text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Learning Objectives
- [ ] 3.1.1 Represent data in binary and hexadecimal forms.
- [ ] 3.1.2 Write programs to perform the conversion of positive integers between different number bases: denary, binary and hexadecimal forms; and display results.
- [ ] 3.2.1 Give examples of where or how Unicode is used.
- [ ] 3.2.2 Use ASCII code in programs.
- [ ] 3.3.9 Understand the need for privacy and integrity of data.
- [ ] 3.3.10 Describe methods to protect data.
- [ ] 3.3.11 Explain the difference between backup and archive.
- [ ] 3.3.12 Describe the need for version control and naming convention.
- [ ] 3.3.13 Explain how data in Singapore is protected under the Personal Data Protection Act to govern the collection, use and disclosure of personal data.
# References
1. Leadbetter, C., Blackford, R., & Piper, T. (2012). Cambridge international AS and A level computing coursebook. Cambridge: Cambridge University Press.
2. https://www.geeksforgeeks.org/tree-traversals-inorder-preorder-and-postorder/
3. https://www.dcode.fr/babylonian-numbers
4. https://www.joelonsoftware.com/2003/10/08/the-absolute-minimum-every-software-developer-absolutely-positively-must-know-about-unicode-and-character-sets-no-excuses/
5. CPDD
# 13.0 Representation of Positive Integers
All of us are familiar with the use of positive integers. In particular, we grew up learning how to count from a young age and (hopefully) none of us consider counting from 1 to 100 to be difficult. To count, we need to first understand the meanings of the symbols 0, 1, 2, . . ., 9 and also, the meaning of positions two identical in the integer. This way of writing out numbers is termed **positional notation / number system**. For example, when we write the number `107` to represent **one hundred and seven**, we actually mean $$1\times 10^2 +0\times 10^1 +7\times 10^0.$$
In particular, note that in our modern use of Hindu-Arabic numbers, we have 10 different symbols that we could mix and match to generate more numbers with by switching their positions in our writing. The number of such different symbols in a number system is called the **base** or **radix** of the number system. Thus, with this definition, our usual number system is base-10. This number system is also called **decimal** number system or **denary** system.
The Babylonians are generally credited as the first civilization that came up with the positional system.
<center>
<img src="https://upload.wikimedia.org/wikipedia/commons/thumb/d/d6/Babylonian_numerals.svg/1920px-Babylonian_numerals.svg.png" width="600" align="center"/><br>
By Josell7 - File:Babylonian_numerals.jpg, CC BY-SA 4.0, https://commons.wikimedia.org/w/index.php?curid=9862983
</center>
However, from the diagram of symbols above, we see that they actually have more than 10 symbols in their system. In fact, Babylonian number system is more akin to our modern base-60 system. We still see their influence in some of the measurements in our daily life like 60 seconds in a 1 minute, 60 minutes in an hour; 6 * 60 degrees in 1 revolution of a circle.
For the subsequent parts of this section, we will denote base-$k$ number with subscript when the meaning of the number is ambigous. So, for example, if $13_{10}$ is the integer `13` in base-10, $101_{2}$ is the integer `5` in base-2.
## 13.1.1 Base-2 Number System
The number system with two symbols only (0,1) is called the **binary** number system. In this system, every integer $n$ as we know it can uniquely represented by a number that consists only of 0 and 1s. For example, $2_{10}$ is $10_{2}$, $3_{10}$ is $11_{2}$.
Let $\left(a_{n-1}a_{n-2}\cdots a_{1}a_{0}\right)_{2}$ be a $n$ digit binary number, so for every $i$, 0\leq i\leq n-1, $a_i$ is either 0 or 1, that represents $x_{10}$ in denary. Then, $$x_{10}=a_{n-1}2^{n-1}+a_{n-2}2^{n-2}+\cdots+a_{1}2^{1}+a_{0}2^{0}.$$
For example,
- $3_{10}=1*2^{1}+1*2^{0}=11_{2}$,
- $65_{10}=1*2^{6}+0*2^{5}+0*2^{4}+0*2^{3}+0*2^{2}+0*2^{1}+1*2^{0}=(1000001)_{2}$.
## Exercise
- Without using Python's inbuilt converter, write a function `den_to_bin()` that takes in an integer $n$ in denary representation and returns the binary representation of the integer. E.g. `den_to_bin(65)` returns `1000001`.
- Without using Python's inbuilt converter, write a function `bin_to_den()` that takes in an integer $n$ in binary representation and returns the denary representation of the integer. E.g. `bin_to_den(1000001)` returns `65`.
```
#YOUR_ANSWER_HERE
import math
def den_to_bin(n):
highest_power=math.floor(math.log(n,2))
representation=[]
while highest_power>=0:
representation.append(n//(2**highest_power))
n=n%(2**highest_power)
highest_power-= 1
return int(''.join([str(i) for i in representation]))
den_to_bin(1025) == int(bin(1025)[2:])
def bin_to_den(n):
highest_power=len(str(n))
digits=[int(i) for i in list(str(n))]
weights=[2**(highest_power-i) for i in range(1,highest_power+1)]
return sum([digits[i]*weights[i] for i in range(highest_power)])
bin_to_den(10000000001)
den_to_bin(1025)
int('10000000001',2)
```
## 13.1.2 Base-16 Number System
The number system with 16 symbols is called the **hexadecimal** ( hex for short) number system. Hexadecimal system uses the decimal numbers and six extra symbols. There are no numerical symbols that represent values greater than nine, so letters taken from the English alphabet are used, specifically `A`, `B`, `C`, `D`, `E` and `F`. $A_{16} = 10_{10}$, $B_{16} = 11_{10}$, and so on so forth, up to $F_{16} = 15_{10}$.
Let $\left(a_{n-1}a_{n-2}\cdots a_{1}a_{0}\right)_{2}$ be a $n$ digit hexadecimal number, so for every $i$, 0\leq i\leq n-1, $a_i$ is either 0, 1, .., A,..F, that represents $x_{10}$ in denary. Then, $$x_{10}=b_{n-1}16^{n-1}+b_{n-2}16^{n-2}+\cdots+b_{1}16^{1}+b_{0}16^{0},$$
where
- $b_i=a_i$ if $0\leq a_i\leq 9$,
- Also, $b_{i}=\begin{cases}
10 & ,\text{if }a_{i}=A\\
11 & ,\text{if }a_{i}=B\\
12 & ,\text{if }a_{i}=C\\
13 & ,\text{if }a_{i}=D\\
14 & ,\text{if }a_{i}=E\\
15 & ,\text{if }a_{i}=F
\end{cases}$
For example,
- $3_{10}=3*16^{0}=3_{16}$,
- $65_{10}=4*16^{1}+1*16^{0}=41_{16}$.
- $FF_{16}=15*16^{1}+15*16^{0}=255_{10}$.
## Exercise
- Without using Python's inbuilt converter, write a function `den_to_hex()` that takes in an integer $n$ in denary representation and returns the hexadecimal representation of the integer. E.g. `den_to_hex(65)` returns `41`.
- Without using Python's inbuilt converter, write a function `hex_to_den()` that takes in an integer $n$ in hexadecimal representation and returns the denary representation of the integer. E.g. `hex_to_den(41)` returns `65`.
```
#YOUR_ANSWER_HERE
import math
def den_to_hex(n):
highest_power=math.floor(math.log(n,16))
representation=[]
while highest_power>=0:
representation.append(n//(16**highest_power))
n=n%(16**highest_power)
highest_power-= 1
symbols='0123456789ABCDEF'
representation=[symbols[i] for i in representation]
return int(''.join(representation))
#den_to_bin(1025) == int(bin(1025)[2:])
def hex_to_den(n):
highest_power=len(str(n))
digits=list
weights=[16**(highest_power-i) for i in range(1,highest_power+1)]
return sum([digits[i]*weights[i] for i in range(highest_power)])
bin_to_den(10000000001)
den_to_hex(1025)
# int('10000000001',2)
hex(1025)
```
# 13.1 Data Representation
Values are stored inside a computer as a series of 0s and 1s. A single 0 or 1 is called a **binary digit** or **bit**. Any group of 0s and 1s can be used to represent a specific character, for example, a letter. The number of bits used to store one character is called a **byte**. The complete set of characters that the computer uses is known as its **character set**. For example, the character `A` could be stored as `00000001`. The representation of characters as in terms of `0`s and `1`s are called **binary representation**.
Binary representation is a suitable representation of values in a computer as the circuits in a computer's processor are made up of billions of transistors, tiny switches that are activated by the electronic signals it receives and building blocks of modern computers. The digits 1 and 0 used in binary reflect the on and off states of a transistor.
<center>
<img src="https://image.shutterstock.com/z/stock-photo-brown-motherboard-pcb-close-up-shot-with-dual-bios-chips-surface-mount-transistors-capasitors-and-1864279093.jpg" width="300" align="center"/><br>
By Anton Bushinskiy, Shutterstock
</center>
Another commonly used representation of values in computer is hexacadecimal representation, which is intended to simplify the binary representation. Furthermore, note that each hex digit is a 4-bit binary sequence. For example, `11010100` in binary would be `D4` in hex.
A good way of thinking about the character set for a normal computer is to have a look at the characters that are available on its keyboard. For this character set, eight bits are used to store a single character. Using a byte (eight bits) for a character, we can store up to 256 unique characters (i.e. 256 different eight-bit binary values). This is enough to represent each of the characters on a standard keyboard (Most Windows based keyboard has 104 keys).
# 13.1.1 American Standard Code for Information Interchange (ASCII)
In the 1960s, a meeting in the USA agreed a standard set of codes which allows different computers to store 'standard' characters in a uniform manner. Before this convention, different machines could store same characters is stored differently. E.g, some computer designer might prefer `10000000` to store the character `A`. This caused documents created on one of the computers to be unreadable on another.
Most computer systems today use the ASCII coding system, so you can be fairly sure that when you type in `A` on any computer, it is stored in the computer’s memory as $01000001_2$ and will be read as `A` by any other computer.
These binary numbers in the ASCII coding system are termed **ASCII code points**.
The mapping of code points to a sequence of code units is termed **character encoding**.
The ASCII coding system uses **seven** (not eight) bits to represent each character and the eighth bit as a means of checking the rest (called the **parity bit**). For ASCII, these 8 bits is called a **code unit**. Parity bit for ASCII is usually found in the leading bit of the code unit. This means that 128 different characters can be represented in the standard ASCII character set.
Not all ASCII code points represents a written symbol. These code points are termed `control characters`. They are used to cause effects other than the addition of a symbol to the text. E.g.
- $00000000_2$ or $0_16$ represents `Null`, which does nothing,
- $01111111_2$ or $7F_{16}$ represents `Delete`, which is designed to erase incorrect characters.
- $01000001_2$ or $41_{16}$ represents the character `A`,
- $01000010_2$ or $42_{16}$ represents the character `B`,
- $01100001_2$ or $61_{16}$ represents the character `a`, etc.
The full list of ASCII code points can be found at https://en.wikipedia.org/wiki/ASCII
The function `ord()` in Python takes a character as a parameter and returns the *denary* representation of the ASCII code point associated with the character. For example,
>```
> ord('A')
>> 65
>```
In the other direction, there's also another function `chr()` that takes in an integer and return a character.
# Exercise
- Find the sum of the ASCII character codes for `INDONESIA`.
- Find the character represented by the integer `9786`.
```
#YOUR_ANSWER_HERE
```
# 13.1.2 Unicode and Unicode Transformation Formats (UTF)
Unicode is a more recent set of code to store characters in a uniform manner. The Unicode standard describes how characters are represented by code points. Similar to ASCII, each character has a Unicode code point associated with it. Unicode code points has the form `U+hhhh` where `h` is a hexadecimal value.
**Unicode character encoding** refers to the mapping of the Unicode code points to the memory representation of it.
The most commonly used Unicode encodings are
- `UTF-8` uses one to four-byte (8-bit) code units to represent a character.
- Used by 97% of web pages and also python
- In UTF-8, every code point from 0-127 (in denary) is stored in a single byte. Only code points 128 and above are stored using 2, 3, in fact, up to 6 bytes.
- compatible with ASCII, i.e., in other words, the character `k`, represented by $01101011_2$, in ASCII is also represented by the same byte in UTF-8.
- `UTF-16` uses one to two 16-bit code units to represent a character.
- incompatible with ASCII
The full list of Unicode characters can be found at https://en.wikipedia.org/wiki/List_of_Unicode_characters
## Example
The string `Hello` has the code points
>```
>U+0048 U+0065 U+006C U+006C U+006F
>```
If we try to store those numbers in two bytes each, we have
>```
>00 48 00 65 00 6C 00 6C 00 6F
>```
## Benefits of using Unicode
- Using 16 bits makes it possible to represent over 65,000 characters. This means that all the characters used by the world’s languages can be represented in Unicode.
- It is widely used to handle documents, particularly if a single document needs to be written in, for example, English, Arabic and Chinese.
- Unicode also allows the localisation of software, where standard software can be adapted for use by different cultures by modifying the layout and features of the interface.
## Example
Run the following print statement.
```
print("\u3053\u3093\u306B\u3061\u306F\U0001F600")
```
# 13.2 Data Management
## 13.2.1 Data Privacy
Data privacy is the requirement for data to be accessed by or disclosed to authorised persons only. It is important that unauthorised people do not have access to data they are not supposed to have. Unfortunately, in today’s digitised society, it is easier than ever to gather a person’s data and use it to obtain valuable insights, track his/her movements or to commit fraud.
For instance, data on which websites you visit often can reveal which products you are more likely to purchase as a shopper. This information can be highly valuable to an advertiser. As more services become available online, the risk of fraudent use of data increases. For instance, with a photo of your identity card, a person can impersonate you and register for a new phone line on a telco website. (Previously such a transaction would have required the person to personally register over the counter with the physical identity card.) As technology becomes increasingly powerful, machines can gather information on a person easily, like performing facial recognition on surveillance videos to track down the whereabouts of an individual in a particular area.
We want our personal data to be private so that unauthorised people cannot use our personal data for unauthorised use. In Singapore, personal data is protected under the Personal Data Protection Act (PDPA).
NB: Personal data refers to data, whether true or not, about an individual who can be identified from that data; or from that data and other information to which the organisation has or is likely to have access.
### Exercise
- Why is data privacy necessary?
- Suppose you take a picture of yourself outside your home in your school uniform, and post it on social media to show everyone that you just returned home from school. What personal information can be gathered from the photo?
```
#YOUR_ANSWER_HERE
```
## 13.2.2 Personal Data Protection Act (PDPA)
The PDPA is a data protection law comprising various rules that govern the collection, use, disclosure and care of personal data. It recognises both the rights of individuals to protect their personal data, including rights of access and correction, as well as the needs of organisations to collect, use or disclose personal data for legitimate and reasonable purposes.
It takes into account the following:
- **Consent** – Organisations must obtain an individual's knowledge and consent to collect, use or disclose his/her personal data (with some exceptions).
- **Notification** – Organisations must inform individuals of the purposes for collecting, using or disclosing their personal data.
- **Appropriateness** – Organisations may collect, use or disclose personal data only for purposes that would be considered appropriate to a reasonable person under the given circumstances.
- **Accountability** – Organisations must make information about their personal data protection policies available on request. They should also make available the business contact information of the representatives responsible for answering questions relating to the organisations’ collection, use or disclosure of personal data.
To administer and enforce the PDPA, Singapore set up the Personal Data Protection Commission (PDPC) in 2013.
### 13.2.2.1 Do Not Call Registry
Have you received calls from unknown companies who seem to know your name and perhaps try to sell products to you? Your telephone number could have been gathered from unexpected sources, such as a lucky draw form that you filled up long ago. With technology, companies can easily gather and consolidate personal information. In fact, it can even automate the making of such calls.
To prevent you from getting unnecessary marketing calls, you can register in the Do Not Call (DNC) Registry to opt out of marketing messages and calls. The PDPA prohibits organisations from sending marketing messages to Singapore telephone numbers, including mobile, fixed-line, residential and business numbers that are registered with the DNC Registry.
There are three DNC registers that individuals can choose to register in:
- No Voice Call Register
- No Text Message Register
- No Fax Message Register
Registering the phone number in each register is to opt out of receiving marketing messages through voice calls, text messages and fax messages.
Note that organisations which have an ongoing relationship with a subscriber or user of a Singapore telephone number may send marketing messages on similar or related products, services and memberships to that Singapore telephone number via text or fax without checking against the DNC Registry. However, each exempt message must also contain an opt-out facility that the recipient may use to opt out from receiving such telemarketing messages. If a recipient opts out, organisations must stop sending such messages to his/her Singapore telephone number after 30 days. This means that, for instance, you can get a marketing message from your mobile phone company even if you listed the number in the DNC Registry. However, the message will include a link for you to opt out from receiving their marketing messages. Once you opt out, the mobile phone company can no longer send you marketing messages on that number.
### 13.2.2.2 Use of NRIC
The Singapore National Registration Identification Card (NRIC) number is a unique idenfier assigned to Singapore citizens and permanent residents. Similarly, the Foreign Identification Number (FIN) is a unique identifier that is assigned to foreigners. The NRIC/FIN contains personal information about the person, such as his/her date of birth and address. As unique identifiers like NRIC and FIN are permament, irreplaceable and used in a variety of government transactions, we need to be careful with such data.
Individuals should not readily provide their NRIC/FIN and personal particulars to companies/strangers. Consent is required before organisations can obtain a person’s data. Under the PDPA, from 1 September 2019, organisations are generally not allowed to collect, use or disclose NRIC numbers (or copies of NRIC) except in the following circumstances:
- Collection, use or disclosure of NRIC numbers (or copies of NRIC) is required under the law (or an exception under the PDPA applies); or
- Collection, use or disclosure of NRIC numbers (or copies of NRIC) is necessary to accurately establish or verify the identities of the individuals to a high degree of fidelity.
For example, a medical clinic needs to see the NRIC of a patient to identify the person. The clinic will need to keep the name, address, NRIC and contact number of the person with the medical notes for future reference. The PDPA allows for that. However, a shopping mall cannot collect the photographs of NRICs of all the shoppers that want to participate in their lucky draw. It is unnecessary to collect the photographs to verify the lucky draw participant. Instead, the participants can be identified with their mobile number, or be asked to give the last 4 characters of the NRIC (i.e. partial NRIC) for verification purposes. This reduces the security risks if the data collected is unintentionally revealed.
NB: Note that PDPA does not apply to public agencies and organisations acting on behalf of them, thus, for example the police can collect your personal information, including NRIC/FIN. Data collected by public agencies are protected by other acts. See https://www.mci.gov.sg/pressroom/news-and-stories/pressroom/2019/2/mcis-response-to-pq-on-public-agencies-exemption-from-pdpa for more information.
### Exercise
- When can a handphone company ask for your NRIC?
- What should the company do to ensure that your data is protected?
```
#YOUR_ANSWER_HERE
```
### 13.2.3 Data Obligations
Organisations are required to abide by the following 9 main personal data obilgations:
1. **Consent Obligation**<br>
Only collect, use or disclose personal data for purposes for which an individual has given his or her consent.
2. **Purpose Limitation Obligation**<br>
An organisation may collect, use or disclose personal data about an individual for the purposes that a reasonable person would consider appropriate in the circumstances and for which the individual has given consent.
3. **Notification Obligation**<br>
Notify individuals of the purposes for which your organisation is intending to collect, use or disclose their personal data on or before such collection, use or disclosure of personal data.
4. **Access and Correction Obligation**<br>
Upon request, the personal data of an individual and information about the ways in which his or her personal data has been or may have been used or disclosed within a year before the request should be provided. However, organisations are prohibited from providing an individual access if the provision of the personal data or other information could reasonably be expected to cause harmful effects. Organisations are also required to correct any error or omission in an individual’s personal data that is raised by the individual.
5. **Accuracy Obligation**<br>
Make reasonable effort to ensure that personal data collected by or on behalf of your organisation is accurate and complete, if it is likely to be used to make a decision that affects the individual, or if it is likely to be disclosed to another organisation.
6. **Protection Obligation**<br>
Make reasonable security arrangements to protect the personal data that your organisation possesses or controls to prevent unauthorised access, collection, use, disclosure or similar risks.
7. **Retention Limitation Obligation**<br>
Cease retention of personal data or remove the means by which the personal data can be associated with particular individuals when it is no longer necessary for any business or legal purpose.
8. **Transfer Limitation Obligation**<br>
Transfer personal data to another country only according to the requirements prescribed under the regulations, to ensure that the standard of protection provided to the personal data so transferred will be comparable to the protection under the PDPA, unless exempted by the PDPC.
9. **Accountability Obligation**<br>
Make information about your data protection policies, practices and complaints process available on request. Designate a Data Protection Officer to ensure that your organisation complies with the PDPA.
More information on PDPA are available at the PDPC website: http://www.pdpc.gov.sg/
### 13.2.4 Protecting your Personal Data
You can take various measures to protect your personal data.
- Don’t reveal your personal data to unknown sources. For phone calls, ensure that the caller is who he or she is before giving your personal information. For websites and applications, read the privacy or data protection policies of the website to understand how your data is used. Websites/applications require you to explicitly agree with the terms when you submit the data. An example is shown below.
<center>
<img src="/images/data-consent.png" width="300" align="center"/>
</center>
Please note that once you agreed, the company can contact you, even if your number is listed in the Do Not Call Registry. If you have queries on personal data, or to withdraw consent, you can contact the data protection officer (DPO). Under PDPA, companies are required to appoint one or more persons to be DPO to oversee the data protection responsibilities within the organisation and ensure compliance with the PDPA.
- Also be careful when throwing away paper containing your personal data such as application forms or letters from schools/banks. Tear or shred the paper so that people cannot use the paper to obtain personal data about yourself.
### Exercise
Read the following case and answer the questions that follow.
>
>A pre-school organised a school trip for interested pre-school students and their parents. To verify that only authorised parents turned up for the school trip, the pre-school teacher collected the parent’s personal data (like identity card numbers).<br>
>A few days before the school trip, the teacher sent a photograph of the consolidated name list to the parent’s WhatsApp chat group to remind those who signed up about the school trip. The photograph contained a table which included the names of the students, with the contact number and identity card numbers of the parents attending.
>
- (a) Was the Personal Data Protection Act (PDPA) breached? If yes, in what ways? If no, why?
- (b) Give examples of the personal data obtained by the teacher.
- (c) What precautions can the teacher take to prevent a similar incident from happening?
```
#YOUR_CODE_HERE
```
# 13.3 Social, Ethical, Legal and Economic Issues
UNDER CONSTRUCTION
| github_jupyter |
The following example illustrates the effect of scaling the regularization parameter when using Support Vector Machines for classification. For SVC classification, we are interested in a risk minimization for the equation:

where
* is used to set the amount of regularization
* is a loss function of our samples and our model parameters.
* is a penalty function of our model parameters
If we consider the loss function to be the individual error per sample, then the data-fit term, or the sum of the error for each sample, will increase as we add more samples. The penalization term, however, will not increase.
When using, for example, cross validation, to set the amount of regularization with C, there will be a different amount of samples between the main problem and the smaller problems within the folds of the cross validation.
Since our loss function is dependent on the amount of samples, the latter will influence the selected value of C. The question that arises is How do we optimally adjust C to account for the different amount of training samples?
The figures below are used to illustrate the effect of scaling our C to compensate for the change in the number of samples, in the case of using an l1 penalty, as well as the l2 penalty.
### l1-penalty case
In the l1 case, theory says that prediction consistency (i.e. that under given hypothesis, the estimator learned predicts as well as a model knowing the true distribution) is not possible because of the bias of the l1. It does say, however, that model consistency, in terms of finding the right set of non-zero parameters as well as their signs, can be achieved by scaling C1.
### l2-penalty case
The theory says that in order to achieve prediction consistency, the penalty parameter should be kept constant as the number of samples grow.
### Simulations
The two figures below plot the values of C on the x-axis and the corresponding cross-validation scores on the y-axis, for several different fractions of a generated data-set.
In the l1 penalty case, the cross-validation-error correlates best with the test-error, when scaling our C with the number of samples, n, which can be seen in the first figure.
For the l2 penalty case, the best result comes from the case where C is not scaled.
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
### Version
```
import sklearn
sklearn.__version__
```
### Imports
This tutorial imports [LinearSVC](http://scikit-learn.org/stable/modules/generated/sklearn.svm.LinearSVC.html#sklearn.svm.LinearSVC), [ShuffleSplit](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.ShuffleSplit.html#sklearn.model_selection.ShuffleSplit), [GridSearchCV](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html#sklearn.model_selection.GridSearchCV) and [check_random_state](http://scikit-learn.org/stable/modules/generated/sklearn.utils.check_random_state.html#sklearn.utils.check_random_state).
```
import plotly.plotly as py
import plotly.graph_objs as go
from plotly import tools
import numpy as np
from sklearn.svm import LinearSVC
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import GridSearchCV
from sklearn.utils import check_random_state
from sklearn import datasets
```
### Calculations
```
rnd = check_random_state(1)
# set up dataset
n_samples = 100
n_features = 300
```
l1 data (only 5 informative features)
```
X_1, y_1 = datasets.make_classification(n_samples=n_samples,
n_features=n_features, n_informative=5,
random_state=1)
```
l2 data: non sparse, but less features
```
y_2 = np.sign(.5 - rnd.rand(n_samples))
X_2 = rnd.randn(n_samples, n_features / 5) + y_2[:, np.newaxis]
X_2 += 5 * rnd.randn(n_samples, n_features / 5)
clf_sets = [(LinearSVC(penalty='l1', loss='squared_hinge', dual=False,
tol=1e-3),
np.logspace(-2.3, -1.3, 10), X_1, y_1),
(LinearSVC(penalty='l2', loss='squared_hinge', dual=True,
tol=1e-4),
np.logspace(-4.5, -2, 10), X_2, y_2)]
colors = ['navy', 'cyan', 'darkorange']
lw = 2
data = []
titles= []
```
### Plot Results
```
for fignum, (clf, cs, X, y) in enumerate(clf_sets):
# set up the plot for each regressor
data.append([[],[]])
for k, train_size in enumerate(np.linspace(0.3, 0.7, 3)[::-1]):
param_grid = dict(C=cs)
# To get nice curve, we need a large number of iterations to
# reduce the variance
grid = GridSearchCV(clf, refit=False, param_grid=param_grid,
cv=ShuffleSplit(train_size=train_size,
n_splits=250, random_state=1))
grid.fit(X, y)
scores = grid.cv_results_['mean_test_score']
scales = [(1, 'No scaling'),
((n_samples * train_size), '1/n_samples'),
]
for subplotnum, (scaler, name) in enumerate(scales):
grid_cs = cs * float(scaler) # scale the C's
trace = go.Scatter(x=grid_cs, y=scores,
name="fraction %.2f" %
train_size,
mode='lines',
line=dict(color=colors[k], width=lw))
data[fignum][subplotnum].append(trace)
titles.append('scaling=%s, penalty=%s, loss=%s' %
(name, clf.penalty, clf.loss))
```
### Plot l1-penalty
```
fig1 = tools.make_subplots(rows=2, cols=1,
subplot_titles=tuple(titles[ :2]))
for i in range(0, len(data[0][0])):
fig1.append_trace(data[0][0][i], 1, 1)
for i in range(0, len(data[0][1])):
fig1.append_trace(data[0][1][i], 2, 1)
for i in map(str, range(1, 3)):
y = 'yaxis' + i
x = 'xaxis' + i
fig1['layout'][y].update(title='CV Score')
fig1['layout'][x].update(title='C', type='log')
fig1['layout'].update(height=700)
py.iplot(fig1)
```
### Plot l2-penalty
```
fig2 = tools.make_subplots(rows=2, cols=1,
subplot_titles=tuple(titles[6 : 8]))
for i in range(0, len(data[1][0])):
fig2.append_trace(data[1][0][i], 1, 1)
for i in range(0, len(data[1][1])):
fig2.append_trace(data[1][1][i], 2, 1)
for i in map(str, range(1, 3)):
y = 'yaxis' + i
x = 'xaxis' + i
fig2['layout'][y].update(title='CV Score')
fig2['layout'][x].update(title='C', type='log')
fig2['layout'].update(height=700)
py.iplot(fig2)
```
### License
Author:
Andreas Mueller <amueller@ais.uni-bonn.de>
Jaques Grobler <jaques.grobler@inria.fr>
License:
BSD 3 clause
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'Scaling the Regularization Parameter for SVCs.ipynb', 'scikit-learn/plot-svm-scale-c/', 'Scaling the Regularization Parameter for SVCs | plotly',
' ',
title = 'Scaling the Regularization Parameter for SVCs | plotly',
name = 'Scaling the Regularization Parameter for SVCs',
has_thumbnail='true', thumbnail='thumbnail/scale.jpg',
language='scikit-learn', page_type='example_index',
display_as='vector_machines', order=12,
ipynb= '~Diksha_Gabha/3590')
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import torch
import torchvision
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from matplotlib import pyplot as plt
%matplotlib inline
mu1 = np.array([3,3,3,3,0])
sigma1 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu2 = np.array([4,4,4,4,0])
sigma2 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu3 = np.array([10,5,5,10,0])
sigma3 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu4 = np.array([-10,-10,-10,-10,0])
sigma4 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu5 = np.array([-21,4,4,-21,0])
sigma5 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu6 = np.array([-10,18,18,-10,0])
sigma6 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu7 = np.array([4,20,4,20,0])
sigma7 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu8 = np.array([4,-20,-20,4,0])
sigma8 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu9 = np.array([20,20,20,20,0])
sigma9 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
mu10 = np.array([20,-10,-10,20,0])
sigma10 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
sample1 = np.random.multivariate_normal(mean=mu1,cov= sigma1,size=500)
sample2 = np.random.multivariate_normal(mean=mu2,cov= sigma2,size=500)
sample3 = np.random.multivariate_normal(mean=mu3,cov= sigma3,size=500)
sample4 = np.random.multivariate_normal(mean=mu4,cov= sigma4,size=500)
sample5 = np.random.multivariate_normal(mean=mu5,cov= sigma5,size=500)
sample6 = np.random.multivariate_normal(mean=mu6,cov= sigma6,size=500)
sample7 = np.random.multivariate_normal(mean=mu7,cov= sigma7,size=500)
sample8 = np.random.multivariate_normal(mean=mu8,cov= sigma8,size=500)
sample9 = np.random.multivariate_normal(mean=mu9,cov= sigma9,size=500)
sample10 = np.random.multivariate_normal(mean=mu10,cov= sigma10,size=500)
# mu1 = np.array([3,3,0,0,0])
# sigma1 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu2 = np.array([4,4,0,0,0])
# sigma2 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu3 = np.array([10,5,0,0,0])
# sigma3 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu4 = np.array([-10,-10,0,0,0])
# sigma4 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu5 = np.array([-21,4,0,0,0])
# sigma5 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu6 = np.array([-10,18,0,0,0])
# sigma6 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu7 = np.array([4,20,0,0,0])
# sigma7 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu8 = np.array([4,-20,0,0,0])
# sigma8 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu9 = np.array([20,20,0,0,0])
# sigma9 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu10 = np.array([20,-10,0,0,0])
# sigma10 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# sample1 = np.random.multivariate_normal(mean=mu1,cov= sigma1,size=500)
# sample2 = np.random.multivariate_normal(mean=mu2,cov= sigma2,size=500)
# sample3 = np.random.multivariate_normal(mean=mu3,cov= sigma3,size=500)
# sample4 = np.random.multivariate_normal(mean=mu4,cov= sigma4,size=500)
# sample5 = np.random.multivariate_normal(mean=mu5,cov= sigma5,size=500)
# sample6 = np.random.multivariate_normal(mean=mu6,cov= sigma6,size=500)
# sample7 = np.random.multivariate_normal(mean=mu7,cov= sigma7,size=500)
# sample8 = np.random.multivariate_normal(mean=mu8,cov= sigma8,size=500)
# sample9 = np.random.multivariate_normal(mean=mu9,cov= sigma9,size=500)
# sample10 = np.random.multivariate_normal(mean=mu10,cov= sigma10,size=500)
X = np.concatenate((sample1,sample2,sample3,sample4,sample5,sample6,sample7,sample8,sample9,sample10),axis=0)
Y = np.concatenate((np.zeros((500,1)),np.ones((500,1)),2*np.ones((500,1)),3*np.ones((500,1)),4*np.ones((500,1)),
5*np.ones((500,1)),6*np.ones((500,1)),7*np.ones((500,1)),8*np.ones((500,1)),9*np.ones((500,1))),axis=0).astype(int)
print(X.shape,Y.shape)
# plt.scatter(sample1[:,0],sample1[:,1],label="class_0")
# plt.scatter(sample2[:,0],sample2[:,1],label="class_1")
# plt.scatter(sample3[:,0],sample3[:,1],label="class_2")
# plt.scatter(sample4[:,0],sample4[:,1],label="class_3")
# plt.scatter(sample5[:,0],sample5[:,1],label="class_4")
# plt.scatter(sample6[:,0],sample6[:,1],label="class_5")
# plt.scatter(sample7[:,0],sample7[:,1],label="class_6")
# plt.scatter(sample8[:,0],sample8[:,1],label="class_7")
# plt.scatter(sample9[:,0],sample9[:,1],label="class_8")
# plt.scatter(sample10[:,0],sample10[:,1],label="class_9")
# plt.legend(bbox_to_anchor=(1.05, 1), loc='upper left')
class SyntheticDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, x, y):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.x = x
self.y = y
#self.fore_idx = fore_idx
def __len__(self):
return len(self.y)
def __getitem__(self, idx):
return self.x[idx] , self.y[idx] #, self.fore_idx[idx]
trainset = SyntheticDataset(X,Y)
# testset = torchvision.datasets.MNIST(root='./data', train=False, download=True, transform=transform)
classes = ('zero','one','two','three','four','five','six','seven','eight','nine')
foreground_classes = {'zero','one','two'}
fg_used = '012'
fg1, fg2, fg3 = 0,1,2
all_classes = {'zero','one','two','three','four','five','six','seven','eight','nine'}
background_classes = all_classes - foreground_classes
background_classes
trainloader = torch.utils.data.DataLoader(trainset, batch_size=100, shuffle=True)
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=100
for i in range(50):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]])
j+=1
else:
image_list.append(foreground_data[fg_idx])
label = foreground_label[fg_idx] - fg1 # minus fg1 because our fore ground classes are fg1,fg2,fg3 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 3000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
list_set_labels = []
for i in range(desired_num):
set_idx = set()
np.random.seed(i)
bg_idx = np.random.randint(0,3500,8)
set_idx = set(background_label[bg_idx].tolist())
fg_idx = np.random.randint(0,1500)
set_idx.add(foreground_label[fg_idx].item())
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
list_set_labels.append(set_idx)
def create_avg_image_from_mosaic_dataset(mosaic_dataset,labels,foreground_index,dataset_number):
"""
mosaic_dataset : mosaic_dataset contains 9 images 32 x 32 each as 1 data point
labels : mosaic_dataset labels
foreground_index : contains list of indexes where foreground image is present so that using this we can take weighted average
dataset_number : will help us to tell what ratio of foreground image to be taken. for eg: if it is "j" then fg_image_ratio = j/9 , bg_image_ratio = (9-j)/8*9
"""
avg_image_dataset = []
for i in range(len(mosaic_dataset)):
img = torch.zeros([5], dtype=torch.float64)
for j in range(9):
if j == foreground_index[i]:
img = img + mosaic_dataset[i][j]*dataset_number/9
else :
img = img + mosaic_dataset[i][j]*(9-dataset_number)/(8*9)
avg_image_dataset.append(img)
return torch.stack(avg_image_dataset) , torch.stack(labels) , foreground_index
def calculate_loss(dataloader,model,criter):
model.eval()
r_loss = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = model(inputs)
loss = criter(outputs, labels)
r_loss += loss.item()
return r_loss/i
class MosaicDataset1(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list, mosaic_label,fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] , self.fore_idx[idx]
batch = 250
msd = MosaicDataset1(mosaic_list_of_images, mosaic_label, fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
```
**Focus Net**
```
class Focus_deep(nn.Module):
'''
deep focus network averaged at zeroth layer
input : elemental data
'''
def __init__(self,inputs,output,K,d):
super(Focus_deep,self).__init__()
self.inputs = inputs
self.output = output
self.K = K
self.d = d
self.linear1 = nn.Linear(self.inputs,100) #,self.output)
self.linear2 = nn.Linear(100,self.output)
def forward(self,z):
batch = z.shape[0]
x = torch.zeros([batch,self.K],dtype=torch.float64)
y = torch.zeros([batch,self.d], dtype=torch.float64)
x,y = x.to("cuda"),y.to("cuda")
for i in range(self.K):
x[:,i] = self.helper(z[:,i] )[:,0] # self.d*i:self.d*i+self.d
x = F.softmax(x,dim=1) # alphas
x1 = x[:,0]
for i in range(self.K):
x1 = x[:,i]
y = y+torch.mul(x1[:,None],z[:,i]) # self.d*i:self.d*i+self.d
return y , x
def helper(self,x):
x = F.relu(self.linear1(x))
x = self.linear2(x)
return x
```
**Classification Net**
```
class Classification_deep(nn.Module):
'''
input : elemental data
deep classification module data averaged at zeroth layer
'''
def __init__(self,inputs,output):
super(Classification_deep,self).__init__()
self.inputs = inputs
self.output = output
self.linear1 = nn.Linear(self.inputs,300)
self.linear2 = nn.Linear(300,self.output)
def forward(self,x):
x = F.relu(self.linear1(x))
x = self.linear2(x)
return x
```
```
where = Focus_deep(5,1,9,5).double()
what = Classification_deep(5,3).double()
where = where.to("cuda")
what = what.to("cuda")
def calculate_attn_loss(dataloader,what,where,criter):
what.eval()
where.eval()
r_loss = 0
alphas = []
lbls = []
pred = []
fidices = []
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels,fidx = data
lbls.append(labels)
fidices.append(fidx)
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
avg,alpha = where(inputs)
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
alphas.append(alpha.cpu().numpy())
mx,_ = torch.max(alpha,1)
entropy = np.mean(-np.log2(mx.cpu().detach().numpy()))
# print("entropy of batch", entropy)
loss = criter(outputs, labels) + entropy
r_loss += loss.item()
alphas = np.concatenate(alphas,axis=0)
pred = np.concatenate(pred,axis=0)
lbls = np.concatenate(lbls,axis=0)
fidices = np.concatenate(fidices,axis=0)
#print(alphas.shape,pred.shape,lbls.shape,fidices.shape)
analysis = analyse_data(alphas,lbls,pred,fidices)
return r_loss/i,analysis
def analyse_data(alphas,lbls,predicted,f_idx):
'''
analysis data is created here
'''
batch = len(predicted)
amth,alth,ftpt,ffpt,ftpf,ffpf = 0,0,0,0,0,0
for j in range (batch):
focus = np.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
amth +=1
else:
alth +=1
if(focus == f_idx[j] and predicted[j] == lbls[j]):
ftpt += 1
elif(focus != f_idx[j] and predicted[j] == lbls[j]):
ffpt +=1
elif(focus == f_idx[j] and predicted[j] != lbls[j]):
ftpf +=1
elif(focus != f_idx[j] and predicted[j] != lbls[j]):
ffpf +=1
#print(sum(predicted==lbls),ftpt+ffpt)
return [ftpt,ffpt,ftpf,ffpf,amth,alth]
print("--"*40)
criterion = nn.CrossEntropyLoss()
optimizer_where = optim.Adam(where.parameters(),lr =0.001)
optimizer_what = optim.Adam(what.parameters(), lr=0.001)
acti = []
loss_curi = []
analysis_data = []
epochs = 1000
running_loss,anlys_data = calculate_attn_loss(train_loader,what,where,criterion)
loss_curi.append(running_loss)
analysis_data.append(anlys_data)
print('epoch: [%d ] loss: %.3f' %(0,running_loss))
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
what.train()
where.train()
for i, data in enumerate(train_loader, 0):
# get the inputs
inputs, labels,_ = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_where.zero_grad()
optimizer_what.zero_grad()
# forward + backward + optimize
avg, alpha = where(inputs)
outputs = what(avg)
mx,_ = torch.max(alpha,1)
entropy = np.mean(-np.log2(mx.cpu().detach().numpy()))
# print("entropy of batch", entropy)
loss = criterion(outputs, labels) + entropy
# loss = criterion(outputs, labels)
# print statistics
running_loss += loss.item()
loss.backward()
optimizer_where.step()
optimizer_what.step()
running_loss,anls_data = calculate_attn_loss(train_loader,what,where,criterion)
analysis_data.append(anls_data)
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
loss_curi.append(running_loss) #loss per epoch
if running_loss<=0.06:
break
print('Finished Training')
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
images, labels,_ = data
images = images.double()
images, labels = images.to("cuda"), labels.to("cuda")
avg, alpha = where(images)
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 3000 train images: %d %%' % ( 100 * correct / total))
analysis_data = np.array(analysis_data)
plt.figure(figsize=(6,6))
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,0],label="ftpt")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,1],label="ffpt")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,2],label="ftpf")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,3],label="ffpf")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.savefig("trends_synthetic_300_300.png",bbox_inches="tight")
plt.savefig("trends_synthetic_300_300.pdf",bbox_inches="tight")
analysis_data[-1,:2]/3000
running_loss,anls_data = calculate_attn_loss(train_loader,what,where,criterion)
print(running_loss, anls_data)
what.eval()
where.eval()
alphas = []
max_alpha =[]
alpha_ftpt=[]
alpha_ffpt=[]
alpha_ftpf=[]
alpha_ffpf=[]
argmax_more_than_half=0
argmax_less_than_half=0
cnt =0
with torch.no_grad():
for i, data in enumerate(train_loader, 0):
inputs, labels, fidx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
avg, alphas = where(inputs)
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
batch = len(predicted)
mx,_ = torch.max(alphas,1)
max_alpha.append(mx.cpu().detach().numpy())
for j in range (batch):
cnt+=1
focus = torch.argmax(alphas[j]).item()
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if (focus == fidx[j].item() and predicted[j].item() == labels[j].item()):
alpha_ftpt.append(alphas[j][focus].item())
# print(focus, fore_idx[j].item(), predicted[j].item() , labels[j].item() )
elif (focus != fidx[j].item() and predicted[j].item() == labels[j].item()):
alpha_ffpt.append(alphas[j][focus].item())
elif (focus == fidx[j].item() and predicted[j].item() != labels[j].item()):
alpha_ftpf.append(alphas[j][focus].item())
elif (focus != fidx[j].item() and predicted[j].item() != labels[j].item()):
alpha_ffpf.append(alphas[j][focus].item())
np.mean(-np.log2(mx.cpu().detach().numpy()))
a = np.array([0.8,0.9])
-np.log2(a)
np.mean(-np.log2(a))
max_alpha = np.concatenate(max_alpha,axis=0)
print(max_alpha.shape, cnt)
np.array(alpha_ftpt).size, np.array(alpha_ffpt).size, np.array(alpha_ftpf).size, np.array(alpha_ffpf).size
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(max_alpha,bins=50,color ="c")
plt.title("alpha values histogram")
plt.savefig("attention_model_2_hist")
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(np.array(alpha_ftpt),bins=50,color ="c")
plt.title("alpha values in ftpt")
plt.savefig("attention_model_2_hist")
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(np.array(alpha_ffpt),bins=50,color ="c")
plt.title("alpha values in ffpt")
plt.savefig("attention_model_2_hist")
```
| github_jupyter |
# Handwritten Chinese and Japanese OCR
In this tutorial, we perform optical character recognition (OCR) for handwritten Chinese (simplified) and Japanese. An OCR tutorial using the Latin alphabet is available in [notebook 208](../208-optical-character-recognition/208-optical-character-recognition.ipynb). This model is capable of processing only one line of symbols at a time.
The models used in this notebook are [handwritten-japanese-recognition](https://docs.openvino.ai/latest/omz_models_model_handwritten_japanese_recognition_0001.html) and [handwritten-simplified-chinese](https://docs.openvino.ai/latest/omz_models_model_handwritten_simplified_chinese_recognition_0001.html). To decode model outputs as readable text [kondate_nakayosi](https://github.com/openvinotoolkit/open_model_zoo/blob/master/data/dataset_classes/kondate_nakayosi.txt) and [scut_ept](https://github.com/openvinotoolkit/open_model_zoo/blob/master/data/dataset_classes/scut_ept.txt) charlists are used. Both models are available on [Open Model Zoo](https://github.com/openvinotoolkit/open_model_zoo/).
## Imports
```
from collections import namedtuple
from itertools import groupby
from pathlib import Path
import cv2
import matplotlib.pyplot as plt
import numpy as np
from openvino.runtime import Core
```
## Settings
Set up all constants and folders used in this notebook
```
# Directories where data will be placed
model_folder = "model"
data_folder = "data"
charlist_folder = f"{data_folder}/charlists"
# Precision used by model
precision = "FP16"
```
To group files, you have to define the collection. In this case, you can use `namedtuple`.
```
Language = namedtuple(
typename="Language", field_names=["model_name", "charlist_name", "demo_image_name"]
)
chinese_files = Language(
model_name="handwritten-simplified-chinese-recognition-0001",
charlist_name="chinese_charlist.txt",
demo_image_name="handwritten_chinese_test.jpg",
)
japanese_files = Language(
model_name="handwritten-japanese-recognition-0001",
charlist_name="japanese_charlist.txt",
demo_image_name="handwritten_japanese_test.png",
)
```
## Select Language
Depending on your choice you will need to change a line of code in the cell below.
If you want to do Japanese OCR, this line should be set to ```language = 'japanese'``` for Chinese set ```language = 'chinese'```.
```
# Select language by using either language='chinese' or language='japanese'
language = "chinese"
languages = {"chinese": chinese_files, "japanese": japanese_files}
selected_language = languages.get(language)
```
## Download Model
In addition to images and charlists, we need to download the model file. In the sections below there are cells for downloading either the Chinese or Japanese model.
If it is your first time running the notebook, the model will download. It may take a few minutes.
We use `omz_downloader`, which is a command-line tool from the `openvino-dev` package. `omz_downloader` automatically creates a directory structure and downloads the selected model.
```
path_to_model_weights = Path(f'{model_folder}/intel/{selected_language.model_name}/{precision}/{selected_language.model_name}.bin')
if not path_to_model_weights.is_file():
download_command = f'omz_downloader --name {selected_language.model_name} --output_dir {model_folder} --precision {precision}'
print(download_command)
! $download_command
```
## Load Network and Execute
When all files are downloaded and language is selected, you need to read and compile the network to run inference. The path to the model is defined based on the selected language.
```
ie = Core()
path_to_model = path_to_model_weights.with_suffix(".xml")
model = ie.read_model(model=path_to_model)
```
### Select Device Name
You may choose to run the network on multiple devices by default it will load the model on the CPU (you can choose manually CPU, GPU etc.) or let the engine choose the best available device (AUTO).
To list all available devices that you can use, uncomment and run the line ```print(ie.available_devices)```.
```
# To check available device names run the line below
# print(ie.available_devices)
compiled_model = ie.compile_model(model=model, device_name="CPU")
```
## Fetch Information About Input and Output Layers
Now that the model is loaded, you need to fetch information about input and output layers. This is information about the input shape and the output.
```
recognition_output_layer = compiled_model.output(0)
recognition_input_layer = compiled_model.input(0)
```
## Load an Image
The next step is to load an image.
The model expects a single-channel image as input, which is why we read the image in grayscale.
After loading the input image, the next step is getting information that you will use for calculating the scale ratio. This describes the ratio between required input layer height and the current image height. In the cell below, the image will be resized and padded to keep letters proportional and meet input shape.
```
# Read file name of demo file based on the selected model
file_name = selected_language.demo_image_name
# Text detection models expects an image in grayscale format
# IMPORTANT!!! This model allows to read only one line at time
# Read image
image = cv2.imread(filename=f"{data_folder}/{file_name}", flags=cv2.IMREAD_GRAYSCALE)
# Fetch shape
image_height, _ = image.shape
# B,C,H,W = batch size, number of channels, height, width
_, _, H, W = recognition_input_layer.shape
# Calculate scale ratio between input shape height and image height to resize image
scale_ratio = H / image_height
# Resize image to expected input sizes
resized_image = cv2.resize(
image, None, fx=scale_ratio, fy=scale_ratio, interpolation=cv2.INTER_AREA
)
# Pad image to match input size, without changing aspect ratio
resized_image = np.pad(
resized_image, ((0, 0), (0, W - resized_image.shape[1])), mode="edge"
)
# Reshape to network the input shape
input_image = resized_image[None, None, :, :]
```
## Visualise Input Image
After preprocessing you can display the image.
```
plt.figure(figsize=(20, 1))
plt.axis("off")
plt.imshow(resized_image, cmap="gray", vmin=0, vmax=255);
```
## Prepare Charlist
The model is loaded and the image is ready. The only element left is the charlist which is downloaded, but before we use it, there is one more step. You must add a blank symbol at the beginning of the charlist. This is expected for both the Chinese and Japanese models.
```
# Get dictionary to encode output, based on model documentation
used_charlist = selected_language.charlist_name
# With both models, there should be blank symbol added at index 0 of each charlist
blank_char = "~"
with open(f"{charlist_folder}/{used_charlist}", "r", encoding="utf-8") as charlist:
letters = blank_char + "".join(line.strip() for line in charlist)
```
## Run Inference
Now run inference. `compiled_model()` takes a list with input(s) in the same order as model input(s). Then we can fetch the output from output tensors.
```
# Run inference on the model
predictions = compiled_model([input_image])[recognition_output_layer]
```
## Process Output Data
The output of model format is W x B x L, where:
* W - output sequence length
* B - batch size
* L - confidence distribution across the supported symbols in Kondate and Nakayosi.
To get a more human-readable format, select a symbol with the highest probability. When you hold a list of indexes that are predicted to have the highest probability, due to limitations in [CTC Decoding](https://towardsdatascience.com/beam-search-decoding-in-ctc-trained-neural-networks-5a889a3d85a7), you will remove concurrent symbols and then remove the blanks.
The last step is getting the symbols from corresponding indexes in the charlist.
```
# Remove batch dimension
predictions = np.squeeze(predictions)
# Run argmax to pick the symbols with the highest probability
predictions_indexes = np.argmax(predictions, axis=1)
# Use groupby to remove concurrent letters, as required by CTC greedy decoding
output_text_indexes = list(groupby(predictions_indexes))
# Remove grouper objects
output_text_indexes, _ = np.transpose(output_text_indexes, (1, 0))
# Remove blank symbols
output_text_indexes = output_text_indexes[output_text_indexes != 0]
# Assign letters to indexes from output array
output_text = [letters[letter_index] for letter_index in output_text_indexes]
```
## Print Output
Now you have a list of letters predicted by the model. The only thing left to do is display the image with predicted text printed below.
```
plt.figure(figsize=(20, 1))
plt.axis("off")
plt.imshow(resized_image, cmap="gray", vmin=0, vmax=255)
print("".join(output_text))
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import math
import sklearn
from sklearn.cross_validation import cross_val_score
from subprocess import check_output
from sklearn.metrics import make_scorer, mean_squared_error
from sklearn.cross_validation import train_test_split
from sklearn.preprocessing import normalize
from sklearn.linear_model import SGDRegressor
from sklearn.preprocessing import OneHotEncoder
enc = OneHotEncoder()
def rmsle_func(actual, predicted):
return np.sqrt(msle(actual, predicted))
def msle(actual, predicted):
return np.mean(sle(actual, predicted))
def sle(actual, predicted):
return (np.power(np.log(np.array(actual)+1) -
np.log(np.array(predicted)+1), 2))
dtypes = {'Semana' : 'int32',
'Agencia_ID' :'int32',
'Canal_ID' : 'int32',
'Ruta_SAK' : 'int32',
'Cliente-ID' : 'int32',
'Producto_ID':'int32',
'Venta_hoy':'float32',
'Venta_uni_hoy': 'int32',
'Dev_uni_proxima':'int32',
'Dev_proxima':'float32',
'Demanda_uni_equil':'int32'}
model = SGDRegressor(loss='squared_loss', penalty='l2', alpha=0.0001,
fit_intercept=True, n_iter=10, shuffle=True, verbose=0,
epsilon=0.1, learning_rate='invscaling',
eta0=0.01, power_t=0.25, warm_start=True, average=False)
from sklearn.feature_extraction import FeatureHasher
h = FeatureHasher(n_features=8000, input_type = 'string')
# Cliente_ID: # of unique = 880604 - многовато, дропаем
df_train = pd.read_csv('train.csv', dtype = dtypes, usecols=["Semana", "Agencia_ID", "Canal_ID", 'Ruta_SAK',
'Producto_ID','Demanda_uni_equil'], chunksize=900000)
i = 1
num = 30000
def logg (x):
return math.log(x+1)
#pd.concat([train, pd.get_dummies(train['Semana'],sparse=True)], axis=1, join_axes=[train.index])
for chunk in df_train:
if i < num :
X_chunk = h.fit_transform(chunk[["Semana", "Agencia_ID", "Canal_ID", 'Ruta_SAK', 'Producto_ID']].astype('string').as_matrix())
y_chunk = np.log(np.ravel(chunk[['Demanda_uni_equil']].as_matrix()) +1)
model.partial_fit(X_chunk, y_chunk)
i = i + 1
elif i == num:
X_chunk = h.fit_transform(chunk[["Semana", "Agencia_ID", "Canal_ID", 'Ruta_SAK','Producto_ID']].astype('string').values)
y_chunk = np.log(np.ravel(chunk[['Demanda_uni_equil']].values) + 1)
#print 'rmsle: ', rmsle_func(y_chunk, model.predict(X_chunk))
print 'rmsle ', math.sqrt(sklearn.metrics.mean_squared_error(y_chunk, model.predict(X_chunk)))
i = i + 1
else:
break
print 'Finished the fitting'
# Now make predictions with trained model
X_test = pd.read_csv('test.csv',dtype = dtypes,usecols=['id', "Semana", "Agencia_ID", "Canal_ID", 'Ruta_SAK',
'Producto_ID'])
ids = X_test['id']
X_test.drop(['id'], axis =1, inplace = True)
y_predicted = np.exp(model.predict(h.fit_transform(X_test.astype('string').values)))-1
submission = pd.DataFrame({"id":ids, "Demanda_uni_equil": y_predicted})
def nonnegative(x):
if (x > 0) or (x == 0):
return x
else:
return 3.9
y_predicted = map(nonnegative, y_predicted)
submission = pd.DataFrame({"id":ids, "Demanda_uni_equil": y_predicted})
cols = ['id',"Demanda_uni_equil"]
submission = submission[cols]
submission.to_csv("submission.csv", index=False)
print('Completed!')
submission = pd.DataFrame({"id":ids, "Demanda_uni_equil": y_predicted})
y_predicted = map(nonnegative, y_predicted)
submission = pd.DataFrame({"id":ids, "Demanda_uni_equil": y_predicted})
cols = ['id',"Demanda_uni_equil"]
submission = submission[cols]
submission.to_csv("submission.csv", index=False)
print('Completed!')
k = submission.Demanda_uni_equil.values
i
```
| github_jupyter |
# Introduction
IPython, pandas and matplotlib have a number of useful options you can use to make it easier to view and format your data. This notebook collects a bunch of them in one place. I hope this will be a useful reference.
The original blog posting is on http://pbpython.com/ipython-pandas-display-tips.html
## Import modules and some sample data
First, do our standard pandas, numpy and matplotlib imports as well as configure inline displays of plots.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
```
One of the simple things we can do is override the default CSS to customize our DataFrame output.
This specific example is from - [Brandon Rhodes' talk at pycon](https://www.youtube.com/watch?v=5JnMutdy6Fw "Pandas From The Ground Up")
For the purposes of the notebook, I'm defining CSS as a variable but you could easily read in from a file as well.
```
CSS = """
body {
margin: 0;
font-family: Helvetica;
}
table.dataframe {
border-collapse: collapse;
border: none;
}
table.dataframe tr {
border: none;
}
table.dataframe td, table.dataframe th {
margin: 0;
border: 1px solid white;
padding-left: 0.25em;
padding-right: 0.25em;
}
table.dataframe th:not(:empty) {
background-color: #fec;
text-align: left;
font-weight: normal;
}
table.dataframe tr:nth-child(2) th:empty {
border-left: none;
border-right: 1px dashed #888;
}
table.dataframe td {
border: 2px solid #ccf;
background-color: #f4f4ff;
}
"""
```
Now add this CSS into the current notebook's HTML.
```
from IPython.core.display import HTML
HTML('<style>{}</style>'.format(CSS))
SALES=pd.read_csv("../data/sample-sales-tax.csv", parse_dates=True)
SALES.head()
```
You can see how the CSS is now applied to the DataFrame and how you could easily modify it to customize it to your liking.
Jupyter notebooks do a good job of automatically displaying information but sometimes you want to force data to display. Fortunately, ipython provides and option. This is especially useful if you want to display multiple dataframes.
```
from IPython.display import display
display(SALES.head(2))
display(SALES.tail(2))
display(SALES.describe())
```
## Using pandas settings to control output
Pandas has many different options to control how data is displayed.
You can use max_rows to control how many rows are displayed
```
pd.set_option("display.max_rows",4)
SALES
```
Depending on the data set, you may only want to display a smaller number of columns.
```
pd.set_option("display.max_columns",6)
SALES
```
You can control how many decimal points of precision to display
```
pd.set_option('precision',2)
SALES
pd.set_option('precision',7)
SALES
```
You can also format floating point numbers using float_format
```
pd.set_option('float_format', '{:.2f}'.format)
SALES
```
This does apply to all the data. In our example, applying dollar signs to everything would not be correct for this example.
```
pd.set_option('float_format', '${:.2f}'.format)
SALES
```
## Third Party Plugins
Qtopian has a useful plugin called qgrid - https://github.com/quantopian/qgrid
Import it and install it.
```
import qgrid
qgrid.nbinstall(overwrite=True)
```
Showing the data is straighforward.
```
qgrid.show_grid(SALES, remote_js=True)
```
The plugin is very similar to the capability of an Excel autofilter. It can be handy to quickly filter and sort your data.
## Improving your plots
I have mentioned before how the default pandas plots don't look so great. Fortunately, there are style sheets in matplotlib which go a long way towards improving the visualization of your data.
Here is a simple plot with the default values.
```
SALES.groupby('name')['quantity'].sum().plot(kind="bar")
```
We can use some of the matplolib styles available to us to make this look better.
http://matplotlib.org/users/style_sheets.html
```
plt.style.use('ggplot')
SALES.groupby('name')['quantity'].sum().plot(kind="bar")
```
You can see all the styles available
```
plt.style.available
plt.style.use('bmh')
SALES.groupby('name')['quantity'].sum().plot(kind="bar")
plt.style.use('fivethirtyeight')
SALES.groupby('name')['quantity'].sum().plot(kind="bar")
```
Each of the different styles have subtle (and not so subtle) changes. Fortunately it is easy to experiment with them and your own plots.
You can find other articles at [Practical Business Python](http://pbpython.com)
This notebook is referenced in the following post - http://pbpython.com/ipython-pandas-display-tips.html
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Filter/filter_eq.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Filter/filter_eq.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Filter/filter_eq.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Filter/filter_eq.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The following script checks if the geehydro package has been installed. If not, it will install geehydro, which automatically install its dependencies, including earthengine-api and folium.
```
import subprocess
try:
import geehydro
except ImportError:
print('geehydro package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geehydro'])
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once.
```
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
states = ee.FeatureCollection('TIGER/2018/States')
selected = states.filter(ee.Filter.eq("NAME", 'California'))
Map.centerObject(selected, 6)
Map.addLayer(ee.Image().paint(selected, 0, 2), {'palette': 'yellow'}, 'Selected')
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
### Data Loaders Pytorch Class
```
import torch
import torchvision
from torch.utils.data import DataLoader, Dataset
import numpy as np
import pandas as pd
```
> **Point** - gradient computation is not efficient for the whole dataset. So the data must be splited in `so` called `batches`
> **Training Loop** -with the data splitted into batches
```
for epoch in range(epochs):
## loop over all batches
for batch in range(total_batches):
batch_x, batch_y = ...
```
#### Terms
1. **epoch** - one forward pass to **ALL** trainning sample.
2. **Batch_size** - number of trainning sample used in one backward/forward pass
3. **number_of_iterations** = number of passes each epoch pass (forward + backward) ``100 samples, batch_size =20: iterations = 100/20 =5 for each epoch``
### The DataLoader
> The dataloader can do the `batch_size` and other computation for us.
#### Creation a custom Dataset.
There are 2 steps to do this:
* inherit the Dataset Class
* implement the following
* `__init__`
* `__getitem__`
* `__len__`
```
class Wine(Dataset):
def __init__(self):
# Xy = np.loadtxt('wine.csv', delimiter=',', dtype='float32', skiprows=1)
# print(Xy[0])
xy = pd.read_csv('wine.csv').values
self.n_samples = xy.shape[0]
self.X = torch.from_numpy(xy[:, 1:].astype('float32')) # all the columns except the first one
self.y = torch.from_numpy(xy[:, 0:1].astype('float32')) # the first column
# print(self.y[:3])
# To allow indexing such as dataset[i]
def __getitem__(self, index):
return self.X[index], self.y[index]
# when we call len(dataset)
def __len__(self):
return self.n_samples
wine = Wine()
wine
len(wine), wine[:2]
```
### Load Wine dataset using the DataLoader class
```
train = DataLoader(dataset=wine,
batch_size=4,
shuffle=True,
)
```
### Loading data from `torchvision.datasets`
```
train_dataset = torchvision.datasets.MNIST(
'',
train = True,
transform=torchvision.transforms.ToTensor(),
download=True
)
test_dataset = torchvision.datasets.MNIST(
'',
train = False,
transform=torchvision.transforms.ToTensor(),
download=True
)
```
### Creating Loaders
```
train_loader = DataLoader(dataset=train_dataset,
batch_size = 10,
shuffle = True
)
test_loader = DataLoader(dataset=test_dataset,
batch_size = 10,
shuffle = True
)
```
### Iterating over datasets
```
for data in test_loader:
X, y = data
print(X, y)
break
```
> Do tehe training magic with the data.
| github_jupyter |
```
import pandas as pd
import numpy as np
from scipy.stats import norm
# 입력으로 주어지는 데이터는 모두 numpy array라고 가정하고 구현
# 입력 특성의 개수에 관계없이 동작하도록 구현
class MyGaussianNB:
def __init__(self):
self.distXy0 = []
self.distXy1 = []
self.prior0 = 0
self.prior1 = 0
self.m0 = 0 # 훈련 데이터에서 레이블이 0인 샘플 개수
self.m1 = 0 # 훈련 데이터에서 레이블이 0인 샘플 개수
self.num_col = 0 # 특성벡터의 차원
def fit(self, X_train, y_train):
self.num_col = X_train.shape[1]
X_train0 = X_train[y_train == 0] ###################여기를 채우세요
X_train1 = X_train[y_train == 1] ###################여기를 채우세요
self.m0 = (y_train == 0).sum() ###################여기를 채우세요
self.m1 = (y_train == 1).sum() ###################여기를 채우세요
# self.fit_dist() method는 코드 맨 끝에 정의되어 있음
for i in range(self.num_col):
self.distXy0.append(self.fit_dist(X_train0,i)) # X_train0의 column index i열에 대한 정규분포를 distXy0 리스트 i번째에 추가
self.distXy1.append(self.fit_dist(X_train1,i)) # X_train1의 column index i열에 대한 정규분포를 distXy1 리스트 i번째에 추가
self.prior0 = self.m0/(self.m0 + self.m1) ###################여기를 채우세요
self.prior1 = self.m1/(self.m0 + self.m1) ###################여기를 채우세요
def predict_proba(self, X):
prob0 = self.prior0
prob1 = self.prior1
for i in range(self.num_col):
prob0 *= self.distXy0[i].pdf(X[:,i]) ###################여기를 채우세요
prob1 *= self.distXy1[i].pdf(X[:,i]) ###################여기를 채우세요
result = np.array([prob0, prob1])/ (prob0+prob1)
return result.T
def predict(self, X):
return np.argmax(self.predict_proba(X), axis=1)
def score(self, X, y):
y_hat = self.predict(X)
return (y == y_hat).sum()/len(y) ###################여기를 채우세요
def fit_dist(self,data, col_idx):
mu = data[:,col_idx].mean()
sigma = data[:,col_idx].std(ddof=0)
dist = norm(mu, sigma)
return dist
data = pd.read_csv('./dataset/PimaIndiansDiabetes.csv')
data_subset = data.loc[1:,['Blood Glucose', 'BMI', 'Class']]
data_subset[data_subset.isna() == True].sum()
bg_mask = data_subset.loc[:,"Blood Glucose"]!=0
bmi_mask = data_subset.loc[:,"BMI"]!=0
clean_data_subset = data_subset[bg_mask & bmi_mask]
X = clean_data_subset.loc[:, ['Blood Glucose', 'BMI']]
y = clean_data_subset.loc[:, 'Class']
ratio = 0.8
total_num = len(clean_data_subset)
train_num = int(ratio*total_num)
np.random.seed(42)
shuffled_idx = np.arange(total_num)
np.random.shuffle(shuffled_idx)
X_train = X.iloc[shuffled_idx[:train_num]]
y_train = y.iloc[shuffled_idx[:train_num]]
X_test = X.iloc[shuffled_idx[train_num:]]
y_test = y.iloc[shuffled_idx[train_num:]]
# 준비된 데이터가 pd.DataFrame일 때, 모두 to_numpy() 메소드를 이용하여 numpy array로 바꾼 뒤에 사용
ClassifierNB = MyGaussianNB()
ClassifierNB.fit(X_train.to_numpy(), y_train.to_numpy())
ClassifierNB.predict(X_test.to_numpy())[:22]
ClassifierNB.predict_proba(X_test.to_numpy())[:5]
ClassifierNB.score(X_train.to_numpy(), y_train.to_numpy())
ClassifierNB.score(X_test.to_numpy(), y_test.to_numpy())
```
| github_jupyter |
```
import spiceypy as spice
import pvl
import os
import re
import subprocess
from ale import util
from itertools import chain
import io
import networkx as nx
# These should be provided when running this script.
cube = "/home/acpaquette/B10_013341_1010_XN_79S172W.cub"
output_dir = "/Users/jmapel/ale/nb_test" # Output dir for created kernel files
data_dir = "/usgs/cpkgs/isis3/data/" # Dir of where to pull original kernels from
ckslicer_loc = "/Users/jmapel/ale/ckslicer"
def merge_intervals(intervals):
"""
Merge a set of intervals. The intervals are assumed to be closed, that is they include the end-points.
Parameters
----------
intervals : list
The input list of intrvals where each interval is a tuple of (start, end)
Returns
-------
: list
A sorted list of the merged intervals
"""
sorted_intervals = sorted(intervals, key=lambda tup: tup[0])
merged = [sorted_intervals[0]]
for interval in sorted_intervals[1:]:
# No intersection
if interval[0] > merged[-1][1]:
merged.append(interval)
# Intersection, but new interval isn't wholey contained
elif interval[1] > merged[-1][1]:
merged[-1] = (merged[-1][0], interval[1])
return merged
def add_light_time_correction(cube_info, padding=5):
"""
Compute the time intervals for the image and any light time correction
Parameters
----------
cube_info : ordered dict
The cube info from ale.util.generate_kernels_from_cube
padding : float
Time padding in seconds to add to each interval
Returns
-------
: list
A sorted list of the intervals as (start_et, stop_et)
"""
image_start_et = spice.scs2e(cube_info['SpacecraftID'], cube_info['SpacecraftClockCount'])
image_end_et = image_start_et + cube_info['ExposureDuration'] * cube_info['Lines']
inst_state, inst_lt = spice.spkez(cube_info['SpacecraftID'], image_start_et, 'J2000', 'LT+S', 0)
target_state, target_lt = spice.spkez(cube_info['TargetID'], image_start_et, 'J2000', 'LT+S', 0)
sun_state, sun_lt = spice.spkez(10, image_start_et, 'J2000', 'LT+S', cube_info['TargetID'])
intervals = [
(image_start_et - padding, image_end_et + padding),
(image_start_et - padding - inst_lt, image_end_et + padding - inst_lt),
(image_start_et - padding - target_lt, image_end_et + padding - target_lt),
(image_start_et - padding - sun_lt, image_end_et + padding - sun_lt)]
return merge_intervals(intervals)
# These are the processing steps. This will make use of the cube provided further up to create smaller,
# more manageable kernel files for ale testing purposes. This currently only handles ck and spk files.
# Get dictionary of kernel lists from cube
cube_info = util.generate_kernels_from_cube(cube, format_as = 'dict')
# Replace path variables with absolute paths for kernels
for kernel_list in cube_info:
for index, kern in enumerate(cube_info[kernel_list]):
if kern is not None:
cube_info[kernel_list][index] = data_dir + kern.strip('$')
# Create ordered list of kernels for furnishing
kernels = [kernel for kernel in chain.from_iterable(cube_info.values()) if isinstance(kernel, str)]
spice.furnsh(kernels)
# Loads cube as pvl to extract rest of data
cube_pvl = pvl.load(cube)
# Save other necesary info in cube_info dict
cube_info.update(Lines = cube_pvl['IsisCube']['Core']['Dimensions']['Lines'])
cube_info.update(Lines = 400)
cube_info.update(SpacecraftClockCount = cube_pvl['IsisCube']['Instrument']['SpacecraftClockCount'])
cube_info.update(ExposureDuration = cube_pvl['IsisCube']['Instrument']['LineExposureDuration'].value * 0.001)
cube_info.update(TargetID = spice.bods2c(cube_pvl['IsisCube']['Instrument']['TargetName']))
cube_info.update(SpacecraftID = spice.bods2c(cube_pvl['IsisCube']['Instrument']['SpacecraftName']))
# Account for light time correction
intervals = add_light_time_correction(cube_info)
# For each binary ck kernel specified in cube, run the ckslicer, comment and to-transfer commands
for ck in [k for k in kernels if k.lower().endswith('.bc')]:
ck_path, ck_file_extension = os.path.splitext(ck)
ck_basename = os.path.basename(ck_path)
for index, interval in enumerate(intervals):
for frame in util.get_ck_frames(ck):
output_basename = os.path.join(output_dir, ck_basename + '_' + str(index) + '_sliced_' + str(frame))
output_kern = output_basename + ck_file_extension
output_comments = output_basename + '.cmt'
start_sclk = spice.sce2s(cube_info['SpacecraftID'], interval[0])
end_sclk = spice.sce2s(cube_info['SpacecraftID'], interval[1])
# Create new sliced ck kernel
ckslicer_command = [ckslicer_loc,
'-LSK {}'.format(cube_info['LeapSecond'][0]),
'-SCLK {}'.format(cube_info['SpacecraftClock'][0]),
'-INPUTCK {}'.format(ck),
'-OUTPUTCK {}'.format(output_kern),
'-ID {}'.format(frame),
'-TIMETYPE {}'.format('SCLK'),
'-START {}'.format(start_sclk),
'-STOP {}'.format(end_sclk)]
subprocess.run(ckslicer_command, check=True)
# Remove old comments from new ck kernel
commnt_command = ['commnt', '-d {}'.format(output_kern)]
subprocess.run(commnt_command, check=True)
with open(output_comments, 'w+') as comment_file:
comment_file.write("This CK is for testing with the image: {}\n".format(cube))
comment_file.write("\nThis CK was generated using the following command: {}\n")
comment_file.write(" ".join(ckslicer_command))
# Add new comments to new ck kernel
new_commnts_command = ["commnt", "-a {}".format(output_kern), output_comments]
subprocess.run(new_commnts_command, check=True)
# Create the transfer file of the new ck kernel
subprocess.run(["toxfr", output_kern], check=True)
# Create the config file for the spkmerge command
for index, interval in enumerate(intervals):
output_spk_basename = os.path.join(output_dir, os.path.basename(os.path.splitext(cube)[0]) + '_' + str(index))
output_spk = output_spk_basename + '.bsp'
start_utc = spice.et2utc(interval[0], 'c', 3)
end_utc = spice.et2utc(interval[1], 'c', 3)
spk_dep_tree = util.create_spk_dependency_tree([k for k in kernels if k.lower().endswith('.bsp')])
config_string = util.spkmerge_config_string(spk_dep_tree,
output_spk,
[cube_info['TargetID'], cube_info['SpacecraftID'], 10],
cube_info['LeapSecond'][0],
start_utc,
end_utc)
with open(output_spk_basename + '.conf', 'w+') as spk_config:
spk_config.write(config_string)
# Create the new SPK
spkmerge_command = ["spkmerge", spk_config.name]
subprocess.run(spkmerge_command, check=True)
# Create the transfer file of the new SPK kernel
subprocess.run(["toxfr", output_spk], check=True)
```
| github_jupyter |
```
import os
import tensorflow as tf
from keras import backend as K
os.environ["CUDA_VISIBLE_DEVICES"] = '3'
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
K.set_session(sess)
from keras.applications import Xception
from keras.layers import Dense, Flatten, GlobalAveragePooling2D, Activation
from keras.models import Model, load_model
from keras.preprocessing.image import ImageDataGenerator
from keras.optimizers import Adam
from keras.callbacks import EarlyStopping, ReduceLROnPlateau
from sklearn import metrics
nb_classes = 1 # number of classes
img_width, img_height = 224, 224 # change based on the shape/structure of your images
batch_size = 32 # try 4, 8, 16, 32, 64, 128, 256 dependent on CPU/GPU memory capacity (powers of 2 values).
nb_epoch = 50 # number of iteration the algorithm gets trained.
learn_rate = 1e-5 # sgd learning rate
train_dir = '/home/skkulab/ICCV/Dataset/preprocessed_dataset/train'
validation_dir = '/home/skkulab/ICCV/Dataset/preprocessed_dataset/validation'
test_dir = '/home/skkulab/ICCV/Dataset/preprocessed_dataset/test'
```
### Xception Model
```
base_model = Xception(weights='imagenet', include_top=False, input_shape=(img_height, img_width, 3))
# Add fully connected layer
x = base_model.output
x = GlobalAveragePooling2D()(x)
x = Dense(nb_classes, activation=None)(x)
x = Activation('sigmoid')(x)
model = Model(base_model.input, x)
print(model.summary())
print(len(model.trainable_weights))
```
### Preparing train, validation data
```
train_datagen = ImageDataGenerator(rescale=1./255)
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(train_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
shuffle=True,
class_mode='binary')
validation_generator = test_datagen.flow_from_directory(validation_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
shuffle=False,
class_mode='binary')
test_generator = test_datagen.flow_from_directory(test_dir,
target_size=(img_height, img_width),
batch_size=32,
shuffle=False,
class_mode='binary')
test_classes = test_generator.classes
len(test_classes[test_classes == 0])
```
### Train model(weight unfreezed)
```
for layer in base_model.layers:
layer.trainable = True
model.compile(optimizer=Adam(lr=learn_rate),
loss='binary_crossentropy',
metrics=['accuracy'])
print(len(model.trainable_weights))
callback_list = [EarlyStopping(monitor='val_acc', patience=5),
ReduceLROnPlateau(monitor='val_loss', factor=0.1, patience=3)]
history = model.fit_generator(train_generator,
steps_per_epoch=len(train_generator),
epochs=nb_epoch,
validation_data=validation_generator,
validation_steps=len(validation_generator),
callbacks=callback_list,
verbose=1)
model.save('/home/skkulab/ICCV/models/xception_v3.h5')
```
### Evaluate test data
```
test_loss, test_acc = model.evaluate_generator(test_generator, steps=len(test_generator))
print('test acc:', test_acc)
print('test_loss:', test_loss)
predictions = model.predict_generator(test_generator, steps=len(test_generator))
predictions[predictions > 0.5] = 1
predictions[predictions <= 0.5] = 0
true_classes = test_generator.classes
report = metrics.classification_report(true_classes,predictions)
print(report)
train_classes = train_generator.classes
validation_predictions = model.predict_generator(validation_generator, steps=len(validation_generator))
validation_predictions[validation_predictions > 0.5] = 0
validation_predictions[validation_predictions <= 0.5] = 1
validation_predictions
true_classes = validation_generator.classes
true_classes
report = metrics.classification_report(true_classes,validation_predictions)
print(report)
```
### Draw plot
```
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
plt.savefig('/home/skkulab/ICCV/models//xcetion_v1.png')
```
### Model train(weight unfreezed)
```
for layer in base_model.layers:
layer.trainable = True
model.compile(optimizer=Adam(lr=learn_rate),
loss='binary_crossentropy',
metrics=['accuracy'])
print(len(model.trainable_weights))
history = model.fit_generator(train_generator,
steps_per_epoch=len(train_generator),
epochs=nb_epoch,
validation_data=validation_generator,
validation_steps=len(validation_generator),
verbose=1)
model.save('/home/skkulab/ICCV/models/xception_v2.h5')
```
### Draw plot
```
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
```
### Load trained model
```
loaded_model = load_model('/home/skkulab/ICCV/models/xception_v1.h5')
loaded_model.summary()
```
### Preparing test data
```
test_generator = test_datagen.flow_from_directory(test_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
shuffle=False,
class_mode='binary')
test_loss, test_acc = loaded_model.evaluate_generator(test_generator, steps=len(test_generator))
print('test acc:', test_acc)
print('test_loss:', test_loss)
predictions = loaded_
```
| github_jupyter |
# What is the % odorant vapor in the air stream, as a function of odorant properties, jar dimensions, and air flow rate?
A cynlindrical jar with cross-sectional area $A$ and height $h$ contains a volume $V_{sol}$ of odorant solution (diluted in a solvent with zero vapor pressure), and the remaining volume, above the solution, is called $V_{head}$.
The flow rate of air into the jar, in mol/s, is $r_{air,in}$. The gas flow out of the jar consists of the air component $r_{air, out}$ and the odorant component $r_{odor, out}$, also in mol/s.
The interface between the solution and the jar headspace has odorant molecules leaving with rate $r_{evap}$ and molecules condensing at rate $r_{condense}$, also in mol/s.
Therefore, the amount of odorant $n_o$ (in moles) in the jar headspace evolves according to:
\begin{equation}
\tag{1}\frac{dn_{odor}}{dt} = r_{evap} - r_{condense} - r_{odor, out}
\end{equation}
And the amount of air $n_{air}$ in the jar headspace evolves according to:
\begin{equation}
\tag{2}\frac{dn_{air}}{dt} = r_{air,in} - r_{air, out}
\end{equation}
Assuming that mixing with odorant vapor in the jar is instantaneous. Then the ratio of odorant efflux out of the jar to air efflux out of the jar is equal to the ratio of odor to air in the headspace of the jar at that moment.
\begin{equation}
\tag{3}\frac{n_{odor}}{n_{air}} = \frac{r_{odor,out}}{r_{air, out}}
\end{equation}
If evaporation is slow compared to the flow rate, we can assume that:
\begin{equation}
\tag{4}r_{air,in} = r_{air,out} + r_{odor, out}
\end{equation}
Combining equations 3 and 4 gives:
$$r_{odor,out} = r_{air,out}\frac{n_{odor}}{n_{air}} = (r_{air,in} - r_{odor, out})\frac{n_{odor}}{n_{air}}$$
$$r_{odor,out}(1+\frac{n_{odor}}{n_{air}}) = r_{air,in}\frac{n_{odor}}{n_{air}}$$
\begin{equation}
\tag{5}r_{odor,out} = \frac{r_{air,in}n_{odor}}{n_{air}+n_{odor}}
\end{equation}
And lastly, if both odorant and air are ideal gases, and air flow does not substantially change pressure in the jar, then the combined air and gas molecules fill the headspace at room temperature and ambient pressure according to:
\begin{equation}
\tag{6}n_{odor} + n_{air} = \frac{P_{room}V_{head}}{RT_{room}} \sim \frac{V_{head}}{22.4 Liters}
\end{equation}
Substituting (6) into (5) gives:
\begin{equation}
\tag{7}r_{odor, out} = \frac{r_{air,in}n_{odor}RT_{room}}{P_{room}V_{head}}
\end{equation}
The evaporation rate is assumed to be indepedent of time, depending (through a function $F_{evap}$ obtained empirically and described later) only upon the partial pressure $P^*_o$ of the liquid odorant, and the surface area $A$ of the liquid-vapor interface, where the partial pressure is equal to the mole fraction $f_o$ of the odorant in the liquid times its intrinsic vapor pressure $P_o$ (at this temperature), according to Raoult's law.
\begin{equation}
\tag{8}r_{evap} = F_{evap}(P^*_{odor})A
\end{equation}
\begin{equation}
\tag{9}P^*_{odor} = P_{odor}f_{odor}
\end{equation}
Meanwhile, the condensation rate depends upon the molar concentration C_o of odorant vapor in the headspace and the headspace volume according to a constant $k_{condense}$:
\begin{equation}
\tag{10}r_{condense} = k_{condense}AC_{odor}
\end{equation}
\begin{equation}
\tag{11}C_{odor} = n_{odor}/V_{head}
\end{equation}
Substituting (7-11) back into equation (1) gives a first order, linear differential equation in $n_{odor}$:
$$\frac{dn_{odor}}{dt} = F_{evap}(P_{odor},f_{odor})A - \frac{k_{condense}An_{odor}}{V_{head}} - \frac{r_{air,in}n_{odor}RT_{room}}{P_{room}V_{head}}$$
\begin{equation}
\tag{12}\frac{dn_{odor}}{dt} = F_{evap}(P_{odor},f_{odor})A - n_{odor}(\frac{1}{V_{head}}(k_{condense}A + \frac{r_{air,in}RT_{room}}{P_{room}}))
\end{equation}
We can define:
\begin{equation}
\tag{13}u = \frac{1}{V_{head}}(k_{condense}A + \frac{r_{air,in}RT_{room}}{P_{room}})
\end{equation}
and if the headspace volume changes very slowly, we can assume that this is independent of time.
The integrating factor is:
\begin{equation}
e^{\int{u}dt} = e^{ut}\tag{14}
\end{equation}
The canonical solution to (12) is thus:
\begin{equation}
\tag{15}n_{odor}(t) = e^{-ut}(\int{e^{ut}F_{evap}(P_{odor},f_{odor})Adt} + constant)
\end{equation}
Integrating gives:
\begin{equation}
n_{odor}(t) = \frac{F_{evap}(P_{odor},f_{odor})A}{u} + constant*e^{-ut}\tag{16}
\end{equation}
At steady state flow ($t = \infty$), the number of odorant molecules in the headspace is proportional to the evaporation function $F_{evap}$, which depends on the vapor pressure of the odorant, its mole fraction in solution, and the surface area of the liquid-vapor interface). The number of odorant molecules in the headspace is inversely proportional to $u$, which includes a weighted sum of the condensation rate constant and the air inflow rate.
\begin{equation}
\tag{17}
n_{odor}(t=\infty) = \frac{F_{evap}(P_{odor},f_{odor})A}{u}
\end{equation}
At the starting time ($t=0$), we will assume that the jar is already in equilibrium, and the number of odorant molecules in the headspace is determined by the odorant's partial pressure:
\begin{equation}
\tag{18}
n_{odor}(t=0) = \frac{P_{odor}f_{odor}V_{head}}{RT_{room}} = \frac{F_{evap}(P_{odor},f_{odor})A}{u} + constant
\end{equation}
So the constant is equal to:
\begin{equation}
\tag{19}
constant = \frac{P_{odor}f_{odor}V_{head}}{RT_{room}} - \frac{F_{evap}(P_{odor},f_{odor})A}{u}
\end{equation}
Substituting back into (16) gives:
\begin{equation}
n_{odor}(t) = \frac{F_{evap}(P_{odor},f_{odor})A}{u} + (\frac{P_{odor}f_{odor}V_{head}}{RT_{room}} - \frac{F_{evap}(P_{odor},f_{odor})A}{u})*e^{-ut}\tag{20}
\end{equation}
And substituting (13) into (20) gives:
\begin{equation}
n_{odor}(t) = \frac{V_{head}F_{evap}(P_{odor},f_{odor})A}{(k_{condense}A + \frac{r_{air,in}RT_{room}}{P_{room}})} + (\frac{P_{odor}f_{odor}V_{head}}{RT_{room}} - \frac{V_{head}F_{evap}(P_{odor},f_{odor})A}{(k_{condense}A + \frac{r_{air,in}RT_{room}}{P_{room}})})*e^{-(\frac{1}{V_{head}}(k_{condense}A + \frac{r_{air,in}RT_{room}}{P_{room}}))t}\tag{21}
\end{equation}
Subsituting (21) into (5) gives:
\begin{equation}
\frac{r_{odor,out}}{r_{air,in}} = \frac{\frac{V_{head}F_{evap}(P_{odor},f_{odor})A}{(k_{condense}A + \frac{r_{air,in}RT_{room}}{P_{room}})} + (\frac{P_{odor}f_{odor}V_{head}}{RT_{room}} - \frac{V_{head}F_{evap}(P_{odor},f_{odor})A}{(k_{condense}A + \frac{r_{air,in}RT_{room}}{P_{room}})})*e^{-(\frac{1}{V_{head}}(k_{condense}A + \frac{r_{air,in}RT_{room}}{P_{room}}))t}}{\frac{P_{room}V_{head}}{RT_{room}}}
\end{equation}
\begin{equation}
\tag{22}
\frac{r_{odor,out}}{r_{air,in}} = \frac{F_{evap}(P_{odor},f_{odor})}{(\frac{k_{condense}P_{room}}{RT_{room}} + \frac{r_{air,in}}{A})} + (\frac{P_{odor}f_{odor}}{P_{room}} - \frac{F_{evap}(P_{odor},f_{odor})}{(\frac{k_{condense}P_{room}}{RT_{room}} + \frac{r_{air,in}}{A})})*e^{-(\frac{1}{V_{head}}(k_{condense}A + \frac{r_{air,in}RT_{room}}{P_{room}}))t}
\end{equation}
The evaporation rate $F_{evap}$ depends on $P_{odor}$, $f_{odor}$, and $A$. According to *Mackay, D., & van Wesenbeeck, I. (2014). Correlation of Chemical Evaporation Rate with Vapor Pressure. Environmental Science & Technology, 48(17), 10259–10263.
doi:10.1021/es5029074* the pure odorant evaporation rate per unit area has the empirical form:
\begin{equation}
\tag{23}
F^*_{evap}(P_{odor}) = e^{1.0243 ln(\frac{P_{odor}}{P_{unity}} - 15.08)} * F_{unity}
\end{equation}
where $P_{unity} = 1 Pa$ and $F_{unity}= 1\frac{mol}{m^2s}$
Assuming linear dependence on mole fraction in solution:
\begin{equation}
\tag{25}
F_{evap}(P_{odor},f_{odor}) = F^*_{evap}(P_{odor})f_{odor} = f_{odor}e^{1.0243 ln(\frac{P_{odor}}{P_{unity}} - 15.08)} * F_{unity}
\end{equation}
At equilibrium, the condensation rate is equal to the evaporation rate, resulting in a $n_{odor}(t=0)$ given by the partial pressure $P^*_{odor} = P_{odor}f_{odor}$. Setting (8) and (10) equal and subsituting (18) gives:
\begin{equation}
F_{evap}(P_{odor},f_{odor})A = k_{condense}A\frac{n_{odor}(t=0)}{V_{head}}
\end{equation}
\begin{equation}
\tag{26}
F^*_{evap}(P_{odor})f_{odor}A = k_{condense}A\frac{P_{odor}f_{odor}V_{head}}{V_{head}RT_{room}}
\end{equation}
\begin{equation}
\tag{27}
k_{condense} = \frac{RT_{room}F^*_{evap}(P_{odor})}{P_{odor}}
\end{equation}
Substituting (25) and (26) into (22) gives:
\begin{equation}
\frac{r_{odor,out}}{r_{air,in}}(t) = \frac{F^*_{evap}(P_{odor})f_{odor}}{(\frac{\frac{RT_{room}F^*_{evap}(P_{odor})}{P_{odor}}P_{room}}{RT_{room}} + \frac{r_{air,in}}{A})} + (\frac{P_{odor}f_{odor}}{P_{room}} - \frac{F^*_{evap}(P_{odor})f_{odor}}{(\frac{\frac{RT_{room}F^*_{evap}(P_{odor})}{P_{odor}}P_{room}}{RT_{room}} + \frac{r_{air,in}}{A})})e^{-(\frac{1}{V_{head}}(\frac{RT_{room}F^*_{evap}(P_{odor})}{P_{odor}}A + \frac{r_{air,in}RT_{room}}{P_{room}}))t}
\end{equation}
\begin{equation}
\frac{r_{odor,out}}{r_{air,in}}(t) = f_{odor}(\frac{F^*_{evap}(P_{odor})}{(\frac{F^*_{evap}(P_{odor})}{P_{odor}}P_{room} + \frac{r_{air,in}}{A})} + (\frac{P_{odor}}{P_{room}} - \frac{F^*_{evap}(P_{odor})}{(\frac{F^*_{evap}(P_{odor})}{P_{odor}}P_{room} + \frac{r_{air,in}}{A})})e^{-(\frac{RT_{room}}{V_{head}}(\frac{F^*_{evap}(P_{odor})}{P_{odor}}A + \frac{r_{air,in}}{P_{room}}))t})
\end{equation}
\begin{equation}
\tag{28}
\frac{r_{odor,out}}{r_{air,in}}(t) = f_{odor}(\frac{1}{(\frac{P_{room}}{P_{odor}} + \frac{r_{air,in}}{AF^*_{evap}(P_{odor})})} + (\frac{P_{odor}}{P_{room}} - \frac{1}{(\frac{P_{room}}{P_{odor}} + \frac{r_{air,in}}{AF^*_{evap}(P_{odor})})})e^{-(\frac{RT_{room}}{V_{head}}(\frac{F^*_{evap}(P_{odor})}{P_{odor}}A + \frac{r_{air,in}}{P_{room}}))t})
\end{equation}
This equation, describing the molar fraction of odorant in the air stream, has the functional form:<br><br>
\begin{equation}
\tag{29}
\frac{r_{odor,out}}{r_{air,in}}(t) = f_{odor}(a + (b-a)e^{-ct})
\end{equation}
<br>where $f_{odor}$ is the mole fraction of the odorant in solution, $f_{odor}a$ describes the steady-state fraction in the exiting vapor at $t=0$, $f_{odor}b$ describes the steady-state fraction in the exiting vapor at $t=\infty$, and $\frac{1}{c}$ is the time constant.
In the vapor pressure regime of most odorants (0.1 - 10 mm Hg), the ratio $\frac{F^*_{evap}(P_{odor})}{P_{odor}}$ does not differ more than $\sim5\%$ from $k_{evap} \sim3.2*10^{-7} \frac{mol}{m^2sPa}$, i.e. $F^*_{evap}$ is linear in $P_{odor}$. This means that the time constant $\frac{1}{c}$ is largely independent of the odorant in question.
Plugging in numbers:
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import quantities as pq
from quantities.constants.statisticalmechanics import R
from IPython.display import Markdown
sns.set(font_scale=1.5)
jar_diameter = 6 * pq.cm
jar_height = 5 * pq.cm
height_filled = 1*pq.cm
A = np.pi * (jar_diameter / 2)**2
V_head = A * (jar_height - height_filled)
f_odor = 0.001 # A 0.1% solution
P_room = 1*pq.atm # 1 atmosphere
P_odor = 10.5 * pq.mmHg # Hexanal vapor pressure
T_room = (22 + 273.15)*pq.Kelvin
r_air_in = (1.0*pq.L/pq.min)*P_room/(R*T_room) # 1 L/min convered to mol/s
def F_star_evap(vp):
# Units of Pascals
vp = vp.rescale(pq.Pa)
# Strip units for logarithm
vp /= vp.units
# Evaporation rate
er = np.exp(1.0243*np.log(vp) - 15.08)
if isinstance(er, np.ndarray):
er[vp==0] = 0
# Attach units
er *= pq.mol / (pq.m**2 * pq.s)
return er
t = np.linspace(0,5,1000) * pq.s # 1000 ms
# Use the form:
# ratio = f_odor*(a + (b - a)exp(-ct))
a = (1/(P_room/P_odor + r_air_in/(A*F_star_evap(P_odor)))).rescale(pq.dimensionless)
b = (P_odor/P_room).rescale(pq.dimensionless)
c = ((R*T_room/V_head)*(A*F_star_evap(P_odor)/P_odor + r_air_in/P_room)).rescale(1/pq.s)
ratio = f_odor*(a + (b - a)*np.exp(-c*t))
Markdown(r"a = %.3g<br>b= %.3g<br>c = %.3g($s^{-1}$)" % (a, b, c))
```
Here is the decay curve for the fraction of odorant (by mole) in the vapor leaving the jar over time:
```
plt.plot(t, ratio)
plt.xlabel('Time (s)')
plt.ylabel('Volume fraction odorant\nin air stream')
plt.ylim(0, ratio.max()*1.1);
vps = (np.logspace(-1,1,100) * pq.mmHg).rescale(pq.Pa)
ers = np.zeros(vps.shape)
for i, vp in enumerate(vps):
ers[i] = F_star_evap(vp)
plt.scatter(vps, ers/vps)
plt.xscale('log')
plt.ylim(3e-7,3.5e-7)
plt.xlabel(r'Vapor pressure ($Pa$)')
plt.ylabel(r'Evaporation rate ($\frac{mol}{m^2s}$)');
r_air_ins_volume = np.logspace(-4,2,100)*pq.L/pq.min
r_air_ins_mole = (r_air_ins_volume*P_room/(R*T_room)).rescale(pq.mol/pq.s)
k_evap = 3.2e-7*pq.mol/(pq.m**2*pq.s*pq.Pa)
cs = ((R*T_room/V_head)*(A*k_evap + r_air_ins_mole/P_room)).rescale(1/pq.s)
plt.scatter(r_air_ins_volume, 1/cs)
plt.xlabel(r'Air flow rate ($\frac{L}{min}$)')
plt.ylabel(r'Odorant depletion time constant ($s$)')
plt.xscale('log')
plt.yscale('log')
plt.xlim(1e-4,1e2)
plt.ylim(1e-2,1e2);
```
Using the linear form of $F^*_{evap}$, the dependence on $P_{odor}$ of the ratio between the initial and steady-state odorant enrichments disappears:
\begin{equation}
\tag{30}
\frac{b}{a} =
\frac{\frac{P_{odor}}{P_{room}}}{\frac{1}{(\frac{P_{room}}{P_{odor}} + \frac{r_{air,in}}{AF^*_{evap}(P_{odor})})}} =
\frac{\frac{P_{odor}}{P_{room}}}{\frac{1}{(\frac{P_{room}}{P_{odor}} + \frac{r_{air,in}}{Ak_{evap}P_{odor}})}} =
\frac{P_{room}+\frac{r_{air,in}}{Ak_{evap}}}{P_{room}} =
1+\frac{r_{air,in}}{Ak_{evap}P_{room}}
\end{equation}
The only way to keep this fraction close to 1 is to have a low air-flow rate $r_{air,in}$ or a large solution-vapor interface surface area $A$, i.e. a wide jar.
```
b_a_ratios = 1 + (r_air_ins_mole/(A*k_evap*P_room)).rescale(pq.dimensionless)
plt.scatter(r_air_ins_volume, b_a_ratios)
plt.xlabel(r'Air flow rate ($\frac{L}{min}$)')
plt.ylabel(r'Odorant enrichment ratio $\frac{initial}{steadystate}$')
plt.xscale('log')
plt.yscale('log')
plt.xlim(1e-4,1e2)
plt.ylim(0.9,1e3);
```
In the interval between stimuli, if air flow through the jar is turned off, the odorant concentration increases again towards its intitial concentration. Here we assume that the pressure change inside the sealed jar due to the odorant realizing its partial pressure is small.<br><br>
\begin{equation}
\tag{31}
\frac{dn_{odor}}{dt} = r_{evap} - r_{condense} = f_{odor}F*_{evap}(P_{odor})A - \frac{RT_{room}F^*_{evap}(P_{odor})An_{odor}}{P_{odor}V_{head}}
\end{equation}
We can define:
\begin{equation}
\tag{32}u^* = \frac{RT_{room}F^*_{evap}(P_{odor})A}{P_{odor}V_{head}}
\end{equation}
with a solution just like (21), except without the term for air influx:<br><br>
\begin{equation}
\tag{32}
n_{odor}(t) = \frac{V_{head}F_{evap}(P_{odor},f_{odor})A}{k_{condense}A} + (\frac{P_{odor}f_{odor}V_{head}}{RT_{room}} - \frac{V_{head}F_{evap}(P_{odor},f_{odor})A}{k_{condense}A})*e^{-(\frac{1}{V_{head}}(k_{condense}A))t}
\end{equation}
\begin{equation}
n_{odor}(t) = e^{-ut}(\int{e^{ut}f_{odor}F^*_{evap}(P_{odor})Adt} + constant)
\end{equation}
\begin{equation}
n_{odor}(t) = \frac{f_{odor}F^*_{evap}(P_{odor})A}{u} + constant*e^{-ut}
\end{equation}
<br>
\begin{equation}
n_{odor}(t) = \frac{f_{odor}F^*_{evap}(P_{odor})A}{\frac{RT_{room}F^*_{evap}(P_{odor})A}{P_{odor}V_{head}}} + constant*e^{-ut}
\end{equation}
<br>
\begin{equation}
\tag{33}n_{odor}(t) = \frac{f_{odor}P_{odor}V_{head}}{RT_{room}} + constant*e^{-\frac{RT_{room}F^*_{evap}(P_{odor})A}{P_{odor}V_{head}}t}
\end{equation}
The value of the constant term depends on how depleted the odorant in the headspace is before the airflow is turned off. Without less of generality we can simply define it as:
\begin{equation}
\tag{34}
constant = n_{odor}(t=0) - \frac{f_{odor}P_{odor}V_{head}}{RT_{room}}
\end{equation}
giving us:
\begin{equation}
\tag{35}n_{odor}(t) = \frac{f_{odor}P_{odor}V_{head}}{RT_{room}} + (n_{odor}(t=0) - \frac{f_{odor}P_{odor}V_{head}}{RT_{room}})*e^{-\frac{RT_{room}F^*_{evap}(P_{odor})A}{P_{odor}V_{head}}t}
\end{equation}
Therefore, a scenario of a stimulus of duration $T_on$ followed by an interval of duration $T_off$ exhibits the following behavior: The quantity of odorant in the jar headspace is initially $\frac{f_{odor}P_{odor}V_{head}}{RT_{room}}$, which follows trivially from Raoult's law and the ideal gas law. Once the air flow is turned on, it declines by a factor of $\frac{b}{a} = 1+\frac{r_{air,in}}{Ak_{evap}P_{room}}$ with inverse time constant $c = \frac{RT_{room}}{V_{head}}(\frac{F^*_{evap}(P_{odor})}{P_{odor}}A + \frac{r_{air,in}}{P_{room}})) \sim \frac{RT_{room}}{V_{head}}(k_{evap}A + \frac{r_{air,in}}{P_{room}}))$. When the air is turned off, it returns to its initial value with inverse time constant $c = \frac{RT_{room}F^*_{evap}(P_{odor})A}{P_{odor}V_{head}} \sim \frac{RT_{room}k_{evap}A}{V_{head}}$.
```
dt = 0.001
T_on = 2
T_off = 8
n_cycles = 10
t = np.arange(0,(T_on+T_off)*n_cycles,dt) * pq.s
t_on = int(T_on / dt)
t_off = int(T_off / dt)
ratios = np.zeros(t.shape)
# Use the form:
# ratio = f_odor*(a + (b - a)exp(-ct))
a = (1/(P_room/P_odor + r_air_in/(A*F_star_evap(P_odor)))).rescale(pq.dimensionless)
b = (P_odor/P_room).rescale(pq.dimensionless)
c_decay = ((R*T_room/V_head)*(A*F_star_evap(P_odor)/P_odor + r_air_in/P_room)).rescale(1/pq.s)
c_recover = ((R*T_room/V_head)*(A*F_star_evap(P_odor)/P_odor)).rescale(1/pq.s)
ratios[0:t_on] = f_odor*(a + (b - a)*np.exp(-c_decay*t[:t_on]))
ratios[t_on:t_on+t_off] = f_odor*(b + (ratios[t_on-1]/f_odor - b)*np.exp(-c_recover*t[:t_off]))
for cycle in range(1,n_cycles):
ratios[cycle*t_on+cycle*t_off:(cycle+1)*t_on+cycle*t_off] = f_odor*(a + (ratios[cycle*t_on+cycle*t_off-1]/f_odor - a)*np.exp(-c_decay*t[:t_on]))
ratios[(cycle+1)*t_on+cycle*t_off:(cycle+1)*t_on+(cycle+1)*t_off] = f_odor*(b + (ratios[(cycle+1)*t_on+cycle*t_off-1]/f_odor - b)*np.exp(-c_recover*t[:t_off]))
```
Repeated stimuli (with on time T_on and off time T_off will have an odorant mole fraction in the vapor that changes as follows):
```
plt.plot(t, ratios)
plt.xlabel('Time ($s$)')
plt.ylabel('Mole fraction of odorant\nin headspace vapor');
plt.ylim(0, ratios[0]*1.1);
print("Depletion time constant is %.3g s" % (1/c_decay))
print("Recovery time constant is %.3g s" % (1/c_recover))
```
| github_jupyter |
# The _fast & parallel_ Virtual Brain
## A fast implementation of The Virtual Brain brain network simulator
* written in C using a host of optimizations that make brain simulation reeeallllyy fast
* parallelized (multithreading)
* containerized (can be conveniently run e.g. through Docker, Shifter or Singularity, without the need to install dependencies or set up environment)
* uses the Deco-Wang (aka "ReducedWongWang") neural mass model to simulate local brain region activity as described in Deco et al., 2014, Journal of Neuroscience or Schirner et al., 2018, eLife
### In this example we show how to start a containerized version of TVB on a supercomputer.
* First, we will upload our custom brain model to Collab Storage
* Then, we will copy the brain model into the virtual filesystem underlying the Python notebook container and from there onto the supercomputer
* Then, we will set up a job for the supercomputer batch system and submit it.
* Finally, we will download simulation results from the supercomputer.
### Note on the parallelization
In this version of TVB a simulation of a single parameter set is split up and distributed over several threads, that are then executed on multiple cores. For example, if a brain network model consists of 400 nodes and one uses 8 threads for simulation, each thread computes the activity of 50 nodes. Since the threads operate on shared memory, there is no additional overhead for distributing data between threads.
### To use this example notebook with your own data copy it to another Collab, adapt parameters below according to your setup, and then run all cells.
### Citation
If you use this code, please cite this publication where it was used first:
**Schirner, Michael, Anthony Randal McIntosh, Viktor Jirsa, Gustavo Deco, and Petra Ritter. "Inferring multi-scale neural mechanisms with brain network modelling." Elife 7 (2018): e28927.**
### Note: the fastTVB Docker Container is also freely available as a standalone and can be directly pulled from https://hub.docker.com/r/thevirtualbrain/fast_tvb and used on local (super)computers.
Michael Schirner, May 2020.
michael.schirner@charite.de
petra.ritter@charite.de
# 1. Prepare your brain model
Brain model data consists of a structural connectome (weights and distance files) and a parameter set (parameter file). Unlike the Python TVB version where brain model input data (i.e. the structural connectome) is stored as a Zip or hdf5 file, for this implementation a brain model simply consists of two ASCII text files and, additionally, one ASCII text files that contains the simulation parameters.
All three files must be stored in a folder with the name `input`. Additionally, we need a folder where outputs are stored called `output`.
__Naming conventions of weights and distance matrix files__
Connectome files must have the suffixes `"_SC_weights.txt"` or `"_SC_distances.txt"`
connection weights file: `<sub_id>_SC_weights.txt`
connection distances file: `<sub_id>_SC_distances.txt`
The subject identifier `<sub_id>` can be any arbitrary short (alphanumeric) string, e.g. `"sub002"`, and must be provided as a parameter to the batch file (which allows to have multiple different brain models stored in the same folder).
Example:
```
sub002_SC_weights.txt
```
and
```
sub002_SC_distances.txt
```
__Formatting conventions of weights and distance matrix files__
ASCII text files that contain the connection weight and distance matrices as floating point numbers separated by white-spaces (columns) and line breaks/newline characters (rows). The unit of distances is _mm_ while connection weights are dimensionless and depend on the global coupling scaling factor parameter that is set in the parameter file.
Example (weights file):
```
0 0.012 0.0 0.119 0.0 1.1234 ...
0.012 0 0 0.0 1.34 ...
...
...
```
__Naming conventions of parameter file__
Arbitrary short string with the suffix `".txt"`. Must be provided as a parameter to the batch file (which allows to have multiple different parameter sets stored in the same folder). The file must have the suffix `".txt"` (actually, the TVB container is able to work with arbitrary parameter file names, but HBP Collab Storage prevents upload of files without suffix).
Example:
```
param_set_042.txt
```
__Formatting conventions of parameter file__
ASCII text file that contains the parameters as floating point or integer numbers separated by white spaces.
The sorting of the parameter file is:
```
nodes, G, J_NMDA, w_plus, Ji, sigma, time_steps, BOLD_TR, global_trans_v, rand_num_seed
```
**Important**: Correct formatting of floating point vs. integer numbers is mandatory. Integer parameters (like "number of nodes") must not be formatted as floating point numbers (don't format the integer "2" with a radix point like this "2.0")! For correct number format please refer to the table below.
Example:
```
379 1.000 0.150 1.400 1.000 0.0100 10000 720 12.5000 1403
```
__Parameter description__
For a detailed description of parameters see Deco et al. (2014) JNeuro or Schirner et al. (2018) eLife
Parameter | Description | Number format
:---: | :---: | :---:
nodes | number of nodes in brain network model | Integer
G | global coupling scaling factor | Float
J_NMDA | strength of excitatory (NMDA) synpases | Float
w_plus | strength of local excitatory recurrence | Float
Ji | strength of local inhibitory (GABA) synapses | Float
sigma | noise strength | Float
time_steps | length of the simulation (ms) | Integer
BOLD_TR | TR of simulated fMRI BOLD signal (ms) | Integer
global_trans_v | transmission velocity of large-scale network (m/s) | Float
rand_num_seed | Seed to initialize random number generator | Integer
__Create input/output folders__
As final step, we create the folders `input` and `output` and copy the two connectome files and the parameter file into the folder `input`. The brain model is now ready to be used if you pull the Docker container into your (super)computer (e.g. `docker pull thevirtualbrain/fast_tvb`). If this is your goal, please refer to usage instructions on the Dockerhub page (https://hub.docker.com/r/thevirtualbrain/fast_tvb). In order to use it through EBRAINS Collabs, you can continue with the steps below.
## 2. Upload your brain model to the EBRAINS Collab
1. Navigate to the [Collabs](https://wiki.ebrains.eu/bin/view/Collabs/) page (https://wiki.ebrains.eu/bin/view/Collabs/)
2. Click on the "Create a Collab" button and fill out the form.
4. Click on "Drive" in the left sidebar menu. You might need to wait for a few seconds and refresh the page before it is visible.
5. Download this Ipython notebook to your local file system (e.g. by clicking on File->Save notebook as...) and upload it into your newly created Collab by using the "Upload" button on the "Drive" page.
6. Click on New to create a folder for your brain network model data (we will call it ```input``` here) and then upload your weights, distances and parameter files into the newly created folder (or drag & drop them into the new folder).
7. Make sure that the three files were successfully uploaded. They should now appear as the contents of the new folder.
In this example we uploaded the files
```
gavg_SC_distances.txt
gavg_SC_weights.txt
param_set.txt
```
Depending on whether you used a private or public repository the files will end up in either of the following folders in the filesystem of the EBRAINS Jupyter Hub at https://lab.ebrains.eu/
```
public_drive = 'drive/Shared with all'
private_drive = 'drive/Shared with groups'
```
## 3. Upload brain model to supercomputer
Move over to EBRAINS Jupyter Hub to adapt this notebook according to your Collab file system.
Build the path to the three files in your Collab like in the following code snippet.
```
# ADJUSTABLE PARAMETERS
################################
# paths for public and private drives
public_drive = 'drive/Shared with all'
private_drive = 'drive/Shared with groups'
which_drive = private_drive # is your Collab public or private?
subject_id = 'gavg' # prefix of weights and distances files
parameter_file = 'param_set.txt' # name of parameter set file
# Collab name
collab = 'TVB C -- High-speed parallel brain network models' # name of Collab
# data folder name
path = 'input' # the folder where you uploaded the three brain model files
################################
weights_file = subject_id + '_SC_weights.txt' # weights file must have this suffix
distances_file = subject_id + '_SC_distances.txt' # distances file must have this suffix
# get path of home folder
import os
home = os.getenv('HOME')
# full path
weights_path = os.path.join(home, which_drive, collab, path, weights_file)
distances_path = os.path.join(home, which_drive, collab, path, distances_file)
params_path = os.path.join(home, which_drive, collab, path, parameter_file)
print(weights_path)
print(distances_path)
print(params_path)
# Check whether the files are really there
print(os.path.exists(weights_path))
print(os.path.exists(distances_path))
print(os.path.exists(params_path))
```
If we end up with "True" three times in a row, we have the correct paths.
Now it's time to upload our brain model to the supercomputer. Therefore, we create a PyUnicore client.
First, we update PyUnicore, if necessary. Then, we import it. Finally, we connect with PizDaint. To see which other supercomputers are there, and to learn their ID run the commented
```
r.site_urls
```
To select a different supercomputer replace the supercomputer identifier string in
```
site_client = r.site('DAINT-CSCS')
```
with your preferred supercomputer.
```
# use the pyunicore library
!pip install pyunicore --upgrade
import pyunicore.client as unicore_client
tr = unicore_client.Transport(clb_oauth.get_token())
r = unicore_client.Registry(tr, unicore_client._HBP_REGISTRY_URL)
site_client = r.site('DAINT-CSCS')
```
Next, we start an "empty" interactive job to get a workspace on PizDaint
```
job_description = {}
job = site_client.new_job(job_description)
storage = job.working_dir
#storage.properties
```
First, let's check the contents of the folder
```
storage.listdir()
```
Good, it's empty. If it's not empty we can remove files or folders with `storage.rm(filename)` or `storage.rmdir(foldername)`. Run `help(storage)` for more information.
Now, let's create our `input` and `output` directories and then copy the three brain model files into `input`.
```
storage.mkdir('input')
storage.mkdir('output')
storage.listdir()
```
Great, the two folders exist. While we are here, let's extract the path of our current working directory (mount point), we need it later when we generate the job script.
```
mp = (storage.properties['mountPoint']).encode('ascii').decode('utf-8')
mp
```
Now, let's copy the three brain model files into input. With the last two lines we check whether the folder `input` contains our three files.
```
storage.upload(input_name=weights_path, destination = "input/" + weights_file)
storage.upload(input_name=distances_path, destination = "input/" + distances_file)
storage.upload(input_name=params_path, destination = "input/" + parameter_file)
r=storage.stat("input")
r.properties
```
We see that the folder "input" has three children -- our three uploaded files. Great, the entire brain model is now copied to the supercomputer. What remains to be done is generating a batch script for SLURM (the job manager; installed on many supercomputers), copying the script into our current working directory and submitting the job to the queue.
## 6. Create SLURM job script for supercomputer
### Specify input data set, output folder and simulation parameters
HBP supercomputers use SLURM to manage job queues. Below we create a SLURM submission script that loads required modules and then posts a simulation job to the queue.
### Configure the following parameters:
* paths of input/output folders
* data set ID
* path of parameter file
* number of parallel threads
* optimum depends on the supercomputer architecture and the size of the brain model
Note: instead of directly starting a job on the batch system, we use PyUnicore to submit a job on the login node, which in turn submits a job for the batch system. This gives us a greater flexibility to configure our job, we don't have to learn so much PyUnicore (although it's great!) and are falisafe if PyUnicore misses bindings for certain job managers. Note that before we run the container, we make sure that the image is up to date, or, if non-existent, gets pulled for the first time.
Supercomputers typically don't have Docker installed due to security reasons, but more secure alternatives like Sarus. Usage is usually very similar, but permissions are more restricted.
Below is a brief outline of the Sarus run command. For a great in-depth tutorial check out the help pages of the Swiss CSCS supercomputing site: https://user.cscs.ch/tools/containers/
```
sarus run <container_name>
```
is the standard way of running a container.
Here, we additionally use the `mount` command to directly mount the `input` and the `output` folders into the container's filesystem's top-level directories `/input` and `/output`.
Example:
```
srun sarus run --mount=type=bind,source=/path/to/output,target=/output --mount=type=bind,source=/path/to/input,target=/input thevirtualbrain/fast_tvb /start_simulation.sh <Arguments>
```
**Important**: the container assumes that the external folders mounted to `/input` and `/output` in its virtual file system contain the input data, respectively, are the place where output data shall be stored. If there is no rw access to these folders container execution will fail!
Output will be stored into the `output` folder following the naming schema
```
<sub_id>_<paramset_file>_fMRI.txt
```
The entrypoint script in the root folder of the container is called `start_simulation.sh` and it needs three arguments: the name of the parameter file, the subject ID and the number of threads.
The command format of the entry-point script is
Usage:
```
/start_simulation.sh <paramset_file> <sub_id> <#threads>
```
Example:
```
/start_simulation.sh param_set42.txt sub0014 8
```
For an in-depth discussion of Sarus (Shifter, Singularity) usage, check out this documentation:
https://user.cscs.ch/tools/containers/
```
# ADJUSTABLE PARAMETERS
################################################
wall_time = "00:10:00" # ADJUST wall time of job
cpu_per_task = "36" # ADJUST according to supercomputer architecture
num_threads = "4" # ADJUST according to size of the model
################################################
# FIXED PARAMETERS
################################################
job_script = "job_script" # name of the job script file
input_folder = mp + "input"
output_folder = mp + "output"
################################################
with open(job_script, "w") as f:
f.write("#!/bin/bash -l\n")
f.write("#SBATCH --time=" + wall_time + "\n")
f.write("#SBATCH --output=slurm-" + job_script + ".out\n")
f.write("#SBATCH --nodes=1\n")
f.write("#SBATCH --ntasks-per-core=1\n")
f.write("#SBATCH --ntasks-per-node=1\n")
f.write("#SBATCH --cpus-per-task=" + cpu_per_task + "\n")
f.write("#SBATCH --partition=normal\n")
f.write("#SBATCH --constraint=mc\n")
f.write("#SBATCH --hint=nomultithread\n") # disable hyperthreading such that all cores become available for multithreading
f.write("export OMP_NUM_THREADS=$SLURM_CPUS_PER_TASK\n")
f.write("module load /apps/daint/UES/easybuild/modulefiles/daint-mc\n")
f.write("module load /apps/daint/system/modulefiles/sarus/1.1.0\n\n")
f.write("srun sarus pull thevirtualbrain/fast_tvb\n")
f.write("srun sarus run --mount=type=bind,source=" + output_folder + \
",target=/output --mount=type=bind,source=" + input_folder + \
",target=/input thevirtualbrain/fast_tvb /start_simulation.sh " + \
parameter_file + " " + subject_id + " " + num_threads + "\n")
```
Let's check the generated script
```
!cat job_script
```
Looking good!
## 7. Upload SLURM script to supercomputer
Here we use the same upload function as used previously for the brain model data followed by a quick check whether the file arrived.
```
storage.upload(input_name=job_script, destination = job_script)
storage.listdir()
```
Nice!
## 8. Launching the simulation on the supercomputer
The big moment has come, we will now finally launch the simulation. We do this by executing the the SLURM command `sbatch`, which will evaluate our batch file and generate a job out of it that is added to the queue.
After we executed the job, we extract the working directory of this job (which is different from the other working directory that we created above, where the folders `input` and `output` are located).
```
rr=site_client.execute("sbatch " + mp + "job_script")
rr.working_dir.properties['mountPoint']
```
Let's peak into the working directory of the job we just started
```
rr.working_dir.listdir()
```
Now that we made sure that the simulation is running smoothly, we finished quite an amount of work. Congratulations so far! I think you earned yourself a short break. Go outside for a few minutes and get some fresh air and light!
I'll wait.
...
...
...
Are you back? Great! There's not much left to do now. What we gonna do is: check whether the simulation runs or ran smoothly, copy the result into our virtual file system of the notebook, plot the result and finally copy it into storage so you can download it.
First, let's peek into the SLURM output file -- we defined its name above when we generated the batch script. We used the job script name to define it.
Most important are the last 19 lines of the output file. Important is the line `MT-TVBii finished. Execution took X s for Y nodes. Goodbye!`, which indicates that the simulation finished successfully. The table below shows you when the job started, when it finished and how long it took. If these lines are missing the job is either still running or something went wrong.
```
job_out = rr.working_dir.stat("slurm-" + job_script + ".out")
job_out.raw().readlines()[-30:]
```
## 9. Collecting the results
In our example everything went smoothly, so let's collect the results and plot them.
```
result_filename = subject_id + "_" + parameter_file + "_fMRI.txt"
storage.stat("output/" + result_filename).download(result_filename)
import numpy as np
fMRI = np.genfromtxt(result_filename)
%matplotlib inline
from matplotlib.pyplot import *
figure(figsize=(15, 10))
plt = plot(fMRI.T, alpha=0.1)
title("simulated fMRI time series")
xlabel('Time (TR)')
ylabel('fMRI Amplitude (1)')
```
Beautiful!
We're done!
Your simulation results should await you under the filename ```result_filename``` in the folder of this notebook. Right-click and "Download" and you can download the data to your local computer.
**If you would like to use fast_TVB on your local (super)computer without the intermediary Collab, just pull it from Dockerhub into your local machine**
https://hub.docker.com/r/thevirtualbrain/fast_tvb
```
docker pull thevirtualbrain/fast_tvb
```
Feedback may be addressed to:
michael.schirner@charite.de
or
petra.ritter@charite.de
Here's the end. Enjoy your day!
| github_jupyter |
# COMP 135 day01: Intro to Numerical Python
A Python-based version of the "Ch 2 lab" from James et al.'s "Introduction to Statistical Learning" textbook
Based on original notebook: https://nbviewer.jupyter.org/github/emredjan/ISL-python/blob/master/labs/lab_02.3_introduction.ipynb
# What to Do
Students should run this notebook locally, interactively modifying cells as needed to understand concepts and have hands-on practice.
Try to make sure you can *predict* what a function will do. Build a mental model of NumPy will make you a better programmer and ML engineer.
Ask questions like:
* what should the result's output type be?
* what should the result's *shape* be?
* what should the result's *values* be?
# Outline
* [Data types](#data_types)
* [Dimension and shape](#dimension_and_shape)
* [Reshaping](#reshaping)
* [Elementwise multiplication](#elementwise_multiplication)
* [Matrix multiplication](#matrix_multiplication)
* [Useful functions](#useful_functions)
-- linspace, logspace
-- arange
-- allclose
* [Reductions](#reductions)
-- min
-- max
-- sum
-- prod
* [Indexing](#indexing)
# Key Takeaways
* Numpy array types (`np.array`) have a DIMENSION, a SHAPE, and a DATA-TYPE (dtype)
* * Know what you are using!
* Consider using standard notation to avoid confusion
* * 1-dim arrays of size N could be named `a_N` or `b_N` instead of `a` or `b`
* * 2-dim arrays of size (M,N) could be named `a_MN` instead of `a`
* * With this notation, it is far more clear that `np.dot(a_MN, b_N)` will work, but `np.dot(a_MN.T, b_N)` will not
* Broadcasting is key
* * See https://numpy.org/doc/stable/user/basics.broadcasting.html
* Always use np.array, avoid np.matrix
* * Why? Array is more flexible (can be 1-dim, 2-dim, 3-dim, 4-dim, and more!)
* * Also, np.matrix will be deprecated soon <https://numpy.org/doc/stable/reference/generated/numpy.matrix.html>
# Further Reading
* Stefan van der Walt, S. Chris Colbert, Gaël Varoquaux. The NumPy array: a structure for efficientnumerical computation. Computing in Science and Engineering, Institute of Electrical and Electronics Engineers, 2011.
<https://hal.inria.fr/inria-00564007/document>
* https://realpython.com/numpy-array-programming/
```
# import numpy (array library)
import numpy as np
```
# Basic array creation and manipulation
We use `np.array(...)` function to create arrays
```
x = np.array([1.0, 6.0, 2.4]);
print(x);
x + 2 # basic element-wise addition
x * 2 # basic element-wise multiplication
x / 2 # basic element-wise division
x + x # can operate on two arrays of SAME size
x / x # element-wise division
```
<a id="data_types"></a>
# Data types
Arrays have *data types* (or "dtypes")
```
y = np.array([1., 4, 3]) # with decimal point in "1.", defaults to 'float' type
print(y)
print(y.dtype)
y_int = np.array([1, 4, 3]) # without decimal point, defaults to 'int' type
print(y_int)
print(y_int.dtype)
y_float32 = np.array([1, 4, 3], dtype=np.float32) # use optional keyword argument (aka 'kwarg') to specify data type
print(y_float32)
print(y_float32.dtype)
z = y + y_float32 # What happens when you add float32 to float64? *upcast* to highest precision
print(z)
print(z.dtype)
```
<a id="dimension_and_shape"></a>
# Dimension and Shape
Arrays have DIMENSION and SHAPE
Dimension = an integer value : number of integers needed to index a unique entry of the array
Shape = a tuple of integers : each entry gives the size of the corresponding dimension
```
y.ndim
y.shape
# Create 2D 3x3 array 'M' as floats
M = np.asarray([[1, 4, 7.0], [2, 5, 8], [3, 6, 9]])
print(M)
print(M.ndim)
print(M.shape)
# Create 2D *rectangular* array
M_35 = np.asarray([[1, 4, 7.0, 10, 13], [2, 5, 8, 11, 14], [3, 6, 9, 12, 15]])
print(M_35)
```
<a id="reshaping"></a>
# Reshaping
Sometimes, we want to transforming from 1-dim to 2-dim arrays
We can either use:
* the *reshape* function
* indexing with the "np.newaxis" built-in <https://numpy.org/doc/stable/reference/constants.html#numpy.newaxis>
#### Demo of reshape
```
y = np.array([1.0, 4, 3])
print(y)
print(y.shape)
y_13 = np.reshape(y, (1,3)) # use '_AB' suffix to denote an array with shape (A, B)
print(y_13)
print(y_13.shape)
y_31 = np.reshape(y, (3, 1)) # use '_AB' suffix to denote an array with shape (A, B)
print(y_31)
print(y_31.shape)
```
#### Demo of newaxis
```
y = np.array([1.0, 4, 3])
print(y)
print(y.shape)
y_13 = y[np.newaxis,:] # use '_AB' suffix to denote an array with shape (A, B)
print(y_13)
print(y_13.shape)
y_31 = y[:, np.newaxis] # use '_AB' suffix to denote an array with shape (A, B)
print(y_31)
print(y_31.shape)
y_311 = y[:, np.newaxis, np.newaxis] # use '_AB' suffix to denote an array with shape (A, B)
print(y_311)
print(y_311.shape)
```
<a id="elementwise_multiplication"></a>
# Elementwise Multiplication
To perform *element-wise* multiplication, use '*' symbol
```
print(y)
print(M)
R = M * M
print(R)
# What happens when we multiply (3,3) shape by a (3,) shape?
# y is implicitly expanded to (1,3) and thus multiplied element-wise to each row
M * y
M * y[np.newaxis,:]
M * y[:,np.newaxis] # this makes y multiplied to each column
```
In NumPy, multiplying an (M,N) array by an (M,) array is known as *broadcasting*
NumPy's implicit rules for what happens are defined here:
https://numpy.org/doc/stable/user/basics.broadcasting.html
<a id="matrix_multiplication"></a>
# Matrix multiplication
To do matrix multiplication, use np.dot
```
np.dot(M, y)
np.dot(M, M)
np.dot(y,y) # when applied to a 1-dim array, this is an inner product
np.dot(y[np.newaxis,:], y[:,np.newaxis])
np.sum(np.square(y))
```
<a id="pseudorandom_number_generation"></a>
# Pseudorandom number generation
```
x = np.random.uniform(size=15) # Float values uniformly distributed between 0 and 1
print(x)
x = np.random.normal(size=15) # Float values normally distributed according to 'standard' normal (mean 0, variance 1)
print(x)
```
To make *repeatable* pseudo-randomness, use a generator with the same seed!
```
seedA = 0
seedB = 1111
prng = np.random.RandomState(seedA)
prng.uniform(size=10)
prng = np.random.RandomState(seedA)
prng.uniform(size=10)
prng = np.random.RandomState(seedB)
prng.uniform(size=10)
```
<a id="useful_functions"></a>
# Useful functions
#### linspace and logspace
```
# Linearly spaced numbers
x_N = np.linspace(-2, 2, num=5)
for a in x_N:
print(a)
# Logarithmically spaced numbers
x_N = np.logspace(-2, 2, base=10, num=5)
for a in x_N:
print(a)
```
#### arange
```
# Start at 0 (default), count up by 1 (default) until you get to 4 (exclusive)
x = np.arange(4)
print(x)
# Start at negative PI, count up by increments of pi/4 until you get to + PI (exclusive)
y = np.arange(start=-np.pi, stop=np.pi, step=np.pi/4)
print(y)
# Start at negative PI, count up by increments of pi/4 until you get to PI + very small number (exclusive)
y = np.arange(start=-np.pi, stop=np.pi + 0.0000001, step=np.pi/4)
print(y)
```
#### allclose
Useful when checking if entries in an array are "close enough" to some reference value
E.g. sometimes due to numerical issues of representation, we would consider 5.00002 as good as "5"
```
x_N = np.arange(4)
print(x_N)
x2_N = x_N + 0.000001
print(x2_N)
np.all(x_N == x2_N)
np.allclose(x_N, x2_N, atol=0.01) # 'atol' is *absolute tolerance*
np.allclose(x_N, x2_N, atol=1e-7) # trying with too small a tolerance will result in False
```
<a id="reductions"></a>
# Reductions
Some numpy functions like 'sum' or 'prod' or 'max' or 'min' that take in many values and produce fewer values.
These kinds of operations are known as "reductions".
Within numpy, any reduction function takes an optional 'axis' kwarg to specify specific dimensions to apply the reduction to
```
# 2D array creation
# R equivalent of matrix(1:16, 4 ,4))
A = np.arange(1, 17).reshape(4, 4).transpose()
A
np.sum(A) # sum of all entries of A
np.sum(A, axis=0) # sum across dim with index 0 (across rows)
np.sum(A, axis=1) # sum across dim with index 1 (across cols)
np.min(A, axis=1) # compute minimum across dim with index 1
```
<a id="indexing"></a>
# Indexing
```
# 2D array creation
# R equivalent of matrix(1:16, 4 ,4))
A = np.arange(1, 17).reshape(4, 4).transpose()
A
# Show the first row
A[0]
# Show the first col
A[:, 0]
# Grab the second row, third column
A[1,2]
# select a range of rows and columns
A[0:3, 1:4]
# select a range of rows and all columns
A[0:2,:]
# select the *last* row
A[-1]
# select the *second to last* column
A[:, -2]
```
| github_jupyter |
<center>
<img src="https://drive.google.com/uc?id=12fkBBarn5tldtws1MLZ8aZ_Tw87kPnBp"/>
<h1> City of Rochester - Business Analytics Project </h1>
<h2> Xiaodan Ding, Pin Li, Jiawen Liang, Ruiling Shen, Chenxi Tao </h2>
</center>
# 1. Overview
In the codebook, we walk you through the data cleaning, data augmentation, and predictive modeling process through which we select out the targeted brand **Walgreens** for downtown Rochester.
The outline of this codebook is as followed: </br>
* Data Cleaning
- Reverse geocode
- Format zipcode
* Get American Community Service Data
* Data Augmentation
* Brand-selection Predictive Model
* Walgreens-prediction Model
# 2. Data Cleaning
## 2.1 Reverse Geocode
### 2.1.1 Reverse Geocode Zipcode
```
# Install Package to reverse geocoding.
!pip install geopy
# Load Packages
import numpy as np
import pandas as pd
# Load Business Location Data
sm = pd.read_csv(".../POIN_MASTER_010319.csv")
# pre-selected targetted list from the business location data.
# in the following analysis, our group only focus on those stores.
chain_store_list = ['GEOCVS','GEODD','GEODICK','GEOHDPT','GEOHGOOD','GEOJCP',
'GEOKOHL','GEOKRGR','GEOLOWE','GEOMACYS','GEOMASS','GEOODEPT',
'GEOROSS','GEOSAFEW','GEOSAV','GEOSEAR','GEOTJ','GEOTRGT',
'GEOWALG','GEOWMT']
# remove unrelevant business from the original data frame.
sm = sm[sm['gitext'].isin(chain_store_list)]
sm.info()
```
In this data frame, there are 7392 rows that have zipcode information as NaN. Since we will use zip code as our primary granularity, we have to adopt the **reverse-geocoding** method to retrieve the zip codes.
However, since the free reverse geocoding package is time-consuming, we do not need to retrieve the zipcode information that is already available in the data frame. Thus, we only reverse-geocoded those rows that have no zip code information.
```
sm_isna = pd.read_csv(".../temp_sm_isna.csv")
sm_isnotna = sm[sm['zip'].isna() == False]
sm_isna.info()
```
-- reverse geocode is the method to retrieve zip code information from spatial coordinates
we use the geopy.geocoders packages
``` Python
# Import Packages
from geopy.geocoders import Nominatim
import geopy.geocoders
from geopy.geocoders import Nominatim
geolocator = Nominatim(timeout=3)
from geopy.extra.rate_limiter import RateLimiter
geocode = RateLimiter(geolocator.reverse, min_delay_seconds=1)
for i in range(len(df)):
try:
df.loc[i,'zipcode'] = geocode(str(df.iloc[i]['latitude']) + ',' + str(df.iloc[i]['longitude'])).raw['address']['postcode']
print(str(i) + ' '+ str(df.iloc[i]['zipcode']))
except KeyError:
print(KeyError)
continue
```
We get the full dataset after utilizing the above method on those rows missing zip code information, as well as hardcoding those zip codes that cannot be retrieved from the above method.
```
sm_new = pd.concat([sm_isna,sm_isnotna],axis=0)
sm_new.drop(columns=['Unnamed: 0'],inplace=True)
# sm_new.to_csv("reverse_geocodone.csv")
```
### 2.1.2 Reverse Geocode City & States
Although our primary focus is on zipcode-level data, we still need to retrieve city and state information for our target stores in order to conduct peer-city analysis introduced in our pitch deck.
To retrieve city and state data we use a different reverse geocode method instead. Below are the Python codes:
```Python
!pip install reverse_geocoder
!pip install pprint
import reverse_geocoder as rg
import pprint
import time
df['cor'] = list(zip(df.latitude, df.longitude)) # format the zipcode in a standard form.
def find_city_name(cor):
city_name = rg.search(cor)[0]['name']
return city_name
def find_state(cor):
state_name = rg.search(cor)[0]['admin1']
return state_name
```
Our final dataset for peer city analysis can be found in our hand-in file: **peercity.csv**
## 2.2 Format zipcode
-- In this section, we further processed the data. We found out some exotic zipcode patterns and we need to adjust them into a uniform format or to drop them. Those exotic zip codes can be summarized by the following formats:
1. xxxxx-xxxx
2. canadian zip codes
3. xxxx and xxx (missing 0 before the first digit)
```
# sm_new = pd.read_csv(".../reverse_geocodone.csv")
# sm_new.drop(columns=['Unnamed: 0'],inplace=True)
f = sm_new[['gitext', 'poiname', 'addr1', 'city', 'state','zip','latitude','longitude']]
# Remove canadian zipcode
f = f[~f.zip.str.match('[a-zA-Z]')]
# select out the rows that zip is like xxxxx-xxxx
a = f[f.zip.str.match('^\d{5}-')]
a['zip'] = a['zip'].apply(lambda x:x[:5])
# select out the rows that zip is like xxxxx:xxxxx
b = f[f.zip.str.match('^\d{5}:\d{5}')]
b['zip'] = b['zip'].apply(lambda x:x[:5])
# select out the rows that zip length is less than 5 and add 0 in front of it
c = f[f['zip'].apply(lambda x: len(x)<5)]
c['zip'] = c['zip'].apply(lambda x: '{0:0>5}'.format(x))
# select out the rows that zip is like xxxxx:...
d = f[f.zip.str.match('^\d{5};')]
d['zip'] = d['zip'].apply(lambda x:x[:5])
# zipcode that is already in a desired format
e = f[f.zip.str.match('^\d{5}$')]
# concat them together to get the final business location dataset
final = pd.concat([a,b,c,d,e],axis=0)
# final.to_csv("full_zip.csv")
final.info()
```
# 3. Get American Community Survey Data
The American Community Survey (ACS) is an ongoing survey by the U.S. Census Bureau. It regularly gathers information previously contained only in the long form of the decennial census, such as ancestry, citizenship, educational attainment, income, language proficiency, migration, disability, employment, and housing characteristics. These data are used by many public-sector, private-sector, and not-for-profit stakeholders to allocate funding, track shifting demographics, plan for emergencies, and learn about local communities. Sent to approximately 295,000 addresses monthly (or 3.5 million per year), it is the largest household survey that the Census Bureau administers.
