adata,
color="leiden",
size=200,
palette=sc.pl.palettes.default_102,
legend_loc='right margin',
show=False,
ax=axs,
)
plt.tight_layout()
```
We also visualize the annotated cell types in UMAP space.
```
fig, axs = plt.subplots(figsize=(8, 7))
sc.pl.umap(
adata,
color="celltype",
size=200,
palette=sc.pl.palettes.default_102,
legend_loc='right margin',
show=False,
ax=axs,
)
plt.tight_layout()
```
## Cell type marker genes
Lastly, we compute the differentially expressed (DE) genes of each cell type for visualization.
```
sc.tl.rank_genes_groups(adata, groupby='celltype', method='t-test')
sc.pl.rank_genes_groups(adata, n_genes=20, ncols=3, sharey=False)
marker_genes = pd.DataFrame(adata.uns['rank_genes_groups']['names']).iloc[:10,:]
marker_genes
```
Now, we focus on a specific cell type, here BJ for demonstration.
We visualize the expression levels of the first-ranked DE gene of BJ in UMAP space.
```
fig, axs = plt.subplots(figsize=(8, 7))
sc.pl.umap(
adata,
color=marker_genes.iloc[0,2],
size=200,
palette=sc.pl.palettes.default_20,
legend_loc='right margin',
show=False,
ax=axs,
)
plt.tight_layout()
```
Then, we visualize the expression pattern of top 3 DE genes of each cell type.
```
fig, axs = plt.subplots(figsize=(9, 10))
sc.pl.dotplot(
adata,
var_names=marker_genes.iloc[0:3, :].to_numpy().T.reshape(-1),
groupby='celltype',
expression_cutoff=5,
dot_min=0.8,
swap_axes=True,
show=False,
ax=axs,
)
plt.tight_layout()
```
| github_jupyter |
```
import numpy as np
from mlp.layers import BatchNormalizationLayer
test_inputs = np.array([[-1.38066782, -0.94725498, -3.05585424, 2.28644454, 0.85520889,
0.10575624, 0.23618609, 0.84723205, 1.06569909, -2.21704034],
[ 0.11060968, -0.0747448 , 0.56809029, 2.45926149, -2.28677816,
-0.9964566 , 2.7356007 , 1.98002308, -0.39032315, 1.46515481]])
test_grads_wrt_outputs = np.array([[-0.43857052, 1.00380109, -1.18425494, 0.00486091, 0.21470207,
-0.12179054, -0.11508482, 0.738482 , -1.17249238, 0.69188295],
[ 1.07802015, 0.69901145, 0.81603688, -1.76743026, -1.24418692,
-0.65729963, -0.50834305, -0.49016145, 1.63749743, -0.71123104]])
#produce BatchNorm fprop and bprop
activation_layer = BatchNormalizationLayer(input_dim=10)
beta = np.array(10*[0.3])
gamma = np.array(10*[0.5])
activation_layer.params = [gamma, beta]
BN_fprop = activation_layer.fprop(test_inputs)
BN_bprop = activation_layer.bprop(
test_inputs, BN_fprop, test_grads_wrt_outputs)
BN_grads_wrt_params = activation_layer.grads_wrt_params(
test_inputs, test_grads_wrt_outputs)
true_fprop_outputs = np.array([[-0.1999955 , -0.19998686, -0.19999924, -0.1996655 , 0.79999899,
0.79999177, -0.1999984 , -0.19999221, 0.79999528, -0.19999926],
[ 0.7999955 , 0.79998686, 0.79999924, 0.7996655 , -0.19999899,
-0.19999177, 0.7999984 , 0.79999221, -0.19999528, 0.79999926]])
assert BN_fprop.shape == true_fprop_outputs.shape, (
'Layer bprop returns incorrect shaped array. '
'Correct shape is \n\n{0}\n\n but returned shape is \n\n{1}.'
.format(true_fprop_outputs.shape, BN_fprop.shape)
)
assert np.allclose(np.round(BN_fprop, decimals=2), np.round(true_fprop_outputs, decimals=2)), (
'Layer bprop does not return correct values. '
'Correct output is \n\n{0}\n\n but returned output is \n\n{1}\n\n difference is \n\n{2}'
.format(true_fprop_outputs, BN_fprop, BN_fprop-true_fprop_outputs)
)
print("Batch Normalization F-prop test passed")
true_bprop_outputs = np.array([[ -9.14558020e-06, 9.17665617e-06, -8.40575535e-07,
6.85384297e-03, 9.40668131e-07, 7.99795574e-06,
5.03719464e-07, 1.69038704e-05, -1.82061629e-05,
5.62083224e-07],
[ 9.14558020e-06, -9.17665617e-06, 8.40575535e-07,
-6.85384297e-03, -9.40668131e-07, -7.99795574e-06,
-5.03719464e-07, -1.69038704e-05, 1.82061629e-05,
-5.62083224e-07]])
assert BN_bprop.shape == true_bprop_outputs.shape, (
'Layer bprop returns incorrect shaped array. '
'Correct shape is \n\n{0}\n\n but returned shape is \n\n{1}.'
.format(true_bprop_outputs.shape, BN_bprop.shape)
)
assert np.allclose(np.round(BN_bprop, decimals=2), np.round(true_bprop_outputs, decimals=2)), (
'Layer bprop does not return correct values. '
'Correct output is \n\n{0}\n\n but returned output is \n\n{1}\n\n difference is \n\n{2}'
.format(true_bprop_outputs, BN_bprop, BN_bprop-true_bprop_outputs)
)
print("Batch Normalization B-prop test passed")
grads_wrt_gamma, grads_wrt_beta = BN_grads_wrt_params
true_grads_wrt_gamma = np.array(([ 1.51657703, -0.30478163, 2.00028878, -1.77110552, 1.45888603,
0.53550028, -0.39325697, -1.2286243 , -2.8099633 , -1.40311192]))
true_grads_wrt_beta = np.array([ 0.63944963, 1.70281254, -0.36821806, -1.76256935, -1.02948485,
-0.77909018, -0.62342786, 0.24832055, 0.46500505, -0.01934809])
assert grads_wrt_gamma.shape == true_grads_wrt_gamma.shape, (
'Layer bprop returns incorrect shaped array. '
'Correct shape is \n\n{0}\n\n but returned shape is \n\n{1}.'
.format(true_grads_wrt_gamma.shape, grads_wrt_gamma.shape)
)
assert np.allclose(np.round(grads_wrt_gamma, decimals=2), np.round(true_grads_wrt_gamma, decimals=2)), (
'Layer bprop does not return correct values. '
'Correct output is \n\n{0}\n\n but returned output is \n\n{1}\n\n difference is \n\n{2}'
.format(true_grads_wrt_gamma, grads_wrt_gamma, grads_wrt_gamma-true_grads_wrt_gamma)
)
assert grads_wrt_beta.shape == true_grads_wrt_beta.shape, (
'Layer bprop returns incorrect shaped array. '
'Correct shape is \n\n{0}\n\n but returned shape is \n\n{1}.'
.format(true_grads_wrt_beta.shape, grads_wrt_beta.shape)
)
assert np.allclose(np.round(grads_wrt_beta, decimals=2), np.round(true_grads_wrt_beta, decimals=2)), (
'Layer bprop does not return correct values. '
'Correct output is \n\n{0}\n\n but returned output is \n\n{1}\n\n difference is \n\n{2}'
.format(true_grads_wrt_beta, grads_wrt_beta, grads_wrt_beta-true_grads_wrt_beta)
)
print("Batch Normalization grads wrt to params test passed")
```
| github_jupyter |
# Задание 2.2 - Введение в PyTorch
Для этого задания потребуется установить версию PyTorch 1.0
https://pytorch.org/get-started/locally/
В этом задании мы познакомимся с основными компонентами PyTorch и натренируем несколько небольших моделей.<br>
GPU нам пока не понадобится.
Основные ссылки:
https://pytorch.org/tutorials/beginner/deep_learning_60min_blitz.html
https://pytorch.org/docs/stable/nn.html
https://pytorch.org/docs/stable/torchvision/index.html
```
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision.datasets as dset
from torch.utils.data.sampler import SubsetRandomSampler, Sampler
from torchvision import transforms
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
device = torch.device("cuda:0") # Let's make sure GPU is available!
```
## Как всегда, начинаем с загрузки данных
PyTorch поддерживает загрузку SVHN из коробки.
```
# First, lets load the dataset
data_train = dset.SVHN('./data/', split='train', download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=[0.43,0.44,0.47],
std=[0.20,0.20,0.20])
])
)
data_test = dset.SVHN('./data/', split='test', download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=[0.43,0.44,0.47],
std=[0.20,0.20,0.20])
]))
#plt.imshow(data_train.__getitem__(5)[0].permute(1, 2, 0))
```
Теперь мы разделим данные на training и validation с использованием классов `SubsetRandomSampler` и `DataLoader`.
`DataLoader` подгружает данные, предоставляемые классом `Dataset`, во время тренировки и группирует их в батчи.
Он дает возможность указать `Sampler`, который выбирает, какие примеры из датасета использовать для тренировки. Мы используем это, чтобы разделить данные на training и validation.
Подробнее: https://pytorch.org/tutorials/beginner/data_loading_tutorial.html
```
batch_size = 64
data_size = data_train.data.shape[0]
validation_split = .2
split = int(np.floor(validation_split * data_size))
indices = list(range(data_size))
np.random.shuffle(indices)
train_indices, val_indices = indices[split:], indices[:split]
train_sampler = SubsetRandomSampler(train_indices)
val_sampler = SubsetRandomSampler(val_indices)
train_loader = torch.utils.data.DataLoader(data_train, batch_size=batch_size,
sampler=train_sampler)
val_loader = torch.utils.data.DataLoader(data_train, batch_size=batch_size,
sampler=val_sampler)
```
В нашей задаче мы получаем на вход изображения, но работаем с ними как с одномерными массивами. Чтобы превратить многомерный массив в одномерный, мы воспользуемся очень простым вспомогательным модулем `Flattener`.