In this section, we show how we use census Application Programming Interface (API) tools to extract our pre-selected ACS attributes for zipcodes that are generated through the reverse geocoding process.
The main Python Code is shown below:
``` Python
import requests
import json
url = f'https://api.census.gov/data/2018/acs/acs5?key={apiKey}&get={filed}&for=zip%20code%20tabulation%20area:{a}'
response = requests.get(url)
#load the response into a JSON, ignoring the first element which is just field labels
formattedResponse = json.loads(response.text)[1:]
#flip the order of the response from [population, zipcode] -> [zipcode, population]
formattedResponse = [item[::-1] for item in formattedResponse]
```
**Notes**:
1. you can get your apiKey through: https://www.census.gov/developers/ and click the Request a KEY button on the left.
2. in the url: filed is the pre-selected attributes that you want to get, and attributes of ACS can be found in this link: https://api.census.gov/data/2018/acs/acs5/variables.html
3. You can also extract data on a different granularity rather than zipcode.
4. The maximum number of zipcode you can retrive from one api call is 1000. So in the code below I write a for loop to retrive data for 10200 zipcode.
```
# load the zipcodes from that are generated from the reverse-geocoding process.
fl = open(".../zip.txt",'w')
for i in zipz:
fl.write(i+'\n')
fl.close()
zips = open('.../zip.txt', 'r').readlines()
zips = [z.replace('\n', '') for z in zips]
# print(len(zips))
# load the pre-selected attributes.
filed = open('.../field of interest.txt', 'r').readlines()
filed = [h.replace('\n', '') for h in filed]
print(len(filed))
filed = ','.join(filed)
df = pd.DataFrame(columns=["zipcode","B02001_001E","B02001_002E","B02001_003E","B02001_004E","B02001_005E",
"B02001_006E","B02001_007E","B02001_008E","B01003_001E","B08301_002E","B08301_003E",
"B08301_004E","B08301_005E","B08301_006E","B08301_007E","B08301_008E","B08301_009E",
"B08301_010E","B08301_011E","B08301_012E","B08301_013E","B08301_014E","B08301_015E",
"B08301_016E","B08301_017E","B08301_018E","B08301_019E","B08301_020E","B08301_021E",
"B08014_002E","B08014_003E","B08014_004E","B08014_005E","B08014_006E","B08014_007E"])
for i in range(102):
a = zips[i*100:(1+i)*100]
a = ','.join(a)
url = f'https://api.census.gov/data/2018/acs/acs5?key={apiKey}&get={filed}&for=zip%20code%20tabulation%20area:{a}'
response = requests.get(url)
#load the response into a JSON, ignoring the first element which is just field labels
formattedResponse = json.loads(response.text)[1:]
#flip the order of the response from [population, zipcode] -> [zipcode, population]
formattedResponse = [item[::-1] for item in formattedResponse]
#store the response in a dataframe
zippp = pd.DataFrame(columns=["zipcode","B02001_001E","B02001_002E","B02001_003E","B02001_004E","B02001_005E",
"B02001_006E","B02001_007E","B02001_008E","B01003_001E","B08301_002E","B08301_003E",
"B08301_004E","B08301_005E","B08301_006E","B08301_007E","B08301_008E","B08301_009E",
"B08301_010E","B08301_011E","B08301_012E","B08301_013E","B08301_014E","B08301_015E",
"B08301_016E","B08301_017E","B08301_018E","B08301_019E","B08301_020E","B08301_021E",
"B08014_002E","B08014_003E","B08014_004E","B08014_005E","B08014_006E","B08014_007E"],
data=formattedResponse)
df = pd.concat([df,zippp],axis=0)
df = df[df['B02001_001E'].isna() == False]
```
#4. Data Augmentation
In this section, we combine the ACS data, mosaic data, and business location data into a format for our modelling process.
```
ma = pd.read_csv(".../mosaic_byZip.csv")
# format the mosaic data's zipcode column into standard xxxxx
ma['zip'] = ma['zip'].apply(lambda x: '{0:0>5}'.format(x))
cs_and_ma = df.merge(ma,how='left',left_on='zipcode',right_on='zip')
cs_and_ma.drop(columns=['zip'],inplace=True)
# Creating wide table for further multi-labeled machine learning
final = final[final['zip'].isin(zip_final)]
final = final[['gitext','zip']]
final = final.astype(str)
final['existence'] = [True] * len(final)
final_wide = final.pivot_table(index='zip',columns='gitext',values='existence')
final_wide.fillna(False,inplace = True)
# Combine the business location data with the census + mosaic dataset
final = final_wide.reset_index()
final = final.merge(cs_and_ma,how='left',left_on='zip',right_on='zipcode')
final.drop(columns=['zipcode'],inplace=True)
# final.to_csv("fulldata.csv")
```
# 5. Brand-selection Predictive Model
In this section we find an optimal machine learning model to make a prediction for the probability of our 20 targeted brands on Rochester downtown zipcodes: 14604, 14608, 14614.
## 5.1 Data-processing before modelling
```
raw = final
# we drop columns where either do not contain ACS information or Mosaic information.
raw.dropna(axis=0,inplace=True) # there are 9855 zipcodes left.
# create our multi-lable dependent variables
label = raw.loc[:,'GEOCVS':'GEOWMT']
y = label*1
# y.shape
# create our independent variabels into X
X = raw.loc[:,'B02001_001E':'political_affiliation_p111975_n']
X.shape
```
## 5.2 Feature Scaling
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_stan = scaler.fit_transform(X)
```
## 5.3 Model Setup
In this step we tried random forest multi-lable classification and several deep neural network to find the optimal predicitive model.
```
# Train test split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_stan, y, test_size=0.2, random_state=101)
from sklearn.metrics import accuracy_score,roc_curve, auc, roc_auc_score
from sklearn.model_selection import KFold, cross_val_score
from sklearn import metrics
from sklearn.metrics import classification_report
n_folds = 10
def get_CVacc(model):
"""
Return the accuracy score
"""
# Set KFold to shuffle data before the split
kf = KFold(n_folds, shuffle=True, random_state=42)
# Get accuracy score
accuracy_score = cross_val_score(model, X_stan, y, scoring="accuracy", cv=kf)
return accuracy_score.mean()
def get_acc(model):
"""
Return accuracy score
"""
model.fit(X_train, y_train)
predictions = model.predict(X_test)
acc = accuracy_score(y_test,predictions)
print(classification_report(y_test, predictions))
return acc
```
### 5.3.1 Random Forest
```
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(n_estimators=500)
print("accuracy_rfc: ", get_acc(rfc))
```
### 5.3.2 Deep Neural Net
```
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation,Dropout,BatchNormalization
from tensorflow.keras.constraints import max_norm
# Model 1:
model = Sequential()
# input layer
model.add(Dense(246, activation='sigmoid'))
model.add(Dropout(0.5))
# hidden layer
model.add(Dense(64, activation='sigmoid'))
model.add(Dropout(0.5))
# hidden layer
model.add(Dense(128, activation='sigmoid'))
model.add(Dropout(0.5))
# hidden layer
model.add(Dense(32, activation='sigmoid'))
model.add(Dropout(0.5))
# output layer
model.add(Dense(units=20,activation='sigmoid'))
# Compile model
model.compile(loss='binary_crossentropy', optimizer='adam',metrics=['accuracy'])
# Model 2:
model2 = Sequential()
model2.add(Dense(246, input_dim= 246, activation='sigmoid'))
model2.add(Dense(128, activation='sigmoid'))
model2.add(Dense(64, activation='sigmoid'))
model2.add(Dropout(0.5))
model2.add(Dense(64, activation='sigmoid'))
model2.add(Dense(32, activation='sigmoid'))
model2.add(Dropout(0.5))
model2.add(BatchNormalization())
model2.add(Dense(20, activation='softmax'))
# Compile model
model2.compile(loss='binary_crossentropy', optimizer='adam',metrics=['accuracy'])
# Model 3:
model3 = Sequential()
model3.add(Dense(246, kernel_initializer="uniform", activation='sigmoid'))
model3.add(Dense(64, activation='sigmoid'))
model3.add(Dense(32, activation='sigmoid'))
model3.add(Dense(20, activation='sigmoid'))
model3.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# After comparison we use the first model:
model.fit(x=X_train,
y=y_train,
epochs=20,
batch_size=128,
validation_data=(X_test, y_test),
)
```
## 5.4 Predict Brand
```
test = pd.read_csv('.../roc.csv')
testset = test.loc[:,'B02001_001E':'political_affiliation_p111975_n']
model.fit(X_stan, y)
predictions_nn = model.predict(testset)
output_nn = pd.DataFrame({
'14608': predictions_nn[0],
'14604': predictions_nn[1],
'14614': predictions_nn[2]},index=y.columns)
output_nn['mean']=np.mean(output_nn,axis=1)
output_nn.sort_values(by='mean',ascending=False,inplace=True)
output_nn
```

# 6. Walgreens-prediction Model
In this selection, combining with our peer-city analysis, we choose Walgreens as our final target.
We will use binary classification machine learning models to justify that it is promising to open Walgreens at downtown Rochester, and to offer some insights on the site-selection process introduced in our pitch deck.
```
label2 = raw.loc[:,'GEOWALG']
y2 = label2*1
x_train, x_test, y2_train, y2_test = train_test_split(X_stan, y2, test_size=0.2, random_state=101)
def plotROC(model):
"""
1. Plot ROC AUC
2. Return the best threshold
"""
model.fit(x_train, y2_train)
predictions = model.predict(x_test)
probs = model.predict_proba(x_test)
preds = probs[:,1]
fpr, tpr, threshold = roc_curve(y2_test, preds)
roc_auc = auc(fpr, tpr)
# Plot ROC AUC
plt.title('Receiver Operating Characteristic')
plt.plot(fpr, tpr, 'b', label = 'AUC = %0.2f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# report
acc = accuracy_score(y2_test,predictions)
print(classification_report(y2_test, predictions))
# Find optimal threshold
rocDf = pd.DataFrame({'fpr': fpr, 'tpr':tpr, 'threshold':threshold})
rocDf['tpr - fpr'] = rocDf.tpr - rocDf.fpr
optimalThreshold = rocDf.threshold[rocDf['tpr - fpr'].idxmax()]
return acc
```
## 6.1 Random Forest Classifier
```
rfc2 = RandomForestClassifier(n_estimators=500)
print("accuracy_rfc: ", plotROC(rfc2))
```
## 6.2 LightGBM
```
from lightgbm import LGBMClassifier
lgb = LGBMClassifier(objective='binary',
learning_rate=0.049,
n_estimators=1500,
num_leaves=8,
min_data_in_leaf=4,
max_depth=3,
max_bin=41,
bagging_fraction=0.845,
bagging_freq=5,
feature_fraction=0.24,
feature_fraction_seed=9,
bagging_seed=9,
min_sum_hessian_in_leaf=11)
print("accuracy_lgb: ", plotROC(lgb))
```
Since the LightGBM generates a slightly higher result, we choose LightGBM as our final walgreen-prediction model.
```
lgb.fit(X_stan, y2)
predictions = lgb.predict_proba(testset)[:, 1]
predictions
output = pd.DataFrame({'zipcode': test.zipcode,
'probability': predictions})
output
```
It turns out that three downtown zip codes all have very high probabilities on opening Walgreens, which justifies our previous brand selection model.
# 7. Conclusion
Thanks for your patience to read through our codebook, we sincerely wish that our explanation of our data analytical process is clear to you. Meanwhile, we wish this codebook serves as a strong appendix to our final pitch deck presentation.
```
```
| github_jupyter |
## Cycle GAN in PyTorch
```
%load_ext autoreload
%matplotlib inline
%autoreload 2
from IPython import display
from utils import Logger
import torch
from torch import nn, optim
from torch.autograd.variable import Variable
from torchvision import transforms, datasets
DATA_FOLDER = './torch_data/CycleGAN'
import os
import urllib, zipfile
from tqdm import tqdm
import zipfile
VALID_DATA_NAMES = ["ae_photos", "apple2orange", "summer2winter_yosemite", "horse2zebra", "monet2photo", "cezanne2photo", "ukiyoe2photo", "vangogh2photo", "maps", "cityscapes","facades","iphone2dslr_flower","mini", "mini_pix2pix", "mini_colorization"]
URL = 'https://people.eecs.berkeley.edu/~taesung_park/CycleGAN/datasets/{}.zip'
class DownloadProgressBar(tqdm):
def update_to(self, b=1, bsize=1, tsize=None):
if tsize is not None:
self.total = tsize
self.update(b * bsize - self.n)
def rename_images(rootdir):
for subdir, dirs, files in os.walk(rootdir):
idx = 0
for file in files:
file_path = os.path.join(subdir, file)
new_file_path = os.path.join(subdir, "{}.jpg".format(idx))
os.rename(file_path, new_file_path)
idx = idx + 1
def download_cyclegan_dataset(filename, path, force=False):
# Validate dataset filename is valid.
assert(filename in VALID_DATA_NAMES)
# Return if path exists.
file_path = "{}/{}.zip".format(path, filename)
if(os.path.exists(file_path) and not force): return
# Otherwise download.
# Make path directory if missing.
os.makedirs(path, exist_ok=True)
# Download data
url = URL.format(filename)
with DownloadProgressBar(unit='B', unit_scale=True,
miniters=1, desc=url.split('/')[-1]) as t:
urllib.request.urlretrieve(url, file_path, reporthook=t.update_to)
with zipfile.ZipFile(file_path, 'r') as zip_obj:
zip_obj.extractall(path)
rename_images(os.path.join(path, filename))
for name in VALID_DATA_NAMES:
download_cyclegan_dataset(name, DATA_FOLDER, force=False)
from torch.utils.data import Dataset
from PIL import Image
from skimage import io
class MyDataset(Dataset):
"""My dataset."""
def __init__(self, path, transform=None):
self._path = path
self.num_files = len(os.listdir(self._path))
self.transform = transform
def __len__(self):
return self.num_files
def filename(self, idx):
for directory in ["trainA", "testA"]:
possible_path = os.path.join(self._path, directory, "{}.jpg".format(str(idx)))
if os.path.exists(possible_path):
return possible_path
def __getitem__(self, idx):
# Handle vectors.
if torch.is_tensor(idx):
idx = idx.tolist()
img_name = self.filename(idx)
print(img_name)
image = Image.fromarray(io.imread(img_name))
if self.transform:
image = self.transform(image)
return image
def visualize(self, idx):
np_img = np.transpose(self[idx].numpy(), (1,2, 0))
plt.imshow(np_img)
```
### Load Data
```
def my_data():
compose = transforms.Compose(
[transforms.Resize((80, 80)),
transforms.ToTensor(),
transforms.Normalize((.5,), (.5,))
])
return MyDataset(DATA_FOLDER + "/ae_photos", transform=compose)
import matplotlib.pyplot as plt
import numpy as np
# data = MyDataset(DATA_FOLDER + "/ae_photos")
data = my_data()
# Create loader with data, so that we can iterate over it.
# data_loader = torch.utils.data.DataLoader(my_data(), batch_size=10, shuffle=True)
# plt.imshow(np.flip(my_data()[48], 1))
# plt.imshow(my_data()[48][1])
data.visualize(101)
```
#### Discriminator
```
import torch
import torch.nn as nn
import torch.nn.functional as F
class DiscriminatorNet(torch.nn.Module):
def __init__(self):
super(DiscriminatorNet, self).__init__()
self.conv_0 = nn.Sequential(
nn.Conv2d(in_channels=3, out_channels=64,
kernel_size=(4, 4), stride=2),
nn.LeakyReLU(negative_slope=0.2, inplace=True)
)
self.conv_1 = nn.Sequential(
nn.Conv2d(in_channels=64, out_channels=128,
kernel_size=(4, 4), stride=2),
nn.BatchNorm2d(128),
nn.LeakyReLU(negative_slope=0.2, inplace=True)
)
self.conv_2 = nn.Sequential(
nn.Conv2d(in_channels=128, out_channels=256,
kernel_size=(4, 4), stride=2),
nn.BatchNorm2d(256),
nn.LeakyReLU(negative_slope=0.2, inplace=True)
)
self.conv_3 = nn.Sequential(
nn.Conv2d(in_channels=256, out_channels=512,
kernel_size=(4, 4), stride=2),
nn.BatchNorm2d(512),
nn.LeakyReLU(negative_slope=0.2, inplace=True)
)
self.conv_out = nn.Conv2d(in_channels=512, out_channels=1,
kernel_size=(4, 4), stride=2)
def forward(self, x):
x = self.conv_0(x)
x = self.conv_1(x)
x = self.conv_2(x)
x = self.conv_3(x)
x = self.conv_out(x)
d = DiscriminatorNet()
```
#### Generator net
```
import torch
import torch.nn as nn
import torch.nn.functional as F
class GeneratorNet(torch.nn.Module):
def __init__(self):
super(GeneratorNet, self).__init__()
# c7s1-64
self.conv_0 = nn.Sequential(
nn.Conv2d(in_channels=3, out_channels=64, kernel_size=(7, 7), stride=1,
padding_mode='reflect'),
nn.BatchNorm2d(64),
nn.ReLU(inplace=True)
)
# d128
self.conv_1 = nn.Sequential(
nn.Conv2d(in_channels=64, out_channels=128, kernel_size=(3, 3), stride=2,
padding_mode='reflect'),
nn.BatchNorm2d(64),
nn.ReLU(inplace=True)
)
# d256
self.conv_2 = nn.Sequential(
nn.Conv2d(in_channels=128, out_channels=256, kernel_size=(3, 3), stride=2,
padding_mode='reflect'),
nn.BatchNorm2d(64),
nn.ReLU(inplace=True)
)
# R256
self.res_3 = nn.Conv2d(in_channels=256, out_channels=256, kernel_size=(3, 3),
stride=2),
# R256
self.res_4 = nn.Conv2d(in_channels=256, out_channels=256, kernel_size=(3, 3),
stride=2),
# R256
self.res_5 = nn.Conv2d(in_channels=256, out_channels=256, kernel_size=(3, 3),
stride=2),
# R256
self.res_6 = nn.Conv2d(in_channels=256, out_channels=256, kernel_size=(3, 3),
stride=2),
# R256
self.res_7 = nn.Conv2d(in_channels=256, out_channels=256, kernel_size=(3, 3),
stride=2),
# R256
self.res_8 = nn.Conv2d(in_channels=256, out_channels=256, kernel_size=(3, 3),
stride=2),
# u128
self.ups_9 = nn.Sequential(
nn.ConvTranspose2d(in_channels=256, out_channels=128,
kernel_size=(3, 3), stride=0.5),
nn.BatchNorm2d(128),
nn.ReLU(inplace=True)
)
# u64
self.ups_10 = nn.Sequential(
nn.ConvTranspose2d(in_channels=128, out_channels=64, kernel_size=(3, 3),
stride=0.5),
nn.BatchNorm2d(64),
nn.ReLU(inplace=True)
)
# c7s1-3
self.conv_0 = nn.Sequential(
nn.Conv2d(in_channels=3, out_channels=3, kernel_size=(7, 7), stride=1,
padding_mode='reflect'),
nn.BatchNorm2d(64),
nn.ReLU(inplace=True)
)
# TODO(diegoalejogm): Add skip layer connections.
def forward(self, x):
x = self.conv_0(x)
x = self.conv_1(x)
x = self.conv_2(x)
x = self.res_3(x)
x = self.res_4(x)
x = self.res_5(x)
x = self.res_6(x)
x = self.res_7(x)
x = self.res_8(x)
x = self.ups_9(x)
x = self.ups_10(x)
x = self.conv_11(x)
return x
g = GeneratorNet()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import random as rd
import matplotlib.pyplot as plt
from IPython.core.interactiveshell import InteractiveShell # 显示所有输出
InteractiveShell.ast_node_interactivity = "all"
data = pd.read_csv("films.csv")
# X = data[["Category","Time"]]
from sklearn import preprocessing
le = preprocessing.LabelEncoder()
features = ['Category']
for feature in features:
#非数字型和数字型标签值标准化
le.fit(data[feature])
data[feature] = le.transform(data[feature])
data['Category']
X = data[["Category","Time"]]
#Visualise data points
plt.figure(dpi=150)
plt.scatter(X["Category"],X["Time"],c='blue')
plt.xlabel('Category')
plt.ylabel('Time (minute)')
plt.savefig('C:\\Users\AoSun\Desktop\cluster\C-T cluster1.png',bbox_inches = 'tight')
plt.show()
from sklearn import preprocessing
X_scale = X.values[:,1]
X_scale
scaler = preprocessing.StandardScaler().fit(X_scale.reshape(-1,1))
print(scaler.mean_)
print(scaler.scale_)
X_scaled = scaler.transform(X_scale.reshape(-1,1))
X_scale.reshape(-1,1).shape
X_scaled.shape
X_scaled_csv = pd.Series(X_scaled.reshape(1,-1)[0])
# 将其重新格式化以导出到Kaggle
X['Time'] = pd.Series(X_scaled.reshape(1, -1)[0])
#Visualise data points
plt.figure(dpi=150)
plt.scatter(X["Category"],X["Time"],c='blue')
plt.xlabel('Category')
plt.ylabel('Time')
plt.savefig('C:\\Users\AoSun\Desktop\cluster\C-T cluster2.png')
plt.show()
K=3
# Select random observation as centroids
Centroids = (X.sample(n=K)) # 随机抽样三个点
plt.figure(dpi=150)
plt.scatter(X["Category"],X["Time"],c='blue')
plt.scatter(Centroids["Category"],Centroids["Time"],c='red')
plt.xlabel('Category')
plt.ylabel('Time')
plt.savefig('C:\\Users\AoSun\Desktop\cluster\C-T cluster3.png')
plt.show()
diff = 1
j=0
while(diff!=0):
XD=X
i=1
for index1,row_c in Centroids.iterrows():
ED=[] # 记录每个点离质心的距离
for index2,row_d in XD.iterrows():
d1=(row_c["Category"]-row_d["Category"])**2
d2=(row_c["Time"]-row_d["Time"])**2
d=np.sqrt(d1+d2)
ED.append(d)
X[i]=ED # 各个点离质心距离
i=i+1
C=[]
for index,row in X.iterrows(): # 判断点离哪个质心最近
min_dist=row[1]
pos=1
for i in range(K): # i从0开始
if row[i+1] < min_dist:
min_dist = row[i+1]
pos=i+1
C.append(pos)
X["Cluster"]=C
Centroids_new = X.groupby(["Cluster"]).mean()[["Time","Category"]] # 求各簇的质心
if j == 0:
diff=1
j=j+1
else:
diff = (Centroids_new['Time'] - Centroids['Time']).sum() + (Centroids_new['Category'] - Centroids['Category']).sum()
print(diff.sum()) # 质心不再改变时停止
Centroids = X.groupby(["Cluster"]).mean()[["Time","Category"]]
# Step 3 - Assign all the points to the closest cluster centroid
# Step 4 - Recompute centroids of newly formed clusters
# Step 5 - Repeat step 3 and 4
plt.figure(dpi=150)
color=['blue','green','cyan']
for k in range(K):
data=X[X["Cluster"]==k+1]
plt.scatter(data["Category"],data["Time"],c=color[k])
plt.scatter(Centroids["Category"],Centroids["Time"],c='red')
plt.xlabel('Category')
plt.ylabel('Time')
plt.savefig('C:\\Users\AoSun\Desktop\cluster\C-T cluster4.png')
plt.show()
```
| github_jupyter |
[musicinformationretrieval.com](https://musicinformationretrieval.com)
Jupyter Basics
=======================
You are looking at a **Jupyter Notebook**, an interactive Python shell inside of a web browser. With it, you can run individual Python commands and immediately view their output. It's basically like the Matlab Desktop or Mathematica Notebook but for Python.
To start an interactive Jupyter notebook on your local machine, read the [instructions at the GitHub `README` for this repository](https://github.com/stevetjoa/stanford-mir#how-to-use-this-repo).
If you are reading this notebook on <http://musicinformationretrieval.com>, you are viewing a read-only version of the notebook, not an interactive version. Therefore, the instructions below do not apply.
## Tour
If you're new, we recommend that you take the *User Interface Tour* in the Help Menu above.
## Cells
A Jupyter Notebook is comprised of **cells**. Cells are just small units of code or text. For example, the text that you are reading is inside a *Markdown* cell. (More on that later.)
*Code* cells allow you to edit, execute, and analyze small portions of Python code at a time. Here is a code cell:
```
1+2
```
## Modes
The Jupyter Notebook has two different keyboard input modes.
In **Edit Mode**, you type code/text into a cell. Edit Mode is indicated by a *green* cell border.
To enter Edit Mode from Command Mode, press `Enter`. You can also double-click on a cell.
To execute the code inside of a cell and move to the next cell, press **`Shift-Enter`**. (`Ctrl-Enter` will run the current cell without moving to the next cell. This is useful for rapidly tweaking the current cell.)
In **Command Mode**, you can perform notebook level actions such as navigating among cells, selecting cells, moving cells, saving notebooks, displaying help. Command Mode is indicated by a *grey* cell border.
To enter Command Mode from Edit Mode, press **`Esc`**. Other commands can also enter Command Mode, e.g. `Shift-Enter`.
To display the Help Menu from Command Mode, press **`h`**. *Use it often*; `h` is your best friend.
## Saving
Your code goes directly into a Jupyter notebook. To save your changes, click on the "Save" icon in the menu bar, or type **`s`** in command mode.
If this notebook is in a Git repo, use `git checkout -- <file>` to revert a saved edit.
## Writing Text in Markdown
Markdown is simply a fancy way of formatting plain text. It is a markup language that is a superset of HTML. The Markdown specification is found here: http://daringfireball.net/projects/markdown/basics/
A cell may contain Python code or Markdown code. To convert any Python cell to a Markdown cell, press **`m`**. To convert from a Markdown cell to a Python cell, press **`y`**.
For headings, we recommend that you use Jupyter's keyboard shortcuts. To change the text in a cell to a level-3 header, simply press `3`. For similar commands, press **`h`** to view the Help menu.
## Writing Text in $\LaTeX$
In a Markdown cell, you can also use $\LaTeX$ syntax. Example input:
$$ \max_{||w||=1} \sum_{i=1}^{N} \big| \langle w, x_i - m \rangle \big|^2 $$
Output:
$$ \max_{||w||=1} \sum_{i=1}^{N} \big| \langle w, x_i - m \rangle \big|^2 $$
## Imports
You may encounter the following imports while using this website:
```
import numpy
import scipy
import pandas
import sklearn
import seaborn
import matplotlib
import matplotlib.pyplot as plt
import librosa
import librosa.display
import IPython.display as ipd
```
You can also combine imports on one line:
```
import numpy, scipy, pandas
```
## Tab Autocompletion
Tab autocompletion works in Command Window and the Editor. After you type a few letters, press the `Tab` key and a popup will appear and show you all of the possible completions, including variable names and functions. This prevents you from mistyping the names of variables -- a big time saver!
For example, type `scipy.` and then press `Tab`. You should see a list of members in the Python package `scipy`.
Or type `scipy.sin`, then press `Tab` to view members that begin with `sin`.
```
# Press Tab at the end of the following line
scipy.sin
```
## Inline Documentation
To get help on a certain Python object, type `?` after the object name, and run the cell:
```
# Run this cell.
int?
```
In addition, if you press `Shift-Tab` in a code cell, a help dialog will also appear. For example, in the cell above, place your cursor after `int`, and press `Shift-Tab`. Press `Shift-Tab` twice to expand the help dialog.
## More Documentation: NumPy, SciPy, Matplotlib
In the top menu bar, click on Help, and you'll find a prepared set of documentation links for IPython, NumPy, SciPy, Matplotlib, and Pandas.
## Experimenting
Code cells are meant to be interactive. We may present you with several options for experimentation, e.g. choices of variables, audio files, and algorithms. For example, if you see a cell like this, then try all of the possible options by uncommenting the desired line(s) of code. (To run the cell, select "Cell" and "Run" from the top menu, or press `Shift-Enter`.)
```
x = scipy.arange(50)
# Try these too:
# x = scipy.randn(50)
# x = scipy.linspace(0, 1, 50, endpoint=False)
x
```
| github_jupyter |
```
!wget = 'https://raw.githubusercontent.com/Doodies/Github-Stars-Predictor/master/PreprocessData.csv'
!ls -lh
```
# Importing required modules
```
!pip install catboost ipywidgets
!pip install xgboost
!pip install keras
# Handle table-like data and matrices
import numpy as np
import pandas as pd
# Visualisation
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
import seaborn as sns
# Configure visualisations
%matplotlib inline
color = sns.color_palette()
pd.options.mode.chained_assignment = None
pd.options.display.max_columns = 999
# mpl.style.use( 'ggplot' )
sns.set_style( 'whitegrid' )
pylab.rcParams[ 'figure.figsize' ] = 10,8
seed = 7
# importing libraries
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_squared_error
from sklearn.ensemble import GradientBoostingRegressor
from catboost import CatBoostRegressor
from sklearn.ensemble import RandomForestRegressor
from keras.models import Sequential
from keras.layers import Dense, Dropout, BatchNormalization
from keras.wrappers.scikit_learn import KerasRegressor
from sklearn.model_selection import cross_val_score, KFold
from sklearn.pipeline import Pipeline
np.random.seed(seed)
data = pd.read_csv('../dataset/PreprocessData.csv').iloc[:, 1:]
X = data.drop(['stars'] , axis =1)
y = data.stars
```
# Feature normalization
```
s = StandardScaler()
X = s.fit_transform(X)
X
```
# train test data splitting
```
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.10, random_state=42)
training_scores = []
test_scores = []
models = []
```
# Training Models
## 1. Gradient Boost
```
models.append("gradient boost")
reg = GradientBoostingRegressor(verbose = 1, n_estimators = 500)
reg.fit(X_train , y_train)
training_score = reg.score(X_train, y_train)
test_score = reg.score(X_test, y_test)
training_scores.append(training_score)
test_scores.append(test_score)
print("training set performance: ", training_score)
print("test set performance: ", test_score)
pred = reg.predict(X_test).astype(int)
temp1 = y_test.values > 0
# plt.axis((0,1000,0,1000))
plt.scatter(y_test.values[temp1], pred[temp1])
plt.xlabel("original", fontsize=12)
plt.ylabel("predictions", fontsize=12)
plt.show()
```
## 2. Cat Boost
```
models.append("cat boost")
model = CatBoostRegressor(iterations= 440 , depth= 8 , learning_rate= 0.1 , loss_function='RMSE' , use_best_model=True)
model.fit(X_train[:90503], y_train[:90503] , eval_set=(X_train[90503:], y_train[90503:]),plot=True)
y_train_pred = model.predict(X_train)
y_pred = model.predict(X_test)
train_score = r2_score(y_train , y_train_pred)
test_score = r2_score(y_test, y_pred)
training_scores.append(training_score)
test_scores.append(test_score)
print("Training score - " + str(train_score))
print("Test score - " + str(test_score))
```
## 3. Random Forest
```
models.append("random forest")
model = RandomForestRegressor(n_jobs=-1, n_estimators=10, verbose=1, random_state=seed)
model.fit(X_train, y_train)
training_score = model.score(X_train, y_train)
test_score = model.score(X_test, y_test)
print("training score: ", training_score)
print("test score: ", test_score)
training_scores.append(training_score)
test_scores.append(test_score)
```
## 4. Neural Network
```
models.append("neural network")
def baseline_model():
model = Sequential()
model.add(Dense(100,input_dim=54, activation='relu', kernel_initializer='glorot_normal'))
model.add(Dropout(0.2))
model.add(Dense(80, activation='relu', kernel_initializer='glorot_normal'))
model.add(Dropout(0.2))
model.add(Dense(60, activation='relu', kernel_initializer='glorot_normal'))
model.add(Dropout(0.2))
model.add(Dense(40, activation='relu', kernel_initializer='glorot_normal'))
model.add(Dropout(0.2))
model.add(Dense(20, activation='relu', kernel_initializer='glorot_normal'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='relu', kernel_initializer='glorot_normal'))
model.add(Dropout(0.2))
model.add(Dense(1, kernel_initializer='glorot_normal'))
model.compile(loss='mean_squared_error', optimizer='adam')
return model
estimator = KerasRegressor(build_fn=baseline_model, nb_epoch=200, epochs=10, batch_size=32, verbose=True)
# kfold = KFold(n_splits=10, random_state=seed)
# results = cross_val_score(estimator, X_train.values, y_train.values, cv=kfold)
# print("Results: %.2f (%.2f) MSE" % (results.mean(), results.std()))
estimator.fit(X_train, y_train)
train_pred = estimator.predict(X_train)
test_pred = estimator.predict(X_test)
training_score = r2_score(y_train, train_pred)
test_score = r2_score(y_test, test_pred)
print("training score: ", training_score)
print("test score: ", test_score)
training_scores.append(training_score)
test_scores.append(test_score)
```
# Comparing all model's results
```
print(models)
print(training_scores)
print(test_scores)
SMALL_SIZE = 8
MEDIUM_SIZE = 10
BIGGER_SIZE = 12
mpl.rc('font', size=MEDIUM_SIZE) # controls default text sizes
mpl.rc('axes', titlesize=18) # fontsize of the axes title
mpl.rc('axes', labelsize=16) # fontsize of the x and y labels
mpl.rc('xtick', labelsize=MEDIUM_SIZE) # fontsize of the tick labels
mpl.rc('ytick', labelsize=MEDIUM_SIZE) # fontsize of the tick labels
mpl.rc('legend', fontsize=BIGGER_SIZE) # legend fontsize
mpl.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
y_pos = np.arange(len(models))
rects1 = plt.bar(y_pos-0.12, training_scores, 0.35, alpha=0.8, color=color[2], label="train score")
rects2 = plt.bar(y_pos+0.13, test_scores, 0.35, alpha=0.8, color=color[5], label="test score")
plt.xticks(y_pos, models)
plt.xlabel('Models')
plt.title('R2 scores of different models')
plt.legend()
def autolabel(rects):
"""
Attach a text label above each bar displaying its height
"""
for rect in rects:
height = rect.get_height()
plt.text(rect.get_x() + rect.get_width()/2., 1.01*height,
'%.2f' % float(height),
ha='center', va='bottom')
autolabel(rects1)
autolabel(rects2)
plt.show()
```
| github_jupyter |
# Initialization
```
import os
import itertools
import sys
from math import factorial as fac
sys.path.append("D:/TU_Delft/Msc_Building_Technology/Semester_3/Graduation/Aditya_Graduation_Project_BT/06_Libraries")
import topogenesis as tg
import pyvista as pv
import trimesh as tm
import numpy as np
import networkx as nx
np.random.seed(0)
np.set_printoptions(threshold=sys.maxsize)
import networkx as nx
from itertools import combinations
from itertools import permutations
import pickle
import pandas as pd
lattice_path = os.path.relpath('Zoning_envelop.csv')
Zoning_matrix_sequential = tg.lattice_from_csv(lattice_path)
lattice_path_availability_matrix = os.path.relpath('voxelized_envelope_6m_voxel_size.csv')
avail_lattice_base = tg.lattice_from_csv(lattice_path_availability_matrix)
avail_lattice = avail_lattice_base*1
init_avail_lattice = tg.to_lattice(np.copy(avail_lattice*1), avail_lattice)
```
# Stencils
```
stencil_von_neumann = tg.create_stencil("von_neumann", 1, 1)
stencil_von_neumann.set_index([0,0,0], 0)
stencil_cuboid = tg.create_stencil("moore", 1, 1)
stencil_cuboid.set_index([0,0,0], 0)
# creating neighborhood definition
stencil_squareness = tg.create_stencil("moore", 1, 1)
# Reshaping the moore neighbourhood
stencil_squareness[0,:,:] = 0
stencil_squareness[2,:,:] = 0
stencil_squareness.set_index([0,0,0], 0)
stencil_squareness_t = np.transpose(stencil_squareness)
#print(stencil_squareness_t)
```
# Initial Vizualization
```
p = pv.Plotter(notebook=True)
base_lattice = Zoning_matrix_sequential
# Set the grid dimensions: shape + 1 because we want to inject our values on the CELL data
grid = pv.UniformGrid()
grid.dimensions = np.array(base_lattice.shape) + 1
# The bottom left corner of the data set
grid.origin = base_lattice.minbound - base_lattice.unit * 0.5
# These are the cell sizes along each axis
grid.spacing = base_lattice.unit
# adding the boundingbox wireframe
p.add_mesh(grid.outline(), color="grey", label="Domain")
init_avail_lattice.fast_vis(p)
# adding axes
p.add_axes()
p.show_bounds(grid="back", location="back", color="#aaaaaa")
# Add the data values to the cell data
grid.cell_arrays["Agents"] = Zoning_matrix_sequential.flatten(order="F").astype(int) # Flatten the array!
# filtering the voxels
threshed = grid.threshold([101,210])
# adding the voxels
p.add_mesh(threshed, name='sphere', show_edges=True, opacity=1.0, show_scalar_bar=False)
p.show(use_ipyvtk=True)
```
# Cleanup
```
def cleanup_algorithm (Zoning_matrix,stencil_1,stencil_2,a_id):
#First Cleanup
Zoning_matrix_flat = Zoning_matrix.flatten()
all_indices_for_agent = np.argwhere(Zoning_matrix_flat ==a_id).flatten()
all_vox_neighs_inds = Zoning_matrix.find_neighbours(stencil_1)[np.argwhere(Zoning_matrix_flat ==a_id)]
retrieved_neighs= Zoning_matrix_flat[all_vox_neighs_inds]
#Second Cleanup
for_improvement = np.copy(Zoning_matrix_flat)
for_improvement[np.argwhere(Zoning_matrix_flat==0)] =a_id
all_vox_neighs_inds_improv = Zoning_matrix.find_neighbours(stencil_2)[np.argwhere(Zoning_matrix_flat ==a_id)]
retrieved_neighs_improv = for_improvement[all_vox_neighs_inds_improv]
indexing =[]
for i, item in enumerate(retrieved_neighs_improv):
flattened_list = item.flatten()
#print(flattened_list)
if np.all(flattened_list[[0,1,2,7,3]] == a_id) or np.all(flattened_list[[3,4,5,6,7]] == a_id) or np.all(flattened_list[[1,2,3,4,5]] == a_id) or np.all(flattened_list[[5,6,7,0,1]] == a_id):
a = i
else:
indexing.append(i)
#print("truthy")
lattice_indexes= all_indices_for_agent[indexing]
return retrieved_neighs,all_indices_for_agent, lattice_indexes
PH_Unoccupy_indexes = cleanup_algorithm(Zoning_matrix_sequential,stencil_cuboid,stencil_squareness_t,200)
```
# Unoccupy process
```
def unoccupy_based_on_number(lattice,removal_indices,a_id,number):
list_of_neighbour_repetitions = (np.sum(np.count_nonzero(removal_indices[0] == a_id,axis =1),axis=1))
removal_indices= removal_indices[1][np.argwhere(list_of_neighbour_repetitions <= number)].flatten()
lattice[np.unravel_index(removal_indices,avail_lattice.shape)] = 0
unoccupy_based_on_number (Zoning_matrix_sequential,PH_Unoccupy_indexes,200,6)
#def unoccupy_based_on_stencil(lattice,removal_indices):
Zoning_matrix_sequential[np.unravel_index(PH_Unoccupy_indexes[2],Zoning_matrix_sequential.shape)] = 0
#unoccupy_based_on_stencil( Zoning_matrix_sequential,PH_Unoccupy_indexes[2])
PH_Unoccupy_indexes[2]
```
# Final Viz after cleanup
```
p = pv.Plotter(notebook=True)
base_lattice = Zoning_matrix_sequential
# Set the grid dimensions: shape + 1 because we want to inject our values on the CELL data
grid = pv.UniformGrid()
grid.dimensions = np.array(base_lattice.shape) + 1
# The bottom left corner of the data set
grid.origin = base_lattice.minbound - base_lattice.unit * 0.5
# These are the cell sizes along each axis
grid.spacing = base_lattice.unit
# adding the boundingbox wireframe
p.add_mesh(grid.outline(), color="grey", label="Domain")
init_avail_lattice.fast_vis(p)
# adding axes
p.add_axes()
p.show_bounds(grid="back", location="back", color="#aaaaaa")
# Add the data values to the cell data
grid.cell_arrays["Agents"] = Zoning_matrix_sequential.flatten(order="F").astype(int) # Flatten the array!
# filtering the voxels
threshed = grid.threshold([101,210])
# adding the voxels
p.add_mesh(threshed, name='sphere', show_edges=True, opacity=1.0, show_scalar_bar=False)
p.show(use_ipyvtk=True)
n = 8
n1 = 5
perms =[]
for x in itertools.combinations( range(n), n1 ) :
perms.append([ 1 if i in x else 0 for i in range(n)] )
perms
perms[0]
```
| github_jupyter |
```
import numpy as np
from sklearn.datasets import fetch_openml
from sklearn.utils.extmath import softmax
import matplotlib.pyplot as plt
from matplotlib import pyplot
from sklearn import metrics
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from mpl_toolkits.axes_grid1 import make_axes_locatable
plt.rcParams['font.family'] = 'serif'
plt.rcParams['font.serif'] = ['Times New Roman'] + plt.rcParams['font.serif']
```
## Load and display MNIST handwritten digits dataset
```
# Load data from https://www.openml.org/d/554
X, y = fetch_openml('mnist_784', version=1, return_X_y=True)
# X = X.values ### Uncomment this line if you are having type errors in plotting. It is loading as a pandas dataframe, but our indexing is for numpy array.
X = X / 255.
print('X.shape', X.shape)
print('y.shape', y.shape)
'''
Each row of X is a vectroization of an image of 28 x 28 = 784 pixels.
The corresponding row of y holds the true class label from {0,1, .. , 9}.
'''
# see how many images are there for each digit
for j in np.arange(10):
idx = np.where(y==str(j))
idx = np.asarray(idx)[0,:]
print('digit %i length %i' % (j, len(idx)))
# Plot some sample images
ncols = 10
nrows = 4
fig, ax = plt.subplots(nrows=nrows, ncols=ncols, figsize=[15, 6.5])
for j in np.arange(ncols):
for i in np.arange(nrows):
idx = np.where(y==str(j)) # index of all images of digit 'j'
idx = np.asarray(idx)[0,:] # make idx from tuple to array
idx_subsampled = np.random.choice(idx, nrows)
ax[i,j].imshow(X[idx_subsampled[i],:].reshape(28,28))
# ax[i,j].title.set_text("label=%s" % y[idx_subsampled[j]])
if i == 0:
# ax[j,i].set_ylabel("label=%s" % y[idx_subsampled[j]])
ax[i,j].set_title("label$=$%s" % y[idx_subsampled[i]], fontsize=14)
# ax[i].legend()
plt.subplots_adjust(wspace=0.3, hspace=-0.1)
plt.savefig('MNIST_ex1.pdf', bbox_inches='tight')
# Split the dataset into train and test sets
X_train = []
X_test = []
y_test = []
y_train = []
for i in np.arange(X.shape[0]):
# for each example i, make it into train set with probabiliy 0.8 and into test set otherwise
U = np.random.rand() # Uniform([0,1]) variable
if U<0.8:
X_train.append(X[i,:])
y_train.append(y[i])
else:
X_test.append(X[i,:])
y_test.append(y[i])
X_train = np.asarray(X_train)
X_test = np.asarray(X_test)
y_train = np.asarray(y_train)
y_test = np.asarray(y_test)
print('X_train.shape', X_train.shape)
print('X_test.shape', X_test.shape)
print('y_train.shape', y_train.shape)
print('y_test.shape', y_test.shape)
def sample_binary_MNIST(list_digits=['0','1'], full_MNIST=None, noise_rate=0):
# get train and test set from MNIST of given two digits
# e.g., list_digits = ['0', '1']
if full_MNIST is not None:
X, y = full_MNIST
else:
X, y = fetch_openml('mnist_784', version=1, return_X_y=True)
X = X / 255.
idx = [i for i in np.arange(len(y)) if y[i] in list_digits] # list of indices where the label y is in list_digits
X01 = X[idx,:]
y01 = y[idx]
X_train = []
X_test = []
y_test = [] # list of integers 0 and 1s
y_train = [] # list of integers 0 and 1s
for i in np.arange(X01.shape[0]):
# for each example i, make it into train set with probabiliy 0.8 and into test set otherwise
U = np.random.rand() # Uniform([0,1]) variable
label = 0
if y01[i] == str(list_digits[1]):
label = 1
if U<0.8:
# add noise to the sampled images
if noise_rate > 0:
for j in np.arange(X01.shape[1]):
U1 = np.random.rand()
if U1 < noise_rate:
X01[i,j] += np.random.rand()
X_train.append(X01[i,:])
y_train.append(label)
else:
X_test.append(X01[i,:])
y_test.append(label)
X_train = np.asarray(X_train)
X_test = np.asarray(X_test)
y_train = np.asarray(y_train).reshape(-1,1)
y_test = np.asarray(y_test).reshape(-1,1)
return X_train, X_test, y_train, y_test
X_train, X_test, y_train, y_test = sample_binary_MNIST(list_digits=['0','1'], full_MNIST=[X, y], noise_rate=0.5)
print('X_train.shape', X_train.shape)
print('X_test.shape', X_test.shape)
print('y_train.shape', y_train.shape)
print('y_test.shape', y_test.shape)
print('y_test', y_test)
# plot corrupted images
ncols = 4
fig, ax = plt.subplots(nrows=1, ncols=ncols, figsize=[15, 6.5])
for j in np.arange(ncols):
id = np.random.choice(np.arange(X_train.shape[0]))
ax[j].imshow(X_train[id,:].reshape(28,28))
plt.savefig('MNIST_ex_corrupted1.pdf', bbox_inches='tight')
def list2onehot(y, list_classes):
"""
y = list of class lables of length n
output = n x k array, i th row = one-hot encoding of y[i] (e.g., [0,0,1,0,0])
"""
Y = np.zeros(shape = [len(y), len(list_classes)], dtype=int)
for i in np.arange(Y.shape[0]):
for j in np.arange(len(list_classes)):
if y[i] == list_classes[j]:
Y[i,j] = 1
return Y
def sample_multiclass_MNIST(list_digits=['0','1', '2'], full_MNIST=None):
# get train and test set from MNIST of given digits
# e.g., list_digits = ['0', '1', '2']
if full_MNIST is not None:
X, y = full_MNIST
else:
X, y = fetch_openml('mnist_784', version=1, return_X_y=True)
X = X / 255.
Y = list2onehot(y.tolist(), list_digits)
idx = [i for i in np.arange(len(y)) if y[i] in list_digits] # list of indices where the label y is in list_digits
X01 = X[idx,:]
y01 = Y[idx,:]
X_train = []
X_test = []
y_test = [] # list of one-hot encodings (indicator vectors) of each label
y_train = [] # list of one-hot encodings (indicator vectors) of each label
for i in np.arange(X01.shape[0]):
# for each example i, make it into train set with probabiliy 0.8 and into test set otherwise
U = np.random.rand() # Uniform([0,1]) variable
if U<0.8:
X_train.append(X01[i,:])
y_train.append(y01[i,:].copy())
else:
X_test.append(X01[i,:])
y_test.append(y01[i,:].copy())
X_train = np.asarray(X_train)
X_test = np.asarray(X_test)
y_train = np.asarray(y_train)
y_test = np.asarray(y_test)
return X_train, X_test, y_train, y_test
# test
X_train, X_test, y_train, y_test = sample_multiclass_MNIST(list_digits=['0','1', '2'], full_MNIST=[X, y])
print('X_train.shape', X_train.shape)
print('X_test.shape', X_test.shape)
print('y_train.shape', y_train.shape)
print('y_test.shape', y_test.shape)
print('y_test', y_test)
```
## Logistic Regression
```
# sigmoid and logit function
def sigmoid(x):
return np.exp(x)/(1+np.exp(x))
# plot sigmoid function
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=[10,3])
x = np.linspace(-7, 7, 100)
ax.plot(x, sigmoid(x), color='blue', label="$y=\sigma(x)=\exp(x)/(1+\exp(x))$")
plt.axhline(y=1, color='g', linestyle='--')
plt.axvline(x=0, color='g', linestyle='--')
ax.legend()
plt.savefig('sigmoid_ex.pdf', bbox_inches='tight')
def fit_LR_GD(Y, H, W0=None, sub_iter=100, stopping_diff=0.01):
'''
Convex optimization algorithm for Logistic Regression using Gradient Descent
Y = (n x 1), H = (p x n) (\Phi in lecture note), W = (p x 1)
Logistic Regression: Y ~ Bernoulli(Q), Q = sigmoid(H.T @ W)
MLE -->
Find \hat{W} = argmin_W ( sum_j ( log(1+exp(H_j.T @ W) ) - Y.T @ H.T @ W ) )
'''
if W0 is None:
W0 = np.random.rand(H.shape[0],1) #If initial coefficients W0 is None, randomly initialize
W1 = W0.copy()
i = 0
grad = np.ones(W0.shape)
while (i < sub_iter) and (np.linalg.norm(grad) > stopping_diff):
Q = 1/(1+np.exp(-H.T @ W1)) # probability matrix, same shape as Y
# grad = H @ (Q - Y).T + alpha * np.ones(W0.shape[1])
grad = H @ (Q - Y)
W1 = W1 - (np.log(i+1) / (((i + 1) ** (0.5)))) * grad
i = i + 1
print('iter %i, grad_norm %f' %(i, np.linalg.norm(grad)))
return W1
def fit_LR_NR(Y, H, W0=None, sub_iter=100, stopping_diff=0.01):
'''
Convex optimization algorithm for Logistic Regression using Newton-Ralphson algorithm.
Y = (n x 1), H = (p x n) (\Phi in lecture note), W = (p x 1)
Logistic Regression: Y ~ Bernoulli(Q), Q = sigmoid(H.T @ W)
MLE -->
Find \hat{W} = argmin_W ( sum_j ( log(1+exp(H_j.T @ W) ) - Y.T @ H.T @ W ) )
'''
### Implement by yourself.
# fit logistic regression using GD
X_train, X_test, y_train, y_test = sample_binary_MNIST(['0', '1'], full_MNIST = [X,y])
# Feature matrix of size (p x n) = (feature dim x samples)
H_train = np.vstack((np.ones(X_train.shape[0]), X_train.T)) # add first row of 1's for bias features
W = fit_LR_GD(Y=y_train, H=H_train/400)
plt.imshow(W[1:,:].reshape(28,28))
# plot fitted logistic regression curve
digit_list_list = [['0','1'],['0','7'],['2','3'],['2', '8']] # list of list of two digits
# fit LR for each cases
W_array = []
for i in np.arange(len(digit_list_list)):
L = digit_list_list[i]
X_train, X_test, y_train, y_test = sample_binary_MNIST(list_digits=L, full_MNIST = [X,y])
H_train = np.vstack((np.ones(X_train.shape[0]), X_train.T)) # add first row of 1's for bias features
W = fit_LR_GD(Y=y_train, H=H_train)
W = fit_LR_GD(Y=y_train, H=H_train)
W_array.append(W.copy())
W_array = np.asarray(W_array)
# make plot
fig, ax = plt.subplots(nrows=1, ncols=len(digit_list_list), figsize=[16, 4])
for i in np.arange(len(digit_list_list)):
L = digit_list_list[i]
W = W_array[i]
im = ax[i].imshow(W[1:,:].reshape(28,28), vmin=np.min(W_array), vmax=np.max(W_array))
ax[i].title.set_text("LR coeff. for %s vs. %s" % (L[0], L[1]))
# ax[i].legend()
fig.subplots_adjust(right=0.9)
cbar_ax = fig.add_axes([0.92, 0.15, 0.01, 0.7])
fig.colorbar(im, cax=cbar_ax)
plt.savefig('LR_MNIST_training_ex.pdf', bbox_inches='tight')
def compute_accuracy_metrics(Y_test, P_pred, use_opt_threshold=False, verbose=False):
# y_test = binary label
# P_pred = predicted probability for y_test
# compuate various binary classification accuracy metrics
fpr, tpr, thresholds = metrics.roc_curve(Y_test, P_pred, pos_label=None)
mythre = thresholds[np.argmax(tpr - fpr)]
myauc = metrics.auc(fpr, tpr)
# print('!!! auc', myauc)
# Compute classification statistics
threshold = 0.5
if use_opt_threshold:
threshold = mythre
Y_pred = P_pred.copy()
Y_pred[Y_pred < threshold] = 0
Y_pred[Y_pred >= threshold] = 1
mcm = confusion_matrix(Y_test, Y_pred)
tn = mcm[0, 0]
tp = mcm[1, 1]
fn = mcm[1, 0]
fp = mcm[0, 1]
accuracy = (tp + tn) / (tp + tn + fp + fn)
sensitivity = tn / (tn + fp)
specificity = tp / (tp + fn)
precision = tp / (tp + fp)
fall_out = fp / (fp + tn)
miss_rate = fn / (fn + tp)
# Save results
results_dict = {}
results_dict.update({'Y_test': Y_test})
results_dict.update({'Y_pred': Y_pred})
results_dict.update({'AUC': myauc})
results_dict.update({'Opt_threshold': mythre})
results_dict.update({'Accuracy': accuracy})
results_dict.update({'Sensitivity': sensitivity})
results_dict.update({'Specificity': specificity})
results_dict.update({'Precision': precision})
results_dict.update({'Fall_out': fall_out})
results_dict.update({'Miss_rate': miss_rate})
if verbose:
for key in [key for key in results_dict.keys()]:
print('% s ===> %.3f' % (key, results_dict.get(key)))
return results_dict
# fit logistic regression using GD and compute binary classification accuracies
# Get train and test data
digits_list = ['4', '7']
X_train, X_test, y_train, y_test = sample_binary_MNIST(digits_list, full_MNIST = [X,y])
# Feature matrix of size (p x n) = (feature dim x samples)
list_train_size = [1,10, 30, 100]
# train the regression coefficients for all cases
W_list = []
results_list = []
for i in np.arange(len(list_train_size)):
size = list_train_size[i]
idx = np.random.choice(np.arange(len(y_train)), size)
X_train0 = X_train[idx, :]
y_train0 = y_train[idx]
# Train the logistic regression model
H_train0 = np.vstack((np.ones(X_train0.shape[0]), X_train0.T)) # add first row of 1's for bias features
W = fit_LR_GD(Y=y_train0, H=H_train0)
W_list.append(W.copy()) # make sure use copied version of W since the same name is overrided in the loop
# Get predicted probabilities
H_test = np.vstack((np.ones(X_test.shape[0]), X_test.T))
Q = 1 / (1 + np.exp(-H_test.T @ W)) # predicted probabilities for y_test
# Compute binary classification accuracies
results_dict = compute_accuracy_metrics(Y_test=y_test, P_pred = Q)
results_dict.update({'train size':X_train0.shape[0]}) # add the train data size to the results dictionary
results_list.append(results_dict.copy())
# Print out the results
"""
keys_list = [i for i in results_dict.keys()]
for key in keys_list:
if key not in ['Y_test', 'Y_pred']:
print('%s = %f' % (key, results_dict.get(key)))
"""
# make plot
fig, ax = plt.subplots(nrows=1, ncols=len(list_train_size), figsize=[16, 4])
for i in np.arange(len(list_train_size)):
result_dict = results_list[i]
W = W_list[i][1:,:]
im = ax[i].imshow(W.copy().reshape(28,28), vmin=np.min(W_list), vmax=np.max(W_list))
subtitle = ""
keys_list = [i for i in results_list[i].keys()]
for key in keys_list:
if key not in ['Y_test', 'Y_pred', 'AUC', 'Opt_threshold']:
subtitle += "\n" + str(key) + " = " + str(np.round(results_list[i].get(key),3))
# print('%s = %f' % (key, results_list[i].get(key)))
ax[i].set_title('Opt. regression coeff.', fontsize=13)
ax[i].set_xlabel(subtitle, fontsize=20)
fig.subplots_adjust(right=0.9)
fig.suptitle("MNIST Binary Classification by LR for %s vs. %s" % (digits_list[0], digits_list[1]), fontsize=20, y=1.05)
cbar_ax = fig.add_axes([0.92, 0.15, 0.01, 0.7])
fig.colorbar(im, cax=cbar_ax)
plt.savefig('LR_MNIST_test_ex1.pdf', bbox_inches='tight')
```
## Multiclass Logistic Regression
```
def fit_MLR_GD(Y, H, W0=None, sub_iter=100, stopping_diff=0.01):
'''
Convex optimization algorithm for Multiclass Logistic Regression using Gradient Descent
Y = (n x k), H = (p x n) (\Phi in lecture note), W = (p x k)
Multiclass Logistic Regression: Y ~ vector of discrete RVs with PMF = sigmoid(H.T @ W)
MLE -->
Find \hat{W} = argmin_W ( sum_j ( log(1+exp(H_j.T @ W) ) - Y.T @ H.T @ W ) )
'''
k = Y.shape[1] # number of classes
if W0 is None:
W0 = np.random.rand(H.shape[0],k) #If initial coefficients W0 is None, randomly initialize
W1 = W0.copy()
i = 0
grad = np.ones(W0.shape)
while (i < sub_iter) and (np.linalg.norm(grad) > stopping_diff):
Q = 1/(1+np.exp(-H.T @ W1)) # probability matrix, same shape as Y
# grad = H @ (Q - Y).T + alpha * np.ones(W0.shape[1])
grad = H @ (Q - Y)
W1 = W1 - (np.log(i+1) / (((i + 1) ** (0.5)))) * grad
i = i + 1
# print('iter %i, grad_norm %f' %(i, np.linalg.norm(grad)))
return W1
def custom_softmax(a):
"""
given an array a = [a_1, .. a_k], compute the softmax distribution p = [p_1, .. , p_k] where p_i \propto exp(a_i)
"""
a1 = a - np.max(a)
p = np.exp(a1)
if type(a) is list:
p = p/np.sum(p)
else:
row_sum = np.sum(p, axis=1)
p = p/row_sum[:, np.newaxis]
return p
print(np.sum(custom_softmax([1,20,30,50])))
a= np.ones((2,3))
print(softmax(a))
def multiclass_accuracy_metrics(Y_test, P_pred, class_labels=None, use_opt_threshold=False):
# y_test = multiclass one-hot encoding labels
# Q = predicted probability for y_test
# compuate various classification accuracy metrics
results_dict = {}
y_test = []
y_pred = []
for i in np.arange(Y_test.shape[0]):
for j in np.arange(Y_test.shape[1]):
if Y_test[i,j] == 1:
y_test.append(j)
if P_pred[i,j] == np.max(P_pred[i,:]):
# print('!!!', np.where(P_pred[i,:]==np.max(P_pred[i,:])))
y_pred.append(j)
confusion_mx = metrics.confusion_matrix(y_test, y_pred)
results_dict.update({'confusion_mx':confusion_mx})
results_dict.update({'Accuracy':np.trace(confusion_mx)/np.sum(np.sum(confusion_mx))})
print('!!! confusion_mx', confusion_mx)
print('!!! Accuracy', results_dict.get('Accuracy'))
return results_dict
# fit multiclass logistic regression using GD
list_digits=['0', '1', '2']
X_train, X_test, y_train, y_test = sample_multiclass_MNIST(list_digits=list_digits, full_MNIST = [X,y])
# Feature matrix of size (p x n) = (feature dim x samples)
H_train = np.vstack((np.ones(X_train.shape[0]), X_train.T)) # add first row of 1's for bias features
W = fit_MLR_GD(Y=y_train, H=H_train)
print('!! W.shape', W.shape)
# Get predicted probabilities
H_test = np.vstack((np.ones(X_test.shape[0]), X_test.T))
Q = softmax(H_test.T @ W.copy()) # predicted probabilities for y_test # Uses sklearn's softmax for numerical stability
print('!!! y_test.shape', y_test.shape)
print('!!! Q.shape', Q.shape)
results_dict = multiclass_accuracy_metrics(Y_test=y_test, P_pred=Q)
confusion_mx = results_dict.get('results_dict')
# make plot
fig, ax = plt.subplots(nrows=1, ncols=len(list_digits), figsize=[12, 4])
for i in np.arange(len(list_digits)):
L = list_digits[i]
im = ax[i].imshow(W[1:,i].reshape(28,28), vmin=np.min(W), vmax=np.max(W))
ax[i].title.set_text("MLR coeff. for %s" % L )
# ax[i].legend()
# if i == len(list_digits) - 1:
cbar_ax = fig.add_axes([0.92, 0.15, 0.01, 0.7])
fig.colorbar(im, cax=cbar_ax)
plt.savefig('MLR_MNIST_ex1.pdf', bbox_inches='tight')
# fit multiclass logistic regression using GD and compute multiclass classification accuracies
# Get train and test data
digits_list = ['0', '1', '2', '3', '4']
X_train, X_test, y_train, y_test = sample_multiclass_MNIST(digits_list, full_MNIST = [X,y])
# Feature matrix of size (p x n) = (feature dim x samples)
list_train_size = [1,10, 30, 100]
# train the regression coefficients for all cases
W_list = []
results_list = []
for i in np.arange(len(list_train_size)):
size = list_train_size[i]
idx = np.random.choice(np.arange(len(y_train)), size)
X_train0 = X_train[idx, :]
y_train0 = y_train[idx, :]
# Train the multiclass logistic regression model
H_train0 = np.vstack((np.ones(X_train0.shape[0]), X_train0.T)) # add first row of 1's for bias features
W = fit_MLR_GD(Y=y_train0, H=H_train0)
W_list.append(W.copy()) # make sure use copied version of W since the same name is overrided in the loop
# Get predicted probabilities
H_test = np.vstack((np.ones(X_test.shape[0]), X_test.T))
Q = softmax(H_test.T @ W.copy()) # predicted probabilities for y_test # Uses sklearn's softmax for numerical stability
results_dict = multiclass_accuracy_metrics(Y_test=y_test, P_pred=Q)
results_dict.update({'train size':X_train0.shape[0]}) # add the train data size to the results dictionary
results_list.append(results_dict.copy())
# make plot
fig, ax = plt.subplots(nrows=len(list_train_size), ncols=len(digits_list)+1, figsize=[15, 10])
for i in np.arange(len(list_train_size)):
for j in np.arange(len(digits_list)+1):
if j < len(digits_list):
L = digits_list[j]
W = W_list[i]
im = ax[i,j].imshow(W[1:,j].reshape(28,28), vmin=np.min(W), vmax=np.max(W))
ax[i,j].title.set_text("MLR coeff. for %s" % L )
if j == 0:
ax[i,j].set_ylabel("train size = %i" % results_list[i].get("train size"), fontsize=13)
divider = make_axes_locatable(ax[i,j])
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im, cax=cax)
else:
confusion_mx = results_list[i].get("confusion_mx")
im_confusion = ax[i,j].matshow(confusion_mx)
# ax[i,j].set_title("Confusion Matrix")
ax[i,j].set_xlabel("Confusion Matrix", fontsize=13)
# ax[i].legend()
# if i == len(list_digits) - 1:
divider = make_axes_locatable(ax[i,j])
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im_confusion, cax=cax)
plt.subplots_adjust(wspace=0.3, hspace=0.3)
plt.savefig('MLR_MNIST_test_ex2.pdf', bbox_inches='tight')
```
## Probit Regression
```
# probit function
from scipy.stats import norm
def probit(x):
return norm.cdf(x) # Yes, it is exactly the standard normal CDF.
# plot probit and sigmoid function
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=[10,3])
x = np.linspace(-7, 7, 100)
ax.plot(x, sigmoid(x), color='blue', label="$y=\sigma(x)=\exp(x)/(1+\exp(x))$")
ax.plot(x, probit(x), color='red', label="$y=\psi(x)=Probit(x)$")
plt.axhline(y=1, color='g', linestyle='--')
plt.axvline(x=0, color='g', linestyle='--')
ax.legend()
plt.savefig('probit_ex.pdf', bbox_inches='tight')
def fit_PR_GD(Y, H, W0=None, sub_iter=100, stopping_diff=0.01):
'''
Convex optimization algorithm for Probit Regression using Gradient Descent
Y = (n x 1), H = (p x n) (\Phi in lecture note), W = (p x 1)
Logistic Regression: Y ~ Bernoulli(Q), Q = Probit(H.T @ W)
'''
if W0 is None:
W0 = 1-2*np.random.rand(H.shape[0],1) #If initial coefficients W0 is None, randomly initialize from [-1,1]
W1 = W0.copy()
i = 0
grad = np.ones(W0.shape)
while (i < sub_iter) and (np.linalg.norm(grad) > stopping_diff):
Q = norm.pdf(H.T @ W1) * ( (1-Y)/norm.cdf(-H.T @ W1) - Y/norm.cdf(H.T @ W1) )
grad = H @ Q
W1 = W1 - (np.log(i+1) / (((i + 1) ** (0.5)))) * grad
i = i + 1
# print('iter %i, grad_norm %f' %(i, np.linalg.norm(grad)))
return W1
# plot fitted probit regression curve
digit_list_list = [['0','1'],['0','7'],['2','3'],['2', '8']] # list of list of two digits
# fit LR for each cases
W_array = []
for i in np.arange(len(digit_list_list)):
L = digit_list_list[i]
X_train, X_test, y_train, y_test = sample_binary_MNIST(list_digits=L, full_MNIST = [X,y], noise_rate=0.5)
H_train = np.vstack((np.ones(X_train.shape[0]), X_train.T)) # add first row of 1's for bias features
W = fit_PR_GD(Y=y_train, H=H_train/1000)
W = fit_PR_GD(Y=y_train, H=H_train/1000)
W_array.append(W.copy())
W_array = np.asarray(W_array)
# make plot
fig, ax = plt.subplots(nrows=1, ncols=len(digit_list_list), figsize=[16, 4])
for i in np.arange(len(digit_list_list)):
L = digit_list_list[i]
W = W_array[i]
im = ax[i].imshow(W[1:,:].reshape(28,28), vmin=np.min(W_array), vmax=np.max(W_array))
ax[i].title.set_text("LR coeff. for %s vs. %s" % (L[0], L[1]))
# ax[i].legend()
fig.subplots_adjust(right=0.9)
cbar_ax = fig.add_axes([0.92, 0.15, 0.01, 0.7])
fig.colorbar(im, cax=cbar_ax)
plt.savefig('PR_MNIST_training_ex.pdf', bbox_inches='tight')
# fit probit regression using GD and compute binary classification accuracies
# Get train and test data
digits_list = ['4', '7']
X_train, X_test, y_train, y_test = sample_binary_MNIST(digits_list, full_MNIST = [X,y], noise_rate=0.5)
# Feature matrix of size (p x n) = (feature dim x samples)
list_train_size = [1,10, 30, 100]
# train the regression coefficients for all cases
W_list = []
results_list = []
for i in np.arange(len(list_train_size)):
size = list_train_size[i]
idx = np.random.choice(np.arange(len(y_train)), size)
X_train0 = X_train[idx, :]
y_train0 = y_train[idx]
# Train the logistic regression model
H_train0 = np.vstack((np.ones(X_train0.shape[0]), X_train0.T)) # add first row of 1's for bias features
W = fit_PR_GD(Y=y_train0, H=H_train0/100) # reduce the scale of H for numerical stability
W_list.append(W.copy()) # make sure use copied version of W since the same name is overrided in the loop
# Get predicted probabilities
H_test = np.vstack((np.ones(X_test.shape[0]), X_test.T))
Q = 1 / (1 + np.exp(-H_test.T @ W)) # predicted probabilities for y_test
# Compute binary classification accuracies
results_dict = compute_accuracy_metrics(Y_test=y_test, P_pred = Q)
results_dict.update({'train size':X_train0.shape[0]}) # add the train data size to the results dictionary
results_list.append(results_dict.copy())
# Print out the results
"""
keys_list = [i for i in results_dict.keys()]
for key in keys_list:
if key not in ['Y_test', 'Y_pred']:
print('%s = %f' % (key, results_dict.get(key)))
"""
# make plot
fig, ax = plt.subplots(nrows=1, ncols=len(list_train_size), figsize=[16, 4])
for i in np.arange(len(list_train_size)):
result_dict = results_list[i]
W = W_list[i][1:,:]
im = ax[i].imshow(W.copy().reshape(28,28), vmin=np.min(W_list), vmax=np.max(W_list))
subtitle = ""
keys_list = [i for i in results_list[i].keys()]
for key in keys_list:
if key not in ['Y_test', 'Y_pred', 'AUC', 'Opt_threshold']:
subtitle += "\n" + str(key) + " = " + str(np.round(results_list[i].get(key),3))
# print('%s = %f' % (key, results_list[i].get(key)))
ax[i].set_title('Opt. regression coeff.', fontsize=13)
ax[i].set_xlabel(subtitle, fontsize=20)
fig.subplots_adjust(right=0.9)
fig.suptitle("MNIST Binary Classification by LR for %s vs. %s" % (digits_list[0], digits_list[1]), fontsize=20, y=1.05)
cbar_ax = fig.add_axes([0.92, 0.15, 0.01, 0.7])
fig.colorbar(im, cax=cbar_ax)
plt.savefig('PR_MNIST_test_ex1.pdf', bbox_inches='tight')
```
| github_jupyter |
```
import sys, os, time
from solc import compile_source, compile_files, link_code
from ethjsonrpc import EthJsonRpc
print("Using environment in "+sys.prefix)
print("Python version "+sys.version)
# Initiate connection to ethereum node
# Requires a node running with an RPC connection available at port 8545
c = EthJsonRpc('127.0.0.1', 8545)
print(c.web3_clientVersion())
print("Block number %s"%c.eth_blockNumber())
source = """pragma solidity ^0.4.2;
contract mortal {
/* Define variable owner of the type address*/
address owner;
/* this function is executed at initialization and sets the owner of the contract */
function mortal() { owner = msg.sender; }
/* Function to recover the funds on the contract */
function kill() { if (msg.sender == owner) selfdestruct(owner); }
}
contract greeter is mortal {
/* define variable greeting of the type string */
string greeting;
address greetername;
/* this runs when the contract is executed */
function greeter(string _greeting) public {
greeting = _greeting;
greetername = msg.sender;
}
/* main function */
function greetme() constant returns (string) {
return greeting;
}
function originator() constant returns (address ret) {
return greetername;
}
function greettwo() constant returns (string) {
return greeting;
}
}"""
```
- *i* participants/nodes
- *t* time periods
- *k* ADMM iteration
- *z* global ADMM estimate at iteration *k*
- *x_i* local variable estimates for node *i*
Pseudocode:
- Variables:
- Array of *i* addresses (permanent)
- Current iteration (int)
- ADMM variable estimates from each participant: *i* entries, each holding ~3n variables
- Array of people for whom we are still expecting an update this iteration
- Schedule - array, *t* time steps by *i* nodes
- Functions:
- Initialize (list of addresses):
- Initialize the global variable estimates
- Set the schedule to none
- Set the list of people we want to be everyone
- Set the iteration to be 1
- Set the tolerance
- Recieve update (variable estimate)
- Check that iteration is current- if not throw
- Remove message sender from waiting list
- Store estimate in array
- If waiting list is now empty, call updater
- Updater
- Compute the average value
- Compute the change in average value
- If the deviation is above the tolerance
- Increment the current iteration
- Else,
- Save the schedule
- Set the iteration to throw an error
- Get schedule:
- If the schedule is empty, throw
- If we have a schedule, return the schedule for the message sender
Workflow:
- Device submits variable estimate
- Once all devices have submitted estimates, global variable is updated
- Devices poll contract until they see the the iteration number increment
- Process loops until global variable change is less than the tolerance
- Schedule is saved
- Device
Private:
- Check if still waiting:
- index goes until whitelist length
-
- Loop through whitelist
- if any of those are still waiting, return true
```
source = """pragma solidity ^0.4.2;
contract aggregator {
address owner;
uint8 public iteration;
address[] public whitelist;
mapping (address => bool) public waiting;
/* CONSTRUCTOR */
function aggregator (address[] _whitelist) public{
whitelist = _whitelist;
iteration = 1;
resetWaiting(); // Set the waiting flag to 1 for everybody
}
function stillWaiting () returns (bool) {
for (uint8 i=0; i<whitelist.length; i++){
if (waiting[whitelist[i]]){ return true; }
}
return false;
}
function resetWaiting () {
// Reset the flag for each address
for(uint8 i=0; i<whitelist.length; i++){
waiting[ whitelist[i] ] = true;
}
}
}
"""
compiled['<stdin>:aggregator']['abi']
# Basic contract compiling process.
# Requires that the creating account be unlocked.
# Note that by default, the account will only be unlocked for 5 minutes (300s).
# Specify a different duration in the geth personal.unlockAccount('acct','passwd',300) call, or 0 for no limit
compiled = compile_source(source)
compiledCode = compiled['<stdin>:aggregator']['bin']
compiledCode = '0x'+compiledCode # This is a hack which makes the system work
addressList = [x[2:] for x in c.eth_accounts()]
contractTx = c.create_contract(c.eth_coinbase(), compiledCode, gas=3000000,sig='aggregator(address[])',args=[addressList])
# contractTx = c.create_contract(c.eth_coinbase(), compiledCode, gas=3000000,sig='greeter(string)',args=['Hello World!'])
print("Contract transaction id is "+contractTx)
print("Waiting for the contract to be mined into the blockchain...")
while c.eth_getTransactionReceipt(contractTx) is None:
time.sleep(1)
contractAddr = c.get_contract_address(contractTx)
print("Contract address is "+contractAddr)
c.call('0x3f211b2256bc7b64f365cbc7bbff4ae77e22a151','indexer()',[],['int8'])
c.call('0x3f211b2256bc7b64f365cbc7bbff4ae77e22a151','names(uint256)',[2],['uint256'])
```
# Other Stuff
Stackoverflow question at https://stackoverflow.com/questions/44373531/contract-method-not-returning-the-value-while-using-ethjsonrpc-and-pyethapp
```
source = """pragma solidity ^0.4.2;
contract Example {
// Device Registry
mapping (uint => string) public _registry;
uint nIndex= 0;
function set_s(string new_s) {
_registry[nIndex] = new_s;
nIndex = nIndex + 1;
}
function get_s(uint number) returns (string) {
return _registry[number];
}
}
"""
compiled = compile_source(source)
# compiled = compile_files(['Solidity/ethjsonrpc_tutorial.sol']) #Note: Use this to compile from a file
compiledCode = compiled['Example']['bin']
compiledCode = '0x'+compiledCode # This is a hack which makes the system work
# Put the contract in the pool for mining, with a gas reward for processing
contractTx = c.create_contract(c.eth_coinbase(), compiledCode, gas=3000000)
print("Contract transaction id is "+contractTx)
print("Waiting for the contract to be mined into the blockchain...")
while c.eth_getTransactionReceipt(contractTx) is None:
time.sleep(1)
contractAddr = c.get_contract_address(contractTx)
print("Contract address is "+contractAddr)
tx = c.call_with_transaction(c.eth_coinbase(), contractAddr, 'set_s(string)', ['Dinesh'])
while c.eth_getTransactionReceipt(tx) is None:
time.sleep(1)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/harry418/EmotionRecog/blob/master/training/training_4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from google.colab import drive
drive.mount('/content/gdrive',force_remount=True)
```
# Imported importanat libraries
```
# baseline model with dropout and data augmentation on the cifar10 dataset
import sys
import tensorflow
from matplotlib import pyplot
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential,Model
from tensorflow.keras.layers import Conv2D,MaxPooling2D,Dense,Flatten,Dropout,BatchNormalization,AveragePooling2D
from tensorflow.keras.optimizers import SGD,Adam
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from sklearn.model_selection import train_test_split
from tensorflow.keras.preprocessing.image import img_to_array
from tensorflow.keras.preprocessing.image import load_img
from imutils import paths
import matplotlib.pyplot as plt
import numpy as np
import random
import os
from sklearn.preprocessing import LabelEncoder
from tensorflow.keras.layers import Input
```
# load preprocessed data
```
# storing preprocessed images and labels for further use
data = np.load('/content/gdrive/My Drive/emotion_recog/data.npy')
labels_value = np.load('/content/gdrive/My Drive/emotion_recog/labels_value.npy')
```
# train and test splitting with sklearn
```
trainX, testX,trainY, testY = train_test_split(data, labels_value,test_size=0.2, random_state=42,shuffle = True)
```
# Training and plotting accuracy and loss
```
# plot diagnostic learning curves
import matplotlib.pyplot as plt
def summarize_diagnostics(hist):
plt.plot(hist.history["accuracy"])
plt.plot(hist.history['val_accuracy'])
plt.plot(hist.history['loss'])
plt.plot(hist.history['val_loss'])
plt.title("model accuracy")
plt.ylabel("Accuracy")
plt.xlabel("Epoch")
plt.legend(["Accuracy","Validation Accuracy","loss","Validation Loss"])
plt.show()
from keras.applications.vgg16 import VGG16
from keras.preprocessing import image
from keras.layers import GlobalAveragePooling2D
inp = Input(shape = (224,224,3))
model_mobile = VGG16(input_shape=(224,224,3), include_top=False, weights='imagenet')
x1 = model_mobile(inp)
x2 = GlobalAveragePooling2D()(x1)
#x3 = Dense(128,activation='relu')(x2)
out = Dense(6, activation='softmax')(x2)
INIT_LR = 1e-4
print("[INFO] compiling model...")
opt = Adam(lr=INIT_LR, decay=INIT_LR /100)
model = Model(inputs = inp, outputs = out)
model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
model.summary()
# batch size and epochs
EPOCHS = 100
BS = 32
datagen = ImageDataGenerator(width_shift_range=0.1, height_shift_range=0.1, horizontal_flip=True,zoom_range=0.1)
# prepare iterator
it_train = datagen.flow(trainX, trainY, batch_size=BS)
# fit model
steps = int(trainX.shape[0] / BS)
hist = model.fit_generator(it_train, steps_per_epoch=steps, epochs=EPOCHS, validation_data=(testX, testY), verbose=1)
# evaluate model
_, acc = model.evaluate(testX, testY, verbose=1)
print('> %.3f' % (acc * 100.0))
summarize_diagnostics(hist)
```
# Confusion matrix and classification report
```
from sklearn.metrics import confusion_matrix,classification_report
y_pred = model.predict(testX)
y_p = np.argmax(y_pred,axis=1)
y_true = np.argmax(testY,axis=1)
print(confusion_matrix(y_true,y_p))
print('Classification report')
print(classification_report(y_true,y_p))
```
# CNN + SVM
```
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
# extracting features using Transfer Learning
model_new = Model(inputs = model.input,outputs = model.get_layer('global_average_pooling2d_3').output)
train_new = sc.fit_transform(model_new.predict(trainX))
test_new = sc.fit_transform(model_new.predict(testX))
from sklearn.svm import SVC
svm = SVC(kernel='rbf')
svm.fit(train_new,np.argmax(trainY,axis=1))
sc1 = svm.score(train_new,np.argmax(trainY,axis=1))
sc2 = svm.score(test_new,np.argmax(testY,axis=1))
print('training accuracy of svm is : ',sc1)
print('testing accuracy of svm is : ',sc2)
```
# CNN + XGBOOST
```
from xgboost import XGBClassifier
xg = XGBClassifier()
xg.fit(train_new,np.argmax(trainY,axis=1))
sc3 = xg.score(train_new,np.argmax(trainY,axis=1))
sc4 = xg.score(test_new,np.argmax(testY,axis=1))
print('training accuracy of xgboost is : ',sc3)
print('testing accuracy of xgboost is : ',sc4)
```
| github_jupyter |
```
def displayList (myList):
print(myList)
myList = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
displayList(myList)
input('Please enter a value: ')
result = input('Please enter a number: ')
type(result)
int(result)
result = int(input("Please enter a number: "))
type(result)
result = int(input("Please enter a number: "))
def userChoice():
'''
User inputs a number between 0 to 10 and we reutrn
Integer number of it.
'''
choice = input("Please input a number (0 - 10): ")
return int(choice)
userChoice()
result = userChoice()
type(result)
someInput = '10'
someInput.isdigit()
def userChoice():
choice = 'wrong'
while choice.isdigit() == False:
choice = input("Choose a Number: ")
return int(choice)
userChoice
userChoice()
def userChoice():
choice = "wrong"
while choice.isdigit() == False:
choice = input("Chose a Number:")
if choice.isdigit() == False:
print("Sorry, but you did not enter an enter. Please try again")
return int(choice)
userChoice()
from IPython.display import clear_output
clear_output()
def userChoice():
choice = "wrong"
while choice.isdigit() == False:
choice = input("Choice a Number: ")
if choice.isdigit() == False:
clear_output()
print("Sorry, you did not enter an integer, Try again !!")
return int(choice)
userChoice()
result = "wrong value"
acceptable_values = ['0', '1', '2']
result in acceptable_values
result not in acceptable_values
from IPython.display import clear_output
clear_output
def userChoice():
choice = "Hey"
while choice not in ['0', '1', '2']:
choice = input("Choose one of these numbers (0, 1, 2): ")
if choice not in ['0', '1', '2']:
clear_output()
print("Sorry, but you did not choose a value in the correct range (0, 1, 2)")
return int(choice)
userChoice()
def userChoice():
choice = "WRONG"
within_range = False
while choice.isdigit() == False or within_range == False:
choice = input("Please enter a number (0 - 10): ")
if choice.isdigit() == False:
print("Sorry that is not a digit!")
if choice.isdigit() == True:
if int(choice) in range(0, 10):
within_range = True
else:
within_range = False
return int(choice)
userChoice()
clear()
gameList = [0, 1, 2]
def displayGame(gameList):
print("Here is the current list")
print(gameList)
displayGame(gameList)
def positionChoice():
choice = "Wrong"
while choice not in ['0', '1', '2']:
choice = input("Pick a position to replace (0, 1, 2): ")
if choice not in ['0', '1', '2']:
clear_output()
print("Sorry, but you did not choose a valid position (0, 1, 2)")
return int(choice)
def replacementChoice(gameList, position):
userPlacement = input("Type a string to place the position")
gameList[position] = userPlacement
return gameList
def gameonChoice():
choice = "wrong"
while choice not in ['Y', 'N']:
choice = input("Would you like to keep playing? Y or N")
if choice not in ['Y', 'N']:
clear_output()
print("Sorry, I didn't understand. Please make sure to choose Y or N")
if choice == "Y":
return True
else:
return False
gameOn = True
gameList = [0, 1, 2]
while gameOn:
clear_output()
displayGame(gameList)
position = positionChoice()
gameList = replacementChoice(gameList, position)
clear_output()
displayGame(gameList)
gameOn = gameonChoice()
```
| github_jupyter |
Adapted from https://pytorch.org/tutorials/intermediate/torchvision_tutorial.html
# Prepare Training Script
In this notebook, we create the training script of the Mask R-CNN model that will be tuned. We first define the custom dataset class and the model that finetunes a pre-trained Mask R-CNN for our dataset. The training script is created by appending some notebook cells in turn so it is essential that you run the notebook's cells in order for the script to run correctly.
## Define dataset class and transformations
```
%%writefile scripts/XMLDataset.py
import os
import xml.etree.ElementTree as ET
import torch
import transforms as T
from PIL import Image
class BuildDataset(torch.utils.data.Dataset):
def __init__(self, root, transforms=None):
self.root = root
self.transforms = transforms
# load all image files
self.imgs = list(sorted(os.listdir(os.path.join(root, "Data/JPEGImages"))))
def __getitem__(self, idx):
img_path = os.path.join(self.root, "Data/JPEGImages", self.imgs[idx])
xml_path = os.path.join(
self.root, "Data/Annotations", "{}.xml".format(self.imgs[idx].strip(".jpg"))
)
img = Image.open(img_path).convert("RGB")
# parse XML annotation
tree = ET.parse(xml_path)
t_root = tree.getroot()
# get bounding box coordinates
boxes = []
for obj in t_root.findall("object"):
bnd_box = obj.find("bndbox")
xmin = float(bnd_box.find("xmin").text)
xmax = float(bnd_box.find("xmax").text)
ymin = float(bnd_box.find("ymin").text)
ymax = float(bnd_box.find("ymax").text)
boxes.append([xmin, ymin, xmax, ymax])
num_objs = len(boxes)
boxes = torch.as_tensor(boxes, dtype=torch.float32)
# there is only one class
labels = torch.ones((num_objs,), dtype=torch.int64)
image_id = torch.tensor([idx])
# area of the bounding box, used during evaluation with the COCO metric for small, medium and large boxes
area = (boxes[:, 3] - boxes[:, 1]) * (boxes[:, 2] - boxes[:, 0])
# suppose all instances are not crowd
iscrowd = torch.zeros((num_objs,), dtype=torch.int64)
target = {}
target["boxes"] = boxes
target["labels"] = labels
target["image_id"] = image_id
target["area"] = area
target["iscrowd"] = iscrowd
if self.transforms is not None:
img, target = self.transforms(img, target)
return img, target
def __len__(self):
return len(self.imgs)
def get_transform(train):
transforms = []
transforms.append(T.ToTensor())
if train:
transforms.append(T.RandomHorizontalFlip(0.5))
return T.Compose(transforms)
```
## Define model
```
%%writefile scripts/maskrcnn_model.py
import torchvision
from torchvision.models.detection.faster_rcnn import FastRCNNPredictor
from torchvision.models.detection.rpn import AnchorGenerator
from torchvision.models.detection.rpn import RPNHead
def get_model(
num_classes,
anchor_sizes,
anchor_aspect_ratios,
rpn_nms_threshold,
box_nms_threshold,
box_score_threshold,
num_box_detections,
):
# load pre-trained mask R-CNN model
model = torchvision.models.detection.maskrcnn_resnet50_fpn(
pretrained=True,
rpn_nms_thresh=rpn_nms_threshold,
box_nms_thresh=box_nms_threshold,
box_score_thresh=box_score_threshold,
box_detections_per_img=num_box_detections,
)
# get number of input features for the classifier
in_features = model.roi_heads.box_predictor.cls_score.in_features
# replace the pre-trained head with a new one
model.roi_heads.box_predictor = FastRCNNPredictor(in_features, num_classes)
anchor_sizes = tuple([float(i) for i in anchor_sizes.split(",")])
anchor_aspect_ratios = tuple([float(i) for i in anchor_aspect_ratios.split(",")])
# create an anchor_generator for the FPN which by default has 5 outputs
anchor_generator = AnchorGenerator(
sizes=tuple([anchor_sizes for _ in range(5)]),
aspect_ratios=tuple([anchor_aspect_ratios for _ in range(5)]),
)
model.rpn.anchor_generator = anchor_generator
# get number of input features for the RPN returned by FPN (256)
in_channels = model.backbone.out_channels
# replace the RPN head
model.rpn.head = RPNHead(
in_channels, anchor_generator.num_anchors_per_location()[0]
)
# turn off masks since dataset only has bounding boxes
model.roi_heads.mask_roi_pool = None
return model
```
## Define the training script and its arguments
We will use some of the below arguments as hyperparameters to tune the object detection model later. See following for all [arguments of MaskRCNN](https://github.com/pytorch/vision/blob/7716aba57e6e12a544c42136b274508955526163/torchvision/models/detection/mask_rcnn.py#L20).
```
%%writefile scripts/train.py
import os
import sys
sys.path.append("./cocoapi/PythonAPI/")
import torch
import argparse
import utils
from XMLDataset import BuildDataset, get_transform
from maskrcnn_model import get_model
from engine import train_one_epoch, evaluate
if __name__ == "__main__":
parser = argparse.ArgumentParser(description="PyTorch Object Detection Training")
parser.add_argument(
"--data_path", default="./Data/", help="the path to the dataset"
)
parser.add_argument("--batch_size", default=2, type=int)
parser.add_argument(
"--epochs", default=10, type=int, help="number of total epochs to run"
)
parser.add_argument(
"--workers", default=4, type=int, help="number of data loading workers"
)
parser.add_argument(
"--learning_rate", default=0.005, type=float, help="initial learning rate"
)
parser.add_argument("--momentum", default=0.9, type=float, help="momentum")
parser.add_argument(
"--weight_decay",
default=0.0005,
type=float,
help="weight decay (default: 1e-4)",
)
parser.add_argument(
"--lr_step_size", default=3, type=int, help="decrease lr every step-size epochs"
)
parser.add_argument(
"--lr_gamma",
default=0.1,
type=float,
help="decrease lr by a factor of lr-gamma",
)
parser.add_argument("--print_freq", default=10, type=int, help="print frequency")
parser.add_argument("--output_dir", default="outputs", help="path where to save")
parser.add_argument("--anchor_sizes", default="16", type=str, help="anchor sizes")
parser.add_argument(
"--anchor_aspect_ratios", default="1.0", type=str, help="anchor aspect ratios"
)
parser.add_argument(
"--rpn_nms_thresh",
default=0.7,
type=float,
help="NMS threshold used for postprocessing the RPN proposals",
)
parser.add_argument(
"--box_nms_thresh",
default=0.5,
type=float,
help="NMS threshold for the prediction head. Used during inference",
)
parser.add_argument(
"--box_score_thresh",
default=0.05,
type=float,
help="during inference only return proposals"
"with a classification score greater than box_score_thresh",
)
parser.add_argument(
"--box_detections_per_img",
default=100,
type=int,
help="maximum number of detections per image, for all classes",
)
args = parser.parse_args()
```
## Load data
```
%%writefile --append scripts/train.py
data_path = args.data_path
# use our dataset and defined transformations
dataset = BuildDataset(data_path, get_transform(train=True))
dataset_test = BuildDataset(data_path, get_transform(train=False))
# split the dataset in train and test set
indices = torch.randperm(len(dataset)).tolist()
dataset = torch.utils.data.Subset(dataset, indices[:-100])
dataset_test = torch.utils.data.Subset(dataset_test, indices[-100:])
batch_size = args.batch_size
workers = args.workers
# define training and validation data loaders
data_loader = torch.utils.data.DataLoader(
dataset,
batch_size=2,
shuffle=True,
num_workers=workers,
collate_fn=utils.collate_fn,
)
data_loader_test = torch.utils.data.DataLoader(
dataset_test,
batch_size=2,
shuffle=False,
num_workers=workers,
collate_fn=utils.collate_fn,
)
```
## Create model
```
%%writefile --append scripts/train.py
# our dataset has two classes only - background and out of stock
num_classes = 2
model = get_model(
num_classes,
args.anchor_sizes,
args.anchor_aspect_ratios,
args.rpn_nms_thresh,
args.box_nms_thresh,
args.box_score_thresh,
args.box_detections_per_img,
)
```
## Train model
```
%%writefile --append scripts/train.py
# train on the GPU or on the CPU, if a GPU is not available
device = torch.device("cuda") if torch.cuda.is_available() else torch.device("cpu")
# move model to the right device
model.to(device)
learning_rate = args.learning_rate
momentum = args.momentum
weight_decay = args.weight_decay
# construct an optimizer
params = [p for p in model.parameters() if p.requires_grad]
optimizer = torch.optim.SGD(
params, lr=learning_rate, momentum=momentum, weight_decay=weight_decay
)
lr_step_size = args.lr_step_size
lr_gamma = args.lr_gamma
# and a learning rate scheduler
lr_scheduler = torch.optim.lr_scheduler.StepLR(
optimizer, step_size=lr_step_size, gamma=lr_gamma
)
# number of training epochs
num_epochs = args.epochs
print_freq = args.print_freq
for epoch in range(num_epochs):
# train for one epoch, printing every 10 iterations
train_one_epoch(model, optimizer, data_loader, device, epoch, print_freq=print_freq)
# update the learning rate
lr_scheduler.step()
# evaluate on the test dataset after every epoch
evaluate(model, data_loader_test, device=device)
# save model
if not os.path.exists(args.output_dir):
os.makedirs(args.output_dir)
torch.save(model.state_dict(), os.path.join(args.output_dir, "model_latest.pth"))
print("That's it!")
```
In the next notebook, we [train the model locally and visualize its predictions](02_PytorchEstimatorLocalRun.ipynb).
| github_jupyter |
# A deep dive into our NLP solution
In this notebook we will see how does our model react in concrete situation, for that we will use only one PDF and some raw text.\
Feel free to play with our model ;)
First of all, let's import our packages:
- `os`: Various interfaces for the operating system
- `sys`: System specific parameters and fonctions
- `metaData`: Fetching informations from raw text and PDF
- `dataExtraction`: Work with pdf and raw data extraction
- `TrQuestions` : Work with transformer for QnA
- `TrSentymentAnalysis` : Work with transformer for Sentyment Analysis
- `Numpy` : Fundamental package for scientific computing with Python
- `pyplot` : Collection of functions that make matplotlib work like MATLAB
- `transformers` : Hugging face's transformer library
```
import os
import sys
import metaData
import dataExtraction
import TrQuestions
import TrSentymentAnalysis
import numpy as np
import matplotlib.pyplot as plt
from transformers import pipeline
```
**Now let's load and parse our report!**
- `dataExtraction.PDFToText(path)`: convert pdf to raw text
- `metaData.getInfo(data, from_text=bool)`: parse raw text to extract usefull informations
```
report = dataExtraction.PDFToText("exemple_report.pdf")
reportInfos = metaData.getInfo(report, from_text=True)
```
Let's play with report's data !\
First we will see what informations did we fetch for the part *Pupils' achievements*:
```
print(reportInfos[4])
```
We can see our data fetching worked pretty well, let's play with it by asking some questions.\
What about "What is the quality of pupils", pretty straight forward right?
- `pipeline('question-answering')`: load QnA Transformer model
- `nlp_qa(context=text, question=text)`: ask to the model the sentiment of the sentence
```
nlp_qa = pipeline('question-answering')
nlp_qa(context=reportInfos[4], question='What is the quality of pupils')
```
Impressive ! Our model succed to answer right ou question, but is a bit to specific.\
How does it handle a verry vague question: "Who ?"
```
nlp_qa(context=reportInfos[4], question='Who ?')
```
Well, it succed once again !\
Let's complexify it one more time by testing our own custom text.
**PS: Feel free to test with your own text/questions**
```
text = """
PoC is a Student Innovation Center, currently based at EPITECH, which aims to promote Innovation and Open Source through its projects and events.
We work in innovation through three axes:
- Our internal projects: Funded by PoC and carried out by our members, in partnership with foundations and research actors.
- Our services: For innovative companies in all sectors
- Our events: Workshops, talks or hackathons on the theme of technological innovation
"""
print(nlp_qa(context=text, question='What is PoC ?'))
print(nlp_qa(context=text, question='What is PoC main goal ?'))
```
Ok, now we know ouu model is confident and pretty efficient with QnA, but what about Sentiments analysis ?\
Let's check with our report once again ;)
- `pipeline('sentiment-analysis')`: load sentiment analysis Transformer model
- `nlp_sentence_classif(text)`: do sentiment analysis on provided text
```
nlp_sentence_classif = pipeline('sentiment-analysis')
nlp_sentence_classif(reportInfos[4])
```
For the last test we will use a custom text and once again feel free to try with your own text.
```
text = "I'm soo glad to be here !"
TrSentymentAnalysis.getSentiment(text)
```
| github_jupyter |
<img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
# Python for Finance (2nd ed.)
**Mastering Data-Driven Finance**
© Dr. Yves J. Hilpisch | The Python Quants GmbH
<img src="http://hilpisch.com/images/py4fi_2nd_shadow.png" width="300px" align="left">
# Model Calibration
## The Data
```
import numpy as np
import pandas as pd
import datetime as dt
from pylab import mpl, plt
plt.style.use('seaborn')
mpl.rcParams['font.family'] = 'serif'
%config InlineBackend.figure_format = 'svg'
import sys
sys.path.append('../')
sys.path.append('../dx')
dax = pd.read_csv('../../source/tr_eikon_option_data.csv',
index_col=0)
for col in ['CF_DATE', 'EXPIR_DATE']:
dax[col] = dax[col].apply(lambda date: pd.Timestamp(date))
dax.info()
dax.set_index('Instrument').head(7)
initial_value = dax.iloc[0]['CF_CLOSE']
calls = dax[dax['PUTCALLIND'] == 'CALL'].copy()
puts = dax[dax['PUTCALLIND'] == 'PUT '].copy()
calls.set_index('STRIKE_PRC')[['CF_CLOSE', 'IMP_VOLT']].plot(
secondary_y='IMP_VOLT', style=['bo', 'rv'], figsize=(10, 6));
# plt.savefig('../../images/ch21/dx_cal_01.png');
ax = puts.set_index('STRIKE_PRC')[['CF_CLOSE', 'IMP_VOLT']].plot(
secondary_y='IMP_VOLT', style=['bo', 'rv'], figsize=(10, 6))
ax.get_legend().set_bbox_to_anchor((0.25, 0.5));
# plt.savefig('../../images/ch21/dx_cal_02.png');
```
## Model Calibration
### Relevant Market Data
```
limit = 500
option_selection = calls[abs(calls['STRIKE_PRC'] - initial_value) < limit].copy()
option_selection.info()
option_selection.set_index('Instrument').tail()
option_selection.set_index('STRIKE_PRC')[['CF_CLOSE', 'IMP_VOLT']].plot(
secondary_y='IMP_VOLT', style=['bo', 'rv'], figsize=(10, 6));
# plt.savefig('../../images/ch21/dx_cal_03.png');
```
### Option Modeling
```
from valuation_mcs_european import valuation_mcs_european
from jump_diffusion import jump_diffusion
from market_environment import market_environment
from constant_short_rate import constant_short_rate
from derivatives_position import derivatives_position
from derivatives_portfolio import derivatives_portfolio
pricing_date = option_selection['CF_DATE'].max()
me_dax = market_environment('DAX30', pricing_date)
maturity = pd.Timestamp(calls.iloc[0]['EXPIR_DATE'])
me_dax.add_constant('initial_value', initial_value)
me_dax.add_constant('final_date', maturity)
me_dax.add_constant('currency', 'EUR')
me_dax.add_constant('frequency', 'B')
me_dax.add_constant('paths', 10000)
csr = constant_short_rate('csr', 0.01)
me_dax.add_curve('discount_curve', csr)
me_dax.add_constant('volatility', 0.2)
me_dax.add_constant('lambda', 0.8)
me_dax.add_constant('mu', -0.2)
me_dax.add_constant('delta', 0.1)
dax_model = jump_diffusion('dax_model', me_dax)
me_dax.add_constant('strike', initial_value)
me_dax.add_constant('maturity', maturity)
payoff_func = 'np.maximum(maturity_value - strike, 0)'
dax_eur_call = valuation_mcs_european('dax_eur_call',
dax_model, me_dax, payoff_func)
dax_eur_call.present_value()
option_models = {}
for option in option_selection.index:
strike = option_selection['STRIKE_PRC'].loc[option]
me_dax.add_constant('strike', strike)
option_models[strike] = valuation_mcs_european(
'eur_call_%d' % strike,
dax_model,
me_dax,
payoff_func)
def calculate_model_values_old(p0):
''' Returns all relevant option values.
Parameters
===========
p0: tuple/list
tuple of kappa, theta, volatility
Returns
=======
model_values: dict
dictionary with model values
'''
volatility, lamb, mu, delta = p0
dax_model.update(volatility=volatility, lamb=lamb, mu=mu, delta=delta)
model_values = {}
for strike in option_models:
model_values[strike] = option_models[strike].present_value(fixed_seed=True)
return model_values
def calculate_model_values(p0):
''' Returns all relevant option values.
Parameters
===========
p0: tuple/list
tuple of kappa, theta, volatility
Returns
=======
model_values: dict
dictionary with model values
'''
volatility, lamb, mu, delta = p0
dax_model.update(volatility=volatility, lamb=lamb,
mu=mu, delta=delta)
return {
strike: model.present_value(fixed_seed=True)
for strike, model in option_models.items()
}
calculate_model_values((0.1, 0.1, -0.4, 0.0))
```
### Calibration Procedure
```
i = 0
def mean_squared_error(p0):
''' Returns the mean-squared error given
the model and market values.
Parameters
===========
p0: tuple/list
tuple of kappa, theta, volatility
Returns
=======
MSE: float
mean-squared error
'''
global i
model_values = np.array(list(calculate_model_values(p0).values()))
market_values = option_selection['CF_CLOSE'].values
option_diffs = model_values - market_values
MSE = np.sum(option_diffs ** 2) / len(option_diffs)
if i % 75 == 0:
if i == 0:
print('%4s %6s %6s %6s %6s --> %6s' %
('i', 'vola', 'lambda', 'mu', 'delta', 'MSE'))
print('%4d %6.3f %6.3f %6.3f %6.3f --> %6.3f' %
(i, p0[0], p0[1], p0[2], p0[3], MSE))
i += 1
return MSE
mean_squared_error((0.1, 0.1, -0.4, 0.0))
import scipy.optimize as spo
%%time
i = 0
opt_global = spo.brute(mean_squared_error,
((0.10, 0.201, 0.025), # range for volatility
(0.10, 0.80, 0.10), # range for jump intensity
(-0.40, 0.01, 0.10), # range for average jump size
(0.00, 0.121, 0.02)), # range for jump variability
finish=None)
mean_squared_error(opt_global)
%%time
i = 0
opt_local = spo.fmin(mean_squared_error, opt_global,
xtol=0.00001, ftol=0.00001,
maxiter=200, maxfun=550)
i = 0
mean_squared_error(opt_local)
calculate_model_values(opt_local)
option_selection['MODEL'] = np.array(list(calculate_model_values(opt_local).values()))
option_selection['ERRORS_EUR'] = (option_selection['MODEL'] -
option_selection['CF_CLOSE'])
option_selection['ERRORS_%'] = (option_selection['ERRORS_EUR'] /
option_selection['CF_CLOSE']) * 100
option_selection[['MODEL', 'CF_CLOSE', 'ERRORS_EUR', 'ERRORS_%']]
round(option_selection['ERRORS_EUR'].mean(), 3)
round(option_selection['ERRORS_%'].mean(), 3)
fix, (ax1, ax2, ax3) = plt.subplots(3, sharex=True, figsize=(10, 10))
strikes = option_selection['STRIKE_PRC'].values
ax1.plot(strikes, option_selection['CF_CLOSE'], label='market quotes')
ax1.plot(strikes, option_selection['MODEL'], 'ro', label='model values')
ax1.set_ylabel('option values')
ax1.legend(loc=0)
wi = 15
ax2.bar(strikes - wi / 2., option_selection['ERRORS_EUR'], width=wi)
ax2.set_ylabel('errors [EUR]')
ax3.bar(strikes - wi / 2., option_selection['ERRORS_%'], width=wi)
ax3.set_ylabel('errors [%]')
ax3.set_xlabel('strikes');
# plt.savefig('../../images/ch21/dx_cal_04.png');
```
## Market-Based Valuation
### Modeling Option Positions
```
me_dax = market_environment('me_dax', pricing_date)
me_dax.add_constant('initial_value', initial_value)
me_dax.add_constant('final_date', pricing_date)
me_dax.add_constant('currency', 'EUR')
me_dax.add_constant('volatility', opt_local[0])
me_dax.add_constant('lambda', opt_local[1])
me_dax.add_constant('mu', opt_local[2])
me_dax.add_constant('delta', opt_local[3])
me_dax.add_constant('model', 'jd')
payoff_func = 'np.maximum(strike - instrument_values, 0)'
shared = market_environment('share', pricing_date)
shared.add_constant('maturity', maturity)
shared.add_constant('currency', 'EUR')
option_positions = {}
option_environments = {}
for option in option_selection.index:
option_environments[option] = market_environment(
'am_put_%d' % option, pricing_date)
strike = option_selection['STRIKE_PRC'].loc[option]
option_environments[option].add_constant('strike', strike)
option_environments[option].add_environment(shared)
option_positions['am_put_%d' % strike] = \
derivatives_position(
'am_put_%d' % strike,
quantity=np.random.randint(10, 50),
underlying='dax_model',
mar_env=option_environments[option],
otype='American',
payoff_func=payoff_func)
```
### The Options Portfolio
```
val_env = market_environment('val_env', pricing_date)
val_env.add_constant('starting_date', pricing_date)
val_env.add_constant('final_date', pricing_date)
val_env.add_curve('discount_curve', csr)
val_env.add_constant('frequency', 'B')
val_env.add_constant('paths', 25000)
underlyings = {'dax_model' : me_dax}
portfolio = derivatives_portfolio('portfolio', option_positions,
val_env, underlyings)
%time results = portfolio.get_statistics(fixed_seed=True)
results.round(1)
results[['pos_value','pos_delta','pos_vega']].sum().round(1)
```
<img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
<a href="http://tpq.io" target="_blank">http://tpq.io</a> | <a href="http://twitter.com/dyjh" target="_blank">@dyjh</a> | <a href="mailto:training@tpq.io">training@tpq.io</a>
| github_jupyter |
# StructN2V - 2D Example for Synthetic Membrane Data
Clean signal simulated/provided by [Alex Dibrov]("Alexandr Dibrov" <dibrov@mpi-cbg.de>)
```
# We import all our dependencies
from n2v.models import N2VConfig, N2V
import numpy as np
from csbdeep.utils import plot_history
from n2v.utils.n2v_utils import manipulate_val_data, autocorrelation
from n2v.internals.N2V_DataGenerator import N2V_DataGenerator
from matplotlib import pyplot as plt
import urllib
import os
import zipfile
```
# Training Data Preparation
```
# create a folder for our data
if not os.path.isdir('./data'):
os.mkdir('data')
# check if data has been downloaded already
dataPath="data/gt.npy"
if not os.path.exists(dataPath):
_ = urllib.request.urlretrieve('https://cloud.mpi-cbg.de/index.php/s/9LawR2GwE5WGrxw/download', dataPath)
X = np.load(dataPath).astype(np.float32)
plt.imshow(X[0]) ## clean signal simulated fluorescent cell membranes in 2D epithelium
## compute the [autocorrelation](https://en.wikipedia.org/wiki/Autocorrelation) for each 2D image
xautocorr = np.array([autocorrelation(_x) for _x in X])
## notice faint hexagonal symmetry of cells
x = xautocorr.mean(0)
def crop_square_center(x,w=20):
a,b = x.shape
x = x[a//2-w:a//2+w,b//2-w:b//2+w]
return x
plt.imshow(crop_square_center(x,18))
## generate synthetic structured noise by convolving pixelwise independent noise with a small 3x1 kernel.
## Then add this noise to the clean signal to generate our `noisy_dataset`.
from scipy.ndimage import convolve
purenoise = []
noise_kernel = np.array([[1,1,1]])/3 ## horizontal correlations
a,b,c = X.shape
for i in range(a):
noise = np.random.rand(b,c)*1.5
noise = convolve(noise,noise_kernel)
purenoise.append(noise)
purenoise = np.array(purenoise)
purenoise = purenoise - purenoise.mean()
noisy_dataset = X + purenoise
plt.imshow(noisy_dataset[20])
## Autocorrelation (top row) vs Data (bottom row)
## Notice how the autocorrelation of the noise (far right) reveals the horizontal shape of `noise_kernel` used above.
## Also see how the autocorrelation of the `noisy_dataset` (center top) is a combination of that of the signal and the noise?
fig,axs = plt.subplots(2,3, gridspec_kw = {'wspace':0.025, 'hspace':0.025}, figsize=(18,12))
def ac_and_crop(x):
x = autocorrelation(x)
a,b = x.shape
x = x[a//2-20:a//2+20, b//2-20:b//2+20]
return x
x1,x2,x3 = ac_and_crop(X[0]), ac_and_crop(noisy_dataset[0]), ac_and_crop(purenoise[0])
axs[0,0].imshow(x1)
axs[0,1].imshow(x2)
axs[0,2].imshow(x3)
axs[1,0].imshow(X[0])
axs[1,1].imshow(noisy_dataset[0])
axs[1,2].imshow(purenoise[0])
for a in axs.flat: a.axis('off')
## shuffle and randomly split the data into training and validation sets
inds = np.arange(X.shape[0])
np.random.shuffle(inds)
X_val = noisy_dataset[inds[:800]][...,None]
X_train = noisy_dataset[inds[800:]][...,None]
```
# Configure
```
config = N2VConfig(X_train, unet_kern_size=3,
train_steps_per_epoch=10, train_epochs=30, train_loss='mse', batch_norm=True,
train_batch_size=128, n2v_perc_pix=0.198, n2v_patch_shape=(64, 64),
unet_n_first = 96,
unet_residual = True,
n2v_manipulator='normal_withoutCP', n2v_neighborhood_radius=2,
structN2Vmask = [[0,1,1,1,1,1,0]]) ## mask should be wide enough to cover most of the noise autocorrelation
# Let's look at the parameters stored in the config-object.
vars(config)
# a name used to identify the model
model_name = 'structn2v_membrane_sim_normal_withoutCP'
# the base directory in which our model will live
basedir = 'models'
# We are now creating our network model.
model = N2V(config, model_name, basedir=basedir)
model.prepare_for_training(metrics=())
```
# Training
Training the model will likely take some time. We recommend to monitor the progress with TensorBoard, which allows you to inspect the losses during training. Furthermore, you can look at the predictions for some of the validation images, which can be helpful to recognize problems early on.
You can start TensorBoard in a terminal from the current working directory with tensorboard --logdir=. Then connect to http://localhost:6006/ with your browser.
```
# We are ready to start training now.
history = model.train(X_train, X_val)
```
### After training, lets plot training and validation loss.
```
print(sorted(list(history.history.keys())))
plt.figure(figsize=(16,5))
plot_history(history,['loss','val_loss']);
```
# Compute PSNR to GT
```
def PSNR(gt, img):
mse = np.mean(np.square(gt - img))
return 20 * np.log10(1.0) - 10 * np.log10(mse)
pred = []
psnrs = []
for gt, img in zip(X, noisy_dataset):
p_ = model.predict(img.astype(np.float32), 'YX');
pred.append(p_)
psnrs.append(PSNR(gt, p_))
psnrs = np.array(psnrs)
pred = np.array(pred)
print("PSNR: {:.3f} {:.3f}".format(psnrs.mean(), psnrs.std()))
print("-------------------")
print("Means: {:.3f} {:.3f} {:.3f}".format(X.mean(),noisy_dataset.mean(),pred.mean()))
print("Stds: {:.3f} {:.3f} {:.3f}".format(X.std(),noisy_dataset.std(),pred.std()))
fig,axs = plt.subplots(1,3,figsize=(6*3,6))
axs[0].imshow(noisy_dataset[2], interpolation='nearest')
axs[1].imshow(X[2], interpolation='nearest')
axs[2].imshow(pred[2], interpolation='nearest')
fig.subplots_adjust(wspace=0.025, hspace=0.025)
for a in axs.flat: a.axis('off')
```
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy
import requests
import urllib3
```
## Log scales with Numpy
En el siguiente ejemplo, compararemos la frecuencia de palabras en dos fragmentos: uno del Martín Fierro y uno de Don Quijote de la Mancha. Primero leeremos los textos y obtendremos las palabras más frecuentes, junto con su número de ocurrencias, para ambas obras.
```
martin_fierro_url = 'https://cs.famaf.unc.edu.ar/~mteruel/datasets/diplodatos/martin_fierro.txt'
quijote_url = 'https://cs.famaf.unc.edu.ar/~mteruel/datasets/diplodatos/quijote.txt'
def get_frequent_words(target_url):
words = []
http = urllib3.PoolManager()
for line in urllib3.PoolManager().request('GET', target_url).data.decode('utf-8').split():
if len(line) > 2:
words.extend(line.split())
unique_words, counts = numpy.unique(words, return_counts=True)
sorted_counts = numpy.argsort(counts * -1)
most_frequent_words = [unique_words[i] for i in sorted_counts]
most_frequent_counts = counts[sorted_counts]
return most_frequent_words, most_frequent_counts
words_m, counts_m = get_frequent_words(martin_fierro_url)
words_q, counts_q = get_frequent_words(quijote_url)
[x for x in zip(words_m, counts_m)][:10]
[x for x in zip(words_q, counts_q)][:10]
```
Sabemos que la distribución de palabras en lenguaje natural sigue una distribución exponencial (Ley de Zipf), al igual que la mayoría de los fenómenos humanos. Si graficamos la distribución de las frecuencias, obtenemos un gráfico que no nos permite comparar fácilmente los valores más pequeños.
```
plt.figure(figsize=(10,5))
plt.bar(numpy.arange(150), counts_q[:150])
plt.xlabel('Word frequency')
plt.ylabel('Count of words')
plt.title('Distribution of word frequency in Don Quijote de la Mancha')
```
Para solucionar este problema, podemos usar una escala logarítmica en el eje de las y, en lugar de una lineal. En el siguiente ejemplo vemos como se programa un gráfico de barras dobre con **matplotlib** y cómo le agregamos la escala logarítmica.
```
import matplotlib.pyplot as plt
import numpy as np
data = [_ for _ in zip(counts_m, counts_q)][:150]
dimw = 0.75 / len(data[0]) # Width of the bars
fig, ax = plt.subplots()
fig.set_size_inches(10, 5)
x = np.arange(len(data))
for i in range(len(data[0])):
y = [d[i] for d in data]
b = ax.bar(x + (i * dimw) - dimw / 2, y, dimw)
ax.set_yscale('log')
ax.set_xlabel('Most frequent words')
ax.set_ylabel('Frequency')
```
----
## Múltiples variables con Seaborn
Credits to https://gist.github.com/mwaskom/8224591
En la siguiente sección vamos a ver un ejemplo para graficar la distribución de una dataset incluyendo tantas variables como sea posible. El dataset del titanic es muy conocido para data science y machine learning, encontrarán muchas descripciones y ejemplos.
```
import pandas
import seaborn
seaborn.set_style('whitegrid')
seaborn.set_context('talk', font_scale=1.5)
titanic = pandas.read_csv(
'https://cs.famaf.unc.edu.ar/~mteruel/datasets/diplodatos/titanic_train.csv')
titanic[:5]
palette ={'female':'#FF686B', 'male':'#84DCC6'}
seaborn.factorplot(
'Survived', 'Age', data=titanic, hue='Sex',
row='Pclass', col='Embarked', kind='bar',
palette=palette)
seaborn.despine()
```
---
# Titanic plots with R
En las siguientes celdas veremos como utilizar gráficos de R en una notebook de python. Las instrucciones de instalación están en el archivo README.md
```
import rpy2
%load_ext rpy2.ipython
```
El %% indica un magic, esto quiere decir una instrucción sobre la ejecución del código. En este caso, indica que se debe utilizar el kernel de R.
Debemos importar también las librerías de R.
```
%%R
library(ggplot2)
library(reshape2)
```
Pueden encontrar un tutorial completo de ggPlot en https://github.com/crscardellino/MeetupDSCba2017/blob/master/python_with_ggplot.ipynb
Leemos la variable de python donde hemos guardado el dataset con -i
```
%%R -i titanic -w 10 -h 4 -u in
ggplot(titanic, aes(x=Age, fill=Sex)) +
geom_histogram(binwidth=2, aes(y = ..density..)) +
geom_density(alpha=0.5) + labs(x='Edad', y='Densidad de población')+
facet_wrap(~Sex) + scale_fill_brewer(palette='Set1', name='Sexo')
%%R -i titanic
ggplot(titanic, aes(x=Age, y=Fare)) + geom_count()
%%R -i titanic
ggplot(titanic, aes(x=Sex, y=Age)) + geom_boxplot() + geom_dotplot(binaxis='y',
stackdir='center',
dotsize = .2,
fill="red")
```
---
## Instalando librerías en R
En esta sección utilizaresmos una librería nueva para R, llamada ggparallel. Utilizaremos ggparallel para generar diagramas Sankey.
Se pueden instalar nuevas librerías directamente desde la notebook con un comando como el siguiente:
```
%%R
install.packages("ggparallel")
%%R
library(ggparallel)
```
En este caso, en lugar de utilizar el dataset del Titanic que ya habíamos leído con pandas, usaremos el que viene dado con la distribución de R. Al igual que pandas, R almacena los datos en dataframes con funciones y accesores muy similares.
```
%%R -w 10 -h 8 -u in
data(Titanic)
titanic <- as.data.frame(Titanic)
ggparallel(names(titanic)[c(1,4,2,1)], order=0, titanic, weight="Freq") +
scale_fill_brewer(palette="Paired", guide="none") +
scale_colour_brewer(palette="Paired", guide="none") +
theme(text=element_text(size=15), axis.text=element_text(size=15))
```
---
# Conflictos mundiales
En esta sección analizaremos un nuevo dataset sobre conflictos mundiales. Combinaremos dos dataset: uno descripto en [este documento](http://www.pcr.uu.se/digitalAssets/63/a_63324-f_Codebook_UCDP_PRIO_Armed_Conflict_Dataset_v4_2011.pdf) con un reporte de los conflicto nacionales e internaciones, y otro con un listado de países actuales y sus continentes.
Nota: las claves de region tiene el siguiente significado
1 Europe,
2 Middle east,
3 Asia,
4 Africa,
5 Americas
```
import pandas
countries = pandas.read_csv('https://cs.famaf.unc.edu.ar/~mteruel/datasets/diplodatos/Countries-Continents.csv')
conflicts = pandas.read_csv('https://cs.famaf.unc.edu.ar/~mteruel/datasets/diplodatos/armed_conflicts.csv')
```
Imprimimos muestras de ambos datasets
```
conflicts[:5]
countries[:5]
```
Combinamos la información tomando como clave el nombre del país y agregando el nombre del continente correspondiente. Notar que estamos filtrando todas las filas donde SideA o SideB no estén listados como países actuales.
```
country_conflicts = conflicts[conflicts.SideB.isin(countries.Country)][
['SideA', 'SideB', 'Location', 'Terr', 'Region', 'YEAR']]
country_conflicts = country_conflicts.merge(
countries[['Continent', 'Country']].rename(columns={'Continent': 'ContinentA'}),
left_on=['SideA'], right_on=['Country']).merge(
countries[['Continent', 'Country']].rename(columns={'Continent': 'ContinentB'}),
left_on=['SideB'], right_on=['Country'])#.drop(columns=['Country_x', 'Country_y'])
country_conflicts[:3]
```
Generamos el diagrama de Sankey. Definimos los colores manualmente para que se repitan en ambos lados
```
%%R -i country_conflicts -w 10 -h 8 -u in
colors <- c("#999999", "#E69F00", "#56B4E9", "#009E73", "#F0E442")
ggparallel(list("ContinentA", "ContinentB", "Region"),
data=country_conflicts, text.angle=0, alpha=0.25) +
theme(legend.position="none") +
scale_fill_manual(values = rep(colors, 14)) +
scale_colour_manual(values = rep(colors, 14)) +
theme(text=element_text(size=15), axis.text=element_text(size=15))
```
Nota: En las filminas pueden encontrar más información sobre los errores que tiene este gráfico, incluyendo el preproceso de datos.
En lugar de un diagrama Sankey, un heatmap o diagrama de calor también muestra la misma información.
Para crear un heatmap utilizando seaborn, necesitamos los datos como una matriz. Por suerte, pandas viene con una función que nos simplica este cálculo.
```
continent_joint_freq = pandas.crosstab(
index=country_conflicts["ContinentA"], columns=country_conflicts["ContinentB"])
continent_joint_freq
seaborn.heatmap(continent_joint_freq, annot=True)
```
En ggplot, por otra parte, la matriz debe tener solamente 3 columnas. Cada fila es una tripla (valor del eje x, valor del eje y, valor de la celda x,y)
```
continent_joint_freq = country_conflicts[["ContinentA", "ContinentB", "Region"]].groupby(
["ContinentA", "ContinentB"]).count().reset_index().rename(columns={'Region': 'Count'})
%%R -i continent_joint_freq
ggplot(data = continent_joint_freq, aes(x=ContinentA, y=ContinentB, fill=Count)) +
geom_tile() + scale_fill_gradient(low = "white", high = "steelblue")
```
Puede haber muchos problemas con el dataset, comenzando por el filtrado de paises. Plotearemos ahora la cantidad de conflictos por pais, con respecto a la zona donde se desarrollaron.
La principal desventaja de este método es que, al haber tantos países, debemos limitarnos y mostrar sólo un subconjunto. En este caso, sólo aquellos que tengan más de 10 conflictos.
```
conflicts_region = conflicts[["SideA", "SideB", "Region"]].groupby(
["SideA", "Region"]).count().reset_index().rename(columns={'SideB': 'Count'})
conflicts_region = conflicts_region[conflicts_region.Count > 10]
%%R -i conflicts_region
ggplot(data = conflicts_region, aes(x=Region, y=SideA, fill=Count)) +
geom_tile() + scale_fill_gradient(low = "white", high = "steelblue") +
theme(axis.text.x = element_text(angle = 90, hjust = 1))
```
---
# Visualizaciones con D3
Solamente vamos a dejar los datos procesados para poder leerlos desde el javascript
Es recomendable luego checkear los nombres de los paises, ya que el archivo que usaremos como origen de las coordenadas no tiene los mismos nombres. Por ej: United States vs United States of America.
```
conflicts.SideA.value_counts().to_json('countries_conflicts.json')
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import os, math
import numpy as np, pandas as pd
import matplotlib.pyplot as plt, seaborn as sns
from pandas_summary import DataFrameSummary
from tqdm import tqdm, tqdm_notebook
from pathlib import Path
pd.set_option('display.max_columns', 1000)
pd.set_option('display.max_rows', 400)
sns.set()
os.chdir('../..')
from src import utils
DATA = Path('data')
RAW = DATA/'raw'
INTERIM = DATA/'interim'
PROCESSED = DATA/'processed'
SUBMISSIONS = DATA/'submissions'
from surprise import dump
_, svd = dump.load(PROCESSED/'svd.dump')
_, nmf = dump.load(PROCESSED/'nmf.dump')
week_labels = [20180226, 20180305, 20180312, 20180319,
20180326, 20180402, 20180409, 20180416, 20180423]
%%time
weeks = []
for name in week_labels:
weeks.append(pd.read_feather(PROCESSED/f'week_{name % 10000:04}_diffscount.feather'))
```
## SVD features
```
uid = svd.trainset._raw2inner_id_users
iid = svd.trainset._raw2inner_id_items
user_bias = lambda x: svd.bu[uid[x]]
item_bias = lambda x: svd.bi[iid[x]]
recommend = lambda r: svd.predict(r.CustomerIdx, r.IsinIdx).est
%%time
from tqdm._tqdm_notebook import tqdm_notebook
tqdm_notebook.pandas()
for w in weeks:
w['SVD_CustomerBias'] = w.CustomerIdx.apply(user_bias)
w['SVD_IsinBias'] = w.IsinIdx.apply(item_bias)
w['SVD_Recommend'] = w.apply(recommend, axis=1)
%%time
for n, w in zip(week_labels, weeks):
print(n)
customer_factors = np.array([svd.pu[uid[cIdx]] for cIdx in w.CustomerIdx])
isin_factors = np.array([svd.qi[iid[iIdx]] for iIdx in w.IsinIdx])
for i in range(customer_factors.shape[1]):
w[f'SVD_CustomerFactor{i:02}'] = customer_factors[:,i]
for i in range(isin_factors.shape[1]):
w[f'SVD_IsinFactor{i:02}'] = isin_factors[:,i]
%%time
for name, w in zip(week_labels, weeks):
w.to_feather(PROCESSED/f'week_{name % 10000:04}_SVD_diffscount.feather')
```
## Preprocessing
```
from functools import cmp_to_key
from src.utils import composite_rating_cmp
isin = pd.read_csv(RAW/'Isin.csv', low_memory=False)
ratings = list(isin.CompositeRating.value_counts().index)
ratings = sorted(ratings, key=cmp_to_key(composite_rating_cmp), reverse=True)
rank = {k: i for i, k in enumerate(ratings)}
%%time
for n, w in zip(week_labels, weeks):
print(n)
w['CompositeRating'] = w.CompositeRating.apply(lambda x: rank[x])
cat_cols = ['BuySell', 'Sector', 'Subsector', 'Region_x', 'Country',
'TickerIdx', 'Seniority', 'Currency', 'ActivityGroup',
'Region_y', 'Activity', 'RiskCaptain', 'Owner',
'IndustrySector', 'IndustrySubgroup', 'MarketIssue', 'CouponType']
id_cols = ['TradeDateKey', 'CustomerIdx', 'IsinIdx']
target_col = 'CustomerInterest'
pred_col = 'PredictionIdx'
from src.utils import apply_cats
for col in cat_cols:
weeks[-1][col] = weeks[-1][col].astype('category').cat.as_ordered()
for w in weeks[:-1]:
apply_cats(w, weeks[-1])
for w in weeks:
for col in cat_cols:
w[col] = w[col].cat.codes
```
## Model
```
from src.utils import run_model
from lightgbm import LGBMClassifier
metric_names = ['auc']
for i, w in enumerate(weeks[1:]):
train, val, test = weeks[i], w, weeks[-1]
print(train['TradeDateKey'].unique(),
val['TradeDateKey'].unique(),
test['TradeDateKey'].unique())
%%time
results = None
output = []
for i, w in enumerate(weeks[1:]):
train, val, test = weeks[i], weeks[-2], weeks[-1]
X_train, y_train = train.drop(id_cols + [target_col], axis=1), \
train[target_col]
if pred_col in val.columns: # when test acts as validation
X_val, y_val = None, None
else:
X_val, y_val = val.drop(id_cols + [target_col], axis=1), \
val[target_col]
X_test = test.drop(id_cols + [target_col, pred_col], axis=1)
y_test, _, results, model = run_model(
LGBMClassifier(n_estimators=100),
X_train, y_train, X_val, y_val, X_test,
metric_names, results,
params_desc='n_estimators=100',
dataset_desc=f'{week_labels[i]}_diffcounts',
early_stopping=False)
output.append([y_test, model])
results
# first 5 predictions (2018 data)
np.array([x[0] for x in output])[:,:5]
test[target_col] = np.mean([x[0] for x in output], axis=0)
```
## Submission
```
submission = pd.read_csv(RAW/'sample_submission.csv', low_memory=False)
submission = pd.merge(submission[['PredictionIdx']], test[['PredictionIdx', target_col]],
how='left', on='PredictionIdx')
submission[target_col].describe()
submission.head()
submission.to_csv(SUBMISSIONS/'18-lgbm_8weeks_SVD_diffscounts_0226-0416.csv', index=False)
```
## Feature importance
```
from lightgbm import plot_importance
plot_importance(output[0][1], figsize=(5,10))
plot_importance(output[-1][1], figsize=(5, 10))
```
| github_jupyter |
# Session 10: Unsupervised clustering with K-means
------------------------------------------------------
Introduction to Data Science & Machine Learning
*Pablo M. Olmos olmos@tsc.uc3m.es*
------------------------------------------------------
# Unsupervised Learning
In this notebook, we will study a particularly simple method to perform unsupervised learning over a dataset. In this case, we will use [K-means](https://www.datascience.com/blog/k-means-clustering) to divide our population in a set of **clusters**. In unsupervised learning, we expect that the structure behind our data can be interpreted or exploited for future applications.
This post is inspired by the excellent book [Python Data Science Handbook by Jake VanderPlas](https://jakevdp.github.io/PythonDataScienceHandbook/index.html). I mostly added all the theoretical background.
## Clustering with K-means
The K-means algorithm is rather simple. Given a **distance metric** $d(\mathbf{x},\mathbf{x}')$ and a number $K$ of clusters selected in advance, we perform as follows.
Until convergence:
1. Initialize $K$ *centroids* at random among points in the dataset. Denote the centroids as $\mathbf{c}_k$, $k=1,\ldots,K$.
2. For $i=1,\ldots,N$, assign point $\mathbf{x}^{(i)}$ to the cluster with the closest centroid. If $A(\mathbf{x}^{(i)})\in\{1,\ldots,K\}$ is the cluster assigned to point $\mathbf{x}^{(i)}$ then
\begin{align}
A(\mathbf{x}^{(i)}) = \arg \min_{\{1,\ldots,K\}} d(\mathbf{x}^{(i)},\mathbf{c}_k)
\end{align}
3. Recompute centroids as the mean across all points assigned to the same cluster. For $k=1,\ldots,K$,
\begin{align}
\mathbf{c}_k = \frac{1}{N_k} \sum_{i: A(\mathbf{x}^{(i)})= k} \mathbf{x}^{(i)},
\end{align}
where $\frac{1}{N_k}$ is the number of points assigned to cluster $k$.
4. Declare convergence if no changes between two consecutive iterations or maximum number of iterations achieved.
```
import matplotlib.pyplot as plt
import numpy as np
import scipy.io
from sklearn.datasets.samples_generator import make_blobs
from sklearn.cluster import KMeans,MiniBatchKMeans
%matplotlib inline
```
### Lets create a toy data set
```
X, y_true = make_blobs(n_samples=500, centers=6,
cluster_std=0.6, random_state=22)
plt.scatter(X[:, 0], X[:, 1], s=50);
plt.xlabel('$x_1$')
plt.ylabel('$x_2$')
#Lets run Kmeans clustering
K=4
kmeans = KMeans(n_clusters=K)
kmeans.fit(X)
y_kmeans = kmeans.predict(X)
```
Let's visualize the results by plotting the data colored by these labels. We will also plot the cluster centers as determined by the k-means estimator:
```
plt.scatter(X[:, 0], X[:, 1], c=y_kmeans, s=50, cmap='viridis')
centers = kmeans.cluster_centers_
plt.scatter(centers[:, 0], centers[:, 1], c='black', s=200, alpha=0.5)
```
## Selecting the number of clusters
The number $K$ of clusters represent the model complexity. When we set large number of clusters, we are assuming tha the data structure is very complex, and we might be certainly **overfitting** our data.
How to diagnose overfitting in unsupervised clustering? There are several approaches. A common one is to set a model performance metric and analyze how it depends on the model complexity.
In $K$-means, the natural choice is to analyze the **average distance of points w.r.t. their cluster**. This is in fact the function that we minimize to derive the $K$-means algorithm!
\begin{align}
\mathcal{L}(\mathbf{c}_1,\ldots,\mathbf{c}_K) = \frac{1}{K} \sum_{k=1}^K \sum_{i: A(\mathbf{x}^{(i)})= k} \frac{d(\mathbf{x}^{(i)},\mathbf{c}_k)}{N_k}
\end{align}
If we optimize the above function w.r.t. $\mathbf{c}_k$, $k=1,\ldots,K$, by implementing a [**coordinate gradient descent**](https://en.wikipedia.org/wiki/Coordinate_descent) we derive the $K$-means algorithm.
Lets plot the evolution of $\mathcal{L}(\mathbf{c}_1,\ldots,\mathbf{c}_K)$ for different values of $K$.
```
kmeans.inertia_
K_list = range(2,15)
L = []
for k in K_list:
kmeans = KMeans(n_clusters=k)
kmeans.fit(X)
L.append((kmeans.inertia_)/(0.0+k*X.shape[0]))
plt.figure()
plt.plot(K_list,L)
plt.xlabel('$K$')
plt.ylabel('$L$')
plt.grid()
```
Clearly, the cost function becomes flat for large $K$ values. This means that we are making our model **more complex** but we are not essentially explaining our data better! A sign of **overfitting**.
Given the above figure, it would make sense to select a value of $K$ around 6 (which is also the true value in this case).
Be aware that this plot is dependent of the number of data points. Also, that the $K$-means solution is **highly dependent** on the intialization. Actually, it is usual to run it several times and select the solution that provides the smallest $\mathcal{L}(\mathbf{c}_1,\ldots,\mathbf{c}_K)$ function. In the [**Sklearn K-means library**](http://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html), it is controlled by the parameter *n_init*.
```
N_list =[20,50,100,500]
K_list = range(2,8)
plt.figure()
for n in N_list:
X, y_true = make_blobs(n_samples=n, centers=6,
cluster_std=0.6, random_state=22)
L = []
for k in K_list:
kmeans = KMeans(n_clusters=k,n_init=5)
kmeans.fit(X)
L.append((kmeans.inertia_)/(0.0+k*X.shape[0]))
plt.plot(K_list,L,label='N_points='+str(n))
plt.xlabel('$K$')
plt.ylabel('$L$')
plt.grid()
plt.legend()
```
## Mini-batch $K$-means
Updating centroids at every $K$-means update using all datapoints can be costly for very large databases. An scalable and effective solution uses small **mini-batches** of data at every iteration. This is known as **mini-batch** $K$-means, and it is also provided in the [**sklearn library**](http://scikit-learn.org/stable/modules/generated/sklearn.cluster.MiniBatchKMeans.html#sklearn.cluster.MiniBatchKMeans).
Compare the running times of the following two examples.
```
# Without mini-batches. Full K-means
n =int(1e5)
K_list = range(2,8)
X, y_true = make_blobs(n_samples=n, centers=6,
cluster_std=0.6, random_state=22)
L = []
batch = 100
for k in K_list:
kmeans = MiniBatchKMeans(n_clusters=k,batch_size=batch,n_init=5)
kmeans.fit(X)
L.append((kmeans.inertia_)/(0.0+k*X.shape[0]))
plt.figure()
plt.plot(K_list,L)
plt.xlabel('$K$')
plt.ylabel('$L$')
plt.grid()
```
**Summary**: $K$-means is a simple and cheap algorithm for unsupervised clustering. However, it has fundamental weaknesses that we will explain when we introduce probabilisitic clustering using **Gaussian Mixture Models** and the **Expectation Maximization Algorithm**.
| github_jupyter |
## Import Libraries and Connect to STK/ODTK
```
# Import Python Libraries
# !pip install "C:\Program Files\AGI\STK 12\bin\AgPythonAPI\agi.stk12-12.2.0-py3-none-any.whl"
# !pip install opencv-python
import numpy as np
import pandas as pd
import cv2
import os
import shutil
import imageio
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import time
import win32com.client as w32c
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score
from skimage.color import rgb2gray
from scipy.spatial.transform import Rotation as R
from skimage.feature import peak_local_max
from astropy import units as u
from astropy.coordinates import Angle
%matplotlib inline
# Import STK Libraries
from agi.stk12.stkdesktop import STKDesktop
from agi.stk12.stkobjects import *
from agi.stk12.stkutil import *
from agi.stk12.vgt import *
from agi.stk12.utilities.colors import Color, Colors
# Import functions from EOIRProcessesing.py
from EOIRProcessingLib import *
# Attatch to open instance of STK
stk = STKDesktop.AttachToApplication() # attach to exisiting instance of stk
root = stk.Root
root.Isolate()
root.UnitPreferences.Item('DateFormat').SetCurrentUnit('EpSec')
sc = root.CurrentScenario
# Attach to exisiting instance of ODTK
try:
#
app = w32c.GetActiveObject("ODTK7.Application")
ODTK = app.Personality
except:
print('Did not connect to ODTK')
```
## Examples and Inputs
Setup Considerations
* Use a sensor type of Fixed or Fixed in Axes if you are updating the sensor pointing based on observations
* Using updateSensorMethod = 'previousMeasurementDirection' requires azStart and elStart to be defined
* If your satellite is maneuevering or changing attitude, you might want to use a frame other than the body frame
* Depending on what type of sensor you are modeling, it may be beneficial to set constrains such as Sun exclusion angle, target must be lit, space only backgrounds, in STK and/or ODTK
* If you plan on running ODTK ensure STK and ODTK are using the same force models.See: https://help.agi.com/ODTK/index.htm#../LinkedDocuments/astrodynamicsConsistency.pdf
* Right now the measurements passed to ODTK are assumed to be spacebased and passed in the form of a .geosc. The file format supports non-spacedbased measurements (facility observations), but the code to write the .geosc file would need to be updated to support this, see the function RADECToMeasurementFileLine in EOIRProcessingLib.py
* You may want to modify the file paths of where images/videos are saved
```
# Set Up
sensorPath = '*/Satellite/LEO/Sensor/LEOLaunchTracker'
tstart = 0
tstop = 120
tstep = 2
# Image Processing
minSNR = 3
percentofmax = 0.25
method = 'localpeaks' # 'localpeaks','minSNR','percentofmax','kmeans'
maxObjects = 1
k = 1/3 # Common Values: 1/3(enhances dark spots), 1 (simply normalizes), and 2 (enhances bright spots)
# Update Sensor Position
updateSensorMethod = 'previousMeasurementDirection' # '' (this will skip updating), 'previousmeasurementdirection','odtk'
azStart = -90
elStart = 42
# # Set Up
# sensorPath = '*/Satellite/LEO/Sensor/LEODetector'
# tstart = 0
# tstop = 500
# tstep = 10
# # Image Processing
# minSNR = 3
# percentofmax = 0.25
# method = 'localpeaks' # 'localpeaks','minSNR','percentofmax','kmeans'
# maxObjects = 1
# k = 1/3 # Common Values: 1/3(enhances dark spots), 1 (simply normalizes), and 2 (enhances bright spots)
# differenceImages = False # Only use for staring sensors
# # Update Sensor Position
# updateSensorMethod = '' # '' (this will skip updating), 'previousmeasurementdirection','odtk'
# # Set Up
# sensorPath = '*/Aircraft/UAV/Sensor/HGVTracker'
# tstart = 1000
# tstop = 1300
# tstep = 5 # 10
# # Image Processing
# minSNR = 2.5
# percentofmax = 0.25
# method = 'localpeaks' # 'localpeaks','minSNR','percentofmax','kmeans'
# maxObjects = 1
# k = 1/3 # Common Values: 1/3(enhances dark spots), 1 (simply normalizes), and 2 (enhances bright spots)
# # Update Sensor Position
# updateSensorMethod = 'previousMeasurementDirection' # '' (this will skip updating), 'previousmeasurementdirection','odtk'
# azStart = 170
# elStart = 0
# # Set Up
# sensorPath = '*/Satellite/HEO/Sensor/HEOLaunch'
# tstart = 0
# tstop = 40
# tstep = 0.2
# # Image Processing
# minSNR = 5
# percentofmax = 0.25
# method = 'localpeaks' # 'localpeaks','minSNR','percentofmax','kmeans'
# maxObjects = 1
# k = 1/3 # Common Values: 1/3(enhances dark spots), 1 (simply normalizes), and 2 (enhances bright spots)
# differenceImages = False # Only use for staring sensors
# deltaTime = 3 # Use a multiple of the time step
# # Update Sensor Position
# updateSensorMethod = '' # '' (this will skip updating), 'previousmeasurementdirection','odtk'
# # Set Up, #Use the scal imagery
# sensorPath = '*/Aircraft/UAV/Sensor/LaunchTracker'
# tstart = 0.1
# tstop = 60.1
# tstep = .5
# # Image Processing
# minSNR = 3
# percentofmax = 0.25
# method = 'localpeaks' # 'localpeaks','minSNR','percentofmax','kmeans'
# maxObjects = 1
# k = 1/3 # Common Values: 1/3(enhances dark spots), 1 (simply normalizes), and 2 (enhances bright spots)
# # Update Sensor Position
# updateSensorMethod = 'previousMeasurementDirection' # '' (this will skip updating), 'previousmeasurementdirection','odtk'
# azStart = 100
# elStart = 6
# # Set Up, #Use the scal imagery
# sensorPath = '*/Satellite/GEO/Sensor/Stare'
# tstart = 0
# tstop = 500
# tstep = 10
# sensorPath = '*/Satellite/GEO/Sensor/StareHGV'
# tstart = 1000
# tstop = 1400
# tstep = 10
# # Image Processing
# minSNR = 3
# percentofmax = 0.25
# method = 'localpeaks' # 'localpeaks','minSNR','percentofmax','kmeans'
# maxObjects = 1
# k = 1/3 # Common Values: 1/3(enhances dark spots), 1 (simply normalizes), and 2 (enhances bright spots)
# # Update Sensor Position
# updateSensorMethod = '' # '' (this will skip updating), 'previousmeasurementdirection','odtk'
# # Set Up
# sensorPath = '*/Satellite/RPO/Sensor/Detector'
# tstart = 4800
# tstop = 12800
# tstep = 60
# # Image Processing
# minSNR = 5
# percentofmax = 0.25
# method = 'localpeaks' # 'localpeaks','minSNR','percentofmax','kmeans'
# maxObjects = 1
# k = 1 # Common Values: 1/3(enhances dark spots), 1 (simply normalizes), and 2 (enhances bright spots)
# # Update Sensor Position
# updateSensorMethod = 'odtk'# '' (this will skip updating), 'previousmeasurementdirection','odtk'
# Additional Inputs
# Output/Runtime Settings
writeVideo = False # Combine the tagged images into a video
realTimeRate = 5 # (tstop-tstart)/realTimeRate = videoLength assuming no frames are missing, how fast the final video should play relative to real time
reuseFiles = True # Reuse existing eoir images/text files if they already exist
# Limit Observations
useAccessConstraintsToLimitObs = False # Limit imaging times to be within access intervals
targetPath = '*/Satellite/TargetODTK' # Access object, usually the object you are trying to observe/detect
# Plotting
plotImage = True # Show the tagged images
# ODTK settings, Modify these as needed if you are running ODTK
ODSatName = 'RPO' # Odtk satellite name with the sensor
ODFilterName = 'Filter1' # Filter name
ODTargetName = 'Target' # Object you are trying to perform OD on
targetID = 1001 # Target tracking ID in ODTK
sensorID = 1000 # Sensor tracking ID in ODTK
RAstd = 0.00 # 1 standard deviation in arcSec, format is xx.xx, setting to 0 uses ODTK's value
DECstd = 0.00 # 1 standard deviation in arcSec, format is xx.xx, setting to 0 uses ODTK's value
usePixelSizeForStd = False # overrides the RAstd and DECstd values based on pixel size
visualizeOD = False # Set to True if you want to see the filter run in STK as it goes
# Look for differences in images instead of the image itself, Only use for staring sensors
differenceImages = False
deltaTime = tstep # Time to difference images, use a multiple of the time step
```
## Execute Imaging Over Time
```
# Get handles and setup for intiial config
sensor = root.GetObjectFromPath(sensorPath)
vectors = sensor.Vgt.Vectors
sensorName = sensorPath.split('/')[-1]
if differenceImages == True: # Differencing only works for staring images, don't update the sensor pointing
updateSensorMethod = ''
# Use a sensor type of Fixed or Fixed in Axes if you are updating the sensor pointing
if sensor.PointingType == AgESnPointing.eSnPtFixedAxes:
axes = sensor.Pointing.ReferenceAxes[:-5]
else:
axes = '/'.join(sensorPath.split('/')[1:3]) + ' Body'
print('Assuming Parent Body axes and the sensor pointing type is Fixed')
# Set up ODTK
if updateSensorMethod.lower() == 'odtk':
# Get ODTK handles
ODScenario = ODTK.Scenario(0)
ODSat = ODScenario.Satellite(ODSatName)
ODFilter = ODScenario.Filter(ODFilterName)
ODObject = ODScenario.Satellite(ODTargetName)
# Add measurement file
measurementFile = sensorName+".geosc"
open(measurementFile, 'w').close() # Clear RADecfile
measurementFiles = ODScenario.Measurements.Files
measurementFiles.clear()
newFile = measurementFiles.NewElem()
newFile.Filename = os.getcwd()+'\\'+measurementFile
measurementFiles.push_back(newFile)
ODScenario.Measurements.Files(0).Enabled = True
# Run filter with no Obs
ODFilter.ProcessControl.StopMode = 'TimeSpan'
ODFilter.ProcessControl.TimeSpan= (tstop-tstart)/3600
ODFilter.Go()
ODFilter.ProcessControl.StopMode = 'LastMeasurement'
predictionTimeSpan = 3600 # sec
ODFilter.Output.STKEphemeris.Predict.TimeSpan = predictionTimeSpan/60 # Convert to mins
ephemerisFile = ODFilter.Output.STKEphemeris.Files(0).FileName()
# Set up VGT
points = sensor.Vgt.Points
if not points.Contains('EstimatedTargetLocation'):
point = points.Factory.Create('EstimatedTargetLocation','Estimated Pointing Location',AgECrdnPointType.eCrdnPointTypeFile)
point.Filename = ephemerisFile
point = points.Item('EstimatedTargetLocation')
if not vectors.Contains('PointingDirection'):
vector = vectors.Factory.Create('PointingDirection','Estimated Pointing Direction',AgECrdnVectorType.eCrdnVectorTypeDisplacement)
vector.Destination.SetPath(sensorPath[2:]+' EstimatedTargetLocation')
# Set up filter run visualization
if visualizeOD == True:
if sc.Children.Contains(AgESTKObjectType.eSatellite,targetPath.split('/')[-1]+'Filter'):
satVis = root.GetObjectFromPath(targetPath+'Filter')
else:
satVis = sc.Children.New(AgESTKObjectType.eSatellite,targetPath.split('/')[-1]+'Filter')
satVis.SetPropagatorType(AgEVePropagatorType.ePropagatorStkExternal)
satVis.Propagator.Filename = ephemerisFile
satVis.VO.Covariance.Attributes.IsVisible = True
satVis.VO.OrbitSystems.RemoveAll()
satVis.VO.OrbitSystems.InertialByWindow.IsVisible = False
satVis.VO.OrbitSystems.Add(targetPath.split('*/')[-1]+' RIC System')
# Set up measurement vector
if not vectors.Contains('MeasurementDirection'):
vectors.Factory.Create('MeasurementDirection','Measurement Direction',AgECrdnVectorType.eCrdnVectorTypeFixedInAxes)
vector = vectors.Item('MeasurementDirection')
vector.ReferenceAxes.SetPath(axes)
# Update initial pointing
if updateSensorMethod.lower() == 'previousmeasurementdirection':
sensor.Pointing.Orientation.AssignAzEl(azStart,elStart,AgEAzElAboutBoresight.eAzElAboutBoresightRotate)
# Get the access handle
if useAccessConstraintsToLimitObs == True:
access = sensor.GetAccess(targetPath)
# Get pixel size
horizontalPixels,verticalPixels,horizontalAngle,verticalAngle = getSensorFOVAndPixels(sensor)
horizontalDegPerPixel = horizontalAngle/horizontalPixels
verticalDegPerPixel = verticalAngle/verticalPixels
horizontalCenter = horizontalPixels/2+0.5
verticalCenter = verticalPixels/2+0.5
# Update RA and Dec based on pixel size
if usePixelSizeForStd == True:
RAstd = horizontalDegPerPixel*3600
DECstd = verticalDegPerPixel*3600 # Be careful if this exceeds 99 arc Seconds, should add a check, could also leave blank to use ODTK values
if RAstd >=100 or DECstd >=100:
print('Setting standard deviations to use ODTK values')
RAstd = 00.0
DECstd = 00.0
# Set up times and measurements
times = np.arange(tstart,tstop,tstep)
times = np.append(times,tstop).round(3) # round to 1/1000th of a sec get rid of numerical issues
timesWithTaggedImages = []
measurementsAzElErrors = []
meaurementsAzEl = []
sensorPointingHistory = []
imageFolder = os.getcwd()+'\\Images'
t1 = time.time()
# Loop through time
for t in times:
# Set Animation time
print('Time:',t)
t = float(t)
root.CurrentTime = t
# If running ODTK, update sensor pointing to predicted target location
if updateSensorMethod.lower() == 'odtk':
# Have to force a filereload, there may be a better way, or could switch between the copy to save a bit of time
shutil.copyfile(ephemerisFile, ephemerisFile.split('.')[0]+'Copy.e') #copy src to dst
point.Filename = ephemerisFile.split('.')[0]+'Copy.e'
point.Filename = ephemerisFile
if visualizeOD == True:
satVis.Propagator.Filename = ephemerisFile.split('.')[0]+'Copy.e'
satVis.Propagator.Filename = ephemerisFile
satVis.Propagator.Propagate()
_,az,el = getPointingDirection(sensor,t,t,tstep,axes=axes)
sensor.Pointing.Orientation.AssignAzEl(az,el,AgEAzElAboutBoresight.eAzElAboutBoresightRotate)
print('Pointing Update:',az,el)
sensorPointingHistory.append((t,az,el))
# Limit imagine times to when the object has access, this will take into account pointing updates
if useAccessConstraintsToLimitObs == True:
access.SpecifyAccessTimePeriod(t,t)
access.ComputeAccess()
skipAccess = False if access.ComputedAccessIntervalTimes.Count != 0 else True
else:
skipAccess = False
# Image Processing and Tagging
if skipAccess == False:
# Construct file names
imageName = '"{}\\{}Time{}.jpg"'.format(imageFolder,sensorName,str(t).replace('.','_'))
textName = '"{}\\{}Time{}.txt"'.format(imageFolder,sensorName,str(t).replace('.','_'))
# Run EOIR
getEOIRImages(root,sensorPath,imageName='',textName=textName,reuseFiles=reuseFiles) # set imageName=imageName if you want EOIR to generate the image
# Image processing
data = np.loadtxt(textName.replace('"','')) # Load EOIR Image
# Difference Images if needed
if differenceImages == True:
textNamePrevious = '"{}\\{}Time{}.txt"'.format(imageFolder,sensorName,str(t-deltaTime).replace('.','_'))
if os.path.exists(textNamePrevious.replace('"','')):
dataPrevious = np.loadtxt(textNamePrevious.replace('"',''))
data = data-dataPrevious
else:
continue # skip to the next iteration if not previous image exists
timesWithTaggedImages.append(t) # Time where an image processing attempt was made
image = normalizeImage(data,k=k,convertToInt=False,plotImage=False)
# Get the object centers
maxSNR = getMaxSNR(image)
objectCenters = getObjectCenters(image,method=method,minSNR=minSNR,percentofmax=percentofmax,maxObjects=maxObjects)
if len(objectCenters) > 0:
objectCenters,objectSNRS = sortBySNR(image,objectCenters) # Puts highest SNR last
else:
objectSNRS = []
print('Object Centers:',objectCenters)
print('Object SNRs:',objectSNRS)
# Store Az El measurements
for objectCenter in objectCenters:
azError = (objectCenter[1]-horizontalCenter)*horizontalDegPerPixel
elError = (objectCenter[0]-verticalCenter)*verticalDegPerPixel
measurementsAzElErrors.append((t,azError,elError))
# Update pointing and measurements if an object is detected
if len(objectCenters) > 0:
print('Errors (deg): ',azError,elError)
# Obs Association: The image detection process can detect multiple objects and figuring out which obs corresponds to which object is non-trival. This is an area of future improvment, with Python or lettings ODTK/OAT help out
# Common issues: Mistags, missed observations/undetectable signals,false positives (find peaks which are not the actual object, stars, clouds, background clutter.) Be aware of this, better image processing or sensors could resolve this.
# For now the script assumes a very simple object association: the highest SNR is the specified target object and will use this for pointing updates
# Get current rotation matrix for sensor body frame to specified axes
timeRotationZYXs = computeSensorBodyToParentRotations(sensor,t,t,tstep,axes=axes)
# Can also compute true values aif desired
# trueAzElError = computeTrueSensorAzElError(sensor,t,t,tstep,target=targetPath.split('/')[-1])
# Calculate target direction in axes from az el offsets in sensor image
azMeas,elMeas,targetVec = updatePointingDir(measurementsAzElErrors[-1][1],measurementsAzElErrors[-1][2],timeRotationZYXs[0,:])
meaurementsAzEl.append((t,azMeas,elMeas))
# Update the measurement direction vector
vector.Direction.AssignXYZ(targetVec[0],targetVec[1],targetVec[2])
# Convert the measurement direction vector to RA and Dec measurements
if updateSensorMethod.lower() == 'odtk':
_,ra,dec = getRADECMeasurements(sensor,t,t,tstep,useMeasurementDirection=True)
with open(measurementFile,"a+") as f:
line = RADECToMeasurementFileLine(root,t,ra,dec,targetID=1001,sensorID=1000,RAstd=RAstd,DECstd=DECstd)
f.write(line)
print('RA Dec:',ra,dec)
ODFilter.Go()
if updateSensorMethod.lower() == 'previousmeasurementdirection':
sensor.Pointing.Orientation.AssignAzEl(azMeas,elMeas,AgEAzElAboutBoresight.eAzElAboutBoresightRotate)
sensorPointingHistory.append((t,azMeas,elMeas))
# Alternative approaches/future improvements: could estimate position and velocity, can do simple regression for velocity, could look at multiple images(over time and different sensors), could do missle model and kalman filter
# Plotting
if plotImage == True:
plt.figure(figsize=(8,8))
if differenceImages == True:
data = np.loadtxt(textName.replace('"','')) # reload data for saving the image
image = normalizeImage(data,k=k,convertToInt=False,plotImage=False)
plt.imshow(image)
plt.imshow(image,cmap='gray')
if len(objectCenters)>0:
plt.plot(objectCenters[:,1],objectCenters[:,0],'ro',markersize=12,markerfacecolor='none')
plt.title('Time: '+str(t))
plt.savefig(imageName.replace('"','').split('.')[0]+'Tagged.png',dpi=500)
plt.show()
# continue loop
print('RunTime: ',time.time()-t1)
else:
print('Skipped Time:',t) # No access
# Create a history of sensor pointing
writeSensorPointingFile(sensorPointingHistory,fileName='{}SensorPointing.sp'.format(sensorName),axes=axes)
print('Wrote ','{}SensorPointing.sp'.format(sensorName))
# Create a video
if writeVideo == True:
createVideo(sensorName,timesWithTaggedImages,tstep,imageFolder,realTimeRate=realTimeRate)
print('Wrote ','{}.mp4'.format(sensorName))
print(time.time()-t1)
```
| github_jupyter |
```
import silence_tensorflow.auto
import time
import numpy as np
import tensorflow as tf
import tensorflow.compat.v1 as tf_v1
from segmenter.slicer import Slice
start_time = time.time()
class ML:
model = ''
def __init__(self, model, vocabulary,input_dir,image,classification,seq):
self.model=model
self.vocabulary=vocabulary
self.input_dir=input_dir
self.slice_dir=slice_dir
self.classification=classification
self.seq=seq
def setup(self):
# Read the dictionary
dict_file = open(self.voc_file,'r')
dict_list = dict_file.read().splitlines()
self.int2word = dict()
for word in dict_list:
word_idx = len(int2word)
int2word[word_idx] = word
dict_file.close()
tf.reset_default_graph()
saver = tf_v1.train.import_meta_graph(self.model)
saver.restore(sess,self.model[:-5])
graph = tf_v1.get_default_graph()
self.input = graph.get_tensor_by_name("model_input:0")
self.seq_len = graph.get_tensor_by_name("seq_lengths:0")
self.rnn_keep_prob = graph.get_tensor_by_name("keep_prob:0")
self.height_tensor = graph.get_tensor_by_name("input_height:0")
self.width_reduction_tensor = graph.get_tensor_by_name("width_reduction:0")
self.logits = graph.get_tensor_by_name("fully_connected/BiasAdd:0")
self.decoded, _ = tf_v1.nn.ctc_greedy_decoder(self.logits, self.seq_len)
self.WIDTH_REDUCTION = 16
return self.sess = tf.InteractiveSession()
def predict(self,image,seq_lengths):
slices = Slice(image)
print("SLICE COMPLETED in: " + str(time.time() - start_time) )
prediction = session.run(model.decoded,
feed_dict={
self.input: image,
self.seq_len: seq_lengths,
self.rnn_keep_prob: 1.0,
})
print("PREDICTION COMPLETED in: " + str(time.time() - start_time) )
return prediction, slices
import os
import cv2
import numpy as np
from PIL import Image
#import tensorflow as tf
#import tensorflow.compat.v1 as tf
from segmenter.slicer import Slice
from PIL import Image
from PIL import ImageFont
import ctc_utils
from PIL import ImageDraw
from flask_ngrok import run_with_ngrok
from flask import Flask,request,send_from_directory,render_template
from ml_model import ML
import config
from apputil import normalize, resize, sparse_tensor_to_strs
from matplotlib.pyplot import imshow
# GLOBAL ACCESS
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
#THIS_FOLDER = os.path.dirname(os.path.abspath(__file__))
# SETUP APPLICATION
#app = Flask(__name__, static_url_path='')
#run_with_ngrok(app)
model = ML(config.model,config.voc_file,config.input_dir,
config.slice_dir,config.classification,config.seq)
session = model.setup()
#f = request.files['file']
#read image file string data
#filestr = request.files['file'].read()
#print(filestr)
#convert string data to numpy array
IMG_PATH = 'C:/Users/aroue/Downloads/Documents/@ML/Sheet Music/[easy/holynight.png'
image_string = open(IMG_PATH, 'rb')
#npimg = np.fromstring(image_string, np.uint8)
img = Image.open(image_string)
img = np.fromfile(image_string, np.uint8)
pre_image = cv2.imread(str(IMG_PATH),0)
#img = Image.open(request.files['file'])
#x = np.frombuffer(buffer(s), dtype='int8')
#npimg = normalize(npimg)
# convert numpy array to image
#img = cv2.imdecode(npimg,cv2.IMREAD_COLOR)IMREAD_UNCHANGED
#cvimg = cv2.imdecode(img, cv2.IMREAD_UNCHANGED)
pre_image = ctc_utils.resize(pre_image, 128)
pre_image = ctc_utils.normalize(pre_image)
#model_image = np.asarray(pre_image).reshape(1,pre_image.shape[0],-1,1)
shape = model_image.shape[2]
input_image = Image.fromarray(pre_image)
#if im.mode != 'RGB':
#im = im.convert('RGB')
#im.save('img/out.png')
#im.show()
%matplotlib inline
#pil_im = Image.open('data/empire.jpg', 'r')
#imshow(np.asarray(im))
#imshow(image)
#image = np.asarray(cvimg).reshape(1,cvimg.shape[0],-1,1)
#image = np.asarray(cvimg).reshape(1,cvimg.shape[0],cvimg.shape[1],1)
#img = Image.open(request.files['file'])
prediction, slices = model.predict(input_image, model_image, shape)
# str_predictions = sparse_tensor_to_strs(prediction)
# print(str_predictions)
# array_of_notes = []
# for w in str_predictions[0]:
# array_of_notes.append(model.int2word[w])
# notes=[]
# for i in array_of_notes:
# if i[0:5]=="note-":
# if not i[6].isdigit():
# notes.append(i[5:7])
# else:
# notes.append(i[5])
# img = Image.open(img).convert('L')
# size = (img.size[0], int(img.size[1]*1.5))
# layer = Image.new('RGB', size, (255,255,255))
# layer.paste(img, box=None)
# img_arr = np.array(layer)
# height = int(img_arr.shape[0])
# width = int(img_arr.shape[1])
# print(img_arr.shape[0])
# draw = ImageDraw.Draw(layer)
# # font = ImageFont.truetype(<font-file>, <font-size>)
# font = ImageFont.truetype("Aaargh.ttf", 16)
# # draw.text((x, y),"Sample Text",(r,g,b))
# j = width / 9
# for i in notes:
# draw.text((j, height-40), i, (0,0,0), font=font)
# j+= (width / (len(notes) + 4))
# layer.save("img/annotated.png")
# return render_template('result.html')
@app.route('/predict', methods = ['GET', 'POST'])
def predict():
if request.method == 'POST':
print("POST SUCCESS")
if 'file' not in request.files:
print("FILE DOES NOT EXIST")
return "No file part in request", 400
f = request.files['file']
print("READING FILE")
if f.filename == '':
print("No selected file")
return "No selected file", 400
if f and allowed_file(file.filename):
print("PREDICTING FILE")
f.save(os.path.join(app.config['UPLOAD_FOLDER'], f.filename))
img = Image.open(request.files['file'].stream).convert('RGB')
np_img = np.array(img)
cv_img = cv2.cvtColor(np_img, cv2.COLOR_RGB2BGR)
all_predictions = model.predict(cv_img)
generateWAV(all_predictions, "false")
print("PREDICTION COMPLETE")
return redirect(url_for('uploaded_file', filename=f.filename))
else:
print("FORMAT FAILURE")
else:
return 'EXIT'
import shutil
def compress(output_filename, dir_name):
shutil.make_archive(output_filename, 'zip', dir_name)
if __name__=="__main__":
compress('data/compressed/archive','data/melody')
#return fl.send_file(
#data,
#mimetype='application/zip',
#as_attachment=True,
#attachment_filename='data/compressed/archive.zip'
#)
#return redirect(url_for('uploaded_file', filename=f.filename))
```
| github_jupyter |
# 简单理解 RHF 含频极化率及其与 TD-HF 间的关系
> 创建日期:2020-01-02
>
> 最后修改:2020-06-10
含频极化率在非线性光学中有所应用。这里指的频率是入射激发光频率,而非分子振动频率。含频极化率英文是 Frequency-Dependent Polarizability,也有时使用动态 Dynamic 替代含频 Frequency-Dependent;相对地,没有入射激发光给出的极化率称为静态 Static 极化率。
但这也只是道听途说。对于我来讲更直接的意义会是,含频极化率对坐标的一阶导数可以用于计算含频 Raman 光谱。
写这篇文档一开始的原因是,曾经在尝试计算简单的 SERS 光谱时,发现 Valley, Schatz et al. [^Valley-Schatz.JPCL.2013] 的含频 Raman 光谱计算的文章提到使用 TD-DFT (time-dependent density functional theory);其它文献也几乎无一例外。这多少对我来说有点意外。Raman 光谱的计算通过求取极化率对简正坐标的导数 (不管是解析的还是数值的) 得到,而极化率则可以通过 CP-HF (coupled-perturbated Hartree-Fock) 方程给出。此前我确实地成功得到了 Gaussian 所给出的 RKS (GGA level) 极化率,并且并没有使用 TD-DFT 计算,而是 CP-KS (coupled-perturbated Kohn-Sham) 方程计算得到的极化率;我曾经一度以为这是 ADF 软件与 Gaussian 软件两者的区别。后来在各路同学的提醒下,才渐渐明白极化率与含时分析 (TD) 之间的关系。
这篇文档将会忽略大部分与公式推导有关的问题;这是由于 TD-DFT 或 TD-HF 的公式推导并不简单;在短时间之内我重复不出让我自己信服的推导。这篇文档不讨论与复数有关的话题,变量与公式全部采用实数与实函数。
我们会使用非对称的双氧水分子作为演示分子,基组使用 6-31G。计算程序使用 PySCF 提供电子积分,并与 Gaussian 的含频极化率、PySCF 的激发频率计算结果作对应。
```
%matplotlib notebook
import numpy as np
import scipy
from pyscf import gto, scf, tdscf
from functools import partial
import matplotlib.pyplot as plt
from matplotlib import patches
from formchk_interface import FormchkInterface
np.einsum = partial(np.einsum, optimize=["greedy", 1024 ** 3 * 2 / 8])
np.set_printoptions(5, linewidth=150, suppress=True)
```
全文使用以下记号:
- $p, q, r, s, m$ 表示全部轨道
- $i, j$ 表示占据分子轨道
- $a, b$ 表示非占分子轨道
- $\mu, \nu, \kappa, \lambda$ 表示原子轨道
- $t, s$ 在不引起歧义的情况下表示空间取向 $x, y, z$
- $P, Q, R, S$ 在这篇文档表示类似于 $ai$ 的组合下标
- $n$ 表示 TD-HF 激发态
全文使用简化与不严格的 Einstein Summation Convention。
下面补充一个原子单位能量 $E_\mathrm{h}$ 到波数 $\mathrm{cm}^{-1}$ 的换算 `Eh_cm`:
$$
1 \, E_\mathrm{h} = 219474.6 \, \mathrm{cm}^{-1}
$$
```
from scipy.constants import physical_constants
Eh_cm = physical_constants["hartree-inverse meter relationship"][0] / 100
Eh_cm
```
## 分子体系与标准结果
:::{admonition} 阅读提示
我们会花很长的时间进行分子体系与标准结果的定义。如果对代码与文本阅读能力有信心,这段可以跳过。
:::
### PySCF 体系定义
在进入下面的讨论前,我们先定义如下变量:
- `mol` PySCF 分子实例
```
mol = gto.Mole()
mol.atom = """
O 0.0 0.0 0.0
O 0.0 0.0 1.5
H 1.0 0.0 0.0
H 0.0 0.7 1.0
"""
mol.basis = "6-31G"
mol.verbose = 0
mol.build()
```
- `nao` 轨道数量 $n_\mathrm{AO}$, `nocc` 占据轨道数 $n_\mathrm{occ}$, `nvir` 非占轨道数 $n_\mathrm{vir}$
- `so` 占据轨道分割,`sv` 非占轨道分割,`sa` 全轨道分割
- `eri0_ao` 原子轨道基组双电子积分 ERI (electron repulsion integral)
$$
(\mu \nu | \kappa \lambda) = \int \phi_\mu (\boldsymbol{r}) \phi_\nu (\boldsymbol{r}) \frac{1}{|\boldsymbol{r} - \boldsymbol{r}'|} \phi_\kappa (\boldsymbol{r}') \phi_\lambda (\boldsymbol{r}') \, \mathrm{d} \boldsymbol{r} \, \mathrm{d} \boldsymbol{r}'
$$
- `d_ao` 偶极积分,其中下述的 $t$ 或后文会出现的 $s$ 表示偶极积分的方向 $x, y$ 或 $z$
$$
d_{\mu \nu}^t = - \langle \mu | t | \nu \rangle = \int \phi_\mu (\boldsymbol{r}) t \phi_\nu (\boldsymbol{r}) \, \mathrm{d} \boldsymbol{r}
$$
```
nao = nmo = mol.nao
nocc = mol.nelec[0]
nvir = nmo - nocc
so, sv, sa = slice(0, nocc), slice(nocc, nmo), slice(0, nmo)
eri0_ao = mol.intor("int2e")
d_ao = - mol.intor("int1e_r")
```
- `scf_eng` PySCF 的 RHF 计算实例
```
scf_eng = scf.RHF(mol).run()
```
- `C` $C_{\mu p}$ 分子轨道系数
- `e` $e_p$ RHF 轨道能量
- `D` $D_{\mu \nu} = 2 C_{\mu i} C_{\nu i}$ RHF 电子态密度
```
C, e = scf_eng.mo_coeff, scf_eng.mo_energy
D = 2 * C[:, so] @ C[:, so].T
```
- `eri0_mo` 分子轨道双电子 ERI $(pq|rs) = C_{\mu p} C_{\nu q} (\mu \nu | \kappa \lambda) C_{\kappa r} C_{\lambda s}$
- `d_mo` 分子轨道偶极积分 $d^t_{pq} = C_{\mu p} d^t_{\mu \nu} C_{\nu q}$
- `d_ia` 占据-非占的分子轨道偶极积分 $d^t_{ia}$
- `d_P` 以双下标 $P = ia$ 为记号的占据-非占分子轨道偶极积分 $d^t_P$
```
eri0_mo = np.einsum("up, vq, uvkl, kr, ls -> pqrs", C, C, eri0_ao, C, C)
d_mo = np.einsum("up, tuv, vq -> tpq", C, d_ao, C)
d_ia = d_mo[:, so, sv]
d_P = d_ia.reshape(3, nocc*nvir)
```
- `scf_td` PySCF 的 TD-RHF 计算实例
```
scf_td = tdscf.TDHF(scf_eng)
scf_td.nstates = nvir * nocc
scf_td.run()
```
就我目前所知,PySCF 可以计算极化率;但与含频极化率有关的计算,我在这里采用下文所述的、与 Gaussian 可以大致匹配结果的程序。
### Gaussian 计算含频极化率
我们需要一个可以核验结果的结果与工具。Gaussian 事实上提供了含频的极化率的选项。这一小段我们简单了解如何使用 Gaussian 来计算含频极化率。
我们首先给给出一个示范的例子。这个例子只是演示,并不能代表真实的物理。
Gaussian 的输入卡 {download}`assets/H2O2_freq_polar_example.gjf` 如下:
```
with open("assets/H2O2_freq_polar_example.gjf", "r") as f:
print(f.read()[:-1])
```
事实上这段程序不只计算了含频极化率,还计算了含频 Raman;但这份文档只讨论极化率问题。这里的“含频”指的是两个频率,其一为静态 (static) 极化率,其二是入射光线为 $\omega = 1 \, \mathrm{nm}$ 频率下的极化率。后者是一个相当极端的例子,因为通常可能没有人会想到用 X-Ray 照射一个普通的液体分子。
Gaussian 程序默认会输出 .out 或 .log 文件作为文本信息输出,其输出文件在 {download}`assets/H2O2_freq_polar_example.out` 中。我们还要求 Gaussian 输出 .chk 文件,该文件只包含单纯的计算数据信息,其通过 Gaussian Utility `formchk` 导出为 ASCII 格式的文件在 {download}`assets/H2O2_freq_polar_example.fch`。
对于 .out 文件,我们可以用下述命令仅查看含频极化率:
```
with open("assets/H2O2_freq_polar_example.out", "r") as f:
while f.readable():
line = f.readline()
if "Alpha(-w,w) frequency" in line:
print(line[:-1])
for _ in range(4):
print(f.readline()[:-1])
if "Beta(-w,w,0) frequency" in line:
break
```
对于每个频率,程序会给出一个 $3 \times 3$ 大小的矩阵;这就是极化率张量 $\alpha_{ts} (-\omega, \omega)$,其中 $t, s$ 可以是 $x, y, z$。以后的文档中,我们会简记 $\alpha_{ts} (-\omega, \omega)$ 为 $\alpha_{ts} (\omega)$。
极化率单位是原子单位;关于极化率原子单位与 SI 单位制的转换,参考下述 [NIST 网页](https://www.physics.nist.gov/cgi-bin/cuu/Value?auepol)。
上述出现的两个频率值 `0.000000`, `45.563353` 并不是以 $\mathrm{nm}$ 为单位,而是以 $E_\mathrm{h}$ Hartree 为单位。关于上述单位换算的过程,我们采用下述代码来说明:
```
1 / Eh_cm * 1e7
```
上面出现的 `Eh_cm` 已经在文档开头有所解释。这样我们就完成了 .out 文件的含频极化率的读取。
但在后面的文档中,我们将会利用 .chk 文件 (或者几乎等价的 .fch 文件) 的信息,来给出 Gaussian 计算的标准结果。在这份文档中,我们会利用到的键值有两段:`Frequencies for FD properties` 储存了以 $E_\mathrm{h}$ Hartree 为单位的频率 $\omega$,`Alpha(-w,w)` 储存了以原子单位储存的极化率张量 $\alpha_{ts} (\omega)$。
```
with open("assets/H2O2_freq_polar_example.fch", "r") as f:
print_flag = False
while f.readable():
line = f.readline()
if "Frequencies for FD properties" in line:
print_flag = True
if "Beta(-w,w,0)" in line:
break
if print_flag is True:
print(line[:-1])
```
我们可以看到,至少从程序输出的角度来讲,包含频率的极化率很可能与不含极化率的频率值相去甚远。当然,至于在 $1 \, \mathrm{nm}$ 如此高的光能量下,这个含频极化率是否在物理上正确,不是这篇文档讨论的问题。
通过我们导入的 `FormchkInterface` (取自 [pyxdh 项目](https://github.com/ajz34/Py_xDH/blob/master/pyxdh/Utilities/formchk_interface.py)),我们也可以对上述数值导入到 numpy 向量中。譬如我们需要提取所有频率,那么下面的代码就可以给出:
```
fchk_helper = FormchkInterface("assets/H2O2_freq_polar_example.fch")
fchk_helper.key_to_value("Frequencies for FD properties")
```
下面是 Gaussian 所给出的静态极化率 `ref_alpha_static` $\alpha_{ts} (-\omega, \omega)$。下一段我们会先回顾如何通过 CP-HF 方程,得到静态极化率。下述的矩阵将可以是我们计算结果的参考值。
```
ref_alpha_static = fchk_helper.key_to_value("Alpha(-w,w)")[:9].reshape(3, 3)
ref_alpha_static
```
再之后的段落,我们会需要计算含频极化率。Gaussian 一次性至多计算 100 个频率下的光学性质,否则程序会报错;因此我们的输入文件将会是多个文件,这里不列举其超链接。我们用 `freq_all_list` 表示含频极化率对应的频率,而 `alpha_all_list` 表示这些频率下的极化率。
```
freq_full_list = []
alpha_full_list = []
for idx in (1, 2, 3):
fchk_helper = FormchkInterface("assets/H2O2_freq_polar_{:1d}.fch".format(idx))
freq_full_list.append(fchk_helper.key_to_value("Frequencies for FD properties")[1:])
alpha_full_list.append(fchk_helper.key_to_value("Alpha(-w,w)").reshape(-1, 3, 3)[1:])
freq_full_list = np.concatenate(freq_full_list)
alpha_full_list = np.concatenate(alpha_full_list)
```
将所获得的含频极化率 (仅绘制其中一个分量 $\alpha_{zz} (\omega)$) 绘图可以得到:
```
fig, ax = plt.subplots()
ax.plot(freq_full_list, alpha_full_list[:, 2, 2])
rect = patches.Rectangle((0.184, -24), 0.01, 78, linewidth=1, edgecolor='C1', facecolor='C1', alpha=.25)
ax.add_patch(rect)
ax.set_ylim(-25, 75)
ax.set_xlabel(r"$\omega$ / $E_\mathrm{h}$")
ax.set_ylabel(r"$\alpha_{zz} (\omega)$ / a.u.")
ax.set_title("Frequency-Dependent Polarizability of $\mathrm{H_2O_2}$ (RHF/6-31G)")
fig.show()
```
含频极化率的图像在处于分子的激发态区域会呈剧烈振荡。在后续文档中,我们会先更多地看上图中橙色区域表示的前两个激发态。由于上图对橙色部分的描述不很精细,我们下面做一份更精细的极化率图绘制。
```
fchk_helper = FormchkInterface("assets/H2O2_freq_polar_small_range.fch")
freq_small_list = fchk_helper.key_to_value("Frequencies for FD properties")[1:]
alpha_small_list = fchk_helper.key_to_value("Alpha(-w,w)").reshape(-1, 3, 3)[1:]
fig, ax = plt.subplots()
ax.plot(freq_small_list, alpha_small_list[:, 2, 2])
ax.set_xlabel(r"$\omega$ / $E_\mathrm{h}$")
ax.set_ylabel(r"$\alpha_{zz} (\omega)$ / a.u.")
ax.set_title("Frequency-Dependent Polarizability of $\mathrm{H_2O_2}$ (RHF/6-31G)\nFor First Two Excited States")
fig.show()
```
从这两张图的纵坐标的缩放关系来看,事实上,对于每一个振荡峰,其振荡是趋于无穷大的;并且其对应的频率恰好是分子 TD-HF 计算得到的激发能。这将会在后面的文档中叙述并验证。
## TD-HF 方程过程回顾
### TD-HF 方程与激发能
一般会认为,TD 方法是用于求解与电子激发过程有关的方法。最常用的应用即是求解激发能与跃迁偶极矩。我们在这一段先回顾这两者的计算过程。
在进行后续的描述前,我们会定义下述与 TD-HF 方程有关的张量或矩阵 `A` $\mathbb{A}_{ia, jb}$ 与 `B` $\mathbb{B}_{ia, jb}$:
$$
\begin{align}
\mathbb{A}_{ia, jb} &= (\varepsilon_a - \varepsilon_i) \delta_{ij} \delta_{ab} + 2 (ia|jb) - (ij|ab) \\
\mathbb{B}_{ia, jb} &= 2 (ia|jb) - (ib|ja)
\end{align}
$$
其中两个辅助变量为:
- `delta_ij` $\delta_{ij}$ 为占据轨道数维度的单位矩阵
- `delta_ab` $\delta_{ab}$ 为非占轨道数维度的单位矩阵
```
delta_ij, delta_ab = np.eye(nocc), np.eye(nvir)
A_iajb = (
np.einsum("ia, ij, ab -> iajb", - e[so, None] + e[sv], delta_ij, delta_ab)
+ 2 * eri0_mo[so, sv, so, sv]
- eri0_mo[so, so, sv, sv].swapaxes(1, 2))
B_iajb = (
+ 2 * eri0_mo[so, sv, so, sv]
- eri0_mo[so, sv, so, sv].swapaxes(1, 3))
```
为了后文的代码方便,我们把双下标的矩阵记为 `A` $A_{PQ}$ 与 `B` $B_{PQ}$:
```
A = A_iajb.reshape(nocc*nvir, nocc*nvir)
B = B_iajb.reshape(nocc*nvir, nocc*nvir)
```
根据 TD-DFT 中的 Casida 方程 (TD-DFT 可以看作是 TD-HF 情形的扩展),我们可以写出 TD-HF 的频率及其对应的本征向量为 $\omega_n, X_{ia}^n, Y_{ia}^n$,或者双下标记号的 $X_{P}^n, Y_{P}^n$。其中,$X_{ia}^n$ 有时称为第 $n$ 个激发态的激发矩阵,$Y_{ia}^n$ 则称退激发矩阵。这几者之间满足下述 TD-HF 矩阵方程。
$$
\begin{pmatrix} \mathbb{A} & \mathbb{B} \\ - \mathbb{B} & - \mathbb{A} \end{pmatrix}
\begin{pmatrix} \mathbf{X}^n \\ \mathbf{Y}^n \end{pmatrix}
= \omega_n \begin{pmatrix} \mathbf{X}^n \\ \mathbf{Y}^n \end{pmatrix}
$$
我们在程序上,记等式左边的大矩阵为 `AB`。
```
AB = np.block([
[ A, B],
[- B, - A]
])
AB.shape
```
我们首先解出上述矩阵的本征值 `eigs` 与本征向量 `xys`:
```
eigs, xys = np.linalg.eig(AB)
```
但我们会发现,我们本来预期的在 6-31G 基组下可解的激发态数量只有 $n_\mathrm{occ} n_\mathrm{vir} = 117$,但本征值数量却是 234 个。我们需要舍去所有负值的本征值。事实上,负值的本征值与正值的本征值之间有一一对应的关系。
```
(eigs < 0).sum()
```
我们舍去负本征值及其对应的本征向量,并对本征值作排序,得到正的本征值 `eigs_sorted` 及其相对应的本征向量 `xys_sorted`:
```
eigs_sorted = eigs[eigs.argsort()[int(eigs.size / 2):]]
xys_sorted = xys[:, eigs.argsort()[int(eigs.size / 2):]]
```
我们应当可以验证,上述本征值与本征向量确实满足 TD-HF 矩阵方程:
```
np.allclose(AB @ xys_sorted, eigs_sorted * xys_sorted)
```
最后,我们用 `td_eig` $\omega_n$、`td_x_unnormed` 未归一化的 $X^n_P$、`td_y_unnormed` 未归一化的 $Y^n_P$ 来重新整理上述的结果 `eigs_sorted` 与 `xys_sorted`。需要注意,变量 `td_x_unnormed` 的两个维度中,第一个维度为激发态 $n$,第二个维度为双下标 $P = ia$;尽管两个维度的大小都是 $n_\mathrm{occ} n_\mathrm{vir} = 117$,但意义完全不同。
```
td_eig = eigs_sorted
td_x_unnormed = xys_sorted.T[:, :nvir*nocc]
td_y_unnormed = xys_sorted.T[:, nvir*nocc:]
```
我们简单看一下最低激发能的几个激发态的能级大小,单位是原子单位或 Hartree $E_\mathrm{h}$:
```
eigs_sorted[:10]
```
我们能看到最低的激发态中,有 0.187 与 0.191;这恰好与上面 Gaussian 绘制出来的含频极化率图中的两个振荡峰位置恰好吻合。这并非是偶然,并且我们会在后文进行更详细的描述。
### TD-HF 跃迁偶极矩
从基态波函数 $| 0 \rangle$ 到激发态波函数 $| n \rangle$ 的跃迁偶极矩可以写作 $\langle 0 | \hat d{}^t | n \rangle$ 或等价的 $- \langle 0 | t | n \rangle$;留意到 $t \in \{ x, y, z \}$。但实施上,我们尚不能写出激发态波函数 $| n \rangle$ 的具体形式。这个激发态波函数需要通过激发矩阵 $X_{ia}^n$ 与退激发矩阵 $Y_{ia}^n$ 来描述。
刚才的计算中,我们得到的本征向量是未经归一化的;它乘以任何非零常数,仍然会是 TD-HF 矩阵方程的本征向量。但我们可以使用激发与退激发,赋予这个本征向量以物理含义。其归一化条件是,态 $| n \rangle$ 的电子数守恒,即与 $| 0 \rangle$ 的电子数相同。在 RHF 问题下,这要求
$$
(X_{ia}^n)^2 - (Y_{ia}^n)^2 = 2
$$
我们令归一化过程中的中间量为 `td_renorm` $N_n = \frac{1}{2} \left( (X_{ia}^n)^2 - (Y_{ia}^n)^2 \right)$:
```
td_renorm = ((td_x_unnormed**2).sum(axis=1) - (td_y_unnormed**2).sum(axis=1)) / 2
```
那么重新归一化后的 `X` $X_P^n$ 与 `Y` $Y_P^n$ 为
```
X = td_x_unnormed / np.sqrt(td_renorm)[:, None]
Y = td_y_unnormed / np.sqrt(td_renorm)[:, None]
```
为了处理一些问题的便利,我们声明变量 `X_ia` $X_{ia}^n$ 与 `Y_ia` $Y_{ia}^n$;它们的维度均是 $(n, i, a)$:
```
X_ia = X.reshape(nocc*nvir, nocc, nvir)
Y_ia = Y.reshape(nocc*nvir, nocc, nvir)
X_ia.shape
```
以此为基础,我们可以写出 TD-HF 的跃迁偶极矩 `td_transdip`
$$
\langle 0 | \hat d{}^t | n \rangle = d_{ia}^t (X_{ia}^n + Y_{ia}^n)
$$
我们会打印出最低能级的 5 个激发态的跃迁偶极矩:
```
td_transdip = np.einsum("tia, nia -> nt", d_ia, X_ia + Y_ia)
td_transdip[:5]
```
这与 PySCF 所给出的跃迁偶极矩几乎是相同的,但符号上会有差异。我们认为这已经完整并成功地重复了跃迁偶极矩了。
```
scf_td.transition_dipole()[:5]
```
需要注意,这可能与 Gaussian 计算得到的跃迁偶极矩的值接近但并不完全相等。这可能与 Gaussian 默认的 TD-HF 精度偏低有关。
## 静态极化率
### 偶极微扰下的 CP-HF 方程
这篇文档的一个目的是将 CP-HF 方程与 TD-HF 方程之间的关系作一个联系。因此,我们需要首先了解 CP-HF 方程在静态极化率中的工作过程。
:::{attention}
尽管我们确实可以用后续文档中的代码或公式计算得到一些结果,但这并不意味着成型的量化软件也使用这些算法。譬如对于 RHF 下静态极化率计算,通常更高效的做法是使用类似于 CP-HF 方程的 Z-Vector 方程。
:::
由此,我们会写偶极微扰下的 CP-HF 方程为
$$
A'_{ia, jb} U^t_{jb} = d^t_{ia}
$$
下面简单但不严谨地回顾 CP-HF 方程的推导思路。我们在分子体系上,外加一个微扰偶极场,其大小是分子轨道基组下的 $d_{pq}^t$ 偶极矩阵,微扰哈密顿量为 $t$ (即单位方向为 $t$ 的电场微扰)。根据 RHF 的变分条件,任何外加微扰的哈密顿量 $t$ 都应该满足
$$
\frac{\partial F_{pq}}{\partial t} = 0
$$
通过该式,几乎可以直接得到 CP-HF 方程。方程的左边定义上是偶极积分,右边的 `A_p` $A'_{ia, jb}$ 为
$$
A'_{ia, jb} = (\varepsilon_a - \varepsilon_i) \delta_{ij} \delta_{ab} + 4 (ia|jb) - (ij|ab) - (ib|ja)
$$
而 `U_ia` $U_{jb}^t$ 称为 U 矩阵,表示的是与电子态密度在外加偶极微扰影响下的变化有关的量;一种导出式如下:
$$
\frac{\partial D_{pq}}{\partial t} = D_{pm} U^t_{mq} + D_{mq} U^t_{mp}
$$
因此,CP-HF 的一种直观的解释思路是,它求取的是分子受到外加偶极的微扰 $d^t_{ia}$ 下,电子态密度形变的大小,而这个大小是由 $U_{jb}^t$ U 矩阵刻画的。很容易想到的性质是,若外加偶极微扰趋于零,那么外加的形变微扰也趋于零矩阵。
下面我们来求取 CP-HF 方程,给出 `A_p` $A'_{PQ} = A'_{ia, jb}$ 与 `U_ia` $U_{jb}^t$。需要注意在这份文档中,角标顺序是 $ia, jb$ 而非 $ai, bj$;这可能与其它课本或文档的顺序不太相同,在一些矩阵的正负号上也可能存在差异。
```
A_p = (
+ np.einsum("ia, ij, ab -> iajb", - e[so, None] + e[sv], delta_ij, delta_ab)
+ 4 * eri0_mo[so, sv, so, sv]
- eri0_mo[so, so, sv, sv].swapaxes(1, 2)
- eri0_mo[so, sv, so, sv].swapaxes(1, 3)
).reshape(nvir*nocc, nvir*nocc)
U_ia = np.einsum("PQ, sQ -> sP", np.linalg.inv(A_p), d_P)
U_ia.shape = (3, nocc, nvir)
```
随后,根据求导法则与矩阵的对称性、反对称性的应用,应当可以得到静态极化率表达式为
$$
\alpha_{ts} (0) = \frac{\partial^2 E_\mathrm{RHF}}{\partial t \partial s} = \frac{\partial D_{ij} d^t_{ij} \delta_{ij}}{\partial s} = 4 d^t_{ia} U^s_{ia}
$$
```
4 * np.einsum("tia, sia -> ts", d_ia, U_ia)
```
上述的结果与 Gaussian 计算所得到的静态极化率 `ref_alpha_static` 完全一致。
```
np.allclose(
4 * np.einsum("tia, sia -> ts", d_ia, U_ia),
ref_alpha_static)
```
### 矩阵求逆直接获得静态极化率
根据我们所写的 CP-HF 方程
$$
A'_{ia, jb} U^t_{jb} = d^t_{ia}
$$
我们应当很容易地想到,如果我们有足够的计算能力,可以对四脚标矩阵 $A'_{ia, jb}$ 求逆,那么我们不一定需要明确写出 U 矩阵,也一样可以求得静态极化率:
$$
\alpha_{ts} (0) = 4 d^t_{ia} (A'{}^{-1})_{ia, jb} d^s_{ia}
$$
当然,上面的计算过程实际上是用双下标 ($P = ia$, $Q = jb$) 表达式实现的:
$$
\alpha_{ts} (0) = 4 d^t_{P} (A'{}^{-1})_{PQ} d^s_{Q}
$$
```
np.allclose(4 * np.einsum("tP, PQ, sQ -> ts", d_P, np.linalg.inv(A_p), d_P), ref_alpha_static)
```
### 跃迁偶极矩获得静态极化率
CP-HF 方程求解静态极化率的思路是非常直观的;但静态极化率还可以通过 TD-HF 的方式求得。表面上,这两种推导思路和前提几乎完全不同;但我们却可以得到数值上 **完全** (而非近似) 相等的静态极化率。这里我们会作说明。
我们首先不加说明地直接给出 TD-HF 方式给出的静态极化率公式:
$$
\alpha_{ts} (0) = 2 \frac{\langle 0 | \hat d{}^t | n \rangle \langle n | \hat d{}^s | 0 \rangle}{\omega_n}
$$
它很容易化为程序表达式:
```
2 * np.einsum("nt, n, ns -> ts", td_transdip, 1 / td_eig, td_transdip)
```
我们与上面用 CP-HF 方式计算得到的静态极化率作对比,不难发现两者的值是完全相等的。可以用下述程序与 Gaussian 的计算结果作对比:
```
np.allclose(2 * np.einsum("nt, n, ns -> ts", td_transdip, 1 / td_eig, td_transdip), ref_alpha_static)
```
这说明,对于静态极化率问题,TD-HF 与 CP-HF 方法之间有着确实的联系。我们下面就使用 TD-HF 方程来导出 CP-HF 方程的结果,或者说从简单的线性代数角度证明:
$$
\alpha_{ts} (0) = 2 \frac{\langle 0 | \hat d{}^t | n \rangle \langle n | \hat d{}^s | 0 \rangle}{\omega_n} = 4 d^t_{P} (A'{}^{-1})_{PQ} d^s_{Q}
$$
### 静态极化率下 TD-HF 方程与 CP-HF 方程的等价推导
这里我们会给出静态情况下,TD-HF 与 CP-HF 方程的推演过程。对于动态 (含频) 过程的推演,我们会放在文档的后面描述。
首先,我们会说明 TD-HF 方程的 `A` $\mathbb{A}_{PQ}$ 与 `B` $\mathbb{B}_{PQ}$ 之和,恰好是 CP-HF 方程的 `A_p` $A'_{PQ}$:
$$
A'_{PQ} = \mathbb{A}_{PQ} + \mathbb{B}_{PQ}
$$
```
np.allclose(A + B, A_p)
```
利用 $\langle 0 | \hat d{}^t | n \rangle = d_{ia}^t (X_{ia}^n + Y_{ia}^n)$,我们重新用 $X_P^n, Y_P^n$ 的形式写一下 TD-HF 方程所给出的极化率公式:
$$
\alpha_{ts} (0) = 2 \frac{d_P^t d_Q^s (X_P^n + Y_P^n) (X_Q^n + Y_Q^n)}{\omega_n}
$$
```
np.allclose(2 * np.einsum("tP, sQ, nP, nQ, n -> ts", d_P, d_P, X + Y, X + Y, 1 / td_eig), ref_alpha_static)
```
回顾到 TD-HF 方程
$$
\begin{pmatrix} \mathbb{A} & \mathbb{B} \\ - \mathbb{B} & - \mathbb{A} \end{pmatrix}
\begin{pmatrix} \mathbf{X}^n \\ \mathbf{Y}^n \end{pmatrix}
= \omega_n \begin{pmatrix} \mathbf{X}^n \\ \mathbf{Y}^n \end{pmatrix}
$$
我们可以推知,$(\mathbb{A} + \mathbb{B}) (\mathbf{X}^n + \mathbf{Y}^n) = \omega_n (\mathbf{X}^n - \mathbf{Y}^n)$,或者写为角标求和的形式,
$$
(\mathbb{A} + \mathbb{B})_{RQ} (\mathbf{X}^n + \mathbf{Y}^n)_Q = \omega_n (\mathbf{X}^n - \mathbf{Y}^n)_R
$$
那么我们可以将上式,以及矩阵逆关系 $(\mathbb{A} + \mathbb{B})^{-1} (\mathbb{A} + \mathbb{B}) = \mathbb{1}$ 即
$$
(\mathbb{A} + \mathbb{B})^{-1}_{SR} (\mathbb{A} + \mathbb{B})_{RQ} = \delta_{SQ}
$$
代入到上面提到的 TD-HF 的极化率公式中,得到
$$
\begin{align}
\alpha_{ts} (0)
&= 2 \frac{d_P^t d_Q^s (X_P^n + Y_P^n) (\mathbb{A} + \mathbb{B})^{-1}_{QR} (\mathbb{A} + \mathbb{B})_{RQ} (X_Q^n + Y_Q^n)}{\omega_n} \\
&= 2 d_P^t d_Q^s (X_P^n + Y_P^n) (\mathbb{A} + \mathbb{B})^{-1}_{QR} (X_R^n - Y_R^n)
\end{align}
$$
随后我们需要利用 $\mathbf{X}^n, \mathbf{Y}^n$ 的正交化条件。正交化条件我们曾经在给出 $\mathbf{X}^n, \mathbf{Y}^n$ 时确实利用到过,但其更有用的推论是 $(\mathbf{X} + \mathbf{Y})^\dagger (\mathbf{X} - \mathbf{Y}) = 2 \cdot \mathbb{1}$
$$
(\mathbf{X}^n + \mathbf{Y}^n)_P (\mathbf{X}^n - \mathbf{Y}^n)_R = 2 \delta_{PR}
$$
我们可以用下面的程序来说明这一问题:
```
np.einsum("nP, nQ -> PQ", X + Y, X - Y)
```
那么我们就可以将上面的极化率公式化为
$$
\begin{align}
\alpha_{ts} (0)
&= 2 d_P^t d_Q^s (\mathbb{A} + \mathbb{B})^{-1}_{QR} \cdot 2 \delta_{PR} \\
&= 4 d_Q^s (\mathbb{A} + \mathbb{B})^{-1}_{QP} d_P^t \\
&= 4 d_Q^s (A'{}^{-1})_{QP} d^t_P
\end{align}
$$
我们知道,上式的等式左边是对 $Q, P$ 双下标进行求和。作为被求和的两个下标是可以被交换的,因此我们可以将上式写为
$$
\alpha_{ts} (0) = 4 d_P^s (A'{}^{-1})_{PQ} d^t_Q
$$
```
np.allclose(4 * np.einsum("sP, PQ, tQ -> ts", d_P, np.linalg.inv(A_p), d_P), ref_alpha_static)
```
上述推导并没有结束。我们回顾到刚才 CP-HF 所给出的极化率并不是上述的表达式,而是交换了 $t, s$ 两者的极化率:
$$
\alpha_{ts} (0) = 4 d_P^t (A'{}^{-1})_{PQ} d^s_Q
$$
```
np.allclose(4 * np.einsum("tP, PQ, sQ -> ts", d_P, np.linalg.inv(A_p), d_P), ref_alpha_static)
```
从极化率作为能量的二阶梯度的角度来说,这是因为被求导量可交换,因此极化率具有 Hermite 性质:
$$
\alpha_{ts} (\omega) = \alpha_{st} (\omega)
$$
到这一步为止,从 TD-HF 方程给出的极化率,成功地推导出了 CP-HF 方程所给出的极化率。
## 含频极化率
### 跃迁偶极矩获得含频极化率
我们先不对含频极化率作公式上的分析,先只看数值的结果,并与 Gaussian 输出的结果进行比较。
我们仍然不加说明地直接给出 TD-HF 方式给出的含频极化率公式:
$$
\alpha_{ts} (\omega_n) = \frac{\langle 0 | \hat d{}^t | n \rangle \langle n | \hat d{}^s | 0 \rangle}{\omega_n - \omega} + \frac{\langle 0 | \hat d{}^t | n \rangle \langle n | \hat d{}^s | 0 \rangle}{\omega_n + \omega}
$$
需要留意的是,$\omega_n$ 为分子通过 TD-HF 方程解出来的激发能,而 $\omega$ 是外加的、任意的激发光束频率;两者除了单位一致外几乎完全无关。
上述公式中,前一项称为共振项 (resonance term),后一项称为非共振项。这两项在不同频率下的行为可以很容易地用图片表示出来;当 $\omega$ 接近激发频率 $\omega_n$ 时产生断点行为的项是共振项。
它也很容易化为程序表达式。我们用下述函数 `freq_to_alpha`,输入 `omega` $\omega$ 来返回含频频率 $\alpha_{ts} (\omega_n)$;并将其中的共振项与非共振项拆分为函数 `freq_to_res` 与 `freq_to_nonres`:
```
freq_to_res = lambda omega: np.einsum("nt, n, ns -> ts", td_transdip, 1 / (td_eig - omega), td_transdip)
freq_to_nonres = lambda omega: np.einsum("nt, n, ns -> ts", td_transdip, 1 / (td_eig + omega), td_transdip)
freq_to_alpha = lambda omega: freq_to_res(omega) + freq_to_nonres(omega)
```
若外加激发光束频率为 0,那么将退化到静态极化率的情形中:
```
freq_to_alpha(0)
np.allclose(freq_to_alpha(0), ref_alpha_static)
```
在静态情况下,共振项与非共振项对总极化率的贡献是相等的:
```
freq_to_res(0)
```
我们现在想要尝试与 Gaussian 的数据进行核对并绘制图片。回顾到我们曾经定义过 `freq_full_list` 为较广的频率范围,其对应的 Gaussian 计算的极化率在 `alpha_full_list`。我们将用 `freq_to_alpha` 函数绘制的 $\alpha_{zz} (\omega)$ 极化率分量放在 numpy 张量列表 `alpha_zz_full_calc` 中;并将其共振项与非共振项分别放在 `alpha_zz_full_res` 与 `alpha_zz_full_nonres`:
```
alpha_zz_full_calc = np.vectorize(lambda omega: freq_to_alpha (omega)[2, 2])(freq_full_list)
alpha_zz_full_res = np.vectorize(lambda omega: freq_to_res (omega)[2, 2])(freq_full_list)
alpha_zz_full_nonres = np.vectorize(lambda omega: freq_to_nonres(omega)[2, 2])(freq_full_list)
```
下面我们绘制在这个频率区间内,Gaussian 计算得到的结果与我们用上面跃迁偶极矩的公式获得的结果作比较。尽管在断点附近两者表现略有不同,但稳定区间的含频极化率与 Gaussian 的结果基本上是一致的。
```
fig, ax = plt.subplots()
ax.plot(freq_full_list, alpha_full_list[:, 2, 2], label="Gaussian")
ax.plot(freq_full_list, alpha_zz_full_res, linestyle="-.", c="C2", label="Resonance")
ax.plot(freq_full_list, alpha_zz_full_nonres, linestyle="-.", c="C3", label="Non-Resonance")
ax.plot(freq_full_list, alpha_zz_full_calc, linestyle=":", label="Calculated")
rect = patches.Rectangle((0.184, -24), 0.01, 78, linewidth=1, edgecolor='C4', facecolor='C4', alpha=.25)
ax.add_patch(rect)
ax.set_ylim(-25, 75)
ax.set_xlabel(r"$\omega$ / $E_\mathrm{h}$")
ax.set_ylabel(r"$\alpha_{zz} (\omega)$ / a.u.")
ax.set_title("Frequency-Dependent Polarizability of $\mathrm{H_2O_2}$ (RHF/6-31G)")
ax.legend()
fig.show()
```
对于前两个激发态能量的窄区间中,我们的结果在非断点附近其实与 Gaussian 的结果也基本一致:
```
alpha_zz_small_calc = np.vectorize(lambda omega: freq_to_alpha (omega)[2, 2])(freq_small_list)
alpha_zz_small_res = np.vectorize(lambda omega: freq_to_res (omega)[2, 2])(freq_small_list)
alpha_zz_small_nonres = np.vectorize(lambda omega: freq_to_nonres(omega)[2, 2])(freq_small_list)
fig, ax = plt.subplots()
ax.plot(freq_small_list, alpha_small_list[:, 2, 2], label="Gaussian")
ax.plot(freq_small_list, alpha_zz_small_res, linestyle="-.", c="C2", label="Resonance")
ax.plot(freq_small_list, alpha_zz_small_nonres, linestyle="-.", c="C3", label="Non-Resonance")
ax.plot(freq_small_list, alpha_zz_small_calc, linestyle=":", label="Calculated")
ax.set_xlabel(r"$\omega$ / $E_\mathrm{h}$")
ax.set_ylabel(r"$\alpha_{zz} (\omega)$ / a.u.")
ax.set_title("Frequency-Dependent Polarizability of $\mathrm{H_2O_2}$ (RHF/6-31G)\nFor First Two Excited States")
ax.legend()
fig.show()
```
我们认为我们确实正确计算了含频极化率。在断点附近的行为应当被认为是数值上的微小差别;并且我们认为,在断点 (共振) 处附近产生的极化率的主要贡献部分应为极化率的共振项所产生。
### TD-HF 方程含频极化率及其与跃迁偶极矩的关联
上一大段中,我们仅仅是用了 TD-HF 给出的跃迁偶极矩结果,反推出了 CP-HF 的公式。但我们并没有介绍过最原始的 TD-HF 极化率表达式。下面我们会从最普遍的公式,推导含频极化率的表达式。下面的推导过程中,程序的部分会少一些。
我们从 TD-DFT 的 Casida 方程开始;Casida 方程可以简单地退化到 TD-HF 的情形。我们上面在推演激发频率 $\omega_n$ 时,也提到了 Casida 方程;
$$
\begin{align}
\begin{pmatrix} \mathbb{A} & \mathbb{B} \\ - \mathbb{B} & - \mathbb{A} \end{pmatrix}
\begin{pmatrix} \mathbf{X}^n \\ \mathbf{Y}^n \end{pmatrix}
= \omega_n \begin{pmatrix} \mathbf{X}^n \\ \mathbf{Y}^n \end{pmatrix}
\tag{1}
\end{align}
$$
但下面的 Casida 方程具有更为广泛的适用情形。我们引入外加的偶极微扰 `d_P` $\mathbf{d}^t$ 与外加激发光束频率 `omega` $\omega$ 的微扰,则有
$$
\begin{align}
\begin{pmatrix} \mathbb{A} & \mathbb{B} \\ \mathbb{B} & \mathbb{A} \end{pmatrix}
\begin{pmatrix} \mathbf{X}'{}^t \\ \mathbf{Y}'{}^t \end{pmatrix}
= \omega \begin{pmatrix} \mathbf{X}'{}^t \\ - \mathbf{Y}'{}^t \end{pmatrix} +
\begin{pmatrix} 2 \mathbf{d}^t \\ 2 \mathbf{d}^t \end{pmatrix}
\tag{2}
\end{align}
$$
这里有不少符号上的区别。首先,(1) 式的激发态频率 $\omega_n$ 与 (2) 式的外加光束的频率 $\omega$ 并不相同;但 (2) 在一种情形下确实地可以退化到 (1) 式。若现在没有外加偶极微扰 $\mathbf{d}^t$,那么外加光束必须要恰好处于分子电子的激发频率上,分子的电子云微扰变化才能被允许 (即使时间非常短,这对应的是紫外光谱电子态的平均寿命)。而电子的激发频率不一定只有一个,因此会产生上下标 $n$ 表示不同的激发频率;第 $n$ 个激发态电子云微扰的形变大小和取向由 $\mathbf{X}^n$ 与 $\mathbf{Y}^n$ 共同决定;它们将会产生第 $n$ 个激发态的跃迁密度 $\rho^n (\boldsymbol{r}, \omega)$ (下式对 $i, a$ 求和):
$$
\rho^n (\boldsymbol{r}, \omega_n) = (X_{ia}^n + Y_{ia}^n) \phi_i (\boldsymbol{r}) \phi_a (\boldsymbol{r})
$$
其次,(2) 式的 $\mathbf{X}'{}^t$ 若不看偶极激发方向 $t$,它还比 (1) 式的 $\mathbf{X}^n$ 少了 $n$ 并多了一撇;多的一撇是为了区分两者。之所以这里没有 $n$,我们可以这样考虑:在某一个特定的外加偶极微扰 $\mathbf{d}^t$ 与频率 $\omega$ 微扰下,分子的电子云确实会发生改变;但这种改变的方式一般是唯一的 (简并情况我们不作讨论)。这种形变所产生的密度形式也是类似的 (下式对 $i, a$ 求和):
$$
\rho (\boldsymbol{r}, \mathbf{d}^t, \omega) = (X_{ia}'{}^t + Y_{ia}'{}^t) \phi_i (\boldsymbol{r}) \phi_a (\boldsymbol{r})
$$
含频极化率可以通过上述的形变密度在偶极算符的作用下给出:
$$
\alpha_{ts} (\omega)
= \int -s \cdot \rho (\boldsymbol{r}, \mathbf{d}^t, \omega) \, \mathrm{d} \boldsymbol{r}
= (X_{ia}'{}^t + Y_{ia}'{}^t) \int -s \cdot \phi_i (\boldsymbol{r}) \phi_a (\boldsymbol{r}) \, \mathrm{d} \boldsymbol{r}
= (X_{ia}'{}^t + Y_{ia}'{}^t) d_{ia}^s = (X_P'{}^t + Y_P'{}^t) d_P^s
$$
其中关于 $X_P'{}^t, Y_P'{}^t$ 需要通过 Casida 方程求取。(2) 式经过简单的代数处理后得到 (3) 式:
$$
\begin{align}
\begin{pmatrix} \mathbb{A} - \omega \mathbb{1} & \mathbb{B} \\ - \mathbb{B} & - \mathbb{A} - \omega \mathbb{1} \end{pmatrix}
\begin{pmatrix} \mathbf{X}'{}^t \\ \mathbf{Y}'{}^t \end{pmatrix}
= \begin{pmatrix} 2 \mathbf{d}^t \\ - 2 \mathbf{d}^t \end{pmatrix}
\tag{3}
\end{align}
$$
那么我们有
$$
\begin{align}
\alpha_{ts} (\omega) = (X_P'{}^t + Y_P'{}^t) d_P^s
= \begin{pmatrix} \mathbf{d}^s & \mathbf{d}^s \end{pmatrix} \begin{pmatrix} \mathbf{X}'{}^t \\ \mathbf{Y}'{}^t \end{pmatrix}
= \begin{pmatrix} \mathbf{d}^s & \mathbf{d}^s \end{pmatrix}
\begin{pmatrix} \mathbb{A} - \omega \mathbb{1} & \mathbb{B} \\ - \mathbb{B} & - \mathbb{A} - \omega \mathbb{1} \end{pmatrix}^{-1}
\begin{pmatrix} 2 \mathbf{d}^t \\ - 2 \mathbf{d}^t \end{pmatrix}
\tag{4}
\end{align}
$$
对于不带频率的情形,我们可以用下面的代码验证:
```
np.einsum("tP, PQ, sQ -> ts",
np.concatenate([d_P, d_P], axis=1),
np.linalg.inv(AB),
np.concatenate([2 * d_P, - 2 * d_P], axis=1))
```
而对于带频率的情形,我们可以举一个 $\omega = 0.186 \, E_\mathrm{h}$ 的例子:
```
omega = 0.186
np.einsum("tP, PQ, sQ -> ts",
np.concatenate([d_P, d_P], axis=1),
np.linalg.inv(AB - np.eye(nvir*nocc*2) * omega),
np.concatenate([2 * d_P, - 2 * d_P], axis=1))
```
但这样的表达式 (4) 并没有出现跃迁偶极矩。下面我们需要简化表达式。
首先,我们回顾式 (3),得到方程组
$$
\begin{align}
\mathbb{A} \mathbf{X}'{}^t + \mathbb{B} \mathbf{Y}'{}^t - \omega \mathbf{X}'{}^t &= 2 \mathbf{d}^t \\
\mathbb{B} \mathbf{X}'{}^t + \mathbb{A} \mathbf{Y}'{}^t + \omega \mathbf{Y}'{}^t &= 2 \mathbf{d}^t
\end{align}
$$
两式加减后,可以得到
$$
\begin{align}
(\mathbb{A} + \mathbb{B}) (\mathbf{X}'{}^t + \mathbf{Y}'{}^t) - \omega (\mathbf{X}'{}^t - \mathbf{Y}'{}^t) &= 4 \mathbf{d}^t \tag{5} \\
(\mathbb{A} - \mathbb{B}) (\mathbf{X}'{}^t - \mathbf{Y}'{}^t) &= \omega (\mathbf{X}'{}^t + \mathbf{Y}'{}^t) \tag{6}
\end{align}
$$
利用 (6) 式替换 (5) 式中出现的 $(\mathbf{X}'{}^t - \mathbf{Y}'{}^t)$,有
$$
(\mathbb{A} - \mathbb{B}) (\mathbb{A} + \mathbb{B}) (\mathbf{X}'{}^t + \mathbf{Y}'{}^t) - \omega^2 (\mathbf{X}'{}^t + \mathbf{Y}'{}^t) = 4 (\mathbb{A} - \mathbb{B}) \mathbf{d}^t
$$
或者,等价地,
$$
(\mathbf{X}'{}^t + \mathbf{Y}'{}^t) = 4 \left( (\mathbb{A} - \mathbb{B}) (\mathbb{A} + \mathbb{B}) - \omega^2 \mathbb{1} \right)^{-1} (\mathbb{A} - \mathbb{B}) \mathbf{d}^t
$$
两边再乘上 $\mathbf{d}^s$,就得到了含频极化率:
$$
\begin{align}
\alpha_{ts} (\omega) = 4 \mathbf{d}^s{}^\dagger \left( (\mathbb{A} - \mathbb{B}) (\mathbb{A} + \mathbb{B}) - \omega^2 \mathbb{1} \right)^{-1} (\mathbb{A} - \mathbb{B}) \mathbf{d}^t \tag{7}
\end{align}
$$
以 $\omega = 0.186 \, E_\mathrm{h}$ 来表达上式,则有
```
4 * np.einsum("tP, PR, RQ, sQ -> ts", d_P, np.linalg.inv((A - B) @ (A + B) - omega**2 * np.eye(nvir*nocc)), A - B, d_P)
```
上述结果与通过式 (4) 给出的结果一致。
下面我们引入一个技巧。我们将会对矩阵 $(\mathbb{A} - \mathbb{B}) (\mathbb{A} + \mathbb{B}) - \omega^2 \mathbb{1}$ 作逆矩阵分解。我们采用的做法是分析矩阵的特征值与特征向量。依据我们对式 (1) 形式的 Casida 方程的讨论,我们应当容易推知
$$
\begin{align}
(\mathbb{A} - \mathbb{B}) (\mathbb{A} + \mathbb{B}) (\mathbf{X}^n + \mathbf{Y}^n) = \omega_n^2 (\mathbf{X}^n + \mathbf{Y}^n) \tag{8}
\end{align}
$$
```
np.allclose((A - B) @ (A + B) @ (X + Y).T, td_eig**2 * (X + Y).T)
```
我们曾经指出过正交条件为 $(\mathbf{X} + \mathbf{Y})^\dagger (\mathbf{X} - \mathbf{Y}) = 2 \cdot \mathbb{1}$,因此,若将 $\mathbf{X}, \mathbf{Y}$ 分别看作是横向维度 $n$ 表示激发态,纵向维度 $P$ 表示激发和退激发分量的方形矩阵:
$$
\mathbf{X} = \begin{pmatrix} \mathbf{X}^1{}^\dagger \\ \mathbf{X}^2{}^\dagger \\ \vdots \\ \mathbf{X}^{n_\mathrm{occ} n_\mathrm{vir}}{}^\dagger \end{pmatrix}
$$
那么 $(\mathbf{X} + \mathbf{Y})$ 与 $(\mathbf{X} - \mathbf{Y})$ 之间存在互逆关系:
$$
(\mathbf{X} + \mathbf{Y})^{-1} = \frac{1}{2} (\mathbf{X} - \mathbf{Y})^\dagger
$$
```
np.allclose(np.linalg.inv(X + Y), 0.5 * (X - Y).T)
```
因此,一定程度上,我们可以认为 $(\mathbf{X} + \mathbf{Y})$ 与 $(\mathbf{X} - \mathbf{Y})$ 两者是相互对偶的向量组。注意区别这里的 $\mathbf{X}$ 引入的目的是为了刻画 $\mathbb{A}, \mathbb{B}$ 的性质,而与 $\mathbf{X}'{}^t$ 没有直接关联。
这种向量组满足下述特征向量分解公式:
$$
(\mathbb{A} - \mathbb{B}) (\mathbb{A} + \mathbb{B}) = \frac{1}{2} (\mathbf{X} + \mathbf{Y}) \mathbf{\Omega}^2 (\mathbf{X} - \mathbf{Y})^\dagger
$$
其中,
$$
\mathbf{\Omega} =
\begin{pmatrix}
\omega_1 & 0 & \cdots & 0 \\
0 & \omega_2 & \cdots & 0 \\
\vdots & \vdots & \ddots & \vdots \\
0 & 0 & \cdots & \omega_{n_\mathrm{occ} n_\mathrm{vir}}
\end{pmatrix}
$$
```
np.allclose(
0.5 * np.einsum("nP, n, nQ -> PQ", X + Y, td_eig**2, X - Y),
(A - B) @ (A + B))
```
下面我们来讨论矩阵 $(\mathbb{A} - \mathbb{B}) (\mathbb{A} + \mathbb{B}) - \omega^2 \mathbb{1}$。我们容易从式 (8) 推得
$$
\left( (\mathbb{A} - \mathbb{B}) (\mathbb{A} + \mathbb{B}) - \omega^2 \mathbb{1} \right) (\mathbf{X}^n + \mathbf{Y}^n) = (\omega_n^2 - \omega^2) (\mathbf{X}^n + \mathbf{Y}^n)
$$
即上述方程的本征向量仍然是 $(\mathbf{X}^n + \mathbf{Y}^n)$,但本征值却变为 $\omega_n^2 - \omega^2$。因此,
$$
\begin{align}
\left( (\mathbb{A} - \mathbb{B}) (\mathbb{A} + \mathbb{B}) - \omega^2 \mathbb{1} \right)^{-1} = \left( \frac{1}{2} (\mathbf{X} + \mathbf{Y}) (\mathbf{\Omega}^2 - \omega^2 \mathbb{1}) (\mathbf{X} - \mathbf{Y})^\dagger \right)^{-1} = \frac{1}{2} (\mathbf{X} + \mathbf{Y}) (\mathbf{\Omega}^2 - \omega^2 \mathbb{1})^{-1} (\mathbf{X} - \mathbf{Y})^\dagger \tag{9}
\end{align}
$$
上式中 $(\mathbf{\Omega}^2 - \omega^2 \mathbb{1})^{-1}$ 是一个极为容易计算的对角矩阵。$\omega = 0.186 \, E_\mathrm{h}$ 下,程序表示上述过程则为
```
np.allclose(
0.5 * np.einsum("nP, n, nQ -> PQ", X + Y, 1 / (td_eig**2 - omega**2), X - Y),
np.linalg.inv((A - B) @ (A + B) - omega**2 * np.eye(nvir*nocc)))
```
将式 (9) 代入到式 (7),得到
$$
\alpha_{ts} (\omega) = 2 \mathbf{d}^s{}^\dagger (\mathbf{X} + \mathbf{Y}) (\mathbf{\Omega}^2 - \omega^2 \mathbb{1})^{-1} (\mathbf{X} - \mathbf{Y})^\dagger (\mathbb{A} - \mathbb{B}) \mathbf{d}^t
$$
利用极化率的 Hermite 性,并将上述矩阵表达式展开为求和式,得到
$$
\alpha_{ts} (\omega) = 2 d_P^t (\mathbf{X}^n + \mathbf{Y}^n)_P \cdot \frac{1}{\omega_n^2 - \omega^2} \cdot (\mathbf{X}^n - \mathbf{Y}^n)_R (\mathbb{A} - \mathbb{B})_{RQ} d_Q^s
$$
```
2 * np.einsum("tP, nP, n, nR, RQ, sQ", d_P, X + Y, 1 / (td_eig**2 - omega**2), X - Y, A - B, d_P)
```
若要化简上式,首先需要利用 $(\mathbb{A} - \mathbb{B})_{RQ}$ 事实上恰好是一个 Hermite 矩阵:
```
np.allclose(A - B, (A - B).T)
```
随后注意到 $(\mathbb{A} - \mathbb{B})_{QR} (\mathbf{X}^n - \mathbf{Y}^n)_R = \omega_n (\mathbf{X}^n + \mathbf{Y}^n)_Q$;于是上式化为
$$
\alpha_{ts} (\omega) = 2 d_P^t (\mathbf{X}^n + \mathbf{Y}^n)_P \cdot \frac{\omega_n}{\omega_n^2 - \omega^2} \cdot (\mathbf{X}^n + \mathbf{Y}^n)_Q d_Q^s
$$
我们留意到跃迁偶极矩的定义是 $\langle 0 | \hat d{}^t | n \rangle = d_P^t (\mathbf{X}^n + \mathbf{Y}^n)_P$,并且利用下面的小技巧:
$$
\frac{\omega_n}{\omega_n^2 - \omega^2} = \frac{1}{2} \left( \frac{1}{\omega_n - \omega} - \frac{1}{\omega_n + \omega} \right)
$$
我们就可以推知,
$$
\alpha_{ts} (\omega) = \frac{\langle 0 | \hat d{}^t | n \rangle \langle n | \hat d{}^s | 0 \rangle}{\omega_n - \omega} + \frac{\langle 0 | \hat d{}^t | n \rangle \langle n | \hat d{}^s | 0 \rangle}{\omega_n + \omega}
$$
```
(
+ np.einsum("tP, nP, n, nQ, sQ -> ts", d_P, X + Y, 1 / (td_eig - omega), X + Y, d_P)
+ np.einsum("tP, nP, n, nQ, sQ -> ts", d_P, X + Y, 1 / (td_eig + omega), X + Y, d_P)
)
```
这就完成了从普适的 Casida 方程推演得到跃迁偶极矩表示的极化率的公式了。
若接受 Casida 方程的假设前提,那么上述的推演将会是严格的。
### TD-HF 方程含频极化率与 CP-HF 方程间的关系
我们先回顾静态极化率求取时所使用的 CP-HF 方程:
$$
A'_{ia, jb} U^t_{jb} = d^t_{ia}
$$
写为双下标的形式,则为
$$
A'_{PQ} U^t_Q = d^t_P
$$
写为矩阵形式,则为
$$
\mathbf{A}' \mathbf{U}^t = \mathbf{d}^t
$$
留意到 $A'_{PQ} = (\mathbb{A} + \mathbb{B})_{PQ}$,因此上式这对应到 Casida 方程的一个导出式 (5)。若 $\omega = 0$,则
$$
(\mathbb{A} + \mathbb{B}) (\mathbf{X}'{}^t + \mathbf{Y}'{}^t) = 4 \mathbf{d}^t
$$
因此,在静态情形 $\omega = 0$ 下,$\mathbf{U}^t = \frac{1}{4} (\mathbf{X}' + \mathbf{Y}')$。
但若 $\omega \neq 0$,那么式 (5) 应写作
$$
\left( (\mathbb{A} + \mathbb{B}) - \omega^2 (\mathbb{A} - \mathbb{B})^{-1} \right) (\mathbf{X}'{}^t + \mathbf{Y}'{}^t) = 4 \mathbf{d}^t
$$
若我们拓展 CP-HF 方程为含频形式
$$
\mathbf{A}' (\omega) \mathbf{U}^t (\omega) = \mathbf{d}^t
$$
并且极化率可以写为
$$
\alpha_{ts} (\omega) = 4 \mathbf{U}^t (\omega)^\dagger \mathbf{d}^t
$$
那么
$$
\begin{align}
\mathbf{A}' (\omega) &= (\mathbb{A} + \mathbb{B}) - \omega^2 (\mathbb{A} - \mathbb{B})^{-1} \\
\mathbf{U}^t (\omega) &= \frac{1}{4} (\mathbf{X}'{}^t + \mathbf{Y}'{}^t)
\end{align}
$$
需要留意尽管我们之前一直没有引入 $(\omega)$ 记号来强调,但 $\mathbf{X}'{}^t, \mathbf{Y}'{}^t$ 是随频率变化而变化的。
```
omega = 0.186
A_p_omega = (A + B) - omega**2 * np.linalg.inv(A - B)
U_omega = np.einsum("PQ, tQ -> tP", np.linalg.inv(A_p_omega), d_P)
4 * np.einsum("tP, sP -> ts", U_omega, d_P)
```
这就在含频情形下,将 CP-HF 与 TD-HF 的公式联系在了一起。
## 总结
这篇文档我们简单且不太严格和完整地回顾了静态与含频极化率的计算,通过 Casida 方程推导了含频极化率,并将 TD-HF 分析方法与 CP-HF 方法联系起来。一些主要的结论和拓展思路会是:
- TD-HF 方程与 CP-HF 方程在静态极化率情形下有极为紧密的联系,两种表达式完全等价;而含频极化率情形下,TD-HF 方程 (Casida 方程) 可以导出与 CP-HF 方程形式类似的方程。这可能是 Frequency-Dependent CP-HF 方程的原型。
- 从 CP 的角度讲,根据下式
$$
(\mathbb{A} + \mathbb{B}) (\mathbf{X}'{}^t + \mathbf{Y}'{}^t) = 4 \mathbf{d}^t + \omega (\mathbb{A} - \mathbb{B})^{-1} (\mathbf{X}'{}^t + \mathbf{Y}'{}^t)
$$
或
$$
\mathbf{A}' (\omega) \mathbf{U}^t (\omega) = \mathbf{d}^t
$$
我们可以想到等式左边是电子云的弛豫过程;等式右边是电子云所受到的外加微扰场。因此电子云的激发过程可以看作外场微扰。
若退化为没有偶极微扰的分子激发过程,从 CP 的角度看,电子云变形的弛豫效应 (等式左边) 应当恰好能补足电子云激发所导致的变形外场 (等式右边)。当两者自洽时,电子云能态的激发成立。
- 上述所述的 TD-HF 方程是 TD 分析中的 Linear Response 关系。事实上,TD 还具有更高阶的分析过程,但这里我们并没有涉及。
一个显而易见的问题会是,电子云已经在外加偶极场中,因此电子云的性质应当已经发生变化;但 Linear Response 给出的 TD-HF 方程 (Casida 方程) 却告诉我们体系的激发能 $\omega_n$ 并没有变化,无论偶极场的值有多大。这显然违背我们的物理直觉。这也可能意味着以 MP2、CC 为底层量化方法,若外加的相关能密度微扰近似场没有把握住物理实在,那么含频极化率也可能面临激发能 $\omega_n$ 还仍然是 TD-HF 激发能的情况。
- 文档中没有画出,但确实存在的情况是,若入射频率恰好处在分子的激发态密集区,那么单个入射频率下的含频极化率的计算很可能是没有意义的;因为在激发态密集区,含频极化率曲线的振荡情况及其严重,不仅近乎毫无规律可言,其绝对值也大到非常离谱。
Raman 光谱可以认为是含频极化率对分子的简正坐标导数得来。从实验的角度来讲,这可能是表面增强 Raman (SERS) 中化学增强效应的一个很好的性质。若有机物负载在银原子簇上,入射的激发光处于银原子簇的激发能带但又不破坏有机物分子,或让银原子簇与有机物分子产生电荷转移时,Raman 信号确实可以成千上万倍地增强。但从计算的角度来讲,这可能是非常令人绝望的,因为计算过程中的误差把控近乎于不存在;若使用不同的密度泛函近似,即使得到相差 0.1 eV 的激发能带差,即使定性上结果正确,但计算得到的 Raman 信号也可能会有成百上千的误差。
因此,这可能需要我们对入射光作某种频率增宽,以平缓极化率断点,得到相对定量的数值结果;或者将极化率当作原子可加性的物理性质,进行类似于 AIM (Atomic in Molecule) 的极化率拆分的定性近似分析。
[^Valley-Schatz.JPCL.2013]: Valley, N.; Greeneltch, N.; Duyne, R. P. V.; Schatz, G. C. A Look at the Origin and Magnitude of the Chemical Contribution to the Enhancement Mechanism of Surface-Enhanced Raman Spectroscopy (SERS): Theory and Experiment. *J. Phys. Chem. Lett.* **2013**, *4* (16), 2599–2604. doi: [10.1021/jz4012383](https://doi.org/10.1021/jz4012383).
| github_jupyter |
Consider using Cairo for plotting...
```
import math
import cairo # see https://www.cairographics.org/samples/
from IPython.display import Image
if (cairo.HAS_SVG_SURFACE and cairo.HAS_PNG_FUNCTIONS):
print ('Cairo: {c}'.format(c=cairo.CAIRO_VERSION_STRING))
with cairo.SVGSurface("./img/example.svg", 100, 100) as surface:
context = cairo.Context(surface)
x, y, x1, y1 = 0.1, 0.5, 0.4, 0.9
x2, y2, x3, y3 = 0.6, 0.1, 0.9, 0.5
context.scale(100, 100)
context.set_line_width(0.04)
context.move_to(x, y)
context.curve_to(x1, y1, x2, y2, x3, y3)
context.stroke()
context.set_source_rgba(1, 0.2, 0.2, 0.6)
context.set_line_width(0.02)
context.move_to(x, y)
context.line_to(x1, y1)
context.move_to(x2, y2)
context.line_to(x3, y3)
context.stroke()
```

```
anno_str = 'Illumina-450k-Anno.{rev}.{ext}'.format(rev='hg19',ext='pkl')
annotation = load_obj(anno_str[:-4]) # load the saved annotation file
print (len(annotation.probe))
def _range(probes, interval):
'''
Return an unsorted range of probes.
Equivalent to df.head() at any starting location.
'''
return dict(list(probes.items())
[interval.start:interval.end])
def sort_range_by_refid(probes):
'''
Return dict of range of probes, sorted by refid.
Call _range function first to limit scope.
'''
return dict(sorted(list(probes.items())))
def sort_range_by_coordinate(probes):
'''
Return dict of range of probes, sorted by coordinate.
Call _range function first to limit scope.
'''
return dict(sorted(list(probes.items()),
key=lambda item: item[1].cord))
def get_probes_by_chr(probes, interval):
"""
Return a dict of probes by chromosome.
Call _range function first to limit scope.
"""
return {p: probes[p] for p in probes
if probes[p].chr == interval.chr}
def get_probes_by_location(probes, interval):
"""
Return a dict of probes by location.
Call _range function first to limit scope.
"""
chrom = interval.chr
probe_dict = {k: probes[k] for k in probes if
probes[k].chr == chrom and start < probes[k].cord < end}
return probe_dict
chrom = 'Y'
start = 0
end = 5
_slice = Interval(chrom, start, end, '+')
probe_slice = _range(annotation.probe, _slice) # dict of the first 10 annotation entries.
# Cord Chr ID*
# 8553009 Y cg00035864
# 9363356 Y cg00050873
# 25314171 Y cg00061679
# 22741795 Y cg00063477
# 21664296 Y cg00121626
sort_range_by_refid(probe_slice)
# 'cg00035864' 8553009
# 'cg00050873' 9363356
# 'cg00061679' 25314171
# 'cg00063477' 22741795
# 'cg00121626' 21664296
sort_range_by_coordinate(probe_slice)
# 'cg00035864' 8553009
# 'cg00050873' 9363356
# 'cg00121626' 21664296
# 'cg00063477' 22741795
# 'cg00061679' 25314171
get_probes_by_chr(probe_slice, _slice)
sorted_locations = sorted([probes[k].cord for k in probes])
sorted_locations = sorted([probes[k].cord for k in probes])
midway = int((interval.start + interval.end)/2)
if midway in range(sorted_locations[0], sorted_locations[-1]):
print ('found')
#
#probes_by_key = sorted([probe for probe in probes]) # sorts on REF_ID by default
#probes_by_key
sorted_locations = sorted([probes[k].cord for k in probes])
midway = int((interval.start + interval.end)/2)
if midway in range(sorted_locations[0], sorted_locations[-1]):
print ('found')
mid in range(sorted_locations[0],sorted_locations[-1])
mid
len(range(range_tup[0], range_tup[1]))
int((range_tup[0] + range_tup[1])/2)
def get_probes_in_range(start, stop):
return dict(sorted(list(annotate.probe.items())[start:stop]))
probes = get_probes_in_range(0,5) # dict of the first 5 entries in annotate.probe
chrom = 'Y'
start = 8443000
stop = 8572220
# key: cg00035864 cg00050873 cg00061679 cg00063477 cg00121626
# cord: 8553009 9363356 25314171 22741795 21664296
# chr: Y Y Y Y Y
[probes[k] for k in probes if probes[k].chr == chrom and start < probes[k].cord < stop]
# %load '../methylator/annotation/annotate_450k.py'
import os
class Probe:
"""
Holds Illumina 450k probe info for a single CpG site.
"""
def __init__(self):
self.id = None
self. seq = None
self.name = None
self.chr = None
self.cord = None
self.strand = None
self.gene = None
self.refseq = None
self.tour = None
self.loc = None
class Interval:
"""
Define a genomic interval by chromsome and strand orientation.
"""
def __init__(self, chromosome, start, end, strand):
self.chr = chromosome
self.start = start
self.end = end
self.strand = strand
class Location:
"""
Define a Probe location.
"""
BODY = "Body"
TSS200 = "TSS200"
TSS1500 = "TSS1500"
UTR5 = "5'UTR"
UTR3 = "3'UTR"
EXON = "Exon"
class CpG_location:
"""
Defines a CpG location.
"""
ISLAND = "Island"
NSHORE = "N_Shore"
SSHORE = "S_Shore"
NSHELF = "N_Shelf"
SSHELF = "S_Shelf"
class SNP:
"""
Defines the SNPs in probes. Used to filter probes.
"""
def __init__(self):
self.probeid = None
self.snpid = None
class Annotate_450k:
"""
Parse and hold information about Illumina probes.
"""
def __init__(self):
for i in open(anno_file, mode="r"):
self.ann = os.path.join("../../data/", i.strip("\n").strip("\r"))
self.probe = {}
self.__run__()
def __run__(self):
"""
A static function to setup the Probe classes.
"""
for i in open(self.ann, mode="r"):
if i.startswith("cg"):
data = i.split(",")
# Assign probe information.
new_probe = Probe()
new_probe.id = data[0]
new_probe.name = data[1]
new_probe.seq = data[13]
new_probe.chr = str(data[11])
new_probe.cord = int(data[12])
new_probe.strand = data[16]
new_probe.gene = data[21].split(";")
new_probe.refseq = data[22]
locs = data[23].split(";")
list_locs = []
for i in locs:
if i not in list_locs:
list_locs.append(i)
new_probe.loc = list_locs
new_probe.tour = data[25]
newcpg = {new_probe.id: new_probe}
self.probe.update(newcpg)
def get_probe(self, probe_id): #WORKS
"""
Return probe info associated with an reference.
"""
try:
probe = self.probe[probe_id]
except Exception as ex:
probe = None
print("WARNING: No probe with ref-id of %s found." % probe_id)
return probe
def get_all_probes(self):
"""
Return list of all probes.
"""
probe_list = []
for probe in self.probe.keys():
probe_list.append(self.get_probe(probe))
return probe_list
def get_probes_by_list(self, list_of_ids):
"""
Return a list of probes from a list of references.
"""
out_list = []
for probe_id in list_of_ids:
out_list.append(self.get_probe(probe_id))
return out_list
def get_probe_refs_by_gene(self, gene_name):
"""
Get all probe references associated with a gene.
"""
probes = {k: self.probe[k] for k in self.probe if gene_name in self.probe[k].gene}
return self.get_keys(probes.keys())
def get_probe_refs_by_location(self, probe_loc):
"""
Get all probe references associated with a genomic location.
"""
probes = {k: self.probe[k] for k in self.probe if probe_loc in self.probe[k].loc}
return self.get_keys(probes.keys())
def get_keys(self, dic_keys):
"""
Get Probe reference from probe dictionaries.
"""
l = []
for i in dic_keys:
l.append(i)
return l
def get_probes_by_gene(self, gene_name):
"""
Return list of probes for an associated gene.
"""
return self.get_probes_by_list(self.get_probe_refs_by_gene(gene_name))
def get_probes_by_location(self, loc):
"""
Return list of probes from genomic location.
"""
return self.get_probes_by_list(self.get_probe_refs_by_location(loc))
def get_probes_by_cpg(self, cpg_loc):
"""
Get a list probes from cpg location.
FIXME
"""
return self.get_probes_by_list(self.get_probes_by_cpg(cpg_loc))
def get_probes_by_chr(self, chr_loc):
"""
Get a list of probes within a certain genomic region
FIXME
"""
print (chr_loc.chr)
probes = {k: self.probe[k] for k in self.probe if
self.probe[k].chr == chr_loc.chr}
def get_probes_by_chr_and_loc(self, chr_loc):
"""
Get a list of probes within a certain genomic region
FIXME
"""
chrom = chr_loc.chr
start = int(chr_loc.start)
end = int(chr_loc.end)
#print (chrom, start, stop)
probes = {k: self.probe[k] for k in self.probe if
self.probe[k].chr == chrom and start < self.probe[k].cord < end}
return probes
def get_probe_keys_by_chr_and_loc(self, chr_loc):
"""
Get a list of probe reference *keys* within a genomic region
FIXME
"""
probes = self.get_probes_by_chr_and_loc(chr_loc)
return self.get_keys(probes)
def get_number(self):
"""
Return total number of probes.
"""
number = 0
for probe_id in self.probe.keys():
number += 1
return number
def get_coord(self, probe):
"""
Get genomic coordinate of a single probe.
"""
return probe.cord
def get_sorted_probes_by_id(self):
"""
Sort probes according to probe id.
"""
sorted_keys = sorted(list(self.probe.keys()))
return sorted_keys
def get_sorted_probes_by_chr(self):
"""
Sort probes according to probe id.
"""
return sorted(self.get_all_probes(), key=lambda x: x.chr)
def remove_snp_probes(self):
"""
Removes all SNPs associated with probes.
"""
snp_list = []
snp_file = open("../../data/humanmethylation450_dbsnp137.snpupdate.table.v2.sorted.txt", "r")
for line in snp_file:
if line.startswith("cg"):
line = line.strip("\n").strip("\r").split("\t")
new_snp = SNP()
new_snp.probeid = line[0]
new_snp.snpid = line[1]
snp_list.append(new_snp)
for snp in snp_list:
self.probe.pop(snp.probeid)
anno_file = os.path.abspath("../../data/config.ini") # Illumina probe manifest. Note: This (large) file is not
# in the repository.
# Functions to save/load dictionary objects.
import _pickle as pickle
def save_obj(obj, name):
with open('../../data/pickle/'+ name + '.pkl', 'wb+') as f:
pickle.dump(obj, f)
def load_obj(name):
with open('../../data/pickle/' + name + '.pkl', 'rb') as f:
return pickle.load(f)
```
| github_jupyter |
### In this notebook we will compile interesting questions that we can answer using the data from this table.
### We can refer to data_exploration.ipynb to figure out what kind of information we already have.
1. On an average, how often do people order from Instacart?
2. What product was ordered most often?
3. At what time during the day do people order most? Can we have a plot for how busy different times during the day are?
4. How often do people order ice-cream? How often do they order alcohol?
5. If we classify all food-related orders under food groups (dairy, vegetables, protein) - what groups are represented more than others in terms of number of items?
6. What kind of alcohol is most popular? lol
7. What aisles are the most popular? Can we draw any conclusions about eating habits based on this?
8. What is usually the first item people put into their cart?
### Q1. On an average, how often do people order from Instacart?
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
orders = pd.read_csv('./instacart-market-basket-analysis/orders.csv')
# plan: take the average of the row days_since_prior_order, but don't include NaNs.
# pandas ignores NaNs while taking average, so we only need to call the average method on the series.
print('On an average, people order once every ', orders['days_since_prior_order'].mean(), 'days')
```
### Distribution of orders with respect to days
```
order_by_date = orders.groupby(by='days_since_prior_order').count()
fig = plt.figure(figsize = [15, 7.5])
ax = fig.add_subplot()
order_by_date['order_id'].plot.bar(color = '0.75')
ax.set_xticklabels(ax.get_xticklabels(), fontsize= 15)
plt.yticks(fontsize=16)
ax.get_xaxis().set_major_formatter(matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x))))
ax.get_yaxis().set_major_formatter(matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x/1000))))
ax.set_xlabel('Days since previous order', fontsize=16)
ax.set_ylabel('Number of orders / 1000', fontsize=16)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
# ax.spines['bottom'].set_visible(False)
# ax.spines['left'].set_visible(False)
# setting 7 and 30 to a solid blue color
ax.get_children()[7].set_color('0.1')
ax.get_children()[14].set_color('0.1')
ax.get_children()[21].set_color('0.1')
ax.get_children()[30].set_color('0.1')
my_yticks = ax.get_yticks()
plt.yticks([my_yticks[-2]], visible=True)
plt.xticks(rotation = 'horizontal');
```
### Another cool plot
```
# along with this, we need the average size of the order for each day since previous order value
# for order products table, join on order_id, then you get order_id many times and different product ids.
orders = pd.read_csv('./instacart-market-basket-analysis/orders.csv')
order_products_prior = pd.read_csv('./instacart-market-basket-analysis/order_products__prior.csv')
order_id_count_products = order_products_prior.groupby(by='order_id').count()
orders_with_count = order_id_count_products.merge(orders, on='order_id')
# take above table and group by days_since_prior_order
df_mean_order_size = orders_with_count.groupby(by='days_since_prior_order').mean()['product_id']
df_mean_order_renamed = df_mean_order_size.rename('average_order_size')
bubble_plot_dataframe = pd.concat([order_by_date['order_id'], df_mean_order_renamed], axis=1)
bubble_plot_dataframe['average_order_size'].index.to_numpy()
import seaborn as sns
fig = plt.figure(figsize=[15,7.5])
ax = fig.add_subplot()
plt.scatter(bubble_plot_dataframe['average_order_size'].index.to_numpy(), bubble_plot_dataframe['order_id'].values, s=((bubble_plot_dataframe['average_order_size'].values/bubble_plot_dataframe['average_order_size'].values.mean())*10)**3.1, alpha=0.5, c = '0.5')
plt.xticks(np.arange(0, 31, 1.0));
ax.xaxis.grid(True)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.set_xlabel('Days since previous order', fontsize=16)
ax.set_ylabel('Number of orders / 1000', fontsize=16)
ax.get_xaxis().set_major_formatter(matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x))))
ax.get_yaxis().set_major_formatter(matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x/1000))))
my_yticks = ax.get_yticks()
plt.yticks([my_yticks[-2], my_yticks[0]], visible=True);
fig = plt.figure(figsize=[10,9])
ax = fig.add_subplot()
plt.scatter(bubble_plot_dataframe['average_order_size'].index.to_numpy()[:8], bubble_plot_dataframe['order_id'].values[:8], s=((bubble_plot_dataframe['average_order_size'].values[:8]/bubble_plot_dataframe['average_order_size'].values.mean())*10)**3.1, alpha=0.5, c = '0.5')
plt.xticks(np.arange(0, 8, 1.0));
ax.xaxis.grid(True)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.set_xlabel('Days since previous order', fontsize=16)
ax.set_ylabel('Number of orders / 1000', fontsize=16)
ax.get_xaxis().set_major_formatter(matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x))))
ax.get_yaxis().set_major_formatter(matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x/1000))))
my_yticks = ax.get_yticks()
plt.yticks([my_yticks[-2], my_yticks[0]], visible=True);
```
### Q2. What product was ordered most often?
```
# plan - we use two tables: order_products__train.csv and order_products__prior.csv. We assume these have different values, and we check if they are the same. Best to check the kaggle dataset for any hints as to what these files have.
# for simplicity, we take just one file for this question - prior.
order_products_prior = pd.read_csv('./instacart-market-basket-analysis/order_products__prior.csv')
# the table looks like so:
order_products_prior
order_id_count_products = order_products_prior.groupby(by='order_id').count()
order_id_count_products
products = pd.read_csv('./instacart-market-basket-analysis/products.csv')
# for each order, we want each product to be considered only once in our calculations. So we should remove duplicates of the same order_id and product_id for this question.
df = order_products_prior.drop_duplicates(subset=['order_id', 'product_id'])
df_with_product_description = df.merge(products, on = 'product_id', how='inner')
df_with_product_description
# now if we group by product_id and show count along with product_id, that should be quite interesting
# but product_id tells us nothing, so best to also join the result to products table.
df_with_product_description.groupby(['product_name']).count().sort_values(by = 'order_id', ascending=False)['order_id'].head()
```
Bar plots
Just the head
```
print('Bananas were ordered most often, followed Strawberries, Baby Spinach, and Avocados')
```
### Q3. At what time during the day do people order most often?
```
# plan - take orders table and group by order time of day
time_of_day = orders.groupby(by='order_hour_of_day').count().sort_values(by='order_hour_of_day', ascending=True)
plt.figure(figsize=(10,5))
time_of_day['order_id'].plot.bar()
plt.ylabel('Number of orders');
```
### Q4. What departments are the most popular? What can we say about food habits based on this?
```
# plan: Which departments are most popular?
# which departments have the most products ordered from them.
# from order product table, group by department. Merge with department table.
department = pd.read_csv('./instacart-market-basket-analysis/departments.csv')
df_with_product_description_dept = df_with_product_description.merge(department, on='department_id') # contains all the info for this question
df_with_product_description_dept
how_many_items_per_department = df_with_product_description_dept.groupby(by='department').count()['order_id'].sort_values(ascending = False)
how_many_items_per_department.plot.pie(y='order_id', figsize = [10,10], title = 'Share of items per department (version 1)');
```
Too crowded.
### Q8. What is usually the first item that shoppers put into their carts?
### filter the dataset such that you only have those orders which satisfy add_to_cart_order == 1
```
first_in_cart = df_with_product_description['add_to_cart_order'] == 1
first_in_cart_products = df_with_product_description[first_in_cart]
first_in_cart_products.groupby(by='product_name').count()['order_id'].sort_values(ascending = False)[:15]
```
The first thing people put in their carts is generally produce.
Why do people order produce so often? Because it is perishable. You buy it in small quantities, because if you buy too much at one time, it goes bad. Another reason is that it is genuinely popular. Fruits make great snacks!
Why organic: People who can afford instacart can also afford organic might be a reason for this trend.
| github_jupyter |
## 1. Longest Substring Without Repeating Characters
Given a string, find the length of the longest substring without repeating characters.
```
def lengthOfLongestSubstring(s):
stack = []
maxL = 0
for st in s:
if st in stack:
stack = stack[stack.index(st)+1:]
stack.append(st)
maxL = max(maxL, len(stack))
return maxL
s = "abcabcbb"
lengthOfLongestSubstring(s)
```
## 2. Generate Parentheses
Given n pairs of parentheses, write a function to generate all combinations of well-formed parentheses.
```
def generateParenthesis(n):
def gen(p, l, r, par=[]):
if l:
gen(p+'(', l-1, r)
if l < r:
gen(p + ')',l, r-1)
if r == 0:
par += [p]
return par
return gen('',n,n)
n = 3
generateParenthesis(n)
# dp solution
def generateParenthesis(n):
dp = [[] for i in range(n+1)]
dp[0].append('')
for i in range(n+1):
for j in range(i):
dp[i] += ['(' + x + ')' + y for x in dp[j] for y in dp[i-j-1]]
return dp[n]
n = 3
generateParenthesis(n)
```
## 3. Letter Combinations of a Phone Number
Given a string containing digits from 2-9 inclusive, return all possible letter combinations that the number could represent.
A mapping of digit to letters (just like on the telephone buttons) is given below. Note that 1 does not map to any letters.
```
def letterCombinations(digits):
dic = {'2':'abc', '3':'def', '4':'ghi', '5':'jkl','6':'mno','7':'pqs', '8':'tuv', '9':'wxyz'}
if not digits:
return []
if len(digits) == 1:
return list(dic[digits])
ite = letterCombinations(digits[:-1])
add = dic[digits[-1]]
return [i + a for i in ite for a in add]
# look at the primary loop for iterative method
digits = '23'
letterCombinations(digits)
def letterCombinations(digits):
dic = {'2':"abc", '3':"def", '4':"ghi", '5':"jkl", '6':"mno", '7': "pqrs", '8':"tuv", '9':"wxyz"}
cmb = [''] if digits else []
for d in digits:
cmb = [p + q for p in cmb for q in dic[d]]
return cmb
digits = '23'
letterCombinations(digits)
```
## 4. Length of Last Word
Given a string s consists of upper/lower-case alphabets and empty space characters ' ', return the length of last word (last word means the last appearing word if we loop from left to right) in the string.
If the last word does not exist, return 0.
```
def lengthOfLastWord(s):
return len(s.rstrip(' ').split(' ')[-1])
s = "Hello World"
lengthOfLastWord(s)
```
## 5. Group Anagrams
Given an array of strings, group anagrams together.
```
def groupAnagrams(strs):
dic = {}
for word in strs:
key = ''.join(sorted(word))
dic[key] = dic.get(key, []) + [word]
return list(dic.values())
strs = ["eat", "tea", "tan", "ate", "nat", "bat"]
groupAnagrams(strs)
```
## 6. Interleaving String
Given s1, s2, s3, find whether s3 is formed by the interleaving of s1 and s2.
```
# 1D dp solution, could be 2D dp array solution where dp[i][j] is boolean
# and represents if (i+j+2)-prefix of s3 could be create by (i+1)-prefix of s1
# and (j+1)-prefix of s2
def isInterleave(s1, s2, s3):
l1, l2, l3= len(s1), len(s2), len(s3)
if l1+l2 != l3:
return False
dp = [True for _ in range(l2+1)]
for j in range(1, l2+1):
dp[j] = dp[j-1] and s2[j-1] == s3[j-1]
for i in range(1, l1+1):
dp[0] = dp[0] and s1[i-1] == s3[i-1]
for j in range(1, l2+1):
dp[j] = (dp[j] and s1[i-1] == s3[i-1+j]) or (dp[j-1] and s2[j-1] == s3[i-1+j])
return dp[-1]
s1 = "aabcc"
s2 = "dbbca"
s3 = "aadbbcbcac"
isInterleave(s1, s2, s3)
```
## 7. Remove Element
Given an array nums and a value val, remove all instances of that value in-place and return the new length.
Do not allocate extra space for another array, you must do this by modifying the input array in-place with O(1) extra memory.
The order of elements can be changed. It doesn't matter what you leave beyond the new length.
```
def removeElement(nums, val):
l, r = 0, len(nums)-1
while l <= r:
if nums[l] == val:
nums[l], nums[r], r = nums[r], nums[l], r-1
else:
l += 1
return l
# careful with nums = single target case
nums = [3,2,2,3]
val = 3
removeElement(nums, val)
```
## 8. Simplify Path
Given an absolute path for a file (Unix-style), simplify it. Or in other words, convert it to the canonical path.
In a UNIX-style file system, a period . refers to the current directory. Furthermore, a double period .. moves the directory up a level.
Note that the returned canonical path must always begin with a slash /, and there must be only a single slash / between two directory names. The last directory name (if it exists) must not end with a trailing /. Also, the canonical path must be the shortest string representing the absolute path.
```
def simplifyPath(path):
stack = []
for place in path.split("/"):
if place == "..":
if stack:
stack.pop()
elif place and place != '.':
stack.append(place)
return '/' + '/'.join(stack)
path = "/home//foo/"
simplifyPath(path)
path = 'a/b/c/..//.//'
path.split('/')
```
## 9. Reverse Words in a String
Given an input string, reverse the string word by word.
```
def reverseWords(s):
stack = []
for w in s.split(' ')[::-1]:
if w:
stack.append(w)
return ' '.join(stack)
s = "a good example"
reverseWords(s)
```
## 10. Integer to Roman
Roman numerals are represented by seven different symbols: I, V, X, L, C, D and M.
For example, two is written as II in Roman numeral, just two one's added together. Twelve is written as, XII, which is simply X + II. The number twenty seven is written as XXVII, which is XX + V + II.
Roman numerals are usually written largest to smallest from left to right. However, the numeral for four is not IIII. Instead, the number four is written as IV. Because the one is before the five we subtract it making four. The same principle applies to the number nine, which is written as IX. There are six instances where subtraction is used:
- I can be placed before V (5) and X (10) to make 4 and 9.
- X can be placed before L (50) and C (100) to make 40 and 90.
- C can be placed before D (500) and M (1000) to make 400 and 900.
Given an integer, convert it to a roman numeral. Input is guaranteed to be within the range from 1 to 3999.
```
def intToRoman(num):
dic = {1000:'M', 900:'CM', 500:'D', 400:'CD', 100:'C', 90:'XC', 50:'L',
40:'XL', 10:'X', 9:'IX', 5:'V', 4:'IV', 1:'I'}
ans = ''
for i in dic:
if num >= i:
ans += dic[i] * (num//i)
num %= i
return ans
num = 4
intToRoman(num)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/MIT-LCP/sccm-datathon/blob/master/01_explore_patients.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# eICU Collaborative Research Database
# Notebook 1: Exploring the patient table
The aim of this notebook is to get set up with access to a demo version of the [eICU Collaborative Research Database](http://eicu-crd.mit.edu/). The demo is a subset of the full database, limited to ~1000 patients.
We begin by exploring the `patient` table, which contains patient demographics and admission and discharge details for hospital and ICU stays. For more detail, see: http://eicu-crd.mit.edu/eicutables/patient/
## Prerequisites
- If you do not have a Gmail account, please create one at http://www.gmail.com.
- If you have not yet signed the data use agreement (DUA) sent by the organizers, please do so now to get access to the dataset.
## Load libraries and connect to the data
Run the following cells to import some libraries and then connect to the database.
```
# Import libraries
import numpy as np
import os
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.path as path
# Make pandas dataframes prettier
from IPython.display import display, HTML
# Access data using Google BigQuery.
from google.colab import auth
from google.cloud import bigquery
```
Before running any queries, you need to first authenticate yourself by running the following cell. If you are running it for the first time, it will ask you to follow a link to log in using your Gmail account, and accept the data access requests to your profile. Once this is done, it will generate a string of verification code, which you should paste back to the cell below and press enter.
```
auth.authenticate_user()
```
We'll also set the project details.
```
project_id='sccm-datathon'
os.environ["GOOGLE_CLOUD_PROJECT"]=project_id
```
# "Querying" our database with SQL
Now we can start exploring the data. We'll begin by running a simple query to load all columns of the `patient` table to a Pandas DataFrame. The query is written in SQL, a common language for extracting data from databases. The structure of an SQL query is:
```sql
SELECT <columns>
FROM <table>
WHERE <criteria, optional>
```
`*` is a wildcard that indicates all columns
# BigQuery
Our dataset is stored on BigQuery, Google's database engine. We can run our query on the database using some special ("magic") [BigQuery syntax](https://googleapis.dev/python/bigquery/latest/magics.html).
```
%%bigquery patient
SELECT *
FROM `physionet-data.eicu_crd_demo.patient`
```
We have now assigned the output to our query to a variable called `patient`. Let's use the `head` method to view the first few rows of our data.
```
# view the top few rows of the patient data
patient.head()
```
## Questions
- What does `patientunitstayid` represent? (hint, see: http://eicu-crd.mit.edu/eicutables/patient/)
- What does `patienthealthsystemstayid` represent?
- What does `uniquepid` represent?
```
# select a limited number of columns to view
columns = ['uniquepid', 'patientunitstayid','gender','age','unitdischargestatus']
patient[columns].head()
```
- Try running the following query, which lists unique values in the age column. What do you notice?
```
# what are the unique values for age?
age_col = 'age'
patient[age_col].sort_values().unique()
```
- Try plotting a histogram of ages using the command in the cell below. What happens? Why?
```
# try plotting a histogram of ages
patient[age_col].plot(kind='hist', bins=15)
```
Let's create a new column named `age_num`, then try again.
```
# create a column containing numerical ages
# If ‘coerce’, then invalid parsing will be set as NaN
agenum_col = 'age_num'
patient[agenum_col] = pd.to_numeric(patient[age_col], errors='coerce')
patient[agenum_col].sort_values().unique()
patient[agenum_col].plot(kind='hist', bins=15)
```
## Questions
- Use the `mean()` method to find the average age. Why do we expect this to be lower than the true mean?
- In the same way that you use `mean()`, you can use `describe()`, `max()`, and `min()`. Look at the admission heights (`admissionheight`) of patients in cm. What issue do you see? How can you deal with this issue?
```
adheight_col = 'admissionheight'
patient[adheight_col].describe()
# set threshold
adheight_col = 'admissionheight'
patient[patient[adheight_col] < 10] = None
```
| github_jupyter |
#### Dataset
```
import numpy as np
np.random.seed(42)
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.datasets import load_breast_cancer
dataset = load_breast_cancer()
print(f"ClassNames: {dataset.target_names}")
print(f"DESCR:\n{dataset.DESCR}")
x = dataset.data
y = dataset.target
print(f"x-shape: {x.shape}")
```
#### PCA
```
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
scaler = StandardScaler()
scaler.fit(x)
# StandardScaler.fit(x)
x_mean = np.mean(x, axis=0)
x_std = np.std(x, axis=0)
print(f"x mean:\n{x_mean}")
print(f"x std:\n{x_std}")
x_standardized = scaler.transform(x)
print(x_standardized[0])
# StandardScaler.transform(x)
x_ = (x - x_mean) / x_std
print(x_[0])
n_components = 15
pca = PCA(n_components=n_components, copy=True)
pca.fit(x_standardized)
x_pca = pca.transform(x_standardized)
print(f"Components:\n{pca.components_}")
print(f"Explained Variance:\n{pca.explained_variance_}")
print(f"Explained Variance Ratio:\n{pca.explained_variance_ratio_}")
print(f"Sum of Exmplained Variance Ratio:\n{sum(pca.explained_variance_ratio_)}")
colors = ["red", "blue"]
for index, point in enumerate(x_pca):
plt.scatter(point[0], point[1], color=colors[y[index]])
plt.show()
```
### Coding Exercise
```
# Aufgabe 1:
# Finde die Anzahl an Dimensionen, um 90% der Varianz "zu erklären"
for n_components in range(1, 29):
pca = PCA(n_components=n_components, copy=True)
pca.fit(x_standardized)
explained_variance_ratio = sum(pca.explained_variance_ratio_)
print(f"Sum of Explained Variance Ratio: {round(explained_variance_ratio, 4)} with: {n_components} components.")
if explained_variance_ratio > 0.90:
break
else:
best_explained_variance_ratio = explained_variance_ratio
# Aufgabe 2:
# Wende das gefundene Setup auf die Daten an
n_components = 7
pca = PCA(n_components=n_components, copy=True)
pca.fit(x_standardized)
x_pca = pca.transform(x_standardized)
# Aufgabe 3:
# Split das Dataset in ein Train and Testset
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x_pca, y, test_size=0.30)
# Aufgabe 4:
# Wende das KNN-Verfahren an
from sklearn.neighbors import KNeighborsClassifier
best_score = 0.0
for i in range(1, 11):
n_neighbors = i
neigh = KNeighborsClassifier(n_neighbors=n_neighbors)
neigh.fit(x_train, y_train)
score = neigh.score(x_test, y_test)
if score > best_score:
best_score = score
print("Score: ", score, " with: ", n_neighbors, " neighbors.")
# Aufgabe 5:
# Wende das KNN ohne Normalisierung und PCA an
dataset = load_breast_cancer()
x = dataset.data
y = dataset.target
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.30)
best_score = 0.0
for i in range(1, 11):
n_neighbors = i
neigh = KNeighborsClassifier(n_neighbors=n_neighbors)
neigh.fit(x_train, y_train)
score = neigh.score(x_test, y_test)
if score > best_score:
best_score = score
print("Score: ", score, " with: ", n_neighbors, " neighbors.")
```
| github_jupyter |
```
import os, sys, argparse
sys.path.append(os.path.dirname(os.path.abspath(os.path.dirname('..'))))
from torchvision.models import resnet50
from thop import profile
import torch
import torch.nn as nn
from models import TSN
from opts import parser
train_path="data/something_train.txt"
val_path="data/something_val.txt"
dataset_name="something"
netType1="TSM"
netType2="MS"
batch_size=1
learning_rate=0.01
num_segments_8=8
num_segments_16=16
num_segments_32=32
num_segments_128=128
mode=1
dropout=0.3
iter_size=1
num_workers=5
input1 = torch.rand(num_segments_8,3,224,224).cuda()
input2 = torch.rand(num_segments_16,3,224,224).cuda()
input3 = torch.rand(num_segments_32,3,224,224).cuda()
input4 = torch.rand(num_segments_128,3,224,224).cuda()
sys.argv = ['main.py', dataset_name, 'RGB', train_path, val_path, '--arch',
str(netType1), '--num_segments', str(num_segments_8), '--mode', str(mode),
'--gd', '200', '--lr', str(learning_rate), '--lr_steps',
'20', '30', '--epochs', '35', '-b', str(batch_size), '-i',
str(iter_size), '-j', str(num_workers), '--dropout',
str(dropout),
'--consensus_type', 'avg', '--eval-freq', '1', '--rgb_prefix', 'img_',
'--pretrained_parts', 'finetune', '--no_partialbn',
'-p', '20', '--nesterov', 'True']
args = parser.parse_args()
args_dict = args.__dict__
print("------------------------------------")
print(args.arch+" Configurations:")
for key in args_dict.keys():
print("- {}: {}".format(key, args_dict[key]))
print("------------------------------------")
if args.dataset == 'ucf101':
num_class = 101
rgb_read_format = "{:05d}.jpg"
elif args.dataset == 'hmdb51':
num_class = 51
rgb_read_format = "{:05d}.jpg"
elif args.dataset == 'kinetics':
num_class = 400
rgb_read_format = "{:05d}.jpg"
elif args.dataset == 'something':
num_class = 174
rgb_read_format = "{:05d}.jpg"
elif args.dataset == 'tinykinetics':
num_class = 150
rgb_read_format = "{:05d}.jpg"
else:
raise ValueError('Unknown dataset '+args.dataset)
TSM_8frame = TSN(num_class, args.num_segments, args.pretrained_parts, args.modality,
base_model=netType1,
consensus_type=args.consensus_type, dropout=args.dropout, partial_bn=not args.no_partialbn).cuda()
MS_8frame = TSN(num_class, args.num_segments, args.pretrained_parts, args.modality,
base_model=netType2,
consensus_type=args.consensus_type, dropout=args.dropout, partial_bn=not args.no_partialbn).cuda()
args.num_segments=num_segments_16
TSM_16frame = TSN(num_class, args.num_segments, args.pretrained_parts, args.modality,
base_model=netType1,
consensus_type=args.consensus_type, dropout=args.dropout, partial_bn=not args.no_partialbn).cuda()
MS_16frame = TSN(num_class, args.num_segments, args.pretrained_parts, args.modality,
base_model=netType2,
consensus_type=args.consensus_type, dropout=args.dropout, partial_bn=not args.no_partialbn).cuda()
# temperature
temperature = 100
flops1, params1 = profile(TSM_8frame, inputs=(input1, temperature), verbose=False)
flops2, params2 = profile(MS_8frame, inputs=(input1, temperature), verbose=False)
flops3, params3 = profile(TSM_16frame, inputs=(input2, temperature), verbose=False)
flops4, params4 = profile(MS_16frame, inputs=(input2, temperature), verbose=False)
def human_format(num):
magnitude = 0
while abs(num) >= 1000:
magnitude += 1
num /= 1000.0
# add more suffixes if you need them
return '%.3f%s' % (num, ['', 'K', 'M', 'G', 'T', 'P'][magnitude])
print('Models\tFrames\tFLOPs\tParams')
print('='*30)
print('%s\t%d\t%s\t%s' % (netType1, num_segments_8, human_format(flops1), human_format(params1)))
print('%s\t%d\t%s\t%s' % (netType2, num_segments_8, human_format(flops2), human_format(params2)))
print('%s\t%d\t%s\t%s' % (netType1, num_segments_16, human_format(flops3), human_format(params3)))
print('%s\t%d\t%s\t%s' % (netType2, num_segments_16, human_format(flops4), human_format(params4)))
```
| github_jupyter |
```
import torch
import torchvision
from torch import nn
from torch.utils.data import DataLoader
from torchvision import transforms
from torchvision.datasets import MNIST
from matplotlib import pyplot as plt
import torch.nn.functional as F
from torchflare.experiments import Experiment, ModelConfig
%load_ext nb_black
train_dataset = MNIST(root='./mnist_data/', train=True, transform=transforms.ToTensor(), download=True)
test_dataset = MNIST(root='./mnist_data/', train=False, transform=transforms.ToTensor(), download=False)
train_dataset, val_dataset = torch.utils.data.random_split(
train_dataset, [50000, 10000]
)
bs = 64
train_loader = torch.utils.data.DataLoader(
dataset=train_dataset, batch_size=bs, shuffle=True
)
val_loader = torch.utils.data.DataLoader(
dataset=val_dataset, batch_size=bs, shuffle=True
)
test_loader = torch.utils.data.DataLoader(
dataset=test_dataset, batch_size=bs, shuffle=False
)
class VAE(nn.Module):
def __init__(self, d):
super().__init__()
self.d = d
self.encoder = nn.Sequential(
nn.Linear(784, self.d ** 2), nn.ReLU(), nn.Linear(self.d ** 2, self.d * 2)
)
self.decoder = nn.Sequential(
nn.Linear(self.d, self.d ** 2),
nn.ReLU(),
nn.Linear(self.d ** 2, 784),
nn.Sigmoid(),
)
def reparameterise(self, mu, logvar):
if self.training:
std = logvar.mul(0.5).exp_()
eps = std.data.new(std.size()).normal_()
return eps.mul(std).add_(mu)
else:
return mu
def forward(self, x):
mu_logvar = self.encoder(x.view(-1, 784)).view(-1, 2, self.d)
mu = mu_logvar[:, 0, :]
logvar = mu_logvar[:, 1, :]
z = self.reparameterise(mu, logvar)
return self.decoder(z), mu, logvar
# Reconstruction + β * KL divergence losses summed over all elements and batch
def loss_function(x_hat, x, mu, logvar, β=1):
BCE = nn.functional.binary_cross_entropy(
x_hat, x.view(-1, 784), reduction='sum'
)
KLD = 0.5 * torch.sum(logvar.exp() - logvar - 1 + mu.pow(2))
return BCE + β * KLD
class VAEExperiment(Experiment):
def batch_step(self):
self.preds = self.state.model(self.batch[self.input_key])
x_hat, mu, logvar = self.preds
self.loss = self.state.criterion(x_hat, self.batch[self.input_key], mu, logvar)
self.loss_per_batch = {"loss": self.loss.item()}
config = ModelConfig(
nn_module=VAE,
module_params={"d": 20},
optimizer="Adam",
optimizer_params={"lr": 3e-3},
criterion=loss_function,
)
exp = VAEExperiment(num_epochs=30, fp16=False, device="cuda", seed=42)
exp.compile_experiment(model_config=config)
exp.fit_loader(train_loader, val_loader)
# Displaying routine
def display_images(in_, out, n=1, label=None, count=False):
for N in range(n):
if in_ is not None:
in_pic = in_.data.cpu().view(-1, 28, 28)
plt.figure(figsize=(18, 4))
plt.suptitle(label + ' – real test data / reconstructions', color='w', fontsize=16)
for i in range(4):
plt.subplot(1,4,i+1)
plt.imshow(in_pic[i+4*N])
plt.axis('off')
out_pic = out.data.cpu().view(-1, 28, 28)
plt.figure(figsize=(18, 6))
for i in range(4):
plt.subplot(1,4,i+1)
plt.imshow(out_pic[i+4*N])
plt.axis('off')
if count: plt.title(str(4 * N + i), color='w')
N = 16
device = "cuda"
z = torch.randn((N, 20)).to(device)
sample = exp.state.model.decoder(z)
display_images(None, sample, N // 4, count=True)
```
| github_jupyter |
# Artificial Intelligence Nanodegree
## Machine Translation Project
In this notebook, sections that end with **'(IMPLEMENTATION)'** in the header indicate that the following blocks of code will require additional functionality which you must provide. Please be sure to read the instructions carefully!
## Introduction
In this notebook, you will build a deep neural network that functions as part of an end-to-end machine translation pipeline. Your completed pipeline will accept English text as input and return the French translation.
- **Preprocess** - You'll convert text to sequence of integers.
- **Models** Create models which accepts a sequence of integers as input and returns a probability distribution over possible translations. After learning about the basic types of neural networks that are often used for machine translation, you will engage in your own investigations, to design your own model!
- **Prediction** Run the model on English text.
```
%load_ext autoreload
%aimport helper, tests
%autoreload 1
import collections
import helper
import numpy as np
import project_tests as tests
import tensorflow as tf
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Model
from keras.layers import GRU, Input, Dense, TimeDistributed, Activation, RepeatVector, Bidirectional
from keras.layers.embeddings import Embedding
from keras.optimizers import Adam
from keras.losses import sparse_categorical_crossentropy
```
### Verify access to the GPU
The following test applies only if you expect to be using a GPU, e.g., while running in a Udacity Workspace or using an AWS instance with GPU support. Run the next cell, and verify that the device_type is "GPU".
- If the device is not GPU & you are running from a Udacity Workspace, then save your workspace with the icon at the top, then click "enable" at the bottom of the workspace.
- If the device is not GPU & you are running from an AWS instance, then refer to the cloud computing instructions in the classroom to verify your setup steps.