```
sample, label = data_train[0]
print("SVHN data sample shape: ", sample.shape)
# As you can see, the data is shaped like an image
# We'll use a special helper module to shape it into a tensor
class Flattener(nn.Module):
def forward(self, x):
batch_size, *_ = x.shape
return x.view(batch_size, -1)
```
И наконец, мы создаем основные объекты PyTorch:
- `model` - собственно, модель с нейросетью
- `loss` - функцию ошибки, в нашем случае `CrossEntropyLoss`
- `optimizer` - алгоритм оптимизации, в нашем случае просто `SGD`
```
def display_history(loss_history, train_history, val_history):
plt.figure(figsize=(14, 5))
plt.subplot('121')
plt.title("Train/Validation accuracy")
plt.plot(train_history, label='train')
plt.plot(val_history, label='val')
plt.legend()
plt.subplot('122')
plt.title("Loss")
plt.plot(loss_history, label='loss')
plt.legend();
model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.ReLU(inplace=True),
nn.Linear(100, 10)
)
model.type(torch.cuda.FloatTensor)
model.to(device)
loss = nn.CrossEntropyLoss().type(torch.cuda.FloatTensor)
optimizer = optim.SGD(model.parameters(), lr=1e-2, weight_decay=1e-1)
```
## Тренируем!
Ниже приведена функция `train_model`, реализующая основной цикл тренировки PyTorch.
Каждую эпоху эта функция вызывает функцию `compute_accuracy`, которая вычисляет точность на validation, эту последнюю функцию предлагается реализовать вам.
```
# This is how to implement the same main train loop in PyTorch. Pretty easy, right?
def train_model(model, train_loader, val_loader, loss, optimizer, num_epochs, scheduler=None):
loss_history = []
train_history = []
val_history = []
for epoch in range(num_epochs):
if scheduler is not None:
scheduler.step()
model.train() # Enter train mode
loss_accum = 0
correct_samples = 0
total_samples = 0
for i_step, (x, y) in enumerate(train_loader):
x_gpu = x.to(device)
y_gpu = y.to(device)
prediction = model(x_gpu)
loss_value = loss(prediction, y_gpu)
optimizer.zero_grad()
loss_value.backward()
optimizer.step()
_, indices = torch.max(prediction, 1)
correct_samples += torch.sum(indices == y_gpu)
total_samples += y.shape[0]
loss_accum += loss_value
ave_loss = loss_accum / (i_step + 1)
train_accuracy = float(correct_samples) / total_samples
val_accuracy = compute_accuracy(model, val_loader)
loss_history.append(float(ave_loss))
train_history.append(train_accuracy)
val_history.append(val_accuracy)
print("Epoch: %d, Average loss: %f, Train accuracy: %f, Val accuracy: %f" %
(epoch+1, ave_loss, train_accuracy, val_accuracy))
return loss_history, train_history, val_history
def compute_accuracy(model, loader):
"""
Computes accuracy on the dataset wrapped in a loader
Returns: accuracy as a float value between 0 and 1
"""
model.eval() # Evaluation mode
# TODO: Implement the inference of the model on all of the batches from loader,
# and compute the overall accuracy.
# Hint: torch doesn't have a dedicated argmax function,
# but you can use torch.max instead (see the documentation).
correct_samples = 0
total_samples = 0
for i_step, (x, y) in enumerate(loader):
x_gpu = x.to(device)
y_gpu = y.to(device)
prediction = model(x_gpu)
_, indices = torch.max(prediction, 1)
correct_samples += torch.sum(indices == y_gpu)
total_samples += y.shape[0]
return float(correct_samples) / total_samples
loss_history, train_history, val_history = train_model(model, train_loader, val_loader, loss, optimizer, 20)
display_history(loss_history, train_history, val_history)
```
## После основного цикла
Посмотрим на другие возможности и оптимизации, которые предоставляет PyTorch.
Добавьте еще один скрытый слой размера 100 нейронов к модели
```
# Since it's so easy to add layers, let's add some!
# TODO: Implement a model with 2 hidden layers of the size 100
model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.ReLU(inplace=True),
nn.Linear(100, 100),
nn.ReLU(inplace=True),
nn.Linear(100, 10)
)
model.type(torch.cuda.FloatTensor)
model.to(device)
optimizer = optim.SGD(model.parameters(), lr=1e-2, weight_decay=1e-1)
loss_history, train_history, val_history = train_model(model, train_loader, val_loader, loss, optimizer, 20)
```
Добавьте слой с Batch Normalization
```
# We heard batch normalization is powerful, let's use it!
# TODO: Add batch normalization after each of the hidden layers of the network, before or after non-linearity
# Hint: check out torch.nn.BatchNorm1d
model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.BatchNorm1d(100),
nn.ReLU(inplace=True),
nn.Linear(100, 100),
nn.BatchNorm1d(100),
nn.ReLU(inplace=True),
nn.Linear(100, 10),
)
model.type(torch.cuda.FloatTensor)
model.to(device)
optimizer = optim.SGD(model.parameters(), lr=1e-3, weight_decay=1e-1)
loss_history, train_history, val_history = train_model(model, train_loader, val_loader, loss, optimizer, 20)
```
Добавьте уменьшение скорости обучения по ходу тренировки.
```
# Learning rate annealing
# Reduce your learning rate 2x every 2 epochs
# Hint: look up learning rate schedulers in PyTorch. You might need to extend train_model function a little bit too!
model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.BatchNorm1d(100),
nn.ReLU(inplace=True),
nn.Linear(100, 100),
nn.BatchNorm1d(100),
nn.ReLU(inplace=True),
nn.Linear(100, 10),
)
model.type(torch.cuda.FloatTensor)
model.to(device)
optimizer = optim.SGD(model.parameters(), lr=1e-3, weight_decay=1e-2)
scheduler = optim.lr_scheduler.LambdaLR(optimizer, lr_lambda=lambda epoch: .98**((epoch+1) // 2))
loss_history, train_history, val_history = train_model(model, train_loader, val_loader,
loss, optimizer, 20, scheduler)
```
# Визуализируем ошибки модели
Попробуем посмотреть, на каких изображениях наша модель ошибается.
Для этого мы получим все предсказания модели на validation set и сравним их с истинными метками (ground truth).
Первая часть - реализовать код на PyTorch, который вычисляет все предсказания модели на validation set.
Чтобы это сделать мы приводим код `SubsetSampler`, который просто проходит по всем заданным индексам последовательно и составляет из них батчи.
Реализуйте функцию `evaluate_model`, которая прогоняет модель через все сэмплы validation set и запоминает предсказания модели и истинные метки.
```
class SubsetSampler(Sampler):
r"""Samples elements with given indices sequentially
Arguments:
indices (ndarray): indices of the samples to take
"""
def __init__(self, indices):
self.indices = indices
def __iter__(self):
return (self.indices[i] for i in range(len(self.indices)))
def __len__(self):
return len(self.indices)
def evaluate_model(model, dataset, indices):
"""
Computes predictions and ground truth labels for the indices of the dataset
Returns:
predictions: np array of ints - model predictions
grount_truth: np array of ints - actual labels of the dataset
"""
model.eval() # Evaluation mode
# TODO: Evaluate model on the list of indices and capture predictions
# and ground truth labels
# Hint: SubsetSampler above could be useful!
#raise Exception("Not implemented")
sampler = SubsetSampler(indices)
loader = torch.utils.data.DataLoader(dataset, batch_size=batch_size, sampler=sampler)
predictions = []
ground_truth = []
for i_step, (x, y) in enumerate(loader):
x_gpu = x.to(device)
y_gpu = y.to(device)
prediction = model(x_gpu)
_, ind = torch.max(prediction, 1)
predictions += list(ind.cpu().data.numpy())
ground_truth += list(y.data.numpy())
return np.array(predictions), np.array(ground_truth)
# Evaluate model on validation
predictions, gt = evaluate_model(model, data_train, val_indices)
assert len(predictions) == len(val_indices)
assert len(gt) == len(val_indices)
assert gt[100] == data_train[val_indices[100]][1]
assert np.any(np.not_equal(gt, predictions))
```
## Confusion matrix
Первая часть визуализации - вывести confusion matrix (https://en.wikipedia.org/wiki/Confusion_matrix ).
Confusion matrix - это матрица, где каждой строке соответствуют классы предсказанный, а столбцу - классы истинных меток (ground truth). Число с координатами `i,j` - это количество сэмплов класса `j`, которые модель считает классом `i`.

Для того, чтобы облегчить вам задачу, ниже реализована функция `visualize_confusion_matrix` которая визуализирует такую матрицу.
Вам осталось реализовать функцию `build_confusion_matrix`, которая ее вычислит.
Результатом должна быть матрица 10x10.
```
def visualize_confusion_matrix(confusion_matrix):
"""
Visualizes confusion matrix
confusion_matrix: np array of ints, x axis - predicted class, y axis - actual class
[i][j] should have the count of samples that were predicted to be class i,
but have j in the ground truth
"""
# Adapted from
# https://stackoverflow.com/questions/2897826/confusion-matrix-with-number-of-classified-misclassified-instances-on-it-python
assert confusion_matrix.shape[0] == confusion_matrix.shape[1]
size = confusion_matrix.shape[0]
fig = plt.figure(figsize=(10,10))
plt.title("Confusion matrix")
plt.ylabel("predicted")
plt.xlabel("ground truth")
res = plt.imshow(confusion_matrix, cmap='GnBu', interpolation='nearest')
cb = fig.colorbar(res)
plt.xticks(np.arange(size))
plt.yticks(np.arange(size))
for i, row in enumerate(confusion_matrix):
for j, count in enumerate(row):
plt.text(j, i, count, fontsize=14, horizontalalignment='center', verticalalignment='center')
def build_confusion_matrix(predictions, ground_truth):
"""
Builds confusion matrix from predictions and ground truth
predictions: np array of ints, model predictions for all validation samples
ground_truth: np array of ints, ground truth for all validation samples
Returns:
np array of ints, (10,10), counts of samples for predicted/ground_truth classes
"""
confusion_matrix = np.zeros((10,10), np.int)
# TODO: Implement filling the prediction matrix
data = np.vstack((predictions, ground_truth)).T
for i in range(10):
for j in range(10):
confusion_matrix[i][j] = len(data[(data[:, 0] == i) & (data[:, 1] == j)])
return confusion_matrix
confusion_matrix = build_confusion_matrix(predictions, gt)
visualize_confusion_matrix(confusion_matrix)
```
Наконец, посмотрим на изображения, соответствующие некоторым элементам этой матрицы.