```
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
# Add-on : Check framework versions being used
import tensorflow as tf
import keras as K
import sys
print("Python", sys.version)
print("Tensorflow version", tf.__version__)
print("Keras version", K.__version__)
```
## Dataset
We begin by investigating the dataset that will be used to train and evaluate your pipeline. The most common datasets used for machine translation are from [WMT](http://www.statmt.org/). However, that will take a long time to train a neural network on. We'll be using a dataset we created for this project that contains a small vocabulary. You'll be able to train your model in a reasonable time with this dataset.
### Load Data
The data is located in `data/small_vocab_en` and `data/small_vocab_fr`. The `small_vocab_en` file contains English sentences with their French translations in the `small_vocab_fr` file. Load the English and French data from these files from running the cell below.
```
# Remark : Encoding UTF-8 has been fixed in helper.py
# Load English data
english_sentences = helper.load_data('data/small_vocab_en')
# Load French data
french_sentences = helper.load_data('data/small_vocab_fr')
print('Dataset Loaded')
```
### Files
Each line in `small_vocab_en` contains an English sentence with the respective translation in each line of `small_vocab_fr`. View the first two lines from each file.
```
for sample_i in range(2):
print('small_vocab_en Line {}: {}'.format(sample_i + 1, english_sentences[sample_i]))
print('small_vocab_fr Line {}: {}'.format(sample_i + 1, french_sentences[sample_i]))
```
From looking at the sentences, you can see they have been preprocessed already. The puncuations have been delimited using spaces. All the text have been converted to lowercase. This should save you some time, but the text requires more preprocessing.
### Vocabulary
The complexity of the problem is determined by the complexity of the vocabulary. A more complex vocabulary is a more complex problem. Let's look at the complexity of the dataset we'll be working with.
```
english_words_counter = collections.Counter([word for sentence in english_sentences for word in sentence.split()])
french_words_counter = collections.Counter([word for sentence in french_sentences for word in sentence.split()])
print('{} English words.'.format(len([word for sentence in english_sentences for word in sentence.split()])))
print('{} unique English words.'.format(len(english_words_counter)))
print('10 Most common words in the English dataset:')
print('"' + '" "'.join(list(zip(*english_words_counter.most_common(10)))[0]) + '"')
print()
print('{} French words.'.format(len([word for sentence in french_sentences for word in sentence.split()])))
print('{} unique French words.'.format(len(french_words_counter)))
print('10 Most common words in the French dataset:')
print('"' + '" "'.join(list(zip(*french_words_counter.most_common(10)))[0]) + '"')
```
For comparison, _Alice's Adventures in Wonderland_ contains 2,766 unique words of a total of 15,500 words.
## Preprocess
For this project, you won't use text data as input to your model. Instead, you'll convert the text into sequences of integers using the following preprocess methods:
1. Tokenize the words into ids
2. Add padding to make all the sequences the same length.
Time to start preprocessing the data...
### Tokenize (IMPLEMENTATION)
For a neural network to predict on text data, it first has to be turned into data it can understand. Text data like "dog" is a sequence of ASCII character encodings. Since a neural network is a series of multiplication and addition operations, the input data needs to be number(s).
We can turn each character into a number or each word into a number. These are called character and word ids, respectively. Character ids are used for character level models that generate text predictions for each character. A word level model uses word ids that generate text predictions for each word. Word level models tend to learn better, since they are lower in complexity, so we'll use those.
Turn each sentence into a sequence of words ids using Keras's [`Tokenizer`](https://keras.io/preprocessing/text/#tokenizer) function. Use this function to tokenize `english_sentences` and `french_sentences` in the cell below.
Running the cell will run `tokenize` on sample data and show output for debugging.
```
def tokenize(x):
"""
Tokenize x
:param x: List of sentences/strings to be tokenized
:return: Tuple of (tokenized x data, tokenizer used to tokenize x)
"""
# TODO: Implement
x_tk = Tokenizer()
x_tk.fit_on_texts(x)
return x_tk.texts_to_sequences(x), x_tk
tests.test_tokenize(tokenize)
# Tokenize Example output
text_sentences = [
'The quick brown fox jumps over the lazy dog .',
'By Jove , my quick study of lexicography won a prize .',
'This is a short sentence .']
text_tokenized, text_tokenizer = tokenize(text_sentences)
print(text_tokenizer.word_index)
print()
for sample_i, (sent, token_sent) in enumerate(zip(text_sentences, text_tokenized)):
print('Sequence {} in x'.format(sample_i + 1))
print(' Input: {}'.format(sent))
print(' Output: {}'.format(token_sent))
```
### Padding (IMPLEMENTATION)
When batching the sequence of word ids together, each sequence needs to be the same length. Since sentences are dynamic in length, we can add padding to the end of the sequences to make them the same length.
Make sure all the English sequences have the same length and all the French sequences have the same length by adding padding to the **end** of each sequence using Keras's [`pad_sequences`](https://keras.io/preprocessing/sequence/#pad_sequences) function.
```
def pad(x, length=None):
"""
Pad x
:param x: List of sequences.
:param length: Length to pad the sequence to. If None, use length of longest sequence in x.
:return: Padded numpy array of sequences
"""
# TODO: Implement
if length is None:
# Find the length of the longest sequence/sentence
length = max([len(seq) for seq in x])
return pad_sequences(sequences=x, maxlen=length, padding='post')
tests.test_pad(pad)
# Pad Tokenized output
test_pad = pad(text_tokenized)
for sample_i, (token_sent, pad_sent) in enumerate(zip(text_tokenized, test_pad)):
print('Sequence {} in x'.format(sample_i + 1))
print(' Input: {}'.format(np.array(token_sent)))
print(' Output: {}'.format(pad_sent))
```
### Preprocess Pipeline
Your focus for this project is to build neural network architecture, so we won't ask you to create a preprocess pipeline. Instead, we've provided you with the implementation of the `preprocess` function.
```
def preprocess(x, y):
"""
Preprocess x and y
:param x: Feature List of sentences
:param y: Label List of sentences
:return: Tuple of (Preprocessed x, Preprocessed y, x tokenizer, y tokenizer)
"""
preprocess_x, x_tk = tokenize(x)
preprocess_y, y_tk = tokenize(y)
preprocess_x = pad(preprocess_x)
preprocess_y = pad(preprocess_y)
# Keras's sparse_categorical_crossentropy function requires the labels to be in 3 dimensions
preprocess_y = preprocess_y.reshape(*preprocess_y.shape, 1)
return preprocess_x, preprocess_y, x_tk, y_tk
preproc_english_sentences, preproc_french_sentences, english_tokenizer, french_tokenizer =\
preprocess(english_sentences, french_sentences)
max_english_sequence_length = preproc_english_sentences.shape[1]
max_french_sequence_length = preproc_french_sentences.shape[1]
english_vocab_size = len(english_tokenizer.word_index)
french_vocab_size = len(french_tokenizer.word_index)
print('Data Preprocessed')
print("Max English sentence length:", max_english_sequence_length)
print("Max French sentence length:", max_french_sequence_length)
print("English vocabulary size:", english_vocab_size)
print("French vocabulary size:", french_vocab_size)
```
## Models
In this section, you will experiment with various neural network architectures.
You will begin by training four relatively simple architectures.
- Model 1 is a simple RNN
- Model 2 is a RNN with Embedding
- Model 3 is a Bidirectional RNN
- Model 4 is an optional Encoder-Decoder RNN
After experimenting with the four simple architectures, you will construct a deeper architecture that is designed to outperform all four models.
### Ids Back to Text
The neural network will be translating the input to words ids, which isn't the final form we want. We want the French translation. The function `logits_to_text` will bridge the gab between the logits from the neural network to the French translation. You'll be using this function to better understand the output of the neural network.
```
def logits_to_text(logits, tokenizer):
"""
Turn logits from a neural network into text using the tokenizer
:param logits: Logits from a neural network
:param tokenizer: Keras Tokenizer fit on the labels
:return: String that represents the text of the logits
"""
index_to_words = {id: word for word, id in tokenizer.word_index.items()}
index_to_words[0] = '<PAD>'
return ' '.join([index_to_words[prediction] for prediction in np.argmax(logits, 1)])
print('`logits_to_text` function loaded.')
# Add-on : Collect logs for Tensorboard
from keras.callbacks import TensorBoard
from time import time
tensorboard = TensorBoard(log_dir="logs/{}".format(time()), histogram_freq=1, write_graph=True)
```
### Model 1: RNN (IMPLEMENTATION)

A basic RNN model is a good baseline for sequence data. In this model, you'll build a RNN that translates English to French.
```
def simple_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a basic RNN on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Build the layers
input_seq = Input(shape=input_shape[1:])
rnn = GRU(units=english_vocab_size, return_sequences=True)(input_seq)
logits = TimeDistributed(Dense(units=french_vocab_size))(rnn)
model = Model(input_seq, Activation('softmax')(logits))
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(lr=1e-3),
metrics=['accuracy'])
return model
tests.test_simple_model(simple_model)
# Pad and Reshape the input to work with a basic RNN
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
# Train the neural network
simple_rnn_model = simple_model(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size+1,
french_vocab_size+1)
simple_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024, epochs=10, validation_split=0.2)
# Print prediction(s)
print(logits_to_text(simple_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
```
### Model 2: Embedding (IMPLEMENTATION)

You've turned the words into ids, but there's a better representation of a word. This is called word embeddings. An embedding is a vector representation of the word that is close to similar words in n-dimensional space, where the n represents the size of the embedding vectors.
In this model, you'll create a RNN model using embedding.
```
def embed_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a RNN model using word embedding on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Implement
#print("Debug input_shape =" , input_shape, " Input length=", input_shape[1:][0])
#print("Debug output_sequence_length =" , output_sequence_length)
#print("Debug english_vocab_size =" , english_vocab_size)
#print("Debug french_vocab_size =" , french_vocab_size)
# Hyperparameters
embedding_size = 128
rnn_cells = 200
dropout = 0.0
learning_rate = 1e-3
# Sequential Model
#from keras.models import Sequential
#model = Sequential()
#model.add(Embedding(english_vocab_size, embedding_size, input_length=input_shape[1:][0]))
#model.add(GRU(rnn_cells, dropout=dropout, return_sequences=True))
#model.add(Dense(french_vocab_size, activation='softmax'))
#print(model.summary())
# model's Functional equivalent
input_seq = Input(shape=input_shape[1:])
embedded_seq = Embedding(input_dim = english_vocab_size,
output_dim = embedding_size,
input_length=input_shape[1:][0])(input_seq)
rnn = GRU(units=rnn_cells, dropout=dropout, return_sequences=True)(embedded_seq)
logits = TimeDistributed(Dense(units=french_vocab_size))(rnn)
model = Model(input_seq, Activation('softmax')(logits))
#print(model.summary())
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(lr=learning_rate),
metrics=['accuracy'])
return model
tests.test_embed_model(embed_model)
# Pad the input to work with the Embedding layer
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
#print("Debug tmp_x shape=", tmp_x.shape )
# Train the neural network
embed_rnn_model = embed_model(input_shape = tmp_x.shape,
output_sequence_length = max_french_sequence_length,
english_vocab_size = english_vocab_size+1,
french_vocab_size = french_vocab_size+1)
embed_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024, epochs=10, validation_split=0.2)
# Print prediction(s)
print(logits_to_text(embed_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
```
### Model 3: Bidirectional RNNs (IMPLEMENTATION)

One restriction of a RNN is that it can't see the future input, only the past. This is where bidirectional recurrent neural networks come in. They are able to see the future data.
```
def bd_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a bidirectional RNN model on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Implement
#print("Debug input_shape =" , input_shape)
#print("Debug output_sequence_length =" , output_sequence_length)
#print("Debug english_vocab_size =" , english_vocab_size)
#print("Debug french_vocab_size =" , french_vocab_size)
# Hyperparameters
dropout = 0.0
learning_rate = 1e-3
# Choose Sequential or Functional API implementation ('seq' or 'func')
impl='seq'
if impl=='func':
# Sequential Model
print("Using Sequential model (Note: this version makes the unitary test to fail: Disable tests to use it)")
from keras.models import Sequential
model = Sequential()
model.add(Bidirectional(GRU(english_vocab_size, dropout=dropout, return_sequences=True)))
model.add(Dense(french_vocab_size, activation='softmax'))
else:
# model's Functional equivalent
# Note : we could have also used "Bidirectional(GRU(...))" instead of buidling the Bidirectional RNNS manually
print("Using Functional API")
from keras.layers import concatenate, add
input_seq = Input(shape=input_shape[1:])
right_rnn = GRU(units=english_vocab_size, return_sequences=True, go_backwards=False)(input_seq)
left_rnn = GRU(units=english_vocab_size, return_sequences=True, go_backwards=True)(input_seq)
# Choose how to merge the 2 rnn layers : add or concatenate
#logits = TimeDistributed(Dense(units=french_vocab_size))(add([right_rnn, left_rnn]))
logits = TimeDistributed(Dense(units=french_vocab_size))(concatenate([right_rnn, left_rnn]))
model = Model(input_seq, Activation('softmax')(logits))
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(lr=learning_rate),
metrics=['accuracy'])
return model
tests.test_bd_model(bd_model)
# TODO: Train and Print prediction(s)
# Pad and Reshape the input to work with a RNN without an Embedding layer
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
#print("Debug tmp_x shape=", tmp_x.shape )
# Train the neural network
bd_rnn_model = bd_model(input_shape = tmp_x.shape,
output_sequence_length = max_french_sequence_length,
english_vocab_size = english_vocab_size+1,
french_vocab_size = french_vocab_size+1)
#print(model.summary())
bd_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024, epochs=10, validation_split=0.2)
# Print prediction(s)
print(logits_to_text(bd_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
```
### Model 4: Encoder-Decoder (OPTIONAL)
Time to look at encoder-decoder models. This model is made up of an encoder and decoder. The encoder creates a matrix representation of the sentence. The decoder takes this matrix as input and predicts the translation as output.
Create an encoder-decoder model in the cell below.
```
def encdec_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train an encoder-decoder model on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# OPTIONAL: Implement
#print("Debug input_shape =" , input_shape, " input_shape[1:] =", input_shape[1:], " Input length=", input_shape[1:][0])
#print("Debug output_sequence_length =" , output_sequence_length)
#print("Debug english_vocab_size =" , english_vocab_size)
#print("Debug french_vocab_size =" , french_vocab_size)
# Hyperparameters
embedding_size = 128
rnn_cells = 200
dropout = 0.0
learning_rate = 1e-3
from keras.layers import LSTM
# Input
encoder_input_seq = Input(shape=input_shape[1:], name="enc_input")
# Encoder (Return the internal states of the RNN -> 1 hidden state for GRU cells, 2 hidden states for LSTM cells))
encoder_output, state_t = GRU(units=rnn_cells,
dropout=dropout,
return_sequences=False,
return_state=True,
name="enc_rnn")(encoder_input_seq)
#or for LSTM cells: encoder_output, state_h, state_c = LSTM(...)
# Decoder Input
decoder_input_seq = RepeatVector(output_sequence_length)(encoder_output)
# Decoder RNN (Take the encoder returned states as initial states)
decoder_out = GRU(units=rnn_cells,
dropout=dropout,
return_sequences=True,
return_state=False)(decoder_input_seq, initial_state=state_t)
#or for LSTM cells: (decoder_input_seq, initial_state=[state_h, state_c])
# Decoder output
logits = TimeDistributed(Dense(units=french_vocab_size))(decoder_out)
# Model
model = Model(encoder_input_seq, Activation('softmax')(logits))
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(lr=learning_rate),
metrics=['accuracy'])
return model
# Unitary tests
tests.test_encdec_model(encdec_model)
# OPTIONAL: Train and Print prediction(s)
# Pad and Reshape the input to work with the Embedding layer
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
#print("Debug tmp_x shape=", tmp_x.shape )
# Train the neural network
encdec_rnn_model = encdec_model(input_shape = tmp_x.shape,
output_sequence_length = max_french_sequence_length,
english_vocab_size = english_vocab_size+1,
french_vocab_size = french_vocab_size+1)
#print(encdec_rnn_model.summary())
encdec_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024, epochs=10, validation_split=0.2) # callbacks=[tensorboard]
# Print prediction(s)
print(logits_to_text(encdec_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
```
### Model 5: Custom (IMPLEMENTATION)
Use everything you learned from the previous models to create a model that incorporates embedding and a bidirectional rnn into one model.
```
def model_final(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a model that incorporates embedding, encoder-decoder, and bidirectional RNN on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Train the final model
#print("Debug input_shape =" , input_shape, " input_shape[1:] =", input_shape[1:], " Input length=", input_shape[1:][0])
#print("Debug output_sequence_length =" , output_sequence_length)
#print("Debug english_vocab_size =" , english_vocab_size)
#print("Debug french_vocab_size =" , french_vocab_size)
## Model_final is a seq2seq model (encoder-decoder) using embedding and bidirectional LSTM
# Hyperparameters
embedding_size = 128
rnn_cells = 300
dropout = 0.2
learning_rate = 1e-3
from keras.layers import LSTM, concatenate
# Input and embedding
encoder_input_seq = Input(shape=input_shape[1:])
embedded_input_seq = Embedding(input_dim = english_vocab_size,
output_dim = embedding_size,
input_length=input_shape[1:][0])(encoder_input_seq)
## Note Bidirectional LSTM is used on the encoder side (see https://arxiv.org/pdf/1609.08144.pdf )
## Alternate version : Encoder RNN Bidirectional layer (Using the Keras Bidirectional layer wrappers)
#encoder_output, forward_state_h, forward_state_c, backward_state_h, backward_state_c = Bidirectional(LSTM(units=rnn_cells,
# dropout=dropout,
# return_sequences=False,
# return_state=True))(embedded_input_seq)
# Encoder Forward RNN layer
encoder_forward_output, forward_state_h, forward_state_c = LSTM(units=rnn_cells,
dropout=dropout,
return_sequences=False,
return_state=True,
go_backwards=False)(embedded_input_seq)
# Encoder backward RNN layer
encoder_backward_output, backward_state_h, backward_state_c = LSTM(units=rnn_cells,
dropout=dropout,
return_sequences=False,
return_state=True,
go_backwards=True)(embedded_input_seq)
# Encoder output and states : Merge the LSTM Forward and Backward ouputs (using 'concatenate' method)
state_h = concatenate([forward_state_h, backward_state_h])
state_c = concatenate([forward_state_c, backward_state_c])
encoder_output = concatenate([encoder_forward_output, encoder_backward_output])
# Decoder Input
decoder_input_seq = RepeatVector(output_sequence_length)(encoder_output)
# Decoder RNN layer
# Note : we need twice more LSTM cells as we have concatenated backward and forward LSTM layers in encoder
decoder_output = LSTM(units=rnn_cells*2,
dropout=dropout,
return_sequences=True,
return_state=False,
go_backwards=False)(decoder_input_seq, initial_state=[state_h, state_c])
# Decoder output
logits = TimeDistributed(Dense(units=french_vocab_size))(decoder_output)
# Model
model = Model(encoder_input_seq, Activation('softmax')(logits))
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(lr=learning_rate),
metrics=['accuracy'])
return model
tests.test_model_final(model_final)
print('Final Model Loaded\n')
# Train and Print prediction(s)
# Pad the input to work with the Embedding layer
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
#print("Debug tmp_x shape=", tmp_x.shape )
# Train the neural network
final_rnn_model = model_final(input_shape = tmp_x.shape,
output_sequence_length = max_french_sequence_length,
english_vocab_size = english_vocab_size+1,
french_vocab_size = french_vocab_size+1)
print(final_rnn_model.summary(line_length=125))
final_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024, epochs=10, validation_split=0.2)
# Print prediction(s)
print(logits_to_text(final_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
```
## Prediction (IMPLEMENTATION)
```
def final_predictions(x, y, x_tk, y_tk):
"""
Gets predictions using the final model
:param x: Preprocessed English data
:param y: Preprocessed French data
:param x_tk: English tokenizer
:param y_tk: French tokenizer
"""
# TODO: Train neural network using model_final
model = model_final(input_shape = x.shape,
output_sequence_length = y.shape[1],
english_vocab_size = len(x_tk.word_index)+1,
french_vocab_size = len(y_tk.word_index)+1)
model.fit(x, y, batch_size=1024, epochs=20, validation_split=0.2)
## DON'T EDIT ANYTHING BELOW THIS LINE
y_id_to_word = {value: key for key, value in y_tk.word_index.items()}
y_id_to_word[0] = '<PAD>'
sentence = 'he saw a old yellow truck'
sentence = [x_tk.word_index[word] for word in sentence.split()]
sentence = pad_sequences([sentence], maxlen=x.shape[-1], padding='post')
sentences = np.array([sentence[0], x[0]])
predictions = model.predict(sentences, len(sentences))
print('Sample 1:')
print(' '.join([y_id_to_word[np.argmax(x)] for x in predictions[0]]))
print('Il a vu un vieux camion jaune')
print('Sample 2:')
print(' '.join([y_id_to_word[np.argmax(x)] for x in predictions[1]]))
print(' '.join([y_id_to_word[np.max(x)] for x in y[0]]))
final_predictions(preproc_english_sentences, preproc_french_sentences, english_tokenizer, french_tokenizer)
```
## Submission
When you're ready to submit, complete the following steps:
1. Review the [rubric](https://review.udacity.com/#!/rubrics/1004/view) to ensure your submission meets all requirements to pass
2. Generate an HTML version of this notebook
- Run the next cell to attempt automatic generation (this is the recommended method in Workspaces)
- Navigate to **FILE -> Download as -> HTML (.html)**
- Manually generate a copy using `nbconvert` from your shell terminal
```
$ pip install nbconvert
$ python -m nbconvert machine_translation.ipynb
```
3. Submit the project
- If you are in a Workspace, simply click the "Submit Project" button (bottom towards the right)
- Otherwise, add the following files into a zip archive and submit them
- `helper.py`
- `machine_translation.ipynb`
- `machine_translation.html`
- You can export the notebook by navigating to **File -> Download as -> HTML (.html)**.
### Generate the html
**Save your notebook before running the next cell to generate the HTML output.** Then submit your project.
```
# Save before you run this cell!
!!jupyter nbconvert *.ipynb
```
## Optional Enhancements
This project focuses on learning various network architectures for machine translation, but we don't evaluate the models according to best practices by splitting the data into separate test & training sets -- so the model accuracy is overstated. Use the [`sklearn.model_selection.train_test_split()`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) function to create separate training & test datasets, then retrain each of the models using only the training set and evaluate the prediction accuracy using the hold out test set. Does the "best" model change?
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
S, P = np.load("../data/dataset.npy")
molecules = np.load("../data/molecules.npy")
def extract_triu(A):
"""Extracts the upper triangular part of the matrix.
Input can be matrix, will be reshaped if it is not.
"""
return A.reshape(dim, dim)[np.triu_indices(dim)]
def reconstruct_from_triu(A_flat):
"""Reconstructus the full symmetric matrix (dim x dim, not
flattened out) from the flattend elements of the upper
triag of a symmetric matrix!"""
result = np.zeros((dim, dim))
result[np.triu_indices(dim)] = A_flat
return result + result.T - np.diag(np.diag(result))
from SCFInitialGuess.utilities.dataset import Dataset
dim = 26
dim_triu = int(dim * (dim + 1) / 2)
ind_cut = 150
index = np.arange(200)
np.random.shuffle(index)
S_triu = list(map(extract_triu, S))
P_triu = list(map(extract_triu, P))
S_test = np.array(S_triu)[index[150:]]
P_test = np.array(P_triu)[index[150:]]
molecules_test = [molecules[index[i]] for i in range(150, 200)]
S_train = np.array(S_triu)[index[:150]]
P_train = np.array(P_triu)[index[:150]]
molecules_train = [molecules[index[i]] for i in range(150)]
dataset = Dataset(np.array(S_train), np.array(P_train), split_test=0.0)
dataset.testing = (Dataset.normalize(S_test, mean=dataset.x_mean, std=dataset.x_std)[0], P_test)
from SCFInitialGuess.nn.networks import EluTrNNN
from SCFInitialGuess.nn.training import Trainer
from SCFInitialGuess.nn.cost_functions import RegularizedMSE
trainer = Trainer(
EluTrNNN([dim_triu, 400, 400, 400, 400, dim_triu]),
cost_function=RegularizedMSE(alpha=1e-7),
optimizer=tf.train.AdamOptimizer(learning_rate=1e-3)
)
trainer.setup()
network, sess = trainer.train(
dataset,
convergence_threshold=1e-7
)
graph = trainer.graph
#from SCFInitialGuess.utilities.analysis import prediction_scatter
import matplotlib.pyplot as plt
with graph.as_default():
plt.scatter(
dataset.testing[1].flatten(),
network.run(sess, dataset.testing[0]).flatten()
)
plt.show()
def mc_wheeny_purification(p,s):
p = p.reshape(dim, dim)
s = s.reshape(dim, dim)
return (3 * np.dot(np.dot(p, s), p) - np.dot(np.dot(np.dot(np.dot(p, s), p), s), p)) / 2
def multi_mc_wheeny(p, s, n_max=4):
for i in range(n_max):
p = mc_wheeny_purification(p, s)
return p
def idemp_error(p, s):
p = p.reshape(dim, dim)
s = s.reshape(dim, dim)
return np.mean(np.abs(np.dot(np.dot(p, s), p) - 2 * p))
P_NN_multi = []
for (s, p) in zip(*dataset.testing):
#for (s, p) in zip(S_test, P_test):
#s_norm = s.reshape(1, dim**2)
s_raw = reconstruct_from_triu(dataset.inverse_input_transform(s))
p_raw = reconstruct_from_triu(p)
print("Orig: {:0.3E}".format(idemp_error(p_raw, s_raw)))
with graph.as_default():
p_nn = reconstruct_from_triu(network.run(sess, s.reshape(1, dim_triu)))
print("NN: {:0.3E}".format(idemp_error(p_nn, s_raw)))
print("NN pruified: {:0.3E}".format(idemp_error(mc_wheeny_purification(p_nn, s_raw), s_raw)))
p_nn_multi = multi_mc_wheeny(p_nn, s_raw, n_max=5)
P_NN_multi.append(extract_triu(p_nn_multi))
print("NN multified: {:0.3E}".format(idemp_error(p_nn_multi, s_raw)))
print("Error NN before: {:0.3E}".format(np.mean(np.abs(p_raw - p_nn))))
print("Error NN multifed: {:0.3E}".format(np.mean(np.abs(p_raw - p_nn_multi))))
print("---------------------------")
with graph.as_default():
plt.scatter(
dataset.testing[1].flatten(),
network.run(sess, dataset.testing[0]).flatten(),
label="orig"
)
plt.scatter(
dataset.testing[1].flatten(),
np.array(P_NN_multi).flatten(),
label="pruified"
)
plt.legend()
plt.show()
from pyscf.scf import hf
from SCFInitialGuess.utilities.analysis import prediction_scatter
keys = ["minao", "noise", "nn", "nn_purified"]
iterations = {}
for k in keys:
iterations[k] = []
for i, (molecule, p) in enumerate(zip(molecules_test, P_test)):
mol = molecule.get_pyscf_molecule()
print("Calculating: " + str(i + 1) + "/" + str(len(molecules_test)))
guesses = {}
s_raw = hf.get_ovlp(mol)
s_norm = dataset.input_transformation(extract_triu(s_raw)).reshape(1, dim_triu)
# pyscf init guess
p_minao = hf.init_guess_by_minao(mol)
guesses["minao"] = p_minao
# P_actual wi noise
p_raw = reconstruct_from_triu(p)
p_noise = p_raw + np.random.randn(dim, dim) * 1e-3
guesses["noise"] = p_noise
with graph.as_default():
p_orig = reconstruct_from_triu(network.run(sess, s_norm)).astype('float64')
guesses["nn"] = p_orig
p_purified = multi_mc_wheeny(p_orig, s_raw, n_max=5)
guesses["nn_purified"] = p_purified
# check errors
print("Accuracy (MSE):")
print(" -Noise: {:0.3E}".format(np.mean(np.abs(p_raw - p_noise))))
print(" -NN: {:0.3E}".format(np.mean(np.abs(p_raw - p_orig))))
print(" -Pruif: {:0.3E}".format(np.mean(np.abs(p_raw - p_purified))))
print(" -minao: {:0.3E}".format(np.mean(np.abs(p_raw - p_minao))))
print("Idempotency:")
print(" -Noise: {:0.3E}".format(idemp_error(p_noise, s_raw)))
print(" -Orig: {:0.3E}".format(idemp_error(p_orig, s_raw)))
print(" -Purif: {:0.3E}".format(idemp_error(p_purified, s_raw)))
print(" -minao: {:0.3E}".format(idemp_error(p_minao, s_raw)))
for (key, guess) in guesses.items():
mf = hf.RHF(mol)
mf.diis = None
mf.verbose = 1
mf.kernel(dm0=guess)
iterations[key].append(mf.iterations)
for k in keys:
iterations[k] = np.array(iterations[k])
for key, val in iterations.items():
print(key + ": " + str(val.mean()))
from SCFInitialGuess.utilities.analysis import iterations_histogram
for key, val in iterations.items():
hist, bins = np.histogram(val)
center = (bins[:-1] + bins[1:]) / 2
#axes[i].bar(center, hist, label=name)
plt.bar(center, hist, label=key)
plt.legend()
plt.show()
plt.show()
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Model Remediation Case Study
<div class="devsite-table-wrapper"><table class="tfo-notebook-buttons" align="left">
<td><a target="_blank" href="https://tensorflow.org/responsible_ai/model_remediation/min_diff/tutorials/min_diff_keras">
<img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/model-remediation/blob/master/docs/min_diff/tutorials/min_diff_keras.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png">Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/model-remediation/blob/master/docs/min_diff/tutorials/min_diff_keras.ipynb">
<img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">View source on GitHub</a>
</td>
<td>
<a target="_blank" href="https://storage.googleapis.com/tensorflow_docs/model-remediation/docs/min_diff/tutorials/min_diff_keras.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table></div>
In this notebook, we’ll train a text classifier to identify written content that could be considered toxic or harmful, and apply MinDiff to remediate some fairness concerns. In our workflow, we will:
1. Evaluate our baseline model’s performance on text containing references to sensitive groups.
2. Improve performance on any underperforming groups by training with MinDiff.
3. Evaluate the new model’s performance on our chosen metric.
Our purpose is to demonstrate usage of the MinDiff technique with a very minimal workflow, not to lay out a principled approach to fairness in machine learning. As such, our evaluation will only focus on one sensitive category and a single metric. We also don’t address potential shortcomings in the dataset, nor tune our configurations. In a production setting, you would want to approach each of these with rigor. For more information on evaluating for fairness, see [this guide](https://www.tensorflow.org/responsible_ai/fairness_indicators/guide/guidance).
## Setup
We begin by installing Fairness Indicators and TensorFlow Model Remediation.
```
#@title Installs
!pip install --upgrade tensorflow-model-remediation
!pip install --upgrade fairness-indicators
```
Import all necessary components, including MinDiff and Fairness Indicators for evaluation.
```
#@title Imports
import copy
import os
import requests
import tempfile
import zipfile
import tensorflow_model_remediation.min_diff as md
from tensorflow_model_remediation.tools.tutorials_utils import min_diff_keras_utils
from fairness_indicators.tutorial_utils import util as fi_util
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
from tensorflow_model_analysis.addons.fairness.view import widget_view
```
We use a utility function to download the preprocessed data and prepare the labels to match the model’s output shape. The function also downloads the data as TFRecords to make later evaluation quicker. Alternatively, you may convert the Pandas DataFrame into TFRecords with any available utility conversion function.
```
# We use a helper utility to preprocessed data for convenience and speed.
data_train, data_validate, validate_tfrecord_file, labels_train, labels_validate = min_diff_keras_utils.download_and_process_civil_comments_data()
```
We define a few useful constants. We will train the model on the `’comment_text’` feature, with our target label as `’toxicity’`. Note that the batch size here is chosen arbitrarily, but in a production setting you would need to tune it for best performance.
```
TEXT_FEATURE = 'comment_text'
LABEL = 'toxicity'
BATCH_SIZE = 512
```
Set random seeds. (Note that this does not fully stabilize results.)
```
#@title Seeds
np.random.seed(1)
tf.random.set_seed(1)
```
## Define and train the baseline model
To reduce runtime, we use a pretrained model by default. It is a simple Keras sequential model with an initial embedding and convolution layers, outputting a toxicity prediction. If you prefer, you can change this and train from scratch using our utility function to create the model. (Note that since your environment is likely different from ours, you would need to customize the tuning and evaluation thresholds.)
```
use_pretrained_model = True #@param {type:"boolean"}
if use_pretrained_model:
URL = 'https://storage.googleapis.com/civil_comments_model/baseline_model.zip'
BASE_PATH = tempfile.mkdtemp()
ZIP_PATH = os.path.join(BASE_PATH, 'baseline_model.zip')
MODEL_PATH = os.path.join(BASE_PATH, 'tmp/baseline_model')
r = requests.get(URL, allow_redirects=True)
open(ZIP_PATH, 'wb').write(r.content)
with zipfile.ZipFile(ZIP_PATH, 'r') as zip_ref:
zip_ref.extractall(BASE_PATH)
baseline_model = tf.keras.models.load_model(
MODEL_PATH, custom_objects={'KerasLayer' : hub.KerasLayer})
else:
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
loss = tf.keras.losses.BinaryCrossentropy()
baseline_model = min_diff_keras_utils.create_keras_sequential_model()
baseline_model.compile(optimizer=optimizer, loss=loss, metrics=['accuracy'])
baseline_model.fit(x=data_train[TEXT_FEATURE],
y=labels_train,
batch_size=BATCH_SIZE,
epochs=20)
```
We save the model in order to evaluate using [Fairness Indicators](https://www.tensorflow.org/responsible_ai/fairness_indicators).
```
base_dir = tempfile.mkdtemp(prefix='saved_models')
baseline_model_location = os.path.join(base_dir, 'model_export_baseline')
baseline_model.save(baseline_model_location, save_format='tf')
```
Next we run Fairness Indicators. As a reminder, we’re just going to perform sliced evaluation for comments referencing one category, *religious groups*. In a production environment, we recommend taking a thoughtful approach to determining which categories and metrics to evaluate across.
To compute model performance, the utility function makes a few convenient choices for metrics, slices, and classifier thresholds.
```
# We use a helper utility to hide the evaluation logic for readability.
base_dir = tempfile.mkdtemp(prefix='eval')
eval_dir = os.path.join(base_dir, 'tfma_eval_result')
eval_result = fi_util.get_eval_results(
baseline_model_location, eval_dir, validate_tfrecord_file)
```
### Render Evaluation Results
```
widget_view.render_fairness_indicator(eval_result)
```
Let’s look at the evaluation results. Try selecting the metric false positive rate (FPR) with threshold 0.450. We can see that the model does not perform as well for some religious groups as for others, displaying a much higher FPR. Note the wide confidence intervals on some groups because they have too few examples. This makes it difficult to say with certainty that there is a significant difference in performance for these slices. We may want to collect more examples to address this issue. We can, however, attempt to apply MinDiff for the two groups that we are confident are underperforming.
We’ve chosen to focus on FPR, because a higher FPR means that comments referencing these identity groups are more likely to be incorrectly flagged as toxic than other comments. This could lead to inequitable outcomes for users engaging in dialogue about religion, but note that disparities in other metrics can lead to other types of harm.
## Define and Train the MinDiff Model
Now, we’ll try to improve the FPR for underperforming religious groups. We’ll attempt to do so using [MinDiff](https://arxiv.org/abs/1910.11779), a remediation technique that seeks to balance error rates across slices of your data by penalizing disparities in performance during training. When we apply MinDiff, model performance may degrade slightly on other slices. As such, our goals with MinDiff will be:
* Improved performance for underperforming groups
* Limited degradation for other groups and overall performance
### Prepare your data
To use MinDiff, we create two additional data splits:
* A split for non-toxic examples referencing minority groups: In our case, this will include comments with references to our underperforming identity terms. We don’t include some of the groups because there are too few examples, leading to higher uncertainty with wide confidence interval ranges.
* A split for non-toxic examples referencing the majority group.
It’s important to have sufficient examples belonging to the underperforming classes. Based on your model architecture, data distribution, and MinDiff configuration, the amount of data needed can vary significantly. In past applications, we have seen MinDiff work well with 5,000 examples in each data split.
In our case, the groups in the minority splits have example quantities of 9,688 and 3,906. Note the class imbalances in the dataset; in practice, this could be cause for concern, but we won’t seek to address them in this notebook since our intention is just to demonstrate MinDiff.
We select only negative examples for these groups, so that MinDiff can optimize on getting these examples right. It may seem counterintuitive to carve out sets of ground truth *negative* examples if we’re primarily concerned with disparities in *false positive rate*, but remember that a false positive prediction is a ground truth negative example that’s incorrectly classified as positive, which is the issue we’re trying to address.
#### Create MinDiff DataFrames
```
# Create masks for the sensitive and nonsensitive groups
minority_mask = data_train.religion.apply(
lambda x: any(religion in x for religion in ('jewish', 'muslim')))
majority_mask = data_train.religion.apply(lambda x: x == "['christian']")
# Select nontoxic examples, so MinDiff will be able to reduce sensitive FP rate.
true_negative_mask = data_train['toxicity'] == 0
data_train_main = copy.copy(data_train)
data_train_sensitive = data_train[minority_mask & true_negative_mask]
data_train_nonsensitive = data_train[majority_mask & true_negative_mask]
```
We also need to convert our Pandas DataFrames into Tensorflow Datasets for MinDiff input. Note that unlike the Keras model API for Pandas DataFrames, using Datasets means that we need to provide the model’s input features and labels together in one Dataset. Here we provide the `'comment_text'` as an input feature and reshape the label to match the model's expected output.
We batch the Dataset at this stage, too, since MinDiff requires batched Datasets. Note that we tune the batch size selection the same way it is tuned for the baseline model, taking into account training speed and hardware considerations while balancing with model performance. Here we have chosen the same batch size for all three datasets but this is not a requirement, although it’s good practice to have the two MinDiff batch sizes be equivalent.
#### Create MinDiff Datasets
```
# Convert the pandas DataFrames to Datasets.
dataset_train_main = tf.data.Dataset.from_tensor_slices(
(data_train_main['comment_text'].values,
data_train_main.pop(LABEL).values.reshape(-1,1) * 1.0)).batch(BATCH_SIZE)
dataset_train_sensitive = tf.data.Dataset.from_tensor_slices(
(data_train_sensitive['comment_text'].values,
data_train_sensitive.pop(LABEL).values.reshape(-1,1) * 1.0)).batch(BATCH_SIZE)
dataset_train_nonsensitive = tf.data.Dataset.from_tensor_slices(
(data_train_nonsensitive['comment_text'].values,
data_train_nonsensitive.pop(LABEL).values.reshape(-1,1) * 1.0)).batch(BATCH_SIZE)
```
### Train and evaluate the model
To train with MinDiff, simply take the original model and wrap it in a MinDiffModel with a corresponding `loss` and `loss_weight`. We are using 1.5 as the default `loss_weight`, but this is a parameter that needs to be tuned for your use case, since it depends on your model and product requirements. You can experiment with changing the value to see how it impacts the model, noting that increasing it pushes the performance of the minority and majority groups closer together but may come with more pronounced tradeoffs.
Then we compile the model normally (using the regular non-MinDiff loss) and fit to train.
#### Train MinDiffModel
```
use_pretrained_model = True #@param {type:"boolean"}
base_dir = tempfile.mkdtemp(prefix='saved_models')
min_diff_model_location = os.path.join(base_dir, 'model_export_min_diff')
if use_pretrained_model:
BASE_MIN_DIFF_PATH = tempfile.mkdtemp()
MIN_DIFF_URL = 'https://storage.googleapis.com/civil_comments_model/min_diff_model.zip'
ZIP_PATH = os.path.join(BASE_PATH, 'min_diff_model.zip')
MIN_DIFF_MODEL_PATH = os.path.join(BASE_MIN_DIFF_PATH, 'tmp/min_diff_model')
DIRPATH = '/tmp/min_diff_model'
r = requests.get(MIN_DIFF_URL, allow_redirects=True)
open(ZIP_PATH, 'wb').write(r.content)
with zipfile.ZipFile(ZIP_PATH, 'r') as zip_ref:
zip_ref.extractall(BASE_MIN_DIFF_PATH)
min_diff_model = tf.keras.models.load_model(
MIN_DIFF_MODEL_PATH, custom_objects={'KerasLayer' : hub.KerasLayer})
min_diff_model.save(min_diff_model_location, save_format='tf')
else:
min_diff_weight = 1.5 #@param {type:"number"}
# Create the dataset that will be passed to the MinDiffModel during training.
dataset = md.keras.utils.input_utils.pack_min_diff_data(
dataset_train_main, dataset_train_sensitive, dataset_train_nonsensitive)
# Create the original model.
original_model = min_diff_keras_utils.create_keras_sequential_model()
# Wrap the original model in a MinDiffModel, passing in one of the MinDiff
# losses and using the set loss_weight.
min_diff_loss = md.losses.MMDLoss()
min_diff_model = md.keras.MinDiffModel(original_model,
min_diff_loss,
min_diff_weight)
# Compile the model normally after wrapping the original model. Note that
# this means we use the baseline's model's loss here.
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
loss = tf.keras.losses.BinaryCrossentropy()
min_diff_model.compile(optimizer=optimizer, loss=loss, metrics=['accuracy'])
min_diff_model.fit(dataset, epochs=20)
min_diff_model.save_original_model(min_diff_model_location, save_format='tf')
```
Next we evaluate the results.
```
min_diff_eval_subdir = os.path.join(base_dir, 'tfma_eval_result')
min_diff_eval_result = fi_util.get_eval_results(
min_diff_model_location,
min_diff_eval_subdir,
validate_tfrecord_file,
slice_selection='religion')
```
To ensure we evaluate a new model correctly, we need to select a threshold the same way that we would the baseline model. In a production setting, this would mean ensuring that evaluation metrics meet launch standards. In our case, we will pick the threshold that results in a similar overall FPR to the baseline model. This threshold may be different from the one you selected for the baseline model. Try selecting false positive rate with threshold 0.400. (Note that the subgroups with very low quantity examples have very wide confidence range intervals and don’t have predictable results.)
```
widget_view.render_fairness_indicator(min_diff_eval_result)
```
Reviewing these results, you may notice that the FPRs for our target groups have improved. The gap between our lowest performing group and the majority group has improved from .024 to .006. Given the improvements we’ve observed and the continued strong performance for the majority group, we’ve satisfied both of our goals. Depending on the product, further improvements may be necessary, but this approach has gotten our model one step closer to performing equitably for all users.
| github_jupyter |
# Load the Dataset
```
import numpy as np
import matplotlib.pyplot as plt
import warnings
from matplotlib.colors import ListedColormap
%matplotlib inline
warnings.filterwarnings('ignore')
# for plotting
cmap2 = ListedColormap(['r', 'k'])
cmap4 = ListedColormap(['k', 'r', 'g', 'b'])
plt.rc("font",family="sans-serif",size=20)
plt.rcParams["font.sans-serif"] = "Arial"
#hold data
mice = list()
synthetic = list()
```
### Synthetic Dataset
* There are 4 clusters in the target dataset (but we do not know their labels *a priori*.
* In the background, all the data pts are from the same distribution, which has different variances in three subspaces.
```
from scipy.stats import ortho_group
np.random.seed(0) # for reproducibility
# In A there are four clusters.
N = 400; D = 30; gap=1.5
rotation = ortho_group.rvs(dim=D)
target_ = np.zeros((N, D))
target_[:,0:10] = np.random.normal(0,10,(N,10))
# group 1
target_[0:100, 10:20] = np.random.normal(-gap,1,(100,10))
target_[0:100, 20:30] = np.random.normal(-gap,1,(100,10))
# group 2
target_[100:200, 10:20] = np.random.normal(-gap,1,(100,10))
target_[100:200, 20:30] = np.random.normal(gap,1,(100,10))
# group 3
target_[200:300, 10:20] = np.random.normal(2*gap,1,(100,10))
target_[200:300, 20:30] = np.random.normal(-gap,1,(100,10))
# group 4
target_[300:400, 10:20] = np.random.normal(2*gap,1,(100,10))
target_[300:400, 20:30] = np.random.normal(gap,1,(100,10))
target_ = target_.dot(rotation)
sub_group_labels_ = [0]*100+[1]*100+[2]*100+[3]*100
background_ = np.zeros((N, D))
background_[:,0:10] = np.random.normal(0,10,(N,10))
background_[:,10:20] = np.random.normal(0,3,(N,10))
background_[:,20:30] = np.random.normal(0,1,(N,10))
background_ = background_.dot(rotation)
data_ = np.concatenate((background_, target_))
labels_ = len(background_)*[0] + len(target_)*[1]
```
### Mice Protein Dataset
```
data = np.genfromtxt('datasets/Data_Cortex_Nuclear.csv',delimiter=',',
skip_header=1,usecols=range(1,78),filling_values=0)
classes = np.genfromtxt('datasets/Data_Cortex_Nuclear.csv',delimiter=',',
skip_header=1,usecols=range(78,81),dtype=None)
```
* Target consists of mice that have been stimulated by shock therapy. Some have Down Syndrome, others don't, but we assume this label is not known to us *a priori*
* Background consists of mice that have not been stimulated by shock therapy, and do not have Down Syndrome
```
target_idx_A = np.where((classes[:,-1]==b'S/C') & (classes[:,-2]==b'Saline') & (classes[:,-3]==b'Control'))[0]
target_idx_B = np.where((classes[:,-1]==b'S/C') & (classes[:,-2]==b'Saline') & (classes[:,-3]==b'Ts65Dn'))[0]
sub_group_labels = len(target_idx_A)*[0] + len(target_idx_B)*[1]
target_idx = np.concatenate((target_idx_A,target_idx_B))
target = data[target_idx]
target = (target-np.mean(target,axis=0)) / np.std(target,axis=0) # standardize the dataset
background_idx = np.where((classes[:,-1]==b'C/S') & (classes[:,-2]==b'Saline') & (classes[:,-3]==b'Control'))
# background_idx = np.where((classes[:,-1]==b'C/S') & (classes[:,-2]==b'Saline') & (classes[:,-3]==b'Ts65Dn'))
background = data[background_idx]
background = (background-np.mean(background,axis=0)) / np.std(background,axis=0) # standardize the dataset
labels = len(background)*[0] + len(target)*[1]
data = np.concatenate((background, target))
```
# Comparing cPCA to Other Dimensionality Reduction Techniques
### PCA and cPCA
(PCA corresponds to the first column, since it is equivalent to cPCA with $\alpha=0$)
```
from contrastive import CPCA
mdl = CPCA()
mdl.fit_transform(target, background, plot=True, active_labels=sub_group_labels)
projected_data = mdl.fit_transform(target, background, plot=False, active_labels=sub_group_labels)
mdl.fit_transform(target_, background_, plot=True, active_labels=sub_group_labels_)
projected_data_ = mdl.fit_transform(target_, background_, plot=False, active_labels=sub_group_labels_)
mice.append(projected_data[1])
mice.append(projected_data[0])
synthetic.append(projected_data_[2])
synthetic.append(projected_data_[0])
```
* PCA is unable to resolve the subgroups of interest in the mice data, while cPCA is able to separate the 2 subgroups with an appropriate (and automatically discovered) value of $\alpha$
* PCA is unable to resolve the subgroups in the synthetic data, while cPCA is able to resolve all 4 with an appropriate (and automatically discovered) value of $\alpha$ (2.15). An alternative value of $\alpha$ (151.18) discovers another interesting projection.
### Supervised PCA
```
from supervised import SupervisedPCAClassifier
mdl = SupervisedPCAClassifier(n_components=2)
projected_data = mdl.fit(data, labels).get_transformed_data(target)
plt.figure()
plt.scatter(*projected_data.T, c=sub_group_labels, cmap=cmap2)
plt.title('Supervised PCA: Mice Data')
projected_data_ = mdl.fit(data_, labels_).get_transformed_data(target_)
plt.figure()
plt.scatter(*projected_data_.T, c=sub_group_labels_, cmap=cmap4)
plt.title('Supervised PCA: Synthetic Data')
mice.append(projected_data)
synthetic.append(projected_data_)
```
* Supervised PCA is unable to resolve the subgroups of interest in the mice data
* Supervised PCA is unable to resolve the subgroups in the synthetic data
### Linear Discriminant Analysis (LDA)
Note: because LDA returns at most $c-1$ components, where $c$ is the number of classes in the data, in this case, the target data is projected onto exactly 1 dimension (the x-dimension). For ease of visualization, we assign a random value to the y-dimension of each data point, and plot them in 2 dimensions.
```
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
mdl = LDA()
projected_data = mdl.fit(data, labels).transform(target)
random_y_values = np.random.random(size=projected_data.shape)
plt.figure()
plt.scatter(projected_data, random_y_values, c=sub_group_labels, cmap=cmap2)
plt.title('Linear Discriminant Analysis: Mice Data')
mice.append(np.array([projected_data,random_y_values]).T)
projected_data_ = mdl.fit(data_, labels_).transform(target_)
random_y_values_ = np.random.random(size=projected_data_.shape)
plt.figure()
plt.scatter(projected_data_, random_y_values_, c=sub_group_labels_, cmap=cmap4)
plt.title('Linear Discriminant Analysis: Synthetic Data')
synthetic.append(np.array([projected_data_,random_y_values_]).T)
```
* LDA is unable to resolve the subgroups of interest in the mice data
* LDA is able to somewhat resolve two pairs of subgroups in the synthetic data, but not all four
### Quadratic Discriminant Analysis (QDA)
Note: Unlike the PCA or LDA, the QDA does not return a subspace on which to project the data for dimensionality reduction. Instead, we plotted each point based according to its posterior class probability.
```
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis as QDA
from numpy import inf
mdl = QDA()
projected_data = mdl.fit(data, labels).predict_log_proba(target)[:,0]
projected_data[projected_data == -inf] = -800
random_y_values = np.random.random(size=projected_data.shape)
plt.figure()
# print(projected_data.shape,random_y_values.shape)
plt.scatter(projected_data, random_y_values, c=sub_group_labels, cmap=cmap2)
plt.title('Quadratic Discriminant Analysis: Mice Data')
mice.append(np.array([projected_data,random_y_values]).T)
projected_data_ = mdl.fit(data_, labels_).predict_log_proba(target_)[:,0]
random_y_values_ = np.random.random(size=projected_data_.shape)
plt.figure()
plt.scatter(projected_data_, random_y_values_, c=sub_group_labels_, cmap=cmap4)
plt.title('Quadratic Discriminant Analysis: Synthetic Data')
synthetic.append(np.array([projected_data_,random_y_values_]).T)
```
* QDA is unable to resolve the subgroups of interest in the mice data
* QDA is unable to resolve the subgroups in the synthetic data
### Linear Regression + PCA
```
from sklearn.linear_model import LinearRegression
from sklearn.decomposition import PCA
lr = LinearRegression()
lr.fit(data, labels)
idx = np.where(np.abs(lr.coef_)>0.0001)[0] # get significant directions
target_reduced = target[:,idx]
mdl = PCA(n_components=2)
projected_data = mdl.fit_transform(target_reduced)
plt.figure()
plt.scatter(*projected_data.T, c=sub_group_labels, cmap=cmap2)
plt.title('Limma: Mice Data')
mice.append(projected_data)
lr = LinearRegression()
lr.fit(data_, labels_)
idx = np.where(np.abs(lr.coef_)>0.0001)[0] # get significant directions
target_reduced_ = target_[:,idx]
mdl = PCA(n_components=2)
projected_data_ = mdl.fit_transform(target_reduced_)
plt.figure()
plt.scatter(*projected_data_.T, c=sub_group_labels_, cmap=cmap4)
plt.title('Limma: Synthetic Data')
synthetic.append(projected_data_)
```
### Multidimensional Scaling (MDS)
```
from sklearn.manifold import MDS
np.random.seed(0) # for reproducibility
mdl = MDS(n_components=2)
projected_data = mdl.fit_transform(target)
plt.figure()
plt.scatter(*projected_data.T, c=sub_group_labels, cmap=cmap2)
plt.title('Multidimensional Scaling: Mice Data')
projected_data_ = mdl.fit_transform(target_)
plt.figure()
plt.scatter(*projected_data_.T, c=sub_group_labels_, cmap=cmap4)
plt.title('Multidimensional Scaling: Synthetic Data')
mice.append(projected_data)
synthetic.append(projected_data_)
```
* MDS is mostly able to resolve the subgroups of interest in the mice data, although not as well as cPCA (if the same plot had been plotted without the subgroups colored differently, it would be harder to resolve the subgroups)
* MDS is unable to resolve the subgroups in the synthetic data
### Principal Component Pursuit
```
from pursuit import R_pca
from sklearn.decomposition import PCA
mdl = PCA(n_components=2) # this will be used to select the top 2 principal pursuit components
rpca = R_pca(target)
L, S = rpca.fit(max_iter=10000) #L is the low-rank structure we are interested in
projected_data = mdl.fit_transform(L)
plt.figure()
plt.scatter(*projected_data.T, c=sub_group_labels, cmap=cmap2)
rpca_ = R_pca(target_)
L_, S_ = rpca_.fit(max_iter=10000) #L is the low-rank structure we are interested in
projected_data_ = mdl.fit_transform(L_)
plt.figure()
plt.scatter(*projected_data_.T, c=sub_group_labels_, cmap=cmap4)
mice.append(projected_data)
synthetic.append(projected_data_)
```
* PCP is unable to resolve the subgroups of interest in the mice data (again, consider if the subgroups had not been color-coded), although it does a better job than PCA
* PCP is unable to resolve the subgroups in the synthetic data perfectly
### Factor Analysis
```
from sklearn.decomposition import FactorAnalysis as FA
mdl = FA(n_components=2)
projected_data = mdl.fit_transform(target)
plt.figure()
plt.scatter(*projected_data.T, c=sub_group_labels, cmap=cmap2)
plt.title('Factor Analysis: Mice Data')
projected_data_ = mdl.fit_transform(target_)
plt.figure()
plt.scatter(*projected_data_.T, c=sub_group_labels_, cmap=cmap4)
plt.title('Factor Analysis: Synthetic Data')
mice.append(projected_data)
synthetic.append(projected_data_)
```
* FA is unable to resolve the subgroups of interest in the mice data (again, consider if the subgroups had not been color-coded), although it does a better job than PCA
* FA is unable to resolve the subgroups in the synthetic data perfectly
### Independent Component Analysis
```
from sklearn.decomposition import FastICA as ICA
mdl = ICA(n_components=2)
projected_data = mdl.fit_transform(target)
plt.figure()
plt.scatter(*projected_data.T, c=sub_group_labels, cmap=cmap2)
plt.title('ICA: Mice Data')
projected_data_ = mdl.fit_transform(target_)
plt.figure()
plt.scatter(*projected_data_.T, c=sub_group_labels_, cmap=cmap4)
plt.title('ICA: Synthetic Data')
mice.append(projected_data)
synthetic.append(projected_data_)
```
* ICA is unable to resolve the subgroups of interest in the mice data
* ICA is unable to resolve the subgroups in the synthetic datam
# Comprehensive Plots
```
method_names = ['cPCA','PCA','Supervised PCA','LDA','QDA','LR+PCA','MDS','PC Pursuit','FA','ICA']
plt.figure(figsize=[15, 20])
k = len(mice)
for i in range(k):
#ax = plt.subplot((k+1)/2,2,i+1)
ax = plt.subplot(4,3,i+1+2*(i>0)+(i==0))
plt.scatter(*mice[i].T, c=sub_group_labels, cmap=cmap2, alpha=0.6)
plt.title(method_names[i])
plt.tight_layout()
plt.savefig('mice10.jpg')
method_names = ['cPCA','PCA','Supervised PCA','LDA','QDA','LR+PCA','MDS','PC Pursuit','FA','ICA']
plt.figure(figsize=[15, 20])
k = len(mice)
for i in range(k):
#ax = plt.subplot((k+1)/2,2,i+1)
ax = plt.subplot(4,3,i+1+2*(i>0)+(i==0))
plt.scatter(*synthetic[i].T, c=sub_group_labels_, cmap=cmap4, alpha=0.6)
plt.title(method_names[i])
plt.tight_layout()
plt.savefig('synthetic10.jpg')
```
| github_jupyter |
```
#Import Library :
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.preprocessing import image
from tensorflow.keras.optimizers import RMSprop
from keras.optimizers import adam_v2
from keras import models
from keras import layers
import tensorflow as tf
import matplotlib.pyplot as plt
import cv2
import numpy as np
import os
#Menghubungkan Google Colab dengan Google Drive :
from google.colab import drive
drive.mount('/content/drive')
#Path data train dan data testingnya :
validation_path ='/content/drive/MyDrive/Colab Notebooks/Machine Learning Proyek/validation'
training_path ='/content/drive/MyDrive/Colab Notebooks/Machine Learning Proyek/training'
#melihat salah satu gambar dari data validation
img = image.load_img('/content/drive/MyDrive/Colab Notebooks/Machine Learning Proyek/validation/Apple___Apple_scab/00075aa8-d81a-4184-8541-b692b78d398a___FREC_Scab 3335_new30degFlipLR.JPG')
plt.imshow(img)
#melihat ukuran pixel dari salah satu gambar :
cv2.imread('/content/drive/MyDrive/Colab Notebooks/Machine Learning Proyek/validation/Apple___Apple_scab/00075aa8-d81a-4184-8541-b692b78d398a___FREC_Scab 3335_new30degFlipLR.JPG').shape
#Normalisasi Data :
training = ImageDataGenerator(rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
validation_split=0.2)
validation = ImageDataGenerator(rescale=1./255)
training_dataset = training.flow_from_directory(training_path,
target_size=(200,200),
batch_size=32,
class_mode='sparse')
validation_dataset = validation.flow_from_directory(validation_path,
target_size=(200,200),
batch_size=32,
class_mode='sparse')
#Cek Klasifikasi pada tiap tiap label
training_dataset.class_indices
#Melihat Klasifikasi pada tiap gambar
training_dataset.classes
#CNN MODEL
model = tf.keras.models.Sequential([
#Feature Extraction Layer
tf.keras.layers.Conv2D(32,(3,3),activation = 'relu', input_shape=(200,200,3)),
tf.keras.layers.MaxPool2D(2,2),
tf.keras.layers.Conv2D(32,(3,3),activation = 'relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(64,(3,3),activation = 'relu'),
tf.keras.layers.MaxPool2D(2,2),
# Flatten feature map
tf.keras.layers.Flatten(),
# Fully Connected Layer
tf.keras.layers.Dense(64,activation='relu'),
tf.keras.layers.Dense(4,activation='softmax')
])
#print model summary
model.summary()
#Compile dengan menggunakan optimzer adam dan sparse_crossentropy untuk menghitung lossnya
adam = adam_v2.Adam(learning_rate=0.001)
model.compile(adam, loss='sparse_categorical_crossentropy', metrics=['accuracy'])
history = model.fit(training_dataset,
steps_per_epoch=10,
epochs=120,
validation_data = validation_dataset)
#save weightnya
filepath="/content/drive/MyDrive/Colab Notebooks/Machine Learning Proyek/model_last.hdf5"
model.save(filepath)
#model.load_weights('/content/drive/MyDrive/Colab Notebooks/Machine Learning Proyek/machineLearning_model_save.hdf5')
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
#Train and validation accuracy
plt.plot(epochs, acc, 'b', label='Training accurarcy')
plt.plot(epochs, val_acc, 'r', label='Validation accurarcy')
plt.title('Training and Validation accurarcy')
plt.legend()
plt.figure()
#Train and validation loss
plt.plot(epochs, loss, 'b', label='Training loss')
plt.plot(epochs, val_loss, 'r', label='Validation loss')
plt.title('Training and Validation loss')
plt.legend()
plt.show()
print("[INFO] Calculating model accuracy")
scores = model.evaluate(validation_dataset)
print(f"Test Accuracy: {scores[1]*100}")
#Memasukkan Jenis jenis data ke dalam variable li
class_dict = training_dataset.class_indices
li = list(class_dict.keys())
# predicting an image
image_path = '/content/drive/MyDrive/Colab Notebooks/Machine Learning Proyek/training/Apple___Apple_scab/00075aa8-d81a-4184-8541-b692b78d398a___FREC_Scab 3335.JPG'
new_img = image.load_img(image_path, target_size=(200, 200))
img = image.img_to_array(new_img)
img = np.expand_dims(img, axis=0)
img = img/255 #normalisasi
print("Hasil Prediksi :")
prediction = model.predict(img)
# decode the results into a list of tuples (class, description, probability)
# (one such list for each sample in the batch)
d = prediction.flatten()
j = d.max()
for index,item in enumerate(d):
if item == j:
class_name = li[index]
##Another way
# img_class = classifier.predict_classes(img)
# img_prob = classifier.predict_proba(img)
# print(img_class ,img_prob )
#ploting image with predicted class name
plt.figure(figsize = (4,4))
plt.imshow(new_img)
#plt.axis('off')
plt.title(class_name)
plt.show()
#prediksi dalam beberapa gambar
image_path = '/content/drive/MyDrive/Colab Notebooks/Machine Learning Proyek/testing'
for i in os.listdir(image_path):
prediction = model.predict(img)
new_img = image.load_img(image_path+'//'+i, target_size=(200, 200))
img = image.img_to_array(new_img)
img = np.expand_dims(img, axis=0)
img = img/255
d = prediction.flatten()
j = d.max()
for index,item in enumerate(d):
if item == j:
class_name = li[index]
plt.figure(figsize = (4,4))
plt.imshow(new_img)
#plt.axis('off')
plt.title(class_name)
plt.show()
```
| github_jupyter |
# 新闻数据采集器(国内板块)
## 吴泓骏 [拂晓工作室](https://github.com/Errrneist/Alchemist)
* 此程序将对指定股票数据进行收集并整理
* 同时创建SFrame友好的CSV文件以及对数据进行清洗
# 参考资料
* [1] [Basics of SFrame](https://apple.github.io/turicreate/docs/api/generated/turicreate.SFrame.html#turicreate.SFrame)
* [2] [Remove Multiple Substring from String](https://stackoverflow.com/questions/31273642/better-way-to-remove-multiple-words-from-a-string)
* [3] [楼老师的Python分析红楼梦](https://zhuanlan.zhihu.com/p/29209681)
* [4] [如何使用pyltp包进行中文分词](https://blog.csdn.net/sinat_26917383/article/details/77067515)
# 导入库
```
# 导入库
import urllib
import re
import os
import csv
import time
import datetime
import turicreate as tc
from bs4 import BeautifulSoup
```
# 准备阶段
* 1. 定义用于搜寻的板块
* 2. 创建一个包含25个链接的搜寻目录
* 3. 创建一个包含20个Class的搜寻目录
```
# 新闻板块(同时也会加入热门新闻的数据)
china = 'cgnjj' # 国内
international = 'cgjjj' # 国际
# 定义基本常量
url = 'http://finance.eastmoney.com/news/' + china + '.html' # 主目录板块
# 创建一个二十五页的list
page_list = []
counter = 1
while counter <= 25:
pageurl = 'http://finance.eastmoney.com/news/' + china
if counter != 1:
pageurl = pageurl + '_' + str(counter) + '.html'
page_list.append(pageurl)
else:
pageurl = pageurl + '.html'
page_list.append(pageurl)
counter += 1
print('成功创建包含 ' + str(len(page_list)) + ' 个页面链接的目录!')
# 创建一个包含20个class的list
counter = 0
class_list = []
while counter < 20:
class_list.append('newsTr' + str(counter))
counter += 1
print('成功创建包含 ' + str(len(class_list)) + ' 个Class的目录!')
# print(class_list) # Debug
```
# 获得所有文章的链接
```
# 初始化urllist
urllist = []
year = '2018' # Separate parameter
counter = 1
# 创造urllist
print('-----------------获取所有文章超链接程序-----------------')
print('分析任务开始!')
for url in page_list:
req = urllib.request.Request(url)
response = urllib.request.urlopen(req)
html = response.read()
soup = BeautifulSoup(html, "lxml")
for each_url in soup.find_all('a', href=True):
if 'http://finance.eastmoney.com/news/' in each_url['href']:
if year in each_url['href']:
urllist.append(each_url['href'])
print('第' + str(counter) + '页分析完成!现在一共获得了' + str(len(urllist)) + '篇文章的链接!')
counter += 1
print('全部分析完成!正在查重...')
urllist = list(set(urllist))
print('任务完成!共获得了' + str(len(urllist)) + '篇文章的链接!')
print('---------------------------------------------')
```
# 将各个网页的资料抓取进SFrame
* 1. 新建用于抓取的函数
* 2. 实施抓取
* 3. 清洗数据
# 创建洗词库
```
# 清洗数据的词库
banned_info = ['责任编辑','原标题']
banned_words = ['摘要\n', '\n', '\r','\u3000','(中国新闻网)',
'来源','以下简称', '(新华社)', '>>>',
'商务微新闻带你一图了解!','-', '经济日报', '中国经济网',
'附件:', '>>>>', '>>', '>>>>>', '(新华网)',
'(第一财经)', '据新华社报道,', '▼', '▲', '【','】',' ']
```
# 创建抓取与清洗函数
```
# 抓取新闻URL函数
def collectNews(news, url, counter):
# 准备请求数据
req = urllib.request.Request(url)
print('请求链接连接成功!')
response = urllib.request.urlopen(req)
print('收到反馈信号!')
html = response.read()
# print('HTML生成完毕!')
soup = BeautifulSoup(html, "lxml")
# print('SOUP创建完成!')
# 获得文章的发表时间
time = soup.find(class_="time").get_text()
# print('时间获取完毕!')
# 获得文章的标题
title = soup.find('h1').get_text()
# 鉴于有的文章将标题重复一遍 如此加入洗词库把标题洗掉
banned_words.append(title)
# 获得文章的内容
content = soup.find(id="ContentBody").get_text()
# 清洗文章末尾编辑原文信息
for banned_information in banned_info:
if banned_information in content:
content = content[0:re.search(banned_information, content).span()[0]-1].strip()
# 清洗文章中冗余文字
for banned_word in banned_words:
content = content.replace(banned_word, '')
# print('文章内容获取完成!')
# 将标题从洗词库中去除
banned_words.remove(title)
# 获取相关主题
related_stocks = []
for each in soup.find_all(class_='keytip'):
related_stocks.append(each.get_text())
# 相关主题查重
related_stocks = list(set(related_stocks))
# print('相关主题单获取完成!')
# 写入SFrame
temp_sframe = tc.SFrame({'year':[str(time[0:4])],
'month':[str(time[5:7])],
'day':[str(time[8:10])],
'date':[str(time[0:4]) + str(time[5:7]) + str(time[8:10])],
'title':[title],
'contents':[content],
'related':[related_stocks]})
news = news.append(temp_sframe)
# print('SFrame写入完成!')
# 释放内存
del(req, response, html, soup, time, content, related_stocks)
# 刷新计数
counter += 1
# print('计数刷新完成!')
print('页面数据获取完毕!')
return news
```
# 执行抓取与数据清洗函数
```
# 初始化计数器
counter = 1
# 获取总任务数
total = len(urllist)
# 初始化SFrame
news = tc.SFrame({'year':['0000'],'month':['00'],'day':['00'],'date':['00000000'],'title':['Null Title'],'contents':['Null Contents'],'related':[['Null', 'Null']]})
# 下载数据
for each_article in urllist:
print('=============================================')
print('正在获取第' + str(counter) + '篇文章,共' + str(total) + '篇.')
print('---------------------------------------------')
news = collectNews(news, each_article, counter)
counter += 1
print('获取完毕!共获取了' + str(len(news['title'])) + '篇文章!')
# 删除占位符
news = news[1:len(news['title'])]
# 若仍然存在 则原地去世
if(news[0]['year'] == '0000'):
print('未移除前:' + news[0]['title'])
news = news[1:len(news['title'])]
print('移除后:' + news[0]['title'])
else:
print('占位符已确认移除!')
# 最后一次查重
news = news.unique()
```
# 盖上时间戳并保存数据
```
# 保存数据
filepath = '../DataSets/Eastmoney/News/China/'
date = '20' + str(datetime.datetime.now().strftime("%y%m%d-%H%M"))
news.save(filepath + 'CHINA' + date + '.csv', format='csv')
print('成功保存数据文件!数据路径:' + filepath + 'CHINA' + '-' + date + '.csv')
# 打印时间戳
print('程序运行时间戳:20'
+ str(datetime.datetime.now().strftime("%y")) + '年'
+ str(datetime.datetime.now().strftime("%m")) + '月'
+ str(datetime.datetime.now().strftime("%d")) + '日'
+ str(datetime.datetime.now().strftime("%H")) + '时'
+ str(datetime.datetime.now().strftime("%M")) + '分'
+ str(datetime.datetime.now().strftime("%S")) + '秒')
```
| github_jupyter |
```
from utils.utils import load_model
from prompts.generic_prompt import load_prefix, generate_response_interactive, select_prompt_interactive
from prompts.generic_prompt_parser import load_prefix as load_prefix_parse
from prompts.persona_chat import convert_sample_to_shot_persona
from prompts.persona_chat_memory import convert_sample_to_shot_msc, convert_sample_to_shot_msc_interact
from prompts.persona_parser import convert_sample_to_shot_msc as convert_sample_to_shot_msc_parse
from prompts.emphatetic_dialogue import convert_sample_to_shot_ed
from prompts.daily_dialogue import convert_sample_to_shot_DD_prefix, convert_sample_to_shot_DD_inference
from prompts.skill_selector import convert_sample_to_shot_selector
import random
import torch
import pprint
pp = pprint.PrettyPrinter(indent=4)
args = type('', (), {})()
args.multigpu = False
device = 4
## To use GPT-Jumbo (178B) set this to true and input your api-key
## Visit https://studio.ai21.com/account for more info
## AI21 provides 10K tokens per day, so you can try only for few turns
api = False
api_key = ''
## This is the config dictionary used to select the template converter
mapper = {
"persona": {"shot_converter":convert_sample_to_shot_persona,
"shot_converter_inference": convert_sample_to_shot_persona,
"file_data":"data/persona/","with_knowledge":None,
"shots":{1024:[0,1,2],2048:[0,1,2,3,4,5]},"max_shot":{1024:2,2048:3},
"shot_separator":"\n\n",
"meta_type":"all","gen_len":50,"max_number_turns":5},
"msc": {"shot_converter":convert_sample_to_shot_msc,
"shot_converter_inference": convert_sample_to_shot_msc_interact,
"file_data":"data/msc/session-2-","with_knowledge":None,
"shots":{1024:[0,1],2048:[0,1,3]},"max_shot":{1024:1,2048:3},
"shot_separator":"\n\n",
"meta_type":"all","gen_len":50,"max_number_turns":3},
"ed": {"shot_converter":convert_sample_to_shot_ed,
"shot_converter_inference": convert_sample_to_shot_ed,
"file_data":"data/ed/","with_knowledge":None,
"shots":{1024:[0,1,7],2048:[0,1,17]},"max_shot":{1024:7,2048:17},
"shot_separator":"\n\n",
"meta_type":"none","gen_len":50,"max_number_turns":5},
"DD": {"shot_converter":convert_sample_to_shot_DD_prefix,
"shot_converter_inference": convert_sample_to_shot_DD_inference,
"file_data":"data/dailydialog/","with_knowledge":False,
"shots":{1024:[0,1,2],2048:[0,1,6]},"max_shot":{1024:2,2048:6},
"shot_separator":"\n\n",
"meta_type":"all_turns","gen_len":50,"max_number_turns":5},
"msc-parse": {"shot_converter":convert_sample_to_shot_msc_parse, "max_shot":{1024:1,2048:2},
"file_data":"data/msc/parse-session-1-","level":"dialogue", "retriever":"none",
"shots":{1024:[0,1],2048:[0, 1, 2]},"shot_separator":"\n\n",
"meta_type":"incremental","gen_len":50,"max_number_turns":3},
}
## This is the config dictionary used to select the template converter
mapper_safety = {
"safety_topic": {"file_data":"data/safety_layers/safety_topic.json","with_knowledge":None,
"shots":{1024:[0,1,2],2048:[0,1,2,3,4,5]},"max_shot":{1024:2,2048:3},
"shot_separator":"\n\n",
"meta_type":"all","gen_len":50,"max_number_turns":2},
"safety_nonadv": {"file_data":"data/safety_layers/safety_nonadv.json","with_knowledge":None,
"shots":{1024:[0,1,2],2048:[0,1,2,3,4,5]},"max_shot":{1024:2,2048:3},
"shot_separator":"\n\n",
"meta_type":"all","gen_len":50,"max_number_turns":2},
"safety_adv": {"file_data":"data/safety_layers/safety_adv.json","with_knowledge":None,
"shots":{1024:[0,1,2],2048:[0,1,2,3,4,5]},"max_shot":{1024:2,2048:3},
"shot_separator":"\n\n",
"meta_type":"all","gen_len":50,"max_number_turns":2},
}
## Load LM and tokenizer
## You can try different LMs:
## gpt2
## gpt2-medium
## gpt2-large
## gpt2-xl
## EleutherAI/gpt-neo-1.3B
## EleutherAI/gpt-neo-2.7B
## EleutherAI/gpt-j-6B
## So far the largest I could load is gpt2-large
model_checkpoint = "EleutherAI/gpt-neo-1.3B"
model, tokenizer, max_seq = load_model(args,model_checkpoint,device)
available_datasets = mapper.keys()
prompt_dict = {}
prompt_parse = {}
prompt_skill_selector = {}
for d in available_datasets:
if "parse" in d:
prompt_parse[d] = load_prefix_parse(tokenizer=tokenizer, shots_value=mapper[d]["shots"][max_seq],
shot_converter=mapper[d]["shot_converter"],
file_shot=mapper[d]["file_data"]+"valid.json",
name_dataset=d, level=mapper[d]["level"],
shot_separator=mapper[d]["shot_separator"],sample_times=1)[0]
else:
prompt_skill_selector[d] = load_prefix(tokenizer=tokenizer, shots_value=[6],
shot_converter=convert_sample_to_shot_selector,
file_shot= mapper[d]["file_data"]+"train.json" if "smd" in d else mapper[d]["file_data"]+"valid.json",
name_dataset=d, with_knowledge=None,
shot_separator=mapper[d]["shot_separator"],sample_times=1)[0]
prompt_dict[d] = load_prefix(tokenizer=tokenizer, shots_value=mapper[d]["shots"][max_seq],
shot_converter=mapper[d]["shot_converter"],
file_shot=mapper[d]["file_data"]+"valid.json",
name_dataset=d, with_knowledge=mapper[d]["with_knowledge"],
shot_separator=mapper[d]["shot_separator"],sample_times=1)[0]
## add safety prompts
for d in mapper_safety.keys():
prompt_skill_selector[d] = load_prefix(tokenizer=tokenizer, shots_value=[6],
shot_converter=convert_sample_to_shot_selector,
file_shot= mapper_safety[d]["file_data"],
name_dataset=d, with_knowledge=None,
shot_separator=mapper_safety[d]["shot_separator"],sample_times=1)[0]
def run_parsers(args, model, tokenizer, device, max_seq, dialogue, skill, prefix_dict):
dialogue["user_memory"].append([])
if skill not in ["msc"]: return dialogue
# if d == "dialKG":
# dialogue["KG"].append([])
### parse
d_p = f"{skill}-parse"
# print(f"Parse with {d_p}")
prefix = prefix_dict[d_p].get(mapper[d_p]["max_shot"][max_seq])
query = generate_response_interactive(model, tokenizer, shot_converter=mapper[d_p]["shot_converter"],
dialogue=dialogue, prefix=prefix,
device=device, with_knowledge=None,
meta_type=None, gen_len=50,
beam=1, max_seq=max_seq, eos_token_id=198,
do_sample=False, multigpu=False, api=api, api_key=api_key)
# print(f"Query: {query}")
# if d == "wow":
# dialogue["KB_wiki"].append([retrieve_K])
# elif d == "dialKG":
# dialogue["KG"][-1] = [retrieve_K]
# elif d == "wit":
# dialogue["KB_internet"].append([retrieve_K])
# dialogue["query"].append([query])
if skill == "msc":
if "none" != query:
dialogue["user"].append(query)
dialogue["user_memory"][-1] = [query]
return dialogue
max_number_turns = 3
dialogue = {"dialogue":[],"meta":[],"user":[],"assistant":[],"user_memory":[]}
## This meta information is the persona of the FSB
dialogue["meta"] = dialogue["assistant"] = [
"i am the smartest chat-bot around .",
"my name is FSB . ",
"i love chatting with people .",
"my creator is Andrea"
]
t = 10
while t>0:
t -= 1
user_utt = input(">>> ")
dialogue["dialogue"].append([user_utt,""])
## run the skill selector
skill = select_prompt_interactive(model, tokenizer,
shot_converter=convert_sample_to_shot_selector,
dialogue=dialogue, prompt_dict=prompt_skill_selector,
device=device, max_seq=max_seq, max_shot=6)
if "safety" in skill:
response = "Shall we talk about something else?"
print(f"FSB (Safety) >>> {response}")
else:
## parse user dialogue history ==> msc-parse
dialogue = run_parsers(args, model, tokenizer, device=device, max_seq=max_seq,
dialogue=dialogue, skill=skill,
prefix_dict=prompt_parse)
## generate response based on skills
prefix = prompt_dict[skill].get(mapper[skill]["max_shot"][max_seq])
response = generate_response_interactive(model, tokenizer, shot_converter=mapper[skill]["shot_converter_inference"],
dialogue=dialogue, prefix=prefix,
device=device, with_knowledge=mapper[skill]["with_knowledge"],
meta_type=mapper[skill]["meta_type"], gen_len=50,
beam=1, max_seq=max_seq, eos_token_id=198,
do_sample=True, multigpu=False, api=api, api_key=api_key)
print(f"FSB ({skill}) >>> {response}")
dialogue["dialogue"][-1][1] = response
dialogue["dialogue"] = dialogue["dialogue"][-max_number_turns:]
dialogue["user_memory"] = dialogue["user_memory"][-max_number_turns:]
print("This is the conversation history with its meta-data!")
print(pp.pprint(dialogue))
```
| github_jupyter |
```
# Import the numpy package under the name "np"
import numpy as np
# Print the numpy version and the configuration
print(np.__version__)
print(np.show_config())
# Create a null vector of size 10
np.zeros(10)
# How to find the memory size of any array
arr = np.arange(10)
arr.size * arr.itemsize # Number of elements * byte size of element
# How to get the documentation of the numpy add function from the command line
np.info("add")
# Create a null vector of size 10 but the fifth value which is 1
arr = np.zeros(10)
arr[4] = 1
arr
# Create a vector with values ranging from 10 to 49
np.arange(10, 50)
# Reverse a vector (first element becomes last)
np.arange(10)[::-1]
# Create a 3x3 matrix with values ranging from 0 to 8
np.arange(9).reshape(3,3)
# Find indices of non-zero elements from [1, 2, 0, 0, 4, 0]
np.nonzero([1, 2, 0, 0, 4, 0])
# Create a 3x3 identity matrix
np.eye(3)
# Create a 3x3x3 array with random values
np.random.rand(3,3,3)
# Create a 10x10 array with random values and find the minimum and maximum values
arr = np.random.rand(10,10)
print(arr)
print("Min: {}".format(np.min(arr)))
print("Max: {}".format(np.max(arr)))
# Create a random vector of size 30 and find the mean value
arr = np.random.rand(30)
print(arr)
print("Mean: {}".format(np.mean(arr)))
# Create a 2d array with 1 on the border and 0 inside
n = 10
arr1 = np.ones(n)
arr2 = np.ones(n)
arr3 = np.ones(n)
arr2[1:-1] = 0
arr = np.array([arr1, arr2, arr3])
arr
# How to add a border (filled with 0s) around an existing array?
existing_arr = np.arange(1,10).reshape(3,3)
np.pad(existing_arr, 1, "constant")
# What is the result of the following expression?
0 * np.nan # nan
np.nan == np.nan # False
np.inf > np.nan # False
np.nan - np.nan # nan
0.3 == 3 * 0.1 # False
# Create a 5x5 matrix with values 1,2,3,4 just below the diagonal
arr = np.zeros((5,5))
arr[1, 0] = 1
arr[2, 1] = 2
arr[3, 2] = 3
arr[4, 3] = 4
arr
# Create a 8x8 matrix and fill it with a checkerboard pattern
arr = np.zeros((8,8))
arr[1::2, ::2] = 1
arr[::2, 1::2] = 1
arr
# Consider a (6, 7, 8) shape array, whatis the index (x, y, z) of the 100th element?
np.unravel_index(100, (6,7,8))
# Create a checkerboard 8x8 matrix using the tile function
arr = np.array([[0, 1], [1, 0]])
np.tile(arr, (4,4))
# Normalize a 5x5 random matrix
arr = np.random.rand(5,5)
print("Original 5x5 random matrix: \n{}\n".format(arr))
mx = np.max(arr)
mn = np.min(arr)
for i in range(arr[0].size):
for j in range(arr[i].size):
arr[i, j] = (arr[i, j] - mn) / (mx - mn)
print("Normalized 5x5 matrix: \n{}".format(arr))
# Create a custom dtype that describes a color as four unsigned bytes (RGBA)
np.dtype([('r', np.uint), ('g', np.uint), ('b', np.uint), ('a', np.uint)])
# Multiply a 5x3 matrix by a 3x2 matrix (real matrix product)
a = np.random.rand(5,3)
b = np.random.rand(3,2)
print("{}\n".format(a))
print("{}\n".format(b))
print("Matrix product of the above two matrices:")
print(np.dot(a, b)) # Or a @ b
# Given a 1D array, negate all elements which are between 3 and 8, in place
arr = np.arange(30)
np.random.shuffle(arr)
for i in range(len(arr)):
if 3 <= arr[i] <= 8:
arr[i] = -arr[i]
arr
# What is the output of the following script?
print(sum(range(5),-1)) # 10
from numpy import *
print(sum(range(5),-1)) # 10
# Consider an integer vector Z, which of these expressions are legal?
# All of the below expressions are legal!
Z = 5
Z**Z # 3125
2 << Z >> 2 # 16
Z <- Z # False
1j*Z # 5j
Z/1/1 # 5.0
Z<Z>Z # False
# What is the result of the following expression?
np.array(0) / np.array(0) # nan (RuntimeWarning: invalid value encountered in true_divide)
np.array(0) // np.array(0) # 0 (RuntimeWarning: divide by zero encountered in floor_divide)
np.array([np.nan]).astype(int).astype(float) # array([-9.22337204e+18])
# How to round away from zero a float array?
arr = np.random.uniform(-5, 5, 10)
arr2 = np.round(arr + np.copysign(0.5, arr))
print(arr)
print(arr2)
# How to find common values between two arrays?
arr = np.random.randint(-5, 5, 10)
arr2 = np.random.randint(-5, 5, 10)
arr3 = np.intersect1d(arr, arr2)
print(arr)
print(arr2)
print(arr3)
# How to ignore all numpy warnings (not recommended)?
# Default: np.seterr(all="warn")
np.seterr(all="ignore")
# Is the following expression true?
# False
np.sqrt(-1) == np.emath.sqrt(-1)
# How to get the dates of yesterday, today, and tomorrow?
today = np.datetime64("today")
yesterday = today - np.timedelta64(1)
tomorrow = today + np.timedelta64(1)
print("Today: {}".format(today))
print("Yesterday: {}".format(yesterday))
print("Tomorrow: {}".format(tomorrow))
# How to get all the dates corresponding to the month of July 2016?
np.arange("2016-07", "2016-08", dtype="datetime64[D]")
# How to compute ((A+B)\*(-A/2)) in place (without copy)?
A = np.random.uniform(-5, 5, 5)
B = np.random.uniform(-5, 5, 5)
print(A)
print(B)
np.add(A, B, out=B)
np.negative(A, out=A)
np.multiply(A, 0.5, out=A) # np.divide(A, 2.0, out=A) causes a TypeError
np.multiply(A, B, out=A)
print(A)
# Extract the integer part of a random array using 5 different methods
arr = np.random.uniform(-15, 15, 10)
print(arr)
print(arr - arr % 1) # Only handles positives correctly, negatives round away
print(np.floor(arr))
print(np.ceil(arr) - 1)
print(arr.astype(int).astype(float)) # Handles positives and negatives correctly
print(np.trunc(arr))
# Create a 5x5 matrix with row values ranging from 0 to 4
arr = np.zeros((5,5))
arr += np.arange(5)
print(arr)
# Consider a generator function that generates 10 integers and use it to build an array
n = 10
iterable = (n * np.random.rand() for x in range(n))
np.fromiter(iterable, float)
# Create a vector of size 10 with values ranging from 0 to 1, both excluded
arr = np.random.uniform(0, 1, 10) # Random values in (0, 1)
arr = np.linspace(0, 1, 11, endpoint=False)[1:] # Equally distributed values in (0, 1)
arr
# Create a random vector of size 10 and sort it
arr = np.random.randint(-15, 15, 10)
np.sort(arr)
# How to sum a small array faster than np.sum?
arr = np.random.randint(-10, 10, 5)
arr_sum = np.add.reduce(arr)
print(arr)
print(arr_sum)
# Consider two random arrays A and B, check if they are equal
A = np.random.randint(0, 2, 4)
B = np.random.randint(0, 2, 4)
result1 = np.allclose(A, B) # Assumes identical shape of arrays, and a tolerance for comparison of values
result2 = np.array_equal(A, B) # Checks for both identical array shape and values, no tolerance for comparison
print(A)
print(B)
print(result1)
print(result2)
# Make an array immutable (read-only)
arr = np.random.randint(-5, 5, 10)
arr.flags.writeable = False
arr
# Any future changes to arr will cause a ValueError
# Consider a random 10x2 matrix representing cartesian coordinates, convert them to polar coordinates
arr = np.random.randint(0, 16, 20).reshape(10,2)
x = arr[:,0]
y = arr[:,1]
r = np.sqrt(x ** 2 + y ** 2) # r = sqrt(x^2 + y^2)
theta = np.arctan2(y, x) * 180 / np.pi # theta = [180 * arctan(y/x)] / pi
print("[X Y] (Cartesian coordinates)")
print(arr)
print("\n[r theta] (Polar coordinates)")
print(np.column_stack((r, theta)))
# Create random vector of size 10 and replace the maximum value by 0
arr = np.random.uniform(-10, 10, 10)
print(arr)
print("Max: {} at index {}".format(max(arr), np.argmax(arr)))
arr[np.argmax(arr)] = 0
print(arr)
# Create a structured array with x and y coordinates covering the [0,1] x [1,0] area
arr = np.zeros((5,5), [("x", float), ("y", float)])
arr["x"], arr["y"] = np.meshgrid(np.linspace(0, 1, 5), np.linspace(0, 1, 5))
print(arr)
# Given two arrays, X and Y, construct the Cauchy matrix C (Cij = 1/(xi - yj))
X = np.random.uniform(-5, 5, 5)
Y = np.random.uniform(-5, 5, 5)
C = 1.0 / np.subtract.outer(X, Y)
C
# Print the minimum and maximum representable value for each numpy scalar type
for dtype in [np.uint8, np.int8, np.uint16, np.int16, np.uint32, np.int32, np.uint64, np.int64]:
print(dtype)
print("Min: {}".format(np.iinfo(dtype).min))
print("Max: {}".format(np.iinfo(dtype).max))
print()
for dtype in [np.float16, np.float32, np.float64, np.float128]:
print(dtype)
print("Min: {}".format(np.finfo(dtype).min))
print("Max: {}".format(np.finfo(dtype).max))
print("Eps: {}".format(np.finfo(dtype).eps))
print()
# How to print all the values of an array?
# Sets the total number of array elements which trigger summarization rather than a full representation to NaN.
np.set_printoptions(threshold=np.nan)
arr = np.arange(50)
print(arr)
# How to find the closest value (to a given scalar) in a vector?
arr = np.arange(100) # Array with elements [0, 100)
scalar = np.random.uniform(0, 100) # A random scalar from [0, 100)
print(arr)
print(scalar)
print(arr[(np.abs(arr - scalar)).argmin()])
# Create a structured array representing a position (x,y) and a color (r,g,b)
dtype = [
("position", [("x", float, 1),
("y", float, 1)]),
("color", [("r", float, 1),
("g", float, 1),
("b", float, 1)])
]
np.zeros(1, dtype)
# Consider a random vector with shape (100,2) representing coordinates, find point by point distances
arr = np.random.randint(-5, 5, (100,2))
x,y = np.atleast_2d(arr[:,0], arr[:,1])
distance = np.sqrt((x - x.T) ** 2 + (y - y.T) ** 2) # T = transpose()
print(arr)
print(distance)
# How to convert a float (32 bits) array into an integer (32 bits) in place?
arr = np.random.uniform(-10, 10, 10)
print(arr)
print(arr.astype(int, copy=False))
# How to read the following file?
# Creating a "file" using StringIO module that contains the text
from io import StringIO
s = StringIO("""1, 2, 3, 4, 5\n6, , , 7, 8\n , , 9,10,11\n""")
print(s.getvalue())
print(np.genfromtxt(s, delimiter=",", dtype=np.int))
# What is the equivalent of enumerate for numpy arrays?
arr = np.random.randint(-10, 10, 9).reshape(3,3)
print(arr)
print("\nUsing np.ndenumerate:")
for i,v in np.ndenumerate(arr):
print(i, v)
print("\nUsing np.ndindex:")
for i in np.ndindex(arr.shape):
print(i, arr[i])
# Generate a generic 2D Gaussian-like array
x,y = np.meshgrid(np.linspace(-1, 1, 10), np.linspace(-1, 1, 10))
distance = np.sqrt(x ** 2 + y ** 2)
sigma = 1.0
mu = 0.0
gaussian = np.exp(-((distance - mu) ** 2 / (2.0 * sigma ** 2)))
print(gaussian)
# How to randomly place p elements in a 2D array?
n = 10
p = 3
arr = np.zeros((n,n))
np.put(arr, np.random.choice(range(n ** 2), p, replace=False), 1)
print(arr)
# Subtract the mean of each row of a matrix
arr = np.random.randint(-10, 10, (5,10))
new_arr = arr - arr.mean(axis=1, keepdims=True)
print("Initial random array: \n{}".format(arr))
print("\nMean of each row of initial array:")
for row in arr:
print(row.mean())
print("\nNew array with mean of each row subtracted from each element: \n{}".format(new_arr))
# How to sort an array by the nth column?
arr = np.random.randint(-10, 10, (3,3))
print("Initial random array: \n{}\n".format(arr))
print("Sorted array by the nth column: \n{}".format(arr[arr[:,1].argsort()]))
# How to tell if a given 2D array has null columns (columns with all 0s)?
arr = np.random.randint(0, 3, (3,10))
print(arr)
print((~arr.any(axis=0)).any())
# Find the nearest value from a given value in an array
arr = np.random.rand(10)
val = 0.5
nearest = arr.flat[np.abs(arr - val).argmin()]
print(arr)
print(nearest)
# Considering two arrays with shape (1,3) and (3,1), how to compute their sum using an iterator?
arr1 = np.arange(3).reshape(1,3)
arr2 = np.arange(3).reshape(3,1)
iterator = np.nditer([arr1, arr2, None])
for x,y,z in iterator:
z[...] = x + y
print(arr1)
print(arr2)
print(iterator.operands[2])
```
| github_jupyter |
[this doc on github](https://github.com/dotnet/interactive/tree/master/samples/notebooks/csharp/Samples)
# Machine Learning over House Prices with ML.NET
```
#r "nuget:Microsoft.ML,1.4.0"
#r "nuget:Microsoft.ML.AutoML,0.16.0"
#r "nuget:Microsoft.Data.Analysis,0.1.0"
using Microsoft.Data.Analysis;
using XPlot.Plotly;
using Microsoft.AspNetCore.Html;
Formatter<DataFrame>.Register((df, writer) =>
{
var headers = new List<IHtmlContent>();
headers.Add(th(i("index")));
headers.AddRange(df.Columns.Select(c => (IHtmlContent) th(c.Name)));
var rows = new List<List<IHtmlContent>>();
var take = 20;
for (var i = 0; i < Math.Min(take, df.RowCount); i++)
{
var cells = new List<IHtmlContent>();
cells.Add(td(i));
foreach (var obj in df[i])
{
cells.Add(td(obj));
}
rows.Add(cells);
}
var t = table(
thead(
headers),
tbody(
rows.Select(
r => tr(r))));
writer.Write(t);
}, "text/html");
using System.IO;
using System.Net.Http;
string housingPath = "housing.csv";
if (!File.Exists(housingPath))
{
var contents = new HttpClient()
.GetStringAsync("https://raw.githubusercontent.com/ageron/handson-ml2/master/datasets/housing/housing.csv").Result;
File.WriteAllText("housing.csv", contents);
}
var housingData = DataFrame.LoadCsv(housingPath);
housingData
housingData.Description()
Chart.Plot(
new Graph.Histogram()
{
x = housingData["median_house_value"],
nbinsx = 20
}
)
var chart = Chart.Plot(
new Graph.Scattergl()
{
x = housingData["longitude"],
y = housingData["latitude"],
mode = "markers",
marker = new Graph.Marker()
{
color = housingData["median_house_value"],
colorscale = "Jet"
}
}
);
chart.Width = 600;
chart.Height = 600;
display(chart);
static T[] Shuffle<T>(T[] array)
{
Random rand = new Random();
for (int i = 0; i < array.Length; i++)
{
int r = i + rand.Next(array.Length - i);
T temp = array[r];
array[r] = array[i];
array[i] = temp;
}
return array;
}
int[] randomIndices = Shuffle(Enumerable.Range(0, (int)housingData.RowCount).ToArray());
int testSize = (int)(housingData.RowCount * .1);
int[] trainRows = randomIndices[testSize..];
int[] testRows = randomIndices[..testSize];
DataFrame housing_train = housingData[trainRows];
DataFrame housing_test = housingData[testRows];
display(housing_train.RowCount);
display(housing_test.RowCount);
using Microsoft.ML;
using Microsoft.ML.Data;
using Microsoft.ML.AutoML;
#!time
var mlContext = new MLContext();
var experiment = mlContext.Auto().CreateRegressionExperiment(maxExperimentTimeInSeconds: 15);
var result = experiment.Execute(housing_train, labelColumnName:"median_house_value");
var scatters = result.RunDetails.Where(d => d.ValidationMetrics != null).GroupBy(
r => r.TrainerName,
(name, details) => new Graph.Scattergl()
{
name = name,
x = details.Select(r => r.RuntimeInSeconds),
y = details.Select(r => r.ValidationMetrics.MeanAbsoluteError),
mode = "markers",
marker = new Graph.Marker() { size = 12 }
});
var chart = Chart.Plot(scatters);
chart.WithXTitle("Training Time");
chart.WithYTitle("Error");
display(chart);
Console.WriteLine($"Best Trainer:{result.BestRun.TrainerName}");
var testResults = result.BestRun.Model.Transform(housing_test);
var trueValues = testResults.GetColumn<float>("median_house_value");
var predictedValues = testResults.GetColumn<float>("Score");
var predictedVsTrue = new Graph.Scattergl()
{
x = trueValues,
y = predictedValues,
mode = "markers",
};
var maximumValue = Math.Max(trueValues.Max(), predictedValues.Max());
var perfectLine = new Graph.Scattergl()
{
x = new[] {0, maximumValue},
y = new[] {0, maximumValue},
mode = "lines",
};
var chart = Chart.Plot(new[] {predictedVsTrue, perfectLine });
chart.WithXTitle("True Values");
chart.WithYTitle("Predicted Values");
chart.WithLegend(false);
chart.Width = 600;
chart.Height = 600;
display(chart);
#!lsmagic
new [] { 1,2,3 }
new { foo ="123" }
#!fsharp
[1;2;3]
b("hello").ToString()
```
| github_jupyter |
#### Copyright 2017 Google LLC.
```
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# TensorFlow Programming Concepts
**Learning Objectives:**
* Learn the basics of the TensorFlow programming model, focusing on the following concepts:
* tensors
* operations
* graphs
* sessions
* Build a simple TensorFlow program that creates a default graph, and a session that runs the graph
**Note:** Please read through this tutorial carefully. The TensorFlow programming model is probably different from others that you have encountered, and thus may not be as intuitive as you'd expect.
## Overview of Concepts
TensorFlow gets its name from **tensors**, which are arrays of arbitrary dimensionality. Using TensorFlow, you can manipulate tensors with a very high number of dimensions. That said, most of the time you will work with one or more of the following low-dimensional tensors:
* A **scalar** is a 0-d array (a 0th-order tensor). For example, `"Howdy"` or `5`
* A **vector** is a 1-d array (a 1st-order tensor). For example, `[2, 3, 5, 7, 11]` or `[5]`
* A **matrix** is a 2-d array (a 2nd-order tensor). For example, `[[3.1, 8.2, 5.9][4.3, -2.7, 6.5]]`
TensorFlow **operations** create, destroy, and manipulate tensors. Most of the lines of code in a typical TensorFlow program are operations.
A TensorFlow **graph** (also known as a **computational graph** or a **dataflow graph**) is, yes, a graph data structure. A graph's nodes are operations (in TensorFlow, every operation is associated with a graph). Many TensorFlow programs consist of a single graph, but TensorFlow programs may optionally create multiple graphs. A graph's nodes are operations; a graph's edges are tensors. Tensors flow through the graph, manipulated at each node by an operation. The output tensor of one operation often becomes the input tensor to a subsequent operation. TensorFlow implements a **lazy execution model,** meaning that nodes are only computed when needed, based on the needs of associated nodes.
Tensors can be stored in the graph as **constants** or **variables**. As you might guess, constants hold tensors whose values can't change, while variables hold tensors whose values can change. However, what you may not have guessed is that constants and variables are just more operations in the graph. A constant is an operation that always returns the same tensor value. A variable is an operation that will return whichever tensor has been assigned to it.
To define a constant, use the `tf.constant` operator and pass in its value. For example:
```
x = tf.constant(5.2)
```
Similarly, you can create a variable like this:
```
y = tf.Variable([5])
```
Or you can create the variable first and then subsequently assign a value like this (note that you always have to specify a default value):
```
y = tf.Variable([0])
y = y.assign([5])
```
Once you've defined some constants or variables, you can combine them with other operations like `tf.add`. When you evaluate the `tf.add` operation, it will call your `tf.constant` or `tf.Variable` operations to get their values and then return a new tensor with the sum of those values.
Graphs must run within a TensorFlow **session**, which holds the state for the graph(s) it runs:
```
with tf.Session() as sess:
initialization = tf.global_variables_initializer()
print(y.eval())
```
When working with `tf.Variable`s, you must explicitly initialize them by calling `tf.global_variables_initializer` at the start of your session, as shown above.
**Note:** A session can distribute graph execution across multiple machines (assuming the program is run on some distributed computation framework). For more information, see [Distributed TensorFlow](https://www.tensorflow.org/deploy/distributed).
### Summary
TensorFlow programming is essentially a two-step process:
1. Assemble constants, variables, and operations into a graph.
2. Evaluate those constants, variables and operations within a session.
## Creating a Simple TensorFlow Program
Let's look at how to code a simple TensorFlow program that adds two constants.
### Provide import statements
As with nearly all Python programs, you'll begin by specifying some `import` statements.
The set of `import` statements required to run a TensorFlow program depends, of course, on the features your program will access. At a minimum, you must provide the `import tensorflow` statement in all TensorFlow programs:
```
import tensorflow as tf
```
**Don't forget to execute the preceding code block (the `import` statements).**
Other common import statements include the following:
```
import matplotlib.pyplot as plt # Dataset visualization.
import numpy as np # Low-level numerical Python library.
import pandas as pd # Higher-level numerical Python library.
```
TensorFlow provides a **default graph**. However, we recommend explicitly creating your own `Graph` instead to facilitate tracking state (e.g., you may wish to work with a different `Graph` in each cell).
```
from __future__ import print_function
import tensorflow as tf
# Create a graph.
g = tf.Graph()
# Establish the graph as the "default" graph.
with g.as_default():
# Assemble a graph consisting of the following three operations:
# * Two tf.constant operations to create the operands.
# * One tf.add operation to add the two operands.
x = tf.constant(8, name="x_const")
y = tf.constant(5, name="y_const")
my_sum = tf.add(x, y, name="x_y_sum")
# Now create a session.
# The session will run the default graph.
with tf.Session() as sess:
print(my_sum.eval())
```
## Exercise: Introduce a Third Operand
Revise the above code listing to add three integers, instead of two:
1. Define a third scalar integer constant, `z`, and assign it a value of `4`.
2. Add `z` to `my_sum` to yield a new sum.
**Hint:** See the API docs for [tf.add()](https://www.tensorflow.org/api_docs/python/tf/add) for more details on its function signature.
3. Re-run the modified code block. Did the program generate the correct grand total?
### Solution
Click below for the solution.
```
# Create a graph.
g = tf.Graph()
# Establish our graph as the "default" graph.
with g.as_default():
# Assemble a graph consisting of three operations.
# (Creating a tensor is an operation.)
x = tf.constant(8, name="x_const")
y = tf.constant(5, name="y_const")
my_sum = tf.add(x, y, name="x_y_sum")
# Task 1: Define a third scalar integer constant z.
z = tf.constant(4, name="z_const")
# Task 2: Add z to `my_sum` to yield a new sum.
new_sum = tf.add(my_sum, z, name="x_y_z_sum")
# Now create a session.
# The session will run the default graph.
with tf.Session() as sess:
# Task 3: Ensure the program yields the correct grand total.
print(new_sum.eval())
```
## Further Information
To explore basic TensorFlow graphs further, experiment with the following tutorial:
* [Mandelbrot set](https://www.tensorflow.org/tutorials/non-ml/mandelbrot)
| github_jupyter |
## Data Pre-processing and Exploratory Data Analysis
```
##import the libraries
import numpy as np
import pandas as pd
#Load the Data
patient_data=pd.read_csv("Health_Data.csv")
##use the head function to get a glimpse data
patient_data.head()
mydata=pd.read_csv("Health_Data.csv")
X=mydata.iloc[:,1:9]
y=mydata.iloc[:,9]
##New Admission type
A_type=pd.get_dummies(X.iloc[:,1],drop_first=True,prefix='Atype')
##New Gender
New_gender=pd.get_dummies(X.iloc[:,4],drop_first=True,prefix='Gender')
##New Pre Existing Disease Variable
Pre_exdis=pd.get_dummies(X.iloc[:,2],drop_first=True,prefix='PreExistDis')
## Drop the original categorical columns
X.drop(['Admission_type','PreExistingDisease','Gender'],axis=1,inplace=True)
##Concat the new transformed data to X dataframe
X=pd.concat([X,A_type,New_gender,Pre_exdis],axis=1)
##Split The Data into Train and Test
from sklearn.model_selection import train_test_split
xtrain,xtest,ytrain,ytest= train_test_split(X, y, test_size=0.25, random_state=711)
##Initialize StandardScaler
from sklearn.preprocessing import StandardScaler
sc=StandardScaler()
#Transform the training data
xtrain=sc.fit_transform(xtrain)
xtrain=pd.DataFrame(xtrain,columns=xtest.columns)
#Transform the testing data
xtest=sc.transform(xtest)
xtest=pd.DataFrame(xtest,columns=xtrain.columns)
#Convert DataFrame to Numpy array
x_train=xtrain.values
x_test=xtest.values
y_train=ytrain.values
y_test=ytest.values
```
## Train a Neural Network and find its accuracy
```
##Import the relevant Keras libraries
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
##Initiate the Model with Sequential Class
model=Sequential()
## Add the 1st dense layer and Dropout Layer
model.add(Dense(units=6,activation='relu',kernel_initializer='uniform',input_dim=11))
model.add(Dropout(rate=0.3))
##Add the 2nd dense Layer and Dropout Layer
model.add(Dense(units=6,activation='relu',kernel_initializer='uniform'))
model.add(Dropout(rate=0.3))
##Add Output Dense Layer
model.add(Dense(units=1,activation='sigmoid',kernel_initializer='uniform'))
#Compile the Model
model.compile(optimizer='adam',loss='binary_crossentropy',metrics=['accuracy'])
#Fit the Model
model.fit(x_train,y_train,epochs=200,batch_size=20,)
#y_pred_class is the predcition & y_pred_prob is probabilities of the prediction
y_pred_class=model.predict(x_test)
y_pred_prob=model.predict_proba(x_test)
##Set threshold all values above threshold are 1 and #below 0
y_pred_class=y_pred_class>0.5
#Calculate accuracy
from sklearn.metrics import accuracy_score
accuracy_score(y_test,y_pred_class)
```
## Compute the Null Accuracy
```
# Use the value_count function to calculate distinct class values
ytest.value_counts()
##use head function and divide it by lenght of ytest
ytest.value_counts().head(1)/len(ytest)
```
| github_jupyter |
```
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import pandas as pd
import pickle as pkl
from pathlib import Path
import am_sim as ams
from utilities.analyze_inference import best_par_in_df
# load inferred parameter set
inference_path = Path('inference_results/t_final_search_history.csv')
search_df = pd.read_csv(inference_path, index_col=0)
par = best_par_in_df(search_df)
# save folder path
save_path = Path('figures/fig_4')
```
### Panel A - asymptotic evolution
```
from utilities.asymptotic_evolution import evolve_pop_at_constant_C, asymptotic_phi_and_u
C_const = 30 # Ag concentration
T_skip = 10 # pre-evolve the initial distribution for some rounds
T_save = 60 # number of evolution rounds to save to be then plotted in panel A
# simulate evolution with constant Ag concentration
results = evolve_pop_at_constant_C(C_const, par, T_skip, T_save)
# extract results
t, N, avg_eps, distr_y, distr_x= [results[lab] for lab in ['t', 'N', 'avg_eps', 'distr_y', 'distr_x']]
# evaluate asymptotic growth rate and travelling wave speed
phi, u = asymptotic_phi_and_u(C_const, par, T_max=1000)
# plot panel A - setup figure
fig, ax = plt.subplots(2,2, figsize = (8,5), constrained_layout=True)
# setup colormap
cmap = mpl.cm.cool
norm = mpl.colors.Normalize(vmin=t.min(), vmax=t.max())
mapp = mpl.cm.ScalarMappable(norm=norm, cmap=cmap)
mapp.set_array(t)
# plot pop. size evolution
ax[0,0].scatter(t, N, color=cmap(norm(t)), zorder = 3)
# plot pop. size evolution
ax[1,0].scatter(t, avg_eps, color=cmap(norm(t)), zorder = 3)
# select times to be displayed
plt_idsx = range(len(t))[::10]
# for each time
for idx in plt_idsx:
# evaluate population function
pop_f = distr_y[idx] * N[idx]
# plot evolution of population function
ax[0,1].plot(distr_x, pop_f, c=cmap(norm(t[idx])))
# evaluate rescaled population function
shifted_x = distr_x - u * t[idx]
rescaled_pop_f = pop_f / np.exp(phi * t[idx])
# plot evolution of rescaled population function
ax[1,1].plot(shifted_x, rescaled_pop_f, c=cmap(norm(t[idx])))
# set axes limits, scales and ticks
ax[0,0].set_yscale('log')
ax[0,1].set_ylim(bottom=0)
ax[1,1].set_ylim(bottom=0)
ax[0,1].set_xlim(-22.5, -10)
ax[1,1].set_xlim(-22.5, -10)
ax[0,1].set_xticks([-20, -15, -10])
# set labels
ax[0,0].set_xlabel('t (rounds)')
ax[0,0].set_ylabel(r'$N_B$')
ax[1,0].set_xlabel('t (rounds)')
ax[1,0].set_ylabel(r'$\langle \epsilon \rangle$')
ax[0,1].set_xlabel(r'$\epsilon$')
ax[0,1].set_ylabel(r'$\rho_t(\epsilon)$')
ax[1,1].set_xlabel(r'$\epsilon - u t$')
ax[1,1].set_ylabel(r'$\rho_t(\epsilon) / \exp \{\phi t\}$')
# write Ag concentration
ax[0,0].text(0.1, 0.85, f'C = {C_const}', transform=ax[0,0].transAxes)
# plot colorbar
plt.colorbar(mapp, ax=ax, label='t (rounds)', aspect=50.)
plt.savefig(save_path / 'panel_A.pdf')
plt.savefig(save_path / 'panel_A.svg')
plt.show()
```
### Panel B - pahse diagram
```
# concentration range on which to draw the phase diagram
C_range = np.logspace(0,4,20)
# maximum allowed number of iterations for the function that evaluates u and phi
T_max_sim = 10000
# evaluate u and phi for all specified concentrations (NB: if too long results can be loaded by executing the next cell)
phi_list = []
u_list = []
for num_c, C in enumerate(C_range):
print(f'test {num_c + 1}/{len(C_range)} at ag. concentration C = {C:.5}')
phi, u = asymptotic_phi_and_u(C, par, T_max_sim)
phi_list.append(phi)
u_list.append(u)
# save results
with open(save_path / 'phi_u_results.pkl', 'wb') as f:
pkl.dump([phi_list, u_list], f)
f.close()
# load results
with open(save_path / 'phi_u_results.pkl', 'rb') as f:
phi_list, u_list = pkl.load(f)
f.close()
# find the two critical concentrations by interpolation
import scipy.optimize as spo
import scipy.interpolate as spi
log_C = np.log(C_range)
# find C^* with an interpolation of phi
interp_phi = spi.interp1d(log_C, phi_list, bounds_error=True)
log_Cs = spo.brentq(interp_phi, log_C.min(), log_C.max())
# find C^** with an interpolation of u
interp_u = spi.interp1d(log_C, u_list, bounds_error=True)
log_Css = spo.brentq(interp_u, log_C.min(), log_C.max())
# plot panel B - setup figure
fig, ax = plt.subplots(2,1, sharex=True, figsize=(4,4))
for ax_i in ax:
# for both axes draw an horizontal line on zero
ax_i.axhline(0, c='k')
# and draw two vertical axes at the critical concentrations
ax_i.axvline(np.exp(log_Cs), c='C3', ls='--',label=r'$C^*$')
ax_i.axvline(np.exp(log_Css), c='C4', ls='--', label=r'$C^{**}$')
# plot phi and u
ax[0].plot(C_range, phi_list, 'C2.-')
ax[1].plot(C_range, u_list, 'C1.-')
# labels
ax[0].set_ylabel(r'$\phi$')
ax[1].set_ylabel(r'$u$')
ax[1].set_xlabel(r'$C$')
ax[1].legend()
# set logarithmic scale on concentration
plt.xscale('log')
plt.tight_layout()
plt.savefig(save_path / 'panel_B.pdf')
plt.savefig(save_path / 'panel_B.svg')
plt.show()
```
| github_jupyter |
# Seaborn Workshop
Seaborn is a Python data visualization library based on matplotlib.
It provides a high-level interface for drawing attractive and informative statistical graphics.
___
Installing Seaborn (conda installation recommend)
https://seaborn.pydata.org/installing.html
___
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns # seaborn library
```
For this session, you will need the data set named ```heart.csv```, which can be downloaded from our [GitHub repository](https://github.com/IC-Computational-Biology-Society/Pandas_Matplotlib_session.git) dedicated to today's workshop. Make sure you save it in the same directory as this Jupyter notebook.
___
## Getting Started
```
df = pd.read_csv('heart.csv')
display(df.head())
```
**Dataset description**
- ```age```: The patient's age
- ```gender```: 0 = female and 1 = male
- ```cp```: The chest pain experienced (Value 1: typical angina, Value 2: atypical angina, Value 3: non-anginal pain, Value 4: asymptomatic)
- ```trestbps```: The patient's resting blood pressure (mm Hg on admission to the hospital)
- ```chol```: The patient's cholesterol measurement in mg/dl
- ```fbs```: The patient's fasting blood sugar (> 120 mg/dl, 1 = true; 0 = false)
- ```restecg```: Resting electrocardiographic measurement (0 = normal, 1 = having ST-T wave abnormality, 2 = showing probable or definite left ventricular hypertrophy by Estes' criteria)
- ```thalach```: The patient's maximum heart rate achieved
- ```exang```: Exercise induced angina (0 = no, 1 = yes)
- ```oldpeak```: ST depression induced by exercise relative to rest ('ST' relates to positions on the ECG plot. See more here)
- ```slope```: the slope of the peak exercise ST segment (Value 1: upsloping, Value 2: flat, Value 3: downsloping)
- ```ca```: The number of major vessels (0-3)
- ```thal```: A blood disorder called thalassemia (3 = normal; 6 = fixed defect; 7 = reversable defect)
- ```target```: Heart disease (0 = no, 1 = yes)
```
# get the number of patients (number of rows) and columns of the dataset
print ("number of patients :", len(df))
print ("number of columns :", len(df.columns))
# check if any values are missing
```
## Task 1
Plot a histogram using ```sns.histplot``` function of the patients age distribution.
Set the paramete ```kde``` to ```True``` inlcude the kernel density estimate.
Don't forget to include the plot's title.
___
```
### Enter code below
```
Create a new histogram using again the ```sns.histplot``` function but showing in different colour the patients with disease (target = 1) and patients without disease (target = 0)
____
```
### Enter code below
```
## Task 2
Similar to histograms are kernel density estimate (KDE) plots, which can be used for visualising the distribution of observations in a dataset. KDE represents the data using a continuous probability density curve.
___
Use the ```sns.kdeplot``` function to visualise to the distribution of resting blood pressure ```trestbps``` with the patients ```gender``` (0 = female, 1 = male)
Rename the legend to 'female' and 'male'
___
```
### Enter code below
```
Create a new figure that contains two subplots, one showing the distribution of resting blood pressure ```trestbps``` with gender and the other showing the distribution of cholesterol ```chol``` with gender.
Include a title to each of the subplots and rename the legends to 'female' and 'male'.
___
```
### Enter code below
```
## Task 3
Use the ```sns.countplot``` function to visualise the counts of patients with and without the disease based on their gender.
Rename the x ticks labels to 'female' and 'male' and the legend values to 'disease' and 'no disease'.
___
```
### Enter code below
```
## Task 4
Correlation indicates how the features are related to each other or to the target variable.
The correlation may be positive (increase in one value of the feature increases the value of the target variable) or negative (increase in one value of the feature decreases the value of the target variable).
Plot a correlation matrix using the ```sns.heatmap``` showing the correlation of the features to each other and the target value.
**Hint**
The correlation between the variable in the data can be caluated using ``` df.corr()```, which needs to be added as the data parameter of the heatmap function.
```
### Enter code below
```
| github_jupyter |
```
import tensorflow as tf
import numpy as np
import pickle as pkl
import random
from Data_converter import convert_to_one_hot
```
Initialize parameters of the system
--------------------------
```
POOL_SIZE = 5
INPUT_CHANNEL = 220
OUTPUT_CLASSES = 16
KERNEL_1 = 21
FILTERS_1 = 20
HIDDEN_LAYER_1 = 100
```
Loading data
----------
```
f = pkl.load(open('our_data.pkl','rb'))
data = f['data']
```
Make next_batch function
---------------------
```
def batch(data,k):
batch = random.sample(data, k)
inputs =[]
targets = []
for a,b in batch:
inputs.append(a)
targets.append(b)
# targets = convert_to_one_hot(targets,OUTPUT_CLASSES)
return inputs, targets
x = tf.placeholder("float",name='x',shape=([None,INPUT_CHANNEL,1,1]))
y = tf.placeholder("float",name='y',shape=([None,OUTPUT_CLASSES]))
```
Defining layers for our network
-----------------------
```
# """ define function """
# def weight_variable(shape):
# init = tf.truncated_normal(shape,stddev=1.0)
# return tf.Variable(init)
# def bias_variable(shape):
# init = tf.constant(0.1,shape=shape)
# return tf.Variable(init)
# def conv_2d(x,W):
# return tf.nn.conv2d(x,W,strides = [1,1,1,1], padding='VALID')
# def max_pool_5x1(x):
# return tf.nn.max_pool(x,ksize=[1,POOL_SIZE,1,1],strides=[1,POOL_SIZE,1,1],padding='SAME') # check k-size
```
Making the model
----------
__conv--ReLU--maxpool--fc__
```
def inference(input_data):
with tf.variable_scope('h_conv1') as scope:
weights = tf.get_variable('weights', shape=[KERNEL_1,1,1,FILTERS_1],
initializer=tf.contrib.layers.xavier_initializer_conv2d())
biases = tf.get_variable('biases', shape=[FILTERS_1], initializer=tf.constant_initializer(0.05))
z = tf.nn.conv2d(input_data, weights, strides=[1, 1, 1, 1], padding='VALID')
h_conv1 = tf.nn.relu(z+biases, name=scope.name)
h_pool1 = tf.nn.max_pool(h_conv1, ksize=[1, POOL_SIZE, 1, 1],
strides=[1, POOL_SIZE, 1, 1], padding='SAME', name='h_pool1')
h_pool2_flat = tf.reshape(h_pool2, [-1, 40*1*20])
with tf.variable_scope('h_fc') as scope:
weights = tf.get_variable('weights', shape=[HIDDEN_LAYER_1, OUTPUT_CLASSES],
initializer=tf.contrib.layers.xavier_initializer())
biases = tf.get_variable('biases', shape=[OUTPUT_CLASSES])
logits = (tf.matmul(h_pool2_flat, weights) + biases)
return logits
def calc_loss(logits, labels):
"""Calculates the loss from the logits and the labels.
Args:
logits: Logits tensor, float - [batch_size, NUM_CLASSES].
labels: Labels tensor, int32 - [batch_size].
Returns:
loss: Loss tensor of type float.
"""
labels = tf.to_float(labels)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(logits, labels, name='xentropy')
loss = tf.reduce_mean(cross_entropy, name='xentropy_mean')
return loss
def training(loss, learning_rate=5e-4):
"""Sets up the training Ops.
Creates a summarizer to track the loss over time in TensorBoard.
Creates an optimizer and applies the gradients to all trainable variables.
The Op returned by this function is what must be passed to the
`sess.run()` call to cause the model to train.
Args:
loss: Loss tensor, from loss().
learning_rate: The learning rate to use for gradient descent.
Returns:
train_op: The Op for training.
"""
# Add a scalar summary for the snapshot loss.
tf.scalar_summary(loss.op.name, loss)
# Create the Adam optimizer with the given learning rate.
optimizer = tf.train.AdamOptimizer(learning_rate)
# Create a variable to track the global step.
global_step = tf.Variable(0, name='global_step', trainable=False)
# Use the optimizer to apply the gradients that minimize the loss
# (and also increment the global step counter) as a single training step.
train_op = optimizer.minimize(loss, global_step=global_step)
return train_op
# def evaluation(logits, labels, topk=1):
# """Evaluate the quality of the logits at predicting the label.
# Args:
# logits: Logits tensor, float - [batch_size, NUM_CLASSES].
# labels: Labels tensor, int32 - [batch_size], with values in the
# range [0, NUM_CLASSES).
# topk: the number k for 'top-k accuracy'
# Returns:
# A scalar int32 tensor with the number of examples (out of batch_size)
# that were predicted correctly.
# """
# # For a classifier model, we can use the in_top_k Op.
# # It returns a bool tensor with shape [batch_size] that is true for
# # the examples where the label is in the top k (here k=1)
# # of all logits for that example.
# correct = tf.nn.in_top_k(logits, tf.reshape(tf.slice(labels, [0,1], [int(labels.get_shape()[0]), 1]),[-1]), topk)
# # Return the number of true entries.
# return tf.reduce_sum(tf.cast(correct, tf.int32))
def placeholder_inputs(batch_size):
"""Generate placeholder variables to represent the input tensors.
These placeholders are used as inputs by the rest of the model building
code and will be fed from the downloaded data in the .run() loop, below.
Args:
batch_size: The batch size will be baked into both placeholders.
Returns:
images_placeholder: Images placeholder.
labels_placeholder: Labels placeholder.
"""
# Note that the shapes of the placeholders match the shapes of the full
# image and label tensors, except the first dimension is now batch_size
# rather than the full size of the train or test data sets.
images_placeholder = tf.placeholder(tf.float32, shape=([None,INPUT_CHANNEL,1,1]))
labels_placeholder = tf.placeholder(tf.int32, shape=([None,OUTPUT_CLASSES]))
return images_placeholder, labels_placeholder
#UPDATE current_img_ind
def fill_feed_dict(data_set, images_pl, labels_pl, current_img_ind, batch_size):
"""Fills the feed_dict for training the given step.
A feed_dict takes the form of:
feed_dict = {
<placeholder>: <tensor of values to be passed for placeholder>,
....
}
Args:
data_set: The set of images and labels, from input_data.read_data_sets()
images_pl: The images placeholder, from placeholder_inputs().
labels_pl: The labels placeholder, from placeholder_inputs().
current_img_ind: The current position of the index in the dataset
Returns:
feed_dict: The feed dictionary mapping from placeholders to values.
current_img_ind: The updated position of the index in the dataset
data_set: updated data_set
"""
# Create the feed_dict for the placeholders filled with the next
# `batch size ` examples.
batch, current_img_ind, data_set= next_batch(batch_size, data_set, current_img_ind)
feed_dict = {
images_pl: batch[0],
labels_pl: batch[1]
}
return feed_dict, current_img_ind, data_set
def do_eval(sess, eval_correct, images_placeholder, labels_placeholder, data_set, batch_size):
"""Runs one evaluation against the full epoch of data.
Args:
sess: The session in which the model has been trained.
eval_correct: The Tensor that returns the number of correct predictions.
images_placeholder: The images placeholder.
labels_placeholder: The labels placeholder.
data_set: The set of images and labels to evaluate, from
input_data.read_data_sets().
"""
# And run one epoch of eval.
true_count = 0 # Counts the number of correct predictions.
steps_per_epoch = len(data_set) // batch_size
num_examples = steps_per_epoch * batch_size
current_img_ind = 0
for step in xrange(steps_per_epoch):
feed_dict, current_img_ind, data_set = fill_feed_dict(data_set, images_placeholder,
labels_placeholder, current_img_ind, batch_size)
true_count += sess.run(eval_correct, feed_dict=feed_dict)
precision = true_count / num_examples
print(' Num examples: %d Num correct: %d Precision @ 1: %0.04f' %
(num_examples, true_count, precision))
```
| github_jupyter |
# [译]音乐信息获取(MIR)之学习对音频的节拍追踪(Beat Tracking)
原文出处:[audio-beat-tracking-for-music-information-retrieval](https://www.analyticsvidhya.com/blog/2018/02/audio-beat-tracking-for-music-information-retrieval/)
本文在忠实原文的基础上补充了部分内容(如绘图,参考文献等)
## 介绍
音乐无处不在。当我们听到那种打动人心的音乐,我们就会整个沉浸在其中。同时,我们随着听到的节奏而打着拍子。你也一定注意到过你的腿不由自主的随着音乐的节拍而抖动。在这方面没有任何逻辑上可以解释我们为何会这样做,可能仅仅是因为我们沉浸在了音乐的律动中,我们的大脑开始对这种曲调产生共鸣。

假如我们能训练一个人工系统像我们人一样来捕捉这样的律动呢?可以想象,一个很酷的应用可能是构建一个具有表现力的类人机器人,它运行实时的节拍跟踪算法,使其在跳舞时能够与音乐保持同步。

原视频地址:[DARwin-OP dancing with beat tracking](https://www.youtube.com/watch?v=AJ--LrnkR6Y)
很有趣,对吧?
在本文中,我们将了解节拍(beats)的概念,以及在跟踪它时所面临的挑战。然后我们将学习解决这些问题的方法,和行业内最先进的解决方案。
注意:本文假设您具有python中音频数据分析的基本知识。如果没有,你可以阅读[这篇文章](https://www.analyticsvidhya.com/blog/2017/08/audio-voice-processing-deep-learning/),然后继续阅读。
## 目录
- 节拍追踪概览
- 什么是节拍追踪?
- 节拍追踪的应用
- 节拍追踪中的挑战
- 解决节拍追踪的方法
- 动态规划
- 循环神经网络 + 深度置信网
## 节拍追踪概览
### 什么是节拍追踪(Beat Tracking)
音频节拍追踪通常定义为确定音频记录中的时间实例,人类听众可能会随着音乐轻拍自己的脚。音频节拍跟踪技术使音乐的“拍同步”分析成为可能。
作为音乐信息获取(MIR)的一项重要和相关的任务,这一领域的研究十分活跃。自动节拍跟踪任务的目标是跟踪声音文件集合中的所有节拍位置,并为每个文件输出这些节拍开始时间。
为了给你一个对此任务的直观感受,下载并聆听下面的音频:
[原始音频](./res/teasure-trimed.wav)
[添加节拍注释的音频](./res/teasure-trimed-annotated.wav)
### 节拍追踪的应用
> 下面列举几个例子
1. 灯光随音乐节奏变化
2. Music-driven content的需要。音乐驱动的创作内容,如拍摄的视频短片,现在流行的一些音乐视频软件等,在融合音乐的时候都需要对节拍进行卡点。
3. 作为音乐分类的重要特征,等。
### 节拍追踪中的挑战
节拍跟踪可能听起来像一个简单的概念,但直到现在它实际上还是一个未解决的问题。 对于简单的曲调,算法可以很容易地从音乐中找出节拍。 但通常在现实生活中,音频要复杂得多,噪音也很大。 例如,可能存在来自环境的噪声,其可能会混淆算法并导致其在检测节拍时产生误报(False positive)。
从技术上讲,处理节拍跟踪有三个主要挑战:
1. 脉冲水平不清晰
2. 突然的节奏改变
3. 模糊/嘈杂的信息
## 节拍追踪中的可行方法
现在我们知道了一些有关节拍追踪的一些要点,下面我们来了解一下用来解决这个问题的一些方法。
音乐信息检索(music information retrival)算法的年度评估活动和ISMIR会议一起被称作音乐信息检索评估交换(MIREX)。其中有一个叫做音频街拍追踪的任务,研究人员参加MIREX并提交他们的方法,下面将介绍两种方法,第一种是简单而原始的,第二种是最先进的方法。
### 方法1: 突发点(onset)检测与动态规划
例如我们有下面的一段音频。

我们可以找到那种声音的突发点(也叫做onset)的位置,并且标注这些时间点,得到

这很可能是节拍的表现形式。 但它会包含许多误报(False positive),例如人的声音或背景噪音。 因此,为了最大限度地减少这些误报,我们可以找到这些起始点的**最长公共子序列**来识别节拍。 如果您想了解动态编程的工作原理,可以参考[这篇文章](https://www.analyticsvidhya.com/blog/2016/05/ase-studies-10x-faster-using-dynamic-programming/)。
下面的代码实现让你有一个更清晰的认识。
```
import librosa
import IPython.display as ipd
# read audio file
y, sr = librosa.load('./res/treasure-trimed.wav')
ipd.Audio(y, rate=sr)
# method 1 - onset detection and dynamic programming
tempo, beat_times = librosa.beat.beat_track(y, sr=sr, units='time')
clicks = librosa.clicks(beat_times,sr=sr,length=len(y))
ipd.Audio(y + clicks, rate=sr)
```
- 参考文献
> [1] Ellis, Daniel PW. “Beat tracking by dynamic programming.” Journal of New Music Research 36.1 (2007): 51-60. http://labrosa.ee.columbia.edu/projects/beattrack/
## 方法2: 集成循环神经网络(RNN)和动态贝叶斯网络
> 我们应该更多关注这部分
我们可以使用机器学习/深度学习方法,而不是手动地依赖声音的提示。 下面展示了用于解决节拍跟踪的框架的体系结构。 要了解它的细节,你可以阅读官方研究论文。
方法的要点是 - 我们预处理音频信号,然后使用递归神经网络找出这些节拍时间的最可能的值。 研究人员还使用了一系列循环神经网络(RNN),然后使用贝叶斯网络对其输出进行整合。如下图所示:

madmom库包含在节拍跟踪领域中各种最先进算法的实现,可在[github](https://github.com/CPJKU/madmom)上获得源码。 它结合了基于机器学习方法的低级特征提取和高级特征分析。 方法2的代码可以从此仓库中获得,其实现了从输入音频文件输出节拍位置。
下面我们看一下方法2的代码实现。
```
import madmom
# method 2 - dbn tracker
proc = madmom.features.beats.DBNBeatTrackingProcessor(fps=100, min_bpm=60)
act = madmom.features.beats.RNNBeatProcessor()('./res/treasure-trimed.wav')
beat_times = proc(act)
print(beat_times)
clicks = librosa.clicks(beat_times, sr=sr, length=len(y))
ipd.Audio(y + clicks, rate=sr)
```
- 参考文献
> [1] Sebastian Böck and Markus Schedl, “Enhanced Beat Tracking with Context-Aware Neural Networks”, Proceedings of the 14th International Conference on Digital Audio Effects (DAFx), 2011.
>
> [2] Florian Krebs, Sebastian Böck and Gerhard Widmer, “An Efficient State Space Model for Joint Tempo and Meter Tracking”, Proceedings of the 16th International Society for Music Information Retrieval Conference (ISMIR), 2015.
下面我们分析下方法2的处理步骤:
#### 1. 预处理音频信号

与所有非结构化数据一样,人工系统不容易掌握音频信号的概念。 因此必须以一种可以解释为机器学习模型的格式(即预处理数据)进行转换。 如下图所示,你可以使用很多方法来预处理音频数据。

1. 时域特征,如:波形的RMSE(均方根误差)
2. 频域特征,如:单个频率振幅
3. 感知特征,如:MFCC(梅尔频率倒谱系数)
4. 窗口特征,如:窗口的汉明距离
#### 2. 训练和微调RNN模型

现在你已经预处理了数据,你可以应用机器学习/深度学习模型来学习到数据中的模式。
从理论上讲,如果你训练一个具有足够训练样例和适当架构的深度学习模型,它可以很好地解决问题。 但由于训练数据可用性不足等原因,这并非总是可行的。因此,为了提高性能,我们可以做的是在**单一类型的音乐文件上训练多个RNN模型**,以便它可以捕获模式的这种类型(genre)本身。 这有助于缓解一些数据不足问题。
对于这种方法,我们首先训练LSTM模型(RNN的改进版本)并将其设置为我们的基本模型。 然后我们微调从我们的基础模型派生的多个LSTM模型。 这种微调是在不同类型的音乐序列上完成的。 在测试时,我们传递来自所有模型的音频信号。
#### 3. 选择最佳的RNN模型

在这一步,我们只是选择所有模型中误差最小的模型,将它们与我们从基础参照模型得到的分数进行比较。
#### 4. 应用动态贝叶斯网络

无论你使用一个LSTM还是多个LSTM,都存在一个根本性的缺点:在选择节拍的最终位置时,最终的峰值发现阶段并不试图找到全局最优值。它确定了作品(或一定长度的片段)的主要节奏,然后根据这个节奏排列节拍位置,简单地选择最好的起始位置,然后逐步地将节拍定位在预先确定的位置周围某个区域的激活函数值最高的位置。
为了避免这个问题,我们将所选神经网络模型的输出馈送到动态贝叶斯网络(DBN)中,该网络共同推断节拍序列的节奏和相位。使用DBN的另一个优点是我们能够对节拍状态和非节拍状态进行建模,这表明其性能优于仅对节拍状态进行建模的情况。我不会详细介绍DBN的工作原理,但如果您有兴趣,可以参考[此视频](https://youtu.be/lecy8kEjC3Q)。
这就是我们如何从原始音频信号中获得音乐序列的节拍的大致方法。
## 结语
我希望这篇文章能让你直观地了解如何在python中解决节拍跟踪问题。 这在音乐信息检索领域具有潜力的应用,我们可以使用我们在跟踪检测它时获得的节拍,来识别相似类型的音乐。
| github_jupyter |
# Paraphrase
<div class="alert alert-info">
This tutorial is available as an IPython notebook at [Malaya/example/paraphrase](https://github.com/huseinzol05/Malaya/tree/master/example/paraphrase).
</div>
```
%%time
import malaya
from pprint import pprint
```
### List available T5 models
```
malaya.paraphrase.available_t5()
```
### Load T5 models
```python
def t5(
model: str = 'base',
compressed: bool = True,
optimized: bool = False,
**kwargs,
):
"""
Load T5 model to generate a paraphrase given a string.
Parameters
----------
model : str, optional (default='base')
Model architecture supported. Allowed values:
* ``'base'`` - T5 BASE parameters.
* ``'small'`` - T5 SMALL parameters.
compressed: bool, optional (default=True)
Load compressed model, but this not able to utilize malaya-gpu function.
This only compressed model size, but when loaded into VRAM / RAM, size uncompressed and compressed are the same.
We prefer un-compressed model due to compressed model prone to error.
optimized : bool, optional (default=False)
if True, will load optimized uncompressed model, remove unnecessary nodes and fold batch norm to reduce model size.
Optimized model not necessary faster, totally depends on the machine.
We have no concrete proof optimized model maintain same accuracy as uncompressed model.
```
**For malaya-gpu user, compressed t5 very fragile and we suggest use `compressed=False`. Uncompressed model also can utilise GPU usage more efficient**.
```
t5 = malaya.paraphrase.t5()
```
### Paraphrase simple string
To paraphrase, simply use `paraphrase` method.
```
string = "Beliau yang juga saksi pendakwaan kesembilan berkata, ia bagi mengelak daripada wujud isu digunakan terhadap Najib."
pprint(string)
pprint(t5.paraphrase(string))
```
### Paraphrase longer string
```
string = """
PELETAKAN jawatan Tun Dr Mahathir Mohamad sebagai Pengerusi Parti Pribumi Bersatu Malaysia (Bersatu) ditolak di dalam mesyuarat khas Majlis Pimpinan Tertinggi (MPT) pada 24 Februari lalu.
Justeru, tidak timbul soal peletakan jawatan itu sah atau tidak kerana ia sudah pun diputuskan pada peringkat parti yang dipersetujui semua termasuk Presiden, Tan Sri Muhyiddin Yassin.
Bekas Setiausaha Agung Bersatu Datuk Marzuki Yahya berkata, pada mesyuarat itu MPT sebulat suara menolak peletakan jawatan Dr Mahathir.
"Jadi ini agak berlawanan dengan keputusan yang kita sudah buat. Saya tak faham bagaimana Jabatan Pendaftar Pertubuhan Malaysia (JPPM) kata peletakan jawatan itu sah sedangkan kita sudah buat keputusan di dalam mesyuarat, bukan seorang dua yang buat keputusan.
"Semua keputusan mesti dibuat melalui parti. Walau apa juga perbincangan dibuat di luar daripada keputusan mesyuarat, ini bukan keputusan parti.
"Apa locus standy yang ada pada Setiausaha Kerja untuk membawa perkara ini kepada JPPM. Seharusnya ia dibawa kepada Setiausaha Agung sebagai pentadbir kepada parti," katanya kepada Harian Metro.
Beliau mengulas laporan media tempatan hari ini mengenai pengesahan JPPM bahawa Dr Mahathir tidak lagi menjadi Pengerusi Bersatu berikutan peletakan jawatannya di tengah-tengah pergolakan politik pada akhir Februari adalah sah.
Laporan itu juga menyatakan, kedudukan Muhyiddin Yassin memangku jawatan itu juga sah.
Menurutnya, memang betul Dr Mahathir menghantar surat peletakan jawatan, tetapi ditolak oleh MPT.
"Fasal yang disebut itu terpakai sekiranya berhenti atau diberhentikan, tetapi ini mesyuarat sudah menolak," katanya.
Marzuki turut mempersoal kenyataan media yang dibuat beberapa pimpinan parti itu hari ini yang menyatakan sokongan kepada Perikatan Nasional.
"Kenyataan media bukanlah keputusan rasmi. Walaupun kita buat 1,000 kenyataan sekali pun ia tetap tidak merubah keputusan yang sudah dibuat di dalam mesyuarat. Kita catat di dalam minit apa yang berlaku di dalam mesyuarat," katanya.
"""
import re
# minimum cleaning, just simply to remove newlines.
def cleaning(string):
string = string.replace('\n', ' ')
string = re.sub(r'[ ]+', ' ', string).strip()
return string
string = cleaning(string)
pprint(string)
```
#### T5 model
```
pprint(t5.paraphrase(string))
```
You can see `Griffin` out-of-context, this is because the model trying to predict who is `katanya`, so it simply pulled random name from training set. To solve this problem, you need to do sliding windows. If we have 5 strings, simply give [s1, s2], [s2, s3] and so on the model, at least the model got some context from previous string.
```
pprint(t5.paraphrase(string, split_fullstop = False))
```
When you try to paraphrase entire string, the output is quite good, a summary!
### List available LM Transformer models
Problem with T5 models, it built on top of mesh-tensorflow, so the input must size of 1. So we use Tensor2Tensor library to train exact model as T5 with dynamic size of batch.
**But, we found out, our pretrained LM Transformer not good as T5**, we might skipped some literature in t5 papers.
```
malaya.paraphrase.available_transformer()
```
### Load Transformer
```
model = malaya.paraphrase.transformer()
```
### Load Quantized model
To load 8-bit quantized model, simply pass `quantized = True`, default is `False`.
We can expect slightly accuracy drop from quantized model, and not necessary faster than normal 32-bit float model, totally depends on machine.
```
quantized_model = malaya.paraphrase.transformer(quantized = True)
```
#### decoder mode
LM Transformer provided 3 different decoder for summarization,
1. greedy decoder, simply argmax,
```python
model.summarization([string], decoder = 'greedy')
```
2. beam decoder, Beam width size 3, alpha 0.5 .
```python
model.summarization([string], decoder = 'beam')
```
3. nucleus sampling decoder, Beam width size 1, with nucleus sampling.
```python
model.summarization([string], decoder = 'nucleus', top_p = 0.7)
```
default is `greedy`,
```python
def paraphrase(
self,
strings: List[str],
decoder: str = 'greedy',
top_p: float = 0.7,
):
"""
Summarize strings.
Parameters
----------
decoder: str
mode for summarization decoder. Allowed values:
* ``'greedy'`` - Beam width size 1, alpha 0.
* ``'beam'`` - Beam width size 3, alpha 0.5 .
* ``'nucleus'`` - Beam width size 1, with nucleus sampling.
top_p: float, (default=0.7)
cumulative distribution and cut off as soon as the CDF exceeds `top_p`.
this is only useful if use `nucleus` decoder.
```
```
string = """
PELETAKAN jawatan Tun Dr Mahathir Mohamad sebagai Pengerusi Parti Pribumi Bersatu Malaysia (Bersatu) ditolak di dalam mesyuarat khas Majlis Pimpinan Tertinggi (MPT) pada 24 Februari lalu.
Justeru, tidak timbul soal peletakan jawatan itu sah atau tidak kerana ia sudah pun diputuskan pada peringkat parti yang dipersetujui semua termasuk Presiden, Tan Sri Muhyiddin Yassin.
Bekas Setiausaha Agung Bersatu Datuk Marzuki Yahya berkata, pada mesyuarat itu MPT sebulat suara menolak peletakan jawatan Dr Mahathir.
"Jadi ini agak berlawanan dengan keputusan yang kita sudah buat. Saya tak faham bagaimana Jabatan Pendaftar Pertubuhan Malaysia (JPPM) kata peletakan jawatan itu sah sedangkan kita sudah buat keputusan di dalam mesyuarat, bukan seorang dua yang buat keputusan.
"Semua keputusan mesti dibuat melalui parti. Walau apa juga perbincangan dibuat di luar daripada keputusan mesyuarat, ini bukan keputusan parti.
"Apa locus standy yang ada pada Setiausaha Kerja untuk membawa perkara ini kepada JPPM. Seharusnya ia dibawa kepada Setiausaha Agung sebagai pentadbir kepada parti," katanya kepada Harian Metro.
Beliau mengulas laporan media tempatan hari ini mengenai pengesahan JPPM bahawa Dr Mahathir tidak lagi menjadi Pengerusi Bersatu berikutan peletakan jawatannya di tengah-tengah pergolakan politik pada akhir Februari adalah sah.
Laporan itu juga menyatakan, kedudukan Muhyiddin Yassin memangku jawatan itu juga sah.
Menurutnya, memang betul Dr Mahathir menghantar surat peletakan jawatan, tetapi ditolak oleh MPT.
"Fasal yang disebut itu terpakai sekiranya berhenti atau diberhentikan, tetapi ini mesyuarat sudah menolak," katanya.
Marzuki turut mempersoal kenyataan media yang dibuat beberapa pimpinan parti itu hari ini yang menyatakan sokongan kepada Perikatan Nasional.
"Kenyataan media bukanlah keputusan rasmi. Walaupun kita buat 1,000 kenyataan sekali pun ia tetap tidak merubah keputusan yang sudah dibuat di dalam mesyuarat. Kita catat di dalam minit apa yang berlaku di dalam mesyuarat," katanya.
"""
import re
# minimum cleaning, just simply to remove newlines.
def cleaning(string):
string = string.replace('\n', ' ')
string = re.sub(r'[ ]+', ' ', string).strip()
return string
string = cleaning(string)
splitted = malaya.text.function.split_into_sentences(string)
' '.join(splitted[:2])
model.paraphrase([' '.join(splitted[:2])], decoder = 'greedy')
quantized_model.paraphrase([' '.join(splitted[:2])], decoder = 'greedy')
model.paraphrase([' '.join(splitted[:2])], decoder = 'beam')
quantized_model.paraphrase([' '.join(splitted[:2])], decoder = 'beam')
model.paraphrase([' '.join(splitted[:2])], decoder = 'nucleus', top_p = 0.7)
```
| github_jupyter |
```
import PyPDF2
import re
from nltk.stem import PorterStemmer
from sklearn.feature_extraction.text import CountVectorizer
import pandas as pd
from sklearn.metrics import silhouette_score
from sklearn.cluster import KMeans
import matplotlib.pyplot as plt
from sklearn.preprocessing import normalize
import os,json
import numpy as np
import nltk
class convertCVtoText:
@staticmethod
def startConversion(fileName):
pdfFileObj = open(fileName,'rb') #'rb' for read binary mode
pdfReader = PyPDF2.PdfFileReader(pdfFileObj)
noOfPages=pdfReader.numPages
text=""
for pages in range(0,pdfReader.numPages):
pageObj = pdfReader.getPage(pages) #'9' is the page number
text+=pageObj.extractText()
return text
class textCleaner:
def __init__(self):
self.text=""
self.cleanText=""
def normalizeText(self):
norText=""
returnText=""
norText+= re.sub(r'[^a-zA-Z ]',r' ',self.text)
returnText+=re.sub(' +',' ',norText)
self.cleanText+=re.sub(r'([A-Z])', lambda pat: pat.group(1).lower(), returnText)
class CV:
def __init__(self,name,path,post):
self.fileName=name
self.filePath=path
self.CVCategory=post
self.textHandeller=textCleaner()
self.textHandeller.text=convertCVtoText.startConversion(self.filePath)
self.textHandeller.normalizeText()
self.featureVector=[]
self.score=None
relevantWords1=[
'ip',
'firewall',
'layer',
'wan',
'protocol',
'router',
'switch',
'traffic',
'css',
'design',
'html',
'javascript',
'jquery',
'mysql',
'ajax',
'php',
'unity','game','team','computer','engine','software','programming','developer','microsoft','project',
'animation','adobe','flash','character','art','illustrator','design','animator','effects','maya','photoshop',
"software","skills","application","developer","server",
"systems","framework","net","visual",
"algorithm","analyst","aws","datasets","clustering","intelligence","logistic","mining","neural","regression","scikit"
]
relevantWords=list(set(relevantWords1))
print(len(relevantWords1))
print(len(relevantWords))
def readCSVFile(fileName,row=None,col=None,):
pd.set_option('display.max_columns', col)
pd.set_option('display.max_rows', row)
#pd.set_option('display.max_colwidth',1000)
return pd.read_csv(fileName,names=set(relevantWords))
import numpy as np
from sklearn.metrics.pairwise import pairwise_distances
from skfeature.utility.util import reverse_argsort
class refiefFAlgo:
def __init__(self,mode=None):
self.scoreList=[]
self.wordIndex=[]
self.mode=mode
def feature_ranking(self):
"""
Rank features in descending order according to reliefF score, the higher the reliefF score, the more important the
feature is
"""
for scorePerCluster in self.scoreList:
temp=np.asarray(scorePerCluster)
idx = np.argsort(temp, 0)
#print(idx)
self.wordIndex.append(idx[::-1])
#return idx[::-1]
def score_ranking(self):
"""
Rank features in descending order according to reliefF score, the higher the reliefF score, the more important the
feature is
"""
for scorePerCluster in self.scoreList:
temp=np.asarray(scorePerCluster)
idx = np.argsort(temp, 0)
#print(idx)
self.wordIndex.append(idx[::-1])
#return idx[::-1]
def reliefF(self,X, y,**kwargs):
"""
This function implements the reliefF feature selection
Input
-----
X: {numpy array}, shape (n_samples, n_features)
input data
y: {numpy array}, shape (n_samples,)
input class labels
kwargs: {dictionary}
parameters of reliefF:
k: {int}
choices for the number of neighbors (default k = 5)
Output
------
score: {numpy array}, shape (n_features,)
reliefF score for each feature
Reference
---------
Robnik-Sikonja, Marko et al. "Theoretical and empirical analysis of relieff and rrelieff." Machine Learning 2003.
Zhao, Zheng et al. "On Similarity Preserving Feature Selection." TKDE 2013.
"""
if "k" not in list(kwargs.keys()):
k = 5
else:
k = kwargs["k"]
n_samples, n_features = X.shape
# calculate pairwise distances between instances
distance = pairwise_distances(X, metric='manhattan')
# the number of sampled instances is equal to the number of total instances
for idx in range(n_samples):
score = np.zeros(n_features)
near_hit = []
near_miss = dict()
self_fea = X[idx, :]
c = np.unique(y).tolist()
stop_dict = dict()
for label in c:
stop_dict[label] = 0
del c[c.index(y[idx])]
p_dict = dict()
p_label_idx = float(len(y[y == y[idx]]))/float(n_samples)
for label in c:
p_label_c = float(len(y[y == label]))/float(n_samples)
p_dict[label] = p_label_c/(1-p_label_idx)
near_miss[label] = []
distance_sort = []
distance[idx, idx] = np.max(distance[idx, :])
for i in range(n_samples):
distance_sort.append([distance[idx, i], int(i), y[i]])
distance_sort.sort(key=lambda x: x[0])
for i in range(n_samples):
# find k nearest hit points
if distance_sort[i][2] == y[idx]:
if len(near_hit) < k:
near_hit.append(distance_sort[i][1])
elif len(near_hit) == k:
stop_dict[y[idx]] = 1
else:
# find k nearest miss points for each label
if len(near_miss[distance_sort[i][2]]) < k:
near_miss[distance_sort[i][2]].append(distance_sort[i][1])
else:
if len(near_miss[distance_sort[i][2]]) == k:
stop_dict[distance_sort[i][2]] = 1
stop = True
for (key, value) in list(stop_dict.items()):
if value != 1:
stop = False
if stop:
break
# update reliefF score
near_hit_term = np.zeros(n_features)
for ele in near_hit:
near_hit_term = np.array(abs(self_fea-X[ele, :]))+np.array(near_hit_term)
near_miss_term = dict()
for (label, miss_list) in list(near_miss.items()):
near_miss_term[label] = np.zeros(n_features)
for ele in miss_list:
near_miss_term[label] = np.array(abs(self_fea-X[ele, :]))+np.array(near_miss_term[label])
score += near_miss_term[label]/(k*p_dict[label])
score -= near_hit_term/k
self.scoreList.append(score)
self.feature_ranking()
if self.mode == 'raw':
print("here")
print(score)
return score
elif self.mode == 'index':
print("herew")
return feature_ranking(score)
elif self.mode == 'rank':
print("hereq")
return reverse_argsort(feature_ranking(score), X.shape[1])
class CBRAlgo:
def __init__(self):
self.CVScoreList=[]
self.topWords=None
self.clusterWiseTopWordList=[]
self.overAllWeight=[]
def getTopWords(self,vocabulary,clusterCenters,noOfFormedClusters):
x=[]
for key,values in vocabulary.items():
x.append(values)
self.topWords=refiefFAlgo()
self.topWords.reliefF(clusterCenters,np.asarray(x),k=noOfFormedClusters-1)
def getOverallWeightOfRelevantWords(self):
self.overAllWeight=np.average([x for x in self.topWords.scoreList],axis=0)
def calculateCVScoreViaCluster(self,documentMatrix,vocabulary,clusteringInfo):
self.getTopWords(vocabulary,clusteringInfo.kMeans.cluster_centers_,clusteringInfo.bestClusterToForm)
self.getTopWordsPerCluster(clusteringInfo.kMeans.cluster_centers_,vocabulary)
self.getOverallWeightOfRelevantWords()
featureVector=documentMatrix.toarray()
for cvNumber,clusterNumber in enumerate(clusteringInfo.kMeans.labels_):
score=0
for wordFrequency,weight in zip(featureVector[cvNumber],self.topWords.scoreList[clusterNumber]):
score+=wordFrequency*weight
self.CVScoreList.append(score)
def calculateCVScore(self,documentMatrix,vocabulary,clusteringInfo):
self.getTopWords(vocabulary,clusteringInfo.kMeans.cluster_centers_,clusteringInfo.bestClusterToForm)
self.getTopWordsPerCluster(clusteringInfo.kMeans.cluster_centers_,vocabulary)
self.getOverallWeightOfRelevantWords()
featureVector=documentMatrix.toarray()
for cvNumber,clusterNumber in enumerate(clusteringInfo.kMeans.labels_):
score=0
for wordFrequency,weight in zip(featureVector[cvNumber],self.overAllWeight):
score+=wordFrequency*weight
self.CVScoreList.append(score)
def getTopWordsPerCluster(self,clusterCenters,vocabulary):
for clusterNo,impFeaturesRow in enumerate(clusterCenters):
WordList={}
for indexNo in self.topWords.wordIndex[clusterNo]:
WordList.update({list(vocabulary.keys())[list(vocabulary.values()).index(indexNo)]:self.topWords.scoreList[clusterNo][indexNo]})
self.clusterWiseTopWordList.append(WordList)
def plotTopWordsPerCluster(self):
topwords=10
width =1
import matplotlib.pyplot as plt
for index,clusterTopWord in enumerate(self.clusterWiseTopWordList):
ig,ax = plt.subplots()
lists = [(key,value) for (key,value) in clusterTopWord.items()] # sorted by key, return a list of tuples
key, value = zip(*lists)
x = np.arange(topwords)
plt.barh(x[:topwords],value[:topwords],align='center')
plt.yticks(x, key[:topwords])
#plt.rcParams["figure.figsize"] = (10,5)
plt.title('top %d words in %d cluster'%(topwords,index))
plt.ylabel('words')
plt.xlabel('weight')
plt.show()
def plotOverAllWeight(self,vocabulary):
import matplotlib.pyplot as pl
ig,ax = plt.subplots()
x = np.arange(0,len(self.overAllWeight))
pl.barh(x,self.overAllWeight,align='center')
pl.yticks(x,vocabulary)
#pl.rcParams["figure.figsize"] = (20,20)
pl.title('weight bar graph of Relevant words')
pl.ylabel('words')
pl.xlabel('weight')
pl.show()
class NLTKHelper:
def __init__(self):
self.documentMatrix=None
self.vocabulary=None
self.normalizedFeatureSet=[]
def findDocumentMatrix(self,totalCVText,minFrequency,vocab):
#vectorizer=CountVectorizer(stop_words='english',min_df=minFrequency)
vectorizer=CountVectorizer(stop_words='english',vocabulary=vocab)
#vectorizer=CountVectorizer(stop_words='english')
self.documentMatrix=vectorizer.fit_transform(totalCVText)
self.vocabulary=vectorizer.vocabulary_
#return documentMatrix,vocabulary
self.normalizeMatrix()
def normalizeMatrix(self):
self.normalizedFeatureSet=normalize(self.documentMatrix.toarray().astype('float64'))
class Clustering:
def __init__(self):
self.kMeans=None
self.bestClusterToForm=None
self.silCoeffInfo={}
self.minCluster=2
self.maxCluster=10
def findSilCoeff(self,data):
for n_cluster in range(self.minCluster, self.maxCluster):
kmeans = KMeans(n_clusters=n_cluster).fit(data)
label = kmeans.labels_
sil_coeff = silhouette_score(data, label, metric='euclidean')
self.silCoeffInfo.update({n_cluster:float(sil_coeff)})
maxSilCoeff=max(self.silCoeffInfo.values())
maxSilCoeffkeys = [k for k, v in self.silCoeffInfo.items() if v == maxSilCoeff]
if(len(maxSilCoeffkeys)==1):
for x in maxSilCoeffkeys:
self.bestClusterToForm=x
else:
print("2 keys,confusion")
def clusterData(self,data):
self.findSilCoeff(data)
self.kMeans=KMeans(n_clusters=self.bestClusterToForm).fit(data)
class CVManager:
def __init__(self):
self.CVList=[]
self.cvsFile="documentMatrix.csv"
self.CVFileName=[]
self.fileNamesWithPath=[]
self.cvPostList=[]
self.CVTextColl=[]
self.noOfTopCV=10
self.languageProcessing=None
self.clusteringInfo=None
self.CVRanker=None
def list_CVs(self,rootPath):
for root, dirs, files in os.walk(rootPath):
for name in files:
self.CVFileName.append(name)
self.fileNamesWithPath.append(os.path.join(root, name))
self.cvPostList.append(os.path.basename(os.path.dirname(os.path.join(root,name))))
def collectCV(self):
for cvFilePath,cvFileName,cvPost in zip(self.fileNamesWithPath,self.CVFileName,self.cvPostList):
try:
newCV=CV(cvFileName,cvFilePath,cvPost)
self.CVList.append(newCV)
except Exception as e:
print(cvFileName)
print("in collection of CV \t"+str(e))
def collectCVText(self):
self.CVTextColl=[]
for cv in self.CVList:
self.CVTextColl.append(cv.textHandeller.cleanText)
def findDocumentMatrix(self,minFrequency,vocab):
self.collectCVText()
self.languageProcessing=NLTKHelper()
self.languageProcessing.findDocumentMatrix(self.CVTextColl,minFrequency,vocab)
#df = pd.DataFrame(self.languageProcessing.documentMatrix.toarray())
#df.to_csv(self.cvsFile)
self.assignFeatureVector()
def assignFeatureVector(self):
for cv,cvNum in zip(self.CVList,range(0,len(self.CVFileName)-1)):
for featureRow in range(0,len(self.languageProcessing.vocabulary)-1):
cv.featureVector.append(self.languageProcessing.normalizedFeatureSet[cvNum][featureRow])
def makeGraph(self,data):
lists = sorted(data) # sorted by key, return a list of tuples
x, y = zip(*lists) # unpack a list of pairs into two tuples
plt.plot(x, y)
plt.show()
def clusterData(self):
self.clusteringInfo=Clustering()
self.clusteringInfo.clusterData(self.languageProcessing.normalizedFeatureSet)
self.makeGraph(self.clusteringInfo.silCoeffInfo.items())
def rankCV(self):
self.CVRanker=CBRAlgo()
self.CVRanker.calculateCVScore(self.languageProcessing.documentMatrix,self.languageProcessing.vocabulary,self.clusteringInfo)
for cv,cvScore in zip(self.CVList,self.CVRanker.CVScoreList):
cv.score=cvScore
def showAnalytics(self):
self.CVRanker.plotTopWordsPerCluster()
self.CVRanker.plotOverAllWeight(self.languageProcessing.vocabulary)
def showTopCVPerPost(self,post):
cvlist={}
cvScore=[]
cvData=[]
if post is None:
for cvCategery in set(self.cvPostList):
cvlist={}
print("cv of %s"%cvCategery)
for cv in self.CVList:
if(cv.CVCategory==cvCategery):
cvlist.update({cv.fileName:cv.score})
temp=[(value,key) for key,value in cvlist.items()]
temp.sort()
temp.reverse()
temp=[(key,value) for value,key in temp]
cvData=temp
return cvData[:self.noOfTopCV]
else:
print("cv of post %s"%post)
for cv in self.CVList:
if(cv.CVCategory==post):
cvlist.update({cv.filePath:cv.score})
temp=[(value,key) for key,value in cvlist.items()]
temp.sort()
temp.reverse()
temp=[(key,value) for value,key in temp]
cvData=temp
return cvData[:self.noOfTopCV]
class communicationInformation:
def __init__(self):
self.directoryPath=""
self.relevantWords=[]
self.jobSelected=""
self.workFlow=True
relevantWords1=[
'ip',
'firewall',
'layer',
'wan',
'protocol',
'router',
'switch',
'traffic',
'css',
'design',
'html',
'javascript',
'jquery',
'mysql',
'ajax',
'php',
'unity','game','team','computer','engine','software','programming','developer','microsoft','project',
'animation','adobe','flash','character','art','illustrator','design','animator','effects','maya','photoshop',
"software","skills","application","developer","server",
"systems","framework","net","visual",
"algorithm","analyst","aws","datasets","clustering","intelligence","logistic","mining","neural","regression","scikit"
]
self.relevantWords=list(set(relevantWords1))
import webbrowser
class buttonHandler:
def __init__(self, master,buttonDataList):
self.buttonList=[]
self.frame = Frame(master)
self.frame.pack()
for buttonName in buttonDataList:
buttonToShow=Button(self.frame,text=str(buttonName[0][50:]),command=lambda:self.openPdf(buttonName[0]))
buttonToShow.pack(padx=5,pady=10)
self.buttonList.append(buttonToShow)
def openPdf(self,link):
webbrowser.open_new_tab(link)
def destoringButtons(self):
self.frame.destroy()
from tkinter import*
import tkinter.filedialog
from tkinter import Tk, StringVar, ttk
import webbrowser
class GUI:
def __init__(self,root):
self.passingInfo=communicationInformation()
self.root=root
self.run=False
self.frame = Frame()
self.frame.pack(fill=X)
self.directoryLabel = Label(self.frame,text="Path", width=10)
self.directoryLabel.pack(side=LEFT)
self.directoryEntry = Entry(self.frame,width=100)
self.directoryEntry.pack(side=LEFT, padx=0,pady=10 ,expand=True)
self.addDirectoryButton = Button(self.root, text =" Add ",command=self.pathAdd)
self.addDirectoryButton.pack(padx=5,pady=10)
self.buttonList=None
self.jobReqFrame = Frame()
self.jobReqFrame.pack(side=LEFT,fill=X)
self.jobRequirementLabel = Label(self.jobReqFrame, text="Jobs Requirements", width=50)
self.jobRequirementLabel.pack(expand=True, padx=5, pady=5)
self.relevantWordsText = Text(self.jobReqFrame,width=40,height=15)
self.relevantWordsText.pack( side=LEFT,pady=5, padx=5)
self.relevantWordsText.insert('1.0',' '.join(self.passingInfo.relevantWords))
self.CVTitleFrame=Frame()
self.CVTitleFrame.pack(fill=X)
self.CVTitleLabel = Label(self.CVTitleFrame, text="Selecte Post", width=50)
self.CVTitleLabel.pack( anchor=N, padx=0, pady=0)
self.topCVDisplay = Frame()
self.topCVDisplay.pack(fill=X,padx=50)
self.box_value = StringVar()
self.box = ttk.Combobox(self.topCVDisplay, textvariable=self.box_value,state='readonly')
self.box['values'] = ('game developer', 'animator', 'network engineer','web developer','DataScientist','Software developer')
self.box.grid(column=0, row=0)
#process button
self.processButton = Button(self.jobReqFrame, text =" Process ",command=self.processExe)
self.processButton.pack(side=LEFT,padx=10,pady=10)
self.exitButton = Button(self.root, text =" Exit ",command=self.root.destroy)
self.exitButton.pack(side=LEFT,padx=10,pady=10)
self.manager=CVManager()
def pathAdd(self):
directoryPath=filedialog.askdirectory()
self.directoryEntry.insert(0, directoryPath)
def processExe(self):
if(not(self.run)):
self.passingInfo.directoryPath=self.directoryEntry.get()
self.passingInfo.jobSelected=self.box.get()
self.passingInfo.relevantWords=self.relevantWordsText.get('1.0',END).strip().split(" ")
# if self.passingInfo.relevantWords:
# if self.passingInfo.jobSelected:
# if self.passingInfo.directoryPath:
# self.passingInfo.workFlow=True
if self.passingInfo.workFlow:
try:
self.run=True
print(self.passingInfo.relevantWords)
self.manager.list_CVs(self.passingInfo.directoryPath)
self.manager.collectCV()
self.manager.findDocumentMatrix(None,relevantWords)
self.manager.clusterData()
self.manager.rankCV()
self.manager.showAnalytics()
CVDataList=self.manager.showTopCVPerPost(self.passingInfo.jobSelected)
self.createLinkToCV(CVDataList)
except Exception as e:
print("error")
print("processing \t"+str(e))
else:
print("select all necessary info")
else:
self.passingInfo.jobSelected=self.box.get()
CVDataList=self.manager.showTopCVPerPost(self.passingInfo.jobSelected)
self.buttonList.destoringButtons()
self.createLinkToCV(CVDataList)
def createLinkToCV(self,dataList):
self.buttonList=buttonHandler(self.root,dataList)
root=Tk()
graphics=GUI(root)
root.geometry("700x600")
root.mainloop()
manager=CVManager()
manager.list_CVs("CV coll")
manager.collectCV()
from tkinter import*
def changebutton():
but.destroy()
secondbut=Button(root,text="changed")
secondbut.pack()
if __name__=='__main__':
root=Tk()
global but
but= Button(root,text="button",command=changebutton)
but.pack()
root.mainloop()
#use this function to test the feature vector given by relevant words
#manager.languageProcessing.findDocumentMatrix(None,relevantWords)
#use this function to find the most ferquently words as given in function
#it will help to find the relevant word for that post
#manager.findDocumentMatrix(70,None)
#manager.vocabulary
#manager.normalizedFeatureSet[0]
readCSVFile(manager.cvsFile)
manager.clusterData()
manager.rankCV()
manager.showAnalytics()
a=[("ramdsfaaaaaaaaaaaaaaaas","harisadfffffffff"),("ritaasadfffffffffff","sitaasfddddddddddd")]
for item in a:
print(item[0][10:])
from tkinter import *
class App:
def __init__(self, master):
frame = Frame(master)
frame.pack()
self.button = Button(frame,
text="QUIT", fg="red",
command=quit)
self.button.pack(side=LEFT)
self.slogan = Button(frame,
text="Hello",
command=self.write_slogan)
self.slogan.pack(side=LEFT)
def write_slogan(self):
print("Tkinter is easy to use!")
root = Tk()
app = App(root)
root.mainloop()
a=[]
if a:
print("fs")
a.append("ra_")
# import numpy as np
# import matplotlib.pyplot as plt
# from sklearn.cluster import KMeans
# x = [916,684,613,612,593,552,487,484,475,474,438,431,421,418,409,391,389,388,
# 380,374,371,369,357,356,340,338,328,317,316,315,313,303,283,257,255,254,245,
# 234,232,227,227,222,221,221,219,214,201,200,194,169,155,140]
# kmeans = KMeans(n_clusters=4)
# a = kmeans.fit(np.reshape(x,(len(x),1)))
# centroids = kmeans.cluster_centers_
# labels = kmeans.labels_
# print(centroids)
# print(labels)
# colors = ["g.","r.","y.","b."]
# for i in centroids:
# plt.plot( [0, len(x)-1],[i,i], "k" )
# for i in range(len(x)):
# plt.plot(i, x[i], colors[labels[i]], markersize = 10)
# plt.show()
# import matplotlib.pyplot as plt
# topwords=10
# width =1
# for index,clusterTopWord in enumerate(manager.CVRanker.clusterWiseTopWordList):
# ig,ax = plt.subplots()
# lists = [(key,value) for (key,value) in clusterTopWord.items()] # sorted by key, return a list of tuples
# key, value = zip(*lists)
# x = np.arange(topwords)
# plt.barh(x[:topwords],value[:topwords],align='center')
# plt.yticks(x, key[:topwords])
# plt.rcParams["figure.figsize"] = (10,5)
# plt.title('top %d words in %d cluster'%(topwords,index))
# plt.ylabel('words')
# plt.xlabel('weight')
# plt.show()
# #for index,weight in enumerate(manager.overAllWeight)
# import matplotlib.pyplot as pl
# ig,ax = plt.subplots()
# #lists = [(key,value) for (key,value) in clusterTopWord.items()] # sorted by key, return a list of tuples
# #key, value = zip(*lists)
# x = np.arange(0,len(manager.CVRanker.overAllWeight))
# print(x)
# pl.barh(x,manager.CVRanker.overAllWeight,align='center')
# pl.yticks(x,manager.languageProcessing.vocabulary)
# pl.rcParams["figure.figsize"] = (20,20)
# pl.title('weight bar graph of Relevant words')
# pl.ylabel('words')
# pl.xlabel('weight')
# pl.show()
# import matplotlib.pyplot as plt; plt.rcdefaults()
# import numpy as np
# import matplotlib.pyplot as plt
# objects = ['Python', 'C++', 'Java', 'Perl', 'Scala', 'Lisp']
# y_pos = np.arange(len(objects))
# print(y_pos)
# performance = [10,8,6,4,2,1]
# plt.bar(y_pos, performance, align='center')
# plt.xticks(y_pos, objects)
# plt.ylabel('Usage')
# plt.title('Programming language usage')
# plt.show()
# print(np.__version__)
# x = [1, 5, 1.5, 8, 1, 9]
# y = [2, 8, 1.8, 8, 0.6, 11]
# plt.scatter(x,y)
# plt.show()
# colors = ["g.","r.","c.","y."]
# for i in range(len(X)):
# print("coordinate:",X[i], "label:", labels[i])
# plt.plot(X[i][0], X[i][1], colors[labels[i]], markersize = 10)
# plt.scatter(centroids[:, 0],centroids[:, 1], marker = "x", s=150, linewidths = 5, zorder = 10)
# plt.show()
# x={1:3,2:1,4:56}
# y=list(x.values())
# type(max(y))
import tkinter as tk
counter = 0
def counter_label(label):
counter = 0
def count():
global counter
counter += 1
label.config(text=str(counter))
label.after(1000, count)
count()
root = tk.Tk()
root.title("Counting Seconds")
label = tk.Label(root, fg="dark green")
label.pack()
counter_label(label)
button = tk.Button(root, text='Stop', width=25, command=root.destroy)
button.pack()
root.mainloop()
```
| github_jupyter |
# CNTK 103: Part A - MNIST Data Loader
This tutorial is targeted to individuals who are new to CNTK and to machine learning. We assume you have completed or are familiar with CNTK 101 and 102. In this tutorial, we will download and pre-process the MNIST digit images to be used for building different models to recognize handwritten digits. We will extend CNTK 101 and 102 to be applied to this data set. Additionally, we will introduce a convolutional network to achieve superior performance. This is the first example, where we will train and evaluate a neural network based model on real world data.
CNTK 103 tutorial is divided into multiple parts:
- Part A: Familiarize with the [MNIST](http://yann.lecun.com/exdb/mnist/) database that will be used later in the tutorial
- Subsequent parts in this 103 series will be using the MNIST data with different types of networks.
```
# Import the relevant modules to be used later
from __future__ import print_function
import gzip
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import numpy as np
import os
import shutil
import struct
import sys
try:
from urllib.request import urlretrieve
except ImportError:
from urllib import urlretrieve
# Config matplotlib for inline plotting
%matplotlib inline
```
## Data download
We will download the data to the local machine. The MNIST database contains standard handwritten digits that have been widely used for training and testing of machine learning algorithms. It has a training set of 60,000 images and a test set of 10,000 images with each image being 28 x 28 pixels. This set is easy to use visualize and train on any computer.
```
# Functions to load MNIST images and unpack into train and test set.
# - loadData reads a image and formats it into a 28x28 long array
# - loadLabels reads the corresponding label data, one for each image
# - load packs the downloaded image and label data into a combined format to be read later by
# the CNTK text reader
def loadData(src, cimg):
print ('Downloading ' + src)
gzfname, h = urlretrieve(src, './delete.me')
print ('Done.')
try:
with gzip.open(gzfname) as gz:
n = struct.unpack('I', gz.read(4))
# Read magic number.
if n[0] != 0x3080000:
raise Exception('Invalid file: unexpected magic number.')
# Read number of entries.
n = struct.unpack('>I', gz.read(4))[0]
if n != cimg:
raise Exception('Invalid file: expected {0} entries.'.format(cimg))
crow = struct.unpack('>I', gz.read(4))[0]
ccol = struct.unpack('>I', gz.read(4))[0]
if crow != 28 or ccol != 28:
raise Exception('Invalid file: expected 28 rows/cols per image.')
# Read data.
res = np.fromstring(gz.read(cimg * crow * ccol), dtype = np.uint8)
finally:
os.remove(gzfname)
return res.reshape((cimg, crow * ccol))
def loadLabels(src, cimg):
print ('Downloading ' + src)
gzfname, h = urlretrieve(src, './delete.me')
print ('Done.')
try:
with gzip.open(gzfname) as gz:
n = struct.unpack('I', gz.read(4))
# Read magic number.
if n[0] != 0x1080000:
raise Exception('Invalid file: unexpected magic number.')
# Read number of entries.
n = struct.unpack('>I', gz.read(4))
if n[0] != cimg:
raise Exception('Invalid file: expected {0} rows.'.format(cimg))
# Read labels.
res = np.fromstring(gz.read(cimg), dtype = np.uint8)
finally:
os.remove(gzfname)
return res.reshape((cimg, 1))
def try_download(dataSrc, labelsSrc, cimg):
data = loadData(dataSrc, cimg)
labels = loadLabels(labelsSrc, cimg)
return np.hstack((data, labels))
```
## Download the data
The MNIST data is provided as a training and test set. Training set has 60000 images while the test set has 10000 images. Let us download the data.
```
# URLs for the train image and label data
url_train_image = 'http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz'
url_train_labels = 'http://yann.lecun.com/exdb/mnist/train-labels-idx1-ubyte.gz'
num_train_samples = 60000
print("Downloading train data")
train = try_download(url_train_image, url_train_labels, num_train_samples)
# URLs for the test image and label data
url_test_image = 'http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz'
url_test_labels = 'http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz'
num_test_samples = 10000
print("Downloading test data")
test = try_download(url_test_image, url_test_labels, num_test_samples)
```
## Visualize the data
```
# Plot a random image
sample_number = 5001
plt.imshow(train[sample_number,:-1].reshape(28,28), cmap="gray_r")
plt.axis('off')
print("Image Label: ", train[sample_number,-1])
```
## Save the images
Save the images in a local directory. While saving the data we flatten the images to a vector (28x28 image pixels becomes an array of length 784 data points).

The labels are encoded as [1-hot]( https://en.wikipedia.org/wiki/One-hot) encoding (label of 3 with 10 digits becomes `0001000000`, where the first index corresponds to digit `0` and the last one corresponds to digit `9`.

```
# Save the data files into a format compatible with CNTK text reader
def savetxt(filename, ndarray):
dir = os.path.dirname(filename)
if not os.path.exists(dir):
os.makedirs(dir)
if not os.path.isfile(filename):
print("Saving", filename )
with open(filename, 'w') as f:
labels = list(map(' '.join, np.eye(10, dtype=np.uint).astype(str)))
for row in ndarray:
row_str = row.astype(str)
label_str = labels[row[-1]]
feature_str = ' '.join(row_str[:-1])
f.write('|labels {} |features {}\n'.format(label_str, feature_str))
else:
print("File already exists", filename)
# Save the train and test files (prefer our default path for the data)
data_dir = os.path.join("..", "Examples", "Image", "DataSets", "MNIST")
if not os.path.exists(data_dir):
data_dir = os.path.join("data", "MNIST")
print ('Writing train text file...')
savetxt(os.path.join(data_dir, "Train-28x28_cntk_text.txt"), train)
print ('Writing test text file...')
savetxt(os.path.join(data_dir, "Test-28x28_cntk_text.txt"), test)
print('Done')
```
**Suggested Explorations**
One can do data manipulations to improve the performance of a machine learning system. We suggest you first use the data generated in this tutorial and run the classifier in subsequent parts of the CNTK 103 tutorial series. Once you have a baseline with classifying the data in its original form, you can use the different data manipulation techniques to further improve the model.
There are several ways to alter (transform) the data using CNTK readers. However, to get a feel for how these transforms can impact training and test accuracies, we strongly encourage individuals to try different transformation options.
- Shuffle the training data (change the ordering of the rows). Hint: Use `permute_indices = np.random.permutation(train.shape[0])`. Then, run Part B of the tutorial with this newly permuted data.
- Adding noise to the data can often improve the [generalization error](https://en.wikipedia.org/wiki/Generalization_error). You can augment the training set by adding noise (generated with numpy, hint: use `numpy.random`) to the training images.
- Distort the images with [affine transformation](https://en.wikipedia.org/wiki/Affine_transformation) (translations or rotations).
| github_jupyter |
```
sc
spark = SparkSession.builder.appName('summerRain').getOrCreate()
from io import StringIO
import itertools
import numpy as np
import pandas as pd
from pandas import Series
from pandas import concat
from pandas.plotting import lag_plot
from pandas.plotting import autocorrelation_plot
import statsmodels.api as sm
from statsmodels.graphics.tsaplots import plot_acf
from statsmodels.tsa.ar_model import AR
from sklearn.metrics import mean_squared_error
import matplotlib.pyplot as plt
%matplotlib inline
fig = plt.figure(figsize=(40,10))
plt.style.use('fivethirtyeight')
warnings.filterwarnings("ignore")
%%time
def readFiles (feature):
def process(tup):
def convertTuple(tup):
str = ''.join(tup)
return str
# Driver code
str = convertTuple(tup[0:2])
df = pd.read_csv(StringIO(str), header=None)
# slicing the output and storing filename in "filename dataframe"
filename = df[0:1]
filename = filename.iloc[0][0].replace('txtncols 180','')
filename = filename[-9:-1]
# storing the content of file in "content dataframe"
content = df[6:]
counter = 0
allValues = 0
# iterate through all the rows and columns of the file - better than for loop which took 40 mins
modPandasDF=content.iloc[:,0].str.split(' ',expand=True).replace("-9999",float('NaN'))
modPandasDF=modPandasDF.astype('float')
modPandasDF=modPandasDF.values
xDF=np.nanmean(modPandasDF)
mean=np.nanmean(xDF)
return filename, mean
# daily grid data for rainfall
if(feature=="rainfall" or feature=="maximum-temperature"):
path = "hdfs:/data/HCP053/climate/gridded-land-obs-daily/grid/"+feature+"/*"
elif(feature=="sunshine" or feature=="snow-falling"):
path = "hdfs:/data/HCP053/climate/gridded-land-obs-monthly/"+feature+"/*"
# Get rdd containing one record for each file.
files_rdd = sc.wholeTextFiles(path, minPartitions=20)
print(type(files_rdd))
print('Number of records (months):', files_rdd.count())
print('Number of partitions:', files_rdd.getNumPartitions())
# map lines to n_words
records = files_rdd.map(lambda n : process(n))
# collect the RDD to a list
llist = records.collect()
# two arrays
values = []
months = []
# store the filename in months array and its corrosponding value(mean) in values array
for line in llist:
values.append(line[1])
months.append(line[0])
# converting arrays to dataframes
valuesdf = pd.DataFrame({'DailyMean':values[:]})
yearmonthdf = pd.DataFrame({'YearMonth':months[:]})
# merging two dataframes into one
finaldf = pd.merge(yearmonthdf, valuesdf, left_index=True, right_index=True)
# sort the index
finaldf = finaldf.sort_values('YearMonth')
# converting first column to integer format
finaldf['YearMonth'] = finaldf['YearMonth'].astype('int')
# converting first column from integer to datetime format
finaldf['YearMonth'] = pd.to_datetime(finaldf['YearMonth'].astype(str), format='%Y%m%d')
# extracting year and month
finaldf['Year'] = finaldf['YearMonth'].dt.year
finaldf['Month'] = finaldf['YearMonth'].dt.month
print(finaldf.head())
return finaldf
%%time
finaldf = readFiles("rainfall")
finaldf.plot(x = 'YearMonth', y = 'DailyMean')
plt.title('Daily Rainfall in UK (1910 - 2016)')
plt.xlabel('Date')
plt.ylabel('Rainfall (mm)')
plt.xticks(rotation='vertical')
plt.show()
# Observation from plot: just plotting daily rainfall vs years makes no sense as it is not possible to observe any trends
z=pd.DataFrame(finaldf['YearMonth'])
z.insert(1,'DailyMean',finaldf['DailyMean'])
z.to_csv('SGD1.csv', index=False)
# BEST LINE FIT to look for any trend in data if present
y_values = finaldf['DailyMean']
#create a set of intervals equal to the number of dates
x_values = np.linspace(0,1,len(finaldf.loc[:, "DailyMean"]))
poly_degree = 3
coeffs = np.polyfit(x_values, y_values, poly_degree)
poly_eqn = np.poly1d(coeffs)
y_hat = poly_eqn(x_values)
fig = plt.figure(figsize=(40,10))
plt.xlabel('Year-Month')
plt.title('Daily Rainfall in the UK (1910 - 2016)')
plt.ylabel('Rainfall (mm)')
plt.xticks(rotation='vertical')
plt.plot(finaldf.loc[:, "YearMonth"], finaldf.loc[:,"DailyMean"], "ro")
plt.plot(finaldf.loc[:, "YearMonth"], y_hat)
y=pd.DataFrame(y_hat)
y.insert(1,'Year',finaldf['YearMonth'])
y.to_csv('SGD2.csv', index=False)
# it can be observed from the plot, the slope of best line fit is almost zero entirely
# hence it is not ideal to consider amount of rainfall as the deciding factor to prove summers will get drier
finaldf.head()
# creating a backup(tempdf) of finaldf
tempdf = finaldf
tempdf.head()
# plotting rainfall on monthly basis
rainfalldf = tempdf.loc[tempdf['Month'].isin(['1','2','3','4','5','6','7','8','9','10','11','12'])]
plt.xlabel('Months')
plt.title('Monthly Rainfall in UK (1960 - 2016)')
plt.ylabel('Rainfall (mm)')
rainfalldf.groupby('Month')['DailyMean'].mean().plot.bar()
# we can observe from the plot below, the rainfall during summer is lower than other seasons
# plotting rainfall for months of summer (Jun=6, July=7, Aug=8)
rainfalldf = tempdf.loc[tempdf['Month'].isin(['6','7','8'])]
plt.xlabel('Months')
plt.title('Rainfall during Summer in UK (1960 - 2016)')
plt.ylabel('Rainfall (mm)')
rainfalldf.groupby('Month')['DailyMean'].mean().plot.bar()
# dataframe to hold number of days having rainfall less than 1 mm
daysbelowthresholdrainfall = rainfalldf.loc[(rainfalldf['DailyMean'] <= 1)]
print(daysbelowthresholdrainfall.dtypes)
print(daysbelowthresholdrainfall.tail())
# number of days having rainfall less than 1 mm during summer grouped by year
daysbelowthresholdrainfall['DailyMean'].groupby([daysbelowthresholdrainfall.Year, daysbelowthresholdrainfall.Month]).agg('count')
daysbelowthresholdrainfall.plot(x = 'YearMonth', y = 'DailyMean')
plt.title('Rainfall below 1 mm in a day during summer (1960 - 2016)')
plt.xlabel('Date')
plt.ylabel('Rainfall (mm)')
plt.xticks(rotation='vertical')
# Observations: the plot shows days having rainfall recorded less than 1mm, but does not show a specific increasing or decreasing trend
# To make the plot more sensible we do the next operations
# BEST LINE FIT - the slope of this plot is better (increasing/decreasing) than the previous one
y_values = daysbelowthresholdrainfall['DailyMean']
#create a set of intervals equal to the number of dates
x_values = np.linspace(0,1,len(daysbelowthresholdrainfall.loc[:, "DailyMean"]))
poly_degree = 3
coeffs = np.polyfit(x_values, y_values, poly_degree)
poly_eqn = np.poly1d(coeffs)
y_hat = poly_eqn(x_values)
fig = plt.figure(figsize=(40,10))
plt.xlabel('Year-Month')
plt.title('Rainfall below 1 mm in a day (1960 - 2016)')
plt.ylabel('Rainfall (mm)')
plt.xticks(rotation='vertical')
plt.plot(daysbelowthresholdrainfall.loc[:, "YearMonth"], daysbelowthresholdrainfall.loc[:,"DailyMean"], "ro")
plt.plot(daysbelowthresholdrainfall.loc[:, "YearMonth"], y_hat)
# dropping redundant columns that are not required for prediction stage
daysbelowthresholdrainfall = daysbelowthresholdrainfall.drop(['Year', 'Month'], axis=1)
daysbelowthresholdrainfall.dtypes
# Count the number of days in a year having rainfall < 1 mm
count = daysbelowthresholdrainfall['YearMonth'].groupby([daysbelowthresholdrainfall.YearMonth.dt.year]).agg({'count'}).reset_index()
print(count.dtypes)
# Create a new dataframe with year and count as two columns
newdf = pd.DataFrame(count)
newdf.columns = ['Year','Number_of_days']
newdf.dtypes
newdf.plot(x = 'Year', y = 'Number_of_days')
plt.title('Rainfall below 1 mm in a day (1960 - 2016)')
plt.xlabel('Date')
plt.ylabel('Number_of_days')
plt.xticks(rotation='vertical')
z=pd.DataFrame(newdf['Year'])
z.insert(1,'Data',newdf['Number_of_days'])
z.to_csv('SGD3.csv', index=False)
# BEST LINE FIT - to observe the rate of increment in trend of the above plot
y_values = newdf['Number_of_days']
#create a set of intervals equal to the number of dates
x_values = np.linspace(0,1,len(newdf.loc[:, "Number_of_days"]))
poly_degree = 3
coeffs = np.polyfit(x_values, y_values, poly_degree)
poly_eqn = np.poly1d(coeffs)
y_hat = poly_eqn(x_values)
fig = plt.figure(figsize=(40,10))
plt.xlabel('Year')
plt.title('Rainfall below 1 mm in a day (1960 - 2016)')
plt.ylabel('Rainfall (mm)')
plt.xticks(rotation='vertical')
plt.plot(newdf.loc[:, "Year"], newdf.loc[:,"Number_of_days"], "ro")
plt.plot(newdf.loc[:, "Year"], y_hat)
z=pd.DataFrame(y_hat)
z.insert(1,'Year',newdf['Year'])
z.to_csv('SGD4.csv', index=False)
newdf = newdf.astype({"Number_of_days": float})
newdf.to_csv('RainfallBelow1.csv', index=False)
df = pd.read_csv('RainfallBelow1.csv', index_col='Year')
df.index = pd.to_datetime(df.index, format='%Y')
print(df.head())
print(df.dtypes)
p = d = q = range(0, 2)
pdq = list(itertools.product(p, d, q))
seasonal_pdq = [(x[0], x[1], x[2], 12) for x in list(itertools.product(p, d, q))]
print('Examples of parameter combinations for Seasonal ARIMA...')
print('SARIMAX: {} x {}'.format(pdq[1], seasonal_pdq[1]))
print('SARIMAX: {} x {}'.format(pdq[1], seasonal_pdq[2]))
print('SARIMAX: {} x {}'.format(pdq[2], seasonal_pdq[3]))
print('SARIMAX: {} x {}'.format(pdq[2], seasonal_pdq[4]))
# goal here is to use a “grid search” to find the optimal set of parameters(p, d, q) that yields the best performance for our model.
for param in pdq:
for param_seasonal in seasonal_pdq:
mod = sm.tsa.statespace.SARIMAX(df,
order=param,
seasonal_order=param_seasonal,
enforce_stationarity=False,
enforce_invertibility=False)
results = mod.fit()
print('ARIMA{}x{}12 - AIC:{}'.format(param, param_seasonal, results.aic))
# ARIMA(1, 1, 1)x(1, 1, 0, 12)12 - AIC:245.13442269741506
# the above AIC (Akaike Information Critera) value is the lowest of all, so we should consider its corrosponding values as the optimal
# fitting the arima model
mod = sm.tsa.statespace.SARIMAX(df,
order=(1, 1, 1),
seasonal_order=(1, 1, 0, 12),
enforce_stationarity=False,
enforce_invertibility=False)
results = mod.fit()
print(results.summary().tables[1])
# model diagnostics to investigate any unusual behavior
results.plot_diagnostics(figsize=(16, 8))
plt.show()
# validating forecasts from 2010-01-01 to the end date 2016-01-01
pred = results.get_prediction(start=pd.to_datetime('2010-01-01'), dynamic=False)
pred_ci = pred.conf_int()
ax = df['1960':].plot(label='Observed')
pred.predicted_mean.plot(ax=ax, label='One-step ahead Forecast', alpha=.7, figsize=(14, 7))
ax.fill_between(pred_ci.index,
pred_ci.iloc[:, 0],
pred_ci.iloc[:, 1], color='k', alpha=.2)
plt.title('Training from 1960 to 2016, Testing from 2010 to 2016')
ax.set_xlabel('Year')
ax.set_ylabel('Number of Days')
plt.legend()
plt.show()
df['1960':]
# observations: the predicted plot(orange) is almost similar to the expected plot(blue)
# validating forecasts from 2010-01-01 to the end date 2016-01-01
pred = results.get_prediction(start=pd.to_datetime('2010-01-01'), dynamic=False)
pred_ci = pred.conf_int()
ax = df['1960':'2010'].plot(label='Observed')
pred.predicted_mean.plot(ax=ax, label='One-step ahead Forecast', alpha=.7, figsize=(14, 7))
ax.fill_between(pred_ci.index,
pred_ci.iloc[:, 0],
pred_ci.iloc[:, 1], color='k', alpha=.2)
plt.title('Training from 1960 to 2009, Testing from 2010 to 2016')
ax.set_xlabel('Year')
ax.set_ylabel('Number of Days')
plt.legend()
plt.show()
df['1960':'2010']
# forecasting data for future
pred_uc = results.get_forecast(steps=50)
pred_ci = pred_uc.conf_int()
ax = df.plot(label='Observed', figsize=(14, 7))
pred_uc.predicted_mean.plot(ax=ax, label='Forecast')
ax.fill_between(pred_ci.index,
pred_ci.iloc[:, 0],
pred_ci.iloc[:, 1], color='k', alpha=.25)
plt.title('Number of Days having rainfall < 1 mm during Summer')
ax.set_xlabel('Year')
ax.set_ylabel('Number of Days')
plt.legend()
plt.show()
pred_uc.predicted_mean
# observations: the prediction model seems to follow the learned trend and keeps on decreasing.
# This contradicts our stated hypothesis as we were expecting number of days to increase
# Extract the predicted and true values of our time series
y_forecasted = pred_uc.predicted_mean
y_truth = df['2010-01-01':]
# Compute the mean square error
mse = ((y_forecasted - y_truth) ** 2).mean()
print('The Mean Squared Error of our forecasts is {}'.format(round(mse, 2)))
# the results are NaN - why?
```
| github_jupyter |
# データサイエンス100本ノック(構造化データ加工編) - R
## はじめに
- 初めに以下のセルを実行してください
- 必要なライブラリのインポートとデータベース(PostgreSQL)からのデータ読み込みを行います
- 利用が想定されるライブラリは以下セルでインポートしています
- その他利用したいライブラリがあればinstall.packages()で適宜インストールしてください
- 名前、住所等はダミーデータであり、実在するものではありません
```
require('RPostgreSQL')
require('tidyr')
require('dplyr')
require('stringr')
require('caret')
require('lubridate')
require('rsample')
require('recipes')
require('themis')
host <- 'db'
port <- Sys.getenv()["PG_PORT"]
dbname <- Sys.getenv()["PG_DATABASE"]
user <- Sys.getenv()["PG_USER"]
password <- Sys.getenv()["PG_PASSWORD"]
con <- dbConnect(PostgreSQL(), host=host, port=port, dbname=dbname, user=user, password=password)
df_customer <- dbGetQuery(con,"SELECT * FROM customer")
df_category <- dbGetQuery(con,"SELECT * FROM category")
df_product <- dbGetQuery(con,"SELECT * FROM product")
df_receipt <- dbGetQuery(con,"SELECT * FROM receipt")
df_store <- dbGetQuery(con,"SELECT * FROM store")
df_geocode <- dbGetQuery(con,"SELECT * FROM geocode")
```
# 演習問題
---
> R-001: レシート明細データフレーム(df_receipt)から全項目の先頭10件を表示し、どのようなデータを保有しているか目視で確認せよ。
---
> R-002: レシート明細データフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、10件表示させよ。
---
> R-003: レシート明細データフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、10件表示させよ。ただし、sales_ymdはsales_dateに項目名を変更しながら抽出すること。
---
> R-004: レシート明細データフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
> - 顧客ID(customer_id)が"CS018205000001"
---
> R-005: レシート明細データフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
> - 顧客ID(customer_id)が"CS018205000001"
> - 売上金額(amount)が1,000以上
---
> R-006: レシート明細データフレーム(receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上数量(quantity)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
> - 顧客ID(customer_id)が"CS018205000001"
> - 売上金額(amount)が1,000以上または売上数量(quantity)が5以上
---
> R-007: レシート明細のデータフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
> - 顧客ID(customer_id)が"CS018205000001"
> - 売上金額(amount)が1,000以上2,000以下
---
> R-008: レシート明細データフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
> - 顧客ID(customer_id)が"CS018205000001"
> - 商品コード(product_cd)が"P071401019"以外
---
> R-009: 以下の処理において、出力結果を変えずにORをANDに書き換えよ。
`
df_store %>%
filter(!(prefecture_cd == "13" | floor_area > 900))
`
---
> R-010: 店舗データフレーム(df_store)から、店舗コード(store_cd)が"S14"で始まるものだけ全項目抽出し、10件だけ表示せよ。
---
> R-011: 顧客データフレーム(df_customer)から顧客ID(customer_id)の末尾が1のものだけ全項目抽出し、10件だけ表示せよ。
---
> R-012: 店舗データフレーム(df_store)から横浜市の店舗だけ全項目表示せよ。
---
> R-013: 顧客データフレーム(df_customer)から、ステータスコード(status_cd)の先頭がアルファベットのA〜Fで始まるデータを全項目抽出し、10件だけ表示せよ。
---
> R-014: 顧客データフレーム(df_customer)から、ステータスコード(status_cd)の末尾が数字の1〜9で終わるデータを全項目抽出し、10件だけ表示せよ。
---
> R-015: 顧客データフレーム(df_customer)から、ステータスコード(status_cd)の先頭がアルファベットのA〜Fで始まり、末尾が数字の1〜9で終わるデータを全項目抽出し、10件だけ表示せよ。
---
> R-016: 店舗データフレーム(df_store)から、電話番号(tel_no)が3桁-3桁-4桁のデータを全項目表示せよ。
---
> R-017: 顧客データフレーム(df_customer)を生年月日(birth_day)で高齢順にソートし、先頭10件を全項目表示せよ。
---
> R-018: 顧客データフレーム(df_customer)を生年月日(birth_day)で若い順にソートし、先頭10件を全項目表示せよ。
---
> R-019: レシート明細データフレーム(df_receipt)に対し、1件あたりの売上金額(amount)が高い順にランクを付与し、先頭10件を抽出せよ。項目は顧客ID(customer_id)、売上金額(amount)、付与したランクを表示させること。なお、売上金額(amount)が等しい場合は同一順位を付与するものとする。
---
> R-020: レシート明細データフレーム(df_receipt)に対し、1件あたりの売上金額(amount)が高い順にランクを付与し、先頭10件を抽出せよ。項目は顧客ID(customer_id)、売上金額(amount)、付与したランクを表示させること。なお、売上金額(amount)が等しい場合でも別順位を付与すること。
---
> R-021: レシート明細データフレーム(df_receipt)に対し、件数をカウントせよ。
---
> R-022: レシート明細データフレーム(df_receipt)の顧客ID(customer_id)に対し、ユニーク件数をカウントせよ。
---
> R-023: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)と売上数量(quantity)を合計せよ。
---
> R-024: レシート明細データフレーム(df_receipt)に対し、顧客ID(customer_id)ごとに最も新しい売上日(sales_ymd)を求め、10件表示せよ。
---
> R-025: レシート明細データフレーム(df_receipt)に対し、顧客ID(customer_id)ごとに最も古い売上日(sales_ymd)を求め、10件表示せよ。
---
> R-026: レシート明細データフレーム(df_receipt)に対し、顧客ID(customer_id)ごとに最も新しい売上日(sales_ymd)と古い売上日を求め、両者が異なるデータを10件表示せよ。
---
> R-027: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の平均を計算し、降順でTOP5を表示せよ。
---
> R-028: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の中央値を計算し、降順でTOP5を表示せよ。
---
> R-029: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに商品コード(product_cd)の最頻値を求めよ。
---
> R-030: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の標本分散を計算し、降順でTOP5を表示せよ。
---
> R-031: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の標本標準偏差を計算し、降順でTOP5を表示せよ。
---
> R-032: レシート明細データフレーム(df_receipt)の売上金額(amount)について、25%刻みでパーセンタイル値を求めよ。
---
> R-033: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の平均を計算し、330以上のものを抽出せよ。
---
> R-034: レシート明細データフレーム(df_receipt)に対し、顧客ID(customer_id)ごとに売上金額(amount)を合計して全顧客の平均を求めよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。
---
> R-035: レシート明細データフレーム(df_receipt)に対し、顧客ID(customer_id)ごとに売上金額(amount)を合計して全顧客の平均を求め、平均以上に買い物をしている顧客を抽出せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。なお、データは10件だけ表示させれば良い。
---
> R-036: レシート明細データフレーム(df_receipt)と店舗データフレーム(df_store)を内部結合し、レシート明細データフレームの全項目と店舗データフレームの店舗名(store_name)を10件表示させよ。
---
> R-037: 商品データフレーム(df_product)とカテゴリデータフレーム(df_category)を内部結合し、商品データフレームの全項目とカテゴリデータフレームの小区分名(category_small_name)を10件表示させよ。
---
> R-038: 顧客データフレーム(df_customer)とレシート明細データフレーム(df_receipt)から、各顧客ごとの売上金額合計を求めよ。ただし、売上実績がない顧客については売上金額を0として表示させること。また、顧客は性別コード(gender_cd)が女性(1)であるものを対象とし、非会員(顧客IDが"Z"から始まるもの)は除外すること。なお、結果は10件だけ表示させれば良い。
---
> R-039: レシート明細データフレーム(df_receipt)から売上日数の多い顧客の上位20件と、売上金額合計の多い顧客の上位20件を抽出し、完全外部結合せよ。ただし、非会員(顧客IDが"Z"から始まるもの)は除外すること。
---
> R-040: 全ての店舗と全ての商品を組み合わせると何件のデータとなるか調査したい。店舗(df_store)と商品(df_product)を直積した件数を計算せよ。
---
> R-041: レシート明細データフレーム(df_receipt)の売上金額(amount)を日付(sales_ymd)ごとに集計し、前日からの売上金額増減を計算せよ。なお、計算結果は10件表示すればよい。
---
> R-042: レシート明細データフレーム(df_receipt)の売上金額(amount)を日付(sales_ymd)ごとに集計し、各日付のデータに対し、1日前、2日前、3日前のデータを結合せよ。結果は10件表示すればよい。
---
> R-043: レシート明細データフレーム(df_receipt)と顧客データフレーム(df_customer)を結合し、性別(gender)と年代(ageから計算)ごとに売上金額(amount)を合計した売上サマリデータフレーム(df_sales_summary)を作成せよ。性別は0が男性、1が女性、9が不明を表すものとする。
>
> ただし、項目構成は年代、女性の売上金額、男性の売上金額、性別不明の売上金額の4項目とすること(縦に年代、横に性別のクロス集計)。また、年代は10歳ごとの階級とすること。
---
> R-044: 前設問で作成した売上サマリデータフレーム(df_sales_summary)は性別の売上を横持ちさせたものであった。このデータフレームから性別を縦持ちさせ、年代、性別コード、売上金額の3項目に変換せよ。ただし、性別コードは男性を"00"、女性を"01"、不明を"99"とする。
---
> R-045: 顧客データフレーム(df_customer)の生年月日(birth_day)は日付型でデータを保有している。これをYYYYMMDD形式の文字列に変換し、顧客ID(customer_id)とともに抽出せよ。データは10件を抽出すれば良い。
---
> R-046: 顧客データフレーム(df_customer)の申し込み日(application_date)はYYYYMMDD形式の文字列型でデータを保有している。これを日付型に変換し、顧客ID(customer_id)とともに抽出せよ。データは10件を抽出すれば良い。
---
> R-047: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)はYYYYMMDD形式の数値型でデータを保有している。これを日付型に変換し、レシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。データは10件を抽出すれば良い。
---
> R-048: レシート明細データフレーム(df_receipt)の売上エポック秒(sales_epoch)は数値型のUNIX秒でデータを保有している。これを日付型に変換し、レシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。データは10件を抽出すれば良い。
---
> R-049: レシート明細データフレーム(df_receipt)の売上エポック秒(sales_epoch)を日付型に変換し、「年」だけ取り出してレシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。データは10件を抽出すれば良い。
---
> R-050: レシート明細データフレーム(df_receipt)の売上エポック秒(sales_epoch)を日付型に変換し、「月」だけ取り出してレシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。なお、「月」は0埋め2桁で取り出すこと。データは10件を抽出すれば良い。
---
> R-051: レシート明細データフレーム(df_receipt)の売上エポック秒を日付型に変換し、「日」だけ取り出してレシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。なお、「日」は0埋め2桁で取り出すこと。データは10件を抽出すれば良い。
---
> R-052: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計の上、売上金額合計に対して2,000円以下を0、2,000円より大きい金額を1に2値化し、顧客ID、売上金額合計とともに10件表示せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。
---
> R-053: 顧客データフレーム(df_customer)の郵便番号(postal_cd)に対し、東京(先頭3桁が100〜209のもの)を1、それ以外のものを0に2値化せよ。さらにレシート明細データフレーム(df_receipt)と結合し、全期間において売上実績がある顧客数を、作成した2値ごとにカウントせよ。
---
> R-054: 顧客データフレーム(df_customer)の住所(address)は、埼玉県、千葉県、東京都、神奈川県のいずれかとなっている。都道府県毎にコード値を作成し、顧客ID、住所とともに抽出せよ。値は埼玉県を11、千葉県を12、東京都を13、神奈川県を14とすること。結果は10件表示させれば良い。
---
> R-055: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、その合計金額の四分位点を求めよ。その上で、顧客ごとの売上金額合計に対して以下の基準でカテゴリ値を作成し、顧客ID、売上金額合計とともに表示せよ。カテゴリ値は上から順に1〜4とする。結果は10件表示させれば良い。
>
> - 最小値以上第一四分位未満
> - 第一四分位以上第二四分位未満
> - 第二四分位以上第三四分位未満
> - 第三四分位以上
---
> R-056: 顧客データフレーム(df_customer)の年齢(age)をもとに10歳刻みで年代を算出し、顧客ID(customer_id)、生年月日(birth_day)とともに抽出せよ。ただし、60歳以上は全て60歳代とすること。年代を表すカテゴリ名は任意とする。先頭10件を表示させればよい。
---
> R-057: 前問題の抽出結果と性別(gender)を組み合わせ、新たに性別×年代の組み合わせを表すカテゴリデータを作成せよ。組み合わせを表すカテゴリの値は任意とする。先頭10件を表示させればよい。
---
> R-058: 顧客データフレーム(df_customer)の性別コード(gender_cd)をダミー変数化し、顧客ID(customer_id)とともに抽出せよ。結果は10件表示させれば良い。
---
> R-059: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を平均0、標準偏差1に標準化して顧客ID、売上金額合計とともに表示せよ。標準化に使用する標準偏差は、不偏標準偏差と標本標準偏差のどちらでも良いものとする。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。結果は10件表示させれば良い。
---
> R-060: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を最小値0、最大値1に正規化して顧客ID、売上金額合計とともに表示せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。結果は10件表示させれば良い。
---
> R-061: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を常用対数化(底=10)して顧客ID、売上金額合計とともに表示せよ(ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること)。結果は10件表示させれば良い。
---
> R-062: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を自然対数化(底=e)して顧客ID、売上金額合計とともに表示せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。結果は10件表示させれば良い。
---
> R-063: 商品データフレーム(df_product)の単価(unit_price)と原価(unit_cost)から、各商品の利益額を算出せよ。結果は10件表示させれば良い。
---
> R-064: 商品データフレーム(df_product)の単価(unit_price)と原価(unit_cost)から、各商品の利益率の全体平均を算出せよ。 ただし、単価と原価にはNULLが存在することに注意せよ。
---
> R-065: 商品データフレーム(df_product)の各商品について、利益率が30%となる新たな単価を求めよ。ただし、1円未満は切り捨てること。そして結果を10件表示させ、利益率がおよそ30%付近であることを確認せよ。ただし、単価(unit_price)と原価(unit_cost)にはNULLが存在することに注意せよ。
---
> R-066: 商品データフレーム(df_product)の各商品について、利益率が30%となる新たな単価を求めよ。今回は、1円未満を丸めること(四捨五入または偶数への丸めで良い)。そして結果を10件表示させ、利益率がおよそ30%付近であることを確認せよ。ただし、単価(unit_price)と原価(unit_cost)にはNULLが存在することに注意せよ。
---
> R-067: 商品データフレーム(df_product)の各商品について、利益率が30%となる新たな単価を求めよ。今回は、1円未満を切り上げること。そして結果を10件表示させ、利益率がおよそ30%付近であることを確認せよ。ただし、単価(unit_price)と原価(unit_cost)にはNULLが存在することに注意せよ。
---
> R-068: 商品データフレーム(df_product)の各商品について、消費税率10%の税込み金額を求めよ。1円未満の端数は切り捨てとし、結果は10件表示すれば良い。ただし、単価(unit_price)にはNULLが存在することに注意せよ。
---
> R-069: レシート明細データフレーム(df_receipt)と商品データフレーム(df_product)を結合し、顧客毎に全商品の売上金額合計と、カテゴリ大区分(category_major_cd)が"07"(瓶詰缶詰)の売上金額合計を計算の上、両者の比率を求めよ。抽出対象はカテゴリ大区分"07"(瓶詰缶詰)の売上実績がある顧客のみとし、結果は10件表示させればよい。
---
> R-070: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)に対し、顧客データフレーム(df_customer)の会員申込日(application_date)からの経過日数を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。
---
> R-071: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)に対し、顧客データフレーム(df_customer)の会員申込日(application_date)からの経過月数を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。1ヶ月未満は切り捨てること。
---
> R-072: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)に対し、顧客データフレーム(df_customer)の会員申込日(application_date)からの経過年数を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。1年未満は切り捨てること。
---
> R-073: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)に対し、顧客データフレーム(df_customer)の会員申込日(application_date)からのエポック秒による経過時間を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。なお、時間情報は保有していないため各日付は0時0分0秒を表すものとする。
---
> R-074: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)に対し、当該週の月曜日からの経過日数を計算し、顧客ID、売上日、当該週の月曜日付とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値でデータを保持している点に注意)。
---
> R-075: 顧客データフレーム(df_customer)からランダムに1%のデータを抽出し、先頭から10件データを抽出せよ。
---
> R-076: 顧客データフレーム(df_customer)から性別(gender_cd)の割合に基づきランダムに10%のデータを層化抽出し、性別ごとに件数を集計せよ。
---
> R-077: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客単位に合計し、合計した売上金額の外れ値を抽出せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。なお、ここでは外れ値を平均から3σ以上離れたものとする。結果は10件表示させれば良い。
---
> R-078: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客単位に合計し、合計した売上金額の外れ値を抽出せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。なお、ここでは外れ値を第一四分位と第三四分位の差であるIQRを用いて、「第一四分位数-1.5×IQR」よりも下回るもの、または「第三四分位数+1.5×IQR」を超えるものとする。結果は10件表示させれば良い。
---
> R-079: 商品データフレーム(df_product)の各項目に対し、欠損数を確認せよ。
---
> R-080: 商品データフレーム(df_product)のいずれかの項目に欠損が発生しているレコードを全て削除した新たなdf_product_1を作成せよ。なお、削除前後の件数を表示させ、前設問で確認した件数だけ減少していることも確認すること。
---
> R-081: 単価(unit_price)と原価(unit_cost)の欠損値について、それぞれの平均値で補完した新たなdf_product_2を作成せよ。なお、平均値については1円未満を丸めること(四捨五入または偶数への丸めで良い)。補完実施後、各項目について欠損が生じていないことも確認すること。
---
> R-082: 単価(unit_price)と原価(unit_cost)の欠損値について、それぞれの中央値で補完した新たなdf_product_3を作成せよ。なお、中央値については1円未満を丸めること(四捨五入または偶数への丸めで良い)。補完実施後、各項目について欠損が生じていないことも確認すること。
---
> R-083: 単価(unit_price)と原価(unit_cost)の欠損値について、各商品の小区分(category_small_cd)ごとに算出した中央値で補完した新たなdf_product_4を作成せよ。なお、中央値については1円未満を丸めること(四捨五入または偶数への丸めで良い)。補完実施後、各項目について欠損が生じていないことも確認すること。
---
> R-084: 顧客データフレーム(df_customer)の全顧客に対し、全期間の売上金額に占める2019年売上金額の割合を計算せよ。ただし、売上実績がない場合は0として扱うこと。そして計算した割合が0超のものを抽出せよ。 結果は10件表示させれば良い。また、作成したデータにNAやNANが存在しないことを確認せよ。
---
> R-085: 郵便番号(postal_cd)を用いて経度緯度変換用データフレーム(df_geocode)を紐付け、新たなdf_customer_1を作成せよ。ただし、複数紐づく場合は経度(longitude)、緯度(latitude)それぞれ平均を算出すること。
---
> R-086: 前設問で作成した緯度経度つき顧客データフレーム(df_customer_1)に対し、申込み店舗コード(application_store_cd)をキーに店舗データフレーム(df_store)と結合せよ。そして申込み店舗の緯度(latitude)・経度情報(longitude)と顧客の緯度・経度を用いて距離(km)を求め、顧客ID(customer_id)、顧客住所(address)、店舗住所(address)とともに表示せよ。計算式は簡易式で良いものとするが、その他精度の高い方式を利用したライブラリを利用してもかまわない。結果は10件表示すれば良い。
$$
緯度(ラジアン):\phi \\
経度(ラジアン):\lambda \\
距離L = 6371 * arccos(sin \phi_1 * sin \phi_2
+ cos \phi_1 * cos \phi_2 * cos(\lambda_1 − \lambda_2))
$$
---
> R-087: 顧客データフレーム(df_customer)では、異なる店舗での申込みなどにより同一顧客が複数登録されている。名前(customer_name)と郵便番号(postal_cd)が同じ顧客は同一顧客とみなし、1顧客1レコードとなるように名寄せした名寄顧客データフレーム(df_customer_u)を作成せよ。ただし、同一顧客に対しては売上金額合計が最も高いものを残すものとし、売上金額合計が同一もしくは売上実績がない顧客については顧客ID(customer_id)の番号が小さいものを残すこととする。
---
> R-088: 前設問で作成したデータを元に、顧客データフレームに統合名寄IDを付与したデータフレーム(df_customer_n)を作成せよ。ただし、統合名寄IDは以下の仕様で付与するものとする。
> - 重複していない顧客:顧客ID(customer_id)を設定
> - 重複している顧客:前設問で抽出したレコードの顧客IDを設定
---
> R-089: 売上実績がある顧客に対し、予測モデル構築のため学習用データとテスト用データに分割したい。それぞれ8:2の割合でランダムにデータを分割せよ。
---
> R-090: レシート明細データフレーム(df_receipt)は2017年1月1日〜2019年10月31日までのデータを有している。売上金額(amount)を月次で集計し、学習用に12ヶ月、テスト用に6ヶ月のモデル構築用データを3セット作成せよ。
---
> R-091: 顧客データフレーム(df_customer)の各顧客に対し、売上実績がある顧客数と売上実績がない顧客数が1:1となるようにアンダーサンプリングで抽出せよ。
---
> R-092: 顧客データフレーム(df_customer)では、性別に関する情報が非正規化の状態で保持されている。これを第三正規化せよ。
---
> R-093: 商品データフレーム(df_product)では各カテゴリのコード値だけを保有し、カテゴリ名は保有していない。カテゴリデータフレーム(df_category)と組み合わせて非正規化し、カテゴリ名を保有した新たな商品データフレームを作成せよ。
---
> R-094: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。なお、出力先のパスはdata配下とする。
> - ファイル形式はCSV(カンマ区切り)
> - ヘッダ有り
> - 文字コードはUTF-8
---
> R-095: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。なお、出力先のパスはdata配下とする。
> - ファイル形式はCSV(カンマ区切り)
> - ヘッダ有り
> - 文字コードはCP932
---
> R-096: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。なお、出力先のパスはdata
配下とする。
> - ファイル形式はCSV(カンマ区切り)
> - ヘッダ無し
> - 文字コードはUTF-8
---
> R-097: 先に作成した以下形式のファイルを読み込み、データフレームを作成せよ。また、先頭3件を表示させ、正しくとりまれていることを確認せよ。
> - ファイル形式はCSV(カンマ区切り)
> - ヘッダ有り
> - 文字コードはUTF-8
---
> R-098: 先に作成した以下形式のファイルを読み込み、データフレームを作成せよ。また、先頭3件を表示させ、正しくとりまれていることを確認せよ。
> - ファイル形式はCSV(カンマ区切り)
> - ヘッダ無し
> - 文字コードはUTF-8
---
> R-099: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。なお、出力先のパスはdata配下とする。
> - ファイル形式はTSV(タブ区切り)
> - ヘッダ有り
> - 文字コードはUTF-8
---
> R-100: 先に作成した以下形式のファイルを読み込み、データフレームを作成せよ。また、先頭3件を表示させ、正しくとりまれていることを確認せよ。
> - ファイル形式はTSV(タブ区切り)
> - ヘッダ有り
> - 文字コードはUTF-8
# これで100本終わりです。おつかれさまでした!
| github_jupyter |
<a href="https://colab.research.google.com/github/Kristina140699/Practice_100Codes/blob/main/Python100Codes/Part_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **100 Codes in Python Programing Part 2**
# **This file contains 26-50 codes**
# Code 26 of 100
Displaying the highest common factor (H.C.F) or greatest common divisor (G.C.D) of two numbers which is the largest positive integer that perfectly divides the two given numbers.
```
def hcfFn(x, y):
if x > y:
smaller = y
else:
smaller = x
for i in range(1, smaller+1):
if((x % i == 0) and (y % i == 0)):
hcf = i
return hcf
num1 = int(input("Enter the first number: "))
num2 = int(input("Enter the second number: "))
print("The H.C.F. is", hcfFn(num1, num2))
```
# Code 27 of 100
#Computing the LCM-least common multiple of two numbers
```
def lcmFn(x, y):
if x > y:
greater = x
else:
greater = y
while(True):
if((greater % x == 0) and (greater % y == 0)):
lcm = greater
break
greater += 1
return lcm
num1 = int(input("Enter the first number: "))
num2 = int(input("Enter the second number: "))
print("The L.C.M. is", lcmFn(num1, num2))
```
#Note:
There is a convension that if we know the **HCF** then we can easily find out the **LCM** of any two given numbers.
**Formula:** Number1 X Number2 = L.C.M. X G.C.D.
#Code 28 of 100
# Computing LCM Using HCF
```
def hcfFn(x, y):
while(y):
x, y = y, x % y
print("\nThe HCF of the given numbers is: ",x)
return x
def compute_lcm(x, y):
lcm = (x*y)//hcfFn(x,y)
print("The product of the given numbers is: ",x*y)
return lcm
num1 = int(input("Enter the first number: "))
num2 = int(input("Enter the second number: "))
print("The L.C.M. is", compute_lcm(num1, num2))
```
#Code 29 of 100
#Shuffling Deck of Cards using random module.
Shuffling a deck of cards has been the evergreen game since forever
but how do we do it in python?
Well that can be very easily done using the random module so let's have a look at the following code
```
import itertools, random
deck = list(itertools.product(range(1,14),['Spade','Heart','Diamond','Club']))
random.shuffle(deck)
print("You got:")
for i in range(5):
print(deck[i][0], "of", deck[i][1])
```
#Code 30 of 100
#Printing a calender
```
import calendar
yy = int(input("Enter the year: "))
mm = int(input("Enter the month: "))
print("\n")
c = calendar.TextCalendar(calendar.SUNDAY) #Sunday is the first day in the calendar
st = c.formatmonth(yy, mm, 0, 0) #(year, month, horizontal space, vertical space)
print(st)
```
# Code 31 of 100
Let us suppose that I want to have meetUps with my team on the first Friday of every month but i want to figure out when is the first friday of every month!!
This can be done with the following program:
```
import calendar
yy = int(input("Enter the year: "))
print("\n Team meetings will be on: ")
for m in range(1, 13):
cal = calendar.monthcalendar(yy, m)
weekone = cal[0]
weektwo = cal[1]
if weekone[calendar.FRIDAY] != 0:
meetday = weekone[calendar.FRIDAY]
else:
meetday = weektwo[calendar.FRIDAY]
print("%2d of %s" % ( meetday, calendar.month_name[m]))
```
#Code 32 of 100
#Analysing the datetime module of python
```
#this program shows my current date-time ie. at the time when I ran this program, when you run this code it will show you your current date-time.
from datetime import date
from datetime import time
from datetime import datetime
today= date.today()
print("Today's date is :", today)
print("Date components: ", today.day,":",today.month,":",today.year)
print("Today's weekday number is: ", today.weekday())
days = ["monday","tuesday","wednesday","thursday","friday","saturday","sunday"]
print("Which is a", days[today.weekday()])
today = datetime.now()
print("The current date and time is ", today)
t = datetime.time(datetime.now())
print("The current time is", t)
```
#Code 33 of 100
#Date-Time module
```
from datetime import datetime
n = datetime.now()
#### Date Formatting ####
# %y/%Y - Year, %a/%A - weekday, %b/%B - month, %d - day of month
print(n.strftime("The current year is: %Y"))
print(n.strftime("The date today is %d %B %Y" ))
print(n.strftime("%A %d %B %Y"))
# %c - locale's date and time, %x - locale's date, %X - locale's time
print(n.strftime("Locale date and time: %c")) # %C make no sense!
print(n.strftime("Locale date: %x"))
print(n.strftime("Locale time: %X"))
#### Time Formatting ####
# %I/%H - 12/24 Hour, %M - minute, %S - second, %p - locale's AM/PM
print(n.strftime("The current time is: %I:%M:%S %p"))
print(n.strftime("The current time is: %H:%M"))
```
#Code 34 of 100
#timedelta module
```
from datetime import date
from datetime import time
from datetime import datetime
from datetime import timedelta
now = datetime.now()
print ("Today is: " + str(now))
#using timedelta to print the date
print ("\nOne year from now it will be: " + str(now + timedelta(days=365)))
#timedelta with more than one argument
print ("\nIn two weeks and 3 days it will be: " + str(now + timedelta(weeks=2, days=3)))
#using strftime to format string and calculate the date 1 week ago
t = datetime.now() - timedelta(weeks=1)
s = t.strftime("%A %B %d, %Y")
print ("\nA week ago it was " + s)
#Example printing: How many days until April Fools' Day?
today = date.today() #get today's date
afd = date(today.year, 4, 1) #get April Fool's date for the same year
#use date comparison to see if April Fool's has already gone for this year
#if it has, use the replace() function to get the date for next year
if afd < today:
print ("\nApril Fool's day already went by %d days ago" % ((today-afd).days))
afd = afd.replace(year=today.year + 1) # if so, get the date for next year
# Now calculate the amount of time until April Fool's Day
time_to_afd = afd - today
print ("It's just", time_to_afd.days, "days until next April Fools' Day!")
```
# **Using Recursion**
#Code 35 of 100
#Displaying Fibonacci Sequence Using Recursion
```
def recur_fibo(n):
if n <= 1:
return n
else:
return(recur_fibo(n-1) + recur_fibo(n-2))
nterms = int(input("Enter the number of terms you want to see in the series: "))
if nterms <= 0:
print("Plese enter a positive integer")
else:
print("\nFibonacci sequence:")
for i in range(nterms):
print(recur_fibo(i), end=" ")
```
#Code 36 of 100
# Factorial of a number using recursion
```
def recur_factorial(n):
if n == 1:
return n
else:
print(n, end=" x ")
return n*recur_factorial(n-1)
num = int(input("Enter the number whose factorial is to be printed: "))
print("The factorial of", num, "is", end=" ")
if num < 0:
print("Sorry, factorial does not exist for negative numbers")
elif num == 0:
print("The factorial of 0 is 1")
else:
print(1,"=", recur_factorial(num)) # 3 x 2 x 1 = 6
```
#Code 37 of 100
# Converting Decimal to Binary Using Recursion
# Converting Decimal to Octal Using Recursion
```
## Converting Decimal to Binary Using Recursion
def convertToBinary(n):
if n > 1:
convertToBinary(n//2)
print(n % 2,end = '')
dec = int(input("Enter a decimal number: "))
print("Binary equivalent of", dec, "is: ", end="")
convertToBinary(dec)
## Converting Decimal to Octal Using Recursion
def convertToOctal(n):
if n > 1:
convertToOctal(n//8)
print(n % 8,end = '')
dec = int(input("\n\nEnter a decimal number: "))
print("Octal equivalent of", dec, "is: ", end="")
convertToOctal(dec)
```
#Code 38 of 100
#Simple Calculator operations on two matrices
```
def two_d_matrix(m, n):
Outp = []
for i in range(m):
row = []
for j in range(n):
num = int(input(f"Index [{i}][{j}]: "))
row.append(num)
Outp.append(row)
return Outp
def sum(A, B):
output = []
print("\nSum of the matrix is :")
for i in range(len(A)):
row = []
for j in range(len(A[0])):
row.append(A[i][j] + B[i][j])
output.append(row)
return output
def minus(A, B):
output = []
print("\nDifference of the matrix is :")
for i in range(len(A)):
row = []
for j in range(len(A[0])):
row.append(A[i][j] + B[i][j])
output.append(row)
return output
def pro(A, B):
output = []
print("\nProduct of the matrix is :")
for i in range(len(A)):
row = []
for j in range(len(A[0])):
row.append(A[i][j] * B[i][j])
output.append(row)
return output
def div(A, B):
output = []
print("\nQuotient of the matrix is :")
for i in range(len(A)):
row = []
for j in range(len(A[0])):
row.append(A[i][j] * B[i][j])
output.append(row)
return output
m = int(input("Enter the value of Rows: "))
n = int(input("Enter the value of Columns: "))
print("\nThe matrices are of size: ", m, "x", n)
print("\nEnter values for the First matrix ")
A = two_d_matrix(m, n)
print("\nThe first matrix :")
print(A)
print("\nEnter values for the Second matrix ")
B = two_d_matrix(m, n)
print("\nThe second matrix: ")
print(B)
print("\n Select an operation: ")
print("1.Add")
print("2.Subtract")
print("3.Multiply")
print("4.Divide")
while True:
choice = input("Enter choice(1/2/3/4): ")
if choice in ('1', '2', '3', '4'):
if choice == '1':
s= sum(A, B)
print(s)
elif choice == '2':
print(minus(A, B))
elif choice == '3':
print(pro(A, B))
elif choice == '4':
print(div(A, B))
break
else:
print("Invalid Input")
```
#Code 39 of 100
#Transpose of a 3x3 Matrix using Nested Loop
```
def two_d_matrix(m, n):
Output = []
for i in range(m):
row = []
for j in range(n):
num = int(input(f"Index [{i}][{j}]: "))
row.append(num)
Output.append(row)
return Output
m = int(input("Enter the value of Rows: "))
n = int(input("Enter the value of Columns: "))
print("\nThe matrices are of size: ", m, "x", n)
print("\nEnter values for the Matrix ")
X = two_d_matrix(m, n)
print("\nThe first matrix :")
print(X)
result = [[0,0,0],
[0,0,0],
[0,0,0]]
print("\nThe transpose of the matrix: ")
for i in range(len(X)):
for j in range(len(X[0])):
result[j][i] = X[i][j]
for r in result:
print(r)
```
#Code 40 of 100
#Check if it is a palindrome in numeric form!
```
n=int(input("Enter number:"))
temp=n
rev=0
while(n>0):
dig=n%10
rev=rev*10+dig
n=n//10
if(temp==rev):
print("The number is a palindrome!")
else:
print("The number isn't a palindrome!")
```
#Code 41 of 100
#Check if it is a palindrome in String form!
```
def isPalindrome(n):
return n == n[::-1]
n=(input("Enter a String value: "))
ans = isPalindrome(n)
if ans:
print("The number is a palindrome!")
else:
print("The number isn't a palindrome!")
```
# **Let's Play with some Patterns!!**
# Code 42 of 100
# Printing Asterisk pyramid pattern Part 1
#Left Aligned
```
for num in range(8):
for i in range(num):
print ("*",end=" ") #print *
print("\r")
```
# Code 43 of 100
# Printing Asterisk pyramid pattern Part 2
# Center Aligned
```
def triangle(n):
k = n - 1
for i in range(0, n):
for j in range(0, k):
print(end=" ")
k = k - 1
for j in range(0, i+1):
print("* ", end="")
print("\r")
n = int(input("Enter the height of the pyramid: "))
print("\n")
triangle(n)
```
# Code 44 of 100
# Printing Asterisk pyramid pattern Part 3
# Right Aligned
```
def pypart2(n):
k = 2*n - 2
for i in range(0, n):
for j in range(0, k):
print(end=" ")
k = k - 2
for j in range(0, i+1):
print("* ", end="")
print("\r")
n = int(input("Enter the height of the pyramid: "))
print("\n")
pypart2(n)
```
# Code 45 of 100
# Printing Number pyramid pattern Part 1
# Horizontal
```
for num in range(6):
for i in range(num):
print (num,end=" ") #print number
print("\r")
```
# Code 46 of 100
# Printing Number pyramid pattern Part 2
# Vertical
```
num = 1
for i in range(0, 5):
num = 1
for j in range(0, i+1):
print(num, end=" ")
num = num + 1
print("\r")
```
# Code 47 of 100
# Printing Alphabet pyramid pattern Part 1
#Single Alphabet
```
for num in range(6):
for i in range(num):
print ("A",end=" ") #print number
print("\r")
```
# Code 48 of 100
# Printing Alphabet pyramid pattern Part 2
#Multi-Alphabet
```
num = 65
for i in range(0, 5):
for j in range(0, i+1):
ch = chr(num)
print(ch, end=" ")
num = num + 1
print("\r")
```
#Code 49 of 100
# Removing Punctuations From a String and Converting it into lower case.
```
punctuations = '''”!()-[]{};:'"\,<>./?@#$%^&*_~“'''
my_string = input("Enter some string: \n")
no_punct = ""
for char in my_string:
if char not in punctuations:
no_punct = no_punct + char
print("\nRemoving Punctuations: ")
print(no_punct)
print("\nPrinting in lower case: ")
print(my_string.lower())
```
#Code 50 of 100
#Sorting Words in Alphabetic Order
```
my_string = input("Enter a string: ")
words = [word.lower() for word in my_string.split()]
words.sort()
print("\nThe sorted words are:")
for word in words:
print(word)
```
| github_jupyter |

# Activity 4: Grapevines in a warming world
___
In the last lesson, you learned about pandas, dataframes, and seaborn. You learned that the harvest dates of grapevines in Europe have been recorded for centuries and you read the data in to Jupyter to analyze using `pandas`.
In the cell below, import `pandas` again, use the `pd.read_csv()` function to read the [data](https://github.com/DanChitwood/PlantsAndPython/blob/master/grape_harvest.csv) (`grape_harvest.csv`) in using the `pd.read_csv()` function, and make sure the data is ready to analyze by printing the outputs of `.head()`, `.tail()`, `.describe()`, and `.columns` functions in the cell below.
Remember to import `pandas`!
```
# Read in the grape_harvest.csv dataset here
# Use pandas functions to verify that the dataset has been read in correctly
# Print the outputs of the .head(), .tail(), .describe(), and .columns functions
# Remember to import pandas!
```
Now that your data is read in, let's use masking, data visualization, and line fitting to explore the relationship between grape harvest date and climate over the centuries.
___
## Masking
### Determining the earliest and latest harvest dates and where they occurred
We'll start off our exploration of grape harvest dates by figuring out when and where the earliest and latest harvest dates occurred.
The pandas dataframe you just created should have four columns, which are:
* **'year'**: the year the data was collected
* **'region'**: the region in Europe that the data was collected from
* **'harvest'**: the harvest date recorded. Harvest date is defined as number of days after August 31st. A negative number means the grapes were harvested before August 31st that year, and a positive number after.
* **'anomaly'**: the temperature anomaly. For a given year, this number represents how much colder (negative) or hotter (positive) Europe was compared to a long term reference value, in degrees Celsius
Below, print out statements answering the following questions using masking techniques that you have learned:
1) **Which year did the earliest harvest happen, which region did it occur in, and how early was the harvest?**
2) **Which year did the latest harvest happen, which region did it occur in, and how late was the harvest?**
**Hint**: Remember, a mask is a Boolean statement. But that Boolean statement can be combined with pandas functions, like .min() or .max(). Also remember that masking, the Boolean statement, and pandas functions can be combined with specific columns by name.
**Second hint**: After implementing your mask, you can append to it the .values() function within your print statement. This will allow you to print the values retrieved using your mask.
```
# Put your code here to print which year the earliest harvest occured,
# the region it occurred in, and how early the harvest was
# Put your code here to print which year the latest harvest occured,
# the region it occurred in, and how late the harvest was
```
### Finding median harvest dates in 50 year intervals
You want to know if the grape harvest date is changing, and if so, is it getting earlier or later?
You decide that you would like to know the median grape harvest date for the following 50 year intervals, as well as the median since the year 2000:
* 1800-1849
* 1850-1899
* 1900-1949
* 1950-1999
* 2000-2007
**For each of the above intervals, calculate the median grape harvest date. For each interval print out statements saying "The median harvest date for years (*insert interval here*) is: x."**
**Hint:** You can write Boolean statements for values within an interval as:
``` python
data["column"] >= value & data["column"] <= value
```
**Is the harvest date for grapes getting earlier or later?**
```
# Put your code here
```
____
## Visualization and correlation
Now that you understand a bit about the overall trends in the data, you want to examine other factors that might influence grape harvest date besides historical changes in climate.
You see that the data comes from many regions, all the way from sunny Spain to Germany. You wonder if these latitudinal differences would have any effect on the grape harvest dates.
**Make a boxplot comparing the distributions of grape harvest dates where the x-axis is "region" and the y-axis "harvest". The regions will be ordered by latitude, from the most southern to northern. This will allow you to assess visually if latitude is affecting grape harvest date.**
Your plot should:
1. include axis labels (use `matplotlib` functions that you have already learned about)
2. have a title (use `matplotlib` functions that you have already learned about)
3. keep the figure size and x ticks commands provided in the code below
4. import `seaborn`
5. finally, arrange the regions by latitude, from the most southern to the most northern. To do this, use the provided list `latitude_order`. Within the `seaborn` boxplot function, specify the `order` argument as follows: `order = latitude_order`. This will arrange the regions in your boxplot from the most southern to northern.
**Hint:** Notice that you can combine `matplotlib` and `seaborn`! You can call the `seaborn` boxplot function and apply the styles that you like, but use the `matplotlib` functions you already know to modify the title, axes labels, and other plot attributes. The best of both worlds!
```
# Import matplotlib and seaborn
# Remember the line of code to set matplotlib figures to inline in Jupyter
## DO NOT DELETE THE PROVIDED LINES OF CODE
## (they are included to make sure you get decent looking plots)
plt.figure(figsize=(15,4))
#a list of regions, from the most southern to northern
latitude_order = ['spain','maritime_alps','languedoc','various_south_east',
'gaillac_south_west','southern_rhone_valley','bordeaux',
'northern_rhone_valley','auvergne','savoie','northern_italy',
'beaujolais_maconnais','vendee_poitou_charente','switzerland_leman_lake',
'jura','high_loire_valley','low_loire_valley','burgundy','auxerre_avalon',
'champagne_2','southern_lorraine','alsace','northern_lorraine',
'germany','ile_de_france','champagne_1','luxembourg']
# Put your code here
plt.xticks(rotation=90) #Rotates x axis labels so that they are readable
```
**Based on your graph, do you believe that latitude affects the harvest date?** Explain your reasoning. If harvest date changes going south to north, how does this impact your analysis of the effect of history and climate change on harvest date? If harvest date is not affected by latitude, what are the implications for your analysis then?
```
# Provide your interpretation from your plot of the relationship between harvest date and latitude
```
### Looking for correlation in harvest date between Burgundy and Switzerland
You wonder if the grape harvest dates in different regions are correlated with each other. Two regions with some of the longest records of grape harvest dates are Burgundy ("burgundy") and Switzerland ("switzerland_leman_lake"). You would like to examine the correlation between the grape harvest dates at these two locations. But first, you need to make sure there is a matching grape harvest date for every year between these regions.
Your task below is to:
**1. Create two masked dataframes, one named "burgundy" with only "burgundy" data and the other named "switzerland", with only "switzerland_leman_lake" data.**
**2. Then, using the "merge" function code provided below, create a dataset where every year of recorded "burgundy" data has a matching year or recorded "switzerland" data. Be sure to inspect the column names**
**3. Using your new merged dataframe named "burgundy_switzerland", print out the earliest and latest year in your new merged data.**
```
# DO NOT DELETE THE PROVIDED CODE FOR MERGING DATAFRAMES
# Below this point, first create two masked datasets, "burgundy" and "switzerland", that contain only the data for
# "burgundy" and "switzerland_leman_lake" regions, respectively by using masks
# Put your code here for the "burgundy" and "switzerland" masked dataframes
burgandy =
switzerland =
# After creating your masked dataframes above, use the merge function below to create a single dataframe
# Check the column names of your new merged dataset. The "burgundy" data will have "x" after it and the
# "switzerland" data "y". "year" will have neither "x" nor "y" because it is shared between the dataframes
burgundy_switzerland = burgundy.merge(switzerland, on='year')
print(burgundy_switzerland.columns)
# Next, find the minimum and maximum years represented in this new merged dataset
# Below, print out statements of what the earliest and latest years are
# Put your code here for the minimum and maximum years in your merged "burgundy_switzerland" dataframe
```
Now that you have data that is matched, with exactly one "burgundy" and one "switzerland_leman_lake" observation for each year, you can examine if the grape harvest dates are correlated.
Using the seaborn `jointplot()` function (documentation [here](https://seaborn.pydata.org/generated/seaborn.jointplot.html)), make a scatterplot with distributions on the sides to look at this correaltion. **Make sure that your plot includes the regression line!**
```
### Put your code here
```
**Question**: Correlation is the relationship between two variables: for example, when one variable increases in value, the other increases as well (positive correlation) or when one variable increases in value the other decreases (negative correlation). Variability is how spread the data is or how clustered.
Do you think that harvest date between the two regions is correlated? Positive or negative? How variable is the data? Is the variability constant across harvest dates? What can you learn from your graph?
```
# Provide your interpretation of your plot here
```
____
## Graphing temperature anomaly vs year and harvest date vs temperature anomaly
You have been wondering this whole time: is the temperature increasing with time?
To find out the answer to this question, **below create a scatter plot with "anomaly" on the y-axis and "year" on the x-axis.** Make sure your plots have labels and a title.
**Remember:** If you have already imported in `seaborn`, if you use `matplotlib` functions, your plots will still have the style of `seaborn` and you can refer to dataframes and specific columns within `matplotlib` functions that you have already learned about! Use `matplotlib` functions and refer to your dataframe and columns by name and see what happens!
```
## DO NOT DELETE THE PROVIDED LINES OF CODE
## (they are included to make sure you get decent looking plots)
plt.figure(figsize=(15,4))
# Put your code here
```
You see that indeed, temperature is increasing with time. The classic "hockey stick" pattern. You want to see if the harvest date is impacted by the temperature anomaly. As above, **below create a scatterplot with "harvest date" on the y-axis and "anomaly" on the x-axis.** Make sure your plots have labels and a title.
**Hint**: you are plotting lots of data and it will be hard to see the underlying relationship because of overplotting. You can insert the `alpha` argument into the `scatter()` function to create transparency and see your data better. You can start with an alpha as low as 0.1 (e.g., `alpha=0.1`) and adjust it higher if you like.
```
## DO NOT DELETE THE PROVIDED LINES OF CODE
## (they are included to make sure you get decent looking plots)
plt.figure(figsize=(8,5))
# Put your code here
```
**Question:** Based on your graph, is there a correaltion between harvest date and temperature anamoly? If so, is it positive or negative? Do you think the relationship is linear or is curved, like your temperature anomaly vs. year graph?
```
# Write your thoughts here about the relationship between harvest date and temperature here
```
### Modeling harvest date as a function of temperature anomaly using `seaborn`
You wonder if "harvest" date can be modeled as a function of temperature "anomaly". You even suspect, after plotting "harvest" vs. "anomaly" above, if this relationship might be linear. You are amazed to learn that in fact others suggest this relationship is linear! In the [publication](https://www.clim-past.net/8/1403/2012/cp-8-1403-2012.pdf) where you got this data from, it suggests that others have found that *for every 1C rise in temperature, harvest date is on average 10 days earlier!*
You set out to create a linear model predicting "harvest" as a function of "anomaly" and realize that this is wasy to do using the `sns.lmplot()` function! Read the documentation for `sns.lmplot()` [here](https://seaborn.pydata.org/generated/seaborn.lmplot.html) and make a plot, with a linear fit displayed, of harvest date versus temperature anomaly.
```
# Make your sns.lmplot() here
```
There seems to be a lot of overplotting with too many points! In the cell below, use the `scatter=False` argument to remove the datapoints for us to see only the fitted line!
```
# Remove the datapoints from your plot in this cell
```
Look at your fitted line and try to estimate the slope. Remember, the slope is the unit change in $y$ values divided by the unit change in $x$ values. For each degree Celsius, by how many days does the harvest change approximately?
```
# Put your estimate of how many days the harvest changes per each degree of temperature anomaly in Celsius.
```
Remember in Activity 1, that you indexed the global temperature anomaly during your lifetime to see how much global temperatures have changed since you were born. As a rough estimate, let's see by how many days such a temperature increase could have affected the harvest of grapes.
In the cell below, the temperature anamoly values starting at year 1900 to 2020 are provided in a list called `temp_anomaly`. These data are from [NASA](https://climate.nasa.gov/vital-signs/global-temperature/). Execute the cell below to create the list.
**Remember:** This list starts at 1900 to 2020 and indexing in python starts at 0.
```
temp_anomaly = [-0.19,-0.23,-0.25,-0.28,-0.3,-0.33,-0.36,-0.37,-0.39,-0.4,-0.41,-0.38,
-0.35,-0.32,-0.31,-0.3,-0.29,-0.29,-0.29,-0.29,-0.27,-0.26,-0.25,-0.24,
-0.23,-0.22,-0.21,-0.2,-0.19,-0.19,-0.19,-0.19,-0.18,-0.17,-0.16,-0.14,
-0.11,-0.06,-0.01,0.03,0.06,0.09,0.11,0.1,0.07,0.04,0,-0.04,-0.07,-0.08,
-0.08,-0.07,-0.07,-0.07,-0.07,-0.06,-0.05,-0.04,-0.01,0.01,0.03,0.01,
-0.01,-0.03,-0.04,-0.05,-0.06,-0.05,-0.03,-0.02,0,0,0,0,0.01,0.02,0.04,
0.07,0.12,0.16,0.2,0.21,0.22,0.21,0.21,0.22,0.24,0.27,0.31,0.33,0.34,
0.33,0.33,0.34,0.34,0.37,0.4,0.43,0.45,0.48,0.51,0.53,0.55,0.59,0.61,
0.62,0.63,0.64,0.65,0.65,0.65,0.67,0.7,0.74,0.79,0.83,0.88,0.91,0.95,
0.98,1.01]
```
In the cell below, index the list from the year of your birth to 2020. Look at the anomaly difference from your birth to 2020 and roughly estimate by how much such a change in temperature would affect the harvest date of grapevines.
**Hint:** Remember that -1 can index the last element of a list.
```
# Put your answer here
```
Reexamining your graph of temperature anomaly versus year, does your result make sense? What does this say about the pace of climate change and implications for agriculture?
```
# Write your thoughts about the pace of climate change and its implications
```
That's all for this activity! Thank you for participating!
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Estimators
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/estimator"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/estimator.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/estimator.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/estimator.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
This document introduces `tf.estimator`—a high-level TensorFlow
API. Estimators encapsulate the following actions:
* training
* evaluation
* prediction
* export for serving
You may either use the pre-made Estimators we provide or write your
own custom Estimators. All Estimators—whether pre-made or custom—are
classes based on the `tf.estimator.Estimator` class.
For a quick example try [Estimator tutorials](../tutorials/estimator/linear.ipynb). For an overview of the API design, see the [white paper](https://arxiv.org/abs/1708.02637).
## Advantages
Similar to a `tf.keras.Model`, an `estimator` is a model-level abstraction. The `tf.estimator` provides some capabilities currently still under development for `tf.keras`. These are:
* Parameter server based training
* Full [TFX](http://tensorflow.org/tfx) integration.
## Estimators Capabilities
Estimators provide the following benefits:
* You can run Estimator-based models on a local host or on a distributed multi-server environment without changing your model. Furthermore, you can run Estimator-based models on CPUs, GPUs, or TPUs without recoding your model.
* Estimators provide a safe distributed training loop that controls how and when to:
* load data
* handle exceptions
* create checkpoint files and recover from failures
* save summaries for TensorBoard
When writing an application with Estimators, you must separate the data input
pipeline from the model. This separation simplifies experiments with
different data sets.
## Pre-made Estimators
Pre-made Estimators enable you to work at a much higher conceptual level than the base TensorFlow APIs. You no longer have to worry about creating the computational graph or sessions since Estimators handle all the "plumbing" for you. Furthermore, pre-made Estimators let you experiment with different model architectures by making only minimal code changes. `tf.estimator.DNNClassifier`, for example, is a pre-made Estimator class that trains classification models based on dense, feed-forward neural networks.
### Structure of a pre-made Estimators program
A TensorFlow program relying on a pre-made Estimator typically consists of the following four steps:
#### 1. Write one or more dataset importing functions.
For example, you might create one function to import the training set and another function to import the test set. Each dataset importing function must return two objects:
* a dictionary in which the keys are feature names and the values are Tensors (or SparseTensors) containing the corresponding feature data
* a Tensor containing one or more labels
For example, the following code illustrates the basic skeleton for an input function:
```
def input_fn(dataset):
... # manipulate dataset, extracting the feature dict and the label
return feature_dict, label
```
See [data guide](../../guide/data.md) for details.
#### 2. Define the feature columns.
Each `tf.feature_column` identifies a feature name, its type, and any input pre-processing. For example, the following snippet creates three feature columns that hold integer or floating-point data. The first two feature columns simply identify the feature's name and type. The third feature column also specifies a lambda the program will invoke to scale the raw data:
```
# Define three numeric feature columns.
population = tf.feature_column.numeric_column('population')
crime_rate = tf.feature_column.numeric_column('crime_rate')
median_education = tf.feature_column.numeric_column(
'median_education',
normalizer_fn=lambda x: x - global_education_mean)
```
For further information, see the [feature columns tutorial](https://www.tensorflow.org/tutorials/keras/feature_columns).
#### 3. Instantiate the relevant pre-made Estimator.
For example, here's a sample instantiation of a pre-made Estimator named `LinearClassifier`:
```
# Instantiate an estimator, passing the feature columns.
estimator = tf.estimator.LinearClassifier(
feature_columns=[population, crime_rate, median_education])
```
For further information, see the [linear classifier tutorial](https://www.tensorflow.org/tutorials/estimator/linear).
#### 4. Call a training, evaluation, or inference method.
For example, all Estimators provide a `train` method, which trains a model.
```
# `input_fn` is the function created in Step 1
estimator.train(input_fn=my_training_set, steps=2000)
```
You can see an example of this below.
### Benefits of pre-made Estimators
Pre-made Estimators encode best practices, providing the following benefits:
* Best practices for determining where different parts of the computational graph should run, implementing strategies on a single machine or on a
cluster.
* Best practices for event (summary) writing and universally useful
summaries.
If you don't use pre-made Estimators, you must implement the preceding features yourself.
## Custom Estimators
The heart of every Estimator—whether pre-made or custom—is its *model function*, which is a method that builds graphs for training, evaluation, and prediction. When you are using a pre-made Estimator, someone else has already implemented the model function. When relying on a custom Estimator, you must write the model function yourself.
## Recommended workflow
1. Assuming a suitable pre-made Estimator exists, use it to build your first model and use its results to establish a baseline.
2. Build and test your overall pipeline, including the integrity and reliability of your data with this pre-made Estimator.
3. If suitable alternative pre-made Estimators are available, run experiments to determine which pre-made Estimator produces the best results.
4. Possibly, further improve your model by building your own custom Estimator.
```
import tensorflow as tf
import tensorflow_datasets as tfds
```
## Create an Estimator from a Keras model
You can convert existing Keras models to Estimators with `tf.keras.estimator.model_to_estimator`. Doing so enables your Keras
model to access Estimator's strengths, such as distributed training.
Instantiate a Keras MobileNet V2 model and compile the model with the optimizer, loss, and metrics to train with:
```
keras_mobilenet_v2 = tf.keras.applications.MobileNetV2(
input_shape=(160, 160, 3), include_top=False)
keras_mobilenet_v2.trainable = False
estimator_model = tf.keras.Sequential([
keras_mobilenet_v2,
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(1)
])
# Compile the model
estimator_model.compile(
optimizer='adam',
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
metrics=['accuracy'])
```
Create an `Estimator` from the compiled Keras model. The initial model state of the Keras model is preserved in the created `Estimator`:
```
est_mobilenet_v2 = tf.keras.estimator.model_to_estimator(keras_model=estimator_model)
```
Treat the derived `Estimator` as you would with any other `Estimator`.
```
IMG_SIZE = 160 # All images will be resized to 160x160
def preprocess(image, label):
image = tf.cast(image, tf.float32)
image = (image/127.5) - 1
image = tf.image.resize(image, (IMG_SIZE, IMG_SIZE))
return image, label
def train_input_fn(batch_size):
data = tfds.load('cats_vs_dogs', as_supervised=True)
train_data = data['train']
train_data = train_data.map(preprocess).shuffle(500).batch(batch_size)
return train_data
```
To train, call Estimator's train function:
```
est_mobilenet_v2.train(input_fn=lambda: train_input_fn(32), steps=500)
```
Similarly, to evaluate, call the Estimator's evaluate function:
```
est_mobilenet_v2.evaluate(input_fn=lambda: train_input_fn(32), steps=10)
```
For more details, please refer to the documentation for `tf.keras.estimator.model_to_estimator`.
| github_jupyter |
Copyright (c) 2019 [윤기태]
https://github.com/yoonkt200/python-data-analysis
[MIT License](https://github.com/yoonkt200/python-data-analysis/blob/master/LICENSE.txt)
# (가제) 파이썬 데이터 분석
-----
# 3.1) 국내 프로야구 연봉 예측
### 바로가기
- [<Step1. 탐색> 프로야구 연봉 데이터 살펴보기](#<Step1.-탐색>-프로야구-연봉-데이터-살펴보기)
- [프로야구 연봉 데이터셋의 기본 정보]
- [회귀 분석에 사용할 피처 살펴보기]
- [<Step2. 예측> : 투수의 연봉 예측하기](#<Step2.-예측>-:-투수의-연봉-예측하기)
- [피처들의 단위 맞춰주기 : 피처 스케일링]
- [피처들의 단위 맞춰주기 : one-hot-encoding]
- [피처들의 상관관계 분석]
- [회귀 분석 적용하기]
- [<Step3. 평가> : 예측 모델 평가하기](#<Step3.-평가>-:-예측-모델-평가하기)
- [어떤 피처가 가장 영향력이 강한 피처일까]
- [예측 모델의 평가]
- [회귀 분석 예측 성능을 높이기 위한 방법 : 다중 공선성 확인]
- [믿을만한 피처로 다시 학습하기]
- [<Step4. 시각화> : 분석 결과의 시각화](#<Step4.-시각화>-:-분석-결과의-시각화)
- [예상 연봉과 실제 연봉 비교]
-----
```
# -*- coding: utf-8 -*-
%matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
```
# <Step1. 탐색> 프로야구 연봉 데이터 살펴보기
### [프로야구 연봉 데이터셋의 기본 정보]
```
# Data Source : http://www.statiz.co.kr/
picher_file_path = '../data/picher_stats_2017.csv'
batter_file_path = '../data/batter_stats_2017.csv'
picher = pd.read_csv(picher_file_path)
batter = pd.read_csv(batter_file_path)
picher.columns
picher.head()
print(picher.shape)
```
-----
### `[참고 - 한글을 사용하는 그래프]`
- 파이썬에서 matplotlib 기반의 시각화 툴을 이용할 때 한글 폰트를 사용하는 방법
- 자신의 개발환경에 설치된 폰트 확인
- `set(sorted([f.name for f in mpl.font_manager.fontManager.ttflist]))`
- 한글 폰트를 설정한 뒤, 축에 적용
- 예시 :`mpl.rc('font', family='08SeoulHangang')`
- 한글 폰트가 없다면, http://hangeul.naver.com/2017/nanum 에서 설치
```
import matplotlib as mpl
set(sorted([f.name for f in mpl.font_manager.fontManager.ttflist])) # 현재 OS 내에 설치된 폰트를 확인합니다.
mpl.rc('font', family='NanumGothicOTF') # 자신의 OS에 존재하는 한글 폰트를 선택합니다. 없는경우, 위의 링크에서 한글폰트 설치 후 실행합니다.
```
###### 예측할 대상인 '연봉'에 대한 정보
```
picher['연봉(2018)'].describe()
picher['연봉(2018)'].hist(bins=100) # 2018년 연봉 분포를 출력합니다.
picher.boxplot(column=['연봉(2018)']) # 연봉의 Boxplot을 출력합니다.
```
-----
### [회귀 분석에 사용할 피처 살펴보기]
```
picher_features_df = picher[['승', '패', '세', '홀드', '블론', '경기', '선발', '이닝', '삼진/9',
'볼넷/9', '홈런/9', 'BABIP', 'LOB%', 'ERA', 'RA9-WAR', 'FIP', 'kFIP', 'WAR',
'연봉(2018)', '연봉(2017)']]
# 피처 각각에 대한 histogram을 출력합니다.
def plot_hist_each_column(df):
plt.rcParams['figure.figsize'] = [20, 16]
fig = plt.figure(1)
# df의 column 갯수 만큼의 subplot을 출력합니다.
for i in range(len(df.columns)):
ax = fig.add_subplot(5, 5, i+1)
plt.hist(df[df.columns[i]], bins=50)
ax.set_title(df.columns[i])
plt.show()
plot_hist_each_column(picher_features_df)
```
-----
# <Step2. 예측> : 투수의 연봉 예측하기
### [피처들의 단위 맞춰주기 : 피처 스케일링]
```
# pandas 형태로 정의된 데이터를 출력할 때, scientific-notation이 아닌 float 모양으로 출력되게 해줍니다.
pd.options.mode.chained_assignment = None
# 피처 각각에 대한 scaling을 수행하는 함수를 정의합니다.
def standard_scaling(df, scale_columns):
for col in scale_columns:
series_mean = df[col].mean()
series_std = df[col].std()
df[col] = df[col].apply(lambda x: (x-series_mean)/series_std)
return df
# 피처 각각에 대한 scaling을 수행합니다.
scale_columns = ['승', '패', '세', '홀드', '블론', '경기', '선발', '이닝', '삼진/9',
'볼넷/9', '홈런/9', 'BABIP', 'LOB%', 'ERA', 'RA9-WAR', 'FIP', 'kFIP', 'WAR', '연봉(2017)']
picher_df = standard_scaling(picher, scale_columns)
picher_df = picher_df.rename(columns={'연봉(2018)': 'y'})
picher_df.head(5)
```
-----
### [피처들의 단위 맞춰주기 : one-hot-encoding]
```
# 팀명 피처를 one-hot encoding으로 변환합니다.
team_encoding = pd.get_dummies(picher_df['팀명'])
picher_df = picher_df.drop('팀명', axis=1)
picher_df = picher_df.join(team_encoding)
team_encoding.head(5)
picher_df.head()
```
-----
### [회귀 분석 적용하기]
##### 회귀 분석을 위한 학습, 테스트 데이터셋 분리
```
from sklearn import linear_model
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from math import sqrt
# 학습 데이터와 테스트 데이터로 분리합니다.
X = picher_df[picher_df.columns.difference(['선수명', 'y'])]
y = picher_df['y']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=19)
```
##### 회귀 분석 계수 학습 & 학습된 계수 출력
```
# 회귀 분석 계수를 학습합니다 (회귀 모델 학습)
lr = linear_model.LinearRegression()
model = lr.fit(X_train, y_train)
# 학습된 계수를 출력합니다.
print(lr.coef_)
picher_df.columns
```
-----
# <Step3. 평가> : 예측 모델 평가하기
### [어떤 피처가 가장 영향력이 강한 피처일까]
```
!pip install statsmodels
import statsmodels.api as sm
# statsmodel 라이브러리로 회귀 분석을 수행합니다.
X_train = sm.add_constant(X_train)
model = sm.OLS(y_train, X_train).fit()
model.summary()
# 한글 출력을 위한 사전 설정 단계입니다.
mpl.rc('font', family='AppleGothic')
plt.rcParams['figure.figsize'] = [20, 16]
# 회귀 계수를 리스트로 반환합니다.
coefs = model.params.tolist()
coefs_series = pd.Series(coefs)
# 변수명을 리스트로 반환합니다.
x_labels = model.params.index.tolist()
# 회귀 계수를 출력합니다.
ax = coefs_series.plot(kind='bar')
ax.set_title('feature_coef_graph')
ax.set_xlabel('x_features')
ax.set_ylabel('coef')
ax.set_xticklabels(x_labels)
```
-----
### [예측 모델의 평가]
```
# 학습 데이터와 테스트 데이터로 분리합니다.
X = picher_df[picher_df.columns.difference(['선수명', 'y'])]
y = picher_df['y']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=19)
# 회귀 분석 모델을 학습합니다.
lr = linear_model.LinearRegression()
model = lr.fit(X_train, y_train)
```
##### R2 score
```
# 회귀 분석 모델을 평가합니다.
print(model.score(X_train, y_train)) # train R2 score를 출력합니다.
print(model.score(X_test, y_test)) # test R2 score를 출력합니다.
```
##### RMSE score
```
# 회귀 분석 모델을 평가합니다.
y_predictions = lr.predict(X_train)
print(sqrt(mean_squared_error(y_train, y_predictions))) # train RMSE score를 출력합니다.
y_predictions = lr.predict(X_test)
print(sqrt(mean_squared_error(y_test, y_predictions))) # test RMSE score를 출력합니다.
```
-----
### `[미니 퀴즈 - 3.1]`
- `Train score, 그리고 Test score의 차이점은 무엇일까요? 그리고 어떤 점수가 더 높아야 할까요?`
- 훈련 데이터셋을 통해 계산한 점수와 테스트 데이터셋을 통해 계산한 점수간에는 어떤 차이가 있는 것인지 적어 봅시다. 그리고 이 두 점수의 차이가 크다면, 어떤 상황을 의미하는 것인지 생각해 봅시다.
- `대답 예시` : 훈련 데이터셋은 오로지 모델을 학습하고, 비용함수(오차함수)를 평가하는데 사용됩니다. 반면, 테스트 데이터셋은 모델에 영향을 미치지 않습니다. 결과를 예측하기 위한 입력데이터로만 활용됩니다. 가장 이상적인 경우는 Train score, Test score가 차이가 없는 것입니다. 이 차이가 벌어질수록 모델이 훈련 데이터셋에 과적합된 것입니다. 일반적으로는 Train score가 약간 더 높습니다.
-----
### [피처들의 상관관계 분석]
```
import seaborn as sns
# 피처간의 상관계수 행렬을 계산합니다.
corr = picher_df[scale_columns].corr(method='pearson')
show_cols = ['win', 'lose', 'save', 'hold', 'blon', 'match', 'start',
'inning', 'strike3', 'ball4', 'homerun', 'BABIP', 'LOB',
'ERA', 'RA9-WAR', 'FIP', 'kFIP', 'WAR', '2017']
# corr 행렬 히트맵을 시각화합니다.
plt.rc('font', family='NanumGothicOTF')
sns.set(font_scale=1.5)
hm = sns.heatmap(corr.values,
cbar=True,
annot=True,
square=True,
fmt='.2f',
annot_kws={'size': 15},
yticklabels=show_cols,
xticklabels=show_cols)
plt.tight_layout()
plt.show()
```
-----
### [회귀분석 예측 성능을 높이기 위한 방법 : 다중공선성 확인]
```
from statsmodels.stats.outliers_influence import variance_inflation_factor
# 피처마다의 VIF 계수를 출력합니다.
vif = pd.DataFrame()
vif["VIF Factor"] = [variance_inflation_factor(X.values, i) for i in range(X.shape[1])]
vif["features"] = X.columns
vif.round(1)
```
-----
### `[미니 퀴즈 - 3.2]`
- `적절한 피처를 선정하여 다시 학습해 보세요.`
- 사용할 피처를 다시 고른 뒤, 다시 학습을 진행했을 때의 Train score, Test score가 어떻게 향상되는지 살펴봅니다.
- 저자의 경우 아래의 과정을 거쳐 ['FIP', 'WAR', '볼넷/9', '삼진/9', '연봉(2017)'] 피처를 선정했을 때, 더욱 좋은 결과를 나타냈습니다.
- 선정 과정
- 1. VIF 계수가 높은 피처들을 우선적으로 제거합니다. 단, (FIP, kFIP)와 같이 유사한 두가지 피처중에는 하나만을 제거합니다.
- 2. 다시 공선성을 검증합니다. 이전 단계에서는 VIF 계수가 높았던 볼넷, 삼진 등의 피처의 VIF 계수가 낮아진 것을 확인할 수 있습니다. VIF 계수가 비정상적으로 높은 피처를 제거해주면, 다른 피처들의 공선성도 자연스레 감소하기 때문입니다.
- 3. 여전히 VIF 계수가 높은 피처들을 제거합니다.
- 4. 남은 피처를 토대로 다시한번 회귀분석을 실시합니다. 분석 결과에서 p-value가 유의하면서도 영향력이 큰 피처들을 선정합니다.
- train_score, test_score를 비교했을 때, 기존보다 overfit이 비교적 덜 된것으로 보입니다
- test rmse 역시 감소한 것을 볼 수 있습니다.
### [적절한 피처로 다시 학습하기]
```
# 피처를 재선정합니다.
X = picher_df[['FIP', 'WAR', '볼넷/9', '삼진/9', '연봉(2017)']]
y = picher_df['y']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=19)
# 모델을 학습합니다.
lr = linear_model.LinearRegression()
model = lr.fit(X_train, y_train)
# 결과를 출력합니다.
print(model.score(X_train, y_train)) # train R2 score를 출력합니다.
print(model.score(X_test, y_test)) # test R2 score를 출력합니다.
# 회귀 분석 모델을 평가합니다.
y_predictions = lr.predict(X_train)
print(sqrt(mean_squared_error(y_train, y_predictions))) # train RMSE score를 출력합니다.
y_predictions = lr.predict(X_test)
print(sqrt(mean_squared_error(y_test, y_predictions))) # test RMSE score를 출력합니다.
# 피처마다의 VIF 계수를 출력합니다.
X = picher_df[['FIP', 'WAR', '볼넷/9', '삼진/9', '연봉(2017)']]
vif = pd.DataFrame()
vif["VIF Factor"] = [variance_inflation_factor(X.values, i) for i in range(X.shape[1])]
vif["features"] = X.columns
vif.round(1)
```
-----
# <Step4. 시각화> : 분석 결과의 시각화
### [예상 연봉과 실제 연봉 비교]
```
# 2018년 연봉을 예측하여 데이터프레임의 column으로 생성합니다.
X = picher_df[['FIP', 'WAR', '볼넷/9', '삼진/9', '연봉(2017)']]
predict_2018_salary = lr.predict(X)
picher_df['예측연봉(2018)'] = pd.Series(predict_2018_salary)
# 원래의 데이터 프레임을 다시 로드합니다.
picher = pd.read_csv(picher_file_path)
picher = picher[['선수명', '연봉(2017)']]
# 원래의 데이터 프레임에 2018년 연봉 정보를 합칩니다.
result_df = picher_df.sort_values(by=['y'], ascending=False)
result_df.drop(['연봉(2017)'], axis=1, inplace=True, errors='ignore')
result_df = result_df.merge(picher, on=['선수명'], how='left')
result_df = result_df[['선수명', 'y', '예측연봉(2018)', '연봉(2017)']]
result_df.columns = ['선수명', '실제연봉(2018)', '예측연봉(2018)', '작년연봉(2017)']
# 재계약하여 연봉이 변화한 선수만을 대상으로 관찰합니다.
result_df = result_df[result_df['작년연봉(2017)'] != result_df['실제연봉(2018)']]
result_df = result_df.reset_index()
result_df = result_df.iloc[:10, :]
result_df.head(10)
# 선수별 연봉 정보(작년 연봉, 예측 연봉, 실제 연봉)를 bar 그래프로 출력합니다.
mpl.rc('font', family='NanumGothicOTF')
result_df.plot(x='선수명', y=['작년연봉(2017)', '예측연봉(2018)', '실제연봉(2018)'], kind="bar")
```
| github_jupyter |
# About: Notebooks for Hadoop Clusters README
Literate Computing for Reproducible Infrastructure: インフラ運用をJupyter + Ansibleでおこなう際のお手本Notebookです。(Hadoop版)
このリポジトリでは、HDP(Hortonworks Data Platform, https://jp.hortonworks.com/products/data-center/hdp/ )を利用してHadoopクラスタを構築し、運用するためのNotebook例を紹介しています。
**なお、これらのNotebookはNIIクラウドチーム内で行っている作業の考え方を示すためのもので、環境によってはそのままでは動作しないものもあります。**
----
[](http://creativecommons.org/licenses/by/4.0/)
Literate-computing-Hadoop (c) by National Institute of Informatics
Literate-computing-Hadoop is licensed under a
Creative Commons Attribution 4.0 International License.
You should have received a copy of the license along with this
work. If not, see <http://creativecommons.org/licenses/by/4.0/>.