Как и раньше, вам дана функция `visualize_images`, которой нужно воспрользоваться при реализации функции `visualize_predicted_actual`. Эта функция должна вывести несколько примеров, соответствующих заданному элементу матрицы.
Визуализируйте наиболее частые ошибки и попробуйте понять, почему модель их совершает.
```
data_train_images = dset.SVHN('./data/', split='train')
def visualize_images(indices, data, title='', max_num=10):
"""
Visualizes several images from the dataset
indices: array of indices to visualize
data: torch Dataset with the images
title: string, title of the plot
max_num: int, max number of images to display
"""
to_show = min(len(indices), max_num)
fig = plt.figure(figsize=(10,1.5))
fig.suptitle(title)
for i, index in enumerate(indices[:to_show]):
plt.subplot(1,to_show, i+1)
plt.axis('off')
sample = data[index][0]
plt.imshow(sample)
def visualize_predicted_actual(predicted_class, gt_class, predictions, ground_truth, val_indices, val_data):
"""
Visualizes images of a ground truth class which were predicted as the other class
predicted: int 0-9, index of the predicted class
gt_class: int 0-9, index of the ground truth class
predictions: np array of ints, model predictions for all validation samples
ground_truth: np array of ints, ground truth for all validation samples
val_indices: np array of ints, indices of validation samples
"""
# TODO: Implement visualization using visualize_images above
# predictions and ground_truth are provided for validation set only, defined by val_indices
# Hint: numpy index arrays might be helpful
# https://docs.scipy.org/doc/numpy/user/basics.indexing.html#index-arrays
# Please make the title meaningful!
#raise Exception("Not implemented")
data = np.vstack((val_indices, predictions, ground_truth)).T
indices = data[(data[:, 1] == predicted_class) & (data[:, 2] == gt_class)][:,0]
title = f'Failing samples. Predicted: {predicted_class}, Actual: {gt_class}'
visualize_images(indices, val_data, title=title, max_num=10)
visualize_predicted_actual(6, 8, predictions, gt, np.array(val_indices), data_train_images)
visualize_predicted_actual(1, 7, predictions, gt, np.array(val_indices), data_train_images)
```
# Переходим к свободным упражнениям!
Натренируйте модель как можно лучше - экспериментируйте сами!
Что следует обязательно попробовать:
- перебор гиперпараметров с помощью валидационной выборки
- другие оптимизаторы вместо SGD
- изменение количества слоев и их размеров
- наличие Batch Normalization
Но ограничиваться этим не стоит!
Точность на валидацонной выборке должна быть доведена до **60%**
За лучший результат в группе вы получите дополнительные баллы :)
```
# Experiment here!
#Best: Epoch: 2, Average loss: 0.273843, Train accuracy: 0.921066, Val accuracy: 0.869702 (pretrained)
model = nn.Sequential(
Flattener(),
#---
nn.Linear(3*32*32, 200),
nn.BatchNorm1d(200),
nn.ReLU(inplace=True),
#---
nn.Linear(200, 200),
nn.BatchNorm1d(200),
nn.ReLU(inplace=True),
#---
nn.Linear(200, 10),
)
model.type(torch.cuda.FloatTensor)
model.to(device)
optimizer = optim.Adam(model.parameters(), lr=2.5e-4, weight_decay=2e-4)
scheduler = optim.lr_scheduler.LambdaLR(optimizer, lr_lambda=lambda epoch: .985**((epoch+1) // 2))
loss_history, train_history, val_history = train_model(model, train_loader, val_loader,
loss, optimizer, 20, scheduler)
#Epoch: 1, Average loss: 1.422093, Train accuracy: 0.554073, Val accuracy: 0.707802
#Epoch: 5, Average loss: 0.661490, Train accuracy: 0.793025, Val accuracy: 0.800833
#Epoch: 10, Average loss: 0.524439, Train accuracy: 0.835324, Val accuracy: 0.834278
#Epoch: 15, Average loss: 0.462645, Train accuracy: 0.854622, Val accuracy: 0.845744
#Epoch: 20, Average loss: 0.412639, Train accuracy: 0.869245, Val accuracy: 0.847997
optimizer = optim.Adam(model.parameters(), lr=5e-5, weight_decay=4e-4)
loss_history, train_history, val_history = train_model(model, train_loader, val_loader, loss, optimizer, 3)
optimizer = optim.Adam(model.parameters(), lr=1e-5, weight_decay=8e-4)
loss_history, train_history, val_history = train_model(model, train_loader, val_loader, loss, optimizer, 3)
optimizer = optim.Adam(model.parameters(), lr=5e-6, weight_decay=1.6e-3)
loss_history, train_history, val_history = train_model(model, train_loader, val_loader, loss, optimizer, 3)
optimizer = optim.Adam(model.parameters(), lr=.5e-6, weight_decay=1.6e-3)
loss_history, train_history, val_history = train_model(model, train_loader, val_loader, loss, optimizer, 3)
# Как всегда, в конце проверяем на test set
#Best: 8503
test_loader = torch.utils.data.DataLoader(data_test, batch_size=batch_size)
test_accuracy = compute_accuracy(model, test_loader)
print("Test accuracy: %2.4f" % test_accuracy)
predictions, gt = evaluate_model(model, data_train, val_indices)
confusion_matrix = build_confusion_matrix(predictions, gt)
visualize_confusion_matrix(confusion_matrix)
visualize_predicted_actual(5, 3, predictions, gt, np.array(val_indices), data_train_images)
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import sys
from pathlib import Path
sys.path.append(str(Path('.').resolve().parents[0]))
from pprint import pprint
from collections import Counter
import numpy as np
import pandas as pd
import sklearn
from imblearn.under_sampling import RandomUnderSampler, NearMiss, EditedNearestNeighbours, RepeatedEditedNearestNeighbours, AllKNN
from sklearn.metrics import make_scorer
from sklearn.model_selection import GridSearchCV, StratifiedKFold, cross_validate
from xgboost import XGBClassifier, plot_importance
from src import timer
from src.data import FeatherReadWriter, JDataTrainTestReadWriter
from src.models import monkey_patch
from src.models.metrics import get_jdata_scoring, jdata_fscorer
import matplotlib
matplotlib.use('Agg')
%matplotlib inline
from matplotlib import pyplot
pd.set_option('display.max_columns', 10000)
monkey_patch.run()
```
## choose an appropriate sampler and sampling ratio by the submission
```
def get_X_y(dataset):
X = dataset.drop(columns=["user_id", "label"]).fillna(-1).values
y = dataset.label.values
return X, y
def sampler(name, ratio, random_state=0, return_indices=True, **kwargs):
if name == "rus":
sampler = RandomUnderSampler(
ratio=ratio,
return_indices=return_indices,
random_state=random_state,
**kwargs,
)
elif name == "nm":
sampler = NearMiss(
ratio=ratio,
return_indices=return_indices,
random_state=random_state,
**kwargs,
)
elif name == "enn":
sampler = EditedNearestNeighbours(
return_indices=return_indices, random_state=random_state, **kwargs
)
elif name == "renn":
sampler = RepeatedEditedNearestNeighbours(
return_indices=return_indices, random_state=random_state, **kwargs
)
elif name == "allknn":
sampler = AllKNN(
return_indices=return_indices, random_state=random_state, **kwargs
)
elif name == "tl":
sampler = TomekLinks(
return_indices=return_indices, random_state=random_state, **kwargs
)
else:
raise ValueError
return sampler
def merge_scoring_metrics(scores, scorer):
df = pd.DataFrame(scores)
custom_metrics = scorer.get(filter=None)
for metric, scores in custom_metrics.items():
df["test_{0}".format(metric)] = scores[::2]
df["train_{0}".format(metric)] = scores[1::2]
return df
def score_whole_dataset(clf, dataset, pre_train=True):
if not ("label" in dataset):
raise ValueError("dataset must include the label column")
X, y = get_X_y(dataset)
if not pre_train:
clf.fit(X, y)
scoring, scorer = get_jdata_scoring(dataset)
scoring["custom_index"](clf, X, y, np.arange(X.shape[0]))
metrics = {}
for k, v in scorer.get(filter=None).items():
metrics["test_{}".format(k)] = v
return pd.DataFrame(metrics)
# load training dataset
frw = FeatherReadWriter()
train = frw.read(dir="processed", name=frw.extend_name("all_merged_1.0"), nthreads=4)
label = frw.read(
dir="processed", name=frw.extend_name("all_merged_1.0.label"), nthreads=4
)
train[label.columns] = label
X, y = get_X_y(train)
# load online dataset for submission
online_train = frw.read(dir="processed", name=frw.extend_name("all_merged_online"), nthreads=4)
online_label = frw.read(
dir="processed", name=frw.extend_name("all_merged_online.label"), nthreads=4
)
online_train[online_label.columns] = online_label
sampling_paras = [
("rus", 0.1),
("rus", 0.01),
("rus", 0.001),
("nm", 0.1),
("nm", 0.01),
("nm", 0.001),
("tl", None),
("enn", None),
("renn", None),
("allknn", None),
]
fpath = str(PROJECT_DIR.joinpath("reports/metrics_by_samplers.csv"))
whole_dataset_metrics = pd.DataFrame()
for method, ratio in sampling_paras:
with timer(f"method: {method}, ratio: {ratio}"):
sampler_setting = {"name": method, "ratio": ratio, "n_jobs": 4}
s = sampler(**sampler_setting)
res_X, res_y, indices = s.fit_sample(X, y)
print("Distribution of class labels after resampling {}".format(Counter(res_y)))
clf = XGBClassifier(nthread=-1)
with timer("start training"):
clf.fit(res_X, res_y, verbose=3)
score_df = score_whole_dataset(clf, online_train)
score_df = score_df.set_index([["{0}-{1}".format(method, ratio)]])
whole_dataset_metrics = pd.concat([whole_dataset_metrics, score_df])
whole_dataset_metrics.to_csv(fpath)
frw.write(
pd.DataFrame({"index": indices}),
"processed",
frw.extend_name(f"all_merged_1.0_{method}_{ratio}_indices"),
)
```
```Distribution of class labels after resampling Counter({0: 21240, 1: 2124})
method: rus, ratio: 0.1: 18.471 sec
Distribution of class labels after resampling Counter({0: 212400, 1: 2124})
method: rus, ratio: 0.01: 39.256 sec
Distribution of class labels after resampling Counter({0: 2124000, 1: 2124})
method: rus, ratio: 0.001: 654.605 sec
Distribution of class labels after resampling Counter({0: 21240, 1: 2124})
method: nm, ratio: 0.1: 866.883 sec
Distribution of class labels after resampling Counter({0: 212400, 1: 2124})
method: nm, ratio: 0.01: 798.771 sec
Distribution of class labels after resampling Counter({0: 2124000, 1: 2124})
method: nm, ratio: 0.001: 1384.717 sec```
we cannot get the results of ("tl", None), ("renn", None), ("allknn", None), ("enn", None)
TODO: maybe we should standardize our datasets to speed up these processes
finally, we choose random under sampling with ratio 0.01.