## 関連資料
- [Jupyter notebook を用いた文芸的インフラ運用のススメ - SlideShare](http://www.slideshare.net/nobu758/jupyter-notebook-63167604)
- [Literate Automation(文芸的自動化)についての考察 - めもめも](http://enakai00.hatenablog.com/entry/2016/04/22/204125)
# お手本Notebook
お手本NotebookはこのNotebookと同じディレクトリにあります。Notebookは目的に応じて以下のような命名規則に則って名前がつけられています。
- D(NN)\_(Notebook名) ... インストール関連Notebook
- O(NN)\_(Notebook名) ... 運用関連Notebook
- T(NN)\_(Notebook名) ... テスト関連Notebook
特に、**[D00_Prerequisites for Literate Computing via Notebooks](D00_Prerequisites for Literate Computing via Notebooks.ipynb)は、お手本Notebookが適用可能なNotebook環境、Bind対象であるかどうかを確認するためのNotebook**です。はじめに実施して、これらのお手本Notebookが利用可能な状態かを確認してみてください。
## お手本Notebookの構成
このNotebookは、大きく分けて以下のような構成になっています。
1. 収容設計の準備Notebook
2. Hadoopマシン準備Notebook
3. HadoopインストールNotebook
4. Hadoop運用Notebook
5. Hadoop動作確認Notebook
これらのNotebookと、構築・運用する対象の関係は、以下のようになります。

*収容設計の準備Notebook* は、構築・運用対象とするシステム全体の収容設計をおこないます。ここで、どのようなマシンがあって(あるいは用意する必要があって)、それぞれのマシンにどのような役割を割り当てるかを明確化します。このプロセスによりAnsibleのInventory, 変数群を生成します。**ここで、Ansibleなど自動化ツールに与えるパラメータと、収容設計の関連を明確化**します。
*Hadoopマシン準備Notebook* は、Hadoopのインストール対象のマシンを準備し、**Hadoop(HDP)のインストールに必要な前提条件を満たすよう各マシンのOS設定を調整**します。それぞれの要件については各Notebookを参照してください。
*HadoopインストールNotebook* は、Hadoopおよびその周辺ツールのインストール手順を定義します。マシンの準備Notebookにより**前提条件が満たされた状態のマシンに対して各種ツールをインストールする**ことができます。
*Hadoop運用Notebook* は、構築した環境を保守する手順を定義します。実際にクラスタを構築すると、ハードウェア故障が生じたりさまざまなトラブルが発生するので、それらの状態変化に応じてクラスタの健康状態を維持する必要があります。**公開したこれらのNotebookは運用における手順や考え方を紹介する一例であり、障害の具体的な発生状況に応じて修正しながら実施する必要があります。**
*Hadoop動作確認Notebook* は、構築した環境に実際にデータをアップロードしてみたり、ジョブを実行してみたりする例を提供します。**HadoopインストールNotebookを通してインストールされた環境の使い方**を示す役割もあります。
### 収容設計の準備Notebook
Hadoopのように複数ノードからなるクラスタを構築・運用する際は、それぞれのノードにどのようなロール(役割)を割り振って、リソースを割り当てていくか(収容設計)が重要なポイントになります。収容設計を明確にしながらシステムを構築・運用していくことで、的確なシステムのスケールや、スムーズなトラブル対応をすることが可能になります。
そこで、インストールの前に以下のNotebookにより、収容設計の確認をしながら、AnsibleのInventoryやgroup_varsといった各種パラメータを生成しています。
- [D10_Hadoop - Set! Inventory](D10_Hadoop - Set! Inventory.ipynb)
ここで生成されるパラメータは、以降の手順のNotebookで設定ファイル生成の際に利用されます。
[](http://interactive.blockdiag.com/?compression=deflate&encoding=base64&src=eJyFk0FPhDAQhe_7K-peuLCoN5MVExMvXtRkvenGFDpAQ2lJO0tEs__dFkxoEOp5Zt57-V4mEyqvGacl-d4QUimDJslNR95MRVtIpUI47u0kOgBekEfZgUSl-3f5ZCeZUnU0TO-l4ZkAUmp1aj86qk1kJZDmNbB4LvUArVB9Y5UmGW__79KLoP3iUsUKszMcIflsRJwkiedK5rbPLWiKXMmAq7ezZjoxut0FsLijXkDKFKJNI2gGItVQgP6HJ9ndrfAcNRrFeNEfV7G7VGvwvBRxoQRz5Xgpg_W4WGHdk1ko2FtzCkuNjdclSMcebDCJhn9Ben0VKs6prdQ1i7Pg6SAFjyML6XJEHU2Jbn6pDboD9OFtCBmPtq9Ul4Dk0BuEZrsfRgv2bnDenH8AoFQymA)
ここで定義されるパラメータには、以下のようなものがあります。
- 各サービスに割り当てるリソース量の情報
- 各サービスのインストールに使用する外部リソースの情報
Hadoop構築Notebookでは、各マシンにどのサービスを載せて、どの程度のメモリやディスクを割り当てていくかは、 [hosts.csv](hosts.csv) のような表形式で定義しています。この情報に基づいて、AnsibleのPlaybookに与えるパラメータを定義しています。
また、Hadoopのインストール時にどのリポジトリからパッケージを取ってくるかという情報も、システムの素性を理解するうえで重要な情報です。このような情報も含めてNotebookの形で明確化、証跡を残すようにしています。
### Hadoopマシン準備Notebook
Hadoopをインストールする対象となるマシンの準備に関するNotebookです。
#### VMの作成手順
NIIクラウドチームでは、プライベートなベアメタルクラウド(仮想マシンではなく物理マシンを貸すIaaS)を持っています。このクラウド上でベアメタル(物理マシン)を複数台用意し、そこにHadoopをインストールする形を採っています。
ベアメタルの設定手順の事例紹介は難しいのですが、テスト用クラスタを構築する際に使っているKVM環境の準備と、VMの構築用Notebookをここでは紹介しています。
この例では、以下のものを準備しています。
- [D03_KVM - Ready! on CentOS](D03_KVM - Ready! on CentOS.ipynb)
- [D03b_KVM - Set! CentOS6](D03b_KVM - Set! CentOS6.ipynb)
- [D03c_KVM - Go! VM](D03c_KVM - Go! VM.ipynb)
また、Google Compute Engineでのマシン確保を想定したNotebookも添付しています。
- [D01_GCE - Set! Go! (Google Compute Engine)](D01_GCE - Set! Go! %28Google Compute Engine%29.ipynb)
- [O03_GCE - Destroy VM (Google Compute Engine)](O03_GCE - Destroy VM %28Google Compute Engine%29.ipynb)
#### VMの設定手順
作成したVMは、MinimalなOSがインストールされた状態ですので、(当然のことながら)セキュリティ上必要な設定を施す必要があります。
- [D90_Postscript - Operational Policy Settings; Security etc. (to be elaborated)](D90_Postscript - Operational Policy Settings; Security etc. %28to be elaborated%29.ipynb)
また、HDPをインストールするために必要な推奨設定などもあります。これらの設定を施すNotebookも紹介しています。
- [D11_Hadoop Prerequisites - Ready! on CentOS6](D11_Hadoop Prerequisites - Ready! on CentOS6.ipynb)
これらの設定としてどのような項目が必要かは、VMの初期設定により異なってきます。
### HadoopインストールNotebook
Hadoopのインストールは以下のような手順で実施しています。
HDPで提供されていないものについては、HDPインストール用Notebookとは別にNotebookを用意しています。
- [D12_Hadoop - Ready! on CentOS6 - ZK,HDFS,YARN,HBase,Hive,Spark](D12_Hadoop - Ready! on CentOS6 - ZK,HDFS,YARN,HBase,Hive,Spark.ipynb)
- [D13a_Hadoop Swimlanes - Ready! on Tez](D13a_Hadoop Swimlanes - Ready! on Tez.ipynb)
- [D13b_Hivemall - Ready! on Hive](D13b_Hivemall - Ready! on Hive.ipynb)
### Hadoop運用Notebook
Hadoopの運用に関しては、以下のようなNotebookを用意しています。
- [O12a_Hadoop - Start the services - ZK,HDFS,YARN,HBase,Spark](O12a_Hadoop - Start the services - ZK,HDFS,YARN,HBase,Spark.ipynb)
- [O12b_Hadoop - Stop the services - Spark,HBase,YARN,HDFS,ZK](O12b_Hadoop - Stop the services - Spark,HBase,YARN,HDFS,ZK.ipynb)
- [O12c_Hadoop - Decommission DataNode](O12c_Hadoop - Decommission DataNode.ipynb)
- [O12d_Hadoop - Restore a Slave Node](O12d_Hadoop - Restore a Slave Node.ipynb)
### Hadoop動作確認Notebook
動作確認のためのNotebookとして、以下のものを用意しています。
- [T12a_Hadoop - Confirm the services are alive - ZK,HDFS,YARN,HBase,Spark](T12a_Hadoop - Confirm the services are alive - ZK,HDFS,YARN,HBase,Spark.ipynb)
- [T12b_Hadoop - Simple YARN for Test job](T12b_Hadoop - Simple YARN job for Test.ipynb)
- [T12c_Hadoop - Simple HBase query for Test.ipynb](T12c_Hadoop - Simple HBase query for Test.ipynb)
- [T12d_Hadoop - Simple Spark script for Test](T12d_Hadoop - Simple Spark script for Test.ipynb)
- [T13b_Hadoop - Simple Hivemall query for Test](T13b_Hadoop - Simple Hivemall query for Test.ipynb)
## お手本Notebookの一覧
現在、このNotebook環境からアクセス可能なNotebookの一覧を参照するには、以下のセルを実行(`Run cell`)してください。Notebookファイルへのリンクが表示されます。
```
import re
import os
from IPython.core.display import HTML
ref_notebooks = filter(lambda m: m, map(lambda n: re.match(r'([A-Z][0-9][0-9a-z]+_.*)\.ipynb', n), os.listdir('.')))
ref_notebooks = sorted(ref_notebooks, key=lambda m: m.group(1))
HTML(''.join(map(lambda m: '<div><a href="{name}" target="_blank">{title}</a></div>'.format(name=m.group(0), title=m.group(1)),
ref_notebooks)))
```
## お手本Notebookと証跡Notebook
お手本Notebookを使う場合は、お手本をコピーし、そのコピーを開きます。このように、**お手本と作業証跡は明確に分けながら作業をおこないます。**
また、お手本をコピーする際は、 `YYYYMMDD_NN_` といった実施日を示すプレフィックスを付加することで、後で整理しやすくしています。
## 実際にお手本Notebookを使ってみる
以下のJavaScriptを実行することで、簡単にお手本から作業用Notebookを作成することもできます。
以下のセルを実行すると、Notebook名のドロップダウンリストと[作業開始]ボタンが現れます。
[作業開始]ボタンを押すと、お手本Notebookのコピーを作成した後、自動的にブラウザでコピーが開きます。
Notebookの説明を確認しながら実行、適宜修正しながら実行していってください。
```
from datetime import datetime
import shutil
def copy_ref_notebook(src):
prefix = datetime.now().strftime('%Y%m%d') + '_'
index = len(filter(lambda name: name.startswith(prefix), os.listdir('.'))) + 1
new_notebook = '{0}{1:0>2}_{2}'.format(prefix, index, src)
shutil.copyfile(src, new_notebook)
print(new_notebook)
frags = map(lambda m: '<option value="{name}">{title}</option>'.format(name=m.group(0), title=m.group(1)),
ref_notebooks)
HTML('''
<script type="text/Javascript">
function copy_otehon() {
var sel = document.getElementById('selector');
IPython.notebook.kernel.execute('copy_ref_notebook("' + sel.options[sel.selectedIndex].value + '")',
{'iopub': {'output': function(msg) {
window.open(msg.content.text, '_blank')
}}});
}
</script>
<select id="selector">''' + ''.join(frags) + '</select><button onclick="copy_otehon()">作業開始</button>')
```
## お手本のアーカイブ
以下のセルで、お手本NotebookのZIPアーカイブを作成できます。
```
ref_notebooks = filter(lambda m: m, map(lambda n: re.match(r'([A-Z][0-9][0-9a-z]+_.*)\.ipynb', n), os.listdir('.')))
ref_notebooks = sorted(ref_notebooks, key=lambda m: m.group(1))
!zip ref_notebooks-{datetime.now().strftime('%Y%m%d')}.zip README.ipynb hosts.csv {' '.join(map(lambda n: '"' + n.group(0) + '"', ref_notebooks))} scripts/* images/* group_vars/.gitkeep
```
こいつを・・・以下のURLからダウンロードできます。
```
HTML('<a href="../files/{filename}" target="_blank">{filename}</a>' \
.format(filename='ref_notebooks-' + datetime.now().strftime('%Y%m%d') + '.zip'))
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from sklearn import preprocessing
from sklearn.linear_model import RidgeCV
from sklearn.metrics import fbeta_score
from sklearn import linear_model
cancerFrame = pd.read_csv("../DataSet/CancerDataSet.txt", delimiter="\t")
cancerFrame.head()
```
# On garde pour X les variables contenues dans les colonnes 1 à -3
# On garde pour y les variables contenues dans -2
```
X = cancerFrame.iloc[:,1 :-3]
y = cancerFrame.iloc[:, -2]
```
## La regularisation permet d'éviter le sur apprentissage du modèle pour une regression linéaire
On ajoute une fonction objective (la somme des carrés des erreurs appellée baseline) un temre de regularisation qui mesure la complexité du modèle.
## Pour la regression Ridge la standardisation des valeurs est importante
```
X_reshape = np.array(X).reshape(-1, 1)
std_scaler = preprocessing.StandardScaler().fit(X)
X_std = std_scaler.transform(X)
y_reshape = np.array(y).reshape(-1, 1)
std_scaler = preprocessing.StandardScaler().fit(y_reshape)
y_std = std_scaler.transform(y_reshape)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_std, y, test_size=0.25)
lnr = linear_model.LinearRegression()
lnr.fit(X_train, y_train)
Xmean = np.mean(X_std)
Ymean = np.mean(y_std)
xycov = (X_std - Xmean) * (y_std-Ymean)
xvar = (X_std - Xmean)**2
beta = xycov.sum()/xvar.sum()
#betaWithNp = np.polyfit(X_reshape.reshape(1,-1), y_reshape, 1)[0]
print("beta", beta)
#print("betaWithNp", betaWithNp)
#erreur Quadratique au carré entre les données d'entrainement et de test
baseline_error = np.mean((lnr.predict(X_test)-y_test) **2)
baseline_error
# paramètre de regularisation
n_alphas = 100
alphas = np.logspace(-5, 5, n_alphas)
coefs = []
errors = []
ridge = linear_model.Ridge()
for a in alphas:
ridge.set_params(alpha=a)
ridge.fit(X_train, y_train)
errors.append(np.mean((ridge.predict(X_test)-y_test)**2))
coefs.append(ridge.coef_)
import matplotlib.pyplot as plt
ax = plt.gca()
ax.plot(alphas, errors, [10**-5, 10**5], [baseline_error, baseline_error])
ax.plot(baseline_error)
ax.set_xscale("log")
#index de l'erreur la plus petite
errorsIndex = np.argmin(errors)
#valeur de l'erreur la plus petite
errors[errorsIndex]
#coefficient alpha associé
alphas[errorsIndex]
## chemin de regularisation
chemin = plt.gca()
chemin.plot(alphas, coefs)
chemin.set_xscale("log")
#beta est en ordonnée et la solution de la regularisation est quand beta = 0
```
### Modèle Partimonieux pouvant mettre la ponderation de certaines varibles à zero
Lasso Least Absolute Shrinkage and Selection operator
```
lasso = linear_model.Lasso()
# paramètre de regularisation
n_alphas = 100
alphas = np.logspace(-5, 5, n_alphas)
coefs = []
errors = []
for a in alphas:
lasso.set_params(alpha=a)
lasso.fit(X_train, y_train)
coefs.append(lasso.coef_)
errors.append([baseline_error, (np.mean(lasso.predict(X_test) - y_test)**2)])
## chemin de regularisation
chemin = plt.gca()
chemin.plot(alphas, coefs)
chemin.set_xscale("log")
graph = plt.gca()
graph.plot(alphas, errors)
graph.set_xscale("log")
```
| github_jupyter |
# 3. Train-Predict Mix3model
## Result:
- Kaggle score:
## Tensorboard
- Input at command: tensorboard --logdir=./log
- Input at browser: http://127.0.0.1:6006
## Reference
- https://www.kaggle.com/codename007/a-very-extensive-landmark-exploratory-analysis
## Import PKGs
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
%matplotlib inline
from IPython.display import display
import os
import gc
import time
import zipfile
import pickle
import math
import pdb
import h5py
from PIL import Image
import shutil
from tqdm import tqdm
import multiprocessing
```
## Run name
```
project_name = 'Google_LandMark_Rec'
step_name = '3. Train-Predict_Mix3model'
time_str = time.strftime("%Y%m%d_%H%M%S", time.localtime())
run_name = project_name + '_' + step_name + '_' + time_str
print('run_name: ' + run_name)
```
## 项目文件夹
```
cwd = os.getcwd()
input_folder = os.path.join(cwd, 'input')
output_folder = os.path.join(cwd, 'output')
model_folder = os.path.join(cwd, 'model')
feature_folder = os.path.join(cwd, 'feature')
post_pca_feature_folder = os.path.join(cwd, 'post_pca_feature')
log_folder = os.path.join(cwd, 'log')
print('input_folder: \t\t\t' + input_folder)
print('output_folder: \t\t\t' + output_folder)
print('model_folder: \t\t\t' + model_folder)
print('feature_folder: \t\t' + feature_folder)
print('post_pca_feature_folder: \t' + post_pca_feature_folder)
print('log_folder: \t\t\t' + log_folder)
org_train_folder = os.path.join(input_folder, 'org_train')
org_test_folder = os.path.join(input_folder, 'org_test')
train_folder = os.path.join(input_folder, 'data_train')
test_folder = os.path.join(input_folder, 'data_test')
test_sub_folder = os.path.join(test_folder, 'test')
if not os.path.exists(post_pca_feature_folder):
os.mkdir(post_pca_feature_folder)
print('Create folder: %s' % post_pca_feature_folder)
train_csv_file = os.path.join(input_folder, 'train.csv')
test_csv_file = os.path.join(input_folder, 'test.csv')
sample_submission_folder = os.path.join(input_folder, 'sample_submission.csv')
```
## 获取landmark_id集合
```
train_csv = pd.read_csv(train_csv_file)
print('train_csv.shape is {0}.'.format(train_csv.shape))
display(train_csv.head(10))
test_csv = pd.read_csv(test_csv_file)
print('test_csv.shape is {0}.'.format(test_csv.shape))
display(test_csv.head(10))
train_id = train_csv['id']
train_landmark_id = train_csv['landmark_id']
unique_landmark_ids = list(set(train_landmark_id))
len_unique_landmark_ids = len(unique_landmark_ids)
print(unique_landmark_ids[:10]) # 确认landmark_id是从0开始
print('len(unique_landmark_ids)=%d' % len_unique_landmark_ids)
```
## 预览sample_submission.csv
```
sample_submission_csv = pd.read_csv(sample_submission_folder)
print('sample_submission_csv.shape is {0}.'.format(sample_submission_csv.shape))
display(sample_submission_csv.head(2))
```
## 加载feature
```
def load_h5(file_name):
print(file_name)
with h5py.File(file_name, 'r') as h:
x_train = np.array(np.array(h['train']))
y_train = np.array(h['train_labels'])
# x_val.append(np.array(h['val']))
# y_val = np.array(h['val_labels'])
x_test = np.array(np.array(h['test']))
return x_train, y_train, x_test
%%time
model_name = 'InceptionV3'
time_str = '200_20180312-050926'
feature_Xception = os.path.join(feature_folder, 'feature_%s_%s.h5' % (model_name, time_str))
x_train, y_train, x_test = load_h5(feature_Xception)
print(x_train.shape)
print(y_train.shape)
print(x_test.shape)
len_data = x_train.shape[0]
len_test = x_test.shape[0]
print('len_data: %s' % len_data)
print('len_test: %s' % len_test)
print(y_train[:10])
%%time
from sklearn.utils import shuffle
data_indeces = list(range(len_data))
print(data_indeces[:10])
data_indeces = shuffle(data_indeces)
print(data_indeces[:10])
from sklearn.model_selection import train_test_split
train_indeces, val_indeces = train_test_split(data_indeces[:30*10000], test_size=0.01, random_state=2018, shuffle=False)
print('len(train_indeces)=%s' % len(train_indeces))
print(train_indeces[:10])
print('len(val_indeces)=%s' % len(val_indeces))
print(val_indeces[:10])
def load_h5_data(feature_folder, file_reg, model_name, time_str):
x_data = {}
y_data = {}
feature_model = os.path.join(feature_folder, file_reg % (model_name, time_str))
with h5py.File(feature_model, 'r') as h:
x_data = np.array(h['train'])
y_data = np.array(h['train_labels'])
return x_data, y_data
def load_h5_test(feature_folder, file_reg, model_name, time_str):
x_test = {}
feature_model = os.path.join(feature_folder, file_reg % (model_name, time_str))
with h5py.File(feature_model, 'r') as h:
x_test = np.array(h['test'])
return x_test
def is_files_existed(feature_folder, file_reg, model_names, time_strs):
for model_name in model_names:
for time_str in time_strs:
file_name = file_reg % (model_name, time_str)
file_path = os.path.join(feature_folder, file_name)
if not os.path.exists(file_path):
print('File not existed: %s' % file_path)
return False
else:
print('File existed: %s' % file_path)
return True
# Test
file_reg = 'feature_%s_%s.h5'
model_names = [
# 'MobileNet',
# 'VGG16',
# 'VGG19',
# 'ResNet50',
# 'DenseNet121',
# 'DenseNet169',
# 'DenseNet201',
'Xception',
'InceptionV3',
'InceptionResNetV2'
]
time_strs = [
'200_20180312-050926',
'150_20180311-151108'
]
print(is_files_existed(feature_folder, file_reg, model_names, time_strs))
def time_str_generator(time_strs):
while(1):
for time_str in time_strs:
print(' ' + time_str)
yield time_str
# Test
time_str_gen = time_str_generator(time_strs)
for i in range(10):
next(time_str_gen)
%%time
from keras.utils.np_utils import to_categorical
from sklearn.utils import shuffle
def load_time_str_feature_data(data_indeces, feature_folder, file_reg, model_names, time_str):
x_data_time_strs = []
y_data_time_strs = None
for model_name in model_names:
x_data_time_str, y_data_time_str = load_h5_data(feature_folder, file_reg, model_name, time_str)
x_data_time_str, y_data_time_str = x_data_time_str[data_indeces], y_data_time_str[data_indeces]
# Around data to 3 decimals to calculate computation
# x_data_time_str = np.round(x_data_time_str, decimals=3) # 导致后面的全链接神经网络分类器不收敛
x_data_time_strs.append(x_data_time_str)
y_data_time_strs = y_data_time_str
x_data_time_strs = np.concatenate(x_data_time_strs, axis=-1)
# print(x_data_time_strs.shape)
# print(y_data_time_strs.shape)
return x_data_time_strs, y_data_time_strs
def data_generator_folder(data_indeces, feature_folder, file_reg, model_names, time_strs, batch_size, num_classes):
assert is_files_existed(feature_folder, file_reg, model_names, time_strs)
time_str_gen = time_str_generator(time_strs)
x_data, y_data = load_time_str_feature_data(data_indeces, feature_folder, file_reg, model_names, next(time_str_gen))
len_x_data = len(x_data)
start_index = 0
end_index = 0
while(1):
end_index = start_index + batch_size
if end_index < len_x_data:
# print(start_index, end_index, end=' ')
x_batch = x_data[start_index: end_index, :]
y_batch = y_data[start_index: end_index]
y_batch_cat = to_categorical(y_batch, num_classes)
start_index = start_index + batch_size
# print(x_batch.shape, y_batch_cat.shape)
yield x_batch, y_batch_cat
else:
end_index = end_index-len_x_data
# print(start_index, end_index, end=' ')
x_data_old = np.array(x_data[start_index:, :], copy=True)
y_data_old = np.array(y_data[start_index:], copy=True)
# Load new datas
x_data, y_data = load_time_str_feature_data(data_indeces, feature_folder, file_reg, model_names, next(time_str_gen))
# x_data, y_data = shuffle(x_data, y_data, random_state=2018)
len_x_data = len(x_data)
gc.collect()
x_batch = np.vstack((x_data_old, x_data[:end_index, :]))
y_batch = np.concatenate([y_data_old, y_data[:end_index]])
y_batch_cat = to_categorical(y_batch, num_classes)
start_index = end_index
# print(x_batch.shape, y_batch_cat.shape)
yield x_batch, y_batch_cat
# x_train = np.concatenate([x_train_Xception, x_train_InceptionV3, x_train_InceptionResNetV2], axis=-1)
num_classes = len_unique_landmark_ids
print('num_classes: %s' % num_classes)
file_reg = 'feature_%s_%s.h5'
model_names = [
# 'MobileNet',
# 'VGG16',
# 'VGG19',
# 'ResNet50',
# 'DenseNet121',
# 'DenseNet169',
# 'DenseNet201',
# 'Xception',
# 'InceptionV3',
'InceptionResNetV2'
]
time_strs = [
'200_20180312-050926',
# '150_20180311-151108'
]
batch_size = 8
print('*' * 60)
timesteps = len(model_names)
len_train_csv = len(train_indeces)
steps_per_epoch_train = int(len_train_csv/batch_size)
print('timesteps: %s' % timesteps)
print('len(train_data): %s' % len_train_csv)
print('batch_size: %s' % batch_size)
print('steps_per_epoch_train: %s' % steps_per_epoch_train)
train_gen = data_generator_folder(train_indeces, feature_folder, file_reg, model_names, time_strs, batch_size, num_classes)
batch_data = next(train_gen)
print(batch_data[0].shape, batch_data[1].shape)
batch_data = next(train_gen)
print(batch_data[0].shape, batch_data[1].shape)
# for i in range(steps_per_epoch_train*5):
# next(train_gen)
print('*' * 60)
len_val_csv = len(val_indeces)
steps_per_epoch_val = int(len_val_csv/batch_size) + 1
print('len(val_data): %s' % len_val_csv)
print('batch_size: %s' % batch_size)
print('steps_per_epoch_val: %s' % steps_per_epoch_val)
val_gen = data_generator_folder(val_indeces, feature_folder, file_reg, model_names, time_strs, batch_size, num_classes)
batch_data = next(val_gen)
print(batch_data[0].shape, batch_data[1].shape)
batch_data = next(val_gen)
print(batch_data[0].shape, batch_data[1].shape)
print('*' * 80)
data_dim = batch_data[0].shape[-1]
print('data_dim: %s' % data_dim)
%%time
def load_time_str_feature_test(feature_folder, file_reg, model_names, time_str):
x_test_time_strs = []
for model_name in model_names:
file_name = file_reg % (model_name, time_str)
file_path = os.path.join(feature_folder, file_name)
x_test_time_str = load_h5_test(feature_folder, file_reg, model_name, time_str)
# x_test_time_str = np.round(x_test_time_str, decimals=3) # 导致后面的全链接神经网络分类器不收敛
x_test_time_strs.append(x_test_time_str)
x_test_time_strs = np.concatenate(x_test_time_strs, axis=-1)
# print(x_test_time_strs.shape)
return x_test_time_strs
x_test = load_time_str_feature_test(feature_folder, file_reg, model_names, time_strs[0])
print(x_test.shape)
print(x_train.shape)
print(y_train.shape)
print(x_test.shape)
len_data = x_train.shape[0]
len_test = x_test.shape[0]
print('len_data: %s' % len_data)
print('len_test: %s' % len_test)
```
## Build NN
```
from sklearn.metrics import confusion_matrix
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
from keras.models import Sequential
from keras.layers import Dense, Dropout, Input, Flatten, Conv2D, MaxPooling2D, BatchNormalization
from keras.optimizers import Adam
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import LearningRateScheduler, TensorBoard
def get_lr(x):
lr = round(3e-4 * 0.97 ** x, 6)
if lr < 5e-4:
lr = 5e-4
print(lr, end=' ')
return lr
# annealer = LearningRateScheduler(lambda x: 1e-3 * 0.9 ** x)
annealer = LearningRateScheduler(get_lr)
callbacks = []
callbacks = [annealer]
log_dir = os.path.join(log_folder, run_name)
print('log_dir:' + log_dir)
tensorBoard = TensorBoard(log_dir=log_dir)
model = Sequential()
model.add(Dense(8192, input_shape=x_test.shape[1:]))
model.add(BatchNormalization())
model.add(Dropout(0.3))
model.add(Dense(4096, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(0.3))
model.add(Dense(len_unique_landmark_ids, activation='softmax'))
model.compile(
optimizer=Adam(lr=1e-4),
loss='categorical_crossentropy',
metrics=['accuracy']
)
model.summary()
# %%time
# hist = model.fit(
# x=x_train,
# y=y_train,
# batch_size=batch_size,
# epochs=20, #Increase this when not on Kaggle kernel
# verbose=1, #1 for ETA, 0 for silent
# callbacks=callbacks)
%%time
hist = model.fit_generator(
train_gen,
steps_per_epoch=steps_per_epoch_train,
epochs=20, #Increase this when not on Kaggle kernel
verbose=1, #1 for ETA, 0 for silent
callbacks=callbacks,
max_queue_size=2,
workers=4,
use_multiprocessing=False,
validation_data=val_gen,
validation_steps=steps_per_epoch_val
)
final_loss, final_acc = model.evaluate_generator(val_gen, steps=steps_per_epoch_val)
print("Final loss: {0:.4f}, final accuracy: {1:.4f}".format(final_loss, final_acc))
run_name_acc = run_name + '_' + str(int(final_acc*10000)).zfill(4)
histories = pd.DataFrame(hist.history)
histories['epoch'] = hist.epoch
print(histories.columns)
histories_file = os.path.join(model_folder, run_name_acc + '.csv')
histories.to_csv(histories_file, index=False)
import matplotlib.pyplot as plt
%matplotlib inline
plt.plot(hist.history['loss'], color='b')
plt.plot(hist.history['val_loss'], color='r')
plt.show()
plt.plot(hist.history['acc'], color='b')
plt.plot(hist.history['val_acc'], color='r')
plt.show()
def saveModel(model, run_name):
cwd = os.getcwd()
modelPath = os.path.join(cwd, 'model')
if not os.path.isdir(modelPath):
os.mkdir(modelPath)
weigthsFile = os.path.join(modelPath, run_name + '.h5')
model.save(weigthsFile)
saveModel(model, run_name_acc)
```
## Predict
```
y_pred = model.predict(x_test, batch_size=batch_size)
print(y_pred.shape)
# y_pred_file = os.path.join(model_folder, '%s.npy' % run_name_acc)
# np.save(y_pred_file, y_pred)
# y_pred_reload = np.load(y_pred_file)
# print(y_pred_file)
# print(y_pred_reload.shape)
# 这里证明os.listdir()得到的图片名称list不正确
files = os.listdir(os.path.join(cwd, 'input', 'data_test', 'test'))
print(files[:10])
# 这里证明ImageDataGenerator()得到的图片名称list才是正确
gen = ImageDataGenerator()
image_size = (299, 299)
# batch_size = 128
test_generator = gen.flow_from_directory(test_folder, image_size, shuffle=False, batch_size=batch_size)
print('test_generator')
print(len(test_generator.filenames))
print(test_generator.filenames[:10])
%%time
max_indexes = np.argmax(y_pred, -1)
print(max_indexes.shape)
test_dict = {}
for i, paire in enumerate(zip(test_generator.filenames, max_indexes)):
image_name, indx = paire[0], paire[1]
image_id = image_name[5:-4]
# test_dict[image_id] = '%d %.4f' % (indx, y_pred[i, indx])
test_dict[image_id] = '%d %.4f' % (indx, y_pred[i, indx])
#确认图片的id是否能与ImageDataGenerator()对应上
for key in list(test_dict.keys())[:10]:
print('%s %s' % (key, test_dict[key]))
display(sample_submission_csv.head(2))
%%time
len_sample_submission_csv = len(sample_submission_csv)
print('len(len_sample_submission_csv)=%d' % len_sample_submission_csv)
count = 0
for i in range(len_sample_submission_csv):
image_id = sample_submission_csv.iloc[i, 0]
# landmarks = sample_submission_csv.iloc[i, 1]
if image_id in test_dict:
pred_landmarks = test_dict[image_id]
# print('%s %s' % (image_id, pred_landmarks))
sample_submission_csv.iloc[i, 1] = pred_landmarks
else:
# print(image_id)
# sample_submission_csv.iloc[i, 1] = '9633 1.0' # 属于9633的类最多,所以全都设置成这个类,可能会比设置成空得到的结果好
sample_submission_csv.iloc[i, 1] = '' # 设置成空
count += 1
if count % 10000 == 0:
print(int(count/10000), end=' ')
display(sample_submission_csv.head(2))
pred_file = os.path.join(output_folder, 'pred_' + run_name_acc + '.csv')
sample_submission_csv.to_csv(pred_file, index=None)
print(run_name_acc)
print('Done !')
```
| github_jupyter |
# 1장. 머신러닝 개요
*아래 링크를 통해 이 노트북을 주피터 노트북 뷰어(nbviewer.org)로 보거나 구글 코랩(colab.research.google.com)에서 실행할 수 있습니다.*
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://nbviewer.org/github/rickiepark/handson-gb/blob/main/Chapter01/Gradient_Boosting_in_Machine_Learning.ipynb"><img src="https://jupyter.org/assets/share.png" width="60" />주피터 노트북 뷰어로 보기</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/rickiepark/handson-gb/blob/main/Chapter01/Gradient_Boosting_in_Machine_Learning.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
</td>
</table>
```
# 노트북이 코랩에서 실행 중인지 체크합니다.
import sys
if 'google.colab' in sys.modules:
!pip install -q --upgrade xgboost
!wget -q https://raw.githubusercontent.com/rickiepark/handson-gb/main/Chapter01/bike_rentals.csv
```
## 데이터 랭글링
### 데이터셋 1 - 자전거 대여
```
# 판다스를 임포트합니다.
import pandas as pd
# 'bike_rentals.csv'를 데이터프레임으로 읽습니다.
df_bikes = pd.read_csv('bike_rentals.csv')
# 처음 다섯 개 행을 출력합니다.
df_bikes.head()
```
### 데이터 이해하기
#### describe()
```
# df_bikes의 통계를 출력합니다.
df_bikes.describe()
```
#### info()
```
# df_bikes 정보를 출력합니다.
df_bikes.info()
```
### 누락된 값 처리하기
#### 누락된 값의 개수 계산하기
```
# 누락된 값의 개수 더하기
df_bikes.isna().sum().sum()
```
#### 누락된 값 출력하기
```
# df_bikes에 있는 누락된 값을 출력합니다.
df_bikes[df_bikes.isna().any(axis=1)]
```
#### 누락된 값 고치기
##### 중간값이나 평균으로 바꾸기
```
# windspeed의 누락된 값을 중간값으로 채웁니다.
df_bikes['windspeed'].fillna((df_bikes['windspeed'].median()), inplace=True)
# 인덱스 56과 81의 행을 출력합니다.
df_bikes.iloc[[56, 81]]
```
##### 중간값이나 평균으로 그룹바이 하기
```
# season으로 groupby한 중간값을 얻습니다.
df_bikes.groupby(['season']).median()
# 'hum' 열의 누락된 값을 season의 중간값으로 바꿉니다.
df_bikes['hum'] = df_bikes['hum'].fillna(df_bikes.groupby('season')['hum'].transform('median'))
```
##### 특정 행에서 중간값이나 평균을 구하기
```
# 'temp' 열의 누락된 값을 확인합니다.
df_bikes[df_bikes['temp'].isna()]
# temp와 atemp의 평균을 계산합니다.
mean_temp = (df_bikes.iloc[700]['temp'] + df_bikes.iloc[702]['temp'])/2
mean_atemp = (df_bikes.iloc[700]['atemp'] + df_bikes.iloc[702]['atemp'])/2
# 누락된 값을 평균 온도로 대체합니다.
df_bikes['temp'].fillna((mean_temp), inplace=True)
df_bikes['atemp'].fillna((mean_atemp), inplace=True)
```
##### 날짜 추정하기
```
# 'dteday' 열을 datetime 객체로 바꿉니다.
df_bikes['dteday'] = pd.to_datetime(df_bikes['dteday'])
df_bikes['dteday'].apply(pd.to_datetime, infer_datetime_format=True, errors='coerce')
# datetime을 임포트합니다.
import datetime as dt
df_bikes['mnth'] = df_bikes['dteday'].dt.month
# 마지막 다섯 개 행을 출력합니다.
df_bikes.tail()
# 인덱스 730 행의 'yr' 열을 1.0으로 바꿉니다.
df_bikes.loc[730, 'yr'] = 1.0
# 마지막 다섯 개 행을 출력합니다.
df_bikes.tail()
```
#### 수치형이 아닌 열 삭제하기
```
# 'dteday' 열을 삭제합니다.
df_bikes = df_bikes.drop('dteday', axis=1)
```
## 회귀 모델 만들기
### 자전거 대여 예측하기
```
# 'casual', 'registered' 열을 삭제합니다.
df_bikes = df_bikes.drop(['casual', 'registered'], axis=1)
```
### 나중을 위해서 데이터 저장하기
```
# 'bike_rentals_cleaned.csv' 파일로 저장합니다.
df_bikes.to_csv('bike_rentals_cleaned.csv', index=False)
```
### 특성과 타깃 준비하기
```
# X와 y로 데이터를 나눕니다.
X = df_bikes.iloc[:,:-1]
y = df_bikes.iloc[:,-1]
```
### 사이킷런 사용하기
```
# train_test_split 함수를 임포트합니다.
from sklearn.model_selection import train_test_split
# LinearRegression 클래스를 임포트합니다.
from sklearn.linear_model import LinearRegression
# 데이터를 훈련 세트와 테스트 세트로 나눕니다.
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=2)
```
### 경고 끄기
```
# 경고를 표시하지 않습니다.
import warnings
warnings.filterwarnings('ignore')
import xgboost as xgb
xgb.set_config(verbosity=0)
```
### 선형 회귀 모델 만들기
```
# LinearRegression 모델 객체를 만듭니다.
lin_reg = LinearRegression()
# 훈련 데이터로 lin_reg 객체를 훈련합니다.
lin_reg.fit(X_train, y_train)
# lin_reg 객체를 사용해 X_test에 대한 예측값을 만듭니다.
y_pred = lin_reg.predict(X_test)
# mean_squared_error 함수를 임포트합니다.
from sklearn.metrics import mean_squared_error
# numpy를 임포트합니다.
import numpy as np
# mean_squared_error 함수로 평균 제곱 오차를 계산합니다.
mse = mean_squared_error(y_test, y_pred)
# 평균 제곱근 오차를 계산합니다.
rmse = np.sqrt(mse)
# 평균 제곱근 오차를 출력합니다.
print("RMSE: %0.2f" % (rmse))
mean_squared_error(y_test, y_pred, squared=False)
# 'cnt' 열의 통계를 출력합니다.
df_bikes['cnt'].describe()
```
### XGBRegressor
```
# XGBRegressor를 임포트합니다.
from xgboost import XGBRegressor
# XGBRegressor의 객체 xg_reg를 만듭니다.
xg_reg = XGBRegressor()
# 훈련 데이터로 xg_reg 객체를 훈련합니다.
xg_reg.fit(X_train, y_train)
# 테스트 세트의 레이블을 예측합니다.
y_pred = xg_reg.predict(X_test)
# 평균 제곱 오차를 계산합니다.
mse = mean_squared_error(y_test, y_pred)
# 평균 제곱근 오차를 계산합니다.
rmse = np.sqrt(mse)
# 평균 제곱근 오차를 출력합니다.
print("RMSE: %0.2f" % (rmse))
```
### 교차 검증
#### 선형 회귀 교차 검증
```
# cross_val_score 함수를 임포트합니다.
from sklearn.model_selection import cross_val_score
# LinearRegression 클래스 객체를 만듭니다.
model = LinearRegression()
# 10-폴드 교차 검증으로 평균 제곱 오차를 구합니다.
scores = cross_val_score(model, X, y, scoring='neg_mean_squared_error', cv=10)
# 이 점수의 제곱근을 계산합니다.
rmse = np.sqrt(-scores)
# 평균 제곱근 오차를 출력합니다.
print('회귀 rmse:', np.round(rmse, 2))
# 평균 점수를 출력합니다.
print('RMSE 평균: %0.2f' % (rmse.mean()))
-np.mean(cross_val_score(model, X, y, scoring='neg_root_mean_squared_error', cv=10))
from sklearn.model_selection import cross_validate
cv_results = cross_validate(model, X, y, scoring='neg_root_mean_squared_error', cv=10)
-np.mean(cv_results['test_score'])
```
#### XGBoost 교차 검증
```
# XGBRegressor 객체를 만듭니다.
model = XGBRegressor(objective="reg:squarederror")
# 10-폴드 교차 검증으로 평균 제곱 오차를 구합니다.
scores = cross_val_score(model, X, y, scoring='neg_mean_squared_error', cv=10)
# 이 점수의 제곱근을 계산합니다.
rmse = np.sqrt(-scores)
# 평균 제곱근 오차를 출력합니다.
print('회귀 rmse:', np.round(rmse, 2))
# 평균 점수를 출력합니다.
print('RMSE 평균: %0.2f' % (rmse.mean()))
```
## 분류 모델 만들기
### 데이터 랭글링
#### 데이터 적재
```
# UCI 머신러닝 저장소에서 인구 조사 데이터셋(adult)을 로드합니다.
df_census = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data')
# 처음 다섯 개 행을 출력합니다.
df_census.head()
# 헤더가 없는 데이터셋을 로드합니다.
df_census = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data', header=None)
# 처음 다섯 개 행을 출력합니다.
df_census.head()
# df_census 열 이름을 정의합니다.
df_census.columns = ['age', 'workclass', 'fnlwgt', 'education', 'education-num', 'marital-status', 'occupation',
'relationship', 'race', 'sex', 'capital-gain', 'capital-loss', 'hours-per-week', 'native-country',
'income']
# 처음 다섯 개 행을 출력합니다.
df_census.head()
```
#### 누락된 값
```
# df_census 정보를 출력합니다.
df_census.info()
```
#### 수치형이 아닌 열
```
# education 열을 삭제합니다.
df_census = df_census.drop(['education'], axis=1)
# get_dummies를 사용해 숫자가 아닌 열을 바꿉니다.
df_census = pd.get_dummies(df_census)
# 처음 다섯 개 행을 출력합니다.
df_census.head()
```
#### 특성과 타깃 데이터
```
# 'income_ <=50K' 열을 삭제합니다.
df_census = df_census.drop('income_ <=50K', axis=1)
# 데이터를 X와 y로 나눕니다.
X = df_census.iloc[:,:-1]
y = df_census.iloc[:,-1]
```
### 로지스틱 회귀
```
# LogisticRegression을 임포트합니다.
from sklearn.linear_model import LogisticRegression
```
#### 교차 검증 함수
```
# classifier와 num_splits 매개변수를 가진 cross_val 함수를 정의합니다.
def cross_val(classifier, num_splits=10):
# 분류 모델 생성
model = classifier
# 교차 검증 점수 얻기
scores = cross_val_score(model, X, y, cv=num_splits)
# 정확도 출력
print('정확도:', np.round(scores, 2))
# 평균 정확도 출력
print('평균 정확도: %0.2f' % (scores.mean()))
# LogisticRegression으로 cross_val 함수를 호출합니다.
cross_val(LogisticRegression())
```
### XGBClassifier
```
# XGBClassifier를 임포트합니다.
from xgboost import XGBClassifier
# XGBClassifier로 cross_val 함수를 호출합니다.
cross_val(XGBClassifier(n_estimators=5))
```
| github_jupyter |
# Introduction to Python programming.
### [Gerard Gorman](http://www.imperial.ac.uk/people/g.gorman), [Christian Jacobs](http://christianjacobs.uk/)
### Updated for MPECDT by [David Ham](http://www.imperial.ac.uk/people/david.ham)
# Lecture 1: Computing with formulas
## Learning objectives:
* Execute a Python statement from within IPython.
* Learn what a program variable is and how to express a mathematical expression in code.
* Print program outputs.
* Access mathematical functions from a Python module.
## Programming a mathematical formula
Here is a formula for the position of a ball in vertical motion, starting at ground level (i.e. $y=0$) at time $t=0$:
$$ y(t) = v_0t- \frac{1}{2}gt^2 $$
where:
* $y$ is the height (position) as a function of time $t$
* $v_0$ is the initial velocity (at $t=0$)
* $g$ is the acceleration due to gravity
The computational task is: given $v_0$, $g$ and $t$, compute the value $y$.
**How do we program this task?** A program is a sequence of instructions given to the computer. However, while a programming language is much **simpler** than a natural language, it is more **pedantic**. Programs must have correct syntax, i.e., correct use of the computer language grammar rules, and no misprints.
So let's execute a Python statement based on this example. Evaluate $y(t) = v_0t- \frac{1}{2}gt^2$ for $v_0=5$, $g=9.81$ and $t=0.6$. If you were doing this on paper you would probably write something like this: $$ y = 5\cdot 0.6 - {1\over2}\cdot 9.81 \cdot 0.6^2.$$ Happily, writing this in Python is very similar:
```
print(5*0.6 - 0.5*9.81*0.6**2)
```
Go ahead and mess with the code above to see what happens when you change values and rerun. To see what I mean about programming being pedantic, see what happens if you replace `**` with `^`:
```
print(5*0.6 - 0.5*9.81*0.6**2)
```
or `write` rather than `print`:
```
write (5*0.6 - 0.5*9.81*0.6**2)
```
While a human might still understand these statements, they do not mean anything to the Python interpreter. Rather than throwing your hands up in the air whenever you get an error message like the above (you are going to see many during the course of these lectures!!!) train yourself to read the message patiently to get an idea what it is complaining about and re-read your code from the perspective of the pedantic Python interpreter.
Error messages can look bewildering (frustrating etc.) at first, but it gets much **easier with practise**.
## Storing numbers in variables
From mathematics you are already familiar with variables (e.g. $v_0=5,\quad g=9.81,\quad t=0.6,\quad y = v_0t -{1\over2}gt^2$) and you already know how important they are for working out complicated problems. Similarly, you can use variables in a program to make it easier to read and understand.
```
v0 = 5
g = 9.81
t = 0.6
y = v0*t - 0.5*g*t**2
print(y)
a=2
print(type(a))
a=2.5
print(type(a))
```
This program spans several lines of text and uses variables, otherwise the program performs the same calculations and gives the same output as the previous program.
In mathematics we usually use one letter for a variable, resorting to using the Greek alphabet and other characters for more clarity. The main reason for this is to avoid becoming exhausted from writing when working out long expressions or derivations. However, when programming you should use more descriptive names for variable names. This might not seem like an important consideration for the trivial example here but it becomes increasingly important as the program gets more complicated and if someone else has to read your code. **Good variable names make a program easier to understand!**
Permitted variable names include:
* One-letter symbols.
* Words or abbreviation of words.
* Variable names can contain a-z, A-Z, underscore ("'_'") and digits 0-9, **but** the name cannot start with a digit.
* In Python 3, variable names can also include letters from other alphabets, such as α or π.
Variable names are case-sensitive (i.e. "'a'" is different from "'A'"). Let's rewrite the previous example using more descriptive variable names:
```
initial_velocity = 5
g = 9.81
TIME = 0.6
VerticalPositionOfBall = initial_velocity*TIME - 0.5*g*TIME**2
print(VerticalPositionOfBall)
from math import pi as π
radius = 2
area = π * radius ** 2
print(area)
```
Certain words have are **reserved** in Python and **cannot be used as variable names**. These are: *and, as, assert, break, class, continue, def, del, elif, else, except, exec, finally, for, from, global, if, import, in, is, lambda, not, or, pass, print, raise, return, try, with, while,* and *yield*.
## Adding comments to code
Not everything written in a computer program is intended for execution. In Python anything on a line after the '#' character is ignored and is known as a **comment**. You can write whatever you want in a comment. Comments are intended to be used to explain what a snippet of code is intended for. It might for example explain the objective or provide a reference to the data or algorithm used. This is both useful for you when you have to understand your code at some later stage, and indeed for whoever has to read and understand your code later.
```
# Program for computing the height of a ball in vertical motion.
v0 = 5 # Set initial velocity in m/s.
g = 9.81 # Set acceleration due to gravity in m/s^2.
t = 0.6 # Time at which we want to know the height of the ball in seconds.
y = v0*t - 0.5*g*t**2 # Calculate the vertical position
print(y)
```
## <span style="color:blue">Exercise: Convert from meters to Imperial length units</span>
Make a program where you set a length given in meters and then compute and write out the corresponding length measured in inches, in feet, in yards, and in miles. Use the fact that one inch is 2.54 cm, one foot is 12 inches, one yard is 3 feet, and one British mile is 1760 yards. As a verification, a length of 640 meters corresponds to 25196.85 inches, 2099.74 feet, 699.91 yards, or 0.3977 miles.
```
length = 640
print(length*100/(2.54))
print(length*100/(2.54*12))
print(length*100/(2.54*12*3))
print(length*100/(2.54*12*3*1760))
```
## Formatting numbers as strings
Often we want to print out results using a combination of text and numbers, e.g. "'At t=0.6 s, y is 1.23 m'". Particularly when printing out floating point numbers we should **never** quote numbers to a higher accuracy than they were measured. Python provides a *printf formatting* syntax exactly for this purpose. We can see in the following example that the *slot* `%g` was used to express the floating point number with the minimum number of significant figures, and the *slot* `%.2f` specified that only two digits are printed out after the decimal point.
```
print("At t=%gs, y is %.2fm." % (t, y))
```
Notice in this example how the values in the tuple `(t, y)` are inserted into the *slots*.
Sometimes we want a multi-line output. This is achieved using a triple quotation (*i.e.* `"""`):
```
print("""At t=%f s, a ball with
initial velocity v0=%.3E m/s
is located at the height %.2f m.
""" % (t, v0, y))
```
## <span style="color:blue">Exercise: Compute the air resistance on a football</span>
The drag force, due to air resistance, on an object can be expressed as
$$F_d = \frac{1}{2}C_D\rho AV^2$$
where $\rho$ is the density of the air, $V$ is the velocity of the object, $A$ is the cross-sectional area (normal to the velocity direction), and $C_D$ is the drag coefficient, which depends heavily on the shape of the object and the roughness of the surface.</br></br>
The gravity force on an object with mass $m$ is $F_g = mg$, where $g = 9.81ms^{−2}$.</br></br>
Write a program that computes the drag force and the gravity force on an object. Write out the forces with one decimal in units of Newton ($N = kgm/s^2$). Also print the ratio of the drag force and the gravity force. Define $C_D$, $\rho$, $A$, $V$, $m$, $g$, $F_d$, and $F_g$ as variables, and put a comment with the corresponding unit.</br></br>
As a computational example, you can initialize all variables with values relevant for a football kick. The density of air is $\rho = 1.2 kg m^{−3}$. For any ball, we have obviously that $A = \pi a^2$, where $a$ is the radius of the ball, which can be taken as $11cm$ for a football. The mass of the ball is $0.43kg$. $C_D$ can be taken as $0.2$.</br></br>
Use the program to calculate the forces on the ball for a hard kick, $V = 120km/h$ and for a soft kick, $V = 10km/h$ (it is easy to make the mistake of mixing inconsistent units, so make sure you compute with V expressed in m/s). Make sure you use the *printf* formatting style introduced above.
```
from math import pi as π
a = 0.11 #m
A = π*a**2 #m^2
C_d = 0.2
rho = 1.2 #kg/m^3
g = 9.81 #m/s^2
m = 0.43 #kg
Vh = 120 *1000/(60 *60)
Vs = 10 * 1000/(60*60)
F_dh = 0.5 * C_d * rho * A * Vh ** 2
F_ds = 0.5 * C_d * rho * A * Vs ** 2
F_g = m * g
print("""With a hard kick the drag force on the ball is %.1fN and a soft kick the drag force is %.1eN.
The gravitational force is %.1fN. The ratio for the hard kick is %.1f.
""" % (F_dh, F_ds, F_g, F_dh/F_g))
```
## How are arithmetic expressions evaluated?
Consider the random mathematical expression, ${5\over9} + 2a^4/2$, implemented in Python as `5.0/9 + 2*a**4/2`.
The rules for evaluating the expression are the same as in mathematics: proceed term by term (additions/subtractions) from the left, compute powers first, then multiplication and division. Therefore in this example the order of evaluation will be:
1. `r1 = 5.0/9`
2. `r2 = a**4`
3. `r3 = 2*r2`
4. `r4 = r3/2`
5. `r5 = r1 + r4`
Use parenthesis to override these default rules. Indeed, many programmers use parenthesis for greater clarity.
## <span style="color:blue">Exercise: Compute the growth of money in a bank</span>
Let *p* be a bank's interest rate in percent per year. An initial amount *A* has then grown to $$A\left(1+\frac{p}{100}\right)^n$$ after *n* years. Write a program for computing how much money 1000 euros have grown to after three years with a 5% interest rate.
## Standard mathematical functions
What if we need to compute $\sin x$, $\cos x$, $\ln x$, etc. in a program? Such functions are available in Python's *math module*. In fact there is a vast universe of functionality for Python available in modules. We just *import* in whatever we need for the task at hand.
In this example we compute $\sqrt{2}$ using the *sqrt* function in the *math* module:
```
import math
r = math.sqrt(2)
print(r)
```
or:
```
from math import sqrt
r = sqrt(2)
print(r)
```
or:
```
from math import * # import everything in math
r = sqrt(2)
print(r)
```
Another example:
```
from math import sin, cos, log
x = 1.2
print(sin(x)*cos(x) + 4*log(x)) # log is ln (base e)
```
## <span style="color:blue">Exercise: Evaluate a Gaussian function</span>
The bell-shaped Gaussian function,
$$f(x)=\frac{1}{\sqrt{2\pi}s}\exp\left(-\frac{1}{2} \left(\frac{x-m}{s}\right)^2\right)$$
is one of the most widely used functions in science and technology. The parameters $m$ and $s$ are real numbers, where $s$ must be greater than zero. Write a program for evaluating this function when $m = 0$, $s = 2$, and $x = 1$. Verify the program's result by comparing with hand calculations on a calculator.
```
dir(math)
help(math.exp)
math.acos?
s = 2
1/sqrt(2*math.pi*s)*exp(-0.5*((1/s)**2))
```
| github_jupyter |
# Generate Reactions
This script performs the same task as the script in `scripts/generateReactions.py` but in visual ipynb format.
It can also evaluate the reaction forward and reverse rates at a user selected temperature.
```
from rmgpy.rmg.main import RMG
from rmgpy.rmg.model import CoreEdgeReactionModel
from rmgpy import settings
from IPython.display import display
from arkane.output import prettify
```
Declare database variables here by changing the thermo and reaction libraries, or restrict to certain reaction families.
```
database = """
database(
thermoLibraries = ['BurkeH2O2','primaryThermoLibrary','DFT_QCI_thermo','CBS_QB3_1dHR','Narayanaswamy','Chernov'],
reactionLibraries = [],
seedMechanisms = [],
kineticsDepositories = ['training'],
kineticsFamilies = [
'H_Abstraction',
'R_Addition_MultipleBond',
'intra_H_migration',
'Intra_R_Add_Endocyclic',
'Intra_R_Add_Exocyclic'
],
kineticsEstimator = 'rate rules',
)
options(
verboseComments=True, # Set to True for detailed kinetics comments
)
"""
```
List all species you want reactions between
```
species_list = """
species(
label='i1',
reactive=True,
structure=adjacencyList(
\"""
multiplicity 2
1 C u0 p0 c0 {3,S} {4,S} {10,S} {11,S}
2 C u0 p0 c0 {4,S} {12,S} {13,S} {14,S}
3 C u0 p0 c0 {1,S} {5,B} {6,B}
4 C u1 p0 c0 {1,S} {2,S} {15,S}
5 C u0 p0 c0 {3,B} {8,B} {19,S}
6 C u0 p0 c0 {3,B} {9,B} {20,S}
7 C u0 p0 c0 {8,B} {9,B} {17,S}
8 C u0 p0 c0 {5,B} {7,B} {16,S}
9 C u0 p0 c0 {6,B} {7,B} {18,S}
10 H u0 p0 c0 {1,S}
11 H u0 p0 c0 {1,S}
12 H u0 p0 c0 {2,S}
13 H u0 p0 c0 {2,S}
14 H u0 p0 c0 {2,S}
15 H u0 p0 c0 {4,S}
16 H u0 p0 c0 {8,S}
17 H u0 p0 c0 {7,S}
18 H u0 p0 c0 {9,S}
19 H u0 p0 c0 {5,S}
20 H u0 p0 c0 {6,S}
\"""
)
)
"""
# Write input file to disk
with open('temp/input.py','w') as input_file:
input_file.write(database)
input_file.write(species_list)
# Execute generate reactions
from rmgpy.tools.generate_reactions import RMG, execute
kwargs = {
'walltime': '00:00:00:00',
'kineticsdatastore': True
}
rmg = RMG(input_file='temp/input.py', output_directory='temp')
rmg = execute(rmg, **kwargs)
# Pick some temperature to evaluate the forward and reverse kinetics
T = 623.0 # K
for rxn in rmg.reaction_model.output_reaction_list:
print('=========================')
display(rxn)
print('Reaction Family = {0}'.format(rxn.family))
print('')
print('Reactants')
for reactant in rxn.reactants:
print('Label: {0}'.format(reactant.label))
print('SMILES: {0}'.format(reactant.molecule[0].to_smiles()))
print('')
print('Products')
for product in rxn.products:
print('Label: {0}'.format(product.label))
print('SMILES: {0}'.format(product.molecule[0].to_smiles()))
print('')
print(rxn.to_chemkin())
print('')
print('Heat of Reaction = {0:.2F} kcal/mol'.format(rxn.get_enthalpy_of_reaction(623.0)/4184))
print('Forward kinetics at {0} K: {1:.2E}'.format(T, rxn.get_rate_coefficient(T)))
reverseRate = rxn.generate_reverse_rate_coefficient()
print('Reverse kinetics at {0} K: {1:.2E}'.format(T, reverseRate.get_rate_coefficient(T)))
```
| github_jupyter |
<br><br><font color="gray">DOING COMPUTATIONAL SOCIAL SCIENCE<br>MODULE 4 <strong>PROBLEM SETS</strong></font>
# <font color="#49699E" size=40>MODULE 4 </font>
# What You Need to Know Before Getting Started
- **Every notebook assignment has an accompanying quiz**. Your work in each notebook assignment will serve as the basis for your quiz answers.
- **You can consult any resources you want when completing these exercises and problems**. Just as it is in the "real world:" if you can't figure out how to do something, look it up. My recommendation is that you check the relevant parts of the assigned reading or search for inspiration on [https://stackoverflow.com](https://stackoverflow.com).
- **Each problem is worth 1 point**. All problems are equally weighted.
- **The information you need for each problem set is provided in the blue and green cells.** General instructions / the problem set preamble are in the blue cells, and instructions for specific problems are in the green cells. **You have to execute all of the code in the problem set, but you are only responsible for entering code into the code cells that immediately follow a green cell**. You will also recognize those cells because they will be incomplete. You need to replace each blank `▰▰#▰▰` with the code that will make the cell execute properly (where # is a sequentially-increasing integer, one for each blank).
- Most modules will contain at least one question that requires you to load data from disk; **it is up to you to locate the data, place it in an appropriate directory on your local machine, and replace any instances of the `PATH_TO_DATA` variable with a path to the directory containing the relevant data**.
- **The comments in the problem cells contain clues indicating what the following line of code is supposed to do.** Use these comments as a guide when filling in the blanks.
- **You can ask for help**. If you run into problems, you can reach out to John (john.mclevey@uwaterloo.ca) or Pierson (pbrowne@uwaterloo.ca) for help. You can ask a friend for help if you like, regardless of whether they are enrolled in the course.
Finally, remember that you do not need to "master" this content before moving on to other course materials, as what is introduced here is reinforced throughout the rest of the course. You will have plenty of time to practice and cement your new knowledge and skills.
<div class='alert alert-block alert-danger'>As you complete this assignment, you may encounter variables that can be assigned a wide variety of different names. Rather than forcing you to employ a particular convention, we leave the naming of these variables up to you. During the quiz, submit an answer of 'USER_DEFINED' (without the quotation marks) to fill in any blank that you assigned an arbitrary name to. In most circumstances, this will occur due to the presence of a local iterator in a for-loop.</b></div>
## Package Imports
```
import pandas as pd
import numpy as np
from pprint import pprint
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score, silhouette_samples
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
%config Completer.use_jedi = False
```
## Defaults
```
seed = 7
```
## Problem 1:
<div class="alert alert-block alert-info">
In this exercise, we're going to ask you to supply the names of the Pandas methods you'll need to (1) load the .csv from disk and (2) preview a random sample of 5 rows.
</div>
<div class="alert alert-block alert-success">
In the code block below, fill in the blanks to insert the functions, methods, or variable names needed to load the .csv and draw a random sample of 5 rows.
</div>
```
# Load vdem_subset.csv as a dataframe
df = pd.▰▰1▰▰(PATH_TO_DATA/'vdem_subset.csv', low_memory=False, index_col=0)
# Draw random sample of 5 rows from vdem dataframe
df.▰▰2▰▰(▰▰3▰▰, random_state = 7)
```
## Problem 2:
<div class="alert alert-block alert-info">
You may have noticed that many of the cells in the dataframe we created have 'NaN' values. It's useful for us to know just how many values in our dataset are missing or not defined. Let's do that now:
</div>
<div class="alert alert-block alert-success">
In the code block below, fill in the blanks to insert the functions, methods, or variable names needed to create a Pandas series of the missing values for each column and then sort it.
</div>
```
# Sum together all NaN values to produce series with numerical values indicating number of missing entires
missing = df.▰▰1▰▰().▰▰2▰▰()
# Sort the `missing` series
missing = missing.▰▰3▰▰()
print(missing)
print("Total missing values: " + str(sum(missing)))
```
## Problem 3:
<div class="alert alert-block alert-info">
The list below contains a number of variables, including mid-level indicators that go into the 5 high-level democracy indexes that were used in the assigned readings. In this problem, we'll subset our data in two ways - first conceptually by selecting only the mid-level indicators, and then empirically by selecting the indicators that heave the least missing data.
</div>
<div class="alert alert-block alert-success">
Use the list of column names we've provided to filter the large dataframe into a subset. Fill in the blanks to insert the functions, methods, or variable names needed.
</div>
```
vd_meta_vars = ['country_name', 'year', 'e_regiongeo']
vd_index_vars = ['v2x_freexp_altinf', 'v2x_frassoc_thick', 'v2x_suffr', 'v2xel_frefair', 'v2x_elecoff', # electoral democracy index
'v2xcl_rol', 'v2x_jucon', 'v2xlg_legcon', # liberal democracy index
'v2x_cspart', 'v2xdd_dd', 'v2xel_locelec', 'v2xel_regelec', 'v2x_polyarchy', # participatory democracy index
'v2dlreason', 'v2dlcommon', 'v2dlcountr', 'v2dlconslt', 'v2dlengage', # deliberative democracy index
'v2xeg_eqprotec', 'v2xeg_eqaccess', 'v2xeg_eqdr'] # egalitarian democracy index
# filter `df` so that it only includes columns from the two lists above
sdf = df[▰▰1▰▰ ▰▰2▰▰ vd_index_vars]
sdf.describe()
```
## Problem 4:
<div class="alert alert-block alert-info">
One useful thing that using Pandas dataframes enables us to do is group data based on one or more the columns and then work with the resulting grouped dataframe (in much the same way we would with an un-grouped dataframe). Using the VDEM data, we'll only import a subset of the data, using the 'columns_to_use' variable. At the same time, we're going to replace the numerical values in the 'e_regionpol_6c' variable with easy-to-read string representations. Finally, we'll filter the resulting dataset to include only those rows from the year 2015.
</div>
<div class="alert alert-block alert-success">
In this next code block, we're going to load in a dataset, filtering our dataframe to include only those rows where the year is 2015. Fill in the blanks to continue.
</div>
```
columns_to_use = [
'country_name',
'country_id',
'year',
'e_area',
'e_regionpol_6C',
'v2x_polyarchy',
'v2x_libdem',
'v2x_partipdem',
'v2x_delibdem',
'v2x_egaldem'
]
# Load the dataset as a dataframe
df = pd.▰▰1▰▰(
PATH_TO_DATA/"vdem_subset.csv",
usecols = ▰▰2▰▰,
low_memory = False
)
df['e_regionpol_6C'].replace({
1.0: "East Europe and Central Asia",
2.0: "Latin America and Carribean",
3.0: "Middle East and North Africa",
4.0: "Sub-Saharan Africa",
5.0: "West Europe and North America",
6.0: 'Asia and Pacific'
}, inplace=True)
# Subset the dataframe to include only those rows from 2015
df_2015 = df.▰▰3▰▰("year ▰▰4▰▰ 2015")
df_2015
```
## Problem 5:
<div class="alert alert-block alert-info">
Now, we're going to use the Pandas Dataframe's `groupby` method to combine each nation into the region it belongs to. As you would have read in the accompanying chapter, the Pandas groupby method only preserves columns that you give it instructions for; everything else is dropped in the resulting dataframe.
<br><br>
In order to figure out how to aggregate each of our columns, let's think through them together. First up, we have 'country_name' and 'country_ID'. Since we're going to be grouping our data into only 6 rows (one for each of the 6 politico-geographical regions), it doesn't make sense to keep either of these columns. The same goes for 'year', since we will have already filtered our dataset to only include rows that are from 2015. We're going to be using 'e_regionpol_6C' as the basis for our groupings, so it doesn't make sense to keep it as a data column any longer.
<br><br>
That leaves us with 'e_area' and the 5 democracy indices. Since we're interested in knowing the total area of each region, it would make sense to <b>add</b> each country's area together. We could do something similar for the 5 democracy indices, but we'll leave them alone for now. In order to make things easier on ourselves, we're going to start by filtering out all of the columns we don't want in our final dataset, which will make aggregating what's left much easier.
</div>
<div class="alert alert-block alert-success">
In the following code cell, we're going to filter out most of the columns in `df_2015` so that only 'e_regionpol_6C' and 'e_area' remain, and store the resulting filtered dataframe as `df_area`. Then, we're going to run a `groupby` operation on the `e_regionpol_6C` column of and sum the `e_area` column in the `df_area` dataframe. Fill in the blanks to continue.
</div>
```
# Filter out all columns except 'e_regionpol_6C', 'e_area'
df_area = ▰▰1▰▰[['e_regionpol_6C', 'e_area']]
# group by political region and sum remaining columns
df_grouped_area = df_area.▰▰2▰▰('e_regionpol_6C').▰▰3▰▰()
df_grouped_area
```
## Problem 6:
<div class="alert alert-block alert-info">
In the last question, we explored how we could use Pandas to group rows of a dataframe according to a variable's value, and to handle a subset of the remaining columns according to some kind of aggregation logic (such as adding the values or averaging over them). This time, rather than lumping countries together by region, we're going to drill deeper on how an individual nation has changed over time. For this exercise, we're going to look at how democratic norms in Costa Rica have developed in the decades since the Second World War. Since we already have the full dataframe stored in memory (as 'df'), we'll start by filtering our dataset to include only those rows pertaining to Costa Rica (across all years, not just 2015).
<br><br>
If you examine the resulting dataframe, you might notice that Costa Rica does not have any scores for the 5 democratic indices the earlier years for which it is present in the dataset. This should come as no surprise; even for a group as capable as the VDEM project, constructing a democratic index for the year 1839 would involve enough guesswork to render the result meaningless. As such, we're going to immediately filter our Costa Rica-only dataframe to weed out any rows that don't have scores for the 5 democratic indices.
</div>
<div class="alert alert-block alert-success">
Find the first year for which we have a complete set of the democratic indices for Costa Rica. Fill in the blanks to continue.
</div>
```
# Filter the dataframe to include only rows pertaining to Costa Rica
df_cr = df.▰▰1▰▰("▰▰2▰▰ ▰▰3▰▰ 'Costa Rica'")
# Drop each row with one or more missing values
df_cr_filtered = df_cr.▰▰4▰▰(subset=[
'v2x_polyarchy',
'v2x_libdem',
'v2x_partipdem',
'v2x_delibdem',
'v2x_egaldem'])
# Find first year for which VDEM has a complete set of indices for Costa Rica
first_year = ▰▰5▰▰(df_cr_filtered[▰▰6▰▰])
```
## Problem 7:
<div class="alert alert-block alert-info">
Now our data is ready to be plotted! In this part of the exercise, we're going to plot two of Costa Rica's democratic indices against the 'year' variable to see how its democratic norms have evolved over time. We'll accomplish this by using Seaborn and taking advantage of the fact that the columns in Pandas Dataframes can be individually 'pulled out' as a Series (which operate similarly to Numpy arrays, for most intents and purposes). In the following code cell, we'll create the plot for you so you can see how it's done and what it should look like. It won't be graded, and there aren't any blanks to fill in.
<br><br>Despite being as simple as can be, that doesn't look half bad! It's always a good idea to label your axes and give the plot a title so that anyone encoutering it for the first time can rapidly determine what the plot represents.
<br><br>A quick note; if you want to see what the first label-less plot looks like before adding labels to the second plot, you can comment out each of the lines below the first instance of <code>figure.show()</code>.
</div>
<div class="alert alert-block alert-success">
Add useful labels to the x-axis and y-axis of the second plot produced by the code cell below, along with a title describing what the plot is. Fill in the blanks to continue.
</div>
```
cr_years = df_cr_filtered['year']
cr_polyarchy = df_cr_filtered['v2x_polyarchy']
figure = plt.figure(figsize=(10, 6))
sns.lineplot(x = cr_years, y = cr_polyarchy)
figure.show()
figure = plt.figure(figsize=(10, 6))
sns.lineplot(x = cr_years, y = cr_polyarchy)
# Label y-axis
plt.ylabel(▰▰1▰▰)
# Label x-axis
plt.▰▰2▰▰(▰▰3▰▰)
# Add title
▰▰4▰▰.▰▰5▰▰("Polyarchy over Time, Costa Rica")
figure.show()
```
## Problem 8:
<div class="alert alert-block alert-info">
In this exercise, we're going to work through how to combine multiple pandas dataframes. This will come in handy whenever you want to explore the relationships between variables that come from different datasets, but which can be linked according to some underlying relationship.
<br><br>
Earlier, we used addition to aggregate the land area of every nation in a politico-geographic region to give us a sense of how large each region was. In this exercise, we're going to turn our attention to the 5 democracy indices. Using addition (which is what we did with area) to aggregate the 5 democracy indices doesn't make as much sense, though: that might lead us to conclude that regions with more countries would be 'more democratic' than those with only a small number of nations. Instead, we'll *average* over these indicators, which will give us a sense of how democratic each region is, taken together.
</div>
<div class="alert alert-block alert-success">
Create a dataframe that only includes the columns we care about (the region variable and the 5 democratic indices), group the result by region, and take the average across each score.
</div>
```
df_democracy = df_2015[['v2x_polyarchy',
'v2x_libdem',
'v2x_partipdem',
'v2x_delibdem',
'v2x_egaldem',
'e_regionpol_6C']]
# Group by region and take average of other variables
df_grouped_democracy = ▰▰1▰▰.▰▰2▰▰('▰▰3▰▰').▰▰4▰▰()
df_grouped_democracy
```
## Problem 9:
<div class="alert alert-block alert-info">
If you compare the 'df_grouped_democracy' dataframe and the 'df_grouped_area' dataframe, you might notice that the bolded columns on the left are identical. You may recall that the bold column on the left of a dataframe is the 'index', and we can take advantage of its special status to join the two dataframes together. The result will be one dataframe with the same number of rows, but with all 6 of the columns we aggregated: area and the 5 democratic indices.
</div>
<div class="alert alert-block alert-success">
In the following code block, we're going to concatenate `df_grouped_democracy` and `df_grouped`area. Fill in the blanks to continue.
</div>
```
# Concatenate df_grouped_democracy and df_grouped_area on rows
df_full_rows = pd.▰▰1▰▰([df_grouped_democracy, df_grouped_area], ▰▰2▰▰=1)
df_full_rows
```
## Problem 10:
<div class="alert alert-block alert-info">
In the above exercise, we combined dataframes along their rows, using the row index to guide how the data was combined. We can do much the same with columns. To demonstrate how, let's return to our Costa Rica dataframe and add another country to it. Since Costa Rica and Nicaragua are geographic neighbours, it makes sense to compare them directly.
</div>
<div class="alert alert-block alert-success">
Time to give Nicaragua the same treatment as we did to Costa Rica! Once that's done, we're going to concatenate `df_nicaragua_filtered` and `df_nicaragua_cr`, column-wise. Fill in the blanks to continue.
</div>
```
# Create a dataframe only containing rows pertaining to Nicaragua
df_nicaragua = df.▰▰1▰▰("country_name ▰▰2▰▰ 'Nicaragua'")
# Drop rows in the Nicaragua dataframe that contain NaNs in the 5 index columns
df_nicaragua_filtered = df_nicaragua.▰▰3▰▰(subset=[
'v2x_polyarchy',
'v2x_libdem',
'v2x_partipdem',
'v2x_delibdem',
'v2x_egaldem'])
# Concatenate df_grouped_democracy and df_grouped_area on columns
df_nicaragua_cr = pd.▰▰4▰▰([df_nicaragua_filtered, ▰▰5▰▰], axis = ▰▰6▰▰)
df_nicaragua_cr
```
## Problem 11:
<div class="alert alert-block alert-info">
Now that the data for these two countries has been combined into a single dataframe, we can easily create plots that allow us to compare them. Again, we'll be using the Seaborn package to do our plotting for us. Even though all of our data is lumped together, Seaborn allows us to use the 'hue' variable to differentiate the data we're plotting based on some categorical variable (which, in this case, is the country variable -- it's what differentiates between Costa Rica and Nicaragua).
</div>
<div class="alert alert-block alert-success">
Create a line plot that contains separate lines for both Nicaragua's and Costa Rica's polyarchy score by year. We'll also include labels for the x-axis, y-axis, and plot title. Fill in the blanks to continue.
</div>
```
concat_years = df_nicaragua_cr['year']
concat_polyarchy = df_nicaragua_cr['v2x_polyarchy']
concat_country = df_nicaragua_cr['country_name']
figure = plt.figure(figsize=(10,6))
ax = sns.▰▰1▰▰(x=▰▰2▰▰,
y=▰▰3▰▰,
hue=▰▰4▰▰
)
ax.set(xlabel='Year', ylabel='Polyarchy', title="Polyarchy over Time")
figure.show()
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#@title MIT License
#
# Copyright (c) 2017 François Chollet
#
# Permission is hereby granted, free of charge, to any person obtaining a
# copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
```
# Train your first neural network: basic classification
<table class="tfo-notebook-buttons" align="left"><td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/models/blob/master/samples/core/get_started/basic_classification.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" /><span>Run in Google Colab</span></a>
</td><td>
<a target="_blank" href="https://github.com/tensorflow/models/blob/master/samples/core/get_started/basic_classification.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /><span>View source on GitHub</span></a></td></table>
In this guide, we will train a neural network model to classify images of clothing, like sneakers and shirts. It's fine if you don't understand all the details, this is a fast-paced overview of a complete TensorFlow program with the details explained as we go.
This guide uses [tf.keras](https://www.tensorflow.org/guide/keras), a high-level API to build and train models in TensorFlow.
```
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
```
## Import the Fashion MNIST dataset
This guide uses the [Fashion MNIST](https://github.com/zalandoresearch/fashion-mnist) dataset which contains 70,000 grayscale images in 10 categories. The images show individual articles of clothing at low resolution (28 by 28 pixels), as seen here:
<table>
<tr><td>
<img src="https://tensorflow.org/images/fashion-mnist-sprite.png"
alt="Fashion MNIST sprite" width="600">
</td></tr>
<tr><td align="center">
<b>Figure 1.</b> <a href="https://github.com/zalandoresearch/fashion-mnist">Fashion-MNIST samples</a> (by Zalando, MIT License).<br/>
</td></tr>
</table>
Fashion MNIST is intended as a drop-in replacement for the classic [MNIST](http://yann.lecun.com/exdb/mnist/) dataset—often used as the "Hello, World" of machine learning programs for computer vision. The MNIST dataset contains images of handwritten digits (0, 1, 2, etc) in an identical format to the articles of clothing we'll use here.
This guide uses Fashion MNIST for variety, and because it's a slightly more challenging problem than regular MNIST. Both datasets are relatively small and are useful to verify that an algorithm works as expected. They're good starting points to test and debug code.
We will use 60,000 images to train the network and 10,000 images to evaluate how accurately the network learned to classify images. You can acess the Fashon MNIST directly from TensorFlow, just import and load the data:
```
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
```
Loading the dataset returns four NumPy arrays:
* The `train_images` and `train_labels` arrays are the *training set*, this is the data the model uses to learn.
* The model is tested against the *test set*, the `test_images` and `test_labels` arrays.
The images are 28x28 numpy arrays, with pixel values ranging between 0 and 255. The *labels* are an array of integers, ranging from 0 to 9. These correspond to the *class* of clothing the image represents:
<table>
<tr>
<th>Label</th>
<th>Class</th>
</tr>
<tr>
<td>0</td>
<td>T-shirt/top</td>
</tr>
<tr>
<td>1</td>
<td>Trouser</td>
</tr>
<tr>
<td>2</td>
<td>Pullover</td>
</tr>
<tr>
<td>3</td>
<td>Dress</td>
</tr>
<tr>
<td>4</td>
<td>Coat</td>
</tr>
<tr>
<td>5</td>
<td>Sandal</td>
</tr>
<tr>
<td>6</td>
<td>Shirt</td>
</tr>
<tr>
<td>7</td>
<td>Sneaker</td>
</tr>
<tr>
<td>8</td>
<td>Bag</td>
</tr>
<tr>
<td>9</td>
<td>Ankle boot</td>
</tr>
</table>
Each image is mapped to a single label. Since the *class names* are not included with the dataset, store them here to use later when plotting the images:
```
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
```
## Explore the data
Let's explore the format of the dataset before training the model. The following shows there are 60,000 images in the training set, with each image represented as 28 x 28 pixels:
```
train_images.shape
```
Likewise, there are 60,000 labels in the training set:
```
len(train_labels)
```
Each label is an integer between 0 and 9:
```
train_labels
```
There are 10,000 images in the test set. Again, each image is represented as 28 x 28 pixels:
```
test_images.shape
```
And the test set contains 10,000 images labels:
```
len(test_labels)
```
## Preprocess the data
The data must be preprocessed before training the network. If you inspect the first image in the training set, you will see that the pixel values fall in the range of 0 to 255:
```
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.gca().grid(False)
```
We will scale these values to a range of 0 to 1 before feeding to the neural network model. For this, cast the datatype of the image components from and integer to a float, and divide by 255. Here's the function to preprocess the images:
It's important that the *training set* and the *testing set* are preprocessed in the same way:
```
train_images = train_images / 255.0
test_images = test_images / 255.0
```
Display the first 25 images from the *training set* and display the class name below each image. Verify that the data is in the correct format and we're ready to build and train the network.
```
import matplotlib.pyplot as plt
%matplotlib inline
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid('off')
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
```
## Build the model
Building the neural network requires configuring the layers of the model, then compiling the model.
### Setup the layers
The basic building block of a neural network is the *layer*. Layers extract representations from the data fed into them. And, hopefully, these representations are more meaningful for the problem at hand.
Most of deep learning consists of chaining together simple layers. Most layers, like `tf.keras.layers.Dense`, have parameters that are learned during training.
```
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation=tf.nn.relu),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
```
The first layer in this network, `tf.keras.layers.Flatten`, transforms the format of the images from a 2d-array (of 28 by 28 pixels), to a 1d-array of 28 * 28 = 784 pixels. Think of this layer as unstacking rows of pixels in the image and lining them up. This layer has no parameters to learn, it only reformats the data.
After the pixels are flattened, the network consists of a sequence of two `tf.keras.layers.Dense` layers. These are densely-connected, or fully-connected, neural layers. The first `Dense` layer has 128 nodes, or neurons. The second (and last) layer is a 10-node *softmax* layer—this returns an array of 10 probability scores that sum to 1. Each node contains a score that indicates the probability that the current image belongs to one of the 10 digit classes.
### Compile the model
Before the model is ready for training, it needs a few more settings. These are added during the model's *compile* step:
* *Loss function* —This measures how accurate the model is during training. We want to minimize this function to "steer" the model in the right direction.
* *Optimizer* —This is how the model is updated based on the data it sees and its loss function.
* *Metrics* —Used to monitor the training and testing steps. The following example uses *accuracy*, the fraction of the images that are correctly classified.
```
model.compile(optimizer=tf.train.AdamOptimizer(),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
```
## Train the model
Training the neural network model requires the following steps:
1. Feed the training data to the model—in this example, the `train_images` and `train_labels` arrays.
2. The model learns to associate images and labels.
3. We ask the model to make predictions about a test set—in this example, the `test_images` array. We verify that the predictions match the labels from the `test_labels` array..
To start training, call the `model.fit` method—the model is "fit" to the training data:
```
model.fit(train_images, train_labels, epochs=5)
```
As the model trains, the loss and accuracy metrics are displayed. This model reaches an accuracy of about 0.88 (or 88%) on the training data.
## Evaluate accuracy
Next, compare how the model performs on the test dataset:
```
test_loss, test_acc = model.evaluate(test_images, test_labels)
print('Test accuracy:', test_acc)
```
It turns out, the accuracy on the test dataset is a little less than the accuracy on the training dataset. This gap between training accuracy and test accuracy is an example of *overfitting*. Overfitting is when a machine learning model performs worse on new data than on their training data.
## Make predictions
With the model trained, we can use it to make predictions about some images.
```
predictions = model.predict(test_images)
```
Here, the model has predicted the label for each image in the testing set. Let's take a look at the first prediction:
```
predictions[0]
```
A prediction is an array of 10 numbers. These describe the "confidence" of the model that the image corresponds to each of the 10 different articles of clothing. We can see see which label has the highest confidence value:
```
np.argmax(predictions[0])
```
So the model is most confident that this image is an ankle boot, or `class_names[9]`. And we can check the test label to see this is correct:
```
test_labels[0]
```
Let's plot several images with their predictions. Correct prediction labels are green and incorrect prediction labels are red.
```
# Plot the first 25 test images, their predicted label, and the true label
# Color correct predictions in green, incorrect predictions in red
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid('off')
plt.imshow(test_images[i], cmap=plt.cm.binary)
predicted_label = np.argmax(predictions[i])
true_label = test_labels[i]
if predicted_label == true_label:
color = 'green'
else:
color = 'red'
plt.xlabel("{} ({})".format(class_names[predicted_label],
class_names[true_label]),
color=color)
```
Finally, use the trained model to make a prediction about a single image.
```
# Grab an image from the test dataset
img = test_images[0]
print(img.shape)
```
`tf.keras` models are optimized to make predictions on a *batch*, or collection, of examples at once. So even though we're using a single image, we need to add it to a list:
```
# Add the image to a batch where it's the only member.
img = (np.expand_dims(img,0))
print(img.shape)
```
Now predict the image:
```
predictions = model.predict(img)
print(predictions)
```
`model.predict` returns a list of lists, one for each image in the batch of data. Grab the predictions for our (only) image in the batch:
```
prediction = predictions[0]
np.argmax(prediction)
```
And, as before, the model predicts a label of 9.
| github_jupyter |
# Prepare Dataset for RoBERTa
# Feature Transformation with Amazon a SageMaker Processing Job and Scikit-Learn
Typically a machine learning (ML) process consists of few steps. First, gathering data with various ETL jobs, then pre-processing the data, featurizing the dataset by incorporating standard techniques or prior knowledge, and finally training an ML model using an algorithm.
Often, distributed data processing frameworks such as Scikit-Learn are used to pre-process data sets in order to prepare them for training. In this notebook we'll use Amazon SageMaker Processing, and leverage the power of Scikit-Learn in a managed SageMaker environment to run our processing workload.
# Setup Environment
Let's start by specifying:
* The S3 bucket and prefixes that you use for training and model data. Use the default bucket specified by the Amazon SageMaker session.
* The IAM role ARN used to give processing and training access to the dataset.
```
import sagemaker
import boto3
sagemaker_session = sagemaker.Session()
role = sagemaker.get_execution_role()
bucket = sagemaker_session.default_bucket()
region = boto3.Session().region_name
sm = boto3.Session().client(service_name='sagemaker', region_name=region)
```
# Retrieve S3 Path for Raw Input Data
```
%store -r raw_input_data_s3_uri
print(raw_input_data_s3_uri)
!aws s3 ls $raw_input_data_s3_uri/
```
# Run the Processing Job using Amazon SageMaker
Next, use the Amazon SageMaker Python SDK to submit a processing job using our custom python script.
# Review the Processing Script
```
!pygmentize ./src/prepare_data.py
```
Run this script as a processing job. You also need to specify one `ProcessingInput` with the `source` argument of the Amazon S3 bucket and `destination` is where the script reads this data from `/opt/ml/processing/input` (inside the Docker container.) All local paths inside the processing container must begin with `/opt/ml/processing/`.
Also give the `run()` method a `ProcessingOutput`, where the `source` is the path the script writes output data to. For outputs, the `destination` defaults to an S3 bucket that the Amazon SageMaker Python SDK creates for you, following the format `s3://sagemaker-<region>-<account_id>/<processing_job_name>/output/<output_name>/`. You also give the `ProcessingOutput` value for `output_name`, to make it easier to retrieve these output artifacts after the job is run.
The arguments parameter in the `run()` method are command-line arguments in our `prepare_data.py` script.
Note that we sharding the data using `ShardedByS3Key` to spread the transformations across all worker nodes in the cluster.
# Set the Processing Job Hyper-Parameters
```
processing_instance_type='ml.c5.2xlarge'
processing_instance_count=2
train_split_percentage=0.90
validation_split_percentage=0.05
test_split_percentage=0.05
balance_dataset=True
from sagemaker.sklearn.processing import SKLearnProcessor
processor = SKLearnProcessor(framework_version='0.20.0',
role=role,
instance_type=processing_instance_type,
instance_count=processing_instance_count,
max_runtime_in_seconds=7200)
from sagemaker.processing import ProcessingInput, ProcessingOutput
processor.run(code='./src/prepare_data.py',
inputs=[
ProcessingInput(source=raw_input_data_s3_uri,
destination='/opt/ml/processing/input/data/',
s3_data_distribution_type='ShardedByS3Key')
],
outputs=[
ProcessingOutput(s3_upload_mode='EndOfJob',
output_name='sentiment-train',
source='/opt/ml/processing/output/sentiment/train'),
ProcessingOutput(s3_upload_mode='EndOfJob',
output_name='sentiment-validation',
source='/opt/ml/processing/output/sentiment/validation'),
ProcessingOutput(s3_upload_mode='EndOfJob',
output_name='sentiment-test',
source='/opt/ml/processing/output/sentiment/test')
],
arguments=['--train-split-percentage', str(train_split_percentage),
'--validation-split-percentage', str(validation_split_percentage),
'--test-split-percentage', str(test_split_percentage),
'--balance-dataset', str(balance_dataset)
],
logs=True,
wait=False)
scikit_processing_job_name = processor.jobs[-1].describe()['ProcessingJobName']
print(scikit_processing_job_name)
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="https://console.aws.amazon.com/sagemaker/home?region={}#/processing-jobs/{}">Processing Job</a></b>'.format(region, scikit_processing_job_name)))
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="https://console.aws.amazon.com/cloudwatch/home?region={}#logStream:group=/aws/sagemaker/ProcessingJobs;prefix={};streamFilter=typeLogStreamPrefix">CloudWatch Logs</a> After About 5 Minutes</b>'.format(region, scikit_processing_job_name)))
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="https://s3.console.aws.amazon.com/s3/buckets/{}/{}/?region={}&tab=overview">S3 Output Data</a> After The Processing Job Has Completed</b>'.format(bucket, scikit_processing_job_name, region)))
```
# Monitor the Processing Job
```
running_processor = sagemaker.processing.ProcessingJob.from_processing_name(processing_job_name=scikit_processing_job_name,
sagemaker_session=sagemaker_session)
processing_job_description = running_processor.describe()
print(processing_job_description)
running_processor.wait(logs=False)
```
# _Please Wait Until the ^^ Processing Job ^^ Completes Above._
# Inspect the Processed Output Data
Take a look at a few rows of the transformed dataset to make sure the processing was successful.
```
processing_job_description = running_processor.describe()
output_config = processing_job_description['ProcessingOutputConfig']
for output in output_config['Outputs']:
if output['OutputName'] == 'sentiment-train':
processed_train_data_s3_uri = output['S3Output']['S3Uri']
if output['OutputName'] == 'sentiment-validation':
processed_validation_data_s3_uri = output['S3Output']['S3Uri']
if output['OutputName'] == 'sentiment-test':
processed_test_data_s3_uri = output['S3Output']['S3Uri']
print(processed_train_data_s3_uri)
print(processed_validation_data_s3_uri)
print(processed_test_data_s3_uri)
!aws s3 ls $processed_train_data_s3_uri/
!aws s3 ls $processed_validation_data_s3_uri/
!aws s3 ls $processed_test_data_s3_uri/
```
# Pass Variables to the Next Notebook(s)
```
%store raw_input_data_s3_uri
%store train_split_percentage
%store validation_split_percentage
%store test_split_percentage
%store balance_dataset
%store processed_train_data_s3_uri
%store processed_validation_data_s3_uri
%store processed_test_data_s3_uri
%store
```
# Release Resources
```
%%html
<p><b>Shutting down your kernel for this notebook to release resources.</b></p>
<button class="sm-command-button" data-commandlinker-command="kernelmenu:shutdown" style="display:none;">Shutdown Kernel</button>
<script>
try {
els = document.getElementsByClassName("sm-command-button");
els[0].click();
}
catch(err) {
// NoOp
}
</script>
%%javascript
try {
Jupyter.notebook.save_checkpoint();
Jupyter.notebook.session.delete();
}
catch(err) {
// NoOp
}
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Distributed PyTorch with DistributedDataParallel
In this tutorial, you will train a PyTorch model on the [CIFAR-10](https://www.cs.toronto.edu/~kriz/cifar.html) dataset using distributed training with PyTorch's `DistributedDataParallel` module across a GPU cluster.
## Prerequisites
* If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, go through the [Configuration](../../../../configuration.ipynb) notebook to install the Azure Machine Learning Python SDK and create an Azure ML `Workspace`
```
# Check core SDK version number
import azureml.core
print("SDK version:", azureml.core.VERSION)
```
## Diagnostics
Opt-in diagnostics for better experience, quality, and security of future releases.
```
from azureml.telemetry import set_diagnostics_collection
set_diagnostics_collection(send_diagnostics=True)
```
## Initialize workspace
Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`.
```
from azureml.core.workspace import Workspace
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep='\n')
```
## Create or attach existing AmlCompute
You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, we use Azure ML managed compute ([AmlCompute](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)) for our remote training compute resource. Specifically, the below code creates an `STANDARD_NC6` GPU cluster that autoscales from `0` to `4` nodes.
**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace, this code will skip the creation process.
As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# choose a name for your cluster
cluster_name = 'gpu-cluster'
try:
compute_target = ComputeTarget(workspace=ws, name=cluster_name)
print('Found existing compute target.')
except ComputeTargetException:
print('Creating a new compute target...')
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6',
max_nodes=4)
# create the cluster
compute_target = ComputeTarget.create(ws, cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# use get_status() to get a detailed status for the current AmlCompute.
print(compute_target.get_status().serialize())
```
The above code creates GPU compute. If you instead want to create CPU compute, provide a different VM size to the `vm_size` parameter, such as `STANDARD_D2_V2`.
## Prepare dataset
Prepare the dataset used for training. We will first download and extract the publicly available CIFAR-10 dataset from the cs.toronto.edu website and then create an Azure ML FileDataset to use the data for training.
### Download and extract CIFAR-10 data
```
import urllib
import tarfile
import os
url = 'https://www.cs.toronto.edu/~kriz/cifar-10-python.tar.gz'
filename = 'cifar-10-python.tar.gz'
data_root = 'cifar-10'
filepath = os.path.join(data_root, filename)
if not os.path.isdir(data_root):
os.makedirs(data_root, exist_ok=True)
urllib.request.urlretrieve(url, filepath)
with tarfile.open(filepath, "r:gz") as tar:
tar.extractall(path=data_root)
os.remove(filepath) # delete tar.gz file after extraction
```
### Create Azure ML dataset
The `upload_directory` method will upload the data to a datastore and create a FileDataset from it. In this tutorial we will use the workspace's default datastore.
```
from azureml.core import Dataset
datastore = ws.get_default_datastore()
dataset = Dataset.File.upload_directory(
src_dir=data_root, target=(datastore, data_root)
)
```
## Train model on the remote compute
Now that we have the AmlCompute ready to go, let's run our distributed training job.
### Create a project directory
Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script and any additional files your training script depends on.
```
project_folder = './pytorch-distr'
os.makedirs(project_folder, exist_ok=True)
```
### Prepare training script
Now you will need to create your training script. In this tutorial, the script for distributed training on CIFAR-10 is already provided for you at `train.py`. In practice, you should be able to take any custom PyTorch training script as is and run it with Azure ML without having to modify your code.
Once your script is ready, copy the training script `train.py` into the project directory.
```
import shutil
shutil.copy('train.py', project_folder)
```
### Create an experiment
Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed PyTorch tutorial.
```
from azureml.core import Experiment
experiment_name = 'pytorch-distr'
experiment = Experiment(ws, name=experiment_name)
```
### Create an environment
In this tutorial, we will use one of Azure ML's curated PyTorch environments for training. [Curated environments](https://docs.microsoft.com/azure/machine-learning/how-to-use-environments#use-a-curated-environment) are available in your workspace by default. Specifically, we will use the PyTorch 1.6 GPU curated environment.
```
from azureml.core import Environment
pytorch_env = Environment.get(ws, name='AzureML-PyTorch-1.6-GPU')
```
### Configure the training job
To launch a distributed PyTorch job on Azure ML, you have two options:
1. Per-process launch - specify the total # of worker processes (typically one per GPU) you want to run, and
Azure ML will handle launching each process.
2. Per-node launch with [torch.distributed.launch](https://pytorch.org/docs/stable/distributed.html#launch-utility) - provide the `torch.distributed.launch` command you want to
run on each node.
For more information, see the [documentation](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-train-pytorch#distributeddataparallel).
Both options are shown below.
#### Per-process launch
To use the per-process launch option in which Azure ML will handle launching each of the processes to run your training script,
1. Specify the training script and arguments
2. Create a `PyTorchConfiguration` and specify `node_count` and `process_count`. The `process_count` is the total number of processes you want to run for the job; this should typically equal the # of GPUs available on each node multiplied by the # of nodes. Since this tutorial uses the `STANDARD_NC6` SKU, which has one GPU, the total process count for a 2-node job is `2`. If you are using a SKU with >1 GPUs, adjust the `process_count` accordingly.
Azure ML will set the `MASTER_ADDR`, `MASTER_PORT`, `NODE_RANK`, `WORLD_SIZE` environment variables on each node, in addition to the process-level `RANK` and `LOCAL_RANK` environment variables, that are needed for distributed PyTorch training.
```
from azureml.core import ScriptRunConfig
from azureml.core.runconfig import PyTorchConfiguration
# create distributed config
distr_config = PyTorchConfiguration(process_count=2, node_count=2)
# create args
args = ["--data-dir", dataset.as_download(), "--epochs", 25]
# create job config
src = ScriptRunConfig(source_directory=project_folder,
script='train.py',
arguments=args,
compute_target=compute_target,
environment=pytorch_env,
distributed_job_config=distr_config)
```
#### Per-node launch with `torch.distributed.launch`
If you would instead like to use the PyTorch-provided launch utility `torch.distributed.launch` to handle launching the worker processes on each node, you can do so as well.
1. Provide the launch command to the `command` parameter of ScriptRunConfig. For PyTorch jobs Azure ML will set the `MASTER_ADDR`, `MASTER_PORT`, and `NODE_RANK` environment variables on each node, so you can simply just reference those environment variables in your command. If you are using a SKU with >1 GPUs, adjust the `--nproc_per_node` argument accordingly.
2. Create a `PyTorchConfiguration` and specify the `node_count`. You do not need to specify the `process_count`; by default Azure ML will launch one process per node to run the `command` you provided.
Uncomment the code below to configure a job with this method.
```
'''
from azureml.core import ScriptRunConfig
from azureml.core.runconfig import PyTorchConfiguration
# create distributed config
distr_config = PyTorchConfiguration(node_count=2)
# define command
launch_cmd = ["python -m torch.distributed.launch --nproc_per_node 1 --nnodes 2 " \
"--node_rank $NODE_RANK --master_addr $MASTER_ADDR --master_port $MASTER_PORT --use_env " \
"train.py --data-dir", dataset.as_download(), "--epochs 25"]
# create job config
src = ScriptRunConfig(source_directory=project_folder,
command=launch_cmd,
compute_target=compute_target,
environment=pytorch_env,
distributed_job_config=distr_config)
'''
```
### Submit job
Run your experiment by submitting your `ScriptRunConfig` object. Note that this call is asynchronous.
```
run = experiment.submit(src)
print(run)
```
### Monitor your run
You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes. You can see that the widget automatically plots and visualizes the loss metric that we logged to the Azure ML run.
```
from azureml.widgets import RunDetails
RunDetails(run).show()
```
Alternatively, you can block until the script has completed training before running more code.
```
run.wait_for_completion(show_output=True) # this provides a verbose log
```
| github_jupyter |
## Gaussian Mixture Model for Density Estimation
This notebook demonstrates Gaussian mixture models (GMMs) in 2D. We can see that the GMM can model quite complicated distributions, but in certain situations be unnecessarily parameterised.
The approximation of a large GMM (i.e. with a large number of components) with a smaller one is a challenging task. This can be achieved using the well-known Expectation Maximisation algorithm for GMMs as applied to a _sample_ from such a model, but no analytic optimisation procedure is known. A poor-man's version might be the Variational Boosting algorithm (Miller, Foti, and Adams - ICML 2017) which greedily fits Gaussian components via KL minimisation, but this is reasonably expensive and by no means optimal.
The below code generates a random Gaussian mixture, draws samples from it, and then fits a smaller GMM to this synthetic data.
```
import numpy as np
from sklearn.mixture import GaussianMixture
import matplotlib.pyplot as plt
def gaussian_2D_level_curve(mu, sigma, alpha=2, ncoods=100, plot=True):
# (Alex Bird 2016)
# (Ported from my matlab utils / pyalexutil)
assert isinstance(mu, np.ndarray) and isinstance(sigma, np.ndarray), "mu/sigma must be numpy arrays."
assert mu.shape == (2, ), 'mu must be vector in R^2'
assert sigma.shape == (2, 2), 'sigma must be 2x2 array'
U, S, V = np.linalg.svd(sigma)
sd = np.sqrt(S)
coods = np.linspace(0, 2 * np.pi, ncoods)
coods = np.vstack((sd[0] * np.cos(coods), sd[1] * np.sin(coods))) * alpha
# project onto basis of ellipse
coods = (V @ coods).T
# add mean
coods += mu
if plot:
plt.plot(*coods.T)
return coods
def gen_mix_mvn(n_components=30, d=2, n_samples=400):
# Generate random multivariate Gaussians
pi = np.random.dirichlet([0.8]*n_components)
mu = np.random.multivariate_normal(np.zeros(d), np.eye(d)*25, size=(n_components))
mu = np.random.rand(n_components, d) * 10 - 5
sigma = np.zeros((n_components, d, d))
for n in range(n_components):
_tmpmat = np.random.rand(d,d)
Q, _junk = np.linalg.qr(_tmpmat)
lam = np.random.exponential(1, d)
sigma[n] = Q @ np.diag(lam) @ Q.T
# Draw samples
z = np.random.multinomial(n_samples, pi)
smps = np.zeros((n_samples, d))
indexes = np.stack((np.cumsum(np.concatenate(([0], z[:-1]))),
np.cumsum(z)), axis=1)
for ixs, n, m, s in zip(indexes, z, mu, sigma):
smps[slice(*ixs)] = np.random.multivariate_normal(m, s, size=n)
return smps, (pi, mu, sigma)
n_components = 30 # number of Gaussians in original mixture
n_approx = 5 # number of Gaussians to approximate with
n_samples = 800 # number of samples to draw from mixture for visualisation.
# generate random Gaussian mixture
smps, pars = gen_mix_mvn(30, n_samples=n_samples)
f, axs = plt.subplots(2, 2)
f.set_size_inches(12,12)
# plot level curves of random Gaussian mixture
axs[0,0].set_title('Original Gaussian Mixture (alpha=weight)')
for pi, m, s in zip(*pars):
axs[0,0].plot(*gaussian_2D_level_curve(m, s, plot=False).T, alpha=pi/max(pars[0]))
# sample from this Gaussian mixture
axs[0,1].scatter(*smps.T, alpha=0.4)
axs[0,1].set_title('Sample from Original Density')
# fit a new Gaussian mixture with 5 components:
gmm = GaussianMixture(n_components=n_approx)
gmm.fit(smps)
axs[1,0].set_title('Approximated Gaussian Mixture (alpha=weight)')
maxw = max(gmm.weights_)
for pi, m, s in zip(gmm.weights_ ,gmm.means_, gmm.covariances_):
axs[1,0].plot(*gaussian_2D_level_curve(m, s, plot=False).T, alpha=pi/maxw)
# sample from this Gaussian mixture
smps_approx = gmm.sample(n_samples)[0]
axs[1,1].scatter(*smps_approx.T, alpha=0.4)
axs[1,1].set_title('Sample from Approximated Density');
```
#### ^^ Above
[**<span style='color:blue'>Top left</span>**] The original (generated) Gaussian mixture. The 2 sigma level curves are drawn (note that this corresponds only to approx 63% of density in 2D) and their weight in the mixture denoted by the alpha value (transparency). Often a fairly large number of low weight components are present (this occurs e.g. in Gaussian Sum filtering).<br>
[**<span style='color:blue'>Top right</span>**] A sample (default: 800) points are drawn from this mixture. The generative model is to choose the component proportionally to the weights, and then draw a sample from the relevant Gaussian. In a certain sense the sample appears less complicated than the original mixture.<br>
[**<span style='color:blue'>Bottom left</span>**] The fitted GMM (using _sklearn_'s version of the EM algorithm). This is implemented reasonably well and converges quickly. However, this may take some time in high dimensions - it is an iterative co-ordinate descent algorithm and is particularly prone to slow convergence if boundaries between components are ill-defined.<br>
[**<span style='color:blue'>Bottom right</span>**] A sample from the approximate GMM. This often appears superficially very similar to the original density and gives credence to the idea that small GMMs are capable of capturing the salient features of complicated densities.
### Comparison with standard Gaussian Sum collapse heuristics
```
ixs_top4 = np.flip(np.argsort(pars[0]), axis=0)[:n_approx-1]
ixs_other = np.array(list(set(np.argsort(pars[0])) - set(ixs_top4)))
pi_approx = np.concatenate((pars[0][ixs_top4], [sum(pars[0][ixs_other])]))
mu_approx = pars[1][ixs_top4]
mu_other = np.dot(pars[0][ixs_other], pars[1][ixs_other])/pi_approx[-1]
sigma_approx = pars[2][ixs_top4]
sigma_other = -np.outer(mu_other, mu_other)
for i in ixs_other:
sigma_other += pars[0][i] * (pars[2][i] + np.outer(pars[1][i], pars[1][i]))
mu_approx = np.concatenate((mu_approx, mu_other[None,:]), axis=0)
sigma_approx = np.concatenate((sigma_approx, sigma_other[None,:]), axis=0)
f, axs = plt.subplots(1, 2)
f.set_size_inches(12,6)
# plot level curves of random Gaussian mixture
axs[0].set_title('Heuristic Approximate Gaussian Mixture (alpha=weight)')
for pi, m, s in zip(pi_approx, mu_approx, sigma_approx):
axs[0].plot(*gaussian_2D_level_curve(m, s, plot=False).T, alpha=pi/max(pi_approx))
# sample from this Gaussian mixture
gmm.means_ = mu_approx; gmm.covariances_ = sigma_approx; gmm.weights_ = pi_approx
smps_approx = gmm.sample(n_samples)[0]
axs[1].scatter(*smps_approx.T, alpha=0.4)
axs[1].set_title('Sample from Heuristic Approximation');
```
| github_jupyter |
```
#hide
from dash_oop_components.core import *
```
# Tracking state of your app in url querystrings
> instructions on how to track the state of your dashboard in the url querystring
## Make shareable dashboards by tracking state in url querystrings
For a lot of analytical web apps it can be super useful to be able to share the state of a dashboard with others through a url. Imagine you have done a particular analysis on a particular tab, setting certain dropdowns and toggles and you wish to share these with a co-worker.
You could tell them to go to the dashboard with instructions to set the exact same dropdowns and toggles. But it would be much easier if you can simply send a url that rebuild the dashboard exactly as you saw it!
This can be done by storing the state of the dashboard in the querystring:

## Tracking state with `dash_oop_components`
Thanks to the modular nature and tree structure of `DashComponents` it is relatively straightforward to keep track of
which elements should be tracked in the url querystring, and rebuild the page in accordance with the state of the querystring.
An example dashboard that demonstrates how to build a dashboard with querystrings included can be found at [github.com/oegedijk/dash_oop_demo](https://github.com/oegedijk/dash_oop_demo) and has been deployed to [https://dash-oop-demo.herokuapp.com/](https://dash-oop-demo.herokuapp.com/)
## Basic summary instructions:
In order to add querystring support to your app all you need is to:
1. Pass `querystrings=True` parameters to `DashApp`
2. Change the `def layout(self)` method to `def layout(self, params=None)`
3. Inside your `DashComponents` wrap the elements that you want to track in `self.querystring(params)(...)`:
- i.e. change
```python
dcc.Input(id='input-'+self.name)
```
to
```python
self.querystring(params)(dcc.Input)(id='input-'+self.name')
```
4. pass down `params` to all subcomponent layouts:
```python
def layout(self, params=None):
return html.Div([self.subcomponent.layout(params)])
```
**note:** it is important to assign a proper `.name` to components with querystring elements, as otherwise the elements will get a different random uuid `id` each time you reboot the dashboard, breaking old querystrings.
### Step 1: Turning on querystrings in Dashapp
In order to turn on the tracking of querystrings you need to start `DashApp`
with the `querystrings=True` parameter, e.g.:
```python
dashboard = CovidDashboard(plot_factory)
app = DashApp(dashboard, querystrings=True, bootstrap=dbc.themes.FLATLY)
```
### Step 2: Building `DashComponent` with `layout(params)` and `self.querystring()`
The example [dashboard](https://github.com/oegedijk/dash_oop_demo) consists of four tabs that each contain the layout of a `CovidComposite` subcomponent:
- `self.europe`: a tab with only european countries
- `self.asia`: a tab with only Asian countries
- `self.cases_only`: a tab with only cases data (for the whole world)
- `self.deaths_only`: a tab with only deaths data (for the whole world)
In order to keep track of an attribute of a layout element we simply wrap it inside a `self.querystring()(element_func)(params)` wrapper:
```python
self.querystring(params)(dcc.Tabs)(id='tabs', ...)`
```
This will make sure that the `value` attribute of the `dcc.Tabs` element with `id='tabs'` is tracked in the querystring, so that users will start on the same tab when you send them a link.
Other querystring parameters get tracked inside the subcomponent definition of `DashComposite`. In order to make sure that these subcomponents also receive the `params` we need to pass those params down to the layout of our subcomponents as well:
```python
dcc.Tab(..., children=self.europe.layout(params))
dcc.Tab(..., children=self.asia.layout(params))
dcc.Tab(..., children=self.cases_only.layout(params))
dcc.Tab(..., children=self.deaths_only.layout(params))
```
Note that we set the `name` of the tabs to `"eur"`, `"asia"`, `"cases"` and `"deaths"`
**Full definition of `CovidDashboard`:**
```python
class CovidDashboard(DashComponent):
def __init__(self, plot_factory,
europe_countries = ['Italy', 'Spain', 'Germany', 'France',
'United_Kingdom', 'Switzerland', 'Netherlands',
'Belgium', 'Austria', 'Portugal', 'Norway'],
asia_countries = ['China', 'Vietnam', 'Malaysia', 'Philippines',
'Taiwan', 'Myanmar', 'Thailand', 'South_Korea', 'Japan']):
super().__init__(title="Covid Dashboard")
self.europe = CovidComposite(self.plot_factory, "Europe",
include_countries=self.europe_countries, name="eur")
self.asia = CovidComposite(self.plot_factory, "Asia",
include_countries=self.asia_countries, name="asia")
self.cases_only = CovidComposite(self.plot_factory, "Cases Only",
metric='cases', hide_metric_dropdown=True,
countries=['China', 'Italy', 'Brazil'], name="cases")
self.deaths_only = CovidComposite(self.plot_factory, "Deaths Only",
metric='deaths', hide_metric_dropdown=True,
countries=['China', 'Italy', 'Brazil'], name="deaths")
def layout(self, params=None):
return dbc.Container([
dbc.Row([
html.H1("Covid Dashboard"),
]),
dbc.Row([
dbc.Col([
self.querystring(params)(dcc.Tabs)(id="tabs", value=self.europe.name,
children=[
dcc.Tab(label=self.europe.title,
id=self.europe.name,
value=self.europe.name,
children=self.europe.layout(params)),
dcc.Tab(label=self.asia.title,
id=self.asia.name,
value=self.asia.name,
children=self.asia.layout(params)),
dcc.Tab(label=self.cases_only.title,
id=self.cases_only.name,
value=self.cases_only.name,
children=self.cases_only.layout(params)),
dcc.Tab(label=self.deaths_only.title,
id=self.deaths_only.name,
value=self.deaths_only.name,
children=self.deaths_only.layout(params)),
]),
])
])
], fluid=True)
```
## Step 3: tracking parameters in subcomponents:
A `CovidComposite` `DashComponent` consists of a `CovidTimeSeries`, a `CovidPieChart` and two dropdowns for metric and country selection. The value of the dropdowns get passed to the corresponding dropdowns of the subcomponents, which are hidden through the config params.
We would like to keep track of the state of these dropdowns so we wrap them inside a `self.querystring()`:
For the metric dropdown:
```python
self.querystring(params)(dcc.Dropdown)(id='dashboard-metric-dropdown-'+self.name, ...)
```
For the country dropdown:
```python
self.querystring(params)(dcc.Dropdown)(id='dashboard-country-dropdown-'+self.name, ...)
```
And we also make sure that parameters can be passed down the layout with
```
def layout(self, params=None):
...
```
**Full definition of `CovidComposite`:**
```python
class CovidComposite(DashComponent):
def __init__(self, plot_factory, title="Covid Analysis",
hide_country_dropdown=False,
include_countries=None, countries=None,
hide_metric_dropdown=False,
include_metrics=None, metric='cases', name=None):
super().__init__(title=title)
if not self.include_countries:
self.include_countries = self.plot_factory.countries
if not self.countries:
self.countries = self.include_countries
if not self.include_metrics:
self.include_metrics = self.plot_factory.metrics
if not self.metric:
self.metric = self.include_metrics[0]
self.timeseries = CovidTimeSeries(
plot_factory,
hide_country_dropdown=True, countries=self.countries,
hide_metric_dropdown=True, metric=self.metric)
self.piechart = CovidPieChart(
plot_factory,
hide_country_dropdown=True, countries=self.countries,
hide_metric_dropdown=True, metric=self.metric)
def layout(self, params=None):
return dbc.Container([
dbc.Row([
dbc.Col([
html.H1(self.title),
self.make_hideable(
self.querystring(params)(dcc.Dropdown)(
id='dashboard-metric-dropdown-'+self.name,
options=[{'label': metric, 'value': metric} for metric in self.include_metrics],
value=self.metric,
), hide=self.hide_metric_dropdown),
self.make_hideable(
self.querystring(params)(dcc.Dropdown)(
id='dashboard-country-dropdown-'+self.name,
options=[{'label': metric, 'value': metric} for metric in self.include_countries],
value=self.countries,
multi=True,
), hide=self.hide_country_dropdown),
], md=6),
], justify="center"),
dbc.Row([
dbc.Col([
self.timeseries.layout(),
], md=6),
dbc.Col([
self.piechart.layout(),
], md=6)
])
], fluid=True)
def component_callbacks(self, app):
@app.callback(
Output('timeseries-country-dropdown-'+self.timeseries.name, 'value'),
Output('piechart-country-dropdown-'+self.piechart.name, 'value'),
Input('dashboard-country-dropdown-'+self.name, 'value'),
)
def update_timeseries_plot(countries):
return countries, countries
@app.callback(
Output('timeseries-metric-dropdown-'+self.timeseries.name, 'value'),
Output('piechart-metric-dropdown-'+self.piechart.name, 'value'),
Input('dashboard-metric-dropdown-'+self.name, 'value'),
)
def update_timeseries_plot(metric):
return metric, metric
```
## Addendum: Tracking querystring params of current tab only
When you define dashboard with lots of tabs, lots of components and lots of elements, the size of the querystring can explode rapidly, resulting in clumsy long urls to copy-paste. One solution if to only keep track of the parameters in the current open tab.
The downside is that the rest of the dashboard will take default values, but the upside is significantly smaller querystrings.
In order to implement this, you can make sure of the `DashComponentTabs` as a stand-in replacement for `dcc.Tabs`.
You simply replace
```python
self.querystring(params)(dcc.Tabs)(id="tabs", value=self.europe.name,
children=[
dcc.Tab(label=self.europe.title,
id=self.europe.name,
value=self.europe.name,
children=self.europe.layout(params)),
dcc.Tab(label=self.asia.title,
id=self.asia.name,
value=self.asia.name,
children=self.asia.layout(params)),
dcc.Tab(label=self.cases_only.title,
id=self.cases_only.name,
value=self.cases_only.name,
children=self.cases_only.layout(params)),
dcc.Tab(label=self.deaths_only.title,
id=self.deaths_only.name,
value=self.deaths_only.name,
children=self.deaths_only.layout(params)),
]),
```
with
```python
self.querystring(params)(DashComponentTabs)(id="tabs",
tabs=[self.europe, self.asia, self.cases_only, self.deaths_only],
params=params, component=self, single_tab_querystrings=True)
```
And automatically all parameters from tabs other than the current tab will be excluded from the url querystring
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/JavaScripts/Image/PixelLonLat.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/JavaScripts/Image/PixelLonLat.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/JavaScripts/Image/PixelLonLat.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as geemap
except:
import geemap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google MapS`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/basemaps.py) can be added using the `Map.add_basemap()` function.
```
Map = geemap.Map(center=[40,-100], zoom=4)
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
# Draws 60 lat/long lines per degree using the pixelLonLat() function.
# Create an image in which the value of each pixel is its
# coordinates in minutes.
img = ee.Image.pixelLonLat().multiply(60.0)
# Get the decimal part and check if it's less than a small delta.
img = img.subtract(img.floor()).lt(0.05)
# The pixels less than the delta are the grid, in both directions.
grid = img.select('latitude').Or(img.select('longitude'))
# Draw the grid.
Map.setCenter(-122.09228, 37.42330, 12)
Map.addLayer(grid.updateMask(grid), {'palette': '008000'}, 'Graticule')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
# Deep Reinforcement Learning <em> in Action </em>
## N-Armed Bandits
### Chapter 2
```
import numpy as np
import torch as th
from torch.autograd import Variable
from matplotlib import pyplot as plt
import random
%matplotlib inline
```
This defines the main contextual bandit class we'll be using as our environment/simulator to train a neural network.
```
class ContextBandit:
def __init__(self, arms=10):
self.arms = arms
self.init_distribution(arms)
self.update_state()
def init_distribution(self, arms):
# Num states = Num Arms to keep things simple
self.bandit_matrix = np.random.rand(arms,arms)
#each row represents a state, each column an arm
def reward(self, prob):
reward = 0
for i in range(self.arms):
if random.random() < prob:
reward += 1
return reward
def get_state(self):
return self.state
def update_state(self):
self.state = np.random.randint(0,self.arms)
def get_reward(self,arm):
return self.reward(self.bandit_matrix[self.get_state()][arm])
def choose_arm(self, arm):
reward = self.get_reward(arm)
self.update_state()
return reward
```
Here we define our simple neural network model using PyTorch
```
def softmax(av, tau=1.12):
n = len(av)
probs = np.zeros(n)
for i in range(n):
softm = ( np.exp(av[i] / tau) / np.sum( np.exp(av[:] / tau) ) )
probs[i] = softm
return probs
def one_hot(N, pos, val=1):
one_hot_vec = np.zeros(N)
one_hot_vec[pos] = val
return one_hot_vec
arms = 10
# N is batch size; D_in is input dimension;
# H is hidden dimension; D_out is output dimension.
N, D_in, H, D_out = 1, arms, 100, arms
model = th.nn.Sequential(
th.nn.Linear(D_in, H),
th.nn.ReLU(),
th.nn.Linear(H, D_out),
th.nn.ReLU(),
)
loss_fn = th.nn.MSELoss(size_average=False)
env = ContextBandit(arms)
```
Next we define the training function, which accepts an instantiated ContextBandit object.
```
def train(env):
epochs = 5000
#one-hot encode current state
cur_state = Variable(th.Tensor(one_hot(arms,env.get_state())))
reward_hist = np.zeros(50)
reward_hist[:] = 5
runningMean = np.average(reward_hist)
learning_rate = 1e-2
optimizer = th.optim.Adam(model.parameters(), lr=learning_rate)
plt.xlabel("Plays")
plt.ylabel("Mean Reward")
for i in range(epochs):
y_pred = model(cur_state) #produce reward predictions
av_softmax = softmax(y_pred.data.numpy(), tau=2.0) #turn reward distribution into probability distribution
av_softmax /= av_softmax.sum() #make sure total prob adds to 1
choice = np.random.choice(arms, p=av_softmax) #sample an action
cur_reward = env.choose_arm(choice)
one_hot_reward = y_pred.data.numpy().copy()
one_hot_reward[choice] = cur_reward
reward = Variable(th.Tensor(one_hot_reward))
loss = loss_fn(y_pred, reward)
if i % 50 == 0:
runningMean = np.average(reward_hist)
reward_hist[:] = 0
plt.scatter(i, runningMean)
reward_hist[i % 50] = cur_reward
optimizer.zero_grad()
# Backward pass: compute gradient of the loss with respect to model
# parameters
loss.backward()
# Calling the step function on an Optimizer makes an update to its
# parameters
optimizer.step()
cur_state = Variable(th.Tensor(one_hot(arms,env.get_state())))
train(env)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/butchland/fastai_nb_explorations/blob/master/fastai_scratch_with_tpu_mnist_4_experiment2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import os
assert os.environ['COLAB_TPU_ADDR'], 'Make sure to select TPU from Edit > Notebook settings > Hardware accelerator'
!curl https://course.fast.ai/setup/colab | bash
VERSION = "20200325" #@param ["1.5" , "20200325", "nightly"]
!curl https://raw.githubusercontent.com/pytorch/xla/master/contrib/scripts/env-setup.py -o pytorch-xla-env-setup.py
!python pytorch-xla-env-setup.py --version $VERSION
!pip freeze | grep torchvision
!pip install fastcore --upgrade
!pip install fastai2 --upgrade
pip install fastai --upgrade
from google.colab import drive
drive.mount('/content/drive')
%cd /content/drive/My\ Drive/course-v4/
!pwd
!pip install -r requirements.txt
%cd nbs
!pwd
```
### Start of import libraries
```
from fastai2.vision.all import *
from utils import *
path = untar_data(URLs.MNIST_SAMPLE)
Path.BASE_PATH = path
path.ls()
```
### Import torch xla libraries
```
import torch
import torch_xla
import torch_xla.core.xla_model as xm
```
### define load data tensors in cpu
```
def load_tensors(dpath):
return torch.stack([tensor(Image.open(o))
for o in dpath.ls().sorted()]
).float()/255.
def count_images(dpath):
return len(dpath.ls())
train_x = torch.cat([load_tensors(path/'train'/'3'),
load_tensors(path/'train'/'7')]).view(-1,28*28)
valid_x = torch.cat([load_tensors(path/'valid'/'3'),
load_tensors(path/'valid'/'7')]).view(-1,28*28)
(train_x.device, valid_x.device)
(train_x.shape, valid_x.shape)
train_y = tensor([1]*count_images(path/'train'/'3') + [0]*count_images(path/'train'/'7')).unsqueeze(1)
valid_y = tensor([1]*count_images(path/'valid'/'3') + [0]*count_images(path/'valid'/'7')).unsqueeze(1)
(train_y.shape, valid_y.shape, train_y.device, valid_y.device)
train_dl = DataLoader(list(zip(train_x, train_y)),batch_size=256)
valid_dl = DataLoader(list(zip(valid_x, valid_y)), batch_size=256)
```
### Get TPU Device
```
tpu_dev = xm.xla_device()
tpu_dev
```
## Fix Model
```
torch.manual_seed(42)
np.random.seed(42)
```
### Loss function
```
# define loss function using sigmoid to return a val between 0.0 and 1
def mnist_loss_sigmoid(qpreds, qtargs):
qqpreds = qpreds.sigmoid()
return torch.where(qtargs==1, 1.-qqpreds, qqpreds).mean()
```
### Forward Pass + Back Propagation
```
# forward prop + back prop
def calc_grad(xb,yb,m):
qpreds = m(xb)
qloss = mnist_loss_sigmoid(qpreds,yb)
qloss.backward()
```
### Basic Optimizer
```
class BasicOptimizer:
def __init__(self, params,lr): self.lr, self.params = lr,list(params)
def step(self, *args, **kwargs):
for p in self.params: p.data -= p.grad.data * self.lr
def zero_grad(self, *args, **kwargs):
for p in self.params: p.grad = None
```
### Train Epoch
```
def train_epoch(qdl,qmodel,qopt, dev):
for xb,yb in qdl:
calc_grad(xb.to(dev),yb.to(dev),qmodel)
# qopt.step()
# replace optimizer step with xla device step computation
xm.optimizer_step(qopt, barrier=True)
qopt.zero_grad()
```
### Compute Metrics
```
def batch_accuracy(qpreds, qtargets):
qqpreds = qpreds.sigmoid()
correct = (qqpreds > 0.5) == qtargets
return correct.float().mean()
def validate_epoch(qmodel, qdl, dev):
accs = [batch_accuracy(qmodel(xb.to(dev)), yb.to(dev)) for xb,yb in qdl]
return round(torch.stack(accs).mean().item(),4)
def train_model(qtrain_dl, qvalid_dl, qmodel, qopt, epochs, dev):
for i in range(epochs):
train_epoch(qtrain_dl, qmodel, qopt, dev)
print(validate_epoch(qmodel, qvalid_dl, dev), end=' ')
```
### Build and Train Model
```
model = nn.Linear(28*28,1).to(tpu_dev)
optim = BasicOptimizer(model.parameters(),0.5)
# use basic Optimizer
train_model(train_dl, valid_dl, model, optim, 50, tpu_dev)
train_model(train_dl, valid_dl, model, SGD(model.parameters(),0.1), 50, tpu_dev)
simple_net = nn.Sequential(
nn.Linear(28*28,30),
nn.ReLU(),
nn.Linear(30,1)
).to(tpu_dev)
sgd_optim1 = SGD(simple_net.parameters(),0.1)
train_model(train_dl, valid_dl, simple_net, sgd_optim1, 50, tpu_dev)
resnet18_model = resnet18(pretrained=True).to(tpu_dev)
sgd_optim18 = SGD(resnet18_model.parameters(), 1e-2)
train_model(train_dl, valid_dl, resnet18_model, sgd_optim18, 1, tpu_dev)
```
| github_jupyter |
<img src="../../images/qiskit-heading.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
## _*Visualizing a Quantum Data and States*_
The latest version of this notebook is available on https://github.com/qiskit/qiskit-tutorial.
***
### Contributors
Jay Gambetta and Andrew Cross
```
# useful additional packages
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
from pprint import pprint
from scipy import linalg as la
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister
from qiskit import execute
# import state tomography functions
from qiskit.tools.visualization import plot_histogram, plot_state
from qiskit import Aer
def ghz_state(q, c, n):
# Create a GHZ state
qc = QuantumCircuit(q, c)
qc.h(q[0])
for i in range(n-1):
qc.cx(q[i], q[i+1])
return qc
def superposition_state(q, c):
# Create a Superposition state
qc = QuantumCircuit(q, c)
qc.h(q)
return qc
```
### The outcomes of a quantum circuit
In quantum physics you generally cannot measure a state without disturbing it. The act of measurement changes the state. After performing a quantum measurement, a qubit's quantum information reduces to a classical bit. In our system, as is standard, measurements are performed in the computational basis. For each qubit, the measurement either results in the value 0 if the qubit is measured in state $\left| 0\right\rangle$, or in the value 1 if the qubit is measured in state $\left| 1\right\rangle$.
In a given run of a quantum circuit that concludes with measurements of all $n$ qubits, the result will be one of the possible $n$-bit binary strings. If the experiment is run a second time, even if the measurement is perfect and has no error, the outcome may be different due to the fundamental randomness of quantum physics. The measurement results from many executions of the quantum circuit can be represented as a probability distribution over the possible outcomes. For a quantum circuit which has previously run on a backend with name `circuit`, a histogram visualizing the probability distribution can be obtained provided you have imported ```tools.visualization```. The histogram is generated by
```
plot_histogram(result.get_counts('circuit'), number)
```
The generated bar graph is simple to understand. The height of each bar represents the fraction of instances the corresponding outcome is obtained within the total number of shots on the backend. Only those outcomes that occurred at least once are displayed. The optional parameter `number` specifies the total number of bars to be displayed. All remaining probability is collected into a single bar labeled "rest". Quantum circuits for most quantum algorithm will typically have one outcome (representing the desired solution to the problem), or at most a few outcomes. Only circuits that produce superpositions of many computational states as their final state will give many outcomes, and collecting those could require an exponentially large number of measurements. To demonstrate this, we study two circuits: a GHZ state and a superposition state involving 3 qubits.
```
# Build the quantum circuit. We are going to build two circuits a GHZ over 3 qubits and a
# superpositon over all 3 qubits
n = 3 # number of qubits
q = QuantumRegister(n)
c = ClassicalRegister(n)
# quantum circuit to make a GHZ state
ghz = ghz_state(q, c, n)
# quantum circuit to make a superposition state
superposition = superposition_state(q, c)
measure_circuit = QuantumCircuit(q,c)
measure_circuit.measure(q, c)
# execute the quantum circuit
backend = Aer.get_backend('qasm_simulator') # the device to run on
circuits = [ghz+measure_circuit, superposition+measure_circuit]
job = execute(circuits, backend, shots=1000)
plot_histogram(job.result().get_counts(circuits[0]))
plot_histogram(job.result().get_counts(circuits[1]),options={'number_to_keep': 15})
```
### Method for visualizing a quantum state
For educational and debugging purposes, it is useful to visualize the quantum state given by a density matrix $\rho$. This can not be obtained from a single shot of an experiment or simulation of quantum circuit. However, in Qiskit we have also provided a unitary simulator, which provided you don't place measurements these backends will return the quantum state.
```
n = 2 # number of qubits
q = QuantumRegister(n)
c = ClassicalRegister(n)
qc = QuantumCircuit(q, c)
qc.h(q[1])
# execute the quantum circuit
backend = Aer.get_backend('statevector_simulator')
job = execute(qc, backend)
state_superposition = job.result().get_statevector(qc)
state_superposition
```
#### Pure states
A pure state $\left|\psi\right\rangle$ is an element of the Hilbert space $\mathcal{H}$. For $n$ qubits, the Hilbert space is the complex vector space $\mathbb{C}^{d}$ with dimension $d=2^n$. We denote the inner product between quantum states by $\left\langle \phi \right| \psi \rangle$. We denote the canonical set of orthonormal basis vectors spanning $\mathcal{H}$ as $\left| i\right\rangle$ where $i\in {0,...,d-1}$ and $\left\langle j \right| i \rangle =\delta_{i,j}$. This allows us to define an arbitrary pure state, state vector, or ket as
$$\left|\psi\right\rangle = \sum_{i=0}^{d-1}\psi_i \left| i\right\rangle,$$
where the $\psi_i$ are complex numbers. The dual vector, or bra, is defined as
$$\left\langle\psi\right| = \sum_{i=0}^{d-1}\psi_i^* \left\langle i\right|.$$
With this, the inner product takes the form
$$\left\langle \phi \right| \psi \rangle = \sum_{i=0}^{d-1} \phi^*_i \psi_i.$$
We require the state vector to be normalized, $\left\langle \psi \right| \psi \rangle = \sum_{i=0}^{d-1} |\psi_i|^2 =1$. As a result, $(d - 1)$ complex numbers are necessary to describe an arbitrary pure state.
```
def overlap(state1, state2):
return round(np.dot(state1.conj(), state2))
print(state_superposition)
overlap(state_superposition, state_superposition)
```
#### Operators
In order to relate the state to quantities of physical interest, we need to introduce operators. An operator is an object which maps a state from Hilbert space onto another state. Hence, in Dirac notation an operator can be written as
$$A = \sum_{i,j} a_{i,j} \left|i\right\rangle \left\langle j\right|$$
where $a_{ij}$ are complex numbers. For a given state $\left|\psi \right\rangle$, the mean or expectation value of $A$ is written as
$$\langle A \rangle = \left\langle \psi \right|A \left|\psi\right\rangle. $$
```
def expectation_value(state, Operator):
return round(np.dot(state.conj(), np.dot(Operator, state)).real)
X = np.array([[0, 1], [1, 0]])
Z = np.array([[1, 0], [0, -1]])
IZ = np.kron(np.eye(2), Z)
ZI = np.kron(Z, np.eye(2))
IX = np.kron(np.eye(2), X)
XI = np.kron(X, np.eye(2))
print("Operator Z on qubit 0 is " + str(expectation_value(state_superposition, IZ)))
print("Operator Z on qubit 1 is " + str(expectation_value(state_superposition, ZI)))
print("Operator X on qubit 0 is " + str(expectation_value(state_superposition, IX)))
print("Operator X on qubit 1 is " + str(expectation_value(state_superposition, XI)))
```
#### Mixed states
Consider the expectation value with respect to a statistical mixture of states
$$\langle A \rangle = \sum_k P_k \langle \psi_k |A |\psi_k\rangle = \mathrm{Tr}[A \rho ]. $$
Here,
$$\rho = \sum_k p_k |\psi_k\rangle\langle \psi_k |.$$
captures the most general type of quantum state known as a mixed state. $\rho$ is called the density matrix or state operator. A quantum state operator must obey the following three constraints:
1. Normalization, $\mathrm{Tr}(\rho) = 1$.
2. Hermiticity, $\rho^\dagger = \rho$.
3. Positive semi-definiteness, i.e. eigenvalues of $\rho$ must be non-negative.
The real and imaginary matrix elements $\rho_{i,j}$ represent the standard representation of a quantum state. Qiskit provides the following function to plot them:
```
plot_state(rho, method="city")
```
This draws two 2-dimensional bargraphs (for the real and the imaginary part of $\rho$). Note that the diagonal is necessarily real and sums to 1 (due to normalization), and $\rho_{i,j} =\rho_{j,i}^*$ due to Hermiticity.
It can be useful to interpret operators as vectors in a complex space of dimensions $d^2$ ($\mathbb{C}^{d^2}$). To access the element $a_{ij}$, we use $a_p$ where $p = i + jd$. The indices $i$ and $j$ associated with a given $p$ are given by $i = p \% d$ and $j = \mathrm{floor}(p/d)$. We will use double-ket notation $\mid A \rangle$ to represent the matrix $A$ as a vector in the operator vector space. In this operator space, we also define an inner product in the standard way:
$$
\newcommand{\llangle}{\langle\!\langle}
\newcommand{\rrangle}{\rangle\!\rangle}
\llangle A \mid B\rrangle = \sum_{p=0}^{d^2-1}a^*_pb_p=\sum_{i,j=0}^{d-1} a_{ij}^* b_{ij}=\mathrm{Tr}[A^\dagger B].$$
Employing an orthonormal basis $\{\mid A_j \rrangle\}$ of the operator space, we can decompose $\rho$:
$$\mid \rho \rrangle = \sum_{j=0}^{d^2-1} \mid A_j\rrangle\llangle A_j\mid\rho\rrangle = \sum_{j=0}^{d^2-1} \rho_j \mid A_j\rrangle.$$
(Here, some of the basis states may represent operators that are not measurable.) The Pauli basis, consisting of the $4^n$ operators formed by all tensor products of the Pauli operators ${I,X,Y,Z}$, provides a special basis in which $\rho$ has only real-valued coefficients $\rho_q$,
$$\mid \rho \rrangle = \frac{1}{d}\sum_{q=0}^{d^2-1} \rho_q \mid P_q\rrangle. $$
To display a bar graph of these coefficients, Qiskit provides the method
```
plot_state(rho, method="paulivec"),
```
as well as the method
```
plot_state(rho, method="qsphere")
```
which plots the "qspheres" of the quantum state as follows. Since $\rho$ is a hermitian operator, we can diagonalize it:
$$\rho = \sum_k \lambda_k \left|\lambda_k\right\rangle\!\left\langle \lambda_k \right|.$$
For each eigenvalue $\lambda_k$, the corresponding pure state $\left|\lambda_k\right\rangle$ is plotted on a "qsphere". Each "qsphere" is divided into $n+1$ levels. Each such level is used to represent the weight (total number of 1s) of the binary outcome. The top level corresponds to the $\left|0\ldots0\right\rangle$ state, the next level includes all states with a single 1 ($\left|10\ldots0\right\rangle$, $\left|010\ldots0\right\rangle$, etc.), the level after that comprises all states with two 1s, and so on. Finally, the bottom level represents the state $\left|1\ldots1\right\rangle$. The contrast of each line is set to $\left|\langle i\mid\lambda\rangle\right|^2$, and the color represents the phase via the angle $\angle(\langle i\mid\lambda\rangle)$, with the global phase normalized to the maximum amplitude.
This visualization gives a useful and compact representation for quantum states that are close to pure states.
As an example we consider the same states as above.
```
def state_2_rho(state):
return np.outer(state, state.conj())
rho_superposition=state_2_rho(state_superposition)
print(rho_superposition)
plot_state(rho_superposition,'city')
plot_state(rho_superposition,'paulivec')
plot_state(rho_superposition,'qsphere')
plot_state(rho_superposition,'bloch')
n = 2 # number of qubits
q = QuantumRegister(n)
c = ClassicalRegister(n)
qc2 = QuantumCircuit(q, c)
qc2.h(q[1])
qc2.z(q[1])
# execute the quantum circuit
backend = Aer.get_backend('statevector_simulator')
job = execute(qc2, backend)
state_neg_superposition = job.result().get_statevector(qc2)
rho_neg_superposition=state_2_rho(state_neg_superposition)
plot_state(rho_neg_superposition, 'qsphere')
plot_state(0.5*rho_neg_superposition + 0.5* rho_superposition, 'qsphere')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/BNkosi/Zeus/blob/master/Zeus.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Zeus.py
## Contents
1. First installation
2. Imports
3. Data
4. Data cleaning and Preprocessing
5. Retriever
6. Reader
7. Finder
8. Prediction
```
# First instalation
!pip install git+https://github.com/deepset-ai/haystack.git
!pip install torch==1.5.1+cu101 torchvision==0.6.1+cu101 -f https://download.pytorch.org/whl/torch_stable.html
# Make sure you have a GPU running
!nvidia-smi
```
## Imports
```
# Minimum imports
from haystack import Finder
from haystack.indexing.cleaning import clean_wiki_text
from haystack.indexing.utils import convert_files_to_dicts, fetch_archive_from_http
from haystack.reader.farm import FARMReader
from haystack.reader.transformers import TransformersReader
from haystack.utils import print_answers
from haystack.database.faiss import FAISSDocumentStore
from haystack.retriever.dense import DensePassageRetriever
```
## Load Data
```
def fetch_data_from_repo(doc_dir = "data5/website_data/",
s3_url = "https://github.com/Thabo-5/Chatbot-scraper/raw/master/txt_files.zip",
doc_store=FAISSDocumentStore()):
"""
Function to download data from s3 bucket/ github
Parameters
----------
doc_dir (str): path to destination folder
s3_url (str): path to download zipped data
doc_store (class): Haystack document store
Returns
-------
document_store (object): Haystack document store object
"""
document_store=doc_store
fetch_archive_from_http(url=s3_url, output_dir=doc_dir)
import os
for filename in os.listdir(path=doc_dir):
with open(os.path.join(doc_dir, filename), 'r', encoding='utf-8', errors='replace') as file:
text = file.read()
file.close()
with open(os.path.join(doc_dir, filename), 'w', encoding='utf-8', errors='replace') as file:
file.write(text)
file.close()
# Convert files to dicts
dicts = convert_files_to_dicts(dir_path=doc_dir, clean_func=clean_wiki_text, split_paragraphs=True)
# Now, let's write the dicts containing documents to our DB.
document_store.write_documents(dicts)
return document_store
document_store = fetch_data_from_repo()
```
## Initialize Retriver, Reader and Finder
```
def initFinder():
"""
Function to initiate retriever, reader and finder
Parameters
----------
Returns
-------
finder (object): Haystack finder
"""
retriever = DensePassageRetriever(document_store=document_store,
query_embedding_model="facebook/dpr-question_encoder-single-nq-base",
passage_embedding_model="facebook/dpr-ctx_encoder-single-nq-base",
use_gpu=False,
embed_title=True,
max_seq_len=256,
batch_size=16,
remove_sep_tok_from_untitled_passages=True)
# Important:
# Now that after we have the DPR initialized, we need to call update_embeddings() to iterate over all
# previously indexed documents and update their embedding representation.
# While this can be a time consuming operation (depending on corpus size), it only needs to be done once.
# At query time, we only need to embed the query and compare it the existing doc embeddings which is very fast.
document_store.update_embeddings(retriever)
reader = FARMReader(model_name_or_path="deepset/roberta-base-squad2", use_gpu=False)
return Finder(reader, retriever)
finder = initFinder()
def getAnswers(retrieve=3, read=5, num_answers=1):
while(True):
query = input("You: ")
if query == "bye":
print("Goodbye!")
break
prediction = finder.get_answers(question=query, top_k_retriever=retrieve, top_k_reader=read)
for i in range(0, num_answers):
print(f"\nAnswer\t: {prediction['answers'][i]['answer']}")
print(f"Context\t: {prediction['answers'][i]['context']}")
print(f"Document name\t: {prediction['answers'][i]['meta']['name']}")
print(f"Probability\t: {prediction['answers'][i]['probability']}\n\n")
getAnswers()
getAnswers(5,3,1)
```
| github_jupyter |
# Table widgets in the napari viewer
Before we talk about tables and widgets in napari, let's create a viewer, a simple test image and a labels layer:
```
import numpy as np
import napari
import pandas
from napari_skimage_regionprops import regionprops, add_table, get_table
viewer = napari.Viewer()
viewer.add_image(np.asarray([[1,2],[2,2]]))
viewer.add_labels(np.asarray([[1,2],[3,3]]))
```
Now, let's perform a measurement of `size` and `intensity` of the labeled objects in the given image. A table with results will be automatically added to the viewer
```
regionprops(
viewer.layers[0],
viewer.layers[1],
viewer,
size=True,
intensity=True
)
napari.utils.nbscreenshot(viewer)
```
We can also get the widget representing the table:
```
# The table is associated with a given labels layer:
labels = viewer.layers[1]
table = get_table(labels, viewer)
table
```
You can also read the content from the table as a dictionary. It is recommended to convert it into a pandas `DataFrame`:
```
content = pandas.DataFrame(table.get_content())
content
```
The content of this table can be changed programmatically. This also changes the `properties` of the associated layer.
```
new_values = {'A': [1, 2, 3],
'B': [4, 5, 6]
}
table.set_content(new_values)
napari.utils.nbscreenshot(viewer)
```
You can also append data to an existing table through the `append_content()` function: Suppose you have another measurement for the labels in your image, i.e. the "double area":
```
table.set_content(content.to_dict('list'))
double_area = {'label': content['label'].to_numpy(),
'Double area': content['area'].to_numpy() * 2.0}
```
You can now append this as a new column to the existing table:
```
table.append_content(double_area)
napari.utils.nbscreenshot(viewer)
```
*Note*: If the added data has columns in common withh the exisiting table (for instance, the labels columns), the tables will be merged on the commonly available columns. If no common columns exist, the data will simply be added to the table and the non-intersecting row/columns will be filled with NaN:
```
tripple_area = {'Tripple area': content['area'].to_numpy() * 3.0}
table.append_content(tripple_area)
napari.utils.nbscreenshot(viewer)
```
Note: Changing the label's `properties` does not invoke changes of the table...
```
new_values = {'C': [6, 7, 8],
'D': [9, 10, 11]
}
labels.properties = new_values
napari.utils.nbscreenshot(viewer)
```
But you can refresh the content:
```
table.update_content()
napari.utils.nbscreenshot(viewer)
```
You can remove the table from the viewer like this:
```
viewer.window.remove_dock_widget(table)
napari.utils.nbscreenshot(viewer)
```
Afterwards, the `get_table` method will return None:
```
get_table(labels, viewer)
```
To add the table again, just call `add_table` again. Note that the content of the properties of the labels have not been changed.
```
add_table(labels, viewer)
napari.utils.nbscreenshot(viewer)
```
| github_jupyter |
```
import numpy as np
import time
import sys
#
from matplotlib import pyplot as plt
%matplotlib inline
```
### Real Space LPT ###
In this notebook we give some examples to compute real-space 1-loop halo/matter pothird_orderra in LPT, as well as cross spectra of the "component fields" that comprise the emulator in Modi et al. 2019 (https://arxiv.org/abs/1910.07097).
This is done using the CLEFT class, which is the basic object in the LPT modules.
```
from velocileptors.LPT.cleft_fftw import CLEFT
# To match the plots in Chen, Vlah & White (2020) let's
# work at z=0.8, and scale our initial power spectrum
# to that redshift:
z,D,f = 0.8,0.6819,0.8076
klin,plin = np.loadtxt("pk.dat",unpack=True)
plin *= D**2
# Initialize the class -- with no wisdom file passed it will
# experiment to find the fastest FFT algorithm for the system.
start= time.time()
cleft = CLEFT(klin,plin)
print("Elapsed time: ",time.time()-start," seconds.")
# You could save the wisdom file here if you wanted:
# mome.export_wisdom(wisdom_file_name)
```
### Halo-Halo Autospectrum in Real Space ###
This is the basic application of CLEFT, so comes with its own auxiliary function.
All we need to do is make a power spectrum table and call it.
```
# The parameters we feed it are: b1, b2, bs, b3, alpha, and sn
# The first four are deterministic Lagrangian bias up to third order
# While alpha and sn are the counterterm and stochastic term (shot noise)
pars = [0.70, -1.3, -0.06, 0, 7.4, 1.9e3]
#
start= time.time()
cleft.make_ptable(nk=200)
kv, pk = cleft.combine_bias_terms_pk(*pars)
print("Elapsed time: ",time.time()-start," seconds.")
plt.plot(kv, kv * pk)
plt.xlim(0,0.25)
plt.ylim(850,1120)
plt.ylabel(r'k $P_{hh}(k)$ [h$^{-2}$ Mpc$^2$]')
plt.xlabel('k [h/Mpc]')
plt.show()
```
### Lagrangian Component Spectra ###
All spectra in LPT can be thought of as sums of cross spectra of bias operators $delta_X(q)$ shifted to their observed positions $x = q + \Psi$.
At up to third order these operators are $\{1, b_1, b_2, b_s, b_3\}$, not including derivative bias (which is roughly $b_1 \times k^2$) and stochastic contributions (e.g. shot noise).
```
# Let's explicitly list the components
# Note that the cross spectra are multiplied by a factor of one half.
kv = cleft.pktable[:,0]
spectra = {\
r'$(1,1)$':cleft.pktable[:,1],\
r'$(1,b_1)$':0.5*cleft.pktable[:,2], r'$(b_1,b_1)$': cleft.pktable[:,3],\
r'$(1,b_2)$':0.5*cleft.pktable[:,4], r'$(b_1,b_2)$': 0.5*cleft.pktable[:,5], r'$(b_2,b_2)$': cleft.pktable[:,6],\
r'$(1,b_s)$':0.5*cleft.pktable[:,7], r'$(b_1,b_s)$': 0.5*cleft.pktable[:,8], r'$(b_2,b_s)$':0.5*cleft.pktable[:,9], r'$(b_s,b_s)$':cleft.pktable[:,10],\
r'$(1,b_3)$':0.5*cleft.pktable[:,11],r'$(b_1,b_3)$': 0.5*cleft.pktable[:,12]}
# Plot some of them!
plt.figure(figsize=(15,10))
spec_names = spectra.keys()
for spec_name in spec_names:
plt.loglog(kv, spectra[spec_name],label=spec_name)
plt.ylim(10,3e4)
plt.legend(ncol=4)
plt.xlabel('k [h/Mpc]')
plt.ylabel(r'$P_{ab}$ [(Mpc/h)$^3$]')
plt.show()
```
### Bonus: Cross Spectra with Matter in Real Space ###
In the language of the component spectra the matter field is just "1."
This means we have straightforwardly
$P_{mm} = P_{11}$
and
$P_{hm} = P_{11} + b_1 P_{1b_1} + b_2 P_{1b_2} + b_s P_{1b_s} + b_3 P_{1b_3} + $ eft corrections.
```
# Note that if desired one can also add subleading k^n
# type stochastic corrections to these
def combine_bias_terms_pk_matter(alpha):
kv = cleft.pktable[:,0]
ret = cleft.pktable[:,1] + alpha*kv**2 * cleft.pktable[:,13]
return kv, ret
def combine_bias_terms_pk_crossmatter(b1,b2,bs,b3,alpha):
kv = cleft.pktable[:,0]
ret = cleft.pktable[:,1] + 0.5*b1*cleft.pktable[:,2] \
+ 0.5*b2*cleft.pktable[:,4] + 0.5*bs*cleft.pktable[:,7] + 0.5*b3*cleft.pktable[:,11]\
+ alpha*kv**2 * cleft.pktable[:,13]
return kv, ret
plt.figure(figsize=(10,5))
alpha_mm = 2
b1, b2, bs, b3, alpha_hm = 0.70, -1.3, -0.06, 0, 5
kv, phm = combine_bias_terms_pk_crossmatter(b1,b2,bs,b3,alpha_hm)
kv, pmm = combine_bias_terms_pk_matter(alpha_mm)
plt.plot(kv, kv * pk, label='hh')
plt.plot(kv, kv * phm, label='hm')
plt.plot(kv, kv * pmm, label='mm')
plt.xlim(0,0.25)
plt.ylim(0,1220)
plt.ylabel(r'$k P(k)$ [h$^{-2}$ Mpc$^2$]')
plt.xlabel('k [h/Mpc]')
plt.legend()
plt.show()
```
| github_jupyter |
```
import os
import math
import configparser
import numpy as np
import pandas as pd
from sklearn.neighbors import NearestNeighbors
import tensorflow as tf
from tensorflow import keras
import py
import mylib
import cv2 as cv
import pytesseract
from tqdm import tqdm
from typing import Optional, List, Dict, Set, Tuple
from scml.nlp import strip_punctuation, to_ascii_str
IMAGE = True
TITLE = True
PHASH = True
OCR = False
MODEL = 'efficientnetb3'
pd.set_option("use_inf_as_na", True)
pd.set_option("display.max_columns", 9999)
pd.set_option("display.max_rows", 9999)
pd.set_option('max_colwidth', 9999)
#os.environ["OMP_THREAD_LIMIT"] = "1"
CONF = configparser.ConfigParser()
CONF.read("app.ini")
resolution = int(CONF[MODEL]["resolution"])
print(f"resolution={resolution}")
train = pd.read_csv("input/train.csv", engine="c", low_memory=False)
train["target"] = mylib.target_label(train)
train["image_path"] = "input/train_images/" + train["image"]
posting_ids = train["posting_id"].tolist()
train.info()
%%time
# required for post-processing
train["title_p"] = train.apply(mylib.preprocess("title"), axis=1)
imap = {}
for t in tqdm(train.itertuples()):
pid = getattr(t, "posting_id")
title = getattr(t, "title_p")
imap[pid] = mylib.extract(title)
```
# PHash
th=.25, f1=.586 | th=.30, f1=.586 | th=.35, f1=.587 | th=.40, f1=.583
```
%%time
if PHASH:
train["phash_matches"] = mylib.phash_matches(train, threshold=0.3)
```
# Title
```
%%time
if TITLE:
st_name = "stsb-distilbert-base"
#st_name = "paraphrase-distilroberta-base-v1"
#st_name = "paraphrase-xlm-r-multilingual-v1"
train["title_matches"] = mylib.sbert_matches(
model_path=f"pretrained/sentence-transformers/{st_name}",
sentences=train["title_p"].tolist(),
posting_ids=posting_ids,
threshold=0.5
)
```
# Image
```
if IMAGE:
model_dir = "models/eb3_arc_20210510_1800"
m0 = keras.models.load_model(f"{model_dir}/trial_0/model.h5")
m0 = keras.models.Model(inputs=m0.input[0], outputs=m0.get_layer("embedding_output").output)
m0.summary()
if IMAGE:
idg = keras.preprocessing.image.ImageDataGenerator(
rescale=1./255,
data_format="channels_last",
dtype=np.float32
)
data = idg.flow_from_dataframe(
dataframe=train,
x_col="image",
y_col="label_group",
directory="input/train_images",
target_size=(resolution, resolution),
color_mode="rgb",
batch_size=1024,
shuffle=False,
class_mode="raw",
interpolation="nearest",
)
y0 = m0.predict(data, verbose=1)
#y1 = m1.predict(data, verbose=1)
#y2 = m2.predict(data, verbose=1)
#y3 = m3.predict(data, verbose=1)
#y4 = m4.predict(data, verbose=1)
#assert y0.shape == y1.shape == y2.shape == y3.shape == y4.shape
#print(f"y0.shape={y0.shape}")
em = y0.astype(np.float32)
print(f"em.shape={em.shape}")
#res = []
#for i in range(len(y0)):
#a = np.vstack((y0[i], y1[i], y2[i], y3[i], y4[i]))
#a = np.vstack((y0[i], y1[i]))
#m = np.mean(a, axis=0)
#res.append(m)
#em = np.array(res, dtype=np.float32)
#assert y0.shape == em.shape
#print(f"em.shape={em.shape}")
%%time
if IMAGE:
threshold = 1e-4
nn = NearestNeighbors(
n_neighbors=min(49, len(posting_ids) - 1), metric="euclidean", n_jobs=-1
)
nn.fit(em)
distances, indices = nn.kneighbors()
res: List[List[str]] = [[] for _ in range(len(indices))]
for i in range(len(indices)):
for j in range(len(indices[0])):
if distances[i][j] > threshold:
break
res[i].append(posting_ids[indices[i][j]])
train["image_matches"] = res
```
# OCR
```
def erode_dilate(img):
kernel = np.ones((2, 2), np.uint8)
img = cv.erode(img, kernel, iterations=1)
img = cv.dilate(img, kernel, iterations=1)
return img
def image_to_text(img_path, mode: str, timeout: float, neighbours: int=41, psm: int=3) -> Optional[str]:
config = f"--psm {psm}"
s1, s2 = None, None
img = cv.imread(img_path, cv.IMREAD_GRAYSCALE)
#img = cv.resize(img, None, fx=0.5, fy=0.5, interpolation=cv.INTER_AREA)
img = cv.medianBlur(img, 3)
if mode == "binary_inverted" or mode == "binary":
th = cv.adaptiveThreshold(img, 255, cv.ADAPTIVE_THRESH_GAUSSIAN_C, cv.THRESH_BINARY, neighbours, 2)
th = erode_dilate(th)
try:
s1 = pytesseract.image_to_string(th, timeout=timeout, config=config)
except:
s1 = None
if mode == "binary_inverted" or mode == "inverted":
th = cv.adaptiveThreshold(img, 255, cv.ADAPTIVE_THRESH_GAUSSIAN_C, cv.THRESH_BINARY_INV, neighbours, 2)
th = erode_dilate(th)
try:
s2 = pytesseract.image_to_string(th, timeout=timeout, config=config)
except:
s2 = None
if s1 is None and s2 is None:
return None
tokens = []
if s1 is not None:
s1 = to_ascii_str(s1)
s1 = strip_punctuation(s1)
tokens += s1.split()
if s2 is not None:
s2 = to_ascii_str(s2)
s2 = strip_punctuation(s2)
tokens += s2.split()
return " ".join(tokens)
if OCR:
res = []
n_timeout = 0
for t in tqdm(train.itertuples()):
img_path = getattr(t, "image_path")
s = image_to_text(img_path, mode="inverted", timeout=0.4, neighbours=41, psm=11)
if s is None:
s = ""
n_timeout += 1
res.append(s)
print(f"n_timeout={n_timeout}")
if OCR:
train["itext"] = res
train["text"] = train["title"] + " " + train["itext"]
cols = ["text", "itext", "title"]
train[cols].head()
%%time
if OCR:
train["text_p"] = train.apply(mylib.preprocess("text"), axis=1)
if OCR:
st_name = "stsb-distilbert-base"
#st_name = "paraphrase-distilroberta-base-v1"
#st_name = "paraphrase-xlm-r-multilingual-v1"
train["text_matches"] = mylib.sbert_matches(
model_path=f"pretrained/sentence-transformers/{st_name}",
sentences=train["text_p"].tolist(),
posting_ids=posting_ids,
threshold=0.5
)
```
# Result
```
fs = []
if IMAGE:
fs.append("image_matches")
if TITLE:
fs.append("title_matches")
if PHASH:
fs.append("phash_matches")
if OCR:
fs.append("text_matches")
train["matches"] = train.apply(mylib.combine_as_list(
fs,
imap=imap,
brand_threshold=0.5,
measurement_threshold=0.5,
), axis=1)
train["f1"] = train.apply(mylib.metric_per_row("matches"), axis=1)
print(f"Combined score={train.f1.mean():.3f}")
res = [
{
"score": 0.654,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 1e-6,
"image_pretrained": "enb3",
"brand_theshold": 0.3,
"measurement_threshold": 0.3,
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.654,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 1e-5,
"image_pretrained": "enb3",
"brand_theshold": 0.3,
"measurement_threshold": 0.3,
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.654,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 1e-4,
"image_pretrained": "enb3",
"brand_theshold": 0.3,
"measurement_threshold": 0.3,
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.645,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 1e-3,
"image_pretrained": "enb3",
"brand_theshold": 0.3,
"measurement_threshold": 0.3,
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.656,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 5e-3,
"image_pretrained": "enb3",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.522,
"phash_threshold": None,
"title_threshold": None,
"image_threshold": 5e-3,
"image_pretrained": "enb3",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.473,
"phash_threshold": None,
"title_threshold": None,
"image_threshold": 0.01,
"image_pretrained": "enb3",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.502,
"phash_threshold": None,
"title_threshold": None,
"image_threshold": 1e-3,
"image_pretrained": "enb3",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.651,
"phash_threshold": 0.2,
"title_threshold": 0.5,
"image_threshold": 1e-4,
"image_pretrained": "enb3",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.654,
"phash_threshold": 0.2,
"title_threshold": 0.5,
"image_threshold": 1e-5,
"image_pretrained": "enb3",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.658,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 1e-5,
"image_pretrained": "enb3",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.656,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 1e-4,
"image_pretrained": "enb3",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.562,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 0.001,
"image_pretrained": "enb3",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.514,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 0.001,
"image_pretrained": "enb0",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.498,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 0.01,
"image_pretrained": "enb0",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.136,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": 0.05,
"image_pretrained": "enb0",
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
},
{
"score": 0.674,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"text_threshold": 0.5,
"image_threshold": None,
"image_pretrained": None,
"ocr_threshold": "inverted",
"ocr_timeout": 0.4,
"ocr_neighbours": 41,
"ocr_psm": 11
},
{
"score": 0.674,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"text_threshold": 0.5,
"image_threshold": None,
"image_pretrained": None,
"ocr_threshold": "binary",
"ocr_timeout": 0.4,
"ocr_neighbours": 41,
"ocr_psm": 11
},
{
"score": 0.674,
"phash_threshold": 0.3,
"title_threshold": 0.5,
"image_threshold": None,
"image_pretrained": None,
"text_threshold": None,
"ocr_threshold": None,
"ocr_timeout": None,
"ocr_neighbours": None,
"ocr_psm": None
}
]
df = pd.DataFrame.from_records(res)
df.sort_values("score", ascending=False, inplace=True, ignore_index=True)
df.T.head(30)
cols = ["f1", "target", "matches"] + fs
train[cols].head(30)
df = train.sort_values("f1", ascending=True, ignore_index=True)
df[cols].head()
```
| github_jupyter |
# Markdown Cells
Text can be added to IPython Notebooks using Markdown cells. Markdown is a popular markup language that is a superset of HTML. Its specification can be found here:
<http://daringfireball.net/projects/markdown/>
## Markdown basics
You can make text *italic* or **bold**.
You can build nested itemized or enumerated lists:
* One
- Sublist
- This
- Sublist
- That
- The other thing
* Two
- Sublist
* Three
- Sublist
Now another list:
1. Here we go
1. Sublist
2. Sublist
2. There we go
3. Now this
You can add horizontal rules:
---
Here is a blockquote:
> Beautiful is better than ugly.
> Explicit is better than implicit.
> Simple is better than complex.
> Complex is better than complicated.
> Flat is better than nested.
> Sparse is better than dense.
> Readability counts.
> Special cases aren't special enough to break the rules.
> Although practicality beats purity.
> Errors should never pass silently.
> Unless explicitly silenced.
> In the face of ambiguity, refuse the temptation to guess.
> There should be one-- and preferably only one --obvious way to do it.
> Although that way may not be obvious at first unless you're Dutch.
> Now is better than never.
> Although never is often better than *right* now.
> If the implementation is hard to explain, it's a bad idea.
> If the implementation is easy to explain, it may be a good idea.
> Namespaces are one honking great idea -- let's do more of those!
And shorthand for links:
[IPython's website](http://ipython.org)
## Headings
If you want, you can add headings using Markdown's syntax:
# Heading 1
# Heading 2
## Heading 2.1
## Heading 2.2
**BUT most of the time you should use the Notebook's Heading Cells to organize your Notebook content**, as they provide meaningful structure that can be interpreted by other tools, not just large bold fonts.
## Embedded code
You can embed code meant for illustration instead of execution in Python:
def f(x):
"""a docstring"""
return x**2
or other languages:
if (i=0; i<n; i++) {
printf("hello %d\n", i);
x += 4;
}
## LaTeX equations
Courtesy of MathJax, you can include mathematical expressions both inline:
$e^{i\pi} + 1 = 0$ and displayed:
$$e^x=\sum_{i=0}^\infty \frac{1}{i!}x^i$$
## Github flavored markdown (GFM)
The Notebook webapp support Github flavored markdown meaning that you can use triple backticks for code blocks
<pre>
```python
print "Hello World"
```
```javascript
console.log("Hello World")
```
</pre>
Gives
```python
print "Hello World"
```
```javascript
console.log("Hello World")
```
And a table like this :
<pre>
| This | is |
|------|------|
| a | table|
</pre>
A nice Html Table
| This | is |
|------|------|
| a | table|
## General HTML
Because Markdown is a superset of HTML you can even add things like HTML tables:
<table>
<tr>
<th>Header 1</th>
<th>Header 2</th>
</tr>
<tr>
<td>row 1, cell 1</td>
<td>row 1, cell 2</td>
</tr>
<tr>
<td>row 2, cell 1</td>
<td>row 2, cell 2</td>
</tr>
</table>
## Local files
If you have local files in your Notebook directory, you can refer to these files in Markdown cells directly:
[subdirectory/]<filename>
For example, in the images folder, we have the Python logo:
<img src="../images/python_logo.svg" />
<img src="../images/python_logo.svg" />
and a video with the HTML5 video tag:
<video controls src="images/animation.m4v" />
<video controls src="images/animation.m4v" />
These do not embed the data into the notebook file, and require that the files exist when you are viewing the notebook.
### Security of local files
Note that this means that the IPython notebook server also acts as a generic file server
for files inside the same tree as your notebooks. Access is not granted outside the
notebook folder so you have strict control over what files are visible, but for this
reason it is highly recommended that you do not run the notebook server with a notebook
directory at a high level in your filesystem (e.g. your home directory).
When you run the notebook in a password-protected manner, local file access is restricted
to authenticated users unless read-only views are active.
| github_jupyter |
Siu expression autocompletion: _.cyl.\<tab\>
==========================================
Note: this is document is based on [PR 248](https://github.com/machow/siuba/pull/248) by [@tmastny](https://github.com/tmastny), and all the discussion there!
(Drafted on 7 August 2020)
tl;dr. Implementing autocompletion requires 3 components: identifying the DataFrame to complete, understanding IPython autocompletion, and plugging in to it. The approach we took is to use a user's execution history to identify the DataFrame, and to modify `IPCompleter._jedi_matches`. As discussed in this [PR](https://github.com/machow/siuba/pull/258), a useful approach in the future would be to use a simple regex, like RStudio does.
## Problem
The `_` is meant as a lazy way of representing your data.
Currently, a user autocompleting with _.\<tab\> will not receive suggestions for the data they have in mind.
After importing siuba's mtcars data, a user might want to filter by cylinder, but forget its exact name.
Autocomplete to the rescue! They'd be able to press tab to receive handy suggestions, including column names.

**While an exciting feature, this requires solving hard problems**. There are significant technical challenges related to (1) getting the DataFrame to complete, and (2) plugging into the autocomplete architecture.
For example, the most common approach used for autocompletion--and one used by pandas--is to define a `__dir__` method.
This method then lists out everything you want the user to see when they autocomplete.
However, because the `_` object doesn't know anything about DataFrames, it doesn't return anything useful.
```
from siuba.siu import _
dir(_)[:6]
```
In this ADR, I will review how we can find the right DataFrame to autocomplete, the state of autocompletion in IPython, and three potential solutions.
## Key questions
I'll review each of these questions below.
* **framing**: How do we know what DataFrame (e.g. mtcars) the user wants completions for?
* **IPython autocompletion**: What are the key technical hurdles in the existing autocomplete API?
* **glory road**: What are three ways to get completions like in the gif?
## Framing: what DataFrame are users looking for?
Two possibilities come to mind:
1. The DataFrame being used at the start of a pipe.
2. The last DataFrame they defined or used.
### Start of a pipe
```python
(mtcars
>> filter(_.<tab> == 6) # note the tab!
>> mutate(hp2 = _.hp*2)
)
```
A big challenge here is that this code is not valid python (since it has `_. == 6`). We would likely need to use regex to analyze it. Alternatively, looking at the code they've already run, rather then the code they're on, might be a better place to start.
### Last defined or used
The last defined or used DataFrame is likely impossible to identify, since it'd require knowing the order variables get defined and accessed. However, **static analysis of code history** would let us take a guess. For example, the code below shows some different cases. In each case, we could pick the mtcars or cars2 is being used.
```python
# import mtcars
from siuba.data import mtcars
# assign cars2
cars2 = mtcars
# attribute access cars2
cars2.cyl + 1
```
```
import ast
class FrameDetector(ast.NodeVisitor):
def __init__(self):
self.results = []
super().__init__()
def visit_Name(self, node):
# visit any children
self.generic_visit(node)
# store name as a result
self.results.append(node.id)
visitor = FrameDetector()
visitor.visit(ast.parse("""
from siuba.data import mtcars
cars2 = mtcars + 1
"""))
visitor.results
```
The tree is traversed depth first, and can be dumped out for inspection. See [greentreesnakes](https://greentreesnakes.readthedocs.io/en/latest/) for a nice python AST primer.
```
ast.dump(ast.parse("cars2 = mtcars"))
```
Just knowing the variable names is not enough. We also need to know **which ones are DataFrames**. For our guess, we can use what type of object a variable is at the time the user pressed tab (may differ from when the code is run!).
Here is an example of one way that can be done in IPython.
```
import pandas as pd
shell = get_ipython()
[k for k,v in shell.user_ns.items() if isinstance(v, pd.DataFrame)]
```
Last DataFrame defined or used seems ideal!
Once we know the DataFrame the user has in mind, we need to work it into the autocompletion machinery somehow, so that `_.<tab>` returns the same results as if that DataFrame were being autocompleted.
## IPython Autocompletion
This section will go into great detail about how IPython's autocomplete works, to set the stage for technical solutions. Essentially, when a user interacts with autocompletion, there are 3 main libraries involved: ipykernel, IPython, and jedi. This is shown in the dependency graph below.

Essentially, our challenge is figuring how where autocomplete could fit in. Just to set the stage, the IPython IPCompleter uses some of its own useful completion strategies, but the bulk of where we benefit comes from its use of the library jedi.
In the sections below, I'll first give a quick preview of how jedi works, followed by two sequence diagrams of how it's intergrated into the ipykernel.
### Jedi completion
At it's core, jedi is easy to use, and does a mix of static analysis and object evaluation. It's super handy!
The code below shows how it might autocomplete a DataFrame called `zzz`, where we define `zzz` to really be the `mtcars` data.
```
import jedi
from siuba.data import mtcars
interpreter = jedi.Interpreter('zzz.m', [{'zzz': mtcars}])
completions = list(interpreter.complete())
entry = completions[0]
entry, entry.type
```
Notice that it knows the suggestion `mad` is a function! For a column of the data, it knows that it's not a function, but an instance.
The IPython shell has an instance of it's IPCompleter class, and it's `_jedi_matches` method is responsible for doing the jedi stuff.
```
from siuba.data import mtcars
shell = get_ipython()
df_auto = list(shell.Completer._jedi_matches(7, 0, "mtcars."))
df_auto[:5]
```
While this simple description captures the main thrust of how autocomplete works, the full dynamics include some more features such as entry hooks, and some shuffling things around (since the IPCompleter is deprecating its old methods for completing).
The sequence diagrams below show how the kernel sets up autocomplete, and how a specific autocomplete event is run.
Links to code used for diagrams:
* [ipykernel 5.3.4](https://github.com/ipython/ipykernel/blob/5.3.4/ipykernel/ipkernel.py#L64)
* [IPython 7.17.0 - completer.py](https://github.com/ipython/ipython/blob/7.17.0/IPython/core/completer.py#L1861)
* [IPython interactive shell](https://github.com/ipython/ipython/blob/7.17.0/IPython/core/interactiveshell.py#L676)
#### IPython hooks
ipykernel sets everything up, and also exposes methods for using IPCompleter hooks.
[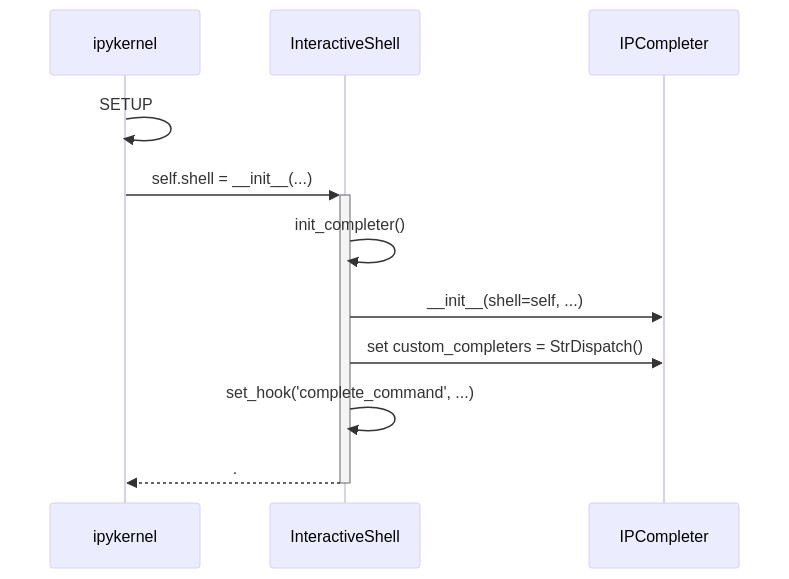](https://mermaid-js.github.io/mermaid-live-editor/#/edit/eyJjb2RlIjoic2VxdWVuY2VEaWFncmFtXG4gIGlweWtlcm5lbC0-PmlweWtlcm5lbDogU0VUVVBcbiAgaXB5a2VybmVsLT4-K0ludGVyYWN0aXZlU2hlbGw6IHNlbGYuc2hlbGwgPSBfX2luaXRfXyguLi4pXG4gIEludGVyYWN0aXZlU2hlbGwtPj5JbnRlcmFjdGl2ZVNoZWxsOiBpbml0X2NvbXBsZXRlcigpXG4gIEludGVyYWN0aXZlU2hlbGwtPj5JUENvbXBsZXRlcjogX19pbml0X18oc2hlbGw9c2VsZiwgLi4uKVxuICBJbnRlcmFjdGl2ZVNoZWxsLT4-SVBDb21wbGV0ZXI6IHNldCBjdXN0b21fY29tcGxldGVycyA9IFN0ckRpc3BhdGNoKClcbiAgSW50ZXJhY3RpdmVTaGVsbC0-PkludGVyYWN0aXZlU2hlbGw6IHNldF9ob29rKCdjb21wbGV0ZV9jb21tYW5kJywgLi4uKVxuICBJbnRlcmFjdGl2ZVNoZWxsLS0-Pi1pcHlrZXJuZWw6IC5cbiAgIiwibWVybWFpZCI6eyJ0aGVtZSI6ImRlZmF1bHQifSwidXBkYXRlRWRpdG9yIjpmYWxzZX0)
(Note that InteractiveShell and IPCompleter come from IPython)
A key here is that one hook, set by the `set_hooks` method is configured using something called `StrDispatch`
```
from IPython.utils.strdispatch import StrDispatch
dis = StrDispatch()
dis.add_s('hei', lambda: 1)
dis.add_re('_\\..*', lambda: 2)
# must be exactly hei
list(dis.flat_matches('hei'))
```
This let's you set hooks that only fire when a specific match is in the code being completed. For example...
```
# needs to match regex abc.*
list(dis.flat_matches('_.abc'))
```
For example, the code below should make `_.a<tab>` complete to `_.ab`.
```
shell = get_ipython()
shell.set_hook('complete_command', lambda shell, event: ['_.ab'], re_key = '_\\.a.*')
```
This would be a really promising avenue. However, as I'll show in the next section, hooks must return a list of strings, so cannot give the nice color info with completions, even if they use jedi under the hood.
#### IPython _jedi_matches
The following diagrams illustrates what the path through a single autocompletion event (e.g. pressing tab) looks like. Note that because IPCompleter is transitioning to a new setup, there is some shuffling around that goes on (e.g. do_complete calls _experimental_do_complete, etc..).
[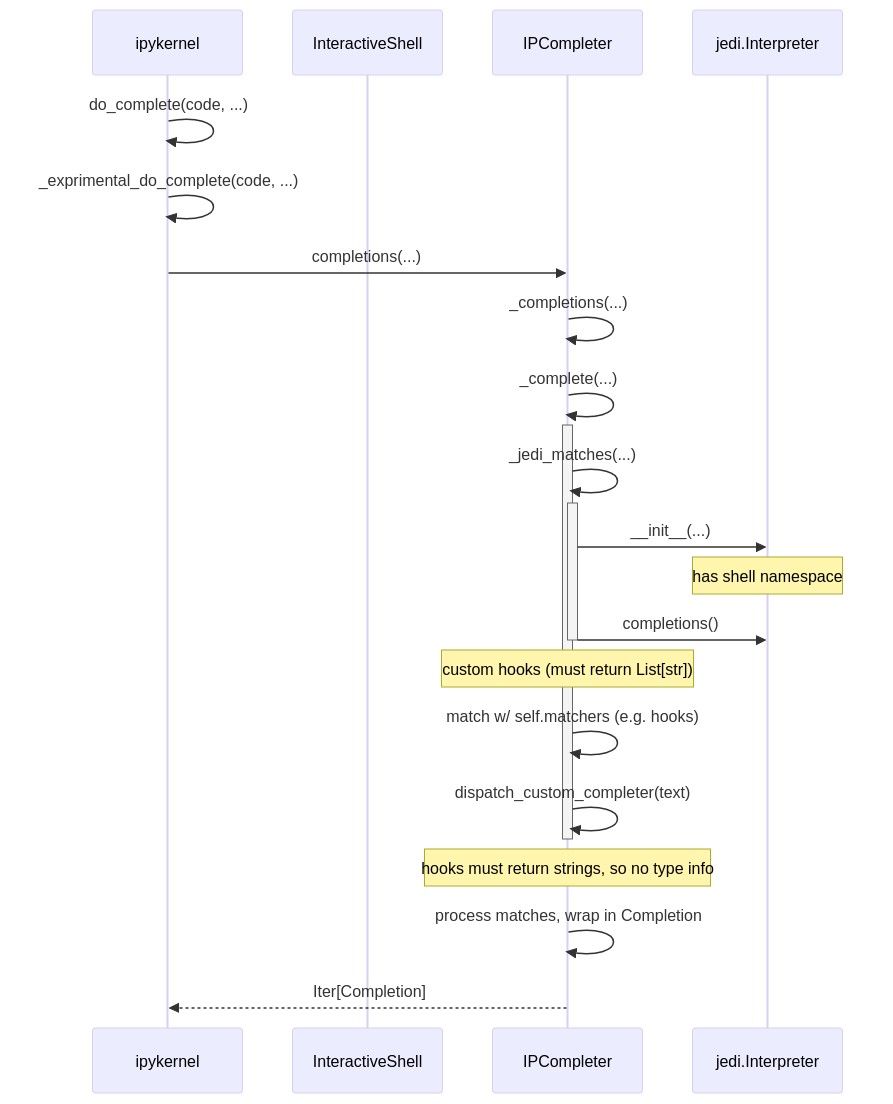](https://mermaid-js.github.io/mermaid-live-editor/#/edit/eyJjb2RlIjoic2VxdWVuY2VEaWFncmFtXG4gIFxuICBcbiAgaXB5a2VybmVsLT4-aXB5a2VybmVsOiBkb19jb21wbGV0ZShjb2RlLCAuLi4pXG4gIGlweWtlcm5lbC0-PmlweWtlcm5lbDogX2V4cHJpbWVudGFsX2RvX2NvbXBsZXRlKGNvZGUsIC4uLilcblxuICBwYXJ0aWNpcGFudCBJbnRlcmFjdGl2ZVNoZWxsXG5cbiAgaXB5a2VybmVsLT4-SVBDb21wbGV0ZXI6IGNvbXBsZXRpb25zKC4uLilcbiAgSVBDb21wbGV0ZXItPj5JUENvbXBsZXRlcjogX2NvbXBsZXRpb25zKC4uLilcbiAgSVBDb21wbGV0ZXItPj4rSVBDb21wbGV0ZXI6IF9jb21wbGV0ZSguLi4pXG4gIElQQ29tcGxldGVyLT4-K0lQQ29tcGxldGVyOiBfamVkaV9tYXRjaGVzKC4uLilcbiAgSVBDb21wbGV0ZXItPj5qZWRpLkludGVycHJldGVyOiBfX2luaXRfXyguLi4pXG4gIG5vdGUgb3ZlciBqZWRpLkludGVycHJldGVyOiBoYXMgc2hlbGwgbmFtZXNwYWNlXG4gIElQQ29tcGxldGVyLT4-LWplZGkuSW50ZXJwcmV0ZXI6IGNvbXBsZXRpb25zKClcbiAgbm90ZSBvdmVyIElQQ29tcGxldGVyOiBjdXN0b20gaG9va3MgKG11c3QgcmV0dXJuIExpc3Rbc3RyXSlcbiAgSVBDb21wbGV0ZXItPj5JUENvbXBsZXRlcjogbWF0Y2ggdy8gc2VsZi5tYXRjaGVycyAoZS5nLiBob29rcylcbiAgSVBDb21wbGV0ZXItPj4tSVBDb21wbGV0ZXI6IGRpc3BhdGNoX2N1c3RvbV9jb21wbGV0ZXIodGV4dClcbiAgbm90ZSBvdmVyIElQQ29tcGxldGVyOiBob29rcyBtdXN0IHJldHVybiBzdHJpbmdzLCBzbyBubyB0eXBlIGluZm9cbiAgSVBDb21wbGV0ZXItPj5JUENvbXBsZXRlcjogcHJvY2VzcyBtYXRjaGVzLCB3cmFwIGluIENvbXBsZXRpb25cblxuICBcbiAgSVBDb21wbGV0ZXItLT4-aXB5a2VybmVsOiBJdGVyW0NvbXBsZXRpb25dXG5cbiIsIm1lcm1haWQiOnsidGhlbWUiOiJkZWZhdWx0In0sInVwZGF0ZUVkaXRvciI6ZmFsc2V9)
Intriguingly, ipykernel also jumps over InteractiveShell, accessing the shell's Completer instance directly. Then, essentially 3 critical steps are run: jedi completions, two kinds of hooks, and wrapping each result in a simple Completion class.
## Glory road: three technical solutions
Essentially, the dynamics described above leave us with three potential solutions for autocomplete:
* hooks (without type info)
* modify siu's `Symbolic.__dir__` method
* monkey patch Completer's _jedi_matches method
To foreshadow, the last is the only one that will give us those sweet colored type annotations, so is preferred!
### Option 1: IPython.Completer hooks
While hooks are an interesting approach, they currently require you to return a list of strings. Only `Completer._jedi_matches` can return the enriched suggestions, and it requires strings from hooks.
(**NOTE:** if you make changes to the code below, you may need to restart your kernel and re-run the cell's code.)
```
#TODO: make workable
from siuba.data import mtcars
from siuba import _
import sys
# will use zzz.<tab> for this example
zzz = _
def hook(shell, event):
# change the completers namespace, then change it back at the end
# would likely need to be done in a context manager, down the road!
old_ns = shell.Completer.namespace
target_df = shell.user_ns["mtcars"]
shell.Completer.namespace = {**old_ns, "zzz": target_df}
# then, run completion method
col_num, line_num = len(event.symbol), 0
completions = shell.Completer._jedi_matches(col_num, line_num, event.symbol)
# change namespace back
shell.Completer.namespace = old_ns
# get suggestions
suggestions = [event.command + x.name for x in completions]
# should be able to see these in the terminal for debugging
with open('/dev/stdout', 'w') as f:
print(suggestions, file = f)
return suggestions
shell = get_ipython()
shell.set_hook('complete_command', hook, re_key = '.*zzz.*')
# uncomment and press tab
#zzz.
```
<div style="width: 200px;">

</div>
### Option 2: monkey patching siuba.siu.Symbolic
Finally, you could imagine that we replace some part of the Symbolic class, so that it does the autocomplete. This is shown below (using a new class rather than monkey patching).
```
from siuba.siu import Symbolic
from siuba.data import mtcars
class Symbolic2(Symbolic):
def __dir__(self):
return dir(mtcars)
```
However, a problem here is that when Jedi completes on a DataFrame (vs something with a `__dir__` method that spits out DataFrame info), it can add type information. With the `__dir__` method, Jedi does not know we want it to think of Symbolic2 as a DataFrame.
```
bbb = Symbolic2()
from siuba import _
import jedi
interpreter = jedi.Interpreter('bbb.', [{'bbb': bbb, 'mtcars': mtcars}])
completions = list(interpreter.complete())
entry = completions[0]
entry.name, entry.type
```
This is why in the output above it doesn't know that `abs` is a function, so reports it as an instance.
### Option 3: monkey patching `IPython.Completer._jedi_matches`
This approach is similar to the above, where we replace `_` in the Completer's namespace with the target DataFrame. However, we do the replacement by manually copying the code of the `_jedi_matches` method, and making the replacement at the very beginning.
Alternatively, you could just wrap _jedi_matches to change `shell.Completer.namespace` as in the hook example.
```
import types
from functools import wraps
from siuba.data import mtcars
from siuba import _
# using aaa for this example
aaa = _
def _jedi_matches_wrapper(obj):
f = obj._jedi_matches
@wraps(f)
def wrapper(self, *args, **kwargs):
# store old namespace (should be context manager)
old_ns = self.namespace
target_df = self.namespace["mtcars"]
self.namespace = {**old_ns, "aaa": target_df}
res = f(*args, **kwargs)
# set namespace back
self.namespace = old_ns
# return results
return res
return types.MethodType(wrapper, obj)
#shell = get_ipython()
#shell.Completer._jedi_matches = _jedi_matches_wrapper(shell.Completer)
from IPython.core.completer import IPCompleter, provisionalcompleter
shell = get_ipython()
completer = IPCompleter(shell, shell.user_ns)
completer._jedi_matches = _jedi_matches_wrapper(shell.Completer)
with provisionalcompleter():
completions = list(completer.completions('aaa.', 4))
completions[:3]
```
Alternatively, you could manually copy the `_jedi_matches` function, and modify it to pass the edited namespace instead.
```python
def _jedi_matches(self, cursor_column:int, cursor_line:int, text:str) -> Iterable[Any]:
# THIS CONTENT IS INSERTED ----
# do namespace stuff...
# THIS CONTENT ORIGINAL ----
# do original stuff in function
shell = get_ipython()
if shell is not None:
shell.Completer._jedi_matches = functools.partial(_jedi_matches, shell.Completer)
```
This has the advantage of not changing state on the Completer, but essentially locks us into using whatever `_jedi_matches` was when we copied it.
## Summary
Autocomplete for `_` that returns info about a DataFrame, requires two things.
1. Identifying what DataFrame the user has in mind
2. Implementing code around IPython's IPCompleter class.
For identifying the right DataFrame, we can use static analysis over a user's code history, along with python's built-in `ast` package.
For implementing the autocomplete itself, our best bet for now is to wrap `IPCompleter._jedi_matches`. In the long run it's worth opening an issue on IPython to discuss how we could get the colored type annotations without this kind of patch, or whether they know of better options!
| github_jupyter |
# **pix2pix**
---
<font size = 4>pix2pix is a deep-learning method allowing image-to-image translation from one image domain type to another image domain type. It was first published by [Isola *et al.* in 2016](https://arxiv.org/abs/1611.07004). The image transformation requires paired images for training (supervised learning) and is made possible here by using a conditional Generative Adversarial Network (GAN) architecture to use information from the input image and obtain the equivalent translated image.
<font size = 4> **This particular notebook enables image-to-image translation learned from paired dataset. If you are interested in performing unpaired image-to-image translation, you should consider using the CycleGAN notebook instead.**
---
<font size = 4>*Disclaimer*:
<font size = 4>This notebook is part of the *Zero-Cost Deep-Learning to Enhance Microscopy* project (https://github.com/HenriquesLab/DeepLearning_Collab/wiki). Jointly developed by the Jacquemet (link to https://cellmig.org/) and Henriques (https://henriqueslab.github.io/) laboratories.
<font size = 4>This notebook is based on the following paper:
<font size = 4> **Image-to-Image Translation with Conditional Adversarial Networks** by Isola *et al.* on arXiv in 2016 (https://arxiv.org/abs/1611.07004)
<font size = 4>The source code of the PyTorch implementation of pix2pix can be found here: https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix
<font size = 4>**Please also cite this original paper when using or developing this notebook.**
# **License**
---
```
#@markdown ##Double click to see the license information
#------------------------- LICENSE FOR ZeroCostDL4Mic------------------------------------
#This ZeroCostDL4Mic notebook is distributed under the MIT licence
#------------------------- LICENSE FOR CycleGAN ------------------------------------
#Copyright (c) 2017, Jun-Yan Zhu and Taesung Park
#All rights reserved.
#Redistribution and use in source and binary forms, with or without
#modification, are permitted provided that the following conditions are met:
#* Redistributions of source code must retain the above copyright notice, this
# list of conditions and the following disclaimer.
#* Redistributions in binary form must reproduce the above copyright notice,
# this list of conditions and the following disclaimer in the documentation
# and/or other materials provided with the distribution.
#THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS"
#AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE
#IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE
#DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT HOLDER OR CONTRIBUTORS BE LIABLE
#FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL
#DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR
#SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER
#CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY,
#OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE
#OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
#--------------------------- LICENSE FOR pix2pix --------------------------------
#BSD License
#For pix2pix software
#Copyright (c) 2016, Phillip Isola and Jun-Yan Zhu
#All rights reserved.
#Redistribution and use in source and binary forms, with or without
#modification, are permitted provided that the following conditions are met:
#* Redistributions of source code must retain the above copyright notice, this
# list of conditions and the following disclaimer.
#* Redistributions in binary form must reproduce the above copyright notice,
# this list of conditions and the following disclaimer in the documentation
# and/or other materials provided with the distribution.
#----------------------------- LICENSE FOR DCGAN --------------------------------
#BSD License
#For dcgan.torch software
#Copyright (c) 2015, Facebook, Inc. All rights reserved.
#Redistribution and use in source and binary forms, with or without modification, are permitted provided that the following conditions are met:
#Redistributions of source code must retain the above copyright notice, this list of conditions and the following disclaimer.
#Redistributions in binary form must reproduce the above copyright notice, this list of conditions and the following disclaimer in the documentation and/or other materials provided with the distribution.
#Neither the name Facebook nor the names of its contributors may be used to endorse or promote products derived from this software without specific prior written permission.
#THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS" AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT HOLDER OR CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
```
# **How to use this notebook?**
---
<font size = 4>Video describing how to use our notebooks are available on youtube:
- [**Video 1**](https://www.youtube.com/watch?v=GzD2gamVNHI&feature=youtu.be): Full run through of the workflow to obtain the notebooks and the provided test datasets as well as a common use of the notebook
- [**Video 2**](https://www.youtube.com/watch?v=PUuQfP5SsqM&feature=youtu.be): Detailed description of the different sections of the notebook
---
###**Structure of a notebook**
<font size = 4>The notebook contains two types of cell:
<font size = 4>**Text cells** provide information and can be modified by douple-clicking the cell. You are currently reading the text cell. You can create a new text by clicking `+ Text`.
<font size = 4>**Code cells** contain code and the code can be modfied by selecting the cell. To execute the cell, move your cursor on the `[ ]`-mark on the left side of the cell (play button appears). Click to execute the cell. After execution is done the animation of play button stops. You can create a new coding cell by clicking `+ Code`.
---
###**Table of contents, Code snippets** and **Files**
<font size = 4>On the top left side of the notebook you find three tabs which contain from top to bottom:
<font size = 4>*Table of contents* = contains structure of the notebook. Click the content to move quickly between sections.
<font size = 4>*Code snippets* = contain examples how to code certain tasks. You can ignore this when using this notebook.
<font size = 4>*Files* = contain all available files. After mounting your google drive (see section 1.) you will find your files and folders here.
<font size = 4>**Remember that all uploaded files are purged after changing the runtime.** All files saved in Google Drive will remain. You do not need to use the Mount Drive-button; your Google Drive is connected in section 1.2.
<font size = 4>**Note:** The "sample data" in "Files" contains default files. Do not upload anything in here!
---
###**Making changes to the notebook**
<font size = 4>**You can make a copy** of the notebook and save it to your Google Drive. To do this click file -> save a copy in drive.
<font size = 4>To **edit a cell**, double click on the text. This will show you either the source code (in code cells) or the source text (in text cells).
You can use the `#`-mark in code cells to comment out parts of the code. This allows you to keep the original code piece in the cell as a comment.
#**0. Before getting started**
---
<font size = 4> For pix2pix to train, **it needs to have access to a paired training dataset**. This means that the same image needs to be acquired in the two conditions and provided with indication of correspondence.
<font size = 4> Therefore, the data structure is important. It is necessary that all the input data are in the same folder and that all the output data is in a separate folder. The provided training dataset is already split in two folders called Training_source and Training_target. Information on how to generate a training dataset is available in our Wiki page: https://github.com/HenriquesLab/ZeroCostDL4Mic/wiki
<font size = 4>**We strongly recommend that you generate extra paired images. These images can be used to assess the quality of your trained model (Quality control dataset)**. The quality control assessment can be done directly in this notebook.
<font size = 4> **Additionally, the corresponding input and output files need to have the same name**.
<font size = 4> Please note that you currently can **only use .PNG files!**
<font size = 4>Here's a common data structure that can work:
* Experiment A
- **Training dataset**
- Training_source
- img_1.png, img_2.png, ...
- Training_target
- img_1.png, img_2.png, ...
- **Quality control dataset**
- Training_source
- img_1.png, img_2.png
- Training_target
- img_1.png, img_2.png
- **Data to be predicted**
- **Results**
---
<font size = 4>**Important note**
<font size = 4>- If you wish to **Train a network from scratch** using your own dataset (and we encourage everyone to do that), you will need to run **sections 1 - 4**, then use **section 5** to assess the quality of your model and **section 6** to run predictions using the model that you trained.
<font size = 4>- If you wish to **Evaluate your model** using a model previously generated and saved on your Google Drive, you will only need to run **sections 1 and 2** to set up the notebook, then use **section 5** to assess the quality of your model.
<font size = 4>- If you only wish to **run predictions** using a model previously generated and saved on your Google Drive, you will only need to run **sections 1 and 2** to set up the notebook, then use **section 6** to run the predictions on the desired model.
---
# **1. Initialise the Colab session**
---
## **1.1. Check for GPU access**
---
By default, the session should be using Python 3 and GPU acceleration, but it is possible to ensure that these are set properly by doing the following:
<font size = 4>Go to **Runtime -> Change the Runtime type**
<font size = 4>**Runtime type: Python 3** *(Python 3 is programming language in which this program is written)*
<font size = 4>**Accelerator: GPU** *(Graphics processing unit)*
```
#@markdown ##Run this cell to check if you have GPU access
import tensorflow as tf
if tf.test.gpu_device_name()=='':
print('You do not have GPU access.')
print('Did you change your runtime ?')
print('If the runtime setting is correct then Google did not allocate a GPU for your session')
print('Expect slow performance. To access GPU try reconnecting later')
else:
print('You have GPU access')
!nvidia-smi
```
## **1.2. Mount your Google Drive**
---
<font size = 4> To use this notebook on the data present in your Google Drive, you need to mount your Google Drive to this notebook.
<font size = 4> Play the cell below to mount your Google Drive and follow the link. In the new browser window, select your drive and select 'Allow', copy the code, paste into the cell and press enter. This will give Colab access to the data on the drive.
<font size = 4> Once this is done, your data are available in the **Files** tab on the top left of notebook.
```
#@markdown ##Play the cell to connect your Google Drive to Colab
#@markdown * Click on the URL.
#@markdown * Sign in your Google Account.
#@markdown * Copy the authorization code.
#@markdown * Enter the authorization code.
#@markdown * Click on "Files" site on the right. Refresh the site. Your Google Drive folder should now be available here as "drive".
# mount user's Google Drive to Google Colab.
from google.colab import drive
drive.mount('/content/gdrive')
```
# **2. Install pix2pix and dependencies**
---
```
Notebook_version = ['1.12']
#@markdown ##Install pix2pix and dependencies
#Here, we install libraries which are not already included in Colab.
import sys
before = [str(m) for m in sys.modules]
!git clone https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix
import os
os.chdir('pytorch-CycleGAN-and-pix2pix/')
!pip install -r requirements.txt
!pip install fpdf
import imageio
from skimage import data
from skimage import exposure
from skimage.exposure import match_histograms
import glob
import os.path
# ------- Common variable to all ZeroCostDL4Mic notebooks -------
import numpy as np
from matplotlib import pyplot as plt
import urllib
import os, random
import shutil
import zipfile
from tifffile import imread, imsave
import time
import sys
from pathlib import Path
import pandas as pd
import csv
from glob import glob
from scipy import signal
from scipy import ndimage
from skimage import io
from sklearn.linear_model import LinearRegression
from skimage.util import img_as_uint
import matplotlib as mpl
from skimage.metrics import structural_similarity
from skimage.metrics import peak_signal_noise_ratio as psnr
from astropy.visualization import simple_norm
from skimage import img_as_float32
from skimage.util import img_as_ubyte
from tqdm import tqdm
from fpdf import FPDF, HTMLMixin
from datetime import datetime
import subprocess
from pip._internal.operations.freeze import freeze
# Colors for the warning messages
class bcolors:
WARNING = '\033[31m'
#Disable some of the tensorflow warnings
import warnings
warnings.filterwarnings("ignore")
print('----------------------------')
print("Libraries installed")
# Check if this is the latest version of the notebook
Latest_notebook_version = pd.read_csv("https://raw.githubusercontent.com/HenriquesLab/ZeroCostDL4Mic/master/Colab_notebooks/Latest_ZeroCostDL4Mic_Release.csv")
if Notebook_version == list(Latest_notebook_version.columns):
print("This notebook is up-to-date.")
if not Notebook_version == list(Latest_notebook_version.columns):
print(bcolors.WARNING +"A new version of this notebook has been released. We recommend that you download it at https://github.com/HenriquesLab/ZeroCostDL4Mic/wiki")
def pdf_export(trained = False, augmentation = False, pretrained_model = False):
class MyFPDF(FPDF, HTMLMixin):
pass
pdf = MyFPDF()
pdf.add_page()
pdf.set_right_margin(-1)
pdf.set_font("Arial", size = 11, style='B')
Network = 'pix2pix'
day = datetime.now()
datetime_str = str(day)[0:10]
Header = 'Training report for '+Network+' model ('+model_name+')\nDate: '+datetime_str
pdf.multi_cell(180, 5, txt = Header, align = 'L')
# add another cell
if trained:
training_time = "Training time: "+str(hour)+ "hour(s) "+str(mins)+"min(s) "+str(round(sec))+"sec(s)"
pdf.cell(190, 5, txt = training_time, ln = 1, align='L')
pdf.ln(1)
Header_2 = 'Information for your materials and method:'
pdf.cell(190, 5, txt=Header_2, ln=1, align='L')
all_packages = ''
for requirement in freeze(local_only=True):
all_packages = all_packages+requirement+', '
#print(all_packages)
#Main Packages
main_packages = ''
version_numbers = []
for name in ['tensorflow','numpy','torch']:
find_name=all_packages.find(name)
main_packages = main_packages+all_packages[find_name:all_packages.find(',',find_name)]+', '
#Version numbers only here:
version_numbers.append(all_packages[find_name+len(name)+2:all_packages.find(',',find_name)])
cuda_version = subprocess.run('nvcc --version',stdout=subprocess.PIPE, shell=True)
cuda_version = cuda_version.stdout.decode('utf-8')
cuda_version = cuda_version[cuda_version.find(', V')+3:-1]
gpu_name = subprocess.run('nvidia-smi',stdout=subprocess.PIPE, shell=True)
gpu_name = gpu_name.stdout.decode('utf-8')
gpu_name = gpu_name[gpu_name.find('Tesla'):gpu_name.find('Tesla')+10]
#print(cuda_version[cuda_version.find(', V')+3:-1])
#print(gpu_name)
shape = io.imread(Training_source+'/'+os.listdir(Training_source)[1]).shape
dataset_size = len(os.listdir(Training_source))
text = 'The '+Network+' model was trained from scratch for '+str(number_of_epochs)+' epochs on '+str(dataset_size)+' paired image patches (image dimensions: '+str(shape)+', patch size: ('+str(patch_size)+','+str(patch_size)+')) with a batch size of '+str(batch_size)+' and a vanilla GAN loss function, using the '+Network+' ZeroCostDL4Mic notebook (v '+Notebook_version[0]+') (von Chamier & Laine et al., 2020). Key python packages used include tensorflow (v '+version_numbers[0]+'), numpy (v '+version_numbers[1]+'), torch (v '+version_numbers[2]+'), cuda (v '+cuda_version+'). The training was accelerated using a '+gpu_name+'GPU.'
if Use_pretrained_model:
text = 'The '+Network+' model was trained for '+str(number_of_epochs)+' epochs on '+str(dataset_size)+' paired image patches (image dimensions: '+str(shape)+', patch size: ('+str(patch_size)+','+str(patch_size)+')) with a batch size of '+str(batch_size)+' and a vanilla GAN loss function, using the '+Network+' ZeroCostDL4Mic notebook (v '+Notebook_version[0]+') (von Chamier & Laine et al., 2020). The model was retrained from a pretrained model. Key python packages used include tensorflow (v '+version_numbers[0]+'), numpy (v '+version_numbers[1]+'), torch (v '+version_numbers[2]+'), cuda (v '+cuda_version+'). The training was accelerated using a '+gpu_name+'GPU.'
pdf.set_font('')
pdf.set_font_size(10.)
pdf.multi_cell(190, 5, txt = text, align='L')
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.ln(1)
pdf.cell(28, 5, txt='Augmentation: ', ln=0)
pdf.set_font('')
if augmentation:
aug_text = 'The dataset was augmented by a factor of '+str(Multiply_dataset_by)+' by'
if rotate_270_degrees != 0 or rotate_90_degrees != 0:
aug_text = aug_text+'\n- rotation'
if flip_left_right != 0 or flip_top_bottom != 0:
aug_text = aug_text+'\n- flipping'
if random_zoom_magnification != 0:
aug_text = aug_text+'\n- random zoom magnification'
if random_distortion != 0:
aug_text = aug_text+'\n- random distortion'
if image_shear != 0:
aug_text = aug_text+'\n- image shearing'
if skew_image != 0:
aug_text = aug_text+'\n- image skewing'
else:
aug_text = 'No augmentation was used for training.'
pdf.multi_cell(190, 5, txt=aug_text, align='L')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(1)
pdf.cell(180, 5, txt = 'Parameters', align='L', ln=1)
pdf.set_font('')
pdf.set_font_size(10.)
if Use_Default_Advanced_Parameters:
pdf.cell(200, 5, txt='Default Advanced Parameters were enabled')
pdf.cell(200, 5, txt='The following parameters were used for training:')
pdf.ln(1)
html = """
<table width=40% style="margin-left:0px;">
<tr>
<th width = 50% align="left">Parameter</th>
<th width = 50% align="left">Value</th>
</tr>
<tr>
<td width = 50%>number_of_epochs</td>
<td width = 50%>{0}</td>
</tr>
<tr>
<td width = 50%>patch_size</td>
<td width = 50%>{1}</td>
</tr>
<tr>
<td width = 50%>batch_size</td>
<td width = 50%>{2}</td>
</tr>
<tr>
<td width = 50%>initial_learning_rate</td>
<td width = 50%>{3}</td>
</tr>
</table>
""".format(number_of_epochs,str(patch_size)+'x'+str(patch_size),batch_size,initial_learning_rate)
pdf.write_html(html)
#pdf.multi_cell(190, 5, txt = text_2, align='L')
pdf.set_font("Arial", size = 11, style='B')
pdf.ln(1)
pdf.cell(190, 5, txt = 'Training Dataset', align='L', ln=1)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(30, 5, txt= 'Training_source:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = Training_source, align = 'L')
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(29, 5, txt= 'Training_target:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = Training_target, align = 'L')
#pdf.cell(190, 5, txt=aug_text, align='L', ln=1)
pdf.ln(1)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(22, 5, txt= 'Model Path:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = model_path+'/'+model_name, align = 'L')
pdf.ln(1)
pdf.cell(60, 5, txt = 'Example Training pair', ln=1)
pdf.ln(1)
exp_size = io.imread('/content/TrainingDataExample_pix2pix.png').shape
pdf.image('/content/TrainingDataExample_pix2pix.png', x = 11, y = None, w = round(exp_size[1]/8), h = round(exp_size[0]/8))
pdf.ln(1)
ref_1 = 'References:\n - ZeroCostDL4Mic: von Chamier, Lucas & Laine, Romain, et al. "ZeroCostDL4Mic: an open platform to simplify access and use of Deep-Learning in Microscopy." BioRxiv (2020).'
pdf.multi_cell(190, 5, txt = ref_1, align='L')
ref_2 = '- pix2pix: Isola, Phillip, et al. "Image-to-image translation with conditional adversarial networks." Proceedings of the IEEE conference on computer vision and pattern recognition. 2017.'
pdf.multi_cell(190, 5, txt = ref_2, align='L')
if augmentation:
ref_3 = '- Augmentor: Bloice, Marcus D., Christof Stocker, and Andreas Holzinger. "Augmentor: an image augmentation library for machine learning." arXiv preprint arXiv:1708.04680 (2017).'
pdf.multi_cell(190, 5, txt = ref_3, align='L')
pdf.ln(3)
reminder = 'Important:\nRemember to perform the quality control step on all newly trained models\nPlease consider depositing your training dataset on Zenodo'
pdf.set_font('Arial', size = 11, style='B')
pdf.multi_cell(190, 5, txt=reminder, align='C')
pdf.output(model_path+'/'+model_name+'/'+model_name+"_training_report.pdf")
def qc_pdf_export():
class MyFPDF(FPDF, HTMLMixin):
pass
pdf = MyFPDF()
pdf.add_page()
pdf.set_right_margin(-1)
pdf.set_font("Arial", size = 11, style='B')
Network = 'pix2pix'
day = datetime.now()
datetime_str = str(day)[0:10]
Header = 'Quality Control report for '+Network+' model ('+QC_model_name+')\nDate: '+datetime_str
pdf.multi_cell(180, 5, txt = Header, align = 'L')
all_packages = ''
for requirement in freeze(local_only=True):
all_packages = all_packages+requirement+', '
pdf.set_font('')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(2)
pdf.cell(190, 5, txt = 'Development of Training Losses', ln=1, align='L')
pdf.ln(1)
exp_size = io.imread(full_QC_model_path+'/Quality Control/SSIMvsCheckpoint_data.png').shape
pdf.image(full_QC_model_path+'/Quality Control/SSIMvsCheckpoint_data.png', x = 11, y = None, w = round(exp_size[1]/8), h = round(exp_size[0]/8))
pdf.ln(2)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.ln(3)
pdf.cell(80, 5, txt = 'Example Quality Control Visualisation', ln=1)
pdf.ln(1)
exp_size = io.imread(full_QC_model_path+'/Quality Control/QC_example_data.png').shape
if Image_type == 'RGB':
pdf.image(full_QC_model_path+'/Quality Control/QC_example_data.png', x = 16, y = None, w = round(exp_size[1]/5), h = round(exp_size[0]/5))
if Image_type == 'Grayscale':
pdf.image(full_QC_model_path+'/Quality Control/QC_example_data.png', x = 16, y = None, w = round(exp_size[1]/8), h = round(exp_size[0]/8))
pdf.ln(1)
pdf.set_font('')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(1)
pdf.cell(180, 5, txt = 'Quality Control Metrics', align='L', ln=1)
pdf.set_font('')
pdf.set_font_size(10.)
pdf.ln(1)
for checkpoint in os.listdir(full_QC_model_path+'/Quality Control'):
if os.path.isdir(os.path.join(full_QC_model_path,'Quality Control',checkpoint)) and checkpoint != 'Prediction':
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(70, 5, txt = 'Metrics for checkpoint: '+ str(checkpoint), align='L', ln=1)
html = """
<body>
<font size="8" face="Courier New" >
<table width=95% style="margin-left:0px;">"""
with open(full_QC_model_path+'/Quality Control/'+str(checkpoint)+'/QC_metrics_'+QC_model_name+str(checkpoint)+'.csv', 'r') as csvfile:
metrics = csv.reader(csvfile)
header = next(metrics)
image = header[0]
mSSIM_PvsGT = header[1]
mSSIM_SvsGT = header[2]
header = """
<tr>
<th width = 60% align="left">{0}</th>
<th width = 20% align="center">{1}</th>
<th width = 20% align="center">{2}</th>
</tr>""".format(image,mSSIM_PvsGT,mSSIM_SvsGT)
html = html+header
for row in metrics:
image = row[0]
mSSIM_PvsGT = row[1]
mSSIM_SvsGT = row[2]
cells = """
<tr>
<td width = 60% align="left">{0}</td>
<td width = 20% align="center">{1}</td>
<td width = 20% align="center">{2}</td>
</tr>""".format(image,str(round(float(mSSIM_PvsGT),3)),str(round(float(mSSIM_SvsGT),3)))
html = html+cells
html = html+"""</body></table>"""
pdf.write_html(html)
pdf.ln(2)
else:
continue
pdf.ln(1)
pdf.set_font('')
pdf.set_font_size(10.)
ref_1 = 'References:\n - ZeroCostDL4Mic: von Chamier, Lucas & Laine, Romain, et al. "ZeroCostDL4Mic: an open platform to simplify access and use of Deep-Learning in Microscopy." BioRxiv (2020).'
pdf.multi_cell(190, 5, txt = ref_1, align='L')
ref_2 = '- pix2pix: Isola, Phillip, et al. "Image-to-image translation with conditional adversarial networks." Proceedings of the IEEE conference on computer vision and pattern recognition. 2017.'
pdf.multi_cell(190, 5, txt = ref_2, align='L')
pdf.ln(3)
reminder = 'To find the parameters and other information about how this model was trained, go to the training_report.pdf of this model which should be in the folder of the same name.'
pdf.set_font('Arial', size = 11, style='B')
pdf.multi_cell(190, 5, txt=reminder, align='C')
pdf.output(full_QC_model_path+'/Quality Control/'+QC_model_name+'_QC_report.pdf')
# Exporting requirements.txt for local run
!pip freeze > ../requirements.txt
after = [str(m) for m in sys.modules]
# Get minimum requirements file
#Add the following lines before all imports:
# import sys
# before = [str(m) for m in sys.modules]
#Add the following line after the imports:
# after = [str(m) for m in sys.modules]
from builtins import any as b_any
def filter_files(file_list, filter_list):
filtered_list = []
for fname in file_list:
if b_any(fname.split('==')[0] in s for s in filter_list):
filtered_list.append(fname)
return filtered_list
df = pd.read_csv('../requirements.txt', delimiter = "\n")
mod_list = [m.split('.')[0] for m in after if not m in before]
req_list_temp = df.values.tolist()
req_list = [x[0] for x in req_list_temp]
# Replace with package name
mod_name_list = [['sklearn', 'scikit-learn'], ['skimage', 'scikit-image']]
mod_replace_list = [[x[1] for x in mod_name_list] if s in [x[0] for x in mod_name_list] else s for s in mod_list]
filtered_list = filter_files(req_list, mod_replace_list)
file=open('../pix2pix_requirements_simple.txt','w')
for item in filtered_list:
file.writelines(item + '\n')
file.close()
```
# **3. Select your parameters and paths**
---
## **3.1. Setting main training parameters**
---
<font size = 4>
<font size = 5> **Paths for training, predictions and results**
<font size = 4>**`Training_source:`, `Training_target`:** These are the paths to your folders containing the Training_source and Training_target training data respecively. To find the paths of the folders containing the respective datasets, go to your Files on the left of the notebook, navigate to the folder containing your files and copy the path by right-clicking on the folder, **Copy path** and pasting it into the right box below.
<font size = 4>**`model_name`:** Use only my_model -style, not my-model (Use "_" not "-"). Do not use spaces in the name. Avoid using the name of an existing model (saved in the same folder) as it will be overwritten.
<font size = 4>**`model_path`**: Enter the path where your model will be saved once trained (for instance your result folder).
<font size = 5>**Training parameters**
<font size = 4>**`number_of_epochs`:**Input how many epochs (rounds) the network will be trained. Preliminary results can already be observed after a few (10) epochs, but a full training should run for 200 epochs or more. Evaluate the performance after training (see 5). **Default value: 200**
<font size = 5>**Advanced Parameters - experienced users only**
<font size = 4>**`patch_size`:** pix2pix divides the image into patches for training. Input the size of the patches (length of a side). The value should be smaller than the dimensions of the image and divisible by 8. **Default value: 512**
<font size = 4>**When choosing the patch_size, the value should be i) large enough that it will enclose many instances, ii) small enough that the resulting patches fit into the RAM.**<font size = 4>
<font size =4>**`batch_size:`** This parameter defines the number of patches seen in each training step. Reducing or increasing the **batch size** may slow or speed up your training, respectively, and can influence network performance. **Default value: 1**
<font size = 4>**`initial_learning_rate`:** Input the initial value to be used as learning rate. **Default value: 0.0002**
```
#@markdown ###Path to training images:
Training_source = "" #@param {type:"string"}
InputFile = Training_source+"/*.png"
Training_target = "" #@param {type:"string"}
OutputFile = Training_target+"/*.png"
#Define where the patch file will be saved
base = "/content"
# model name and path
#@markdown ###Name of the model and path to model folder:
model_name = "" #@param {type:"string"}
model_path = "" #@param {type:"string"}
# other parameters for training.
#@markdown ###Training Parameters
#@markdown Number of epochs:
number_of_epochs = 200#@param {type:"number"}
#@markdown ###Advanced Parameters
Use_Default_Advanced_Parameters = True #@param {type:"boolean"}
#@markdown ###If not, please input:
patch_size = 512#@param {type:"number"} # in pixels
batch_size = 1#@param {type:"number"}
initial_learning_rate = 0.0002 #@param {type:"number"}
if (Use_Default_Advanced_Parameters):
print("Default advanced parameters enabled")
batch_size = 1
patch_size = 512
initial_learning_rate = 0.0002
#here we check that no model with the same name already exist, if so delete
if os.path.exists(model_path+'/'+model_name):
print(bcolors.WARNING +"!! WARNING: "+model_name+" already exists and will be deleted in the following cell !!")
print(bcolors.WARNING +"To continue training "+model_name+", choose a new model_name here, and load "+model_name+" in section 3.3")
#To use pix2pix we need to organise the data in a way the network can understand
Saving_path= "/content/"+model_name
#Saving_path= model_path+"/"+model_name
if os.path.exists(Saving_path):
shutil.rmtree(Saving_path)
os.makedirs(Saving_path)
imageA_folder = Saving_path+"/A"
os.makedirs(imageA_folder)
imageB_folder = Saving_path+"/B"
os.makedirs(imageB_folder)
imageAB_folder = Saving_path+"/AB"
os.makedirs(imageAB_folder)
TrainA_Folder = Saving_path+"/A/train"
os.makedirs(TrainA_Folder)
TrainB_Folder = Saving_path+"/B/train"
os.makedirs(TrainB_Folder)
# Here we disable pre-trained model by default (in case the cell is not ran)
Use_pretrained_model = False
# Here we disable data augmentation by default (in case the cell is not ran)
Use_Data_augmentation = False
# This will display a randomly chosen dataset input and output
random_choice = random.choice(os.listdir(Training_source))
x = imageio.imread(Training_source+"/"+random_choice)
#Find image XY dimension
Image_Y = x.shape[0]
Image_X = x.shape[1]
Image_min_dim = min(Image_Y, Image_X)
#Hyperparameters failsafes
if patch_size > min(Image_Y, Image_X):
patch_size = min(Image_Y, Image_X)
print (bcolors.WARNING + " Your chosen patch_size is bigger than the xy dimension of your image; therefore the patch_size chosen is now:",patch_size)
# Here we check that patch_size is divisible by 4
if not patch_size % 4 == 0:
patch_size = ((int(patch_size / 4)-1) * 4)
print (bcolors.WARNING + " Your chosen patch_size is not divisible by 4; therefore the patch_size chosen is now:",patch_size)
# Here we check that patch_size is at least bigger than 256
if patch_size < 256:
patch_size = 256
print (bcolors.WARNING + " Your chosen patch_size is too small; therefore the patch_size chosen is now:",patch_size)
y = imageio.imread(Training_target+"/"+random_choice)
f=plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.imshow(x, interpolation='nearest')
plt.title('Training source')
plt.axis('off');
plt.subplot(1,2,2)
plt.imshow(y, interpolation='nearest')
plt.title('Training target')
plt.axis('off');
plt.savefig('/content/TrainingDataExample_pix2pix.png',bbox_inches='tight',pad_inches=0)
```
## **3.2. Data augmentation**
---
<font size = 4>
<font size = 4>Data augmentation can improve training progress by amplifying differences in the dataset. This can be useful if the available dataset is small since, in this case, it is possible that a network could quickly learn every example in the dataset (overfitting), without augmentation. Augmentation is not necessary for training and if your training dataset is large you should disable it.
<font size = 4>Data augmentation is performed here by [Augmentor.](https://github.com/mdbloice/Augmentor)
<font size = 4>[Augmentor](https://github.com/mdbloice/Augmentor) was described in the following article:
<font size = 4>Marcus D Bloice, Peter M Roth, Andreas Holzinger, Biomedical image augmentation using Augmentor, Bioinformatics, https://doi.org/10.1093/bioinformatics/btz259
<font size = 4>**Please also cite this original paper when publishing results obtained using this notebook with augmentation enabled.**
```
#Data augmentation
Use_Data_augmentation = True #@param {type:"boolean"}
if Use_Data_augmentation:
!pip install Augmentor
import Augmentor
#@markdown ####Choose a factor by which you want to multiply your original dataset
Multiply_dataset_by = 2 #@param {type:"slider", min:1, max:30, step:1}
Save_augmented_images = False #@param {type:"boolean"}
Saving_path = "" #@param {type:"string"}
Use_Default_Augmentation_Parameters = True #@param {type:"boolean"}
#@markdown ###If not, please choose the probability of the following image manipulations to be used to augment your dataset (1 = always used; 0 = disabled ):
#@markdown ####Mirror and rotate images
rotate_90_degrees = 0 #@param {type:"slider", min:0, max:1, step:0.1}
rotate_270_degrees = 0 #@param {type:"slider", min:0, max:1, step:0.1}
flip_left_right = 0 #@param {type:"slider", min:0, max:1, step:0.1}
flip_top_bottom = 0 #@param {type:"slider", min:0, max:1, step:0.1}
#@markdown ####Random image Zoom
random_zoom = 0 #@param {type:"slider", min:0, max:1, step:0.1}
random_zoom_magnification = 0 #@param {type:"slider", min:0, max:1, step:0.1}
#@markdown ####Random image distortion
random_distortion = 0 #@param {type:"slider", min:0, max:1, step:0.1}
#@markdown ####Image shearing and skewing
image_shear = 0 #@param {type:"slider", min:0, max:1, step:0.1}
max_image_shear = 10 #@param {type:"slider", min:1, max:25, step:1}
skew_image = 0 #@param {type:"slider", min:0, max:1, step:0.1}
skew_image_magnitude = 0 #@param {type:"slider", min:0, max:1, step:0.1}
if Use_Default_Augmentation_Parameters:
rotate_90_degrees = 0.5
rotate_270_degrees = 0.5
flip_left_right = 0.5
flip_top_bottom = 0.5
if not Multiply_dataset_by >5:
random_zoom = 0
random_zoom_magnification = 0.9
random_distortion = 0
image_shear = 0
max_image_shear = 10
skew_image = 0
skew_image_magnitude = 0
if Multiply_dataset_by >5:
random_zoom = 0.1
random_zoom_magnification = 0.9
random_distortion = 0.5
image_shear = 0.2
max_image_shear = 5
skew_image = 0.2
skew_image_magnitude = 0.4
if Multiply_dataset_by >25:
random_zoom = 0.5
random_zoom_magnification = 0.8
random_distortion = 0.5
image_shear = 0.5
max_image_shear = 20
skew_image = 0.5
skew_image_magnitude = 0.6
list_files = os.listdir(Training_source)
Nb_files = len(list_files)
Nb_augmented_files = (Nb_files * Multiply_dataset_by)
if Use_Data_augmentation:
print("Data augmentation enabled")
# Here we set the path for the various folder were the augmented images will be loaded
# All images are first saved into the augmented folder
#Augmented_folder = "/content/Augmented_Folder"
if not Save_augmented_images:
Saving_path= "/content"
Augmented_folder = Saving_path+"/Augmented_Folder"
if os.path.exists(Augmented_folder):
shutil.rmtree(Augmented_folder)
os.makedirs(Augmented_folder)
#Training_source_augmented = "/content/Training_source_augmented"
Training_source_augmented = Saving_path+"/Training_source_augmented"
if os.path.exists(Training_source_augmented):
shutil.rmtree(Training_source_augmented)
os.makedirs(Training_source_augmented)
#Training_target_augmented = "/content/Training_target_augmented"
Training_target_augmented = Saving_path+"/Training_target_augmented"
if os.path.exists(Training_target_augmented):
shutil.rmtree(Training_target_augmented)
os.makedirs(Training_target_augmented)
# Here we generate the augmented images
#Load the images
p = Augmentor.Pipeline(Training_source, Augmented_folder)
#Define the matching images
p.ground_truth(Training_target)
#Define the augmentation possibilities
if not rotate_90_degrees == 0:
p.rotate90(probability=rotate_90_degrees)
if not rotate_270_degrees == 0:
p.rotate270(probability=rotate_270_degrees)
if not flip_left_right == 0:
p.flip_left_right(probability=flip_left_right)
if not flip_top_bottom == 0:
p.flip_top_bottom(probability=flip_top_bottom)
if not random_zoom == 0:
p.zoom_random(probability=random_zoom, percentage_area=random_zoom_magnification)
if not random_distortion == 0:
p.random_distortion(probability=random_distortion, grid_width=4, grid_height=4, magnitude=8)
if not image_shear == 0:
p.shear(probability=image_shear,max_shear_left=20,max_shear_right=20)
if not skew_image == 0:
p.skew(probability=skew_image,magnitude=skew_image_magnitude)
p.sample(int(Nb_augmented_files))
print(int(Nb_augmented_files),"matching images generated")
# Here we sort through the images and move them back to augmented trainning source and targets folders
augmented_files = os.listdir(Augmented_folder)
for f in augmented_files:
if (f.startswith("_groundtruth_(1)_")):
shortname_noprefix = f[17:]
shutil.copyfile(Augmented_folder+"/"+f, Training_target_augmented+"/"+shortname_noprefix)
if not (f.startswith("_groundtruth_(1)_")):
shutil.copyfile(Augmented_folder+"/"+f, Training_source_augmented+"/"+f)
for filename in os.listdir(Training_source_augmented):
os.chdir(Training_source_augmented)
os.rename(filename, filename.replace('_original', ''))
#Here we clean up the extra files
shutil.rmtree(Augmented_folder)
if not Use_Data_augmentation:
print(bcolors.WARNING+"Data augmentation disabled")
```
## **3.3. Using weights from a pre-trained model as initial weights**
---
<font size = 4> Here, you can set the the path to a pre-trained model from which the weights can be extracted and used as a starting point for this training session. **This pre-trained model needs to be a pix2pix model**.
<font size = 4> This option allows you to perform training over multiple Colab runtimes or to do transfer learning using models trained outside of ZeroCostDL4Mic. **You do not need to run this section if you want to train a network from scratch**.
```
# @markdown ##Loading weights from a pre-trained network
Use_pretrained_model = False #@param {type:"boolean"}
#@markdown ###If yes, please provide the path to the model folder:
pretrained_model_path = "" #@param {type:"string"}
# --------------------- Check if we load a previously trained model ------------------------
if Use_pretrained_model:
h5_file_path = os.path.join(pretrained_model_path, "latest_net_G.pth")
# --------------------- Check the model exist ------------------------
if not os.path.exists(h5_file_path):
print(bcolors.WARNING+'WARNING: Pretrained model does not exist')
Use_pretrained_model = False
print(bcolors.WARNING+'No pretrained network will be used.')
if os.path.exists(h5_file_path):
print("Pretrained model "+os.path.basename(pretrained_model_path)+" was found and will be loaded prior to training.")
else:
print(bcolors.WARNING+'No pretrained network will be used.')
```
#**4. Train the network**
---
## **4.1. Prepare the training data for training**
---
<font size = 4>Here, we use the information from Section 3 to prepare the training data into a suitable format for training. **Your data will be copied in the google Colab "content" folder which may take some time depending on the size of your dataset.**
```
#@markdown ##Prepare the data for training
# --------------------- Here we load the augmented data or the raw data ------------------------
if Use_Data_augmentation:
Training_source_dir = Training_source_augmented
Training_target_dir = Training_target_augmented
if not Use_Data_augmentation:
Training_source_dir = Training_source
Training_target_dir = Training_target
# --------------------- ------------------------------------------------
print("Data preparation in progress")
if os.path.exists(model_path+'/'+model_name):
shutil.rmtree(model_path+'/'+model_name)
os.makedirs(model_path+'/'+model_name)
#--------------- Here we move the files to trainA and train B ---------
print('Copying training source data...')
for f in tqdm(os.listdir(Training_source_dir)):
shutil.copyfile(Training_source_dir+"/"+f, TrainA_Folder+"/"+f)
print('Copying training target data...')
for f in tqdm(os.listdir(Training_target_dir)):
shutil.copyfile(Training_target_dir+"/"+f, TrainB_Folder+"/"+f)
#---------------------------------------------------------------------
#--------------- Here we combined A and B images---------
os.chdir("/content")
!python pytorch-CycleGAN-and-pix2pix/datasets/combine_A_and_B.py --fold_A "$imageA_folder" --fold_B "$imageB_folder" --fold_AB "$imageAB_folder"
# pix2pix uses EPOCH without lr decay and EPOCH with lr decay, here we automatically choose half and half
number_of_epochs_lr_stable = int(number_of_epochs/2)
number_of_epochs_lr_decay = int(number_of_epochs/2)
if Use_pretrained_model :
for f in os.listdir(pretrained_model_path):
if (f.startswith("latest_net_")):
shutil.copyfile(pretrained_model_path+"/"+f, model_path+'/'+model_name+"/"+f)
#Export of pdf summary of training parameters
pdf_export(augmentation = Use_Data_augmentation, pretrained_model = Use_pretrained_model)
print('------------------------')
print("Data ready for training")
```
## **4.2. Start Training**
---
<font size = 4>When playing the cell below you should see updates after each epoch (round). Network training can take some time.
<font size = 4>* **CRITICAL NOTE:** Google Colab has a time limit for processing (to prevent using GPU power for datamining). Training time must be less than 12 hours! If training takes longer than 12 hours, please decrease the number of epochs or number of patches or continue the training in a second Colab session. **Pix2pix will save model checkpoints every 5 epochs.**
<font size = 4>Once training is complete, the trained model is automatically saved on your Google Drive, in the **model_path** folder that was selected in Section 3. It is however wise to download the folder as all data can be erased at the next training if using the same folder.
```
#@markdown ##Start training
start = time.time()
os.chdir("/content")
#--------------------------------- Command line inputs to change pix2pix paramaters------------
# basic parameters
#('--dataroot', required=True, help='path to images (should have subfolders trainA, trainB, valA, valB, etc)')
#('--name', type=str, default='experiment_name', help='name of the experiment. It decides where to store samples and models')
#('--gpu_ids', type=str, default='0', help='gpu ids: e.g. 0 0,1,2, 0,2. use -1 for CPU')
#('--checkpoints_dir', type=str, default='./checkpoints', help='models are saved here')
# model parameters
#('--model', type=str, default='cycle_gan', help='chooses which model to use. [cycle_gan | pix2pix | test | colorization]')
#('--input_nc', type=int, default=3, help='# of input image channels: 3 for RGB and 1 for grayscale')
#('--output_nc', type=int, default=3, help='# of output image channels: 3 for RGB and 1 for grayscale')
#('--ngf', type=int, default=64, help='# of gen filters in the last conv layer')
#('--ndf', type=int, default=64, help='# of discrim filters in the first conv layer')
#('--netD', type=str, default='basic', help='specify discriminator architecture [basic | n_layers | pixel]. The basic model is a 70x70 PatchGAN. n_layers allows you to specify the layers in the discriminator')
#('--netG', type=str, default='resnet_9blocks', help='specify generator architecture [resnet_9blocks | resnet_6blocks | unet_256 | unet_128]')
#('--n_layers_D', type=int, default=3, help='only used if netD==n_layers')
#('--norm', type=str, default='instance', help='instance normalization or batch normalization [instance | batch | none]')
#('--init_type', type=str, default='normal', help='network initialization [normal | xavier | kaiming | orthogonal]')
#('--init_gain', type=float, default=0.02, help='scaling factor for normal, xavier and orthogonal.')
#('--no_dropout', action='store_true', help='no dropout for the generator')
# dataset parameters
#('--dataset_mode', type=str, default='unaligned', help='chooses how datasets are loaded. [unaligned | aligned | single | colorization]')
#('--direction', type=str, default='AtoB', help='AtoB or BtoA')
#('--serial_batches', action='store_true', help='if true, takes images in order to make batches, otherwise takes them randomly')
#('--num_threads', default=4, type=int, help='# threads for loading data')
#('--batch_size', type=int, default=1, help='input batch size')
#('--load_size', type=int, default=286, help='scale images to this size')
#('--crop_size', type=int, default=256, help='then crop to this size')
#('--max_dataset_size', type=int, default=float("inf"), help='Maximum number of samples allowed per dataset. If the dataset directory contains more than max_dataset_size, only a subset is loaded.')
#('--preprocess', type=str, default='resize_and_crop', help='scaling and cropping of images at load time [resize_and_crop | crop | scale_width | scale_width_and_crop | none]')
#('--no_flip', action='store_true', help='if specified, do not flip the images for data augmentation')
#('--display_winsize', type=int, default=256, help='display window size for both visdom and HTML')
# additional parameters
#('--epoch', type=str, default='latest', help='which epoch to load? set to latest to use latest cached model')
#('--load_iter', type=int, default='0', help='which iteration to load? if load_iter > 0, the code will load models by iter_[load_iter]; otherwise, the code will load models by [epoch]')
#('--verbose', action='store_true', help='if specified, print more debugging information')
#('--suffix', default='', type=str, help='customized suffix: opt.name = opt.name + suffix: e.g., {model}_{netG}_size{load_size}')
# visdom and HTML visualization parameters
#('--display_freq', type=int, default=400, help='frequency of showing training results on screen')
#('--display_ncols', type=int, default=4, help='if positive, display all images in a single visdom web panel with certain number of images per row.')
#('--display_id', type=int, default=1, help='window id of the web display')
#('--display_server', type=str, default="http://localhost", help='visdom server of the web display')
#('--display_env', type=str, default='main', help='visdom display environment name (default is "main")')
#('--display_port', type=int, default=8097, help='visdom port of the web display')
#('--update_html_freq', type=int, default=1000, help='frequency of saving training results to html')
#('--print_freq', type=int, default=100, help='frequency of showing training results on console')
#('--no_html', action='store_true', help='do not save intermediate training results to [opt.checkpoints_dir]/[opt.name]/web/')
# network saving and loading parameters
#('--save_latest_freq', type=int, default=5000, help='frequency of saving the latest results')
#('--save_epoch_freq', type=int, default=5, help='frequency of saving checkpoints at the end of epochs')
#('--save_by_iter', action='store_true', help='whether saves model by iteration')
#('--continue_train', action='store_true', help='continue training: load the latest model')
#('--epoch_count', type=int, default=1, help='the starting epoch count, we save the model by <epoch_count>, <epoch_count>+<save_latest_freq>, ...')
#('--phase', type=str, default='train', help='train, val, test, etc')
# training parameters
#('--n_epochs', type=int, default=100, help='number of epochs with the initial learning rate')
#('--n_epochs_decay', type=int, default=100, help='number of epochs to linearly decay learning rate to zero')
#('--beta1', type=float, default=0.5, help='momentum term of adam')
#('--lr', type=float, default=0.0002, help='initial learning rate for adam')
#('--gan_mode', type=str, default='lsgan', help='the type of GAN objective. [vanilla| lsgan | wgangp]. vanilla GAN loss is the cross-entropy objective used in the original GAN paper.')
#('--pool_size', type=int, default=50, help='the size of image buffer that stores previously generated images')
#('--lr_policy', type=str, default='linear', help='learning rate policy. [linear | step | plateau | cosine]')
#('--lr_decay_iters', type=int, default=50, help='multiply by a gamma every lr_decay_iters iterations'
#---------------------------------------------------------
#----- Start the training ------------------------------------
if not Use_pretrained_model:
!python pytorch-CycleGAN-and-pix2pix/train.py --dataroot "$imageAB_folder" --name $model_name --model pix2pix --batch_size $batch_size --preprocess scale_width_and_crop --load_size $Image_min_dim --crop_size $patch_size --checkpoints_dir "$model_path" --no_html --n_epochs $number_of_epochs_lr_stable --n_epochs_decay $number_of_epochs_lr_decay --lr $initial_learning_rate --display_id 0 --save_epoch_freq 5
if Use_pretrained_model:
!python pytorch-CycleGAN-and-pix2pix/train.py --dataroot "$imageAB_folder" --name $model_name --model pix2pix --batch_size $batch_size --preprocess scale_width_and_crop --load_size $Image_min_dim --crop_size $patch_size --checkpoints_dir "$model_path" --no_html --n_epochs $number_of_epochs_lr_stable --n_epochs_decay $number_of_epochs_lr_decay --lr $initial_learning_rate --display_id 0 --save_epoch_freq 5 --continue_train
#---------------------------------------------------------
print("Training, done.")
# Displaying the time elapsed for training
dt = time.time() - start
mins, sec = divmod(dt, 60)
hour, mins = divmod(mins, 60)
print("Time elapsed:",hour, "hour(s)",mins,"min(s)",round(sec),"sec(s)")
# Export pdf summary after training to update document
pdf_export(trained = True, augmentation = Use_Data_augmentation, pretrained_model = Use_pretrained_model)
```
# **5. Evaluate your model**
---
<font size = 4>This section allows the user to perform important quality checks on the validity and generalisability of the trained model.
<font size = 4>**We highly recommend to perform quality control on all newly trained models.**
## **5.1. Choose the model you want to assess**
```
# model name and path
#@markdown ###Do you want to assess the model you just trained ?
Use_the_current_trained_model = False #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
QC_model_folder = "" #@param {type:"string"}
#Here we define the loaded model name and path
QC_model_name = os.path.basename(QC_model_folder)
QC_model_path = os.path.dirname(QC_model_folder)
if (Use_the_current_trained_model):
QC_model_name = model_name
QC_model_path = model_path
full_QC_model_path = QC_model_path+'/'+QC_model_name+'/'
if os.path.exists(full_QC_model_path):
print("The "+QC_model_name+" network will be evaluated")
else:
W = '\033[0m' # white (normal)
R = '\033[31m' # red
print(R+'!! WARNING: The chosen model does not exist !!'+W)
print('Please make sure you provide a valid model path and model name before proceeding further.')
```
## **5.2. Identify the best checkpoint to use to make predictions**
<font size = 4> Pix2pix save model checkpoints every five epochs. Due to the stochastic nature of GAN networks, the last checkpoint is not always the best one to use. As a consequence, it can be challenging to choose the most suitable checkpoint to use to make predictions.
<font size = 4>This section allows you to perform predictions using all the saved checkpoints and to estimate the quality of these predictions by comparing them to the provided ground truths images. Metric used include:
<font size = 4>**1. The SSIM (structural similarity) map**
<font size = 4>The SSIM metric is used to evaluate whether two images contain the same structures. It is a normalized metric and an SSIM of 1 indicates a perfect similarity between two images. Therefore for SSIM, the closer to 1, the better. The SSIM maps are constructed by calculating the SSIM metric in each pixel by considering the surrounding structural similarity in the neighbourhood of that pixel (currently defined as window of 11 pixels and with Gaussian weighting of 1.5 pixel standard deviation, see our Wiki for more info).
<font size=4>**mSSIM** is the SSIM value calculated across the entire window of both images.
<font size=4>**The output below shows the SSIM maps with the mSSIM**
<font size = 4>**2. The RSE (Root Squared Error) map**
<font size = 4>This is a display of the root of the squared difference between the normalized predicted and target or the source and the target. In this case, a smaller RSE is better. A perfect agreement between target and prediction will lead to an RSE map showing zeros everywhere (dark).
<font size =4>**NRMSE (normalised root mean squared error)** gives the average difference between all pixels in the images compared to each other. Good agreement yields low NRMSE scores.
<font size = 4>**PSNR (Peak signal-to-noise ratio)** is a metric that gives the difference between the ground truth and prediction (or source input) in decibels, using the peak pixel values of the prediction and the MSE between the images. The higher the score the better the agreement.
<font size=4>**The output below shows the RSE maps with the NRMSE and PSNR values.**
```
#@markdown ##Choose the folders that contain your Quality Control dataset
import glob
import os.path
Source_QC_folder = "" #@param{type:"string"}
Target_QC_folder = "" #@param{type:"string"}
Image_type = "Grayscale" #@param ["Grayscale", "RGB"]
# average function
def Average(lst):
return sum(lst) / len(lst)
# Create a quality control folder
if os.path.exists(QC_model_path+"/"+QC_model_name+"/Quality Control"):
shutil.rmtree(QC_model_path+"/"+QC_model_name+"/Quality Control")
os.makedirs(QC_model_path+"/"+QC_model_name+"/Quality Control")
# Create a quality control/Prediction Folder
QC_prediction_results = QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction"
if os.path.exists(QC_prediction_results):
shutil.rmtree(QC_prediction_results)
os.makedirs(QC_prediction_results)
# Here we count how many images are in our folder to be predicted and we had a few
Nb_files_Data_folder = len(os.listdir(Source_QC_folder)) +10
# List images in Source_QC_folder
# This will find the image dimension of a randomly choosen image in Source_QC_folder
random_choice = random.choice(os.listdir(Source_QC_folder))
x = imageio.imread(Source_QC_folder+"/"+random_choice)
#Find image XY dimension
Image_Y = x.shape[0]
Image_X = x.shape[1]
Image_min_dim = min(Image_Y, Image_X)
# Here we need to move the data to be analysed so that pix2pix can find them
Saving_path_QC= "/content/"+QC_model_name+"_images"
if os.path.exists(Saving_path_QC):
shutil.rmtree(Saving_path_QC)
os.makedirs(Saving_path_QC)
Saving_path_QC_folder = Saving_path_QC+"/QC"
if os.path.exists(Saving_path_QC_folder):
shutil.rmtree(Saving_path_QC_folder)
os.makedirs(Saving_path_QC_folder)
imageA_folder = Saving_path_QC_folder+"/A"
os.makedirs(imageA_folder)
imageB_folder = Saving_path_QC_folder+"/B"
os.makedirs(imageB_folder)
imageAB_folder = Saving_path_QC_folder+"/AB"
os.makedirs(imageAB_folder)
testAB_folder = Saving_path_QC_folder+"/AB/test"
os.makedirs(testAB_folder)
testA_Folder = Saving_path_QC_folder+"/A/test"
os.makedirs(testA_Folder)
testB_Folder = Saving_path_QC_folder+"/B/test"
os.makedirs(testB_Folder)
QC_checkpoint_folders = "/content/"+QC_model_name
if os.path.exists(QC_checkpoint_folders):
shutil.rmtree(QC_checkpoint_folders)
os.makedirs(QC_checkpoint_folders)
for files in os.listdir(Source_QC_folder):
shutil.copyfile(Source_QC_folder+"/"+files, testA_Folder+"/"+files)
for files in os.listdir(Target_QC_folder):
shutil.copyfile(Target_QC_folder+"/"+files, testB_Folder+"/"+files)
#Here we create a merged folder containing only imageA
os.chdir("/content")
!python pytorch-CycleGAN-and-pix2pix/datasets/combine_A_and_B.py --fold_A "$imageA_folder" --fold_B "$imageB_folder" --fold_AB "$imageAB_folder"
# This will find the image dimension of a randomly choosen image in Source_QC_folder
random_choice = random.choice(os.listdir(Source_QC_folder))
x = imageio.imread(Source_QC_folder+"/"+random_choice)
#Find image XY dimension
Image_Y = x.shape[0]
Image_X = x.shape[1]
Image_min_dim = int(min(Image_Y, Image_X))
patch_size_QC = Image_min_dim
if not patch_size_QC % 256 == 0:
patch_size_QC = ((int(patch_size_QC / 256)) * 256)
print (" Your image dimensions are not divisible by 256; therefore your images have now been resized to:",patch_size_QC)
if patch_size_QC < 256:
patch_size_QC = 256
Nb_Checkpoint = len(glob.glob(os.path.join(full_QC_model_path, '*G.pth')))
print(Nb_Checkpoint)
## Initiate list
Checkpoint_list = []
Average_ssim_score_list = []
for j in range(1, len(glob.glob(os.path.join(full_QC_model_path, '*G.pth')))+1):
checkpoints = j*5
if checkpoints == Nb_Checkpoint*5:
checkpoints = "latest"
print("The checkpoint currently analysed is ="+str(checkpoints))
Checkpoint_list.append(checkpoints)
# Create a quality control/Prediction Folder
QC_prediction_results = QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)
if os.path.exists(QC_prediction_results):
shutil.rmtree(QC_prediction_results)
os.makedirs(QC_prediction_results)
# Create a quality control/Prediction Folder
QC_prediction_results = QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)
if os.path.exists(QC_prediction_results):
shutil.rmtree(QC_prediction_results)
os.makedirs(QC_prediction_results)
#---------------------------- Predictions are performed here ----------------------
os.chdir("/content")
!python pytorch-CycleGAN-and-pix2pix/test.py --dataroot "$imageAB_folder" --name "$QC_model_name" --model pix2pix --epoch $checkpoints --no_dropout --preprocess scale_width --load_size $patch_size_QC --crop_size $patch_size_QC --results_dir "$QC_prediction_results" --checkpoints_dir "$QC_model_path" --direction AtoB --num_test $Nb_files_Data_folder
#-----------------------------------------------------------------------------------
#Here we need to move the data again and remove all the unnecessary folders
Checkpoint_name = "test_"+str(checkpoints)
QC_results_images = QC_prediction_results+"/"+QC_model_name+"/"+Checkpoint_name+"/images"
QC_results_images_files = os.listdir(QC_results_images)
for f in QC_results_images_files:
shutil.copyfile(QC_results_images+"/"+f, QC_prediction_results+"/"+f)
os.chdir("/content")
#Here we clean up the extra files
shutil.rmtree(QC_prediction_results+"/"+QC_model_name)
#-------------------------------- QC for RGB ------------------------------------
if Image_type == "RGB":
# List images in Source_QC_folder
# This will find the image dimension of a randomly choosen image in Source_QC_folder
random_choice = random.choice(os.listdir(Source_QC_folder))
x = imageio.imread(Source_QC_folder+"/"+random_choice)
def ssim(img1, img2):
return structural_similarity(img1,img2,data_range=1.,full=True, multichannel=True)
# Open and create the csv file that will contain all the QC metrics
with open(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)+"/"+"QC_metrics_"+QC_model_name+str(checkpoints)+".csv", "w", newline='') as file:
writer = csv.writer(file)
# Write the header in the csv file
writer.writerow(["image #","Prediction v. GT mSSIM","Input v. GT mSSIM"])
# Initiate list
ssim_score_list = []
# Let's loop through the provided dataset in the QC folders
for i in os.listdir(Source_QC_folder):
if not os.path.isdir(os.path.join(Source_QC_folder,i)):
print('Running QC on: '+i)
shortname_no_PNG = i[:-4]
# -------------------------------- Target test data (Ground truth) --------------------------------
test_GT = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), shortname_no_PNG+"_real_B.png"))
# -------------------------------- Source test data --------------------------------
test_source = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints),shortname_no_PNG+"_real_A.png"))
# -------------------------------- Prediction --------------------------------
test_prediction = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints),shortname_no_PNG+"_fake_B.png"))
#--------------------------- Here we normalise using histograms matching--------------------------------
test_prediction_matched = match_histograms(test_prediction, test_GT, multichannel=True)
test_source_matched = match_histograms(test_source, test_GT, multichannel=True)
# -------------------------------- Calculate the metric maps and save them --------------------------------
# Calculate the SSIM maps
index_SSIM_GTvsPrediction, img_SSIM_GTvsPrediction = ssim(test_GT, test_prediction_matched)
index_SSIM_GTvsSource, img_SSIM_GTvsSource = ssim(test_GT, test_source_matched)
ssim_score_list.append(index_SSIM_GTvsPrediction)
#Save ssim_maps
img_SSIM_GTvsPrediction_8bit = (img_SSIM_GTvsPrediction* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/SSIM_GTvsPrediction_"+shortname_no_PNG+'.tif',img_SSIM_GTvsPrediction_8bit)
img_SSIM_GTvsSource_8bit = (img_SSIM_GTvsSource* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/SSIM_GTvsSource_"+shortname_no_PNG+'.tif',img_SSIM_GTvsSource_8bit)
writer.writerow([i,str(index_SSIM_GTvsPrediction),str(index_SSIM_GTvsSource)])
#Here we calculate the ssim average for each image in each checkpoints
Average_SSIM_checkpoint = Average(ssim_score_list)
Average_ssim_score_list.append(Average_SSIM_checkpoint)
#------------------------------------------- QC for Grayscale ----------------------------------------------
if Image_type == "Grayscale":
def ssim(img1, img2):
return structural_similarity(img1,img2,data_range=1.,full=True, gaussian_weights=True, use_sample_covariance=False, sigma=1.5)
def normalize(x, pmin=3, pmax=99.8, axis=None, clip=False, eps=1e-20, dtype=np.float32):
mi = np.percentile(x,pmin,axis=axis,keepdims=True)
ma = np.percentile(x,pmax,axis=axis,keepdims=True)
return normalize_mi_ma(x, mi, ma, clip=clip, eps=eps, dtype=dtype)
def normalize_mi_ma(x, mi, ma, clip=False, eps=1e-20, dtype=np.float32):#dtype=np.float32
if dtype is not None:
x = x.astype(dtype,copy=False)
mi = dtype(mi) if np.isscalar(mi) else mi.astype(dtype,copy=False)
ma = dtype(ma) if np.isscalar(ma) else ma.astype(dtype,copy=False)
eps = dtype(eps)
try:
import numexpr
x = numexpr.evaluate("(x - mi) / ( ma - mi + eps )")
except ImportError:
x = (x - mi) / ( ma - mi + eps )
if clip:
x = np.clip(x,0,1)
return x
def norm_minmse(gt, x, normalize_gt=True):
if normalize_gt:
gt = normalize(gt, 0.1, 99.9, clip=False).astype(np.float32, copy = False)
x = x.astype(np.float32, copy=False) - np.mean(x)
#x = x - np.mean(x)
gt = gt.astype(np.float32, copy=False) - np.mean(gt)
#gt = gt - np.mean(gt)
scale = np.cov(x.flatten(), gt.flatten())[0, 1] / np.var(x.flatten())
return gt, scale * x
# Open and create the csv file that will contain all the QC metrics
with open(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)+"/"+"QC_metrics_"+QC_model_name+str(checkpoints)+".csv", "w", newline='') as file:
writer = csv.writer(file)
# Write the header in the csv file
writer.writerow(["image #","Prediction v. GT mSSIM","Input v. GT mSSIM", "Prediction v. GT NRMSE", "Input v. GT NRMSE", "Prediction v. GT PSNR", "Input v. GT PSNR"])
# Let's loop through the provided dataset in the QC folders
for i in os.listdir(Source_QC_folder):
if not os.path.isdir(os.path.join(Source_QC_folder,i)):
print('Running QC on: '+i)
ssim_score_list = []
shortname_no_PNG = i[:-4]
# -------------------------------- Target test data (Ground truth) --------------------------------
test_GT_raw = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), shortname_no_PNG+"_real_B.png"))
test_GT = test_GT_raw[:,:,2]
# -------------------------------- Source test data --------------------------------
test_source_raw = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints),shortname_no_PNG+"_real_A.png"))
test_source = test_source_raw[:,:,2]
# Normalize the images wrt each other by minimizing the MSE between GT and Source image
test_GT_norm,test_source_norm = norm_minmse(test_GT, test_source, normalize_gt=True)
# -------------------------------- Prediction --------------------------------
test_prediction_raw = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints),shortname_no_PNG+"_fake_B.png"))
test_prediction = test_prediction_raw[:,:,2]
# Normalize the images wrt each other by minimizing the MSE between GT and prediction
test_GT_norm,test_prediction_norm = norm_minmse(test_GT, test_prediction, normalize_gt=True)
# -------------------------------- Calculate the metric maps and save them --------------------------------
# Calculate the SSIM maps
index_SSIM_GTvsPrediction, img_SSIM_GTvsPrediction = ssim(test_GT_norm, test_prediction_norm)
index_SSIM_GTvsSource, img_SSIM_GTvsSource = ssim(test_GT_norm, test_source_norm)
ssim_score_list.append(index_SSIM_GTvsPrediction)
#Save ssim_maps
img_SSIM_GTvsPrediction_8bit = (img_SSIM_GTvsPrediction* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/SSIM_GTvsPrediction_"+shortname_no_PNG+'.tif',img_SSIM_GTvsPrediction_8bit)
img_SSIM_GTvsSource_8bit = (img_SSIM_GTvsSource* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/SSIM_GTvsSource_"+shortname_no_PNG+'.tif',img_SSIM_GTvsSource_8bit)
# Calculate the Root Squared Error (RSE) maps
img_RSE_GTvsPrediction = np.sqrt(np.square(test_GT_norm - test_prediction_norm))
img_RSE_GTvsSource = np.sqrt(np.square(test_GT_norm - test_source_norm))
# Save SE maps
img_RSE_GTvsPrediction_8bit = (img_RSE_GTvsPrediction* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/RSE_GTvsPrediction_"+shortname_no_PNG+'.tif',img_RSE_GTvsPrediction_8bit)
img_RSE_GTvsSource_8bit = (img_RSE_GTvsSource* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/RSE_GTvsSource_"+shortname_no_PNG+'.tif',img_RSE_GTvsSource_8bit)
# -------------------------------- Calculate the RSE metrics and save them --------------------------------
# Normalised Root Mean Squared Error (here it's valid to take the mean of the image)
NRMSE_GTvsPrediction = np.sqrt(np.mean(img_RSE_GTvsPrediction))
NRMSE_GTvsSource = np.sqrt(np.mean(img_RSE_GTvsSource))
# We can also measure the peak signal to noise ratio between the images
PSNR_GTvsPrediction = psnr(test_GT_norm,test_prediction_norm,data_range=1.0)
PSNR_GTvsSource = psnr(test_GT_norm,test_source_norm,data_range=1.0)
writer.writerow([i,str(index_SSIM_GTvsPrediction),str(index_SSIM_GTvsSource),str(NRMSE_GTvsPrediction),str(NRMSE_GTvsSource),str(PSNR_GTvsPrediction),str(PSNR_GTvsSource)])
#Here we calculate the ssim average for each image in each checkpoints
Average_SSIM_checkpoint = Average(ssim_score_list)
Average_ssim_score_list.append(Average_SSIM_checkpoint)
# All data is now processed saved
# -------------------------------- Display --------------------------------
# Display the IoV vs Threshold plot
plt.figure(figsize=(20,5))
plt.plot(Checkpoint_list, Average_ssim_score_list, label="SSIM")
plt.title('Checkpoints vs. SSIM')
plt.ylabel('SSIM')
plt.xlabel('Checkpoints')
plt.legend()
plt.savefig(full_QC_model_path+'/Quality Control/SSIMvsCheckpoint_data.png',bbox_inches='tight',pad_inches=0)
plt.show()
# -------------------------------- Display RGB --------------------------------
from ipywidgets import interact
import ipywidgets as widgets
if Image_type == "RGB":
random_choice_shortname_no_PNG = shortname_no_PNG
@interact
def show_results(file=os.listdir(Source_QC_folder), checkpoints=Checkpoint_list):
random_choice_shortname_no_PNG = file[:-4]
df1 = pd.read_csv(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)+"/"+"QC_metrics_"+QC_model_name+str(checkpoints)+".csv", header=0)
df2 = df1.set_index("image #", drop = False)
index_SSIM_GTvsPrediction = df2.loc[file, "Prediction v. GT mSSIM"]
index_SSIM_GTvsSource = df2.loc[file, "Input v. GT mSSIM"]
#Setting up colours
cmap = None
plt.figure(figsize=(15,15))
# Target (Ground-truth)
plt.subplot(3,3,1)
plt.axis('off')
img_GT = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_real_B.png"), as_gray=False, pilmode="RGB")
plt.imshow(img_GT, cmap = cmap)
plt.title('Target',fontsize=15)
# Source
plt.subplot(3,3,2)
plt.axis('off')
img_Source = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_real_A.png"), as_gray=False, pilmode="RGB")
plt.imshow(img_Source, cmap = cmap)
plt.title('Source',fontsize=15)
#Prediction
plt.subplot(3,3,3)
plt.axis('off')
img_Prediction = io.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_fake_B.png"))
plt.imshow(img_Prediction, cmap = cmap)
plt.title('Prediction',fontsize=15)
#SSIM between GT and Source
plt.subplot(3,3,5)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_SSIM_GTvsSource = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "SSIM_GTvsSource_"+random_choice_shortname_no_PNG+".tif"))
imSSIM_GTvsSource = plt.imshow(img_SSIM_GTvsSource, cmap = cmap, vmin=0, vmax=1)
#plt.colorbar(imSSIM_GTvsSource,fraction=0.046, pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsSource,3)),fontsize=14)
plt.ylabel('SSIM maps',fontsize=20, rotation=0, labelpad=75)
#SSIM between GT and Prediction
plt.subplot(3,3,6)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_SSIM_GTvsPrediction = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "SSIM_GTvsPrediction_"+random_choice_shortname_no_PNG+".tif"))
imSSIM_GTvsPrediction = plt.imshow(img_SSIM_GTvsPrediction, cmap = cmap, vmin=0,vmax=1)
#plt.colorbar(imSSIM_GTvsPrediction,fraction=0.046, pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsPrediction,3)),fontsize=14)
plt.savefig(full_QC_model_path+'/Quality Control/QC_example_data.png',bbox_inches='tight',pad_inches=0)
# -------------------------------- Display Grayscale --------------------------------
if Image_type == "Grayscale":
random_choice_shortname_no_PNG = shortname_no_PNG
@interact
def show_results(file=os.listdir(Source_QC_folder), checkpoints=Checkpoint_list):
random_choice_shortname_no_PNG = file[:-4]
df1 = pd.read_csv(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)+"/"+"QC_metrics_"+QC_model_name+str(checkpoints)+".csv", header=0)
df2 = df1.set_index("image #", drop = False)
index_SSIM_GTvsPrediction = df2.loc[file, "Prediction v. GT mSSIM"]
index_SSIM_GTvsSource = df2.loc[file, "Input v. GT mSSIM"]
NRMSE_GTvsPrediction = df2.loc[file, "Prediction v. GT NRMSE"]
NRMSE_GTvsSource = df2.loc[file, "Input v. GT NRMSE"]
PSNR_GTvsSource = df2.loc[file, "Input v. GT PSNR"]
PSNR_GTvsPrediction = df2.loc[file, "Prediction v. GT PSNR"]
plt.figure(figsize=(20,20))
# Currently only displays the last computed set, from memory
# Target (Ground-truth)
plt.subplot(3,3,1)
plt.axis('off')
img_GT = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_real_B.png"))
plt.imshow(img_GT, norm=simple_norm(img_GT, percent = 99))
plt.title('Target',fontsize=15)
# Source
plt.subplot(3,3,2)
plt.axis('off')
img_Source = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_real_A.png"))
plt.imshow(img_Source, norm=simple_norm(img_Source, percent = 99))
plt.title('Source',fontsize=15)
#Prediction
plt.subplot(3,3,3)
plt.axis('off')
img_Prediction = io.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_fake_B.png"))
plt.imshow(img_Prediction, norm=simple_norm(img_Prediction, percent = 99))
plt.title('Prediction',fontsize=15)
#Setting up colours
cmap = plt.cm.CMRmap
#SSIM between GT and Source
plt.subplot(3,3,5)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_SSIM_GTvsSource = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "SSIM_GTvsSource_"+random_choice_shortname_no_PNG+".tif"))
img_SSIM_GTvsSource = img_SSIM_GTvsSource / 255
imSSIM_GTvsSource = plt.imshow(img_SSIM_GTvsSource, cmap = cmap, vmin=0, vmax=1)
plt.colorbar(imSSIM_GTvsSource,fraction=0.046, pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsSource,3)),fontsize=14)
plt.ylabel('SSIM maps',fontsize=20, rotation=0, labelpad=75)
#SSIM between GT and Prediction
plt.subplot(3,3,6)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_SSIM_GTvsPrediction = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "SSIM_GTvsPrediction_"+random_choice_shortname_no_PNG+".tif"))
img_SSIM_GTvsPrediction = img_SSIM_GTvsPrediction / 255
imSSIM_GTvsPrediction = plt.imshow(img_SSIM_GTvsPrediction, cmap = cmap, vmin=0,vmax=1)
plt.colorbar(imSSIM_GTvsPrediction,fraction=0.046, pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsPrediction,3)),fontsize=14)
#Root Squared Error between GT and Source
plt.subplot(3,3,8)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_RSE_GTvsSource = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "RSE_GTvsSource_"+random_choice_shortname_no_PNG+".tif"))
img_RSE_GTvsSource = img_RSE_GTvsSource / 255
imRSE_GTvsSource = plt.imshow(img_RSE_GTvsSource, cmap = cmap, vmin=0, vmax = 1)
plt.colorbar(imRSE_GTvsSource,fraction=0.046,pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('NRMSE: '+str(round(NRMSE_GTvsSource,3))+', PSNR: '+str(round(PSNR_GTvsSource,3)),fontsize=14)
#plt.title('Target vs. Source PSNR: '+str(round(PSNR_GTvsSource,3)))
plt.ylabel('RSE maps',fontsize=20, rotation=0, labelpad=75)
#Root Squared Error between GT and Prediction
plt.subplot(3,3,9)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_RSE_GTvsPrediction = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "RSE_GTvsPrediction_"+random_choice_shortname_no_PNG+".tif"))
img_RSE_GTvsPrediction = img_RSE_GTvsPrediction / 255
imRSE_GTvsPrediction = plt.imshow(img_RSE_GTvsPrediction, cmap = cmap, vmin=0, vmax=1)
plt.colorbar(imRSE_GTvsPrediction,fraction=0.046,pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('NRMSE: '+str(round(NRMSE_GTvsPrediction,3))+', PSNR: '+str(round(PSNR_GTvsPrediction,3)),fontsize=14)
plt.savefig(full_QC_model_path+'/Quality Control/QC_example_data.png',bbox_inches='tight',pad_inches=0)
#Make a pdf summary of the QC results
qc_pdf_export()
```
# **6. Using the trained model**
---
<font size = 4>In this section the unseen data is processed using the trained model (in section 4). First, your unseen images are uploaded and prepared for prediction. After that your trained model from section 4 is activated and finally saved into your Google Drive.
## **6.1. Generate prediction(s) from unseen dataset**
---
<font size = 4>The current trained model (from section 4.2) can now be used to process images. If you want to use an older model, untick the **Use_the_current_trained_model** box and enter the name and path of the model to use. Predicted output images are saved in your **Result_folder** folder as PNG images.
<font size = 4>**`Data_folder`:** This folder should contain the images that you want to use your trained network on for processing.
<font size = 4>**`Result_folder`:** This folder will contain the predicted output images.
<font size = 4>**`checkpoint`:** Choose the checkpoint number you would like to use to perform predictions. To use the "latest" checkpoint, input "latest".
```
#@markdown ### Provide the path to your dataset and to the folder where the predictions are saved, then play the cell to predict outputs from your unseen images.
import glob
import os.path
latest = "latest"
Data_folder = "" #@param {type:"string"}
Result_folder = "" #@param {type:"string"}
# model name and path
#@markdown ###Do you want to use the current trained model?
Use_the_current_trained_model = True #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
Prediction_model_folder = "" #@param {type:"string"}
#@markdown ###What model checkpoint would you like to use?
checkpoint = latest#@param {type:"raw"}
#Here we find the loaded model name and parent path
Prediction_model_name = os.path.basename(Prediction_model_folder)
Prediction_model_path = os.path.dirname(Prediction_model_folder)
#here we check if we use the newly trained network or not
if (Use_the_current_trained_model):
print("Using current trained network")
Prediction_model_name = model_name
Prediction_model_path = model_path
#here we check if the model exists
full_Prediction_model_path = Prediction_model_path+'/'+Prediction_model_name+'/'
if os.path.exists(full_Prediction_model_path):
print("The "+Prediction_model_name+" network will be used.")
else:
W = '\033[0m' # white (normal)
R = '\033[31m' # red
print(R+'!! WARNING: The chosen model does not exist !!'+W)
print('Please make sure you provide a valid model path and model name before proceeding further.')
Nb_Checkpoint = len(glob.glob(os.path.join(full_Prediction_model_path, '*G.pth')))+1
if not checkpoint == "latest":
if checkpoint < 10:
checkpoint = 5
if not checkpoint % 5 == 0:
checkpoint = ((int(checkpoint / 5)-1) * 5)
print (bcolors.WARNING + " Your chosen checkpoints is not divisible by 5; therefore the checkpoints chosen is now:",checkpoints)
if checkpoint == Nb_Checkpoint*5:
checkpoint = "latest"
if checkpoint > Nb_Checkpoint*5:
checkpoint = "latest"
# Here we need to move the data to be analysed so that pix2pix can find them
Saving_path_prediction= "/content/"+Prediction_model_name
if os.path.exists(Saving_path_prediction):
shutil.rmtree(Saving_path_prediction)
os.makedirs(Saving_path_prediction)
imageA_folder = Saving_path_prediction+"/A"
os.makedirs(imageA_folder)
imageB_folder = Saving_path_prediction+"/B"
os.makedirs(imageB_folder)
imageAB_folder = Saving_path_prediction+"/AB"
os.makedirs(imageAB_folder)
testAB_Folder = Saving_path_prediction+"/AB/test"
os.makedirs(testAB_Folder)
testA_Folder = Saving_path_prediction+"/A/test"
os.makedirs(testA_Folder)
testB_Folder = Saving_path_prediction+"/B/test"
os.makedirs(testB_Folder)
for files in os.listdir(Data_folder):
shutil.copyfile(Data_folder+"/"+files, testA_Folder+"/"+files)
shutil.copyfile(Data_folder+"/"+files, testB_Folder+"/"+files)
# Here we create a merged A / A image for the prediction
os.chdir("/content")
!python pytorch-CycleGAN-and-pix2pix/datasets/combine_A_and_B.py --fold_A "$imageA_folder" --fold_B "$imageB_folder" --fold_AB "$imageAB_folder"
# Here we count how many images are in our folder to be predicted and we had a few
Nb_files_Data_folder = len(os.listdir(Data_folder)) +10
# This will find the image dimension of a randomly choosen image in Data_folder
random_choice = random.choice(os.listdir(Data_folder))
x = imageio.imread(Data_folder+"/"+random_choice)
#Find image XY dimension
Image_Y = x.shape[0]
Image_X = x.shape[1]
Image_min_dim = min(Image_Y, Image_X)
#-------------------------------- Perform predictions -----------------------------
#-------------------------------- Options that can be used to perform predictions -----------------------------
# basic parameters
#('--dataroot', required=True, help='path to images (should have subfolders trainA, trainB, valA, valB, etc)')
#('--name', type=str, default='experiment_name', help='name of the experiment. It decides where to store samples and models')
#('--gpu_ids', type=str, default='0', help='gpu ids: e.g. 0 0,1,2, 0,2. use -1 for CPU')
#('--checkpoints_dir', type=str, default='./checkpoints', help='models are saved here')
# model parameters
#('--model', type=str, default='cycle_gan', help='chooses which model to use. [cycle_gan | pix2pix | test | colorization]')
#('--input_nc', type=int, default=3, help='# of input image channels: 3 for RGB and 1 for grayscale')
#('--output_nc', type=int, default=3, help='# of output image channels: 3 for RGB and 1 for grayscale')
#('--ngf', type=int, default=64, help='# of gen filters in the last conv layer')
#('--ndf', type=int, default=64, help='# of discrim filters in the first conv layer')
#('--netD', type=str, default='basic', help='specify discriminator architecture [basic | n_layers | pixel]. The basic model is a 70x70 PatchGAN. n_layers allows you to specify the layers in the discriminator')
#('--netG', type=str, default='resnet_9blocks', help='specify generator architecture [resnet_9blocks | resnet_6blocks | unet_256 | unet_128]')
#('--n_layers_D', type=int, default=3, help='only used if netD==n_layers')
#('--norm', type=str, default='instance', help='instance normalization or batch normalization [instance | batch | none]')
#('--init_type', type=str, default='normal', help='network initialization [normal | xavier | kaiming | orthogonal]')
#('--init_gain', type=float, default=0.02, help='scaling factor for normal, xavier and orthogonal.')
#('--no_dropout', action='store_true', help='no dropout for the generator')
# dataset parameters
#('--dataset_mode', type=str, default='unaligned', help='chooses how datasets are loaded. [unaligned | aligned | single | colorization]')
#('--direction', type=str, default='AtoB', help='AtoB or BtoA')
#('--serial_batches', action='store_true', help='if true, takes images in order to make batches, otherwise takes them randomly')
#('--num_threads', default=4, type=int, help='# threads for loading data')
#('--batch_size', type=int, default=1, help='input batch size')
#('--load_size', type=int, default=286, help='scale images to this size')
#('--crop_size', type=int, default=256, help='then crop to this size')
#('--max_dataset_size', type=int, default=float("inf"), help='Maximum number of samples allowed per dataset. If the dataset directory contains more than max_dataset_size, only a subset is loaded.')
#('--preprocess', type=str, default='resize_and_crop', help='scaling and cropping of images at load time [resize_and_crop | crop | scale_width | scale_width_and_crop | none]')
#('--no_flip', action='store_true', help='if specified, do not flip the images for data augmentation')
#('--display_winsize', type=int, default=256, help='display window size for both visdom and HTML')
# additional parameters
#('--epoch', type=str, default='latest', help='which epoch to load? set to latest to use latest cached model')
#('--load_iter', type=int, default='0', help='which iteration to load? if load_iter > 0, the code will load models by iter_[load_iter]; otherwise, the code will load models by [epoch]')
#('--verbose', action='store_true', help='if specified, print more debugging information')
#('--suffix', default='', type=str, help='customized suffix: opt.name = opt.name + suffix: e.g., {model}_{netG}_size{load_size}')
#('--ntest', type=int, default=float("inf"), help='# of test examples.')
#('--results_dir', type=str, default='./results/', help='saves results here.')
#('--aspect_ratio', type=float, default=1.0, help='aspect ratio of result images')
#('--phase', type=str, default='test', help='train, val, test, etc')
# Dropout and Batchnorm has different behavioir during training and test.
#('--eval', action='store_true', help='use eval mode during test time.')
#('--num_test', type=int, default=50, help='how many test images to run')
# rewrite devalue values
# To avoid cropping, the load_size should be the same as crop_size
#parser.set_defaults(load_size=parser.get_default('crop_size'))
#------------------------------------------------------------------------
#---------------------------- Predictions are performed here ----------------------
os.chdir("/content")
!python pytorch-CycleGAN-and-pix2pix/test.py --dataroot "$imageAB_folder" --name "$Prediction_model_name" --model pix2pix --no_dropout --preprocess scale_width --load_size $Image_min_dim --crop_size $Image_min_dim --results_dir "$Result_folder" --checkpoints_dir "$Prediction_model_path" --num_test $Nb_files_Data_folder --epoch $checkpoint
#-----------------------------------------------------------------------------------
Checkpoint_name = "test_"+str(checkpoint)
Prediction_results_folder = Result_folder+"/"+Prediction_model_name+"/"+Checkpoint_name+"/images"
Prediction_results_images = os.listdir(Prediction_results_folder)
for f in Prediction_results_images:
if (f.endswith("_real_B.png")):
os.remove(Prediction_results_folder+"/"+f)
```
## **6.2. Inspect the predicted output**
---
```
# @markdown ##Run this cell to display a randomly chosen input and its corresponding predicted output.
import os
# This will display a randomly chosen dataset input and predicted output
random_choice = random.choice(os.listdir(Data_folder))
random_choice_no_extension = os.path.splitext(random_choice)
x = imageio.imread(Result_folder+"/"+Prediction_model_name+"/test_"+str(checkpoint)+"/images/"+random_choice_no_extension[0]+"_real_A.png")
y = imageio.imread(Result_folder+"/"+Prediction_model_name+"/test_"+str(checkpoint)+"/images/"+random_choice_no_extension[0]+"_fake_B.png")
f=plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.imshow(x, interpolation='nearest')
plt.title('Input')
plt.axis('off');
plt.subplot(1,2,2)
plt.imshow(y, interpolation='nearest')
plt.title('Prediction')
plt.axis('off');
```
## **6.3. Download your predictions**
---
<font size = 4>**Store your data** and ALL its results elsewhere by downloading it from Google Drive and after that clean the original folder tree (datasets, results, trained model etc.) if you plan to train or use new networks. Please note that the notebook will otherwise **OVERWRITE** all files which have the same name.
#**Thank you for using pix2pix!**
| github_jupyter |
# ML Pipeline Preparation
Follow the instructions below to help you create your ML pipeline.
### 1. Import libraries and load data from database.
- Import Python libraries
- Load dataset from database with [`read_sql_table`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_sql_table.html)
- Define feature and target variables X and Y
```
# import libraries
from sqlalchemy import create_engine
import pandas as pd
import re
import numpy as np
import pickle
from sklearn.multioutput import MultiOutputClassifier
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.pipeline import Pipeline
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import LinearSVC
from sklearn.metrics import f1_score, classification_report
from lightgbm import LGBMClassifier
import nltk
from nltk.corpus import stopwords
from nltk.stem.wordnet import WordNetLemmatizer
from nltk.tokenize import word_tokenize
nltk.download('punkt')
nltk.download('stopwords')
nltk.download('wordnet')
# load data from database
engine = create_engine('sqlite:///data/DisasterResponse.db')
df = pd.read_sql('select * from messages', engine)
df.head()
# selecting feature column
X = df['message']
X.head()
# selecting label columns
Y = df[[col_name for col_name in df.columns if col_name not in ['id', 'message', 'original', 'genre']]]
Y.head()
```
### 2. Write a tokenization function to process your text data
```
def tokenize(text):
'''
Create word tokens using a input text.
- Lower case
- Remove stopwords
- Remove ponctuation
- Lemmatize
Parameters
----------
text : string
Text you want to tokenize.
Returns
-------
tokens : list
List of tokens
'''
# put all letter to lower case
text = text.lower()
# substitute everything that is not letters or numbers
text = re.sub('[^a-z 0-9]', ' ', text)
# create tokens using nltk
tokens = word_tokenize(text)
# load stopwords
stop = stopwords.words('english')
# remove stopwords and lemmatize
tokens = [WordNetLemmatizer().lemmatize(word) for word in tokens if word not in stop]
return tokens
# testing tokenize function
print(X[0])
print(tokenize(X[0]))
```
### 3. Build a machine learning pipeline
This machine pipeline should take in the `message` column as input and output classification results on the other 36 categories in the dataset. You may find the [MultiOutputClassifier](http://scikit-learn.org/stable/modules/generated/sklearn.multioutput.MultiOutputClassifier.html) helpful for predicting multiple target variables.
```
pipeline = Pipeline([('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', MultiOutputClassifier(RandomForestClassifier()))])
```
### 4. Train pipeline
- Split data into train and test sets
- Train pipeline
```
# spliting and fitting the model
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.33, random_state=42)
pipeline.fit(X_train, y_train)
```
### 5. Test your model
Report the f1 score, precision and recall for each output category of the dataset. You can do this by iterating through the columns and calling sklearn's `classification_report` on each.
```
# predicting test data
y_pred = pipeline.predict(X_test)
# printing f1_score
print(classification_report(y_test, y_pred))
```
### 6. Improve your model
Use grid search to find better parameters.
```
# create a grid search to test hyperparameters, I have to test a little because training is so slow
parameters = {'clf__estimator__n_estimators': [10, 50, 100]}
cv = GridSearchCV(pipeline, parameters)
cv.fit(X_train, y_train)
```
### 7. Test your model
Show the accuracy, precision, and recall of the tuned model.
Since this project focuses on code quality, process, and pipelines, there is no minimum performance metric needed to pass. However, make sure to fine tune your models for accuracy, precision and recall to make your project stand out - especially for your portfolio!
```
# show results of grid search
cv.cv_results_
```
### 8. Try improving your model further. Here are a few ideas:
* try other machine learning algorithms
* add other features besides the TF-IDF
```
# trying linear SVC model
pipeline_svc = Pipeline([('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', MultiOutputClassifier(LinearSVC()))])
# fitting model. Note: I have to drop 'child alone' label for this model, it was getting an error
pipeline_svc.fit(X_train, y_train.drop(labels=['child_alone'], axis=1))
# predicting test data
y_pred_svc = pipeline_svc.predict(X_test)
# printing results
print(classification_report(y_test.drop(labels=['child_alone'], axis=1), y_pred_svc))
# testing another hyperparameters using grid search
parameters = {'clf__estimator__penalty': ['l1', 'l2'],
'clf__estimator__C': [0.001, 0.01, 0.1, 1, 10, 100, 1000, 10000]}
cv = GridSearchCV(pipeline_svc, parameters)
cv.fit(X_train, y_train.drop(labels=['child_alone'], axis=1))
# printing f1_score for best model
y_pred_svc = cv.predict(X_test)
print(classification_report(y_test.drop(labels=['child_alone'], axis=1), y_pred_svc))
# printing results of grid search
cv.cv_results_
# well it looks like the best model was standard hyperparameter of the model
cv.best_params_
```
### 8.b. Trying LightGBM
```
# try using LightGBM
pipeline_lgbm = Pipeline([('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', MultiOutputClassifier(LGBMClassifier()))])
# fitting model
pipeline_lgbm.fit(X_train, y_train)
# predicting test data and printing f1_score
y_pred_lgbm = pipeline_lgbm.predict(X_test)
print(classification_report(y_test, y_pred_lgbm))
# trying some change in hyperparameters
parameters = {'clf__estimator__boosting_type': ['gbdt', 'dart', 'goss', 'rf']}
cv = GridSearchCV(pipeline_lgbm, parameters)
cv.fit(X_train, y_train)
# checking the best hyperparameters after grid search
cv.cv_results_
```
### 9. Export your model as a pickle file
```
pickle.dump(pipeline_lgbm, open('model.pkl', 'wb'))
```
### 10. Use this notebook to complete `train.py`
Use the template file attached in the Resources folder to write a script that runs the steps above to create a database and export a model based on a new dataset specified by the user.
| github_jupyter |
[](https://www.ibm.com/demos/collection/db2-database/)
<a id="top">
# Using the Open Db2 Data Management Console RESTful Service APIs
Welcome to this Db2 Data Management Console lab that highlights the RESTful services of the console. This lab uses Jupyter notebooks to demonstrate these features. If you are not familiar with the use of Jupyter notebooks or Python, the following notebooks will guide you through their usage. You can find a copy of these notebooks at https://github.com/Db2-DTE-POC/db2dmc.