```
indices = frw.read("processed", frw.extend_name(f"all_merged_1.0_rus_0.01_indices"))
res_train = train.iloc[indices['index'], :]
res_X, res_y = get_X_y(res_train)
Counter(res_y)
```
## use cross validation to compare metrics
```
kfold = StratifiedKFold(n_splits=5, shuffle=True, random_state=41)
```
### sklearn default metrics
```
clf = XGBClassifier(nthread=-1)
scoring = {
"precision": "precision",
"recall": "recall",
"f1": "f1",
"neg_log_loss": "neg_log_loss",
"roc_auc": "roc_auc",
}
scores = cross_validate(clf, res_X, res_y, cv=kfold, scoring=scoring, return_train_score=True, verbose=1)
pd.DataFrame(scores)
```
### The difference between JData Fscore and sklearn metrics
```
clf = XGBClassifier(nthread=-1)
scoring, scorer = get_jdata_scoring(res_train)
%time scores = cross_validate(clf, res_X, res_y, cv=kfold, scoring=scoring, return_train_score=True, verbose=1)
pd.DataFrame(scores)[['test_custom_index', 'train_custom_index']]
# test metrics
pd.DataFrame(scorer.get())
merge_scoring_metrics(scores, scorer)
```
## find best estimator by gridsearchcv and use custom jdata score function
```
scoring, scorer = get_jdata_scoring(res_train)
scoring = {'custom': scoring["custom_index"]}
refit = 'custom'
clf = XGBClassifier(nthread=-1)
n_estimators = range(50, 400, 50)
param_grid = dict(n_estimators=n_estimators)
grid_search = GridSearchCV(clf, param_grid, scoring=scoring, cv=kfold, refit=refit, return_train_score=True)
%time grid_result = grid_search.fit(res_X, res_y)
# summarize results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
means = grid_result.cv_results_["mean_test_custom"]
stds = grid_result.cv_results_["std_test_custom"]
params = grid_result.cv_results_["params"]
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# Plot
pyplot.errorbar(n_estimators, means, yerr=stds)
pyplot.title("XGBoost n_estimators vs JData score")
pyplot.xlabel('n_estimators')
pyplot.ylabel('JData score')
# pd.DataFrame(grid_result.cv_results_).to_csv('../reports/search_n_estimators_all_merged_1.0_rus_0.01.csv', index=False)
pd.DataFrame(grid_result.cv_results_)
scoring, scorer = get_jdata_scoring(res_train)
scoring = {'custom': scoring["custom_index"]}
refit = 'custom'
clf = XGBClassifier(nthread=-1)
max_depth = range(1, 11, 2)
print(max_depth)
param_grid = dict(max_depth=max_depth)
grid_search = GridSearchCV(clf, param_grid, scoring=scoring, cv=kfold, refit=refit, return_train_score=True)
%time grid_result = grid_search.fit(res_X, res_y)
# summarize results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
means = grid_result.cv_results_["mean_test_custom"]
stds = grid_result.cv_results_["std_test_custom"]
params = grid_result.cv_results_["params"]
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# plot
pyplot.errorbar(max_depth, means, yerr=stds)
pyplot.title("XGBoost max_depth vs JData score")
pyplot.xlabel('max_depth')
pyplot.ylabel('JData score')
# pd.DataFrame(grid_result.cv_results_).to_csv('../reports/search_max_depth_all_merged_1.0_rus_0.01.csv', index=False)
pd.DataFrame(grid_result.cv_results_)
scoring, scorer = get_jdata_scoring(res_train)
scoring = {'custom': scoring["custom_index"]}
refit = 'custom'
clf = XGBClassifier(nthread=-1)
n_estimators = range(150, 400, 50)
max_depth = range(3, 9, 2)
param_grid = dict(max_depth=max_depth, n_estimators=n_estimators)
grid_search = GridSearchCV(clf, param_grid, scoring=scoring, cv=kfold, refit=refit, verbose=1, return_train_score=True)
%time grid_result = grid_search.fit(res_X, res_y)
# summarize results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
means = grid_result.cv_results_["mean_test_custom"]
stds = grid_result.cv_results_["std_test_custom"]
params = grid_result.cv_results_["params"]
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# plot results
scores = np.array(means).reshape(len(max_depth), len(n_estimators))
for i, value in enumerate(max_depth):
pyplot.plot(n_estimators, scores[i], label='depth: ' + str(value))
pyplot.legend()
pyplot.xlabel('n_estimators')
pyplot.ylabel('JData score')
# pd.DataFrame(grid_result.cv_results_).to_csv('../reports/search_estimators_and_max_depth_all_merged_1.0_rus_0.01.csv', index=False)
pd.DataFrame(grid_result.cv_results_).sort_values('rank_test_custom')
```
## traning whole dataset -> worse
```
param = {'max_depth': 3, 'n_estimators': 350}
clf = XGBClassifier(nthread=-1, **param)
X, y = get_X_y(train)
%time clf.fit(X, y)
clf
# use our best model to evalute result of the whole local data
scoring, scorer = get_jdata_scoring(train)
scores = scoring['custom_index'](clf, X, y, np.arange(X.shape[0]))
print(f'whole local result: {scores}')
```
## use model trained by sampling dataset to submit -> not good enough
```
param = {'max_depth': 3, 'n_estimators': 350}
clf = XGBClassifier(nthread=-1, **param)
%time clf.fit(res_X, res_y)
scoring, scorer = get_jdata_scoring(train)
scores = scoring['custom_index'](clf, X, y, np.arange(X.shape[0]))
print(f'whole local result: {scores}')
score_df = score_whole_dataset(clf, online_train)
score_df
# trytry place
# rank3
param = {'max_depth': 5, 'n_estimators': 350}
clf = XGBClassifier(nthread=-1, **param)
%time clf.fit(res_X, res_y)
scoring, scorer = get_jdata_scoring(train)
scores = scoring['custom_index'](clf, X, y, np.arange(X.shape[0]))
print(f'whole local result: {scores}')
score_df = score_whole_dataset(clf, online_train)
score_df
## 增加深度 local score 增加但是 online 分數沒有增加,代表 feature 無法提供足夠的資訊讓 model 預測?
## overfitting?
## ensemble to reduce overfitting
## sampling 3 sample datasets to ensemble results
import random
rslt = []
for i in range(3):
rs = {}
method = 'rus'
ratio = 0.01
random_state = random.randint(0, 10000)
sampler_setting = {"name": method, "ratio": ratio, "random_state": random_state}
print(sampler_setting)
s = sampler(**sampler_setting)
res_X, res_y, indices = s.fit_sample(X, y)
rs['method'] = method
rs['ratio'] = ratio
rs['random_state'] = random_state
rs['indices'] = indices
param = {'max_depth': 3, 'n_estimators': 350}
clf = XGBClassifier(nthread=-1, **param)
%time clf.fit(res_X, res_y)
rs['param'] = param
rs['model'] = clf
scoring, scorer = get_jdata_scoring(train)
scores = scoring['custom_index'](clf, X, y, np.arange(X.shape[0]))
print(f'whole local result: {scores}')
rs['scoring'] = scoring
rs['scorer'] = scorer
rslt.append(rs)
from sklearn.ensemble import VotingClassifier
eclf = VotingClassifier(estimators=[('xgb1', rslt[0]['model']), ('xgb2', rslt[1]['model']), ('xgb3', rslt[2]['model'])], voting='soft')
%time eclf = eclf.fit(res_X, res_y)
score_whole_dataset(eclf, online_train)