<div style="font-family: 'IBM Plex Sans';">
<table style="float:left; width: 620px; height: 235px; border-spacing: 10px; border-collapse: separate; table-layout: fixed">
<td style="padding: 15px; text-align:left; vertical-align: text-top; background-color:#F7F7F7; width: 300px; height: 250px;">
<div style="height: 75px"><p style="font-size: 24px">An Introduction to Jupyter Notebooks</div>
<div style="height: 125px"><p style="font-size: 14px">
If you are not familiar with the use of Jupyter notebooks or Python, the following notebook
will guide you through their usage.
</div>
<div style="height: 25px"><p style="font-size: 12px; text-align: right">
<img style="display: inline-block;"src="./media/clock.png"> 10min
</div>
<div style="height: 10px"><p style="font-size: 12px; text-align: right">
<a href="http://localhost:8888/notebooks/An_Introduction_to_Jupyter_Notebooks.ipynb">
<img style="display: inline-block;"src="./media/arrowblue.png"></a>
</div>
</td>
<td style="padding: 15px; text-align:left; vertical-align: text-top; background-color:#F7F7F7; width: 300px; height:250px">
<div style="height: 75px"><p style="font-size: 24px">Db2 Magic Commands</div>
<div style="height: 125px"><p style="font-size: 14px">
Db2 Magic commands are used in all of the notebooks used in this lab.
The following notebook provides a tutorial on basics of using the Db2 magic commands.
</div>
<div style="height: 25px"><p style="font-size: 12px; text-align: right">
<img style="display: inline-block;"src="./media/clock.png"> 10min
</div>
<div style="height: 10px"><p style="font-size: 12px; text-align: right">
<a href="http://localhost:8888/notebooks/Db2_Jupyter_Extensions_Tutorial.ipynb">
<img style="display: inline-block;"src="./media/arrowblue.png"></a>
</div>
</td>
<td style="padding: 15px; text-align:left; vertical-align: text-top; background-color:#F7F7F7; width: 300px; height:250px">
<div style="height: 75px"><p style="font-size: 24px">The Db2 Data Management Console</div>
<div style="height: 125px"><p style="font-size: 14px">
This is an introduction to the Db2 Data Management Console.
It is more than a graphical user interface.
It is a set of microservices that you can use to build custom solutions to automate Db2.
</div>
<div style="height: 25px"><p style="font-size: 12px; text-align: right">
<img style="display: inline-block;"src="./media/clock.png"> 10min
</div>
<div style="height: 10px"><p style="font-size: 12px; text-align: right">
<a href="http://localhost:8888/notebooks/Db2_Data_Management_Console_Introduction.ipynb">
<img style="display: inline-block;"src="./media/arrowblue.png"></a>
</div>
</td>
</table>
</div>
Everything we do through the Db2 Data Management Console interface goes through a new RESTful API that is open and available. That includes APIs to run SQL, Manage Database Objects and Privileges, Monitor performance, load data and files and configure all aspects of the console. The following labs illustrates the use of these RESTful APIs.
<!-- Row 1 -->
<div style="font-family: 'IBM Plex Sans';">
<table style="float:left; width: 620px; height: 235px; border-spacing: 10px; border-collapse: separate; table-layout: fixed">
<td style="padding: 15px; text-align:left; vertical-align: text-top; background-color:#F7F7F7; width: 300px; height: 250px;">
<div style="height: 75px"><p style="font-size: 24px">
<!-- Title -->
Db2 SQL with RESTful Services
</div>
<div style="height: 125px"><p style="font-size: 14px">
<!-- Description -->
The Db2 Data Management Console includes a number of RESTful services including those which allow for SQL execution. This lab takes the user through the steps required to query Db2 using RESTful APIs.
</div>
<div style="height: 25px"><p style="font-size: 12px; text-align: right">
<img style="display: inline-block;"src="./media/clock.png">
<!-- Duration -->
15 min
</div>
<div style="height: 10px"><p style="font-size: 12px; text-align: right">
<!-- URL -->
<a href="http://localhost:8888/notebooks/Db2_RESTful_APIS.ipynb">
<img style="display: inline-block;"src="./media/arrowblue.png"></a>
</div>
</td>
<td style="padding: 15px; text-align:left; vertical-align: text-top; background-color:#F7F7F7; width: 300px; height: 250px;">
<div style="height: 75px"><p style="font-size: 24px">
<!-- Title -->
Automate Db2 with Open Console Services
</div>
<div style="height: 120px"><p style="font-size: 14px">
<!-- Description -->
This lab contains examples of how to use the Open APIs and composable interfaces that is available through the Db2 Data Management Console service. Everything in the User Interface is also available through an open and fully documented RESTful Services API.
</div>
<div style="height: 25px"><p style="font-size: 12px; text-align: right">
<img style="display: inline-block;"src="./media/clock.png">
<!-- Duration -->
30 min
</div>
<div style="height: 10px"><p style="font-size: 12px; text-align: right">
<!-- URL -->
<a href="http://localhost:8888/notebooks/Db2_Data_Management_Console_Overview.ipynb">
<img style="display: inline-block;"src="./media/arrowblue.png"></a>
</div>
</td>
<td style="padding: 15px; text-align:left; vertical-align: text-top; background-color:#F7F7F7; width: 300px; height: 250px;">
<div style="height: 75px"><p style="font-size: 24px">
<!-- Title -->
Analyzing SQL Workloads
</div>
<div style="height: 125px"><p style="font-size: 14px">
<!-- Description -->
This lab uses more adanced techniques to analyze SQL workloads and individual statements. Run workloads in batch across databases to compare performance across Db2 services. Learn how to embed the interactive Visual Explain tool into your Jupyter notebooks.
</div>
<div style="height: 25px"><p style="font-size: 12px; text-align: right">
<img style="display: inline-block;"src="./media/clock.png">
<!-- Duration -->
30 min
</div>
<div style="height: 10px"><p style="font-size: 12px; text-align: right">
<!-- URL -->
<a href="http://localhost:8888/notebooks/Db2_Data_Management_Console_SQL.ipynb">
<img style="display: inline-block;"src="./media/arrowblue.png"></a>
</div>
</td>
</table>
</div>
If you are administering the Db2 Console for your team, the next lesson walks you through some of the most important tasks to setup and maintain the DB2 Console.
<!-- Row 1 -->
<div style="font-family: 'IBM Plex Sans';">
<table style="float:left; width: 310px; height: 235px; border-spacing: 10px; border-collapse: separate; table-layout: fixed">
<td style="padding: 15px; text-align:left; vertical-align: text-top; background-color:#F7F7F7; width: 300px; height: 250px;">
<div style="height: 75px"><p style="font-size: 24px">
<!-- Title -->
Managing the Console Settings
</div>
<div style="height: 125px"><p style="font-size: 14px">
<!-- Description -->
This Jupyter Notebook contains examples of how to setup and manage the Db2 Data Management Console. It covers how to add additional users using database authentication, how to explore and manage connections and setup and manage monitoring profiles.
</div>
<div style="height: 25px"><p style="font-size: 12px; text-align: right">
<img style="display: inline-block;"src="./media/clock.png">
<!-- Duration -->
30 min
</div>
<div style="height: 10px"><p style="font-size: 12px; text-align: right">
<!-- URL -->
<a href="http://localhost:8888/notebooks/Db2_Data_Management_Console_Management.ipynb">
<img style="display: inline-block;"src="./media/arrowblue.png"></a>
</div>
</td>
<td style="padding: 15px; text-align:left; vertical-align: text-top; background-color:#F7F7F7; width: 300px; height: 250px;">
<div style="height: 75px"><p style="font-size: 24px">
<!-- Title -->
Applying a Fix Pack to the Db2 Console
</div>
<div style="height: 125px"><p style="font-size: 14px">
<!-- Description -->
Fix Pack updates are regularly available for the Db2 Data Management Console. This notebook shows you how to upgrade from Version 3.1 to 3.1.1.
</div>
<div style="height: 25px"><p style="font-size: 12px; text-align: right">
<img style="display: inline-block;"src="./media/clock.png">
<!-- Duration -->
15 min
</div>
<div style="height: 10px"><p style="font-size: 12px; text-align: right">
<!-- URL -->
<a href="http://localhost:8888/notebooks/Db2_Data_Management_Console_FPUpgrade.ipynb">
<img style="display: inline-block;"src="./media/arrowblue.png"></a>
</div>
</td>
<td style="padding: 15px; text-align:left; vertical-align: text-top; background-color:#F7F7F7; width: 300px; height: 250px;">
<div style="height: 75px"><p style="font-size: 24px">
<!-- Title -->
Using cURL to work with the Db2 Console
</div>
<div style="height: 125px"><p style="font-size: 14px">
<!-- Description -->
This Jupyter Notebook contains examples of how to setup and manage the Db2 Data Management Console using cURL in simple BASH scripts. cURL is a command-line tool for getting or sending data, including RESTful API calls, using URL syntax
</div>
<div style="height: 25px"><p style="font-size: 12px; text-align: right">
<img style="display: inline-block;"src="./media/clock.png">
<!-- Duration -->
10 min
</div>
<div style="height: 10px"><p style="font-size: 12px; text-align: right">
<!-- URL -->
<a href="http://localhost:8888/notebooks/Using%20CURL.ipynb#">
<img style="display: inline-block;"src="./media/arrowblue.png"></a>
</div>
</td>
</table>
</div>
You can even create customized versions of the Db2 Console with a bit of HTML and CSS.
<!-- Row 1 -->
<div style="font-family: 'IBM Plex Sans';">
<table style="float:left; width: 310px; height: 235px; border-spacing: 10px; border-collapse: separate; table-layout: fixed">
<td style="padding: 15px; text-align:left; vertical-align: text-top; background-color:#F7F7F7; width: 300px; height: 250px;">
<div style="height: 75px"><p style="font-size: 24px">
<!-- Title -->
Building a Custom Db2 Console Webpage
</div>
<div style="height: 125px"><p style="font-size: 14px">
<!-- Description -->
This Jupyter Notebook contains examples of how to build a custom web page out of parts of the Db2 Console microservice user interface. In just a few minutes you will customize your own web page and install Apache to share it.
</div>
<div style="height: 25px"><p style="font-size: 12px; text-align: right">
<img style="display: inline-block;"src="./media/clock.png">
<!-- Duration -->
15 min
</div>
<div style="height: 10px"><p style="font-size: 12px; text-align: right">
<!-- URL -->
<a href="http://localhost:8888/notebooks/ComposeConsoleWebpage.ipynb">
<img style="display: inline-block;"src="./media/arrowblue.png"></a>
</div>
</td>
</table>
</div>
#### Questions and Comments: Peter Kohlmann [kohlmann@ca.ibm.com], George Baklarz [baklarz@ca.ibm.com]
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
data = pd.read_csv('/Users/zengxin/Study/Econ2355/result/corr.csv', index_col="Unnamed: 0")
data
data_corr = data.drop(['ticker', 'date'], axis=1)
corr = data[[i for i in data_corr.columns]].corr()
ax = sns.heatmap(
corr,
vmin=-1, vmax=1, center=0,
cmap=sns.diverging_palette(20, 220, n=200),
square=True
)
ax.set_xticklabels(
ax.get_xticklabels(),
rotation=45,
horizontalalignment='right'
);
corr.loc["sentiment_score"]
plt.figure(figsize=(10, 7))
data = data.sort_values(by="date")
plt.plot(range(data.shape[0]), data["sentiment_score"].values, label="Sentimental Score")
plt.plot(range(data.shape[0]), data["return_1"].values, label="1-day return")
plt.ylabel('Value')
plt.xlabel('Data Points')
plt.title("Data Visulization")
plt.legend()
plt.figure(figsize=(10, 7))
data = data.sort_values(by="date")
plt.plot(range(data.shape[0]), data["sentiment_score"].values, label="Sentimental Score")
plt.plot(range(data.shape[0]), data["return_10"].values, label="10-day return")
plt.ylabel('y')
plt.xlabel('x')
plt.title("Data Visulization")
plt.legend()
plt.figure(figsize=(10, 7))
plt.plot(range(data.shape[0]), data["sentiment_score"].values, label="Sentimental Score")
plt.plot(range(data.shape[0]), data["return_100"].values, label="100-day return")
plt.xlabel('date')
plt.title("Data Visulization")
plt.legend()
plt.figure(figsize=(10, 7))
# plt.scatter(x = range(data.shape[0]), y = data["sentiment_score"].values, label="Sentimental Score")
# plt.scatter(x = range(data.shape[0]), y = data["return_1"].values, label="1-day return")
plt.plot(range(data.shape[0]), data["sentiment_score"].values, label="Sentimental Score")
plt.plot(range(data.shape[0]), data["return_60"].values, label="60-day return")
plt.xlabel('date')
plt.title("Data Visulization")
plt.legend()
plt.figure(figsize=(10, 7))
# plt.scatter(x = range(data.shape[0]), y = data["sentiment_score"].values, label="Sentimental Score")
# plt.scatter(x = range(data.shape[0]), y = data["return_1"].values, label="1-day return")
index = np.argwhere(~np.isnan(data["return_30"].values))
plt.plot(range(data["sentiment_score"].values[index].shape[0]), data["sentiment_score"].values[index], label="Sentimental Score")
plt.plot(range(data["sentiment_score"].values[index].shape[0]), data["return_30"].values[index], label="30-day return")
plt.xlabel('date')
plt.title("Sentimental Scores and 30-day Return")
plt.legend()
plt.figure(figsize=(10, 7))
# plt.scatter(x = range(data.shape[0]), y = data["sentiment_score"].values, label="Sentimental Score")
# plt.scatter(x = range(data.shape[0]), y = data["return_1"].values, label="1-day return")
index = np.argwhere(~np.isnan(data["return_1"].values))
plt.plot(range(data["sentiment_score"].values[index].shape[0]), data["sentiment_score"].values[index], label="Sentimental Score")
plt.plot(range(data["sentiment_score"].values[index].shape[0]), data["return_5"].values[index], label="5-day return")
plt.xlabel('date')
plt.title("Sentimental Scores and 5-day Return")
plt.legend()
plt.scatter(data["sentiment_score"].values, data["return_20"].values, c=data["date"]//10000)
plt.ylabel('20 day-Return')
plt.xlabel('Sentimental Score')
plt.colorbar()
data = pd.read_csv('/Users/zengxin/Study/Econ2355/result/corr_final.csv', index_col="Unnamed: 0")
data_corr = data.drop(['ticker', 'date'], axis=1)
corr = data[[i for i in data_corr.columns]].corr()
ax = sns.heatmap(
corr,
vmin=-1, vmax=1, center=0,
cmap=sns.diverging_palette(20, 220, n=200),
square=True
)
ax.set_xticklabels(
ax.get_xticklabels(),
rotation=45,
horizontalalignment='right'
);
data_after_2018 = data[data.date >= 20190101]
data_corr = data_after_2018.drop(['ticker', 'date'], axis=1)
corr_2018 = data_after_2018[[i for i in data_corr.columns]].corr()
ax = sns.heatmap(
corr,
vmin=-1, vmax=1, center=0,
cmap=sns.diverging_palette(20, 220, n=200),
square=True
)
ax.set_xticklabels(
ax.get_xticklabels(),
rotation=45,
horizontalalignment='right'
);
corr_2018.loc["sentiment_score"][:9]
plt.plot(corr_2018.loc["sentiment_score"].values[2:9])
plt.xticks(np.arange(7), corr_2018.index[2:9])
plt.ylabel('Correlation')
plt.scatter(data_after_2018["sentiment_score"].values, data_after_2018["return_20"].values, c=data_after_2018["date"]//10000)
plt.ylabel('20 day-Return')
plt.xlabel('Sentimental Score')
plt.colorbar()
plt.scatter(data_after_2018["sentiment_score"].values, data_after_2018["return_20"].values, c=data_after_2018["ticker"].astype(int))
plt.ylabel('20 day-Return')
plt.xlabel('Sentimental Score')
plt.colorbar()
plt.scatter(data_after_2018["sentiment_score"].values, data_after_2018["return_30"].values, c=data_after_2018["date"]//10000)
plt.ylabel('30 day-Return')
plt.xlabel('Sentimental Score')
plt.colorbar()
plt.scatter(data_after_2018["sentiment_score"].values, data_after_2018["return_30"].values, c=data_after_2018["ticker"].astype(int))
plt.ylabel('30 day-Return')
plt.xlabel('Sentimental Score')
plt.colorbar()
```
| github_jupyter |
# Delay-and-Sum Beamformer - Linear Array of Infinite Length
*This Jupyter notebook is part of a [collection of notebooks](../index.ipynb) in the masters course Selected Topics in Audio Signal Processing, Communications Engineering, Universität Rostock. Please direct questions and suggestions to [Sascha.Spors@uni-rostock.de](mailto:Sascha.Spors@uni-rostock.de).*
## Beampattern
In this example the beampattern of a delay-and-sum (DSB) beamformer for a linear array of infinite length is computed and plotted for various steering angles. For numerical evaluation the array of infinite length is approximated by a long array of finite length. First, two functions are defined for computation and illustration of the beampattern, respectively.
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
dx = 0.1 # spatial sampling interval (distance between microphones)
c = 343 # speed of sound
om = 2*np.pi * np.linspace(100, 8000, 1000) # angular frequencies
theta_pw = np.linspace(0, np.pi, 181) # angles of the incident plane waves
def compute_dsb_beampattern(theta, theta_pw, om, dx, nmic=5000):
"Compute beampattern of a delay-and-sub beamformer for given steering angle"
B = np.zeros(shape=(len(om), len(theta_pw)), dtype=complex)
for n in range(len(om)):
for mu in range(-nmic//2, nmic//2+1):
B[n, :] += np.exp(-1j * om[n]/c * mu*dx * (np.cos(theta_pw) - np.cos(theta)))
return B/nmic
def plot_dsb_beampattern(B, theta_pw, om):
"Plot beampattern of a delay-and-sub beamformer"
plt.figure(figsize=(10,10))
plt.imshow(20*np.log10(np.abs(B)), aspect='auto', vmin=-50, vmax=0, origin='lower', \
extent=[0, 180, om[0]/(2*np.pi), om[-1]/(2*np.pi)], cmap='viridis')
plt.xlabel(r'$\theta_{pw}$ in deg')
plt.ylabel('$f$ in Hz')
plt.title('Beampattern')
cb = plt.colorbar()
cb.set_label(r'$|\bar{P}(\theta, \theta_{pw}, \omega)|$ in dB')
```
### Steering Angle $\theta = 90^\mathrm{o}$
```
B = compute_dsb_beampattern(np.pi/2, theta_pw, om, dx)
plot_dsb_beampattern(B, theta_pw, om)
```
### Steering Angle $\theta = 45^\mathrm{o}$
```
B = compute_dsb_beampattern(np.pi/4, theta_pw, om, dx)
plot_dsb_beampattern(B, theta_pw, om)
```
### Steering Angle $\theta = 0^\mathrm{o}$
```
B = compute_dsb_beampattern(0, theta_pw, om, dx)
plot_dsb_beampattern(B, theta_pw, om)
```
**Copyright**
This notebook is provided as [Open Educational Resources](https://en.wikipedia.org/wiki/Open_educational_resources). Feel free to use the notebook for your own purposes. The text/images/data are licensed under [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/), the code of the IPython examples under the [MIT license](https://opensource.org/licenses/MIT). Please attribute the work as follows: *Sascha Spors, Selected Topics in Audio Signal Processing - Supplementary Material, 2017*.
| github_jupyter |
<a href="https://qworld.net" target="_blank" align="left"><img src="../qworld/images/header.jpg" align="left"></a>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
$ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
$ \newcommand{\greenbit}[1] {\mathbf{{\color{green}#1}}} $
$ \newcommand{\bluebit}[1] {\mathbf{{\color{blue}#1}}} $
$ \newcommand{\redbit}[1] {\mathbf{{\color{red}#1}}} $
$ \newcommand{\brownbit}[1] {\mathbf{{\color{brown}#1}}} $
$ \newcommand{\blackbit}[1] {\mathbf{{\color{black}#1}}} $
<font style="font-size:28px;" align="left"><b> Entanglement and Superdense Coding </b></font>
<br>
_prepared by Abuzer Yakaryilmaz_
<br><br>
[<img src="../qworld/images/watch_lecture.jpg" align="left">](https://youtu.be/ZzRcItzUF2U)
<br><br><br>
Asja has a qubit, initially set to $ \ket{0} $.
Balvis has a qubit, initially set to $ \ket{0} $.
<h3> Entanglement </h3>
Asja applies Hadamard operator to her qubit.
The quantum state of Asja's qubit is $ \stateplus $.
Then, Asja and Balvis combine their qubits. Their quantum state is
$ \stateplus \otimes \vzero = \myvector{ \frac{1}{\sqrt{2}} \\ 0 \\ \frac{1}{\sqrt{2}} \\ 0 } $.
Asja and Balvis apply CNOT operator on two qubits.
The new quantum state is
$ \CNOT \myvector{ \frac{1}{\sqrt{2}} \\ 0 \\ \frac{1}{\sqrt{2}} \\ 0 } = \myvector{ \frac{1}{\sqrt{2}} \\ 0 \\0 \\ \frac{1}{\sqrt{2}} } = \frac{1}{\sqrt{2}}\ket{00} + \frac{1}{\sqrt{2}}\ket{11} $.
At this moment, Asja's and Balvis' qubits are correlated to each other.
If we measure both qubits, we can observe either state $ \ket{00} $ or state $ \ket{11} $.
Suppose that Asja observes her qubit secretly.
<ul>
<li> When Asja sees the result $ \ket{0} $, then Balvis' qubit also collapses to state $ \ket{0} $. Balvis cannot observe state $ \ket{1} $. </li>
<li> When Asja sees the result $ \ket{1} $, then Balvis' qubit also collapses to state $ \ket{1} $. Balvis cannot observe state $ \ket{0} $. </li>
</ul>
Experimental results have confirmed that this happens even if there is a physical distance between Asja's and Balvis' qubits.
It seems correlated quantum particles can "affect each other" instantly, even if they are in the different part of the universe.
If two qubits are correlated in this way, then we say that they are <b>entangled</b>.
<i> <u>Technical note</u>:
If the quantum state of two qubits can be written as $ \ket{u} \otimes \ket{v} $, then two qubits are not correlated, where $ \ket{u} $ and $ \ket{v} $ are the quantum states of the first and second qubits.
On the other hand, if the quantum state of two qubits cannot be written as $ \ket{u} \otimes \ket{v} $, then there is an entanglement between the qubits.
</i>
<b> Entangled qubits can be useful </b>
<h3> Quantum communication </h3>
After having the entanglement, Balvis takes his qubit and goes away.
Asja will send two classical bits of information by only sending her qubit.
<img src="images/superdense-coding.jpg" align="left" width="800px">
Now, we describe this protocol.
Asja has two bits of classical information: $ a,b \in \{0,1\} $.
There are four possible values for the pair $ (a,b) $: $ (0,0), (0,1), (1,0),\mbox{ or } (1,1) $.
If $a$ is 1, then Asja applies z-gate, i.e., $ Z = \Z $, to her qubit.
If $b$ is 1, then Asja applies x-gate (NOT operator) to her qubit.
Then, Asja sends her qubit to Balvis.
<h3> After the communication </h3>
Balvis has both qubits.
Balvis applies cx-gate (CNOT operator), where Asja's qubit is the controller.
Then, Balvis applies h-gate (Hadamard operator) to Asja's qubit.
Balvis measures both qubits.
The measurement result will be exactly $ (a,b) $.
<h3> Task 1</h3>
Verify the correctness of the above protocol.
For each pair of $ (a,b) \in \left\{ (0,0), (0,1), (1,0),(1,1) \right\} $:
- Create a quantum circuit with two qubits: Asja's and Balvis' qubits
- Both are initially set to $ \ket{0} $
- Apply h-gate (Hadamard) to Asja's qubit
- Apply cx-gate as CNOT(Asja's-qubit,Balvis'-qubit)
Assume that both qubits are separated from each other.
<ul>
<li> If $ a $ is 1, then apply z-gate to Asja's qubit. </li>
<li> If $ b $ is 1, then apply x-gate (NOT) to Asja's qubit. </li>
</ul>
Assume that Asja sends her qubit to Balvis.
- Apply cx-gate as CNOT(Asja's-qubit,Balvis'-qubit)
- Apply h-gate (Hadamard) to Asja's qubit
- Measure both qubits and compare the results with pair $ (a,b) $
```
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
all_pairs = ['00','01','10','11']
#
# your code is here
#
```
<a href="Q72_Superdense_Coding_Solutions.ipynb#task1">click for our solution</a>
<h3> Task 2 </h3>
Verify each case by tracing the state vector (on paper).
_Hint: Representing quantum states as the linear combinations of basis states makes calculation easier._
<h3> Task 3</h3>
Can the above set-up be used by Balvis?
Verify that the following modified protocol allows Balvis to send two classical bits by sending only his qubit.
For each pair of $ (a,b) \in \left\{ (0,0), (0,1), (1,0),(1,1) \right\} $:
- Create a quantum circuit with two qubits: Asja's and Balvis' qubits
- Both are initially set to $ \ket{0} $
- Apply h-gate (Hadamard) to Asja's qubit
- Apply cx-gate as CNOT(Asja's-qubit,Balvis'-qubit)
Assume that both qubits are separated from each other.
<ul>
<li> If $ a $ is 1, then apply z-gate to Balvis' qubit. </li>
<li> If $ b $ is 1, then apply x-gate (NOT) to Balvis' qubit. </li>
</ul>
Assume that Balvis sends his qubit to Asja.
- Apply cx-gate as CNOT(Asja's-qubit,Balvis'-qubit)
- Apply h-gate (Hadamard) to Asja's qubit
- Measure both qubits and compare the results with pair $ (a,b) $
```
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
all_pairs = ['00','01','10','11']
#
# your code is here
#
```
<a href="Q72_Superdense_Coding_Solutions.ipynb#task3">click for our solution</a>
<h3> Task 4 </h3>
Verify each case by tracing the state vector (on paper).
_Hint: Representing quantum states as the linear combinations of basis states makes calculation easier._
| github_jupyter |
# Named Entity Recognition by fine-tuning Keras BERT on SageMaker
## Setup
We'll begin with some necessary imports, and get an Amazon SageMaker session to help perform certain tasks, as well as an IAM role with the necessary permissions.
```
import os
import json
import time
from datetime import datetime
import numpy as np
import pandas as pd
import boto3
import sagemaker
from sagemaker import get_execution_role
from sagemaker.tensorflow import TensorFlow
from sagemaker.tensorflow.serving import TensorFlowModel
import logging
role = get_execution_role()
%matplotlib inline
```
### SageMaker variables and S3 bucket
```
#Creating a sagemaker session
sagemaker_session = sagemaker.Session()
#We'll be using the sagemaker default bucket
BUCKET = sagemaker_session.default_bucket()
PREFIX = 'graph-nerc-blog' #Feel free to change this
DATA_FOLDER = 'tagged-data'
#Using default region, same as where this notebook is, change if needed
REGION = sagemaker_session.boto_region_name
INPUTS = 's3://{}/{}/{}/'.format(BUCKET,PREFIX,DATA_FOLDER)
print("Using region: {}".format(REGION))
print('Bucket: {}'.format(BUCKET))
print("Using prefix : {}".format(INPUTS))
```
# Downloading dataset
We will be using the Kaggle entity-annotated-corpus that can be found at https://www.kaggle.com/abhinavwalia95/entity-annotated-corpus
To be able to download it, you will be required to create a Kaggle account.
Once the zip folder is downloaded, unzip it locally and upload the file ner_dataset.csv in the folder of this notebook. (notebooks/)
# 1. Data exploration and preparation
The dataset consists of 47959 news article sentences (1048575 words) with tagged entities representing:
- geo = Geographical Entity
- org = Organization
- per = Person
- gpe = Geopolitical Entity
- tim = Time indicator
- art = Artifact
- eve = Event
- nat = Natural Phenomenon
```
ner_dataset = pd.read_csv('ner_dataset.csv', encoding = 'latin')
# Here is an example sentence. We will only be using the Sentence #, Word and Tag columns
ner_dataset.head(24)
# These are the following entities we have in the data
ner_dataset.Tag.unique()
ner_dataset.Tag = ner_dataset.Tag.fillna('O')
```
### Split data to train and test
We split the data into train, validation and test set, taking the first 45000 sentences for training, the next 2000 sentences for validation and the last 959 sentences for testing.
```
index = ner_dataset['Sentence #'].index[~ner_dataset['Sentence #'].isna()].values.tolist()
train_index = index[45000]
val_index = index[47000]
train_df = ner_dataset[:train_index]
val_df = ner_dataset[train_index:val_index]
test_df = ner_dataset[val_index:]
```
### Save data to s3
```
train_df.to_csv(INPUTS + 'train.csv')
val_df.to_csv(INPUTS + 'val.csv')
test_df.to_csv(INPUTS + 'test.csv')
```
# 2. Training BERT model using Sagemaker
For fine-tuning the Keras BERT for Named Entity Recognition, the whole code is in the folder code/
The folder contains the train.py script that will be executed within a SageMaker training job to launch the training. The train.py imports modules for found in code/source/
```
!pygmentize ../code/train.py
```
The data on which we train are the outputs of part 1: Data Exploration and Preparation
**NOTA: If you change where you save the train, validation and test csv files please reflect those changes in the INPUTS variable**
## Single Training job
### Job name and instance type
```
JOB_NAME = 'ner-bert-keras'
INSTANCE_TYPE = 'ml.p3.2xlarge'
# INSTANCE_TYPE = "local_gpu"
```
### Hyperparameters:
```
EPOCHS = 20
BATCH_SIZE = 16
MAX_SEQUENCE_LENGTH = 64 # This correspond to the input size of BERT that we want (The training time is quadratically increasing with input size)
DROP_OUT = 0.1
LEARNING_RATE = 4.0e-05
BERT_PATH = 'https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/3'
OUTPUT_PATH = 's3://{}/{}/training-output/'.format(BUCKET,PREFIX)
```
### Defining training job
By providing a *framework_version* "2.3" and *py_version* "py37" in the TensorFlow object, we'll be calling a managed ECR image provided by AWS.
Training on a gpu instance in the region eu-west-1, this is the same as providing the explicit image_uri: *training_image_uri = "763104351884.dkr.ecr.eu-west-1.amazonaws.com/tensorflow-training:2.3.1-gpu-py37-cu110-ubuntu18.04".*
Using *framework_version* and *py_version* the TensorFlow (Estimator) object manages this for you by ensuring that the right image_uri is used depending on the region of your sagemaker session, as well as the type of instance used for training.
Refer to https://github.com/aws/deep-learning-containers/blob/master/available_images.md for more details
```
hyperparameters = {'epochs': EPOCHS,
'batch_size' : BATCH_SIZE,
'max_sequence_length': MAX_SEQUENCE_LENGTH,
'drop_out': DROP_OUT,
'learning_rate': LEARNING_RATE,
'bert_path':BERT_PATH
}
# Use either framework_version and py_version or an explicit image_uri
estimator = TensorFlow(base_job_name=JOB_NAME,
source_dir='../code',
entry_point='train.py',
role=role,
framework_version='2.3',
py_version='py37',
# image_uri=training_image_uri,
hyperparameters=hyperparameters,
instance_count=1,
script_mode=True,
metric_definitions=[
{'Name': 'train loss', 'Regex': 'loss: (.*?) -'},
{'Name': 'train accuracy', 'Regex': ' accuracy: (.*?) -'},
{'Name': 'val loss', 'Regex': 'val_loss: (.*?) -'},
{'Name': 'val accuracy', 'Regex': 'val_accuracy: (.*?)$'}
],
output_path=OUTPUT_PATH,
instance_type=INSTANCE_TYPE)
REMOTE_INPUTS = {'train' : INPUTS,
'validation' : INPUTS,
'eval' : INPUTS}
dt = datetime.now()
estimator.fit(REMOTE_INPUTS, wait = False) # Set to True if you want to see the logs here
```
The training can take between 40min and 1h. The following cells can be run to check the status. Once the status is 'Completed' you can go ahead and Deploy an Inference Endpoint
```
print(estimator.model_data)
sm_client = boto3.client('sagemaker')
response = sm_client.describe_training_job(
TrainingJobName=estimator._current_job_name
)
response.get('TrainingJobStatus')
```
### Run the next cells only once TrainingJobStatus response is 'Completed'
# 3. Deploy an Inference Endpoint
```
!pygmentize ../code/inference.py
MODEL_ARTEFACTS_S3_LOCATION = response.get('ModelArtifacts').get('S3ModelArtifacts')
INSTANCE_TYPE = "ml.g4dn.xlarge"
print(MODEL_ARTEFACTS_S3_LOCATION)
# Use either framework_version or an explicit image_uri. Refer to https://github.com/aws/deep-learning-containers/blob/master/available_images.md for more details
model = TensorFlowModel(entry_point='inference.py',
source_dir='../code',
framework_version='2.3',
# image_uri = inference_image_uri,
role=role,
model_data=MODEL_ARTEFACTS_S3_LOCATION,
sagemaker_session=sagemaker_session,
env = {'SAGEMAKER_MODEL_SERVER_TIMEOUT' : '300' }
)
predictor = model.deploy(initial_instance_count=1, instance_type=INSTANCE_TYPE, wait=True)
```
### Testing the endpoint
```
test_set = pd.read_csv(INPUTS + 'test.csv')
df = test_set.copy()
df = df.fillna(method='ffill')
d = (df.groupby('Sentence #')
.apply(lambda x: list(x['Word']))
.to_dict())
test_list = []
for (k, v) in d.items():
article = {'id': k, 'sentence':' '.join(v)}
test_list.append(article)
test_list[:10]
start_time = time.time()
test_endpoint = predictor.predict(test_list[:1000])
print("--- %s seconds ---" % (time.time() - start_time))
test_endpoint[-10:]
```
### Running predictions for the whole dataset (example)
Endpoints, which are built for doing real-time inference, and are by design meant to run for a maximum of 60 seconds per request.
To test the endpoint on a big dataset, we can send it in requests of 1000 sentences each to avoid long inferences that could make the endpoint time out.
```
# train_set = pd.read_csv(INPUTS + 'train.csv')
# val_set = pd.read_csv(INPUTS + 'val.csv')
# df = pd.concat([train_set,val_set,test_set])
# df = df.fillna(method='ffill')
# d = (df.groupby('Sentence #')
# .apply(lambda x: list(x['Word']))
# .to_dict())
# test_list = []
# for (k, v) in d.items():
# article = {'id': k, 'sentence':' '.join(v)}
# test_list.append(article)
# start_time = time.time()
# preds = []
# for k in range (0,round(len(test_list)/1000)):
# preds.append(predictor.predict(test_list[k*1000:(k+1)*1000]))
# print("--- %s seconds ---" % (time.time() - start_time))
# preds_flat = [item for sublist in preds for item in sublist]
# preds_flat[-10:]
# with open('data_with_entities.json', 'w', encoding='utf-8') as f:
# json.dump(preds_flat, f, ensure_ascii=False, indent=4)
```
### Writing output to s3
```
import json
import boto3
s3 = boto3.resource('s3')
s3object = s3.Object(BUCKET, PREFIX + '/data_with_entities.json')
s3object.put(
Body=(bytes(json.dumps(test_endpoint).encode('UTF-8')))
)
```
### Delete the endpoint
```
# predictor.delete_endpoint()
```
| github_jupyter |
<center> <font size=5> <h1>Define working environment</h1> </font> </center>
The following cells are used to:
- Import needed libraries
- Set the environment variables for Python, Anaconda, GRASS GIS and R statistical computing
- Define the ["GRASSDATA" folder](https://grass.osgeo.org/grass73/manuals/helptext.html), the name of "location" and "mapset" where you will to work.
**Import libraries**
```
## Import libraries needed for setting parameters of operating system
import os
import sys
```
<center> <font size=3> <h3>Environment variables when working on Linux Mint</h3> </font> </center>
**Set 'Python' and 'GRASS GIS' environment variables**
Here, we set [the environment variables allowing to use of GRASS GIS](https://grass.osgeo.org/grass64/manuals/variables.html) inside this Jupyter notebook. Please change the directory path according to your own system configuration.
```
### Define GRASS GIS environment variables for LINUX UBUNTU Mint 18.1 (Serena)
# Check is environmental variables exists and create them (empty) if not exists.
if not 'PYTHONPATH' in os.environ:
os.environ['PYTHONPATH']=''
if not 'LD_LIBRARY_PATH' in os.environ:
os.environ['LD_LIBRARY_PATH']=''
# Set environmental variables
os.environ['GISBASE'] = '/home/tais/SRC/GRASS/grass_trunk/dist.x86_64-pc-linux-gnu'
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'bin')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'script')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'lib')
#os.environ['PATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'etc','python')
os.environ['PYTHONPATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'etc','python')
os.environ['PYTHONPATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'etc','python','grass')
os.environ['PYTHONPATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'etc','python','grass','script')
os.environ['PYTHONLIB'] = '/usr/lib/python2.7'
os.environ['LD_LIBRARY_PATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'lib')
os.environ['GIS_LOCK'] = '$$'
os.environ['GISRC'] = os.path.join(os.environ['HOME'],'.grass7','rc')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['HOME'],'.grass7','addons')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['HOME'],'.grass7','addons','bin')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['HOME'],'.grass7','addons')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['HOME'],'.grass7','addons','scripts')
## Define GRASS-Python environment
sys.path.append(os.path.join(os.environ['GISBASE'],'etc','python'))
```
**Import GRASS Python packages**
```
## Import libraries needed to launch GRASS GIS in the jupyter notebook
import grass.script.setup as gsetup
## Import libraries needed to call GRASS using Python
import grass.script as gscript
from grass.script import core as grass
```
**-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
**Display current environment variables of your computer**
```
## Display the current defined environment variables
for key in os.environ.keys():
print "%s = %s \t" % (key,os.environ[key])
```
**-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
<center> <font size=5> <h1>Define functions</h1> </font> </center>
This section of the notebook is dedicated to defining functions which will then be called later in the script. If you want to create your own functions, define them here.
### Function for computing processing time
The "print_processing_time" is used to calculate and display the processing time for various stages of the processing chain. At the beginning of each major step, the current time is stored in a new variable, using [time.time() function](https://docs.python.org/2/library/time.html). At the end of the stage in question, the "print_processing_time" function is called and takes as argument the name of this new variable containing the recorded time at the beginning of the stage, and an output message.
```
## Import library for managing time in python
import time
## Function "print_processing_time()" compute processing time and printing it.
# The argument "begintime" wait for a variable containing the begintime (result of time.time()) of the process for which to compute processing time.
# The argument "printmessage" wait for a string format with information about the process.
def print_processing_time(begintime, printmessage):
endtime=time.time()
processtime=endtime-begintime
remainingtime=processtime
days=int((remainingtime)/86400)
remainingtime-=(days*86400)
hours=int((remainingtime)/3600)
remainingtime-=(hours*3600)
minutes=int((remainingtime)/60)
remainingtime-=(minutes*60)
seconds=round((remainingtime)%60,1)
if processtime<60:
finalprintmessage=str(printmessage)+str(seconds)+" seconds"
elif processtime<3600:
finalprintmessage=str(printmessage)+str(minutes)+" minutes and "+str(seconds)+" seconds"
elif processtime<86400:
finalprintmessage=str(printmessage)+str(hours)+" hours and "+str(minutes)+" minutes and "+str(seconds)+" seconds"
elif processtime>=86400:
finalprintmessage=str(printmessage)+str(days)+" days, "+str(hours)+" hours and "+str(minutes)+" minutes and "+str(seconds)+" seconds"
return finalprintmessage
```
### Function for creation of configuration file for r.li (landscape units provided as polygons) (multiprocessed)
```
##### Function that create the r.li configuration file for a list of landcover raster.
### It enable to create in one function as many configuration file as the number of raster provided in 'listoflandcoverraster'.
### It could be use only in case study with a several landcover raster and only one landscape unit layer.
### So, the landscape unit layer if fixed and there are the landcover raster which change.
# 'listoflandcoverraster' wait for a list with the name (string) of landcover rasters.
# 'landscape_polygons' wait for the name (string) of the vector layer containing the polygons to be used as landscape units.
# 'returnlistpath' wait for a boolean value (True/False) according to the fact that a list containing the path to the configuration files is desired.
# 'ncores' wait for a integer corresponding to the number of desired cores to be used for parallelization.
# Import libraries for multiprocessing
import multiprocessing
from multiprocessing import Pool
from functools import partial
# Function that copy the landscape unit raster masks on a new layer with name corresponding to the current 'landcover_raster'
def copy_landscapeunitmasks(current_landcover_raster,base_landcover_raster,landscape_polygons,landscapeunit_bbox,cat):
### Copy the landscape units mask for the current 'cat'
# Define the name of the current "current_landscapeunit_rast" layer
current_landscapeunit_rast=current_landcover_raster.split("@")[0]+"_"+landscape_polygons.split("@")[0]+"_"+str(cat)
base_landscapeunit_rast=base_landcover_raster.split("@")[0]+"_"+landscape_polygons.split("@")[0]+"_"+str(cat)
# Copy the the landscape unit created for the first landcover map in order to match the name of the current landcover map
gscript.run_command('g.copy', overwrite=True, quiet=True, raster=(base_landscapeunit_rast,current_landscapeunit_rast))
# Add the line to the maskedoverlayarea variable
maskedoverlayarea="MASKEDOVERLAYAREA "+current_landscapeunit_rast+"|"+landscapeunit_bbox[cat]
return maskedoverlayarea
# Function that create the r.li configuration file for the base landcover raster and then for all the binary rasters
def create_rli_configfile(listoflandcoverraster,landscape_polygons,returnlistpath=True,ncores=2):
# Check if 'listoflandcoverraster' is not empty
if len(listoflandcoverraster)==0:
sys.exit("The list of landcover raster is empty and should contain at least one raster name")
# Check if raster exists to avoid error in mutliprocessing
for cur_rast in listoflandcoverraster:
try:
mpset=cur_rast.split("@")[1]
except:
mpset=""
if cur_rast not in gscript.list_strings(type='raster',mapset=mpset):
sys.exit('Raster <%s> not found' %categorical_raster)
# Get the version of GRASS GIS
version=grass.version()['version'].split('.')[0]
# Define the folder to save the r.li configuration files
if sys.platform=="win32":
rli_dir=os.path.join(os.environ['APPDATA'],"GRASS"+version,"r.li")
else:
rli_dir=os.path.join(os.environ['HOME'],".grass"+version,"r.li")
if not os.path.exists(rli_dir):
os.makedirs(rli_dir)
## Create an ordered list with the 'cat' value of landscape units to be processed.
list_cat=[int(x) for x in gscript.parse_command('v.db.select', quiet=True, map=landscape_polygons, column='cat', flags='c')]
list_cat.sort()
# Declare a empty dictionnary which will contains the north, south, east, west values for each landscape unit
landscapeunit_bbox={}
# Declare a empty list which will contain the path of the configation files created
listpath=[]
# Declare a empty string variable which will contains the core part of the r.li configuration file
maskedoverlayarea=""
# Duplicate 'listoflandcoverraster' in a new variable called 'tmp_list'
tmp_list=list(listoflandcoverraster)
# Set the current landcover raster as the first of the list
base_landcover_raster=tmp_list.pop(0) #The pop function return the first item of the list and delete it from the list at the same time
# Loop trough the landscape units
for cat in list_cat:
# Extract the current landscape unit polygon as temporary vector
tmp_vect="tmp_"+base_landcover_raster.split("@")[0]+"_"+landscape_polygons.split("@")[0]+"_"+str(cat)
gscript.run_command('v.extract', overwrite=True, quiet=True, input=landscape_polygons, cats=cat, output=tmp_vect)
# Set region to match the extent of the current landscape polygon, with resolution and alignement matching the landcover raster
gscript.run_command('g.region', vector=tmp_vect, align=base_landcover_raster)
# Rasterize the landscape unit polygon
landscapeunit_rast=tmp_vect[4:]
gscript.run_command('v.to.rast', overwrite=True, quiet=True, input=tmp_vect, output=landscapeunit_rast, use='cat', memory='3000')
# Remove temporary vector
gscript.run_command('g.remove', quiet=True, flags="f", type='vector', name=tmp_vect)
# Set the region to match the raster landscape unit extent and save the region info in a dictionary
region_info=gscript.parse_command('g.region', raster=landscapeunit_rast, flags='g')
n=str(round(float(region_info['n']),5)) #the config file need 5 decimal for north and south
s=str(round(float(region_info['s']),5))
e=str(round(float(region_info['e']),6)) #the config file need 6 decimal for east and west
w=str(round(float(region_info['w']),6))
# Save the coordinates of the bbox in the dictionary (n,s,e,w)
landscapeunit_bbox[cat]=n+"|"+s+"|"+e+"|"+w
# Add the line to the maskedoverlayarea variable
maskedoverlayarea+="MASKEDOVERLAYAREA "+landscapeunit_rast+"|"+landscapeunit_bbox[cat]+"\n"
# Compile the content of the r.li configuration file
config_file_content="SAMPLINGFRAME 0|0|1|1\n"
config_file_content+=maskedoverlayarea
config_file_content+="RASTERMAP "+base_landcover_raster+"\n"
config_file_content+="VECTORMAP "+landscape_polygons+"\n"
# Create a new file and save the content
configfilename=base_landcover_raster.split("@")[0]+"_"+landscape_polygons.split("@")[0]
path=os.path.join(rli_dir,configfilename)
listpath.append(path)
f=open(path, 'w')
f.write(config_file_content)
f.close()
# Continue creation of r.li configuration file and landscape unit raster the rest of the landcover raster provided
while len(tmp_list)>0:
# Reinitialize 'maskedoverlayarea' variable as an empty string
maskedoverlayarea=""
# Set the current landcover raster as the first of the list
current_landcover_raster=tmp_list.pop(0) #The pop function return the first item of the list and delete it from the list at the same time
# Copy all the landscape units masks for the current landcover raster
p=Pool(ncores) #Create a pool of processes and launch them using 'map' function
func=partial(copy_landscapeunitmasks,current_landcover_raster,base_landcover_raster,landscape_polygons,landscapeunit_bbox) # Set fixed argument of the function
maskedoverlayarea=p.map(func,list_cat) # Launch the processes for as many items in the list and get the ordered results using map function
p.close()
p.join()
# Compile the content of the r.li configuration file
config_file_content="SAMPLINGFRAME 0|0|1|1\n"
config_file_content+="\n".join(maskedoverlayarea)+"\n"
config_file_content+="RASTERMAP "+current_landcover_raster+"\n"
config_file_content+="VECTORMAP "+landscape_polygons+"\n"
# Create a new file and save the content
configfilename=current_landcover_raster.split("@")[0]+"_"+landscape_polygons.split("@")[0]
path=os.path.join(rli_dir,configfilename)
listpath.append(path)
f=open(path, 'w')
f.write(config_file_content)
f.close()
# Return a list of path of configuration files creates if option actived
if returnlistpath:
return listpath
```
### Function for creation of binary raster from a categorical raster (multiprocessed)
```
###### Function creating a binary raster for each category of a base raster.
### The function run within the current region. If a category do not exists in the current region, no binary map will be produce
# 'categorical_raster' wait for the name of the base raster to be used. It is the one from which one binary raster will be produced for each category value
# 'prefix' wait for a string corresponding to the prefix of the name of the binary raster which will be produced
# 'setnull' wait for a boolean value (True, False) according to the fact that the output binary should be 1/0 or 1/null
# 'returnlistraster' wait for a boolean value (True, False) regarding to the fact that a list containing the name of binary raster is desired as return of the function
# 'category_list' wait for a list of interger corresponding to specific category of the base raster to be used
# 'ncores' wait for a integer corresponding to the number of desired cores to be used for parallelization
# Import libraries for multiprocessing
import multiprocessing
from multiprocessing import Pool
from functools import partial
def create_binary_raster(categorical_raster,prefix="binary",setnull=False,returnlistraster=True,category_list=None,ncores=2):
# Check if raster exists to avoid error in mutliprocessing
try:
mpset=categorical_raster.split("@")[1]
except:
mpset=""
if categorical_raster not in gscript.list_strings(type='raster',mapset=mpset):
sys.exit('Raster <%s> not found' %categorical_raster)
# Check for number of cores doesnt exceed available
nbcpu=multiprocessing.cpu_count()
if ncores>=nbcpu:
ncores=nbcpu-1
returnlist=[] #Declare empty list for return
#gscript.run_command('g.region', raster=categorical_raster, quiet=True) #Set the region
null='null()' if setnull else '0' #Set the value for r.mapcalc
minclass=1 if setnull else 2 #Set the value to check if the binary raster is empty
if category_list == None: #If no category_list provided
category_list=[cl for cl in gscript.parse_command('r.category',map=categorical_raster,quiet=True)]
for i,x in enumerate(category_list): #Make sure the format is UTF8 and not Unicode
category_list[i]=x.encode('UTF8')
category_list.sort(key=float) #Sort the raster categories in ascending.
p=Pool(ncores) #Create a pool of processes and launch them using 'map' function
func=partial(get_binary,categorical_raster,prefix,null,minclass) # Set the two fixed argument of the function
returnlist=p.map(func,category_list) # Launch the processes for as many items in the 'functions_name' list and get the ordered results using map function
p.close()
p.join()
if returnlistraster:
return returnlist
#### Function that extract binary raster for a specified class (called in 'create_binary_raster' function)
def get_binary(categorical_raster,prefix,null,minclass,cl):
binary_class=prefix+"_"+cl
gscript.run_command('r.mapcalc', expression=binary_class+'=if('+categorical_raster+'=='+str(cl)+',1,'+null+')',overwrite=True, quiet=True)
if len(gscript.parse_command('r.category',map=binary_class,quiet=True))>=minclass: #Check if created binary is not empty
return binary_class
else:
gscript.run_command('g.remove', quiet=True, flags="f", type='raster', name=binary_class)
```
### Function for computation of spatial metrics at landscape level (multiprocessed)
```
##### Function that compute different landscape metrics (spatial metrics) at landscape level.
### The metric computed are "dominance","pielou","renyi","richness","shannon","simpson".
### It is important to set the computation region before runing this script so that it match the extent of the 'raster' layer.
# 'configfile' wait for the path (string) to the configuration file corresponding to the 'raster' layer.
# 'raster' wait for the name (string) of the landcover map on which landscape metrics will be computed.
# 'returnlistresult' wait for a boolean value (True/False) according to the fact that a list containing the path to the result files is desired.
# 'ncores' wait for a integer corresponding to the number of desired cores to be used for parallelization.
# Import libraries for multiprocessing
import multiprocessing
from multiprocessing import Pool
from functools import partial
def compute_landscapelevel_metrics(configfile, raster, spatial_metric):
filename=raster.split("@")[0]+"_%s" %spatial_metric
outputfile=os.path.join(os.path.split(configfile)[0],"output",filename)
if spatial_metric=='renyi': # The alpha parameter was set to 2 as in https://en.wikipedia.org/wiki/R%C3%A9nyi_entropy
gscript.run_command('r.li.%s' %spatial_metric, overwrite=True,
input=raster,config=configfile,alpha='2', output=filename)
else:
gscript.run_command('r.li.%s' %spatial_metric, overwrite=True,
input=raster,config=configfile, output=filename)
return outputfile
def get_landscapelevel_metrics(configfile, raster, returnlistresult=True, ncores=2):
# Check if raster exists to avoid error in mutliprocessing
try:
mpset=raster.split("@")[1]
except:
mpset=""
if raster not in gscript.list_strings(type='raster',mapset=mpset):
sys.exit('Raster <%s> not found' %raster)
# Check if configfile exists to avoid error in mutliprocessing
if not os.path.exists(configfile):
sys.exit('Configuration file <%s> not found' %configfile)
# List of metrics to be computed
spatial_metric_list=["dominance","pielou","renyi","richness","shannon","simpson"]
# Check for number of cores doesnt exceed available
nbcpu=multiprocessing.cpu_count()
if ncores>=nbcpu:
ncores=nbcpu-1
if ncores>len(spatial_metric_list):
ncores=len(spatial_metric_list) #Adapt number of cores to number of metrics to compute
#Declare empty list for return
returnlist=[]
# Create a new pool
p=Pool(ncores)
# Set the two fixed argument of the 'compute_landscapelevel_metrics' function
func=partial(compute_landscapelevel_metrics,configfile, raster)
# Launch the processes for as many items in the 'functions_name' list and get the ordered results using map function
returnlist=p.map(func,spatial_metric_list)
p.close()
p.join()
# Return list of paths to result files
if returnlistresult:
return returnlist
```
### Function for computation of spatial metrics at class level (multiprocessed)
```
##### Function that compute different landscape metrics (spatial metrics) at class level.
### The metric computed are "patch number (patchnum)","patch density (patchdensity)","mean patch size(mps)",
### "coefficient of variation of patch area (padcv)","range of patch area size (padrange)",
### "standard deviation of patch area (padsd)", "shape index (shape)", "edge density (edgedensity)".
### It is important to set the computation region before runing this script so that it match the extent of the 'raster' layer.
# 'configfile' wait for the path (string) to the configuration file corresponding to the 'raster' layer.
# 'raster' wait for the name (string) of the landcover map on which landscape metrics will be computed.
# 'returnlistresult' wait for a boolean value (True/False) according to the fact that a list containing the path to the result files is desired.
# 'ncores' wait for a integer corresponding to the number of desired cores to be used for parallelization.
# Import libraries for multiprocessing
import multiprocessing
from multiprocessing import Pool
from functools import partial
def compute_classlevel_metrics(configfile, raster, spatial_metric):
filename=raster.split("@")[0]+"_%s" %spatial_metric
gscript.run_command('r.li.%s' %spatial_metric, overwrite=True,
input=raster,config=configfile,output=filename)
outputfile=os.path.join(os.path.split(configfile)[0],"output",filename)
return outputfile
def get_classlevel_metrics(configfile, raster, returnlistresult=True, ncores=2):
# Check if raster exists to avoid error in mutliprocessing
try:
mpset=raster.split("@")[1]
except:
mpset=""
if raster not in [x.split("@")[0] for x in gscript.list_strings(type='raster',mapset=mpset)]:
sys.exit('Raster <%s> not found' %raster)
# Check if configfile exists to avoid error in mutliprocessing
if not os.path.exists(configfile):
sys.exit('Configuration file <%s> not found' %configfile)
# List of metrics to be computed
spatial_metric_list=["patchnum","patchdensity","mps","padcv","padrange","padsd","shape","edgedensity"]
# Check for number of cores doesnt exceed available
nbcpu=multiprocessing.cpu_count()
if ncores>=nbcpu:
ncores=nbcpu-1
if ncores>len(spatial_metric_list):
ncores=len(spatial_metric_list) #Adapt number of cores to number of metrics to compute
# Declare empty list for return
returnlist=[]
# Create a new pool
p=Pool(ncores)
# Set the two fixed argument of the 'compute_classlevel_metrics' function
func=partial(compute_classlevel_metrics,configfile, raster)
# Launch the processes for as many items in the 'functions_name' list and get the ordered results using map function
returnlist=p.map(func,spatial_metric_list)
p.close()
p.join()
# Return list of paths to result files
if returnlistresult:
return returnlist
```
**-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
<center> <font size=5> <h1>User inputs</h1> </font> </center>
```
## Define a empty dictionnary for saving user inputs
user={}
## Enter the path to GRASSDATA folder
user["gisdb"] = "/home/tais/Documents/GRASSDATA_Spie2017subset_Ouaga"
## Enter the name of the location (existing or for a new one)
user["location"] = "SPIE_subset"
## Enter the EPSG code for this location
user["locationepsg"] = "32630"
## Enter the name of the mapset to use for segmentation
user["mapsetname"] = "test_rli"
```
**-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# Compute spatial metrics for deriving land use in street blocs
**Launch GRASS GIS working session**
```
## Set the name of the mapset in which to work
mapsetname=user["mapsetname"]
## Launch GRASS GIS working session in the mapset
if os.path.exists(os.path.join(user["gisdb"],user["location"],mapsetname)):
gsetup.init(os.environ['GISBASE'], user["gisdb"], user["location"], mapsetname)
print "You are now working in mapset '"+mapsetname+"'"
else:
print "'"+mapsetname+"' mapset doesn't exists in "+user["gisdb"]
```
**-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
### Create binary rasters from the base landcover map
```
# Save time for computing processin time
begintime=time.time()
# Set the name of the 'base' landcover map
baselandcoverraster="classif@test_rli"
# Create as many binary raster layer as categorical values existing in the base landcover map
gscript.run_command('g.region', raster=baselandcoverraster, quiet=True) #Set the region
pref=baselandcoverraster.split("@")[0]+"_cl" #Set the prefix
raster_list=[] # Initialize a empty list for results
raster_list=create_binary_raster(baselandcoverraster,
prefix=pref,setnull=True,returnlistraster=True,
category_list=None,ncores=15) #Extract binary raster
# Compute and print processing time
print_processing_time(begintime,"Extraction of binary rasters achieved in ")
# Insert the name of the base landcover map at first position in the list
raster_list.insert(0,baselandcoverraster)
# Display the raster to be used for landscape analysis
raster_list
```
## Create r.li configuration file for a list of landcover rasters
```
# Save time for computing processin time
begintime=time.time()
# Set the name of the vector polygon layer containing the
landscape_polygons="streetblocks@PERMANENT"
# Run creation of r.li configuration file and associated raster layers
list_configfile=create_rli_configfile(raster_list,landscape_polygons,returnlistpath=True,ncores=20)
# Compute and print processing time
print_processing_time(begintime,"Extraction of binary rasters achieved in ")
# Display the path to the configuration files created
list_configfile
```
**-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
## Compute spatial metrics at landscape level
```
# Initialize an empty list which will contains the resultfiles
resultfiles=[]
# Save time for computing processin time
begintime=time.time()
# Get the path to the configuration file for the base landcover raster
currentconfigfile=list_configfile[0]
# Get the name of the base landcover raster
currentraster=raster_list[0]
# Set the region to match the extent of the base raster
gscript.run_command('g.region', raster=currentraster, quiet=True)
# Launch the processes for as many items in the 'functions_name' list and get the ordered results using map function
resultfiles.append(get_landscapelevel_metrics(currentconfigfile, currentraster, returnlistresult=True, ncores=15))
# Compute and print processing time
print_processing_time(begintime,"Computation of spatial metric achieved in ")
resultfiles
```
## Compute spatial metrics at class level
```
# Save time for computing processin time
begintime=time.time()
# Get a list with paths to the configuration file for class level metrics
classlevelconfigfiles=list_configfile[1:]
# Get a list with name of binary landcover raster for class level metrics
classlevelrasters=raster_list[1:]
for x,currentraster in enumerate(classlevelrasters[:]):
# Get the path to the configuration file for the base landcover raster
currentconfigfile=classlevelconfigfiles[x]
# Set the region to match the extent of the base raster
gscript.run_command('g.region', raster=currentraster, quiet=True)
# Launch the processes for as many items in the 'functions_name' list and get the ordered results using map function
resultfiles.append(get_classlevel_metrics(currentconfigfile, currentraster, returnlistresult=True, ncores=10))
# Compute and print processing time
print_processing_time(begintime,"Computation of spatial metric achieved in ")
resultfiles
# Flat the 'resultfiles' list which contains several lists
resultfiles=[item for sublist in resultfiles for item in sublist]
resultfiles
```
## Compute some special metrics
### Mean and standard deviation of NDVI
### Mean and standard deviation of SAR textures
### Mean and standard deviation of building's height
# Importing the NDVI layer
```
break
## Saving current time for processing time management
begintime_ndvi=time.time()
## Import nDSM imagery
print ("Importing NDVI raster imagery at " + time.ctime())
gscript.run_command('r.import',
input="/media/tais/data/MAUPP/WorldView3_Ouagadougou/Orthorectified/mosaique_georef/NDVI/ndvi_georef_ordre2.TIF",
output="ndvi", overwrite=True)
# Mask null/nodata values
gscript.run_command('r.null', map="ndvi")
print_processing_time(begintime_ndvi, "imagery has been imported in ")
```
# Importing the nDSM layer
```
break
## Saving current time for processing time management
begintime_ndsm=time.time()
## Import nDSM imagery
print ("Importing nDSM raster imagery at " + time.ctime())
grass.run_command('r.import',
input="/media/tais/data/MAUPP/WorldView3_Ouagadougou/Orthorectified/mosaique_georef/nDSM/nDSM_mosaik_georef_ordre2.tif",
output="ndsm", overwrite=True)
## Define null value for specific value in nDSM raster. Adapt the value to your own data.
# If there is no null value in your data, comment the next line
grass.run_command('r.null', map="ndsm", setnull="-999")
# Make histogram equalisation on grey color.
grass.run_command('r.colors', flags='e', map='ndsm', color='grey')
print_processing_time(begintime_ndsm, "nDSM imagery has been imported in ")
```
### Masking the nDSM artifacts
```
break
# Import vector with nDSM artifacts zones
grass.run_command('v.in.ogr', overwrite=True,
input="/media/tais/data/MAUPP/WorldView3_Ouagadougou/Masque_artifacts_nDSM/Ouaga_mask_artifacts_nDSM.shp",
output="mask_artifacts_ndsm")
## Set computational region to match the default region
grass.run_command('g.region', overwrite=True, raster="ndsm")
# Rasterize the vector layer, with value "0" on the artifacts zones
grass.run_command('v.to.rast', input='mask_artifacts_ndsm', output='mask_artifacts_ndsm',
use='val', value='0', memory='5000')
## Set computational region to match the default region
grass.run_command('g.region', overwrite=True, raster="ndsm")
## Create a new nDSM with artifacts filled with '0' value
formula='tmp_artifact=nmin(ndsm,mask_artifacts_ndsm)'
grass.mapcalc(formula, overwrite=True)
## Remove the artifact mask
grass.run_command('g.remove', flags='f', type='raster', name="mask_artifacts_ndsm")
## Rename the new nDSM
grass.run_command('g.rename', raster='tmp_artifact,ndsm', overwrite=True)
## Remove the intermediate nDSM layer
grass.run_command('g.remove', flags='f', type='raster', name="tmp_artifact")
```
**-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# Define input raster for computing statistics of segments
```
## Display the name of rasters available in PERMANENT and CLASSIFICATION mapset
print grass.read_command('g.list',type="raster", mapset="PERMANENT", flags='rp')
print grass.read_command('g.list',type="raster", mapset=user["classificationA_mapsetname"], flags='rp')
## Define the list of raster layers for which statistics will be computed
inputstats=[]
inputstats.append("opt_blue")
inputstats.append("opt_green")
inputstats.append("opt_red")
inputstats.append("opt_nir")
inputstats.append("ndsm")
inputstats.append("ndvi")
print "Layer to be used to compute raster statistics of segments:\n"+'\n'.join(inputstats)
## Define the list of raster statistics to be computed for each raster layer
rasterstats=[]
rasterstats.append("min")
rasterstats.append("max")
rasterstats.append("range")
rasterstats.append("mean")
rasterstats.append("stddev")
#rasterstats.append("coeff_var") # Seems that this statistic create null values
rasterstats.append("median")
rasterstats.append("first_quart")
rasterstats.append("third_quart")
rasterstats.append("perc_90")
print "Raster statistics to be computed:\n"+'\n'.join(rasterstats)
## Define the list of area measures (segment's shape statistics) to be computed
areameasures=[]
areameasures.append("area")
areameasures.append("perimeter")
areameasures.append("compact_circle")
areameasures.append("compact_square")
areameasures.append("fd")
print "Area measures to be computed:\n"+'\n'.join(areameasures)
```
**-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
<center> <font size=5> <h1>Compute objects' statistics</h1> </font> </center>
```
## Saving current time for processing time management
begintime_computeobjstat=time.time()
```
## Define the folder where to save the results and create it if necessary
In the next cell, please adapt the path to the directory where you want to save the .csv output of i.segment.uspo.
```
## Folder in which save processing time output
outputfolder="/media/tais/My_Book_1/MAUPP/Traitement/Ouagadougou/Segmentation_fullAOI_localapproach/Results/CLASSIF/stats_optical"
## Create the folder if does not exists
if not os.path.exists(outputfolder):
os.makedirs(outputfolder)
print "Folder '"+outputfolder+"' created"
```
### Copy data from other mapset to the current mapset
Some data need to be copied from other mapsets into the current mapset.
### Remove current mask
```
## Check if there is a raster layer named "MASK"
if not grass.list_strings("rast", pattern="MASK", mapset=mapsetname, flag='r'):
print 'There is currently no MASK'
else:
## Remove the current MASK layer
grass.run_command('r.mask',flags='r')
print 'The current MASK has been removed'
```
***Copy segmentation raster***
```
## Copy segmentation raster layer from SEGMENTATION mapset to current mapset
grass.run_command('g.copy', overwrite=True,
raster="segmentation_raster@"+user["segmentation_mapsetname"]+",segments")
```
***Copy morphological zone (raster)***
```
## Copy segmentation raster layer from SEGMENTATION mapset to current mapset
grass.run_command('g.copy', overwrite=True,
raster="zone_morpho@"+user["segmentation_mapsetname"]+",zone_morpho")
```
***Copy morphological zone (vector)***
```
## Copy segmentation raster layer from SEGMENTATION mapset to current mapset
grass.run_command('g.copy', overwrite=True,
vector="zone_morpho@"+user["segmentation_mapsetname"]+",zone_morpho")
```
**-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# Compute statistics of segments (Full AOI extend)
### Compute statistics of segment using i.segment.stats
The process is make to compute statistics iteratively for each morphological zones, used here as tiles.
This section uses the ['i.segment.stats' add-on](https://grass.osgeo.org/grass70/manuals/addons/i.segment.stats.html) to compute statistics for each object.
```
## Save name of the layer to be used as tiles
tile_layer='zone_morpho'+'@'+mapsetname
## Save name of the segmentation layer to be used by i.segment.stats
segment_layer='segments'+'@'+mapsetname
## Save name of the column containing area_km value
area_column='area_km2'
## Save name of the column containing morphological type value
type_column='type'
## Save the prefix to be used for the outputfiles of i.segment.stats
prefix="Segstat"
## Save the list of polygons to be processed (save the 'cat' value)
listofregion=list(grass.parse_command('v.db.select', map=tile_layer,
columns='cat', flags='c'))[:]
for count, cat in enumerate(listofregion):
print str(count)+" cat:"+str(cat)
```
```
## Initialize a empty string for saving print outputs
txtcontent=""
## Running i.segment.stats
messagetoprint="Start computing statistics for segments to be classified, using i.segment.stats on "+time.ctime()+"\n"
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
begintime_isegmentstats=time.time()
## Compute total area to be processed for process progression information
processed_area=0
nbrtile=len(listofregion)
attributes=grass.parse_command('db.univar', flags='g', table=tile_layer.split("@")[0], column=area_column, driver='sqlite')
total_area=float(attributes['sum'])
messagetoprint=str(nbrtile)+" region(s) will be processed, covering an area of "+str(round(total_area,3))+" Sqkm."+"\n\n"
print (messagetoprint)
txtcontent+=messagetoprint
## Save time before looping
begintime_isegmentstats=time.time()
## Start loop on morphological zones
count=1
for cat in listofregion[:]:
## Save current time at loop' start.
begintime_current_id=time.time()
## Create a computional region for the current polygon
condition="cat="+cat
outputname="tmp_"+cat
grass.run_command('v.extract', overwrite=True, quiet=True,
input=tile_layer, type='area', where=condition, output=outputname)
grass.run_command('g.region', overwrite=True, vector=outputname, align=segment_layer)
grass.run_command('r.mask', overwrite=True, raster=tile_layer, maskcats=cat)
grass.run_command('g.remove', quiet=True, type="vector", name=outputname, flags="f")
## Save size of the current polygon and add it to the already processed area
size=round(float(grass.read_command('v.db.select', map=tile_layer,
columns=area_column, where=condition,flags="c")),2)
## Print
messagetoprint="Computing segments's statistics for tile n°"+str(cat)
messagetoprint+=" ("+str(count)+"/"+str(len(listofregion))+")"
messagetoprint+=" corresponding to "+str(size)+" km2"
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
## Define the csv output file name, according to the optimization function selected
outputcsv=os.path.join(outputfolder,prefix+"_"+str(cat)+".csv")
## Compute statistics of objets using i.segment.stats only with .csv output (no vectormap output).
grass.run_command('i.segment.stats', overwrite=True, map=segment_layer,
rasters=','.join(inputstats), raster_statistics=','.join(rasterstats),
area_measures=','.join(areameasures), csvfile=outputcsv, processes='20')
## Add the size of the zone to the already processed area
processed_area+=size
## Print
messagetoprint=print_processing_time(begintime_current_id,
"i.segment.stats finishes to process th current tile in ")
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
remainingtile=nbrtile-count
if remainingtile>0:
messagetoprint=str(round((processed_area/total_area)*100,2))+" percent of the total area processed. "
messagetoprint+="Still "+str(remainingtile)+" zone(s) to process."+"\n"
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
else:
messagetoprint="\n"
print (messagetoprint)
txtcontent+=messagetoprint
## Adapt the count
count+=1
## Remove current mask
grass.run_command('r.mask', flags='r')
## Compute processing time and print it
messagetoprint=print_processing_time(begintime_isegmentstats, "Statitics computed in ")
print (messagetoprint)
txtcontent+=messagetoprint
#### Write text file with log of processing time
## Create the .txt file for processing time output and begin to write
f = open(os.path.join(outputfolder,mapsetname+"_processingtime_isegmentstats.txt"), 'w')
f.write(mapsetname+" processing time information for i.segment.stats"+"\n\n")
f.write(txtcontent)
f.close()
## print
print_processing_time(begintime_computeobjstat,"Object statistics computed in ")
```
## Concatenate individuals .csv files and replace unwanted values
BE CAREFUL! Before runing the following cells, please check your data to be sure that it makes sens to replace the 'nan', 'null', or 'inf' values with "0"
```
## Define the outputfile for .csv containing statistics for all segments
outputfile=os.path.join(outputfolder,"all_segments_stats.csv")
print outputfile
# Create a dictionary with 'key' to be replaced by 'values'
findreplacedict={}
findreplacedict['nan']="0"
findreplacedict['null']="0"
findreplacedict['inf']="0"
# Define pattern of file to concatenate
pat=prefix+"_*.csv"
sep="|"
## Initialize a empty string for saving print outputs
txtcontent=""
## Saving current time for processing time management
begintime_concat=time.time()
## Print
messagetoprint="Start concatenate individual .csv files and replacing unwanted values."
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
# Concatenate and replace unwanted values
messagetoprint=concat_findreplace(outputfolder,pat,sep,findreplacedict,outputfile)
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
## Compute processing time and print it
messagetoprint=print_processing_time(begintime_concat, "Process achieved in ")
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
#### Write text file with log of processing time
## Create the .txt file for processing time output and begin to write
filepath=os.path.join(outputfolder,mapsetname+"_processingtime_concatreplace.txt")
f = open(filepath, 'w')
f.write(mapsetname+" processing time information for concatenation of individual .csv files and replacing of unwanted values."+"\n\n")
f.write(txtcontent)
f.close()
```
# Create new database in postgresql
```
# User for postgresql connexion
dbuser="tais"
# Password of user
dbpassword="tais"
# Host of database
host="localhost"
# Name of the new database
dbname="ouaga_fullaoi_localsegment"
# Set name of schema for objects statistics
stat_schema="statistics"
# Set name of table with statistics of segments - FOR OPTICAL
object_stats_table="object_stats_optical"
break
from psycopg2.extensions import ISOLATION_LEVEL_AUTOCOMMIT
# Connect to postgres database
db=None
db=pg.connect(dbname='postgres', user=dbuser, password=dbpassword, host=host)
# Allow to create a new database
db.set_isolation_level(ISOLATION_LEVEL_AUTOCOMMIT)
# Execute the CREATE DATABASE query
cur=db.cursor()
#cur.execute('DROP DATABASE IF EXISTS ' + dbname) #Comment this to avoid deleting existing DB
cur.execute('CREATE DATABASE ' + dbname)
cur.close()
db.close()
```
### Create PostGIS Extension in the database
```
break
# Connect to the database
db=pg.connect(database=dbname, user=dbuser, password=dbpassword, host=host)
# Open a cursor to perform database operations
cur=db.cursor()
# Execute the query
cur.execute('CREATE EXTENSION IF NOT EXISTS postgis')
# Make the changes to the database persistent
db.commit()
# Close connection with database
cur.close()
db.close()
```
<center> <font size=4> <h2>Import statistics of segments in a Postgresql database</h2> </font> </center>
## Create new schema in the postgresql database
```
schema=stat_schema
from psycopg2.extensions import ISOLATION_LEVEL_AUTOCOMMIT
# Connect to postgres database
db=None
db=pg.connect(dbname=dbname, user='tais', password='tais', host='localhost')
# Allow to create a new database
db.set_isolation_level(ISOLATION_LEVEL_AUTOCOMMIT)
# Execute the CREATE DATABASE query
cur=db.cursor()
#cur.execute('DROP SCHEMA IF EXISTS '+schema+' CASCADE') #Comment this to avoid deleting existing DB
try:
cur.execute('CREATE SCHEMA '+schema)
except Exception as e:
print ("Exception occured : "+str(e))
cur.close()
db.close()
```
## Create a new table
```
# Connect to an existing database
db=pg.connect(database=dbname, user=dbuser, password=dbpassword, host=host)
# Open a cursor to perform database operations
cur=db.cursor()
# Drop table if exists:
cur.execute("DROP TABLE IF EXISTS "+schema+"."+object_stats_table)
# Make the changes to the database persistent
db.commit()
import csv
# Create a empty list for saving of column name
column_name=[]
# Create a reader for the first csv file in the stack of csv to be imported
pathtofile=os.path.join(outputfolder, outputfile)
readercsvSubset=open(pathtofile)
readercsv=csv.reader(readercsvSubset, delimiter='|')
headerline=readercsv.next()
print "Create a new table '"+schema+"."+object_stats_table+"' with header corresponding to the first row of file '"+pathtofile+"'"
## Build a query for creation of a new table with auto-incremental key-value (thus avoiding potential duplicates of 'cat' value)
# All column data-types are set to 'text' in order to be able to import some 'nan', 'inf' or 'null' values present in statistics files
# This table will allow to import all individual csv files in a single Postgres table, which will be cleaned after
query="CREATE TABLE "+schema+"."+object_stats_table+" ("
query+="key_value serial PRIMARY KEY"
query+=", "+str(headerline[0])+" text"
column_name.append(str(headerline[0]))
for column in headerline[1:]:
if column[0] in ('1','2','3','4','5','6','7','8','9','0'):
query+=","
query+=" "+"W"+str(column)+" double precision"
column_name.append("W"+str(column))
else:
query+=","
query+=" "+str(column)+" double precision"
column_name.append(str(column))
query+=")"
# Execute the CREATE TABLE query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
# Close cursor and communication with the database
cur.close()
db.close()
```
## Copy objects statistics from csv to Postgresql database
```
# Connect to an existing database
db=pg.connect(database=dbname, user=dbuser, password=dbpassword, host=host)
# Open a cursor to perform database operations
cur=db.cursor()
## Initialize a empty string for saving print outputs
txtcontent=""
## Saving current time for processing time management
begintime_copy=time.time()
## Print
messagetoprint="Start copy of segments' statistics in the postgresql table '"+schema+"."+object_stats_table+"'"
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
# Create query for copy data from csv, avoiding the header, and updating only the column which are in the csv (to allow auto-incremental key value to wokr)
query="COPY "+schema+"."+object_stats_table+"("+', '.join(column_name)+") "
query+=" FROM '"+str(pathtofile)+"' HEADER DELIMITER '|' CSV;"
# Execute the COPY FROM CSV query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
## Compute processing time and print it
messagetoprint=print_processing_time(begintime_copy, "Process achieved in ")
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
#### Write text file with log of processing time
## Create the .txt file for processing time output and begin to write
filepath=os.path.join(outputfolder,mapsetname+"_processingtime_PostGimport.txt")
f = open(filepath, 'w')
f.write(mapsetname+" processing time information for importation of segments' statistics in the PostGreSQL Database."+"\n\n")
f.write(txtcontent)
f.close()
# Close cursor and communication with the database
cur.close()
db.close()
```
# Drop duplicate values of CAT
Here, we will find duplicates. Indeed, as statistics are computed for each tile (morphological area) and computational region aligned to the pixels raster, some objets could appear in two different tile resulting on duplicates on "CAT" column.
We firs select the "CAT" of duplicated objets and then puting them in a list. Then, for each duplicated "CAT", we select the key-value (primary key) of the smallest object (area_min). The row corresponding to those key-values are then remoed using the "DELETE FROM" query.
```
# Connect to an existing database
db=pg.connect(database=dbname, user=dbuser, password=dbpassword, host=host)
# Open a cursor to perform database operations
cur=db.cursor()
## Initialize a empty string for saving print outputs
txtcontent=""
## Saving current time for processing time management
begintime_removeduplic=time.time()
## Print
messagetoprint="Start removing duplicates in the postgresql table '"+schema+"."+object_stats_table+"'"
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
# Find duplicated 'CAT'
find_duplicated_cat()
# Remove duplicated
count_pass=1
count_removedduplic=0
while len(cattodrop)>0:
messagetoprint="Removing duplicates - Pass "+str(count_pass)
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
find_duplicated_key()
remove_duplicated_key()
messagetoprint=str(len(keytodrop))+" duplicates removed."
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
count_removedduplic+=len(keytodrop)
# Find again duplicated 'CAT'
find_duplicated_cat()
count_pass+=1
messagetoprint="A total of "+str(count_removedduplic)+" duplicates were removed."
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
## Compute processing time and print it
messagetoprint=print_processing_time(begintime_removeduplic, "Process achieved in ")
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
#### Write text file with log of processing time
## Create the .txt file for processing time output and begin to write
filepath=os.path.join(outputfolder,mapsetname+"_processingtime_RemoveDuplic.txt")
f = open(filepath, 'w')
f.write(mapsetname+" processing time information for removing duplicated objects."+"\n\n")
f.write(txtcontent)
f.close()
# Vacuum the current Postgresql database
vacuum(db)
```
# Change the primary key from 'key_value' to 'cat'
```
# Connect to an existing database
db=pg.connect(database=dbname, user=dbuser, password=dbpassword, host=host)
# Open a cursor to perform database operations
cur=db.cursor()
# Build a query to drop the current constraint on primary key
query="ALTER TABLE "+schema+"."+object_stats_table+" \
DROP CONSTRAINT "+object_stats_table+"_pkey"
# Execute the query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
# Build a query to change the datatype of 'cat' to 'integer'
query="ALTER TABLE "+schema+"."+object_stats_table+" \
ALTER COLUMN cat TYPE integer USING cat::integer"
# Execute the query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
# Build a query to add primary key on 'cat'
query="ALTER TABLE "+schema+"."+object_stats_table+" \
ADD PRIMARY KEY (cat)"
# Execute the query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
# Build a query to drop column 'key_value'
query="ALTER TABLE "+schema+"."+object_stats_table+" \
DROP COLUMN key_value"
# Execute the query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
# Vacuum the current Postgresql database
vacuum(db)
# Close cursor and communication with the database
cur.close()
db.close()
```
### Show first rows of statistics
```
# Connect to an existing database
db=pg.connect(database=dbname, user=dbuser, password=dbpassword, host=host)
# Number of line to show (please limit to 100 for saving computing time)
nbrow=15
# Query
query="SELECT * FROM "+schema+"."+object_stats_table+" \
ORDER BY cat \
ASC LIMIT "+str(nbrow)
# Execute query through panda
df=pd.read_sql(query, db)
# Show dataframe
df.head(15)
```
<left> <font size=4> <b> End of classification part </b> </font> </left>
```
print("The script ends at "+ time.ctime())
print_processing_time(begintime_segmentation_full, "Entire process has been achieved in ")
```
**-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
| github_jupyter |
## Using SageMaker Debugger and SageMaker Experiments for iterative model pruning
This notebook demonstrates how we can use [SageMaker Debugger](https://docs.aws.amazon.com/sagemaker/latest/dg/train-debugger.html) and [SageMaker Experiments](https://docs.aws.amazon.com/sagemaker/latest/dg/experiments.html) to perform iterative model pruning. Let's start first with a quick introduction into model pruning.
State of the art deep learning models consist of millions of parameters and are trained on very large datasets. For transfer learning we take a pre-trained model and fine-tune it on a new and typically much smaller dataset. The new dataset may even consist of different classes, so the model is basically learning a new task. This process allows us to quickly achieve state of the art results without having to design and train our own model from scratch. However, it may happen that a much smaller and simpler model would also perform well on our dataset. With model pruning we identify the importance of weights during training and remove the weights that are contributing very little to the learning process. We can do this in an iterative way where we remove a small percentage of weights in each iteration. Removing means to eliminate the entries in the tensor so its size shrinks.
We use SageMaker Debugger to get weights, activation outputs and gradients during training. These tensors are used to compute the importance of weights. We will use SageMaker Experiments to keep track of each pruning iteration: if we prune too much we may degrade model accuracy, so we will monitor number of parameters versus validation accuracy.
```
! pip -q install sagemaker
! pip -q install sagemaker-experiments
```
### Get training dataset
Next we get the [Caltech101](http://www.vision.caltech.edu/Image_Datasets/Caltech101/) dataset. This dataset consists of 101 image categories.
```
import tarfile
import requests
import os
filename = '101_ObjectCategories.tar.gz'
data_url = os.path.join("https://s3.us-east-2.amazonaws.com/mxnet-public", filename)
r = requests.get(data_url, stream=True)
with open(filename, 'wb') as f:
for chunk in r.iter_content(chunk_size=1024):
if chunk:
f.write(chunk)
print('Extracting {} ...'.format(filename))
tar = tarfile.open(filename, "r:gz")
tar.extractall('.')
tar.close()
print('Data extracted.')
```
And upload it to our SageMaker default bucket:
```
import sagemaker
import boto3
def upload_to_s3(path, directory_name, bucket, counter=-1):
print("Upload files from" + path + " to " + bucket)
client = boto3.client('s3')
for path, subdirs, files in os.walk(path):
path = path.replace("\\","/")
print(path)
for file in files[0:counter]:
client.upload_file(os.path.join(path, file), bucket, directory_name+'/'+path.split("/")[-1]+'/'+file)
boto_session = boto3.Session()
sagemaker_session = sagemaker.Session(boto_session=boto_session)
bucket = sagemaker_session.default_bucket()
upload_to_s3("101_ObjectCategories", directory_name="101_ObjectCategories_train", bucket=bucket)
#we will compute saliency maps for all images in the test dataset, so we will only upload 4 images
upload_to_s3("101_ObjectCategories_test", directory_name="101_ObjectCategories_test", bucket=bucket, counter=4)
```
### Load and save ResNet model
First we load a pre-trained [ResNet](https://arxiv.org/abs/1512.03385) model from PyTorch model zoo.
```
import torch
from torchvision import models
from torch import nn
model = models.resnet18(pretrained=True)
```
Let's have a look on the model architecture:
```
model
```
As we can see above, the last Linear layer outputs 1000 values, which is the number of classes the model has originally been trained on. Here, we will fine-tune the model on the Caltech101 dataset: as it has only 101 classes, we need to set the number of output classes to 101.
```
nfeatures = model.fc.in_features
model.fc = torch.nn.Linear(nfeatures, 101)
```
Next we store the model definition and weights in an output file.
**IMPORTANT**: the model file will be used by the training job. To avoid version conflicts, you need to ensure that your notebook is running a Jupyter kernel with PyTorch version 1.6.
```
checkpoint = {'model': model,
'state_dict': model.state_dict()}
torch.save(checkpoint, 'src/model_checkpoint')
```
The following code cell creates a SageMaker experiment:
```
import boto3
from datetime import datetime
from smexperiments.experiment import Experiment
sagemaker_boto_client = boto3.client("sagemaker")
#name of experiment
timestep = datetime.now()
timestep = timestep.strftime("%d-%m-%Y-%H-%M-%S")
experiment_name = timestep + "-model-pruning-experiment"
#create experiment
Experiment.create(
experiment_name=experiment_name,
description="Iterative model pruning of ResNet trained on Caltech101",
sagemaker_boto_client=sagemaker_boto_client)
```
The following code cell defines a list of tensor names that be used to compute filter ranks. The lists are defined in the Python script `model_resnet`.
```
import model_resnet
activation_outputs = model_resnet.activation_outputs
gradients = model_resnet.gradients
```
### Iterative model pruning: step by step
Before we jump into the code for running the iterative model pruning we will walk through the code step by step.
#### Step 0: Create trial and debugger hook coonfiguration
First we create a new trial for each pruning iteration. That allows us to track our training jobs and see which models have the lowest number of parameters and best accuracy. We use the `smexperiments` library to create a trial within our experiment.
```
from smexperiments.trial import Trial
trial = Trial.create(
experiment_name=experiment_name,
sagemaker_boto_client=sagemaker_boto_client
)
```
Next we define the experiment_config which is a dictionary that will be passed to the SageMaker training.
```
experiment_config = { "ExperimentName": experiment_name,
"TrialName": trial.trial_name,
"TrialComponentDisplayName": "Training"}
```
We create a debugger hook configuration to define a custom collection of tensors to be emitted. The custom collection contains all weights and biases of the model. It also includes individual layer outputs and their gradients which will be used to compute filter ranks. Tensors are saved every 100th iteration where an iteration represents one forward and backward pass.
```
from sagemaker.debugger import DebuggerHookConfig, CollectionConfig
debugger_hook_config = DebuggerHookConfig(
collection_configs=[
CollectionConfig(
name="custom_collection",
parameters={ "include_regex": ".*relu|.*weight|.*bias|.*running_mean|.*running_var|.*CrossEntropyLoss",
"save_interval": "100" })])
```
#### Step 1: Start training job
Now we define the SageMaker PyTorch Estimator. We will train the model on an `ml.p2.xlarge` instance. The model definition plus training code is defined in the entry_point file `train.py`.
```
import sagemaker
from sagemaker.pytorch import PyTorch
estimator = PyTorch(role=sagemaker.get_execution_role(),
train_instance_count=1,
train_instance_type='ml.p3.2xlarge',
train_volume_size=400,
source_dir='src',
entry_point='train.py',
framework_version='1.6',
py_version='py3',
metric_definitions=[ {'Name':'train:loss', 'Regex':'loss:(.*?)'}, {'Name':'eval:acc', 'Regex':'acc:(.*?)'} ],
enable_sagemaker_metrics=True,
hyperparameters = {'epochs': 10},
debugger_hook_config=debugger_hook_config
)
```
Once we have defined the estimator object we can call `fit` which creates a ml.p2.xlarge instance on which it starts the training. We pass the experiment_config which associates the training job with a trial and an experiment. If we don't specify an `experiment_config` the training job will appear in SageMaker Experiments under `Unassigned trial components`
```
estimator.fit(inputs={'train': 's3://{}/101_ObjectCategories_train'.format(bucket),
'test': 's3://{}/101_ObjectCategories_test'.format(bucket)},
experiment_config=experiment_config)
```
#### Step 2: Get gradients, weights, biases
Once the training job has finished, we will retrieve its tensors, such as gradients, weights and biases. We use the `smdebug` library which provides functions to read and filter tensors. First we create a [trial](https://github.com/awslabs/sagemaker-debugger/blob/master/docs/analysis.md#Trial) that is reading the tensors from S3.
For clarification: in the context of SageMaker Debugger a trial is an object that lets you query tensors for a given training job. In the context of SageMaker Experiments a trial is part of an experiment and it presents a collection of training steps involved in a single training job.
```
from smdebug.trials import create_trial
path = estimator.latest_job_debugger_artifacts_path()
smdebug_trial = create_trial(path)
```
To access tensor values, we only need to call `smdebug_trial.tensor()`. For instance to get the outputs of the first ReLU activation at step 0 we run `smdebug_trial.tensor('layer4.1.relu_0_output_0').value(0, mode=modes.TRAIN)`. Next we compute a filter rank for the convolutions.
Some defintions: a filter is a collection of kernels (one kernel for every single input channel) and a filter produces one feature map (output channel). In the image below the convolution creates 64 feature maps (output channels) and uses a kernel of 5x5. By pruning a filter, an entire feature map will be removed. So in the example image below the number of feature maps (output channels) would shrink to 63 and the number of learnable parameters (weights) would be reduced by 1x5x5.

#### Step 3: Compute filter ranks
In this notebook we compute filter ranks as described in the article ["Pruning Convolutional Neural Networks for Resource Efficient Inference"](https://arxiv.org/pdf/1611.06440.pdf) We basically identify filters that are less important for the final prediction of the model. The product of weights and gradients can be seen as a measure of importance. The product has the dimension `(batch_size, out_channels, width, height)` and we get the average over `axis=0,2,3` to have a single value (rank) for each filter.
In the following code we retrieve activation outputs and gradients and compute the filter rank.
```
import numpy as np
from smdebug import modes
def compute_filter_ranks(smdebug_trial, activation_outputs, gradients):
filters = {}
for activation_output_name, gradient_name in zip(activation_outputs, gradients):
for step in smdebug_trial.steps(mode=modes.TRAIN):
activation_output = smdebug_trial.tensor(activation_output_name).value(step, mode=modes.TRAIN)
gradient = smdebug_trial.tensor(gradient_name).value(step, mode=modes.TRAIN)
rank = activation_output * gradient
rank = np.mean(rank, axis=(0,2,3))
if activation_output_name not in filters:
filters[activation_output_name] = 0
filters[activation_output_name] += rank
return filters
filters = compute_filter_ranks(smdebug_trial, activation_outputs, gradients)
```
Next we normalize the filters:
```
def normalize_filter_ranks(filters):
for activation_output_name in filters:
rank = np.abs(filters[activation_output_name])
rank = rank / np.sqrt(np.sum(rank * rank))
filters[activation_output_name] = rank
return filters
filters = normalize_filter_ranks(filters)
```
We create a list of filters, sort it by rank and retrieve the smallest values:
```
def get_smallest_filters(filters, n):
filters_list = []
for layer_name in sorted(filters.keys()):
for channel in range(filters[layer_name].shape[0]):
filters_list.append((layer_name, channel, filters[layer_name][channel], ))
filters_list.sort(key = lambda x: x[2])
filters_list = filters_list[:n]
print("The", n, "smallest filters", filters_list)
return filters_list
filters_list = get_smallest_filters(filters, 100)
```
#### Step 4 and step 5: Prune low ranking filters and set new weights
Next we prune the model, where we remove filters and their corresponding weights.
```
step = smdebug_trial.steps(mode=modes.TRAIN)[-1]
model = model_resnet.prune(model,
filters_list,
smdebug_trial,
step)
```
#### Step 6: Start next pruning iteration
Once we have pruned the model, the new architecture and pruned weights will be saved under src and will be used by the next training job in the next pruning iteration.
```
# save pruned model
checkpoint = {'model': model,
'state_dict': model.state_dict()}
torch.save(checkpoint, 'src/model_checkpoint')
#clean up
del model
```
#### Overall workflow
The overall workflow looks like the following:

### Run iterative model pruning
After having gone through the code step by step, we are ready to run the full worfklow. The following cell runs 1 pruning iteration for a tutorial purpose. Change the range of the for loop to 10 to replicate the same result shown in the [Pruning machine learning models with Amazon SageMaker Debugger and Amazon SageMaker Experiments blog](https://aws.amazon.com/blogs/machine-learning/pruning-machine-learning-models-with-amazon-sagemaker-debugger-and-amazon-sagemaker-experiments/) and the figure below the cell. In each iteration a new SageMaker training job is started, where it emits gradients and activation outputs to Amazon S3. Once the job has finished, filter ranks are computed and the 100 smallest filters are removed.
```
# start iterative pruning
for pruning_step in range(1):
#create new trial for this pruning step
smexperiments_trial = Trial.create(
experiment_name=experiment_name,
sagemaker_boto_client=sagemaker_boto_client
)
experiment_config["TrialName"] = smexperiments_trial.trial_name
print("Created new trial", smexperiments_trial.trial_name, "for pruning step", pruning_step)
#start training job
estimator = PyTorch(role=sagemaker.get_execution_role(),
train_instance_count=1,
train_instance_type='ml.p3.2xlarge',
train_volume_size=400,
source_dir='src',
entry_point='train.py',
framework_version='1.6',
py_version='py3',
metric_definitions=[ {'Name':'train:loss', 'Regex':'loss:(.*?)'}, {'Name':'eval:acc', 'Regex':'acc:(.*?)'} ],
enable_sagemaker_metrics=True,
hyperparameters = {'epochs': 10},
debugger_hook_config = debugger_hook_config
)
#start training job
estimator.fit(inputs={'train': 's3://{}/101_ObjectCategories_train'.format(bucket),
'test': 's3://{}/101_ObjectCategories_test'.format(bucket)},
experiment_config=experiment_config)
print("Training job", estimator.latest_training_job.name, " finished.")
# read tensors
path = estimator.latest_job_debugger_artifacts_path()
smdebug_trial = create_trial(path)
# compute filter ranks and get 100 smallest filters
filters = compute_filter_ranks(smdebug_trial, activation_outputs, gradients)
filters_normalized = normalize_filter_ranks(filters)
filters_list = get_smallest_filters(filters_normalized, 100)
#load previous model
checkpoint = torch.load("src/model_checkpoint")
model = checkpoint['model']
model.load_state_dict(checkpoint['state_dict'])
#prune model
step = smdebug_trial.steps(mode=modes.TRAIN)[-1]
model = model_resnet.prune(model,
filters_list,
smdebug_trial,
step)
print("Saving pruned model")
# save pruned model
checkpoint = {'model': model,
'state_dict': model.state_dict()}
torch.save(checkpoint, 'src/model_checkpoint')
#clean up
del model
```
As the iterative model pruning is running, we can track and visualize our experiment in SageMaker Studio. In our training script we use SageMaker debugger's `save_scalar` method to store the number of parameters in the model and the model accuracy. So we can visualize those in Studio or use the `ExperimentAnalytics` module to read and plot the values directly in the notebook.
Initially the model consisted of 11 million parameters. After 11 iterations, the number of parameters was reduced to 270k, while accuracy increased to 91% and then started dropping after 8 pruning iteration.
This means that the best accuracy can be reached if the model has a size of about 4 million parameters, while shrinking model size about 3x!

### Additional: run iterative model pruning with custom rule
In the previous example, we have seen that accuracy drops when the model has less than 22 million parameters. Clearly, we want to stop our experiment once we reach this point. We can define a custom rule that returns `True` if the accuracy drops by a certain percentage. You can find an example implementation in `custom_rule/check_accuracy.py`. Before we can use the rule we have to define a custom rule configuration:
```python
from sagemaker.debugger import Rule, CollectionConfig, rule_configs
check_accuracy_rule = Rule.custom(
name='CheckAccuracy',
image_uri='759209512951.dkr.ecr.us-west-2.amazonaws.com/sagemaker-debugger-rule-evaluator:latest',
instance_type='ml.c4.xlarge',
volume_size_in_gb=400,
source='custom_rule/check_accuracy.py',
rule_to_invoke='check_accuracy',
rule_parameters={"previous_accuracy": "0.0",
"threshold": "0.05",
"predictions": "CrossEntropyLoss_0_input_0",
"labels":"CrossEntropyLoss_0_input_1"},
)
```
The rule reads the inputs to the loss function, which are the model predictions and the labels. It computes the accuracy and returns `True` if its value has dropped by more than 5% otherwise `False`.
In each pruning iteration, we need to pass the accuracy of the previous training job to the rule, which can be retrieved via the `ExperimentAnalytics` module.
```python
from sagemaker.analytics import ExperimentAnalytics
trial_component_analytics = ExperimentAnalytics(experiment_name=experiment_name)
accuracy = trial_component_analytics.dataframe()['scalar/accuracy_EVAL - Max'][0]
```
And overwrite the value in the rule configuration:
```python
check_accuracy_rule.rule_parameters["previous_accuracy"] = str(accuracy)
```
In the PyTorch estimator we need to add the argument `rules = [check_accuracy_rule]`.
We can create a CloudWatch alarm and use a Lambda function to stop the training. Detailed instructions can be found [here](https://github.com/awslabs/amazon-sagemaker-examples/tree/master/sagemaker-debugger/tensorflow_action_on_rule). In each iteration we check the job status and if the previous job has been stopped, we exit the loop:
```python
job_name = estimator.latest_training_job.name
client = estimator.sagemaker_session.sagemaker_client
description = client.describe_training_job(TrainingJobName=job_name)
if description['TrainingJobStatus'] == 'Stopped':
break
```
| github_jupyter |
Deep Learning with TensorFlow
=============
Credits: Forked from [TensorFlow](https://github.com/tensorflow/tensorflow) by Google
Setup
------------
Refer to the [setup instructions](https://github.com/donnemartin/data-science-ipython-notebooks/tree/feature/deep-learning/deep-learning/tensor-flow-exercises/README.md).
Exercise 2
------------
Previously in `1_notmnist.ipynb`, we created a pickle with formatted datasets for training, development and testing on the [notMNIST dataset](http://yaroslavvb.blogspot.com/2011/09/notmnist-dataset.html).
The goal of this exercise is to progressively train deeper and more accurate models using TensorFlow.
```
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
import cPickle as pickle
import numpy as np
import tensorflow as tf
```
First reload the data we generated in `1_notmist.ipynb`.
```
pickle_file = 'notMNIST.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset = save['train_dataset']
train_labels = save['train_labels']
valid_dataset = save['valid_dataset']
valid_labels = save['valid_labels']
test_dataset = save['test_dataset']
test_labels = save['test_labels']
del save # hint to help gc free up memory
print 'Training set', train_dataset.shape, train_labels.shape
print 'Validation set', valid_dataset.shape, valid_labels.shape
print 'Test set', test_dataset.shape, test_labels.shape
```
Reformat into a shape that's more adapted to the models we're going to train:
- data as a flat matrix,
- labels as float 1-hot encodings.
```
image_size = 28
num_labels = 10
def reformat(dataset, labels):
dataset = dataset.reshape((-1, image_size * image_size)).astype(np.float32)
# Map 0 to [1.0, 0.0, 0.0 ...], 1 to [0.0, 1.0, 0.0 ...]
labels = (np.arange(num_labels) == labels[:,None]).astype(np.float32)
return dataset, labels
train_dataset, train_labels = reformat(train_dataset, train_labels)
valid_dataset, valid_labels = reformat(valid_dataset, valid_labels)
test_dataset, test_labels = reformat(test_dataset, test_labels)
print 'Training set', train_dataset.shape, train_labels.shape
print 'Validation set', valid_dataset.shape, valid_labels.shape
print 'Test set', test_dataset.shape, test_labels.shape
```
We're first going to train a multinomial logistic regression using simple gradient descent.
TensorFlow works like this:
* First you describe the computation that you want to see performed: what the inputs, the variables, and the operations look like. These get created as nodes over a computation graph. This description is all contained within the block below:
with graph.as_default():
...
* Then you can run the operations on this graph as many times as you want by calling `session.run()`, providing it outputs to fetch from the graph that get returned. This runtime operation is all contained in the block below:
with tf.Session(graph=graph) as session:
...
Let's load all the data into TensorFlow and build the computation graph corresponding to our training:
```
# With gradient descent training, even this much data is prohibitive.
# Subset the training data for faster turnaround.
train_subset = 10000
graph = tf.Graph()
with graph.as_default():
# Input data.
# Load the training, validation and test data into constants that are
# attached to the graph.
tf_train_dataset = tf.constant(train_dataset[:train_subset, :])
tf_train_labels = tf.constant(train_labels[:train_subset])
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
# These are the parameters that we are going to be training. The weight
# matrix will be initialized using random valued following a (truncated)
# normal distribution. The biases get initialized to zero.
weights = tf.Variable(
tf.truncated_normal([image_size * image_size, num_labels]))
biases = tf.Variable(tf.zeros([num_labels]))
# Training computation.
# We multiply the inputs with the weight matrix, and add biases. We compute
# the softmax and cross-entropy (it's one operation in TensorFlow, because
# it's very common, and it can be optimized). We take the average of this
# cross-entropy across all training examples: that's our loss.
logits = tf.matmul(tf_train_dataset, weights) + biases
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
# We are going to find the minimum of this loss using gradient descent.
optimizer = tf.train.GradientDescentOptimizer(0.5).minimize(loss)
# Predictions for the training, validation, and test data.
# These are not part of training, but merely here so that we can report
# accuracy figures as we train.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(
tf.matmul(tf_valid_dataset, weights) + biases)
test_prediction = tf.nn.softmax(tf.matmul(tf_test_dataset, weights) + biases)
```
Let's run this computation and iterate:
```
num_steps = 801
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1))
/ predictions.shape[0])
with tf.Session(graph=graph) as session:
# This is a one-time operation which ensures the parameters get initialized as
# we described in the graph: random weights for the matrix, zeros for the
# biases.
tf.initialize_all_variables().run()
print 'Initialized'
for step in xrange(num_steps):
# Run the computations. We tell .run() that we want to run the optimizer,
# and get the loss value and the training predictions returned as numpy
# arrays.
_, l, predictions = session.run([optimizer, loss, train_prediction])
if (step % 100 == 0):
print 'Loss at step', step, ':', l
print 'Training accuracy: %.1f%%' % accuracy(
predictions, train_labels[:train_subset, :])
# Calling .eval() on valid_prediction is basically like calling run(), but
# just to get that one numpy array. Note that it recomputes all its graph
# dependencies.
print 'Validation accuracy: %.1f%%' % accuracy(
valid_prediction.eval(), valid_labels)
print 'Test accuracy: %.1f%%' % accuracy(test_prediction.eval(), test_labels)
```
Let's now switch to stochastic gradient descent training instead, which is much faster.
The graph will be similar, except that instead of holding all the training data into a constant node, we create a `Placeholder` node which will be fed actual data at every call of `sesion.run()`.
```
batch_size = 128
graph = tf.Graph()
with graph.as_default():
# Input data. For the training data, we use a placeholder that will be fed
# at run time with a training minibatch.
tf_train_dataset = tf.placeholder(tf.float32,
shape=(batch_size, image_size * image_size))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
weights = tf.Variable(
tf.truncated_normal([image_size * image_size, num_labels]))
biases = tf.Variable(tf.zeros([num_labels]))
# Training computation.
logits = tf.matmul(tf_train_dataset, weights) + biases
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.5).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(
tf.matmul(tf_valid_dataset, weights) + biases)
test_prediction = tf.nn.softmax(tf.matmul(tf_test_dataset, weights) + biases)
```
Let's run it:
```
num_steps = 3001
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
print "Initialized"
for step in xrange(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = train_dataset[offset:(offset + batch_size), :]
batch_labels = train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print "Minibatch loss at step", step, ":", l
print "Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels)
print "Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels)
print "Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels)
```
---
Problem
-------
Turn the logistic regression example with SGD into a 1-hidden layer neural network with rectified linear units (nn.relu()) and 1024 hidden nodes. This model should improve your validation / test accuracy.
---
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#@title MIT License
#
# Copyright (c) 2017 François Chollet
#
# Permission is hereby granted, free of charge, to any person obtaining a
# copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
```
# 첫 번째 신경망 훈련하기: 기초적인 분류 문제
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/keras/basic_classification"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />TensorFlow.org에서 보기</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/ko/tutorials/keras/basic_classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/ko/tutorials/keras/basic_classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />깃허브(GitHub) 소스 보기</a>
</td>
</table>
Note: 이 문서는 텐서플로 커뮤니티에서 번역했습니다. 커뮤니티 번역 활동의 특성상 정확한 번역과 최신 내용을 반영하기 위해 노력함에도
불구하고 [공식 영문 문서](https://www.tensorflow.org/?hl=en)의 내용과 일치하지 않을 수 있습니다.
이 번역에 개선할 부분이 있다면
[tensorflow/docs](https://github.com/tensorflow/docs) 깃헙 저장소로 풀 리퀘스트를 보내주시기 바랍니다.
문서 번역이나 리뷰에 지원하려면 [이 양식](https://bit.ly/tf-translate)을
작성하거나
[docs@tensorflow.org](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs)로
메일을 보내주시기 바랍니다.
이 튜토리얼에서는 운동화나 셔츠 같은 옷 이미지를 분류하는 신경망 모델을 훈련합니다. 상세 내용을 모두 이해하지 못해도 괜찮습니다. 여기서는 완전한 텐서플로(TensorFlow) 프로그램을 빠르게 살펴 보겠습니다. 자세한 내용은 앞으로 배우면서 더 설명합니다.
여기에서는 텐서플로 모델을 만들고 훈련할 수 있는 고수준 API인 [tf.keras](https://www.tensorflow.org/guide/keras)를 사용합니다.
```
from __future__ import absolute_import, division, print_function, unicode_literals
# tensorflow와 tf.keras를 임포트합니다
import tensorflow as tf
from tensorflow import keras
# 헬퍼(helper) 라이브러리를 임포트합니다
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
```
## 패션 MNIST 데이터셋 임포트하기
10개의 범주(category)와 70,000개의 흑백 이미지로 구성된 [패션 MNIST](https://github.com/zalandoresearch/fashion-mnist) 데이터셋을 사용하겠습니다. 이미지는 해상도(28x28 픽셀)가 낮고 다음처럼 개별 옷 품목을 나타냅니다:
<table>
<tr><td>
<img src="https://tensorflow.org/images/fashion-mnist-sprite.png"
alt="Fashion MNIST sprite" width="600">
</td></tr>
<tr><td align="center">
<b>그림 1.</b> <a href="https://github.com/zalandoresearch/fashion-mnist">패션-MNIST 샘플</a> (Zalando, MIT License).<br/>
</td></tr>
</table>
패션 MNIST는 컴퓨터 비전 분야의 "Hello, World" 프로그램격인 고전 [MNIST](http://yann.lecun.com/exdb/mnist/) 데이터셋을 대신해서 자주 사용됩니다. MNIST 데이터셋은 손글씨 숫자(0, 1, 2 등)의 이미지로 이루어져 있습니다. 여기서 사용하려는 옷 이미지와 동일한 포맷입니다.
패션 MNIST는 일반적인 MNIST 보다 조금 더 어려운 문제이고 다양한 예제를 만들기 위해 선택했습니다. 두 데이터셋은 비교적 작기 때문에 알고리즘의 작동 여부를 확인하기 위해 사용되곤 합니다. 코드를 테스트하고 디버깅하는 용도로 좋습니다.
네트워크를 훈련하는데 60,000개의 이미지를 사용합니다. 그다음 네트워크가 얼마나 정확하게 이미지를 분류하는지 10,000개의 이미지로 평가하겠습니다. 패션 MNIST 데이터셋은 텐서플로에서 바로 임포트하여 적재할 수 있습니다:
```
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
```
load_data() 함수를 호출하면 네 개의 넘파이(NumPy) 배열이 반환됩니다:
* `train_images`와 `train_labels` 배열은 모델 학습에 사용되는 *훈련 세트*입니다.
* `test_images`와 `test_labels` 배열은 모델 테스트에 사용되는 *테스트 세트*입니다.
이미지는 28x28 크기의 넘파이 배열이고 픽셀 값은 0과 255 사이입니다. *레이블*(label)은 0에서 9까지의 정수 배열입니다. 이 값은 이미지에 있는 옷의 *클래스*(class)를 나타냅니다:
<table>
<tr>
<th>레이블</th>
<th>클래스</th>
</tr>
<tr>
<td>0</td>
<td>T-shirt/top</td>
</tr>
<tr>
<td>1</td>
<td>Trouser</td>
</tr>
<tr>
<td>2</td>
<td>Pullover</td>
</tr>
<tr>
<td>3</td>
<td>Dress</td>
</tr>
<tr>
<td>4</td>
<td>Coat</td>
</tr>
<tr>
<td>5</td>
<td>Sandal</td>
</tr>
<tr>
<td>6</td>
<td>Shirt</td>
</tr>
<tr>
<td>7</td>
<td>Sneaker</td>
</tr>
<tr>
<td>8</td>
<td>Bag</td>
</tr>
<tr>
<td>9</td>
<td>Ankle boot</td>
</tr>
</table>
각 이미지는 하나의 레이블에 매핑되어 있습니다. 데이터셋에 *클래스 이름*이 들어있지 않기 때문에 나중에 이미지를 출력할 때 사용하기 위해 별도의 변수를 만들어 저장합니다:
```
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
```
## 데이터 탐색
모델을 훈련하기 전에 데이터셋 구조를 살펴보죠. 다음 코드는 훈련 세트에 60,000개의 이미지가 있다는 것을 보여줍니다. 각 이미지는 28x28 픽셀로 표현됩니다:
```
train_images.shape
```
비슷하게 훈련 세트에는 60,000개의 레이블이 있습니다:
```
len(train_labels)
```
각 레이블은 0과 9사이의 정수입니다:
```
train_labels
```
테스트 세트에는 10,000개의 이미지가 있습니다. 이 이미지도 28x28 픽셀로 표현됩니다:
```
test_images.shape
```
테스트 세트는 10,000개의 이미지에 대한 레이블을 가지고 있습니다:
```
len(test_labels)
```
## 데이터 전처리
네트워크를 훈련하기 전에 데이터를 전처리해야 합니다. 훈련 세트에 있는 첫 번째 이미지를 보면 픽셀 값의 범위가 0~255 사이라는 것을 알 수 있습니다:
```
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
plt.show()
```
신경망 모델에 주입하기 전에 이 값의 범위를 0~1 사이로 조정하겠습니다. 이렇게 하려면 255로 나누어야 합니다. *훈련 세트*와 *테스트 세트*를 동일한 방식으로 전처리하는 것이 중요합니다:
```
train_images = train_images / 255.0
test_images = test_images / 255.0
```
*훈련 세트*에서 처음 25개 이미지와 그 아래 클래스 이름을 출력해 보죠. 데이터 포맷이 올바른지 확인하고 네트워크 구성과 훈련할 준비를 마칩니다.
```
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
plt.show()
```
## 모델 구성
신경망 모델을 만들려면 모델의 층을 구성한 다음 모델을 컴파일합니다.
### 층 설정
신경망의 기본 구성 요소는 *층*(layer)입니다. 층은 주입된 데이터에서 표현을 추출합니다. 아마도 문제를 해결하는데 더 의미있는 표현이 추출될 것입니다.
대부분 딥러닝은 간단한 층을 연결하여 구성됩니다. `tf.keras.layers.Dense`와 같은 층들의 가중치(parameter)는 훈련하는 동안 학습됩니다.
```
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation=tf.nn.relu),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
```
이 네트워크의 첫 번째 층인 `tf.keras.layers.Flatten`은 2차원 배열(28 x 28 픽셀)의 이미지 포맷을 28 * 28 = 784 픽셀의 1차원 배열로 변환합니다. 이 층은 이미지에 있는 픽셀의 행을 펼쳐서 일렬로 늘립니다. 이 층에는 학습되는 가중치가 없고 데이터를 변환하기만 합니다.
픽셀을 펼친 후에는 두 개의 `tf.keras.layers.Dense` 층이 연속되어 연결됩니다. 이 층을 밀집 연결(densely-connected) 또는 완전 연결(fully-connected) 층이라고 부릅니다. 첫 번째 `Dense` 층은 128개의 노드(또는 뉴런)를 가집니다. 두 번째 (마지막) 층은 10개의 노드의 *소프트맥스*(softmax) 층입니다. 이 층은 10개의 확률을 반환하고 반환된 값의 전체 합은 1입니다. 각 노드는 현재 이미지가 10개 클래스 중 하나에 속할 확률을 출력합니다.
### 모델 컴파일
모델을 훈련하기 전에 필요한 몇 가지 설정이 모델 *컴파일* 단계에서 추가됩니다:
* *손실 함수*(Loss function)-훈련 하는 동안 모델의 오차를 측정합니다. 모델의 학습이 올바른 방향으로 향하도록 이 함수를 최소화해야 합니다.
* *옵티마이저*(Optimizer)-데이터와 손실 함수를 바탕으로 모델의 업데이트 방법을 결정합니다.
* *지표*(Metrics)-훈련 단계와 테스트 단계를 모니터링하기 위해 사용합니다. 다음 예에서는 올바르게 분류된 이미지의 비율인 *정확도*를 사용합니다.
```
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
```
## 모델 훈련
신경망 모델을 훈련하는 단계는 다음과 같습니다:
1. 훈련 데이터를 모델에 주입합니다-이 예에서는 `train_images`와 `train_labels` 배열입니다.
2. 모델이 이미지와 레이블을 매핑하는 방법을 배웁니다.
3. 테스트 세트에 대한 모델의 예측을 만듭니다-이 예에서는 `test_images` 배열입니다. 이 예측이 `test_labels` 배열의 레이블과 맞는지 확인합니다.
훈련을 시작하기 위해 `model.fit` 메서드를 호출하면 모델이 훈련 데이터를 학습합니다:
```
model.fit(train_images, train_labels, epochs=5)
```
모델이 훈련되면서 손실과 정확도 지표가 출력됩니다. 이 모델은 훈련 세트에서 약 0.88(88%) 정도의 정확도를 달성합니다.
## 정확도 평가
그다음 테스트 세트에서 모델의 성능을 비교합니다:
```
test_loss, test_acc = model.evaluate(test_images, test_labels)
print('테스트 정확도:', test_acc)
```
테스트 세트의 정확도가 훈련 세트의 정확도보다 조금 낮습니다. 훈련 세트의 정확도와 테스트 세트의 정확도 사이의 차이는 *과대적합*(overfitting) 때문입니다. 과대적합은 머신러닝 모델이 훈련 데이터보다 새로운 데이터에서 성능이 낮아지는 현상을 말합니다.
## 예측 만들기
훈련된 모델을 사용하여 이미지에 대한 예측을 만들 수 있습니다.
```
predictions = model.predict(test_images)
```
여기서는 테스트 세트에 있는 각 이미지의 레이블을 예측했습니다. 첫 번째 예측을 확인해 보죠:
```
predictions[0]
```
이 예측은 10개의 숫자 배열로 나타납니다. 이 값은 10개의 옷 품목에 상응하는 모델의 신뢰도(confidence)를 나타냅니다. 가장 높은 신뢰도를 가진 레이블을 찾아보죠:
```
np.argmax(predictions[0])
```
모델은 이 이미지가 앵클 부츠(`class_name[9]`)라고 가장 확신하고 있습니다. 이 값이 맞는지 테스트 레이블을 확인해 보죠:
```
test_labels[0]
```
10개의 신뢰도를 모두 그래프로 표현해 보겠습니다:
```
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
```
0번째 원소의 이미지, 예측, 신뢰도 점수 배열을 확인해 보겠습니다.
```
i = 0
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions, test_labels)
plt.show()
i = 12
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions, test_labels)
plt.show()
```
몇 개의 이미지의 예측을 출력해 보죠. 올바르게 예측된 레이블은 파란색이고 잘못 예측된 레이블은 빨강색입니다. 숫자는 예측 레이블의 신뢰도 퍼센트(100점 만점)입니다. 신뢰도 점수가 높을 때도 잘못 예측할 수 있습니다.
```
# 처음 X 개의 테스트 이미지와 예측 레이블, 진짜 레이블을 출력합니다
# 올바른 예측은 파랑색으로 잘못된 예측은 빨강색으로 나타냅니다
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions, test_labels)
plt.show()
```
마지막으로 훈련된 모델을 사용하여 한 이미지에 대한 예측을 만듭니다.
```
# 테스트 세트에서 이미지 하나를 선택합니다
img = test_images[0]
print(img.shape)
```
`tf.keras` 모델은 한 번에 샘플의 묶음 또는 *배치*(batch)로 예측을 만드는데 최적화되어 있습니다. 하나의 이미지를 사용할 때에도 2차원 배열로 만들어야 합니다:
```
# 이미지 하나만 사용할 때도 배치에 추가합니다
img = (np.expand_dims(img,0))
print(img.shape)
```
이제 이 이미지의 예측을 만듭니다:
```
predictions_single = model.predict(img)
print(predictions_single)
plot_value_array(0, predictions_single, test_labels)
_ = plt.xticks(range(10), class_names, rotation=45)
```
`model.predict`는 2차원 넘파이 배열을 반환하므로 첫 번째 이미지의 예측을 선택합니다:
```
np.argmax(predictions_single[0])
```
이전과 마찬가지로 모델의 예측은 레이블 9입니다.
| github_jupyter |
```
import numpy as np
import pandas as pd
import loompy
import velocyto as vcy
import matplotlib.pyplot as plt
import matplotlib as mpl
import copy
import igraph as ig
import louvain
import umap
import networkx
import community
import pandas as pd
import scanpy as sc
import seaborn as sns
from collections import Counter
import numpy as np
from scipy.integrate import simps
from numpy import trapz
import seaborn as sns; sns.set()
from sklearn.decomposition import PCA
import pickle
sns.set_style("dark")
cd revisions0/FinalNotebooks/
ALL_DATA = sc.read_h5ad("ALL_VITRO_TIMECOURSE_DATA_RAW.h5ad")
D00 = ALL_DATA[ALL_DATA.obs["DAY"]=="D0"].copy()
D07 = ALL_DATA[ALL_DATA.obs["DAY"]=="D7"].copy()
D14 = ALL_DATA[ALL_DATA.obs["DAY"]=="D14"].copy()
D30 = ALL_DATA[ALL_DATA.obs["DAY"]=="D30"].copy()
D38 = ALL_DATA[ALL_DATA.obs["DAY"]=="D38"].copy()
D45 = ALL_DATA[ALL_DATA.obs["DAY"]=="D45"].copy()
D60 = ALL_DATA[ALL_DATA.obs["DAY"]=="D60"].copy()
datasets = {"D00":D00, "D07":D07, "D14":D14, "D30":D30, "D38":D38, "D45":D45, "D60":D60}
all_vlm = vcy.VelocytoLoom("ALL_VITRO_TIMECOURSE_DATA_RAW.loom")
all_vlm.S.sum(0)
all_vlm._normalize_S(relative_size=all_vlm.initial_cell_size,
target_size=np.mean(all_vlm.initial_cell_size))
all_days = ["D0", "D7", "D14", "D30", "D38", "D45", "D60"]
ms_cum_dict = {"Trapezoid":{}, "Simpsons":{}, "Cumsum":{}}
cs_cum_dict = {"Trapezoid":{}, "Simpsons":{}, "Cumsum":{}}
ms_noncum_dict = {"Trapezoid":{}, "Simpsons":{}, "Cumsum":{}}
cs_noncum_dict = {"Trapezoid":{}, "Simpsons":{}, "Cumsum":{}}
res_dict = {"Trapezoid":{}, "Simpsons":{}, "Cumsum":{}}
ngs_dict = {"Trapezoid":{}, "Simpsons":{}, "Cumsum":{}}
# Get copy of normalized filtered split time point datasets
cell_filtered_days = {d:{} for d in all_days}
for curr_day in all_days:
print(curr_day)
curr_vlm_norm = copy.deepcopy(all_vlm)
cell_filter_array = np.array(curr_vlm_norm.ca["DAY"] == curr_day)
curr_vlm_norm.filter_cells(bool_array=cell_filter_array)
curr_vlm_norm.S_norm = curr_vlm_norm.S_norm[:, cell_filter_array]
cell_filtered_days[curr_day]["counts"] = curr_vlm_norm.S_norm
cell_filtered_days[curr_day]["rows"] = curr_vlm_norm.ra["Gene"]
cell_filtered_days[curr_day]["columns"] = curr_vlm_norm.ca["CellID"]
from sklearn.svm import SVR
import numpy as np
import matplotlib.pyplot as plt
def filter_cv_vs_mean(S: np.ndarray, N: int, svr_gamma: float=None, plot: bool=True, min_expr_cells: int=2,
max_expr_avg: float=20, min_expr_avg: float=0) -> np.ndarray:
muS = S.mean(1)
detected_bool = ((S > 0).sum(1) > min_expr_cells) & (muS < max_expr_avg) & (muS > min_expr_avg)
Sf = S[detected_bool, :]
mu = Sf.mean(1)
sigma = Sf.std(1, ddof=1)
cv = sigma / mu
log_m = np.log2(mu)
log_cv = np.log2(cv)
if svr_gamma is None:
svr_gamma = 150. / len(mu)
svr = SVR(gamma=svr_gamma)
svr.fit(log_m[:, None], log_cv)
fitted_fun = svr.predict
ff = fitted_fun(log_m[:, None])
score = log_cv - ff
xnew = np.linspace(np.min(log_m), np.max(log_m))
ynew = svr.predict(xnew[:, None])
nth_score = np.sort(score)[::-1][N]
if plot:
plt.scatter(log_m[score > nth_score], log_cv[score > nth_score], s=3, alpha=0.4, c="tab:red")
plt.scatter(log_m[score <= nth_score], log_cv[score <= nth_score], s=3, alpha=0.4, c="tab:blue")
mu_linspace = np.linspace(np.min(log_m), np.max(log_m))
plt.plot(mu_linspace, fitted_fun(mu_linspace[:, None]), c="k")
plt.xlabel("log2 mean S")
plt.ylabel("log2 CV S")
cv_mean_score = np.zeros(detected_bool.shape)
cv_mean_score[~detected_bool] = np.min(score) - 1e-16
cv_mean_score[detected_bool] = score
cv_mean_selected = cv_mean_score >= nth_score
return cv_mean_selected, cv_mean_score
# Get copy of scores separate time point unnormalized datasets
for key in datasets.keys():
print(key)
ds = datasets[key]
sc.pp.normalize_total(ds)
cv_mean_selected, cv_mean_score = filter_cv_vs_mean(ds.X.T.toarray(), N=2000, max_expr_avg=50)
ds.var["cv_mean_score"] = cv_mean_score
ds.var["cv_mean_selected"] = cv_mean_selected
datasets[key] = ds
```
# CV score theshold changes, trapezoid rule AUC
```
THRESHOLD = 0.3
# Different CV score thresholds for keeping genes
print("THRESHOLD =", THRESHOLD)
ngs = []
selected_genes = {}
for key in datasets.keys():
curr_vlm = datasets[key]
keep_genes = list(curr_vlm.var.index[curr_vlm.var["cv_mean_score"] > THRESHOLD])
ngs.append(keep_genes)
selected_genes.update({g: True for g in keep_genes})
print(len(selected_genes))
for i in ngs:
print(len(i))
ms_cum = []
cs_cum = []
ms_noncum = []
cs_noncum = []
# Get AUC for each individual time point
NUM_PCS = 1000
for curr_day in all_days:
counts = cell_filtered_days[curr_day]["counts"]
genes = cell_filtered_days[curr_day]["rows"]
cells = cell_filtered_days[curr_day]["columns"]
gene_filter_array = np.array([g in selected_genes for g in genes])
np.random.seed(0)
subset = {c:True for c in np.random.choice(cells, NUM_PCS, replace=False)}
cell_filter_array = np.array([i in subset for i in cells])
counts = counts[gene_filter_array, :][:, cell_filter_array]
pca = PCA()
pcs = pca.fit_transform(counts.T)
evr = pca.explained_variance_ratio_
# Cumulative sum AUC
measured_cum = trapz(np.cumsum(evr))
control_cum = trapz([i/NUM_PCS for i in range(0, NUM_PCS)]) # control
# Non-cumulative sum AUC
measured_noncum = trapz(evr)
control_noncum = trapz([1/NUM_PCS]*NUM_PCS) # control
print(curr_day, measured_cum, control_cum, measured_noncum, control_noncum)
ms_cum.append(measured_cum)
cs_cum.append(control_cum)
ms_noncum.append(measured_noncum)
cs_noncum.append(control_noncum)
ms_cum_dict["Trapezoid"][THRESHOLD] = ms_cum
cs_cum_dict["Trapezoid"][THRESHOLD] = cs_cum
ms_noncum_dict["Trapezoid"][THRESHOLD] = ms_noncum
cs_noncum_dict["Trapezoid"][THRESHOLD] = cs_noncum
ngs_dict["Trapezoid"][THRESHOLD] = ngs
thresholds = list(ms_cum_dict["Trapezoid"].keys())
thresholds
plt.figure(None, (8, 6))
for thres in thresholds:
ms = ms_cum_dict["Trapezoid"][thres]
cs = cs_cum_dict["Trapezoid"][thres]
plt.plot(range(0, 7), np.array(ms) - np.array(cs), label=thres)
plt.xticks(range(0, 7), labels=all_days)
plt.xlabel("Differentiation Day")
plt.ylabel("Cumulative Area Under Curve (AUC)")
plt.title("Cumulative AUC during differentiation time course, 1K cells")
plt.legend(loc=1)
#plt.savefig("PCA variance AUC during differentiation time course, 700 cells.png", dpi=300)
ms_cum_dict["TrapezoidGSS_Final"] = {}
cs_cum_dict["TrapezoidGSS_Final"] = {}
ms_noncum_dict["TrapezoidGSS_Final"] = {}
cs_noncum_dict["TrapezoidGSS_Final"] = {}
ngs_dict["TrapezoidGSS_Final"] = {}
print("THRESHOLD =", THRESHOLD)
ngs = []
selected_genes = {}
for key in datasets.keys():
curr_vlm = datasets[key]
keep_genes = list(curr_vlm.var.index[curr_vlm.var["cv_mean_score"] > THRESHOLD])
ngs.append(keep_genes)
selected_genes.update({g: True for g in keep_genes})
print(len(selected_genes))
ms_cum = {d:[] for d in all_days}
cs_cum = {d:[] for d in all_days}
ms_noncum = {d:[] for d in all_days}
cs_noncum = {d:[] for d in all_days}
# Get AUC for each individual time point
NUM_CELLS = 800
NUM_GENES = int(round(len(selected_genes)/2))
NUM_ITERS = 100
print(NUM_CELLS, NUM_GENES, NUM_ITERS)
for curr_day in all_days:
for i in range(0, NUM_ITERS):
counts = cell_filtered_days[curr_day]["counts"]
genes = cell_filtered_days[curr_day]["rows"]
cells = cell_filtered_days[curr_day]["columns"]
gene_subset = np.random.choice(list(selected_genes.keys()), NUM_GENES, replace=False)
gene_filter_array = np.array([i in gene_subset for i in genes])
cell_subset = {c:True for c in np.random.choice(cells, NUM_CELLS, replace=False)}
cell_filter_array = np.array([i in cell_subset for i in cells])
counts = counts[gene_filter_array, :][:, cell_filter_array]
pca = PCA()
pcs = pca.fit_transform(counts.T)
evr = pca.explained_variance_ratio_
# Cumulative sum AUC
assert(NUM_CELLS <= NUM_GENES)
NUM_PCS = min(NUM_CELLS, NUM_GENES)
measured_cum = trapz(np.cumsum(evr))
control_cum = trapz([i/NUM_PCS for i in range(0, NUM_PCS)]) # control
# Non-cumulative sum AUC
measured_noncum = trapz(evr)
control_noncum = trapz([1/NUM_PCS]*NUM_PCS) # control
if i%100==0:
print("ITER =", i, curr_day, measured_cum, control_cum, measured_noncum, control_noncum)
ms_cum[curr_day].append(measured_cum)
cs_cum[curr_day].append(control_cum)
ms_noncum[curr_day].append(measured_noncum)
cs_noncum[curr_day].append(control_noncum)
ms_cum_dict["TrapezoidGSS_Final"][THRESHOLD] = ms_cum
cs_cum_dict["TrapezoidGSS_Final"][THRESHOLD] = cs_cum
ms_noncum_dict["TrapezoidGSS_Final"][THRESHOLD] = ms_noncum
cs_noncum_dict["TrapezoidGSS_Final"][THRESHOLD] = cs_noncum
ngs_dict["TrapezoidGSS_Final"][THRESHOLD] = ngs
#AUC 800 cells, 1K iter, thres=0.3
sc.settings.set_figure_params(vector_friendly=True)
sc.settings.set_figure_params(dpi=120)
sns.set_style("dark")
ms = ms_cum_dict["TrapezoidGSS_Final"][0.3]
cs = cs_cum_dict["TrapezoidGSS_Final"][0.3]
ds = []
for day in all_days:
ds.append(np.array(ms[day]) - np.array(cs[day]))
parts = plt.violinplot(ds, showmeans=True)
for i in range(0, len(parts['bodies'])):
pc = parts["bodies"][i]
c = ['#1f77b4', '#ff7f0e', '#279e68', '#d62728', '#aa40fc', '#8c564b', '#e377c2'][i]
pc.set_color(c)
plt.xticks(range(1, 8), labels=all_days)
plt.xlabel("Differentiation Day")
plt.ylabel("Cumulative AUC")
plt.title("AUC of Principal Component Variance")
plt.tight_layout()
plt.show()
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Загрузка данных NumPy
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/load_data/numpy"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Смотрите на TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/ru/tutorials/load_data/numpy.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Запустите в Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/ru/tutorials/load_data/numpy.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Изучайте код на GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/ru/tutorials/load_data/numpy.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Скачайте ноутбук</a>
</td>
</table>
Note: Вся информация в этом разделе переведена с помощью русскоговорящего Tensorflow сообщества на общественных началах. Поскольку этот перевод не является официальным, мы не гарантируем что он на 100% аккуратен и соответствует [официальной документации на английском языке](https://www.tensorflow.org/?hl=en). Если у вас есть предложение как исправить этот перевод, мы будем очень рады увидеть pull request в [tensorflow/docs](https://github.com/tensorflow/docs) репозиторий GitHub. Если вы хотите помочь сделать документацию по Tensorflow лучше (сделать сам перевод или проверить перевод подготовленный кем-то другим), напишите нам на [docs-ru@tensorflow.org list](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ru).
В этом руководстве приведен пример загрузки данных из массивов NumPy в `tf.data.Dataset`.
Этот пример загружает датасет MNIST из файла `.npz`. Однако источник массивов NumPy не важен.
## Setup
```
try:
# %tensorflow_version существует только в Colab.
%tensorflow_version 2.x
except Exception:
pass
from __future__ import absolute_import, division, print_function, unicode_literals
import numpy as np
import tensorflow as tf
```
### Load from `.npz` file
```
DATA_URL = 'https://storage.googleapis.com/tensorflow/tf-keras-datasets/mnist.npz'
path = tf.keras.utils.get_file('mnist.npz', DATA_URL)
with np.load(path) as data:
train_examples = data['x_train']
train_labels = data['y_train']
test_examples = data['x_test']
test_labels = data['y_test']
```
## Загрузите массивы NumPy с `tf.data.Dataset`
Представьте что у вас есть массив примеров и соответствующий массив меток, передайте эти два массива кортежом в `tf.data.Dataset.from_tensor_slices` чтобы создать `tf.data.Dataset`.
```
train_dataset = tf.data.Dataset.from_tensor_slices((train_examples, train_labels))
test_dataset = tf.data.Dataset.from_tensor_slices((test_examples, test_labels))
```
## Используйте датасеты
### Перемешайте датасеты и разбейте их на пакеты
```
BATCH_SIZE = 64
SHUFFLE_BUFFER_SIZE = 100
train_dataset = train_dataset.shuffle(SHUFFLE_BUFFER_SIZE).batch(BATCH_SIZE)
test_dataset = test_dataset.batch(BATCH_SIZE)
```
### Постройте и обучите модель
```
model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer=tf.keras.optimizers.RMSprop(),
loss=tf.keras.losses.SparseCategoricalCrossentropy(),
metrics=[tf.keras.metrics.SparseCategoricalAccuracy()])
model.fit(train_dataset, epochs=10)
model.evaluate(test_dataset)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.