# TODO
# remove unseen sku?
```
# plot importance
```
feature_names = train.drop(columns=["user_id", "label"]).columns
feature_mapping = dict([('f{}'.format(i), feature_names[i]) for i in range(len(feature_names))])
from sklearn import preprocessing
def plot_importance(model, feature_mapping, n=30):
# Get xgBoost importances
importance_dict = {}
for import_type in ['weight', 'gain', 'cover']:
importance_dict['xgBoost-{}'.format(import_type)] = model.get_booster().get_score(importance_type=import_type)
# MinMax scale all importances
importance_df = pd.DataFrame(importance_dict).fillna(0)
importance_df = pd.DataFrame(
preprocessing.MinMaxScaler().fit_transform(importance_df),
columns=importance_df.columns,
index=importance_df.index
)
# replace index
importance_df.index = importance_df.index.map(mapper=feature_mapping)
# Create mean column
importance_df['mean'] = importance_df.mean(axis=1)
# Plot the feature importances
importance_df.sort_values('mean').head(n).plot(kind='bar', figsize=(20, 10))
plot_importance(clf, feature_mapping)
```
| github_jupyter |
# Test notebook Meteorites
```
from pathlib import Path
import numpy as np
import pandas as pd
import requests
from IPython.display import display
from IPython.utils.capture import capture_output
import pandas_profiling
from pandas_profiling.utils.cache import cache_file
file_name = cache_file(
"meteorites.csv",
"https://data.nasa.gov/api/views/gh4g-9sfh/rows.csv?accessType=DOWNLOAD",
)
df = pd.read_csv(file_name)
# Note: Pandas does not support dates before 1880, so we ignore these for this analysis
df["year"] = pd.to_datetime(df["year"], errors="coerce")
# Example: Constant variable
df["source"] = "NASA"
# Example: Boolean variable
df["boolean"] = np.random.choice([True, False], df.shape[0])
# Example: Mixed with base types
df["mixed"] = np.random.choice([1, "A"], df.shape[0])
# Example: Highly correlated variables
df["reclat_city"] = df["reclat"] + np.random.normal(scale=5, size=(len(df)))
# Example: Duplicate observations
duplicates_to_add = pd.DataFrame(df.iloc[0:10])
duplicates_to_add["name"] = duplicates_to_add["name"] + " copy"
df = df.append(duplicates_to_add, ignore_index=True)
# Inline report without saving
with capture_output() as out:
pr = df.profile_report(
sort=None,
html={"style": {"full_width": True}},
progress_bar=False,
minimal=True,
)
display(pr)
assert len(out.outputs) == 2
assert out.outputs[0].data["text/plain"] == "<IPython.core.display.HTML object>"
assert all(
s in out.outputs[0].data["text/html"]
for s in ["<iframe", "Profile report generated with the `pandas-profiling`"]
)
assert out.outputs[1].data["text/plain"] == ""
# There should also 2 progress bars in minimal mode
with capture_output() as out:
pfr = df.profile_report(
html={"style": {"full_width": True}},
minimal=True,
progress_bar=True,
lazy=False,
)
assert all(
any(v in s.data["text/plain"] for v in ["%|", "FloatProgress"]) for s in out.outputs
)
assert len(out.outputs) == 2
# Write to a file
with capture_output() as out:
pfr.to_file("/tmp/example.html")
assert all(
any(v in s.data["text/plain"] for v in ["%|", "FloatProgress"]) for s in out.outputs
)
assert len(out.outputs) == 2
# Print existing ProfileReport object inline
with capture_output() as out:
display(pfr)
assert len(out.outputs) == 2
assert out.outputs[0].data["text/plain"] == "<IPython.core.display.HTML object>"
assert all(
s in out.outputs[0].data["text/html"]
for s in ["<iframe", "Profile report generated with the `pandas-profiling`"]
)
assert out.outputs[1].data["text/plain"] == ""
```
| github_jupyter |
#### MicroSoft MSVC cl
```
//---------------------------
//%runinterm
//%term:c:\Windows\System32\cmd.exe /c start
//%execfile:src\test.exe
//---------------------------
//%ccompiler:cl
//%cflags: /Fe:src\test.exe /source-charset:utf-8
//%ldflags:/execution-charset:utf-8
//---------------------------
//%overwritefile
//%file:src/test.c
///%noruncode
//%log:0
//---------------------------
#include <stdio.h>
int main(){
printf("Hello World!\n");
}
##%overwritefile
##%file:print256colours.sh
##%runprg:gnome-terminal
##%runprgargs:l -- /bin/bash print256colours.sh &
##%loadurl:https://gist.github.com/danieldietrich/1606ca62a93c94dfb3b44f81303e744a/raw/e50a28ec54188d2413518788de6c6367ffcea4f7/print256colours.sh
get_char()
{
SAVEDSTTY=`stty -g`
stty -echo
stty cbreak
dd if=/dev/tty bs=1 count=1 2> /dev/null
stty -raw
stty echo
stty $SAVEDSTTY
}
echo ""
echo "Press any key to start...or Press Ctrl+c to cancel"
char=`get_char`
echo "hello world"
//%//runmode:repl
//%replsetip:Please enter text
//%file:test0.c
//%cflags: -g -o test0.out
//%runinterm
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <unistd.h>
#include <signal.h>
#include <sys/prctl.h>
int getsline(char *result)
{
int point = 0;
int word;
long i=0;
while(1)
{
i++;
usleep(15);
word = getc(stdin);//等待用户输入或从缓存中读一个字符
if(word != '\n')//读到回车符就认为一行指令被读完了
{
*result = word;//记录这个字符
result ++;
point ++;
//putc(word,stdout);
}
else
{
result = '\0';//给指针末尾添加一个结束符
result = result - point;//让指针指回字符串的头
return 0;
}
}
return 0;
}
int main()
{
// char buff[3];
// memset( buff, '\0', sizeof( buff ));
// fprintf(stdout, "Going to set full buffering on\n");
// setvbuf(stdout, buff, _IOFBF, 3);
char *line;
line = malloc(150);
// sleep(60);
// printf("3Please enter 1text\r\n");
// printf("3Please enter 2text\r\n");
printf("Please enter text\r\n");
// fflush(stdout);
getsline(line);
printf("You enter:%s\r\n",line);
// sleep(4);
// printf("3Please enter 3text\r\n");
// sleep(4);
// printf("4Please enter 4text\r\n");
// fflush(stdout);
free(line);
// sleep(60);
// prctl(PR_SET_PDEATHSIG,SIGHUP);
// raise(SIGCHLD);
return 0;
}
```
#### Cygwin GCC
```
//---------------------------
//%runinterm
///%term:c:\Windows\System32\cmd.exe /c start
//%cflags: -o src/test.exe
///%ldflags:
//----------------------------------
//%overwritefile
//%file:src/test.c
///%noruncode
//%log:0
//----------------------------------
#include <stdio.h>
int main(){
printf("Hello World!中文测试\n");
}
```
MSYS2 GCC
```
//----------------------------------
//%noruncode
//%overwritefile
//%file:inc/add.h
///%noruncode
//%log:1
//----------------------------------
int add(int,int);
//----------------------------------
//%noruncode
//%overwritefile
//%file:src/add.c
///%noruncode
//%log:1
//----------------------------------
#include <add.h>
int add(int a,int b)
{
return a+b;
}
//---------------------------
//%runinterm
///%term:c:\Windows\System32\cmd.exe /c start
//%cflags: -o src/test.exe
///%ldflags:
//----------------------------------
//%overwritefile
//%file:src/test.c
///%noruncode
//%log:0
//----------------------------------
#include <stdio.h>
#include <stdlib.h>
#include "add.h"
int main(int argc,char *argv[])//char **argv
{
int i=1;
printf("%d\n",argc);
while(*argv)
{
printf("%s\n",*argv++);
}
i=2;
i++;
i++;
i=add(i,10);
printf("i=%d\n",i);
// system("pause");
}
//%rungdb
help
//%rungdb
quit
//---------------------------
//%runinterm
//%term:c:\Windows\System32\cmd.exe /c start
//%execfile:src\test.exe
//---------------------------
//%ccompiler:cl
//%cflags: /Fe:src\test.exe /source-charset:utf-8
//%ldflags:/execution-charset:utf-8
//---------------------------
//%overwritefile
//%file:src/test.c
///%noruncode
//%log:0
//---------------------------
#include <stdio.h>
#include <string.h>
#include <locale.h>
#include <stddef.h>
#include <wchar.h>
int main(void){
setlocale(LC_ALL,"zh_CN.UTF-8");
wchar_t* s=L"你好";
wprintf(L"你好c test %s\n",s);
return 0;
}
//---------------------------
// %runinterm
// %term:mintty
//%cflags:-finput-charset=UTF-8 -o src/test.exe
//%ldflags:-fexec-charset=UTF-8
// %ldflags: -fwide-exec-charset=UTF-8
//----------------------------------
//%overwritefile
//%file:src/test.c
///%noruncode
//%log:1
//----------------------------------
#include <stdio.h>
#include <string.h>
#include <locale.h>
#include <stddef.h>
#include <wchar.h>
int main(void){
setlocale(LC_ALL,"zh_CN.UTF-8");
wprintf (L"%ls \n", L"A wide string");
wprintf (L"%ls \n", L"中文显示");
wprintf (L"%ls \n", L"chinese show 中文显示");
wchar_t* s=L"你好";
wprintf(L"你好c test %ls \n",s);
return 0;
}
```
| github_jupyter |
# Section 4: Case Study I - U-Net for Building Mapping
Now let's move in to a little advance model call U-Net. U-Net is popular in satellite image analysis (remote sensing) community. It’s very elegant and simple model that can be used to perform semantic segmentation task (labelling each pixel) well.
In this section, we will use U-Net to identify buildings from satellite images (2 class classification problem).
1) U-Net for Building Mapping
<hr>
<hr>
<hr>
```
'''first, let's import libraries '''
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.python.keras import Model
from tensorflow.python.keras.layers import Input, Conv2D, MaxPooling2D, Conv2DTranspose, Concatenate, Dropout
```
## 1) U-Net for Building Mapping
Input data are RGB satellite images. And output are binary images. Pixel value is 0 for non-buildings and pixel value is 1 for buildings.
```
'''loading data'''
# data is already randomized and split in to training / test sets. So we can go ahead and use them as it is.
x_train = np.load('./data/SpaceNet/sat_train.npy').astype('float32')
y_train= np.load('./data/SpaceNet/bul_train.npy').astype('float32')
x_test = np.load('./data/SpaceNet/sat_test.npy').astype('float32')
y_test = np.load('./data/SpaceNet/bul_test.npy').astype('float32')
print("x_train shape", x_train.shape)
print("y_train shape", y_train.shape)
print("y_test shape", x_test.shape)
print("y_test shape", y_test.shape)
# Let's plot a sample input RGB image and output image with buildings
plt.imshow(x_test[10, :, :, :].astype('uint8'))
plt.show()
plt.imshow(y_test[10, :, :, 0].astype('uint8'))
plt.show()
```
#### First little bit about U-Net
Architecture of U-Net is in following figure,
<img src="./graphics/U-Net.PNG" width="60%"/>
<sub>Source of the figure: https://arxiv.org/pdf/1505.04597.pdf</sub>
It's very elegant and symmetric architecture. U-Net was first developed for biomedical image segmentation in a paper called "U-Net: Convolutional Networks for Biomedical Image Segmentation" (https://arxiv.org/pdf/1505.04597.pdf). And later, this architecture was widely adapted in satellite image analysis (remote sensing) community.
As in our last examples, in the left side of the network (Encoder), convolutional and max pooling operations down-sample the input image. And in the right side of the network (Decoder), transpose convolutional operations with and without strides up-sample output of the Encoder producing another image with same size as input image as the final output.
Another unique thing here is each corresponding down-sampling and up-sampling outputs with same sizes are connected (skip-connections) by __*Concatenation*__ operation. This skip-connections allow gradient (information) to pass through different levels of the network efficiently, avoiding the bottleneck at the middle of the U-Net architecture. Now a days, these skip connections are widely used in various neural network architectures and they significantly improves performance of neural networks.
For skip-connections, we are using inbuilt __*Concatenate*__ function from Tensorflow (Keras) library. Due to this skip-connections, intermediate layers of the U-Net are connected. So U-Net can't be built as a __*Sequential*__ model as in out all past examples. Hence, we are using non-sequential model from Tensorflow (Keras) library for U-Net where input and output of each layer in the neural network is assigned to different variables.
Now, let's go ahead and define the U-Net model, fit, prediction and validate the output.
```
x_in = Input(shape=(128, 128, 3))
'''Encoder'''
x_temp = Conv2D(32, (3, 3), activation='relu', padding='same')(x_in)
x_temp = Dropout(0.25)(x_temp)
x_skip1 = Conv2D(32, (3, 3), activation='relu', padding='same')(x_temp)
x_temp = MaxPooling2D((2,2))(x_skip1)
x_temp = Conv2D(32, (3, 3), activation='relu', padding='same')(x_temp)
x_temp = Dropout(0.25)(x_temp)
x_skip2 = Conv2D(32, (3, 3), activation='relu', padding='same')(x_temp)
x_temp = MaxPooling2D((2,2))(x_skip2)
x_temp = Conv2D(64, (3, 3), activation='relu', padding='same')(x_temp)
x_temp = Dropout(0.25)(x_temp)
x_skip3 = Conv2D(64, (3, 3), activation='relu', padding='same')(x_temp)
x_temp = MaxPooling2D((2,2))(x_skip3)
x_temp = Conv2D(64, (3, 3), activation='relu', padding='same')(x_temp)
x_temp = Dropout(0.5)(x_temp)
x_temp = Conv2D(64, (3, 3), activation='relu', padding='same')(x_temp)
'''Decoder'''
x_temp = Conv2DTranspose(64, (3, 3), activation='relu', padding='same')(x_temp)
x_temp = Dropout(0.5)(x_temp)
x_temp = Conv2DTranspose(64, (3, 3), strides=(2, 2), activation='relu', padding='same')(x_temp)
x_temp = Concatenate()([x_temp, x_skip3])
x_temp = Conv2DTranspose(64, (3, 3), activation='relu', padding='same')(x_temp)
x_temp = Dropout(0.5)(x_temp)
x_temp = Conv2DTranspose(64, (3, 3), strides=(2, 2), activation='relu', padding='same')(x_temp)
x_temp = Concatenate()([x_temp, x_skip2])
x_temp = Conv2DTranspose(32, (3, 3), activation='relu', padding='same')(x_temp)
x_temp = Dropout(0.5)(x_temp)
x_temp = Conv2DTranspose(32, (3, 3), strides=(2, 2), activation='relu', padding='same')(x_temp)
x_temp = Concatenate()([x_temp, x_skip1])
x_temp = Conv2DTranspose(32, (3, 3), activation='relu', padding='same')(x_temp)
x_temp = Dropout(0.5)(x_temp)
x_temp = Conv2DTranspose(32, (3, 3), activation='relu', padding='same')(x_temp)
'''Use 1 by 1 Convolution to get desired output bands'''
x_temp = Conv2D(32, (1, 1), activation='relu', padding='same')(x_temp)
x_temp = Conv2D(32, (1, 1), activation='relu', padding='same')(x_temp)
x_out = Conv2D(1, (1, 1), activation='sigmoid', padding='same')(x_temp)
# use sigmoid activation here because output values are either 0 or 1
model = Model(inputs=x_in, outputs=x_out)
model.compile(loss='mean_squared_error', optimizer='adam')
model.summary()
```
Another new concept here use of multiple __*Dropout*__ layers in the middle of the network. This helps reducing over-fitting during training process. Simply it randomly drops out (ignores) portion of layer outputs (in our case 25%). This allow making robust network architecture finally reducing over-fitting. Short description about __*Dropout*__ can be seen in this nice short video.(https://www.youtube.com/watch?v=NhZVe50QwPM)
```
history = model.fit(x_train, y_train, validation_data=(x_test, y_test), epochs=100, batch_size=10, verbose=0)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
'''Prediction over the test dataset'''
pred_test = model.predict(x_test)
#let's comare random predicted and actial y values
plt.imshow(pred_test[20, :, :, 0])
plt.show()
plt.imshow(y_test[20,:,:,0])
plt.show()
```
This is not an operational model with high accuracy. But with more layers and with more data, we can develop this architecture in to an operational model with high accuracy.
| github_jupyter |
# バッチ推論サービスを作成する
健康クリニックは一日中患者の測定を取り、各患者の詳細を別々のファイルに保存すると想像してください。その後、一晩で糖尿病予測モデルを使用して、その日のすべての患者データをバッチとして処理し、翌朝待つ予測を生成し、糖尿病のリスクがあると予測される患者をフォローアップできるようにします。Azure Machine Learning では、*バッチ推論パイプライン*を作成することでこれを実現できます。そして、この演習ではそれを実施します。
## ワークスペースに接続する
作業を開始するには、ワークスペースに接続します。
> **注**: Azure サブスクリプションでまだ認証済みのセッションを確立していない場合は、リンクをクリックして認証コードを入力し、Azure にサインインして認証するよう指示されます。
```
import azureml.core
from azureml.core import Workspace
# 保存された構成ファイルからワークスペースを読み込む
ws = Workspace.from_config()
print('Ready to use Azure ML {} to work with {}'.format(azureml.core.VERSION, ws.name))
```
## モデルをトレーニングして登録する
それでは、バッチ推論パイプラインにデプロイするモデルをトレーニングして登録してみましょう。
```
from azureml.core import Experiment
from azureml.core import Model
import pandas as pd
import numpy as np
import joblib
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
# ワークスペースで Azure 実験を作成する
experiment = Experiment(workspace=ws, name='mslearn-train-diabetes')
run = experiment.start_logging()
print("Starting experiment:", experiment.name)
# 糖尿病データセットを読み込む
print("Loading Data...")
diabetes = pd.read_csv('data/diabetes.csv')
# 特徴とラベルを分離する
X, y = diabetes[['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']].values, diabetes['Diabetic'].values
# データをトレーニング セットとテスト セットに分割する
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
# デシジョン ツリー モデルをトレーニングする
print('Training a decision tree model')
model = DecisionTreeClassifier().fit(X_train, y_train)
# 精度を計算する
y_hat = model.predict(X_test)
acc = np.average(y_hat == y_test)
print('Accuracy:', acc)
run.log('Accuracy', np.float(acc))
# AUC を計算する
y_scores = model.predict_proba(X_test)
auc = roc_auc_score(y_test,y_scores[:,1])
print('AUC: ' + str(auc))
run.log('AUC', np.float(auc))
# トレーニング済みモデルを保存する
model_file = 'diabetes_model.pkl'
joblib.dump(value=model, filename=model_file)
run.upload_file(name = 'outputs/' + model_file, path_or_stream = './' + model_file)
# 実行を完了する
run.complete()
# モデルを登録する
run.register_model(model_path='outputs/diabetes_model.pkl', model_name='diabetes_model',
tags={'Training context':'Inline Training'},
properties={'AUC': run.get_metrics()['AUC'], 'Accuracy': run.get_metrics()['Accuracy']})
print('Model trained and registered.')
```
## バッチ データを生成およびアップロードする
この演習用の新しいデータを取得するために、スタッフが揃って患者がいる診療所は実在しないため、糖尿病の CSV ファイルからランダムなサンプルを生成し、そのデータを Azure Machine Learning ワークスペースのデータストアにアップロードし、そのためのデータセットを登録します。
```
from azureml.core import Datastore, Dataset
import pandas as pd
import os
# 既定のデータ ストアを設定する
ws.set_default_datastore('workspaceblobstore')
default_ds = ws.get_default_datastore()
# すべてのデータストアを列挙し、どちらが既定かを示す
for ds_name in ws.datastores:
print(ds_name, "- Default =", ds_name == default_ds.name)
# 糖尿病データを読み込む
diabetes = pd.read_csv('data/diabetes2.csv')
# 特徴列の 100 項目のサンプルを取得する (糖尿病ラベルではない)
sample = diabetes[['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']].sample(n=100).values
# フォルダーを作成する
batch_folder = './batch-data'
os.makedirs(batch_folder, exist_ok=True)
print("Folder created!")
# 各サンプルを個別のファイルとして保存する
print("Saving files...")
for i in range(100):
fname = str(i+1) + '.csv'
sample[i].tofile(os.path.join(batch_folder, fname), sep=",")
print("files saved!")
# 既定のデータストアにファイルをアップロードする
print("Uploading files to datastore...")
default_ds = ws.get_default_datastore()
default_ds.upload(src_dir="batch-data", target_path="batch-data", overwrite=True, show_progress=True)
# 入力データ用データセットを登録する
batch_data_set = Dataset.File.from_files(path=(default_ds, 'batch-data/'), validate=False)
try:
batch_data_set = batch_data_set.register(workspace=ws,
name='batch-data',
description='batch data',
create_new_version=True)
except Exception as ex:
print(ex)
print("Done!")
```
## コンピューティングを作成する
パイプラインのコンピューティング コンテキストが必要なので、次のコードを使用して Azure Machine Learning コンピューティング クラスター (存在しない場合は作成されます) を指定します。
> **重要**: 実行する前に、以下のコードで *your-compute-cluster* をコンピューティング クラスターの名前に変更してください。クラスター名は、長さが 2 〜 16 文字のグローバルに一意の名前である必要があります。英字、数字、- の文字が有効です。
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
cluster_name = "your-compute-cluster"
try:
# Check for existing compute target
inference_cluster = ComputeTarget(workspace=ws, name=cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
# If it doesn't already exist, create it
try:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_DS11_V2', max_nodes=2)
inference_cluster = ComputeTarget.create(ws, cluster_name, compute_config)
inference_cluster.wait_for_completion(show_output=True)
except Exception as ex:
print(ex)
```
> **注**: コンピューティング インスタンスとクラスターは、スタンダードの Azure 仮想マシンのイメージに基づいています。この演習では、コストとパフォーマンスの最適なバランスを実現するために、*Standard_DS11_v2* イメージが推薦されます。サブスクリプションにこのイメージを含まないクォータがある場合は、別のイメージを選択してください。 ただし、画像が大きいほどコストが高くなり、小さすぎるとタスクが完了できない場合があることに注意してください。Azure 管理者にクォータを拡張するように依頼していただくことも可能です。
## バッチ推論用パイプラインを作成する
これで、バッチ推論に使用するパイプラインを定義する準備が整いました。パイプラインではバッチ推論を実行するために Python コードが必要となるので、パイプラインで使用されるすべてのファイルを保存できるフォルダーを作成しましょう。
```
import os
# 実験ファイル用フォルダーを作成する
experiment_folder = 'batch_pipeline'
os.makedirs(experiment_folder, exist_ok=True)
print(experiment_folder)
```
次に、実際の作業を行う Python スクリプトを作成し、パイプライン フォルダーに保存します。
```
%%writefile $experiment_folder/batch_diabetes.py
import os
import numpy as np
from azureml.core import Model
import joblib
def init():
# Runs when the pipeline step is initialized
global model
# load the model
model_path = Model.get_model_path('diabetes_model')
model = joblib.load(model_path)
def run(mini_batch):
# This runs for each batch
resultList = []
# process each file in the batch
for f in mini_batch:
# Read the comma-delimited data into an array
data = np.genfromtxt(f, delimiter=',')
# Reshape into a 2-dimensional array for prediction (model expects multiple items)
prediction = model.predict(data.reshape(1, -1))
# Append prediction to results
resultList.append("{}: {}".format(os.path.basename(f), prediction[0]))
return resultList
```
パイプラインには実行する環境が必要になるため、コードが使用するパッケージを含む Conda 仕様を作成します。
```
%%writefile $experiment_folder/batch_environment.yml
name: batch_environment
dependencies:
- python=3.6.2
- scikit-learn
- pip
- pip:
- azureml-defaults
```
次に、Conda 環境を含む実行コンテキストを定義します。
```
from azureml.core import Environment
from azureml.core.runconfig import DEFAULT_CPU_IMAGE
# 実験用の環境を作成する
batch_env = Environment.from_conda_specification("experiment_env", experiment_folder + "/batch_environment.yml")
batch_env.docker.base_image = DEFAULT_CPU_IMAGE
print('Configuration ready.')
```
パイプラインを使用して、バッチ予測スクリプトを実行し、入力データから予測を生成し、結果をテキスト ファイルとして出力フォルダーに保存します。これを行うには、**ParallelRunStep** を使用して、バッチ データを並列処理し、結果を *parallel_run_step.txt* という名前の単一の出力ファイルに照合することができます。
```
from azureml.pipeline.steps import ParallelRunConfig, ParallelRunStep
from azureml.data import OutputFileDatasetConfig
from azureml.core.runconfig import DockerConfiguration
output_dir = OutputFileDatasetConfig(name='inferences')
parallel_run_config = ParallelRunConfig(
source_directory=experiment_folder,
entry_script="batch_diabetes.py",
mini_batch_size="5",
error_threshold=10,
output_action="append_row",
environment=batch_env,
compute_target=inference_cluster,
node_count=2)
parallelrun_step = ParallelRunStep(
name='batch-score-diabetes',
parallel_run_config=parallel_run_config,
inputs=[batch_data_set.as_named_input('diabetes_batch')],
output=output_dir,
arguments=[],
allow_reuse=True
)
print('Steps defined')
```
次は、ステップをパイプラインに入れて実行します。
> **注**: これには時間がかかる場合があります。
```
from azureml.core import Experiment
from azureml.pipeline.core import Pipeline
pipeline = Pipeline(workspace=ws, steps=[parallelrun_step])
pipeline_run = Experiment(ws, 'mslearn-diabetes-batch').submit(pipeline)
pipeline_run.wait_for_completion(show_output=True)
```
パイプラインの実行が終了すると、結果の予測は、パイプラインの最初の (そして唯一の) ステップに関連付けられた実験の出力に保存されます。次のように取得できます。
```
import pandas as pd
import shutil
# 以前の実行で残っている場合は、ローカル結果フォルダーを削除する
shutil.rmtree('diabetes-results', ignore_errors=True)
# 最初のステップの実行を取得し、その出力をダウンロードする
prediction_run = next(pipeline_run.get_children())
prediction_output = prediction_run.get_output_data('inferences')
prediction_output.download(local_path='diabetes-results')
# フォルダー階層を走査して、結果ファイルを見つける
for root, dirs, files in os.walk('diabetes-results'):
for file in files:
if file.endswith('parallel_run_step.txt'):
result_file = os.path.join(root,file)
# クリーンアップ出力形式
df = pd.read_csv(result_file, delimiter=":", header=None)
df.columns = ["File", "Prediction"]
# 最初の 20 件の結果を表示する
df.head(20)
```
## パイプラインを公開し、その REST インターフェイスを使用する
バッチ推論用の作業パイプラインができたので、それを公開し、REST エンドポイントを使用してアプリケーションから実行できます。
```
published_pipeline = pipeline_run.publish_pipeline(
name='diabetes-batch-pipeline', description='Batch scoring of diabetes data', version='1.0')
published_pipeline
```
公開されたパイプラインにはエンドポイントがあり、Azure portal で確認できます。また、公開されたパイプライン オブジェクトのプロパティとして見つけることもできます。
```
rest_endpoint = published_pipeline.endpoint
print(rest_endpoint)
```
エンドポイントを使用するには、クライアント アプリケーションが HTTP 経由で REST 呼び出しを行う必要があります。この要求は認証される必要があるため、Authorization ヘッダーが必要です。これをテストするには、現在の接続から Azure ワークスペースへの Authorization ヘッダーを使用します。これは、次のコードを使用して取得できます。
> **注**: 実際のアプリケーションには、認証に使用するサービス プリンシパルが必要です。
```
from azureml.core.authentication import InteractiveLoginAuthentication
interactive_auth = InteractiveLoginAuthentication()
auth_header = interactive_auth.get_authentication_header()
print('Authentication header ready.')
```
これで、REST インターフェイスを呼び出す準備ができました。パイプラインは非同期で実行されるため、識別子を取得します。実行中のパイプライン実験を追跡するために使用できます。
```
import requests
rest_endpoint = published_pipeline.endpoint
response = requests.post(rest_endpoint,
headers=auth_header,
json={"ExperimentName": "mslearn-diabetes-batch"})
run_id = response.json()["Id"]
run_id
```
実行 ID があるため、**RunDetails** ウィジェットを使用して、実行中の実験を表示できます。
```
from azureml.pipeline.core.run import PipelineRun
from azureml.widgets import RunDetails
published_pipeline_run = PipelineRun(ws.experiments['mslearn-diabetes-batch'], run_id)
# 実行が完了するまでブロックする
published_pipeline_run.wait_for_completion(show_output=True)
```
パイプラインの実行が完了するのを待ってから、以下のセルを実行して結果を確認します。
前と同様に、結果は最初のパイプライン ステップの出力にあります。
```
import pandas as pd
import shutil
# 以前の実行で残っている場合は、ローカル結果フォルダーを削除する
shutil.rmtree('diabetes-results', ignore_errors=True)
# 最初のステップの実行を取得し、その出力をダウンロードする
prediction_run = next(pipeline_run.get_children())
prediction_output = prediction_run.get_output_data('inferences')
prediction_output.download(local_path='diabetes-results')
# フォルダー階層を走査して、結果ファイルを見つける
for root, dirs, files in os.walk('diabetes-results'):
for file in files:
if file.endswith('parallel_run_step.txt'):
result_file = os.path.join(root,file)
# クリーンアップ出力形式
df = pd.read_csv(result_file, delimiter=":", header=None)
df.columns = ["File", "Prediction"]
# 最初の 20 件の結果を表示する
df.head(20)
```
これで、毎日の患者データをバッチ処理するために使用できるパイプラインが作成されました。
**詳細情報**: バッチ推論用パイプラインの使用方法の詳細については、Azure Machine Learning のドキュメントの[「バッチ予測の実行方法」](https://docs.microsoft.com/azure/machine-learning/how-to-run-batch-predictions)を参照してください。
| github_jupyter |
#$EXERCISE_PREAMBLE$
As before, don't forget to run the setup code below before jumping into question 1.
```
# SETUP. You don't need to worry for now about what this code does or how it works.
from learntools.core import binder; binder.bind(globals())
from learntools.python.ex2 import *
print('Setup complete.')
```
# Exercises
## 1.
Complete the body of the following function according to its docstring.
HINT: Python has a builtin function `round`
```
def round_to_two_places(num):
"""Return the given number rounded to two decimal places.
>>> round_to_two_places(3.14159)
3.14
"""
# Replace this body with your own code.
# ("pass" is a keyword that does literally nothing. We used it as a placeholder
# because after we begin a code block, Python requires at least one line of code)
pass
q1.check()
#%%RM_IF(PROD)%%
q1.assert_check_unattempted()
#%%RM_IF(PROD)%%
def round_to_two_places(num):
"""Return the given number rounded to two decimal places.
>>> round_to_two_places(3.14159)
3.14
"""
return round(num, 2)
q1.assert_check_passed()
#%%RM_IF(PROD)%%
def round_to_two_places(num):
"""Return the given number rounded to two decimal places.
>>> round_to_two_places(3.14159)
3.14
"""
return round(num, 3)
q1.assert_check_failed()
# Uncomment the following for a hint
#_COMMENT_IF(PROD)_
q1.hint()
# Or uncomment the following to peek at the solution
#_COMMENT_IF(PROD)_
q1.solution()
```
## 2.
The help for `round` says that `ndigits` (the second argument) may be negative.
What do you think will happen when it is? Try some examples in the following cell?
Can you think of a case where this would be useful?
```
# Put your test code here
#_COMMENT_IF(PROD)_
q2.solution()
```
## 3.
In a previous programming problem, the candy-sharing friends Alice, Bob and Carol tried to split candies evenly. For the sake of their friendship, any candies left over would be smashed. For example, if they collectively bring home 91 candies, they'll take 30 each and smash 1.
Below is a simple function that will calculate the number of candies to smash for *any* number of total candies.
Modify it so that it optionally takes a second argument representing the number of friends the candies are being split between. If no second argument is provided, it should assume 3 friends, as before.
Update the docstring to reflect this new behaviour.
```
def to_smash(total_candies):
"""Return the number of leftover candies that must be smashed after distributing
the given number of candies evenly between 3 friends.
>>> to_smash(91)
1
"""
return total_candies % 3
q3.check()
#%%RM_IF(PROD)%%
def to_smash(total_candies, n_friends=3):
return n_friends % total_candies
q3.assert_check_failed()
#%%RM_IF(PROD)%%
def to_smash(total_candies, n_friends=3):
return total_candies % n_friends
q3.assert_check_passed()
#_COMMENT_IF(PROD)_
q3.hint()
#_COMMENT_IF(PROD)_
q3.solution()
```
## 4.
It may not be fun, but reading and understanding error messages will be an important part of your Python career.
Each code cell below contains some commented-out buggy code. For each cell...
1. Read the code and predict what you think will happen when it's run.
2. Then uncomment the code and run it to see what happens. (**Tip**: In the kernel editor, you can highlight several lines and press `ctrl`+`/` to toggle commenting.)
3. Fix the code (so that it accomplishes its intended purpose without throwing an exception)
<!-- TODO: should this be autochecked? Delta is probably pretty small. -->
```
# ruound_to_two_places(9.9999)
# x = -10
# y = 5
# # Which of the two variables above has the smallest absolute value?
# smallest_abs = min(abs(x, y))
# def f(x):
# y = abs(x)
# return y
# print(f(5))
```
#$KEEP_GOING$
| github_jupyter |
Sascha Spors,
Professorship Signal Theory and Digital Signal Processing,
Institute of Communications Engineering (INT),
Faculty of Computer Science and Electrical Engineering (IEF),
University of Rostock,
Germany
# Data Driven Audio Signal Processing - A Tutorial with Computational Examples
Winter Semester 2021/22 (Master Course #24512)
- lecture: https://github.com/spatialaudio/data-driven-audio-signal-processing-lecture
- tutorial: https://github.com/spatialaudio/data-driven-audio-signal-processing-exercise
Feel free to contact lecturer frank.schultz@uni-rostock.de
## Exercise 1: Introduction
[Introduction](exercise01.ipynb)
## Exercise 2: Audio Features I (Segmentation, STFT, Spectrogram, Periodogram)
[Audio Features I](exercise02.ipynb)
## Exercise 3: Audio Features II (Segmentation, RMS/(True)Peak/Crest Factor, R128 loudness)
[Audio Features II](exercise03.ipynb)
## Exercise 4: SVD / 4 Subspaces / Left Inverse
- [SVD and 4 Subspaces](exercise04_svd.ipynb)
- [SVD and Left Inverse](exercise04_leftinv.ipynb)
- [SVD and Right Inverse](exercise04_rightinv.ipynb)
## Exercise 5: Column Space Singular Vectors of a Multitrack Audio Matrix
- [exercise05.ipynb](exercise05.ipynb)
## Exercise 6: Principal Component Analysis (PCA)
- Matlab code:
- [exercise06_pca_2D.m](exercise06_pca_2D.m)
- [exercise06_pca_3D.m](exercise06_pca_3D.m)
- Python notebooks TBD
## Exercise 7: QR, SVD, Linear Regression vs. SVD Regression
- [exercise07_QR.ipynb](exercise07_QR.ipynb)
- [exercise07_left_inverse_SVD_QR.ipynb](exercise07_left_inverse_SVD_QR.ipynb)
- [exercise07_linear_regression_LS_vs_SVD.ipynb](exercise07_linear_regression_LS_vs_SVD.ipynb)
## Exercise 8: Ridge Regression / Bias vs. Variance
- [exercise08_ridge_regression.ipynb](exercise08_ridge_regression.ipynb)
- [exercise08_bias_variance.ipynb](exercise08_bias_variance.ipynb)
## Exercise 9: Gradient Descent (Steepest Descent)
- [exercise09_gradient_descent.m](exercise09_gradient_descent.m)
## Exercise 10: Perceptron / Neural Networks
- The XOR mapping is a popular example to motivate non-linearities in models, as linear regression cannot solve this simple problem in [exercise10_xor_example.m](exercise10_xor_example.m)
- Our own implementation of simple **0/1 classification** using only **one layer** with **sigmoid activation** function [exercise10_binary_logistic_regression.py](exercise10_binary_logistic_regression.py)
We should not miss these brilliant resources to start with neural networks
- [https://pythonalgos.com/create-a-neural-network-from-scratch-in-python-3/](https://pythonalgos.com/create-a-neural-network-from-scratch-in-python-3/)
- [https://playground.tensorflow.org](https://playground.tensorflow.org)
- https://www.tensorflow.org/tutorials/keras/overfit_and_underfit (and the other tutorials found there)
## Exercise 11: Binary Classification
- With [exercise11_binary_logistic_regression_tf.py](exercise11_binary_logistic_regression_tf.py) we **compare our** above implementation [exercise10_binary_logistic_regression.py](exercise10_binary_logistic_regression.py) **against a TF model**
- Next, we create more complex models in [exercise11_binary_logistic_regression_tf_with_hidden_layers.py](exercise11_binary_logistic_regression_tf_with_hidden_layers.py) using **hidden layers**, but still with **manually tuned hyper parameters**
## Exercise 12: Multiclass Classification
- With [exercise12_MulticlassClassification_CategoricalCrossentropy.ipynb](exercise12_MulticlassClassification_CategoricalCrossentropy.ipynb) we expand the example [exercise11_binary_logistic_regression_tf_with_hidden_layers.py](exercise11_binary_logistic_regression_tf_with_hidden_layers.py) towards **classification of more than two classes** using **softmax activation** function in the output layer
- With [exercise12_HyperParameterTuning.ipynb](exercise12_HyperParameterTuning.ipynb) we introduce
- data split into train, validate, test data sets
- hyper parameter tuning
- one hot encoding
- training of best model with re-set weights using train / val data set
- final prediction on unseen test data set compared to predictions on train / val data sets
- confusion matrix and visualization of predictions
- Finally we apply all this to a music genre classification application in [exercise12_MusicGenreClassification.ipynb](exercise12_MusicGenreClassification.ipynb)
- feature design (loudness, crest, peak, rms, spectral weight)
- feature inspection / avoiding NaNs
- feature normalization
- balancing data set wrt class occurence
We could move on with dropout layers, regularization...
## Exercise 13: CNN
TBD
[exercise13_CNN.py](exercise13_CNN.py)
## Textbook Recommendations
- Gilbert **Strang**: *Linear Algebra and Learning from Data*, Wellesley, 2019, consider to buy your own copy of this brilliant book
- Gareth **James**, Daniela Witten, Trevor Hastie, Rob Tibshirani: *An Introduction to Statistical Learning* with Applications in R, Springer, 2nd ed., 2021, [free pdf e-book](https://www.statlearning.com/)
- Trevor **Hastie**, Robert Tibshirani, Jerome Friedman: *The Elements of Statistical Learning: Data Mining, Inference, and Prediction*, Springer, 2nd ed., 2009, [free pdf e-book](https://hastie.su.domains/ElemStatLearn/)
- Sergios **Theodoridis**: *Machine Learning*, Academic Press, 2nd ed., 2020, check your university library service for free pdf e-book
- Kevin P. **Murphy**: *Machine Learning*, MIT Press, 2012, check your university library service for free pdf e-book
- Marc Peter **Deisenroth**, A. Aldo Faisal, Cheng Soon Ong: *Mathemathics for Machine Learning*, Cambridge University Press, 2020, [free pdf e-book](https://mml-book.github.io/)
- Steven L. **Brunton**, J. Nathan Kutz: *Data Driven Science & Engineering - Machine Learning, Dynamical Systems, and Control*, Cambridge University Press, 2020, check your university library service for free pdf e-book
- Aurélien **Géron**: *Hands-on machine learning with Scikit-Learn, Keras and TensorFlow*. O’Reilly, 2nd ed., 2019
## Open Course Ware Recommendations
- Online Course by Andrew **Ng** et al. at https://www.coursera.org/ and https://www.deeplearning.ai/
- Online Course by Gilbert **Strang** et al. at https://ocw.mit.edu/courses/mathematics/18-065-matrix-methods-in-data-analysis-signal-processing-and-machine-learning-spring-2018/
- Online Course/Material by Aurélien **Géron** https://github.com/ageron
- Online Course by Meinard **Müller** https://www.audiolabs-erlangen.de/resources/MIR/FMP/B/B_GetStarted.html (focus on music information retrieval)
## Autorship
- University of Rostock
- Frank Schultz
- Sascha Spors
## Copyright
- the notebooks are provided as [Open Educational Resources](https://en.wikipedia.org/wiki/Open_educational_resources)
- feel free to use the notebooks for your own purposes
- the text is licensed under [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/)
- the code of the IPython examples is licensed under under the [MIT license](https://opensource.org/licenses/MIT)
- please attribute the work as follows: *Frank Schultz, Data Driven Audio Signal Processing - A Tutorial Featuring Computational Examples, University of Rostock* ideally with relevant file(s), github URL https://github.com/spatialaudio/data-driven-audio-signal-processing-exercise, commit number and/or version tag, year.
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.